code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Let's work with the collection of state CSVs
# +
import csv
import numpy as np
import pandas as pd
from pandas import (DataFrame, Series)
# +
import glob
glob.glob("census_2010_sf1/state*")
# +
# use http://pandas.pydata.org/pandas-docs/stable/generated/pandas.read_csv.html instead
# of DataFrame.from_csv to use dtype
df = pd.read_csv("census_2010_sf1/state_population.csv", dtype={'FIPS': str})
df.head()
# -
# use some of the pre-written code
from census_api_utils import entropy
df = pd.read_csv("census_2010_sf1/state_diversity_measures.csv", dtype={'FIPS': str})
df.head()
# https://docs.scipy.org/doc/numpy/reference/generated/numpy.testing.assert_array_almost_equal.html
np.testing.assert_array_almost_equal(
df[["White", "Black", "Asian", "Hispanic", "Other"]].apply(entropy, axis=1),
df.entropy5)
df.sort('entropy5', ascending=False).head()
| 05_Examples_Calculations_with_CSVs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/jamestheengineer/data-science-from-scratch-Python/blob/master/Chapter_22.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="pxsJpb6Al8qY"
# !git clone https://github.com/jamestheengineer/data-science-from-scratch-Python.git
# %cd data-science-from-scratch-Python/
# !pip install import-ipynb
import import_ipynb
# + id="dkXYd2_PWkcn"
from typing import NamedTuple
class User(NamedTuple):
id: int
name: str
users = [User(0, "Hero"), User(1, "Dunn"), User(2, "Sue"), User(3, "Chi"),
User(4, "Thor"), User(5, "Clive"), User(6, "Hicks"),
User(7, "Devin"), User(8, "Kate"), User(9, "Klein")]
friend_pairs = [(0, 1), (0, 2), (1, 2), (1, 3), (2, 3), (3, 4),
(4, 5), (5, 6), (5, 7), (6, 8), (7, 8), (8, 9)]
from typing import Dict, List
# type alias for keeping track ofr Friendships
Friendships = Dict[int, List[int]]
friendships: Friendships = {user.id: [] for user in users}
for i, j in friend_pairs:
friendships[i].append(j)
friendships[j].append(i)
assert friendships[4] == [3, 5]
assert friendships[8] == [6,7,9]
# + id="N2SOEIZAXeLC"
from collections import deque
Path = List[int]
def shortest_paths_from(from_user_id: int,
friendships: Friendships) -> Dict[int, List[Path]]:
# A dictionary from user_id to *all* shortest paths to that user.
shortest_paths_to: Dict[int, List[Path]] = {from_user_id: [[]]}
# A queue of (previous user, next user) that we need to check.
# Starts out with all pairs(from_user, friend_of_from_user).
frontier = deque((from_user_id, friend_id)
for friend_id in friendships[from_user_id])
# Keep going until we empty the queue
while frontier:
# Remove the pair that's next in the queue.
prev_user_id, user_id = frontier.popleft()
# Because of the way we're adding to the queue,
# necessarily we already know some shortest paths to prev_user.
paths_to_prev_user = shortest_paths_to[prev_user_id]
new_paths_to_user = [path + [user_id] for path in paths_to_prev_user]
# It's possible we already know a shortest path to user_id.
old_paths_to_user = shortest_paths_to.get(user_id, [])
# What's the shortest path to here that we've seen so far?
if old_paths_to_user:
min_path_length = len(old_paths_to_user[0])
else:
min_path_length = float('inf')
# Only keep paths that aren't too long and are actually new.
new_paths_to_user = [path
for path in new_paths_to_user
if len(path) <= min_path_length
and path not in old_paths_to_user]
shortest_paths_to[user_id] = old_paths_to_user + new_paths_to_user
# Add never seen neighbors to the frontier.
frontier.extend((user_id, friend_id)
for friend_id in friendships[user_id]
if friend_id not in shortest_paths_to)
return shortest_paths_to
# + id="zzhU7rfogwbl"
# For each from_user, for each to_user, a list of shortest paths
shortest_paths = {user.id: shortest_paths_from(user.id, friendships)
for user in users}
# + id="eSNBQ2ATQG_g"
betweenness_centrality = {user.id: 0.0 for user in users}
for source in users:
for target_id, paths in shortest_paths[source.id].items():
if source.id < target_id: # don't double count
num_paths = len(paths)
contrib = 1 / num_paths
for path in paths:
for between_id in path:
if between_id not in [source.id, target_id]:
betweenness_centrality[between_id] += contrib
print(betweenness_centrality)
# + id="xYRF8G1TQ7-j"
# Closeness centrality
def farness(user_id: int) -> float:
"""The sum of the lengths of the shortest paths to each other user"""
return sum(len(paths[0])
for paths in shortest_paths[user_id].values())
closeness_centrality = {user.id: 1 / farness(user.id) for user in users}
print(closeness_centrality)
# + id="zXBsWyDKUmzM"
# Computing shortest paths is kinda a pain, thus not used much on large networks. Usually, we use eigenvector centrality
from Chapter_04 import Matrix, make_matrix, shape
def matrix_times_matrix(m1: Matrix, m2: Matrix) -> Matrix:
nr1, nc1 = shape(m1)
nr2, nc2 = shape(m2)
assert nc1 == nr2, "must have (# of columns in m1) == (# of rows in m2)"
def entry_fn(i: int, j: int) -> float:
"""dot product of i-th row of m1 with j-th column of m2"""
return sum(m1[i][k] * m2[k][j] for k in range(nc1))
return make_matrix(nr1, nc2, entry_fn)
from Chapter_04 import Vector, dot
def matrix_times_vector(m: Matrix, v: Vector) -> Vector:
nr, nc = shape(m)
n = len(v)
assert nc == n, "must hae (# cols in m) == (# elements in v)"
return [dot(row, v) for row in m] # output has length nr
from typing import Tuple
import random
from Chapter_04 import magnitude, distance
def find_eigenvector(m: Matrix,
tolerance: float = 0.00001) -> Tuple[Vector, float]:
guess = [random.random() for _ in m]
while True:
result = matrix_times_vector(m, guess) # transform guess
norm = magnitude(result) # compute norm
next_guess = [x / norm for x in result] # rescale
if distance(guess, next_guess) < tolerance:
# convergence so return (eigenvector, eigenvalue)
return next_guess, norm
guess = next_guess
def entry_fn(i: int, j: int):
return 1 if (i,j) in friend_pairs or (j,i) in friend_pairs else 0
n = len(users)
adjacency_matrix = make_matrix(n,n, entry_fn)
[x,y] = find_eigenvector(adjacency_matrix)
print(x,y)
# + id="uk085k0_lDRa"
# Directed graphs and pagerank
endorsements = [(0, 1), (1, 0), (0, 2), (2, 0), (1, 2),
(2, 1), (1, 3), (2, 3), (3, 4), (5, 4),
(5, 6), (7, 5), (6, 8), (8, 7), (8, 9)]
from collections import Counter
endorsement_counts = Counter(target for source, target in endorsements)
import tqdm
def page_rank(users: List[User],
endorsements: List[Tuple[int, int]],
damping: float = 0.85,
num_iters: int = 100) -> Dict[int, float]:
# Compute how many people each person endorses
outgoing_counts = Counter(target for source, target in endorsements)
print(outgoing_counts)
# Initially distribute PageRank evenly
num_users = len(users)
pr = {user.id : 1 / num_users for user in users}
# Small fraction of PageRank that each node gets each iteration
base_pr = (1 - damping) / num_users
for iter in tqdm.trange(num_iters):
next_pr = {user.id : base_pr for user in users} # start with base_pr
for source, target in endorsements:
# Add damped fraction of source pr to target
next_pr[target] += damping * pr[source] / outgoing_counts[source]
pr = next_pr
return pr
# + id="6Bf733HthnHO" outputId="1286ce26-2b54-45ab-ffa6-f656ad08e360" colab={"base_uri": "https://localhost:8080/", "height": 68}
pr = page_rank(users, endorsements)
print(pr)
assert pr[4] > max(page_rank for user_id, page_rank in pr.items() if user_id != 4)
# + id="Mfom-XNMidx_" outputId="c4a96e02-6061-4032-884d-f6bdc696620c" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Thor (user_id 4) has higher page rank than anyone else
assert pr[4] > max(page_rank
for user_id, page_rank in pr.items()
if user_id != 4)
print(pr)
# + id="8VOfWGNAkkmj"
| Chapter_22.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import nannyml as nml
import pandas as pd
reference, analysis, analysis_gt = nml.datasets.load_synthetic_binary_classification_dataset()
display(reference.head(3))
# -
reference.salary_range.value_counts()
metadata = nml.extract_metadata(data = reference, model_name='wfh_predictor', model_type='classification_binary', exclude_columns=['identifier'])
metadata.is_complete()
metadata.target_column_name = 'work_home_actual'
metadata.is_complete()
metadata.to_df()
print(metadata.to_df().to_markdown(tablefmt="grid"))
| docs/example_notebooks/Dataset Binary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # part 6 - Synonyms of Tags
# To create the synonyms table, we will use wordnet to conduct the operation.The package we use is wordnet.
from nltk.corpus import wordnet as wn
import pandas as pd
# import tag information as dataframe
twitterTag = pd.read_csv("Final_Data/twitterTag.csv",encoding = 'latin-1')
youtubeData = pd.read_csv("Final_Data/youtubeData_clean.csv",encoding = 'latin-1')
# show tag structure
twitterTag[0:1000:10]
tag=twitterTag['tag']
tag[0:1000:100]
# +
#syns_list=[]
#syns = wordnet.synsets("program")
#syns_list.append(syns)
# -
# print all the existing synsets of tags in the form of csv
tag_synonyms= pd.DataFrame(columns=['tag', 'synonyms','2','3','4','5'])
times=1
for i in range(0,2524):
#syns_list=[]
syns = wordnet.synsets(tag[i])
if(syns!=[]):
print(i,end=',')
print(tag[i],end=',')
for j in syns:
print(j.name(),end=',')
print('')
# Then we copy those data into csv tables and conduct the initial data cleaning.
# # Data Cleaning Part-1
# import the roughly cleaned datasets and remove duplicates and fill null values.
twitterSynonyms = pd.read_csv("Final_Data/synonym1.csv",encoding = 'utf-8')
# check current data
twitterSynonyms[0:1000:100]
# create a function to remove irrelevant symbols
def normalization(synonym):
for r in range(0,729):
if (synonym[r]!='none'):
synonym[r]=str(synonym[r])
location=synonym[r].find('.')
if(location!=-1):
synonym[r]=synonym[r][0:location]
# We will apply these function to each column of the data frame and output a csv table to avoid repeatedly calling this time consuming function.
normalization(twitterSynonyms['synonym_1'])
normalization(twitterSynonyms['synonym_2'])
normalization(twitterSynonyms['synonym_3'])
normalization(twitterSynonyms['synonym_4'])
normalization(twitterSynonyms['synonym_5'])
twitterSynonyms.to_csv('Final_Data/synonym2.csv')
# # Data Cleaning Part-2
# import the csv we output in the previous part
twitterSynonyms=pd.read_csv('Final_Data/synonym2.csv',index_col=0)
#check the data
twitterSynonyms[0:1000:100]
# Now we will be fill the null value and drop the duplicated value of the synonym dataframe.
#
# Also, we need to refresh the dataframe's index value to prevent error.
twitterSynonyms=twitterSynonyms.fillna('none')
twitterSynonyms=twitterSynonyms.drop_duplicates()
twitterSynonyms.index=range(len(twitterSynonyms))
#!!!! this function will be manully repeat(change column name),you can't not only apply it once, change parameter and do it over again
# aim is to avoid repeat value as the former data display shows.(e.g first row)
# here I have manualy changed the value in twitterSynonyms['synonym_3'] to twitterSynonyms['synonym_2'] .etc.
#The aim is to cover the similar value in column with the after element.
tag= twitterSynonyms['synonym_3']
synonym= twitterSynonyms['synonym_4']
synonym2=twitterSynonyms['synonym_5']
for r in range(0,459):
if (synonym[r]==tag[r]):
synonym[r]=synonym2[r]
twitterSynonyms[0:1000:100]
twitterSynonyms['synonym_4'].value_counts()[0:10]
# Noticing that most value in the 5h column is none, not to mention 6th column, we will delete these 2 columns.
del twitterSynonyms['synonym_4']
del twitterSynonyms['synonym_5']
#output the data again to operate a next-stage data cleaning in excel
twitterSynonyms.to_csv('Final_Data/synonym3.csv')
#read the table we just out put
twitterSynonyms=pd.read_csv('Final_Data/synonym3.csv',index_col=0)
# change the index value
twitterSynonyms.index=range(len(twitterSynonyms))
# check the final data
twitterSynonyms
# Import the new data to database
# Ruijia connect databse
import pandas as pd
from sqlalchemy import create_engine
# Ruijia connect databse
engine=create_engine('mysql+mysqldb://root:fbfs1stuacc@localhost:3306/final_project?charset=latin1')
twitterSynonyms.to_sql(name='synonym',con=engine,if_exists='replace')
import MySQLdb
db=MySQLdb.connect("localhost","root","fbfs1stuacc","final_project")
cursor=db.cursor()
cursor.execute("""
select * from synonym
limit 20
""")
print(cursor.fetchall())
| part 6- Synonyms Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/tancik/fourier-feature-networks/blob/master/Experiments/1d_regression.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="pE8BiDAW0lVt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="caf8b3a6-54fc-410b-ed17-356f57ba82d5"
# !pip install -qq neural_tangents==0.2.2
# + id="Wcn1yDT30vMF" colab_type="code" colab={}
import jax
from jax import random, grad, jit, vmap
from jax.config import config
from jax.lib import xla_bridge
import jax.numpy as np
import neural_tangents as nt
from neural_tangents import stax
from jax.experimental import optimizers
import os
import matplotlib
import matplotlib.pylab as pylab
import matplotlib.pyplot as plt
from matplotlib.lines import Line2D
from tqdm.notebook import tqdm as tqdm
import time
import numpy as onp
# + id="gnC7achEw60P" colab_type="code" colab={}
# Utils
fplot = lambda x : np.fft.fftshift(np.log10(np.abs(np.fft.fft(x))))
# Signal makers
def sample_random_signal(key, decay_vec):
N = decay_vec.shape[0]
raw = random.normal(key, [N, 2]) @ np.array([1, 1j])
signal_f = raw * decay_vec
signal = np.real(np.fft.ifft(signal_f))
return signal
def sample_random_powerlaw(key, N, power):
coords = np.float32(np.fft.ifftshift(1 + N//2 - np.abs(np.fft.fftshift(np.arange(N)) - N//2)))
decay_vec = coords ** -power
decay_vec = onp.array(decay_vec)
decay_vec[N//4:] = 0
return sample_random_signal(key, decay_vec)
# Network
def make_network(num_layers, num_channels, ntk_params=True, num_outputs=1):
layers = []
for i in range(num_layers-1):
if ntk_params:
layers.append(stax.Dense(num_channels, parameterization='standard'))
else:
layers.append(stax.Dense(num_channels, parameterization='standard'))
layers.append(stax.Relu(do_backprop=True))
layers.append(stax.Dense(num_outputs, parameterization='standard'))
return stax.serial(*layers)
# Encoding
def compute_ntk(x, avals, bvals, kernel_fn):
x1_enc = input_encoder(x, avals, bvals)
x2_enc = input_encoder(np.array([0.], dtype=np.float32), avals, bvals)
out = np.squeeze(kernel_fn(x1_enc, x2_enc, 'ntk'))
return out
input_encoder = lambda x, a, b: np.concatenate([a * np.sin((2.*np.pi*x[...,None]) * b),
a * np.cos((2.*np.pi*x[...,None]) * b)], axis=-1) / np.linalg.norm(a)
def predict_psnr_basic(kernel_fn, train_fx, test_fx, train_x, train_y, test_x, test_y, t_final, eta=None):
g_dd = kernel_fn(train_x, train_x, 'ntk')
g_td = kernel_fn(test_x, train_x, 'ntk')
train_predict_fn = nt.predict.gradient_descent_mse(g_dd, train_y[...,None], g_td)
train_theory_y, test_theory_y = train_predict_fn(t_final, train_fx[...,None], test_fx[...,None])
calc_psnr = lambda f, g: -10. * np.log10(np.mean((f-g)**2))
return calc_psnr(test_y, test_theory_y[:,0]), calc_psnr(train_y, train_theory_y[:,0])
predict_psnr_basic = jit(predict_psnr_basic, static_argnums=(0,))
def train_model(rand_key, network_size, lr, iters,
train_input, test_input, test_mask, optimizer, ab, name=''):
if ab is None:
ntk_params = False
else:
ntk_params = True
init_fn, apply_fn, kernel_fn = make_network(*network_size, ntk_params=ntk_params)
if ab is None:
run_model = jit(lambda params, ab, x: np.squeeze(apply_fn(params, x[...,None] - .5)))
else:
run_model = jit(lambda params, ab, x: np.squeeze(apply_fn(params, input_encoder(x, *ab))))
model_loss = jit(lambda params, ab, x, y: .5 * np.sum((run_model(params, ab, x) - y) ** 2))
model_psnr = jit(lambda params, ab, x, y: -10 * np.log10(np.mean((run_model(params, ab, x) - y) ** 2)))
model_grad_loss = jit(lambda params, ab, x, y: jax.grad(model_loss)(params, ab, x, y))
opt_init, opt_update, get_params = optimizer(lr)
opt_update = jit(opt_update)
if ab is None:
_, params = init_fn(rand_key, (-1, 1))
else:
_, params = init_fn(rand_key, (-1, input_encoder(train_input[0], *ab).shape[-1]))
opt_state = opt_init(params)
pred0 = run_model(get_params(opt_state), ab, test_input[0])
pred0_f = np.fft.fft(pred0)
train_psnrs = []
test_psnrs = []
theories = []
xs = []
errs = []
for i in tqdm(range(iters), desc=name):
opt_state = opt_update(i, model_grad_loss(get_params(opt_state), ab, *train_input), opt_state)
if i % 20 == 0:
train_psnr = model_psnr(get_params(opt_state), ab, *train_input)
test_psnr = model_psnr(get_params(opt_state), ab, test_input[0][test_mask], test_input[1][test_mask])
if ab is None:
train_fx = run_model(get_params(opt_state), ab, train_input[0])
test_fx = run_model(get_params(opt_state), ab, test_input[0][test_mask])
theory = predict_psnr_basic(kernel_fn, train_fx, test_fx, train_input[0][...,None]-.5, train_input[1], test_input[0][test_mask][...,None], test_input[1][test_mask], i*lr)
else:
test_x = input_encoder(test_input[0][test_mask], *ab)
train_x = input_encoder(train_input[0], *ab)
train_fx = run_model(get_params(opt_state), ab, train_input[0])
test_fx = run_model(get_params(opt_state), ab, test_input[0][test_mask])
theory = predict_psnr_basic(kernel_fn, train_fx, test_fx, train_x, train_input[1], test_x, test_input[1][test_mask], i*lr)
train_psnrs.append(train_psnr)
test_psnrs.append(test_psnr)
theories.append(theory)
pred = run_model(get_params(opt_state), ab, train_input[0])
errs.append(pred - train_input[1])
xs.append(i)
return get_params(opt_state), train_psnrs, test_psnrs, errs, np.array(theories), xs
# + [markdown] id="MXMEEOql0_Hn" colab_type="text"
# # Make fig 2
# + id="tWMfr1H50_jd" colab_type="code" colab={}
N_train = 32
data_power = 1
network_size = (4, 1024)
learning_rate = 1e-5
sgd_iters = 50001
# + id="TjOhNefO1C5z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 414, "referenced_widgets": ["958e2c696e6e40b7990717cf2d9b5c19", "820da148a1644d67b68c123a2d80085c", "948bb76079de420a81978221ad89b153", "332a533d1df240ae8a895fbda284e6b6", "99b0a159dadf4436b19463819bbcd7f4", "0849582555764ae1a38007fd5ce6008b", "acb028b70e3b4e40b78db78f75338df5", "<KEY>", "<KEY>", "98ad0622b53b4bb4be39178351c0e520", "<KEY>", "e7368ed6802f486592333443221a7441", "<KEY>", "<KEY>", "415fa3c0246446b88eda63112b5dafe7", "<KEY>", "3c6e31ec064e4e588611c49cf7f6e552", "<KEY>", "2e9d70f319604168b119062e9661ae43", "<KEY>", "f9660eeaf88a40ba9eec974935cb4b05", "be93709dc1a94a8c8298cf3a1454e734", "d18914187f574a12a58de8c7003b666e", "<KEY>", "33298a3f247441178f567eb45d99a624", "<KEY>", "bba323c0d8de42beab6e0fa061680e7c", "<KEY>", "b44d67a65b00463e9b06ed1c8282f5b7", "c3af70b17eb843d0869bd0241c32af68", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "8ef062845e844c3982e0914af1902d5f", "13fafc1c60af433b941ddd44a86e791c", "<KEY>", "<KEY>", "72ee4f39626a4d98829405c802b4614a", "772d63ec6557410ab945c3703d0215e2", "<KEY>", "<KEY>", "<KEY>", "7edc5de5591d4524a081141daad8eada", "<KEY>", "<KEY>", "<KEY>", "145f0588df734a378818d50d0e3f8d34", "<KEY>", "ac17850207f54051be736886f4ea8dea", "<KEY>", "<KEY>", "<KEY>", "<KEY>"]} outputId="40a49daf-b9f0-44a4-9a00-e2949667e5e2"
rand_key = random.PRNGKey(0)
config.update('jax_disable_jit', False)
# Signal
M = 8
N = N_train
x_test = np.float32(np.linspace(0,1.,N*M,endpoint=False))
x_train = x_test[::M]
test_mask = onp.ones(len(x_test), onp.bool)
test_mask[np.arange(0,x_test.shape[0],M)] = 0
s = sample_random_powerlaw(rand_key, N*M, data_power)
s = (s-s.min()) / (s.max()-s.min()) - .5
# Kernels
bvals = np.float32(np.arange(1, N//2+1))
ab_dict = {}
# ab_dict = {r'$p = {}$'.format(p) : (bvals**-np.float32(p), bvals) for p in [0, 1]}
ab_dict = {r'$p = {}$'.format(p) : (bvals**-np.float32(p), bvals) for p in [0, 0.5, 1, 1.5, 2]}
ab_dict[r'$p = \infty$'] = (np.eye(bvals.shape[0])[0], bvals)
ab_dict['No mapping'] = None
# Train the networks
rand_key, *ensemble_key = random.split(rand_key, 1 + len(ab_dict))
outputs = {k : train_model(key, network_size, learning_rate, sgd_iters,
(x_train, s[::M]), (x_test, s), test_mask,
optimizer=optimizers.sgd, ab=ab_dict[k], name=k) for k, key in zip(ab_dict, ensemble_key)}
ab_dict.update({r'$p = {}$'.format(p) : (bvals**-np.float32(p), bvals) for p in [0.5, 1.5, 2]})
# + [markdown] id="tV6zZaI2t6T9" colab_type="text"
# # Fig 2
# + id="WYhgkC9v8hqM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 367} outputId="51649bc1-c96d-4a2c-8107-cdbb82dd7620"
prop_cycle = plt.rcParams['axes.prop_cycle']
colors = prop_cycle.by_key()['color']
params = {'legend.fontsize': 24,
'axes.labelsize': 22,
'axes.titlesize': 26,
'xtick.labelsize':20,
'ytick.labelsize':20}
pylab.rcParams.update(params)
matplotlib.rcParams['mathtext.fontset'] = 'cm'
matplotlib.rcParams['mathtext.rm'] = 'serif'
plt.rcParams["font.family"] = "cmr10"
names = ['$p = 0$',
'$p = 0.5$',
'$p = 1$',
'$p = 1.5$',
'$p = 2$',
'$p = \\infty$']
N_kernels = len(names)
colors_k = np.array([[0.8872, 0.4281, 0.1875],
[0.8136, 0.6844, 0.0696],
[0.2634, 0.6634, 0.4134],
[0.0943, 0.5937, 0.8793],
[0.3936, 0.2946, 0.6330],
[0.7123, 0.2705, 0.3795]])
linewidth = 3
line_alpha = .8
title_offset = -0.3
xs = outputs[names[0]][-1]
t_final = learning_rate * sgd_iters
init_fn, apply_fn, kernel_fn = make_network(*network_size)
run_model = jit(lambda params, ab, x: np.squeeze(apply_fn(params, input_encoder(x, *ab))))
fig3 = plt.figure(constrained_layout=True, figsize=(22,4))
gs = fig3.add_gridspec(1, 4, width_ratios=[1,1,1.3,1.3])
### Plot NTK stuff
H_rows = {k : compute_ntk(x_train, *ab_dict[k], kernel_fn) for k in names}
samples = 100
x_no_encoding = np.linspace(-np.pi, np.pi, samples)
x_basic = np.stack([np.sin(x_no_encoding),np.cos(x_no_encoding)], axis=-1)
relu_NTK = kernel_fn(x_no_encoding[:,None], x_no_encoding[:,None], 'ntk')
basic_NTK = kernel_fn(x_basic, x_basic, 'ntk')
ax = fig3.add_subplot(gs[0, 0])
ax.imshow(relu_NTK, cmap='inferno', extent=[-.5,.5,.5,-.5])
ax.xaxis.tick_top()
extent = [-.5,.5]
ax.set_xticks([-.5,.5])
ax.set_yticks([-.5,.5])
ax.set_xticklabels([fr'${t:g}$' for t in extent])
ax.set_yticklabels([fr'${t:g}$' for t in extent])
xtick = ax.get_xticks()
ax.set_xticks(xtick)
ax.set_xticklabels([fr'${t:g}$' for t in xtick])
ax.set_title('(a) No mapping NTK', y=title_offset)
ax = fig3.add_subplot(gs[0, 1])
ax.imshow(basic_NTK, cmap='inferno', extent=[-.5,.5,.5,-.5])
ax.xaxis.tick_top()
ax.set_xticks([-.5,.5])
ax.set_yticks([-.5,.5])
ax.set_xticklabels([fr'${t:g}$' for t in extent])
ax.set_yticklabels([fr'${t:g}$' for t in extent])
ax.set_title('(b) Basic mapping NTK', y=title_offset)
ax = fig3.add_subplot(gs[0, 2])
for c, k in zip(colors_k, H_rows):
ntk_spatial = np.fft.fftshift(H_rows[k])
ax.plot(np.linspace(-.5, .5, 33, endpoint=True), np.append(ntk_spatial, ntk_spatial[0]), label=k, color=c, alpha=line_alpha, linewidth=linewidth)
ax.set_title('(c) NTK spatial', y=title_offset)
xtick = ax.get_xticks()
ax.set_xticks(xtick)
ax.set_xticklabels([fr'${t:g}$' for t in xtick])
plt.grid(True, which='both', alpha=.3)
plt.autoscale(enable=True, axis='x', tight=True)
ax = fig3.add_subplot(gs[0, 3])
for c, k in zip(colors_k, H_rows):
ntk_spectrum = 10**fplot(H_rows[k])
plt.semilogy(np.append(ntk_spectrum, ntk_spectrum[0]), label=k, color=c, alpha=line_alpha, linewidth=linewidth)
ax.set_title('(d) NTK Fourier spectrum', y=title_offset)
plt.xticks([0,8,16,24,32], ['$-\pi$','$-\pi/2$','$0$','$\pi/2$','$\pi$'])
plt.autoscale(enable=True, axis='x', tight=True)
plt.grid(True, which='major', alpha=.3)
plt.legend(loc='center left', bbox_to_anchor=(1,.5), handlelength=1)
plt.savefig('1D_fig2.pdf', bbox_inches='tight', pad_inches=0)
plt.show()
# + [markdown] id="qafttdJwNRLa" colab_type="text"
# # Fig 3
# + id="7ALq8TDYt_3G" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 727} outputId="09b378d3-06e6-43d8-8d18-ca866227eff7"
import matplotlib.patches as patches
prop_cycle = plt.rcParams['axes.prop_cycle']
colors = prop_cycle.by_key()['color']
params = {'legend.fontsize': 20,
'axes.labelsize': 18,
'axes.titlesize': 22,
'xtick.labelsize':16,
'ytick.labelsize':16}
pylab.rcParams.update(params)
matplotlib.rcParams['mathtext.fontset'] = 'cm'
matplotlib.rcParams['mathtext.rm'] = 'serif'
plt.rcParams["font.family"] = "cmr10"
names = list(outputs.keys())
names = [n for n in names if 'opt' not in n]
names = [
'$p = 0$',
# '$p = 0.5$',
'$p = 1$',
# '$p = 1.5$',
# '$p = 2$',
'$p = \\infty$',
'No mapping'
]
N_kernels = len(names)
colors_k = np.array([[0.8872, 0.4281, 0.1875],
# [0.8136, 0.6844, 0.0696],
[0.2634, 0.6634, 0.4134],
[0.0943, 0.5937, 0.8793],
[0.3936, 0.2946, 0.6330],
[0.7123, 0.2705, 0.3795]])
linewidth = 4
line_alpha = .7
title_offset = -0.2
xs = outputs[names[0]][-1]
t_final = learning_rate * sgd_iters
_, apply_fn, kernel_fn = make_network(*network_size)
_, apply_fn_standard, kernel_fn_standard = make_network(*network_size, ntk_params=False)
run_model = jit(lambda params, ab, x: np.squeeze(apply_fn(params, input_encoder(x, *ab))))
fig3 = plt.figure(constrained_layout=True, figsize=(20,9))
gs = fig3.add_gridspec(6, 8)
H_rows = {k : compute_ntk(x_train, *ab_dict[k], kernel_fn) for k in names if k not in 'No mapping'}
### Plot learned fns
max_size = x_test.shape[0]
i0, i1 = 0.5, .75 ### These numbers are arbitrary, just taking a slice
i0 = (int(max_size * i0)//M)*M
i1 = (int(max_size * i1)//M)*M
ax = fig3.add_subplot(gs[0:4, 0:6])
for c, k in zip(colors_k, names):
params = outputs[k][0]
if k in 'No mapping':
ax.plot(x_test[i0:i1], apply_fn_standard(params, x_test[..., None]-.5)[i0:i1], label=k, color=c, linewidth=linewidth, alpha=line_alpha, zorder=1)
else:
ax.plot(x_test[i0:i1], run_model(params, ab_dict[k], x_test)[i0:i1], label=k, color=c, linewidth=linewidth, alpha=line_alpha, zorder=1)
plt.plot(x_test[i0:i1], s[i0:i1], label='Target signal', color='k', linewidth=linewidth, alpha=line_alpha, zorder=1)
plt.autoscale(enable=True, axis='x', tight=True)
ax.scatter(x_train[i0//M:i1//M], s[i0:i1:M], color='w', edgecolors='k', linewidths=2, s=150, linewidth=3, label='Training points', zorder=2)
ax.set_title('(a) Final learned functions', y=title_offset+.08)
ax.set_xticks([])
ax.set_yticks([])
ax.legend(loc='lower left', ncol=2)
plt.xlabel('$x$')
plt.ylabel('$y$')
plt.grid(True, which='both', alpha=.3)
### Plot loss curves
xvals = xs # np.log10(np.array(xs))
ax = fig3.add_subplot(gs[3:6, 6:8])
for c, k in zip(colors_k, names):
ax.semilogy(xvals, 10**(np.array(outputs[k][1])/-10), color=c, label=k, alpha=line_alpha, linewidth=linewidth)
# if k in 'no encoding':
# continue
ax.semilogy(xvals, 10**(np.array(outputs[k][4][:,1])/-10), '--', color=c, alpha=line_alpha, linewidth=linewidth)
plt.autoscale(enable=True, axis='x', tight=True)
ax.set_xticks([0,10000,20000,30000,40000,50000])
ax.set_xticklabels(['$0$','','','','','$50000$'])
plt.xlabel('Iteration', labelpad=-13)
plt.ylabel('Mean squared error')
plt.xlabel('Iteration')
ax.set_title('(d) Train loss', y=title_offset)
ax.set_ylim(top=.1)
plt.grid(True, which='major', alpha=.3)
custom_lines = [Line2D([], [], color='gray', linestyle='--', linewidth=linewidth),
Line2D([], [], color='gray', linestyle='-', linewidth=linewidth)]
ax.legend(custom_lines, ['Theory', 'Observed'], loc='lower left', ncol=1, framealpha=.95, handlelength=1.75)
ax = fig3.add_subplot(gs[0:3, 6:8])
for c, k in zip(colors_k, names):
ax.semilogy(xvals, 10**(np.array(outputs[k][2])/-10), color=c, label=k, alpha=line_alpha, linewidth=linewidth)
ax.semilogy(xvals, 10**(np.array(outputs[k][4][:,0])/-10), '--', color=c, alpha=line_alpha, linewidth=linewidth)
plt.autoscale(enable=True, axis='x', tight=True)
ax.set_xticks([0,10000,20000,30000,40000,50000])
ax.set_xticklabels(['$0$','','','','','$50000$'])
plt.xlabel('Iteration', labelpad=-13)
plt.ylabel('Mean squared error')
plt.xlabel('Iteration')
ax.set_title('(b) Test loss', y=title_offset)
ax.set_yticklabels([], minor=True)
plt.grid(True, which='major', alpha=.3)
### Plot freq components
letters = ['f','g','h']
prefix = ['Low', 'Mid', 'High']
for i, j in enumerate([1, 5, 15]):
err_lines = {k : np.abs(np.fft.fft(np.array(outputs[k][3]))[:,j]) for k in names}
ax = fig3.add_subplot(gs[4:6, 0+2*i:0+2*i+2])
for c, k in zip(colors_k, err_lines):
ax.semilogy(xs, err_lines[k], label=k, color=c, alpha=line_alpha, linewidth=linewidth)
plt.autoscale(enable=True, axis='x', tight=True)
props = dict(boxstyle='round,pad=0.35,rounding_size=.1', facecolor='white', edgecolor='gray', linewidth=.3, alpha=0.95)
plt.text(0.5, 0.87, prefix[i], horizontalalignment='center',
verticalalignment='center', transform=ax.transAxes, bbox=props, fontsize=18)
if i == 0:
plt.ylabel('Absolute Error')
if i == 1:
ax.set_title(f'(c) Train loss frequency components', y=title_offset-.08)
ax.set_xticks([0,10000,20000,30000,40000,50000])
ax.set_xticklabels(['$0$','','','','','$50000$'])
plt.xlabel('Iteration', labelpad=-13)
ax.set_yticklabels([], minor=True)
plt.grid(True, which='major', alpha=.3)
plt.savefig('1D_fig2_5.pdf', bbox_inches='tight', pad_inches=0)
plt.show()
# + [markdown] id="E9YFDzniOAlq" colab_type="text"
# # Supp (fig 3 expanded)
# + id="_UdvsJSiATNp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 727} outputId="b111796a-ce92-4cc6-a3ff-710bb1d8ea41"
import matplotlib.patches as patches
prop_cycle = plt.rcParams['axes.prop_cycle']
colors = prop_cycle.by_key()['color']
params = {'legend.fontsize': 20,
'axes.labelsize': 18,
'axes.titlesize': 22,
'xtick.labelsize':16,
'ytick.labelsize':16}
pylab.rcParams.update(params)
matplotlib.rcParams['mathtext.fontset'] = 'cm'
matplotlib.rcParams['mathtext.rm'] = 'serif'
plt.rcParams["font.family"] = "cmr10"
names = list(outputs.keys())
names = [n for n in names if 'opt' not in n]
names = [
'$p = 0$',
'$p = 0.5$',
'$p = 1$',
'$p = 1.5$',
'$p = 2$',
'$p = \\infty$',
'No mapping'
]
N_kernels = len(names)
colors_k = np.array([[0.8872, 0.4281, 0.1875],
# [0.8136, 0.6844, 0.0696],
[0.2634, 0.6634, 0.4134],
[0.0943, 0.5937, 0.8793],
[0.3936, 0.2946, 0.6330],
[0.7123, 0.2705, 0.3795]])
linewidth = 4
line_alpha = .7
title_offset = -0.2
xs = outputs[names[0]][-1]
t_final = learning_rate * sgd_iters
_, apply_fn, kernel_fn = make_network(*network_size)
_, apply_fn_standard, kernel_fn_standard = make_network(*network_size, ntk_params=False)
run_model = jit(lambda params, ab, x: np.squeeze(apply_fn(params, input_encoder(x, *ab))))
fig3 = plt.figure(constrained_layout=True, figsize=(20,9))
gs = fig3.add_gridspec(6, 8)
H_rows = {k : compute_ntk(x_train, *ab_dict[k], kernel_fn) for k in names if k not in 'No mapping'}
### Plot learned fns
max_size = x_test.shape[0]
i0, i1 = 0.5, .75 ### These numbers are arbitrary, just taking a slice
i0 = (int(max_size * i0)//M)*M
i1 = (int(max_size * i1)//M)*M
ax = fig3.add_subplot(gs[0:4, 0:6])
for c, k in zip(colors_k, names):
params = outputs[k][0]
if k in 'No mapping':
ax.plot(x_test[i0:i1], apply_fn_standard(params, x_test[..., None]-.5)[i0:i1], label=k, color=c, linewidth=linewidth, alpha=line_alpha, zorder=1)
else:
ax.plot(x_test[i0:i1], run_model(params, ab_dict[k], x_test)[i0:i1], label=k, color=c, linewidth=linewidth, alpha=line_alpha, zorder=1)
plt.plot(x_test[i0:i1], s[i0:i1], label='Target signal', color='k', linewidth=linewidth, alpha=line_alpha, zorder=1)
plt.autoscale(enable=True, axis='x', tight=True)
ax.scatter(x_train[i0//M:i1//M], s[i0:i1:M], color='w', edgecolors='k', linewidths=2, s=150, linewidth=3, label='Training points', zorder=2)
ax.set_title('(a) Final learned functions', y=title_offset+.08)
ax.set_xticks([])
ax.set_yticks([])
ax.legend(loc='lower left', ncol=2)
plt.xlabel('$x$')
plt.ylabel('$y$')
plt.grid(True, which='both', alpha=.3)
### Plot loss curves
xvals = xs # np.log10(np.array(xs))
ax = fig3.add_subplot(gs[3:6, 6:8])
for c, k in zip(colors_k, names):
ax.semilogy(xvals, 10**(np.array(outputs[k][1])/-10), color=c, label=k, alpha=line_alpha, linewidth=linewidth)
# if k in 'no encoding':
# continue
ax.semilogy(xvals, 10**(np.array(outputs[k][4][:,1])/-10), '--', color=c, alpha=line_alpha, linewidth=linewidth)
plt.autoscale(enable=True, axis='x', tight=True)
ax.set_xticks([0,10000,20000,30000,40000,50000])
ax.set_xticklabels(['$0$','','','','','$50000$'])
plt.xlabel('Iteration', labelpad=-13)
plt.ylabel('Mean squared error')
plt.xlabel('Iteration')
ax.set_title('(d) Train loss', y=title_offset)
ax.set_ylim(top=.1)
plt.grid(True, which='major', alpha=.3)
custom_lines = [Line2D([], [], color='gray', linestyle='--', linewidth=linewidth),
Line2D([], [], color='gray', linestyle='-', linewidth=linewidth)]
ax.legend(custom_lines, ['Theory', 'Observed'], loc='lower left', ncol=1, framealpha=.95, handlelength=1.75)
ax = fig3.add_subplot(gs[0:3, 6:8])
for c, k in zip(colors_k, names):
ax.semilogy(xvals, 10**(np.array(outputs[k][2])/-10), color=c, label=k, alpha=line_alpha, linewidth=linewidth)
ax.semilogy(xvals, 10**(np.array(outputs[k][4][:,0])/-10), '--', color=c, alpha=line_alpha, linewidth=linewidth)
plt.autoscale(enable=True, axis='x', tight=True)
ax.set_xticks([0,10000,20000,30000,40000,50000])
ax.set_xticklabels(['$0$','','','','','$50000$'])
plt.xlabel('Iteration', labelpad=-13)
plt.ylabel('Mean squared error')
plt.xlabel('Iteration')
ax.set_title('(b) Test loss', y=title_offset)
ax.set_yticklabels([], minor=True)
plt.grid(True, which='major', alpha=.3)
### Plot freq components
letters = ['f','g','h']
prefix = ['Low', 'Mid', 'High']
for i, j in enumerate([1, 5, 15]):
err_lines = {k : np.abs(np.fft.fft(np.array(outputs[k][3]))[:,j]) for k in names}
ax = fig3.add_subplot(gs[4:6, 0+2*i:0+2*i+2])
for c, k in zip(colors_k, err_lines):
ax.semilogy(xs, err_lines[k], label=k, color=c, alpha=line_alpha, linewidth=linewidth)
plt.autoscale(enable=True, axis='x', tight=True)
props = dict(boxstyle='round,pad=0.35,rounding_size=.1', facecolor='white', edgecolor='gray', linewidth=.3, alpha=0.95)
plt.text(0.5, 0.87, prefix[i], horizontalalignment='center',
verticalalignment='center', transform=ax.transAxes, bbox=props, fontsize=18)
if i == 0:
plt.ylabel('Absolute Error')
if i == 1:
ax.set_title(f'(c) Train loss frequency components', y=title_offset-.08)
ax.set_xticks([0,10000,20000,30000,40000,50000])
ax.set_xticklabels(['$0$','','','','','$50000$'])
plt.xlabel('Iteration', labelpad=-13)
ax.set_yticklabels([], minor=True)
plt.grid(True, which='major', alpha=.3)
plt.savefig('1D_fig_supp.pdf', bbox_inches='tight', pad_inches=0)
plt.show()
# + id="jl98aFkC1L03" colab_type="code" colab={}
| Experiments/1d_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ConvNet Stacked Bidirectional LSTM for Sentiment Classification
# This ConvNet Stacked Bi-LSTM performs sentiment analysis on the IMDB review dataset.
import keras
from keras.datasets import imdb
from keras.preprocessing.sequence import pad_sequences
from keras.models import Sequential
from keras.layers import Conv1D, MaxPooling1D, LSTM, Embedding, Dense, Flatten, Dropout, Activation, SpatialDropout1D
from keras.layers.wrappers import Bidirectional
from keras.callbacks import ModelCheckpoint
import os
from sklearn.metrics import roc_auc_score
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# #### Set Hyperparameters
# +
output_dir = 'model_output/cnnLstm'
epochs = 4
batch_size = 128
n_dim = 64
n_unique_words = 10000
max_review_length = 200
pad_type = trunc_type = 'pre'
drop_embed = 0.2
n_conv = 64
k_conv = 3
mp_size = 4
n_lstm = 64
drop_lstm = 0.2
# -
# #### Load Data
(X_train, y_train), (X_valid, y_valid) = imdb.load_data(num_words=n_unique_words)
# #### Preprocess Data
X_train = pad_sequences(X_train, maxlen=max_review_length, padding=pad_type, truncating=trunc_type, value=0)
X_valid = pad_sequences(X_valid, maxlen=max_review_length, padding=pad_type, truncating=trunc_type, value=0)
# #### Design Deep Net Architecture
# +
model = Sequential()
model.add(Embedding(n_unique_words, n_dim, input_length=max_review_length))
model.add(SpatialDropout1D(drop_embed))
model.add(Conv1D(n_conv, k_conv, activation='relu'))
model.add(MaxPooling1D(mp_size))
model.add(Bidirectional(LSTM(n_lstm, dropout=drop_lstm)))
model.add(Dense(1, activation='sigmoid'))
# -
model.summary()
# #### Configure the Model
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
modelcheckpoint = ModelCheckpoint(filepath=output_dir+'/weights.{epoch:02d}.hdf5')
if not os.path.exists(output_dir):
os.makedirs(output_dir)
# ### Train the Model
model.fit(X_train, y_train, batch_size=batch_size, epochs=epochs, verbose=1, validation_split=0.3)#, callbacks=[modelcheckpoint])
# #### Evaluate
#model.load_weights(output_dir+'/weights.03.hdf5')
print('Not loading weights.')
y_hat = model.predict_proba(X_valid)
y_hat[0]
plt.hist(y_hat)
_ = plt.axvline(x=0.5, color='orange')
pct_auc = roc_auc_score(y_valid, y_hat) * 100
'{:0.2f}'.format(pct_auc)
| notebooks/nlp/conv_stacked_bidirectional_lstm_in_keras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Collaboration and Competition
#
# ---
#
# Congratulations for completing the third project of the [Deep Reinforcement Learning Nanodegree](https://www.udacity.com/course/deep-reinforcement-learning-nanodegree--nd893) program! In this notebook, you will learn how to control agents in a more challenging environment, where the goal is to train a team of agents to play soccer. **Note that this exercise is optional!**
#
# ### 1. Start the Environment
#
# We begin by importing the necessary packages. If the code cell below returns an error, please revisit the project instructions to double-check that you have installed [Unity ML-Agents](https://github.com/Unity-Technologies/ml-agents/blob/master/docs/Installation.md) and [NumPy](http://www.numpy.org/).
from unityagents import UnityEnvironment
import numpy as np
# Next, we will start the environment! **_Before running the code cell below_**, change the `file_name` parameter to match the location of the Unity environment that you downloaded.
#
# - **Mac**: `"path/to/Soccer.app"`
# - **Windows** (x86): `"path/to/Soccer_Windows_x86/Soccer.exe"`
# - **Windows** (x86_64): `"path/to/Soccer_Windows_x86_64/Soccer.exe"`
# - **Linux** (x86): `"path/to/Soccer_Linux/Soccer.x86"`
# - **Linux** (x86_64): `"path/to/Soccer_Linux/Soccer.x86_64"`
# - **Linux** (x86, headless): `"path/to/Soccer_Linux_NoVis/Soccer.x86"`
# - **Linux** (x86_64, headless): `"path/to/Soccer_Linux_NoVis/Soccer.x86_64"`
#
# For instance, if you are using a Mac, then you downloaded `Soccer.app`. If this file is in the same folder as the notebook, then the line below should appear as follows:
# ```
# env = UnityEnvironment(file_name="Soccer.app")
# ```
env = UnityEnvironment(file_name="...")
# Environments contain **_brains_** which are responsible for deciding the actions of their associated agents. Here we obtain separate brains for the striker and goalie agents.
# +
# print the brain names
print(env.brain_names)
# set the goalie brain
g_brain_name = env.brain_names[0]
g_brain = env.brains[g_brain_name]
# set the striker brain
s_brain_name = env.brain_names[1]
s_brain = env.brains[s_brain_name]
# -
# ### 2. Examine the State and Action Spaces
#
# Run the code cell below to print some information about the environment.
# +
# reset the environment
env_info = env.reset(train_mode=True)
# number of agents
num_g_agents = len(env_info[g_brain_name].agents)
print('Number of goalie agents:', num_g_agents)
num_s_agents = len(env_info[s_brain_name].agents)
print('Number of striker agents:', num_s_agents)
# number of actions
g_action_size = g_brain.vector_action_space_size
print('Number of goalie actions:', g_action_size)
s_action_size = s_brain.vector_action_space_size
print('Number of striker actions:', s_action_size)
# examine the state space
g_states = env_info[g_brain_name].vector_observations
g_state_size = g_states.shape[1]
print('There are {} goalie agents. Each receives a state with length: {}'.format(g_states.shape[0], g_state_size))
s_states = env_info[s_brain_name].vector_observations
s_state_size = s_states.shape[1]
print('There are {} striker agents. Each receives a state with length: {}'.format(s_states.shape[0], s_state_size))
# -
# ### 3. Take Random Actions in the Environment
#
# In the next code cell, you will learn how to use the Python API to control the agents and receive feedback from the environment.
#
# Once this cell is executed, you will watch the agents' performance, if they select actions at random with each time step. A window should pop up that allows you to observe the agents.
#
# Of course, as part of the project, you'll have to change the code so that the agents are able to use their experiences to gradually choose better actions when interacting with the environment!
for i in range(2): # play game for 2 episodes
env_info = env.reset(train_mode=False) # reset the environment
g_states = env_info[g_brain_name].vector_observations # get initial state (goalies)
s_states = env_info[s_brain_name].vector_observations # get initial state (strikers)
g_scores = np.zeros(num_g_agents) # initialize the score (goalies)
s_scores = np.zeros(num_s_agents) # initialize the score (strikers)
while True:
# select actions and send to environment
g_actions = np.random.randint(g_action_size, size=num_g_agents)
s_actions = np.random.randint(s_action_size, size=num_s_agents)
actions = dict(zip([g_brain_name, s_brain_name],
[g_actions, s_actions]))
env_info = env.step(actions)
# get next states
g_next_states = env_info[g_brain_name].vector_observations
s_next_states = env_info[s_brain_name].vector_observations
# get reward and update scores
g_rewards = env_info[g_brain_name].rewards
s_rewards = env_info[s_brain_name].rewards
g_scores += g_rewards
s_scores += s_rewards
# check if episode finished
done = np.any(env_info[g_brain_name].local_done)
# roll over states to next time step
g_states = g_next_states
s_states = s_next_states
# exit loop if episode finished
if done:
break
print('Scores from episode {}: {} (goalies), {} (strikers)'.format(i+1, g_scores, s_scores))
# When finished, you can close the environment.
env.close()
# ### 4. It's Your Turn!
#
# Now it's your turn to train your own agent to solve the environment! When training the environment, set `train_mode=True`, so that the line for resetting the environment looks like the following:
# ```python
# env_info = env.reset(train_mode=True)[brain_name]
# ```
| p3_collab-compet/Soccer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import pandas as pd
import numpy as np
import subprocess
import os
os.chdir('../..')
data_dir='derivatives/Tensor_ICA'
import re
import nilearn.image as image
f=[]
for root,directories,filenames in os.walk(data_dir):
for x in filenames:
if "zscore" in x:
f.append(os.path.join(root,x))
sub=[]
for file in f:
sub.append(re.search('Tensor_ICA/sub-(.*)/preproc_task-', file).group(1))
sub=list(set(sub))
list_of_final_files=[]
for sub in sub:
print('sub-%s'%(sub))
mylist=[s for s in f if 'sub-%s'%(sub) in s]
mylist.sort()
if len(mylist)==4:
mylist.sort(key=lambda x: x[x.find('stim-'):])
print(mylist)
full_rest_filename="derivatives/Tensor_ICA/sub-%s/sub-%s_fullrest-stim-BRTSHVL.nii.gz"%(sub,sub)
list_of_final_files.append(full_rest_filename)
full_rest=image.concat_imgs(mylist)
full_rest.to_filename(full_rest_filename)
df=pd.DataFrame(list_of_final_files)
df.to_csv("concatonated_rest_files.csv",sep=',',index=False)
f.sort(key=lambda x: x[x.find('stim-'):])
print type(f)
np.savetxt("zimages_all4stims.csv", f, delimiter=",", fmt='%s')
| code/Tensor_Analysis/Concat_those_zimages.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from celery_test import func
stockcodes=['300251',]
date=['2016-02-29',]
p1=list(range(150,250,10))
p2=list(range(50,100,10))
p3=list(range(150,250,10))
p4=list(range(20,40,5))
func(stockcodes,date,p1,p2,p3,p4)
date=('2016-02-29',)
p1=range(150,250,10)
p2=range(50,100,10)
p3=range(150,250,10)
p4=range(20,40,5)
func(stockcodes,date,p1,p2,p3,p4)
date
p5=list(range(150,250,10))
for i in p5:
print(i)
| celerybacktest/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from sklearn import tree
from sklearn.datasets import load_iris
import numpy as np
from sklearn import preprocessing
import pandas as pd
clf = tree.DecisionTreeClassifier()
filename = "credit.txt"
data = np.genfromtxt(filename, delimiter=None,dtype=str)
print(data)
df = pd.DataFrame(data[1:9])
df.columns=data[0]
print(df)
print(df.columns)
# one_hot_data = pd.get_dummies(df.columns,drop_first=False)
# print(one_hot_data)
# print(df['Risk'])
# clf.fit(one_hot_data, df['Risk'])
#data = pd.DataFrame()
#df = pd.read_fwf(filename)
#data = pd.read_csv(filename, sep=" ", header=None)
#print(df)
#data.columns = ["a", "b", "c", "etc."]
# # Name Debt Income Married? Owns_Property Gender Risk
# # Tim low low no no male low
# # Joe high high yes yes male low
# # Sue low high yes no female low
# # John medium low no no male high
# # Mary high low yes no female high
# # Fred low low yes no male high
# # Pete low medium no yes male low
# # Jacob high medium yes yes male low
# # Sofia medium low no no female low
# data = pd.DataFrame()
# data['Name'] = ['Tim', 'Joe', '']
# one_hot_data = pd.get_dummies(data[['A','B','C']],drop_first=True)
# tree.fit(one_hot_data, data['Class'])
# clf = clf.fit(X, Y)
# # 训练完成后,我们可以用 export_graphviz 将树导出为 Graphviz 格式,存到文件iris.dot中
# with open("credit.dot", 'w') as f:
# f = tree.export_graphviz(clf, out_file=f)
# [['Name' 'Debt' 'Income' 'Married?' 'Owns_Property' 'Gender' 'Risk']
# ['Tim' 'low' 'low' 'no' 'no' 'male' 'low']
# ['Joe' 'high' 'high' 'yes' 'yes' 'male' 'low']
# ['Sue' 'low' 'high' 'yes' 'no' 'female' 'low']
# ['John' 'medium' 'low' 'no' 'no' 'male' 'high']
# ['Mary' 'high' 'low' 'yes' 'no' 'female' 'high']
# ['Fred' 'low' 'low' 'yes' 'no' 'male' 'high']
# ['Pete' 'low' 'medium' 'no' 'yes' 'male' 'low']
# ['Jacob' 'high' 'medium' 'yes' 'yes' 'male' 'low']
# ['Sofia' 'medium' 'low' 'no' 'no' 'female' 'low']]
# -
# !pip install pandas
| hw1/test/.ipynb_checkpoints/Untitled-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# !pip install python-dotenv
from dotenv import load_dotenv, find_dotenv
#find .env automatically by walking up directories until it's found
dotenv_path = find_dotenv()
#load up the entries as environment variables
load_dotenv(dotenv_path)
#extracting environment variables using os.environment.get
import os
KAGGLE_USERNAME = os.environ.get("KAGGLE_USERNAME")
print(KAGGLE_USERNAME)
#imports
import requests
from requests import session
import os
from dotenv import load_dotenv, find_dotenv
os.environ.get("KAGGLE_USERNAME")
# !pip install kaggle
# !kaggle competitions download -c titanic
#file paths
raw_data_path = os.path.join(os.path.pardir,'data','raw')
train_data_path = os.path.join(raw_data_path,'train.csv')
test_data_path = os.path.join(raw_data_path,'test.csv')
train_data_path
| notebooks/titanic_project-data_extract.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# PIKE: I will analyze the data in blair-l's 'meetings' folder
# +
# PIKE: First, I will read in the files from this folder and store their pathnames in a new list
import os
root_path = '/home/EDSE-brandmr-workspace/Enron_dataset/maildir/blair-l/meetings'
count_emails = 0
full_filepath = []
for dirs, subdirs, files in os.walk(root_path):
count_emails += len(files)
for f in files:
full_filepath.append(os.path.join(dirs,f))
print("Number of emails: ",len(full_filepath))
# +
# PIKE: Next, I will have the computer read through each of the emails from each filepath and pull out relevant data like
# To, From, Subject, Date, and Body of the email and store them in a dictionary
from email.parser import Parser
to_read = ""
emails_to = []
emails_from = []
emails_subject = []
emails_body = []
emails_path = []
emails_date = []
emails = {}
i = -1
for path in full_filepath:
i += 1
to_read = path
with open(to_read, "r") as f:
data = f.read()
email = Parser().parsestr(data)
to1 = email['to']
emails_to.append(to1)
if to1 is None:
to1 = "To: N/A"
else:
to1 = "To: "+to1
from1 = email['from']
emails_from.append(from1)
from1 = "From: "+from1
subject1 = email['subject']
emails_subject.append(subject1)
if subject1 !="":
subject1 = "Subject: "+subject1
else:
subject1 = "Subject: N/A"
date1 = email['date']
emails_date.append(date1)
date1 = "Date: "+date1
body1 = email.get_payload()
emails_body.append(body1)
body1 = "Body: "+body1
emails_path.append(path)
emails[i] = from1,to1,subject1,date1,
#(I know I never used the dictionary in this example, but I spent a lot of time to create it so I'm keeping it!)
# +
#PIKE: Then I will create a pandas dataframe from the collected data
# (this code pulled from geeksforgeeks.org/create-a-pandas-dataframe-from-lists/)
import pandas as pd
Emails_DataFrame = pd.DataFrame(list(zip(emails_path,emails_date,emails_from,emails_to,emails_subject,emails_body)),
columns = ["Path","Date","From","To","Subject","Body"])
Emails_DataFrame.head()
# +
#PIKE: Let's see when these emails were sent...
unique_date = set(emails_date)
unique_date
# +
#PIKE: And then let's graph the distribution of these emails by date...
# (this code is from https://stackoverflow.com/questions/28418988/how-to-make-histogram-from-a-list-of-strings-in-python)
from collections import Counter
email_date_counter = Counter(emails_date)
email_date_counter_DF = pd.DataFrame.from_dict(email_date_counter, orient = 'index')
email_date_counter_DF.plot(kind='bar')
# +
#PIKE: And we see that the majority of the emails (by a considerable margin) are from the same date: 27 JUN 2001
| Homework 8_Mohr, Brandon.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Generating a training set
# - The goal of this notebook is to generate a training set for a ML algorithm.
# - The first approach will be simple.
# - Using the DQ as the label, train up a binary classifier to find cosmic rays
# %matplotlib notebook
from astropy.io import fits
import pandas as pd
import numpy as np
import os
import glob
import datetime as dt
import matplotlib.pyplot as plt
import matplotlib.patches as patches
import numpy as np
from astropy.visualization import SqrtStretch
from astropy.visualization import LogStretch, LinearStretch, ZScaleInterval
from astropy.visualization.mpl_normalize import ImageNormalize
from sklearn.preprocessing import StandardScaler
from scipy import ndimage
import sys
import skimage.segmentation as segment
from photutils.utils import random_cmap
plt.style.use('ggplot')
sys.path.append('/Users/nmiles/hst_cosmic_rays/lib/')
from CosmicRayLabel import CosmicRayLabel
from ComputeStats import ComputeStats
fname = './../data/jd4wemc8q_flt.fits'
c = CosmicRayLabel(fname)
c.generate_label()
with fits.open(fname) as hdu:
sci2 = hdu[1].data
sci1 = hdu[4].data
sci = np.concatenate([sci2, sci1])
sci
stats = ComputeStats(fname,c.label)
sizes = stats.compute_size()
sizes
max_size_idx = np.where(np.asarray(list(sizes.values())) > 15.)
max_size_idx = max_size_idx[0] + 1
idx = max_size_idx[1]
sizes[idx]
stats.cr_locs[idx-1]
box_data = sci[stats.cr_locs[idx-1]]
box_data.shape
coords = np.where(box_data > 4*np.median(box_data))
coords = list(zip(coords[0], coords[1]))
box_data_dq = c.dq[stats.cr_locs[idx-1]]
def mk_patch(r, c='red'):
CR_center = patches.Rectangle((r[1]-0.5,r[0]-0.5),
width=1, height=1,
alpha=1.0, fill=False,
linewidth=1.75, color=c)
return CR_center
norm = ImageNormalize(sci, stretch=LogStretch(a=5.), interval=ZScaleInterval())
fig = plt.figure(figsize=(5,3))
ax1 = fig.add_subplot(1,2,1)
ax2 = fig.add_subplot(1,2,2,sharex=ax1, sharey=ax1)
ax1.imshow(box_data, cmap='gray', origin='lower', norm=norm)
ax2.imshow(box_data_dq, cmap='bone', interpolation='nearest', origin='lower')
for coord in coords:
patch1 = mk_patch(coord)
patch2 = mk_patch(coord)
ax1.add_patch(patch1)
ax2.add_patch(patch2)
np.where(box_data < -10)
num_pix = len(box_data.flatten())
data = {'sci':box_data.flatten(),'dq':box_data_dq.flatten(),'pix':np.linspace(1, num_pix, num_pix)}
data
print(len(data['sci']), len(data['dq']), len(data['pix']))
df = pd.DataFrame(data)
df.plot(kind='scatter',x='pix',y='sci',c='dq',colormap=plt.get_cmap('inferno_r'), alpha=0.65)
d = [[1,2,3,4,5],[6,7,8,9,10]]
d = np.asarray(d)
d.flatten()
# ### Generate a multipage pdf of cutouts for cosmic rays to show what cosmic rays of different sizes look like
avg_size = np.nanmean(list(sizes.values()))
std_size = np.nanstd(list(sizes.values()))
avg_size, std_size
max_size_idx = np.where(np.asarray(list(sizes.values())) > 5.)
| pipeline/prototyping_parameters.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Image Classifier - MIDAS INTERNSHIP CHALLENGE
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
# ## Explanation - 1
# The following code is used to import the our 3rd Party Deep Learning Libraries like *Pytorch* as well as checks, if the GPU is set or not.
#
# <img src ="https://cdn-images-1.medium.com/max/2600/1*aqNgmfyBIStLrf9k7d9cng.jpeg"/>
from fastai.vision import *
import csv
from fastai.metrics import error_rate
from lr_finder import LRFinder
import pandas as pd
import time
from torch.utils.data.sampler import SubsetRandomSampler
import os
from torch.autograd import Variable
import matplotlib.image as mpimg
from PIL import Image
import torch
import copy
from torch.utils.data.dataset import Dataset
from torch.utils.data import DataLoader
import torch
import torchvision
import torchvision.transforms as transforms
import torch.nn.functional as F
from torch import nn, optim
import torch.optim as optim
import numpy as np
import torch
import pickle
import os
import torchvision
use_cuda = torch.cuda.is_available()
device = torch.device("cuda:0" if use_cuda else "cpu")
print("Device is usage: {0}".format(device))
# ## Explanation 2
#
# ### Intuition
# The code block below defines few global variables that we would like to take care of, for example the batch size for our Deep Learning Pipeline as well as the paths of the training data/labels as well as the test_data
# +
data_path = "./data"
train_images_path = "{0}/train_image.pkl".format(data_path)
test_images_path = "{0}/test_image.pkl".format(data_path)
train_labels_path = "{0}/train_label.pkl".format(data_path)
sanitized_labels_path = "{0}/train_labels_sanitized.pkl".format(data_path)
batch_size = 64
random_seed = 43
validation_split = 0.2
shuffle_dataset = True
torch.manual_seed(random_seed)
with open(test_images_path, "rb") as file:
test_data = torch.ByteTensor(pickle.load(file)).view(-1, 28, 28)
# +
def path_exists(path):
"""
Function to verfiy if, file path defined is correct
"""
return os.path.exists(path)
valid_path = [path_exists(_) for _ in (train_images_path, test_images_path, train_labels_path)]
valid_path
# +
def sanitize_labels(labels_path):
"""
The labels given in the file are 0, 2, 3, 6
Each label should be mapped to required class, as 0 -> 0, 2 -> 2, 3 -> 2, 6 -> 3
"""
mapping = {0: 0, 2: 1, 3: 2, 6: 3}
with open(labels_path, 'rb') as file:
data = pickle.load(file)
result = [mapping[i] for i in data]
return result
result = sanitize_labels(train_labels_path)
with open(sanitized_labels_path, "wb") as file:
pickle.dump(result, file)
path_exists(sanitized_labels_path)
# -
# ## Deep Learning Pipeline - Data Pre-processing
#
# <img src="https://cdn-images-1.medium.com/max/1200/1*ZX05x1xYgaVoa4Vn2kKS9g.png" />
#
# ### What's happening ?
# In the code block we have defined a basic class which takes care of our pre-processing pipeline.
#
# #### Why did we **inherit** the `Dataset` class ?
# We are using PyTorch as our DeepLearning framework, PyTorch provides a very simple API to build the Deep Learnign PipeLine, we can do make our own pipeline by inheriting the **Dataset** class, which has internal methods to take of the things we need to take care of pre-processing.
#
# Here in our case I have divided the pipelin in the following parts.
# - Load data from pickle dump.
# - Build an API to *clean the data*, or arrange the data in a format that PyTorch understands, this has been done by the `__getitem__()` function.
# - This functions iteratively returns a single dataset and the following label for our Neural Net to work on.
# - We also incorporate transformations in the class, which helps us normalising our data.
# - **Normalising** input data is important, as this helps *Gradient Descent* to run faster, which means we can use a higher learning rate for our Neural Network, this helps us the reacht the minima faster, the contour of the cost functions are symmetrical when data is normalised.
#
class DataSetLoader(Dataset):
'''Dataset Loader'''
def __init__(self, train_path, labels_path, transform=None, train=True, target_transform=None):
"""
Args:
train_path (string): Path to the training data file
labels_path (string): Path to the labels present for the training data
transform (callable): Optional transform to apply to sample
"""
self.test_lables = []
self.train=train
if self.train:
data = self._load_from_pickle(train_path)
self.train_data = torch.ByteTensor(data).view(-1, 28, 28)
self.train_labels = self._load_from_pickle(labels_path)
else:
data = self._load_from_pickle(train_path)
self.test_data = torch.ByteTensor(data).view(-1, 28, 28)
self.test_lables = [i for i in range(len(data))]
del data
self.transform = transform
self.target_transform = target_transform
def __len__(self):
"""
Returns the length of whole Dataset fed into the Neural Net.
"""
if self.train:
return len(self.train_data)
else:
return len(self.test_data)
def __getitem__(self, index):
"""
Returns a single training/test example after applying the required normalisation/transformations techniques.
As well as the label.
ret: (image, label)
"""
if self.train:
img, target = self.train_data[index], self.train_labels[index]
else:
img, target = self.test_data[index], self.test_lables[index]
# return a PIL Image
img = Image.fromarray(img.numpy(), mode='L')
if self.transform is not None:
img = self.transform(img)
if self.target_transform is not None:
target = self.target_transform(target)
return img, target
def _load_from_pickle(self, file_path):
"""
file_path: File path to load data, returns an ndarray
ret: Loaded dump, in primitive data format.
"""
with open(file_path, 'rb') as file:
data = pickle.load(file)
return data
def get_labels(self):
"""
Return an array of labels present in the data set.
"""
if self.train:
return np.unique(self.train_labels)
return []
def visualise_data_set(self):
"""
Utility function to randomly display images from the dataset
"""
fig = plt.figure(figsize=(8,8))
columns = 4
rows = 5
if self.train:
data_set = self.train_data
else:
data_set = self.test_data
for i in range(1, columns * rows +1):
img_xy = np.random.randint(len(data_set));
img = data_set[img_xy][0][0,:,:]
fig.add_subplot(rows, columns, i)
plt.title(labels_map[data_set[img_xy][1]])
plt.axis('off')
plt.imshow(img)
plt.show()
# +
# Creating a labels map for storing the labels
# Define the normalisation/tranformation techniques we need during pre-processing.
img_transforms = transforms.Compose([transforms.ToTensor(),
transforms.Normalize((0.5,0.5,0.5), (0.5,0.5,0.5))])
# +
# Create training and test data sets.
train_dataset = DataSetLoader(train_images_path, sanitized_labels_path, train=True, transform=img_transforms)
test_dataset = DataSetLoader(test_images_path, "", train=False, transform=img_transforms)
dataset_size = len(train_dataset)
indices = list(range(dataset_size))
split = int(np.floor(validation_split * dataset_size))
if shuffle_dataset :
np.random.seed(random_seed)
np.random.shuffle(indices)
train_indices, val_indices = indices[split:], indices[:split]
train_sampler = SubsetRandomSampler(train_indices)
valid_sampler = SubsetRandomSampler(val_indices)
# -
# ## Looking at the data
#
# The following code blocks gives us some insights about what kind of images our data set has.
# By looking at the training set, we have 4 labels, *0, 2 ,3, 6*, these labels are numerically labelled in our training labelled data set.
#
# Just for readibilty I have created a dictionary of mapping each numerical label to a physical label, like *Shirt*, *T-Shirt* etc.
#
# ```
# labels = train_dataset.get_labels()
# ```
# The code above gives us the unique labels present in our training data.
# ```
# labels_map = {0 : 'T-Shirt', 1 : 'Pullover', 2 : 'Dress', 3 : 'Shirt'}
# ```
# Initialises a dictionary to map each numerical label to a physical entity.
labels = train_dataset.get_labels()
labels_map = {0 : 'T-Shirt', 1 : 'Pullover', 2 : 'Dress', 3 : 'Shirt'}
reverse_mapping = {0: 0, 1: 2, 2: 3, 3: 6}
for label in labels:
print("Label: {0}, value: {1}".format(label, labels_map[label]))
class_names = ['T-Shirt','Pullover', 'Dress', 'Shirt']
fig = plt.figure(figsize=(8,8));
columns = 4;
rows = 5;
for i in range(1, columns*rows +1):
img_xy = np.random.randint(len(train_dataset));
img = train_dataset[img_xy][0][0,:,:]
fig.add_subplot(rows, columns, i)
plt.title(labels_map[train_dataset[img_xy][1]])
plt.axis('off')
plt.imshow(img)
plt.show()
# ### Explanation of code block below
#
# For our data pre-processing we have cleaned our data, applied required transformations, as well as wrote a simple API that allows us to do the steps gracefully.
# What we need now is another API which can help us in iterating over our training/test datasets efficiently.
#
# The following code does the same, **DataLoader** class provides us with an *Python* `iterator` object which allows for traverse over our dataset very efficiently, in specific batch size we want.
#
# ```
# for i_batch, sample_batched in enumerate(trainloader):
# do_something(sample_batched)
# ```
#
# Since trainloader is an iterator we can easily now iterate over our data set, you can also the see the code example above.
#
# #### What's my intuition behind this ?
#
# The most basic intuition in making an iterator here is we need to carry the basic Neural Net operations:
# - Forward Propagation
# - Backward Propagation
#
# Since we want to carry out **Mini Batch Gradient Descent** for our Neural Network to learn the hyper-params, we are using this iterator to automatically create the required batches.
# +
"""
We built our data set using DataSetLoader class we defined above, now we need the the following operations
1. Divide our dataset the into the batch sizes.
2. Shuffle the dataset accordingly for randomness.
"""
trainloader = torch.utils.data.DataLoader(dataset=train_dataset, batch_size=batch_size,
sampler=train_sampler, num_workers=2)
validationloader = torch.utils.data.DataLoader(dataset=train_dataset,batch_size=batch_size,
sampler=valid_sampler, num_workers=2)
testloader = torch.utils.data.DataLoader(dataset=test_dataset,
batch_size=batch_size,
shuffle=True, num_workers=2)
# -
# ## Model Creation
#
# ### What is code block doing ?
# The code block below defines the architecture of our Convolutional Neural Network. Our input is an image of dimensions (28 x 28), usually images have 3 channels, **RED, BLUE AND GREEN**, but in the current data set we have only 1 channel, due to this we initialise our Convolutional Layer accordingly.
#
# In the first Convolutional block of our CNN we use only a single channel that is 1, becaue input image has only 1 channel.
# ```
# class CNN(nn.Module):
# def __init__(self):
# .....
# nn.Conv2d(1, 16, kernel_size=4, padding=2)
# .....
# ```
#
# Similarly.
#
# ```
# class CNN(nn.Module):
#
# def __init__(self):
# .....
# nn.Conv2d(16, 32, kernel_size=6, padding=2),
# ....
# ```
# In the second Convolutional Block, the number of input channels are 16 as the number of channels that will be received from CNN block 1 are 16.
#
# The code block above specifies that our Convolutional Blocks have a padding of 2 and kernel size of 5.
# We use BatchNormalisation to smoothen our loss landscape, and we use Dropout as a regularisation technique to keep overfitting in check.
#
# ### My Intuition
#
# The basic intuition behind the CNN architecture is to involve techniques like Batch Normalization and Regularisation techniques like Dropout, on one hand, as well know Gradient Descent will converge to a minima for the training set,
# but if the number of epochs are high, then this can lead to overfitting.
#
# One another important factor to keep in-check is how we initialise weights, if weights are too small, gradients can vanish easily, if weights are too large, gradients can explode, to take care of the this issue, we initialise the weights with according to ideas presented in the paper ["Understanding the difficulty of training deep feedforward neural networks"](http://proceedings.mlr.press/v9/glorot10a/glorot10a.pdf), if we initialise the weights of our Neural Netowk using the techniques presented in the paper, we can help reduce the chances of vanishing/exploding gradients.
#
# ```
# def weights_initialisation(m):
# if isinstance(m, nn.Conv2d):
# torch.nn.init.xavier_uniform_(m.weight)
# if m.bias is not None:
# torch.nn.init.zeros_(m.bias)
# ```
#
# The code snippet above initialises the weights of our convolutional filters with a variance of `(2/n)`, where n is the size of input of the layer, as per the ideas ideas presented in the paper ["Understanding the difficulty of training deep feedforward neural networks"](http://proceedings.mlr.press/v9/glorot10a/glorot10a.pdf).
#
#
# therefore:
# - Dropout helps in regularising our weights to reduce overfitting, I have kept the probability of regularisation as 0.5
# - Batch Normalization smoothens the loss landscape and thus allows to converge faster to minima, using Gradient Descent.
# - Xavier Initialisation, can help in reducing vanishing/exploding gradients.
# +
def weights_initialisation(m):
if isinstance(m, nn.Conv2d):
torch.nn.init.xavier_uniform_(m.weight)
if m.bias is not None:
torch.nn.init.zeros_(m.bias)
class CNN(nn.Module):
"""
CNN architecture:
CONV -> BatchNorm -> Relu -> MaxPool -> CONV -> BatchNorm -> Relu -> Dropout -> MaxPool -> Linear -> Output
"""
def __init__(self):
super(CNN, self).__init__()
self.layer1 = nn.Sequential(
nn.Conv2d(1, 16, kernel_size=5, padding=2),
# 16 is the number of channels
nn.BatchNorm2d(16),
nn.ReLU(),
nn.MaxPool2d(2))
self.layer2 = nn.Sequential(
nn.Conv2d(16, 32, kernel_size=5, padding=2),
nn.BatchNorm2d(32),
nn.ReLU(),
nn.Dropout2d(p=0.5),
nn.MaxPool2d(2))
self.layer3 = nn.Sequential(
nn.Conv2d(32, 32, kernel_size=5, padding=2),
nn.BatchNorm2d(32),
nn.ReLU(),
nn.Dropout2d(p=0.5),
nn.MaxPool2d(2))
# Number of classes is 4
self.fc = nn.Linear(3 * 3 * 32, 4)
def forward(self, x):
out = self.layer1(x)
out = self.layer2(out)
out = self.layer3(out)
out = out.view(out.size(0), -1)
out = self.fc(out)
return out
# -
# ## Finding Hyper-parmeters
#
# Finding hyper-paramters are very important for buidling a good classifier, one of the most important one is *learning rate*
# There can two approaches we can take here to find the appropriate learning rate.
# - We can do a grid search of learning rate vs epochs to find the combination for the model with the best accuracy on our validation set.
# - This is might take great lenghts of time to find the right learning rate.
# - Or we can do use the ideas from the paper: [Cyclical Learning Rates for Training Neural Networks](https://arxiv.org/pdf/1506.01186.pdf), to find the a good learning rate range for our model.
#
# The code below depicts what can be possible learning rates for our CNN, given our training and validation set.
# Find learning rate using Exponential decay
net = CNN().cuda().apply(weights_initialisation)
optimizer = optim.Adam(net.parameters(), lr=1e-5)
lr_finder = LRFinder(net, optimizer, nn.CrossEntropyLoss().cuda(), device="cuda")
lr_finder.range_test(trainloader, val_loader=validationloader, end_lr=10)
lr_finder.plot()
history = lr_finder.history
lr_finder.reset()
# From the graphs above, we can consider the range of rates where the graph steeps downwards continously, that range can be taken as an approximate learning rate for our network, why? because the loss is calculated on our validation set `validationloader`, which is being passed in our code above, as the loss is continously decreasing, it makes sense to use that range as our range of learning rate.
#
# Therefore, we can use an approximate learning rate of from 0.001 to 0.01, here.
# ## Model Training
#
# The code below is an abstraction over how we will be training our model on our training data.
#
# ```
# class ModelRunner:
#
# def __init__(self, learning_rate, epochs):
# .....
#
# def run_model(self):
# .....
#
# def plot_losses(self):
# .....
# ```
# The code snippets above gives us the simple API to train our selected model on specific hyper-paramters we want.
# We have already seen the range of learning rate we need to keep so that our model can converge as well as reduce over-fitting.
#
# We now need to find the right range of epochs for which we need to train our model.
# We will use a technique called *Grid Search* to find the right set of paramters.
#
# Once our models are trained on those hyper-paramters, we will use our validation-set, `vaidationloader` to find the model which has the better accuracy.
#
# I have wrriten to this code snippet, so that I can easily do a grid search, save the data from the grid-search and then use our *validation set* to find which set of parameters are the best.
class ModelRunner:
"""
Class to abstract how we train our model on training and validation data sets.
"""
def __init__(self, learning_rate, epochs):
self.learning_rate = learning_rate
self.epochs = epochs
self.training_losses = []
def initialise_model(self):
self.model = CNN().cuda()
self.criterion = nn.CrossEntropyLoss().cuda()
self.optimizer = torch.optim.Adam(self.model.parameters(), lr=self.learning_rate)
def run_model(self):
self.initialise_model()
print("-------Running Model, with rate: {0} and epochs: {1} -------".format(self.learning_rate, self.epochs))
for epoch in range(self.epochs):
for i, (images, labels) in enumerate(trainloader):
images = Variable(images.to(device).float())
labels = Variable(labels.cuda())
self.optimizer.zero_grad()
outputs = self.model(images)
loss = self.criterion(outputs, labels)
loss.backward()
self.optimizer.step()
self.training_losses.append(loss.data.item())
if (i+1) % 100 == 0:
print('Epoch : %d/%d, Iter : %d/%d, Training Loss: %.4f'
%(epoch+1, self.epochs, i+1, len(train_dataset)//batch_size, loss.data.item()))
self.plot_losses()
print("\n \n")
return self.model
def plot_losses(self):
plt.xlabel('Epoch #')
plt.ylabel('Loss')
plt.plot(self.training_losses[::100])
plt.show()
"""
Doing a grid search over possible learning rates as well as epochs.
"""
models = []
learning_rates = [1e-3, 3e-3, 5e-3]
epochs = [40, 50, 60]
for rate in learning_rates:
for epoch in epochs:
runner = ModelRunner(rate, epoch)
models.append({"model": runner.run_model(), "rate":rate, "epoch": epoch})
# ## Model Evaluation
# Since now we have found our set of models over our grid-search, we need to evaluate which model is better for use.
# Now since we have trained on our training set, we cannot validate our models on that set.
#
# What we need is our validation-set, `validationloader`, we will test the accuracy of each model on our validation-set and find which one of them, is the most accurate.
#
# I have written 2 functions for this.
#
# ```
# def evaluate_model(model):
# ....
# ```
# and
#
# ```
# def find_best_model(models):
# ....
# ```
# `evaluate_model(model)`, shows the accuracy of a single model on our validation set.
#
# `find_best_model(models)`, takes a collection of models as input and returns the best model, with the correct combination of epochs and learning rate.
# +
np.random.seed(random_seed)
def evaluate_model(model_obj):
model_obj["model"].eval()
correct = 0
total = 0
print("\n --- Epoch and learning rate are: {0} and {1} ----".format(model_obj["epoch"], model_obj["rate"]))
for images, labels in validationloader:
images, labels = images.to(device), labels.to(device)
images = Variable(images.float())
outputs = model_obj["model"](images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum()
print('Validation Accuracy of the model on validation images: %.4f %%' % (100 * correct / total))
return (100 * correct / total)
def find_best_model(models):
best_model = None
best_metric = 0.0
for model in models:
metric = evaluate_model(model)
if metric > best_metric:
best_model = model
best_metric = metric
return best_model
# +
best_model = find_best_model(models)
epoch, rate = best_model["epoch"], best_model["rate"]
print("\n\nBest combination of Number of epochs and learning rate found are: {0}, {1}".format(epoch, rate))
torch.save(best_model["model"].state_dict(), "model_weights.pt")
# +
"""
Helper functions to help us visualize our predictions
"""
def imshow(inp, title=None):
"""Imshow for Tensor."""
inp = inp.numpy().transpose((1, 2, 0))
mean = np.array([0.485, 0.456, 0.406])
std = np.array([0.229, 0.224, 0.225])
inp = std * inp + mean
inp = np.clip(inp, 0, 1)
plt.imshow(inp)
if title is not None:
plt.title(title)
plt.pause(0.001) # pause a bit so that plots are updated
def visualize_model(model, num_images=6):
was_training = model.training
model.eval()
images_so_far = 0
fig = plt.figure()
with torch.no_grad():
for i, (inputs, labels) in enumerate(validationloader):
inputs = inputs.to(device)
labels = labels.to(device)
outputs = model(inputs)
_, preds = torch.max(outputs, 1)
for j in range(inputs.size()[0]):
images_so_far += 1
ax = plt.subplot(num_images//2, 2, images_so_far)
ax.axis('off')
ax.set_title('predicted: {}'.format(class_names[preds[j]]))
imshow(inputs.cpu().data[j])
if images_so_far == num_images:
model.train(mode=was_training)
return
model.train(mode=was_training)
visualize_model(best_model["model"])
# -
# ## Inference
# The following code below is used to give more insights on how good our model is.
# +
class_correct = list(0. for i in range(4))
class_total = list(0. for i in range(4))
label_decoder = dict(zip(range(4), class_names))
print("|Item|Accuracy (%)|")
print("|-+-|")
with torch.no_grad():
for images, labels in validationloader:
images, labels = images.to(device), labels.to(device)
images = Variable(images.float())
outputs = best_model["model"](images)
_, predicted = torch.max(outputs, 1)
c = (predicted == labels).squeeze()
for i in range(len(labels)):
label = labels[i]
class_correct[label.item()] += c[i].item()
class_total[label.item()] += 1
for i in range(4):
print('|{}|{:.1f}'.format(
label_decoder[i], 100 * class_correct[i] / class_total[i]))
# -
# Our model has good performance over T-Shirt, Pullover and Dress, but has fails to classify Shirts to a greater accuracy, this can be a result of class imbalance in our training set, we can solve this by sampling our training set in such a way that each class as particular number of examples, so that our classifier can understand the differences more.
# ## Testing and saving lables of our test set
#
# The following code below loads our model, from the best state of model we saved previously and saves the predicted class in a csv file.
# +
model = CNN()
model.load_state_dict(torch.load("model_weights.pt"))
model.eval()
count = 0
predictions = open('AKSHAY_SHARMA.csv', 'w')
with predictions:
fnames = ['index', 'image_class']
writer = csv.DictWriter(predictions, fieldnames=fnames)
writer.writeheader()
for i, (images, labels) in enumerate(testloader):
images = Variable(images.float())
outputs = model(images)
_, predicted = torch.max(outputs, 1)
for index, value in enumerate(predicted):
writer.writerow({'index':count, 'image_class': reverse_mapping[value.item()]})
count += 1
# -
# # Conclusion
# We have successfully created an image classifier, that can classify give image data into 4 classes.
#
# To recap:
# - We pre-prprocessed the data, including normalisation of images and as well making training, validation and test set.
# - We defined a CNN architecture, used Xavier Initialisation, Batch Normalization and Dropout regularisation as part of our architecture.
# - We did our hyper-parameter search, to find the correct learning rate as well as the number of epochs.
# - Used grid-search, and then found the better model, by using our validation set.
# - We saw how our classifier is working on different classes, from the data used from our validation set.
# - In the end, we dumped the our predictions of our test set in a csv file.
#
# **By <NAME>**, <<EMAIL>>
| CV_PROBLEM/Image Classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PyCharm (pycon2015_tutorial322)
# language: python
# name: pycharm-cef1f773
# ---
# # Exercises for "Hands-on with Pydata: How to Build a Minimal Recommendation Engine"
# # Systems check: imports and files
import numpy as np
import pandas as pd
# # Pandas questions: Series and DataFrames
# ## 1. Adding a column in a DataFrame
# given the following DataFrame, add a new column to it
df = pd.DataFrame({'col1': [1,2,3,4]})
df['col2'] = [5,6,7,8]
df
# ## 2. Deleting a row in a DataFrame
# given the following DataFrame, delete row 'd' from it
df = pd.DataFrame({'col1': [1,2,3,4]}, index = ['a','b','c','d'])
df.drop('d', inplace=True)
df
# ## 3. Creating a DataFrame from a few Series
# given the following three Series, create a DataFrame such that it holds them as its columns
ser_1 = pd.Series(np.random.randn(6))
ser_2 = pd.Series(np.random.randn(6))
ser_3 = pd.Series(np.random.randn(6))
pd.DataFrame({'ser_1': ser_1, 'ser_2': ser_2, 'ser_3': ser_3})
# # Pandas questions: Indexing
# ## 1. Indexing into a specific column
# given the following DataFrame, try to index into the 'col_2' column
df = pd.DataFrame(data={'col_1': [0.12, 7, 45, 10], 'col_2': [0.9, 9, 34, 11]},
columns=['col_1', 'col_2', 'col_3'],
index=['obs1', 'obs2', 'obs3', 'obs4'])
df
# + pycharm={"name": "#%%\n"}
df.col_2
# -
# ## 2. Label-based indexing
# + pycharm={"name": "#%%\n"}
# using the same DataFrame, index into the row whose index is 'obs3'
df.loc['obs3']
# -
# ## 2. Position-based indexing
# using the same DataFrame, index into into its first row
df.iloc[:1]
# # Mini-Challenge prep: data loading
#
# ## 1. How to load the `users` and `movies` portions of MovieLens
# +
import pandas as pd
users = pd.read_table('data/ml-1m/users.dat',
sep='::', header=None,
names=['user_id', 'gender', 'age', 'occupation', 'zip'], engine='python')
movies = pd.read_table('data/ml-1m/movies.dat',
sep='::', header=None,
names=['movie_id', 'title', 'genres'], engine='python')
# -
# ## 2. How to load the training and testing subsets
# subset version (hosted notebook)
movielens_train = pd.read_csv('data/movielens_train.csv', index_col=0)
movielens_test = pd.read_csv('data/movielens_test.csv', index_col=0)
movielens_train.head()
movielens_test.head()
# # Mini-Challenge prep: evaluation functions
#
# These are the two functions that you will need to test your `estimate` method.
def compute_rmse(y_pred, y_true):
""" Compute Root Mean Squared Error. """
return np.sqrt(np.mean(np.power(y_pred - y_true, 2)))
def evaluate(estimate_f):
""" RMSE-based predictive performance evaluation with pandas. """
ids_to_estimate = zip(movielens_test.user_id, movielens_test.movie_id)
estimated = np.array([estimate_f(u,i) for (u,i) in ids_to_estimate])
real = movielens_test.rating.values
return compute_rmse(estimated, real)
# Test a dummy solution!
# This is a baseline that just gives an average rating to everything
def my_estimate_func(user_id, movie_id):
return 3.0
# You can test for performance with the following line, which assumes that your function is called `my_estimate_func`:
print('RMSE for my estimate function: %s' % evaluate(my_estimate_func))
# + [markdown] pycharm={"name": "#%% md\n"}
# The above is the starting baseline.
# Want to improve from here.
# 0 error means the recommendation is perfect.
# 4 (5-1) is the maximum amount it could be off.
# -
# # Reco systems questions: Minimal reco engine v1.0
# ## 1. Simple collaborative filtering using mean ratings
# +
# write an 'estimate' function that computes the mean rating of a particular user
def collab_mean(user_id, movie_id):
""" Simple collaborative filter based on mean ratings. """
user_condition = movielens_train.user_id != user_id
movie_condition = movielens_train.movie_id == movie_id
ratings_by_others = movielens_train.loc[user_condition & movie_condition]
if ratings_by_others.empty:
return 3.0
else:
return ratings_by_others.rating.mean()
# try it out for a user_id, movie_id pair
collab_mean(4653, 2648)
# + pycharm={"name": "#%%\n"}
print(f'RMSE for estimate2 is: {evaluate(collab_mean)}.')
# -
# # Mini-Challenge: first round
# Implement an `estimate` function of your own using other similarity notions, eg.:
#
# - collaborative filter based on age similarities
# - collaborative filter based on zip code similarities
# - collaborative filter based on occupation similarities
# - content filter based on movie genre
# + pycharm={"name": "#%%\n"}
user_info = users.set_index('user_id')
user_info.head(3)
# + pycharm={"name": "#%%\n"}
class CollaborativeRecommendation:
""" Collaborative filtering using an implicit sim(u,u'). """
def __init__(self):
""" Prepare data structures for estimation. """
self.means_by_gender = movielens_train.pivot_table('rating', index='movie_id', columns='gender')
self.means_by_age = movielens_train.pivot_table('rating', index='movie_id', columns='age')
self.means_by_zip = movielens_train.pivot_table('rating', index='movie_id', columns='zip')
self.means_by_occupation = movielens_train.pivot_table('rating', index='movie_id', columns='occupation')
def estimate_gender(self, user_id, movie_id):
""" Mean ratings by other users of the same gender. """
if movie_id not in self.means_by_gender.index:
return 3.0
user_gender = user_info.loc[user_id, 'gender']
if ~np.isnan(self.means_by_gender.loc[movie_id, user_gender]):
return self.means_by_gender.loc[movie_id, user_gender]
else:
return self.means_by_gender.loc[movie_id].mean()
def estimate_age(self, user_id, movie_id):
""" Mean ratings by other users of the same age. """
if movie_id not in self.means_by_age.index:
return 3.0
user_age = user_info.loc[user_id, 'age']
if ~np.isnan(self.means_by_age.loc[movie_id, user_age]):
return self.means_by_age.loc[movie_id, user_age]
else:
return self.means_by_age.loc[movie_id].mean()
def estimate_zip(self, user_id, movie_id):
""" Mean ratings by other users of the same zip code. """
if movie_id not in self.means_by_zip.index:
return 3.0
user_zip = user_info.loc[user_id, 'zip']
if ~np.isnan(self.means_by_zip.loc[movie_id, user_zip]):
return self.means_by_zip.loc[movie_id, user_zip]
else:
return self.means_by_zip.loc[movie_id].mean()
def estimate_occupation(self, user_id, movie_id):
""" Mean ratings by other users of the same occupation. """
if movie_id not in self.means_by_occupation.index:
return 3.0
user_occupation = user_info.loc[user_id, 'occupation']
if ~np.isnan(self.means_by_occupation.loc[movie_id, user_occupation]):
return self.means_by_occupation.loc[movie_id, user_occupation]
else:
return self.means_by_occupation.loc[movie_id].mean()
# + pycharm={"name": "#%%\n"}
reco = CollaborativeRecommendation()
print('RMSE for Gender: %s' % evaluate(reco.estimate_gender))
print('RMSE for Age: %s' % evaluate(reco.estimate_age))
print('RMSE for Zip: %s' % evaluate(reco.estimate_zip))
print('RMSE for Occupation: %s' % evaluate(reco.estimate_occupation))
# + pycharm={"name": "#%%\n"}
class CollaborativeRecommendation2:
""" Collaborative filtering using an implicit sim(u,u'). """
def __init__(self, feature):
""" Prepare data structures for estimation. """
self._feature = feature
self.means_by_feature = movielens_train.pivot_table('rating', index='movie_id', columns=self.feature)
@property
def feature(self):
return self._feature
def estimate(self, user_id, movie_id):
""" Mean ratings by other users of the same feature. """
if movie_id not in self.means_by_feature.index:
return 3.0
user_feature = user_info.loc[user_id, self.feature]
if ~np.isnan(self.means_by_feature.loc[movie_id, user_feature]):
return self.means_by_feature.loc[movie_id, user_feature]
else:
return self.means_by_feature.loc[movie_id].mean()
# + pycharm={"name": "#%%\n"}
gender = CollaborativeRecommendation2('gender')
print('RMSE for Gender: %s' % evaluate(gender.estimate))
age = CollaborativeRecommendation2('age')
print('RMSE for Age: %s' % evaluate(age.estimate))
zip_code = CollaborativeRecommendation2('zip')
print('RMSE for Zip: %s' % evaluate(zip_code.estimate))
occupation = CollaborativeRecommendation2('occupation')
print('RMSE for Occupation: %s' % evaluate(occupation.estimate))
# -
# # Mini-Challenge: second round
# Implement an `estimate` function of your own using other custom similarity notions, eg.:
#
# - euclidean
# - cosine
# + pycharm={"name": "#%%\n"}
from scipy.special import logsumexp
# Euclidean 'similarity'
def euclidean(s1, s2):
"""Take two pd.Series objects and return their euclidean 'similarity'."""
diff = s1 - s2
return 1 / (1 + np.sqrt(np.sum(diff ** 2)))
# Cosine similarity
def cosine(s1, s2):
"""Take two pd.Series objects and return their cosine similarity."""
return np.sum(s1 * s2) / np.sqrt(np.sum(s1 ** 2) * np.sum(s2 ** 2))
# Pearson correlation
def pearson(s1, s2):
"""Take two pd.Series objects and return a pearson correlation."""
s1_c = s1 - s1.mean()
s2_c = s2 - s2.mean()
return np.sum(s1_c * s2_c) / np.sqrt(np.sum(logsumexp(s1_c ** 2)) * np.sum(logsumexp(s2_c ** 2)))
# Jaccard similarity
def jaccard(s1, s2):
dotp = np.sum(s1 * s2)
return dotp / (np.sum(s1 ** 2) + np.sum(s2 ** 2) - dotp)
def binjaccard(s1, s2):
dotp = s1.index.intersection(s2.index).size
return dotp / (s1.sum() + s2.sum() - dotp)
# + pycharm={"name": "#%%\n"}
class CollaborativeSimilarityRecommendation:
""" Collaborative filtering using a custom sim(u,u'). """
def __init__(self, similarity=None):
""" Prepare datastructures for estimation. """
self.all_user_profiles = movielens_train.pivot_table('rating', index='movie_id', columns='user_id')
self._similarity = similarity
@property
def similarity(self):
return self._similarity
@similarity.setter
def similarity(self, value):
self._similarity = value
def estimate(self, user_id, movie_id):
""" Ratings weighted by correlation similarity. """
user_condition = movielens_train.user_id != user_id
movie_condition = movielens_train.movie_id == movie_id
ratings_by_others = movielens_train.loc[user_condition & movie_condition]
if ratings_by_others.empty:
return 3.0
ratings_by_others.set_index('user_id', inplace=True)
their_ids = ratings_by_others.index
their_ratings = ratings_by_others.rating
their_profiles = self.all_user_profiles[their_ids]
user_profile = self.all_user_profiles[user_id]
sims = their_profiles.apply(lambda profile: self.similarity(profile, user_profile), axis=0)
ratings_sims = pd.DataFrame({'sim': sims, 'rating': their_ratings})
ratings_sims = ratings_sims[ratings_sims.sim > 0]
if ratings_sims.empty:
return their_ratings.mean()
else:
return np.average(ratings_sims.rating, weights=ratings_sims.sim)
# + pycharm={"name": "#%%\n"}
reco = CollaborativeSimilarityRecommendation(pearson)
print('RMSE for Pearson: %s' % evaluate(reco.estimate))
# + pycharm={"name": "#%%\n"}
reco = CollaborativeSimilarityRecommendation(euclidean)
print('RMSE for Euclidean: %s' % evaluate(reco.estimate))
# + pycharm={"name": "#%%\n"}
reco = CollaborativeSimilarityRecommendation(cosine)
print('RMSE for Cosine: %s' % evaluate(reco.estimate))
# + pycharm={"name": "#%%\n"}
reco = CollaborativeSimilarityRecommendation(jaccard)
print('RMSE for Jaccard: %s' % evaluate(reco.estimate))
# + pycharm={"name": "#%%\n"}
reco = CollaborativeSimilarityRecommendation(binjaccard)
print('RMSE for Bin Jaccard: %s' % evaluate(reco.estimate))
# + pycharm={"name": "#%%\n"}
from functools import reduce
class Recommender(object):
def __init__(self):
self.overall_mean = movielens_train['rating'].mean()
self.all_user_profiles = movielens_train.pivot_table('rating', index='movie_id', columns='user_id')
def estimate_movie(self, user_id, movie_id):
all_ratings = movielens_train.loc[movielens_train.movie_id == movie_id]
if all_ratings.empty:
return self.overall_mean
all_ratings.set_index('user_id', inplace=True)
their_ids = all_ratings.index
their_ratings = all_ratings.rating
their_profiles = self.all_user_profiles[their_ids]
user_profile = self.all_user_profiles[user_id]
sims = their_profiles.apply(lambda profile: pearson(profile, user_profile), axis=0)
ratings_sims = pd.DataFrame({'sim': sims, 'rating': their_ratings})
ratings_sims = ratings_sims[ratings_sims.sim > 0]
if ratings_sims.empty:
return their_ratings.mean()
else:
return np.average(ratings_sims.rating, weights=ratings_sims.sim)
def estimate_user(self, user_id):
similar = movielens_train.loc[reduce(np.logical_and,
[movielens_train.gender == users.loc[user_id].gender,
movielens_train.occupation == users.loc[user_id].occupation,
abs(movielens_train.age - users.loc[user_id].age) < 10 ])]
if similar.empty:
return self.overall_mean
else:
return similar['rating'].mean()
def estimate(self, user_id, movie_id):
return 0.5 * self.estimate_user(user_id) + 0.5 * self.estimate_movie(user_id, movie_id)
rec = Recommender()
print('RMSE for recommender estimate class: %s' % evaluate(rec.estimate))
| questions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Skater
import warnings
warnings.filterwarnings('ignore')
import sys
sys.path.append("../")
import spopt
import geopandas as gpd
import libpysal
import numpy as np
from sklearn.metrics import pairwise as skm
from spopt.region.skater import Skater
# ##### Airbnb Spots Clustering in Chicago
#
# Cluster 77 communities into 10 regions such that each region consists of at least 3 communities and homogeneity in the number of Airbnb spots in communities is maximized.
from libpysal.examples import load_example
load_example('AirBnB')
# +
pth = libpysal.examples.get_path('airbnb_Chicago 2015.shp')
chicago = gpd.read_file(pth)
chicago
# -
chicago.plot(column='community', categorical=True, figsize=(12,8), edgecolor='w')
# Initialize the parameters
w = libpysal.weights.Queen.from_dataframe(chicago)
attrs_name = ['num_spots']
n_clusters = 10
floor = 3
trace = False
islands = "increase"
spanning_forest_kwds = dict(dissimilarity=skm.manhattan_distances, affinity=None, reduction=np.sum, center=np.mean)
model = Skater(chicago, w, attrs_name, n_clusters, floor, trace, islands, spanning_forest_kwds)
model.solve()
chicago['skater_new'] = model.labels_
chicago.plot(column='skater_new', categorical=True, figsize=(12,8), edgecolor='w')
| notebooks/skater.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import numpy as np
from sklearn.model_selection import train_test_split
# %matplotlib inline
# +
def to_one_hot(Y):
n_col = np.amax(Y) + 1
binarized = np.zeros((len(Y),n_col))
for i in range(len(Y)):
binarized[i,Y[i]] = 1
return binarized
def from_one_hot(Y):
arr = np.zeros((len(Y),1))
for i in range(len(Y)):
arr[i] = np.nonzero(Y[i]) + 1
return arr
def sigmoid(x, deriv = False):
if(deriv==True):
return x*(1-x)
return 1/(1+np.exp(-x))
def normalize(X, axis=-1, order=2):
l2 = np.atleast_1d(np.linalg.norm(X, order, axis))
l2[l2 == 0] = 1
return X / np.expand_dims(l2, axis)
# -
iris = pd.read_csv("Datasets/Iris.csv")
sns.pairplot(iris.drop("Id", axis=1), hue="Species", height=2, diag_kind="kde")
# +
iris['Species'].replace(['Iris-setosa', 'Iris-virginica', 'Iris-versicolor'], [0, 1, 2], inplace=True)
#Get input
columns = ['SepalLengthCm', 'SepalWidthCm', 'PetalLengthCm', 'PetalWidthCm']
x = pd.DataFrame(iris, columns=columns)
x = normalize(x.values)
#Get Output, flatten and encode to one-hot
columns = ['Species']
y = pd.DataFrame(iris, columns=columns)
y = y.values
y = y.flatten()
y = to_one_hot(y)
# -
X_train, X_test, y_train, y_test = train_test_split(x, y, test_size=0.33)
# +
#Weights
w0 = 2*np.random.random((4, 5)) - 1 #for input - 4 inputs, 3 outputs
w1 = 2*np.random.random((5, 3)) - 1 #for layer 1 - 5 inputs, 3 outputs
#learning rate
n = 0.1
#Errors - for graph later
errors = []
#Train
for i in range(300000):
#Feed forward
layer0 = X_train
layer1 = sigmoid(np.dot(layer0, w0))
layer2 = sigmoid(np.dot(layer1, w1))
#Back propagation using gradient descent
layer2_error = y_train - layer2
layer2_delta = layer2_error * sigmoid(layer2, deriv=True)
layer1_error = layer2_delta.dot(w1.T)
layer1_delta = layer1_error * sigmoid(layer1, deriv=True)
print(layer2_delta.shape, layer1_delta.shape, layer1.shape)
w1 += layer1.T.dot(layer2_delta) * n
w0 += layer0.T.dot(layer1_delta) * n
error = np.mean(np.abs(layer2_error))
errors.append(error)
accuracy = (1 - error) * 100
#Plot the accuracy chart
plt.plot(errors)
plt.xlabel('Training')
plt.ylabel('Error')
plt.show()
print("Training Accuracy " + str(round(accuracy,2)) + "%")
# +
#Validate
layer0 = X_test
layer1 = sigmoid(np.dot(layer0, w0))
layer2 = sigmoid(np.dot(layer1, w1))
layer2_error = y_test - layer2
error = np.mean(np.abs(layer2_error))
accuracy = (1 - error) * 100
print("Validation Accuracy " + str(round(accuracy,2)) + "%")
# -
| .ipynb_checkpoints/Kaggle Numpy NN-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import pystorm
from pystorm.hal import HAL
from pystorm.hal.net_builder import NetBuilder
from pystorm.hal.run_control import RunControl
from pystorm.hal.calibrator import Calibrator
# +
# call the function
hal = HAL()
nb = NetBuilder(hal)
cal = Calibrator(hal)
bad_syns, dbg = cal.get_bad_syns()
# +
# this was basically my scratchpad to write the function, does basically the same thing
# compare to function output afterwards
plt.figure()
plt.title('pulse widths by position')
pw = dbg['pulse_widths']
plt.imshow(pw)
med = np.median(pw)
std = np.std(pw)
print(med, std)
plt.figure()
plt.title('pulse widths, lines are median +/- 1 std')
plt.hist(pw.flatten(), bins=20)
ax = plt.gca()
plt.axvline(med, color='r')
plt.axvline(med - std, color='r')
plt.axvline(med + std, color='r')
plt.figure()
plt.title('pulse widths, lines are attrition cutoffs')
plt.hist(pw.flatten(), bins=20)
ax = plt.gca()
pwflat = pw.flatten()
fmaxes = 1 / pwflat
order = np.argsort(fmaxes)
fmax_for_attritions = []
attritions = np.linspace(0, .5, 11)
for attrition in attritions:
cutoff_idx = order[int(attrition * len(fmaxes))]
val = fmaxes[cutoff_idx]
plt.axvline(1 / val, color='r')
print('fmax at attrition of ' + "{:0.2f}".format(attrition) + ' is ' + str(val))
fmax_for_attritions.append(val)
plt.figure()
plt.title('fmax vs attrition')
plt.plot(attritions, fmax_for_attritions)
plt.xlabel('attrition')
plt.ylabel('fmax')
# +
# study correlation of high bias and slow pulse synapses
hb = dbg['high_bias']
plt.figure()
plt.title('high bias syns')
plt.imshow(hb)
bad_pw = fmaxes < fmax_for_attritions[1]
plt.figure()
plt.title('slow pulse syns')
plt.imshow(bad_pw.reshape(32, 32))
plt.figure()
plt.figure('both')
plt.imshow(bad_pw.reshape(32, 32) & hb)
redo_bad_syn = bad_pw.reshape(32, 32) | hb
print(np.sum(redo_bad_syn), "lost total neurons")
# -
# compare to output of call
plt.figure()
plt.title('XOR of function output and work in this notebook')
plt.imshow(bad_syns & redo_bad_syn | ~bad_syns & ~redo_bad_syn) # xor
| pystorm/test/test_determine_bad_syns.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# ... ***CURRENTLY UNDER DEVELOPMENT*** ...
#
# ## Validation of the synthetic waves and level
#
# inputs required:
# * historical wave conditions
# * emulator output - synthetic wave conditions
# * emulator output - synthetic wave conditions associated to the future TCs probability
# * Projected Sea Level Rise at the Site for the intermediate SLR scenario, +1m (to take into account in the TWL)
#
# in this notebook:
# * Comparison of the extreme distributions
# * Analysis of the DWT resposible of extreme TWL events (from simulation with SLR and simulation with future TCs prob. and SLR)
#
#
# +
# #!/usr/bin/env python
# -*- coding: utf-8 -*-
# common
import os
import os.path as op
# pip
import numpy as np
import xarray as xr
from datetime import datetime
import matplotlib.pyplot as plt
# DEV: override installed teslakit
import sys
sys.path.insert(0, op.join(os.path.abspath(''), '..', '..', '..', '..'))
# teslakit
from teslakit.database import Database
from teslakit.climate_emulator import Climate_Emulator
from teslakit.extremes import Peaks_Over_Threshold as POT
from teslakit.util.time_operations import xds_reindex_daily
from teslakit.plotting.extremes import Plot_ReturnPeriodValidation_CC
from teslakit.plotting.estela import Plot_DWTs_Probs
from teslakit.plotting.wts import Plot_Probs_WT_WT
from teslakit.plotting.outputs import Plot_LevelVariables_Histograms
# -
#
# ## Database and Site parameters
# +
# --------------------------------------
# Teslakit database
p_data = r'/media/administrador/HD/Dropbox/Guam/teslakit/data'
# p_data=r'/Users/laurac/Dropbox/Guam/teslakit/data'
# offshore
db = Database(p_data)
db.SetSite('GUAM')
# climate change - S4
db_S4 = Database(p_data)
db_S4.SetSite('GUAM_CC_S4')
# +
# --------------------------------------
# Load complete hourly data for extremes analysis
# Historical
HIST_C_h = db.Load_HIST_OFFSHORE(decode_times=True)
# Simulation
SIM_C_h = db.Load_SIM_OFFSHORE_all(decode_times=True, use_cftime=True)
# Simulation climate change
SIM_C_h_CChange = db_S4.Load_SIM_OFFSHORE_all(decode_times=True, use_cftime=True)
# +
# --------------------------------------
# Load SLR file
SLR = db.Load_SeaLevelRise()
SLR['time'] = SLR.time.dt.round('H')
# Select intermediate scenario (+1.0)
SLR = SLR.sel(scenario='1.0')
# +
# def. some auxiliar function to select all dataset variables at vn max by groups
def grouped_max(ds, vn=None, dim=None):
return ds.isel(**{dim: ds[vn].argmax(dim)})
# -
#
# ## AWL - Annual Maxima Calculation
# +
# Historical AWL Annual Maxima
# remove nans before and after AWL
ix_nonan = np.squeeze(np.argwhere(~np.isnan(HIST_C_h['AWL'].values[:])))
HIST_C_nonan = HIST_C_h.isel(time = ix_nonan)
# calculate AWL annual maxima dataset
hist_AMax = HIST_C_nonan.groupby('time.year').apply(grouped_max, vn='AWL', dim='time')
# +
# Simulation AWL Annual Maxima
# calculate AWL annual maxima dataset
sim_AMax = SIM_C_h.groupby('time.year').apply(grouped_max, vn='AWL', dim='time')
# +
# Simulation climate change AWL Annual Maxima
# calculate AWL annual maxima dataset
sim_AMax_CChange = SIM_C_h_CChange.groupby('time.year').apply(grouped_max, vn='AWL', dim='time')
# -
#
# ## AWL - Annual Maxima Return Period
# +
# AWL Annual Maxima Return Period (historical vs. simulations)
Plot_ReturnPeriodValidation_CC(hist_AMax['AWL'], sim_AMax['AWL'].transpose(), sim_AMax_CChange['AWL'].transpose(), label_1='Simulation', label_2 = 'Simulation TCs');
# +
# Hs and Tp Return Period (historical vs. simulations) at AWL Annual Maxima
# Hs at AWL Annual Maxima
Plot_ReturnPeriodValidation_CC(hist_AMax['Hs'], sim_AMax['Hs'].transpose(), sim_AMax_CChange['Hs'].transpose(), label_1='Simulation', label_2 = 'Simulation TCs');
# Tp at AWL Annual Maxima
Plot_ReturnPeriodValidation_CC(hist_AMax['Tp'], sim_AMax['Tp'].transpose(), sim_AMax_CChange['Tp'].transpose(), label_1='Simulation', label_2 = 'Simulation TCs');
# -
#
# ## AWL - Annual Maxima Probabilistic Plots
# +
# Probabilistic plots parameters
n_clusters_AWT = 6 # number of AWT clusters
n_clusters_DWT = 42 # number of DWT clusters
# Select one simulation DWTs - WAVEs simulation
n_sim = 0
sim_AMax_n = sim_AMax.sel(n_sim=0)
sim_AMax_n_CChange = sim_AMax_CChange.sel(n_sim=0)
# +
# Plot Annual Maxima DWT probabilities by month
# Simulation
Plot_DWTs_Probs(sim_AMax_n['DWT'].values, sim_AMax_n.time.values, n_clusters_DWT);
# Simulation climate change
Plot_DWTs_Probs(sim_AMax_n_CChange['DWT'].values, sim_AMax_n_CChange.time.values, n_clusters_DWT);
# +
# Plot Annual Maxima AWTs/DWTs Probabilities
# Simulation
Plot_Probs_WT_WT(
sim_AMax_n['AWT'].values[:] - 1, sim_AMax_n['DWT'].values[:] - 1,
n_clusters_AWT, n_clusters_DWT, wt_colors=True, ttl = 'Simulation',
);
# Simulation climate change
Plot_Probs_WT_WT(
sim_AMax_n_CChange['AWT'].values[:] - 1, sim_AMax_n_CChange['DWT'].values[:] - 1,
n_clusters_AWT, n_clusters_DWT, wt_colors=True, ttl = 'Simulation Climate Change (TCs)',
);
# -
#
# ## AWL - Peaks Over Threshold Calculation
# +
# POT plots parameters
n_clusters_AWT = 6 # number of AWT clusters
n_clusters_DWT = 42 # number of DWT clusters
# Select one simulation DWTs - WAVEs simulation
n_sim = 0
SIM_C_h_n = SIM_C_h.sel(n_sim=0)
SIM_C_h_n_CChange = SIM_C_h_CChange.sel(n_sim=0)
# TODO: update POT to work with hourly data
_, ix = np.unique(SIM_C_h_n['time'], return_index=True)
SIM_C_h_n = SIM_C_h_n.isel(time=ix)
_, ix = np.unique(SIM_C_h_n_CChange['time'], return_index=True)
SIM_C_h_n_CChange = SIM_C_h_n_CChange.isel(time=ix)
# Parse data to daily
SIM_C_d_n = xds_reindex_daily(SIM_C_h_n)# TODO: check possible bug if this puts NAN inside AWL data
SIM_C_d_n_CChange = xds_reindex_daily(SIM_C_h_n_CChange)# TODO: check possible bug if this puts NAN inside AWL data
# -
SIM_C_d_n_CChange
# +
# Get independent event peaks (99 percentile)
# Simulation
times_POT = POT(SIM_C_d_n, 'AWL', 99).time
sim_POT = SIM_C_d_n.sel(time = times_POT)
# Simulation climate change
SIM_C_d_n_CChange.AWL[np.where(np.isnan(SIM_C_d_n_CChange.AWL)==True)[0]]=np.nanmean(SIM_C_d_n_CChange.AWL)
times_POT_CChange = POT(SIM_C_d_n_CChange, 'AWL', percentile=99).time
sim_POT_CChange = SIM_C_d_n_CChange.sel(time = times_POT)
# -
#
# ## AWL - Peaks Over Threshold Probabilistic Plots
# +
# Plot Peaks Over Threshold DWT probabilities by month
# Simulation
Plot_DWTs_Probs(sim_POT['DWT'].values, sim_POT.time.values, n_clusters_DWT);
# Simulation climate change
Plot_DWTs_Probs(sim_POT_CChange['DWT'].values, sim_POT_CChange.time.values, n_clusters_DWT);
# +
# Plot Peaks Over Threshold AWTs/DWTs Probabilities
# Simulation
Plot_Probs_WT_WT(
sim_POT['AWT'].values[:] - 1, sim_POT['DWT'].values[:] - 1,
n_clusters_AWT, n_clusters_DWT, wt_colors=True, ttl = 'Simulation',
);
# Simulation climate change
Plot_Probs_WT_WT(
sim_POT_CChange['AWT'].values[:] - 1, sim_POT_CChange['DWT'].values[:] - 1,
n_clusters_AWT, n_clusters_DWT, wt_colors=True, ttl = 'Simulation Climate Change (TCs)',
);
# -
#
# ## Level Variables (level, AT, MMSL, TWL) - Histograms
# +
# Plot level, AT, MMSL and TWL histogram comparison between historical and simulated data
Plot_LevelVariables_Histograms(SIM_C_h.sel(n_sim = 0), SIM_C_h_CChange.sel(n_sim = 0), label_1='Simulation', label_2 = 'Simulation TCs', );
# -
# # Level variables taking into account SLR in the TWL (100 yrs. of simulated data)
#
# +
# Plot level variables of climate change simulated data: SLR vs ENSO+SLR
# select 100 yrs of simulation
SIM_C_h_SLR = SIM_C_h.isel(time=slice(0, len(SLR.time))).copy(deep=True) # 100 years
SIM_C_h_CChange_SLR = SIM_C_h_CChange.isel(time=slice(0, len(SLR.time))).copy(deep=True) # 100 years
# Add SLR to TWL
for n in range(0, len(SIM_C_h_SLR.n_sim)):
SIM_C_h_SLR['TWL'][:,n] = SIM_C_h_SLR['TWL'].isel(n_sim=n) + SLR['SLR'].values
SIM_C_h_CChange_SLR['TWL'][:,n] = SIM_C_h_CChange_SLR['TWL'].isel(n_sim=n) + SLR['SLR'].values
# Plot level, AT, MMSL and TWL histogram comparison between historical and simulated data
Plot_LevelVariables_Histograms(SIM_C_h_SLR.sel(n_sim = 0), SIM_C_h_CChange_SLR.sel(n_sim = 0), label_1='Simulation SLR', label_2 = 'Simulation TCs & SLR');
# -
#
# ## TWL - Annual Maxima. Simulations SLR vs. simulations SLR+TCs
# +
# Plot TWL annual maxima
# calculate Annual Maxima values for historical and simulated data
hist_A = HIST_C_h['TWL'].groupby('time.year').max(dim='time')
sim_A = SIM_C_h_SLR['TWL'].groupby('time.year').max(dim='time')
sim_B = SIM_C_h_CChange_SLR['TWL'].groupby('time.year').max(dim='time')
# Return Period historical vs. simulations
Plot_ReturnPeriodValidation_CC(hist_A, sim_A.transpose(), sim_B.transpose(), label_1='Simulation SLR', label_2 = 'Simulation TCs & SLR');
# -
| notebooks/GUAM/GUAM/03_ClimateChange/S4_SLR_TCs/02_14_Validation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # MediZen Strain Recommendation API Model
#
# ## Version 1.1 - 2019-11-19
# ---
#
# ## Imports and Config
# General imports
import pandas as pd
import janitor
import os
# NLP Imports
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.neighbors import NearestNeighbors
# Configure pandas to display entire text of column
pd.set_option('max_colwidth', 200)
pd.set_option('max_columns', 200) # Display up to 200 columns
# ---
#
# ## Data Loading and First Looks
# +
# Load the data into pd.DataFrame
filepath = "/Users/Tobias/workshop/buildbox/medizen_ds_api/data/"
data_filename = "cannabis.csv"
data_filepath = os.path.join(filepath, data_filename)
df1 = pd.read_csv(data_filepath)
# -
df1.head()
# ---
#
# ## Data Wrangling and Feature Engineering
#
# The end result that should be passed into the model is a single long string.
# Therefore, the new feature will simply be a concatenation of all three current features:
#
# - `type`
# - `effects`
# - `flavor`
#
# The cell below uses [pyjanitor](pyjanitor.readthedocs.io/) method-chaining to:
#
# 1. Clean up the feature names, which in this case only makes them lowercase
# 2. Concatenate the three features into one one, comma-separated feature
# 3. Remove all of the features except the new one
# +
# User pyjanitor to wrangle the data
df2 = (df1
.clean_names() # In this case, fixes Title Case
.concatenate_columns(
# Explanation above - create one feature for NLP analysis
column_names=["type", "effects", "flavor"],
new_column_name="type_effects_flavor",
sep=",", # Staying consistent with comma-separation
)
.remove_columns(column_names=[
"rating",
"description",
"type",
"effects",
"flavor",
]))
# -
# Look at the resulting dataframe
print(df2.shape)
df2.head()
# Look at null values - hint: there shouldn't be any
# because they all have values in at least one of the three columns
df2.isnull().sum()
# ---
#
# ## TF-IDF
#
# TF-IDF is a method of finding unique aspects of documents (strings).
# The more common a word is across the documents the lower the score.
# The result is the unique topics rising to the top. (They are called _top_-ics, after all)...
# +
# Instantiate the vectorizer object
tfidf = TfidfVectorizer(stop_words="english")
# Create a vocabulary from the new feature
dtm = tfidf.fit(df2["type_effects_flavor"])
# Create vectorized version of wrangled dataframe
sparse = tfidf.transform(df2["type_effects_flavor"])
# The result is a sparse matrix, which can be converted back to a dataframe
vdtm = pd.DataFrame(sparse.todense(), columns=tfidf.get_feature_names())
# -
# ## Get Similarities with Nearest Neighbor (K-NN)
#
# In order to get a list of strains that are similar to a given input string,
# the K-Nearest Neighbor model will be used. This model uses a tree-based approach
# to calculate the distances between points and recursively clusters them until
# to find the desired number (k) of neighboring data points.
# +
# Instantiate the knn model
nn = NearestNeighbors(n_neighbors=10, algorithm='ball_tree')
# Fit (train) the model on the TF-IDF vector dataframe created above
nn.fit(vdtm)
# -
# ### Running the KNN Model, Part 1
# Example string to demonstrate the model
# This is a realistic example of what will be passed
# into the model once it is integrated into the API
ex_1_str = "indica,happy,relaxed,hungry,talkative,citrus,tangy,flowery"
# Create the input vector
ex_1_vec = tfidf.transform([ex_1_str])
# Pass that vector into the trained knn model, specifying the number of neighbors to return
# This returns a list of two arrays: one is a measure of each neighbors 'near-ness'
# the other (the one we want) contains the indexes for the neighbors
rec_array = nn.kneighbors(ex_1_vec.todense(), n_neighbors=10)
rec_array
# Extract the second array - the list of strain ids (indexes) that are 'closest' to input
rec_id_list = rec_array[1][0]
rec_id_list
# Although the API will return only this list of indexes,
# for the purposes of this demo I'll hydrate that list with the rest
# of the strain data from the original (pre-wrangled) dataframe
recommendations = df1.iloc[rec_id_list]
recommendations
# ---
#
# ## The FuncZone
#
# In order to more easily integrate this recommendation process into the Flask API,
# the steps can be grouped into a function that will take in a request and return the recommendations.
def recommend(req, n=10):
"""Function to recommend top n strains given a request."""
# Create vector from request
req_vec = tfidf.transform([req])
# Access the top n indexes
top_id = nn.kneighbors(req_vec.todense(), n_neighbors=n)[1][0]
# Index-locate the neighbors in original dataframe
top_df = df1.iloc[top_id]
return top_df
# +
# Another example request to test out the above function
ex_2_str = "hybrid,euphoric,energetic,creative,woody,earthy"
# Run the function, this time asking for the top 5 recommendations
ex_2_recs = recommend(ex_2_str, 5)
ex_2_recs
# -
# The API should return a JSON object with only the ids
# Here's a slightly modified version to accomplish that
def recommend_json(req, n=10):
"""Function to recommend top n strains given a request."""
# Create vector from request
req_vec = tfidf.transform([req])
# Access the top n indexes
rec_id = nn.kneighbors(req_vec.todense(), n_neighbors=n)[1][0]
# Convert np.ndarray to pd.Series then to JSON
rec_json = pd.Series(rec_id).to_json(orient="records")
return rec_json
# ## Pickling
#
# In order to use the model in the Flask app, it can be pickled.
# The pickle module, and the pickle file format, allows Python objects
# to be serialized and de-serialized. In this case, the trained vectorizer
# and model can be made into pickle files, which are then loaded into the
# Flask app for use in the recommendation API.
# Create pickle func to make pickling (a little) easier
def picklizer(to_pickle, filename, path):
"""
Creates a pickle file.
Parameters
----------
to_pickle : Python object
The trained / fitted instance of the
transformer or model to be pickled.
filename : string
The desired name of the output file,
not including the '.pkl' extension.
path : string or path-like object
The path to the desired output directory.
"""
import os
import pickle
# Create the path to save location
picklepath = os.path.join(path, filename)
# Use context manager to open file
with open(picklepath, "wb") as p:
pickle.dump(to_pickle, p)
# +
# Picklize!
# Export vectorizer as pickle
picklizer(dtm, "vect_02.pkl", filepath)
# Export knn model as pickle
picklizer(nn, "knn_02.pkl", filepath)
# -
# ### Running the KNN Model, Part 2 (this time with pickles)
# +
import pickle
# Load the vdtm pickle into new object for testing
vv_path = os.path.join(filepath, "vect_02.pkl")
# Use context manager to open and load pickle file
with open(vv_path, "rb") as p:
vv = pickle.load(p)
# +
# Load the knn pickle into new object for testing
knn_path = os.path.join(filepath, "knn_02.pkl")
# Use context manager to open and load pickle file
with open(knn_path, "rb") as p:
knn = pickle.load(p)
# +
# Another slightly modified version that uses test objects
def rec_pickle(req, n=10):
"""Function to recommend top n strains given a request."""
# Create vector from request
req_vec = vv.transform([req])
# Access the top n indexes
rec_id = knn.kneighbors(req_vec.todense(), n_neighbors=n)[1][0]
# Convert np.ndarray to pd.Series then to JSON
rec_json = pd.Series(rec_id).to_json(orient="records")
return rec_json
# +
# Test out the modified function / pickle objects
ex_3_str = "sativa,energetic,talkative,euphoric,creative,focused,spicy,tangy,sweet"
# This time I'll pass in 8, because why not?
ex_3_rec = rec_pickle(ex_3_str, 8)
ex_3_rec
# -
| docs/notebooks/03-medizen-recommendations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
import os
os.chdir('..')
import os
os.environ["CUDA_VISIBLE_DEVICES"]="1,2"
# +
dataset_params = {
'dataset': 'dsprites',
'batch_size': 32,
'cuda': True,
'root': 'data'
}
from src.utils import get_datasets
train_loader, test_loader, (width, height, channels) = get_datasets(
**dataset_params)
# +
# %%time
from src.models.conv_vsc import ConvolutionalVariationalSparseCoding
model_params = {
'dataset': 'dsprites',
'width': width,
'height': height,
'channels': channels,
'kernel_szs': '32,32,64,64',
'hidden_sz': 256,
'latent_sz': 100,
'learning_rate': 3e-4,
'alpha': 0.01,
'beta': 2,
'beta_delta': 0,
'c': 50,
'c_delta': 1e-3,
'device': 'cuda',
'log_interval': 5000,
'normalize': False,
'flatten': False
}
convvsc = ConvolutionalVariationalSparseCoding(**model_params)
# -
training_params = {
'train_loader': train_loader,
'test_loader': test_loader,
'epochs': 15,
'report_interval': 3,
'sample_sz': 64,
'reload_model': False,
'checkpoints_path': 'results/checkpoints',
'logs_path': 'results/logs',
'images_path': 'results/images'
}
# +
import logging
for handler in logging.root.handlers[:]:
logging.root.removeHandler(handler)
logging.basicConfig(
filename='notebooks/train_dsprites.log',
format='%(asctime)s - %(levelname)s - %(message)s',
datefmt='%d-%b-%y %H:%M:%S',
level=logging.INFO
)
# -
# %%time
try:
convvsc.run_training(logging_func=logging.info,
**training_params)
except:
logging.exception("Exception occurred")
train_losses = pd.DataFrame(convvsc.train_losses)
train_losses.tail()
train_losses[['LOSS', 'BCE', 'PRIOR']].rolling(window=1000).mean().plot();
# %%time
try:
convvsc.run_training(logging_func=logging.info, start_epoch=16,
**training_params)
except:
logging.exception("Exception occurred")
train_losses = pd.DataFrame(convvsc.train_losses)
train_losses.tail()
train_losses[['LOSS', 'BCE', 'PRIOR']].rolling(window=5000).mean().plot();
# ## 90 epochs
training_params = {
'train_loader': train_loader,
'test_loader': test_loader,
'epochs': 60,
'report_interval': 5,
'sample_sz': 64,
'reload_model': False,
'checkpoints_path': 'results/checkpoints',
'logs_path': 'results/logs',
'images_path': 'results/images'
}
# %%time
try:
convvsc.run_training(logging_func=logging.info, start_epoch=31,
**training_params)
except:
logging.exception("Exception occurred")
train_losses = pd.DataFrame(convvsc.train_losses)
train_losses.tail()
train_losses[['LOSS', 'BCE', 'PRIOR']].iloc[:1550000] \
.rolling(window=5000).mean().plot(figsize=(10,5));
train_losses[['PRIOR']].iloc[:1550000] \
.rolling(window=5000).mean().plot(figsize=(10,5));
| notebooks/Retrain - Conv - dSprites.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Ridge (L2) / Lasso (L1) / ElasticNet Regression
#
# ______
# ## Environment Set-Up
#
# ### Load relevant Python Packages
# +
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.linear_model import Ridge
from sklearn.linear_model import Lasso
from sklearn.linear_model import ElasticNet
from sklearn.metrics import mean_squared_error, r2_score
# -
# ### Data Import
#data has been saved using a .pkl file
path = './data/df_model.pkl'
df_model = pd.read_pickle(path)
df_model.head(2)
# ### Setting Up Training & Test Dataframes
#
# The dataframe is split into a training set (80%) and a test set (20%).
# define features and target
X = df_model.drop('power_W', axis=1)
y = df_model.power_W
# train-test-split
X_train, X_test, y_train, y_test = train_test_split(X, y,test_size = .2, random_state=105)
# Let's check the shape of our dataframes
print(X_train.shape)
print(y_train.shape)
print(X_test.shape)
print(y_test.shape)
# General information: Ridge Regression and Lasso Regression can be used to reduce model complexity and prevent overfitting which may result from simple linear regression.
# ## Ridge Regression (L2 regularization)
# A commenly used alternative to the "normal" linear regression model is Ridge regression. It is also a linear model that uses basically the same formula that is used for ordinary least squares. However, our ridge regression model will also try to keep the magnitude of coefficients to be as small as possible. In other words, all entries of w should be close to zero. We can also say, each feature should have as little effect on the outcome as possible while still predicting well.
#
# Performing L2 regularization with different alpha values:
# initialize and train model with (default value) alpha = 1
ridge = Ridge(alpha=1)
ridge.fit(X_train, y_train)
# predict on test-set
y_pred_ridge = ridge.predict(X_test)
# R-squared scores for train and test set
train_score_ridge = ridge.score(X_train, y_train)
test_score_ridge = ridge.score(X_test, y_test)
print("Train score: {:.2f}".format(train_score_ridge))
print("Test score: {:.2f}".format(test_score_ridge))
# RMSE of test set
print("MSE:", round((mean_squared_error(y_test, y_pred_ridge)), 3))
# initialize and train model with alpha = 10
ridge_10 = Ridge(alpha=10)
ridge_10.fit(X_train, y_train)
# predict on test-set
y_pred_ridge_10 = ridge_10.predict(X_test)
# R-squared scores for train and test set
train_score_ridge_10 = ridge_10.score(X_train, y_train)
test_score_ridge_10 = ridge_10.score(X_test, y_test)
print("Train score: {:.2f}".format(train_score_ridge_10))
print("Test score: {:.2f}".format(test_score_ridge_10))
# RMSE of test set
print("MSE:", round(mean_squared_error(y_test, y_pred_ridge_10), 3))
# initialize and train model with alpha = 0.1
ridge_01 = Ridge(alpha=0.1)
ridge_01.fit(X_train, y_train)
# predict on test-set
y_pred_ridge_01 = ridge_01.predict(X_test)
# R-squared scores for train and test set
train_score_ridge_01 = ridge_01.score(X_train, y_train)
test_score_ridge_01 = ridge_01.score(X_test, y_test)
print("Train score: {:.2f}".format(train_score_ridge_01))
print("Test score: {:.2f}".format(test_score_ridge_01))
# RMSE of test set
print("MSE:", round(mean_squared_error(y_test, y_pred_ridge_01), 3))
# ## Lasso Regression (L1 regularization)
# An alternative to Ridge is Lasso regression. Similarly to ridge regression lasso restricts coefficients to be close to zero. It does so in a slightly different way so that when using lasso some coefficients become exactly zero. This means some features are entirely ignored by the model. It can be seen as an automatic feature selection which makes models often easier to interpret and can reveal the most important features.
