code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/tjHendrixx/DS-Unit-1-Sprint-1-Dealing-With-Data/blob/master/DS_Unit_1_Sprint_Challenge_1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="NooAiTdnafkz" colab_type="text"
# # Data Science Unit 1 Sprint Challenge 1
#
# ## Loading, cleaning, visualizing, and analyzing data
#
# In this sprint challenge you will look at a dataset of the survival of patients who underwent surgery for breast cancer.
#
# http://archive.ics.uci.edu/ml/datasets/Haberman%27s+Survival
#
# Data Set Information:
# The dataset contains cases from a study that was conducted between 1958 and 1970 at the University of Chicago's Billings Hospital on the survival of patients who had undergone surgery for breast cancer.
#
# Attribute Information:
# 1. Age of patient at time of operation (numerical)
# 2. Patient's year of operation (year - 1900, numerical)
# 3. Number of positive axillary nodes detected (numerical)
# 4. Survival status (class attribute)
# -- 1 = the patient survived 5 years or longer
# -- 2 = the patient died within 5 year
#
# Sprint challenges are evaluated based on satisfactory completion of each part. It is suggested you work through it in order, getting each aspect reasonably working, before trying to deeply explore, iterate, or refine any given step. Once you get to the end, if you want to go back and improve things, go for it!
# + [markdown] id="5wch6ksCbJtZ" colab_type="text"
# ## Part 1 - Load and validate the data
#
# - Load the data as a `pandas` data frame.
# - Validate that it has the appropriate number of observations (you can check the raw file, and also read the dataset description from UCI).
# - Validate that you have no missing values.
# - Add informative names to the features.
# - The survival variable is encoded as 1 for surviving >5 years and 2 for not - change this to be 0 for not surviving and 1 for surviving >5 years (0/1 is a more traditional encoding of binary variables)
#
# At the end, print the first five rows of the dataset to demonstrate the above.
# + id="83WvQtjwuu2L" colab_type="code" colab={}
import requests
import pandas as pd
import io
urlData = requests.get('http://archive.ics.uci.edu/ml/machine-learning-databases/haberman/haberman.data').content
rawData = pd.read_csv(io.StringIO(urlData.decode('utf-8')), header=None)
df = rawData
# + id="9p3lAXTbu4Qc" colab_type="code" outputId="1bb3258a-5c0b-44ec-c537-ace2606fb62d" colab={"base_uri": "https://localhost:8080/", "height": 204}
df.head()
# + id="IOI_XtMTu9cV" colab_type="code" outputId="44ff52b0-a634-4937-eeac-c14a8732fa45" colab={"base_uri": "https://localhost:8080/", "height": 102}
df.isnull().sum()
# + id="FNd4xXv2vOhJ" colab_type="code" colab={}
df.columns = ['Age', 'Year of Operation', '# of +Aux Nodes', 'Survival Status']
# + id="i-9S7R8AvYlk" colab_type="code" outputId="b4381950-8c10-4f26-a4db-561134960c63" colab={"base_uri": "https://localhost:8080/", "height": 204}
df.head()
# + id="hRK79Pduvb-K" colab_type="code" outputId="9bba5e69-79ee-4570-b033-6360a035afb4" colab={"base_uri": "https://localhost:8080/", "height": 102}
df.dtypes
# + id="v5KffBrxvfzu" colab_type="code" colab={}
df['Survival Status'].replace(1,0, inplace=True)
# + id="p4qzXA3Kvi-8" colab_type="code" colab={}
df['Survival Status'].replace(2,1, inplace=True)
# + id="schTPYeRvlxt" colab_type="code" outputId="381494a3-3bc9-40b1-d70f-385bbf896fd9" colab={"base_uri": "https://localhost:8080/", "height": 204}
df.head()
# + [markdown] id="G7rLytbrO38L" colab_type="text"
# ## Part 2 - Examine the distribution and relationships of the features
#
# Explore the data - create at least *2* tables (can be summary statistics or crosstabulations) and *2* plots illustrating the nature of the data.
#
# This is open-ended, so to remind - first *complete* this task as a baseline, then go on to the remaining sections, and *then* as time allows revisit and explore further.
#
# Hint - you may need to bin some variables depending on your chosen tables/plots.
# + id="QQvvBd8I2pFL" colab_type="code" colab={}
import numpy as np
# + id="GIdsc0-ey-Yo" colab_type="code" outputId="858647d1-089a-44cb-d81c-b13d31b0e10d" colab={"base_uri": "https://localhost:8080/", "height": 297}
df.describe()
# + id="bdGZFLnNzKXP" colab_type="code" outputId="fdbc2bb9-723c-4d8f-d354-99f767ce1412" colab={"base_uri": "https://localhost:8080/", "height": 171}
pd.crosstab(df['Survival Status'], df['# of +Aux Nodes'])
# + id="EPfyrf5A4ALQ" colab_type="code" outputId="ef976b82-19e9-4792-8afd-dfdfceb151d4" colab={"base_uri": "https://localhost:8080/", "height": 1041}
pd.crosstab(df['# of +Aux Nodes'], df['Survival Status'])
# + id="IAkllgCIFVj0" colab_type="code" colab={}
import matplotlib.pyplot as plt
import numpy as np
# + id="dcTamcbbvwPn" colab_type="code" outputId="6b3e647f-dde8-4c68-b122-628d3e0a7750" colab={"base_uri": "https://localhost:8080/", "height": 300}
y = df['Age']
x = df['Survival Status']
plt.ylabel('AGE')
plt.xlabel('Survival Status')
plt.bar(x,y)
plt.show
# + id="lMMBBz_Qv0yZ" colab_type="code" outputId="36ad4044-f145-4f45-9966-fd0bc5846230" colab={"base_uri": "https://localhost:8080/", "height": 300}
y1 = df['# of +Aux Nodes']
plt.ylabel('# of +Aux Nodes')
plt.xlabel('Age')
plt.plot(y,y1)
plt.show
# + id="6dsfo1-Sw0ot" colab_type="code" outputId="de7e17f1-bb23-444c-95a3-2545f346b3a4" colab={"base_uri": "https://localhost:8080/", "height": 300}
plt.xlabel('# of +Aux Nodes')
plt.ylabel('Survival Status')
plt.bar(y1,x)
plt.show
# + [markdown] id="ZM8JckA2bgnp" colab_type="text"
# ## Part 3 - Analysis and Interpretation
#
# Now that you've looked at the data, answer the following questions:
#
# - What is at least one feature that looks to have a positive relationship with survival?
# - What is at least one feature that looks to have a negative relationship with survival?
# - How are those two features related with each other, and what might that mean?
#
# Answer with text, but feel free to intersperse example code/results or refer to it from earlier.
# + [markdown] id="DUjCb6xAxWq_" colab_type="text"
#
#
# 1. The # of +Aux Nodes when above 25 the chances of survival past 5 years drops sharply.
#
#
# 2. When the number of # of +Aux Nodes is below 25 the chances of Survival increase greatly.
#
# 3. The presence of +Aux Nodes can be a determining factor in whether a surgery is a successful one or not .
#
#
#
#
#
#
#
# + [markdown] id="rShhctzfxfZe" colab_type="text"
#
| DS_Unit_1_Sprint_Challenge_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Notas para contenedor de docker:**
# Comando de docker para ejecución de la nota de forma local:
#
# nota: cambiar `dir_montar` por la ruta de directorio que se desea mapear a `/datos` dentro del contenedor de docker.
#
# ```
# dir_montar=<ruta completa de mi máquina a mi directorio>#aquí colocar la ruta al directorio a montar, por ejemplo:
# #dir_montar=/Users/erick/midirectorio.
# ```
#
# Ejecutar:
#
# ```
# $docker run --rm -v $dir_montar:/datos --name jupyterlab_prope_r_kernel_tidyverse -p 8888:8888 -d palmoreck/jupyterlab_prope_r_kernel_tidyverse:2.1.4
#
# ```
# Ir a `localhost:8888` y escribir el password para jupyterlab: `<PASSWORD>`
#
# Detener el contenedor de docker:
#
# ```
# docker stop jupyterlab_prope_r_kernel_tidyverse
# ```
#
# Documentación de la imagen de docker `palmoreck/jupyterlab_prope_r_kernel_tidyverse:2.1.4` en [liga](https://github.com/palmoreck/dockerfiles/tree/master/jupyterlab/prope_r_kernel_tidyverse).
# ---
# Para ejecución de la nota usar:
#
# [docker](https://www.docker.com/) (instalación de forma **local** con [Get docker](https://docs.docker.com/install/)) y ejecutar comandos que están al inicio de la nota de forma **local**.
#
# O bien dar click en alguno de los botones siguientes:
# [](https://mybinder.org/v2/gh/palmoreck/dockerfiles-for-binder/jupyterlab_prope_r_kernel_tidyerse?urlpath=lab/tree/Propedeutico/Python/clases/1_introduccion/4_modulos_numpy_matplotlib.ipynb) esta opción crea una máquina individual en un servidor de Google, clona el repositorio y permite la ejecución de los notebooks de jupyter.
# [](https://repl.it/languages/python3) esta opción no clona el repositorio, no ejecuta los notebooks de jupyter pero permite ejecución de instrucciones de Python de forma colaborativa con [repl.it](https://repl.it/). Al dar click se crearán nuevos ***repl*** debajo de sus users de ***repl.it***.
#
# # Revisar y ejecutar los ejemplos de las secciones 1.5 a 1.8 del libro de texto "Numerical Methods in Engineering with Python3" de <NAME>
# Temas:
#
# * Módulo numpy.
# * Gráficas con matplotlib.pyplot.
# * Scope de variables.
# * Escritura y ejecución de programas.
#siguiente ejemplo de la sección 1.6 del libro de texto
import matplotlib.pyplot as plt
from numpy import arange,sin,cos
x = arange(0.0,6.2,0.2)
x
plt.plot(x,sin(x),'o-',x,cos(x),'^-')
plt.xlabel('x')
plt.legend(('sine','cosine'),loc = 0)
plt.grid(True)
plt.show()
plt.subplot(2,1,1)
plt.plot(x,sin(x),'o-')
plt.xlabel('x');plt.ylabel('sin(x)')
plt.grid(True)
plt.subplot(2,1,2)
plt.plot(x,cos(x),'^-')
plt.xlabel('x');plt.ylabel('cos(x)')
plt.grid(True)
| Python/clases/1_introduccion/4_modulos_numpy_matplotlib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.0.0
# language: julia
# name: julia-1.0
# ---
using LatPhysBandstructures
using LatticePhysics
using HDF5
using LatPhysBandstructuresPlottingPyPlot
# # Bond Hamiltonians
# +
function saveBondHamiltonian(
hb :: HB,
fn :: AbstractString,
group :: AbstractString = "bond_hamiltonian"
;
append :: Bool = false
) where {L,N,HB<:AbstractBondHamiltonian{L,N}}
# print an error because implementation for concrete type is missing
error("not implemented interface function 'saveBondHamiltonian' for bond Hamiltonian type " * string(typeof(hb)))
end
# convinience function for standard type
function loadBondHamiltonian(
fn :: AbstractString,
group :: AbstractString = "bond_hamiltonian"
)
# find out the type
T = getBondHamiltonianType(Val(Symbol(h5readattr(fn, group)["type"])))
# return the loaded bond hamiltonian
return loadBondHamiltonian(T, fn, group)
end
function getBondHamiltonianType(
::Val{HB}
) where {HB}
error("Type $(HB) could not be identified, i.e. getBondHamiltonianType(Val{:$(HB)}) is missing")
end
# +
function getBondHamiltonianType(
::Val{:BondHoppingHamiltonianSimple}
)
# return the type
return BondHoppingHamiltonianSimple
end
function saveBondHamiltonian(
hb :: HB,
fn :: AbstractString,
group :: AbstractString = "bond_hamiltonian"
;
append :: Bool = false
) where {L,HB<:BondHoppingHamiltonianSimple{L}}
# determine the mode based on if one wants to append stuff
if append
mode = "r+"
else
mode = "w"
end
# open the file in mode
h5open(fn, mode) do file
# create the group in which the bonds are saved
group_hb = g_create(file, group)
# save the type identifier
attrs(group_hb)["type"] = "BondHoppingHamiltonianSimple"
# save the parameters
attrs(group_hb)["N"] = 1
attrs(group_hb)["L"] = string(L)
# save the Float64 coupling
group_hb["coupling"] = hb.coupling
end
# return nothing
return nothing
end
function loadBondHamiltonian(
::Type{HB},
fn :: AbstractString,
group :: AbstractString = "bond_hamiltonian"
) where {LI,HB<:Union{BondHoppingHamiltonianSimple{LI},BondHoppingHamiltonianSimple}}
# read attribute data
attr_data = h5readattr(fn, group)
# determine D based on this
L = Meta.eval(Meta.parse(attr_data["L"]))
N = attr_data["N"]
# load all remaining data
coupling = h5read(fn, group*"/coupling")
# return the new bond hamiltonian
return BondHoppingHamiltonianSimple{L}(coupling)
end
# -
hb = BondHoppingHamiltonianSimple{Int64}(3.0)
saveBondHamiltonian(hb, "test.h5")
hb
loadBondHamiltonian("test.h5")
# +
function getBondHamiltonianType(
::Val{:BondHoppingHamiltonianSimpleNN}
)
# return the type
return BondHoppingHamiltonianSimpleNN
end
function saveBondHamiltonian(
hb :: HB,
fn :: AbstractString,
group :: AbstractString = "bond_hamiltonian"
;
append :: Bool = false
) where {L,HB<:BondHoppingHamiltonianSimpleNN{L}}
# determine the mode based on if one wants to append stuff
if append
mode = "r+"
else
mode = "w"
end
# open the file in mode
h5open(fn, mode) do file
# create the group in which the bonds are saved
group_hb = g_create(file, group)
# save the type identifier
attrs(group_hb)["type"] = "BondHoppingHamiltonianSimpleNN"
# save the parameters
attrs(group_hb)["N"] = 1
attrs(group_hb)["L"] = string(L)
# save the Float64 coupling
group_hb["coupling"] = hb.coupling
if L <: Number
group_hb["label"] = hb.label
else
group_hb["label"] = string(hb.label)
end
end
# return nothing
return nothing
end
function loadBondHamiltonian(
::Type{HB},
fn :: AbstractString,
group :: AbstractString = "bond_hamiltonian"
) where {LI,HB<:Union{BondHoppingHamiltonianSimpleNN{LI},BondHoppingHamiltonianSimpleNN}}
# read attribute data
attr_data = h5readattr(fn, group)
# determine D based on this
L = Meta.eval(Meta.parse(attr_data["L"]))
N = attr_data["N"]
# load coupling
coupling = h5read(fn, group*"/coupling")
# load label
label = L(h5read(fn, group*"/label"))
# return the new bond hamiltonian
return BondHoppingHamiltonianSimpleNN{L}(coupling, label)
end
# -
hb = BondHoppingHamiltonianSimpleNN{String}(3.0, "test")
saveBondHamiltonian(hb, "test.h5")
hb
loadBondHamiltonian("test.h5")
# +
function getBondHamiltonianType(
::Val{:BondHoppingHamiltonianDict}
)
# return the type
return BondHoppingHamiltonianDict
end
function saveBondHamiltonian(
hb :: HB,
fn :: AbstractString,
group :: AbstractString = "bond_hamiltonian"
;
append :: Bool = false
) where {L,HB<:BondHoppingHamiltonianDict{L}}
# determine the mode based on if one wants to append stuff
if append
mode = "r+"
else
mode = "w"
end
# open the file in mode
h5open(fn, mode) do file
# create the group in which the bonds are saved
group_hb = g_create(file, group)
# save the type identifier
attrs(group_hb)["type"] = "BondHoppingHamiltonianDict"
# save the parameters
attrs(group_hb)["N"] = 1
attrs(group_hb)["L"] = string(L)
# save the labels
if L <: Number
group_hb["labels"] = [p[1] for p in hb.couplings]
else
group_hb["labels"] = [string(p[1]) for p in hb.couplings]
end
# save the Float64 couplings
group_hb["couplings"] = [p[2] for p in hb.couplings]
end
# return nothing
return nothing
end
function loadBondHamiltonian(
::Type{HB},
fn :: AbstractString,
group :: AbstractString = "bond_hamiltonian"
) where {LI,HB<:Union{BondHoppingHamiltonianDict{LI},BondHoppingHamiltonianDict}}
# read attribute data
attr_data = h5readattr(fn, group)
# determine D based on this
L = Meta.eval(Meta.parse(attr_data["L"]))
N = attr_data["N"]
# load coupling
couplings = h5read(fn, group*"/couplings")
# load label
labels = L.(h5read(fn, group*"/labels"))
# return the new bond hamiltonian
return BondHoppingHamiltonianDict{L}(Dict([(labels[i], couplings[i]) for i in 1:length(labels)]))
end
# -
hb = BondHoppingHamiltonianDict{String}(Dict("x"=>0.5, "y"=>-0.5))
saveBondHamiltonian(hb, "test.h5")
hb
loadBondHamiltonian("test.h5")
# # Hamiltonian
# +
function saveHamiltonian(
h :: H,
fn :: AbstractString,
group :: AbstractString = "hamiltonian"
;
append :: Bool = false
) where {L,UC<:AbstractUnitcell,N,HB<:AbstractBondHamiltonian{L,N}, H<:Hamiltonian{L,UC,HB}}
# determine the mode based on if one wants to append stuff
if append
mode = "r+"
else
mode = "w"
end
# group for bond hamiltonian
group_hb = group*"/bond_hamiltonian"
# group for unitcell
group_uc = group*"/unitcell"
# open the file in mode
h5open(fn, mode) do file
# create the group in which the bonds are saved
group_h = g_create(file, group)
# save the parameters
attrs(group_h)["N"] = N
attrs(group_h)["L"] = string(L)
# save the groups
attrs(group_h)["bond_hamiltonian"] = group_hb
attrs(group_h)["unitcell"] = group_uc
end
# save bond hamiltonian and unitcell in respective groups (append!)
saveBondHamiltonian(bondHamiltonian(h), fn, group_hb, append=true)
saveUnitcell(unitcell(h), fn, group_uc, append=true)
# return nothing
return nothing
end
function loadHamiltonian(
::Type{H},
fn :: AbstractString,
group :: AbstractString = "hamiltonian"
) where {L,UC<:AbstractUnitcell,N,HB<:AbstractBondHamiltonian{L,N}, H<:Union{Hamiltonian{L,UC,HB}, Hamiltonian}}
# read attribute data
attr_data = h5readattr(fn, group)
# load bond hamiltonian
hb = loadBondHamiltonian(fn, attr_data["bond_hamiltonian"])
# load unitcell
uc = loadUnitcell(fn, attr_data["unitcell"])
# return the new hamiltonian
return Hamiltonian(uc, hb)
end
function loadHamiltonian(
fn :: AbstractString,
group :: AbstractString = "hamiltonian"
)
return loadHamiltonian(Hamiltonian, fn, group)
end
# -
uc = getUnitcellHoneycomb(Symbol, String)
hb = BondHoppingHamiltonianSimpleNN{String}(2.0, "l")
h = Hamiltonian(uc, hb)
saveHamiltonian(h, "test.h5")
h
loadHamiltonian("test.h5")
# # Bandstructure
# +
function saveBandstructure(
bs :: BS,
fn :: AbstractString,
group :: AbstractString = "bandstructure"
;
append :: Bool = false
) where {RP, P<:AbstractReciprocalPath{RP}, L,UC,HB,H<:AbstractHamiltonian{L,UC,HB}, BS<:Bandstructure{P,H}}
# determine the mode based on if one wants to append stuff
if append
mode = "r+"
else
mode = "w"
end
# group for hamiltonian
group_h = group*"/hamiltonian"
# group for path
group_p = group*"/path"
# group for energy bands
group_e = group*"/bands"
# open the file in mode
h5open(fn, mode) do file
# create the group in which the bonds are saved
group_bs = g_create(file, group)
# write number of segments
attrs(group_bs)["segments"] = length(bs.bands)
# save the groups
attrs(group_bs)["hamiltonian"] = group_h
attrs(group_bs)["path"] = group_p
attrs(group_bs)["bands"] = group_e
end
# save hamiltonian
saveHamiltonian(hamiltonian(bs), fn, group_h, append=true)
# save reciprocal path
saveReciprocalPath(path(bs), fn, group_p, append=true)
# save the individual segments within the bands group
h5open(fn, "r+") do file
# create the group in which the bonds are saved
group_bands = g_create(file, group_e)
# save the individual segments
for si in 1:length(bs.bands)
s = bs.bands[si]
# reformat segment and save as matrix
#segmat = [s[j][i] for i in 1:length(s[1]) for j in 1:length(s)]
segmat = zeros(length(s[1]), length(s))
for i in 1:length(s[1])
for j in 1:length(s)
segmat[i,j] = s[j][i]
end
end
# save
group_bands["segment_$(si)"] = segmat
end
end
# return nothing
return nothing
end
function loadBandstructure(
::Type{BS},
fn :: AbstractString,
group :: AbstractString = "bandstructure"
) where {RP, P<:AbstractReciprocalPath{RP}, L,UC,HB,H<:AbstractHamiltonian{L,UC,HB}, BS<:Union{Bandstructure, Bandstructure{P,H}}}
# read attribute data
attr_data = h5readattr(fn, group)
# group for hamiltonian
group_h = attr_data["hamiltonian"]
# group for path
group_p = attr_data["path"]
# group for energy bands
group_e = attr_data["bands"]
# load number of segments
segments = attr_data["segments"]
# load hamiltonian
h = loadHamiltonian(fn, group_h)
# load path
p = loadReciprocalPath(fn, group_p)
# load all energy bands
bds = Vector{Vector{Float64}}[]
# iterate over all expected segments
for s in 1:segments
# load segment as matrix
segmat = h5read(fn, group_e*"/segment_$(s)")
# reformat segment and push to list
segment = Vector{Float64}[
Float64[segmat[i,b] for i in 1:size(segmat,1)]
for b in 1:size(segmat,2)
]
# push to the list
push!(bds, segment)
end
# return the new Bandstructure
bs = getBandstructure(h,p,recalculate=false)
bs.bands = bds
return bs
end
function loadBandstructure(
fn :: AbstractString,
group :: AbstractString = "bandstructure"
)
return loadBandstructure(Bandstructure, fn, group)
end
# -
uc = getUnitcellHoneycomb()
bs = getBandstructure(uc, :Gamma, :K, :M, :Gamma, :Mp);
saveBandstructure(bs, "test.h5")
plotBandstructure(bs);
bsp = loadBandstructure("test.h5")
plotBandstructure(bsp);
| devel/saveload.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Probability | IMDB 5000 Movies | <NAME>
import pandas as pd
import numpy as np
import matplotlib as plt
from datetime import date
df = pd.read_csv('./movie_metadata.csv', error_bad_lines=False)
df = df.drop_duplicates()
df.head()
# # What's the probability that a movie was longer than an hour and a half? Two hours?
more_than_90 = df[df.duration > 90.0]
prob_more_than_90 = more_than_90.count() / df.count()
'{:.2f}%'.format(prob_more_than_90['duration'] * 100)
more_than_120 = df[df.duration > 120.0]
prob_more_than_120 = more_than_120.count() / df.count()
'{:.2f}%'.format(prob_more_than_120['duration'] * 100)
df.duration[(df.duration < 200) & (df.duration > 30)].groupby(df.duration).count().plot()
# # What's the probability that a movie was directed by <NAME>?
director_steve = df[df.director_name == '<NAME>']
prob_director_steve = director_steve.count() / df.count()
'{:.2f}%'.format(prob_director_steve['director_name'] * 100)
# # What's the probability that a movie directed by <NAME> will gross under budget?
director_clint = df[df.director_name == '<NAME>']
gross_clint = director_clint[director_clint.gross < director_clint.budget]
prob_gross_clint = gross_clint.count() / director_clint.count()
'{:.2f}%'.format(prob_gross_clint['gross'] * 100)
# # What's the probability that a movie generally grossed more than its budget?
gross = df[df.gross > df.budget]
prob_gross_df = gross.count() / df.count()
'{:.2f}%'.format(prob_gross_df['gross'] * 100)
# # What's the probability that a movie grossed over the average gross of this data set?
avg_gross = df.gross.mean()
over_gross = df[df.gross > avg_gross].count()
prob_over_gross = over_gross / df.count()
'{:.2f}%'.format(prob_over_gross['gross'] * 100)
# # For ratings we'll consider a movie with at least a 6/10 to be worth renting, if not seeing in theaters. A false positive would be a movie that was highly-rated but did poorly in the box office (gross < budget). A false negative would be a movie that was poorly-rated but did great in the box office (gross > budget).
# # In the IMDB dataset, what are the false positive and false negative rates? Can you provide some examples of each?
rating_high = df[df.imdb_score >= 6.0]
rating_low = df[df.imdb_score < 6.0]
rating_high[rating_high.gross < rating_high.budget].head()
rating_low[rating_low.gross > rating_low.budget].head()
# # If I’m a production studio exec and <NAME> is starring in my movie but I’m feeling uncertain about whether we should keep him (will he make as much money as we want?), tell me should I keep him in the movie or switch him out for <NAME>?
actor_tom = df[(df.actor_1_name == '<NAME>') | (df.actor_2_name == '<NAME>') | (df.actor_3_name == '<NAME>')]
gross_tom = actor_tom[actor_tom.gross > actor_tom.budget]
prob_gross_tom = gross_tom.count() / actor_tom.count()
'{:.2f}%'.format(prob_gross_tom['gross'] * 100)
actor_ford = df[(df.actor_1_name == '<NAME>') | (df.actor_2_name == '<NAME>') | (df.actor_3_name == '<NAME>')]
gross_ford = actor_ford[actor_ford.gross > actor_ford.budget]
prob_gross_ford = gross_ford.count() / actor_ford.count()
'{:.2f}%'.format(prob_gross_ford['gross'] * 100)
# # Same as above, but I’m judging on the ratings of the movie instead of the gross/budget.
actor_tom = df[(df.actor_1_name == '<NAME>') | (df.actor_2_name == '<NAME>') | (df.actor_3_name == '<NAME>')]
actor_tom['imdb_score'].mean()
actor_ford = df[(df.actor_1_name == '<NAME>') | (df.actor_2_name == '<NAME>') | (df.actor_3_name == '<NAME>')]
actor_ford['imdb_score'].mean()
# # What’s the probability that a movie’s length will be between 1hr 10mins and 1h 30mins?
between_70_90 = df[(df.duration < 90.0) & (df.duration > 70.0)]
prob_between_70_90 = between_70_90.count() / df.count()
'{:.2f}%'.format(prob_between_70_90['duration'] * 100)
# # How does the distribution of movie budgets compare to the movie gross values?
df[['gross', 'budget']][df.budget < 1e9].plot(x='budget', y='gross', kind='scatter')
# # Which genre trends toward the highest gross-to-budget ratio? You may have to do some extra parsing to answer this question.
df['gross_budget'] = df['gross'] / df['budget']
genres_table = df[['genres', 'gross_budget']].groupby(df['genres']).mean()
genres_table.sort_values('gross_budget', ascending=False).head()
# +
# Megan did this
genre_gross = df[['genres', 'gross_budget']]
all_genres = list(set('|'.join(df.genres.values).split('|')))
groups = genre_gross.genres.map(lambda cell: tuple(genre in cell for genre in all_genres))
print('presence of specific genres in a movie\n')
groups.head()
# -
genre_gross.index = pd.MultiIndex.from_tuples(groups.values, names=all_genres)
print('convert tuples to indexes')
genre_gross.head() # note: all the indexes are actually filled!
# Create a new data from the new genre data
genre_data = {'gross_budget_mean': [], 'genre': []}
for g in all_genres:
genre_data['gross_budget_mean'].append(genre_gross.xs(True, level=g).gross_budget.mean())
genre_data['genre'].append(g)
pd.DataFrame(genre_data).sort_values('gross_budget_mean', ascending=False).head()
# Thanks Megan
pd.DataFrame(genre_data).sort_values('gross_budget_mean', ascending=False).plot(x='genre', kind='bar')
# # <NAME> is known for starring in some pretty bad movies. Are his movies statistically significantly worse (i.e. in rating) than the rest of the IMDB 5000+?
df['imdb_score'].mean()
actor_nick = df[(df.actor_1_name == '<NAME>') | (df.actor_2_name == '<NAME>') | (df.actor_3_name == '<NAME>')]
actor_nick['imdb_score'].mean()
# # Have any years grossed a statistically-significant higher amount than the other years?
df[['gross', 'title_year']].groupby(df.title_year).sum().plot()
| imdb-notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: solaris
# language: python
# name: solaris
# ---
# # Scoring model performance with the `solaris` python API
#
# This tutorial describes how to run evaluation of a proposal (CSV or .geojson) for a single chip against a ground truth (CSV or .geojson) for the same chip.
#
# ---
# ## CSV Eval
#
# ### Steps
# 1. Imports
# 2. Load ground truth CSV
# 3. Load proposal CSV
# 4. Perform evaluation
#
# ### Imports
#
# For this test case we will use the `eval` submodule within `solaris`.
# imports
import os
import solaris as sol
from solaris.data import data_dir
import pandas as pd # just for visualizing the outputs
# ---
#
#
# ### Load ground truth CSV
#
# We will first instantiate an `Evaluator()` object, which is the core class `eval` uses for comparing predicted labels to ground truth labels. `Evaluator()` takes one argument - the path to the CSV or .geojson ground truth label object. It can alternatively accept a pre-loaded `GeoDataFrame` of ground truth label geometries.
# +
ground_truth_path = os.path.join(data_dir, 'sample_truth.csv')
evaluator = sol.eval.base.Evaluator(ground_truth_path)
evaluator
# -
# At this point, `evaluator` has the following attributes:
#
# - `ground_truth_fname`: the filename corresponding to the ground truth data. This is simply `'GeoDataFrame'` if a GDF was passed during instantiation.
#
# - `ground_truth_GDF`: GeoDataFrame-formatted geometries for the ground truth polygon labels.
#
# - `ground_truth_GDF_Edit`: A deep copy of `eval_object.ground_truth_GDF` which is edited during the process of matching ground truth label polygons to proposals.
#
# - `ground_truth_sindex`: The RTree/libspatialindex spatial index for rapid spatial referencing.
#
# - `proposal_GDF`: An _empty_ GeoDataFrame instantiated to hold proposals later.
#
# ---
#
# ### Load proposal CSV
#
# Next we will load in the proposal CSV file. Note that the `proposalCSV` flag must be set to true for CSV data. If the CSV contains confidence column(s) that indicate confidence in proprosals, the name(s) of the column(s) should be passed as a list of strings with the `conf_field_list` argument; because no such column exists in this case, we will simply pass `conf_field_list=[]`. There are additional arguments available (see [the method documentation](https://cw-eval.readthedocs.io/en/latest/api.html#cw_eval.baseeval.EvalBase.load_proposal)) which can be used for multi-class problems; those will be covered in another recipe. The defaults suffice for single-class problems.
proposals_path = os.path.join(data_dir, 'sample_preds.csv')
evaluator.load_proposal(proposals_path, proposalCSV=True, conf_field_list=[])
# ---
#
#
# ### Perform evaluation
#
# Evaluation iteratively steps through the proposal polygons in `eval_object.proposal_GDF` and determines if any of the polygons in `eval_object.ground_truth_GDF_Edit` have IoU overlap > `miniou` (see [the method documentation](https://cw-eval.readthedocs.io/en/latest/api.html#cw_eval.baseeval.EvalBase.eval_iou)) with that proposed polygon. If one does, that proposal polygon is scored as a true positive. The matched ground truth polygon with the highest IoU (in case multiple had IoU > `miniou`) is removed from `eval_object.ground_truth_GDF_Edit` so it cannot be matched against another proposal. If no ground truth polygon matches with IoU > `miniou`, that proposal polygon is scored as a false positive. After iterating through all proposal polygons, any remaining ground truth polygons in `eval_object.ground_truth_GDF_Edit` are scored as false negatives.
#
# There are several additional arguments to this method related to multi-class evaluation which will be covered in a later recipe. See [the method documentation](https://cw-eval.readthedocs.io/en/latest/api.html#cw_eval.baseeval.EvalBase.eval_iou) for usage.
#
# The prediction outputs a `list` of `dict`s for each class evaluated (only one `dict` in this single-class case). The `dict`(s) have the following keys:
#
# - `'class_id'`: The class being scored in the dict, `'all'` for single-class scoring.
#
# - `'iou_field'`: The name of the column in `eval_object.proposal_GDF` for the IoU score for this class. See [the method documentation](https://cw-eval.readthedocs.io/en/latest/api.html#cw_eval.baseeval.EvalBase.eval_iou) for more information.
#
# - `'TruePos'`: The number of polygons in `eval_object.proposal_GDF` that matched a polygon in `eval_object.ground_truth_GDF_Edit`.
#
# - `'FalsePos'`: The number of polygons in `eval_object.proposal_GDF` that had no match in `eval_object.ground_truth_GDF_Edit`.
#
# - `'FalseNeg'`: The number of polygons in `eval_object.ground_truth_GDF_Edit` that had no match in `eval_object.proposal_GDF`.
#
# - `'Precision'`: The [precision statistic](https://en.wikipedia.org/wiki/Precision_and_recall) for IoU between the proposals and the ground truth polygons.
#
# - `'Recall'`: The [recall statistic](https://en.wikipedia.org/wiki/Precision_and_recall) for IoU between the proposals and the ground truth polygons.
#
# - `'F1Score'`: Also known as the [SpaceNet Metric](https://medium.com/the-downlinq/the-spacenet-metric-612183cc2ddb), the [F<sub>1</sub> score](https://en.wikipedia.org/wiki/F1_score) for IoU between the proposals and the ground truth polygons.
evaluator.eval_iou(calculate_class_scores=False)
# In this case, the score is perfect because the polygons in the ground truth CSV and the proposal CSV are identical. At this point, a new proposal CSV can be loaded (for example, for a new nadir angle at the same chip location) and scoring can be repeated.
# ---
#
# ## GeoJSON Eval
#
# The same operation can be completed with .geojson-formatted ground truth and proposal files. See the example below, and see the detailed explanation above for a description of each step's operations.
# +
ground_truth_geojson = os.path.join(data_dir, 'gt.geojson')
proposal_geojson = os.path.join(data_dir, 'pred.geojson')
evaluator = sol.eval.base.Evaluator(ground_truth_geojson)
evaluator.load_proposal(proposal_geojson, proposalCSV=False, conf_field_list=[])
evaluator.eval_iou(calculate_class_scores=False)
# -
# (Note that the above comes from a different chip location and different proposal than the CSV example, hence the difference in scores)
| docs/tutorials/notebooks/api_evaluation_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: deeprl
# language: python
# name: deeprl
# ---
# +
import torch
from unityagents import UnityEnvironment
import numpy as np
from ddpg_agent import Agent
env = UnityEnvironment(file_name='.\Reacher_Windows_x86_64_20agents\Reacher')
# get the default brain
brain_name = env.brain_names[0]
brain = env.brains[brain_name]
env_info = env.reset(train_mode=False)[brain_name]
# number of agents
num_agents = len(env_info.agents)
print('Number of agents:', num_agents)
# size of each action
action_size = brain.vector_action_space_size
print('Size of each action:', action_size)
# examine the state space
states = env_info.vector_observations
state_size = states.shape[1]
print("Comment lines 91,95-101 in ddpg_agent.py before running this cell")
agent = Agent(state_size=state_size, action_size=action_size,num_agents=num_agents, random_seed=0)
agent.actor_local.load_state_dict(torch.load('checkpoint_actor.pth'))
agent.critic_local.load_state_dict(torch.load('checkpoint_critic.pth'))
states = env_info.vector_observations
for t in range(2000):
actions = agent.act(states, add_noise=False)
actions = np.clip(actions, -1, 1) # all actions between -1 and 1
env_info = env.step(actions)[brain_name] # send all actions to tne environment
next_states = env_info.vector_observations # get next state (for each agent)
rewards = env_info.rewards # get reward (for each agent)
dones = env_info.local_done # see if episode finished
states = next_states # roll over states to next time step
if np.any(dones): # exit loop if episode finished
break
env.close()
# -
| DDPG Continuous Control/Results/watch_agent.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # 1次元有限要素法における2次要素の剛性行列の確認
#
# 1次元有限要素法の2次要素の剛性行列が必要になったためGetfem++で作成しました。
import getfem as gf
import numpy as np
# メッシュ作成の際に、'regular simplices'オプションで次数を2次にします。
m = gf.Mesh('regular simplices', np.arange(0, 2, 1),'degree',2)
print m
# 変位用オブジェクトとデータ用オブジェクトを作成します。
# +
mfu = gf.MeshFem(m, 1)
mfu.set_fem(gf.Fem('FEM_PK(1,2)'))
print mfu
mfd = gf.MeshFem(m, 1)
mfd.set_fem(gf.Fem('FEM_PK(1,2)'))
print mfd
# -
# 積分法は'GAUSS1D'の2次とします。
mim = gf.MeshIm(m, gf.Integ('IM_GAUSS1D(2)'))
print mim
# 剛性行列は以下の様になりました。
Lambda = 1.000
Mu = 1.000
K = gf.asm_linear_elasticity(mim, mfu, mfd, np.repeat([Lambda], mfu.nbdof()), np.repeat([Mu], mfu.nbdof()))
K.full()
# 今回は応力の計算までやってみます。
P = m.pts()
print P
# 1要素で上端に力を加え、下端を固定にします。
cbot = (abs(P[0,:]-np.min(P)) < 1.000e-06)
ctop = (abs(P[0,:]-np.max(P)) < 1.000e-06)
print P[0,:]
print cbot
print ctop
pidbot = np.compress(cbot,range(0,m.nbpts()))
pidtop = np.compress(ctop,range(0,m.nbpts()))
print pidbot
print pidtop
fbot = m.faces_from_pid(pidbot)
ftop = m.faces_from_pid(pidtop)
print fbot
print ftop
BOTTOM = 1
TOP = 2
m.set_region(BOTTOM, fbot)
m.set_region(TOP, ftop)
print m
nbdof = mfd.nbdof()
print nbdof
F = gf.asm_boundary_source(TOP, mim, mfu, mfd, np.repeat([[1.0]], nbdof,1))
print F
(H,R) = gf.asm_dirichlet(BOTTOM, mim, mfu, mfd, mfd.eval('[1]'), mfd.eval('[0]'))
print 'H = ', H.full()
print 'R = ', R
(N, U0) = H.dirichlet_nullspace(R)
print 'N = ', N.full()
print 'U0 = ', U0
Nt = gf.Spmat('copy', N)
Nt.transpose()
print 'Nt = ', Nt.full()
# 準備ができたので連立方程式を解きます。
KK = Nt*K*N
print 'KK = ', KK.full()
FF = Nt*F
print 'FF = ', FF
P = gf.Precond('ildlt',KK)
UU = gf.linsolve_cg(KK,FF,P)
U = N*UU+U0
print U
# gf.compute_gradientで変位の傾きを計算すると、応力$\sigma$は$1$になります。
DU = gf.compute_gradient(mfu,U,mfd)
print 'DU = ', DU
sigma = (Lambda+2.0*Mu)*DU
print 'sigma = ', sigma
| doc/demo_2nddegree_FEM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# We have already seen lists and how they can be used. Now that you have some more background I will go into more detail about lists. First we will look at more ways to get at the elements in a list and then we will talk about copying them.
#
#
# Here are some examples of using indexing to access a single element of an list. Execute the cells below in order:
#
list = ['zero', 'one', 'two', 'three', 'four', 'five']
list[0]
list[4]
list[5]
# You should have gotten the result 'zero', 'four', 'five'.
#
# All those examples should look familiar to you. If you want the first item in the list just look at index 0. The second item is index 1 and so on through the list. However, what if you want the last item in the list? One way could be to use the `len` function like `list[len(list)-1]`. This way works since the `len` function always returns the last index plus one. The second from the last would then be `list[len(list)-2]`. There is an easier way to do this. In Python the last item is always index -1. The second to the last is index -2 and so on. Here are some more examples:
#
list[len(list)-1]
# + active=""
# 'five'
# -
list[len(list)-2]
# + active=""
# 'four'
# -
list[-1]
# + active=""
# 'five'
# -
list[-2]
# + active=""
# 'four'
# -
list[-6]
# + active=""
# 'zero'
# -
#
# Thus any item in the list can be indexed in two ways: from the front and from the back.
#
#
# Another useful way to get into parts of lists is using slices. Here is another example to give you an idea what they can be used for:
#
list = [0, 'Fred', 2, 'S.P.A.M.', 'Stocking', 42, "Jack", "Jill"]
list[0]
# + active=""
# 0
# -
list[7]
# + active=""
# 'Jill'
# -
list[0:8]
# + active=""
# [0, 'Fred', 2, 'S.P.A.M.', 'Stocking', 42, 'Jack', 'Jill']
# -
list[2:4]
# + active=""
# [2, 'S.P.A.M.']
# -
list[4:7]
# + active=""
# ['Stocking', 42, 'Jack']
# -
list[1:5]
# + active=""
# ['Fred', 2, 'S.P.A.M.', 'Stocking']
# -
#
# Slices are used to return part of a list. The slice operator is in the form `list[first_index:following_index]`. The slice goes from the `first_index` to the index before the `following_index`. You can use both types of indexing:
#
list[-4:-2]
# + active=""
# ['Stocking', 42]
# -
list[-4]
# + active=""
# 'Stocking'
# -
list[-4:6]
# + active=""
# ['Stocking', 42]
# -
#
# Another trick with slices is the unspecified index. If the first index is not specified the beginning of the list is assumed. If the following index is not specified the whole rest of the list is assumed. Here are some examples:
#
list[:2]
# + active=""
# [0, 'Fred']
# -
list[-2:]
# + active=""
# ['Jack', 'Jill']
# -
list[:3]
# + active=""
# [0, 'Fred', 2]
# -
list[:-5]
# + active=""
# [0, 'Fred', 2]
# -
#
# Here is a program example:
#
# +
poem = ["<B>", "Jack", "and", "Jill", "</B>", "went", "up", "the", "hill",\
"to", "<B>", "fetch", "a", "pail", "of", "</B>", "water.", "Jack",\
"fell", "<B>", "down", "and", "broke", "</B>", "his", "crown", "and",\
"<B>", "Jill", "came", "</B>", "tumbling", "after"]
def get_bolds(list):
## is_bold tells whether or not the we are currently looking at
## a bold section of text.
is_bold = False
## start_block is the index of the start of either an unbolded
## segment of text or a bolded segment.
start_block = 0
for index in range(len(list)):
##Handle a starting of bold text
if list[index] == "<B>":
if is_bold:
print("Error: Extra Bold")
##print "Not Bold:", list[start_block:index]
is_bold = True
start_block = index+1
##Handle end of bold text
##Remember that the last number in a slice is the index
## after the last index used.
if list[index] == "</B>":
if not is_bold:
print("Error: Extra Close Bold")
print("Bold [", start_block, ":", index, "] ",\
list[start_block:index])
is_bold = False
start_block = index+1
get_bolds(poem)
# -
# with the output being:
#
# + active=""
# Bold [ 1 : 4 ] ['Jack', 'and', 'Jill']
# Bold [ 11 : 15 ] ['fetch', 'a', 'pail', 'of']
# Bold [ 20 : 23 ] ['down', 'and', 'broke']
# Bold [ 28 : 30 ] ['Jill', 'came']
# -
#
#
# The `get_bold` function takes in a list that is broken into words
# and token's. The tokens that it looks for are `<B>` which starts
# the bold text and `<\B>` which ends bold text. The function
# `get_bold` goes through and searches for the start and end
# tokens.
#
#
# The next feature of lists is copying them. If you try something simple like:
#
a = [1, 2, 3]
b = a
print(b)
# + active=""
# [1, 2, 3]
# -
b[1] = 10
print(b)
# + active=""
# [1, 10, 3]
# -
print(a)
# + active=""
# [1, 10, 3]
# -
#
# This probably looks surprising since a modification to b
# resulted in a being changed as well. What happened is that the
# statement `b = a` makes b a *reference* to the same list that a is a reference to.
# This means that b and a are different names for the same list.
# Hence any modification to b changes a as well. However
# some assignments don't create two names for one list:
#
a = [1, 2, 3]
b = a*2
print(a)
# + active=""
# [1, 2, 3]
# -
print(b)
# + active=""
# [1, 2, 3, 1, 2, 3]
# -
a[1] = 10
print(a)
# + active=""
# [1, 10, 3]
# -
print(b)
# + active=""
# [1, 2, 3, 1, 2, 3]
# -
#
#
# In this case, b is not a reference to a since the
# expression `a*2` creates a new list. Then the statement
# `b = a*2` gives b a reference to `a*2` rather than a
# reference to a. All assignment operations create a reference.
# When you pass a list as a argument to a function you create a
# reference as well. Most of the time you don't have to worry about
# creating references rather than copies. However when you need to make
# modifications to one list without changing another name of the list
# you have to make sure that you have actually created a copy.
#
#
# There are several ways to make a copy of a list. The simplest that
# works most of the time is the slice operator since it always makes a
# new list even if it is a slice of a whole list:
#
a = [1, 2, 3]
b = a[:]
b[1] = 10
print(a)
# + active=""
# [1, 2, 3]
# -
print(b)
# + active=""
# [1, 10, 3]
# -
#
#
# Taking the slice [:] creates a new copy of the list. However it
# only copies the outer list. Any sublist inside is still a references
# to the sublist in the original list. Therefore, when the list
# contains lists the inner lists have to be copied as well. You could
# do that manually but Python already contains a module to do it. You
# use the deepcopy function of the copy module:
#
import copy
a = [[1, 2, 3], [4, 5, 6]]
b = a[:]
c = copy.deepcopy(a)
b[0][1] = 10
c[1][1] = 12
print(a)
# + active=""
# [[1, 10, 3], [4, 5, 6]]
# -
print(b)
# + active=""
# [[1, 10, 3], [4, 5, 6]]
# -
print(c)
# + active=""
# [[1, 2, 3], [4, 12, 6]]
# -
#
# First of all, notice that a is an array of arrays. Then notice
# that when `b[0][1] = 10` is run both a and b are
# changed, but c is not. This happens because the inner arrays
# are still references when the slice operator is used. However, with
# deepcopy, c was fully copied.
#
#
# So, should I worry about references every time I use a function or
# `=`? The good news is that you only have to worry about
# references when using dictionaries and lists. Numbers and strings
# create references when assigned but every operation on numbers and
# strings that modifies them creates a new copy so you can never modify
# them unexpectedly. You do have to think about references when you are
# modifying a list or a dictionary.
#
#
# By now you are probably wondering why are references used at all? The
# basic reason is speed. It is much faster to make a reference to a
# thousand element list than to copy all the elements. The other reason
# is that it allows you to have a function to modify the inputed list
# or dictionary. Just remember about references if you ever have some
# weird problem with data being changed when it shouldn't be.
#
| tutorial12.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/amjadraza/learn-ml-with-spark/blob/main/Spark_on_Local_Computers.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="7JCIUZNbhFGw"
# ## **How to use Spark on your local computer**
# + [markdown] id="iAHg3AIxhW_o"
# #### Using Conda
# + [markdown] id="Wjt625gdhecf"
# Conda is an open-source package management and environment management system which is a part of the Anaconda distribution. It is both cross-platform and language agnostic. In practice, Conda can replace both pip and virtualenv. You can download the anaconda packages using the link [here](https://www.anaconda.com/products/individual).
# + [markdown] id="GfmADWOQhg7q"
# 1. Anaconda for windows:
#
# 
#
# 2. For Linux and Mac OS, we have the following options available for downloading anaconda presented at the end of the same page.
#
# 
#
# + [markdown] id="MjLaPRvyhpr5"
# After downloading Anaconda, you have to create a conda environment. Open your terminal and use the command shown below.
#
# 
# + [markdown] id="yQU2QYDQh5LS"
# After the virtual environment is created, it should be visible under the list of Conda environments which can be seen using the following command:
#
# 
#
# + [markdown] id="hkTYxxwliGIE"
# Now activate the newly created environment with the following command:
#
# 
#
# + [markdown] id="tmazFnqviJVl"
# You can install pyspark by Using PyPI to install PySpark in the newly created environment, for example as below. It will install PySpark under the new virtual environment pyspark_env created above.
#
# 
# + [markdown] id="6yH10YB6iOaU"
# Alternatively, you can install PySpark from Conda itself as below:
#
# 
# + [markdown] id="_NXA0GVJiVe1"
# **Manually Downloading PySpark**
#
# PySpark is included in the distributions available at the [Apache Spark website](https://spark.apache.org/downloads.html). You can download a distribution you want from the site. After that, uncompress the tar file into the directory where you want to install Spark, for example, as below:
#
# + id="MDAgCvB7czES" colab={"base_uri": "https://localhost:8080/"} outputId="a9abf381-386c-4ed1-b698-ef9f85b9b01a"
# #!apt-get install openjdk-8-jdk-headless -qq > /dev/null
# !java -version
# + id="EJWXOC5z7c4g"
# !wget -q http://apache.osuosl.org/spark/spark-3.2.0/spark-3.2.0-bin-hadoop3.2.tgz
# + id="CMeH3tin7iTC"
# !tar xf spark-3.2.0-bin-hadoop3.2.tgz
# + [markdown] id="s_2dD3zW7_Jo"
# Ensure the SPARK_HOME environment variable points to the directory where the tar file has been extracted. Update PYTHONPATH environment variable such that it can find the PySpark and Py4J under SPARK_HOME/python/lib. One example of doing this is shown below:
#
# + id="FVoEKyzO7r0q"
import os
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-11-openjdk-amd64"
os.environ["SPARK_HOME"] = "/content/spark-3.2.0-bin-hadoop3.2"
# + id="EBfJzTQZ8Dw8"
| notebooks/week-1/Spark_on_Local_Computers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py3
# language: python
# name: py3
# ---
import numpy as np
from testCases import *
import sklearn
import sklearn.datasets
import sklearn.linear_model
from planar_utils import plot_decision_boundary, sigmoid, load_planar_dataset, load_extra_datasets
# %matplotlib inline
X, Y = load_planar_dataset()
X.shape
plt.scatter(X[0, :], X[1, :], c=Y[0], cmap=plt.cm.Spectral)
plt.show()
# ----
# # 1. Logistic Regression
clf = sklearn.linear_model.LogisticRegressionCV()
clf.fit(X.T, Y.T)
yy.ravel()
np.c_[xx.ravel(), yy.ravel(), yy.ravel()]
# +
x1_min, x1_max = X[0, :].min() - 1, X[0, :].max() + 1
x2_min, x2_max = X[1, :].min() - 1, X[1, :].max() + 1
step = 0.01
mesh_x1 = np.arange(x1_min, x1_max, step)
mesh_x2 = np.arange(x1_min, x2_max, step)
xx1, xx2 = np.meshgrid(mesh_x1, mesh_x2)
plot_x1x2 = np.c_[xx1.ravel(), xx2.ravel()]
Z = clf.predict(plot_x1x2)
Z = Z.reshape(xx1.shape)
plt.contourf(xx1, xx2, Z, cmap=plt.cm.Spectral)
plt.xlabel('x1')
plt.ylabel('x2')
plt.scatter(X[0, :], X[1, :], c=Y[0], cmap=plt.cm.Spectral)
plt.show()
# -
# np.dot(Y[0], pred): 1 = 1 정답
# np.dot(1 - Y[0], 1 - pred): 0 = 0 정답
pred = clf.predict(X.T)
NumOfAns = np.dot(Y[0], pred) + np.dot(1 - Y[0], 1 - pred)
accuracy = NumOfAns / Y[0].size * 100
print("accuracy: ", accuracy, "%")
# # 2. Neural Network Model
# ## Dimension
# - W: (자기 feature =자기 unit) * (앞레이어 feature = 앞레이어 unit)
# ### 1) set structure
def layer_size(X, Y):
# n_h: Num of Unit of HiddenLayer
n_x = X.shape[0]
n_h = 4
n_y = Y.shape[0]
print("num of feature: {}, num of unit of hidden Layer: {}, out put: {}".\
format(n_x, n_h, n_y))
return (n_x, n_h, n_y)
n_x, n_h, n_y = layer_size(X, Y)
# ### 2) initialize params
def initialize_params(n_x, n_h, n_y):
np.random.seed(2)
W1 = np.random.randn(n_h, n_x) * 0.01
b1 = np.zeros(shape=(n_h, 1))
W2 = np.random.randn(n_y, n_h) * 0.01
b2 = np.zeros(shape=(n_y, 1))
assert (W1.shape == (n_h, n_x))
assert (b1.shape == (n_h, 1))
assert (W2.shape == (n_y, n_h))
assert (b2.shape == (n_y, 1))
params = {"W1": W1,
"b1": b1,
"W2": W2,
"b2": b2}
return params
params = initialize_params(n_x, n_h, n_y)
params
# ### 3) loop: forward propagation
def forward_propagation(X, params):
W1 = params['W1']
b1 = params['b1']
W2 = params['W2']
b2 = params['b2']
Z1 = np.dot(W1, X) + b1
A1 = np.tanh(Z1)
Z2 = np.dot(W2, A1) + b2
A2 = sigmoid(Z2)
assert (A2.shape == (1, X.shape[1]))
cache = {"Z1": Z1,
"A1": A1,
"Z2": Z2,
"A2": A2}
return A2, cache
A2, cache = forward_propagation(X, params)
A2.shape
# ### 4) Cost
# $$J = - \frac{1}{m} \sum\limits_{i = 0}^{m} \large{(} \small y^{(i)}\log\left(a^{[2] (i)}\right) + (1-y^{(i)})\log\left(1- a^{[2] (i)}\right) \large{)} $$
#
def compute_cost(A2, Y):
m = Y.shape[1] # Num of sample
logprob = np.multiply(np.log(A2), Y) + np.multiply((1- Y), np.log(1 - A2))
loss = - logprob
cost = 1/m * np.sum(loss)
cost = np.squeeze(cost) # makes sure cost is the dimension we expect.
# E.g., turns [[17]] into 17
assert(isinstance(cost, float))
return cost
compute_cost(A2, Y)
# ### 5) loop: backward propagation
def backward_propagation(params, cache, X, Y):
m = X.shape[1]
W1 = params['W1']
W2 = params['W2']
A1 = cache['A1']
A2 = cache['A2']
d_Z2 = A2 - Y
d_W2 = (1 / m) * np.dot(d_Z2, A1.T)
d_b2 = (1 / m) * np.sum(d_Z2)
d_Z1 = np.multiply(np.dot(W2.T, d_Z2), 1 - np.power(A1, 2))
d_W1 = (1 / m) * np.dot(d_Z1, X.T)
d_b1 = (1 / m) * np.sum(d_Z1, axis=1, keepdims=True)
grads = {"d_W1": d_W1,
"d_b1": d_b1,
"d_W2": d_W2,
"d_b2": d_b2,}
return grads
grads = backward_propagation(params, cache, X, Y)
grads
# ## 6) update_params
def update_params(params, grads, learning_rate=1.2):
# before
W1 = params['W1']
b1 = params['b1']
W2 = params['W2']
b2 = params['b2']
d_W2 = grads['d_W2']
d_b2 = grads['d_b2']
d_W1 = grads['d_W1']
d_b1 = grads['d_b1']
# after
W2 = W2 - learning_rate * d_W2
b2 = b2 - learning_rate * d_b2
W1 = W1 - learning_rate * d_W1
b1 = b1 - learning_rate * d_b1
params = {"W1": W1,
"b1": b1,
"W2": W2,
"b2": b2}
return params
update_params(params, grads)
# ## 7) Build Model
# - forward propagation: get $Z, \sigma(Z), \text{ and cost}$
# - backward propagation: get **dW, db** and update
def nn_model(X, Y, n_h, num_iteration=10000, print_cost=False):
np.random.seed(3)
n_x = layer_size(X, Y)[0]
n_y = layer_size(X, Y)[2]
params = initialize_params(n_x, n_h, n_y)
W1 = params['W1']
b1 = params['b1']
W2 = params['W2']
b2 = params['b2']
# gradient descent
for i in range(0, num_iteration):
A2, cache = forward_propagation(X, params)
cost = compute_cost(A2, Y)
grads = backward_propagation(params, cache, X, Y)
params = update_params(params, grads)
if print_cost and i % 1000 == 0:
print("Cost after iteration {}: {}".format(i, cost))
return params
nn_model(X, Y, 4, print_cost=True)
# ## 8) prediction
# $y_{prediction} = \mathbb 1 \text{{activation > 0.5}} = \begin{cases} 1 & \text{if}\ activation > 0.5 \\ 0 & \text{otherwise} \end{cases}$
def predict(params, X):
A2, cache = forward_propagation(X, params)
pred = np.round(A2)
return pred
predict(params, X)
# ## eg.
X.shape
# +
# Build model
params = nn_model(X, Y, n_h=4, num_iteration=10000, print_cost=True)
# Plotting
x1_min, x1_max = X[0, :].min() - 1, X[0, :].max() + 1
x2_min, x2_max = X[1, :].min() - 1, X[1, :].max() + 1
step = 0.01
mesh_x1 = np.arange(x1_min, x1_max, step)
mesh_x2 = np.arange(x1_min, x2_max, step)
xx1, xx2 = np.meshgrid(mesh_x1, mesh_x2)
plot_x1x2 = np.c_[xx1.ravel(), xx2.ravel()]
Z = predict(params, plot_x1x2.T)
Z = Z.reshape(xx1.shape)
plt.contourf(xx1, xx2, Z, cmap=plt.cm.Spectral)
plt.xlabel('x1')
plt.ylabel('x2')
plt.scatter(X[0, :], X[1, :], c=Y[0], cmap=plt.cm.Spectral)
plt.show()
# +
plt.figure(figsize=(16, 32))
hidden_layer_depth = [1, 2, 3, 4, 5, 40 ,50]
for i, n_h in enumerate(hidden_layer_depth):
plt.subplot(5, 2, i+1)
plt.title("Hidden layer depth: {}".format(n_h))
params = nn_model(X, Y, n_h, num_iteration=5000)
plot_decision_boundary(lambda x: predict(params, x.T), X, Y)
pred = predict(params, X)
ans = np.dot(Y, pred.T) + np.dot(1 - Y, 1 - pred.T)
accuracy = float(ans) / float(Y.size) * 100
print("Accuracy for {} hidden units: {} %".format(n_h, accuracy))
# -
# - 질문
# - prediction에서 왜 Transpose?
# reference
# - https://github.com/Kulbear/deep-learning-coursera
| 07.DeepLearning/01_MLandDeepLearning_BuildingDeepNeuralNetwork.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ADDRCODE Mapping for Dengue Case Data
# +
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
import os
import pandas as pd
from collections import Counter
# +
df_DHF = pd.read_csv(os.path.join('..','..','data','dengue-cases','2016.csv'))
df_DHF = df_DHF.set_index('ID')
df_DHF = df_DHF.loc[df_DHF['PROVINCE'] == 80]
df_DHF = df_DHF.astype(str)
df_DHF['CODE'] = df_DHF['PROVINCE'] + df_DHF['ADDRCODE']
df_DHF['CODE'] = df_DHF['CODE'].str[:-2]
df_DHF = pd.DataFrame.from_records(Counter(df_DHF['CODE'].values).most_common())
df_DHF.columns = ['Subdistrict_Code', 'Cases']
df_DHF = df_DHF.set_index('Subdistrict_Code')
df_DHF.head()
len(df_DHF)
# -
df_code = pd.read_csv(os.path.join('..','..','data','dengue-cases','province_dist_sub_code.csv'))
df_code['Subdistrict_Code'] = df_code['Subdistrict_Code'].astype(str)
df_code = df_code.set_index('Subdistrict_Code')
df_code = df_code.loc[df_code['Province_Code'] == 80]
df_code.head()
len(df_code)
df_dengue_caces = df_code.join(df_DHF)
df_dengue_caces = df_dengue_caces.fillna(0)
df_dengue_caces = df_dengue_caces.dropna(how='any', axis=0)
df_dengue_caces = df_dengue_caces.reset_index()
df_dengue_caces = df_dengue_caces.drop(['Subdistrict_Code', 'District_Code', 'Province_Code'], axis=1)
df_dengue_caces.columns = ['subdist', 'district', 'province', 'cases']
df_dengue_caces.head()
df_dengue_caces.to_csv(os.path.join('..','..','data','dengue-cases','dengue_caces_2016.csv'), index=None)
| src/preprocess/addrcode-mapping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/skywalker0803r/c620/blob/main/notebook/Integration_and_Test.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="YEiUzu5_2x3I"
import joblib
import os
import numpy as np
import pandas as pd
pd.options.display.max_rows = 9999
# !pip install autorch > log.txt
import matplotlib.pyplot as plt
import autorch
from autorch.function import sp2wt
import random
random.seed(11)
np.random.seed(11)
# + colab={"base_uri": "https://localhost:8080/"} id="3YPjXHkR3Eso" outputId="d789e8c0-cae0-4e43-a094-46e85489b81a"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="nuzZj4KB6mQB"
# # columns name
# + colab={"base_uri": "https://localhost:8080/"} id="vD1E0WtI3OKC" outputId="7632cdb3-6566-4b0b-ded9-c229b265a6b2"
icg_c = joblib.load('/content/drive/MyDrive/台塑輕油案子/data/c620/col_names/icg_col_names.pkl')
c620_c = joblib.load('/content/drive/MyDrive/台塑輕油案子/data/c620/col_names/c620_col_names.pkl')
c660_c = joblib.load('/content/drive/MyDrive/台塑輕油案子/data/c620/col_names/c660_col_names.pkl')
t651_c = joblib.load('/content/drive/MyDrive/台塑輕油案子/data/c620/col_names/t651_col_names.pkl')
c670_c = joblib.load('/content/drive/MyDrive/台塑輕油案子/data/c620/col_names/c670_col_names.pkl')
print(icg_c.keys())
print(c620_c.keys())
print(c660_c.keys())
print(c670_c.keys())
print(t651_c.keys())
# + [markdown] id="x8BgCUXhB_Qe"
# # DataFrame
# + id="TjsU3C9bB_er" colab={"base_uri": "https://localhost:8080/"} outputId="920a146f-f92d-41ce-9d18-b51af95b5d58"
icg_df = pd.read_csv('/content/drive/MyDrive/台塑輕油案子/data/c620/cleaned/icg_train.csv',index_col=0)
c620_df = pd.read_csv('/content/drive/MyDrive/台塑輕油案子/data/c620/cleaned/c620_train.csv',index_col=0)
c660_df = pd.read_csv('/content/drive/MyDrive/台塑輕油案子/data/c620/cleaned/c660_train.csv',index_col=0)
c670_df = pd.read_csv('/content/drive/MyDrive/台塑輕油案子/data/c620/cleaned/c670_train.csv',index_col=0)
t651_df = pd.read_csv('/content/drive/MyDrive/台塑輕油案子/data/c620/cleaned/t651_train.csv',index_col=0)
idx = list(set(icg_df.index)&
set(c620_df.index)&
set(c660_df.index)&
set(c670_df.index)&
set(t651_df.index))
len(idx)
# + id="kxeTF9Z5LYQV" colab={"base_uri": "https://localhost:8080/", "height": 377} outputId="fcf9360e-bec9-4039-e408-b7afee18b949"
icg_df.loc[idx].head()
# + id="2KP7j-9PLUel" colab={"base_uri": "https://localhost:8080/", "height": 306} outputId="a8dc9030-679b-4a9c-ab74-207def7fc206"
c620_df.loc[idx][c620_c['case']].head()
# + id="xuehZMotRCYO" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="5b575359-315c-4df2-c4e7-2958b48cc343"
c660_df.loc[idx][c660_c['case']].head()
# + [markdown] id="eT4-lodHox-u"
# # Input data
# + id="3EOUCMOXB2Y1"
# icg
icg_input = icg_df.loc[idx,icg_c['x']]
icg_input = icg_input.join(c620_df.loc[idx,'Tatoray Stripper C620 Operation_Specifications_Spec 2 : Distillate Rate_m3/hr'])
icg_input = icg_input.join(c660_df.loc[idx,'Benzene Column C660 Operation_Specifications_Spec 3 : Toluene in Benzene_ppmw'])
icg_input = icg_input.join(c620_df.loc[idx].filter(regex='Receiver Temp'))
# c620
c620_feed = c620_df.loc[idx,c620_c['x41']]
# t651
t651_feed = t651_df.loc[idx,t651_c['x41']]
# + id="hTyJ6JVm3Xzv" colab={"base_uri": "https://localhost:8080/", "height": 377} outputId="7ba79811-bc65-4171-86ad-79cf23bee0aa"
icg_input.head()
# + [markdown] id="mp1GqvJ0o0k1"
# # Output data
# + id="uHCoq80jo0ss"
c620_op = c620_df.loc[idx,c620_c['density']+c620_c['yRefluxRate']+c620_c['yHeatDuty']+c620_c['yControl']]
c620_wt = c620_df.loc[idx,c620_c['vent_gas_x']+c620_c['distillate_x']+c620_c['sidedraw_x']+c620_c['bottoms_x']]
c660_op = c660_df.loc[idx,c660_c['density']+c660_c['yRefluxRate']+c660_c['yHeatDuty']+c660_c['yControl']]
c660_wt = c660_df.loc[idx,c660_c['vent_gas_x']+c660_c['distillate_x']+c660_c['sidedraw_x']+c660_c['bottoms_x']]
c670_op = c670_df.loc[idx,c670_c['density']+c670_c['yRefluxRate']+c670_c['yHeatDuty']+c670_c['yControl']]
c670_wt = c670_df.loc[idx,c670_c['distillate_x']+c670_c['bottoms_x']]
# + [markdown] id="h8VBB0aG-e0c"
# # config
# + id="0Ktejan2r_U7"
config = {
# simple op_col
'c620_simple_op_col':'/content/drive/MyDrive/台塑輕油案子/data/c620/map_dict/c620_simple_op_col.pkl',
'c660_simple_op_col':'/content/drive/MyDrive/台塑輕油案子/data/c620/map_dict/c660_simple_op_col.pkl',
'c670_simple_op_col':'/content/drive/MyDrive/台塑輕油案子/data/c620/map_dict/c670_simple_op_col.pkl',
# model paht
'icg_model_path':'/content/drive/MyDrive/台塑輕油案子/data/c620/model/c620_icg_svr.pkl',
'c620_model_path':'/content/drive/MyDrive/台塑輕油案子/data/c620/model/c620.pkl',
'c660_model_path':'/content/drive/MyDrive/台塑輕油案子/data/c620/model/c660.pkl',
'c670_model_path':'/content/drive/MyDrive/台塑輕油案子/data/c620/model/c670.pkl',
# real data model path
'icg_model_path_real_data':'/content/drive/MyDrive/台塑輕油案子/data/c620/model/c620_icg_svr_real_data.pkl',
'c620_model_path_real_data':'/content/drive/MyDrive/台塑輕油案子/data/c620/model/c620_real_data.pkl',
'c660_model_path_real_data':'/content/drive/MyDrive/台塑輕油案子/data/c620/model/c660_real_data.pkl',
'c670_model_path_real_data':'/content/drive/MyDrive/台塑輕油案子/data/c620/model/c670_real_data.pkl',
# real data linear model path
'c620_model_path_real_data_linear':'/content/drive/MyDrive/台塑輕油案子/data/c620/model/c620_real_data_lassocv.pkl',
'c660_model_path_real_data_linear':'/content/drive/MyDrive/台塑輕油案子/data/c620/model/c660_real_data_lassocv.pkl',
'c670_model_path_real_data_linear':'/content/drive/MyDrive/台塑輕油案子/data/c620/model/c670_real_data_lassocv.pkl',
# col_names
'icg_col_path':'/content/drive/MyDrive/台塑輕油案子/data/c620/col_names/icg_col_names.pkl',
'c620_col_path':'/content/drive/MyDrive/台塑輕油案子/data/c620/col_names/c620_col_names.pkl',
'c660_col_path':'/content/drive/MyDrive/台塑輕油案子/data/c620/col_names/c660_col_names.pkl',
'c670_col_path':'/content/drive/MyDrive/台塑輕油案子/data/c620/col_names/c670_col_names.pkl',
# Special column (0.9999 & 0.0001)
'index_9999_path':'/content/drive/MyDrive/台塑輕油案子/data/c620/cleaned/index_9999.pkl',
'index_0001_path':'/content/drive/MyDrive/台塑輕油案子/data/c620/cleaned/index_0001.pkl',
# sp
'c620_wt_always_same_split_factor_dict':'/content/drive/MyDrive/台塑輕油案子/data/c620/map_dict/c620_wt_always_same_split_factor_dict.pkl',
'c660_wt_always_same_split_factor_dict':'/content/drive/MyDrive/台塑輕油案子/data/c620/map_dict/c660_wt_always_same_split_factor_dict.pkl',
'c670_wt_always_same_split_factor_dict':'/content/drive/MyDrive/台塑輕油案子/data/c620/map_dict/c670_wt_always_same_split_factor_dict.pkl',
}
# + [markdown] id="MoSO4Cbx6oxP"
# # define F
# + id="oiu5SRYL5NOr"
import joblib
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import autorch
from autorch.function import sp2wt
class F(object):
def __init__(self,config):
# simulation data model
self.icg_model = joblib.load(config['icg_model_path'])
self.c620_model = joblib.load(config['c620_model_path'])
self.c660_model = joblib.load(config['c660_model_path'])
self.c670_model = joblib.load(config['c670_model_path'])
# real data model
self.icg_real_data_model = joblib.load(config['icg_model_path_real_data'])
self.c620_real_data_model = joblib.load(config['c620_model_path_real_data'])
self.c660_real_data_model = joblib.load(config['c660_model_path_real_data'])
self.c670_real_data_model = joblib.load(config['c670_model_path_real_data'])
# real data linear model
self.c620_real_data_model_linear = joblib.load(config['c620_model_path_real_data_linear'])
self.c660_real_data_model_linear = joblib.load(config['c660_model_path_real_data_linear'])
self.c670_real_data_model_linear = joblib.load(config['c670_model_path_real_data_linear'])
# columns name
self.icg_col = joblib.load(config['icg_col_path'])
self.c620_col = joblib.load(config['c620_col_path'])
self.c660_col = joblib.load(config['c660_col_path'])
self.c670_col = joblib.load(config['c670_col_path'])
# simple op_col
self.c620_simple_op_col = joblib.load(config['c620_simple_op_col'])
self.c660_simple_op_col = joblib.load(config['c660_simple_op_col'])
self.c670_simple_op_col = joblib.load(config['c670_simple_op_col'])
# other infomation
self.c620_wt_always_same_split_factor_dict = joblib.load(config['c620_wt_always_same_split_factor_dict'])
self.c660_wt_always_same_split_factor_dict = joblib.load(config['c660_wt_always_same_split_factor_dict'])
self.c670_wt_always_same_split_factor_dict = joblib.load(config['c670_wt_always_same_split_factor_dict'])
self.index_9999 = joblib.load(config['index_9999_path'])
self.index_0001 = joblib.load(config['index_0001_path'])
self.V615_density = 0.8626
self.C820_density = 0.8731
self.T651_density = 0.8749
# user can set two mode
self.Recommended_mode = False
self.real_data_mode = False
self._Post_processing = True
self._linear_model = False
def ICG_loop(self,Input):
while True:
if self.real_data_mode == True:
output = pd.DataFrame(self.icg_real_data_model.predict(Input[self.icg_col['x']].values),
index=Input.index,columns=['Simulation Case Conditions_C620 Distillate Rate_m3/hr'])
if self.real_data_mode == False:
output = pd.DataFrame(self.icg_model.predict(Input[self.icg_col['x']].values),
index=Input.index,columns=['Simulation Case Conditions_C620 Distillate Rate_m3/hr'])
dist_rate = output['Simulation Case Conditions_C620 Distillate Rate_m3/hr'].values[0]
na_in_benzene = Input['Simulation Case Conditions_Spec 2 : NA in Benzene_ppmw'].values[0]
print('current Distillate Rate_m3/hr:{} NA in Benzene_ppmw:{}'.format(dist_rate,na_in_benzene))
if output['Simulation Case Conditions_C620 Distillate Rate_m3/hr'].values[0] > 0:
return output,Input
else:
Input['Simulation Case Conditions_Spec 2 : NA in Benzene_ppmw'] -= 30
print('NA in Benzene_ppmw -= 30')
def __call__(self,icg_input,c620_feed,t651_feed):
# get index
idx = icg_input.index
# c620_case
c620_case = pd.DataFrame(index=idx,columns=self.c620_col['case'])
# c620_case(Receiver Temp_oC) = user input
c620_case['Tatoray Stripper C620 Operation_Specifications_Spec 1 : Receiver Temp_oC'] = icg_input['Tatoray Stripper C620 Operation_Specifications_Spec 1 : Receiver Temp_oC'].values
if self.Recommended_mode == True:
icg_input['Simulation Case Conditions_Spec 2 : NA in Benzene_ppmw'] = 980.0
icg_input['Simulation Case Conditions_Spec 1 : Benzene in C620 Sidedraw_wt%'] = 70.0
icg_output,icg_input = self.ICG_loop(icg_input)
c620_case['Tatoray Stripper C620 Operation_Specifications_Spec 2 : Distillate Rate_m3/hr'] = icg_output.values
c620_case['Tatoray Stripper C620 Operation_Specifications_Spec 3 : Benzene in Sidedraw_wt%'] = icg_input['Simulation Case Conditions_Spec 1 : Benzene in C620 Sidedraw_wt%'].values
if self.Recommended_mode == False:
c620_case['Tatoray Stripper C620 Operation_Specifications_Spec 2 : Distillate Rate_m3/hr'] = icg_input['Tatoray Stripper C620 Operation_Specifications_Spec 2 : Distillate Rate_m3/hr'].values
c620_case['Tatoray Stripper C620 Operation_Specifications_Spec 3 : Benzene in Sidedraw_wt%'] = icg_input['Simulation Case Conditions_Spec 1 : Benzene in C620 Sidedraw_wt%'].values
# c620_input(c620_case&c620_feed)
c620_input = c620_case.join(c620_feed)
# c620 output(op&wt)
c620_input = c620_case.join(c620_feed)
c620_output = self.c620_model.predict(c620_input)
c620_sp,c620_op = c620_output.iloc[:,:41*4],c620_output.iloc[:,41*4:]
# update by c620 real data model?
if self.real_data_mode == True:
if self._linear_model == True:
c620_op_real = self.c620_real_data_model_linear.predict(c620_input)[:,41*4:]
c620_op_real = pd.DataFrame(c620_op_real,index=c620_input.index,columns=self.c620_simple_op_col)
c620_sp_real = self.c620_real_data_model_linear.predict(c620_input)[:,:41*4]
c620_sp_real = pd.DataFrame(c620_sp_real,index=c620_input.index,columns=c620_sp.columns)
if self._linear_model == False:
c620_op_real = self.c620_real_data_model.predict(c620_input).iloc[:,41*4:]
c620_sp_real = self.c620_real_data_model.predict(c620_input).iloc[:,:41*4]
c620_op.update(c620_op_real)
c620_sp.update(c620_sp_real)
# c620 sp後處理
if self._Post_processing:
for i in self.c620_wt_always_same_split_factor_dict.keys():
c620_sp[i] = self.c620_wt_always_same_split_factor_dict[i]
# 計算 c620_wt
s1,s2,s3,s4 = c620_sp.iloc[:,:41].values,c620_sp.iloc[:,41:41*2].values,c620_sp.iloc[:,41*2:41*3].values,c620_sp.iloc[:,41*3:41*4].values
w1,w2,w3,w4 = sp2wt(c620_feed,s1),sp2wt(c620_feed,s2),sp2wt(c620_feed,s3),sp2wt(c620_feed,s4)
wt = np.hstack((w1,w2,w3,w4))
c620_wt = pd.DataFrame(wt,index=idx,columns=self.c620_col['vent_gas_x']+self.c620_col['distillate_x']+self.c620_col['sidedraw_x']+self.c620_col['bottoms_x'])
# 如果是線性模式就再update c620 wt一次,放在後處理前
if self._linear_model:
c620_wt_real = self.c620_real_data_model_linear.predict(c620_input)[:,:41*4]
c620_wt_real = pd.DataFrame(c620_wt_real,index=c620_input.index,columns=c620_wt.columns)
c620_wt.update(c620_wt_real)
# c620_wt 後處理 為了在輸出之前滿足業主給的約束條件
if self._Post_processing:
bz_idx = c620_wt.columns.tolist().index('Tatoray Stripper C620 Operation_Sidedraw Production Rate and Composition_Benzene_wt%')
other_idx = [i for i in range(41*2,41*3,1) if i != bz_idx]
other_total = (100 - c620_input['Tatoray Stripper C620 Operation_Specifications_Spec 3 : Benzene in Sidedraw_wt%'].values).reshape(-1,1)
c620_wt.iloc[:,bz_idx] = c620_input['Tatoray Stripper C620 Operation_Specifications_Spec 3 : Benzene in Sidedraw_wt%'].values
c620_wt.iloc[:,other_idx] = (c620_wt.iloc[:,other_idx].values /
c620_wt.iloc[:,other_idx].values.sum(axis=1).reshape(-1,1))*other_total
# c620 input mass flow rate m3 to ton
V615_Btm_m3 = icg_input['Simulation Case Conditions_Feed Rate_Feed from V615 Btm_m3/hr'].values.reshape(-1,1)
C820_Dist_m3 = icg_input['Simulation Case Conditions_Feed Rate_Feed from C820 Dist_m3/hr'].values.reshape(-1,1)
V615_Btm_ton = V615_Btm_m3*self.V615_density
C820_Dist_ton = C820_Dist_m3*self.C820_density
c620_feed_rate_ton = V615_Btm_ton+C820_Dist_ton
# c620 output mass flow ton
c620_mf_side = np.sum(c620_feed_rate_ton*c620_feed.values*s3*0.01,axis=1,keepdims=True)
c620_mf_bot = np.sum(c620_feed_rate_ton*c620_feed.values*s4*0.01,axis=1,keepdims=True)
# t651 feed mass flow rate(ton)
t651_mf = (icg_input['Simulation Case Conditions_Feed Rate_Feed from T651_m3/hr']*self.T651_density).values.reshape(-1,1)
# c660 input mass flow(ton)
c660_mf = t651_mf + c620_mf_side
t651_mf_p ,c620_mf_side_p = t651_mf/c660_mf ,c620_mf_side/c660_mf
# c660 input(feed & case)
c660_feed = c620_wt[self.c620_col['sidedraw_x']].values*c620_mf_side_p + t651_feed.values*t651_mf_p
c660_feed = pd.DataFrame(c660_feed,index=idx,columns=self.c660_col['x41'])
c660_case = pd.DataFrame(index=idx,columns=self.c660_col['case'])
c660_case['Benzene Column C660 Operation_Specifications_Spec 2 : NA in Benzene_ppmw'] = icg_input['Simulation Case Conditions_Spec 2 : NA in Benzene_ppmw'].values
if self.Recommended_mode == True:
# fix Toluene in Benzene_ppmw = 10
c660_case['Benzene Column C660 Operation_Specifications_Spec 3 : Toluene in Benzene_ppmw'] = 10.0
if self.Recommended_mode == False:
# Toluene in Benzene_ppmw = user input
c660_case['Benzene Column C660 Operation_Specifications_Spec 3 : Toluene in Benzene_ppmw'] = icg_input['Benzene Column C660 Operation_Specifications_Spec 3 : Toluene in Benzene_ppmw'].values
c660_input = c660_case.join(c660_feed)
# c660 output(op&wt)
c660_output = self.c660_model.predict(c660_input)
c660_sp,c660_op = c660_output.iloc[:,:41*4],c660_output.iloc[:,41*4:]
# update by c660 real data model?
if self.real_data_mode == True:
if self._linear_model == True:
c660_op_real = self.c660_real_data_model_linear.predict(c660_input)[:,41*4:]
c660_op_real = pd.DataFrame(c660_op_real,index=c660_input.index,columns=self.c660_simple_op_col)
c660_sp_real = self.c660_real_data_model_linear.predict(c660_input)[:,:41*4]
c660_sp_real = pd.DataFrame(c660_sp_real,index=c660_input.index,columns=c660_sp)
if self._linear_model == False:
c660_op_real = self.c660_real_data_model.predict(c660_input).iloc[:,41*4:] #操作條件放後面
c660_sp_real = self.c660_real_data_model.predict(c660_input).iloc[:,:41*4] #分離係數放前面
c660_op.update(c660_op_real)
c660_sp.update(c660_sp_real)
# c660 sp後處理
if self._Post_processing:
for i in self.c660_wt_always_same_split_factor_dict.keys():
c660_sp[i] = self.c660_wt_always_same_split_factor_dict[i]
# 計算 c660_wt
s1,s2,s3,s4 = c660_sp.iloc[:,:41].values,c660_sp.iloc[:,41:41*2].values,c660_sp.iloc[:,41*2:41*3].values,c660_sp.iloc[:,41*3:41*4].values
w1,w2,w3,w4 = sp2wt(c660_feed,s1),sp2wt(c660_feed,s2),sp2wt(c660_feed,s3),sp2wt(c660_feed,s4)
wt = np.hstack((w1,w2,w3,w4))
c660_wt = pd.DataFrame(wt,index=idx,columns=self.c660_col['vent_gas_x']+self.c660_col['distillate_x']+self.c660_col['sidedraw_x']+self.c660_col['bottoms_x'])
# 如果是線性模式就再update c660 wt一次,放在後處理前
if self._linear_model:
c660_wt_real = self.c660_real_data_model_linear.predict(c660_input)[:,:41*4]
c660_wt_real = pd.DataFrame(c660_wt_real,index=c660_input.index,columns=c660_wt.columns)
c660_wt.update(c660_wt_real)
# c660_wt 後處理 為了在輸出之前滿足業主給的約束條件
if self._Post_processing:
na_idx = [1,2,3,4,5,6,8,9,11,13,14,15,20,22,29]
other_idx = list(set([*range(41)])-set(na_idx))
na_total = (c660_input['Benzene Column C660 Operation_Specifications_Spec 2 : NA in Benzene_ppmw'].values/10000).reshape(-1,1)
other_total = 100 - na_total
c660_wt.iloc[:,41*2:41*3].iloc[:,na_idx] = (c660_wt.iloc[:,41*2:41*3].iloc[:,na_idx].values/
c660_wt.iloc[:,41*2:41*3].iloc[:,na_idx].values.sum(axis=1).reshape(-1,1))*na_total
c660_wt.iloc[:,41*2:41*3].iloc[:,other_idx] = (c660_wt.iloc[:,41*2:41*3].iloc[:,other_idx].values/
c660_wt.iloc[:,41*2:41*3].iloc[:,other_idx].values.sum(axis=1).reshape(-1,1))*other_total
# c660 output mass flow (ton)
c660_mf_bot = np.sum(c660_mf*c660_feed.values*s4*0.01,axis=1,keepdims=True)
# c670 input mass flow
c670_mf = c620_mf_bot + c660_mf_bot
c620_mf_bot_p,c660_mf_bot_p = c620_mf_bot/c670_mf , c660_mf_bot/c670_mf
# c670 feed wt%
c670_feed = c620_wt[self.c620_col['bottoms_x']].values*c620_mf_bot_p + c660_wt[self.c660_col['bottoms_x']].values*c660_mf_bot_p
c670_feed = pd.DataFrame(c670_feed,index=idx,columns=self.c670_col['combined'])
c670_bf = pd.DataFrame(index=idx,columns=self.c670_col['upper_bf'])
c620_bot_x = c620_wt[self.c620_col['bottoms_x']].values
c660_bot_x = c660_wt[self.c660_col['bottoms_x']].values
upper_bf = (c660_bot_x*c660_mf_bot)/(c620_bot_x*c620_mf_bot+c660_bot_x*c660_mf_bot)
upper_bf = pd.DataFrame(upper_bf,index=idx,columns=self.c670_col['upper_bf'])
upper_bf[list(set(self.index_9999)&set(upper_bf.columns))] = 0.9999
upper_bf[list(set(self.index_0001)&set(upper_bf.columns))] = 0.0001
# c670 input (feed%bf)
c670_input = c670_feed.join(upper_bf)
c670_output = self.c670_model.predict(c670_input)
c670_sp,c670_op = c670_output.iloc[:,:41*2],c670_output.iloc[:,41*2:]
# update by c670 real data model?
if self.real_data_mode == True:
if self._linear_model == True:
c670_op_real = self.c670_real_data_model_linear.predict(c670_input)[:,41*2:]
c670_op_real = pd.DataFrame(c670_op_real,index=c670_input.index,columns=self.c670_simple_op_col)
c670_sp_real = self.c670_real_data_model_linear.predict(c670_input)[:,:41*2]
c670_sp_real = pd.DataFrame(c670_sp_real,index=c670_input.index,columns=c670_sp.columns)
if self._linear_model == False:
c670_op_real = self.c670_real_data_model.predict(c670_input).iloc[:,41*2:] #操作條件放後面
c670_sp_real = self.c670_real_data_model.predict(c670_input).iloc[:,:41*2] #分離係數放前面
c670_op.update(c670_op_real)
c670_sp.update(c670_sp_real)
# c670 sp後處理
if self._Post_processing:
for i in self.c670_wt_always_same_split_factor_dict.keys():
c670_sp[i] = self.c670_wt_always_same_split_factor_dict[i]
s1 = c670_sp[self.c670_col['distillate_sf']].values
s2 = c670_sp[self.c670_col['bottoms_sf']].values
w1 = sp2wt(c670_feed,s1)
w2 = sp2wt(c670_feed,s2)
c670_wt = np.hstack((w1,w2))
c670_wt = pd.DataFrame(c670_wt,index = idx,columns=self.c670_col['distillate_x']+self.c670_col['bottoms_x'])
# 如果是線性模式就再update c670 wt一次,放在後處理前
if self._linear_model:
c670_wt_real = self.c670_real_data_model_linear.predict(c670_input)[:,:41*2]
c670_wt_real = pd.DataFrame(c670_wt_real,index=c670_input.index,columns=c670_wt.columns)
c670_wt.update(c670_wt_real)
# c670wt沒有後處理
return c620_wt,c620_op,c660_wt,c660_op,c670_wt,c670_op
# + [markdown] id="Cb7TJSTp444q"
# # 試算模式測試
# + id="bEL2f8Kmq9ow"
f = F(config)
f.Recommended_mode = False
f.real_data_mode = False
f._Post_processing = True
f._linear_model = False
# + id="0cjAoL-Z4q4z"
commom_idx = list(set(icg_df.index)&
set(c620_df.index)&
set(c660_df.index)&
set(c670_df.index)&
set(t651_df.index))
idx = np.random.choice(commom_idx,size=100,replace=False,p=None)
# minibatch input
icg_input = icg_input.loc[idx]
c620_feed = c620_feed.loc[idx]
t651_feed = t651_feed.loc[idx]
# minibatch output
c620_op = c620_op.loc[idx]
c620_wt = c620_wt.loc[idx]
c660_op = c660_op.loc[idx]
c660_wt = c660_wt.loc[idx]
c670_op = c670_op.loc[idx]
c670_wt = c670_wt.loc[idx]
# + [markdown] id="gEW6WdmS-m55"
# # predict output
# + id="z23Si7irsA5P"
c620_wt_,c620_op_,c660_wt_,c660_op_,c670_wt_,c670_op_ = f(icg_input,c620_feed,t651_feed)
# + [markdown] id="7X3v7hYJ6jfh"
# # c620 show_metrics
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="qrGCMeFvsBEV" outputId="9edd5068-7b6d-4068-b3de-d75e6ad5dddb"
f.c620_model.show_metrics(c620_wt,c620_wt_,e=2e-2)
# + colab={"base_uri": "https://localhost:8080/", "height": 390} id="I8RmO0CosBG9" outputId="3be4e919-4b7f-45ef-e073-b41ec2103d10"
f.c620_model.show_metrics(c620_op,c620_op_,e=2e-2)
# + [markdown] id="P37pbl4F-wJE"
# # c660 show_metrics
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="A6SzCgpXsBJr" outputId="a6e458ac-e8d5-47c2-94cc-7970e142da0e"
f.c660_model.show_metrics(c660_wt,c660_wt_,e=2e-2)
# + colab={"base_uri": "https://localhost:8080/", "height": 390} id="pYJcYfOsztlz" outputId="b950ce5f-463e-43c7-f2e7-2835bedd685a"
f.c660_model.show_metrics(c660_op,c660_op_,e=2e-2)
# + [markdown] id="ojwKS0TP-yTa"
# # c670 show_metrics
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="odK0RWomzwNE" outputId="199619ec-7645-4dd6-c06a-5ff8d713156c"
f.c670_model.show_metrics(c670_wt,c670_wt_,e=2e-2)
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="AF7PspgF29yO" outputId="34b5c385-5bc3-4322-970b-12f18c20cb31"
f.c670_model.show_metrics(c670_op,c670_op_,e=2e-2)
# + [markdown] id="xHtEOnkCnomJ"
# # 推薦模式 測試
# + id="Zmu8FCxhVIri" colab={"base_uri": "https://localhost:8080/"} outputId="4b614ca1-74e0-4139-8157-396e13b3b66f"
# change mode
f.Recommended_mode = True
f.Recommended_mode
# + id="J2fSD-la4UZT" colab={"base_uri": "https://localhost:8080/", "height": 185} outputId="179ddbdc-cc07-4ed1-96ce-ee8e4c864984"
# select one sample c620_side == 70 & NA in Benzene == 980
cond = (icg_input['Simulation Case Conditions_Spec 1 : Benzene in C620 Sidedraw_wt%']==70)&(icg_input['Simulation Case Conditions_Spec 2 : NA in Benzene_ppmw']==980)
sample = icg_input.sample(1)
sample.head()
# + id="mAuevmR23Zsq" colab={"base_uri": "https://localhost:8080/"} outputId="8d6717fa-dd40-4e23-c612-e8e8f3951248"
idx = sample.index
print(idx)
# + id="NdLYvIBYnR-Q"
demo = {
# input
'icg_input':icg_input.loc[idx],
'c620_feed':c620_feed.loc[idx],
't651_feed':t651_feed.loc[idx],
# output
'c620_op':c620_op.loc[idx],
'c620_wt':c620_wt.loc[idx],
'c660_op':c660_op.loc[idx],
'c660_wt':c660_wt.loc[idx],
'c670_op':c670_op.loc[idx],
'c670_wt':c670_wt.loc[idx],
}
# + id="K8M9MvKrUEZP"
icg_input = demo['icg_input'].copy()
c620_feed = demo['c620_feed'].copy()
t651_feed = demo['t651_feed'].copy()
# + id="3dhFIyj2VDvY" colab={"base_uri": "https://localhost:8080/"} outputId="5a9dc1c9-4f72-477f-b4ce-838a7695c62e"
c620_wt_,c620_op_,c660_wt_,c660_op_,c670_wt_,c670_op_ = f(icg_input,c620_feed,t651_feed)
# + id="vBaxMbP5fn-t" colab={"base_uri": "https://localhost:8080/"} outputId="17ec18c7-7fe9-4802-ea4c-1423f64b8aa7"
for i in [c620_wt_,c620_op_,c660_wt_,c660_op_,c670_wt_,c670_op_]:
print(i.isnull().sum().sum())
# + id="oBGAZznfdAH8" colab={"base_uri": "https://localhost:8080/", "height": 185} outputId="e27c2874-e2bb-4556-e7c6-e1a295cce6b5"
demo['icg_input'].head()
# + id="Ur4DKaAloVW1" colab={"base_uri": "https://localhost:8080/"} outputId="3e85ef2d-a165-4865-da61-91463931bb6f"
import joblib
joblib.dump(demo,"/content/drive/MyDrive/台塑輕油案子/data/c620/demo/demo.pkl")
# + id="nba31qndo6hT"
| notebook/Integration_and_Test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # FAQ
# An uncleaned list of questions with raw answers that have been asked during the course.
# ## Pandas
How do you remove a row from a serie?
https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.Series.pop.html
Note that under normal circumstances you won't be mutating Series like this
Another option: https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.Series.drop.html
Series.drop
I'm used to work with Pycharm to make python scripts, is it also possible to debug in jupyter?
if you use iloc you define always first the rows and then the columns?
rows then columns
can you only show 2 decimals in a DataFrame?
https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.round.html
what does the "axis" argument exactly mean in the context of the dataframe?
https://i.stack.imgur.com/h1alT.jpg
Can I import multiple files to a list , for instance to loop through each data frame?
data = []
filenames = [...]
for fn in filenames:
data.append(pd.read_csv(fn))
# +
Remark: iteratively appending rows to a DataFrame can be more computationally intensive than a single concatenate. A better solution is to append those rows to a list and then concatenate the list with the original DataFrame all at once.
if we want to use this remark then the row indeces will be ruined?
if the row indeces are integers you can use ignore_index = True to let them keep counting. If you know how the keys will look like you can use also an argument key that accepts a list of let's say strings that will be your row indeces
# -
# ## Visualizations
what does sns.set_context("paper") do exactly?
scaling: https://seaborn.pydata.org/generated/seaborn.plotting_context.html#seaborn.plotting_context
# ## BioPython
How do I search for a method for instance, Now that I see that append doesn't work for a seq object. How can I find the correct method?
dir(...)
# +
so, how can we see all the things in annotations?
basically you can call the annotations and see everytihng that is in there print(record.annotations. But you'll indeed see that not everything is in there. So this link will give you more information.
https://biopython.readthedocs.io/en/latest/chapter_seq_annot.html
(scroll down to references)
r = record.annotations['references']
dir(r[0])
# -
'Parsing' basically means reading in?
Parsing means to "understand" the structure of the data
Is it also possible to do a multiple alignment with specific proteins?
Can you use qblast() to aling a fasta sequence against other fasta sequences instead aling the first one agains the all data base?
# ## General questions
Question regarding the usage of R and Python, how to know which is best to use for the any given task?
Following the literature in your field helps. What are the rest of the academic people in your field using moistly. This language likely has the most libraries that could be useful for you. For intstance, I work mostly with single cell omics. Most of the articles also give a link to the code they used for the analsys and I can see that they mostly use R and occasionally python. With th hints about the algorithms used, you can start browsing github to see various tools
| solutions/FAQ.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DQNs on GCP
#
# Reinforcement Learning (RL) Agents can be quite fickle. This is because the environment for an Agent is different than that of Supervised and Unsupervised algorithms.
#
# | Supervised / Unsupervised | Reinforcement Learning |
# | ----------- | ----------- |
# | Data is previously gathered | Data needs to be simulated |
# | Big Data: Many examples covering many siutations | Sparse Data: Agent trades off between exploring and exploiting |
# | The environment is assumed static | The environment may change in response to the agent |
#
# Because of this, hyperparameter tuning is even more crucial in RL as it not only impacts the training of the agent's neural network, but it also impacts how the data is gathered through simulation.
#
# ## Setup
#
# Hypertuning takes some time, and in this case, it can take anywhere between **10 - 30 minutes**. If this hasn't been done already, run the cell below to kick off the training job now. We'll step through what the code is doing while our agents learn.
# + language="bash"
# BUCKET=<your-bucket-here> # Change to your bucket name
# JOB_NAME=dqn_on_gcp_$(date -u +%y%m%d_%H%M%S)
# REGION='us-central1' # Change to your bucket region
# IMAGE_URI=gcr.io/qwiklabs-resources/rl-qwikstart/dqn_on_gcp@sha256:326427527d07f30a0486ee05377d120cac1b9be8850b05f138fc9b53ac1dd2dc
#
# gcloud ai-platform jobs submit training $JOB_NAME \
# --staging-bucket=gs://$BUCKET \
# --region=$REGION \
# --master-image-uri=$IMAGE_URI \
# --scale-tier=BASIC_GPU \
# --job-dir=gs://$BUCKET/$JOB_NAME \
# --config=hyperparam.yaml
# -
# The above command sends a [hyperparameter tuning job](https://cloud.google.com/ml-engine/docs/hyperparameter-tuning-overview) to the [Google Cloud AI Platform](https://cloud.google.com/sdk/gcloud/reference/ai-platform/jobs/submit/training). It's a service that sets up scaling distributed training so data scientists and machine learning engineers do not have to worry about technical infrastructure. Usually, it automatically selects the [container environment](https://cloud.google.com/ml-engine/docs/runtime-version-list), but we're going to take advantage of a feature to specify our own environment with [Docker](https://www.docker.com/resources/what-container). Not only will this allow us to install our game environment to be deployed to the cloud, but it will also significantly speed up hyperparameter tuning time as each worker can skip the library installation steps.
#
# The <a href="Dockerfile">Dockerfile</a> in this directory shows the steps taken to build this environment. First, we copy from a [Google Deep Learning Container](https://cloud.google.com/ai-platform/deep-learning-containers/docs/choosing-container) which already has Google Cloud Libraries installed. Then, we install our other desired modules and libraries. `ffmpeg`, `xvfb`, and `python-opengl` are needed in order to get video output from the server. Machines on the cloud don't typically have a display (why would they need one?), so we'll make a virtual display of our own.
#
# After we copy our code, we tell the container to be configured as an executable so we can pass our hyperparameter tuning flags to it with the [ENTRYPOINT](https://stackoverflow.com/questions/21553353/what-is-the-difference-between-cmd-and-entrypoint-in-a-dockerfile) command. In order to set up our virtual display, we can use the [xvfb-run](http://manpages.ubuntu.com/manpages/trusty/man1/xvfb-run.1.html) command. Unfortunately, Docker strips quotes from specified commands in ENTRYPOINT, so we'll make a super simple shell script, <a href="train_model.sh">train_model.sh</a>, to specify our virtual display parameters. The `"@"` parameter is used to pass the flags called against the container to our python module, `trainer.trainer`.
#
# ## CartPole-v0
#
# So what is the game we'll be solving for? We'll be playing with [AI Gym's CartPole Environment](https://gym.openai.com/envs/CartPole-v1/). As MNIST is the "Hello World" of image classification, CartPole is the "Hello World" of Deep Q Networks. Let's install [OpenAI Gym](https://gym.openai.com/) and play with the game ourselves!
# !python3 -m pip freeze | grep gym || python3 -m pip install --user gym==0.12.5
# !python3 -m pip freeze | grep 'tensorflow==2\|tensorflow-gpu==2' || \
# python3 -m pip install --user tensorflow==2
# ###### Note: Restart the kernel if the above libraries needed to be installed
#
# The `gym` library hosts a number of different gaming environments that our agents (and us humans) can play around in. To make an environment, we simply need to pass it what game we'd like to play with the `make` method.
#
# This will create an environment object with a number of useful methods and properties.
# * The `observation_space` parameter is the structure of observations about the environment.
# - Each "state" or snapshot or our environment will follow this structure
# * The `action_space` parameter is the possible actions the agent can take
#
# So for example, with CartPole, there are 4 observation dimensions which represent `[Cart Position, Cart Velocity, Pole Angle, Pole Velocity At Tip]`. For the actions, there are 2 possible actions to take: 0 pushes the cart to the left, and 1 pushes the cart to the right. More detail is described in the game's code [here](https://github.com/openai/gym/blob/master/gym/envs/classic_control/cartpole.py).
# +
from collections import deque
import random
import gym
import numpy as np
import tensorflow as tf
from tensorflow.keras import layers, models
env = gym.make('CartPole-v0')
print("The observation space is", env.observation_space)
print("The observation dimensions are", env.observation_space.shape)
print("The action space is", env.action_space)
print("The number of possible actions is", env.action_space.n)
# -
# * The `reset` method will restart the environment and return a starting state.
# * The `step` method takes an action, applies it to the environment and returns a new state. Each step returns a new state, the transition reward, whether the game is over or not, and game specific information. For CartPole, there is no extra info, so it returns a blank dictionary.
# +
def print_state(state, step, reward=None):
format_string = 'Step {0} - Cart X: {1:.3f}, Cart V: {2:.3f}, Pole A: {3:.3f}, Pole V:{4:.3f}, Reward:{5}'
print(format_string.format(step, *tuple(state), reward))
state = env.reset()
step = 0
print_state(state, step)
# -
action = 0
state_prime, reward, done, info = env.step(action)
step += 1
print_state(state_prime, step, reward)
print("The game is over." if done else "The game can continue.")
print("Info:", info)
# Run the cell below repeatedly until the game is over, changing the action to push the cart left (0) or right (1). The game is considered "won" when the pole can stay up for an average of steps 195 over 100 games. How far can you get? An agent acting randomly can only survive about 10 steps.
# +
action = 1 # Change me: 0 Left, 1 Right
state_prime, reward, done, info = env.step(action)
step += 1
print_state(state_prime, step, reward)
print("The game is over." if done else "The game can continue.")
# -
# We can make our own policy and create a loop to play through an episode (one full simulation) of the game. Below, actions are generated to alternate between pushing the cart left and right. The code is very similar to how our agents will be interacting with the game environment.
# +
# [0, 1, 0, 1, 0, 1, ...]
actions = [x % 2 for x in range(200)]
state = env.reset()
step = 0
episode_reward = 0
done = False
while not done and step < len(actions):
action = actions[step] # In the future, our agents will define this.
state_prime, reward, done, info = env.step(action)
episode_reward += reward
step += 1
state = state_prime
print_state(state, step, reward)
end_statement = "Game over!" if done else "Ran out of actions!"
print(end_statement, "Score =", episode_reward)
# -
# It's a challenge to get to 200! We could repeatedly experiment to find the best heuristics to beat the game, or we could leave all that work to the robot. Let's create an intelligence to figure this out for us.
#
# ## The Theory Behind Deep Q Networks
#
# The fundamental principle behind RL is we have two entities: the **agent** and the **environment**. The agent takes state and reward information about the envionment and chooses an action. The environment takes that action and will change to be in a new state.
#
# <img src="images/agent_and_environment.jpg" width="476" height="260">
#
# RL assumes that the environment follows a [Markov Decision Process (MDP)](https://en.wikipedia.org/wiki/Markov_decision_process). That means the state is dependent partially on the agent's actions, and partially on chance. MDPs can be represented by a graph, with states and actions as nodes, and rewards and path probabilities on the edges.
#
# <img src="images/mdp.jpg" width="471" height="243">
#
# So what would be the best path through the graph above? Or perhaps a more difficult question, what would be our expected winnings if we played optimally? The probability introduced in this problem has inspired multiple strategies over the years, but all of them boil down to the idea of discounted future rewards.
#
# Would you rather have `$100` now or `$105` a year from now? With inflation, there's no definitive answer, but each of us has a threshold that we use to determine the value of something now versus the value of something later. In psychology, this is called [Delayed Gratification](https://en.wikipedia.org/wiki/Delayed_gratification). <NAME> expressed this theory in an equation widely used in RL called the [Bellman Equation](https://en.wikipedia.org/wiki/Bellman_equation). Let's introduce some vocab to better define it.
#
# | Symbol | Name | Definition | Example |
# | - | - | - | - |
# | | agent | An entity that can act and transition between states | Us when we play CartPole |
# | s | state | The environmental parameters describing where the agent is | The position of the cart and angle of the pole |
# | a | action | What the agent can do within a state | Pushing the cart left or right |
# | t | time / step | One transition between states | One push of the cart |
# || episode | One full simulation run | From the start of the game to game over |
# | v, V(s) | value | How much a state is worth | V(last state dropping the pole) = 0
# | r, R(s, a) | reward | Value gained or lost transitioning between states through an action | R(keeping the pole up) = 1 |
# | γ | gamma | How much to value a current state based on a future state | Coming up soon |
# | 𝜋, 𝜋(s) | policy |The recommended action to the agent based on the current state | π(in trouble) = honesty |
#
# Bellman realized this: The value of our current state should the discounted value of the next state the agent will be in plus any rewards picked up along the way, given the agent takes the best action to maximize this.
#
# Using all the symbols from above, we get:
#
# <img src="images/bellman_equation.jpg" width="260" height="50">
#
# However, this is assuming we know all the states, their corresponding actions, and their rewards. If we don't know this in advance, we can explore and simulate this equation with what is called the [Q equation](https://en.wikipedia.org/wiki/Q-learning):
#
# <img src="https://wikimedia.org/api/rest_v1/media/math/render/svg/47fa1e5cf8cf75996a777c11c7b9445dc96d4637">
#
# Here, the value function is replaced with the Q value, which is a function of a state and action. The learning rate is how much we want to change our old Q value with new information found during simulation. Visually, this results in a Q-table, where rows are the states, actions are the columns, and each cell is the value found through simulation.
#
# || Meal | Snack | Wait |
# |-|-|-|-|
# | Hangry | 1 | .5 | -1 |
# | Hungry | .5 | 1 | 0 |
# | Full | -1 | -.5 | 1.5 |
#
# So this is cool and all, but how exactly does this fit in with CartPole? Here, MDPs are discrete states. CartPole has multidimensional states on a continuous scale. This is where neural networks save the day! Rather than categorize each state, we can feed state properties into our network. By having the same number of output nodes as possible actions, our network can be used to predict the value of the next state given the current state and action.
#
# ## Building the Agent
#
# These networks can be configured with the same architectures and tools as other problems, such as CNNs and LSTMs. However, the one gotcha is that uses a specialized loss function. We'll instead be using the derivative of the Bellman Equation. Let's go ahead and define our model function as it is in trainer/model.py
def deep_q_network(
state_shape, action_size, learning_rate, hidden_neurons):
"""Creates a Deep Q Network to emulate Q-learning.
Creates a two hidden-layer Deep Q Network. Similar to a typical nueral
network, the loss function is altered to reduce the difference between
predicted Q-values and Target Q-values.
Args:
space_shape: a tuple of ints representing the observation space.
action_size (int): the number of possible actions.
learning_rate (float): the nueral network's learning rate.
hidden_neurons (int): the number of neurons to use per hidden
layer.
"""
state_input = layers.Input(state_shape, name='frames')
actions_input = layers.Input((action_size,), name='mask')
hidden_1 = layers.Dense(hidden_neurons, activation='relu')(state_input)
hidden_2 = layers.Dense(hidden_neurons, activation='relu')(hidden_1)
q_values = layers.Dense(action_size)(hidden_2)
masked_q_values = layers.Multiply()([q_values, actions_input])
model = models.Model(
inputs=[state_input, actions_input], outputs=masked_q_values)
optimizer = tf.keras.optimizers.RMSprop(lr=learning_rate)
model.compile(loss='mse', optimizer=optimizer)
return model
# Notice any other atypical aspects of this network?
#
# Here, we take in both state and actions as inputs to our network. The states are fed in as normal, but the actions are used to "mask" the output. This is actually used for faster training, as we'd only want to update the nodes correspnding to the action that we simulated.
#
# The Bellman Equation actually isn't in the network. That's because this is only the "brain" of our agent. As an intelligence, it has much more! Before we get to how exactly the agent learns, let's looks at the other aspects of its body: "Memory" and "Exploration".
#
# Just like other neural network algorithms, we need data to train on. However, this data is the result of our simulations, not something previously stored in a table. Thus, we're going to give our agent a memory where we can store state - action - new state transitions to learn on.
#
# Each time the agent takes a step in gym, we'll save `(state, action, reward, state_prime, done)` to our buffer, which is defined like so.
class Memory():
"""Sets up a memory replay buffer for a Deep Q Network.
A simple memory buffer for a DQN. This one randomly selects state
transitions with uniform probability, but research has gone into
other methods. For instance, a weight could be given to each memory
depending on how big of a difference there is between predicted Q values
and target Q values.
Args:
memory_size (int): How many elements to hold in the memory buffer.
batch_size (int): The number of elements to include in a replay batch.
gamma (float): The "discount rate" used to assess Q values.
"""
def __init__(self, memory_size, batch_size, gamma):
self.buffer = deque(maxlen=memory_size)
self.batch_size = batch_size
self.gamma = gamma
def add(self, experience):
"""Adds an experience into the memory buffer.
Args:
experience: a (state, action, reward, state_prime, done) tuple.
"""
self.buffer.append(experience)
def sample(self):
"""Uniformally selects from the replay memory buffer.
Uniformally and randomly selects experiences to train the nueral
network on. Transposes the experiences to allow batch math on
the experience components.
Returns:
(list): A list of lists with structure [
[states], [actions], [rewards], [state_primes], [dones]
]
"""
buffer_size = len(self.buffer)
index = np.random.choice(
np.arange(buffer_size), size=self.batch_size, replace=False)
# Columns have different data types, so numpy array would be awkward.
batch = np.array([self.buffer[i] for i in index]).T.tolist()
states_mb = tf.convert_to_tensor(np.array(batch[0], dtype=np.float32))
actions_mb = np.array(batch[1], dtype=np.int8)
rewards_mb = np.array(batch[2], dtype=np.float32)
states_prime_mb = np.array(batch[3], dtype=np.float32)
dones_mb = batch[4]
return states_mb, actions_mb, rewards_mb, states_prime_mb, dones_mb
# Let's make a fake buffer and play around with it! We'll add the memory into our game play code to start collecting experiences.
# +
test_memory_size = 20
test_batch_size = 4
test_gamma = .9 # Unused here. For learning.
test_memory = Memory(test_memory_size, test_batch_size, test_gamma)
# +
actions = [x % 2 for x in range(200)]
state = env.reset()
step = 0
episode_reward = 0
done = False
while not done and step < len(actions):
action = actions[step] # In the future, our agents will define this.
state_prime, reward, done, info = env.step(action)
episode_reward += reward
test_memory.add((state, action, reward, state_prime, done)) # New line here
step += 1
state = state_prime
print_state(state, step, reward)
end_statement = "Game over!" if done else "Ran out of actions!"
print(end_statement, "Score =", episode_reward)
# -
# Now, let's sample the memory by running the cell below multiple times. It's different each call, and that's on purpose. Just like with other neural networks, it's important to randomly sample so that our agent can learn from many different situations.
#
# The use of a memory buffer is called [Experience Replay](https://arxiv.org/pdf/1511.05952.pdf). The above technique of a uniform random sample is a quick and computationally efficient way to get the job done, but RL researchers often look into other sampling methods. For instance, maybe there's a way to weight memories based on their rarity or loss when the agent learns with it.
test_memory.sample()
# But before the agent has any memories and has learned anything, how is it supposed to act? That comes down to [Exploration vs Exploitation](https://en.wikipedia.org/wiki/Multi-armed_bandit). The trouble is that in order to learn, risks with the unknown need to be made. There's no right answer, but there is a popular answer. We'll start by acting randomly, and over time, we will slowly decay our chance to act randomly.
#
# Below is a partial version of the agent.
class Partial_Agent():
"""Sets up a reinforcement learning agent to play in a game environment."""
def __init__(self, network, memory, epsilon_decay, action_size):
"""Initializes the agent with DQN and memory sub-classes.
Args:
network: A neural network created from deep_q_network().
memory: A Memory class object.
epsilon_decay (float): The rate at which to decay random actions.
action_size (int): The number of possible actions to take.
"""
self.network = network
self.action_size = action_size
self.memory = memory
self.epsilon = 1 # The chance to take a random action.
self.epsilon_decay = epsilon_decay
def act(self, state, training=False):
"""Selects an action for the agent to take given a game state.
Args:
state (list of numbers): The state of the environment to act on.
traning (bool): True if the agent is training.
Returns:
(int) The index of the action to take.
