code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# +
from tensorflow.keras.preprocessing.text import Tokenizer
samples = [
"the cat sat on the mat",
"the dog ate my homework"
]
tokenizer = Tokenizer(num_words=1000)
tokenizer.fit_on_texts(samples)
tokenizer.word_index
# + tags=[]
from tensorflow.keras.datasets import imdb
from tensorflow.keras import preprocessing
max_features = 10000
max_length = 20
(X_train, Y_train), (X_test, Y_test) = imdb.load_data(num_words=max_features)
X_train = preprocessing.sequence.pad_sequences(X_train, maxlen=max_length)
X_test = preprocessing.sequence.pad_sequences(X_test, maxlen=max_length)
# -
# WORD EMBEDDINGS
# one hot vectors are wasteful of space, and can not scale to very large differences in values.
# word embeddings are associating every word with a float vector.
# This allows us to represent a very large variety of words with much less space (quasi logarithmic)
# A good word embedding should show structure shown in the words and their relationships.
#
# Example:
# If (dog, wolf, book, notes) are in the dataset, their vectors in n-D space should be representing similarity.
# dog and wolf should have less euclidean distance than suppose say dog and book.
# similarly book and notes should have less distance.
#
#
# Initially, the Embeddings are assigned randomly, and with training process, we use backpropagation to get better embeddings.
# If we are working on a document task, and we can get a good word embeddings trained on millions of sentences, we should use that.
#
# + tags=[]
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Flatten, Dense, Embedding
model = Sequential()
model.add(Embedding(max_features, 8, input_length=max_length))
model.add(Flatten())
model.add(Dense(1, activation='sigmoid'))
model.compile(
optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['acc']
)
model.summary()
# + tags=[]
history = model.fit(X_train, Y_train,
epochs=10,
batch_size=32,
validation_split=0.2
)
# -
# This can easily be increased upto around 90% accuracy by just increasing max_length = 200 words instead.
# But since this network has only one Dense neuron,
# it is essentially weighted sum of everything before.
# sum(x(i) * w(i)) does not take into account relationships between words
# +
from os.path import join as pjoin
raw = "aclImdb"
base_dir = pjoin(os.getcwd(), raw)
train_dir = pjoin(base_dir, 'train')
test_dir = pjoin(base_dir, 'test')
labels = []
texts = []
for label_type in ['neg', 'pos']:
dirname = pjoin(train_dir, label_type)
for filename in os.listdir(dirname):
if filename[-4:] == '.txt':
contents = open(pjoin(dirname, filename),encoding='utf-8').read()
texts.append(contents)
labels.append(1 if label_type == 'pos' else 0)
# + tags=[]
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
import numpy as np
maxlen = 100
training_samples = 200
validation_samples = 10000
max_words = 10000
tokenizer = Tokenizer(num_words=max_words)
# assign number to each word
tokenizer.fit_on_texts(texts)
# convert every sentence into an array of numbers
sequences = tokenizer.texts_to_sequences(texts)
print(f"Found {len(tokenizer.word_index)} words")
# -
data = pad_sequences(sequences, maxlen=maxlen)
labels = np.asarray(labels)
data.shape, labels.shape
# +
# We will train on only 200 samples, and use a pretrained word embedding to get good performance
indices = np.arange(data.shape[0])
np.random.shuffle(indices)
data = data[indices]
labels = labels[indices]
X_train = data[:training_samples]
Y_train = labels[:training_samples]
X_val = data[training_samples: training_samples + validation_samples]
Y_val = labels[training_samples: training_samples + validation_samples]
# + tags=[]
# Download the pretrained word embedding -> https://nlp.stanford.edu/projects/glove/
glove_dir = pjoin(os.getcwd(), 'glove.6B')
embeddings_index = {}
with open(pjoin(glove_dir, "glove.6B.100d.txt"), encoding='utf-8') as reader:
for line in reader:
values = line.split()
word = values[0]
coefficients = values[1:]
embeddings_index[word] = coefficients
print(f"Found {len(embeddings_index)} word vectors")
# +
word_index = tokenizer.word_index
embedding_dim = 100
embedding_matrix = np.zeros((max_words, embedding_dim))
for word, i in word_index.items():
if i < max_words:
embedding_vector = embeddings_index.get(word)
if embedding_vector is not None:
embedding_matrix[i] = embedding_vector
# + tags=[]
model = Sequential()
model.add(Embedding(max_words, embedding_dim, input_length=maxlen))
model.add(Flatten())
model.add(Dense(32, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
model.summary()
# -
model.layers[0].set_weights([embedding_matrix])
model.layers[0].trainable = False
model.compile(
optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['acc']
)
# + tags=[]
history = model.fit(
X_train, Y_train,
epochs=10,
batch_size=32,
validation_data=(X_val, Y_val)
)
# -
| books/DeepLearningWithKeras/RNN/WordEncoding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
sys.path.insert(0, '..')