#
# Performing L1 regularization with different alpha values:
# +
# initialize and train model with (default value) alpha = 1.0
lasso = Lasso(alpha=1, max_iter=10e5)
lasso.fit(X_train,y_train)
# predict on test-set
y_pred_lasso = lasso.predict(X_test)
# R-squared scores for train and test set
train_score_lasso = lasso.score(X_train, y_train)
test_score_lasso = lasso.score(X_test, y_test)
print("Train score: {:.2f}".format(train_score_lasso))
print("Test score: {:.2f}".format(test_score_lasso))
# RMSE of test set
print("MSE:", round(mean_squared_error(y_test, y_pred_lasso), 3))
# number of features used
coeff_used = np.sum(lasso.coef_!=0)
print("# features: ", coeff_used)
# -
# +
# initialize and train model with alpha 0.01
# We'll also increase the amount of max_iter otherwise it will raise a warning.
lasso_01 = Lasso(alpha=0.01, max_iter=1000000)
lasso_01.fit(X_train,y_train)
# predict on test-set
y_pred_lasso_01 = lasso_01.predict(X_test)
# R-squared scores for train and test set
train_score_lasso_01 = lasso_01.score(X_train, y_train)
test_score_lasso_01 = lasso_01.score(X_test, y_test)
print("Train score: {:.2f}".format(train_score_lasso_01))
print("Test score: {:.2f}".format(test_score_lasso_01))
# RMSE of test set
print("MSE:", round(mean_squared_error(y_test, y_pred_lasso_01), 3))
# number of features used
coeff_used = np.sum(lasso_01.coef_!=0)
print("# features: ", coeff_used)
# -
# +
# initialize and train model with alpha 0.0001
lasso_0001 = Lasso(alpha=0.0001, max_iter=1000000)
lasso_0001.fit(X_train,y_train)
# predict on test-set
y_pred_lasso_0001 = lasso_0001.predict(X_test)
# R-squared scores for train and test set
train_score_lasso_0001 = lasso_0001.score(X_train, y_train)
test_score_lasso_0001 = lasso_0001.score(X_test, y_test)
print("Train score: {:.2f}".format(train_score_lasso_0001))
print("Test score: {:.2f}".format(test_score_lasso_0001))
# RMSE of test set
print("MSE:", round(mean_squared_error(y_test, y_pred_lasso_0001), 3))
# number of features used
coeff_used = np.sum(lasso_0001.coef_!=0)
print("# features: ", coeff_used)
# -
# ## ElasticNet Regression
# The ElasticNet regression combines the penalties of Lasso and Ridge. Often this combination works best, though at the price of having two parameters to adjust: one for the L1 regularization, and one for the L2 regularization.
# +
# initialize and train model with (default value) alpha = 1.0
elastic = ElasticNet(alpha=1, l1_ratio=0.6, max_iter=10e5)
elastic.fit(X_train,y_train)
# predict on test-set
y_pred_elastic = elastic.predict(X_test)
# R-squared scores for train and test set
train_score_elastic = elastic.score(X_train, y_train)
test_score_elastic = elastic.score(X_test, y_test)
print("Train score: {:.2f}".format(train_score_elastic))
print("Test score: {:.2f}".format(test_score_elastic))
# RMSE of test set
print("MSE:", round(mean_squared_error(y_test, y_pred_elastic)), 3)
# number of features used
coeff_used = np.sum(elastic.coef_!=0)
print("# features: ", coeff_used)
print('This cell was last run on: ')
print(datetime.now())
# -
# ----
# ## Conclusion
# Now you saw three different ways to prevent a linear regression model from overfitting and for prediction. In practice, ridge regression is usually the first choice. But if there are a large amount of features and want to improve the interpretability of a model it makes sense to go for lasso regression since it will eliminate some of your features.
# With the ElasticNet both can be combined.
# All in all and with a look on the outcome for the three typs - the results are not satisfying!
# For this reason go on with: [CyPer_TensorFlow](CyPer_TensorFlow.ipynb).
| CyPer_Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# dependencies
import tensorflow as tf
import numpy as np
from sklearn.cross_validation import train_test_split
import time
import matplotlib.pyplot as plt
import pickle
def read_dataset(filepath):
with open(filepath, 'rb') as fp:
return pickle.load(fp)
# read dataset
dataset_location = "./data.p"
X, Y, l1_word2idx, l1_idx2word, l1_vocab, l2_word2idx, l2_idx2word, l2_vocab = read_dataset(dataset_location)
print(l1_vocab[:30])
print(l2_vocab[:30])
# +
# data processing
# data padding
def data_padding(x, y, length = 20):
for i in range(len(x)):
x[i] = x[i] + (length - len(x[i])) * [l1_word2idx['<pad>']]
y[i] = [l2_word2idx['<go>']] + y[i] + [l2_word2idx['<eos>']] + (length-len(y[i])) * [l2_word2idx['<pad>']]
data_padding(X, Y)
# data splitting
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.1)
del X
del Y
# +
# build a model
input_seq_len = 20
output_seq_len = 22
l1_vocab_size = len(l1_vocab) + 2 # + <pad>, <ukn>
l2_vocab_size = len(l2_vocab) + 4 # + <pad>, <ukn>, <eos>, <go>
# placeholders
encoder_inputs = [tf.placeholder(dtype = tf.int32, shape = [None], name = 'encoder{0}'.format(i)) for i in range(input_seq_len)]
decoder_inputs = [tf.placeholder(dtype = tf.int32, shape = [None], name = 'decoder{0}'.format(i)) for i in range(output_seq_len)]
targets = [decoder_inputs[i+1] for i in range(output_seq_len-1)]
# add one more target
targets.append(tf.placeholder(dtype = tf.int32, shape = [None], name = 'last_target'))
target_weights = [tf.placeholder(dtype = tf.float32, shape = [None], name = 'target_w{0}'.format(i)) for i in range(output_seq_len)]
# output projection
size = 512
w_t = tf.get_variable('proj_w', [l2_vocab_size, size], tf.float32)
b = tf.get_variable('proj_b', [l2_vocab_size], tf.float32)
w = tf.transpose(w_t)
output_projection = (w, b)
outputs, states = tf.nn.seq2seq.embedding_attention_seq2seq(
encoder_inputs,
decoder_inputs,
tf.nn.rnn_cell.BasicLSTMCell(size),
num_encoder_symbols = l1_vocab_size,
num_decoder_symbols = l2_vocab_size,
embedding_size = 80,
feed_previous = False,
output_projection = output_projection,
dtype = tf.float32)
# +
# define our loss function
# sampled softmax loss - returns: A batch_size 1-D tensor of per-example sampled softmax losses
def sampled_loss(logits, labels):
return tf.nn.sampled_softmax_loss(
weights = w_t,
biases = b,
labels = tf.reshape(labels, [-1, 1]),
inputs = logits,
num_sampled = 512,
num_classes = l2_vocab_size)
# Weighted cross-entropy loss for a sequence of logits
loss = tf.nn.seq2seq.sequence_loss(outputs, targets, target_weights, softmax_loss_function = sampled_loss)
# +
# simple softmax function
def softmax(x):
n = np.max(x)
e_x = np.exp(x - n)
return e_x / e_x.sum()
# feed data into placeholders
def feed_dict(x, y, batch_size = 64):
feed = {}
idxes = np.random.choice(len(x), size = batch_size, replace = False)
for i in range(input_seq_len):
feed[encoder_inputs[i].name] = np.array([x[j][i] for j in idxes])
for i in range(output_seq_len):
feed[decoder_inputs[i].name] = np.array([y[j][i] for j in idxes])
feed[targets[len(targets)-1].name] = np.full(shape = [batch_size], fill_value = l2_word2idx['<pad>'])
for i in range(output_seq_len-1):
batch_weights = np.ones(batch_size, dtype = np.float32)
target = feed[decoder_inputs[i+1].name]
for j in range(batch_size):
if target[j] == l2_word2idx['<pad>']:
batch_weights[j] = 0.0
feed[target_weights[i].name] = batch_weights
feed[target_weights[output_seq_len-1].name] = np.zeros(batch_size, dtype = np.float32)
return feed
# decode output sequence
def decode_output(output_seq):
words = []
for i in range(output_seq_len):
smax = softmax(output_seq[i])
idx = np.argmax(smax)
words.append(l2_idx2word[idx])
return words
# +
# ops and hyperparameters
learning_rate = 3e-3
batch_size = 8
steps = 40000
# ops for projecting outputs
outputs_proj = [tf.matmul(outputs[i], output_projection[0]) + output_projection[1] for i in range(output_seq_len)]
# training op
optimizer = tf.train.RMSPropOptimizer(learning_rate, decay=0.99).minimize(loss)
# init op
init = tf.global_variables_initializer()
# forward step
def forward_step(sess, feed):
output_sequences = sess.run(outputs_proj, feed_dict = feed)
return output_sequences
# training step
def backward_step(sess, feed):
sess.run(optimizer, feed_dict = feed)
# +
# we will use this list to plot losses through steps
losses = []
# save a checkpoint so we can restore the model later
saver = tf.train.Saver()
print('------------------TRAINING------------------')
with tf.Session() as sess:
sess.run(init)
t = time.time()
for step in range(steps):
feed = feed_dict(X_train, Y_train)
backward_step(sess, feed)
if step % 50 == 49 or step == 0:
loss_value = sess.run(loss, feed_dict = feed)
print('step: {}, loss: {}'.format(step, loss_value))
losses.append(loss_value)
if step % 1000 == 999:
saver.save(sess, './checkpoints/', global_step=step)
print('Checkpoint is saved')
print('Training time for {} steps: {}s'.format(steps, time.time() - t))
# +
# plot losses
with plt.style.context('fivethirtyeight'):
plt.plot(losses, linewidth = 1)
plt.xlabel('Steps')
plt.ylabel('Losses')
plt.ylim((0, 12))
plt.show()
# -
encoded_test = {'X':X_test, 'Y':Y_test}
pickle.dump(encoded_test, open("./encoded_test.p", mode='wb'))
| Hindi to English/Model Train.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_tensorflow_p36
# language: python
# name: conda_tensorflow_p36
# ---
# # TensorFlow script mode training and serving
#
# Script mode is a training script format for TensorFlow that lets you execute any TensorFlow training script in SageMaker with minimal modification. The [SageMaker Python SDK](https://github.com/aws/sagemaker-python-sdk) handles transferring your script to a SageMaker training instance. On the training instance, SageMaker's native TensorFlow support sets up training-related environment variables and executes your training script. In this tutorial, we use the SageMaker Python SDK to launch a training job and deploy the trained model.
#
# Script mode supports training with a Python script, a Python module, or a shell script. In this example, we use a Python script to train a classification model on the [MNIST dataset](http://yann.lecun.com/exdb/mnist/). In this example, we will show how easily you can train a SageMaker using TensorFlow 1.x and TensorFlow 2.0 scripts with SageMaker Python SDK. In addition, this notebook demonstrates how to perform real time inference with the [SageMaker TensorFlow Serving container](https://github.com/aws/sagemaker-tensorflow-serving-container). The TensorFlow Serving container is the default inference method for script mode. For full documentation on the TensorFlow Serving container, please visit [here](https://github.com/aws/sagemaker-python-sdk/blob/master/src/sagemaker/tensorflow/deploying_tensorflow_serving.rst).
#
# # Set up the environment
#
# Let's start by setting up the environment:
# +
import os
import sagemaker
from sagemaker import get_execution_role
sagemaker_session = sagemaker.Session()
role = get_execution_role()
region = sagemaker_session.boto_session.region_name
# -
# ## Training Data
#
# The MNIST dataset has been loaded to the public S3 buckets ``sagemaker-sample-data-<REGION>`` under the prefix ``tensorflow/mnist``. There are four ``.npy`` file under this prefix:
# * ``train_data.npy``
# * ``eval_data.npy``
# * ``train_labels.npy``
# * ``eval_labels.npy``
training_data_uri = "s3://sagemaker-sample-data-{}/tensorflow/mnist".format(region)
# # Construct a script for distributed training
#
# This tutorial's training script was adapted from TensorFlow's official [CNN MNIST example](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/examples/tutorials/layers/cnn_mnist.py). We have modified it to handle the ``model_dir`` parameter passed in by SageMaker. This is an S3 path which can be used for data sharing during distributed training and checkpointing and/or model persistence. We have also added an argument-parsing function to handle processing training-related variables.
#
# At the end of the training job we have added a step to export the trained model to the path stored in the environment variable ``SM_MODEL_DIR``, which always points to ``/opt/ml/model``. This is critical because SageMaker uploads all the model artifacts in this folder to S3 at end of training.
#
# Here is the entire script:
# +
# !pygmentize 'mnist.py'
# TensorFlow 2.1 script
# !pygmentize 'mnist-2.py'
# -
# # Create a training job using the `TensorFlow` estimator
#
# The `sagemaker.tensorflow.TensorFlow` estimator handles locating the script mode container, uploading your script to a S3 location and creating a SageMaker training job. Let's call out a couple important parameters here:
#
# * `py_version` is set to `'py3'` to indicate that we are using script mode since legacy mode supports only Python 2. Though Python 2 will be deprecated soon, you can use script mode with Python 2 by setting `py_version` to `'py2'` and `script_mode` to `True`.
#
# * `distributions` is used to configure the distributed training setup. It's required only if you are doing distributed training either across a cluster of instances or across multiple GPUs. Here we are using parameter servers as the distributed training schema. SageMaker training jobs run on homogeneous clusters. To make parameter server more performant in the SageMaker setup, we run a parameter server on every instance in the cluster, so there is no need to specify the number of parameter servers to launch. Script mode also supports distributed training with [Horovod](https://github.com/horovod/horovod). You can find the full documentation on how to configure `distributions` [here](https://github.com/aws/sagemaker-python-sdk/tree/master/src/sagemaker/tensorflow#distributed-training).
#
#
# +
from sagemaker.tensorflow import TensorFlow
mnist_estimator = TensorFlow(
entry_point="mnist.py",
role=role,
instance_count=2,
instance_type="ml.p3.2xlarge",
framework_version="1.15.2",
py_version="py3",
distribution={"parameter_server": {"enabled": True}},
)
# -
# You can also initiate an estimator to train with TensorFlow 2.1 script. The only things that you will need to change are the script name and ``framewotk_version``
mnist_estimator2 = TensorFlow(
entry_point="mnist-2.py",
role=role,
instance_count=2,
instance_type="ml.p3.2xlarge",
framework_version="2.1.0",
py_version="py3",
distribution={"parameter_server": {"enabled": True}},
)
# ## Calling ``fit``
#
# To start a training job, we call `estimator.fit(training_data_uri)`.
#
# An S3 location is used here as the input. `fit` creates a default channel named `'training'`, which points to this S3 location. In the training script we can then access the training data from the location stored in `SM_CHANNEL_TRAINING`. `fit` accepts a couple other types of input as well. See the API doc [here](https://sagemaker.readthedocs.io/en/stable/estimators.html#sagemaker.estimator.EstimatorBase.fit) for details.
#
# When training starts, the TensorFlow container executes mnist.py, passing `hyperparameters` and `model_dir` from the estimator as script arguments. Because we didn't define either in this example, no hyperparameters are passed, and `model_dir` defaults to `s3://<DEFAULT_BUCKET>/<TRAINING_JOB_NAME>`, so the script execution is as follows:
# ```bash
# python mnist.py --model_dir s3://<DEFAULT_BUCKET>/<TRAINING_JOB_NAME>
# ```
# When training is complete, the training job will upload the saved model for TensorFlow serving.
mnist_estimator.fit(training_data_uri)
# Calling fit to train a model with TensorFlow 2.1 script.
mnist_estimator2.fit(training_data_uri)
# # Deploy the trained model to an endpoint
#
# The `deploy()` method creates a SageMaker model, which is then deployed to an endpoint to serve prediction requests in real time. We will use the TensorFlow Serving container for the endpoint, because we trained with script mode. This serving container runs an implementation of a web server that is compatible with SageMaker hosting protocol. The [Using your own inference code]() document explains how SageMaker runs inference containers.
predictor = mnist_estimator.deploy(initial_instance_count=1, instance_type="ml.p2.xlarge")
# Deployed the trained TensorFlow 2.1 model to an endpoint.
predictor2 = mnist_estimator2.deploy(initial_instance_count=1, instance_type="ml.p2.xlarge")
# # Invoke the endpoint
#
# Let's download the training data and use that as input for inference.
# +
import numpy as np
# !aws --region {region} s3 cp s3://sagemaker-sample-data-{region}/tensorflow/mnist/train_data.npy train_data.npy
# !aws --region {region} s3 cp s3://sagemaker-sample-data-{region}/tensorflow/mnist/train_labels.npy train_labels.npy
train_data = np.load("train_data.npy")
train_labels = np.load("train_labels.npy")
# -
# The formats of the input and the output data correspond directly to the request and response formats of the `Predict` method in the [TensorFlow Serving REST API](https://www.tensorflow.org/serving/api_rest). SageMaker's TensforFlow Serving endpoints can also accept additional input formats that are not part of the TensorFlow REST API, including the simplified JSON format, line-delimited JSON objects ("jsons" or "jsonlines"), and CSV data.
#
# In this example we are using a `numpy` array as input, which will be serialized into the simplified JSON format. In addtion, TensorFlow serving can also process multiple items at once as you can see in the following code. You can find the complete documentation on how to make predictions against a TensorFlow serving SageMaker endpoint [here](https://github.com/aws/sagemaker-python-sdk/blob/master/src/sagemaker/tensorflow/deploying_tensorflow_serving.rst#making-predictions-against-a-sagemaker-endpoint).
predictions = predictor.predict(train_data[:50])
for i in range(0, 50):
prediction = predictions["predictions"][i]["classes"]
label = train_labels[i]
print(
"prediction is {}, label is {}, matched: {}".format(prediction, label, prediction == label)
)
# Examine the prediction result from the TensorFlow 2.1 model.
predictions2 = predictor2.predict(train_data[:50])
for i in range(0, 50):
prediction = np.argmax(predictions2["predictions"][i])
label = train_labels[i]
print(
"prediction is {}, label is {}, matched: {}".format(prediction, label, prediction == label)
)
# # Delete the endpoint
#
# Let's delete the endpoint we just created to prevent incurring any extra costs.
predictor.delete_endpoint()
# Delete the TensorFlow 2.1 endpoint as well.
predictor2.delete_endpoint()
| sagemaker-python-sdk/tensorflow_script_mode_training_and_serving/tensorflow_script_mode_training_and_serving.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial #1: Train an raw sound classification model with Azure Machine Learning
# In this tutorial, you train a machine learning model on remote compute resources. You'll use the training and deployment workflow for Azure Machine Learning service (preview) in a Python Jupyter notebook. You can then use the notebook as a template to train your own machine learning model with your own data. This tutorial is part one of a two-part tutorial series.
#
# This tutorial trains a CNN model using raw sound dataset captured by [SoundCaptureModule](https://github.com/ms-iotkithol-jp/MicCaptureIoTSoundSample) on Azure IoT Edge device with Azure Machine Learning. Sound dataset consist from raw format and csv format file for each timespan. The goal is to create a multi-class classifier to identify the major or minor code of guiter equipment.
#
# Learn how to:
#
# - Set up your development environment
# - Access and examine the data
# - Train a CNN model on a remote cluster
# - Review training results, find and register the best model
# - You'll learn how to select a model and deploy it in part two of this tutorial later.
#
# ## Prerequisites
# See prerequisites in the Azure Machine Learning documentation.
# ## Set up your development environment
# All the setup for your development work can be accomplished in a Python notebook. Setup includes:
#
# - Importing Python packages
# - Connecting to a workspace to enable communication between your local computer and remote resources
# - Creating an experiment to track all your runs
# - Creating a remote compute target to use for training
# - Import packages
# - Import Python packages you need in this session. Also display the Azure Machine Learning SDK version.
# l-1
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
# +
import azureml.core
from azureml.core import Workspace
# check core SDK version number
print("Azure ML SDK Version: ", azureml.core.VERSION)
# -
# ## Connect to workspace
# Create a workspace object from the existing workspace. Workspace.from_config() reads the file config.json and loads the details into an object named ws.
# load workspace configuration from the config.json file in the current folder.
ws = Workspace.from_config()
print(ws.name, ws.location, ws.resource_group, sep='\t')
# ## Create experiment
# Create an experiment to track the runs in your workspace. A workspace can have muliple experiments.
# +
experiment_name = 'raw-sound-major-miner-cnn'
from azureml.core import Experiment
exp = Experiment(workspace=ws, name=experiment_name)
# -
# Specify dataset and testset names and positions
# +
# l-2
dataset_name = 'sound_csv_data'
testset_name = 'sound_csv_test'
data_folder_path = 'data-csv'
test_folder_path = 'test-csv'
soundDataDefFile = 'sounddata-csv.yml'
dataSrorageConfigFile = 'data-storage-config.yml'
# -
# ## Create or Attach existing compute resource
# By using Azure Machine Learning Compute, a managed service, data scientists can train machine learning models on clusters of Azure virtual machines. Examples include VMs with GPU support. In this tutorial, you create Azure Machine Learning Compute as your training environment. The code below creates the compute clusters for you if they don't already exist in your workspace.
#
# <b>Creation of compute takes approximately 5 minutes</b>. If the AmlCompute with that name is already in your workspace the code will skip the creation process.
# +
from azureml.core.compute import AmlCompute
from azureml.core.compute import ComputeTarget
import os
# choose a name for your cluster
compute_name = os.environ.get("AML_COMPUTE_CLUSTER_NAME", "cpu-cluster")
compute_min_nodes = os.environ.get("AML_COMPUTE_CLUSTER_MIN_NODES", 0)
compute_max_nodes = os.environ.get("AML_COMPUTE_CLUSTER_MAX_NODES", 4)
# This example uses CPU VM. For using GPU VM, set SKU to STANDARD_NC6
vm_size = os.environ.get("AML_COMPUTE_CLUSTER_SKU", "STANDARD_D3_V2")
if compute_name in ws.compute_targets:
compute_target = ws.compute_targets[compute_name]
if compute_target and type(compute_target) is AmlCompute:
print("found compute target: " + compute_name)
else:
print("creating new compute target...")
provisioning_config = AmlCompute.provisioning_configuration(vm_size = vm_size,
min_nodes = compute_min_nodes,
max_nodes = compute_max_nodes)
# create the cluster
compute_target = ComputeTarget.create(ws, compute_name, provisioning_config)
# can poll for a minimum number of nodes and for a specific timeout.
# if no min node count is provided it will use the scale settings for the cluster
compute_target.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)
# For a more detailed view of current AmlCompute status, use get_status()
print(compute_target.get_status().serialize())
# -
# You now have the necessary packages and compute resources to train a model in the cloud.
#
# ## Explore data
# Before you train a model, you need to understand the data that you are using to train it. In this section you learn how to:
#
# - Download the captured raw sound is CSV files dataset
# - Split each csv file into chunks when captured and convert it to a format that is suitable for CNN training
# - Create label dataset using user specified csv file. the csv file should be provided by user apart from sound csv files.
# - Display some sounds
#
# The format of sound scv file name is...
# <i>deviceid</i>-sound-<i>yyyyMMddHHmmssffffff</i>.csv
# The part of <i>yyyyMMddHHmmss</i> means that the start time of sound capturing so that user can specify the appropriate files by specifying the start and end times.
# So the csv file for label dataset specification consist from following format record.
# <i>label_name</i>,<i>start timestamp</i>,<i>end timestamp</i>
#
# both timestamp format shold be following.
# <i>yyyy</i>/<i>MM</i>/<i>dd</i> <i>HH</i>:<i>mm</i>:<i>ss</i>
#
# In the following code label dataset definition csv files name are
# - for training 'train-label-range.csv'
# - for testing 'test-label-range.csv'
#
# The following cell should be run only when dataset is ready or updated.
#
# ※ You can use sound dataset stored in [sample guitar raw sound dataset](https://egstorageiotkitvol5.blob.core.windows.net/sound-ml-data/raw-sound-data.zip). please use a dataset that uploads a set of files that be extracted from the zip file to the blob which you create by your own Azure account.
# +
# l-3
import os
train_label_range_csv_file = 'train-label-range.csv'
test_label_range_csv_file = 'test-label-range.csv'
csv_files = [train_label_range_csv_file, test_label_range_csv_file]
parsed_specs =[]
for csv in csv_files:
specs ={}
parsed_specs.append(specs)
with open(csv,"rt") as f:
ranges = f.readlines()
for r in ranges:
spec = r.rstrip().split(',')
if not (spec[0] in specs):
specs[spec[0]] = []
specs[spec[0]].append([spec[1],spec[2]])
duration_for_train = parsed_specs[0]
duration_for_test = parsed_specs[1]
for i, ps in enumerate(parsed_specs):
print('spec for - ' + csv_files[i])
for d in duration_for_train.keys():
for s in duration_for_train[d]:
print(' {0}:{1}-{2}'.format(d,s[0],s[1]))
# -
# ## Download guitar sound dataset from your own blob container
# This code download only csv files that meet label specificated timespan criteria. Before you run following code, please set source_azure_storage_account_connection_string and source_container_name to match your storage account that contains the sound data files. The source_azure_storage_account_connection_string is configured into data-storage-config.yml
#
# The files satisfied criteria will be downloaded in data and test folder.
# +
# l-4
# #!pip install -U azure-storage-blob>=12.2.0
# #!pip list
import os
import datetime
import yaml
yml = {}
with open(dataSrorageConfigFile,'r') as ymlfile:
yml.update(yaml.safe_load(ymlfile))
#print('config - {}'.format(yml))
# Specify datasource
source_azure_storage_account_connection_string = yml['blob_connection_string']
#source_azure_storage_account_connection_string = '< - your azure storage connection string - >'
source_container_name = 'edgesounds'
# Specify start and end time of duration for each chord
# duration_for_train = {
# 'major':[['2020/02/09 16:58:34', '2020/02/09 16:58:40'], ['2020/02/09 17:06:02','2020/02/09 17:06:10'], ['2020/02/09 16:59:42','2020/02/09 16:59:50'], ['2020/02/09 17:00:41','2020/02/09 17:00:49'],['2020/02/18 11:27:20', '2020/02/18 11:27:26'],['2020/02/18 11:28:05', '2020/02/18 11:28:11'],['2020/02/18 11:28:41', '2020/02/18 11:28:44'],['2020/02/18 11:29:18', '2020/02/18 11:29:20'],['2020/02/18 11:29:51', '2020/02/18 11:29:57'],['2020/02/18 11:30:25', '2020/02/18 11:30:29'],['2020/02/18 11:31:05', '2020/02/18 11:31:12']
# ],
# 'minor':[['2020/02/09 16:58:49', '2020/02/09 16:59:00'], ['2020/02/09 17:06:26','2020/02/09 17:06:36'], ['2020/02/09 16:59:57','2020/02/09 17:00:05'], ['2020/02/09 17:00:56','2020/02/09 17:01:03'],['2020/02/18 11:27:41', '2020/02/18 11:27:47'],['2020/02/18 11:28:22', '2020/02/18 11:28:26'],['2020/02/18 11:28:59', '2020/02/18 11:29:03'],['2020/02/18 11:29:33', '2020/02/18 11:29:38'],['2020/02/18 11:30:07', '2020/02/18 11:30:13'],['2020/02/18 11:30:45', '2020/02/18 11:30:49'],['2020/02/18 11:31:24', '2020/02/18 11:31:35']
# ]
# }
# duration_for_test = {
# 'major':[['2020/02/18 11:36:37', '2020/02/18 11:36:44'],['2020/02/18 11:37:12', '2020/02/18 11:37:18'],['2020/02/18 11:37:43', '2020/02/18 11:37:49'],['2020/02/18 11:38:18', '2020/02/18 11:38:23'],['2020/02/18 11:38:58', '2020/02/18 11:39:06'],['2020/02/18 11:39:36', '2020/02/18 11:39:40'],['2020/02/18 11:40:14', '2020/02/18 11:40:20']],
# 'minor':[['2020/02/18 11:36:56', '2020/02/18 11:37:01'],['2020/02/18 11:37:25', '2020/02/18 11:37:33'],['2020/02/18 11:38:00', '2020/02/18 11:38:08'],['2020/02/18 11:38:35', '2020/02/18 11:38:42'],['2020/02/18 11:39:15', '2020/02/18 11:39:21'],['2020/02/18 11:39:53', '2020/02/18 11:39:59'],['2020/02/18 11:40:34', '2020/02/18 11:40:41']]
# }
def pickup_target_files(target_folder_name, duration_for_target):
folder_for_label = {}
condition_for_label = {}
target_data = {}
# data store for traning
data_folder = os.path.join(os.getcwd(), target_folder_name)
for dflk in duration_for_target.keys():
folder_for_key = os.path.join(data_folder, dflk)
folder_for_label[dflk] = folder_for_key
os.makedirs(folder_for_key, exist_ok=True)
condition_for_label[dflk] = []
durs = duration_for_target[dflk]
for dur in durs:
dur_se = []
while len(dur)>0:
t = dur.pop(0)
ttime = datetime.datetime.strptime(t, '%Y/%m/%d %H:%M:%S')
tnum = ttime.strftime('%Y%m%d%H%M%S') + '000000'
dur_se.append(int(tnum))
condition_for_label[dflk].append(dur_se)
target_data[dflk] = []
return folder_for_label, condition_for_label, target_data
#data_folder_name = 'data-csv'
#test_folder_name = 'test-csv'
train_folder_for_label, train_condition_for_label, train_data = pickup_target_files(data_folder_path, duration_for_train)
test_folder_for_label, test_condition_for_label, test_data = pickup_target_files(test_folder_path, duration_for_test)
from azure.storage.blob import BlobServiceClient
import datetime
import re
import numpy as np
# Connect to our blob via the BlobService
blobServiceClient = BlobServiceClient.from_connection_string(source_azure_storage_account_connection_string)
containerClient = blobServiceClient.get_container_client(source_container_name)
soundDataDefFile = 'sounddata-csv.yml'
def load_targeted_blobs(container, condition_for_target, folder_for_target, data_for_target ):
with open(soundDataDefFile, "wb") as ymlFile:
blobClient = containerClient.get_blob_client(soundDataDefFile)
stream = blobClient.download_blob()
blobContent = stream.readall()
ymlFile.write(blobContent)
target_blobs = []
loaded_num_of_files = {}
for blob in containerClient.list_blobs():
matching = re.findall('sound-[0-9]+.csv', blob.name)
if len(matching)>0:
target_blobs.append({'blob':blob, 'num-of-ts':int(re.findall('[0-9]+',blob.name)[0])})
for l in condition_for_target.keys():
for cfl in condition_for_target[l]:
filtered = list(filter(lambda b: cfl[0] <= b['num-of-ts'] and b['num-of-ts'] <= cfl[1], target_blobs))
data_for_target[l].append(filtered)
for l in data_for_target.keys():
num_of_files = 0
print('Label - '+l)
for dft in data_for_target[l]:
for ltd in dft:
blobClient = containerClient.get_blob_client(ltd['blob'])
stream = blobClient.download_blob()
csvFilePath = os.path.join(folder_for_target[l], ltd['blob'].name)
print(' Downloading - ' + ltd['blob'].name)
with open(csvFilePath,"wb") as csvFile:
blobContent = stream.readall()
csvFile.write(blobContent)
num_of_files = num_of_files + 1
loaded_num_of_files[l] = num_of_files
return loaded_num_of_files
result = load_targeted_blobs(containerClient, train_condition_for_label, train_folder_for_label, train_data)
for k in result.keys():
print('Loaded file for train:{0} - {1}'.format(k, result[k]))
result = load_targeted_blobs(containerClient, test_condition_for_label, test_folder_for_label, test_data)
for k in result.keys():
print('Loaded file for test:{0} - {1}'.format(k, result[k]))
import shutil
for fldr in [data_folder_path, test_folder_path]:
destFName = os.path.join(fldr,soundDataDefFile)
shutil.copy(soundDataDefFile, destFName)
# -
# ### Create Training and Test data
# reform data for training and test
# +
# %%writefile loadsounds.py
# l-5
import os
import csv
import numpy as np
import random
import yaml
def load_csvdata(file):
with open(file) as f:
reader = csv.reader(f)
l = [row for row in reader]
l.pop(0)
return np.array(l).T.astype(np.int16) / 32768
# data_folder = 'data'
# data_folder_path = os.path.join(os.getcwd(), data_folder)
def parse_file(file_path, labeled_dataset, data_chunk, by_channel=False):
csvdata = load_csvdata(file_path)
if by_channel:
if len(csvdata) > 0:
num_of_block = int(len(csvdata[0]) / data_chunk)
if num_of_block > 0 :
labeled_dataset = np.ndarray(shape=(len(csvdata),num_of_block,data_chunk))
num_of_chunk = 0
for index, umicdata in enumerate(csvdata):
if len(umicdata) % data_chunk == 0:
micdata1024 = np.split(umicdata, len(umicdata) / data_chunk)
if by_channel:
labeled_dataset[index] = micdata1024
else:
if len(labeled_dataset) == 0:
labeled_dataset = micdata1024
else:
labeled_dataset = np.append(labeled_dataset, micdata1024,axis=0)
num_of_chunk = num_of_chunk + len(micdata1024)
# print('{}:{} units'.format(file_path, num_of_chunk))
return num_of_chunk, labeled_dataset
def load_data_definition(data_def_file_path):
definition = {}
with open(data_def_file_path, "r") as ymlFile:
yml = yaml.safe_load(ymlFile)
definition.update(yml)
return definition
def reshape_dataset(sound_data, data_chunk):
dataset = np.zeros((len(sound_data), 1, 1, data_chunk))
for index, d in enumerate(sound_data):
dataset[index][0][0] = d
return dataset
def load_data(data_folder_path, data_def_file):
data_chunk = 1024
tdata ={}
num_of_data = 0
for df in os.listdir(path=data_folder_path):
if df == data_def_file:
ddefFile = os.path.join(data_folder_path, df)
data_chunk = load_data_definition(ddefFile)['data-chunk']
print('data_chunk - {}'.format(data_chunk))
continue
tdata[df] = np.array([])
ldata_folder_path = os.path.join(data_folder_path,df)
for datafile in os.listdir(path=ldata_folder_path):
datafile_path = os.path.join(ldata_folder_path, datafile)
num_of_chunks, tdata[df] = parse_file(datafile_path, tdata[df], data_chunk)
num_of_data = num_of_data + num_of_chunks
data_of_sounds = np.zeros((num_of_data, 1, 1, data_chunk))
label_of_sounds = np.zeros(num_of_data,dtype=int)
label_matrix_of_sounds = np.zeros((num_of_data, len(tdata.keys())))
labelname_of_sounds = np.empty(num_of_data,dtype=object)
index = 0
lindex = 0
labeling_for_train = {}
for l in tdata.keys():
for micdata1024 in tdata[l]:
data_of_sounds[index][0][0] = micdata1024
label_of_sounds[index] = lindex
label_matrix_of_sounds[index, lindex] = 1.
labelname_of_sounds[index] = l
index = index + 1
labeling_for_train[l] = lindex
lindex = lindex + 1
indexx = np.arange(num_of_data)
random.shuffle(indexx)
data_of_sounds = data_of_sounds[indexx]
label_matrix_of_sounds = label_matrix_of_sounds[indexx]
label_of_sounds = label_of_sounds[indexx]
labelname_of_sounds = labelname_of_sounds[indexx]
# train_dataset is labeled sound data set
train_dataset = [data_of_sounds, label_of_sounds, labelname_of_sounds]
return train_dataset
# -
# ### Check sound data for training
# You can confirm that content of sound data by following code.
# +
# l-6
from loadsounds import load_data
# data_chunk shoud be same value for sound capturing chunk
data_def_file = 'sounddata-csv.yml'
print('loading train data...')
train_dataset = load_data(data_folder_path, data_def_file)
print('loading test data...')
test_dataset = load_data(test_folder_path, data_def_file)
print(train_dataset)
import matplotlib.pyplot as plt
sample_size = 4
figure = plt.figure(figsize=(5,4))
fig, grps = plt.subplots(ncols=sample_size)
for d in range(0,sample_size):
grps[d].plot(train_dataset[0][d][0][0])
grps[d].set_title(train_dataset[2][d])
plt.show()
# -
# ## Create Data store for training on remote computer
# This logic execution is necessary for only when data is updated.
#
# +
from azureml.core import Workspace, Datastore, Dataset
# retrieve current datastore in the workspace
datastore = ws.get_default_datastore()
# Upload files to dataset on datastore
datastore.upload(src_dir=data_folder_path,
target_path= dataset_name,
overwrite=True,
show_progress=True)
datastore.upload(src_dir=test_folder_path,
target_path= testset_name,
overwrite=True,
show_progress=True)
# create a FileDataset pointing to files in 'animals' folder and its subfolders recursively
datastore_paths = [(datastore, dataset_name)]
sound_ds = Dataset.File.from_files(path=datastore_paths)
teststore_paths = [(datastore, testset_name)]
sound_ts = Dataset.File.from_files(path=teststore_paths)
print(sound_ds)
# Register dataset to current workspace
sound_dataset = sound_ds.register(workspace=ws,
name=dataset_name,
description='sound classification training data')
sound_testset = sound_ts.register(workspace=ws,
name=testset_name,
description='sound classification test data')
# -
# ### Construct CNN model
#
# Following cell show a sample of CNN model for sound classification and training.
# The cell logic will be run on this computing environment.
#
# +
# l-7
# https://www.tensorflow.org/tutorials/images/intro_to_cnns?hl=ja
import tensorflow as tf
from tensorflow.keras import datasets, layers, models
print('tensorflow version - '+tf.__version__)
model = models.Sequential()
model.add(layers.Conv2D(16,input_shape=(1,1,1024),kernel_size=(1,8),padding='same', strides=(1,4), activation='relu'))
model.add(layers.MaxPooling2D(pool_size=(1,2), padding='same'))
model.add(layers.Conv2D(filters=16,input_shape=(1,1,128),kernel_size=(1,8),padding='same', strides=(1,4), activation='relu'))
model.add(layers.MaxPooling2D(pool_size=(1,2), padding='same'))
model.add(layers.Conv2D(filters=16,input_shape=(1,1,16),kernel_size=(1,8),padding='same', strides=(1,4), activation='relu'))
model.add(layers.MaxPooling2D(pool_size=(1,2), padding='same'))
model.add(layers.Flatten())
model.add(layers.Dense(10, activation='relu'))
model.add(layers.Dense(2, activation='sigmoid'))
# Above code part is same as training logic on remote computer
model.summary()
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
hist = model.fit(train_dataset[0], train_dataset[1], epochs=5, validation_data=(test_dataset[0], test_dataset[1]))
test_loss, test_acc = model.evaluate(test_dataset[0],test_dataset[1])
print('Test accuracy - '+str(test_acc))
for hk in hist.history.keys():
print(hk)
import matplotlib.pyplot as plt
epoch_list = list(range(1, len(hist.history['accuracy']) + 1))
plt.plot(epoch_list, hist.history['accuracy'], epoch_list, hist.history['val_accuracy'])
plt.legend(('Training Accuracy', "Validation Accuracy"))
plt.show()
predictions = model.predict(test_dataset[0])
# Plot a random sample of 10 test images, their predicted labels and ground truth
figure = plt.figure(figsize=(20, 8))
for i, index in enumerate(np.random.choice(test_dataset[1], size=15, replace=False)):
ax = figure.add_subplot(3, 5, i + 1, xticks=[], yticks=[])
# Display each image
ax.plot(test_dataset[0][index][0][0])
predict_index = np.argmax(predictions[index])
true_index = np.argmax(test_dataset[1][index])
# print('{}-{}'.format(predict_index, true_index))
# Set the title for each image
ax.set_title("{} ({})".format(test_dataset[2][predict_index],
test_dataset[2][true_index]),
color=("green" if predict_index == true_index else "red"))
# When you need h5 format file, change .pkl -> .h5
model_path = 'sound-classification-csv-model'
#model_path_ext = '.h5'
model_path_ext = '.pkl'
output_dir = 'outputs'
os.makedirs(output_dir, exist_ok=True)
model_pathname = os.path.join(output_dir, model_path + model_path_ext)
model.save(model_pathname)
#model.save('outputs/sound-classification-model.h5')
# -
# ### Export learned model
#
# exported model will be used in IoT Edge SoundClassifierService module
# +
import os
import datetime
import tarfile
def compress_files(top, archive, dest_folder):
tarfilename = archive + '.tar.gz'
topbase = os.path.basename(top)
if tarfilename is None:
now = datetime.datetime.now()
tarfilename = '{0}-{1:%Y%m%d%H%M%S}.tar.gz'.format(topbase, now)
if dest_folder is not None:
tarfilename = os.path.join(dest_folder, tarfilename)
os.makedirs(dest_folder,exist_ok=True)
tar = tarfile.open(tarfilename, "w:gz")
for root, dirs, files in os.walk(top):
for filename in files:
parent = root[len(top):]
if parent.startswith('\\'):
parent = parent[1:]
archnameraw = os.path.join(parent,filename)
archname = os.path.join(topbase, archnameraw).replace('\\','/',-1)
tar.add(os.path.join(root, filename),archname)
tar.close()
return tarfilename
export_folder_path = 'export'
os.makedirs(export_folder_path,exist_ok=True)
if model_path_ext == '.pkl':
compressed = compress_files(model_pathname, model_path + model_path_ext, export_folder_path)
print('Learned model is exproted as ' + compressed)
# -
# ### Convert and create tensorflow lite model for micro edge device.
# Please refer https://www.tensorflow.org/lite/guide/inference and [/SoundAIonMicroEdge/README.md](../../SoundAIonMicroEdge/README.md) to use the converted file.
# +
import tensorflow as tf
import os
model_file = 'sound-classification-csv-model.pkl' # this value should be same name of above logic
output_dir = 'outputs'
model_pathname = os.path.join(output_dir, model_file)
converter = tf.lite.TFLiteConverter.from_saved_model(model_pathname)
converter.optimizations = [tf.lite.Optimize.DEFAULT]
quantized_model = converter.convert()
tflite_model_file = 'sound-classification-csv-model.tflite'
export_folder_path = 'export'
os.makedirs(export_folder_path,exist_ok=True)
tflite_model_pathname = os.path.join(export_folder_path, tflite_model_file)
open(tflite_model_pathname, "wb").write(quantized_model)
# -
# ### Create Training script
import os
script_folder = os.path.join(os.getcwd(), "sklearn-script")
os.makedirs(script_folder, exist_ok=True)
# ## CNN Model traing script
# The following cell is the script of model definition, model training for running on remote computing cluster.
# +
# %%writefile $script_folder/train.py
import argparse
import os
import numpy as np
import glob
#from sklearn.externals import joblib
from azureml.core import Run
from loadsounds import load_data
import tensorflow as tf
from tensorflow.keras import datasets, layers, models
parser = argparse.ArgumentParser()
parser.add_argument('--data-folder', type=str, dest='data_folder', help='data folder mounting point')
parser.add_argument('--test-folder', type=str, dest='test_folder', help='test folder mounting point')
#parser.add_argument('--regularization', type=float, dest='reg', default=0.01, help='regularization rate')
args = parser.parse_args()
data_folder = args.data_folder
test_folder = args.test_folder
print('Data folder:{0}, Test folder:{1}'.format( data_folder, test_folder))
data_def_file = 'sounddata-csv.yml'
train_dataset = load_data(data_folder,data_def_file)
test_dataset = load_data(test_folder, data_def_file)
# get hold of the current run
run = Run.get_context()
print('tensorflow version - '+tf.__version__)
model = models.Sequential()
model.add(layers.Conv2D(16,input_shape=(1,1,1024),kernel_size=(1,8),padding='same', strides=(1,4), activation='relu'))
model.add(layers.MaxPooling2D(pool_size=(1,2), padding='same'))
model.add(layers.Conv2D(filters=16,input_shape=(1,1,128),kernel_size=(1,8),padding='same', strides=(1,4), activation='relu'))
model.add(layers.MaxPooling2D(pool_size=(1,2), padding='same'))
model.add(layers.Conv2D(filters=16,input_shape=(1,1,16),kernel_size=(1,8),padding='same', strides=(1,4), activation='relu'))
model.add(layers.MaxPooling2D(pool_size=(1,2), padding='same'))
model.add(layers.Flatten())
model.add(layers.Dense(10, activation='relu'))
model.add(layers.Dense(2, activation='sigmoid'))
# Above code part is same as training logic on remote computer
model.summary()
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
hist = model.fit(train_dataset[0], train_dataset[1], epochs=5, validation_data=(test_dataset[0], test_dataset[1]))
test_loss, test_acc = model.evaluate(test_dataset[0],test_dataset[1])
print('Test accuracy - '+str(test_acc))
for hk in hist.history.keys():
print(hk)
epoch_list = list(range(1, len(hist.history['accuracy']) + 1))
#import matplotlib.pyplot as plt
# plt.plot(epoch_list, hist.history['accuracy'], epoch_list, hist.history['val_accuracy'])
# plt.legend(('Training Accuracy', "Validation Accuracy"))
# plt.show()
# predictions = model.predict(test_dataset[0])
# Plot a random sample of 10 test images, their predicted labels and ground truth
# figure = plt.figure(figsize=(20, 8))
# for i, index in enumerate(np.random.choice(test_dataset[1], size=15, replace=False)):
# ax = figure.add_subplot(3, 5, i + 1, xticks=[], yticks=[])
# # Display each image
# ax.plot(test_dataset[0][index][0][0])
# predict_index = np.argmax(predictions[index])
# true_index = np.argmax(test_dataset[1][index])
# print('{}-{}'.format(predict_index, true_index))
# Set the title for each image
# ax.set_title("{} ({})".format(test_dataset[2][predict_index],
# test_dataset[2][true_index]),
# color=("green" if predict_index == true_index else "red"))
# Above code result can't be shown in Azure ML Studio
model_path = 'sound-classification-csv-model'
model_path_ext = '.pkl'
output_dir = 'outputs'
os.makedirs(output_dir, exist_ok=True)
model_pathname = os.path.join(output_dir, model_path + model_path_ext)
model.save(model_pathname)
# -
import shutil
shutil.copy('loadsounds.py', script_folder)
# ### Training and Learning!
# train CNN model with sound dataset
# +
from azureml.core.environment import Environment
from azureml.core.conda_dependencies import CondaDependencies
# to install required packages
env = Environment('my_env')
cd = CondaDependencies.create(pip_packages=['azureml-sdk','scikit-learn','azureml-dataprep[pandas,fuse]>=1.1.14','tensorflow==2.1.0','matplotlib','pyyaml'])
env.python.conda_dependencies = cd
# +
from azureml.train.sklearn import SKLearn
from azureml.core import Dataset, Run
# Get a dataset by name
sound_dataset = Dataset.get_by_name(workspace=ws, name=dataset_name)
data_mount = sound_dataset.as_named_input('sound_data').as_mount()
test_dataset = Dataset.get_by_name(workspace=ws, name=testset_name)
test_mount = sound_dataset.as_named_input('sound_test').as_mount()
script_params = {
# to mount files referenced by mnist dataset
'--data-folder': data_mount,
'--test-folder': test_mount
}
est = SKLearn(source_directory=script_folder,
script_params=script_params,
compute_target=compute_target,
environment_definition=env,
entry_script='train.py')
# -
# ### For debug
# Following three blocks are used to check specified parameters and dataset validity.
# When you don't need to debug, please go forward to ["Submit the job to the cluster"](#Submit-the-job-to-the-cluster)
# +
# %%writefile $script_folder/testargs.py
import argparse
parser = argparse.ArgumentParser()
parser.add_argument('--data-folder', type=str, dest='data_folder', help='data folder mounting point')
parser.add_argument('--test-folder', type=str, dest='test_folder', help='test folder mounting point')
#parser.add_argument('--regularization', type=float, dest='reg', default=0.01, help='regularization rate')
args = parser.parse_args()
data_folder = args.data_folder
test_folder = args.test_folder
print('Data folder:{0}', 'Test folder:{1}'.format( data_folder, test_folder))
import os
chdir = os.getcwd()
print('Current Dir - '+chdir)
folders = {'data':data_folder,'test':test_folder}
for fld in folders.keys():
cfld = folders[fld]
print('Check content of {0} - {1}'.format(fld, cfld))
for f in os.listdir(cfld):
print(' '+f)
cdir = os.path.join(data_folder, f)
if os.path.isdir(cdir):
for cf in os.listdir(cdir):
print(' '+cf)
# +
from azureml.core.environment import Environment
from azureml.core.conda_dependencies import CondaDependencies
# to install required packages
env = Environment('my_env')
cd = CondaDependencies.create(pip_packages=['azureml-sdk','matplotlib','pyyaml','azureml-dataprep[pandas,fuse]>=1.1.14'])
env.python.conda_dependencies = cd
# +
from azureml.train.sklearn import SKLearn
from azureml.core import Dataset, Run
# Get a dataset by name
sound_dataset = Dataset.get_by_name(workspace=ws, name=dataset_name)
data_mount = sound_dataset.as_named_input('sound_data').as_mount()
test_dataset = Dataset.get_by_name(workspace=ws, name=testset_name)
test_mount = sound_dataset.as_named_input('sound_test').as_mount()
print(data_mount)
print(test_mount)
script_params = {
# to mount files referenced by mnist dataset
'--data-folder': data_mount,
'--test-folder': test_mount
}
est = SKLearn(source_directory=script_folder,
script_params=script_params,
compute_target=compute_target,
environment_definition=env,
entry_script='testargs.py')
# -
# ### Local Debug
# +
import numpy as np
from loadsounds import parse_file, load_data_definition, reshape_dataset
import os
import random
#data_def_file = 'sounddata.yml'
#datafile = 'cherry-sound-20200218113643319806.csv'
data_chunk = load_data_definition(data_def_file)['data-chunk']
#csv_dataset = parse_file(datafile,np.array([]),data_chunk)
#sound_dataset = np.zeros((csv_dataset[0], 1, 1, data_chunk))
#index = 0
#for d in csv_dataset[1]:
# sound_dataset[index][0][0] = d
# index = index + 1
test_data_files = []
for d in os.listdir(test_folder_path):
dname = os.path.join(test_folder_path, d)
if os.path.isdir(dname):
for f in os.listdir(dname):
if f.rfind('.csv') >= 0:
test_data_files.append(os.path.join(dname,f))
random.shuffle(test_data_files)
csv_dataset = parse_file(test_data_files[0], np.array([]), data_chunk, by_channel=True)
import tensorflow as tf
from tensorflow.keras import datasets, layers, models
print('tensorflow version - '+tf.__version__)
#model_file_path ='outputs/sound-classification-model.h5'
model_file_path ='outputs/sound-classification-model.pkl'
# model name should be used other style
model = models.load_model(model_file_path)
for channel, csvds in enumerate(csv_dataset[1]):
sound_dataset = reshape_dataset(csv_dataset[1][channel], data_chunk['data-chunk'])
predicted = model.predict(sound_dataset)
print('channel:{}'.format(channel))
result = predicted.tolist()
for r in result:
print('{0}<->{1}'.format(r[0],r[1]))
# -
# ### Submit the job to the cluster
# Run the experiment by submitting the estimator object. And you can navigate to Azure portal to monitor the run.
run = exp.submit(config=est)
run
# ### Since the call is asynchronous, it returns a **Preparing** or **Running** state as soon as the job is started.
#
# ## Monitor a remote run
#
# In total, the first run takes **approximately 10 minutes**. But for subsequent runs, as long as the dependencies (`conda_packages` parameter in the above estimator constructor) don't change, the same image is reused and hence the container start up time is much faster.
#
# Here is what's happening while you wait:
#
# - **Image creation**: A Docker image is created matching the Python environment specified by the estimator. The image is built and stored in the ACR (Azure Container Registry) associated with your workspace. Image creation and uploading takes **about 5 minutes**.
#
# This stage happens once for each Python environment since the container is cached for subsequent runs. During image creation, logs are streamed to the run history. You can monitor the image creation progress using these logs.
#
# - **Scaling**: If the remote cluster requires more nodes to execute the run than currently available, additional nodes are added automatically. Scaling typically takes **about 5 minutes.**
#
# - **Running**: In this stage, the necessary scripts and files are sent to the compute target, then data stores are mounted/copied, then the entry_script is run. While the job is running, stdout and the files in the ./logs directory are streamed to the run history. You can monitor the run's progress using these logs.
#
# - **Post-Processing**: The ./outputs directory of the run is copied over to the run history in your workspace so you can access these results.
#
#
# You can check the progress of a running job in multiple ways. This tutorial uses a Jupyter widget as well as a `wait_for_completion` method.
#
# ### Jupyter widget
#
# Watch the progress of the run with a Jupyter widget. Like the run submission, the widget is asynchronous and provides live updates every 10-15 seconds until the job completes.
from azureml.widgets import RunDetails
RunDetails(run).show()
run.wait_for_completion(show_output=True) # specify True for a verbose log
# ### Evaluate the model output
print(run.get_metrics())
print(run.get_file_names())
# ## Are you happy with the model??? Register it in Azure Machine Learning to manage
# register model
model = run.register_model(model_name='sound_clasification_model', model_path='outputs/')
print(model.name, model.id, model.version, sep = '\t')
# ## Next step
# In this Azure Machine Learning tutorial, you used Python to:
#
# > * Set up your development environment
# > * Access and examine the data
# > * Train multiple models on a remote cluster using the tensorflow keras machine learning library
# > * Review training details and register the best model
#
# You are ready to deploy this registered model using the instructions in the next part of the tutorial series:
#
# > [Tutorial 2 - Deploy models](ai-sound-major-miner-classification-part2-deploy.ipynb)
| SoundAI/notebook/ai-sound-major-miner-classification-part1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Supporting workings - Navigating census data
#
# As support for the main workbook, I first need an effective way of selecting which categories to select in the first instance from the extremely large datasets. From the raw metadata file for the 2016 datapack, there was 15536 different data points for each statistical area across all 59 tables. Some of these were totals of other lines, some were the total population, some had combinations of classifications available elsewhere in the overall population, etc. so a mechanism of refinement is clearly necessary.
#
# Ideally I would like to be able to import data based on some simple aggregations where possible (e.g. by sex), or otherwise taking the "highest" level data possible given the input constraints. E.g. I would like to say "Import data from tables G09, G52 and G53, but split by sex where possible", if "sex" is not available in one of the tables, it will still return the data from the table, but from the most aggregated column(s) only. Additionally, if "Sex" is only available as a sub-category of another split (e.g. the measure is "Sex by Age") I would like to be able to perform my own aggregation, to abstract away the unneccessary other details imbedded in the data (e.g. sum all the "Age" categories together, grouping by sex).
# +
# Import statements
# Declare Imports
import numpy as np
import pandas as pd
import os
import re
import operator
import csv
# Set a variable for current notebook's path for various loading/saving mechanisms
nb_path = os.getcwd()
# -
# Generalise this importation method to allow easy imports based on folder name (for SA level)
# and a list of datapack files you want to amalgamate into a single dataframe
def load_census_csv(table_list, statistical_area_code):
statistical_area_code = statistical_area_code.upper()
for index, table in enumerate(table_list):
if index==0:
df = pd.read_csv('{}\Data\{}\AUST\\2016Census_{}_AUS_{}.csv'.format(nb_path,
statistical_area_code,
table,
statistical_area_code
),
engine='python')
else:
temp_df = pd.read_csv('{}\Data\{}\AUST\\2016Census_{}_AUS_{}.csv'.format(nb_path,
statistical_area_code,
table,
statistical_area_code
),
engine='python')
merge_col = df.columns[0]
df = pd.merge(df, temp_df, on=merge_col)
return df
# ### Begin importing and exploring data
# Import metadata sheets
df_meta_tables = pd.read_excel('{}\Data\Metadata\Metadata_2016_GCP_DataPack.xlsx'.format(nb_path),
sheet_name = 'Table number, name, population',
skiprows=9)
df_meta_measures = pd.read_excel('{}\Data\Metadata\Metadata_2016_GCP_DataPack.xlsx'.format(nb_path),
sheet_name = 'Cell descriptors information',
skiprows=10)
df_meta_tables.tail()
df_meta_measures['Table number'] = df_meta_measures['DataPack file'].str[:3]
df_meta_measures = pd.merge(df_meta_measures, df_meta_tables, on='Table number')
df_meta_measures.shape
df_meta_measures.tail(20)
# #### Quick notes:
# I noticed that very conveniently a lot of the capitalisation in the "Long" field works as a delimiter between different population characteristics. Working through some exceptions below, split the Long fields based on capitalisation (and a couple of other hueristics) to get down to a much shorter list of 911 characteristics. I turned this information into a csv file which I then did some tedious review on in order to come up with a much shorter list of 61 categories and created a new "Meta" file which I import after this section.
def lower_all_except_first_char(str_item):
str_replace = '{}{}'.format(str_item[:1],str_item[1:].lower())
df_meta_measures['Long'] = df_meta_measures['Long'].str.replace(str_item, str_replace)
# +
replace_capitalisation_list = ['Not_stated',
'None',
'No_children',
'Aboriginal_and_or_Torres_Strait_Islander',
'Both_Aboriginal_and_Torres_Strait_Islander',
'Torres_Strait_Islander',
'Non_Indigenous',
'New_Zealand',
'South_Africa',
'Sri_Lanka',
'Bosnia_and_Herzegovina',
'China_excludes_SARs_and_Taiwan',
'Hong_Kong_SAR_of_China',
'Korea_Republic_of_South',
'Northern_Ireland',
'Papua_New_Guinea',
'South_Eastern_Europe',
'Total_Responses',
'Census_Night',
'Elsewhere_in_Australia',
'Birthplace_Australia',
'Birthplace_Elsewhere',
'Other_Language',
'Age_of_Persons',
'Count_of_Persons',
'Average_number_of_Persons_per_bedroom',
'Visitor_from_Different_SA2',
'Visitor_from_Same_Statistical_Area_Level_2_SA2',
'Western_Australia',
'Northern_Territory',
'Australian_Capital_Territory',
'South_Australia',
'New_South_Wales',
'Other_Territories',
'China_excl_SARs_and_Taiwan',
'United_Kingdom_Channel_Islands_and_Isle_of_Man',
'The_Former_Yugoslav_Republic_of_Macedonia',
'United_States_of_America',
'Year_of_arrival_Before',
'Speaks_English',
'speaks_English',
'Proficiency_in_English',
'Proficiency_in_english',
'_Before_2000',
'Total_Year_of_arrival_not_stated',
'Australian_Indigenous_Languages',
'Chinese_Languages_Cantonese',
'Chinese_Languages_Mandarin',
'Chinese_Languages_Other',
'Chinese_languages_Other',
'Chinese_Languages_Total',
'Indo_Aryan_Languages_Bengali',
'Indo_Aryan_Languages_Hindi',
'Indo_Aryan_Languages_Punjabi',
'Indo_Aryan_Languages_Sinhalese',
'Indo_Aryan_Languages_Urdu',
'Indo_Aryan_Languages_Other',
'Indo_Aryan_Languages_Total',
'Persian_excluding_Dari',
'Southeast_Asian_Austronesian_Languages_Filipino',
'Southeast_Asian_Austronesian_Languages_Indonesian',
'Southeast_Asian_Austronesian_Languages_Tagalog',
'Southeast_Asian_Austronesian_Languages_Other',
'Southeast_Asian_Austronesian_Languages_Total',
'Christianity_Anglican',
'Christianity_Assyrian_Apostolic',
'Christianity_Baptist',
'Christianity_Brethren',
'Christianity_Catholic',
'Christianity_Churches_of_Christ',
'Christianity_Eastern_Orthodox',
'Christianity_Jehovahs_Witnesses',
'Christianity_Latter_day_Saints',
'Christianity_Lutheran',
'Christianity_Oriental_Orthodox',
'Christianity_Other_Protestant',
'Christianity_Pentecostal',
'Christianity_Presbyterian_and_Reformed',
'Christianity_Salvation_Army',
'Christianity_Seventh_day_Adventist',
'Christianity_Uniting_Church',
'Christianity_Christianity_nfd',
'Christianity_Other_Christian',
'Christianity_Total',
'Other_Religions_Australian_Aboriginal_Traditional_Religions',
'Other_Religions_Sikhism',
'Other_Religions_Other',
'Other_Religions_Total',
'Secular_Beliefs_and_Other_Spiritual_Beliefs_and_No_Religious_Affiliation_No_Religion_So_Described',
'Secular_Beliefs_and_Other_Spiritual_Beliefs_and_No_Religious_Affiliation_Secular_Beliefs',
'Secular_Beliefs_and_Other_Spiritual_Beliefs_and_No_Religious_Affiliation_Other_Spiritual_Beliefs',
'Secular_Beliefs_and_Other_Spiritual_Beliefs_and_No_Religious_Affiliation_Total',
'Infants_Primary',
'Other_Non_Government',
'Technical_or_Further_Educational_institution',
'Full_Part_time',
'University_or_other_Tertiary_Institution',
'Males_Negative_Nil_income', #maybe look at this whole section
'Females_Negative_Nil_income', #maybe look at this whole section
'Persons_Negative_Nil_income', #maybe look at this whole section
'Males_Personal_income_not_stated',#maybe look at this whole section
'Feales_Personal_income_not_stated',#maybe look at this whole section
'Persons_Personal_income_not_stated',#maybe look at this whole section
'Cared_for_Own',
'Cared_for_Other',
'Visitor_from_within_Australia',
'born_in_Australia',
'Real_Estate_Agent',
'with_Children',
'with__No_children',
'Flat_or_Apartment',
'Graduate_Diploma_and_Graduate_Certificate_Level_Graduate_Diploma_Level',
'Advanced_Diploma_and_Diploma_Level_Advanced_Diploma_and_Associate_Degree_Level',
'Advanced_Diploma_and_Diploma_Advanced_Diploma_and_Associate_Degree_Level',
'Certificate_Level_Certificate_III_and_IV_Level',
'Certificate_Level_Certificate_I_and_II_Level',
'Certificate_Level_Certificate_Level_nfd',
'Graduate_Diploma_and_Graduate_Certificate_Level',
'Advanced_Diploma_and_Diploma_Level_Diploma_Level',
'Occupation_Inadequately_described',
'Postgraduate_Degree_Level',
'Master_Degree_Level',
'Doctoral_Degree_Level',
'Certificate_I_and_II_Level',
'Certificate_III_and_IV_Level',
'Advanced_Diploma_and_Diploma_Level',
'Graduate_Certificate_Level',
'Bachelor_Degree_Level',
'Certificate_Level',
'Different_SA2',
'Same_Statistical_Area_Level_2',
'Lone_Parent',
'Worked_Full_Time',
'Worked_Part_Time',
'Away_From_Work',
'Age_Of_Dependent_Children',
'4_Years',
'9_Years',
'2_Years',
'7_Years',
'0_Years',
'Hours_Worked',
'Labour_Force_Status_Not_Stated',
'Not_In_The_Labour_Force',
'Labour_Force',
'Looking_For_Full_Time_Work',
'Looking_For_Part_Time_Work',
'Natural_and_Physical_Sciences',
'Information_Technology',
'Engineering_and_Related_Technologies',
'Architecture_and_Building',
'Agriculture_Environmental_and_Related_Studies',
'Management_and_Commerce',
'Society_and_Culture',
'Creative_Arts',
'Food_Hospitality_and_Personal_Services',
'Mixed_Field_Programmes',
'Male_Parent',
'Looking_For',
'Hours_Worked_Not_Stated',
'Inadequately_described_Not_stated',
'Number_of_hours_worked_None',
'Number_of_hours_worked_Not_stated',
'Dependent_children_In_Couple_Families',
'Negative_Nil',
'Never_Married',
'_Census_night',
'Speaks_English',
'Speaks_english',
'Speaks_other'
]
# note: there were still some areas I missed and had to clean up in the tedious manual work on the csv file
# -
replace_capitalisation_list.sort(key = len, reverse=True)
for correction in replace_capitalisation_list:
lower_all_except_first_char(correction)
df_meta_measures.tail(20)
# +
# replace any words following the word "and" with a lowercase version
replace_ands = set()
for cats in df_meta_measures.Long.unique():
try:
word = re.search('(?<=_and_)\w+', cats).group(0).split('_')[0]
replace_ands.add('_and_{}'.format(word))
except:
pass
for repl in replace_ands:
df_meta_measures['Long'] = df_meta_measures['Long'].str.replace(repl,repl.lower())
# replace any words following the word "Occupation" with a lowercase version
replace_occs = set()
for cats in df_meta_measures.Long.unique():
try:
word = re.search('(?<=ccupation_)\w+', cats).group(0).split('_')[0]
replace_occs.add('ccupation_{}'.format(word))
except:
pass
for repl in replace_occs:
df_meta_measures['Long'] = df_meta_measures['Long'].str.replace(repl,repl.lower())
# -
measure_cats = []
for category in df_meta_measures.Long.tolist():
measure_cats.append(re.findall('[A-Z][^A-Z]*', category))
#export list to csv for framework to build reference table
with open("out.csv", "w", newline="") as f:
writer = csv.writer(f)
writer.writerows(measure_cats)
# ### Begin working with new metaclasses file
# This file has a kind of "OneHotEncoding" to assist infer which classes particular measures fall into, e.g. do they have detail for "Age", "Sex", "Place of Birth", etc.
df_meta = pd.read_csv('{}\Data\Metadata\Metadata_2016_refined.csv'.format(nb_path))
df_meta.head()
#Thoughts on algorithms to select line items
# For table in table_list
# slice meta based on table
# for category in filter_cats
# slice based on category >0
# select rows with lowest value in "Number of Classes Excl Total" field
test_tbl = 'G57'
test_cats = ['Sex','Age'] # testing for rows that have a "Total" row first
meta_df_select = df_meta[df_meta['Profile table'].str[:3] == test_tbl].copy()
for cat in test_cats:
meta_df_select = meta_df_select[meta_df_select[cat]>0]
min_fields = meta_df_select['Number of Classes Excl Total'].min()
meta_df_select = meta_df_select[meta_df_select['Number of Classes Excl Total'] == min_fields]
print(meta_df_select.shape)
meta_df_select
# Select the table file(s) to import
import_table_list = meta_df_select['DataPack file'].unique()
# Import the data table itself
df_data = load_census_csv(import_table_list, 'SA3')
# Select only columns included in the meta-sliced table above
df_data.set_index(df_data.columns[0], inplace=True)
refined_columns = meta_df_select.Short.tolist()
df_data = df_data[refined_columns]
df_data.shape
# aggregate data by:
# transposing the dataframe
df_data_t = df_data.T.reset_index()
df_data_t.rename(columns={ df_data_t.columns[0]: 'Short' }, inplace = True)
# merging with the refined meta_df to give table name, "Measures" and "Categories" fields
meta_merge_ref = meta_df_select[['Short','Table name','Measures','Categories']]
df_data_t = df_data_t.merge(meta_merge_ref, on='Short')
df_data_t.head()
# +
#get positions in measures field for selected categories
position_list = []
for i, j in enumerate(df_data_t.Categories.iloc[0].split("|")):
if j in test_cats:
position_list.append(i)
print(position_list)
table_name = df_data_t['Table name'].iloc[0].replace(" ","_") + "|"
# +
# from the "Categories" field, you should be able to split an individual entry by the "|" character
# to give the index of the measure you are interested in grouping by
# create a new column based on splitting the "Measure" field and selecting the value of this index/
# or textjoining indices where there are multiple categories selected
# also remove any instance of the word "total" and any multiple underscores
# Merge above with the table name to form "[Table_Name]|[groupby_value]" to have a good naming convention
# eg "Method_of_Travel_to_Work_by_Sex|Three_methods_Females"
#p = table_name + '_'.join([df_data_t.Measures.iloc[0].split("|")[i] for i in position_list])
def custom_name(table_namer, string_item, measure_positions):
return table_namer + '_'.join([string_item.split("|")[i] for i in measure_positions])
df_data_t['Test_name'] = df_data_t.apply(lambda x: custom_name(x['Table name'], x['Measures'], position_list), axis=1)
df_data_t.head()
# -
# then groupby this new column
# then transpose again and either create the base data_df for future merges or merge with the already existing data_df
df_data_t = df_data_t.drop(['Short','Table name','Measures','Categories'], axis=1)
df_data_t = df_data_t.groupby(['Test_name']).sum()
df_data_t.head()
df_data = df_data_t.T
df_data.head()
# +
# Consolidate into a single function
def refine_measure_name(table_namer, string_item, category_item, category_list):
'''Simple function for generating measure names based on custom metadata information on ABS measures'''
position_list = []
for i, j in enumerate(category_item.split("|")):
if j in category_list:
position_list.append(i)
return table_namer + '|' + '_'.join([string_item.split("|")[i] for i in position_list])
def load_table_refined(table_ref, category_list, statistical_area_code='SA3'):
'''
Function for loading ABS census data tables, and refining/aggregating by a set of defined categories
(e.g. age, sex, occupation, English proficiency, etc.) where available.
INPUTS
table_ref: STRING - the ABS Census Datapack table to draw information from (G01-G59)
category_list: LIST of STRING objects - Cetegorical informatio to slice/aggregate information from (e.g. Age)
statistical_area_code: STRING - the ABS statistical area level of detail required (SA1-SA3)
'''
df_meta = pd.read_csv('{}\Data\Metadata\Metadata_2016_refined.csv'.format(os.getcwd()))
# slice meta based on table
meta_df_select = df_meta[df_meta['Profile table'].str[:3] == table_ref].copy()
# for category in filter_cats, slice based on category >0
for cat in category_list:
# First, check if there *are* any instances of the given category
if meta_df_select[cat].sum() > 0:
# If so, apply the filter
meta_df_select = meta_df_select[meta_df_select[cat]>0]
else:
pass # If not, don't apply (otherwise you will end up with no selections)
# select rows with lowest value in "Number of Classes Excl Total" field
min_fields = meta_df_select['Number of Classes Excl Total'].min()
meta_df_select = meta_df_select[meta_df_select['Number of Classes Excl Total'] == min_fields]
# Select the table file(s) to import
import_table_list = meta_df_select['DataPack file'].unique()
# Import the SA data tables
df_data = load_census_csv(import_table_list, statistical_area_code.upper())
# Select only columns included in the meta-sliced table above
df_data.set_index(df_data.columns[0], inplace=True)
refined_columns = meta_df_select.Short.tolist()
df_data = df_data[refined_columns]
# aggregate data by:
# transposing the dataframe
df_data_t = df_data.T.reset_index()
df_data_t.rename(columns={ df_data_t.columns[0]: 'Short' }, inplace = True)
# merging with the refined meta_df to give table name, "Measures" and "Categories" fields
meta_merge_ref = meta_df_select[['Short','Table name','Measures','Categories']]
df_data_t = df_data_t.merge(meta_merge_ref, on='Short')
# from the "Categories" field, you should be able to split an individual entry by the "|" character
# to give the index of the measure you are interested in grouping by
# create a new column based on splitting the "Measure" field and selecting the value of this index/indices
# Merge above with the table name to form "[Table_Name]|[groupby_value]" to have a good naming convention
# eg "Method_of_Travel_to_Work_by_Sex|Three_methods_Females"
df_data_t['Test_name'] = df_data_t.apply(lambda x: refine_measure_name(x['Table name'],
x['Measures'],
x['Categories'],
category_list), axis=1)
# then groupby this new column
# then transpose again and either create the base data_df for future merges or merge with the already existing data_df
df_data_t = df_data_t.drop(['Short','Table name','Measures','Categories'], axis=1)
df_data_t = df_data_t.groupby(['Test_name']).sum()
return df_data_t.T
# -
# Test aggregation on import on category detail without "Total" fields
df_no_total = load_table_refined('G01', ['Education'])
df_no_total.head()
# Test import on table with no category detail matching
df_no_match = load_table_refined('G01', ['Occupation'])
df_no_match.head()
# Stitch it together to import multiple tables and multiple categories
def load_tables_specify_cats(table_list, category_list, statistical_area_code='SA3'):
for index, table in enumerate(table_list):
if index==0:
df = load_table_refined(table, category_list, statistical_area_code)
df.reset_index(inplace=True)
else:
temp_df = load_table_refined(table, category_list, statistical_area_code)
temp_df.reset_index(inplace=True)
merge_col = df.columns[0]
df = pd.merge(df, temp_df, on=merge_col)
return df
df_y = load_tables_specify_cats(['G01','G57','G58'],['Age','Occupation','Number of Commuting Methods'])
df_y.head()
# ### Remaining Challenges
# Ideally I would like to also have a mechanism to bring in sub-sets where available, and if not available still bring in the super-set (e.g. bring in "Religious Sect" detail where possible [Christianity] and for those religions without "Sect" breakdowns, still bring them in). Currently if you choose both "Religious Sect" and "Religion" as a category, you will get a double up of the "Total" lines of the "Religious Sect" subset, which could confuse some algorithms using the data, and would need to be manually picked out and removed.
#
# It would also be nice to have a simple mechanism for showing which categories are available within which tables.
| .ipynb_checkpoints/Census data navigation workings-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from scipy.stats import wishart, gamma
import matplotlib.pyplot as plt
df, scale = 7, 3
x = np.linspace(1e-5, 24, 100)
ws = wishart.pdf(x, df=df, scale=scale) # chi2.pdf(x, df)
print(ws[:5])
plt.plot(x, ws)
gm = gamma.pdf(x, a=df/2, scale=scale * 2)
print(gm[:5])
plt.plot(x, gm)
| scratchpad.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Chapter 8 - Tree-based Methods
# - [8.1.1 Regression Trees](#8.1.1-Regression-Trees)
# - [8.1.2 Classification Trees](#8.1.2-Classification-Trees)
# - [Lab: 8.3.1 Fitting Classification Trees](#8.3.1-Fitting-Classification-Trees)
# - [Lab: 8.3.2 Fitting Regression Trees](#8.3.2-Fitting-Regression-Trees)
# - [Lab: 8.3.3 Bagging and Random Forests](#8.3.3-Bagging-and-Random-Forests)
# - [Lab: 8.3.4 Boosting](#8.3.4-Boosting)
# +
# # %load ../standard_import.txt
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import pydot
from IPython.display import Image
from sklearn.model_selection import train_test_split, cross_val_score
from six import StringIO
from sklearn.tree import DecisionTreeRegressor, DecisionTreeClassifier, export_graphviz
from sklearn.ensemble import BaggingClassifier, RandomForestClassifier, BaggingRegressor, RandomForestRegressor, GradientBoostingRegressor
from sklearn.metrics import mean_squared_error,confusion_matrix, classification_report
# %matplotlib inline
plt.style.use('seaborn-white')
# -
# This function creates images of tree models using pydot
def print_tree(estimator, features, class_names=None, filled=True):
tree = estimator
names = features
color = filled
classn = class_names
dot_data = StringIO()
export_graphviz(estimator, out_file=dot_data, feature_names=features, class_names=classn, filled=filled)
graph = pydot.graph_from_dot_data(dot_data.getvalue())
return(graph)
# ### 8.1.1 Regression Trees
# In R, I exported the dataset from package 'ISLR' to a csv file.
df = pd.read_csv('../Datasets/Hitters.csv').dropna()
df.info()
# +
X = df[['Years', 'Hits']].to_numpy()
y = np.log(df.Salary.to_numpy())
fig, (ax1, ax2) = plt.subplots(1,2, figsize=(11,4))
ax1.hist(df.Salary.to_numpy())
ax1.set_xlabel('Salary')
ax2.hist(y)
ax2.set_xlabel('Log(Salary)');
# -
regr = DecisionTreeRegressor(max_leaf_nodes=3)
regr.fit(X, y)
# ### Figure 8.1
graph, = print_tree(regr, features=['Years', 'Hits'])
Image(graph.create_png())
# ### Figure 8.2
df.plot('Years', 'Hits', kind='scatter', color='orange', figsize=(7,6))
plt.xlim(0,25)
plt.ylim(ymin=-5)
plt.xticks([1, 4.5, 24])
plt.yticks([1, 117.5, 238])
plt.vlines(4.5, ymin=-5, ymax=250)
plt.hlines(117.5, xmin=4.5, xmax=25)
plt.annotate('R1', xy=(2,117.5), fontsize='xx-large')
plt.annotate('R2', xy=(11,60), fontsize='xx-large')
plt.annotate('R3', xy=(11,170), fontsize='xx-large');
# ### Pruning
# This is currently not supported in scikit-learn. See first point under 'disadvantages of decision trees in the <A href='http://scikit-learn.github.io/dev/modules/tree.html#'>documentation</A>. Implementation has been <A href='https://github.com/scikit-learn/scikit-learn/pull/941'>discussed</A> but Random Forests have better predictive qualities than a single pruned tree anyway if I understand correctly.
#
# ### 8.1.2 Classification Trees
# Dataset available on http://www-bcf.usc.edu/~gareth/ISL/data.html
df2 = pd.read_csv('../Datasets/Heart.csv').drop('Unnamed: 0', axis=1).dropna()
df2.info()
df2.ChestPain = pd.factorize(df2.ChestPain)[0]
df2.Thal = pd.factorize(df2.Thal)[0]
X2 = df2.drop('AHD', axis=1)
y2 = pd.factorize(df2.AHD)[0]
clf = DecisionTreeClassifier(max_depth=None, max_leaf_nodes=6, max_features=3)
clf.fit(X2,y2)
clf.score(X2,y2)
graph2, = print_tree(clf, features=X2.columns, class_names=['No', 'Yes'])
Image(graph2.create_png())
# ## Lab
# ### 8.3.1 Fitting Classification Trees
# In R, I exported the dataset from package 'ISLR' to a csv file.
#df3 = pd.read_csv('Carseats.csv').drop('Unnamed: 0', axis=1) #Unnamed does not exist in the provided file
df3 = pd.read_csv('../Datasets/Carseats.csv')
df3.head()
# +
df3['High'] = df3.Sales.map(lambda x: 1 if x>8 else 0)
df3.ShelveLoc = pd.factorize(df3.ShelveLoc)[0]
df3.Urban = df3.Urban.map({'No':0, 'Yes':1})
df3.US = df3.US.map({'No':0, 'Yes':1})
df3.info()
# -
df3.head(5)
# +
X = df3.drop(['Sales', 'High'], axis=1)
y = df3.High
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.5, random_state=0)
# -
clf = DecisionTreeClassifier(max_depth=6)
clf.fit(X, y)
print(classification_report(y, clf.predict(X)))
graph3, = print_tree(clf, features=X.columns, class_names=['No', 'Yes'])
Image(graph3.create_png())
clf.fit(X_train, y_train)
pred = clf.predict(X_test)
cm = pd.DataFrame(confusion_matrix(y_test, pred).T, index=['No', 'Yes'], columns=['No', 'Yes'])
cm.index.name = 'Predicted'
cm.columns.name = 'True'
cm
# Precision of the model using test data is 74%
print(classification_report(y_test, pred))
# Pruning not implemented in scikit-learn.
# ### 8.3.2 Fitting Regression Trees
# In R, I exported the dataset from package 'MASS' to a csv file.
boston_df = pd.read_csv('../Datasets/Boston.csv')
boston_df.info()
# +
X = boston_df.drop('medv', axis=1)
y = boston_df.medv
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.5, random_state=0)
# -
# Pruning not supported. Choosing max depth 3)
regr2 = DecisionTreeRegressor(max_depth=3)
regr2.fit(X_train, y_train)
pred = regr2.predict(X_test)
graph, = print_tree(regr2, features=X.columns)
Image(graph.create_png())
plt.scatter(pred, y_test, label='medv')
plt.plot([0, 1], [0, 1], '--k', transform=plt.gca().transAxes)
plt.xlabel('pred')
plt.ylabel('y_test')
mean_squared_error(y_test, pred)
# ### 8.3.3 Bagging and Random Forests
# There are 13 features in the dataset
X.shape
# Bagging: using all features
regr1 = RandomForestRegressor(max_features=13, random_state=1)
regr1.fit(X_train, y_train)
# +
pred = regr1.predict(X_test)
plt.scatter(pred, y_test, label='medv')
plt.plot([0, 1], [0, 1], '--k', transform=plt.gca().transAxes)
plt.xlabel('pred')
plt.ylabel('y_test')
# -
mean_squared_error(y_test, pred)
# Random forests: using 6 features
regr2 = RandomForestRegressor(max_features=6, random_state=1)
regr2.fit(X_train, y_train)
pred = regr2.predict(X_test)
mean_squared_error(y_test, pred)
Importance = pd.DataFrame({'Importance':regr2.feature_importances_*100}, index=X.columns)
Importance.sort_values('Importance', axis=0, ascending=True).plot(kind='barh', color='r', )
plt.xlabel('Variable Importance')
plt.gca().legend_ = None
# ### 8.3.4 Boosting
regr = GradientBoostingRegressor(n_estimators=500, learning_rate=0.01, random_state=1)
regr.fit(X_train, y_train)
feature_importance = regr.feature_importances_*100
rel_imp = pd.Series(feature_importance, index=X.columns).sort_values(inplace=False)
print(rel_imp)
rel_imp.T.plot(kind='barh', color='r', )
plt.xlabel('Variable Importance')
plt.gca().legend_ = None
mean_squared_error(y_test, regr.predict(X_test))
| Trees_tryout/trees_binder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Q.1)# list1=[10,20,30,40,50]
# list2=[5,15,25,35,45,60]
# Merge the two lists and give output as follows with new list
# Note:-
# #You have to use any one loop only one time
# #Don’t use in built functions of list i’es sort and sorted
# Without using this write the code and give the output
#
# +
def Merge(list1, list2):
final_list = list1 + list2
final_list.sort()
return(final_list)
# Driver Code
list1 = [10, 20, 30, 40, 50]
list2 = [5, 15, 25, 35, 45, 60]
print(Merge(list1, list2))
# -
| DAY 7 ASSIGNMENT .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # matplotlib-applied
#
# * Applying Matplotlib Visualizations to Kaggle: Titanic
# * Bar Plots, Histograms, subplot2grid
# * Normalized Plots
# * Scatter Plots, subplots
# * Kernel Density Estimation Plots
# ## Applying Matplotlib Visualizations to Kaggle: Titanic
# Prepare the titanic data to plot:
# +
# %matplotlib inline
import pandas as pd
import numpy as np
import pylab as plt
import seaborn
# Set the global default size of matplotlib figures
plt.rc('figure', figsize=(10, 5))
# Set seaborn aesthetic parameters to defaults
seaborn.set()
# +
df_train = pd.read_csv('../data/titanic/train.csv')
def clean_data(df):
# Get the unique values of Sex
sexes = np.sort(df['Sex'].unique())
# Generate a mapping of Sex from a string to a number representation
genders_mapping = dict(zip(sexes, range(0, len(sexes) + 1)))
# Transform Sex from a string to a number representation
df['Sex_Val'] = df['Sex'].map(genders_mapping).astype(int)
# Get the unique values of Embarked
embarked_locs = np.sort(df['Embarked'].unique())
# Generate a mapping of Embarked from a string to a number representation
embarked_locs_mapping = dict(zip(embarked_locs,
range(0, len(embarked_locs) + 1)))
# Transform Embarked from a string to dummy variables
df = pd.concat([df, pd.get_dummies(df['Embarked'], prefix='Embarked_Val')], axis=1)
# Fill in missing values of Embarked
# Since the vast majority of passengers embarked in 'S': 3,
# we assign the missing values in Embarked to 'S':
if len(df[df['Embarked'].isnull()] > 0):
df.replace({'Embarked_Val' :
{ embarked_locs_mapping[np.nan] : embarked_locs_mapping['S']
}
},
inplace=True)
# Fill in missing values of Fare with the average Fare
if len(df[df['Fare'].isnull()] > 0):
avg_fare = df['Fare'].mean()
df.replace({ None: avg_fare }, inplace=True)
# To keep Age in tact, make a copy of it called AgeFill
# that we will use to fill in the missing ages:
df['AgeFill'] = df['Age']
# Determine the Age typical for each passenger class by Sex_Val.
# We'll use the median instead of the mean because the Age
# histogram seems to be right skewed.
df['AgeFill'] = df['AgeFill'] \
.groupby([df['Sex_Val'], df['Pclass']]) \
.apply(lambda x: x.fillna(x.median()))
# Define a new feature FamilySize that is the sum of
# Parch (number of parents or children on board) and
# SibSp (number of siblings or spouses):
df['FamilySize'] = df['SibSp'] + df['Parch']
return df
df_train = clean_data(df_train)
# -
# ## Bar Plots, Histograms, subplot2grid
# +
# Size of matplotlib figures that contain subplots
figsize_with_subplots = (10, 10)
# Set up a grid of plots
fig = plt.figure(figsize=figsize_with_subplots)
fig_dims = (3, 2)
# Plot death and survival counts
plt.subplot2grid(fig_dims, (0, 0))
df_train['Survived'].value_counts().plot(kind='bar',
title='Death and Survival Counts',
color='r',
align='center')
# Plot Pclass counts
plt.subplot2grid(fig_dims, (0, 1))
df_train['Pclass'].value_counts().plot(kind='bar',
title='Passenger Class Counts')
# Plot Sex counts
plt.subplot2grid(fig_dims, (1, 0))
df_train['Sex'].value_counts().plot(kind='bar',
title='Gender Counts')
plt.xticks(rotation=0)
# Plot Embarked counts
plt.subplot2grid(fig_dims, (1, 1))
df_train['Embarked'].value_counts().plot(kind='bar',
title='Ports of Embarkation Counts')
# Plot the Age histogram
plt.subplot2grid(fig_dims, (2, 0))
df_train['Age'].hist()
plt.title('Age Histogram')
# +
# Get the unique values of Embarked and its maximum
family_sizes = np.sort(df_train['FamilySize'].unique())
family_size_max = max(family_sizes)
df1 = df_train[df_train['Survived'] == 0]['FamilySize']
df2 = df_train[df_train['Survived'] == 1]['FamilySize']
plt.hist([df1, df2],
bins=family_size_max + 1,
range=(0, family_size_max),
stacked=True)
plt.legend(('Died', 'Survived'), loc='best')
plt.title('Survivors by Family Size')
# -
# ## Normalized Plots
# +
pclass_xt = pd.crosstab(df_train['Pclass'], df_train['Survived'])
# Normalize the cross tab to sum to 1:
pclass_xt_pct = pclass_xt.div(pclass_xt.sum(1).astype(float), axis=0)
pclass_xt_pct.plot(kind='bar',
stacked=True,
title='Survival Rate by Passenger Classes')
plt.xlabel('Passenger Class')
plt.ylabel('Survival Rate')
# Plot survival rate by Sex
females_df = df_train[df_train['Sex'] == 'female']
females_xt = pd.crosstab(females_df['Pclass'], df_train['Survived'])
females_xt_pct = females_xt.div(females_xt.sum(1).astype(float), axis=0)
females_xt_pct.plot(kind='bar',
stacked=True,
title='Female Survival Rate by Passenger Class')
plt.xlabel('Passenger Class')
plt.ylabel('Survival Rate')
# Plot survival rate by Pclass
males_df = df_train[df_train['Sex'] == 'male']
males_xt = pd.crosstab(males_df['Pclass'], df_train['Survived'])
males_xt_pct = males_xt.div(males_xt.sum(1).astype(float), axis=0)
males_xt_pct.plot(kind='bar',
stacked=True,
title='Male Survival Rate by Passenger Class')
plt.xlabel('Passenger Class')
plt.ylabel('Survival Rate')
# -
# ## Scatter Plots, subplots
# +
# Set up a grid of plots
fig, axes = plt.subplots(2, 1, figsize=figsize_with_subplots)
# Histogram of AgeFill segmented by Survived
df1 = df_train[df_train['Survived'] == 0]['Age']
df2 = df_train[df_train['Survived'] == 1]['Age']
max_age = max(df_train['AgeFill'])
axes[1].hist([df1, df2],
bins=max_age / 10,
range=(1, max_age),
stacked=True)
axes[1].legend(('Died', 'Survived'), loc='best')
axes[1].set_title('Survivors by Age Groups Histogram')
axes[1].set_xlabel('Age')
axes[1].set_ylabel('Count')
# Scatter plot Survived and AgeFill
axes[0].scatter(df_train['Survived'], df_train['AgeFill'])
axes[0].set_title('Survivors by Age Plot')
axes[0].set_xlabel('Survived')
axes[0].set_ylabel('Age')
# -
# ## Kernel Density Estimation Plots
# +
# Get the unique values of Pclass:
passenger_classes = np.sort(df_train['Pclass'].unique())
for pclass in passenger_classes:
df_train.AgeFill[df_train.Pclass == pclass].plot(kind='kde')
plt.title('Age Density Plot by Passenger Class')
plt.xlabel('Age')
plt.legend(('1st Class', '2nd Class', '3rd Class'), loc='best')
| matplotlib/matplotlib-applied.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Time Series Prediction with Stateful LSTM
#
# ### Abstract
# This Jupyter notebook demonstrates the use of Google Tensorflow to perform a time series prediction task.
#
# Here, a recurrent neural network, more specifically a stateful Long Short Term Memory (LSTM) is used.
#
# <b>Tested on Google TensorFlow V1.0 (https://github.com/tensorflow/tensorflow)</b>
#
# #### References:
# 1. <NAME>'s Keras example using <i>stateful LSTM</i> (https://github.com/fchollet/keras/blob/master/examples/stateful_lstm.py)
#
# #### Dependencies
# 1. Python 3.5.2
# 2. TensorFlow (GPU) 1.0.1
# 3. Keras 2.0.2 (refer to https://keras.io/backend/ to configure backend)
#
# ##### Overview:
# 1. Import Libraries
# 2. Setup Environment
# 3. Configure Parameters
# 4. Prepare Data
# 5. Configure LSTM Network
# 6. Train
# 7. Evaluate
# 8. Visualize Prediction
# ### Step 1: Import Libraries
# +
# Generic Libraries
import os, sys, warnings
import numpy as np
import math
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.preprocessing import MinMaxScaler
from sklearn.metrics import mean_squared_error
# ML Libraries
import keras
import tensorflow as tf
# Keras Libraries
from keras.models import Sequential
from keras.layers import Dense, LSTM
from keras.callbacks import EarlyStopping, ModelCheckpoint, TensorBoard
# -
# ### Step 2: Setup Environment
# +
# enable inline plotting of graphs within Jupyter notebook
# %matplotlib inline
# ensure reproducible output
np.random.seed(7)
# suppress non-essential warnings
os.environ['TF_CPP_MI_LOG_LEVEL'] = '3' # tensorflow
#warnings.filterwarnings("ignore") # numpy
# confirm versions of key libraries
print("python:{}, \nkeras:{}, \ntensorflow:{}".format(sys.version, keras.__version__, tf.__version__))
# -
# ### Step 3: Configure Parameters
# Hyper-parameters
timesteps = 1 # using stateful LSTM
batch_size = 1 # using stateful LSTM
epochs = 10
#lookahead = 1 # number of elements ahead to predict
num_features = 1 # number of features
data_size = 21600
# ### Step 4: Prepare Data
#
# 1. Load Data
# 2. Normalize OR Standardize Data - state difference
# 3. Prepare Train/Test Samples
def load_dataset(filename, header, usecols, skipfooter, delimiter=None, delim_whitespace=False):
df = pd.read_csv(
filename,
header=header,
usecols=usecols,
skipfooter=skipfooter,
engine='python',
delimiter=delimiter,
delim_whitespace=delim_whitespace
)
print(df.describe())
plt.figure(figsize=(15,4))
plt.title('Selected Data')
plt.plot(df[0:data_size])
plt.show()
dataset = df.values
dataset = dataset.astype('float32')
return dataset
# +
# load the dataset
# https://datamarket.com/data/set/22u3/international-airline-passengers-monthly-totals-in-thousands-jan-49-dec-60#!ds=22u3&display=line
#dataset = load_dataset(filename='data/international-airline-passengers.csv', header='infer', usecols=[1], skipfooter=2)
#dataset = load_dataset(filename='data/sp500.csv', header=None, usecols=[0], skipfooter=0)
#dataset = load_dataset(filename='data/household_power_consumption_cleansed.txt', header='infer', usecols=[2], skipfooter=0, delimiter=';')
dataset = load_dataset(filename='data/mitdbx_mitdbx_108.txt', header=None, usecols=[1], skipfooter=0, delim_whitespace=True)
# -
# normalize using MinMax to obtain a range of (0,1)
def normalize_dataset(dataset):
scaler = MinMaxScaler(feature_range=(0, 1))
return scaler, scaler.fit_transform(dataset)
scaler, dataset_norm = normalize_dataset(dataset)
print('Dataset normalized.')
# convert an array of values into train and test data
def split_train_test_data(dataset, look_back, split):
dataX, dataY = [], []
for i in range(len(dataset) - look_back):
dataX.append(dataset[i:(i + look_back), 0])
dataY.append(dataset[i + look_back, 0])
dataX = np.array(dataX)
dataY = np.array(dataY)
# split into train and test sets
train_size = int(len(dataset) * split)
test_size = len(dataset) - train_size
trainX, testX = dataX[0:train_size,:], dataX[train_size:len(dataX),:]
trainY, testY = dataY[0:train_size], dataY[train_size:len(dataY)]
# reshape input to be [samples, time steps, features]
trainX = np.reshape(trainX, (trainX.shape[0], trainX.shape[1], 1))
testX = np.reshape(testX, (testX.shape[0], testX.shape[1], 1))
# add last entry from dataset to be able to predict unknown testY
temp = dataset[len(dataset) - timesteps:len(dataset), 0]
temp = np.reshape(temp, (1, temp.shape[0]))
temp = np.reshape(temp, (temp.shape[0], temp.shape[1], 1))
testX = np.append(testX, temp, 0)
return trainX, trainY, testX, testY
trainX, trainY, testX, testY = split_train_test_data(dataset_norm[0:data_size,:], look_back=timesteps, split=0.67)
print('Data split into train and test set.')
# ### Step 5: Configure LSTM Network
def build_model():
model = Sequential()
model.add(LSTM(4,
batch_input_shape=(batch_size, timesteps, num_features),
stateful=True,
return_sequences=True))
model.add(LSTM(4, # There is no need for an batch_input_shape in the second layer
return_sequences=False,
stateful=True))
model.add(Dense(1))
model.compile(loss='mean_squared_error', optimizer='adam')
return model
# +
print('Model Summary')
model = build_model()
model.summary()
# implement callbacks
# 1. Early Stopping
early_stopping = EarlyStopping(monitor='val_loss', patience=10)
# 2. Model Checkpoint
checkpointer = ModelCheckpoint(filepath='keras_weights.hdf5', verbose=1, save_best_only=True)
# 3. TensorBoard
# to monitor: tensorboard --logdir "complete_path_to\Graph"
tensorboard = TensorBoard(log_dir='./Graph', histogram_freq=0, write_graph=True, write_images=True)
# -
# ### Step 6: Train
print('Start training...')
loss_history = []
val_loss_history = []
for i in range(epochs):
print('Epoch', i+1, '/', epochs)
# Note that the last state for sample i in a batch will
# be used as initial state for sample i in the next batch.
# Thus we are simultaneously training on batch_size series with
# lower resolution than the original series contained in cos.
# Each of these series are offset by one step and can be
# extracted with cos[i::batch_size].
history = model.fit(
trainX,
trainY,
epochs=1,
batch_size=batch_size,
validation_split=0.08,
verbose=2,
shuffle=False,
callbacks=[early_stopping, checkpointer, tensorboard]
)
loss_history.append(history.history['loss'])
val_loss_history.append(history.history['val_loss'])
model.reset_states() # required for stateful LSTMs
# summarize history for loss
plt.plot(loss_history)
plt.plot(val_loss_history)
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train loss', 'val loss'], loc='upper left')
plt.show()
# ### Step 7: Evaluate
# +
# make predictions on train and test
# train data
trainPredict = model.predict(trainX, batch_size=batch_size)
# reset states
model.reset_states()
# test data
testPredict = model.predict(testX, batch_size=batch_size)
print('Prediction performed against train and test data.')
# +
# invert predictions to revert to initial scale
trainPredict = scaler.inverse_transform(trainPredict)
trainY = scaler.inverse_transform([trainY])
testPredict = scaler.inverse_transform(testPredict)
testY = scaler.inverse_transform([testY])
# -
# print next predicted value
print('Prediction of the next value:', round(testPredict[len(testPredict) - 1, 0],3))
# calculate root mean squared error
trainScore = math.sqrt(mean_squared_error(trainY[0], trainPredict[:,0]))
print('Train Score: %.2f RMSE' % (trainScore))
# ### Step 8: Visualize
# +
print('Visualize results')
# shift train predictions for plotting
trainPredictPlot = np.empty_like(dataset_norm[0:data_size,:]) # create array with same shape as dataset
trainPredictPlot[:, :] = np.nan # fill with nan
trainPredictPlot[timesteps:len(trainPredict) + timesteps, :] = trainPredict
# shift test predictions for plotting
testPredictPlot = np.empty_like(dataset_norm[0:data_size,:]) # create array with same shape as dataset
testPredictPlot[:, :] = np.nan # fill with nan
test = ['nan']
test = np.reshape(test, (len(test), 1))
testPredictPlot = np.append(testPredictPlot, test, 0)
testPredictPlot[len(trainPredict) + timesteps:len(dataset_norm[0:data_size,:]) + 1, :] = testPredict
# plot baseline and predictions
plt.figure(figsize=(15,4))
plt.title('All Data')
plt.plot(scaler.inverse_transform(dataset_norm[0:data_size,:]), color='b', label='Actual')
plt.plot(trainPredictPlot, color='g', label='Training')
plt.plot(testPredictPlot, color='r', label='Prediction')
plt.grid(True)
plt.grid(b=True, which='minor', axis='both')
plt.minorticks_on()
plt.legend()
plt.show()
# zoom in on the test predictions
plt.figure(figsize=(15,4))
plt.title('Test Predictions')
testYPlot = np.reshape(testY, [testY.shape[1], 1]) # testY is [0, n] need to change shape to [n, 0] to plot it
testYPlot = testYPlot[testYPlot.shape[0] - len(testPredict) + 1:testYPlot.shape[0]] # +1, because testY has one less value than testPredict
plt.plot(testYPlot, color='b', label='testY')
testPredictPlot = testPredict[len(testPredict) - len(testPredict):len(testPredict), :]
plt.plot(testPredictPlot, color='r', label='Prediction')
plt.grid(True)
plt.grid(b=True, which='minor', axis='both')
plt.minorticks_on()
plt.legend()
plt.show()
# -
| Time Series/LSTM/Stateful_LSTM_Time Series.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Round 2 - Convolutional Neural Networks
# Authors: <NAME> and <NAME>
# Welcome to second round of the course "Deep Learning with Python". In the previous round (Round 1 - "ANN"), you have learnt how to use simple aritifical neural networks to classify images according to what type of fashion item they show. In this round, you will learn about an important class of deep neural network which are particularly suitable for image data. This class of deep neural network is referred to as convolutional neural networks (CNN).
#
# **Learning Goals.** In this round you will learn:
#
# - about the basic network structure of CNNs.
# - how to use the "padding" and "stride" parameters in CNN.
# - how to determine CNN parameters required for a given data set.
# - how to visualize the **activation** (outputs) of different neurons within a CNN.
#
# +
# tensorflow library provides functions for deep neural networks
import tensorflow as tf
# for reproducibility
from numpy.random import seed
seed(1)
tf.random.set_seed(1)
from tensorflow.keras.datasets import fashion_mnist # to load the fashion mnist data set
# import plt library which provides functios to visualize data
import matplotlib.pyplot as plt
# import numpy library which provides functions for matrix computations
import numpy as np
# to get a text report showing the main classification metrics for each class
from sklearn.metrics import classification_report
# %matplotlib inline
# -
# # Dataset
# In this notebook we will be using same dataset as for round 1, i.e., the **"Fashion-MNIST dataset"**. Each data point is represented by a gray scale image of size 28x28. Each image represent the specific fashion item. And each data point is associated with a label taking on values $y=0,...,9$ according to **10 different classes**
#
# We can load this dataset using the command **tf.keras.datasets.fashion_mnist.load_data()**
# load dataset:
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data()
# So, we can see that dataset consists of well defined train and test set.
# shape of train and test image
print(f'Number of training examples: {train_images.shape}')
print(f'Number of test examples: {test_images.shape}')
# <div class='alert alert-info' > Let's further split the train_images and train_labels into train and validation data and normalize it. </div>
# +
# splitting the train set into train and validation set and normalize both dataset
X_train=train_images[:-5000]/255.0
y_train=train_labels[:-5000]
X_valid=train_images[-5000:]/255.0
y_valid=train_labels[-5000:]
test_images=test_images/255.0
print(f'Number of training examples: {X_train.shape[0]}')
print(f'Number of validation examples: {X_valid.shape[0]}')
# -
# Reshaping training and validation set:
X_train=X_train.reshape(-1,28,28,1)
X_valid=X_valid.reshape(-1,28,28,1)
# ### Problem with Simple Artificial Neural Network
# In previous notebook, we saw how we flattened **(28 *28 * 1)** images into one dimensional **(784 * 1)** vector and feed into densly connected neural network.
#
# We got a good result with this approach with **28 x 28** image data.
#
# However, this approach does not scale up for larger images.
# - With larger images, the number of total parameters can grow to very high. For example, flattening **(200 * 200 * 3)** (3 refers to RGB values) and feeding to fully connected neural network, the numbers of parameters (weights) to be learned during training becomes **120,000** in a single fully connected layer. This growing number of parameters can leads to overfitting.
# ### What is Convolutional Neural Network (CNN/ConvNet) ?
# Convolutional Neural Network (CNN) is a special type of neural network architecture that are primarily used with Image data. CNN takes image as an input and assign importance (learnable weights and biases) to various aspects/objects in the image so that they can differentiate one from the other.
#
# Convolutional Neural Networks is extension of traditional Multi-layer Perceptron, based on 3 ideas:
# - Local receptive fields
# - Shared weights
# - Spatial / temporal sub-sampling
# **student feedback Explanation needed of these terms**
# ## CNN architecture
# A CNN consists of sequence of different kinds of layers. The three main types of layers used to form CNN are :
# - Convolution Layer(conv)
# - Pooling Layer(pooling)
# - Fully Connected Layer(FC)
#
#
#
# A typical CNN architecture looks like:
#
# **Input --> Conv --> pooling --> Conv -- Pooling --> ---- --> Fully connected layer --> output**
#
#
# <figure>
# <img src='./convolution/cnn_net.svg' alt='missing' align='middle' />
# <figcaption style="text-align:center"><u> <i> Simple CNN architecture </u> </i>
# </figcaption>
# </figure>
#
# <br>
# <br>
#
#
# Now, Let's briefly describe each layer.
#
# **Input:**
# Unlike in ANN where the input are 1-dimensional vectors, the input to CNN are 2-dimensional matrix like images. To be specific, CNN operates over volume. Below we have an example of image matrix (volume) of dimension 4x4x3.
#
# <figure>
# <img src='./convolution/volume.png' alt='missing' />
# <figcaption style="text-align:center"> <u> <i>Example of input volume to CNN. Image of 4x4x3 dimension with 3 color channel. Image from </u> </i>
# <a href="https://towardsdatascience.com/a-comprehensive-guide-to-convolutional-neural-networks-the-eli5-way-3bd2b1164a53">blog</a>
# </figcaption>
# </figure>
#
# **Convolution Layer:**
# This layer performs the operation called 'convolution'. In context of CNN, convolution is linear mathematical operation between input and fixed size 2-dimensional weights. In CNN, this 2-dimensional weights is called **filter or kernel**. Convolutional layer performs dot product between filter (of size smaller than input) and small region in input (called as **Receptive Fields**) in sliding window fashion. This process of scaning the kernel across image is called convolving. After sliding the filters across whole image results in 2-dimensional scaler valued output. We then apply elementwise non-linear activation function to this 2d output. This resulting 2d output is then called **activation map or feature map**. The convolution layer in CNN acts as an feature extraction layer. The filter learn the particular feature from image. Hence, Convolutional neural networks do not have only single filter. They in fact have multiple filters learning multiple features in parallel for a given input. The important point to remember while doing convolution operation is that the the depth of a filter/kernel should match that of the input depth. Meaning if you have a image of volume 32x32x3 as an input ( where 3 represents 3 color channel: red, blue and green), filter should also be of same depth, for example 5x5x3.
# In CNN, filters are initialized randomly and become parameters (weights) in network which will be learned by the network subsequently.
#
#
# After the brief introduction about convolution layer now let's demonstrate the concept of convolution operation using figures.
# Let's us consider a gray scale image of shape 5x5x1(height x weight x channel) is convolved with a filter of size 3x3x1. We do elementwise multiplication between filter and image pixel values overlapped by filter and sum up the numbers to get a single value of feature map. The first step in convolution looks like below.
#
# <figure>
# <img src='./convolution/c1.png' alt='missing' />
# $$1*1+2*3+3*5+2*2+0*(-1)+0*9+3*1+4*1+4*(-1)= 29$$
# <figcaption style="text-align:center"> <u> <i>First convolution step. We do elementwise multiplication between filter and image pixel values overlapped by filter and sum up the numbers to get a single value of feature map.</u> </i></figcaption>
# </figure>
#
# <br>
# <br>
#
# Next, kernel slide to right by 1 pixel and produces the new value of feature map as shown below.
# <figure>
# <img src="./convolution/c2.png"
# alt="">
# <figcaption style="text-align:center"><u> <i> Second convolution step. This time we move kernel to right by one step </u> </i></figcaption>
# </figure>
#
# <br>
# <br>
#
#
# In similar fashion, kernel slides over whole images (right and down) and produces the complete feature map.
# <figure>
# <img src="./convolution/c3.png"
# alt="">
# <figcaption style="text-align:center"> <u> <i>Final convolution step. Now we have complete feature map.</u> </i></figcaption>
# </figure>
#
# <br>
# <br>
#
#
# In above convolution operation we saw that kernel is traversing across the image by one pixel at a time. We call a number of pixel( or step size) by which kernel traversed in each slide a **stride**. Also, we saw that size of output from convolution operation, i.e feature map is smaller than input image size. This means we are losing some pixel values around the perimeter of image. Since CNN might consists of number of convolutional layer, loss of pixels values in each successive convolution layer might result in loss of important features from image. So, to get the input sized output, we employ a technique called **zero padding**. Padding is a technique in which we add zero valued pixel around the image symmetrically.
#
# Below we show an example of zero padding.
# <figure>
# <img src="./convolution/padding.png"
# alt="">
# <figcaption style="text-align:center"><u> <i> Example of zero padding. In left we have original image and in right we
# have zero padded image where size of zero padding is one. </u> </i>
# </figcaption>
# </figure>
#
# In order to get output volume of same spatial dimension as input volume given the **stride=1**, we can find the size of zero padding needed with following formula.
# $$zero\,padding = \frac{(F-1)}{2}$$
# <center> Where: F is filter/kernel size</center>
#
# **Mutlifilter multichannel convolution**
#
# Further, we briefly discussed earlier that convolution operation can involve multiple filters. Below we show the example of convolution operation involving 2 filters in 3 channeled image.
#
# <figure>
# <img src="./convolution/standford_CNN.gif"
# alt="">
# <figcaption style="text-align:center">Convolution demo from <a href="http://cs231n.github.io/convolutional-networks/">standford</a>
# with K=2,F=3,S=2,P=1
# Where:
# <ul>
# <li>K = Number of filters,</li>
# <li>F= filter size ,</li>
# <li>S = stride length,</li>
# <li>P= amount of zero padding .</li>
# </ul>
# </figcaption>
# </figure>
#
# <br>
#
# In above animation we have a 7x7x3 image and image is zero padded once and we use a filter of size 3x3x3 ( notice that the depth of the convolution filter matches the depth of the image, both being 3). When the filter is at a particular location it covers a small volume of the input, and we perform the convolution operation. The only difference is this time we do the sum of matrix multiply (and in this case, we have added bias term also) in 3D instead of 2D, but the result is still a scalar.Since 3D volumes are hard to visualize, all the volumes (the input volume (in blue), the weight volumes (in red), the output volume (in green)) are visualized with each depth slice stacked in rows. We slide the filter over the input like above and perform the convolution at every location aggregating the result in a feature map. This feature map is of size 3x3x1, shown as the green slice on the right. Since we used 2 different filters we have 2 feature maps of size 3x3x1 and stacking them along the depth dimension would give us the final output of the convolution layer: a volume of size 3x3x2. Here, the output volume is visualized with each depth stacked in row.
#
# Another look into convolution operation is provided below.
# <figure>
# <img src="./convolution/cnn.gif"
# alt="">
# <figcaption style="text-align:center"> <u> <i> Convolution operation on a MxNx3 image matrix with a 3x3x3 Kernel.
# GIF from <a href="<a href="https://towardsdatascience.com/a-comprehensive-guide-to-convolutional-neural-networks-the-eli5-way-3bd2b1164a53">this blog post.</a></u> </i>
# </figure>
# <div class="alert alert-info">
# <b><u>To summarize, the Conv layer:</u></b>
# <ul>
# <li>It has following hyperparameters.</li>
# <ul>
# <li>Number of filters, K</li>
# <li>Stride length, S</li>
# <li>Amount of Zero padding, P</li>
# </ul>
# <li>It accepts the input volume of size: $W_{in}$x$H_{in}$x$D_{in}$.</li>
# <li>Output the volume of size: $W_{out}$x$H_{out}$x$D_{out}$</li>
# <br>
# where
# $W_{out}$x$H_{out}$x$D_{out}$ = $[(\frac{W_{in}+2P-F}{S} +1,\frac{H_{in}+2P-F}{S } +1,K]$
# <li> Convolution layer can be implemented using <b>tf.keras.layers.Conv2D</b> api using Keras in tensorflow 2. </li>
# </ul>
#
# </div>
#
# <div class="alert alert-info">
# The given link below visually shows what effect different Kernels have when applied over image. Try changing kernel and see the effect.
#
# <br>
#
# http://setosa.io/ev/image-kernels/
#
# </div>
#
#
# **Pooling Layer**
#
# Pooling layers reduces the spatial size of feature map. This enables to reduce the number of parameters in layers, helping to avoid overfitting. Pooling layers are also useful for extracting dominant features which are rotational and positional invariant, thus maintaining effective training of the model. Two most common pooling operations are
#
# - Max pooling and
# - average pooling
#
# Similar to convolution layer, in pooling layer also we slide the kernel of certain size over image. During this process, Max pooling takes the largest value from the section of the feature map overlapped by the kernel, while average pooling takes the average of all values in the window.
# Let's now visualize the max pooling operation.
# <figure>
# <img src="./convolution/m1.png"
# alt="">
# <figcaption style="text-align:center"> <u> <i>Max pooling step 1 with <b>kernel size of 2 and stride 1</b>. We take the largest value from the window of the feature map overlapped by the kernel.</u> </i>
# </figcaption>
# </figure>
# <br>
# <br>
#
# Similar to convolution operation, we now move kernel to 1 step right.
# <figure>
# <img src="./convolution/m2.png"
# alt="">
# <figcaption style="text-align:center"> <u> <i>Max pooling step 2 with <b>kernel size of 2 and stride 1</b>.</u> </i>
# </figcaption>
# </figure>
#
# <br>
# <br>
#
# Finally, we get complete max pooled feature map
# <figure>
# <img src="./convolution/m3.png"alt="">
# <figcaption style="text-align:center"> <u> <i>Max pooling final step with <b>kernel size of 2 and stride 1.</b></u> </i>
# </figcaption>
# </figure>
# <br>
# <br>
#
#
# <div class="alert alert-info">
# <b><u>To summarize the Pooling layer</u></b>:
# <ul>
# <li>It has following hyperparameters:</li>
# <ul>
# <li> Filter size,F</li>
# <li>Stride length, S</li>
# </ul>
# <li>It accepts the input volume of size: $W_{in}$x$H_{in}$x$D_{in}$.</li>
# <li>Output the volume of size: $W_{out}$x$H_{out}$x$D_{out}$</li>
#
# <br>
# where
# $W_{out}$x$H_{out}$x$D_{out}$ = $[(\frac{W_{in}-F}{S} +1,\frac{H_{in}-F}{S } +1,D_{in}]$
# <li>Max pooling can be implemented using <b>tf.keras.layers.MaxPool2D</b> api using Keras in tensorflow 2 </li>
# </ul>
# </div>
#
# <br>
# <br>
# <div class="alert alert-info">
# In CNN, max pooling is used much more often than average pooling. And two most common hyperparameter choices are
# F=3,S=2, and F=2,S=2 (later one being even more common)
# <a href='https://www.youtube.com/watch?v=8oOgPUO-TBY'>[1]</a>
# <a href='http://cs231n.github.io/convolutional-networks/'>[2]</a>.
# </div>
#
#
# **Fully-Connected layer**
#
# In this layer, feature map from last convolution or pooling layer is flattened into single vector of values and feed it into a fully connected layer. Fully connected layers are just like Artificial Neural Networks and perform the same mathematical operations. After passing through the fully connected layers, the final layer uses the softmax activation function which gives the probabilities of the input belonging to a particular class.
# <br>
# <br>
# <figure>
# <img src="./convolution/fully_connected.png"alt="">
# <figcaption style="text-align:center"> <u> <i>Fully connected network in CNN. <br> Feature map from final convolution or pooling layer is unrolled into column vector and feed into fully connected layers.</u> </i>
# </figcaption>
# </figure>
# ## Keras Implementation of CNN using tensorflow 2.0
# Now, we will build and train a Convolutional Neural Network using sequential API from Keras. Our network architecture following the sequence of following layers.
#
# Input → 2 * (Conv → Conv → Pool) → Flatten → Dense → Dense.
# define the model architecture
model = tf.keras.models.Sequential([
tf.keras.layers.Conv2D(filters=32, kernel_size=3, padding="same", activation="relu", input_shape=[28,28,1]),
tf.keras.layers.Conv2D(filters=32, kernel_size=3, padding="same", activation="relu"),
tf.keras.layers.MaxPool2D(pool_size=2),
tf.keras.layers.Conv2D(filters=64, kernel_size=3, padding="same", activation="relu"),
tf.keras.layers.Conv2D(filters=64, kernel_size=3, padding="same", activation="relu"),
tf.keras.layers.MaxPool2D(pool_size=2),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(128, activation="relu"),
tf.keras.layers.Dense(10, activation="softmax")
])
# <div class='alert alert-info'>
# We use <b>Conv2d</b> class method to define convolution layer. This method requires the numbers of parameters. The parameters we defined are:
#
# - filters: the numbers of filters the layers will learn.
# - Kernel size: integer specfifying the 2D convolution window size.
# - padding: types of padding to apply
# - activation: string specifying the activation function to apply after performing the convolution.
# <br>
#
# Similarly, we apply max pooling in our pooling layer by using <b>MaxPool2D</b> class method. And the parameter 'pool_size' determines the kernel size or filter size.
# Both Conv2d and and MaxPool2d have few other parameters. So, we request you to go through their documentation to understand them if you are interested.
# </div>
#
#
#
# **NOTE**
# <div class="alert alert-warning">
# In keras when creating a conv2d layer padding parameter is defined as:
# <li>padding = 'valid'. Meaning input volume is not zero-padded thus reducing the spatial dimension</li>
# <li>padding = 'same'. Meaning output volume size matches input volume size</li>
#
# </div>
# ### Calculating the number of parameters in CNN layer
# Deep learning model learns 100 of thousands of parameters (weights and bias). Knowing the total numbers of learnable parameters helps to determine the required sample size (number of training data points) to avoid overfitting.
# In CNN also we can calculate the learnable parameters in each layer by using following formula.
# <div class="alert alert-info">
# $$Number\,of\,params =({kernel\_ width }* kernel\_height * channels\_in + 1 (for\_bias)) * channels\_out$$
# </div>
#
#
#
# Using the above formula, let's calculate the numbers of parameters in the CNN we defined above.
#
#
# param_calculation| input|output|layer|param
# ---|---|----|----|---
# (3 * 3 * 1 + 1)* 32 |28x28x1 |28x28x32 | Conv2D |320
# (3 * 3 * 32 + 1) * 32 |28x28x32 |28x28x32 | Conv2D |9248
# 0 |28x28x32 |14x14x32 | MaxPool2D |0
# (3 * 3 * 32 + 1) * 64 |14x14x32 |14x14x64 | Conv2D |18494
# (3 * 3 * 64 + 1) * 64 |14x14x64 |14x14x64 |Conv2D |36928
# 0 |7x7x64 |7x7x64 | MaxPool2D |0
# 0 |7x7x64 |3136 |Flatten |0
# (3136 + 1) * 128 |3136 |128 |Dense |401536
# (128 + 1) * 10 |128 |10 | Dense |1290
#
# **NOTE**
# <div class="alert alert-warning">
# In CNN, pooling layers do not have any learnable parameter.
# </div>
#
# #### Comparing manually computed parameter number with the result from .summary() function
model.summary()
# <div class="alert alert-info">
# Comparing the above two result, we can see that out manual calculation and output from summary() function yields same results regarding the total number of parameters in CNN.
# </div>
# #### Plotting the graph of the model
# Lets plot the graph of the model
tf.keras.utils.plot_model(
model,
to_file='model.png',show_shapes=True, show_layer_names=True)
# <div class="alert alert-info">
# Did you notice at above graph that spatial volume of output is decreasing whereas the number of filters learned is increasing as we go deeper into the network? This is a common practice in designing CNN architectures. Regarding choosing the suitable numbers of filters, it is advised to chose the powers of 2 values and kernel size to be a odd integer value <a href= 'https://www.pyimagesearch.com/2018/12/31/keras-conv2d-and-convolutional-layers/'>[3]</a>.
#
# </div>
#
# Next, we'll configure the specifications for model training. We will train our model with the sparse_categorical_crossentropy, because it's a multiclass classification problem and our final activation is a Softmax. We will use the rmsprop optimizer with a learning rate of 0.001. During training, we will want to monitor classification accuracy.
#
# <div class="alert alert-info">
# NOTE: RMSprop optimization algorithm is preferable to stochastic gradient descent (SGD), because RMSprop automates learning-rate tuning for us. (Other optimizers, such as Adam and Adagrad, also automatically adapt the learning rate during training, and would work equally well here.)
# </div>
#
model.compile(loss="sparse_categorical_crossentropy",
optimizer=tf.keras.optimizers.RMSprop(lr=0.001),
metrics=["accuracy"])
# Let's train for 20 epochs -- this may take a few minutes to run.
#
# The Loss and Accuracy are a great indication of progress of training. It's making a guess as to the classification of the training data, and then measuring it against the known label, calculating the result. Accuracy is the portion of correct guesses.
history = model.fit(X_train, y_train, epochs=20,
# We pass validation data to
# monitor validation loss and accuracy
# at the end of each epoch
validation_data=(X_valid, y_valid))
#-----------------------------------------------------------
# Retrieve a list of list results on training and validation data
# sets for each training epoch
#-----------------------------------------------------------
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1,len(acc)+1) # Get number of epochs
#------------------------------------------------
# Plot training and validation accuracy per epoch
#------------------------------------------------
plt.plot(epochs, acc, label='Training accuracy')
plt.plot(epochs, val_acc, label='Validation accuracy')
plt.title('Training and validation accuracy')
plt.xticks(epochs)
plt.xlabel('epoch')
plt.ylabel('accuracy')
plt.legend();
#------------------------------------------------
# Plot training and validation loss per epoch
#------------------------------------------------
plt.plot(epochs, loss, label='Training Loss')
plt.plot(epochs, val_loss, label='Validation Loss')
plt.title('Training and validation loss')
plt.xticks(epochs)
plt.xlabel('epoch')
plt.ylabel('loss')
plt.legend();
# ### test set accuracy
# Now, let's check the accuracy of model in test set, which is done by calling 'evalute' method on model. Test the accuracy of model on test set. Use batch size of 128.
#
test_loss, test_accuracy = model.evaluate(test_images.reshape(-1,28,28,1), test_labels, batch_size=128)
print('Accuracy on test dataset:', test_accuracy)
# # Visualizing the activations in CNN
# In this section we will visualize the activations of a particular layer in a CNN. Visualizing the activation of individual layer helps to understand how the input is decomposed into some relevant pixel patterns within an image. Each filter encodes relatively independent features of input image.
# first let's display the image whose feature map we will visualize
plt.imshow(test_images[0]);
# +
# Now, lets extract the outputs from all layers:
layer_outputs = [layer.output for layer in model.layers]
# Create a model that will return these outputs, given the model input:
activation_model = tf.keras.models.Model(inputs = model.input, outputs = layer_outputs)
activation = activation_model.predict(test_images[0].reshape(1, 28, 28, 1))
# -
# let's take an activation of first convolution layer for our input.
first_layer_activation = activation[0]
print(first_layer_activation.shape)
# So, the feature map from first convolution layer is 28 x 28 in dimension with 32 channels. Now, lets visualize what kind of feature few channels have learned.
channels = [1,3] # let's select first and third channel.
for channel in channels:
plt.matshow(first_layer_activation[0, :, :, channel], cmap='viridis')
plt.show()
# We can see from above visualization of feature map from first
# layer is that initial layers in network learned the lower level features from image, in this case, various edges.
## Now, let's go deeper into the network and select the feature map
# from 5th layer
fifth_layer_activation = activation[4]
print(fifth_layer_activation .shape)
channels = [10, 20, 32,63] # select the channel to visualize
for channel in channels:
plt.matshow(fifth_layer_activation[0, :, :, channel], cmap='viridis')
plt.show()
# Above visualization of activations from deeper layers reveals that as we go higher-up in the network, the feature learned becomes less visually interpretable, meaning encoding the higher level feature of object.
#
# **student feedback - explain what the higher level feature of object mean ?**
# # Prediction on test and visualizing the result
#get the predictions for the test data
predicted_classes = model.predict_classes(test_images.reshape(-1,28,28,1))
#get true test_label
y_true=test_labels
# +
#to get the total correct and incorrect prediction from the predict class
correct=np.nonzero(predicted_classes==y_true)[0]
correct.shape[0]
incorrect=np.nonzero(predicted_classes!=y_true)[0]
print("Correct predicted classes:",correct.shape[0])
print("Incorrect predicted classes:",incorrect.shape[0])
# +
# let's map each images labels to item name
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
# -
target_names = ["Class {} ({}) :".format(i,class_names[i]) for i in range(10)]
print(classification_report(y_true, predicted_classes, target_names=target_names))
# +
def plot_images(data_index):
'''
function to plot correctly and incorrectly classified images.
Parameters:
data_index (int): index of either correctly or incorrectly classified images.