"""
if training:
# Random actions until enough simulations to train the model.
if len(self.memory.buffer) >= self.memory.batch_size:
self.epsilon *= self.epsilon_decay
if self.epsilon > np.random.rand():
print("Exploration!")
return random.randint(0, self.action_size-1)
# If not acting randomly, take action with highest predicted value.
print("Exploitation!")
state_batch = np.expand_dims(state, axis=0)
predict_mask = np.ones((1, self.action_size,))
action_qs = self.network.predict([state_batch, predict_mask])
return np.argmax(action_qs[0])
# Let's define the agent and get a starting state to see how it would act without any training.
# +
state = env.reset()
# Define "brain"
space_shape = env.observation_space.shape
action_size = env.action_space.n
# Feel free to play with these
test_learning_rate = .2
test_hidden_neurons = 10
test_epsilon_decay = .95
test_network = deep_q_network(
space_shape, action_size, test_learning_rate, test_hidden_neurons)
test_agent = Partial_Agent(
test_network, test_memory, test_epsilon_decay, action_size)
# -
# Run the cell below multiple times. Since we're decaying the random action rate after every action, it's only a matter a time before the agent exploits more than it explores.
action = test_agent.act(state, training=True)
print("Push Right" if action else "Push Left")
# Memories, a brain, and a healthy dose of curiosity. We finally have all the ingredient for our agent to learn. After all, as the Scarecrow from the Wizard of Oz said:
#
# "Everything in life is unusual until you get accustomed to it."
# ~L. <NAME>
#
# Below is the code used by our agent to learn, where the Bellman Equation at last makes an appearance. We'll run through the following steps.
#
# 1. Pull a batch from memory
# 2. Get the Q value (the output of the neural network) based on the memory's ending state
# - Assume the Q value of the action with the highest Q value (test all actions)
# 4. Update these Q values with the Bellman Equations
# - `target_qs = (next_q_mb * self.memory.gamma) + reward_mb`
# - If the state is the end of the game, set the target_q to the reward for entering the final state.
# 5. Reshape the target_qs to match the networks output
# - Only learn on the memory's corresponding action by setting all action nodes to zero besides the action node taken.
# 6. Fit Target Qs as the label to our model against the memory's starting state and action as the inputs.
# +
def learn(self):
"""Trains the Deep Q Network based on stored experiences."""
batch_size = self.memory.batch_size
if len(self.memory.buffer) < batch_size:
return None
# Obtain random mini-batch from memory.
state_mb, action_mb, reward_mb, next_state_mb, done_mb = (
self.memory.sample())
# Get Q values for next_state.
predict_mask = np.ones(action_mb.shape + (self.action_size,))
next_q_mb = self.network.predict([next_state_mb, predict_mask])
next_q_mb = tf.math.reduce_max(next_q_mb, axis=1)
# Apply the Bellman Equation
target_qs = (next_q_mb * self.memory.gamma) + reward_mb
target_qs = tf.where(done_mb, reward_mb, target_qs)
# Match training batch to network output:
# target_q where action taken, 0 otherwise.
action_mb = tf.convert_to_tensor(action_mb, dtype=tf.int32)
action_hot = tf.one_hot(action_mb, self.action_size)
target_mask = tf.multiply(tf.expand_dims(target_qs, -1), action_hot)
return self.network.train_on_batch(
[state_mb, action_hot], target_mask, reset_metrics=False
)
Partial_Agent.learn = learn
test_agent = Partial_Agent(
test_network, test_memory, test_epsilon_decay, action_size)
# -
# Nice! We finally have an intelligence that can walk and talk and... well ok, this intelligence is too simple to be able to do those things, but maybe it can learn to push a cart with a pole on it. Let's update our training loop to use our new agent.
#
# Run the below cell over and over up to ten times to train the agent.
# +
state = env.reset()
step = 0
episode_reward = 0
done = False
while not done:
action = test_agent.act(state, training=True)
state_prime, reward, done, info = env.step(action)
episode_reward += reward
test_agent.memory.add((state, action, reward, state_prime, done)) # New line here
step += 1
state = state_prime
print_state(state, step, reward)
test_agent.learn()
print(end_statement, "Game over! Score =", episode_reward)
# -
# ## Hypertuning
#
# Chances are, at this point, the agent is having a tough time learning. Why is that? Well, remember that hyperparameter tuning job we kicked off at the start of this notebook?
#
# The are many parameters that need adjusting with our agent. Let's recap:
# * The number of `episodes` or full runs of the game to train on
# * The neural networks `learning_rate`
# * The number of `hidden_neurons` to use in our network
# * `gamma`, or how much we want to discount the future value of states
# * How quickly we want to switch from explore to exploit with `explore_decay`
# * The size of the memory buffer, `memory_size`
# * The number of memories to pull from the buffer when training, `memory_batch_size`
#
# These all have been added as flags to pass to the model in `trainer/trainer.py`'s `_parse_arguments` method. For the most part, `trainer/trainer.py` follows the structure of the training loop that we have above, but it does have a few extra bells and whistles, like a hook into TensorBoard and video output.
def _parse_arguments(argv):
"""Parses command-line arguments."""
parser = argparse.ArgumentParser()
parser.add_argument(
'--game',
help='Which open ai gym game to play',
type=str,
default='CartPole-v0')
parser.add_argument(
'--episodes',
help='The number of episodes to simulate',
type=int,
default=200)
parser.add_argument(
'--learning_rate',
help='Learning rate for the nueral network',
type=float,
default=0.2)
parser.add_argument(
'--hidden_neurons',
help='The number of nuerons to use per layer',
type=int,
default=30)
parser.add_argument(
'--gamma',
help='The gamma or "discount" factor to discount future states',
type=float,
default=0.5)
parser.add_argument(
'--explore_decay',
help='The rate at which to decay the probability of a random action',
type=float,
default=0.1)
parser.add_argument(
'--memory_size',
help='Size of the memory buffer',
type=int,
default=100000)
parser.add_argument(
'--memory_batch_size',
help='The amount of memories to sample from the buffer while training',
type=int,
default=8)
parser.add_argument(
'--job-dir',
help='Directory where to save the given model',
type=str,
default='models/')
parser.add_argument(
'--print_rate',
help='How often to print the score, 0 if never',
type=int,
default=0)
parser.add_argument(
'--eval_rate',
help="""While training, perform an on-policy simulation and record
metrics to tensorboard every <record_rate> steps, 0 if never. Use
higher values to avoid hyperparameter tuning "too many metrics"
error""",
type=int,
default=20)
return parser.parse_known_args(argv)
# Geez, that's a lot. And like with other machine learning methods, there's no hard and fast rule and is problem dependent. Plus, there are many more paramaters we could explore, like the number of layers, learning rate decay, and so on.
#
# We can tell Google Cloud how to explore the hyperparameter tuning space with a [config file](https://cloud.google.com/ml-engine/reference/rest/v1/projects.jobs#HyperparameterSpec). The `hyperparam.yaml` file in this directory is exactly that. It specifies which parameter to tune on (in this case, the `episode_reward`) and the range for the different flags we want to tune on.
#
# In our code, we'll add the following
#
# `import hypertune #From cloudml-hypertune library`
#
# `hpt = hypertune.HyperTune() # Initialized before looping through episodes`
#
# `# Placed right before the end of the training loop
# hpt.report_hyperparameter_tuning_metric(
# hyperparameter_metric_tag='episode_reward',
# metric_value=reward,
# global_step=episode)`
#
# This way, at the end of every episode, we can send information to the tuning service on how the agent is doing. The service can only handle so much information being thrown at it at once, so we'll add a `eval_rate` flag to throttle information to every `eval_rate` episodes.
#
# It is definately a worthwhile exercise to try and find the optimal set of parameters on one's on, but if life is too short, and there isn't time for that, the hyperparameter tuning job should now be complete. Head on over to [Google Cloud's AI Platform](https://console.cloud.google.com/ai-platform/jobs) to see the job labeled `dqn_on_gcp_<time_this_lab_was_started>`
#
# **TODO insert image**
#
# Click on the job name to see the results. Information comes in as each trial is complete, and the best performing trial will be listed on the top.
#
# <img src="images/hypertune_trials.jpg" width="966" height="464">
# Logs can be invaluable when debugging. Click the three dots to the right of one of the trials to filter logs by that particular trial.
#
# At last, let's see the results of the best trial. Keep in mind the best trial number and navigate over to [your bucket](https://console.cloud.google.com/storage/browser). The results will be in a file with the same Job Name as your hyperparameter tuning job. In that folder, there will be a number of subfolders equal to the number of hyperparameter tuning trials. Select the folder with your best performing `Trial Id`
#
# <img src="images/best_trial.jpg" width="956" height="456">
#
# There should be a number of goodies in the file including TensorBoard information in `/train`, a saved model in `saved_model.pb`, and a recording of the model in `recording.mp4`.
#
# Open the [Google Cloud Shell](https://console.cloud.google.com/home/dashboard?cloudshell=true&_ga=2.207467987.-157492093.1570741979) and run Tensorboard with
#
# `tensorboard --logdir=gs://<your-bucket>/<job-name>/<path-best-trial>`
#
# The episode rewards and training loss are displayed for the trial in intervals of 20 episodes.
#
# <img src="images/tensorboard.jpg" width="910" height="708">
#
# Click `recording.mp4` in your bucket to visually see how the model performed! How did it do? If you're not proud of your little robot, check out the recordings of the other trials to see how it decimates the competition.
#
# Congratulations on making a Deep Q Agent! That's it for now, but this is just scratching the surface for Reinforcement Learning. AI Gym has plenty of other [environments](https://gym.openai.com/envs/#classic_control), see if you can conquer them with your new skills!
# Copyright 2019 Google Inc.
# Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
| quests/rl/dqn/dqns_on_gcp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import os
from PIL import Image
import matplotlib.pyplot as plt
# this is nemo's "core" package
import nemo
# this is nemos's collection of GAN-related modules used for this example
import nemo.collections.simple_gan as nemo_simple_gan
# -
# Define model parameters
batch_size = 64
data_root = "."
# ### Instantiate necessary Neural Modules
mnist_data = nemo_simple_gan.MnistGanDataLayer(
batch_size=batch_size,
shuffle=True,
train=True,
root=data_root
)
generator = nemo_simple_gan.SimpleGenerator(batch_size=batch_size)
discriminator = nemo_simple_gan.SimpleDiscriminator()
interpolater = nemo_simple_gan.InterpolateImage()
# Loss_D = D(interpolated) - D(real) + lambda * GP
disc_loss = nemo_simple_gan.DiscriminatorLoss()
neg_disc_loss = nemo_simple_gan.DiscriminatorLoss(neg=True)
disc_grad_penalty = nemo_simple_gan.GradientPenalty(lambda_=10)
# ### Describe how Neural Modules are connected together
# Create generator DAG
# Grab data from data layer
latents, real_data, _ = mnist_data()
# Generate image from latents
generated_image = generator(latents=latents)
# Define D(G(z)) where z represents the latents
# generator_decision is a tensor that represents D(G(z))
generator_decision = discriminator(image=generated_image)
# Define loss_G = - mean(D(G(z)))
generator_loss = neg_disc_loss(decision=generator_decision)
# +
# Create discriminator DAG
# Create interpolated image that is somewhere inbetween the real_image
# and the generated image
# Note: we reuse the generated_image tensor from the generator DAG
interpolated_image = interpolater(image1=real_data, image2=generated_image)
# Define D(x~) where x~ is the interpolated image
interpolated_decision = discriminator(image=interpolated_image)
# Define D(x) where x is the real image
real_decision = discriminator(image=real_data)
# Define the components of the discriminator loss
# interpolated_loss = mean(D(x~))
interpolated_loss = disc_loss(decision=interpolated_decision)
# real_loss = - mean(D(x))
real_loss = neg_disc_loss(decision=real_decision)
# grad_penalty = mean(lambda* (|gradients| - 1) ** 2)
grad_penalty = disc_grad_penalty(
interpolated_image=interpolated_image,
interpolated_decision=interpolated_decision)
# Note the final loss_D = interpolated_loss + real_loss + grad_penalty
# -
# Create Eval DAG
# Create a new datalayer that samples from the latent distribution
random_data = nemo_simple_gan.RandomDataLayer(batch_size=batch_size)
# Create a new NmTensor to get data from the data layer
latents_e = random_data()
# Generate from latents
generated_image_e = generator(latents=latents_e)
# ### Run training
# Create a neural factory
neural_factory = nemo.core.NeuralModuleFactory(backend=nemo.core.Backend.PyTorch)
trainer = neural_factory._trainer
# Define the losses
losses_G = [generator_loss]
losses_D = [interpolated_loss, real_loss, grad_penalty]
# Since we want optimizers to only operate on a subset of the model, we need
# to manually create optimizers
# For single loss and single optimizer, the following steps can be skipped
# and an optimizer will be created in trainer.train()
optimizer_G = trainer.create_optimizer(
optimizer="adam",
things_to_optimize=[generator],
optimizer_params={
"lr": 1e-4,
"betas": (0.5, 0.9),
})
optimizer_D = trainer.create_optimizer(
optimizer="adam",
things_to_optimize=[discriminator],
optimizer_params={
"lr": 1e-4,
"betas": (0.5, 0.9),
})
# +
# Define some helper functions to log generated samples
def save_image(global_vars):
images = global_vars["image"]
image = images[0].squeeze(0).detach().cpu().numpy() * 255
plt.imshow(image, cmap="gray")
plt.show()
def put_tensor_in_dict(tensors, global_vars):
global_vars["image"] = tensors[generated_image_e.unique_name][0]
# +
# Define a callback that generates samples
eval_callback = nemo.core.EvaluatorCallback(
eval_tensors=[generated_image_e],
user_iter_callback=put_tensor_in_dict,
user_epochs_done_callback=lambda x: save_image(x),
eval_step=1000,
)
# Define our training loop. Here we optimize take 3 discriminator steps
# prior to taking the generator step
tensors_to_optimize = [
(optimizer_D, losses_D),
(optimizer_D, losses_D),
(optimizer_D, losses_D),
(optimizer_G, losses_G),
]
# Finally, call train with our training loop and callbacks
trainer.train(
tensors_to_optimize=tensors_to_optimize,
callbacks=[eval_callback],
optimization_params={"num_epochs": 10})
# -
| examples/image/simple_gan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Determining genotypes
#
# This excersize aims at applying Bayesian statistics to calculate the probability of a specific genotype. In particular we will:
# 1. Combine two .
# 2. Compare the computed results with simple simulations.
#
# The exercise is made to play around with a simulated sample of genes and DNA. The goal is to calculate the probability of an original state, based on data and prior probability.
#
# ### References:
# - Lecture on Bayesian statistics
# - Lecture on Markov Chains
#
# ### Authors:
# - <NAME> (Niels Bohr Institute)
#
# ### Date:
# - 29-12-2021 (latest update)
#
# ***
# As always we will include a collection of libraries (Standard)
import numpy as np # Matlab like syntax for linear algebra and functions
from numpy.linalg import matrix_power # This is to raise matrices to powers
import matplotlib.pyplot as plt # Plots and figures like you know them from Matlab
import seaborn as sns # Make the plots nicer to look at
from iminuit import Minuit # The actual fitting tool, better than scipy's
import sys # Module to see files and folders in directories
from scipy import stats
# We will also (big surprise...) need a random generator:
r = np.random # Random generator
r.seed(42) # Set a random seed (but a fixed one)
# And now for the actual problem: You are given a sample of a particular gene (for simplicity say eyecolor). This gene is found in two versions: A and B. Given that we all have DNA from our fathers and mothers, there are 3 possibilities of combinations for genotypes: AA, AB and BB.
#
# So say you measure everything optimally, if you have 8 versions of the gene, coming from AB it might look like:
# - [A,B,A,B,A,A,A,B]
#
# However, you don't have the best equipment and there is a possibility that you measure a gene wron. This is given by a matrix, XX.
#
#
# .............................................................................................................................................................................................................
#
# Assume you get the sequence: [A,A,A,A,B]. We want to compute the probability for each of the three genotypes given sequences of data.
#
# .............................................................................................................................................................................................................
#
# Start by writing on paper, thinking about how you would calculate this probability.
# Information 1: The matrix of transformations (meaning the probability that your machine measures A if it was really a B), takes the form:
XX = np.zeros((2,2)); XX[0,0] = 0.85; XX[0,1] = 0.15; XX[1,0] = 0.05; XX[1,1] = 0.95;
print("XX (i.e. the matrix of transformations error) = ")
print(XX)
# Information 2: You know that in the population, there are the probabilities for each combination:
# - AA = 0.45
# - BB = 0.40
# - AB = 0.15
Pri = np.zeros(3); Pri[0] = 0.45; Pri[1] = 0.15; Pri[2] = 0.40;
print("Prior = ")
print(Pri)
S = np.zeros((2,3));
S[0,0] = 2./3; S[0,1] = 1./3; S[1,1] = 1./3; S[1,2] = 2./3 ## Matrix to end up in a genotype
print("S = ")
print(S)
# We can now construct a full matrix of the seven states:
M = np.zeros((7,7)); M[0:2,2:4] = np.transpose(XX); M[2:4,4:] = S;
print("M = ")
print(M)
# So what is the probability a measured base B comes from the genotype BB?
V1 = np.zeros(2); V1[1] = 1;
XX2 = np.transpose(XX)
P = np.matmul(V1,np.matmul(XX2,S));
print(P)
# Think about why do we use the transposed matrix?
#
# So now the probability that the sequence [A,A,A,A,B] comes from the 3 genotypes must bu the product of these probabilities (think about why this has to be the case):
Seq = [0,0,0,0,1]
LLout = np.ones(3)
LLoutFull = np.ones(3)
for igeno in range(3):
V2 = np.zeros(3); V2[igeno]= 1;
V2Full = np.zeros(7); V2Full[4+igeno]= 1;
for iseq in range(len(Seq)):
V1Full = np.zeros([1,7]); V1Full[0,Seq[iseq]] = 1;
LLoutFull[igeno] *= np.matmul(np.matmul(V1Full,np.linalg.matrix_power(M,2)),V2Full)
V1 = np.zeros([1,2]); V1[0,Seq[iseq]] = 1;
LLout[igeno] *= np.matmul(np.matmul(V1,np.matmul(XX2,S)),V2)
# Note that we could use the full matrix and square it, and we could use the combination of our XX matrix and S. These give the same:
print("The likelihood using the squared matrix is:")
print(LLoutFull)
print("The likelihood using the two submatrices is:")
print(LLout)
# Now we should produce a probability based on this likelihood. We can get a probability by including the prior or not - but these results differ of course!
print("Probabilities with the included prior ")
for i in range(3):
print(LLout[i]/np.dot(LLout,Pri)*Pri[i])
print(" ")
print("Probabilities without the included prior ")
for i in range(3):
print(LLout[i]/np.sum(LLout))
# So now we have obtained the predicted probabilites based on our calculations.
# Lets make a simulation to test if this is true!
# +
c = 0 ## counter
nsave = 100 ## 100 datapoints to save - only for plotting!
lseq = 5 ## length of the sequence
Ntest = 10000 ## Number of points to generate
PlAr = np.zeros((nsave,3))
ntrue = np.zeros(Ntest) ## array where we save the actual genotype that gave rise to a sequence
ns = np.zeros(5) ## Here we put the test-sequence for all runs
click = 1
while(c<Ntest):
A = np.random.random() ## This is a random number to determine genotype
if (A<Pri[0]): ## For genotype AA
ntrue[c] = 0
for i in range(lseq):
B = np.random.random()
if (B<XX[0,0]):
ns[i] = 0
else:
ns[i] = 1
elif (A<Pri[0]+Pri[1]): ## For genotype AB
ntrue[c] = 1
for i in range(lseq):
if (np.random.random()<0.5):
B = np.random.random()
if (B<XX[0,0]):
ns[i] = 0
else:
ns[i] = 1
else:
B = np.random.random()
if (B<XX[1,0]):
ns[i] = 0
else:
ns[i] = 1
else: ## For genotype BB
ntrue[c] = 2
for i in range(lseq):
B = np.random.random()
if (B<XX[1,0]):
ns[i] = 0
else:
ns[i] = 1
if (np.sum(ns) == 1): ## If our simulated sequence matches the desired sequence
c+= 1
if (c > click*Ntest/nsave):
l1 = np.ones(c)
PlAr[click,0] = np.sum(l1[ntrue[0:c]==0])/len(ntrue[0:c])
PlAr[click,1] = np.sum(l1[ntrue[0:c]==1])/len(ntrue[0:c])
PlAr[click,2] = np.sum(l1[ntrue[0:c]==2])/len(ntrue[0:c])
click+=1
fig, ax = plt.subplots(3)
ax[0].plot(np.linspace(1*Ntest/nsave,Ntest,nsave-1),PlAr[1:,0],'-g')
ax[0].set_xlabel('# of tested sequence')
ax[0].set_ylabel('Probability')
ax[0].set_title('Probability of state AA')
ax[1].plot(np.linspace(1*Ntest/nsave,Ntest,nsave-1),PlAr[1:,1],'-b')
ax[1].set_xlabel('# of tested sequence')
ax[1].set_ylabel('Probability')
ax[1].set_title('Probability of state AB')
ax[2].plot(np.linspace(1*Ntest/nsave,Ntest,nsave-1),PlAr[1:,2],'-r')
ax[2].set_xlabel('# of tested sequence')
ax[2].set_ylabel('Probability')
ax[2].set_title('Probability of state BB')
fig.tight_layout()
fig.set_size_inches(18.5, 10.5)
# -
l1 = np.ones(len(ntrue))
print("When we measure a sequence [A,A,A,A,B] the fractions of genotypes are:")
print(np.sum(l1[ntrue==0])/len(ntrue))
print(np.sum(l1[ntrue==1])/len(ntrue))
print(np.sum(l1[ntrue==2])/len(ntrue))
# Clearly this agrees well with the calculated result when we use the prior!
# Below here, this code can be used to calculate the probabilities by updating the prior:
######### Try to make an updated prior ##########
Seq = [0,0,0,0,1]
LLout = np.ones(3)
XX2 = np.transpose(XX)
Pri0 = np.zeros(3); Pri0[0] = 0.45; Pri0[1] = 0.15; Pri0[2] = 0.40;
for iseq in range(len(Seq)):
V1 = np.zeros([1,2]); V1[0,Seq[iseq]] = 1;
LLout = np.matmul(V1,np.matmul(XX2,S))
Pout = 0 ####### You write here but use Pri0
# This piece of code can be used to generate sequences of length Nseq in order to calculate the probabilities for the questions.
# +
######### Generate probabilities of arbitrary length ##########
Nseq = 8
ProbAA = np.zeros((Nseq,Nseq+1))
ProbAB = np.zeros((Nseq,Nseq+1))
ProbBB = np.zeros((Nseq,Nseq+1))
MaxProb = np.zeros((Nseq,Nseq+1))
for tseq in range(Nseq):
Seq = np.zeros(tseq, dtype=int)
for vseq in range(tseq+1):
if (vseq > 0):
Seq[vseq-1] = 1
LLout = np.ones(3)
for iseq in range(tseq):
V1 = np.zeros([1,2]); V1[0,Seq[iseq]] = 1;
L1 = np.matmul(V1,np.matmul(XX2,S))
LLout *= L1.flatten()
ProbAA[tseq,vseq] = LLout[0]/np.dot(LLout,Pri)*Pri[0]
ProbAB[tseq,vseq] = LLout[1]/np.dot(LLout,Pri)*Pri[1]
ProbBB[tseq,vseq] = LLout[2]/np.dot(LLout,Pri)*Pri[2]
MaxProb[tseq,vseq] = np.max(LLout/np.dot(LLout,Pri)*Pri)
fig, ax = plt.subplots(3)
ax[0].set_title('Probability for AA')
ax[0].imshow(ProbAA)
ax[0].set_aspect('equal')
ax[0].set_xlabel('Number of B in sequence')
ax[0].set_ylabel('Sequence length')
ax[1].set_title('Probability for AB')
ax[1].imshow(ProbAB)
ax[1].set_aspect('equal')
ax[1].set_xlabel('Number of B in sequence')
ax[1].set_ylabel('Sequence length')
ax[2].set_title('Probability for BB')
ax[2].imshow(ProbBB)
ax[2].set_aspect('equal')
ax[2].set_xlabel('Number of B in sequence')
ax[2].set_ylabel('Sequence length')
#fig.tight_layout()
fig.set_size_inches(28.5, 20.5)
# -
# For the original values - why is the P(BB) band more narrow than the P(AA)?
#
# Here we can calculate some relevant probabilities, for instance the maximal probability of AB and the minimal value of the most probable sequence of bases for each length of sequences (make sure you understand this).
MinProb = np.zeros(Nseq)
MaxProbAB = np.zeros(Nseq)
for i in range(Nseq):
MinProb[i] = np.min(MaxProb[i,0:i+1])
MaxProbAB[i] = np.max(ProbAB[i,0:i+1])
plt.plot(MinProb,'--xr')
plt.plot(MaxProbAB,'--+g')
plt.title('Probabilities for specific values at different sequence lengths')
plt.xlabel('Probability')
plt.ylabel('Sequence length')
# # Questions:
#
# This excersize should make you think of how to combine two different types of uncertainty with the concept of a bayesian prior.
#
# 1) Try to change the sequence of length 5 so no genotype probability is larger than 0.66? Can you make a sequence of length 10 that does not have any probability larger than 0.66?
#
# 2) For the sequence of length 5: What happens if you update the prior for each base you are reading? Does it give the same result?
#
# 3) Assume for this question that the priors P(AA) = P(BB) = 0.49. How long a sequence do you need to make the call (p>0.95) of a the genotype AB if you have equal number of A's and B's?
#
# 4) Assume again that the priors P(AA) = P(BB) = 0.49 but that the matrix XX now has the values
# - XX[0,0] = 0.99.
# - XX[1,0] = 0.01.
# - XX[0,1] = 0.01.
# - XX[1,1] = 0.99.
#
# For the sequence: [A,A,A,A,B] - does the prior change the result in an important way?
#
# 5) Lets say you are in charge of deciding the strategy of the lab department that should determine genotypes. Use the original prior.
# - How much do you need to improve the measurement accuracy, if you want p>0.95 for all sequences of length 10?
#
# ### Advanced question:
#
# 6) Assume that you had a 10% error in one of the measurement estimates (i.e. the entries in the matrix XX). Which one would afffect the probabilites the most?
#
| AppStat2022/Week6/original/GenotypeDetermination/Determining_Genotypes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 彩票预测
# 本项目使用RNN进行彩票预测。
#
# 使用人工智能技术来预测彩票,是这次的主题,那么预测哪种彩票呢?我们先选择简单一些的,就是排列组合少一些的,如果证明我们的模型work,再扩展到其他的彩票上。最终我选择了[`排列三`](https://baike.baidu.com/item/排列三/343981?fr=aladdin), 从000-999的数字中选取1个3位数,一共有1000种,中奖概率就是千分之一,够简单了吧。
#
# 历史数据在[`这里`](https://datachart.500.com/)。
#
# 数据是按照每期一组数的顺序排列的,从第一期到最新的一期,实际上是时间序列的数据。跟回归预测有很大的区别,因为特征上没有特殊的意义,不具备一组特征x映射到label y的条件。但是按照时间序列来训练的话就不一样了,输入x是一期的开奖结果,要学习的y是下一期的开奖结果。
#
# 彩票的开奖结果是一个随机分布,跟投骰子、抛硬币差不多,从数学角度看没有规律可言。我的预期是,虽然数学模型是随机的,但是一旦跟现实世界的物体发生关系,总会受到某种影响吧,比如量子纠缠,万有引力,动力学,空气阻力,空气湿度,开奖时刻的机器电压强度,开奖器材的损耗,每个球的质量的差异,吹球设备的物理特性,装球器皿的特定形状等等因素所产生的规律性的东西。
#
# 看得出来以上所列出的和没列出的都是增加不确定性、随机性的因素,但是也有可能每次开奖这些相同的特点可能造成某种规律性的结果出来,比如根据这些物理特性的影响,某个球特别容易开出来。一旦是这样,那么就有迹可循,让我们的学习器学到规律。
#
#
#
# ## LSTM介绍
# 我们需要从过往的历史数据中寻找规律,[`LSTM`](https://en.wikipedia.org/wiki/Long_short-term_memory)再适合不过了。如果你对LSTM不熟悉的话,以下几篇文章建议你阅读:
#
# [`Understanding LSTM Networks`](http://colah.github.io/posts/2015-08-Understanding-LSTMs/)
#
# [`[译] 理解 LSTM 网络`](http://www.jianshu.com/p/9dc9f41f0b29)
#
# [`RNN以及LSTM的介绍和公式梳理`](http://blog.csdn.net/Dark_Scope/article/details/47056361)
#
import tensorflow as tf
import os
# 加载数据集
def load_data(path):
input_file = os.path.join(path)
with open(input_file, "r") as f:
data = f.read()
return data
# ## 加载数据集
data_dir = './data/cp.txt'
text = load_data(data_dir)
# 一共4656条记录,4600多期了。共出现了988个不重复的结果,就是说还有(1000 – 988)12组数到现在还没有开出来过。文件中第一行是最新的一期,第二行是之前的一期,。。。,最后一行是第一期。
#
# 我们可以把三个数组合成一组数,就像数据集中体现的那样,并且把一组数当作一个数或者说当作一个单词。这样在预处理数据集的时候会简单一些,从索引到单词(0 -> ‘000’)和从单词到索引(‘012’-> 12)其实都是同一个数。
# ## 预测网络介绍
# 网络的输入是每一期的开奖结果,总共有1000组数,用one hot编码是一个1000维的稀疏向量:
#
# <img src="assets/OneHot.png">
# 使用one hot稀疏向量在输入层与网络第一层做矩阵乘法时会很没有效率,因为向量里面大部分都是0, 矩阵乘法浪费了大量的计算,最终矩阵运算得出的结果是向量中值为1的列所对应的矩阵中的行向量。
# <img src="assets/Matrix_multiplication.png">
#
# 这看起来很像用索引查表一样,one hot向量中值为1的位置作为下标,去索引参数矩阵中的行向量。
# 为了代替矩阵乘法,我们将参数矩阵当作一个查找表(lookup table)或者叫做嵌入矩阵(embedding matrix),将每组开奖数据所对应的数作为索引,比如“958”,对应索引就是958,然后在查找表中找第958行。
# <img src="assets/lookup_table.png">
# 这其实跟替换之前的模型没有什么不同,嵌入矩阵就是参数矩阵,嵌入层仍然是隐层。查找表只是矩阵乘法的一种便捷方式,它会像参数矩阵一样被训练,是要学习的参数。
# 下面就是我们要构建的网络架构,从嵌入层输出的向量进入LSTM层进行时间序列的学习,然后经过softmax预测出下一期的开奖结果。
# <img src="assets/model.png">
# 网络训练的代码,使用了几个trick,在下文<[`构建计算图`](#构建计算图)>和<[`训练`](#训练)>章节会做说明,<[`结论`](#结论)>在最后。
#
# 几个图表的位置:
# - [`真实值在预测值中的距离图表`](#真实值在预测值中的距离图表)
# - [`显示训练Loss`](#显示训练Loss)
# - [`显示测试Loss`](#显示测试Loss)
# - [`显示准确率`](#显示准确率)
# - [`显示预测结果和实际开奖结果`](#显示预测结果和实际开奖结果)
# ## 编码实现
# ### 实现数据预处理
# 我们需要先准备好彩票开奖记录和ID之间的转换关系。在这个函数中,创建并返回两个字典:
# - 单词到ID的转换字典: `vocab_to_int`
# - ID到单词的转换字典: `int_to_vocab`
#
# +
import numpy as np
def create_lookup_tables():
vocab_to_int = {str(ii).zfill(3) : ii for ii in range(1000)}
int_to_vocab = {ii : str(ii).zfill(3) for ii in range(1000)}
return vocab_to_int, int_to_vocab
# -
# ### 处理所有数据并保存
# 将每期结果按照从第一期开始的顺序保存到文件中。
# +
import pickle
text = load_data(data_dir)
words = [word for word in text.split()]
reverse_words = [text.split()[idx] for idx in (range(len(words)-1, 0, -1))]
vocab_to_int, int_to_vocab = create_lookup_tables()
int_text = [vocab_to_int[word] for word in reverse_words]
pickle.dump((int_text, vocab_to_int, int_to_vocab), open('preprocess.p', 'wb'))
# -
# 读取保存的数据
int_text, vocab_to_int, int_to_vocab = pickle.load(open('preprocess.p', mode='rb'))
#
def get_batches(int_text, batch_size, seq_length):
batchCnt = len(int_text) // (batch_size * seq_length)
int_text_inputs = int_text[:batchCnt * (batch_size * seq_length)]
int_text_targets = int_text[1:batchCnt * (batch_size * seq_length)+1]
result_list = []
x = np.array(int_text_inputs).reshape(1, batch_size, -1)
y = np.array(int_text_targets).reshape(1, batch_size, -1)
x_new = np.dsplit(x, batchCnt)
y_new = np.dsplit(y, batchCnt)
for ii in range(batchCnt):
x_list = []
x_list.append(x_new[ii][0])
x_list.append(y_new[ii][0])
result_list.append(x_list)
return np.array(result_list)
# ### 超参数
# +
# 训练迭代次数
epochs = 50
# 批次大小
batch_size = 32
# RNN的大小(隐藏节点的维度)
rnn_size = 512
# 嵌入层的维度
embed_dim = 512
# 序列的长度,始终为1
seq_length = 1
# 学习率
learning_rate = 0.01
# 过多少batch以后打印训练信息
show_every_n_batches = 10
save_dir = './save'
# -
# ### 构建计算图
# 使用实现的神经网络构建计算图。
#
# 使用normalized_embedding做相似度上距离的计算。
# +
import tensorflow as tf
tf.reset_default_graph()
train_graph = tf.Graph()
with train_graph.as_default():
vocab_size = len(int_to_vocab)
# 定义输入、目标和学习率占位符
input_text = tf.placeholder(tf.int32, [None, None], name="input")
targets = tf.placeholder(tf.int32, [None, None], name="targets")
lr = tf.placeholder(tf.float32)
input_data_shape = tf.shape(input_text)
# 构建RNN单元并初始化
# 将一个或多个BasicLSTMCells 叠加在MultiRNNCell中,这里我们使用2层LSTM cell
cell = tf.contrib.rnn.MultiRNNCell([tf.contrib.rnn.BasicLSTMCell(num_units=rnn_size) for _ in range(2)])
initial_state = cell.zero_state(input_data_shape[0], tf.float32)
initial_state = tf.identity(initial_state, name="initial_state")
# embed_matrix是嵌入矩阵,后面计算相似度(距离)的时候会用到
embed_matrix = tf.Variable(tf.random_uniform([vocab_size, embed_dim], -1, 1))
# embed_layer是从嵌入矩阵(查找表)中索引到的向量
embed_layer = tf.nn.embedding_lookup(embed_matrix, input_text)
# 使用RNN单元构建RNN
outputs, state = tf.nn.dynamic_rnn(cell, embed_layer, dtype=tf.float32)
final_state = tf.identity(state, name="final_state")
logits = tf.layers.dense(outputs, vocab_size)
probs = tf.nn.softmax(logits, name='probs')
cost = tf.contrib.seq2seq.sequence_loss(
logits,
targets,
tf.ones([input_data_shape[0], input_data_shape[1]]))
norm = tf.sqrt(tf.reduce_sum(tf.square(embed_matrix), 1, keep_dims=True))
normalized_embedding = embed_matrix / norm
optimizer = tf.train.AdamOptimizer(lr)
gradients = optimizer.compute_gradients(cost)
capped_gradients = [(tf.clip_by_value(grad, -1., 1.), var) for grad, var in gradients if grad is not None] #clip_by_norm
train_op = optimizer.apply_gradients(capped_gradients)
correct_pred = tf.equal(tf.argmax(probs, 2), tf.cast(targets, tf.int64))#logits <--> probs tf.argmax(targets, 1) <--> targets
accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32), name='accuracy')
# -
# ## 训练
# 在预处理过的数据上训练神经网络。
#
# 这里除了保存预测准确率之外,还保存了三类准确率:
#
# - Top K准确率:
#
# 预测结果中,前K个结果的预测准确率。
#
# - 与预测结果距离最近的Top K准确率:
#
# 先得到预测结果,使用嵌入矩阵计算与预测结果Top 1距离最近的相似度向量,取这个相似度向量中前K个结果的预测准确率。
#
# - 浮动距离中位数范围K准确率:
#
# 得到预测结果之后,计算正确结果在预测结果中的距离中位数,这个距离实际上是元素在向量中的位置与第一个元素位置的距离。这个距离数据告诉我们真正的结果在我们的预测向量中的位置在哪。每次训练之后,距离中位数都会有变化,所以是浮动的,当然也可以考虑使用众数或均值。使用中位数表示真正的结果通常会在我们的预测向量中大部分时候(平均、或者说更具代表性的)位置在哪。所以这个准确率就是以中位数为中心,范围K为半径预测准确的概率。
#
# 这里距离中位数准确率我分别在预测结果向量和与预测结果Top 1距离最近的相似度向量中都做了统计,从结果来看在相似度向量中的距离中位数准确率要稍好一些。
#
# 浮动距离中位数的概率越高,说明我们的模型训练的不好,理想情况下应该是Top K准确率越来越高,说明模型预测的越来越准确。一旦模型预测的很差,那么预测向量中一定会有一部分区域是热点区域,也就是距离中位数指示的区域,这样可以通过距离中位数来进行预测。我们使用距离中位数来帮助我们进行预测,相当于为预测做了第二套方案,一旦模型预测不准确的时候,可以尝试使用距离中位数来预测。
#
# 这三类准确率都是范围的,我们只能知道在某个范围内猜中的概率会高一些,但是到底是范围内的哪一个是准确值则很难说。
#
#
# - batches:是训练批数据
# - test_batches:是测试批数据
# - topk_acc:是预测结果的Top K准确率
# - sim_topk_acc:是与预测结果距离最近的Top K准确率
# - range_k:表示k值是一个范围,不像Top K是最开始的K个。
# - floating_median_acc_range_k:是以每次训练得出的距离中位数为中心,以范围K为半径的准确率,使用预测结果向量。
# - floating_median_sim_acc_range_k:同上,使用的是相似度向量。
# - losses:保存训练损失和测试损失
# - accuracies:保存各类准确率
# +
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
import seaborn as sns
import matplotlib.pyplot as plt
batches = get_batches(int_text[:-(batch_size+1)], batch_size, seq_length)
test_batches = get_batches(int_text[-(batch_size+1):], batch_size, seq_length)
top_k = 10
topk_acc_list = []
topk_acc = 0
sim_topk_acc_list = []
sim_topk_acc = 0
range_k = 5
floating_median_idx = 0
floating_median_acc_range_k = 0
floating_median_acc_range_k_list = []
floating_median_sim_idx = 0
floating_median_sim_acc_range_k = 0
floating_median_sim_acc_range_k_list = []
losses = {'train':[], 'test':[]}
accuracies = {'accuracy':[], 'topk':[], 'sim_topk':[], 'floating_median_acc_range_k':[], 'floating_median_sim_acc_range_k':[]}
with tf.Session(graph=train_graph) as sess:
sess.run(tf.global_variables_initializer())
saver = tf.train.Saver()
for epoch_i in range(epochs):
state = sess.run(initial_state, {input_text: batches[0][0]})
#训练的迭代,保存训练损失
for batch_i, (x, y) in enumerate(batches):
feed = {
input_text: x,
targets: y,
initial_state: state,
lr: learning_rate}
train_loss, state, _ = sess.run([cost, final_state, train_op], feed) #
losses['train'].append(train_loss)
# Show every <show_every_n_batches> batches
if (epoch_i * len(batches) + batch_i) % show_every_n_batches == 0:
print('Epoch {:>3} Batch {:>4}/{} train_loss = {:.3f}'.format(
epoch_i,
batch_i,
len(batches),
train_loss))
#使用测试数据的迭代
acc_list = []
prev_state = sess.run(initial_state, {input_text: np.array([[1]])})#test_batches[0][0]
for batch_i, (x, y) in enumerate(test_batches):
# Get Prediction
test_loss, acc, probabilities, prev_state = sess.run(
[cost, accuracy, probs, final_state],
{input_text: x,
targets: y,
initial_state: prev_state}) #
#保存测试损失和准确率
acc_list.append(acc)
losses['test'].append(test_loss)
accuracies['accuracy'].append(acc)
print('Epoch {:>3} Batch {:>4}/{} test_loss = {:.3f}'.format(
epoch_i,
batch_i,
len(test_batches),
test_loss))
#利用嵌入矩阵和生成的预测计算得到相似度矩阵sim
valid_embedding = tf.nn.embedding_lookup(normalized_embedding, np.squeeze(probabilities.argmax(2)))
similarity = tf.matmul(valid_embedding, tf.transpose(normalized_embedding))
sim = similarity.eval()
#保存预测结果的Top K准确率和与预测结果距离最近的Top K准确率
topk_acc = 0
sim_topk_acc = 0
for ii in range(len(probabilities)):
nearest = (-sim[ii, :]).argsort()[0:top_k]
if y[ii] in nearest:
sim_topk_acc += 1
if y[ii] in (-probabilities[ii]).argsort()[0][0:top_k]:
topk_acc += 1
topk_acc = topk_acc / len(y)
topk_acc_list.append(topk_acc)
accuracies['topk'].append(topk_acc)
sim_topk_acc = sim_topk_acc / len(y)
sim_topk_acc_list.append(sim_topk_acc)
accuracies['sim_topk'].append(sim_topk_acc)
#计算真实值在预测值中的距离数据
realInSim_distance_list = []
realInPredict_distance_list = []
for ii in range(len(probabilities)):
sim_nearest = (-sim[ii, :]).argsort()
idx = list(sim_nearest).index(y[ii])
realInSim_distance_list.append(idx)
nearest = (-probabilities[ii]).argsort()[0]
idx = list(nearest).index(y[ii])
realInPredict_distance_list.append(idx)
print('真实值在预测值中的距离数据:')
print('max distance : {}'.format(max(realInPredict_distance_list)))
print('min distance : {}'.format(min(realInPredict_distance_list)))
print('平均距离 : {}'.format(np.mean(realInPredict_distance_list)))
print('距离中位数 : {}'.format(np.median(realInPredict_distance_list)))
print('距离标准差 : {}'.format(np.std(realInPredict_distance_list)))
print('真实值在预测值相似向量中的距离数据:')
print('max distance : {}'.format(max(realInSim_distance_list)))
print('min distance : {}'.format(min(realInSim_distance_list)))
print('平均距离 : {}'.format(np.mean(realInSim_distance_list)))
print('距离中位数 : {}'.format(np.median(realInSim_distance_list)))
print('距离标准差 : {}'.format(np.std(realInSim_distance_list)))
# sns.distplot(realInPredict_distance_list, rug=True) #, hist=False
#plt.hist(np.log(realInPredict_distance_list), bins=50, color='steelblue', normed=True )
#计算以距离中位数为中心,范围K为半径的准确率
floating_median_sim_idx = int(np.median(realInSim_distance_list))
floating_median_sim_acc_range_k = 0
floating_median_idx = int(np.median(realInPredict_distance_list))
floating_median_acc_range_k = 0
for ii in range(len(probabilities)):
nearest_floating_median = (-probabilities[ii]).argsort()[0][floating_median_idx - range_k:floating_median_idx + range_k]
if y[ii] in nearest_floating_median:
floating_median_acc_range_k += 1
nearest_floating_median_sim = (-sim[ii, :]).argsort()[floating_median_sim_idx - range_k:floating_median_sim_idx + range_k]
if y[ii] in nearest_floating_median_sim:
floating_median_sim_acc_range_k += 1
floating_median_acc_range_k = floating_median_acc_range_k / len(y)
floating_median_acc_range_k_list.append(floating_median_acc_range_k)
accuracies['floating_median_acc_range_k'].append(floating_median_acc_range_k)
floating_median_sim_acc_range_k = floating_median_sim_acc_range_k / len(y)
floating_median_sim_acc_range_k_list.append(floating_median_sim_acc_range_k)
accuracies['floating_median_sim_acc_range_k'].append(floating_median_sim_acc_range_k)
print('Epoch {:>3} floating median sim range k accuracy {} '.format(epoch_i, np.mean(floating_median_sim_acc_range_k_list)))#:.3f
print('Epoch {:>3} floating median range k accuracy {} '.format(epoch_i, np.mean(floating_median_acc_range_k_list)))#:.3f
print('Epoch {:>3} similar top k accuracy {} '.format(epoch_i, np.mean(sim_topk_acc_list)))#:.3f
print('Epoch {:>3} top k accuracy {} '.format(epoch_i, np.mean(topk_acc_list)))#:.3f
print('Epoch {:>3} accuracy {} '.format(epoch_i, np.mean(acc_list)))#:.3f
# Save Model
saver.save(sess, save_dir) #, global_step=epoch_i
print('Model Trained and Saved')
embed_mat = sess.run(normalized_embedding)
# -
# ### 真实值在预测值中的距离图表
# 真实值在预测值相似向量中的距离数据:
# max distance : 965
# min distance : 20
# 平均距离 : 472.71875
# 距离中位数 : 562.0
# 距离标准差 : 311.20714909596387
sns.distplot(realInSim_distance_list, rug=True)
# 真实值在预测值中的距离数据:
# max distance : 946
# min distance : 7
# 平均距离 : 556.65625
# 距离中位数 : 570.0
# 距离标准差 : 274.8418419490335
sns.distplot(realInPredict_distance_list, rug=True)
# 从最后一次的距离图表来看,模型训练的还不够好,从0到1000的距离都有可能,也就是没有学到规律。
# ## 显示训练Loss
plt.plot(losses['train'], label='Training loss')
plt.legend()
_ = plt.ylim()
# ## 显示测试Loss
# 测试损失始终没有降下去。。。,epochs高一点的话会出现下降-上升-下降的波浪形曲线。
plt.plot(losses['test'], label='Test loss')
plt.legend()
_ = plt.ylim()
# ## 显示准确率
# - 测试准确率
# - Top K准确率
# - 相似度Top K准确率
# - 浮动距离中位数Range K准确率
plt.plot(accuracies['accuracy'], label='Accuracy')
plt.plot(accuracies['topk'], label='Top K')
plt.plot(accuracies['sim_topk'], label='Similar Top K')
plt.plot(accuracies['floating_median_acc_range_k'], label='Floating Median Range K Acc')
plt.plot(accuracies['floating_median_sim_acc_range_k'], label='Floating Median Sim Range K Acc')
plt.legend()
_ = plt.ylim()
# ## 显示预测结果和实际开奖结果
# 感觉从趋势上看起来还行,实际结果是一个都没有猜中 :P
#
# 有的简直错的离谱,南辕北辙
for batch_i, (x, y) in enumerate(test_batches):
plt.plot(y, label='Targets')
plt.plot(np.squeeze(probabilities.argmax(2)), label='Prediction')
plt.legend()
_ = plt.ylim()
# ## 保存参数
# 保存 `seq_length` 和 `save_dir` 在生成预测时使用。
pickle.dump((seq_length, save_dir), open('params.p', 'wb'))
# ## 加载参数
# +
import tensorflow as tf
import numpy as np
_, vocab_to_int, int_to_vocab = pickle.load(open('preprocess.p', mode='rb'))
seq_length, load_dir = pickle.load(open('params.p', mode='rb'))
# -
# ## 实现生成预测函数
# ### 获取 Tensors
def get_tensors(loaded_graph):
inputs = loaded_graph.get_tensor_by_name("input:0")
initial_state = loaded_graph.get_tensor_by_name("initial_state:0")
final_state = loaded_graph.get_tensor_by_name("final_state:0")
probs = loaded_graph.get_tensor_by_name("probs:0")
return inputs, initial_state, final_state, probs
# ### 选择号码
# 实现 `pick_word()` 函数从概率向量 `probabilities`或相似度向量`sim`中选择号码。
#
# pred_mode是选择预测的种类:
# - sim:从相似度向量Top K中选号。
# - median:从浮动距离中位数(相似度向量)Range K中选号。
# - topk:从概率向量Top K中选号。
# - max:从概率向量中选择最大概率的号码。
def pick_word(probabilities, sim, int_to_vocab, top_n = 5, pred_mode = 'sim'):
vocab_size = len(int_to_vocab)
if pred_mode == 'sim':
p = np.squeeze(sim)
p[np.argsort(p)[:-top_n]] = 0
p = p / np.sum(p)
c = np.random.choice(vocab_size, 1, p=p)[0]
return int_to_vocab[c]
elif pred_mode == 'median':
p = np.squeeze(sim)
p[np.argsort(p)[:floating_median_sim_idx - top_n]] = 0
p[np.argsort(p)[floating_median_sim_idx + top_n:]] = 0
p = np.abs(p) / np.sum(np.abs(p))
c = np.random.choice(vocab_size, 1, p=p)[0]
return int_to_vocab[c]
elif pred_mode == 'topk':
p = np.squeeze(probabilities)
p[np.argsort(p)[:-top_n]] = 0
p = p / np.sum(p)
c = np.random.choice(vocab_size, 1, p=p)[0]
return int_to_vocab[c]
elif pred_mode == 'max':
return int_to_vocab[probabilities.argmax()]
# ## 生成彩票号码
# 开始进行预测彩票了。
# - `gen_length` 作为你想生成多少期的号码。
# - `prime_word` 是前一期号码
# +
gen_length = 17
prime_word = ["623", "891", "262", "761", "900", "598", "306", "580", "243", "202"]
loaded_graph = tf.Graph()
with tf.Session(graph=loaded_graph) as sess:
loader = tf.train.import_meta_graph(load_dir + '.meta')
loader.restore(sess, load_dir)
input_text, initial_state, final_state, probs = get_tensors(loaded_graph)
normalized_embedding = loaded_graph.get_tensor_by_name("truediv:0")
gen_sentences = []
prev_state = sess.run(initial_state, {input_text: np.array([[1]])})
x = np.zeros((1, 1))
for word in prime_word:
x[0,0] = vocab_to_int[word]
probabilities, prev_state = sess.run(
[probs, final_state],
{input_text: x, initial_state: prev_state})
valid_embedding = tf.nn.embedding_lookup(normalized_embedding, probabilities.argmax())
valid_embedding = tf.reshape(valid_embedding, (1, embed_dim))
similarity = tf.matmul(valid_embedding, tf.transpose(normalized_embedding))
sim = similarity.eval()
pred_word = pick_word(probabilities, sim, int_to_vocab, 5, 'topk') # median topk max sim
gen_sentences.append(pred_word)
for n in range(gen_length):
x[0,0] = pred_word
probabilities, prev_state = sess.run(
[probs, final_state],
{input_text: x, initial_state: prev_state})
valid_embedding = tf.nn.embedding_lookup(normalized_embedding, probabilities.argmax())
valid_embedding = tf.reshape(valid_embedding, (1, embed_dim))
similarity = tf.matmul(valid_embedding, tf.transpose(normalized_embedding))
sim = similarity.eval()
pred_word = pick_word(probabilities, sim, int_to_vocab, 5, 'topk') # median topk max sim
gen_sentences.append(pred_word)
cp_script = ' '.join(gen_sentences)
cp_script = cp_script.replace('\n ', '\n')
cp_script = cp_script.replace('( ', '(')
print(cp_script)
# -
# # 结论
#
# 先从数据上说,训练的最后打印出的准确率如下:
# Epoch 49 floating median sim range k accuracy 0.01125
#
# Epoch 49 floating median range k accuracy 0.02875
#
# Epoch 49 similar top k accuracy 0.0
#
# Epoch 49 top k accuracy 0.004375
#
# Epoch 49 accuracy 0.0
# 正常的开奖概率是1‰。
#
# 准确率和相似度向量top k一样都是0,一个都没猜中。。。
#
# top k是4.3‰左右,但因为是top 10,所以实际上是0.43‰左右。
#
# 浮动中位数准确率在11.25‰~28.75‰之间,但由于这个范围range 10,所以实际上是1.125~2.875‰之间。
#
# 真没比正常开奖概率好多少。
# 从训练结果打印出的准确率,和往期开奖的相互之间的距离图都可以看得出来,想进行彩票预测实际上是不可行的。在排列三如此简单的、排列组合只有1000(样本空间已经足够小了)的等概率事件上进行预测都如此的困难,这也印证了数学的奇妙之处。都说了彩票是等概率,那么出任何一种号码都是有可能的,没有规律可言。惊不惊喜?意不意外?
# ## 新的思路
# 既然不能准确的预测,唯一能给我们提供思路的就是学习器学到的趋势,来看看下面的代码。
# - int_sentences:里面保存着上面生成的若干期号码
# - val_data:是最新几期的开奖号码,作为validate数据集
# +
int_sentences = [int(words) for words in gen_sentences]
int_sentences = int_sentences[1:]
val_data = [[103],[883],[939],[36],[435],[173],[572],[828],[509],[723],[145],[621],[535],[385],[98],[321],[427]]
plt.plot(int_sentences, label='History')
plt.plot(val_data, label='val_data')
plt.legend()
_ = plt.ylim()
# -
# 看得出来,虽然每期预测的号码不对,但是下一期号码的大概范围以及若干期号码的变化趋势学习的还可以,剩下的就要靠运气了:)
# 今天的分享就到这里,大家洗洗睡吧 :)
# +
# 913 909 997 006 278 455 016 260 349 606 271 992 049 957 970 723 571 412
# +
int_sentences = [int(words) for words in gen_sentences]
int_sentences = int_sentences[1:]
val_data = [[103],[883],[939],[36],[435],[173],[572],[828],[509],[723],[145],[621],[535],[385],[98],[321],[427]]
plt.plot(int_sentences, label='History')
plt.plot(val_data, label='val_data')
plt.legend()
_ = plt.ylim()
| cp_generation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Minicurso - Análise exploratória
#
# ## <NAME> - <NAME>
#
# Este notebook se dedica à análise exploratória de diferentes bases de dados. Serão comentadas distribuições, análise de outliers, valores ausentes, correlações, entre outros.
import numpy as np
import pandas as pd
import seaborn as sns
import scipy.stats as ss
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler, StandardScaler
# ### Descrição dos dados
# +
# Dataset do kaggle : iot (predict user transportation mode)
# link : https://www.kaggle.com/fschwartzer/tmd-dataset-5-seconds-sliding-window
# -
data = pd.read_csv('data/dataset_5secondWindow%5B1%5D.csv')
data.head()
# Dimensão dos dados
data.shape
# Ver infos sobre nossas variáveis numéricas
data.describe()
# Tipos das variáveis
data.dtypes
# Checar dados nulos
data.isna().sum()
# Variavel dependente
data.target.unique()
# Distribuição de valores
data.target.value_counts()
# Para datasets com classes desbalanceadas :
# https://www.kaggle.com/rafjaa/resampling-strategies-for-imbalanced-datasets
# ### Correlação
# Time é só uma aferição, também é uma variável que pode ser dropada
data.drop('time', axis = 1, inplace = True)
# Correlação (por matriz)
cormatrix=data.corr()
fig, ax = plt.subplots(figsize=(16, 8))
sns.heatmap(cormatrix, annot=True ,square=True)
plt.show()
# Triangular superior da matrix de correlação
fig, ax = plt.subplots(figsize=(16, 8))
upper = cormatrix.where(np.triu(np.ones(cormatrix.shape), k=1).astype(np.bool))
sns.heatmap(upper, annot=True ,square=True)
plt.show()
# Considerar somente as colunas cuja correlação abs dela com qualquer outra seja superior a cidx
cidx = 0.5
to_drop = [column for column in upper.columns if any(abs(upper[column]) > cidx)]
to_drop
# Só para ter noção da correlação (correlacao, p_valor)
coef = ss.pearsonr(data['sound#mean'], data['sound#min'])
coef = coef[0]
plt.title('Correlação', fontsize = 14)
plt.scatter(data['sound#mean'], data['sound#min'], label = str(round(coef, 3)))
plt.xlabel('sound#mean', fontsize = 14)
plt.ylabel('sound#min', fontsize = 14)
plt.xticks(fontsize = 14)
plt.yticks(fontsize = 14)
plt.legend()
plt.show()
# Dropar colunas acima para correlação
data.drop(to_drop, axis=1, inplace = True) # variavel alvo é categórica
cormatrix_less_cor = data.drop('target', axis = 1).corr()
fig, ax = plt.subplots(figsize=(16, 8))
sns.heatmap(cormatrix_less_cor, annot=True ,square=True)
plt.show()
# Da para fazer um pairplot agora que tem menos colunas
sns.set()
g = sns.pairplot(data, height = 2.5)
for ax in g.axes.flat:
ax.set_xlabel(ax.get_xlabel(), rotation = 75)
ax.set_ylabel(ax.get_ylabel(), rotation = 75)
plt.show()
data.head()
# ### Outliers
# Outliers !
for column in data.columns[:-1]:
plt.figure()
data.boxplot([column])
xdata = data.drop('target', axis = 1)
ydata = data[data.columns[-1]]
# +
# Distancia inter-quartil
Q1 = xdata.quantile(0.25)
Q3 = xdata.quantile(0.75)
IQR = Q3 - Q1
v=1.5 # ponderamento de IQR
# Remove outliers
df_no_out_x=xdata[~((xdata < (Q1 - v * IQR)) | (xdata > (Q3 + v * IQR))).any(axis=1)]
df_no_out_y=ydata[~((xdata < (Q1 - v * IQR)) | (xdata > (Q3 + v * IQR))).any(axis=1)]
# -
df_no_out_x.shape
df_no_out_y.shape
# Repete o processo
for column in df_no_out_x:
plt.figure()
df_no_out_x.boxplot([column])
# Melhorou bastante ;)
# Reconstituindo o dataframe sem outliers
df = pd.concat([df_no_out_x, df_no_out_y], axis = 1)
df.head()
# ### Normalização e padronização
# Podemos agora normalizar ou padronizar os dados (de entrada) -> trazer todo mundo para a mesma escala !
scaler_ss = StandardScaler()
scaler_mm = MinMaxScaler()
X = np.array(df[df.columns[:-1]])
Y = np.array(df[df.columns[-1]])
X
Y
X_ss = scaler_ss.fit_transform(X)
X_mm = scaler_mm.fit_transform(X)
# Olhar, por exemplo, a diferença que surgiu no android.sensor.accelerometer#mean (variavel 1)
plt.figure()
plt.title('Coluna 1 sem scaler', fontsize = 14)
sns.distplot(X[:,0])
plt.xticks(fontsize = 14)
plt.yticks(fontsize = 14)
plt.show()
# Olhar, por exemplo, a diferença que surgiu no android.sensor.accelerometer#mean (variavel 1)
plt.figure()
plt.title('Coluna 1 Standard Scaler', fontsize = 14)
sns.distplot(X_ss[:,0])
plt.xticks(fontsize = 14)
plt.yticks(fontsize = 14)
plt.show()
print('Media : ', round(X_ss[:,0].mean(), 3))
print('Desv. Padrão : ', X_ss[:,0].std())
print('Max : ', X_ss[:,0].max())
print('Min : ', X_ss[:,0].min())
# Olhar, por exemplo, a diferença que surgiu no android.sensor.accelerometer#mean (variavel 1)
plt.figure()
plt.title('Coluna 1 MaxMin', fontsize = 14)
sns.distplot(X_mm[:,0])
plt.xticks(fontsize = 14)
plt.yticks(fontsize = 14)
plt.show()
print('Media : ', round(X_mm[:,0].mean(), 3))
print('Desv. Padrão : ', round(X_mm[:,0].std(), 3))
print('Max : ', X_mm[:,0].max())
print('Min : ', X_mm[:,0].min())
# Percebam que a distribuição não mudou !
# Teste de normalidade (para modelos paramétricos)
plt.figure()
plt.title('Coluna 1 MaxMin', fontsize = 14)
sns.distplot(X_mm[:,0], fit = ss.norm)
plt.xticks(fontsize = 14)
plt.yticks(fontsize = 14)
plt.show()
ss.skew(X_mm[:,0]) # Positivo é deslocado para esquerda
ss.kurtosis(X_mm[:,0]) # Positivo é deslocado para cima
# Teste de normalidade -> quanto mais em cima da reta -> melhor (QQ plot - quantile-quantile)
fig = plt.figure()
ss.probplot(X_mm[:,0], plot=plt) # normal probaxbility plot
plt.show()
x_bc = ss.boxcox(1 + X_mm[:,0]) # Precisa ser positivo, seg. argumento é o lambda que maximiza a transformacao
# Não esquecer de transformar e somar 1 nos dados de teste também
# Olhar, por exemplo, a diferença que surgiu no android.sensor.accelerometer#mean (variavel 1)
plt.figure()
plt.title('Coluna 1 MaxMin', fontsize = 14)
sns.distplot(x_bc[0], fit = ss.norm)
plt.xticks(fontsize = 14)
plt.yticks(fontsize = 14)
plt.show()
ss.skew(x_bc[0])
ss.kurtosis(x_bc[0])
# Se aproximaram de zero ! Portanto a transformação foi, de certo modo, eficiente
# Teste de normalidade -> quanto mais em cima da reta -> melhor
fig = plt.figure()
ss.probplot(x_bc[0], plot=plt) # normal probaxbility plot
plt.show()
# Podemos adicionar essa ultima transformação aos nossos dados normalizados, então
X_mm.shape
X_bc = X_mm.copy()
for i in range(X_mm.shape[1]):
X_bc[:,i] = ss.boxcox(1 + X_mm[:,i])[0]
fig, ax = plt.subplots(2, 2, figsize = (16,8))
for i, ax in zip(range(X_bc.shape[1]), ax.flat):
ax.set_title(df.columns[i])
sns.distplot(X_bc[:,i], fit = ss.norm, ax=ax)
plt.show()
# Tem casos que isso não resolve. Solução : achar outra maneira de transformar os dados, tentar modelos paramétricos que não exijam distribuições normais ou simplesmente tentar modelos não paramétricos.
# ### Divisão treinamento e teste
# Por fim : divisão dos dados em treinamento e teste para treinar o modelo
p = 0.7
train_x, test_x, train_y, test_y = train_test_split(X_ss, Y, test_size = 1-p, random_state = 42)
# Lembrando o shape original e o que se espera
print('Dimensão original : ', df.shape, '\n')
print('Qte linhas esperadas : ')
print(str(df.shape[0])+'x0.7 = '+ str(round(0.7*df.shape[0], 2)))
print(str(df.shape[0])+'x0.3 = '+ str(round(0.3*df.shape[0], 2)))
print('Dimensão matriz de entrada de treinamento : ',train_x.shape)
print('Dimensão matriz de entrada de teste : ',test_x.shape)
print('Dimensão matriz de saída de treinamento : ',train_y.shape)
print('Dimensão matriz de saída de teste : ',test_y.shape)
# ### Valores ausentes
# +
# Dataset kaggle : titanic (passageiro sobreviveu ou não ?)
# Link : https://www.kaggle.com/c/titanic
# -
df = pd.read_csv('data/train.csv')
df.head()
df.shape
df.Survived.unique()
df.Survived.value_counts()
# Variáveis inúteis : PassengerId, Name, Ticket, Cabin (são apenas identificadores)
df.drop(['Name', 'Ticket', 'Cabin', 'PassengerId'], axis=1, inplace = True)
df.head()
# +
# Valores nulos (vamos formar um dataframe relacionando a contagem e porcentagem)
# Total por feature
total = df.isnull().sum().sort_values(ascending=False)
# Porcentagem de ausentes por feature
percent = (df.isnull().sum()/df.isnull().count()).sort_values(ascending=False)
# Concatenar em um dataframe
missing_data = pd.concat([total, percent], axis=1, keys=['Total', 'Percent'])
# -
missing_data.head()
# Como há menos de 40% de valores nulos nas features que possuem valores nulos, vamos substituir os valores nulos por não nulos. Para dados numéricos (Age) será utilizada a media e para dados categóricos (Embarked), como há apenas duas instâncias nulas as mesmas serão descartadas. Vale ressaltar que para valores categóricos, os mesmos podem ser substituídos pela moda.
# Tratar variaveis continuas
df['Age'].fillna((df['Age'].mean()), inplace=True)
# Tratar variáveis categóricas
df.dropna(inplace = True) # Os unicos ausentes são do Embarked agora
# Ou substituir pela moda (descomentar linhas abaixo e comentar linha acima antes de executar a célula)
#mode = ss.mode(df['Embarked'].dropna()).mode.item()
#df['Embarked'].fillna(mode, inplace=True)
df.isnull().sum()
df.shape # Diminuiu dois pelo drop
# ### Variáveis categóricas independentes
df.head()
df.Sex.unique()
df.Embarked.unique()
# Get dummies em Sex e Embarked !
df_dum = pd.get_dummies(df)
df_dum.head()
# Variáveis categóricas com apenas duas classes dá para deixar uma única coluna.
| analise_exploratoria.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Methods to Handle Missing Values
# 1. Imputation
# 2. Droping
# Importing the libraries
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# loading the dataset to a pandas DataFrame
dataset = pd.read_csv("Downloads/Placement_Dataset.csv")
dataset.head()
dataset.shape
dataset.isnull().sum()
# Central Tendencies
# 1. Mean
# 2. Median
# 3. Mode
# analyse the distribution of data in the salary
fig, ax = plt.subplots(figsize=(8,8))
# analyse the distribution of data in the salary
fig, ax = plt.subplots(figsize=(8,8))
sns.distplot(dataset.salary)
dataset['salary'].fillna(dataset['salary'].median(), inplace=True)
dataset.isnull().sum()
dataset.head()
# filling missing values with Mean values
dataset['salary'].fillna(dataset['salary'].mean(), inplace=True)
# filling missing values with Mode values
dataset['salary'].fillna(dataset['salary'].mode(), inplace=True)
dataset.head()
# # Dropping Method
salary_dataset = pd.read_csv("C:/Users/siddharth/Downloads/Placement_Dataset.csv")
salary_dataset.head()
salary_dataset.shape
salary_dataset.isnull().sum()
# drop the missing values
salary_dataset = salary_dataset.dropna(how='any')
salary_dataset.isnull().sum()
salary_dataset.shape
| MINI PROJECTS/Machine Learning/Handling Missing values.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Práctica 2: Aprendizaje automático
#
# __Fecha de entrega: 16 de mayo de 2021__
#
# El objetivo de esta práctica es aplicar los distintos algoritmos de aprendizaje automático disponibles en la scikit-learn [sklearn](https://scikit-learn.org/stable/) sobre varios conjuntos de datos y aprender a interpretar los resultados obtenidos. La práctica consta de 3 notebooks que se entregarán simultáneamente en la tarea de entrega habilitada en el Campus Virtual.
#
# Lo más importante en esta práctica no es el código Python, sino el análisis de los datos y modelos que construyas y las explicaciones razonadas de cada una de las decisiones que tomes. __No se valorarán trozos de código o gráficas sin ningún tipo de contexto o explicación__.
#
# Finalmente, recuerda establecer el parámetro `random_state` en todas las funciones que tomen decisiones aleatorias para que los resultados sean reproducibles (los resultados no varíen entre ejecuciones).
# Fijamos el parámetro RANDOM_STATE
RANDOM_STATE = 333
# # Parte 3: Regresión
# __Número de grupo: 15__
#
# __Nombres de los estudiantes:__
# - <NAME>
# - <NAME>
# - <NAME>
# En este notebook trabajaremos con una colección de datos de alquileres de Airbnb en Amsterdam. El objetivo de este problema es entrenar una red neuronal capaz de predecir el precio del alojamiento a partir de un conjunto de características.
# ## 1) Descripción de los datos
#
# Carga el fichero de datos `airbnb.csv` en un dataframe. Describe el conjunto de datos y trata de interpretar el significado de cada una de las variables. Estudia la distribución de precios.
#
# ### Descripción del conjunto de datos
# +
import pandas as pd
df = pd.read_csv (r'airbnb.csv')
df
# -
df.describe().transpose()
# Tenemos los datos de unos 14.998 apartamentos disponibles para alquiler en Amsterdam, y de cada uno tenemos 15 características que queremos usar para predecir el precio de cualquier apartamento de la ciudad.
#
# ### Significado de cada variable
# Para deducir el significado de algunas de estas variables nos ha servido consultar foros y páginas oficiales de Airbnb.
# - **Accomodates:** Número de huéspedes que caben o pueden quedarse en el apartamento.
# - **Bathrooms:** Número de cuartos de baño del apartamento.
# - **Bedrooms:** Número de dormitorios.
# - **Guests included:** Número de invitados que pueden traer los huéspedes.
# - **Host listings count:** Puede que el número de anfitriones
# - **Latitude & Longitude:** Coordenadas del apartamento en latitud y longitud respectivamente.
# - **Minimum nights:** Mínimo número de noches de una estancia.
# - **Number of reviews:** Número de valoraciones del apartamento.
# - **Distance centre:** Distancia al centro de la ciudad (probablemente en kilómetros).
# - **Instant bookable:** 1 si se puede reservar el apartamento inmediatamente, 0 en caso contrario.
# - **Entire home:** 1 si se alquila la casa entera, 0 en caso contrario.
# - **Private room:** 1 si se tiene una habitación privada, 0 en caso contrario.
# - **Shared room:** 1 si la habitación es compartida, 0 en caso contrario.
# - **Price:** Precio del alquiler en € (posiblemente sea el precio por noche).
#
# ### Estudio de la distribución de precios
# Como queremos predecir los precios, estudiemos su distribución:
#df.price.hist()
df['price'].plot(kind="hist", bins=20)
# Observamos una distribución asimétrica a la derecha, ya que muchos apartamentos cuestan alrededor de 130€ la noche (que es la media) pero hay valores de alquileres más separados de la media a la derecha. Es decir, la "cola" a la derecha de la media es más larga que la de la izquierda.
# Como podemos observar de la representación, la mayoría de los alquileres de nuestros datos cuestan entre 50 y 200 Euros.
# Todo esto es importante saberlo ya que es necesario comprobar si la variable sigue una distribución normal o si los valores que predecirá nuestro algoritmo serán adecuados. Como podemos observar tenemos pocos datos de alquileres de precio alto y al ser tantas variables, el algoritmo podrá predecir mal los precios en estos rangos.
# ## 2) Selección de variables
#
# Calcula los coeficientes de correlación de Pearson entre la variable de salida y el resto de variables. Crea un nuevo dataframe que contenga el precio y, además, las variables que estén relacionadas con él por un valor de correlacción de al menos 0.2 (de forma directa o inversa).
# Calculamos correlaciones 2 a 2
correl = df.corr(method='pearson')
correl
# Solo correlaciones con price superiores o iguales a 0.2
best_correls = correl['price'][abs(correl['price']) >= 0.2]
# Nos quedamos con las columnas con una correlación mínima de 0.2
final_df = df.loc[:, best_correls.index]
final_df
import seaborn as sns
##Dibujamos el diagrama de correlaciones para observar mejor los valores
Ce= set(correl.columns) - set(final_df.columns)
Ce = list(Ce)
bestC = correl.copy()
bestC = bestC.drop(Ce, axis=1)
bestC = bestC.drop(Ce, axis=0)
sns.heatmap(bestC,
xticklabels=bestC.columns,
yticklabels=bestC.columns,
cmap='RdBu_r',
annot=True,
linewidth=0.000000001)
import seaborn as sns
##Como nos importa con el precio, podemos solo ver esta columna
Ce2= list(set(final_df.columns)- set(['price']))
bestC2 = bestC.copy()
bestC2 = bestC2.drop(Ce2, axis=1)
sns.heatmap(bestC2,
xticklabels=bestC2.columns,
yticklabels=Ce2,
cmap='RdBu_r',
annot=True,
linewidth=0.000000001)
from scipy.stats import pearsonr
##Podemos ver las relaciones entre cada una de las variables importantes
## y el precio
sns.jointplot(x='accommodates', y='price', data=final_df, kind='reg')
sns.jointplot(x='bedrooms', y='price', data=final_df, kind='reg')
sns.jointplot(x='private_room', y='price', data=final_df, kind='reg')
sns.jointplot(x='bathrooms', y='price', data=final_df, kind='reg')
sns.jointplot(x='distance_centre', y='price', data=final_df, kind='reg')
sns.jointplot(x='entire_home', y='price', data=final_df, kind='reg')
sns.jointplot(x='guests_included', y='price', data=final_df, kind='reg')
# ### Análisis
# Con estos valores, podemos estimar lo que nuestro algoritmo analizará para predecir los precios:
# - Price-Accomodates: es una relación directa. Tiene cierta tendencia que a mayor número de huéspedes, mayor será el precio del alquiler.
# - Price-Bedrooms: relación directa. Tiene tendencia que a mayor número de cuartos, mayor es el precio.
# - Price-private_room: Esta relación es algo curiosa. La relación indica si no hay habitación privada, cuesta menos. Esto no tiene mucho sentido.
# - Price-bathrooms: relación directa, tiene tendencia que a mayor cantidad de baños, mayor es el precio.
# - Price-distance_centre: relacion inversa. Mientras mayor sea la distancia al centro de la ciudad, menor es el precio del alquiler.
# - Price-entire_home: relación directa. Si se alquila la casa completa, mayor será el precio del alquiler.
# - Price-guest_included: relación directa. Tiene cierta tendencia que a mayor número de huéspedes invitados, mayor será el precio.
#
# Todos estas relaciones,menos price-private_room, son cuestiones de sentido común o conocimiento que nosotros aceptamos pero que ahora están corroboradas por estos datos. Por lo cual podemos asumir que el algoritmo podría predecir más o menos bien los valores de los precios.
# ## 3) Normalización
#
# Decide si debes o no normalizar los datos. En caso afirmativo elige razonadamente entre escalarlos o estandarizarlos.
#
# Si decides escalar los datos deberás crear dos `scalers` distintos, uno para el array con la descripción de los pisos y otro para el array con el precio. Lo hacemos de este modo porque así podremos desescalar las predicciones más fácilmente.
#
# ---
#
# El MLP puede tratar con variables sin normalizar o sin estandarizar.
# Sin embargo, la normalización o estandarización de las variables disminuye el tiempo de entrenamiento y evita los “mínimos locales” en la optimización del error.
# Cuando trabajamos con redes neuronales suele ser una buena idea reescalar los datos para facilitar que el entrenamiento converja más rápidamente.
# +
import statistics as stats
# Para obtener media, mediana y moda
i=0
print(final_df.columns.values[i])
print("Media: ", stats.mean(final_df.iloc[:,i]))
print("Mediana: ", stats.median(final_df.iloc[:,i]))
print("Moda: ", stats.mode(final_df.iloc[:,i]))
df[final_df.columns.values[i]].plot.density()
# -
i=1
print(final_df.columns.values[i])
print("Media: ", stats.mean(final_df.iloc[:,i]))
print("Mediana: ", stats.median(final_df.iloc[:,i]))
print("Moda: ", stats.mode(final_df.iloc[:,i]))
df[final_df.columns.values[i]].plot.density()
i=2
print(final_df.columns.values[i])
print("Media: ", stats.mean(final_df.iloc[:,i]))
print("Mediana: ", stats.median(final_df.iloc[:,i]))
print("Moda: ", stats.mode(final_df.iloc[:,i]))
df[final_df.columns.values[i]].plot.density()
i=3
print(final_df.columns.values[i])
print("Media: ", stats.mean(final_df.iloc[:,i]))
print("Mediana: ", stats.median(final_df.iloc[:,i]))
print("Moda: ", stats.mode(final_df.iloc[:,i]))
df[final_df.columns.values[i]].plot.density()
i=4
print(final_df.columns.values[i])
print("Media: ", stats.mean(final_df.iloc[:,i]))
print("Mediana: ", stats.median(final_df.iloc[:,i]))
print("Moda: ", stats.mode(final_df.iloc[:,i]))
df[final_df.columns.values[i]].plot.density()
i=5
print(final_df.columns.values[i])
print("Media: ", stats.mean(final_df.iloc[:,i]))
print("Mediana: ", stats.median(final_df.iloc[:,i]))
print("Moda: ", stats.mode(final_df.iloc[:,i]))
df[final_df.columns.values[i]].plot.density()
i=6
print(final_df.columns.values[i])
print("Media: ", stats.mean(final_df.iloc[:,i]))
print("Mediana: ", stats.median(final_df.iloc[:,i]))
print("Moda: ", stats.mode(final_df.iloc[:,i]))
df[final_df.columns.values[i]].plot.density()
i=7
print(final_df.columns.values[i])
print("Media: ", stats.mean(final_df.iloc[:,i]))
print("Mediana: ", stats.median(final_df.iloc[:,i]))
print("Moda: ", stats.mode(final_df.iloc[:,i]))
df[final_df.columns.values[i]].plot.density()
# Como se puede ver, los valores no siguen una distribución normal en muchos casos ya que la media, mediana y moda no se parecen mucho. Por tanto debemos reescalar los datos y no estandarizarlos.
#
# Transformamos las variables de entrada:
# +
from sklearn.preprocessing import MinMaxScaler
# Creamos una copia de los datos originales para no modificarlos
df_sin_precio = final_df.copy().drop(['price'], axis=1)
# MinMaxScaler escala los datos al intervalo [0-1] sin modificar su distribución
scaler = MinMaxScaler() # lo usamos porque los puntos no siguen una distribución normal
scaler.fit(df_sin_precio)
scaled_data = scaler.transform(df_sin_precio) # cambia los datos a un sistema de coordendas scaled
scaled_data
# -
# Y transformamos la variable de salida.
# +
# Usamos una copia de los datos originales para no modificarlos
df_price = final_df.copy().price.to_numpy().reshape(-1,1)
# MinMaxScaler escala los datos al intervalo [0-1] sin modificar su distribución
scaler_precio = MinMaxScaler() # lo uso porque los puntos no siguen distribucion normal
scaler_precio.fit(df_price)
precios_scaled = scaler_precio.transform(df_price) # cambia los datos a un sistema de coordendas escaladas
precios_scaled
# -
# ## 4) Entrenamiento y selección
#
# Crea dos redes neuronales de tipo Perceptrón Multicapa:
# - La primera con una capa oculta de 200 neuronas
# - La segunda con dos capas ocultas cada una de 100 neuronas
#
# Pinta la curva de aprendizaje para cada red variando el parámetro `alpha` que controla el coeficiente de regularización L2 y determina el valor óptimo usando validación cruzada. Asegúrate de que no salen warnings indicando que no se ha alcanzado la convergencia durante el entrenamiento (basta con poner un número de max_iter suficientemente grande).
#
# ¿Alguna de las dos redes domina a la otra? ¿Por qué crees que se producen las diferencias?
# +
from sklearn.neural_network import MLPRegressor
from sklearn.model_selection import cross_val_score
from sklearn.metrics import mean_squared_error
import numpy as np
# Entrenar y validar distintas configuraciones
# Definido aquí porque lo necesitaremos también para dibujar la curva de aprendizaje
regul_param_range = 10.0 ** -np.arange(1, 7) # [0.1, 0.01, ..., 0.0000001]
# Definimos una función para entrenar un MLP dados unos datos, el target y el tamaño de las capas ocultas
def trainMLP(scaled_data, scaled_target, hidden_layer_sizes_tuple):
cv_scores_mean = []
cv_scores_std = []
for regul_param in regul_param_range:
# Aumentamos el parámetro max_iter hasta que converja
mlp = MLPRegressor(hidden_layer_sizes=hidden_layer_sizes_tuple,
activation='relu',
solver='adam',
alpha=regul_param,
batch_size=100,
learning_rate='constant',
learning_rate_init=0.001,
max_iter=500,
random_state=RANDOM_STATE)
# El error es neg_mean_squared_error el MSE en negativo!! Más pequeño peor.
scores = cross_val_score(mlp, scaled_data, scaled_target.ravel(),
cv=5, scoring='neg_mean_squared_error')
cv_scores_mean.append(scores.mean())
cv_scores_std.append(scores.std())
return cv_scores_mean, cv_scores_std
# -
# Entrenamos los 2 modelos: uno de una capa de 200 neuronas (modelo 1), y otro de 2 capas de 10 neuronas cada una (modelo 2).
# Definimos las capas ocultas
capas_ocultas1 = (200,)
capas_ocultas2 = (100,100,)
cv_scores_mean1, cv_scores_std1 = trainMLP(scaled_data, precios_scaled, capas_ocultas1)
cv_scores_mean2, cv_scores_std2 = trainMLP(scaled_data, precios_scaled, capas_ocultas2)
# Definimos una función para dibujar los resultados obtenidos y poder compararlos:
# +
import matplotlib.pyplot as plt
def plot_compare_MLP_data(cv_scores_means, cv_scores_stds, labels):
colors = ['r','g','b','c','y','k','m',]
# Suponemos que cv_scores_means y cv_scores_stds tienen la misma longitud
num_models = len(cv_scores_means)
for i in range(num_models):
# Dibujamos la línea de la exactitud media en las partes de test
plt.plot(np.log10(regul_param_range), cv_scores_means[i],
color=colors[i%len(colors)], label="Test " + labels[i])
# Dibujamos la banda de la desviación típica
lower_limit = np.array(cv_scores_means[i]) - np.array(cv_scores_stds[i])
upper_limit = np.array(cv_scores_means[i]) + np.array(cv_scores_stds[i])
plt.fill_between(np.log10(regul_param_range), lower_limit, upper_limit,
color=colors[i%len(colors)], alpha=0.2)
# Creamos el gráfico
plt.title("Curva de aprendizaje")
plt.xlabel("Regularización 10^{-X}"), plt.ylabel("negative MSE"), plt.legend(loc="best")
plt.tight_layout()
plt.show()
# -
plot_compare_MLP_data([cv_scores_mean1, cv_scores_mean2],
[cv_scores_std1, cv_scores_std2],
["Modelo 1", "Modelo 2"])
# Lo que nos interesa es minimizar el valor absoluto del MSE (es decir, que se acerque a 0) y el de la regularización L2. Por tanto, como vemos que el segundo modelo es el que menor error tiene en la gran mayoría de casos (de hecho, cuanto más pequeña es la regularización más se nota que el modelo 2 domina al modelo 1), concluimos que este debe de ser el mejor de los 2.
#
# La superioridad del modelo 2 se debe a usar 2 capas ocultas de 100 neuronas cada una en vez de una sola capa oculta de 200 neuronas. Tener más capas ocultas permite abstraer más los datos de entrada y añadir "representaciones" de los mismos, lo que se supone que ayuda a encontrar más rápidamente soluciones a partir de los datos, y esto explica que el modelo 2 se comporte mejor.
#
# Queremos el $\alpha$ que minimice el MSE positivo, pero no termina de quedar claro cuál es el mejor valor por el comportamiento estable del MSE en el rango $[10^{-3}, 10^{-5}]$
# Usamos max() porque cv_scores_mean2 tiene valores negativos, así que el máximo de estos es el más cercano a 0
alpha = 10 ** -(cv_scores_mean2.index(max(cv_scores_mean2)) + 1)
alpha
# Elejimos, por tanto, $\alpha = 10^{-4}$.
# ## 5) Medición del error
#
# Elige la mejor configuración del apartado anterior y usa la función `cross_val_predict` para realizar una predicción del valor de todos los establecimientos usando validación cruzada. ¿Cuál es el error medio del modelo en euros? ¿Crees que el modelo es suficientemente bueno?
#
# Pinta la distribución del error en euros y el diagrama de dispersión de la predicción frente al valor real. ¿El modelo comete los mismos tipos de errores en establecimientos de distintos precios? ¿Por qué crees que se producen esos errores?
# Creamos el clasificador con la regularización L2 (alpha) que consideramos mejor y las capas ocultas del
# modelo 2
mlp = MLPRegressor(hidden_layer_sizes=capas_ocultas2,
activation='relu',
solver='adam',
alpha=alpha,
batch_size=100,
learning_rate='constant',
learning_rate_init=0.001,
max_iter=500,
random_state=RANDOM_STATE)
# Estudiemos ahora el error `cross_val_predict`.
# +
from sklearn.model_selection import cross_val_predict
# Estimamos
predicts_scaled = cross_val_predict(mlp, scaled_data, precios_scaled.ravel(), cv=10)
# Desescalamos las predicciones para que estén en la escala original
predicts = scaler_precio.inverse_transform(predicts_scaled.reshape(-1,1))
# -
# Para calcular el error necesitamos trabajar con arrays, así que convertimos los precios y las predicciones.
# 'predicts' tiene una lista de listas, pero necesitamos que sea solo una, así que las concatenamos
flat_predicts = [item for sublist in predicts for item in sublist]
# Aquí obtenemos un array de los precios
prices = final_df['price'].values
# +
errors = np.abs(prices - flat_predicts) # error en cada ejemplo
mean_error = np.mean(errors)
std_error = np.std(errors)
mean_price = np.mean(prices)
#std_price = np.std(prices)
print('Precio medio:', mean_price)
#print('Desviacion típica del precio:', std_price)
print('Error medio:', mean_error)
print('Desviación típica del error:', std_error)
print('Error medio en %:', mean_error/mean_price*100)
# -
# El modelo tiene un error medio de unos 32.00€ y un error relativo del 24.60% además de una desviación típica de 32.59. Vemos que el error es bastante considerable y con una desviación típica grande, por lo que el modelo no parece ser precisamente bueno.
# Dibujamos el histograma de los errores
fig1 = plt.figure()
adf_sin_precio = plt.axes()
adf_sin_precio.hist(errors, bins=10)
plt.xlabel('Error en valor absoluto')
plt.ylabel('Frecuencia')
plt.title('Distribución del error en valor absoluto')
plt.show()
# Vemos que buena parte de los errores no son excesivamente grandes, pero una parte muy significativa de los errores más comunes son errores, por ejemplo, de alrededor de 50€ o 100€, lo que es una cantidad alta teniendo en cuenta que el precio medio es de 130.05€.
#
# Veamos una gráfica que represente las distancias entre los valores predichos y los reales. La recta en negro simboliza un error de 0.
# +
fig2 = plt.figure()
ax2 = plt.axes()
# dibujamos los errores: la distancia a la recta indica el nivel de error
ax2.scatter(prices, flat_predicts)
# Pintamos la referencia de la predicción perfecta
ax2.plot(prices, prices, color='k')
# Ponemos nombres a los ejes
plt.xlabel('Precio real')
plt.ylabel('Prediccion')
plt.show()
# -
# El modelo predice con un error considerable pero estable los precios de los apartamentos de la mitad izquierda de la gráfica, pero en la otra mitad tiene una fuerte tendencia a predecir precios mucho menores al precio real, por lo que para apartamentos más caros tiene una desviación típica muy alta.
#
# Creemos que el modelo se comporta peor con apartamentos caros porque hay muy pocos en comparación con apartamentos de precios más cercanos a la media, así que necesitaríamos una muestra más grande para conseguir predicciones más fiables para estos apartamentos.
#
# De todos modos el error sigue siendo muy alto para los apartamentos más baratos, lo que sugiere que, o bien nos faltan parámetros que ayuden a predecir el precio del alquiler, o bien que simplemente este depende más de lo que piense el dueño que de las características del apartamento.
| IA2/Practica2/P3_regresion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Project: Write a Data Sceince blog post
#
# In this project data has been fetched from the Kaggle website
#
# #### Data set details
#
# Data set : India - Trade data
#
# Data Credit : Kaggle
#
# Brief description : Trade statistics of India for export and import of commodities from 1996-2018
# +
# Import necessary libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sb
# %matplotlib inline
from sklearn.model_selection import train_test_split
from sklearn.metrics import r2_score, mean_squared_error, median_absolute_error
from sklearn.ensemble import RandomForestRegressor
# -
# ## 1. Business Understanding
#
# Being an Indian, i always would like to see what major factors are driving Indian economy and how fast the growth is happening on regular interval.
#
# Thanks to Kaggle datasets, i found the data related to Indian export and import trade value with commodity name and the corresponding value in US$ million.
#
# With this data i would like to focus on
#
# 1) What are the countries that are heavily relying on India and vice versa?
#
# 2) What are the top commodities that India getting profitted from and vice versa?
#
# 3) Progress over years (Since 2000 till 2018) on both the Import and Export front as well.
# ## 2. Data Understanding
#
# Now, let us read the data and do basic checks
# Import the data (Two files - one for Export and other for Import)
df_import = pd.read_csv("Input/2018-2010_import.csv")
df_export = pd.read_csv("Input/2018-2010_export.csv")
print('Shape of the Import data'+str(df_import.shape)+ "\n" + 'Shape of the Export data'+str(df_export.shape))
# Print first few observations
df_import.head()
# Print first few observations
df_export.head()
# Checking the data format of each column
print(str(df_import.info()) +str(df_export.info()))
# Both the files contains 5 columns information, majorly holding HSCode stands for Harmonized System code, Commodity name from HS system, Value, Country and Year of the export / Import.
#
# Value appeared to be missing in both the files (Might be due to less data coverage)
# Checking the missing values
print("- For Import data - \n"+str(df_import.isnull().sum()) +"\n\n"+ "- For Export data - \n"+str(df_export.isnull().sum()))
# Graphical representation of the missing values from Import data set
na_counts = df_import.isna().sum()
base_color = sb.color_palette()[0]
sb.barplot(na_counts.index.values, na_counts, color = base_color)
plt.xlabel('Fields')
plt.ylabel('# Missing reocrds')
plt.title('Field wise information for missing records');
# Checking the distribution of the numerical columns
print("- For Import data - \n"+str(df_import.describe()) +"\n\n"+ "- For Export data - \n"+str(df_export.describe()))
# Based on the above description of the values, we can see that Year is captured between 2010 and 2018 only
#
# Value has a wide range from 0 till 32k or 19k (for Import and Export)
#
# HS Code is ranging from 1 till 99 only
# ## 3. Prepare Data
#
# Since the data is clean and ready to use no need to drop any unused columns.
#
# But, we may need to check on areas like
#
# 1) Duplicate values
#
# 2) Unknown or missing country names
#
# 3) Missing value for Export / Import value
# Checking the duplicate values in both the files
print("- For Import data - \n"+str(df_import.duplicated().sum()) +"\n\n"+ "- For Export data - \n"+str(df_export.duplicated().sum()))
# Checking on the missing values of imported value
df_import[df_import['value'].isnull()].head()
# Checking on the missing values of exported value
df_export[df_export['value'].isnull()].head()
# Would like to check further on the country level as well how many coutries are having missing values of value of export
# Distribution of the records which are having null value at country level
pd.DataFrame(df_export[df_export['value'].isnull()]).country.value_counts()
# From the above distribution, we can see that there is one entry in country name with "UNSPECIFIED". We would need treat this as well
# Country names distribution
df_import.country.value_counts()
#Check the number of unspecified countries
print("- For Import data - \n"+str(df_import[df_import['country']=='UNSPECIFIED']['country'].count()) +"\n\n"
+ "- For Export data - \n"+str(df_export[df_export['country']=='UNSPECIFIED']['country'].count()))
# Check whether the unspecified countries hold any value associated with Export or Import
print("- For Import data - \n"+str(df_import[df_import['country']=='UNSPECIFIED']['value'].sum()) +"\n\n"
+ "- For Export data - \n"+str(df_export[df_export['country']=='UNSPECIFIED']['value'].sum()))
# Since these are having significant value, it is not good idea to drop them, rather we can use this
# We need to clean the dataset with related to value = 0 and removing the duplicates
def df_clean(df):
df = df[df['value']!=0]
df.dropna(inplace=True)
# df.year = pd.Categorical(df.year)
df.drop_duplicates(keep="first",inplace=True)
return df
# Create copies of Import and export and then apply the cleaning function
df_import1=df_import.copy()
df_export1=df_export.copy()
df_import1=df_clean(df_import1)
df_export1=df_clean(df_export1)
# Check once back with criteria
# Check the Duplicates in the data set
print("- For Import data - \n"+str(df_import1.duplicated().sum()) +"\n\n"+ "- For Export data - \n"+str(df_export1.duplicated().sum()))
# Check for the null values in the column value
print("- For Import data - \n"+str(df_import1.isnull().sum()) +"\n\n"+ "- For Export data - \n"+str(df_export1.isnull().sum()))
# ### 4. Answer Questions based on dataset
# 1) What are the countries that are heavily relying on India?
# Since the unspecified country is also being listed, remove that and show the graph
plt.figure(figsize = (20,8))
df_export1['value']=df_export1['value']/1000
df_export1[df_export1['country']!='UNSPECIFIED'].groupby('country')['value'].sum().nlargest(10).plot('bar')
plt.xlabel('Country')
plt.ylabel('Value of Export/Import (In Billion $)')
plt.title('Top 10 Countries - Export from India');
# 1) What are the countries on which India being relied upon?
plt.figure(figsize = (20,8))
df_import1.groupby('country')['value'].sum().nlargest(10).plot('bar')
plt.title('Top 10 Countries - Import to India');
# Let us take a look at the countries to which India is exporting commodities alot by checking whether India is getting import from those and if so, how much
# Create data frames to have the data at country and type of business level
country_import=df_import1.groupby('country')['value'].sum().reset_index()
country_import['Type']='Import'
country_export=df_export1.groupby('country')['value'].sum().reset_index()
country_export['Type']='Export'
country_wise=country_export.append(country_import)
country_wise['Type'].value_counts()
val_df=country_wise.groupby(['country', 'Type'])['value'].sum().reset_index()
export_ctr=df_export1[df_export1['country']!='UNSPECIFIED'].groupby('country')['value'].sum().nlargest(10).reset_index()
export_ctr
exp_imp_top=pd.merge(val_df,export_ctr,on='country',how='inner')
exp_imp_top['value_x']=exp_imp_top['value_x']/1000
exp_imp_top
# Plot a Bar chart to see the how the top 10 export nations are doing business with India related to Import
plt.figure(figsize = [14, 5])
exp_imp_top=exp_imp_top.sort_values(['value_y','Type'],ascending=[False,True])
sb.barplot(x='country', y='value_x', hue='Type', data=exp_imp_top,palette="Set2");
plt.legend(loc = 2, ncol = 1, framealpha = 1, title = 'Type of Business')
plt.xlabel('Country')
plt.ylabel('Value of Export/Import (In Billion $)')
plt.title('India - Export and Import');
# From the above bar graph we can notice that, Even though India is exporting most of it's commodities to these countries, at the same time India is relying heavily on countries like China, USA and UAE & Saudi arabia.
#
# Further, we can check what is that India getting importing from China and Saudi Arabia
# Look at china data
china = df_import1[df_import1['country']=='CHINA P RP']
china.head()
china.groupby(['year'])['value'].sum()
china_c=china.groupby(['Commodity'])['value'].sum().reset_index()
china_c['value']=china_c['value']/1000
china_c.sort_values('value',ascending=False)
plt.figure(figsize = (10,7))
china_c.groupby('Commodity')['value'].sum().nlargest(5).sort_values().plot('barh')
plt.title('Top 10 Commodities (HSA) - Import from China');
# +
# Look at Saudi Arabia data
sab = df_import1[df_import1['country']=='SAUDI ARAB']
sab.head()
sab_c=sab.groupby(['Commodity'])['value'].sum().reset_index()
sab_c['value']=sab_c['value']/1000
sab_c.sort_values('value',ascending=False)
plt.figure(figsize = (10,7))
sab_c.groupby('Commodity')['value'].sum().nlargest(5).sort_values().plot('barh')
plt.title('Top 10 Commodities (HSA) - Import from Saudi Arabia');
# -
# 2) What are the top commodities that India getting profitted from?
plt.figure(figsize = (10,7))
df_export1.groupby('Commodity')['value'].sum().nlargest(10).sort_values().plot('barh')
plt.title('Top 10 Commodities (HSA) - Export from India');
plt.figure(figsize = (10,7))
df_import1.groupby('Commodity')['value'].sum().nlargest(10).sort_values().plot('barh')
plt.title('Top 10 Commodities (HSA) - Import to India');
# Based on the above charts, we can see that top exported commodities are as well part of the top imported commodities as well.
# But the trade value is the difference between the Export and Import of these top commodities.
#
# Vehicle other than railway or tramway and Cotton are the goods that are not part of top 10 import categories.
# 3) Progress over years (Since 2010 till 2018) on both the Import and Export front as well.
# Calculate the export and import value at year level and merge the data
year_wise=pd.merge(((df_import1.groupby('year')['value'].sum().reset_index()).rename(columns={'value':'Import_Value'})),
(((df_export1.groupby('year')['value'].sum().reset_index()).rename(columns={'value':'Export_Value'}))))
year_wise
year_wise['Import_Value'] = year_wise['Import_Value'] / 1000
year_wise['Export_Value'] = year_wise['Export_Value'] / 1000
year_wise
plt.figure(figsize=(10, 8))
plt.plot(year_wise['year'], year_wise['Import_Value'], 'g', label = 'Import')
plt.plot(year_wise['year'], year_wise['Export_Value'], 'b', label = 'Export')
plt.xlabel('Year of Export / Import'); plt.ylabel('Value of Export/Import (In Billion $)');
plt.title('India - Export and Import')
plt.legend();
# With the help of above line chart, we can able to clearly see that India is importing goods which are having value larger than that of the export goods.
#
# With 2014 being the election year there has been dip in the export but post that we can see clear upward trends
# Create data frames to have the data at country and type of business level
year_to_import=df_import1.groupby(['year','country'])['value'].sum().reset_index()
year_to_import['Type']='Import'
year_to_export=df_export1.groupby(['year','country'])['value'].sum().reset_index()
year_to_export['Type']='Export'
year_to_wise=year_to_export.append(year_to_import)
year_to_wise['Type'].value_counts()
year_to_wise_top=pd.merge(year_to_wise,export_ctr,on='country',how='inner')
year_to_wise_top['value_x']=year_to_wise_top['value_x']/1000
year_to_wise_top
year_to_wise_top=year_to_wise_top.sort_values(['value_y','Type'],ascending=[False,True])
plt.figure(figsize=(20,9))
ax = sb.boxplot(x="country", y="value_x", hue="Type",data=year_to_wise_top, palette="Set2")
plt.show()
| Project_1_Blog Post.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="hMNVigZDu9rT"
# # Install packages and mount Google-Drive
#
# Run the cell -> click the link -> Login with @gmail.com e-mail account.
# + [markdown] id="h4TLEuOWo8eM" outputId="7a7d4660-dd18-42f8-acdf-1e9f960d4142"
# # # %%capture
# # # !pip install pandas matplotlib google-cloud-storage
#
# import os
# from google.colab import drive
# drive.mount('/content/drive')
#
# # Point environment variable `GOOGLE_APPLICATION_CREDENTIALS` to
# # location of service account file 'dtu-course-02456-students.json'.
# os.environ['GOOGLE_APPLICATION_CREDENTIALS'] = "/content/drive/My Drive/Woodsense/Tech/Software/Deep Learning Course DTU/Students Folder/gcp-service-accounts/dtu-course-02456-students.json"
# + id="3iYPb7WOkXRB" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1609285430210, "user_tz": -60, "elapsed": 20231, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}} outputId="8cf88741-b90d-4c7d-98ae-faa840def2df"
# %%capture
# !pip install pandas google-cloud-storage
import os
from google.colab import drive
drive.mount('/content/drive')
# Point environment variable `GOOGLE_APPLICATION_CREDENTIALS` to
# location of service account file 'dtu-course-02456-students.json'.
os.environ['GOOGLE_APPLICATION_CREDENTIALS'] = "/content/drive/My Drive/Woodsense/Tech/Software/Deep Learning Course DTU/Students Folder/gcp-service-accounts/dtu-course-02456-students.json"
# + [markdown] id="ZZwDr-yVe4ME"
# # Load dataset
#
# The dataset is simply time series grouped by sensor ID's containing the parameters described in the Readme document.
#
# For further questions please use the Slack channel `#p10-woodsense` or email <EMAIL>.
# + id="-Z5NY-dVbk1n" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1609285432360, "user_tz": -60, "elapsed": 22371, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}} outputId="9d251534-47cd-4556-f202-d4a58877e50e"
import io
import json
import seaborn as sns
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
import matplotlib
import pandas as pd
#df = pd.read_csv('/content/drive/My Drive/Colab Notebooks/DL/Woodsense/data/woodsense-sensor-data-2020-10-27.csv')
df = pd.read_csv('/content/drive/My Drive/WoodSense/data/woodsense-sensor-data-2020-11-30-cleaned.csv') #Tue destination
df.timestamp = pd.to_datetime(df.timestamp, errors='coerce')
#Count the length of the timeseres whihc might be used later
sensor_seq_len = df.sensor_id.value_counts()
df = df.drop(['weather_wind_min', 'ohms'], axis=1)
df['tod_sin'] = np.sin(df['timestamp'].dt.hour / 24 * 2 * np.pi)
df['tod_cos'] = np.cos(df['timestamp'].dt.hour / 24 * 2 * np.pi)
df['doy_sin'] = np.sin(df['timestamp'].dt.dayofyear / 365 * 2 * np.pi)
df['doy_cos'] = np.cos(df['timestamp'].dt.dayofyear / 365 * 2 * np.pi)
print(len(df))
# + [markdown] id="HWfD5QNXLrCb"
# #Data analysis
# + id="xTTSE8CFT71p" executionInfo={"status": "ok", "timestamp": 1609285435634, "user_tz": -60, "elapsed": 25629, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}}
#Imports
import torch
import torch.nn as nn
import torch.nn.functional as F
import torchvision.datasets as datasets
from torch.utils.data import DataLoader
import torchvision.transforms as transforms
from torch.utils.data.sampler import SubsetRandomSampler
from torchvision.transforms import ToTensor
# + id="gyJkZBnqkDK7" executionInfo={"status": "ok", "timestamp": 1609285435635, "user_tz": -60, "elapsed": 25627, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}}
# Generators
import random
def sensor_split(data, test_size = 0.15, test_sensors = None):
'''
Splits the sensors into two groups, test and train.
test_size = size of test set
test_sensors = if True, sensors are given as inputs.
'''
sensors = data.sensor_id.unique().tolist()
n_sensors = len(sensors)
k = int(test_size*n_sensors)
random.seed = 42
if test_sensors == None:
test_sensors = random.choices(sensors,k=k)
train = data[~data.sensor_id.isin(test_sensors)]
test = data[data.sensor_id.isin(test_sensors)]
train = train.sort_values(['sensor_id','timestamp'])
test = test.sort_values(['sensor_id','timestamp'])
return train, test
# + id="qyRfHhKLkDK8" executionInfo={"status": "ok", "timestamp": 1609285435635, "user_tz": -60, "elapsed": 25624, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}}
#Split sensors
test_sensors = [20, 25, 26, 27, 50, 51]
validation_sensors = [4, 5, 10, 11, 12, 34, 37, 41, 45, 52, 55, 65]
df_train, df_test = sensor_split(df, 0.20, test_sensors = test_sensors)
df_train, df_val = sensor_split(df_train, 0.20, test_sensors = validation_sensors)
# + id="a9nAUbzfkDK-" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1609285435636, "user_tz": -60, "elapsed": 25622, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}} outputId="02075505-d6e2-4d7b-bd16-f6e16db5f159"
df_train['sensor_id'].unique()
# + id="g5e1zyBskDK_" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1609285435636, "user_tz": -60, "elapsed": 25606, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}} outputId="a302fc81-2509-421e-cb11-204311e285a6"
df_test['sensor_id'].unique()
# + colab={"base_uri": "https://localhost:8080/"} id="NjjJLBACyGMS" executionInfo={"status": "ok", "timestamp": 1609285435637, "user_tz": -60, "elapsed": 25591, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}} outputId="76918990-86f8-419f-d1a3-7582fbf4e30e"
df_val['sensor_id'].unique()
# + [markdown] id="d-AqaEabOBmr"
# ## Data loader
# Here we create the data loader
# + id="093jm0jpq54a" executionInfo={"status": "ok", "timestamp": 1609285435637, "user_tz": -60, "elapsed": 25573, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}}
import numpy as np
def standardise_depant(df_data, df_mean_std):
'''
Standardise data based on the inputs
df_data = dataframe
df_mean_std = dataframe containing the mean and std for each feature'''
df_data = df_data.drop([ 'sensor_id'], axis=1)
for index, row in df_mean_std.iterrows():
measure = row['measure']
mean = row['mean']
std = row['std']
df_data[measure] = (df_data[measure]-mean)/std
return df_data
# + id="98dPEXXdOPja" executionInfo={"status": "ok", "timestamp": 1609285435638, "user_tz": -60, "elapsed": 25571, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}}
import numpy as np
def create_inout_sequences(input_data, tw, target_size = 3, step_size = 1, keep_sensor_id = False, df_mean_and_std = None):
'''Function that generates sequences based on the current time steps
input_data = The data from Woodsens
tw = timewindow/timestemp we want to look back on
target_size = how many hours we want to predict // For anomaly detection this would most likely be 1
step_size = how many jumps we want to create between each data sample
keep_sensor_id = If True the sensor_id's are not dropped
df_mean_and_std = if given use the mean and std to standardize
'''
inout_seq = []
label = []
train = []
for sensors in df['sensor_id'].unique():
data = input_data[input_data['sensor_id'] == sensors]
L = len(data)
data = data.drop(['timestamp'], axis=1)
data = standardise_depant(data, df_mean_and_std)
label_data = data.drop(['tod_sin', 'tod_cos', 'doy_sin', 'doy_cos', 'weather_humidity', 'weather_pressure','weather_temp_dew', 'weather_temp_dry','weather_precip_past10min', 'weather_wind_max', 'weather_wind_speed'], axis=1)
data = data.astype(np.float32).to_numpy().tolist()
data = torch.FloatTensor(data)
label_data = label_data.astype(np.float32).to_numpy().tolist()
label_data = torch.FloatTensor(label_data)
for i in range(0,L-tw-target_size, step_size):
train_seq = data[i:i+tw]
train_label = label_data[i+tw:i+tw+target_size]
label.append(train_label)
train.append(train_seq)
if keep_sensor_id == False:
inout_seq.append((train_seq ,train_label))
else:
inout_seq.append((sensors, train_seq ,train_label))
return train, label, inout_seq
# + colab={"base_uri": "https://localhost:8080/", "height": 359} id="6rahcPZyucCn" executionInfo={"status": "ok", "timestamp": 1609285435638, "user_tz": -60, "elapsed": 25566, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}} outputId="1421e4af-337f-4a55-9eaf-5869abc6f4e7"
#Determine mean and std's for the training set
param_standard = df.columns[2:12]
mean_and_std = []
for param in param_standard:
mean = df_train[param].mean()
std = df_train[param].std()
mean_and_std.append((param, mean, std))
df_mean_std = pd.DataFrame(mean_and_std)
df_mean_std.columns = ['measure', 'mean', 'std']
df_mean_std
# + id="bIrBMIyHkDLF" executionInfo={"status": "ok", "timestamp": 1609285437584, "user_tz": -60, "elapsed": 27495, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}}
#Sequence generation
window = 24
train_seq, train_label, train_inout_seq = create_inout_sequences(df_train, tw = window, target_size = 1, step_size = 1, keep_sensor_id = False, df_mean_and_std = df_mean_std)
val_seq, val_label, val_inout_seq = create_inout_sequences(df_val, tw = window, target_size = 1, step_size = 1, keep_sensor_id = True, df_mean_and_std = df_mean_std)
# + id="RQikYgcosTVi" executionInfo={"status": "ok", "timestamp": 1609285437584, "user_tz": -60, "elapsed": 27492, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}}
import torch
class Dataset(torch.utils.data.Dataset):
'Characterizes a dataset for PyTorch'
def __init__(self, data, labels):
'Initialization'
self.labels = labels
self.data = data
def __len__(self):
'Denotes the total number of samples'
return len(self.data)
def __getitem__(self, index):
'Generates one sample of data'
# Load data and get label
X = self.data[index]
y = self.labels[index]
return X, y
# + id="dtJfZGzdsZvU" executionInfo={"status": "ok", "timestamp": 1609285437585, "user_tz": -60, "elapsed": 27490, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}}
# Generators
batch_size = 32
training_set = Dataset(train_seq, train_label)
training_generator = torch.utils.data.DataLoader(training_set, batch_size=batch_size, shuffle=True, num_workers=0)
validation_set = Dataset(val_seq, val_label)
validation_generator = torch.utils.data.DataLoader(validation_set, num_workers=0)
# + [markdown] id="OIPXkTJMzqU_"
# ### Model
# + id="RkWcl-Rv0GgA" executionInfo={"status": "ok", "timestamp": 1609285437585, "user_tz": -60, "elapsed": 27486, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}}
class Network_2layers(nn.Module):
def __init__(self, features=14, tw=24, target_features=3, target_size=1, filters1=32, filters2=32, kernel1=3, kernel2=3):
super(Network_2layers, self).__init__()
padding1 = int(kernel1/2)
padding2 = int(kernel2/2)
self.target_features = target_features
self.target_size = target_size
dimensions_conv_out = int(tw/(2*2))
self.convolutional = nn.Sequential(
nn.Conv1d(features, filters1, kernel1, stride=1, padding=padding1), # 32x24
#nn.Dropout2d(0.5), #50 % probability
nn.MaxPool1d(2, stride=2), #32x12
nn.ReLU(),
nn.Conv1d(filters1, filters2, kernel2, stride=1, padding=padding2), #32x12
#nn.Dropout2d(0.2), #50 % probability
nn.MaxPool1d(2, stride=2), # 32 x 6
nn.ReLU()
)
self.fully_connected = nn.Sequential(
nn.Linear(filters2*dimensions_conv_out, target_features*target_size),
#nn.Dropout(p=0.2),
#nn.ReLU(),
#nn.Linear(50, 4)
)
def forward(self, x):
x = torch.transpose(x, 2, 1)
x = self.convolutional(x)
#reshape x so it becomes flat, except for the first dimension (which is the minibatch)
x = x.view(x.size(0), -1)
x = self.fully_connected(x)
x_shape = x.shape
x = torch.reshape(x, (x_shape[0], self.target_features, self.target_size) )
return x
# + id="DfeliDUjBD-o" executionInfo={"status": "ok", "timestamp": 1609285437586, "user_tz": -60, "elapsed": 27484, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}}
X, Y = next(iter(training_generator))
# + colab={"base_uri": "https://localhost:8080/"} id="YSY_Ct3pNBu6" executionInfo={"status": "ok", "timestamp": 1609285437839, "user_tz": -60, "elapsed": 27735, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}} outputId="fbe492df-80d3-4cf5-c77f-912e14582efe"
print(Y.shape)
print(X.shape)
net = Network_2layers(features=14, tw=24, target_features=3, target_size=1, filters1=128, filters2=128, kernel1=11, kernel2=11)
output = net(X)
print(output.shape)
criterion = nn.L1Loss()
# + colab={"base_uri": "https://localhost:8080/"} id="uNTeVR6Jdmeo" executionInfo={"status": "ok", "timestamp": 1609285437840, "user_tz": -60, "elapsed": 27721, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}} outputId="9cf4180d-dcf5-4a5f-8fdf-d503f3731dd3"
print(output.shape)
print(torch.transpose(Y, 2, 1).shape)
# + id="nu4SJ90mFEri" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1609285437840, "user_tz": -60, "elapsed": 27706, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}} outputId="fb85cba7-aa3b-49b9-cc3e-99c5379d51ee"
#Usa GPU
if torch.cuda.is_available():
print("The code will run on GPU.")
else:
print("The code will run on CPU. Go to Edit->Notebook Settings and choose GPU as the hardware accelerator")
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
# + [markdown] id="hSQiTNsCzylw"
# ### Model traning
# + id="JFHz6zJX_3qt" executionInfo={"status": "ok", "timestamp": 1609285448021, "user_tz": -60, "elapsed": 37871, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}}
#We define the training as a function so we can easily re-use it.
from tqdm.notebook import tqdm
import torch.optim as optim
net = Network_2layers(features=14, tw=24, target_features=3, filters1=32, filters2=32, kernel1=3, kernel2=3)
net.to(device)
optimizer = optim.Adam(net.parameters(), lr=0.001)
criterion = nn.L1Loss()
def train(model, optimizer, num_epochs=10):
out_dict = {
'train_loss': [],
'test_loss': []}
for epoch in tqdm(range(num_epochs), unit='epoch'):
model.train()
#For each epoch
train_loss = []
for minibatch_no, (data, target) in enumerate(training_generator): # tqdm(enumerate(training_generator), total=len(training_generator)):
#FOR GPU
data, target = data.to(device), target.to(device)
#Zero the gradients computed for each weight
optimizer.zero_grad()
#Forward pass your image through the network
model.train()
output = model(data)
#Compute the loss
target = torch.transpose(target, 2, 1)
loss = criterion(output, target)
#Backward pass through the network
loss.backward()
#Update the weights
optimizer.step()
train_loss.append(loss.item())
#Compute how many were correctly classified
#Comput the test accuracy
test_loss = []
model.eval()
for data, target in validation_generator: #tqdm(enumerate(validation_generator), total=len(validation_generator)):
data, target = data.to(device), target.to(device)
target = torch.transpose(target, 2, 1)
with torch.no_grad():
model.eval()
output = model(data)
test_loss.append(criterion(output, target).cpu().item())
out_dict['train_loss'].append(np.mean(train_loss))
out_dict['test_loss'].append(np.mean(test_loss))
print("Loss train: ", np.mean(train_loss), "\t test: ", np.mean(test_loss))
return out_dict
# + id="4-wAsfBnD5e4" colab={"base_uri": "https://localhost:8080/", "height": 236, "referenced_widgets": ["ae069edfaaff4bc3b01512249b135dad", "de1795f0aa0b4ac38e86bec34a3a63d1", "488e34fffdd64e55a28e5d782e7f2d87", "5c487198adb24e3eaa0f085b9a78529a", "6e07d86e9e8340378bc45c255d5616a7", "82663da2759c422faa10190e0548c628", "a6c26f474aaf4d91a529c6f638ef6ba3", "ceda371ce5064cb8be4b896d2b9dbf83"]} executionInfo={"status": "ok", "timestamp": 1609285558853, "user_tz": -60, "elapsed": 148699, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}} outputId="e6520675-cbdc-4e02-a507-a82ae0bd8490"
D = train(net, optimizer, num_epochs=10)
test_loss = D["test_loss"]
train_loss = D["train_loss"]
# + [markdown] id="RiFpHTB1pxKq"
# ## Visualization on test set
# Functions for vizualtsation and determining treshold on the testset
# + id="Dk_mkpSz7FGc" executionInfo={"status": "ok", "timestamp": 1609285559234, "user_tz": -60, "elapsed": 149065, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}}
tw = train_seq[0].shape[0] # Get size of window
parameters_predicted = ['temp', 'humid', 'moist']
#parameters_predicted = [ 'moist']
# Convert test data to sequences
test_seq, test_label, test_inout_seq = create_inout_sequences(df_test, tw = window, target_size = 1, step_size = 1, keep_sensor_id = True, df_mean_and_std = df_mean_std)
# + id="0Q9hg8Yi6Z9v" executionInfo={"status": "ok", "timestamp": 1609285559616, "user_tz": -60, "elapsed": 149445, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}}
def euclidean_dist(x, y): # Calculates euclidean dist
if (type(x) == float):
return abs(x-y)
return np.sqrt(sum((y-x)**2))
# + id="NS8qLpO2qRLq" executionInfo={"status": "ok", "timestamp": 1609285571210, "user_tz": -60, "elapsed": 161036, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}}
# Use trained network to predict next point, and compute loss
net = net.to(device)
combined_data = [];
for i in range(len(test_inout_seq)):
sensor, X, y = test_inout_seq[i]
X, y = X.to(device), y.to(device)
with torch.no_grad():
X = X.reshape(1, tw, 14)
output = net(X)
#y = torch.transpose(y, 1, 0)
y2= y
y = y.reshape(output.shape)
loss = criterion(y, output) # same loss as used in training
loss = float(loss.cpu().detach().numpy())
output = output.cpu().detach().numpy()
output = np.squeeze(output)
y = y.cpu().detach().numpy()
y = np.squeeze(y)
if y.size == 1:
euc_dist = abs(y-output);
temp_list = [sensor, loss, euc_dist, euc_dist, y, output]
else:
euc_dist = euclidean_dist(y,output) # euclidean distance
temp_list = [sensor, loss, euc_dist] # list to hold sensor id, losses, values
for y_i, output_i in zip(y, output): # iterate through the predicted features
temp_list.append(abs(output_i-y_i))
#print(y)
y = torch.transpose(y2, 1, 0)
y = y.cpu().detach().numpy()
#print(y)
temp_list.extend(y)
temp_list.extend(output)
combined_data.append(temp_list)
# + id="oc3S6Liukupu" colab={"base_uri": "https://localhost:8080/", "height": 204} executionInfo={"status": "ok", "timestamp": 1609285571211, "user_tz": -60, "elapsed": 161033, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}} outputId="65d415c2-96fe-45bd-e996-c0bb776a62d9"
# Convert to dataframe
column_names = ['sensor_id', 'loss', 'euclidean_dist']
for i, param in enumerate(parameters_predicted):
column_names.append('loss_' + param)
for i, param in enumerate(parameters_predicted):
column_names.append('true_' + param)
for i, param in enumerate(parameters_predicted):
column_names.append('pred_' + param)
df_combined = pd.DataFrame(combined_data)
df_combined.columns = column_names
df_combined.head()
# + id="Yyx46FENJzVD" executionInfo={"status": "ok", "timestamp": 1609285571211, "user_tz": -60, "elapsed": 161018, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}}
#Create temporary df for plotting
tmp_df = df_combined[df_combined['sensor_id'] == 25]
df_new = pd.concat([tmp_df['sensor_id'],tmp_df['euclidean_dist'].explode(),
tmp_df['loss_temp'].explode(),
tmp_df['loss_humid'].explode(),
tmp_df['loss_moist'].explode(),
tmp_df['true_temp'].explode(),
tmp_df['true_humid'].explode(),
tmp_df['true_moist'].explode(),
tmp_df['pred_temp'].explode(),
tmp_df['pred_humid'].explode(),
tmp_df['pred_moist'].explode()],
axis=1, keys=['sensor_id','euclidean_dist','loss_temp','loss_humid','loss_moist','true_temp','true_humid','true_moist','pred_temp','pred_humid', 'pred_moist'])
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="MhJLlMuuR7Rj" executionInfo={"status": "ok", "timestamp": 1609285573077, "user_tz": -60, "elapsed": 162879, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}} outputId="ef8745e0-4fe3-4a21-ab9c-f8b7c9ee45a5"
import plotly.graph_objects as go
sensor = 25
fig = go.Figure()
fig.add_trace(go.Scatter(y=df_new['true_moist'][df_new['sensor_id'] == sensor],
mode='lines',
name='True'))
fig.add_trace(go.Scatter(y=df_new['pred_moist'][df_new['sensor_id'] == sensor],
mode='lines',
name='DeepAnT'))
fig.show()
# + [markdown] id="daHk2tjZT78l"
# This part only works for a target of size 1
# + id="WKKC4zl8T6Kl" executionInfo={"status": "ok", "timestamp": 1609285573078, "user_tz": -60, "elapsed": 162865, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}}
from itertools import compress
def plot_losses(idx_worst, idx_best, df, window = 24, param = 'temp'):
'''
The function takes as input two array of indexes, that define where the 5 best
and the 5 worst losses were obtained over a 24 hour period. The function then
plots the 24 hour windows for a chosen parameter.