from typing import Any, Dict, List
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
from matplotlib.ticker import FormatStrFormatter
from data_utils import DATAFILE_LIST, NUM_CLASSES_DICT, prepare_data, CIFAR100_CLASSES, FIGURE_DIR
from models import BetaBernoulli, ClasswiseEce
# %matplotlib inline
LINEWIDTH = 13.97
import matplotlib;matplotlib.rcParams['font.size'] = 10
import matplotlib;matplotlib.rcParams['font.family'] = 'serif'
# +
def vstripe(ax: mpl.axes.Axes,
x: np.ndarray,
labels: List[str] = None,
limit: int = None,
color: str = 'red') -> None:
num_columns = x.shape[0]
labels = labels if labels is not None else list(range(num_columns))
# Apply limit
sentinel = np.empty((1, 3))
sentinel[:] = np.nan
if limit is not None:
x = np.concatenate((x[:limit], sentinel, x[-limit:]), axis=0)
labels = labels[:limit] + ['...'] + labels[-limit:]
num_columns = x.shape[0]
# Plot
for i, row in enumerate(x):
low, mid, high = row.tolist()
ax.plot((i, i), (low, high), color = color)
ax.plot((i - .2, i + .2), (mid, mid), color = color)
# Add labels
ax.set_xlim(-1, num_columns)
ax.set_xticks(np.arange(num_columns))
ax.set_xticklabels(labels, rotation=40, fontsize=12)
ax.yaxis.set_major_formatter(FormatStrFormatter('%.1f'))
def plot_figure_1(accuracy: np.ndarray,
ece: np.ndarray,
labels: List[str] = None,
limit: int = None,
reverse: bool = False):
assert accuracy.shape == ece.shape
# Sort the datapoints and labels
sort_indices = np.argsort(accuracy[:, 1])
if reverse:
sort_indices = sort_indices[::-1]
accuracy = accuracy[sort_indices]
ece = ece[sort_indices]
if labels is not None:
labels = [labels[i] for i in sort_indices]
# Plot
fig, axes = plt.subplots(nrows=2, dpi=300, sharex=True, figsize=(LINEWIDTH*0.5, 5),
gridspec_kw = {'wspace':0.25, 'hspace':0.2})
vstripe(axes[0], accuracy, labels=labels, limit=limit, color='blue')
axes[0].set_ylim(top=1.0)
axes[0].set_ylabel('Accuracy', fontsize=12)
axes[0].tick_params(bottom=False)
# axes[1].vlines(0, -1, ece.shape[0] + 1, colors='#777777', linewidth=1, linestyle='dashed')
vstripe(axes[1], ece, labels=labels, limit=limit, color='red')
axes[1].set_ylim(bottom=0)
axes[1].set_ylabel('ECE', fontsize=12)
axes[1].set_xlabel("CIFAR-100 Labels", fontsize=14)
return fig, axes
# +
dataset = 'cifar100'
num_samples = 1000
limit=10
datafile = DATAFILE_LIST[dataset]
num_classes = NUM_CLASSES_DICT[dataset]
categories, observations, confidences, idx2category, category2idx, labels = prepare_data(datafile, False)
# accuracy models
accuracy_model = BetaBernoulli(k=num_classes, prior=None)
accuracy_model.update_batch(categories, observations)
# ece models for each class
ece_model = ClasswiseEce(num_classes, num_bins=10, pseudocount=2)
ece_model.update_batch(categories, observations, confidences)
# draw samples from posterior of classwise accuracy
accuracy_samples = accuracy_model.sample(num_samples) # (num_categories, num_samples)
ece_samples = ece_model.sample(num_samples) # (num_categories, num_samples)
accuracy = np.array([np.quantile(accuracy_samples, 0.025, axis=1),
np.quantile(accuracy_samples, 0.5, axis=1),
np.quantile(accuracy_samples, 0.975, axis=1)]).T
ece = np.array([np.quantile(ece_samples, 0.025, axis=1),
np.quantile(ece_samples, 0.5, axis=1),
np.quantile(ece_samples, 0.975, axis=1)]).T
fig, axes = plot_figure_1(accuracy, ece, labels=CIFAR100_CLASSES, limit=limit, reverse=True)
fig.tight_layout()
# -
fig.savefig(FIGURE_DIR + 'figure1.pdf', bbox_inches="tight", pad_inches=0.05)
# +
######################################CONSTANTS######################################
DEFAULT_PLOT_KWARGS = {
'color': 'blue',
'linewidth': 1
}
DEFAULT_RC = {
'font.size': 8,
'font.family': 'serif',
'font.serif': ['Times'],
# 'text.usetex': True,
'axes.titlesize': 8,
'axes.labelsize': 8,
'figure.titlesize': 8
}
COLUMN_WIDTH = 3.25 # Inches
######################################CONSTANTS######################################
import sys
sys.path.insert(0, '..')
from typing import Any, Dict, List
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
from matplotlib.ticker import FormatStrFormatter
from data_utils import DATAFILE_LIST, NUM_CLASSES_DICT, prepare_data, CIFAR100_CLASSES, FIGURE_DIR
from models import BetaBernoulli, ClasswiseEce
def hstripe(ax: mpl.axes.Axes,
x: np.ndarray,
labels: List[str] = None,
limit: int = None,
plot_kwargs: Dict[str, Any] = {}) -> None:
"""
Plots a horizontal stripe plot in the given axis.
Parameters
===
ax : matplotlib.axes.Axes
The axis to add the plot to.
x : numpy.ndarray
An array of shape (n_classes, 3) where:
x[:, 0] is the lower bounds
x[:, 1] is the midpoints
x[:, 2] is the upper bounds
labels : list
A list containing `n_classes` labels. Default: class indices are used.
limit : int
Limits the number of data points displayed; the middle data points are skipped.
Default: all data points plotted.
plot_kwargs : dict
Keyword arguments passed to the plot.
"""
num_rows = x.shape[0]
labels = labels if labels is not None else list(range(num_rows))
# Combine default and custom kwargs
# TODO: @rloganiv - find a clearner way to merge dictionaries
_plot_kwargs = DEFAULT_PLOT_KWARGS.copy()
_plot_kwargs.update(plot_kwargs)
# Apply limit
sentinel = np.empty((1, 3))
sentinel[:] = np.nan
if limit is not None:
x = np.concatenate((x[:limit], sentinel, x[-limit:]), axis=0)
labels = labels[:limit] + ['...'] + labels[-limit:]
num_rows = x.shape[0]
# Plot
for i, row in enumerate(x):
low, mid, high = row.tolist()
ax.plot((low, high), (i, i), **_plot_kwargs)
ax.plot((mid, mid), (i - .2, i + .2), **_plot_kwargs)
# Add labels
ax.set_ylim(-1, num_rows)
ax.set_yticks(np.arange(num_rows))
ax.set_yticklabels(labels)
ax.xaxis.set_major_formatter(FormatStrFormatter('%.1f'))
def plot_figure_1(accuracy: np.ndarray,
ece: np.ndarray,
labels: List[str] = None,
limit: int = None,
reverse: bool = False):
"""
Replicates Figure 1 in [CITE PAPER].
Parameters
===
accuracy : np.ndarray
An array of shape (n_classes, 3) where:
x[:, 0] is the lower bounds
x[:, 1] is the midpoints
x[:, 2] is the upper bounds
Contains the accuracy data plotted in the left plot.
ece : np.ndarray
Simlar to left, except data is plotted in the right plot.
labels : list
A list containing `n_classes` labels. Default: class indices are used.
limit : int
Limits the number of data points displayed; the middle data points are skipped.
Default: all data points plotted.
reverse : bool
Whether to reverse the vertical ordering. Default: highest value to be at top.
Returns
===
fig, axes : The generated matplotlib Figure and Axes.