'''
# Plot the sample images now
f, ax = plt.subplots(3,3, figsize=(10,10))
for i, indx in enumerate(data_index[:9]):
ax[i//3, i%3].imshow(test_images[indx].reshape(28,28))
ax[i//3, i%3].axis('off')
ax[i//3, i%3].set_title("True:{} Pred:{}".format(class_names[test_labels[indx]],class_names[predicted_classes[indx]]))
plt.show()
plot_images(correct)
# -
plot_images(incorrect)
# +
# plot the predicted label and true label of image
def plot_image(i, true_labels, images):
''' function to plot prediction result along with class probability
arguments :
i : image index
true_labels : true class label of test set images
images : test set images
'''
img=(images[i])
img=np.array([img])
predictions_array=model.predict(img.reshape(1,28,28,1))
true_label=true_labels[i]
fig, (ax1,ax2) = plt.subplots(figsize=(6,9),ncols=2)
ax1.grid(False)
ax1.set_xticks([])
ax1.set_yticks([])
ax1.imshow(img.reshape(28,28), cmap=plt.cm.binary)
predicted_label = np.argmax(predictions_array)
if predicted_label == true_label:
color = 'blue'
else:
color = 'red'
ax1.set_xlabel("Predicted : {} {:2.0f}% \n True label:({})".format(class_names[predicted_label],
100*np.max(predictions_array),
class_names[true_label]),
color=color)
ax2.barh(np.arange(10), predictions_array.squeeze())
ax2.set_aspect(0.1)
ax2.set_yticks(np.arange(10))
ax2.set_yticklabels(class_names, size='small');
ax2.set_title('Class Probability')
ax2.set_xlim(0, 1.1)
plt.tight_layout()
# -
plot_image(1123,test_labels,test_images)
# ## Refrences
# 1. https://www.youtube.com/watch?v=XTzDMvMXuAk
# 2. http://cs231n.github.io/convolutional-networks/
# 3. https://www.pyimagesearch.com/2018/12/31/keras-conv2d-and-convolutional-layers/
# **Additional material**
# - [Convolutional explained visually](http://setosa.io/ev/image-kernels/)
# - [Convolutional Neural Networks (CNNs / ConvNets), Standford](http://cs231n.github.io/convolutional-networks/)
# - [A Beginner's Guide To Understanding Convolutional Neural Networks,<NAME>](https://adeshpande3.github.io/adeshpande3.github.io/A-Beginner's-Guide-To-Understanding-Convolutional-Neural-Networks/)
| ANN_Demo/.ipynb_checkpoints/Round2_CNN-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="qD2vxNk3a5mX" colab_type="code" outputId="5ae300a6-2a79-4daf-91a6-46ec739ccf6a" colab={"base_uri": "https://localhost:8080/", "height": 34}
'''
'''
# + id="eT4koJpFDTwf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 122} outputId="b981fbc0-8f1d-4c81-8cdf-941767711802"
from google.colab import drive
drive.mount('/content/gdrive')
# + id="PziA1cW1XMBx" colab_type="code" outputId="1b1671b5-2f59-47f9-a1c0-c47c86feb6ff" colab={"base_uri": "https://localhost:8080/", "height": 306}
# !nvidia-smi
# + id="jYkHHJALobDu" colab_type="code" outputId="f3a57d5a-7e60-46fc-add3-31769e08aa7c" colab={"base_uri": "https://localhost:8080/", "height": 34}
# %tensorflow_version 1.x
import tensorflow as tf
# + id="C8xL9ualobVq" colab_type="code" outputId="c4406035-db9d-44aa-ce24-23d08d9a150a" colab={"base_uri": "https://localhost:8080/", "height": 173}
# !pip install bert-tensorflow
import bert
from bert import run_classifier
from bert import optimization
from bert import tokenization
# + id="wdIu6lnmpCK9" colab_type="code" outputId="c498393c-6291-4a48-b318-8c3843150b33" colab={"base_uri": "https://localhost:8080/", "height": 204}
# !wget https://storage.googleapis.com/bert_models/2018_11_23/multi_cased_L-12_H-768_A-12.zip
# + id="fA_D5h70vT23" colab_type="code" outputId="1910a70c-528c-4314-d56c-51cab7401de3" colab={"base_uri": "https://localhost:8080/", "height": 34}
# !ls
# + id="0RmhjbtWqP7l" colab_type="code" outputId="2fc82e37-c8fc-4949-934c-cf9b16cebf6a" colab={"base_uri": "https://localhost:8080/", "height": 136}
# !unzip multi_cased_L-12_H-768_A-12.zip
# + id="BMGsSaNCqWKG" colab_type="code" outputId="b7284a55-8f80-4d72-c77e-f519b262d2bc" colab={"base_uri": "https://localhost:8080/", "height": 51}
# !ls
# + id="6YjL7tWva7KB" colab_type="code" colab={}
import pandas as pd
file = '/content/gdrive/My Drive/multilingual grounded/final.csv'
df = pd.read_csv(file)
# + id="qZ5_sQwDaKFM" colab_type="code" colab={}
df = df.head(500)
# + id="JnEibMrF-0-_" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
train, test = train_test_split(df, test_size=0.1,random_state = 42,shuffle = True)
# + id="mPOe9tf9a7M-" colab_type="code" colab={}
def get_data_eng_hindi(a):
b_ = list(a['gold_label'])
lab = []
"""
lab = []
for i in b_:
lab.append(i-1)
"""
for i in b_:
if i=='contradiction':
lab.append(0)
elif i=='neutral':
lab.append(1)
elif i== 'entailment':
lab.append(2)
else:
lab.append(3)
sentence_1 = list(a['premise'])
sentence_2 = list(a['hypo_hindi'])
raw_data_train = {'sentence1_eng': sentence_1,
'sentence2_hindi': sentence_2,
'label': lab}
df = pd.DataFrame(raw_data_train, columns = ['sentence1_eng','sentence2_hindi','label'])
return df
# + id="OXPP1J0Ya7QJ" colab_type="code" colab={}
def get_data_hindi_eng(a):
b_ = list(a['gold_label'])
lab = []
"""
lab = []
for i in b_:
lab.append(i-1)
"""
for i in b_:
if i=='contradiction':
lab.append(0)
elif i=='neutral':
lab.append(1)
elif i== 'entailment':
lab.append(2)
else:
lab.append(3)
sentence_1 = list(a['premise_hindi'])
sentence_2 = list(a['hypo'])
raw_data_train = {'sentence1_hindi': sentence_1,
'sentence2_eng': sentence_2,
'label': lab}
df = pd.DataFrame(raw_data_train, columns = ['sentence1_hindi','sentence2_eng','label'])
return df
# + id="2ihgc-mMa7Vc" colab_type="code" colab={}
train_eng_hindi = get_data_eng_hindi(train)
train_hindi_eng = get_data_hindi_eng(train)
# + id="6YrgZRZM_Bwk" colab_type="code" colab={}
test_eng_hindi = get_data_eng_hindi(test)
test_hindi_eng = get_data_hindi_eng(test)
# + id="dUujUqjsqgxz" colab_type="code" outputId="3f909b4b-7a54-40f1-e721-67e4973aba9f" colab={"base_uri": "https://localhost:8080/", "height": 359}
train_eng_hindi[0:10]
# + id="oNgE0azkwvH0" colab_type="code" outputId="4b6d3a53-944b-4f10-fdca-67002c00a0fe" colab={"base_uri": "https://localhost:8080/", "height": 359}
train_hindi_eng[0:10]
# + id="Fm4CAUdkwI5E" colab_type="code" outputId="5394397e-9dfd-47fb-d9da-26b0d61ac192" colab={"base_uri": "https://localhost:8080/", "height": 359}
test_eng_hindi[0:10]
# + id="l0yZjAVuABTY" colab_type="code" outputId="3dd376ee-cf14-42fe-d224-09a229ca9154" colab={"base_uri": "https://localhost:8080/", "height": 359}
test_hindi_eng[0:10]
# + id="9g9gnIn5dg9X" colab_type="code" colab={}
label_list = [0,1,2,3]
# + id="H5AOE0Kka7YR" colab_type="code" colab={}
train_InputExamples_eng = train_eng_hindi.apply(lambda x: bert.run_classifier.InputExample(guid=None, # Globally unique ID for bookkeeping, unused in this example
text_a = x['sentence1_eng'],
text_b = x['sentence2_hindi'],
label = x['label']), axis = 1)
train_InputExamples_hindi = train_hindi_eng.apply(lambda x: bert.run_classifier.InputExample(guid=None, # Globally unique ID for bookkeeping, unused in this example
text_a = x['sentence1_hindi'],
text_b = x['sentence2_eng'],
label = x['label']), axis = 1)
# + id="2cbAnc9BAFZ0" colab_type="code" colab={}
test_InputExamples_eng = test_eng_hindi.apply(lambda x: bert.run_classifier.InputExample(guid=None, # Globally unique ID for bookkeeping, unused in this example
text_a = x['sentence1_eng'],
text_b = x['sentence2_hindi'],
label = x['label']), axis = 1)
test_InputExamples_hindi = test_hindi_eng.apply(lambda x: bert.run_classifier.InputExample(guid=None, # Globally unique ID for bookkeeping, unused in this example
text_a = x['sentence1_hindi'],
text_b = x['sentence2_eng'],
label = x['label']), axis = 1)
# + id="dV0e5pV7a7hw" colab_type="code" outputId="5d7f2ea9-ab2e-4298-da2d-36812c09f1db" colab={"base_uri": "https://localhost:8080/", "height": 71}
vocab_file = "multi_cased_L-12_H-768_A-12/vocab.txt"
def create_tokenizer_from_hub_module():
return bert.tokenization.FullTokenizer(
vocab_file=vocab_file, do_lower_case=True)
tokenizer = create_tokenizer_from_hub_module()
# + id="GUZbgivOqqFe" colab_type="code" outputId="f4325b8a-0915-4523-8766-8b9dba53ef05" colab={"base_uri": "https://localhost:8080/", "height": 34}
tokenizer.tokenize("how are you")
# + id="rekQnmuRw-Cw" colab_type="code" outputId="6db1e687-39fb-41b7-c2c3-03f702fe2e80" colab={"base_uri": "https://localhost:8080/", "height": 306}
tokenizer.tokenize("एक आदमी गोरा सिर वाली महिला से बात कर रहा है।")
# + id="oPXlGCfZa7fH" colab_type="code" outputId="e295c98c-8aad-4ae4-f7d4-3b18ddc9223d" colab={"base_uri": "https://localhost:8080/", "height": 1000}
MAX_SEQ_LENGTH = 128
# Convert our train and test features to InputFeatures that BERT understands.
train_features_eng = bert.run_classifier.convert_examples_to_features(train_InputExamples_eng, label_list, MAX_SEQ_LENGTH, tokenizer)
train_features_hindi = bert.run_classifier.convert_examples_to_features(train_InputExamples_hindi, label_list, MAX_SEQ_LENGTH, tokenizer)
# + id="zjU-gih6AjFc" colab_type="code" outputId="9432fb56-54f8-4481-a73c-f0ab63d47653" colab={"base_uri": "https://localhost:8080/", "height": 1000}
MAX_SEQ_LENGTH = 128
# Convert our train and test features to InputFeatures that BERT understands.
test_features_eng = bert.run_classifier.convert_examples_to_features(test_InputExamples_eng, label_list, MAX_SEQ_LENGTH, tokenizer)
test_features_hindi = bert.run_classifier.convert_examples_to_features(test_InputExamples_hindi, label_list, MAX_SEQ_LENGTH, tokenizer)
# + id="hkXg5ovaa7ca" colab_type="code" colab={}
def create_model(bert_config, is_training, input_ids, input_mask, segment_ids,
labels, num_labels, use_one_hot_embeddings):
"""Creates a classification model."""
model = bert.run_classifier.modeling.BertModel(
config=bert_config,
is_training=is_training,
input_ids=input_ids,
input_mask=input_mask,
token_type_ids=segment_ids,
use_one_hot_embeddings=use_one_hot_embeddings)
# In the demo, we are doing a simple classification task on the entire
# segment.
#
# If you want to use the token-level output, use model.get_sequence_output()
# instead.
output_layer = model.get_pooled_output()
hidden_size = output_layer.shape[-1].value
output_weights = tf.get_variable(
"output_weights", [num_labels, hidden_size],
initializer=tf.truncated_normal_initializer(stddev=0.02))
output_bias = tf.get_variable(
"output_bias", [num_labels], initializer=tf.zeros_initializer())
with tf.variable_scope("loss"):
if is_training:
# I.e., 0.1 dropout
output_layer = tf.nn.dropout(output_layer, keep_prob=0.9)
logits = tf.matmul(output_layer, output_weights, transpose_b=True)
logits = tf.nn.bias_add(logits, output_bias)
probabilities = tf.nn.softmax(logits, axis=-1)
log_probs = tf.nn.log_softmax(logits, axis=-1)
predicted_labels = tf.squeeze(tf.argmax(log_probs, axis=-1, output_type=tf.int32))
one_hot_labels = tf.one_hot(labels, depth=num_labels, dtype=tf.float32)
per_example_loss = -tf.reduce_sum(one_hot_labels * log_probs, axis=-1)
loss = tf.reduce_mean(per_example_loss)
return (loss, per_example_loss, logits, probabilities,predicted_labels,output_layer)
# + id="Qg6sR0Ar31NO" colab_type="code" colab={}
import numpy as np
# + id="uvZyC3BFeSA1" colab_type="code" colab={}
def model_fn_builder(bert_config, num_labels, init_checkpoint, learning_rate,
num_train_steps, num_warmup_steps, use_tpu,
use_one_hot_embeddings):
"""Returns `model_fn` closure for TPUEstimator."""
def model_fn(features, labels, mode, params): # pylint: disable=unused-argument
"""The `model_fn` for TPUEstimator."""
tf.logging.info("*** Features ***")
for name in sorted(features.keys()):
tf.logging.info(" name = %s, shape = %s" % (name, features[name].shape))
input_ids = features["input_ids"]
input_mask = features["input_mask"]
segment_ids = features["segment_ids"]
label_ids = features["label_ids"]
is_real_example = None
if "is_real_example" in features:
is_real_example = tf.cast(features["is_real_example"], dtype=tf.float32)
else:
is_real_example = tf.ones(tf.shape(label_ids), dtype=tf.float32)
is_training = (mode == tf.estimator.ModeKeys.TRAIN)
(total_loss, per_example_loss, logits, probabilities,predicted_labels,hidden_context) = create_model(
bert_config, is_training, input_ids, input_mask, segment_ids, label_ids,
num_labels, use_one_hot_embeddings)
tvars = tf.trainable_variables()
initialized_variable_names = {}
scaffold_fn = None
if init_checkpoint:
(assignment_map, initialized_variable_names
) = bert.run_classifier.modeling.get_assignment_map_from_checkpoint(tvars, init_checkpoint)
if use_tpu:
def tpu_scaffold():
tf.train.init_from_checkpoint(init_checkpoint, assignment_map)
return tf.train.Scaffold()
scaffold_fn = tpu_scaffold
else:
tf.train.init_from_checkpoint(init_checkpoint, assignment_map)
"""
tf.logging.info("**** Trainable Variables ****")
for var in tvars:
init_string = ""
if var.name in initialized_variable_names:
init_string = ", *INIT_FROM_CKPT*"
tf.logging.info(" name = %s, shape = %s%s", var.name, var.shape,
init_string)
"""
output_spec = None
if mode == tf.estimator.ModeKeys.TRAIN:
train_op = optimization.create_optimizer(
total_loss, learning_rate, num_train_steps, num_warmup_steps, use_tpu)
output_spec = tf.estimator.EstimatorSpec(
mode=mode,
loss=total_loss,
train_op=train_op)
elif mode == tf.estimator.ModeKeys.EVAL:
def metric_fn(per_example_loss, label_ids, logits, is_real_example):
predictions = tf.argmax(logits, axis=-1, output_type=tf.int32)
accuracy = tf.metrics.accuracy(
labels=label_ids, predictions=predictions, weights=is_real_example)
loss = tf.metrics.mean(values=per_example_loss, weights=is_real_example)
return {
"eval_accuracy": accuracy,
"eval_loss": loss
}
eval_metrics = metric_fn(per_example_loss, label_ids, logits, is_real_example)
output_spec = tf.estimator.EstimatorSpec(
mode=mode,
loss=total_loss,
eval_metric_ops=eval_metrics)
else:
output_spec = tf.estimator.EstimatorSpec(
mode=mode,
predictions={"probabilities": probabilities,"labels": predicted_labels, "hidden_context": hidden_context})
return output_spec
return model_fn
# + id="XPQALfA9CFbW" colab_type="code" colab={}
def create_model_img(img_features,bert_config, is_training, input_ids, input_mask, segment_ids,
labels, num_labels, use_one_hot_embeddings):
"""Creates a classification model."""
model = bert.run_classifier.modeling.BertModel(
config=bert_config,
is_training=is_training,
input_ids=input_ids,
input_mask=input_mask,
token_type_ids=segment_ids,
use_one_hot_embeddings=use_one_hot_embeddings)
# In the demo, we are doing a simple classification task on the entire
# segment.
#
# If you want to use the token-level output, use model.get_sequence_output()
# instead.
output_layer = model.get_pooled_output()
hidden_size = output_layer.shape[-1].value
old_size = img_features.shape[-1].value
output_weights = tf.get_variable(
"output_weights", [num_labels, hidden_size],
initializer=tf.truncated_normal_initializer(stddev=0.02))
output_bias = tf.get_variable(
"output_bias", [num_labels], initializer=tf.zeros_initializer())
output_weights_img = tf.get_variable(
"output_weights_img", [hidden_size,old_size],
initializer=tf.truncated_normal_initializer(stddev=0.02))
output_bias_img = tf.get_variable(
"output_bias_img", [hidden_size], initializer=tf.zeros_initializer())
with tf.variable_scope("loss"):
if is_training:
# I.e., 0.1 dropout
output_layer = tf.nn.dropout(output_layer, keep_prob=0.9)
img_features = tf.matmul(img_features, output_weights_img, transpose_b=True)
img_features = tf.nn.bias_add(img_features, output_bias_img)
img_features = tf.nn.relu(img_features)
output_layer = tf.math.multiply(output_layer,img_features)
logits = tf.matmul(output_layer, output_weights, transpose_b=True)
logits = tf.nn.bias_add(logits, output_bias)
probabilities = tf.nn.softmax(logits, axis=-1)
log_probs = tf.nn.log_softmax(logits, axis=-1)
predicted_labels = tf.squeeze(tf.argmax(log_probs, axis=-1, output_type=tf.int32))
one_hot_labels = tf.one_hot(labels, depth=num_labels, dtype=tf.float32)
per_example_loss = -tf.reduce_sum(one_hot_labels * log_probs, axis=-1)
loss = tf.reduce_mean(per_example_loss)
return (loss, per_example_loss, logits, probabilities,predicted_labels,output_layer)
# + id="AmjDdjWoCFi3" colab_type="code" colab={}
def model_fn_builder_img(bert_config, num_labels, init_checkpoint, learning_rate,
num_train_steps, num_warmup_steps, use_tpu,
use_one_hot_embeddings):
"""Returns `model_fn` closure for TPUEstimator."""
def model_fn(features, labels, mode, params): # pylint: disable=unused-argument
"""The `model_fn` for TPUEstimator."""
tf.logging.info("*** Features ***")
for name in sorted(features.keys()):
tf.logging.info(" name = %s, shape = %s" % (name, features[name].shape))
input_ids = features["input_ids"]
input_mask = features["input_mask"]
segment_ids = features["segment_ids"]
label_ids = features["label_ids"]
img_features = features["img_features"]
is_real_example = None
if "is_real_example" in features:
is_real_example = tf.cast(features["is_real_example"], dtype=tf.float32)
else:
is_real_example = tf.ones(tf.shape(label_ids), dtype=tf.float32)
is_training = (mode == tf.estimator.ModeKeys.TRAIN)
(total_loss, per_example_loss, logits, probabilities,predicted_labels,hidden_context) = create_model_img(
img_features, bert_config, is_training, input_ids, input_mask, segment_ids, label_ids,
num_labels, use_one_hot_embeddings)
tvars = tf.trainable_variables()
initialized_variable_names = {}
scaffold_fn = None
if init_checkpoint:
(assignment_map, initialized_variable_names
) = bert.run_classifier.modeling.get_assignment_map_from_checkpoint(tvars, init_checkpoint)
if use_tpu:
def tpu_scaffold():
tf.train.init_from_checkpoint(init_checkpoint, assignment_map)
return tf.train.Scaffold()
scaffold_fn = tpu_scaffold
else:
tf.train.init_from_checkpoint(init_checkpoint, assignment_map)
"""
tf.logging.info("**** Trainable Variables ****")
for var in tvars:
init_string = ""
if var.name in initialized_variable_names:
init_string = ", *INIT_FROM_CKPT*"
tf.logging.info(" name = %s, shape = %s%s", var.name, var.shape,
init_string)
"""
output_spec = None
if mode == tf.estimator.ModeKeys.TRAIN:
train_op = optimization.create_optimizer(
total_loss, learning_rate, num_train_steps, num_warmup_steps, use_tpu)
output_spec = tf.estimator.EstimatorSpec(
mode=mode,
loss=total_loss,
train_op=train_op)
elif mode == tf.estimator.ModeKeys.EVAL:
def metric_fn(per_example_loss, label_ids, logits, is_real_example):
predictions = tf.argmax(logits, axis=-1, output_type=tf.int32)
accuracy = tf.metrics.accuracy(
labels=label_ids, predictions=predictions, weights=is_real_example)
loss = tf.metrics.mean(values=per_example_loss, weights=is_real_example)
return {
"eval_accuracy": accuracy,
"eval_loss": loss
}
eval_metrics = metric_fn(per_example_loss, label_ids, logits, is_real_example)
output_spec = tf.estimator.EstimatorSpec(
mode=mode,
loss=total_loss,
eval_metric_ops=eval_metrics)
else:
output_spec = tf.estimator.EstimatorSpec(
mode=mode,
predictions={"probabilities": probabilities,"labels": predicted_labels, "hidden_context": hidden_context})
return output_spec
return model_fn
# + id="jD-zW6FwfOV7" colab_type="code" colab={}
def create_model_progressive(bert_config, is_training, input_ids, input_mask, segment_ids,
labels, num_labels, use_one_hot_embeddings,hidden_context):
"""Creates a classification model."""
model = bert.run_classifier.modeling.BertModel(
config=bert_config,
is_training=is_training,
input_ids=input_ids,
input_mask=input_mask,
token_type_ids=segment_ids,
use_one_hot_embeddings=use_one_hot_embeddings)
# In the demo, we are doing a simple classification task on the entire
# segment.
#
# If you want to use the token-level output, use model.get_sequence_output()
# instead.
output_layer = model.get_pooled_output()
hidden_size = output_layer.shape[-1].value
output_weights = tf.get_variable(
"output_weights", [num_labels, hidden_size],
initializer=tf.truncated_normal_initializer(stddev=0.02))
output_bias = tf.get_variable(
"output_bias", [num_labels], initializer=tf.zeros_initializer())
with tf.variable_scope("loss"):
if is_training:
# I.e., 0.1 dropout
output_layer = tf.nn.dropout(output_layer, keep_prob=0.9)
output_layer_probs = tf.nn.softmax(output_layer,axis = -1)
#loss = y_true * log(y_true / y_pred)
hidden_context = tf.nn.softmax(hidden_context,axis = -1)
per_example_kd_loss = tf.keras.losses.KLD(hidden_context,output_layer_probs)
logits = tf.matmul(output_layer, output_weights, transpose_b=True)
logits = tf.nn.bias_add(logits, output_bias)
probabilities = tf.nn.softmax(logits, axis=-1)
log_probs = tf.nn.log_softmax(logits, axis=-1)
predicted_labels = tf.squeeze(tf.argmax(log_probs, axis=-1, output_type=tf.int32))
one_hot_labels = tf.one_hot(labels, depth=num_labels, dtype=tf.float32)
per_example_loss = -tf.reduce_sum(one_hot_labels * log_probs, axis=-1)
kd_loss_weight = 0.2 #hyperparameter
per_example_kd_loss = kd_loss_weight*per_example_kd_loss
per_example_loss += per_example_kd_loss
loss = tf.reduce_mean(per_example_loss)
return (loss, per_example_loss, logits, probabilities,predicted_labels)
# + id="0_JJrlOgfOaw" colab_type="code" colab={}
def model_fn_builder_progressive(bert_config, num_labels, init_checkpoint, learning_rate,
num_train_steps, num_warmup_steps, use_tpu,
use_one_hot_embeddings):
"""Returns `model_fn` closure for TPUEstimator."""
def model_fn(features, labels, mode, params): # pylint: disable=unused-argument
"""The `model_fn` for TPUEstimator."""
tf.logging.info("*** Features ***")
for name in sorted(features.keys()):
tf.logging.info(" name = %s, shape = %s" % (name, features[name].shape))
input_ids = features["input_ids"]
input_mask = features["input_mask"]
segment_ids = features["segment_ids"]
label_ids = features["label_ids"]
hidden_context = features["hidden_context"]
is_real_example = None
if "is_real_example" in features:
is_real_example = tf.cast(features["is_real_example"], dtype=tf.float32)
else:
is_real_example = tf.ones(tf.shape(label_ids), dtype=tf.float32)
is_training = (mode == tf.estimator.ModeKeys.TRAIN)
(total_loss, per_example_loss, logits, probabilities,predicted_labels) = create_model_progressive(
bert_config, is_training, input_ids, input_mask, segment_ids, label_ids,
num_labels, use_one_hot_embeddings,hidden_context)
tvars = tf.trainable_variables()
initialized_variable_names = {}
scaffold_fn = None
if init_checkpoint:
(assignment_map, initialized_variable_names
) = bert.run_classifier.modeling.get_assignment_map_from_checkpoint(tvars, init_checkpoint)
if use_tpu:
def tpu_scaffold():
tf.train.init_from_checkpoint(init_checkpoint, assignment_map)
return tf.train.Scaffold()
scaffold_fn = tpu_scaffold
else:
tf.train.init_from_checkpoint(init_checkpoint, assignment_map)
"""
tf.logging.info("**** Trainable Variables ****")
for var in tvars:
init_string = ""
if var.name in initialized_variable_names:
init_string = ", *INIT_FROM_CKPT*"
tf.logging.info(" name = %s, shape = %s%s", var.name, var.shape,
init_string)
"""
output_spec = None
if mode == tf.estimator.ModeKeys.TRAIN:
train_op = optimization.create_optimizer(
total_loss, learning_rate, num_train_steps, num_warmup_steps, use_tpu)
output_spec = tf.estimator.EstimatorSpec(
mode=mode,
loss=total_loss,
train_op=train_op)
elif mode == tf.estimator.ModeKeys.EVAL:
def metric_fn(per_example_loss, label_ids, logits, is_real_example):
predictions = tf.argmax(logits, axis=-1, output_type=tf.int32)
accuracy = tf.metrics.accuracy(
labels=label_ids, predictions=predictions, weights=is_real_example)
loss = tf.metrics.mean(values=per_example_loss, weights=is_real_example)
return {
"eval_accuracy": accuracy,
"eval_loss": loss,
}
eval_metrics = metric_fn(per_example_loss, label_ids, logits, is_real_example)
output_spec = tf.estimator.EstimatorSpec(
mode=mode,
loss=total_loss,
eval_metric_ops=eval_metrics)
else:
output_spec = tf.estimator.EstimatorSpec(
mode=mode,
predictions={"probabilities": probabilities,"labels": predicted_labels})
return output_spec
return model_fn
# + id="5dbRv0rEEacS" colab_type="code" colab={}
def create_model_progressive_img(img_features,bert_config, is_training, input_ids, input_mask, segment_ids,
labels, num_labels, use_one_hot_embeddings,hidden_context):
"""Creates a classification model."""
model = bert.run_classifier.modeling.BertModel(
config=bert_config,
is_training=is_training,
input_ids=input_ids,
input_mask=input_mask,
token_type_ids=segment_ids,
use_one_hot_embeddings=use_one_hot_embeddings)
# In the demo, we are doing a simple classification task on the entire
# segment.
#
# If you want to use the token-level output, use model.get_sequence_output()
# instead.
output_layer = model.get_pooled_output()
hidden_size = output_layer.shape[-1].value
old_size = img_features.shape[-1].value
output_weights = tf.get_variable(
"output_weights", [num_labels, hidden_size],
initializer=tf.truncated_normal_initializer(stddev=0.02))
output_bias = tf.get_variable(
"output_bias", [num_labels], initializer=tf.zeros_initializer())
output_weights_img = tf.get_variable(
"output_weights_img", [hidden_size,old_size],
initializer=tf.truncated_normal_initializer(stddev=0.02))
output_bias_img = tf.get_variable(
"output_bias_img", [hidden_size], initializer=tf.zeros_initializer())
with tf.variable_scope("loss"):
if is_training:
# I.e., 0.1 dropout
output_layer = tf.nn.dropout(output_layer, keep_prob=0.9)
img_features = tf.matmul(img_features, output_weights_img, transpose_b=True)
img_features = tf.nn.bias_add(img_features, output_bias_img)
img_features = tf.nn.relu(img_features)
print('shape of img features {}'.format(np.shape(img_features)))
output_layer = tf.math.multiply(img_features,output_layer)
output_layer_probs = tf.nn.softmax(output_layer,axis = -1)
#loss = y_true * log(y_true / y_pred)
hidden_context = tf.nn.softmax(hidden_context,axis = -1)
per_example_kd_loss = tf.keras.losses.KLD(hidden_context,output_layer_probs)
logits = tf.matmul(output_layer, output_weights, transpose_b=True)
logits = tf.nn.bias_add(logits, output_bias)
probabilities = tf.nn.softmax(logits, axis=-1)
log_probs = tf.nn.log_softmax(logits, axis=-1)
predicted_labels = tf.squeeze(tf.argmax(log_probs, axis=-1, output_type=tf.int32))
one_hot_labels = tf.one_hot(labels, depth=num_labels, dtype=tf.float32)
per_example_loss = -tf.reduce_sum(one_hot_labels * log_probs, axis=-1)
kd_loss_weight = 0.2 #hyperparameter
per_example_kd_loss = kd_loss_weight*per_example_kd_loss
per_example_loss += per_example_kd_loss
loss = tf.reduce_mean(per_example_loss)
return (loss, per_example_loss, logits, probabilities,predicted_labels)
# + id="YAyXjSYBEaYr" colab_type="code" colab={}
def model_fn_builder_img_progressive(bert_config, num_labels, init_checkpoint, learning_rate,
num_train_steps, num_warmup_steps, use_tpu,
use_one_hot_embeddings):
"""Returns `model_fn` closure for TPUEstimator."""
def model_fn(features, labels, mode, params): # pylint: disable=unused-argument
"""The `model_fn` for TPUEstimator."""
tf.logging.info("*** Features ***")
for name in sorted(features.keys()):
tf.logging.info(" name = %s, shape = %s" % (name, features[name].shape))
input_ids = features["input_ids"]
input_mask = features["input_mask"]
segment_ids = features["segment_ids"]
label_ids = features["label_ids"]
hidden_context = features["hidden_context"]
img_features = features["img_features"]
is_real_example = None
if "is_real_example" in features:
is_real_example = tf.cast(features["is_real_example"], dtype=tf.float32)
else:
is_real_example = tf.ones(tf.shape(label_ids), dtype=tf.float32)
is_training = (mode == tf.estimator.ModeKeys.TRAIN)
(total_loss, per_example_loss, logits, probabilities,predicted_labels) = create_model_progressive_img(
img_features,bert_config, is_training, input_ids, input_mask, segment_ids, label_ids,
num_labels, use_one_hot_embeddings,hidden_context)
tvars = tf.trainable_variables()
initialized_variable_names = {}
scaffold_fn = None
if init_checkpoint:
(assignment_map, initialized_variable_names
) = bert.run_classifier.modeling.get_assignment_map_from_checkpoint(tvars, init_checkpoint)
if use_tpu:
def tpu_scaffold():
tf.train.init_from_checkpoint(init_checkpoint, assignment_map)
return tf.train.Scaffold()
scaffold_fn = tpu_scaffold
else:
tf.train.init_from_checkpoint(init_checkpoint, assignment_map)
"""
tf.logging.info("**** Trainable Variables ****")
for var in tvars:
init_string = ""
if var.name in initialized_variable_names:
init_string = ", *INIT_FROM_CKPT*"
tf.logging.info(" name = %s, shape = %s%s", var.name, var.shape,
init_string)
"""
output_spec = None
if mode == tf.estimator.ModeKeys.TRAIN:
train_op = optimization.create_optimizer(
total_loss, learning_rate, num_train_steps, num_warmup_steps, use_tpu)
output_spec = tf.estimator.EstimatorSpec(
mode=mode,
loss=total_loss,
train_op=train_op)
elif mode == tf.estimator.ModeKeys.EVAL:
def metric_fn(per_example_loss, label_ids, logits, is_real_example):
predictions = tf.argmax(logits, axis=-1, output_type=tf.int32)
accuracy = tf.metrics.accuracy(
labels=label_ids, predictions=predictions, weights=is_real_example)
loss = tf.metrics.mean(values=per_example_loss, weights=is_real_example)
return {
"eval_accuracy": accuracy,
"eval_loss": loss,
}
eval_metrics = metric_fn(per_example_loss, label_ids, logits, is_real_example)
output_spec = tf.estimator.EstimatorSpec(
mode=mode,
loss=total_loss,
eval_metric_ops=eval_metrics)
else:
output_spec = tf.estimator.EstimatorSpec(
mode=mode,
predictions={"probabilities": probabilities,"labels": predicted_labels})
return output_spec
return model_fn
# + id="CqDUOvGGFNOO" colab_type="code" colab={}
def input_fn_builder(features, hidden_context,seq_length, is_training, drop_remainder):
"""Creates an `input_fn` closure to be passed to TPUEstimator."""
all_input_ids = []
all_input_mask = []
all_segment_ids = []
all_label_ids = []
for feature in features:
all_input_ids.append(feature.input_ids)
all_input_mask.append(feature.input_mask)
all_segment_ids.append(feature.segment_ids)
all_label_ids.append(feature.label_id)
def input_fn(params):
"""The actual input function."""
batch_size = params["batch_size"]
num_examples = len(features)
hidden_shape = hidden_context.shape[-1]
# This is for demo purposes and does NOT scale to large data sets. We do
# not use Dataset.from_generator() because that uses tf.py_func which is
# not TPU compatible. The right way to load data is with TFRecordReader.
d = tf.data.Dataset.from_tensor_slices({
"input_ids":
tf.constant(
all_input_ids, shape=[num_examples, seq_length],
dtype=tf.int32),
"input_mask":
tf.constant(
all_input_mask,
shape=[num_examples, seq_length],
dtype=tf.int32),
"segment_ids":
tf.constant(
all_segment_ids,
shape=[num_examples, seq_length],
dtype=tf.int32),
"label_ids":
tf.constant(all_label_ids, shape=[num_examples], dtype=tf.int32),
"hidden_context":
tf.constant(hidden_context, shape = [num_examples,hidden_shape], dtype = tf.float32),
})
if is_training:
d = d.repeat()
d = d.shuffle(buffer_size=100)
d = d.batch(batch_size=batch_size, drop_remainder=drop_remainder)
return d
return input_fn
# + id="BQTfzZYw_qHr" colab_type="code" colab={}
def input_fn_builder_img(img_features,features,seq_length, is_training, drop_remainder):
"""Creates an `input_fn` closure to be passed to TPUEstimator."""
all_input_ids = []
all_input_mask = []
all_segment_ids = []
all_label_ids = []
for feature in features:
all_input_ids.append(feature.input_ids)
all_input_mask.append(feature.input_mask)
all_segment_ids.append(feature.segment_ids)
all_label_ids.append(feature.label_id)
def input_fn(params):
"""The actual input function."""
batch_size = params["batch_size"]
num_examples = len(features)
hidden_shape_img = img_features.shape[-1]
# This is for demo purposes and does NOT scale to large data sets. We do
# not use Dataset.from_generator() because that uses tf.py_func which is
# not TPU compatible. The right way to load data is with TFRecordReader.
d = tf.data.Dataset.from_tensor_slices({
"input_ids":
tf.constant(
all_input_ids, shape=[num_examples, seq_length],
dtype=tf.int32),
"input_mask":
tf.constant(
all_input_mask,
shape=[num_examples, seq_length],
dtype=tf.int32),
"segment_ids":
tf.constant(
all_segment_ids,
shape=[num_examples, seq_length],
dtype=tf.int32),
"label_ids":
tf.constant(all_label_ids, shape=[num_examples], dtype=tf.int32),
"img_features":
tf.constant(img_features, shape = [num_examples,hidden_shape_img], dtype = tf.float32),
})
if is_training:
d = d.repeat()
d = d.shuffle(buffer_size=100)
d = d.batch(batch_size=batch_size, drop_remainder=drop_remainder)
return d
return input_fn
# + id="liiTqnPCAabw" colab_type="code" colab={}
# + id="zVek2cNV_q_B" colab_type="code" colab={}
def input_fn_builder_pr_img(img_features,features,hidden_context,seq_length, is_training, drop_remainder):
"""Creates an `input_fn` closure to be passed to TPUEstimator."""
all_input_ids = []
all_input_mask = []
all_segment_ids = []
all_label_ids = []
for feature in features:
all_input_ids.append(feature.input_ids)
all_input_mask.append(feature.input_mask)
all_segment_ids.append(feature.segment_ids)
all_label_ids.append(feature.label_id)
def input_fn(params):
"""The actual input function."""
batch_size = params["batch_size"]
num_examples = len(features)
hidden_shape_img = img_features.shape[-1]
hidden_shape = hidden_context.shape[-1]
# This is for demo purposes and does NOT scale to large data sets. We do
# not use Dataset.from_generator() because that uses tf.py_func which is
# not TPU compatible. The right way to load data is with TFRecordReader.
d = tf.data.Dataset.from_tensor_slices({
"input_ids":
tf.constant(
all_input_ids, shape=[num_examples, seq_length],
dtype=tf.int32),
"input_mask":
tf.constant(
all_input_mask,
shape=[num_examples, seq_length],
dtype=tf.int32),
"segment_ids":
tf.constant(
all_segment_ids,
shape=[num_examples, seq_length],
dtype=tf.int32),
"label_ids":
tf.constant(all_label_ids, shape=[num_examples], dtype=tf.int32),
"img_features":
tf.constant(img_features, shape = [num_examples,hidden_shape_img], dtype = tf.float32),
"hidden_context":
tf.constant(hidden_context, shape = [num_examples,hidden_shape], dtype = tf.float32),
})
if is_training:
d = d.repeat()
d = d.shuffle(buffer_size=100)
d = d.batch(batch_size=batch_size, drop_remainder=drop_remainder)
return d
return input_fn
# + id="a_kkHv98MVLG" colab_type="code" colab={}
def train(output_dir,input_fn,input_fn_builder_progressive = False,hidden_context = None):
CONFIG_FILE = "multi_cased_L-12_H-768_A-12/bert_config.json"
INIT_CHECKPOINT = "multi_cased_L-12_H-768_A-12/bert_model.ckpt"
BATCH_SIZE = 28
LEARNING_RATE = 2e-5
NUM_TRAIN_EPOCHS = 2
# Warmup is a period of time where hte learning rate
# is small and gradually increases--usually helps training.
WARMUP_PROPORTION = 0.1
# Model configs
SAVE_CHECKPOINTS_STEPS = 6000
SAVE_SUMMARY_STEPS = 100
OUTPUT_DIR = output_dir
# Compute # train and warmup steps from batch size
num_train_steps = int(len(input_fn) / BATCH_SIZE * NUM_TRAIN_EPOCHS)
num_warmup_steps = int(num_train_steps * WARMUP_PROPORTION)
print(num_train_steps)
run_config = tf.estimator.RunConfig(
model_dir=OUTPUT_DIR,
save_summary_steps=SAVE_SUMMARY_STEPS,
save_checkpoints_steps=SAVE_CHECKPOINTS_STEPS)
# Specify outpit directory and number of checkpoint steps to save
if input_fn_builder_progressive==False:
model_fn = model_fn_builder(
bert_config=bert.run_classifier.modeling.BertConfig.from_json_file(CONFIG_FILE),
num_labels=4, #number of unique labels
init_checkpoint=INIT_CHECKPOINT,
learning_rate=LEARNING_RATE,
num_train_steps=num_train_steps,
num_warmup_steps=num_warmup_steps,
use_tpu=False,
use_one_hot_embeddings=False
)
estimator = tf.estimator.Estimator(
model_fn=model_fn,
config=run_config,
params={"batch_size": BATCH_SIZE})
train_input_fn = bert.run_classifier.input_fn_builder(
features=input_fn,
seq_length=MAX_SEQ_LENGTH,
is_training=True,
drop_remainder=False)
else:
model_fn_pr = model_fn_builder_progressive(
bert_config=bert.run_classifier.modeling.BertConfig.from_json_file(CONFIG_FILE),
num_labels=4, #number of unique labels
init_checkpoint=INIT_CHECKPOINT,
learning_rate=LEARNING_RATE,
num_train_steps=num_train_steps,
num_warmup_steps=num_warmup_steps,
use_tpu=False,
use_one_hot_embeddings=False
)
estimator = tf.estimator.Estimator(
model_fn=model_fn_pr,
config=run_config,
params={"batch_size": BATCH_SIZE})
train_input_fn = input_fn_builder(
features=input_fn,
hidden_context=hidden_context,
seq_length=MAX_SEQ_LENGTH,
is_training=True,
drop_remainder=False)
print(f'Beginning Training!')
# %timeit
estimator.train(input_fn=train_input_fn, max_steps=num_train_steps)
return estimator
# + id="SE8NmyRD96MR" colab_type="code" colab={}
def train_img(img_features,output_dir,input_fn,input_fn_builder_progressive = False,hidden_context = None):
CONFIG_FILE = "multi_cased_L-12_H-768_A-12/bert_config.json"
INIT_CHECKPOINT = "multi_cased_L-12_H-768_A-12/bert_model.ckpt"
BATCH_SIZE = 28
LEARNING_RATE = 2e-5
NUM_TRAIN_EPOCHS = 2
# Warmup is a period of time where hte learning rate
# is small and gradually increases--usually helps training.
WARMUP_PROPORTION = 0.1
# Model configs
SAVE_CHECKPOINTS_STEPS = 6000
SAVE_SUMMARY_STEPS = 100
OUTPUT_DIR = output_dir
# Compute # train and warmup steps from batch size
num_train_steps = int(len(input_fn) / BATCH_SIZE * NUM_TRAIN_EPOCHS)
num_warmup_steps = int(num_train_steps * WARMUP_PROPORTION)
print(num_train_steps)
run_config = tf.estimator.RunConfig(
model_dir=OUTPUT_DIR,
save_summary_steps=SAVE_SUMMARY_STEPS,
save_checkpoints_steps=SAVE_CHECKPOINTS_STEPS)
# Specify outpit directory and number of checkpoint steps to save
if input_fn_builder_progressive==False:
model_fn = model_fn_builder_img(
bert_config=bert.run_classifier.modeling.BertConfig.from_json_file(CONFIG_FILE),
num_labels=4, #number of unique labels
init_checkpoint=INIT_CHECKPOINT,
learning_rate=LEARNING_RATE,
num_train_steps=num_train_steps,
num_warmup_steps=num_warmup_steps,
use_tpu=False,
use_one_hot_embeddings=False
)
estimator = tf.estimator.Estimator(
model_fn=model_fn,
config=run_config,
params={"batch_size": BATCH_SIZE})
train_input_fn = input_fn_builder_img(
img_features = img_features,
features=input_fn,
seq_length=MAX_SEQ_LENGTH,
is_training=True,
drop_remainder=False)
else:
model_fn_pr = model_fn_builder_img_progressive(
bert_config=bert.run_classifier.modeling.BertConfig.from_json_file(CONFIG_FILE),
num_labels=4, #number of unique labels
init_checkpoint=INIT_CHECKPOINT,
learning_rate=LEARNING_RATE,
num_train_steps=num_train_steps,
num_warmup_steps=num_warmup_steps,
use_tpu=False,
use_one_hot_embeddings=False
)
estimator = tf.estimator.Estimator(
model_fn=model_fn_pr,
config=run_config,
params={"batch_size": BATCH_SIZE})
train_input_fn = input_fn_builder_pr_img(
img_features = img_features,
features=input_fn,
hidden_context=hidden_context,
seq_length=MAX_SEQ_LENGTH,
is_training=True,
drop_remainder=False)
print(f'Beginning Training!')
# %timeit
estimator.train(input_fn=train_input_fn, max_steps=num_train_steps)
return estimator
# + id="ZagTHpcmMWPX" colab_type="code" colab={}
def evaluate_and_get_hidden_context(estimator,input_fn_for_test,input_fn_for_hidden,is_progressive = False,hidden_context=None):
MAX_SEQ_LENGTH = 128
if not is_progressive:
test_input_fn = run_classifier.input_fn_builder(
features=input_fn_for_test,
seq_length=MAX_SEQ_LENGTH,
is_training=False,
drop_remainder=False)
estimator.evaluate(input_fn=test_input_fn, steps=None)
hidden_input_fn = run_classifier.input_fn_builder(
features=input_fn_for_hidden,
seq_length=MAX_SEQ_LENGTH,
is_training=False,
drop_remainder=False)
res = estimator.predict(hidden_input_fn)
hidden_context = []
for i in res:
hidden_context.append(i["hidden_context"])
hidden_context = np.array(hidden_context)
return hidden_context
else:
test_input_fn = input_fn_builder(
features=input_fn_for_test,
hidden_context=hidden_context,
seq_length=MAX_SEQ_LENGTH,
is_training=False,
drop_remainder=False)
estimator.evaluate(input_fn=test_input_fn, steps=None)
# + id="SqdpaiSQ7xbI" colab_type="code" colab={}
#IMG
def evaluate_and_get_hidden_context_img(img_features_for_test,img_features,estimator,input_fn_for_test,input_fn_for_hidden,is_progressive = False,hidden_context=None):
MAX_SEQ_LENGTH = 128
if not is_progressive:
test_input_fn = input_fn_builder_img(
features=input_fn_for_test,
img_features = img_features_for_test,
seq_length=MAX_SEQ_LENGTH,
is_training=False,
drop_remainder=False)
estimator.evaluate(input_fn=test_input_fn, steps=None)
hidden_input_fn = input_fn_builder_img(
features=input_fn_for_hidden,
img_features = img_features,
seq_length=MAX_SEQ_LENGTH,
is_training=False,
drop_remainder=False)
res = estimator.predict(hidden_input_fn)
hidden_context = []
for i in res:
hidden_context.append(i["hidden_context"])
hidden_context = np.array(hidden_context)
return hidden_context
else:
test_input_fn = input_fn_builder_pr_img(
img_features = img_features_for_test,
features=input_fn_for_test,
hidden_context=hidden_context,
seq_length=MAX_SEQ_LENGTH,
is_training=False,
drop_remainder=False)
estimator.evaluate(input_fn=test_input_fn, steps=None)
# + id="NJ7432nv7xYQ" colab_type="code" colab={}
# + id="2MvA0A3I7xWs" colab_type="code" colab={}
# + id="-IpQuhsF7xTn" colab_type="code" colab={}
# + id="r8gVyoiA7xPv" colab_type="code" colab={}
# + id="qwpnFuAgRvsm" colab_type="code" outputId="84490fb2-91fc-47f1-99f2-2ea9fb24748d" colab={"base_uri": "https://localhost:8080/", "height": 1000}
'''
this function trains on english premise and hindi hypothesis
'''
estimator = train('out_dir_train_eng',train_features_eng,input_fn_builder_progressive = False,hidden_context = None)
# + id="O1c3hPMtJjhL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 459} outputId="7e9d1e9d-cf04-4942-ef70-50ea0e3b0632"
'''
this function evaluates and gets hidden context on english premise and hindi hypo
'''
hidden_context_eng = evaluate_and_get_hidden_context(estimator,input_fn_for_test = test_features_eng,input_fn_for_hidden = train_features_eng,is_progressive = False)
# + id="AaIU_UNcJfVT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 445} outputId="ad96ce9c-fe3e-4e93-eaab-b1379da645f4"
'''
this function trains on hindi premise and english hypo
'''
estimator = train('out_dir_train_hindi',train_features_hindi,input_fn_builder_progressive = False,hidden_context = None)
# + id="O295FWzuKLSX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 459} outputId="f5e8b591-47f1-48e0-cb83-213e14b407af"
'''
evaluates on hindi premise and english hypo
'''
hidden_context_hindi = evaluate_and_get_hidden_context(estimator,input_fn_for_test = test_features_hindi,input_fn_for_hidden = train_features_hindi,is_progressive = False)
# + id="bcXafp0NKdQ5" colab_type="code" colab={}
#progressive training and evaluation
# + id="_VoHbzytKdWs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 462} outputId="c2d834d1-76ac-46bd-bb44-dd6208c30a13"
'''
progressively trains on eng-hindi
'''
estimator = train('out_dir_train_eng_pr',train_features_eng,input_fn_builder_progressive = True,hidden_context = hidden_context_hindi)
# + id="R3GvjWJnKdU0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 289} outputId="5a08e5e5-bf54-4881-d918-652204a09464"
'''
progressive evaluation on eng-hindi
'''
dummy = np.random.randn(50,768)
h = evaluate_and_get_hidden_context(estimator,input_fn_for_test = test_features_eng,input_fn_for_hidden = train_features_eng,is_progressive = True,hidden_context=dummy)
# + id="Rw23CFuXTDCp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="429dedbf-884e-493d-bc24-06daae65b026"
print(h)
# + id="0jKDRHVSL9Ur" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 462} outputId="7ba304ca-1761-4876-e3a5-099022d6e747"
'''
progressively trains on hindi-eng
'''
estimator = train('out_dir_train_hindi_pr',train_features_hindi,input_fn_builder_progressive = True,hidden_context = hidden_context_eng)
# + id="vI0kLJF2ME1b" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 289} outputId="48b55b9e-9507-4b98-b842-d67328f5f357"
'''
progressive evaluation on hindi-english
'''
dummy = np.random.randn(50,768)
#dummy = hidden_context_eng[22000:26000,:]
h = evaluate_and_get_hidden_context(estimator,input_fn_for_test = test_features_hindi,input_fn_for_hidden = train_features_hindi,is_progressive = True,hidden_context=dummy)
# + id="sHp-_GYVfcpD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5818df32-7d58-461d-d760-7d85ea00cb14"
hidden_context_eng.shape
# + id="wn8gP81vgGP9" colab_type="code" colab={}
h
# + id="rQs73q4ciUPl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 289} outputId="c38dc0fe-b9bc-45ea-b5d2-e374092a3035"
# !nvidia-smi
# + id="O8TH26DTCGY8" colab_type="code" colab={}
# + id="pUc7IgVwGJ0h" colab_type="code" colab={}
# + id="0vlIVJpdGJyZ" colab_type="code" colab={}
# + id="CpfYjkDxGJve" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 462} outputId="8895cb09-3307-4278-d43b-1ff797863172"
'''
this function trains on english premise and hindi hypothesis with image
'''
train_features = np.random.randn(450,1024)
estimator = train_img(train_features,'out_dir_train_eng_im',train_features_eng,input_fn_builder_progressive = False,hidden_context = None)
# + id="iDz-gxlBGJr6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 493} outputId="f0877f93-99b4-4f38-9d06-e0f9209b427c"
'''
this function evaluates and gets hidden context on english premise and hindi hypo with image
'''
test_features = np.random.randn(50,1024)
evaluate_features = np.random.randn(450,1024)
hidden_context_eng = evaluate_and_get_hidden_context_img(test_features,evaluate_features,estimator,input_fn_for_test = test_features_eng,input_fn_for_hidden = train_features_eng,is_progressive = False)
# + id="so33xLAoHpEY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 496} outputId="ada6f683-8705-4e8f-dd42-a3f545bdd38e"
'''
progressively trains on hindi-eng
'''
train_features = np.random.randn(450,1024)
estimator = train_img(train_features,'out_dir_train_hindi_pr1',train_features_hindi,input_fn_builder_progressive = True,hidden_context = hidden_context_eng)
# + id="Y2f-SKV0HpBQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 323} outputId="ba7a91fd-c18b-4cf7-f77f-5c78d0596204"
'''
progressive evaluation on hindi-english
'''
dummy = np.random.randn(50,768)
test_features = np.random.randn(50,1024)
evaluate_features = np.random.randn(450,1024)
h = evaluate_and_get_hidden_context_img(test_features,evaluate_features,estimator,input_fn_for_test = test_features_hindi,input_fn_for_hidden = train_features_hindi,is_progressive = True,hidden_context=dummy)
# + id="Oq3eFanvf0IB" colab_type="code" colab={}
h
# + id="DBlozuaV4ptP" colab_type="code" colab={}
| multilingual_bert_infomax_with_image_final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:fastai]
# language: python
# name: conda-env-fastai-py
# ---
# https://github.com/fastai/course-v3/blob/master/nbs/dl1/lesson3-planet.ipynb
#
# Read in parquet: https://www.kaggle.com/sohier/getting-started-reading-the-data
# +
from fastai.vision import *
import pandas as pd
from tqdm import tqdm
import matplotlib.pyplot as plt
import numpy as np
import pretrainedmodels
from sklearn.model_selection import StratifiedKFold
import ImageDataAugmentation as ida
import cv2
# +
# #!pip install pyarrow==0.15.*
# #!pip install nbdev
# #!pip install pretrainedmodels
# +
# #!apt-get install -y libsm6 libxext6 libxrender-dev
# #!pip install opencv-python
# -
source = Path('/home/kaggle/bengaliai-cv19/input')
source
df_label = pd.read_csv(source/'train.csv')
df_label.head()
cls_count=[df_label.grapheme_root.nunique(), df_label.vowel_diacritic.nunique(), df_label.consonant_diacritic.nunique()]
cls_count
# +
train = pd.DataFrame(df_label['image_id'])
#train['tags'] = df_label['grapheme_root'].astype(str) #+' '+ df_label['vowel_diacritic'].astype(str) +' '+ df_label['consonant_diacritic'].astype(str)
#train['tags'] = df_label['vowel_diacritic'].astype(str)
#train['tags'] = df_label['consonant_diacritic'].astype(str)
# train.groupby(['grapheme_root','vowel_diacritic','consonant_diacritic']).count()
train = df_label.drop('grapheme', axis=1)
train['tag'] = train['grapheme_root'].astype(str)+'_'+train['vowel_diacritic'].astype(str)+'_'+train['consonant_diacritic'].astype(str)
# +
n_splits=5
skf = StratifiedKFold(n_splits=n_splits, shuffle=True, random_state=2020)
for fold, (train_idx, val_idx) in enumerate(skf.split(X=train, y=train['tag'].values)):
train['valid_kf_'+str(fold)] = 0
train.loc[val_idx, 'valid_kf_'+str(fold)] = 1
# -
train.head()
# +
#train.describe()
#train.groupby(['grapheme_root','vowel_diacritic','consonant_diacritic'])['valid_kf_1'].sum()
# -
# # Load all
# +
def read_all(file='train_image_data_crop_scaled_'):
df = (pd.read_feather(source/(file+'0.feather'))
.append(pd.read_feather(source/(file+'1.feather')))
.append(pd.read_feather(source/(file+'2.feather')))
.append(pd.read_feather(source/(file+'3.feather')))
)
df.reset_index(inplace=True,drop = True)
return df
df_train = read_all('train_image_data_crop_scaled_')
# -
# # Load one
# +
def read_one(file='train_image_data_crop_scaled_0.feather'):
df = pd.read_feather(source/(file))
df.reset_index(inplace=True,drop = True)
return df
df_train = read_one('train_image_data_crop_scaled_0.feather') #train_image_data_crop_scaled_0
#df_train = read_one('train_image_data_resize_only_0.feather')#'train_image_data_crop_scaled_0.feather') #train_image_data_crop_scaled_0
#df_train = pd.read_parquet('input/train_image_data_0.parquet')
#df_train.reset_index(inplace=True,drop = True)#
# +
#df_train=df_train.append(pd.read_feather(source/('train_image_data_resize_only_0.feather')))
#df_train=df_train.append(pd.read_feather(source/('train_image_data_resize_only_1.feather')))
#df_train=df_train.append(pd.read_feather(source/('train_image_data_resize_only_2.feather')))
#df_train=df_train.append(pd.read_feather(source/('train_image_data_resize_only_3.feather')))
# -
df_train.shape
# + active=""
#
# +
#train.groupby(['grapheme_root','vowel_diacritic','consonant_diacritic']).count().sort_values('image_id', ascending=False)
# -
# # Merge with tags
df_train = df_train.merge(train, on='image_id')
df_train['fn'] = df_train.index
df_train.head(10)
# +
#df_train[df_train['grapheme_root']==48][df_train['vowel_diacritic']==4][df_train['consonant_diacritic']==0]
# +
#data.get(4526)
# +
#### df_train.groupby(['tag'])['valid_kf_4'].sum()
# -
SIZE = 128
# +
tfms = get_transforms(do_flip=False,max_zoom=1, xtra_tfms=[cutout(n_holes=(1,4), length=(16, 16), p=.5)]) #,rand_zoom(scale=(.8,1))
#tfms = ([cutout(n_holes=(1,4), length=(16, 16), p=.5)],[])
tfms[0][0].kwargs['padding_mode'] = 'zeros'
tfms[1][0].kwargs['padding_mode'] = 'zeros'
# -
tfms
#https://www.kaggle.com/melissarajaram/model-ensembling-and-transfer-learning
class PixelImageItemList(ImageList):
def open(self,fn):
img_pixel = self.inner_df.loc[self.inner_df['fn'] == int(fn[2:])].values[0,1:(SIZE*SIZE+1)] #.values[0,1:32333]#
img_pixel = img_pixel.reshape(SIZE,SIZE) #.reshape(137,236)#
#img_pixel = np.stack((img_pixel,)*3,axis=-1)
return vision.Image((pil2tensor(img_pixel,np.float32).div_(255)-1).abs_())
class PixelImageItemList_full(ImageList):
def __init__(self, myimages = {}, *args, **kwargs):
super().__init__(*args, **kwargs)
self.myimages = myimages
def open(self,fn):
return self.myimages.get(fn)
@classmethod
def from_df(cls, df:DataFrame, path:PathOrStr, cols:IntsOrStrs=0, folder:PathOrStr=None, suffix:str='', **kwargs)->'ItemList':
"Get the filenames in `cols` of `df` with `folder` in front of them, `suffix` at the end."
res = super().from_df(df, path=path, cols=cols, **kwargs)
# full load of all images
for i, row in tqdm(df.drop(labels=['image_id','fn','grapheme_root','vowel_diacritic','consonant_diacritic',
'tag','valid_kf_0','valid_kf_1','valid_kf_2','valid_kf_3','valid_kf_4'],axis=1).iterrows()):
# Numpy to Image conversion from
# https://www.kaggle.com/heye0507/fastai-1-0-with-customized-itemlist
img_pixel = row.values.reshape(SIZE,SIZE) #.reshape(137,236)
img_pixel = np.stack((img_pixel,)*1,axis=-1)
'''
lvl = 0
go_on = True
while(go_on):
if np.sum(img_pixel[-1,:,0])==0:
lvl+=1
img_pixel=np.roll(img_pixel,1,axis=0)
else:
go_on=False
#img_pixel[lvl:,:,0]=img_pixel[:-lvl,:,0]
#img_pixel[:lvl,:,0]=0
# img_pixel = np.stack((img_pixel,)*5,axis=-1)
## override Top
img_pixel[:16+lvl,:,1]=0
## Bottom
img_pixel[16+lvl:,:,2]=0
## Left
img_pixel[:,14:,3]=0#[:,14:]=0
## Right
img_pixel[:,:14,4]=0#[:,:14]=0
# remove original layer
# img_pixel=img_pixel[:,:,1:]
'''
#https://discuss.pytorch.org/t/how-to-calculate-the-gradient-of-images/1407/5
#Black and white input image x, 1x1xHxW
a = torch.Tensor([[1, 0, -1],
[2, 0, -2],
[1, 0, -1]])
x=pil2tensor(img_pixel,np.float32).div_(255).unsqueeze(0)
a = a.view((1,1,3,3))
G_x = F.conv2d(x, a, padding=1)
b = torch.Tensor([[1, 2, 1],
[0, 0, 0],
[-1, -2, -1]])
b = b.view((1,1,3,3))
G_y = F.conv2d(x, b, padding=1)
G = torch.sqrt(torch.pow(G_x,2)+ torch.pow(G_y,2))
#res = P(G)
#
PI = torch.acos(torch.Tensor([-1]))
#D = ((torch.round((torch.atan2(G_y, G_x) * 180 / PI) / 45) * 45) + 180) % 180
D1 = ((torch.round((torch.atan2(G_x, G_y) * 180 / PI) / 45) * 45) + 180) % 180
D2 = ((torch.round((torch.atan2(G_y, G_x) * 180 / PI) / 45) * 45) + 180) % 180
limg = torch.cat((pil2tensor(img_pixel,np.float32).div_(255),D1.squeeze(0).div_(180),D2.squeeze(0).div_(180)))
#print(D)
# vision.Image(pil2tensor(img_pixel,np.float32).div_(255)).show()
#vision.Image(G_x.squeeze(0)).show()
#vision.Image(G_y.squeeze(0)).show()
#vision.Image(G.squeeze(0)).show()
# vision.Image(D1.squeeze(0)).show()
# vision.Image(D2.squeeze(0)).show()
#print(img_pixel[:,:,1][6])
#print(img_pixel[:,:,2][6])
#print(img_pixel[:,:,3][6])
#print(img_pixel[:,:,4][6])
# break
res.myimages[res.items[i]]=vision.Image(limg)
#res.myimages[res.items[i]]=vision.Image(pil2tensor(img_pixel,np.float32).div_(255))
return res
# %%time
piil = PixelImageItemList.from_df(df=df_train,path='.',cols='fn')
#piil = PixelImageItemList_full.from_df(df=df_train,path='.',cols='fn')
# +
#df_train.tags
# +
bs = 128 #64
valid_col = 'valid_kf_2'
data = (piil#
.split_by_rand_pct(0.2)
#.label_from_df(cols='tags')#,label_delim=' ')
#.split_from_df(col = valid_col)
.label_from_df(cols=['grapheme_root','vowel_diacritic','consonant_diacritic'])
.transform(tfms, padding_mode='zeros')#, size=(96,168)) #([cutout(n_holes=(1,4), length=(16, 16), p=.5),rand_zoom(scale=(.8,1))],[]), padding_mode='zeros')#tfms) #, size=128)
.databunch(bs=bs).normalize(imagenet_stats)) #mnist_stats
'''(piil
.split_by_rand_pct(0.2)
.label_from_df(cols='tags',label_delim=' ')
.databunch(bs=32))'''
# +
#data.train_dl.dl.dataset.tfms
# -
data.show_batch(rows=3, figsize=(12,9))
# +
#data
# -
leak=0.1
best_architecture = nn.Sequential(
conv_layer(3,32,stride=1,ks=3,leaky=leak),
conv_layer(32,32,stride=2,ks=3,leaky=leak),
conv_layer(32,32,stride=2,ks=5,leaky=leak),
nn.Dropout(0.4),
conv_layer(32,32,stride=1,ks=3,leaky=leak),
conv_layer(32,32,stride=2,ks=3,leaky=leak),
conv_layer(32,32,stride=2,ks=5,leaky=leak),
nn.Dropout(0.4),
conv_layer(32,64,stride=1,ks=3,leaky=leak),
conv_layer(64,64,stride=2,ks=3,leaky=leak),
conv_layer(64,64,stride=2,ks=5,leaky=leak),
nn.Dropout(0.4),
Flatten(),
nn.Linear(64, 32),#
relu(inplace=True),
nn.BatchNorm1d(32),
nn.Dropout(0.4),
nn.Linear(32,168)
)
# +
batch_size = 5
nb_digits = 10
# Dummy input that HAS to be 2D for the scatter (you can use view(-1,1) if needed)
y = torch.LongTensor(batch_size).random_() % nb_digits
# One hot encoding buffer that you create out of the loop and just keep reusing
y_onehot = torch.FloatTensor(batch_size, nb_digits)
print(y, y.unsqueeze(1))
# In your for loop
y_onehot.zero_()
y_onehot.scatter_(1, y.unsqueeze(1),1)
print(y_onehot)
F.one_hot(y, num_classes=10)
# +
'''#https://discuss.pytorch.org/t/convert-int-into-one-hot-format/507/4
def y_onehot(y, cardinality):
#print('in1H:',y, batch_size, cardinality)
batch_size = y.shape[0]
out = torch.FloatTensor(batch_size, cardinality).cuda()
out.zero_()
try:
out.scatter_(1, y.unsqueeze(1),1)
except:
print(out,y)
print(out.shape,y.shape)
return out'''
#https://www.kaggle.com/mnpinto/bengali-ai-fastai2-starter-lb0-9598
# Loss function
'''class Loss_combine(Module):
def __init__(self, func=F.cross_entropy, weights=[2, 1, 1]):
self.func, self.w = func, weights
def forward(self, xs, *ys):
for i, w, x, y in zip(range(len(xs)), self.w, xs, ys):
print(x, y)
if i == 0: loss = w*self.func(x, y)
else: loss += w*self.func(x, y)
return loss'''
#https://www.kaggle.com/iafoss/grapheme-fast-ai-starter-lb-0-964
class Loss_combine(nn.Module):
def __init__(self):
super().__init__()
def forward(self, input, target,reduction='mean'):
x1,x2,x3 = input
x1,x2,x3 = x1.float(),x2.float(),x3.float()
y = target.long()
return 0.5*F.cross_entropy(x1,y[:,0],reduction=reduction) + 0.25*F.cross_entropy(x2,y[:,1],reduction=reduction) + \
0.25*F.cross_entropy(x3,y[:,2],reduction=reduction)
class Metric_idx(Callback):
def __init__(self, idx, average='macro'):
super().__init__()
self.idx = idx
self.n_classes = 0
self.average = average
self.cm = None
self.eps = 1e-9
def on_epoch_begin(self, **kwargs):
self.tp = 0
self.fp = 0
self.cm = None
def on_batch_end(self, last_output:Tensor, last_target:Tensor, **kwargs):
last_output = last_output[self.idx]
last_target = last_target[:,self.idx]
preds = last_output.argmax(-1).view(-1).cpu()
targs = last_target.long().cpu()
if self.n_classes == 0:
self.n_classes = last_output.shape[-1]
self.x = torch.arange(0, self.n_classes)
cm = ((preds==self.x[:, None]) & (targs==self.x[:, None, None])) \
.sum(dim=2, dtype=torch.float32)
if self.cm is None: self.cm = cm
else: self.cm += cm
def _weights(self, avg:str):
if self.n_classes != 2 and avg == "binary":
avg = self.average = "macro"
warn("average=`binary` was selected for a non binary case. \
Value for average has now been set to `macro` instead.")
if avg == "binary":
if self.pos_label not in (0, 1):
self.pos_label = 1
warn("Invalid value for pos_label. It has now been set to 1.")
if self.pos_label == 1: return Tensor([0,1])
else: return Tensor([1,0])
elif avg == "micro": return self.cm.sum(dim=0) / self.cm.sum()
elif avg == "macro": return torch.ones((self.n_classes,)) / self.n_classes
elif avg == "weighted": return self.cm.sum(dim=1) / self.cm.sum()
def _recall(self):
rec = torch.diag(self.cm) / (self.cm.sum(dim=1) + self.eps)
if self.average is None: return rec
else:
if self.average == "micro": weights = self._weights(avg="weighted")
else: weights = self._weights(avg=self.average)
return (rec * weights).sum()
def on_epoch_end(self, last_metrics, **kwargs):
return add_metrics(last_metrics, self._recall())
Metric_grapheme = partial(Metric_idx,0)
Metric_vowel = partial(Metric_idx,1)
Metric_consonant = partial(Metric_idx,2)
class Metric_tot(Callback):
def __init__(self):
super().__init__()
self.grapheme = Metric_idx(0)
self.vowel = Metric_idx(1)
self.consonant = Metric_idx(2)
def on_epoch_begin(self, **kwargs):
self.grapheme.on_epoch_begin(**kwargs)
self.vowel.on_epoch_begin(**kwargs)
self.consonant.on_epoch_begin(**kwargs)
def on_batch_end(self, last_output:Tensor, last_target:Tensor, **kwargs):
self.grapheme.on_batch_end(last_output, last_target, **kwargs)
self.vowel.on_batch_end(last_output, last_target, **kwargs)
self.consonant.on_batch_end(last_output, last_target, **kwargs)
def on_epoch_end(self, last_metrics, **kwargs):
return add_metrics(last_metrics, 0.5*self.grapheme._recall() +
0.25*self.vowel._recall() + 0.25*self.consonant._recall())
# -
acc_02 = partial(accuracy_thresh, thresh=0.2)
f_score = partial(fbeta, thresh=0.2)
# +
#https://github.com/osmr/imgclsmob/blob/master/pytorch/pytorchcv/models/common.py
def conv1x1(in_channels,
out_channels,
stride=1,
groups=1,
bias=False):
"""
Convolution 1x1 layer.
Parameters:
----------
in_channels : int
Number of input channels.
out_channels : int
Number of output channels.
stride : int or tuple/list of 2 int, default 1
Strides of the convolution.
groups : int, default 1
Number of groups.
bias : bool, default False
Whether the layer uses a bias vector.
"""
return nn.Conv2d(
in_channels=in_channels,
out_channels=out_channels,
kernel_size=1,
stride=stride,
groups=groups,
bias=bias)
class SEBlock(nn.Module):
"""
Squeeze-and-Excitation block from 'Squeeze-and-Excitation Networks,' https://arxiv.org/abs/1709.01507.
Parameters:
----------
channels : int
Number of channels.
reduction : int, default 16
Squeeze reduction value.
round_mid : bool, default False
Whether to round middle channel number (make divisible by 8).
activation : function, or str, or nn.Module, default 'relu'
Activation function after the first convolution.
out_activation : function, or str, or nn.Module, default 'sigmoid'
Activation function after the last convolution.
"""
def __init__(self,
channels,
reduction=16,
round_mid=False,
mid_activation=(lambda: nn.ReLU(inplace=True)),
out_activation=(lambda: nn.Sigmoid())):
super(SEBlock, self).__init__()
mid_channels = channels // reduction if not round_mid else round_channels(float(channels) / reduction)
self.pool = nn.AdaptiveAvgPool2d(output_size=1)
self.conv1 = conv1x1(
in_channels=channels,
out_channels=mid_channels,
bias=True)
self.activ = nn.ReLU(inplace=True)
self.conv2 = conv1x1(
in_channels=mid_channels,
out_channels=channels,
bias=True)
self.sigmoid = nn.Sigmoid()
def forward(self, x):
identity = x
w = self.pool(x)
w = self.conv1(w)
w = self.activ(w)
w = self.conv2(w)
w = self.sigmoid(w)
x = x * w
x = x + identity
return x
# +
#arch = models.densenet121 #resnet50
import torchvision.models as torch_models
arch = torch_models.resnext50_32x4d #
#arch = torch_models.resnext101_32x8d #
def se_resnext50(pretrained=True):
pretrained = 'imagenet' if pretrained else None
model = pretrainedmodels.__dict__['se_resnext50_32x4d'](num_classes=1000, pretrained=pretrained)
return model
#arch = models.cadene_models.se_resnext50_32x4d(pretrained=True)
# -
class SAI(nn.Module):
''' SelfAttention with Identity '''
def __init__(self, nf):
super(SAI, self).__init__()
self.sa = PooledSelfAttention2d(nf)
self.bn = nn.BatchNorm2d(nf)
self.do = nn.Dropout(0.4),
def forward(self, x):
ident = x
out = self.sa(x)
out = ident + out
out = self.do(self.bn(out)) #
return nn.ReLU(out)
# +
class HeadBlock0(nn.Module):
def __init__(self, nf, nc):
super(HeadBlock0, self).__init__()
self.head1 = create_head(nf, 1024, bn_final = True)
self.at1 = SelfAttention(nf//4)
self.rl1 = nn.LeakyReLU(0.1, inplace=True)
self.lin1 = nn.Linear(1024, nc[0])
self.head2 = create_head(nf, 1024, bn_final = True)
self.at2 = SelfAttention(nf//4)
self.rl2 = nn.LeakyReLU(0.1, inplace=True)
self.lin2 = nn.Linear(1024, nc[1])
self.head3 = create_head(nf, 1024, bn_final = True)
self.at3 = SelfAttention(nf//4)
self.rl3 = nn.LeakyReLU(0.1, inplace=True)
self.lin3 = nn.Linear(1024, nc[2])
def forward(self, x):
x1 = self.head1(x)
x1 = self.at1(x1)
x1 = self.rl1(x1)
x1 = self.lin1(x1)
x2 = self.head2(x)
x2 = self.at2(x2)
x2 = self.rl2(x2)
x2 = self.lin2(x2)
x3 = self.head3(x)
x3 = self.at3(x3)
x3 = self.rl3(x3)
x3 = self.lin3(x3)
return x1, x2, x3
class HeadBlock(nn.Module):
def __init__(self, nf, nc):
super(HeadBlock, self).__init__()
self.se1 = SEBlock(nf//2)
#self.at1 = SEBlock(nf//2)
self.head1 = create_head(nf, nc[0])
#self.sa1 = SelfAttention(nf//4)
self.se2 = SEBlock(nf//2)
#self.at2 = SEBlock(nf//2)
self.head2 = create_head(nf, nc[1])
#self.sa2 = SelfAttention(nf//4)
self.se3 = SEBlock(nf//2)
#self.at3 = SEBlock(nf//2)
self.head3 = create_head(nf, nc[2])
#self.sa3 = SelfAttention(nf//4)
def forward(self, x):
x1 = self.se1(x)
##x1 = self.at1(x1)
x1 = self.head1(x1)
x2 = self.se2(x)
##x2 = self.at2(x2)
x2 = self.head2(x2)
x3 = self.se3(x)
##x3 = self.at3(x3)
x3 = self.head3(x3)
return x1, x2, x3
class HeadBlock2(nn.Module):
def __init__(self, nf, nc):
super(HeadBlock2, self).__init__()
self.head1 = create_head(nf, nc[0])
self.head2 = create_head(nf, nc[1])
self.head3 = create_head(nf, nc[2])
self.rl1 = nn.LeakyReLU(0.1, inplace=True)
self.rl2 = nn.LeakyReLU(0.1, inplace=True)
self.rl3 = nn.LeakyReLU(0.1, inplace=True)
self.bn1 = nn.BatchNorm1d(nc[0])
self.bn2 = nn.BatchNorm1d(nc[1])
self.bn3 = nn.BatchNorm1d(nc[2])
#self.flatt1 = Flatten(nc[0])
#self.flatt2 = Flatten(nc[1])
#self.flatt3 = Flatten(nc[2])
self.att = SelfAttention(sum(nc))
self.head4 = nn.Linear(sum(nc)+nc[0], nc[0])
self.head5 = nn.Linear(sum(nc)+nc[1], nc[1])
self.head6 = nn.Linear(sum(nc)+nc[2], nc[2])
def forward(self, x):
x1 = self.head1(x)
x2 = self.head2(x)
x3 = self.head3(x)
x1 = self.bn1(self.rl1(x1))
x2 = self.bn2(self.rl2(x2))
x3 = self.bn3(self.rl3(x3))
f = torch.cat([(x1),(x2),(x3)],1)
b = self.att(f)
x4 = self.head4(torch.cat([x1 , b], 1))
x5 = self.head5(torch.cat([x2 , b], 1))
x6 = self.head6(torch.cat([x3 , b], 1))
return x4, x5, x6
''' (0): AdaptiveConcatPool2d(
(ap): AdaptiveAvgPool2d(output_size=1)
(mp): AdaptiveMaxPool2d(output_size=1)
)
(1): Flatten()
(2): BatchNorm1d(4096, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(3): Dropout(p=0.25, inplace=False)
(4): Linear(in_features=4096, out_features=512, bias=True)
(5): ReLU(inplace=True)
(6): BatchNorm1d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(7): Dropout(p=0.5, inplace=False)
(8): Linear(in_features=512, out_features=7, bias=True)
'''
def threeHeads(nf, nc):
layers =[HeadBlock0(nf,nc)]
return nn.Sequential(*layers)
# +
#https://www.kaggle.com/iafoss/grapheme-fast-ai-starter-lb-0-964#MixUp
class MixUpLoss(Module):
"Adapt the loss function `crit` to go with mixup."
def __init__(self, crit, reduction='mean'):
super().__init__()
if hasattr(crit, 'reduction'):
self.crit = crit
self.old_red = crit.reduction
setattr(self.crit, 'reduction', 'none')
else:
self.crit = partial(crit, reduction='none')
self.old_crit = crit
self.reduction = reduction
def forward(self, output, target):
if len(target.shape) == 2 and target.shape[1] == 7:
loss1, loss2 = self.crit(output,target[:,0:3].long()), self.crit(output,target[:,3:6].long())
d = loss1 * target[:,-1] + loss2 * (1-target[:,-1])
else: d = self.crit(output, target)
if self.reduction == 'mean': return d.mean()
elif self.reduction == 'sum': return d.sum()
return d
def get_old(self):
if hasattr(self, 'old_crit'): return self.old_crit
elif hasattr(self, 'old_red'):
setattr(self.crit, 'reduction', self.old_red)
return self.crit
class MixUpCallback(LearnerCallback):
"Callback that creates the mixed-up input and target."
def __init__(self, learn:Learner, alpha:float=0.4, stack_x:bool=False, stack_y:bool=True):
super().__init__(learn)
self.alpha,self.stack_x,self.stack_y = alpha,stack_x,stack_y
def on_train_begin(self, **kwargs):
if self.stack_y: self.learn.loss_func = MixUpLoss(self.learn.loss_func)
def on_batch_begin(self, last_input, last_target, train, **kwargs):
"Applies mixup to `last_input` and `last_target` if `train`."
if not train: return
lambd = np.random.beta(self.alpha, self.alpha, last_target.size(0))
lambd = np.concatenate([lambd[:,None], 1-lambd[:,None]], 1).max(1)
lambd = last_input.new(lambd)
shuffle = torch.randperm(last_target.size(0)).to(last_input.device)
x1, y1 = last_input[shuffle], last_target[shuffle]
if self.stack_x:
new_input = [last_input, last_input[shuffle], lambd]
else:
out_shape = [lambd.size(0)] + [1 for _ in range(len(x1.shape) - 1)]
new_input = (last_input * lambd.view(out_shape) + x1 * (1-lambd).view(out_shape))
if self.stack_y:
new_target = torch.cat([last_target.float(), y1.float(), lambd[:,None].float()], 1)
else:
if len(last_target.shape) == 2:
lambd = lambd.unsqueeze(1).float()
new_target = last_target.float() * lambd + y1.float() * (1-lambd)
return {'last_input': new_input, 'last_target': new_target}
def on_train_end(self, **kwargs):
if self.stack_y: self.learn.loss_func = self.learn.loss_func.get_old()
# +
class CutMixCallback(LearnerCallback):
"Callback that creates the cutmixed input and target."
def __init__(self, learn:Learner, α:float=1., stack_y:bool=True, stack_x:bool=True):
super().__init__(learn)
self.α,self.stack_y,self.stack_x = α,stack_y,stack_x
def on_train_begin(self, **kwargs):
if self.stack_y: self.learn.loss_func = MixUpLoss(self.learn.loss_func)
def on_batch_begin(self, last_input, last_target, train, **kwargs):
if not train: return
self.learn.loss_func = MixUpLoss(self.learn.loss_func)
lambd = np.random.beta(self.α, self.α)
lambd = max(lambd, 1- lambd)
shuffle = torch.randperm(last_target.size(0)).to(last_input.device)
x1, y1 = last_input[shuffle], last_target[shuffle]
#Get new input
last_input_size = last_input.shape
bbx1, bby1, bbx2, bby2 = rand_bbox(last_input.size(), lambd)
new_input = last_input.clone()
new_input[:, ..., bby1:bby2, bbx1:bbx2] = last_input[shuffle, ..., bby1:bby2, bbx1:bbx2]
lambd = last_input.new([lambd])
if self.stack_x:
lambd = 1 - ((bbx2 - bbx1) * (bby2 - bby1) / (last_input_size[-1] * last_input_size[-2]))
lambd = last_input.new([lambd])
if self.stack_y:
new_target = torch.cat([last_target.float(), y1.float(),
lambd.repeat(last_input_size[0]).unsqueeze(1).float()], 1)
else:
if len(last_target.shape) == 2:
lambd = lambd.unsqueeze(1).float()
new_target = last_target.float() * lambd + y1.float() * (1-lambd)
return {'last_input': new_input, 'last_target': new_target}
def on_train_end(self, **kwargs):
if self.stack_y: self.learn.loss_func = self.learn.loss_func.get_old()
def rand_bbox(last_input_size, λ):
'''lambd is always between .5 and 1'''
W = last_input_size[-1]
H = last_input_size[-2]
cut_rat = np.sqrt(1. - λ) # 0. - .707
cut_w = np.int(W * cut_rat)
cut_h = np.int(H * cut_rat)
# uniform
cx = np.random.randint(W)
cy = np.random.randint(H)
bbx1 = np.clip(cx - cut_w // 2, 0, W)
bby1 = np.clip(cy - cut_h // 2, 0, H)
bbx2 = np.clip(cx + cut_w // 2, 0, W)
bby2 = np.clip(cy + cut_h // 2, 0, H)
return bbx1, bby1, bbx2, bby2
# -
class Loss_dist(nn.Module):
def __init__(self):
super().__init__()
def forward(self, input, target,reduction='mean'):
x1,x2,x3 = input
#print(x2.shape,target.shape)
#print(target.long())
x1,x2,x3 = x1.float(),x2.float(),x3.float()
y = target.long()
#print(x2,y[:,1])
#return 0.5*F.cross_entropy(x1,y[:,0],reduction=reduction) + 0.25*F.cross_entropy(x2,y[:,1],reduction=reduction) + \
# 0.25*F.cross_entropy(x3,y[:,2],reduction=reduction)
#print('COS: ',F.cosine_similarity(x2,y_onehot(y[:,1], x2.shape[1]),1))
#print('ENT: ',F.log_softmax(x2,1))
bs = x1.shape[0]
t = 1
#print('bs', bs)
#print('OHE',F.one_hot(y[:,0], num_classes = x1.shape[1]))
#print('cos_sim',F.cosine_similarity(x1,F.one_hot(y[:,0], num_classes = x1.shape[1]),1).exp())
loss = [None]*3
for i, x_i in enumerate([x1,x2,x3]):
num_classes = x_i.shape[1]
cos_sim_exp_1 = (F.cosine_similarity(x_i,F.one_hot(y[:,i], num_classes = num_classes),1)*t).exp()
#cos_sim_exp_1 = torch.pow(x_i-F.one_hot(y[:,i], num_classes = num_classes), 2).sum(0).exp()
cos_sim_exp_1_sum = cos_sim_exp_1.sum()
#cos_sim_exp_2 = (-F.cosine_similarity(x_i,F.one_hot(y[:,i], num_classes = num_classes),1)).exp()
#cos_sim_exp_2_sum = cos_sim_exp_2.sum()
softmax_1 = cos_sim_exp_1 / (cos_sim_exp_1_sum-cos_sim_exp_1)
#softmax_2 = cos_sim_exp_2 / (cos_sim_exp_2_sum-cos_sim_exp_2)
ll_1 = -softmax_1.log()
#ll_2 = -softmax_2.log()
loss[i] = (ll_1.sum()/bs)*0.5 + (0.5*F.cross_entropy(x1,y[:,0],reduction=reduction) + 0.25*F.cross_entropy(x2,y[:,1],reduction=reduction) + \
0.25*F.cross_entropy(x3,y[:,2],reduction=reduction))*0.5
#print('sm_1', softmax_1)
#print('ll_1',ll_1)
#print('new_loss', loss)
#print('softmax',F.softmax(F.cosine_similarity(x1,F.one_hot(y[:,0], num_classes = x1.shape[1]),1),dim=0))
#print('c_ent',F.cross_entropy(x1,y[:,0],reduction=reduction))
return 0.5*loss[0]+0.25*loss[1]+0.25*loss[2]
#print('XXX 1hot:', y_onehot(y[:,1], bs, x2.shape[1]))
#return 0.5*F.cosine_embedding_loss(x1,y_onehot(y[:,0], x1.shape[1]),y[:,0]) + \
# 0.25*F.cosine_embedding_loss(x2,y_onehot(y[:,1], x2.shape[1]),y[:,1]) + \
# 0.25*F.cosine_embedding_loss(x3,y_onehot(y[:,2], x3.shape[1]),y[:,2])
#print(F.cosine_similarity(x1,y_onehot(y[:,0], bs, x1.shape[1]),1).unsqueeze(1))
#print('##',F.cosine_similarity(x1,y_onehot(y[:,0], bs, x1.shape[1]),1).unsqueeze(1).shape,'##',x1, x1.shape)
#print('##mean##',F.cosine_similarity(x1,y_onehot(y[:,0], bs, x1.shape[1]),1).mean())
#print('##cos_l##',F.cosine_embedding_loss(x1,y_onehot(y[:,0], bs, x1.shape[1]),y[:,0]))
#print('##cent##',F.cross_entropy(x1,y[:,0],reduction=reduction))
#return 0.5*F.cosine_similarity(x1,y_onehot(y[:,0], x1.shape[1]),1).abs().mean() + \
# 0.25*F.cosine_similarity(x2,y_onehot(y[:,1], x2.shape[1]),1).abs().mean() + \
# 0.25*F.cosine_similarity(x3,y_onehot(y[:,2], x3.shape[1]),1).abs().mean()
# +
#arch
# +
# create learner
#Loss_combine(),torch.nn.BCEWithLogitsLoss()
#se_resnext50 #arch
learn = cnn_learner(data, se_resnext50, cut=-2, custom_head=threeHeads(4096, cls_count), loss_func = Loss_combine(),metrics=[Metric_grapheme(),Metric_vowel(),Metric_consonant(),Metric_tot()]).to_fp16()
#learn.freeze()
learn.clip_grad = 1.0
#learn.split([learn.model[1]])
#learn.unfreeze()
# -
learn.unfreeze()
# +
#learn.model
# +
learn.unfreeze()
for i in range(3):
#learn.model[0][4][i].add_module('self_se1',SEBlock(256))
#learn.model[0][4][i].add_module('self_att1',PooledSelfAttention2d(256)) #PooledSelfAttention2d
learn.model[0][4][i].add_module('self_att1',SAI(256)) #PooledSelfAttention2d
for i in range(4):
#learn.model[0][5][i].add_module('self_se2',SEBlock(512))
#learn.model[0][5][i].add_module('self_att2',PooledSelfAttention2d(512)) #PooledSelfAttention2d
learn.model[0][5][i].add_module('self_att2',SAI(512)) #PooledSelfAttention2d
for i in range(6): #23 #6
#learn.model[0][6][i].add_module('self_se3',SEBlock(512))
#learn.model[0][6][i].add_module('self_att3',PooledSelfAttention2d(512)) #PooledSelfAttention2d
learn.model[0][6][i].add_module('self_att3',SAI(512)) #PooledSelfAttention2d
for i in range(3):
#learn.model[0][7][i].add_module('self_se4',SEBlock(512))
#learn.model[0][7][i].add_module('self_att4',PooledSelfAttention2d(512)) #PooledSelfAttention2d
learn.model[0][7][i].add_module('self_att4',SAI(512)) #PooledSelfAttention2d
# +
#learn.model
# -
#learn.freeze()
#learn.to_fp32()
learn.lr_find()
learn.recorder.plot(suggestion=True)
lr = 1e-2 #2
n_cycle = 10
# +
csvlogger = callbacks.CSVLogger(learn,'learn_log_2')
reduceLR = callbacks.ReduceLROnPlateauCallback(learn=learn, monitor = 'valid_loss', mode = 'auto', patience = 2, factor = 0.2, min_delta = 0)
#learn.fit(1,1e-4) #slice(lr)
learn.fit_one_cycle(n_cycle, slice(lr), callbacks=[reduceLR, MixUpCallback(learn), csvlogger]) #,pct_start=0.0)
# -
lr = 1e-3 #2
n_cycle = 10
# +
csvlogger = callbacks.CSVLogger(learn,'learn_log_3')
reduceLR = callbacks.ReduceLROnPlateauCallback(learn=learn, monitor = 'valid_loss', mode = 'auto', patience = 2, factor = 0.2, min_delta = 0)
#learn.fit(1,1e-4) #slice(lr)
learn.fit_one_cycle(n_cycle, slice(lr), callbacks=[reduceLR, MixUpCallback(learn), csvlogger]) #,pct_start=0.0)
# -
learn.recorder.plot_losses()
learn.recorder.plot_lr()
learn.recorder.plot_metrics()
learn.save('mymod_10E_SE_mix_50d_Inv_head0_2cos')
learn.export('export_mymod_10E_SE_mix_50d_Inv_head0_2cos.pkl')
# # Pred train and valid
# https://github.com/fastai/fastai/blob/master/fastai/basic_train.py#L370
def my_validate(model:nn.Module, dl:DataLoader, loss_func:OptLossFunc=None, cb_handler:Optional[CallbackHandler]=None,
pbar:Optional[PBar]=None, average=False, n_batch:Optional[int]=None)->Iterator[Tuple[Union[Tensor,int],...]]:
"Calculate `loss_func` of `model` on `dl` in evaluation mode."
model.eval()
with torch.no_grad():
val_losses,nums = [],[]
grapheme_root, vowel_diacritic, consonant_diacritic = torch.Tensor(),torch.Tensor(),torch.Tensor()# = [],[],[]
if cb_handler: cb_handler.set_dl(dl)
for xb,yb in progress_bar(dl, parent=pbar, leave=(pbar is not None)):
if cb_handler: xb, yb = cb_handler.on_batch_begin(xb, yb, train=False)
val_loss = loss_batch(model, xb, yb, loss_func, cb_handler=cb_handler)[0]
#val_losses.append(val_loss)
#print(val_loss[0].shape,val_loss[1].shape,val_loss[2].shape)
#grapheme_root.append(val_loss[0])
#vowel_diacritic.append(val_loss[1])
#consonant_diacritic.append(val_loss[2])
grapheme_root = torch.cat((grapheme_root,val_loss[0]))
vowel_diacritic = torch.cat((vowel_diacritic,val_loss[1]))
consonant_diacritic = torch.cat((consonant_diacritic,val_loss[2]))
if not is_listy(yb): yb = [yb]
nums.append(first_el(yb).shape[0])
if cb_handler and cb_handler.on_batch_end(grapheme_root[-1]): break
#if cb_handler and cb_handler.on_batch_end(val_losses[-1]): break
if n_batch and (len(nums)>=n_batch): break
nums = np.array(nums, dtype=np.float32)
#if average: return (to_np(torch.stack(val_losses)) * nums).sum() / nums.sum()
#else:
return grapheme_root, vowel_diacritic, consonant_diacritic
# +
output = pd.DataFrame()
'''test_data = (piil#
#.split_by_rand_pct(0.2)
#.label_from_df(cols='tags')#,label_delim=' ')
.split_from_df(col = valid_col)
.label_from_df(cols=['grapheme_root','vowel_diacritic','consonant_diacritic'])
.transform(tfms, padding_mode='zeros')#, size=224)#([cutout(n_holes=(1,4), length=(16, 16), p=.5),rand_zoom(scale=(.8,1))],[]), padding_mode='zeros')#tfms) #, size=128)
.databunch(bs=bs).normalize(imagenet_stats)) #mnist_stats
'''
learn.data.add_test(piil)
learn=learn.to_fp32()
grapheme_root, vowel_diacritic, consonant_diacritic = my_validate(learn.model, learn.dl(ds_type=DatasetType.Test))
df_g = pd.DataFrame({'image_id': df_train['image_id'], 'grapheme_root': grapheme_root.numpy().argmax(axis=1)})
df_v = pd.DataFrame({'image_id': df_train['image_id'], 'vowel_diacritic': vowel_diacritic.numpy().argmax(axis=1)})
df_c = pd.DataFrame({'image_id': df_train['image_id'], 'consonant_diacritic': consonant_diacritic.numpy().argmax(axis=1)})
#if i==0:
# output = (df_g.append(df_v).append(df_c)).sort_values(by=['rn','row_id'])
#else:
#output = output.append((df_g.append(df_v).append(df_c)).sort_values(by=['rn','row_id']))
output = df_g.merge(df_v.merge(df_c,on='image_id'), on='image_id')
#del df_g
#del df_c
#del df_v
#del learn
del grapheme_root
del vowel_diacritic
del consonant_diacritic
gc.collect()
#del piil
gc.collect()
# -
output.head(20)
output.to_csv('mod2_train_inference', index=False)
temp = pd.read_csv('mod2_train_inference')
temp
# +
import numpy as np
import sklearn.metrics
# +
rs1=sklearn.metrics.recall_score(df_label[['grapheme_root']], output[['grapheme_root']], average='macro')
rs2=sklearn.metrics.recall_score(df_label[['vowel_diacritic']], output[['vowel_diacritic']], average='macro')
rs3=sklearn.metrics.recall_score(df_label[['consonant_diacritic']], output[['consonant_diacritic']], average='macro')
mtrcs = [rs1, rs2, rs3, np.average([rs1, rs2, rs3], weights=[2,1,1])]
# -
df_m = pd.DataFrame(mtrcs)
df_m.to_csv("mtrcs.csv")
# # xx
# +
#https://github.com/fastai/fastai/blob/master/fastai/basic_train.py#L370
def my_predict(self,item:ItemBase, return_x:bool=False, batch_first:bool=True, with_dropout:bool=False, **kwargs):
"Return predicted class, label and probabilities for `item`."
batch = self.data.one_item(item)
res = self.pred_batch(batch=batch, with_dropout=with_dropout)
raw_pred,x = grab_idx(res,0,batch_first=batch_first),batch[0]
norm = getattr(self.data,'norm',False)
if norm:
x = self.data.denorm(x)
if norm.keywords.get('do_y',False): raw_pred = self.data.denorm(raw_pred)
ds = self.data.single_ds
pred = raw_pred #ds.y.analyze_pred(raw_pred, **kwargs)
x = ds.x.reconstruct(grab_idx(x, 0))
# y = ds.y.reconstruct(pred, x) if has_arg(ds.y.reconstruct, 'x') else ds.y.reconstruct(pred)
return (x, y, pred, raw_pred) if return_x else (y, pred, raw_pred)
#return raw_pred, x
a,b,r = my_predict(learn,learn.data.train_ds[2][0])
r[0].argmax(-1).cpu(),r[1].argmax(-1).cpu(),r[2].argmax(-1).cpu(),a,b
# -
# https://github.com/fastai/fastai/blob/master/fastai/basic_train.py#L370
def my_validate(model:nn.Module, dl:DataLoader, loss_func:OptLossFunc=None, cb_handler:Optional[CallbackHandler]=None,
pbar:Optional[PBar]=None, average=False, n_batch:Optional[int]=None)->Iterator[Tuple[Union[Tensor,int],...]]:
"Calculate `loss_func` of `model` on `dl` in evaluation mode."
model.eval()
with torch.no_grad():
val_losses,nums = [],[]
grapheme_root, vowel_diacritic, consonant_diacritic = torch.Tensor(),torch.Tensor(),torch.Tensor()# = [],[],[]
if cb_handler: cb_handler.set_dl(dl)
for xb,yb in progress_bar(dl, parent=pbar, leave=(pbar is not None)):
if cb_handler: xb, yb = cb_handler.on_batch_begin(xb, yb, train=False)
val_loss = loss_batch(model, xb, yb, loss_func, cb_handler=cb_handler)[0]
#val_losses.append(val_loss)
#print(val_loss[0].shape,val_loss[1].shape,val_loss[2].shape)
#grapheme_root.append(val_loss[0])
#vowel_diacritic.append(val_loss[1])
#consonant_diacritic.append(val_loss[2])
grapheme_root = torch.cat((grapheme_root,val_loss[0]))
vowel_diacritic = torch.cat((vowel_diacritic,val_loss[1]))
consonant_diacritic = torch.cat((consonant_diacritic,val_loss[2]))
if not is_listy(yb): yb = [yb]
nums.append(first_el(yb).shape[0])
if cb_handler and cb_handler.on_batch_end(grapheme_root[-1]): break
#if cb_handler and cb_handler.on_batch_end(val_losses[-1]): break
if n_batch and (len(nums)>=n_batch): break
nums = np.array(nums, dtype=np.float32)
#if average: return (to_np(torch.stack(val_losses)) * nums).sum() / nums.sum()
#else:
return grapheme_root, vowel_diacritic, consonant_diacritic
#https://github.com/fastai/fastai/blob/master/fastai/basic_train.py
def my_get_preds(model:nn.Module, dl:DataLoader, pbar:Optional[PBar]=None, cb_handler:Optional[CallbackHandler]=None,
activ:nn.Module=None, loss_func:OptLossFunc=None, n_batch:Optional[int]=None) -> List[Tensor]:
"Tuple of predictions and targets, and optional losses (if `loss_func`) using `dl`, max batches `n_batch`."
res = [to_float(torch.cat(o).cpu()) for o in
zip(*my_validate(model, dl, cb_handler=cb_handler, pbar=pbar, average=False, n_batch=n_batch))]
if loss_func is not None:
with NoneReduceOnCPU(loss_func) as lf: res.append(lf(res[0], res[1]))
if activ is not None: res[0] = activ(res[0])
return res
def my_learner_get_preds(self=learn, ds_type:DatasetType=DatasetType.Valid, activ:nn.Module=None,
with_loss:bool=False, n_batch:Optional[int]=None, pbar:Optional[PBar]=None) -> List[Tensor]:
"Return predictions and targets on `ds_type` dataset."
#lf = self.loss_func if with_loss else None
#activ = ifnone(activ, _loss_func2activ(self.loss_func))
print('yahoo')
if not getattr(self, 'opt', False): self.create_opt(defaults.lr, self.wd)
callbacks = [cb(self) for cb in self.callback_fns + listify(defaults.extra_callback_fns)] + listify(self.callbacks)
return my_get_preds(self.model, self.dl(ds_type), cb_handler=CallbackHandler(callbacks),
activ=activ, loss_func=None, n_batch=n_batch, pbar=pbar)
# +
# mokey patching
setattr(learn, "get_preds", my_learner_get_preds)
# -
interp = ClassificationInterpretation.from_learner(learn)
# %debug
interp.most_confused(min_val=4)
# confusion matrix
display(interp.plot_confusion_matrix())
| notes/backups/bck_Bengaliai_CV19-Mod2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 (tensorflow)
# language: python
# name: py36
# ---
# !pip install -q tf-models-official==2.3.0
# +
import os
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
import tensorflow_hub as hub
os.environ["TFHUB_CACHE_DIR"] = "gs://nts2020-tpu"
from official import nlp
from official.modeling import tf_utils
from official.nlp import bert
# Load the required submodules
import official.nlp.optimization
import official.nlp.bert.bert_models
import official.nlp.bert.configs
import official.nlp.bert.run_classifier
import official.nlp.bert.tokenization
import official.nlp.data.classifier_data_lib
import official.nlp.modeling.losses
import official.nlp.modeling.models
import official.nlp.modeling.networks
import json
# -
try:
tpu = tf.distribute.cluster_resolver.TPUClusterResolver(tpu='tpu-quickstart')
tf.config.experimental_connect_to_cluster(tpu)
tf.tpu.experimental.initialize_tpu_system(tpu)# TPU detection
print('Running on TPU ', tpu.cluster_spec().as_dict()['worker'])
except ValueError:
raise BaseException('ERROR: Not connected to a TPU runtime; please see the previous cell in this notebook for instructions!')
tf.__version__
def single_file_dataset(input_file, name_to_features, num_samples=None):
"""Creates a single-file dataset to be passed for BERT custom training."""
# For training, we want a lot of parallel reading and shuffling.
# For eval, we want no shuffling and parallel reading doesn't matter.
d = tf.data.TFRecordDataset(input_file)
if num_samples:
d = d.take(num_samples)
d = d.map(
lambda record: decode_record(record, name_to_features),
num_parallel_calls=tf.data.experimental.AUTOTUNE)
# When `input_file` is a path to a single file or a list
# containing a single path, disable auto sharding so that
# same input file is sent to all workers.
if isinstance(input_file, str) or len(input_file) == 1:
options = tf.data.Options()
options.experimental_distribute.auto_shard_policy = (
tf.data.experimental.AutoShardPolicy.OFF)
d = d.with_options(options)
return d
def decode_record(record, name_to_features):
"""Decodes a record to a TensorFlow example."""
example = tf.io.parse_single_example(record, name_to_features)
# tf.Example only supports tf.int64, but the TPU only supports tf.int32.
# So cast all int64 to int32.
for name in list(example.keys()):
t = example[name]
if t.dtype == tf.int64:
t = tf.cast(t, tf.int32)
example[name] = t
return example
def create_classifier_dataset(file_path,
seq_length,
batch_size,
task_id,
is_training=True,
input_pipeline_context=None,
label_type=tf.int64,
lang_id = 0,
include_sample_weights=False,
num_samples=None):
"""Creates input dataset from (tf)records files for train/eval."""
name_to_features = {
'input_ids': tf.io.FixedLenFeature([seq_length], tf.int64),
'input_mask': tf.io.FixedLenFeature([seq_length], tf.int64),
'segment_ids': tf.io.FixedLenFeature([seq_length], tf.int64),
'label_ids': tf.io.FixedLenFeature([], label_type),
}
if include_sample_weights:
name_to_features['weight'] = tf.io.FixedLenFeature([], tf.float32)
dataset = single_file_dataset(file_path, name_to_features,
num_samples=num_samples)
# The dataset is always sharded by number of hosts.
# num_input_pipelines is the number of hosts rather than number of cores.
if input_pipeline_context and input_pipeline_context.num_input_pipelines > 1:
dataset = dataset.shard(input_pipeline_context.num_input_pipelines,
input_pipeline_context.input_pipeline_id)
def _select_data_from_record(record):
x = {
'input_word_ids': record['input_ids'],
'input_mask': record['input_mask'],
'input_type_ids': record['segment_ids'],
'lang_id' : lang_id
}
#pdb.set_trace()
y = record['label_ids']
if include_sample_weights:
w = record['weight']
return (x, y, w)
default = tf.constant(-1, dtype=tf.int32)
if task_id ==0:
return (x, (y, default))
if task_id == 1:
return (x, (default,y))
if is_training:
dataset = dataset.shuffle(100)
dataset = dataset.repeat()
dataset = dataset.map(
_select_data_from_record,
num_parallel_calls=tf.data.experimental.AUTOTUNE)
#dataset = dataset.batch(batch_size, drop_remainder=is_training)
dataset = dataset.prefetch(tf.data.experimental.AUTOTUNE)
return dataset
# +
config_dict = {
"attention_probs_dropout_prob": 0.1,
"directionality": "bidi",
"hidden_act": "gelu",
"hidden_dropout_prob": 0.1,
"hidden_size": 768,
"initializer_range": 0.02,
"intermediate_size": 3072,
"max_position_embeddings": 512,
"num_attention_heads": 12,
"num_hidden_layers": 12,
"pooler_fc_size": 768,
"pooler_num_attention_heads": 12,
"pooler_num_fc_layers": 3,
"pooler_size_per_head": 128,
"pooler_type": "first_token_transform",
"type_vocab_size": 2,
"vocab_size": 119547
}
bert_config = bert.configs.BertConfig.from_dict(config_dict)
# +
tf_records_filenames = ["gs://nts2020/xtereme/pawsx/train.en.tfrecords", "gs://nts2020/xtereme/xnli/train.en.tfrecords"]
sampling_factor = []
for fn in tf_records_filenames:
c = 0
for record in tf.compat.v1.python_io.tf_record_iterator(fn):
c += 1
sampling_factor.append(c)
print(c)
c = sum(sampling_factor)
for i in range(0, len(sampling_factor)):
sampling_factor[i] = sampling_factor[i]/c
sampling_factor
# -
tf_records_filenames = ["gs://nts2020/xtreme/translate_train/train.ar.tfrecords", "gs://nts2020/xtreme/translate_train/train.bg.tfrecords", "gs://nts2020/xtreme/translate_train/train.de.tfrecords",
"gs://nts2020/xtreme/translate_train/train.el.tfrecords","gs://nts2020/xtreme/translate_train/train.es.tfrecords","gs://nts2020/xtreme/translate_train/train.fr.tfrecords",
"gs://nts2020/xtreme/translate_train/train.hi.tfrecords","gs://nts2020/xtreme/translate_train/train.ru.tfrecords","gs://nts2020/xtreme/translate_train/train.sw.tfrecords",
"gs://nts2020/xtreme/translate_train/train.th.tfrecords","gs://nts2020/xtreme/translate_train/train.tr.tfrecords","gs://nts2020/xtreme/translate_train/train.ur.tfrecords",
"gs://nts2020/xtreme/translate_train/train.vi.tfrecords","gs://nts2020/xtreme/translate_train/train.zh.tfrecords"]
# +
tf_records_filenames = ["gs://nts2020/xtreme/translate_train/train.ar.tfrecords", "gs://nts2020/xtreme/translate_train/train.bg.tfrecords", "gs://nts2020/xtreme/translate_train/train.de.tfrecords",
"gs://nts2020/xtreme/translate_train/train.el.tfrecords","gs://nts2020/xtreme/translate_train/train.es.tfrecords","gs://nts2020/xtreme/translate_train/train.fr.tfrecords",
"gs://nts2020/xtreme/translate_train/train.hi.tfrecords","gs://nts2020/xtreme/translate_train/train.ru.tfrecords","gs://nts2020/xtreme/translate_train/train.sw.tfrecords",
"gs://nts2020/xtreme/translate_train/train.th.tfrecords","gs://nts2020/xtreme/translate_train/train.tr.tfrecords","gs://nts2020/xtreme/translate_train/train.ur.tfrecords",
"gs://nts2020/xtreme/translate_train/train.vi.tfrecords","gs://nts2020/xtreme/translate_train/train.zh.tfrecords"]
other_langs_sampling_factor = []
for fn in tf_records_filenames:
c = 0
for record in tf.compat.v1.python_io.tf_record_iterator(fn):
c += 1
other_langs_sampling_factor.append(c)
print(c)
c = sum(other_langs_sampling_factor)
for i in range(0, len(other_langs_sampling_factor)):
other_langs_sampling_factor[i] = other_langs_sampling_factor[i]/c
other_langs_sampling_factor
# -
other_lang_count = len(tf_records_filenames)
other_lang_aggregate_weight = 0.9
train_sampling_factor = []
for i in sampling_factor:
train_sampling_factor.append((i* (1-other_lang_aggregate_weight))/ sum(sampling_factor))
for i in other_langs_sampling_factor:
train_sampling_factor.append((i * other_lang_aggregate_weight))
train_sampling_factor[0] = 0.03
train_sampling_factor = [0.03,
0.08882590708500053,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428]
if sum(train_sampling_factor)!=1:
train_sampling_factor[1]+= 1- sum(train_sampling_factor)
def _loss_with_filter(y_true, y_pred):
num_labels = y_pred.get_shape().as_list()[-1]
log_probs = tf.nn.log_softmax(y_pred, axis=-1)
log_probs = tf.reshape(log_probs, [-1, num_labels])
labels = tf.reshape(y_true, [-1])
one_hot_labels = tf.one_hot(labels, depth=num_labels, dtype=tf.float32)
per_example_loss = -tf.reduce_sum(one_hot_labels * log_probs, axis=-1)
loss = tf.reduce_mean(per_example_loss)
return loss
import tensorflow.keras.backend as K
def accuracy_mod(y_true, y_pred):
# Squeeze the shape to (None, ) from (None, 1) as we want to apply operations directly on y_true
if K.ndim(y_true) == K.ndim(y_pred):
y_true = K.squeeze(y_true, -1)
# Normalize the y_pred values first and then take the arg at which we have a maximum value (This is the predicted label)
y_pred = K.softmax(y_pred, axis = -1)
y_pred = K.argmax(y_pred, axis = -1)
# Since the ground labels can also have -1s for which we don't wanna calculate accuracy, we are filtering them off
defa = K.constant([0], dtype=tf.float32)
#Creating a boolean tensor for labels greater or equal to 0
is_valid = K.greater_equal(y_true, defa)
#Get the corresponding indices
indices = tf.where(is_valid)
#Gather the results of y_true and y_pred at the indices we calculated above
fil_y_true = K.gather(y_true, K.reshape(indices, [-1]))
fil_y_pred = K.gather(y_pred, K.reshape(indices, [-1]))
# K.print_tensor(res, message='res = ')
# K.print_tensor(comp, message='comp = ')
fil_y_true = K.cast(fil_y_true, K.floatx())
fil_y_pred = K.cast(fil_y_pred, K.floatx())
#pdb.set_trace()
return K.cast(K.equal(fil_y_true, fil_y_pred), K.floatx())
epochs = 3
batch_size = 64
eval_batch_size = 64
max_seq_length = 128
# +
paws_training_dataset = create_classifier_dataset(
"gs://nts2020/xtereme/pawsx/train.en.tfrecords",
128,
batch_size,
task_id = 0,
is_training=True)
xnli_training_dataset = create_classifier_dataset(
"gs://nts2020/xtereme/xnli/train.en.tfrecords",
128,
batch_size,
task_id =1,
is_training=True)
paws_eval_dataset = create_classifier_dataset(
"gs://nts2020/xtereme/pawsx/eval.en.tfrecords",
128,
batch_size,
task_id = 0,
is_training=False)
xnli_eval_dataset = create_classifier_dataset(
"gs://nts2020/xtereme/xnli/eval.en.tfrecords",
128,
batch_size,
task_id = 1,
is_training=False)
# -
tf_records_filenames = ["gs://nts2020/xtreme/translate_train/train.ar.tfrecords", "gs://nts2020/xtreme/translate_train/train.bg.tfrecords", "gs://nts2020/xtreme/translate_train/train.de.tfrecords",
"gs://nts2020/xtreme/translate_train/train.el.tfrecords","gs://nts2020/xtreme/translate_train/train.es.tfrecords","gs://nts2020/xtreme/translate_train/train.fr.tfrecords",
"gs://nts2020/xtreme/translate_train/train.hi.tfrecords","gs://nts2020/xtreme/translate_train/train.ru.tfrecords","gs://nts2020/xtreme/translate_train/train.sw.tfrecords",
"gs://nts2020/xtreme/translate_train/train.th.tfrecords","gs://nts2020/xtreme/translate_train/train.tr.tfrecords","gs://nts2020/xtreme/translate_train/train.ur.tfrecords",
"gs://nts2020/xtreme/translate_train/train.vi.tfrecords","gs://nts2020/xtreme/translate_train/train.zh.tfrecords"]
xnli_ar_training_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/translate_train/train.ar.tfrecords",
128,
batch_size,
task_id =1,
lang_id =1,
is_training=True)
xnli_bg_training_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/translate_train/train.bg.tfrecords",
128,
batch_size,
task_id =1,
lang_id =2,
is_training=True)
xnli_de_training_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/translate_train/train.de.tfrecords",
128,
batch_size,
task_id =1,
lang_id =3,
is_training=True)
xnli_el_training_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/translate_train/train.el.tfrecords",
128,
batch_size,
task_id =1,
lang_id =4,
is_training=True)
xnli_es_training_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/translate_train/train.es.tfrecords",
128,
batch_size,
task_id =1,
lang_id =5,
is_training=True)
xnli_fr_training_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/translate_train/train.fr.tfrecords",
128,
batch_size,
task_id =1,
lang_id =6,
is_training=True)
xnli_hi_training_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/translate_train/train.hi.tfrecords",
128,
batch_size,
task_id =1,
lang_id =7,
is_training=True)
xnli_ru_training_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/translate_train/train.ru.tfrecords",
128,
batch_size,
task_id =1,
lang_id =8,
is_training=True)
xnli_sw_training_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/translate_train/train.sw.tfrecords",
128,
batch_size,
task_id =1,
lang_id =9,
is_training=True)
xnli_th_training_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/translate_train/train.th.tfrecords",
128,
batch_size,
task_id =1,
lang_id =10,
is_training=True)
xnli_tr_training_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/translate_train/train.tr.tfrecords",
128,
batch_size,
task_id =1,
lang_id =11,
is_training=True)
xnli_ur_training_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/translate_train/train.ur.tfrecords",
128,
batch_size,
task_id =1,
lang_id =12,
is_training=True)
xnli_vi_training_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/translate_train/train.vi.tfrecords",
128,
batch_size,
task_id =1,
lang_id =13,
is_training=True)
xnli_zh_training_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/translate_train/train.zh.tfrecords",
128,
batch_size,
task_id =1,
lang_id =14,
is_training=True)
xnli_ar_eval_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/xnli_w_dev/eval_ar.tfrecords",
max_seq_length,
batch_size,
task_id =1,
is_training=False)
xnli_bg_eval_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/xnli_w_dev/eval_bg.tfrecords",
max_seq_length,
batch_size,
task_id =1,
is_training=False)
xnli_de_eval_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/xnli_w_dev/eval_de.tfrecords",
max_seq_length,
batch_size,
task_id =1,
is_training=False)
xnli_el_eval_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/xnli_w_dev/eval_el.tfrecords",
max_seq_length,
batch_size,
task_id =1,
is_training=False)
xnli_es_eval_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/xnli_w_dev/eval_es.tfrecords",
max_seq_length,
batch_size,
task_id =1,
is_training=False)
xnli_fr_eval_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/xnli_w_dev/eval_fr.tfrecords",
max_seq_length,
batch_size,
task_id =1,
is_training=False)
xnli_hi_eval_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/xnli_w_dev/eval_hi.tfrecords",
max_seq_length,
batch_size,
task_id =1,
is_training=False)
xnli_ru_eval_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/xnli_w_dev/eval_ru.tfrecords",
max_seq_length,
batch_size,
task_id =1,
is_training=False)
xnli_sw_eval_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/xnli_w_dev/eval_sw.tfrecords",
max_seq_length,
batch_size,
task_id =1,
is_training=False)
xnli_th_eval_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/xnli_w_dev/eval_th.tfrecords",
max_seq_length,
batch_size,
task_id =1,
is_training=False)
xnli_tr_eval_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/xnli_w_dev/eval_tr.tfrecords",
max_seq_length,
batch_size,
task_id =1,
is_training=False)
xnli_ur_eval_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/xnli_w_dev/eval_ur.tfrecords",
max_seq_length,
batch_size,
task_id =1,
is_training=False)
xnli_vi_eval_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/xnli_w_dev/eval_vi.tfrecords",
max_seq_length,
batch_size,
task_id =1,
is_training=False)
xnli_zh_eval_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/xnli_w_dev/eval_zh.tfrecords",
max_seq_length,
batch_size,
task_id =1,
is_training=False)
# +
delta = 0.50
for i in range(0, len(train_sampling_factor)):
train_sampling_factor[i] = train_sampling_factor[i] * (1 - delta)
train_sampling_factor[13] += delta
# -
if sum(train_sampling_factor)!=1:
train_sampling_factor[1]+= 1- sum(train_sampling_factor)
# +
train_sampling_factor = 16 * [None]
train_sampling_factor[0] = 0.01
train_sampling_factor[1:] = [0.54957585, 0.00897124, 0.00887785, 0.00799495, 0.0105019 ,
0.00910511, 0.00809789, 0.33139596, 0.00878845, 0.0100313 ,
0.00971826, 0.00991482, 0.00882816, 0.00819859, 0.00999967]
train_sampling_factor[1] -= 0.04
if sum(train_sampling_factor)!=1:
train_sampling_factor[1]+= 1- sum(train_sampling_factor)
# -
train_sampling_factor = [0.03,
0.30,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.50,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428]
if sum(train_sampling_factor)!=1:
train_sampling_factor[1]+= 1- sum(train_sampling_factor)
training_dataset = tf.data.experimental.sample_from_datasets(
[paws_training_dataset, xnli_training_dataset, xnli_ar_training_dataset, xnli_bg_training_dataset, xnli_de_training_dataset, xnli_el_training_dataset, xnli_es_training_dataset,
xnli_fr_training_dataset, xnli_hi_training_dataset, xnli_ru_training_dataset, xnli_sw_training_dataset, xnli_th_training_dataset,
xnli_tr_training_dataset, xnli_ur_training_dataset, xnli_vi_training_dataset, xnli_zh_training_dataset], weights=tf.constant([train_sampling_factor[0], train_sampling_factor[1],
train_sampling_factor[2], train_sampling_factor[3],
train_sampling_factor[4],train_sampling_factor[5],
train_sampling_factor[6],train_sampling_factor[7],
train_sampling_factor[8],train_sampling_factor[9],
train_sampling_factor[10],train_sampling_factor[11],
train_sampling_factor[12],train_sampling_factor[13],
train_sampling_factor[14],train_sampling_factor[15]]))
# +
# iterator = training_dataset.as_numpy_iterator()
# -
# training_dataset = tf.data.experimental.sample_from_datasets(
# [paws_training_dataset, xnli_sw_training_dataset], weights = tf.constant([0.5,0.5]))
evaluation_sampling_factor = [0.3 , 0.7]
# evaluation_dataset = tf.data.experimental.sample_from_datasets(
# [paws_eval_dataset, xnli_eval_dataset, xnli_ar_eval_dataset, xnli_bg_eval_dataset, xnli_de_eval_dataset, xnli_el_eval_dataset, xnli_es_eval_dataset,
# xnli_fr_eval_dataset, xnli_hi_eval_dataset, xnli_ru_eval_dataset, xnli_sw_eval_dataset, xnli_th_eval_dataset,
# xnli_tr_eval_dataset, xnli_ur_eval_dataset, xnli_vi_eval_dataset, xnli_zh_eval_dataset], weights=tf.constant([train_sampling_factor[0], train_sampling_factor[1],
# train_sampling_factor[2], train_sampling_factor[3],
# train_sampling_factor[4],train_sampling_factor[5],
# train_sampling_factor[6],train_sampling_factor[7],
# train_sampling_factor[8],train_sampling_factor[9],
# train_sampling_factor[10],train_sampling_factor[11],
# train_sampling_factor[12],train_sampling_factor[13],
# train_sampling_factor[14],train_sampling_factor[15]]))
evaluation_dataset = tf.data.experimental.sample_from_datasets(
[paws_eval_dataset,xnli_hi_eval_dataset], weights=tf.constant([evaluation_sampling_factor[0],evaluation_sampling_factor[1]]))
# +
# evaluation_dataset = tf.data.experimental.sample_from_datasets(
# [paws_eval_dataset, xnli_eval_dataset], weights=tf.constant([sampling_factor[0], sampling_factor[1]])
# )
# -
class myCallback(tf.keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs={}):
if(logs.get('val_output1_accuracy_mod') > 0.71):
print("\nWe have reached %2.2f%% accuracy, so we will stopping training." %(acc_thresh*100))
self.model.stop_training = True
callbacks = myCallback()
# +
# resolver = tf.distribute.cluster_resolver.TPUClusterResolver.connect(tpu='tpu-quickstart', project = 'moana-intern-fall-2020')
# tf.config.experimental_connect_to_cluster(resolver)
# tf.tpu.experimental.initialize_tpu_system(resolver)
# strategy = tf.distribute.TPUStrategy(resolver)
strategy = tf.distribute.TPUStrategy(tpu)
with strategy.scope():
max_seq_length = 128
initializer = tf.keras.initializers.TruncatedNormal(
stddev=bert_config.initializer_range)
bert_encoder = bert.bert_models.get_transformer_encoder(
bert_config, max_seq_length)
input_word_ids = tf.keras.layers.Input(
shape=(max_seq_length,), dtype=tf.int32, name='input_word_ids')
input_mask = tf.keras.layers.Input(
shape=(max_seq_length,), dtype=tf.int32, name='input_mask')
input_type_ids = tf.keras.layers.Input(
shape=(max_seq_length,), dtype=tf.int32, name='input_type_ids')
bert_model = hub.KerasLayer("https://tfhub.dev/tensorflow/bert_multi_cased_L-12_H-768_A-12/2",
trainable=True)
#bert_model = hub.KerasLayer(hub_url_bert, trainable=True)
pooled_output, seq_output = bert_model([input_word_ids, input_mask, input_type_ids])
output1 = tf.keras.layers.Dropout(rate=bert_config.hidden_dropout_prob)(
pooled_output)
output1 = tf.keras.layers.Dense(
2, kernel_initializer=initializer, name='output1')(
output1)
output2 = tf.keras.layers.Dropout(rate=bert_config.hidden_dropout_prob)(
pooled_output)
output2 = tf.keras.layers.Dense(
3, kernel_initializer=initializer, name='output2')(
output2)
model = tf.keras.Model(
inputs={
'input_word_ids': input_word_ids,
'input_mask': input_mask,
'input_type_ids': input_type_ids
},
outputs=[output1, output2])
# Set up epochs and steps
# get train_data_size from metadata
train_data_size = c
steps_per_epoch = int(train_data_size / batch_size)
num_train_steps = steps_per_epoch * epochs
warmup_steps = int(epochs * train_data_size * 0.1 / batch_size)
# creates an optimizer with learning rate schedule
optimizer = nlp.optimization.create_optimizer(
2e-5, num_train_steps=num_train_steps, num_warmup_steps=warmup_steps)
training_dataset = training_dataset.batch(batch_size)
evaluation_dataset = evaluation_dataset.batch(batch_size, drop_remainder=True)
model.compile(optimizer = optimizer, loss = [_loss_with_filter, _loss_with_filter], metrics = [accuracy_mod])
history = model.fit(training_dataset, batch_size = batch_size, epochs= 35, steps_per_epoch = 1000, validation_data=evaluation_dataset, callbacks = [callbacks])
# -
xnli_hi_test_dataset = create_classifier_dataset(
"gs://nts2020/xtreme/xnli_w_dev/test_hi.tf_record",
max_seq_length,
batch_size,
task_id =1,
is_training=False)
xnli_hi_test_dataset = xnli_hi_test_dataset.batch(batch_size, drop_remainder = True)
model.evaluate(xnli_hi_test_dataset)
iterat = xnli_hi_test_dataset.as_numpy_iterator()
iterat.next()
sw_impulse = create_classifier_dataset(
"gs://nts2020/xtreme/xnli_w_dev/eval_sw.tfrecords",
max_seq_length,
batch_size,
task_id =1,
is_training=False)
sw_impulse = sw_impulse.batch(batch_size)
iterat = sw_impulse.as_numpy_iterator()
iterat.next()
# +
strategy = tf.distribute.TPUStrategy(tpu)
train_data_size = c
steps_per_epoch = int(train_data_size / batch_size)
num_train_steps = steps_per_epoch * epochs
warmup_steps = int(epochs * train_data_size * 0.1 / batch_size)
# creates an optimizer with learning rate schedule
optimizer = nlp.optimization.create_optimizer(
2e-5, num_train_steps=num_train_steps, num_warmup_steps=warmup_steps)
# -
with strategy.scope():
model.save('gs://nts2020/XNLI_SW_uniform_10epochs_tf',save_format='tf')
with strategy.scope():
loaded_model = tf.keras.models.load_model('gs://nts2020/XNLI_SW_uniform_10epochs_tf',
compile=False,custom_objects={"accuracy_mod": accuracy_mod})
# +
delta = 0.20
for i in range(0, len(train_sampling_factor)):
train_sampling_factor[i] = train_sampling_factor[i] * (1 - delta)
train_sampling_factor[1] += delta
# -
if sum(train_sampling_factor)!=1:
train_sampling_factor[1]+= 1- sum(train_sampling_factor)
train_sampling_factor = [0.03,
0.08882590708500053,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428,
0.06428571428571428]
train_sampling_factor
new_weights = get_new_static_weights(0.66, tf.Variable(train_sampling_factor[1:]))
new_weights.numpy()
new_weights.numpy()
new_weights.numpy()
new_weights.numpy()
new_weights.numpy()
new_weights.numpy()
train_sampling_factor = 15 * [None]
train_sampling_factor[0] = 0.01
train_sampling_factor[1:] = new_weights.numpy()
# train_sampling_factor[1:] = [0.49758378, 0.01234279, 0.01297586, 0.01206473, 0.0130334 ,
# 0.01228484, 0.01269346, 0.0109312 , 0.01205256, 0.01229929,
# 0.34325856, 0.01188709, 0.01400499, 0.01077152, 0.01181594]
# train_sampling_factor[1] -= 0.04
if sum(train_sampling_factor)!=1:
train_sampling_factor[1]+= 1- sum(train_sampling_factor)
training_dataset = tf.data.experimental.sample_from_datasets(
[paws_training_dataset, xnli_training_dataset, xnli_ar_training_dataset, xnli_bg_training_dataset, xnli_de_training_dataset, xnli_el_training_dataset, xnli_es_training_dataset,
xnli_fr_training_dataset, xnli_hi_training_dataset, xnli_ru_training_dataset, xnli_sw_training_dataset, xnli_th_training_dataset,
xnli_tr_training_dataset, xnli_ur_training_dataset, xnli_vi_training_dataset, xnli_zh_training_dataset], weights=tf.constant([train_sampling_factor[0], train_sampling_factor[1],
train_sampling_factor[2], train_sampling_factor[3],
train_sampling_factor[4],train_sampling_factor[5],
train_sampling_factor[6],train_sampling_factor[7],
train_sampling_factor[8],train_sampling_factor[9],
train_sampling_factor[10],train_sampling_factor[11],
train_sampling_factor[12],train_sampling_factor[13],
train_sampling_factor[14],train_sampling_factor[15]]))
untouched_dataset = training_dataset
training_dataset = training_dataset.batch(batch_size)
evaluation_dataset = tf.data.experimental.sample_from_datasets(
[paws_eval_dataset,xnli_hi_eval_dataset], weights=tf.constant([evaluation_sampling_factor[0],evaluation_sampling_factor[1]]))
evaluation_dataset = evaluation_dataset.batch(batch_size, drop_remainder=True)
model.fit(training_dataset, batch_size = batch_size, epochs= 15, steps_per_epoch = 1000, validation_data=evaluation_dataset)
loaded_model.compile(optimizer = optimizer, loss = [_loss_with_filter, _loss_with_filter], metrics = [accuracy_mod])
loaded_model.evaluate(evaluation_dataset)
loaded_model.fit(training_dataset, batch_size = batch_size, epochs= 2, steps_per_epoch = 1000, validation_data=evaluation_dataset)
loaded_model.fit(training_dataset, batch_size = batch_size, epochs= 8, steps_per_epoch = 1000, validation_data=evaluation_dataset)
itera = None
rl_dataset = untouched_dataset#.batch(batch_size)
rl_dataset = rl_dataset.batch(32)
itera = rl_dataset.as_numpy_iterator()
from collections import defaultdict
def get_batch_lang(iterator):
appearances = defaultdict(int)
for curr in iterator.next()[0]['lang_id']:
appearances[curr] += 1
batch_lang_count = 15 *[None]
for i in range(15):
batch_lang_count[i] = appearances[i]
return batch_lang_count
# # initializer = tf.keras.initializers.RandomUniform(minval=0., maxval=1.)
# # values = initializer(shape=(1, 15))
# phi = tf.Variable(train_sampling_factor[1:])
# +
# initializer = tf.keras.initializers.RandomUniform(minval=0., maxval=1.)
# input2 = initializer(shape=(1, 15))
# +
# tf.squeeze(tf.nn.softmax(input2, axis = -1))
# +
# input1 = phi
# -
import random
target_lang = 7
cce = tf.keras.losses.CategoricalCrossentropy()
d = 15
# +
# def train(opt, input1, batch_lang_count, R, loss):
# loss = 0
# with tf.GradientTape() as tape:
# tape.watch(input1)
# for i, val in enumerate(batch_lang_count):
# loss += val * R * cce(tf.one_hot(i, depth =d), tf.squeeze(tf.nn.softmax(input1, axis = -1)))
# gradients = tape.gradient(loss, input1)
# opt.apply_gradients(zip([gradients], [input1]))
# #print(loss)
# print(tf.nn.softmax(input1, axis = -1))
# return input1
# +
# def train(opt, input1, batch_lang_count, R, loss, priority, flag):
# loss = 0
# with tf.GradientTape() as tape:
# #tape.watch(input1)
# for i, val in enumerate(batch_lang_count):
# loss += val * (100 - R) * cce(tf.one_hot(i, depth =d), tf.squeeze(tf.nn.softmax(input1, axis = -1)))
# gradients = tape.gradient(loss, input1)
# #print(gradients)
# opt.apply_gradients(zip([gradients], [input1]))
# print(loss)
# print(tf.nn.softmax(input1, axis = -1))
# return input1
# -
opt = tf.keras.optimizers.SGD(learning_rate = 0.001,clipvalue= 0.001)
loss = 0
def get_new_static_weights(reward, input1):
epsilon = 0.05
epsilon2 = 0.50
acc = 0
# initializer = tf.keras.initializers.RandomUniform(minval=0., maxval=1.)
# input1 = tf.Variable(initializer(shape=(1, 15))[0])
#input1 = tf.Variable(train_sampling_factor[1:])
flag = 0
batch_data = get_batch_lang(itera)
for i in range(100):
draw = random.uniform(0.0, 1.0)
batch_data = get_batch_lang(itera)
if draw <= epsilon:
#print("a")
for i in range(15):
batch_data[i] = random.random()
#if draw > epsilon and draw < epsilon2:
#print("b")
# batch_data = get_batch_lang(itera)
if draw >= epsilon2:
#print("c")
ans = [i for i in range(0, len(batch_data))]
ind = list(set(ans).difference([0, target_lang]))
for i in ind:
batch_data[i] = 0
input1 = trainstep(opt, input1, batch_data, reward * 100, loss)
acc += tf.nn.softmax(input1, axis = -1)
return acc/100
# +
# initializer = tf.keras.initializers.RandomUniform(minval=0., maxval=1.)
# test1 = tf.Variable(initializer(shape=(1, 15))[0])
# +
# opt1 = tf.keras.optimizers.SGD(learning_rate = 0.001)
# loss = 0
# -
def trainstep(opt, input1, batch_lang_count, R, loss):
loss = 0
with tf.GradientTape() as tape:
#tape.watch(input1)
for i, val in enumerate(batch_lang_count):
loss += val * (R) * cce(tf.one_hot(i, depth =d), tf.squeeze(tf.nn.softmax(input1, axis = -1)))
#loss += cce(input1, tf.cast(batch_data, dtype= tf.float64))
print(tf.nn.softmax(input1, axis = -1))
gradients = tape.gradient(loss, input1)
#gradients, _ = tf.clip_by_global_norm(gradients, 5.0)
#print(gradients)
opt.apply_gradients(zip([gradients], [input1]))
return input1
#print(loss)
# +
# train_sampling_factor
# -
# for i in range(0, 100):
# #if i%10 == 0:
# batch_data = get_batch_lang(itera)
# test1 = trainstep(opt1, test1, batch_data, 0.4 * 100, loss)
# +
# batch_data
# +
# def get_new_static_weights(reward):
# refresh()
# epsilon = 0.05
# epsilon2 = 0.75
# initializer = tf.keras.initializers.RandomUniform(minval=0., maxval=1.)
# input1 = tf.Variable(initializer(shape=(1, 15))[0])
# flag = 0
# batch_data = get_batch_lang()
# for i in range(100):
# draw = random.uniform(0.0, 1.0)
# #batch_data = get_batch_lang()
# if draw <= epsilon:
# print("a")
# initializer = tf.keras.initializers.RandomUniform(minval=0., maxval=1.)
# input1 = tf.Variable(initializer(shape=(1, 15))[0])
# ans = [i for i in range(0, len(batch_data))]
# rind = list(set(ans).difference(random.sample(range(0, 15), 2)))
# for i in rind:
# batch_data[i] = 0
# if draw > epsilon and draw < epsilon2:
# print("b")
# phi = tf.Variable(train_sampling_factor[1:])
# input1 = phi
# ans = [i for i in range(0, len(batch_data))]
# find = list(set(ans).difference(random.sample(range(0, 15), 2)))
# for i in find:
# batch_data[i] = 0
# if draw >= epsilon2:
# print("c")
# ans = [i for i in range(0, len(batch_data))]
# ind = list(set(ans).difference([0, target_lang]))
# for i in ind:
# batch_data[i] = 0
# input1 = trainstep(opt, input1, batch_data, reward * 100, loss)
# return tf.nn.softmax(input1, axis = -1), max1, max2
# -
paws_batched_eval_data = paws_eval_dataset.batch(batch_size)
xnli_batched_eval_data = xnli_eval_dataset.batch(batch_size)
model.evaluate(paws_batched_eval_data)
model.evaluate(xnli_batched_eval_data)
xnli_ar_eval_dataset = xnli_ar_eval_dataset.batch(batch_size)
xnli_bg_eval_dataset = xnli_bg_eval_dataset.batch(batch_size)
xnli_de_eval_dataset = xnli_de_eval_dataset.batch(batch_size)
xnli_el_eval_dataset = xnli_el_eval_dataset.batch(batch_size)
xnli_es_eval_dataset = xnli_es_eval_dataset.batch(batch_size)
xnli_fr_eval_dataset = xnli_fr_eval_dataset.batch(batch_size)
xnli_hi_eval_dataset = xnli_hi_eval_dataset.batch(batch_size)
xnli_ru_eval_dataset = xnli_ru_eval_dataset.batch(batch_size)
xnli_sw_eval_dataset = xnli_sw_eval_dataset.batch(batch_size)
xnli_th_eval_dataset = xnli_th_eval_dataset.batch(batch_size)
xnli_tr_eval_dataset = xnli_tr_eval_dataset.batch(batch_size)
xnli_ur_eval_dataset = xnli_ur_eval_dataset.batch(batch_size)
xnli_vi_eval_dataset = xnli_vi_eval_dataset.batch(batch_size)
xnli_zh_eval_dataset = xnli_zh_eval_dataset.batch(batch_size)
model.evaluate(xnli_ar_eval_dataset)
model.evaluate(xnli_bg_eval_dataset)
model.evaluate(xnli_de_eval_dataset)
model.evaluate(xnli_el_eval_dataset)
model.evaluate(xnli_es_eval_dataset)
model.evaluate(xnli_fr_eval_dataset)
model.evaluate(xnli_hi_eval_dataset)
model.evaluate(xnli_ru_eval_dataset)
model.evaluate(xnli_sw_eval_dataset)
model.evaluate(xnli_th_eval_dataset)
model.evaluate(xnli_tr_eval_dataset)
model.evaluate(xnli_ur_eval_dataset)
model.evaluate(xnli_vi_eval_dataset)
model.evaluate(xnli_zh_eval_dataset)
# +
# resolver = tf.distribute.cluster_resolver.TPUClusterResolver.connect(tpu='tpu-quickstart', project = 'moana-intern-fall-2020')
# tf.config.experimental_connect_to_cluster(resolver)
# tf.tpu.experimental.initialize_tpu_system(resolver)
# strategy = tf.distribute.TPUStrategy(resolver)
# with strategy.scope():
# max_seq_length = 128
# initializer = tf.keras.initializers.TruncatedNormal(
# stddev=bert_config.initializer_range)
# bert_encoder = bert.bert_models.get_transformer_encoder(
# bert_config, max_seq_length)
# input_word_ids = tf.keras.layers.Input(
# shape=(max_seq_length,), dtype=tf.int32, name='input_word_ids')
# input_mask = tf.keras.layers.Input(
# shape=(max_seq_length,), dtype=tf.int32, name='input_mask')
# input_type_ids = tf.keras.layers.Input(
# shape=(max_seq_length,), dtype=tf.int32, name='input_type_ids')
# bert_model = hub.KerasLayer("https://tfhub.dev/tensorflow/bert_multi_cased_L-12_H-768_A-12/2",
# trainable=True)
# #bert_model = hub.KerasLayer(hub_url_bert, trainable=True)
# pooled_output, seq_output = bert_model([input_word_ids, input_mask, input_type_ids])
# output1 = tf.keras.layers.Dropout(rate=bert_config.hidden_dropout_prob)(
# pooled_output)
# output1 = tf.keras.layers.Dense(
# 2, kernel_initializer=initializer, name='output1')(
# output1)
# output2 = tf.keras.layers.Dropout(rate=bert_config.hidden_dropout_prob)(
# pooled_output)
# output2 = tf.keras.layers.Dense(
# 3, kernel_initializer=initializer, name='output2')(
# output2)
# model = tf.keras.Model(
# inputs={
# 'input_word_ids': input_word_ids,
# 'input_mask': input_mask,
# 'input_type_ids': input_type_ids
# },
# outputs=[output1, output2])
# # Set up epochs and steps
# epochs = 3
# batch_size = 64
# eval_batch_size = 64
# # get train_data_size from metadata
# train_data_size = c
# steps_per_epoch = int(train_data_size / batch_size)
# num_train_steps = steps_per_epoch * epochs
# warmup_steps = int(epochs * train_data_size * 0.1 / batch_size)
# # creates an optimizer with learning rate schedule
# optimizer = nlp.optimization.create_optimizer(
# 2e-5, num_train_steps=num_train_steps, num_warmup_steps=warmup_steps)
# paws_training_dataset = create_classifier_dataset(
# "gs://nts2020/xtereme/pawsx/train.en.tfrecords",
# 128,
# batch_size,
# task_id = 0,
# is_training=True)
# xnli_training_dataset = create_classifier_dataset(
# "gs://nts2020/xtereme/xnli/train.en.tfrecords",
# 128,
# batch_size,
# task_id =1,
# is_training=True)
# paws_eval_dataset = create_classifier_dataset(
# "gs://nts2020/xtereme/pawsx/eval.en.tfrecords",
# 128,
# batch_size,
# task_id = 0,
# is_training=False)
# xnli_eval_dataset = create_classifier_dataset(
# "gs://nts2020/xtereme/xnli/eval.en.tfrecords",
# 128,
# batch_size,
# task_id = 1,
# is_training=False)
# training_dataset = tf.data.experimental.sample_from_datasets(
# [paws_training_dataset, xnli_training_dataset], weights=tf.constant([sampling_factor[0], sampling_factor[1]]))
# evaluation_dataset = tf.data.experimental.sample_from_datasets(
# [paws_eval_dataset, xnli_eval_dataset], weights=tf.constant([sampling_factor[0], sampling_factor[1]]))
# training_dataset = training_dataset.batch(batch_size)
# evaluation_dataset = evaluation_dataset.batch(batch_size)
# model.compile(optimizer = optimizer, loss = [_loss_with_filter, _loss_with_filter], metrics = [accuracy_mod])
# model.fit(training_dataset, batch_size = batch_size, epochs= 13, steps_per_epoch = 1000, validation_data=evaluation_dataset)
| dev/dev_mt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from espn_api.football import League, Team, Player
from copy import copy
from collections import Counter
from tabulate import tabulate as table
# +
year = 2020
login = pd.read_csv('login.csv')
manager, league_name, league_id, swid, espn_s2 = login.iloc[1]
# +
league = League(league_id=league_id,
year=year,
swid=swid,
espn_s2=espn_s2)
week = 4
league.load_roster_week(week)
team = league.teams[0]
# box_scores = league.box_scores(week)
# box_score = box_scores[0]
# -
# Get a dictionary of the starting roster slots and number of each for the League (Week 1 must have passed already)
starting_roster_slots = Counter([p.slot_position for p in league.box_scores(1)[0].home_lineup if p.slot_position not in ['BE', 'IR']])
league.name
# ## Analytic functions
# +
def get_lineup(league: League, team: Team, week: int, box_scores=None):
''' Return the lineup of the given team during the given week '''
# Get the lineup for the team during the specified week
if box_scores is None: box_scores = league.box_scores(week)
for box_score in box_scores:
if team == box_score.home_team:
return box_score.home_lineup
elif team == box_score.away_team:
return box_score.away_lineup
def get_top_players(lineup: list, slot: str, n: int):
''' Takes a list of players and returns a list of the top n players based on points scored. '''
# Gather players of the desired position
eligible_players = []
for player in lineup:
if slot in player.eligibleSlots:
eligible_players.append(player)
return sorted(eligible_players, key=lambda x: x.points, reverse=True)[:n]
def get_best_lineup(lineup: list):
''' Returns the best possible lineup for team during the loaded week. '''
# Save full roster
saved_roster = copy(lineup)
# Find Best Lineup
best_lineup = []
for slot in sorted(starting_roster_slots.keys(), key=len): # Get best RB before best RB/WR/TE
num_players = starting_roster_slots[slot]
best_players = get_top_players(saved_roster, slot, num_players)
best_lineup.extend(best_players)
# Remove selected players from consideration for other slots
for player in best_players:
saved_roster.remove(player)
return np.sum([player.points for player in best_lineup])
def get_best_trio(lineup: list):
''' Returns the the sum of the top QB/RB/Reciever trio for a team during the loaded week. '''
qb = get_top_players(lineup, 'QB', 1)[0].points
rb = get_top_players(lineup, 'RB', 1)[0].points
wr = get_top_players(lineup, 'WR', 1)[0].points
te = get_top_players(lineup, 'TE', 1)[0].points
best_trio = round(qb + rb + max(wr, te), 2)
return best_trio
def get_lineup_efficiency(lineup: list):
max_score = get_best_lineup(lineup)
real_score = np.sum([player.points for player in lineup if player.slot_position not in ('BE', 'IR')])
return real_score / max_score
def get_weekly_finish(league: League, team: Team, week: int):
''' Returns the rank of a team compared to the rest of the league by points for (for the loaded week) '''
league_scores = [tm.scores[week-1] for tm in league.teams]
league_scores = sorted(league_scores, reverse=True)
return league_scores.index(team.scores[week-1]) + 1
def get_num_out(lineup: list):
''' Returns the (esimated) number of players who did not play for a team for the loaded week (excluding IR slot players). '''
num_out = 0
# TODO: write new code based on if player was injured
return num_out
def avg_slot_score(lineup: list, slot: str):
'''
Returns the average score for starting players of a specified slot.