'''
n_subplots = len(idx_best)
fig, ax = plt.subplots(1, n_subplots, figsize=(25, 5), sharey = True)
for count, idx in enumerate(idx_worst):
sensor_id = df.iloc[int(idx)].sensor_id
ax[count].plot(range(window), df['true_'+param][int(idx) : int(idx)+window], label = 'True')
ax[count].plot(range(window), df['pred_'+param][int(idx) : int(idx)+window], label = 'Predicted')
#ax[count].title('Sensor')
fig.suptitle('5 worst losses', fontsize=24)
for ax in fig.axes:
ax.grid()
ax.legend()
plt.show()
fig, ax = plt.subplots(1, n_subplots, figsize=(25, 5), sharey = True)
for count, idx in enumerate(idx_best):
sensor_id = df.iloc[int(idx)].sensor_id
ax[count].plot(range(window), df['true_'+param][int(idx) : int(idx)+window], label = 'True')
ax[count].plot(range(window), df['pred_'+param][int(idx) : int(idx)+window], label = 'Predicted')
#ax[count].title('Sensor')
fig.suptitle('5 best losses', fontsize = 24)
for ax in fig.axes:
ax.grid()
ax.legend()
plt.show()
def plot_sensor(df):
'''
Makes a plot of predicted and actual values for all features for all sensor
over the entire period
'''
n_subplots = len(df['sensor_id'].unique())
# Convert parameters_predicted to corresponding plot titles
all_params = ['temp', 'humid', 'ohms', 'moist']
all_titles = ['Temperature', 'Humidity', 'Ohms', 'Moisture']
titles = list(compress(all_titles, [x in parameters_predicted for x in all_params]))
for idx, param in enumerate(parameters_predicted):
fig, ax = plt.subplots(1, n_subplots, figsize=(25, 5), sharey = True)
for count, sensor in enumerate(df['sensor_id'].unique()):
ax[count].plot(df['true_'+param][df['sensor_id'] == sensor], label = 'True')
ax[count].plot(df['pred_'+param][df['sensor_id'] == sensor], label = 'Predicted')
ax[count].set_title('Sensor '+ str(int(sensor)))
fig.suptitle(titles[idx], fontsize=24)
for ax in fig.axes:
ax.grid()
ax.legend()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 904} id="1ZFvHoWSUCsy" executionInfo={"status": "ok", "timestamp": 1609285575507, "user_tz": -60, "elapsed": 165290, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}} outputId="4f79e5f7-a3cc-4b91-e786-021e0d45f16c"
plot_sensor(df_combined)
# + [markdown] id="o6XZ8DeO2MDY"
# To plot the five best and the five worst predictions, we first five the best and worst 24 hour windows of predictions. We calculate the average Euclidean distance over a 24 hour period, and choose the windows with highest and lowest losses.
# + id="VgWbNCQ8KKVY" executionInfo={"status": "ok", "timestamp": 1609285575856, "user_tz": -60, "elapsed": 165623, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}}
# Conmpute mean loss for each sensor for each 24 hour window
losses = []; # list of losses
for sensor in df_combined['sensor_id'].unique():
df_sensor = df_combined[df_combined['sensor_id'] == sensor]
length = len(df_sensor)
for i in range(int(length/24)):
idx = df_sensor.iloc[i*24].name # row in dataframe
loss = df_sensor[i*24:(i+1)*24]['loss'].mean()
euc_dist = df_sensor[i*24:(i+1)*24]['euclidean_dist'].mean()
temp_list = [idx, loss, euc_dist]
for param in parameters_predicted:
loss_param = df_sensor[i*24:(i+1)*24]['loss_'+param].mean()
temp_list.append(loss_param)
losses.append(temp_list)
# + id="aD1lH2ErPC8M" colab={"base_uri": "https://localhost:8080/", "height": 609} executionInfo={"status": "ok", "timestamp": 1609285577289, "user_tz": -60, "elapsed": 167052, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}} outputId="f9835a02-e0f2-438a-a752-73e2272151af"
# Determine the five best and five worst windows
loss_array = np.array(losses)
# sort loss_array based on one of the average losses
# sort by 1 = criterion function, 2 = euclidean dist, 3 = temp, 4 = humid, 5 = moist
loss_array = loss_array[np.argsort(loss_array[:, 1])]
n = 5
idx_worst = loss_array[-n:,0]
idx_best = loss_array[0:n,0]
plot_losses(idx_worst, idx_best, df_combined, window = 24, param = 'moist')
# + [markdown] id="5hvi6x9F20RZ"
# In order to use the model for anomaly detection, we determine the 99th percentile of the Euclidean distances on the validation set. We also determine the 99th percentile for the absolute error on for each of the features.
# + id="nTBxmszSc33S" executionInfo={"status": "ok", "timestamp": 1609285584435, "user_tz": -60, "elapsed": 174183, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}}
#Use trained network to predict next point, and compute loss
net = net.to(device)
losses_train = [];
for i in range(len(val_inout_seq)):
s, X, y = val_inout_seq[i]
X, y = X.to(device), y.to(device)
with torch.no_grad():
X = X.reshape(1, tw, 14)
output = net(X)
y = y.reshape(output.shape)
loss = criterion(y, output)
loss = float(loss.cpu().detach().numpy())
output = output.cpu().detach().numpy()
output = np.squeeze(output)
y = y.cpu().detach().numpy()
y = np.squeeze(y)
if y.size == 1:
euc_dist = abs(y-output);
temp_list = [loss, euc_dist, euc_dist, y, output]
else:
euc_dist = euclidean_dist(y,output) # euclidean distance
temp_list = [loss, euc_dist] # list to hold sensor id, losses, values
for y_i, output_i in zip(y, output): # iterate through the predicted features
temp_list.append(abs(output_i-y_i))
temp_list.extend(y)
temp_list.extend(output)
losses_train.append(temp_list)
# + id="PHz04h_Hc-5_" colab={"base_uri": "https://localhost:8080/", "height": 204} executionInfo={"status": "ok", "timestamp": 1609285584438, "user_tz": -60, "elapsed": 174182, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}} outputId="ccd8d618-ca26-4051-f181-0c8cc93a6cf9"
# Convert to dataframe
df_losses = pd.DataFrame(losses_train)
df_losses.columns = column_names[1:]
df_losses.head()
# + id="zHi1MLCZ84t4" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1609285584439, "user_tz": -60, "elapsed": 174169, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}} outputId="e91c3021-beb7-489f-a3d9-29c85fe36eca"
param_thresholds = np.zeros((len(parameters_predicted)))
threshold = df_losses['euclidean_dist'].quantile(0.99)
for i, param in enumerate(parameters_predicted):
param_thresholds[i] = df_losses['loss_'+param].quantile(0.99)
param_thresholds
# + [markdown] id="lqolWzcF5jgK"
# ## Save model
# + id="_5f37KN4-Kyd" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1609285585645, "user_tz": -60, "elapsed": 175359, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "05119137911773041381"}} outputId="12622540-5825-4cc1-cfeb-e534757371c2"
def save_model_all(model, save_dir, model_name, epoch):
"""
:param model: nn model
:param save_dir: save model direction
:param model_name: model name
:param epoch: epoch
:return: None
"""
if not os.path.isdir(save_dir):
os.makedirs(save_dir)
save_prefix = os.path.join(save_dir, model_name)
save_path = '{}_epoch_{}.pt'.format(save_prefix, epoch)
print("save all model to {}".format(save_path))
output = open(save_path, mode="wb")
torch.save(model, output)
# torch.save(model.state_dict(), save_path)
output.close()
save_model_all([net, [threshold, param_thresholds]], '/content/drive/My Drive/WoodSense/notebooks/Stine', 'temp.pt', 100)
| Code/DeepAntT.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/nickprock/corso_data_science/blob/devs/intro_librerie_python/01_numpy.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="2ZOadGSbwWZC" colab_type="text"
# # NumPy
# [Numpy](http://www.numpy.org/) è una libreria open source che consente di svolgere mol te operazioni matematiche, in particolare è utilizzata molto per l'algebra lineare e il lavoro con vettori e matrici.
#
# <br>
# <img src='https://scipy-lectures.org/_images/numpy_indexing.png' width=400 heght=400>
# <br>
#
# [**Image Credit**](https://scipy-lectures.org/intro/numpy/array_object.html)
# <br>
#
# Per approfondire e tenere sempre a portata di mano le funzionalità più comuni consultare il [cheatsheet](https://www.datacamp.com/community/blog/python-numpy-cheat-sheet) ufficiale.
# + id="JT64jeGfvHM6" colab_type="code" colab={}
import numpy as np
print('numpy: {}'.format(np.__version__))
# + [markdown] id="X6gq_WqMyf9R" colab_type="text"
# ## Creazione degli array
#
# <br>
# <img src='http://community.datacamp.com.s3.amazonaws.com/community/production/ckeditor_assets/pictures/332/content_arrays-axes.png' width=500 heght=500>
# <br>
#
# [**Image Credit**](https://www.datacamp.com/community/tutorials/python-numpy-tutorial)
# + [markdown] id="7VIY-U6n3pYN" colab_type="text"
# ### Creare un array da una lista
# + id="YdMWwc2BxrFT" colab_type="code" colab={}
mylist = [1, 2, 3]
myarray = np.array(mylist)
print("dimensioni array originale: ", myarray.shape)
# modificare la dimensione di una array
myarray.resize(3,3)
print("dimensioni array modificate: ", myarray.shape)
print("array modificato: ", "\n", myarray)
# + [markdown] id="fUna02tG3u4o" colab_type="text"
# ### Creare un array di dimensioni definite e valori zero (o uno)
# + id="i2Fiu21zy2sw" colab_type="code" colab={}
print("array di zero: ", "\n", np.zeros((3,3)))
print("\n")
print("array di uno: ", "\n", np.ones((1,3)))
# + [markdown] id="HPksMJZu32ON" colab_type="text"
# ### Altri metodi per la creazione di un array
# + id="9yw-Keez0w06" colab_type="code" colab={}
# utilizzare la funziona arange
print(np.arange(1,10))
print("\n")
# creare una matrice di zero con la diagonale di uno
print(np.eye(4))
print("\n")
# creare un array per ripetizione
print(np.array([1,3,5] * 3))
print("\n")
# creare un array utilizzando repeat
print(np.repeat([1,2,3], 3))
# + [markdown] id="T_7k24W7VWFl" colab_type="text"
# ### Combinare gli array
# + id="jIucSzy_UoJy" colab_type="code" colab={}
a = np.array([1,2,3])
# combinare gli array in verticale
av = np.vstack([a, a*2])
print(av)
print("\n")
# combinare gli array in orizzontale
ah = np.hstack([a, a*2])
print(ah)
# + [markdown] colab_type="text" id="daJxCmJJogU1"
# ## Operazioni sugli array
# <br>
#
# <img src='http://scipy-lectures.org/_images/numpy_broadcasting.png'>
# <br>
#
# [Image Credit](http://scipy-lectures.org/intro/numpy/operations.html)
# + id="40wnrgm4WM0Z" colab_type="code" colab={}
x=np.array([1, 2, 3])
y=np.array([4, 5, 6])
print(x + y) # addizione [1 2 3] + [4 5 6] = [5 7 9]
print(x - y) # sottrazione [1 2 3] - [4 5 6] = [-3 -3 -3]
print(x * y) # moltiplicazione elemento per elemento [1 2 3] * [4 5 6] = [4 10 18]
print(x / y) # divisione elemento per elemento [1 2 3] / [4 5 6] = [0.25 0.4 0.5]
print(x**2) # potenza [1 2 3] ^2 = [1 4 9]
print(x.dot(y)) # prodotto cartesiano 1*4 + 2*5 + 3*6
# + id="F_omHwyosr0x" colab_type="code" colab={}
# numpy ha diverse funzioni per svolgere operazioni matematiche sui vettori, di seguito alcune tra le più comuni
x.sum() # somma gli elementi del vettore 1 + 2 + 3 = 6
x.max() # restituisce l'elemento dell'array con valore massimo 3 (analogo per il minimo)
x.mean() # media dei valori nel vettore 2
x.std() # deviazione strandard 0.8164
# argmax e argmin restituiscono l'indice del valore massimo e minimo nel vettore
x.argmax() # 2
x.argmin() # 0
# + [markdown] id="JFKXV72VqEbY" colab_type="text"
# ### Trasporre un vettore
# I vettori che abbiamo creato sono 1D, per trasporre bisogna avere almeno 2D
# + id="HAub9gZkpl72" colab_type="code" colab={}
# dimensioni originali
print(y,"\n")
print("dimensioni vettore originale: ", y.shape , "\n")
# aggiungere una dimensione
y.resize([1,3])
print("nuove dimensioni vettore: ", y.shape, "\n")
# trasporre
print("vettore trasposto: ", "\n", y.T)
# + [markdown] id="Ro16e_GHx5Bi" colab_type="text"
# ## Navigare un array
# Quando di và a navigare un array ricordare sempre che gli indicii partono da zero e l'ultimo indice non farà parte della selezione, come spiegato nell'immagine.
#
# <br>
# <img src='http://s8.picofile.com/file/8353147750/numpy_math2.png' width=300, height=200>
# <br>
#
# [Image Credit](https://www.stechies.com/numpy-indexing-slicing/)
# + _uuid="82ca4b616a46de3280b5a50df4c0114298d07aea" id="Sqz9Ev3Our1w" colab_type="code" colab={}
myarray = np.arange(12)**2
myarray
# + id="OqheT7ALscDo" colab_type="code" colab={}
# usare un indice per trovare un valore
myarray[4] # 16
myarray[-1] #121
# + id="0MRiTtUD3-Jk" colab_type="code" colab={}
# usare un indice di start e uno di stop
myarray[5:8]
# + id="eVAgK3Da4D9B" colab_type="code" colab={}
# selezionare da un elemento fino alla fine dell'array
myarray[4:]
# selezionare fino a un elemento dell'array
myarray[:7]
# + id="YfLFfC1i4Vvp" colab_type="code" colab={}
# utilizzare gli step
## scorrere l'array a passo 2
myarray[::2]
# + id="amlc6KtU4h0q" colab_type="code" colab={}
# array 2D
myarray.resize((2,6))
print(myarray)
print("\n")
myarray[:,2:]
# + id="uiFb5MUV6Ase" colab_type="code" colab={}
# selezionare solo gli elementi maggiori (o minori) di un certo valore
myarray[(myarray > 50) & (myarray < 100)]
# + [markdown] id="HcUX5iKDwpOU" colab_type="text"
# ## Copiare i dati
#
# Ci sono diversi modi di copiare i dati da un array, bisogna fare attenzione a quale si usa perchè potrebbe portare a conseguenze indesiderate.
# + id="9TVgIEtqw2fR" colab_type="code" colab={}
myarray = np.arange(1,10)
myarray.resize((3,3))
myarray
# + [markdown] id="FKnfTcU8xDPT" colab_type="text"
# Creaiamo un nuovo array come sottoinsieme di myarray.
# + id="k_6OOoY1xIfw" colab_type="code" colab={}
myarray2 = myarray[:2,:2]
myarray2
# + [markdown] id="R0g6EXp4xfdL" colab_type="text"
# Ora sostituiamo i valori in myarray2 con zero.
# + id="h_O_DE1oxllQ" colab_type="code" colab={}
myarray2[:] = 0
myarray2
# + [markdown] id="DrRQ0nSpxsCL" colab_type="text"
# Vediamo cosa succede sul vettore originale.
# + id="DhTOY8hbxwhw" colab_type="code" colab={}
myarray
# + [markdown] id="6-seCY2UxzMV" colab_type="text"
# Questo succede perchè myarray2 è subset di myarray, per copiare le informazioni bisogna usare *copy* che crea un vettore nuovo indipendente dall'originale.
# + id="wLSNASnpyINQ" colab_type="code" colab={}
myarray = np.arange(1,10)
myarray.resize((3,3))
myarray3 = myarray[:2,:2].copy()
print(myarray3)
myarray3[:]=0
# + [markdown] id="uAXm116IyoDv" colab_type="text"
# Vediamo cosa è successo all'originale e alla copia.
# + id="0h-eK8kpysXh" colab_type="code" colab={}
print("vettore originale: ", "\n", myarray, "\n")
print("vettore copia: ", "\n", myarray3, "\n")
# + [markdown] colab_type="text" id="sySx0_7jzMlp"
# ## Diversi tipi di iteratori
#
# In questo paragrafo vedremo diversi modi di *scorrere* gli elementi del vettore mediante l'utilizzo del ***ciclo for***.
#
# Per prima cosa creare un vettore, il vettore di test sarà 4x3 di interi tra 0 e 9.
#
# + id="3F0tROVqzv4w" colab_type="code" colab={}
test = np.random.randint(0, 10, (4,3))
test
# + id="3fWONu6R0q5j" colab_type="code" colab={}
# iterare per ogni riga
for row in test:
print(row)
print("\n")
# iterare per indice
for i in range(len(test)):
print(test[i])
print("\n")
# iterare con doppio indice usando enumerate
for i, row in enumerate(test):
print("row ", i, " is ", row)
# + id="YITu72WL2rJV" colab_type="code" colab={}
# iterare su due array mediante zip
test2 = test*2
for i, j in zip(test, test2):
print(i,'+',j,'=',i+j)
# + id="Ip9iHueU3ER9" colab_type="code" colab={}
# controllare i progressi del codice mediante tqdm
from tqdm import tqdm
for i in tqdm(range(len(test))):
for j in range(len(test[i])):
print ("riga: ", i, " colonna: ", j, " valore: ", test[i,j])
| intro_librerie_python/01_numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Uf0Pazc2SSXu"
# # Hitting your data with a Cue Ball:
# ## Introduction to Whitening or Sphering a Dataset
#
#
# ### (Re)Sources:
# - http://ufldl.stanford.edu/tutorial/unsupervised/PCAWhitening/
# - https://learndataa.com/2020/09/15/data-preprocessing-whitening-or-sphering-in-python/
#
# ### Introduction for the Reviewer:
#
# Sphereing or Whitening is one of those things that many machine learning practitioners see as a blackbox way of getting thier data in order. But visualizing what is under the hood is important here, especially since sphering can be a powerful tool when used in conjunction with various kernal tricks. In high dimensional data it can be easy to just abstract spehering away. Here we will try to make things as concrete as possible by visualizing sphering using 2 and 3D projections of high dimensional data.
#
# ### Target Learner:
# The Target Learner for this tutorial wants to be able to use sphering/whitening tools without treating them as a black box. In particular the target learner wants to use basic PCA based sphering to prepare data for further processing. Since they often deal with high dimensional data, they want to be able to see how sphering changes thier data using projections onto 2 or 3 dimensions.
#
# ### Assumptions:
# We assume the learner is proficient in linear algebra and statistics as well as python, basic numpy, and visualization libraries like Matplotlib. It is assumed that they understand what a given library is (generally) doing when the eigenvalues and eigenvectors of a matrix are retrieved.
#
# ### Learning Objectives:
# By the end of this course the learner will create a basic sphering function as well as a visualizer for the results of that function by selecting 2 or 3 dimensions or a high dimensional dataset.
#
#
# ### Syllabus:
#
# TODO FIND BETTER SAMPLE DATASET!
# TODO generate high dim gaussian data
# TODO built-in 3d visualizer picker using matplotlib
#
#
#
#
#
| Deeplearningai Curriculum Architect Course March 2022/Josh Week 1 Intermediate (TODO redo syllabus).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="GYiRsFGD6iUC"
# # 0 TorchText
# + [markdown] id="tp5IzBGsPGHs"
# ## Dataset Preview
#
# Your first step to deep learning in NLP. We will be mostly using PyTorch. Just like torchvision, PyTorch provides an official library, torchtext, for handling text-processing pipelines.
#
# We will be using previous session tweet dataset. Let's just preview the dataset.
# + colab={"base_uri": "https://localhost:8080/"} id="SKSTi9GepUmJ" outputId="9584e13f-b872-4b4b-e71b-c7a3fa9fde33"
from google.colab import drive
drive.mount('/content/gdrive/')
# + id="yhrCTg2Qphmm"
# !cp '/content/gdrive/My Drive/EVA/stanfordSentimentTreebank.zip' stanfordSentimentTreebank.zip
# !unzip -q -o stanfordSentimentTreebank.zip -d stanfordSentimentTreebank
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="o1-Yz-5RRFYc" outputId="1e09214b-c0f7-44b4-83dd-46c768402464"
import pandas as pd
df = pd.read_csv('/content/stanfordSentimentTreebank/stanfordSentimentTreebank/datasetSentences.txt',sep='\t')
df.tail()
# + colab={"base_uri": "https://localhost:8080/"} id="R7JdpCW-YbAG" outputId="09ecff92-b014-45f3-e869-a23c2ea20a22"
df.shape
# + colab={"base_uri": "https://localhost:8080/"} id="zqRsoF6xYdgl" outputId="70d80499-92c1-4ead-ed81-495f56df30d5"
df.info()
# + [markdown] id="XJ6o_79ISSVb"
# ## Defining Fields
# + id="AgaTy-qJ_ATn"
from torch.utils.data import DataLoader
from torch.optim.lr_scheduler import ReduceLROnPlateau
# + id="WoEXPawa_3tN"
class StanfordDatasetReader():
def __init__(self, sst_dir, split_idx):
merged_dataset = self.get_merged_dataset(sst_dir)
merged_dataset['sentiment values'] = merged_dataset['sentiment values'].astype(float)
self.dataset = merged_dataset[merged_dataset["splitset_label"] == split_idx]
# self.dataset["Revised_Sentiment"] = self.discretize_label(self.dataset.iloc[5])
self.dataset['Revised_sentiment values'] = self.dataset.apply(lambda x: labelfunc(x["sentiment values"]), axis=1)
# train_st_data['Revised_sentiment values'] = train_st_data.apply(lambda x: myfunc(x["sentiment values"]), axis=1)
# https://github.com/iamsimha/conv-sentiment-analysis/blob/master/code/dataset_reader.py
def get_merged_dataset(self, sst_dir):
sentiment_labels = pd.read_csv(os.path.join(sst_dir, "sentiment_labels.txt"), sep="|")
sentence_ids = pd.read_csv(os.path.join(sst_dir, "datasetSentences.txt"), sep="\t")
dictionary = pd.read_csv(os.path.join(sst_dir, "dictionary.txt"), sep="|", names=['phrase', 'phrase ids'])
train_test_split = pd.read_csv(os.path.join(sst_dir, "datasetSplit.txt"))
sentence_phrase_merge = pd.merge(sentence_ids, dictionary, left_on='sentence', right_on='phrase')
sentence_phrase_split = pd.merge(sentence_phrase_merge, train_test_split, on='sentence_index')
return pd.merge(sentence_phrase_split, sentiment_labels, on='phrase ids').sample(frac=1)
def discretize_label(self, label):
print(type(label))
if label <= 0.2: return 0
if label <= 0.4: return 1
if label <= 0.6: return 2
if label <= 0.8: return 3
return 4
def word_to_index(self, word):
if word in self.w2i:
return self.w2i[word]
else:
return self.w2i["<OOV>"]
def __len__(self):
return self.dataset.shape[0]
# def __getitem__(self, idx):
# return {"sentence": [self.word_to_index(x) for x in self.dataset.iloc[idx, 1].split()],
# "label": self.discretize_label(self.dataset.iloc[idx, 5])}
def labelfunc(label):
if label <= 0.2: return 0
if label <= 0.4: return 1
if label <= 0.6: return 2
if label <= 0.8: return 3
return 4
def get_data(self):
return self.dataset
def __getitem__(self, idx):
return {"sentence": [x for x in self.dataset.iloc[idx, 1].split()],
"label": self.discretize_label(self.dataset.iloc[idx, 5])}
# + id="G1BWSC8jd7AZ"
def labelfunc(label):
if label <= 0.2: return 0
if label <= 0.4: return 1
if label <= 0.6: return 2
if label <= 0.8: return 3
return 4
# + id="rBxRHuT5Fhyb"
import os
def load_data(sst_dir="/content/stanfordSentimentTreebank/stanfordSentimentTreebank/"):
train_st_data_cl = StanfordDatasetReader(sst_dir, 1).get_data()
# train_st_data_cl['Revised_sentiment values'] = train_st_data.apply(lambda x: labelfunc(x["sentiment values"]), axis=1)
test_st_data_cl = StanfordDatasetReader(sst_dir, 2).get_data()
# test_st_data_cl['Revised_sentiment values'] = test_st_data_cl.apply(lambda x: labelfunc(x["sentiment values"]), axis=1)
validation_st_data_cl = StanfordDatasetReader(sst_dir, 3).get_data()
# validation_st_data_cl['Revised_sentiment values'] = validation_st_data_cl.apply(lambda x: labelfunc(x["sentiment values"]), axis=1)
return train_st_data_cl,test_st_data_cl,validation_st_data_cl
# + colab={"base_uri": "https://localhost:8080/"} id="ABfKELBlJf2v" outputId="8b2611cf-47fa-44f8-8107-a2032249b6f1"
train_st_data,test_st_data,validation_st_data = load_data()
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="c10qU_CBoOZM" outputId="0c10a80d-bda8-41dd-a319-c06e816200a2"
train_st_data.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="fwFAQsWogYg-" outputId="3ecaa80e-11a5-41f8-cdad-082efa5daff2"
test_st_data.tail()
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="HzM5HDWkgcip" outputId="3082c7c3-9898-4598-f82a-658d58074340"
validation_st_data.head()
# + [markdown] id="dFCfiRpdGnQD"
# ### Further NLP Augemnattion
# + colab={"base_uri": "https://localhost:8080/"} id="4Ubb2QEEG0in" outputId="4f48496c-204c-422b-f055-c9cf950e1b94"
# !pip install nlpaug
# + id="PMI8CXdrIMGy"
# # !pip install transformers
# + id="TGXDQghJEqmV"
## Lets do the NLP data augmentation
import nlpaug.augmenter.char as nac
import nlpaug.augmenter.word as naw
import nlpaug.augmenter.sentence as nas
import nlpaug.flow as nafc
from nlpaug.util import Action
# + [markdown] id="ed8FaP3MG-kc"
# ##### Some basic examples for understanding and then further data augmentation by these
# - Substitute word by WordNet's synonym
# - Swap word randomly
# - Delete a set of contunous word will be removed randomly
# - Delete word randomly augemnattion
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="9gledVX9KtEj" outputId="9991300a-7372-46d9-ff5c-1136cd52ded6"
# validation_st_data.head()
train_st_data['sentence'].iloc[0]
# + colab={"base_uri": "https://localhost:8080/"} id="B0gu3UXtXqH4" outputId="93a40002-16ce-4efc-fcd7-182d9d6dee98"
aug = naw.SynonymAug(aug_src='wordnet') ## Substitute word by WordNet's synonym¶
augmented_text = aug.augment(train_st_data['sentence'].iloc[0])
print("Original:")
print(train_st_data['sentence'].iloc[0])
print("Augmented Text:")
print(augmented_text)
train_st_data_SynonymAug_aug = train_st_data
train_st_data_SynonymAug_aug['sentence_aug'] = train_st_data_SynonymAug_aug.apply(lambda x: aug.augment(x['sentence']),axis=1) ## Swap word randomly¶
# + colab={"base_uri": "https://localhost:8080/"} id="vBoPakkKX3ly" outputId="573425bf-18d8-43a1-cac1-138215df59d1"
aug = naw.RandomWordAug(action="swap") # Swap word randomly¶
augmented_text = aug.augment(train_st_data['sentence'].iloc[0])
print("Original:")
print(train_st_data['sentence'].iloc[0])
print("Augmented Text:")
print(augmented_text)
train_st_data_swap_aug = train_st_data
train_st_data_swap_aug['sentence_aug'] = train_st_data_swap_aug.apply(lambda x: aug.augment(x['sentence']),axis=1) ## Swap word randomly¶
# + id="n24qk3VIYIR3"
# aug = naw.RandomWordAug(action='crop',aug_p=0.5, aug_min=0)
# augmented_text = aug.augment(train_st_data['sentence'].iloc[0]) ## Delete a set of contunous word will be removed randomly¶
# print("Original:")
# print(train_st_data['sentence'].iloc[0])
# print("Augmented Text:")
# print(augmented_text)
# train_st_data_crop_aug = train_st_data
# train_st_data_crop_aug['sentence_aug'] = train_st_data_crop_aug.apply(lambda x: aug.augment(x['sentence']),axis=1) ## Delete a set of contunous word will be removed randomly¶
# + id="mcDwOTqxNXkc"
text = 'The quick brown fox jumps over the lazy dog .'
# Augmenter that apply random word operation to textual input.Augmenter that apply randomly behavior for augmentation.
aug = naw.RandomWordAug()
augmented_data = aug.augment(text)
augmented_data
train_st_data_delete_aug = train_st_data
# train_st_data_aug[sentence_aug] = aug.augment(train_st_data_aug.loc["sentence"] )
#--Using position to slice Email using a lambda function
train_st_data_delete_aug['sentence_aug'] = train_st_data_delete_aug.apply(lambda x: aug.augment(x['sentence']),axis=1) ## Delete word randomly augemnattion
# + colab={"background_save": true} id="fFRlJ07oZnwX" outputId="b0a17e9d-3dcb-4e32-b72a-924570ba0ee9"
print("Original:")
print(train_st_data_delete_aug['sentence'].iloc[0])
print("Augmented Text:")
print(train_st_data_delete_aug['sentence_aug'].iloc[0])
# + colab={"background_save": true} id="2zMubaWhVP2w" outputId="290a72b0-7fcf-42fd-8de1-5808bcfae097"
train_st_data_delete_aug.head()
# + id="Qw_pqEU9csk0"
## Now I need to add all these data frames
combined_data_aug = pd.concat([train_st_data_delete_aug, train_st_data_swap_aug, train_st_data_SynonymAug_aug], axis=0)
## after this, now I need to drop the sentence column and rename sentence_aug to sentence
combined_data_aug.drop('sentence', axis=1, inplace=True)
combined_data_aug.rename(columns = {'sentence_aug':'sentence'}, inplace = True)
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="quNy-dDMzfeL" outputId="f7ae9958-1107-45ba-c3d0-87eada7e4576"
combined_data_aug.head()
# + [markdown] id="9ZkxFvYtGs7K"
# ### Final Data Preparation
# + id="arFy0s73nImt"
def get_final_data(train_st_data,test_st_data,validation_st_data,combined_data_aug):
train_st_data_final = train_st_data.drop(['sentence_index','phrase','phrase ids','splitset_label','sentiment values'],axis=1)
train_st_data_final.rename(columns = {'Revised_sentiment values': 'sentiment'}, inplace = True)
combined_data_aug.drop(['sentence_index','phrase','phrase ids','splitset_label','sentiment values'],axis=1,inplace=True)
combined_data_aug.rename(columns = {'Revised_sentiment values': 'sentiment'}, inplace = True)
train_st_data_final_mixed = pd.concat([combined_data_aug, train_st_data_final], axis=0)
train_st_data_final_mixed = train_st_data_final_mixed.reset_index(drop=True) ## This is being done because data.Example.fromlist was failing
test_st_data_final = test_st_data.drop(['sentence_index','phrase','phrase ids','splitset_label','sentiment values'],axis=1)
test_st_data_final.rename(columns = {'Revised_sentiment values': 'sentiment'}, inplace = True)
test_st_data_final = test_st_data_final.reset_index(drop=True) ## This is being done because data.Example.fromlist was failing
validation_st_data_final = validation_st_data.drop(['sentence_index','phrase','phrase ids','splitset_label','sentiment values'],axis=1)
validation_st_data_final.rename(columns = {'Revised_sentiment values': 'sentiment'} , inplace = True)
validation_st_data_final = validation_st_data_final.reset_index(drop=True) ## This is being done because data.Example.fromlist was failing
return train_st_data_final_mixed, test_st_data_final, validation_st_data_final
# + id="tjmsNd8wJiIj"
train_st_data_final, test_st_data_final, validation_st_data_final = get_final_data(train_st_data,test_st_data,validation_st_data,combined_data_aug)
train_st_data_final.drop(['sentence_aug'],axis=1,inplace=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="iWZytpY6pE2S" outputId="36687e07-be66-4df1-917b-e1a2729e0c69"
train_st_data_final.head()
# + id="v0p26-wQLbV4"
# train_st_data_final.to_csv(r'train_st_data_final.csv', index = False)
# + colab={"base_uri": "https://localhost:8080/"} id="--Z1M-vssT03" outputId="95c277a9-d28b-4b46-d02c-e0c475d2905f"
train_st_data_final.shape
# + colab={"base_uri": "https://localhost:8080/"} id="TCq6t89LsXvI" outputId="1916b624-b8dd-477a-d21c-eeb88eefee7e"
train_st_data_final.sentiment.value_counts()
# + colab={"base_uri": "https://localhost:8080/"} id="9dQTLWRqVxME" outputId="e01c352b-c0b4-4006-cfdb-b2e7cba3848e"
validation_st_data_final.sentiment.value_counts()
# + [markdown] id="e63g08ijOrf7"
# Now we shall be defining LABEL as a LabelField, which is a subclass of Field that sets sequen tial to False (as it’s our numerical category class). TWEET is a standard Field object, where we have decided to use the spaCy tokenizer and convert all the text to lower‐ case.
# + colab={"base_uri": "https://localhost:8080/"} id="qk8IP4SK1Lrp" outputId="321d5274-32dc-4695-b498-b6b9aeb2028c"
# Import Library
import random
import torch, torchtext
from torchtext import data
import pandas as pd
# Manual Seed
SEED = 43
torch.manual_seed(SEED)
# + id="u6bKQax2Mf_U"
Sentence = data.Field(sequential = True, tokenize = 'spacy', batch_first =True, include_lengths=True)
Sentiment = data.LabelField(tokenize ='spacy',is_target=True, batch_first =True, sequential =False)
# + [markdown] id="mX-lYIe_O7Vy"
# Having defined those fields, we now need to produce a list that maps them onto the list of rows that are in the CSV:
# + id="VawdWq36O6td"
fields = [('sentence', Sentence),('sentiment',Sentiment)]
# + id="w0UOsNaNO-hp"
# saving the dataframe
train_st_data_final.to_csv('train_st_data_final.csv', index=False)
test_st_data_final.to_csv('test_st_data_final.csv', index=False)
validation_st_data_final.to_csv('validation_st_data_final.csv', index=False)
# + id="ruDvWc02eG-N"
# !cp '/content/gdrive/My Drive/EVA/train_st_data_final_fine_grained.csv' train_st_data_final_fine_grained.csv
# !cp '/content/gdrive/My Drive/EVA/validation_st_data_final_fine_grained.csv' validation_st_data_final_fine_grained.csv
# !cp '/content/gdrive/My Drive/EVA/test_st_data_final_fine_grained.csv' test_st_data_final_fine_grained.csv
# + id="xPXcc-LifvmN"
train_st_data_final = pd.read_csv("train_st_data_final_fine_grained.csv")
test_st_data_final = pd.read_csv("test_st_data_final_fine_grained.csv")
validation_st_data_final = pd.read_csv("validation_st_data_final_fine_grained.csv")
# + id="pjwYTMZG6h9d" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="25d4752b-3782-4f3c-c5ba-6c713d926adc"
train_st_data_final.head()
# train_st_data_final.drop(['sentence_aug'],axis=1,inplace=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="Pa_O5Wqq6s63" outputId="171f224e-9ec4-41b1-86ff-dbe6cdaffa54"
test_st_data_final.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="oqIJCrH_0HZ4" outputId="e5f7e2b7-705a-428c-8dc7-f9e4f16aacda"
import matplotlib.pyplot as plt
ax = train_st_data_final['sentiment'].value_counts(sort=False).plot(kind='barh')
ax.set_xlabel("Number of Samples in training Set")
ax.set_ylabel("sentiment")
# + [markdown] id="q10sQx1G7Ms4"
# It is clear that most of the training samples belong to classes 0 and 3 (the weakly negative/positive classes). A sizeable number of samples belong to the neutral class. Barely 12% of the samples are from the strongly negative class 1, which is something to keep in mind as we evaluate our classifier accuracy.
# + id="sgH1g8jTMPuH"
# # saving the dataframe
# train_st_data_final.to_csv('train_st_data_final.csv', index=False)
# test_st_data_final.to_csv('test_st_data_final.csv', index=False)
# validation_st_data_final.to_csv('validation_st_data_final.csv', index=False)
# + [markdown] id="qIlOGo2HvOHi"
#
# + [markdown] id="ZbtZ-Ph2P1xL"
# Armed with our declared fields, lets convert from pandas to list to torchtext. We could also use TabularDataset to apply that definition to the CSV directly but showing an alternative approach too.
# + id="L3OLcJ5B7rHz"
example_trng = [data.Example.fromlist([train_st_data_final.sentence[i],train_st_data_final.sentiment[i]], fields) for i in range(train_st_data_final.shape[0])]
example_val = [data.Example.fromlist([validation_st_data_final.sentence[i],validation_st_data_final.sentiment[i]], fields) for i in range(validation_st_data_final.shape[0])]
# + id="nT-flpH-P1cd"
# Creating dataset
#twitterDataset = data.TabularDataset(path="tweets.csv", format="CSV", fields=fields, skip_header=True)
# twitterDataset = data.Dataset(example, fields)
train = data.Dataset(example_trng, fields)
valid = data.Dataset(example_val, fields)
# + [markdown] id="g6ZnyCPaR08F"
# Finally, we can split into training, testing, and validation sets by using the split() method:
# + id="uPYXyuKhRpBk"
# (train, valid) = twitterDataset.split(split_ratio=[0.85, 0.15], random_state=random.seed(SEED))
# + colab={"base_uri": "https://localhost:8080/"} id="ykvsCGQMR6UD" outputId="d7cbe283-8265-4dd2-8c5d-022a102c1827"
(len(train), len(valid))
# + [markdown] id="kix8P2IKSBaV"
# An example from the dataset:
# + colab={"base_uri": "https://localhost:8080/"} id="dUpEOQruR9JL" outputId="203cf75e-aada-4068-b24c-ed601bf33b7a"
vars(train.examples[10])
# + [markdown] id="AKdllP3FST4N"
# ## Building Vocabulary
# + [markdown] id="SuvWQ-SpSmSz"
# At this point we would have built a one-hot encoding of each word that is present in the dataset—a rather tedious process. Thankfully, torchtext will do this for us, and will also allow a max_size parameter to be passed in to limit the vocabu‐ lary to the most common words. This is normally done to prevent the construction of a huge, memory-hungry model. We don’t want our GPUs too overwhelmed, after all.
#
# Let’s limit the vocabulary to a maximum of 5000 words in our training set:
#
# + colab={"base_uri": "https://localhost:8080/"} id="IGH3R6IYcY_f" outputId="a3c639da-4635-4055-faf2-123d47cb83e5"
MAX_VOCAB_SIZE = 25_000
Sentence.build_vocab(train,
max_size = MAX_VOCAB_SIZE,
vectors = "glove.6B.200d",
# vectors = "glove.6B.300d",
unk_init = torch.Tensor.normal_)
# + id="u9CVBX6IhZtL"
Sentiment.build_vocab(train)
# https://github.com/shayneobrien/sentiment-classification/blob/master/notebooks/11-cnn-conv2d-cbow-glove.ipynb
# https://github.com/shayneobrien/sentiment-classification
# + id="mx955u93SGeY"
# Sentence.build_vocab(train)
# Sentiment.build_vocab(train)
# + [markdown] id="xvyEeEjXTGhX"
# By default, torchtext will add two more special tokens, <unk> for unknown words and <pad>, a padding token that will be used to pad all our text to roughly the same size to help with efficient batching on the GPU.
# + colab={"base_uri": "https://localhost:8080/"} id="rA3tIESdcJdN" outputId="a1d383b5-b757-42ba-de41-b4c93dccd47b"
print('Size of input vocab : ', len(Sentence.vocab))
print('Size of label vocab : ', len(Sentiment.vocab))
print('Top 10 words appreared repeatedly :', list(Sentence.vocab.freqs.most_common(10)))
print('Labels : ', Sentiment.vocab.stoi)
# + colab={"base_uri": "https://localhost:8080/"} id="BOgXGAD3fqQs" outputId="b052cd1d-0088-41d8-f0bb-9578e80d2236"
print('Size of input vocab : ', len(Sentence.vocab))
print('Size of label vocab : ', len(Sentiment.vocab))
print('Top 10 words appreared repeatedly :', list(Sentence.vocab.freqs.most_common(10)))
print('Labels : ', Sentiment.vocab.stoi)
# + [markdown] id="rwjD2-ebTeUX"
# **Lots of stopwords!!**
# + [markdown] id="sLWW221gTpNs"
# Now we need to create a data loader to feed into our training loop. Torchtext provides the BucketIterator method that will produce what it calls a Batch, which is almost, but not quite, like the data loader we used on images.
# + [markdown] id="EQqMhMoDUDmn"
# But at first declare the device we are using.
# + id="Zfo2QhGJUK4l"
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# + id="zK2ORoqdTNsM"
train_iterator, valid_iterator = data.BucketIterator.splits((train, valid), batch_size = 32,
sort_key = lambda x: len(x.sentence),
sort_within_batch=True, device = device)
# + [markdown] id="Gg7gTFQO4fby"
# Save the vocabulary for later use
# + id="niE9Cc6-2bD_"
import os, pickle
with open('tokenizer.pkl', 'wb') as tokens:
pickle.dump(Sentence.vocab.stoi, tokens)
# + [markdown] id="1AbsQwqkVyAy"
# ## Defining Our Model
# + [markdown] id="E4PED4HJWH4t"
# We use the Embedding and LSTM modules in PyTorch to build a simple model for classifying tweets.
#
# In this model we create three layers.
# 1. First, the words in our tweets are pushed into an Embedding layer, which we have established as a 300-dimensional vector embedding.
# 2. That’s then fed into a 2 stacked-LSTMs with 100 hidden features (again, we’re compressing down from the 300-dimensional input like we did with images). We are using 2 LSTMs for using the dropout.
# 3. Finally, the output of the LSTM (the final hidden state after processing the incoming tweet) is pushed through a standard fully connected layer with three outputs to correspond to our three possible classes (negative, positive, or neutral).
# + id="43pVRccMT0bT"
import torch.nn as nn
import torch.nn.functional as F
class classifier(nn.Module):
# Define all the layers used in model
def __init__(self, vocab_size, embedding_dim, hidden_dim, output_dim, n_layers, dropout, bidirectional,pad_idx):
super().__init__()
# Embedding layer
self.embedding = nn.Embedding(vocab_size, embedding_dim,padding_idx = pad_idx)
# LSTM layer
self.encoder = nn.LSTM(embedding_dim,
hidden_dim,
num_layers=n_layers,
dropout=dropout,
bidirectional = bidirectional,
batch_first=True)
# try using nn.GRU or nn.RNN here and compare their performances
# try bidirectional and compare their performances
self.dropout = nn.Dropout(dropout)
# Dense layer
self.fc = nn.Linear(hidden_dim, output_dim)
def forward(self, text, text_lengths):
# text = [batch size, sent_length]
embedded = self.embedding(text)
# embedded = [batch size, sent_len, emb dim]
# packed sequence
packed_embedded = nn.utils.rnn.pack_padded_sequence(embedded, text_lengths.cpu(), batch_first=True)
packed_output, (hidden, cell) = self.encoder(packed_embedded)
#hidden = [batch size, num layers * num directions,hid dim]
#cell = [batch size, num layers * num directions,hid dim]
# Hidden = [batch size, hid dim * num directions]
dense_outputs = self.dropout(self.fc(hidden))
# Final activation function softmax
output = F.softmax(dense_outputs[0], dim=1)
# output = F.softmax(dense_outputs, dim=1)
return output
# + id="qGXHblEXX5GJ"
import torch.nn as nn
import torch.nn.functional as F
class classifier(nn.Module):
# Define all the layers used in model
def __init__(self, vocab_size, embedding_dim, hidden_dim, output_dim, n_layers, dropout, bidirectional,pad_idx):
super().__init__()
# Embedding layer
self.embedding = nn.Embedding(vocab_size, embedding_dim,padding_idx = pad_idx)
# LSTM layer
self.encoder = nn.GRU(embedding_dim,
hidden_dim,
num_layers=n_layers,
dropout=dropout,
bidirectional = bidirectional,
batch_first=True)
# try using nn.GRU or nn.RNN here and compare their performances
# try bidirectional and compare their performances
self.dropout = nn.Dropout(dropout)
# Dense layer
self.fc = nn.Linear(hidden_dim, output_dim)
def forward(self, text, text_lengths):
# text = [batch size, sent_length]
embedded = self.embedding(text)
# embedded = [batch size, sent_len, emb dim]
# packed sequence
packed_embedded = nn.utils.rnn.pack_padded_sequence(embedded, text_lengths.cpu(), batch_first=True)
packed_output, (hidden) = self.encoder(packed_embedded)
#hidden = [batch size, num layers * num directions,hid dim]
#cell = [batch size, num layers * num directions,hid dim]
# Hidden = [batch size, hid dim * num directions]
dense_outputs = self.dropout(self.fc(hidden))
# Final activation function softmax
output = F.softmax(dense_outputs[0], dim=1)
# output = F.softmax(dense_outputs, dim=1)
return output
# + id="rwBoGE_X_Fl8"
# Define hyperparameters
size_of_vocab = len(Sentence.vocab)
embedding_dim = 200
num_hidden_nodes = 256
num_output_nodes = 5
num_layers = 5
dropout = 0.4
bidirectional = True
PAD_IDX = Sentence.vocab.stoi[Sentence.pad_token]
# Instantiate the model
model = classifier(size_of_vocab, embedding_dim, num_hidden_nodes, num_output_nodes, num_layers, dropout, bidirectional,PAD_IDX)
# + colab={"base_uri": "https://localhost:8080/"} id="O-pOMqzJ3eTv" outputId="bccbebe0-041e-4a08-dd5e-bb5d7bf2ee32"
print(model)
#No. of trianable parameters
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
print(f'The model has {count_parameters(model):,} trainable parameters')
# + [markdown] id="eXajorf5Xz7t"
# ## Model Training and Evaluation
# + [markdown] id="PrE9RpMtZ1Vs"
# First define the optimizer and loss functions
# + id="-u86JWdlXvu5"
import torch.optim as optim
# define optimizer and loss
optimizer = optim.Adam(model.parameters(), lr=2e-3)
criterion = nn.CrossEntropyLoss()
# define metric
def binary_accuracy(preds, y):
#round predictions to the closest integer
_, predictions = torch.max(preds, 1)
correct = (predictions == y).float()
acc = correct.sum() / len(correct)
return acc
# + colab={"base_uri": "https://localhost:8080/"} id="Rt_e2IkGdRW8" outputId="9b635a33-358b-4552-925b-035ec05cefcf"
pretrained_embeddings = Sentence.vocab.vectors
print(pretrained_embeddings.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="t1bHOjdUdY2a" outputId="27328700-845a-4f19-f5be-c543f78f010e"
model.embedding.weight.data.copy_(pretrained_embeddings)
# + colab={"base_uri": "https://localhost:8080/"} id="ASWyq5tgiyLt" outputId="01a05239-17ed-4e04-8807-00f09ebbb7a0"
UNK_IDX = Sentence.vocab.stoi[Sentence.unk_token]
model.embedding.weight.data[UNK_IDX] = torch.zeros(embedding_dim)
model.embedding.weight.data[PAD_IDX] = torch.zeros(embedding_dim)
print(model.embedding.weight.data)
# + id="GiVj8tajdTTC"
# push to cuda if available
model = model.to(device)
criterion = criterion.to(device)
# + [markdown] id="3VCJtNb3Zt8w"
# The main thing to be aware of in this new training loop is that we have to reference `batch.tweets` and `batch.labels` to get the particular fields we’re interested in; they don’t fall out quite as nicely from the enumerator as they do in torchvision.
# + [markdown] id="2WjEPLKsAiS_"
# **Training Loop**
# + id="HDWNnGK3Y5oJ"
def train(model, iterator, optimizer, criterion):
# initialize every epoch
epoch_loss = 0
epoch_acc = 0
# set the model in training phase
model.train()
for batch in iterator:
# resets the gradients after every batch
optimizer.zero_grad()
# retrieve text and no. of words
tweet, tweet_lengths = batch.sentence
# convert to 1D tensor
predictions = model(tweet, tweet_lengths).squeeze()
# compute the loss
loss = criterion(predictions, batch.sentiment)
# compute the binary accuracy
acc = binary_accuracy(predictions, batch.sentiment)
# backpropage the loss and compute the gradients
loss.backward()
# update the weights
optimizer.step()
# loss and accuracy
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
# + [markdown] id="CZcHhkkvAsCt"
# **Evaluation Loop**
# + id="zHEe-zSVAriL"
def evaluate(model, iterator, criterion):
# initialize every epoch
epoch_loss = 0
epoch_acc = 0
# deactivating dropout layers
model.eval()
# deactivates autograd
with torch.no_grad():
for batch in iterator:
# retrieve text and no. of words
tweet, tweet_lengths = batch.sentence
# convert to 1d tensor
predictions = model(tweet, tweet_lengths).squeeze()
# compute loss and accuracy
loss = criterion(predictions, batch.sentiment)
acc = binary_accuracy(predictions, batch.sentiment)
# keep track of loss and accuracy
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
# + [markdown] id="L6LJFW7HaJoV"
# **Let's Train and Evaluate**
# + colab={"base_uri": "https://localhost:8080/"} id="tq330XlnaEU9" outputId="6c9e1643-a123-4ea8-cb76-9aba8310f23c"
N_EPOCHS = 15
best_valid_loss = float('inf')
#freeze embeddings
model.embedding.weight.requires_grad = unfrozen = False
for epoch in range(N_EPOCHS):
# train the model
train_loss, train_acc = train(model, train_iterator, optimizer, criterion)
# evaluate the model
valid_loss, valid_acc = evaluate(model, valid_iterator, criterion)
# save the best model
if valid_loss < best_valid_loss:
best_valid_loss = valid_loss
torch.save(model.state_dict(), 'saved_weights.pt')
print(f'\tTrain Loss: {train_loss:.3f} | Train Acc: {train_acc*100:.2f}%')
print(f'\t Val. Loss: {valid_loss:.3f} | Val. Acc: {valid_acc*100:.2f}% \n')
# + id="fs9COVJlW_JI"
path='./saved_weights.pt'
model.load_state_dict(torch.load(path));
# + colab={"base_uri": "https://localhost:8080/"} id="W6Z7TKg_W6s4" outputId="daae17f4-ab37-45e2-a781-038ac36e093d"
# N_EPOCHS = 15
# best_valid_loss = float('inf')
# #freeze embeddings
# model.embedding.weight.requires_grad = unfrozen = True
# for epoch in range(N_EPOCHS):
# # train the model
# train_loss, train_acc = train(model, train_iterator, optimizer, criterion)
# # evaluate the model
# valid_loss, valid_acc = evaluate(model, valid_iterator, criterion)
# # save the best model
# if valid_loss < best_valid_loss:
# best_valid_loss = valid_loss
# torch.save(model.state_dict(), 'saved_weights.pt')
# print(f'\tTrain Loss: {train_loss:.3f} | Train Acc: {train_acc*100:.2f}%')
# print(f'\t Val. Loss: {valid_loss:.3f} | Val. Acc: {valid_acc*100:.2f}% \n')
# + [markdown] id="LZgzB0ZkHVTI"
# ## Model Testing
# + id="aZZfnWo0abRx"
#load weights and tokenizer
path='./saved_weights.pt'
model.load_state_dict(torch.load(path));
model.eval();
tokenizer_file = open('./tokenizer.pkl', 'rb')
tokenizer = pickle.load(tokenizer_file)
#inference
import spacy
nlp = spacy.load('en')
def classify_tweet(tweet):
categories = {0: "Negative", 1:"Positive", 2:"Neutral"}
# tokenize the tweet
tokenized = [tok.text for tok in nlp.tokenizer(tweet)]
# convert to integer sequence using predefined tokenizer dictionary
indexed = [tokenizer[t] for t in tokenized]
# compute no. of words
length = [len(indexed)]
# convert to tensor
tensor = torch.LongTensor(indexed).to(device)
# reshape in form of batch, no. of words
tensor = tensor.unsqueeze(1).T
# convert to tensor
length_tensor = torch.LongTensor(length)
# Get the model prediction
prediction = model(tensor, length_tensor)
_, pred = torch.max(prediction, 1)
return categories[pred.item()]
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="yTkHLEipIlM9" outputId="da80d345-e226-4d45-fde0-b95d9821d8da"
classify_tweet("A valid explanation for why Trump won't let women on the golf course.")
# + [markdown] id="WVjCuKK_LVEF"
# ## Discussion on Data Augmentation Techniques
#
# You might wonder exactly how you can augment text data. After all, you can’t really flip it horizontally as you can an image! :D
#
# In contrast to data augmentation in images, augmentation techniques on data is very specific to final product you are building. As its general usage on any type of textual data doesn't provides a significant performance boost, that's why unlike torchvision, torchtext doesn’t offer a augmentation pipeline. Due to powerful models as transformers, augmentation tecnhiques are not so preferred now-a-days. But its better to know about some techniques with text that will provide your model with a little more information for training.
#
# ### Synonym Replacement
#
# First, you could replace words in the sentence with synonyms, like so:
#
# The dog slept on the mat
#
# could become
#
# The dog slept on the rug
#
# Aside from the dog's insistence that a rug is much softer than a mat, the meaning of the sentence hasn’t changed. But mat and rug will be mapped to different indices in the vocabulary, so the model will learn that the two sentences map to the same label, and hopefully that there’s a connection between those two words, as everything else in the sentences is the same.
# + [markdown] id="T_uEfWJpL6Nq"
# ### Random Insertion
# A random insertion technique looks at a sentence and then randomly inserts synonyms of existing non-stopwords into the sentence n times. Assuming you have a way of getting a synonym of a word and a way of eliminating stopwords (common words such as and, it, the, etc.), shown, but not implemented, in this function via get_synonyms() and get_stopwords(), an implementation of this would be as follows:
#
# + id="7Alm5D7WIvAC"
def random_insertion(sentence, n):
words = remove_stopwords(sentence)
for _ in range(n):
new_synonym = get_synonyms(random.choice(words))
sentence.insert(randrange(len(sentence)+1), new_synonym)
return sentence
# + [markdown] id="gqLWzwJ3Mm8h"
# ## Random Deletion
# As the name suggests, random deletion deletes words from a sentence. Given a probability parameter p, it will go through the sentence and decide whether to delete a word or not based on that random probability. Consider of it as pixel dropouts while treating images.
# + id="-7Dz7JJfMqyC"
def random_deletion(words, p=0.5):
if len(words) == 1: # return if single word
return words
remaining = list(filter(lambda x: random.uniform(0,1) > p,words))
if len(remaining) == 0: # if not left, sample a random word
return [random.choice(words)]
else:
return remaining
# + [markdown] id="zOIbi5WzO5OU"
# ### Random Swap
# The random swap augmentation takes a sentence and then swaps words within it n times, with each iteration working on the previously swapped sentence. Here we sample two random numbers based on the length of the sentence, and then just keep swapping until we hit n.
# + id="LnkbG15HO3Yj"
def random_swap(sentence, n=5):
length = range(len(sentence))
for _ in range(n):
idx1, idx2 = random.sample(length, 2)
sentence[idx1], sentence[idx2] = sentence[idx2], sentence[idx1]
return sentence
# + [markdown] id="599NpwfMR5Vm"
# For more on this please go through this [paper](https://arxiv.org/pdf/1901.11196.pdf).
# + [markdown] id="a5aeKuNCRGip"
# ### Back Translation
#
# Another popular approach for augmenting text datasets is back translation. This involves translating a sentence from our target language into one or more other languages and then translating all of them back to the original language. We can use the Python library googletrans for this purpose.
# + colab={"base_uri": "https://localhost:8080/"} id="B2a5FZkoDSyL" outputId="9269405b-950c-4377-d333-f33ce2d323b8"
# !pip install googletrans==3.1.0a0
# + colab={"base_uri": "https://localhost:8080/"} id="pHhNBbYrRXNy" outputId="8b97e6f2-836d-4590-ced6-85c7ddc435a1"
import random
import googletrans
from googletrans import Translator
# import googletrans.Translator
translator = Translator()
sentence = ['The dog slept on the rug']
available_langs = list(googletrans.LANGUAGES.keys())
trans_lang = random.choice(available_langs)
print(f"Translating to {googletrans.LANGUAGES[trans_lang]}")
translations = translator.translate(sentence, dest=trans_lang)
t_text = [t.text for t in translations]
print(t_text)
translations_en_random = translator.translate(t_text, src=trans_lang, dest='en')
en_text = [t.text for t in translations_en_random]
print(en_text)
| Stanford_Sentiment_Treebank_Sentiment_Analysis/src/END_NLP_Assignment_7_StanfordSentimentAnalysis_GRU_FineGrained_2_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# -
data=pd.read_csv("C:\\Users\\91760\\Downloads\\indian_liver_patient.csv")
data.head(10)
## Checking null values
data.isnull().sum()
data.rename(columns={"Dataset":"Target"},inplace=True)
sns.countplot(data=data,x="Target",label="Count")
data.info()
data.describe()
sns.factorplot(x="Age",y="Gender",hue="Target",data=data)
g=sns.FacetGrid(data,col="Target",row="Gender",margin_titles=True)
g.map(plt.hist,"Age")
plt.subplots_adjust(top=0.9)
g.fig.suptitle('Disease by Age and Gender')
g=sns.FacetGrid(data,col="Gender",row="Target",margin_titles=True)
g.map(plt.scatter,"Direct_Bilirubin","Total_Bilirubin",edgecolor="w")
plt.subplots_adjust(top=0.9)
data["Albumin_and_Globulin_Ratio"] = data.Albumin_and_Globulin_Ratio.fillna(data['Albumin_and_Globulin_Ratio'].mean())
data["Albumin_and_Globulin_Ratio"]
X=data.drop(['Gender','Target'],axis=1)
y=data['Target']
liver_corr = X.corr()
plt.figure(figsize=(18,18))
sns.heatmap(liver_corr, cbar = True, square = True, annot=True, fmt= '.2f',annot_kws={'size': 15},
cmap= 'coolwarm')
plt.title('Correlation between features');
sns.jointplot("Total_Protiens", "Albumin", data=data, kind="reg")
| Liver-EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install schedule
# +
from selenium import webdriver
import schedule
import time
import sqlite3
from bs4 import BeautifulSoup
def job():
driver = webdriver.Chrome('./chromedriver.exe')
conn = sqlite3.connect('./db.stock')
c = conn.cursor()
c.execute("CREATE TABLE IF NOT EXISTS article (id INTEGER PRIMARY KEY AUTOINCREMENT, date TEXT, time TEXT, title TEXT, content TEXT, press TEXT , href TEXT, stock TEXT, posi_nega TEXT)")
uri = 'https://finance.naver.com/news/market_special.nhn?&page='
# fnews = []
for page in range(1, 3):
target = uri+str(page)
driver.get(target)
try:
for n in range(1, 24):
# time.sleep(2)
driver.implicitly_wait(5)
try:
driver.find_element_by_xpath('//*[@id="contentarea_left"]/div[2]/table/tbody/tr[%s]/td[1]/a' %n).click()
html = driver.page_source
soup = BeautifulSoup(html, 'html.parser')
presses = soup.select('span.press > img')
press = presses[0]['title']
datas = soup.select('div.article_info > h3')
title = datas[0].text.strip()
dates = soup.select('div.article_sponsor > span')
date = dates[0].text.strip()
contents = soup.select('div#content')
content1 = contents[0]
for tag in content1.find_all(['li', 'h3']):
tag.replace_with('')
content = content1.text.strip()
driver.back()
html = driver.page_source
soup1 = BeautifulSoup(html, 'html.parser')
times = soup1.select('td.wdate')
time1 = times[int(n)-1]
time = time1.text.strip()[-5:]
hrefs = soup1.select('td.publicSubject > a')
href1 = hrefs[int(n)-1]
href = 'https://finance.naver.com'+ href1['href']
# a = [date , time , press , title, content, href]
# fnews.append(a)
c.execute("INSERT INTO article( date , time , title, content, press, href, stock , posi_nega ) VALUES(?,?,?,?,?,?,?,?)",(date, time, title, content, press, href,'stock','posi_nega'))
except:
pass
except:
pass
conn.commit()
c.close()
# print(fnews)
schedule.every().day.at("06:00").do(job)
while True:
schedule.run_pending()
time.sleep(5)
# -
| scraping/naver_finance_news_result_autoStart.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys, os
import argparse
sys.path.insert(0, os.path.abspath(os.path.join(os.getcwd(), "./../../../../")))
sys.path.insert(0, os.path.abspath(os.path.join(os.getcwd(), "./../../../")))
sys.path.insert(0, os.path.abspath(os.path.join(os.getcwd(), "")))
# +
from fedml_api.data_preprocessing.cifar10.data_loader import load_partition_data_cifar10
from fedml_api.standalone.fedavg.my_model_trainer_classification import MyModelTrainer as MyModelTrainerCLS
# from fedml_api.model.cv.resnet import resnet56
from fedml_api.model.contrastive_cv.resnet_with_embedding import Resnet56
import torch
from torch import nn
from collections import OrderedDict
import torch.nn.functional as F
import matplotlib.pyplot as plt
from sklearn.manifold import TSNE
import numpy as np
import random
import pickle
# +
dataset = 'cifar10'
data_dir = "./../../../data/cifar10"
partition_method = 'hetero'
# partition_method = 'homo'
partition_alpha = 100
client_num_in_total = 10
batch_size = 100
total_epochs = 500
save_model_path = 'model/client_{0}_triplet_epochs_{1}.pt'
device = 'cuda:1'
# +
train_data_num, test_data_num, train_data_global, test_data_global, \
train_data_local_num_dict, train_data_local_dict, test_data_local_dict, \
class_num,traindata_cls_counts = load_partition_data_cifar10(dataset, data_dir, partition_method,
partition_alpha, client_num_in_total, batch_size)
# -
dataset = [train_data_num, test_data_num, train_data_global, test_data_global, \
train_data_local_num_dict, train_data_local_dict, test_data_local_dict, \
class_num,traindata_cls_counts]
with open(f'dataset_{partition_method}_{partition_alpha}_{client_num_in_total}_with_cls_counts.pickle', 'wb') as f:
pickle.dump(dataset, f)
# pickle.save()
# +
# print(train_data_num)
# print(train_data_local_dict)
# -
with open(f'dataset_{partition_method}_{client_num_in_total}.pickle', 'rb') as f:
dataset = pickle.load(f)
class Client(object):
def __init__(self, client_index, train_data_local_dict, train_data_local_num_dict, test_data_local_dict, device, model):
self.id = client_index
self.train_data = train_data_local_dict[self.id]
self.local_sample_number = train_data_local_num_dict[self.id]
self.test_local = test_data_local_dict[self.id]
self.device = device
self.model = model
model = Resnet56(class_num=dataset[-1], neck='bnneck')
model.load_state_dict(torch.load(str.format('model/cs_{0}_{1}_client_{2}_triplet_epochs_{3}.pt', client_num_in_total, partition_method, 0, 300)))
# [train_data_num, test_data_num, train_data_global, test_data_global, \
# # train_data_local_num_dict, train_data_local_dict, test_data_local_dict, \
# # class_num]
# model.load_state_dict(torch.load(str.format('model/client_{0}_triplet_epochs_{1}.pt', 0, 399)))
client_1 = Client(0, dataset[5], dataset[4], dataset[6], device, model)
# client_1.train_data.shuffle = False
# +
def extract_features(model, data_loader,device):
model.to(device)
model.eval()
features = []
labels = []
with torch.no_grad():
for batch_idx, (x, l) in enumerate(data_loader):
x, l = x.to(device), l.to(device)
score, feats = model(x)
# print(feats.shape)
for feat, label in zip(feats, l):
features.append(feat.cpu())
labels.append(label.cpu())
return features, labels
features, labels = extract_features(client_1.model, client_1.test_local, device)
# +
def extract_cnn_feature(model, input, modules):
# model.eval()
outputs = OrderedDict()
handles = []
# outputs = None
for m in modules:
# print(id(m))
outputs[id(m)] = None
def func(m, i, o): outputs[id(m)] = o.data.cpu()
handles.append(module.register_forward_hook(func))
model(input)
for h in handles:
h.remove()
# print(outputs.values())
#return: [1, 64, 256, 8, 8]
return list(outputs.values())[0]
def extract_features(model, data_loader,device, module):
model.to(device)
model.eval()
features = []
labels = []
with torch.no_grad():
for batch_idx, (x, l) in enumerate(data_loader):
x, l = x.to(device), l.to(device)
#[batch_size, channel, w, h]
outputs = extract_cnn_feature(model, x, module)
# features.append(outputs)
# labels.append(labels)
# print(len(outputs))
for output, label in zip(outputs, l):
# features[batch_idx] = output
# labels[batch_idx] = label
# print(output.shape)
# output: [64, 256, 8, 8]
features.append(output.view(-1, 256*8*8))
labels.append(label)
# print(len(outputs))
# print(outputs[0].shape)
#[79, 64, 256, 8, 8]
return features, labels
module = None
for name, m in client_1.model.named_modules():
if name == 'layer3.5.relu':
module = m
print(module)
# features, labels = extract_features(client_1.model, test_data_global, device, [module])
features, labels = extract_features(client_1.model, client_1.train_data, device, [module])
# -
from collections import Counter
ls = np.array([int(l) for l in labels])
counter_result = Counter(ls)
print(Counter(ls))
# +
# class imbalance sample
# ks = counter_result.keys()
def get_index(lst, item):
return [i for i in range(len(lst)) if lst[i]==item]
sampled_idx = []
# ids = np.random.randint(0, 9, size=5)
# print(idx)
for k, v in counter_result.items():
# print(f'{k}-{v}')
idx = get_index(ls, k)
# print(len(idx))
# print(k)
sampled_idx.extend(random.sample(idx, 100 if v>=100 else 0))
# if k in ids:
# sampled_idx.extend(random.sample(idx, 100 if v>=100 else 0))
# print(len(sampled_idx))
# +
# print(shape(features))
# print(np.array(features[0][0]))
# print(features[0])
# print(np.array(features[0][0]).shape)
# print(torch.stack(labels).shape)
plot_features = torch.squeeze(torch.stack(features),dim=1)
plot_features = plot_features.numpy()[sampled_idx]
# for idx in sampled_idx:
# plt_f.append(plot_features[idx])
print(len(plot_features))
source_features = F.normalize(torch.from_numpy(plot_features), dim=1)
tsne = TSNE(n_components=2, init='pca', perplexity=30)
Y = tsne.fit_transform(source_features)
plt.scatter(Y[:, 0], Y[:, 1], c=ls[sampled_idx])
plt.show()
# -
model = Resnet56(class_num=dataset[-1], neck='bnneck')
model.load_state_dict(torch.load(str.format('model/cs_{0}_{1}_client_{2}_oral_epochs_{3}.pt', client_num_in_total, partition_method, 0, 200)))
# [train_data_num, test_data_num, train_data_global, test_data_global, \
# # train_data_local_num_dict, train_data_local_dict, test_data_local_dict, \
# # class_num]
# model.load_state_dict(torch.load(str.format('model/client_{0}_triplet_epochs_{1}.pt', 0, 399)))
client_2 = Client(0, dataset[5], dataset[4], dataset[6], device, model)
features2, labels2 = extract_features(client_2.model, client_2.test_local, device)
ls = np.array([int(l) for l in labels2])
counter_result = Counter(ls)
print(Counter(ls))
# +
# # class imbalance sample
# # ks = counter_result.keys()
# def get_index(lst, item):
# return [i for i in range(len(lst)) if lst[i]==item]
# sampled_idx = []
# # ids = np.random.randint(0, 9, size=5)
# # print(idx)
# for k, v in counter_result.items():
# # print(f'{k}-{v}')
# idx = get_index(ls, k)
# # print(len(idx))
# # print(k)
# sampled_idx.extend(random.sample(idx, 100 if v>=100 else 0))
# # if k in ids:
# # sampled_idx.extend(random.sample(idx, 100 if v>=100 else 0))
# # print(len(sampled_idx))
# +
# print(shape(features))
# print(np.array(features[0][0]))
# print(features[0])
# print(np.array(features[0][0]).shape)
# print(torch.stack(labels).shape)
plot_features = torch.squeeze(torch.stack(features2),dim=1)
plot_features = plot_features.numpy()[sampled_idx]
# for idx in sampled_idx:
# plt_f.append(plot_features[idx])
print(len(plot_features))
source_features = F.normalize(torch.from_numpy(plot_features), dim=1)
tsne = TSNE(n_components=2, init='pca', perplexity=30)
Y = tsne.fit_transform(source_features)
plt.scatter(Y[:, 0], Y[:, 1], c=ls[sampled_idx])
plt.show()
| fedml_experiments/distributed/contrastive_fed/.ipynb_checkpoints/Untitled2-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# In this example, we will be using
# - MNIST Dataset
# - Keras Library: which is a high-level neural networks API, written in Python and capable of running on top of either TensorFlow or Theano. It was developed with a focus on enabling fast experimentation.
# +
#Imports
from keras.datasets import mnist
import matplotlib.pyplot as plt
from keras.utils import to_categorical
from keras.models import Sequential
from keras.layers import Dense, Conv2D, Flatten
from keras.utils import plot_model
import keras
# -
(X_train,y_train),(X_test,y_test) = mnist.load_data()
plt.imshow(X_train[0])
X_train[0].shape
# +
# Reshape
## The Keras reshape function takes four arguments:
## number of training images, pixel size, and image depth—use 1 to indicate a grayscale image
X_train = X_train.reshape(60000,28,28,1)
X_test = X_test.reshape(10000, 28,28,1)
# +
# We will do One-hot encoding for the target variable
## one column for each classification, 10 columns will be created if 10 values are present,
### and one of them will contain "1"
y_train = to_categorical(y_train)
y_test = to_categorical(y_test)
print(y_train[0].shape)
y_train[0]
# -
# ## Feature Maps:
# - The most important idea in CNN is the moving filter, which passes through the image
# - The weights within the filter could be any combination of values depending on how the filters are trained.
# - These weights are held constant as the filter moves throughout the image. This way it learns to identify features such as lines, edges or a specific shape.
# ## Pooling:
# It is similar to the feature maps, just that instead of a sliding window of matrix, we here apply some statistical function over the values in the window. Example: Max-pooling means it takes he maximun value from the pool, other examples could be mean pooling etc.
# - It reduces the parameters int he network, also known as downsampling
# - To make feature detection more robust, we make it inaffective to scale and orientation shanges
# ## Strides and down-sampling:
# So above we saw that the feature maps go 1 block in X or Y direction, which leds to an overlap of filter areas. With Max pooling the stride is usually set so that there is no overlap.
# ## Padding
# Pooling is used in convolutional neural networks to make the detection of certain features in the input invariant to scale and orientation changes.
# - The network shoudl detect the Image no matter the oriwntation of it, we need such a neural network. This is what padding helps us with.
# - pooling coupled with convolutional filters attempt to detect objects within an image.
# ## Callbacks
# Callbacks in Keras are objects that are called at different points during training (at the start of an epoch, at the end of a batch, at the end of an epoch, etc. We can use it to create checkpoints which saves model at different stages in the training to help in avoiding the work loss. It is passed to the **.fit()** function
#
# ### The model type that we will be using is Sequential. Sequential is the easiest way to build a model in Keras. It allows you to build a model layer by layer.
# +
# create model using Sequential
model = Sequential()
# +
# adding model layers
# the first two layers are Conv2D—2-dimensional convolutional layers, our input images, which are seen as 2-dimensional matrices.
## it takes 4 arguments
## Number of neural nodes per layer, kernel size, activation, input_shape (1 is for grey_scale)
## we have 64 in first layer, 32 in 2nd layer, which can be adjusted higer or lower according to dataset
## The Kernel size is the size of the filter matrix for our convolution,
## kernel size of 3 means we will have a 3x3 filter matrix.
## Activation function used is ReLU, or Rectified Linear Activation
## "Softmax" activation makes the output sum up to 1 so the output can be interpreted as probabilities
## input_shape is (28,28,1) - where 1 signifies greyscale
model.add(Conv2D(64, kernel_size =3, activation = "relu", input_shape= (28,28,1) ))
model.add(Conv2D(32, kernel_size =3, activation = "relu"))
# Add a ‘Flatten’ layer,
## which takes the output of the two convolution layers
## and turns it into a format that can be used by the final, densely connected neural layer.
model.add(Flatten())
# Add the final layer of type ‘Dense’, a densely-connected neural layer which will generate the final prediction
# Parameters: Number of output nodes—10, activation function - 10
model.add(Dense(10, activation='softmax'))
# -
# **Note**: Each of the convolution layers reduces the depth, width and height of each feature, this is equivalent to the pooling/downsampling stage in the CNN model.
# +
# now we complile the model
## Compiling the model takes three parameters: optimizer, loss and metrics.
## adam is a good optimizer, adjusts the learning rate through training
#compile model using accuracy to measure model performance
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
# +
#train the model
# parameters: training data (train_X), target data (train_y), validation data, and the number of epochs.
## epochs is the number of times the model will cycle through the data.
model.fit(X_train, y_train, validation_data=(X_test, y_test), epochs=3)
# -
# Summary of the model
model.summary()
# +
## check the predictions
#predict first 4 images in the test set
model.predict(X_test[:4])
# -
import matplotlib.pyplot as plt
plt.plot(hist.history["acc"])
plt.plot(hist.history['val_acc'])
plt.plot(hist.history['loss'])
plt.plot(hist.history['val_loss'])
plt.title("model accuracy")
plt.ylabel("Accuracy")
plt.xlabel("Epoch")
plt.legend(["Accuracy","Validation Accuracy","loss","Validation Loss"])
plt.show()
| Build a CNN for MNIST Dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="m8U-WGj90Jl7"
# # CNN Training Loop Refactoring (Simultaneous Hyperparameter Testing)
# + [markdown] id="pC6QAjtSpLm6"
# When we [last](https://saptarshidatta.in/2020/10/06/PyTorch_CIFAR10_TB.html) trained our network, we built out quite a lot of functionality that allowed us to experiment with many different parameters and values, and we also made the calls need inside our training loop that would get our results into TensorBoard.
#
# All of this work has helped, but our training loop is quite crowded now. In this exercise, we're going to clean up our training loop and set the stage for more experimentation up by using the `RunBuilder` class that we built last time and by building a new class called `RunManager`.
#
# I also find this way of Hyperparameter Tuning more intuitive than TensorBoard. Also, as our number of parameters and runs get larger, TensorBoard will start to breakdown as a viable solution for reviewing our results.
#
# However, calls have been made inside our `RunManager` class to TensorBoard, so it can be used as an added functionality. For reference, on how to use TensorBoard with PyTorch inside Google Collab, plese refer [here](https://saptarshidatta.in/2020/10/06/PyTorch_CIFAR10_TB.html).
#
# The code also generates results in `csv` and `json` format, which can be used gor further analysis.
# + id="28sB7uXPimG8"
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import torchvision
import torchvision.transforms as transforms
from torch.utils.data import DataLoader
from torch.utils.tensorboard import SummaryWriter
from IPython.display import display, clear_output
import pandas as pd
import time
import json
from itertools import product
from collections import namedtuple
from collections import OrderedDict
# + [markdown] id="uIoI5SyHtAHH"
# ## Designing the Neural Network
# + id="JeGzwSVqipb1"
class Network(nn.Module):
def __init__(self):
super(Network,self).__init__()
self.conv1 = nn.Conv2d(in_channels=3, out_channels=6, kernel_size=5)
self.conv2 = nn.Conv2d(in_channels=6, out_channels=16, kernel_size=5)
self.fc1 = nn.Linear(in_features=16*5*5, out_features=120)
self.fc2 = nn.Linear(in_features=120, out_features=84)
self.out = nn.Linear(in_features=84, out_features=10)
def forward(self, t):
#Layer 1
t = t
#Layer 2
t = self.conv1(t)
t = F.relu(t)
t = F.max_pool2d(t, kernel_size=2, stride=2)#output shape : (6,14,14)
#Layer 3
t = self.conv2(t)
t = F.relu(t)
t = F.max_pool2d(t, kernel_size=2, stride=2)#output shape : (16,5,5)
#Layer 4
t = t.reshape(-1, 16*5*5)
t = self.fc1(t)
t = F.relu(t)#output shape : (1,120)
#Layer 5
t = self.fc2(t)
t = F.relu(t)#output shape : (1, 84)
#Layer 6/ Output Layer
t = self.out(t)#output shape : (1,10)
return t
# + [markdown] id="LyhvwLLar0YM"
# ## `RunBuilder` class
# + id="M8MekfxKi0v6"
class RunBuilder():
@staticmethod
def get_runs(params):
Run = namedtuple('Run', params.keys())
runs = []
for v in product(*params.values()):
runs.append(Run(*v))
return runs
# + [markdown] id="3_JggKs1sG30"
# ## `RunManager` class
# + id="y8opwcSpi7vy"
class RunManager():
def __init__(self):
self.epoch_count = 0
self.epoch_loss = 0
self.epoch_num_correct = 0
self.epoch_start_time = None
self.run_params = None
self.run_count = 0
self.run_data = []
self.run_start_time = None
self.network = None
self.loader = None
self.tb = None
def begin_run(self, run, network, loader):
self.run_start_time = time.time()
self.run_params = run
self.run_count += 1
self.network = network
self.loader = loader
self.tb = SummaryWriter(comment=f'-{run}')
images, labels = next(iter(self.loader))
grid = torchvision.utils.make_grid(images)
self.tb.add_image('images', grid)
self.tb.add_graph(
self.network
,images.to(getattr(run, 'device', 'cpu'))
)
def end_run(self):
self.tb.close()
self.epoch_count = 0
def begin_epoch(self):
self.epoch_start_time = time.time()
self.epoch_count += 1
self.epoch_loss = 0
self.epoch_num_correct = 0
def end_epoch(self):
epoch_duration = time.time() - self.epoch_start_time
run_duration = time.time() - self.run_start_time
loss = self.epoch_loss / len(self.loader.dataset)
accuracy = self.epoch_num_correct / len(self.loader.dataset)
self.tb.add_scalar('Loss', loss, self.epoch_count)
self.tb.add_scalar('Accuracy', accuracy, self.epoch_count)
for name, param in self.network.named_parameters():
self.tb.add_histogram(name, param, self.epoch_count)
self.tb.add_histogram(f'{name}.grad', param.grad, self.epoch_count)
results = OrderedDict()
results["run"] = self.run_count
results["epoch"] = self.epoch_count
results['loss'] = loss
results["accuracy"] = accuracy
results['epoch duration'] = epoch_duration
results['run duration'] = run_duration
for k,v in self.run_params._asdict().items(): results[k] = v
self.run_data.append(results)
df = pd.DataFrame.from_dict(self.run_data, orient='columns')
clear_output(wait=True)
display(df)
def track_loss(self, loss, batch):
self.epoch_loss += loss.item() * batch[0].shape[0]
def track_num_correct(self, preds, labels):
self.epoch_num_correct += self._get_num_correct(preds, labels)
def _get_num_correct(self, preds, labels):
return preds.argmax(dim=1).eq(labels).sum().item()
def save(self, fileName):
pd.DataFrame.from_dict(
self.run_data
,orient='columns'
).to_csv(f'{fileName}.csv')
with open(f'{fileName}.json', 'w', encoding='utf-8') as f:
json.dump(self.run_data, f, ensure_ascii=False, indent=4)
# + [markdown] id="qMWgRpmrscfT"
# ## Loading the CIFAR-10 data and pre-processing
# + id="zl2SMDGVi_OK" outputId="c6b24208-acab-41cd-9b28-43bd7d05cc6d" colab={"base_uri": "https://localhost:8080/", "height": 34}
transform =transforms.Compose([transforms.ToTensor(), transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
train_set = torchvision.datasets.CIFAR10(
root='./data'
,train=True
,download=True
,transform=transform
)
# + [markdown] id="XVvHIScdsM5j"
# ## Training the Nueral Network
# + id="FxpoZAwyjAim" outputId="eb650cf8-8f7a-47d3-9836-5dabeb779b67" colab={"base_uri": "https://localhost:8080/", "height": 235}
params = OrderedDict(
lr = [.01]
,batch_size = [1000]
,shuffle = [True]
,num_workers = [0, 1, 2, 4, 8, 16]
)
m = RunManager()
for run in RunBuilder.get_runs(params):
network = Network()
loader = DataLoader(train_set, batch_size=run.batch_size, shuffle=run.shuffle, num_workers=run.num_workers)
optimizer = optim.Adam(network.parameters(), lr=run.lr)
m.begin_run(run, network, loader)
for epoch in range(1):
m.begin_epoch()
for batch in loader:
images, labels = batch
preds = network(images) # Pass Batch
loss = F.cross_entropy(preds, labels) # Calculate Loss
optimizer.zero_grad() # Zero Gradients
loss.backward() # Calculate Gradients
optimizer.step() # Update Weights
m.track_loss(loss, batch)
m.track_num_correct(preds, labels)
m.end_epoch()
m.end_run()
m.save('results')
# + [markdown] id="c_kqfMJJppzH"
# ## Experimenting with DataLoader `num_workers` attribute
#
# The `num_workers` attribute tells the data loader instance how many sub-processes to use for data loading. By default, the `num_workers` value is set to zero, and a value of zero tells the loader to load the data inside the main process.
#
# This means that the training process will work sequentially inside the main process. After a batch is used during the training process and another one is needed, we read the batch data from disk.
#
# Now, if we have a worker process, we can make use of the facility that our machine has multiple cores. This means that the next batch can already be loaded and ready to go by the time the main process is ready for another batch. This is where the speed up comes from. The batches are loaded using additional worker processes and are queued up in memory.
#
# The main take-away from these results is that having a single worker process in addition to the main process resulted in a speed up of about twenty percent. However, adding additional worker processes after the first one didn't really show any further improvements.
#
# Additionally, we can see with higher number of num_workers results in higher run times. Please go through this [link](https://discuss.pytorch.org/t/guidelines-for-assigning-num-workers-to-dataloader/813/5) to know more.
# + [markdown] id="A_aSpj25uKtd"
# ## Summary
#
# We have introduced a way to experiment with Hyperparameters to extract maximum efficiency for our model. This code can be scaled up or scaled down to change the Hyperparameters we wish to experiment upon.
#
# This may be noted that, accuracy is not that high as we have trained our model for 1 epoch with each set of parameters. This has been purely done for experimentation purpose.
#
# However, we might need to change our network architecture i.e. a deeper network for higher efficiency.
| _notebooks/2020-10-11-PyTorch_CIFAR_10_Refactoring.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from ipysheet import from_array, to_array
arr = np.random.randn(6, 10)
sheet = from_array(arr)
sheet
# +
arr = np.array([True, False, True])
sheet = from_array(arr)
sheet
# -
to_array(sheet)
| examples/numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Hidden Markov Model (and Filters) notes
# Ref: meta_concepts_probabilistic.ipynb
# * Used to analyze or predict time series (which involves noise/uncertainty)
# * Uses Bayes Nets/Markov Model
# * Markov Chain
# * Stationary Distribution
# * Maximum Likelihood
# * Laplacian Smoothing
# * HMM
# * Prediction
# * State Estimation
| ai_hidden_markov_models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Pyplot Mathtext
#
#
# Use mathematical expressions in text labels. For an overview over MathText
# see :doc:`/tutorials/text/mathtext`.
#
# +
import numpy as np
import matplotlib.pyplot as plt
t = np.arange(0.0, 2.0, 0.01)
s = np.sin(2*np.pi*t)
plt.plot(t, s)
plt.title(r'$\alpha_i > \beta_i$', fontsize=20)
plt.text(1, -0.6, r'$\sum_{i=0}^\infty x_i$', fontsize=20)
plt.text(0.6, 0.6, r'$\mathcal{A}\mathrm{sin}(2 \omega t)$',
fontsize=20)
plt.xlabel('time (s)')
plt.ylabel('volts (mV)')
plt.show()
# -
# ------------
#
# References
# """"""""""
#
# The use of the following functions, methods, classes and modules is shown
# in this example:
#
#
import matplotlib
matplotlib.pyplot.text
matplotlib.axes.Axes.text
| matplotlib/gallery_jupyter/pyplots/pyplot_mathtext.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import os
import json
import glob
import sqlite3, sqlalchemy
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.linear_model import LinearRegression
from sklearn.ensemble import RandomForestRegressor
from sklearn.feature_extraction.text import CountVectorizer
# %matplotlib inline
#utiliser le json indiquer les dossiers dans lesquels l'on veut travailler
with open('../parameters.json') as json_data:
params = json.load(json_data)
DATA_DIR = params['DIRECTORIES']['DATA_DIR']
TMP_DIR = params['DIRECTORIES']['TMP_DIR']
PRICE_FILES = params['FILES']['PRICE_FILE']
STATION_FILES = params['FILES']['STATION_FILE']
SERVICE_FILES = params['FILES']['SERVICE_FILE']
GAS_DB = os.path.join(DATA_DIR, params['DATABASES']['GAS_DB'])
params
dfall = pd.read_pickle(os.path.join(DATA_DIR, PRICE_FILES))
dfstations = pd.read_pickle(os.path.join(DATA_DIR, STATION_FILES))
dfservices = pd.read_pickle(os.path.join(DATA_DIR, SERVICE_FILES))
dfall['date_releve'] = pd.to_datetime(dfall.date_releve)
moyenne_hebdo_df = (dfall
.dropna(subset=['date_releve'])
.groupby(['libele_carburant', pd.Grouper(key='date_releve', freq='1w')])
.agg({'prix': np.mean})).reset_index()
model_df = (dfall
.dropna(subset=['date_releve'])
.groupby(['id_station', 'code_postal','type_station','libele_carburant', pd.Grouper(key='date_releve', freq='1w')])
.agg({'prix': np.mean})
.reset_index()
.merge(moyenne_hebdo_df,how='inner', on=['libele_carburant','date_releve'],suffixes=('_station','_moyen'))
.assign(indice = lambda _df: _df.prix_station/_df.prix_moyen)
)
model_df.shape
model_df.head()
model_df.indice.describe()
model_df.indice.hist(bins=50)
#### Construction du vecteur de service.
_tokenize = lambda x : x.split(',')
vectorize = CountVectorizer(tokenizer=_tokenize, lowercase=False)
vectorize.fit_transform(dfservices.type_service.fillna('Rien'))
### Créer un sparse dataFrame pour les types de services proposés par la station.
matrice_services_df = pd.SparseDataFrame(vectorize.fit_transform(dfservices.type_service.fillna('Rien')),
dfservices.id_station,
vectorize.get_feature_names(),
default_fill_value=0
).reset_index()
matrice_services_df.head()
model_service_df = model_df.merge(matrice_services_df, how='inner', on=['id_station'])
model_df.merge(matrice_services_df, how='inner', on=['id_station']).head()
# +
(model_df
.merge(matrice_services_df, how='right', on=['id_station'], indicator=True)
['_merge']
.value_counts())
# -
#### Régression
from sklearn.linear_model import LinearRegression
lr = LinearRegression()
lr.fit(X=model_service_df[list(set(matrice_services_df.columns)-{'id_station'})], y=model_service_df.indice)
lr.coef_
[(f,c) for f, c in zip(list(set(matrice_services_df.columns)-{'id_station'}), lr.coef_)]
### Histogramme des résultats
resultats = pd.DataFrame(
[(f,c) for f, c in zip(list(set(matrice_services_df.columns)-{'id_station'}), lr.coef_)],
columns=['type_service', 'coefficients']).sort_values('coefficients', ascending=False)
resultats.set_index('type_service').plot(kind='barh')
### Utiliser une autre méthode que la régression linéaire, mais en random forest
rf = RandomForestRegressor()
rf.fit(X=model_service_df[list(set(matrice_services_df.columns)-{'id_station'})], y=model_service_df.indice)
rf.feature_importances_
resultats_rf = pd.DataFrame(
[(f,c) for f, c in zip(list(set(matrice_services_df.columns)-{'id_station'}), rf.feature_importances_)],
columns=['type_service', 'coefficients']).sort_values('coefficients', ascending=False)
resultats_rf.set_index('type_service').plot(kind='barh')
| src/Transformation/Transformation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
dataset = pd.read_csv('datasets/Churn_Modelling.csv')
dataset.head()
X = dataset.iloc[:, 3:13].values
y = dataset.iloc[:, 13].values
from sklearn.preprocessing import LabelEncoder, OneHotEncoder
labelencoder_X_1 = LabelEncoder()
X[:, 1] = labelencoder_X_1.fit_transform(X[:, 1])
labelencoder_X_2 = LabelEncoder()
X[:, 2] = labelencoder_X_2.fit_transform(X[:, 2])
onehotencoder = OneHotEncoder(categorical_features = [1])
X = onehotencoder.fit_transform(X).toarray()
X = X[:, 1:]
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2)
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
import keras
from keras.models import Sequential
from keras.layers import Dense
classifier = Sequential()
# 11 input neurons
classifier.add(Dense(units = 6, kernel_initializer = 'uniform', activation = 'relu', input_dim = 11))
# 6 hidden neurons
classifier.add(Dense(units = 6, kernel_initializer = 'uniform', activation = 'relu'))
# 1 output neuron
classifier.add(Dense(units = 1, kernel_initializer = 'uniform', activation = 'sigmoid'))
classifier.compile(optimizer = 'adam', loss = 'binary_crossentropy', metrics = ['accuracy'])
classifier.fit(X_train, y_train, batch_size = 10, nb_epoch = 100)
y_pred = classifier.predict(X_test)
y_pred = (y_pred > 0.5)
from sklearn.metrics import accuracy_score
score = accuracy_score(y_test, y_pred)
score
| experiments/tests/ANN vs PGSO/.ipynb_checkpoints/Churn prediction with ANN-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import warnings
warnings.filterwarnings('ignore') # Hide warnings
import datetime as dt
import pandas as pd
pd.core.common.is_list_like = pd.api.types.is_list_like
import pandas_datareader.data as web
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from mpl_finance import candlestick_ohlc
import matplotlib.dates as mdates
# +
#Getting stock price data
#SRCL
start = dt.datetime(2020, 1, 1)
end = dt.datetime.now()
df = web.DataReader("SRCL", 'yahoo', start, end)
df.reset_index(inplace=True)
df.set_index("Date", inplace=True)
#prices in USD
# -
df.head()
plt.figure(figsize=(12, 8))
plt.plot(df[["High","Low","Open","Close"]])
plt.legend(labels=df[["High","Low","Open","Close"]])
plt.title('Stock Trend 2020', fontsize = 12)
plt.xlabel('Time')
plt.ylabel('Stock Value')
# +
df_open_close= df.copy()
df_open_close['Close_10ma'] = df['Close'].rolling(window=10,min_periods=0).mean()
df_open_close['Open_10ma'] = df['Open'].rolling(window=10,min_periods=0).mean()
df_open_close.head()
# -
plt.figure(figsize=(20, 12))
plt.plot(df_open_close[["Close_10ma","Open_10ma","Open","Close"]])
plt.legend(labels=df_open_close[["Close_10ma","Open_10ma","Open","Close"]])
plt.title('Stock Trend Open Close 2020 10 Day MA', fontsize = 20)
plt.xlabel('Time')
plt.ylabel('Stock Value')
# +
#40days Moving Average
df_open_close['Close_40ma'] = df['Close'].rolling(window=40,min_periods=0).mean()
df_open_close['Open_40ma'] = df['Open'].rolling(window=40,min_periods=0).mean()
df_open_close.head()
# -
plt.figure(figsize=(20, 12))
plt.plot(df_open_close[["Close_40ma","Open_40ma","Open","Close"]])
plt.legend(labels=df_open_close[["Close_40ma","Open_40ma","Open","Close"]])
plt.title('Stock Trend Open Close 2020 40 Day MA', fontsize = 20)
plt.xlabel('Time')
plt.ylabel('Stock Value')
| Stericycle/SRCL Stock Trend 2020.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Examples for AbsLine class (v1.6)
# +
# suppress warnings for these examples
import warnings
warnings.filterwarnings('ignore')
# import
import astropy.units as u
from linetools.spectralline import AbsLine, SpectralLine
from linetools import spectralline as ltsp
from linetools import line_utils as ltlu
from linetools.spectra import io as lsio
from linetools.spectra.xspectrum1d import XSpectrum1D
# -
# ## Generate a line
abslin = AbsLine(1548.195*u.AA)
abslin
# ### Data
abslin.data
# ### As dict
abslin = AbsLine(1548.195*u.AA)
tmp = abslin.to_dict()
tmp
# ### From dict
tmp2 = SpectralLine.from_dict(tmp)
tmp2
# ### From old dict (using analy['vlim']) -- Backwards compatability
tmp.pop('limits')
tmp['analy']['vlim'] = [-150,150]*u.km/u.s
tmp['attrib']['z'] = 0.5
tmp3 = SpectralLine.from_dict(tmp)
# ## List of AbsLines
abslines = [abslin, AbsLine('HI 1215')]
# ### Parse
wrest_values = ltlu.parse_speclines(abslines, 'wrest')
wrest_values
# ### Transition Table
tbl = ltlu.transtable_from_speclines(abslines)
tbl
# ## Measure an EW
# Set spectrum
abslin.analy['spec'] = XSpectrum1D.from_file('../../linetools/spectra/tests/files/UM184_nF.fits')
# Set analysis range
abslin.limits.set([6080.78, 6087.82]*u.AA) # wvlim
# Measure
abslin.measure_ew() # Observer frame
print('EW = {:g} with error {:g}'.format(abslin.attrib['EW'],abslin.attrib['sig_EW']))
# ## Measure AODM
#
abslin.analy['spec'] = lsio.readspec('../../linetools/spectra/tests/files/UM184_nF.fits')
abslin.setz(2.92929)
abslin.limits.set((-150., 150.)*u.km/u.s) # vlim
abslin.measure_aodm()
N, sigN, flgN = [abslin.attrib[key] for key in ['N','sig_N','flag_N']]
print('logN = {:g}, siglogN = {:g}'.format(abslin.attrib['logN'], abslin.attrib['sig_logN']))
# ### Another Table
tbl = ltlu.transtable_from_speclines(abslines)
tbl
| docs/examples/AbsLine_examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
pip install opencv-python
import cv2
# # Color to Black and White image
img = cv2.imread('child.jpg')
#to show the image
cv2.imshow('img',img)
cv2.waitKey(0)
# to convert color to black and white
out = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
# +
# TO SHOW THE converted image
# -
cv2.imshow("out",out)
cv2.waitKey(0)
# # Resizing the image
img = cv2.imread('man.png')
img.shape
#Using dsize
out = cv2.resize(img,(600,600))
out.shape
# +
#Using fx and fy
# -
out1 = cv2.resize(img,None,fx=2,fy=2)
out1.shape
# # Noise Removal
# + active=""
# Salt and Pepper Removal Technique :- Median filtering
# Gaussian Noise Removal Technique :- Gaussian Blurring
# -
# salt-and-pepper-noise
img = cv2.imread('salt-and-pepper-noise.png')
cv2.imshow('img',img)
cv2.waitKey(0)
out = cv2.medianBlur(img,5) #5 is the kernal size
# # Skew Correction
img = cv2.imread('skew.png')
cv2.imshow('img',img)
cv2.waitKey(0)
# Detect the box
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
ret, thresh = cv2.threshold(gray,0,255,cv2.THRESH_BINARY_INV + cv2.THRESH_OTSU)
contours, _ = cv2.findContours(thresh, cv2.RETR_LIST,cv2.CHAIN_APPROX_SIMPLE)
contours = sorted(contours, key = cv2.contourArea, reverse = True)
max_cnt = contours[0]
# Find the skew angle
angle = cv2.minAreaRect(max_cnt)[-1]
if angle < -45:
angle = 90 + angle
angle
# +
# Re-rotatation of image
height, width, _ =img.shape
center = (width//2, height//2)
M = cv2.getRotationMatrix2D(center, angle,1)
dst = cv2.warpAffine(img, M, (width,height),flags = cv2.INTER_LINEAR,borderMode = cv2.BORDER_REPLICATE)
# -
cv2.imshow('img',img)
cv2.waitKey(0)
| Image Preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Overview
#
#
# - **4.0. Overfitting and Underfitting**
# - 4.0.1. *Network size*
# - 4.0.2. *Weights regularization*
# - 4.0.3. *Dropouts*
# <br><br>
#
# - **4.1. TensorboardX**
#
#
#
#
#
# # 4.0. Over-/Underfitting
#
# **Optimization:** process of adjusting a model to get the best performance possible on the training data
#
# **Generalization:** how well trained model performs on data it has never seen before
#
# #### What to do when model is underfitting (not optimal)?
# - Train longer
#
# #### How to prevent overfitting (i.e. generalize)?
# - Get more data
# - Modulate quantity of information fed to model
#
#
#
# # 4.0.1. Network Size
#
#
# > *“A **small network**, with say one hidden unit is **likely to be biased**, since the repertoire of available functions spanned by 𝑓(𝑥,𝑤) over allowable weights will in this case be quite limited.”*
#
# > “if we **overparameterize, via a large number of hidden units and associated weights**, then bias will be reduced (… with enough weights and hidden units, the network will interpolate the data) but there is then the **danger of significant variance** contribution to the mean-square error”
#
#
# ([German et al. 1992](http://web.mit.edu/6.435/www/Geman92.pdf))
#
# Managing the size/capacity of your network in the model, more accurately the no. of learnable parameters (i.e. no. of layers x no. of units per layer), will (i) ***optimize the network to converge faster*** and/or (ii) ***prevent the network from overfitting to the training data***. To illustrate this point, lets go back to our XOR example.
#
# +
from tqdm import tqdm
import numpy as np
import torch
from torch import nn
from torch import tensor
from torch import optim
#from tensorboardX import SummaryWriter
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style("darkgrid")
sns.set(rc={'figure.figsize':(12, 8)})
torch.manual_seed(42)
device = 'cuda' if torch.cuda.is_available() else 'cpu'
# XOR gate inputs and outputs.
X = xor_input = tensor([[0,0], [0,1], [1,0], [1,1]]).float().to(device)
Y = xor_output = tensor([[0],[1],[1],[0]]).float().to(device)
# Use tensor.shape to get the shape of the matrix/tensor.
num_data, input_dim = X.shape
print('Inputs Dim:', input_dim) # i.e. n=2
num_data, output_dim = Y.shape
print('Output Dim:', output_dim)
print('No. of Data:', num_data) # i.e. n=4
# -
# ## This time lets try different hidden dimensions!
# +
def MLP(hidden_dim, num_epochs=5000):
# Step 1: Initialization.
# Use Sequential to define a simple feed-forward network.
model = nn.Sequential(
# Use nn.Linear to get our simple perceptron.
nn.Linear(input_dim, hidden_dim),
# Use nn.Sigmoid to get our sigmoid non-linearity.
nn.Sigmoid(),
# Second layer neurons.
nn.Linear(hidden_dim, output_dim),
nn.Sigmoid()
).to(device)
# Initialize the optimizer
learning_rate = 0.3
optimizer = optim.SGD(model.parameters(), lr=learning_rate)
# Initialize the loss function.
criterion = nn.MSELoss()
# Initialize the stopping criteria
# For simplicity, just stop training after certain no. of epochs.
losses = [] # Keeps track of the loses.
# Step 2-4 of training routine.
for _e in tqdm(range(num_epochs)):
# Reset the gradient after every epoch.
optimizer.zero_grad()
# Step 2: Foward Propagation
predictions = model(X)
# Step 3: Back Propagation
# Calculate the cost between the predictions and the truth.
loss = criterion(predictions, Y)
# Remember to back propagate the loss you've computed above.
loss.backward()
# Step 4: Optimizer take a step and update the weights.
optimizer.step()
# Log the loss value as we proceed through the epochs.
if _e % 100 == 0:
losses.append(loss.data.item())
return losses
losses_10 = MLP(hidden_dim=10)
losses_100 = MLP(hidden_dim=100)
losses_200 = MLP(hidden_dim=200)
# -
#
#
# ## This seems to fit what we learn,
#
# Small network with limited memorization capacity underfits and takes longer to coverge (i.e. find the minimum loss).
# +
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
sns.set_style("darkgrid")
sns.set(rc={'figure.figsize':(12, 8)})
# -
plt.plot(losses_10, label='hidden = 10')
plt.plot(losses_100, label='hidden = 100')
plt.plot(losses_200, label='hidden = 200')
plt.legend(loc='upper right')
plt.show()
# ## Now, let's try some larger no. of units
losses_300 = MLP(hidden_dim=300)
losses_1000 = MLP(hidden_dim=1000)
losses_2000 = MLP(hidden_dim=2000)
# ## Hey now the curves of larger networks look funky =)
#
# Larger network will converge faster until a certain capacity sweetspot; `hidden_dim=200` converges faster than `hidden_dim=100`, and once it gets to `hidden>=300`, the training loss gets funky.
plt.plot(losses_10, label='hidden = 10')
plt.plot(losses_100, label='hidden = 100')
plt.plot(losses_200, label='hidden = 200')
plt.plot(losses_300, label='hidden = 300')
plt.plot(losses_1000, label='hidden = 1000')
plt.plot(losses_2000, label='hidden = 2000')
plt.legend(loc='upper right')
plt.show()
plt.scatter(range(len(losses_1000)), losses_1000, label='hidden = 1000', color='purple')
plt.scatter(range(len(losses_2000)), losses_2000, label='hidden = 2000', color='brown')
plt.legend(loc='lower left')
plt.show()
#
#
# ## Summary:
#
# 1. Small network <u>underfits</u> (longer to converge)
# 2. larger network easier to converge
# 3. Too large network <u>overfits</u>
# 4. Too too large network don't train properly at all.
# # 4.0.2. Weights regularization
#
#
# <img src="http://3.bp.blogspot.com/_oqH68z1KYWk/Stc15KGFpCI/AAAAAAAAARI/ARojEj9rH7o/s1600/occams_professor.gif" width="500" align="left">
#
# The simpler solution is better because makes fewer assumptions. Simpler models are less likely to overfit than complex ones.
#
# By adding a cost associated with large weights, **weights regularization** forces the model weights to take smaller values.
#
# - ***L1 regularization***: Cost is proportional to the absolute value of the weight coefficient.
#
# - ***L2 regularization***: Cost is proportional to the square value of the weight coefficient. (L2 regularization is also call *weight decay*
#
# In `PyTorch`, the regularization is a parameter in the optimizer, e.g.
#
# ```python
# # Using the L2 regularization.
# optimizer = optim.SGD(model.parameters(), lr=learning_rate, weight_decay=1e-5)
# ```
#
# IMHO, in practice, I (personally) never tune weights regularization. Other regularization methods gives you better result. L1 Loss is messy in PyTorch, so we'll skip that for now =)
#
def MLP_L2Loss(hidden_dim, num_epochs=5000, weight_decay=1e-4):
# Step 1: Initialization.
# Use Sequential to define a simple feed-forward network.
model = nn.Sequential(
# Use nn.Linear to get our simple perceptron.
nn.Linear(input_dim, hidden_dim),
# Use nn.Sigmoid to get our sigmoid non-linearity.
nn.Sigmoid(),
# Second layer neurons.
nn.Linear(hidden_dim, output_dim),
nn.Sigmoid()
).to(device)
# Initialize the optimizer
learning_rate = 0.3
optimizer = optim.SGD(model.parameters(), lr=learning_rate, weight_decay=weight_decay)
# Initialize the loss function.
criterion = nn.MSELoss()
# Initialize the stopping criteria
# For simplicity, just stop training after certain no. of epochs.
losses = [] # Keeps track of the loses.
# Step 2-4 of training routine.
for _e in tqdm(range(num_epochs)):
# Reset the gradient after every epoch.
optimizer.zero_grad()
# Step 2: Foward Propagation
predictions = model(X)
# Step 3: Back Propagation
# Calculate the cost between the predictions and the truth.
loss = criterion(predictions, Y)
# Remember to back propagate the loss you've computed above.
loss.backward()
# Step 4: Optimizer take a step and update the weights.
optimizer.step()
# Log the loss value as we proceed through the epochs.
if _e % 100 == 0:
losses.append(loss.data.item())
return losses
# Not much different, but subjected to some randomness...
loss_normal = MLP(200)
loss_l2 = MLP_L2Loss(200)
plt.plot(loss_normal, label='normal')
plt.plot(loss_l2, label='L2 loss')
plt.legend(loc='upper right')
plt.show()
# # 4.0.3. Dropout
#
# **Dropout** is the poster boy of neural network regularization. <NAME> on bank's fraud prevention mechanism:
#
# > *"I went to my bank. The tellers kept changing and I asked one of them why. He said he didn't know but they got moved around a lot.