"""
assert accuracy.shape == ece.shape
# Sort the datapoints and labels
sort_indices = np.argsort(accuracy[:, 1])
if reverse:
sort_indices = sort_indices[::-1]
accuracy = accuracy[sort_indices]
ece = ece[sort_indices]
if labels is not None:
labels = [labels[i] for i in sort_indices]
# Plot
with mpl.rc_context(rc=DEFAULT_RC):
fig, axes = plt.subplots(ncols=2, figsize=(3, 3), dpi=300, sharey=True)
plot_kwargs = {'color': 'tab:blue'}
hstripe(axes[0], accuracy, labels=labels, limit=limit, plot_kwargs=plot_kwargs)
axes[0].set_xlim(right=1.0)
axes[0].set_title('Accuracy')
# axes[1].vlines(0, -1, ece.shape[0] + 1, colors='#777777', linewidth=1, linestyle='dashed')
plot_kwargs = {'color': 'tab:red'}
hstripe(axes[1], ece, labels=labels, limit=limit, plot_kwargs=plot_kwargs)
axes[1].set_xlim(left=0)
axes[1].tick_params(left=False)
axes[1].set_title('ECE')
#axes[0].text(-0.5, 10, "CIFAR-100 Labels", verticalalignment='center', rotation=90)
return fig, axes
dataset = 'cifar100'
num_samples = 1000
datafile = DATAFILE_LIST[dataset]
num_classes = NUM_CLASSES_DICT[dataset]
categories, observations, confidences, idx2category, category2idx, labels = prepare_data(datafile, False)
# accuracy models
accuracy_model = BetaBernoulli(k=num_classes, prior=None)
accuracy_model.update_batch(categories, observations)
# ece models for each class
ece_model = ClasswiseEce(num_classes, num_bins=10, pseudocount=2)
ece_model.update_batch(categories, observations, confidences)
# draw samples from posterior of classwise accuracy
accuracy_samples = accuracy_model.sample(num_samples) # (num_categories, num_samples)
ece_samples = ece_model.sample(num_samples) # (num_categories, num_samples)
accuracy = np.array([np.quantile(accuracy_samples, 0.025, axis=1),
np.quantile(accuracy_samples, 0.5, axis=1),
np.quantile(accuracy_samples, 0.975, axis=1)]).T
ece = np.array([np.quantile(ece_samples, 0.025, axis=1),
np.quantile(ece_samples, 0.5, axis=1),
np.quantile(ece_samples, 0.975, axis=1)]).T
# +
fig, axes = plot_figure_1(accuracy, ece, labels=CIFAR100_CLASSES, limit=10, reverse=False)
fig.tight_layout()
fig.subplots_adjust(bottom=-0.2, wspace=0.35)
fig.set_size_inches(COLUMN_WIDTH, 1.85)
fig.savefig(FIGURE_DIR + 'figure1.pdf', bbox_inches="tight", pad_inches=0.0)
| src/plot/figure_intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.preprocessing import Imputer,LabelEncoder,OneHotEncoder
from sklearn.model_selection import train_test_split
import warnings
warnings.filterwarnings('ignore')
sns.set_style('whitegrid')
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
# +
# read-in data and merge on 'parcelid'
train = pd.read_csv('train_2016_v2.csv')
train_prop = pd.read_csv('properties_2016.csv')
df_train = pd.merge(train,train_prop,on='parcelid',how='inner')
del train,train_prop
# -
df_train.shape
# define function to reduce memory
def reduce_memory(df,verbose=True):
num_dtypes = 'int16 int32 int64 float16 float32 float64'.split()
start_mem = df.memory_usage().sum() / 1024**2
for col in df.columns:
col_type = df[col].dtypes
if col_type in num_dtypes:
col_min = df[col].min()
col_max = df[col].max()
# check if first letters start with 'flo'
if str(col_type)[:3] == 'int':
if col_min > np.iinfo(np.int8).min and col_max < np.iinfo(np.int8).max:
df[col] = df[col].astype(np.int8)
elif col_min > np.iinfo(np.int16).min and col_max < np.iinfo(np.int16).max:
df[col] = df[col].astype(np.int16)
elif col_min > np.iinfo(np.int32).min and col_max < np.iinfo(np.int32).max:
df[col] = df[col].astype(np.int32)
elif col_min > np.iinfo(np.int64).min and col_max < np.iinfo(np.int64).max:
df[col] = df[col].astype(np.int64)
else:
if col_min > np.finfo(np.float16).min and col_max < np.finfo(np.float16).max:
df[col] = df[col].astype(np.float16)
elif col_min > np.finfo(np.float32).min and col_max < np.finfo(np.float32).max:
df[col] = df[col].astype(np.float32)
else:
df[col] = df[col].astype(np.float64)
end_mem = df.memory_usage().sum() / 1024 ** 2
if verbose:
print('Memory usage decreased to {:5.2f}Mb ({:.1f}% reduction)'.format(end_mem,100*(start_mem - end_mem) / start_mem))
return df
reduce_memory(df_train)
# Missing data
# cols with more than 50 % missing values
na_count = df_train.isnull().sum().sort_values(ascending=False)
na_perc = na_count / df_train.shape[0]
na_perc = na_perc[na_perc > 0.5]
pd.DataFrame(na_perc,columns=['% missing'])
mis_col_count = len(na_perc[na_perc > 0])
print(f'There are {mis_col_count} columns with missing values.')
# Drop columns with more than 50% missing values
mis_cols = na_perc.index
df_train = df_train.drop(labels=mis_cols,axis=1)
df_train.shape
# Drop `parcelid` column
df_train = df_train.drop(labels='parcelid',axis=1)
# Get lists of numerical/categorical features with missing data.
# +
# get all the numerical columns/features
num_cols = df_train.dtypes[df_train.dtypes != 'object'].index
num_na_cols = [col for col in df_train[num_cols].columns if df_train[col].isnull().sum() > 0]
# get all the categorical features
cat_cols = df_train.dtypes[df_train.dtypes == 'object'].index
cat_na_cols = [col for col in df_train[cat_cols].columns if df_train[col].isnull().sum() > 0]
# -
# #### Imputing data
# +
# impute numerical features
impute_num = Imputer(missing_values=np.nan,strategy='mean',axis=0)
impute_num = impute_num.fit(df_train[num_na_cols])
df_train[num_na_cols] = impute_num.transform(df_train[num_na_cols])
# Manually impute categorical features
# 'propertycountylandusecode' feature
most_frequent = df_train['propertycountylandusecode'].value_counts().index[0]
df_train['propertycountylandusecode'] = df_train['propertycountylandusecode'].fillna(most_frequent)
# 'propertyzoningdesc' feature
most_frequent = df_train['propertyzoningdesc'].value_counts().index[0]
df_train['propertyzoningdesc'] = df_train['propertyzoningdesc'].fillna(most_frequent)
# -
df_train.isnull().sum()