`lineup` is either BoxScore().away_lineup or BoxScore().home_lineup (a list of BoxPlayers)
'''
return np.mean([player.points for player in lineup if player.slot_position == slot])
def sum_bench_points(lineup: list):
'''
Returns the total score for bench players
`lineup` is either BoxScore().away_lineup or BoxScore().home_lineup (a list of BoxPlayers)
'''
return np.sum([player.points for player in lineup if player.slot_position == 'BE'])
def print_weekly_stats(league: League, team: Team, week: int):
''' Print the weekly stats for the team during a given week. '''
lineup = get_lineup(league, team, week)
stats_table = [['Week Score: ', team.scores[week-1]],
['Best Possible Lineup: ', get_best_lineup(lineup)],
['Opponent Score: ', team.schedule[week-1].scores[week-1]],
['Weekly Finish: ', get_weekly_finish(league, team, week)],
['Best Trio: ', get_best_trio(lineup)],
['Number of Injuries: ', get_num_out(lineup)],
['Starting QB pts: ', avg_slot_score(lineup, 'QB')],
['Avg. Starting RB pts: ', avg_slot_score(lineup, 'RB')],
['Avg. Starting WR pts: ', avg_slot_score(lineup, 'WR')],
['Starting TE pts: ', avg_slot_score(lineup, 'TE')],
['Starting Flex pts: ', avg_slot_score(lineup, 'RB/WR/TE')],
['Starting DST pts: ', avg_slot_score(lineup, r'D/ST')],
['Starting K pts: ', avg_slot_score(lineup, 'K')],
['Total Bench pts: ', sum_bench_points(lineup)]]
print('\n', table(stats_table, headers = ['Week ' + str(week), ''], numalign = 'left'))
print_weekly_stats(league, team, 4)
# +
# box_scores = league.box_scores(week)
# # %time [get_lineup(league, team, week) for i in range(10)]
# # %time [get_lineup(league, team, week, box_scores) for i in range(10)]
# -
# ## Advanced stat functions
league.power_rankings(week)
# +
def get_weekly_luck_index(league: League, team: Team, week: int):
'''
This function returns an index quantifying how 'lucky' a team was in a given week
Luck index:
50% probability of playing a team with a lower record
25% your play compared to previous weeks
25% opp's play compared to previous weeks
'''
opp = team.schedule[week-1]
num_teams = len(league.teams)
# Luck Index based on where the team and its opponent finished compared to the rest of the league
rank = get_weekly_finish(league, team, week)
opp_rank = get_weekly_finish(league, opp, week)
if rank < opp_rank: # If the team won...
luck_index = 5 * (rank - 1) / (num_teams - 2) # Odds of this team playing a team with a higher score than it
elif rank > opp_rank: # If the team lost or tied...
luck_index = -5 * (num_teams - rank) / (num_teams - 2) # Odds of this team playing a team with a lower score than it
# If the team tied...
elif rank < (num_teams / 2):
luck_index = -2.5 * (num_teams - rank - 1) / (num_teams - 2) # They are only half as unlucky, because tying is not as bad as losing
else:
luck_index = 2.5 * (rank - 1) / (num_teams - 2) # They are only half as lucky, because tying is not as good as winning
# Update luck index based on how team played compared to normal
team_score = team.scores[week - 1]
team_avg = np.mean(team.scores[:week])
team_std = np.std(team.scores[:week])
if team_std != 0:
# Get z-score of the team's performance
z = (team_score - team_avg) / team_std
# Noramlize the z-score so that a performance 3 std dev's away from the mean has an effect of 2 points on the luck index
z_norm = z / (3*team_std) * 2.5
luck_index += z_norm
# Update luck index based on how opponent played compared to normal
opp_score = opp.scores[week - 1]
opp_avg = np.mean(opp.scores[:week])
opp_std = np.std(opp.scores[:week])
if team_std != 0:
# Get z-score of the team's performance
z = (opp_score - opp_avg) / opp_std
# Noramlize the z-score so that a performance 3 std dev's away from the mean has an effect of 2 points on the luck index
z_norm = z / (3*opp_std) * 2.5
luck_index -= z_norm
return luck_index / 10
def get_season_luck_indices(league: League, week: int):
''' This function returns an index quantifying how 'lucky' a team was all season long (up to a certain week) '''
luck_indices = {team:0 for team in league.teams}
for wk in range(1, week + 1):
# Update luck_index for each team
for team in league.teams:
luck_indices[team] += get_weekly_luck_index(league, team, week)
return luck_indices
# +
def sort_lineups_by_func(league: League, week: int, func, box_scores=None, **kwargs):
'''
Sorts league teams according to function.
Values are sorted ascending.
DOES NOT ACCOUNT FOR TIES
'''
if box_scores is None: box_scores = league.box_scores(week)
return sorted(league.teams, key=lambda x:func(get_lineup(league, x, week, box_scores), **kwargs))
def print_weekly_stats(league: League, week: int):
''' Prints weekly stat report for a league during a given week '''
# Load box scores for specified week
box_scores = league.box_scores(week)
statsTable = [['Most Points Scored: ', sorted(league.teams, key=lambda x:x.scores[week-1], reverse=True)[0].owner],
['Least Points Scored: ', sorted(league.teams, key=lambda x:x.scores[week-1])[0].owner],
['Best Possible Lineup: ', sort_lineups_by_func(league, week, get_best_lineup, box_scores)[-1].owner],
['Best Trio: ', sort_lineups_by_func(league, week, get_best_trio, box_scores)[-1].owner],
['Worst Trio: ', sort_lineups_by_func(league, week, get_best_trio, box_scores)[0].owner],
['Best Lineup Setter', sort_lineups_by_func(league, week, get_lineup_efficiency, box_scores)[-1].owner],
['Worst Lineup Setter', sort_lineups_by_func(league, week, get_lineup_efficiency, box_scores)[0].owner],
['---------------------','----------------'],
['Best QBs: ', sort_lineups_by_func(league, week, avg_slot_score, box_scores, slot='QB')[-1].owner],
['Best RBs: ', sort_lineups_by_func(league, week, avg_slot_score, box_scores, slot='RB')[-1].owner],
['Best WRs: ', sort_lineups_by_func(league, week, avg_slot_score, box_scores, slot='WR')[-1].owner],
['Best TEs: ', sort_lineups_by_func(league, week, avg_slot_score, box_scores, slot='TE')[-1].owner],
['Best Flex: ', sort_lineups_by_func(league, week, avg_slot_score, box_scores, slot=r'RB/WR/TE')[-1].owner],
['Best DST: ', sort_lineups_by_func(league, week, avg_slot_score, box_scores, slot=r'D/ST')[-1].owner],
['Best K: ', sort_lineups_by_func(league, week, avg_slot_score, box_scores, slot='K')[-1].owner],
['Best Bench:', sort_lineups_by_func(league, week, sum_bench_points, box_scores)[-1].owner],
['---------------------','----------------'],
['Worst QBs: ', sort_lineups_by_func(league, week, avg_slot_score, box_scores, slot='QB')[0].owner],
['Worst RBs: ', sort_lineups_by_func(league, week, avg_slot_score, box_scores, slot='RB')[0].owner],
['Worst WRs: ', sort_lineups_by_func(league, week, avg_slot_score, box_scores, slot='WR')[0].owner],
['Worst TEs: ', sort_lineups_by_func(league, week, avg_slot_score, box_scores, slot='TE')[0].owner],
['Worst Flex: ', sort_lineups_by_func(league, week, avg_slot_score, box_scores, slot=r'RB/WR/TE')[0].owner],
['Worst DST: ', sort_lineups_by_func(league, week, avg_slot_score, box_scores, slot=r'D/ST')[0].owner],
['Worst K: ', sort_lineups_by_func(league, week, avg_slot_score, box_scores, slot='K')[0].owner],
['Worst Bench:', sort_lineups_by_func(league, week, sum_bench_points, box_scores)[0].owner],
]
print('\n', table(statsTable, headers = ['Week ' + str(week), '']))
print_weekly_stats(league, 12)
# -
league.standings()
league.scoringPeriodId
# +
def print_current_standings(league: League):
''' Inputs: None
Outputs: table (prints current standings)
This function prints the current standings for a league.
This function does NOT account for tiebreakers.
'''
standings = league.standings()
results_table = []
for team in standings:
results_table += [[ team.team_name, team.wins, team.losses, team.ties, team.points_for, team.owner ]]
print('\nWeek {}\n'.format(league.currentMatchupPeriod),
table(results_table,
headers = ['Team', 'Wins', 'Losses', 'Ties', 'Points Scored', 'Owner'],
floatfmt = '.2f'))
print_current_standings(league)
# -
# +
# value_max = df.groupby([by, col])\
# .count()['player_id']\
# .unstack()\
# .fillna(0)\
# .idxmax(axis=1)
# value_count = df.groupby([by, col])\
# .count()['player_id']\
# .unstack()\
# .fillna(0)\
# .max(axis=1)
# value_counts = pd.concat([value_max, value_count], axis=1).rename(columns={0:'value', 1:'count'})
def get_draft_details(league: League, primary_slots: list = None):
draft = pd.DataFrame()
if primary_slots is None:
# Get a dictionary of the starting roster slots and number of each for the League (Week 1 must have passed already)
primary_slots = [slot for slot in starting_roster_slots.keys() if ('/' not in slot) or (slot == 'D/ST')]
for i, player in enumerate(league.draft):
draft.loc[i, 'year'] = league.year
draft.loc[i, 'team_owner'] = player.team.owner
draft.loc[i, 'team_id'] = player.team.team_id
draft.loc[i, 'player_name'] = player.playerName
draft.loc[i, 'player_id'] = player.playerId
draft.loc[i, 'round_num'] = player.round_num
draft.loc[i, 'round_pick'] = player.round_pick
try:
# Get more player details (can take 1.5 min)
player = league.player_info(playerId=draft.loc[i, 'player_id'])
draft.loc[i, 'pro_team'] = player.proTeam
draft.loc[i, 'proj_points'] = player.projected_total_points
draft.loc[i, 'total_points'] = player.total_points
draft.loc[i, 'position'] = [slot for slot in player.eligibleSlots if slot in primary_slots][0]
except AttributeError:
print('Pick {} missing.'.format(i+1))
draft.loc[i, 'player_name'] = ''
draft.loc[i, 'player_id'] = ''
draft.loc[i, 'round_num'] = 99
draft.loc[i, 'round_pick'] = 99
except:
print(i, player, league.draft[i-2:i+2])
draft.loc[i, 'position'] = player.eligibleSlots[0]
draft['first_letter'] = draft.player_name.str[0]
draft['points_surprise'] = draft.total_points - draft.proj_points
draft['positive_surprise'] = draft.points_surprise > 0
draft['pick_num'] = (draft.round_num - 1) * len(draft.team_id.unique()) + draft.round_pick
draft_pick_values = pd.read_csv('./pick_value.csv')
draft = pd.merge(draft, draft_pick_values, left_on='pick_num', right_on='pick', how='left').drop(columns=['pick'])
return draft
def get_multiple_drafts(league: League, start_year: int = 2020, end_year: int = 2021, swid=None, espn_s2=None):
draft = pd.DataFrame()
for year in range(start_year, end_year+1):
print('Fetching {} draft...'.format(year), end='')
try:
draft_league = League(league_id=league.league_id,
year=year,
swid=swid,
espn_s2=espn_s2)
except: continue
# Get a dictionary of the starting roster slots and number of each for the League (Week 1 must have passed already)
# try:
# starting_roster_slots = Counter([p.slot_position for p in draft_league.box_scores(1)[0].home_lineup if p.slot_position not in ['BE', 'IR']])
# except:
# starting_roster_slots = Counter([p.slot_position for p in league.box_scores(1)[0].home_lineup if p.slot_position not in ['BE', 'IR']])
try:
primary_slots = [slot for slot in starting_roster_slots.keys() if ('/' not in slot) or (slot == 'D/ST')]
draft = pd.concat([draft, get_draft_details(draft_league, primary_slots)])
print('Done.')
except: continue
return draft
def get_team_max(df, col, by='team_owner', keep=None):
'''
`by` = 'team_id', 'team_owner'
'''
def get_maxs(s):
return ' | '.join(s[s == s.max()].index.values)
value_counts = df.groupby([by, col])\
.count()['player_id']\
.unstack()\
.fillna(0)
value_counts['max_value'] = value_counts.apply(get_maxs, axis=1)
value_counts['max_count'] = value_counts.max(axis=1)
value_counts = value_counts.iloc[:, -2:]
if keep is not None:
return value_counts[value_counts.index.isin(keep)]
else: return value_counts
# -
# # %time draft = get_draft_details(league) 2015
# %time draft = get_multiple_drafts(league, 2021, 2021, swid, espn_s2)
draft.head()
league = League(league_id=league_id,
year=2021,
swid=swid,
espn_s2=espn_s2)
draft['first_letter'].value_counts().plot(kind='bar', rot=0);
get_team_max(draft, 'first_letter', keep=[team.owner for team in league.teams])
draft[(draft.first_letter == 'J') & (draft.team_owner == '<NAME>')]
draft['position'].value_counts().plot(kind='bar', rot=0);
draft['position'].value_counts()
get_team_max(draft, 'position', keep=[team.owner for team in league.teams])
draft['pro_team'].value_counts().plot(kind='bar', rot=90);
draft['pro_team'].value_counts()
get_team_max(draft[draft.round_num <= 20], 'pro_team', keep=[team.owner for team in league.teams])
get_team_max(draft.dropna(how='any'), 'player_name', keep=[team.owner for team in league.teams])
draft['points_surprise'].plot(kind='hist', rot=0, bins=25);
best_pick = draft[draft.year > 2017].sort_values(by='points_surprise', ascending=False).groupby('team_owner').first()
best_pick = best_pick[best_pick.index.isin([team.owner for team in league.teams])]
best_pick
best_pick = draft[draft.year > 2017].sort_values(by='points_surprise', ascending=False).groupby('position').first()
best_pick
worst_pick = draft[draft.year > 2017].sort_values(by='points_surprise', ascending=False).groupby('position').last()
worst_pick
best_player = draft[draft.year == 2020].sort_values(by='total_points', ascending=False).groupby('position').first()
best_player
worst_player = draft[draft.year == 2020].sort_values(by='total_points', ascending=False).groupby('position').last()
worst_player
# +
perc_boom = draft[draft.year > 2017].sort_values(by='positive_surprise', ascending=False).groupby('team_owner').mean()
perc_boom = perc_boom[['positive_surprise', 'proj_points', 'total_points', 'points_surprise']]
perc_boom.rename(columns={'positive_surprise':'Boom %',
'proj_points':'Avg. Projected Points',
'total_points':'Avg. Total Points',
'points_surprise': 'Avg. Points Surprise'},
inplace=True)
perc_boom = perc_boom.sort_values(by='Boom %', ascending=False)
perc_boom = perc_boom[perc_boom.index.isin([team.owner for team in league.teams])]
perc_boom
# +
def plot_surprise(draft, year: int, owner: str, sort_by='round_num', asc=False):
'''
sort_by = {'round_num', 'points_surprise'}
'''
if sort_by == 'points_surprise': asc = True
sub = draft[(draft.year == year) & (draft.team_owner == owner)].sort_values(by=sort_by, ascending=asc)
fig, ax = plt.subplots(figsize=(15, 10))
sub.plot(x='player_name',
y='points_surprise',
kind='barh',
color=(sub.points_surprise > 0).map({True: 'g', False: 'r'}),
legend=None,
ax=ax)
plt.axvline(0, color='k', lw=1)
ax.set_ylabel('')
if sort_by == 'points_surprise': ax.set_yticks([])
else: ax.set_yticklabels('Rd ' + sub.round_num.astype(str).str[:-2])
for i, (name, height) in enumerate(zip(sub.player_name, sub.points_surprise)):
ha = 'right' if height < 0 else 'left'
ax.text(height, i, ' {} '.format(name),
ha=ha, va='center', rotation=0, fontsize=10)
xmin, xmax = ax.get_xlim()
xmin -= len(sub.player_name.iloc[0])*3
xmax += len(sub.player_name.iloc[-1])*2
ax.set_xlim(xmin, xmax)
ax.set_title(owner, fontsize='x-large')
fig.show();
def plot_top_surprise(draft, year: int, top: int = 5):
sub = draft[draft.year == year].sort_values(by='points_surprise', ascending=True).dropna(how='any')
sub = pd.concat([sub.head(top), sub.tail(5)])
fig, ax = plt.subplots(figsize=(15, 10))
sub.plot(x='player_name',
y='points_surprise',
kind='barh',
color=(sub.points_surprise > 0).map({True: 'g', False: 'r'}),
legend=None,
ax=ax)
plt.axvline(0, color='k', lw=1)
ax.set_ylabel('')
ax.set_yticks([])
ax.set_yticklabels([])
# ax.set_title('Points Surpise', size='xx-large')
# Add player name
for i, (name, height) in enumerate(zip(sub.player_name, sub.points_surprise)):
ha = 'right' if height < 0 else 'left'
text = ' {} '.format(name)
ax.text(height, i, text, ha=ha, va='center', rotation=0, fontsize=15)
# Add team owner
for i, (owner, height) in enumerate(zip(sub.team_owner, sub.points_surprise)):
ha = 'right' if height > 0 else 'left'
text = ' {} '.format(owner)
ax.text(height, i, text, ha=ha, va='center', rotation=0, fontsize=13, weight='bold', color='white')
# Add round number
for i, (rd, height) in enumerate(zip(sub.round_num, sub.points_surprise)):
ha = 'right' if height < 0 else 'left'
text = ' Rd {:.0f} '.format(rd, owner)
height = 1 if height > 0 else -1
ax.text(height, i, text, ha=ha, va='center', rotation=0, fontsize=10, weight='bold', color='white')
xmin, xmax = ax.get_xlim()
xmin -= len(sub.player_name.iloc[0])*7
xmax += len(sub.player_name.iloc[-1])*7
ax.set_xlim(xmin, xmax)
ax.text(xmin / 2,
int(top*1.5),
'Top / bottom 5 players based on the\ndifference between projected and\nactual season point totals',
ha='center',
fontsize='xx-large')
ax.set_facecolor('lightgray')
fig.show();
# plot_surprise(draft, 2020, '<NAME>', 'points_surprise')
# plot_surprise(draft, 2020, '<NAME>', 'round_num')
plot_top_surprise(draft, 2020)
# -
for name in draft[draft.year == 2020].team_owner.unique():
plot_surprise(draft, 2020, name)
draft[draft.year > 2017].positive_surprise.mean()
# league.load_roster_week(18)
| .ipynb_checkpoints/new espn-checkpoint.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 9.4
# language: sagemath
# metadata:
# cocalc:
# description: Open-source mathematical software system
# priority: 10
# url: https://www.sagemath.org/
# name: sage-9.4
# resource_dir: /ext/jupyter/kernels/sage-9.4
# ---
# Nastavimo p za linearni program in definiramo spremenljivko k.
p = MixedIntegerLinearProgram(maximization=True)
k = p.new_variable(real=True)
# Z r, x, y označimo polmer ter koordinati središča.
r, x, y = (k[i] for i in "rxy")
# Maksimiziramo r
p.set_objective(r)
# Dodamo pogoj, da je polmer nenegativen.
p.add_constraint(r >= 0)
# S for zanko se sprehodimo po polravninah lika ter za vsako dodamo pogoj.
for ai, bi, ci in zip(a, b, c):
p.add_constraint(r * sqrt(ai^2 + bi^2) <= ci -ai*x-bi*y)
p.solve()
res = p.get_values(k)
res
| Krog2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7
# language: python
# name: python3
# ---
# # Decision Tree Induction with scikit-learn
# + [markdown] tags=["remove-cell"]
# **CS5483 Data Warehousing and Data Mining**
# ___
# + deletable=false editable=false init_cell=true nbgrader={"cell_type": "code", "checksum": "851cc20d47e07396bc98793d282e9860", "grade": false, "grade_id": "init", "locked": true, "schema_version": 3, "solution": false, "task": false} slideshow={"slide_type": "-"}
# %reset -f
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn import datasets, tree
# produce vector inline graphics
from IPython.display import set_matplotlib_formats
set_matplotlib_formats('svg')
# -
# ## Decision Tree Induction
# We first import the [*iris dataset*](https://en.wikipedia.org/wiki/Iris_flower_data_set) from [`sklearn.datasets` package](https://scikit-learn.org/stable/datasets/index.html)
# +
from sklearn import datasets
import pandas as pd
iris = datasets.load_iris()
# -
# Recall that the classification task is to train a model that can classify the spieces (*target*) automatically based on the lengths and widths of the petals and sepals (*input features*).
# To build a decision tree, we simply create a tree using `DecisionTreeClassifier` from `sklearn.tree` and apply its method `fit` on the training set.
clf_gini = tree.DecisionTreeClassifier(random_state=0).fit(iris.data, iris.target)
# To display the decision tree, we can use the function `plot_tree` from `sklearn.tree`:
# +
# to make the tree look better
options = {'feature_names': iris.feature_names,
'class_names': iris.target_names,
'filled': True,
'node_ids': True,
'rounded': True,
'fontsize': 6}
plt.figure(figsize=(9,6))
tree.plot_tree(clf_gini, **options)
plt.show()
# -
# For each node:
# - `___ <= ___` is the splitting criterion for internal nodes, satisfied only by samples going left.
# - `gini = ...` shows the impurity index. By default, the algorithm uses Gini impurity index to find the best binary split. Observe that the index decreases down the tree towards the leafs.
# - `value = [_, _, _]` shows the number of examples for each of the three classes, and `class = ...` indicates a majority class, which may be used as the decision for a leaf node. The majority classes are also color coded. Observe that the color gets lighter towards the root, as the class distribution is more impure.
#
# In particular, check that iris setosa is distinguished immediately after checking the petal width/length.
# All the information of the decision is stored in the `tree_` attribute of the classifer. For more details:
# + tags=["remove-output"]
help(clf_gini.tree_)
# -
# **Exercise** Assign to `clf_entropy` the decision tree classifier created using *entropy* as the impurity measure. You can do so with the keyword argument `criterion='entropy'` in `DecisionTreeClassifier`. Furthermore, Use `random_state=0` and fit the classifier on the entire iris dataset. Check whether the resulting decision tree same as the one created using the Gini impurity index.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "2b5af6f05d8ce6f84d7a2178a5ed5a53", "grade": false, "grade_id": "tree-entropy", "locked": false, "schema_version": 3, "solution": true, "task": false} tags=["remove-output"]
# YOUR CODE HERE
raise NotImplementedError()
plt.figure(figsize=(9, 6))
tree.plot_tree(clf_entropy, **options)
plt.show()
# + [markdown] deletable=false nbgrader={"cell_type": "markdown", "checksum": "b33243c1128464d72060ac89b3646dd1", "grade": true, "grade_id": "same-tree-as-gini", "locked": false, "points": 1, "schema_version": 3, "solution": true, "task": false}
# YOUR ANSWER HERE
# -
# It is important to note that, although one can specify whether to use Gini impurity or entropy, `sklearn` implements neither C4.5 nor CART algorithms. In particular, it supports only binary splits on numeric input attributes, unlike C4.5 which supports multi-way splits using information gain ratio.
# (See some [workarounds][categorical].)
#
# [categorical]: https://stackoverflow.com/questions/38108832/passing-categorical-data-to-sklearn-decision-tree
# ## Compute Splitting Criterion
# To understand how the decision tree is generated, we will implements the computation of the splitting criterion.
# ### Basic data analysis using `pandas`
# To have an idea of qualities of the features, create a [`pandas`](https://pandas.pydata.org/docs/user_guide/index.html) [`DataFrame`](https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.html?highlight=dataframe#pandas.DataFrame)
# to operate on the dataset.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="9gQINfrjsb4M" outputId="77b77a38-2712-4c93-c503-219e74f354fd" slideshow={"slide_type": "-"}
# write the input features first
iris_df = pd.DataFrame(data=iris.data, columns=iris.feature_names)
# append the target values to the last column
iris_df['target'] = iris.target
iris_df.target = iris_df.target.astype('category')
iris_df.target.cat.categories = iris.target_names
iris_df
# -
# To display some statistics of the input features for different classes:
iris_df.groupby('target').boxplot(rot=90, layout=(1,3))
iris_df.groupby('target').agg(['mean','std']).round(2)
# **Exercise** Identify good feature(s) based on the above statistics. Does you choice agree with the decision tree generated by `DecisionTreeClassifier`?
# + [markdown] deletable=false nbgrader={"cell_type": "markdown", "checksum": "23015178e587e6f2ad5fd85735da2221", "grade": true, "grade_id": "good-features", "locked": false, "points": 1, "schema_version": 3, "solution": true, "task": false}
# YOUR ANSWER HERE
# -
# ### Compute impurity
# Given a distribution $\boldsymbol{p}=(p_1,p_2,\dots)$ where $p_k\in [0,1]$ and $\sum_k p_k =1$, the Gini impurity index is defined as:
#
# \begin{align}
# \operatorname{Gini}(\boldsymbol{p}) = \operatorname{Gini}(p_1,p_2,\dots) &:= \sum_k p_k(1-p_k)\\
# &= 1- \sum_k p_k^2.
# \end{align}
# We can represent a distribution simply as a `numpy` array. To return the empirical class distributions of the iris dataset:
# +
def dist(values):
'''Returns the empirical distribution of the given 1D array of values as a
1D array of probabilites.'''
counts = np.unique(values, return_counts=True)[-1]
return counts / counts.sum()
print(f"Distribution of target: {dist(iris.target).round(4)}")
# -
# The Gini impurity index can be computed as follows:
# +
def gini(p):
'''Returns the Gini impurity of distribution p.'''
return 1 - (p**2).sum()
print(f"Gini impurity of target: {gini(dist(iris.target)):.4g}")
# -
# **Exercise** Complete the following function to compute the entropy of a distribution:
#
# \begin{align}
# h(\boldsymbol{p}) = h(p_1,p_2,\dots) &= \sum_k - p_k \log_2 p_k\\
# &= \sum_{k:p_k>0} - p_k \log_2 p_k.
# \end{align}
#
# You may use the function `log2` from `numpy` to calculate the logarithm base 2. Note that logarithm of $0$ is undefined, we use the last expression of the entropy to avoid taking the limit $\lim_{p\to 0} p\log p=0$.
#
# You solution should look like:
# ```Python
# def entropy(p):
# ...
# return (p * ___ * ___).sum()
# ```
# + deletable=false nbgrader={"cell_type": "code", "checksum": "f18b27cbcc409dcd4a353adfe721bbc2", "grade": false, "grade_id": "entropy", "locked": false, "schema_version": 3, "solution": true, "task": false} tags=["remove-output"]
def entropy(p):
'''Returns the entropy of distribution p.'''
p = np.array(p)
p = p[(p > 0) & (p < 1)] # 0 log 0 = 1 log 1 = 0
# YOUR CODE HERE
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "6413a8489e09d23de03ae05263557271", "grade": true, "grade_id": "test-entropy", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} tags=["remove-output"]
# tests
assert np.isclose(entropy([1/2, 1/2]), 1)
assert np.isclose(entropy([1, 0]), 0)
# -
# ### Compute drop in impurity and best split
# Now, to compute the drop in impurity for given splitting criterion:
#
# \begin{align}
# \Delta \operatorname{Gini}_{X\leq s}(Y) = \operatorname{Gini}(P_Y) - \left[\Pr\{X\leq s\} \operatorname{Gini}(P_{Y|X\leq s}) + \Pr\{X> s\}\operatorname{Gini}(P_{Y|X> s})\right]
# \end{align}
# +
def drop_in_gini(X, Y, split_pt):
'''Returns the drop in Gini impurity of Y for the split X <= split_pt.
Parameters
----------
X: 1D array the values of a feature for different instances
Y: 1D array of the corresponding target values
split_pt: the value of the split point
'''
S = X <= split_pt
q = S.mean()
return gini(dist(Y)) - q * gini(dist(Y[S])) - (1 - q) * gini(dist(Y[~S]))
X, Y = iris_df['petal width (cm)'], iris_df.target
print(f"Drop in Gini: {drop_in_gini(X, Y, 0.8):.4g}")
# -
# To compute the best split point for a feature, we check every consecutive mid-points of the possible feature values:
# +
def find_best_split_pt(X, Y, gain_function):
'''Return the best split point s and the maximum gain evaluated using
gain_function for the split X <= s and target Y.
Parameters
----------
X: 1D array the values of a feature for different instances
Y: 1D array of the corresponding target values
gain_function: a function such as drop_in_gini for evaluating a split
Returns
-------
A tuple (s, g) where s is the best split point and g is the maximum gain.
See also
--------
drop_in_gini
'''
values = np.sort(np.unique(X))
split_pts = (values[1:] + values[:-1]) / 2
gain = np.array([gain_function(X, Y, s) for s in split_pts])
max_index = np.argmax(gain)
return split_pts[max_index], gain[max_index]
print('''Best split point: {0:.4g}
Maximum gain: {1:.4g}'''.format(*find_best_split_pt(X, Y, drop_in_gini)))
# -
# The following ranks the features according to the gains of their best binary splits:
rank_by_gini = pd.DataFrame({
'feature': feature,
**(lambda s, g: {
'split point': s,
'gain': g
})(*find_best_split_pt(iris_df[feature], iris_df.target, drop_in_gini))
} for feature in iris.feature_names).sort_values(by='gain', ascending=False)
rank_by_gini
# **Exercise** Complete the following function to calculate the *information gain* for a binary split $X\leq s$ and target $Y$:
#
# \begin{align}
# \operatorname{Gain}_{X\leq s}(Y) := h(P_Y) - \left[\Pr(X\leq s) h(P_{Y|X\leq s}) + \Pr(X> s) h(P_{Y|X> s})\right].
# \end{align}
#
# You may use `dist` and `entropy` defined previously.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "b9b4d69805318711d8c61770cafe6922", "grade": false, "grade_id": "info-gain", "locked": false, "schema_version": 3, "solution": true, "task": false} tags=["remove-output"]
def info_gain(X, Y, split_pt):
'''Returns the information Gain of Y for the split X <= split_pt.
Parameters
----------
X: 1D array the values of a feature for different instances
Y: 1D array of the corresponding target values
split_pt: the value of the split point
'''
S = X <= split_pt
q = S.mean()
# YOUR CODE HERE
raise NotImplementedError()
print(f"Information gain: {info_gain(X, Y, 0.8):.4g}")
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "10a933cf9c02c5de17b080f633d449f3", "grade": true, "grade_id": "test-info-gain", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} tags=["remove-output"]
# tests
rank_by_entropy = pd.DataFrame({
'feature':
feature,
**(lambda s, g: {
'split point': s,
'gain': g
})(*find_best_split_pt(iris_df[feature], iris_df.target, info_gain))
} for feature in iris.feature_names).sort_values(by='gain', ascending=False)
rank_by_entropy
# -
# **Exercise** Complete the following function to calculate the *information gain ratio* for a binary split $X\leq s$ and target $Y$:
#
# \begin{align}
# \operatorname{GainRatio}_{X\leq s}(Y) &:= \frac{\operatorname{Gain}_{X\leq s}(Y)}{\operatorname{SplitInfo}(X\leq s)} \qquad\text{where}\\
# \operatorname{SplitInfo}(X\leq s) &:= h(\Pr(X\leq s),\Pr(X> s)).
# \end{align}
#
# You may use `entropy` and `info_gain` defined previously.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "4d741235b484b516c1bf725fb66ef845", "grade": false, "grade_id": "info-gain-ratio", "locked": false, "schema_version": 3, "solution": true, "task": false} tags=["remove-output"]
def info_gain_ratio(X, Y, split_pt):
S = X <= split_pt
q = S.mean()
# YOUR CODE HERE
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "fc8476135900c3ab349e9d4643fd3640", "grade": true, "grade_id": "test-info-gain-ratio", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} tags=["remove-output"]
# tests
rank_by_info_gain_ratio = pd.DataFrame({
'feature':
feature,
**(lambda s, g: {
'split point': s,
'gain': g
})(*find_best_split_pt(iris_df[feature], iris_df.target, info_gain_ratio))
} for feature in iris.feature_names).sort_values(by='gain', ascending=False)
rank_by_info_gain_ratio
# -
# **Exercise** Does the information gain ratio give a different ranking of the features? Why?
# Information gain ratio gives the same ranking as information gain in this case. This is because the split is restricted to be binary and so the normalization by split information has less effect on the ranking.
| Tutorial3/Decision Tree Induction with scikit-learn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tmv
# language: python
# name: tmv
# ---
# +
s = ["US","EU","AR","BR","CA","JP","ML","SA","ZA","TH","US"]
ghgs = [14.75, 9.33, 0.74, 2.25, 1.63, 2.99, 0.06, 1.21, 1.13, 0.82]
print(sum(ghgs))
ns = ["AZ","CN","EG","KE","KG","MG","MV","SW","TZ"]
ghg_ns = [0.15, 27.51, 0.6, 0.13, 0.03, 0.06, 0.00, 0.11, 0.18]
print(sum(ghg_ns))
print(sum(ghgs+ghg_ns))
| code/solutions_emissions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <!-- HTML file automatically generated from DocOnce source (https://github.com/doconce/doconce/)
# doconce format html week43.do.txt --no_mako -->
# <!-- dom:TITLE: Week 43: Deep Learning: Recurrent Neural Networks and other Deep Learning Methods. Principal Component analysis -->
# # Week 43: Deep Learning: Recurrent Neural Networks and other Deep Learning Methods. Principal Component analysis
# **<NAME>**, Department of Physics, University of Oslo and Department of Physics and Astronomy and National Superconducting Cyclotron Laboratory, Michigan State University
#
# Date: **Nov 2, 2021**
#
# Copyright 1999-2021, <NAME>. Released under CC Attribution-NonCommercial 4.0 license
# ## Plans for week 43
#
# * Thursday: Summary of Convolutional Neural Networks from week 42 and Recurrent Neural Networks
#
# * [Video of Lecture](https://www.uio.no/studier/emner/matnat/fys/FYS-STK3155/h21/forelesningsvideoer/LectureOctober28.mp4?vrtx=view-as-webpage)
#
# * Friday: Recurrent Neural Networks and other Deep Learning methods such as Generalized Adversarial Neural Networks. Start discussing Principal component analysis
#
# * [Video of Lecture](https://www.uio.no/studier/emner/matnat/fys/FYS-STK3155/h21/forelesningsvideoer/LectureOctober29.mp4?vrtx=view-as-webpage)
#
# **Excellent lectures on CNNs and RNNs.**
#
# * [Video on Convolutional Neural Networks from MIT](https://www.youtube.com/watch?v=iaSUYvmCekI&ab_channel=AlexanderAmini)
#
# * [Video on Recurrent Neural Networks from MIT](https://www.youtube.com/watch?v=SEnXr6v2ifU&ab_channel=AlexanderAmini)
#
# * [Video on Deep Learning](https://www.youtube.com/playlist?list=PLZHQObOWTQDNU6R1_67000Dx_ZCJB-3pi)
#
# **More resources.**
#
# * [IN5400 at UiO Lecture](https://www.uio.no/studier/emner/matnat/ifi/IN5400/v20/material/week10/in5400_2020_week10_recurrent_neural_network.pdf)
#
# * [CS231 at Stanford Lecture](https://www.youtube.com/watch?v=6niqTuYFZLQ&list=PLzUTmXVwsnXod6WNdg57Yc3zFx_f-RYsq&index=10&ab_channel=StanfordUniversitySchoolofEngineering)
# ## Reading Recommendations
#
# * Goodfellow et al, chapter 10 on Recurrent NNs, chapters 11 and 12 on various practicalities around deep learning are also recommended.
#
# * <NAME>, chapter 14 on RNNs.
# ## Summary on Deep Learning Methods
#
# We have studied fully connected neural networks (also called artifical nueral networks) and convolutional neural networks (CNNs).
#
# The first type of deep learning networks work very well on homogeneous and structured input data while CCNs are normally tailored to recognizing images.
# ## CNNs in brief
#
# In summary:
#
# * A CNN architecture is in the simplest case a list of Layers that transform the image volume into an output volume (e.g. holding the class scores)
#
# * There are a few distinct types of Layers (e.g. CONV/FC/RELU/POOL are by far the most popular)
#
# * Each Layer accepts an input 3D volume and transforms it to an output 3D volume through a differentiable function
#
# * Each Layer may or may not have parameters (e.g. CONV/FC do, RELU/POOL don’t)
#
# * Each Layer may or may not have additional hyperparameters (e.g. CONV/FC/POOL do, RELU doesn’t)
#
# For more material on convolutional networks, we strongly recommend
# the course
# [IN5400 – Machine Learning for Image Analysis](https://www.uio.no/studier/emner/matnat/ifi/IN5400/index-eng.html)
# and the slides of [CS231](http://cs231n.github.io/convolutional-networks/) which is taught at Stanford University (consistently ranked as one of the top computer science programs in the world). [<NAME>'s book is a must read, in particular chapter 6 which deals with CNNs](http://neuralnetworksanddeeplearning.com/chap6.html).
#
# However, both standard feed forwards networks and CNNs perform well on data with unknown length.
#
# This is where recurrent nueral networks (RNNs) come to our rescue.
# ## Recurrent neural networks: Overarching view
#
# Till now our focus has been, including convolutional neural networks
# as well, on feedforward neural networks. The output or the activations
# flow only in one direction, from the input layer to the output layer.
#
# A recurrent neural network (RNN) looks very much like a feedforward
# neural network, except that it also has connections pointing
# backward.
#
# RNNs are used to analyze time series data such as stock prices, and
# tell you when to buy or sell. In autonomous driving systems, they can
# anticipate car trajectories and help avoid accidents. More generally,
# they can work on sequences of arbitrary lengths, rather than on
# fixed-sized inputs like all the nets we have discussed so far. For
# example, they can take sentences, documents, or audio samples as
# input, making them extremely useful for natural language processing
# systems such as automatic translation and speech-to-text.
# ## Set up of an RNN
#
# More to text to be added
# ## A simple example
# +
# %matplotlib inline
# Start importing packages
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow.keras import datasets, layers, models
from tensorflow.keras.layers import Input
from tensorflow.keras.models import Model, Sequential
from tensorflow.keras.layers import Dense, SimpleRNN, LSTM, GRU
from tensorflow.keras import optimizers
from tensorflow.keras import regularizers
from tensorflow.keras.utils import to_categorical
# convert into dataset matrix
def convertToMatrix(data, step):
X, Y =[], []
for i in range(len(data)-step):
d=i+step
X.append(data[i:d,])
Y.append(data[d,])
return np.array(X), np.array(Y)
step = 4
N = 1000
Tp = 800
t=np.arange(0,N)
x=np.sin(0.02*t)+2*np.random.rand(N)
df = pd.DataFrame(x)
df.head()
plt.plot(df)
plt.show()
values=df.values
train,test = values[0:Tp,:], values[Tp:N,:]
# add step elements into train and test
test = np.append(test,np.repeat(test[-1,],step))
train = np.append(train,np.repeat(train[-1,],step))
trainX,trainY =convertToMatrix(train,step)
testX,testY =convertToMatrix(test,step)
trainX = np.reshape(trainX, (trainX.shape[0], 1, trainX.shape[1]))
testX = np.reshape(testX, (testX.shape[0], 1, testX.shape[1]))
model = Sequential()
model.add(SimpleRNN(units=32, input_shape=(1,step), activation="relu"))
model.add(Dense(8, activation="relu"))
model.add(Dense(1))
model.compile(loss='mean_squared_error', optimizer='rmsprop')
model.summary()
model.fit(trainX,trainY, epochs=100, batch_size=16, verbose=2)
trainPredict = model.predict(trainX)
testPredict= model.predict(testX)
predicted=np.concatenate((trainPredict,testPredict),axis=0)
trainScore = model.evaluate(trainX, trainY, verbose=0)
print(trainScore)
index = df.index.values
plt.plot(index,df)
plt.plot(index,predicted)
plt.axvline(df.index[Tp], c="r")
plt.show()
# -
# ## An extrapolation example
#
# The following code provides an example of how recurrent neural
# networks can be used to extrapolate to unknown values of physics data
# sets. Specifically, the data sets used in this program come from
# a quantum mechanical many-body calculation of energies as functions of the number of particles.
# +
# For matrices and calculations
import numpy as np
# For machine learning (backend for keras)
import tensorflow as tf
# User-friendly machine learning library
# Front end for TensorFlow
import tensorflow.keras
# Different methods from Keras needed to create an RNN
# This is not necessary but it shortened function calls
# that need to be used in the code.
from tensorflow.keras import datasets, layers, models
from tensorflow.keras.layers import Input
from tensorflow.keras import regularizers
from tensorflow.keras.models import Model, Sequential
from tensorflow.keras.layers import Dense, SimpleRNN, LSTM, GRU
# For timing the code
from timeit import default_timer as timer
# For plotting
import matplotlib.pyplot as plt
# The data set
datatype='VaryDimension'
X_tot = np.arange(2, 42, 2)
y_tot = np.array([-0.03077640549, -0.08336233266, -0.1446729567, -0.2116753732, -0.2830637392, -0.3581341341, -0.436462435, -0.5177783846,
-0.6019067271, -0.6887363571, -0.7782028952, -0.8702784034, -0.9649652536, -1.062292565, -1.16231451,
-1.265109911, -1.370782966, -1.479465113, -1.591317992, -1.70653767])
# -
# ## Formatting the Data
#
# The way the recurrent neural networks are trained in this program
# differs from how machine learning algorithms are usually trained.
# Typically a machine learning algorithm is trained by learning the
# relationship between the x data and the y data. In this program, the
# recurrent neural network will be trained to recognize the relationship
# in a sequence of y values. This is type of data formatting is
# typically used time series forcasting, but it can also be used in any
# extrapolation (time series forecasting is just a specific type of
# extrapolation along the time axis). This method of data formatting
# does not use the x data and assumes that the y data are evenly spaced.
#
# For a standard machine learning algorithm, the training data has the
# form of (x,y) so the machine learning algorithm learns to assiciate a
# y value with a given x value. This is useful when the test data has x
# values within the same range as the training data. However, for this
# application, the x values of the test data are outside of the x values
# of the training data and the traditional method of training a machine
# learning algorithm does not work as well. For this reason, the
# recurrent neural network is trained on sequences of y values of the
# form ((y1, y2), y3), so that the network is concerned with learning
# the pattern of the y data and not the relation between the x and y
# data. As long as the pattern of y data outside of the training region
# stays relatively stable compared to what was inside the training
# region, this method of training can produce accurate extrapolations to
# y values far removed from the training data set.
#
# <!-- -->
# <!-- The idea behind formatting the data in this way comes from [this resource](https://machinelearningmastery.com/time-series-prediction-lstm-recurrent-neural-networks-python-keras/) and [this one](https://fairyonice.github.io/Understand-Keras%27s-RNN-behind-the-scenes-with-a-sin-wave-example.html). -->
# <!-- -->
# <!-- The following method takes in a y data set and formats it so the "x data" are of the form (y1, y2) and the "y data" are of the form y3, with extra brackets added in to make the resulting arrays compatable with both Keras and Tensorflow. -->
# <!-- -->
# <!-- Note: Using a sequence length of two is not required for time series forecasting so any lenght of sequence could be used (for example instead of ((y1, y2) y3) you could change the length of sequence to be 4 and the resulting data points would have the form ((y1, y2, y3, y4), y5)). While the following method can be used to create a data set of any sequence length, the remainder of the code expects the length of sequence to be 2. This is because the data sets are very small and the higher the lenght of the sequence the less resulting data points. -->
# +
# FORMAT_DATA
def format_data(data, length_of_sequence = 2):
"""
Inputs:
data(a numpy array): the data that will be the inputs to the recurrent neural
network
length_of_sequence (an int): the number of elements in one iteration of the
sequence patter. For a function approximator use length_of_sequence = 2.
Returns:
rnn_input (a 3D numpy array): the input data for the recurrent neural network. Its
dimensions are length of data - length of sequence, length of sequence,
dimnsion of data
rnn_output (a numpy array): the training data for the neural network
Formats data to be used in a recurrent neural network.
"""
X, Y = [], []
for i in range(len(data)-length_of_sequence):
# Get the next length_of_sequence elements
a = data[i:i+length_of_sequence]
# Get the element that immediately follows that
b = data[i+length_of_sequence]
# Reshape so that each data point is contained in its own array
a = np.reshape (a, (len(a), 1))
X.append(a)
Y.append(b)
rnn_input = np.array(X)
rnn_output = np.array(Y)
return rnn_input, rnn_output
# ## Defining the Recurrent Neural Network Using Keras
#
# The following method defines a simple recurrent neural network in keras consisting of one input layer, one hidden layer, and one output layer.
def rnn(length_of_sequences, batch_size = None, stateful = False):
"""
Inputs:
length_of_sequences (an int): the number of y values in "x data". This is determined
when the data is formatted
batch_size (an int): Default value is None. See Keras documentation of SimpleRNN.
stateful (a boolean): Default value is False. See Keras documentation of SimpleRNN.
Returns:
model (a Keras model): The recurrent neural network that is built and compiled by this
method
Builds and compiles a recurrent neural network with one hidden layer and returns the model.
"""
# Number of neurons in the input and output layers
in_out_neurons = 1
# Number of neurons in the hidden layer
hidden_neurons = 200
# Define the input layer
inp = Input(batch_shape=(batch_size,
length_of_sequences,
in_out_neurons))
# Define the hidden layer as a simple RNN layer with a set number of neurons and add it to
# the network immediately after the input layer
rnn = SimpleRNN(hidden_neurons,
return_sequences=False,
stateful = stateful,
name="RNN")(inp)
# Define the output layer as a dense neural network layer (standard neural network layer)
#and add it to the network immediately after the hidden layer.
dens = Dense(in_out_neurons,name="dense")(rnn)
# Create the machine learning model starting with the input layer and ending with the
# output layer
model = Model(inputs=[inp],outputs=[dens])
# Compile the machine learning model using the mean squared error function as the loss
# function and an Adams optimizer.
model.compile(loss="mean_squared_error", optimizer="adam")
return model
# -
# ## Predicting New Points With A Trained Recurrent Neural Network
# +
def test_rnn (x1, y_test, plot_min, plot_max):
"""
Inputs:
x1 (a list or numpy array): The complete x component of the data set
y_test (a list or numpy array): The complete y component of the data set
plot_min (an int or float): the smallest x value used in the training data
plot_max (an int or float): the largest x valye used in the training data
Returns:
None.
Uses a trained recurrent neural network model to predict future points in the
series. Computes the MSE of the predicted data set from the true data set, saves
the predicted data set to a csv file, and plots the predicted and true data sets w
while also displaying the data range used for training.
"""
# Add the training data as the first dim points in the predicted data array as these
# are known values.
y_pred = y_test[:dim].tolist()
# Generate the first input to the trained recurrent neural network using the last two
# points of the training data. Based on how the network was trained this means that it
# will predict the first point in the data set after the training data. All of the
# brackets are necessary for Tensorflow.
next_input = np.array([[[y_test[dim-2]], [y_test[dim-1]]]])
# Save the very last point in the training data set. This will be used later.
last = [y_test[dim-1]]
# Iterate until the complete data set is created.
for i in range (dim, len(y_test)):
# Predict the next point in the data set using the previous two points.
next = model.predict(next_input)
# Append just the number of the predicted data set
y_pred.append(next[0][0])
# Create the input that will be used to predict the next data point in the data set.
next_input = np.array([[last, next[0]]], dtype=np.float64)
last = next
# Print the mean squared error between the known data set and the predicted data set.
print('MSE: ', np.square(np.subtract(y_test, y_pred)).mean())
# Save the predicted data set as a csv file for later use
name = datatype + 'Predicted'+str(dim)+'.csv'
np.savetxt(name, y_pred, delimiter=',')
# Plot the known data set and the predicted data set. The red box represents the region that was used
# for the training data.
fig, ax = plt.subplots()
ax.plot(x1, y_test, label="true", linewidth=3)
ax.plot(x1, y_pred, 'g-.',label="predicted", linewidth=4)
ax.legend()
# Created a red region to represent the points used in the training data.
ax.axvspan(plot_min, plot_max, alpha=0.25, color='red')
plt.show()
# Check to make sure the data set is complete
assert len(X_tot) == len(y_tot)
# This is the number of points that will be used in as the training data
dim=12
# Separate the training data from the whole data set
X_train = X_tot[:dim]
y_train = y_tot[:dim]
# Generate the training data for the RNN, using a sequence of 2
rnn_input, rnn_training = format_data(y_train, 2)
# Create a recurrent neural network in Keras and produce a summary of the
# machine learning model
model = rnn(length_of_sequences = rnn_input.shape[1])
model.summary()
# Start the timer. Want to time training+testing
start = timer()
# Fit the model using the training data genenerated above using 150 training iterations and a 5%
# validation split. Setting verbose to True prints information about each training iteration.
hist = model.fit(rnn_input, rnn_training, batch_size=None, epochs=150,
verbose=True,validation_split=0.05)
for label in ["loss","val_loss"]:
plt.plot(hist.history[label],label=label)
plt.ylabel("loss")
plt.xlabel("epoch")
plt.title("The final validation loss: {}".format(hist.history["val_loss"][-1]))
plt.legend()
plt.show()
# Use the trained neural network to predict more points of the data set
test_rnn(X_tot, y_tot, X_tot[0], X_tot[dim-1])
# Stop the timer and calculate the total time needed.
end = timer()
print('Time: ', end-start)
# -
# ## Other Things to Try
#
# Changing the size of the recurrent neural network and its parameters
# can drastically change the results you get from the model. The below
# code takes the simple recurrent neural network from above and adds a
# second hidden layer, changes the number of neurons in the hidden
# layer, and explicitly declares the activation function of the hidden
# layers to be a sigmoid function. The loss function and optimizer can
# also be changed but are kept the same as the above network. These
# parameters can be tuned to provide the optimal result from the
# network. For some ideas on how to improve the performance of a
# [recurrent neural network](https://danijar.com/tips-for-training-recurrent-neural-networks).
# +
def rnn_2layers(length_of_sequences, batch_size = None, stateful = False):
"""
Inputs:
length_of_sequences (an int): the number of y values in "x data". This is determined
when the data is formatted
batch_size (an int): Default value is None. See Keras documentation of SimpleRNN.
stateful (a boolean): Default value is False. See Keras documentation of SimpleRNN.
Returns:
model (a Keras model): The recurrent neural network that is built and compiled by this
method
Builds and compiles a recurrent neural network with two hidden layers and returns the model.
"""
# Number of neurons in the input and output layers
in_out_neurons = 1
# Number of neurons in the hidden layer, increased from the first network
hidden_neurons = 500
# Define the input layer
inp = Input(batch_shape=(batch_size,
length_of_sequences,
in_out_neurons))
# Create two hidden layers instead of one hidden layer. Explicitly set the activation
# function to be the sigmoid function (the default value is hyperbolic tangent)
rnn1 = SimpleRNN(hidden_neurons,
return_sequences=True, # This needs to be True if another hidden layer is to follow
stateful = stateful, activation = 'sigmoid',
name="RNN1")(inp)
rnn2 = SimpleRNN(hidden_neurons,
return_sequences=False, activation = 'sigmoid',
stateful = stateful,
name="RNN2")(rnn1)
# Define the output layer as a dense neural network layer (standard neural network layer)
#and add it to the network immediately after the hidden layer.
dens = Dense(in_out_neurons,name="dense")(rnn2)
# Create the machine learning model starting with the input layer and ending with the
# output layer
model = Model(inputs=[inp],outputs=[dens])
# Compile the machine learning model using the mean squared error function as the loss
# function and an Adams optimizer.
model.compile(loss="mean_squared_error", optimizer="adam")
return model
# Check to make sure the data set is complete
assert len(X_tot) == len(y_tot)
# This is the number of points that will be used in as the training data
dim=12
# Separate the training data from the whole data set
X_train = X_tot[:dim]
y_train = y_tot[:dim]
# Generate the training data for the RNN, using a sequence of 2
rnn_input, rnn_training = format_data(y_train, 2)
# Create a recurrent neural network in Keras and produce a summary of the
# machine learning model
model = rnn_2layers(length_of_sequences = 2)
model.summary()
# Start the timer. Want to time training+testing
start = timer()
# Fit the model using the training data genenerated above using 150 training iterations and a 5%
# validation split. Setting verbose to True prints information about each training iteration.
hist = model.fit(rnn_input, rnn_training, batch_size=None, epochs=150,
verbose=True,validation_split=0.05)
# This section plots the training loss and the validation loss as a function of training iteration.
# This is not required for analyzing the couple cluster data but can help determine if the network is
# being overtrained.
for label in ["loss","val_loss"]:
plt.plot(hist.history[label],label=label)
plt.ylabel("loss")
plt.xlabel("epoch")
plt.title("The final validation loss: {}".format(hist.history["val_loss"][-1]))
plt.legend()
plt.show()
# Use the trained neural network to predict more points of the data set
test_rnn(X_tot, y_tot, X_tot[0], X_tot[dim-1])
# Stop the timer and calculate the total time needed.
end = timer()
print('Time: ', end-start)
# -
# ## Other Types of Recurrent Neural Networks
#
# Besides a simple recurrent neural network layer, there are two other
# commonly used types of recurrent neural network layers: Long Short
# Term Memory (LSTM) and Gated Recurrent Unit (GRU). For a short
# introduction to these layers see <https://medium.com/mindboard/lstm-vs-gru-experimental-comparison-955820c21e8b>
# and <https://medium.com/mindboard/lstm-vs-gru-experimental-comparison-955820c21e8b>.
#
# The first network created below is similar to the previous network,
# but it replaces the SimpleRNN layers with LSTM layers. The second
# network below has two hidden layers made up of GRUs, which are
# preceeded by two dense (feeddorward) neural network layers. These
# dense layers "preprocess" the data before it reaches the recurrent
# layers. This architecture has been shown to improve the performance
# of recurrent neural networks (see the link above and also
# <https://arxiv.org/pdf/1807.02857.pdf>.
# +
def lstm_2layers(length_of_sequences, batch_size = None, stateful = False):
"""
Inputs:
length_of_sequences (an int): the number of y values in "x data". This is determined
when the data is formatted
batch_size (an int): Default value is None. See Keras documentation of SimpleRNN.
stateful (a boolean): Default value is False. See Keras documentation of SimpleRNN.
Returns:
model (a Keras model): The recurrent neural network that is built and compiled by this
method
Builds and compiles a recurrent neural network with two LSTM hidden layers and returns the model.
"""
# Number of neurons on the input/output layer and the number of neurons in the hidden layer
in_out_neurons = 1
hidden_neurons = 250
# Input Layer
inp = Input(batch_shape=(batch_size,
length_of_sequences,
in_out_neurons))
# Hidden layers (in this case they are LSTM layers instead if SimpleRNN layers)
rnn= LSTM(hidden_neurons,
return_sequences=True,
stateful = stateful,
name="RNN", use_bias=True, activation='tanh')(inp)
rnn1 = LSTM(hidden_neurons,
return_sequences=False,
stateful = stateful,
name="RNN1", use_bias=True, activation='tanh')(rnn)
# Output layer
dens = Dense(in_out_neurons,name="dense")(rnn1)
# Define the midel
model = Model(inputs=[inp],outputs=[dens])
# Compile the model
model.compile(loss='mean_squared_error', optimizer='adam')
# Return the model
return model
def dnn2_gru2(length_of_sequences, batch_size = None, stateful = False):
"""
Inputs:
length_of_sequences (an int): the number of y values in "x data". This is determined
when the data is formatted
batch_size (an int): Default value is None. See Keras documentation of SimpleRNN.
stateful (a boolean): Default value is False. See Keras documentation of SimpleRNN.
Returns:
model (a Keras model): The recurrent neural network that is built and compiled by this
method
Builds and compiles a recurrent neural network with four hidden layers (two dense followed by
two GRU layers) and returns the model.
"""
# Number of neurons on the input/output layers and hidden layers
in_out_neurons = 1
hidden_neurons = 250
# Input layer
inp = Input(batch_shape=(batch_size,
length_of_sequences,
in_out_neurons))
# Hidden Dense (feedforward) layers
dnn = Dense(hidden_neurons/2, activation='relu', name='dnn')(inp)
dnn1 = Dense(hidden_neurons/2, activation='relu', name='dnn1')(dnn)
# Hidden GRU layers
rnn1 = GRU(hidden_neurons,
return_sequences=True,
stateful = stateful,
name="RNN1", use_bias=True)(dnn1)
rnn = GRU(hidden_neurons,
return_sequences=False,
stateful = stateful,
name="RNN", use_bias=True)(rnn1)
# Output layer
dens = Dense(in_out_neurons,name="dense")(rnn)
# Define the model
model = Model(inputs=[inp],outputs=[dens])
# Compile the mdoel
model.compile(loss='mean_squared_error', optimizer='adam')
# Return the model
return model
# Check to make sure the data set is complete
assert len(X_tot) == len(y_tot)
# This is the number of points that will be used in as the training data
dim=12
# Separate the training data from the whole data set
X_train = X_tot[:dim]
y_train = y_tot[:dim]
# Generate the training data for the RNN, using a sequence of 2
rnn_input, rnn_training = format_data(y_train, 2)
# Create a recurrent neural network in Keras and produce a summary of the
# machine learning model
# Change the method name to reflect which network you want to use
model = dnn2_gru2(length_of_sequences = 2)
model.summary()
# Start the timer. Want to time training+testing
start = timer()
# Fit the model using the training data genenerated above using 150 training iterations and a 5%
# validation split. Setting verbose to True prints information about each training iteration.
hist = model.fit(rnn_input, rnn_training, batch_size=None, epochs=150,
verbose=True,validation_split=0.05)
# This section plots the training loss and the validation loss as a function of training iteration.
# This is not required for analyzing the couple cluster data but can help determine if the network is
# being overtrained.
for label in ["loss","val_loss"]:
plt.plot(hist.history[label],label=label)
plt.ylabel("loss")
plt.xlabel("epoch")
plt.title("The final validation loss: {}".format(hist.history["val_loss"][-1]))
plt.legend()
plt.show()
# Use the trained neural network to predict more points of the data set
test_rnn(X_tot, y_tot, X_tot[0], X_tot[dim-1])
# Stop the timer and calculate the total time needed.
end = timer()
print('Time: ', end-start)
# ### Training Recurrent Neural Networks in the Standard Way (i.e. learning the relationship between the X and Y data)
#
# Finally, comparing the performace of a recurrent neural network using the standard data formatting to the performance of the network with time sequence data formatting shows the benefit of this type of data formatting with extrapolation.
# Check to make sure the data set is complete
assert len(X_tot) == len(y_tot)
# This is the number of points that will be used in as the training data
dim=12
# Separate the training data from the whole data set
X_train = X_tot[:dim]
y_train = y_tot[:dim]
# Reshape the data for Keras specifications
X_train = X_train.reshape((dim, 1))
y_train = y_train.reshape((dim, 1))
# Create a recurrent neural network in Keras and produce a summary of the
# machine learning model
# Set the sequence length to 1 for regular data formatting
model = rnn(length_of_sequences = 1)
model.summary()
# Start the timer. Want to time training+testing
start = timer()
# Fit the model using the training data genenerated above using 150 training iterations and a 5%
# validation split. Setting verbose to True prints information about each training iteration.
hist = model.fit(X_train, y_train, batch_size=None, epochs=150,
verbose=True,validation_split=0.05)
# This section plots the training loss and the validation loss as a function of training iteration.
# This is not required for analyzing the couple cluster data but can help determine if the network is
# being overtrained.
for label in ["loss","val_loss"]:
plt.plot(hist.history[label],label=label)
plt.ylabel("loss")
plt.xlabel("epoch")
plt.title("The final validation loss: {}".format(hist.history["val_loss"][-1]))
plt.legend()
plt.show()
# Use the trained neural network to predict the remaining data points
X_pred = X_tot[dim:]
X_pred = X_pred.reshape((len(X_pred), 1))
y_model = model.predict(X_pred)
y_pred = np.concatenate((y_tot[:dim], y_model.flatten()))
# Plot the known data set and the predicted data set. The red box represents the region that was used
# for the training data.
fig, ax = plt.subplots()
ax.plot(X_tot, y_tot, label="true", linewidth=3)
ax.plot(X_tot, y_pred, 'g-.',label="predicted", linewidth=4)
ax.legend()
# Created a red region to represent the points used in the training data.
ax.axvspan(X_tot[0], X_tot[dim], alpha=0.25, color='red')
plt.show()
# Stop the timer and calculate the total time needed.
end = timer()
print('Time: ', end-start)
# -
# ## Generative Models
#
# **Generative models** describe a class of statistical models that are a contrast
# to **discriminative models**. Informally we say that generative models can
# generate new data instances while discriminative models discriminate between
# different kinds of data instances. A generative model could generate new photos
# of animals that look like 'real' animals while a discriminative model could tell
# a dog from a cat. More formally, given a data set $x$ and a set of labels /
# targets $y$. Generative models capture the joint probability $p(x, y)$, or
# just $p(x)$ if there are no labels, while discriminative models capture the
# conditional probability $p(y | x)$. Discriminative models generally try to draw
# boundaries in the data space (often high dimensional), while generative models
# try to model how data is placed throughout the space.
#
# **Note**: this material is thanks to <NAME>.
# ## Generative Adversarial Networks
#
# **Generative Adversarial Networks** are a type of unsupervised machine learning
# algorithm proposed by [Goodfellow et. al](https://arxiv.org/pdf/1406.2661.pdf)
# in 2014 (short and good article).
#
# The simplest formulation of
# the model is based on a game theoretic approach, *zero sum game*, where we pit
# two neural networks against one another. We define two rival networks, one
# generator $g$, and one discriminator $d$. The generator directly produces
# samples
# <!-- Equation labels as ordinary links -->
# <div id="_auto1"></div>
#
# $$
# \begin{equation}
# x = g(z; \theta^{(g)})
# \label{_auto1} \tag{1}
# \end{equation}
# $$
# ## Discriminator
# The discriminator attempts to distinguish between samples drawn from the
# training data and samples drawn from the generator. In other words, it tries to
# tell the difference between the fake data produced by $g$ and the actual data
# samples we want to do prediction on. The discriminator outputs a probability
# value given by
# <!-- Equation labels as ordinary links -->
# <div id="_auto2"></div>
#
# $$
# \begin{equation}
# d(x; \theta^{(d)})
# \label{_auto2} \tag{2}
# \end{equation}
# $$
# indicating the probability that $x$ is a real training example rather than a
# fake sample the generator has generated. The simplest way to formulate the
# learning process in a generative adversarial network is a zero-sum game, in
# which a function
# <!-- Equation labels as ordinary links -->
# <div id="_auto3"></div>
#
# $$
# \begin{equation}
# v(\theta^{(g)}, \theta^{(d)})
# \label{_auto3} \tag{3}
# \end{equation}
# $$
# determines the reward for the discriminator, while the generator gets the
# conjugate reward
# <!-- Equation labels as ordinary links -->
# <div id="_auto4"></div>
#
# $$
# \begin{equation}
# -v(\theta^{(g)}, \theta^{(d)})
# \label{_auto4} \tag{4}
# \end{equation}
# $$
# ## Learning Process
#
# During learning both of the networks maximize their own reward function, so that
# the generator gets better and better at tricking the discriminator, while the
# discriminator gets better and better at telling the difference between the fake
# and real data. The generator and discriminator alternate on which one trains at
# one time (i.e. for one epoch). In other words, we keep the generator constant
# and train the discriminator, then we keep the discriminator constant to train
# the generator and repeat. It is this back and forth dynamic which lets GANs
# tackle otherwise intractable generative problems. As the generator improves with
# training, the discriminator's performance gets worse because it cannot easily
# tell the difference between real and fake. If the generator ends up succeeding
# perfectly, the the discriminator will do no better than random guessing i.e.
# 50\%. This progression in the training poses a problem for the convergence
# criteria for GANs. The discriminator feedback gets less meaningful over time,
# if we continue training after this point then the generator is effectively
# training on junk data which can undo the learning up to that point. Therefore,
# we stop training when the discriminator starts outputting $1/2$ everywhere.
# ## More about the Learning Process
#
# At convergence we have
# <!-- Equation labels as ordinary links -->
# <div id="_auto5"></div>
#
# $$
# \begin{equation}
# g^* = \underset{g}{\mathrm{argmin}}\hspace{2pt}
# \underset{d}{\mathrm{max}}v(\theta^{(g)}, \theta^{(d)})
# \label{_auto5} \tag{5}
# \end{equation}
# $$
# The default choice for $v$ is
# <!-- Equation labels as ordinary links -->
# <div id="_auto6"></div>
#
# $$
# \begin{equation}
# v(\theta^{(g)}, \theta^{(d)}) = \mathbb{E}_{x\sim p_\mathrm{data}}\log d(x)
# + \mathbb{E}_{x\sim p_\mathrm{model}}
# \log (1 - d(x))
# \label{_auto6} \tag{6}
# \end{equation}
# $$
# The main motivation for the design of GANs is that the learning process requires
# neither approximate inference (variational autoencoders for example) nor
# approximation of a partition function. In the case where
# <!-- Equation labels as ordinary links -->
# <div id="_auto7"></div>
#
# $$
# \begin{equation}
# \underset{d}{\mathrm{max}}v(\theta^{(g)}, \theta^{(d)})
# \label{_auto7} \tag{7}
# \end{equation}
# $$
# is convex in $\theta^{(g)} then the procedure is guaranteed to converge and is
# asymptotically consistent
# ( [<NAME> on QuGANs](https://arxiv.org/pdf/1804.09139.pdf) ).
# ## Additional References
# This is in
# general not the case and it is possible to get situations where the training
# process never converges because the generator and discriminator chase one
# another around in the parameter space indefinitely. A much deeper discussion on
# the currently open research problem of GAN convergence is available
# [here](https://www.deeplearningbook.org/contents/generative_models.html). To
# anyone interested in learning more about GANs it is a highly recommended read.
# Direct quote: "In this best-performing formulation, the generator aims to
# increase the log probability that the discriminator makes a mistake, rather than
# aiming to decrease the log probability that the discriminator makes the correct
# prediction." [Another interesting read](https://arxiv.org/abs/1701.00160)
# ## Writing Our First Generative Adversarial Network
# Let us now move on to actually implementing a GAN in tensorflow. We will study
# the performance of our GAN on the MNIST dataset. This code is based on and
# adapted from the
# [google tutorial](https://www.tensorflow.org/tutorials/generative/dcgan)
#
# First we import our libraries
import os
import time
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
from tensorflow.keras import layers
from tensorflow.keras.utils import plot_model
# Next we define our hyperparameters and import our data the usual way
# +
BUFFER_SIZE = 60000
BATCH_SIZE = 256
EPOCHS = 30
data = tf.keras.datasets.mnist.load_data()
(train_images, train_labels), (test_images, test_labels) = data
train_images = np.reshape(train_images, (train_images.shape[0],
28,
28,
1)).astype('float32')
# we normalize between -1 and 1
train_images = (train_images - 127.5) / 127.5
training_dataset = tf.data.Dataset.from_tensor_slices(
train_images).shuffle(BUFFER_SIZE).batch(BATCH_SIZE)
# -
# ## MNIST and GANs
#
# Let's have a quick look
plt.imshow(train_images[0], cmap='Greys')
plt.show()
# Now we define our two models. This is where the 'magic' happens. There are a
# huge amount of possible formulations for both models. A lot of engineering and
# trial and error can be done here to try to produce better performing models. For
# more advanced GANs this is by far the step where you can 'make or break' a
# model.
#
# We start with the generator. As stated in the introductory text the generator
# $g$ upsamples from a random sample to the shape of what we want to predict. In
# our case we are trying to predict MNIST images ($28\times 28$ pixels).
def generator_model():
"""
The generator uses upsampling layers tf.keras.layers.Conv2DTranspose() to
produce an image from a random seed. We start with a Dense layer taking this
random sample as an input and subsequently upsample through multiple
convolutional layers.
"""
# we define our model
model = tf.keras.Sequential()
# adding our input layer. Dense means that every neuron is connected and
# the input shape is the shape of our random noise. The units need to match
# in some sense the upsampling strides to reach our desired output shape.
# we are using 100 random numbers as our seed
model.add(layers.Dense(units=7*7*BATCH_SIZE,
use_bias=False,
input_shape=(100, )))
# we normalize the output form the Dense layer
model.add(layers.BatchNormalization())
# and add an activation function to our 'layer'. LeakyReLU avoids vanishing
# gradient problem
model.add(layers.LeakyReLU())
model.add(layers.Reshape((7, 7, BATCH_SIZE)))
assert model.output_shape == (None, 7, 7, BATCH_SIZE)
# even though we just added four keras layers we think of everything above
# as 'one' layer
# next we add our upscaling convolutional layers
model.add(layers.Conv2DTranspose(filters=128,
kernel_size=(5, 5),
strides=(1, 1),
padding='same',
use_bias=False))
model.add(layers.BatchNormalization())
model.add(layers.LeakyReLU())
assert model.output_shape == (None, 7, 7, 128)
model.add(layers.Conv2DTranspose(filters=64,
kernel_size=(5, 5),
strides=(2, 2),
padding='same',
use_bias=False))
model.add(layers.BatchNormalization())
model.add(layers.LeakyReLU())
assert model.output_shape == (None, 14, 14, 64)
model.add(layers.Conv2DTranspose(filters=1,
kernel_size=(5, 5),
strides=(2, 2),
padding='same',
use_bias=False,
activation='tanh'))
assert model.output_shape == (None, 28, 28, 1)
return model
# And there we have our 'simple' generator model. Now we move on to defining our
# discriminator model $d$, which is a convolutional neural network based image
# classifier.
def discriminator_model():
"""
The discriminator is a convolutional neural network based image classifier
"""
# we define our model
model = tf.keras.Sequential()
model.add(layers.Conv2D(filters=64,
kernel_size=(5, 5),
strides=(2, 2),
padding='same',
input_shape=[28, 28, 1]))
model.add(layers.LeakyReLU())
# adding a dropout layer as you do in conv-nets
model.add(layers.Dropout(0.3))
model.add(layers.Conv2D(filters=128,
kernel_size=(5, 5),
strides=(2, 2),
padding='same'))
model.add(layers.LeakyReLU())
# adding a dropout layer as you do in conv-nets
model.add(layers.Dropout(0.3))
model.add(layers.Flatten())
model.add(layers.Dense(1))
return model
# ## Other Models
# Let us take a look at our models. **Note**: double click images for bigger view.
generator = generator_model()
plot_model(generator, show_shapes=True, rankdir='LR')
discriminator = discriminator_model()
plot_model(discriminator, show_shapes=True, rankdir='LR')
# Next we need a few helper objects we will use in training
cross_entropy = tf.keras.losses.BinaryCrossentropy(from_logits=True)
generator_optimizer = tf.keras.optimizers.Adam(1e-4)
discriminator_optimizer = tf.keras.optimizers.Adam(1e-4)
# The first object, *cross_entropy* is our loss function and the two others are
# our optimizers. Notice we use the same learning rate for both $g$ and $d$. This
# is because they need to improve their accuracy at approximately equal speeds to
# get convergence (not necessarily exactly equal). Now we define our loss
# functions
def generator_loss(fake_output):
loss = cross_entropy(tf.ones_like(fake_output), fake_output)
return loss
def discriminator_loss(real_output, fake_output):
real_loss = cross_entropy(tf.ones_like(real_output), real_output)
fake_loss = cross_entropy(tf.zeros_liks(fake_output), fake_output)
total_loss = real_loss + fake_loss
return total_loss
# Next we define a kind of seed to help us compare the learning process over
# multiple training epochs.
noise_dimension = 100
n_examples_to_generate = 16
seed_images = tf.random.normal([n_examples_to_generate, noise_dimension])
# ## Training Step
#
# Now we have everything we need to define our training step, which we will apply
# for every step in our training loop. Notice the @tf.function flag signifying
# that the function is tensorflow 'compiled'. Removing this flag doubles the
# computation time.
@tf.function
def train_step(images):
noise = tf.random.normal([BATCH_SIZE, noise_dimension])
with tf.GradientTape() as gen_tape, tf.GradientTape() as disc_tape:
generated_images = generator(noise, training=True)
real_output = discriminator(images, training=True)
fake_output = discriminator(generated_images, training=True)
gen_loss = generator_loss(fake_output)
disc_loss = discriminator_loss(real_output, fake_output)
gradients_of_generator = gen_tape.gradient(gen_loss,
generator.trainable_variables)
gradients_of_discriminator = disc_tape.gradient(disc_loss,
discriminator.trainable_variables)
generator_optimizer.apply_gradients(zip(gradients_of_generator,
generator.trainable_variables))
discriminator_optimizer.apply_gradients(zip(gradients_of_discriminator,
discriminator.trainable_variables))
return gen_loss, disc_loss
# Next we define a helper function to produce an output over our training epochs
# to see the predictive progression of our generator model. **Note**: I am including
# this code here, but comment it out in the training loop.
def generate_and_save_images(model, epoch, test_input):
# we're making inferences here
predictions = model(test_input, training=False)
fig = plt.figure(figsize=(4, 4))
for i in range(predictions.shape[0]):
plt.subplot(4, 4, i+1)
plt.imshow(predictions[i, :, :, 0] * 127.5 + 127.5, cmap='gray')
plt.axis('off')
plt.savefig(f'./images_from_seed_images/image_at_epoch_{str(epoch).zfill(3)}.png')
plt.close()
#plt.show()
# ## Checkpoints
# Setting up checkpoints to periodically save our model during training so that
# everything is not lost even if the program were to somehow terminate while
# training.
# Setting up checkpoints to save model during training
checkpoint_dir = './training_checkpoints'
checkpoint_prefix = os.path.join(checkpoint_dir, 'ckpt')
checkpoint = tf.train.Checkpoint(generator_optimizer=generator_optimizer,
discriminator_optimizer=discriminator_optimizer,
generator=generator,
discriminator=discriminator)
# Now we define our training loop
def train(dataset, epochs):
generator_loss_list = []
discriminator_loss_list = []
for epoch in range(epochs):
start = time.time()
for image_batch in dataset:
gen_loss, disc_loss = train_step(image_batch)
generator_loss_list.append(gen_loss.numpy())
discriminator_loss_list.append(disc_loss.numpy())
#generate_and_save_images(generator, epoch + 1, seed_images)
if (epoch + 1) % 15 == 0:
checkpoint.save(file_prefix=checkpoint_prefix)
print(f'Time for epoch {epoch} is {time.time() - start}')
#generate_and_save_images(generator, epochs, seed_images)
loss_file = './data/lossfile.txt'
with open(loss_file, 'w') as outfile:
outfile.write(str(generator_loss_list))
outfile.write('\n')
outfile.write('\n')
outfile.write(str(discriminator_loss_list))
outfile.write('\n')
outfile.write('\n')
# To train simply call this function. **Warning**: this might take a long time so
# there is a folder of a pretrained network already included in the repository.
train(train_dataset, EPOCHS)
# And here is the result of training our model for 100 epochs
#
# <!-- dom:MOVIE: [images_from_seed_images/generation.gif] -->
# <!-- begin movie -->
from IPython.display import HTML
_s = """
<embed src="images_from_seed_images/generation.gif" autoplay="false" loop="true"></embed>
<p><em></em></p>
"""
HTML(_s)
# <!-- end movie -->
#
# Now to avoid having to train and everything, which will take a while depending
# on your computer setup we now load in the model which produced the above gif.
# +
checkpoint.restore(tf.train.latest_checkpoint(checkpoint_dir))
restored_generator = checkpoint.generator
restored_discriminator = checkpoint.discriminator
print(restored_generator)
print(restored_discriminator)
# -
# ## Exploring the Latent Space
#
# We have successfully loaded in our latest model. Let us now play around a bit
# and see what kind of things we can learn about this model. Our generator takes
# an array of 100 numbers. One idea can be to try to systematically change our
# input. Let us try and see what we get
# +
def generate_latent_points(number=100, scale_means=1, scale_stds=1):
latent_dim = 100
means = scale_means * tf.linspace(-1, 1, num=latent_dim)
stds = scale_stds * tf.linspace(-1, 1, num=latent_dim)
latent_space_value_range = tf.random.normal([number, latent_dim],
means,
stds,
dtype=tf.float64)
return latent_space_value_range
def generate_images(latent_points):
# notice we set training to false because we are making inferences
generated_images = restored_generator.predict(latent_points)
return generated_images
# -
def plot_result(generated_images, number=100):
# obviously this assumes sqrt number is an int
fig, axs = plt.subplots(int(np.sqrt(number)), int(np.sqrt(number)),
figsize=(10, 10))
for i in range(int(np.sqrt(number))):
for j in range(int(np.sqrt(number))):
axs[i, j].imshow(generated_images[i*j], cmap='Greys')
axs[i, j].axis('off')
plt.show()
generated_images = generate_images(generate_latent_points())
plot_result(generated_images)
# ## Getting Results
# We see that the generator generates images that look like MNIST
# numbers: $1, 4, 7, 9$. Let's try to tweak it a bit more to see if we are able
# to generate a similar plot where we generate every MNIST number. Let us now try
# to 'move' a bit around in the latent space. **Note**: decrease the plot number if
# these following cells take too long to run on your computer.
# +
plot_number = 225
generated_images = generate_images(generate_latent_points(number=plot_number,
scale_means=5,
scale_stds=1))
plot_result(generated_images, number=plot_number)
generated_images = generate_images(generate_latent_points(number=plot_number,
scale_means=-5,
scale_stds=1))
plot_result(generated_images, number=plot_number)
generated_images = generate_images(generate_latent_points(number=plot_number,
scale_means=1,
scale_stds=5))
plot_result(generated_images, number=plot_number)
# -
# Again, we have found something interesting. *Moving* around using our means
# takes us from digit to digit, while *moving* around using our standard
# deviations seem to increase the number of different digits! In the last image
# above, we can barely make out every MNIST digit. Let us make on last plot using
# this information by upping the standard deviation of our Gaussian noises.
plot_number = 400
generated_images = generate_images(generate_latent_points(number=plot_number,
scale_means=1,
scale_stds=10))
plot_result(generated_images, number=plot_number)
# A pretty cool result! We see that our generator indeed has learned a
# distribution which qualitatively looks a whole lot like the MNIST dataset.
# ## Interpolating Between MNIST Digits
# Another interesting way to explore the latent space of our generator model is by
# interpolating between the MNIST digits. This section is largely based on
# [this excellent blogpost](https://machinelearningmastery.com/how-to-interpolate-and-perform-vector-arithmetic-with-faces-using-a-generative-adversarial-network/)
# by <NAME>.
#
# So let us start by defining a function to interpolate between two points in the
# latent space.
def interpolation(point_1, point_2, n_steps=10):
ratios = np.linspace(0, 1, num=n_steps)
vectors = []
for i, ratio in enumerate(ratios):
vectors.append(((1.0 - ratio) * point_1 + ratio * point_2))
return tf.stack(vectors)
# Now we have all we need to do our interpolation analysis.
# +
plot_number = 100
latent_points = generate_latent_points(number=plot_number)
results = None
for i in range(0, 2*np.sqrt(plot_number), 2):
interpolated = interpolation(latent_points[i], latent_points[i+1])
generated_images = generate_images(interpolated)
if results is None:
results = generated_images
else:
results = tf.stack((results, generated_images))
plot_results(results, plot_number)
# -
# ## Basic ideas of the Principal Component Analysis (PCA)
#
# The principal component analysis deals with the problem of fitting a
# low-dimensional affine subspace $S$ of dimension $d$ much smaller than
# the total dimension $D$ of the problem at hand (our data
# set). Mathematically it can be formulated as a statistical problem or
# a geometric problem. In our discussion of the theorem for the
# classical PCA, we will stay with a statistical approach.
# Historically, the PCA was first formulated in a statistical setting in order to estimate the principal component of a multivariate random variable.
#
# We have a data set defined by a design/feature matrix $\boldsymbol{X}$ (see below for its definition)
# * Each data point is determined by $p$ extrinsic (measurement) variables
#
# * We may want to ask the following question: Are there fewer intrinsic variables (say $d << p$) that still approximately describe the data?
#
# * If so, these intrinsic variables may tell us something important and finding these intrinsic variables is what dimension reduction methods do.
#
# A good read is for example [<NAME> and Sastry](https://www.springer.com/gp/book/9780387878102).
# ## Introducing the Covariance and Correlation functions
#
# Before we discuss the PCA theorem, we need to remind ourselves about
# the definition of the covariance and the correlation function. These are quantities
#
# Suppose we have defined two vectors
# $\hat{x}$ and $\hat{y}$ with $n$ elements each. The covariance matrix $\boldsymbol{C}$ is defined as
# $$
# \boldsymbol{C}[\boldsymbol{x},\boldsymbol{y}] = \begin{bmatrix} \mathrm{cov}[\boldsymbol{x},\boldsymbol{x}] & \mathrm{cov}[\boldsymbol{x},\boldsymbol{y}] \\
# \mathrm{cov}[\boldsymbol{y},\boldsymbol{x}] & \mathrm{cov}[\boldsymbol{y},\boldsymbol{y}] \\
# \end{bmatrix},
# $$
# where for example
# $$
# \mathrm{cov}[\boldsymbol{x},\boldsymbol{y}] =\frac{1}{n} \sum_{i=0}^{n-1}(x_i- \overline{x})(y_i- \overline{y}).
# $$
# With this definition and recalling that the variance is defined as
# $$
# \mathrm{var}[\boldsymbol{x}]=\frac{1}{n} \sum_{i=0}^{n-1}(x_i- \overline{x})^2,
# $$
# we can rewrite the covariance matrix as
# $$
# \boldsymbol{C}[\boldsymbol{x},\boldsymbol{y}] = \begin{bmatrix} \mathrm{var}[\boldsymbol{x}] & \mathrm{cov}[\boldsymbol{x},\boldsymbol{y}] \\
# \mathrm{cov}[\boldsymbol{x},\boldsymbol{y}] & \mathrm{var}[\boldsymbol{y}] \\
# \end{bmatrix}.
# $$
# ## More on the covariance
# The covariance takes values between zero and infinity and may thus
# lead to problems with loss of numerical precision for particularly
# large values. It is common to scale the covariance matrix by
# introducing instead the correlation matrix defined via the so-called
# correlation function
# $$
# \mathrm{corr}[\boldsymbol{x},\boldsymbol{y}]=\frac{\mathrm{cov}[\boldsymbol{x},\boldsymbol{y}]}{\sqrt{\mathrm{var}[\boldsymbol{x}] \mathrm{var}[\boldsymbol{y}]}}.
# $$
# The correlation function is then given by values $\mathrm{corr}[\boldsymbol{x},\boldsymbol{y}]
# \in [-1,1]$. This avoids eventual problems with too large values. We
# can then define the correlation matrix for the two vectors $\boldsymbol{x}$
# and $\boldsymbol{y}$ as
# $$
# \boldsymbol{K}[\boldsymbol{x},\boldsymbol{y}] = \begin{bmatrix} 1 & \mathrm{corr}[\boldsymbol{x},\boldsymbol{y}] \\
# \mathrm{corr}[\boldsymbol{y},\boldsymbol{x}] & 1 \\
# \end{bmatrix},
# $$
# In the above example this is the function we constructed using **pandas**.
# ## Reminding ourselves about Linear Regression
# In our derivation of the various regression algorithms like **Ordinary Least Squares** or **Ridge regression**
# we defined the design/feature matrix $\boldsymbol{X}$ as
# $$
# \boldsymbol{X}=\begin{bmatrix}
# x_{0,0} & x_{0,1} & x_{0,2}& \dots & \dots x_{0,p-1}\\
# x_{1,0} & x_{1,1} & x_{1,2}& \dots & \dots x_{1,p-1}\\
# x_{2,0} & x_{2,1} & x_{2,2}& \dots & \dots x_{2,p-1}\\
# \dots & \dots & \dots & \dots \dots & \dots \\
# x_{n-2,0} & x_{n-2,1} & x_{n-2,2}& \dots & \dots x_{n-2,p-1}\\
# x_{n-1,0} & x_{n-1,1} & x_{n-1,2}& \dots & \dots x_{n-1,p-1}\\
# \end{bmatrix},
# $$
# with $\boldsymbol{X}\in {\mathbb{R}}^{n\times p}$, with the predictors/features $p$ refering to the column numbers and the
# entries $n$ being the row elements.
# We can rewrite the design/feature matrix in terms of its column vectors as
# $$
# \boldsymbol{X}=\begin{bmatrix} \boldsymbol{x}_0 & \boldsymbol{x}_1 & \boldsymbol{x}_2 & \dots & \dots & \boldsymbol{x}_{p-1}\end{bmatrix},
# $$
# with a given vector
# $$
# \boldsymbol{x}_i^T = \begin{bmatrix}x_{0,i} & x_{1,i} & x_{2,i}& \dots & \dots x_{n-1,i}\end{bmatrix}.
# $$
# ## Simple Example
# With these definitions, we can now rewrite our $2\times 2$
# correlation/covariance matrix in terms of a moe general design/feature
# matrix $\boldsymbol{X}\in {\mathbb{R}}^{n\times p}$. This leads to a $p\times p$
# covariance matrix for the vectors $\boldsymbol{x}_i$ with $i=0,1,\dots,p-1$
# $$
# \boldsymbol{C}[\boldsymbol{x}] = \begin{bmatrix}
# \mathrm{var}[\boldsymbol{x}_0] & \mathrm{cov}[\boldsymbol{x}_0,\boldsymbol{x}_1] & \mathrm{cov}[\boldsymbol{x}_0,\boldsymbol{x}_2] & \dots & \dots & \mathrm{cov}[\boldsymbol{x}_0,\boldsymbol{x}_{p-1}]\\
# \mathrm{cov}[\boldsymbol{x}_1,\boldsymbol{x}_0] & \mathrm{var}[\boldsymbol{x}_1] & \mathrm{cov}[\boldsymbol{x}_1,\boldsymbol{x}_2] & \dots & \dots & \mathrm{cov}[\boldsymbol{x}_1,\boldsymbol{x}_{p-1}]\\
# \mathrm{cov}[\boldsymbol{x}_2,\boldsymbol{x}_0] & \mathrm{cov}[\boldsymbol{x}_2,\boldsymbol{x}_1] & \mathrm{var}[\boldsymbol{x}_2] & \dots & \dots & \mathrm{cov}[\boldsymbol{x}_2,\boldsymbol{x}_{p-1}]\\
# \dots & \dots & \dots & \dots & \dots & \dots \\
# \dots & \dots & \dots & \dots & \dots & \dots \\
# \mathrm{cov}[\boldsymbol{x}_{p-1},\boldsymbol{x}_0] & \mathrm{cov}[\boldsymbol{x}_{p-1},\boldsymbol{x}_1] & \mathrm{cov}[\boldsymbol{x}_{p-1},\boldsymbol{x}_{2}] & \dots & \dots & \mathrm{var}[\boldsymbol{x}_{p-1}]\\
# \end{bmatrix},
# $$
# ## The Correlation Matrix
#
# and the correlation matrix
# $$
# \boldsymbol{K}[\boldsymbol{x}] = \begin{bmatrix}
# 1 & \mathrm{corr}[\boldsymbol{x}_0,\boldsymbol{x}_1] & \mathrm{corr}[\boldsymbol{x}_0,\boldsymbol{x}_2] & \dots & \dots & \mathrm{corr}[\boldsymbol{x}_0,\boldsymbol{x}_{p-1}]\\
# \mathrm{corr}[\boldsymbol{x}_1,\boldsymbol{x}_0] & 1 & \mathrm{corr}[\boldsymbol{x}_1,\boldsymbol{x}_2] & \dots & \dots & \mathrm{corr}[\boldsymbol{x}_1,\boldsymbol{x}_{p-1}]\\
# \mathrm{corr}[\boldsymbol{x}_2,\boldsymbol{x}_0] & \mathrm{corr}[\boldsymbol{x}_2,\boldsymbol{x}_1] & 1 & \dots & \dots & \mathrm{corr}[\boldsymbol{x}_2,\boldsymbol{x}_{p-1}]\\
# \dots & \dots & \dots & \dots & \dots & \dots \\
# \dots & \dots & \dots & \dots & \dots & \dots \\
# \mathrm{corr}[\boldsymbol{x}_{p-1},\boldsymbol{x}_0] & \mathrm{corr}[\boldsymbol{x}_{p-1},\boldsymbol{x}_1] & \mathrm{corr}[\boldsymbol{x}_{p-1},\boldsymbol{x}_{2}] & \dots & \dots & 1\\
# \end{bmatrix},
# $$
# ## Numpy Functionality
#
# The Numpy function **np.cov** calculates the covariance elements using
# the factor $1/(n-1)$ instead of $1/n$ since it assumes we do not have
# the exact mean values. The following simple function uses the
# **np.vstack** function which takes each vector of dimension $1\times n$
# and produces a $2\times n$ matrix $\boldsymbol{W}$
# $$
# \boldsymbol{W}^T = \begin{bmatrix} x_0 & y_0 \\
# x_1 & y_1 \\
# x_2 & y_2\\
# \dots & \dots \\
# x_{n-2} & y_{n-2}\\
# x_{n-1} & y_{n-1} &
# \end{bmatrix},
# $$
# which in turn is converted into into the $2\times 2$ covariance matrix
# $\boldsymbol{C}$ via the Numpy function **np.cov()**. We note that we can also calculate
# the mean value of each set of samples $\boldsymbol{x}$ etc using the Numpy
# function **np.mean(x)**. We can also extract the eigenvalues of the
# covariance matrix through the **np.linalg.eig()** function.
# Importing various packages
import numpy as np
n = 100
x = np.random.normal(size=n)
print(np.mean(x))
y = 4+3*x+np.random.normal(size=n)
print(np.mean(y))
W = np.vstack((x, y))
C = np.cov(W)
print(C)
# ## Correlation Matrix again
#
# The previous example can be converted into the correlation matrix by
# simply scaling the matrix elements with the variances. We should also
# subtract the mean values for each column. This leads to the following
# code which sets up the correlations matrix for the previous example in
# a more brute force way. Here we scale the mean values for each column of the design matrix, calculate the relevant mean values and variances and then finally set up the $2\times 2$ correlation matrix (since we have only two vectors).
import numpy as np
n = 100
# define two vectors
x = np.random.random(size=n)
y = 4+3*x+np.random.normal(size=n)
#scaling the x and y vectors
x = x - np.mean(x)
y = y - np.mean(y)
variance_x = np.sum(x@x)/n
variance_y = np.sum(y@y)/n
print(variance_x)
print(variance_y)
cov_xy = np.sum(x@y)/n
cov_xx = np.sum(x@x)/n
cov_yy = np.sum(y@y)/n
C = np.zeros((2,2))
C[0,0]= cov_xx/variance_x
C[1,1]= cov_yy/variance_y
C[0,1]= cov_xy/np.sqrt(variance_y*variance_x)
C[1,0]= C[0,1]
print(C)
# We see that the matrix elements along the diagonal are one as they
# should be and that the matrix is symmetric. Furthermore, diagonalizing
# this matrix we easily see that it is a positive definite matrix.
#
# The above procedure with **numpy** can be made more compact if we use **pandas**.
# ## Using Pandas
#
# We whow here how we can set up the correlation matrix using **pandas**, as done in this simple code
import numpy as np
import pandas as pd
n = 10
x = np.random.normal(size=n)
x = x - np.mean(x)
y = 4+3*x+np.random.normal(size=n)
y = y - np.mean(y)
X = (np.vstack((x, y))).T
print(X)
Xpd = pd.DataFrame(X)
print(Xpd)
correlation_matrix = Xpd.corr()
print(correlation_matrix)
# ## And then the Franke Function
#
# We expand this model to the Franke function discussed above.
# +
# Common imports
import numpy as np
import pandas as pd
def FrankeFunction(x,y):
term1 = 0.75*np.exp(-(0.25*(9*x-2)**2) - 0.25*((9*y-2)**2))
term2 = 0.75*np.exp(-((9*x+1)**2)/49.0 - 0.1*(9*y+1))
term3 = 0.5*np.exp(-(9*x-7)**2/4.0 - 0.25*((9*y-3)**2))
term4 = -0.2*np.exp(-(9*x-4)**2 - (9*y-7)**2)
return term1 + term2 + term3 + term4
def create_X(x, y, n ):
if len(x.shape) > 1:
x = np.ravel(x)
y = np.ravel(y)
N = len(x)
l = int((n+1)*(n+2)/2) # Number of elements in beta
X = np.ones((N,l))
for i in range(1,n+1):
q = int((i)*(i+1)/2)
for k in range(i+1):
X[:,q+k] = (x**(i-k))*(y**k)
return X
# Making meshgrid of datapoints and compute Franke's function
n = 4
N = 100
x = np.sort(np.random.uniform(0, 1, N))
y = np.sort(np.random.uniform(0, 1, N))
z = FrankeFunction(x, y)
X = create_X(x, y, n=n)
Xpd = pd.DataFrame(X)
# subtract the mean values and set up the covariance matrix
Xpd = Xpd - Xpd.mean()
covariance_matrix = Xpd.cov()
print(covariance_matrix)
# -
# We note here that the covariance is zero for the first rows and
# columns since all matrix elements in the design matrix were set to one
# (we are fitting the function in terms of a polynomial of degree $n$). We would however not include the intercept
# and wee can simply
# drop these elements and construct a correlation
# matrix without them by centering our matrix elements by subtracting the mean of each column.
# ## Lnks with the Design Matrix
#
# We can rewrite the covariance matrix in a more compact form in terms of the design/feature matrix $\boldsymbol{X}$ as
# $$
# \boldsymbol{C}[\boldsymbol{x}] = \frac{1}{n}\boldsymbol{X}^T\boldsymbol{X}= \mathbb{E}[\boldsymbol{X}^T\boldsymbol{X}].
# $$
# To see this let us simply look at a design matrix $\boldsymbol{X}\in {\mathbb{R}}^{2\times 2}$
# $$
# \boldsymbol{X}=\begin{bmatrix}
# x_{00} & x_{01}\\
# x_{10} & x_{11}\\
# \end{bmatrix}=\begin{bmatrix}
# \boldsymbol{x}_{0} & \boldsymbol{x}_{1}\\
# \end{bmatrix}.
# $$
# ## Computing the Expectation Values
#
# If we then compute the expectation value
# $$
# \mathbb{E}[\boldsymbol{X}^T\boldsymbol{X}] = \frac{1}{n}\boldsymbol{X}^T\boldsymbol{X}=\begin{bmatrix}
# x_{00}^2+x_{01}^2 & x_{00}x_{10}+x_{01}x_{11}\\
# x_{10}x_{00}+x_{11}x_{01} & x_{10}^2+x_{11}^2\\
# \end{bmatrix},
# $$
# which is just
# $$
# \boldsymbol{C}[\boldsymbol{x}_0,\boldsymbol{x}_1] = \boldsymbol{C}[\boldsymbol{x}]=\begin{bmatrix} \mathrm{var}[\boldsymbol{x}_0] & \mathrm{cov}[\boldsymbol{x}_0,\boldsymbol{x}_1] \\
# \mathrm{cov}[\boldsymbol{x}_1,\boldsymbol{x}_0] & \mathrm{var}[\boldsymbol{x}_1] \\
# \end{bmatrix},
# $$
# where we wrote $$\boldsymbol{C}[\boldsymbol{x}_0,\boldsymbol{x}_1] = \boldsymbol{C}[\boldsymbol{x}]$$ to indicate that this the covariance of the vectors $\boldsymbol{x}$ of the design/feature matrix $\boldsymbol{X}$.
#
# It is easy to generalize this to a matrix $\boldsymbol{X}\in {\mathbb{R}}^{n\times p}$.
# ## Towards the PCA theorem
#
# We have that the covariance matrix (the correlation matrix involves a simple rescaling) is given as
# $$
# \boldsymbol{C}[\boldsymbol{x}] = \frac{1}{n}\boldsymbol{X}^T\boldsymbol{X}= \mathbb{E}[\boldsymbol{X}^T\boldsymbol{X}].
# $$
# Let us now assume that we can perform a series of orthogonal transformations where we employ some orthogonal matrices $\boldsymbol{S}$.
# These matrices are defined as $\boldsymbol{S}\in {\mathbb{R}}^{p\times p}$ and obey the orthogonality requirements $\boldsymbol{S}\boldsymbol{S}^T=\boldsymbol{S}^T\boldsymbol{S}=\boldsymbol{I}$. The matrix can be written out in terms of the column vectors $\boldsymbol{s}_i$ as $\boldsymbol{S}=[\boldsymbol{s}_0,\boldsymbol{s}_1,\dots,\boldsymbol{s}_{p-1}]$ and $\boldsymbol{s}_i \in {\mathbb{R}}^{p}$.
#
# Assume also that there is a transformation $\boldsymbol{S}^T\boldsymbol{C}[\boldsymbol{x}]\boldsymbol{S}=\boldsymbol{C}[\boldsymbol{y}]$ such that the new matrix $\boldsymbol{C}[\boldsymbol{y}]$ is diagonal with elements $[\lambda_0,\lambda_1,\lambda_2,\dots,\lambda_{p-1}]$.
#
# That is we have
# $$
# \boldsymbol{C}[\boldsymbol{y}] = \mathbb{E}[\boldsymbol{S}^T\boldsymbol{X}^T\boldsymbol{X}T\boldsymbol{S}]=\boldsymbol{S}^T\boldsymbol{C}[\boldsymbol{x}]\boldsymbol{S},
# $$
# since the matrix $\boldsymbol{S}$ is not a data dependent matrix. Multiplying with $\boldsymbol{S}$ from the left we have
# $$
# \boldsymbol{S}\boldsymbol{C}[\boldsymbol{y}] = \boldsymbol{C}[\boldsymbol{x}]\boldsymbol{S},
# $$
# and since $\boldsymbol{C}[\boldsymbol{y}]$ is diagonal we have for a given eigenvalue $i$ of the covariance matrix that
# $$
# \boldsymbol{S}_i\lambda_i = \boldsymbol{C}[\boldsymbol{x}]\boldsymbol{S}_i.
# $$
# ## More on the PCA Theorem
#
# In the derivation of the PCA theorem we will assume that the eigenvalues are ordered in descending order, that is
# $\lambda_0 > \lambda_1 > \dots > \lambda_{p-1}$.
#
# The eigenvalues tell us then how much we need to stretch the
# corresponding eigenvectors. Dimensions with large eigenvalues have
# thus large variations (large variance) and define therefore useful
# dimensions. The data points are more spread out in the direction of
# these eigenvectors. Smaller eigenvalues mean on the other hand that
# the corresponding eigenvectors are shrunk accordingly and the data
# points are tightly bunched together and there is not much variation in
# these specific directions. Hopefully then we could leave it out
# dimensions where the eigenvalues are very small. If $p$ is very large,
# we could then aim at reducing $p$ to $l << p$ and handle only $l$
# features/predictors.
# ## The Algorithm before theorem
#
# Here's how we would proceed in setting up the algorithm for the PCA, see also discussion below here.
# * Set up the datapoints for the design/feature matrix $\boldsymbol{X}$ with $\boldsymbol{X}\in {\mathbb{R}}^{n\times p}$, with the predictors/features $p$ referring to the column numbers and the entries $n$ being the row elements.
# $$
# \boldsymbol{X}=\begin{bmatrix}
# x_{0,0} & x_{0,1} & x_{0,2}& \dots & \dots x_{0,p-1}\\
# x_{1,0} & x_{1,1} & x_{1,2}& \dots & \dots x_{1,p-1}\\
# x_{2,0} & x_{2,1} & x_{2,2}& \dots & \dots x_{2,p-1}\\
# \dots & \dots & \dots & \dots \dots & \dots \\
# x_{n-2,0} & x_{n-2,1} & x_{n-2,2}& \dots & \dots x_{n-2,p-1}\\
# x_{n-1,0} & x_{n-1,1} & x_{n-1,2}& \dots & \dots x_{n-1,p-1}\\
# \end{bmatrix},
# $$
# * Center the data by subtracting the mean value for each column. This leads to a new matrix $\boldsymbol{X}\rightarrow \overline{\boldsymbol{X}}$.
#
# * Compute then the covariance/correlation matrix $\mathbb{E}[\overline{\boldsymbol{X}}^T\overline{\boldsymbol{X}}]$.
#
# * Find the eigenpairs of $\boldsymbol{C}$ with eigenvalues $[\lambda_0,\lambda_1,\dots,\lambda_{p-1}]$ and eigenvectors $[\boldsymbol{s}_0,\boldsymbol{s}_1,\dots,\boldsymbol{s}_{p-1}]$.
#
# * Order the eigenvalue (and the eigenvectors accordingly) in order of decreasing eigenvalues.
#
# * Keep only those $l$ eigenvalues larger than a selected threshold value, discarding thus $p-l$ features since we expect small variations in the data here.
# ## Writing our own PCA code
#
# We will use a simple example first with two-dimensional data
# drawn from a multivariate normal distribution with the following mean and covariance matrix (we have fixed these quantities but will play around with them below):
# $$
# \mu = (-1,2) \qquad \Sigma = \begin{bmatrix} 4 & 2 \\
# 2 & 2
# \end{bmatrix}
# $$
# Note that the mean refers to each column of data.
# We will generate $n = 10000$ points $X = \{ x_1, \ldots, x_N \}$ from
# this distribution, and store them in the $1000 \times 2$ matrix $\boldsymbol{X}$. This is our design matrix where we have forced the covariance and mean values to take specific values.
# ## Implementing it
# The following Python code aids in setting up the data and writing out the design matrix.
# Note that the function **multivariate** returns also the covariance discussed above and that it is defined by dividing by $n-1$ instead of $n$.
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from IPython.display import display
n = 10000
mean = (-1, 2)
cov = [[10, 0.02], [0.02, 0.05]]
X = np.random.multivariate_normal(mean, cov, n)
# Now we are going to implement the PCA algorithm. We will break it down into various substeps.
# ## First Step
#
# The first step of PCA is to compute the sample mean of the data and use it to center the data. Recall that the sample mean is
# $$
# \mu_n = \frac{1}{n} \sum_{i=1}^n x_i
# $$
# and the mean-centered data $\bar{X} = \{ \bar{x}_1, \ldots, \bar{x}_n \}$ takes the form
# $$
# \bar{x}_i = x_i - \mu_n.
# $$
# When you are done with these steps, print out $\mu_n$ to verify it is
# close to $\mu$ and plot your mean centered data to verify it is
# centered at the origin!
# The following code elements perform these operations using **pandas** or using our own functionality for doing so. The latter, using **numpy** is rather simple through the **mean()** function.
df = pd.DataFrame(X)
# Pandas does the centering for us
df = df -df.mean()
# we center it ourselves
X_centered = X - X.mean(axis=0)
# ## Scaling
# Alternatively, we could use the functions we discussed
# earlier for scaling the data set. That is, we could have used the
# **StandardScaler** function in **Scikit-Learn**, a function which ensures
# that for each feature/predictor we study the mean value is zero and
# the variance is one (every column in the design/feature matrix). You
# would then not get the same results, since we divide by the
# variance. The diagonal covariance matrix elements will then be one,
# while the non-diagonal ones need to be divided by $2\sqrt{2}$ for our
# specific case.
# ## Centered Data
#
# Now we are going to use the mean centered data to compute the sample covariance of the data by using the following equation
# $$
# \Sigma_n = \frac{1}{n-1} \sum_{i=1}^n \bar{x}_i^T \bar{x}_i = \frac{1}{n-1} \sum_{i=1}^n (x_i - \mu_n)^T (x_i - \mu_n)
# $$
# where the data points $x_i \in \mathbb{R}^p$ (here in this example $p = 2$) are column vectors and $x^T$ is the transpose of $x$.
# We can write our own code or simply use either the functionaly of **numpy** or that of **pandas**, as follows
print(df.cov())
print(np.cov(X_centered.T))
# Note that the way we define the covariance matrix here has a factor $n-1$ instead of $n$. This is included in the **cov()** function by **numpy** and **pandas**.
# Our own code here is not very elegant and asks for obvious improvements. It is tailored to this specific $2\times 2$ covariance matrix.
# extract the relevant columns from the centered design matrix of dim n x 2
x = X_centered[:,0]
y = X_centered[:,1]
Cov = np.zeros((2,2))
Cov[0,1] = np.sum(x.T@y)/(n-1.0)
Cov[0,0] = np.sum(x.T@x)/(n-1.0)
Cov[1,1] = np.sum(y.T@y)/(n-1.0)
Cov[1,0]= Cov[0,1]
print("Centered covariance using own code")
print(Cov)
plt.plot(x, y, 'x')
plt.axis('equal')
plt.show()
# ## Exploring
#
# Depending on the number of points $n$, we will get results that are close to the covariance values defined above.
# The plot shows how the data are clustered around a line with slope close to one. Is this expected? Try to change the covariance and the mean values. For example, try to make the variance of the first element much larger than that of the second diagonal element. Try also to shrink the covariance (the non-diagonal elements) and see how the data points are distributed.
# ## Diagonalize the sample covariance matrix to obtain the principal components
#
# Now we are ready to solve for the principal components! To do so we
# diagonalize the sample covariance matrix $\Sigma$. We can use the
# function **np.linalg.eig** to do so. It will return the eigenvalues and
# eigenvectors of $\Sigma$. Once we have these we can perform the
# following tasks:
#
# * We compute the percentage of the total variance captured by the first principal component
#
# * We plot the mean centered data and lines along the first and second principal components
#
# * Then we project the mean centered data onto the first and second principal components, and plot the projected data.
#
# * Finally, we approximate the data as
# $$
# x_i \approx \tilde{x}_i = \mu_n + \langle x_i, v_0 \rangle v_0
# $$
# where $v_0$ is the first principal component.
# ## Collecting all Steps
#
# Collecting all these steps we can write our own PCA function and
# compare this with the functionality included in **Scikit-Learn**.
#
# The code here outlines some of the elements we could include in the
# analysis. Feel free to extend upon this in order to address the above
# questions.
# diagonalize and obtain eigenvalues, not necessarily sorted
EigValues, EigVectors = np.linalg.eig(Cov)
# sort eigenvectors and eigenvalues
#permute = EigValues.argsort()
#EigValues = EigValues[permute]
#EigVectors = EigVectors[:,permute]
print("Eigenvalues of Covariance matrix")
for i in range(2):
print(EigValues[i])
FirstEigvector = EigVectors[:,0]
SecondEigvector = EigVectors[:,1]
print("First eigenvector")
print(FirstEigvector)
print("Second eigenvector")
print(SecondEigvector)
#thereafter we do a PCA with Scikit-learn
from sklearn.decomposition import PCA
pca = PCA(n_components = 2)
X2Dsl = pca.fit_transform(X)
print("Eigenvector of largest eigenvalue")
print(pca.components_.T[:, 0])
# This code does not contain all the above elements, but it shows how we can use **Scikit-Learn** to extract the eigenvector which corresponds to the largest eigenvalue. Try to address the questions we pose before the above code. Try also to change the values of the covariance matrix by making one of the diagonal elements much larger than the other. What do you observe then?
# ## Classical PCA Theorem
#
# We assume now that we have a design matrix $\boldsymbol{X}$ which has been
# centered as discussed above. For the sake of simplicity we skip the
# overline symbol. The matrix is defined in terms of the various column
# vectors $[\boldsymbol{x}_0,\boldsymbol{x}_1,\dots, \boldsymbol{x}_{p-1}]$ each with dimension
# $\boldsymbol{x}\in {\mathbb{R}}^{n}$.
#
# The PCA theorem states that minimizing the above reconstruction error
# corresponds to setting $\boldsymbol{W}=\boldsymbol{S}$, the orthogonal matrix which
# diagonalizes the empirical covariance(correlation) matrix. The optimal
# low-dimensional encoding of the data is then given by a set of vectors
# $\boldsymbol{z}_i$ with at most $l$ vectors, with $l << p$, defined by the
# orthogonal projection of the data onto the columns spanned by the
# eigenvectors of the covariance(correlations matrix).
# ## The PCA Theorem
#
# To show the PCA theorem let us start with the assumption that there is one vector $\boldsymbol{s}_0$ which corresponds to a solution which minimized the reconstruction error $J$. This is an orthogonal vector. It means that we now approximate the reconstruction error in terms of $\boldsymbol{w}_0$ and $\boldsymbol{z}_0$ as
#
# We are almost there, we have obtained a relation between minimizing
# the reconstruction error and the variance and the covariance
# matrix. Minimizing the error is equivalent to maximizing the variance
# of the projected data.
#
# We could trivially maximize the variance of the projection (and
# thereby minimize the error in the reconstruction function) by letting
# the norm-2 of $\boldsymbol{w}_0$ go to infinity. However, this norm since we
# want the matrix $\boldsymbol{W}$ to be an orthogonal matrix, is constrained by
# $\vert\vert \boldsymbol{w}_0 \vert\vert_2^2=1$. Imposing this condition via a
# Lagrange multiplier we can then in turn maximize
# $$
# J(\boldsymbol{w}_0)= \boldsymbol{w}_0^T\boldsymbol{C}[\boldsymbol{x}]\boldsymbol{w}_0+\lambda_0(1-\boldsymbol{w}_0^T\boldsymbol{w}_0).
# $$
# Taking the derivative with respect to $\boldsymbol{w}_0$ we obtain
# $$
# \frac{\partial J(\boldsymbol{w}_0)}{\partial \boldsymbol{w}_0}= 2\boldsymbol{C}[\boldsymbol{x}]\boldsymbol{w}_0-2\lambda_0\boldsymbol{w}_0=0,
# $$
# meaning that
# $$
# \boldsymbol{C}[\boldsymbol{x}]\boldsymbol{w}_0=\lambda_0\boldsymbol{w}_0.
# $$
# **The direction that maximizes the variance (or minimizes the construction error) is an eigenvector of the covariance matrix**! If we left multiply with $\boldsymbol{w}_0^T$ we have the variance of the projected data is
# $$
# \boldsymbol{w}_0^T\boldsymbol{C}[\boldsymbol{x}]\boldsymbol{w}_0=\lambda_0.
# $$
# If we want to maximize the variance (minimize the construction error)
# we simply pick the eigenvector of the covariance matrix with the
# largest eigenvalue. This establishes the link between the minimization
# of the reconstruction function $J$ in terms of an orthogonal matrix
# and the maximization of the variance and thereby the covariance of our
# observations encoded in the design/feature matrix $\boldsymbol{X}$.
#
# The proof
# for the other eigenvectors $\boldsymbol{w}_1,\boldsymbol{w}_2,\dots$ can be
# established by applying the above arguments and using the fact that
# our basis of eigenvectors is orthogonal, see [Murphy chapter
# 12.2](https://mitpress.mit.edu/books/machine-learning-1). The
# discussion in chapter 12.2 of Murphy's text has also a nice link with
# the Singular Value Decomposition theorem. For categorical data, see
# chapter 12.4 and discussion therein.
#
# For more details, see for example [Vidal, Ma and Sastry, chapter 2](https://www.springer.com/gp/book/9780387878102).
# ## Geometric Interpretation and link with Singular Value Decomposition
#
# For a detailed demonstration of the geometric interpretation, see [Vidal, Ma and Sastry, section 2.1.2](https://www.springer.com/gp/book/9780387878102).
#
# Principal Component Analysis (PCA) is by far the most popular dimensionality reduction algorithm.
# First it identifies the hyperplane that lies closest to the data, and then it projects the data onto it.
#
# The following Python code uses NumPy’s **svd()** function to obtain all the principal components of the
# training set, then extracts the first two principal components. First we center the data using either **pandas** or our own code
# +
import numpy as np
import pandas as pd
from IPython.display import display
np.random.seed(100)
# setting up a 10 x 5 vanilla matrix
rows = 10
cols = 5
X = np.random.randn(rows,cols)
df = pd.DataFrame(X)
# Pandas does the centering for us
df = df -df.mean()
display(df)
# we center it ourselves
X_centered = X - X.mean(axis=0)
# Then check the difference between pandas and our own set up
print(X_centered-df)
#Now we do an SVD
U, s, V = np.linalg.svd(X_centered)
c1 = V.T[:, 0]
c2 = V.T[:, 1]
W2 = V.T[:, :2]
X2D = X_centered.dot(W2)
print(X2D)
# -
# PCA assumes that the dataset is centered around the origin. Scikit-Learn’s PCA classes take care of centering
# the data for you. However, if you implement PCA yourself (as in the preceding example), or if you use other libraries, don’t
# forget to center the data first.
#
# Once you have identified all the principal components, you can reduce the dimensionality of the dataset
# down to $d$ dimensions by projecting it onto the hyperplane defined by the first $d$ principal components.
# Selecting this hyperplane ensures that the projection will preserve as much variance as possible.
W2 = V.T[:, :2]
X2D = X_centered.dot(W2)
# ## PCA and scikit-learn
#
# Scikit-Learn’s PCA class implements PCA using SVD decomposition just like we did before. The
# following code applies PCA to reduce the dimensionality of the dataset down to two dimensions (note
# that it automatically takes care of centering the data):
#thereafter we do a PCA with Scikit-learn
from sklearn.decomposition import PCA
pca = PCA(n_components = 2)
X2D = pca.fit_transform(X)
print(X2D)
# After fitting the PCA transformer to the dataset, you can access the principal components using the
# components variable (note that it contains the PCs as horizontal vectors, so, for example, the first
# principal component is equal to
pca.components_.T[:, 0]
# Another very useful piece of information is the explained variance ratio of each principal component,
# available via the $explained\_variance\_ratio$ variable. It indicates the proportion of the dataset’s
# variance that lies along the axis of each principal component.
# ## Back to the Cancer Data
# We can now repeat the above but applied to real data, in this case our breast cancer data.
# Here we compute performance scores on the training data using logistic regression.
# +
import matplotlib.pyplot as plt
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.datasets import load_breast_cancer
from sklearn.linear_model import LogisticRegression
cancer = load_breast_cancer()
X_train, X_test, y_train, y_test = train_test_split(cancer.data,cancer.target,random_state=0)
logreg = LogisticRegression()
logreg.fit(X_train, y_train)
print("Train set accuracy from Logistic Regression: {:.2f}".format(logreg.score(X_train,y_train)))
# We scale the data
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(X_train)
X_train_scaled = scaler.transform(X_train)
X_test_scaled = scaler.transform(X_test)
# Then perform again a log reg fit
logreg.fit(X_train_scaled, y_train)
print("Train set accuracy scaled data: {:.2f}".format(logreg.score(X_train_scaled,y_train)))
#thereafter we do a PCA with Scikit-learn
from sklearn.decomposition import PCA
pca = PCA(n_components = 2)
X2D_train = pca.fit_transform(X_train_scaled)
# and finally compute the log reg fit and the score on the training data
logreg.fit(X2D_train,y_train)
print("Train set accuracy scaled and PCA data: {:.2f}".format(logreg.score(X2D_train,y_train)))
# -
# We see that our training data after the PCA decomposition has a performance similar to the non-scaled data.
#
# Instead of arbitrarily choosing the number of dimensions to reduce down to, it is generally preferable to
# choose the number of dimensions that add up to a sufficiently large portion of the variance (e.g., 95%).
# Unless, of course, you are reducing dimensionality for data visualization — in that case you will
# generally want to reduce the dimensionality down to 2 or 3.
# The following code computes PCA without reducing dimensionality, then computes the minimum number
# of dimensions required to preserve 95% of the training set’s variance:
pca = PCA()
pca.fit(X)
cumsum = np.cumsum(pca.explained_variance_ratio_)
d = np.argmax(cumsum >= 0.95) + 1
# You could then set $n\_components=d$ and run PCA again. However, there is a much better option: instead
# of specifying the number of principal components you want to preserve, you can set $n\_components$ to be
# a float between 0.0 and 1.0, indicating the ratio of variance you wish to preserve:
pca = PCA(n_components=0.95)
X_reduced = pca.fit_transform(X)
# ## Incremental PCA
#
# One problem with the preceding implementation of PCA is that it requires the whole training set to fit in
# memory in order for the SVD algorithm to run. Fortunately, Incremental PCA (IPCA) algorithms have
# been developed: you can split the training set into mini-batches and feed an IPCA algorithm one minibatch
# at a time. This is useful for large training sets, and also to apply PCA online (i.e., on the fly, as new
# instances arrive).
# ### Randomized PCA
#
# Scikit-Learn offers yet another option to perform PCA, called Randomized PCA. This is a stochastic
# algorithm that quickly finds an approximation of the first d principal components. Its computational
# complexity is $O(m \times d^2)+O(d^3)$, instead of $O(m \times n^2) + O(n^3)$, so it is dramatically faster than the
# previous algorithms when $d$ is much smaller than $n$.
# ### Kernel PCA
#
# The kernel trick is a mathematical technique that implicitly maps instances into a
# very high-dimensional space (called the feature space), enabling nonlinear classification and regression
# with Support Vector Machines. Recall that a linear decision boundary in the high-dimensional feature
# space corresponds to a complex nonlinear decision boundary in the original space.
# It turns out that the same trick can be applied to PCA, making it possible to perform complex nonlinear
# projections for dimensionality reduction. This is called Kernel PCA (kPCA). It is often good at
# preserving clusters of instances after projection, or sometimes even unrolling datasets that lie close to a
# twisted manifold.
# For example, the following code uses Scikit-Learn’s KernelPCA class to perform kPCA with an
from sklearn.decomposition import KernelPCA
rbf_pca = KernelPCA(n_components = 2, kernel="rbf", gamma=0.04)
X_reduced = rbf_pca.fit_transform(X)
# ## Other techniques
#
# There are many other dimensionality reduction techniques, several of which are available in Scikit-Learn.
#
# Here are some of the most popular:
# * **Multidimensional Scaling (MDS)** reduces dimensionality while trying to preserve the distances between the instances.
#
# * **Isomap** creates a graph by connecting each instance to its nearest neighbors, then reduces dimensionality while trying to preserve the geodesic distances between the instances.
#
# * **t-Distributed Stochastic Neighbor Embedding** (t-SNE) reduces dimensionality while trying to keep similar instances close and dissimilar instances apart. It is mostly used for visualization, in particular to visualize clusters of instances in high-dimensional space (e.g., to visualize the MNIST images in 2D).
#
# * Linear Discriminant Analysis (LDA) is actually a classification algorithm, but during training it learns the most discriminative axes between the classes, and these axes can then be used to define a hyperplane onto which to project the data. The benefit is that the projection will keep classes as far apart as possible, so LDA is a good technique to reduce dimensionality before running another classification algorithm such as a Support Vector Machine (SVM) classifier discussed in the SVM lectures.
| Lectures/week43.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Gressling/examples/blob/master/25_1_SQL_with_a_relational_database.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="TLGCzrNPNLJ4"
# SQL with a relational database
# author: <NAME>
# license: MIT License # code: github.com/gressling/examples
# activity: single example # index: 25-1
# + id="ooTiqrigNQG7"
import psycopg2
conn = psycopg2.connect("dbname='example_db' user='dbuser'
host='localhost' password='password'")
cur = conn.cursor()
# + id="IyUJDE2tNTrb"
cur.execute("""SELECT * from results where yield>80""")
rows = cur.fetchall()
# + id="qiwHaynnNVyz"
for row in rows:
print(" ", row[0])
| 25_1_SQL_with_a_relational_database.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sqlalchemy import create_engine
import spotipy
from spotipy.oauth2 import SpotifyClientCredentials
from env_vars import *
import pandas as pd
import sqlite3
from sqlalchemy import engine
import pickle
import numpy as np
# +
def authorize():
client_credentials_manager = SpotifyClientCredentials(client_id=CLIENT_ID,client_secret=CLIENT_SECRET)
sp = spotipy.Spotify(client_credentials_manager=client_credentials_manager)
client_credentials_manager = SpotifyClientCredentials(client_id=CLIENT_ID,client_secret=CLIENT_SECRET)
sp = spotipy.Spotify(client_credentials_manager=client_credentials_manager)
def get_features(trackID):
return sp.audio_features(trackID)[0]
def get_popularity(trackID):
return sp.track(trackID)['popularity']
def get_markets(trackID):
foo = sp.track(trackID)['available_markets']
if foo != None:
return 1
else:
return 0
# -
con = sqlite3.connect("song_list_v3_raw.db")
# Load the data into a DataFrame
df = pd.read_sql_query("SELECT * from songs", con)
con.close()
X = df.loc[df['index'][202912:274019]]
X = X.drop('level_0', axis=1)
engine = create_engine('sqlite:///song_list_v3_11.db', echo=False)
X.to_sql('songs', con=engine)
engine.execute('SELECT * FROM songs;').fetchone()
for row_index in X['songid']:
songid = row_index
#print(songid)
popularity = get_popularity(songid)
#print(popularity)
markets = get_markets(songid)
#print(markets)
try:
engine.execute(f'UPDATE songs SET popularity = "{popularity}", availability = "{markets}" WHERE songid = \"{songid}\"')
authorize()
except:
print(f"couldn't insert {markets} or {popularity} into table")
continue
| data_collection/split_workload_try.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas_datareader.data as reader
import matplotlib.pyplot as plt
import datetime as dt
import seaborn as sns
end = dt.datetime.now()
start = dt.date(end.year - 5,end.month,end.day)
portfolio = ['BMW.DE', 'DAI.DE', 'VOW3.DE', '^GDAXI', 'TSLA']
df = reader.get_data_yahoo(portfolio,start,end)['Adj Close']
df
returns = df.pct_change()
returns.cov()
returns.var()
returns.corr()
sns.heatmap(returns.corr())
| Covariance and Correlation Matrix of German carstock returns with Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import scipy
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
import statsmodels
import statsmodels.api as sm
from statsmodels.formula.api import ols
# -
# # one-way anova
# ### Example 1
#
# Assume partner_status has no effect on conformity.
# Do a one-way anova.
# If PR(>F) <= 0.05, reject the assumption.
# +
moore = sm.datasets.get_rdataset("Moore", "car", cache=True)
data = moore.data
data = data.rename(columns={"partner.status" :"partner_status"})
data.head()
# +
# r = 'conformity ~ C(fcategory, Sum)*C(partner_status, Sum)'
r = 'conformity ~ C(partner_status)'
moore_lm = ols(r, data=data).fit()
moore_lm.params
# -
moore_lm.bse
fig, ax = plt.subplots()
fig = sm.graphics.plot_fit(moore_lm, 1, ax=ax)
# +
table = sm.stats.anova_lm(moore_lm, typ=2) # Type 2 ANOVA DataFrame
print(table)
# -
# ### Example 2
#
# Assume gender has no effect on result.
# Do a one-way anova.
# If PR(>F) <= 0.05, reject the assumption.
#
# +
df = pd.DataFrame(
{'result': [0,0,0,0,0,1, 1,1,1,1,1,0],
'gender': ['M','M','M','M','M','M', 'F','F','F','F','F','F'],
'age': ['<50','<50','<50','>=50','>=50','>=50','<50','<50','<50','>=50','>=50','>=50']}
)
# -
r = 'result ~ C(gender)'
results = ols(r, data=df).fit()
results.params
fig, ax = plt.subplots()
fig = sm.graphics.plot_fit(result_lm, 1, ax=ax)
# +
table = sm.stats.anova_lm(results, typ=2) # Type 2 ANOVA DataFrame
print(table)
# +
def esq_sm(aov_table):
return aov_table['sum_sq'][0]/(aov_table['sum_sq'][0]+aov_table['sum_sq'][1])
esq_sm(table)
# -
# # two-way anova
# ### Example 1
data = pd.read_csv('https://raw.githubusercontent.com/plotly/datasets/master/tooth_growth_csv')
data[0:10]
formula = 'len ~ C(supp) + C(dose) + C(supp):C(dose)'
model = ols(formula, data).fit()
aov_table = sm.stats.anova_lm(model, typ=2)
print(aov_table)
model.summary()
model.params
# +
# Q-Q plot
res = model.resid
fig = sm.qqplot(res, line='s')
# -
# ### Example 2
# +
df = pd.DataFrame(
{'result': [0,0,0,0,0,1, 1,1,1,1,1,0],
'gender': ['M','M','M','M','M','M', 'F','F','F','F','F','F'],
'age': ['<50','<50','<50','<50','<50','>=50','>=50','>=50','>=50','>=50','>=50','>=50'],
'location': np.random.choice(['Tokyo', 'Shinjuku'], 12)}
)
df
# -
formula = """
result ~
C(gender) + C(age) + C(location) + C(gender):C(age) + C(gender):C(location) + C(location):C(age)
"""
model = ols(formula, df).fit()
aov_table = sm.stats.anova_lm(model, typ=2)
print(aov_table)
model.summary()
| anova.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 项目提交
#
# 准备好提交项目时,意味着你已经根据 [审阅标准](https://review.udacity.com/#!/rubrics/1427/view) 自行做了检查,确保已完成所有任务并回答了所有问题。 然后,就可以压缩文件并提交你的项目了!