# >
# > I figured it must be because it would require cooperation between employees to succesfully defraud the bank.
# >
# > This made me realize that randomly removing a difference subsset of neurons on each example would prevent conspiracies and thus reduce overfitting"
#
# [<NAME> (in Reddit AMA)](https://www.reddit.com/r/MachineLearning/comments/4w6tsv/ama_we_are_the_google_brain_team_wed_love_to/)
#
#
# To prevent the model from over-memorizing the data, adding random zero-ing noise to the output values of the layer will prevent *"conspiracies among neurons"*.
def MLP_Dropout(hidden_dim, num_epochs=5000, dropout=0.1):
# Step 1: Initialization.
# Use Sequential to define a simple feed-forward network.
model = nn.Sequential(
# Use nn.Linear to get our simple perceptron.
nn.Linear(input_dim, hidden_dim),
nn.Dropout(dropout),
# Use nn.Sigmoid to get our sigmoid non-linearity.
nn.Sigmoid(),
# Second layer neurons.
nn.Linear(hidden_dim, output_dim),
nn.Sigmoid()
).to(device)
# Initialize the optimizer
learning_rate = 0.3
optimizer = optim.SGD(model.parameters(), lr=learning_rate)
# Initialize the loss function.
criterion = nn.MSELoss()
# Initialize the stopping criteria
# For simplicity, just stop training after certain no. of epochs.
losses = [] # Keeps track of the loses.
# Step 2-4 of training routine.
for _e in tqdm(range(num_epochs)):
# Reset the gradient after every epoch.
optimizer.zero_grad()
# Step 2: Foward Propagation
predictions = model(X)
# Step 3: Back Propagation
# Calculate the cost between the predictions and the truth.
loss = criterion(predictions, Y)
# Remember to back propagate the loss you've computed above.
loss.backward()
# Step 4: Optimizer take a step and update the weights.
optimizer.step()
# Log the loss value as we proceed through the epochs.
if _e % 100 == 0:
losses.append(loss.data.item())
return losses
loss_normal = MLP(1000)
loss_droptout = MLP_Dropout(1000, dropout=0.5)
# Even at higher no. of parameter,
# training is stable with quite a high dropout.
plt.plot(loss_normal, label='normal')
plt.plot(loss_droptout, label='Dropout')
plt.legend(loc='upper right')
plt.show()
import sys
sys.executable
# ! pip3 install -U --user tensorboard tensorboardX
# # TensorboardX
#
# First, install `Tensorboard`:
#
# ```
# pip3 install -U --user tensorboard
# ```
#
#
# Using `TensorboardX` ( https://github.com/lanpa/tensorboardX ), a PyTorch wrapper for https://www.tensorflow.org/guide/summaries_and_tensorboard
#
#
# In command line:
#
# ```
# python3 -m tensorboard.main --logdir='./logs' --reload_interval --port 6006
# ```
#
#
# Then open a page on the browser to:
#
# ```
# localhost:6006
# ```
#
# Then run the code below
# +
from tensorboardX import SummaryWriter
gradients = []
hidden_dim = 200
num_epochs = 5000
#writer = SummaryWriter('./logs')
# Step 1: Initialization.
# Use Sequential to define a simple feed-forward network.
model = nn.Sequential(
# Use nn.Linear to get our simple perceptron.
nn.Linear(input_dim, hidden_dim),
# Use nn.Sigmoid to get our sigmoid non-linearity.
nn.Sigmoid(),
# Second layer neurons.
nn.Linear(hidden_dim, output_dim),
nn.Sigmoid()
)
# Initialize the optimizer
learning_rate = 0.3
optimizer = optim.SGD(model.parameters(), lr=learning_rate)
# Initialize the loss function.
criterion = nn.MSELoss()
#writer.add_graph(model, torch.zeros(*X.shape), True)
# Initialize the stopping criteria
# For simplicity, just stop training after certain no. of epochs.
losses = [] # Keeps track of the loses.
# Step 2-4 of training routine.
for _e in tqdm(range(num_epochs)):
# Reset the gradient after every epoch.
optimizer.zero_grad()
# Step 2: Foward Propagation
predictions = model(X)
#for name, param in model.named_parameters():
# writer.add_histogram(name, param.clone().cpu().data.numpy(), _e)
# Step 3: Back Propagation
# Calculate the cost between the predictions and the truth.
loss = criterion(predictions, Y)
# Remember to back propagate the loss you've computed above.
loss.backward()
# Step 4: Optimizer take a step and update the weights.
optimizer.step()
# Log the loss value as we proceed through the epochs.
if _e % 100 == 0:
losses.append(loss.data.item())
#writer.add_scalar('Train/Loss', loss.data.item(), _e)
#writer.close()
# -
# # [Optional] Now lets use TensorboardX on Word2Vec
#
# Go back to Session3 code, recode these components:
#
# - Get the data from `Language is never ever random`.
# - Recode the `Word2VecDataset`
# - Recode the `CBOW` and `Skipgram` models
# - Recode the training routine, `model(x); loss.backward(); optimizer.step()` etc.
#
# On top of that launch the TensorboardX and see explore.
| completed/Session 4 - Nuts and Bolts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="ju4Ilo4o5Np9" colab_type="text"
# # The Jupyter Notebook
# + [markdown] id="XGS-144K5Np-" colab_type="text"
# This is a jupyter notebook, one of the environments in which you can run Python code.
#
# It is comprised of "cells" that can be executed.
#
# The cells can be one of two types, markdown or code. Markdown cells are just for static text and equations. Code cells execute python code. You can change the type with the combo box above or using keyboard shortcuts
#
# Keyboard shortcuts are your friends. Learn the various shortcuts on your system, especially the "run cell" shortcut, usually ctrl-enter. alt-enter runs the cell and makes a new one after it!
#
# When having trouble, tinker, and ask google (often solutions are on stack overflow)
# + [markdown] id="GzfrskLm5Np_" colab_type="text"
# **Best to start simple, and play around!**
#
#
#
# + id="wdCuC3AZ5NqA" colab_type="code" colab={}
1
# + id="iIdqdxSR5NqD" colab_type="code" colab={}
3
# + id="-2k0GGni5NqF" colab_type="code" colab={}
'b'
# + id="lA7KbvCK5NqJ" colab_type="code" colab={}
type(1)
# + id="hW8dTlbc5NqM" colab_type="code" colab={}
type('b')
# + id="CcBP9uK95NqP" colab_type="code" colab={}
type(0.2323)
# + id="vdAJkBUg5NqS" colab_type="code" colab={}
int(0.232)
# + id="5cpED7_q5NqU" colab_type="code" colab={}
int(343.99)
# + id="IUd8DmUg5NqX" colab_type="code" colab={}
s='eggs'
# + id="lCip4skmO8je" colab_type="code" colab={}
s[0]
# + id="bksMXxAePIds" colab_type="code" colab={}
s[0:3]
# + id="l2pbCKQ9PxYk" colab_type="code" colab={}
print(s)
# + id="e5vX9wknP32g" colab_type="code" colab={}
print(type(s))
print(type(s[0]))
# + [markdown] id="MrsUljjM7QOO" colab_type="text"
# **Let's step back: Where did python's name come from?**
#
# a. A programmer who loved a snake?
#
# b. Monte Python
#
# c. Someone's favorite neck tatoo
#
#
#
# When he began implementing Python, <NAME> was also reading the published scripts from “Monty Python’s Flying Circus”, a BBC comedy series from the 1970s. Van Rossum thought he needed a name that was short, unique, and slightly mysterious, so he decided to call the language Python.
# + [markdown] id="6Po4h51k9B6X" colab_type="text"
# **Python was meant to be "fun":**
#
# [Zen of Python and the guiding principles](https://en.wikipedia.org/wiki/Zen_of_Python)
#
# Beautiful is better than ugly.
#
# Explicit is better than implicit.
#
# Simple is better than complex.
#
# Complex is better than complicated.
#
# Flat is better than nested.
#
# Sparse is better than dense.
#
# Readability counts.
#
# Special cases aren't special enough to break the rules.
#
# Although practicality beats purity.
#
# Errors should never pass silently.
#
# Unless explicitly silenced.
#
# In the face of ambiguity, refuse the temptation to guess.
#
# There should be one—and preferably only one—obvious way to do it.
#
# Although that way may not be obvious at first unless you're Dutch.
#
# Now is better than never.
#
# Although never is often better than right now.
#
# If the implementation is hard to explain, it's a bad idea.
#
# If the implementation is easy to explain, it may be a good idea.
#
# Namespaces are one honking great idea—let's do more of those!
# + [markdown] id="VNcdFQfC7eTX" colab_type="text"
# **Back to some coding**
#
# #**Math**
#
# **Operators**
#
# Operators for integers:
# `+ - * / // % **`
#
# Operators for floats:
# `+ - * / **`
#
# Boolean expressions:
# * keywords: `True` and `False` (note capitalization)
# * `==` equals: `5 == 5` yields `True`
# * `!=` does not equal: `5 != 5` yields `False`
# * `>` greater than: `5 > 4` yields `True`
# * `>=` greater than or equal: `5 >= 5` yields `True`
# * Similarly, we have `<` and `<=`.
#
# Logical operators:
# * `and`, `or`, and `not`
# * `True and False`
# * `True or False`
# * `not True`
# + id="okBL7w1x74rl" colab_type="code" colab={}
sin(pi/2)
# + id="T6MgCdOI7zOQ" colab_type="code" colab={}
import math
sin(pi/2)
# + id="-XTjEF848PtX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 166} executionInfo={"status": "error", "timestamp": 1593458309216, "user_tz": 240, "elapsed": 332, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="6717b650-3487-42e8-fa7e-3f4ee163d56c"
math.sin(math.pi/2)
# + id="LUfboTVz7p60" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1593458345870, "user_tz": 240, "elapsed": 430, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="2c7a37ee-7dab-4600-f365-e4afdf6a52ff"
import math
from math import sin, pi
math.sin(math.pi/2) # Note, in collaboratory, if you just try this code from the beginning it will work; math is automatically imported (here)
# + id="11kEEgZ5oW10" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1593458211119, "user_tz": 240, "elapsed": 764, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="6af4f598-6cf3-4735-d3ac-8e555d702fd7"
10/5
# + id="qfZKYOWYgsyO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1593458220438, "user_tz": 240, "elapsed": 512, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="73e03134-5ae2-419c-98dc-6eb5d938859c"
10%5
# + id="BsmaRF_HgvJz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 87} executionInfo={"status": "ok", "timestamp": 1593458271051, "user_tz": 240, "elapsed": 385, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="ba4040f3-b115-4e7f-8656-7e2e4af48b7f"
print(10//5)
print(10%5)
print(10//4)
print(10%4)
# + [markdown] colab_type="text" id="ogAIHRdvR9tA"
# #**Basic Variables Overview**
# + id="YebulT_u4_AW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} executionInfo={"status": "ok", "timestamp": 1592952863529, "user_tz": 240, "elapsed": 611, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="01b123ec-04d8-4541-b166-0426e341f5e8"
t=55
print(f'{t} is a {type(t)}') #formatted string literal or f string
# + id="4Z0bpk-zZG63" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1592952913224, "user_tz": 240, "elapsed": 608, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="54ee31ea-044f-41de-eb7b-6617fe3e48cb"
type(t)
# + id="W4JsRJLp5Rsm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1592952892653, "user_tz": 240, "elapsed": 543, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="66e9b64a-7a7a-4b0b-9bbe-bfa4fa0b2633"
strings="hi mom"
type(strings)
# + id="GMfl3VAF5Xjy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1592952896770, "user_tz": 240, "elapsed": 580, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="1043a8fe-46d2-4430-f75e-4d9397a47ccd"
t1=44.5
type(t1)
# + id="yhflg9-b5csk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1592952898667, "user_tz": 240, "elapsed": 631, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="bd2738e1-6f5a-4e04-ef1b-ff7ba6c1c5ba"
bl=False
type(bl)
# + [markdown] colab_type="text" id="FnUhFCVz47uA"
# #**Strings**
# + id="gzZhcMH7ACp9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} executionInfo={"status": "ok", "timestamp": 1593350915621, "user_tz": 240, "elapsed": 614, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="a86b400e-ad98-45dc-87bd-336a5b34a511"
str1="test"
str2='case'
print(str1)
print(str2)
# + id="Jusx5DyxAZbY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} executionInfo={"status": "ok", "timestamp": 1593350916766, "user_tz": 240, "elapsed": 506, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="2fc8c9e1-b3ad-4e1a-a61f-dc8ad37e8f2c"
#concatenation
str3=str1+str2+'!'
print(str1+str2)
print(str3)
# + id="E6quVujdAwag" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 69} executionInfo={"status": "ok", "timestamp": 1593350918889, "user_tz": 240, "elapsed": 517, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="ec59e4f2-e7f6-4081-cb84-daba0d421481"
x=20
y="40"
print("twice {x} is {y}")
print(f"twice {x} is {y}") # f string is an expression evaluated at runtime
print(f"twice {x} is {2*x}")
# + id="Stmu6PoqCp2g" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} executionInfo={"status": "ok", "timestamp": 1593539640740, "user_tz": 240, "elapsed": 628, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="79d5d3ba-7648-4815-d2ce-4558ed90a669"
str1 = "a: %s" % "string"
print(str1)
str2 = "b: %f, %s, %d" % (1.0, 'hello', 5)
print(str2)
str3 = "c: {}".format(3.14)
print(str3)
mylist = [1,2,3]
print("A list: %s" % mylist)
# + id="h3xb9RXqESro" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} executionInfo={"status": "ok", "timestamp": 1593545955062, "user_tz": 240, "elapsed": 1009, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="7acbc4d6-5b61-4129-e1d0-0434ccae751a"
str1 = "Hello, World!"
print(str1)
print(str1.upper())
print(str1.lower())
print(str1[0])
print(str1[len(str1)-1])
# + id="cYIknYohEva9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 163} executionInfo={"status": "error", "timestamp": 1593545956887, "user_tz": 240, "elapsed": 629, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="0e7b79a1-463e-42cc-ad00-09321f683228"
print(str1[len(str1)])
# + id="Pg5is8gdEy3c" colab_type="code" colab={}
str1.replace('l','p')
# + [markdown] colab_type="text" id="U4Kj8LATSoGs"
#
# #**Lists Tuple Set and Dict**
#
# List: collection of arbitrary objects, like array-- ordered, arbitrary, individually accessible, nestable, mutable, dynamic
#
#
# + id="ihxX8imdylNI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1592946962830, "user_tz": 240, "elapsed": 611, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="fa34a402-f159-44be-ba52-95e054487b0e"
# a List is an ordered set of objects ... foreshadow--numpy arrays
list1 = [4, 5, 'hi mom', 6.343]
print(list1)
# + id="snvXwaqCzIeR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} executionInfo={"status": "ok", "timestamp": 1592875969550, "user_tz": 240, "elapsed": 790, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="c62b2b16-575d-47ca-8029-8bd8add4b500"
list1[0]
print('length of list is', len(list1))
print('element 0 is', list1[0])
# + id="4d0gIpiCy3Ee" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} executionInfo={"status": "ok", "timestamp": 1592875823990, "user_tz": 240, "elapsed": 507, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="43d6377c-af25-496b-b92e-428af6ca00bf"
for x in list1:
print(x)
# + id="r9Dqp-QjCGsy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1592946965173, "user_tz": 240, "elapsed": 649, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="4fe3f0dd-9c8e-44b0-81c0-87f4892a6992"
list1.remove(4)
print(list1)
# + id="zEB0Xh5qCdiu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1593539884559, "user_tz": 240, "elapsed": 580, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="76f26f7c-5ea4-43c0-df45-9cb6662b4aad"
b=[] # b is an object of class list
for i in range(5):
b.append(i**2) # append is a 'method'
print(b)
# + id="tXpPzzY-ZvP8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} executionInfo={"status": "ok", "timestamp": 1592953201805, "user_tz": 240, "elapsed": 731, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="f3341958-e962-4e4b-db8d-1db9374cab5e"
# nesting in a list
x = ['a', ['bb', ['ccc', 'ddd'], 'ee', 'ff'], 'g', ['hh', 'ii'], 'j']
print(len(x))
print(x[0])
print(x[1])
print(x[1][0])
print(x[1][1])
print(x[1][1][0])
# + id="-dqoZCKVZM_y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1592953264973, "user_tz": 240, "elapsed": 311, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="391ead05-319f-48b3-c2f2-c21117ede458"
#tuple is ordered, indexed and immutable
tpl=(1,4,8)
tpl[0]
# + id="9gDHW8Znae-H" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 166} executionInfo={"status": "error", "timestamp": 1592953276609, "user_tz": 240, "elapsed": 537, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="8ac69cdf-d46f-4e64-8ae0-6c6b071507c8"
tpl[0]=5
# + id="2amZSwkrajKV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1592953341823, "user_tz": 240, "elapsed": 545, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="8156798f-e233-4d58-b3de-69f76456a748"
#set is unordered, unindexed, mutable, not allowed to have duplicate elements
st={0, 3, 5, 99}
st
# + id="3bNF1RBDaxOS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 166} executionInfo={"status": "error", "timestamp": 1592953348999, "user_tz": 240, "elapsed": 604, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="11a6aad5-fbb5-426b-8750-4eff5479ad4d"
st[0]
# + id="6HHPgsEea21Q" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} executionInfo={"status": "ok", "timestamp": 1592953461864, "user_tz": 240, "elapsed": 608, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="7180fb9d-97eb-4e2f-9a78-886369d8c140"
#dictionary (dict) unordered, indexed, mutable, no duplicate keys
dct={}
dct[5]=55
dct['somekey']=999
dct["anotherkey"]='hello'
print(dct)
print(dct['somekey'])
# + [markdown] id="jfFZ0uPIXpK6" colab_type="text"
# Create an array named ar1 that has the numbers 1-99 in it
# + id="viPSDQkFXoG7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 55} executionInfo={"status": "ok", "timestamp": 1593539926037, "user_tz": 240, "elapsed": 721, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="124e637d-d1f5-442d-cdba-e46e5abfe988"
ar1=[]
for i in range(99):
ar1.append(i+1)6
print(ar1)
# + [markdown] colab_type="text" id="TWbAd6axSGN8"
# #**Looping and flow control**
# + id="O1dzVDAkFDUt" colab_type="code" colab={}
# if, elif, else
xx=5
yy=True
if xx==5:
print("xx is 5 broh")
elif xx==yy:
print("xx is yy")
else:
print("nada")
# + id="bGtI-gQv17BC" colab_type="code" colab={}
xx=5
yy=True
if xx!=5:
print("xx is not 5 broh")
elif xx==yy:
print("xx is yy")
else:
print("nada")
# + id="E95BfrQu2P4y" colab_type="code" colab={}
# for loops
print(range(10))
for x in range(10):
print(x)
# + id="w6vZ_J_O2dmZ" colab_type="code" colab={}
x=66
while x<100:
print(x)
x+=0.1*x
# + id="JpizDBOU2wPX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 544} executionInfo={"status": "ok", "timestamp": 1593609883941, "user_tz": 240, "elapsed": 380, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="9def15be-1687-4edc-f26e-2126befb7702"
n=64
for num in range(2,n):
if num % 2 == 0:
continue # sends you to loop beginning again
print(f"{num} is an odd number")
# + id="PIJzPwBm3p0c" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} executionInfo={"status": "ok", "timestamp": 1593610034989, "user_tz": 240, "elapsed": 388, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="7ce5e552-9410-4a89-9b20-e395eeed86cd"
n=64
range(2,n)
for x in range(2,n):
if n%x == 0:
print(f'{n} equals {x} * {n // x}')
# + id="tGW8X0oY35Yh" colab_type="code" colab={}
if True==False:
pass
else:
print('True does not equal false')
# + [markdown] colab_type="text" id="swGW0kUOSGoQ"
# #**Functions**
# + id="fr0C9wVevyfm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1593610425258, "user_tz": 240, "elapsed": 391, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="f69094ab-b9d7-44e4-83bf-c531d36ef698"
# Use def to define a function
def rect_area(length, width):
return length*width
rect_area(5, 6)
# + id="aMJbuDdHwW1Y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 316} executionInfo={"status": "error", "timestamp": 1593610428242, "user_tz": 240, "elapsed": 561, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="be6b3fc0-1e7b-4d23-e04e-a5de579eeea8"
def rect_area(length, width):
if length<=0 or width<=0:
raise ValueError("length or width cannot be negative")
return length*width
print(rect_area(-1, 6))
# + id="RQMI51srxQX0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1592875567240, "user_tz": 240, "elapsed": 506, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="a8c10c9a-e8d9-4467-9af7-c8665a60c9cb"
# Any object can be passed to a fucntion
def f(x):
return x*x
def g(f, x):
return(f(f(x)))
g(f,5) # Calls f(x) on itself, so will return cube of x argument in g(f, x)
# + id="vUZKq8IWyLr1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1592875647207, "user_tz": 240, "elapsed": 548, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="df716141-264c-4247-8847-25e94bcd72f1"
def g(a, x, b=0):
return a * x + b
g(2,5,1)
# + id="OAS9A79MyYaR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1592875660739, "user_tz": 240, "elapsed": 298, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="76e39844-2b2b-49bd-fa3d-069c430546ae"
g(2,5)
# + [markdown] colab_type="text" id="1oByHKbJjsxE"
# #**Exceptions**
# + id="fG5ZkMauC-ai" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 166} executionInfo={"status": "error", "timestamp": 1592947115471, "user_tz": 240, "elapsed": 581, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="9ddf1569-1423-42d2-cd9d-c206aecf1e0a"
5/0
# + id="JDV5LjAvDBh5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1592947647083, "user_tz": 240, "elapsed": 614, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="f415832e-84b1-4a03-f4a5-03db5d6e06df"
try:
5/0
except ZeroDivisionError:
print("Don't divide by zero buddy")
# + [markdown] colab_type="text" id="MghTPHCfSw2H"
# #**Classes**
# + id="PXy01zcvbi40" colab_type="code" colab={}
class Materials:
def somefunc(self):
print("Hello!")
# + id="Iiv15nexcxkn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1592954040310, "user_tz": 240, "elapsed": 590, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="a5fbe731-b355-41e3-bddf-47e6b614995b"
ttt=Materials()
ttt.somefunc()
# + id="LJD23DoPet1e" colab_type="code" colab={}
# Example pulled from the web:
# Class that holds fractions r = p / q
class Rational:
def __init__(self, p, q=1):
if q == 0:
raise ValueError('Denominator must not be zero')
if not isinstance(p, int):
raise ValueError('Numerator must be an integer')
if not isinstance(q, int):
raise ValueError('Denominator must be an integer')
g = math.gcd(p, q)
self.p = p // g # integer division
self.q = q // g
# method to convert rational to float
def __float__(self):
return self.p / self.q
# method to convert rational to string for printing
def __str__(self):
return f'{self.p}/{self.q}'
def __repr__(self):
return f'Rational({self.p}, {self.q})'
# + id="PzjnaKK-e5Mm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} executionInfo={"status": "ok", "timestamp": 1593605856811, "user_tz": 240, "elapsed": 307, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="bd4bc36c-0d3f-4b56-ce29-ae240fcb8741"
import math
a=Rational(6,4)
print(f"a = {a}")
print(type(a))
print(f"float(a) = {float(a)}")
print(f"str(a) = {str(a)}")
print(f"repr(a) = {repr(a)}")
print(type(str(a)))
# + id="H-G7dSSAhoJy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} executionInfo={"status": "ok", "timestamp": 1593605866490, "user_tz": 240, "elapsed": 331, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiLyGI84VptrbFbYavfYil7QChJttVIfRd6CWPQeg=s64", "userId": "01470415962470640809"}} outputId="800d59ad-52d6-45fb-dc8e-058503d9c32d"
b=Rational(3,2)
print(str(b))
print(type(b))
# + [markdown] colab_type="text" id="HRNrWwmCS0Ll"
# #**Assignment 1**
#
# Please save your assignment in your personal google drive folder that is shared with the Profs as *Assignmnet1_[your last name]*, such as *Assignment1_Strandwitz*
#
# 1. Write a function that when called, prints every power of 2 that is less than the value passed to the function
# 2. Write a function that take inputs a, b, c and returns the difference of the sum of the squares of a and b and the square of c
# 3. create a list with elements 'this', 'is', 'a', 'list'. Then use insert() to add a period at the end of your list and then use remove() to take out any element of the list.
#
#
#
| lectures/1_Intro Part 1/MAT495_Intro to Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # PanLex Dataset Preparation
#
# This notebook is dedicated at making language pairs from the PanLex Dataset for word-level traduction
#
# The number of languages will be dictated by a pre-filtered list of languages obtained (kind of arbitrarilly) during the Universal Dependencies Conllu Dataset exploration
# A first dataset exploration of the different versions (CSV, SQL and JSON) shows that the sql one seems to lack a few things available in the other ones
# note that as the PanLex dataset defines the languages by 3 characters, the 2 characters codes had to be transformed to 3 and some of the 2 characters ones could not be found in 3 char definition, so there is not the same number of langs in 3char and 2char
# +
import os
import sys
import numpy as np
import scipy as sp
import pandas as pd
import matplotlib.pyplot as plt
import csv
import json
import sqlite3
import pickle
# %matplotlib inline
# -
PANLEX_BASEDIR= "/home/leo/projects/Datasets/text/PanLex"
PANLEX_CSV_DIR= os.path.join(PANLEX_BASEDIR,"panlex-20200601-csv")
PANLEX_SQLITE= os.path.join(PANLEX_BASEDIR,"panlex_lite/db.sqlite")
PANLEX_TRANSLATIONS = os.path.join(PANLEX_BASEDIR, "translations_tuples.pkl")
langs_2char = ['af', 'be', 'bg', 'ca', 'cs', 'cy', 'da', 'de', 'el', 'en', 'es', 'et', 'eu', 'fi', 'fr', 'fro', 'ga', 'gd', 'gl', 'grc', 'gsw', 'he', 'hr', 'hu', 'hy', 'it', 'la', 'lt', 'lv', 'mt', 'nl', 'no', 'pl', 'pt', 'ro', 'ru', 'sk', 'sl', 'sr', 'sv', 'tr', 'uk']
langs_3char = ('afr', 'bel', 'bul', 'cat', 'ces', 'cym', 'dan', 'deu', 'ell', 'eng', 'spa', 'est', 'eus', 'fin', 'fra', 'gle', 'gla', 'glg', 'heb', 'hrv', 'hun', 'hye', 'ita', 'lat', 'lit', 'lav', 'mlt', 'nld', 'nor', 'pol', 'por', 'ron', 'rus', 'slk', 'slv', 'srp', 'swe', 'tur', 'ukr')
len(langs_2char), len(langs_3char)
scripts = ('Latn', 'Cyrl', 'Grek', 'Hebr')
conn = sqlite3.connect(PANLEX_SQLITE)
c = conn.cursor()
q_langcodes = f"SELECT DISTINCT * from langvar where lang_code in {langs_3char} AND script_expr_txt in {scripts};"
q_langcodes
r_langcodes = list(c.execute(q_langcodes))
lang_ids = tuple([i[0] for i in r_langcodes])
q_expr_red = f"SELECT DISTINCT * from expr WHERE langvar in {lang_ids}"
q_expr_red
r_expr_red = list(c.execute(q_expr_red))
q_denot_red = f"SELECT meaning, expr, langvar FROM denotationx WHERE langvar IN {lang_ids};"
q_denot_red
r_denot_red = list(c.execute(q_denot_red))
# +
denotation = {}
for k,eid,lid in r_denot_red:
if k in denotation:
denotation[k].append((eid, lid))
else:
denotation[k] = [(eid, lid)]
# -
len(list(denotation.keys()))
expr = {k:(lid, txt) for (k,lid,txt) in r_expr_red}
len(r_expr_red), len(list(expr.keys()))
langvar = {i[0]:i for i in r_langcodes}
len(r_langcodes), len(list(langvar.keys()))
len(r_denot_red)
r_denot_red[:10], r_expr_red[:10], r_langcodes[:10]
# +
# denotation = pd.DataFrame(r_denot_red)
# del(denotation)
# +
# expressions = pd.DataFrame(r_expr_red)
# langvar = pd.DataFrame(r_langcodes)
# del(expressions)
# del(lanvar)
# +
# import gc
# gc.collect()
# +
# %%time
expr_tuples = []
expr_errs = []
for v in denotation.values():
transl = []
for item in v:
eid,lid = item
try:
lang = langvar[lid][1]
ex = expr[eid][1]
transl.append((lang, ex))
except:
expr_errs.append((eid, lid))
expr_tuples.append(transl)
# -
len(expr_errs)
len(expr_tuples)
expr_tuples[1256:1265]
expr_errs[:10]
expr[43387]
# +
# f = open(PANLEX_TRANSLATIONS, 'wb')
# pickle.dump(expr_tuples, f)
# -
f.close()
| utf8/notebooks/PanLex_preparation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
df = pd.read_excel('default_of_credit_card_clients.xls')
df.columns
df.head()
df['ID'].nunique()
df.shape
id_counts = df['ID'].value_counts()
id_counts.head()
id_counts.value_counts()
import numpy as np
np.random.seed(seed=24)
random_integers = np.random.randint(low=1,high=5,size=100)
is_equal_to_3 = random_integers == 3
is_equal_to_3[:5]
sum(is_equal_to_3)
random_integers[is_equal_to_3]
dupe_mask = id_counts == 2
dupe_mask[0:5]
id_counts.index[0:5]
dupe_ids = id_counts.index[dupe_mask]
dupe_ids = list(dupe_ids)
len(dupe_ids)
dupe_ids[0:5]
df.loc[df['ID'].isin(dupe_ids[0:3]),:].head(10)
df_zero_mask = df == 0
feature_zero_mask = df_zero_mask.iloc[:,1:].all(axis=1)
sum(feature_zero_mask)
df_clean_1 = df.loc[~feature_zero_mask,:].copy()
df_clean_1.shape
df_clean_1['ID'].nunique()
df_clean_1.info()
df_clean_1['PAY_1'].head()
df_clean_1['PAY_1'].value_counts()
valid_pay_1_mask = df_clean_1['PAY_1'] != 'Not available'
valid_pay_1_mask[0:5]
sum(valid_pay_1_mask)
df_clean_2 = df_clean_1.loc[valid_pay_1_mask,:].copy()
df_clean_2.shape
df_clean_2['PAY_1'].value_counts()
df_clean_2['PAY_1'] = df_clean_2['PAY_1'].astype('int64')
df_clean_2[['PAY_1', 'PAY_2']].info()
# +
import matplotlib.pyplot as plt #import plotting package
#render plotting automatically
# %matplotlib inline
import matplotlib as mpl #additional plotting functionality
mpl.rcParams['figure.dpi'] = 400 #high resolution figures
# -
df_clean_2[['LIMIT_BAL', 'AGE']].describe()
df_clean_2['EDUCATION'].value_counts()
df_clean_2['EDUCATION'].replace(to_replace=[0, 5, 6], value=4, inplace=True)
df_clean_2['EDUCATION'].value_counts()
df_clean_2['MARRIAGE'].value_counts()
df_clean_2['MARRIAGE'].replace(to_replace=0, value=3, inplace=True)
df_clean_2['MARRIAGE'].value_counts()
df_clean_2.groupby('EDUCATION').agg({'default payment next month':'mean'}).plot.bar(legend=False)
plt.ylabel('Default rate')
plt.xlabel('Education level: ordinal encoding')
df_clean_2['EDUCATION_CAT'] = 'none'
df_clean_2[['EDUCATION', 'EDUCATION_CAT']].head(10)
cat_mapping = {
1: "graduate school",
2: "university",
3: "high school",
4: "others"
}
df_clean_2['EDUCATION_CAT'] = df_clean_2['EDUCATION'].map(cat_mapping)
df_clean_2[['EDUCATION', 'EDUCATION_CAT']].head(10)
edu_ohe = pd.get_dummies(df_clean_2['EDUCATION_CAT'])
edu_ohe.head(10)
df_with_ohe = pd.concat([df_clean_2, edu_ohe], axis=1)
df_with_ohe[['EDUCATION_CAT', 'graduate school',
'high school', 'university', 'others']].head(10)
df_with_ohe.to_csv('cleaned_data.csv', index=False)
df = pd.read_csv('cleaned_data.csv')
pay_feats = ['PAY_1', 'PAY_2', 'PAY_3', 'PAY_4', 'PAY_5', 'PAY_6']
df[pay_feats].describe()
df[pay_feats[0]].value_counts().sort_index()
df[pay_feats[0]].hist()
pay_1_bins = np.array(range(-2,10)) - 0.5
pay_1_bins
df[pay_feats[0]].hist(bins=pay_1_bins)
plt.xlabel('PAY_1')
plt.ylabel('Number of accounts')
mpl.rcParams['font.size'] = 4
df[pay_feats].hist(bins=pay_1_bins, layout=(2,3))
df.loc[df['PAY_2']==2, ['PAY_2', 'PAY_3']].head()
| Mini-Project-2/Project 2/Credit_Card_Data_Exploration_and_Cleaning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <form action="https://github.com/prmiles/pymcmcstat_examples">
# <input type="submit" value="Return to Index" style="background-color: green; color: white; width: 150px; height: 35px; float: right"/>
# </form>
#
# # Estimating the Error Variance
#
# Author(s): <NAME> | Date Created: August 21, 2018
#
# Included in the [pymcmcstat](https://github.com/prmiles/pymcmcstat/wiki) package is the ability to estimate the error variance as part of the sampling process. Furthermore, when using multiple data sets to inform parameter values, you can estimate error variances for each data set separately.
#
# For more details regarding how to estimate the error variance please refer to:
# - <NAME>. (2013). Uncertainty quantification: theory, implementation, and applications (Vol. 12). SIAM.
# - <NAME>., & <NAME>. (2000). A simple method for generating gamma variables. ACM Transactions on Mathematical Software (TOMS), 26(3), 363-372. [https://doi.org/10.1145/358407.358414](https://doi.org/10.1145/358407.358414)
# import required packages
import numpy as np
from pymcmcstat.MCMC import MCMC
import matplotlib.pyplot as plt
# # Define Model and Sum-of-Squares Functions
# - Note, the sum-of-squares function is designed to loop through the data sets.
# define model function
def modelfun(xdata, theta):
m = theta[0]
b = theta[1]
nrow, ncol = xdata.shape
y = np.zeros([nrow, 1])
y[:,0] = m*xdata.reshape(nrow,) + b
return y
# define sum-of-squares function
def ssfun(theta, data):
n = len(data.xdata)
ss = np.zeros([n])
for ii in range(n):
xdata = data.xdata[ii]
ydata = data.ydata[ii]
# eval model
ymodel = modelfun(xdata, theta)
# calc sos
ss[ii] = sum((ymodel[:, 0] - ydata[:, 0])**2)
return ss
# # Define Data Set - Plot
# We consider two simulated data sets. Both are linear, but each one has a different level of observation errors. The first data set has $\varepsilon_i \sim N(0, 0.1)$, whereas the second data set has $\varepsilon_i \sim N(0, 0.5)$.
# +
# Add data
nds = 100
m = 2.
b = 3.
x1 = np.linspace(2, 3, num=nds).reshape(nds, 1)
y1 = m*x1 + b + 0.1*np.random.standard_normal(x1.shape)
res1 = y1 - modelfun(x1, [m, b])
x2 = np.linspace(1, 5, num=nds).reshape(nds, 1)
y2 = m*x2 + b + 0.5*np.random.standard_normal(x2.shape)
res2 = y2 - modelfun(x2, [m, b])
plt.figure(figsize=(8, 4))
plt.subplot(1, 2, 1)
plt.plot(x1, y1, '.b');
plt.plot(x1, modelfun(x1, [m, b]), '-r', linewidth=3);
plt.xlabel('$x_1$'); plt.ylabel('$y_1$');
plt.subplot(1, 2, 2)
plt.plot(x1, res1, '.g');
mr = res1.mean()
plt.plot([x1[0], x1[-1]], [mr, mr], '-k', linewidth=3)
plt.xlabel('$x_1$')
plt.ylabel(s=str('Residual, ($\\mu$ = {:5.4e})'.format(mr)));
plt.suptitle('Data Set 1')
plt.tight_layout(rect=[0, 0.03, 1, 0.95])
plt.figure(figsize=(8, 4))
plt.suptitle('Data Set 2')
plt.subplot(1, 2, 1)
plt.plot(x2, y2, '.b');
plt.plot(x2, modelfun(x2, [m, b]), '-r', linewidth = 3);
plt.xlabel('$x_2$')
plt.ylabel('$y_2$');
plt.subplot(1, 2, 2)
plt.plot(x2, res2, '.g');
mr = res2.mean()
plt.plot([x2[0], x2[-1]], [mr, mr], '-k', linewidth = 3)
plt.xlabel('$x_2$')
plt.ylabel(s=str('Residual, ($\\mu$ = {:5.4e})'.format(mr)));
plt.tight_layout(rect=[0, 0.03, 1, 0.95])
# -
# # Initialize MCMC Object and Setup Simulation
# - We call the `add_data_set` method twice to add each data set to the MCMC data structure.
# - The `updatesigma` flag must be turned on in order to include the observation errors in the sampling process.
mcstat = MCMC()
mcstat.data.add_data_set(x1, y1)
mcstat.data.add_data_set(x2, y2)
mcstat.simulation_options.define_simulation_options(
nsimu=int(5.0e4),
updatesigma=True,
method='dram')
mcstat.model_settings.define_model_settings(sos_function=ssfun)
mcstat.parameters.add_model_parameter(name='m', theta0=1.0)
mcstat.parameters.add_model_parameter(name='b', theta0=2.0)
# # Run Simulation
# run mcmc
mcstat.run_simulation()
# Extract results
results = mcstat.simulation_results.results
chain = results['chain']
s2chain = results['s2chain']
names = results['names']
# define burnin
burnin = int(results['nsimu']/2)
# display chain statistics
mcstat.chainstats(chain[burnin:, :], results)
# # Plot Parameter Chains, Posteriors, and Observation Standard Deviations
mcpl = mcstat.mcmcplot # initialize plotting methods
mcpl.plot_density_panel(chain[burnin:,:], names);
mcpl.plot_chain_panel(chain[burnin:,:], names);
mcpl.plot_density_panel(np.sqrt(s2chain[burnin:,:]), ['$\\sigma_1$', '$\\sigma_2$']);
print(np.mean(np.sqrt(s2chain[burnin:,:]), axis = 0))
# We observe that the estimated observation error standard deviations have mean values around 0.11 and 0.50. As the data was generated assuming 0.10 and 0.50, this result is in good agreement.
# # Plot Prediction/Credible Intervals for Multiple Data Sets
# generate prediction intervals
def pred_modelfun(preddata, theta):
return modelfun(preddata.xdata[0], theta)
mcstat.PI.setup_prediction_interval_calculation(
results=results,
data=mcstat.data,
modelfunction=pred_modelfun,
burnin=burnin)
mcstat.PI.generate_prediction_intervals(calc_pred_int=True)
# plot prediction intervals
data_display = dict(marker='o', color='b', mfc='w')
model_display = dict(color='r', linestyle='--')
figh, axh = mcstat.PI.plot_prediction_intervals(
adddata=True,
plot_pred_int=True,
model_display=model_display,
data_display=data_display)
for ii, axi in enumerate(axh):
axi.set_xlabel(str('$x_{}$'.format(ii + 1)), fontsize=22)
axi.set_ylabel(str('$y_{}$'.format(ii + 1)), fontsize=22)
axi.set_title(str('Data Set {}'.format(ii + 1)), fontsize=22)
axi.tick_params(labelsize=22)
# # Discussion
# We observe the expected behavior as about 95% of the data points are contained within the 95% prediction interval. The prediction intervals are generated by propagating uncertainty from the parameters and the observation error chain, so this result further supports that our algorithm successfully estimates the observation errors in the data.
| tutorials/estimating_error_variance/estimating_error_variance_for_mutliple_data_sets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ODEs (Lecture 7)
# + tags=["hide_input"]
# This cell just imports the relevant modules
import numpy as np
from math import pi, exp
from sympy import init_printing, sin, cos, Function, Symbol, diff, integrate, dsolve, checkodesol, solve, ode_order, classify_ode, pprint
import mpmath
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
from matplotlib import animation, rc
from IPython.display import HTML
# -
# ## Order of an ODE
#
# **Slide 9**
#
# Use `sympy` to define dependent and independent variables, constants, ODE, and to find the order of ODEs.
# +
t = Symbol('t') # Independent variable
eta = Symbol('eta') # Constant
v = Function('v')(t) # Dependent variable v(t)
ode = diff(v,t) + eta*v # The ODE we wish to solve. Make sure the RHS is equal to zero.
print("ODE #1:")
pprint(ode)
print("The order of ODE #1 is", ode_order(ode, v))
x = Function('x')(t) # Dependent variable x(t)
m = Symbol('m') # Constant
k = Symbol('k') # Constant
ode = m*diff(x,t,2) + k*x
print("ODE #2:")
pprint(ode)
print("The order of ODE #2 is", ode_order(ode, x))
y = Function('y')(t) # Dependent variable y(t)
ode = diff(y,t,4) - diff(y,t,2)
print("ODE #3:")
pprint(ode)
print("The order of ODE #3 is", ode_order(ode, y))
# -
# # Analytical solutions
#
# **Slide 14**
#
# Solving ODEs analytically using `sympy.dsolve`
# +
x = Symbol('x') # Independent variable
y = Function('y')(x) # Dependent variable y(x)
# The ODE we wish to solve. Make sure the RHS is equal to zero.
ode = diff(y,x) - 2*x*(1-y)
solution = dsolve(ode, y) # Solve the ode for function y(x).
print("ODE #4:")
pprint(ode)
print("The solution to ODE #4 is: ", solution)
# + tags=["hide_input"]
x_3d = np.arange(-5, 5, 0.01)
y_3d = np.arange(-5, 5, 0.01)
X, Y = np.meshgrid(x_3d, y_3d)
dydx = 2 * X * (1-Y)
x = np.linspace(-5, 5, 1000)
y = 1 + np.exp(-x**2)
fig = plt.figure(figsize=(16,8))
ax1 = fig.add_subplot(121, projection='3d')
ax1.plot_surface(X, Y, dydx, cmap='seismic', edgecolor='k', lw=0.25)
ax1.set_xlabel('x')
ax1.set_ylabel('y(x)')
ax1.set_zlabel('dy/dx')
ax1.set_title('dy/dx = 2x(1-y)', fontsize=14)
ax2 = fig.add_subplot(122)
ax2.plot(x, y, 'k', label='y(x) = 1 + exp(-x**2)')
ax2.set_xlabel('x')
ax2.set_ylabel('y(x)')
ax2.set_title("Solution to ODE 4", fontsize=14)
ax2.legend(loc='best')
ax2.grid(True)
fig.tight_layout()
plt.show()
# -
# ```{note}
#
# The function `checkodesol` checks that the result from `dsolve` is indeed a solution to the ode. It substitutes in 'solution' into 'ode' and checks that the RHS is zero. If it is, the function returns 'True'.
#
# ```
# +
print("Checking solution using checkodesol...")
check = checkodesol(ode, solution)
print("Output from checkodesol:", check)
if(check[0] == True):
print("y(x) is indeed a solution to ODE #4")
else:
print("y(x) is NOT a solution to ODE #4")
# -
# ```{note}
#
# The mpmath module can handle initial conditions (x0, y0) when solving an initial value problem, using the
# odefun function. However, this will not give you an analytical solution to the ODE, only a numerical solution.
#
# ```
# +
f = mpmath.odefun(lambda x, y: 2*x*(1-y), x0=0, y0=2)
# compares the numerical solution f(x) with the values of the (already known) analytical solution
# between x=0 and x=10
for x in np.linspace(0, 10, 101):
print("x=%.1f" % (x), ",", f(x), ",", 1+exp(-x**2))
# -
# # Separation of variables
#
# **Slide 20**
#
# We can solve ODEs via separation of variables in Python using `sympy.dsolve` by passing the `hint` argument. Note that the optional `hint` argument here has been used to tell SymPy how to solve the ODE. However, it is usually smart enough to work it out for itself.
# +
x = Symbol('x') # Independent variable
y = Function('y')(x) # Dependent variable y(x)
# The ODE we wish to solve.
ode = (1.0/y)*diff(y,x) - cos(x)
print("ODE #5:")
pprint(ode)
# Solve the ode for function y(x).using separation of variables.
solution = dsolve(ode, y, hint='separable')
print("The solution to ODE #5 is:")
pprint(solution)
# + tags=["hide_input"]
x_3d = np.arange(-5, 5, 0.01)
y_3d = np.arange(-5, 5, 0.01)
X, Y = np.meshgrid(x_3d, y_3d)
dydx = Y * np.cos(X)
x = np.linspace(-5, 5, 1000)
y = np.exp(np.sin(x))
fig = plt.figure(figsize=(16, 8))
ax1 = fig.add_subplot(121, projection='3d')
ax1.plot_surface(X, Y, dydx, cmap='seismic', edgecolor='k', lw=0.25)
ax1.set_xlabel('x')
ax1.set_ylabel('y(x)')
ax1.set_zlabel('dy/dx')
ax1.set_title('dy/dx = ycos(x)', fontsize=14)
ax2 = fig.add_subplot(122)
ax2.plot(x, y, 'k', label='y(x) = exp(sin(x))')
ax2.set_xlabel('x')
ax2.set_ylabel('y(x)')
ax2.set_title("Solution of ODE 5", fontsize=14)
ax2.legend(loc='best')
ax2.grid(True)
fig.tight_layout()
plt.show()
# -
# # Integration factor
#
# **Slide 23**
# +
x = Symbol('x') # Independent variable
y = Function('y')(x) # Dependent variable y(x)
# The ODE we wish to solve.
ode = diff(y,x) - 2*x + 2*x*y
print("ODE #6:")
pprint(ode)
# Solve the ode for function y(x).using separation of variables
solution = dsolve(ode, y)
print("The solution to ODE #6 is:", solution)
# -
# ## Application
#
# ### Radioactive decay
#
# **Slide 26**
# +
t = Symbol('t') # Independent variable
N = Function('N')(t) # Dependent variable N(t)
l = Symbol('l') # Constant
# The ODE we wish to solve:
ode = diff(N,t) + l*N
print("ODE #7:")
pprint(ode)
solution = dsolve(ode, N)
print("The solution to ODE #7 is:")
pprint(solution)
# -
# **Example:** 1 mole of carbon-14 at t=0
# + tags=["hide_input"]
l = 3.8394e-12
C = 6.02e23 * np.exp(l)
t_3d = np.arange(0, 1e12, 1e9)
n_3d = np.arange(0, 6.02e23, 6.02e20)
N, T = np.meshgrid(n_3d, t_3d)
dNdt = -l * N
t = np.linspace(0, 1e12, 1000)
n = C * np.exp(-l*t)
t_years = t/(3600*24*365.25)
fig = plt.figure(figsize=(16, 8))
ax1 = fig.add_subplot(121, projection='3d')
ax1.plot_surface(T, N, dNdt, cmap='seismic', edgecolor='k', lw=0.25)
ax1.set_xlabel('t')
ax1.set_ylabel('N(t)')
ax1.set_zlabel('dN/dt')
ax1.set_title('Rate of change in number of C-14 nuclei', fontsize=14)
ax2 = fig.add_subplot(122)
ax2.plot(t_years, n, 'k', label='N(t) = Cexp(-lt)')
ax2.set_xlabel('Time (years)')
ax2.set_ylabel('Number of C-14 nuclei')
ax2.set_title("Number of C-14 nuclei over time", fontsize=14)
ax2.legend(loc='best', fontsize=12)
ax2.grid(True)
fig.tight_layout()
plt.show()
# -
# ```{note}
#
# The plane in the first graph shows that radioactive decay is independent of time, but only dependent on the number of radioactive nuclei present.
#
# ```
# ### Particle settling
#
# **Slide 31**
# +
t = Symbol('t') # Independent variable - time
v = Function('v')(t) # Dependent variable v(t) - the particle velocity
# Physical constants
rho_f = Symbol('rho_f') # Fluid density
rho_p = Symbol('rho_p') # Particle density
eta = Symbol('eta') # Viscosity
g = Symbol('g') # Gravitational acceleration
a = Symbol('a') # Particle radius
# The ODE we wish to solve.
ode = diff(v,t) - ((rho_p - rho_f)/rho_p)*g + (9*eta/(2*(a**2)*rho_p))*v
print("ODE #8:")
pprint(ode)
solution = dsolve(ode, v)
print("The solution to ODE #8 is:")
pprint(solution)
# -
# **Example**: sand grain with density 2650kg/m3 and radius 1mm sinking in water.
#
# Initial conditions: v=0 when t=0
# + tags=["hide_input"]
rho_f = 1000
rho_p = 2650
eta = 0.89
g = 9.81
a = 1e-3
C = -(2*a**2*rho_p)/(9*eta) * np.log((rho_p-rho_f)/rho_p)
v_ode = np.linspace(0, 10, 1000)
dvdt = (rho_p-rho_f)/rho_p - (9*eta*v_ode)/(2*a**2*rho_p)
t = np.arange(0, 0.005, 0.00005)
v = -2*a**2*g*rho_f + 2*a**2*g*rho_p + np.exp(eta*(C - 9*t/(2*a**2*rho_p)))/(9*eta)
fig = plt.figure(figsize=(16, 8))
ax1 = fig.add_subplot(121)
ax1.plot(v_ode, dvdt, 'r')
ax1.set_xlabel('v')
ax1.set_ylabel('dv/dt')
ax1.set_title("Plot of dv/dt vs v")
ax1.grid(True)
ax2 = fig.add_subplot(122)
ax2.plot(t, v, 'b')
ax2.set_xlabel('time (seconds)')
ax2.set_ylabel('velocity (m/s)')
ax2.set_title("Plot of velocity over time")
ax2.grid(True)
fig.tight_layout()
plt.show()
# -
| notebooks/a_modules/math_methods_1/8_ODEs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:us-east-1:081325390199:image/datascience-1.0
# ---
# <h1>Model Deployment</h1>
# Once we have built and trained our models for feature engineering (using Amazon SageMaker Processing and SKLearn) and binary classification (using the XGBoost open-source container for Amazon SageMaker), we can choose to deploy them in a pipeline on Amazon SageMaker Hosting, by creating an Inference Pipeline.
# https://docs.aws.amazon.com/sagemaker/latest/dg/inference-pipelines.html
#
# This notebook demonstrates how to create a pipeline with the SKLearn model for feature engineering and the XGBoost model for binary classification.
#
# Let's define the variables first.
# +
import sagemaker
import sys
import IPython
# Let's make sure we have the required version of the SM PySDK.
required_version = '2.46.0'
def versiontuple(v):
return tuple(map(int, (v.split("."))))
if versiontuple(sagemaker.__version__) < versiontuple(required_version):
# !{sys.executable} -m pip install -U sagemaker=={required_version}
IPython.Application.instance().kernel.do_shutdown(True)
# -
import sagemaker
print(sagemaker.__version__)
# +
import boto3
role = sagemaker.get_execution_role()
region = boto3.Session().region_name
sagemaker_session = sagemaker.Session()
bucket_name = sagemaker_session.default_bucket()
prefix = 'endtoendmlsm'
print(region)
print(role)
print(bucket_name)
# -
# ## Retrieve model artifacts
# First, we need to create two Amazon SageMaker **Model** objects, which associate the artifacts of training (serialized model artifacts in Amazon S3) to the Docker container used for inference. In order to do that, we need to get the paths to our serialized models in Amazon S3.
# <ul>
# <li>For the SKLearn model, in Step 02 (data exploration and feature engineering) we defined the path where the artifacts are saved</li>
# <li>For the XGBoost model, we need to find the path based on Amazon SageMaker's naming convention. We are going to use a utility function to get the model artifacts of the last training job matching a specific base job name.</li>
# </ul>
# +
from notebook_utilities import get_latest_training_job_name, get_training_job_s3_model_artifacts
# SKLearn model artifacts path.
sklearn_model_path = 's3://{0}/{1}/output/sklearn/model.tar.gz'.format(bucket_name, prefix)
# XGBoost model artifacts path.
training_base_job_name = 'end-to-end-ml-sm-xgb'
latest_training_job_name = get_latest_training_job_name(training_base_job_name)
xgboost_model_path = get_training_job_s3_model_artifacts(latest_training_job_name)
print('SKLearn model path: ' + sklearn_model_path)
print('XGBoost model path: ' + xgboost_model_path)
# -
# ## SKLearn Featurizer Model
# Let's build the SKLearn model. For hosting this model we also provide a custom inference script, that is used to process the inputs and outputs and execute the transform.
#
# The inference script is implemented in the `sklearn_source_dir/inference.py` file. The custom script defines:
#
# - a custom `input_fn` for pre-processing inference requests. Our input function accepts only CSV input, loads the input in a Pandas dataframe and assigns feature column names to the dataframe
# - a custom `predict_fn` for running the transform over the inputs
# - a custom `output_fn` for returning either JSON or CSV
# - a custom `model_fn` for deserializing the model
# !pygmentize data_transformation_dir/inference.py
# Now, let's create a `Model` object, by providing the custom script as input.
# +
import time
from sagemaker.model import Model
code_location = 's3://{0}/{1}/code'.format(bucket_name, prefix)
sklearn_model = SKLearnModel(name='end-to-end-ml-sm-skl-model-{0}'.format(str(int(time.time()))),
model_data=sklearn_model_path,
entry_point='inference.py',
source_dir='sklearn_source_dir/',
code_location=code_location,
role=role,
sagemaker_session=sagemaker_session,
framework_version='0.20.0',
py_version='py3')
# -
# ## XGBoost Model
# Similarly to the previous steps, we can create an `XGBoost` model object. Also here, we have to provide a custom inference script.
#
# The inference script is implemented in the `xgboost_source_dir/inference.py` file. The custom script defines:
#
# - a custom `input_fn` for pre-processing inference requests. This input function is able to handle JSON requests, plus all content types supported by the default XGBoost container. For additional information please visit: https://github.com/aws/sagemaker-xgboost-container/blob/master/src/sagemaker_xgboost_container/encoder.py. The reason for adding the JSON content type is that the container-to-container default request content type in an inference pipeline is JSON.
# - a custom `model_fn` for deserializing the model
# !pygmentize xgboost_source_dir/inference.py
# Now, let's create the `XGBoostModel` object, by providing the custom script and S3 model artifacts as input.
# +
import time
from sagemaker.xgboost import XGBoostModel
code_location = 's3://{0}/{1}/code'.format(bucket_name, prefix)
xgboost_model = XGBoostModel(name='end-to-end-ml-sm-xgb-model-{0}'.format(str(int(time.time()))),
model_data=xgboost_model_path,
entry_point='inference.py',
source_dir='xgboost_source_dir/',
code_location=code_location,
framework_version='0.90-2',
py_version='py3',
role=role,
sagemaker_session=sagemaker_session)
# -
# ## Pipeline Model
# Once we have models ready, we can deploy them in a pipeline, by building a `PipelineModel` object and calling the `deploy()` method.
# +
import sagemaker
import time
from sagemaker.pipeline import PipelineModel
pipeline_model_name = 'end-to-end-ml-sm-xgb-skl-pipeline-{0}'.format(str(int(time.time())))
pipeline_model = PipelineModel(
name=pipeline_model_name,
role=role,
models=[
sklearn_model,
xgboost_model],
sagemaker_session=sagemaker_session)
endpoint_name = 'end-to-end-ml-sm-pipeline-endpoint-{0}'.format(str(int(time.time())))
print(endpoint_name)
pipeline_model.deploy(initial_instance_count=1,
instance_type='ml.m5.xlarge',
endpoint_name=endpoint_name)
# -
# <span style="color: red; font-weight:bold">Please take note of the endpoint name, since it will be used in the next workshop module.</span>
# ## Getting inferences
# Finally we can try invoking our pipeline of models and get some inferences:
# +
from sagemaker.serializers import CSVSerializer
from sagemaker.deserializers import JSONDeserializer
from sagemaker.predictor import Predictor
predictor = Predictor(
endpoint_name=endpoint_name,
sagemaker_session=sagemaker_session,
serializer=CSVSerializer(),
deserializer=JSONDeserializer())
payload = "TID008,HAWT,64,80,46,21,55,55,7,34,SE"
print(predictor.predict(payload))
payload = "TID008,HAWT,64,2,46,21,55,55,7,2,SE"
print(predictor.predict(payload))
payload = "TID008,HAWT,1,2,46,21,55,55,7,2,SE"
print(predictor.predict(payload))
# -
predictor.delete_endpoint()
# Once we have tested the endpoint, we can move to the next workshop module. Please access the module <a href="https://github.com/giuseppeporcelli/end-to-end-ml-sm/tree/master/05_API_Gateway_and_Lambda" target="_blank">05_API_Gateway_and_Lambda</a> on GitHub to continue.
| 04_deploy_model/04_deploy_model_dw_fs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [anaconda]
# language: python
# name: Python [anaconda]
# ---
from bs4 import BeautifulSoup
import os
import spacy
nlp = spacy.load('en_core_web_sm')
import logging
logging.basicConfig(format='%(asctime)s : %(levelname)s : %(message)s', level=logging.INFO)
def get_sentences(PATH):
'''For each paras.txt extract sentences for each paragraph using bs4.'''
sent_file = open(os.path.join(PATH,'sentences.txt'),'w')
with open(os.path.join(PATH,'paras.txt')) as fobj:
#read the file line-by-line
for line in fobj:
if line != '\n' and line.strip().startswith('<p>'):
try:
#extract tagless paragraph-text from <p> tags.
soup = BeautifulSoup(line.strip(),"lxml")
#divide this tagless paragraphs into proper sentences using NLP via spacy.
doc = nlp(soup.p.text)
except:
#If the line can't be parsed then log the line and continue to next line.
logging.warning(PATH,":",line,"can't be parsed.")
continue
#If parsed then write each sentence in the file 'sentences.txt'.
for each in doc.sents:
text = each.text+'\n'
sent_file.write(each.text+'\n')
sent_file.close()
dirs = os.listdir('/Volumes/Seagate Expansion Drive/stackexchange')
#print len(dirs)
for each in dirs:
if each == '.DS_Store':
continue
PATH = os.path.join('/Volumes/Seagate Expansion Drive/stackexchange',each)
logging.info('Generating sentences for ' + each)
get_sentences(PATH)
logging.info('SUCCESS')
| Model/Sentence_Extraction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### This notebook shows some lower-level functionality in `flopy` for working with shapefiles
# including:
# * `recarray2shp` convience function for writing a numpy record array to a shapefile
# * `shp2recarray` convience function for quickly reading a shapefile into a numpy recarray
# * `utils.geometry` classes for writing shapefiles of model input/output. For example, quickly writing a shapefile of model cells with errors identified by the checker
# * demonstration of how the `epsgRef` class works for retrieving projection file information (WKT text) from spatialreference.org, and caching that information locally for when an internet connection isn't available
# * how to reset `epsgRef` if it becomes corrupted
# * examples of how the `Point` and `LineString` classes can be used to quickly plot pathlines and endpoints from MODPATH (these are also used by the `PathlineFile` and `EndpointFile` classes to write shapefiles of this output)
# +
import os
import sys
import shutil
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
# run installed version of flopy or add local path
try:
import flopy
except:
fpth = os.path.abspath(os.path.join('..', '..'))
sys.path.append(fpth)
import flopy
from flopy.utils.geometry import Polygon, LineString, Point
from flopy.export.shapefile_utils import recarray2shp, shp2recarray
from flopy.utils.modpathfile import PathlineFile, EndpointFile
from flopy.utils import geometry
from flopy.utils.reference import epsgRef
ep = epsgRef()
ep.reset()
print(sys.version)
print('numpy version: {}'.format(np.__version__))
print('matplotlib version: {}'.format(mpl.__version__))
print('flopy version: {}'.format(flopy.__version__))
# -
# ### write a numpy record array to a shapefile
# in this case, we want to visualize output from the checker
# first make a toy model
m = flopy.modflow.Modflow('toy_model', model_ws='data')
botm = np.zeros((2, 10, 10))
botm[0, :, :] = 1.5
botm[1, 5, 5] = 4 # negative layer thickness!
botm[1, 6, 6] = 4
dis = flopy.modflow.ModflowDis(nrow=10, ncol=10,
nlay=2, delr=100, delc=100,
top=3, botm=botm, model=m)
# + active=""
# ### set coordinate information
# -
grid = m.modelgrid
grid.set_coord_info(xoff=600000, yoff=5170000, proj4='EPSG:26715', angrot=45)
chk = dis.check()
chk.summary_array
# ### make geometry objects for the cells with errors
# * geometry objects allow the shapefile writer to be simpler and agnostic about the kind of geometry
get_vertices = m.modelgrid.get_cell_vertices # function to get the referenced vertices for a model cell
geoms = [Polygon(get_vertices(i, j)) for i, j in chk.summary_array[['i', 'j']]]
geoms[0].type
geoms[0].exterior
geoms[0].bounds
geoms[0].plot() # this feature requires descartes
# ### write the shapefile
# * the projection (.prj) file can be written using an epsg code
# * or copied from an existing .prj file
recarray2shp(chk.summary_array, geoms, 'data/test.shp', epsg=26715)
shutil.copy('data/test.prj', 'data/26715.prj')
recarray2shp(chk.summary_array, geoms, 'data/test.shp', prj='data/26715.prj')
# ### read it back in
# * flopy geometry objects representing the shapes are stored in the 'geometry' field
ra = shp2recarray('data/test.shp')
ra
ra.geometry[0].plot()
# ### How the epsg feature works
# * requires an internet connection the first time to get the prj text from [spatialreference.org](https://spatialreference.org/) using ```requests```
# * if it doesn't exist, ```epsgref.json``` is created in the user's data directory
# * the prj text is then stashed in this JSON file hashed by the EPSG numeric code
from flopy.utils.reference import epsgRef
ep = epsgRef()
prj = ep.to_dict()
prj
from flopy.utils.reference import getprj, epsgRef
getprj(4326)
prj = ep.to_dict()
for k, v in prj.items():
print('{}:\n{}\n'.format(k, v))
# ### working with the ```flopy.utils.reference.epsgRef``` handler
ep = epsgRef()
ep.add(9999, 'junk')
epsgRef.show()
# #### remove an entry
ep.remove(9999)
epsgRef.show()
# #### start over with a new file
ep.reset()
prj = ep.to_dict()
prj
len(prj.keys())
# ## Other geometry types
#
# ### Linestring
# * create geometry objects for pathlines from a MODPATH simulation
# * plot the paths using the built in plotting method
pthfile = PathlineFile('../data/mp6/EXAMPLE-3.pathline')
pthdata = pthfile._data.view(np.recarray)
# +
length_mult = 1. # multiplier to convert coordinates from model to real world
rot = 0 # grid rotation
particles = np.unique(pthdata.particleid)
geoms = []
for pid in particles:
ra = pthdata[pthdata.particleid == pid]
x, y = geometry.rotate(ra.x * length_mult,
ra.y * length_mult,
grid.xoffset,
grid.yoffset,
rot)
z = ra.z
geoms.append(LineString(list(zip(x, y, z))))
# -
geoms[0]
geoms[0].plot()
fig, ax = plt.subplots()
for g in geoms:
g.plot(ax=ax)
ax.autoscale()
ax.set_aspect(1)
# ## Points
eptfile = EndpointFile('../data/mp6/EXAMPLE-3.endpoint')
eptdata = eptfile.get_alldata()
# +
x, y = geometry.rotate(eptdata['x0'] * length_mult,
eptdata['y0'] * length_mult,
grid.xoffset,
grid.yoffset,
rot)
z = eptdata['z0']
geoms = [Point(x[i], y[i], z[i]) for i in range(len(eptdata))]
# -
fig, ax = plt.subplots()
for g in geoms:
g.plot(ax=ax)
ax.autoscale()
ax.set_aspect(2e-6)
| examples/Notebooks/flopy3_shapefile_features.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:mllanguage] *
# language: python
# name: conda-env-mllanguage-py
# ---
# # Sequence Processing with Recurrent Neural Networks
#
# So far, we have seen how word vectors can be constructed from corpus statistics, and how they can be utilized to infer latent semantic content either in isolation (e.g. genders from names) or in relation to one another (e.g. similarities and analogies).
#
# For tasks involving larger linguistic units such as phrases, sentences and dialogues, we need machinery capable of processing _sequences_ or _structures_ of words.
#
# Recurrent Neural Networks are an example of such machinery; for this assignment, you will construct a recurrent neural network that annotates each word of a sentence with a linguistically informative marker. In the simple case (and in this assignment), these markers will be POS tags. However, they can also be morphosyntactic informative [categories](https://en.wikipedia.org/wiki/Combinatory_categorial_grammar) (supertags).
#
# In both cases, the task is a case of sequence labeling. A good reference point is Jurafsky and Martin [Chapter 9](https://web.stanford.edu/~jurafsky/slp3/9.pdf). For a fuller view of the picture, a good reference point is <NAME>' [dissertation](https://www.cs.toronto.edu/~graves/preprint.pdf).
#
# We will take a gradual approach, first inspecting recurrent neural networks, then moving on to data processing using high-grade word vectors before finally moving to the problem at hand.
# ---
# ## Recurrent Neural Networks
# Recurrent Neural Networks are a particularly interesting class of neural networks. Unlike standard fully-connected networks, which accept a fixed-size input and produce a fixed-size output over a predefined number of computational steps (i.e. network layers), RNNs instead operate on sequences of vectors.
#
# Computationally, feedforward networks may be seen as a trainable (but parametrically fixed) function, whereas RNNs act as continuous, stateful programs operating on sequences of inputs.
# Cognitively, this may be viewed as enhancing our system's perceptive and computational abilities with a notion of memory.
# In the general case, this statefulness is captured by an intermediate hidden vector which is adjusted throughout the computation, affected by both the immediately previous version of itself __and__ the current input.
#
# RNNs are nowadays established as the core machinery of neural sequence processing.
#
# A simple recurrent network (SRN or Elman network) is described by the equations:
# * $h_t = \theta_h (W_h x_t + U_h h_{t-1} + b_h ) $
# * $y_t = \theta_y (W_y h_t + b_y) $
#
# where (at timestep $t$) $x_t$, $h_t$, $y_t$ are the network's input, hidden and output representations respectively, $\theta_h$, $\theta_y$ its hidden and output activation functions, and $W_h$, $U_h$, $b_h$, $W_y$, $b_y$ parametric tensors to be learned.
import torch
from torch import FloatTensor, LongTensor
from typing import Tuple, List, Callable, Optional
# ### Assignment 2.0: Our own SRN
# Let's make our own simple recurrent network from scratch, to get an idea of its inner workings. To make our life just a bit simpler, we will use `torch.nn.Linear` to model the internal transformations.
#
# Complete the `mySRN` class, which is initialized with the input $d_i$, hidden $d_h$ and output $d_o$ dimensionalities, as well as two non-linear functions $\theta_h$ and $\theta_y$, and constructs a SRN implementing three `torch.nn.Linear` layers:
# 1. `x_to_h`: a layer that takes $x_t$ and produces $W_h x_t$
# 2. `h_to_h`: a layer that takes $h_{t-1}$ and produces $U_h h_{t-1} + b_h$
# 3. `h_to_y`: a layer that takes $h_t$ and produces $W_y h_t + b_y$
#
# Implement the function `step` that performs a computational step, accepting $x_t$ and $h_{t-1}$ and producing $h_t$ and $y_t$.
#
# Implement the function forward that accepts a List of inputs $X$, an initial hidden vector $h_{-1}$ and iteratively applies `step` until the input sequence is exhausted, returning a List of outputs $Y$ (of the same length as $X$).
#
# _Hint_: Note that `x_to_h` does not have a bias term $b$, since we will incorporate it into `h_to_h`
class mySRN(torch.nn.Module):
def __init__(self, input_dim: int, hidden_dim: int, output_dim: int,
hidden_activation: Callable[[FloatTensor], FloatTensor],
output_activation: Callable[[FloatTensor], FloatTensor],
device: str):
super(mySRN, self).__init__()
self.device = device
self.input_dim = input_dim
self.hidden_dim = hidden_dim
self.output_dim = output_dim
self.hidden_activation = hidden_activation
self.output_activation = output_activation
self.x_to_h = torch.nn.Linear(in_features=self.input_dim, out_features=self.hidden_dim, bias=False)
self.h_to_h = torch.nn.Linear(in_features=self.hidden_dim, out_features=self.hidden_dim)
self.h_to_y = torch.nn.Linear(in_features=self.hidden_dim, out_features=self.output_dim)
def step(self, x: FloatTensor, h: FloatTensor) -> Tuple[FloatTensor, FloatTensor]:
h_t = self.hidden_activation(self.x_to_h(self.input_dim, self.hidden_dim))
y_t = self.output_activation(self.h_to_y(self.input_dim, self.output_dim))
return(h_t, y_t)
def forward(self, X: List[FloatTensor], h: FloatTensor) -> List[FloatTensor]:
Y = []
for x in X:
Y.append(step(x, h))
return(Y)
# In practice, we do not need to write our own functions for common RNN architectures.
# Torch already provides the [necessary abstractions](https://pytorch.org/docs/stable/nn.html#recurrent-layers).
#
# The [RNN](https://pytorch.org/docs/stable/nn.html#rnn) wrapper implements highly optimized forward routines to compute the hidden representations of a full input sequence.
#
# Some pointers:
# * Unlike our naive implementation, RNN accepts a 3-dimensional tensor of shape (seq_len, batch_shape, input_dim) rather than a list of 2-dimensional tensors
# * If no initial hidden state is provided, it defaults to a zero tensor
# * The class produces just the RNN hidden states; it is up to us to define the `h_to_y` transformation on top of them
# * The non-linearity argument is a string; our only two choices are either `'tanh'` or `'relu'` (shorthands for `torch.nn.Tanh` and `torch.nn.ReLU` respectively)
#
# Read the documentation (!) for further details.
#
# A brief example is given below.
rnn = torch.nn.RNN(input_size=16, hidden_size=48, nonlinearity='tanh')
X = torch.rand(10, 32, 16)
h, _ = rnn(X)
print(h.shape)
# So, for a random input tensor of shape (seq_len, batch_size, input_dim), we get back an output tensor of shape (seq_len, batch_size, hidden_dim)
del mySRN, rnn, X, h
# ### Assignment 2.1: A faster version of the SRN
# Now let's wrap an `RNN` into a custom module `myFastSRN` that implements it aside the `h_to_y` transformation.
class fastSRN(torch.nn.Module):
def __init__(self, input_dim: int, hidden_dim: int, output_dim: int,
hidden_activation: str,
output_activation: Callable[[FloatTensor], FloatTensor],
device: str):
super(fastSRN, self).__init__()
self.device = device
self.output_activation = output_activation
self.rnn = torch.nn.RNN(input_size=input_dim,
hidden_size=hidden_dim,
nonlinearity=hidden_activation)
self.h_to_y = torch.nn.Linear(in_features=hidden_dim,
out_features=output_dim)
def forward(self, X:FloatTensor, h: Optional[FloatTensor]=None) -> FloatTensor:
h, _ = self.rnn(X)
out = self.h_to_y(h)
out = self.output_activation(out)
return(out)
# Let's see our new implementation in action.
#
# Initialize a random input tensor $X$ that would correspond to 32 sequences, each of length 10, with each item having 16 features, and a `fastSRN` fit to process it, producing 42-dimensional hidden states and 2-dimension output vectors for each sequence item.
#
# Run the SRN on the tensor and make sure the output shape is as expected.
myFastSRN = fastSRN(input_dim=16,
hidden_dim=48,
hidden_activation='tanh',
output_activation=torch.nn.Softmax(dim=-1),
output_dim=2,
device='cpu')
X = torch.rand(10, 32, 16)
h = myFastSRN(X) # why does h, _ = myFastSRN(X) not work?
# Hopefully everything should be in order.
#
# You may have noticed a minor complication: in order to utilize batching, we need our input sequences to be of the same **length**.
#
# This however is very rarely the case in practice. A common trick against this problem is _padding_; that is, appending zero tensors to all input sequences shorter than the maximum in-batch length to make them all equally long.
#
# As usual, torch already does the hard work for us via [pad_sequence](https://pytorch.org/docs/stable/nn.html?highlight=pad%20_sequence#torch.nn.utils.rnn.pad_sequence). Given a list of $N$ 2-dimensional tensors, each of shape (seq_len$_n$, input_dim), it will construct a 3-d tensor of shape ($max_{n \in N}${seq_len$_n$}, N, input_dim).
#
# An example:
# +
x_1 = torch.rand(1, 16) # a sequence of 1, 16-dimensional item
x_2 = torch.rand(7, 16) # a sequence of 7, 16-dimensional items
x_3 = torch.rand(5, 16) # a sequence of 5, 16-dimensional items
X = torch.nn.utils.rnn.pad_sequence([x_1, x_2, x_3])
# Can you guess what the shape of X is?
print(X.shape)
# -
del x_1, x_2, x_3, X
# ## Pretrained Word Embeddings
# Moving on-- last assignment, we saw how to train our own word embeddings using a miniature toy corpus. Now, we will see how to easily employ high-quality pretrained word vectors and, later on, how to utilize them for further downstream tasks.
#
# We are going to use [spaCy](https://spacy.io/). SpaCy is a high-level NLP library that provides a ton of useful functionalities, but we will only focus on its pretrained embeddings for this assignment.
#
# Before proceeding, [install spacy](https://spacy.io/usage) using your python package manager (e.g. `pip install spacy`).
# !pip install spacy
import spacy
# SpaCy comes with a lot of different-size models for different languages.
#
# We will need to download the small english model for the exercises to follow. You can either do it on a new terminal window (optimal, if you are running this assignment through a virtual environment) or by simply running the magic command below.
# !python3 -m spacy download en_core_web_lg
# After having downloaded the model, we can load it as follows (you may need to restart your notebook after the download is complete):
nlp = spacy.load('en_core_web_lg')
# We can then use the loaded model to process a sentence and obtain its word vectors, a List of 300-dimensional numpy arrays.
doc = nlp('this is a sentence of 7 words') # the processed sentence
vectors = list(map(lambda x: x.vector, doc)) # its vectors
print('We have {} vectors..'.format(len(vectors)))
print('..each of shape {}'.format(vectors[0].shape))
# And then finally convert them into torch tensors.
torch_vectors = torch.tensor(vectors)
print(torch_vectors.shape)
# Or, in the case of multiple sentences:
# +
# Example sentences
sentences = ['This is a sentence', 'This is another sentence.']
# Parallel processing with spacy
docs = list(map(nlp, sentences))
# Convert each processed sentence into a list of vectors
vectors = map(lambda doc: [word.vector for word in doc], docs)
# Convert each list of vectors into a 2-d torch tensor
tensors = list(map(lambda sentence_vectors: torch.tensor(sentence_vectors), vectors))
# -
# ## POS Tagging
# Given our pretrained embeddings, we may represent sentences as _sequences of vectors_, which is exactly the format expected by an RNN.
# We will now try to train an SRN to iterate over a sentence and assign part of speech tags to each of its words.
# ### Assignment 2.2: Why use an RNN?
# In the context of POS tagging, what is the advantage of using a recurrent network over a feedforward network that processes each word individually?
# When dealing with language we are dealing with a sequence of words that must be processed sequentially. In this context, feedforward networks process all the relevant aspects of an example at once by employing fixed-size input vectors (with associated weights). The approach of the feedforward network makes it difficult to deal with sequences of varying length and fails to capture temporal aspects of the language. One workaround is using a fixed-sized windows of tokens as input and sliding these windows over the input while making prediction. However, this window-based method has some drawbacks:
# * It limits the context from which information can be extracted to the *window-size* "length";
# * It makes it difficult for a network to learn systematic patterns arising from phenomena (like constituency).
#
# Recurrent neural networks (RNN) deal with these challenges by dealing directly with the temporal aspect of language allowing variable length inputs without using fixed-sized windows and therefore caputring the temporal nature of language. RNN process words sequences a word at a time predicting the next word in a sentence by using the current words and the previous hiddent state as input, thus avoiding the context constraints of feedforward networks.
#
# First, let's load and inspect our datafiles.
#
# The pickle file contains three items:
# 1. `sentences`: a List of strings (-sentences)
# 1. `postags`: a List of Lists of strings (-POS tags)
# 2. `pos_to_int`: a Dictionary from strings to ints (mapping each POS tag to a unique identifier)
#
import pickle
with open('TRAIN.p', 'rb') as f:
sentences, postags, pos_to_int = pickle.load(f)
# Now, let us take a moment to understand the data a bit more.
# The POS tags in this dataset are in the style of the Penn Treebank. Find the top 20 most common tags and plot a histogram of their frequencies. Find out what these tags mean linguisically! https://www.ling.upenn.edu/courses/Fall_2003/ling001/penn_treebank_pos.html
# +
from collections import Counter
most_common = {}
for i in range(len(postags)):
for tag in postags[i]:
if tag in most_common.keys():
most_common[tag] += 1
else:
most_common[tag] = 1
k = Counter(most_common)
top_20 = k.most_common(20)
# +
print("20 highest values:")
print("Keys: Values")
for i in top_20:
print(i[0]," :",i[1]," ")
# -
# As expected, the highest count among POS tags is of singular nouns ('NN'), prepositions or subordinate conjunctions ("because", "before", ...) and proper, singular nouns. Interestingly, verbs only appear at the 12th position with the past tense: I would have expected them to appear in a higher position with in its base form or in the third person present singular.
# Next, we need to convert our data to numeric form.
# ### Assignment 2.3: Tensorizing sentences
# Convert sentences to their tensor format, as done earlier (this may take a while).
#
# <div class="alert alert-block alert-warning">
# <b>Important!</b>
# Since the sentences are pre-tokenized (i.e. they are provided as sequences of words rather than strings), we need to change the processing call to ensure the output vectors are aligned with our tokenization.
# </div>
# +
docs = list(map(lambda sentence: spacy.tokens.doc.Doc(nlp.vocab, words=sentence.split()), sentences))
# The coaches should REALLY have made a public notification about the lack of ".split()". I lost hours
#trying to understand what was going on
doc_vectors = list(map(lambda doc: [word.vector for word in doc], docs))
doc_tensors = list(map(lambda sentence_vectors: torch.tensor(sentence_vectors), doc_vectors))
# We no longer need the docs and numpy arrays
del doc_vectors, docs
# -
# Similarly, we will use `pos_to_int` to convert the POS sequences into tensors.
pos_numeric = list(map(lambda pos_sequence: [pos_to_int[pos] for pos in pos_sequence], postags))
pos_tensors = list(map(lambda pos_num_sequence: torch.tensor(pos_num_sequence), pos_numeric))
# In the first assignment, we saw how to split our dataset into a training and a validation set.
#
# Do the same here, splitting the sentences, postags and their corresponding tensors into a training and a validation set.
# +
from sklearn.model_selection import train_test_split
sentences_train, sentences_val, postags_train, postags_val, X_train, X_val, Y_train, Y_val = train_test_split(
doc_tensors, pos_tensors, doc_tensors, pos_tensors, test_size = 0.2)
assert len(X_train) == len(Y_train) == len(sentences_train)
assert len(X_val) == len(Y_val) == len(sentences_val)
# -
# Again, following along the first assignment, we will wrap our tensors into a `Dataset` and a `DataLoader`.
#
# Since our data are not Tensors but rather Lists of Tensors of uneven lengths, we need to write our own Dataset wrapper.
# The wrapper only needs to implement two functions; `__len__`, which expects no arguments and returns the number of samples in the dataset, and `__getitem__`, which accepts an index `idx` and returns the input-output pair `X[idx]`, `Y[idx]`.
#
# Similarly, the Dataloader needs to process the list of input-output pairs produced by the Dataset using `pad_sequence`, as seen earlier.
# ### Assignment 2.4: Padding
# #### a) What is the advantage to applying padding on the batch rather than the entire dataset?
# Applying padding on the batch rather than on the entire dataset allows us to reduce the amount of zeros that will be padded and therefore decreases memory usage. As a matter of fact, the padding gets created equal to the maximum sequence length within one batch which, in the example at hand, is a list of 32 tensors. If we would pad the entire dataset of tensors at the same lengths one would use much more space uselessly as every tensor would have to be padded at the maximum sequence length across the *entire dataset*.
# #### b) Fill in the code for `UnevenLengthDataset` class, implementing its two core functions.
#
# Then, complete the function `pad_batch` which takes a list of (x$_i$, y$_i$) pairs and produces the pair of their paddings: (X, Y).
#
# Given the two, the `DataLoader` object defined can iterate over the Dataset yielding uniform batches ready to be consumed by an RNN.
# +
from torch.nn.utils.rnn import pad_sequence
from torch.utils.data import Dataset, DataLoader
class UnevenLengthDataset(Dataset):
def __init__(self, X: List[FloatTensor], Y: List[LongTensor]) -> None:
self.X = X
self.Y = Y
def __len__(self) -> int:
return(len(self.X))
def __getitem__(self, idx: int) -> Tuple[FloatTensor, LongTensor]:
return(self.X[idx], self.Y[idx])
def pad_batch(batch: List[Tuple[FloatTensor, LongTensor]]) -> Tuple[FloatTensor, LongTensor]:
tokens, tags = zip(*batch)
x = torch.nn.utils.rnn.pad_sequence(tokens, padding_value=0) #necessary padding_value?
y = torch.nn.utils.rnn.pad_sequence(tags, padding_value=0) #necessary padding_value?
return (x, y)
train_dataset = UnevenLengthDataset(X_train, Y_train)
train_dataloader = DataLoader(train_dataset,
collate_fn=pad_batch,
shuffle=True,
batch_size=32)
val_dataset = UnevenLengthDataset(X_val, Y_val)
val_dataloader = DataLoader(val_dataset,
collate_fn=pad_batch,
shuffle=False,
batch_size=32)
# -
# What does a batch look like, shape-wise?
for batch_x, batch_y in train_dataloader:
print(batch_x.shape)
print(batch_y.shape)
break
# So far so good. On to the network.
#
# ### Assignment 2.5: Utility Functions
# Remember how we defined our training and validation functions for the first assignment?
#
# You will need to do the same here.
# Note that while you can use the given code as a guideline, just copying it won't do the trick; unlike a feedforward net, a recurrent network produces a 3rd order output tensor, of shape (max_seq_len, batch_size, num_output_classes).
#
# Similarly, our target Y is a 2nd order tensor of shape (max_seq_len, batch_size).
#
# You will need to properly treat the extra dimensional of both the output and the target, since loss functions expect an order 2 output tensor and an order 1 target tensor.
#
# Complete the functions `train_batch`, `train_epoch`, `eval_batch` and `eval_epoch`.
# +
def accuracy(predictions: LongTensor, truth: LongTensor, ignore_idx: int) -> Tuple[int, int]:
"""
Given a tensor containing the network's predictions and a tensor containing the true values, as well
as an output value to ignore (e.g. the padding value), computes and returns the total count of non-
ignored values as well the total count of correctly predicted values.
predictions: The network's predictions.
truth: The true output labels.
ignore_idx: The output padding value, to be ignored in accuracy calculation.
"""
correct_words = torch.ones(predictions.size())
correct_words[predictions != truth] = 0
correct_words[truth == ignore_idx] = 1
num_correct_words = correct_words.sum().item()
num_masked_words = len(truth[truth == ignore_idx])
return predictions.shape[0] * predictions.shape[1] - num_masked_words, num_correct_words - num_masked_words
def measure_accuracy(network: torch.nn.Module,
dataloader: DataLoader,
device: str) -> float:
"""
Given a network, a dataloader and a device, iterates over the dataset and returns the network's accuracy.
"""
correct = 0
total = 0
for x_batch, y_batch in dataloader:
pred = network(x_batch.to(device))
local_total, local_correct = accuracy(pred.argmax(dim=-1), y_batch.to(device), ignore_idx=0)
correct+= local_correct
total+= local_total
return correct/total
def train_batch(network: torch.nn.Module,
X_batch: FloatTensor,
Y_batch: LongTensor,
loss_fn: Callable[[FloatTensor, FloatTensor], FloatTensor],
optimizer: torch.optim.Optimizer) -> float:
network.train()
pred = network(X_batch)
pred = pred.view(pred.shape[0]*pred.shape[1], pred.shape[2])
batch_loss = loss_fn(pred, Y_batch.view(-1)) # loss calculation
batch_loss.backward() # gradient computation
optimizer.step() # back-propagation
optimizer.zero_grad() # gradient reset
return batch_loss.item()
def train_epoch(network: torch.nn.Module,
dataloader: DataLoader,
loss_fn: Callable[[FloatTensor, FloatTensor], FloatTensor],
optimizer: torch.optim.Optimizer,
device: str) -> float:
loss = 0.
for i, (x_batch, y_batch) in enumerate(dataloader):
x_batch = x_batch.to(device) # convert back to your chosen device
y_batch = y_batch.to(device)
loss += train_batch(network=network, X_batch=x_batch, Y_batch=y_batch, loss_fn=loss_fn, optimizer=optimizer)
loss /= (i+1) # divide loss by number of batches for consistency
return loss
def eval_batch(network: torch.nn.Module,
X_batch: FloatTensor,
Y_batch: LongTensor,
loss_fn: Callable[[FloatTensor, LongTensor], FloatTensor]) -> float:
network.eval()
#because we are in eval mode no backprop is needed
with torch.no_grad():
pred = network(X_batch) # forward pass
pred = pred.view(pred.shape[0]*pred.shape[1], pred.shape[2])
batch_loss = loss_fn(pred, Y_batch.view(-1)) # loss calculation
return batch_loss.item()
def eval_epoch(network: torch.nn.Module,
# a list of data points x
dataloader: DataLoader,
loss_fn: Callable[[FloatTensor, LongTensor], FloatTensor],
device: str) -> float:
loss = 0.
# Copied from above with modification due to being eval mode and not training
for i, (x_batch, y_batch) in enumerate(dataloader):
x_batch = x_batch.to(device) # convert back to your chosen device
y_batch = y_batch.to(device)
loss += eval_batch(network=network, X_batch=x_batch, Y_batch=y_batch, loss_fn=loss_fn)
loss /= (i+1) # divide loss by number of batches for consistency
return loss
# -
# ### Assignment 2.6: SRN POS tagging
# Define a simple recurrent network, with input size compatible with the vector dimensionality, output size compatible with the number of output classes (the number of different POS tags + 1) and a hidden size of your own choice. What is a reasonale choice?
# Input size = 300; output size = 37; hidden size = between 300 and 49.
# #### a) Why do we need to add 1 to the number of output classes?
# Because we need to take into consideration that the network wants us to start at 0 with the class numbers, therefore we have to add one more class.
# #### b) Implementation
#
# Use `"tanh"` as your hidden layer activation, and choose **an appropriate combination of output activation and loss function** (consider the task at hand, and refer to the documentation if in doubt- refer to tutorial as well!).
#
# Then instantiate an optimizer over your network, and train it for a number of epochs, measuring and printing all metrics in the process (train and validation loss and accuracy).
#
# _Hint_: Use `measure_accuracy` (defined earlier) to obtain accuracy.
#
# Plot the loss curves over the training process.
# +
srn = fastSRN(input_dim=300,
hidden_dim=150,
hidden_activation='tanh',
output_activation=torch.nn.LogSoftmax(dim=-1), #
output_dim=49,
device='cpu')
opt = torch.optim.Adam(srn.parameters(), lr=0.003)
loss_fn = torch.nn.NLLLoss()
# | K-class Classification | LogSoftmax | NLLLoss or CrossEntropyLoss - combines SoftMax and NLLLoss |
# +
# Your training script here
NUM_EPOCHS = 50
device = 'cpu'
train_losses = []
val_losses = []
train_accuracies = []
val_accuracies = []
for i in range(NUM_EPOCHS):
train_loss = train_epoch(srn, train_dataloader, optimizer=opt, loss_fn=loss_fn, device=device)
val_loss = eval_epoch(srn, val_dataloader, loss_fn, device=device)
train_accuracy = measure_accuracy(srn, train_dataloader, device=device)
val_accuracy = measure_accuracy(srn, val_dataloader, device=device)
if i%10 == 0:
print('Epoch {};'.format(i))
print(' Training Loss: {};'.format(train_loss))
print(' Validation Loss: {};'.format(val_loss))
print(' Training accuracy:', train_accuracy)
print(' Validation accuracy:', val_accuracy)
train_losses.append(train_loss)
val_losses.append(val_loss)
train_accuracies.append(train_accuracy)
val_accuracies.append(val_accuracy)
# +
# %matplotlib inline
from matplotlib import pyplot as plt
plt.plot(train_losses)
plt.plot(val_losses)
plt.plot(train_accuracies)
plt.plot(val_accuracies)
plt.legend(['Training loss', 'Validation loss', 'Train accuracy', 'Validation accuracy'])
plt.show()
# -
# The test accuracy on this task should be well over 90%. If you are getting an accuracy much below this, play with your hyperparameters and try to improve.
#
# #### c) What is your final accuracy?
# 0.9680520442986152
# #### d) A little error analysis
# Minimally, find a few instances of sentences with wrong tags. Can you say why these mistakes are made?
# Optionally, feel free is do a full error analysis. What are the most commonly confused tags for an English POS tagger?
srn.parameters()
test = srn(['In their house, everything comes in pairs. There’s his car and her car, his towels and her towels, and his library and hers.'])
print(test)
# **Congratulations!** 🎉 You are done with the required part! Now for some fun:
# ### Optional[CCG Supertagging]
# See blackboard!
| courses/mlhvl/2-assignments/4-ass-4/Sequence_RNN_2_final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:dgs]
# language: python
# name: conda-env-dgs-py
# ---
import sys
sys.path.append("../")
from dgs.data.denotation_graph import DenotationGraph, visualize_denotation_graph
help(DenotationGraph)
dg = DenotationGraph(graph_folder_path="../../../flicker30k_denotation_graph/flicker30k/graph")
len(dg.leaf_nodes)
len(dg.nodes)
len(dg.edges)
dg.get_node_info(0).keys()
dg.get_node_info(0)["text"]
len(dg.get_node_info(0)["images"])
dg.get_edge_info((0,54))
dg.leaf_nodes[0]
# there are a few leaf nodes which are not the full captions too. Not sure why though!
dg.get_node_info(dg.leaf_nodes[0])["text"]
# There are also a few leaf nodes which contains the full captions
print(dg.leaf_nodes[-1])
print(dg.get_node_info(dg.leaf_nodes[-1]))
dg.get_in_edges(dg.leaf_nodes[0])
# This should be an empty list as leaf does not have any children
dg.get_children_nodes(dg.leaf_nodes[0])
dg.get_parent_nodes(dg.leaf_nodes[0])
assert dg.leaf_nodes[0] in dg.get_children_nodes(1105)
len(dg.get_children_nodes(1105))
assert len(dg.get_children_nodes(1105)) == len(dg.get_out_edges(1105))
len(dg.get_descendents(0))
len(dg.get_ancestors(dg.leaf_nodes[0]))
# find the sibling nodes
siblings = dg.get_sibling_nodes(5635)
list(dg.get_parent_nodes(5635))
len(siblings)
assert 5635 not in siblings[1105]
assert 5635 not in siblings[14]
cousins = dg.get_cousin_nodes(dg.leaf_nodes[-1])
len(cousins)
dg.edges[-1]
def get_immediate_in_out_edges(node_idx, graph):
edges = graph.get_in_edges(node_idx)
edges.extend(graph.get_out_edges(node_idx))
return edges
# find all the incomming and outgoing edges for a random node.
# useful for visualization
edges= get_immediate_in_out_edges(7996, dg)
len(edges)
subgraph = dg.get_subgraph_from_edges(edges[:10]) # create a subgraph from a random subset of 10 edges
len(subgraph.edges)
visualize_denotation_graph(subgraph, "./subgraph.png")
from IPython import display
display.Image(filename="./subgraph.png")
| notebooks/using_denotation_graph.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Before you turn this problem in, make sure everything runs as expected. First, **restart the kernel** (in the menubar, select Kernel$\rightarrow$Restart) and then **run all cells** (in the menubar, select Cell$\rightarrow$Run All).
#
# Make sure you fill in any place that says `YOUR CODE HERE` or "YOUR ANSWER HERE", as well as your name and collaborators below:
NAME = ""
COLLABORATORS = ""
# ---
# <!--NOTEBOOK_HEADER-->
# *This notebook contains material from [PyRosetta](https://RosettaCommons.github.io/PyRosetta.notebooks);
# content is available [on Github](https://github.com/RosettaCommons/PyRosetta.notebooks.git).*
# <!--NAVIGATION-->
# < [`GALigandDock` Protocol with `pyrosetta.distributed` Using the `beta_cart.wts` Scorefunction](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/08.02-Ligand-Docking-pyrosetta.distributed.ipynb) | [Contents](toc.ipynb) | [Index](index.ipynb) | [Working With Density](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/11.00-Working-With-Density.ipynb) ><p><a href="https://colab.research.google.com/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/10.00-Working-With-Symmetry.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open in Google Colaboratory"></a>
# # Working With Symmetry
# Keywords: symmetry, asymmetric, SetupForSymmetryMover, virtual
#
# ## Overview
# Symmetry is an important concept to learn when working with biomolecules. When a protein is crystalized, it is in the precense of its symmetrical neighbors - which can be important if testing particular protocols or using crystal density for refinement or full structure building.
#
# Symmetry can also be useful for designing symmetrical structures or large repeating meta-proteins like protein cages.
#
# ### Symmetry In Rosetta
# So why do we care if our protein is symmetrical or not when it comes to Rosetta? Each residue and atom that is loaded into Rosetta takes time to both load, and time to score. Since scoring can happen thousands of times - even in a short protocol, anything we can do to speed this up becomes important. The most expensive operation in Rosetta is minimization, and by using symmetry - we can reduce the minimization time exponentially by minimizing a single copy instead of ALL copies. We will get into the details about how this works below.
#
# When we use symmetry in Rosetta - we are basically telling rosetta that the symmetrical partners are 'special', however, the total number of residues is now ALL residues, including symmetrical partners. Upon setting up symmety in Rosetta, Rosetta will replace the `Conformation` within the pose with a **Symmetrical** version, called the `SymmetricConformation`. If you know anything about classes, this `SymmetricConformation` is derived from the actual `Conformation` object, but contains extra information about the pose and some functions are replaced.
#
# ### Symmetric Scoring and Moving
# Ok, so now lets assume that we have our symmetric pose. Now what? Well, the symmetric copies are all tied to their real counterparts. Once you move a chain, residue, or atom by packing or minimization, the symmetric copies of that residue are all moved in the same way.
#
# Cool. But what about scoring? Scoring works very similarly - instead of scoring each and every residue in our pose, Rosetta will score just our assymetric unit, and multiply that out to the number of symmetric copies we have. Intelligently, Rosetta will also figure out the symmetric interfaces that arise from the interactions of our assymetric unit to the symmetric copies and score them appropriately.
#
# ### Symmetry-aware movers
# Most of our common movers are symmetry-aware. At one point there were different symmetric and non-symmetric versions of particular code, such as MinMover and PackRotamersMover. Now though, Rosetta will automatically use the pose to figure out what needs to be done. You should seek original documentation (and contact the author if not explicit) to make sure that an uncommon protocol you are using is symmetry-aware.
#
# ## Documentation
# More information on RosettaSymmetry can be found in the following places:
# - https://www.rosettacommons.org/docs/latest/rosetta_basics/structural_concepts/symmetry
# - https://www.rosettacommons.org/demos/latest/tutorials/Symmetry/Symmetry
# - https://www.rosettacommons.org/docs/latest/application_documentation/utilities/make-symmdef-file
# - https://www.rosettacommons.org/docs/latest/scripting_documentation/RosettaScripts/Movers/movers_pages/SetupForSymmetryMover
# - https://www.rosettacommons.org/docs/latest/scripting_documentation/RosettaScripts/Movers/movers_pages/ExtractAsymmetricUnitMover
#
# Notebook setup
import sys
if 'google.colab' in sys.modules:
# !pip install pyrosettacolabsetup
import pyrosettacolabsetup
pyrosettacolabsetup.setup()
print ("Notebook is set for PyRosetta use in Colab. Have fun!")
# Here, we will use a few specific options. The first three options make Rosetta a bit more robust to input structures. The `-load_PDB_components` cannot be used with glycans, unfortunately, and our structure has a few very important glycans. Finally, we load a bunch of glycan-specific options, which we will cover in the next tutorial.
# +
from pyrosetta import *
from pyrosetta.rosetta import *
from pyrosetta.teaching import *
import os
init('-ignore_unrecognized_res -load_PDB_components false -ignore_zero_occupancy false @inputs/glycan_flags')
# -
# ## Creating a SymDef file
#
# Here, we will start with how to create a basic symdef file for cyrstal symmetry. Note that there are ways to do this without a symdef file, but these do not currently work for glycan structures, which we will be using here.
#
# The `make_symdef_file.pl` file is within Rosetta3. To use it, you will need to download and licence Rosetta3. The code is in the `Rosetta/main/src/apps/public` directory. In the interest of reducing code drift, this file is NOT included in the tutorial directory as we may then have version drift.
#
# If you have done this, we can use the following command to create the symdef file. Here, the radius of symmetrical partners is 12A, which is certainly fairly large, but produces a very well represented crystal.
pdb = "inputs/1jnd.pdb"
base_cmd = f'cd inputs && make_symmdef_file.pl -r 12 -m CRYST -p {pdb}.pdb > {pdb}_crys.symm && cd -'
print(base_cmd)
# Use this base command and the `os.system(cmd)` function to run the code or use the provided symdef file.
os.system('cp inputs/1jnd_crys.symm .')
# Take a look at the symmetrized structure in pymol (`inputs/1jnd_symm.pdb`). What would happen if we increased the radius to 24 instead of 12?
# ## Setup a Symmetrized Pose
#
# Here, we will run a basic Rosetta protocol with symmetry. There are much more complicated things you can do with symmetry, but for now, we just want to symmetrically pack the protein. Please see the docs for more on symmetry. The full Rosetta C++ tutorial for symmetry is a great place to go from here: - https://www.rosettacommons.org/demos/latest/tutorials/Symmetry/Symmetry
#
# Lets first create a pose, and then use the `SetupForSymmetryMover` on the pose. Note this is an unrefined input structure. This is so that minmover will actually do something. A pareto-optimal refined structure can be found in the inputs as `1jnd_refined.pdb.gz`
p = pose_from_pdb('inputs/1jnd.pdb')
original = p.clone()
p.total_residue()
type(p.conformation())
symmetrize = rosetta.protocols.symmetry.SetupForSymmetryMover("1jnd_crys.symm")
symmetrize.apply(p)
print(p.total_residue())
print(type(p.conformation()))
# How many symmetric copies do we have in our pose?
# How do the scores compare for our original pose and our symmetrized version?
# Now lets use some of the functionality to understand how this all works. We can use the `SymetricInfo` object that is part of the `SymmetricConformation` to get at some info. Lets take a look at all residues and find the assymetric unit residues and equivalent residues for the rest.
print("AssymUnit? equivalent_res")
sym_info = p.conformation().Symmetry_Info()
for i in range(1, p.size()+1):
print(i, sym_info.bb_is_independent(i), sym_info.bb_follows(i))
# Which residues are our original pose residues? Note that the final residues are called `Virtual` residues. Virtual residues are not scored. They have coordinates, and can move, but simply result in a score of zero. They are useful in some contexts to hide a part of the pose from the scoring machinery, and there are movers that can change residues to and from virtual. In this case, they are used for the FoldTree - in order to allow refinement of the full crystal environment. They allow relative movement of each subunit relative to each other. There are two virtual residues for each subunit
print(p.residue(3654))
print("Total Subunits:", (3654-18)/404)
print("Total Subunits:", sym_info.subunits())
score = get_score_function()
print(score(original))
print(score(p))
# ## Running Protocols with Symmetry
#
# Now, lets try running a minimization with symmetry on.
mm = MoveMap()
mm.set_bb(True)
mm.set_chi(True)
minmover = rosetta.protocols.minimization_packing.MinMover()
minmover.score_function(score)
minmover.set_movemap(mm)
if not os.getenv("DEBUG"):
minmover.apply(p)
score(p)
# How does our pose look? For being such a large pose, how was the speed of minimization?
#
# How does this compare to our refined pose? Try to copy a subunit to a new object in PyMol. Then use the align command to align it to our assymetric unit. What is the RMSD?
# Now lets pack with our symmetric structure.
# +
from rosetta.core.pack.task import *
from rosetta.core.pack.task.operation import *
packer = PackRotamersMover()
tf = rosetta.core.pack.task.TaskFactory()
tf.push_back(RestrictToRepacking())
tf.push_back(IncludeCurrent())
packer.task_factory(tf)
p = original.clone()
symmetrize.apply(p)
# -
if not os.getenv("DEBUG"):
packer.apply(p)
print("packed", score(p))
# ## Conclusions
#
# Symmetry is a useful tool in the Rosetta Library. There are also selectors and movers that you may find useful, such as the `AsymmetricUnitSelector` in `rosetta.core.select.residue_selectors` and the `ExtractAsymmetricUnitMover`, which will give you back just the single subunit, without any asymetric partners, and the `ExtractAsymmetricPoseMover`, which will remove 'symmetry' information and give you back a pose with all the subunits. The later of these can be found by importing `rosetta.protocols.symmetry`.
#
# Note that not ALL protocols will respect symmetry - so please check the original documentation to see if symmetry is supported. If you are unsure, please email the developer.
# **Chapter contributors:**
#
# - <NAME> (Scripps; Institute for Protein Innovation)
# <!--NAVIGATION-->
# < [`GALigandDock` Protocol with `pyrosetta.distributed` Using the `beta_cart.wts` Scorefunction](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/08.02-Ligand-Docking-pyrosetta.distributed.ipynb) | [Contents](toc.ipynb) | [Index](index.ipynb) | [Working With Density](http://nbviewer.jupyter.org/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/11.00-Working-With-Density.ipynb) ><p><a href="https://colab.research.google.com/github/RosettaCommons/PyRosetta.notebooks/blob/master/notebooks/10.00-Working-With-Symmetry.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open in Google Colaboratory"></a>
| student-notebooks/10.00-Working-With-Symmetry.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.10 64-bit
# language: python
# name: python3
# ---
# +
import numpy as np
import collections
import pandas as pd
import os
import math
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.metrics.pairwise import cosine_similarity
from sklearn.model_selection import train_test_split
from scipy.sparse.linalg import svds
import warnings; warnings.simplefilter('ignore')
from scipy import sparse
from typing import List
import scipy.sparse as sp
for dirname, _, filenames in os.walk('/home/ebcffhh/Documents/personal/Masters/Thesis'):
for filename in filenames:
print(os.path.join(dirname, filename))
# -
data=pd.read_csv("/home/ebcffhh/Documents/personal/Masters/Thesis/ratings_Beauty.csv", names = ["userId", "ProductId", "Ratings", "Timestamp"])
counts=data.userId.value_counts()
dataset_final=data[data.userId.isin(counts[counts>=25].index)]
print('Number of users who have rated 25 or more items =', len(dataset_final))
print('Number of unique users in the final data = ', dataset_final['userId'].nunique())
print('Number of unique products in the final data = ', dataset_final['ProductId'].nunique())
rated_products = data.groupby(by='userId',as_index=False)['Ratings'].count()
print(rated_products)
rated_products = rated_products[rated_products['Ratings'] < 20]
new_dataset = data.loc[~((data.userId.isin(rated_products['userId']))),:]
no_of_rated_products_per_user = new_dataset.groupby(by='userId')['Ratings'].count().sort_values(ascending=False)
print(no_of_rated_products_per_user)
print(new_dataset.ProductId.nunique())
print(new_dataset.userId.nunique())
n_users = new_dataset.userId.unique().shape[0]
n_products = new_dataset.ProductId.unique().shape[0]
product_list = new_dataset.ProductId.unique().tolist()
print(n_users)
from collections import defaultdict
def GetTopN(predictions, n=10, minimumRating=4.0):
topN = defaultdict(list)
for userID, productId, actualRating, estimatedRating, _ in predictions:
if (estimatedRating >= minimumRating):
topN[userID].append((productId, estimatedRating))
for userID, ratings in topN.items():
ratings.sort(key=lambda x: x[1], reverse=True)
topN[userID] = ratings[:n]
return topN
def get_customer_satisfaction(pred_u,k):
edt = {}
rating_list = defaultdict(list)
pred = pred_u.copy().groupby(['userId'])
for userId in pred.groups.keys():
sorted_pred_group = pred.get_group(userId).sort_values(['prediction'], ascending = False)
top_k = sorted_pred_group[:k]
top_k_g = top_k.groupby(by='userId')
for userId in top_k_g.groups.keys():
top_k_user_list = top_k_g.get_group(userId)
for _, groups in top_k_user_list.iterrows():
diff_ratings = groups['prediction'] - groups['actual']
rating_list.setdefault(groups['userId'], []).append(diff_ratings)
edt[userId] = (np.sum(rating_list.get(userId)))
return edt
def prediction_coverage(predicted: List[list], catalog: list) -> float:
predicted_flattened = [p for sublist in predicted for p in sublist]
unique_predictions = len(set(predicted_flattened))
prediction_coverage = round(unique_predictions/(len(catalog)* 1.0)*100,2)
return prediction_coverage
# +
def recommender_precision(predicted: List[list], actual: List[list]) -> int:
def calc_precision(predicted, actual):
prec = [value for value in predicted if value in actual]
prec = np.round(float(len(prec)) / float(len(predicted)), 4)
return prec
precision_list = list(map(calc_precision, predicted, actual))
precision = np.mean(precision_list)
return precision, precision_list
def recommender_recall(predicted: List[list], actual: List[list]) -> int:
def calc_recall(predicted, actual):
reca = [value for value in predicted if value in actual]
reca = np.round(float(len(reca)) / float(len(actual)), 4)
return reca
recall_list = list(map(calc_recall, predicted, actual))
recall = np.mean(recall_list)
return recall, recall_list
# -
def personalization(predicted: List[list]) -> float:
"""
Personalization measures recommendation similarity across users.
A high score indicates good personalization (user's lists of recommendations are different).
A low score indicates poor personalization (user's lists of recommendations are very similar).
A model is "personalizing" well if the set of recommendations for each user is different.
Parameters:
----------
predicted : a list of lists
Ordered predictions
example: [['X', 'Y', 'Z'], ['X', 'Y', 'Z']]
Returns:
-------
The personalization score for all recommendations.
"""
def make_rec_matrix(predicted: List[list]) -> sp.csr_matrix:
df = pd.DataFrame(data=predicted).reset_index().melt(
id_vars='index', value_name='item',
)
df = df[['index', 'item']].pivot(index='index', columns='item', values='item')
df = pd.notna(df)*1
rec_matrix = sp.csr_matrix(df.values)
return rec_matrix
#create matrix for recommendations
predicted = np.array(predicted)
rec_matrix_sparse = make_rec_matrix(predicted)
#calculate similarity for every user's recommendation list
similarity = cosine_similarity(X=rec_matrix_sparse, dense_output=False)
avg_sim = similarity.mean(axis=1)
#get indicies for upper right triangle w/o diagonal
upper_right = np.triu_indices(similarity.shape[0], k=1)
#calculate average similarity score of all recommended items in list
ils_single_user = np.mean(similarity[upper_right])
return avg_sim, (1 - ils_single_user)
#return similarity
# +
from surprise import Dataset
from surprise import Reader
from surprise.model_selection import train_test_split
reader = Reader()
rating_data = Dataset.load_from_df(new_dataset[['userId', 'ProductId', 'Ratings']], reader)
trainset, testset = train_test_split(rating_data, test_size=0.2,random_state=100)
# +
from surprise import SVD
from surprise import KNNWithMeans
from surprise import accuracy
k = [5, 10, 15, 20, 30, 40, 50, 60, 70, 80, 90, 100]
mae_svd = list()
for i in k:
algo = SVD(n_factors=i, n_epochs=200)
algo.fit(trainset)
test_pred = algo.test(testset)
mae_svd.append(accuracy.mae(test_pred))
print("Mean Absolute Error for value k {} is ".format(i), accuracy.mae(test_pred))
# +
from surprise import SVD
from surprise import KNNWithMeans
from surprise import Dataset
from surprise.model_selection import cross_validate
from surprise import Reader
from surprise.model_selection import train_test_split
from surprise import accuracy
k = [5, 10, 15, 20, 30, 40, 50, 60, 70, 80, 90, 100]
reader = Reader()
rating_data = Dataset.load_from_df(new_dataset[['userId', 'ProductId', 'Ratings']], reader)
trainset, testset = train_test_split(rating_data, test_size=0.2,random_state=100)
mae_knn = list()
for i in k:
algo = KNNWithMeans(k=i, sim_options={'name':'pearson','user_based': True})
algo.fit(trainset)
test_pred = algo.test(testset)
mae_knn.append(accuracy.mae(test_pred))
print("Mean Absolute Error for value k {} is ".format(i), accuracy.mae(test_pred))
# +
from scipy.stats import entropy
def get_f1_score(predictions, k):
threshold = 4
# First map the predictions to each user.
user_est_rating = defaultdict(list)
for index, row in predictions.iterrows():
user_est_rating[row['userId']].append((row['prediction'], row['actual']))
# Then sort the predictions for each user and retrieve the k highest ones.
f1_score = dict()
for uid, user_ratings in user_est_rating.items():
user_ratings.sort(key=lambda x:x[0], reverse=True)
# Number of relevant items
n_rel = sum((r_ui >= threshold) for (_, r_ui) in user_ratings)
if math.isnan(n_rel):
print("nan value for rel")
# Number of recommended items in top k
n_rec_k = sum((est >= threshold) for (est, _) in user_ratings[:k])
if math.isnan(n_rec_k):
print("nan value for rel")
# Number of relevant and recommended items in top k
n_rel_and_rec_k = sum(((true_r >= threshold) and (est >= threshold))
for (est, true_r) in user_ratings[:k])
if math.isnan(n_rel_and_rec_k):
print("nan value for rel and rec")
precision = n_rel_and_rec_k / n_rec_k if n_rec_k != 0 else 0
recall = n_rel_and_rec_k / n_rel if n_rel != 0 else 0
f1_score[uid] = 2 * ((precision * recall)/(precision+recall)) if (precision + recall) != 0 else 0
return f1_score
def get_cus(predictions, k):
threshold = 4
# First map the predictions to each user.
user_est_rating = defaultdict(list)
for index, row in predictions.iterrows():
user_est_rating[row['userId']].append((row['prediction'], row['actual']))
# Then sort the predictions for each user and retrieve the k highest ones.
cus = defaultdict(list)
for uid, user_ratings in user_est_rating.items():
user_ratings.sort(key=lambda x:x[0], reverse=True)
for est, r_ui in user_ratings[:k]:
diff = r_ui - est
cus[uid].append(diff)
customerSatisfaction = {}
for key in cus:
customerSatisfaction[key] = np.sum(cus.get(key))/k
return customerSatisfaction
def get_f1_score_nn(predictions, k):
threshold = 4
# First map the predictions to each user.
user_est_rating = defaultdict(list)
#for uid, iid, r_ui, est in predictions:
# user_est_rating[uid].append((est, r_ui))
for index, row in predictions.iterrows():
user_est_rating[row['userId']].append((row['prediction'], row['actual']))
# Then sort the predictions for each user and retrieve the k highest ones.
f1_score = dict()
for uid, user_ratings in user_est_rating.items():
user_ratings.sort(key=lambda x:x[0], reverse=True)
# Number of relevant items
n_rel = sum((r_ui >= threshold) for (_, r_ui) in user_ratings)
if math.isnan(n_rel):
print("nan value for rel")
# Number of recommended items in top k
n_rec_k = sum((est >= threshold) for (est, _) in user_ratings[:k])
if math.isnan(n_rec_k):
print("nan value for rel")
# Number of relevant and recommended items in top k
n_rel_and_rec_k = sum(((true_r >= threshold) and (est >= threshold))
for (est, true_r) in user_ratings[:k])
if math.isnan(n_rel_and_rec_k):
print("nan value for rel and rec")
precision = n_rel_and_rec_k / n_rec_k if n_rec_k != 0 else 0
recall = n_rel_and_rec_k / n_rel if n_rel != 0 else 0
f1_score[uid] = 2 * ((precision * recall)/(precision+recall)) if (precision + recall) != 0 else 0
return f1_score
def cal_f1(test_pred, k):
f1_scores = get_f1_score(test_pred, k)
average_f1_score = sum(score for score in f1_scores.values())/ len(f1_scores)
return f1_scores , average_f1_score
def get_accuracy(predictions,k):
user_est_rating = defaultdict(list)
for index, row in predictions.iterrows():
user_est_rating[row['userId']].append((row['prediction'], row['actual']))
accuracy_scores = dict()
for uid, user_ratings in user_est_rating.items():
scores = list()
user_ratings.sort(key=lambda x:x[0], reverse=True)
for (est, actual) in user_ratings[:k]:
diff = abs(actual - est)
scores.append(diff)
accuracy_scores[uid] = sum(score for score in scores)/len(scores)
return accuracy_scores
def cal_accuracy(test_pred, k):
accuracy = get_accuracy(test_pred, k)
average_accuracy = sum(score for score in accuracy.values())/len(accuracy)
return accuracy , average_accuracy
def get_shannon_entropy(predictions, product_list, no_of_recommendations):
recommendation_items = [item for sublist in predictions for item in sublist]
products = set(recommendation_items)
count_recommendation_items = collections.Counter(recommendation_items)
print(count_recommendation_items)
pi = list()
for product in products:
#if product in count_recommendation_items.keys():
pi.append(count_recommendation_items.get(product)/len(set(product_list)))
#e = -np.sum(pi*np.log(pi)/np.log(no_of_recommendations))
e = -np.sum(pi*np.log(pi))
return e
#print("Average diversity using shannon entropy for {} no of recommendations is {} \n".format(no_of_recommendations, -np.sum(pi*np.log(pi)/np.log(no_of_recommendations))))
def get_shannon_entropy_new(predictions, product_list, no_of_recommendations):
recommendation_items = [item for sublist in predictions for item in sublist]
products = set(recommendation_items)
count_recommendation_items = collections.Counter(recommendation_items)
n_rec = sum(count_recommendation_items.values())
c = np.fromiter(count_recommendation_items.values(), dtype=int)
pi = c/n_rec
shannon_entropy = -np.sum(pi * np.log2(pi))
return shannon_entropy
def get_s_entropy(predictions, count_recommended_products, total_products):
entropy_list = []
for recommended_product_list in predictions:
probability_list_each_user = []
for recommended_product in recommended_product_list:
probability_list_each_user.append(count_recommended_products.get(recommended_product)/len(total_products))
entropy_list.append(entropy(probability_list_each_user))
return np.mean(entropy_list)
# -
import re
def sorted_nicely( l ):
""" Sorts the given iterable in the way that is expected.