# Now we have dealt with all the missing data.
# Drop columns: `propertycountylandusecode`,`propertylandusetypeid`, `transactiondate`
drop_cols = ['propertycountylandusecode','propertyzoningdesc','transactiondate']
df_train = df_train.drop(labels=drop_cols,axis=1)
df_train.shape
# #### Correlation heatmap of the features
plt.figure(figsize=(12,8))
sns.heatmap(df_train.corr(),cmap='coolwarm')
# Train/Test split data
indep_cols = list(df_train.columns)
indep_cols.remove('logerror')
X = df_train[indep_cols]
y = df_train['logerror']
# split data
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=12)
# Train Linear model
# import linear regression and instantiate
from sklearn.linear_model import LinearRegression
linear_model = LinearRegression()
# train/fit model
linear_model.fit(X_train,y_train)
coef = linear_model.coef_
pd.DataFrame(data=coef,columns=['coefficients'])
# Testing model/predicting data:
# predict using test data
prediction = linear_model.predict(X_test)
# Residuals
res = y_test - prediction
plt.scatter(x=prediction,y=res,alpha=0.2)
plt.xlabel('prediction')
plt.ylabel('residual');
# import metrics
from sklearn import metrics
print('MAE',metrics.mean_absolute_error(y_test,prediction))
# ## Regularization
from sklearn.linear_model import Lasso,Ridge,ElasticNet
# Lasso:
lasso = Lasso(alpha=0.01)
lasso_pred = lasso.fit(X_train,y_train).predict(X_test)
res = y_test - lasso_pred
plt.scatter(x=lasso_pred,y=res,alpha=0.2)
plt.xlabel('prediction')
plt.ylabel('residual');
# Columns that were selected out:
pd.Series(lasso.fit(X_train,y_train).coef_,index=X_train.columns)
print(metrics.mean_absolute_error(y_test,lasso_pred))
# Ridge:
ridge = Ridge(alpha=0.01)
ridge_pred = ridge.fit(X_train,y_train).predict(X_test)
res = y_test - ridge_pred
plt.scatter(x=ridge_pred,y=res,alpha=0.2)
plt.xlabel('prediction')
plt.ylabel('residual');
print(metrics.mean_absolute_error(y_test,ridge_pred))
# Elastic Net:
en = ElasticNet(alpha=0.01)
en_pred = en.fit(X_train,y_train).predict(X_test)
res = y_test - en_pred
plt.scatter(x=en_pred,y=res,alpha=0.2)
plt.xlabel('prediction')
plt.ylabel('residual');
print(metrics.mean_absolute_error(y_test,en_pred))
# Lasso performed slightly better compared to other models
| src/zillow-house-value-prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Differential Diagnosis of COVID-19 with Bayesian Belief Networks
#
# Let's see if a Bayesian Belief Network (`BBN`) is able to diagnose the COVID-19 virus with any reasonable success. The idea is that a patients presents some symptoms, and we must diagnostically reason from the `symptoms` back to the `cause`. The `BBN` is taken from [BayesiaLab's](https://www.bayesia.com/) [Differential Diagnosis](https://forum.bayesia.us/t/83hhnxy/covid-19-websimulator-for-differential-diagnosis) model.
# ## Data
#
# The data is taken from the [Hubei dataset](https://github.com/beoutbreakprepared/nCoV2019/tree/master/covid19/raw-data). We will first load both sets of data.
# +
import pandas as pd
inside = pd.read_csv('./covid/data/00/COVID19_2020_open_line_list - Hubei.csv', low_memory=False)
outside = pd.read_csv('./covid/data/00/COVID19_2020_open_line_list - outside_Hubei.csv', low_memory=False)
outside = outside.drop(['data_moderator_initials'], axis=1)
data = pd.concat([inside, outside])
# -
# ## Data Transformation
#
# We will apply transformations to the data, primarily on the symptoms. There are only about 200 unique symptoms on all the COVID-19 patients. We map these 200 unique symptoms in a many-to-many approach to 32 broad symptom categories. The following are the 32 broad symptom categories.
#
# * abdominal_pain
# * anorexia
# * anosmia
# * chest
# * chills
# * coronary
# * diarrhoea
# * digestive
# * discharge
# * dizziness
# * dry_cough
# * dryness
# * dyspnea
# * eye
# * fatigue
# * fever
# * headache
# * lungs
# * malaise
# * mild
# * muscle
# * myelofibrosis
# * nasal
# * nausea
# * respiratory
# * running_nose
# * sneezing
# * sore_throat
# * sputum
# * sweating
# * walking
# * wheezing
# +
import json
import itertools
from datetime import datetime
with open('./covid/data/00/symptom-mapping.json', 'r') as f:
symptom_map = json.load(f)
def tokenize(s):
if s is None or isinstance(s, float) or len(s) < 1 or pd.isna(s):
return None
try:
delim = ';' if ';' in s else ','
return [t.strip().lower() for t in s.split(delim) if len(t.strip()) > 0]
except:
return s
def map_to_symptoms(s):
if s.startswith('fever') or s.startswith('low fever'):
return ['fever']
return [k for k, v in symptom_map.items() if s in v]
d = data[['symptoms']].dropna(how='all').copy(deep=True)
print(d.shape)
d.symptoms = d.symptoms.apply(lambda s: tokenize(s))
d.symptoms = d.symptoms.apply(lambda tokens: [map_to_symptoms(s) for s in tokens] if tokens is not None else None)
d.symptoms = d.symptoms.apply(lambda arrs: None if arrs is None else list(itertools.chain(*arrs)))
for s in symptom_map.keys():
d[s] = d.symptoms.apply(lambda arr: 0 if arr is None else 1 if s in arr else 0)
d = d.drop(['symptoms'], axis=1)
print(d.shape)
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
plt.style.use('seaborn')
v = [d[d[c] == 1].shape[0] for c in d.columns]
s = pd.Series(v, d.columns)
fig, ax = plt.subplots(figsize=(15, 5))
_ = s.plot(kind='bar', ax=ax, title=f'Frequency of symptoms, n={d.shape[0]}')
plt.tight_layout()
# -
# ## Bayesian Belief Network
#
# The BBN structure is a result of assuming independence between the symptoms, and we know this assumption is wrong. However, we know that if we do not assume independence between the symptoms, there are more parameters to estimate and/or provide. As for the parameters, according to the original authors of this BBN, the parameters are taken from a variety of sources.