#
# 请参考以下步骤:
# 1. 所有代码单元格都已在notebook 2和notebook 3中*运行*过(并且已保存进度)。
# 2. 这些notebook中的所有问题都已得到解答。
# 3. `model.py`中的架构是经过测试选出的最佳架构。
#
# 在进行下一步之前,请确保你的所有代码都已保存,同时,无需更改这些单元格中的任何代码,因为此代码仅用于帮助你提交项目。
#
# ---
#
# 我们要做的第一件事就是,将你的notebook转换成`.html`文件。这些文件将保存每个单元格的输出以及已修改并保存在这些notebook中的所有代码与文本。 请注意,不包括第一个notebook(译者注:原文的复数notebooks是否有问题),因为其中的内容不会影响你的项目审阅。
# !jupyter nbconvert "2_Training.ipynb"
# !jupyter nbconvert "3_Inference.ipynb"
# ### 压缩项目文件
#
# 接下来,我们将所有notebook文件和`model.py`文件压缩到一个名为`project2.zip`的压缩存档中。
#
# 完成这个步骤后,你应该会在home目录中看到此压缩文件。你可以从下面的图像中下载该文件,方法是从列表中选择它并单击**“下载”**。此步骤可能需要一两分钟才能完成。
#
# <img src='images/download_ex.png' width=50% height=50%/>
!!apt-get -y update && apt-get install -y zip
# !zip project2.zip -r . -i@filelist.txt
# ### 提交项目
#
# 创建并下载压缩文件后,单击`Submit`按钮并按照提交`project2.zip`文件的说明进行操作即可。恭喜你完成这个项目哦,希望你学有所得,学有所乐!
| 4_Zip Your Project Files and Submit-zh.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# <NAME>
# separate into functions and have more configurability
import matplotlib.pyplot as plt
import numpy as np
import numpy.linalg as lin
import matplotlib.patches as mpatches
import scipy as sypy
from scipy import signal
from scipy import io
from numpy import ndarray
# For testing function.
# maxtrain=6; #maximum training images
# iter = 1; #maximum iterations
# eta=0.01; # learning rate
# n_fl=10;
# # # %%select the pooling
# # pool='maxpool';
# pool= 'avgpool';
# trained_parameter_file = 'trained_parameters'+'_maxtrain'+str(maxtrain)+'_iter'+str(iter)+'_eta'+str(eta)+ pool+'.mat';
# [trainlabels, trainimages, testlabels, testimages] = cnnload()
from ipynb.fs.full.cnn import cnnload
from ipynb.fs.full.avgpool import avgpool
from ipynb.fs.full.avgpool import maxpool
def cnn_inference(testlabels,testimages,pool,trained_parameter_file):
fn = 4; # number of kernels for layer 1
ks = 5; # size of kernel
[n,h,w]=np.shape(testimages)
numtest=n;
h1 = h-ks+1;
w1 = w-ks+1;
A1 = np.zeros((fn,h1,w1));
h2 = h1//2;
w2 = w1//2;
I2 = np.zeros((fn,h2,w2));
A2 = np.zeros((fn,h2,w2));
A3 = np.zeros(10);
tr_pr_fl=sypy.io.loadmat(trained_parameter_file)
W1=tr_pr_fl['W1']
W3=tr_pr_fl['W3']
B1=tr_pr_fl['B1']
B2=tr_pr_fl['B2']
B3=tr_pr_fl['B3']
S2=tr_pr_fl['S2']
maxtrain=tr_pr_fl['maxtrain']
it= tr_pr_fl['it']
eta= tr_pr_fl['eta']
err= tr_pr_fl['err']
# normalize data to [-1,1] range
nitest = (testimages / 255) * 2 - 1;
miss = 0;
missimages = np.zeros(numtest);
misslabels = np.zeros(numtest);
for im in range(0,numtest):
for fm in range (0,fn):
A1[fm,:,:] = sypy.signal.convolve2d(nitest[im,:,:],W1[fm, ::-1, ::-1], 'valid') + B1[:,fm]
Z1 = np.tanh(A1);
# % Layer 2: max or average (both subsample) with scaling and bias
for fm in range(0,fn):
if(pool=='maxpool'):
I2[fm,:,:] = maxpool(Z1[fm,:,:]);
elif(pool=='avgpool'):
I2[fm,:,:] = avgpool(Z1[fm,:,:]);
A2[fm,:,:] = I2[fm,:,:] * S2[:,fm] + B2[:,fm];
Z2 = np.tanh(A2);
# % Layer 3: fully connected
for cl in range(0,10):
A3[cl] = sypy.signal.convolve(Z2,W3[cl, ::-1, ::-1, ::-1],'valid') + B3[:,cl]
Z3 = np.tanh(A3); # Final output
pm = np.max(Z3);
pl= np.argmax(Z3);
if (pl != testlabels[im]+1):
miss = miss + 1;
missimages[miss] = im;
misslabels[miss] = pl - 1;
print(['Miss: ' + str(miss) +' out of ' +str(numtest)]);
return missimages, misslabels
# -
| ML_Notebook/hdr_om/cnn_inference.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from collections import Counter
DEBUG = True
# -
# ## Multiclass Classifier with Decision Tree using ID3 algorithm
# ### Utility functions
# Min-Max normalization function
# + code_folding=[]
def normalize(matrix):
"""
Min Max scaling
matrix must be a np.ndarry
"""
return (matrix - matrix.min(axis=0))/(matrix.max(axis=0) - matrix.min(axis=0))
# -
# Function to calculate Entropy
# + code_folding=[]
def entropy(y) -> float:
m = len(y)
class_freq_dict = Counter(y)
entropy = -1 * sum([(class_count/m)*np.log2(class_count/m)
for class_count in class_freq_dict.values()])
return entropy
# -
# Function to load csv file and return X features and y target matrix
# + code_folding=[]
def load_csv(file, unpack=True):
data = np.genfromtxt(file, delimiter=',', encoding=None, dtype=str)
X, y = data[:, :-1], data[:, -1].reshape(-1,1)
return (np.array(X).astype(np.float), np.array(y)) if unpack else data
# -
# Function to randomly shuffle the array using the numpy.random.shuffle()
# + code_folding=[]
def shuffle_array(matrix, seed=None):
"""
np.random.shuffle() shuffles a multidimensional arr using only the first dimension
Alternative implementation of shuffle
def shuffle_array(matrix):
# shuffles an array in place based on the first dimension
for i in range(len(matrix)):
rd_idx = np.random.randint(i,len(matrix))
matrix[i], matrix[rd_idx] = matrix[rd_idx], matrix[i]
"""
np.random.seed(seed)
np.random.shuffle(matrix)
# -
# Function to generate k fold cross validation training and testing lists
# + code_folding=[]
def generate_kfold_train_test_set(X, y, test_frac=0.10):
"""
Generates kfold train test sets from X_feat, y_target
Returns X_train_list, y_train_list, X_test_list, y_test_list
X must be np.array of ints or floats
y is the target or label array
"""
m = len(X)
norm_X = normalize(X)
test_data_count = int(test_frac*m)
start, end = 0, test_data_count
# sets to store features and labels of training and testing data
X_train_list, y_train_list, X_test_list, y_test_list = [], [], [], []
# 10-fold cross-validation:
for _ in range(10):
X_test = norm_X[start:end]
X_train = np.concatenate([norm_X[:start], norm_X[end:]], axis=0)
X_test_list.append(X_test)
X_train_list.append(X_train)
y_test = y[start:end].flatten()
y_train = np.concatenate([y[:start], y[end:]], axis=0).flatten()
y_test_list.append(y_test)
y_train_list.append(y_train)
# update test set start and end pointers
start += test_data_count
end += test_data_count
# return the feature, label fold lists for both training and testing set.
return X_train_list, y_train_list, X_test_list, y_test_list
# -
# Function to generate attr_dict
# + code_folding=[]
def get_attrib_dict_with_class_labels(X, y):
"""
Builds a dict where,
the keys are the feature idx,
the values are feature value and corresponding feature labels two-element lists
"""
attr_dict = {}
for feat_idx in range(X.shape[1]):
feat_vals = X[:, feat_idx]
attr_list = []
for data_idx, val in enumerate(feat_vals):
attr_list.append([val, y[data_idx]])
attr_dict[feat_idx] = attr_list
return attr_dict
# -
# Decision Tree Node Class
# + code_folding=[]
class Node:
def __init__(self, class_label, left, right, threshold):
self.class_label = class_label
self.left = left
self.right = right
self.threshold = threshold
def __repr__(self):
return f"{self.class_label}, {self.left}, {self.right}, {self.threshold}"
# -
# Decision Tree Class
class DecisionTree():
def __init__(self):
self.root = None
def get_major_class_label(self, class_values):
try:
return Counter(class_values).most_common(1)[0][0]
except Exception as e:
print(e)
return None
def get_best_threshold_perm_sorting(self, attri_list):
"""
Calculates best threshold value for each feature
attri_list are the values of the attr_dict
WARNING: perm sorts the training data. Shuffling is discarded
"""
attri_list.sort()
orig_entropy = entropy([label for val, label in attri_list])
max_info_gain = 0
optimal_threshold = 0
label_after_split_list = []
optimal_left_idx_list, optimal_right_idx_list = [], []
for i in range(len(attri_list)-1):
if attri_list[i][1] == attri_list[i+1][1]:
continue
# candidate threshold is the midpoint between cur data and next data
threshold = (attri_list[i][0] + attri_list[i+1][0])/2
# lists that store index, value less than threshold
lt_idx_list, lt_val_list = list(range(i+1)), [label for val, label in attri_list[:i+1]]
# lists that store index, value greater than threshold
gt_idx_list, gt_val_list = list(range(i+1,len(attri_list))), [label for val, label in attri_list[i+1:len(attri_list)]]
# calculate the entropy of the "less" list
entropy_lt_list = entropy(lt_val_list)
# calculate the entropy of the "greater" list
entropy_gt_list = entropy(gt_val_list)
# calculate the info gain using the formula
info_gain = orig_entropy - (
entropy_lt_list*(len(lt_val_list)/len(attri_list)) +
entropy_gt_list*(len(gt_val_list)/len(attri_list)))
# if current info gain > max info gan
if info_gain > max_info_gain:
max_info_gain = info_gain
optimal_threshold = threshold
optimal_left_idx_list = lt_idx_list
optimal_right_idx_list = gt_idx_list
return max_info_gain, optimal_threshold, optimal_left_idx_list, optimal_right_idx_list
def get_best_threshold(self, attri_list):
"""
Calculates best threshold value for each feature
attri_list are the values of the attr_dict
"""
# extract data vals and label vals from sorted attri_list
data_list, class_label_list = map(list, zip(*sorted(attri_list)))
orig_entropy = entropy(class_label_list)
max_info_gain = 0
optimal_threshold = 0
label_after_split_list = []
optimal_left_idx_list, optimal_right_idx_list = [], []
for i in range(len(data_list)-1):
# IMPORTANT speeding step. If the i and the i+1th class values are the same, don't split
if class_label_list[i] == class_label_list[i+1]:
continue
# candidate threshold is the midpoint between cur data and next data
threshold = (data_list[i] + data_list[i+1])/2
# lists that store index, value less than threshold
lt_idx_list, lt_val_list = [], []
# lists that store index, value greater than threshold
gt_idx_list, gt_val_list = [], []
# for each index and value in attri_list
for j, data_tuple in enumerate(attri_list):
data, label = data_tuple
# if value less or equal than the current theta:
if data < threshold:
# update the "less" list of index and value
lt_idx_list.append(j)
lt_val_list.append(label)
else:
# update the "greater" list of index and value
gt_idx_list.append(j)
gt_val_list.append(label)
# calculate the entropy of the "less" list
entropy_lt_list = entropy(lt_val_list)
# calculate the entropy of the "greater" list
entropy_gt_list = entropy(gt_val_list)
# calculate the info gain using the formula
info_gain = orig_entropy - (
entropy_lt_list*(len(lt_val_list)/len(attri_list)) +
entropy_gt_list*(len(gt_val_list)/len(attri_list)))
# if current info gain > max info gan
if info_gain > max_info_gain:
max_info_gain = info_gain
optimal_threshold = threshold
optimal_left_idx_list = lt_idx_list
optimal_right_idx_list = gt_idx_list
return max_info_gain, optimal_threshold, optimal_left_idx_list, optimal_right_idx_list
def get_best_feature(self, attr_dict):
"""
Method to select the optimum feature out of all the features.
For each feature, get the optimum threshold and information gain
"""
best_feat = None
best_info_gain = -1
best_threshold = 0
best_left_idx_list = []
best_right_idx_list = []
# get max_info_gain using each feat as split to get the best_feat to split on
for feat in attr_dict.keys():
info_gain, threshold, left_idx_list, right_idx_list = self.get_best_threshold(
attr_dict[feat])
if info_gain > best_info_gain:
best_feat = feat
best_threshold = threshold
best_info_gain = info_gain
best_left_idx_list = left_idx_list
best_right_idx_list = right_idx_list
return [best_feat, best_threshold, best_left_idx_list, best_right_idx_list]
def get_remainder_dict(self, attr_dict, index_split):
split_dict = {}
exclude_index_split = set(index_split)
for feat in attr_dict.keys():
class_label_list = []
modified_list = []
feat_val_list = attr_dict[feat]
# feat_val_list = [[1.2,'iris-setosa'],[2.2,'iris-verisicolor'],....]
for i, dl_tuple in enumerate(feat_val_list):
if i in index_split:
class_label_list.append(dl_tuple[1])
modified_list.append(dl_tuple)
split_dict[feat] = modified_list
return split_dict, class_label_list
def create_decision_tree(self, attr_dict, y_train, n_min_val):
"""
Grow decision tree and return the root node
"""
# if all the class labels are same, node is pure
if len(set(y_train)) == 1:
return Node(y_train[0], None, None, 0)
# if num class vales are less than threshold, we assign the class with max values as the class label
elif len(y_train) < n_min_val:
max_value_class = self.get_major_class_label(y_train)
return Node(max_value_class, None, None, 0)
else:
feat, threshold, left_idx_split, right_idx_split = self.get_best_feature(
attr_dict)
left_tree_dict, left_tree_class_labels = self.get_remainder_dict(
attr_dict, left_idx_split)
right_tree_dict, right_tree_class_labels = self.get_remainder_dict(
attr_dict, right_idx_split)
left_node = self.create_decision_tree(
left_tree_dict, left_tree_class_labels, n_min_val)
right_node = self.create_decision_tree(
right_tree_dict, right_tree_class_labels, n_min_val)
return Node(feat, left_node, right_node, threshold)
def fit(self, X_train, y_train, eta_min_val):
attr_dict = get_attrib_dict_with_class_labels(X_train, y_train)
self.root = self.create_decision_tree(
attr_dict, y_train, eta_min_val)
def classify(self, data_row):
cur_node = self.root
# while cur_node is not a leaf node
while cur_node and (cur_node.left or cur_node.right):
if data_row[cur_node.class_label] <= cur_node.threshold:
cur_node = cur_node.left
else:
cur_node = cur_node.right
return cur_node.class_label
def predict(self, X):
"""
Predicts labels for the X_feat data
"""
return [self.classify(data_row) for data_row in X]
# Evaluation of model
def get_accuracy(y_actual, y_pred):
return len([1 for actual,pred in zip(y_actual, y_pred) if actual==pred])/len(y_pred)
# + code_folding=[]
def run_decision_trees(X, y, n_min):
"""
function to run decision trees with k-Fold Cross validation
"""
m, n = X.shape[0], X.shape[1]
if isinstance(n_min, int):
n_min_val = n_min # Use values for node cutoff
elif 0.0 < n_min < 1.0:
n_min_val = round(n_min*m) # Use fractions for node cutoff
else:
raise ValueError("Fractions must be in range (0.0, 1.0)")
# concatenate, shuffle and split X,y
concatenated_data = np.concatenate([X,y], axis=1)
shuffle_array(concatenated_data)
X,y = concatenated_data[:,:-1].astype('float'), concatenated_data[:,-1].reshape(-1,1)
X_train_list, y_train_list, X_test_list, y_test_list = generate_kfold_train_test_set(
X, y)
decision_tree = DecisionTree()
accuracy_list = []
for i in range(10):
X_train, y_train = X_train_list[i], y_train_list[i]
X_test, y_test = X_test_list[i], y_test_list[i]
# Fit the decision tree model
decision_tree.fit(X_train, y_train, n_min_val)
y_pred = decision_tree.predict(X_test)
accuracy = get_accuracy(y_test, y_pred)
accuracy_list.append(accuracy)
print("Accuracy is ", "{:.4f}".format(accuracy))
print("Average accuracy across 10-cross validation when cutoff is",
n_min, "nodes is", "{:.4f}".format(np.mean(accuracy_list)))
print("Standard deviation across 10-cross validation when cutoff is",
n_min, "nodes is", "{:.4f}".format(np.std(accuracy_list)))
# +
# %%time
# n_min = minimum frac/num of examples for a node to become a leaf
# n_min_list_frac = [0.05,0.10,0.15,0.20,0.25]
n_min_list_val = [5, 10, 15, 20]
iris_data = "../data/iris.csv"
X,y = load_csv(iris_data)
for n_min in n_min_list_val:
run_decision_trees(X, y, n_min)
# +
# %%time
# eta_min_list_val = [5, 10, 15, 20, 25]
eta_min_list_val = [5]
iris_data = "../data/spambase.csv"
X,y = load_csv(iris_data)
for eta_min in eta_min_list_val:
run_decision_trees(X, y, eta_min)
| notebooks/decision_tree.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Import Python Modules
import numpy as np
import matplotlib.pyplot as plt
np.seterr(divide='ignore', invalid='ignore')
from ipywidgets import interact, interactive, fixed, interact_manual
import ipywidgets as widgets
# # Image resolution and Field of View example
# +
# load in data
imdata = np.genfromtxt('data/kspace_imdata.csv', delimiter=',')
# generate k-space
kdata = np.fft.fftshift(np.fft.fft2(imdata))
# Define base resolution and FOV
FOV = 200 # units are mm
Nx, Ny = imdata.shape
dx = FOV/Nx # units are mm
grid = np.meshgrid(np.arange(-(FOV/2-dx),(FOV/2)+dx,dx), np.arange(-(FOV/2-dx),(FOV/2)+dx,dx))
# Define a grid of k-space sample locations
FOVk = 2*np.pi/(dx)
dk = 2*np.pi/FOV
kgrid = np.meshgrid(np.arange(-(FOVk/2-dk),(FOVk/2)+dk,dk), np.arange(-(FOVk/2-dk),(FOVk/2)+dk,dk))
# -
# ## Use the sliders to change the k-space sampling
# Let's set FOV and resolution only in one direction
# There is an option to apply to both directions
#
# There are three interactive controls below
# 1. The first slider below can be used to reduce the image resolution (i.e. increase the voxel size)
# 2. The second slider below reduces the *encoded* FOV by the factor shown, only integers allowed
# 3. The sliders apply only to one dimension in k-space. Tick the checkbox below to apply to both
@interact(reduce_resolution = (0.1,1,0.1),
reduce_FOV = (1,4,1),
resample_both_dimensions = True)
def kspace_enc(reduce_resolution, reduce_FOV, resample_both_dimensions):
# based on above make a filter for k-space data
kfilter1 = np.zeros(kdata.shape)
kfilter2 = np.zeros(kdata.shape)
ksample = np.arange(-(FOVk/2-dk),(FOVk/2)+dk,dk)
kfilter1[:,np.where(abs(ksample)<((FOVk/2)*reduce_resolution))]=1
kfilter2[:,0:-1:reduce_FOV]=1
kfilter = kfilter1 * kfilter2
if resample_both_dimensions:
# also apply in AP direction
kfilter = kfilter * kfilter.conj().transpose()
print("Total fraction of k-space sampled = {}".format(100*np.size(np.where(kfilter != 0))/(Nx*Ny)))
kdata_sampled = kfilter * kdata
# invert FT
imencoded = np.fft.ifft2(np.fft.ifftshift(kdata_sampled))
fig, axes = plt.subplots(1, 2, figsize=(8, 4))
ax = axes.ravel()
ax[0].imshow(abs(kdata_sampled), cmap='gray', extent=[-(FOVk/2-dk), (FOVk/2)+dk, -(FOVk/2-dk), (FOVk/2)+dk],vmin=0, vmax=10)
ax[0].set_xlabel('k_x (rad/mm)')
ax[0].set_ylabel('k_y (rad/mm)')
ax[0].set_title('K-space Data')
ax[1].imshow(abs(imencoded), cmap='gray', extent=[0, 1/reduce_FOV, 0, 1/reduce_FOV])
ax[1].set_title('Reconstructed image')
ax[1].axis('off')
fig.tight_layout()
plt.show()
| MRI_kspace_encoding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="9goLeU8ewzry"
# # EM算法
# -
# 如果概率模型的变量都是观测变量(数据中可见的变量),则可以直接用极大似然估计,或者用贝叶斯估计模型参数。但是,当模型含有隐变量(数据中看不到的变量)时,就不能简单地使用这些估计方法,而应该使用含有隐变量的概率模型参数的极大似然估计法,EM算法
# EM算法就是个迭代优化的方法,之所以叫EM,是因为计算方法中每一次迭代都分两部,先求**期望步**再求**极大步**。基本思想是:首先根据己经给出的观测数据,估计出模型参数的值;然后再依据上一步估计出的参数值估计缺失数据的值,再根据估计出的缺失数据加上之前己经观测到的数据重新再对参数值进行估计,然后反复迭代,直至最后收敛,迭代结束。
# EM算法的推导和原理不难,需要掌握**极大似然估计**和**Jensen不等式**这两个基础知识。极大似然估计(MLE)在去年公司的数据分析课程里我有讲到^-^。接下来简单介绍下**Jensen不等式**。
# ## Jensen不等式
# 首先我们需要知道什么是凸函数,定义就是:
# - 设$f$是定义域为实数的函数,如果对所有的实数$x$,$f(x)$的二阶导数都大于0,那么$f$是凸函数。
# Jensen不等式的定义是:
# - 如果$f$是凸函数,$X$是随机变量,那么:$E[f(X)] \ge f(E[X])$ 。当且仅当$X$是常量时,该式取等号。其中,$E(X)$表示$X$的数学期望。
#
# Notice: 当Jensen不等式应用于凹函数时,不等号方向反向。当且仅当$X$是常量时,该不等式取等号
# 
# 这个例子就一目了然了。
# 现在举个经典例子:
# 有ABC三枚硬币,单次投掷出现正面的概率分别为$\pi$、$p$、$q$。利用这三枚硬币进行如下实验:
#
# 1、第一次先投掷A,若出现正面则投掷B,否则投掷C
#
# 2、记录第二次投掷的硬币出现的结果,正面记作1,反面记作0
#
# 独立重复1和2十次,产生如下观测结果:
#
# 1 1 0 1 0 0 1 0 1 1
#
# 假设只能观测到掷硬币的最终结果,无法观测第一次投掷的是哪一枚硬币,求$\pi$、$p$、$q$,即三硬币模型的参数。
#
# 记模型参数为$\theta=(\pi,p,q)$,无法观测的第一次投掷的硬币为随机变量$z$,可以观测的第二次投掷的硬币为随机变量$y$,则观测数据的似然函数为:$$P(Y|\theta)=\sum_{Z}P(Z|\theta)P(Y|Z,\theta)$$
# 意思就是,两个事件,第一个事件选出那枚看不到的硬币,第二个事件利用这枚硬币进行一次投掷。利用硬币结果只可能是0或1这个特性,可以将这个式子展开为:$$P(Y|\theta)=\prod^{n}_{j=1}[\pi p^{y_{j}}(1-p)^{1-y_{j}}+(1-\pi)q^{y_{j}}(1-q)^{1-y_{j}}]$$
#
# $y$的观测序列给定了,怎么找出一个模型参数,使得这个序列的概率(似然函数的值)最大呢,也就是求模型参数的极大似然估计:
# $$\tilde\theta = argmax[logP(Y|\theta)]$$
#
# 这个可以算是个NP问题,首先,给定模型参数我们可以直接MLE,但是参数组合其实挺多的,无法确定目前的是否是最优的。
# # EM的简单理解
# EM是个迭代算法(一定程度上更像个近似算法而不是精确算法)。可以大致分为三步:
# - 选取模型参数的初值:$\theta^{(0)}=(\pi^{(0)},p^{(0)},q^{(0)})$,循环如下两步迭代
# - E步(expectation):计算在当前迭代的模型参数下,观测数据$y$来自硬币$B$的概率:$$\mu^{(i+1)}=\frac{\pi^{i}(p^{(i)})^{y_{j}}(1-p^{(i)})^{1-y_{j}}}{\pi^{i}(p^{(i)})^{y_{j}}(1-p^{(i)})^{1-y_{j}}+(1-\pi^{i})(q^{(i)})^{y_{j}}(1-q^{(i)})^{1-y_{j}}}$$ 分子是选定B进行一次投掷实验的概率,分母是选定B或C,最终结果是结果来自B的概率。
# - M步(Maximization):估算下一个迭代的新的模型估算值,$$\pi^{(i+1)}=\frac{1}{n}\sum^{n}_{j=1}\mu^{(i+1)}_{j}$$ $$p^{(i+1)}=\frac{\sum^{n}_{j=1}\mu^{(i+1)}_{j}y_{j}}{\sum^{n}_{j=1}\mu^{(i+1)}_{j}}$$
# $$q^{(i+1)}=\frac{\sum^{n}_{j=1}(1-\mu^{(i+1)}_{j})y_{j}}{\sum^{n}_{j=1}(1-\mu^{(i+1)}_{j})}$$这个就是把这n个{试验结果来自B的概率}求和得到期望,平均后,得到B出正面的似然估计,同理有p和q。
#
# 重复迭代,直到收敛为止。
#
# 这个模型中,观测数据Y和隐数据Z组合在一起称为完全数据,单独的观测数据Y称为不完全数据。在隐数据未知的情况,无法直接估计Y的概率分布。但当模型概率给定时,就可以估计Y的条件概率分布了。
# ## EM的定义
# 输入:观测变量数据$Y$,隐变量数据$Z$,联合分布 $P(Y,Z|\theta)$,条件分布$P(Z|Y,\theta)$;
#
# 输出:模型参数$\theta$
# (1) 选择参数初始值$\theta^{(0)}$,开始迭代;
#
# (2) E步:记$\theta^{(i)}$为第i次迭代参数$\theta$的估计值,在第i+1次迭代的E步,计算$$Q(\theta,\theta^{(i)})=E_{Z}[logP(Y,Z|\theta)|Y,\theta^{(i)}]=\sum_{Z}logP(Y,Z|\theta)P(Z|Y,\theta^{(i)})$$
#
# $P(Z|Y,\theta^{(i)})$在这里表示在给定观测数据Y和当前的参数估计$\theta^{(i)}$的情况下,隐变量数据z的条件概率分布;
#
# (3) M步:求使$Q(\theta,\theta^{(i)})$极大化的$\theta$,确定第i+1次迭代的参数的估计值$\theta^{(i+1)}$ $$\theta^{(i+1)}=argmaxQ(\theta,\theta^{(i)})$$
#
# (4) 重复第(2)和第(3)步,直到收敛。
# ### 这个Q函数($Q(\theta,\theta{(i)})$)是EM算法最重要的。
# Definition:完全数据的对数似然函数$logP(Y,Z|\theta)$关于在给定观测数据Y和当前参数$\theta^{{i}}$下对未观测数据Z的条件概率分布$P(Z|Y,\theta^{(i)})$的期望称为Q函数
# ## 关于EM算法需要注意的
# - 步骤(1)参数的初值可以任意选择,但需注意EM算法对初值是敏感的
# - 步骤(2)E步求$Q(\theta,\theta^{(i)})$。$Q$函数式中$Z$是未观测数据,$Y$是观测数据。注意,$Q(\theta,\theta^{(i)})$的第1个变元表示要极大化的参数,第2个变元表示参数的当前估计值。每次迭代实际在求Q函数及其极大。
# - 步骤(3)M步求$Q(\theta,\theta^{(i)})$的极大化,得到$\theta^{(i)}$,完成一次迭代$\theta^{(i)}$到$\theta^{(i+1)}$。后面将证明每次迭代使似然函数增大或达到局部极值。
# - 步骤(4)给出停止迭代的条件,一般是对较小的正数$\epsilon_{1},\epsilon_{2}$,若满足则停止迭代。
# # EM算法手写计算的技巧
#
# EM算法真的还是有难度的,具体的一些计算的技巧和推倒需要网上找找,篇幅挺大的,什么Jensen不等式等各种技巧有很多。
# EM算法是通过不断求解下界的极大化逼近求解对数似然函数极大化的算法。在一个迭代内保证对数似然函数的增加的,迭代结束时无法保证对数似然函数是最大的。也就是说,EM算法不能保证找到全局最优值。
#
# 关于为什么估计大家也不会感兴趣,如果感兴趣的话可以上网找找,关键词是**EM算法的收敛性**。
#
# 事实证明呢,EM算法不能保证找到全局最优解,而且初值会影响最终结果。。。。。
#
# 那怎么办呢,只有多选几个初始值,然后都试下,最后选最好的那个。
# # EM的Python实现
# 上面是三硬币,接下来是双硬币模型:
# 假设有两枚硬币A、B,以相同的概率随机选择一个硬币,进行如下的抛硬币实验:共做5次实验,每次实验独立的抛十次,结果如图中a所示,例如某次实验产生了H、T、T、T、H、H、T、H、T、H,H代表正面朝上。
#
# 假设试验数据记录员可能是实习生,业务不一定熟悉,造成a和b两种情况
#
# a表示实习生记录了详细的试验数据,我们可以观测到试验数据中每次选择的是A还是B
#
# b表示实习生忘了记录每次试验选择的是A还是B,我们无法观测实验数据中选择的硬币是哪个
#
# 问在两种情况下分别如何估计两个硬币正面出现的概率?
# ---------------
# a是用极大似然做,b则是用EM做,如下:
# a既然能观测到试验数据是哪枚硬币产生的,就可以统计正反面的出现次数,直接利用最大似然估计即可。
#
# b情况就无法直接进行最大似然估计了,只能用EM算法。
# # python实现
import numpy as np
from scipy import stats
# 我们先将EM算法原论文代码拆开。如下所示:
# 首先,采集数据,用1表示H(正面),0表示T(反面):
# 硬币投掷结果观测序列
observations = np.array([[1, 0, 0, 0, 1, 1, 0, 1, 0, 1],
[1, 1, 1, 1, 0, 1, 1, 1, 1, 1],
[1, 0, 1, 1, 1, 1, 1, 0, 1, 1],
[1, 0, 1, 0, 0, 0, 1, 1, 0, 0],
[0, 1, 1, 1, 0, 1, 1, 1, 0, 1]])
# 接下来,我们需要选定初始值: $$\theta^{0}_{A}=0.6$$ $$\theta^{0}_{B}=0.5$$
# #### 第一次迭代的E
# 我们看到第一行数据,正反各5次;其次,抛硬币是二项分布,那么
coin_A_pmf_observation_1 = stats.binom.pmf(5,10,0.6)
coin_A_pmf_observation_1
# 同理,也可以求出第一行由B产生的概率
coin_B_pmf_observation_1 = stats.binom.pmf(5,10,0.5)
coin_B_pmf_observation_1
# 然后normalize一下
normalized_coin_A_pmf_observation_1 = coin_A_pmf_observation_1/(coin_A_pmf_observation_1+coin_B_pmf_observation_1)
print ("%0.2f" %normalized_coin_A_pmf_observation_1)
# 接下来算是剩余四行的$\mu$了
# ### 更新在当前参数下A、B硬币产生的正反面次数
# counts['A']['H'] += weight_A * num_heads <br/>
# counts['A']['T'] += weight_A * num_tails <br/>
# counts['B']['H'] += weight_B * num_heads <br/>
# counts['B']['T'] += weight_B * num_tails
# #### 第一个迭代的M步
# new_theta_A = counts['A']['H'] / (counts['A']['H'] + counts['A']['T']) <br/>
# new_theta_B = counts['B']['H'] / (counts['B']['H'] + counts['B']['T']
# 到这就是单次迭代EM算法的逻辑,原代码如下:
def em_single(priors, observations):
"""
EM算法单次迭代
Arguments
---------
priors先验 : [theta_A, theta_B]
observations : [m X n matrix]
Returns
--------
new_priors更新后的先验: [new_theta_A, new_theta_B]
:param priors:
:param observations:
:return:
"""
counts = {'A': {'H': 0, 'T': 0}, 'B': {'H': 0, 'T': 0}}
theta_A = priors[0]
theta_B = priors[1]
# E step
for observation in observations:
len_observation = len(observation)
num_heads = observation.sum()
num_tails = len_observation - num_heads
contribution_A = stats.binom.pmf(num_heads, len_observation, theta_A)
contribution_B = stats.binom.pmf(num_heads, len_observation, theta_B) # 两个二项分布
weight_A = contribution_A / (contribution_A + contribution_B) # A的百分比
weight_B = contribution_B / (contribution_A + contribution_B) # B的百分比
# 更新在当前参数下A、B硬币产生的正反面次数
counts['A']['H'] += weight_A * num_heads
counts['A']['T'] += weight_A * num_tails
counts['B']['H'] += weight_B * num_heads
counts['B']['T'] += weight_B * num_tails
# M step
new_theta_A = counts['A']['H'] / (counts['A']['H'] + counts['A']['T'])
new_theta_B = counts['B']['H'] / (counts['B']['H'] + counts['B']['T'])
return [new_theta_A, new_theta_B]
em_single([0.6,0.5],observations)
# ## EM的主循环
# 给定循环的两个终止条件:模型参数变化小于阈值;循环达到最大次数,就可以写出EM算法的主循环了:
def em(observations, prior, tol=1e-6, iterations=10000):
"""
EM算法
:param observations: 观测数据
:param prior: 模型初值
:param tol: 迭代结束阈值
:param iterations: 最大迭代次数
:return: 局部最优的模型参数
"""
import math
iteration = 0
while iteration < iterations:
new_prior = em_single(prior, observations)
delta_change = np.abs(prior[0] - new_prior[0])
if delta_change < tol:
break
else:
prior = new_prior
iteration += 1
return [new_prior, iteration]
em(observations, [0.6, 0.5])
# 第十四次停止收敛,$$\tilde\theta^{(10)}_{A} \approx 0.80$$ $$\tilde\theta^{(10)}_{B} \approx 0.52$$
# 试着改变一下初始值,试验初值对EM的影响
em(observations, [0.5,0.6])
em(observations, [0.3,0.9])
em(observations, [0.4,0.4])
em(observations, [0.9999,0.0001])
# EM还是挺靠谱的
# -------
# # 下面这个例子选修
# + [markdown] colab_type="text" id="Oewoj4r315fN"
# ##### 高斯混合模型 GMM
# 这一例子是求GMM的参数近似值
# + colab={} colab_type="code" id="_OrYQBcP104s"
class GMM:
def __init__(self, K, weights=None,means=None,covars=None, sigma=0.0000001):
"""
:K: component
:weigths: init weights
:means: Gaussain mean
:covars: Gaussain covars
"""
self.K = K
if weights is not None:
self.weights = weights
else:
self.weights = np.random.rand(self.K)
self.weights /= np.sum(self.K) # normlize, to make sure sum = 1
self.means = means
self.covars = covars
self.sigma = sigma
def _init_param(self, X):
col = X.shape[1]
if self.means is None:
self.means = []
for k in range(self.K):
mean = np.random.rand(col)
self.means.append(mean)
if self.covars is None:
self.covars = []
for k in range(self.K):
cov = np.random.rand(col, col)
self.covars.append(cov)
def Gaussian(self, x, mean, cov):
dim = np.shape(cov)[0]
# cov的行列式为零时的措施
covdet = np.linalg.det(cov + np.eye(dim) * 0.01) #协方差矩阵的行列式
covinv = np.linalg.inv(cov + np.eye(dim) * 0.01) #协方差矩阵的逆
xdiff = (x - mean).reshape((1,dim))
# 概率密度
prob = 1.0/(np.power(np.power(2*np.pi,dim)*np.abs(covdet),0.5))* np.exp(-0.5*xdiff.dot(covinv).dot(xdiff.T))[0][0]
return prob
def fit(self, X):
self._init_param(X)
loglikelyhood = 0.
oldloglikelyhood = 1.
length,dim = np.shape(X)
# gamma表示第n个样本属于第k个混合高斯的概率
gammas = [np.zeros(self.K) for i in range(length)]
while np.abs(loglikelyhood-oldloglikelyhood) > self.sigma:
oldloglikelyhood = loglikelyhood
# E-step
for n in range(length):
# respons是GMM的EM算法中的权重w,即后验概率
respons = [self.weights[k] * self.Gaussian(X[n], self.means[k], self.covars[k])
for k in range(self.K)]
sum_respons = np.sum(respons)
gammas[n] = respons/sum_respons
# M-step
for k in range(self.K):
#nk表示N个样本中有多少属于第k个高斯
nk = np.sum([gammas[n][k] for n in range(length)])
# 更新每个高斯分布的概率
self.weights[k] = 1.0 * nk / length
# 更新高斯分布的均值
self.means[k] = (1.0/nk) * np.sum([gammas[n][k] * X[n] for n in range(length)], axis=0)
xdiffs = X - self.means[k]
# 更新高斯分布的协方差矩阵
self.covars[k] = (1.0/nk)*np.sum([gammas[n][k]*xdiffs[n].reshape((dim,1)).dot(xdiffs[n].reshape((1,dim))) for n in range(length)],axis=0)
loglikelyhood = []
for n in range(length):
tmp = [np.sum(self.weights[k]*self.Gaussian(X[n],self.means[k],self.covars[k])) for k in range(self.K)]
tmp = np.log(np.array(tmp))
loglikelyhood.append(list(tmp))
loglikelyhood = np.sum(loglikelyhood)
for i in range(length):
gammas[i] = gammas[i]/np.sum(gammas[i])
self.posibility = gammas
self.prediction = [np.argmax(gammas[i]) for i in range(length)]
return self.weights, self.means, self.covars
#https://blog.csdn.net/qq_30091945/article/details/81134598
# + [markdown] colab_type="text" id="FAzKJKu6W2C7"
# ##### iris数据集
# + colab={"base_uri": "https://localhost:8080/", "height": 125} colab_type="code" id="19XKdlFhAPnA" outputId="1008307f-c3f2-4073-a937-9fe9a7495d72"
# 导入Iris数据集
iris = load_iris()
label = np.array(iris.target)
data = np.array(iris.data)
print("Iris label:\n",label)
# + colab={"base_uri": "https://localhost:8080/", "height": 281} colab_type="code" id="ghZH6RnpUUP1" outputId="57242957-ccc8-448b-bdf4-9e77e2899b96"
# 对数据进行预处理
data = Normalizer().fit_transform(data)
# 数据可视化
plt.scatter(data[:,0],data[:,1],c = label)
plt.title("Iris")
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 373} colab_type="code" id="Ov1PVnJQUk9L" outputId="8b872c16-a8e7-4d5d-df20-1f289f779de3"
# GMM模型
K = 3
gmm = GMM(K)
gmm.fit(data)
y_pre = gmm.prediction
print("GMM预测结果:\n",y_pre)
print("GMM正确率为:\n",accuracy_score(label,y_pre))
plt.scatter(data[:, 0], data[:, 1], c=y_pre)
plt.title("GMM")
plt.show()
| EM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# <hr>
# ### Kalman Filters and Pairs Trading
#
# There are a few Python packages out there for Kalman filters, but we're adapting this example and the Kalman filter class code from [this article](https://www.quantstart.com/articles/kalman-filter-based-pairs-trading-strategy-in-qstrader) and demonstrating how you can implement similar ideas using QuantConnect!
#
# Briefly, a Kalman filter is a [state-space model](https://en.wikipedia.org/wiki/State-space_representation) applicable to linear dynamic systems -- systems whose state is time-dependent and state variations are represented linearly. The model is used to estimate unknown states of a variable based on a series of past values. The procedure is two-fold: a prediction (estimate) is made by the filter of the current state of a variable and the uncertainty of the estimate itself. When new data is available, these estimates are updated. There is a lot of information available about Kalman filters, and the variety of their applications is pretty astounding, but for now, we're going to use a Kalman filter to estimate the hedge ratio between a pair of equities.
#
# The idea behind the strategy is pretty straightforward: take two equities that are cointegrated and create a long-short portfolio. The premise of this is that the spread between the value of our two positions should be mean-reverting. Anytime the spread deviates from its expected value, one of the assets moved in an unexpected direction and is due to revert back. When the spread diverges, you can take advantage of this by going long or short on the spread.
#
# To illustrate, imagine you have a long position in AAPL worth \\$2000 and a short position in IBM worth \\$2000. This gives you a net spread of \\$0. Since you expected AAPL and IBM to move together, then if the spread increases significantly above \\$0, you would short the spread in the expectation that it will return to \\$0, it's natural equilibrium. Similarly, if the value drops significantly below \\$0, you would long the spread and capture the profits as its value returns to \\$0. In our application, the Kalman filter will be used to track the hedging ratio between our equities to ensure that the portfolio value is stationary, which means it will continue to exhibit mean-reversion behavior.
# ##### Note: Run the final cell first so the remaining cells will execute
# +
# QuantBook Analysis Tool
# For more information see [https://www.quantconnect.com/docs/research/overview]
import numpy as np
from math import floor
import matplotlib.pyplot as plt
from KalmanFilter import KalmanFilter
qb = QuantBook()
symbols = [qb.AddEquity(x).Symbol for x in ['VIA', 'VIAB']]
# -
# Now, we initialize the Kalman Filter, grab our data, and then run the Kalman Filter update process over the data.
kf = KalmanFilter()
history = qb.History(qb.Securities.Keys, datetime(2019, 1, 1), datetime(2019, 1, 11), Resolution.Daily)
prices = history.unstack(level=1).close.transpose()
for index, row in prices.iterrows():
via = row.loc[str(symbols[0].ID)]
viab = row.loc[str(symbols[1].ID)]
forecast_error, prediction_std_dev, hedge_quantity = kf.update(via, viab)
print(f'{forecast_error} :: {prediction_std_dev} :: {hedge_quantity}')
# In an algorithm, the <em>kf.qty</em> variable is the number of shares to invested in VIAB, and <em>hedge_quantity</em> is the amount to trade in the opposite direction for VIA
# ##### Code for the Kalman Filter
# +
import numpy as np
from math import floor
class KalmanFilter:
def __init__(self):
self.delta = 1e-4
self.wt = self.delta / (1 - self.delta) * np.eye(2)
self.vt = 1e-3
self.theta = np.zeros(2)
self.P = np.zeros((2, 2))
self.R = None
self.qty = 2000
def update(self, price_one, price_two):
# Create the observation matrix of the latest prices
# of TLT and the intercept value (1.0)
F = np.asarray([price_one, 1.0]).reshape((1, 2))
y = price_two
# The prior value of the states \theta_t is
# distributed as a multivariate Gaussian with
# mean a_t and variance-covariance R_t
if self.R is not None:
self.R = self.C + self.wt
else:
self.R = np.zeros((2, 2))
# Calculate the Kalman Filter update
# ----------------------------------
# Calculate prediction of new observation
# as well as forecast error of that prediction
yhat = F.dot(self.theta)
et = y - yhat
# Q_t is the variance of the prediction of
# observations and hence \sqrt{Q_t} is the
# standard deviation of the predictions
Qt = F.dot(self.R).dot(F.T) + self.vt
sqrt_Qt = np.sqrt(Qt)
# The posterior value of the states \theta_t is
# distributed as a multivariate Gaussian with mean
# m_t and variance-covariance C_t
At = self.R.dot(F.T) / Qt
self.theta = self.theta + At.flatten() * et
self.C = self.R - At * F.dot(self.R)
hedge_quantity = int(floor(self.qty*self.theta[0]))
return et, sqrt_Qt, hedge_quantity
# -
| Research2Production/Python/04 Kalman Filters and Pairs Trading.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/paulcodrea/dissertation/blob/main/5a_LSTM_prediction_BTC_price.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="DnXDwfZHA9pZ" outputId="328e5ed5-e03e-499a-e69b-26f91160a714"
# !pip install wandb --quiet
# + colab={"base_uri": "https://localhost:8080/"} id="MoL6M_NEFGKS" outputId="578b7d3f-992b-4f03-8dc5-d2e1db269df0"
# !wandb login
# + id="B4cHGHj8G1XC"
import os
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import wandb
from wandb.keras import WandbCallback
from time import time
from keras.models import Sequential, load_model
from keras.layers.core import Dense, Dropout
from keras.layers import LSTM
from time import time
from keras.callbacks import EarlyStopping
from sklearn.preprocessing import MinMaxScaler
# + colab={"base_uri": "https://localhost:8080/", "height": 110} id="Snl-yrkWBbq2" outputId="a78add3f-2937-45d3-d45a-41cae145a034"
wandb.init(project="5a_LSTM-precition_BTC_price", entity="paulcodrea")
# + id="bKD0YDNFCk_U"
wandb.config = {
"learning_rate": 0.001,
"epochs": 40,
"batch_size": 4,
"train_p": 0.55,
"val_p": 0.05,
"LSTM_layer": [50, 100],
"Dropout_layer": [0.15, 0.2],
"activation": 'tanh',
"timesteps": 1,
}
# Construct the metrics to store the results
metrics_df = pd.DataFrame(columns=['script','epoch', 'batch_size','timesteps',
'train_p', 'val_p', 'test_p', 'loss', 'val_loss',
'mse', 'rmse', 'mae', 'mape', 'runtime',
'cross_correlation'])
# + id="TIAuBmM0ISML"
df = pd.read_csv('/content/drive/MyDrive/COMP30030_Dissertation_paul.codrea/Market-prediction/btc_only_specific_time.csv', parse_dates=True, index_col="time")
df.drop(columns=['Unnamed: 0'], inplace=True)
# + id="OmsP9D6IIgN_"
# Data normalization. This is one of the first steps to normalize the values.
# The goal is to change the values of numeric columns in the data set to a common
# scale, wihtout distorting differeces in the randes of values.
df_values = df.values
sc = MinMaxScaler()
data_scaled = sc.fit_transform(df_values)
# + id="z9F_IGvLQqgN"
# Splitting data into training and testing data
train_index = int(wandb.config['train_p'] * len(data_scaled))
val_index = train_index + int(wandb.config['val_p'] *
len(data_scaled))
train = data_scaled[:train_index]
val = data_scaled[train_index:val_index]
test = data_scaled[val_index:]
# + colab={"base_uri": "https://localhost:8080/"} id="q_9sp2nbhpPi" outputId="137602c4-fde9-412c-8335-159d74b5854e"
print("train,test,val",train.shape, test.shape, val.shape)
# + id="mc0zxz7LiQWH"
xtrain , ytrain = train[:, :1], train[:, 0]
xtest, ytest = test[:, :1], test[:, 0]
xval , yval = val[:, :1], val[:, 0]
# + colab={"base_uri": "https://localhost:8080/"} id="daLgJ_8OTICC" outputId="3d17e51b-6f75-47a7-d39c-753f18d792b8"
# Samples -> these are the rows in the data.
# Number of hours in the future?
# Timesteps -> these are the past observations for a feature (such as lag variable).
# input_shape is the shape of the training dataset.
timesteps = wandb.config["timesteps"]
n_features = 1 # Columns in the data. Should be set to 1 since we only have 'close' column.
train_len = len(xtrain) - timesteps
test_len = len(xtest) - timesteps
val_len = len(xval) - timesteps
x_train = np.zeros((train_len, timesteps, n_features))
y_train = np.zeros((train_len))
for i in range(train_len):
ytemp = i + timesteps
x_train[i] = xtrain[i: ytemp]
y_train[i] = ytrain[ytemp]
print("x_train", x_train.shape)
print("y_train", y_train.shape)
x_test = np.zeros((test_len, timesteps, n_features))
y_test = np.zeros((test_len))
for i in range(test_len):
ytemp = i + timesteps
x_test[i] = xtest[i: ytemp]
y_test[i] = ytest[ytemp]
print("x_test", x_test.shape)
print("y_test", y_test.shape)
x_val = np.zeros((val_len, timesteps, n_features))
y_val = np.zeros((val_len))
for i in range(val_len):
ytemp = i + timesteps
x_val[i] = xval[i: ytemp]
y_val[i] = yval[ytemp]
print("x_val", x_val.shape)
print("y_val", y_val.shape)
################################################################################
# 2 hidden layers with 50 neurons each and a dropout between every one of them.
# Start with Sequencial class.
model = Sequential()
# return_sequence will return a sequence rather than a single value for each input.
# Sequential model -> as apipeline with raw data fed in
model.add(LSTM(wandb.config['LSTM_layer'][0], input_shape = (timesteps, n_features),
return_sequences=True, activation=wandb.config['activation']))
# Dropout Regularisation - method of ignoring and dropping random units during training.
# This is essential to prevent overfitting. e.g. Dropout of 15%
model.add(Dropout(wandb.config['Dropout_layer'][0]))
model.add(LSTM(wandb.config['LSTM_layer'][1], activation=wandb.config['activation']))
model.add(Dropout(wandb.config['Dropout_layer'][1]))
model.add(Dense(1)) # This layer is at the end of the architecture and it is used for outputting a prediction.
print(model.summary())
# mean-squared-error loss function and Adam optimiser. MSE is a standard loss function for a regression model.
# adam -> optimiser algorithm
model.compile(loss = 'mean_squared_error', optimizer = 'adam')
# Too many epochs can lead to overfitting of the training dataset, whereas too few may result in an underfit model.
# Early stopping is a method that allows you to specify an arbitrary large number of training epochs and stop training
# once the model performance stops improving on a hold out validation dataset.
earlystop = EarlyStopping(monitor='val_loss', min_delta=0.0001, patience=80,
verbose=1, mode='min')
start = time()
print("start:",0)
history = model.fit(x_train, y_train, epochs = wandb.config['epochs'],
batch_size=wandb.config['batch_size'],
validation_data=(x_val, y_val), verbose = 1,
shuffle = False, callbacks=[WandbCallback(), earlystop])
# Print the time it took to run the code
runtime = time()-start
print("Time: %.4f" % runtime)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="UyGCR4EZiqtF" outputId="dba9d5f5-2e28-411f-d163-178d67862ea2"
# Plotting data
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(loss))
plt.figure(figsize=(10, 8))
plt.plot(epochs, loss, 'b', label='Training loss')
plt.plot(epochs, val_loss, 'orange', label='Validation loss')
plt.title("Training and Validation loss")
plt.legend()
plt.show()
# Predict the model
y_pred = model.predict(x_test)
# Print out Mean Squared Error (MSE)
mse = np.mean((y_pred - y_test)**2)
print("MSE: %.4f" % mse)
# Print out Root Mean Squared Error (RMSE)
rmse = np.sqrt(mse)
print("RMSE: %.4f" % rmse)
# Print out Mean Absolute Error (MAE)
mae = np.mean(np.abs(y_pred - y_test))
print("MAE: %.4f" % mae)
# Print out Mean Absolute Percentage Error (MAPE)
mape = np.mean(np.abs((y_pred - y_test) / y_test)) * 100
print("MAPE (percentage): %.4f" % mape)
# Invers Scaling
actual_price = sc.inverse_transform([y_test])
actual_price = np.reshape(actual_price, (actual_price.shape[1], 1))
predicted_price = sc.inverse_transform(y_pred)
# Plotting the prediction
plt.figure(figsize=(18,8))
plt.plot(actual_price, '.-', color='red', label='Real market values', alpha=0.5)
plt.plot(predicted_price, '.-', color='blue', label='Predicted values', alpha=1)
# fill between actual price and x-axis
# plt.fill_between(range(len(actual_price)), actual_price, predicted_price, color='blue', alpha=0.2)
plt.title("Bitcoin Price Prediction using RNN-LSTM")
plt.xlabel("Units of time")
plt.ylabel("Price")
plt.legend()
plt.show()
# + [markdown] id="MNdEeny9Uz7r"
# ## Cross-correlation analysis
# + id="WlHmxF4YU2gS"
actual_price_df = pd.DataFrame(y_test, columns=['price'])
predicted_price_df = pd.DataFrame(y_pred, columns=['price'])
# + id="3ReLNFjPUrOD"
def crosscorr(datax, datay, lag=0, method="pearson"):
""" Lag-N cross correlation.
Parameters
—------—
lag : int, default 0
datax, datay : pandas.Series objects of equal length
Returns
—------—
crosscorr : float
"""
return datax.corr(datay.shift(lag), method=method)
# + colab={"base_uri": "https://localhost:8080/", "height": 715} id="4BqY1MIaUuJE" outputId="1efcd4d2-0fa7-4c10-cd98-00b9d57e3c06"
# Calculate the Pearson Cross Correlation for lag 0
curr_corr = crosscorr(predicted_price_df['price'], actual_price_df['price'],
method="pearson")
# Print out the Correlation for lag 0
print(f"Pearson CC for lag 0: {curr_corr}")
# Process of improving the CC value
xcov = [crosscorr(predicted_price_df['price'], actual_price_df['price'], lag=i, method="pearson") for i in range(-len(actual_price_df), len(actual_price_df))]
# Identify the lag that maximizes the correlation
lag_max = np.argmax(xcov)
print(f"Lag that maximizes the correlation {lag_max}\n\n")
plt.figure(figsize=(18,8))
plt.plot(xcov, '.-', color='blue', label='Cross-correlation', alpha=1)
plt.title("Cross-correlation between actual and predicted values")
plt.xlabel("Units of time")
plt.ylabel("Cross-correlation")
plt.legend()
plt.show()
# If the Lag is different from 0 then shift the predicted price df and plot again
if lag_max != 0:
# Use the lag value to shift the predicted values to align with the actual values
predicted_price_df['price'] = predicted_price_df['price'].shift(lag_max)
plt.figure(figsize=(18,8))
plt.plot(predicted_price_df, '.-', color='blue', label='Predicted values', alpha=1)
plt.plot(actual_price_df, '.-', color='red', label='Real market values', alpha=0.5)
plt.title("Predicted values")
plt.xlabel("Units of time")
plt.ylabel("Price")
plt.legend()
else:
print("\n\n No changes to current plotting")
# + [markdown] id="s6I9TVnI-bN8"
# ### Save metrics
# + id="MeymLX3s-efS"
# add values to metrics_df
metrics_df = metrics_df.append({'script': 'LSTM_price',
'epoch': wandb.config['epochs'],
'batch_size': wandb.config['batch_size'],
'timesteps': wandb.config['timesteps'],
'train_p': wandb.config['train_p'] * 100,
'val_p': wandb.config['val_p'] * 100,
'test_p': (1 - wandb.config['train_p'] - wandb.config['val_p']) * 100,
'loss': min(loss), 'val_loss': min(val_loss),
'mse': mse, 'rmse': rmse, 'mae': mae, 'mape': mape,
'runtime': runtime, 'cross_correlation':curr_corr},
ignore_index=True)
# + [markdown] id="ZNan1CD1dKM_"
# ## Forecast upcoming 'n_hours'
# + id="I8vSGGpvtuQz"
n_hours = 4
# Prepare dataset based on xtest
xtest_copy = xtest.copy()
xtest_copy = np.append(xtest_copy, np.zeros((n_hours, n_features)), axis=0)
# Forecast the price based on the model build
y_pred_forecast = model.predict(xtest_copy)
forecast_price = sc.inverse_transform(y_pred_forecast)
# + colab={"base_uri": "https://localhost:8080/", "height": 513} id="buOFOQAcfmbZ" outputId="62817cff-2f14-4e70-db79-77ac98664774"
plt.figure(figsize=(18,8))
plt.plot(actual_price, '.-', color='red', label='Real market values', alpha=0.5)
plt.plot(forecast_price, '.-', color='blue', label='Predicted values', alpha=1)
plt.title("Bitcoin Price Prediction using RNN-LSTM")
plt.xlabel("Time")
plt.ylabel("Price")
plt.legend()
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="zo6dJU1qqnFW" outputId="8b6464d8-16c3-449d-d191-67efeed9a162"
# Print out statistics of the forecasted price
print(f"Forecasted price: {forecast_price[-1]}")
# Compare the price based on the n_hours before the actual price
print(f"Actual price: {actual_price[-1]}")
# Print out if the price will go down or up
if forecast_price[-1] > actual_price[-1]:
print(f"Price will go up in the following {n_hours} hours")
else:
print(f"Price will go down in the following {n_hours} hours")
| 5a_LSTM_prediction_BTC_price.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # BigQuery ML models with feature engineering
#
# In this notebook, we will use BigQuery ML to build more sophisticated models for taxifare prediction.
#
# This is a continuation of our [first models](../../02_bqml/solution/first_model.ipynb) we created earlier with BigQuery ML but now with more feature engineering.
#
# ## Learning Objectives
# 1. Apply transformations using SQL to prune the taxi cab dataset
# 2. Create and train a new Linear Regression model with BigQuery ML
# 3. Evaluate and predict with the linear model
# 4. Create a feature cross for day-hour combination using SQL
# 5. Examine ways to reduce model overfitting with regularization
# 6. Create and train a DNN model with BigQuery ML
#
# Each learning objective will correspond to a __#TODO__ in the [student lab notebook](../labs/feateng_bqml.ipynb) -- try to complete that notebook first before reviewing this solution notebook.
# + language="bash"
# export PROJECT=$(gcloud config list project --format "value(core.project)")
# echo "Your current GCP Project Name is: "$PROJECT
# +
import os
PROJECT = "your-gcp-project-here" # REPLACE WITH YOUR PROJECT NAME
REGION = "us-central1" # REPLACE WITH YOUR BUCKET REGION e.g. us-central1
# Do not change these
os.environ["PROJECT"] = PROJECT
os.environ["REGION"] = REGION
os.environ["BUCKET"] = PROJECT # DEFAULT BUCKET WILL BE PROJECT ID
if PROJECT == "your-gcp-project-here":
print("Don't forget to update your PROJECT name! Currently:", PROJECT)
# -
# ## Create a BigQuery Dataset and Google Cloud Storage Bucket
#
# A BigQuery dataset is a container for tables, views, and models built with BigQuery ML. Let's create one called __serverlessml__ if we have not already done so in an earlier lab. We'll do the same for a GCS bucket for our project too.
# + language="bash"
#
# ## Create a BigQuery dataset for serverlessml if it doesn't exist
# datasetexists=$(bq ls -d | grep -w serverlessml)
#
# if [ -n "$datasetexists" ]; then
# echo -e "BigQuery dataset already exists, let's not recreate it."
#
# else
# echo "Creating BigQuery dataset titled: serverlessml"
#
# bq --location=US mk --dataset \
# --description 'Taxi Fare' \
# $PROJECT:serverlessml
# echo "\nHere are your current datasets:"
# bq ls
# fi
#
# ## Create GCS bucket if it doesn't exist already...
# exists=$(gsutil ls -d | grep -w gs://${PROJECT}/)
#
# if [ -n "$exists" ]; then
# echo -e "Bucket exists, let's not recreate it."
#
# else
# echo "Creating a new GCS bucket."
# gsutil mb -l ${REGION} gs://${PROJECT}
# echo "\nHere are your current buckets:"
# gsutil ls
# fi
# -
# ## Model 4: With some transformations
#
# BigQuery ML automatically scales the inputs. so we don't need to do scaling, but human insight can help.
#
# Since we we'll repeat this quite a bit, let's make a dataset with 1 million rows.
# +
# %%bigquery
CREATE OR REPLACE TABLE serverlessml.feateng_training_data AS
SELECT
(tolls_amount + fare_amount) AS fare_amount,
pickup_datetime,
pickup_longitude AS pickuplon,
pickup_latitude AS pickuplat,
dropoff_longitude AS dropofflon,
dropoff_latitude AS dropofflat,
passenger_count*1.0 AS passengers
FROM `nyc-tlc.yellow.trips`
# The full dataset has 1+ Billion rows, let's take only 1 out of 1,000 (or 1 Million total)
WHERE MOD(ABS(FARM_FINGERPRINT(CAST(pickup_datetime AS STRING))), 1000) = 1
AND
trip_distance > 0
AND fare_amount >= 2.5
AND pickup_longitude > -78
AND pickup_longitude < -70
AND dropoff_longitude > -78
AND dropoff_longitude < -70
AND pickup_latitude > 37
AND pickup_latitude < 45
AND dropoff_latitude > 37
AND dropoff_latitude < 45
AND passenger_count > 0
# +
# %%bigquery
# Tip: You can CREATE MODEL IF NOT EXISTS as well
CREATE OR REPLACE MODEL serverlessml.model4_feateng
TRANSFORM(
* EXCEPT(pickup_datetime)
, SQRT( (pickuplon-dropofflon)*(pickuplon-dropofflon) + (pickuplat-dropofflat)*(pickuplat-dropofflat) ) AS euclidean
, CAST(EXTRACT(DAYOFWEEK FROM pickup_datetime) AS STRING) AS dayofweek
, CAST(EXTRACT(HOUR FROM pickup_datetime) AS STRING) AS hourofday
)
OPTIONS(input_label_cols=['fare_amount'], model_type='linear_reg')
AS
SELECT * FROM serverlessml.feateng_training_data
# -
# Once the training is done, visit the [BigQuery Cloud Console](https://console.cloud.google.com/bigquery) and look at the model that has been trained. Then, come back to this notebook.
# Note that BigQuery automatically split the data we gave it, and trained on only a part of the data and used the rest for evaluation. We can look at eval statistics on that held-out data:
# %%bigquery
SELECT *, SQRT(loss) AS rmse FROM ML.TRAINING_INFO(MODEL serverlessml.model4_feateng)
# %%bigquery
SELECT SQRT(mean_squared_error) AS rmse FROM ML.EVALUATE(MODEL serverlessml.model4_feateng)
# Yippee! We're now below our target of 6 dollars in RMSE.
# We are now beating our goals, and with just a linear model.
#
# ## Making predictions with BigQuery ML
#
# This is how the prediction query would look that we saw earlier [heading 1.3 miles uptown](https://www.google.com/maps/dir/'40.742104,-73.982683'/'40.755174,-73.983766'/@40.7481394,-73.993579,15z/data=!3m1!4b1!4m9!4m8!1m3!2m2!1d-73.982683!2d40.742104!1m3!2m2!1d-73.983766!2d40.755174) in New York City.
# %%bigquery
SELECT * FROM ML.PREDICT(MODEL serverlessml.model4_feateng, (
SELECT
-73.982683 AS pickuplon,
40.742104 AS pickuplat,
-73.983766 AS dropofflon,
40.755174 AS dropofflat,
3.0 AS passengers,
TIMESTAMP('2019-06-03 04:21:29.769443 UTC') AS pickup_datetime
))
# ## Improving the model with feature crosses
#
# Let's do a [feature cross](https://developers.google.com/machine-learning/crash-course/feature-crosses/video-lecture) of the day-hour combination instead of using them raw
# +
# %%bigquery
CREATE OR REPLACE MODEL serverlessml.model5_featcross
TRANSFORM(
* EXCEPT(pickup_datetime)
, SQRT( (pickuplon-dropofflon)*(pickuplon-dropofflon) + (pickuplat-dropofflat)*(pickuplat-dropofflat) ) AS euclidean
, ML.FEATURE_CROSS(STRUCT(CAST(EXTRACT(DAYOFWEEK FROM pickup_datetime) AS STRING) AS dayofweek,
CAST(EXTRACT(HOUR FROM pickup_datetime) AS STRING) AS hourofday)) AS day_hr
)
OPTIONS(input_label_cols=['fare_amount'], model_type='linear_reg')
AS
SELECT * FROM serverlessml.feateng_training_data
# -
# %%bigquery
SELECT *, SQRT(loss) AS rmse FROM ML.TRAINING_INFO(MODEL serverlessml.model5_featcross)
# %%bigquery
SELECT SQRT(mean_squared_error) AS rmse FROM ML.EVALUATE(MODEL serverlessml.model5_featcross)
# Sometimes (not the case above), the training RMSE is quite reasonable, but the evaluation RMSE is terrible. This is an indication of overfitting.
# When we do feature crosses, we run into the risk of overfitting (for example, when a particular day-hour combo doesn't have enough taxirides).
#
# ## Reducing overfitting
#
# Let's add [L2 regularization](https://developers.google.com/machine-learning/glossary/#L2_regularization) to help reduce overfitting. Let's set it to 0.1
# +
# %%bigquery
CREATE OR REPLACE MODEL serverlessml.model6_featcross_l2
TRANSFORM(
* EXCEPT(pickup_datetime)
, SQRT( (pickuplon-dropofflon)*(pickuplon-dropofflon) + (pickuplat-dropofflat)*(pickuplat-dropofflat) ) AS euclidean
, ML.FEATURE_CROSS(STRUCT(CAST(EXTRACT(DAYOFWEEK FROM pickup_datetime) AS STRING) AS dayofweek,
CAST(EXTRACT(HOUR FROM pickup_datetime) AS STRING) AS hourofday)) AS day_hr
)
OPTIONS(input_label_cols=['fare_amount'], model_type='linear_reg', l2_reg=0.1)
AS
SELECT * FROM serverlessml.feateng_training_data
# -
# %%bigquery
SELECT SQRT(mean_squared_error) AS rmse FROM ML.EVALUATE(MODEL serverlessml.model6_featcross_l2)
# These sorts of experiment would have taken days to do otherwise. We did it in minutes, thanks to BigQuery ML! The advantage of doing all this in the TRANSFORM is the client code doing the PREDICT doesn't change. Our model improvement is transparent to client code.
# %%bigquery
SELECT * FROM ML.PREDICT(MODEL serverlessml.model6_featcross_l2, (
SELECT
-73.982683 AS pickuplon,
40.742104 AS pickuplat,
-73.983766 AS dropofflon,
40.755174 AS dropofflat,
3.0 AS passengers,
TIMESTAMP('2019-06-03 04:21:29.769443 UTC') AS pickup_datetime
))
# ## Let's try feature crossing the locations too
#
# Because the lat and lon by themselves don't have meaning, but only in conjunction, it may be useful to treat the fields as a pair instead of just using them as numeric values. However, lat and lon are continuous numbers, so we have to discretize them first. That's what ML.BUCKETIZE does.
#
# Here are some of the preprocessing functions in BigQuery ML:
# * ML.FEATURE_CROSS(STRUCT(features)) does a feature cross of all the combinations
# * ML.POLYNOMIAL_EXPAND(STRUCT(features), degree) creates x, x^2, x^3, etc.
# * ML.BUCKETIZE(f, split_points) where split_points is an array
# +
# %%bigquery
-- BQML chooses the wrong gradient descent strategy here. It will get fixed in (b/141429990)
-- But for now, as a workaround, explicitly specify optimize_strategy='BATCH_GRADIENT_DESCENT'
CREATE OR REPLACE MODEL serverlessml.model7_geo
TRANSFORM(
fare_amount
, SQRT( (pickuplon-dropofflon)*(pickuplon-dropofflon) + (pickuplat-dropofflat)*(pickuplat-dropofflat) ) AS euclidean
, ML.FEATURE_CROSS(STRUCT(CAST(EXTRACT(DAYOFWEEK FROM pickup_datetime) AS STRING) AS dayofweek,
CAST(EXTRACT(HOUR FROM pickup_datetime) AS STRING) AS hourofday), 2) AS day_hr
, CONCAT(
ML.BUCKETIZE(pickuplon, GENERATE_ARRAY(-78, -70, 0.01)),
ML.BUCKETIZE(pickuplat, GENERATE_ARRAY(37, 45, 0.01)),
ML.BUCKETIZE(dropofflon, GENERATE_ARRAY(-78, -70, 0.01)),
ML.BUCKETIZE(dropofflat, GENERATE_ARRAY(37, 45, 0.01))
) AS pickup_and_dropoff
)
OPTIONS(input_label_cols=['fare_amount'], model_type='linear_reg', l2_reg=0.1, optimize_strategy='BATCH_GRADIENT_DESCENT')
AS
SELECT * FROM serverlessml.feateng_training_data
# -
# %%bigquery
SELECT SQRT(mean_squared_error) AS rmse FROM ML.EVALUATE(MODEL serverlessml.model7_geo)
# Yippee! We're now below our target of 6 dollars in RMSE.
# ## DNN
#
# You could, of course, train a more sophisticated model. Change "linear_reg" above to "dnn_regressor" and see if it improves things.
#
# __Note: This takes 20 - 25 minutes to run.__
# +
# %%bigquery
-- This is alpha and may not work for you.
CREATE OR REPLACE MODEL serverlessml.model8_dnn
TRANSFORM(
fare_amount
, SQRT( (pickuplon-dropofflon)*(pickuplon-dropofflon) + (pickuplat-dropofflat)*(pickuplat-dropofflat) ) AS euclidean
, CONCAT(CAST(EXTRACT(DAYOFWEEK FROM pickup_datetime) AS STRING),
CAST(EXTRACT(HOUR FROM pickup_datetime) AS STRING)) AS day_hr
, CONCAT(
ML.BUCKETIZE(pickuplon, GENERATE_ARRAY(-78, -70, 0.01)),
ML.BUCKETIZE(pickuplat, GENERATE_ARRAY(37, 45, 0.01)),
ML.BUCKETIZE(dropofflon, GENERATE_ARRAY(-78, -70, 0.01)),
ML.BUCKETIZE(dropofflat, GENERATE_ARRAY(37, 45, 0.01))
) AS pickup_and_dropoff
)
-- at the time of writing, l2_reg wasn't supported yet.
OPTIONS(input_label_cols=['fare_amount'], model_type='dnn_regressor', hidden_units=[32, 8])
AS
SELECT * FROM serverlessml.feateng_training_data
# -
# %%bigquery
SELECT SQRT(mean_squared_error) AS rmse FROM ML.EVALUATE(MODEL serverlessml.model8_dnn)
# We really need the L2 reg (recall that we got 4.77 without the feateng). It's time to do [Feature Engineering in Keras](../../06_feateng_keras/labs/taxifare_fc.ipynb).
# Copyright 2019 Google Inc.
# Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
| quests/serverlessml/05_feateng/solution/feateng_bqml.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # An Introduction to SageMaker ObjectToVec model for sequence-sequence embedding
#
# ## Table of contents
#
# 1. [Background](#Background)
# 1. [Download datasets](#Download-datasets)
# 1. [Preprocessing](#Preprocessing)
# 1. [Model training and inference](#Model-training-and-inference)
# 1. [Transfer learning with object2vec](#Transfer-learning)
# 1. [How to enable the optimal training result](#How-to-enable-the-optimal-training-result)
# 1. [Hyperparameter Tuning (Advanced)](#Hyperparameter-Tuning-(Advanced))
# # Background
#
# *Object2Vec* is a highly customizable multi-purpose algorithm that can learn embeddings of pairs of objects. The embeddings are learned in a way that it preserves their pairwise **similarities**
# - **Similarity** is user-defined: users need to provide the algorithm with pairs of objects that they define as similar (1) or dissimilar (0); alternatively, the users can define similarity in a continuous sense (provide a real-valued similarity score for reach object pair)
# - The learned embeddings can be used to compute nearest neighbors of objects, as well as to visualize natural clusters of related objects in the embedding space. In addition, the embeddings can also be used as features of the corresponding objects in downstream supervised tasks such as classification or regression
# ### Using Object2Vec to Encode Sentences into Fixed Length Embeddings
# In this notebook, we will demonstrate how to train *Object2Vec* to encode sequences of varying length into fixed length embeddings.
#
# As a specific example, we will represent each sentence as a sequence of integers, and we will show how to learn an encoder to embed these sentences into fixed-length vectors. To this end, we need pairs of sentences with labels that indicate their similarity. The Stanford Natural Language Inference data set (https://nlp.stanford.edu/projects/snli/), which consists
# of pairs of sentences labeled as "entailment", "neutral" or "contradiction", comes close to our requirements; we will pick this data set as our training dataset in this notebook example.
#
# Once the model is trained on this data,
# the trained encoders can be used to convert any new English sentences into fixed length embeddings. We will measure the quality of learned sentence embeddings on new sentences, by computing similarity of sentence pairs in the embedding space from the STS'16 dataset (http://alt.qcri.org/semeval2016/task1/), and evaluating against human-labeled ground-truth ratings.
# <img style="float:middle" src="image_snli.png" width="480">
# ### Before running the notebook
# - Please use a Python 3 kernel for the notebook
# - Please make sure you have `jsonlines` and `nltk` packages installed
# ##### (If you haven't done it) install jsonlines and nltk
# !pip install -U nltk
# !pip install jsonlines
# ## Download datasets
# Please be aware of the following requirements about acknowledgment, copyright and availability, cited from the [dataset description page](https://nlp.stanford.edu/projects/snli/).
# > The Stanford Natural Language Inference Corpus by The Stanford NLP Group is licensed under a Creative Commons Attribution-ShareAlike 4.0 International License.
# Based on a work at http://shannon.cs.illinois.edu/DenotationGraph
import os
import requests
import io
import numpy as np
from zipfile import ZipFile
from datetime import datetime
# +
SNLI_PATH = "snli_1.0"
STS_PATH = "sts2016-english-with-gs-v1.0"
if not os.path.exists(SNLI_PATH):
url_address = "https://nlp.stanford.edu/projects/snli/snli_1.0.zip"
request = requests.get(url_address)
zfile = ZipFile(io.BytesIO(request.content))
zfile.extractall()
zfile.close()
if not os.path.exists(STS_PATH):
url_address = (
"http://alt.qcri.org/semeval2016/task1/data/uploads/sts2016-english-with-gs-v1.0.zip"
)
request = requests.get(url_address)
zfile = ZipFile(io.BytesIO(request.content))
zfile.extractall()
# -
# ## Preprocessing
import boto3
import sys, os
import jsonlines
import json
from collections import Counter
from itertools import chain, islice
from nltk.tokenize import TreebankWordTokenizer
# +
# constants
BOS_SYMBOL = "<s>"
EOS_SYMBOL = "</s>"
UNK_SYMBOL = "<unk>"
PAD_SYMBOL = "<pad>"
PAD_ID = 0
TOKEN_SEPARATOR = " "
VOCAB_SYMBOLS = [PAD_SYMBOL, UNK_SYMBOL, BOS_SYMBOL, EOS_SYMBOL]
LABEL_DICT = {"entailment": 0, "neutral": 1, "contradiction": 2}
# +
#### Utility functions
def read_jsonline(fname):
"""
Reads jsonline files and returns iterator
"""
with jsonlines.open(fname) as reader:
for line in reader:
yield line
def sentence_to_integers(sentence, tokenizer, word_dict):
"""
Converts a string of tokens to a list of integers
TODO: Better handling of the case
where token is not in word_dict
"""
return [word_dict[token] for token in get_tokens(sentence, tokenizer) if token in word_dict]
def get_tokens(line, tokenizer):
"""
Yields tokens from input string.
:param line: Input string.
:return: Iterator over tokens.
"""
for token in tokenizer.tokenize(line):
if len(token) > 0:
yield token
def get_tokens_from_snli(input_dict, tokenizer):
iter_list = list()
for sentence_key in ["sentence1", "sentence2"]:
sentence = input_dict[sentence_key]
iter_list.append(get_tokens(sentence, tokenizer))
return chain(iter_list[0], iter_list[1])
def get_tokens_from_sts(input_sentence_pair, tokenizer):
iter_list = list()
for s in input_sentence_pair:
iter_list.append(get_tokens(s, tokenizer))
return chain(iter_list[0], iter_list[1])
def resolve_snli_label(raw_label):
"""
Converts raw label to integer
"""
return LABEL_DICT[raw_label]
# -
# #### Functions to build vocabulary from SNLI corpus
def build_vocab(
data_iter, dataname="snli", num_words=50000, min_count=1, use_reserved_symbols=True, sort=True
):
"""
Creates a vocabulary mapping from words to ids. Increasing integer ids are assigned by word frequency,
using lexical sorting as a tie breaker. The only exception to this are special symbols such as the padding symbol
(PAD).
:param data_iter: Sequence of sentences containing whitespace delimited tokens.
:param num_words: Maximum number of words in the vocabulary.
:param min_count: Minimum occurrences of words to be included in the vocabulary.
:return: word-to-id mapping.
"""
vocab_symbols_set = set(VOCAB_SYMBOLS)
tokenizer = TreebankWordTokenizer()
if dataname == "snli":
raw_vocab = Counter(
token
for line in data_iter
for token in get_tokens_from_snli(line, tokenizer)
if token not in vocab_symbols_set
)
elif dataname == "sts":
raw_vocab = Counter(
token
for line in data_iter
for token in get_tokens_from_sts(line, tokenizer)
if token not in vocab_symbols_set
)
else:
raise NameError(f"Data name {dataname} is not recognized!")
print("Initial vocabulary: {} types".format(len(raw_vocab)))
# For words with the same count, they will be ordered reverse alphabetically.
# Not an issue since we only care for consistency
pruned_vocab = sorted(((c, w) for w, c in raw_vocab.items() if c >= min_count), reverse=True)
print("Pruned vocabulary: {} types (min frequency {})".format(len(pruned_vocab), min_count))
# truncate the vocabulary to fit size num_words (only includes the most frequent ones)
vocab = islice((w for c, w in pruned_vocab), num_words)
if sort:
# sort the vocabulary alphabetically
vocab = sorted(vocab)
if use_reserved_symbols:
vocab = chain(VOCAB_SYMBOLS, vocab)
word_to_id = {word: idx for idx, word in enumerate(vocab)}
print("Final vocabulary: {} types".format(len(word_to_id)))
if use_reserved_symbols:
# Important: pad symbol becomes index 0
assert word_to_id[PAD_SYMBOL] == PAD_ID
return word_to_id
# #### Functions to convert SNLI data to pairs of sequences of integers
def convert_snli_to_integers(data_iter, word_to_id, dirname=SNLI_PATH, fname_suffix=""):
"""
Go through snli jsonline file line by line and convert sentences to list of integers
- convert entailments to labels
"""
fname = "snli-integer-" + fname_suffix + ".jsonl"
path = os.path.join(dirname, fname)
tokenizer = TreebankWordTokenizer()
count = 0
max_seq_length = 0
with jsonlines.open(path, mode="w") as writer:
for in_dict in data_iter:
# in_dict = json.loads(line)
out_dict = dict()
rlabel = in_dict["gold_label"]
if rlabel in LABEL_DICT:
rsentence1 = in_dict["sentence1"]
rsentence2 = in_dict["sentence2"]
for idx, sentence in enumerate([rsentence1, rsentence2]):
# print(count, sentence)
s = sentence_to_integers(sentence, tokenizer, word_to_id)
out_dict[f"in{idx}"] = s
count += 1
max_seq_length = max(len(s), max_seq_length)
out_dict["label"] = resolve_snli_label(rlabel)
writer.write(out_dict)
else:
count += 1
print(f"There are in total {count} invalid labels")
print(f"The max length of converted sequence is {max_seq_length}")
# ### Generate vocabulary from SNLI data
# +
def make_snli_full_vocab(dirname=SNLI_PATH, force=True):
vocab_path = os.path.join(dirname, "snli-vocab.json")
if not os.path.exists(vocab_path) or force:
data_iter_list = list()
for fname_suffix in ["train", "test", "dev"]:
fname = "snli_1.0_" + fname_suffix + ".jsonl"
data_iter_list.append(read_jsonline(os.path.join(dirname, fname)))
data_iter = chain(data_iter_list[0], data_iter_list[1], data_iter_list[2])
with open(vocab_path, "w") as write_file:
word_to_id = build_vocab(
data_iter, num_words=50000, min_count=1, use_reserved_symbols=False, sort=True
)
json.dump(word_to_id, write_file)
make_snli_full_vocab(force=False)
# -
# ### Generate tokenized SNLI data as sequences of integers
# - We use the SNLI vocabulary as a lookup dictionary to convert SNLI sentence pairs into sequences of integers
# +
def make_snli_data(dirname=SNLI_PATH, vocab_file="snli-vocab.json", outfile_suffix="", force=True):
for fname_suffix in ["train", "test", "validation"]:
outpath = os.path.join(dirname, f"snli-integer-{fname_suffix}-{outfile_suffix}.jsonl")
if not os.path.exists(outpath) or force:
if fname_suffix == "validation":
inpath = os.path.join(dirname, f"snli_1.0_dev.jsonl")
else:
inpath = os.path.join(dirname, f"snli_1.0_{fname_suffix}.jsonl")
data_iter = read_jsonline(inpath)
vocab_path = os.path.join(dirname, vocab_file)
with open(vocab_path, "r") as f:
word_to_id = json.load(f)
convert_snli_to_integers(
data_iter,
word_to_id,
dirname=dirname,
fname_suffix=f"{fname_suffix}-{outfile_suffix}",
)
make_snli_data(force=False)
# -
# ## Model training and inference
# ### Training
# +
def get_vocab_size(vocab_path):
with open(vocab_path) as f:
word_to_id = json.load(f)
return len(word_to_id.keys())
vocab_path = os.path.join(SNLI_PATH, "snli-vocab.json")
vocab_size = get_vocab_size(vocab_path)
print("There are {} words in vocabulary {}".format(vocab_size, vocab_path))
# -
# For the runs in this notebook, we will use the Hierarchical CNN architecture to encode each of the sentences into fixed length embeddings. Some of the other hyperparameters are shown below.
## Define hyperparameters and define S3 input path
DEFAULT_HP = {
"enc_dim": 4096,
"mlp_dim": 512,
"mlp_activation": "linear",
"mlp_layers": 2,
"output_layer": "softmax",
"optimizer": "adam",
"learning_rate": 0.0004,
"mini_batch_size": 32,
"epochs": 20,
"bucket_width": 0,
"early_stopping_tolerance": 0.01,
"early_stopping_patience": 3,
"dropout": 0,
"weight_decay": 0,
"enc0_max_seq_len": 82,
"enc1_max_seq_len": 82,
"enc0_network": "hcnn",
"enc1_network": "enc0",
"enc0_token_embedding_dim": 300,
"enc0_layers": "auto",
"enc0_cnn_filter_width": 3,
"enc1_token_embedding_dim": 300,
"enc1_layers": "auto",
"enc1_cnn_filter_width": 3,
"enc0_vocab_file": "",
"enc1_vocab_file": "",
"enc0_vocab_size": vocab_size,
"enc1_vocab_size": vocab_size,
"num_classes": 3,
"_num_gpus": "auto",
"_num_kv_servers": "auto",
"_kvstore": "device",
}
# ### Define input data channel and output path in S3
# + tags=["parameters"]
import sagemaker
bucket = sagemaker.Session().default_bucket()
# -
## Input data bucket and prefix
prefix = "object2vec/input/"
input_path = os.path.join("s3://", bucket, prefix)
print(f"Data path for training is {input_path}")
## Output path
output_prefix = "object2vec/output/"
output_bucket = bucket
output_path = os.path.join("s3://", output_bucket, output_prefix)
print(f"Trained model will be saved at {output_path}")
# ### Initialize Sagemaker estimator
# - Get IAM role ObjectToVec algorithm image
# +
import sagemaker
from sagemaker import get_execution_role
sess = sagemaker.Session()
role = get_execution_role()
print(role)
## Get docker image of ObjectToVec algorithm
from sagemaker.amazon.amazon_estimator import get_image_uri
container = get_image_uri(boto3.Session().region_name, "object2vec")
# +
import sagemaker
from sagemaker.session import s3_input
def set_training_environment(
bucket,
prefix,
base_hyperparameters=DEFAULT_HP,
is_quick_run=True,
is_pretrain=False,
use_all_vocab={},
):
input_channels = {}
s3_client = boto3.client("s3")
for split in ["train", "validation"]:
if is_pretrain:
fname_in = f"all_vocab_datasets/snli-integer-{split}-pretrain.jsonl"
fname_out = f"{split}/snli-integer-{split}-pretrain.jsonl"
else:
fname_in = os.path.join(SNLI_PATH, f"snli-integer-{split}-.jsonl")
fname_out = f"{split}/snli-integer-{split}.jsonl"
s3_client.upload_file(fname_in, bucket, os.path.join(prefix, fname_out))
input_channels[split] = s3_input(
input_path + fname_out,
distribution="ShardedByS3Key",
content_type="application/jsonlines",
)
print("Uploaded {} data to {}".format(split, input_path + fname_out))
hyperparameters = base_hyperparameters.copy()
if use_all_vocab:
hyperparameters["enc0_vocab_file"] = "all_vocab.json"
hyperparameters["enc1_vocab_file"] = "all_vocab.json"
hyperparameters["enc0_vocab_size"] = use_all_vocab["vocab_size"]
hyperparameters["enc1_vocab_size"] = use_all_vocab["vocab_size"]
if is_pretrain:
## set up auxliary channel
aux_path = os.path.join(prefix, "auxiliary")
# upload auxiliary files
assert os.path.exists("GloVe/glove.840B-trim.txt"), "Pretrained embedding does not exist!"
s3_client.upload_file(
"GloVe/glove.840B-trim.txt", bucket, os.path.join(aux_path, "glove.840B-trim.txt")
)
if use_all_vocab:
s3_client.upload_file(
"all_vocab_datasets/all_vocab.json",
bucket,
os.path.join(aux_path, "all_vocab.json"),
)
else:
s3_client.upload_file(
"snli_1.0/snli-vocab.json", bucket, os.path.join(aux_path, "snli-vocab.json")
)
input_channels["auxiliary"] = s3_input(
"s3://" + bucket + "/" + aux_path,
distribution="FullyReplicated",
content_type="application/json",
)
print(
"Uploaded auxiliary data for initializing with pretrain-embedding to {}".format(
aux_path
)
)
# add pretrained_embedding_file name to hyperparameters
for idx in [0, 1]:
hyperparameters[f"enc{idx}_pretrained_embedding_file"] = "glove.840B-trim.txt"
if is_quick_run:
hyperparameters["mini_batch_size"] = 8192
hyperparameters["enc_dim"] = 16
hyperparameters["epochs"] = 2
else:
hyperparameters["mini_batch_size"] = 256
hyperparameters["enc_dim"] = 8192
hyperparameters["epochs"] = 20
return hyperparameters, input_channels
# -
# ### Train without using pretrained embedding
# +
## get estimator
regressor = sagemaker.estimator.Estimator(
container,
role,
train_instance_count=1,
train_instance_type="ml.p2.xlarge",
output_path=output_path,
sagemaker_session=sess,
)
## set up training environment
"""
- To get good training result, set is_quick_run to False
- To test-run the algorithm quickly, set is_quick_run to True
"""
hyperparameters, input_channels = set_training_environment(
bucket, prefix, is_quick_run=True, is_pretrain=False, use_all_vocab={}
)
regressor.set_hyperparameters(**hyperparameters)
regressor.hyperparameters()
# -
regressor.fit(input_channels)
# ### Plot evaluation metrics for training job
#
# Evaluation metrics for the completed training job are available in CloudWatch. We can pull the cross entropy metric of the validation data set and plot it to see the performance of the model over time.
# +
# %matplotlib inline
from sagemaker.analytics import TrainingJobAnalytics
latest_job_name = regressor.latest_training_job.job_name
metric_name = "validation:cross_entropy"
metrics_dataframe = TrainingJobAnalytics(
training_job_name=latest_job_name, metric_names=[metric_name]
).dataframe()
plt = metrics_dataframe.plot(
kind="line", figsize=(12, 5), x="timestamp", y="value", style="b.", legend=False
)
plt.set_ylabel(metric_name);
# -
# ### Deploy trained algorithm and set input-output configuration for inference
# +
from sagemaker.predictor import json_serializer, json_deserializer
# deploy model and create endpoint and with customer-defined endpoint_name
predictor1 = regressor.deploy(initial_instance_count=1, instance_type="ml.m4.xlarge")
# -
# define encode-decode format for inference data
predictor1.serializer = json_serializer
predictor1.deserializer = json_deserializer
# ### Invoke endpoint and do inference with trained model
# - Suppose we deploy our trained model with the endpoint_name "seqseq-prelim-with-pretrain-3". Now we demonstrate how to do inference using our earlier model
def calc_prediction_accuracy(predictions, labels):
loss = 0
for idx, s_and_l in enumerate(zip(predictions["predictions"], labels)):
score, label = s_and_l
plabel = np.argmax(score["scores"])
loss += int(plabel != label["label"])
return 1 - loss / len(labels)
# ### Send mini-batches of SNLI test data to the endpoint and evaluate our model
# +
import math
import sagemaker
from sagemaker.predictor import json_serializer, json_deserializer
# load SNLI test data
snli_test_path = os.path.join(SNLI_PATH, "snli-integer-test-.jsonl")
test_data_content = list()
test_label = list()
for line in read_jsonline(snli_test_path):
test_data_content.append({"in0": line["in0"], "in1": line["in1"]})
test_label.append({"label": line["label"]})
print("Evaluating test results on SNLI without pre-trained embedding...")
batch_size = 100
n_test = len(test_label)
n_batches = math.ceil(n_test / float(batch_size))
start = 0
agg_acc = 0
for idx in range(n_batches):
if idx % 10 == 0:
print(f"Evaluating the {idx+1}-th batch")
end = (start + batch_size) if (start + batch_size) <= n_test else n_test
payload = {"instances": test_data_content[start:end]}
acc = calc_prediction_accuracy(predictor1.predict(payload), test_label[start:end])
agg_acc += acc * (end - start + 1)
start = end
print(f"The test accuracy is {agg_acc/n_test}")
# -
# # Transfer learning
# - We evaluate the trained model directly on STS16 **question-question** task
# - See SemEval-2016 Task 1 paper (http://www.aclweb.org/anthology/S16-1081) for an explanation of the evaluation method and benchmarking results
# The cells below provide details on how to combine vocabulary for STS and SNLI,and how to get glove pretrained embedding
# ### Functions to generate STS evaluation set (from sts-2016-test set)
# +
def loadSTSFile(fpath=STS_PATH, datasets=["question-question"]):
data = {}
for dataset in datasets:
sent1 = []
sent2 = []
for line in (
io.open(fpath + f"/STS2016.input.{dataset}.txt", encoding="utf8").read().splitlines()
):
splitted = line.split("\t")
sent1.append(splitted[0])
sent2.append(splitted[1])
raw_scores = np.array(
[
x
for x in io.open(fpath + f"/STS2016.gs.{dataset}.txt", encoding="utf8")
.read()
.splitlines()
]
)
not_empty_idx = raw_scores != ""
gs_scores = [float(x) for x in raw_scores[not_empty_idx]]
sent1 = np.array(sent1)[not_empty_idx]
sent2 = np.array(sent2)[not_empty_idx]
data[dataset] = (sent1, sent2, gs_scores)
return data
def get_sts_data_iterator(fpath=STS_PATH, datasets=["question-question"]):
data = loadSTSFile(fpath, datasets)
for dataset in datasets:
sent1, sent2, _ = data[dataset]
for s1, s2 in zip(sent1, sent2):
yield [s1, s2]
## preprocessing unit for STS test data
def convert_single_sts_to_integers(s1, s2, gs_label, tokenizer, word_dict):
converted = []
for s in [s1, s2]:
converted.append(sentence_to_integers(s, tokenizer, word_dict))
converted.append(gs_label)
return converted
def convert_sts_to_integers(sent1, sent2, gs_labels, tokenizer, word_dict):
for s1, s2, gs in zip(sent1, sent2, gs_labels):
yield convert_single_sts_to_integers(s1, s2, gs, tokenizer, word_dict)
def make_sts_data(
fpath=STS_PATH,
vocab_path_prefix=SNLI_PATH,
vocab_name="snli-vocab.json",
dataset="question-question",
):
"""
prepare test data; example: test_data['left'] = [{'in0':[1,2,3]}, {'in0':[2,10]}, ...]
"""
test_data = {"left": [], "right": []}
test_label = list()
tokenizer = TreebankWordTokenizer()
vocab_path = os.path.join(vocab_path_prefix, vocab_name)
with open(vocab_path) as f:
word_dict = json.load(f)
data = loadSTSFile(fpath=fpath, datasets=[dataset])
for s1, s2, gs in convert_sts_to_integers(*data[dataset], tokenizer, word_dict):
test_data["left"].append({"in1": s1})
test_data["right"].append({"in1": s2})
test_label.append(gs)
return test_data, test_label
# -
# Note, in `make_sts_data`, we pass both inputs (s1 and s2 to a single encoder; in this case, we pass them to 'in1'). This makes sure that both inputs are mapped by the same encoding function (we empirically found that this is crucial to achieve competitive embedding performance)
# ### Build vocabulary using STS corpus
# +
def make_sts_full_vocab(dirname=STS_PATH, datasets=["question-question"], force=True):
vocab_path = os.path.join(dirname, "sts-vocab.json")
if not os.path.exists(vocab_path) or force:
data_iter = get_sts_data_iterator(dirname, datasets)
with open(vocab_path, "w") as write_file:
word_to_id = build_vocab(
data_iter,
dataname="sts",
num_words=50000,
min_count=1,
use_reserved_symbols=False,
sort=True,
)
json.dump(word_to_id, write_file)
make_sts_full_vocab(force=False)
# -
# Define functions for embedding evaluation on STS16 question-question task
# +
from scipy.stats import pearsonr, spearmanr
import math
def wrap_sts_test_data_for_eval(
fpath=STS_PATH, vocab_path_prefix=".", vocab_name="all_vocab.json", dataset="question-question"
):
"""
Prepare data for evaluation
"""
test_data, test_label = make_sts_data(fpath, vocab_path_prefix, vocab_name, dataset)
input1 = {"instances": test_data["left"]}
input2 = {"instances": test_data["right"]}
return [input1, input2, test_label]
def get_cosine_similarity(vec1, vec2):
assert len(vec1) == len(vec2), "Vector dimension mismatch!"
norm1 = 0
norm2 = 0
inner_product = 0
for v1, v2 in zip(vec1, vec2):
norm1 += v1 ** 2
norm2 += v2 ** 2
inner_product += v1 * v2
return inner_product / math.sqrt(norm1 * norm2)
def eval_corr(predictor, eval_data):
"""
input:
param: predictor: Sagemaker deployed model
eval_data: a list of [input1, inpu2, gs_scores]
Evaluate pearson and spearman correlation between algorithm's embedding and gold standard
"""
sys_scores = []
input1, input2, gs_scores = (
eval_data[0],
eval_data[1],
eval_data[2],
) # get this from make_sts_data
embeddings = []
for data in [input1, input2]:
prediction = predictor.predict(data)
embeddings.append(prediction["predictions"])
for emb_pair in zip(embeddings[0], embeddings[1]):
emb1 = emb_pair[0]["embeddings"]
emb2 = emb_pair[1]["embeddings"]
sys_scores.append(get_cosine_similarity(emb1, emb2)) # TODO: implement this
results = {
"pearson": pearsonr(sys_scores, gs_scores),
"spearman": spearmanr(sys_scores, gs_scores),
"nsamples": len(sys_scores),
}
return results
# -
# ### Check overlap between SNLI and STS vocabulary
# +
snli_vocab_path = os.path.join(SNLI_PATH, "snli-vocab.json")
sts_vocab_path = os.path.join(STS_PATH, "sts-vocab.json")
with open(sts_vocab_path) as f:
sts_v = json.load(f)
with open(snli_vocab_path) as f:
snli_v = json.load(f)
sts_v_set = set(sts_v.keys())
snli_v_set = set(snli_v.keys())
print(len(sts_v_set))
not_captured = sts_v_set.difference(snli_v_set)
print(not_captured)
print(f"\nThe number of words in STS not included in SNLI is {len(not_captured)}")
print(
f"\nThis is {round(float(len(not_captured)/len(sts_v_set)), 2)} percent of the total STS vocabulary"
)
# -
# #### Since the percentage of vocabulary in STS not covered by SNLI is pretty large, we are going to include the uncovered words into our vocabulary and use the *GloVe* pretrained embedding to initialize our network.
#
# ##### Intuitive reasoning for why this works
#
# * Our algorithm will not have seen the ***uncovered words*** during training
# * If we directly use integer representation of words during training, the unseen words will have zero correlation with words seen.
# - This means the model cannot embed the unseen words in a manner that takes advantage of its training knowledge
# * However, if we use pre-trained word embedding, then we expect that some of the unseen words will be close to the words that the algorithm has seen in the embedding space
# +
def combine_vocabulary(vocab_paths, new_vocab_path):
wd_count = 0
all_vocab = set()
new_vocab = {}
for vocab_path in vocab_paths:
with open(vocab_path) as f:
vocab = json.load(f)
all_vocab = all_vocab.union(vocab.keys())
for idx, wd in enumerate(all_vocab):
new_vocab[wd] = idx
print(f"The new vocabulary size is {idx+1}")
with open(new_vocab_path, "w") as f:
json.dump(new_vocab, f)
vocab_paths = [snli_vocab_path, sts_vocab_path]
new_vocab_path = "all_vocab.json"
combine_vocabulary(vocab_paths, new_vocab_path)
# -
# ## Get pre-trained GloVe word embedding and upload it to S3
#
# - Our notebook storage is not enough to host the *GloVe* file. Fortunately, we have extra space in the `/tmp` folder that we can utilize: https://docs.aws.amazon.com/sagemaker/latest/dg/howitworks-create-ws.html
# - You may use the bash script below to download and unzip *GloVe* in the `/tmp` folder and remove it after use
# + language="bash"
# # download glove file from website
# mkdir /tmp/GloVe
# curl -Lo /tmp/GloVe/glove.840B.zip http://nlp.stanford.edu/data/glove.840B.300d.zip
# unzip /tmp/GloVe/glove.840B.zip -d /tmp/GloVe/
# rm /tmp/GloVe/glove.840B.zip
# -
# We next trim the original *GloVe* embedding file so that it just covers our combined vocabulary, and then we save the trimmed glove file in the newly created *GloVe* directory
# !mkdir GloVe
# +
import json
# credit: This preprocessing function is modified from the w2v preprocessing script in Facebook infersent codebase
# Infersent code license can be found at: https://github.com/facebookresearch/InferSent/blob/master/LICENSE
def trim_w2v(in_path, out_path, word_dict):
# create word_vec with w2v vectors
lines = []
with open(out_path, "w") as outfile:
with open(in_path) as f:
for line in f:
word, vec = line.split(" ", 1)
if word in word_dict:
lines.append(line)
print("Found %s(/%s) words with w2v vectors" % (len(lines), len(word_dict)))
outfile.writelines(lines)
in_path = "/tmp/GloVe/glove.840B.300d.txt"
out_path = "GloVe/glove.840B-trim.txt"
with open("all_vocab.json") as f:
word_dict = json.load(f)
trim_w2v(in_path, out_path, word_dict)
# -
# remember to remove the original GloVe embedding folder since it takes up a lot of space
# !rm -r /tmp/GloVe/
# ## Reprocess training data (SNLI) with the combined vocabulary
# Create a new directory called `all_vocab_datasets`, and copy snli raw json files and all_vocab file to it
# + language="bash"
#
# mkdir all_vocab_datasets
#
# for SPLIT in train dev test
# do
# cp snli_1.0/snli_1.0_${SPLIT}.jsonl all_vocab_datasets/
# done
#
# cp all_vocab.json all_vocab_datasets/
# -
# Convert snli data to integers using the all_vocab file
make_snli_data(
dirname="all_vocab_datasets",
vocab_file="all_vocab.json",
outfile_suffix="pretrain",
force=False,
)
# Let's see the size of this new vocabulary
all_vocab_path = "all_vocab.json"
all_vocab_size = get_vocab_size(all_vocab_path)
print("There are {} words in vocabulary {}".format(all_vocab_size, all_vocab_path))
# ### Reset training environment
# Note that when we combine the vocabulary of our training and test data, we should not fine-tune the GloVE embeddings, but instead, keep them fixed. Otherwise, it amounts to a bit of cheating -- training on test data! Thankfully, our hyper-parameter `enc0/1_freeze_pretrained_embedding` is set to `True` by default. Note that in the earlier training where we did not use pretrained embeddings, this parameter is inconsequential.
# +
hyperparameters_2, input_channels_2 = set_training_environment(
bucket,
prefix,
is_quick_run=True,
is_pretrain=True,
use_all_vocab={"vocab_size": all_vocab_size},
)
# attach a new regressor to the old one using the previous training job endpoint
# (this will also retrieve the log of the previous training job)
training_job_name = regressor.latest_training_job.name
new_regressor = regressor.attach(training_job_name, sagemaker_session=sess)
new_regressor.set_hyperparameters(**hyperparameters_2)
# -
# fit the new regressor using the new data (with pretrained embedding)
new_regressor.fit(input_channels_2)
# ### Deploy and test the new model
predictor_2 = new_regressor.deploy(initial_instance_count=1, instance_type="ml.m4.xlarge")
predictor_2.serializer = json_serializer
predictor_2.deserializer = json_deserializer
# We first check the test error on SNLI after adding pretrained embedding
# +
# load SNLI test data
snli_test_path = os.path.join("all_vocab_datasets", "snli-integer-test-pretrain.jsonl")
test_data_content = list()
test_label = list()
for line in read_jsonline(snli_test_path):
test_data_content.append({"in0": line["in0"], "in1": line["in1"]})
test_label.append({"label": line["label"]})
print("Evaluating test results on SNLI with pre-trained embedding...")
batch_size = 100
n_test = len(test_label)
n_batches = math.ceil(n_test / float(batch_size))
start = 0
agg_acc = 0
for idx in range(n_batches):
if idx % 10 == 0:
print(f"Evaluating the {idx+1}-th batch")
end = (start + batch_size) if (start + batch_size) <= n_test else n_test
payload = {"instances": test_data_content[start:end]}
acc = calc_prediction_accuracy(predictor_2.predict(payload), test_label[start:end])
agg_acc += acc * (end - start + 1)
start = end
print(f"The test accuracy is {agg_acc/n_test}")
# -
#
# We next test the zero-shot transfer learning performance of our trained model on STS task
# +
eval_data_qq = wrap_sts_test_data_for_eval(
fpath=STS_PATH,
vocab_path_prefix="all_vocab_datasets",
vocab_name="all_vocab.json",
dataset="question-question",
)
results = eval_corr(predictor_2, eval_data_qq)
pcorr = results["pearson"][0]
spcorr = results["spearman"][0]
print(f"The Pearson correlation to gold standard labels is {pcorr}")
print(f"The Spearman correlation to gold standard labels is {spcorr}")
# -
## clean up
sess.delete_endpoint(predictor1.endpoint)
sess.delete_endpoint(predictor_2.endpoint)
# # How to enable the optimal training result
# So far we have been training the algorithm with `is_quick_run` set to `True` (in `set_training_envirnoment` function); this is because we want to minimize the time for you to run through this notebook. If you want to yield the best performance of *Object2Vec* on the tasks above, we recommend setting `is_quick_run` to `False`. For example, with pretrained embedding used, we would re-run the code block under **Reset training environment** as the block below
# <span style="color:red">Run with caution</span>:
# This may take a few hours to complete depending on the machine instance you are using
# +
hyperparameters_2, input_channels_2 = set_training_environment(
bucket,
prefix,
is_quick_run=False, # modify is_quick_run flag here
is_pretrain=True,
use_all_vocab={"vocab_size": all_vocab_size},
)
training_job_name = regressor.latest_training_job.name
new_regressor = regressor.attach(training_job_name, sagemaker_session=sess)
new_regressor.set_hyperparameters(**hyperparameters_2)
# -
# Then we can train and deploy the model as before; similarly, without pretrained embedding, the code block under **Train without using pretrained embedding** can be changed to below to optimize training result
# <span style="color:red">Run with caution</span>:
# This may take a few hours to complete depending on the machine instance you are using
# +
hyperparameters, input_channels = set_training_environment(
bucket,
prefix,
is_quick_run=False, # modify is_quick_run flag here
is_pretrain=False,
use_all_vocab={},
)
regressor.set_hyperparameters(**hyperparameters)
regressor.hyperparameters()
# -
# ### Best training result
#
# With `is_quick_run = False` and without pretrained embedding, our algorithm's test accuracy on SNLI dataset is 78.5%; with pretrained GloVe embedding, we see an improved test accuracy on SNLI dataset to 81.9% ! On STS data, you should expect the Pearson correlation to be around 0.61.
#
# In addition to the training demonstrated in this notebook, we have also done benchmarking experiments on evaluated on both SNLI and STS data, with different hyperparameter configurations, which we include below.
#
# In both charts, we compare against Facebook's Infersent algorithm (https://research.fb.com/downloads/infersent/). The chart on the left shows the additional experiment result on SNLI (using CNN or RNN encoders). The chart on the right shows the best experiment result of Object2Vec on STS.
# <img style="float:left" src="o2v-exp-snli.png" width="430">
# <img style="float:middle" src="o2v-exp-sts.png" width="430">
# # Hyperparameter Tuning (Advanced)
# with Hyperparameter Optimization (HPO) service in Sagemaker
# To yield optimal performance out of any machine learning algorithm often requires a lot of effort on parameter tuning.
# In this notebook demo, we have hidden the hard work of finding a combination of good parameters for the algorithm on SNLI data (again, the optimal parameters are only defined by running `set_training_environment` method with `is_quick_run=False`).
#
# If you are keen to explore how to tune HP on your own, you may find the code blocks below helpful.
# To find the best HP combinations for our task, we can do parameter tuning by launching HPO jobs either from
# - As a simple example, we demonstrate how to find the best `enc_dim` parameter using HPO service here
# +
s3_uri_path = {}
for split in ["train", "validation"]:
s3_uri_path[split] = input_path + f"{split}/snli-integer-{split}.jsonl"
# -
# On a high level, a HPO tuning job is nothing but a collection of multiple training jobs with different HP setups; Sagemaker HPO service compares the performance of different training jobs according to the **HPO tuning metric**, which is specified in the `tuning_job_config`.
#
# - More info on how to manually launch hpo tuning jobs can be found here:
# https://docs.aws.amazon.com/sagemaker/latest/dg/automatic-model-tuning-ex-tuning-job.html
tuning_job_config = {
"ParameterRanges": {
"CategoricalParameterRanges": [],
"ContinuousParameterRanges": [],
"IntegerParameterRanges": [{"MaxValue": "1024", "MinValue": "16", "Name": "enc_dim"}],
},
"ResourceLimits": {"MaxNumberOfTrainingJobs": 3, "MaxParallelTrainingJobs": 3},
"Strategy": "Bayesian",
"HyperParameterTuningJobObjective": {"MetricName": "validation:accuracy", "Type": "Maximize"},
}
# The tuning metric `MetricName` we use here is called `validation:accuracy`, together with `Type` set to `Maximize`, since we are trying to maximize accuracy here (in case you want to minimize mean squared error, you can switch the tuning objective accordingly to `validation:mean_squared_error` and `Minimize`).
# The syntax for defining the configuration of an individual training job in a HPO job is as below
training_job_definition = {
"AlgorithmSpecification": {"TrainingImage": container, "TrainingInputMode": "File"},
"InputDataConfig": [
{
"ChannelName": "train",
"CompressionType": "None",
"ContentType": "application/jsonlines",
"DataSource": {
"S3DataSource": {
"S3DataDistributionType": "FullyReplicated",
"S3DataType": "S3Prefix",
"S3Uri": s3_uri_path["train"],
}
},
},
{
"ChannelName": "validation",
"CompressionType": "None",
"ContentType": "application/jsonlines",
"DataSource": {
"S3DataSource": {
"S3DataDistributionType": "FullyReplicated",
"S3DataType": "S3Prefix",
"S3Uri": s3_uri_path["validation"],
}
},
},
],
"OutputDataConfig": {"S3OutputPath": output_path},
"ResourceConfig": {"InstanceCount": 1, "InstanceType": "ml.p3.8xlarge", "VolumeSizeInGB": 20},
"RoleArn": role,
"StaticHyperParameters": {
#'enc_dim': "16", # do not include enc_dim here as static HP since we are tuning it
"learning_rate": "0.0004",
"mlp_dim": "512",
"mlp_activation": "linear",
"mlp_layers": "2",
"output_layer": "softmax",
"optimizer": "adam",
"mini_batch_size": "8192",
"epochs": "2",
"bucket_width": "0",
"early_stopping_tolerance": "0.01",
"early_stopping_patience": "3",
"dropout": "0",
"weight_decay": "0",
"enc0_max_seq_len": "82",
"enc1_max_seq_len": "82",
"enc0_network": "hcnn",
"enc1_network": "enc0",
"enc0_token_embedding_dim": "300",
"enc0_layers": "auto",
"enc0_cnn_filter_width": "3",
"enc1_token_embedding_dim": "300",
"enc1_layers": "auto",
"enc1_cnn_filter_width": "3",
"enc0_vocab_file": "",
"enc1_vocab_file": "",
"enc0_vocab_size": str(vocab_size),
"enc1_vocab_size": str(vocab_size),
"num_classes": "3",
"_num_gpus": "auto",
"_num_kv_servers": "auto",
"_kvstore": "device",
},
"StoppingCondition": {"MaxRuntimeInSeconds": 43200},
}
# +
import boto3
sm_client = boto3.Session().client("sagemaker")
# -
# ### Disclaimer
#
# Running HPO tuning jobs means dispatching multiple training jobs with different HP setups; this could potentially incur a significant cost on your AWS account if you use the HP combinations that takes long hours to train.
#
tuning_job_name = "hpo-o2v-test-{}".format(datetime.now().strftime("%d%m%Y-%H-%M-%S"))
response = sm_client.create_hyper_parameter_tuning_job(
HyperParameterTuningJobName=tuning_job_name,
HyperParameterTuningJobConfig=tuning_job_config,
TrainingJobDefinition=training_job_definition,
)
# You can then view and track the hyperparameter tuning jobs you launched on the sagemaker console (using the same account that you used to create the sagemaker client to launch these jobs)
| introduction_to_amazon_algorithms/object2vec_sentence_similarity/object2vec_sentence_similarity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!--HEADER-->
# *[Header for the notebooks in the jupyterbookmaker module](https://github.com/rmsrosa/jupyterbookmaker)*
# <!--NAVIGATOR-->
#
# <a href="https://colab.research.google.com/github/rmsrosa/jupyterbookmaker/blob/master/tests/notebooks/01.03-Header_creation.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open and Execute in Google Colaboratory"></a>
#
# <a href="https://mybinder.org/v2/gh/rmsrosa/jupyterbookmaker/master?filepath=tests/notebooks/01.03-Header_creation.ipynb"><img align="left" src="https://mybinder.org/badge.svg" alt="Open in binder" title="Open and Execute in Binder"></a>
#
#
# [<- Creating the Table of Contents](01.02-Toc_creation.ipynb) | [Front page](00.00-Front_Page.ipynb) | [References](BA.00-References.ipynb) | [Creating the navigators ->](01.04-Navigator_creation.ipynb)
#
# ---
#
# # Creating the headers
#
# The method to create, or update, the headers is called `add_headers()` and its definition starts with
# ```python
# def add_headers(header, app_to_notes_path='.'):
# ...
# ```
#
# The argument `add_headers` is required and is a string with the header you want to be displayed on top of each notebook.
#
# The `app_to_notes_path` is a non-required argument with the name of the folder in which both the `toc_nb_name` file and the collection of all notebooks to be listed in the table of contents are expected to be. It should be either an absolute path or a path relative from where the code is being ran. If `app_to_notes_path` is not given, it is assumed to be the current directory.
# <!--NAVIGATOR-->
#
# ---
# [<- Creating the Table of Contents](01.02-Toc_creation.ipynb) | [Front page](00.00-Front_Page.ipynb) | [References](BA.00-References.ipynb) | [Creating the navigators ->](01.04-Navigator_creation.ipynb)
#
# <a href="https://colab.research.google.com/github/rmsrosa/jupyterbookmaker/blob/master/tests/notebooks/01.03-Header_creation.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open and Execute in Google Colaboratory"></a>
#
# <a href="https://mybinder.org/v2/gh/rmsrosa/jupyterbookmaker/master?filepath=tests/notebooks/01.03-Header_creation.ipynb"><img align="left" src="https://mybinder.org/badge.svg" alt="Open in binder" title="Open and Execute in Binder"></a>
#
| tests/notebooks/01.03-Header_creation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import itertools
import codecs
import json
import re
import string
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
import hunspell
hu_spellchecker = hunspell.HunSpell('/home/rgomes/dictionaries/dictionaries/hu/index.dic', '/home/rgomes/dictionaries/dictionaries/hu/index.aff')
it_spellchecker = hunspell.HunSpell('/home/rgomes/dictionaries/dictionaries/it/index.dic', '/home/rgomes/dictionaries/dictionaries/it/index.aff')
# -
# ### Pre processing ###
# 1. Load the datasets
# 2. Format as post and save
# 3. Remove the enconding issues and stopwords
# 4. Analise the frequence of words
# 5. Save the results as bag of words
# +
# TODO check the relevance of language encoding
# see https://docs.python.org/2/library/codecs.html
# see https://www.terena.org/activities/multiling/ml-docs/iso-8859.html
data_hu = pd.read_json('../hungarian.txt')
file = open('../italian.txt', 'r')
data_it = []
for line in file:
data_it.append(json.loads(line))
data_it = pd.DataFrame(data_it)
# -
def replace_numbers(words):
new_words = []
for word in words:
new_word = re.sub(r'[0-9]', '', word)
if new_word != '':
new_words.append(new_word)
return new_words
# +
writer = pd.ExcelWriter('italian.xlsx')
#it_posts.to_excel(writer,'Italian Posts')
writer2 = pd.ExcelWriter('hungarian.xlsx')
#hu_posts.to_excel(writer2,'Hungarian Posts')
# -
# # Mark the words that contains accents not recognized by the spellchecker
# +
# ITALIAN
breakline = '\n===============================================================================\n'
# remove all breaklines
it_posts = data_it["text"].apply(lambda x: x.replace("\n\n\n\n", ""))
it_accented_words = []
for line in it_posts:
words = line.split()
words = replace_numbers(words)
for word in words:
# save the accented words not recognized
if re.search(r'[À-ž\'\`]', word) and not it_spellchecker.spell(word):
it_accented_words.append(word)
print(len(list(it_accented_words)))
print(it_accented_words)
# -
pd_italian_words = pd.DataFrame(it_accented_words)
pd_italian_words.to_csv('italian_ac_words_all.csv')
with open('../italian_ac_words_all.txt','w') as data_file:
for word in it_accented_words:
data_file.write(word + '\n')
with open('../italian_words_all.txt','w') as data_file_all:
for line in it_posts:
words = line.split()
words = replace_numbers(words)
for word in words:
data_file_all.write(word + '\n')
X = data_it["text"].apply(lambda x: x.split(" "))
X = X.values.tolist()
X = list(itertools.chain.from_iterable(X))
spellings = [[x, it_spellchecker.spell(x)] for x in X]
print('Not recognized', len(list(filter(lambda x: x[1] == False, spellings))))
print('Recognized', len(list(filter(lambda x: x[1] == True, spellings))))
# +
# save formatted as posts
breakline = '\n===============================================================================\n'
hu_posts = data_hu["text"]
hu_accented_words = []
for line in hu_posts:
words = line.split()
words = remove_punctuation(words)
words = replace_numbers(words)
for word in words:
if re.match(r'(\w+[À-ž]\w+)', word) and not hu_spellchecker.spell(word):
hu_accented_words.append(word)
print(len(set(hu_accented_words)))
#print(hu_accented_words)
# +
# save formatted as posts
breakline = '\n===============================================================================\n'
# remove all breaklines
it_posts = data_it["text"].apply(lambda x: x.replace("\n\n\n\n", ""))
# save hungarian separated by posts
it_posts_file = open('../italian_posts.txt','w')
for line in it_posts:
#print(line)
it_posts_file.write(line)
it_posts_file.write(breakline)
it_posts_file.close()
# save hungarian separated by posts
hu_posts = data_hu['text']
hu_posts_file = open('../hungarian_posts.txt','w')
for line in hu_posts:
#print(line)
hu_posts_file.write(line)
hu_posts_file.write(breakline)
hu_posts_file.close()
# -
print('%s hungarian posts' % len(data_hu))
data_hu.head()
print('%s italian posts' % len(data_it))
data_it.head(10)
def histogram(list):
d={}
for i in list:
if i in d:
d[i] += 1
else:
d[i] = 1
return d
def itokenize(iterator):
# split to every word
X = map(lambda x: x.split(" "), iterator)
return X
def clearAndTokenize(dataframe):
# replace all non characters letters like !,(,),.,@,[,],breakline'
X = map(lambda x: x.replace("\n", " "), dataframe)
X = map(lambda x: re.sub(r'[\W]', ' ', x), X)
# split to every word
X = map(lambda x: x.split(" "), X)
# transform to list
X = itertools.chain.from_iterable(X)
# remove all words that not contains latin accents
# see http://www.rexegg.com/regex-interesting-character-classes.html#obnoxious
#X = filter(lambda x: re.match(r'[À-ÖØ-öø-ÿ]', x) , X)
# remove the words with one letter, numbers and basic cases
X = filter(lambda i: not str.isdigit(i) and len(i) > 1 and i != 'és', X)
return list(X)
def plotHistogram(data):
h = histogram(X)
h = sorted(h.items(), key=lambda x: x[1], reverse=True)
h = map(lambda x: x[1], h)
# remove the words that appears only once
h = filter(lambda x: x > 1, h)
plt.plot(list(h))
plt.show()
# ## Hunspell ##
# 1. Load the spell checking tools
# 2. Save the not recognized for volunteer analysis
check = clearAndTokenize(data_hu['text'].values.tolist())
posts = data_hu['text'].values.tolist()
hu_list = list(posts)
print(hu_list[:1])
tokens = itokenize(hu_list[:1])
#print(list(tokens))
for word in list(tokens)[0]:
accented_word = re.match(r'(\w+[À-ž]\w+)', word)
if accented_word:
print(word)
print(hu_spellchecker.spell(word))
print(hu_spellchecker.suggest(word))
file = open('../hungarian_suggestions.json','w')
import json
json.dump(list(check), file, indent=1)
file.close()
# +
X = data_it['text']
plotHistogram(X)
print(len(X), 'italian accent')
#print(X[:1000])
# -
# ### Pos processing ###
# 1. Load the volunteer data
# 2. Analyse the distribuition of erros
# 3. Summ the patterns found
| tcc/notebooks/data_exploration_v1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import keras
keras.__version__
# # Overfitting and underfitting
#
# This notebook contains the code samples found in Chapter 3, Section 6 of [Deep Learning with Python](https://www.manning.com/books/deep-learning-with-python?a_aid=keras&a_bid=76564dff). Note that the original text features far more content, in particular further explanations and figures: in this notebook, you will only find source code and related comments.
#
# ----
#
#
# In all the examples we saw in the previous chapter -- movie review sentiment prediction, topic classification, and house price regression --
# we could notice that the performance of our model on the held-out validation data would always peak after a few epochs and would then start
# degrading, i.e. our model would quickly start to _overfit_ to the training data. Overfitting happens in every single machine learning
# problem. Learning how to deal with overfitting is essential to mastering machine learning.
#
# The fundamental issue in machine learning is the tension between optimization and generalization. "Optimization" refers to the process of
# adjusting a model to get the best performance possible on the training data (the "learning" in "machine learning"), while "generalization"
# refers to how well the trained model would perform on data it has never seen before. The goal of the game is to get good generalization, of
# course, but you do not control generalization; you can only adjust the model based on its training data.
#
# At the beginning of training, optimization and generalization are correlated: the lower your loss on training data, the lower your loss on
# test data. While this is happening, your model is said to be _under-fit_: there is still progress to be made; the network hasn't yet
# modeled all relevant patterns in the training data. But after a certain number of iterations on the training data, generalization stops
# improving, validation metrics stall then start degrading: the model is then starting to over-fit, i.e. is it starting to learn patterns
# that are specific to the training data but that are misleading or irrelevant when it comes to new data.
#
# To prevent a model from learning misleading or irrelevant patterns found in the training data, _the best solution is of course to get
# more training data_. A model trained on more data will naturally generalize better. When that is no longer possible, the next best solution
# is to modulate the quantity of information that your model is allowed to store, or to add constraints on what information it is allowed to
# store. If a network can only afford to memorize a small number of patterns, the optimization process will force it to focus on the most
# prominent patterns, which have a better chance of generalizing well.
#
# The processing of fighting overfitting in this way is called _regularization_. Let's review some of the most common regularization
# techniques, and let's apply them in practice to improve our movie classification model from the previous chapter.
# Note: in this notebook we will be using the IMDB test set as our validation set. It doesn't matter in this context.
#
# Let's prepare the data using the code from Chapter 3, Section 5:
# +
from keras.datasets import imdb
import numpy as np
# (train_data, train_labels), (test_data, test_labels) = imdb.load_data(num_words=10000)
import numpy as np
# save np.load
np_load_old = np.load
# modify the default parameters of np.load
np.load = lambda *a,**k: np_load_old(*a, allow_pickle=True, **k)
# call load_data with allow_pickle implicitly set to true
(train_data, train_labels), (test_data, test_labels) = imdb.load_data(num_words=10000)
# restore np.load for future normal usage
np.load = np_load_old
def vectorize_sequences(sequences, dimension=10000):
# Create an all-zero matrix of shape (len(sequences), dimension)
results = np.zeros((len(sequences), dimension))
for i, sequence in enumerate(sequences):
results[i, sequence] = 1. # set specific indices of results[i] to 1s
return results
# Our vectorized training data
x_train = vectorize_sequences(train_data)
# Our vectorized test data
x_test = vectorize_sequences(test_data)
# Our vectorized labels
y_train = np.asarray(train_labels).astype('float32')
y_test = np.asarray(test_labels).astype('float32')
# -
# # Fighting overfitting
#
# ## Reducing the network's size
#
#
# The simplest way to prevent overfitting is to reduce the size of the model, i.e. the number of learnable parameters in the model (which is
# determined by the number of layers and the number of units per layer). In deep learning, the number of learnable parameters in a model is
# often referred to as the model's "capacity". Intuitively, a model with more parameters will have more "memorization capacity" and therefore
# will be able to easily learn a perfect dictionary-like mapping between training samples and their targets, a mapping without any
# generalization power. For instance, a model with 500,000 binary parameters could easily be made to learn the class of every digits in the
# MNIST training set: we would only need 10 binary parameters for each of the 50,000 digits. Such a model would be useless for classifying
# new digit samples. Always keep this in mind: deep learning models tend to be good at fitting to the training data, but the real challenge
# is generalization, not fitting.
#
# On the other hand, if the network has limited memorization resources, it will not be able to learn this mapping as easily, and thus, in
# order to minimize its loss, it will have to resort to learning compressed representations that have predictive power regarding the targets
# -- precisely the type of representations that we are interested in. At the same time, keep in mind that you should be using models that have
# enough parameters that they won't be underfitting: your model shouldn't be starved for memorization resources. There is a compromise to be
# found between "too much capacity" and "not enough capacity".
#
# Unfortunately, there is no magical formula to determine what the right number of layers is, or what the right size for each layer is. You
# will have to evaluate an array of different architectures (on your validation set, not on your test set, of course) in order to find the
# right model size for your data. The general workflow to find an appropriate model size is to start with relatively few layers and
# parameters, and start increasing the size of the layers or adding new layers until you see diminishing returns with regard to the
# validation loss.
#
# Let's try this on our movie review classification network. Our original network was as such:
# +
from keras import models
from keras import layers
original_model = models.Sequential()
original_model.add(layers.Dense(16, activation='relu', input_shape=(10000,)))
original_model.add(layers.Dense(16, activation='relu'))
original_model.add(layers.Dense(1, activation='sigmoid'))
original_model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['acc'])
# -
# Now let's try to replace it with this smaller network:
# +
smaller_model = models.Sequential()
smaller_model.add(layers.Dense(4, activation='relu', input_shape=(10000,)))
smaller_model.add(layers.Dense(4, activation='relu'))
smaller_model.add(layers.Dense(1, activation='sigmoid'))
smaller_model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['acc'])
# -
#
# Here's a comparison of the validation losses of the original network and the smaller network. The dots are the validation loss values of
# the smaller network, and the crosses are the initial network (remember: a lower validation loss signals a better model).
original_hist = original_model.fit(x_train, y_train,
epochs=20,
batch_size=512,
validation_data=(x_test, y_test))
smaller_model_hist = smaller_model.fit(x_train, y_train,
epochs=20,
batch_size=512,
validation_data=(x_test, y_test))
epochs = range(1, 21)
original_val_loss = original_hist.history['val_loss']
smaller_model_val_loss = smaller_model_hist.history['val_loss']
# +
import matplotlib.pyplot as plt
# b+ is for "blue cross"
plt.plot(epochs, original_val_loss, 'b+', label='Original model')
# "bo" is for "blue dot"
plt.plot(epochs, smaller_model_val_loss, 'bo', label='Smaller model')
plt.xlabel('Epochs')
plt.ylabel('Validation loss')
plt.legend()
plt.show()
# -
#
# As you can see, the smaller network starts overfitting later than the reference one (after 6 epochs rather than 4) and its performance
# degrades much more slowly once it starts overfitting.
#
# Now, for kicks, let's add to this benchmark a network that has much more capacity, far more than the problem would warrant:
# +
bigger_model = models.Sequential()
bigger_model.add(layers.Dense(512, activation='relu', input_shape=(10000,)))
bigger_model.add(layers.Dense(512, activation='relu'))
bigger_model.add(layers.Dense(1, activation='sigmoid'))
bigger_model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['acc'])
# -
bigger_model_hist = bigger_model.fit(x_train, y_train,
epochs=20,
batch_size=512,
validation_data=(x_test, y_test))
# Here's how the bigger network fares compared to the reference one. The dots are the validation loss values of the bigger network, and the
# crosses are the initial network.
# +
bigger_model_val_loss = bigger_model_hist.history['val_loss']
plt.plot(epochs, original_val_loss, 'b+', label='Original model')
plt.plot(epochs, bigger_model_val_loss, 'bo', label='Bigger model')
plt.xlabel('Epochs')
plt.ylabel('Validation loss')
plt.legend()
plt.show()
# -
#
# The bigger network starts overfitting almost right away, after just one epoch, and overfits much more severely. Its validation loss is also
# more noisy.
#
# Meanwhile, here are the training losses for our two networks:
# +
original_train_loss = original_hist.history['loss']
bigger_model_train_loss = bigger_model_hist.history['loss']
plt.plot(epochs, original_train_loss, 'b+', label='Original model')
plt.plot(epochs, bigger_model_train_loss, 'bo', label='Bigger model')
plt.xlabel('Epochs')
plt.ylabel('Training loss')
plt.legend()
plt.show()
# -
# As you can see, the bigger network gets its training loss near zero very quickly. The more capacity the network has, the quicker it will be
# able to model the training data (resulting in a low training loss), but the more susceptible it is to overfitting (resulting in a large
# difference between the training and validation loss).
# ## Adding weight regularization
#
#
# You may be familiar with _Occam's Razor_ principle: given two explanations for something, the explanation most likely to be correct is the
# "simplest" one, the one that makes the least amount of assumptions. This also applies to the models learned by neural networks: given some
# training data and a network architecture, there are multiple sets of weights values (multiple _models_) that could explain the data, and
# simpler models are less likely to overfit than complex ones.
#
# A "simple model" in this context is a model where the distribution of parameter values has less entropy (or a model with fewer
# parameters altogether, as we saw in the section above). Thus a common way to mitigate overfitting is to put constraints on the complexity
# of a network by forcing its weights to only take small values, which makes the distribution of weight values more "regular". This is called
# "weight regularization", and it is done by adding to the loss function of the network a _cost_ associated with having large weights. This
# cost comes in two flavors:
#
# * L1 regularization, where the cost added is proportional to the _absolute value of the weights coefficients_ (i.e. to what is called the
# "L1 norm" of the weights).
# * L2 regularization, where the cost added is proportional to the _square of the value of the weights coefficients_ (i.e. to what is called
# the "L2 norm" of the weights). L2 regularization is also called _weight decay_ in the context of neural networks. Don't let the different
# name confuse you: weight decay is mathematically the exact same as L2 regularization.
#
# In Keras, weight regularization is added by passing _weight regularizer instances_ to layers as keyword arguments. Let's add L2 weight
# regularization to our movie review classification network:
# +
from keras import regularizers
l2_model = models.Sequential()
l2_model.add(layers.Dense(16, kernel_regularizer=regularizers.l2(0.001),
activation='relu', input_shape=(10000,)))
l2_model.add(layers.Dense(16, kernel_regularizer=regularizers.l2(0.001),
activation='relu'))
l2_model.add(layers.Dense(1, activation='sigmoid'))
# -
l2_model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['acc'])
# `l2(0.001)` means that every coefficient in the weight matrix of the layer will add `0.001 * weight_coefficient_value` to the total loss of
# the network. Note that because this penalty is _only added at training time_, the loss for this network will be much higher at training
# than at test time.
#
# Here's the impact of our L2 regularization penalty:
l2_model_hist = l2_model.fit(x_train, y_train,
epochs=20,
batch_size=512,
validation_data=(x_test, y_test))
# +
l2_model_val_loss = l2_model_hist.history['val_loss']
plt.plot(epochs, original_val_loss, 'b+', label='Original model')
plt.plot(epochs, l2_model_val_loss, 'bo', label='L2-regularized model')
plt.xlabel('Epochs')
plt.ylabel('Validation loss')
plt.legend()
plt.show()
# -
#
#
# As you can see, the model with L2 regularization (dots) has become much more resistant to overfitting than the reference model (crosses),
# even though both models have the same number of parameters.
#
# As alternatives to L2 regularization, you could use one of the following Keras weight regularizers:
# +
from keras import regularizers
# L1 regularization
regularizers.l1(0.001)
# L1 and L2 regularization at the same time
regularizers.l1_l2(l1=0.001, l2=0.001)
# -
# ## Adding dropout
#
#
# Dropout is one of the most effective and most commonly used regularization techniques for neural networks, developed by Hinton and his
# students at the University of Toronto. Dropout, applied to a layer, consists of randomly "dropping out" (i.e. setting to zero) a number of
# output features of the layer during training. Let's say a given layer would normally have returned a vector `[0.2, 0.5, 1.3, 0.8, 1.1]` for a
# given input sample during training; after applying dropout, this vector will have a few zero entries distributed at random, e.g. `[0, 0.5,
# 1.3, 0, 1.1]`. The "dropout rate" is the fraction of the features that are being zeroed-out; it is usually set between 0.2 and 0.5. At test
# time, no units are dropped out, and instead the layer's output values are scaled down by a factor equal to the dropout rate, so as to
# balance for the fact that more units are active than at training time.
#
# Consider a Numpy matrix containing the output of a layer, `layer_output`, of shape `(batch_size, features)`. At training time, we would be
# zero-ing out at random a fraction of the values in the matrix:
# At training time: we drop out 50% of the units in the output
layer_output *= np.randint(0, high=2, size=layer_output.shape)
#
# At test time, we would be scaling the output down by the dropout rate. Here we scale by 0.5 (because we were previous dropping half the
# units):
# At test time:
layer_output *= 0.5
#
# Note that this process can be implemented by doing both operations at training time and leaving the output unchanged at test time, which is
# often the way it is implemented in practice:
# At training time:
layer_output *= np.randint(0, high=2, size=layer_output.shape)
# Note that we are scaling *up* rather scaling *down* in this case
layer_output /= 0.5
#
# This technique may seem strange and arbitrary. Why would this help reduce overfitting? <NAME> has said that he was inspired, among
# other things, by a fraud prevention mechanism used by banks -- in his own words: _"I went to my bank. The tellers kept changing and I asked
# one of them why. He said he didn’t know but they got moved around a lot. I figured it must be because it would require cooperation
# between employees to successfully defraud the bank. This made me realize that randomly removing a different subset of neurons on each
# example would prevent conspiracies and thus reduce overfitting"_.
#
# The core idea is that introducing noise in the output values of a layer can break up happenstance patterns that are not significant (what
# Hinton refers to as "conspiracies"), which the network would start memorizing if no noise was present.
#
# In Keras you can introduce dropout in a network via the `Dropout` layer, which gets applied to the output of layer right before it, e.g.:
model.add(layers.Dropout(0.5))
# Let's add two `Dropout` layers in our IMDB network to see how well they do at reducing overfitting:
# +
dpt_model = models.Sequential()
dpt_model.add(layers.Dense(16, activation='relu', input_shape=(10000,)))
dpt_model.add(layers.Dropout(0.5))
dpt_model.add(layers.Dense(16, activation='relu'))
dpt_model.add(layers.Dropout(0.5))
dpt_model.add(layers.Dense(1, activation='sigmoid'))
dpt_model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['acc'])
# -
dpt_model_hist = dpt_model.fit(x_train, y_train,
epochs=20,
batch_size=512,
validation_data=(x_test, y_test))
# Let's plot the results:
# +
dpt_model_val_loss = dpt_model_hist.history['val_loss']
plt.plot(epochs, original_val_loss, 'b+', label='Original model')
plt.plot(epochs, dpt_model_val_loss, 'bo', label='Dropout-regularized model')
plt.xlabel('Epochs')
plt.ylabel('Validation loss')
plt.legend()
plt.show()
# -
#
# Again, a clear improvement over the reference network.
#
# To recap: here the most common ways to prevent overfitting in neural networks:
#
# * Getting more training data.
# * Reducing the capacity of the network.
# * Adding weight regularization.
# * Adding dropout.
| 4.4-overfitting-and-underfitting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Creating hurricane tracks using Geoanalytics
#
# The sample code below uses big data analytics (GeoAnalytics) to reconstruct hurricane tracks using data registered on a big data file share in the GIS. Note that this functionality is currently available on ArcGIS Enterprise 10.5 and not yet with ArcGIS Online.
#
# ## Reconstruct tracks
# Reconstruct tracks is a type of data aggregation tool available in the `arcgis.geoanalytics` module. This tool works with a layer of point features or polygon features that are time enabled. It first determines which points belong to a track using an identification number or identification string. Using the time at each location, the tracks are ordered sequentially and transformed into a line representing the path of movement.