Required arguments:
l -- The iterable to be sorted.
"""
convert = lambda text: int(text) if text.isdigit() else text
alphanum_key = lambda key: [convert(c) for c in re.split('([0-9]+)', key)]
return sorted(l, key = alphanum_key)
# +
from surprise import SVD
from surprise import Dataset
from surprise import accuracy
from surprise.model_selection import train_test_split
from collections import Counter
import recmetrics
import csv
ratings_dataset = Dataset.load_from_df(new_dataset[['userId', 'ProductId', 'Ratings']],reader)
trainset, testset = train_test_split(ratings_dataset, test_size=.2)
total_products = set([x[1] for x in testset])
product_list = set()
for inner_pid in trainset.ir.keys():
product_list.add(trainset.to_raw_iid(inner_pid))
algo = SVD(n_factors= 80, n_epochs=200)
algo.fit(trainset)
predictions = algo.test(testset)
test = pd.DataFrame(predictions)
test = test.rename(columns={'uid':'userId', 'iid': 'productId',
'r_ui':'actual', 'est':'prediction'})
pred_user = test.copy().groupby('userId', as_index=False)['productId'].agg({'ratings': (lambda x: list(set(x)))})
pred_user = pred_user.set_index("userId")
cf_model = test.pivot_table(index='userId',
columns='productId', values='prediction').fillna(0)
def get_users_predictions(user_id, n, model):
recommended_items = pd.DataFrame(model.loc[user_id])
recommended_items.columns = ["predicted_rating"]
recommended_items = recommended_items.sort_values('predicted_rating', ascending=False)
recommended_items = recommended_items.head(n)
return recommended_items.index.tolist()
def get_recs(model, k):
recs = []
for user in model.index:
cf_predictions = get_users_predictions(user, k, model)
recs.append(cf_predictions)
return recs
productId_counts = dict(new_dataset.ProductId.value_counts())
userId_counts = test['userId'].value_counts()
diversity_svd = []
novelty_svd = []
coverage_svd = []
f1_score_svd = []
accuracy_svd = []
entropy_svd = []
# Top-n recommendations for each user
no_of_recommendations = [5, 10, 15, 20, 30, 40, 50, 60, 70, 80, 90, 100]
#no_of_recommendations = [5]
for k in no_of_recommendations:
recs = get_recs(cf_model, k)
flat_list_recommendations = [item for sublist in recs for item in sublist]
product_counts = Counter(flat_list_recommendations)
pred_user[f'Top-{k} Recommendation'] = recs
# To calculate entropy
entropy_score = get_s_entropy(recs, product_counts, set(flat_list_recommendations))
print("The entropy score for {} recommendation is {} \n".format(k, entropy_score))
entropy_svd.append(entropy_score)
# To calculate the f1_score
f1_scores_list_svd, average_f1_score_svd = cal_f1(test.copy() ,k)
print("The f1 score for {} recommendation is {} \n".format(k, average_f1_score_svd))
f1_score_svd.append(average_f1_score_svd)
# To calculate accuracy
accuracy_scores_svd, average_accuracy_svd = cal_accuracy(test.copy() ,k)
print("The accuracy score for {} recommendation is {} \n".format(k, average_accuracy_svd))
accuracy_svd.append(average_accuracy_svd)
# To calculate the diversity
diversity_scores_svd, average_diversity_svd = personalization(list(recs))
#diversity = get_shannon_entropy_new(recs, list(product_list), k)
print("The diversity score for {} recommendation is {} \n".format(k, average_diversity_svd))
diversity_svd.append(average_diversity_svd)
# To calculate the novelty
cf_novelty_svd, novelty_list_svd = recmetrics.novelty(recs, productId_counts, len(userId_counts), k)
print("The novelty score for {} recommendation is {} \n".format(k, cf_novelty_svd))
novelty_svd.append(cf_novelty_svd)
# To calculate the coverage
cf_coverage = recmetrics.catalog_coverage(list(recs), product_list, 100)
print("The coverage score for {} recommendation is {} \n".format(k, cf_coverage))
coverage_svd.append(cf_coverage)
# To calculate the customer satisfaction
edt_svd = get_customer_satisfaction(test, k)
print("The cusotmer satisfaction for {} recommendation is {}".format(k, np.mean(list(edt_svd.values()))))
filename = "/home/ebcffhh/thesis/sorted_svd/metrics_svd_%s_recommendations.csv" % k
with open(filename, 'w') as csv_file:
writer = csv.writer(csv_file)
writer.writerow(['userId', 'accuracy', 'f1_score', 'diversity', 'novelty', 'customer_satisfaction'])
for i, nov, (uid, acc_score), (_, f1_score) in zip(diversity_scores_svd, novelty_list_svd, accuracy_scores_svd.items(), f1_scores_list_svd.items()):
if uid in sorted_nicely(edt_svd.keys()):
writer.writerow([uid, acc_score, f1_score, (1 - i[0]), nov, edt_svd.get(uid)])
print(accuracy_svd)
# +
from surprise import KNNWithMeans
from surprise import Dataset
from surprise import accuracy
from surprise.model_selection import train_test_split
import recmetrics
#ratings_dataset = Dataset.load_from_df(new_dataset[['userId', 'ProductId', 'Ratings']],reader)
#trainset, testset = train_test_split(ratings_dataset, test_size=.2)
train_product_list_count = len(trainset.ir.keys())
productId_counts = dict(new_dataset.ProductId.value_counts())
userId_counts = new_dataset['userId'].value_counts()
product_list = set()
for inner_pid in trainset.ir.keys():
product_list.add(trainset.to_raw_iid(inner_pid))
algo = KNNWithMeans(k=100, sim_options={'name':'pearson','user_based': True})
algo.fit(trainset)
predictions = algo.test(testset)
test = pd.DataFrame(predictions)
test = test.rename(columns={'uid':'userId', 'iid': 'productId',
'r_ui':'actual', 'est':'prediction'})
pred_user = test.copy().groupby('userId', as_index=False)['productId'].agg({'ratings': (lambda x: list(set(x)))})
pred_user = pred_user.set_index("userId")
cf_model = test.pivot_table(index='userId',
columns='productId', values='prediction').fillna(0)
def get_users_predictions(user_id, n, model):
recommended_items = pd.DataFrame(model.loc[user_id])
recommended_items.columns = ["predicted_rating"]
recommended_items = recommended_items.sort_values('predicted_rating', ascending=False)
recommended_items = recommended_items.head(n)
return recommended_items.index.tolist()
def get_recs(model, k):
recs = []
for user in model.index:
cf_predictions = get_users_predictions(user, k, model)
recs.append(cf_predictions)
return recs
diversity_knn = []
novelty_knn = []
coverage_knn = []
f1_score_knn = []
accuracy_knn = []
entropy_knn = []
# Top-n recommendations for each user
no_of_recommendations = [5, 10, 15, 20, 30, 40, 50, 60, 70, 80, 90, 100]
for k in no_of_recommendations:
recs = get_recs(cf_model, k)
pred_user[f'Top-{k} Recommendation'] = recs
flat_list_recommendations = [item for sublist in recs for item in sublist]
product_counts = Counter(flat_list_recommendations)
# To calculate entropy
entropy_score = get_s_entropy(recs, product_counts, set(flat_list_recommendations))
print("The entropy score for {} recommendation is {} \n".format(k, entropy_score))
entropy_knn.append(entropy_score)
# To calculate the f1_score
f1_scores_knn, average_f1_score = cal_f1(test ,k)
print("The f1 score for {} recommendation is {} \n".format(k, average_f1_score))
f1_score_knn.append(average_f1_score)
# To calculate accuracy
accuracy_scores_knn, average_accuracy_knn = cal_accuracy(test.copy() ,k)
print("The accuracy score for {} recommendation is {} \n".format(k, average_accuracy_knn))
accuracy_knn.append(average_accuracy_knn)
# To calculate the diversity
diversity_scores_knn, average_diversity_knn = personalization(list(recs))
#diversity = get_shannon_entropy(recs, list(product_list), k)
print("The diversity score for {} recommendation is {} \n".format(k, average_diversity_knn))
diversity_knn.append(average_diversity_knn)
# To calculate the novelty
cf_novelty_knn, novelty_list_knn = recmetrics.novelty(list(recs), productId_counts, len(userId_counts), k)
print("The novelty score for {} recommendation is {} \n".format(k, cf_novelty_knn))
novelty_knn.append(cf_novelty_knn)
# To calculate the coverage
cf_coverage_knn = recmetrics.catalog_coverage(list(recs), product_list, 100)
print("The coverage score for {} recommendation is {} \n".format(k, cf_coverage_knn))
coverage_knn.append(cf_coverage_knn)
# To calculate the customer satisfaction
edt_knn = get_customer_satisfaction(test, k)
print("The cusotmer satisfaction for {} recommendation is {}".format(k, np.mean(list(edt_knn.values()))))
filename = "/home/ebcffhh/thesis/sorted_knn/metrics_knn_%s_recommendations.csv" % k
with open(filename, 'w') as csv_file:
writer = csv.writer(csv_file)
writer.writerow(['userId', 'accuracy', 'f1_score', 'diversity', 'novelty', 'customer_satisfaction'])
for i, nov, (uid, acc_score), (_, f1_score) in zip(diversity_scores_knn, novelty_list_knn, accuracy_scores_knn.items(), f1_scores_knn.items()):
if uid in sorted_nicely(edt_knn.keys()):
writer.writerow([uid, acc_score, f1_score, (1 - i[0]), nov, edt_knn.get(uid)])
# +
from sklearn.preprocessing import LabelEncoder
from keras.preprocessing.text import one_hot
from sklearn.model_selection import train_test_split
train_data, test_data = train_test_split(new_dataset, test_size = 0.2)
user_encoder = LabelEncoder()
product_encoder = LabelEncoder()
train_user_ids = np.array([one_hot(d,10) for d in train_data['userId']])
train_product_ids = np.array([one_hot(d,10) for d in train_data['ProductId']])
test_product_ids = np.array([one_hot(d,10) for d in test_data['ProductId']])
test_user_ids = np.array([one_hot(d,10) for d in test_data['userId']])
num_users= train_user_ids.max()+1
num_products = train_product_ids.max() + 1
print(num_users)
# +
from keras.layers.normalization.batch_normalization import BatchNormalization
import tensorflow as tf
from keras.layers import Input, Embedding, Flatten, Dense, Concatenate, dot
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn.model_selection import train_test_split
from keras.models import Model, Sequential
from keras.layers import Input, Embedding, Flatten, Dense, Concatenate, dot, Multiply, Dropout
from keras.preprocessing.text import one_hot,Tokenizer
import keras.layers
from keras.optimizers import adam_v2
from IPython.display import SVG
from keras.utils.vis_utils import model_to_dot
import tensorflow as tf
def get_ncf_model(no_of_factors):
product_input = Input(shape = [1], name = "Product-Input")
user_input = Input(shape = [1], name = "User-Input")
# Product embedding for GMF
gmf_product_embedding = Embedding(n_products, no_of_factors, name= "GMF-Product-Embedding", embeddings_initializer="he_normal")(product_input)
# User embedding for GMF
gmf_user_embedding = Embedding(n_users, no_of_factors, name = "GMF-User-Embedding", embeddings_initializer="he_normal")(user_input)
# GMF layers
gmf_product_vec = Flatten(name = "GMF-Flatten-Products")(gmf_product_embedding)
gmf_user_vec = Flatten(name = "GMF-Flatten-Users")(gmf_user_embedding)
gmf_output = Multiply()([gmf_user_vec, gmf_product_vec])
# Product embedding for MLP
mlp_product_embedding = Embedding(n_products, no_of_factors, name= "MLP-Product-Embedding", embeddings_initializer="he_normal")(product_input)
# User embedding for MLP
mlp_user_embedding = Embedding(n_users, no_of_factors, name = "MLP-User-Embedding", embeddings_initializer="he_normal")(user_input)
# MLP layers
mlp_product_vec = Flatten(name = "MLP-Flatten-Products")(mlp_product_embedding)
mlp_user_vec = Flatten(name = "MLP-Flatten-Users")(mlp_user_embedding)
#Concatenate features
conc = Concatenate()([mlp_product_vec, mlp_user_vec])
fc1 = Dropout(0.2)(conc)
fc2 = Dense(64, activation='relu')(fc1)
fc3 = BatchNormalization()(fc2)
fc4 = Dropout(0.2)(fc3)
fc5 = Dense(32, activation='relu')(fc4)
fc6 = BatchNormalization()(fc5)
fc7 = Dropout(0.2)(fc6)
fc8 = Dense(16, activation='relu')(fc7)
fc9 = BatchNormalization()(fc8)
fc10 = Dropout(0.2)(fc9)
fc11 = Dense(8, activation='relu')(fc10)
final_conc = Concatenate()([gmf_output, fc11])
output = Dense(1, activation='relu')(final_conc)
#Create model and compile it
opt = keras.optimizers.adam_v2.Adam(learning_rate=0.001)
model = Model([user_input, product_input], output)
model.compile(loss='mean_absolute_error', optimizer=opt, metrics=['accuracy'] )
#model = Model([user_input, product_input], output)
#model.compile('adam', 'mean_absolute_error')
return model
# +
from IPython.display import SVG
model = get_ncf_model(10)
SVG(model_to_dot( model, show_shapes=True, show_layer_names=True).create(prog='dot', format='svg'))
# +
from sklearn.metrics import mean_absolute_error
no_of_factors = [5, 10, 15, 20, 30, 40, 50, 60, 70, 80, 90, 100]
mae_ncf = list()
for k in no_of_factors:
model = get_ncf_model(k)
model.fit([train_user_ids, train_product_ids], train_data['Ratings'], epochs=3)
prediction = model.predict([test_user_ids, test_product_ids])
mae_ncf.append(mean_absolute_error(test_data['Ratings'], prediction))
print("Mean Absolute Error for value k {} is ".format(k), mean_absolute_error(test_data['Ratings'], prediction))
# +
model = get_ncf_model(40)
model.fit([train_user_ids, train_product_ids], train_data['Ratings'], epochs=3)
prediction = model.predict([test_user_ids, test_product_ids])
predicted_df = pd.DataFrame({'userId': test_data['userId'], 'productId': test_data['ProductId'], 'actual': test_data['Ratings']})
predicted_df['prediction'] = prediction
pred_user = predicted_df.copy().groupby('userId', as_index=False)['productId'].agg({'actual': (lambda x: list(set(x)))})
pred_user = pred_user.set_index("userId")
cf_model = predicted_df.pivot_table(index='userId',
columns='productId', values='prediction').fillna(0)
userId_counts = test_data['ProductId'].value_counts()
product_list = train_data.ProductId.unique().tolist()
productId_counts = dict(new_dataset.ProductId.value_counts())
def get_users_predictions(user_id, n, model):
recommended_items = pd.DataFrame(model.loc[user_id])
recommended_items.columns = ["prediction"]
recommended_items = recommended_items.sort_values('prediction', ascending=False)
recommended_items = recommended_items.head(n)
return recommended_items.index.tolist()
def get_recs(model, k):
recs = []
for user in model.index:
cf_predictions = get_users_predictions(user, k, model)
recs.append(cf_predictions)
return recs
diversity_nn = []
novelty_nn = []
coverage_nn = []
f1_score_nn = []
accuracy_nn = []
entropy_nn = []
# Top-n recommendations for each user
no_of_recommendations = [5, 10, 15, 20, 30, 40, 50, 60, 70, 80, 90, 100]
for k in no_of_recommendations:
recs = get_recs(cf_model, k)
preds = pd.DataFrame(index=cf_model.index)
preds[f'Top-{k} Recommendation'] = recs
flat_list_recommendations = [item for sublist in recs for item in sublist]
product_counts = Counter(flat_list_recommendations)
# To calculate entropy
entropy_score = get_s_entropy(recs, product_counts, set(flat_list_recommendations))
print("The entropy score for {} recommendation is {} \n".format(k, entropy_score))
entropy_nn.append(entropy_score)
# To calculate the f1_score
f1_scores, f1_score = cal_f1(predicted_df ,k)
print("The f1 score for {} recommendation is {}".format(k, f1_score))
f1_score_nn.append(f1_score)
# To calculate accuracy
accuracy_scores, accuracy = cal_accuracy(predicted_df ,k)
print("The accuracy score for {} recommendation is {} \n".format(k, accuracy))
accuracy_nn.append(accuracy)
# To calculate the diversity
diversity_scores, diversity = personalization(list(recs))
#diversity = get_shannon_entropy(recs, list(product_list), k)
print("The diversity score for {} recommendation is {} \n".format(k, diversity))
diversity_nn.append(diversity)
# To calculate the novelty
cf_novelty, novelty_list = recmetrics.novelty(list(recs), productId_counts, len(userId_counts), k)
print("The novelty score for {} recommendation is {} \n".format(k, cf_novelty))
novelty_nn.append(cf_novelty)
# To calculate the coverage
cf_coverage = recmetrics.catalog_coverage(list(recs), product_list, 100)
print("The coverage score for {} recommendation is {} \n".format(k, cf_coverage))
coverage_nn.append(cf_coverage)
# To calculate the customer satisfaction
edt = get_customer_satisfaction(predicted_df, k)
print("The cusotmer satisfaction for {} recommendation is {}".format(k, np.mean(list(edt.values()))))
filename = "/home/ebcffhh/thesis/dnn/metrics_dnn_%s_recommendations.csv" % k
with open(filename, 'w') as csv_file:
writer = csv.writer(csv_file)
writer.writerow(['userId', 'accuracy', 'f1_score', 'diversity', 'novelty', 'customer_satisfaction'])
for i, nov, (uid, acc_score), (_, f1_score) in zip(diversity_scores, novelty_list, accuracy_scores.items(), f1_scores.items()):
if uid in sorted(edt.keys()):
writer.writerow([uid, acc_score, f1_score, (1 - i[0]), nov, edt.get(uid)])
# +
k = [5, 10, 15, 20, 30, 40, 50, 60, 70, 80, 90, 100]
plt.xlabel('No of factors')
plt.ylabel('Mean Absolute Error')
ax=plt.gca()
ax.locator_params('y', nbins=10)
plt.locator_params('x', nbins=20)
plt.plot(k, mae_knn, label = "KNN")
plt.plot(k, mae_svd, label = "SVD")
plt.plot(k, mae_ncf, label = "DNN")
plt.scatter(k,mae_knn,s=50,color='red',zorder=2)
plt.scatter(k,mae_svd,s=50,color='green',zorder=2)
plt.scatter(k,mae_ncf,s=50,color='brown',zorder=2)
plt.legend()
plt.show()
print("mae")
print("mae_svd",mae_svd)
print("mae_knn",mae_knn)
print("mae_ncf",mae_ncf)
plt.xlabel('No of Recommendations')
plt.ylabel('Mean Absolute Error')
ax=plt.gca()
ax.locator_params('y', nbins=10)
plt.locator_params('x', nbins=20)
plt.plot(k, accuracy_knn, label = "KNN")
plt.plot(k, accuracy_svd, label = "SVD")
plt.plot(k, accuracy_nn, label = "DNN")
plt.scatter(k,accuracy_knn,s=50,color='red',zorder=2)
plt.scatter(k,accuracy_svd,s=50,color='green',zorder=2)
plt.scatter(k,accuracy_nn,s=50,color='brown',zorder=2)
plt.legend()
plt.show()
print("accuracy")
print("accuracy_svd",accuracy_svd)
print("accuracy_knn",accuracy_knn)
print("accuracy_dnn",accuracy_nn)
plt.xlabel('No of Recommendations')
plt.ylabel('f1_Score')
ax=plt.gca()
ax.locator_params('y', nbins=15)
plt.locator_params('x', nbins=20)
plt.plot(k, f1_score_knn, label = "KNN")
plt.plot(k, f1_score_svd, label = "SVD")
plt.plot(k, f1_score_nn, label = "DNN")
plt.scatter(k,f1_score_knn,s=50,color='red',zorder=2)
plt.scatter(k,f1_score_svd,s=50,color='green',zorder=2)
plt.scatter(k,f1_score_nn,s=50,color='brown',zorder=2)
plt.legend()
plt.show()
print("f1_Score")
print("f1_Score_nn", f1_score_nn)
print("f1_Score_knn", f1_score_knn)
print("f1_Score_svd", f1_score_svd)
print("entropy")
print("entropy_svd",entropy_svd)
print("entropy_knn",entropy_knn)
print("entropy_dnn",entropy_nn)
plt.xlabel('No of Recommendations')
plt.ylabel('Diversity')
ax=plt.gca()
ax.locator_params('y', nbins=25)
plt.locator_params('x', nbins=20)
for i, txt in enumerate(entropy_knn):
ax.annotate(round(txt, 5), (k[i],entropy_knn[i]), fontsize=8)
plt.plot(k, entropy_knn, label = "KNN")
plt.scatter(k,entropy_knn,s=20,color='red',zorder=2)
plt.legend()
plt.show()
plt.xlabel('No of Recommendations')
plt.ylabel('Diversity')
ax=plt.gca()
ax.locator_params('y', nbins=25)
plt.locator_params('x', nbins=20)
for i, txt in enumerate(entropy_svd):
ax.annotate(round(txt, 5), (k[i],entropy_svd[i]), fontsize=8)
plt.plot(k, entropy_svd, label = "SVD")
plt.scatter(k,entropy_svd,s=20,color='red',zorder=2)
plt.legend()
plt.show()
plt.xlabel('No of Recommendations')
plt.ylabel('Diversity')
ax=plt.gca()
ax.locator_params('y', nbins=25)
plt.locator_params('x', nbins=20)
for i, txt in enumerate(entropy_nn):
ax.annotate(round(txt, 5), (k[i],entropy_nn[i]), fontsize=8)
plt.plot(k, entropy_nn, label = "DNN")
plt.scatter(k,entropy_nn,s=20,color='brown',zorder=1)
plt.legend()
plt.show()
plt.xlabel('No of Recommendations')
plt.ylabel('Diversity')
ax=plt.gca()
ax.locator_params('y', nbins=15)
plt.locator_params('x', nbins=20)
plt.plot(k, entropy_knn, label = "KNN")
plt.plot(k, entropy_svd, label = "SVD")
plt.plot(k, entropy_nn, label = "DNN")
plt.scatter(k,entropy_knn,s=50,color='blue',zorder=2)
plt.scatter(k,entropy_svd,s=50,color='yellow',zorder=2)
plt.scatter(k,entropy_nn,s=50,color='brown',zorder=2)
plt.legend()
plt.show()
print("f1_Score")
print("f1_Score_nn", f1_score_nn)
print("f1_Score_knn", f1_score_knn)
print("f1_Score_svd", f1_score_svd)
plt.xlabel('No of Recommendations')
plt.ylabel('Diversity')
ax=plt.gca()
ax.locator_params('y', nbins=25)
plt.locator_params('x', nbins=20)
for i, txt in enumerate(diversity_knn):
ax.annotate(round(txt, 5), (k[i],diversity_knn[i]), fontsize=8)
plt.plot(k, diversity_knn, label = "KNN")
plt.scatter(k,diversity_knn,s=20,color='red',zorder=2)
plt.legend()
plt.show()
plt.xlabel('No of Recommendations')
plt.ylabel('Diversity')
ax=plt.gca()
ax.locator_params('y', nbins=25)
plt.locator_params('x', nbins=20)
for i, txt in enumerate(diversity_svd):
ax.annotate(round(txt, 5), (k[i],diversity_svd[i]), fontsize=8)
plt.plot(k, diversity_svd, label = "SVD")
plt.scatter(k,diversity_svd,s=20,color='red',zorder=2)
plt.legend()
plt.show()
plt.xlabel('No of Recommendations')
plt.ylabel('Diversity')
ax=plt.gca()
ax.locator_params('y', nbins=25)
plt.locator_params('x', nbins=20)
for i, txt in enumerate(diversity_nn):
ax.annotate(round(txt, 5), (k[i],diversity_nn[i]), fontsize=8)
plt.plot(k, diversity_nn, label = "DNN")
plt.scatter(k,diversity_nn,s=20,color='red',zorder=2)
plt.legend()
plt.show()
print("diversity")
print("divesity_knn", diversity_knn)
print("diversity_nn",diversity_nn)
print("diversity_svd",diversity_svd)
plt.xlabel('No of Recommendations')
plt.ylabel('Novelty')
ax=plt.gca()
ax.locator_params('y', nbins=15)
plt.locator_params('x', nbins=20)
plt.plot(k, novelty_knn, label = "KNN")
plt.plot(k, novelty_svd, label = "SVD")
plt.plot(k, novelty_nn, label = "DNN")
plt.scatter(k,novelty_knn,s=50,color='red',zorder=2)
plt.scatter(k,novelty_svd,s=50,color='green',zorder=2)
plt.scatter(k,novelty_nn,s=50,color='black',zorder=2)
plt.legend()
plt.show()
print("novelty")
print("novelty_knn", novelty_knn)
print("novelty_nn",novelty_nn)
print("novelty_svd",novelty_svd)
plt.xlabel('No of Recommendations')
plt.ylabel('Coverage')
ax=plt.gca()
ax.locator_params('y', nbins=15)
plt.locator_params('x', nbins=20)
plt.plot(k, coverage_knn, label = "KNN")
plt.plot(k, coverage_svd, label = "SVD")
plt.plot(k, coverage_nn, label = "DNN")
plt.scatter(k,coverage_knn,s=50,color='blue',zorder=2)
plt.scatter(k,coverage_svd,s=50,color='grey',zorder=2)
plt.scatter(k,coverage_nn,s=50,color='orange',zorder=2)
plt.legend()
plt.show()
| recommendation_thesis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # A report on sales forecasting
# ### Kaggle project
# ## Synopsis
# Sales forecasting is a common problem that can be easily? managed using Machine Learning techniques.
# In this report, sales records made by the Russian software developer, publisher and distributor company *1C Company* have been analysed. Based on past sales, the amount of items that will be sold in a certain shop is predicted.
# The forecast accuracy is XXX% with an error of XXXX.
# ## Data
#
| scripts/.ipynb_checkpoints/repor_1C_v0-checkpoint.ipynb |
/ -*- coding: utf-8 -*-
/ ---
/ jupyter:
/ jupytext:
/ text_representation:
/ extension: .q
/ format_name: light
/ format_version: '1.5'
/ jupytext_version: 1.14.4
/ kernelspec:
/ display_name: Coq
/ language: coq
/ name: coq
/ ---
/ + [markdown] coq_kernel_metadata={"auto_roll_back": true}
/ https://riptutorial.com/it/coq/topic/4007/ricerca-di-un-fatto-esistente-con-ricerca-e-varianti
/ + [markdown] coq_kernel_metadata={"auto_roll_back": true, "cell_id": "be8c834a5e5847eab667a33836c99147", "evaluated": false, "execution_id": "f7653fb90a2840839e3d111eb9ae335d", "rolled_back": false}
/ # Dimostrazione più semplice
/ + coq_kernel_metadata={"auto_roll_back": true, "cell_id": "454809942afa421282ec62bda576708d", "evaluated": true, "execution_id": "1d9ca13b86b0419f81cdd1d7f29aca32", "rolled_back": false}
Theorem my_first_theorem : 1 + 1 = 2.
Proof.
reflexivity.
Qed.
/ + [markdown] coq_kernel_metadata={"auto_roll_back": true}
/ # Prova per induzione
/ + coq_kernel_metadata={"auto_roll_back": true, "cell_id": "016122c5c10440e8bb01cd099433bf23", "evaluated": true, "execution_id": "7c918d1bed904041852f6185e7b4310f", "rolled_back": false}
Require Import Coq.Setoids.Setoid.
Require Import Coq.Arith.Lt.
/ + [markdown] coq_kernel_metadata={"auto_roll_back": true}
/ A number is less than or equal to itself
/ + coq_kernel_metadata={"auto_roll_back": true, "cell_id": "3b38e3fa818c483991fbac1c6be97150", "evaluated": true, "execution_id": "c68e99d911ad4b8da1b1d2ab0e0ab74f", "rolled_back": false}
Theorem aLTEa : forall a, a <= a.
Proof.
auto with arith. (* This follows by simple arithmetic *)
Qed.
/ + coq_kernel_metadata={"auto_roll_back": true, "cell_id": "20fffb8d583d41ca86d348d00a071af5", "evaluated": true, "execution_id": "3fa05f1d86854ec7a14e1757664fa571", "rolled_back": false}
Theorem simplALTE : forall a b,
S a <= S b <-> a <= b. (* If a <= b, then a + 1 <= b + 1 *)
Proof.
Admitted.
/ + coq_kernel_metadata={"auto_roll_back": true, "cell_id": "33be13450bb041abad6f740b15115347", "evaluated": true, "execution_id": "374e969db1d6429b82dec415dc693843", "rolled_back": false}
Theorem ltAlwaysLt: forall a b,
a <= a + b.
Proof.
intros. (* Introduce relevant variables *)
induction a, b. (* Induction on every variable *)
simpl. apply aLTEa. (* 0 <= 0 + S b *)
rewrite -> plus_O_n. auto with arith. (* 0 <= S b *)
rewrite <- plus_n_O. apply aLTEa. (* S a <= S a + 0 *)
rewrite <- simplALTE in IHa. (* IHa: a <= a + S b. Goal: S a <= S a + S b. *)
apply IHa. (* We rewrote the induction hypothesis to be in the same form as the goal, so it applies immediately now *)
Qed.
/ + [markdown] coq_kernel_metadata={"auto_roll_back": true}
/ # Esempio banale di un'analisi del caso
/ + coq_kernel_metadata={"auto_roll_back": true, "cell_id": "3ded1ab731d34547a238e4ccd1b34c54", "evaluated": true, "execution_id": "7cb8b6bad7bc45688f8796c86f5d5ffc", "rolled_back": false}
Require Import Coq.Arith.Lt.
Theorem atLeastZero : forall a,
0 <= a.
Proof.
intros.
destruct a. (* Case analysis *)
reflexivity. (* 0 >= 0 *)
apply le_0_n. (* S a is always greater than zero *)
Qed.
| notebook/GetStarted.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# coding: utf-8
import sys, os
sys.path.append(os.pardir) # 부모 디렉터리의 파일을 가져올 수 있도록
import numpy as np
import pickle
from dataset.mnist import load_mnist
from common.functions import sigmoid, softmax
def get_data():
(x_train, t_train), (x_test, t_test) = load_mnist(normalize=True, flatten=True, one_hot_label=False)
return x_test, t_test
def init_network():
with open("sample_weight.pkl", 'rb') as f:
network = pickle.load(f)
return network
def predict(network, x):
w1, w2, w3 = network['W1'], network['W2'], network['W3']
b1, b2, b3 = network['b1'], network['b2'], network['b3']
a1 = np.dot(x, w1) + b1
z1 = sigmoid(a1)
a2 = np.dot(z1, w2) + b2
z2 = sigmoid(a2)
a3 = np.dot(z2, w3) + b3
y = softmax(a3)
return y
x, t = get_data()
network = init_network()
batch_size = 100 # 배치 크기
accuracy_cnt = 0
for i in range(0, len(x), batch_size):
x_batch = x[i:i+batch_size]
y_batch = predict(network, x_batch)
p = np.argmax(y_batch, axis=1)
accuracy_cnt += np.sum(p == t[i:i+batch_size])
print("Accuracy:" + str(float(accuracy_cnt) / len(x)))
# +
# Result
| 04_Neural_Network_Basics/neuralnet_mnist_batch.ipynb |
# -*- coding: utf-8 -*-
LoadPackage("FrancyMonoids");
# ## Trees
s:=NumericalSemigroup(1);
# ## `DrawTreeOfSonsOfNumericalSemigroup(s,l,gensfunc)`
#
# This function draws the thre of the sons of `s` up to level `l` with respect to `gensfunc` (a function that gets minimal generators according to a given variety of numerical semigroups, for instance, `MinimalGenerators`)
DrawTreeOfSonsOfNumericalSemigroup(s,5,MinimalGenerators);
s:=NumericalSemigroup(4,6,9);
# ## `DrawTreeOfGluingsOfNumericalSemigroup(s,expand...)`
#
# Draws the decomposition of `s` as a gluing of numerical semigroups, and then proceeds recursively with each of the factors. Decomposition is not unique, and this is why a tree is drawn. The optional argument `expand` may be true or false, and it makes the tree fully expandable or not.
DrawTreeOfGluingsOfNumericalSemigroup(s,true);
# ## Hasse Diagrams
s:=NumericalSemigroup(3,5,7);
# ## `DrawHasseDiagramOfNumericalSemigroup(s, A)`
#
# Draws the Hasse diagram of `A` (a set of integers) with respect to the ordering induced by `s`: $a\preceq b$ if $b-a$ is in `s`.
DrawHasseDiagramOfNumericalSemigroup(s,[1..20]);
# Of special interest is the Hasse diagram of an Apéry set or of the Betti elements.
s:=NumericalSemigroup(5,7,9,11);
DrawHasseDiagramOfNumericalSemigroup(s,AperyList(s,10));
# ## `DrawOverSemigroupsNumericalSemigroup(s)`
#
# Draws the Hasse diagram of oversemigroups of the numerical semigroup `s`.
s:=NumericalSemigroup(5,7,11,13);
DrawOverSemigroupsNumericalSemigroup(s);
# ## Graphs
s:=NumericalSemigroup(3,5,7);
f:=FactorizationsElementWRTNumericalSemigroup(30,s);
# ## `DrawFactorizationGraph(f)`
#
# Draws the factorization graph of a set of factorizations `f`, the complete graph with vertices the elements of `f`. Labels are labeled with distances between the nodes they join. It also draws a minimal spanning tree (with minimal distances) and thus one can read the Catenary degree from this tree.
DrawFactorizationGraph(f);
# ## `DrawEliahouGraph(f)`
#
# Draws the Eliahou's graph associated to the set of factorizations `f`. The vertices of the graph are the elements in `f`, and two vertices are joined with an edge if they have common support.
DrawEliahouGraph(f);
# ## `DrawRosalesGraph(n,s)`
#
# Draws the Rosales graph associated to `n` in `s`. The semigroup `s` can be either a numerical or an affine semigroup. The vertices of the graph are the minimal generators `a` of `s` such that `n-a` is in `s`, and edges are pairs `ab` such that `n-(a+b)` is in `s`.
DrawRosalesGraph(10,s);
BettiElements(s);
| notebooks/francy-monoids.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from scipy.linalg import lu_factor, lu_solve
def LU(A, P):
lu, piv = lu_factor(A)
solucao = lu_solve((lu, piv), P)
return solucao
# +
A = np.array([[1, 1, -1],
[1, 1, 4],
[2, -1, 2]], dtype='float16')
P = np.array([[1],
[2],
[3]], dtype='float16')
#Por LU:
s = LU(A, P)
print(f'O vetor solução pelo metodo da decomposição LU é: \n{s}')
# -
| Lab4/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from numba import jit, float64, int64
from numba.experimental import jitclass
import matplotlib.pyplot as plt
# %matplotlib inline
np.set_printoptions(precision=16, suppress=True)
from naginterfaces.library import machine, roots
from naginterfaces.base.utils import NagAlgorithmicWarning
eps = machine.precision()
from numba.core.errors import NumbaDeprecationWarning, NumbaPendingDeprecationWarning, NumbaWarning
import warnings
warnings.simplefilter('ignore', category=NumbaDeprecationWarning)
warnings.simplefilter('ignore', category=NumbaPendingDeprecationWarning)
warnings.simplefilter('ignore', category=NumbaWarning)
warnings.simplefilter('ignore', category=NagAlgorithmicWarning)
# -
# # Using Anderson Acceleration to speed up fixed-point iterations
#
#
# ## Accelerating the fixed point solution of cos(x) = x
#
# Type any real number into a calculator and repeatedly press the cosine button. The result eventually converges to about 0.739085 which is the solution of the equation $x = \cos(x)$. This is an example of a fixed-point iteration. The general case is any computation that looks like this
#
# \begin{align}
# x_{n+1} = f(x_n)
# \end{align}
#
# which is repeated until a convergence criterion is reached. Such computations arise naturally in many areas of finance, physics and engineering.
#
# We could code the cosine fixed-point computation in Python as follows
# +
def fcos(x):
"""Performs one iteration of the cosine fixed point iteration
"""
return np.cos(x)
def fp_driver(x0, func, tol):
x = x0 # starting guess
err = 1000000 # Huge error to start things off
iterations = 0
while err > tol:
nextx = func(x)
err = np.sqrt((x - nextx)**2)
x = nextx
iterations += 1
return iterations, x
fp_driver(3, fcos, 1e-15)
# -
# Demonstrating that we get convergence to a tolerance of 1e-15 in 88 iterations. A natural question to ask is 'Can we accelerate the convergence somehow?' and the answer is 'Yes', using a technique called [Anderson Acceleration](https://epubs.siam.org/doi/abs/10.1137/10078356X). Anderson Acceleration has been known in the chemistry community since the 1960s but until relatively recently it wasn't well known in other fields. The [2011 paper by <NAME> Ni](https://epubs.siam.org/doi/abs/10.1137/10078356X) describing the technique has been cited over 245 times as of late 2019.
#
# NAG recently introduced two Anderson Acceleration routines into the NAG Library following the success of the technique in the evaluation of Nearest correlation matrices (NCMs) (the accelerated nearest correlation matrix routine is implemented in `naginterfaces.library.correg.corrmat_fixed`). Here, we will demonstrate the use of NAG's standard Anderson Acceleration routine `roots.sys_func_aa`.
#
# `roots.sys_func_aa` takes the role of `fp_driver` in the code above. Here's how to use it for the cosine example
# +
def nag_fcos(x):
"""Performs one iteration of the cosine fixed point iteration
"""
# Note that NAG requires f(x) - x to be returned instead of just f(x)
return np.cos(x)-x
m = 0 # When m=0, Acceleration is turned off so this is identical to running a basic fixed-point calculation
tol = 1e-10
nag_cos = roots.sys_func_aa(nag_fcos, 3.0, tol, eps, m).x
nag_cos
# -
# We turn on acceleration by setting `m` to something more than 0.
nag_cos = roots.sys_func_aa(nag_fcos, 3.0, tol, eps, m=1).x
nag_cos
# Unfortunately, the NAG routine does not return the number of iterations required for convergence so we have no idea what the benefit of acceleration is unless we count them ourselves. We do this with a helper class which we pass to the `nag_fcos` function.
# +
class fpinfo:
"""A class used to get information to/from the NAG routine
"""
def __init__(self):
self.iterations = 0
def nag_fcos_withdata(x, data):
"""Performs one iteration of the cosine fixed point iteration
"""
data.iterations += 1
# Note that NAG requires f(x) - x to be returned instead of just f(x)
return np.cos(x)-x
tol = 1e-15
data = fpinfo()
nag_cos = roots.sys_func_aa(nag_fcos_withdata, 3.0, tol, eps, m=0, data=data).x
print(f"Solution found was {nag_cos[0]} in {data.iterations} iterations with acceleration switched off")
data = fpinfo()
nag_cos = roots.sys_func_aa(nag_fcos_withdata, 3.0, tol, eps, m=1, data=data).x
print(f"Solution found was {nag_cos[0]} in {data.iterations} iterations with acceleration switched on")
# -
# An almost 9x improvement in iteration count simply by switching on acceleration is extremely useful. We move on to consider a more challenging example of fixed-point iteration.
# ## Solving the Poisson Equation Using Fixed-Point Iterations
# [Poisson's equation](https://en.wikipedia.org/wiki/Poisson%27s_equation) is an elliptic Partial Differential Equation (PDE) which in two dimensions takes the form
#
# \begin{align}
# \nabla^2 u & = f(x,y)\nonumber \\
# \left( \frac{\partial^2}{\partial x^2} + \frac{\partial^2}{\partial y^2} \right)u(x,y) &= f(x,y)\nonumber
# \end{align}
#
# The special case where $f(x)=0$ is referred to as Laplace's equation. Both Poisson's and Laplace's equations arise in many areas of physics and engineering.
#
# One example which requires the numerical solution of Poisson's equation is finding the steady state of the potential field of two parallel lines of electric charge . One with total charge $+1$ and the other with total charge $-1$. The Dirichlet boundary condition $u(x,y)=0$ is applied at the domain boundary.
# +
def source(N):
# Produces a matrix corresponding to a source term
# Want two line sources with total charge of +/- 1 on each line
# This is our function f(x, y) in the poisson equation
x0 = np.zeros((N, N))
h = 1/(N-1)
source_rows = range((N//4), (3*N//4))
source_col1 = N//4
source_col2 = 3*N//4
x0[source_rows, source_col1] = x0[source_rows, source_col1] + 1/(N*0.5*h**2)
x0[source_rows, source_col2] = x0[source_rows, source_col2] - 1/(N*0.5*h**2)
return x0
def init_problem(N=50):
# Initialises an N x N problem domain with an initialised source
x0 = np.zeros((N, N))
x0 += source(N)
return x0
x0 = init_problem(50)
plt.title('Initial conditions of our example problem on a 50 x 50 grid')
_ = plt.imshow(x0)
# -
# Many introductory courses on the solution of PDEs will demonstrate three methods for the solution of systems such as this:
#
# * The Jacobi Method
# * The Gauss-Seidel Method
# * Successive Over-relaxation
#
# A paper that discusses these methods in depth using similar notation to this notebook is [Parallel S.O.R. iterative methods](https://www.sciencedirect.com/science/article/pii/S0167819184903806) by <NAME>.
# We now look at implementing the three solution methods in turn
#
# ### The Jacobi Method
#
# Starting from the <i>n</i>th configuration of the discretized grid $u^{n}$, we get the next configuration by applying the following formula to every $(i,j)$th grid point other than the very edges where we apply the boundary conditions.
#
# \begin{align}
# u^{n+1}_{j,i} = \frac{1}{4} \left(u^{n}_{j+1,i} +u^{n}_{j-1,i} +u^{n}_{j,i+1} +u^{n}_{j,i-1} \right)+\frac{h^2}{4}f_{j,i}
# \end{align}
#
# An implementation of this iteration in Python is
@jit
def jacobi(x, source, solverinfo):
"""Performs one iteration of the jacobi method
Arguments:
x - N x N Matrix containing the previous iteration of the solution
source - N x N Matrix containing our source function evaluated on the grid
solverinfo - Takes a solverinfo object to get info in/out of the solver
"""
N = x.shape[0]
nextx = np.zeros((N, N))
h = 1/(N-1)
# loop over the grid
# Only iterate over interior points thus keeping the edges untouched and hence enforcing
# the boundary condition that x = 0 at the edges.
for i in range(1, N - 1):
for j in range(1, N - 1):
nextx[j, i] = 0.25 * (x[j+1, i] + x[j, i+1] + x[j-1, i] + x[j, i-1]) + 0.25*h**2*source[j, i]
solverinfo.iterations += 1
return nextx
# Next we need a driver for the Jacobi iteration function. Something that is general enough to drive subsequent methods of solving our problem.
# +
# This spec is required for Numba jit compilation
spec = [
('iterations', int64),
('w', float64)
]
@jitclass(spec)
class solverinfo:
"""A class used to get information to/from a solver
"""
def __init__(self, _w=1):
self.iterations = 0
self.w = 1 # Used for SOR and ignored for everything else
def solve_poisson(x, tol, data, method):
"""Solves Poisson's equation
x - N x N Matrix containing the initial configuration of the problem space
tol - Convergence tolerance
data - A solverinfo object to get arbitrary info in/out of the solver
method - A function that performs one solution iteration
"""
N = int(np.sqrt(x.size))
err = 1 # Big error to start things off with
while err > tol:
nextx = method(x, source(N), data)
err = np.sqrt(np.sum((x - nextx)**2))
x = np.copy(nextx)
return x
# -
# Finally, we can set up and solve an instance of our problem using the Jacobi method.
N = 100
u = init_problem(N)
jacobi_info = solverinfo()
tol = 1e-9
jacobi_sol = solve_poisson(u, tol, jacobi_info, jacobi)
print(f"Gauss-Seidel on a {N} by {N} grid")
print(f"Solution found in {jacobi_info.iterations} iterations")
_ = plt.imshow(jacobi_sol)
# Poisson's equation gives us the potential. For completeness, let's find the electric field by taking the gradient
Ex, Ey = np.gradient(jacobi_sol)
E = np.sqrt(Ex**2+Ey**2) # Magnitude of Electric field
plt.imshow(E)
_ = plt.title('Electric Field')
# 28,734 iterations is a lot and we would hope that we could do better with more sophisticated techniques. Fortunately, we can and 'more sophisticated' isn't quite as sophisticated as you might fear.
#
# ### Gauss--Seidel Method
#
# In the Jacobi method, we kept two independent copies of the solution space. Each element of the **new** solution is computed from the surrounding elements of the **old** solution. In Gauss-Seidel we make use of new grid elements as soon as they become available. That is, when we reach position $(j,i)$ in the grid we have already updated $u_{j-1,i}$ and $u_{j,i-1}$ so we can use those instead of the old values.
#
# \begin{align}
# u^{n+1}_{j,i} = \frac{1}{4} \left(u^{n}_{j+1,i} +u^{n+1}_{j-1,i} +u^{n}_{j,i+1} +u^{n+1}_{j,i-1} \right)+\frac{h^2}{4}f_{j,i}
# \end{align}
#
# The Gauss-Seidel iteration, therefore, looks like this
@jit
def gauss_seidel(x, source, solverinfo):
N = x.shape[0]
nextx = np.copy(x)
h = 1/(N-1)
for i in range(1, N - 1):
for j in range(1, N - 1):
nextx[j, i] = 0.25 * (nextx[j+1, i] + nextx[j, i+1] + nextx[j-1, i] + nextx[j, i-1]) + 0.25*h**2*source[j, i]
solverinfo.iterations += 1
return nextx
# +
N = 100
u = init_problem(N)
gs_info = solverinfo()
tol = 1e-9
gs_sol = solve_poisson(u, tol, gs_info, gauss_seidel)
print(f"Gauss--Seidel on a {N} by {N} grid")
print(f"Solution found in {gs_info.iterations} iterations")
fig, axes = plt.subplots(nrows=1, ncols=2, figsize=(10, 6))
axes[0].imshow(gs_sol)
axes[0].set_title('Potential')
Ex, Ey = np.gradient(gs_sol)
E = np.sqrt(Ex**2+Ey**2) # Magnitude of Electric field
axes[1].imshow(E)
_ = axes[1].set_title('Electric Field')
# -
# The number of iterations has almost halved which is a fantastic return on such a small investment. The standard treatment of this problem has yet another trick up its sleeve with the Successive Over Relaxation method (SOR) which is an extension of the Gauss--Seidel method.
#
# ### Successive Over Relaxation Method (SOR)
#
# Take the Gauss--Seidel update formula above (3) and add and subtract $u_{i,j}^n$ to the right-hand side
#
# \begin{align}
# u^{n+1}_{j,i} & = \frac{1}{4} \left(u^{n}_{j+1,i} +u^{n+1}_{j-1,i} +u^{n}_{j,i+1} +u^{n+1}_{j,i-1} +h^2 f_{j,i} \right) \nonumber \\
# & = u_{i,j}^n + \frac{1}{4} \left(u^{n}_{j+1,i} +u^{n+1}_{j-1,i} +u^{n}_{j,i+1} +u^{n+1}_{j,i-1} + h^2 f_{j,i} - 4 u_{i,j}^n \right) \nonumber \\
# & = u_{i,j}^n + r_{i,j}
# \end{align}
#
# So $r_{i,j}$ is the amount of change of $u_{i,j}$ for one iteration of Gauss_Seidel. If this amount of change is good, maybe more of the same is better and it turns out that the convergence of Gauss--Seidel can be accelerated by making a larger change as follows
#
# \begin{align}
# u^{n+1}_{j,i} & = u_{i,j}^n + \omega r_{i,j}
# \end{align}
#
# where $\omega$ is positive constant called the acceleration factor that in practice lies between 1 and 2. Note that when $\omega = 1$ the SOR reduces to standard Gauss-Seidel.
#
# The Python looks like this
@jit
def SOR(x, source, solverinfo):
N = x.shape[0]
nextx = np.copy(x)
h = 1/(N-1)
w = solverinfo.w
for i in range(1, N - 1):
for j in range(1, N - 1):
new = 0.25 * (nextx[j-1, i] + nextx[j+1, i] + nextx[j, i-1]+ nextx[j, i+1]) + 0.25*h**2*source[j, i]
nextx[j, i] += w * (new - nextx[j, i])
solverinfo.iterations += 1
return nextx
# Using this to solve our problem with $\omega=1.94$ we have
# +
N = 100
u = init_problem(N)
SOR_info = solverinfo()
SOR_info.w = 1.94
tol = 1e-9
SOR_sol = solve_poisson(u, tol, SOR_info, SOR)
print(f"SOR on a {N} by {N} grid")
print(f"Solution found in {SOR_info.iterations} iterations")
fig, axes = plt.subplots(nrows=1, ncols=2, figsize=(10, 6))
axes[0].imshow(SOR_sol)
axes[0].set_title('Potential')
Ex, Ey = np.gradient(SOR_sol)
E = np.sqrt(Ex**2+Ey**2) # Magnitude of Electric field
axes[1].imshow(E)
_ = axes[1].set_title('Electric Field')
# -
# 486 iterations is over 59x faster than our first attempt using the Jacobi method which needed 28,734 iterations to converge.
#
# Choose the SOR $\omega$ parameter badly, however, and you can get performance that's more than 10x slower than when we chose well! It's still several times faster than Jacobi or Gauss--Seidel however.
# +
N = 100
u = init_problem(N)
SOR_info = solverinfo()
SOR_info.w = 1.5
tol = 1e-9
SOR_sol = solve_poisson(u, tol, SOR_info, SOR)
print(f"SOR on a {N} by {N} grid")
print(f"Solution found in {SOR_info.iterations} iterations")
fig, axes = plt.subplots(nrows=1, ncols=2, figsize=(10, 6))
axes[0].imshow(SOR_sol)
axes[0].set_title('Potential')
Ex, Ey = np.gradient(SOR_sol)
E = np.sqrt(Ex**2+Ey**2) # Magnitude of Electric field
axes[1].imshow(E)
_ = axes[1].set_title('Electric Field')
# -
# There are analytic methods available to estimate the optimum $\omega$ but it's often necessary to find it empirically. The optimum value of $\omega$ depends on both the PDE being solved and on the grid resolution. More advanced extensions of SOR exist that have even better convergence properties (Chebyshev acceleration for example). I could also consider parallelisation schemes for these methods such as Red-Black ordering but I need to stop somewhere if I am going to get onto Anderson Acceleration before the deadline of this article!
#
# Before moving on, let's look at the number of iterations taken by SOR for this problem as a function of $\omega$
# +
def SOR_iterations(w):
N = 100
u = init_problem(N)
SOR_info = solverinfo()
SOR_info.w = w
tol = 1e-9
_ = solve_poisson(u, tol, SOR_info, SOR)
return SOR_info.iterations
w = np.arange(1.0, 1.99, 0.01)
iterations = [SOR_iterations(omega) for omega in w]
plt.plot(w, iterations)
plt.title('Number of iterations vs SOR parameter')
plt.xlabel('w')
_ = plt.ylabel('Iterations')
# -
# ## Applying NAG's Anderson Acceleration to these methods
#
# ### Code adaptations required to call Jacobi iterations from NAG
#
# I started on this journey because I noted that all of these methods are fixed-point and wondered if NAG's Anderson acceleration would be useful. Let's go back to Jacobi and recall the definition of each iteration
@jit
def jacobi(x, source, solverinfo): # pylint: disable=function-redefined
"""Performs one iteration of the jacobi method
Arguments:
x - N x N Matrix containing the previous iteration of the solution
source - N x N Matrix containing our source function evaluated on the grid
solverinfo - Takes a solverinfo object to get info in/out of the solver
"""
N = x.shape[0]
nextx = np.zeros((N, N))
h = 1/(N-1)
# loop over the grid
# Only iterate over interior points thus keeping the edges untouched and hence enforcing
# the boundary condition that x = 0 at the edges.
for i in range(1, N - 1):
for j in range(1, N - 1):
nextx[j, i] = 0.25 * (x[j+1, i] + x[j, i+1] + x[j-1, i] + x[j, i-1]) + 0.25*h**2*source[j, i]
solverinfo.iterations += 1
return nextx
# The NAG routine that performs Anderson acceleration is `roots.sys_func_aa` and the [documentation for it](https://www.nag.com/numeric/py/nagdoc_latest/naginterfaces.library.roots.html) tells us that the function to be solved must have the form `fcn(x, data)` where `data` is any arbitrary Python object that can be used to get information in and out of `fcn`. As such, we are going to need to change the `solverinfo` object so that it also holds the `source` term.
#
# Two other constraints are going to change the way this function looks:
#
# * The vector $x$ needs to be one-dimensional. As such, we'll need to change the shape to and from the two-dimensional representation we've been using until now.
# * We need to return `jacobi(x) - x` instead of `jacobi(x)`
#
# With all of this taken into account, the NAG-ified version of `jacobi` and the supporting `solverinfo` class is
# +
# This spec is required for Numba jit compilation
spec = [
('iterations', int64),
('w', float64),
('source', float64[:, :])
]
@jitclass(spec)
class NAG_solverinfo:
"""A class used to get information to/from a solver
"""
def __init__(self, N=50, _w=1):
self.iterations = 0
self.w = 1 # Used for SOR and ignored for everything else
self.source = np.zeros((N, N))
# -
@jit
def NAG_jacobi(x, solverinfo):
"""Performs one iteration of the jacobi method in the format required by NAG
Arguments:
x - N x N Matrix containing the previous iteration of the solution
source - N x N Matrix containing our source function evaluated on the grid
solverinfo - Takes a solverinfo object to get info in/out of the solver
"""
N = int(np.sqrt(x.size))
x.shape = (N, N) # Make x 2D because that's how I think
nextx = np.zeros((N, N))
h = 1/(N-1)
# loop over the grid
# Only iterate over interior points thus keeping the edges untouched and hence enforcing
# the boundary condition that x = 0 at the edges.
for i in range(1, N - 1):
for j in range(1, N - 1):
nextx[j, i] = 0.25 * (x[j+1, i] + x[j, i+1] + x[j-1, i] + x[j, i-1]) + 0.25*h**2*solverinfo.source[j, i]
solverinfo.iterations += 1
nextx -= x # NAG requires this rather than nextx itself
nextx.shape = N*N # Make nextx 1D since that's what NAG needs
return nextx
# We can now call the NAG Anderson Acceleration routine. If we set `m=0`, no acceleration is applied and the routine simply acts as a driver for our fixed point iterations. We should get exactly the same behavior as our original code. We can see that this is the case, right down to the 28,734 iterations required for the Jacobi method to solve.
#
# This gives us confidence that our code changes haven't broken anything in the simulation.
N = 100
x0 = init_problem(N)
x0.shape = N*N
NAG_jacobi_info = NAG_solverinfo()
nag_source = source(N)
NAG_jacobi_info.source = nag_source
tol = 1e-9
m = 0
NAG_jacobi_sol, fvec = roots.sys_func_aa(NAG_jacobi, x0, tol, eps, m, data=NAG_jacobi_info)
print(f"NAG driven Jacobi on a {N} x {N} grid with no acceleration")
print(f"Solution found in {NAG_jacobi_info.iterations} iterations")
fig, axes = plt.subplots(nrows=1, ncols=2, figsize=(10, 6))
NAG_jacobi_sol.shape = (N, N)
axes[0].imshow(NAG_jacobi_sol)
axes[0].set_title('Potential')
Ex, Ey = np.gradient(NAG_jacobi_sol)
E = np.sqrt(Ex**2+Ey**2) # Magnitude of Electric field
axes[1].imshow(E)
_ = axes[1].set_title('Electric Field')
# Set `m=4` as suggested in the NAG documentation and convergence is much faster. In fact, it's almost 3x faster than Gauss Seidel but we didn't need to change the mathematics in the Jacobi iteration at all. As such, we can think of Anderson Acceleration as a Black-box accelerator for at least some types of fixed point iteration.
N = 100
x0 = init_problem(N)
x0.shape = N*N
NAG_jacobi_info = NAG_solverinfo()
NAG_jacobi_info.source = source(N)
tol = 1e-9
m=4
NAG_jacobi_sol, fvec = roots.sys_func_aa(NAG_jacobi, x0, tol, eps, m, data=NAG_jacobi_info)
print(f"Anderson Accelerated Jacobi with m={m} on a {N} by {N} grid")
print(f"Solution found in {NAG_jacobi_info.iterations} iterations")
# We will now investigate how the choice of `m` affects the number of iterations required for The Jacobi method.
def find_best_m(m):
N = 100
x0 = init_problem(N)
x0.shape = N*N
NAG_jacobi_info = NAG_solverinfo()
NAG_jacobi_info.source = source(N)
tol = 1e-9
_ = roots.sys_func_aa(NAG_jacobi, x0, tol, eps, m, data=NAG_jacobi_info)
return NAG_jacobi_info.iterations
mlist = np.arange(1, 200, 1)
iterations = [find_best_m(m) for m in mlist]
plt.plot(mlist, iterations)
plt.title('Anderson Accelerated Jacobi\nNumber of iterations vs m')
plt.xlabel('m')
_ = plt.ylabel('Iterations')
# This is much nicer than the acceleration parameter for SOR. We can just pick something high and be sure of good performance. Let's go even higher than explored above and use `m=250` before moving on.
# +
N = 100
x0 = init_problem(N)
x0.shape = N*N
NAG_jacobi_info = NAG_solverinfo()
NAG_jacobi_info.source = source(N)
tol = 1e-9
m = 250
NAG_jacobi_sol, fvec = roots.sys_func_aa(NAG_jacobi, x0, tol, eps, m, data=NAG_jacobi_info)
print(f"Anderson Accelerated Jacobi with m={m} on a {N} by {N} grid")
print(f"Solution found in {NAG_jacobi_info.iterations} iterations")
fig, axes = plt.subplots(nrows=1, ncols=2, figsize=(10, 6))
NAG_jacobi_sol.shape = (N, N)
axes[0].imshow(NAG_jacobi_sol)
axes[0].set_title('Potential')
Ex, Ey = np.gradient(NAG_jacobi_sol)
E = np.sqrt(Ex**2+Ey**2) # Magnitude of Electric field
axes[1].imshow(E)
_ = axes[1].set_title('Electric Field')
# -
# This is converging over 90x faster than the unaccelerated Jacobi method and even beats the optimal SOR method by a healthy margin.
# ### Anderson Acceleration and Gauss--Seidel
#
# Next we combine Anderson Acceleration with the Gauss-Seidel method. Since Gauss-Seidel is significantly faster than Jacobi, we might hope that this new combination will be even better.
@jit
def NAG_gauss_seidel(x, solverinfo):
N = int(np.sqrt(x.size))
x.shape = (N, N) # Make x 2D because that's how I think
nextx = np.copy(x)
h = 1/(N-1)
for i in range(1, N - 1):
for j in range(1, N - 1):
nextx[j, i] = 0.25 * (nextx[j+1, i] + nextx[j, i+1] + nextx[j-1, i] + nextx[j, i-1]) + 0.25*h**2*solverinfo.source[j, i]
solverinfo.iterations += 1
nextx -= x # NAG requires this rather than nextx itself
nextx.shape = N*N # Make nextx 1D since that's what NAG needs
return nextx
# +
# Define and run the simulation using Gauss--Seidel and NAG Anderson Acceleration
N = 100
x0 = init_problem(N)
x0.shape = N*N
NAG_gs_info = NAG_solverinfo()
NAG_gs_info.source = source(N)
tol = 1e-9
m = 4
NAG_jacobi_sol, fvec = roots.sys_func_aa(NAG_gauss_seidel, x0, tol, eps, m, data=NAG_gs_info)
print(f"Anderson Accelerated Gauss-Seidel with m={m} on a {N} by {N} grid")
print(f"Solution found in {NAG_gs_info.iterations} iterations")
# -
# With the standard choice of `m=4` we are doing almost as well as the best value of $\omega$ we could find for SOR. Switching to `m=7` and we do even better than SOR.
# +
# Define and run the simulation using Gauss--Seidel and NAG Anderson Acceleration
N = 100
x0 = init_problem(N)
x0.shape = N*N
NAG_gs_info = NAG_solverinfo()
NAG_gs_info.source = source(N)
tol = 1e-9
m = 7
NAG_jacobi_sol, fvec = roots.sys_func_aa(NAG_gauss_seidel, x0, tol, eps, m, data=NAG_gs_info)
print(f"Anderson Accelerated Gauss--Seidel with m={m} on a {N} by {N} grid")
print(f"Solution found in {NAG_gs_info.iterations} iterations")
# -
# Once again, we find ourselves needing to find the perfect acceleration parameter. Let's see how the number of iterations varies with m.
def find_best_m(m): # pylint: disable=function-redefined
N = 100
x0 = init_problem(N)
x0.shape = N*N
NAG_gs_info = NAG_solverinfo()
NAG_gs_info.source = source(N)
tol = 1e-9
_ = roots.sys_func_aa(NAG_gauss_seidel, x0, tol, eps, m, data=NAG_gs_info)
return NAG_gs_info.iterations
# +
mlist = np.arange(1, 200, 1)
iterations = [find_best_m(m) for m in mlist]
plt.plot(mlist, iterations)
plt.hlines(486, -1, 200, 'r')
plt.title('Anderson Accelerated Gauss--Seidel\nNumber of iterations vs m')
plt.xlabel('m')
plt.ylabel('Iterations')
label_x = 30
label_y = 1500
arrow_x = 20
arrow_y = 500
arrow_properties = dict(
facecolor="black", width=0.5,
headwidth=4, shrink=0.1)
_ = plt.annotate(
"Best SOR result", xy=(arrow_x, arrow_y),
xytext=(label_x, label_y),
arrowprops=arrow_properties)
# -
# As with the accelerated Jacobi iterations, the situation with the Gauss--Seidel Anderson Acceleration parameter is rather better than the SOR parameter in that, for this simulation at least, we **always** do better than the best SOR result (486 iterations) for all `m>6` with the very best result being 285 iterations at `m=200`. Furthermore, once you get past `m=10` for this simulation, the curve is pretty flat meaning that we are not being excessively punished for a bad choice of `m`.
# +
# Define and run the simulation using Gauss--Seidel and NAG Anderson Acceleration
N = 100
x0 = init_problem(N)
x0.shape = N*N
NAG_gs_info = NAG_solverinfo()
NAG_gs_info.source = source(N)
tol = 1e-9
m = 200
NAG_jacobi_sol, fvec = roots.sys_func_aa(NAG_gauss_seidel, x0, tol, eps, m, data=NAG_gs_info)
print(f"Anderson Accelerated Gauss--Seidel with m={m} on a {N} by {N} grid")
print(f"Solution found in {NAG_gs_info.iterations} iterations")
# -
# ## Combined SOR and Anderson Acceleration
#
# The next natural question to ask is 'Could we combine both acceleration methods and do even better than anything seen so far'?
@jit
def NAG_SOR(x, solverinfo):
N = int(np.sqrt(x.size))
x.shape = (N, N) # Make x 2D because that's how I think
nextx = np.copy(x)
h = 1/(N-1)
w = solverinfo.w
for i in range(1, N - 1):
for j in range(1, N - 1):
new = 0.25 * (nextx[j-1, i] + nextx[j+1, i] + nextx[j, i-1]+ nextx[j, i+1]) + 0.25*h**2*solverinfo.source[j, i]
nextx[j, i] += w * (new - nextx[j, i])
solverinfo.iterations += 1
nextx -= x # NAG requires this rather than nextx itself
nextx.shape = N*N # Make nextx 1D since that's what NAG needs
return nextx
# Define and run the simulation using SOR and NAG Anderson Acceleration
N = 100
x0 = init_problem(N)
x0.shape = N*N
NAG_SOR_info = NAG_solverinfo()
NAG_SOR_info.w = 1.94
NAG_SOR_info.source = source(N)
tol = 1e-9
m = 200
NAG_SOR_sol, fvec = roots.sys_func_aa(NAG_SOR, x0, tol, eps, m, data=NAG_SOR_info)
print(f"Anderson Accelerated SOR with m={m} and w={NAG_SOR_info.w} on a {N} by {N} grid")
print(f"Solution found in {NAG_SOR_info.iterations} iterations")
# We see that using the best values of `m` and $\omega$ found so far gave us a result worse than using either of them independently.
# Scanning over the range of `m` and $\omega$, we have the following results
def find_best_mw(m, w):
N = 100
x0 = init_problem(N)
x0.shape = N*N
NAG_SOR_info = NAG_solverinfo()
NAG_SOR_info.source = source(N)
NAG_SOR_info.w = w
tol = 1e-9
_ = roots.sys_func_aa(NAG_SOR, x0, tol, eps, m, data=NAG_SOR_info)
return NAG_SOR_info.iterations
# This will take a LONG LONG time!
wlist = np.arange(1.01, 1.99, 0.01)
mlist = np.arange(5, 200, 1)
mw_list = [(m, w, find_best_mw(m, w)) for m in mlist for w in wlist]
# Best 5
mw_list.sort(key=lambda x:x[2])
mw_list[0:10]
# Worst 10
mw_list.sort(key=lambda x:x[2], reverse=True)
mw_list[0:10]
# All of the best results have high values of the Anderson Acceleration factor, `m` so we could have restricted our search there to save time. For this simulation, more is better where `m` is concerned. Finding the best value of $\omega$ for SOR, however is less obvious.
#
# By using SOR and Anderson Acceleration we **can** do better than pure SOR or Anderson Accelerated Gauss--Seidel but it is more difficult to choose the best combination of $\omega$ and `m`. Additionally, the returns are increasingly diminished.
# # Conclusions
#
# Anderson Acceleration is a general method for accelerating many fixed point iteration methods. We have shown that it can be used to reduce the number of iterations required in the traditional Jacobi, Gauss--Seidel and SOR methods for solving Laplace's and Poisson's equations in two dimensions.
#
# Anderson Acceleration requires the user to select a parameter `m` which refers to the number of previous iterations used to produce the next one. For this simulation, high values of `m` give good results and getting the value of `m` wrong does not punish us too much.
#
# | Method | Unaccelerated | Accelerated |
# |--------------|---------------|-------------|
# | Jacobi | 28734 | 313 |
# | Gauss--Seidel| 14877 | 285 |
# | SOR | 486 | 230 |
| roots/Anderson_Acceleration_Poisson.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 3. Train-Predict
#
# **Tensorboard**
# - Input at command: tensorboard --logdir=./log
# - Input at browser: http://127.0.0.1:6006
# +
import time
import os
import pandas as pd
project_name = 'SceneClassification2017'
step_name = 'Train'
time_str = time.strftime("%Y%m%d_%H%M%S", time.localtime())
run_name = project_name + '_' + step_name + '_' + time_str
print('run_name: ' + run_name)
cwd = os.getcwd()
model_path = os.path.join(cwd, 'model')
print('model_path: ' + model_path)
# +
import h5py
import numpy as np
from sklearn.utils import shuffle
np.random.seed(2017)
x_train = []
y_train = {}
x_val = []
y_val = {}
x_test = []
cwd = os.getcwd()
feature_cgg16 = os.path.join(cwd, 'model', 'feature_VGG16_{}.h5'.format(171023))
feature_cgg19 = os.path.join(cwd, 'model', 'feature_VGG19_{}.h5'.format(171023))
feature_resnet50 = os.path.join(cwd, 'model', 'feature_ResNet50_{}.h5'.format(171023))
feature_mobilenet = os.path.join(cwd, 'model', 'feature_MobileNet_{}.h5'.format(171023))
feature_xception = os.path.join(cwd, 'model', 'feature_Xception_{}.h5'.format(171023))
feature_inception = os.path.join(cwd, 'model', 'feature_InceptionV3_{}.h5'.format(171023))
for filename in [feature_cgg16, feature_cgg19, feature_resnet50, feature_xception, feature_inception]:
with h5py.File(filename, 'r') as h:
x_train.append(np.array(h['train']))
y_train = np.array(h['train_label'])
x_val.append(np.array(h['val']))
y_val = np.array(h['val_label'])
x_test.append(np.array(h['test']))
# print(x_train[0].shape)
x_train = np.concatenate(x_train, axis=-1)
# y_train = np.concatenate(y_train, axis=0)
x_val = np.concatenate(x_val, axis=-1)
# y_val = np.concatenate(y_val, axis=0)
x_test = np.concatenate(x_test, axis=-1)
print(x_train.shape)
print(x_train.shape[1:])
print(len(y_train))
print(x_val.shape)
print(len(y_val))
print(x_test.shape)
# -
from sklearn.utils import shuffle
(x_train, y_train) = shuffle(x_train, y_train)
# +
from keras.utils.np_utils import to_categorical
y_train = to_categorical(y_train)
y_val = to_categorical(y_val)
print(y_train.shape)
print(y_val.shape)
# +
from keras.models import *
from keras.layers import *
from keras.optimizers import Adam
# model = Sequential()
# model.add(Dense(5000, input_shape=x_train.shape[1:]))
# model.add(Dropout(0.5))
# model.add(Dense(120, activation='softmax'))
# inputs = Input(x_train.shape[1:])
# x = inputs
# x = Dropout(0.5)(x)
# x = Dense(120, activation='softmax')(x)
# model = Model(inputs, x)
model = Sequential()
model.add(Dense(1000, input_shape=x_train.shape[1:]))
model.add(Dropout(0.5))
model.add(Dense(1000, activation='sigmoid'))
model.add(Dropout(0.5))
model.add(Dense(120, activation='softmax'))
model.compile(optimizer=Adam(lr=1e-4),
loss='categorical_crossentropy',
metrics=['accuracy'])
# +
from keras.callbacks import TensorBoard
log_path = os.path.join(model_path, run_name)
print('log_dir:' + log_path)
tensorBoard = TensorBoard(log_dir=log_path)
# -
hist = model.fit(x_train, y_train,
batch_size=16,
epochs=20, #Increase this when not on Kaggle kernel
verbose=2, #1 for ETA, 0 for silent
validation_data=(x_val, y_val),
callbacks=[tensorBoard])
# +
import matplotlib.pyplot as plt
# %matplotlib inline
plt.plot(hist.history['loss'], color='b')
plt.plot(hist.history['val_loss'], color='r')
plt.show()
plt.plot(hist.history['acc'], color='b')
plt.plot(hist.history['val_acc'], color='r')
plt.show()
# -
def saveModel(model, run_name):
cwd = os.getcwd()
modelPath = os.path.join(cwd, 'model')
if not os.path.isdir(modelPath):
os.mkdir(modelPath)
weigthsFile = os.path.join(modelPath, run_name + '.h5')
model.save(weigthsFile)
saveModel(model, run_name)
# ## Predict
# +
# Used to load model directly and skip train
# import os
# from keras.models import load_model
# cwd = os.getcwd()
# model = load_model(os.path.join(cwd, 'model', 'Dog_Breed_Identification_Train_20171024_155154.h5'))
# -
y_pred = model.predict(x_test, batch_size=128)
print(y_pred.shape)
# +
# print(y_pred[:10])
# y_pred = np.clip(y_pred, 0.005, 0.995)
# print(y_pred[:10])
# -
files = os.listdir(os.path.join(cwd, 'input', 'data_test', 'test'))
print(files[:10])
cwd = os.getcwd()
df = pd.read_csv(os.path.join(cwd, 'input', 'labels.csv'))
print('lables amount: %d' %len(df))
df.head()
n = len(df)
breed = set(df['breed'])
n_class = len(breed)
class_to_num = dict(zip(breed, range(n_class)))
num_to_class = dict(zip(range(n_class), breed))
print(breed)
df2 = pd.read_csv('.\\input\\sample_submission.csv')
n_test = len(df2)
print(df2.shape)
for i in range(0, 120):
df2.iloc[:,[i+1]] = y_pred[:,i]
df2.to_csv('.\\output\\pred.csv', index=None)
print('run_name: ' + run_name)
print('Done !')
| SceneClassification2017/3. Train-Predict.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import arviz as az
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import pymc3 as pm
import warnings
from scipy import stats
pd.options.mode.chained_assignment = None
warnings.filterwarnings('ignore', category=FutureWarning)
# -
# %config Inline.figure_format = 'retina'
az.style.use('arviz-darkgrid')
az.rcParams['stats.credible_interval'] = 0.89 # set credible interval for entire notebook
az.rcParams['stats.information_criterion'] = 'waic' # set information criterion to use in `compare`
az.rcParams['stats.ic_scale'] = 'deviance' # set information criterion scale
np.random.seed(0)
# #### Code 8.1
# +
d = pd.read_csv("Data/rugged.csv", delimiter=";")
# make log version of the outcome
d["log_gdp"] = np.log(d["rgdppc_2000"])
# extract countries with GDP data
dd = d.dropna(subset=["log_gdp"])
# rescale variables
dd["log_gdp_std"] = dd["log_gdp"] / dd["log_gdp"].mean()
dd["rugged_std"] = dd["rugged"] / dd["rugged"].max()
# -
# #### Code 8.2
with pm.Model() as m_8_1:
a = pm.Normal("a", 1, 1)
b = pm.Normal("b", 0, 1)
mu = a + b * (dd["rugged_std"] - 0.215)
sigma = pm.Exponential("sigma", 1)
log_gdp_std = pm.Normal("log_gdp_std", mu, sigma, shape=dd.shape[0])
# #### Code 8.3
# +
with m_8_1:
m_8_1_prior = pm.sample_prior_predictive()
# Figure 8.3 is below
# -
# #### Code 8.4
np.sum(np.abs(m_8_1_prior["b"]) > 0.6) / len(m_8_1_prior["b"])
# #### Code 8.5
with pm.Model() as m_8_1t:
a = pm.Normal("a", 1, 0.1)
b = pm.Normal("b", 0, 0.3)
mu = a + b * (dd["rugged_std"] - 0.215)
sigma = pm.Exponential("sigma", 1)
log_gdp_std = pm.Normal("log_gdp_std", mu, sigma, observed=dd["log_gdp_std"])
m_8_1t_trace = pm.sample()
m_8_1t_prior = pm.sample_prior_predictive()
# +
# Figure 8.3
_, (ax1, ax2) = plt.subplots(1, 2, figsize=[7, 4], constrained_layout=True)
n = 100
rugged_plot = np.linspace(-0.1, 1.1)
ax1.plot(
rugged_plot,
m_8_1_prior["a"][:n].T + rugged_plot.reshape(-1, 1) * m_8_1_prior["b"][:n].T,
color="k",
lw=1,
alpha=0.3,
)
ax1.set_title("a ~ Normal(1, 1)\nb ~ Normal(0, 1)")
ax2.plot(
rugged_plot,
m_8_1t_prior["a"][:n].T + rugged_plot.reshape(-1, 1) * m_8_1t_prior["b"][:n].T,
color="k",
lw=1,
alpha=0.3,
)
ax2.set_title("a ~ Normal(1, 0.1)\nb ~ Normal(0, 0.3)")
for ax in (ax1, ax2):
ax.set_xlabel("ruggedness")
ax.set_xlabel("log GDP (prop of mean)")
ax.axhline(0.7, ls="dashed", color="k", lw=1)
ax.axhline(1.3, ls="dashed", color="k", lw=1)
ax.set_ylim(0.5, 1.5)
# -
# #### Code 8.6
az.summary(m_8_1t_trace, kind="stats", round_to=2)
# #### Code 8.7
cid = pd.Categorical(dd["cont_africa"])
# #### Code 8.8
with pm.Model() as m_8_2:
a = pm.Normal("a", 1, 0.1, shape=cid.categories.size)
b = pm.Normal("b", 0, 0.3)
mu = a[cid] + b * (dd["rugged_std"] - 0.215)
sigma = pm.Exponential("sigma", 1)
log_gdp_std = pm.Normal("log_gdp_std", mu, sigma, observed=dd["log_gdp_std"])
m_8_2_trace = pm.sample()
# #### Code 8.9
az.compare({"m_8_1t": m_8_1t_trace, "m_8_2": m_8_2_trace})
# #### Code 8.10
az.summary(m_8_2_trace, kind="stats", round_to=2)
# #### Code 8.11
# +
with m_8_2:
m_8_2_posterior = pm.sample_posterior_predictive(m_8_2_trace, vars=[a, b, sigma])
diff_a0_a1 = m_8_2_posterior["a"][:, 1] - m_8_2_posterior["a"][:, 0]
az.hpd(diff_a0_a1)
# -
# #### Code 8.12
# +
fig, ax = plt.subplots()
rugged_plot = np.linspace(-0.1, 1.1)
ax.scatter(
dd.loc[cid == 0, "rugged_std"],
dd.loc[cid == 0, "log_gdp_std"],
label="Not Africa",
facecolor="w",
lw=1,
edgecolor="k",
)
pred0 = m_8_2_posterior["a"][:, 0] + rugged_plot.reshape(-1, 1) * m_8_2_posterior["b"]
ax.plot(rugged_plot, pred0.mean(1), color="grey")
az.plot_hpd(rugged_plot, pred0.T, color="grey", credible_interval=0.97)
ax.scatter(
dd.loc[cid == 1, "rugged_std"],
dd.loc[cid == 1, "log_gdp_std"],
label="Africa",
color="k",
)
pred1 = m_8_2_posterior["a"][:, 1] + rugged_plot.reshape(-1, 1) * m_8_2_posterior["b"]
ax.plot(rugged_plot, pred1.mean(1), color="k")
az.plot_hpd(rugged_plot, pred1.T, color="grey", credible_interval=0.97)
ax.legend(frameon=True)
ax.set_xlim(-0.1, 1.1)
ax.set_xlabel("ruggedness (standardised)")
ax.set_ylabel("log GDP (as proportion of mean)");
# -
# #### Code 8.13
with pm.Model() as m_8_3:
a = pm.Normal("a", 1, 0.1, shape=cid.categories.size)
b = pm.Normal("b", 0, 0.3, shape=cid.categories.size)
mu = a[cid] + b[cid] * (dd["rugged_std"] - 0.215)
sigma = pm.Exponential("sigma", 1)
log_gdp_std = pm.Normal("log_gdp_std", mu, sigma, observed=dd["log_gdp_std"])
m_8_3_trace = pm.sample()
# #### Code 8.14
az.summary(m_8_3_trace, kind="stats", round_to=2)
# #### Code 8.15
az.compare(
{"m_8_1t": m_8_1t_trace, "m_8_2": m_8_2_trace, "m_8_3": m_8_3_trace}, ic="loo"
)
# #### Code 8.16
# +
m_8_3_loo = az.loo(m_8_3_trace, pointwise=True)
plt.plot(m_8_3_loo.loo_i)
# -
# #### Code 8.17
with m_8_3:
m_8_3_posterior = pm.sample_posterior_predictive(m_8_3_trace, var_names=['a', 'b'])
# +
_, axs = plt.subplots(1, 2, figsize=[7, 4], sharey=True, constrained_layout=True)
ax1, ax0 = axs
rugged_plot = np.linspace(-0.1, 1.1)
ax0.scatter(
dd.loc[cid == 0, "rugged_std"],
dd.loc[cid == 0, "log_gdp_std"],
label="Not Africa",
facecolor="w",
lw=1,
edgecolor="k",
)
# calculating predicted manually because this is a pain with categorical variabiles in PyMC3
pred0 = (
m_8_3_posterior["a"][:, 0] + rugged_plot.reshape(-1, 1) * m_8_3_posterior["b"][:, 0]
)
ax0.plot(rugged_plot, pred0.mean(1), color="grey")
az.plot_hpd(rugged_plot, pred0.T, color="grey", credible_interval=0.97, ax=ax0)
ax0.set_title("Non-African Nations")
ax1.scatter(
dd.loc[cid == 1, "rugged_std"],
dd.loc[cid == 1, "log_gdp_std"],
label="Africa",
color="k",
)
# calculating predicted manually because this is a pain with categorical variabiles in PyMC3
pred1 = (
m_8_3_posterior["a"][:, 1] + rugged_plot.reshape(-1, 1) * m_8_3_posterior["b"][:, 1]
)
ax1.plot(rugged_plot, pred1.mean(1), color="k")
az.plot_hpd(rugged_plot, pred1.T, color="grey", credible_interval=0.97, ax=ax1)
ax1.set_title("African Nations")
ax.set_xlim(-0.1, 1.1)
ax0.set_xlabel("ruggedness (standardised)")
ax1.set_xlabel("ruggedness (standardised)")
ax1.set_ylabel("log GDP (as proportion of mean)");
# -
# #### Code 8.18
# +
rugged_plot = np.linspace(-0.1, 1.1)
delta = pred1 - pred0 # using 'pred' from above
plt.plot(rugged_plot, delta.mean(1))
az.plot_hpd(rugged_plot, delta.T)
plt.axhline(0, ls="dashed", zorder=1, color=(0, 0, 0, 0.5))
plt.xlabel("ruggedness")
plt.ylabel("expected difference log GDP")
plt.xlim(0, 1);
# -
# These numbers are quite different from the book - not sure why.
# #### Code 8.19
d = pd.read_csv("Data/tulips.csv", delimiter=";")
d.head()
# #### Code 8.20
d["blooms_std"] = d["blooms"] / d["blooms"].max()
d["water_cent"] = d["water"] - d["water"].mean()
d["shade_cent"] = d["shade"] - d["shade"].mean()
# #### Code 8.21
a = stats.norm.rvs(0.5, 1, 10000)
sum((a < 0) | (a > 1)) / len(a)
# #### Code 8.22
a = stats.norm.rvs(0.5, 0.25, 10000)
sum((a < 0) | (a > 1)) / len(a)
# #### Code 8.23
with pm.Model() as m_8_4:
a = pm.Normal("a", 0.5, 0.25)
bw = pm.Normal("bw", 0, 0.25)
bs = pm.Normal("bs", 0, 0.25)
mu = a + bw * d["water_cent"] + bs * d["shade_cent"]
sigma = pm.Exponential("sigma", 1)
blooms_std = pm.Normal("blooms_std", mu, sigma, observed=d["blooms_std"])
m_8_4_trace = pm.sample()
m_8_4_post = pm.sample_posterior_predictive(m_8_4_trace, var_names=['a', 'bw', 'bs'])
# #### Code 8.24
with pm.Model() as m_8_5:
a = pm.Normal("a", 0.5, 0.25)
bw = pm.Normal("bw", 0, 0.25)
bs = pm.Normal("bs", 0, 0.25)
bws = pm.Normal("bws", 0, 0.25)
mu = (
a
+ bw * d["water_cent"]
+ bs * d["shade_cent"]
+ bws * d["water_cent"] * d["shade_cent"]
)
sigma = pm.Exponential("sigma", 1)
blooms_std = pm.Normal("blooms_std", mu, sigma, observed=d["blooms_std"])
m_8_5_trace = pm.sample()
m_8_5_post = pm.sample_posterior_predictive(m_8_5_trace, var_names=['a', 'bw', 'bs', 'bws'])
# #### Code 8.25
# +
_, axs = plt.subplots(2, 3, figsize=[9, 5], sharey=True, sharex=True, constrained_layout=True)
n_lines = 20
pred_x = np.array([-1, 1])
for i, shade in enumerate([-1, 0, 1]):
ind = d.shade_cent == shade
for ax in axs[:, i]:
ax.scatter(d.loc[ind, "water_cent"], d.loc[ind, "blooms_std"])
# top row, m_8_4
ax = axs[0, i]
ax.set_title(f"m8.4 post: shade = {shade:.0f}", fontsize=11)
pred_y = (
m_8_4_post["a"][:n_lines]
+ m_8_4_post["bw"][:n_lines] * pred_x.reshape(-1, 1)
+ m_8_4_post["bs"][:n_lines] * shade
)
ax.plot(pred_x, pred_y, lw=1, color=(0, 0, 0, 0.4))
# bottom row, m_8_5
ax = axs[1, i]
ax.set_title(f"m8.5 post: shade = {shade:.0f}", fontsize=11)
pred_y = (
m_8_5_post["a"][:n_lines]
+ m_8_5_post["bw"][:n_lines] * pred_x.reshape(-1, 1)
+ m_8_5_post["bs"][:n_lines] * shade
+ m_8_5_post["bws"][:n_lines] * pred_x.reshape(-1, 1) * shade
)
ax.plot(pred_x, pred_y, lw=1, color=(0, 0, 0, 0.4))
for ax in axs.flat:
if ax.is_first_col():
ax.set_ylabel("blooms")
if ax.is_last_row():
ax.set_xlabel("water");
# -
# #### Code 8.26
# +
with m_8_4:
m_8_4_priors = pm.sample_prior_predictive(var_names=["a", "bw", "bs"])
with m_8_5:
m_8_5_priors = pm.sample_prior_predictive(var_names=["a", "bw", "bs", "bws"])
# +
_, axs = plt.subplots(2, 3, figsize=[9, 5], sharey=True, sharex=True, constrained_layout=True)
n_lines = 20
pred_x = np.array([-1, 1])
for i, shade in enumerate([-1, 0, 1]):
# top row, m_8_4
ax = axs[0, i]
ax.set_title(f"m8.4 prior: shade = {shade:.0f}", fontsize=11)
pred_y = (
m_8_4_priors["a"][:n_lines]
+ m_8_4_priors["bw"][:n_lines] * pred_x.reshape(-1, 1)
+ m_8_4_priors["bs"][:n_lines] * shade
)
ax.plot(pred_x, pred_y, lw=1, color=(0, 0, 0, 0.4))
ax.plot(pred_x, pred_y[:, 0], lw=2, color="k")
# bottom row, m_8_5
ax = axs[1, i]
ax.set_title(f"m8.5 prior: shade = {shade:.0f}", fontsize=11)
pred_y = (
m_8_5_priors["a"][:n_lines]
+ m_8_5_priors["bw"][:n_lines] * pred_x.reshape(-1, 1)
+ m_8_5_priors["bs"][:n_lines] * shade
+ m_8_5_priors["bws"][:n_lines] * pred_x.reshape(-1, 1) * shade
)
ax.plot(pred_x, pred_y, lw=1, color=(0, 0, 0, 0.4))
ax.plot(pred_x, pred_y[:, 0], lw=2, color="k")
for ax in axs.flat:
ax.set_ylim(-0.5, 1.5)
ax.axhline(1, ls="dashed", color=(0, 0, 0, 0.6))
ax.axhline(0, ls="dashed", color=(0, 0, 0, 0.6))
if ax.is_first_col():
ax.set_ylabel("blooms")
if ax.is_last_row():
ax.set_xlabel("water");
# -
# %load_ext watermark
# %watermark -n -u -v -iv -w
| Rethinking_2/Chp_08.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Bayesian Statistics Made Simple
# ===
#
# Code and exercises from my workshop on Bayesian statistics in Python.
#
# Copyright 2016 <NAME>
#
# MIT License: https://opensource.org/licenses/MIT
# +
# If we're running on Colab, install empiricaldist
# https://pypi.org/project/empiricaldist/
import sys
IN_COLAB = 'google.colab' in sys.modules
if IN_COLAB:
# !pip install empiricaldist
# +
import numpy as np
import pandas as pd
import seaborn as sns
sns.set_style('white')
sns.set_context('talk')
import matplotlib.pyplot as plt
from empiricaldist import Pmf
# -
# ### The Euro problem
#
# *"When spun on edge 250 times, a Belgian one-euro coin came up heads 140 times and tails 110. 'It looks very suspicious to me,' said <NAME>, a statistics lecturer at the London School of Economics. 'If the coin were unbiased, the chance of getting a result as extreme as that would be less than 7%.' "*
#
# From “The Guardian” quoted by MacKay, *Information Theory, Inference, and Learning Algorithms*.
#
#
# **Exercise 1:** Write a function called `likelihood_euro` that defines the likelihood function for the Euro problem. Note that `hypo` is in the range 0 to 100.
#
# Here's an outline to get you started.
def likelihood_euro(data, hypo):
""" Likelihood function for the Euro problem.
data: string, either 'H' or 'T'
hypo: prob of heads (0-100)
returns: float probability
"""
#print(data, hypo)
x = hypo / 100
return x if data == 'H' else 1-x
# +
# Solution goes here
# -
# For the prior, we'll start with a uniform distribution from 0 to 100.
def decorate_euro(title):
"""Labels the axes.
title: string
"""
plt.xlabel('Probability of heads')
plt.ylabel('PMF')
plt.title(title)
euro = Pmf.from_seq(range(101))
euro.plot()
decorate_euro('Prior distribution')
# Now we can update with a single heads:
euro.update(likelihood_euro, 'H')
euro.plot()
decorate_euro('Posterior distribution, one heads')
# Another heads:
euro.update(likelihood_euro, 'H')
euro.plot()
decorate_euro('Posterior distribution, two heads')
# And a tails:
euro.update(likelihood_euro, 'T')
euro.plot()
decorate_euro('Posterior distribution, HHT')
# Starting over, here's what it looks like after 7 heads and 3 tails.
# +
euro = Pmf.from_seq(range(101))
for outcome in 'HHHHHHHTTT':
euro.update(likelihood_euro, outcome)
euro.plot()
decorate_euro('Posterior distribution, 7 heads, 3 tails')
# -
# The maximum apostiori probability (MAP) is 70%, which is the observed proportion.
euro.max_prob()
# Here are the posterior probabilities after 140 heads and 110 tails.
# +
euro = Pmf.from_seq(range(101))
evidence = 'H' * 140 + 'T' * 110
for outcome in evidence:
euro.update(likelihood_euro, outcome)
euro.plot()
decorate_euro('Posterior distribution, 140 heads, 110 tails')
# -
# The posterior mean is about 56%
euro.mean()
# So is the MAP.
euro.max_prob()
# And the median (50th percentile).
euro.quantile(0.5)
# The posterior credible interval has a 90% chance of containing the true value (provided that the prior distribution truly represents our background knowledge).
euro.credible_interval(0.9)
# ### Swamping the prior
#
# The following function makes a Euro object with a triangle prior.
def TrianglePrior():
"""Makes a Suite with a triangular prior.
"""
suite = Pmf(name='triangle')
for x in range(0, 51):
suite[x] = x
for x in range(51, 101):
suite[x] = 100-x
suite.normalize()
return suite
# And here's what it looks like:
# +
euro1 = Pmf.from_seq(range(101), name='uniform')
euro1.plot()
euro2 = TrianglePrior()
euro2.plot()
plt.legend()
decorate_euro('Prior distributions')
# -
# **Exercise 9:** Update `euro1` and `euro2` with the same data we used before (140 heads and 110 tails) and plot the posteriors. How big is the difference in the means?
# Solution goes here
evidence = 'H' * 140 + 'T' * 110
for outcome in evidence:
euro1.update(likelihood_euro, outcome)
euro2.update(likelihood_euro, outcome)
euro1.plot()
euro2.plot()
# The posterior distributions are not identical, but with this data, they converge to the point where there is no practical difference, for most purposes.
| BayesMadeSimple/03_euro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="e4rn3W3OYJMu"
# # Deep Q-Learning
# + [markdown] id="TzKNR1XpYJMu"
# Install dependencies for AI gym to run properly (shouldn't take more than a minute). If running on google cloud or running locally, only need to run once. Colab may require installing everytime the vm shuts down.
# + colab={"base_uri": "https://localhost:8080/"} id="3vhcSU8OYJMu" outputId="87ca90f3-7532-4308-85fb-1fcade8b5569"
# !pip3 install gym pyvirtualdisplay
# !sudo apt-get install -y xvfb python-opengl ffmpeg
# + colab={"base_uri": "https://localhost:8080/"} id="i59iSmO7YJMv" outputId="047baa93-7649-4016-9e38-ec7dc025cb60"
# !pip3 install --upgrade setuptools
# !pip3 install ez_setup
# !pip3 install gym[atari]
# + [markdown] id="Vd7PfpL3YJMv"
# For this assignment we will implement the Deep Q-Learning algorithm with Experience Replay as described in breakthrough paper __"Playing Atari with Deep Reinforcement Learning"__. We will train an agent to play the famous game of __Breakout__.
# + colab={"base_uri": "https://localhost:8080/"} id="EG1FXI_-YPne" outputId="a799ed48-67db-49ae-bdf1-25d79d986746"
from google.colab import drive
drive.mount('/content/gdrive')
import os
os.chdir("gdrive/My Drive/Colab Notebooks/assignment5_materials")
# + id="-7xSjWIeYJMv"
# %matplotlib inline
import sys
import gym
import torch
import pylab
import random
import numpy as np
from collections import deque
from datetime import datetime
from copy import deepcopy
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
from torch.autograd import Variable
from utils import find_max_lives, check_live, get_frame, get_init_state
from model import DQN
from config import *
import matplotlib.pyplot as plt
# # %load_ext autoreload
# # %autoreload 2
# + [markdown] id="rAY55N-1YJMv"
# ## Understanding the environment
# + [markdown] id="Q1Uu1k6gYJMv"
# In the following cell, we initialize our game of __Breakout__ and you can see how the environment looks like. For further documentation of the of the environment refer to https://gym.openai.com/envs.
#
# In breakout, we will use 3 actions "fire", "left", and "right". "fire" is only used to reset the game when a life is lost, "left" moves the agent left and "right" moves the agent right.
# + id="s-bGTBfUYJMv"
env = gym.make('BreakoutDeterministic-v4')
state = env.reset()
# + id="41itT10EYJMv"
number_lives = find_max_lives(env)
state_size = env.observation_space.shape
action_size = 3 #fire, left, and right
# + [markdown] id="oZhHUWdJYJMv"
# ## Creating a DQN Agent
# + [markdown] id="V54Nfb9iYJMv"
# Here we create a DQN Agent. This agent is defined in the __agent.py__. The corresponding neural network is defined in the __model.py__. Once you've created a working DQN agent, use the code in agent.py to create a double DQN agent in __agent_double.py__. Set the flag "double_dqn" to True to train the double DQN agent.
#
# __Evaluation Reward__ : The average reward received in the past 100 episodes/games.
#
# __Frame__ : Number of frames processed in total.
#
# __Memory Size__ : The current size of the replay memory.
# + id="6abEqyyhYJMv"
double_dqn = True # set to True if using double DQN agent
if double_dqn:
from agent_double import Agent
else:
from agent import Agent
agent = Agent(action_size)
evaluation_reward = deque(maxlen=evaluation_reward_length)
frame = 0
memory_size = 0
# + [markdown] id="kyunIOlPYJMv"
# ### Main Training Loop
# + [markdown] id="jOTKupI8YJMv"
# In this training loop, we do not render the screen because it slows down training signficantly. To watch the agent play the game, run the code in next section "Visualize Agent Performance"
# + colab={"base_uri": "https://localhost:8080/"} id="DwhM69mjYJMv" outputId="49d8e5c6-746f-4a92-d72e-061d9d4e3b93"
rewards, episodes = [], []
best_eval_reward = 0
for e in range(EPISODES):
done = False
score = 0
history = np.zeros([5, 84, 84], dtype=np.uint8)
step = 0
d = False
state = env.reset()
next_state = state
life = number_lives
get_init_state(history, state)
while not done:
step += 1
frame += 1
# Perform a fire action if ball is no longer on screen to continue onto next life
if step > 1 and len(np.unique(next_state[:189] == state[:189])) < 2:
action = 0
else:
action = agent.get_action(np.float32(history[:4, :, :]) / 255.)
state = next_state
next_state, reward, done, info = env.step(action + 1)
frame_next_state = get_frame(next_state)
history[4, :, :] = frame_next_state
terminal_state = check_live(life, info['ale.lives'])
life = info['ale.lives']
r = np.clip(reward, -1, 1)
r = reward
# Store the transition in memory
agent.memory.push(deepcopy(frame_next_state), action, r, terminal_state)
# Start training after random sample generation
if(frame >= train_frame):
agent.train_policy_net(frame)
# Update the target network only for Double DQN only
if double_dqn and (frame % update_target_network_frequency)== 0:
agent.update_target_net()
score += reward
history[:4, :, :] = history[1:, :, :]
if done:
evaluation_reward.append(score)
rewards.append(np.mean(evaluation_reward))
episodes.append(e)
pylab.plot(episodes, rewards, 'b')
pylab.xlabel('Episodes')
pylab.ylabel('Rewards')
pylab.title('Episodes vs Reward')
pylab.savefig("./save_graph/breakout_dqn.png") # save graph for training visualization
# every episode, plot the play time
print("episode:", e, " score:", score, " memory length:",
len(agent.memory), " epsilon:", agent.epsilon, " steps:", step,
" lr:", agent.optimizer.param_groups[0]['lr'], " evaluation reward:", np.mean(evaluation_reward))
# if the mean of scores of last 100 episode is bigger than 5 save model
### Change this save condition to whatever you prefer ###
if np.mean(evaluation_reward) > 5 and np.mean(evaluation_reward) > best_eval_reward:
torch.save(agent.policy_net, "./save_model/breakout_dqn.pth")
best_eval_reward = np.mean(evaluation_reward)
# + colab={"base_uri": "https://localhost:8080/"} id="tx9sUtOFgSky" outputId="1f60396a-a5a8-416d-bf2a-fd1cc09267e6"
rewards, episodes = [], []
best_eval_reward = 0
for e in range(EPISODES):
done = False
score = 0
history = np.zeros([5, 84, 84], dtype=np.uint8)
step = 0
d = False
state = env.reset()
next_state = state
life = number_lives
get_init_state(history, state)
#agent.scheduler.step()
while not done:
step += 1
frame += 1
# Perform a fire action if ball is no longer on screen to continue onto next life
if step > 1 and len(np.unique(next_state[:189] == state[:189])) < 2:
action = 0
else:
action = agent.get_action(np.float32(history[:4, :, :]) / 255.)
state = next_state
next_state, reward, done, info = env.step(action + 1)
frame_next_state = get_frame(next_state)
history[4, :, :] = frame_next_state
terminal_state = check_live(life, info['ale.lives'])
life = info['ale.lives']
r = np.clip(reward, -1, 1)
r = reward
# Store the transition in memory
agent.memory.push(deepcopy(frame_next_state), action, r, terminal_state)
# Start training after random sample generation
if(frame >= train_frame):
agent.train_policy_net(frame)
# Update the target network only for Double DQN only
if double_dqn and (frame % update_target_network_frequency)== 0:
agent.update_target_net()
score += reward
history[:4, :, :] = history[1:, :, :]
if done:
evaluation_reward.append(score)
rewards.append(np.mean(evaluation_reward))
episodes.append(e)
pylab.plot(episodes, rewards, 'b')
pylab.xlabel('Episodes')
pylab.ylabel('Rewards')
pylab.title('Episodes vs Reward')
pylab.savefig("./save_graph/breakout_ddqn.png") # save graph for training visualization
# every episode, plot the play time
print("episode:", e, " score:", score, " memory length:",
len(agent.memory), " epsilon:", agent.epsilon, " steps:", step,
" lr:", agent.optimizer.param_groups[0]['lr'], " evaluation reward:", np.mean(evaluation_reward))
# if the mean of scores of last 100 episode is bigger than 5 save model
### Change this save condition to whatever you prefer ###
if np.mean(evaluation_reward) > 5 and np.mean(evaluation_reward) > best_eval_reward:
torch.save(agent.policy_net, "./save_model/breakout_ddqn.pth")
best_eval_reward = np.mean(evaluation_reward)
# + [markdown] id="Skhp2P_lYJMw"
# # Visualize Agent Performance
# + [markdown] id="QPGI3m0OYJMw"
# BE AWARE THIS CODE BELOW MAY CRASH THE KERNEL IF YOU RUN THE SAME CELL TWICE.
#
# Please save your model before running this portion of the code.
# + id="AqbC7NgcYJMw"
from gym.wrappers import Monitor
import glob
import io
import base64
from IPython.display import HTML
from IPython import display as ipythondisplay
from pyvirtualdisplay import Display
# Displaying the game live
def show_state(env, step=0, info=""):
plt.figure(3)
plt.clf()
plt.imshow(env.render(mode='rgb_array'))
plt.title("%s | Step: %d %s" % ("Agent Playing",step, info))
plt.axis('off')
ipythondisplay.clear_output(wait=True)
ipythondisplay.display(plt.gcf())
# Recording the game and replaying the game afterwards
def show_video():
mp4list = glob.glob('video/*.mp4')
if len(mp4list) > 0:
mp4 = mp4list[0]
video = io.open(mp4, 'r+b').read()
encoded = base64.b64encode(video)
ipythondisplay.display(HTML(data='''<video alt="test" autoplay
loop controls style="height: 400px;">
<source src="data:video/mp4;base64,{0}" type="video/mp4" />
</video>'''.format(encoded.decode('ascii'))))
else:
print("Could not find video")
def wrap_env(env):
env = Monitor(env, './video', force=True)
return env
# + id="swqwF3OSkNVN"
agent.load_policy_net('save_model/breakout_ddqn.pth')
agent.update_target_net()
# + id="mfdE_HpxYJMw" colab={"base_uri": "https://localhost:8080/", "height": 438} outputId="46ca7bbd-3a8f-4fc1-a54d-2f464acf4f46"
display = Display(visible=0, size=(300, 200))
display.start()
# Load agent
# agent.load_policy_net("./save_model/breakout_dqn.pth")
agent.epsilon = 0.0 # Set agent to only exploit the best action
env = gym.make('BreakoutDeterministic-v4')
env = wrap_env(env)
done = False
score = 0
step = 0
state = env.reset()
next_state = state
life = number_lives
history = np.zeros([5, 84, 84], dtype=np.uint8)
get_init_state(history, state)
while not done:
# Render breakout
env.render()
# show_state(env,step) # uncommenting this provides another way to visualize the game
step += 1
frame += 1
# Perform a fire action if ball is no longer on screen
if step > 1 and len(np.unique(next_state[:189] == state[:189])) < 2:
action = 0
else:
action = agent.get_action(np.float32(history[:4, :, :]) / 255.)
state = next_state
next_state, reward, done, info = env.step(action + 1)
frame_next_state = get_frame(next_state)
history[4, :, :] = frame_next_state
terminal_state = check_live(life, info['ale.lives'])
life = info['ale.lives']
r = np.clip(reward, -1, 1)
r = reward
# Store the transition in memory
agent.memory.push(deepcopy(frame_next_state), action, r, terminal_state)
# Start training after random sample generation
score += reward
history[:4, :, :] = history[1:, :, :]
env.close()
show_video()
display.stop()
# + id="zJFO0g4nYJMw"
| mp5/assignment5_materials/MP5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## SVD DMR: wav file alignment
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" executionInfo={"elapsed": 2251, "status": "ok", "timestamp": 1548951950015, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11966704463856227449"}, "user_tz": 300} id="r80FflgHhCiH" outputId="143411b2-cc11-47a1-c334-a76291219798"
import os
import torch, torchvision
import numpy as np
import matplotlib.pyplot as plt
import soundfile as sf
from pystoi import stoi
# +
# # %matplotlib notebook
# # %matplotlib inline
# + [markdown] colab_type="text" id="2t_9_D3l0Px9"
# #### Machine paths
# -
path = "/home/david/sigMF_ML/RF_SVD/clean_speech/IQ_files/dmr_aligned/dmr_1k_aligned_20210519/" # ace
path_save = "/home/david/sigMF_ML/RF_SVD/clean_speech/IQ_files/dmr_aligned/dmr_1k_aligned_20210519/stoi_save/" # ace
print(path)
# #### reading sigmf meta data and encoder function
def calculate_sdr(source_signal, estimated_signal, offset=None, scale_invariant=True):
# For regular SDR, set “scale_invariant” to False, for SISDR, set it to True.
s = source_signal
y = estimated_signal
# add a batch axis if non-existant
if len(s.shape) != 2:
s = s.unsqueeze(0)
y = y.unsqueeze(0)
# truncate all signals in the batch to match the minimum-length
min_length = min(s.shape[-1], y.shape[-1])
s = s[..., :min_length]
y = y[..., :min_length]
if scale_invariant:
alpha = s.mm(y.T).diag()
alpha /= ((s ** 2).sum(dim=1) + eps)
alpha = alpha.unsqueeze(1) # to allow broadcasting
else:
alpha = 1
e_target = s * alpha
e_res = e_target - y
numerator = (e_target ** 2).sum(dim=1)
denominator = (e_res ** 2).sum(dim=1) + eps
sdr = 10 * torch.log10((numerator / denominator) + eps)
# if `offset` is non-zero, this function returns the relative SDR
# improvement for each signal in the batch
if offset is not None:
sdr -= offset
return sdr
print(path)
os.chdir(path)
eps = np.finfo(np.float32).eps
# clean, fs = sf.read(path+'original_clean_aligned.wav')
# clean, fs = sf.read(path+'1914-133440-0016.flac')
clean, fs = sf.read(path+'dmr_clean_speech_aligned.wav')
c = len(clean)
original_clean = clean
original_c = c
# print('clean length = ', c)
# print('fs original = ', fs)
rank_list = [5,6,7,8,9,10,1000]
d1 = np.zeros(len(rank_list), dtype = float)
SNR_calc = np.zeros(len(rank_list), dtype = float)
m = 0
# print('d1 = ', d1)
for i in rank_list:
# print('rank =', i)
if (i != 1000):
fullpath =path+'dmr_svd'+'{0:02d}'.format(i)+'_aligned.wav'
denoised, fs = sf.read(fullpath)
de = len(denoised)
# print('de length = ', de)
if c > de:
clean = clean[0:de]
# print('clean new length = ', len(clean))
else:
denoised = denoised[0:c]
# print('denoised new length = ', len(denoised))
# Clean and den should have the same length, and be 1D
d1[m] = stoi(clean, denoised, fs, extended=False)
SNR_calc[m] = calculate_sdr(torch.Tensor(clean), torch.Tensor(denoised))
elif (i == 1000): # This is the full vector - NO SVD
denoised, fs = sf.read(path+'dmr_clean_speech_aligned.wav')
de = len(denoised)
if c > de:
clean = clean[0:de]
else:
denoised = denoised[0:c]
d1[m] = stoi(clean, denoised, fs, extended=False)
SNR_calc[m] = calculate_sdr(torch.Tensor(clean), torch.Tensor(denoised))
m=m+1
clean = original_clean
c = original_c
num = 6
np.asarray(rank_list[:num])
d1[:num]
d2 = [0.8603, 0.9161, 0.9154, 0.9170, 0.9124, 0.9217] # stoi results from Minje's matlab code
d2 = np.asarray(d2[:num])
num = 6
os.chdir(path_save)
np.save('rank_list_dmr', np.asarray(rank_list[:num]))
np.save('dmr', d1[:num])
plt.figure(figsize=(9, 6))
fig = plt.figure()
plt.scatter(rank_list[:num],d2,c='r', label='STOI')
plt.legend(loc='lower right')
plt.title('STOI vs SVD vectors (1000 fft)')
plt.xlabel('SVD vectors')
plt.ylabel('STOI')
plt.ylim(.85, .93)
plt.xlim(4.9, 10.1)
plt.grid()
fig.savefig('dmr_stoi.pdf', format="pdf")
plt.show()
plt.figure(figsize=(9, 6))
fig2 = plt.figure()
plt.scatter(rank_list,d1,c='r', label='STOI')
plt.legend(loc='lower right')
plt.title('STOI vs SVD vectors: All')
plt.xlabel('SVD vectors')
plt.ylabel('STOI')
plt.ylim(.75, .8)
plt.xlim(4, 26)
plt.grid()
fig2.savefig('dmr_stoi_all.png', format="png")
plt.show()
| SVD DMR aligned calculation 1MSPS FINAL-20210519 1k fft.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="TXM78u12VWi8"
# # Check and import lirabries/packages
#
# + [markdown] id="eMx7mVxzU82O"
# **1. Checking packpage/libraries version**
# + [markdown] id="Br_eRaWfVKzP"
#
# + colab={"base_uri": "https://localhost:8080/"} id="U88ueLShzEFS" outputId="46af12b7-398a-48ab-fe14-f42d6d927c60"
import sys
import tensorflow
print(f"Python version: {sys.version}")
print(f"Tensorflow version: {tensorflow.__version__}")
# + [markdown] id="cjUg4jwr0Fbs"
# **2. Import libraries**
# + id="YqDcUniCz7Gy"
import tensorflow as tf
import os
import cv2
import numpy as np
import matplotlib.pyplot as plt
import tqdm
from sklearn.preprocessing import LabelBinarizer
# + [markdown] id="ygz8jvFIVefn"
# # Get data from Google Drive
# + [markdown] id="Fj7_gqKS0crP"
# **3. Mount Drive**
# + colab={"base_uri": "https://localhost:8080/"} id="LEy7EaQq0acO" outputId="f64b3e14-6681-4d38-c0e7-b8f17bda29cd"
from google.colab import drive, files
drive.mount('drive',force_remount=True)
# + id="L_4tKKV-08A1"
# checking if Drive is mounted and accessible
# If shared folder does not appear by using the mount command, then reset Colab runtime
# and mount using Web GUI (or vice versa)
# %cd /content/drive/MyDrive/Dataset/
# %ls
# + [markdown] id="-6dSseVl2AiM"
# **4. Setup PATHS & VARIABLES**
# + colab={"base_uri": "https://localhost:8080/"} id="nhCoS-Mk2Hrz" outputId="e3862cfd-d79d-41db-d850-dd44af7c99dc"
# Define Path to the Dataset folder
BASE_PATH = '/content/drive/MyDrive/Dataset/MP4_OUTPUT'
VIDEOS_PATH = os.path.join(BASE_PATH,'**','*.mp4')
print(VIDEOS_PATH)
# Define LSTM sequence length and batch_size
SEQUENCE_LENGTH = 40
BATCH_SIZE = 16
# + [markdown] id="3yweYB1B28pt"
# **5. Sample Video**
# + [markdown] id="qSYw7lu_3Cfm"
# We will not process every frame, but taking Kth sample where = num_frame_in_videos / SEQUENCE_LENGTH
# + id="8qicZT4b2zGA"
# taking Kth sameple function
def frame_generator():
video_paths = tf.io.gfile.glob(VIDEOS_PATH)
np.random.shuffle(video_paths)
for video_path in video_paths:
frames = []
cap = cv2.VideoCapture(video_path)
num_frames = int(cap.get(cv2.CAP_PROP_FRAME_COUNT))
sample_every_frame = max(1, num_frames // SEQUENCE_LENGTH)
current_frame = 0
label = os.path.basename(os.path.dirname(video_path))
max_images = SEQUENCE_LENGTH
while True:
success, frame = cap.read()
if not success:
break
# OpenCV reads in videos in BGR format so we need to rearrange the channels
# to be in RGB format, resize the image, and preprocess it for the CNN
if current_frame % sample_every_frame == 0:
frame = frame[:, :, ::-1]
img = tf.image.resize(frame, (224,224))
img = tf.keras.applications.mobilenet_v2.preprocess_input(img)
max_images -= 1
yield img, video_path
if max_images == 0:
break
current_frame += 1
# + id="-ge7PedS5aQw"
# Load Dataset
dataset = tf.data.Dataset.from_generator(frame_generator,
output_types=(tf.float32,tf.string),
output_shapes=((224,224,3),()))
# set batch_size
dataset = dataset.batch(BATCH_SIZE).prefetch(tf.data.experimental.AUTOTUNE)
# + colab={"base_uri": "https://localhost:8080/"} id="PpBUjSac558j" outputId="58087bff-d9db-44fe-9e81-e66148ecf2f0"
# Print shape and type of dataset
print(dataset)
# + [markdown] id="5eHbpLQFVmgW"
# # Build Feature Extraction Model
# + [markdown] id="5Hj2_LdC6aZd"
# **6. Build Feature Extraction Model**
# + id="39ARk3cJ6HKC"
mobilenet_v2 = tf.keras.applications.mobilenet_v2.MobileNetV2(input_shape=(224,224,3),
include_top=False,
weights='imagenet')
x = mobilenet_v2.output
# Average Pooling - transforming the feature map from 8*8*2048 to 1x2048
pooling_output = tf.keras.layers.GlobalAveragePooling2D()(x)
feature_extraction_model = tf.keras.Model(mobilenet_v2.input,pooling_output)
# + id="QzKsZ2D37IT6"
# ONLY RUN ONCE
# Extract Feature - Generate .npy file for each video
current_path = None
all_features = []
# cycle through each img and extracts its features
# tqdm is a progress bar which updates each time the feature_extraction_model is called
for img, batch_path in tqdm.tqdm(dataset):
batch_features = feature_extraction_model(img)
# Reshape tensor
batch_features = tf.reshape(batch_features,
(batch_features.shape[0], -1))
for features, path in zip(batch_features.numpy(), batch_path.numpy()):
if path != current_path and current_path is not None:
output_path = current_path.decode().replace('.mp4','.npy')
np.save(output_path,all_features)
all_features = []
current_path = path
all_features.append(features)
# + [markdown] id="IUvs2nLt_dDM"
# **7. Create Labels of our classes**
# + colab={"base_uri": "https://localhost:8080/"} id="fIgRYxwK8yf4" outputId="0c325b22-7d65-47ec-ac39-57acbdde30e0"
# load LABELS
LABELS = ['good','heels_off','bent_over','knees_forward','knees_in','shallow']
# Encode to 0- not belong to class Or 1- belong to class using LabelBinarizer()
encoder = LabelBinarizer()
encoder.fit(LABELS )
# + colab={"base_uri": "https://localhost:8080/"} id="yhCjaE6kCHRk" outputId="8597250e-5bf0-4d24-a41e-ce2945066821"
# Checking output of LabelBinarizer()
print(encoder.classes_)
print(encoder.transform(LABELS))
print(encoder.inverse_transform(t))
# + [markdown] id="a0JO9js8DXh-"
# **8. LSTM Model**
# + [markdown] id="IRmMiVRMDf1P"
# Define LSTM Model with following layers:
#
#
# * Layer 1 = Masking Layer ( see keras [Doc](https://keras.io/api/layers/core_layers/masking/) )
# * Layer 2 = Defind what **1** cell of LSTM looks like [LSTM layer](https://keras.io/api/layers/recurrent_layers/lstm/). The total number of cells is defined as SEQUENCE_LENGTH above
# * Layer 3 = FNC (fully-connected layer) relu activation ( see [Dense layer](https://keras.io/api/layers/core_layers/dense/) )
# * Layer 4 = Drouput layer
# * Layer 5 = final decision FNC layer with softmax activation -- output has length of classes
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="GU8IlI1tCWWP" outputId="92482946-0f8a-4b8c-d467-52c6c461ab57"
model = tf.keras.Sequential([
tf.keras.layers.Masking(mask_value=0.),
tf.keras.layers.LSTM(512,dropout=0.5,recurrent_dropout=0.5), # 512 units (hidden-layer)
tf.keras.layers.Dense(256,activation='relu'),
tf.keras.layers.Dropout(0.5),
tf.keras.layers.Dense(6,activation='softmax')
])
print(len(LABELS))
# + id="kS6g_kndHCKz"
# Setup Loss function and metrics (see more at https://keras.io/api/metrics/)
model.compile(loss='categorical_crossentropy', # since we want to classify different type of squats
optimizer='adam',
metrics=['accuracy','top_k_categorical_accuracy']) # focus on accuracy
# + [markdown] id="vrWAKINPVs4d"
# # Split Training/Test Data
#
# + [markdown] id="tbePgF19HevP"
# **9. Splitting Training/Test data**
# + colab={"base_uri": "https://localhost:8080/"} id="JVWuSJmFHZ2z" outputId="b582c46a-6c8a-4898-ef2b-3dd58cea35c6"
data_path=[]
for root,dir,filename in os.walk(BASE_PATH,topdown=True):
for video_path in filename:
if video_path.endswith('.mp4'):
data_path.append(os.path.join(root,video_path))
# Shuffle video_paths and split to training-test(80-20)
import random
print(len(data_path))
random.shuffle(data_path)
train_list = data_path[int((len(data_path)+1)*.20):] # get 80% of data to training
test_list = data_path[:int((len(data_path)+1)*.20)] # the rest go to testing
# number of train
print(f'Training: {len(train_list)}')
# number of test
print(f'Test/Valid: {len(test_list)}')
# + id="hcPk7fL6JUgb"
# define make_generator() that returns a generator which will randomly shuffle the video list
# then building out the list as the .npy feature files
def make_generator(file_list):
def generator():
np.random.shuffle(file_list)
for path in file_list:
full_path = path.replace('.mp4','.npy')
label = os.path.basename(os.path.dirname(path))
features = np.load(full_path)
padded_sequence = np.zeros((SEQUENCE_LENGTH,1280)) # MobileNet feature extractor
padded_sequence[0:len(features)] = np.array(features)
transform_label = encoder.transform([label])
yield padded_sequence, transform_label[0]
return generator
# + id="3NVnLf0FRE1B"
# setting Training/Test(Validation) Data
# Since we would use a new/unseen dataset for testing, thus the testing set here is used for validation
train_dataset = tf.data.Dataset.from_generator(make_generator(train_list),
output_types=(tf.float32,tf.int16),
output_shapes=((SEQUENCE_LENGTH,1280),(len(LABELS))))
train_dataset = train_dataset.batch(BATCH_SIZE,drop_remainder=True).prefetch(tf.data.experimental.AUTOTUNE)
valid_dataset = tf.data.Dataset.from_generator(make_generator(test_list),
output_types=(tf.float32,tf.int16),
output_shapes=((SEQUENCE_LENGTH,1280),(len(LABELS))))
valid_dataset = valid_dataset.batch(BATCH_SIZE,drop_remainder=True).prefetch(tf.data.experimental.AUTOTUNE)
# Print Train and Valid dataset
print(train_dataset)
print(valid_dataset)
# + [markdown] id="npzR1ypmU1Fe"
# # Train LSTM Model
# + [markdown] id="E3yypD4DTn6O"
# **10. Train LSTM**
# + id="oldVX3t7ULG-"
# Create Log_dir for TensorBoard Visualization
ROOT_PATH = '/content/drive/MyDrive/Dataset'
LOG_DIR = os.path.join(ROOT_PATH,'training_log')
# Create new dir if not exists
if not os.path.isdir(LOG_DIR):
# !mkdir training_log
# + id="YIC3ZNa2SkP7"
# Callback function that will store information (checkpoints,etc) used for TensorBoard
tensorboard_callback = tf.keras.callbacks.TensorBoard(log_dir=LOG_DIR,update_freq=1000)
model.fit(train_dataset,epochs=10,callbacks=[tensorboard_callback],verbose=2,validation_data=valid_dataset)
# + id="Fpn664SZVgRo"
# Save model
model.file = os.path.join(BASE_PATH,'feature_extraction.h5')
model.save(model.file)
# + [markdown] id="i7v5U6HdV28c"
# # Tensorboard and Evaluation on test_dataset
# + [markdown] id="UkqCiXVj4hkh"
# **11. Tensorboard Visualization**
# + id="_4X3hGru4GNQ"
# Load the tensorboard notebook
# %load_ext tensorboard
# %tensorboard --logdir {LOG_DIR}
# + [markdown] id="TXu6VqE66I9F"
# **12. Run Evaluation**
# + colab={"base_uri": "https://localhost:8080/"} id="mc77H-eC48X1" outputId="d25ef1cd-065f-4926-8d43-62ae2c8a31e1"
print('----------------')
print('Evaluate on test Data')
# define test_set path
TEST_PATH = ''
test_result = model.evaluate(test_dataset, verbose=1)
print(f'Test Loss, Test Accuracy: {test_result}')
# + [markdown] id="xSFaN3-D6qqC"
# **13. Run Prediction**
# + id="z1kMG-aV6hMf"
prediction = model.predict(valid_dataset,verbose=1)
print(prediction.shape)
print(prediction)
# + id="OSpqwQac7Bsd"
| USquat_FeatureExtraction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <img src="header_anwender.png" align="left"/>
# # Anwendungsbeispiel Regression Wine Quality
#
# Das Ziel des Beispieles ist es, die Qualität eines Weines aus physikalischen Messgrößen zu schätzen. Dazu verwenden wir verschiedene Arten der Regression.