#
# The following are the variables (or nodes) in the BBN.
#
# * anosmia
# * chills
# * diarrhoea
# * dry_cough
# * dyspnea
# * fatigue
# * fever
# * headache
# * muscle
# * nasal
# * nausea
# * running_nose
# * sneezing
# * sore_throat
# * sputum
# * wheezing
#
# Note that all these nodes, except `disease` and `flu_shot` are symptoms.
# ### BBN structure
# +
from pybbn.graph.dag import Bbn
from pybbn.pptc.inferencecontroller import InferenceController
import json
with open('./covid/naive.json', 'r') as f:
bbn = Bbn.from_dict(json.load(f))
join_tree = InferenceController.apply(bbn)
# -
# The following shows the BBN structure. The `disease` node points to all the symptoms, and the `flu_shot` node points to the `disease` node. The `disease` node has the following values/states.
#
# * no_virus
# * rhinovirus
# * hmpv (Metapneumovirus)
# * hrsv (Respiratory syncytial)
# * influenza
# * covid19 (COVID-19)
# +
from pybbn.generator.bbngenerator import convert_for_drawing
import networkx as nx
import warnings
with warnings.catch_warnings():
warnings.simplefilter('ignore')
graph = convert_for_drawing(bbn)
pos = nx.nx_agraph.graphviz_layout(graph, prog='neato')
plt.figure(figsize=(15, 8))
plt.subplot(121)
labels = dict([(k, node.variable.name) for k, node in bbn.nodes.items()])
nx.draw(graph, pos=pos, with_labels=True, labels=labels)
plt.title('BBN DAG')
# -
# ### BBN Parameters
#
# The following shows the marginal posteriors of the nodes.
# +
def potential_to_series(potential):
def get_entry_kv(entry):
arr = [(k, v) for k, v in entry.entries.items()]
arr = sorted(arr, key=lambda tup: tup[0])
return arr[0][1], entry.value
tups = [get_entry_kv(e) for e in potential.entries]
return pd.Series([tup[1] for tup in tups], [tup[0] for tup in tups])
series = [(node, potential_to_series(join_tree.get_bbn_potential(node))) for node in join_tree.get_bbn_nodes()]
n_cols = 3
n_rows = int(len(series) / n_cols)
fig, axes = plt.subplots(n_rows, n_cols, figsize=(10, 20))
axes = np.ravel(axes)
for ax, (node, s) in zip(axes, series):
s.plot(kind='bar', ax=ax, title=f'{node.variable.name}')
plt.tight_layout()
# -
# ## Diagnosis
#
# Now we are ready to make diagnosis using the BBN. The total set of symptoms in the Hubei dataset (as we have transformed them) is 32, however, there are only 16 symptoms modeled into the BBN.
# +
# %%time
from pybbn.graph.jointree import EvidenceBuilder
names = [
'anosmia', 'sputum', 'muscle', 'chills', 'fever',
'wheezing', 'nasal', 'fatigue', 'headache', 'sore_throat',
'dry_cough', 'diarrhoea', 'dyspnea', 'nausea', 'sneezing',
'running_nose'
]
predictions = []
for i, r in d.iterrows():
fields = [name for name in names if r[name] == 1]
join_tree.unobserve_all()
if len(fields) > 0:
bbn_nodes = [join_tree.get_bbn_node_by_name(f) for f in fields]
evidences = [EvidenceBuilder().with_node(n).with_evidence('t', 1.0).build() for n in bbn_nodes]
join_tree.update_evidences(evidences)
disease = join_tree.get_bbn_node_by_name('disease')
disease_potential = join_tree.get_bbn_potential(disease)
s = potential_to_series(disease_potential)
predictions.append(s)
# -
predictions = pd.DataFrame(predictions)
predictions
# ## Diagnosis Performance
#
# All the records/patients in the Hubei dataset are positively-tested COVID-19 patients. Thus, we have no non-COVID-19 patients, and so we will avoid using performance measures that requires negative examples.
# ### Quasi-proper scoring rules
#
# We will try using [average precision](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.average_precision_score.html#sklearn.metrics.average_precision_score) and plot the [precision recall curve](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.precision_recall_curve.html). Note the absurdity of doing so. These performance measures are so-called `quasi-proper scoring rules`.
# +
from sklearn.metrics import average_precision_score
y_true = np.ones(predictions.shape[0])
y_pred = predictions.covid19
ap = average_precision_score(y_true, y_pred)
print(f'average precision score is {ap:.5f}')
# +
from sklearn.metrics import precision_recall_curve
pre, rec, _ = precision_recall_curve(y_true, y_pred)
fig, ax = plt.subplots(figsize=(15, 5))
_ = ax.step(rec, pre, color='b', alpha=0.5, where='post', label='PR curve')
_ = ax.set_xlabel('recall')
_ = ax.set_ylabel('precision')
_ = ax.set_title('Precision-Recall Curve')
# -
# ### Proper scoring rule
#
# Instead, we use a `proper scoring rule` such as the [Brier loss](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.brier_score_loss.html#sklearn.metrics.brier_score_loss). The Brier score is in the range $[0, 1]$, where a value closer to 0 is better. The Brier score essentially is the mean squared difference between the real probability and predicted one. As you can see, the Brier score is about 0.49. Is this value good or bad? It is right smack in the middle; meaning, it is not un-useful, but could be.
# +
from sklearn.metrics import brier_score_loss
bsl = brier_score_loss(y_true, y_pred)
print(f'brier score loss = {bsl:.5f}')
# -
# ### Agreement
#
# Here, we take a different approach to judging the BBN's diagnostic reliability by looking at the counts of predicted patients to have COVID-19 versus the empirical counts.
#
# * First, we create strata based on the observed and unique combinations of symptoms and observe the empirical number of patients with such co-symptoms.
# * Second, for each unique combination of symptoms observed, we present such symptoms as evidence to the model and allow it to give us the probability of having COVID-19.