#
# ## Data used
# For this sample, hurricane data from over a period of 50 years, totalling about 150,000 points split into 5 shape files was used. The [National Hurricane Center](http://www.nhc.noaa.gov/gis/) provides similar datasets that can be used for exploratory purposes.
#
# To illustrate the nature of the data a subset was published as a feature service and can be visualized as below:
# +
from arcgis.gis import GIS
# Create an anonymous connection to ArcGIS Online
arcgis_online = GIS()
hurricane_pts = arcgis_online.content.search("Hurricane_tracks_points AND owner:atma.mani", "Feature Layer")[0]
hurricane_pts
# -
subset_map = arcgis_online.map("USA")
subset_map
subset_map.add_layer(hurricane_pts)
# ## Inspect the data attributes
# Let us query the first layer in hurricane_pts and view its attribute table as a Pandas dataframe.
hurricane_pts.layers[0].query().df.head()
# ## Create a data store
# For the GeoAnalytics server to process your big data, it needs the data to be registered as a data store. In our case, the data is in multiple shape files and we will register the folder containing the files as a data store of type `bigDataFileShare`.
#
# Let us connect to an ArcGIS Enterprise
gis = GIS("https://yourportal.domain.com/webcontext", "username", "password")
# Get the geoanalytics datastores and search it for the registered datasets:
# Query the data stores available
import arcgis
datastores = arcgis.geoanalytics.get_datastores()
bigdata_fileshares = datastores.search()
bigdata_fileshares
# The dataset `hurricanes_all` data is registered as a big data file share with the Geoanalytics datastore, so we can reference it:
data_item = bigdata_fileshares[3]
# If there is no big data file share for hurricane track data registered on the server, we can register one that points to the shared folder containing the shape files.
data_item = datastores.add_bigdata("Hurricane_tracks", r"\\path_to_hurricane_data")
# Once a big data file share is registered, the GeoAnalytics server processes all the valid file types to discern the schema of the data, including information about the geometry in a dataset. If the dataset is time-enabled, as is required to use some GeoAnalytics Tools, the manifest reports the necessary metadata about how time information is stored as well.
#
# This process can take a few minutes depending on the size of your data. Once processed, querying the manifest property returns the schema. As you can see from below, the schema is similar to the subset we observed earlier in this sample.
data_item.manifest['datasets'][0] #for brevity only a portion is printed
# ## Perform data aggregation using reconstruct tracks tool
#
# When you add a big data file share, a corresponding item gets created in your GIS. You can search for it like a regular item and query its layers.
search_result = gis.content.search("", item_type = "big data file share")
search_result
data_item = search_result[4]
data_item
years_50 = data_item.layers[0]
years_50
# ### Reconstruct tracks tool
#
# The `reconstruct_tracks()` function is available in the `arcgis.geoanalytics.summarize_data` module. In this example, we are using this tool to aggregate the numerous points into line segments showing the tracks followed by the hurricanes. The tool creates a feature layer item as an output which can be accessed once the processing is complete.
from arcgis.geoanalytics.summarize_data import reconstruct_tracks
agg_result = reconstruct_tracks(years_50,
track_fields='Serial_Num',
method='GEODESIC')
# ## Inspect the results
# Let us create a map and load the processed result which is a feature service
processed_map = gis.map("USA")
processed_map
processed_map.add_layer(agg_result)
# Thus we transformed a bunch of ponints into tracks that represents paths taken by the hurricanes over a period of 50 years. We can pull up another map and inspect the results a bit more closely
# Our input data and the map widget is time enabled. Thus we can filter the data to represent the tracks from only the years 1860 to 1870
processed_map.set_time_extent('1860', '1870')
# ## What can geoanalytics do for you?
#
# With this sample we just scratched the surface of what big data analysis can do for you. ArcGIS Enterprise at 10.5 packs a powerful set of tools that let you derive a lot of value from your data. You can do so by asking the right questions, for instance, a weather dataset such as this could be used to answer a few interesting questions such as
#
# - did the number of hurricanes per season increase over the years?
# - give me the hurricanes that travelled longest distance
# - give me the ones that stayed for longest time. Do we see a trend?
# - how are wind speed and distance travelled correlated?
# - my assets are located in a tornado corridor. How many times in the past century, was there a hurricane within 50 miles from my assets?
# - my industry is dependent on tourism, which is heavily impacted by the vagaries of weather. From historical weather data, can I correlate my profits with major weather events? How well is my business insulated from freak weather events?
# - over the years do we see any shifts in major weather events - do we notice a shift in when the hurricane season starts?
#
# The ArcGIS API for Python gives you a gateway to easily access the big data tools from your ArcGIS Enterprise. By combining it with other powerful libraries from the pandas and scipy stack and the rich visualization capabilities of the Jupyter notebook, you can extract a lot of value from your data, big or small.
| samples/04_gis_analysts_data_scientists/creating_hurricane_tracks_using_geoanalytics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6.9 64-bit
# name: python3
# ---
# # Model building Pipeline using easyflow feature_encoders module
# This module is a fusion between keras layers and tensorflow feature columns
import pandas as pd
import tensorflow as tf
# local imports
from easyflow.data import TensorflowDataMapper
from easyflow.feature_encoders import FeatureColumnTransformer, FeatureUnionTransformer
from easyflow.feature_encoders import NumericalFeatureEncoder, EmbeddingFeatureEncoder, CategoricalFeatureEncoder
# +
CSV_HEADER = [
"age",
"workclass",
"fnlwgt",
"education",
"education_num",
"marital_status",
"occupation",
"relationship",
"race",
"gender",
"capital_gain",
"capital_loss",
"hours_per_week",
"native_country",
"income_bracket",
]
data_url = (
"https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.data"
)
try:
data_frame = pd.read_csv('adult_features.csv')
labels_binary = pd.read_csv('adult_labels.csv')
except:
data_frame = pd.read_csv(data_url, header=None, names=CSV_HEADER)
labels = data_frame.pop("income_bracket")
labels_binary = 1.0 * (labels == " >50K")
data_frame.to_csv('adult_features.csv', index=False)
labels_binary.to_csv('adult_labels.csv', index=False)
print(f"Train dataset shape: {data_frame.shape}")
# +
batch_size = 256
dataset_mapper = TensorflowDataMapper()
dataset = dataset_mapper.map(data_frame, labels_binary)
train_data_set, val_data_set = dataset_mapper.split_data_set(dataset)
train_data_set = train_data_set.batch(batch_size)
val_data_set = val_data_set.batch(batch_size)
# -
# ## Set up the feature encoding list
# +
NUMERIC_FEATURE_NAMES = [
"age",
"education_num",
"capital_gain",
"capital_loss",
"hours_per_week",
]
CATEGORICAL_FEATURES_NAMES = [
"workclass",
"marital_status",
"relationship",
"race",
"gender"]
EMBEDDING_FEATURES_NAMES = ['education',
'occupation',
'native_country']
# -
feature_encoder_list = [('numerical_features', NumericalFeatureEncoder(), NUMERIC_FEATURE_NAMES),
('categorical_features', CategoricalFeatureEncoder(), CATEGORICAL_FEATURES_NAMES),
('embedding_features_deep', EmbeddingFeatureEncoder(dimension=10), EMBEDDING_FEATURES_NAMES),
('embedding_features_wide', CategoricalFeatureEncoder(), EMBEDDING_FEATURES_NAMES)]
# ## Setting up feature layer and feature encoders
# There are two main column transformer classes namely FeatureColumnTransformer and FeatureUnionTransformer. For this example we are going to build a Wide and Deep model architecture. So we will be using the FeatureColumnTransformer since it gives us more flexibility. FeatureUnionTransformer concatenates all the features in the input layer
feature_layer_inputs, feature_layer = FeatureColumnTransformer(feature_encoder_list).transform(train_data_set)
# +
deep = tf.keras.layers.concatenate([feature_layer['numerical_features'],
feature_layer['categorical_features'],
feature_layer['embedding_features_deep']])
wide = feature_layer['embedding_features_wide']
# +
deep = tf.keras.layers.BatchNormalization()(deep)
for nodes in [128, 64, 32]:
deep = tf.keras.layers.Dense(nodes, activation='relu')(deep)
deep = tf.keras.layers.Dropout(0.5)(deep)
# combine wide and deep layers
wide_and_deep = tf.keras.layers.concatenate([deep, wide])
output = tf.keras.layers.Dense(1, activation='sigmoid')(wide_and_deep)
model = tf.keras.Model(inputs=[v for v in feature_layer_inputs.values()], outputs=output)
model.compile(loss=tf.keras.losses.BinaryCrossentropy(label_smoothing=0.0),
optimizer=tf.keras.optimizers.Adam(learning_rate=0.001),
metrics=[tf.keras.metrics.BinaryAccuracy(name='accuracy'), tf.keras.metrics.AUC(name='auc')])
# -
model.fit(train_data_set, validation_data=val_data_set, epochs=10)
# ## Save and load model
model.save(filepath='tfcolumn_model_example')
del model
loaded_model = tf.keras.models.load_model("tfcolumn_model_example")
loaded_model.predict(val_data_set.take(1))
| notebooks/feature_column_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Dataset
# +
import sys
sys.path.append('../../datasets/')
from prepare_individuals import prepare, germanBats
import matplotlib.pyplot as plt
import torch
import numpy as np
import tqdm
import pickle
classes = germanBats
# +
patch_len = 44 # 88 bei 44100, 44 bei 22050 = 250ms ~ 25ms
X_train, Y_train, X_test, Y_test, X_val, Y_val = prepare("../../datasets/prepared.h5", classes, patch_len)
# -
with open('../call_nocall.indices', 'rb') as file:
indices, labels = pickle.load(file)
train_indices = indices[0][:len(X_train)]
test_indices = indices[1][:len(X_test)]
val_indices = indices[2][:len(X_val)]
X_train = X_train[train_indices]
X_test = X_test[test_indices]
X_val = X_val[val_indices]
Y_train = Y_train[train_indices]
Y_test = Y_test[test_indices]
Y_val = Y_val[val_indices]
print("Total calls:", len(X_train) + len(X_test) + len(X_val))
print(X_train.shape, Y_train.shape)
# +
'''species = [0, 1]
def filterSpecies(s, X, Y):
idx = np.in1d(Y, s)
return X[idx], Y[idx]
X_train, Y_train = filterSpecies(species, X_train, Y_train)
X_test, Y_test = filterSpecies(species, X_test, Y_test)
X_val, Y_val = filterSpecies(species, X_val, Y_val)
classes = {
"Rhinolophus ferrumequinum": 0,
"Rhinolophus hipposideros": 1,
}'''
species = np.asarray([2, 2, 2, 2, 2, 2, 2, 2, 2, 0, 1, 2, 2, 2, 2, 2, 2, 2])
Y_train = species[Y_train]
Y_test = species[Y_test]
Y_val = species[Y_val]
classes = {
"Nyctalus noctula": 0,
"Nyctalus leisleri": 1,
"Other": 2,
}
print("Total calls:", len(X_train) + len(X_test) + len(X_val))
print(X_train.shape, Y_train.shape)
# -
# # Model
import time
import datetime
import tqdm
import torch.nn as nn
import torchvision
from torch.cuda.amp import autocast
from torch.utils.data import TensorDataset, DataLoader
from timm.data.mixup import Mixup
use_stochdepth = False
use_mixedprecision = False
use_imbalancedsampler = False
use_sampler = True
use_cosinescheduler = False
use_reduceonplateu = False
use_nadam = False
use_mixup = False
mixup_args = {
'mixup_alpha': 1.,
'cutmix_alpha': 0.,
'cutmix_minmax': None,
'prob': 1.0,
'switch_prob': 0.,
'mode': 'batch',
'label_smoothing': 0,
'num_classes': len(list(classes))}
mixup_fn = Mixup(**mixup_args)
class Block(nn.Module):
def __init__(self, num_layers, in_channels, out_channels, identity_downsample=None, stride=1):
assert num_layers in [18, 34, 50, 101, 152], "should be a a valid architecture"
super(Block, self).__init__()
self.num_layers = num_layers
if self.num_layers > 34:
self.expansion = 4
else:
self.expansion = 1
# ResNet50, 101, and 152 include additional layer of 1x1 kernels
self.conv1 = nn.Conv2d(in_channels, out_channels, kernel_size=1, stride=1, padding=0)
self.bn1 = nn.BatchNorm2d(out_channels)
if self.num_layers > 34:
self.conv2 = nn.Conv2d(out_channels, out_channels, kernel_size=3, stride=stride, padding=1)
else:
# for ResNet18 and 34, connect input directly to (3x3) kernel (skip first (1x1))
self.conv2 = nn.Conv2d(in_channels, out_channels, kernel_size=3, stride=stride, padding=1)
self.bn2 = nn.BatchNorm2d(out_channels)
self.conv3 = nn.Conv2d(out_channels, out_channels * self.expansion, kernel_size=1, stride=1, padding=0)
self.bn3 = nn.BatchNorm2d(out_channels * self.expansion)
self.relu = nn.ReLU()
self.identity_downsample = identity_downsample
def forward(self, x):
identity = x
if self.num_layers > 34:
x = self.conv1(x)
x = self.bn1(x)
x = self.relu(x)
x = self.conv2(x)
x = self.bn2(x)
x = self.relu(x)
x = self.conv3(x)
x = self.bn3(x)
if self.identity_downsample is not None:
identity = self.identity_downsample(identity)
x = torchvision.ops.stochastic_depth(input=x, p=0.25, mode='batch', training=self.training) # randomly zero input tensor
x += identity
x = self.relu(x)
return x
class ResNet(nn.Module):
def __init__(self, num_layers, block, image_channels, num_classes):
assert num_layers in [18, 34, 50, 101, 152], f'ResNet{num_layers}: Unknown architecture! Number of layers has ' \
f'to be 18, 34, 50, 101, or 152 '
super(ResNet, self).__init__()
if num_layers < 50:
self.expansion = 1
else:
self.expansion = 4
if num_layers == 18:
layers = [2, 2, 2, 2]
elif num_layers == 34 or num_layers == 50:
layers = [3, 4, 6, 3]
elif num_layers == 101:
layers = [3, 4, 23, 3]
else:
layers = [3, 8, 36, 3]
self.in_channels = 64
self.conv1 = nn.Conv2d(image_channels, 64, kernel_size=7, stride=2, padding=3)
self.bn1 = nn.BatchNorm2d(64)
self.relu = nn.ReLU()
self.maxpool = nn.MaxPool2d(kernel_size=3, stride=2, padding=1)
# ResNetLayers
self.layer1 = self.make_layers(num_layers, block, layers[0], intermediate_channels=64, stride=1)
self.layer2 = self.make_layers(num_layers, block, layers[1], intermediate_channels=128, stride=2)
self.layer3 = self.make_layers(num_layers, block, layers[2], intermediate_channels=256, stride=2)
self.layer4 = self.make_layers(num_layers, block, layers[3], intermediate_channels=512, stride=2)
self.avgpool = nn.AdaptiveAvgPool2d((1, 1))
self.fc = nn.Linear(512 * self.expansion, num_classes)
def forward(self, x):
x = self.conv1(x)
x = self.bn1(x)
x = self.relu(x)
x = self.maxpool(x)
x = self.layer1(x)
x = self.layer2(x)
x = self.layer3(x)
x = self.layer4(x)
x = self.avgpool(x)
x = x.reshape(x.shape[0], -1)
x = self.fc(x)
return x
def make_layers(self, num_layers, block, num_residual_blocks, intermediate_channels, stride):
layers = []
identity_downsample = nn.Sequential(nn.Conv2d(self.in_channels, intermediate_channels*self.expansion, kernel_size=1, stride=stride),
nn.BatchNorm2d(intermediate_channels*self.expansion))
layers.append(block(num_layers, self.in_channels, intermediate_channels, identity_downsample, stride))
self.in_channels = intermediate_channels * self.expansion # 256
for i in range(num_residual_blocks - 1):
layers.append(block(num_layers, self.in_channels, intermediate_channels)) # 256 -> 64, 64*4 (256) again
return nn.Sequential(*layers)
def train_epoch(model, epoch, criterion, optimizer, scheduler, dataloader, device):
model.train()
running_loss = 0.0
running_corrects = 0
num_batches = len(dataloader)
num_samples = len(dataloader.dataset)
for batch, (inputs, labels) in enumerate(tqdm.tqdm(dataloader)):
# Transfer Data to GPU if available
inputs, labels = inputs.to(device), labels.to(device)
if use_mixup:
inputs, labels = mixup_fn(inputs, labels)
# Clear the gradients
optimizer.zero_grad()
with autocast(enabled=use_mixedprecision):
# Forward Pass
outputs = model(inputs)
_, predictions = torch.max(outputs, 1)
# Compute Loss
loss = criterion(outputs, labels)
# Calculate gradients
loss.backward()
# Update Weights
optimizer.step()
# Calculate Loss
running_loss += loss.item() * inputs.size(0)
if use_mixup:
running_corrects += (predictions == torch.max(labels, 1)[1]).sum().item()
else:
running_corrects += (predictions == labels).sum().item()
# Perform learning rate step
if use_cosinescheduler:
scheduler.step(epoch + batch / num_batches)
epoch_loss = running_loss / num_samples
epoch_acc = running_corrects / num_samples
return epoch_loss, epoch_acc
def test_epoch(model, epoch, criterion, optimizer, dataloader, device):
model.eval()
num_batches = len(dataloader)
num_samples = len(dataloader.dataset)
with torch.no_grad():
running_loss = 0.0
running_corrects = 0
for batch, (inputs, labels) in enumerate(tqdm.tqdm(dataloader)):
# Transfer Data to GPU if available
inputs, labels = inputs.to(device), labels.to(device)
if use_mixup:
labels = torch.nn.functional.one_hot(labels.to(torch.int64), num_classes=len(list(classes))).float()
# Clear the gradients
optimizer.zero_grad()
# Forward Pass
outputs = model(inputs)
_, predictions = torch.max(outputs, 1)
# Compute Loss
loss = criterion(outputs, labels)
# Update Weights
# optimizer.step()
# Calculate Loss
running_loss += loss.item() * inputs.size(0)
if use_mixup:
running_corrects += (predictions == torch.max(labels, 1)[1]).sum().item()
else:
running_corrects += (predictions == labels).sum().item()
epoch_loss = running_loss / num_samples
epoch_acc = running_corrects / num_samples
return epoch_loss, epoch_acc
batch_size = 64
epochs = 20
lr = 0.01
warmup_epochs = 5
wd = 0.01
# +
from torchsampler import ImbalancedDatasetSampler
from torch.utils.data import WeightedRandomSampler
'''# Experiment: wrong sampling
X = np.concatenate([X_train, X_test, X_val])
Y = np.concatenate([Y_train, Y_test, Y_val])
full_data = TensorDataset(torch.Tensor(np.expand_dims(X, axis=1)), torch.from_numpy(Y))
train_size = int(0.75 * len(full_data))
test_size = len(full_data) - train_size
val_size = int(0.2 * test_size)
test_size -= val_size
train_data, test_data, val_data = torch.utils.data.random_split(full_data, [train_size, test_size, val_size],
generator=torch.Generator().manual_seed(42))'''
if use_mixup and len(X_train) % 2 != 0:
X_train = X_train[:-1]
Y_train = Y_train[:-1]
train_data = TensorDataset(torch.Tensor(np.expand_dims(X_train, axis=1)), torch.from_numpy(Y_train))
test_data = TensorDataset(torch.Tensor(np.expand_dims(X_test, axis=1)), torch.from_numpy(Y_test))
val_data = TensorDataset(torch.Tensor(np.expand_dims(X_val, axis=1)), torch.from_numpy(Y_val))
if use_imbalancedsampler:
train_loader = DataLoader(train_data, sampler=ImbalancedDatasetSampler(train_data), batch_size=batch_size)
test_loader = DataLoader(test_data, sampler=ImbalancedDatasetSampler(test_data), batch_size=batch_size)
val_loader = DataLoader(val_data, sampler=ImbalancedDatasetSampler(val_data), batch_size=batch_size)
elif use_sampler:
def getSampler(y):
_, counts = np.unique(y, return_counts=True)
weights = [len(y)/c for c in counts]
samples_weights = [weights[t] for t in y]
return WeightedRandomSampler(samples_weights, len(y))
train_loader = DataLoader(train_data, sampler=getSampler(Y_train), batch_size=batch_size)
test_loader = DataLoader(test_data, sampler=getSampler(Y_test), batch_size=batch_size)
val_loader = DataLoader(val_data, sampler=getSampler(Y_val), batch_size=batch_size)
else:
train_loader = DataLoader(train_data, batch_size=batch_size)
test_loader = DataLoader(test_data, batch_size=batch_size)
val_loader = DataLoader(val_data, batch_size=batch_size)
# -
model = ResNet(18, Block, image_channels=1, num_classes=len(list(classes)))
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
if torch.cuda.device_count() > 1:
print("Let's use", torch.cuda.device_count(), "GPUs!")
model = nn.DataParallel(model, device_ids=[0, 1])
model.to(device)
print(device)
# +
import wandb
wandb.init(project="BAT-baseline-hierarchical", entity="frankfundel")
wandb.config = {
"learning_rate": lr,
"epochs": epochs,
"batch_size": batch_size
}
criterion = nn.CrossEntropyLoss()
if use_mixup:
criterion = nn.BCEWithLogitsLoss()
optimizer = torch.optim.SGD(model.parameters(), lr=lr)
if use_nadam:
optimizer = torch.optim.NAdam(model.parameters(), lr=lr, weight_decay=wd)
scheduler = None
if use_cosinescheduler:
scheduler = torch.optim.lr_scheduler.CosineAnnealingWarmRestarts(optimizer=optimizer, T_0=warmup_epochs, T_mult=1)
if use_reduceonplateu:
scheduler = torch.optim.lr_scheduler.ReduceLROnPlateau(optimizer)
min_val_loss = np.inf
torch.autograd.set_detect_anomaly(True)
# -
for epoch in range(epochs):
end = time.time()
print(f"==================== Starting at epoch {epoch} ====================", flush=True)
train_loss, train_acc = train_epoch(model, epoch, criterion, optimizer, scheduler, train_loader, device)
print('Training loss: {:.4f} Acc: {:.4f}'.format(train_loss, train_acc), flush=True)
val_loss, val_acc = test_epoch(model, epoch, criterion, optimizer, val_loader, device)
print('Validation loss: {:.4f} Acc: {:.4f}'.format(val_loss, val_acc), flush=True)
if use_reduceonplateu:
scheduler.step(val_loss)
wandb.log({
"train_loss": train_loss,
"train_acc": train_acc,
"val_loss": val_loss,
"val_acc": val_acc,
})
if min_val_loss > val_loss:
print('val_loss decreased, saving model', flush=True)
min_val_loss = val_loss
# Saving State Dict
torch.save(model.state_dict(), 'baseline_nyctalus.pth')
wandb.finish()
model.load_state_dict(torch.load('baseline_nyctalus.pth'))
compiled_model = torch.jit.script(model)
torch.jit.save(compiled_model, 'baseline_nyctalus.pt')
# +
from sklearn.metrics import confusion_matrix
import seaborn as sn
import pandas as pd
Y_pred = []
Y_true = []
corrects = 0
model.eval()
# iterate over test data
for inputs, labels in tqdm.tqdm(test_loader):
output = model(inputs.cuda()) # Feed Network
output = (torch.max(output, 1)[1]).data.cpu().numpy()
Y_pred.extend(output) # Save Prediction
labels = labels.data.cpu().numpy()
Y_true.extend(labels) # Save Truth
# -
# Build confusion matrix
cf_matrix = confusion_matrix(Y_true, Y_pred)
df_cm = pd.DataFrame(cf_matrix / np.sum(cf_matrix, axis=-1), index = [i for i in classes],
columns = [i for i in classes])
plt.figure(figsize = (12,7))
sn.heatmap(df_cm, annot=True)
plt.savefig('baseline_nyctalus_cf.png')
from sklearn.metrics import f1_score
corrects = np.equal(Y_pred, Y_true).sum()
print("Test accuracy:", corrects/len(Y_pred))
print("F1-score:", f1_score(Y_true, Y_pred, average=None).mean())
| models/baseline_hierarchical/baseline-nyctalus.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
def read_csv(S):
f= open(S,'r')
data = f.read()
data_list = data.split("\n")
string_list = data_list[1:-1]
#print(d)
#for i in d :
final_list = []
for i in string_list :
int_fields = []
string_fields = i.split(',')
for y in string_fields :
x = int(y)
int_fields.append(x)
final_list.append(int_fields)
for x in final_list:
return final_list
# +
cdc_list = read_csv("US_births_1994-2003_CDC_NCHS.csv")
print(cdc_list[:10])
# +
def month_births(l) :
births_per_month = {}
for i in l :
month =i[2]
births = i[4]
if month in births_per_month :
births_per_month[month] = births_per_month[month]+births
else :
births_per_month[month] = births
return births_per_month
cdc_month_births = month_births(cdc_list)
print(cdc_month_births)
# +
def dow_births(l) :
births_per_month = {}
for i in l :
month =i[3]
births = i[4]
if month in births_per_month :
births_per_month[month] = births_per_month[month]+births
else :
births_per_month[month] = births
return births_per_month
cdc_day_births = dow_births(cdc_list)
print(cdc_day_births)
# +
def calc_counts(data, column):
births_per_month = {}
if column==3 :
for i in data :
month =i[column]
births = i[4]
if month in births_per_month :
births_per_month[month] = births_per_month[month]+births
else :
births_per_month[month] = births
return births_per_month
if column==2:
for i in data :
month =i[column]
births = i[4]
if month in births_per_month :
births_per_month[month] = births_per_month[month]+births
else :
births_per_month[month] = births
return births_per_month
if column==1:
for i in data :
month =i[column]
births = i[4]
if month in births_per_month :
births_per_month[month] = births_per_month[month]+births
else :
births_per_month[month] = births
return births_per_month
if column == 0 :
for i in data :
month =i[column]
births = i[4]
if month in births_per_month :
births_per_month[month] = births_per_month[month]+births
else :
births_per_month[month] = births
return births_per_month
cdc_year_births = calc_counts(cdc_list,0)
print(cdc_year_births)
cdc_month_births = calc_counts(cdc_list,1)
#print(cdc_month_births)
cdc_dom_births = calc_counts(cdc_list,2)
#print(cdc_dom_births)
cdc_dow_births = calc_counts(cdc_list,3)
#print(cdc_dow_births)
# -
| Guided Project_ Explore U.S. Births/Basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Aula 00 - Intro ao Machine Learning
#
# ### Temas:
# 1. O que é Machine Learning (aprendizagem de máquina)?
# 2. Por que usar Machine Learning?
# 3. Tipos de Machine Learning
# 4. Principais desafios ao trabalhar com Machine Learning
# 5. Como enfrentar um projeto de Machine Learning
#
| lessons/00 - Intro ao Machine Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import tensorflow as tf
import random
data = np.load('mydata.npz')
trX, trY = data[ 'datax' ], data[ 'datay' ]
print("trX.shape:",trX.shape)
print("trY.shape:",trY.shape)
# -
# shuffling the arrays
shuffling = list(zip(trX, trY))
random.shuffle(shuffling)
trX, trY = zip(*shuffling)
trX = np.asarray(trX)
trY = np.asarray(trY)
print("trX.shape:",trX.shape)
print("trY.shape:",trY.shape)
teX, teY = trX [ 3600: ], trY [ 3600: ] # testset
print("teX.shape:",teX.shape)
print("teY.shape:",teY.shape)
trX, trY = trX [ :3600 ], trY [ :3600 ] # trainset
print("trX.shape:",trX.shape)
print("trY.shape:",trY.shape)
teY[0]
trX = trX.reshape( -1, 32, 32, 1)
teX = teX.reshape( -1, 1024)
teY = teY.reshape( -1, 24)
teX.shape
# +
class multinetwork(object):
def __init__(self,x,y,lr):
self.x = x
self.y = y
self.lr = lr
self.opti = multinetwork.optimizer(self)
self.cos = multinetwork.cost(self)
self.acc1,self.acc2 = multinetwork.accuracy(self)
self.scores,self.labels = multinetwork.test(self)
def net(self):
x = tf.reshape(self.x,shape=[-1,32,32,1])
# w0 = tf.Variable(tf.random_normal([3, 3, 1, 32]))
# b0 = tf.Variable(tf.random_normal([32]))
# x_0 = tf.nn.relu(tf.nn.conv2d(x, w0, strides=[1, 1, 1, 1], padding="SAME") + b0)
# p_0 = tf.nn.max_pool(x_0, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='SAME')
w1 = tf.Variable(tf.random_normal([3,3,1,8]))
b1 = tf.Variable(tf.random_normal([8]))
x_1 = tf.nn.tanh(tf.nn.conv2d(x,w1,strides=[1,1,1,1],padding="SAME") + b1)
p_1 = tf.nn.max_pool(x_1,ksize=[1,2,2,1],strides=[1,2,2,1],padding='SAME')
w2 = tf.Variable(tf.random_normal([3,3,8,32]))
b2 = tf.Variable(tf.random_normal([32]))
x_2 = tf.nn.tanh(tf.nn.conv2d(p_1,w2,strides=[1,1,1,1],padding="SAME") + b2)
p_2 = tf.nn.max_pool(x_2,ksize=[1,2,2,1],strides=[1,2,2,1],padding='SAME')
w_f1 = tf.Variable(tf.random_normal([8*8*32,128]))
b_f1 = tf.Variable(tf.random_normal([128]))
p_2f = tf.reshape(p_2,[-1,8*8*32])
f_1 = tf.nn.tanh(tf.matmul(p_2f,w_f1) + b_f1)
f_1d = tf.nn.dropout(f_1,0.5)
w_f2 = tf.Variable(tf.random_normal([128,24]))
b_f2 = tf.Variable(tf.random_normal([24]))
prediction = tf.matmul(f_1d,w_f2) + b_f2
tf.summary.histogram('pred', prediction)
return prediction
def cost(self):
self.score = self.net()
score_split = tf.split(self.score,8,1)
label_split = tf.split(self.y,8,1)
total = 0.0
for i in range ( len(score_split) ):
total += tf.reduce_mean(tf.nn.sigmoid_cross_entropy_with_logits(logits= score_split[i] ,labels= label_split[i] ))
return total/8
def optimizer(self):
return tf.train.AdamOptimizer(learning_rate= self.lr).minimize(self.cost())
def test(self):
self.score = self.net()
score_split = tf.split(self.score, 8, 1)
label_split = tf.split(self.y, 8, 1)
score_split = tf.round(tf.divide(tf.abs(score_split),1e+05,name=None),name=None)
print(score_split,label_split)
return score_split,label_split
def accuracy(self):
self.score = self.net()
score_split = tf.split(self.score, 8, 1)
label_split = tf.split(self.y, 8, 1)
correct_pred1 = tf.equal(tf.argmax(score_split[0], 1), tf.argmax(label_split[0], 1))
correct_pred2 = tf.equal(tf.argmax(score_split[1], 1), tf.argmax(label_split[1], 1))
return correct_pred1, correct_pred2
if __name__ == '__main__':
x = tf.placeholder(tf.float32, [None, 32*32])
y = tf.placeholder(tf.float32, [None, 24])
lr = 0.1
network = multinetwork(x,y,lr)
batch_size = 200
init = tf.global_variables_initializer()
with tf.Session() as sess:
sess.run(init)
index = 0
for batch_i in range(100):
trData_i, trLabel_i = [], []
trData_i .append( trX[ index : index + batch_size ] )
trLabel_i.append( trY[ index : index + batch_size ] )
index += batch_size
if index > ( len(trX) - batch_size+1 ):
index = 0
trData_i = np.reshape(trData_i, (-1, 32 * 32))
trLabel_i = np.reshape(trLabel_i, (-1, 24))
sess.run(network.opti, feed_dict={x: trData_i, y: trLabel_i})
if batch_i % 10 == 0:
cost_tr = sess.run(network.cos, feed_dict={x: trData_i, y: trLabel_i})
cost_te = sess.run(network.cos, feed_dict={x: teX[:3000], y: teY[:3000]})
tf.summary.scalar('train_loss',cost_tr)
tf.summary.scalar('test_loss',cost_te)
# test accuracy
accu1, accu2 = sess.run([network.acc1,network.acc2],
feed_dict={x: teX[:3000], y: teY[:3000]})
# print accu1,accu2
sc,lb = sess.run([network.scores,network.labels],
feed_dict={x: teX[:3000], y: teY[:3000]})
numOfposit = 0.0
for tt in range(accu1.shape[0]):
if accu1[tt] == accu2[tt] or accu1[tt] == True:
numOfposit += 1
test_accu = numOfposit / accu1.shape[0]
accu1, accu2 = sess.run([network.acc1,network.acc2],
feed_dict={x: trData_i, y: trLabel_i})
numOfposit = 0.0
for tt in range(accu1.shape[0]):
if accu1[tt] == accu2[tt] or accu1[tt] == True:
numOfposit += 1
train_accu = numOfposit / accu1.shape[0]
print("%4d, cost_tr: %4.2g , cost_te: %4.2g , trainAccu: %4.2g , testAccu: %4.2g " % (
batch_i, cost_tr, cost_te, train_accu, test_accu))
writer = tf.summary.FileWriter('./logs', sess.graph)
merge_op = tf.summary.merge_all()
# -
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import keras
from IPython.display import SVG
from keras.optimizers import Adam
from keras.utils.vis_utils import model_to_dot
from tqdm import tqdm
from keras import backend as K
from keras.preprocessing.text import Tokenizer
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.feature_extraction.text import CountVectorizer
from numpy import array
from numpy import asarray
from numpy import zeros
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Flatten
from keras.layers import Embedding, Conv1D, MaxPooling1D, Input
from sklearn.metrics import mean_squared_error
from sklearn.metrics import classification_report, confusion_matrix, accuracy_score
from sklearn.model_selection import train_test_split
import gensim
from gensim import utils
import nltk
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings('ignore')
import seaborn as sns
from sklearn.preprocessing import PowerTransformer
from sklearn.preprocessing import QuantileTransformer
import re
plt.xkcd()
# %matplotlib inline
sns.set(style='whitegrid', palette='muted', font_scale=1.2)
# +
from sklearn.dummy import DummyClassifier
from keras.layers import Input, Embedding, Dense, Flatten, Dropout, concatenate
from keras.layers import BatchNormalization, SpatialDropout1D
from keras.callbacks import Callback
from keras.models import Model
from keras.optimizers import Adam
from keras.utils import plot_model
from keras.models import Model
from keras.layers import Input
from keras.layers import Dense
from keras.layers import Flatten
from keras.layers.convolutional import Conv2D
from keras.layers.pooling import MaxPooling2D
from keras.layers.merge import concatenate
from keras.layers.normalization import BatchNormalization
from keras.callbacks import EarlyStopping
from sklearn.preprocessing import Normalizer
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import SVC
from sklearn.gaussian_process import GaussianProcessClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier, AdaBoostClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.utils import class_weight
from sklearn import metrics
from sklearn import preprocessing
# -
df_bills = pd.read_csv('../data/bill_all.csv')
print(df_bills.columns)
df_bills.tail()
df_final = pd.read_csv('../data/df_vote_final.csv')
df_final = df_final[df_final['vote'].isin(['Yea', 'Nay'])]
print(df_final['vote'].unique())
print(df_final.columns)
df_final.tail()
# +
AYE = 1
NAY = -1
vote_map = {'Yea': AYE,'Nay': NAY}
def map_vote(row):
print(row)
return vote_map[row['vote']]
df_final['vote'] = df_final['vote'].apply(lambda x: vote_map[x])
# df_final['vote'] = df_final.vote.apply(map_vote)
print(df_final['vote'].value_counts())
df_final['vote'].value_counts().plot(kind='bar')
# -
word2vec_model = gensim.models.KeyedVectors.load_word2vec_format('/home/sonic/.keras/datasets/GoogleNews-vectors-negative300.bin',
binary=True)
# +
replace_puncts = {'`': "'", '′': "'", '“':'"', '”': '"', '‘': "'"}
strip_chars = [',', '.', '"', ':', ')', '(', '-', '|', ';', "'", '[', ']', '>', '=', '+', '\\', '•', '~', '@',
'·', '_', '{', '}', '©', '^', '®', '`', '<', '→', '°', '€', '™', '›', '♥', '←', '×', '§', '″', '′', 'Â', '█',
'½', 'à', '…', '“', '★', '”', '–', '●', 'â', '►', '−', '¢', '²', '¬', '░', '¶', '↑', '±', '¿', '▾', '═', '¦',
'║', '―', '¥', '▓', '—', '‹', '─', '▒', ':', '¼', '⊕', '▼', '▪', '†', '■', '’', '▀', '¨', '▄', '♫', '☆', 'é',
'¯', '♦', '¤', '▲', 'è', '¸', '¾', 'Ã', '⋅', '‘', '∞', '∙', ')', '↓', '、', '│', '(', '»', ',', '♪', '╩', '╚',
'³', '・', '╦', '╣', '╔', '╗', '▬', '❤', 'ï', 'Ø', '¹', '≤', '‡', '√']
puncts = ['!', '?', '$', '&', '/', '%', '#', '*','£']
def clean_str(x):
x = str(x)
x = x.lower()
x = re.sub(r"(https?:\/\/(?:www\.|(?!www))[a-zA-Z0-9][a-zA-Z0-9-]+[a-zA-Z0-9]\.[^\s]{2,}|www\.[a-zA-Z0-9][a-zA-Z0-9-]+[a-zA-Z0-9]\.[^\s]{2,}|https?:\/\/(?:www\.|(?!www))[a-zA-Z0-9]\.[^\s]{2,}|www\.[a-zA-Z0-9]\.[^\s]{2,})", "url", x)
for k, v in replace_puncts.items():
x = x.replace(k, " {v} ")
for punct in strip_chars:
x = x.replace(punct, ' ')
for punct in puncts:
x = x.replace(punct, ' {punct} ')
x = x.replace(" '", " ")
x = x.replace("' ", " ")
return x
df_bills['billText_clean'] = df_bills['billText'].apply(clean_str)
# -
print(clean_str('u.s \'very" has trump!'))
df_bills['l'] = df_bills['billText_clean'].apply(lambda x: len(str(x).split(' ')))
print('text stats')
# df_bills['l'].plot.hist(bins=5, alpha=0.5)
df_bills['l'].describe()
# +
# %%time
max_words = 20000
MAX_SEQUENCE_LENGTH = 1000
def process_doc(X):
tokenizer = Tokenizer(num_words=max_words,lower=True, split=' ',
filters='"#%&()*+-/<=>@[\\]^_`{|}~\t\n',
char_level=False, oov_token=u'<UNK>')
X_text = X['billText_clean'].values
tokenizer.fit_on_texts(X_text)
print(X.shape)
X_seq = np.array(tokenizer.texts_to_sequences(X_text))
X_seq = pad_sequences(X_seq, maxlen=MAX_SEQUENCE_LENGTH, padding='post')
print('X_seq', X_seq.shape)
count_vect = CountVectorizer()
X_train_counts = count_vect.fit_transform(X_text)
tf_transformer = TfidfTransformer().fit(X_train_counts)
X_train_tf = tf_transformer.transform(X_train_counts)
x_emb = {}
# tokens = nltk.word_tokenize(list(X))
# print('tokens.shape', tokens.shape)
for idx, doc in tqdm(X.iterrows()): #look up each doc in model
# print(doc['legis_num'], doc['billText'])
x_emb[doc['legis_num']] = document_vector(word2vec_model, nltk.word_tokenize(doc['billText_clean'].lower()))
word_index = tokenizer.word_index
print('Found %s unique tokens.' % len(word_index))
return np.array(X_seq), word_index, x_emb, X_train_tf, X_train_counts
def document_vector(word2vec_model, doc):
# remove out-of-vocabulary words
doc = [word for word in doc if word in word2vec_model.vocab]
return np.mean(word2vec_model[doc], axis=0)
def has_vector_representation(word2vec_model, doc):
"""check if at least one word of the document is in the
word2vec dictionary"""
return not all(word not in word2vec_model.vocab for word in doc)
df_bills['billText_clean'] = df_bills['billText_clean'].apply(str)
X_seq, word_index, X_emb, X_train_tf, X_train_counts = process_doc(df_bills)
# df_bills['X_seq'] = X_seq
# df_bills['X_emb'] = X_emb
# df_bills['X_train_tf'] = X_train_tf
# df_bills['X_train_counts'] = X_train_counts
# print(X_emb.shape)
print(X_emb['H R 5010'].shape)
# +
from keras.initializers import glorot_uniform # Or your initializer of choice
from tqdm import tqdm
def plot_history(history):
# print(history.history)
df = pd.DataFrame(history.history)
print(df.describe())
df.plot(xticks=range(epochs))
# print(history.history.keys())
def extract_weights(name, model):
"""Extract weights from a neural network model"""
# Extract weights
weight_layer = model.get_layer(name)
weights = weight_layer.get_weights()[0]
# Normalize
weights = weights / np.linalg.norm(weights, axis = 1).reshape((-1, 1))
return weights
def reset_weights(model):
session = K.get_session()
for layer in model.layers:
if hasattr(layer, 'kernel_initializer'):
layer.kernel.initializer.run(session=session)
def getDataset(df):
dataset = df[['name', 'legis_num',
'vote', 'party',
'sponsor_party', 'sponsor_state',
'sponsor_id']]
# print(df.columns)
dataset['bill_id'] = dataset.legis_num.astype('category').cat.codes.values
dataset['name_id'] = dataset.name.astype('category').cat.codes.values
dataset['vote_orig'] = dataset['vote']
# dataset['vote'] = dataset.vote.astype('category').cat.codes.values
# dataset['vote'] = dataset.vote.apply(map_vote)
dataset['sponsor_party'] = dataset.sponsor_party.astype('category').cat.codes.values
dataset['sponsor_id'] = dataset.sponsor_id.astype('category').cat.codes.values
dataset['sponsor_state'] = dataset.sponsor_state.astype('category').cat.codes.values
# dataset.drop(columns=['name', 'legis_num'], inplace=True)
dataset = dataset.sample(frac=0.9, replace=True)
dataset.reset_index(inplace=True)
return dataset
# -
df_embeding = pd.DataFrame(X_emb)
df_embeding.tail()
# +
scaler = preprocessing.MinMaxScaler( feature_range=(0, 1))
# df_embeding_scaled = scaler.fit_transform(df_embeding)
# DONT normalize
df_embeding_scaled = df_embeding
df_embeding_scaled = pd.DataFrame(df_embeding_scaled, columns=df_embeding.columns)
fig, (ax1, ax2) = plt.subplots(ncols=2, figsize=(6, 5))
ax1.set_title('Before Scaling')
sns.kdeplot(df_embeding, ax=ax1)
ax2.set_title('After Scaling')
sns.kdeplot(df_embeding_scaled, ax=ax2)
plt.show()
df_embeding_scaled.describe()
# +
from keras.layers import Convolution2D, MaxPooling2D
from keras.layers import Input, UpSampling2D
# df_final[df_final['name'] == 'Bateman']
# -
# df_votes_filtered = df_final[df_final['congress'] == 106]
# bill_text = df_embeding_scaled[list(df_votes_filtered['legis_num'].unique())]
# # bill_text.T
# bill_text = np.array(bill_text.T)
# bill_text[1].shape
# df_votes_filtered['legis_num'].unique()
df_final.drop(columns=['Unnamed: 0', 'level_0', 'index', 'Unnamed: 0.1'], inplace=True)
df_final.tail()
df_final = df_final.sort_values('action_date').drop_duplicates(['legis_num','name'], keep='last')
df_final.reset_index(inplace=True)
df_final.tail()
# +
from scipy.stats import norm
from keras.layers import Input, Dense, Lambda
from keras.models import Model
from keras import backend as K
from keras import metrics
# Working
def denoiser_autoencoder(text_all, meta_all, label_all):
print('text_all.shape', text_all.shape)
print('meta_all.shape', meta_all.shape)
input_img = Input(shape=(text_all.shape[1], text_all.shape[2]))
encoded = Dense(256, activation='relu', kernel_initializer='glorot_uniform')(input_img)
encoded = BatchNormalization()(encoded)
encoded = Dense(128, activation='relu')(encoded)
encoded = Dense(64, activation='relu', name='encoded')(encoded)
decoded = Dense(64, activation='relu')(encoded)
decoded = Dense(128, activation='relu')(decoded)
decoded = Dense(text_all.shape[2], activation='sigmoid')(decoded)
autoencoder = Model(input_img, decoded)
loss = 'mean_squared_error'
# loss='binary_crossentropy'
autoencoder.compile(optimizer='RMSprop', loss=loss, metrics=['accuracy', 'binary_crossentropy'])
autoencoder.summary()
encoder = Model(inputs=input_img, outputs= autoencoder.get_layer('encoded').output)
# train
history = autoencoder.fit(meta_all, text_all, epochs=epochs,
batch_size=256, shuffle=True, verbose=1)
plot_history(history)
return autoencoder, encoder
def deep_AE(text_all, meta_all):
print('text_all.shape', text_all.shape)
print('meta_all.shape', meta_all.shape)
input_i = Input(shape=(text_all.shape[1], text_all.shape[2]))
encoded_h1 = Dense(64, activation='tanh')(input_i)
encoded_h2 = Dense(32, activation='tanh')(encoded_h1)
encoded_h3 = Dense(16, activation='tanh')(encoded_h2)
encoded_h4 = Dense(8, activation='tanh')(encoded_h3)
encoded_h5 = Dense(4, activation='tanh')(encoded_h4)
latent = Dense(2, activation='tanh', name='encoded')(encoded_h5)
decoder_h1 = Dense(4, activation='tanh')(latent)
decoder_h2 = Dense(8, activation='tanh')(decoder_h1)
decoder_h3 = Dense(16, activation='tanh')(decoder_h2)
decoder_h4 = Dense(32, activation='tanh')(decoder_h3)
decoder_h5 = Dense(64, activation='tanh')(decoder_h4)
output = Dense(text_all.shape[2], activation='tanh')(decoder_h5)
autoencoder = Model(input_i,output)
autoencoder.compile(optimizer='adadelta', loss='mse')
autoencoder.summary()
encoder = Model(inputs=input_i, outputs= autoencoder.get_layer('encoded').output)
# train
history = autoencoder.fit(meta_all, text_all, epochs=epochs,
batch_size=256, shuffle=True, verbose=1)
plot_history(history)
return autoencoder, encoder
# https://github.com/snatch59/keras-autoencoders/blob/master/variational_autoencoder.py
def VAE(text_all, meta_all, label_all):
original_dim = text_all.shape[1] * text_all.shape[2]
text_all = np.reshape(text_all, [-1, original_dim])
meta_all = np.reshape(meta_all, [-1, original_dim])
print('text_all.shape', text_all.shape)
print('meta_all.shape', meta_all.shape)
batch_size = 100
latent_dim = 2
intermediate_dim = 256
epsilon_std = 1.0
x = Input(shape=(original_dim,))
# x = Input(shape=(text_all.shape[1], text_all.shape[2]))
h = Dense(intermediate_dim, activation='relu')(x)
z_mean = Dense(latent_dim)(h)
z_log_var = Dense(latent_dim)(h)
def sampling(args):
z_mean, z_log_var = args
epsilon = K.random_normal(shape=(K.shape(z_mean)[0], latent_dim), mean=0., stddev=epsilon_std)
return z_mean + K.exp(z_log_var / 2) * epsilon
# note that "output_shape" isn't necessary with the TensorFlow backend
z = Lambda(sampling, output_shape=(latent_dim,))([z_mean, z_log_var])
# we instantiate these layers separately so as to reuse them later
decoder_h = Dense(intermediate_dim, activation='relu')
decoder_mean = Dense(original_dim, activation='sigmoid')
h_decoded = decoder_h(z)
x_decoded_mean = decoder_mean(h_decoded)
# instantiate VAE model
vae = Model(x, x_decoded_mean)
# Compute VAE loss
xent_loss = original_dim * metrics.binary_crossentropy(x, x_decoded_mean)
kl_loss = - 0.5 * K.sum(1 + z_log_var - K.square(z_mean) - K.exp(z_log_var), axis=-1)
vae_loss = K.mean(xent_loss + kl_loss)
vae.add_loss(vae_loss)
vae.compile(optimizer='rmsprop', metrics=['accuracy', 'mse'])
print(vae.summary())
history = vae.fit(meta_all, shuffle=True, epochs=epochs, batch_size=batch_size, verbose=1)
plot_history(history)
# build a model to project inputs on the latent space
encoder = Model(x, z_mean)
return vae, encoder
# +
noise_factor = 0.5
#EPOCHS
epochs = 15
scaler = preprocessing.MinMaxScaler( feature_range=(0, 1))
pred_all = {}
cp_party_all = {}
def runDenoiserAE(df_final):
congress_info = {}
grouped_congress = df_final.groupby('congress')
for congress_session, group in grouped_congress:
print('Processing congress', congress_session)
print('congress shape', group.shape)
cp_party = {}
df_votes_filtered = df_final[df_final['congress'] == congress_session]
num_legistlators = len(df_votes_filtered['name'].unique())
num_bills = len(df_votes_filtered['legis_num'].unique())
print('number of legistlators', num_legistlators)
print('number of bills', num_bills)
dataset = getDataset(df_votes_filtered)
train, test = train_test_split(dataset, test_size=0.2)
print('train.shape', train.shape)
print('test.shape', test.shape)
congress_info[congress_session] = {'num_legislators': num_legistlators,
'num_bills' : num_bills,
'dataset.shape' : dataset.shape[0],
'train.shape': train.shape[0],
'test.shape': test.shape[0]}
# get bill text for the session, this is shared for all CP
bill_text = df_embeding_scaled[list(df_votes_filtered['legis_num'].unique())]
bill_text = np.array(bill_text.T)
print("running embedding mode")
text_all = []
meta_all = []
label_all = []
for name, cp_group in tqdm(train.groupby(['name_id'])):
cp_group.reset_index(inplace=True)
# print(cp_group.shape)
# print(name, cp_group.iloc[0]['name'])
labels = np.random.normal(loc=0.0, scale=1.0, size=num_bills)
# print(labels)
for ind, vote in cp_group.iterrows():
labels[ind] = float(vote['vote'])
# print(labels)
meta = np.multiply(bill_text, np.vstack(labels)) # Eelementwise multiplication, introducing noise
meta = meta + noise_factor * np.random.normal(loc=0.0, scale=1.0, size=meta.shape)
# meta = scaler.fit_transform(meta)
# meta = np.clip(meta, 0., 1.)
# print('bill_text.shape', bill_text.shape)
# print('meta.shape', meta.shape)
# print('labels.shape', labels.shape)
# print(bill_text[0])
# print('**********************')
# print(meta[0])
# plot_emd_meta(bill_text, meta, cp_group.iloc[0]['name'])
text_all.append(bill_text)
meta_all.append(meta)
label_all.append(labels)
# create CP dict
cp_party[cp_group.iloc[0]['name']] = cp_group['party'].unique()[0]
# break
cp_party_all[congress_session] = cp_party
################# VAE ##########################
text_all = np.array(text_all)
original_dim = text_all.shape[1] * text_all.shape[2]
# autoencoder, encoder = VAE(np.asarray(text_all), np.asarray(meta_all), np.asarray(label_all))
# encoded_weights = encoder.predict(np.reshape(text_all, [-1, original_dim]))
#####################################################
# autoencoder, encoder = deep_AE(np.asarray(text_all), np.asarray(meta_all),)
################# DENOISER ##########################
autoencoder, encoder = denoiser_autoencoder(np.asarray(text_all), np.asarray(meta_all), np.asarray(label_all))
encoded_weights = encoder.predict(np.asarray(meta_all))
###########################################
pred_all[congress_session] = encoded_weights
print('encoded_weights.shape', encoded_weights.shape)
print('pred[congress_session].shape', pred_all[congress_session].shape)
break
return congress_info
def plot_emd_meta(embeding, embeding_meta, cp_name):
fig, (ax1, ax2) = plt.subplots(ncols=2, figsize=(6, 5))
ax1.set_title('embedding')
ax1.plot(embeding)
ax2.set_title('meta')
ax2.plot(embeding_meta)
plt.savefig('../data/cp_name/%s.png' % cp_name)
congress_info = runDenoiserAE(df_final)
print(congress_info)
# +
from sklearn import preprocessing
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
import matplotlib
def plot_TNSE_meta(activations, cp_party):
activations = activations.reshape((activations.shape[0], -1))
print(activations.shape)
# transformer = QuantileTransformer(n_quantiles=10, random_state=0)
# transformer = PowerTransformer(method='yeo-johnson')
# activations = transformer.fit_transform(activations)
print('activations.shape', activations.shape)
le = preprocessing.LabelEncoder()
colors = [ 'b','g', 'r']
y = []
# print(cp_party)
for key, value in cp_party.items():
y.append(value)
# print(y)
le.fit(y)
print(le.classes_)
y = le.transform(y)
# print(y)
X_tsne = TSNE(n_components=2, verbose=2).fit_transform(activations)
plt.figure(figsize=(10, 10),)
for color, i, target_name in zip(colors, [0, 1, 2], le.classes_):
plt.scatter(X_tsne[y == i, 0], X_tsne[y == i, 1], cmap=matplotlib.colors.ListedColormap(colors),
color=color, alpha=.6, lw=2, label=target_name)
plt.legend(loc='best', shadow=False, scatterpoints=1)
plt.title('TSNE plot')
plt.show()
# PCA
X_pca = PCA(n_components=2).fit_transform(activations)
plt.figure(figsize=(10, 10),)
for color, i, target_name in zip(colors, [0, 1, 2], le.classes_):
plt.scatter(X_pca[y == i, 0], X_pca[y == i, 1], cmap=matplotlib.colors.ListedColormap(colors),
color=color, alpha=.6, lw=2, label=target_name)
plt.legend(loc='best', shadow=False, scatterpoints=1)
plt.title('PCA plot')
plt.show()
########################################################################
plot_TNSE_meta(pred_all[106], cp_party_all[106])
# -
# pred_all[106]
# print(cp_party_all[106])
print(pred_all[106].shape)
activations = pred_all[106].reshape((pred_all[106].shape[0], -1))
print(activations.shape)
# +
from sklearn.preprocessing import StandardScaler
from sklearn import cluster
activations = pred_all[106]
activations = activations.reshape((activations.shape[0], -1))
plt.figure(figsize=(12, 12))
from sklearn.cluster import KMeans
# normalize dataset for easier parameter selection
activations = StandardScaler().fit_transform(activations)
# Incorrect number of clusters
y_pred = KMeans(n_clusters=2, random_state=42).fit_predict(activations)
plt.subplot(221)
plt.scatter(activations[:, 0], activations[:, 1], c=y_pred)
plt.title("Incorrect Number of Blobs")
plt.show
print(activations.shape)
df = pd.DataFrame(activations)
df.tail()
# -
| neural_collaborative_filtering/AutoEncoder-Denoise-Embedding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Simple Class based generator
# ```
# Py generators allow you to declare a function that behaves as a
# an iterator, hence you needn't return the values immediately
# and retrieve it iteratively.
#
#
# For an object to be an iterator it should implement the __iter__ method which will return the iterator object, the __next__ method will then return the next value in the sequence and possibly might raise the StopIteration exception when there are no values to be returned.
#
#
# Difference Between Generator Functions and Regular Functions
# The main difference between a regular function and generator functions is that the state of generator functions are maintained through the use of the keyword yield and works much like using return, but it has some important differences. the difference is that yield saves the state of the function. The next time the function is called, execution continues from where it left off, with the same variable values it had before yielding, whereas the return statement terminates the function completely. Another difference is that generator functions don’t even run a function, it only creates and returns a generator object. Lastly, the code in generator functions only execute when next() is called on the generator object.
# ```
class Odds:
def __init__(self, max_val, begin=1):
self.n = begin
self.max_val = max_val
def __iter__(self):
return self
def __next__(self):
"""
Here we are trying to generate every odd number in
the sequence until max_val is encountered
"""
if self.n <= self.max_val:
result = self.n
self.n+=2
return result
else:
raise StopIteration
for odd in Odds(10):
print(odd)
# +
odd = Odds(10)
print(next(odd))
print(next(odd))
print(next(odd))
print(next(odd))
# -
# ### Generator Implementation in Python
# +
def get_odds_generator():
"""
The way this works is
successive calls to next, will retrive the value from each yield
"""
n=1
n+=2
yield n
n+=2
yield n
n+=2
yield n
nums = get_odds_generator()
print(next(nums))
print(next(nums))
print(next(nums))
# calling the third time will lead to an error
print(next(nums))
# -
def get_odds_generator(max):
n=1
while n<=max:
yield n
n+=2
else:
raise StopIteration
# +
numbers=get_odds_generator(3)
try:
print(next(numbers))
print(next(numbers))
print(next(numbers))
except RuntimeError as e:
print(e)
# -
# ### creating a power of 2 generator
# +
def power_of_two(max_val=200):
val = 2
while val<=max_val:
result = val
val*=2
yield result
else:
raise StopIteration
num = power_of_two()
print(next(num))
print(next(num))
print(next(num))
# -
# ```
# iterators are useful to work with large stream of data
# which cannot fit into memory at a time
# we can use generator to handle one data at a time
# ```
def fibonacci_generator():
n1 = 0
n2 = 1
while True:
yield n1
n1,n2 = n2, n1+n2
sequence = fibonacci_generator()
print(next(sequence))
print(next(sequence))
print(next(sequence))
print(next(sequence))
# ### using generators to generate a random no. 0f 1s (<=R*C) in a RxC matrix
from random import randint
def random_pattern(M=3,N=3,no_of_ones=3):
while True:
base_mtx = [[0] * M for _ in range(N)]
seen = set()
while len(seen) < no_of_ones:
num = randint(0,N*M-1)
if num not in seen:
seen.add(num)
while seen:
num = seen.pop()
r,c = num//N, num%M
base_mtx[r][c] = 1
yield base_mtx
rand_matrix = random_pattern()
for _ in range(10):
print(next(rand_matrix))
| Basics/Python_For_SDC/Generators.ipynb |
# # Cython Magic Functions
# ## Loading the extension
# IPython had a `cythonmagic` extension that contains a number of magic functions for working with Cython code. This extension can be found in the Cython package now and can be loaded using the `%load_ext` magic as follows:
# %load_ext Cython
# ## The %cython_inline magic
# The `%%cython_inline` magic uses `Cython.inline` to compile a Cython expression. This allows you to enter and run a function body with Cython code. Use a bare `return` statement to return values.
a = 10
b = 20
# %%cython_inline
return a+b
# ## The %cython_pyximport magic
# The `%%cython_pyximport` magic allows you to enter arbitrary Cython code into a cell. That Cython code is written as a `.pyx` file in the current working directory and then imported using `pyximport`. You have to specify the name of the module that the Code will appear in. All symbols from the module are imported automatically by the magic function.
# %%cython_pyximport foo
def f(x):
return 4.0*x
f(10)
# ## The %cython magic
# Probably the most important magic is the `%cython` magic. This is similar to the `%%cython_pyximport` magic, but doesn't require you to specify a module name. Instead, the `%%cython` magic manages everything using temporary files in the `~/.cython/magic` directory. All of the symbols in the Cython module are imported automatically by the magic.
#
# Here is a simple example of a Black-Scholes options pricing algorithm written in Cython. Please note that this example might not compile on non-POSIX systems (e.g., Windows) because of a missing `erf` symbol.
# + language="cython"
# cimport cython
# from libc.math cimport exp, sqrt, pow, log, erf
#
# @cython.cdivision(True)
# cdef double std_norm_cdf_cy(double x) nogil:
# return 0.5*(1+erf(x/sqrt(2.0)))
#
# @cython.cdivision(True)
# def black_scholes_cy(double s, double k, double t, double v,
# double rf, double div, double cp):
# """Price an option using the Black-Scholes model.
#
# s : initial stock price
# k : strike price
# t : expiration time
# v : volatility
# rf : risk-free rate
# div : dividend
# cp : +1/-1 for call/put
# """
# cdef double d1, d2, optprice
# with nogil:
# d1 = (log(s/k)+(rf-div+0.5*pow(v,2))*t)/(v*sqrt(t))
# d2 = d1 - v*sqrt(t)
# optprice = cp*s*exp(-div*t)*std_norm_cdf_cy(cp*d1) - \
# cp*k*exp(-rf*t)*std_norm_cdf_cy(cp*d2)
# return optprice
# -
black_scholes_cy(100.0, 100.0, 1.0, 0.3, 0.03, 0.0, -1)
# For comparison, the same code is implemented here in pure python.
# +
from math import exp, sqrt, pow, log, erf
def std_norm_cdf_py(x):
return 0.5*(1+erf(x/sqrt(2.0)))
def black_scholes_py(s, k, t, v, rf, div, cp):
"""Price an option using the Black-Scholes model.
s : initial stock price
k : strike price
t : expiration time
v : volatility
rf : risk-free rate
div : dividend
cp : +1/-1 for call/put
"""
d1 = (log(s/k)+(rf-div+0.5*pow(v,2))*t)/(v*sqrt(t))
d2 = d1 - v*sqrt(t)
optprice = cp*s*exp(-div*t)*std_norm_cdf_py(cp*d1) - \
cp*k*exp(-rf*t)*std_norm_cdf_py(cp*d2)
return optprice
# -
black_scholes_py(100.0, 100.0, 1.0, 0.3, 0.03, 0.0, -1)
# Below we see the runtime of the two functions: the Cython version is nearly a factor of 10 faster.
# %timeit black_scholes_cy(100.0, 100.0, 1.0, 0.3, 0.03, 0.0, -1)
# %timeit black_scholes_py(100.0, 100.0, 1.0, 0.3, 0.03, 0.0, -1)
# ## External libraries
# Cython allows you to specify additional libraries to be linked with your extension, you can do so with the `-l` flag (also spelled `--lib`). Note that this flag can be passed more than once to specify multiple libraries, such as `-lm -llib2 --lib lib3`. Here's a simple example of how to access the system math library:
# + magic_args="-lm" language="cython"
# from libc.math cimport sin
# print 'sin(1)=', sin(1)
# -
# You can similarly use the `-I/--include` flag to add include directories to the search path, and `-c/--compile-args` to add extra flags that are passed to Cython via the `extra_compile_args` of the distutils `Extension` class. Please see [the Cython docs on C library usage](http://docs.cython.org/src/tutorial/clibraries.html) for more details on the use of these flags.
| examples/Builtin Extensions/Cython Magics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Selección de modelo
#
# El objetivo de realizar selección de modelo es, a partir de una serie de datos $(x,y)$ y cierta clase de hipótesis $\mathcal{H}$. Encontrar el $h\in \mathcal{H}$ que minimiza el error $e(h)=P_{D}[h(x) \neq y]$. Claramente la hipótesis $h$ que minimiza el error en los datos sobre una cantidad suficiente de datos, produce un error $e(h)$ pequeño. El objetivo es balancear la complejidad de $\mathcal{H}$ con el ajuste de $h\in \mathcal{H}$ a los datos de entrenamiento. Es claro que una hipótesis muy simple puede no contener una buena aproximación a la función que queremos aprender, mientras que una $h$ muy compleja puede ajustarse muy bien a los datos pero no ser tan buena en la generalización a nuevos datos. Esto se vuelve particularmente importante cuando se tienen pocos datos o estos resultan muy ruidosos.
#
#
# # Estrategias para selección de modelo
#
# Es importante notar que la complejidad de la clase de modelos es una variable a determinar por el algoritmo de aprendizaje. La selección de modelo se realiza usualmente de la siguiente manera:
# - Se selecciona una función candidata $h_i$ de cierta clase $H_i$, para esto es conveniente, en general, minimizar el error empírico.
# - Se utiliza algún criterio para seleccionar una de las $h\in {h_1,h_2,...}$ de manera que el error $e(h)$ sea lo suficientemente pequeño.
#
#
# # 1. Validación cruzada
#
# Cuando se evalúan diferentes configuraciones de un modelo general, lo que se hace es evaluar diferentes "hiperparámetros" para los estimadores. Tales como $\lambda$ en el caso de regresión lineal con regularización que se realizó previamente. Cuando se escoje un valor de un hiperparáemtetro escogido manuealmente, existe la posibilidad de realizar sobreajuste (overfitting) del conjunto de prueba debido a que los hiperparámetros pueden ser manipulados de manera que el estimador se comporte óptimamente, de manera que se introduce un conocimiento al modelo que invalidala idea de tener un desempeño de generalización. Para resolver este problema, es posible crear OTRO subconjunto de datos de manera que este se pueda mantener "afuera" del conjunto de prueba y funcione como un conjunto de validación durante el proceso de entrenamiento.
#
# Para realizar validación cruzada, se realiza una estimación directa de $e(h_i)$. Para esto, se dividen los datos en conjuntos $(X_{train},Y_{train})$ y $(X_{test},Y_{test})$, donde cada conjunto contiene $1-\alpha$ y $alpha$ veces la cantidad total de datos.
# Con estos datos se halla la hipótesis candidata $h_d\in \mathcal{H}_d$ minimizando el error empírico en el conjunto de datos de entrenamiento (X_{train},Y_{train}). Posteriormente, se selecciona la hipótesis candidata con el menor error empírico en $(X_{test},Y_{test})$.
#
#
#
# \begin{equation}
# h_{d^*}={argmin}_{h_i}{\hat{e}_{(X_{test},Y_{test})} (h_d)}
# \end{equation}
#
# Claramente, la selección de $\alpha$ también tiene consecuencias sobre la estimación y el modelo obtenido. Si se selecciona un $\alpha$ muy pequeño, es posible tener una estimación pobre de $e(h)$ debido a que no se tienen suficientes datos para estimar el error empírico en un conjunto de prueba. Si se selecciona un $\alpha$ muy grande, se corre el riesgo de realizar un aprendizaje pobre debido a que no se tienen los suficientes datos para entrenar el modelo. El criterio de Chernoff es útil determinar qué cantidad $n$ de datos es necesaria para estimar $e(h)$ con precisión $\epsilon$ y confianza $1-\delta$:
#
# \begin{equation}
# n \geq \frac{1}{2\epsilon^2} \ln{\frac{2}{\delta}}
# \end{equation}
#
# Sin embargo, el estimativo de $e(h_d)$ usualmente es ruidoso, por lo que es importante considerar diferentes alternativas para realizar \textit{validación cruzada}.
#
# Sin embargo, al realizar particiones de los datos, se reduce el número de muestras que pueden ser utilizadas para aprender el modelo. Una solución a este problema se llama "validación cruzada (CV)".
#
#
#
# +
#EJEMPLO: SCORES DE CROSS-VALIDATION
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn import datasets
from sklearn import svm
from sklearn.model_selection import cross_val_score
from sklearn import metrics
#Se cargan los datos
iris = datasets.load_iris()
#Se crean las separaciones de train y test.
X_train, X_test, y_train, y_test = train_test_split(iris.data, iris.target, test_size=0.3, random_state=0)
#Se crea un objeto de tipo clasificador
clf = svm.SVC(kernel='linear', C=1)
#Se crea un objeto cross_val_score que recibe como parámetro el modelo,
#los datos y la cantidad de validaciones cruzadas
scores = cross_val_score(clf, X_train , y_train, cv=5)
print("Precisiones en cada subconjunto",scores)
print("Precisión media: %0.2f (+/- %0.2f)" % (scores.mean(), scores.std() * 2))
print(".......")
#Se puede modificar el tipo de scoring que se realiza, para un conjunto de datos balanceado el resultado es muy similar
scores = cross_val_score(clf, X_train, y_train, cv=5, scoring='f1_macro')
print("Precisiones en cada subconjunto",scores)
print("Precisión media: %0.2f (+/- %0.2f)" % (scores.mean(), scores.std() * 2))
# -
# #### Iteradores
# También es posible utilizar diferentes estrategias de validación cruzada pasando un objeto de tipo iterador (por defecto scikit-learn utilizará KFold o StratifiedKFold)
# +
from sklearn.model_selection import ShuffleSplit
cv = ShuffleSplit(n_splits=5, test_size=0.3, random_state=0)
scores=cross_val_score(clf, X_train, y_train, cv=cv)
print("Precisiones en cada subconjunto",scores)
print("Precisión media: %0.2f (+/- %0.2f)" % (scores.mean(), scores.std() * 2))
# -
# # Validación cruzada k-multiple
#
# La idea es obtener un mejor estimativo de $e(h)$. Para esto, teniendo una clase $\mathcal{H}$, el conjunto de datos $S=(X,Y)$ se divide en $S_1,S_2,...,S_k$. Para cada $i=1,2,...k$ se halla $h_i$ minimizando el error empírico en $\cup_{j\neq i} S_j$. Luego, se estima el error calculando el error empírico $\hat{e}_{S_i}(h_i)$. Luego se realiza un promedio de los valores obtenidos
#
# \begin{equation}
# \hat{e}(h_d)=\frac{1}{k} \sum_{i=1}^{k}{\hat{e}s_i (h_i)}
# \end{equation}
#
# Para el $d^*$ que corresponde "fold" con menor valor de riesgo empírico, se halla $h$ minimizando el error empírico en $S$. Es importante mencionar que la validación cruzada k-fold es un proceso costoso computacionalmente y carece de soporte teórico, sin embargo, es ampliamente usado en la práctica.
#
# ## K fold y Leave-one-out.
# En k-fold se segmentan los datos en $k$ grupos, se corren los algoritmos en $k-1$ grupos y se evalúa el riesgo del último grupo. Se repiite esto hasta que todos los grupos se hallan entrenando, por lo que se tienen $k$ números de errores que se deben promediar. Leave-one-out es el caso en el que se tiene $k=n$. La idea es que este método es más robusto, dado que se entrena en una mayor cantidad de datos y todavía se tienen promedios del riesgo estimado. el reto es que ahora se está reutilizando la información en diferentes entrenamientos y conjuntos de validación por lo que los términos en los promedios no son independientes. [1]
#
#
#
# +
import numpy as np
from sklearn.model_selection import KFold
X = np.random.uniform(size=100)
kf = KFold(n_splits=4)
for train, test in kf.split(X): print("%s %s" % (train, test))
# -
# # Comparación de comportamiento de objetos de validación cruzada
# +
#Tomado de https://scikit-learn.org/stable/auto_examples/model_selection/plot_cv_indices.html
from sklearn.model_selection import (TimeSeriesSplit, KFold, ShuffleSplit,
StratifiedKFold, GroupShuffleSplit,
GroupKFold, StratifiedShuffleSplit)
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.patches import Patch
np.random.seed(1338)
cmap_data = plt.cm.Paired
cmap_cv = plt.cm.coolwarm
n_splits = 4
def plot_cv_indices(cv, X, y, group, ax, n_splits, lw=10):
"""Create a sample plot for indices of a cross-validation object."""
# Generate the training/testing visualizations for each CV split
for ii, (tr, tt) in enumerate(cv.split(X=X, y=y, groups=group)):
# Fill in indices with the training/test groups
indices = np.array([np.nan] * len(X))
indices[tt] = 1
indices[tr] = 0
# Visualize the results
ax.scatter(range(len(indices)), [ii + .5] * len(indices),
c=indices, marker='_', lw=lw, cmap=cmap_cv,
vmin=-.2, vmax=1.2)
# Plot the data classes and groups at the end
ax.scatter(range(len(X)), [ii + 1.5] * len(X),
c=y, marker='_', lw=lw, cmap=cmap_data)
ax.scatter(range(len(X)), [ii + 2.5] * len(X),
c=group, marker='_', lw=lw, cmap=cmap_data)
# Formatting
yticklabels = list(range(n_splits)) + ['class', 'group']
ax.set(yticks=np.arange(n_splits+2) + .5, yticklabels=yticklabels,
xlabel='Sample index', ylabel="CV iteration",
ylim=[n_splits+2.2, -.2], xlim=[0, 100])
ax.set_title('{}'.format(type(cv).__name__), fontsize=15)
return ax
# Generate the class/group data
n_points = 100
X = np.random.randn(100, 10)
percentiles_classes = [.1, .3, .6]
y = np.hstack([[ii] * int(100 * perc)
for ii, perc in enumerate(percentiles_classes)])
# Evenly spaced groups repeated once
groups = np.hstack([[ii] * 10 for ii in range(10)])
def visualize_groups(classes, groups, name):
# Visualize dataset groups
fig, ax = plt.subplots()
ax.scatter(range(len(groups)), [.5] * len(groups), c=groups, marker='_',
lw=50, cmap=cmap_data)
ax.scatter(range(len(groups)), [3.5] * len(groups), c=classes, marker='_',
lw=50, cmap=cmap_data)
ax.set(ylim=[-1, 5], yticks=[.5, 3.5],
yticklabels=['Data\ngroup', 'Data\nclass'], xlabel="Sample index")
visualize_groups(y, groups, 'no groups')
cvs = [KFold, GroupKFold, ShuffleSplit, StratifiedKFold,
GroupShuffleSplit, StratifiedShuffleSplit, TimeSeriesSplit]
for cv in cvs:
this_cv = cv(n_splits=n_splits)
fig, ax = plt.subplots(figsize=(6, 3))
plot_cv_indices(this_cv, X, y, groups, ax, n_splits)
ax.legend([Patch(color=cmap_cv(.8)), Patch(color=cmap_cv(.02))],
['Testing set', 'Training set'], loc=(1.02, .8))
# Make the legend fit
plt.tight_layout()
fig.subplots_adjust(right=.7)
plt.show()
# -
# # Aplicaciones:
#
# ## Búsqueda de parámetros utilizando gridsearch:
# Scikit-learn incorpora un objeto que, dados unos datos, calcula el score durante el ajuste de cierto estimador o parámetro y escoge los parámetros que maximizan el score de validación cruzada. Este objeto toma un estimado durante la construcción y devuelve un estimador.
#
# +
from sklearn.model_selection import GridSearchCV, cross_val_score
import numpy as np
from sklearn.model_selection import cross_val_score
from sklearn import datasets, svm
digits = datasets.load_digits()
X = digits.data
y = digits.target
X_train, X_test, y_train, y_test = train_test_split(iris.data, iris.target, test_size=0.4, random_state=0)
svc = svm.SVC(kernel='linear')
Cs = np.logspace(-6, -1, 10)
clf = GridSearchCV(estimator=svc, param_grid=dict(C=Cs),n_jobs=-1,iid=True,cv=5)
clf.fit(X_train,y_train)
#Se puede recuperar el mejor parámetro
print(clf.best_estimator_.C)
#También se puede recuperar el score
clf.score(X_test,y_test)
# +
#Tomado de: https://scikit-learn.org/stable/auto_examples/model_selection/plot_grid_search_digits.html#sphx-glr-auto-examples-model-selection-plot-grid-search-digits-py
from sklearn import datasets
from sklearn.model_selection import train_test_split
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import classification_report
from sklearn.svm import SVC
# Loading the Digits dataset
digits = datasets.load_digits()
# To apply an classifier on this data, we need to flatten the image, to
# turn the data in a (samples, feature) matrix:
n_samples = len(digits.images)
X = digits.images.reshape((n_samples, -1))
y = digits.target
# Split the dataset in two equal parts
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.5, random_state=0)
# Set the parameters by cross-validation
tuned_parameters = [{'kernel': ['rbf'], 'gamma': [1e-3, 1e-4],
'C': [1, 10, 100, 1000]},
{'kernel': ['linear'], 'C': [1, 10, 100, 1000]}]
scores = ['precision', 'recall']
for score in scores:
print("#hyper-parameters para %s" % score)
print()
clf = GridSearchCV(SVC(), tuned_parameters, cv=5,
scoring='%s_macro' % score)
clf.fit(X_train, y_train)
print("Mejores parámetros encontrados:")
print()
print(clf.best_params_)
print()
print("Grid scores on development set:")
print()
means = clf.cv_results_['mean_test_score']
stds = clf.cv_results_['std_test_score']
for mean, std, params in zip(means, stds, clf.cv_results_['params']):
print("%0.3f (+/-%0.03f) for %r"
% (mean, std * 2, params))
print()
print("Reporte detallado de clasificación:")
print()
print("The model is trained on the full development set.")
print("The scores are computed on the full evaluation set.")
print()
y_true, y_pred = y_test, clf.predict(X_test)
print(classification_report(y_true, y_pred))
print()
# -
# ## Eliminación de características:
#
# Es posible realizar una eliminación recursiva de características con una sintonización automática del número de características seleccionadas con cross-validation. Para realizar esta aplicación primero es importante presentar algunas funciones útiles.
#
# ### Nuevo! make_classification
#
# Genera un objeto para solucionar un problema multiclase, en general. Este objeto soluciona el problema creando clusters de puntos normalmente distribuidos sobre los vértices de un n_informativo hipercubo con lados de tamaño $2*class_sep$. Introduce interdependencia entre las características y añade diferentes tipos de ruido a los datos.
# to the data.
#
# ### Nuevo! RFECV
# Realiza RANKING con eliminación recursiva y validación cruzada de características de las mejores características de los modelos.
#
# +
import matplotlib.pyplot as plt
from sklearn.svm import SVC
from sklearn.model_selection import StratifiedKFold
from sklearn.feature_selection import RFECV
from sklearn.datasets import make_classification
# Crea un problema de clasificación, con 3 n_informativos
X, y = make_classification(n_samples=1000, n_features=25, n_informative=3,
n_redundant=2, n_repeated=0, n_classes=8,
n_clusters_per_class=1, random_state=0)
# Crea un objeto de clasificador.
svc = SVC(kernel="linear")
# Crea un objeto de ranking, el ESTIMADOR buscado es el clasificador SVC y el criterio es acuracy.
rfecv = RFECV(estimator=svc, step=1, cv=StratifiedKFold(2),
scoring='accuracy')
rfecv.fit(X, y)
print("Número óptimo de características : %d" % rfecv.n_features_)
# Número de features en función del score de cross validation
plt.figure(figsize=(10,10))
plt.xlabel("Número de características seleccionadas")
plt.ylabel("Puntaje de validación cruzada (Número de clasificaciones correctas)")
plt.plot(range(1, len(rfecv.grid_scores_) + 1), rfecv.grid_scores_)
plt.show()
# -
#
| 10ML/.ipynb_checkpoints/1_ModSel-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import scipy.linalg
def rearrange(matrix):
n = len(matrix)
pivot_matrix = np.eye(n, n)
for index, column in enumerate(np.absolute(matrix.T)):
row = index + np.argmax(column[index:])
if index != row:
pivot_matrix[[index, row]] = pivot_matrix[[row, index]]
return np.dot(pivot_matrix, matrix)
def lu_decomposition(matrix):
n = len(matrix)
lower = np.eye(n, n)
upper = np.zeros((n, n))
rearranged = rearrange(matrix)
for j in range(n):
# вычисляем верхнюю матрицу
for i in range(j + 1):
upper[i, j] = rearranged[i, j] - np.dot(upper[:i, j], lower[i, :i])
# вычисляем нижнюю матрицу
for i in range(j, n):
lower[i, j] = (rearranged[i, j] - np.dot(upper[:i, j], lower[i, :i]))/upper[j, j]
return (lower, upper)
matrix = np.array([[1.00, 0.17, -0.25, 0.54], [0.47, 1.00, 0.67, -0.32], [-0.11, 0.35, 1.00, -0.74], [0.55, 0.43, 0.36, 1.00]])
matrix
lu_decomposition(matrix) # реализованная функция
scipy.linalg.lu(matrix, permute_l=True) # встроенная функция
| Numerical-Methods/LU-decomposition/LU-decomposition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (DenseNet-MURA-PyTorch)
# language: python
# name: myenv
# ---
# # Abnormality Detection in Musculoskeletal Radiographs
# Author: <NAME>
# The objective is to build a machine learning model that can detect an abnormality in the X-Ray radiographs. These models can help towards providing healthcare access to the parts of the world where access to skilled radiologists is limited. According to a study on the Global Burden of Disease and the worldwide impact of all diseases found that, “musculoskeletal conditions affect more than 1.7 billion people worldwide. They are the 2nd greatest cause of disabilities, and have the 4th greatest impact on the overall health of the world population when considering both death and disabilities”. (www.usbji.org, n.d.).
#
# This project attempts to implement deep neural network using DenseNet169 inspired from the Stanford Paper Rajpurkar, et al., 2018.
# ## XR_ELBOW Study Type
# ## Phase 3: Data Preprocessing
# As per the paper, i have normalized the each image to have same mean & std of the images in the ImageNet training set. In the paper, they have used variable-sized images to 320 x 320. But i have chosen to scale 224 x 224. Then i have augmented the data during the training by applying random lateral inversions and rotations of up to 30 degrees using
from keras.applications.densenet import DenseNet169, DenseNet121, preprocess_input
from keras.preprocessing.image import ImageDataGenerator, load_img, image
from keras.models import Sequential, Model, load_model
from keras.layers import Conv2D, MaxPool2D
from keras.layers import Activation, Dropout, Flatten, Dense
from keras.callbacks import EarlyStopping, ReduceLROnPlateau, ModelCheckpoint, Callback
from keras import regularizers
import pandas as pd
from tqdm import tqdm
import os
import numpy as np
import random
from keras.optimizers import Adam
import keras.backend as K
import cv2
import matplotlib.pyplot as plt
# ### 3.1 Data preprocessing
#Utility function to find the list of files in a directory excluding the hidden files.
def listdir_nohidden(path):
for f in os.listdir(path):
if not f.startswith('.'):
yield f
# ### 3.1.1 Creating a csv file containing path to image & csv
def create_images_metadata_csv(category,study_types):
"""
This function creates a csv file containing the path of images, label.
"""
image_data = {}
study_label = {'positive': 1, 'negative': 0}
#study_types = ['XR_ELBOW','XR_FINGER','XR_FOREARM','XR_HAND','XR_HUMERUS','XR_SHOULDER','XR_WRIST']
#study_types = ['XR_ELBOW']
i = 0
image_data[category] = pd.DataFrame(columns=['Path','Count', 'Label'])
for study_type in study_types: # Iterate throught every study types
DATA_DIR = 'data/MURA-v1.1/%s/%s/' % (category, study_type)
patients = list(os.walk(DATA_DIR))[0][1] # list of patient folder names
for patient in tqdm(patients): # for each patient folder
for study in os.listdir(DATA_DIR + patient): # for each study in that patient folder
if(study != '.DS_Store'):
label = study_label[study.split('_')[1]] # get label 0 or 1
path = DATA_DIR + patient + '/' + study + '/' # path to this study
for j in range(len(list(listdir_nohidden(path)))):
image_path = path + 'image%s.png' % (j + 1)
image_data[category].loc[i] = [image_path,1, label] # add new row
i += 1
image_data[category].to_csv(category+"_image_data.csv",index = None, header=False)
#New function create image array by study level
def getImagesInArrayNew(train_dataframe):
images = []
labels = []
for i, data in tqdm(train_dataframe.iterrows()):
img = cv2.imread(data['Path'])
# #random rotation
# angle = random.randint(-30,30)
# M = cv2.getRotationMatrix2D((img_width/2,img_height/2),angle,1)
# img = cv2.warpAffine(img,M,(img_width,img_height))
#resize
img = cv2.resize(img,(img_width,img_height))
img = img[...,::-1].astype(np.float32)
images.append(img)
labels.append(data['Label'])
images = np.asarray(images).astype('float32')
#normalization
mean = np.mean(images[:, :, :])
std = np.std(images[:, :, :])
images[:, :, :] = (images[:, :, :] - mean) / std
labels = np.asarray(labels)
return {'images': images, 'labels': labels}
# #### 3.1.1.1 Variables intialization
img_width, img_height = 224, 224
#Keras ImageDataGenerator to load, transform the images of the dataset
BASE_DATA_DIR = 'data/'
IMG_DATA_DIR = 'MURA-v1.1/'
# ### 3.1.2 XR_ELBOW ImageDataGenertors
# I am going to generate model for every study type and ensemble them. Hence i am preparing data per study type for the model to be trained on.
# +
train_data_dir = BASE_DATA_DIR + IMG_DATA_DIR + 'train/XR_ELBOW'
valid_data_dir = BASE_DATA_DIR + IMG_DATA_DIR + 'valid/XR_ELBOW'
train_datagen = ImageDataGenerator(
rotation_range=30,
horizontal_flip=True
)
test_datagen = ImageDataGenerator(
rotation_range=30,
horizontal_flip=True
)
study_types = ['XR_ELBOW']
create_images_metadata_csv('train',study_types)
create_images_metadata_csv('valid',study_types)
valid_image_df = pd.read_csv('valid_image_data.csv', names=['Path','Count', 'Label'])
train_image_df = pd.read_csv('train_image_data.csv', names=['Path', 'Count','Label'])
dd={}
dd['train'] = train_image_df
dd['valid'] = valid_image_df
valid_dict = getImagesInArrayNew(valid_image_df)
train_dict = getImagesInArrayNew(train_image_df)
train_datagen.fit(train_dict['images'],augment=True)
test_datagen.fit(valid_dict['images'],augment=True)
validation_generator = test_datagen.flow(
x=valid_dict['images'],
y=valid_dict['labels'],
batch_size = 1
)
train_generator = train_datagen.flow(
x=train_dict['images'],
y=train_dict['labels']
)
# -
# ### 3.2 Building a model
# As per the MURA paper, i replaced the fully connected layer with the one that has a single output, after that i applied a sigmoid nonlinearity. In the paper, the optimized weighted binary cross entropy loss. Please see below for the formula,
#
# L(X, y) = -WT,1 * ylog p(Y = 1|X) -WT,0 * (1 - y)log p(Y = 0|X);
# p(Y = 1|X) is the probability that the network assigns to the label i, WT,1 = |NT| / (|AT| + |NT|), and WT,0 = |AT| / (|AT| + |NT|) where |AT|) and |NT|) are the number of abnormal images and normal images of study type T in the training set, respectively.
#
# But i choose to use the default binary cross entropy. The network is configured with Adam using default parameters, batch size of 8, initial learning rate = 0.0001 that is decayed by a factor of 10 each time the validation loss plateaus after an epoch.
# ### 3.2.1 Model paramaters
#model parameters for training
#K.set_learning_phase(1)
nb_train_samples = len(train_dict['images'])
nb_validation_samples = len(valid_dict['images'])
epochs = 10
batch_size = 8
steps_per_epoch = nb_train_samples//batch_size
print(steps_per_epoch)
n_classes = 1
def build_model():
base_model = DenseNet169(input_shape=(None, None,3),
weights='imagenet',
include_top=False,
pooling='avg')
# i = 0
# total_layers = len(base_model.layers)
# for layer in base_model.layers:
# if(i <= total_layers//2):
# layer.trainable = True
# i = i+1
x = base_model.output
predictions = Dense(n_classes,activation='sigmoid')(x)
model = Model(inputs=base_model.input, outputs=predictions)
return model
model = build_model()
#Compiling the model
model.compile(loss="binary_crossentropy", optimizer='adam', metrics=['acc', 'mse'])
#callbacks for early stopping incase of reduced learning rate, loss unimprovement
early_stop = EarlyStopping(monitor='val_loss', patience=8, verbose=1, min_delta=1e-4)
reduce_lr = ReduceLROnPlateau(monitor='val_loss', factor=0.1, patience=1, verbose=1, min_lr=0.0001)
callbacks_list = [early_stop, reduce_lr]
# ### 3.2.2 Training the Model
#train the module
model_history = model.fit_generator(
train_generator,
epochs=epochs,
workers=0,
use_multiprocessing=False,
steps_per_epoch = nb_train_samples//batch_size,
validation_data=validation_generator,
validation_steps=nb_validation_samples //batch_size,
callbacks=callbacks_list
)
model.save("densenet_mura_rs_v3_xr_elbow.h5")
# ### 3.2.3 Visualizing the model
#There was a bug in keras to use pydot in the vis_utils class. In order to fix the bug, i had to comment out line#55 in vis_utils.py file and reload the module
#~/anaconda3/envs/tensorflow_p36/lib/python3.6/site-packages/keras/utils
from keras.utils import plot_model
from keras.utils.vis_utils import *
import keras
import importlib
importlib.reload(keras.utils.vis_utils)
import pydot
plot_model(model, to_file='images/densenet_archi_xr_blow_v3.png', show_shapes=True)
# ### 3.3 Performance Evaluation
# +
#Now we have trained our model, we can see the metrics during the training proccess
plt.figure(0)
plt.plot(model_history.history['acc'],'r')
plt.plot(model_history.history['val_acc'],'g')
plt.xticks(np.arange(0, 10, 1))
plt.rcParams['figure.figsize'] = (8, 6)
plt.xlabel("Num of Epochs")
plt.ylabel("Accuracy")
plt.title("Training Accuracy vs Validation Accuracy")
plt.legend(['train','validation'])
plt.figure(1)
plt.plot(model_history.history['loss'],'r')
plt.plot(model_history.history['val_loss'],'g')
plt.xticks(np.arange(0, 10, 1))
plt.rcParams['figure.figsize'] = (8, 6)
plt.xlabel("Num of Epochs")
plt.ylabel("Loss")
plt.title("Training Loss vs Validation Loss")
plt.legend(['train','validation'])
plt.figure(2)
plt.plot(model_history.history['mean_squared_error'],'r')
plt.plot(model_history.history['val_mean_squared_error'],'g')
plt.xticks(np.arange(0, 10, 1))
plt.rcParams['figure.figsize'] = (8, 6)
plt.xlabel("Num of Epochs")
plt.ylabel("MSE")
plt.title("Training Loss vs Validation Loss")
plt.legend(['train','validation'])
plt.show()
# -
#Now we evaluate the trained model with the validation dataset and make a prediction.
#The class predicted will be the class with maximum value for each image.
ev = model.evaluate_generator(validation_generator, steps=nb_validation_samples, workers=0, use_multiprocessing=False)
ev[1]
#pred = model.predict_generator(validation_generator, steps=1, batch_size=1, use_multiprocessing=False, max_queue_size=25, verbose=1)
validation_generator.reset()
#pred = model.predict_generator(validation_generator,steps=nb_validation_samples)
pred_batch = model.predict_on_batch(valid_dict['images'])
predictions = []
for p in pred_batch:
if(p > 0.5):
predictions+=[1]
else:
predictions+=[0]
error = np.sum(np.not_equal(predictions, valid_dict['labels'])) / valid_dict['labels'].shape[0]
pred = predictions
error
print('Confusion Matrix')
from sklearn.metrics import confusion_matrix, classification_report, cohen_kappa_score
import seaborn as sn
cm = confusion_matrix( pred ,valid_dict['labels'])
plt.figure(figsize = (30,20))
sn.set(font_scale=1.4) #for label size
sn.heatmap(cm, annot=True, annot_kws={"size": 20},cmap="YlGnBu") # font size
plt.show()
print()
print('Classification Report')
print(classification_report(valid_dict['labels'], pred, target_names=["0","1"]))
from sklearn.metrics import confusion_matrix, classification_report, cohen_kappa_score
cohen_kappa_score(valid_dict['labels'], pred)
# ### ROC Curve
from sklearn.metrics import roc_curve
fpr_keras, tpr_keras, thresholds_keras = roc_curve(valid_dict['labels'], pred_batch)
from sklearn.metrics import auc
auc_keras = auc(fpr_keras, tpr_keras)
# +
plt.figure(1)
plt.plot([0, 1], [0, 1], 'k--')
plt.plot(fpr_keras, tpr_keras, label='Keras (area = {:.3f})'.format(auc_keras))
plt.xlabel('False positive rate')
plt.ylabel('True positive rate')
plt.title('ROC curve')
plt.legend(loc='best')
plt.show()
plt.figure(2)
plt.xlim(0.0, 0.2)
plt.ylim(0.65, 0.9)
plt.plot([0, 1], [0, 1], 'k--')
plt.plot(fpr_keras, tpr_keras, label='Keras (area = {:.3f})'.format(auc_keras))
plt.xlabel('False positive rate')
plt.ylabel('True positive rate')
plt.title('ROC curve (zoomed in at top left)')
plt.legend(loc='best')
plt.show()
| src/xr_eblow_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Prudential Life Insurance Assessment
#
# An example of the structured data lessons from Lesson 4 on another dataset.
# +
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
import os
from pathlib import Path
import pandas as pd
import numpy as np
import torch
from torch import nn
import torch.nn.functional as F
from fastai import structured
from fastai.column_data import ColumnarModelData
from fastai.dataset import get_cv_idxs
from sklearn.metrics import cohen_kappa_score
from ml_metrics import quadratic_weighted_kappa
from torch.nn.init import kaiming_uniform, kaiming_normal
# -
PATH = Path('./data/prudential')
PATH.mkdir(exist_ok=True)
# ## Download dataset
# !kaggle competitions download -c prudential-life-insurance-assessment --path={PATH}
for file in os.listdir(PATH):
if not file.endswith('zip'):
continue
# !unzip -q -d {PATH} {PATH}/{file}
train_df = pd.read_csv(PATH/'train.csv')
train_df.head()
# Extra feature engineering taken from the forum
train_df['Product_Info_2_char'] = train_df.Product_Info_2.str[0]
train_df['Product_Info_2_num'] = train_df.Product_Info_2.str[1]
train_df['BMI_Age'] = train_df['BMI'] * train_df['Ins_Age']
med_keyword_columns = train_df.columns[train_df.columns.str.startswith('Medical_Keyword_')]
train_df['Med_Keywords_Count'] = train_df[med_keyword_columns].sum(axis=1)
train_df['num_na'] = train_df.apply(lambda x: sum(x.isnull()), 1)
categorical_columns = 'Product_Info_1, Product_Info_2, Product_Info_3, Product_Info_5, Product_Info_6, Product_Info_7, Employment_Info_2, Employment_Info_3, Employment_Info_5, InsuredInfo_1, InsuredInfo_2, InsuredInfo_3, InsuredInfo_4, InsuredInfo_5, InsuredInfo_6, InsuredInfo_7, Insurance_History_1, Insurance_History_2, Insurance_History_3, Insurance_History_4, Insurance_History_7, Insurance_History_8, Insurance_History_9, Family_Hist_1, Medical_History_2, Medical_History_3, Medical_History_4, Medical_History_5, Medical_History_6, Medical_History_7, Medical_History_8, Medical_History_9, Medical_History_11, Medical_History_12, Medical_History_13, Medical_History_14, Medical_History_16, Medical_History_17, Medical_History_18, Medical_History_19, Medical_History_20, Medical_History_21, Medical_History_22, Medical_History_23, Medical_History_25, Medical_History_26, Medical_History_27, Medical_History_28, Medical_History_29, Medical_History_30, Medical_History_31, Medical_History_33, Medical_History_34, Medical_History_35, Medical_History_36, Medical_History_37, Medical_History_38, Medical_History_39, Medical_History_40, Medical_History_41'.split(', ')
categorical_columns += ['Product_Info_2_char', 'Product_Info_2_num']
cont_columns = 'Product_Info_4, Ins_Age, Ht, Wt, BMI, Employment_Info_1, Employment_Info_4, Employment_Info_6, Insurance_History_5, Family_Hist_2, Family_Hist_3, Family_Hist_4, Family_Hist_5, Medical_History_1, Medical_History_10, Medical_History_15, Medical_History_24, Medical_History_32'.split(', ')
cont_columns += [c for c in train_df.columns if c.startswith('Medical_Keyword_')] + ['BMI_Age', 'Med_Keywords_Count', 'num_na']
train_df[categorical_columns].head()
train_df[cont_columns].head()
train_df = train_df[categorical_columns + cont_columns + ['Response']]
len(train_df.columns)
# ### Convert to categorical
for col in categorical_columns:
train_df[col] = train_df[col].astype('category').cat.as_ordered()
train_df['Product_Info_1'].dtype
train_df.shape
# ### Numericalise and process DataFrame
df, y, nas, mapper = structured.proc_df(train_df, 'Response', do_scale=True)
y = y.astype('float')
num_targets = len(set(y))
# ### Create ColumnData object (instead of ImageClassifierData)
cv_idx = get_cv_idxs(len(df))
cv_idx
model_data = ColumnarModelData.from_data_frame(
PATH, cv_idx, df, y, cat_flds=categorical_columns, is_reg=True)
model_data.trn_ds[0][0].shape[0] + model_data.trn_ds[0][1].shape[0]
model_data.trn_ds[0][1].shape
# ### Get embedding sizes
#
# The formula Jeremy uses for getting embedding sizes is: cardinality / 2 (maxed out at 50).
#
# We reproduce that below:
categorical_column_sizes = [
(c, len(train_df[c].cat.categories) + 1) for c in categorical_columns]
categorical_column_sizes[:5]
embedding_sizes = [(c, min(50, (c+1)//2)) for _, c in categorical_column_sizes]
embedding_sizes[:5]
def emb_init(x):
x = x.weight.data
sc = 2/(x.size(1)+1)
x.uniform_(-sc,sc)
class MixedInputModel(nn.Module):
def __init__(self, emb_sizes, num_cont):
super().__init__()
embedding_layers = []
for size, dim in emb_sizes:
embedding_layers.append(
nn.Embedding(
num_embeddings=size, embedding_dim=dim))
self.embeddings = nn.ModuleList(embedding_layers)
for emb in self.embeddings: emb_init(emb)
self.embedding_dropout = nn.Dropout(0.04)
self.batch_norm_cont = nn.BatchNorm1d(num_cont)
num_emb = sum(e.embedding_dim for e in self.embeddings)
self.fc1 = nn.Linear(
in_features=num_emb + num_cont,
out_features=1000)
kaiming_normal(self.fc1.weight.data)
self.dropout_fc1 = nn.Dropout(p=0.01)
self.batch_norm_fc1 = nn.BatchNorm1d(1000)
self.fc2 = nn.Linear(
in_features=1000,
out_features=500)
kaiming_normal(self.fc2.weight.data)
self.dropout_fc2 = nn.Dropout(p=0.01)
self.batch_norm_fc2 = nn.BatchNorm1d(500)
self.output_fc = nn.Linear(
in_features=500,
out_features=1
)
kaiming_normal(self.output_fc.weight.data)
self.sigmoid = nn.Sigmoid()
def forward(self, categorical_input, continuous_input):
# Add categorical embeddings together
categorical_embeddings = [e(categorical_input[:,i]) for i, e in enumerate(self.embeddings)]
categorical_embeddings = torch.cat(categorical_embeddings, 1)
categorical_embeddings_dropout = self.embedding_dropout(categorical_embeddings)
# Batch normalise continuos vars
continuous_input_batch_norm = self.batch_norm_cont(continuous_input)
# Create a single vector
x = torch.cat([
categorical_embeddings_dropout, continuous_input_batch_norm
], dim=1)
# Fully-connected layer 1
fc1_output = self.fc1(x)
fc1_relu_output = F.relu(fc1_output)
fc1_dropout_output = self.dropout_fc1(fc1_relu_output)
fc1_batch_norm = self.batch_norm_fc1(fc1_dropout_output)
# Fully-connected layer 2
fc2_output = self.fc2(fc1_batch_norm)
fc2_relu_output = F.relu(fc2_output)
fc2_batch_norm = self.batch_norm_fc2(fc2_relu_output)
fc2_dropout_output = self.dropout_fc2(fc2_batch_norm)
output = self.output_fc(fc2_dropout_output)
output = self.sigmoid(output)
output = output * 7
output = output + 1
return output
# +
num_cont = len(df.columns) - len(categorical_columns)
model = MixedInputModel(
embedding_sizes,
num_cont
)
# -
model
from fastai.column_data import StructuredLearner
def weighted_kappa_metric(probs, y):
return quadratic_weighted_kappa(probs[:,0], y[:,0])
learner = StructuredLearner.from_model_data(model, model_data, metrics=[weighted_kappa_metric])
learner.lr_find()
learner.sched.plot()
learner.fit(0.0001, 3, use_wd_sched=True)
learner.fit(0.0001, 5, cycle_len=1, cycle_mult=2, use_wd_sched=True)
learner.fit(0.00001, 3, cycle_len=1, cycle_mult=2, use_wd_sched=True)
# There's either a bug in my implementation, or a NN doesn't do that well at this problem.
| notes/reference/moocs/fast.ai/dl-2018/lesson04-prudential-life-insurance-assessment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Time series Forecasting in Python & R, Part 1 (EDA)
# > Time series forecasting using various forecasting methods in Python & R in one notebook. In the first, part I cover Exploratory Data Analysis (EDA) of the time series using visualizations and statistical methods.
#
# - toc: true
# - badges: true
# - comments: true
# - categories: [forecasting,R,Python,rpy2,altair]
# - hide: false
# ## Overview
#
# This is a quarterly sales data of a French retail company from Prof. <NAME>'s ["Forecasting Methods & Applications"](https://robjhyndman.com/forecasting/) book. I have uploaded the data to my [github](https://github.com/pawarbi/blog/tree/master/data).The goals for this first part are:
#
# 1. Exploratory data analysis of the time series
# 2. Explain the time series behaviour in qualitative and quantitative terms to build intuition for model selection
# 3. Identify the candidate models and possible model parameters that can be used based on the findings in the EDA
# ### Importing libraries
#
# +
#collapse-hide
#Author: <NAME>
#Version: 1.0
#Date Mar 27, 2020
import pandas as pd
import numpy as np
import itertools
#Plotting libraries
import matplotlib.pyplot as plt
import seaborn as sns
import altair as alt
plt.style.use('seaborn-white')
# %matplotlib inline
#statistics libraries
import statsmodels.api as sm
import scipy
from scipy.stats import anderson
from statsmodels.tools.eval_measures import rmse
from statsmodels.tsa.stattools import adfuller
from statsmodels.graphics.tsaplots import month_plot, seasonal_plot, plot_acf, plot_pacf, quarter_plot
from statsmodels.tsa.seasonal import seasonal_decompose
from statsmodels.tsa.holtwinters import ExponentialSmoothing, SimpleExpSmoothing
from statsmodels.stats.diagnostic import acorr_ljungbox as ljung
#from nimbusml.timeseries import SsaForecaster
from statsmodels.tsa.statespace.tools import diff as diff
import pmdarima as pm
from pmdarima import ARIMA, auto_arima
from scipy import signal
from scipy.stats import shapiro
from scipy.stats import boxcox
from sklearn.preprocessing import StandardScaler
#library to use R in Python
import rpy2
from rpy2.robjects import pandas2ri
pandas2ri.activate()
import warnings
warnings.filterwarnings("ignore")
np.random.seed(786)
# -
#
# >Note: I have found that results could be significanlty different if you use different versions of the libraries, especially with statsmodels. If you want to reproduce these results, be sure to use the same versions of these libraries. For this project, I created a conda virtual environment as rpy2 requires specific versions of Pandas & certain R libraries
#
# +
#Printing library versions
print('Pandas:', pd.__version__)
print('Statsmodels:', sm.__version__)
print('Scipy:', scipy.__version__)
print('Rpy2:', rpy2.__version__)
# -
# +
#collapse-hide
# Define some custom functions to help the analysis
def MAPE(y_true, y_pred):
"""
%Error compares true value with predicted value. Lower the better. Use this along with rmse(). If the series has
outliers, compare/select model using MAPE instead of rmse()
"""
y_true, y_pred = np.array(y_true), np.array(y_pred)
return np.mean(np.abs((y_true - y_pred) / y_true)) * 100
def residcheck(residuals, lags):
"""
Function to check if the residuals are white noise. Ideally the residuals should be uncorrelated, zero mean,
constant variance and normally distributed. First two are must, while last two are good to have.
If the first two are not met, we have not fully captured the information from the data for prediction.
Consider different model and/or add exogenous variable.
If Ljung Box test shows p> 0.05, the residuals as a group are white noise. Some lags might still be significant.
Lags should be min(2*seasonal_period, T/5)
plots from: https://tomaugspurger.github.io/modern-7-timeseries.html
"""
resid_mean = np.mean(residuals)
lj_p_val = np.mean(ljung(x=residuals, lags=lags)[1])
norm_p_val = jb(residuals)[1]
adfuller_p = adfuller(residuals)[1]
fig = plt.figure(figsize=(10,8))
layout = (2, 2)
ts_ax = plt.subplot2grid(layout, (0, 0), colspan=2);
acf_ax = plt.subplot2grid(layout, (1, 0));
kde_ax = plt.subplot2grid(layout, (1, 1));
residuals.plot(ax=ts_ax)
plot_acf(residuals, lags=lags, ax=acf_ax);
sns.kdeplot(residuals);
#[ax.set_xlim(1.5) for ax in [acf_ax, kde_ax]]
sns.despine()
plt.tight_layout();
print("** Mean of the residuals: ", np.around(resid_mean,2))
print("\n** Ljung Box Test, p-value:", np.around(lj_p_val,3), "(>0.05, Uncorrelated)" if (lj_p_val > 0.05) else "(<0.05, Correlated)")
print("\n** <NAME>era Normality Test, p_value:", np.around(norm_p_val,3), "(>0.05, Normal)" if (norm_p_val>0.05) else "(<0.05, Not-normal)")
print("\n** <NAME>, p_value:", np.around(adfuller_p,3), "(>0.05, Non-stationary)" if (adfuller_p > 0.05) else "(<0.05, Stationary)")
return ts_ax, acf_ax, kde_ax
def accuracy(y1,y2):
accuracy_df=pd.DataFrame()
rms_error = np.round(rmse(y1, y2),1)
map_error = np.round(np.mean(np.abs((np.array(y1) - np.array(y2)) / np.array(y1))) * 100,1)
accuracy_df=accuracy_df.append({"RMSE":rms_error, "%MAPE": map_error}, ignore_index=True)
return accuracy_df
def plot_pgram(series,diff_order):
"""
This function plots thd Power Spectral Density of a de-trended series.
PSD should also be calculated for a de-trended time series. Enter the order of differencing needed
Output is a plot with PSD on Y and Time period on X axis
Series: Pandas time series or np array
differencing_order: int. Typically 1
"""
#from scipy import signal
de_trended = series.diff(diff_order).dropna()
f, fx = signal.periodogram(de_trended)
freq=f.reshape(len(f),1) #reshape the array to a column
psd = fx.reshape(len(f),1)
# plt.figure(figsize=(5, 4)
plt.plot(1/freq, psd )
plt.title("Periodogram")
plt.xlabel("Time Period")
plt.ylabel("Amplitude")
plt.tight_layout()
# -
# ### Importing Data
# +
path = 'https://raw.githubusercontent.com/pawarbi/datasets/master/timeseries/ts_frenchretail.csv'
#Sales numbers are in thousands, so I am dividing by 1000 to make it easier to work with numbers, especially squared errors
data = pd.read_csv(path, parse_dates=True, index_col="Date").div(1_000)
data.index.freq='Q'
data.head()
# -
# >Note: I have explicitly set the index frequency to *quarterly*. This makes plotting and analyzing data with pandas and statsmodels easier. Many methods in Statsmodels have `freq` argument. Setting the frequency explicitly will pass the value automatically. More date offsets can be found in Pandas [documentation here](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html). `freq='Q-DEC'` in the `index.freq` below shows quarterly data ending in December. Other advantage of setting the `.freq` value is that if the dates are not continous, Pandas will throw an error, which can be used to fix the data quality error and make the series continuos. Other common date offsets are:
# - Monthly Start: `'MS'`
# - Quarterly Start: `'QS'`
# - Weekly: `'W'`
# - Bi Weekly: `'2W'`
# - Business/ Weekday: `'B'`
# - Hourly: `'H'`
#
data.index
# ### Train Test Split:
#
# Before analyzing the data, first split it into train and test (hold-out) for model evaluation. All the EDA and model fitting/selection should be done first using train data. Never look at the test sample until later to avoid any bias. Typically we want at least 3-4 full seasonal cycles for training, and test set length should be no less than the forecast horizon.
#
# In this example, we have 24 observations of the quarterly data, which means 6 full cycles (24/4). Our forecast horizon is 4 quarters. So training set should be more than 16 and less than 20. I will use first 18 observations for training and keep last 6 for validation.Note that I am always selecting the last 6 values for test by using `.iloc[:-6]`. As we get more data, this will ensure that last 6 values are always for validation. Unlike typical train/test split, we can not shuffle the data before splitting to retain the temporal structure.
#
# #### Cross-validation:
# Data can be split using the above method or using cross-validation where the series is split into number of successive segments and the model is tested using one-step ahead forecast. Model accuracy in that case is based on the mean of the cross-validation errors over the number of splits used. This minimizes chances of overfitting. Be sure to include at least 1-2 seasonal periods to capture the seasonality. e.g. in this case, the first training set of the CV should be min 8 values so the model has captured seasonal behaviour from 2 years. This is the preferred method when the time series is short.
#
# Our series has 24 obervations so I can use last 6-8 for validation. When the typical train/test split is used, always check the sensisitivity of the model performance and model parameters to train/test size. If AIC or AICc is used for model evaluation, it approximatley approaches cross-validation error asymptotically. I will cover this in Part 2 with the code and example.
#
# 
# +
#Split into train and test
train = data.iloc[:-6]
test = data.iloc[-6:]
#forecast horizon
h = 6
train_length = len(train)
print('train_length:',train_length, '\n test_length:', len(test) )
# -
train.head()
# ## Exploratory Data Analysis & Modeling Implications
#
# These are some of the questions I ask at various stages of model building.
#
# 1. **Are there any null values? how many? best way to impute the null data?**
# - If null/NaNs are present, first identify why the data is missing and if NaNs mean anything. Missing values can be
# filled by interpolation, forward-fill or backward-fill depending on the data and context. Also make sure null doesnt mean 0, which is acceptable but has modeling implications.
# - It's important to understand how the data was generated (manual entry, ERP system), any transformations, assumptions were made before providing the data.
#
#
# 2. **Are the data/dates continuous?**
# - In this exmaple I am only looking at continous time-series. There other methods that deal with non-continuous data. ETS & ARIMA require the data to be continuous. If the series is not continuous, we can add dummy data or use interpolation.
#
#
# 3. **Are there any duplicate dates, data?**
# - Remove the duplicates or aggregate the data (e.g. average or mean) to treat duplicates
#
#
# 4. **Any 'potential' outliers?**
# - Outliers are defined as observations that differ significantly from the general observations. Identify if the data is
# susceptible to outliers/spikes, if outliers mean anything and how to define outliers. While 'Outlier Detection' is a topic in itself, in forecasting context we want to treat outliers before the data is used for fitting the model. Both
# ETS and ARIMA class of models (especially ARIMA) are not robust to outliers and can provide erroneous forecasts. Data should be analyzed while keeping seasonality in mind. e.g. a sudden spike could be because of the seasonal behaviour and not be outlier. Do not confuse outlier with 'influential data'.
#
# - Few ways to treat outliers:
# - Winsorization: Use Box and whiskers and clip the values tha exceed 1 & 99th percentile (not preferred)
# - Use residual standard deviation and compare against observed values (preferred but can't do *a priori*)
# - Use moving average to check spikes/troughs (iterative and not robust)
#
# - Another important reason to pay close attention to outliers is that we will choose the appropriate error metric based
# on that. There are many error metrics used to assess accuracy of forecasts, *viz.* MAE, MSE, RMSE, %MAPE, %sMAPE. If
# outliers are present, don't use RMSE because the squaring the error at the outlier value can inflate the RMSE. In that
# case model should be selected/assessed using %MAPE or %sMAPE. More on that in part 2.
#
#
# 5. **Visually any trend, seasonality, cyclic behaviour?**
# - This will help us choose the appropriate model (Single, Double, Triple Exponential Smoothing, ARIMA/SARIMA)
# - If cyclic behiour is present (seasonality is short-order variation e.g. month/quarter, cyclicity occurs over 10-20
# years e.g. recession) we will need to use different type of decomposition (X11, STL). Depending on the context and
# purpose of analysis, seasoanlity adjustment may also be needed.
# - If multiple seasonalities are present, ETS or ARIMA cannot be used. SSA, TBATS, harmonic regression are more
# appropriate in that case. FB Prophet can also help with multiple seasonalities.
# - Frequency of seasonality is important. ETS & SARIMAX are not appropriate for high frequency data such as hourly, daily, sub-daily and even weekly. Consider using SSA,TBTAS, FB Prophet, deep learning models.
#
#
# 6. **How does the data change from season to season for each period and period to period and compared to the level?**
# - Does it increas/decrease with the trend? Changes slowly, rapidly or remains constant. This is an important
# observation to be made, especially for ETS model, as it can determine the parametrs to be used & if any preprocessing
# will be needed.
# - De-compose the series into level, trend, seasonal components and residual error. Observe the patterns in the decomposed series.
# - Is the trend constant, growing/slowing linearly or exponentially or some other non-linear function?
# - Is the seasonal pattern repetitive?
# - How is the seasonal pattern changing relative to level? If it is constant relative to level, it shows "additive" seasonality, whereas if it is growing, it's "multiplicative".(Part 2 covers this in detail)
#
#
# 7. **Distribution of the data? will we need any transformations?**
# - While normally distributed data is not a requirement for forecasting and doesnt necessarily improve point forecast accuracy, it can help stablize the variance and narrow the prediction interval.
# - Plot the histogram/KDE for each time period (e.g. each year and each seasona) to get gauge peakedness, spread in the data. It can also help compare different periods and track trends over time.
# - If the data is severely skewed, consider normalizing the data before training the model. Be sure to apply inverse transformation on the forecasts. *Use the same transformation parameters on the train and test sets.* Stabilizing the variance by using Box Cox transformation (special case being log & inverse transform), power law etc can help more than normalizing the data.
# - Watch out for outliers before transformation as it will affect the transformation
# - Plottng distribution also helps track "concept-drift" in the data, *i.e.* does the underlying temporal structure / assumption change over time. If the drift is significant, refit the model or at least re-evaluate. This can be tricky in time series analysis.
# - Uncertainty in the training data will lead to higher uncertainty in the forecast. If the data is highly volatile/uncertain (seen by spread in the distribution, standard deviation, non-constant variance etc), ETS and ARIMA models will not be suitable. Consider GARCH and other methods.
#
#
# 8. **Is the data stationary? Is this a white noise, random walk process?**
#
# - Perhaps the most important concept to keep in mind when doing time series analysis and forecasting is that, time series is a *probabilistic / stochastic process*, and the time series we are analyzing is a *'realization of a stochastic process'*. A time signal could be deterministic or stochastic/probabilistic. In a deterministic process, the future values can be predicted exactly with a mathematical function e.g. y = sin(2$\pi$ft). In our case, the future values can only be expressed in terms of probability distribution. The point estimates are mean/median of the distribution. By definition, the mean has a distribution around it and as such the stakeholders should be made aware of the probabilistic nature of the forecast through uncertainty estimates.
#
# - ***Stationarity***: Statistical stationarity means the time series has constant mean, variance and autocorrelation is insignificant at all lags. Autocorrelation is a mouthful, all it means is the correlation with its past self. e.g. to check if two variables are linearly correlated with each other, we calculate their coeff of correlation (Pearson correlation). Similarly, *autocorrelation* does the same thing but with its past values (i.e lags). More on that later. For a stationary time series, the properties are the same no matter which part of the series (w.r.t time) we look at. This is a core concept of the ARIMA methods, as only stationary processes can be modeled using ARIMA. ETS can handle non-stationary processes.
# - ***White Noise***: If a time series has zero mean and a constant variance , i.e. N(0,$\sigma^2$), it's a white noise. The variables in this case are *independent and identically distributed* (i.i.d) and are uncorrelated. We want the residuals left after fitting the model to be a white noise. White noise can be identified by using ADFuller test and plotting autocorrelation function (ACF) plots. In an ACF plot, the autocorrelation should be insignificant (inside the 95% CI band) at all lags.
# - ***Random Walk***: Random walks are non-stationary. Its mean or variance or both change over time. Random walk cannot be forecast because we have more unknowns than the data so we will end up having way too many parameters in the model. In essence, random walk has no pattern to it, it's last data point plus some random signal (drift). Thus, if the first difference of the time series results in a white noise, it's an indication of a Random Walk. e.g. most equity stocks are random walk but by looking at percent difference (%growth over time) we can study the white noise.
# `Next Data point = Previous Data Point + Random Noise`
# `Random Noise = Next Data Point - Previous Data Point`
#
#
#
# 
#
# >Note: It's easy to mistake randomness for seasonality. In the random walk chart below, it can appear that the data has some seasonality but does not!
# +
#collapse-hide
#create white noise with N(0,1.5), 500 points
np.random.seed(578)
steps = np.random.normal(0,1,500)
noise = pd.DataFrame({"x":steps})
wnoise_chart = alt.Chart(noise.reset_index()).mark_line().encode(
x='index',
y='x').properties(
title="White Noise")
#Create random walk with N(0,1.5), 500 points
steps[0]=0
rwalk = pd.DataFrame({"x":100 + np.cumsum(steps)}).reset_index()
rwalk_chart = alt.Chart(rwalk).mark_line().encode(
x='index',
y=alt.Y('x', scale=alt.Scale(domain=(80,150)))).properties(
title="Random Walk")
wnoise_chart | rwalk_chart
# -
# 9. **Auto-correlation? at what lag?**
# - Study the second order properties (autocorrelation and power spectral density) of the time series along with mean, standard deviation, distribution. More details below.
#
#
# 10. **If trend is present, momentum or mean-reversing?**
# - Time series with momentum indicates the value tends to keep going up or down (relative to trend) depending on the immediate past. Series with mean-reversion indicates it will go up (or down) if it has gone down (or up) in the immediate past. This can be found by examining the coefficients of the ARIMA model. This provides more insight into the process and builds intuition. This doesnt not directly help with forecasting.
#
# 11. **Break-points in the series?**
# - Are there any structural breaks (shifts) in the series. Structural breaks are abrupt changes in the trend. Gather more information about the sudden changes. If the breaks are valid, ETS/ARIMA models wont work. FB Prophet, dynamic regression, deep learning models, adding more features might help. Identify the possible reasons for change, e.g. change in macros, price change, change in customer preferenaces etc. Note structural change persists for some time, while outliers do not. Break points are different from non-stationarity. Read more [here](http://faculty.baruch.cuny.edu/smanzan/FINMETRICS/_book/time-series-models.html#structural-breaks-in-time-series) for examples & explanations. In case of structural break-points, consider modeling the segments of the series separately.
# 
#
#
# 12. **Intermittent demand?**
# - Time series is said to be intermittent when there are several 0 and small values (not nulls) in the series. ETS and ARIMA are not appropriate for this type of time series. It's a common pattern with inventory time series, especially for new items. Croston's method is one approach to use for forecasting intermittent demand.
# - When demand is intermittent, use RMSE rather than %MAPE as the evaluation metric. With %MAPE, the denominator would be 0 leading to erroneous results.
#
#collapse-hide
#creating intermittent demand plot
demand = [10, 12, 0, 3,50,0,0,18,0,4, 12,0,0,8,0,3]
demanddf = pd.DataFrame({'y': demand, 'x': np.arange(2000, 2016) } )
alt.Chart(demanddf).mark_bar().encode(
x='x',
y='y').properties(
title="Example: Intermittent Demand", width = 700)
#
# 13. **Do we need any exogenous variables/external regressors?**
# - It may be necessary to include additional features/variables to accurately capture the time series behaviour. For example, the sales for a retailer might be higher on weekends, holidays, when it's warmer etc. This is different from seasonal pattern. In such cases, using the 'day of the week' or 'is_holiday' feature might provide better forecast. ETS models cannot use exogenous variable. SARIMAX (X is for exogenous), deep learning, XGB models are more suited.
# - Always inspect the residuals after fitting the model. If the residuals are correlated (use ACF/PACF plots, Ljung Box test on residuals), it's an indication that we are not capturing the time series behaviour accurately and could try adding exogenous behaviour.
#
#
# 14. **Are the stakeholders interested in forecast for invidiuals periods or hierarchical forecast?**
# - Typically forecasts are made for individual periods, e.g in this example, we are interested in the forecasts for next 4 quarters. But it's possible that the business leaders might be more interested in the 1 year forecast rather than 4 quarters. We could combine the forecasts from 4 quarters to calculate the forecast for 1 year, but that would be incorrect. As mentioned above, time series is a statistical process with probability distribution. We can get reasonable value by summing the mean forecasts but the uncertainty around those forecasts cannot be added. Also, if the individual forecasts are based on the median (rather than mean), forecasts cannot be added. We will need to calculate "hierarchical forecast* by simulating the future paths and then adding the distributions to get prediction interval.
#
#
# 15. **Are forecast *explainability* & *interpretability* important?**
# - Many traditional and statistical forecasting methods such as ETS, SARIMA are easy to apply, interprete and model parameters/results can help explain the time series behaviour. This can be important in scenarios where such insights can help make business decisions. e.g if an ETS model with damped trend fits better, it can be an indication of slowing growth etc. However, many other models such deep learning, RNN, S2S, LSTM etc are blackbox approaches that may lead to higher accuracy but provide little-to-no explainability.
#
# ## EDA in Pyhton
# ### Data Integrity / Quality
#Any missing data?
print("missing_data:", train.isna().sum())
print("unique dates:", train.index.nunique())
#Counting number of values for each quarter and Year. Columsn are quarters.
#Here each qquarter and year has 1 value, thus no duplicates
pd.crosstab(index=train.index.year, columns=train.index.quarter)
# #### Observations:
# 1. No null values
# 2. Length of the train set is 18 and we have 12 unique dates/quarters so no duplicate dates
# 3. Each year and quarter has 1 observation, so no duplicates and data is continuous
# ### Time Series
# Plotting the time series and the 4 quarter rolling mean using Altair.
# >Tip: Matplotlib and Seaborn create static charts, whereas plots created with Altair are interactive. You can hover over the data points to read tooltips. The most useful feature is the ability to zoom-in and out. Time series data can be dense and it's important to check each time period to get insights. With zoom-in/out, it can be done iteractively without slicing the time series. Altair's [documentation and example library](https://altair-viz.github.io/user_guide/interactions.html) is great.
# +
#collapse-hide
#Create line chart for Training data. index is reset to use Date column
train_chart=alt.Chart(train.reset_index()).mark_line(point=True).encode(
x='Date',
y='Sales',
tooltip=['Date', 'Sales'])
#Create Rolling mean. This centered rolling mean
rolling_mean = alt.Chart(train.reset_index()).mark_trail(
color='orange',
size=1
).transform_window(
rolling_mean='mean(Sales)',
frame=[-4,4]
).encode(
x='Date:T',
y='rolling_mean:Q',
size='Sales'
)
#Add data labels
text = train_chart.mark_text(
align='left',
baseline='top',
dx=5 # Moves text to right so it doesn't appear on top of the bar
).encode(
text='Sales:Q'
)
#Add zoom-in/out
scales = alt.selection_interval(bind='scales')
#Combine everything
(train_chart + rolling_mean +text).properties(
width=600,
title="French Retail Sales & 4Q Rolling mean ( in '000)").add_selection(
scales
)
# -
# #### Sub-series plot
#
# Sub-series plot to show how the series behaves each year in all seasons (quarterly or monthly)
#collapse-hide
alt.Chart(train.reset_index()).mark_line(point=True).encode(
x='quarter(Date)',
y='Sales',
column='year(Date)',
tooltip=['Date', 'Sales']).properties(
title="Sales: Yearly Subseries plot",
width=100).configure_header(
titleColor='black',
titleFontSize=14,
labelColor='blue',
labelFontSize=14
)
#box plot to see distribution of sales in each year
fig, ax = plt.subplots(figsize = (12,8))
sns.boxplot(data=train, x=train.index.year, y = 'Sales', ax = ax, boxprops=dict(alpha=.3));
sns.swarmplot(data=train, x=train.index.year, y = 'Sales');
# #%Growth each year. Excluding 2016 since we have only 2 quarters
growth = train[:'2015'].groupby(train[:'2015'].index.year)["Sales"].sum().pct_change()
growth*100
# #### Observations:
# 1. Sales has gone up each year from 2012-2015 =>Positive Trend present.
# 2. Typically, Sales goes up from Q1 to Q3, peaks in Q3, drops in Q4. Definitely a seasoanl pattern. => Model should capture seasonality and trend.
# 2. Just comparing Q4 peaks, sales has gone up from \\$432K to \\$582K => Trend exists, Model should capture trend. No cyclic behaviour
# 3. Overall data looks clean, no observations outside of IQR => Clean data, no outliers
# 4. No structural breaks, intermittent pattern => ETS and SARIMA may be used
# 5. Notice that the length of the bar in box plot increases from 2012-2015. => Mean & variance increasing, we will need to stabilize the variance by taking log or using Box Cox transform
#
# ### Quarterly trends & distrbution
# Quarterly sub-series plot to see how the series behaves in each quarter across alll years.
#collapse-hide
alt.Chart(train.reset_index()).mark_line(point=True).encode(
x='year(Date)',
y='Sales',
column='quarter(Date)',
tooltip=['Date', 'Sales']).properties(
title="Sales: Quarterly Subseries plot",
width=100).configure_header(
titleColor='black',
titleFontSize=14,
labelColor='blue',
labelFontSize=14
)
# >Tip: Statsmodels has a `quarter_plot()` method that can be used to create similar chart easily.
#Quarterly plot: Shows trend for Q1-Q4 for each of the years. Red line shows mean
quarter_plot(train);
# ##### Distribution of Sales in each year
# +
#collapse-hide
#Distribution plot of each year compared with overall distribution
sns.distplot(train, label='Train', hist=False, kde_kws={"color": "g", "lw": 3, "label": "Train","shade":True})
sns.distplot(train['2012'], label='2012', hist=False)
sns.distplot(train['2013'], label='2013', hist=False)
sns.distplot(train['2014'], label='2014', hist=False)
sns.distplot(train['2015'], label='2015', hist=False);
# -
# In this case the heatmap feels redundant but when the series is long, heatmap can reveal more patterns
#collapse-hide
sns.heatmap(pd.pivot_table(data=train, index=train.index.year, columns=train.index.quarter),
square=True,
cmap='Blues',
xticklabels=["Q1", "Q2", "Q3", "Q4"]);
# Visualizing the quarterly sales for each year as %
# +
#collapse-hide
#As stacked bar chart, in % values.
stack1= alt.Chart(train[:'2015'].reset_index()).mark_bar().encode(
x=alt.X('sum(Sales)'),
y='year(Date):N',
color=alt.Color(
'quarter(Date)',
scale=alt.Scale(scheme='category10')),
tooltip=["Date", "Sales"]).properties(
height=100,
width = 300,
title = "Sum of Sales by each Quarter")
stack2= alt.Chart(train[:'2015'].reset_index()).mark_bar().encode(
x=alt.X('sum(Sales)', stack='normalize'),
y='year(Date):N',
color=alt.Color(
'quarter(Date)',
scale=alt.Scale(scheme='category10')),
tooltip=["Date", "Sales"]
).properties(
height=100,
width = 300,
title = "Sum of Sales as % by each Quarter")
stack1 | stack2
# -
pie= train[:'2015'].groupby(train[:'2015'].index.quarter)["Sales"].sum().plot.bar(
title="Total Sales by Quarter 2012-2015", legend=True, label="Sales each Quarter")
# ##### Seasonality Factor
#
# This will help us understand how much each quarter contributes relative to the average demand. Note that this should be done on a de-trended series(taking first difference) but because we don't have enough data and for a quick demonstration, I am using the series as is.
#
# +
#Groupby Sales by Quarter
#Only use upto 2015 because we have partial data for 2016
train_2015=train[:'2015']
avg_2015= np.int(train[:'2015'].mean())
#Avg sales per quarter
qrt_avg=train_2015.groupby(train_2015.index.quarter)["Sales"].mean()
#Groupby quarter
qrt_table = pd.pivot_table(train_2015, index=train_2015.index.quarter, columns=train_2015.index.year)
#add qrt_avg to qrt_table
qrt_table["avg"] = qrt_avg
#Additive Seasonality Factor: Subtract mean from avg column
qrt_table["additive"] = qrt_table["avg"]-avg_2015
#Multiplicative Seasonality Factor: Subtract mean from avg column
qrt_table["multiplicative"] = (qrt_table["avg"]/avg_2015).round(2)
qrt_table.index.name="Quarters"
qrt_table
# -
# #### Observations:
# 1. Quarter plot & heatmap confirm peak in Q3, drop in Q4.
# 2. For each of the years the upward trend observed in all quarters
# 3. Kenel Density plot shows data looks normally distributed, bi-modal distribution in quarters is because of small sample size. Peaks shift right from 2012 to 2015 indicating increase in average.
# 4. Distribution becomes fatter as the years progress, indicating higher spread/variation (as seen above in boxplot too)
# 5. Though The sales peak in Q3 each year, as a % of annual sales, all quarters contribute roughly the same
# 6. Seasonal factor analysis shows that in 3rd quarter we see that sales jump up by 15% (or $73K) relative to average while in other three quarters it drops by 1-7%. This is great for intuitive understanding of series behaviour. Another key takeaway from this analysis is that sales is not stable, as is evident from the charts above, multiplicative seasonality would capture the pattern better than additive seasonality. This insight will come handy when we create HW/ETS model (part 2). We could also reduce the variance by taking the log so errors are additive.
# ### Decomposition
#
# We will de-compose the time series into trend, seasonal and residuals
# >Tip: *Always* use a semicolon (;) after plotting any results from statsmodels. For some reason if you don't, it will print the plots twice. Also, by default the statsmodels plots are small and do not have a `figsize()` argument. Use `rcParams()` to define the plot size
decompose = seasonal_decompose(train["Sales"])
decompose.plot();
plt.rcParams['figure.figsize'] = (12, 8);
# #### Observations:
#
# 1. Trend is more than linear, notice a small upward take off after 2013-07. Also notice that trend is projecting upward .
# 2. Seasonal pattern is consistent
# 3. Resduals are whetever is left after fitting the trend and seasonal components to the observed data. It's the component we cannot explain. We want the residuals to be *i.i.d* (i.e uncorrelated). If the residuals have a pattern, it means there is still some structural information left to be captured. Residuals are showing some wavy pattern, which is not good. Let's perform Ljung Box test to confirm if they are *i.i.d as a group*.
# 4. We do not want to see any recognizable patterns in the residuals, e.g. waves, upward/downward slope, funnel pattern etc.
#
ljung_p = np.mean(ljung(x=decompose.resid.dropna())[1]).round(3)
print("Ljung Box, p value:", ljung_p, ", Residuals are uncorrelated" if ljung_p>0.05 else ", Residuals are correlated")
# Residuals are uncorrelated. If the residuals are correlated, we can perform transformations to see if it stabilizes the variance. It's also an indication that we may need to use exogenous variable to fully explain the time series behaviour or use higher order models. In this case, the residuals are uncorrelated so that's good.
# >Note: Ljung Box test tests the residuals as a group. Some residuals may have significant lag but as a group, we want to make sure they are uncorrelated.