# Wir verwenden einen Datensatz von Weinen aus Portugal erstellt von Paulo Cortez [1]. Die Details der Erstellung der Daten sind unter folgendem Link zu finden [http://www3.dsi.uminho.pt/pcortez/wine5.pdf](http://www3.dsi.uminho.pt/pcortez/wine5.pdf).
#
# ```
# [1] <NAME>, <NAME>, <NAME>, <NAME> and <NAME>.
# Modeling wine preferences by data mining from physicochemical properties.
# In Decision Support Systems, Elsevier, 47(4):547-553. ISSN: 0167-9236.
# ```
#
#
# ## Methode und Details der Daten
#
#
# - Import der Module
# - Laden der Daten
#
#
#
# Import der Module
#
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn.ensemble import RandomForestRegressor
from sklearn.neural_network import MLPRegressor
from sklearn import metrics
#
# Laden der Daten aus einem CSV File. Der Separator ist hier ein ';'
#
df = pd.read_csv('data/winequality/winequality-red.csv', sep=';')
#
# Anzeige der Dimensionen des Datensatzes
#
print(df.shape)
#
# Anzeige der ersten Datensätze für Kontrolle
#
df.head(20)
df.tail()
#
# Labels werden in x gespeichert, die restlichen Daten in x (ohne quality). Drop löscht ein Feature
#
y_complete = df['quality']
x_complete = df.drop(['quality'], axis=1)
y_complete.head()
x_complete.head()
#
# Aufteilen der Daten in Training Daten und Testdaten
# Beachte auch die neue Schreibweise bei der Zuweisung eines Rückgabewertes der Funktion
#
x_train, x_test, y_train, y_test = train_test_split ( x_complete, y_complete, train_size=0.8, random_state=42 )
x_train.shape
#
# Anlegen eines Modelles für lineare Regression
# Training des Modelles mit Daten (fit)
#
regressor = LinearRegression()
regressor.fit(x_train,y_train)
#
# Kurzer Blick auf die Parameter des Modelles
#
print(regressor.coef_)
#
# Test durch Vorhersage mit dem Modell auf beiden Datensätzen (test und train)
#
prediction_train = regressor.predict(x_train)
prediction_test = regressor.predict(x_test)
prediction_train
#
# Auswertungen der Qualität des Modelles für Regression
# Unterschied zwischen test und train Qualität
#
print('test root mean squared error: {}'.format(np.sqrt(metrics.mean_squared_error(y_test, prediction_test))))
print('train root mean squared error: {}'.format(np.sqrt(metrics.mean_squared_error(y_train, prediction_train))))
#
# Auswertung weiterer Qualitätsparameter für test
#
print('test mean absolute error: {}'.format(metrics.mean_absolute_error(y_test, prediction_test)))
print('test mean squared error: {}'.format(metrics.mean_squared_error(y_test, prediction_test)))
#
# Hilfsfunktion zum zählen
#
def countAccuracy(prediction,y):
prediction_quality_test = np.round_(prediction)
y_test_data = y.values
correct, incorrect = 0,0
for index in range(prediction_test.shape[0]):
if prediction_quality_test[index] == y_test_data[index]:
correct= correct + 1
else:
incorrect= incorrect + 1
print('count accuracy: {}'.format((correct/(correct+incorrect))))
#
# Jetzt Aufruf der Funktion
# Accuracy für Test Daten
#
countAccuracy(prediction_test,y_test)
#
# Accuracy für Training Daten
#
countAccuracy(prediction_train,y_train)
#
# Test eines anderen Modelles für Regression (randomforest regression)
#
random_regressor = RandomForestRegressor(n_estimators = 10, random_state = 42)
random_regressor.fit(x_train, y_train);
# +
prediction_train = random_regressor.predict(x_train)
prediction_test = random_regressor.predict(x_test)
print('test root mean squared error: {}'.format(np.sqrt(metrics.mean_squared_error(y_test, prediction_test))))
print('train root mean squared error: {}'.format(np.sqrt(metrics.mean_squared_error(y_train, prediction_train))))
# -
countAccuracy(prediction_test,y_test)
# +
#
# Wie könnten wir Qualität in diesem Kontext breiter definieren?
#
# -
#
# Hilfsfunktion zum zählen
#
def countAccuracyRelaxed(prediction,y):
prediction_quality_test = np.round_(prediction)
y_test_data = y.values
correct, incorrect = 0,0
for index in range(prediction_test.shape[0]):
if prediction_quality_test[index] == y_test_data[index]:
correct= correct + 1
elif prediction_quality_test[index] == y_test_data[index] + 1:
# wir betrachten die Qualität auch als richtig geschätzt, wenn es die nächsten oder vorherige Klasse war
correct= correct + 1
elif prediction_quality_test[index] == y_test_data[index] - 1:
correct= correct + 1
else:
incorrect= incorrect + 1
print('count accuracy: {}'.format((correct/(correct+incorrect))))
countAccuracyRelaxed(prediction_test,y_test)
# # Test eines Neuronalen Netzwerkes
nn_regressor = MLPRegressor(hidden_layer_sizes=(20,40,10), random_state=42, max_iter=2000, activation='relu')
nn_regressor.fit(x_train, y_train);
# +
prediction_train = nn_regressor.predict(x_train)
prediction_test = nn_regressor.predict(x_test)
print('test root mean squared error: {}'.format(np.sqrt(metrics.mean_squared_error(y_test, prediction_test))))
print('train root mean squared error: {}'.format(np.sqrt(metrics.mean_squared_error(y_train, prediction_train))))
# -
countAccuracy(prediction_test,y_test)
countAccuracyRelaxed(prediction_test,y_test)
| 02 Anwendungsbeispiel Regression Wine Quality.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Confidence-aware UBCF MultiEval Example
# Much is this structure and organization is borrowed from the Lenskit sample evaluation walkthrough
# ## Setup
import sys # set path of locally install lenskit_confidence module
sys.path.insert(0,'C:\\Users\\Name\\Documents\\GitHub\\lenskit_confidence') # Looks like this on my machine
# +
from lenskit.metrics import predict
import pandas as pd
import matplotlib.pyplot as plt
from lenskit.batch_ca import MultiEval
from lenskit.algorithms_ca import user_knn_ca, Recommender # *not* user_knn
from lenskit import topn, datasets, batch_ca # *not* batch
from lenskit import datasets
from lenskit.datasets import MovieLens
from lenskit import crossfold as xf
from lenskit import topn, util #, metrics
from lenskit.crossfold import partition_users, SampleN
# -
# Setting up a progress bar...
from tqdm.notebook import tqdm_notebook as tqdm
tqdm.pandas()
# Setup logging to the notebook...
util.log_to_notebook()
# Pick a dataset to run...
data = MovieLens('../data/ml-1m')
#data = MovieLens('../data/ml-10m')
#data = MovieLens('../data/ml-20m')
#data = MovieLens('../data/jester') # with Jester cleaning, it's the same format a ML datasets, so the ML input function works
# ## Experiment
# Run experiment and store output in the `my-eval` directory.
#
# We're not producing prediction, generating 10-item recommendation lists, and setting up 4 workers.
eval = MultiEval('my-eval', predict = False, recommend = 10, eval_n_jobs = 4)
# We'll use 5-fold CV, partitioning users and putting 5 ratings per user in the test set.
pairs = list(partition_users(data.ratings, 5, SampleN(5)))
# Add the dataset to MultiEval with `add_datasets`.
eval.add_datasets(pairs, name = 'ML1M') # give the added dataset a name
nhbr_range = [25] # We'll use just K=25 for our sample evaluation [10, 25, 50, 75]
# Add the algorithms to MultiEval with `add_algorithms`; the three CUBCF options are listed
eval.add_algorithms([user_knn_ca.UserUserCA(nnbrs = f, aggregate = 'average',
variance_estimator = 'standard-deviation-average') for f in nhbr_range],
attrs = ['nnbrs'], name = 'UserKNN-CA-Average')
eval.add_algorithms([user_knn_ca.UserUserCA(nnbrs = f, aggregate = 'average',
variance_estimator = 'standard-deviation-jackknife-average') for f in nhbr_range],
attrs = ['nnbrs'], name = 'UserKNN-CA-JK-Average')
eval.add_algorithms([user_knn_ca.UserUserCA(nnbrs = f, aggregate = 'average',
variance_estimator = 'standard-deviation-bootstrap-average') for f in nhbr_range],
attrs = ['nnbrs'], name = 'UserKNN-CA-BS-Average')
# Run the experiment...
eval.run(progress = tqdm)
# ## Analyzing Results
#
# We need to read in experiment outputs.
#
# First the run metadata:
runs = pd.read_csv('my-eval/runs.csv')
runs.set_index('RunId', inplace = True)
runs.head() # a quick visual check
# This describes each run - a data set, partition, and algorithm combination. To evaluate, we need to get the actual recommendations, and combine them with this:
recs = pd.read_parquet('my-eval/recommendations.parquet')
recs.head()
recs['score'] = recs['prediction']
recs = recs[['item', 'score', 'user','rank','RunId']]
recs.head()
# Getting the predictions... (this is here for posterity, we're not actually making predictions on test set now)
# +
#preds = pd.read_parquet('my-eval/predictions.parquet')
#preds
# -
# We're going to compute per-(run,user) evaluations of the recommendations *before* combining with metadata.
#
# In order to evaluate the recommendation list, we need to build a combined set of truth data. Since this is a disjoint partition of users over a single data set, we can just concatenate the individual test frames:
truth = pd.concat((p.test for p in pairs), ignore_index = True)
truth.head()
truth.to_csv('my-eval/truth.csv') # saving truth values to a csv for future evaluation
#truth = pd.read_csv('my-eval/truth.csv')
truth = truth[['user', 'item', 'rating']] # just grabbing what we need
truth.head() # a visual check
# Now we can set up an analysis and compute the results.
rla = topn.RecListAnalysis()
rla.add_metric(topn.ndcg) # precision, recall, recip_rank, dcg, ndcg
rla.add_metric(topn.precision)
topn_compute = rla.compute(recs, truth)
topn_compute.head()
# Next, we need to combine this with our run data, so that we know what algorithms and configurations we are evaluating:
topn_results = topn_compute.join(runs[['name', 'nnbrs']], on = 'RunId') #
topn_results.head()
# We can compute the overall average performance for each algorithm configuration
topn_results.fillna(0).groupby(['name', 'nnbrs'])['ndcg','precision'].mean()
| examples/Confidence-aware-UBCF-MultiEval.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from os import listdir
from numpy import array
from keras.preprocessing.text import Tokenizer, one_hot
from keras.preprocessing.sequence import pad_sequences
from keras.models import Model
from keras.utils import to_categorical
from keras.layers import Embedding, TimeDistributed, RepeatVector, LSTM, concatenate , Input, Reshape, Dense, Flatten
from keras.preprocessing.image import array_to_img, img_to_array, load_img
from keras.applications.inception_resnet_v2 import InceptionResNetV2, preprocess_input
import numpy as np
# +
# Load the images and preprocess them for inception-resnet
images = []
all_filenames = listdir('images/')
all_filenames.sort()
for filename in all_filenames:
images.append(img_to_array(load_img('images/'+filename, target_size=(299, 299))))
images = np.array(images, dtype=float)
images = preprocess_input(images)
# Run the images through inception-resnet and extract the features without the classification layer
IR2 = InceptionResNetV2(weights='imagenet', include_top=False)
features = IR2.predict(images)
# +
# We will cap each input sequence to 100 tokens
max_caption_len = 100
# Initialize the function that will create our vocabulary
tokenizer = Tokenizer(filters='', split=" ", lower=False)
# Read a document and return a string
def load_doc(filename):
file = open(filename, 'r')
text = file.read()
file.close()
return text
# Load all the HTML files
X = []
all_filenames = listdir('html/')
all_filenames.sort()
for filename in all_filenames:
X.append(load_doc('html/'+filename))
# Create the vocabulary from the html files
tokenizer.fit_on_texts(X)
# Add +1 to leave space for empty words
vocab_size = len(tokenizer.word_index) + 1
# Translate each word in text file to the matching vocabulary index
sequences = tokenizer.texts_to_sequences(X)
# The longest HTML file
max_length = max(len(s) for s in sequences)
# Intialize our final input to the model
X, y, image_data = list(), list(), list()
for img_no, seq in enumerate(sequences):
for i in range(1, len(seq)):
# Add the entire sequence to the input and only keep the next word for the output
in_seq, out_seq = seq[:i], seq[i]
# If the sentence is shorter than max_length, fill it up with empty words
in_seq = pad_sequences([in_seq], maxlen=max_length)[0]
# Map the output to one-hot encoding
out_seq = to_categorical([out_seq], num_classes=vocab_size)[0]
# Add and image corresponding to the HTML file
image_data.append(features[img_no])
# Cut the input sentence to 100 tokens, and add it to the input data
X.append(in_seq[-100:])
y.append(out_seq)
X, y, image_data = np.array(X), np.array(y), np.array(image_data)
# +
# Create the encoder
image_features = Input(shape=(8, 8, 1536,))
image_flat = Flatten()(image_features)
image_flat = Dense(128, activation='relu')(image_flat)
ir2_out = RepeatVector(max_caption_len)(image_flat)
language_input = Input(shape=(max_caption_len,))
language_model = Embedding(vocab_size, 200, input_length=max_caption_len)(language_input)
language_model = LSTM(256, return_sequences=True)(language_model)
language_model = LSTM(256, return_sequences=True)(language_model)
language_model = TimeDistributed(Dense(128, activation='relu'))(language_model)
# Create the decoder
decoder = concatenate([ir2_out, language_model])
decoder = LSTM(512, return_sequences=False)(decoder)
decoder_output = Dense(vocab_size, activation='softmax')(decoder)
# Compile the model
model = Model(inputs=[image_features, language_input], outputs=decoder_output)
model.compile(loss='categorical_crossentropy', optimizer='rmsprop')
# -
# Train the neural network
model.fit([image_data, X], y, batch_size=64, shuffle=False, epochs=2)
# map an integer to a word
def word_for_id(integer, tokenizer):
for word, index in tokenizer.word_index.items():
if index == integer:
return word
return None
# generate a description for an image
def generate_desc(model, tokenizer, photo, max_length):
# seed the generation process
in_text = 'START'
# iterate over the whole length of the sequence
for i in range(900):
# integer encode input sequence
sequence = tokenizer.texts_to_sequences([in_text])[0][-100:]
# pad input
sequence = pad_sequences([sequence], maxlen=max_length)
# predict next word
yhat = model.predict([photo,sequence], verbose=0)
# convert probability to integer
yhat = np.argmax(yhat)
# map integer to word
word = word_for_id(yhat, tokenizer)
# stop if we cannot map the word
if word is None:
break
# append as input for generating the next word
in_text += ' ' + word
# Print the prediction
print(' ' + word, end='')
# stop if we predict the end of the sequence
if word == 'END':
break
return
# Load and image, preprocess it for IR2, extract features and generate the HTML
test_image = img_to_array(load_img('images/89.jpg', target_size=(299, 299)))
test_image = np.array(test_image, dtype=float)
test_image = preprocess_input(test_image)
test_features = IR2.predict(np.array([test_image]))
generate_desc(model, tokenizer, np.array(test_features), 100)
| local/HTML/HTML.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Tutorial36
# language: python
# name: tutorial
# ---
# # Python Tutorial
# **<NAME>**, Phd Student in Computer Science - ELTE university. Visiting researcher, Aalto university
# ## Agenda
# 1. Motivation
# 3. The Python Programming Language
# # Why to learn Python?
# There are several programming languages... why should you care about learning Python?
# 
# ## 1.- It is widely used in open source projects
# 
# 
# 
# https://octoverse.github.com/
# ## 2.- Relevant open source databases and big data frameworks have a Python SDK
# * PostgreSQL - Psycopg2
# * Apache Spark - pyspark
# * Apache Flink - flink
# * Apache Beam - apache-beam
# * Elasticsearch - elasticsearch
# * Apple's Turi Create - turi-create
# ## 3.- Cloud providers have python SDK too!
# - Amazon Web Services - boto3
# - Microsoft Azure - azure-sdk-for-python
# - Google Cloud - Several python libraries
# ## 4.- The python ecosystem has many libraries that are publicly available
# Popular packages related to data analysis and data mining:
# - **Pandas** - Povides high-performance, easy-to-use data structures and data analysis tools/
# - **Matplotlib** - Plotting library which produces publication quality figures in a variety of hardcopy formats and interactive environments across platforms.
# - **SciPy** - For mathematics, science, and engineering.
# - **NumPy** - Fundamental package for scientific computing with Python. It contains among other things: a powerful N-dimensional array object; sophisticated (broadcasting) functions; tools for integrating C/C++ and Fortran code; useful linear algebra, Fourier transform, and random number capabilities.
# - **TensorFlow** - An open-source machine learning framework for everyone.
# - **Scikit-learn** - Simple and efficient tools for data mining and data analysis.
#
# Popular packages related to web development and crawling:
# - **Django** - A high-level Python Web framework that encourages rapid development and clean, pragmatic design.
# - **Flask** - Micro web framework written in Python.
# - **Uvicorn** - Lightning-fast asyncio server, for Python 3.
# - **Beautiful Soup** - For parsing HTML and XML documents.
# - **Scrapy** - A Fast and Powerful Scraping and Web Crawling Framework
# ## 5.- Platform independent
# - As long as you have a python enviroment and the dependencies are installed in the OS
# - Mobile applications development possible using kiwi or toga
# # So what is Python?
# Reference material: Python Essential Reference (4th Edition) by <NAME>
# - Python is an interpreted high-level programming language for general-purpose programming.
# - Python programs are executed by an interpreter
# - Usually: **python**
# - Often, we use a Python Development Enviroment to simplify the development of Python programs.
# - For example: PyCharm, Jupyter Notebook, Spyder
# - Yes! We can write and run Python in Jupyter Notebook :)
# - There are different versions of python. The most popular are: 2.7, 3.5, 3.6
# - It is suggested to use python 3. Python 2.7 will be depreciated in the year **2020**
# #### Pyton at a Glance
print("Hello World!")
# #### Python is interactive
print(10*3)
# #### Variables
scholarship = 1000 # I am a comment and I just create a variable called balance that contains the number 1000
save_pct = .0 # This is a float
studies_duration = 24 # This is an int
save_each = 3
scholarship
# #### Conditionals
if save_pct <= 0: # Possible conditions: <, <=, ==, >, >=, !=, is, not
print("No savings :-O")
elif save_pct <= .02:
print("Seriously?")
else:
print("Looking good B-)")
# #### Functions
# - The function help is useful to know more about any function
help(help)
help(range)
for month in range(0,studies_duration):
print(month)
if month == 5:
print("Fine I get it")
break
# #### Putting all together
total_savings = 0
for month in range(0,studies_duration):
if (month+1) % save_each == 0: # Modulo operation finds the remainder after division of one number by another
amount_to_save = save_pct * scholarship
print(f"Saving {amount_to_save} in month {month}")
total_savings += amount_to_save
print(f"You saved: {total_savings}, great!") # There are multiple ways to format a string
print("You saved: {}, great!".format(total_savings))
print("You saved: %.1f, great!"%(total_savings))
# #### Structures
# ##### Tuples
a_tuple = (0,1,2,3,4,5,6,6)
print(a_tuple)
print(a_tuple[1])
# ##### Lists
a_list = [0,1,2,3,4,5,4,3,2,1]
print(a_list)
a_list.append(0)
print(a_list)
a_list.remove(0)
print(a_list)
# #### List indexing
# First element
print(a_list[0])
# Last element
print(a_list[-1])
# First two elements
print(a_list[:2])
# Last two elements
print(a_list[-2:])
# ##### Dictionaries
a_dict = {"key1": 1, "key2": 2, "key3": 3}
print(a_dict)
a_dict["key_1"] = 2
a_dict["key_2"] = "any value"
a_dict["key_3"] = {"another_dict":"wow"}
# ##### Sets
a_set = set(a_list)
print(a_set)
# #### Functions
def add_numbers(a,b):
return a+b
print(add_numbers(1,2))
# ##### List Comprehension
my_new_list = [l for l in a_list if l > 2]
print(my_new_list)
my_new_list = [l*2 for l in a_list if l > 2]
print(my_new_list)
# ##### Lambda Functions
g = lambda x: x*x
print(g(5))
help(map)
my_list_tuples = [(0,1),(2,3),(4,5)]
print(my_list_tuples)
for i in map(lambda x: x[0] + x[1], my_list_tuples):
print(i)
# ##### I/O operations
# Reading a file
with open("C:/Users/frede/languages.txt","r") as my_text:
for line in my_text:
print(line)
break
# Writting to a file
with open("C:/Users/frede/count.txt","w") as my_counter:
for i in range(0,10):
my_counter.write(str(i))
my_counter.write("\n")
# #### Installing libraries, and managing enviroments
# Options
# 1. Install packages with the package manager globally as a super user or append --user to install in the home directory
# - Using **pip**
# - sudo pip install pandas
# - sudo pip install --upgrade pandas
# - sudo pip uninstall pandas
# - It is not recommended if you are working in multiple projects in the same machine
# 2. Install the interpreter in your operating system
# - sudo pip install virtualenv
# - Isolate your working enviroments using virtualenv by creating *enviroments*
# - virtualenv -p /usr/bin/python3.6 ~/venv36 # Creates a virtual enviroment using a specific version of python
# - source ~/venv36/bin/activate # Activates the enviroment
# - pip install pandas # This would install pandas in the enviroment
# - deactivate
# 3. Use Anaconda/Mini-conda
# - Install Anaconda: https://anaconda.org/anaconda/python
# - conda create -n env36 python=3.6
# - activate env36 or source activate env36 in linux/mac
# - install packages using conda install instead of pip install. Works better in Windows, as it installs any required DLL
# #### Pandas
# *Not part of DMS course but it's useful to learn Pandas*
import pandas as pd
# %matplotlib inline
df_iris = pd.read_csv("iris.data.txt", names=["sepal_length", "sepal_width", "petal_length", "petal_width", "iris_class"])
df_iris.head()
df_iris.groupby("iris_class").plot.density()
df_iris.query("sepal_length > 5.8").head()
df_iris.describe()
df_iris.groupby("iris_class").describe().transpose().to_csv('iris_data_stats.csv')
| python_tutorial/PythonTutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#hide
# default_exp conda
# -
# # Create conda packages
#
# > Pure python packages created from nbdev settings.ini
# +
#export
from fastcore.script import *
from fastcore.all import *
from fastrelease.core import find_config
import yaml,subprocess
from copy import deepcopy
try: from packaging.version import parse
except ImportError: from pip._vendor.packaging.version import parse
_PYPI_URL = 'https://pypi.org/pypi/'
# -
#export
def pypi_json(s):
"Dictionary decoded JSON for PYPI path `s`"
return urljson(f'{_PYPI_URL}{s}/json')
#export
def latest_pypi(name):
"Latest version of `name` on pypi"
return max(parse(r) for r,o in pypi_json(name)['releases'].items()
if not parse(r).is_prerelease and not o[0]['yanked'])
#export
def _pip_conda_meta(name, path):
ver = str(latest_pypi('sentencepiece'))
pypi = pypi_json(f'{name}/{ver}')
info = pypi['info']
rel = [o for o in pypi['urls'] if o['packagetype']=='sdist'][0]
reqs = ['pip', 'python', 'packaging']
# Work around conda build bug - 'package' and 'source' must be first
d1 = {
'package': {'name': name, 'version': ver},
'source': {'url':rel['url'], 'sha256':rel['digests']['sha256']}
}
d2 = {
'build': {'number': '0', 'noarch': 'python',
'script': '{{ PYTHON }} -m pip install . -vv'},
'test': {'imports': [name]},
'requirements': {'host':reqs, 'run':reqs},
'about': {'license': info['license'], 'home': info['project_url'], 'summary': info['summary']}
}
return d1,d2
#export
def _write_yaml(path, name, d1, d2):
path = Path(path)
p = path/name
p.mkdir(exist_ok=True, parents=True)
yaml.SafeDumper.ignore_aliases = lambda *args : True
with (p/'meta.yaml').open('w') as f:
yaml.safe_dump(d1, f)
yaml.safe_dump(d2, f)
#export
def write_pip_conda_meta(name, path='conda'):
"Writes a `meta.yaml` file for `name` to the `conda` directory of the current directory"
_write_yaml(path, name, *_pip_conda_meta(name))
#export
def _get_conda_meta():
cfg,cfg_path = find_config()
name,ver = cfg.get('lib_name'),cfg.get('version')
url = cfg.get('doc_host') or cfg.get('git_url')
reqs = ['pip', 'python', 'packaging']
if cfg.get('requirements'): reqs += cfg.get('requirements').split()
if cfg.get('conda_requirements'): reqs += cfg.get('conda_requirements').split()
pypi = pypi_json(f'{name}/{ver}')
rel = [o for o in pypi['urls'] if o['packagetype']=='sdist'][0]
# Work around conda build bug - 'package' and 'source' must be first
d1 = {
'package': {'name': name, 'version': ver},
'source': {'url':rel['url'], 'sha256':rel['digests']['sha256']}
}
d2 = {
'build': {'number': '0', 'noarch': 'python',
'script': '{{ PYTHON }} -m pip install . -vv'},
'requirements': {'host':reqs, 'run':reqs},
'test': {'imports': [cfg.get('lib_path')]},
'about': {
'license': 'Apache Software',
'license_family': 'APACHE',
'home': url, 'doc_url': url, 'dev_url': url,
'summary': cfg.get('description')
},
'extra': {'recipe-maintainers': [cfg.get('user')]}
}
return name,d1,d2
#export
def write_conda_meta(path='conda'):
"Writes a `meta.yaml` file to the `conda` directory of the current directory"
_write_yaml(path, *_get_conda_meta())
# This function is used in the `fastrelease_conda_package` CLI command.
#
# **NB**: you need to first of all upload your package to PyPi, before creating the conda package.
#export
@call_parse
def fastrelease_conda_package(path:Param("Path where package will be created", str)='conda',
do_build:Param("Run `conda build` step", bool_arg)=True,
build_args:Param("Additional args (as str) to send to `conda build`", str)='',
do_upload:Param("Run `anaconda upload` step", bool_arg)=True,
upload_user:Param("Optional user to upload package to")=None):
"Create a `meta.yaml` file ready to be built into a package, and optionally build and upload it"
write_conda_meta(path)
cfg,cfg_path = find_config()
out = f"Done. Next steps:\n```\`cd {path}\n"""
name,lib_path = cfg.get('lib_name'),cfg.get('lib_path')
out_upl = f"anaconda upload build/noarch/{lib_path}-{cfg.get('version')}-py_0.tar.bz2"
if not do_build:
print(f"{out}conda build .\n{out_upl}\n```")
return
os.chdir(path)
res = run(f"conda build --output-folder build {build_args} {name}")
if 'anaconda upload' not in res:
print(f"{res}\n\Build failed.")
return
upload_str = re.findall('(anaconda upload .*)', res)[0]
if upload_user: upload_str = upload_str.replace('anaconda upload ', f'anaconda upload -u {upload_user} ')
res = run(upload_str)
# To build and upload a conda package, cd to the root of your repo, and then:
#
# fastrelease_conda_package
#
# Or to do things more manually:
#
# ```
# fastrelease_conda_package --do_build false
# # cd conda
# conda build {name}
# anaconda upload $CONDA_PREFIX/conda-bld/noarch/{name}-{ver}-*.tar.bz2
# ```
#
# Add `--debug` to the `conda build command` to debug any problems that occur. Note that the build step takes a few minutes. Add `-u {org_name}` to the `anaconda upload` command if you wish to upload to an organization, or pass `upload_user` to `fastrelease_conda_package`.
#
# **NB**: you need to first of all upload your package to PyPi, before creating the conda package.
# ## Export-
#hide
from nbdev.export import notebook2script
notebook2script()
| 01_conda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
% matplotlib inline
# +
import numpy as np
class LogisticRegression(object):
"""Single Class Multivariate logistic regression model using gradient descent"""
def __init__(self):
pass
def train(self, x, y, epochs=10, learning_rate=0.0001):
self.theta_array = np.zeros(np.array(x.ndim)+1)
x = self._add_bias(x)
for _ in range(1, epochs):
avg_minibatch_partial_grads = np.average(
(self._sigmoid(x, self.theta_array) - y) * x, axis=1)
self.theta_array -= learning_rate * avg_minibatch_partial_grads
def validate(self, x, y):
self._check_theta_exists('validating')
x = self._add_bias(x)
predicted_y = np.dot(x.transpose(), self.theta_array)
rmse = np.sqrt(np.average(np.square(y- predicted_y))) # Root Mean Square Error (RMSE)
return predicted_y, rmse
def predict(self, x):
self._check_theta_exists('predicting')
x = self._add_bias(x)
predicted_y = self._sigmoid(x, self.theta_array)
return predicted_y
def _add_bias(self, x):
if x.ndim == 1:
x = np.row_stack((x, np.ones(len(x))))
else:
x = np.row_stack((x, np.ones(len(x[0]))))
return x
def _sigmoid(self, x, theta_array):
sigmoid = 1/(1+np.exp(-np.dot(x.transpose(), theta_array)))
return sigmoid
def _avg_minibatch_loss(self, x, theta_array, y):
avg_minibatch_loss = np.sqrt(
np.average(
np.square(
x.transpose().dot(theta_array) - y)))
return avg_minibatch_loss
def _check_theta_exists(self, phrase):
assert hasattr(self, 'theta_array'), ("ValueError: theta is not defined. "
"Please make sure to train the model before {}}.".format(phrase))
# -
logistic_regression = LogisticRegression()
# +
import math
x = np.array(np.arange(100))
delta = np.random.uniform(-10,10, size=(10,))
y = np.concatenate((np.array(np.zeros(int(len(x)/2))), np.array(np.ones(int(len(x)/2)))))
# -
logistic_regression.train(x, y, epochs=100000, learning_rate=0.1)
y_predicted = logistic_regression.predict(x)
# +
import matplotlib.pyplot as plt
% matplotlib inline
plt.scatter(x, y)
plt.plot(x, y_predicted, color='red')
plt.show()
# -
| examples/logistic_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src="images/logo_verde_horz.png" width="600">
# + slideshow={"slide_type": "-"}
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# ## Broadcasting
#
# Las operaciones básicas (suma, resta, etc.) se hacen elemento por elemento y funciona en arrays de diferente tamaño.
#
# La imagen siguiente da un ejemplo de **broadcasting**
#
# <img src="images/broadcasting.png">
a = np.array([[0], [10], [20], [30]])
a
b = np.array([0, 1, 2])
b
a + b
# ## Manipulación de formas
#
# ### Colapsar
#
# Reducir una matriz de múltiples dimensiones y la colapsa a una dimensión.
a = np.array([[1, 2, 3], [4, 5, 6]])
a.ravel()
a.T
a.T.ravel()
# ### Reformar
#
# La operación inversa a colapsar.
b = a.ravel()
b
b.reshape((2,3))
b.reshape((3,2))
# ## Manejo de Archivos
#
# ### Tablas
# Demos una mirada al archivo `/data/populations.txt`
#
# |year|hare|lynx|carrot|
# |----|----|----|------|
# |1900|30000| 4000| 48300|
# |1901|47200| 6100| 48200|
# |1902|70200| 9800| 41500|
# |1903|77400| 35200| 38200|
# Leer datos desde archivo
data = np.loadtxt('data/populations.txt')
data
# ## Ejercicio:
# Genere un gráfico para observar el cambio en la población de liebres y de linces a lo largo de los años
# +
# Almacenar datos en formato texto
# Suponga que se ha cometido un error, y los datos de población de linces,
# reportados en miles, se trataba en realidad de centenares. Corrija la
# columna respectiva y almacene la tabla con un nuevo nombre utilizando:
np.savetxt('data/pop2.txt', data, fmt='%5.i', delimiter='\t')
# -
# ### Formato Numpy
#
# Numpy tiene su propio formato binario.
data = np.ones((3,3))
data
np.save('data/ones.npy', data)
data3 = np.load('data/ones.npy')
data3
# ## Resumen
#
# - Sabe crear arrays: `array`, `arange`, `ones`, `zeros`, `rand`
# - Conoce la forma de un array `shape` y los diferentes métodos de indexado para obtener diferente secciones del array `array[::2]`, etc.
# - Ajustar la forma de un array usando `reshape` o aplanarla con `ravel`
# - Obtener un subconjunto de elementos de un array y modificar sus valores usando mascaras `a[a < 0] = 0`
# - Conocer operaciones miscelanes con arrays, encontrar el máximo `array.max()` o la media `array.mean()`, entre otras.
# - Tiene una idea básica del **broadcasting** y sus reglas.
#
# ## Análisis de Vinos
#
# Usando los resultados de un análisis químico de vinos obtenidos de la misma región en Italia pero de tres diferentes cultivos. Examine el archivo `data/wine.csv`.
#
# Los atributos del dataset son:
#
# 0. Wine (Class)
# 1. Alcohol
# 2. Malic acid
# 3. Ash
# 4. Alcalinity of ash
# 5. Magnesium
# 6. Total phenols
# 7. Flavanoids
# 8. Nonflavanoid phenols
# 9. Proanthocyanins
# 10. Color intensity
# 11. Hue
# 12. OD280/OD315 of diluted wines
# 13. Proline
#
# ---
# #### Truco:
# Use `np.set_printoptions(suppress=True, precision=3)` para imprimer los datos de una manera mas legible.
#
#
# ### Ejercicios
#
# 1. Lea los datos desde la ruta `data/wine.csv`, use el parámetro `delimiter=','` para valores separados por coma.
# 2. Determine entre las 3 clases de vinos, cual tiene el mayor promedio de Alcohol.
# 3. Un vino con concentración de ácido málico mayor a `3.5` se considera demasiado maduro, cuántos vinos tienen una concentración mayor?
# 4. Usando el comando `plt.hist(array)` cree un histograma de ácido málico.
# 5. Normalice entre [0,1] los datos de Alcohol e Intensidad de Color
# 6. Usando el comando `plt.scatter(array1, array2)` cree un gráfico de dispersión de las dos columnas normalizadas.
np.set_printoptions(suppress=True, precision=3)
| Notebooks/nb_10_numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# preprocessing data
'''
Ri= The number of product i produced in regular time in 1000s
Oi= The number of product i produced in overtime in 1000s
Adi= The amount of advertisement expenditure for product i in $1000s
MAX :
(120-66)*(RT+OT)+(150-85)*(RF+OF)+(100-50)*(RC+OC)+(160-80)*(RP+OP)-16*(RT+RF)-18*(OT+OF)-12*(RC+RP)-15*(OC+OP)-Adt-AdF-AdP-AdC
= 38RT+49RF+38RC+68RP+36OT+47OF+35OC+65OP-Adt-AdF-AdC-AdP
Constrint:
Production Capacity in Each Department:
RT+RF<=100
RC+RP<=190
OT+OF<=25
OC+OP<=24
Advertisement Limits
AdT+AdF+AdC+AdP<=18
AdT, AdF, AdC, AdP<=10
RT+OT<=60+60*0.12*AdT/10
RF+OF<=20+20*0.1*AdF/10
RC+OC<=100+100*0.08*AdC/10
RP+OP<=35+35*0.15*AdP/10
Ri, Oi, Adi>= for all i
'''
# objective function coefficientsfrom scipy.optimize import linprog
# 38RT+49RF+38RC+68RP+36OT+47OF+35OC+65OP-Adt-AdF-AdC-AdP
obj = [-38,-49,-38,-68,-36,-47,-35,-65,1, 1, 1, 1]
# ─┬ ─┬ ─┬ ─┬ ─┬ ─┬ ─┬ ─┬ ─┬ ─┬ ─┬ ─┬
# │ │ │ │ │ │ | | | | | └┤ Coefficient for AdP
# │ │ │ │ │ | | | | | └┤ Coefficient for AdC
# │ │ │ │ | | | | | └┤ Coefficient for AdF
# │ │ │ | | | | | └┤ Coefficient for AdT
# │ │ | | | | | └┤ Coefficient for OP
# │ | | | | | └┤ Coefficient for OC
# │ | | | | └┤ Coefficient for OF
# │ | | | └┤ Coefficient for OT
# │ | | └┤Coefficient for RP
# │ | └┤ Coefficient for RC
# │ └┤ Coefficient for RF
# └┤ Coefficient for RT
lhs_ineq = [[1,1,0,0,0,0,0,0,0,0,0,0], # RT+RF
[0,0,1,1,0,0,0,0,0,0,0,0], # RC+RP
[0,0,0,0,1,1,0,0,0,0,0,0], # OT+OF
[0,0,0,0,0,0,1,1,0,0,0,0], # OC+OP
[0,0,0,0,0,0,0,0,1,1,1,1], # AdT+AdF+AdC+AdP
[1,0,0,0,1,0,0,0,0,0,0,0],
[0,1,0,0,0,1,0,0,0,0,0,0],
[0,0,1,0,0,0,1,0,0,0,0,0],
[0,0,0,1,0,0,0,1,0,0,0,0],
[0,0,0,0,0,0,0,0,1,0,0,0], # AdT
[0,0,0,0,0,0,0,0,0,1,0,0], # AdF
[0,0,0,0,0,0,0,0,0,0,1,0], # AdC
[0,0,0,0,0,0,0,0,0,0,0,1]] # AdP
rhs_ineq = [100, # Production Capacity
190, # Production Capacity
25, # Production Capacity
24, # Production Capacity
18, # Advertisement Limits
60, # Advertisement Limits
21, # Advertisement Limits
106.4, # Advertisement Limits
40.25, # Advertisement Limits
10,
10,
10,
10]
# lhs_ineq <= rhs_ineq
bnd = [(0, float("inf")),
(0, float("inf")),
(0, float("inf")),
(0, float("inf")),
(0, float("inf")),
(0, float("inf")),
(0, float("inf")),
(0, float("inf")),
(0, float("inf")),
(0, float("inf")),
(0, float("inf")),
(0, float("inf"))]
opt = linprog(c=obj, A_ub=lhs_ineq, b_ub=rhs_ineq,
# A_eq=lhs_eq, b_eq=rhs_eq,
# bounds=bnd,
method="revised simplex")
opt
# +
# preprocessing data
'''
Ri= The number of product i produced in regular time in 1000s
Oi= The number of product i produced in overtime in 1000s
Adi= The amount of advertisement expenditure for product i in $1000s
MAX :
(120-66)*(RT+OT)+(150-85)*(RF+OF)+(100-50)*(RC+OC)+(160-80)*(RP+OP)-16*(RT+RF)-18*(OT+OF)-12*(RC+RP)-15*(OC+OP)-Adt-AdF-AdP-AdC
= 38RT+49RF+38RC+68RP+36OT+47OF+35OC+65OP-Adt-AdF-AdC-AdP
Constrint:
Production Capacity in Each Department:
RT+RF<=100
RC+RP<=90
OT+OF<=25
OC+OP<=24
Advertisement Limits
AdT+AdF+AdC+AdP<=18
AdT, AdF, AdC, AdP<=10
RT+OT<= 67.2
RF+OF<= 20
RC+OC<= 100
RP+OP<= 40.32
Ri, Oi, Adi>= for all i
'''
# objective function coefficientsfrom scipy.optimize import linprog
# 38RT+49RF+38RC+68RP+36OT+47OF+35OC+65OP-Adt-AdF-AdC-AdP
from scipy.optimize import linprog
obj = [-38,-49,-38,-68,-36,-47,-35,-65, 1, 1, 1, 1]
# ─┬ ─┬ ─┬ ─┬ ─┬ ─┬ ─┬ ─┬ ─┬ ─┬ ─┬ ─┬
# │ │ │ │ │ │ | | | | | └┤ Coefficient for AdP
# │ │ │ │ │ | | | | | └┤ Coefficient for AdC
# │ │ │ │ | | | | | └┤ Coefficient for AdF
# │ │ │ | | | | | └┤ Coefficient for AdT
# │ │ | | | | | └┤ Coefficient for OP
# │ | | | | | └┤ Coefficient for OC
# │ | | | | └┤ Coefficient for OF
# │ | | | └┤ Coefficient for OT
# │ | | └┤Coefficient for RP
# │ | └┤ Coefficient for RC
# │ └┤ Coefficient for RF
# └┤ Coefficient for RT
lhs_ineq = [[1,1,0,0,0,0,0,0,0,0,0,0], # RT+RF
[0,0,1,1,0,0,0,0,0,0,0,0], # RC+RP
[0,0,0,0,1,1,0,0,0,0,0,0], # OT+OF
[0,0,0,0,0,0,1,1,0,0,0,0], # OC+OP
[0,0,0,0,0,0,0,0,1,1,1,1], # AdT+AdF+AdC+AdP
[1,0,0,0,1,0,0,0,0,0,0,0],
[0,1,0,0,0,1,0,0,0,0,0,0],
[0,0,1,0,0,0,1,0,0,0,0,0],
[0,0,0,1,0,0,0,1,0,0,0,0],
[0,0,0,0,0,0,0,0,1,0,0,0], # AdT
[0,0,0,0,0,0,0,0,0,1,0,0], # AdF
[0,0,0,0,0,0,0,0,0,0,1,0], # AdC
[0,0,0,0,0,0,0,0,0,0,0,1]] # AdP
rhs_ineq = [100, # Production Capacity
90, # Production Capacity
25, # Production Capacity
24, # Production Capacity
18, # Advertisement Limits
67.2, # Advertisement Limits
20, # Advertisement Limits
100, # Advertisement Limits
40.32, # Advertisement Limits
10,
10,
10,
10]
# lhs_ineq <= rhs_ineq
bnd = [(0, float("inf")),
(0, float("inf")),
(0, float("inf")),
(0, float("inf")),
(0, float("inf")),
(0, float("inf")),
(0, float("inf")),
(0, float("inf")),
(0, float("inf")),
(0, float("inf")),
(0, float("inf")),
(0, float("inf"))]
opt = linprog(c=obj, A_ub=lhs_ineq, b_ub=rhs_ineq,
# A_eq=lhs_eq, b_eq=rhs_eq,
# bounds=bnd,
method="simplex")
opt
| Solution Code/ICE 2 Q6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="YW_sIvEND5Bh"
# A single perceptron network that simulate an OR gate
from sklearn.linear_model import Perceptron
import matplotlib.pyplot as plt
import numpy as np
from itertools import product
# + id="wjCW4iK9EVdx"
# Data set of different logic gates
data = [[0,0],[0,1],[1,0],[1,1]]
labels = [0,1,1,1]
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="Pl4mdIOYEZSc" outputId="c83cb821-d060-4d71-dfb0-fea6f0e24bc1"
# Scatter Plot of Data
x = [i[0] for i in data]
y = [i[1] for i in data]
c = [i for i in labels]
plt.scatter(x,y,c=labels)
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="vdA3xcktEhc6" outputId="b5aa150d-3c4f-4872-aae9-54dc3faf59ae"
# ML Model
classifier = Perceptron(max_iter=40)
classifier.fit(data, labels)
# Get score
print(classifier.score(data, labels))
# Decision Function
print(classifier.decision_function([[0, 0], [1, 1], [0.5, 0.5]]))
# + id="oGar1F7LEmCe"
# Set Up Heat Map
x_values = np.linspace(0,1,100)
y_values = np.linspace(0,1,100)
point_grid = list(product(x_values, y_values))
distances = classifier.decision_function(point_grid)
abs_distances = [abs(i) for i in distances]
abs_distances_2d = np.reshape(abs_distances, (100,100))
# + colab={"base_uri": "https://localhost:8080/", "height": 269} id="IaM7BzCCEs9X" outputId="e8bb3a09-6635-4c92-8abc-41b546b94a63"
# Draw Map
heatmap = plt.pcolormesh(x_values, y_values, abs_distances_2d)
plt.colorbar(heatmap)
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="W9rrvOPaEyfa" outputId="8c16e27d-3e1e-4e1a-9b4e-d0e01b343347"
# predict
x_test = [[0,0],[1,1],[1,0],[1,0],[0,0]]
y_test = [0,1,1,1,0]
print(classifier.predict(x_test))
print(classifier.score(x_test, y_test))
| deep_learning/logic_simulation/perceptron/single_perceptron.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data visualization and Exploratory Data Analysis (EDA)
#
# In this notebook we'll go over the basics when it comes to data analysis. After getting your data (see the previous
# notebook for an example on how to do that), you are encouraged to explore the dataset for any interesting phenomena.
#
# This process is called Exploratory Data Analysis (EDA) and comprises a lot of different techniques. All of them, in the
# end, are there to provide us, the data scientists, a means to visually detect interesting behaviour in the data.
#
# The two plots generated here are just examples, and you should figure out yourself what interesting data you want to
# look at. For more information on how to create neat plots, see [this](https://seaborn.pydata.org/introduction.html) link.
#
# ### Loading data
# +
import pandas as pd
pd.set_option('display.max_columns', None)
data = pd.read_csv(r"../data/owid-covid-data.csv")
data = data.sort_values('date', ascending=True)
data['date'] = pd.to_datetime(data['date'], format = '%Y-%m-%d')
# -
# ## Scatter plot
# Scatter plots are a good start when you are trying to see if there is some kind of structure to your data. For example,
# your data might be clustered in certain regions or, like in this example, show a trend over time. In the case of the
# latter you could follow it up with a simple line graph (maybe with also averaging the data points per day to reduce
# visual clutter).
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sb
import matplotlib.ticker as ticker
plt.style.use('bmh') # Set the style from a preset of styles
fig = plt.subplots(figsize=(12, 8)) # Create figure and set size
ax = sb.scatterplot(data=data, x='date', y='total_deaths', hue='continent') # Create a scatter plot total deaths by date per continent
ax.set_title('Scatterplot total deaths per continent', size=20) # Set title
ax.set_ylim(ymin=0) # Set starting value of y-axis to 0 - there are no negative deaths
ax.xaxis.set_major_locator(ticker.MultipleLocator(10)) # Set density of x-axis tickers
plt.legend(loc='upper left') # Show legend on the upper left size
plt.setp(ax.get_xticklabels(), rotation=90) # Rotate x-axis tickers 90degrees so that they are readible
print() # removes weird array output filled with 'None's
# -
# There's far too many countries to visualize them in a scatter plot clearly. To increase the likelihood of interesting
# findings and also data interpretability you might want to reduce the search space to a specific continent or even a
# specific country.
#
# In the following code segment we reduce the search to Europe, for no particular reason other than it being the place we
# live.
#
# For more information on using EDA using Pandas, see [this](https://towardsdatascience.com/exploratory-data-analysis-eda-visualization-using-pandas-ca5a04271607)
# link.
# +
# Creating a new dataframe with the rows that correspond to European countries by selecting the continent
# To be able to do this, some indexing is necessary. So save the data with index_col='continent' so that Europe can be used by .loc() method
data_indexed = pd.read_csv(r"../data/owid-covid-data.csv", index_col='continent')
data_europe = data_indexed.loc['Europe']
# + [markdown] pycharm={"name": "#%% md\n"}
# This data can then be used, just like before, to only plot information for this continent, which is what we will do in
# the next section.
#
# ## Bar plot
# Another example of an interesting plot is the bar plot. In the following segment we are looking at a breakdown of the
# number of tests performed per European country.
#
# Some preprocessing, in particular the removal of empty rows, is done before plotting.
# +
# %matplotlib inline
data_europe = data_europe.sort_values('population', ascending=True)
# Remove NAN values in the dataframe
data_europe = data_europe[data_europe['population'].notna()]
data_europe = data_europe[data_europe['total_tests'].notna()]
with plt.style.context('dark_background'):
f, ax = plt.subplots(figsize = (10,15))
sb.barplot(x = 'population', y = 'location', data = data_europe,
label = 'Population', color = 'b', edgecolor = 'w')
sb.set_color_codes('muted')
sb.barplot(x = 'total_tests', y = 'location', data = data_europe,
label = 'Total Tests', color = 'white', edgecolor = 'w')
ax.legend(ncol = 2, loc = 'upper right')
ax.set(xlabel='Population and tests', ylabel='Country')
sb.despine(left = True, bottom = True)
plt.show()
# -
# As expected, countries with higher population counts tested more. What is even more interesting is that Germany tested
# roughly the same amount of people as Spain even though its population is double the amount of Spain's. The same phenomena
# is observed for Russia - having tested the same amount roughly as the United Kingdom but having more than twice their
# population.
#
# ## Conclusion
# By looking at these and other possible graphs you can try to find interesting tidbits of info, hidden in the data. Then,
# you can test theories you've created based on said data and try to make statements about the processes that underlie the
# data.
#
# For more info on that, see our third example notebook.
| notebooks/Example 2 - Exploratory Data Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# <img src="images/utfsm.png" alt="" width="100px" align="right"/>
# # USM Numérica
# ## Licencia y configuración del laboratorio
# Ejecutar la siguiente celda mediante *`Ctr-S`*.
# +
"""
IPython Notebook v4.0 para python 3.0
Librerías adicionales:
Contenido bajo licencia CC-BY 4.0. Código bajo licencia MIT.
(c) <NAME>, <NAME>, <NAME>, <NAME>.
"""
# Configuración para recargar módulos y librerías dinámicamente
# %reload_ext autoreload
# %autoreload 2
# Configuración para graficos en línea
# %matplotlib inline
# Configuración de estilo
from IPython.core.display import HTML
HTML(open("./style/style.css", "r").read())
# -
# ## Introducción a BASH
# Antes de comenzar debemos saber que Bash es un programa informático usado como intérprete de comandos
# o instrucciones dadas por un usuario, las cuales son escritas en alguna interfaz gráfica o comúnmente
# una terminal. Esas instrucciones son interpretadas por Bash para luego enviar dichas órdenes al Núcleo
# o Kernel del sistema operativo.
#
# Cada sistema operativo se encuentra conformado por un Núcleo particular, que se encarga de interactuar
# con la computadora siendo una especie de cerebro capaz de organizar, administrar y distribuir los recursos
# físicos de esta, tales como memoria, procesador, forma de almacenamiento, entre otros.
#
#
#
# <img src="imbash.png"width="700px">
#
#
#
# Bash (Bourne-Again-Shell) es un lenguaje de programación basado en Bourne-Shell, el cual fue creado para
# sistemas Unix en la década de los 70, siendo el sustituto natural y de acceso libre de este a partir del
# año 1987 siendo compatible con la mayoría de sistemas Unix, GNU/Linux y en algunos casos con Microsoft-Windows
# y Apple.
#
# ## Objetivos
# 1. Operaciones básicas para crear, abrir y cambiarse de directorio
# 2. Operaciones para crear un archivo, copiar y cambiarlo de directorio
# 3. Visualizador gráfico de directorios y archivos
# 4. Visualizador de datos y editor de un archivo de texto
# 5. Ejercicio de práctica
#
#
# ### 1. Operaciones para crear, abrir y cambiar de directorio
#
# Este es el tipo de operaciones más básicas que un usuario ejecuta en un sistema operativo, los siguientes comandos nos permiten ubicarnos en alguna carpeta para tener acceso a algún archivo o material en específico, crear carpetas o directorios para almacenar información deseada entre otros.
#
# La acción más simple para comenzar será ingresar a un directorio o carpeta deseada usando el comando *`cd`* como sigue:
# ```
# # cd <directorio>
# ```
# Una extensión de este recurso es la posibilidad de colocar una secuencia de directorios para llegar a la ubicación deseada, separando los nombres por un slash del siguiente modo.
# ```
# # cd <directorio_1>/<subdirectorio_2>/<subdirectorio_3>
# ```
# Podemos visualizar en la terminal el contenido de este directorio con el comando *`ls`* y luego crear un nuevo sub-directorio (o carpeta) dentro del directorio en el cual nos ubicamos con *`mkdir`*:
# ```
# # mkdir <subdirectorio>
# ```
# Mencionamos además una opción con el comando anterior, que nos permite crear varios sub-directorios a la vez escribiendo sus nombres respectivos uno al lado del otro, separados por un espacio.
# ```
# # mkdir <subdirectorio_1> <subdirectorio_2> ... <subdirectorio_N>
# ```
#
# Además como detalle si el nombre de nuestro directorio se conforma por palabras separadas por espacio, entonces conviene por defecto escribir el nombre completo entre comillas, puesto que de lo contrario Bash considerará cada palabra separada por un espacio como un subdirectorio diferente.
# ```
# # mkdir <"nombre subdirectorio">
# ```
#
#
# Si queremos regresar a una ubicación anterior basta los comandos *`cd ..`* o *`cd -`* y si queremos volver al directorio original desde donde se abrió la terminal usamos *`cd ~`*.
#
# Es posible borrar un directorio con su contenido al interior escribiendo el siguiente comando:
# ```Bash
# # rm -r <directorio>
# ```
# Finalmente un comando que nos permite visualizar rápidamente nuestra ubicación actual y las precedentes es *`pwd`*.
# ### 2. Operaciones para crear, copiar y eliminar archivos
# Un paso siguiente a lo visto en el punto anterior es la creación de algún tipo de archivo, realizando operaciones básicas como copiarlo de un directorio a otro, cambiarlo de ubicación o borrarlo.
#
# Para crear un archivo debemos ingresar al directorio en el cual deseamos guardarlo con el comando *`cd`* y luego de esto podemos crear el archivo con el argumento *`>`* de la siguiente manera.
# ```
# > <archivo.tipo>
# ```
# Por defecto el archivo se crea en el directorio actual de nuestra ubicación, recordar que con pwd podemos visualizar la cadena de directorios y subdirectorios hasta la ubicación actual.
#
# Debemos hacer referencia al comando *`echo`*, este consiste en una función interna del intérprete de comandos que nos permite realizar más de una acción al combinarlo de distintas maneras con otros comandos o variables. Uno de los usos más comunes es para la impresión de algún texto en la terminal.
# ```
# # echo <texto a imprimir>
# ```
# También nos permite imprimir un texto en un archivo específico agregando *`echo <texto a imprimir> < <archivo sobre el que se imprime>`*, entre muchas otras opciones que la función *`echo`* nos permite realizar y que es posible profundizar con la práctica, pero estas las postergaremos para la siguiente sección.
#
# Continuamos con el comando *`mv`*, que refiere a "move", el cual sirve para mover algún archivo ya creado a un nuevo directorio.
# ```
# # mv <archivo.tipo> <directorio>
# ```
# También sirve para mover un directorio dentro de otro (mover *directorio_1* al *direcotorio_2*), para que el comando se ejecute correctamente ambos directorios deben estar en una misma ubicación.
# ```
# # mv <directorio_1> <directorio_2>
# ```
# Una operación similar a la anterior es copiar un archivo y llevarlo a un directorio particular, con la diferencia que una vez realizada la acción se tendrán 2 copias del mismo archivo, una en el directorio original y la segunda en el nuevo directorio.
# ```
# # cp <archivo.tipo> <directorio>
# ```
# Supongamos que queremos copiar un archivo existente en otro directorio y reemplazarlo por un archivo del directorio actual, podemos hacer esto de la siguiente manera.
# ```
# # cp ~/directorio_fuente/<archivo_fuente> <archivo_local>
# ```
# Lo anterior se hace desde el directorio al cual deseamos copiar el archivo fuente y *~/directoiro_fuente/* hace alusión al directorio en el cual se encuentra este archivo.
#
# Si por otra parte queremos copiar un archivo fuente y nos encontramos en el directorio en el cual este se encuentra, para realizar la copia en otro directorio, sin necesariamente hacer un reemplazo por otro archivo, se puede con:
# ```
# # cp <archivo_fuente_1> <archivo_fuente_2> ~/directorio_desitno/
# ```
#
# Del mismo modo que para un directorio, si queremos borrar un archivo creado podemos hacerlo con el comando *`rm -r`*.
# ```
# # rm -r <archivo.tipo>
# ```
# Y si queremos borrar una serie de archivos lo hacemos escribiendo consecutivamente.
# ```
# # rm -r <archivo_1.tipo> <archivo_2.tipo> ... <archivo_N.tipo>
# ```
#
# ### 3. Visualizador de estructura de directorios y archivos
# El comando *`tree`* es una forma útil y rápida de visualizar gráficamente la estructura de directorios y archivo pudiendo ver claramente la relación entre estos. Solo debemos escribir el comando para que automáticamente aparezca esta información en pantalla (dentro de la terminal) apareciendo en orden alfabético, por defecto debe ejecutarse ubicándose en el directorio deseado visualizando la estructura dentro de este.
#
# En caso de que este no se encuentre instalado en nuestro sistema operativo, a modo de ejercicio, primero debemos escribir los siguientes comandos.
# ```
# sudo apt-get install <tree>
# ```
#
#
# ### 4. Visualizar, editar y concatenar un archivo de texto
# Para visualizar el contenido de un texto previamente creado, pudiendo hacerlo con el comando visto anteriormente, *`echo > arhcivo.tipo`*, utilizamos el comando *`cat`*.
# ```
# # cat <archivo.tipo>
# ```
# Luego si queremos visualizar varios archivos en la terminal, lo hacemos agregando uno al lado del otro después del comando *`cat`*.
# ```
# # cat <archivo_1.tipo> <archivo_2.tipo> ... <archivo_N.tipo>
# ```
# Existen muchos argumentos que nos permiten visualizar de distinta forma el contenido de un archivo en la terminal, por ejemplo enumerar las filas de algún texto, *`cat -n`*, otra opción sería que solo se enumerara las filas que tienen algún contenido, *`cat -b`*.
#
# En caso de que queramos enumerar solo las filas con texto, pero este tiene demasiadas filas en blanco y buscamos reducirlas a una sola de modo de ahorrar espacio en la terminal, podemos hacerlo agregando el argumento *-s* como sigue.
# ```
# # cat -sb <archivo.tipo>
# ```
# Editar o imprimir un texto en un archivo es posible hacerlo usando la función *`echo`* como sigue.
# ```
# # echo <texto a imprimir> ./archivo.txt
# ```
# Similar a sudo, less es un programa usado como visualizador de archivos de texto que funciona como un comando interpretado desde la terminal. Este permite visualizar completamente el archivo de texto usando por defecto las flechas del teclado para avanzar o retroceder en el visualizador.
#
# Una de las ventajas de un programa como less, es que puede añadirse comandos para ejecutar acciones de forma rápida en modo de comandos que resulta por defecto al ejecutar less, a continuación presentamos algunos comandos básicos.
# ```
# G: permite visualizar el final del texto
# ```
# ```
# g: permite visualizar el inicio del texto
# ```
# ```
# h: nos proporciona ayuda respecto a comandos posibles
# ```
# ```
# q: permite salir de la aplicación dentro del visualizador less
# ```
# Para modificar el texto una de las formas es cargar algún editor de texto como por ejemplo el Visual.
# ```
# v: ejecutar el editor de texto
# ```
# ### 5. Ejercicio de práctica
# Para redondear lo visto en este tutorial se dejara como ejercicio las siguientes instrucciones:
#
# * Crear una carpeta o directorio principal
# * En ella se debe copiar 2 archivos de textos provenientes de cualquier dirección
# * Crear un archivo de texto el cual tenga por nombre "Texto Principal" y se imprima "concatenación de textos"
# * Crear una segunda carpeta dentro de la principal
# * Concatenar los 2 archivos copiados con el archivo creado
# * Mover el archivo "Texto Principal" a la nueva carpeta
# * Eliminar las copias de los archivos concatenados
# * Visualizar con Tree la estructura y relación de archivos y directorios creados
# + language="bash"
| 14_terminal_de_comandos_bash/bash.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# # Alert Investigation - Windows Process Alerts
# <details>
# <summary> <u>Details...</u></summary>
# **Notebook Version:** 1.1<br>
#
# **Data Sources Used**:<br>
# - Log Analytics/Azure Sentinel
# - SecurityAlert
# - SecurityEvent
# <br>
# - Threat Intelligence Providers (Optional)
# - OTX (https://otx.alienvault.com/)
# - VirusTotal (https://www.virustotal.com/)
# - XForce (https://www.ibm.com/security/xforce)
# </details>
#
# This notebook is intended for triage and investigation of security alerts related to process execution. It is specifically targeted at alerts triggered by suspicious process activity on Windows hosts.
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Hunting-Hypothesis" data-toc-modified-id="Hunting-Hypothesis-1"><span class="toc-item-num">1 </span>Hunting Hypothesis</a></span></li><li><span><a href="#Notebook-Initialization" data-toc-modified-id="Notebook-Initialization-2"><span class="toc-item-num">2 </span>Notebook Initialization</a></span><ul class="toc-item"><li><span><a href="#Get-WorkspaceId-and-Authenticate-to-Log-Analytics" data-toc-modified-id="Get-WorkspaceId-and-Authenticate-to-Log-Analytics-2.1"><span class="toc-item-num">2.1 </span>Get WorkspaceId and Authenticate to Log Analytics</a></span></li></ul></li><li><span><a href="#Get-List-of-Alerts" data-toc-modified-id="Get-List-of-Alerts-3"><span class="toc-item-num">3 </span>Get List of Alerts</a></span></li><li><span><a href="#Choose-Alert-to-Investigate" data-toc-modified-id="Choose-Alert-to-Investigate-4"><span class="toc-item-num">4 </span>Choose Alert to Investigate</a></span></li><li><span><a href="#Extract-properties-and-entities-from-Alert" data-toc-modified-id="Extract-properties-and-entities-from-Alert-5"><span class="toc-item-num">5 </span>Extract properties and entities from Alert</a></span></li><li><span><a href="#Entity-Graph" data-toc-modified-id="Entity-Graph-6"><span class="toc-item-num">6 </span>Entity Graph</a></span></li><li><span><a href="#Related-Alerts" data-toc-modified-id="Related-Alerts-7"><span class="toc-item-num">7 </span>Related Alerts</a></span><ul class="toc-item"><li><span><a href="#Show-these-related-alerts-on-a-graph" data-toc-modified-id="Show-these-related-alerts-on-a-graph-7.1"><span class="toc-item-num">7.1 </span>Show these related alerts on a graph</a></span></li><li><span><a href="#Browse-List-of-Related-Alerts" data-toc-modified-id="Browse-List-of-Related-Alerts-7.2"><span class="toc-item-num">7.2 </span>Browse List of Related Alerts</a></span></li></ul></li><li><span><a href="#Get-Process-Tree" data-toc-modified-id="Get-Process-Tree-8"><span class="toc-item-num">8 </span>Get Process Tree</a></span><ul class="toc-item"><li><span><a href="#Process-Time-Line" data-toc-modified-id="Process-Time-Line-8.1"><span class="toc-item-num">8.1 </span>Process Time Line</a></span></li></ul></li><li><span><a href="#Other-Processes-on-Host---Clustering" data-toc-modified-id="Other-Processes-on-Host---Clustering-9"><span class="toc-item-num">9 </span>Other Processes on Host - Clustering</a></span><ul class="toc-item"><li><span><a href="#Clustered-Processes" data-toc-modified-id="Clustered-Processes-9.1"><span class="toc-item-num">9.1 </span>Clustered Processes</a></span></li><li><span><a href="#Variability-in-Command-Lines-and-Process-Names" data-toc-modified-id="Variability-in-Command-Lines-and-Process-Names-9.2"><span class="toc-item-num">9.2 </span>Variability in Command Lines and Process Names</a></span></li><li><span><a href="#Time-Line-of--clustered-processes-data-vs.-original-data" data-toc-modified-id="Time-Line-of--clustered-processes-data-vs.-original-data-9.3"><span class="toc-item-num">9.3 </span>Time Line of clustered processes data vs. original data</a></span></li></ul></li><li><span><a href="#Base64-Decode-and-Check-for-IOCs" data-toc-modified-id="Base64-Decode-and-Check-for-IOCs-10"><span class="toc-item-num">10 </span>Base64 Decode and Check for IOCs</a></span><ul class="toc-item"><li><span><a href="#IoCs-in-the-entire-data-set" data-toc-modified-id="IoCs-in-the-entire-data-set-10.1"><span class="toc-item-num">10.1 </span>IoCs in the entire data set</a></span></li><li><span><a href="#If-any-Base64-encoded-strings,-decode-and-search-for-IoCs-in-the-results." data-toc-modified-id="If-any-Base64-encoded-strings,-decode-and-search-for-IoCs-in-the-results.-10.2"><span class="toc-item-num">10.2 </span>If any Base64 encoded strings, decode and search for IoCs in the results.</a></span></li></ul></li><li><span><a href="#Threat-Intelligence-Lookup" data-toc-modified-id="Threat-Intelligence-Lookup-11"><span class="toc-item-num">11 </span>Threat Intelligence Lookup</a></span></li><li><span><a href="#Alert-command-line---Occurrence-on-other-hosts-in-workspace" data-toc-modified-id="Alert-command-line---Occurrence-on-other-hosts-in-workspace-12"><span class="toc-item-num">12 </span>Alert command line - Occurrence on other hosts in workspace</a></span></li><li><span><a href="#Host-Logons" data-toc-modified-id="Host-Logons-13"><span class="toc-item-num">13 </span>Host Logons</a></span></li><li><span><a href="#Alert-Logon-Account" data-toc-modified-id="Alert-Logon-Account-14"><span class="toc-item-num">14 </span>Alert Logon Account</a></span><ul class="toc-item"><li><span><a href="#All-Host-Logons" data-toc-modified-id="All-Host-Logons-14.1"><span class="toc-item-num">14.1 </span>All Host Logons</a></span></li><li><span><a href="#Comparing-All-Logons-with-Clustered-results-relative-to-Alert-time-line" data-toc-modified-id="Comparing-All-Logons-with-Clustered-results-relative-to-Alert-time-line-14.2"><span class="toc-item-num">14.2 </span>Comparing All Logons with Clustered results relative to Alert time line</a></span></li><li><span><a href="#View-Process-Session-and-Logon-Events-in-Timelines" data-toc-modified-id="View-Process-Session-and-Logon-Events-in-Timelines-14.3"><span class="toc-item-num">14.3 </span>View Process Session and Logon Events in Timelines</a></span></li></ul></li><li><span><a href="#Failed-Logons" data-toc-modified-id="Failed-Logons-15"><span class="toc-item-num">15 </span>Failed Logons</a></span></li><li><span><a href="#Appendices" data-toc-modified-id="Appendices-16"><span class="toc-item-num">16 </span>Appendices</a></span><ul class="toc-item"><li><span><a href="#Available-DataFrames" data-toc-modified-id="Available-DataFrames-16.1"><span class="toc-item-num">16.1 </span>Available DataFrames</a></span></li><li><span><a href="#Saving-Data-to-CSV" data-toc-modified-id="Saving-Data-to-CSV-16.2"><span class="toc-item-num">16.2 </span>Saving Data to CSV</a></span></li><li><span><a href="#Saving-Data-to-Excel" data-toc-modified-id="Saving-Data-to-Excel-16.3"><span class="toc-item-num">16.3 </span>Saving Data to Excel</a></span></li></ul></li><li><span><a href="#Setup-Cell" data-toc-modified-id="Setup-Cell-17"><span class="toc-item-num">17 </span>Setup Cell</a></span><ul class="toc-item"><li><span><a href="#msticpyconfig.yaml-configuration-File" data-toc-modified-id="msticpyconfig.yaml-configuration-File-17.1"><span class="toc-item-num">17.1 </span><code>msticpyconfig.yaml</code> configuration File</a></span></li></ul></li></ul></div>
# -
# <a></a>[Contents](#toc)
# ## Hunting Hypothesis
# Our broad initial hunting hypothesis is that a we have received an alert/indicators involving windows process name which is suspected to be malicious, we will need to hunt from a range of different positions to validate or disprove this hypothesis.
#
# Before you start hunting please run the cells in <a>Setup</a> at the bottom of this Notebook.
# ## Notebook Initialization
# This cell:
#
# - Checks for the correct Python version
# - Checks versions and optionally installs required packages
# - Imports the required packages into the notebook
# - Sets a number of configuration options.
#
# This should complete without errors. If you encounter errors or warnings look at the following two notebooks:
# - [TroubleShootingNotebooks](https://github.com/Azure/Azure-Sentinel-Notebooks/blob/master/TroubleShootingNotebooks.ipynb)
# - [ConfiguringNotebookEnvironment](https://github.com/Azure/Azure-Sentinel-Notebooks/blob/master/ConfiguringNotebookEnvironment.ipynb)
#
# You may also need to do some additional configuration to successfully use functions such as Threat Intelligence service lookup and Geo IP lookup. See the <a href="#Configuration">Configuration</a> section at the end of the notebook and the [ConfiguringNotebookEnvironment](https://github.com/Azure/Azure-Sentinel-Notebooks/blob/master/ConfiguringNotebookEnvironment.ipynb).
#
# +
# Imports
import importlib
import os
import sys
import warnings
from IPython.display import display, HTML, Markdown
display(HTML("<h3>Starting Notebook setup...</h3>"))
warn_mssg = []
err_mssg = []
MISSING_PKG_ERR = """
<h3><font color='red'>Warning {package} is not installed or has an incorrect version</h3></font>
For more details, please go to the <a href="#Setup">Setup section</a> at the end of the notebook.
"""
MIN_REQ_PYTHON = (3, 6)
PANDAS_REQ_VERSION = (0, 25, 0)
MSTICPY_REQ_VERSION = (0, 2, 7)
if sys.version_info < MIN_REQ_PYTHON:
display(HTML("""
<h2><font color='red'>Incorrect notebook kernel version detected</h2></font>
Please check the <b>Kernel->Change Kernel</b> menu and ensure that <b>Python 3.6</b><br>
or later is selected as the active kernel."""))
raise RuntimeError("Python %s.%s or later kernel is required." % MIN_REQ_PYTHON)
print(
"Python kernel version %s.%s.%s" % (
sys.version_info[0], sys.version_info[1], sys.version_info[2]
)
)
try:
import msticpy
mp_version = tuple([int(v) for v in msticpy.__version__.split(".")])
if mp_version < MSTICPY_REQ_VERSION:
raise ImportError("msticpy %s.%s.%s or later is required." % MSTICPY_REQ_VERSION)
print("msticpy imported version %s" % msticpy.__version__)
except ImportError:
display(HTML(MISSING_PKG_ERR.format(package="msticpy")))
resp = input("Install the package now? (y/n)")
if resp.casefold().startswith("y"):
# !pip install --user --upgrade msticpy
warn_mssg.append("msticpy was installed or upgraded.")
if "msticpy" in sys.modules:
importlib.reload(msticpy)
else:
import msticpy
try:
from IPython import get_ipython
import ipywidgets as widgets
from pathlib import Path
import matplotlib.pyplot as plt
from matplotlib import MatplotlibDeprecationWarning
import seaborn as sns
import numpy as np
import pandas as pd
import tqdm
pd_version = tuple([int(v) for v in pd.__version__.split(".")])
if pd_version < PANDAS_REQ_VERSION:
display(HTML(MISSING_PKG_ERR.format(package="pandas")))
resp = input("Install the package now? (y/n)")
if resp.casefold().startswith("y"):
warn_mssg.append("pandas was installed or upgraded.")
# !pip install --user --upgrade pandas
if "pandas" in sys.modules:
importlib.reload(pd)
else:
import pandas as pd
print("pandas imported version %s" % pd.__version__)
from msticpy.data import QueryProvider
from msticpy.nbtools import *
from msticpy.sectools import *
from msticpy.nbtools.foliummap import FoliumMap
from msticpy.nbtools.utility import md, md_warn
from msticpy.nbtools.wsconfig import WorkspaceConfig
from msticpy.nbtools.query_defns import DataFamily
from msticpy.nbtools.entityschema import IpAddress, GeoLocation
additional_packages = []
if additional_packages:
utils.check_and_install_missing_packages(additional_packages)
from dns import reversename, resolver
from ipwhois import IPWhois
except ImportError as imp_err:
display(HTML("""
<h2><font color='red'>One or more missing packages detected</h2>
Please correct these by installing the required packages, restart
the kernel and re-run the notebook.</font>
<i>Package error: %s</i><br>
""" % imp_err))
err_mssg.append("One or more missing packages found.")
else:
mp_path = os.environ.get("MSTICPYCONFIG", "./msticpyconfig.yaml")
if not Path(mp_path).exists():
display(HTML("""
<h3><font color='orange'>Warning: no <i>msticpyconfig.yaml</i> found</h3></font>
Some functionality (such as Threat Intel lookups) will not function without valid configuration
settings.
Please go to the <a href="#Setup">Setup section</a> follow the instructions there.
"""))
warn_mssg.append("msticpyconfig.yaml not found.")
WIDGET_DEFAULTS = {
"layout": widgets.Layout(width="95%"),
"style": {"description_width": "initial"},
}
# Some of our dependencies still use deprecated Matplotlib
# APIs - we can't do anything about it, so suppress them from view
warnings.simplefilter("ignore", category=MatplotlibDeprecationWarning)
warnings.filterwarnings("ignore", category=DeprecationWarning)
sns.set()
pd.set_option("display.max_rows", 100)
pd.set_option("display.max_columns", 50)
pd.set_option("display.max_colwidth", 100)
os.environ["KQLMAGIC_LOAD_MODE"]="silent"
with_errs = "<br>with %s errors" % len(err_mssg) if err_mssg else ""
with_warns = "<br>with %s warnings" % len(warn_mssg) if warn_mssg else ""
if err_mssg:
display(HTML("<font color='red'><h3>Errors:</h3>" + '<br>'.join(err_mssg)))
if warn_mssg:
display(HTML("<font color='red'><h3>Warnings:</h3>" + '<br>'.join(warn_mssg)))
display(HTML("<h3>Notebook setup complete</h3>" + with_errs + with_warns))
# -
# <a></a>[Contents](#toc)
# ### Get WorkspaceId and Authenticate to Log Analytics
# <details>
# <summary> <u>Details...</u></summary>
# If you are using user/device authentication, run the following cell.
# - Click the 'Copy code to clipboard and authenticate' button.
# - This will pop up an Azure Active Directory authentication dialog (in a new tab or browser window). The device code will have been copied to the clipboard.
# - Select the text box and paste (Ctrl-V/Cmd-V) the copied value.
# - You should then be redirected to a user authentication page where you should authenticate with a user account that has permission to query your Log Analytics workspace.
#
# Use the following syntax if you are authenticating using an Azure Active Directory AppId and Secret:
# ```
# # %kql loganalytics://tenant(aad_tenant).workspace(WORKSPACE_ID).clientid(client_id).clientsecret(client_secret)
# ```
# instead of
# ```
# # %kql loganalytics://code().workspace(WORKSPACE_ID)
# ```
#
# Note: you may occasionally see a JavaScript error displayed at the end of the authentication - you can safely ignore this.<br>
# On successful authentication you should see a ```popup schema``` button.
# To find your Workspace Id go to [Log Analytics](https://ms.portal.azure.com/#blade/HubsExtension/Resources/resourceType/Microsoft.OperationalInsights%2Fworkspaces). Look at the workspace properties to find the ID.
# </details>
# +
# To list configured workspaces run WorkspaceConfig.list_workspaces()
# WorkspaceConfig.list_workspaces()
# -
# Authentication
ws_config = WorkspaceConfig()
qry_prov = QueryProvider(data_environment="LogAnalytics")
qry_prov.connect(connection_str=ws_config.code_connect_str)
table_index = qry_prov.schema_tables
# ## Get List of Alerts
#
# We are using an alert as the starting point for this investigation, specify a time range to search for alerts. Once this is set run the following cell to retrieve any alerts in that time window.
# You can change the time range and re-run the queries until you find the alerts that you want to investigate.
alert_q_times = nbwidgets.QueryTime(units='hour',
max_before=72, max_after=1, before=24)
alert_q_times.display()
# +
alert_list = qry_prov.SecurityAlert.list_alerts(
alert_q_times)
alert_counts = qry_prov.SecurityAlert.list_alerts_counts(
alert_q_times)
if isinstance(alert_list, pd.DataFrame) and not alert_list.empty:
print(len(alert_counts), ' distinct alert types')
print(len(alert_list), ' distinct alerts')
# Display alerts on timeline to aid in visual grouping
nbdisplay.display_timeline(
data=alert_list, source_columns=["AlertName", 'CompromisedEntity'], title="Alerts over time", height=300, color="red")
display(alert_counts.head(10)) # remove '.head(10)'' to see the full list grouped by AlertName
else:
display(Markdown('No alerts found.'))
# -
# ## Choose Alert to Investigate
# To focus the investigation select an alert from a list of retrieved alerts.
#
# As you select an alert, the main properties will be shown below the list.
#
# Use the filter box to narrow down your search to any substring in the AlertName.
get_alert = None
alert_select = nbwidgets.AlertSelector(alerts=alert_list, action=nbdisplay.display_alert)
alert_select.display()
# ## Extract properties and entities from Alert
# In order to pivot to data related to the selected security alert we need to identify key data points in the selected alert. This section extracts the alert information and entities into a SecurityAlert object allowing us to query the properties more reliably.
#
# Properties in this object will be used to automatically provide parameters for queries and UI elements.
# Subsequent queries will use properties like the host name and derived properties such as the OS family (Linux or Windows) to adapt the query. Query time selectors like the one above will also default to an origin time that matches the alert selected.
#
# The alert view below shows all of the main properties of the alert plus the extended property dictionary (if any) and JSON representations of the Entity.
# +
# Extract entities and properties into a SecurityAlert class
if alert_select is None or alert_select.selected_alert is None:
raise ValueError("Please select an alert before executing remaining cells.")
else:
security_alert = SecurityAlert(alert_select.selected_alert)
nbdisplay.display_alert(security_alert, show_entities=True)
# -
# ## Entity Graph
# Depending on the type of alert there may be one or more entities attached as properties. Entities are key indicators that we can pivot on during our investigation, such as Host, Account, IpAddress, Process, etc. - essentially the 'nouns' of security investigation.
# Entities are often related to other entities - for example a process will usually have a related file entity (the process image) and an Account entity (the context in which the process was running). Endpoint alerts typically always have a host entity (which could be a physical or virtual machine). In order to more effectively understand the links between related entities we can plot them as a graph.
# Draw the graph using Networkx/Matplotlib
# %matplotlib inline
alertentity_graph = security_alert_graph.create_alert_graph(security_alert)
nbdisplay.draw_alert_entity_graph(alertentity_graph, width=15)
#
# ## Related Alerts
# For certain entities in the alert we can search for other alerts that have that entity in common. Currently this pivot supports alerts with the same Host, Account or Process.
#
# **Notes:**
# - Some alert types do not include all of these entity types.
# - The original alert will be included in the "Related Alerts" set if it occurs within the query time boundary set below.
#
# In order to more effectively identify related alerts the query time boundaries can be adjusted to encompass a longer time frame.
# set the origin time to the time of our alert
query_times = nbwidgets.QueryTime(units='day', origin_time=security_alert.TimeGenerated,
max_before=28, max_after=1, before=5)
query_times.display()
if not security_alert.primary_host:
md_warn('Related alerts is not yet supported for alerts that are not host-based')
related_alerts = None
else:
hostname = security_alert.hostname
if not hostname and "AzureID" in security_alert.primary_host:
hostname = security_alert.primary_host.AzureID.split("/")[-1]
related_alerts = qry_prov.SecurityAlert.list_related_alerts(query_times, security_alert, host_name=hostname)
if related_alerts is not None and not related_alerts.empty:
host_alert_items = related_alerts\
.query('host_match == @True')[['AlertType', 'StartTimeUtc']]\
.groupby('AlertType').StartTimeUtc.agg('count').to_dict()
acct_alert_items = related_alerts\
.query('acct_match == @True')[['AlertType', 'StartTimeUtc']]\
.groupby('AlertType').StartTimeUtc.agg('count').to_dict()
proc_alert_items = related_alerts\
.query('proc_match == @True')[['AlertType', 'StartTimeUtc']]\
.groupby('AlertType').StartTimeUtc.agg('count').to_dict()
def print_related_alerts(alertDict, entityType, entityName):
if len(alertDict) > 0:
print('Found {} different alert types related to this {} (\'{}\')'
.format(len(alertDict), entityType, entityName))
for (k,v) in alertDict.items():
print(' {}, Count of alerts: {}'.format(k, v))
else:
print('No alerts for {} entity \'{}\''.format(entityType, entityName))
print_related_alerts(host_alert_items, 'host', security_alert.hostname)
print_related_alerts(acct_alert_items, 'account',
security_alert.primary_account.qualified_name
if security_alert.primary_account
else None)
print_related_alerts(proc_alert_items, 'process',
security_alert.primary_process.ProcessFilePath
if security_alert.primary_process
else None)
nbdisplay.display_timeline(data=related_alerts, source_columns = ['AlertName'], title='Alerts', height=100)
else:
md('No related alerts found.', styles=["bold","green"])
# ### Show these related alerts on a graph
# To see the how these alerts relate to our original alert, and how these new alerts relate to each other we can graph them.
# +
# Draw a graph of this (add to entity graph)
# %matplotlib notebook
# %matplotlib inline
if related_alerts is not None and not related_alerts.empty:
rel_alert_graph = security_alert_graph.add_related_alerts(related_alerts=related_alerts,
alertgraph=alertentity_graph)
nbdisplay.draw_alert_entity_graph(rel_alert_graph, width=15)
else:
md('No related alerts found.', styles=["bold","green"])
# -
# ### Browse List of Related Alerts
# Once we have understood how these alerts related to each other, we can view the details of each new, related alert.
# +
def disp_full_alert(alert):
global related_alert
related_alert = SecurityAlert(alert)
nbdisplay.display_alert(related_alert, show_entities=True)
if related_alerts is not None and not related_alerts.empty:
related_alerts['CompromisedEntity'] = related_alerts['Computer']
print('Selected alert is available as \'related_alert\' variable.')
rel_alert_select = nbwidgets.AlertSelector(alerts=related_alerts, action=disp_full_alert)
rel_alert_select.display()
else:
md('No related alerts found.', styles=["bold","green"])
# -
# ## Get Process Tree
# If the alert has a process entity this section tries to retrieve the entire process tree to which that process belongs.
#
# Notes:
# - The alert must have a process entity
# - Only processes started within the query time boundary will be included
# - Ancestor and descented processes are retrieved to two levels (i.e. the parent and grandparent of the alert process plus any child and grandchild processes).
# - Sibling processes are the processes that share the same parent as the alert process
# - This can be a long-running query, especially if a wide time window is used! Caveat Emptor!
#
# The source (alert) process is shown in red.
#
# What's shown for each process:
# - Each process line is indented according to its position in the tree hierarchy
# - Top line fields:
# - \[relationship to source process:lev# - where # is the hops away from the source process\]
# - Process creation date-time (UTC)
# - Process Image path
# - PID - Process Id
# - SubjSess - the session Id of the process spawning the new process
# - TargSess - the new session Id if the process is launched in another context/session. If 0/0x0 then the process is launched in the same session as its parent
# - Second line fields:
# - Process command line
# - Account - name of the account context in which the process is running
# set the origin time to the time of our alert
query_times = nbwidgets.QueryTime(units='minute', origin_time=security_alert.origin_time)
query_times.display()
# +
if security_alert.data_family != DataFamily.WindowsSecurity:
raise ValueError('The remainder of this notebook currently only supports Windows. '
'Linux support is in development but not yet implemented.')
def extract_missing_pid(security_alert):
for pid_ext_name in ['Process Id', 'Suspicious Process Id']:
pid = security_alert.ExtendedProperties.get(pid_ext_name, None)
if pid:
return pid
def extract_missing_sess_id(security_alert):
sess_id = security_alert.ExtendedProperties.get('Account Session Id', None)
if sess_id:
return sess_id
for session in [e for e in security_alert.entities if
e['Type'] == 'host-logon-session' or e['Type'] == 'hostlogonsession']:
return session['SessionId']
if (security_alert.primary_process):
# Do some patching up if the process entity doesn't have a PID
pid = security_alert.primary_process.ProcessId
if not pid:
pid = extract_missing_pid(security_alert)
if pid:
security_alert.primary_process.ProcessId = pid
else:
raise ValueError('Could not find the process Id for the alert process.')
# Do the same if we can't find the account logon ID
sess_id = security_alert.get_logon_id()
if sess_id:
sess_id = extract_missing_sess_id(security_alert)
if sess_id and security_alert.primary_account:
security_alert.primary_account.LogonId = sess_id
else:
raise ValueError('Could not find the session Id for the alert process.')
# run the query
process_tree = qry_prov.WindowsSecurity.get_process_tree(
query_times,
security_alert,
process_id=pid,
logon_session_id=sess_id,
host_name=security_alert.primary_host.HostName,
process_name=security_alert.primary_process.ImageFile.FullPath)
if process_tree is not None and not process_tree.empty:
# Print out the text view of the process tree
nbdisplay.display_process_tree(process_tree)
else:
md_warn('No processes were returned so cannot obtain a process tree Skip.')
display(Markdown('Skip to [Other Processes](#process_clustering) later in the notebook to retrieve all processes'))
else:
md_warn('This alert has no process entity so cannot obtain a process tree.')
display(Markdown('Skip to [Other Processes](#process_clustering) later in the notebook to retrieve all processes'))
process_tree = None
# -
# ### Process Time Line
# As well as seeing the processes involved in a tree we want to see the chronology of this process execution. This shows each process in the process tree on a time line view.
# If a large number of processes are involved in this process tree it may take some time to display this time line graphic.
# Show timeline of events
if process_tree is not None and not process_tree.empty:
nbdisplay.display_timeline(data=process_tree, alert=security_alert,
title='Alert Process Session', height=250)
else:
md_warn('This alert has no process entity so cannot obtain a process tree.')
display(Markdown('Skip to [Other Processes](#process_clustering) later in the notebook to retrieve all processes'))
# ## Other Processes on Host - Clustering
# Sometimes you don't have a source process from which to build our investigation. Other times it's just useful to see what other process activity is occurring on the host. This section retrieves all processes on the host within the time bounds
# set in the query times widget.
#
# If you want to view the raw details of this process data display the *processes_on_host* dataframe.
#
# In order to more effectively analyze this process data we can cluster processes into distinct process clusters.
# To do this we process the raw event list output to extract a few features that render strings (such as commandline)into numerical values. The default below uses the following features:
# - commandLineTokensFull - this is a count of common delimiters in the commandline
# (given by this regex r'[\s\-\\/\.,"\'|&:;%$()]'). The aim of this is to capture the commandline structure while ignoring variations on what is essentially the same pattern (e.g. temporary path GUIDs, target IP or host names, etc.)
# - pathScore - this sums the ordinal (character) value of each character in the path (so /bin/bash and /bin/bosh would have similar scores).
# - isSystemSession - 1 if this is a root/system session, 0 if anything else.
#
# Then we run a clustering algorithm (DBScan in this case) on the process list. The result groups similar (noisy) processes together and leaves unique process patterns as single-member clusters.
# ### Clustered Processes
# +
from msticpy.sectools.eventcluster import dbcluster_events, add_process_features
from tqdm.notebook import tqdm
tqdm.pandas(desc="progress")
processes_on_host = None
if security_alert.primary_host:
md("Querying data...")
processes_on_host = qry_prov.WindowsSecurity.list_host_processes(
query_times,
security_alert,
host_name=hostname
)
if processes_on_host is not None and not processes_on_host.empty:
md("Extracting features...")
feature_procs = add_process_features(input_frame=processes_on_host,
path_separator=security_alert.path_separator)
# you might need to play around with the max_cluster_distance parameter.
# decreasing this gives more clusters.
md("Clustering data...")
(clus_events, dbcluster, x_data) = dbcluster_events(data=feature_procs,
cluster_columns=['commandlineTokensFull',
'pathScore',
'isSystemSession'],
max_cluster_distance=0.0001)
print('Number of input events:', len(feature_procs))
print('Number of clustered events:', len(clus_events))
height = int(len(clus_events[clus_events['ClusterSize'] > 1]) / 4)
clus_events[['ClusterSize', 'processName']][clus_events['ClusterSize'] > 1].plot.barh(x='processName',
title='Process names with Cluster > 1',
figsize=(6, height));
if processes_on_host is None or processes_on_host.empty:
md('Unable to obtain any processes for this host. This feature is currently only supported for Windows hosts.', styles=["blue","bold"])
md('If this is a Windows host skip to [Host Logons](#host_logons) later in the notebook to examine logon events.', styles=["bold","large"])
# -
# ### Variability in Command Lines and Process Names
# In this section we display a number of charts highlighting the variability of command lines and processes paths associated with each process.
#
# The top chart shows the variability of command line content for a given process name. The wider the box, the more instances were found with different command line structure. For certain processes such as cmd.exe or powershell.exe a wide variability in command lines could be expected, however with other processes this could be considered abnormal.
#
# Note, the 'structure' in this case is measured by the number of tokens or delimiters in the command line and does not look at content differences. This is done so that commonly varying instances of the same command line are grouped together.<br>
# For example `updatepatch host1.mydom.com` and `updatepatch host2.mydom.com` will be grouped together.
#
#
# The second graph shows processes by variation in the full path associated with the process. This does compare content so `c:\windows\system32\net.exe` and `e:\windows\system32\net.exe` are treated as distinct. You would normally not expect to see any variability in this chart unless you have multiple copies of the same name executable or an executable is trying masquerade as another well-known binary.
# +
# Looking at the variability of commandlines and process image paths
import seaborn as sns
sns.set(style="darkgrid")
if processes_on_host is not None and not processes_on_host.empty:
proc_plot = sns.catplot(y="processName", x="commandlineTokensFull",
data=feature_procs.sort_values('processName'),
kind='box', height=10)
proc_plot.fig.suptitle('Variability of Commandline Tokens', x=1, y=1)
proc_plot = sns.catplot(y="processName", x="pathLogScore",
data=feature_procs.sort_values('processName'),
kind='box', height=10, hue='isSystemSession')
proc_plot.fig.suptitle('Variability of Path', x=1, y=1);
# -
if 'clus_events' in locals() and not clus_events.empty:
resp = input('View the clustered data? y/n')
if resp == 'y':
display(clus_events.sort_values('TimeGenerated')[['TimeGenerated', 'LastEventTime',
'NewProcessName', 'CommandLine',
'ClusterSize', 'commandlineTokensFull',
'pathScore', 'isSystemSession']])
# +
# Look at clusters for individual process names
def view_cluster(exe_name):
display(clus_events[['ClusterSize', 'processName', 'CommandLine', 'ClusterId']][clus_events['processName'] == exe_name])
display(Markdown('You can view the cluster members for individual processes'
'by inserting a new cell and entering:<br>'
'`view_cluster(process_name)`<br></div>'
'where process_name is the unqualified process binary. E.g<br>'
'`view_cluster("reg.exe")`'))
# -
# ### Time Line of clustered processes data vs. original data
# Show timeline of events - clustered events
if 'clus_events' in locals() and not clus_events.empty:
nbdisplay.display_timeline(data=clus_events,
overlay_data=processes_on_host,
alert=security_alert,
title='Distinct Host Processes (bottom) and All Proceses (top)')
# <a></a>[Contents](#toc)
# ## Base64 Decode and Check for IOCs
# This section looks for Indicators of Compromise (IoC) within the data sets passed to it.
#
# The first section looks at the command line for the process related to our original alert (if any). It also looks for Base64 encoded strings within the data - this is a common way of hiding attacker intent. It attempts to decode any strings that look like Base64. Additionally, if the Base64 decode operation returns any items that look like a Base64 encoded string or file, a gzipped binary sequence, a zipped or tar archive, it will attempt to extract the contents before searching for potentially interesting items.
# +
process = security_alert.primary_process
ioc_extractor = IoCExtract()
if process and process["CommandLine"]:
# if nothing is decoded this just returns the input string unchanged
base64_dec_str, _ = base64.unpack_items(input_string=process["CommandLine"])
if base64_dec_str and '<decoded' in base64_dec_str:
print('Base64 encoded items found.')
print(base64_dec_str)
# any IoCs in the string?
iocs_found = ioc_extractor.extract(base64_dec_str)
if iocs_found:
print('\nPotential IoCs found in alert process:')
display(iocs_found)
else:
print('No IoCs found in alert process:')
else:
print('No process command line available in selected alert.')
# -
# ### IoCs in the entire data set
# If we have a process tree or other elements that contain command lines we also want to attempt to extract IoCs from these data sets.
# +
ioc_extractor = IoCExtract()
source_processes = None
# if process tree is populated we use that preferentially
try:
if not process_tree.empty:
source_processes = process_tree
except (NameError, AttributeError):
pass
# If not, use the clustered events from the all sessions
try:
if source_processes is None and not clus_events.empty:
source_processes = clus_events
except (NameError, AttributeError):
pass
if source_processes is not None and not source_processes.empty:
ioc_df = ioc_extractor.extract(data=source_processes,
columns=['CommandLine'],
os_family=security_alert.os_family,
ioc_types=['ipv4', 'ipv6', 'dns', 'url',
'md5_hash', 'sha1_hash', 'sha256_hash'])
if len(ioc_df):
display(HTML("<h3>IoC patterns found in process tree.</h3>"))
display(ioc_df)
else:
ioc_df = None
# -
# ### If any Base64 encoded strings, decode and search for IoCs in the results.
# For simple strings the Base64 decoded output is straightforward. However it is not uncommon to see nested encodings therefore we want to try to extract and decode these nested elements as well.
# +
if source_processes is not None:
dec_df = base64.unpack_items(data=source_processes, column='CommandLine')
if source_processes is not None and (dec_df is not None and not dec_df.empty):
display(HTML("<h3>Decoded base 64 command lines</h3>"))
md_warn("Some binary patterns may be decodable as unicode strings")
display(dec_df[['full_decoded_string', 'original_string', 'decoded_string', 'input_bytes', 'file_hashes']])
ioc_dec_df = ioc_extractor.extract(data=dec_df, columns=['full_decoded_string'])
if len(ioc_dec_df):
display(HTML("<h3>IoC patterns found in base 64 decoded data</h3>"))
display(ioc_dec_df)
if ioc_df is not None:
ioc_df = ioc_df.append(ioc_dec_df ,ignore_index=True)
else:
ioc_df = ioc_dec_df
else:
print("No base64 encodings found.")
# + [markdown] hidden=true
# ## Threat Intelligence Lookup
# Now that we have identified a number of IoCs we want to check to see if they are associated with known mallicious activity. To do this we will query three different Threat Intelligence providers to see if we get results.
#
# We will be using:
# - VirusTotal https://www.virustotal.com/.
# - Alienware OTX https://otx.alienvault.com/
# - IBM X-Force https://exchange.xforce.ibmcloud.com/
#
# If you do not have an API key for any of these providers simply remove their name from the providers list in our lookup_iocs command.
# +
from msticpy.sectools.tiproviders.ti_provider_base import TISeverity
def ti_check_ser_sev(data, threshold):
threshold = TISeverity.parse(threshold)
return data.apply(lambda x: TISeverity.parse(x) >= threshold)
tilookups = TILookup()
if ioc_df is not None and not ioc_df.empty:
ti_results = tilookups.lookup_iocs(data=ioc_df, obs_col='Observable', ioc_type_col='IoCType', providers=["OTX", "VirusTotal", "XForce"])
if not ti_results[ti_check_ser_sev(ti_results['Severity'], 1)].empty:
md("Positive TI Results:",styles=["bold","red","large"])
display(ti_results[ti_check_ser_sev(ti_results['Severity'], 1)])
else:
md("No postive matches found in threat intelligence",styles=["bold","green"])
else:
md_warn("No IOCs to lookup")
# -
# ## Alert command line - Occurrence on other hosts in workspace
# Understanding where else a command line is being run in an environment can give us a good idea of the scope of a security incident, or help us determine whether activity is malicious or expected.
#
# To get a sense of whether the alert process is something that is occuring on other hosts, run this section.
# set the origin time to the time of our alert
query_times = nbwidgets.QueryTime(units='day', before=5, max_before=20,
after=1, max_after=10,
origin_time=security_alert.origin_time)
query_times.display()
# +
# This query needs a commandline parameter which isn't supplied
# by default from the the alert
# - so extract and escape this from the process
proc_match_in_ws = None
if not security_alert.primary_process:
md_warn('This alert has no process entity. This section is not applicable.')
elif not security_alert.primary_process.CommandLine:
md_warn('This alert process entity has no commandline. This section is not applicable.')
elif security_alert.primary_process:
commandline = security_alert.primary_process.CommandLine
commandline = utils.escape_windows_path(commandline)
commandline = commandline.replace('"',"'")
process = security_alert.primary_process.ProcessName
if not process:
raise ValueError("No process name found in selected alert")
process = utils.escape_windows_path(process)
process = process.replace('"',"'")
md(f"Command Line: '{commandline}'", styles=["bold"])
if commandline.strip():
proc_match_in_ws = qry_prov.WindowsSecurity.list_hosts_matching_commandline(start=query_times.start, end=query_times.end, process_name=process,
commandline=commandline)
else:
md('process has empty commandline')
# Check the results
if proc_match_in_ws is None or proc_match_in_ws.empty:
md('No proceses with matching commandline found in on other hosts in workspace', styles=["bold","blue"])
md(f'between: {query_times.start} and {query_times.end}')
else:
hosts = proc_match_in_ws['Computer'].drop_duplicates().shape[0]
processes = proc_match_in_ws.shape[0]
md('{numprocesses} proceses with matching commandline found on {numhosts} hosts in workspace'\
.format(numprocesses=processes, numhosts=hosts))
md(f'between: {query_times.start} and {query_times.end}')
md('To examine these execute the dataframe \'{}\' in a new cell'.format('proc_match_in_ws'))
md(proc_match_in_ws[['TimeCreatedUtc','Computer', 'NewProcessName', 'CommandLine']].head())
# -
# If at this point you wish to investigate a particular host in detail you can use the cells below or you can switch to our Host Investigation Notebooks that provide a deep dive capability for Windows and Linux hosts.
#
# ## Host Logons
# This section retrieves the logon events on the host in the alert.
#
# You may want to use the query times to search over a broader range than the default.
# set the origin time to the time of our alert
query_times = nbwidgets.QueryTime(units='day', origin_time=security_alert.origin_time,
before=1, after=0, max_before=20, max_after=1)
query_times.display()
# + [markdown] hidden=true
# If you wish to investigate a specific host in detail you can use the cells below or switch to our Account investigation notebook.
#
# ### Alert Logon Account
# This returns the account associated with the alert being investigated.
# +
logon_id = security_alert.get_logon_id()
if logon_id:
if logon_id in ['0x3e7', '0X3E7', '-1', -1]:
print('Cannot retrieve single logon event for system logon id '
'- please continue with All Host Logons below.')
else:
logon_event = qry_prov.WindowsSecurity.get_host_logon(provs=[query_times, security_alert], host_name=security_alert.ExtendedProperties['Compromised Host'],start=query_times.start, end= query_times.end,logon_session_id=sess_id)
nbdisplay.display_logon_data(logon_event, security_alert)
else:
print('No account entity in the source alert or the primary account had no logonId value set.')
# -
# ### All Host Logons
# Since the number of logon events may be large and, in the case of system logons, very repetitive, we use clustering to try to identity logons with unique characteristics.
#
# In this case we use the numeric score of the account name and the logon type (i.e. interactive, service, etc.). The results of the clustered logons are shown below along with a more detailed, readable printout of the logon event information. The data here will vary depending on whether this is a Windows or Linux host.
# +
from msticpy.sectools.eventcluster import dbcluster_events, add_process_features, _string_score
if security_alert.primary_host:
md("Querying data...")
host_logons = qry_prov.WindowsSecurity.list_host_logons(
query_times, security_alert, host_name=security_alert.primary_host.HostName
)
host_logons_time = (
host_logons.astype({'LogonType': 'int32'})
.merge(right=pd.Series(data=nbdisplay._WIN_LOGON_TYPE_MAP, name="LogonTypeDesc"),
left_on="LogonType", right_index=True)
.set_index("TimeGenerated")[["LogonTypeDesc"]]
.groupby(["LogonTypeDesc"])
.resample("10T")
.count()
.rename(columns={"LogonTypeDesc": "Count"}).reset_index()
)
fig, ax = plt.subplots()
for l_type, logon_group in host_logons_time.groupby("LogonTypeDesc"):
logon_group.plot.line(x="TimeGenerated", y="Count", label=l_type, title="Number of logons by type", ax=ax)
else:
host_logons = None
md("No data available - alert has no host entity.")
# -
# #### Highest and lowest number of logon types by Account
# +
select_logon_type = widgets.Select(options=nbdisplay._WIN_LOGON_TYPE_MAP.values(), layout=widgets.Layout(height="200px"))
num_items = widgets.IntSlider(min=1, max=200, value=10, description="# logons")
df_output1 = widgets.Output()
df_output2 = widgets.Output()
def display_logons(host_logons, order_column, number_shown, output, title, ascending=True):
pivot_df = (
host_logons[["Account", "LogonType", "EventID"]]
.astype({'LogonType': 'int32'})
.merge(right=pd.Series(data=nbdisplay._WIN_LOGON_TYPE_MAP, name="LogonTypeDesc"),
left_on="LogonType", right_index=True)
.drop(columns="LogonType")
.groupby(["Account", "LogonTypeDesc"])
.count()
.unstack()
.rename(columns={"EventID": "LogonCount"})
.fillna(0)
)
with output:
if ('LogonCount', order_column) in pivot_df.columns:
md(title)
display(
pivot_df
[pivot_df[("LogonCount", order_column)] > 0]
.sort_values(("LogonCount", order_column), ascending=ascending)
.head(number_shown)
.style
.background_gradient(cmap="viridis", low=0.5, high=0)
.format("{0:0>3.0f}")
)
else:
md(f"No logons of type {order_column}")
def show_logons(evt):
del evt
logon_type = select_logon_type.value
list_size = num_items.value
df_output1.clear_output()
df_output2.clear_output()
display_logons(
host_logons,
order_column=logon_type,
number_shown=list_size,
output=df_output1,
title="Most Frequent Logons",
ascending=False,
)
display_logons(
host_logons,
order_column=logon_type,
number_shown=list_size,
output=df_output2,
title="Rarest Logons",
ascending=True
)
select_logon_type.observe(show_logons, names="value")
ctrls = widgets.HBox([select_logon_type, num_items])
outputs = widgets.HBox([df_output1, df_output2])
display(widgets.VBox([ctrls, outputs]))
# + [markdown] hidden=true
# ## Failed Logons
# Failed logons can provide a valuable source of data for investigation so we also want to look at failed logons during the period of our investigation.