# * Third, we multiply the probability by the total number of patients observed across all the strata.
# * Lastly, we compare the `agreement` between the numbers predicted by the BBN and the empirical ones.
# +
def get_symptom_combinations(r):
fields = sorted([name for name in names if r[name] == 1])
return fields
def get_query(combination):
p_tokens = combination.split(',')
n_tokens = [n for n in names if n not in p_tokens]
p_tokens = [f'{t}==1' for t in p_tokens]
n_tokens = [f'{t}==0' for t in n_tokens]
tokens = p_tokens + n_tokens
query = ' and '.join(tokens)
return query
combinations = [get_symptom_combinations(r) for _, r in d.iterrows()]
combinations = [c for c in combinations if len(c) > 0]
combinations = [','.join(c) for c in combinations]
combinations = sorted(list(set(combinations)))
print(f'number of combinations {len(combinations)}')
queries = [get_query(c) for c in combinations]
# we lose 67 patients, they have no symptoms
strata = pd.DataFrame([(c, d.query(q).shape[0]) for c, q in zip(combinations, queries)], columns=['stratum', 'n'])
strata['n_symptoms'] = strata.stratum.apply(lambda s: len(s.split(',')))
print(f'number of patients {strata.n.sum()}')
# -
# This is the distribution of the unique combinations of co-symptoms. Note that some symptoms may show up only by themselves.
fig, ax = plt.subplots(figsize=(20, 5))
s = pd.Series(strata.n.values, strata.stratum.values)
_ = s.plot(kind='bar', ax=ax, title=f'Frequency of all symptom combinations, n={strata.n.sum()}')
# In this graph, we remove strata that have only 1 symptom to remove the effect of visual skewness.
# +
s = strata[strata.n_symptoms > 1]
fig, ax = plt.subplots(figsize=(20, 5))
s = pd.Series(s.n.values, s.stratum.values)
_ = s.plot(kind='bar', ax=ax, title=f'Frequency of symptom combinations (more than 1), n={strata.n.sum()}')
# -
# Now we feed the symptoms in each of the stratum to the BBN and estimate the predicted counts of patients with COVID-19.
# +
import math
predictions = []
for i, r in strata.iterrows():
fields = r.stratum.split(',')
join_tree.unobserve_all()
if len(fields) > 0:
bbn_nodes = [join_tree.get_bbn_node_by_name(f) for f in fields]
evidences = [EvidenceBuilder().with_node(n).with_evidence('t', 1.0).build() for n in bbn_nodes]
join_tree.update_evidences(evidences)
disease = join_tree.get_bbn_node_by_name('disease')
disease_potential = join_tree.get_bbn_potential(disease)
s = potential_to_series(disease_potential)
predictions.append(s)
predictions = pd.DataFrame(predictions)
n = strata.n.sum()
preds = pd.DataFrame([(math.ceil(n * p), c) for p, c in zip(predictions.covid19, strata.n)], columns=['y_pred', 'y_true'])
# -
# Below, we visualize the predicted number of patients with COVID-19 given multiple symptoms with the model versus the empirical numbers. We use [Pearson](https://docs.scipy.org/doc/scipy-0.14.0/reference/generated/scipy.stats.pearsonr.html), [Kendall](https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.kendalltau.html), and [Spearman](https://docs.scipy.org/doc/scipy-0.14.0/reference/generated/scipy.stats.spearmanr.html) correlations. The latter two correlation measures are rank correlations and may be used to gauge at the agreement between the ranked predicted and empirical frequencies/counts. For all these correlation measures, the higher the value, the better the agreement. As can be seen below, there is positive agreement, and in some sense, especially with `Spearman correlation`, the agreement is strong.
#
# Let's note that the few dots to the right correspond to stratum with a single symptom. This observation is not surprising, since the BBN assumes independence between the symptoms; meaning, we should expect agreement between the predicted and empirical counts when it comes to stratum with one symptom.
# +
from scipy.stats import spearmanr, kendalltau, pearsonr
spearman = spearmanr(preds.y_true, preds.y_pred).correlation
kendall = kendalltau(preds.y_true, preds.y_pred).correlation
pearson = pearsonr(preds.y_true, preds.y_pred)[0]
fig, ax = plt.subplots(figsize=(10, 5))
_ = ax.scatter(preds.y_true, preds.y_pred)
_ = ax.set_title(f'Counts of patients predicted to have COVID-19 vs empirical counts\npearson={pearson:.2f}, spearman={spearman:.2f}, kendall={kendall:.2f}')
_ = ax.set_xlabel('empirical counts')
_ = ax.set_ylabel('predicted counts')
# +
x = preds / preds.sum()
spearman = spearmanr(x.y_true, x.y_pred).correlation
kendall = kendalltau(x.y_true, x.y_pred).correlation
pearson = pearsonr(x.y_true, x.y_pred)[0]
fig, ax = plt.subplots(figsize=(10, 5))
_ = ax.scatter(x.y_true, x.y_pred)
_ = ax.set_title(f'Probabilities of patients predicted to have COVID-19 vs empirical counts\npearson={pearson:.2f}, spearman={spearman:.2f}, kendall={kendall:.2f}')
_ = ax.set_xlabel('empirical probability')
_ = ax.set_ylabel('predicted probability')
# -
# Here is the mean squared difference between the predicted probabilities (of frequencies) and the empirical ones. Wow! Almost zero!
x.apply(lambda r: (r.y_pred - r.y_true)**2, axis=1).mean()
# Here is the Brier score for the predicted probabilities. Remember, Brier loss ranges from $[0, 1]$ and the lower the Brier loss, the better. This approach of judging the BBN means that the model is very bad at diagnosing COVID-19.
brier_score_loss(np.ones(x.shape[0]), x.y_pred)
# Here is the Brier score for the empirical probabilities. Whew! These two last results suggest maybe this way of judging the BBN is not correct.
brier_score_loss(np.ones(x.shape[0]), x.y_true)
# ## Misc
#
# Ignore the code below. It will print out all the unique symptoms in the Hubei data. Useful for the symptom mapping exercise.