# ### Second Order Properties of the time series
# We study the second order properties to understand -
# - is the data stationary
# - is the data white noise, random walk? i.e are the lags correlated?
# - quantify seasonal/cyclic behviour
#
#
# #### Stationarity:
# For the series to be stationary, it must have:
# - constant mean
# - constant variance
# - constant covariance (uncorrelated)
#
# We verify this by observing change in mean, variance, autocorrelation and with a statistical test (ADFuller test)
# ##### Is the mean constant?
train.plot(figsize=(12,6), legend=True, label="Train", cmap='gray')
train["Sales"].rolling(4, center=False).mean().plot(legend=True, label="Rolling Mean 4Q");
print("Mean is:", train["Sales"].mean())
# Notice that each year, the difference between the mean and the max in Q3 increases. This can potentially mean multiplicative seasonality.
# ##### Is the variance constant?
train["Sales"].rolling(4).std().plot(legend=True, label="Rolling Std Deviation 4Q");
print("S.D is:", train["Sales"].std().round(1))
# Both mean and standard deviation are increasing, thus not stationary.
# #### Coefficient of Variation:
# Coefficient of variation gives us an idea about the variability in the process, especially when looking at sales and demand. Note that this should be used for relative comparison and does not have a strict statistical defition. It's very common measure in demand planning and inventory analytics.
#
# c.v = s.d/mean
#
# If C.V<0.75 => Low Variability
#
# If 0.75<C.V<1.3 => Medium Variability
#
# If C.V>1.3 => High Variability
#
#
cv = train["Sales"].std()/train["Sales"].mean()
cv.round(2)
# This is a low-variability process.
# ##### Is the covariance constant?
#
#Plot ACF and PACF using statsmodels
plot_acf(train);
plot_pacf(train);
# *ADFuller Test for stationarity*
#
# Augmented Dicky Fuller test is a statistical test for stionarity. If the p value is less than 0.05, the series is stationary, otherwise non-stationary. Use `adfuller()` from statsmodels
#Calculate ad fuller statistic
adf = adfuller(train["Sales"])[1]
print(f"p value:{adf.round(4)}", ", Series is Stationary" if adf <0.05 else ", Series is Non-Stationary")
# #### Observations:
# 1. ACF: ACF plot shows autocorrelation coeff is insignificant at all lag values (within the blue 95%CI band), except lag 1. When studying ACF plots, we look at these 4 things
# - *Are any lags significant*, i.e outside the blue band of 95% CI. If they are, the series is correlated with itself at those lags. Note there is 5% chance that the lag shown as insignificant (ACF=0) is shows as significant. In our case, 1st lag is barely significant, indicating sales last quarter affect the sales this quarter.
# - *How quickly do the bar lenghts change*: If the bars are taping down, that shows presence of trend. Our series has a trend
# - *Pattern*: If the ACF shows up/down repeating pattern, it means seasonality with size equal to length of repetition.
# - *Sign of ACF*: Alternating signs in ACF shows mean-reversing process whereas if all the ACs are positive (or negative), it shows momentum process.
#
# Properties of ACF help us determine the order of the MA process. More on that in part 2.
#
# 2. PACF: Partial autocorrelation, similar to partial correlation, shows correlation after *'partialing out'* previous lags. If a series has PACF significant at lag k, it means controlling for other lags <k, lag k has a significant correlation. PACF plot is used to determine order of AR process.
#
# 2. ADFuller test shows that the series is not stationary. We can try to make it stationary by differencing it.
#De-trending
de_trended = train.diff(1).dropna()
adf2 = adfuller(de_trended)[1]
print(f"p value:{adf2}", ", Series is Stationary" if adf2 <0.05 else ", Series is Non-Stationary")
de_trended.plot();
# By taking the first difference we de-trended the series and it has become stationary!
# *Autocovariance function* vs *autocorrelation fucntion*
# The autocovariance measures the linear dependence between two points on the same series observed at different times. Very smooth series exhibit autocovariance functions that stay large even when the t and s are far apart, whereas choppy series tend to have autocovariance functions that are nearly zero for large separations. autocorrelation functions (ACF) measures the predictability (linear), and is the normalized autocovariance. ACF, just like a correlation coeff, is between [-1,1] and is easier to interprete. Both measure linear dependence between random variables.
#
# For example, the scatterplot below shows the train["Sales"] plotted against it's first lag. We can see a linear, although weak, relationship between them. Use `pandas.plotting.lag_plot()`
# *Lag plot, 1st lag*
pd.plotting.lag_plot(train["Sales"],1);
# *Lag plot, 2nd lag* : Weak relationship
pd.plotting.lag_plot(train["Sales"],2);
# >Note: If you wish to study the lags, you can obtain it by `.shift()` method.
sns.scatterplot(train["Sales"], train["Sales"].shift(-1));
# >Important: Use statsmodels for calculating ACF and NOT pandas `pd.Series.autocorr()`
# Statsmodels and R use mean differencing, i.e subtract the mean of the *entire* series before summing it up, whereas Pandas uses Pearson correlation to calculate the ACF. Pearson correlation uses mean of the *subseries* rather than the entire series. For a long time series, the difference between the two should be negligible but for a short series, the diffrenece could be significant. In most cases, we are more interested in the pattern in the ACF than the actual values so, in a practical sense either would work. But, to be consistent and accurate use statsmodels to calculate and plot the ACF.
#
# ##### Which frequencies are prominent?
#
# We typically look at the time series in the time domain. But, we can also analyze the time series in the frequency domain. It's based on the assumption that it is made up of sine and cosine waves of different frequencies. This helps us detect periodic component of known/unknown frequencies. It can show additional details of the time series that can be easily missed. We do it with a *Periodogram* and *Power Spectral Density* plot.
# *Periodogram*: We analyze frequency and associated intensity of frequency. Note that below I have invrted the frequency to obtain periods `Period,T = 1/frequency`. For example, a monthly time series has 12 seasonal periods, so we would obtain frequency = 1/12 = 0.0833. In our example, we expect to see the intensity to be high at period=4
# >Tip: Periodogram should be plotted for a de-trended time series. Time series can be obtained by differencing the series
plot_pgram(train["Sales"],1);
# *Power Spectral Density*: Periodogram assumes the frequencies to be harmonics of the fundamental frequency, whereas PSD allows the frequency to vary contunuously. PSD is calculated using autocovariance function (ACF seen above). Spectral density is the amount of variance per frequency interval. PSD shows the eaxct same information of the time series as the ACF, just in the frequency domain. We rarely use PSD for business time series analysis. Plot below shows that lower frequency content dominates the time series. If it had another bump at higher frequency, that would indicate cyclic behaviour. Sometime it can be easier to figure out MA vs AR process by looking at the PSD plot.
#Plot PSD
plt.psd(train["Sales"], detrend='linear');
plt.title("PSD Plot");
# #### Is the series Normal?
# As mentioned above, series does not have to be Gaussian for accurate forecasting but if the data is highly skewed it can affect the model selection and forecast uncertainty. In general, if the series is non-gaussian, it should be normalized before any further transformations (differencing, log, Box Cox) at least to check if normalization helps. Normalization will also help if decide to use regression, tree-based models, NN models later. Note that with normalization, we make the z score between [0,1], with *standardization* on the other hand , we center the distribution with mean =0, s.d.=1.
#
# Normality can be checked visually by plotting the density plot, q-q plot and Shapiro-Wilk test.
#Distribution Plot
sns.distplot(train["Sales"]);
#Q-Q Plot
sm.qqplot(train["Sales"], fit=True, line='45', alpha=0.5, dist='norm' );
#Jarque Bera Stastical Test for Normality
from scipy.stats import jarque_bera as jb
is_norm=jb(train["Sales"])[1]
print(f"p value:{is_norm.round(2)}", ", Series is Normal" if is_norm >0.05 else ", Series is Non-Normal")
# ##### Observations:
# 1. Q-Q plot shows the data follows the 45deg line very closely, deviates slightly in the left tail.
# 2. <NAME> test shows the data is from Normal distribution
# ## Summary
# Model explainability is as important as model accuracy. We will keep above insights in mind when choosing, fitting, evaluating and selecting various models. We will choose the model that we can explain based on above insights. It is important to be able to explain the time series behavour in qualitative and quantitative terms.
#
# In Part 2, I will cover model fitting, selection and ensemble forecasting
#
# ## EDA in R
# [Forecasting Principles and Practice](https://otexts.com/fpp2/) by Prof. Hyndmand and Prof. Athanasapoulos is the best and most practical book on time series analysis. Most of the concepts discussed in this blog are from this book. Below is code to run the `forecast()` and `fpp2()` libraries in Python notebook using rpy2
# +
import rpy2
import warnings
warnings.filterwarnings('ignore')
from rpy2.robjects import pandas2ri
import rpy2.rinterface as rinterface
pandas2ri.activate()
# %load_ext rpy2.ipython
# + magic_args="-i data,train -o r_train" language="R"
#
# library(fpp2)
#
# r_train <- ts(train$Sales, start=c(2012,01), frequency=4)
# r_train %>% autoplot() + ggtitle("French Retail") +xlab("Year-Quarter")+ylab("Retail Sales")
#
# -
# %Rpush r_train
# + magic_args="-i r_train" language="R"
#
# r_train %>% ggsubseriesplot()
# -
# ##### Lag Plots
# + language="R"
#
# r_train %>% gglagplot()
# -
# ##### ACF Plots
# + language="R"
#
# r_train %>% ggAcf()
# -
# R shows 3 lags to be significant, whereas in Python we saw only the first lag to be significant. I am not sure why.
# Just to confirm, I am did the analysis in JMP statistical software which I use at work for statistical analysis. Below are the results. It matches with Python's results - 1st lag to be significant, spectral density plot matches too.
# 
# ##### Outlier detection
# + magic_args="-o olier" language="R"
# olier <- r_train %>% tsoutliers()
# -
print(olier, end="")
# using `tsoutliers()` does not return any results, showing there are no statistical outliers
# ## Summary
# Here is a summary of what we have learned about this time series:
# 1. There are no null values, outliers or duplicate values in the series. Series is continuous, non-intermittent. No structural breaks. We don't have to do any cleaning.
# 2. Series has a trend. Sales have increased every year. It looks more than linear but less than exponential. We might need to try lof or BoxCox transform.
# 3. Series has seasonality with seasonal periods = 4. Peak in Q3, drops in Q4, ramps up from Q1 to Q3. No other dominant periods. No cyclic behaviour.
# 4. Avg sales per quarter is 497, and S.D is 111, low variability. We will use this to guage the model error relative to mean and S.D.
# 5. Since it is not a high-frequency data and has fixed seasonality, we can use ETS and SARIMA class of models
# 6. SARIMA could be used. We will need at least 1 differencing as de-trended series was stationary
# 7. Mean, variance, covariance are not constant. Series is not stationary, not white noise and not random-walk.
# 8. Variance increasing with time. Highs also look to be increasing relative to mean (rolling avg). 'multiplicative' seasonality might fit better in ETS model
# 9. Series is normally distributed
# 10. 1st lag was significant, though barely (ACF~0.5)
# 11. We do not have any information on exogenous variables
# 12. As there are no outliers and series is not intermittent, we can use RSME for model evaluation
# 13. We never looked at the test data. All the EDA and model building must be done using the training set. Intuitively, given what we have observed in the training set, we expect the forecast to be on the upward trend with slightly mupltiplicative seasonality.
#
# ### References:
# 1. Forecasting: Principles and Practice, by <NAME>
# 2. Time Series Analysis and its Applications, by <NAME>
# 3. Time Series Analysis and Forecasting, by Montgomery & Jennings
# 4. Introduction to Time Series and Analysis, by Brockwell
# 5. Practial Time Series Forecasting with R, by <NAME>
| _notebooks/2020-04-21-timeseries-part1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Ex2 - Labels Datacube
#
# This notebook provides a quick demo on how to create a datacube of training labels to complement a datacube of imagery. The training label datacube is critical for supervised machine learning. Having the ability to ingest labels in a datacube format provides flexibility for ML engineers to train models easily without worrying about underlying remote sensing image formats, which can be sometimes daunting for non experts.
import os
from pathlib import Path
import glob
import numpy as np
import pandas as pd
import rasterio
from icecube.bin.labels_cube.create_json_labels import CreateLabels
import icecube
# Currently as of version 1.0, `icecube` provides the ability to ingest labels that are both vector and in raster format. These two formats should cover almost all of the use cases needed for training machine learning models for supervised training. Vector labels (e.g. bounding boxes, polygons) are useful for training object detectors while raster images are often (but not always) leveraged for segmentation wokflows.
# set paths here.
icecube_abspath = str(Path(icecube.__file__).parent.parent)
resource_dir = os.path.join(icecube_abspath, "tests/resources")
grd_dir = os.path.join(resource_dir, "grd_stack/")
vector_labels_save_fpath = os.path.join(icecube_abspath, "icecube/dataset/temp/dummy_vector_labels.json")
raster_labels_save_fpath = os.path.join(icecube_abspath, "icecube/dataset/temp/dummy_raster_labels.json")
cube_save_path = os.path.join(icecube_abspath, "icecube/dataset/temp/my_awesome_labels_cube.nc")
Path(str(Path(vector_labels_save_fpath).parent)).mkdir(parents=True, exist_ok=True)
grd_fpaths = glob.glob(grd_dir+"*")
# ## Creating JSON Labels
#
# In order to populate labels in `icecube`, labels must be converted to a specific JSON structure. The section walks one through the script that can be used to create such labels.
# ### 1. Example with vector labels
# Let's first go through an example where we will use the assets in `tests/resources` to demonstrate the example with vector labels.
# The below example showcase ingesting bounding boxes inside datacubes as labels.
# For the examples below, we will use assets inside `tests/resources/grd_stack/*.tif`
# +
# let's create some random bounding boxes for dummy training sample
random_classes = ["rand-a", "rand-b", "rand-c"]
def create_random_bboxes(N, I):
random_bboxes = []
I_shape = I.shape
for _, i in enumerate(range(N)):
xmin, ymin = np.random.randint(0,I_shape[0]), np.random.randint(0,I_shape[1])
xmax, ymax = np.random.randint(xmin, I_shape[0]), np.random.randint(ymin,I_shape[1])
random_bboxes.append([xmin, ymin, xmax, ymax])
return random_bboxes
# +
# For demo purposes, we will generate some random samples for bounding boxes.
labels_collection = []
for grd_fpath in grd_fpaths:
grd_values = rasterio.open(grd_fpath).read(1)
grd_product = rasterio.open(grd_fpath).tags()["PRODUCT_FILE"]
bboxes_seq = create_random_bboxes(np.random.randint(30,45), grd_values)
for each_bbox in bboxes_seq:
labels_collection.append([grd_product, each_bbox[0],
each_bbox[1],
each_bbox[2],
each_bbox[3],
random_classes[np.random.randint(0,3)]])
labels_df = pd.DataFrame(labels_collection, columns=["file_name", "xmin", "ymin", "xmax", "ymax", "class"])
labels_df.head(5)
# +
# Now we can easily convert these bounding boxes to icecube-friendly JSON strucutre using `CreateLabels` class
create_labels = CreateLabels("vector")
for i, df_row in labels_df.T.iteritems():
product_labels_seq = []
product_name = df_row.iloc[0]
# instance label contains bounding box for a unit label.
instance_label = {"xmin": df_row.iloc[1],
"ymin": df_row.iloc[2],
"xmax": df_row.iloc[3],
"ymax": df_row.iloc[4],
}
class_name = df_row.iloc[5]
# product_labels_seq contains sequence of WKT geom vectors
product_labels_seq.append(
create_labels.create_instance_bbox(class_name, instance_label)
)
create_labels.populate_labels(str(product_name), product_labels_seq)
create_labels.write_labels_to_json(vector_labels_save_fpath, ensure_ascii=True)
# -
# Here is a glimpse at how labels look like for a single image in the stack:
create_labels.labels_collection[0]
# Please note that we can follow similar structure for other WKT geometries like Polygons, Points and easily create JSON files with labels. Sample vector labels can be found under `tests/resources/labels/dummy_vector_labels.json` for reference.
# ### 2. Example with Raster Labels
#
# Similar to above example with vector labels, we can ingest rasters as segmentation labels too. This example quickly highlights the workflow to create icecube-friendly raster JSON structure.
# Creating JSON structure for raster labels is relatively straightforward. We simply maintain a dictionary where key represents the product-file (or the image) and value represents the raster as label.
# For this example, we will use the sample masks inside `tests/resources`
# +
masks_dir = os.path.join(resource_dir, "masks/")
raster_dir = os.path.join(resource_dir, "grd_stack/")
raster_names = [
"ICEYE_GRD_SLED_54549_20210427T215124_hollow_10x10pixels_fake_0.tif",
"ICEYE_GRD_SLED_54549_20210427T215124_hollow_10x10pixels_fake_1.tif",
"ICEYE_GRD_SLED_54549_20210427T215124_hollow_10x10pixels_fake_2.tif",
]
masks_names = [
"ICEYE_GRD_SLED_54549_20210427T215124_hollow_10x10pixels_fake_0.png",
"ICEYE_GRD_SLED_54549_20210427T215124_hollow_10x10pixels_fake_1.png",
"ICEYE_GRD_SLED_54549_20210427T215124_hollow_10x10pixels_fake_2.png",
]
masks_fpaths = [os.path.join(masks_dir, fpath) for fpath in masks_names]
# Create a dictionary where key:value pair represents raster:mask
raster_mask_dict = {}
for raster_name, mask_fpath in zip(raster_names, masks_fpaths):
raster_mask_dict[raster_name] = mask_fpath
create_labels = CreateLabels("raster")
for product_name, mask_fpath in raster_mask_dict.items():
seg_mask = create_labels.create_instance_segmentation(mask_fpath)
create_labels.populate_labels(product_name, seg_mask)
create_labels.write_labels_to_json(raster_labels_save_fpath)
# -
# This is how our JSON file looks like for raster labels.
#
# `/home/user/runner/` simply indicates the local filepath of rasters
#
# ```
# [
# {
# "product_file": "ICEYE_GRD_SLED_54549_20210427T215124_hollow_10x10pixels_fake_0.tif",
# "labels": {
# "segmentation": "/home/user/runner/icecube/tests/resources/masks/ICEYE_GRD_SLED_54549_20210427T215124_hollow_10x10pixels_fake_0.png"
# }
# },
# {
# "product_file": "ICEYE_GRD_SLED_54549_20210427T215124_hollow_10x10pixels_fake_1.tif",
# "labels": {
# "segmentation": "/home/user/runner/icecube/tests/resources/masks/ICEYE_GRD_SLED_54549_20210427T215124_hollow_10x10pixels_fake_1.png"
# }
# },
# {
# "product_file": "ICEYE_GRD_SLED_54549_20210427T215124_hollow_10x10pixels_fake_2.tif",
# "labels": {
# "segmentation": "/home/user/runner/icecube/tests/resources/masks/ICEYE_GRD_SLED_54549_20210427T215124_hollow_10x10pixels_fake_2.png"
# }
# }
# ]
# ```
# ## Populating Datacubes with Labels
#
# Once we have create the icecube formatted JSON structure either for vector geometries or for raster labels, it is fairly straightforward to convert them to an `xr.Dataset` or append them to an already created `xr.Dataset`
# First thing first, some imports
from icecube.bin.labels_cube.labels_cube_generator import LabelsDatacubeGenerator
from icecube.bin.config import CubeConfig
from icecube.bin.datacube import Datacube
# +
# Let's create a Datacube from our labels.json file. For demo purposes, we will use only vector labels.
config_dir = os.path.join(resource_dir, "json_config/")
default_config_fpath = os.path.join(config_dir, "config_use_case_default.json")
raster_dir = os.path.join(resource_dir, "grd_stack")
dummy_vector_labels_fpath = os.path.join(
resource_dir, "labels/dummy_vector_labels.json"
)
cc = CubeConfig()
product_type = "GRD"
cc.load_config(default_config_fpath)
labels_datacube = LabelsDatacubeGenerator.build(
cc, product_type, dummy_vector_labels_fpath, raster_dir
)
labels_datacube.to_file(cube_save_path)
# -
# And that was it, we have a labels datacube generated! 🎉
# .
# For inspecting the elements of labels datacube, it is recommended to convert associated `xr.Dataset` to the `Datacube` core class, as it provides ready methods to process datacubes. More details on the `Datacube` core class be found in demo notebook: **[Ex4_Datacube](https://iceye-ltd.github.io/icecube/examples/Ex4_Datacube)**
# We can see that returned object is an instance of class VectorLabels
type(labels_datacube)
# We can throw `labels_datacube.xrDataset` to Datacube core class to easily access useful operations on the cube
dc = Datacube().set_xrdataset(labels_datacube.xrdataset)
print(dc.get_data_variables())
# Finally we can easily see what is inside our datacube easily for one of our products
labels_xrarray = dc.get_xrarray("Labels")
POI = dc.get_all_products(labels_xrarray)[0]
print("Associated labels with product-file: {} are \n".format(POI))
dc.get_product_values(POI, labels_xrarray)
# Great, our vector geometries are preserved inside datacube. Similarly we can show it for our raster labels. We will leave it as an exercise for you to get your hands dirty on the code for that part.
# **Happy Coding :)**
| docs/examples/Ex2_LabelsDatacube.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
class_1=['<NAME>','<NAME>','<NAME>','<NAME>']
class_1
class_2=['<NAME>','<NAME>','<NAME>']
class_2
new_class=class_1+class_2
new_class
new_class.append('<NAME>')
new_class
new_class.remove("<NAME>")
new_class
courses={"Math":65,"English":70,"History":80,"French":70,"Science":60}
courses
#[] for calling dict values
courses["Math"]
total=sum(courses.values())
total
percentage=(total/500)*100
percentage
mathematics={"<NAME>":78,"<NAME>":95,"<NAME>":65,
"<NAME>":50,
"<NAME>":70,
"<NAME>":66,
"<NAME>":75}
mathematics
topper=max(mathematics, key=mathematics.get)
topper
topper.split(" ")
first_name=topper.split()[0]
last_name=topper.split()[1]
# +
#first_name=topper[0:6]
# +
#last_name=topper[7:9]
# -
first_name
last_name
full_name=first_name+" "+last_name
full_name
certificate_name=full_name.upper()
certificate_name
| Student_Management_System/Student_M_S(GA).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py3]
# language: python
# name: conda-env-py3-py
# ---
# Notebook to Test gsw TEOS conversion instead of Matlab
# +
import gsw
import matplotlib.pyplot as plt
import netCDF4 as nc
from salishsea_tools import gsw_calls
from salishsea_tools import LiveOcean_BCs as nancy
# %matplotlib inline
# -
sal_matlab = nc.Dataset('/results/forcing/LiveOcean/boundary_conditions/LiveOcean_v201712_y2018m11d08.nc')
sal_gsw = nc.Dataset('single_LO_y2018m11d09.nc')
fig, axs = plt.subplots(1, 3, figsize=(15, 5))
mesh0 = axs[0].pcolormesh(sal_matlab['vosaline'][0, :, 0, :], vmin=29, vmax=34)
mesh1 = axs[1].pcolormesh(sal_gsw['vosaline'][0, :, 0, :], vmin=29, vmax=34)
mesh2 = axs[2].pcolormesh(sal_gsw['vosaline'][0, :, 0, :] - sal_matlab['vosaline'][0, :, 0, :], cmap='bwr',
vmax=1.6, vmin=-1.6)
fig.colorbar(mesh0, ax=axs[0])
fig.colorbar(mesh1, ax=axs[1])
fig.colorbar(mesh2, ax=axs[2])
for ax in axs:
ax.invert_yaxis()
fig, axs = plt.subplots(1, 3, figsize=(15, 5))
mesh0 = axs[0].pcolormesh(sal_matlab['vosaline'][0, 1, :, :])
mesh1 = axs[1].pcolormesh(sal_gsw['vosaline'][0, 1, :, :])
mesh2 = axs[2].pcolormesh(sal_gsw['vosaline'][0, 1, :, :] - sal_matlab['vosaline'][0, 1, :, :], cmap='bwr',
)
fig.colorbar(mesh0, ax=axs[0])
fig.colorbar(mesh1, ax=axs[1])
fig.colorbar(mesh2, ax=axs[2])
for ax in axs:
ax.invert_yaxis()
# # Dig Deeper
depBC, lonBC, latBC, shape = nancy.load_SalishSea_boundary_grid(imin=376-1, imax=470, rim=10,
meshfilename='/home/sallen/MEOPAR/grid/mesh_mask201702.nc')
# +
# Create metadeta for temperature and salinity (Live Ocean variables, NEMO grid)
var_meta = {'vosaline': {'grid': 'SalishSea2',
'long_name': 'Practical Salinity',
'units': 'psu'},
'votemper': {'grid': 'SalishSea2',
'long_name': 'Potential Temperature',
'units': 'deg C'},
'NO3': {'grid': 'SalishSea2',
'long_name': 'Nitrate',
'units': 'muM'},
'Si': {'grid': 'SalishSea2',
'long_name': 'Nitrate',
'units': 'muM'},
'OXY': {'grid': 'SalishSea2',
'long_name': 'Oxygen',
'units': 'muM'},
'DIC': {'grid': 'SalishSea2',
'long_name': 'Dissolved Inorganic Carbon',
'units': 'muM'},
'TA': {'grid': 'SalishSea2',
'long_name': 'Total Alkalinity',
'units': 'muM'},
}
# Mapping from LiveOcean TS names to NEMO TS names
LO_to_NEMO_var_map = {'salt': 'vosaline',
'temp': 'votemper',
'NO3': 'NO3',
'Si': 'Si',
'oxygen': 'OXY',
'TIC': 'DIC',
'alkalinity': 'TA',}
# -
date = '2018-11-08'
d = nancy.load_LiveOcean(date)
interps = nancy.interpolate_to_NEMO_depths(d, depBC, var_names=(var for var in LO_to_NEMO_var_map if var != 'Si'))
sal_ref = gsw.SR_from_SP(interps['salt'])
mat_sal_ref = gsw_calls.generic_gsw_caller('gsw_SR_from_SP.m', [
interps['salt'][:],
])
mat_sal_ref.shape
fig, axs = plt.subplots(1, 3, figsize=(15, 5))
mesh0 = axs[0].pcolormesh(mat_sal_ref[1, :, :], vmax=32, vmin=30.5)
mesh1 = axs[1].pcolormesh(sal_ref[1, :, :])
mesh2 = axs[2].pcolormesh(sal_ref[1, :, :] - mat_sal_ref[1, :, :], cmap='bwr')
fig.colorbar(mesh0, ax=axs[0])
fig.colorbar(mesh1, ax=axs[1])
fig.colorbar(mesh2, ax=axs[2])
for ax in axs:
ax.invert_yaxis()
temp_cons = gsw.CT_from_pt(sal_ref[:], interps['temp'])
mat_temp_cons = gsw_calls.generic_gsw_caller(
'gsw_CT_from_pt.m', [
mat_sal_ref,
interps['temp'] ,
]
)
fig, axs = plt.subplots(1, 3, figsize=(15, 5))
mesh0 = axs[0].pcolormesh(mat_temp_cons[1, :, :], vmax=7., vmin=11.7)
mesh1 = axs[1].pcolormesh(temp_cons[1, :, :], vmax=7., vmin=11.7)
mesh2 = axs[2].pcolormesh(temp_cons[1, :, :] - mat_temp_cons[1, :, :], cmap='bwr')
fig.colorbar(mesh0, ax=axs[0])
fig.colorbar(mesh1, ax=axs[1])
fig.colorbar(mesh2, ax=axs[2])
for ax in axs:
ax.invert_yaxis()
interps['salt'] = sal_ref
interps['temp'] = temp_cons
interps = nancy.remove_south_of_Tatoosh(interps)
mat_interps = interps.copy()
mat_interps['salt'] = mat_sal_ref
mat_interps['temp'] = mat_temp_cons
mat_interps = nancy.remove_south_of_Tatoosh(mat_interps)
interps = nancy.fill_box(interps)
mat_interps = nancy.fill_box(mat_interps)
fig, axs = plt.subplots(1, 3, figsize=(15, 5))
mesh0 = axs[0].pcolormesh(mat_interps['salt'][1, :, :], vmax=32, vmin=30.5)
mesh1 = axs[1].pcolormesh(interps['salt'][1, :, :], vmax=32, vmin=30.5)
mesh2 = axs[2].pcolormesh(interps['salt'][1, :, :] - mat_interps['salt'][1, :, :], cmap='bwr',)
fig.colorbar(mesh0, ax=axs[0])
fig.colorbar(mesh1, ax=axs[1])
fig.colorbar(mesh2, ax=axs[2])
for ax in axs:
ax.invert_yaxis()
sigma = gsw.sigma0(interps['salt'][:], interps['temp'][:] )
mat_sigma = gsw_calls.generic_gsw_caller('gsw_sigma0.m', [mat_interps['salt'][:], mat_interps['temp'][:] ])
fig, axs = plt.subplots(1, 3, figsize=(15, 5))
mesh0 = axs[0].pcolormesh(mat_sigma[1, :, :],vmax=24.5, vmin=23.2)
mesh1 = axs[1].pcolormesh(sigma[1, :, :], vmax=24.5, vmin=23.2)
mesh2 = axs[2].pcolormesh(sigma[1, :, :] - mat_sigma[1, :, :], cmap='bwr')
fig.colorbar(mesh0, ax=axs[0])
fig.colorbar(mesh1, ax=axs[1])
fig.colorbar(mesh2, ax=axs[2])
for ax in axs:
ax.invert_yaxis()
sigma, interps = nancy.convect(sigma, interps)
mat_sigma, mat_interps = nancy.convect(mat_sigma, mat_interps)
fig, axs = plt.subplots(1, 3, figsize=(15, 5))
mesh0 = axs[0].pcolormesh(mat_sigma[1, :, :],vmax=24.5, vmin=23.2)
mesh1 = axs[1].pcolormesh(sigma[1, :, :], vmax=24.5, vmin=23.2)
mesh2 = axs[2].pcolormesh(sigma[1, :, :] - mat_sigma[1, :, :], cmap='bwr')
fig.colorbar(mesh0, ax=axs[0])
fig.colorbar(mesh1, ax=axs[1])
fig.colorbar(mesh2, ax=axs[2])
for ax in axs:
ax.invert_yaxis()
interps = nancy.extend_to_depth(interps)
mat_interps = nancy.extend_to_depth(mat_interps)
fig, axs = plt.subplots(1, 3, figsize=(15, 5))
mesh0 = axs[0].pcolormesh(mat_interps['salt'][1, :, :], vmax=32, vmin=30.5)
mesh1 = axs[1].pcolormesh(interps['salt'][1, :, :], vmax=32, vmin=30.5)
mesh2 = axs[2].pcolormesh(interps['salt'][1, :, :] - mat_interps['salt'][1, :, :], cmap='bwr')
fig.colorbar(mesh0, ax=axs[0])
fig.colorbar(mesh1, ax=axs[1])
fig.colorbar(mesh2, ax=axs[2])
for ax in axs:
ax.invert_yaxis()
interpl = nancy.interpolate_to_NEMO_lateral(interps, d, lonBC, latBC, shape)
interpl['salt'].shape
mat_interpl = nancy.interpolate_to_NEMO_lateral(mat_interps, d, lonBC, latBC, shape)
fig, axs = plt.subplots(1, 3, figsize=(15, 5))
mesh0 = axs[0].pcolormesh(mat_interpl['salt'][1, :, :],)
mesh1 = axs[1].pcolormesh(interpl['salt'][1, :, :], )
mesh2 = axs[2].pcolormesh(interpl['salt'][1, :, :] - mat_interpl['salt'][1, :, :], cmap='bwr')
fig.colorbar(mesh0, ax=axs[0])
fig.colorbar(mesh1, ax=axs[1])
fig.colorbar(mesh2, ax=axs[2])
for ax in axs:
ax.invert_yaxis()
| notebooks/LiveOcean/TEOS-10, matlab to python gsw.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# Example is taken from François Chollet book: Deep Learning with Python Original code can be found in https://github.com/fchollet/deep-learning-with-python-notebooks/blob/master/6.1-one-hot-encoding-of-words-or-characters.ipynb
# You need to install keras for this example to work. Run following command in the command shell.
#
# conda install -c conda-forge keras
# +
import keras
keras.__version__
# -
# # Using word embeddings
#
#
# Another popular and powerful way to associate a vector with a word is the use of dense "word vectors", also called "word embeddings".
# While the vectors obtained through one-hot encoding are binary, sparse (mostly made of zeros) and very high-dimensional (same dimensionality as the
# number of words in the vocabulary), "word embeddings" are low-dimensional floating point vectors
# (i.e. "dense" vectors, as opposed to sparse vectors).
# Unlike word vectors obtained via one-hot encoding, word embeddings are learned from data.
# It is common to see word embeddings that are 256-dimensional, 512-dimensional, or 1024-dimensional when dealing with very large vocabularies.
# On the other hand, one-hot encoding words generally leads to vectors that are 20,000-dimensional or higher (capturing a vocabulary of 20,000
# token in this case). So, word embeddings pack more information into far fewer dimensions.
# 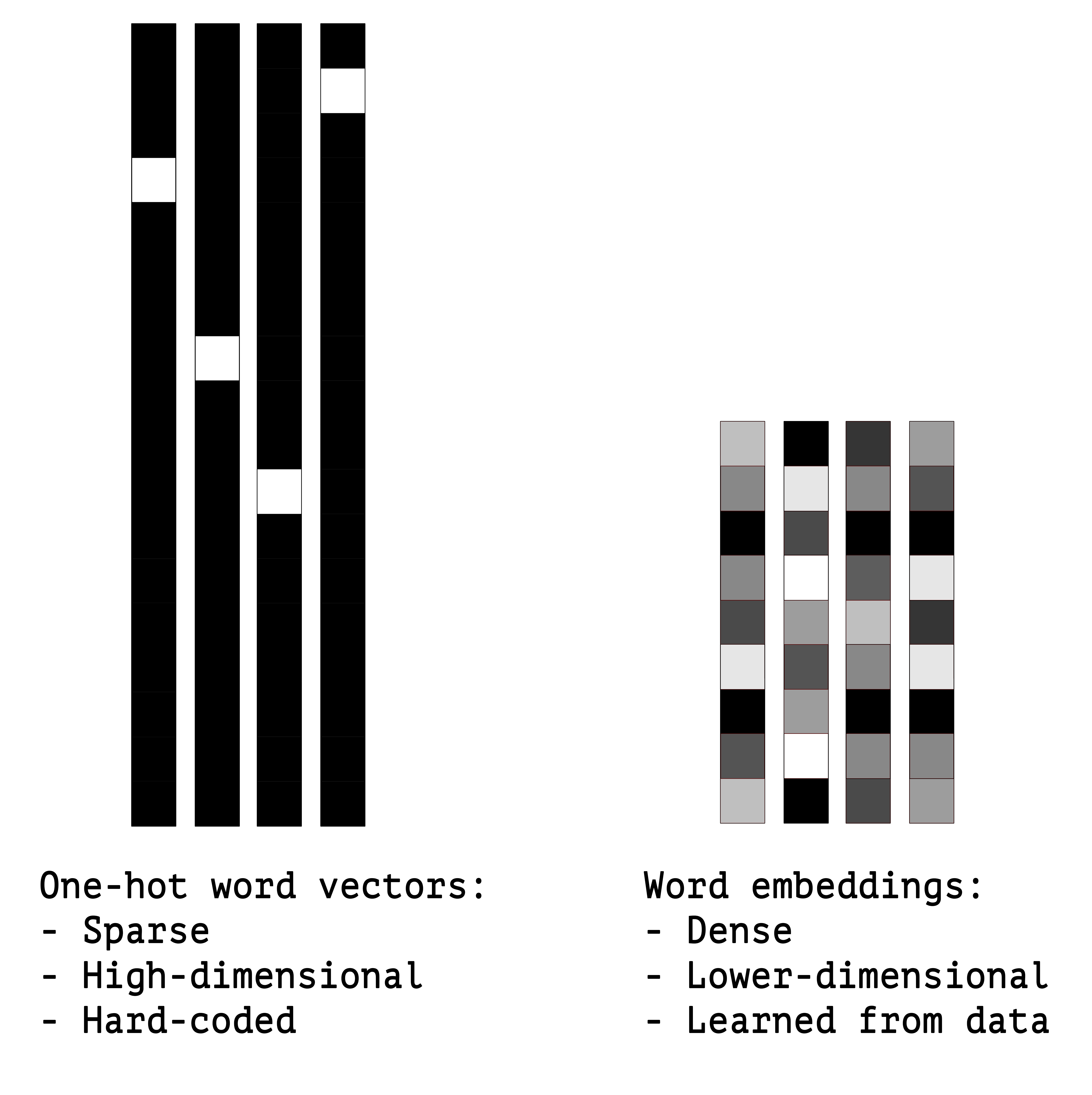
# There are two ways to obtain word embeddings:
#
# * Learn word embeddings
# * Load into your model, pre-trained word embeddings
# ## Learning word embeddings with the `Embedding` layer
#
#
# +
from keras.layers import Embedding
# The Embedding layer takes at least two arguments:
# the number of possible tokens, here 1000 (1 + maximum word index),
# and the dimensionality of the embeddings, here 64.
embedding_layer = Embedding(1000, 64)
# +
from keras.datasets import imdb
from keras import preprocessing
# Number of words to consider as features
max_features = 10000
# Cut texts after this number of words
# (among top max_features most common words)
maxlen = 20
# Load the data as lists of integers.
(x_train, y_train), (x_test, y_test) = imdb.load_data(num_words=max_features)
# This turns our lists of integers
# into a 2D integer tensor of shape `(samples, maxlen)`
x_train = preprocessing.sequence.pad_sequences(x_train, maxlen=maxlen)
x_test = preprocessing.sequence.pad_sequences(x_test, maxlen=maxlen)
# +
from keras.models import Sequential
from keras.layers import Flatten, Dense
model = Sequential()
# We specify the maximum input length to our Embedding layer
# so we can later flatten the embedded inputs
model.add(Embedding(10000, 8, input_length=maxlen))
# After the Embedding layer,
# our activations have shape `(samples, maxlen, 8)`.
# We flatten the 3D tensor of embeddings
# into a 2D tensor of shape `(samples, maxlen * 8)`
model.add(Flatten())
# We add the classifier on top
model.add(Dense(1, activation='sigmoid'))
model.compile(optimizer='rmsprop', loss='binary_crossentropy', metrics=['acc'])
model.summary()
# -
history = model.fit(x_train, y_train,
epochs=10,
batch_size=32,
validation_split=0.2)
# +
import matplotlib.pyplot as plt
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(acc) + 1)
# +
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
# +
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# -
| notebooks/keras-fchollet-using-word-embeddings-learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install -q catalyst==20.12 nltk torchtext==0.8.0 captum
# # Seminar
#
# Hi! Today we starts NLP section in our course. Starting with embeddings and Recurrent Neural Networks.
# +
from catalyst.utils import set_global_seed, get_device
set_global_seed(42)
device = get_device()
# -
# ## Text preprocessing
#
# Text preprocessing is the most important part of NLP. In comparison, an image is usually reshaped and normalized in a preprocessing pipeline. But a text is different. A text consists of words(or tokens), that has a different probability to be written. Words are arrays of characters, and different arrays can be related to one word(E.g. "it" and "It" or "Имя" and "Имени" is one word, but different word form.). That's why texts should be normalized and tokenized.
example = "Hello! My name is <unk> and i'm <unk>."
lower = example.lower()
print(lower)
# +
from nltk.tokenize import WordPunctTokenizer
tokenizer = WordPunctTokenizer()
tokens = tokenizer.tokenize(lower)
print(tokens)
# -
# Another part of the preprocessing is filtration. Every token has to be informative. Punctuation hasn't much information, and it should be deleted. Pronouns, prepositions, articles (and other small words) should be deleted too. Usually, they will not help to solve tasks.
# +
from string import punctuation
filtered = [
token for token in tokens
if ((len(token) >= 3) and (token not in punctuation))
]
print(filtered)
# -
# Last part of preprocessing is adding special tokens. They means begining(`SOS`) or ending(`EOS`) of text/sentences, words out of vocabulary(`UNK`), padding for batching(`PAD`). A nueral networks can have other special tokens. For BERT some tokens should be masked. These tokens are swapped with `MASK` token.
# +
SOS = "<SOS>" # or <SOT>/<BOT>/<BOS>
EOS = "<EOS>" # so on...
PAD = "<PAD>"
UNK = "<UNK>"
# Sometimes
MASK = "<MASK>" # Masked Langueage Models
# -
# ## Text Classification
#
# The most popular task in NLP is text classification. Before 2012, this task is solved by pair of Tf-Idf method and some classification model. But now we have embeddings vector, mapped from tokens to some big continious high dimensions real space. Read more about Embeddings: [NLP course for you](https://lena-voita.github.io/nlp_course/word_embeddings.html).
#
# For text classification we will use mean of embeddings for each text as a feature vector. Let's code this!
# +
import torch
from torchtext.vocab import GloVe
from torchtext.datasets import IMDB
embedding_glove = GloVe(name='glove.twitter.27B.25d', dim=300, unk_init=torch.Tensor.normal_)
train_dataset, test_dataset = IMDB()
vocab = train_dataset.get_vocab()
vocab.load_vectors(embedding_glove)
# -
# Each text has different length, and we will build a batch by adding padding tokens at the end of the text. Effective way to do it by bucketing. However, it's not so easy to implement. So we will create simple padding.
# +
from typing import Tuple
from torch.utils.data import DataLoader
PAD_ID = vocab.stoi['<pad>']
def collate_fn(
batch: Tuple[torch.Tensor, torch.Tensor]
) -> Tuple[torch.Tensor, torch.Tensor]:
texts = []
max_len = max(t.size(0) for _, t in batch)
labels = torch.zeros(len(batch))
for idx, (label, txt) in enumerate(batch):
new_txt = torch.zeros((1, max_len)) + PAD_ID
new_txt[0, : txt.size(0)] = txt
texts.append(new_txt)
labels[idx] = label
return torch.cat(texts).type(torch.LongTensor), labels
batch_size = 256
loaders = {
"train": DataLoader(
train_dataset,
batch_size=batch_size,
shuffle=True,
drop_last=True,
collate_fn=collate_fn,
),
"valid": DataLoader(
test_dataset,
batch_size=batch_size,
shuffle=False,
drop_last=False,
collate_fn=collate_fn,
),
}
# -
# Our algorithm is this:
# - Get embeddings for each word
# - Get mean vector for text
# - Classify text by mean vector
#
# Let's code this.
# +
import torch.nn as nn
from catalyst.contrib.nn import Lambda
class EmbeddingModel(nn.Module):
def __init__(
self,
vocab_size: int,
embedding_size: int = 300,
hidden_size: int = 150,
dropout_p: float = 0.2,
):
super().__init__()
self.embedding = nn.Embedding(vocab_size, embedding_size)
self.out = nn.Sequential(
Lambda(lambda x: x.reshape(x.size(0), embedding_size, x.size(1))),
nn.BatchNorm1d(embedding_size),
nn.Dropout2d(dropout_p),
Lambda(lambda x: x.mean(2)),
nn.Linear(embedding_size, hidden_size),
nn.BatchNorm1d(hidden_size),
nn.Dropout(dropout_p),
nn.ReLU(),
nn.Linear(hidden_size, 1),
)
def forward(self, input_ids: torch.Tensor) -> torch.Tensor:
embedded = self.embedding(input_ids)
return self.out(embedded).reshape(-1)
# -
# In next section, we will intepretate model's prediction. It works best with binary classificator with one output and we will train our model for this by changing criterion to `BCEWithLogitsLoss`.
#
# Create model, optimizer and criterion!
# +
from catalyst.contrib.nn import RAdam
model = EmbeddingModel(len(vocab), dropout_p=0.2)
optimizer = RAdam(model.parameters(), lr=1e-3)
criterion = nn.BCEWithLogitsLoss()
# -
# Embeddings are difficult to train. We will use pretrained one.
model.embedding.weight.data.copy_(vocab.vectors)
# +
from catalyst.dl import (
SupervisedRunner, MultiLabelAccuracyCallback)
runner = SupervisedRunner(device=device)
# +
from datetime import datetime
from pathlib import Path
logdir = Path("emb_logs") / datetime.now().strftime("%Y%m%d-%H%M%S")
# -
# `AccuracyCallback` doesnt' work well with binary classificator. So, we change it to `MultiLabelAccuracyCallback`.
runner.train(
model=model,
optimizer=optimizer,
criterion=criterion,
callbacks=[
MultiLabelAccuracyCallback(threshold=0.5),
],
loaders=loaders,
verbose=True,
num_epochs=10,
logdir=logdir,
)
# ### Model Interpretability
#
# Model's prediction interpretation is one of the ML-Engeenier task. To understand Neural Network prediction, we have great tool by PyTorch: [captum](https://github.com/pytorch/captum). It includes several algorithm, and we will use one (LayerIntegratedGradients, [arxiv](https://arxiv.org/pdf/1805.05492.pdf)) of them to understand which words influence on prediction.
# +
from torchtext.data.utils import get_tokenizer
tokenize = get_tokenizer("basic_english")
# +
from typing import List
from captum.attr import LayerIntegratedGradients, TokenReferenceBase, visualization
token_reference = TokenReferenceBase(reference_token_idx=PAD_ID)
lig = LayerIntegratedGradients(model, model.embedding)
# +
# accumalate couple samples in this array for visualization purposes
vis_data_records_ig = []
def interpret_sentence(
model: nn.Module, sentence: str, min_len: int = 7, label: int = 0
):
model.eval()
text = [tok for tok in tokenize(sentence)]
if len(text) < min_len:
text += ['<pad>'] * (min_len - len(text))
indexed = [vocab.stoi[t] for t in text]
model.zero_grad()
input_indices = torch.tensor(indexed, device=device)
input_indices = input_indices.unsqueeze(0)
# input_indices dim: [sequence_length]
seq_length = min_len
# predict
pred = torch.sigmoid(model(input_indices)).item()
pred_label = "pos" if pred > 0.5 else "neg"
# generate reference indices for each sample
reference_indices = token_reference.generate_reference(
seq_length, device=device
).unsqueeze(0)
# compute attributions and approximation delta using layer integrated gradients
attributions_ig, delta = lig.attribute(
input_indices,
reference_indices,
n_steps=5000,
return_convergence_delta=True,
)
print(f'pred: {pred_label}({pred:.2}), delta: {abs(delta)}')
add_attributions_to_visualizer(
attributions_ig,
text,
pred,
pred_label,
label,
delta,
vis_data_records_ig,
)
def add_attributions_to_visualizer(
attributions: torch.Tensor,
text: str,
pred: int,
pred_ind: str,
label: int,
delta: float,
vis_data_records: List[visualization.VisualizationDataRecord],
):
attributions = attributions.sum(dim=2).squeeze(0)
attributions = attributions / torch.norm(attributions)
attributions = attributions.cpu().detach().numpy()
# storing couple samples in an array for visualization purposes
vis_data_records.append(
visualization.VisualizationDataRecord(
attributions,
pred,
pred_ind,
label,
"pos" if label == 1 else "neg",
attributions.sum(),
text,
delta,
)
)
# -
# We have few sentence for testing
interpret_sentence(model, 'It was a fantastic performance!', label=1)
interpret_sentence(model, 'Best film ever', label=1)
interpret_sentence(model, 'It was a horrible movie', label=0)
interpret_sentence(model, 'It is a disgusting movie!', label=0)
print('Visualize attributions based on Integrated Gradients')
visualization.visualize_text(vis_data_records_ig)
# ## Part-of-Speech
#
# Move on from text classification to token classification. Tokens can include information like year, name, location and e.t.c. Or we try to analisy syntax of sentences by predcting part of speech for each token. Let's solve problem of part of speech prediction by a RNN neural network!
# +
from torchtext.experimental.datasets import CoNLL2000Chunking
train_dataset, test_dataset = CoNLL2000Chunking()
vocab, pos_vocab, _ = train_dataset.get_vocabs()
vocab.load_vectors(embedding_glove)
# -
PAD_ID = vocab.stoi['<pad>']
# +
def collate_fn(
batch: Tuple[torch.Tensor, torch.Tensor]
) -> Tuple[torch.Tensor, torch.Tensor]:
texts = []
token_types = []
max_len = max(t.size(0) for t, _, _ in batch)
for idx, (txt, tt, _) in enumerate(batch):
new_txt = torch.zeros((1, max_len)) + PAD_ID
new_tt = torch.zeros((1, max_len)) + PAD_ID
new_txt[0, : txt.size(0)] = txt
texts.append(new_txt)
new_tt[0, : tt.size(0)] = tt
token_types.append(new_tt)
return (
torch.cat(texts).type(torch.LongTensor),
torch.cat(token_types).type(torch.LongTensor),
)
batch_size = 256
loaders = {
"train": DataLoader(
train_dataset,
batch_size=batch_size,
shuffle=True,
drop_last=True,
collate_fn=collate_fn,
),
"valid": DataLoader(
test_dataset,
batch_size=batch_size,
shuffle=False,
drop_last=False,
collate_fn=collate_fn,
),
}
# -
# Our POS-model now consist of three components:
# - Embeddigns layer
# - (multi or single layer) RNN
# - Classifier for each token
#
# RNN has three main architectures: simple RNN, LSTM and GRU. Choose one of them to solve our task.
class POSModel(nn.Module):
def __init__(
self,
vocab_size: int,
embedding_size: int = 300,
hidden_size: int = 150,
num_classes: int = 2,
dropout_p: float = 0.1,
):
super().__init__()
self.embedding = nn.Embedding(vocab_size, embedding_size)
self.rnn = nn.LSTM(
embedding_size,
hidden_size,
num_layers=2,
dropout=dropout_p,
batch_first=True,
)
self.clf = nn.Linear(hidden_size, num_classes)
def forward(self, input_ids: torch.Tensor) -> torch.Tensor:
embedded = self.embedding(input_ids)
output, _ = self.rnn(embedded)
return self.clf(output)
# Create model, optimizer and criterion. We want predict POS for each token. But, some tokens, like `PAD`, hasn't POS property(we don't know their POS). That's why we will ignore them.
model = POSModel(len(vocab), dropout_p=0.1, num_classes=len(pos_vocab))
optimizer = RAdam(model.parameters(), lr=1e-2)
criterion = nn.CrossEntropyLoss(ignore_index=PAD_ID) # Ignore PAD token
model.embedding.weight.data.copy_(vocab.vectors)
# +
from typing import Dict
class POSRunner(SupervisedRunner):
def _handle_batch(self, batch: Dict[str, torch.Tensor]):
input_ids = batch["features"]
pos_tags = batch["targets"]
output = self.model(input_ids)
self.input = {"input_ids": input_ids, "targets": pos_tags.reshape(-1)}
self.output = {"logits": output.reshape(-1, output.size(2))}
runner = POSRunner(device=device)
# -
logdir = Path("pos_logs") / datetime.now().strftime("%Y%m%d-%H%M%S")
# +
from catalyst.dl import AccuracyCallback
runner.train(
model=model,
optimizer=optimizer,
criterion=criterion,
callbacks=[
AccuracyCallback(),
],
loaders=loaders,
verbose=True,
num_epochs=10,
logdir=logdir,
)
| week-08/seminar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: data_science
# language: python
# name: data_science
# ---
# + [markdown] tags=[]
# # Cloud Seeding
#
# These data were collected in the summer of 1975 from an experiment to investigate the use of silver iodide in cloud seeding to increase rainfall. In the experiment, which was conducted in an area of Florida, 24 days were judged suitable for seeding on the basis that a measured
# suitability criterion, denoted $S-Ne$, was not less than 1.5. On each day, the decision to seed was made randomly. The response variable $Y$
# is the amount of rain (in cubic meters $\times 10^7$) that fell
# in the target area for a 6 hour period on each suitable day. As well as $S - Ne$, the following explanatory variables were also recorded on each suitable day.
#
# - $A$ = Action: an indicator of wheter seeding action occured (1 yes, 0 no)
# - $T$ = Time: number of days after the first day of the experiment (June 1, 1975)
# - $C$ = Echo covarage: the percentage cloud cover in the experimental area, measured using radar
# - $P$ = Pre-wetness: the total rainfall in the target area 1 hour before seeding (in cubic meters $\times 10^7$)
# - $E$ = Echo motion: an indicator showing wheter the radar echo was moving (1) or stationary (2)
#
# The aim is to set up a model to investigate how $Y$ is related to
# the explanatory variables. There are several difficulties; for instance,
# the second data point seems untypical in several ways. The second data point will be highlighted
# separately in histograms as the 'Outlier Point'.
#
# This is the 70th data set from "A Handbook of Small Data Sets".
# + [markdown] tags=[]
# ## 1. Data description
# +
# %matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.patches
import numpy as np
import plotly.express as px
from sklearn.metrics import r2_score
from sklearn.linear_model import LinearRegression
from sklearn.metrics import classification_report
data = pd.read_csv('data.csv')
# -
data
# +
sizes = [data[data['A'] == 0].shape[0], data[data['A'] == 1].shape[0]]
fig, ax = plt.subplots()
ax.set_xticks([0, 1])
ax.set_xticklabels(['Seeding', 'No Seeding'])
ax.set_ylabel('Occurences')
plt.bar([0, 1], sizes)
plt.title("Distribution of seeding action indicators (A)")
plt.show()
# -
# The seeding action indicators are almost evenly distributed, as the data set contains 11 days where seeding occured and 12 days where it didn't.
# +
sizes = [data[data['E'] == 1].shape[0], data[data['E'] == 2].shape[0]]
fig, ax = plt.subplots()
ax.set_xticks([0, 1])
ax.set_xticklabels(['Moving', 'Stationary'])
ax.set_yticks(np.arange(0, max(sizes), 2))
ax.set_ylabel('Occurences')
plt.bar([0, 1], sizes)
plt.title("Distribution of echo motion indicators (E)")
plt.show()
# -
# Meanwhile, the echo motion indicators tell another story. It looks like the echo radar is usually moving, and it being stationary is more of an anomaly, since it
# acoounts for only 17% of data points.
# +
fig, ax = plt.subplots()
plt.title('Distribution of time (T)')
plt.hist([data[data['T'] != 1]['T'],
data[data['T'] == 1]['T']],
bins=12,
stacked=True,
color=('C0', 'r'))
ax.set_ylabel('Occurences')
ax.set_xlabel('Day')
ax.legend(['Usual Points', 'Outlier Point'])
plt.show()
print(f"Time",
f"Mean: {data['T'].mean()}",
f"Std: {data['T'].std()}",
f"Value range: {data['T'].min(), data['T'].max()}",
sep='\n')
# -
# The time data points are distributed evenly enough for our purpose, with a mean of 36.5 and a variance of 25.2, with data points ranging over 84 days.
# +
fig, ax = plt.subplots()
plt.title('Distribution of echo coverage (C)')
plt.hist([data[data['T'] != 1]['C'],
data[data['T'] == 1]['C']],
stacked=True, color=('C0', 'r'))
ax.set_ylabel('Occurences')
ax.set_xlabel('Cloud Cover Percentage')
ax.legend(['Usual Points', 'Outlier Point'])
plt.show()
print("Stats with outlier",
f"Mean: {data['C'].mean()}",
f"Std: {data['C'].std()}",
sep='\n')
print()
print("Stats without outlier",
f"Mean: {data[data['T'] != 1]['C'].mean()}",
f"Std: {data[data['T'] != 1]['C'].std()}",
sep='\n')
print()
print(f"Percentage of points with C<5%: {data[data['C'] <= 5].shape[0] / data.shape[0] * 100}%")
# -
# If we forget about the outlier data point, the echo coverage has a somewhat predictable distribution. As the coverage value is increasing, the less probable of occuring it is. For example, echo coverage under 5% accounts for around 48% of data points.
# +
fig, ax = plt.subplots()
plt.title('Distribution of the suitability criterion (S-Ne)')
plt.hist([data[(data['T'] != 1) & (data['S-Ne'] >= 1.5)]['S-Ne'],
data[(data['T'] == 1) & (data['S-Ne'] >= 1.5)]['S-Ne'],
data[data['S-Ne'] < 1.5]['S-Ne']],
stacked=True,
bins=20,
color=('C0', 'r', 'gray'))
ax.set_ylabel('Occurences')
ax.set_xlabel('S-Ne value')
ax.legend(['Usual Points', 'Outlier Point', 'Misadded Point'])
plt.show()
print()
print(f"Suitability criterion (S-Ne)",
f"Mean: {data['S-Ne'].mean()}",
f"Std: {data['S-Ne'].std()}",
f"Value range: {data['S-Ne'].min(), data['S-Ne'].max()}",
sep='\n')
print()
# -
# The data set was supposed data points where the suitability criterion was at least 1.5, but it contains a point with $S-Ne = 1.3$. Outside this fact, the suitability criterion seems pretty uniformly distributed, with a slight tendency around 3.5.
# +
fig, ax = plt.subplots()
plt.title('Distribution of pre-wetness (P)')
plt.hist([data[data['T'] != 1]['P'],
data[data['T'] == 1]['P']],
stacked=True,
color=('C0', 'r'))
ax.set_ylabel('Occurences')
ax.set_xlabel('Pre-wetness value (cubic meters x 10^7)')
ax.legend(['Usual Points', 'Outlier Point'])
plt.show()
print()
print(f"",
f"Mean: {data['P'].mean()}",
f"Std: {data['P'].std()}",
f"Value range: {data['P'].min(), data['P'].max()}",
sep='\n')
print()
print(f"Percentage of points with P<0.5%: {data[data['P'] <= 0.5].shape[0] / data.shape[0] * 100}%")
# -
# Similarly to the echo coverage, as the pre-wetness value increases, it's less likely to occur. 78.2% of the points in the data set correspond to
# a P value less than 0.5.
# +
fig,ax = plt.subplots()
plt.title('Distribution of rainfall (Y)')
plt.hist([data[data['T'] != 1]['Y'],
data[data['T'] == 1]['Y']],
stacked=True,
color=('C0', 'r'))
ax.set_ylabel('Occurence')
ax.set_xlabel(' value')
ax.legend(['Usual Points', 'Outlier Point'])
plt.show()
print()
print(f"Amount of rain (in cubic meters ×10^7) that fell in the target area for a 6 hour period on each suitable day (Y)",
f"Mean: {data['Y'].mean()}",
f"Std: {data['Y'].std()}",
f"Value range: {data['Y'].min(), data['Y'].max()}",
sep='\n')
print()
# -
# The Y-values seem almost evenly distributed in the interval (0, 7) with 2 data points with Y > 11.
# ## 2. Correlation
data.corr().style.applymap(lambda v: 'opacity: 20%;' if ((v < 0.3) and (v > -0.3)) or (v == 1) else None)
# +
def plot_corr(df,size=8):
f = plt.figure(figsize=(size, size))
corr = df.corr()
plt.matshow(corr, fignum=f.number)
plt.colorbar()
plt.xticks(range(len(corr.columns)), corr.columns)
plt.yticks(range(len(corr.columns)), corr.columns)
plt.title("Covariance matrix")
plot_corr(data)
# -
# ## 2.1 Removing the outlier point
# There are a lot of meaningful covariances that can be worked with, but the outlier point seems to heavily skew the values. Below we can see a difference between the covariance matrices after excluding the outlier point.
# +
diff = data.corr() - data.drop(1, axis=0).corr()
def style_negative(v, props=''):
return props if v < 0 else None
diff = diff.style.set_caption("Covariance matrix difference")\
.applymap(style_negative, props='color:red;')\
.applymap(lambda v: 'opacity: 20%;' if (v < 0.1) and (v > -0.1) else None)
diff
# -
# For example, the outlier point has both a very high pre-wetness and echo coverage indicators relative to the other data points. Therefore, it makes the echo coverage and pre-wetness look much more correlated than they actually are.
# +
f = plt.figure(figsize=(15, 4))
ax1 = f.add_subplot(121)
ax2 = f.add_subplot(122)
ax1.scatter(data['P'], data['C'], label='_nolegend_')
ax1.scatter([data['P'][1]], [data['C'][1]], color='r')
ax1.legend(["Outlier point"])
ax2.scatter(data.drop(1, axis=0)['P'], data.drop(1, axis=0)['C'])
ax1.set_title("With outlier point (p={:.3f})".format(data.corr()['P']['C']))
ax2.set_title("Without outlier point (p={:.3f})".format(data.drop(1, axis=0).corr()['P']['C']))
ax1.set_ylabel("Echo coverage")
ax1.set_xlabel("Pre-wetness")
ax2.set_ylabel("Echo coverage")
ax2.set_xlabel("Pre-wetness")
plt.show()
# -
# Even if the echo coverage of the outlier point is quite high relative to the other data points, it's not an unreasonable value. A day with 38\% cloud coverage corresponds to scattered clouds, which, although not very likely, are entirely possible in summer Florida.
#
# Therefore, since there are not enough data points to really see what happens in more cloudy days, there are two possible ways of going forward:
#
# - Remove the outlier point and focus on sunny days (days with echo coverage percentage less than 20\%) which are more likely in Florida. As a result, the accuracy of the analysis will be higher at the expense of a smaller population considered.
#
# - Keep the outlier point and assume that the linear dependence between cloud coverage and the other variable holds for higher values of cloud coverage (20\% - 40\%). Therefore, interpolation would be fairly accurate and some insight could be gained for this relatively cloud coverage range.
#
# The outlier cloud coverage value is almost triple the second highest value in the data set, so the bold assumption made in the second approach above would be too risky to be used. Since we are more focused on results for typical summer days in Florida, the first approach will be used going forward and the outlier point will be dropped.
data = data.drop(1, axis=0)
# ## 2.2. Correlation without outlier
data.corr().style.applymap(lambda v: 'opacity: 20%;' if ((v < 0.3) and (v > -0.3)) or (v == 1) else None)
plot_corr(data)
# ## 2.3 Echo motion is insignificant
#
# The echo indicator seems to be somewhat correlated to the response variable, but this is a bit misleading since only 17\% percent of the data points have a stationary radar echo. We can't say much with so few stationary data points, but it looks like the echo motion does not really influence the amount of rain $Y$. As a result, the echo motion feature will be dropped going forward.
# +
fig, ax = plt.subplots()
ax.scatter(data[data['E'] == 1]['E'], data[data['E'] == 1]['Y'])
ax.scatter(data[data['E'] == 2]['E'], data[data['E'] == 2]['Y'])
ax.legend(("Moving echo", "Stationary echo"))
ax.set_title("Echo motion (E) vs amount of rainfall (Y)")
ax.set_xlabel("Echo motion (E)")
ax.set_ylabel("Amount of rainfall (Y)")
plt.show()
# -
data = data.drop('E', axis=1)
# ## 2.4 Time, Suitability Criterion and Cloud Coverage
# +
f = plt.figure(figsize=(15, 4))
ax1 = f.add_subplot(121)
ax2 = f.add_subplot(122)
ax1.scatter(data['T'], data['S-Ne'])
ax2.scatter(data['T'], data['C'])
ax1.set_title("Time vs Suitability criterion (p={:.3f})".format(data.corr()['T']['S-Ne']))
ax2.set_title("Time vs Cloud coverage (p={:.3f})".format(data.corr()['T']['C']))
ax1.set_xlabel("Time (T)")
ax1.set_ylabel("Suitability criterion (S-Ne)")
ax2.set_xlabel("Time (T)")
ax2.set_ylabel("Cloud coverage (C)")
plt.show()
# -
ax = plt.axes(projection='3d')
ax.scatter(data['S-Ne'], data['C'], data['T'], c=data['T'], cmap='viridis', linewidth=0.5);
# # 3. Regression model
# +
X = data.drop('Y', axis=1)
Y = data['Y']
reg = LinearRegression().fit(X, Y)
reg.score(X, Y)
| seeding/seeding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # PROBLEMAS DIVERSOS
# <h3>1.</h3>
# Escribí un programa que solicite al usuario ingresar la cantidad de kilómetros recorridos por una motocicleta y la cantidad de litros de combustible que consumió durante ese recorrido. Mostrar el consumo de combustible por kilómetro.
# <code>Kilómetros recorridos: 260
# Litros de combustible gastados: 12.5
# El consumo por kilómetro es de 20.8<code>
# +
kilometros = float (input ('ingresar cantidad de kilometros recorrido por bicicleta'))
litros = float (input ('ingresar cantidad de litros de conbustibles durante el recorrido'))
consumo_por_kilometros = + kilometros / + litros
print (f'el consumo por kilometro es: { consumo_por_kilometros}')
# -
# ### 2.
# Escriba un programa que pida los coeficientes de una ecuación de segundo grado <code>(a x² + b x + c = 0)</code> y escriba la solución.
#
# Se recuerda que una ecuación de segundo grado puede no tener solución, tener una solución única, tener dos soluciones o que todos los números sean solución.
# <img src='https://i.pinimg.com/originals/d3/f7/01/d3f701528ad56ce0f5a98d7c91722fd7.png'>
# Su programa debe indicar:
# - En caso la ecuación cuadrática tenga solución real, su programa debe brindar la solución
# - En caso su ecuación no tenga solución real, su programa debe brindar un mensaje que diga "Ecuación no presenta solución real"
# +
from math import sqrt
A = int(input("Ingrese el coeficiente de la variable cuadrática\n"))
B = int(input("Ingrese el coeficiente de la variable lineal\n"))
C = int(input("Ingrese el término independiente\n"))
x1= 0
x2= 0
if ((B**2)-4*A*C) < 0:
print("La solución de la ecuación es con números complejos")
else:
x1 = (-B+sqrt(B**2-(4*A*C)))/(2*A)
x2 = (-B-sqrt(B**2-(4*A*C)))/(2*A)
print("Las soluciones de la ecuación son:")
print(x1)
print(x2)
| Modulo1/Ejercicios/Problemas Diversos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.6 64-bit (''songs-G2cSMnHZ'': pipenv)'
# name: python3
# ---
# - <https://scikit-learn.org/stable/modules/generated/sklearn.mixture.GaussianMixture.html>
#
# - <https://jakevdp.github.io/PythonDataScienceHandbook/05.12-gaussian-mixtures.html#Generalizing-E%E2%80%93M:-Gaussian-Mixture-Models>
from os import getenv
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.pipeline import Pipeline
from sklearn.model_selection import GridSearchCV
from sklearn.preprocessing import StandardScaler
from sklearn.mixture import GaussianMixture #covariance_type='diag'
from scipy.spatial import distance
kg = pd.read_csv('spotify_tracks_metadata.csv')
kg_slim = kg[['danceability','energy','key','loudness','mode','speechiness','acousticness','instrumentalness','liveness','valence','tempo','duration_ms','time_signature']]
kg_slim.head()
# +
pipe = Pipeline(steps=[('scaler', StandardScaler()), ('gauss', GaussianMixture(
n_components=5, # could tune by minimizing aic or bic
covariance_type='diag', # 'full' is best, 'spherical' is fastest
n_init=10))]) # Hands-On ML warns that default n_init=1 is too low
pipe.fit(kg_slim)
# -
# Like coordinates in multidimensional song-space, relative to clusters;
# Or like affiliation scores to distinct song distributions;
db_song_coords = pipe.predict_proba(kg_slim)
# ## the model (pipe) and what it says about the database songs (db_song_coords) could be saved offline
# ## then, via the app, we provide a new_song (or song_name + artist_name);
# ## the app queries spotify for this new_song's features;
# ## feeds those audio_features to our pre-trained model;
# ## and compares the read-out to its offline analysis of database songs (db_song_coords).
# +
"""Setting up APIs for this notebook"""
from client.spotify_client import *
import spotipy
from spotipy.oauth2 import SpotifyClientCredentials
SPOTIFY_API_KEY = getenv("SPOTIFY_API_KEY")
SPOTIFY_API_KEY_SECRET = getenv("SPOTIFY_API_KEY_SECRET")
# get spotify API
spotify = SpotifyAPI(SPOTIFY_API_KEY, SPOTIFY_API_KEY_SECRET)
# get spotiPy API (wrapper for spotify API)
auth_manager = SpotifyClientCredentials(SPOTIFY_API_KEY, SPOTIFY_API_KEY_SECRET)
spotiPy = spotipy.Spotify(auth_manager=auth_manager)
# +
# html could accept user input, typed into 2 separate fields; hit a button
user_input_song = 'superstition'
user_input_artist = 'stevie wonder'
# app@route('/') takes user_input_song, user_input_artist...feeds APIs
# spotify API -- from (song_name, artist_name) to track's (Spotify ID);
new_song = spotify.search(
{'track':user_input_song, 'artist':user_input_artist}, search_type='track')
new_song_spot_id = new_song['tracks']['items'][0]['id']
# spotiPy API -- from track's (Spotify ID) to its (audio_features):
new_song_features = spotiPy.audio_features(tracks=[new_song_spot_id])
# arrange new_song's audio_features to match feature matrix used to train model;
new_song_ready = pd.DataFrame(new_song_features)[['danceability','energy','key','loudness','mode','speechiness','acousticness','instrumentalness','liveness','valence','tempo','duration_ms','time_signature']]
# app unpickles saved model (pipe) and saved model output (db_song_coords);
# new_song_ready meets the model;
new_song_coords = pipe.predict_proba(new_song_ready)
# use scipy.distance.cdist(...'cosine') to get cosine similarity scores,
# comparing new_song to everything stored in database
distances = distance.cdist(db_song_coords, new_song_coords, 'cosine')
# -
distances.shape
pd.DataFrame(distances).hist()
# Well, that did not work well. Maybe we need more clusters. I chose an arbitrary number, n_components=5.
#
# pipe.fit() took 2 minutes on my laptop. With unlimited time to train a model offline, we can come up with something.
#
# This site shows how to tune n_components, to minimize aic or bic, both built-in attributes of a fitted GaussianMixture model.
# - <https://jakevdp.github.io/PythonDataScienceHandbook/05.12-gaussian-mixtures.html#Generalizing-E%E2%80%93M:-Gaussian-Mixture-Models>
#
# Also, covariance_type could be set to 'full', for a better fitting model.
#
# If GaussianMixture doesn't cut it, any other clustering (unsupervised) model that has a predict_proba method would work well with the general steps outlined in this notebook.
| GaussianMixture_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!--BOOK_INFORMATION-->
# <img align="left" style="padding-right:10px;" src="fig/cover-small.jpg">
#
# *Este notebook es una adaptación realizada por <NAME> del material "[Whirlwind Tour of Python](http://www.oreilly.com/programming/free/a-whirlwind-tour-of-python.csp)" de Jake VanderPlas; tanto el [contenido original](https://github.com/jakevdp/WhirlwindTourOfPython) como la [adpatación actual](https://github.com/rrgalvan/PythonIntroMasterMatemat)] están disponibles en Github.*
#
# *The text and code are released under the [CC0](https://github.com/jakevdp/WhirlwindTourOfPython/blob/master/LICENSE) license; see also the companion project, the [Python Data Science Handbook](https://github.com/jakevdp/PythonDataScienceHandbook).*
#
# <!--NAVIGATION-->
# < [String Manipulation and Regular Expressions](14-Strings-and-Regular-Expressions.ipynb) | [Contents](Index.ipynb) | [Resources for Further Learning](16-Further-Resources.ipynb) >
# # A Preview of Data Science Tools
# If you would like to spring from here and go farther in using Python for scientific computing or data science, there are a few packages that will make your life much easier.
# This section will introduce and preview several of the more important ones, and give you an idea of the types of applications they are designed for.
# If you're using the *Anaconda* or *Miniconda* environment suggested at the beginning of this report, you can install the relevant packages with the following command:
# ```
# $ conda install numpy scipy pandas matplotlib scikit-learn
# ```
# Let's take a brief look at each of these in turn.
# ## NumPy: Numerical Python
#
# NumPy provides an efficient way to store and manipulate multi-dimensional dense arrays in Python.
# The important features of NumPy are:
#
# - It provides an ``ndarray`` structure, which allows efficient storage and manipulation of vectors, matrices, and higher-dimensional datasets.
# - It provides a readable and efficient syntax for operating on this data, from simple element-wise arithmetic to more complicated linear algebraic operations.
#
# In the simplest case, NumPy arrays look a lot like Python lists.
# For example, here is an array containing the range of numbers 1 to 9 (compare this with Python's built-in ``range()``):
import numpy as np
x = np.arange(1, 10)
x
# NumPy's arrays offer both efficient storage of data, as well as efficient element-wise operations on the data.
# For example, to square each element of the array, we can apply the "``**``" operator to the array directly:
x ** 2
# Compare this with the much more verbose Python-style list comprehension for the same result:
[val ** 2 for val in range(1, 10)]
# Unlike Python lists (which are limited to one dimension), NumPy arrays can be multi-dimensional.
# For example, here we will reshape our ``x`` array into a 3x3 array:
M = x.reshape((3, 3))
M
# A two-dimensional array is one representation of a matrix, and NumPy knows how to efficiently do typical matrix operations. For example, you can compute the transpose using ``.T``:
M.T
# or a matrix-vector product using ``np.dot``:
np.dot(M, [5, 6, 7])
# and even more sophisticated operations like eigenvalue decomposition:
np.linalg.eigvals(M)
# Such linear algebraic manipulation underpins much of modern data analysis, particularly when it comes to the fields of machine learning and data mining.
#
# For more information on NumPy, see [Resources for Further Learning](16-Further-Resources.ipynb).
# ## Pandas: Labeled Column-oriented Data
#
# Pandas is a much newer package than NumPy, and is in fact built on top of it.
# What Pandas provides is a labeled interface to multi-dimensional data, in the form of a DataFrame object that will feel very familiar to users of R and related languages.
# DataFrames in Pandas look something like this:
import pandas as pd
df = pd.DataFrame({'label': ['A', 'B', 'C', 'A', 'B', 'C'],
'value': [1, 2, 3, 4, 5, 6]})
df
# The Pandas interface allows you to do things like select columns by name:
df['label']
# Apply string operations across string entries:
df['label'].str.lower()
# Apply aggregates across numerical entries:
df['value'].sum()
# And, perhaps most importantly, do efficient database-style joins and groupings:
df.groupby('label').sum()
# Here in one line we have computed the sum of all objects sharing the same label, something that is much more verbose (and much less efficient) using tools provided in Numpy and core Python.
#
# For more information on using Pandas, see [Resources for Further Learning](16-Further-Resources.ipynb).
# ## Matplotlib MatLab-style scientific visualization
#
# Matplotlib is currently the most popular scientific visualization packages in Python.
# Even proponents admit that its interface is sometimes overly verbose, but it is a powerful library for creating a large range of plots.
#
# To use Matplotlib, we can start by enabling the notebook mode (for use in the Jupyter notebook) and then importing the package as ``plt``"
# run this if using Jupyter notebook
# %matplotlib notebook
import matplotlib.pyplot as plt
plt.style.use('ggplot') # make graphs in the style of R's ggplot
# Now let's create some data (as NumPy arrays, of course) and plot the results:
x = np.linspace(0, 10) # range of values from 0 to 10
y = np.sin(x) # sine of these values
plt.plot(x, y); # plot as a line
# If you run this code live, you will see an interactive plot that lets you pan, zoom, and scroll to explore the data.
#
# This is the simplest example of a Matplotlib plot; for ideas on the wide range of plot types available, see [Matplotlib's online gallery](http://matplotlib.org/gallery.html) as well as other references listed in [Resources for Further Learning](16-Further-Resources.ipynb).
# ## SciPy: Scientific Python
#
# SciPy is a collection of scientific functionality that is built on NumPy.
# The package began as a set of Python wrappers to well-known Fortran libraries for numerical computing, and has grown from there.
# The package is arranged as a set of submodules, each implementing some class of numerical algorithms.
# Here is an incomplete sample of some of the more important ones for data science:
#
# - ``scipy.fftpack``: Fast Fourier transforms
# - ``scipy.integrate``: Numerical integration
# - ``scipy.interpolate``: Numerical interpolation
# - ``scipy.linalg``: Linear algebra routines
# - ``scipy.optimize``: Numerical optimization of functions
# - ``scipy.sparse``: Sparse matrix storage and linear algebra
# - ``scipy.stats``: Statistical analysis routines
#
# For example, let's take a look at interpolating a smooth curve between some data
# +
from scipy import interpolate
# choose eight points between 0 and 10
x = np.linspace(0, 10, 8)
y = np.sin(x)
# create a cubic interpolation function
func = interpolate.interp1d(x, y, kind='cubic')
# interpolate on a grid of 1,000 points
x_interp = np.linspace(0, 10, 1000)
y_interp = func(x_interp)
# plot the results
plt.figure() # new figure
plt.plot(x, y, 'o')
plt.plot(x_interp, y_interp);
# -
# What we see is a smooth interpolation between the points.
# ## Other Data Science Packages
#
# Built on top of these tools are a host of other data science packages, including general tools like [Scikit-Learn](http://scikit-learn.org) for machine learning, [Scikit-Image](http://scikit-image.org) for image analysis, and [Statsmodels](http://statsmodels.sourceforge.net/) for statistical modeling, as well as more domain-specific packages like [AstroPy](http://astropy.org) for astronomy and astrophysics, [NiPy](http://nipy.org/) for neuro-imaging, and many, many more.
#
# No matter what type of scientific, numerical, or statistical problem you are facing, it's likely there is a Python package out there that can help you solve it.
# <!--NAVIGATION-->
# < [String Manipulation and Regular Expressions](14-Strings-and-Regular-Expressions.ipynb) | [Contents](Index.ipynb) | [Resources for Further Learning](16-Further-Resources.ipynb) >
| 15-Preview-of-Data-Science-Tools.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load in
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list the files in the input directory
import os
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
from __future__ import absolute_import, division, print_function
import os
import numpy as np
import matplotlib.pyplot as plt
# + _uuid="dba171cdf87546a2444c89c8e60cbb83cb280d90"
import zipfile
import tensorflow as tf
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Conv2D, Flatten, Dropout, MaxPooling2D
from tensorflow.keras.preprocessing.image import ImageDataGenerator
import glob, os, random
# + _uuid="09e990b9330db24c92a0c665d0012d210a7e3e75"
import numpy as np
from keras.preprocessing import image
import matplotlib.pyplot as plt
base_path = 'dataset/ds/'
img_list = glob.glob(os.path.join(base_path, '*/*.jpg'))
for i, img_path in enumerate(random.sample(img_list, 2)):
img = image.load_img(img_path, target_size=(256, 341))
img = image.img_to_array(img, dtype=np.uint8)
plt.subplot(2, 3, i+1)
plt.imshow(img.squeeze())
# + _uuid="45cf2c15f12b03c448faf1dbc148913f9b141b27"
train_datagen = ImageDataGenerator(
rescale=1./255,
shear_range=0.1,
zoom_range=0.1,
width_shift_range=0.1,
height_shift_range=0.1,
horizontal_flip=True,
vertical_flip=True,
validation_split=0.1
)
test_datagen = ImageDataGenerator(
rescale=1./255,
validation_split=0.1
)
train_generator = train_datagen.flow_from_directory(
base_path,
target_size=(224, 224),
batch_size=16,
class_mode='categorical',
subset='training',
seed=0
)
validation_generator = test_datagen.flow_from_directory(
base_path,
target_size=(224, 224),
batch_size=16,
class_mode='categorical',
subset='validation',
seed=0
)
# + _uuid="fff243361dd423eb00c3789bdc238f0186ac323f"
from keras.applications.resnet152 import ResNet152
IMG_SHAPE = (224,224,3)
base_model = tf.keras.applications.resnet.ResNet152(input_shape=IMG_SHAPE,
include_top=False,
weights='imagenet')
# + _uuid="cb3a30042c960b4d0acc76c9ac0b826de176340f"
base_model.trainable = False
# + _uuid="ca84e1428d78a24ad30a800a4c76ca90906d9256"
from tensorflow import keras
model = tf.keras.Sequential([
base_model,
keras.layers.GlobalAveragePooling2D(),
keras.layers.Dense(2, activation='sigmoid')
])
# + _uuid="34e0dff3be6a2e631a8a0a6d1f1ab44ed649cfc2"
# model.compile(optimizer=tf.keras.optimizers.RMSprop(lr=0.0001),
# loss='categorical_crossentropy',
# metrics=['accuracy'])
model.compile(loss='mean_squared_error', optimizer='adam', metrics=['acc'])
# + _uuid="53b1e7551b6f8393ac422b596089098f976b84da"
batch_size = 32
epochs = 10
steps_per_epoch = train_generator.n // batch_size
validation_steps = validation_generator.n // batch_size
print(steps_per_epoch,validation_steps)
history = model.fit_generator(train_generator,
steps_per_epoch = steps_per_epoch,
epochs=epochs,
workers=4,
validation_data=validation_generator,
validation_steps=validation_steps)
# + _uuid="58ecf13b1352f7b462e36e0211c103652897fda9"
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
plt.figure(figsize=(8, 8))
plt.subplot(2, 1, 1)
plt.plot(acc, label='Training Accuracy')
plt.plot(val_acc, label='Validation Accuracy')
plt.legend(loc='lower right')
plt.ylabel('Accuracy')
plt.ylim([min(plt.ylim()),1])
plt.title('Training and Validation Accuracy')
plt.subplot(2, 1, 2)
plt.plot(loss, label='Training Loss')
plt.plot(val_loss, label='Validation Loss')
plt.legend(loc='upper right')
plt.ylabel('Cross Entropy')
plt.ylim([0,max(plt.ylim())])
plt.title('Training and Validation Loss')
plt.show()
# + [markdown] _uuid="531cd337be0b4c5aca777d9123336ba2c3e01329"
# ## Fine Tuning
# + _uuid="cced640fc5d0afab4e9bdbb025047979535b3a6b"
base_model.trainable = True
# Let's take a look to see how many layers are in the base model
print("Number of layers in the base model: ", len(base_model.layers))
# Fine tune from this layer onwards
fine_tune_at = 100
# Freeze all the layers before the `fine_tune_at` layer
for layer in base_model.layers[:fine_tune_at]:
layer.trainable = False
# + _uuid="454308071374ac8c3826f09b6d89c06d514df342"
model.compile(loss='binary_crossentropy',
optimizer = tf.keras.optimizers.RMSprop(lr=2e-5),
metrics=['accuracy'])
model.summary()
# + _uuid="55201b8cbd4918892298bd6501ce5e0700179567"
history_fine = model.fit_generator(train_generator,
steps_per_epoch = steps_per_epoch,
epochs=epochs,
workers=4,
validation_data=validation_generator,
validation_steps=validation_steps)
# + _uuid="40fc532563619eea26a85f19d69d479f0c63bdb5"
acc += history_fine.history['acc']
val_acc += history_fine.history['val_acc']
loss += history_fine.history['loss']
val_loss += history_fine.history['val_loss']
# + _uuid="348de00d23b6032c9430be8f3f829c889a6e2ef4"
plt.figure(figsize=(8, 8))
plt.subplot(2, 1, 1)
plt.plot(acc, label='Training Accuracy')
plt.plot(val_acc, label='Validation Accuracy')
plt.ylim([0.9, 1])
plt.plot([epochs-1,epochs-1], plt.ylim(), label='Start Fine Tuning')
plt.legend(loc='lower right')
plt.title('Training and Validation Accuracy')
plt.subplot(2, 1, 2)
plt.plot(loss, label='Training Loss')
plt.plot(val_loss, label='Validation Loss')
plt.ylim([0, 0.2])
plt.plot([epochs-1,epochs-1], plt.ylim(), label='Start Fine Tuning')
plt.legend(loc='upper right')
plt.title('Training and Validation Loss')
plt.show()
# + _uuid="28aef1c81ff4657ba064a1323ba2cfb898d12b06"
# -
| transfer-learning-using-mobilenet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Prevalence of Personal Attacks
#
# In this notebook, we do some basic investigation into the frequency of personal attacks on Wikipedia. We will attempt to provide some insight into the following questions:
#
# - What fraction of comments are personal attacks?
#
# - What fraction of users have made a personal attack?
#
# - What fraction of users have been attacked on their user page?
#
# - Are there any temporal trends in the frequency of attacks?
#
#
# We have 2 separate types of data at our disposal. First, we have a random sample of roughly 100k human-labeled comments. Each comment was annotated by 10 separate people as to whether the comment is a personal attack. We take the majority annotation class to get a single label. Second, we have the full history of comments with scores generated by a machine learning model. Due to the construction of the model, these scores can be interpreted as the probability that the majority of annotators would label the comment as an attack.
# +
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
import pandas as pd
from load_utils import *
from analysis_utils import compare_groups
# -
d = load_diffs()
df_events, df_blocked_user_text = load_block_events_and_users()
er_t = 0.425
d['2015']['pred_attack'] = (d['2015']['pred_attack_score_uncalibrated'] > er_t).astype(int)
d['sample']['pred_attack'] = (d['sample']['pred_attack_score_uncalibrated'] > er_t).astype(int)
# ### Q: What fraction of comments are personal attacks?
# +
# calibration based
# -
100 * d['2015']['pred_attack_score_calibrated'].mean()
100 * d['2015'].groupby('ns')['pred_attack_score_calibrated'].mean()
# +
### threshold based
# -
100 * d['2015']['pred_attack'].mean()
100 * d['2015'].groupby('ns')['pred_attack'].mean()
# ### Q: What fraction of users have made/received at least k personal attacks?
# #### 1. threshold based
# +
ks = [1,3,5]
attacker_ys = []
receiver_ys = []
for k in ks:
attacker_ys.append(d['2015'].groupby('user_text')['pred_attack'].sum().apply(lambda x: x >= k).mean() * 100)
for k in ks:
receiver_ys.append(d['2015'].query("ns=='user'and user_text != page_title").groupby('page_title')['pred_attack'].sum().apply(lambda x: x >= k).mean() * 100)
# -
df_sns = pd.DataFrame()
df_sns['k'] = ks
df_sns['attackers'] = attacker_ys
df_sns['victims'] = receiver_ys
# +
plt.figure()
sns.set(font_scale=1.5)
f, (ax1, ax2) = plt.subplots(2, sharex=True)
sns.barplot("k", y="attackers", data=df_sns, ax=ax1, color = 'darkblue' , label = "% of users who made \n at least k attacks")
sns.barplot("k", y="victims", data=df_sns, ax=ax2, color = 'darkred', label = "% of users who received \n at least k attacks")
ax1.set( xlabel = '' , ylabel = '%')
ax2.set( ylabel = '%')
ax1.legend()
ax2.legend()
plt.savefig('../../paper/figs/attacker_and_victim_prevalence.png')
# +
plt.figure()
sns.set(font_scale=1.5)
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(8,4))
sns.barplot("k", y="attackers", data=df_sns, ax=ax1, color = 'darkblue' )
sns.barplot("k", y="victims", data=df_sns, ax=ax2, color = 'darkred')
ax1.set( ylabel = '%')
ax2.set( ylabel = '')
plt.text(- 3.0, 1.65,'% of users \nwho made \nat least k attacks',fontsize = 15)
plt.text(0.6, 1.65,'% of users \nwho received \nat least k attacks',fontsize = 15)
ax2.legend()
plt.savefig('../../paper/figs/attacker_and_victim_prevalence.png')
# -
# #### 2. calibration based
# Take unsampled data. For each comment, let it be an attack with probability equal to the model prediction. Count the number of users that have made at least 1 attack. Repeat.
def simulate_num_attacks_within_group(df, group_col = 'user_text'):
return df.assign( uniform = np.random.rand(df.shape[0], 1))\
.assign(is_attack = lambda x: (x.pred_attack_score_calibrated >= x.uniform).astype(int))\
.groupby(group_col)['is_attack']\
.sum()
def get_within_group_metric_interval(df, group_col = 'user_text', metric = lambda x: (x>=1).astype(int).mean() * 100, iters = 2):
results = []
for i in range(iters):
result = simulate_num_attacks_within_group(df, group_col = group_col)
result = metric(result)
results.append(result)
return np.percentile(results, [2.5, 97.5])
def get_intervals(df, group_col = 'user_text', iters = 10):
ks = range(1,6)
y =[]
lower = []
upper = []
intervals = []
for k in ks:
metric = lambda x: (x>=k).astype(int).mean() * 100
interval = get_within_group_metric_interval(d['2015'], group_col = group_col, iters = iters, metric=metric)
intervals.append(interval)
y.append(interval.mean())
lower.append(interval.mean() - interval[0])
upper.append(interval[1] - interval.mean())
return pd.DataFrame({'k': ks, 'y': y, 'interval': intervals, 'lower': lower, 'upper': upper})
get_intervals(d['2015'])
# ignore anon users
get_intervals(d['2015'].query('not author_anon and not recipient_anon'))
get_intervals(d['2015'].query("ns=='user'"), group_col = 'page_title')
# ignore anon users
get_intervals(d['2015'].query("not author_anon and not recipient_anon and ns=='user'"), group_col = 'page_title')
# ### Q: Has the proportion of attacks changed year over year?
df_span = d['sample'].query('year > 2003 & year < 2016')
plt.figure(figsize=(8,4))
sns.set(font_scale=1.5)
x = 'year'
s = df_span.groupby(x)['pred_attack'].mean() * 100
plt.plot(s.index, s.values)
plt.xlabel(x)
plt.ylabel('Percent of comments that are attacks')
plt.savefig('../../paper/figs/prevalence_by_year.png')
plt.figure(figsize=(8,4))
sns.set(font_scale=1.5)
x = 'year'
s = df_span.groupby(x)['pred_attack_score_calibrated'].mean() * 100
plt.plot(s.index, s.values)
plt.xlabel('')
plt.ylabel('Percent of comments that are attacks')
#plt.savefig('../../paper/figs/prevalence_by_year.png')
# There is a strong yearly pattern. The fraction of attacks peaked in 2008, which is when participation peaked as well.
| src/analysis/Prevalence of Personal Attacks (paper).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Decision Trees and Random Forest
# Import library
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
df = pd.read_csv('kyphosis.csv')
df.head()
df.info()
df['Kyphosis'].value_counts()
sns.pairplot(df,hue='Kyphosis')
from sklearn.cross_validation import train_test_split
X = df.drop('Kyphosis',axis=1)
y = df['Kyphosis']
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=0.3, random_state=0)
from sklearn.tree import DecisionTreeClassifier
dtree = DecisionTreeClassifier()
dtree.fit(X_train,y_train)
prediction = dtree.predict(X_test)
from sklearn.metrics import classification_report, confusion_matrix
print('confusion_matrix\n',confusion_matrix(y_test, prediction))
print('\n classification_report\n',classification_report(y_test, prediction))
from sklearn.tree import export_graphviz
import graphviz
features = list(df.columns[1:])
features
# +
dot_data = export_graphviz(
dtree,out_file=None,
feature_names = features,
class_names = df['Kyphosis'].unique(),
rounded = True,
filled = True
)
graph = graphviz.Source(dot_data)
graph
# +
from IPython.display import Image
from sklearn.externals.six import StringIO
from sklearn.tree import export_graphviz
import pydot
features = list(df.columns[1:])
features
# +
dot_data = StringIO()
export_graphviz(dtree, out_file=dot_data,feature_names=features,filled=True,rounded=True)
graph = pydot.graph_from_dot_data(dot_data.getvalue())
Image(graph[0].create_png())
# -
# ## Random Forest
from sklearn.ensemble import RandomForestClassifier
rfc = RandomForestClassifier(n_estimators=200,random_state=0)
rfc.fit(X_train,y_train)
rfc_pred = rfc.predict(X_test)
print('confusion_matrix \n',confusion_matrix(y_test, rfc_pred))
print('\n classification_report\n',classification_report(y_test, rfc_pred))
# ### Decision Trees and Random Forests Project
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
plt.style.use('ggplot')
df = pd.read_csv('loan_data.csv')
df.head()
df.describe()
df.info()
# ### Exploratory Data analysis
plt.figure(figsize=(10,6))
df[df['credit.policy']==0]['fico'].plot(kind='hist',bins=30,color='r',alpha=0.7,label='credit.policy 0')
df[df['credit.policy']==1]['fico'].plot(kind='hist',bins=30,color='b', alpha=0.7,label='credit.policy 1')
plt.legend()
plt.xlabel('fico')
plt.figure(figsize=(10,6))
df[df['not.fully.paid']==0]['fico'].plot(kind='hist',bins=30,color='g',alpha=0.7,label='not.fully.paid 0')
df[df['not.fully.paid']==1]['fico'].plot(kind='hist',bins=30,color='r', alpha=0.7,label='not.fully.paid 1')
plt.xlabel('fico')
plt.legend()
plt.figure(figsize=(12,6))
sns.countplot(x = 'purpose',data =df,hue ='not.fully.paid')
sns.jointplot(x='fico',y='int.rate', data=df,color='purple')
sns.lmplot(x = 'fico', y='int.rate', col='not.fully.paid', hue='credit.policy',data =df)
cat_feat = ['credit.policy','purpose','inq.last.6mths','delinq.2yrs','pub.rec']
X_new = pd.get_dummies(df,drop_first=True,columns=cat_feat)
y = df['not.fully.paid']
X_new.head()
# ### Train test split
from sklearn.cross_validation import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X_new,y, test_size=0.3,random_state=0)
# ### Decision tree model
from sklearn.tree import DecisionTreeClassifier
dtree = DecisionTreeClassifier(criterion='entropy',random_state=0)
dtree.fit(X_train,y_train)
# ### Prediction and evaluvation
prediction = dtree.predict(X_test)
from sklearn.metrics import classification_report,confusion_matrix
print('confusion_matrix\n',confusion_matrix(y_test,prediction))
print('classification_report\n',classification_report(y_test,prediction))
# ### Train the Random forest model
from sklearn.ensemble import RandomForestClassifier
rfc = RandomForestClassifier(random_state =0)
rfc.fit(X_train,y_train)
# ### Prediction ans Evaluvation
prediction = rfc.predict(X_test)
print('confusion_matrix\n',confusion_matrix(y_test,prediction))
print('classification_report\n',classification_report(y_test,prediction))
# ### Great Job
| Machine_Learning_Bootcamp/Machine Learning Sections/Decision-Trees-and-Random-Forests/Decision_Trees_Random_forest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Fuzzywuzzy
# **Inhalt:** Wie ähnlich sind zwei Strings? II
#
# **Nötige Skills:** keine
#
# **Lernziele:** Eine andere Möglichkeit für fuzzy string matching kennenlernen
# ## About
# Fuzzywuzzy
# - Package https://github.com/seatgeek/fuzzywuzzy
# - Dokumentation: fast nicht vorhanden
#
# Installation:
#
# ```bash
# pip3 install fuzzywuzzy[speedup]
# ```
#
# ## Setup
import pandas as pd
from fuzzywuzzy import fuzz
from fuzzywuzzy import process
# ## Basics
# Die Fuzzywuzzy-Funktionen spucken eine Zahl zwischen 0 und 100 aus, welche die Ähnlichkeit angibt.
fuzz.ratio("a", "b")
fuzz.ratio("a", "a")
# ## Scorer-Funktionen
# ### ratio
#
# Jedes einzelne Zeichen ist wichtig, auch die Reihenfolge.
fuzz.ratio("<NAME>", "<NAME>")
fuzz.ratio("<NAME>", "Dr. <NAME>")
# ### partial_ratio
#
# Partielle Übereinstimmung der Strings ist ok. Allerdings muss die Reihenfolge der Zeichen stimmen:
fuzz.partial_ratio("<NAME>", "Dr. <NAME>")
fuzz.partial_ratio("<NAME>", "<NAME>")
fuzz.partial_ratio("<NAME>", "<NAME>")
# ### token_sort_ratio
#
# Sortiert die Wörter zuerst. Gibt aber Abzüge für partielle Matches:
fuzz.token_sort_ratio("<NAME>", "<NAME>")
fuzz.token_sort_ratio("<NAME>", "<NAME>")
# ### token_set_ratio
#
# Kommt klar mit vertauschter Reihenfolge und partiellen Matches:
fuzz.token_set_ratio("<NAME>", "<NAME>")
fuzz.token_set_ratio("<NAME>", "<NAME>.")
# Abzüge gibt es für falsche Zeichen innerhalb der Wörter:
fuzz.token_set_ratio("<NAME>", "<NAME>.")
# Der Score geht recht rapide runter. Zum Vergleich:
fuzz.token_set_ratio("<NAME>", "<NAME>")
# ## Beispiel in Pandas
#
# Wir wollen die Daten in einer Tabelle "putzen". Das heisst: die verschiedenen Schreibweisen von Begriffen vereinheitlichen (z.B. "<NAME>" - "<NAME>")
df = pd.read_csv('dataprojects/Jellyfish/Words.csv')
df
# ### Score testen
#
# Wir können mit `df.apply()` ähnlich wie vorher den Score eines Vergleichs testen und in einer neuen Spalte festhalten,
df['token_set_ratio'] = df.apply(lambda row: fuzz.token_set_ratio(row['Wort 1'], row['Wort 2']) ,axis=1)
df
df['match'] = df['token_set_ratio'] > 80
df
# **Frage:** Wie steht es hier um die false positives / false negatives?
# +
# false positives:
#
# +
# false negatives:
#
# -
# ### Wörter ersetzen
#
# Was uns aber eigentlich interessiert: die neue Liste.
#
# **a) Alle Varianten eines Begriffs ersetzen**
#
# Die Funktion `process()` ist dafür hilfreich
#
# Sortiert eine Liste/Dictionary entsprechend dem besten Match...
# - Welchen String wollen wir vergleichen? "<NAME>"
# - Welche Liste wollen wir durchsuchen? "Wort 2"
# - Welchen Scorer wollen wir verwenden? token_set_ratio
# - Wie viele Ergebnisse sollen angezeigt werden? 20
#
# ...und retourniert die Liste mit dem Matching-Score:
match_list = process.extract("<NAME>", df['Wort 2'], scorer=fuzz.token_set_ratio, limit=20)
# Wir vergleichen also jede einzelne Zeile in der Tabelle mit dem String "<NAME>":
match_list
# Für unsere Auswahl brauchen wir aber einen bestimmten "Cut-Off-Punkt".
#
# Zum Beispiel können wir sagen, dass wir nur die Schreibweisen ersetzen wollen, die einen Score über 80 erzielen...
#
# ... wir speichern diese Varianten in einer Liste.
variationen = [match[0] for match in match_list if match[1] >= 80]
variationen
# Anhand dieser Liste können wir nun selektiv die Schreibweise in einzelnen Zeilen ändern.
df.loc[df['Wort 2'].isin(variationen), 'New Wort 2'] = "<NAME>"
df
# **b) Ersetzen anhand einer bekannten Liste von richtigen Begriffen**
#
# Vorausgesetzt, wir kennen die Liste der tatsächlichen Begriffe, können wir auch gleich alle auf einmal handhaben:
correct_terms = [
'Tisch',
'<NAME>',
'<NAME>',
'Novartis'
]
# Die Funktion `process.extractOne()` funktioniert wie `process.extract()`, aber gibt nur den besten Match zurück:
process.extractOne("Dorf", correct_terms, scorer=fuzz.token_set_ratio)
# Wir können uns eine "Tester"-Funktion basteln:
# - Findet den besten Match aus der liste der korrekten Strings
# - Falls kein Match gut genug ist, wird das Wort nicht ersetzt
# - Falls der Match gut genug ist, wir das Wort ersetzt
def best_match(word):
match = process.extractOne(word, correct_terms, scorer=fuzz.token_set_ratio)
if match[1] < 80:
best_match = word
else:
best_match = match[0]
return best_match
# Angewendet auf die Tabelle sieht das dann so aus:
df['Wort 2'].apply(best_match)
# Speichern in einer neuen Spalte:
df['New Wort 2'] = df['Wort 2'].apply(best_match)
df
# **Aber:** Was ist, wenn wir die Liste der "richtigen" Schreibweisen gar nicht besitzen???
# +
#
# -
| 19 Geopandas/Fuzzywuzzy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import pyrfume
from pyrfume.odorants import get_cids, from_cids
cas = pd.read_csv('cas.txt', header=None)
cas = cas[0].apply(lambda x: x if '-' in x else None).dropna().values
cas_cid = get_cids(cas)
cas_cid['53896-26-7'] = 8892 # Hexanoic acid
cids = list(cas_cid.values())
molecules = pd.DataFrame(from_cids(cids)).set_index('CID')
molecules.to_csv('molecules.csv')
weights_ = pd.read_csv('dragon-weights.csv') # Extracted directly from supplement
symbols = pd.read_csv('dragon6-symbols.csv') # Looked up in Dragon 6, for comparison to other versions
weights = weights_.join(symbols[['Dragon 6.0 symbol']]).set_index('Dragon 6.0 symbol')
weights = weights.drop('Descriptor', axis=1)
weights.to_csv('weights.csv')
features = pd.read_csv('features.csv').set_index('PubChemID')
assert list(features.columns) == list(weights.index)
weighted_features = features.mul(weights['Weight'])
weighted_features = features.T.mul(weights, axis=0)
weighted_features.to_csv('feautures_weighted.csv')
# +
# Use API to compute distances
#from scipy.spatial.distance import pdist, squareform
#cids = features.index
#distances = pd.DataFrame(index=cids, columns=cids)
#distances[:] = squareform(pdist(weighted_features, metric='euclidean'))
#distances.to_csv('distances.csv')
| haddad_2008/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Train and hyperparameter tune with Scikit-learn
# ## Prerequisites
# - Create an Azure ML Workspace and setup environmnet on local computer following the steps in [Azure README.md](https://gitlab-master.nvidia.com/drobison/aws-sagemaker-gtc-2020/tree/master/azure/README.md )
# +
# verify installation and check Azure ML SDK version
import azureml.core
print('SDK version:', azureml.core.VERSION)
# -
# ## Create a FileDataset
# In this example, we will use 20 million rows (samples) of the [airline dataset](http://kt.ijs.si/elena_ikonomovska/data.html). The [FileDataset](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.data.filedataset?view=azure-ml-py) below references parquet files that have been uploaded to a public [Azure Blob storage](https://docs.microsoft.com/en-us/azure/storage/blobs/storage-blobs-overview), you can download to your local computer or mount the files to your AML compute.
# +
from azureml.core.dataset import Dataset
airline_ds = Dataset.File.from_files('https://airlinedataset.blob.core.windows.net/airline-20m/*')
# larger dataset (10 years of airline data) is also available for multi-GPU option
# airline_ds = Dataset.File.from_files('https://airlinedataset.blob.core.windows.net/airline-10years/*')
# -
# download the dataset as local files
airline_ds.download(target_path='/local/path')
# ## Initialize workspace
# Load and initialize a [Workspace](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#workspace) object from the existing workspace you created in the prerequisites step. `Workspace.from_config()` creates a workspace object from the details stored in `config.json`
# +
from azureml.core.workspace import Workspace
# if a locally-saved configuration file for the workspace is not available, use the following to load workspace
# ws = Workspace(subscription_id=subscription_id, resource_group=resource_group, workspace_name=workspace_name)
ws = Workspace.from_config()
print('Workspace name: ' + ws.name,
'Azure region: ' + ws.location,
'Subscription id: ' + ws.subscription_id,
'Resource group: ' + ws.resource_group, sep = '\n')
datastore = ws.get_default_datastore()
print("Default datastore's name: {}".format(datastore.name))
# -
# ## Upload data
# Upload the dataset to the workspace's default datastore:
path_on_datastore = 'data_airline'
datastore.upload(src_dir='/add/local/path', target_path=path_on_datastore, overwrite=False, show_progress=True)
ds_data = datastore.path(path_on_datastore)
print(ds_data)
# ## Create AML compute
# You will need to create a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) for training your model. In this notebook, we will use Azure ML managed compute ([AmlCompute](https://docs.microsoft.com/azure/machine-learning/service/how-to-set-up-training-targets#amlcompute)) for our remote training using a dynamically scalable pool of compute resources.
#
# This notebook will use 10 nodes for hyperparameter optimization, you can modify `max_node` based on available quota in the desired region. Similar to other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. [This article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) includes details on the default limits and how to request more quota.
# If we could not find the cluster with the given name, then we will create a new cluster here.`vm_size` describes the virtual machine type and size that will be used in the cluster. You will need to specify compute targets from [virtual machines in Azure](https://docs.microsoft.com/en-us/azure/virtual-machines/linux/sizes). Let's create an `AmlCompute` cluster of `Standard_DS5_v2` VMs:
# +
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
# choose a name for your cluster
cpu_cluster_name = "cpu-cluster"
if cpu_cluster_name in ws.compute_targets:
cpu_cluster = ws.compute_targets[cpu_cluster_name]
if cpu_cluster and type(cpu_cluster) is AmlCompute:
print('Found compute target. Will use {0} '.format(cpu_cluster_name))
else:
print("creating new cluster")
provisioning_config = AmlCompute.provisioning_configuration(vm_size = 'Standard_DS5_v2', max_nodes = 10, idle_seconds_before_scaledown = 300)
# create the cluster
cpu_cluster = ComputeTarget.create(ws, cpu_cluster_name, provisioning_config)
# can poll for a minimum number of nodes and for a specific timeout.
# if no min node count is provided it uses the scale settings for the cluster
cpu_cluster.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)
# use get_status() to get a detailed status for the current cluster.
print(cpu_cluster.get_status().serialize())
# -
# ## Prepare training script
# Create a project directory that will contain code from your local machine that you will need access to on the remote resource. This includes the training script and additional files your training script depends on. In this example, the training script is provided `train_sklearn_RF.py`.
# +
import os
project_folder = './train_sklearn'
os.makedirs(project_folder, exist_ok=True)
# -
# We will log some metrics by using the `Run` object within the training script:
#
# ```python
# from azureml.core.run import Run
# run = Run.get_context()
# ```
#
# We will also log the parameters and highest accuracy the model achieves:
#
# ```python
# run.log('Accuracy', np.float(accuracy))
# ```
#
#
# These run metrics will become particularly important when we begin hyperparameter tuning our model in the 'Tune model hyperparameters' section.
# Once your script is ready, copy the training script `train_sklearn_RF.py` into your project directory:
# +
import shutil
shutil.copy('../code/train_sklearn_RF.py', project_folder)
# -
# ## Train model on the remote compute
# Now that you have your data and training script prepared, you are ready to train on your remote compute.
# ### Create an experiment
# Create an [Experiment](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#experiment) to track all the runs in your workspace.
# +
from azureml.core import Experiment
experiment_name = 'train_sklearn'
experiment = Experiment(ws, name=experiment_name)
# -
# ### Create a scikit-learn Estimator
# +
from azureml.train.sklearn import SKLearn
script_params = {
'--data_dir': ds_data.as_mount(),
}
estimator = SKLearn(source_directory=project_folder,
script_params=script_params,
compute_target=cpu_cluster,
entry_script='train_sklearn_RF.py',
pip_packages=['pyarrow'])
# -
# ## Tune model hyperparameters
# We can optimize our model's hyperparameters and improve the accuracy using Azure Machine Learning's hyperparameter tuning capabilities.
# ### Start a hyperparameter sweep
# Let's define the hyperparameter space to sweep over. We will tune `n_estimators`, `max_depth` and `max_features` parameters. In this example we will use random sampling to try different configuration sets of hyperparameters and maximize `Accuracy`.
# +
from azureml.train.hyperdrive.runconfig import HyperDriveConfig
from azureml.train.hyperdrive.sampling import RandomParameterSampling
from azureml.train.hyperdrive.run import PrimaryMetricGoal
from azureml.train.hyperdrive.parameter_expressions import choice, loguniform, uniform
param_sampling = RandomParameterSampling( {
'--n_estimators': choice(range(50, 500)),
'--max_depth': choice(range(5, 19)),
'--max_features': uniform(0.2, 1.0)
}
)
hyperdrive_run_config = HyperDriveConfig(estimator=estimator,
hyperparameter_sampling=param_sampling,
primary_metric_name='Accuracy',
primary_metric_goal=PrimaryMetricGoal.MAXIMIZE,
max_total_runs=100,
max_concurrent_runs=10)
# -
# This will launch the training script with parameters that were specified in the cell above.
# start the HyperDrive run
hyperdrive_run = experiment.submit(hyperdrive_run_config)
# ## Monitor HyperDrive runs
# Monitor and view the progress of the machine learning training run with a [Jupyter widget](https://docs.microsoft.com/en-us/python/api/azureml-widgets/azureml.widgets?view=azure-ml-py).The widget is asynchronous and provides live updates every 10-15 seconds until the job completes.
# +
from azureml.widgets import RunDetails
RunDetails(hyperdrive_run).show()
# +
# hyperdrive_run.wait_for_completion(show_output=True)
# +
# hyperdrive_run.cancel()
# -
# ### Find and register best model
best_run = hyperdrive_run.get_best_run_by_primary_metric()
print(best_run.get_details()['runDefinition']['arguments'])
# List the model files uploaded during the run:
print(best_run.get_file_names())
# Register the folder (and all files in it) as a model named `train-sklearn` under the workspace for deployment
# +
# model = best_run.register_model(model_name='train-sklearn', model_path='outputs/model-sklearn.joblib')
# -
# ## Delete cluster
# +
# delete the cluster
# gpu_cluster.delete()
| azure/notebooks/HPO-SKLearn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: feml
# language: python
# name: feml
# ---
# ## Random Sample Imputation
#
# In this recipe, we will perform random sample imputation using pandas and Feature Engine.
# +
import pandas as pd
import numpy as np
# to split the data sets
from sklearn.model_selection import train_test_split
# to impute missing data with feature-engine
from feature_engine.missing_data_imputers import RandomSampleImputer
# -
# load data
data = pd.read_csv('creditApprovalUCI.csv')
data.head()
# +
# let's separate into training and testing sets
X_train, X_test, y_train, y_test = train_test_split(
data[['A2', 'A3', 'A8', 'A14']], data['A16'], test_size=0.3, random_state=0)
X_train.shape, X_test.shape
# -
X_train.dtypes
# +
# find the percentage of missing data within those variables
X_train.isnull().mean()
# -
# ## Random Sample imputation with Feature Engine
# +
# let's create a random sample imputer
imputer = RandomSampleImputer(random_state=10)
imputer.fit(X_train)
# +
# the imputer stores the train set
imputer.X.head()
# +
# transform the data - replace the missing values
X_train_t = imputer.transform(X_train)
X_test_t = imputer.transform(X_test)
# -
# check that null values were replaced
X_train_t.isnull().mean()
imputer_obs = RandomSampleImputer(random_state=['A8', 'A3'], seed='observation', seeding_method='add')
imputer_obs.fit(X_train)
X_train_tt = imputer_obs.transform(X_train)
X_test_tt = imputer_obs.transform(X_test)
# check that null values were replaced
X_train_tt.isnull().mean()
pd.concat([
X_train_tt[X_train['A2'].isnull()][['A2', 'A3', 'A8']],
X_train_t[X_train['A2'].isnull()]['A2']],
axis=1
)
tmp = pd.DataFrame({'A2':np.nan, 'A3': 3, 'A8':20, 'A14': np.nan}, index=[1])
tmp
for i in range(0,10):
print(imputer.transform(tmp))
for i in range(0,10):
print(imputer_obs.transform(tmp))
for i in range(0,10):
print(imputer_obs.transform(tmp))
| examples/RandomSampleImputer_SeedTest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import sys
sys.path.append("../scripts")
import evaluate
import utils
import tensorflow as tf
import numpy as np
# import matplotlib
import seaborn as sns
import matplotlib.pyplot as plt
import glob, os
import saliency_embed
testset, targets = evaluate.collect_whole_testset('../../datasets/quantitative_data/testset/', coords=True)
np_C, np_X, np_Y = utils.convert_tfr_to_np(testset)
# load and get model layer
run_path = glob.glob('../../trained_models/**/run-20211023_095131-w6okxt01', recursive=True)[0]
layer = -3
model, bin_size = utils.read_model(run_path, compile_model=False)
aux_model = tf.keras.Model(inputs=model.inputs, outputs=model.layers[layer].output)
# +
threshold = 2
cell_line = 13
print(targets[cell_line])
thresholded_C, thresholded_X, thresholded_Y = utils.threshold_cell_line_np(np_C,
np_X,
np_Y,
cell_line,
more_than=threshold)
idr_class = saliency_embed.label_idr_peaks(thresholded_C, cell_line, bedfile1='../../datasets/quantitative_data/testset/sequences.bed',
bedfile2='../../datasets/quantitative_data/cell_line_testsets/cell_line_13/complete/peak_centered/i_2048_w_1.bed',
fraction_overlap=0.5)
predictions = utils.predict_np(thresholded_X, model, batch_size=32, reshape_to_2D=False)
interm_representations = utils.predict_np(thresholded_X, aux_model, batch_size=32, reshape_to_2D=True)
thresholded_Y.shape
# -
embeddings = saliency_embed.get_embeddings(interm_representations)
embeddings['IDR'] = idr_class
embeddings['cell line'] = targets[cell_line]
embeddings.to_csv('../tutorial_outputs/UMAP_embeddings.csv', index=None)
# +
fig, ax = plt.subplots(1,figsize=[6,5])
ax = sns.scatterplot(data=embeddings,
x='UMAP 1', y='UMAP 2',
alpha=0.1,
color='grey',
ax=ax)
ax.set_axis_off()
# +
saliency_cell_line_dir = utils.make_dir('../tutorial_outputs/saliency_maps')
sets_of_ids = {'test_maps':[1000, 10001]}
for label,selected_sample_ids in sets_of_ids.items():
output_path = os.path.join(saliency_cell_line_dir, label+'.svg')
print(output_path)
explainer = saliency_embed.Explainer(model, class_index=cell_line)
X_sample = thresholded_X[selected_sample_ids]
saliency_scores = explainer.saliency_maps(X_sample)
saliency_embed.plot_saliency_logos_oneplot(saliency_scores, X_sample, window=256,
titles = selected_sample_ids,
filename=output_path)
# -
| tutorials/umap_and_saliency.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# single picture
import robosoc2d
import robosoc2dplotty.plotty as r2plotty
import time
sim_handle = robosoc2d.build_simpleplayer_simulator([], 4, [], 4)
robosoc2d.simulator_step_if_playing(sim_handle)
fig = r2plotty.draw(sim_handle, draw_numbers=True)
r2plotty.show(fig)
# +
# whole game (CAREFUL THIS MAY HANG YOUR NOTEBOOK !)
import robosoc2d
import robosoc2dplotty.plotty as r2plotty
sim_handle = robosoc2d.build_simpleplayer_simulator([], 4, [], 4)
r2plotty.play_whole_game_in_notebook(sim_handle)
# +
# equivalent to the previous cell but without using "play_whole_game_in_notebook()", in case you need to process something in the inner loop.
import robosoc2d
import robosoc2dplotty.plotty as r2plotty
import time
sim_handle = robosoc2d.build_simpleplayer_simulator([], 4, [], 4)
fig = r2plotty.draw(sim_handle)
display(fig)
while(robosoc2d.simulator_step_if_playing(sim_handle)):
r2plotty.update(sim_handle, fig)
time.sleep(0.01)
# +
# step by step update clicking a button
import robosoc2d
import robosoc2dplotty.plotty as r2plotty
sim_handle = robosoc2d.build_simpleplayer_simulator([], 4, [], 4)
r2plotty.play_steps_in_notebook(sim_handle)
| pyextension/robosoc2dplotty/test/test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: metoffice_datasets
# language: python
# name: metoffice_datasets
# ---
# # intake_informaticslab
#
# `intake_informaticslab` is an [intake](https://intake.readthedocs.io/en/latest/) catalogue and driver to access hundreds of terabytes of Met Office data presented as coherent [Analysis Ready Cloud Optimised](https://medium.com/informatics-lab/analysis-ready-data-47f7e80cba42) datasets.
#
# For details on the datasets and the licences associated with them see the [README](https://github.com/informatics-lab/intake_informaticslab#readme)
#
# These notebooks demonstrate some of the functionality available through this library and datasets. Many examples have been truncated so they consume less memory so they work on a free [MyBinder](https://mybinder.org/) instance.
#
# These datasets are kindly hosted by Microsoft Azure as part of the [AI For Earth initiative](https://www.microsoft.com/en-us/ai/ai-for-earth).
#
# These datasets are hosted in the Azure East US 2 region. The performance will greatly depend on your connectivity to this region. For best performance, work in the Azure East US 2 region.
# ## Import some things we'll need and set some plot options
# fix for display issue
import xarray as xr
import numpy as np
import intake
import matplotlib
matplotlib.rcParams['figure.figsize'] = (15.0, 15.0)
# ## Data access is as easy as installing a package
# +
## Un-comment to use your prefered package manager. If launched via MyBinder this step can be skipped.
# # %pip install intake_informaticslab --upgrade
# # %conda install -c conda-forge -c informaticslab intake_informaticslab --upgrade
# -
# ## Explore the datasets available
# +
def print_cat_items(cat, indent=0):
if(isinstance(cat, intake.catalog.Catalog)):
for cat_item in list(cat):
print(" "*indent + cat_item)
print_cat_items(cat[cat_item], indent+2)
print_cat_items(intake.cat)
# -
# ## Some datasets might require accepting a licence
# ### A `LicenseNotExceptedError` will be raised if a licence needs accepting
## This line would raise a LicenseNotExceptedError
intake.cat.met_office.weather_forecasts.mogreps_g.single_level.to_dask()
# ### View the dataset's license
intake.cat.met_office.weather_forecasts.mogreps_g.single_level.license
# ### Accept the license with the `license_accepted` flag
intake.cat.met_office.weather_forecasts.mogreps_g.single_level(license_accepted=True).to_dask()
# ## See the other examples for some demonstrations of using the data
| binder/Introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=["intro"]
# # Notes
#
# Different problems give different number of points: 2, 3 or 4.
#
# Please, fill `STUDENT` variable with your name, so that we call collect the results automatically. Each problem contains specific validation details. We will do our best to review your assignments, but please keep in mind, that for this assignment automatic grade (between $0$ an $1$) is the primary source of ground truth.
# -
# %pylab inline
plt.style.use("bmh")
plt.rcParams["figure.figsize"] = (6,6)
import numpy as np
import torch
# + tags=["parameters"]
STUDENT = "<NAME>"
ASSIGNMENT = 2
TEST = False
# -
if TEST:
import solutions
total_grade = 0
MAX_POINTS = 19
# # NumPy broadcasting
# + [markdown] tags=["problem"]
# ### 1. Normalize matrix rows (2 points).
#
# For 2-dimensional array `arr`, calculate an array, in which each row is a normalized version of corresponding row from `arr`.
#
# For example, for `(3,4)` input array, the output is also `(3,4)` and `out_arr[0] = (arr[0] - np.mean(arr[0])) / np.std(arr[0])` and so on for other rows.
#
# Result must be **2-dimensional**, and **will be tested against three random combinations of input array dimensions ($10 \leq n < 100 $)**. Array values will be drawn from a normal distribution (`np.random.normal`) with random mean and standard deviation.
# + tags=["solution"]
def norm_rows(arr):
# your code goes here
return (arr - arr.mean(axis=1).T)/(arr.std(axis=1).T)
# + tags=["validation"]
PROBLEM_ID = 1
if TEST:
total_grade += solutions.check(STUDENT, PROBLEM_ID, norm_rows)
# + [markdown] tags=["problem"]
# ### 2. Normalize matrix columns (2 points).
#
# Similar to Problem 1, but normalization must be performed along columns.
#
# For example, for `(3,4)` input array, the output is also `(3,4)` and `out_arr[:, 0] = (arr[:, 0] - np.mean(arr[:, 0])) / np.std(arr[:, 0])` and so on for other columns.
#
# Result must be **2-dimensional**, and **will be tested against three random combinations of input array dimensions ($10 \leq n < 100 $)**. Array values will be drawn from normal distribution (`np.random.normal`) with random mean and standard deviation.
# + tags=["solution"]
def norm_cols(arr):
# your code goes here
return (arr - arr.mean(axis=0).T)/(arr.std(axis=0).T)
# + tags=["validation"]
PROBLEM_ID = 2
if TEST:
total_grade += solutions.check(STUDENT, PROBLEM_ID, norm_cols)
# + [markdown] tags=["problem"]
# ### 3. Generic normalize routine (2 points).
#
# Similar to Problems 1 and 2, but normalization must be performed according to `axis` argument. `axis=0` means normalization along the columns, and `axis=1` means normalization along the rows.
# + tags=["solution"]
def norm(arr, axis):
# your code goes here
return (arr - arr.mean(axis=axis).T)/(arr.std(axis=axis).T)
# + tags=["validation"]
PROBLEM_ID = 3
if TEST:
total_grade += solutions.check(STUDENT, PROBLEM_ID, norm)
# + [markdown] tags=["problem"]
# ### 4. Dot product of matrix and vector (2 points).
#
# Calculate dot product of 2-dimensional array $M$ of shape $(N,K)$ and 1-dimensional row vector $v$ of shape $(K,)$. You cannot use `np.dot` in this exercise.
#
# Result must be **1-dimensional** of shape $(N,)$, and **will be tested against three random combinations of input arrays dimensions ($10 \leq n < 100 $)**. Arrays values will be drawn from standard normal distribution (`np.random.randn`).
# + tags=["solution"]
def dot(m, v):
# your code goes here
return (m*v).sum(axis=1)
# + tags=["validation"]
PROBLEM_ID = 4
if TEST:
total_grade += solutions.check(STUDENT, PROBLEM_ID, dot)
# + [markdown] tags=["problem"]
# ### 5. Calculate recurrence matrix (3 points).
#
# In signals (or time series) analysis, it's usualy important to quickly assess the structure (if any) of the data. This can be done in many different ways. You can test, whether a signal is stationary or look at Fourier transform to understand the frequency composition of a signal. When you want to understand, whether signal contains some recurring pattern, it's useful to perform what is called *recurrent quantification analysis*.
#
# Imagine a signal $s_i$. Recurrence matrix is then:
#
# $$
# R_{ij} = \left\{
# \begin{array}{l}
# 1, |s_i-s_j|<\varepsilon \\
# 0, |s_i-s_j|\ge\varepsilon \\
# \end{array}
# \right.
# $$
#
# In this exercise you need to implement a function, which calculates recurrence matrix for 1-dimensional array. The function should not use any loops and must leverage broadcasting. For reference, naive loop implementation is provided below. Plot recurrence matrices for some signals to understand, how signal structure reveals itself in the recurrence matrix.
#
# For example, for a signal of shape $(100,)$ result must be of the shape $(100, 100)$. Result must be **2-dimensional**, and **will be tested against three random combinations of input array dimensions ($100 \leq n < 1000 $)** with different signal patterns (noise, $\sin$, noise + randomly-placed recurrent pattern).
# -
def recm_naive(ts, eps):
"""Loop implementation of recurrent matrix."""
ln = len(ts)
rm = np.zeros((ln, ln), dtype=bool)
for i in range(ln):
for j in range(ln):
rm[i, j] = np.abs(ts[i]-ts[j])<eps
return rm
random_signal = np.random.randn(200)
plt.imshow(recm_naive(random_signal, 1e-1), cmap=plt.cm.binary)
sin_signal = np.sin(np.arange(1000))
plt.imshow(recm_naive(sin_signal, 1e-1), cmap=plt.cm.binary)
# +
random_signal = np.random.randn(200)
random_signal[6:21] = 5 * np.ones((15,))
random_signal[93:108] = 5 * np.ones((15,))
random_signal[39:54] = 0.5 * np.ones((15,))
random_signal[162:177] = 0.5 * np.ones((15,))
plt.plot(random_signal)
plt.show()
plt.imshow(recm_naive(random_signal, 5e-1), cmap=plt.cm.binary);
# + tags=["solution"]
def recm(ts, eps):
# your code goes here
pass
# + tags=["validation"]
PROBLEM_ID = 5
if TEST:
total_grade += solutions.check(STUDENT, PROBLEM_ID, recm)
# -
# # PyTorch
# + [markdown] tags=["problem"]
# ### 6. ReLU activation (2 points).
#
# ReLU is the most commonly used activation function in many deep learning application. It's defined as
#
# $$
# ReLU(x) = \max(0, x).
# $$
#
# Outpu must be of the same shape as input, and **will be tested against three random combinations of input array dimensions ($100 \leq n < 1000 $)**, while values of the input are drawn from standard normal distribution. Number of dimensions of the input will also be selected randomly and is either 1, 2 or 3.
# + tags=["solution"]
def relu(arr):
# your code goes here
pass
# + tags=["validation"]
PROBLEM_ID = 6
if TEST:
total_grade += solutions.check(STUDENT, PROBLEM_ID, relu)
# -
# ### 7. Mean squared error (2 points).
#
# In this problem you need to calculate MSE for a pair of tensors `y_true` and `y_pred`. MSE is defined as usual:
#
# $$
# L_{MSE} = \frac{1}{N} \sum_i \left(y_i - \hat y_i\right)^2
# $$
#
# Note, however, that `y_true` and `y_pred`may be of **different shape**. While `y_true` is always $(N,)$, `y_pred` may be $(N,1)$, $(1, N)$ or $(N,)$. Input values are drawn from standard normal distribution and **shape is selected randomly ($100 \leq n < 1000 $)**.
# + tags=["solution"]
def mse(y_true, y_pred):
# your code goes here
pass
# + tags=["validation"]
PROBLEM_ID = 7
if TEST:
total_grade += solutions.check(STUDENT, PROBLEM_ID, mse)
# + [markdown] tags=["problem"]
# ### 8. Character-level encoding (4 points).
#
# In computations in general and in machine learning specifically letters cannot be used directly, as computers only know aboun numbers. Text data may be encoded in many different ways in natural language processing tasks.
#
# One of the simplest ways to encode letters is to use one-hot encoded representation, with letters being "class labels". A letter is represented by a tensor of shape $(26,)$.
#
# Then, for example, word "python" would be transformed into a tensor of shape $(6, 26)$ with all elements being $0$, except $(0, 15)\sim p,\,(1, 24)\sim y,\,(2, 19)\sim t,...$ being $1$. A phrase would be represented with 3-dimensional tensor.
#
# In this problem you need to create a tensor, which represents a list of words `words` of length $N$. The only characters used are those from `string.ascii_lowercase`, and words are of different length $L_i$. Output must be of shape $(N, \max(L_i), 26)$.
#
# Dimension 0 corresponds to words themselves, with `tensor[0]` being a represetation of `words[0]`. Note, that you need to use padding: although trivial in this case, you must remember, that tensor must accomodate for a longest word, thus dimension 1 is $\max(L_i)$.
#
# Note also, that the only loop you need here is a loop over `words`, there's no need to loop over the resulting tensor.
#
# The result will be tested against three predefined lists of word, with all words being lowercase and containing only ASCII characters.
# + tags=["solution"]
def encode(words):
# your code goes here
pass
# + tags=["validation"]
PROBLEM_ID = 8
if TEST:
total_grade += solutions.check(STUDENT, PROBLEM_ID, encode)
# -
# # Your grade
if TEST:
print(f"{STUDENT}: {int(100 * total_grade / MAX_POINTS)}")
| [Py4DP] [Lecture-2] Graded Assignment-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## The following tables/csv files will be created after running this notebook:
# Product
#
# This script reads the product.csv and cleans the rating as well the categories. It overwrites the originall Product.csv.
#
# The original schema contains: 'uniq_id','Product_Name','Manufacturer','Price','average_review_rating','amazon_category_and_sub_category','description'
#
#
# The new schema contains:
# 'uniq_id','Product_Name','Manufacturer','Price','average_review_rating','cat1','cat2','cat3','description'
import pandas as pd
import numpy as np
df = pd.read_csv("product.csv")
df.head()
def clean_rating(x):
if (type(x)==str):
score,*_ = x.split(" ")
return float(score)
def clean_average_review_rating(df):
df["average_review_rating"] = df["average_review_rating"].apply(lambda x:clean_rating(x))
return df
df = clean_average_review_rating(df)
# +
def process_category_value(x,cat1,cat2,cat3):
'''
if x is null, all 3 categories are "All"
if x has more than 3 categories, only first 3 categories are kept and assigned to
cat1, cat2, cat3.
if x has less than 3 categories, "All" will be appended as needed
'''
default = ["All","All","All"]
if (x == x):
all_categories = x.split(" > ")
for i in range(len(all_categories)):
default[i] = all_categories[i]
if (i == 2):
break;
cat1.append(default[0])
cat2.append(default[1])
cat3.append(default[2])
def process_category_columns(df):
'''
create 3 empty lists and apply process_category_value.
append 3 new columns to df and drop the orignal category column
'''
cat1 = []
cat2 = []
cat3 = []
df.amazon_category_and_sub_category.apply(lambda x:process_category_value(x,cat1,cat2,cat3))
df["Cat1"] = cat1
df["Cat2"] = cat2
df["Cat3"] = cat3
df.drop(columns = "amazon_category_and_sub_category",inplace = True)
return df
# -
df = process_category_columns(df)
df.head()
df.to_csv("product.csv")
| data_cleaning/clean_product.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# In this notebook I explore some properties about the generation process for paraphrase generation. I also answer some questions I had.