# + hidden=true
if security_alert.primary_host:
failedLogons = qry_prov.WindowsSecurity.list_host_logon_failures(
query_times, security_alert, host_name=security_alert.primary_host.HostName
)
else:
md_warn("No data available - alert has no host entity.")
failedLogons = None
if failedLogons is None or failedLogons.empty:
md(f'No logon failures recorded for this host between {security_alert.StartTimeUtc} and {security_alert.EndTimeUtc}', styles=["bold","blue"])
else:
md('Failed Logons observed for the host:')
display(failedLogons)
# + [markdown] hidden=true
# ## Appendices
# ### Available DataFrames
# -
print('List of current DataFrames in Notebook')
print('-' * 50)
current_vars = list(locals().keys())
for var_name in current_vars:
if isinstance(locals()[var_name], pd.DataFrame) and not var_name.startswith('_'):
print(var_name)
# ### Saving Data to CSV
# To save the contents of a pandas DataFrame to an CSV
# use the following syntax
# ```
# host_logons.to_csv('host_logons.csv')
# ```
# + [markdown] heading_collapsed=true tags=["todo"]
# ### Saving Data to Excel
# To save the contents of a pandas DataFrame to an Excel spreadsheet
# use the following syntax
# ```
# writer = pd.ExcelWriter('myWorksheet.xlsx')
# my_data_frame.to_excel(writer,'Sheet1')
# writer.save()
# ```
# -
# ## Configuration
#
# ### `msticpyconfig.yaml` configuration File
# You can configure primary and secondary TI providers and any required parameters in the `msticpyconfig.yaml` file. This is read from the current directory or you can set an environment variable (`MSTICPYCONFIG`) pointing to its location.
#
# To configure this file see the [ConfigureNotebookEnvironment notebook](https://github.com/Azure/Azure-Sentinel-Notebooks/blob/master/ConfiguringNotebookEnvironment.ipynb)
| Guided Investigation - Process-Alerts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # `stock_data.py`
# ## `import` statements
# Firstly, we import all the packages that would be used in StockData.py. We used the `import` statement and created an alias for the packages using the `as` statement.
# We import `numpy` because we would be using some of the in-built functions such as np.nan.
# `pandas` package would enable us to read and overwrite our CSV datafiles.
# `matplotlib.pyplot` package would be used for plotting the stock data into graphs.
#
# ```
# import numpy as np
# import pandas as pd
# import matplotlib.pyplot as plt
# ```
# > *Learning points: Package Aliasing*
#
# > *The programmer can create alias for their imported packages so that it would be easier for them to recognize and use the functions in the packages.*
# ## `class` statement
# We create a class name `StockData` which will contain the attributes and functions.
#
# ```
# class StockData():
# ```
# > *Learning point: Classes*
#
# > *Classes are often created because it allows us to bundle data and functionalities together.*
# ## `__ init __(self)` (constructor statement)
# We create a constructor using `__init__` which requires one string parameter: `filepath`.
# The attribute `filepath` stores the parameter `filepath`.
# The attribute `data` stores the pandas dataframe which is extracted from a CSV file found in the `filepath`.
# When constructing a `StockData` object, it will call and run the `check_data()` function.
#
# ```
# 31 self.filepath = filepath
# 32 self.data = pd.read_csv(filepath).set_index('Date')
# 33 self.check_data()
# ```
# > *Learning point: Default Constructor*
#
# > *If you do not create a Constructor, Python will automatically create a default constructor that does not do anything.*
# ## `check_data(self, overwrite=True)`
# We first start by checking for any missing data and then filling in any missing values by interpolation in the csv data. We use the `interpolate()` function to fill in the estimated values. The interpolate function uses a linear interpolation which takes the average of the value before and after the data point to come out with an estimation. We started with this step so that our dataset would be cleaned and have no missing values.
#
# We define a function name `check_data()`. This functions checks and handles missing data by filling in missing values by interpolation. The parameter `(overwrite = True)` takes a boolean value and overwrites the original source stock data .csv file.
#
# ```
# 48 self.data = self.data.interpolate()
# ```
# > *Learning point: Indentation*
#
# > *When creating a function, we would need to make sure there is proper indentation after the colon. All the code that is in the function would need to have the same indentation.*
# The next part is to overwrite the original stock data.csv file. We would use a pandas inbuilt function`to_csv()` with the parameters (self.filepath) as the filepath and (index= overwrite) to overwrite the csv file.
#
# ```
# 49 self.data.to_csv(self.filepath, index=overwrite)
# ```
#
# We then use `return` to send the StockData to any code that calls this function.
#
# ```
# 50 return self
# ```
# > *Learning point: `return` statement*
#
# > `return` statement is often used at the end of the function to returns the results (values) of the expression to the caller. Statements after the return statement are not executed. If the return statement is without any expression, the value returned would be `none`.*
# ## `get_data(self, start_date, end_date)`
# The get_data function to return a subset of the stock data from start_date to end_date inclusive. The parameter `start_date` and `end_date` has a type `str` that is the start date and end_data of stock data range, must be of format YYYY-MM-DD.
#
# The variable self.selected_data would store a dataframe indexed from the specified start to end date inclusive.
#
# ```
# 72 self.selected_data = self.data[str(start_date):str(end_date)]
# ```
#
# We then use `return` to send the `selected_data` that consist of start and end dates to any code that calls this function.
#
# ```
# 73 return self.selected_data
# ```
# ## `get_period(self)`
# The get_period function is used to obtain the earliest and latest date in the `data` dataframe. Since `data` have index based on the date, we can obtain a list of date with `list(self.data.index)`. With the list, we can obtain the first and last index in the list and return them in a tuple.
#
# ```
# 87 index = list(self.data.index)
# 88 (first, last) = (index[0], index[-1])
# 89 return (first, last)
# ```
# > *Learning point: Returning Multiple Variables*
#
# > *If you want to return more than one variable, you can return them in heterogeneous containers like tuple or list.*
# ## `calculate_SMA(self, n)`
# In the calculate_SMA function, we take in 1 parameter: n which is the number of days used to calculate the simple moving average (SMA). With n, we will create a column label named `SMA + n`.
#
# ```
# 194 col_head = 'SMA' + str(n)
# ```
#
# For example, if n is 15, the column label will be named 'SMA15'
# Due to the dataframe of the self.data having an index using the date, we use reset_index() to undo the index and reinclude date into one of the columns.
#
# ```
# 195 df = self.data.reset_index()
# ```
# > *Learning point: Built-in Functions*
#
# > *To speed the working progress, we should use in-built functions provided by packages if it fulfil the requirements.*
# Then we check if the column name `col_head` is found in `df` by using the following code:
#
# ```
# 197 if col_head not in df.columns:
# ```
# > *Learning point: `not` statement*
#
# > *`not` is a logical operator commonly used with conditional statements such as `if else` or `while`.*
# If it is found in `df`, we will `return self` and leave the dataframe untouched as the SMA of `n` number of days has already been calculated. Otherwise, we will begin the calculation.
#
# We begin by retrieving the list of date found in the self.data(portion of the full data) and creating `returnList` which will store the calculated SMA later on by using the following code:
#
# ```
# 200 dateList = self.data.index.values.tolist()
# 201 returnList = []
# ```
# With this list of data, we will do a for loop with each of the date in the list and find the index of each specific date in the full dataset. We will then use these dateIndex to see if there are enough datasets to calculate the SMA. For example, we need 15 data set prior to the current day in order to calculate the SMA of 15 days. If there is not enough data prior to the current date, we will append NaN into the `returnList` to show that we do not have SMA for that current date.
#
# ```
# 202 for date in dateList:
# 204 dateIndex = df[df["Date"]==date].index.values[0]
# 205 if dateIndex < n: # if date index is less than n: append None
# 206 returnList.append(np.nan)
# 207 else:
# 208 sum = 0
# 209 for i in range(n):
# 210 sum += df.iloc[dateIndex-i]["Close"]
# 212 returnList.append(sum/n)
# ```
# If there is enough data, we will do a for loop with `n` number of iterations to calculate the sum of adjusted close values for n number of days which is the SMA value. At the end of the loop, we will append the SMA into `returnList`.
#
# After calculating all the SMA for every date in self.data, we insert the `returnList` containing all the SMA value with a column name stored in `col_head`. At the end of the function, we save the dataframe with SMA into a CSV file.
#
# ```
# 214 self.data[col_head] = returnList
# 216 self.data.to_csv(self.filepath, index=True)
# ```
# ## `calculate_crossover(self, SMAa, SMAb)`
# We first start by creating and defining the shell of the calculate_crossover function:
#
# ```
# 220 def calculate_crossover(self, SMAa,SMAb):
# ...
# 300 return self
# ```
# This function takes in the two SMA values previously calculated in the calculate_sma function as inputs to calculate the crossover locations.
#
# Next we will start to write the code inside the function. We first define the columns we plan to add to the .csv file and extract the all data in the .csv file:
#
# ```
# 244 col_head3 = 'Buy'
# 245 col_head4 = 'Sell'
# 246 df = self.data
# ```
# We convert the data into a list which we will use as a reference to ensure our subsequent calculations have the correct number of elements
#
# ```
# 249 SMAlist = self.data.index.values.tolist()
# ```
# We then use an if, elif, and else statement to assign the lower SMA to SMA1 from the and the higher SMA to SMA2. This is useful later in the calculations to ensure that buy and sell signals are correctly identified.
#
# ```
# 251 if SMAa < SMAb:
# 252 SMA1 = df[SMAa].tolist()
# 253 SMA2 = df[SMAb].tolist()
# 254 elif SMAa > SMAb:
# 255 SMA1 = df[SMAb].tolist()
# 256 SMA2 = df[SMAa].tolist()
# 257 else: # SMAa == SMAb
# 258 raise ValueError(f"Given {SMAa} & {SMAb} are the same. Must be different SMA.")
# ```
# > *Learning point: `if, elif, and else` statements*
#
# > *`elif` is used here because there are multiple distinct different possibilities with how SMAa and SMAb are related. It is common to list the expected possibilities first in the `if` and `elif` statements, and `else` would normally be reserved for unexpected outcomes or errors*
# `df.[SMAa].tolist()` extracts the column `SMAa` from the dataframe `df` and converts it to a list. Likewise for `df.[SMAb].tolist()`. If the two SMA values are equal, the code will raise a value error and the error message.
# We create empty lists for the relative position of the two SMAs (`stockPosition`), the combined list of crossover signals (`stockSignal`), and finally separate lists for the buy and sell signals (`buySignal`, `sellSignal`). These lists will be referenced and used in the next few lines of code.
#
# ```
# 260 stockPosition = []
# 261 stockSignal = []
# 262 buySignal = []
# 263 sellSignal = []
# ```
# To create a list of relative SMA positions, we use a `for` loop:
#
# ```
# 266 for i in range(len(SMAlist)):
# 267 if SMA1[i] > SMA2[i]: stockPosition.append(1)
# 268 elif SMA1[i] < SMA2[i]: stockPosition.append(0)
# 271 elif SMA1[i] == SMA2[i]: stockPosition.append(stockPosition[i-1])
# 272 else: stockPosition.append(np.nan)
# ```
# By setting the range of the for loop to be the length of `SMAlist`, we ensure that the loop iterates over every single element in the dataframe.
#
# Any day that `SMA1` (the smaller one) is higher than `SMA2` will add a `1` to the stockPosition list.
# Days where `SMA2` is higher than SMA1 will add a `0` to the `stockPosition` list. The end result will be a list of 1s and 0s showing which SMA is higher on any given day.
#
# In the unlikely case that the two SMA vaues are equal in a day, the number added will be a repeat of the previous day, as no crossover has occured yet.
#
# On days where either SMA is missing data, such as in the first few days when there is not enough data to compute the SMA, we will add `np.nan` to the list as a filler.
# After getting the full `stockPosition` list, we need to identify the days where crossover occurs. For this, another for loop is used:
#
# ```
# 275 for j in range(len(stockPosition)):
# 278 if j == 0: stockSignal.append(np.nan)
# 280 else: stockSignal.append(stockPosition[j] - stockPosition[j-1])
# ```
# Again we set the range for the loop to be the length of `stockPosition` to ensure the code iterates over every element.
#
# The `stockSignal` list 'lags' behind the stockPosition list by one day, hence we add a `np.nan` as the very first value in the list to align the `stockSignal` list with the `stockPosition` list and ensure that both lists have the same number of elements.
#
# Following that we take the difference between the stockPosition that day and the `stockPosition` the previous day to identify the locations of crossovers. Crossovers show up in the list as `1` for a buy signal, and a `-1` for sell signals. `0` indicates that there has been no crossover that day.
# > *Learning point: indexing*
#
# > *Remember that in python, sequences start with 0, not 1! Hence,* `j == 0` *just refers to the first element in the range*
# > *Learning point: `np.nan`*
#
# > *Remember that any arithmetic operation on `NaN` will result in `NaN`. This allows us to append the list with null values without generating a value error*
# The next step would be to filter out the buy and sell signals, which will be processed separately by the application:
#
# ```
# 283 for k in range(len(stockSignal)):
# 284 if stockSignal[k] == 1:
# 285 value = self.data[SMAa].tolist()[k]
# 286 buySignal.append(value)
# 287 else: buySignal.append(np.nan)
# 288
# 289 for k in range(len(stockSignal)):
# 290 if stockSignal[k] == -1:
# 291 value = self.data[SMAa].tolist()[k]
# 292 sellSignal.append(value)
# 293 else: sellSignal.append(np.nan)
# ```
# Using yet another set of for loops, we identify the crossover locations in the `stockSignal` list. At the crossover locations, we append the average SMA values of that particular day to the appropriate buy or sell list. This value will then be used as the y-axis value that the application uses to plot the crossover signals on the graph.
#
# The else condition appends `np.nan` to the list on days that do not contain the respective crossover signals, and ensures that the signals are correctly aligned to the dates where the crossover occurred.
# Finally, with the locations of buy and sell crossover signals, the function will append the buy and sell signals to the .csv file as new columns while also printing the results in the application:
#
# ```
# 295 self.data[col_head3] = buySignal
# 296 self.data[col_head4] = sellSignal
# 297
# 298 print(self.data)
# 299 self.data.to_csv(self.filepath, index=True)
# ```
# > *Learning point: Testing*
#
# > *The reason why the function prints the results is so we can independently test whether the function works even before the rest of the app is completed. Splitting work up in such a complex application is crucial so you can identify exactly which part of the app is causing errors!*
| report/stock_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Training for Stylizer module
# This notebook will handle the training the stylizer module.
# ## Load Pretrained model
# +
import matplotlib
matplotlib.use('Agg')
import os, sys
import yaml
from argparse import ArgumentParser
from time import gmtime, strftime
from shutil import copy
from frames_dataset import FramesDataset
from my_dataset import MyDataset
from modules.generator import OcclusionAwareGenerator # LCH: refer here for generator
from modules.discriminator import MultiScaleDiscriminator # LCH: refer here for discriminator
from modules.keypoint_detector import KPDetector # LCH: refer here for key point detector
import torch
import torch.nn.functional as F
from train import train # LCH: For training process, everything in this module
from reconstruction import reconstruction
from animate import animate
# +
config_path = "config/anim-256.yaml"
with open(config_path) as f:
# read in the config file
config = yaml.load(f) # config file contains code directions, including training details
checkpoint_path = "pre_trains/vox-cpk.pth.tar"
log_dir = "MyLog/"
if not os.path.exists(log_dir):
os.mkdir(log_dir)
# Copy the config file (*.yaml) into the logging path
if not os.path.exists(os.path.join(log_dir, os.path.basename(config_path))):
copy(config_path, log_dir)
# initialize generator
generator = OcclusionAwareGenerator(**config['model_params']['generator_params'],
**config['model_params']['common_params'])
# initialize discriminator
discriminator = MultiScaleDiscriminator(**config['model_params']['discriminator_params'],
**config['model_params']['common_params'])
# initialize kp detector
kp_detector = KPDetector(**config['model_params']['kp_detector_params'],
**config['model_params']['common_params'])
# If GPU Available, adapt to it
if torch.cuda.is_available():
print("using GPU")
generator.to(0)
discriminator.to(0)
kp_detector.to(0)
# +
# load in the pretrained modules
from logger import Logger
train_params = config['train_params']
if not torch.cuda.is_available():
# remember to adapt to cpu version
checkpoint = torch.load(checkpoint_path, map_location=torch.device('cpu'))
else:
checkpoint = torch.load(checkpoint_path)
generator.load_state_dict(checkpoint['generator'])
discriminator.load_state_dict(checkpoint['discriminator'])
kp_detector.load_state_dict(checkpoint['kp_detector'])
# The following models are used as data pre-processor
generator.eval()
discriminator.eval()
kp_detector.eval()
# -
# ## Dataset Preparation
# +
from frames_dataset import DatasetRepeater
from torch.utils.data import DataLoader
# load original target data
frame_dataset = FramesDataset(is_train=True, **config['dataset_params'])
print("Dataset size: {}, repeat number: {}".format(len(frame_dataset), config['train_params']['num_repeats']))
# load source data
source_dataset = FramesDataset(is_train=True, **config['target_params'])
print("Dataset size: {}, repeat number: {}".format(len(source_dataset), config['train_params']['num_repeats']))
# load combined data
combine_dataset = MyDataset(source_dir=config['target_params']['root_dir'], target_dir=config['dataset_params']['root_dir'],
frame_shape=config['dataset_params']['frame_shape'], id_sampling=config['dataset_params']['id_sampling'],
is_train=True, augmentation_params=config['dataset_params']['augmentation_params'])
print("Dataset size: {}, repeat number: {}".format(len(combine_dataset), config['train_params']['num_repeats']))
if 'num_repeats' in train_params or train_params['num_repeats'] != 1:
# Augment the dataset according to "num_reapeat"
frame_dataset = DatasetRepeater(frame_dataset, train_params['num_repeats'])
print("Repeated Target size: {}, repeat number: {}".format(len(frame_dataset), config['train_params']['num_repeats']))
source_dataset = DatasetRepeater(source_dataset, train_params['num_repeats'])
print("Repeated Source size: {}, repeat number: {}".format(len(source_dataset), config['train_params']['num_repeats']))
combine_dataset = DatasetRepeater(combine_dataset, train_params['num_repeats'])
print("Repeated Combin size: {}, repeat number: {}".format(len(combine_dataset), config['train_params']['num_repeats']))
targetLoader = DataLoader(frame_dataset, batch_size=train_params['batch_size'], shuffle=True, num_workers=2, drop_last=True)
sourceLoader = DataLoader(source_dataset, batch_size=train_params['batch_size'], shuffle=True, num_workers=2, drop_last=True)
combineLoader = DataLoader(combine_dataset, batch_size=train_params['batch_size'], shuffle=True, num_workers=2, drop_last=True)
# +
# declare a model
# declare objects needed by training process
from modules.stylizer import StylizerGenerator
from modules.stylizer_discriminator import StylizerDiscrim
# create network models
stylizer = StylizerGenerator(**config['model_params']['stylizer_params'])
styDiscrim = StylizerDiscrim(**config['model_params']['stylizerDiscrim_params'])
# create optimizers
lr_stylizer = 2.0e-4
lr_styDiscrim = 2.0e-4
optimizer_stylizer = torch.optim.Adam(stylizer.parameters(), lr=lr_stylizer, betas=(0.5, 0.999), weight_decay=1e-2)
optimizer_styDiscrim = torch.optim.Adam(styDiscrim.parameters(), lr=lr_styDiscrim, betas=(0.5, 0.999), weight_decay=1e-2)
# If GPU Available, adapt to it
if torch.cuda.is_available():
print("using GPU")
stylizer.to(0)
styDiscrim.to(0)
# +
# Train following the GAN network process
from tqdm import trange
from my_logger import MyLogger
from tqdm import tqdm
# read in the training parameters
stylize_params = config['stylize_params']
# declare a Logger
styLogger = MyLogger(log_dir, checkpoint_freq=stylize_params['checkpoint_freq'], log_file_name='log.txt')
# read in pre-trained results
# stylizer_checkpoint = "MyLog/checkpoints/0-50epo_onlyMSE/00000049-checkpoint.pth.tar"
stylizer_checkpoint = None
already_trained = 0
if stylizer_checkpoint:
already_trained = styLogger.load_cpk(stylizer_checkpoint, stylizer, styDiscrim, None, optimizer_styDiscrim)
already_trained += 1
# check the dense motion module
if generator.dense_motion_network is None:
print("Error: dense motion network doesn't exist!")
dm_network = generator.dense_motion_network # this model is used for extracting motion features
device = 'cuda' if torch.cuda.is_available() else 'cpu'
# loop over
for epoch in trange(already_trained, already_trained + stylize_params['num_epochs']):
for x in combineLoader:
# ------------------------------- Data Preprocess -------------------------------------#
# first get the key points for both source and driving frames
# Extract data of training data
x['source'] = x['source'].to(device)
kp_source = kp_detector(x['source'])
x['driving'] = x['driving'].to(device)
kp_driving = kp_detector(x['driving'])
# Extract data of target data
x['t_source'] = x['t_source'].to(device)
kp_t_source = kp_detector(x['t_source'])
x['t_driving'] = x['t_driving'].to(device)
kp_t_driving = kp_detector(x['t_driving'])
# second pass through the motion predictor
# plan A: get sparse motion as training data
if dm_network.scale_factor != 1:
src_image = dm_network.down(x['source'])
tar_image = dm_network.down(x['t_source'])
bs, _, h, w = src_image.shape
sparse_motion = dm_network.create_sparse_motions(src_image, kp_driving, kp_source)
sparse_motion_t = dm_network.create_sparse_motions(tar_image, kp_t_driving, kp_t_source) # motion from target data, serve as label
# here we don't need the last key point, which is a identity grid layer added by users
sparse_motion = sparse_motion[:, :-1, :, :, :] # The shape should be: torch.Size([10, 10, 64, 64, 2])
sparse_motion_t = sparse_motion_t[:, :-1, :, :, :]
orig_shape = sparse_motion.shape # record the orginal shape
# NOTE: BE CAREFUL HERE
# apply shape rearrange to the data
sparse_motion = sparse_motion.permute((0, 1, 4, 2, 3)) # NOTE: swap the dimension using INDEX of dimensions
sparse_motion_t = sparse_motion_t.permute((0, 1, 4, 2, 3))
# sparse_motion = sparse_motion.view((orig_shape[0], -1, orig_shape[2], orig_shape[3]))
# sparse_motion_t = sparse_motion_t.view((orig_shape[0], -1, orig_shape[2], orig_shape[3]))
sparse_motion = sparse_motion.reshape((orig_shape[0], orig_shape[1] * orig_shape[4], orig_shape[2], orig_shape[3]))
sparse_motion_t = sparse_motion_t.reshape((orig_shape[0], orig_shape[1] * orig_shape[4], orig_shape[2], orig_shape[3]))
# ------------------------------- Loss Calculation -------------------------------------#
# pass in stylizer network
loss_log = {}
# calculate losses for stylizer
stylized_dict = stylizer(sparse_motion.detach())
loss_values = {}
# for now, using MSE compared with original motion field, and GAN stylizer loss
mse_loss = F.mse_loss(sparse_motion, stylized_dict['prediction'])
loss_values['mse'] = mse_loss * stylize_params['loss_weights']['mse']
# GAN stylizer loss
# Compare real face data
if stylize_params['loss_weights']['gan'] != 0:
feature_real, discrim_real = styDiscrim(sparse_motion_t.detach()) # discriminate on target motion
feature_gene, discrim_gene = styDiscrim(stylized_dict['prediction']) # discriminate on source motion
gan_loss = ((1 - discrim_gene) ** 2).mean()
loss_values['gan'] = gan_loss * stylize_params['loss_weights']['gan']
# TODO: consider add feature matching loss
# Not suitable for this case, there's no one vs one correspondence between two data
if stylize_params['loss_weights']['match'] != 0:
value_total = 0
for i, (a, b) in enumerate(zip(feature_real, feature_gene)):
# calculate feature matching loss for each scale
value = torch.abs(a - b).mean()
value_total += value
loss_values['match'] = value_total * stylize_params['loss_weights']['match']
# Now combine all loss values for stylizer, update stylizer
loss_log.update(loss_values) # this is for logging
loss = sum([val.mean() for val in loss_values.values()])
loss.backward()
optimizer_stylizer.step()
optimizer_stylizer.zero_grad()
# Now deal with discriminator training
if stylize_params['loss_weights']['discrim'] != 0:
loss_values = {}
optimizer_styDiscrim.zero_grad()
_, discrim_real = styDiscrim(sparse_motion_t.detach()) # data that should be judged as True
_, discrim_gene = styDiscrim(stylized_dict['prediction'].detach()) # data that should be judged as False
discrim_loss = (1 - discrim_real) ** 2 + discrim_gene ** 2
loss_values['discrim'] = discrim_loss.mean() * stylize_params['loss_weights']['discrim']
# combine losses for stylizer discriminator, update discriminator
loss_log.update(loss_values) # this is for logging
loss = sum([val.mean() for val in loss_values.values()])
loss.backward()
optimizer_styDiscrim.step()
optimizer_styDiscrim.zero_grad()
# End of iteration, do logging
loss_log = {key: value.mean().detach().data.cpu().numpy() for key, value in loss_log.items()}
styLogger.log_iter(losses=loss_log)
# End of an epoch, do logging
styLogger.log_epoch(epoch,
{'stylizer': stylizer, 'styDiscrim': styDiscrim, 'optimizer_stylizer': optimizer_stylizer, 'optimizer_styDiscrim': optimizer_styDiscrim},
orig_shape, sparse_motion, stylized_dict['prediction'])
# End of training, do logging
styLogger.plot_scores()
styLogger.save_cpk()
# -
# ## Model Embedding
# ## Code snippets part
# +
import numpy as np
import matplotlib.pyplot as plt
inp = np.ones((10, 10, 64, 64, 2))[0]
out = np.ones((10, 10, 64, 64, 2))[0]
shape = inp.shape # the shape should be (num_kp, 64, 64, 2)
fig, axes = plt.subplots(4, shape[0], figsize=(4 * shape[0], 4 * 4))
for kp, ax in enumerate(axes[0]):
# plot the original motion field
ax.set_xlim(0, shape[1])
ax.set_ylim(0, shape[2])
ax.set_title("kp{}".format(kp))
for i in range(shape[1]):
if i % 2 == 0:
continue
for j in range(shape[2]):
if j % 2 == 0:
continue
ax.arrow(i, j, *inp[kp, i, j], color='b', linewidth=0.5, head_width=0.2, head_length=0.2)
for kp, ax in enumerate(axes[1]):
# plot the generated motion field
ax.set_xlim(0, shape[1])
ax.set_ylim(0, shape[2])
ax.set_title("kp{}".format(kp))
for i in range(shape[1]):
if i % 2 == 0:
continue
for j in range(shape[2]):
if j % 2 == 0:
continue
ax.arrow(i, j, *out[kp, i, j], color='r', linewidth=0.5, head_width=0.2, head_length=0.2)
for kp, ax in enumerate(axes[2]):
# plot both original and generated motion fields
ax.set_xlim(0, shape[1])
ax.set_ylim(0, shape[2])
ax.set_title("kp{}".format(kp))
for i in range(shape[1]):
if i % 2 == 0:
continue
for j in range(shape[2]):
if j % 2 == 0:
continue
ax.arrow(i, j, *inp[kp, i, j], color='b', linewidth=0.5, head_width=0.2, head_length=0.2)
ax.arrow(i, j, *out[kp, i, j], color='r', linewidth=0.5, head_width=0.2, head_length=0.2)
for kp, ax in enumerate(axes[3]):
# plot difference motion fields
ax.set_xlim(0, shape[1])
ax.set_ylim(0, shape[2])
ax.set_title("kp{}".format(kp))
for i in range(shape[1]):
if i % 2 == 0:
continue
for j in range(shape[2]):
if j % 2 == 0:
continue
ax.arrow(i, j, *(out[kp, i, j] - inp[kp, i, j]), color='g', linewidth=0.5, head_width=0.2, head_length=0.2)
# save figs
plt.show()
# +
h, w = 4, 4
x = torch.arange(w).type(torch.float)
y = torch.arange(h).type(torch.float)
x = (2 * (x / (w - 1)) - 1)
y = (2 * (y / (h - 1)) - 1)
yy = y.view(-1, 1).repeat(1, w)
xx = x.view(1, -1).repeat(h, 1)
meshed = torch.cat([xx.unsqueeze_(2), yy.unsqueeze_(2)], 2)
print(meshed.shape)
meshed = meshed.view(1, 1, h, w, 2)
meshed.shape
# +
import numpy as np
a = np.array([[1,2,3], [4, 5, 6], [7, 8, 9]])
b = np.array([[10,11,12], [13, 14, 15], [16, 17, 18]])
c = np.array([a, b])
c.shape
d = np.transpose(c, (1, 2, 0))
d.shape
k = []
for i in range(10):
k.append(d)
k = np.array(k)
s = k.shape
k = k.transpose((0, 3, 1, 2))
k.shape
k = k.reshape((20, 3, 3))
k[1]
# -
| train_stylizer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="m7I35Gzz78o1" colab_type="text"
# # Modelling Solar generation across Multiple Sites
#
# This example shows how `timeserio` helps building deep learning models for time series forecasting. Especially,
# we deal with the case of many related timeseries.
#
# We demonstrate some core functionality and concepts, without striving for model accuracy or seeking out additional features like historic weather forecasts.
#
# We will be using the dataset on solar (photo-voltaic, PV) generation potential across Europe, as collected by [SETIS](https://setis.ec.europa.eu/EMHIRES-datasets). The dataset presents solar generation, normalized to the solar capacity installed as of 2015.
# + id="bV5HEW69o3vH" colab_type="code" outputId="07af3434-b124-4735-d98b-c87f7928f5d6" colab={"base_uri": "https://localhost:8080/", "height": 36}
# Restart runtime using 'Runtime' -> 'Restart runtime...'
# %tensorflow_version 1.x
import tensorflow as tf
print(tf.__version__)
# + id="BR8RNvXO78pj" colab_type="code" colab={}
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# + id="fKCWVUdYlcy4" colab_type="code" colab={}
from google.colab import files
import os
import os.path
from os import path
# + id="KPumjlwOBZmz" colab_type="code" colab={}
from google.colab import files
def upload_1_file():
uploaded = files.upload()
for fn in uploaded.keys():
print('User uploaded file "{name}" with length {length} bytes'.format(
name=fn, length=len(uploaded[fn])))
return fn
return ""
# + id="WbVwi1u1dpRA" colab_type="code" cellView="both" colab={}
def download_1_file(http_path):
import requests
import shutil
response = requests.get(http_path, stream=True)
import tempfile
fname = tempfile.mkstemp()[1]
#print(fname)
with open(fname, 'wb') as fin:
shutil.copyfileobj(response.raw, fin)
return fname # Works!
# + id="sH51WPU6jjb1" colab_type="code" colab={}
def download_or_upload_1_file(http_or_file_path):
if (path.exists(http_or_file_path) and (path.isfile(http_or_file_path))):
return http_or_file_path
if (http_or_file_path==""):
encoder_input_file=upload_1_file()
return encoder_input_file
else:
print(http_or_file_path)
return download_1_file(http_or_file_path)
# + [markdown] id="YFrPxsA578pi" colab_type="text"
# ## Load the data from parquet
# + id="b66yLlDxjgDp" colab_type="code" colab={}
selected_path="https://MrYingLee.Github.io/MultiModel/selected.parquet"
# + id="t5KrqfHOkAq4" colab_type="code" outputId="2e221c93-5d2e-42fe-a638-aafd29bc92d1" colab={"base_uri": "https://localhost:8080/", "height": 36}
data_file=download_or_upload_1_file(selected_path)
# + id="wkStn0qBBKdL" colab_type="code" outputId="189856d7-1fcb-4fc8-f30f-1217657b54ea" colab={"base_uri": "https://localhost:8080/", "height": 55}
# %%time
df = pd.read_parquet(data_file)
# + id="4O34kIqTp4D9" colab_type="code" outputId="cf3531d4-c93e-4841-c605-7f80b3dcd6db" colab={"base_uri": "https://localhost:8080/", "height": 355}
df.head(10)
# + [markdown] id="0RxnMQ-W78qM" colab_type="text"
# ## Split into train-test sets
# + id="xp8yaG6_78qN" colab_type="code" outputId="c9d5e562-4486-417d-9014-16d260d15695" colab={"base_uri": "https://localhost:8080/", "height": 36}
df_dev = df.iloc[:100]
df_train, df_test = df[df['Year'] < 2015], df[df['Year'] >= 2015]
len(df_train), len(df_test)
# + id="ehwOPzGDipPI" colab_type="code" outputId="610e53cf-c21e-45e6-af8c-6920586d2414" colab={"base_uri": "https://localhost:8080/", "height": 969}
df.groupby("Year").count().reset_index()
# + id="tpJHQuHjwUiW" colab_type="code" outputId="60e36899-b5fc-41c5-d9a8-c70197cd2dbd" colab={"base_uri": "https://localhost:8080/", "height": 55}
# %%time
df_test.to_parquet("/content/test.parquet")
# + id="UzQcIUAlCRGW" colab_type="code" outputId="1b70a58d-9ebd-4337-9f1f-d597807906aa" colab={"base_uri": "https://localhost:8080/", "height": 73}
# !ls /content -l
# + id="bF9otCaAwUij" colab_type="code" colab={}
#from google.colab import files
#files.download("/content/test.parquet")
# + [markdown] id="Dx7XECPPBKdb" colab_type="text"
# ## Auto-regressive model
#
# In an auto-regressive model, we treat past values of the timeseries as input features to the forecasting model.
# While the functional form of the model is important, deep learning frameworks give us an easy way to try different approaches including CNNs, RNNs, etc.
#
# A key part remains however - we must be able to supply abundant training examples, each consisting of a window of consecutive values, the target, and (optinally) the time between the end of the window and the target (the "forecast horizon"). A long timeseries can be used to generate many examples simply by sampling the windows randomly from the original timeseries - in fact, for a realistic timeseries, pre-generating training examples in memory is prohibitively expensive. `timeserio` provides a way to generate sequence training examples on-demand from data held in memory, or even from datasets partitioned into multiple files.
# + [markdown] id="yrDGFTlTDxqE" colab_type="text"
#
# In [Model 1], we have explored the SETIS PV generation dataset and built a powerful and performant model using
# `timeserio`'s `MultiModel` and datetime feature generation pipelines. In this part, we instead train an auto-regressive model using more advanced batch generator features.
#
# Remember the metrics our previous model achieved on the train/test split (without any parameter tuning):
#
# | | train | test |
# |---|---|---|
# | MSE | 0.0063 | 0.0068 |
# | MAE | 0.0401 | 0.0424 |
#
# In this notebook, we will build a simple model to create short-range predictions (between 1 and 2 hours ahead) based on recent history (say 6h)
# + id="LJWVwl0VmESV" colab_type="code" outputId="4e5d1359-ea87-4490-a93a-212a907b2da8" colab={"base_uri": "https://localhost:8080/", "height": 353}
# !pip install timeserio
# + id="MD0G0K_LBKdd" colab_type="code" outputId="3c462180-f9c6-40cb-d286-0d9f75d3d8c1" colab={"base_uri": "https://localhost:8080/", "height": 36}
from timeserio.batches.chunked.pandas import SequenceForecastBatchGenerator
batchgen_train = SequenceForecastBatchGenerator(
df=df_train, batch_size=2**15,
sequence_length=6,
sequence_columns=["generation", "Time_step"],
last_step_columns=["Time_step"],
forecast_steps_min=1,
forecast_steps_max=2,
batch_offset=True,
id_column="country",
batch_aggregator=1
)
# + id="kWqIg9-PBKdh" colab_type="code" outputId="ff21fd39-a569-4ed0-f2cd-50cff034da49" colab={"base_uri": "https://localhost:8080/", "height": 36}
len(batchgen_train)
# + id="hB2j9pKiBKdo" colab_type="code" outputId="61e5047a-edf9-4f93-f9b0-a15c4d00b916" colab={"base_uri": "https://localhost:8080/", "height": 55}
# %%time
batch = batchgen_train[0]
# + id="0j4v9p6TBKds" colab_type="code" outputId="2f343c69-7efb-46bc-e76f-de92d8bc2f20" colab={"base_uri": "https://localhost:8080/", "height": 141}
batch.head(2)
# + id="prX0tVORBKdy" colab_type="code" outputId="ae5d56bd-8acb-467b-8305-3c55bfc0c8cf" colab={"base_uri": "https://localhost:8080/", "height": 55}
# %%time
batch = batchgen_train[-1]
# + id="bCEsNwHeBKd5" colab_type="code" outputId="157f38a5-760d-4fee-c47a-1bd9ec2f8aa9" colab={"base_uri": "https://localhost:8080/", "height": 161}
batch.head(2)
# + [markdown] id="jslSgu-OBKd9" colab_type="text"
# ### Sequence and Forecast horizon features
# + id="lDYZ_oNEBKd-" colab_type="code" colab={}
from timeserio.pipeline import Pipeline
from timeserio.preprocessing import PandasColumnSelector, PandasValueSelector
class ColumnDifferenceValues:
"""Compute difference feature of two columns"""
def __init__(self, *, col_plus, col_minus):
self.col_plus = col_plus
self.col_minus = col_minus
def fit(self, *args, **kwargs):
return self
def fit_transform(self, df, *args, **kwargs):
return self.transform(df, *args, **kwargs)
def transform(self, df, *args, **kwargs):
return (df[self.col_plus] - df[self.col_minus]).values.reshape(-1, 1)
seq_pipeline = PandasValueSelector("back_generation")
fc_horizon_pipeline = ColumnDifferenceValues(col_plus="Time_step", col_minus="end_of_Time_step")
target_pipeline = PandasValueSelector("fore_generation")
# + [markdown] id="AxA61HJkBKeB" colab_type="text"
# ### Define the Neural Network Architecture
#
# We define a regression network with two inputs: sequence of previous readings, and the forecast horizon
# + id="Nvu1j_9LBKeC" colab_type="code" colab={}
from timeserio.keras.multinetwork import MultiNetworkBase
from keras.layers import Input, Dense, Flatten, Concatenate, Reshape, Permute, Conv1D, BatchNormalization, MaxPool1D, Activation
from keras.models import Model
from keras.optimizers import Adam
from keras.callbacks import EarlyStopping, ReduceLROnPlateau
class ARForecastingNetwork(MultiNetworkBase):
def _model(
self,
*,
seq_length=6, # number of real-valued features
filters=(1, ),
kernel_sizes=(1, ),
strides=(1, ),
pools=(1, ),
hidden_units=(8, 8),
lr=0.01
):
horizon_input = Input(shape=(1,), name='horizon')
seq_input = Input(shape=(seq_length,), name='sequence')
encoding = Reshape(
target_shape=(-1, 1)
)(seq_input)
for idx, (_filters, _kernel_size, _strides, _pool) in enumerate(zip(filters, kernel_sizes, strides, pools)):
encoding = Conv1D(filters=_filters, kernel_size=_kernel_size, strides=_strides, padding="same", name=f"conv_{idx}")(encoding)
encoding = BatchNormalization()(encoding)
encoding = Activation(activation='relu')(encoding)
encoding = MaxPool1D(pool_size=_pool)(encoding)
encoding = Flatten()(encoding)
output = Concatenate(name='concatenate')([encoding, horizon_input])
for idx, _hidden_units in enumerate(hidden_units):
output = Dense(_hidden_units, activation='relu', name=f'dense_{idx}')(output)
output = Dense(2, name='fore_generation', activation='relu')(output) # 1=>2
encoding_model = Model(seq_input, encoding)
forecasting_model = Model([seq_input, horizon_input], output)
optimizer = Adam(lr=lr)
forecasting_model.compile(optimizer=optimizer, loss='mse', metrics=['mae'])
return {'encoder': encoding_model, 'forecast': forecasting_model}
multinetwork = ARForecastingNetwork(seq_length=6, lr=0.001)
# + id="LnOlhZ0xBKeG" colab_type="code" colab={}
from keras.utils.vis_utils import model_to_dot
from IPython.display import SVG
def vis_model(model, show_shapes=False, show_layer_names=True, rankdir='TB'):
"""Visualize model in a notebook."""
return SVG(
model_to_dot(
model, show_shapes=show_shapes, show_layer_names=show_layer_names, rankdir=rankdir
).create(prog='dot', format='svg')
)
# + id="KnaGCVl8BKeM" colab_type="code" outputId="7755d222-6a28-48c4-80ec-b33e185f7598" colab={"base_uri": "https://localhost:8080/", "height": 785}
vis_model(multinetwork.model["forecast"], show_shapes=True, rankdir="LR")
# + [markdown] id="LAyAcdZZBKeQ" colab_type="text"
# ### Connect feature pipelines to the neural network
# + id="chARUG6WBKeR" colab_type="code" colab={}
from timeserio.pipeline import MultiPipeline
# + id="7fZpcQstBKeV" colab_type="code" colab={}
multipipeline = MultiPipeline({
"sequence": seq_pipeline,
"horizon": fc_horizon_pipeline,
"target": target_pipeline
})
# + id="6oAq1WYuBKea" colab_type="code" colab={}
from timeserio.multimodel import MultiModel
manifold = {
# keras_model_name: (input_pipes, output_pipes)
"encoder": ("sequence", None),
"forecast": (["sequence", "horizon"], "target")
}
multimodel = MultiModel(
multinetwork=multinetwork,
multipipeline=multipipeline,
manifold=manifold
)
# + [markdown] id="XMT83C6BBKed" colab_type="text"
# ### Fit model from the batch generator
#
# `multimodel.fit_generator()` will apply pipelines correctly to the training batch generator, and, if `validation_data` is provided in the form of another (pandas) batch generator,
# evaluate the relevant metrics. In addition, feature extraction for each batch will benefit from the `workers` parallelism.
# + id="L6cuAZXADihe" colab_type="code" outputId="76767281-9b3d-40b0-8bcd-5ed58ce35dc5" colab={"base_uri": "https://localhost:8080/", "height": 36}
# !pip install kerashistoryplot
# + [markdown] id="aVwK7Epz-oMR" colab_type="text"
# ### Rolling Generator
# + id="Nn6s34xY-nD0" colab_type="code" colab={}
# + id="zdf1oiTgGV1q" colab_type="code" colab={}
import numpy as np
import pandas as pd
import pytest
import timeserio.ini as ini
from timeserio.data.mock import mock_fit_data, mock_raw_data
from timeserio.batches.single.sequence import (
SamplingForecastBatchGenerator, SequenceForecastBatchGenerator, ForecastBatchGeneratorBase, BatchGenerator
)
from numpy.testing import assert_array_equal
# + id="Aypt_3kbJggA" colab_type="code" colab={}
import abc
import functools
from typing import Union
import numpy as np
from timeserio import ini
from timeserio.preprocessing.pandas import array_to_dataframe
from timeserio.batches.utils import ceiling_division
# + id="ZMyaMEkjiJZ0" colab_type="code" colab={}
class RollingSequenceBatchGenerator(BatchGenerator):
"""Generate batches of sequence forecast examples.
Assume single continuous timeseries.
"""
def __init__(
self,
*,
df,
batch_size: Union[None, int] = None,
lookback_steps=2, # The steps of look back (history)
lookback_columns=[ini.Columns.datetime, ini.Columns.target],
lookback_prefix='back_',
forecast_steps=1, # The steps of forecasting
forecast_columns=[ini.Columns.target],
forecast_prefix='fore_',
id_column=None,
last_step_columns=[ini.Columns.datetime],
last_step_prefix='end_of_',
dt_column=ini.Columns.datetime,
start_time=None
):
self.df = df
self.batch_size = batch_size
self.lookback_steps = lookback_steps
self.lookback_columns = lookback_columns
self.lookback_prefix = lookback_prefix
self.forecast_steps = forecast_steps
self.forecast_columns = forecast_columns
self.forecast_prefix = forecast_prefix
self.last_step_columns = last_step_columns
self.last_step_prefix = last_step_prefix
if lookback_columns:
if not lookback_prefix:
raise ValueError('`lookback_prefix` must be non-empty')
if last_step_columns:
if not (set(last_step_columns) <= set(lookback_columns)):
raise ValueError('`last_step_columns` must be a subset of '
'`lookback_columns`')
if not forecast_columns:
raise ValueError('`forecast_columns` must be non-empty')
if not (set(forecast_columns) <= set(lookback_columns)):
raise ValueError('`forecast_columns` must be a subset of '
'`lookback_columns`')
if not last_step_prefix:
raise ValueError('`last_step_prefix` must be non-empty')
if last_step_prefix == lookback_prefix:
raise ValueError('`last_step_prefix` must be '
'different from `lookback_prefix`')
self.dt_column = dt_column
self.start_time = start_time
@property
def num_points(self):
"""Return number of rows in original timeseries."""
return len(self.df)
@property # type: ignore
@functools.lru_cache(None)
def first_index(self):
if self.start_time is None:
return 0
times = self.df[self.dt_column].dt.time.values
first_idx = np.argmax(times == self.start_time)
if not first_idx and times[0] != self.start_time:
raise ValueError(f'Start time {self.start_time} not found in df')
return first_idx
@property
def num_examples(self):
"""Return number of examples to yield in one epoch."""
return max(0,self.num_points-self.lookback_steps - self.forecast_steps -self.first_index+1)
@property
def _eff_batch_size(self):
return self.batch_size or self.num_examples
def __len__(self):
return ceiling_division(self.num_examples, self._eff_batch_size)
def batch_seq_start_indices(self, batch_idx):
start_indices = np.arange(
self._eff_batch_size * batch_idx,
min(self._eff_batch_size * (batch_idx + 1), self.num_examples)
)
return self.first_index + start_indices
def _get_lookback_values(self, column, start_indices):
values = self.df[column].values
cols = [
values[start_indices + s] for s in range(self.lookback_steps)
]
seq_values = np.vstack(cols).T
return seq_values
def _get_forecast_values(self, column, start_indices):
values = self.df[column].values
cols = [
values[start_indices + self.lookback_steps + s ] for s in range(self.forecast_steps)
]
seq_values = np.vstack(cols).T
return seq_values
def __getitem__(self, batch_idx):
if not len(self):
raise IndexError('Batch index out of range: Empty batch generator')
batch_idx = batch_idx % len(self)
start_indices = self.batch_seq_start_indices(batch_idx)
batch_size = len(start_indices)
end_indices = start_indices + self.lookback_steps
fc_indices = end_indices
cols = []
lookback_columns = self.lookback_columns or []
forecast_columns = self.forecast_columns or []
last_step_columns = self.last_step_columns or []
cols = cols + lookback_columns
batch_df = self.df[cols].iloc[fc_indices].copy()
batch_df.reset_index(drop=True, inplace=True)
for column in lookback_columns:
seq_values = self._get_lookback_values(
column, start_indices
)
seq_col_name = self.lookback_prefix + column
batch_df = array_to_dataframe(
seq_values,
column=seq_col_name,
df=batch_df
)
#if forecast_steps>1:
for column in forecast_columns:
seq_values = self._get_forecast_values(
column, start_indices
)
seq_col_name = self.forecast_prefix + column
batch_df = array_to_dataframe(
seq_values,
column=seq_col_name,
df=batch_df
)
for column in last_step_columns:
seq_col_name = self.lookback_prefix + column
last_step_col_name = self.last_step_prefix + column
batch_df[last_step_col_name] = batch_df[seq_col_name].iloc[:, -1]
return batch_df
# + id="_oqEcw8vjY87" colab_type="code" colab={}
"""Generate batches from pandas DataFrame."""
from typing import Union
from timeserio.batches.chunked.base import ChunkedBatchGenerator
from timeserio import ini
single_sequence= RollingSequenceBatchGenerator
# + id="6zVbcbD5bi_D" colab_type="code" colab={}
class RollingForecastBatchGeneratorChunked(ChunkedBatchGenerator):
def __init__(
self,
*,
df,
batch_size: Union[None, int] = None,
lookback_steps=2, # The steps of look back (history)
lookback_columns=[ini.Columns.datetime, ini.Columns.target],
lookback_prefix='back_',
forecast_steps=1, # The steps of forecasting
forecast_columns=[ini.Columns.target],
forecast_prefix='fore_',
id_column=None,
last_step_columns=[ini.Columns.datetime],
last_step_prefix='end_of_',
dt_column=ini.Columns.datetime,
start_time=None,
batch_aggregator=1
):
super().__init__()
self.df = df
self.batch_size = batch_size
self.lookback_steps = lookback_steps
self.lookback_columns = lookback_columns
self.lookback_prefix = lookback_prefix
self.forecast_steps = forecast_steps
self.forecast_columns = forecast_columns
self.forecast_prefix = forecast_prefix
self.last_step_columns = last_step_columns
self.last_step_prefix = last_step_prefix
if lookback_columns:
if not lookback_prefix:
raise ValueError('`lookback_prefix` must be non-empty')
if last_step_columns:
if not (set(last_step_columns) <= set(lookback_columns)):
raise ValueError('`last_step_columns` must be a subset of '
'`lookback_columns`')
if not forecast_columns:
raise ValueError('`forecast_columns` must be non-empty')
if not (set(forecast_columns) <= set(lookback_columns)):
raise ValueError('`forecast_columns` must be a subset of '
'`lookback_columns`')
if not last_step_prefix:
raise ValueError('`last_step_prefix` must be non-empty')
if last_step_prefix == lookback_prefix:
raise ValueError('`last_step_prefix` must be '
'different from `lookback_prefix`')
self.id_column = id_column
self.dt_column = dt_column
self.start_time = start_time
self.subgens = [] # type: ignore
self.batch_aggregator = batch_aggregator
self.unique_ids = self.df[self.id_column].unique()
@property
def chunks(self):
return self.unique_ids
def make_subgen(self, chunk):
cust_id = chunk
subgen = single_sequence(
df=self.df[self.df[self.id_column] == cust_id],
batch_size=self.batch_size,
id_column=self.id_column,
last_step_columns=self.last_step_columns,
last_step_prefix=self.last_step_prefix,
dt_column=self.dt_column,
start_time=self.start_time,
lookback_steps=self.lookback_steps, # The steps of look back (history)
lookback_columns=self.lookback_columns,
lookback_prefix=self.lookback_prefix,
forecast_steps=self.forecast_steps, # The steps of forecasting
forecast_columns=self.forecast_columns,
forecast_prefix=self.forecast_prefix
)
return subgen
# + [markdown] id="WUROxEiPCMvG" colab_type="text"
# ### Rolling Generator declare
# + id="99muFhJ0-8ju" colab_type="code" colab={}
rolling_chunked=RollingForecastBatchGeneratorChunked(
df=df_train,id_column="country",lookback_steps=6,forecast_steps=2,lookback_columns=["generation", "Time_step"],
forecast_columns=["generation"],
last_step_columns=["Time_step"] ,batch_size=2**15,
)
# + id="HSsEo93RAc5X" colab_type="code" outputId="5bc8be51-dee4-4c5f-d5ad-2d76701c4329" colab={"base_uri": "https://localhost:8080/", "height": 36}
rolling_chunked.chunks
# + id="joeUeh7sAgm9" colab_type="code" outputId="91d39c6f-8e64-4db2-b12f-d2838a1f9594" colab={"base_uri": "https://localhost:8080/", "height": 466}
sub0=rolling_chunked.make_subgen('UK')
sub0[0]
# + id="n49WMi0rBKee" colab_type="code" outputId="4322da9c-107a-4406-a38a-5d4879c23dcc" colab={"base_uri": "https://localhost:8080/", "height": 504}
from kerashistoryplot.callbacks import PlotHistory
#plot_callback = PlotHistory(figsize=(15, 3), n_cols=3, batches=False)
multimodel.fit_generator(
rolling_chunked, model="forecast", verbose=1, epochs=10,
reset_weights=True,
workers=4
#, callbacks=[plot_callback]
)
# + [markdown] id="VfOTeLQ6BKej" colab_type="text"
# persist the model:
# + id="ad8xS6LXmF0w" colab_type="code" colab={}
model_file="/tmp/model.pickle"
network_file="/tmp/network.pickle"
weights_file="/tmp/weights.pickle"
# + id="pmOgE2qFBKek" colab_type="code" colab={}
from timeserio.utils.pickle import loadf, dumpf
dumpf(multimodel, model_file)
dumpf(multinetwork, network_file)
dumpf(multinetwork.weights, weights_file)
# + id="qTpxWNznlsHx" colab_type="code" colab={}
from google.colab import files
files.download(model_file)
files.download(network_file)
files.download(weights_file)
# + id="TEpoZ9dUTTRp" colab_type="code" outputId="50fa5a45-9589-460d-edfe-587734929f94" colab={"base_uri": "https://localhost:8080/", "height": 1000}
multimodel.multinetwork.weights
# + [markdown] id="sEvo9LMxBKep" colab_type="text"
# ### Evaluate performance on test data
# We can evaluate the model on the validation data generator, which can also be out-of-memory:
# + id="Z2WZjvmwBKes" colab_type="code" colab={}
rolling_chunked_test=RollingForecastBatchGeneratorChunked(
df=df_test,id_column="country",lookback_steps=6,forecast_steps=2,lookback_columns=["generation", "Time_step"],
forecast_columns=["generation", "Time_step"],
last_step_columns=["Time_step"],batch_size=2**15,
)
# + id="QOBayckeBKev" colab_type="code" outputId="81317c8f-558f-4949-90a5-cc2d9f6083af" colab={"base_uri": "https://localhost:8080/", "height": 55}
multimodel.evaluate_generator(rolling_chunked_test, model="forecast", verbose=1)
# + [markdown] id="bp7Z1fsMBKey" colab_type="text"
# While the model takes longer to train (and longer still with practical encoder architectures), it can be tuned to achieve higher performanec, especially if encodings are combined with datetime features.
# + id="XdJBSf6QHNGx" colab_type="code" outputId="45afe2b1-d144-4a3e-e761-33de91a2cfbe" colab={"base_uri": "https://localhost:8080/", "height": 225}
from kerashistoryplot.plot import plot_history
history = multimodel.history[-1]["history"]
plot_history(history, figsize=(15, 3), n_cols=3);
| MultiModel/2_Solar_Generation_with_rolling_sequence.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Coin flipping
#
# ***
#
# The ideas behind flipping a coin are important in computing.
#
# Computers rely on bits - a bit is a variable that can take on one of two values, one or zero.
#
# Flipping a coin results in one of two outcomes, heads or tails.
#
# ***
# +
# We'll use numpy and scipy.stats to analyse flipping a coin.
import numpy as np
import scipy.stats as ss
# We'll use this for visualisation.
import matplotlib.pyplot as plt
import seaborn as sns
# -
# This just sets the default plot size to be bigger.
plt.rcParams['figure.figsize'] = (12, 8)
# ***
#
# A fair coin will give a head fifty percent of the time and a tail fifty percent of the time.
#
# We say the probability of a head is 0.5 and the probability of a tail is 0.5.
#
# We can use the following function to simulate this - giving a 1 for a head and 0 for a tail.
#
# ***
# (Number of times to flip a coin, probability of a head, number of times to do this)
np.random.binomial(1, 0.5, 1)
# Flip a fair coin 1000 times - how many heads?
np.random.binomial(1000, 0.5, 1)
# ***
#
# How likely are we to see a certain number of heads when flipping a coin however many times?
#
# ***
# (No. of heads, no. of flips, probability of a head)
ss.binom.pmf(521, 1000, 0.5)
sns.distplot(np.random.binomial(1000, 0.5, 1000))
# ***
#
# What about an unfair coin?
#
# ***
# Flip an unfair coin 10 times - how many heads?
np.random.binomial(10, 0.2, 1)
# ***
#
# Suppose we flip an unfair coin ($p = 0.3$) ten times, what is the probability that the flips are as follows?
#
# $$ HHTTHHHTTT $$
#
# ***
(0.3)*(0.3)*(1.0-0.3)*(1.0-0.3)*(0.3)*(0.3)*(0.3)*(1.0-0.3)*(1.0-0.3)*(1.0-0.3)
# ***
# The probability of $r$ heads when flipping an unfair coin $n$ times is
#
# $$ p(r \mid n , p) = {n \choose r} p^r (1-p)^{(n-r)} $$
#
# ***
# +
noflips = 10
p = 0.3
d = [ss.binom.pmf(i, noflips, p) for i in range(noflips+1)]
d
# -
# ***
#
# $ {n \choose r} $ is the number of ways to select $r$ items from $n$, ignoring the order you select them in.
#
# ***
# +
import math
n = 10
r = 6
choose = lambda x, y: math.factorial(x) / (math.factorial(y) * math.factorial(x-y))
choose(n, r)
# -
# ***
#
# Note the following for ${n \choose 0}$ and ${n \choose n}$.
#
# ***
choose(10, 0)
choose(n, n)
# ***
#
# Even though the chances are, with $p = 0.3$ and $10$ flips, that there are three heads, the most probable outcome is all tails.
#
# ***
(1-0.3)**10
# ***
#
# What has all of this got to do with computers and bits?
#
# Would you consider the following a data set?
#
# ***
import itertools
["".join(seq) for seq in itertools.product("01", repeat=10)]
# ## End
| coin-flip.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Topics:
# * Grouping Data
# * Applying Custom Functions
# # Grouping Data
import numpy as np
import pandas as pd
df = pd.DataFrame(data=[['MI','P1','Male',54,15],
['MI','P2','Female',21,19],
['DD','P3','Male',69,26],
['RR','P4','Female',96,28],
['GT','P5','Male',33,24],
['MI','P6','Female',51,33],
['KNR','P7','Male',24,40],
['GT','P8','Male',36,42],
['RR','P9','Female',78,19],
['KNR','P10','Male',33,17],
['MI','P11','Female',87,20],
['GT','P12','Male',81,21],
['KNR','P13','Female',36,29]],
columns=['Team', 'Player', 'Sex', 'Score', 'Age'])
df
df['Team'].value_counts()
#Show count of players team-wise
df['Team'].unique()
#Gives total unique teams
df['Team'].nunique()
#Show count of unique teams
df['Sex'].value_counts()
byTeam = df.groupby('Team')
#Returns DataFrameGroupBy object
byTeam.sum()
#Sum of scores team-wise
byTeam.mean()
# +
#Pandas calculates sum and mean only on numeric values (score, age) and not other types(player, sex)
# -
df.groupby('Sex').mean()
df.groupby('Team').describe()
#Show various statistical measures
df.groupby('Sex').describe()
# # Applying Custom Functions
df
def congrats(x):
if x > 70:
return "Congratulations "
else:
return ""
df['Score'].apply(congrats)
df['Player'] = df['Score'].apply(congrats) + df['Player']
df
| Data Analysis with Pandas/Grouping Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="0k3TWCdmLepH"
# This is a Word2Vec embedding model with skip-gram and negative sampling method using pure tensorflow 2.
# + [markdown] id="7B3P803wJqos"
# # Making Data Ready
# + id="0KXm_ruQHxds" outputId="421ae833-044a-43e9-ee97-3d6a47878944" colab={"base_uri": "https://localhost:8080/"}
# !pip install tqdm # This is for showing a smart progress meter in any loop
# + id="2EIGNtWfmsMc" outputId="df86aed0-4235-4cdc-8a87-fbf10765b810" colab={"base_uri": "https://localhost:8080/"}
from tqdm.auto import tqdm # This is for showing a smart progress meter in any loop
import gensim.downloader as api
corpus = api.load("text8") # This is tokenized corpus of text from wikipedia
# + id="N11j9MjKmtr3" outputId="e63ae7cb-742f-4408-e24a-fab22f98960d" colab={"base_uri": "https://localhost:8080/"}
# just to show what is inside the corpus
for i in corpus:
print(i[:20])
break
# + [markdown] id="r16RnsXrK3p8"
# Now we want to build our skip-grams which means we should pair every word with its neighbours. For defining neighbour we choose a `window_size`.
#
# For example suppose the sentense "*I want a new car*". its skip-grams with `window_size = 1` would be:
#
# * I , want
# * want , I
# * want , a
# * a , want
# * a , new
# * new , a
# * new , car
# * car , new
#
# And with `window_size = 2` words near `new` would be:
#
# * new , car
# * new , want
# * new , a
#
#
#
#
#
#
#
#
#
#
# + id="2lXK6xCqncVW"
window_size = 2
# + id="cuAxhxnK5nc_" outputId="4f1ceaf7-80da-4d43-fc99-bf53a6cb1305" colab={"base_uri": "https://localhost:8080/", "height": 104, "referenced_widgets": ["e301fbbcddbc4c1d9295620e5bff45b0", "b57aa662139d4efcbc2f7e7a8857a483", "affd720a78754ba5874b6b7b36414344", "1f616f1b81cc447b8c39d48317f14852", "845058b7baba4706a163973585f348cc", "601be8a1b65a42f584377bd7dec1ccb2", "233d898d4eb44a8d88295a058ab34ec2", "a1aae7c823f042ceab8e9030ccfd4e4f"]}
# Now we should build our skip-grams and find neoghbour words, e.g. {('modern', 'era'), ('means', 'to'), ('economics', 'there'), ('while', 'is'),...}
from collections import Counter
neighbour_counter = Counter()
for text in tqdm(corpus):
dump_txt = ['_TEMP_'] * window_size + text + ['_TEMP_'] * window_size # e.g. sentence "I love code" with window_size = 2 would be "_TEMP_ _TEMP_ I love code _TEMP_ _TEMP_"
for i in range(len(dump_txt) - (window_size + 1) * 2):
middle_window = dump_txt[window_size + i]
for j in range(window_size): # we choose words near the center word as its nighbours (context words)
location = window_size + i - 1
if not middle_window == dump_txt[location - j] and not dump_txt[location - j] == '_TEMP_': # This is just for making sure neighbours aren't the same, like (the, the) and we ignore it when it becomes naighbour to "_TEMP_"
neighbour_counter[(middle_window, dump_txt[location - j])] += 1
if not middle_window == dump_txt[location + j] and not dump_txt[location + j] == '_TEMP_': # This is just for making sure neighbours aren't the same, like (the, the) and we ignore it when it becomes naighbour to "_TEMP_"
neighbour_counter[(middle_window, dump_txt[location + j])] += 1
print(len(neighbour_counter))
print(neighbour_counter.most_common(3))
# + id="Yk8k5kOf8WtP" outputId="bc812ee1-231a-4293-da57-abac8b94828e" colab={"base_uri": "https://localhost:8080/"}
neighbours = [i[0] for i in neighbour_counter.items() if i[1] > 5] # Removing neighbours that repeated less than 5 time in our large corpus. They are not useful.
len(neighbours)
# + id="4Q81Z4-ulUNu" outputId="a7fee168-69f0-4b3f-8b04-9d13fdfbe49e" colab={"base_uri": "https://localhost:8080/"}
from collections import defaultdict
dict_of_neighbours = defaultdict(set) # This dict contains any naighbour that a word has in our data
for i, j in neighbours:
dict_of_neighbours[i].add(j)
len(dict_of_neighbours['cat'])
# + id="rGTkFrdnm0yf" outputId="c9886dc9-e464-4e76-f40b-c8dfcfabf1d8" colab={"base_uri": "https://localhost:8080/"}
words_set = list(set([j for i in neighbours for j in (i[0], i[1])])) # list of all tokens
print(len(words_set))
index2word = dict(enumerate(list(words_set))) # A dict that maps an index to a word, e.g. d[3] = 'cat'
word2index = {v: k for k, v in index2word.items()} # Reverse of the dict above. e.g. d['cat'] = 3
# + [markdown] id="-ZGsCdscVuIE"
# Now we should make negative smapling. Which means we should produce fake nighbours. We then lable the true tuple with 1 and fake tuples with 0.
#
# For producing fake neighbours we just randomly sample from our `words_lst`. Do not worry about it randomly choose a true nighbour and not a fake one. We do not need to be perfect for now.
# + id="5mxpWqMCXmCS"
negative_samples = 5
# + id="oQgv-3ZjoSme"
import random
def negative_sampling(true_neighbours): # We lable true_neighbours with 1, and we produce fake neighbours and lable them with 0
center_word = true_neighbours[0]
neighbours_labeled = [(true_neighbours[1], 1)]
for random_word in random.sample(words_set, negative_samples):
neighbours_labeled.append((random_word , 0))
random.shuffle(neighbours_labeled) # Shuffle to make the order random
x, y = zip(*neighbours_labeled) # Now Xs are neighbour and Ys are thir lables (0 or 1)
return center_word, x, y
# + id="Ja1PuZWxkSRf" outputId="ff21591c-f0fe-4d40-b680-4ef70cea9943" colab={"base_uri": "https://localhost:8080/"}
negative_sampling(("cool", "code"))
# + [markdown] id="x6HHhgUSbQeA"
# Now The Final step to have our data ready! (Yoo Hoo!).
#
# We should make a function that return a batch of data everytime we call it. we rather replacethe words with thir indices here (by using `word2index`)
# + id="YIgMBlkYac3s"
def give_us_data(batch_size):
batched_center = []
batched_x = []
batched_y = []
for random_neighbour in random.sample(neighbours, batch_size):
center_word, x , y = negative_sampling(random_neighbour)
batched_y.append(y)
batched_center.append(word2index[center_word]) # Convert the center word to its index (e.g. "cat" to 3)
batched_x.append([word2index[i] for i in x]) # Convert words to its index (e.g. "cat" to 3)
return batched_center, batched_x, batched_y
# + id="UbaWcUj39TLD" outputId="ae439a82-ac61-475c-ea57-e4268bb2fc2a" colab={"base_uri": "https://localhost:8080/"}
give_us_data(2)
# + [markdown] id="7ru-jZjURy9n"
# # MODEL
#
# Now is the time to code the model with pure TF 2
# + id="pXwD9S2OrTEI"
import tensorflow as tf
import numpy as np
embeding_size = 50
batch_size = 128
# + id="kGRBpAK9rgaf"
target_amb = tf.keras.layers.Embedding(len(words_set), embeding_size) # This will be our Word2Vec
context_amb = tf.keras.layers.Embedding(len(words_set), embeding_size) # This is just for the learning phase
optimizer = tf.keras.optimizers.Adam()
# + id="W1mk45uH1GHb" outputId="63d5c872-7f65-4857-eed2-8c59b8d440d6" colab={"base_uri": "https://localhost:8080/", "height": 449, "referenced_widgets": ["fefb7410cb9d42f6b2e7175f047fae02", "60203681914241159d9f211802bf4ca0", "28c2b357616f4b528b46297b252a2e76", "833251bdd02046ba8593f25f5d214a5f", "<KEY>", "<KEY>", "ee6951d02d04410abc3c8fa3273fe254", "117c67d957074b1ebc66c70f88e7cf24"]}
for _ in tqdm(range(10001)): # tqdm is just for a nice progress bar.
center, x, y = give_us_data(batch_size)
center = np.asarray(center)
x = np.asarray(x)
y = np.asarray(y)
with tf.GradientTape() as t:
center_embs = target_amb(center)
neighbor_choices = context_amb(x)
scores = tf.keras.backend.batch_dot(neighbor_choices, center_embs, axes=(2,1)) # dot prudoct vectors of naighbours together
prediction = tf.nn.sigmoid(scores) # we want the model to give us a probability of that two, being real naighbours
loss = tf.keras.losses.categorical_crossentropy(y, prediction)
if not _ % 2000:
print("batch:", _ , " - The mean loss is: " ,tf.reduce_mean(loss).numpy())
print(y[1], prediction[1].numpy())
print("------------------------------------------------")
g_embed, g_context = t.gradient(loss, [target_amb.embeddings, context_amb.embeddings])
optimizer.apply_gradients(zip([g_embed, g_context], [target_amb.embeddings, context_amb.embeddings]))
# + [markdown] id="KSs_zCkqSK9F"
# Wooo Hooo! Congratulation!
#
# Now we have our Word2Vec ready in `target_amb`. Lets analyse is a little bit.
# + [markdown] id="gnhdgX38SjxQ"
# # Model analysis
#
# Here I want to write a function that finds closest words to a word
# + id="vxTzKaq_6rIp"
def find_closest(embeds, word, n=1): # n is for "n closest words"
n = n + 1 # This is becuse the most similar word is definatly that word itself. like the most similar word for "apple" is "apple". so we should look for top n+1 words
main_vec = embeds(word2index[word])
similarities = -tf.keras.losses.cosine_similarity(embeds.embeddings, main_vec)
top_n = tf.math.top_k(similarities, n).indices
words = [index2word[i] for i in top_n.numpy()]
return words[1:] # I did [1:] to remove the word as I mentioned in `n = n + 1` comments
# + id="yyJ0FPcPd0yd" outputId="674c8caf-3fec-4827-c7ce-596b11e49841" colab={"base_uri": "https://localhost:8080/"}
find_closest(target_amb, "two", 10)
| Word2vec.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# # !pip install wordcloud
# +
import csv
from wordcloud import WordCloud
#read first column of csv file to string of words seperated
#by tab
your_list = []
with open('Data-jobs-usa_entryLev_last14_fulltime_.csv', 'rb') as f:
reader = csv.reader(f)
your_list = '\t'.join([i[0] for i in reader])
# Generate a word cloud image
wordcloud = WordCloud().generate(your_list)
# Display the generated image:
# the matplotlib way:
import matplotlib.pyplot as plt
plt.imshow(wordcloud, interpolation='bilinear')
plt.axis("off")
# lower max_font_size
wordcloud = WordCloud(max_font_size=40).generate(your_list)
plt.figure()
plt.imshow(wordcloud, interpolation="bilinear")
plt.axis("off")
plt.show()
# -
| Codes/.ipynb_checkpoints/word_cloud-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# +
import sys
from pathlib import Path
curr_path = str(Path().absolute()) # 当前文件所在绝对路径
parent_path = str(Path().absolute().parent) # 父路径
sys.path.append(parent_path) # 添加路径到系统路径
import numpy as np
# -
# ## 计算高斯核
#
# 高斯核数学表达为:$K(x, x')=\exp\left(-\gamma\|x-x'\|^{2}\right)$
# 用矩阵表达则为:$K(i, j)=\exp\left(-\gamma\|x_i-x_j\|^{2}\right)$
# 其中数据集矩阵$x$的维度为$(m,)$,则高斯核$K$的维度为$(m,m)$
#
def calc_kernel(X_train):
'''
计算核函数
使用的是高斯核 详见“7.3.3 常用核函数” 式7.90
:return: 高斯核矩阵
'''
#初始化高斯核结果矩阵 大小 = 训练集长度m * 训练集长度m
# k[i][j] = Xi * Xj
m = X_train.shape[0]
k = [[0 for i in range(m)] for j in range(m)]
#大循环遍历Xi,Xi为式7.90中的x
for i in range(m):
#得到式7.90中的X
X = X_train[i, :]
#小循环遍历Xj,Xj为式7.90中的Z
# 由于 Xi * Xj 等于 Xj * Xi,一次计算得到的结果可以
# 同时放在k[i][j]和k[j][i]中,这样一个矩阵只需要计算一半即可
#所以小循环直接从i开始
for j in range(i, m):
#获得Z
Z = X_train[j, :]
#先计算||X - Z||^2
result = (X - Z) * (X - Z).T
#分子除以分母后去指数,得到的即为高斯核结果
result = np.exp(-1 * result / (2 * self.sigma**2))
#将Xi*Xj的结果存放入k[i][j]和k[j][i]中
k[i][j] = result
k[j][i] = result
#返回高斯核矩阵
return k
| ml-with-numpy/SVM/SVM_np.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="AEjwD6ekwcZW" colab_type="code" colab={}
# !mkdir work
# !cd work
# !git clone https://github.com/UniversalDependencies/UD_Japanese-GSD
# !git clone https://github.com/uehara1414/japanize-matplotlib
# + id="h7NJyVs9woFL" colab_type="code" colab={}
# !pip install nagisa
# !pip install wordcloud
# + id="uI1hhAhPwvOP" colab_type="code" colab={}
import nagisa
def write_file(fn_in, fn_out):
with open(fn_in, "r") as f:
data = []
words = []
postags = []
for line in f:
line = line.strip()
if len(line) > 0:
prefix = line[0]
if prefix != "#":
tokens = line.split("\t")
word = tokens[1]
postag = tokens[3]
words.append(word)
postags.append(postag)
else:
if (len(words) > 0) and (len(postags) > 0):
data.append([words, postags])
words = []
postags = []
with open(fn_out, "w") as f:
for words, postags in data:
for word, postag in zip(words, postags):
f.write("\t".join([word, postag])+"\n")
f.write("EOS\n")
# files
fn_in_train = "UD_Japanese-GSD/ja_gsd-ud-train.conllu"
fn_out_train = "ja_gsd_ud.train"
# write files for nagisa
write_file(fn_in_train, fn_out_train)
# + id="i3FLc3x6w5JY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 337} outputId="5ce821b0-622c-4dbd-b212-4e1bce691488"
from wordcloud import WordCloud
from IPython.display import Image,display_png
# load a file
fn_in = "ja_gsd_ud.train"
X, Y = nagisa.utils.load_file(fn_in)
# word segmentation
text = ""
for x in X:
s = "".join(x)
nouns = nagisa.extract(s, extract_postags=['名詞'])
text += " ".join(nouns.words)
# make a word cloud
stopwords = ["こと", "ため", "もの", "とも"]
ja_font = "japanize-matplotlib/japanize_matplotlib/fonts/ipaexg.ttf"
wordcloud = WordCloud(width=480, height=320, max_words=50, font_path=ja_font,
stopwords=stopwords, background_color='white', prefer_horizontal=1.)
wordcloud.generate(text)
wordcloud.to_file('wordcloud.png')
display_png(Image('wordcloud.png'))
| notebooks/word_cloud.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to Matplotlib (tutorial)
# This notebook gives a short introduction to *Matplotlib*, Python's most popular package for plotting. Although many different plotting packages exist in the Python ecosystem (see [this talk](https://www.youtube.com/watch?v=FytuB8nFHPQ) for an overview), Matplotlib is arguably the most versatile and flexible. Here, we will give you a short tour of Matplotlib's most important features.
#
# ## Contents
# 1. The state-based approach
# 2. The object-oriented approach
# 3. Subplots (optional)
#
# Most of the plotting functionality is contained in the subpackage `pyplot`, which is usually imported as follows:
import matplotlib.pyplot as plt # nice and short
# Somewhat confusingly, Matplotlib has two interfaces for plotting: a state-based approach that mimicks Matlab's way of plotting and a more "Pythonic" object-oriented approach. As Matplotlib recommends using the object-oriented approach, we will spend most time on this approach. But you'll often see the state-based approach as well, so we'll start with quickly discussing this approach.
# ## The state-based approach
# As mentioned, the state-based approach is a lot like the way plotting is done in Matlab: you call different *functions* that each take care of an aspect of the plot. In Matplotlib, most of these functions are contained in the `pyplot` package. Let's create simple line plot to show how the state-based approach looks like:
# +
x = [0, 1, 2, 3, 4, 5, 6]
y = [1, 2, 3, 4, 5, 6, 7]
plt.plot(x, y) # plot the data
plt.xlabel('x', fontsize=25) # set the x-axis label
plt.ylabel('y', fontsize=25) # set the y-axis label
plt.show() # this actually visualizes the plot
# -
# As you can see, the state-based approach entails a series of function calls (such as `plt.plot` and `plt.xlabel`). After you are done plotting, you just call `plt.show` and the plot will show in your notebook (or an external image viewer if you run it from a script). Note that, technically, the `plt.show` call is not necessary to render the plot in Jupyter notebooks, but we recommend doing it anyway as this is good practice.
#
# The `plt.plot` function is perhaps the most basic function, which can be used to create any plot of paired datapoints (x, y). By default, it creates a line plot (as shown above), but the many (optional) parameters in `plt.plot` allow you to create many different variations! For example, instead of a line, we can plot the data as separate red points by specifying the format in the third argument (here, 'o' to indicate points) and the color (by setting the argument `c`, for **c**olor, to "red"):
plt.plot(x, y, 'o', c='red')
plt.show()
# <div class='alert alert-success'>
# <b>Tip</b>: Note that the third argument, the "format", may be used to specify three things at once: whether you want "markers" (and which type of marker), whether you want a line (and which type of line), and which color the markers/line should have. So, to create red markers, you may specify "or" ("o" for circles as markers and "r" for red). To create a blue line, you may specify "-b". To create a yellow ("y") dotted line ("-.") with stars ("*") as markers, you may use "*-.y".
# </div>
# <div class='alert alert-warning'>
# <b>ToDo</b>: Create the same plot as above, but with a green dashed line with diamonds as markers. Check out the <a href="https://matplotlib.org/api/_as_gen/matplotlib.pyplot.plot.html">plt.plot</a> documentation under "Notes" which particular linestyle you need for this! And check <a href="https://matplotlib.org/api/markers_api.html">this page</a> to see the name of the marker for diamonds! Make sure to write your code <b>above</b> the "<tt>ax2check = plt.gca()</tt>" snippet — we use this to check your plot automatically in the test cell! Also, do not include a <tt>plt.show</tt> call; this is done after the <tt>ax2check</tt> line (calling <tt>plt.show</tt> before <tt>ax2check</tt> will cause to test cell to malfunction).
# </div>
# + nbgrader={"grade": false, "grade_id": "cell-b92e4b8229777658", "locked": false, "schema_version": 3, "solution": true, "task": false} tags=["hide-cell"]
""" Implement the ToDo here. """
### BEGIN SOLUTION
plt.plot(x, y, 'D--g')
### END SOLUTION
# Do not remove the code below!
ax2check = plt.gca()
plt.show()
# + nbgrader={"grade": true, "grade_id": "cell-2ee14e4f54b8e433", "locked": true, "points": 0, "schema_version": 3, "solution": false, "task": false}
""" Tests the ToDo above. """
line = ax2check.get_lines()[0]
if not line._color in ['g', 'green']:
raise ValueError("The line is not green!")
if line._linestyle != '--':
raise ValueError("You didn't use a dashed line!")
if line._marker._marker != 'D':
raise ValueError("You didn't use a dashed line!")
print("Yay! Well done.")
# -
# You can also plot multiple things within a single plot! Just call the `plt.plot` (or any other plotting function) multiple times. Below, we create a new variable (`y_sq`, the values of squared) and plot it in the same plot as our regular (x, y) plot. Importantly, we will include legend with the plot showing what each line represents using `plt.legend`:
# +
y_sq = [yi ** 2 for yi in y] # check out this list comprehension!
plt.plot(x, y, '*b') # only plot markers (*) in blue
plt.plot(x, y_sq, '^--y') # plot both markers (^) and a line (--) in yellow
# Note that the plt.legend function call should come *after* the plotting calls
# and you should give it a *list* with strings
plt.legend(['y', 'y squared'])
plt.show()
# -
# <div class='alert alert-danger'>
# <b>Warning</b>: Importantly, in the context of Jupyter notebooks, each part of the plot should be defined in the same code cell; otherwise, they won't be included in the same figure.
# </div>
# As you can see, Matplotlib automatically creates the right legend! Make sure that the order of your labels (here: `['y', 'y squared']`) matches the order of your plotting calls!
# <div class='alert alert-success'>
# <b>Good to know</b>: If you plot multiple things in the same plot, Matplotlib will automatically choose a different color for the different things (first one is blue, second one is orange, third one is green, etc.).
# </div>
# <div class='alert alert-warning'>
# <b>ToDo</b>: Below, we define some new variables: <tt>z</tt>, the sine of z (<tt>sin_z</tt>), and the cosine of z (<tt>cos_z</tt>). Plot both the (<tt>z</tt>, <tt>sin_z</tt>) and (<tt>z</tt>, <tt>cos_z</tt>) collections as separate dashed lines in whatever (different) colors you like. Make sure to add a legend!
# </div>
# + nbgrader={"grade": false, "grade_id": "cell-926ed397b0cfd7b4", "locked": false, "schema_version": 3, "solution": true, "task": false} tags=["hide-cell"]
""" Implement your ToDo below. """
import math
z = [zi / 10 for zi in list(range(100))]
sin_z = [math.sin(zi) for zi in z]
cos_z = [math.cos(zi) for zi in z]
### BEGIN SOLUTION
plt.plot(z, sin_z)
plt.plot(z, cos_z)
plt.legend(["sin(z)", "cos(z)"])
### END SOLUTION
# Do not remove the code below and implement your code *above* this snippet
ax2check = plt.gca()
plt.show()
# + nbgrader={"grade": true, "grade_id": "cell-84e7cb9e16026c9f", "locked": true, "points": 0, "schema_version": 3, "solution": false, "task": false}
""" Tests the ToDo above. """
lines = ax2check.get_lines()
if len(lines) != 2:
raise ValueError(f"I expected 2 lines but found {len(lines)}!")
leg = ax2check.get_legend()
if leg is None:
raise ValueError("You didn't include a legend!")
n_leg = len(leg.get_lines())
if n_leg != 2:
raise ValueError("I expected two things in the legend but found {n_leg}!")
print("Good job!")
# -
# There are several other things that you can add to or tweak in your plot. For example, you can add a title with `plt.title` or you can change the default ticks and tick labels using `plt.xticks` (for the x-axis ticks/tick labels) and `plt.yticks` (for the y-axis ticks/tick labels). An example:
plt.title("Plot with modified x-axis ticks and tick labels!", fontsize=14)
plt.plot(x, y)
plt.xticks([0, 2, 4, 6], ['0th', '2nd', '4th', '6th'])
plt.show()
# And you can control the range of the axes by the functions `plt.xlim` and `plt.ylim`:
plt.plot(x, y)
plt.xlim(-5, 12)
plt.ylim(-5, 12)
plt.show()
# ### Different plot functions
# Of course, `plt.plot` is not the only plotting functions! There are many different plotting functions in Matplotlib, including scatterplots:
plt.title("A scatterplot!", fontsize=20)
# Note that this is equivalent to plt.plot(x, y, 'o') !
plt.scatter(x, y)
plt.show()
# ... and bar graphs:
# First argument determines the location of the bars on the x-axis
# and the second argument determines the height of the bars
plt.bar(x, x)
plt.show()
# ... and histograms:
# +
# Let's generate some random data
import random
random_unif = [random.uniform(0, 1) for _ in range(100)]
plt.title("A histogram!", fontsize=20)
plt.hist(random_unif)
plt.xlabel("Value", fontsize=15)
plt.ylabel("Frequency", fontsize=15)
plt.show()
# -
# <div class='alert alert-warning'>
# <b>ToDo</b>: Below, we again simulate some random data, but this time not from a uniform distribution, but from a normal distribution (with mean 0.5 and a standard deviation of 0.15). Plot both the uniform data (<tt>random_unif</tt>) and the normal data (<tt>random_norm</tt>) in the same plot, but with different colors (doesn't matter which). Make sure to use 10 bins for each histogram and make sure they are slightly transparent by setting the "alpha" level to 0.5 in both. Also, make sure the ticks and xtick labels are spaced 0.1 apart (i.e., a tick and label at 0, at 0.1, ..., until 1.0). And add a legend! To find out how to set the number of bins and the alpha level, check out the <a href="https://matplotlib.org/3.1.1/api/_as_gen/matplotlib.pyplot.hist.html">documentation</a>!
# </div>
# + nbgrader={"grade": false, "grade_id": "cell-01d2dcba63535062", "locked": false, "schema_version": 3, "solution": true, "task": false} tags=["hide-cell"]
""" Implement the ToDo here! """
random_norm = [random.normalvariate(0.5, 0.15) for _ in range(100)]
### BEGIN SOLUTION
plt.hist(random_unif, alpha=0.5, bins=10)
plt.hist(random_norm, alpha=0.5, bins=10)
plt.legend(['uniform', 'normal'])
plt.xlabel("Value", fontsize=15)
plt.xticks([0, 0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0],
[0, 0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0])
plt.ylabel("Frequency", fontsize=15)
### END SOLUTION
# Do not remove the code below
ax2check = plt.gca()
plt.show()
# + nbgrader={"grade": true, "grade_id": "cell-545d8a139c65258a", "locked": true, "points": 0, "schema_version": 3, "solution": false, "task": false}
""" Tests the ToDo above. """
import matplotlib
rect = [ch for ch in ax2check.get_children() if ch.__class__ == matplotlib.patches.Rectangle]
if len(rect) != 21:
raise ValueError(f"I expected 20 bins in total (10 per histogram), but found {len(rect) - 1}!")
if rect[0]._alpha != 0.5:
raise ValueError(f"The alpha level is not 0.5 (but {rect[0]._alpha})!")
leg = ax2check.get_legend()
if leg is None:
raise ValueError("I couldn't find a legend!")
if len(ax2check.get_xticks()) != 11:
raise ValueError("There should be 11 x-axis ticks and labels!")
if not all(ax2check.get_xticks() == [i / 10 for i in list(range(11))]):
raise ValueError("The x-axis ticks and tick labels are not yet correct ...")
print("Awesome!")
# -
# ## The object-oriented interface
# The state-based plotting approach is easy to learn and pretty straightforward, but when you start creating more complex visualizations, you'll notice that the alternative "object-oriented" approach becomes easier to use. In this section, we will explain this approach by recreating some of the previous plots from the state-based section. We will also discuss some more advanced plotting techniques, such as creating subplots.
#
# Now, within the object-oriented approach, we can explain some of the more technical (but important!) concepts. One of those is that each Matplotlib plot consists of a `Figure` object and one or more `Axes` objects. Essentially, the `Figure` object represents the entire canvas that defines the, well, figure. The `Axes` object(s) contains the actual visualizations that you want to include in the `Figure` (see figure below). Importantly, there may be one *or* multiple `Axes` object within a given `Figure` (e.g., two line plots next to each other).
#
# 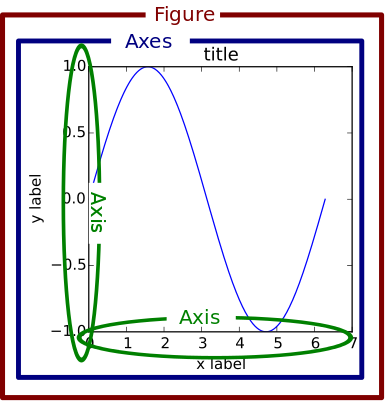
#
# *Figure from <NAME>, from [https://realpython.com/python-matplotlib-guide/](https://realpython.com/python-matplotlib-guide/)*
# <div class='alert alert-danger'>
# <b>Warning</b>: Note that an <tt>Axes</tt> object is something different than the x-axis and y-axis ("axes") of a plot!
# </div>
# Importantly, a `Figure` object by itself doesn't do anything. It just defines the canvas to drawn on, so to speak. `Figure` objects can be initialized using `plt.figure`, which takes several (optional) arguments like `figsize` (width and height in inches) and `dpi` ("dots per inch", i.e., resolution). Let's take a look:
fig = plt.figure(figsize=(8, 4))
plt.show()
# As you can see, nothing happens. We also need an `Axes` object! We can create this using `plt.axes`. Note that, even in the object-oriented appoach, we need the function `plt.show` to render the figure.
fig = plt.figure(figsize=(8, 4))
ax = plt.axes()
plt.show()
# Instead of creating the `Figure` and `Axes` objects separately, we highly recommend using the function `plt.subplots` to create them both at the same time. Like the name suggests, this function also allows you to create multiple subplots (across different `Axes`), which we'll discuss later. For now, we'll just use it to create a `Figure` and `Axes` object at once. Note that all arguments for creating `Figures` using `plt.figure` also work for `plt.subplots`. For example, you can give `plt.subplots` the arguments `figsize=(8, 4)` and `dpi=200` (these must be specified with keywords, however)! Check out the [full documentation] of `plt.subplots` to get an idea about the different arguments it accepts.
#
# Anyway, let's take a look:
fig, ax = plt.subplots(figsize=(8, 4))
plt.show()
# Alright, great, but it's still an empty canvas! Now, we could of course plot some data using the state-based interface (e.g., `plt.plot(x, y)`). Here, however, we will use the object-oriented approach. The only difference between these two approaches is that plotting in the object-oriented approach is done through the *methods* of the `Ax` object instead of the functions from the `pyplot` module. An example:
fig, ax = plt.subplots(figsize=(8, 4))
ax.plot(x, y) # here, `plot` is a method, not a function!
plt.show()
# Basically all functions from the state-based interface are available as methods in the object-oriented approach. For example, to create a legend, run `ax.legend` (instead of `plt.legend`):
fig, ax = plt.subplots(figsize=(8, 4))
ax.plot(x, y)
ax.plot(x, y_sq)
ax.legend(['y', 'y squared'])
plt.show()
# Some `pyplot` functions (like `plt.xlabel`), however, are prefixed with `set_` in the object-oriented interface (e.g., `ax.set_xlabel`):
fig, ax = plt.subplots(figsize=(8, 4))
ax.plot(x, y)
ax.set_xlabel('x', fontsize=15)
ax.set_ylabel('y', fontsize=15)
ax.set_title("Some plot", fontsize=20)
plt.show()
# Okay, time for an exercise!
# <div class='alert alert-warning'>
# <b>ToDo</b>: Below, using a so-called "random walk", we create some random numbers representing two time series. Let's pretend that this data represents the stock price of two companies (Tesla and Shell) across 100 consecutive days. Create a <tt>Figure</tt> of 12 (width) by 4 (height) inches and a single <tt>Axes</tt> onto which you plot this time series data (as lines). Label the axes appropriately and set the range of the x-axis from 1 to 100.
# </div>
# + nbgrader={"grade": false, "grade_id": "cell-26d6937fe9f100ec", "locked": false, "schema_version": 3, "solution": true, "task": false} tags=["hide-cell"]
""" Implement the ToDo here. """
steps1 = [random.uniform(-1, 1) for _ in range(100)]
steps2 = [random.uniform(-1, 1) for _ in range(100)]
tesla = [40 + sum(steps1[:i]) for i in range(100)]
shell = [35 + sum(steps2[:i]) for i in range(100)]
days = list(range(1, 101))
### BEGIN SOLUTION
fig, ax = plt.subplots(figsize=(15, 4))
ax.plot(days, tesla, '-')
ax.plot(days, shell, '-')
ax.set_xlim(1, 100)
ax.set_xlabel('Time (days)', fontsize=15)
ax.set_ylabel('Price', fontsize=15)
ax.legend(['Tesla', 'Shell'])
### END SOLUTION
# Do not remove the code below
ax2check = plt.gca()
plt.show()
# + nbgrader={"grade": true, "grade_id": "cell-df810c6e1ee843cc", "locked": true, "points": 0, "schema_version": 3, "solution": false, "task": false}
""" Tests the above ToDo. """
lines = ax2check.get_lines()
if len(lines) != 2:
raise ValueError(f"I expected two lines, but I saw {len(lines)}!")
if ax2check.get_legend() is None:
raise ValueError("There is no legend!")
if not ax2check.get_xlabel():
raise ValueError("There is no label for the x-axis!")
if not ax2check.get_ylabel():
raise ValueError("There is no label for the y-axis!")
print("YES! Well done!")
# -
# Okay, one last thing we want to show you is how to save figures to disk! In the object-oriented interface, you can save figures using the `Figure` method `savefig` (check out its [documentation](https://matplotlib.org/api/_as_gen/matplotlib.pyplot.savefig.html)). The only mandatory argument is a filename, including an extension. The extension determines as which file type the figure is saved. If you want to save a figure as a PNG file, you can for example do the following:
fig, ax = plt.subplots(figsize=(5, 5))
ax.plot(x, y)
plt.show()
fig.savefig('my_awesome_figure.png')
# This actually created a new file, "my_awesome_figure.png", in our current directory. We can double-check this using the command `!ls`, which is some Jupyter magic that allows us to use a code cell as a terminal temporarily:
# !ls
# ## Subplots (optional)
# In this optional section, we will discuss figures with "subplots", i.e., figures with more than one `Axes` object! The easiest way to do this is to use the `plt.subplots` function. This function accepts the arguments `ncols` and `nrows` to create a figure with multiple `Axes` next to each other (`ncols` > 1) or below each other (`nrows` > 1). For example, suppose that I want to create a figure with three plots next to each other:
# Note that I use the variable name "axes" here instead of "ax" like before
# This is not necessary, but I find it helpful because it tells me this variable contains
# more than one axis
fig, axes = plt.subplots(ncols=3, figsize=(15, 4))
plt.show()
# The variable `axes` is slightly different from what we've seen before. Let's check out its type:
type(axes)
# When you create a figure with more than one `Axes` object, the function `plt.subplots` returns a so-called *numpy* ndarray with `Axes` ("ndarray" stands for *N*-dimensional array). Numpy arrays are data structures that we discuss at length in the last (optional) notebook of this week. For now, you can interpret numpy arrays as (in this case) one or two-dimensional lists. To access the individual `Axes` objects from the numpy array, we can index them as if they are lists. For example:
first_ax = axes[0]
# Now, let's plot some stuff in our different `Axes` objects.
# +
fig, axes = plt.subplots(ncols=3, figsize=(15, 4))
axes[0].plot(x, y)
axes[1].plot(x, [yi ** 2 for yi in y])
axes[2].plot(x, [yi ** 3 for yi in y])
names = ['y', 'y squared', 'y cubed']
for i, name in enumerate(names):
axes[i].set_title(name, fontsize=20)
plt.show()
# -
# <div class='alert alert-warning'>
# <b>ToDo</b>: As you probably have noticed by now, if you don't explicitly give Matplotlib a range for the axes (using <tt>xlim</tt> and <tt>ylim</tt>), it will chose a suitable range itself, which results in separate ranges for the subplots in the above figure. To force the same range across subplots, set the arguments <tt>sharex</tt> and/or <tt>sharey</tt> to <tt>True</tt> in the <tt>plt.subplots</tt> call. Do this in the previous code cell to force the same range across the y-axes!
# </div>
# We can, of course, create figures with multiple columns *and* multiple rows. For example, to create a 2 by 2 grid of `Axes`, we can do the following:
fig, axes = plt.subplots(ncols=2, nrows=2, figsize=(5, 5))
plt.show()
# <div class='alert alert-success'>
# <b>Tip</b>: Sometimes, like in the figure above, subplots may overlap slightly, especially in small figures. Matplotlib has a neat function to fix this: <tt>plt.tight_layout</tt>. Try adding it to the code cell above (after the <tt>plt.subplots</tt> line but before the <tt>plt.show()</tt> line).
# </div>
# Currently, the `axes` variable is a two-dimensional numpy array (because it has both multiple columns and multiple rows). We can double-check this by checking out the `shape` attribute from the numpy array:
axes.shape
# Now, to access the individual `Axes` objects from this numpy array, we need two indices: one to indicate the row and one to indicate the column. For example, to get the upper left `Axes` object (i.e., first row, first column), we do:
upper_left_ax = axes[0, 0]
# To get the upper right `Axes` object (i.e., first row, second column), we do:
upper_right_ax = axes[0, 1]
# <div class='alert alert-warning'>
# <b>ToDo</b>: Extract the lower right <tt>Axes</tt> object and store it in a variable named <tt>lower_right_ax</tt>
# </div>
# + nbgrader={"grade": false, "grade_id": "cell-bc85850323e5ea1d", "locked": false, "schema_version": 3, "solution": true, "task": false} tags=["hide-cell"]
""" Implement the ToDo here. """
### BEGIN SOLUTION
lower_right_ax = axes[1, 1]
### END SOLUTION
# + nbgrader={"grade": true, "grade_id": "cell-76c67b58dfe6f4e9", "locked": true, "points": 0, "schema_version": 3, "solution": false, "task": false}
""" Tests the above ToDo. """
if lower_right_ax != axes.flatten()[-1]:
raise ValueError("That is not the correct Axes object ...")
print("Well done!")
# -
# Alright, there is not much more to subplots that we explained here! Let's finish with a difficult exercise for those that want a challenge. From trigonometry, you may remember we can how to create a sine wave with a particular amplitude and frequency. Below, we included a function, `create_sine_wave`, which takes in a list of timepoints, a desired frequecy, and a desired amplitude:
def create_sine_wave(timepoints, frequency=1, amplitude=1):
""" Creates a sine wave with a given frequency and amplitude for a given set of timepoints.
Parameters
----------
timepoints : list
A list with timepoints (assumed to be in seconds)
frequency : int/float
Desired frequency (in Hz.)
amplitude : int/float
Desired amplitude (arbitrary units)
Returns
-------
sine : list
A list with floats representing the sine wave
"""
sine = [amplitude * math.sin(2 * math.pi * frequency * t) for t in timepoints]
return sine
# Given some timepoints, we can plot its corresponding sine wave:
# +
timepoints = [i / 100 for i in range(500)]
sine = create_sine_wave(timepoints)
fig, ax = plt.subplots(figsize=(8, 2))
ax.plot(timepoints, sine)
ax.set_xlabel("Time")
ax.set_ylabel("sin(x)")
ax.set_xlim(0, max(timepoints))
plt.show()
# -
# We already created a quite complicated figure with 9 subplots (3 rows, 3 columns), which shows a sine wave with increasing frequencies (1, 3, 5) across columns and increasing amplitudes across rows (1, 2, 4). We'll show this figure below:
# 
# <div class='alert alert-warning'>
# <b>ToDo</b>: Try to recreate the figure above with your own code! Use a <tt>figsize</tt> of (10, 10). Good luck! (No test cell)
# </div>
# + nbgrader={"grade": false, "grade_id": "cell-30bc84ade86a2949", "locked": false, "schema_version": 3, "solution": true, "task": false} tags=["hide-cell"]
""" Implement your ToDo here. """
### BEGIN SOLUTION
fig, axes = plt.subplots(ncols=3, nrows=3, figsize=(10, 10), sharex=True, sharey=True)
amps = [1, 2, 4]
freqs = [1, 3, 5]
for i in range(len(amps)):
for ii in range(len(freqs)):
sine = create_sine_wave(timepoints, frequency=freqs[ii], amplitude=amps[i])
axes[i, ii].plot(timepoints, sine)
axes[i, ii].set_title(f"Freq = {freqs[ii]}, amp = {amps[i]}")
axes[i, ii].set_xlim(0, max(timepoints))
if ii == 0:
axes[i, ii].set_ylabel("Activity", fontsize=12)
if i == 2:
axes[i, ii].set_xlabel("Time", fontsize=12)
fig.tight_layout()
plt.show()
fig.savefig('solution_sine_wave_plot.png')
### END SOLUTION
| intropy/solutions/week_1/2_matplotlib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python3 (mle-toolbox)
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# # `mle-logging`: A Lightweight Logger for ML Experiments 📖
# ### Author: [@RobertTLange](https://twitter.com/RobertTLange) [Last Update: January 2022] [](https://colab.research.google.com/github/mle-infrastructure/mle-logging/blob/main/examples/getting_started.ipynb)
#
# There are few things that bring me more joy, than automating and refactoring code, which I use on a daily basis. It feels empowering (when done right) and can lead to some serious time savings. The motto: 'Let's get rid of boilerplate'. One key ingredient to my daily workflow is the logging of neural network training learning trajectories and their diagnostics (predictions, checkpoints, etc.). There are many ways to do this: At the beginning of a project you may only save lists of training/validation/test losses to `.csv` files. After an initial prototype you may switch to a tensorboard setup. Maybe with a time string in front of the filename. But this can become annoying as one wants to compare runs across hyperparameters configurations and aggregate statistics across random seeds. This leads to a fundamental question: **What makes a good logger for Machine Learning Experiments (MLE)?** After looking at some of my previous projects, I drafted a list of desired properties:
#
# 1. **Generality**: The logger should support different types of experiments (multi-seed/multi-configuration) and provide the functionality to store/retrieve their key diagnostics. This includes time-series statistics such as losses or predictive accuracy, network checkpoints of various flavors, generated figures as well as any other objects one might want to save over the course of training.
# 2. **Reproducibility**: The logger should provide all the necessary information in order to reproduce the statistics stored within it. This includes the hyperparameter configuration and random seed of the trained pipeline.
# 3. **Integratability**: Given that we may want to search over hyperparameters and rerun experiments over multiple random seeds, the logger has to be able to easily combine these different sub-logs. Furthermore, fetching an individual run should not be challenging.
# 4. **Usability**: The API has to pleasant and intuitive to use. The log aggregation should require a minimal amount of information regarding the file locations. Plotting individual results should work seamlessly. Finally, it has to be easy to continue a previously interrupted experiment.
#
# Now you may say: "Rob, [Weights&Biases](https://wandb.ai/) provide an awesome service that you may be interested in." And that is certainly true - W&B is awesome. But I am a big fan of simple file systems, which are easily accessible so that loading of results and model checkpoints for post-processing are smooth. Finally, I don't always want to carry around login keys on remote VMs. (Note: I also like to be in low-level control). Based on these considerations and some iterations I came up with the following simple `MLELogger` design:
#
# 
#
# Let's now walk through the individual logging steps, how to visualize & reload the log and how to aggregate multiple logs across random seeds and configurations:
# +
# %load_ext autoreload
# %autoreload 2
# %config InlineBackend.figure_format = 'retina'
# #!pip install -q mle-logging
try:
from mle_logging import MLELogger
except:
# !pip install -q mle-logging
from mle_logging import MLELogger
# -
# # Storing Basic Logging Results (Stats, Checkpoints, Plots)
# We start by creating an instance of the `MLELogger`. The logger takes a set of minimal inputs: `time_to_track`, `what_to_track`. They are both lists of strings which provide the time and statistic variable names, we want to log over time. There are a couple more optional basic ingredients:
#
# | Argument | Type | Description |
# | -------------------------- | -------------------------- | -------------------------------------------------------------- |
# | **`experiment_dir`** | str | Base directory in which you want to store all your results in. |
# | **`config_fname`** | str | Filename of `.json` configuration (to be copied into the `experiment_dir`). |
# | **`config_dict`** | dict | Dictionary of experiment configuration to store in yaml file.
# | **`seed_id`** | int | Random seed of experiment run. |
# | **`model_type`** | str | Specify model type (`jax`, `torch`, `tensorflow`, `sklearn`, `numpy`) to save.|
# | **`use_tboard`** | bool | Boolean indicating whether to log statistics also to TensorBoard. |
# | **`overwrite`** | bool | Whether to overwrite/replace a previously stored log. |
# | **``verbose``** | bool | Whether to print out the most recent updates and logger setup. |
# | **`time_to_print`** | List[str] | Subset of time variables to print out onto the console.|
# | **`what_to_print`** | List[str] | Subset of stats variables to print out onto the console.|
# | **`print_every_k_updates`** | int | How often to print the updated statistics. |
# Instantiate logging to experiment_dir
log = MLELogger(experiment_dir="experiment_dir/",
config_dict={"train_config": {"lrate": 0.01}},
use_tboard=False,
model_type='torch',
print_every_k_updates=1,
verbose=True)
# We can then simply log some a time-series "tick"/timestamp by providing the key, value pairs to `log.update()`. The log will print the last provided statistics using `rich` formatting:
# +
# Save some time series statistics
time_tic = {'num_steps': 10, 'num_epochs': 1}
stats_tic = {'train_loss': 0.1234, 'test_loss': 0.1235}
# Update the log with collected data & save it to .hdf5
log.update(time_tic, stats_tic)
log.save()
# -
# Furthermore, we can save the most current checkpoint via `log.save_model()`. Here is an example for how to store a torch model checkpoint:
# +
# Save a model (torch, sklearn, jax, numpy)
import torch.nn as nn
class DummyModel(nn.Module):
def __init__(self):
super(DummyModel, self).__init__()
self.fc1 = nn.Linear(28*28, 300)
self.fc2 = nn.Linear(300, 100)
self.fc3 = nn.Linear(100, 10)
def forward(self, x):
x = self.fc1(x)
x = self.fc2(x)
x = self.fc3(x)
return x
model = DummyModel()
log.save_model(model)
# -
# If you would like to save a figure that was generated during your training loop, this can be done via `log.save_plot()`. More general objects can be saved as `.pkl` via `log.save_extra`. The log will keep a counter of how many figures or objects were previously saved. If you do not provide an explicit path to the function calls, this counter will be used to archive the files chronologically.
# +
# Save a matplotlib figure as .png
import numpy as np
import matplotlib.pyplot as plt
fig, ax = plt.subplots()
ax.plot(np.random.normal(0, 1, 20))
log.save_plot(fig)
# You can also explicity give a name to the file
# log.save_plot(fig, "some_figure_path.png")
# You can also save (somewhat) arbitrary objects .pkl
some_dict = {"hi": "there"}
log.save_extra(some_dict)
# You can also explicity give a name to the file
# log.save_extra(some_dict, "some_object_path.pkl")
# -
# And obviously you do not need to go through these steps individually, but can also save statistics, model checkpoint, a plot and extra content all in one go:
# Or do everything in one go
log.update(time_tic, stats_tic, model, fig, some_dict, save=True)
# # Reloading for Post-Processing
#
# If your experiments finished and you would like to perform some post-processing, you can load the results via `load_log(<experiment_dir>)`. The reloaded log is a `dotmap` dictionary, which has three subkeys:
#
# 1. `meta`: The meta-information of the experiment. This includes random seed, configuration path, figure paths, experiment directory, etc..
# 2. `time`: The time-series for the `time` variables.
# 3. `stats`: The time-series for the `stats` variables.
#
# The individual data can be accessed via indexing with `.` - have a look:
# +
from mle_logging import load_log
log = load_log("experiment_dir/")
log.meta
# -
log.stats.keys(), log.stats.test_loss
log.time.keys(), log.time.num_updates
# You can also load in the stored model checkpoints for torch, tensorflow, JAX, sklearn, etc. using `load_model`:
# +
from mle_logging import load_model
torch_ckpt = load_model(log.meta.model_ckpt, log.meta.model_type)
torch_ckpt.keys()
# + [markdown] tags=[]
# # Reloading And Resuming An Interrupted Experiment
#
# It can also happen that an experiment was interrupted. Your Colab runtime may run out or for some reason your VM gets interrupted. In that case you would like to be able to resume your experiment and continue to update the log. This can be simply be achieved by using the option `reload=True` and by reloading the previously stored checkpoint:
# +
# Instantiate logging to experiment_dir
log = MLELogger(experiment_dir="experiment_dir/",
use_tboard=True,
model_type='torch',
verbose=True,
reload=True)
log.stats_log.clock_tracked
# -
log.update(time_tic, stats_tic, model, fig, some_dict, save=True)
log.stats_log.stats_tracked
# # Ad-Hoc Plotting of Results
#
# Let's say you have trained a neural network and are now ready to dive into the nitty gritty details. What do you do first? Take a look at your learning curves:
# +
log = MLELogger(time_to_track=['num_updates', 'num_epochs'],
what_to_track=['train_loss', 'test_loss'],
experiment_dir="post_plot_dir/")
for step in range(20):
log.update({'num_updates': step+1, 'num_epochs': 0},
{'train_loss': np.exp(-0.5*step) + np.random.normal(0, 0.1),
'test_loss': np.exp(-0.3*step) + np.random.normal(0, 0.1)}, save=True)
# Load in the log and do a quick inspection of the training loss
log = load_log("post_plot_dir/")
fig, ax = log.plot('train_loss', 'num_updates')
# -
# # Log Different Random Seeds for Same Configuration
# If you provide a `.json` file path and a seed_id, the log will be created in a sub-directory. Furthermore, the `.json` file will be copied for reproducibility. Multiple simultaneous runs (different seeds) can now log to the same directory. Everything else remains the same.
# Check if code is run in Colab: If so -- download configs from repo
try:
import google.colab
IN_COLAB = True
# !wget https://raw.githubusercontent.com/mle-infrastructure/mle-logging/main/examples/config_1.json
# !wget https://raw.githubusercontent.com/mle-infrastructure/mle-logging/main/examples/config_2.json
except:
IN_COLAB = False
# +
# Instantiate logging to experiment_dir for two random seeds
log_seed_1 = MLELogger(experiment_dir="multi_seed_dir/",
config_fname="config_1.json", # Provide path to .json config
seed_id=1) # Provide seed int identifier
log_seed_2 = MLELogger(experiment_dir="multi_seed_dir/",
config_fname="config_1.json", # Provide path to .json config
seed_id=2) # Provide seed int identifier
# Save some time series statistics
for step in range(20):
log_seed_1.update({'num_updates': step+1, 'num_epochs': 0},
{'train_loss': np.exp(-0.5*step) + np.random.normal(0, 0.1),
'test_loss': np.exp(-0.3*step) + np.random.normal(0, 0.1)}, save=True)
log_seed_2.update({'num_updates': step+1, 'num_epochs': 0},
{'train_loss': np.exp(-0.5*step) + np.random.normal(0, 0.1),
'test_loss': np.exp(-0.3*step) + np.random.normal(0, 0.1)}, save=True)
# -
# We can then use `merge_seed_logs` in order to combine both `.hdf5` log files (for the different seeds) into a single file stored in `merged_path`. The `load_log` function afterwards will load this seed-merged log and the first level of the `dotmap` dictionary will give you an overview of the different random seeds:
# +
import os
from mle_logging import merge_seed_logs, load_log
experiment_dir = f"multi_seed_dir/config_1/"
merged_path = os.path.join(experiment_dir, "logs", "seed_aggregated.hdf5")
# Merge different random seeds into one .hdf5 file
merge_seed_logs(merged_path, experiment_dir)
# -
# Load the merged log - Individual seeds can be accessed via log.seed_1, etc.
log = load_log(experiment_dir)
log.eval_ids, log.seed_1.stats.train_loss, log.plot('train_loss', 'num_updates')
# You can also directly aggregate these different random seeds by setting `aggregate_seeds=True`. This will compute the mean, standard deviation as well as different percentiles over the random seeds.