# +
# x = [tokenize(s) for s in data.symptoms if s is not None]
# x = [tokens for tokens in x if tokens is not None and len(tokens) > 0]
# n = len(x)
# x = list(itertools.chain(*[item for item in x]))
# for i, s in enumerate(sorted(list(set(x)))):
# print(f'{i}, {s}')
| sphinx/datascience/source/covid-diagnosis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.12 64-bit (''main'': conda)'
# language: python
# name: python3
# ---
# ## Problem Statement
# > Jill designs solar panels as a hobby.
#
# > On `April 1st`, Jill's `Mark I` design begins generating power: `1Kj/day`.
#
# > On `May 1st`, her `Mark II` design begins generating power: `4Kj` of power per day.
#
# ## Questions
# 1. What day is it when Jill's `Mark II` design has generated as much total energy as the `Mark I` design?
# 1. How much total energy have both generated by that day?
# 1. What would the solutions `1.` and `2.` be if `Mark II` design generated `1Kj` of power per day?
# ## Solutions
#
# ### 1.
# > equation1: e = 1t
#
# > equation2: e = 4(t - 30)
#
# > => 4(t - 30) = 1t
#
# > => 4t - 120 = t
#
# > => 4t - t = 120
#
# > => 3t = 120
#
# > => t = 120/3
#
# > => t = 40 i.e., `May 10th`
#
#
# ### 2.
# > * equation1: e = 1t => 1(40) => `40Kj`
#
# > * equation2: e = 4(t - 30)
# > * => 4(40 - 30)
# > * => 160 - 120
# > * => `40Kj`
#
# > i.e., total `80Kj`
#
#
# ### 3.
# > No Solution
#
| Personal/linear-algebra.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# #!/usr/bin/env python3
# -*- coding: utf-8 -*-
"""
Created on Thu Jun 24 17:17:54 2021
@author: <NAME>
"""
# install the libraries
import sys
# !{sys.executable} -m pip install numpy
# !{sys.executable} -m pip install matplotlib
# !{sys.executable} -m pip install pandas
# !{sys.executable} -m pip install statsmodels
# !{sys.executable} -m pip install sklearn
# import the libraries
import numpy as np
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
import statsmodels.api as sm
from statsmodels.tsa.stattools import adfuller
from statsmodels.tsa.api import VAR
from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
from sklearn.metrics import mean_squared_error
from sklearn.metrics import mean_absolute_error
from sklearn.metrics import r2_score
# +
color_list = ["orange", "green"]
# function to show plot
def Visualize(data):
features = list(df.select_dtypes(include=[np.number]).columns.values)
feature_size = len(features)
fig, axes = plt.subplots(
nrows = int(np.ceil(feature_size/2)),
ncols = 2, figsize = (14, feature_size * 2),
dpi = 150,
facecolor = "w",
edgecolor = "k"
)
for i in range(feature_size):
key = features[i]
c = color_list[i % (len(color_list))]
t_data = data[key]
t_data.head()
ax = t_data.plot(
ax = axes[i % 2],
color = c ,
title = "{}".format(key),
rot = 25
)
ax.legend([key])
plt.tight_layout()
# +
# MAPE
def mean_absolute_percentage_error(y_true, y_pred):
return (np.mean(np.abs((y_pred - y_true) / y_true)) * 100) / 100
# SMAPE
def symmetric_mean_absolute_percentage_error(y_true, y_pred):
return (2.0 * np.mean(np.abs(y_pred - y_true) / (np.abs(y_pred) + np.abs(y_true))) * 100) / 100
# -
# import dataset
df = pd.read_csv('dataset_tondano_winangun.csv', index_col=0, parse_dates=True)
df
Visualize(df)
plt.savefig('data_preprocessing.png')
# +
# cleaning dataset
# change any zero value to NaN and fill NaN with mean value from dataframe
df=df.mask(df==0).fillna(df.mean())
#df = df.dropna() # remove empty Value
#df = df.fillna(0.1) # change NaN to 0.1
#df = df[(df.T != 0).any()] # remove all zero value
# show output
df
# -
Visualize(df)
plt.savefig('data_postprocessing.png')
# ACF Tondano and Winangun
acf_tondano = plot_acf(df['tondano']).legend(['Tondano'])
acf_winangun = plot_acf(df['winangun']).legend(['Winangun'])
plt.show()
# PACF Tondano and Winangun
pacf_tondano = plot_pacf(df['tondano']).legend(['Tondano'])
pacf_winangun = plot_pacf(df['winangun']).legend(['Winangun'])
plt.show()
# check is the series stationary or not using ADF
for i in range(len(df.columns)):
result = adfuller(df[df.columns[i]])
print(f"Test Statistics: {result[0]}")
print(f"P-Value: {result[1]}")
print(f"Critical Values: {result[4]}")
if result[1] > 0.05:
print("{}: Series is not Stationary\n".format(df.columns[i]))
else:
print("{}: Series is Stationary\n".format(df.columns[i]))
# make train data and test data
df_train = df[:int(0.8*(len(df)))]
df_test = df[int(0.8*(len(df))):]
# show dataframe for training
df_train
# show dataframe for test
df_test
# try 10 lags
lags = 10
model = VAR(df_train, freq="D")
for i in range(lags):
results = model.fit(i+1)
print("Order = ", i+1)
print("AIC = ", results.aic)
print("BIC = ", results.bic)
model.select_order(lags).summary()
# from the result above, the lowest BIC is from the lag 1
model = VAR(df_train, freq="D")
results = model.fit(7)
results.summary()
# using lag 6
lag = results.k_ar
print("Lag:",lag)
# +
# result for test and predict
y_test = df_test
y_predict = results.forecast(df_train.values[-lag:], steps=df_test.shape[0])
mape = mean_absolute_percentage_error(y_test, y_predict)
print("MAPE:",mape,"\n")
smape = symmetric_mean_absolute_percentage_error(y_test, y_predict)
print("SMAPE:",smape,"\n")
mae = mean_absolute_error(y_test,y_predict)
print("MAE :",mae)
mse = mean_squared_error(y_test,y_predict)
print("MSE :",mse)
rmse = np.sqrt(mean_squared_error(y_test,y_predict))
print("RMSE:",rmse)
print("R2 :",r2_score(y_test,y_predict))
# -
# generate forecast for next 10 days
data = np.array(results.forecast(df_train.values[-lag:], steps=10))
forecast_output = pd.DataFrame(data=data, columns=['tondano', 'winangun'])
forecast_output['days'] = pd.DataFrame(data=[1,2,3,4,5,6,7,8,9,10])
forecast_output = forecast_output[['days', 'tondano', 'winangun']]
print("10 Days Forecasts:\n=============================")
print(forecast_output.to_string(index=False))
Visualize(forecast_output)
plt.savefig('forecast_output.png')
# write forecast output to csv
forecast_output.to_csv(r'forecast_output_var.csv', index = False)
| model_var.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: .NET (C#)
// language: C#
// name: .net-csharp
// ---
// + dotnet_interactive={"language": "csharp"}
// Use the Sylvester abstract algebra package
//#load "Paket.fsx"
//Paket.Package["Sylvester.AbstractAlgebra"]
//#load "Paket.Generated.Refs.fsx"
#load "C:\\Projects\\Sylvester.git\\notebooks\\MathInclude.fsx"
// -
// # Sylvester.AbstractAlgebra
// The Sylvester abstract algebra library contains types and operations for rigorously defining abstract algebra structures and concepts. The F# language is both functional and object-oriented with infinite sequences as first-class objects, which make it really great at representing mathematical structures like groups which have natural inheritance relations, as well as infinite sequences of objects like the natural numbers and other enumerable sets. The language features of F# together with Sylvester types allow the user to write mathematical expressions that are very close to what you will find in print.