# Note: some of the asserts aren't up to date
# ## Setup
# %load_ext autoreload
# %autoreload 2
# %load_ext line_profiler
# +
## Imports and environment variables
import os
os.environ["TOKENIZERS_PARALLELISM"] = "true" # set to false if not working
# Core imports
import torch, numpy as np, pandas as pd, gc,sys, logging, warnings
from torch.utils.data import DataLoader, RandomSampler
from torch.distributions import Categorical
from datasets import load_dataset, load_metric, load_from_disk, DatasetDict
from transformers import (AutoModelForSeq2SeqLM, AutoModelForSequenceClassification,
AutoTokenizer, AdamW, SchedulerType, get_scheduler)
from sentence_transformers import SentenceTransformer
from sentence_transformers.util import pytorch_cos_sim
from collections import defaultdict
from accelerate import Accelerator, notebook_launcher
from cachetools import cached, LRUCache
from types import MethodType
from timeit import default_timer as timer
import utils; from utils import * # local script
from tqdm.auto import tqdm
import itertools
import copy
import wandb
from pprint import pprint
from undecorated import undecorated
# Dev imports (not needed for final script)
import seaborn as sns
from IPython.display import Markdown
from pprint import pprint
from IPython.core.debugger import set_trace
from GPUtil import showUtilization
import torchsnooper
# -
logging.basicConfig(format='%(message)s')
logger = logging.getLogger("main_logger")
logger.setLevel(logging.INFO)
# +
# options for the pp_model
# 1. tuner007/pegasus_paraphrase
# 2. tdopierre/ProtAugment-ParaphraseGenerator
# 3. eugenesiow/bart-paraphrase
## PEGASUS model
pp_name = "tuner007/pegasus_paraphrase"
pp_tokenizer_pegasus = AutoTokenizer.from_pretrained(pp_name)
pp_model_pegasus = AutoModelForSeq2SeqLM.from_pretrained(pp_name, local_files_only=True)
generate_with_grad = undecorated(pp_model_pegasus.generate)
pp_model_pegasus.generate_with_grad = MethodType(generate_with_grad, pp_model_pegasus)
## BART model
pp_name = "eugenesiow/bart-paraphrase"
pp_tokenizer_bart = AutoTokenizer.from_pretrained(pp_name)
pp_model_bart = AutoModelForSeq2SeqLM.from_pretrained(pp_name, local_files_only=True)
generate_with_grad = undecorated(pp_model_bart.generate)
pp_model_bart.generate_with_grad = MethodType(generate_with_grad, pp_model_bart)
# -
# ## Functions
# +
def get_pp_logp(translated):
"""log(p(pp|orig)) basically.
works for greedy search, will need tweaking for other types probably"""
seq_without_first_tkn = translated.sequences[:, 1:]
attention_mask = pp_model._prepare_attention_mask_for_generation(
seq_without_first_tkn, pp_tokenizer.pad_token_id, pp_tokenizer.eos_token_id
)
scores_log_softmax = torch.stack(translated.scores, 1).log_softmax(2)
seq_token_log_probs = torch.gather(scores_log_softmax,2,seq_without_first_tkn[:,:,None]).squeeze(-1)
del scores_log_softmax
# account for nan values by setting them to 0 (maybe a bit of a hack)
# will also handle inf and -inf values too by default
seq_token_log_probs = torch.nan_to_num(seq_token_log_probs)
# account for the padding tokens at the end
seq_token_log_probs = seq_token_log_probs * attention_mask
seq_log_prob = seq_token_log_probs.sum(-1)
# if np.any(np.isnan(seq_log_prob.detach().cpu()).tolist()):
# warnings.warn(f"Warning: NAN's detected in pp_logp calclulations.\n seq_token_log_probs: {seq_token_log_probs}")
return seq_log_prob
def get_tokens_from_token_ids_batch(tokenizer, ids_batch):
l = []
for i in range(ids_batch.shape[0]):
l.append(tokenizer.convert_ids_to_tokens(ids_batch[i,:]))
return l
def get_start_end_special_token_ids(tokenizer):
"""The token id's that input/output sequences should start and end with"""
d = {}
if pp_tokenizer.name_or_path in ['eugenesiow/bart-paraphrase', 'tdopierre/ProtAugment-ParaphraseGenerator']:
d["input_start_id"] = tokenizer.bos_token_id
d["input_end_id"] = [tokenizer.pad_token_id, tokenizer.eos_token_id]
d["output_start_id"] = tokenizer.eos_token_id
d["output_end_id"] = [tokenizer.pad_token_id, tokenizer.eos_token_id]
elif pp_tokenizer.name_or_path == "tuner007/pegasus_paraphrase":
d["input_start_id"] = None
d["input_end_id"] = [tokenizer.pad_token_id, tokenizer.eos_token_id]
d["output_start_id"] = tokenizer.pad_token_id
d["output_end_id"] = [tokenizer.pad_token_id, tokenizer.eos_token_id]
else:
raise Exception("unrecognised tokenizer")
return d
def check_no_nans_or_infs(x):
assert torch.all(~torch.isnan(x))
assert torch.all(~torch.isneginf(x))
assert torch.all(~torch.isposinf(x))
def assert_start_and_end_tokens_are_correct(tokenizer, orig_token_ids, pp_token_ids):
"""Make sure input sequences (orig) and output sequences (pp) start and end with the
right special tokens (depends on tokenizer)"""
start_end_token_d = get_start_end_special_token_ids(pp_tokenizer)
# Input
if start_end_token_d['input_start_id'] is not None:
assert torch.all(orig_token_ids[:,0] == start_end_token_d['input_start_id'])
# can probs rewrite this to make it nicer but it's fine for now
assert torch.all(torch.logical_or(orig_token_ids[:,-1] == start_end_token_d['input_end_id'][0],
orig_token_ids[:,-1] == start_end_token_d['input_end_id'][1]))
# Output
assert torch.all(pp_token_ids[:,0] == start_end_token_d['output_start_id'])
assert torch.all(torch.logical_or(pp_token_ids[:,-1] == start_end_token_d['output_end_id'][0],
pp_token_ids[:,-1] == start_end_token_d['output_end_id'][1]))
def check_scores_for_posinf_nan_and_unexpected_neginf(scores_stacked):
"""Check we don't have any postive inf or nan, and that all negative inf values are expected"""
assert torch.all(~torch.isposinf(scores_stacked))
assert torch.all(~torch.isnan(scores_stacked))
# We expect to see negative inf for the eos_token when we have not reached min_length.
# But we shouldn't expect it for any other tokens
idx_neginf = torch.nonzero(torch.isneginf(scores_stacked))
assert torch.all(idx_neginf[:,2] == pp_tokenizer.eos_token_id)
# Rough check that all idx before min_length are -inf for all elements in batch
# We do min_length - 1 because sequences are allowed to have length min_length so that idx
# shouldn't be set to -inf
# Not a 100% test but very likely to identify
assert idx_neginf.shape[0] == (pp_model_params["min_length"] -1) * batch_size
# Check that no elements after min_length are -inf
assert torch.all(idx_neginf[:,1] < (pp_model_params["min_length"] -1 ))
def check_scores_log_softmax_sums_and_shape(scores_log_softmax):
sums = scores_log_softmax.exp().sum(2)
# check that the axes is right
# we want to sum over token probabilities at each generation step, so we
# should end up with a shape [batch_size, generated_length]
assert sums.shape[0] == batch_size
assert sums.shape[1] == generated_length - 1
# check that they sum to 1 along the generated_length axis
assert torch.allclose(sums, torch.ones(sums.size()), atol = 1e-4)
def check_seq_token_log_prob_values_are_correct():
"""Just enumerates and checks values
Quite slow for large batches so run as a test rather than an assert in every batch.
"""
l = []
for i_ex in range(batch_size):
for i_step in range(generated_length - 1):
i_tkn = seq_without_first_tkn[i_ex][i_step].item()
l.append(scores_log_softmax[i_ex,i_step, i_tkn] == seq_token_log_probs[i_ex,i_step])
assert all(l)
def pretty_print_pp_batch_and_next_token_probabilities():
"""Goes through each paraphrase and shows at each timestep the next likely tokens.
Only will work for greedy search.
e.g. [
"<pad> ['▁My, 0.289', '▁I, 0.261', '▁Hello, 0.07'] | Entropy: 4.23 ",
"<pad> My ['▁name, 0.935', '▁Name, 0.005', 'name, 0.002'] | Entropy: 0.80 "
]
"""
str_d = defaultdict(list)
for i_tkn in range(0, generated_length-1):
ids = pp_output.sequences[:, :(i_tkn+1)]
partial_pp = pp_tokenizer.batch_decode(ids)
kth_ids,kth_probs = tkn_kmaxidx[:, i_tkn, :], tkn_kmaxprob[:, i_tkn, :]
kth_tkns = get_tokens_from_token_ids_batch(pp_tokenizer, kth_ids)
# enumerates examples in batch
z = zip(partial_pp, kth_tkns, kth_probs, ent.detach())
for i_ex, (ex_sen, ex_next_tkns, ex_next_probs, ex_e) in enumerate(z):
# Form nice formatted string mixing together tokens and probabilities
tkn_tuples_l = [(tkn, round_t(prob,3)) for tkn, prob in zip(ex_next_tkns, ex_next_probs)]
tkn_str = ['%s, %s' % t for t in tkn_tuples_l]
# Add to dict of lists and add on entropy term.
str_d[i_ex].append(f"{ex_sen} {tkn_str} | Entropy: {ex_e[i_tkn]:.2f} ")
for v in str_d.values(): pprint(v)
# -
# ## Greedy search for paraphrase generation
# +
# INPUT AND PARAMETERS
orig_l = [
"Look at the bird over there with the red and yellow stripes.",
"That girl has her hair dyed yellow - how interesting."
]
pp_model_params = {
"num_beams": 1,
"num_return_sequences": 1,
"num_beam_groups": 1,
"diversity_penalty": 0., # must be a float
"temperature": 1.5,
"length_penalty" : 1,
"min_length" : 5
}
## Select which model/tokenizer to use
pp_tokenizer = pp_tokenizer_pegasus
pp_model = pp_model_pegasus
# +
#### TOKENIZER INFORMATION #####
logger.info("\n############################## TOKENIZER ########################################\n")
logger.info(f"We are using the {pp_tokenizer.name_or_path} tokenizer")
logger.info(f"Tokenizer has these special tokens:{pp_tokenizer.all_special_tokens}")
logger.info(f"The bos token is {pp_tokenizer.bos_token} and has id {pp_tokenizer.bos_token_id}")
logger.info(f"The eos token is {pp_tokenizer.eos_token} and has id {pp_tokenizer.eos_token_id}")
logger.info(f"The pad token is {pp_tokenizer.pad_token} and has id {pp_tokenizer.pad_token_id}")
logger.info(f"The unk token is {pp_tokenizer.unk_token} and has id {pp_tokenizer.unk_token_id}")
#### INPUT #####
batch_size = len(orig_l)
orig_tokens = pp_tokenizer(orig_l, return_tensors='pt', padding=True, pad_to_multiple_of=4)
input_length = orig_tokens['input_ids'].size()[1]
orig_l_tokens_list = get_tokens_from_token_ids_batch(pp_tokenizer, orig_tokens['input_ids'])
logger.info("\n############################### INPUT #######################################\n")
logger.info(f"Original text: {orig_l}")
logger.info(f"Batch size is: {batch_size}")
logger.info(f"This is tokenised to get a dict with keys {orig_tokens.keys()} which should be input_ids and attention_mask ")
logger.info(f"The input_ids look like this: {orig_tokens['input_ids']}")
logger.info(f"The tokens are: {orig_l_tokens_list}")
logger.info(f"This has shape {orig_tokens['input_ids'].shape} or [batch_size, input_length], which also\
might be padded to hit a padding multiple (so input_length is not just the longest example length in the batch).")
logger.info(f"Input length is: {input_length}")
logger.info(f"The attention_mask looks like this: {orig_tokens['attention_mask']}")
logger.info(f"This has shape {orig_tokens['attention_mask'].shape} or [batch_size, input_length]")
##### PARAPHRASE #####
pp_output = pp_model.generate_with_grad(**orig_tokens, **pp_model_params, do_sample=False,
return_dict_in_generate=True,
output_scores=True,
remove_invalid_values=False)
generated_length = pp_output.sequences.shape[1]
pp_l = pp_tokenizer.batch_decode(pp_output.sequences, skip_special_tokens=True)
pp_l_with_tokens = pp_tokenizer.batch_decode(pp_output.sequences, skip_special_tokens=False)
pp_l_tokens_list = get_tokens_from_token_ids_batch(pp_tokenizer, pp_output.sequences)
assert_start_and_end_tokens_are_correct(pp_tokenizer, orig_token_ids=orig_tokens['input_ids'],
pp_token_ids=pp_output.sequences)
logger.info("\n#################################### PARAPHRASES ##################################\n")
logger.info(f"Paraphrases: {pp_l}")
logger.info(f"Output has keys {pp_output.keys()}")
logger.info(f"Paraphrases with special tokens: {pp_l_with_tokens}")
logger.info(f"List of pp tokens:{pp_l_tokens_list}")
logger.info(f"Paraphrase token sequences: {pp_output.sequences}")
logger.info(f"Shape of pp token sequences:{pp_output.sequences.shape} or [batch_size, generated_length]")
logger.info(f"Generated length: {generated_length}")
###### SCORES AND PROBABILITIES ########
scores_stacked = torch.stack(pp_output.scores, 1)
# The second argument to stack (i.e. dim) determines which axis the tensors are stacked along.
# It determines the axis that becomes generated_length - 1
# dim=0 gives shape [generated_length-1, batch_size, vocab_size]
# dim=1 gives shape [batch_size, generated_length-1, vocab_size]
# dim=2 gives shape [batch_size, vocab_size, generated_length-1]
# Our scores_stacked is stacked on dim 1 so it should be second
assert scores_stacked.shape == torch.Size([batch_size, (generated_length - 1), pp_tokenizer.vocab_size])
#check_scores_for_posinf_nan_and_unexpected_neginf(scores_stacked)
# These scores are logits
# see some of the docs on this page https://huggingface.co/docs/transformers/v4.16.2/en/main_classes/output#transformers.modeling_outputs.Seq2SeqModelOutput
# so we got to take softmax over them
# but if we take regular softmax then we run into numerical errors
# so instead we take log_softmax
scores_log_softmax = torch.log_softmax(scores_stacked, 2)
check_scores_log_softmax_sums_and_shape(scores_log_softmax)
####### SEQUENCE PROBABILITIES #######
# We select the token probability corresponding to each token
# However because the scores represent transitions we need to remove the first token from each
# sequence to match them up.
seq_without_first_tkn = pp_output.sequences[:,1:]
assert seq_without_first_tkn.shape == torch.Size([batch_size, generated_length - 1])
# Now select prob corresponding to each token
seq_token_log_probs = torch.gather(scores_log_softmax,2,seq_without_first_tkn[:,:,None]).squeeze(-1)
assert seq_token_log_probs.shape == seq_without_first_tkn.shape # probs should be 1-1 with the filtered tkns
#check_no_nans_or_infs(seq_token_log_probs)
# Check that the last token probability corresponds to a possible end token
output_end_ids = get_start_end_special_token_ids(pp_tokenizer)['output_end_id']
assert all([o in scores_log_softmax[:, -1, output_end_ids] for o in seq_token_log_probs[:,-1]])
check_seq_token_log_prob_values_are_correct()
# The attention mask has 1 everywhere except for where padding tokens occur, where it has 0.
# It is used to filter out padding tokens from the sequence probablity because then the sequence
# probability will depend on how many padding tokens there are and the probability of generating them,
# which (a) we don't want and (b) the probability isn't correct anyway
attention_mask = pp_model._prepare_attention_mask_for_generation(
seq_without_first_tkn, pp_tokenizer.pad_token_id, pp_tokenizer.eos_token_id
)
seq_token_log_probs = seq_token_log_probs * attention_mask
# check attention mask only has 0 for padding tokens and not eos tokens or anything else
assert all(seq_without_first_tkn[attention_mask == 0] == pp_tokenizer.pad_token_id)
assert seq_token_log_probs.shape == attention_mask.shape == seq_token_log_probs.shape
#assert torch.all(seq_token_log_probs > -10) # we shouldn't be picking extrememly rare tokens
seq_log_prob = seq_token_log_probs.sum(-1)
assert seq_log_prob.shape == torch.Size([batch_size])
#check_no_nans_or_infs(seq_log_prob)
logger.info("\n########################## SCORES AND PROBABILITIES ####################################\n")
logger.info(f"Scores is a tuple of length {len(pp_output.scores)} which is one less than the generated_length, or \
the number of tokens in the pp token sequences (this has shape {pp_output.sequences.shape}")
logger.info(f"Each score is a tensor of shape {pp_output.scores[0].shape} or [batch_size, vocab_size]")
#logger.info(f"Full shape:{[o.shape for o in pp_output.scores]}")
logger.info(f"We stack them to get a tensor of shape {scores_stacked.shape} or [batch_size, generated_length - 1, vocab_size]")
logger.info(f"Scores are really logits so we have to take softmax to get probabilities. ")
logger.info("But if we take regular softmax then we run into numerical errors so we take log softmax")
logger.info("We then select the token probability corresponding to each token and sum them to get the log \
probability of the sequence.")
############# ENTROPY AND TOKEN PROBABILITIES ####################
ent = Categorical(logits = scores_stacked).entropy()
assert ent.shape == torch.Size([batch_size, generated_length - 1])
scores_softmax = scores_log_softmax.exp()
k=3
tkn_kmaxprob, tkn_kmaxidx = torch.topk(scores_softmax, k=k, dim=2)
tkn_kmaxprob = tkn_kmaxprob.detach() # log these
# The third dimension indexes top1, top2, top3 etc
assert tkn_kmaxprob[:,:,0].shape == torch.Size([batch_size, generated_length - 1])
# I'd naively expect True everywhere for tkn_kmaxidx[:,:,0] == pp_output.sequences[:, 1:] but it turns
# out this is not the case because padding tokens seem to have prob 0 and eos tokens are outputted
# instead by the token generation process and then later replaced by pad
#Uncomment to show how paraphrase is formed.
logger.info("\n########################## ENTROPY AND TOKEN PROBABILITIES ####################################\n")
logger.info(f"Originals: {orig_l}")
pretty_print_pp_batch_and_next_token_probabilities()
# -
# ## Beam search for paraphrase generation
orig_l = [
"Look! A small dog. Isn't it cute?",
"Far out, if I have to write another sentence...it'll be bad."
]
n_seq = 5
pp_model_params = {
"num_beams": 5,
"num_return_sequences": n_seq,
"num_beam_groups": 1,
"diversity_penalty": 0.,
"temperature": 1.5,
"length_penalty" : 0,
"min_length" : 5
}
batch_size = len(orig_l)
logger.info(f"Input: {orig_l}")
orig_tokens = pp_tokenizer(orig_l, return_tensors='pt', padding=True, pad_to_multiple_of=4)
input_length = orig_tokens['input_ids'].shape[1]
pp_output = pp_model.generate_with_grad(**orig_tokens, **pp_model_params, do_sample=False,
return_dict_in_generate=True,
output_scores=True,
remove_invalid_values=False)
logger.info(f"Input: {orig_l}")
# +
generated_length = pp_output.sequences.shape[1]
assert pp_output.sequences.shape == torch.Size([batch_size * n_seq, generated_length ])
pp_l_with_tokens = pp_tokenizer.batch_decode(pp_output.sequences, skip_special_tokens=False)
print("Paraphrases:")
pprint(pp_l_with_tokens)
logger.info(f"Output has keys {pp_output.keys()}")
assert pp_output.sequences_scores.shape == torch.Size([batch_size * n_seq])
assert len(pp_output.scores) == generated_length # different to greedy search: not generated_length - 1
assert pp_output.scores[0].shape == torch.Size([batch_size * n_seq, pp_tokenizer.vocab_size])
# scores_stacked = torch.stack(pp_output.scores, 1)
# assert scores_stacked.shape == torch.Size([batch_size * n_seq, generated_length, pp_tokenizer.vocab_size])
transition_logprobs = pp_model.compute_transition_beam_scores(
pp_output.sequences, pp_output.scores, pp_output.beam_indices, eos_token_id = pp_tokenizer.eos_token_id)
pp_logprobs = transition_logprobs.sum(-1)
assert pp_logprobs.shape == torch.Size([batch_size * n_seq])
print("logprobs (has grad):", pp_logprobs)
baseline_seq_prob = np.log(1/pp_tokenizer.vocab_size)* generated_length
baseline_short_seq_prob = np.log(1/pp_tokenizer.vocab_size)* np.floor(generated_length /2 )
baseline_high_prob_seq = np.log(1000/pp_tokenizer.vocab_size)* generated_length
baseline_high_prob_short_seq = np.log(1000/pp_tokenizer.vocab_size)* np.floor(generated_length /2 )
print("sequences scores (no grad):", pp_output.sequences_scores)
print("baseline prob (selecting token with prob 1/vocab_size every input):",baseline_seq_prob )
print("baseline short seq prob:",baseline_short_seq_prob )
print("baseline high prob seq", baseline_high_prob_seq)
print("baseline high prob short seq", baseline_high_prob_short_seq)
# -
def compare_scores_and_transition_probs():
# This code indicates that the transition probabilities are not the same as the scores.
# It seems they are the same sometimes but other times they are not.
for ex in range(batch_size * n_seq):
for step in range(generated_length):
tkn_id = pp_output.sequences[ex][step].item()
score = pp_output.scores[step][ex][tkn_id]
prob = transition_probs[ex][step]
print("example", ex, "step", step, "tkn_id", tkn_id,
"score", round_t(score), "transition_logprob", round_t(prob))
#compare_scores_and_transition_probs()
# ## Tokenizer differences
# ### Types
# Both the "eugenesiow/bart-paraphrase" model and the "tdopierre/ProtAugment-ParaphraseGenerator" are BART tokenizers and have type BartTokenizerFast. The implementation is identical to RobertaTokenizerFast according to the docs, which in turn was derived from GPT-2. They use byte-level Byte Pair Encoding.
#
# The "tuner007/pegasus_paraphrase" model is a Pegasus tokenizer has type PegasusTokenizerFast. This uses Unigram.
# ### Tokenization differences
# #### Spaces
# The BART tokenizer has been trained to treat spaces like parts of the tokens (a bit like sentencepiece) so a word will be encoded differently whether it is at the beginning of the sentence (without space) or not:
tokens = pp_tokenizer_bart(['hello there',' hello there'], return_tensors='pt')
get_tokens_from_token_ids_batch(pp_tokenizer_bart, tokens['input_ids'])
# The Pegasus tokenizer doesn't do this
tokens = pp_tokenizer_pegasus(['hello there',' hello there'], return_tensors='pt')
get_tokens_from_token_ids_batch(pp_tokenizer_pegasus, tokens['input_ids'])
# ### Representing tokens
# The tokenizers represent tokens differently.
# The BART models use Ġ to indicate start of a word for a token. Its generated tokens look like `['<s>', 'Hello', 'Ġmy', 'Ġname', 'Ġis', 'Ġz', 'f', 'ld', 'lf', 'o', 'q', 'd', '</s>', '<pad>', '<pad>', '<pad>']`
# The Pegasus model uses \_ to indicate start of a word for a token. Its generated tokens look like `['▁Hello', '▁my', '▁name', '▁is', '▁z', 'fl', 'dl', 'fo', 'q', 'd', '</s>', '<pad>']`
# ### Special tokens
# #### BART
logger.info(f"Tokenizer has these special tokens:{pp_tokenizer_bart.all_special_tokens}")
logger.info(f"The bos token is {pp_tokenizer_bart.bos_token} and has id {pp_tokenizer_bart.bos_token_id}")
logger.info(f"The eos token is {pp_tokenizer_bart.eos_token} and has id {pp_tokenizer_bart.eos_token_id}")
logger.info(f"The pad token is {pp_tokenizer_bart.pad_token} and has id {pp_tokenizer_bart.pad_token_id}")
logger.info(f"The unk token is {pp_tokenizer_bart.unk_token} and has id {pp_tokenizer_bart.unk_token_id}")
# #### PEGASUS
logger.info(f"Tokenizer has these special tokens:{pp_tokenizer_pegasus.all_special_tokens}")
logger.info(f"The bos token is {pp_tokenizer_pegasus.bos_token} and has id {pp_tokenizer_pegasus.bos_token_id}")
logger.info(f"The eos token is {pp_tokenizer_pegasus.eos_token} and has id {pp_tokenizer_pegasus.eos_token_id}")
logger.info(f"The pad token is {pp_tokenizer_pegasus.pad_token} and has id {pp_tokenizer_pegasus.pad_token_id}")
logger.info(f"The unk token is {pp_tokenizer_pegasus.unk_token} and has id {pp_tokenizer_pegasus.unk_token_id}")
# ### Special token usage with input and output sequences
# #### BART
# They use this format
# ```
# single sequence: <s> X </s>
# pair of sequences: <s> A </s></s> B </s>
# ```
# #### PEGASUS
# Format:
# ```
# - single sequence: ``X </s>``
# - pair of sequences: ``A B </s>`` (not intended use)
# ```
# BOS token is never used
# #### Differences
# Tokenizers also use different tokens when representing input sequences and generating output sentences. Here is a quick summary:
tokens_d = {
"bart": {
"special_tokens": pp_tokenizer_bart.all_special_tokens,
"input_start": pp_tokenizer_bart.bos_token,
"input_end": [pp_tokenizer_bart.pad_token, pp_tokenizer_bart.eos_token],
"output_start": pp_tokenizer_bart.eos_token,
"output_end": [pp_tokenizer_bart.pad_token, pp_tokenizer_bart.eos_token]
},
"pegasus": {
"special_tokens": pp_tokenizer_pegasus.all_special_tokens,
"input_start": None,
"input_end": [pp_tokenizer_pegasus.pad_token, pp_tokenizer_pegasus.eos_token],
"output_start": pp_tokenizer_pegasus.pad_token,
"output_end": [pp_tokenizer_pegasus.pad_token, pp_tokenizer_pegasus.eos_token],
}
}
tokens_d
# ### Token indexing
def print_tokens_from_ids(tokenizer, start_id=100, end_id=200):
ids = list(range(start_id,end_id))
print(*list(zip(ids, tokenizer.convert_ids_to_tokens(ids))))
# #### BART
# Having a look at generated tokens makes me suspect that they are indexed in whatever order they are encountered in the source text they are trained on. It seems like a rough frequency of english tokens but there are also tokens that are definitely out of order.
#
# The first few are reserved for special tokens, and the other low numbers (e.g. up to 100) are pretty common suffixes and words
print_tokens_from_ids(pp_tokenizer_bart, 0,50)
# Looking at 100 to 200 you can see some words (e.g. Trump at 140, or 2017 at 193) that aren't common enough to be that high. This makes me suspect that words are in encounter order in the text.
print_tokens_from_ids(pp_tokenizer_bart, 100,200)
# Tokens towards the end are gibberish or mispellings encountered in the input. The fifth last token is something labelled <|endoftext|> and I don't know what that is. Then there is a bunch of tokens like "madeupword0001". The last token is the mask token and then token indicies after that return None.
print_tokens_from_ids(pp_tokenizer_bart, pp_tokenizer_bart.vocab_size-20, pp_tokenizer_bart.vocab_size+10)
# #### PEGASUS
# Special tokens make up the first hundred or so. After that there's a token \<n> that seems like some new line thing.
print_tokens_from_ids(pp_tokenizer_pegasus, 0,120)
# Unlike the BART models I can believe that these tokens are in order of frequency. I can't see anything that is obviously out of place.
print_tokens_from_ids(pp_tokenizer_pegasus, 120,250)
# There's nothing special at the end, just looks like isolated tokens and None values after the tokens finish. It's also worth noting that the Pegasus model has ~96100 tokens which is way more than the ~50270 of the BART models (almost double).
print_tokens_from_ids(pp_tokenizer_pegasus, pp_tokenizer_pegasus.vocab_size-20, pp_tokenizer_pegasus.vocab_size+10)
# ## Questions
# ### When does the model generate padding tokens?
# For both models padding tokens are generated after the EOS token. Additionally for Pegasus generated text starts with the padding token.
# ### Why do generated paraphrases start with the EOS token?
# This is only the case for BART models. For pegasus models they use a padding token to start generated paraphrases.
#
# I don't know what the BOS token isn't used for these things. Pegasus has an open issue [here](https://github.com/huggingface/transformers/issues/12474).
#
# Whatever the reason you should just do the default because that is what the preprocessing does and you will get the best results that way.
# ### Does p(PAD) =1 after an eos token?
# For both BART and Pegasus models it appears that probability of outputting a pad token is actually zero at all timesteps. Instead the model outputs the eos token over and over, and there must be some post-processing that takes place that replaces eos token with padding token.
# For Pegasus it appears it is the same behaviour.
# Example code:
print(round_t(scores_softmax[:,:,pp_tokenizer.eos_token_id]))
print(round_t(scores_softmax[:,:,pp_tokenizer.pad_token_id]))
# What is interesting is that there is probability assigned to other tokens other than eos and pad after a eos token is outputted. Again there must be some kind of postprocessing that takes care of this situation because I haven't really seen it in the wild.
# Some models (e.g. GPT2) don't even have a PAD token. Instead they use the eos token on repeat. See this [issue](https://github.com/huggingface/transformers/issues/8452#issuecomment-739008168). What is confusing is seeing this behaviour with models that have a padding token.
# ### Do rows always sum to 1 when looking at token generation scores?
#
# Yes, they should. I put in an assert to check this.
# If you have a nan or an inf then they won't sum to 1. To confirm this
print(torch.isnan(torch.sum(torch.tensor([1,2,3, torch.nan]))))
print(torch.isinf(torch.sum(torch.tensor([1,2,3, torch.inf]))))
# ### Does first row sum to 0? (the one corresponding to the startoff token)
#
# So there is no token scores that correspond to the first token (usually a bos or pad token). The scores are a tuple of length (`generated_length - 1`). So there shouldn't be a "zero" row really.
# I remember seeing something like this at some point so I'll keep an eye out for it.
# ### How are logits containing nan or inf transformed with softmax and log_softmax?
# We can explore this through some code examples.
# #### Vanilla case
# First we look at the case without any nan or inf.
logits = torch.tensor([1.4, -1, 3, 2])
print(logits)
print(torch.softmax(logits,0))
print(torch.log_softmax(logits,0))
# The softmax values are interpreted as probabilities, and the log softmax is just the log of the probabilities, done for numerical stability. We can just take exponents to return to probabilities if needed.
print(torch.log_softmax(logits,0).exp())
# #### Positive inf
# Now let's see what happens if we introduce a positive inf.
logits = torch.tensor([1.4, -1, 3, 2, torch.inf])
print(logits)
print(torch.softmax(logits,0))
print(torch.log_softmax(logits,0))
# We get nan values in the softmax and log_softmax. So if you see nans in the softmax, remember that an inf in the scores is one reason why it may happen.
#
# This is interesting because if we just assume inf is a large positive number, we'd expect a softmax with basically a 1 and all zeros, and a log softmax of a 0 and a lot of negatives. We can try it here:
logits = torch.tensor([1.4, -1, 3, 2, 10000000000])
print(logits)
print(torch.softmax(logits,0))
print(torch.log_softmax(logits,0))
# Basically what we get. So this indicates that if we get a positive inf we might be able to mitigate this problem by clipping it to some kind of maximum value.
# #### Negative inf
logits = torch.tensor([1.4, -1, 3, 2, -torch.inf])
print(logits)
print(torch.softmax(logits,0))
print(torch.log_softmax(logits,0))
# Negative inf behaves a bit differently. The softmax is unaffected and basically just assigns a prob of 0 to the corresponding entry. The log softmax carries the `-inf` through.
#
# Again clipping the -inf to a large negative value can mitigate this problem somewhat:
logits = torch.tensor([1.4, -1, 3, 2, -10000000])
print(logits)
print(torch.softmax(logits,0))
print(torch.log_softmax(logits,0))
# #### nan values
logits = torch.tensor([1.4, -1, 3, 2, torch.nan])
print(logits)
print(torch.softmax(logits,0))
print(torch.log_softmax(logits,0))
# A nan in the logits propagates and affects the entire softmax and log_softmax tensors. The network basically gives up and says "no idea how to deal with this.
#
# This seems to be the case with most torch functions; e.g.
print(torch.sum(logits,0))
print(torch.divide(logits,0.2))
print(logits + logits)
# ### How do you interpret token entropy?
# The token scores (when stacked) are a tensor of dimensions (batch_size, generated_length - 1, vocab_size). We take softmax to get a tensor of probability distributions across all possible tokens. We can also calculate the entropy of each of these probability distributions.
#
# Entropy is a measure of how "peaky" or "flat" a probability distribution is. It is the expected value of the self-information of an event, which is basically a measure of how "surprised" you would be if that event occured.
# If we have a discrete random variable $X$ with probability distribution $P(x) $, entropy is given by $$H(X) = \mathbb{E}_{ X\sim P} [I(x)] = -\mathbb{E}_{X \sim P} [\log P(x)] $$ which is practically calculated by $$H(X) = -\sum_{x=-\infty}^\infty p(x) \log(p(x))$$
#
# The lowest value of entropy is 0, which is when you have $P(x)=1$ for some event.
Categorical(probs = torch.tensor([1,0,0,0])).entropy()
# High values of entropy occur when the probability distribution is very flat.
print(Categorical(probs = torch.tensor([0.20,0.50,0.10,0.20])).entropy()) # spikier, lower entropy
print(Categorical(probs = torch.tensor([0.25,0.25,0.25,0.25])).entropy()) # flatter, higher entropy
# In terms of tokens, we can show two realistic distributions below. The first has two likely tokens, one somewhat likely, and the rest unlikely. The second has many more likely tokens.
l1 = [0.001,0.001,0.001,0.001,0.5,0.001,0.4,0.001,0.001,0.001,0.001,0.001,0.09]
l2 = [0.1,0.1,0.1,0.1,0.025,0.1,0.1,0.1,0.1,0.1,0.025,0.025,0.025]
print(Categorical(probs = torch.tensor(l1)).entropy()) # spikier, lower entropy
print(Categorical(probs = torch.tensor(l2)).entropy()) # flatter, higher entropy
# There isn't a theoretical maximum for entropy, but for tokens you'll be governed by vocab size. Here we show some practical maximums for some different vocab sizes
v_size = 1000
print(Categorical(probs = torch.tensor([1/v_size for i in range(v_size)])).entropy()) # v small vocab
v_size = 10000
print(Categorical(probs = torch.tensor([1/v_size for i in range(v_size)])).entropy()) # small vocab
v_size = 50000
print(Categorical(probs = torch.tensor([1/v_size for i in range(v_size)])).entropy()) # ~BART vocab
v_size = 100000
print(Categorical(probs = torch.tensor([1/v_size for i in range(v_size)])).entropy()) # ~PEGASUS vocab
# These can be a bit hard to interpret so maybe you should also look at some other token-level stats, like max_prob, second_max_prob, third_max_prob, mean, variance or other things like that.
# Some other things to note. First you still get an entropy value if your probability dist sums up to more than 1, so make sure to check this before doing entropy. Secondly if you have nan or inf in the probability values then you will get an error. This is true if you use either `probs` or `logits` in the Categorical function.
print(Categorical(probs = torch.tensor([0.5,0.25,0.25,0.25])).entropy()) # sums to more than 1, gives result
#print(Categorical(probs = torch.tensor([0.5,0.25,0.25,torch.nan])).entropy()) # throws error
#print(Categorical(probs = torch.tensor([0.5,0.25,0.25,torch.inf])).entropy()) # throws error
#print(Categorical(logits = torch.tensor([0.5,0.25,0.25,torch.nan])).entropy()) # throws error
# ### What does it mean to take average of token entropy?
# Average entropy over tokens will depend on how many padding tokens are added. This in turn depends on how many examples there are in the batch.
# The entropy of padding tokens appears to be quite high based on some quick experiments.
# It might or might not depend on sentence length.
# It could be useful as a broad measure to see if the model is getting more "peaky" in selecting tokens at each time step.
# It might be useful when tracking a paraphrase over time and seeing its variations?
# ### What do you calculate KL divergence of?
# When your parameters to the policy network get updated, you calculate change in the output space and work out KL divergence of the prob dists.
#
# This might be hard for your case:
# * if you consider full sequences as the action space, these have such low probabilities that it's hard to see how they change really. you can't get a dist
# * you can get a dist of individual tokens but then you have to have the same paraphrase for that to make any sense whatsoever.
# * you could extract input_ids for a sequence and feed it into pp_model.generate_beam_transition_probabilities() to get the prob of that specific sentence. then you could maybe use this to keep token probs the same?
#
#
# ### How do you get nan and inf introduced into token scores?
#
# My understanding is that you get -inf for the first (min_length - 1) steps when you introduce a min_length parameter for the generated sequences for the eos_token_id slot. This is to stop the token from appearing and truncating the sequence.
#
# I also seem to get -inf for padding, eos tokens and sometimes just random other tokens as well. There doesn't seem to be much rhyme or reason to it.
#
# There is an option to generate to automatically handle these tokens. I haven't been using it though.
#
# You might also get -inf when setting other parameters to the `generate()` function (e.g. bad_words_ids or something similar).
#
# Nan can come in if you multiple -inf by 0 - this is equal to nan under ieee 754 standard.
# ### How big is the action space?
# Each different paraphrase can be considered a different action. This makes the action space a discrete action space rather than continuous. An initial estimate of its size is on the order of `vocab_size ^ generated_length`, but the vast majority of these sequences aren't valid English sentences and have a very low probability of being obtained. In addition the actions available are heavily dependent on the state (i.e. the original paraphrase). It is still a very large space.
# ### Can you log the top X most probable sentences and the probability of obtaining them?
#
# Currently you get your action using greedy search which picks the most likely token at each point. But this might not give the most likely sequence: you could have a high probability token in two timesteps that requires choosing the second-most-probable token to get there. So you can’t say that the greedy-search paraphrase is the most probable sequence.
#
# Beam search will always find outputs with greater than or equal probability to greedy search. However it is not guaranteed to find the most likely output. Yet maybe it can be an estimator of it.
#
#
# The most probable returned sentences will depend heavily on the hyperparameters given to the generate function, such as temperature, diversity_penalty, num_beam_groups, min_length, length_penalty and so on. You can only ever get the most likely tokens for a set of hyperparameters.
#
# So what you can do is generate a bunch of output with beam search and then calculate the sequence probabilities of it (or set length_penalty to 0 and use the sequence_scores). Then this is roughly what you want. But there is no guarantee that the greedy search output will be part of the generated sentences. And if it is then it probably won't be the most likely sentence output either.
#
# This might make more sense if we start using beam search instead of greedy search.
#
# If you can log this you could make a plot tracking how many pp has probs over: 1e-5, 1e-4, 1e-3, 1e-2, 1e-1. would be a good plot. x axis epoch, then ether do (a) for individual examples, or (b) as averages across examples
# ### Why can't we use sampling?
# From what I've read the sampling operation is not differentiable. This means that autograd doesn't carry a gradient through the sampling operation.
# I've read that "RL gets around this" but don't really understand the details at this point. Maybe with a non-differentiable policy gradient method?
#
# Links: [explanations](https://www.google.com/search?q=sampling+non+differentiable&oq=sampling+non+differentiable&aqs=chrome..69i57j69i60.7731j0j4&sourceid=chrome&ie=UTF-8) [hugginface forum post](https://discuss.huggingface.co/t/finetuning-gpt2-with-user-defined-loss/163?page=3) [alt method](https://leolaugier.wp.imt.fr/2019/09/09/workarounds-non-differentiability/)
# ### How are beam search scores and sequence_scores calculated?
# UPDATE: raised a github issue, let's see.
#
# There was a [PR](https://github.com/huggingface/transformers/pull/14654) that was merged in transformers v4.16.0 that seems to fix up the issue of the scores not being correct. Now there is a function [`compute_transition_beam_scores`](https://huggingface.co/docs/transformers/v4.16.2/en/main_classes/model#transformers.generation_utils.GenerationMixin.compute_transition_beam_scores) that seems to give you what you want.
#
# **Previously:**
#
# According to [this post](https://discuss.huggingface.co/t/generation-probabilities-how-to-compute-probabilities-of-output-scores-for-gpt2/3175/15?u=tomroth1001) they are calculated like this:
#
# **`sequence_scores`**: cumulative log probabilities of the `num_beams` most probable beams. It can be formulated as a recursive formula: sequence_scores[k]_i = sequence_score[k]_{i-1} + log_probs[i-1, :])_topk(2)[k] with sequence_score[k]_{i=start_token} = 0` (i being the time step, k being kth beam).
# * Then you divide this by length_penalty i think (or 1 + length penalty?), so a lot of people seem to just set length_penalty to zero
#
# **`scores`**: this is where it becomes confusing. At the moment the scores[i, j] are defined as log_probs[i-1, j] + sequence_score[j % vocab_size]_{i-1} whereas j % vocab_size essentially defines the beam index.
# * don't really know how to interpret this overly much.
#
#
# NOTE: scores for greedy search are not logprobs but rather logits.
#
# ### How does layer-norm affect the probs?
# Not sure. leaving this for now and lumping in with the dropout question.
# ### Given the size of the action space is this a good candidate for differential entropy?
# I don't think so - looks like it's continuous spaces only based on a cursory look
#
# You might need something more specialised - m[this paper](https://arxiv.org/pdf/1806.00589.pdf) talks about entropy for high dimensional spaces
# ### When do you hit floating point threshold for token probabilities? When do nans and inf get introduced?
# You work with log-probabilities exactly so you don't hit this issue.
# ### Does using fp32 affect token calculations?
#
# It shouldn't because you are working with log probabilities.
# ### How does dropout affect generated probabilities? How does train/eval mode affect generated probs for a sentence?
# Based on observations when we put a network into eval mode there is less randomness in the generated probabilities. This makes sense because we have reduced randomness because we remove the stochastic behaviour of the dropout node.
#
# We can have a look at the differences for some examples.
# +
def get_token_probs_for_mode(orig_l, mode):
if mode == "train": pp_model.train()
elif mode == "eval": pp_model.eval()
else: raise Exception("shouldn't get here")
orig_tokens = pp_tokenizer(orig_l, return_tensors='pt', padding=True, pad_to_multiple_of=4)
pp_output = pp_model.generate_with_grad(**orig_tokens, **pp_model_params, do_sample=False,
return_dict_in_generate=True,
output_scores=True,
remove_invalid_values=False)
pp_l_with_tokens = pp_tokenizer.batch_decode(pp_output.sequences, skip_special_tokens=False)
seq_without_first_tkn = pp_output.sequences[:, 1:]
attention_mask = pp_model._prepare_attention_mask_for_generation(
seq_without_first_tkn, pp_tokenizer.pad_token_id, pp_tokenizer.eos_token_id
)
scores_log_softmax = torch.stack(pp_output.scores, 1).log_softmax(2)
seq_token_log_probs = torch.gather(scores_log_softmax,2,seq_without_first_tkn[:,:,None]).squeeze(-1)
del scores_log_softmax
# account for nan values by setting them to 0 (maybe a bit of a hack)
# will also handle inf and -inf values too by default
seq_token_log_probs = torch.nan_to_num(seq_token_log_probs)
# account for the padding tokens at the end
seq_token_log_probs = seq_token_log_probs * attention_mask
seq_log_prob = seq_token_log_probs.sum(-1)
return (pp_l_with_tokens, seq_token_log_probs, seq_log_prob)
orig_l = [
"Hello I am tom",
"yes hello."
]
get_token_probs_for_mode(orig_l, mode ='train')
# -
get_token_probs_for_mode(orig_l, mode ='eval')
# It's hard to make too many inferences from this really.
# ## Glossary
# ### `attention_mask`
#
# The `attention_mask` is used when you have a batch of texts that are different lengths. The shorter one might be padded up to some minimum length with 0’s. The `attention_mask` is a binary vector that tell the model where the padding is. In effect, they can tell the model what to ignore. The 0’s will be padding.
#
# It is used to filter out padding tokens from the sequence probablity because then the sequence
# probability will depend on how many padding tokens there are and the probability of generating them,
# which (a) we don't want and (b) the probability isn't correct anyway
#
# It is returned when you tokenise the input.
#
# I also create it at one point with the code for the paraphrase
# ```
# attention_mask = pp_model._prepare_attention_mask_for_generation(
# seq_without_first_tkn, pp_tokenizer.pad_token_id, pp_tokenizer.eos_token_id
# )
# ```
# so that I can get the right probabilities for the sequence.
# ### `token_type_ids`
#
# **`token_type_ids`:** Some models/tasks use pairs of sentences concatenated together with a [SEP] token (or something) separating them. Tasks that might need this include textual entailment or duplicate detection. The `token_type_ids` is just a vector of 1’s and 0’s that indicate if a given word is part of the first sentence (a 0) or the second sentence (a 1).
#
# Looks something like this:
# ```
# >>> encoded_dict["token_type_ids"]
# [0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 1, 1, 1, 1, 1, 1]
# ```
#
# So you won't need this for sentiment classification but you might for entailment.
# ### `encoder_input_ids`
# Will leave this until I see it
# ### `decoder_input_ids`
# **`decoder_input_ids`:** Only applies to encoder-decoder models. [One source](https://huggingface.co/transformers/glossary.html#decoder-input-ids) puts this as the input id’s that will be fed into the decoder. [Another source](https://huggingface.co/transformers/model_doc/t5.html#training) puts this as the target sequence (shifted by one place) when doing seq2seq training . Often when training you pass in the `labels` attribute and the model figures out what this should be. See [here](https://huggingface.co/transformers/model_doc/t5.html#training)
# ### input_ids
# This is just the tokenized input sequence.
# ### decoder_start_token_id
# Will leave this until I see it.
| archive/exploring_generation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:flyvec] *
# language: python
# name: conda-env-flyvec-py
# ---
#default_exp tokenizer
# %reload_ext autoreload
# %autoreload 2
# # Tokenizer
# > A simple tokenizer for concepts using Gensim
#
# Tokenize words using Gensim. We wanted to avoid sub-word tokenization so that we can understand how the model lumps concepts together
# +
#export
from pathlib import Path
from gensim.corpora import Dictionary
from gensim.utils import simple_preprocess
from gensim.models.phrases import Phrases, Phraser
import os
import regex as re
import string
from cached_property import cached_property
import numpy as np
from typing import *
from fastcore.test import *
from flyvec.downloader import prepare_flyvec_data, get_config_dir, get_model_dir
# -
prepare_flyvec_data()
# ## Preprocessing functions
#
# We want tokens to deal with simple concepts, so we will enforce lowercase ASCII and predominantly split on spaces.
# Our tokenization will work with "lines" -- that is, a sequence of text that can contain multiple sentences, paragraphs, and newlines. For cohesiveness, we want to split these to the sentence and word level.
line = """
Various prior work has demonstrated 100 weaknesses in these models — even highly accurate ones — including reliance on non-salient regions
or on background information only. Explanation methods help identify these pitfalls by providing explanations for model predictions, enabling humans to identify the features on which a model decision is based. However, these methods provide explanations on the image level making it challenging to understand global model behavior or dataset limitations."""
# We first need to check that the line contains actual content and is not a binary string acting as an identifier in most files.
def is_good_line(line):
"""Check if the line is valid"""
return (len(line) > 1) and ("\x00" not in line)
is_good_line(line)
assert is_good_line(line)
assert not is_good_line("\x00\x0033-thegreatdivide.txt\x00")
assert not is_good_line("")
# Split a text by sentence according to the following regex pattern
# +
#export
spattern = re.compile(r"(?<!\w\.\w.)(?<![A-Z][a-z]\.)(?<=\.|\?)\s")
def line2sentences(line):
"""Convert a line into sentences, """
line = line.replace('\n', ' ').strip().lower()
return spattern.split(line)
# -
sentences = line2sentences(line); sentences
# Once we have a sentence, we want to strip all punctuation and unicode
#export
def isascii(s: str):
"""Determine if `s` is an entirely ascii string. Used for back-compatibility with python<3.7"""
try:
s.encode('ascii')
except UnicodeEncodeError:
return False
return True
assert isascii("Hello!")
assert not isascii("Ĉ")
assert isascii("")
#export
def strip_punc_unicode(line):
"""Strip all punctuation and unicode from the line"""
line = line.translate(str.maketrans('', '', string.punctuation))
line = ''.join([c for c in line if isascii(c)])
return line
proc_sentences = [strip_punc_unicode(s) for s in sentences]; proc_sentences
# And remove all instances where there are multiple spaces
# +
#export
space_pat = re.compile("\s+")
def remove_multiple_spaces(sentence):
return space_pat.sub(" ", sentence)
# -
proc_sentences = [remove_multiple_spaces(s) for s in proc_sentences]; proc_sentences
# Before we have our tokens, we will define the concept of 'number' as any ASCII token that contains a digit
#export
def isnum(token):
return any(t.isdigit() for t in token)
# Compiling all these steps into a single function
#export
def process_line(line):
"""Compose all transformations to process a line into tokens as desired"""
sents = line2sentences(line)
out = []
for s in sents:
x = strip_punc_unicode(s)
x = remove_multiple_spaces(x)
xs = x.split()
xs = [x_ if not isnum(x_) else "<NUM>" for x_ in xs]
out.append(xs)
return out
tokens = process_line(line); print(tokens[0])
def process_tok(x, num_tok="xxNUMxx", stop_tok="xxSTOPxx", stopwords=[]):
"""Process a token by replacing numbers and stop tokens with the desired special tokens"""
if isnum(x):
return num_tok
elif x in stopwords:
return stop_tok
return x.strip()
test_eq(process_tok(" "), "")
test_eq(process_tok("abc88"), "xxNUMxx")
test_eq(process_tok("993"), "xxNUMxx")
test_eq(process_tok("the", stopwords=["the", "a", "but"]), "xxSTOPxx")
test_eq(process_tok(" lotsofspace "), "lotsofspace")
[process_tok(t, stopwords=["the", "in", "on", "or", "has"]) for t in tokens[0]]
# And now we can convert an entire file to tokens (naively loading everything into memory)
#export
def file2tokens(fname):
"""Convert a file of text into tokenized sentences"""
with open(fname, 'r', encoding='utf8') as fp:
chunk = fp.readlines()
tokenized = []
for line in chunk:
if is_good_line(line):
tokenized += process_line(line)
return tokenized
# # The Tokenizer
# > Collecting all the helper functions underneath a single class
#hide
#export
PATCH_DICT = {
"<UNK>": 0,
"<NUM>": 1,
}
#export
class GensimTokenizer:
def __init__(self, dictionary, phraser=None, patch_dict=PATCH_DICT):
"""Wrap a Gensim Dictionary, phrase detector, and special tokens for creating tokenization from OWT
Args:
dictionary: The gensim dictionary mapping vocabulary to IDs and back
phraser: If provided, use gensim's phrase detector to lump common concepts together
patch_dict: Patch the dictionary with special tokens
"""
self.dictionary = dictionary
self.phraser = Phrases([[]]) if phraser is None else phraser
self.patch_dict = patch_dict
@classmethod
def from_file(cls, dict_fname, phraser_fname=None):
"""Load tokenizer information from a dictionary file (generated by gensim dictionary.save) and a phraser file."""
d = Dictionary.load(str(dict_fname))
if phraser_fname is not None:
p = Phraser.load(phraser_fname)
else:
p = Phraser(Phrases([[]]))
return cls(d, p)
def add_document_from_fname(self, fname):
"""For training, add the contents of a text file to the dictionary"""
print(f"Adding {fname}")
tokens = self.phraser[file2tokens(fname)]
self.dictionary.add_documents(tokens)
def add_to_phraser_from_fname(self, fname):
"""Detect common phrases from fname for bigramming purposes"""
print(f"Adding {fname} to phraser")
tokens = file2tokens(fname)
self.phraser.add_vocab(tokens)
def get_dictionary(self):
return self.dictionary
def token2id(self, word):
"""Convert a token into an id, converting to UNK ID as necessary"""
d = self.dictionary
return d.token2id.get(word, d.token2id["<UNK>"])
def tokens2ids(self, tokens):
"""Convert a list of tokens into ids, converting to UNK as necessary"""
return [self.token2id(tok) for tok in tokens]
def tokenize(self, s:str):
"""Convert a sentence into its tokens"""
return self.phraser[process_line(s)[0]]
def tokenize_batch(self, lines:List[str]):
"""Convert a batch of lines into their tokens"""
return self.phraser[[process_line(line)[0] for line in lines]]
def encode(self, s):
"""Encode a single sentence into IDs"""
sent_tokens = self.tokenize(s)
return self.tokens2ids(sent_tokens)
def decode(self, ids):
"""Alias for `ids2tokens`"""
return self.ids2tokens(ids)
def id2token(self, id):
"""Convert an id to a token"""
d = self.dictionary
if id == -1: return "<STOPWRD>" # Account for post processing
return d[id] # Add error handling if bad id
def ids2tokens(self, ids):
"""Convert iterable of ids to tokens"""
return [self.id2token(id) for id in ids]
def set_outdir(self, outdir):
"""Useful when training in parallel. If set, will save contents to outdir"""
self.outdir = Path(outdir)
def patch(self, vocab_size, new_vocab, no_below=15, no_above=0.8):
"""Patch the tokenizer with a manually specified list of tokens, after training"""
print("Patching with special tokens...")
self.dictionary.patch_with_special_tokens(self.patch_dict)
print("Filtering vocabulary...")
self.dictionary.filter_extremes(no_below=no_below, no_above=no_above, keep_n=vocab_size)
print(f"Adding {len(new_vocab)} new words to dictionary...")
new_vocab = self.tokenize_batch(new_vocab)
self.dictionary.add_documents(new_vocab)
print(f"Done patching. New vocab size = {self.n_vocab()}")
return new_vocab
def save(self, outfile):
self.dictionary.save(outfile)
def n_vocab(self):
return len(self.vocab)
@cached_property
def vocab(self):
return list(self.dictionary.keys())
@cached_property
def token_vocab(self):
return list(self.dictionary.values())
def __len__(self):
return self.n_vocab()
def encode_sentences_from_fname(self, fname):
"""Tokenize all the sentences from a text file"""
outlist = []
ind_offsets = []
new_start = 0
with open(fname, 'r') as fp:
for line in fp.readlines():
if is_good_line(line):
sents = self.phraser[process_line(line)]
for sent in sents:
ids = self.tokens2ids(sent)
outlist += ids
new_start = new_start + len(ids)
ind_offsets.append(new_start)
return np.asarray(outlist, dtype=np.int32), np.asarray(ind_offsets, dtype=np.uint64)
def encode_and_save_for_mp(self, fname):
"""Save sentences from fname. Needed because a local function can't be used with the MP module"""
if self.outdir is None: raise ValueError("Please `set_outdir` first")
fname = Path(fname)
idarr_outfile = self.outdir / (fname.stem + '.npy')
ind_offsets_outfile = self.outdir / (fname.stem + '_offsets.npy')
idarr, ind_offsets = self.encode_sentences_from_fname(fname)
np.save(idarr_outfile, idarr)
np.save(ind_offsets_outfile, ind_offsets)
# The `GensimTokenizer` is a simple wrapper around gensim's `Dictionary` and `Phraser` classes that aligns them with our simple tokenization rules. You can use the model for converting between tokens and ids as follows:
vocab = get_model_dir() / "tokenizer/gensim1_patched.dict"
tok = GensimTokenizer.from_file(vocab)
tokens = ["apple", "pie", "is", "delicious"]
ids = tok.tokens2ids(tokens); ids
tok.ids2tokens(ids)
# There are several different views into the vocabulary of the model.
# The tokens in the vocabulary
tok.token_vocab[:5]
# The IDs that correspond to those tokens
tok.vocab[:5]
# The dictionary object itself
d = tok.dictionary;
# # Export -
#hide
from nbdev.export import notebook2script
notebook2script()
| nbs/01_Tokenizer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="8IxqSmnawjmq" outputId="34288558-5cd9-4e81-c05b-6315760b97cc"
from google.colab import drive
drive.mount('/content/drive')
# + id="xt9Zn5QhXA5k"
# change to the directory that contains the data
main_path = "/content/drive/MyDrive/UdacityND/project_caps/"
# + id="IfaufJ3owoIv"
jigsaw_path = main_path+"train.csv"
val_data_path = main_path+"validation_data.csv"
comments_to_score_path = main_path+"comments_to_score.csv"
ruddit_path = main_path+"ruddit_with_text.csv"
new_data_path = main_path+"train_data_version2.csv"
# + id="YgbnsOoGwrP7" colab={"base_uri": "https://localhost:8080/"} outputId="c3b8102d-8c06-43f9-d1bd-469d3c0b2d1f"
# Imports
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
import random
random.seed(123)
import sys
import re
import nltk
nltk.download('punkt')
nltk.download('stopwords')
nltk.download('wordnet')
from nltk.tokenize import word_tokenize
from nltk.tokenize import sent_tokenize
from nltk.stem import WordNetLemmatizer
from nltk.corpus import stopwords
import pickle
import joblib
import seaborn as sns
sns.set_theme(style="whitegrid")
# + id="yHvdI9r5wtmT"
#Read in the data, ref: https://www.kaggle.com/andrej0marinchenko/best-score-0-856-jigsaw-for-beginners
def read_jigsaw_toxic_data(train_df_path, fold_num, use_folds = False, approach = 1):
'''
Read in the toxic jigsaw data
Inputs:
train_df_path -> path of the file to read
fold_num -> indicates the number of folds if they are usen
use_folds -> whehter to use folds or not
Output:
balanced_df -> a balanced df that has 3 coloumns, comment_text is the text of the comment, toxic_vs_not whether the comment is toxic or not,
y the score of the toxicity calculated
'''
df = pd.read_csv(train_df_path)
toxic_cols = ['toxic', 'severe_toxic', 'obscene', 'threat','insult', 'identity_hate']
# add a coloumn to indicate whether the comment is toxic or not
# 0 and 1 approach
df['toxic_vs_not'] = np.where(df[toxic_cols].sum(axis=1)==0, 0, 1) #0 if not toxic, 1 if toxic
# another approach instead of 0 and 1
if(approach==1):
df['severe_toxic'] = df.severe_toxic * 2
df['y'] = (df[['toxic', 'severe_toxic', 'obscene', 'threat', 'insult', 'identity_hate']].sum(axis=1) ).astype(int)
df['y'] = df['y']/df['y'].max()
else:
df['obscene'] = df.obscene * 0.16
df['toxic'] = df.toxic * 0.32
df['threat'] = df.threat * 1.5
df['insult'] = df.insult * 0.64
df['severe_toxic'] = df.severe_toxic * 1.5
df['identity_hate'] = df.identity_hate * 1.5
df['y'] = (df[['toxic', 'severe_toxic', 'obscene', 'threat', 'insult', 'identity_hate']].sum(axis=1) ).astype(float)
#df['y'] = df['y']/df['y'].max()
fig, ax = plt.subplots()
ax = sns.countplot(x="toxic_vs_not", data=df)
count_toxic = df['toxic_vs_not'].value_counts()[1]
print(count_toxic)
if(use_folds==False):
balanced_df = pd.concat([df[df.y>0].sample(frac=1, random_state = 10*(fold_num+1)),df[df.y==0].sample(int(count_toxic), random_state = 10*(fold_num+1))], axis=0)
else:
balanced_df = pd.concat([df[df.y>0].sample(frac=0.8, random_state = 10*(fold_num+1)),df[df.y==0].sample(int(0.8*count_toxic), random_state = 10*(fold_num+1))], axis=0)
fig2, ax2 = plt.subplots()
ax2 = sns.histplot(data=balanced_df,x='y',bins = 10, binwidth=0.08)
return df
# + colab={"base_uri": "https://localhost:8080/", "height": 571} id="4nDX86kT-yS5" outputId="ae0d930b-c4f5-47c1-c5d4-7b8fe1600e47"
jigsaw_df = read_jigsaw_toxic_data(jigsaw_path,0 )
# + id="Y4v2XbDd-2NC"
# ref: https://www.kaggle.com/andrej0marinchenko/best-score-0-856-jigsaw-for-beginners
def read_ruddit_toxic_data(ruddit_path, fold_num = 0, use_folds = False):
'''
Read in ruddit toxic data
Inputs:
ruddit_path -> path of the file to read
fold_num -> indicates the number of folds if they are usen
use_folds -> whehter to use folds or not
Output:
df -> a df that has 2 coloumns, comment_text is the text of the comment,
y the score of the toxicity calculated
'''
df = pd.read_csv(ruddit_path)
#print('Shape of Ruddit data is '+str(df.shape))
df = df[['txt', 'offensiveness_score']].rename(columns={'txt': 'comment_text','offensiveness_score':'y'})
fig, ax = plt.subplots()
ax = sns.histplot(data=df,x='y',bins = 10, binwidth=0.08)
df['y'] = (df['y'] - df.y.min()) / (df.y.max() - df.y.min())
if(use_folds==True):
df = df.sample(frac=0.8, random_state = 10*(fold_num+1))
fig, ax = plt.subplots()
ax = sns.histplot(data=df,x='y',bins = 10, binwidth=0.08)
return df[['comment_text','y']]
# + colab={"base_uri": "https://localhost:8080/", "height": 553} id="EIwRJAzQZ-fr" outputId="d9fae10e-c009-46ee-8800-62f21ed35968"
ruddit_df = read_ruddit_toxic_data(ruddit_path)
# + id="SfBLvekIYYfc"
def read_new_toxic_data(new_path, fold_num = 0, use_folds = False):
'''
Read in alternative jigsaw data
Inputs:
new_path -> path of the file to read
fold_num -> indicates the number of folds if they are usen
use_folds -> whehter to use folds or not
Output:
df -> a balanced df that has 2 coloumns, comment_text is the text of the comment,
y the score of the toxicity calculated
'''
df = pd.read_csv(new_path)
df = df[['text','y']].rename(columns={'text': 'comment_text'})
if(use_folds==True):
df = df.sample(frac=0.8, random_state = 10*(fold_num+1))
fig, ax = plt.subplots()
ax = sns.histplot(data=df,x='y',bins = 10, binwidth=0.08)
return df[['comment_text','y']]
# + colab={"base_uri": "https://localhost:8080/", "height": 287} id="z3n-6yW7ZCJL" outputId="6cb5f189-a323-418d-c464-a73a388a1923"
new_toxic_df = read_new_toxic_data(new_data_path)
# + id="11hdlEQdZGeD"
def cleaner(text,use_lemmatizer=True):
'''
A function to clean the text
Inputs:
text -> The text that should be cleaned
Outputs:
clean_tokens -> A list of tokens extracted from text after case normalizing, removing stop words and lemmatizing
'''
url_regex = 'http[s]?://(?:[a-zA-Z]|[0-9]|[$-_@.&+]|[!*\(\),]|(?:%[0-9a-fA-F][0-9a-fA-F]))+'
detected_urls = re.findall(url_regex, text)
for url in detected_urls:
text = text.replace(url, "urlplaceholder")
words = word_tokenize(text)
words = [word for word in words if word.isalpha()] #remove anything that is not alpha numeric
lemmatizer = WordNetLemmatizer()
clean_tokens = []
for tok in words:
if tok not in stopwords.words("english"):
if(use_lemmatizer == True):
clean_tok = lemmatizer.lemmatize(tok).lower().strip()
else:
clean_tok = tok.lower().strip()
clean_tokens.append(clean_tok)
return ' '.join(clean_tokens)
# + id="I5oTeITFoEAd"
# load and save clean jigsaw data
cleaned_jigsaw_text = jigsaw_df['comment_text'].apply(cleaner)
jigsaw_df['comment_text'] = cleaned_jigsaw_text
jigsaw_df.to_csv('jigsaw.csv',index=False)
# + id="K3ksh8y6uSjq"
# load and save clean ruddit data
cleaned_ruddit_text = ruddit_df['comment_text'].apply(cleaner)
ruddit_df['comment_text'] = cleaned_ruddit_text
ruddit_df.to_csv('ruddit.csv',index=False)
# + id="XMcxnQbyvEJt"
# load and save clean new data
cleaned_new_toxic_text = new_toxic_df['comment_text'].apply(cleaner)
new_toxic_df['comment_text'] = cleaned_new_toxic_text
new_toxic_df.to_csv('new_toxic.csv',index=False)
# + id="AHwvfbpAUv-6"
# load, clean and save validation data
all_val_data = pd.read_csv(val_data_path)
less_toxic_data = all_val_data['less_toxic'].apply(cleaner)
more_toxic_data = all_val_data['more_toxic'].apply(cleaner)
all_val_data['less_toxic'] = less_toxic_data
all_val_data['more_toxic'] = more_toxic_data
all_val_data.to_csv('val_data.csv',index=False)
# + id="RNF6Z3w1XinZ"
# load, clean and save new comments that are to be scored
new_comments = pd.read_csv(comments_to_score_path)
temp = new_comments['text'].apply(cleaner)
new_comments['comments'] = temp
new_comments.to_csv('comments_to_score.csv',index=False)
| Data_exploration_and_cleaning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="bODtRcczONme"
# <h2><center> CSCI - UA 9472 - Artificial Intelligence </center></h2>
#
# <h3><center> Assignment 3: Logical reasoning </center></h3>
#
# <center>Given date: November 8
# </center>
# <center><font color='red'>Due date: November 30 </font>
# </center>
# <center><b>Total: 40 pts </b>
# </center>
#
# + [markdown] id="QzVwXVF5ONmo"
# <center>In this third assignment, we will implement a simple Logical agent by relying on the resolution algorithm of Propositional Logic.</center>
# + [markdown] id="CBexOqGKONmo"
# <img src="https://github.com/acosse/Artificial-Intelligence-Fall2021/blob/main/Assignments/Assignment3/simpleVideoGameCave.jpeg?raw=1" width="400" height="300"/>
#
# = = = = = = = = = = = = = = = = = = = = = \
# **Name:** Haoming(<NAME> \
# **NetID:** hl3797 \
# **Date:** Nov. 29th, 2021 \
# = = = = = = = = = = = = = = = = = = = = =
#
# + id="iqLffiCMteyV"
import time
import collections
import numpy as np
from copy import deepcopy
from random import choice
import matplotlib.pyplot as plt
from IPython.display import display, HTML
from matplotlib.animation import FuncAnimation
# + [markdown] id="YH_3BqD6ONmp"
# ### Introduction: logical propositions
#
# The final objective will be to code our logical agent to succeed in a simple world similar to the Wumpus world discussed in the lectures. The final world we will consider is shown below.
# + [markdown] id="_cdk6zR7ONmp"
# <img src="https://github.com/acosse/Artificial-Intelligence-Fall2021/blob/main/Assignments/Assignment3/MazeTotal.png?raw=1" width="400" height="300"/>
# + [markdown] id="H4k8NJpRONmq"
# Before designing the full agent, we will focus on a series of simplified environments (see below). In order to help you in your implementation, you are provided with the class 'Expr' and the associated function 'expr' which can be used to store and process logical propositions. The logical expressions are stored as objects consisting of an operator 'op' which can be of the types '&' (and), '|' (or) '==>' (implication) or '<=>' (double implication) as well as '~' (not). A logical expression such as 'A & B' can be stored as a string by means of the function expr() as expr('A & B') or Expr('&', 'A', 'B').
#
# The function expr() takes operator precedence into account so that the two lines
# + id="8b8vg9P3ONmt"
'''source : AIMA'''
def Symbol(name):
"""A Symbol is just an Expr with no args."""
return Expr(name)
class Expr:
"""source: Artificial Intelligence: A Modern Approach
A mathematical expression with an operator and 0 or more arguments.
op is a str like '+' or 'sin'; args are Expressions.
Expr('x') or Symbol('x') creates a symbol (a nullary Expr).
Expr('-', x) creates a unary; Expr('+', x, 1) creates a binary."""
def __init__(self, op, *args):
self.op = str(op)
self.args = args
# Operator overloads
def __neg__(self):
return Expr('-', self)
def __pos__(self):
return Expr('+', self)
def __invert__(self):
return Expr('~', self)
def __add__(self, rhs):
return Expr('+', self, rhs)
def __sub__(self, rhs):
return Expr('-', self, rhs)
def __mul__(self, rhs):
return Expr('*', self, rhs)
def __pow__(self, rhs):
return Expr('**', self, rhs)
def __mod__(self, rhs):
return Expr('%', self, rhs)
def __and__(self, rhs):
return Expr('&', self, rhs)
def __xor__(self, rhs):
return Expr('^', self, rhs)
def __rshift__(self, rhs):
return Expr('>>', self, rhs)
def __lshift__(self, rhs):
return Expr('<<', self, rhs)
def __truediv__(self, rhs):
return Expr('/', self, rhs)
def __floordiv__(self, rhs):
return Expr('//', self, rhs)
def __matmul__(self, rhs):
return Expr('@', self, rhs)
def __or__(self, rhs):
"""Allow both P | Q, and P |'==>'| Q."""
if isinstance(rhs, Expression):
return Expr('|', self, rhs)
else:
return PartialExpr(rhs, self)
# Reverse operator overloads
def __radd__(self, lhs):
return Expr('+', lhs, self)
def __rsub__(self, lhs):
return Expr('-', lhs, self)
def __rmul__(self, lhs):
return Expr('*', lhs, self)
def __rdiv__(self, lhs):
return Expr('/', lhs, self)
def __rpow__(self, lhs):
return Expr('**', lhs, self)
def __rmod__(self, lhs):
return Expr('%', lhs, self)
def __rand__(self, lhs):
return Expr('&', lhs, self)
def __rxor__(self, lhs):
return Expr('^', lhs, self)
def __ror__(self, lhs):
return Expr('|', lhs, self)
def __rrshift__(self, lhs):
return Expr('>>', lhs, self)
def __rlshift__(self, lhs):
return Expr('<<', lhs, self)
def __rtruediv__(self, lhs):
return Expr('/', lhs, self)
def __rfloordiv__(self, lhs):
return Expr('//', lhs, self)
def __rmatmul__(self, lhs):
return Expr('@', lhs, self)
def __call__(self, *args):
"""Call: if 'f' is a Symbol, then f(0) == Expr('f', 0)."""
if self.args:
raise ValueError('Can only do a call for a Symbol, not an Expr')
else:
return Expr(self.op, *args)
# Equality and repr
def __eq__(self, other):
"""x == y' evaluates to True or False; does not build an Expr."""
return isinstance(other, Expr) and self.op == other.op and self.args == other.args
def __lt__(self, other):
return isinstance(other, Expr) and str(self) < str(other)
def __hash__(self):
return hash(self.op) ^ hash(self.args)
def __repr__(self):
op = self.op
args = [str(arg) for arg in self.args]
if op.isidentifier(): # f(x) or f(x, y)
return '{}({})'.format(op, ', '.join(args)) if args else op
elif len(args) == 1: # -x or -(x + 1)
return op + args[0]
else: # (x - y)
opp = (' ' + op + ' ')
return '(' + opp.join(args) + ')'
def expr(x):
"""
Shortcut to create an Expression. x is a str in which:
- identifiers are automatically defined as Symbols.
- ==> is treated as an infix |'==>'|, as are <== and <=>.
If x is already an Expression, it is returned unchanged. Example:
>>> expr('P & Q ==> Q')
((P & Q) ==> Q)
"""
return eval(expr_handle_infix_ops(x), defaultkeydict(Symbol)) if isinstance(x, str) else x
def expr_handle_infix_ops(x):
"""
Given a str, return a new str with ==> replaced by |'==>'|, etc.
>>> expr_handle_infix_ops('P ==> Q')
"P |'==>'| Q"
"""
infix_ops = '==> <== <=>'.split()
for op in infix_ops:
x = x.replace(op, '|' + repr(op) + '|')
return x
class defaultkeydict(collections.defaultdict):
"""
Like defaultdict, but the default_factory is a function of the key.
>>> d = defaultkeydict(len); d['four']
4
"""
def __missing__(self, key):
self[key] = result = self.default_factory(key)
return result
class PartialExpr:
"""Given 'P |'==>'| Q, first form PartialExpr('==>', P), then combine with Q."""
def __init__(self, op, lhs):
self.op, self.lhs = op, lhs
def __or__(self, rhs):
return Expr(self.op, self.lhs, rhs)
def __repr__(self):
return "PartialExpr('{}', {})".format(self.op, self.lhs)
Number = (int, float, complex)
Expression = (Expr, Number)
# + [markdown] id="frjLqPkdjXlr"
# #### Utility functions
# + id="y1u-NgIcjWO0"
OP_BASE = {'&': True, '|': False}
# credits: https://github.com/aimacode/aima-python
def flatten(op, args):
'''
flatten clause & fetch components
>>> flatten('|', expr('A | (B | C)').args)
[A, B, C]
'''
flat_args = []
def fetch(subargs):
for arg in subargs:
if arg.op == op:
fetch(arg.args)
else:
flat_args.append(arg)
fetch(args)
return flat_args
# credits: https://github.com/aimacode/aima-python
def merge(op, args):
'''
flatten clause & connect args with op
>>> merge('&', expr('(A & B) & (B | C)').args)
(A & B & (B | C))
'''
# print(op, args)
args = flatten(op, args)
# print(args)
if len(args) == 0:
return OP_BASE[op]
elif len(args) == 1:
return args[0]
else:
return Expr(op, *args)
def adjacents(curr_h, curr_w, world_h=5, world_w=5):
'''get valid adjacent cells at given position'''
cells = []
if curr_h-1 >= 0:
cells.append((curr_h-1, curr_w))
if curr_h+1 < world_h:
cells.append((curr_h+1, curr_w))
if curr_w-1 >= 0:
cells.append((curr_h, curr_w-1))
if curr_w+1 < world_w:
cells.append((curr_h, curr_w+1))
return cells
def all_adjacents(pos_list, world_h=5, world_w=5):
'''get valid adjacent cells at multiple position'''
cells = []
for h, w in pos_list:
cells += adjacents(h, w, world_h, world_w)
return cells
def pairs(clauses):
'''iter all possible pairs of clauses'''
val = list(clauses.values())
for i in range(len(val)):
for j in range(i+1, len(val)):
yield (val[i], val[j])
def rules(s1, s2, world_h=5, world_w=5, imp='==>'):
'''init rules with given symbols, implications, and world size'''
ans = []
for h in range(world_h):
for w in range(world_w):
for adj in adjacents(h, w):
c = ' '.join([get_symbol(s1, (h, w)), imp, get_symbol(s2, adj)])
ans.append(expr(c))
# print(ans)
return ans
def display_movie(world, path, delay=1, world_size=(5, 5)):
'''Display a moview with given world & path'''
# init colors
def init_obj_color(key, color_id):
if key not in world:
return
if isinstance(world[key], list):
# print(key, world[key])
for pos in world[key]:
maze[pos] = color_id
else:
maze[world[key]] = color_id
# plot a new frame
def animate(frame_idx):
frame_idx = frame_idx // delay
plot.set_data(maze_frames[frame_idx])
return (plot, )
# init world
print('\ngenerating movie...\n')
maze = np.full(world_size, fill_value=8)
init_obj_color('Shine', 7)
init_obj_color('Oooo', 7)
init_obj_color('Six', 7)
init_obj_color('Ghost', 2)
init_obj_color('Spike', 2)
init_obj_color('Treasure', 1)
init_obj_color('Door', 1)
# generate frames
maze_frames = []
for i in range(len(path)):
maze[path[i]] = 5
maze_with_agent = maze.copy()
maze_with_agent[path[i]] = 3
maze_frames.append(maze_with_agent)
# init first frame
fig, ax = plt.subplots()
plot = ax.matshow(maze_frames[0], cmap=plt.cm.Set2)
# make animation
frame_num = len(maze_frames) * delay
anim = FuncAnimation(fig, animate, frames=frame_num, interval=30, blit=True)
display(HTML(anim.to_html5_video()))
fig.delaxes(ax)
# check raw atomic sentence: expr('A') => True
ras_flag = lambda p: (p.op.isalnum()) and (p.args == ())
# check negated atomic sentence: expr('~A') => True
nas_flag = lambda p: (p.op == '~') and (len(p.args) == 1)
# check atomic sentence: [raw a.s.] or [negated a.s.] => True
atm_flag = lambda p: (ras_flag(p)) or (nas_flag(p) and ras_flag(p.args[0]))
# check disjuction of atomic sentences: expr('(A | ~B)') => True
doa_flag = lambda p: (p.op == '|') and all([atm_flag(q) for q in p.args])
# check CNF components: [atomic] or [disjuction of atomic] => True
cmp_flag = lambda p: atm_flag(p) or doa_flag(p)
# check conj. of disj. of atomic sentences: expr('(A | ~B) & C') => True
cod_flag = lambda p: (p.op == '&') and all([cmp_flag(q) for q in p.args])
# check CNF: [CNF comp.] or [conj. of disj.] => True
cnf_flag = lambda p: cmp_flag(p) or cod_flag(p)
# get symbols of CNF components: expr('(A | ~B)') => [A, ~B]
cmp_symb = lambda p: [p] if atm_flag(p) else list(p.args)
# get CNF components: expr('(A | ~B) & C') => [(A | ~B), C]
cnf_comp = lambda p: [p] if cmp_flag(p) else list(p.args)
# get symbol with given prefix & position
get_symbol = lambda s, pos: '%s%d%d' % (s, pos[0], pos[1])
# get a list with a given item omitted
omit_items = lambda seq, item: [x for x in seq if x != item]
# get the symbol and flag of an atomic sentence (e.g. '~A' => 'A', False)
parse_atomic = lambda p: (str(p)[1:], False) if p.op == '~' else (str(p), True)
# + [markdown] id="2k0yDJ26ONmw"
# #### Question 1: to CNF (7pts)
#
# Now that we can create a knowledge base, in order to implement the resolution algorithm that will ultimately enable our agent to leverage the information from the environment, we need our sentences to written in conjunctive normal form (CNF). That requires a number of steps which are recalled below:
#
# - Biconditional elimination: $(\alpha \Leftrightarrow \beta) \equiv ((\alpha\Rightarrow \beta) \wedge (\beta \Rightarrow \alpha))$
# - Implication elimination $\alpha \Rightarrow \beta \equiv \lnot \alpha \vee \beta$
# - De Morgan's Law $\lnot (A\wedge B) = (\lnot A \vee \lnot B)$, $\lnot(\alpha \vee B) \equiv (\lnot A \wedge \lnot B)$
# - Distributivity of $\vee$ over $\wedge$: $(\alpha \vee (\beta \wedge \gamma)) \equiv ((\alpha \vee \beta) \wedge (\alpha \vee \gamma))$
#
# Relying on the function propositional_exp given above (to avoid having all the questions depend on question 1, we will now rely on this implementation from AIMA), complete the function below which should return the CNF of the logical propostion $p$.
# + id="zzQt9jzdONmx" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1638224000236, "user_tz": 300, "elapsed": 205, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhOzSFkObCl-UMX25oW5lPfwwbD5InBzHMI_DJfSw=s64", "userId": "10852634671235052019"}} outputId="3c5ade02-61b1-423d-d2a4-c82c1aa2f4e0"
def eliminate_imp(p):
'''
implication elimination (i.e. "<=>", "<==", "==>")
Note: single/double cases have been merged for efficiency
>>> eliminate_imp(expr('A <=> B'))
((~A | B) & (~B | A))
'''
# case 1: atomic sentence
if atm_flag(p):
return p
args = tuple(map(eliminate_imp, p.args))
e1, e2 = args[0], args[-1]
# case 2: implications
if p.op == '==>':
return (~e1 | e2)
elif p.op == '<==':
return (~e2 | e1)
elif p.op == '<=>':
return (~e1 | e2) & (~e2 | e1)
# case 3: otherwise
else:
return Expr(p.op, *args)
def move_neg(p):
'''
move '~' inside: De Morgan's Law & neg elimination
>>> move_neg(expr('~(A & ~B)'))
(~A | B)
'''
# case 1: atomic sentence
if atm_flag(p):
return p
# case 2: with '~' to move
elif p.op == '~':
e = p.args[0]
dml = lambda x: move_neg(~x)
if e.op == '~':
# ~~A => A
return move_neg(e.args[0])
else:
# ~(A & B) => (~A | ~B)
# ~(A | B) => (~A & ~B)
new_op = ('&' if e.op == '|' else '|')
return merge(new_op, tuple(map(dml, e.args)))
# case 3: otherwise
else:
return Expr(p.op, *tuple(map(move_neg, p.args)))
def distribute(p):
'''
distributivity of ∨ over ∧
>>> distribute(expr('(A & B) | C'))
((A | C) & (B | C))
'''
def find_first_op(args, op):
for arg in args:
if arg.op == op:
return arg
return None
# case 1: atomic sentence
if atm_flag(p):
return p
# case 2a: there might be '&' inside
if p.op == '|':
p = merge('|', p.args)
if len(p.args) == 1:
return distribute(p.args[0])
conj = find_first_op(p.args, '&')
if conj is None:
return merge(p.op, p.args)
others = [arg for arg in p.args if arg is not conj]
others = (others[0] if len(others) == 1 else merge('|', others))
clause = [(e|others) for e in conj.args]
return merge('&', tuple(map(distribute, clause)))
# case 2b: there might be '&' inside (easier than '|')
elif p.op == '&':
return merge('&', tuple(map(distribute, p.args)))
# otherwise
else:
return p
def beautify_cmp(p):
'''
sort symbol & remove all "A | ~A" in the components
>>> beautify_cmp(expr('A | ~A | B | C'))
(B | C)
'''
if atm_flag(p):
return p
remove_set = set()
symbol_set = set(cmp_symb(p))
# print(symbol_set)
for s in symbol_set:
if ~s in symbol_set:
remove_set = remove_set | {s, ~s}
return merge('|', tuple(sorted(list(symbol_set - remove_set))))
def beautify_cnf(p):
'''beautify all the components inside a CNF'''
if atm_flag(p):
return p
new = tuple(sorted(list(set([beautify_cmp(c) for c in cnf_comp(p)]))))
if (len(new) == 0) or (False in new):
return False
elif len(new) == 1:
return new[0]
else:
return merge('&', new)
def to_CNF(p):
'''return the CNF of proposition p'''
if not isinstance(p, Expr):
p = expr(p)
return beautify_cnf(distribute(move_neg(eliminate_imp(p))))
e = expr('A <=> B')
to_CNF(e)
# + [markdown] id="MCFl_EE6ONmx"
# #### Question 2: The resolution rule (7pts)
#
# Now that you have a function that can turn any logical sentence to a CNF, we will code the resolution rule. For any two propositions $p_1$ and $p_2$ written in conjunctive normal form, write a function Resolution that returns empty if the resolution rule applied to the two sentences cannot produce any new sentence and that returns the set of all propositions $p_i$ following from the resolution of $p_1$ and $p_2$ otherwise.
#
# Study the disjuncts of both $p_1$ and $p_2$. For each of the disjunct in $p_1$ try to find in $p_2$ the negation of that disjunct. If this negation appears, returns the proposition resulting from combining $p_1$ and $p_2$ with a disjunction.
# + id="UJejf19sONmy" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1638224000237, "user_tz": 300, "elapsed": 11, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhOzSFkObCl-UMX25oW5lPfwwbD5InBzHMI_DJfSw=s64", "userId": "10852634671235052019"}} outputId="d5cec187-f0d1-4720-9bfd-9c4d51293d89"
'''
Note: Originally, I implemented this function by the pseudocode in textbook,
which works quite slow. Inspired by Haiyang, I implement the resolution rule
based on symbols and set operations, which works amazingly fast.
This brilliant idea credits to Haiyang.
'''
def resolution_rule(p1, p2):
'''
applies the resolution rule on two propositions p1 and p2
return -1 for invalid cases (e.g. empty clause, insolvable, etc.)
otherwise return the resolution result
'''
def split(p_set):
pos_set = {str(s) for s in p_set if s.op !='~'}
neg_set = {str(s)[1:] for s in p_set if s.op =='~'}
return pos_set, neg_set
def combine(pos_set, neg_set):
pos_lst = [expr(s) for s in sorted(list(pos_set))]
neg_lst = [expr('~' + s) for s in sorted(list(neg_set))]
return pos_lst + neg_lst
# get all symbols & split to pos/neg
p1_set, p2_set = set(cmp_symb(p1)), set(cmp_symb(p2))
p1_pos, p1_neg = split(p1_set)
p2_pos, p2_neg = split(p2_set)
# print(p1_set, p2_set)
# print(p1_pos, p1_neg)
# print(p2_pos, p2_neg)
if len(p1_pos & p2_neg) == 0 and len(p1_neg & p2_pos) == 0:
return -1
# fast resolution on atomic sentences (credits Haiyang)
pos = (p1_pos - p2_neg) | (p2_pos - p1_neg)
neg = (p1_neg - p2_pos) | (p2_neg - p1_pos)
# print(pos, neg)
if len(pos & neg):
return -1
args = combine(pos, neg)
if len(args) == 0:
return Expr('()')
elif len(args) == 1:
return Expr(args[0])
else:
return Expr("|", *args)
# e1 = expr('A')
# e2 = expr('~A')
e1 = expr('A | B')
e2 = expr('~A | C')
resolution_rule(e1, e2)
# + [markdown] id="474KX_DCONmy"
# #### Question 3: The resolution algorithm (6pts)
#
# Now that we have a resolution function we can embed it into a resolution algorithm. Complete the function Resolution below which implements the resolution algorithm. The algorithm takes as input a knowledge base written in conjunctive normal form, and a proposition $\alpha$ and should return true or false (as stored in the variable 'is_entailed') depending on whether $\alpha$ is entailed by the knowledge base KB.
# + id="QeCgqXTiONmy"
# Note: The resolution algorithm has been integrated in the KnowledgeBase class
# + [markdown] id="Mhvnud9UONmz"
# #### Question 4 (8pts): A first logical agent
#
# Given our resolution algorithm, we will finally be able to design our first logical agent. As a first step, we consider a simple agent located on the bottom left cell of a 5 by 5 grid world. The world contains a single threat represented by a ghost which occupies a single cell and emits a loud noise ('OOooo') audible in the immediately adjacent cells.
#
# To implement the agent, we will use the following approach:
#
# Each cell will be represented by its (x,y) coordinate ((0,0) being the bottom leftmost cell). On top of this we will consider the following symbols
#
# - $D_{(x, y)}$ (which you can store as the string 'Dxy' or 'D_xy' as you want) indicating whether there is an exit on the cell or not (when the agent reach the exit, the simulation ends),
#
# - $G_{(x, y)}$ which encodes whether the ghost is located on cell $(x, y)$ or not
#
# - $O_{(x, y)}$ which encodes whether a "Ooooo" is heard on the cell $(x, y)$
#
# Using those three symbols, the state of the world can be defined with a total of 3*25 symbols.
#
# In a while loop running until the door is found, code a simple logical agent that moves at random in the non-threatening cells until it finds the escape cell. Also consider the following:
#
# - The agent should keep track of a knowledge base KB (list of logically true propositions together with a dictionnary storing the symbols that are known to be true) including all the sentences that the agent can generate based on the information it collected at the previous steps.
#
#
# - You might want to have a simple function, which given a symbol returns the adjacent symbols (symbols corresponding to spatially adjacent cells). you can store those as strings
#
#
# - The agent should be equipped with the resolution algorithm that you coded above. Before each new move, the next cell should be determined at random (from the set of all non visited cells) and the agent should use the resolution algorithm to determine whether the ghost is on the cell or not. If there is no indication that the ghost is located on the cell, the agent should make the move.
# + [markdown] id="8F8LSIXvONmz"
# <img src="https://github.com/acosse/Artificial-Intelligence-Fall2021/blob/main/Assignments/Assignment3/MazeGhostb.png?raw=1" width="400" height="300"/>
# + id="aREF3-7YONmz"
class KnowledgeBase(object):
def __init__(self, *clauses):
self.symbols = {} # store T/F of some specific symbols
self.clauses = {} # store all clauses in KB
if len(clauses) > 0:
self.tell(clauses[0] if len(clauses) == 1 else Expr("&", *clauses))
# add a sentence into KB
def tell(self, sentence, to_cnf=False):
# convert to CNF as needed
if not cnf_flag(sentence):
sentence = to_CNF(sentence)
# print('cnf', sentence)
# omit useless sentence
if sentence == False:
return
# add each CNF component
for c in cnf_comp(sentence):
self.tell_comp(c)
# add a CNF component into KB
def tell_comp(self, p):
# print('learn:', p)
# quick way to learn an atomic sentence
if atm_flag(p):
# parse p & backup the knowledge in self.symbols
s, f = parse_atomic(p)
if (s in self.symbols) and (self.symbols.get(s, False) != f):
raise ValueError('Contradiction: symbol [%s].' % s)
self.symbols[s] = f
# pop all useless clauses (e.g. 'A' is true => pop (A | B))
clauses_same = self.find_clauses(str(p))
# print('clauses_same:', clauses_same)
for c in clauses_same:
self.clauses.pop(c)
# print('pop:', self.clauses.pop(c))
# update related clauses (e.g. 'A' is true => change (~A | B) to B)
neg_p = move_neg(~p)
clauses_diff = self.find_clauses(str(neg_p))
# print('clauses_diff:', clauses_diff)
for c in clauses_diff:
new_args = tuple(omit_items(self.clauses[c].args, neg_p))
if len(new_args) == 1:
self.tell_comp(new_args[0])
# print('new finding:', new_args[0])
else:
self.clauses[c] = merge('|', new_args)
# print('updated:', self.clauses[c])
# add p to KB clauses
self.clauses[str(p)] = p
# find all existing clauses with given symbol
def find_clauses(self, p):
return [c for c in self.clauses if c.find(p) != -1]
# query the entailment for some alpha
def ask(self, alpha):
# skip resolution if we already have the answer in self.symbols
if atm_flag(alpha):
s, f = parse_atomic(alpha)
# print(self.symbols)
if s in self.symbols:
return self.symbols[s] == f
# check entailment
ans = self.resolution(alpha)
# learn new knowledge as needed
if ans:
self.tell(alpha)
return ans
# apply resolution algorithm to check entailment
def resolution(self, alpha):
clauses = deepcopy(self.clauses)
query = to_CNF(Expr('~', alpha))
if isinstance(query, bool):
return False
clauses.update({str(i): i for i in cnf_comp(query)})
while True:
new_clauses = {}
# try to resolve all possible pairs
for c1, c2 in pairs(clauses):
resolvent = resolution_rule(c1, c2)
if resolvent == -1:
continue
if resolvent == Expr('()'):
return True
if str(resolvent) not in clauses:
# print('res:', resolvent)
clauses[str(resolvent)] = resolvent
new_clauses[str(resolvent)] = resolvent
# [keep looping] or [stop here with an answer 'no']
if len(new_clauses):
continue
else:
return False
def __len__(self):
return len(self.clauses)
def __repr__(self):
return str(self.clauses.keys())
def __str__(self):
return str(self.clauses.keys())
# + colab={"base_uri": "https://localhost:8080/"} id="yDtvu4P7za80" executionInfo={"status": "ok", "timestamp": 1638224000423, "user_tz": 300, "elapsed": 11, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhOzSFkObCl-UMX25oW5lPfwwbD5InBzHMI_DJfSw=s64", "userId": "10852634671235052019"}} outputId="dd97a993-152e-437b-ee6a-f6c3cf186299"
# set info_flag to True: print out all steps
info_flag = False
world = {
'Door': (4, 4),
'Ghost': [(2, 1)],
'Oooo': all_adjacents([(2, 1)])
}
KB = KnowledgeBase(*rules('Ghost', 'Oooo'))
step = 0
path = []
max_iter = 1000
exit_door = False
agent_h, agent_w = 0, 0
start_time = time.time()
while (not exit_door) and (step < max_iter):
'''Agent should explore the world at random, probing the
knowledge base for any potential threat. if the KB does not
indicate any specific threat, the agent should move in one of
the adacent (cleared) cell. The simulation ends when the agent
reaches the exit door'''
step += 1
curr_pos = agent_h, agent_w
path.append(curr_pos)
# status check
if curr_pos == world['Door']:
print('\nFound exit at %s. You WIN!' % str(world['Door']))
break
if curr_pos in world['Ghost']:
print('\nYou are killed by the Ghost!')
break
# learning
info = 'Step to (%d, %d);' % (agent_h, agent_w)
KB.tell_comp(expr(get_symbol('~Ghost', curr_pos)))
if curr_pos in world['Oooo']:
info = 'Heard Oooo at (%d, %d);' % (agent_h, agent_w)
KB.tell_comp(expr(get_symbol('Oooo', curr_pos)))
else:
KB.tell_comp(expr(get_symbol('~Oooo', curr_pos)))
# print('KB size:', len(KB))
# print(KB)
next_cells = adjacents(agent_h, agent_w)
while True:
cell = choice(next_cells)
query = expr(get_symbol('~Ghost', cell))
# print('query:', query)
if KB.ask(query):
break
next_cells.remove(cell)
if len(next_cells) == 0:
print('\nNowhere to go!')
exit_door = True
break
agent_h, agent_w = cell
if info_flag:
print(info.ljust(22), end='\n' if step % 5 == 0 else ' ')
if step == max_iter:
print('\nMax iter num is reached!')
print('\ntime: %.2fs' % (time.time() - start_time))
print('path:', path)
print('path length:', len(path))
print('KB size:', len(KB))
# + id="-lgxjacDtDVg"
# set movie_flag to True: generate a movie
movie_flag = False
if movie_flag:
display_movie(world, path, delay=2)
# + [markdown] id="svUtoJQcONm0"
# #### Question 5: Getting used to danger.. (6pts)
#
# Now that our agent knows how to handle a ghost, we will increase the level of risk by including spike traps in the environment.
#
# - Our agent has an edge: evolution has endowed it with a sort of additional ($6^{th}$) sense so that it can feel something 'bad' is about to happen when it is in a cell adjacent to a trap. We represent this ability with the eye combined with the exclamation mark.
#
#
# - Since, as we all know, ghosts are purely imaginary entities, the $6th$ sense only works for the spike traps.
#
#
# - The ghost can still be located by means of the noise it generates which can be heard on all adjacent cells.
#
# <img src="https://github.com/acosse/Artificial-Intelligence-Fall2021/blob/main/Assignments/Assignment3/MazeTrapb.png?raw=1" width="400" height="300"/>
#
# Starting from the agent you designed in the previous questions, improve this agent so that it takes into account the spike traps. You should now have a knowledge base defined on a total of 25*5 symbols describing whether each cell contains a ghost, a 'OOoo' noise, a spike trap, activated the agent's'6th sense', or contains the exit door. The search ends when the agent reaches the door.
# + id="36FVJInGONm0" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1638224004047, "user_tz": 300, "elapsed": 3629, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhOzSFkObCl-UMX25oW5lPfwwbD5InBzHMI_DJfSw=s64", "userId": "10852634671235052019"}} outputId="dd45a8ae-40da-482b-c7db-0a8b9f06fb3f"
# set info_flag to True: print out all steps
info_flag = False
'''
The given world is not escapable regardless of max_iter, so I comment it below.
Instead, Another escapable world setting is used for testing here.
'''
# world = {
# 'Door': (4, 4),
# 'Ghost': [(2, 1)],
# 'Oooo': all_adjacents([(2, 1)]),
# 'Spike': [(1, 3), (3, 4)],
# 'Six': all_adjacents([(1, 3), (3, 4)]),
# }
world = {
'Door': (4, 4),
'Ghost': [(2, 1)],
'Oooo': all_adjacents([(2, 1)]),
'Spike': [(0, 4), (3, 4)],
'Six': all_adjacents([(0, 4), (3, 4)]),
}
KB = KnowledgeBase(*(rules('Ghost', 'Oooo') + rules('Spike', 'Six')))
step = 0
path = []
max_iter = 1000
exit_door = False
agent_h, agent_w = 0, 0
start_time = time.time()
while (not exit_door) and (step < max_iter):
'''Complete the loop with the simulation of the ghost
+ spike trap logical agent. The simulation should start
from the bottom leftmost cell'''
step += 1
curr_pos = agent_h, agent_w
path.append(curr_pos)
# status check
if curr_pos == world['Door']:
print('\nFound exit at %s. You WIN!' % str(world['Door']))
break
if curr_pos in world['Ghost']:
print('\nYou are killed by the Ghost!')
break
if curr_pos in world['Spike']:
print('\nYou are killed by the Spike!')
break
# learning
info = 'Step to (%d, %d);' % (agent_h, agent_w)
KB.tell_comp(expr(get_symbol('~Ghost', curr_pos)))
KB.tell_comp(expr(get_symbol('~Spike', curr_pos)))
if curr_pos in world['Oooo']:
info = 'Heard Oooo at (%d, %d);' % (agent_h, agent_w)
KB.tell_comp(expr(get_symbol('Oooo', curr_pos)))
else:
KB.tell_comp(expr(get_symbol('~Oooo', curr_pos)))
if curr_pos in world['Six']:
info = 'Sixth sense at (%d, %d);' % (agent_h, agent_w)
KB.tell_comp(expr(get_symbol('Six', curr_pos)))
else:
KB.tell_comp(expr(get_symbol('~Six', curr_pos)))
# print('KB size:', len(KB))
# print(KB)
next_cells = adjacents(agent_h, agent_w)
while True:
cell = choice(next_cells)
query1 = expr(get_symbol('~Ghost', cell))
query2 = expr(get_symbol('~Spike', cell))
# print('query:', query1, query2)
if KB.ask(query1) and KB.ask(query2):
break
next_cells.remove(cell)
if len(next_cells) == 0:
print('\nNowhere to go!')
exit_door = True
break
agent_h, agent_w = cell
if info_flag:
print(info.ljust(24), end='\n' if step % 5 == 0 else ' ')
if step == max_iter:
print('\nMax iter num is reached!')
print('\ntime: %.2fs' % (time.time() - start_time))
print('path:', path)
print('path length:', len(path))
print('KB size:', len(KB))
# + id="SBgDQcIgtpvV"
# set movie_flag to True: generate a movie
movie_flag = False
if movie_flag:
display_movie(world, path, delay=1)
# + [markdown] id="-ZuIlIOpONm1"
# #### Bonus: For where your treasure is.. (6pts)
#
# We finally consider the whole environment. This environment is composed of all the elements from the previous questions but it now also includes a treasure chest. The final objective this time is to find the chest first and then reach the exit. Although some of the previous symbols are omitted for clarity, the ghost can always be located by means of the sound it produces, the agent can still trust its $6^{\text{th}}$ sense regarding the spike trap and the treasure chest can be perceived in adjacent cells, by means of the shine it produces.
#
# When the knowledge base does not indicate any threat, the agent should move at random in one of the adjacent cells.
#
# The world now contains a total of 25*7 symbols.
#
# + [markdown] id="zA06HnNMONm2"
# <img src="https://github.com/acosse/Artificial-Intelligence-Fall2021/blob/main/Assignments/Assignment3/fullWorldChest.png?raw=1" width="400" height="300"/>
# + id="Kflva-_RONm2" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1638224010970, "user_tz": 300, "elapsed": 6926, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhOzSFkObCl-UMX25oW5lPfwwbD5InBzHMI_DJfSw=s64", "userId": "10852634671235052019"}} outputId="e465ea5c-df5e-469b-acfb-9eb0f53a6113"
# set info_flag to True: print out all steps
info_flag = True
'''
The given world is not escapable regardless of max_iter, so I comment it below.
Instead, Another escapable world setting is used for testing here.
'''
# world = {
# 'Door': (4, 4),
# 'Treasure': (3, 2),
# 'Shine': adjacents(3, 2),
# 'Ghost': [(2, 1)],
# 'Oooo': all_adjacents([(2, 1)]),
# 'Spike': [(1, 2), (2, 0), (3, 4)],
# 'Six': all_adjacents([(1, 2), (2, 0), (3, 4)]),
# }
world = {
'Door': (4, 4),
'Treasure': (3, 2),
'Shine': adjacents(3, 2),
'Ghost': [(2, 1)],
'Oooo': all_adjacents([(2, 1)]),
'Spike': [(2, 0), (3, 4)],
'Six': all_adjacents([(2, 0), (3, 4)]),
}
KB = KnowledgeBase(*(rules('Ghost', 'Oooo') + rules('Spike', 'Six') + \
rules('Treasure', 'Shine')))
step = 0
path = []
max_iter = 1000
found_exit = False
found_treasure = False
agent_h, agent_w = 0, 0
start_time = time.time()
# (not (found_exit and found_treasure))
while step < max_iter:
'''Simulation should complete when the treasure chest
has been found and the exit has been reached.
The agent should start from bottom leftmost cell'''
step += 1
curr_pos = agent_h, agent_w
path.append(curr_pos)
# status check
if (curr_pos == world['Door']) and (not found_exit):
print('\nFound exit at %s.\n' % str(world['Door']))
found_exit = True
if (curr_pos == world['Treasure']) and (not found_treasure):
print('\nFound treasure at %s.\n' % str(world['Treasure']))
found_treasure = True
if found_exit and found_treasure:
break
if curr_pos in world['Ghost']:
print('\nYou are killed by the Ghost!')
break
if curr_pos in world['Spike']:
print('\nYou are killed by the Spike!')
break
# learning
info = 'Step to (%d, %d);' % (agent_h, agent_w)
KB.tell_comp(expr(get_symbol('~Ghost', curr_pos)))
KB.tell_comp(expr(get_symbol('~Spike', curr_pos)))
if curr_pos in world['Oooo']:
info = 'Heard Oooo at (%d, %d);' % (agent_h, agent_w)
KB.tell_comp(expr(get_symbol('Oooo', curr_pos)))
else:
KB.tell_comp(expr(get_symbol('~Oooo', curr_pos)))
if curr_pos in world['Six']:
info = 'Sixth sense at (%d, %d);' % (agent_h, agent_w)
KB.tell_comp(expr(get_symbol('Six', curr_pos)))
else:
KB.tell_comp(expr(get_symbol('~Six', curr_pos)))
if curr_pos in world['Shine']:
info = 'Shine at (%d, %d);' % (agent_h, agent_w)
KB.tell_comp(expr(get_symbol('Shine', curr_pos)))
else:
KB.tell_comp(expr(get_symbol('~Shine', curr_pos)))
# print('KB size:', len(KB))
# print(KB)
next_cells = adjacents(agent_h, agent_w)
while True:
cell = choice(next_cells)
query1 = expr(get_symbol('~Ghost', cell))
query2 = expr(get_symbol('~Spike', cell))
# print('query:', query1, query2)
if KB.ask(query1) and KB.ask(query2):
break
next_cells.remove(cell)
if len(next_cells) == 0:
print('\nNowhere to go!')
exit_door = True
break
agent_h, agent_w = cell
if info_flag:
print(info.ljust(24), end='\n' if step % 5 == 0 else ' ')
if step == max_iter:
print('Max iter num is reached!')
else:
print('You WIN!')
print('\ntime: %.2fs' % (time.time() - start_time))
print('path:', path)
print('path length:', len(path))
print('KB size:', len(KB))
# + colab={"base_uri": "https://localhost:8080/", "height": 379} id="20vuwTbe2tlq" executionInfo={"status": "ok", "timestamp": 1638224025385, "user_tz": 300, "elapsed": 14419, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhOzSFkObCl-UMX25oW5lPfwwbD5InBzHMI_DJfSw=s64", "userId": "10852634671235052019"}} outputId="ff4f148a-d8b3-446c-8f46-410d6ea8f73d"
# set movie_flag to True: generate a movie
movie_flag = True
if movie_flag:
display_movie(world, path, delay=2)
# + [markdown] id="-20nH_Ukvmyi"
# #### **Final Thought**
# To speed up the agent, we can use a list/set to store visited positions and prioritize unvisited cells. Considering the fact that the questions require the agent to move randomly, I didn't implement that part, but it should work anyway.
#
# #### Wish you have a great day! :-)
| assignments/AI_F2021_a3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# name: python3
# ---
# # Differentiation and Searching
# [Numerical differentiation](https://personal.math.ubc.ca/~pwalls/math-python/differentiation/differentiation/) in really straightforward, where we will evaluate the forward, central, and extrapolated-difference methods. Another tool is trial-and-error searching which can be used in root-finding, minimization, and is widely used to solve problems where analytic solutions do not exist or are impractical.
#
# ## Numerical differentiation
# Before implementing a numerical method for differentiation, let's review some basics from Introductory Calculus. The elementery definition was
#
# $\frac{dy(t)}{dt} = \displaystyle \lim_{h\rightarrow 0}\frac{y(t+h)-y(t)}{h}$,
#
# but this is impractical to implement in a computer. The denominator is approaching zero and the finite machine prescision causes oscillations in the solution. Therefore we implement a series of approximations to obtain a practical solution.
#
# ### Forward difference
# The most direct method for numerical differentiaion starts by expanding a function in a Taylor series to obtain its value a small step *h* away:
#
# $y(t+h) = y(t) + h\frac{dy(t)}{dt} + \frac{h^2}{2!}\frac{d^2y(t)}{dt^2} + \frac{h^3}{3!}\frac{d^3y(t)}{dt^3} + \cdots$.
#
# We obtain a *forward-difference* derivative by using the approximation in the elementary definition for $y^\prime(t)$:
#
# $\frac{dy(t)}{dt}\biggr\rvert_{fd} = \frac{y(t+h)-y(t)}{h}$.
#
# An approximation for the error follows from substituting the Taylor series:
#
# $\frac{dy(t)}{dt}\biggr\rvert_{fd} \simeq \frac{dy(t)}{dt} + \frac{h}{2}\frac{d^2y(t)}{dt^2} + \cdots$.
#
# You can think of this approximation as using two points to represent the function by a straight line between $x$ and $x+h$. The approximation has an error proportional to $h$ (the slope of the line). Therefore, we can make the approximation error smaller by taking smaller steps (decreasing $h$), yet there will be some round-off error that is present from the subtraction in the *forward-differenece* derivative. To see how this algorithm works, let $y(t) = a + bt^2$, where the exact derivative is $y^\prime = 2bt$ and the computed derivative is:
#
# $\frac{dy(t)}{dt}\biggr\rvert_{fd} \simeq \frac{y(t+h)-y(t)}{h} = \frac{a+b(t+h)^2 - a - bt^2}{h} = \frac{2bth + bh^2}{h} = 2bt + bh$.
#
# This approximation is only good for small *h*, where the $bh \ll 1$. Implementing the *forward-difference* algorithm in python is super easy. Simply use the *diff* function from **numpy**. Suppose you have an array of *N* values in *y*, then *np.diff(y)* returns an array with length *N-1* containing the consecutive differences (i.e., y[1]-y[0], y[2]-y[1], ...). If the values are sampled at regular intervals in the *t* array, then $y^\prime(t)$ = np.diff(y)/(t[1]-t[0]).
#
#
# ### Central difference
# An improved approximation uses the principles from the Taylor series (i.e., expand relative to a central point). Now, rather than making a single step of *h* forward, we form a *central difference* by stepping forward a half-step and backward a half-step:
#
# $\frac{dy(t)}{dt}\biggr\rvert_{cd} = \frac{y(t+h/2)-y(t-h/2)}{h}$.
#
# We estimate the error by substituting the Taylor series for $y(t\pm h/2)$. **Do you think this will result in a better approximation? Why?** By substitution, we find:
#
# $y(t+\frac{h}{2})-y(t-\frac{h}{2}) \simeq \left[y(t) + \frac{h}{2}y^\prime(t) + \frac{h^2}{8}y^{\prime\prime}(t) + \frac{h^3}{48}y^{\prime\prime\prime}(t) + O(h^4)\right] - \left[y(t) - \frac{h}{2}y^\prime(t) + \frac{h^2}{8}y^{\prime\prime}(t) - \frac{h^3}{48}y^{\prime\prime\prime}(t) + O(h^4)\right]$
#
# and through many cancellations, we find:
#
# $y(t+\frac{h}{2})-y(t-\frac{h}{2}) \simeq hy^\prime(t) + \frac{h^3}{24}y^{\prime\prime\prime}(t) + O(h^5)$,
#
# and the *central-difference* derivative is:
#
# $\frac{dy(t)}{dt}\biggr\rvert_{cd} = y^\prime(t) + \frac{h^2}{24}y^{\prime\prime\prime}(t) + O(h^4)$.
#
# The important difference between the *forward-difference* and *central-difference* algorithms is that the *central-difference* algorithm cancels the terms with an even power of *h* in from each of the Taylor series. This makes the leading-order error of *central-difference* accurate to $h^2$, while the *forward-difference* is only accurate to order *h*. Returning to our parabola example ($y(t) = a + bt^2$), we see that the *central-difference* gives the exact derivative independent of $h$:
#
# $\frac{dy(t)}{dt}\biggr\rvert_{cd} \simeq \frac{y(t+h/2)-y(t-h/2)}{h} = \frac{a+b(t+h/2)^2 - a - b(t-h/2)^2}{h} = \frac{2bth}{h} = 2bt$.
#
# The **numpy** module has a special function called [gradient](https://numpy.org/doc/stable/reference/generated/numpy.gradient.html) that performs a second order central differences method on the input array and requires the stepsize *h* as an argument. From an array with *N* elements $y(t)$, it returns an array of *N* elements for $y^\prime(t)$.
# ### Extrapolated difference
# You may have noticed that different differentiation rules will keep a different number of terms in a Taylor series expansion, which also providesan expression for the error. We can reduce the error further by combining several algorithims together so that the summed errors extrapolate to zero. For example, we could employ two *central-difference* algorithms as a linear combination, where the first algorithm takes half-steps and the second takes quarter-steps. Let's look at the quarter-step approximation ($h\rightarrow h/2$):
#
# $\frac{dy(t)}{dt}\biggr\rvert_{cd}^{h/4} = \frac{y(t+h/4)-y(t-h/4)}{h/2} = y^\prime(t) + \frac{h^2}{96}y^{\prime\prime\prime}(t) + O(h^4)$.
#
# The quarter-step derivative produces a factor of 1/96 instead of the 1/24 on the second order term. If we want to cancel this term, we realize that 96 = 4*24 and so we perform the following subtraction to get:
#
# $4\frac{dy(t)}{dt}\biggr\rvert_{cd}^{h/4} - \frac{dy(t)}{dt}\biggr\rvert_{cd}^{h/2} = 4y^\prime(t) + \frac{h^2}{24}y^{\prime\prime\prime}(t) - y^\prime(t) - \frac{h^2}{24}y^{\prime\prime\prime}(t) = 3 y^\prime(t) + O(h^4)$
#
# Thus, we find for the *extrapolated-difference* the following:
#
# $\frac{dy(t)}{dt}\biggr\rvert_{ed} = \frac{4}{3}\frac{dy(t)}{dt}\biggr\rvert_{cd}^{h/4} - \frac{1}{3}\frac{dy(t)}{dt}\biggr\rvert_{cd}^{h/2} \simeq y^\prime(t) + O(h^4)$
# ## Second Derivatives
# Many problems in physics involve a second derivative, or acceleration. One of the most important equations is Newton's second law that tells us that a force $F$ and acceleration $a$ are linearly related:
#
# $F = ma = m\frac{d^2y}{dt^2}$,
#
# where $m$ is a particle's mass and the acceleration can be expressed as the second derivative of a function $y(t)$. If we can determin the acceleration, then we can determine the force. Using the *central-difference* method for the first derivative, we have:
#
# $\frac{dy(t)}(dt) \simeq \frac{y(t+h/2)-y(t-h/2)}{h}$,
#
# which gives the derivative at t by moving forward and backward from t by a half-step $h/2$. We take the second derivative by applying the same operation again as follows:
#
# $\frac{d^2y(t)}{dt^2} \simeq \frac{y^\prime(t+h/2)-y^\prime(t-h/2)}{h} \simeq \frac{[y(t+h)-y(t)]-[y(t)-y(t-h)]}{h^2} = \frac{y(t+h)-2y(t)+y(t-h)}{h^2}$
#
# ## Root Via Bisection
# Trial-and-error root finding looks for a vlaue of $x$ at which $f(x)=0$, where the $f(x)$ can be polynomials or a trancendental function (e.g., $10\sin x = 3x^3 \rightarrow 10\sin x - 3x^3 = 0$). The search procedure starts with a guess for $x$, substitutes that guess into $f(x)$ (i.e., the trial), and sees how far from zero the evaluated function is (i.e., the error). the program then revises the guess $x$ based on the error and then substitutes the revised guess into $f(x)$ again. The procedure continues until $f(x)\simeq 0$ or to some desired level of precision. Sometimes the procedure is stopped once it appears progress is not being made.
#
# The most elementary trial-and-error technique is the *bisection algorithm*. It is reliable, but slow. If you know some interval in which $f(x)$ changes sign, then the [bisection algorithm](https://personal.math.ubc.ca/~pwalls/math-python/roots-optimization/bisection/) is the way to go. The basis of the bisection algorithm is that we start with two values of $x$ between which a zero occurs (i.e., the bounds). Let us say that $f(x)$ is negative at $a$ and positive at $b$. Thus we start with the interval $a\geq x \geq b$ within which we know a zero occurs. Then we evaluate each of these tasks for a given iteration:
#
# 1. Calculate $c$, the midpoint of the interval, $c = \frac{a+b}{2}$.
# 2. Evaluate the function at the midpoint, $f(c)$.
# 3. If convergence is satisfactory (i.e., $|f(c)|\simeq 0$), return c and stop iterating.
# 4. Examine the sign of $f(c)f(a)$.
# - If negative then set $b = c$ and return to step 1.
# - Else set $a = c$ and return to step 1.
#
# Let's turn these steps into code:
def bisection(f,a,b,N):
'''Approximate solution of f(x)=0 on interval [a,b] by bisection method.
f : function
a,b : The interval in which to search for a solution.
N : The number of iterations to implement.'''
if f(a)*f(b) >= 0: #checking that a zero exist in the interval [a,b]
print("Bisection method fails.")
return None
#initial interval
a_n = a
b_n = b
for n in range(1,N+1):
c = (a_n + b_n)/2 #calculate midpoint
f_c = f(c) #evaluate function at midpoint
if f(a_n)*f_c < 0: #evaluate sign
b_n = c
elif f(b_n)*f_c < 0: #evaluate sign
a_n = c
elif f_c == 0:
print("Found exact solution.")
return m_n
else:
print("Bisection method fails.")
return None
return (a_n + b_n)/2
# Now let's try our function on a problem from quantum mechanics. The most standard problem is to solve for the energies of a particle of mass $m$ bound within a 1-D square well of radius $a$. The potential $V(x) = -V_o$ if $|x|\leq a$, otherwised it is zero. From quantum mechanics textbooks we know there is a [solution](https://ocw.mit.edu/courses/physics/8-04-quantum-physics-i-spring-2016/lecture-notes/MIT8_04S16_LecNotes11.pdf) for the energies of the bound states $E_B$:
#
# $\sqrt{V_o - E_B}\tan(\sqrt{V_o - E_B}) = \sqrt{E_B}$ (even),
#
# and
#
# $\sqrt{V_o - E_B}\cot(\sqrt{V_o - E_B}) = \sqrt{E_B}$ (odd).
#
# Here we have chosen units such that $\hbar=1$, $2m=1$, and $a=1$. Now we want to find several of the bound state energies for the even wave functions. First we have to find a reasonable interval and then apply our algorithm.
# +
import numpy as np
import matplotlib.pyplot as plt
from scipy.optimize import bisect
V_o = 10 #define the potential
def even_func(E_B):
#E_B is a guess at the ground state energy
if (V_o - E_B)>= 0:
temp = np.sqrt(V_o - E_B)
return temp*np.tan(temp) - np.sqrt(E_B)
else:
return np.nan
fig = plt.figure(1,figsize=(8,8))
ax = fig.add_subplot(111)
x_rng = np.arange(0,15,0.1)
for x in x_rng:
y = even_func(x)
ax.plot(x,y,'k.',ms=10)
#root_EB = bisection(even_func,8,10,50)
#scipy_root = bisect(even_func,8,10)
#print("our method = ",root_EB)
#print("scipy method = ",scipy_root)
#ax.plot(root_EB,even_func(root_EB),'r.',ms=10)
ax.grid(True)
ax.set_ylim(-30,30)
ax.set_xlabel("Bound State Energy$ E_B$",fontsize=20)
ax.set_ylabel("1D finite well solution",fontsize=20)
# -
# ## Newton-Raphson searching
# The Newton-Raphson (NR) algorithm is another way to find roots, but it is much quicker than the bisection method. The NR method is equivalent to drawing a straight line $f(x)\simeq mx + b$ to a curve at an $x$ value for which $f(x) \simeq 0$. Then it uses the intercept of the line with the $x$ axis at $-b/m$ as an imporved guess for the root. If the "curve" were a straight line, then the answer would be exact. Otherwise, it is a good approximation if the guess is close enough to the root for $f(x) to be nearly linear.
#
# As an [iteration](https://personal.math.ubc.ca/~pwalls/math-python/roots-optimization/newton/) scheme, we need to start with a guess $x_o$, then find a correction to the guess $\Delta x$, and finally formulate a new guess $x = x_o + \Delta x$. Recall that the equation of a line can be formulated in terms of a Taylor series, keeping on the first two terms:
#
# $f(x) \simeq f(x_o) + \frac{df}{dx}\biggr\rvert_{x_o}\Delta x$.
#
# We then determine the correction $\Delta x$ by calculating the point at which this linear appromation to $f(x)$ would cross the $x$ axis:
#
# $f(x_o) + \frac{df}{dx}\biggr\rvert_{x_o}\Delta x = 0$
#
# $\Delta x = -\frac{f(x_o)}{df/dx\rvert_{x_o}}$.
#
# Notice that the NR method requires the calculation of the first derivative $df/dx$ at each guess. In many cases you may have an analytic expression for the derivative and can build it into the algorithm. However, it is simpler to use a numerical *forward-difference* approximation to the derivative. While a central-difference approximation would be more accurate, it would require addition function evaluations and the overall process would take longer to run.
# ## Problems
# - Complete the following problems in a Jupyter notebook, where you will save your results as an external file (*.png) as needed.
# - Create a LaTex document with:
# - an abstract summary
# - sections for each problem that state the problem, summarize what you did, and display the results
# - include a reference for each solution (this can be textbooks)
#
# 1. Write a program to calculate sin(x) and cos(x). Calculate and plot the numerical derivatives using a) forward-difference, b) central-difference, and c) the analytical derivative.
#
# 2. Write a function to implement the Newton-Raphson method. Apply it to the 1D Quantum well example and compare it to the bisection method. Also compare your implementation to the function from *scipy.optimize*.
#
| docs/_sources/docs/Chapter_5/Differentiation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:.conda-raccoon]
# language: python
# name: conda-env-.conda-raccoon-py
# ---
# Convert to and from Pandas DataFrames
# ====================
# There are no built in methods for the conversions but these functions below should work in most basic instances.
# + pycharm={"is_executing": false}
# remove comment to use latest development version
import sys; sys.path.insert(0, '../')
# -
import raccoon as rc
import pandas as pd
# Raccoon to Pandas
# ----------------
#
def rc_to_pd(raccoon_dataframe):
"""
Convert a raccoon dataframe to pandas dataframe
:param raccoon_dataframe: raccoon DataFrame
:return: pandas DataFrame
"""
data_dict = raccoon_dataframe.to_dict(index=False)
return pd.DataFrame(data_dict, columns=raccoon_dataframe.columns, index=raccoon_dataframe.index)
rc_df = rc.DataFrame({'a': [1, 2, 3], 'b': [4, 5, 6]}, columns=['a', 'b'], index=[7, 8, 9])
print(type(rc_df))
print(rc_df)
pd_df = rc_to_pd(rc_df)
print(type(pd_df))
print(pd_df)
# Pandas to Raccoon
# -----------
def pd_to_rc(pandas_dataframe):
"""
Convert a pandas dataframe to raccoon dataframe
:param pandas_dataframe: pandas DataFrame
:return: raccoon DataFrame
"""
columns = pandas_dataframe.columns.tolist()
data = dict()
pandas_data = pandas_dataframe.values.T.tolist()
for i in range(len(columns)):
data[columns[i]] = pandas_data[i]
index = pandas_dataframe.index.tolist()
index_name = pandas_dataframe.index.name
index_name = 'index' if not index_name else index_name
return rc.DataFrame(data=data, columns=columns, index=index, index_name=index_name)
pd_df = pd.DataFrame({'a': [1, 2, 3], 'b': [4, 5, 6]}, index=[5, 6, 7], columns=['a', 'b'])
print(type(pd_df))
print(pd_df)
rc_df = pd_to_rc(pd_df)
print(type(rc_df))
print(rc_df)
| examples/convert_pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Importing the IRIS Models from S3 and Caching them
#
# This notebook demonstrates how to import a machine learning model file from the S3 and store the Models + Analysis in the redis cache named "**CACHE**".
#
# ## Overview
#
# Download the S3 IRIS Model archive from the configured S3 Bucket + Key and decompress + extract the pickled historcial analysis and previously built models. This includes examples from the IRIS sample dataset and requires you to have a valid S3 Bucket storing the models and are comfortable paying for the download costs for downloading the objects from S3 (https://aws.amazon.com/s3/pricing/).
#
# Once uploaded to the S3 Bucket you should be able to view the files have a similar disk size:
#
# 
#
# After importing, the Models and Analysis are available to any other Sci-pype instance with connectivity to the same redis cache.
#
# ### Command-line Versions
#
# I built this notebook from the importer examples:
#
# https://github.com/jay-johnson/sci-pype/tree/master/bins/ml/importers
# ### 1) Setup the Environment for Importing the IRIS Classifier Model File
# +
# Setup the Sci-pype environment
import sys, os
# Only redis is needed for this notebook:
os.environ["ENV_DEPLOYMENT_TYPE"] = "JustRedis"
# Load the Sci-pype PyCore as a named-object called "core" and environment variables
from src.common.load_ipython_env import *
# -
# ### 2) Setup the Request
#
# Import the Models from S3 and store the extracted Models + Analysis in the Cache.
#
# Please make sure the environment variables are set correctly and the S3 Bucket exists:
#
# ```
# ENV_AWS_KEY=<AWS API Key>
# ENV_AWS_SECRET=<AWS API Secret>
# ```
#
# For docker containers make sure to set these keys in the correct Jupyter env file and restart the container:
#
# ```
# <repo base dir>/justredis/jupyter.env
# <repo base dir>/local/jupyter.env
# <repo base dir>/test/jupyter.env
# ```
# - What's the dataset name?
ds_name = "iris_classifier"
# - Where is the downloaded file getting stored?
data_dir = str(os.getenv("ENV_DATA_SRC_DIR", "/opt/work/data/src"))
if not os.path.exists(data_dir):
os.mkdir(data_dir, 0777)
# - What's the S3 Location (Unique Bucket Name + Key)?
s3_bucket = "unique-bucket-name-for-datasets" # name this something under your AWS Account (This might be open to the public in the future...stay tuned)
s3_key = "dataset_" + core.to_upper(ds_name) + ".cache.pickle.zlib"
s3_loc = str(s3_bucket) + ":" + str(s3_key)
# - Where will the downloaded file be stored?
ml_file = data_dir + "/" + str(s3_key)
# - Check if the Model File needs to be Downloaded
# +
lg("-------------------------------------------------", 6)
lg("Importing Models and Analysis from S3 into Caching Models from CACHE - S3Loc(" + str(s3_loc) + ") File(" + str(ml_file) + ")", 6)
lg("", 6)
if os.path.exists(ml_file) == False:
s3_loc = str(s3_bucket) + ":" + str(s3_key)
lg("Downloading ModelFile S3Loc(" + str(s3_loc) + ")", 6)
download_results = core.s3_download_and_store_file(s3_loc, ml_file, core.get_rds(), core.get_dbs(), debug)
if download_results["Status"] != "SUCCESS":
lg("ERROR: Stopping processing for errror: " + str(download_results["Error"]), 0)
else:
lg("", 6)
lg("Done Downloading ModelFile S3Loc(" + str(s3_loc) + ") File(" + str(download_results["Record"]["File"]) + ")", 5)
ml_file = download_results["Record"]["File"]
else:
lg("", 6)
lg("Continuing with the existing file.", 5)
lg("", 6)
# end of downloading from s3 if it's not locally available
# -
# - Start the Importer to Download + Cache the Models out of S3 or Locally if the file already exists
# +
ra_name = "CACHE"
lg("Importing(" + str(ml_file) + ") Models and Analysis into Redis(" + str(ra_name) + ")", 6)
cache_req = {
"RAName" : ra_name,
"DSName" : str(ds_name),
"TrackingID": "",
"ModelFile" : ml_file,
"S3Loc" : s3_loc
}
upload_results = core.ml_load_model_file_into_cache(cache_req, core.get_rds(), core.get_dbs(), debug)
if upload_results["Status"] == "SUCCESS":
lg("", 6)
lg("Done Loading Model File for DSName(" + str(ds_name) + ") S3Loc(" + str(cache_req["S3Loc"]) + ")", 5)
lg("", 6)
lg("Importing and Caching Completed", 5)
lg("", 6)
else:
lg("", 6)
lg("ERROR: Failed Loading Model File(" + str(cache_req["ModelFile"]) + ") into Cache for DSName(" + str(ds_name) + ")", 6)
lg(upload_results["Error"], 6)
lg("", 6)
# end of if success
# -
# ### 3) Setup to Import and Cache the IRIS Regressor Models and Analysis
ds_name = "iris_regressor"
# ### 4) Import the IRIS Regressor Models from S3 and Store them in Redis
# +
s3_bucket = "unique-bucket-name-for-datasets" # name this something under your AWS Account (This might be open to the public in the future...stay tuned)
s3_key = "dataset_" + core.to_upper(ds_name) + ".cache.pickle.zlib"
s3_loc = str(s3_bucket) + ":" + str(s3_key)
ra_name = "CACHE"
ml_file = data_dir + "/" + str(s3_key)
lg("-------------------------------------------------", 6)
lg("Importing Models and Analysis from S3 into Caching Models from CACHE - S3Loc(" + str(s3_loc) + ") File(" + str(ml_file) + ")", 6)
lg("", 6)
if os.path.exists(ml_file) == False:
s3_loc = str(s3_bucket) + ":" + str(s3_key)
lg("Downloading ModelFile S3Loc(" + str(s3_loc) + ")", 6)
download_results = core.s3_download_and_store_file(s3_loc, ml_file, core.get_rds(), core.get_dbs(), debug)
if download_results["Status"] != "SUCCESS":
lg("ERROR: Stopping processing for errror: " + str(download_results["Error"]), 0)
else:
lg("", 6)
lg("Done Downloading ModelFile S3Loc(" + str(s3_loc) + ") File(" + str(download_results["Record"]["File"]) + ")", 5)
ml_file = download_results["Record"]["File"]
else:
lg("", 6)
lg("Continuing with the existing file.", 5)
lg("", 6)
# end of downloading from s3 if it's not locally available
lg("Importing(" + str(ml_file) + ") Models and Analysis into Redis(" + str(ra_name) + ")", 6)
cache_req = {
"RAName" : ra_name,
"DSName" : str(ds_name),
"TrackingID" : "",
"ModelFile" : ml_file,
"S3Loc" : s3_loc
}
upload_results = core.ml_load_model_file_into_cache(cache_req, core.get_rds(), core.get_dbs(), debug)
if upload_results["Status"] == "SUCCESS":
lg("", 6)
lg("Done Loading Model File for DSName(" + str(ds_name) + ") S3Loc(" + str(cache_req["S3Loc"]) + ")", 5)
lg("", 6)
lg("Importing and Caching Completed", 5)
lg("", 6)
else:
lg("", 6)
lg("ERROR: Failed Loading Model File(" + str(cache_req["ModelFile"]) + ") into Cache for DSName(" + str(ds_name) + ")", 6)
lg(upload_results["Error"], 6)
lg("", 6)
# end of if success
# -
# ### Automation with Lambda - Coming Soon
#
# Native lambda uploading support will be added in the future. Packaging and functionality still need to be figured out. For now, you can extend the command line versions for the extractors below.
# #### Command-line Versions
#
# I built this notebook from the importer examples:
#
# https://github.com/jay-johnson/sci-pype/tree/master/bins/ml/importers
| examples/ML-IRIS-Import-and-Cache-Models-From-S3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
import panel as pn
pn.extension()
# The ``Toggle`` widget allows toggling a single condition between ``True``/``False`` states. This widget is interchangeable with the ``Checkbox`` widget.
#
# For more information about listening to widget events and laying out widgets refer to the [widgets user guide](../../user_guide/Widgets.ipynb). Alternatively you can learn how to build GUIs by declaring parameters independently of any specific widgets in the [param user guide](../../user_guide/Param.ipynb). To express interactivity entirely using Javascript without the need for a Python server take a look at the [links user guide](../../user_guide/Param.ipynb).
#
# #### Parameters:
#
# For layout and styling related parameters see the [customization user guide](../../user_guide/Customization.ipynb).
#
# ##### Core
#
# * **``value``** (boolean): Whether the button is toggled or not
#
# ##### Display
#
# * **``button_type``** (str): A button theme should be one of ``'default'`` (white), ``'primary'`` (blue), ``'success'`` (green), ``'info'`` (yellow), or ``'danger'`` (red)
# * **``disabled``** (boolean): Whether the widget is editable
# * **``name``** (str): The title of the widget
#
# ___
# +
toggle = pn.widgets.Toggle(name='Toggle', button_type='success')
toggle
# -
# ``Toggle.value`` is either True or False depending on whether the button is toggled:
toggle.value
# The color of the ``Toggle`` can be selected using one of the available ``button_type``s:
pn.Column(*(pn.widgets.Toggle(name=p, button_type=p) for p in pn.widgets.Toggle.param.button_type.objects))
# ### Controls
#
# The `Toggle` widget exposes a number of options which can be changed from both Python and Javascript. Try out the effect of these parameters interactively:
pn.Row(toggle.controls(jslink=True), toggle)
| examples/reference/widgets/Toggle.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Notebook to draw the errors of the predicted trip durations from a selected location.
#
# ### Import the required modules.
# +
import os
import arcpy
from spark_esri import spark_start, spark_stop
from pyspark.sql.functions import col, lit
# -
# ### Start a Spark instance.
#
# Note the `config` argument to [configure the Spark instance](https://spark.apache.org/docs/latest/configuration.html).
# +
spark_stop()
config = {"spark.driver.memory":"2G"}
spark = spark_start(config=config)
# -
# ### Create a Spark data frame of the selected predictions features, and create a view named 'v0'.
#
# A new column (`error`) is added which is the square of the difference between `duration` and `duration_predicted`.
# +
fields = ['plon','plat','dlon','dlat','duration','duration_predicted']
schema = ",".join([f"{f} double" for f in fields])
with arcpy.da.SearchCursor("Predictions",fields) as data:
spark\
.createDataFrame(data,schema)\
.withColumn("delta",col("duration")-col("duration_predicted"))\
.withColumn("error", col("delta")*col("delta"))\
.drop("delta")\
.createOrReplaceTempView("v0")
# -
# ### Calculate the average of the pickup locations.
# +
rows = spark\
.sql("""select avg(plon) plon,avg(plat) plat from v0""")\
.collect()
plon,plat = rows[0]
# -
# ### Aggregate the dropoff location at bin locations.
# +
cell1 = 0.05
cell2 = cell1 * 0.5
spark\
.sql(f"""
select
cast(dlon/{cell1} as long) dq,
cast(dlat/{cell1} as long) dr,
error
from v0
""")\
.createOrReplaceTempView('v1')
rows = spark\
.sql(f"""
select
dq*{cell1}+{cell2} dlon,
dr*{cell1}+{cell2} dlat,
avg(error) mse
from v1
group by dq,dr
""")\
.collect()
# -
# ### Create an in-memory linestring features between the avg pickup location and the dropoff bins.
# +
ws = "memory"
nm = "Trips"
fc = os.path.join(ws,nm)
arcpy.management.Delete(fc)
sp_ref = arcpy.SpatialReference(4326)
arcpy.management.CreateFeatureclass(ws,nm,"POLYLINE",spatial_reference=sp_ref)
arcpy.management.AddField(fc, "MSE", "DOUBLE")
with arcpy.da.InsertCursor(fc, ["SHAPE@WKT","MSE"]) as cursor:
for dlon,dlat,mse in rows:
wkt = f"LINESTRING({plon} {plat},{dlon} {dlat})"
cursor.insertRow((wkt,mse))
# -
# ### Stop the spark instance.
spark_stop()
| taxi_trips_duration_error.ipynb |