# Load the merged log and aggregate over random seeds (compute stats)
log = load_log(experiment_dir, aggregate_seeds=True)
log.eval_ids, log.stats.train_loss.keys(), log.plot("train_loss", "num_updates")
log.eval_ids
# ## Log Different Configurations with Different Random Seeds
#
# Next, we can also combine different logs for different hyperparameter configurations and their random seeds. Let's first create two logs for two different configurations:
# +
# Instantiate logging to experiment_dir for two .json configurations and two seeds
log_c1_s1 = MLELogger(time_to_track=['num_updates', 'num_epochs'],
what_to_track=['train_loss', 'test_loss'],
experiment_dir="multi_config_dir/",
config_fname="config_1.json",
seed_id=1)
log_c1_s2 = MLELogger(time_to_track=['num_updates', 'num_epochs'],
what_to_track=['train_loss', 'test_loss'],
experiment_dir="multi_config_dir/",
config_fname="config_1.json",
seed_id=2)
log_c2_s1 = MLELogger(time_to_track=['num_updates', 'num_epochs'],
what_to_track=['train_loss', 'test_loss'],
experiment_dir="multi_config_dir/",
config_fname="config_2.json",
seed_id=1)
log_c2_s2 = MLELogger(time_to_track=['num_updates', 'num_epochs'],
what_to_track=['train_loss', 'test_loss'],
experiment_dir="multi_config_dir/",
config_fname="config_2.json",
seed_id=2)
# Update the logs with collected data & save them to .hdf5
for step in range(20):
log_c1_s1.update({'num_updates': step+1, 'num_epochs': 0},
{'train_loss': np.exp(-0.5*step) + np.random.normal(0, 0.1),
'test_loss': np.exp(-0.3*step) + np.random.normal(0, 0.1)}, save=True)
log_c1_s2.update({'num_updates': step+1, 'num_epochs': 0},
{'train_loss': np.exp(-0.5*step) + np.random.normal(0, 0.1),
'test_loss': np.exp(-0.3*step) + np.random.normal(0, 0.1)}, save=True)
log_c2_s1.update({'num_updates': step+1, 'num_epochs': 0},
{'train_loss': np.exp(-0.5*step) + np.random.normal(0, 0.1),
'test_loss': np.exp(-0.3*step) + np.random.normal(0, 0.1)}, save=True)
log_c2_s2.update({'num_updates': step+1, 'num_epochs': 0},
{'train_loss': np.exp(-0.5*step) + np.random.normal(0, 0.1),
'test_loss': np.exp(-0.3*step) + np.random.normal(0, 0.1)}, save=True)
# -
# We can now first merge different random seeds for both configurations (again via `merge_seed_logs`) and then afterwards, combine the seed-aggregated logs for the two configurations via `merge_config_logs`:
# +
# Merge different random seeds for each config into separate .hdf5 file
merge_seed_logs(f"multi_config_dir/config_1/logs/config_1.hdf5",
f"multi_config_dir/config_1/")
merge_seed_logs(f"multi_config_dir/config_2/logs/config_2.hdf5",
f"multi_config_dir/config_2/")
# Aggregate the different merged configuration .hdf5 files into single meta log
from mle_logging import merge_config_logs
merge_config_logs(experiment_dir="multi_config_dir/",
all_run_ids=["config_1", "config_2"])
# -
# This meta-log can then again be reloaded via `load_log` and specifying its location:
# Afterwards load in the meta log object
meta_log = load_log("multi_config_dir/meta_log.hdf5", aggregate_seeds=True)
meta_log.eval_ids, meta_log.config_1.stats.test_loss.keys(), meta_log.plot(["train_loss", "test_loss"], "num_updates")
# Again `load_log` has the option to `aggregate_seeds` or not:
meta_log = load_log("multi_config_dir/meta_log.hdf5", aggregate_seeds=False)
meta_log.eval_ids, meta_log.config_1.keys()
meta_log.plot(["train_loss", "test_loss"], "num_updates")
# # Logging Every k-th Checkpoint Update
#
# Next up you can choose to not only store the most recent checkpoint, but to also store every k-th one. Simply specify `save_every_k_ckpt` when instantiating the logger and the toolbox will take care of the archiving:
# Instantiate logging to experiment_dir
log = MLELogger(time_to_track=['num_updates', 'num_epochs'],
what_to_track=['train_loss', 'test_loss'],
experiment_dir='every_k_dir/',
model_type='torch',
ckpt_time_to_track='num_updates',
save_every_k_ckpt=2)
# +
time_tic = {'num_updates': 10, 'num_epochs': 1}
log.update(time_tic, stats_tic, model, save=True)
time_tic = {'num_updates': 20, 'num_epochs': 1}
log.update(time_tic, stats_tic, model, save=True)
time_tic = {'num_updates': 30, 'num_epochs': 1}
log.update(time_tic, stats_tic, model, save=True)
time_tic = {'num_updates': 40, 'num_epochs': 1}
log.update(time_tic, stats_tic, model, save=True)
log.model_log.every_k_ckpt_list, log.model_log.every_k_storage_time
# -
# # Logging Top-k Checkpoints Based on Metric
#
# Last but not least we can also choose to keep an archive of the top-k performing networks based on chosen metric. As additional input you have to provide the size of your archive `save_top_k_ckpt`, the metric `top_k_metric_name` and whether to minimize it `top_k_minimize_metric`:
# Instantiate logging to experiment_dir
log = MLELogger(time_to_track=['num_updates', 'num_epochs'],
what_to_track=['train_loss', 'test_loss'],
experiment_dir="top_k_dir/",
model_type='torch',
ckpt_time_to_track='num_updates',
save_top_k_ckpt=2,
top_k_metric_name="test_loss",
top_k_minimize_metric=True)
# +
time_tic = {'num_updates': 10, 'num_epochs': 1}
stats_tic = {'train_loss': 0.1234, 'test_loss': 0.1235}
log.update(time_tic, stats_tic, model, save=True)
time_tic = {'num_updates': 20, 'num_epochs': 1}
stats_tic = {'train_loss': 0.1234, 'test_loss': 0.11}
log.update(time_tic, stats_tic, model, save=True)
time_tic = {'num_updates': 30, 'num_epochs': 1}
stats_tic = {'train_loss': 0.1234, 'test_loss': 0.09}
log.update(time_tic, stats_tic, model, save=True)
time_tic = {'num_updates': 40, 'num_epochs': 1}
stats_tic = {'train_loss': 0.1234, 'test_loss': 0.12}
log.update(time_tic, stats_tic, model, save=True)
log.model_log.top_k_performance, log.model_log.top_k_storage_time
# +
# Reload the previously instantiated logger from the directory
log = MLELogger(time_to_track=['num_updates', 'num_epochs'],
what_to_track=['train_loss', 'test_loss'],
experiment_dir="top_k_dir/",
model_type='torch',
ckpt_time_to_track='num_updates',
save_top_k_ckpt=2,
top_k_metric_name="test_loss",
top_k_minimize_metric=True,
reload=True)
log.model_log.top_k_ckpt_list, log.model_log.top_k_storage_time, log.model_log.top_k_performance
# -
# So this is it. Let me know what you think! If you find a bug or are missing your favourite feature, feel free to contact me [@RobertTLange](https://twitter.com/RobertTLange) or create an issue!
| examples/getting_started.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Loading data into RDD, DataFrame
#
# Most of the tasks in this section will be done in the notebook, some will be done on the cluster, you can find the necessary files and submission scripts here:
#
# ```bash
# # cd 2_LoadingData
# ```
#
# By default, Spark will not override the output folder. For `RDDs`, one can pass a config parameter to alter that `.config("spark.hadoop.validateOutputSpecs", "false")` for `DataFrame`s we will explicitely set the write mode.
# First, initialize spark session and spark context:
import pyspark
try:
sc
except NameError:
spark = pyspark.sql.SparkSession.builder.master("local[*]").appName("BD course").config("spark.hadoop.validateOutputSpecs", "false").getOrCreate()
sc = spark.sparkContext
# ## Load unstructured data
#
# One of the first things we need to learn is how to read the data into Spark RDDs and dataframes. Spark provides a battery-included API for reading structured data in most data formats (CSV, JSON, Parquet) as well as unstructured data (plain text files, server logs, etc).
#
# ### Reading plain text
#
# `textFile` and `wholeTextFiles` are functions to read in plain unstructured text.
#
# 1. `textFile` reads data line by line creating an RDD where each entry corresponds to a line (kind of like readlines() in Python)
# 1. `wholeTextFiles` reads the whole file into a pair RDD: (file path, context of the whole file as string)
#
#
# Following code demonstrate that on an example of the word count.
# +
#from pyspark import SparkContext
import sys
import time
import os
def main1(args):
start = time.time()
#sc = SparkContext(appName="LoadUnstructured")
#By default it assumes file located on hdfs folder,
#but by prefixing "file://" it will search the local file system
#Can specify a folder, can pass list of folders or use wild character
input_rdd = sc.textFile("./data/unstructured/")
counts = input_rdd.flatMap(lambda line: line.split()).map(lambda word: (word, 1)).reduceByKey(lambda a, b: a + b)
print ("\nTaking the 10 most frequent words in the text and corresponding frequencies:")
print (counts.takeOrdered(10, key=lambda x: -x[1]))
counts.map(lambda x: (x[1],x[0])).sortByKey(0).map(lambda x: (x[1],x[0])).repartition(1).saveAsTextFile("./output_loadunstructured1/")
end = time.time()
print ("Elapsed time: ", (end-start))
# -
# Try the record-per-line-input
main1(sys.argv)
def main2(args):
start = time.time()
#Use alternative approach: load the dinitial file into a pair RDD
input_pair_rdd = sc.wholeTextFiles("./data/unstructured/")
counts = input_pair_rdd.flatMap(lambda line: line[1].split()).map(lambda word: (word, 1)).reduceByKey(lambda a, b: a + b)
print ("\nTaking the 10 most frequent words in the text and corresponding frequencies:")
print (counts.takeOrdered(10, key=lambda x: -x[1]))
counts.map(lambda x: (x[1],x[0])).sortByKey(0).map(lambda x: (x[1],x[0])).repartition(1).saveAsTextFile("./output_loadunstructured2/")
end = time.time()
print ("Elapsed time: ", (end-start))
#Use alternative approach: load the initial file into a pair RDD
main2(sys.argv)
# ## Loading CSV
#
# Next, we are going to learn how to load data into structured format like CSV. There is at least two ways to do that:
#
# 1. Read the files line by line with `textFiles()` method, split on delimiter (not recommended). This will produced an RDD which is a data structure optimized for row-oriented analysis and functional primitives like `map` and `filter`
# 1. Read the CSV files using the built in `DataFrameReader` (recommended). This will produce a dataframe, which is a data structure optimized for column-oriented analysis and relational primitives
# +
import csv
import sys
import os
try:
from StringIO import StringIO
except ImportError:
from io import StringIO
#this one is use when you use textFile
def loadRecord(line,header,delimiter):
"""Parse a CSV line"""
input = StringIO(line)
reader = csv.DictReader(input, delimiter=delimiter, fieldnames=header)
return next(reader)
def main_rdd(args):
#sc = SparkContext(appName="LoadCsv")
delimiter = "|"
# Try the record-per-line-input
input = sc.textFile("./data/csv/person_nodes.csv")
header = input.first().split(delimiter)
data = input.filter(lambda x: header[0] not in x).map(lambda x: loadRecord(x,header,delimiter))
data.repartition(1).saveAsTextFile("./output_csv/")
def main_dataframe(args):
delimiter = "|"
#csv into spark dataframe
input_df = spark.read.options(header='true', inferschema='true',delimiter=delimiter).csv('./data/csv/person_nodes.csv')
input_df.write.mode("overwrite").option("header", "true").csv("./output_csv2/")
# -
#Load into a regular RDD using textFile and parsing the CSV file line by line
main_rdd(sys.argv)
#Load into dataframe using the csv reader from Databricks
main_dataframe(sys.argv)
# ### Example: analyzing a diamonds dataset
# In a simialr way as before, we are going to read the CSV file into dataframe.
# Spark DataFrameReader can handle delimiters, escaping, and can optionally skip header line for CSV files.
# Read csv data as DataFrame using spark csv dataframe reader
diamonds = spark.read.options(header='true', inferSchema='true').csv('./data/csv/diamonds.csv')
diamonds.show(10)
diamonds.printSchema()
# Let's try doing some basic queries to understand the dataset better.
diamonds.count()
diamonds.select('color').distinct().show()
# Next, let us try to estimate an average price per carat. As you have noticed, the price column is an integer. This can result in a loss of precision as we do averaging. So first of all we will cast this column to double type:
# +
from pyspark.sql.types import DoubleType
from pyspark.sql.functions import *
# Convert Price column to type DoubleType
diamondsdf = diamonds.withColumn("price", diamonds["price"].cast(DoubleType()))
# -
# We will use "groupby-aggregate function" to calculate the average. This creates a column with default name "avg(price)" which we rename to something easier to type. Finally, we order output by price in descending order:
# +
# Calculate average price per carat value
carat_avgPrice = (diamondsdf
.groupBy("carat")
.avg("price")
.withColumnRenamed("avg(price)", "avgPrice")
.orderBy(desc("avgPrice")))
# View top10 highest average prices and corresponding carat value
carat_avgPrice.show(10)
# -
# ### Analyzing CSV files in Python as RDDs (not recommended approach)
#
# In principle, one can use `RDD` to analyze structured data as well, but it seems to be less concenient, especially if the logic of your analysis can be expressed using SQL-like relational primitives.
#
# We will now convert our diamonds DataFrame into RDD:
# We can convert the DataFrame directly into an RDD
diamonds_rdd = diamonds.rdd
# View first 3 rows of the diamonds RDD
diamonds_rdd.take(3)
# You can now use RDD operations to analyze the data:
# Diamond counts by cuts
countByGroup = diamonds_rdd.map(lambda x: (x.cut, 1)).reduceByKey(lambda x,y: x+y)
print (countByGroup.collect())
# Distinct diamond clarities in dataset
distinctClarity = diamonds_rdd.map(lambda x: x.clarity).distinct()
print (distinctClarity.collect())
# Average price per diamond cut
avgPrice = diamonds_rdd.map(lambda x: (x.cut, float(x.price))).reduceByKey(lambda x,y: (x+y)/2)
print (avgPrice.collect())
# # Exercise: load a CSV file and analyze it
#
# Use what you have learned to load a set of `CSV` datasets. Open **load_csv_exercise.py** and follow the assignment therein.
#
# 1. Actor
# 1. Movie
# 1. Actor playing in movie (relationships)
#
# and find movies where **<NAME>** played in.
#
# Save the answer in the `JSON` format.
# # Loading JSON
#
# The best and probably the only reasonable way to load JSON files is using the Spark DataFrameReader.
# Spark SQL has built in support for reading in JSON files which contain a separate, self-contained JSON object per line.
#
# **Note: Multi-line JSON files are currently not compatible with Spark SQL.**
testJsonData = spark.read.json("./data/json/test.json")
testJsonData.printSchema()
testJsonData.show()
# Spark SQL can infer the schema automatically from your JSON data. To view the schema, use `printSchema`.
# Let's now try doing some basic queries to understand the dataset better.
# Count number of rows in dataset
print (testJsonData.count())
# JSON data can contain nested data structures which can be accessed with a "."
testJsonData.select('dict.key').show()
# We can also perform DataFrame operations such as filtering queries according to some criteria:
testJsonData.filter(testJsonData["int"] > 1).show()
# ### Analyzing JSON files in Python with SQL
# Any DataFrame, including those created with JSON data, can be registered as a Spark SQL table to query with SQL.
# Create a Spark SQL temp table
# Note that temp tables are not global across clusters and will not persist across cluster restarts
testJsonData.registerTempTable("test_json")
# We can run any SQL queries on that table with Spark SQL:
spark.sql("SELECT * FROM test_json").show()
# ### Mini-exercise
#
# Switch to the Adroit cluster work directory, open the file: **load_json.py**
# and follow instructions inline. Submit the jobs to the cluster using **slurm_for_json.cmd** file
| 2_LoadingData/LoadingData.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/MrPr3ntice/da_examples/blob/main/da_example_ot_alignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="8STWCtQgwaM_"
# # Imports
# + id="hDTqMjMzwVsW"
# install Python Optimal Transport
# Reference:
# <NAME> al.
# POT Python Optimal Transport library,
# Journal of Machine Learning Research, 22(78):1−8, 2021.
# Website: https://pythonot.github.io/
# !pip install POT
import ot
from matplotlib import pyplot as plt
import matplotlib
from sklearn.datasets import make_classification
from sklearn.base import clone
from sklearn.svm import SVC
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler, Normalizer
from sklearn.preprocessing import FunctionTransformer as FT
from sklearn.pipeline import Pipeline
from sklearn.dummy import DummyClassifier
import numpy as np
# for better reproducibility
np.random.seed(2022)
# + [markdown] id="ZQIY6FpgwQI9"
# # Helper functions
# + id="xDkNx2e0vvf0"
# define scoring function
def my_scorer(est, X_s, y_s, X_t, y_t):
# randomly split first dataset
X_s_train, X_s_test, y_s_train, y_s_test = train_test_split(
X_s, y_s, test_size=0.50, stratify=y_s
)
est.fit(X_s_train, y_s_train)
print('Train ACC (on source domain): ', est.score(X_s_train, y_s_train))
print('Test ACC (on source domain): ', est.score(X_s_test, y_s_test))
# DA
print('Test ACC (on target domain): ', est.score(X_t, y_t))
print('Base rate ACC:', DummyClassifier(strategy='most_frequent').fit(X_s_train, y_s_train).score(X_t, y_t))
# define plotting function
def plot_datasets(dataset_list, n=1500, ax=None, names=None):
if ax is None:
ax = plt.gca()
markers = ['o', 's', '+', 'v', 'x', '8']
for i, data in enumerate(dataset_list):
x, y = data
labels_unique, labels_for_c = np.unique(y, return_inverse=True)
cmap = plt.cm.rainbow
norm = matplotlib.colors.Normalize(vmin=-1, vmax=len(labels_unique)-1)
if i == 2:
alpha = 1
else:
alpha = 0.4
for j, lab in enumerate(labels_unique):
x_j = x[labels_for_c == j, :]
n_j = min(n, x_j.shape[0])
if names is None:
name_i = str(i)
else:
name_i = names[i]
ax.scatter(x_j[:n_j, 0],
x_j[:n_j, 1],
label='dataset: ' + name_i + ', class: ' + str(lab),
cmap='jet',
marker=markers[np.mod(i, len(markers))],
edgecolor=(0,0,0,1),
color=cmap(norm(lab)),
s=100,
alpha=alpha)
ax.legend()
ax.grid(True)
ax.axis('equal')
def my_split(X, y, prop=[0.5, 0.5]):
n = len(y)
prop = np.asarray(prop)
prop = prop / np.sum(prop)
ns = np.around(prop * n)
ns[-1] += n - sum(ns)
idx = []
for j, i in enumerate(ns):
idx += [j] * int(i)
idx = np.asarray(idx)
np.random.shuffle(idx)
out = []
for i in range(len(prop)):
out.append(X[idx == i])
out.append(y[idx == i])
return out
# defining dataset function
def get_example_dataset(c_classes=2, n_s=500, n_t=100, class_sep=1.0, flip_rot=True):
# generate base dataset
X, y = make_classification(
n_samples = n_s + n_t,
n_features=2,
n_informative=2,
n_redundant=0,
n_repeated=0,
n_classes=c_classes,
n_clusters_per_class=1,
weights=None,
flip_y=0.0,
class_sep=class_sep,
hypercube=True,
shift=0.0,
scale=1.0,
shuffle=False,
random_state=None
)
# split datasets from base dataset
X_s, y_s, X_t, y_t = my_split(X, y, [n_s, n_t])
# (affine) transform 2nd dataset
# random rotation
theta = np.random.rand() * 2 * np.pi
rotMatrix = np.array([[np.cos(theta), -np.sin(theta)],
[np.sin(theta), np.cos(theta)]])
# random noise, shift, flip, shear and scale
rand_flip_vec = np.asarray([np.sign(np.random.normal()), 1])
rand_scale_vec = np.random.normal(1.4, 0.4, 2)
rand_shear_mat = np.eye(2)
rand_shear_mat[0, 1] = np.random.normal(0, 0.5)
rand_shift = np.random.normal(10, 1, 2)
if not flip_rot:
rand_flip_vec = np.asarray([1, 1])
theta = (np.random.rand()-0.5) * 0.2 * 2 * np.pi
rotMatrix = np.array([[np.cos(theta), -np.sin(theta)],
[np.sin(theta), np.cos(theta)]])
print('rot: ', rotMatrix)
print('flip: ', rand_flip_vec)
print('scale: ', rand_scale_vec)
print('shear: ', rand_shear_mat)
print('shift: ', rand_shift)
X_t = np.dot(np.dot((X_t + np.random.normal(0, 0.15, X_t.shape)), rotMatrix) * rand_flip_vec * rand_scale_vec, rand_shear_mat) + rand_shift
# X_t = X_s + + np.random.normal(20, 1, 2)
return X_s, y_s, X_t, y_t
# + [markdown] id="NuwptnbnweY9"
# # Generate datasets
# + colab={"base_uri": "https://localhost:8080/", "height": 809} id="s5LR9X9ewHnd" outputId="237420c5-50ca-4b04-97fc-d1ab613aa1f0"
# new data
X_s, y_s, X_t, y_t = get_example_dataset(c_classes=4, n_s=200, n_t=100, class_sep=2, flip_rot=False)
X_s_train, y_s_train, X_s_test, y_s_test = my_split(X_s, y_s, [0.5, 0.5])
X_t_train, y_t_train, X_t_test, y_t_test = my_split(X_t, y_t, [0.5, 0.5])
# plot
plt.subplots(figsize=(11, 11))
plot_datasets([(X_s, y_s), (X_t_train, y_t_train)], names=['source', 'target'])
plt.title('Source and target domain')
# + [markdown] id="I5Z4qEd751sP"
# # Unsupervised OT
# + colab={"base_uri": "https://localhost:8080/", "height": 855} id="Jw4AjaSzwUgN" outputId="6f0eb378-2052-42a6-df1a-a2947ff08183"
# unsupervised domain adaptation
# Sinkhorn Transport with Group lasso regularization
ot_sinkhorn_un = ot.da.SinkhornLpl1Transport(max_iter=10, max_inner_iter=200, reg_e=0.09, reg_cl=1e0)
scaler_s = StandardScaler().fit(X_s)
scaler_t = StandardScaler().fit(X_t_train)
ot_sinkhorn_un.fit(Xs=scaler_s.transform(X_s), ys=y_s,
Xt=scaler_t.transform(X_t_train))
X_t_test_aligned_sinkhorn_un = scaler_s.inverse_transform(
ot_sinkhorn_un.inverse_transform(Xt=scaler_t.transform(X_t_test)))
# plot
fig, axs = plt.subplots(ncols=2, nrows=1, figsize=(18, 10),
constrained_layout=True)
plot_datasets([(X_s, y_s), (X_t, y_t), (X_t_test_aligned_sinkhorn_un, y_t_test)], ax=axs[0], names=['source', 'target', 't -> s'])
axs[0].set_title('unsupervised DA using OT')
axs[1].imshow(ot_sinkhorn_un.cost_, interpolation='nearest')
axs[1].set_xticks([])
axs[1].set_yticks([])
axs[1].set_title('Cost matrix - unsupervised DA')
# estimate
my_scorer(SVC(), X_s, y_s, X_t_test_aligned_sinkhorn_un, y_t_test)
# + [markdown] id="NbucoJoECwsJ"
# # (Semi-)Supervised OT
# + colab={"base_uri": "https://localhost:8080/", "height": 918} id="Xm8xsyxacFse" outputId="febaf70f-c818-4ced-f052-a8834bdcbb60"
# new data
n_t_train_labeled = 200
n_t_train_unlabeled = 20
n_t_test = 1000
n_s = 500
n_t_train = n_t_train_labeled + n_t_train_unlabeled
n_t = n_t_train + n_t_test
X_s, y_s, X_t, y_t = get_example_dataset(c_classes=4, n_s=n_s, n_t=n_t, class_sep=2, flip_rot=True)
X_t_train, y_t_train, X_t_test, y_t_test = my_split(X_t, y_t, [n_t_train, n_t_test])
y_t_train_orig = y_t_train.copy()
# erase label info from target's labeled train data
idx_labeled = np.asarray([True] * n_t_train_labeled + [False] * n_t_train_unlabeled)
np.random.shuffle(idx_labeled)
y_t_train[idx_labeled] = -1
# plot datasets
plt.subplots(figsize=(13, 13))
plot_datasets([(X_s, y_s), (X_t_train, y_t_train)], names=['source', 'target_train'])
plt.title('Source and target domain')
# + colab={"base_uri": "https://localhost:8080/", "height": 855} id="eY2-fIdyIkxT" outputId="e2bf36dc-4ce4-497f-86c8-dbbb79b1c40f"
# semi-supervised domain adaptation
# Sinkhorn Transport with Group lasso regularization
ot_sinkhorn_semi = ot.da.SinkhornLpl1Transport(max_iter=10, max_inner_iter=200, reg_e=0.1, reg_cl=1)
scaler_s = StandardScaler().fit(X_s)
scaler_t = StandardScaler().fit(X_t_train)
ot_sinkhorn_semi.fit(Xs=scaler_s.transform(X_s), ys=y_s,
Xt=scaler_t.transform(X_t_train), yt=y_t_train)
X_t_test_aligned_sinkhorn_semi = scaler_s.inverse_transform(
ot_sinkhorn_semi.inverse_transform(Xt=scaler_t.transform(X_t_test)))
# plot
fig, axs = plt.subplots(ncols=2, nrows=1, figsize=(18, 10),
constrained_layout=True)
plot_datasets([(X_s, y_s), (X_t_train, y_t_train), (X_t_test_aligned_sinkhorn_semi, y_t_test)], ax=axs[0], names=['source', 'target_train', 't_test -> s'])
axs[0].set_title('unsupervised DA using OT')
axs[1].imshow(ot_sinkhorn_semi.cost_, interpolation='nearest')
axs[1].set_xticks([])
axs[1].set_yticks([])
axs[1].set_title('Cost matrix - unsupervised DA')
# estimate
my_scorer(SVC(), X_s, y_s, X_t_test_aligned_sinkhorn_semi, y_t_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 892} id="MQRoZT0qFqW5" outputId="f3467d74-9545-4875-82af-9e7eba20e53f"
# two-stage semi-supervised domain adaptation
X_t_train_labeled = X_t_train[y_t_train != -1]
y_t_train_labeled = y_t_train[y_t_train != -1]
X_t_train_unlabeled = X_t_train[y_t_train == -1]
y_t_train_unlabeled = y_t_train[y_t_train == -1]
# Sinkhorn Transport with Group lasso regularization
ot_sinkhorn_semi_su = ot.da.SinkhornLpl1Transport(max_iter=10, max_inner_iter=200, reg_e=0.1, reg_cl=1)
ot_sinkhorn_semi_un = ot.da.SinkhornTransport(max_iter=10, reg_e=0.05)
scaler_s = StandardScaler().fit(X_s)
scaler_t = StandardScaler().fit(X_t_train)
ot_sinkhorn_semi_su.fit(Xs=scaler_s.transform(X_s), ys=y_s,
Xt=scaler_t.transform(X_t_train_labeled), yt=y_t_train_labeled)
ot_sinkhorn_semi_un.fit(Xs=scaler_s.transform(X_s),
Xt=ot_sinkhorn_semi_su.inverse_transform(Xt=scaler_t.transform(X_t_train_unlabeled)))
X_t_test_aligned_sinkhorn_semi = scaler_s.inverse_transform(
ot_sinkhorn_semi_su.inverse_transform(
Xt=scaler_t.transform(X_t_test)))
X_t_test_aligned_sinkhorn_semi_two = scaler_s.inverse_transform(
ot_sinkhorn_semi_un.inverse_transform(
Xt=ot_sinkhorn_semi_su.inverse_transform(
Xt=scaler_t.transform(X_t_test))))
# plot
fig, axs = plt.subplots(ncols=2, nrows=1, figsize=(18, 10),
constrained_layout=True)
plot_datasets([(X_s, y_s), (X_t_train, y_t_train), (X_t_test_aligned_sinkhorn_semi, y_t_test)], ax=axs[0], names=['source', 'target_train', 't_test -> s'])
axs[0].set_title('one-stage semi-supervised DA using OT')
#axs[1].imshow(ot_sinkhorn_semi_un.coupling_, interpolation='nearest')
#axs[1].set_xticks([])
#axs[1].set_yticks([])
#axs[1].set_title('Cost matrix - unsupervised DA')
plot_datasets([(X_s, y_s), (X_t_train, y_t_train), (X_t_test_aligned_sinkhorn_semi_two, y_t_test)], ax=axs[1], names=['source', 'target_train', 't_test -> s'])
axs[1].set_title('two-stage semi-supervised DA using OT')
# estimate
my_scorer(SVC(), X_s, y_s, X_t_test_aligned_sinkhorn_semi, y_t_test)
| da_example_ot_alignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.9 64-bit (''env'': venv)'
# language: python
# name: python3
# ---
import pandas as pd
import glob
import os
import tensorflow as tf
import seaborn as sns
import matplotlib.pyplot as plt
data = pd.read_csv("/home/m-joaopr/demo/concat/data.csv")
data = pd.concat(map(pd.read_csv, glob.glob("/home/m-joaopr/demo/data/2018*.csv")))
data.head()
data.describe()
data.info()
data.shape
data.isnull().any().sum()
data.fillna(0, inplace=True)
data.isnull().any().sum()
data.shape
# +
#data.to_csv('/home/m-joaopr/demo/concat/data.csv',index=False)
# -
inputs = data.columns[data.columns.str.contains("input")]
inputs
# +
data.drop(columns=["id"])
ax = sns.heatmap(data, cmap='RdYlGn_r',
robust=True,
fmt='.2f',
annot=True,
linewidths=.5,
annot_kws={'size':11},
cbar_kws={'shrink':.8,
'label':'Precipitation(mm)'})
ax.set_yticklabels(ax.get_yticklabels(), rotation=0, fontsize=10)
ax.set_xticklabels(ax.get_xticklabels(), rotation=0, fontsize=10)
plt.title('Average Precipitations', fontdict={'fontsize':18}, pad=14);
# -
| series.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # WeatherPy
# ----
#
# #### Note
# * Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
# +
# Code to make charts and plots interactive, commented out.
# #%matplotlib notebook
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import requests
import time
import os
from scipy.stats import linregress
# Incorporated citipy to determine city based on latitude and longitude
from citipy import citipy
# For displaying of API data more clearly
from pprint import pprint
# Import API key
from api_keys import weather_api_key
import json
# Output File (CSV)
output_data_file = "output_data/cities.csv"
# Range of latitudes and longitudes
lat_range = (-90, 90)
lng_range = (-180, 180)
# -
# ## Generate Cities List
# +
# List for holding lat_lngs and cities
lat_lngs = []
cities = []
# Create a set of random lat and lng combinations
lats = np.random.uniform(lat_range[0], lat_range[1], size=2000)
lngs = np.random.uniform(lng_range[0], lng_range[1], size=2000)
lat_lngs = zip(lats, lngs)
# Identify nearest city for each lat, lng combination
for lat_lng in lat_lngs:
city = citipy.nearest_city(lat_lng[0], lat_lng[1]).city_name
# Capitalizing the first letter of each city word
city = city.title()
# If the city is unique, then add it to a our cities list
if city not in cities:
cities.append(city)
# Print the city count to confirm sufficient count
randomCities = len(cities)
randomCities
# +
# Setting up the Dataframe to hold 'cities' values
City_DF = pd.DataFrame(cities)
# Copying original list, in case its needed to be referenced later
CityWeather_DF = City_DF
# Changing first column '0' to 'City' which represents the cities generated
CityWeather_DF = CityWeather_DF.rename(columns={0:"City"})
# Exporting initial 'cities data
CityWeather_DF.to_csv('../output_data/OriginalRandomCities.csv')
# Order of columns in new dataframe that will hold location/weather data will be:
# City Country Lat Lng Max Temp Humidity Cloudiness Wind Speed Date
CityWeather_DF.head(20)
# -
# ### Perform API Calls
# * Perform a weather check on each city using a series of successive API calls.
# * Include a print log of each city as it'sbeing processed (with the city number and city name).
#
# +
# SUPPLEMENTAL INFO AND API TESTS
# Creating dynamic URL and checking to see if it works
# Expected output according to API documentation:
# api.openweathermap.org/data/2.5/weather?q={City}&units={unit}&appid={api_key}
# EX) api.openweathermap.org/data/2.5/weather?q=London&units=imperial&appid=APIKEY
# 'units' is 'imperial' for Fahrenheit temp, api key is to be inserted in file "api_keys.py"
# Variables
#baseURL = "http://api.openweathermap.org/data/2.5/find?q="
#cityName = CityWeather_DF["City"][0]
#cityURL = cityName.replace(" ", "+")
#unitSys = "&units=imperial&appid="
#queryURL = (f'{baseURL}{cityURL}{unitSys}{weather_api_key}')
#print(queryURL)
#print()
#JsonWeatherResponse = requests.get(queryURL).json()
#pprint(JsonWeatherResponse)
# + tags=[]
# Start of user facing "data scan"
print()
print(" ----------------------------- ")
print(" BEGINNING DATA RETRIEVAL ")
print(" ----------------------------- ")
print()
# Index variable to iterate through generated citipy list
nextCity = 0
# Counter to keep track of every 50 succussful city data captures to increase setCounter by 1
recordCounter = 1
# Counter to keep track of how many sets of 50 successful city data captures
setCounter = 1
# Counter to keep track of all times a data API attempt was made (successful or not)
allCityCounter = 1
# For loop to go through however many random cities were generated:
for nextCity in range(randomCities):
# Creating API url dynamically for every city json call/response in city list
baseURL = "http://api.openweathermap.org/data/2.5/find?q="
# cityName may have spaces""", so removing them for the url link and replacing them with a "+"
cityName = CityWeather_DF["City"][nextCity]
# using new variable for url to keep the original cityName format for later use in status message
cityURL = cityName.replace(" ", "+")
unitSys = "&units=imperial&appid="
queryURL = (f'{baseURL}{cityURL}{unitSys}{weather_api_key}')
# Copying created API url for current city in another variable to be used in error message
ReviewURL = queryURL
JsonWeatherResponse = requests.get(queryURL).json()
# **Reseting Humidity > 100% checker to defauly False
humidityCheck = False
# Code will try to execute but be cognizant for Exception errors, as well as coded error API checks
try:
# recordCounter will reset itself and add 1 to the "Set" display every 50 successful city entries
if recordCounter == 51:
recordCounter = 0
setCounter += 1
# if API response returns an incomplete dataset (count=0)
# or a dataset with too much data that will errors out (ie count = 5)
# or webpage returns a cod code/Response other than 200 (ie 404, etc).
# this Code will proactively not enter data in and return an error
# message/update to user with api http. link and continue to process through loop and counts.
# These entries/rows will be deleted later
if JsonWeatherResponse["count"] > 3 or \
JsonWeatherResponse["count"] == '0' or \
JsonWeatherResponse["cod"] != "200":
print()
print(f'Data for {cityName} is corrupted. Skipping. Can review URL below...')
print(ReviewURL)
print()
nextCity += 1
allCityCounter += 1
humidityCheck = True
# Create and populate 9 columns with API data and a humidity check for current city (nextCity/index)
else:
CityWeather_DF.loc[nextCity, "Country"] = JsonWeatherResponse['list'][0]["sys"]["country"]
CityWeather_DF.loc[nextCity, "Lat"] = JsonWeatherResponse['list'][0]["coord"]["lat"]
CityWeather_DF.loc[nextCity, "Lng"] = JsonWeatherResponse['list'][0]["coord"]["lon"]
CityWeather_DF.loc[nextCity, "Max Temp"] = JsonWeatherResponse['list'][0]["main"]["temp_max"]
CityWeather_DF.loc[nextCity, "Humidity"] = JsonWeatherResponse['list'][0]["main"]["humidity"]
# ** Will perform a check on the Humidty value here and "mark it down"
CityWeather_DF.loc[nextCity, "Cloudiness"] = JsonWeatherResponse['list'][0]["clouds"]["all"]
CityWeather_DF.loc[nextCity, "Wind Speed"] = JsonWeatherResponse['list'][0]["wind"]["speed"]
# Format data for readability
CityWeather_DF.loc[nextCity, "Date"] = time.strftime(' %m/%d/%Y', time.localtime(JsonWeatherResponse['list'][0]["dt"]))
# **Perform a check on if Humidity % > 100 (to mark for deletion later)
# Assigning value to a new column to track
if CityWeather_DF["Humidity"][nextCity] >= 100.00:
humidityCheck = True
CityWeather_DF.loc[nextCity, "Humidity > 100%"] = humidityCheck
else:
humidityCheck = False
CityWeather_DF.loc[nextCity, "Humidity > 100%"] = humidityCheck
# 'True' columned cities will have their data/ROWS deleted later in a new DF
# 'False' columned cities wll retain original data. New column will then be deleted during clean
# Message progress detailing number of successful sets, all attempts vs full list, and the city
print(f"Processing Record.. { recordCounter } of Set { setCounter } ({allCityCounter} / {randomCities}) | {cityName}")
# Iterate counts
nextCity += 1
recordCounter += 1
allCityCounter += 1
# In the event of an indexerror/incomplet API, no data will be marked and loop/count will be iterated
except IndexError:
print()
print(f'No Data for {cityName}. Skipping entry...')
print()
allCityCounter += 1
nextCity += 1
pass
# End of "data scan"
print()
print(" ----------------------------- ")
print(" DATA RETRIEVAL COMPLETE ")
print(" ----------------------------- ")
print()
# My API key will show in the outputs, but it will be deactivated and deleted. To run a test of your own, a new key will have to be registered at: https://home.openweathermap.org/ and saved in this code's directory "api_keys.py" file.
# -
# Display column counts, calculations overview (including max values ie "Humidity"), and dataframe preview
print(CityWeather_DF.count())
print()
print()
print(CityWeather_DF.describe())
print()
print()
CityWeather_DF.head(20)
# ## Inspect the data and remove the cities where the humidity > 100%.
# ----
# Skip this step if there are no cities that have humidity > 100%.
# +
# _______ CityWeather_DF _______
# Clean the data of any blanks, duplicates, or rows with null/NaN values.
CityWeather_DF.replace("", np.nan, inplace=True)
CityWeather_DF.replace("NaN", np.nan, inplace=True)
CityWeather_DF = CityWeather_DF.drop_duplicates()
CityWeather_DF = CityWeather_DF.dropna()
# Check to see if all rows have the same count and dataframe preview.
#print(CityWeather_DF.count())
#print()
#CityWeather_DF.head(20)
# Create new copy of dataframe.
clean_city_dataDF = CityWeather_DF.copy()
# ** See next code snippet regarding new "100% Humidity" DF
# Once the copy is complete, the orignal dataframe will be cleaned of the extra ["Humidity > 100%"] row
# Displaying cleaned "original/spaceless" DF
del CityWeather_DF["Humidity > 100%"]
print(CityWeather_DF.count())
print()
CityWeather_DF.head(20)
# +
# _______ clean_city_dataDF _______
# Make a new DataFrame equal to the city data to drop all humidity outliers by index.
# **This DF copy will be cleaned further of cities with +100% humidty by removing rows of True
clean_city_dataDF = clean_city_dataDF[clean_city_dataDF["Humidity > 100%"] != True]
# And then Humidity column itself and display DF summary
del clean_city_dataDF["Humidity > 100%"]
print(clean_city_dataDF.count())
print()
clean_city_dataDF.head(20)
# -
# ### Convert Raw Data to DataFrame
# * Export the city data into a .csv.
# * Display the DataFrame
# +
# Summary display of CityWeather_DF dataframe and export to .csv
print(CityWeather_DF.count())
print()
print(CityWeather_DF.describe())
print()
CityWeather_DF.to_csv('../output_data/CityWeather.csv')
print(CityWeather_DF.head(20))
print()
# +
# Summary display of clean_city_dataDF dataframe and export to .csv
print(clean_city_dataDF.count())
print()
print(clean_city_dataDF.describe())
print()
clean_city_dataDF.to_csv('../output_data/CleanCityWeatherData.csv')
print(clean_city_dataDF.head(20))
print()
# -
# ## Plotting the Data
# * Use proper labeling of the plots using plot titles (including date of analysis) and axes labels.
# * Save the plotted figures as .pngs.
# ## Latitude vs. Temperature Plot
# +
# Latitude vs. Max Temperature Plot
plt.scatter(clean_city_dataDF["Lat"], clean_city_dataDF["Max Temp"], facecolor="red", edgecolors="yellow",
s=clean_city_dataDF["Max Temp"])
plt.title("City Latitude vs. Max Temperature (°F) - 03/15/21")
plt.grid()
plt.xlim(-60,80)
plt.ylim(0,110)
plt.xlabel("Latitude")
plt.ylabel("Max Temperature (°F)")
plt.savefig("../Images/Lat_vs_MaxTemp.jpg", dpi=200)
plt.show()
print(" Based on our 'Latitude vs. Max Temperature' scatter plot, we can see that as one gets closer to the equator (0) from the South, the temperature arches upwards in a curve to reflect the rise in temperature before trending down as we move away north from the equator's latitude. Interestingly, the highest temperature are not exactly at the equator but rather off approximately 20 degrees north and south of the Equator.")
# -
# ## Latitude vs. Humidity Plot
# +
# Latitude vs. Humidity Plot
plt.scatter(clean_city_dataDF["Lat"], clean_city_dataDF["Humidity"], facecolor="yellow", edgecolors="black",
alpha=1, s=clean_city_dataDF["Humidity"])
plt.title("City Latitude vs. Humidity (%) - 03/15/21")
plt.grid()
plt.xlim(-60,80)
plt.ylim(0,110)
plt.xlabel("Latitude")
plt.ylabel("Humidity (%)")
plt.savefig("../Images/Lat_vs_Humidity.jpg", dpi=200)
plt.show()
print(" There doesn't seem to be any discernable pattern or relation between Humidity and the Latitude. The humidity % is uniformly scattered, and generally above 60%, no matter how close or far away from the Equator you are.")
# -
# ## Latitude vs. Cloudiness Plot
# +
# Latitude vs. Cloudiness Plot
plt.scatter(clean_city_dataDF["Lat"], clean_city_dataDF["Cloudiness"], facecolor="blue", edgecolors="white",
alpha=1, s=clean_city_dataDF["Cloudiness"])
plt.title("City Latitude vs. Cloudiness (%) - 03/15/21")
plt.grid()
plt.xlim(-60,80)
plt.ylim(0,110)
plt.xlabel("Latitude")
plt.ylabel("Cloudiness (%)")
plt.savefig("../Images/Lat_vs_Cloudiness.jpg", dpi=200)
plt.show()
print(" Just as with the observations regarding Humidity vs Latitude, there seems to be no pattern or trend to signify a direct correlation between Cloudiness and Latitude. Many cities seem to share a high percentage/concentration of cloudiness the world over.")
# -
# ## Latitude vs. Wind Speed Plot
# +
# Latitude vs. Wind Speed Plot
plt.scatter(clean_city_dataDF["Lat"], clean_city_dataDF["Wind Speed"], facecolor="green", edgecolors="gray",
alpha=1)
plt.title("City Latitude vs. Wind Speed (mph) - 03/15/21")
plt.grid()
plt.xlim(-60,80)
plt.ylim(0,30)
plt.xlabel("Latitude")
plt.ylabel("Wind Speed (mph)")
plt.savefig("../Images/Lat_vs_WindSpeed.jpg", dpi=200)
plt.show()
print("The only observation I can see is, more generally, the low wind speed across most cities. The wind speed doesn't really go too much past 20mph, on the high end, regardless of latitude and equatorial distance. The concentration of wind speed seem to be around 3-8 mph today. If any, there are higher wind speed toward the extreme North vs extreme South.")
# -
# ## Linear Regression
# Must separate the plots into a Northern Hemisphere (greater than or equal to 0 degrees latitude) and a Southern Hemisphere (less than 0 degrees latitude) dataframe grouping. Referencing 'Humidity>100%' check code: CityWeather_DF.loc[nextCity, "Humidity > 100%"] = CityWeather_DF["Humidity"][nextCity] >= 100.00:
# Northern Hemispheres Cities
NorthernHemi_DF = clean_city_dataDF.loc[clean_city_dataDF["Lat"] >= 0]
NorthernHemi_DF.head(20) # Test to see if returned Latitude values are > 0, Lat should be positive
#clean_city_dataDF.head(20) # Compare results to normal dataframe, if need.
# Southern Hemispheres Cities
SouthernHemi_DF = clean_city_dataDF.loc[clean_city_dataDF["Lat"] <= 0]
SouthernHemi_DF.head(20) # Test to see if returned Latitude values are < 0, Lat should be negative
#clean_city_dataDF.head(20) # Compare results to normal dataframe, if need.
# +
# Linear Regression formula is going to be:
# (slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
# Will need variables for x/y _values which will be the hemisphere dataframes':
# Lat vs. Max Temp / Humidity / Cloudiness / Wind Speed
# Referencing class exercise on Regression for code:
# x_values = _DF["xxx"]
# y_values = _DF["xxx"]
# (slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
# regress_values = x_values * slope + intercept
# line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
# plt.scatter(x_values,y_values)
# plt.plot(x_values,regress_values,"r-")
# plt.annotate(line_eq,(6,10),fontsize=15,color="red")
# plt.xlabel('xxx')
# plt.ylabel('xxx')
# print(f"The r-squared is: {rvalue**2}")
# plt.show()
# Will plug in values/columns and reuse code format to keep charts consistent
# -
# #### Northern Hemisphere - Max Temp vs. Latitude Linear Regression
# +
plt.title(" North Hemisphere - Latitude vs. Max Temp (°F) - 03/15/21")
plt.grid()
x_values = NorthernHemi_DF["Lat"]
y_values = NorthernHemi_DF["Max Temp"]
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('Latitude')
plt.ylabel('Max Temperature (°F)')
print(f"The r-squared is: {rvalue**2}")
plt.savefig("../Images/MaxTemp_vs_Lat_NorthHemi.jpg", dpi=200)
plt.show()
print("There is a negative linear relationship/regression line between the two variables/axis: as the value of one increases, the value of the other decreases. Note: Correlation is not the same as causation.")
# -
# #### Southern Hemisphere - Max Temp vs. Latitude Linear Regression
# +
plt.title(" South Hemisphere - Latitude vs. Max Temp (°F) - 03/15/21")
plt.grid()
x_values = SouthernHemi_DF["Lat"]
y_values = SouthernHemi_DF["Max Temp"]
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('Latitude')
plt.ylabel('Max Temperature (°F)')
print(f"The r-squared is: {rvalue**2}")
plt.savefig("../Images/MaxTemp_vs_Lat_SouthHemi.jpg", dpi=200)
plt.show()
print("There is a positive linear relationship/regression line between the two variables/axis: as the value of one increases, the value of the other also increases. Note: Correlation is not the same as causation.")
# -
# #### Northern Hemisphere - Humidity (%) vs. Latitude Linear Regression
# +
plt.title(" North Hemisphere - Latitude vs. Humidity (%) - 03/15/21")
plt.grid()
x_values = NorthernHemi_DF["Lat"]
y_values = NorthernHemi_DF["Humidity"]
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('Latitude')
plt.ylabel('Humidity (%)')
print(f"The r-squared is: {rvalue**2}")
plt.savefig("../Images/Humidity_vs_Lat_NorthHemi.jpg", dpi=200)
plt.show()
print("There is a positive linear relationship/regression line between the two variables/axis: as the value of one increases, the value of the other also increases. Note: Correlation is not the same as causation.")
# -
# #### Southern Hemisphere - Humidity (%) vs. Latitude Linear Regression
# +
plt.title(" South Hemisphere - Latitude vs. Humidity (%) - 03/15/21")
plt.grid()
x_values = SouthernHemi_DF["Lat"]
y_values = SouthernHemi_DF["Humidity"]
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('Latitude')
plt.ylabel('Humidity (%)')
print(f"The r-squared is: {rvalue**2}")
plt.savefig("../Images/Humidity_vs_Lat_SouthHemi.jpg", dpi=200)
plt.show()
print("There is a positive linear relationship/regression line between the two variables/axis: as the value of one increases, the value of the other also increases. Note: Correlation is not the same as causation.")
# -
# #### Northern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression
# +
plt.title(" North Hemisphere - Latitude vs. Cloudiness (%) - 03/15/21")
plt.grid()
x_values = NorthernHemi_DF["Lat"]
y_values = NorthernHemi_DF["Cloudiness"]
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('Latitude')
plt.ylabel('Cloudiness (%)')
print(f"The r-squared is: {rvalue**2}")
plt.savefig("../Images/Cloudiness_vs_Lat_NorthHemi.jpg", dpi=200)
plt.show()
print("There is a positive linear relationship/regression line between the two variables/axis: as the value of one increases, the value of the other also increases. Note: Correlation is not the same as causation.")
# -
# #### Southern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression
# +
plt.title(" South Hemisphere - Latitude vs. Cloudiness (%) - 03/15/21")
plt.grid()
x_values = SouthernHemi_DF["Lat"]
y_values = SouthernHemi_DF["Cloudiness"]
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('Latitude')
plt.ylabel('Cloudiness (%)')
print(f"The r-squared is: {rvalue**2}")
plt.savefig("../Images/Cloudiness_vs_Lat_SouthHemi.jpg", dpi=200)
plt.show()
print("There is a positive linear relationship/regression line between the two variables/axis: as the value of one increases, the value of the other also increases. Note: Correlation is not the same as causation.")
# -
# #### Northern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression
# +
plt.title(" North Hemisphere - Latitude vs. Wind Speed (mph) - 03/15/21")
plt.grid()
x_values = NorthernHemi_DF["Lat"]
y_values = NorthernHemi_DF["Wind Speed"]
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('Latitude')
plt.ylabel('Wind Speed (mph)')
print(f"The r-squared is: {rvalue**2}")
plt.savefig("../Images/WindSpeed_vs_Lat_NorthHemi.jpg", dpi=200)
plt.show()
print("The linear regression line is flat, and not sloped really, indicating there is no relationship between the two variables. Note: Correlation is not the same as causation.")
# -
# #### Southern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression
# +
plt.title(" South Hemisphere - Latitude vs. Wind Speed (mph) - 03/15/21")
x_values = SouthernHemi_DF["Lat"]
y_values = SouthernHemi_DF["Wind Speed"]
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('Latitude')
plt.ylabel('Wind Speed (mph)')
print(f"The r-squared is: {rvalue**2}")
plt.savefig("../Images/WindSpeed_vs_Lat_SouthHemi.jpg", dpi=200)
plt.show()
print("There is a negative linear relationship/regression line between the two variables/axis: as the value of one increases, the value of the other decreases. Note: Correlation is not the same as causation.")
# -
| WeatherPy/WeatherPy-Ithamar_Francois.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Forecast with Cold Start Items
#
# Consider the situation where a set of related new items got introduced into the catalog, one needs to forecast the future values for those items. In such a situation, little to none demand history is available for those items. This scenario is coined as "cold-start problem."
#
# <img src="../../common/images/amazon_forecast.png">
#
# # Introduction
#
# In this notebook, we walk through the process of generating forecasts for cold start items. Notice that <b>only DeepAR+ supports cold start.</b> At a high level, the flow can be summarized as follows.
#
# 1. We follow the complete process with non-cold start items as in the previous notebooks such as
# * [4. Getting_started_with_DeepAR+.ipynb](../Getting_started_with_DeepAR+/Getting_started_with_DeepAR+.ipynb) or
# * [6. Incorporating_Related_Time_Series_dataset_to_your_Predictor.ipynb](../Incorporating_Related_Time_Series_dataset_to_your_Predictor/Incorporating_Related_Time_Series_dataset_to_your_Predictor.ipynb).
#
# The major addition is that, *the item meta information for both the cold start and non-cold start items are imported in the system*.
#
# 2. Create another dataset with the cold start items, and create forecasts for those items. Here for simplicity, we only use target time series only dataset, but related time series can be incorporated as well.
#
#
# # Table of Contents
#
# * Step 0: [Setting up](#setup)
# * Step 1: [Preparing the Datasets](#prepare)
# * Step 2: [Importing the Data for Non-Cold Start Items](#import)
# * Step 2a: [Creating a Dataset Group](#create)
# * Step 2b: [Creating a Target Dataset](#target)
# * Step 2c: [Creating a Item Meta Information Dataset](#related)
# * Step 2d: [Update the Dataset Group](#update)
# * Step 2e: [Creating a Target Time Series Dataset Import Job](#targetImport)
# * Step 2f: [Creating a Item Meta Information Dataset Import Job](#relatedImport)
# * Step 3: [Create the DeepAR+ Predictor](#algo)
# * Step 4: [Create a Forecast for non-Cold Start Items](#forecast)
#
# The above steps complete the model training with the non-Cold Start items. Now we ready to import the cold start items, and generate their forecats.
#
# * Step 5: [Create a Cold-Start Target Time Series Dataset Import Job](#coldStartImport)
# * Step 6: [Create a Forecast for the cold start items](#createColdStart)
# * Step 7: [Querying the Forecasts](#query)
# * Step 8: [Exporting the Forecasts](#export)
# * Step 9: [Clearning up your Resources](#cleanup)
# # Step 0: First let us setup Amazon Forecast<a class="anchor" id="setup">
#
# This section sets up the permissions and relevant endpoints.
# +
import sys
import os
import boto3
import pandas as pd
import matplotlib.pyplot as plt
# importing forecast notebook utility from notebooks/common directory
sys.path.insert( 0, os.path.abspath("../../common") )
import util
plt.rcParams['figure.figsize'] = (15.0, 5.0)
# -
# Configure the S3 bucket name and region name for this lesson.
#
# - If you don't have an S3 bucket, create it first on S3.
# - Although we have set the region to us-west-2 as a default value below, you can choose any of the regions that the service is available in.
text_widget_bucket = util.create_text_widget( "bucketName", "input your S3 bucket name" )
text_widget_region = util.create_text_widget( "region", "input region name.", default_value="us-west-2" )
# +
bucketName = text_widget_bucket.value
assert bucketName, "bucketName not set."
region = text_widget_region.value
assert region, "region not set."
# -
session = boto3.Session(region_name=region)
forecast = session.client(service_name='forecast')
forecast_query = session.client(service_name='forecastquery')
# Create the role to provide to Amazon Forecast.
role_name = "ForecastNotebookRole-ColdStart"
role_arn = util.get_or_create_iam_role( role_name = role_name )
# # Step 1: Preparing the Datasets<a class="anchor" id="prepare">
#
#
# Here we use a synthetic dataset based on [electricity]() dataset, which consists of the hourly time series for 370 households (with item id 0 to 369).
#
# In this hypothetical senario, our goal is to generate forecasts for 4 new customers with item id 370 to 373.
# +
zipLocalFilePath = "data/test.csv.gz"
localFilePath = "data/test.csv"
util.extract_gz( zipLocalFilePath, localFilePath )
# -
tdf = pd.read_csv(zipLocalFilePath, dtype = object)
tdf.head()
tdf['target_value'] = tdf['target_value'].astype('float')
# Let us plot one time series first.
tdf[tdf['item_id'] == 'client_1'][-24*7*2:]\
.plot(x='timestamp', y='target_value', figsize=(15, 8));
# Next, we use an item meta information dataset that contains the information for both the non-cold start items (client 0 to 369) and cold start items (client 370 to 373). We call this meta information "type" in this specific case. Only one categorical feature is used in this demo, but in practice one normally has multiple categorical features.
#
# Note that for cold start items where little to none demand history exists, the algorithm can only "transfer" information from the existing items to the new ones through the meta information. Therefore, having informative and high quality meta data is the key for a good cold-start forecast.
# +
# this metadata contains the cold start items' metadata as well.
localItemMetaDataFilePath = "data/itemMetaData.csv"
imdf = pd.read_csv(localItemMetaDataFilePath, dtype = object)
imdf.tail()
# -
# And the following figure shows the histogram of the category "type."
imdf['type'].value_counts().plot(kind='bar');
s3 = session.client('s3')
# +
targetTimeseriesDatakey = "cold-start/test.csv"
s3.upload_file(Filename=localFilePath, Bucket = bucketName, Key = f"{targetTimeseriesDatakey}")
# +
itemMetaDatakey = "cold-start/itemMetaData.csv"
s3.upload_file(Filename=localItemMetaDataFilePath, Bucket = bucketName, Key = f"{itemMetaDatakey}")
# -
project = "coldstart_demo"
# Below, we specify key input data and forecast parameters
freq = "H"
forecast_horizon = 48
timestamp_format = "yyyy-MM-dd HH:mm:ss"
delimiter = ','
# ## Step 2a. Creating a Dataset Group<a class="anchor" id="create">
# First let's create a dataset group and then update it later to add our datasets.
dataset_group = f"{project}_grp"
dataset_arns = []
create_dataset_group_response = forecast.create_dataset_group(Domain="CUSTOM",
DatasetGroupName=dataset_group,
DatasetArns=dataset_arns)
print(f'Creating dataset group {dataset_group}')
dataset_group_arn = create_dataset_group_response['DatasetGroupArn']
forecast.describe_dataset_group(DatasetGroupArn=dataset_group_arn)
# ## Step 2b. Creating a Target Dataset<a class="anchor" id="target">
# In this example, we will define a target time series. This is a required dataset to use the service.
# Below we specify the target time series name af_demo_ts_4.
ts_dataset_name = f"{project}_ts"
print(ts_dataset_name)
# Next, we specify the schema of our dataset below. Make sure the order of the attributes (columns) matches the raw
# data in the files. We follow the same three attribute format as the above example.
ts_schema_val = [{"AttributeName": "timestamp", "AttributeType": "timestamp"},
{"AttributeName": "target_value", "AttributeType": "float"},
{"AttributeName": "item_id", "AttributeType": "string"}]
ts_schema = {"Attributes": ts_schema_val}
print(f'Creating target dataset {ts_dataset_name}')
response = forecast.create_dataset(Domain="CUSTOM",
DatasetType='TARGET_TIME_SERIES',
DatasetName=ts_dataset_name,
DataFrequency=freq,
Schema=ts_schema
)
ts_dataset_arn = response['DatasetArn']
forecast.describe_dataset(DatasetArn=ts_dataset_arn)
# ## Step 2c. Creating ItemMetaData Dataset<a class="anchor" id="related">
# In this example, we will define a Item Metadata Dataset.
# Specify the related time series name af_demo_rts_4.
item_metadata_dataset_name = f"{project}_meta"
print(item_metadata_dataset_name)
# Specify the schema of your dataset here. Make sure the order of columns matches the raw data files. We follow the same three column format as the above example.
meta_schema_val = [{"AttributeName": "item_id", "AttributeType": "string"},
{"AttributeName": "category", "AttributeType": "string"}]
meta_schema = {"Attributes": meta_schema_val}
print(f'Creating related dataset {meta_schema}')
response = forecast.create_dataset(Domain="CUSTOM",
DatasetType='ITEM_METADATA',
DatasetName=item_metadata_dataset_name,
Schema=meta_schema
)
meta_dataset_arn = response['DatasetArn']
forecast.describe_dataset(DatasetArn=meta_dataset_arn)
# ## Step 2d. Updating the dataset group with the datasets we created<a class="anchor" id="update">
# You can have multiple datasets under the same dataset group. Update it with the datasets we created before.
dataset_arns = []
dataset_arns.append(ts_dataset_arn)
dataset_arns.append(meta_dataset_arn)
forecast.update_dataset_group(DatasetGroupArn=dataset_group_arn, DatasetArns=dataset_arns)
forecast.describe_dataset_group(DatasetGroupArn=dataset_group_arn)
# ## Step 2e. Creating a Target Time Series Dataset Import Job<a class="anchor" id="targetImport">
ts_dataset_import_job_response = forecast.create_dataset_import_job(DatasetImportJobName=dataset_group+"_1",
DatasetArn=ts_dataset_arn,
DataSource= {
"S3Config" : {
"Path": f"s3://{bucketName}/{targetTimeseriesDatakey}",
"RoleArn": role_arn
}
},
TimestampFormat=timestamp_format)
ts_dataset_import_job_arn=ts_dataset_import_job_response['DatasetImportJobArn']
status = util.wait(lambda: forecast.describe_dataset_import_job(DatasetImportJobArn=ts_dataset_import_job_arn))
assert status
# ## Step 2f. Creating a Item Meta Data Dataset Import Job<a class="anchor" id="relatedImport">
meta_dataset_import_job_response = forecast.create_dataset_import_job(DatasetImportJobName=dataset_group,
DatasetArn=meta_dataset_arn,
DataSource= {
"S3Config" : {
"Path": f"s3://{bucketName}/{itemMetaDatakey}",
"RoleArn": role_arn
}
})
meta_dataset_import_job_arn=meta_dataset_import_job_response['DatasetImportJobArn']
status = util.wait(lambda: forecast.describe_dataset_import_job(DatasetImportJobArn=meta_dataset_import_job_arn))
assert status
# # Step 3. Create Predictor with the datasets<a class="anchor" id="algo">
algorithm_arn = 'arn:aws:forecast:::algorithm/'
algorithm = 'Deep_AR_Plus'
algorithm_arn_deep_ar_plus = algorithm_arn + algorithm
predictor_name_deep_ar = f'{project}_{algorithm.lower()}'
print(f'[{predictor_name_deep_ar}] Creating predictor {predictor_name_deep_ar} ...')
create_predictor_response = forecast.create_predictor(PredictorName=predictor_name_deep_ar,
AlgorithmArn=algorithm_arn_deep_ar_plus,
ForecastHorizon=forecast_horizon,
PerformAutoML=False,
PerformHPO=False,
InputDataConfig= {"DatasetGroupArn": dataset_group_arn},
FeaturizationConfig= {"ForecastFrequency": freq}
)
predictor_arn_deep_ar = create_predictor_response['PredictorArn']
status = util.wait(lambda: forecast.describe_predictor(PredictorArn=predictor_arn_deep_ar))
assert status
forecast.describe_predictor(PredictorArn=predictor_arn_deep_ar)
# # Step 4. Creating a Forecast<a class="anchor" id="forecast">
#
# Next we re-train with the full dataset, and create the forecast.
print(f"Done fetching accuracy numbers. Creating forecaster for DeepAR+ ...")
forecast_name_deep_ar = f'{project}_deep_ar_plus_1'
create_forecast_response_deep_ar = forecast.create_forecast(ForecastName=forecast_name_deep_ar,
PredictorArn=predictor_arn_deep_ar)
forecast_arn_deep_ar_1 = create_forecast_response_deep_ar['ForecastArn']
status = util.wait(lambda: forecast.describe_forecast(ForecastArn=forecast_arn_deep_ar_1))
assert status
forecast.describe_forecast(ForecastArn=forecast_arn_deep_ar_1)
# # Step 5. Creating a Cold-Start Target Time Series Dataset Import Job<a class="anchor" id="coldStartImport">
#
# Now we are ready to generate the forecasts for the cold start problem. Notice that there is a system constraint such that at least 5 rows needs to be present for each item. Therefore, for the item that has less than 5 observations, we fill in with NaNs. In the following example, both Client 370 and 372 have zero observation, i.e., pure cold-start problem, while the other two have 5 target values.
localColdStartDataFilePath = "data/coldStartTargetData.csv"
# validation logic for at least 5 items
cstdf = pd.read_csv(localColdStartDataFilePath, dtype = object)
cstdf.head(20)
coldStartTargetTimeseriesDatakey = "cold-start/coldStartTargetData.csv"
s3.upload_file(Filename=localColdStartDataFilePath,
Bucket = bucketName,
Key = f"{coldStartTargetTimeseriesDatakey}")
ts_cold_start_dataset_import_job_response = forecast.create_dataset_import_job(DatasetImportJobName=dataset_group+"_2",
DatasetArn=ts_dataset_arn,
DataSource= {
"S3Config" : {
"Path": f"s3://{bucketName}/{coldStartTargetTimeseriesDatakey}",
"RoleArn": role_arn
}
},
TimestampFormat=timestamp_format)
ts_cold_start_dataset_import_job_arn = ts_cold_start_dataset_import_job_response['DatasetImportJobArn']
forecast.describe_dataset_import_job(DatasetImportJobArn=ts_cold_start_dataset_import_job_arn)
status = util.wait(lambda: forecast.describe_dataset_import_job(DatasetImportJobArn=ts_cold_start_dataset_import_job_arn))
assert status
# # Step 6. Create a ColdStart Item Forecast<a class="anchor" id="createColdStart">
#
#
# Now we are ready to create the forecasts for all cold start items.
forecast_name_deep_ar_2 = f'{project}_deep_ar_plus_2'
create_forecast_response_deep_ar_2 = forecast.create_forecast(ForecastName=forecast_name_deep_ar_2,
PredictorArn=predictor_arn_deep_ar)
forecast_arn_deep_ar_2 = create_forecast_response_deep_ar_2['ForecastArn']
status = util.wait(lambda: forecast.describe_forecast(ForecastArn=forecast_arn_deep_ar_2))
assert status
forecast.describe_forecast(ForecastArn=forecast_arn_deep_ar_2)
# # Step 7. Querying the ColdStart Item Forecast<a class="anchor" id="query">
#
# Now we plot the forecast, where the first vertical line is the forecast start date.
# +
forecast_response_deep_2 = forecast_query.query_forecast(
ForecastArn=forecast_arn_deep_ar_2,
Filters={"item_id": "client_370"})
fcst = forecast_response_deep_2['Forecast']['Predictions']
time_stamp = list(map(lambda x: pd.to_datetime(x['Timestamp']), fcst['p10']))
p10_fcst = list(map(lambda x: x['Value'], fcst['p10']))
p50_fcst = list(map(lambda x: x['Value'], fcst['p50']))
p90_fcst = list(map(lambda x: x['Value'], fcst['p90']))
plt.figure(figsize=(15, 10))
plt.plot(time_stamp, p50_fcst)
plt.fill_between(time_stamp, p10_fcst, p90_fcst, alpha=0.2)
plt.title("DeepAR Forecast 2");
# -
# # Step 8. Exporting your Forecasts<a class="anchor" id="export">
forecast_export_name_deep_ar = f'{project}_cold_start_forecast_export_deep_ar_plus'
forecast_export_name_deep_ar_path = f"s3://{bucketName}/{forecast_export_name_deep_ar}"
create_forecast_export_response_deep_ar = forecast.create_forecast_export_job(ForecastExportJobName=forecast_export_name_deep_ar,
ForecastArn=forecast_arn_deep_ar_2,
Destination={
"S3Config" : {
"Path": forecast_export_name_deep_ar_path,
"RoleArn": role_arn
}
})
forecast_export_arn_deep_ar = create_forecast_export_response_deep_ar['ForecastExportJobArn']
status = util.wait(lambda: forecast.describe_forecast_export_job(ForecastExportJobArn = forecast_export_arn_deep_ar))
assert status
# # Step 9. Cleaning up your Resources<a class="anchor" id="cleanup">
# Once we have completed the above steps, we can start to cleanup the resources we created. All delete jobs, except for `delete_dataset_group` are asynchronous, so we have added the helpful `wait_till_delete` function.
# Resource Limits documented <a href="https://docs.aws.amazon.com/forecast/latest/dg/limits.html">here</a>.
# Delete forecast export
util.wait_till_delete(lambda: forecast.delete_forecast_export_job(ForecastExportJobArn = forecast_export_arn_deep_ar))
# Delete forecast
util.wait_till_delete(lambda: forecast.delete_forecast(ForecastArn = forecast_arn_deep_ar_1))
util.wait_till_delete(lambda: forecast.delete_forecast(ForecastArn = forecast_arn_deep_ar_2))
# Delete predictor
util.wait_till_delete(lambda: forecast.delete_predictor(PredictorArn = predictor_arn_deep_ar))
# Delete the target time series and related time series dataset import jobs
util.wait_till_delete(lambda: forecast.delete_dataset_import_job(DatasetImportJobArn=ts_dataset_import_job_arn))
util.wait_till_delete(lambda: forecast.delete_dataset_import_job(DatasetImportJobArn=meta_dataset_import_job_arn))
util.wait_till_delete(lambda: forecast.delete_dataset_import_job(DatasetImportJobArn=ts_cold_start_dataset_import_job_arn))
# Delete the target time series and related time series datasets
util.wait_till_delete(lambda: forecast.delete_dataset(DatasetArn=ts_dataset_arn))
util.wait_till_delete(lambda: forecast.delete_dataset(DatasetArn=meta_dataset_arn))
# Delete dataset group
forecast.delete_dataset_group(DatasetGroupArn=dataset_group_arn)
# Delete IAM role
util.delete_iam_role( role_name )
| notebooks/advanced/Forecast with Cold Start Items/Forecast with Cold Start Items.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python2
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] inputHidden=false outputHidden=false
# #### New to Plotly?
# Plotly's Python library is free and open source! [Get started](https://plot.ly/python/getting-started/) by downloading the client and [reading the primer](https://plot.ly/python/getting-started/).
# <br>You can set up Plotly to work in [online](https://plot.ly/python/getting-started/#initialization-for-online-plotting) or [offline](https://plot.ly/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plot.ly/python/getting-started/#start-plotting-online).
# <br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
# -
# #### Version Check
# Plotly's python package is updated frequently. Run `pip install plotly --upgrade` to use the latest version.
# + inputHidden=false outputHidden=false
import plotly
plotly.__version__
# -
# ### Basic Carpet Plot
#
# Set the `x` and `y` coorindates, using `x` and `y` attributes. If `x` coorindate values are ommitted a cheater plot will be created. To save parameter values use `a` and `b` attributes. To make changes to the axes, use `aaxis` or `baxis` attributes. For a more detailed list of axes attributes refer to [python reference](https://plot.ly/python/reference/#carpet-aaxis).
# +
import plotly.graph_objs as go
import plotly.plotly as py
trace1 = go.Carpet(
a = [0, 1, 2, 3, 0, 1, 2, 3, 0, 1, 2, 3],
b = [4, 4, 4, 4, 5, 5, 5, 5, 6, 6, 6, 6],
x = [2, 3, 4, 5, 2.2, 3.1, 4.1, 5.1, 1.5, 2.5, 3.5, 4.5],
y = [1, 1.4, 1.6, 1.75, 2, 2.5, 2.7, 2.75, 3, 3.5, 3.7, 3.75],
aaxis = dict(
tickprefix = 'a = ',
smoothing = 0,
minorgridcount = 9,
type = 'linear'
),
baxis = dict(
tickprefix = 'b = ',
smoothing = 0,
minorgridcount = 9,
type = 'linear'
)
)
data = [trace1]
layout = go.Layout(
margin = dict(
t = 40,
r = 30,
b = 30,
l = 30
),
yaxis = dict(
range = [0.388,4.361]
),
xaxis = dict(
range = [0.667,5.932]
)
)
fig = go.Figure(data = data, layout = layout)
py.iplot(fig, filename = "contourcarpet/basic")
# -
# ### Add Contours
# + inputHidden=false outputHidden=false
import plotly.graph_objs as go
import plotly.plotly as py
trace1 = go.Contourcarpet(
a = [0, 1, 2, 3, 0, 1, 2, 3, 0, 1, 2, 3],
b = [4, 4, 4, 4, 5, 5, 5, 5, 6, 6, 6, 6],
z = [1, 1.96, 2.56, 3.0625, 4, 5.0625, 1, 7.5625, 9, 12.25, 15.21, 14.0625],
autocontour = False,
contours = dict(
start = 1,
end = 14,
size = 1
),
line = dict(
width = 2,
smoothing = 0
),
colorbar = dict(
len = 0.4,
y = 0.25
)
)
trace2 = go.Carpet(
a = [0, 1, 2, 3, 0, 1, 2, 3, 0, 1, 2, 3],
b = [4, 4, 4, 4, 5, 5, 5, 5, 6, 6, 6, 6],
x = [2, 3, 4, 5, 2.2, 3.1, 4.1, 5.1, 1.5, 2.5, 3.5, 4.5],
y = [1, 1.4, 1.6, 1.75, 2, 2.5, 2.7, 2.75, 3, 3.5, 3.7, 3.75],
aaxis = dict(
tickprefix = 'a = ',
smoothing = 0,
minorgridcount = 9,
type = 'linear'
),
baxis = dict(
tickprefix = 'b = ',
smoothing = 0,
minorgridcount = 9,
type = 'linear'
)
)
data = [trace1, trace2]
layout = go.Layout(
margin = dict(
t = 40,
r = 30,
b = 30,
l = 30
),
yaxis = dict(
range = [0.388,4.361]
),
xaxis = dict(
range = [0.667,5.932]
)
)
fig = go.Figure(data = data, layout = layout)
py.iplot(fig, filename = "contourcarpet/add-contours")
# -
# ### Add Multiple Traces
# + inputHidden=false outputHidden=false
import plotly.graph_objs as go
import plotly.plotly as py
import urllib, json
url = "https://raw.githubusercontent.com/bcdunbar/datasets/master/airfoil_data.json"
response = urllib.urlopen(url)
data = json.loads(response.read())
trace1 = go.Carpet(
a = data[0]['a'],
b = data[0]['b'],
x = data[0]['x'],
y = data[0]['y'],
baxis = dict(
startline = False,
endline = False,
showticklabels = "none",
smoothing = 0,
showgrid = False
),
aaxis = dict(
startlinewidth = 2,
startline = True,
showticklabels = "none",
endline = True,
showgrid = False,
endlinewidth = 2,
smoothing = 0
)
)
trace2 = go.Contourcarpet(
z = data[1]['z'],
autocolorscale = False,
zmax = 1,
name = "Pressure",
colorscale = "Viridis",
zmin = -8,
colorbar = dict(
y = 0,
yanchor = "bottom",
titleside = "right",
len = 0.75,
title = "Pressure coefficient, c<sub>p</sub>"
),
contours = dict(
start = -1,
size = 0.025,
end = 1.000,
showlines = False
),
line = dict(
smoothing = 0
),
autocontour = False,
zauto = False
)
trace3 = go.Contourcarpet(
z = data[2]['z'],
opacity = 0.300,
showlegend = True,
name = "Streamlines",
autocontour = True,
ncontours = 50,
contours = dict(
coloring = "none"
),
line = dict(
color = "white",
width = 1
)
)
trace4 = go.Contourcarpet(
z = data[3]['z'],
showlegend = True,
name = "Pressure<br>contours",
autocontour = False,
line = dict(
color = "rgba(0, 0, 0, 0.5)",
smoothing = 1
),
contours = dict(
size = 0.250,
start = -4,
coloring = "none",
end = 1.000,
showlines = True
)
)
trace5 = go.Scatter(
x = data[4]['x'],
y = data[4]['y'],
legendgroup = "g1",
name = "Surface<br>pressure",
mode = "lines",
hoverinfo = "skip",
line = dict(
color = "rgba(255, 0, 0, 0.5)",
width = 1,
shape = "spline",
smoothing = 1
),
fill = "toself",
fillcolor = "rgba(255, 0, 0, 0.2)"
)
trace6 = go.Scatter(
x = data[5]['x'],
y = data[5]['y'],
showlegend = False,
legendgroup = "g1",
mode = "lines",
hoverinfo = "skip",
line = dict(
color = "rgba(255, 0, 0, 0.3)",
width = 1
)
)
trace7 = go.Scatter(
x = data[6]['x'],
y = data[6]['y'],
showlegend = False,
legendgroup = "g1",
name = "cp",
text = data[6]['text'],
hoverinfo = "text",
mode = "lines",
line = dict(
color = "rgba(255, 0, 0, 0.2)",
width = 0
)
)
data = [trace1,trace2,trace3,trace4,trace5,trace6,trace7]
layout = go.Layout(
yaxis = dict(
zeroline = False,
range = [-1.800,1.800],
showgrid = False
),
dragmode = "pan",
height = 700,
xaxis = dict(
zeroline = False,
scaleratio = 1,
scaleanchor = 'y',
range = [-3.800,3.800],
showgrid = False
),
title = "Flow over a Karman-Trefftz airfoil",
hovermode = "closest",
margin = dict(
r = 60,
b = 40,
l = 40,
t = 80
),
width = 900
)
fig = go.Figure(data=data,layout=layout)
py.iplot(fig, filename = "contourcarpet/airfoil")
# -
# ### Reference
# See https://plot.ly/python/reference/#contourcarpet for more information and chart attribute options!
# +
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
# ! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'contourcarpet.ipynb', 'python/carpet-contour/', 'Carpet Contour Plot',
'How to make carpet contour plots in Python with Plotly.',
title = 'Carpet Contour Plots | Plotly',
has_thumbnail='true', thumbnail='thumbnail/contourcarpet.jpg',
language='python',
# page_type='example_index', // note this is only if you want the tutorial to appear on the main page: plot.ly/python
display_as='scientific', order=27,
ipynb= '~notebook_demo/145')
# -
| _posts/python-v3/scientific/carpet-contour/contourcarpet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Regression Week 2: Multiple Regression (gradient descent)
# In the first notebook we explored multiple regression using Turi Create. Now we will use Turi Create along with numpy to solve for the regression weights with gradient descent.
#
# In this notebook we will cover estimating multiple regression weights via gradient descent. You will:
# * Add a constant column of 1's to a Turi Create SFrame to account for the intercept
# * Convert an SFrame into a Numpy array
# * Write a predict_output() function using Numpy
# * Write a numpy function to compute the derivative of the regression weights with respect to a single feature
# * Write gradient descent function to compute the regression weights given an initial weight vector, step size and tolerance.
# * Use the gradient descent function to estimate regression weights for multiple features
# # Fire up Turi Create
# Make sure you have the latest version of Turi Create
import turicreate
# # Load in house sales data
#
# Dataset is from house sales in King County, the region where the city of Seattle, WA is located.
sales = turicreate.SFrame('home_data.sframe/')
# If we want to do any "feature engineering" like creating new features or adjusting existing ones we should do this directly using the SFrames as seen in the other Week 2 notebook. For this notebook, however, we will work with the existing features.
# # Convert to Numpy Array
# Although SFrames offer a number of benefits to users (especially when using Big Data and built-in Turi Create functions) in order to understand the details of the implementation of algorithms it's important to work with a library that allows for direct (and optimized) matrix operations. Numpy is a Python solution to work with matrices (or any multi-dimensional "array").
#
# Recall that the predicted value given the weights and the features is just the dot product between the feature and weight vector. Similarly, if we put all of the features row-by-row in a matrix then the predicted value for *all* the observations can be computed by right multiplying the "feature matrix" by the "weight vector".
#
# First we need to take the SFrame of our data and convert it into a 2D numpy array (also called a matrix). To do this we use Turi Create's built in .to_dataframe() which converts the SFrame into a Pandas (another python library) dataframe. We can then use Panda's .as_matrix() to convert the dataframe into a numpy matrix.
import numpy as np # note this allows us to refer to numpy as np instead
# Now we will write a function that will accept an SFrame, a list of feature names (e.g. ['sqft_living', 'bedrooms']) and an target feature e.g. ('price') and will return two things:
# * A numpy matrix whose columns are the desired features plus a constant column (this is how we create an 'intercept')
# * A numpy array containing the values of the output
#
# With this in mind, complete the following function (where there's an empty line you should write a line of code that does what the comment above indicates)
def get_numpy_data(data_sframe, features, output):
data_sframe['constant'] = 1 # this is how you add a constant column to an SFrame
# add the column 'constant' to the front of the features list so that we can extract it along with the others:
features = ['constant'] + features # this is how you combine two lists
# select the columns of data_SFrame given by the features list into the SFrame features_sframe (now including constant):
features_sframe = data_sframe[features]
# the following line will convert the features_SFrame into a numpy matrix:
feature_matrix = features_sframe.to_numpy()
# assign the column of data_sframe associated with the output to the SArray output_sarray
output_sarray = data_sframe[output]
# the following will convert the SArray into a numpy array by first converting it to a list
output_array = output_sarray.to_numpy()
return(feature_matrix, output_array)
# For testing let's use the 'sqft_living' feature and a constant as our features and price as our output:
(example_features, example_output) = get_numpy_data(sales, ['sqft_living'], 'price') # the [] around 'sqft_living' makes it a list
print (example_features[0,:]) # this accesses the first row of the data the ':' indicates 'all columns'
print (example_output[0]) # and the corresponding output
# # Predicting output given regression weights
# Suppose we had the weights [1.0, 1.0] and the features [1.0, 1180.0] and we wanted to compute the predicted output 1.0\*1.0 + 1.0\*1180.0 = 1181.0 this is the dot product between these two arrays. If they're numpy arrayws we can use np.dot() to compute this:
my_weights = np.array([1., 1.]) # the example weights
my_features = example_features[0,] # we'll use the first data point
predicted_value = np.dot(my_features, my_weights)
print (predicted_value)
# np.dot() also works when dealing with a matrix and a vector. Recall that the predictions from all the observations is just the RIGHT (as in weights on the right) dot product between the features *matrix* and the weights *vector*. With this in mind finish the following predict_output function to compute the predictions for an entire matrix of features given the matrix and the weights:
def predict_output(feature_matrix, weights):
# assume feature_matrix is a numpy matrix containing the features as columns and weights is a corresponding numpy array
# create the predictions vector by using np.dot()
predictions = np.dot(feature_matrix, weights)
return(predictions)
# If you want to test your code run the following cell:
test_predictions = predict_output(example_features, my_weights)
print (test_predictions[0]) # should be 1181.0
print (test_predictions[1]) # should be 2571.0
# # Computing the Derivative
# We are now going to move to computing the derivative of the regression cost function. Recall that the cost function is the sum over the data points of the squared difference between an observed output and a predicted output.
#
# Since the derivative of a sum is the sum of the derivatives we can compute the derivative for a single data point and then sum over data points. We can write the squared difference between the observed output and predicted output for a single point as follows:
#
# (w[0]\*[CONSTANT] + w[1]\*[feature_1] + ... + w[i] \*[feature_i] + ... + w[k]\*[feature_k] - output)^2
#
# Where we have k features and a constant. So the derivative with respect to weight w[i] by the chain rule is:
#
# 2\*(w[0]\*[CONSTANT] + w[1]\*[feature_1] + ... + w[i] \*[feature_i] + ... + w[k]\*[feature_k] - output)\* [feature_i]
#
# The term inside the paranethesis is just the error (difference between prediction and output). So we can re-write this as:
#
# 2\*error\*[feature_i]
#
# That is, the derivative for the weight for feature i is the sum (over data points) of 2 times the product of the error and the feature itself. In the case of the constant then this is just twice the sum of the errors!
#
# Recall that twice the sum of the product of two vectors is just twice the dot product of the two vectors. Therefore the derivative for the weight for feature_i is just two times the dot product between the values of feature_i and the current errors.
#
# With this in mind complete the following derivative function which computes the derivative of the weight given the value of the feature (over all data points) and the errors (over all data points).
def feature_derivative(errors, feature):
# Assume that errors and feature are both numpy arrays of the same length (number of data points)
# compute twice the dot product of these vectors as 'derivative' and return the value
derivative = 2 * np.dot(errors, feature)
return(derivative)
# To test your feature derivartive run the following:
(example_features, example_output) = get_numpy_data(sales, ['sqft_living'], 'price')
my_weights = np.array([0., 0.]) # this makes all the predictions 0
test_predictions = predict_output(example_features, my_weights)
# just like SFrames 2 numpy arrays can be elementwise subtracted with '-':
errors = test_predictions - example_output # prediction errors in this case is just the -example_output
feature = example_features[:,0] # let's compute the derivative with respect to 'constant', the ":" indicates "all rows"
derivative = feature_derivative(errors, feature)
print (derivative)
print (-np.sum(example_output)*2) # should be the same as derivative
# # Gradient Descent
# Now we will write a function that performs a gradient descent. The basic premise is simple. Given a starting point we update the current weights by moving in the negative gradient direction. Recall that the gradient is the direction of *increase* and therefore the negative gradient is the direction of *decrease* and we're trying to *minimize* a cost function.
#
# The amount by which we move in the negative gradient *direction* is called the 'step size'. We stop when we are 'sufficiently close' to the optimum. We define this by requiring that the magnitude (length) of the gradient vector to be smaller than a fixed 'tolerance'.
#
# With this in mind, complete the following gradient descent function below using your derivative function above. For each step in the gradient descent we update the weight for each feature befofe computing our stopping criteria
from math import sqrt # recall that the magnitude/length of a vector [g[0], g[1], g[2]] is sqrt(g[0]^2 + g[1]^2 + g[2]^2)
def regression_gradient_descent(feature_matrix, output, initial_weights, step_size, tolerance):
converged = False
weights = np.array(initial_weights) # make sure it's a numpy array
while not converged:
# compute the predictions based on feature_matrix and weights using your predict_output() function
predictions = predict_output(feature_matrix, weights)
# compute the errors as predictions - output
errors = predictions - output
gradient_sum_squares = 0 # initialize the gradient sum of squares
# while we haven't reached the tolerance yet, update each feature's weight
for i in range(len(weights)): # loop over each weight
# Recall that feature_matrix[:, i] is the feature column associated with weights[i]
# compute the derivative for weight[i]:
derivative = feature_derivative(errors, feature_matrix[:,i])
# add the squared value of the derivative to the gradient sum of squares (for assessing convergence)
gradient_sum_squares = (derivative * derivative) + gradient_sum_squares
# subtract the step size times the derivative from the current weight
weights[i] = weights[i] - (step_size * derivative)
# compute the square-root of the gradient sum of squares to get the gradient magnitude:
gradient_magnitude = sqrt(gradient_sum_squares)
if gradient_magnitude < tolerance:
converged = True
return(weights)
# A few things to note before we run the gradient descent. Since the gradient is a sum over all the data points and involves a product of an error and a feature the gradient itself will be very large since the features are large (squarefeet) and the output is large (prices). So while you might expect "tolerance" to be small, small is only relative to the size of the features.
#
# For similar reasons the step size will be much smaller than you might expect but this is because the gradient has such large values.
# # Running the Gradient Descent as Simple Regression
# First let's split the data into training and test data.
train_data,test_data = sales.random_split(.8,seed=0)
# Although the gradient descent is designed for multiple regression since the constant is now a feature we can use the gradient descent function to estimate the parameters in the simple regression on squarefeet. The folowing cell sets up the feature_matrix, output, initial weights and step size for the first model:
# let's test out the gradient descent
simple_features = ['sqft_living']
my_output = 'price'
(simple_feature_matrix, output) = get_numpy_data(train_data, simple_features, my_output)
initial_weights = np.array([-47000., 1.])
step_size = 7e-12
tolerance = 2.5e7
# Next run your gradient descent with the above parameters.
weights1 = regression_gradient_descent(simple_feature_matrix, output, initial_weights, step_size, tolerance)
print ("Weights: " + str(weights1))
# How do your weights compare to those achieved in week 1 (don't expect them to be exactly the same)?
#
# **Quiz Question: What is the value of the weight for sqft_living -- the second element of ‘simple_weights’ (rounded to 1 decimal place)?**
# Use your newly estimated weights and your predict_output() function to compute the predictions on all the TEST data (you will need to create a numpy array of the test feature_matrix and test output first:
(test_simple_feature_matrix, test_output) = get_numpy_data(test_data, simple_features, my_output)
# Now compute your predictions using test_simple_feature_matrix and your weights from above.
my_predictions1 = predict_output(test_simple_feature_matrix, np.asarray(weights1))
# **Quiz Question: What is the predicted price for the 1st house in the TEST data set for model 1 (round to nearest dollar)?**
print (my_predictions1[0])
# Now that you have the predictions on test data, compute the RSS on the test data set. Save this value for comparison later. Recall that RSS is the sum of the squared errors (difference between prediction and output).
output = test_data['price']
residuals = my_predictions1 - output
RSS1 = (residuals**2).sum()
# # Running a multiple regression
# Now we will use more than one actual feature. Use the following code to produce the weights for a second model with the following parameters:
model_features = ['sqft_living', 'sqft_living15'] # sqft_living15 is the average squarefeet for the nearest 15 neighbors.
my_output = 'price'
(feature_matrix, output) = get_numpy_data(train_data, model_features, my_output)
initial_weights = np.array([-100000., 1., 1.])
step_size = 4e-12
tolerance = 1e9
# Use the above parameters to estimate the model weights. Record these values for your quiz.
weights2 = regression_gradient_descent(feature_matrix, output, initial_weights, step_size, tolerance)
print (weights2)
# Use your newly estimated weights and the predict_output function to compute the predictions on the TEST data. Don't forget to create a numpy array for these features from the test set first!
my_predictions2 = predict_output(feature_matrix, np.asarray(weights2))
# **Quiz Question: What is the predicted price for the 1st house in the TEST data set for model 2 (round to nearest dollar)?**
print (my_predictions2[0])
# What is the actual price for the 1st house in the test data set?
test_data[0]['price']
# **Quiz Question: Which estimate was closer to the true price for the 1st house on the TEST data set, model 1 or model 2?**
# Now use your predictions and the output to compute the RSS for model 2 on TEST data.
residuals = my_predictions2 - output
RSS2 = (residuals**2).sum()
# **Quiz Question: Which model (1 or 2) has lowest RSS on all of the TEST data? **
print (RSS1 < RSS2)
| Courses/Machine Learning Regression/Regression Week 2 Multiple Regression (gradient descent).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Excercise 1 Task 2
# ## Examination of runtime improvments of ensemble classifiers on a 250k elements dataset in dependence of the number of available cores
# ### This notebook should run on an 8-core server environment to provide similar results
# Load neccessary libraries changed pandas import for convinience
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.datasets import make_classification
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
# creation of a dataset consisting of 250k samples
# with the following parameters
samples = 250*1000
features = 40
informative = 5
redundant=4
X, Y = make_classification(n_samples=samples,
n_features=features,
n_informative=informative,
n_redundant=4)
# Split-out validation dataset
validation_size = 0.20
seed = 7
scoring = 'accuracy'
X_train, X_validation, Y_train, Y_validation = train_test_split(X,
Y,
test_size=validation_size,
random_state=seed)
# ## Using 8 estimators (usage of one per core if 8 cores (jobs) are used)
# ### One RandomForestClassifier (RFC) for each number of jobs (1 to 8 (inclusive)) is instantiated and trained on the training set of 200k elements. During the training the train time is measured with the magic %timeit function and stored in an array.
# Create Random Forest Classifier
estimators = 8 # For mapping one estimator per core in case of max 8 cores
jobs = 8
time_it_results = []
for _ in range(jobs):
rf_class = RandomForestClassifier(n_estimators=estimators, n_jobs=(_+1))
# tr = %timeit -o rf_class.fit(X_train, Y_train)
time_it_results.append(tr)
# best_times are extracted
best_times = [timer.best for timer in time_it_results]
# ## Plot of the training time in seconds of each RFC against the number of used cores (number of jobs)
x = np.arange(1,9)
labels = ['%i. Core' % i for i in x]
fig = plt.figure()
fig.suptitle('Training Time per number of cores')
ax = fig.add_subplot(111)
ax.set_xlabel('Number of cores')
ax.set_ylabel('Training time (s)')
ax.plot(x, best_times)
plt.xticks(x, labels, rotation='vertical')
plt.show()
# Execution time is exponentially decreasing till 4 cpu cores are utilized. Further increase and decrease dependes on mainly two factors.
# - Overhead intruduced by managing multiprocessing
# - Overhead introduced by copying the datasets for processing
#
# A slight increase in runtime between 4 and 7 cores can be experienced till 8 cpu cores are utilized
| notebooks/henrik_ueb01/02_Classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Evaluate a polynomial string
def symbolize(s):
"""
Converts a a string (equation) to a SymPy symbol object
"""
from sympy import sympify
s1=s.replace('.','*')
s2=s1.replace('^','**')
s3=sympify(s2)
return(s3)
def eval_multinomial(s,vals=None,symbolic_eval=False):
"""
Evaluates polynomial at vals.
vals can be simple list, dictionary, or tuple of values.
vals can also contain symbols instead of real values provided those symbols have been declared before using SymPy
"""
from sympy import Symbol
sym_s=symbolize(s)
sym_set=sym_s.atoms(Symbol)
sym_lst=[]
for s in sym_set:
sym_lst.append(str(s))
sym_lst.sort()
if symbolic_eval==False and len(sym_set)!=len(vals):
print("Length of the input values did not match number of variables and symbolic evaluation is not selected")
return None
else:
if type(vals)==list:
sub=list(zip(sym_lst,vals))
elif type(vals)==dict:
l=list(vals.keys())
l.sort()
lst=[]
for i in l:
lst.append(vals[i])
sub=list(zip(sym_lst,lst))
elif type(vals)==tuple:
sub=list(zip(sym_lst,list(vals)))
result=sym_s.subs(sub)
return result
# ### Helper function for flipping binary values of a _ndarray_
def flip(y,p):
import numpy as np
lst=[]
for i in range(len(y)):
f=np.random.choice([1,0],p=[p,1-p])
lst.append(f)
lst=np.array(lst)
return np.array(np.logical_xor(y,lst),dtype=int)
# ### Classification sample generation based on a symbolic expression
def gen_classification_symbolic(m=None,n_samples=100,n_features=2,flip_y=0.0):
"""
Generates classification sample based on a symbolic expression.
Calculates the output of the symbolic expression at randomly generated (Gaussian distribution) points and
assigns binary classification based on sign.
m: The symbolic expression. Needs x1, x2, etc as variables and regular python arithmatic symbols to be used.
n_samples: Number of samples to be generated
n_features: Number of variables. This is automatically inferred from the symbolic expression. So this is ignored
in case a symbolic expression is supplied. However if no symbolic expression is supplied then a
default simple polynomial can be invoked to generate classification samples with n_features.
flip_y: Probability of flipping the classification labels randomly. A higher value introduces more noise and make
the classification problem harder.
Returns a numpy ndarray with dimension (n_samples,n_features+1). Last column is the response vector.
"""
import numpy as np
from sympy import Symbol,sympify
if m==None:
m=''
for i in range(1,n_features+1):
c='x'+str(i)
c+=np.random.choice(['+','-'],p=[0.5,0.5])
m+=c
m=m[:-1]
sym_m=sympify(m)
n_features=len(sym_m.atoms(Symbol))
evals=[]
lst_features=[]
for i in range(n_features):
lst_features.append(np.random.normal(scale=5,size=n_samples))
lst_features=np.array(lst_features)
lst_features=lst_features.T
for i in range(n_samples):
evals.append(eval_multinomial(m,vals=list(lst_features[i])))
evals=np.array(evals)
evals_binary=evals>0
evals_binary=evals_binary.flatten()
evals_binary=np.array(evals_binary,dtype=int)
evals_binary=flip(evals_binary,p=flip_y)
evals_binary=evals_binary.reshape(n_samples,1)
lst_features=lst_features.reshape(n_samples,n_features)
x=np.hstack((lst_features,evals_binary))
return (x)
x=gen_classification_symbolic(m='2*x1+3*x2+5*x3',n_samples=10,flip_y=0.0)
import pandas as pd
df=pd.DataFrame(x)
df
x=gen_classification_symbolic(m='12*x1/(x2+5*x3)',n_samples=10,flip_y=0.2)
df=pd.DataFrame(x)
df
# #### Classification samples with linear separator but no noise
x=gen_classification_symbolic(m='x1-2*x2',n_samples=50,flip_y=0.0)
df=pd.DataFrame(x)
plt.scatter(x=df[0],y=df[1],c=df[2])
plt.show()
# #### Classification samples with linear separator but significant noise (flipped bits)
x=gen_classification_symbolic(m='x1-2*x2',n_samples=50,flip_y=0.15)
df=pd.DataFrame(x)
plt.scatter(x=df[0],y=df[1],c=df[2])
plt.show()
import seaborn as sns
# #### Classification samples with non-linear separator
x=gen_classification_symbolic(m='x1**2-x2**2',n_samples=500,flip_y=0.01)
df=pd.DataFrame(x)
plt.scatter(x=df[0],y=df[1],c=df[2])
plt.show()
x=gen_classification_symbolic(m='x1**2-x2**2',n_samples=500,flip_y=0.01)
df=pd.DataFrame(x)
plt.scatter(x=df[0],y=df[1],c=df[2])
plt.show()
# ### Regression sample generation based on a symbolic expression
def gen_regression_symbolic(m=None,n_samples=100,n_features=2,noise=0.0,noise_dist='normal'):
"""
Generates regression sample based on a symbolic expression. Calculates the output of the symbolic expression
at randomly generated (drawn from a Gaussian distribution) points
m: The symbolic expression. Needs x1, x2, etc as variables and regular python arithmatic symbols to be used.
n_samples: Number of samples to be generated
n_features: Number of variables. This is automatically inferred from the symbolic expression. So this is ignored
in case a symbolic expression is supplied. However if no symbolic expression is supplied then a
default simple polynomial can be invoked to generate regression samples with n_features.
noise: Magnitude of Gaussian noise to be introduced (added to the output).
noise_dist: Type of the probability distribution of the noise signal.
Currently supports: Normal, Uniform, t, Beta, Gamma, Poission, Laplace
Returns a numpy ndarray with dimension (n_samples,n_features+1). Last column is the response vector.
"""
import numpy as np
from sympy import Symbol,sympify
if m==None:
m=''
for i in range(1,n_features+1):
c='x'+str(i)
c+=np.random.choice(['+','-'],p=[0.5,0.5])
m+=c
m=m[:-1]
sym_m=sympify(m)
n_features=len(sym_m.atoms(Symbol))
evals=[]
lst_features=[]
for i in range(n_features):
lst_features.append(np.random.normal(scale=5,size=n_samples))
lst_features=np.array(lst_features)
lst_features=lst_features.T
lst_features=lst_features.reshape(n_samples,n_features)
for i in range(n_samples):
evals.append(eval_multinomial(m,vals=list(lst_features[i])))
evals=np.array(evals)
evals=evals.reshape(n_samples,1)
if noise_dist=='normal':
noise_sample=noise*np.random.normal(loc=0,scale=1.0,size=n_samples)
elif noise_dist=='uniform':
noise_sample=noise*np.random.uniform(low=0,high=1.0,size=n_samples)
elif noise_dist=='beta':
noise_sample=noise*np.random.beta(a=0.5,b=1.0,size=n_samples)
elif noise_dist=='Gamma':
noise_sample=noise*np.random.gamma(shape=1.0,scale=1.0,size=n_samples)
elif noise_dist=='laplace':
noise_sample=noise*np.random.laplace(loc=0.0,scale=1.0,size=n_samples)
noise_sample=noise_sample.reshape(n_samples,1)
evals=evals+noise_sample
x=np.hstack((lst_features,evals))
return (x)
# #### Generate samples with a rational function as input
# ### $$\frac{10x_1}{(3x_2+4x_3)}$$
x=gen_regression_symbolic(m='10*x1/(3*x2+4*x3)',n_samples=10,noise=0.1)
df=pd.DataFrame(x)
df
# #### Generate samples with no symbolic input and with 10 features
x=gen_regression_symbolic(n_features=10,n_samples=10,noise=0.1)
df=pd.DataFrame(x)
df
import matplotlib.pyplot as plt
# #### Generate samples with less noise and plot: $0.2x^2+1.2x+6+f_{noise}(x\mid{N=0.1})$
x=gen_regression_symbolic(m='0.2*x**2+1.2*x+6',n_samples=100,noise=0.1)
df=pd.DataFrame(x)
plt.scatter(df[0],df[1],edgecolor='k',alpha=0.7,c='red',s=150)
plt.show()
# #### Generate samples with more noise and plo: $0.2x^2+1.2x+6+f_{noise}(x\mid{N=10})$
x=gen_regression_symbolic(m='0.2*x**2+1.2*x+6',n_samples=100,noise=10)
df=pd.DataFrame(x)
plt.scatter(df[0],df[1],edgecolor='k',alpha=0.7,c='red',s=150)
plt.show()
# #### Generate samples with larger coefficent for the quadratic term and plot: $1.3x^2+1.2x+6+f_{noise}(x\mid{N=10})$
# #### Generate sample with transcedental or rational functions: $x^2.e^{-0.5x}.sin(x+10)$
x=gen_regression_symbolic(m='x**2*exp(-0.5*x)*sin(x+10)',n_samples=50,noise=1)
df=pd.DataFrame(x)
plt.figure(figsize=(10,4))
plt.scatter(df[0],df[1],edgecolor='k',alpha=0.7,c='red',s=150)
plt.grid(True)
plt.show()
| Symbolic regression classification generator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Ejercicio 3 Prácticas LC. Curso 2020-2021
# ### <NAME>
import nltk
from nltk.corpus import wordnet
from nltk.corpus import stopwords
nltk.download('punkt')
nltk.download('stopwords')
nltk.download('wordnet')
# Para esta tarea hemos seguido la estructura de codigo definida en la guia proporcionada
# +
def compute_overlap(signature, context):
ov = sum(1 for word in context if word in signature)
return ov
def simplified_lesk(word, sentence):
best_sense = None
max_overlap = 0
t_sent = nltk.word_tokenize(sentence)
context = []
for w in t_sent:
if w not in stopwords.words('english'):
context.append(w)
senses = wordnet.synsets(word)
for s in senses:
signature = []
defin = s.definition()
for z in nltk.word_tokenize(defin):
signature.append(z)
examp = s.examples()
for z in examp:
signature.append(z)
overlap = compute_overlap(signature, context)
if overlap > max_overlap:
max_overlap = overlap
best_sense = s
return best_sense
# -
# Usando el ejemplo propuesto de la tarea
w = "bank"
s = "Yesterday I went to the bank to withdraw the money and the credit card did not work"
print(simplified_lesk(w, s))
| LC/lab4/lesk.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:finra]
# language: python
# name: conda-env-finra-py
# ---
# Import local files
from manage_path import *
from get_data import *
from compute_lda import *
data = load_pickle(file_name="FINRA_TRACE_2014.pkl.zip")
data.dtypes
# %%time
bag_of_words = trade_SV_BoW(data)
# bag_of_words = trade_vol_BoW(data,"large")
print(bag_of_words)
# +
print(bag_of_words.isna().sum().sum())
# vals = bag_of_words.values
# for i in range(10):
# print(len(vals[i]))
# -
# volume
# (274150 * 11935) - 3266355180
# frac_out
(287800 * 12013) - 3456142416
data['price'] = (data['ENTRD_VOL_QT'] * data['RPTD_PR'])/100
data['price'].describe()
"""Compute Dc_v4 which is count of bonds on given dealer and day seperated buy and sell"""
data['document_date'] = data['TRD_EXCTN_DTTM'].dt.date.apply(lambda x: str(x))
create_buy_document_no_source_vectorize = np.vectorize(create_buy_document_no_source)
create_sell_document_no_source_vectorize = np.vectorize(create_sell_document_no_source)
client_to_delete_vectorize = np.vectorize(client_to_delete)
print("creating documents ......")
# Add new column Dc_v4_S which is the string representation of report dealer buy on the specific day
data['Dc_v4_S'] = create_sell_document_no_source_vectorize(data['Report_Dealer_Index'].values,data['Contra_Party_Index'].values,data['document_date'].values)
# Add new column Dc_v4_B which is the string representation of report dealer sell on the specific day
data['Dc_v4_B'] = create_buy_document_no_source_vectorize(data['Report_Dealer_Index'].values,data['Contra_Party_Index'].values,data['document_date'].values)
print("documents created!!")
type(data['Dc_v4_S'].iloc[7])
1 != 2
data_sub = data[['Dc_v4_S','Dc_v4_B','BOND_SYM_ID','price']].copy()
data_gb_sell = data_sub[data_sub['Dc_v4_S']!='nan'].groupby(by=['Dc_v4_S','BOND_SYM_ID'])
data_gb_buy = data_sub[data_sub['Dc_v4_B']!='nan'].groupby(by=['Dc_v4_B','BOND_SYM_ID'])
# +
#sell_matrix = data_gb_sell['price'].sum().unstack(fill_value=0)
# -
data_gb_sell['price'].sum().to_sparse().unstack(level=-1)
sell_df = data_gb_sell['price'].sum().to_frame()
sell_df.describe()
sell_df['']
bins=[0,10000,100000,1000000,10000000,sell_df[sell_df['price']>0]['price'].max()]
labels = [i for i in range(len(bins)-1)]
pd.cut(sell_df[sell_df['price']>0]['price'],bins=bins,labels=labels)
pd.DataFrame(preprocessing.minmax_scale(sell_df[sell_df['price']>0],feature_range=(0, 5))).describe()
sell_df.groupby(level=0).count().iloc[12::].describe()
data_gb_sell['price'].sum().describe()
# + active=""
# print("computing Dc_v4 ......")
# Dc_v4 = data_gb_sell['price'].sum().unstack(fill_value=0)
# Dc_v4 = Dc_v4.append(data_gb_buy['price'].sum().unstack(fill_value=0))
# Dc_v4 = Dc_v4.sort_index(axis=1)
# print("computing Dc_v4 done!")
# print("flitering out general client in Dc_v4")
# Dc_v4['to_delete'] = client_to_delete_vectorize(Dc_v4.index)
# Dc_v4 = Dc_v4.loc[Dc_v4['to_delete']!='delete'].drop(['to_delete'],axis=1).copy()
# #Dc_v4 = Dc_v4[Dc_v4.sum(axis=1) > 3].copy()
# #Dc_v4.dropna(axis=1,how='all',inplace=True)
# print("all done!")
# -
from scipy.stats import mstats
data['price_winsor'] = pd.Series(mstats.winsorize(data['price'].values,limits=[0.10,0.10]))
pd.Series(preprocessing.minmax_scale(data['price_winsor'],feature_range=[0,5])).describe()
data['price_winsor']
| TopicModeling/test_compute_lda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# import relevant libraries and modules
import pandas as pd
import matplotlib.pyplot as plt
# create a pandas dataframe containing the means for five exams taken by a set of students
# save in a variable named exam_means
exam_means = pd.DataFrame(data={'exam':[1,2,3,4,5],
'mean':[73.2,71.5,79.0,62.0,84.6]})
# display exam_means
exam_means
# create a bar plot --- plot mean versus exam
plt.bar(exam_means['exam'], exam_means['mean'])
plt.xlabel('Exam')
plt.ylabel('Mean')
plt.show()
| Ex_Files_Python_Data_Functions/Exercise Files/06_03_bar_plots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Trading Lists and Trading Trajectories
#
# ### Introduction
#
# [Almgren and Chriss](https://cims.nyu.edu/~almgren/papers/optliq.pdf) provided a solution to the optimal liquidation problem by assuming the that stock prices follow a discrete arithmetic random walk, and that the permanent and temporary market impact functions are linear functions of the trading rate.
#
# Almgren and Chriss showed that for each value of risk aversion there is a unique optimal execution strategy. This optimal execution strategy is determined by a trading trajectory and its associated trading list. The optimal trading trajectory is given by:
#
# \begin{equation}
# x_j = \frac{\sinh \left( \kappa \left( T-t_j\right)\right)}{ \sinh (\kappa T)}X, \hspace{1cm}\text{ for } j=0,...,N
# \end{equation}
#
# and the associated trading list is given by:
#
# \begin{equation}
# n_j = \frac{2 \sinh \left(\frac{1}{2} \kappa \tau \right)}{ \sinh \left(\kappa T\right) } \cosh \left(\kappa \left(T - t_{j-\frac{1}{2}}\right)\right) X, \hspace{1cm}\text{ for } j=1,...,N
# \end{equation}
#
# where $t_{j-1/2} = (j-\frac{1}{2}) \tau$.
#
# Given some initial parameters, such as the number of shares, the liquidation time, the trader's risk aversion, etc..., the trading list will tell us how many shares we should sell at each trade to minimize our transaction costs.
#
# In this notebook, we will see how the trading list varies according to some initial trading parameters.
#
# ## Visualizing Trading Lists and Trading Trajectories
#
# Let's assume we have 1,000,000 shares that we wish to liquidate. In the code below, we will plot the optimal trading trajectory and its associated trading list for different trading parameters, such as trader's risk aversion, number of trades, and liquidation time.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import utils
# We set the default figure size
plt.rcParams['figure.figsize'] = [17.0, 7.0]
# Set the number of days to sell all shares (i.e. the liquidation time)
l_time = 60
# Set the number of trades
n_trades = 60
# Set the trader's risk aversion
t_risk = 1e-6
# Plot the trading list and trading trajectory. If show_trl = True, the data frame containing the values of the
# trading list and trading trajectory is printed
utils.plot_trade_list(lq_time = l_time, nm_trades = n_trades, tr_risk = t_risk, show_trl = True)
# -
# # Implementing a Trading List
#
# Once we have the trading list for a given set of initial parameters, we can actually implement it. That is, we can sell our shares in the stock market according to the trading list and see how much money we made or lost. To do this, we are going to simulate the stock market with a simple trading environment. This simulated trading environment uses the same price dynamics and market impact functions as the Almgren and Chriss model. That is, stock price movements evolve according to a discrete arithmetic random walk and the permanent and temporary market impact functions are linear functions of the trading rate. We are going to use the same environment to train our Deep Reinforcement Learning algorithm later on.
#
# We will describe the details of the trading environment in another notebook, for now we will just take a look at its default parameters. We will distinguish between financial parameters, such the annual volatility in stock price, and the parameters needed to calculate the trade list using the Almgren and Criss model, such as the trader's risk aversion.
# +
import utils
# Get the default financial and AC Model parameters
financial_params, ac_params = utils.get_env_param()
# -
# ### Default Financial Parameters
financial_params
# ### Parameters for the Almgren and Chriss Model
ac_params
# The code below implements the trading list resulting from different trading parameters, such as trader's risk aversion, number of trades, and liquidation time. All other parameters, such as total number shares to sell, are taken from the simulated trading environment (see above).
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import utils
# We set the default figure size
plt.rcParams['figure.figsize'] = [17.0, 7.0]
# Set the random seed
sd = 1
# Set the number of days to sell all shares (i.e. the liquidation time)
l_time = 60
# Set the number of trades
n_trades = 60
# Set the trader's risk aversion
t_risk = 1e-7
# Implement the trading list for the given parameters
utils.implement_trade_list(seed = sd, lq_time = l_time, nm_trades = n_trades, tr_risk = t_risk)
| finance/.ipynb_checkpoints/Trading Lists-checkpoint.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# formats: ipynb,jl:hydrogen
# text_representation:
# extension: .jl
# format_name: hydrogen
# format_version: '1.3'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.2
# language: julia
# name: julia-1.6
# ---
# %%
function repeatlength(A, i)
k = 1
@inbounds while i + k ≤ lastindex(A) && isequal(A[i+k], A[i])
k += 1
end
k
end
"""
RepVal(A)
Iterator generating the all (repeat length, value)-tuples in `A`.
"""
struct RepVal{T} A::T end
Base.IteratorSize(::Type{RepVal{T}}) where T = Base.SizeUnknown()
Base.eltype(x::RepVal) = Tuple{Int, eltype(x.A)} # (rep.len., value)
Base.iterate(x::RepVal) = iterate(x, firstindex(x.A))
function Base.iterate(x::RepVal, i::Int)
i > lastindex(x.A) && return nothing
k = repeatlength(x.A, i)
(k, x.A[i]), i + k
end
maxrep_maxval(A) = maximum(RepVal(A))
negrep((k, v)) = (-k, v)
maxrep_minval(A) = negrep(minimum(negrep, RepVal(A)))
@doc RepVal
# %%
A = [1, 2, 3, 3, 3, 1, 1, 1, NaN, NaN, NaN, 2, 2, 3]
@show A
RepVal(A) |> collect
# %%
maxrep_maxval(A)
# %%
maxrep_minval(A)
# %%
@code_warntype iterate(RepVal(A), 1)
# %% [markdown]
# https://discourse.julialang.org/t/minimum-mode-problem-minimum-number-maximum-repetition/66749/2
# %%
function findn(x,y=zeros(Int,maximum(x)))
m = 0
fill!(y,0)
@inbounds for i in x
y[i] += 1
m = max(y[i],m)
end
return findfirst(isequal(m),y)
end
# %%
# %%
using StatsBase, DataStructures
function findn_jling(x)
cm = sort!(OrderedDict(countmap(x)); byvalue = true)
last(cm.keys), last(cm.vals)
end
# %%
# %%
function minMode(c)
minVal = -1
i = 0
i = i+1
for row in eachrow(c)
row = collect(row)
temp = filter(x -> x!=0, row)
count = counter(temp)
sortedCollection = sort(collect(count), by=x->x[2], rev=true)
minVal = sortedCollection[1][1]
repetitions = sortedCollection[1][2]
for (key, value) in sortedCollection
if(value == repetitions)
if(key < minVal)
minVal = key
end
end
end
end
return minVal
end
# %%
using StatsBase
swap_negval((val, rep)) = (rep, -val)
inv_swap_negval((rep, val)) = (-val, rep)
minmode(X) = inv_swap_negval(maximum(swap_negval, countmap(X)))k
# %%
X = rand(1:10000:10^8, 10^6)
X'
# %%
using BenchmarkTools
# %%
@btime maxrep_minval($X)
# %%
@btime findn($X)
# %%
@btime minMode($X)
# %%
@btime minmode($X)
# %%
@btime findn_jling($X)
# %%
| 0017/RepVal.ipynb |