// ## Morphisms
// + dotnet_interactive={"language": "fsharp"}
open System
open System.Linq
open Sylvester
open Sylvester.Arithmetic
// + dotnet_interactive={"language": "fsharp"}
// Define a custom symbol type S with a (+) operator and zero
// We could just also use plain strings
type S = S of string with
static member (+) (S l, S r) = S (l + r)
static member Zero = S ""
// Define an infinite sequence of S strings
let Sym = infiniteSeq ofType<S> ((+) 65 >> Char.ConvertFromUtf32 >> S)
Sym
// + dotnet_interactive={"language": "fsharp"}
// Define a monoid using our set and + operator and zero element
let L = Monoid(Sym, (+), S.Zero)
L
// + dotnet_interactive={"language": "csharp"}
// Create 2 S values
let a, b = S "Nancy", S "Drew"
a + b
// + dotnet_interactive={"language": "csharp"}
// Create an L morphism using the PadLeft string function
let Pad = Morph(L, fun l -> let (S s) = l in S(s.PadLeft 20))
let pad = Pad.Map
pad a
// + dotnet_interactive={"language": "csharp"}
// Is pad a homomorphism?
pad a + pad b
// + dotnet_interactive={"language": "csharp"}
pad (a + b)
// + dotnet_interactive={"language": "csharp"}
pad a + pad b = pad (a + b)
// + dotnet_interactive={"language": "csharp"}
let s1 = seq{1..6} |> Seq
let s2 = Seq [5]
let c = s1 |-| s2
c.Powerset.Length
// + dotnet_interactive={"language": "csharp"}
Nz
// -
// ## Lattices
// + dotnet_interactive={"language": "csharp"}
let c1 = infiniteSeq (fun x -> x % 2 = 0) (fun n -> n * 2)
c1
// + dotnet_interactive={"language": "csharp"}
let dice = seq {1..6} |> Seq
let outcomes = dice * dice
outcomes
// + dotnet_interactive={"language": "csharp"}
dice.Powerset
// + dotnet_interactive={"language": "csharp"}
let setal = SetAlgebra(dice)
setal.Set.Length
// + dotnet_interactive={"language": "csharp"}
Z.Set.Prod
// + dotnet_interactive={"language": "csharp"}
Z.Set.Prod |>| fun (x,y) -> x > 16 && x % 2 = 0 && y % 3 = 0
// + dotnet_interactive={"language": "csharp"}
| examples/math/Abstract Algebra.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import shutil
import random
from tqdm import tqdm
DATA_DIR = "/home/jarvis/ML/asl-alphabet"
TRAIN_DIR = os.path.join(DATA_DIR,'asl_alphabet_train')
VAL_DIR = os.path.join(DATA_DIR,'val_images')
NTRAIN_DIR = os.path.join(DATA_DIR,'train_images')
print(os.listdir(TRAIN_DIR))
os.mkdir(VAL_DIR)
os.mkdir(NTRAIN_DIR)
VAL_SIZE = 500
for subd in os.listdir(TRAIN_DIR):
src = os.path.join(TRAIN_DIR,subd)
files = os.listdir(src)
random.shuffle(files)
dest = os.path.join(VAL_DIR,subd)
os.mkdir(dest)
for img in tqdm(files[:VAL_SIZE]):
shutil.copyfile(os.path.join(src,img),os.path.join(dest,img))
dest = os.path.join(NTRAIN_DIR,subd)
os.mkdir(dest)
for img in tqdm(files[VAL_SIZE:]):
shutil.copyfile(os.path.join(src,img),os.path.join(dest,img))
| model-preparation/sep_train_val.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_mxnet_p36
# language: python
# name: conda_mxnet_p36
# ---
# + slideshow={"slide_type": "skip"}
from utils import *
# %matplotlib inline
# + [markdown] slideshow={"slide_type": "slide"}
# # Automatic differentiation with `autograd`
# + [markdown] slideshow={"slide_type": "notes"}
# We train models to get better and better as a function of experience. Usually, getting better means minimizing a loss function. To achieve this goal, we often iteratively compute the gradient of the loss with respect to weights and then update the weights accordingly. While the gradient calculations are straightforward through a chain rule, for complex models, working it out by hand can be a pain.
#
# Before diving deep into the model training, let's go through how MXNet’s `autograd` package expedites this work by automatically calculating derivatives.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Basic usage
# + [markdown] slideshow={"slide_type": "notes"}
# Let's first import the `autograd` package.
# + slideshow={"slide_type": "fragment"}
import mxnet as mx
from mxnet import nd
from mxnet import autograd
# + [markdown] slideshow={"slide_type": "notes"}
# As a toy example, let’s say that we are interested in differentiating a function $f(x) = 2 x^2$ with respect to parameter $x$. We can start by assigning an initial value of $x$.
# + [markdown] slideshow={"slide_type": "slide"}
# differentiate
#
# $f(x) = 2 x^2$
#
# with respect to parameter $x$.
# + attributes={"classes": [], "id": "", "n": "3"} slideshow={"slide_type": "fragment"}
x = nd.array([[1, 2], [3, 4]])
x
# + [markdown] slideshow={"slide_type": "notes"}
# Once we compute the gradient of $f(x)$ with respect to $x$, we’ll need a place to store it. In MXNet, we can tell an NDArray that we plan to store a gradient by invoking its `attach_grad` method.
# + attributes={"classes": [], "id": "", "n": "6"} slideshow={"slide_type": "slide"}
x.attach_grad()
# + [markdown] slideshow={"slide_type": "notes"}
# Now we’re going to define the function $y=f(x)$.
#
# To let MXNet store $y$, so that we can compute gradients later, we need to put the definition inside a `autograd.record()` scope.
# + [markdown] slideshow={"slide_type": "slide"}
# $y=f(x)$
# + slideshow={"slide_type": "fragment"}
def f(x):
return 2 * x**2
# + attributes={"classes": [], "id": "", "n": "7"} slideshow={"slide_type": "fragment"}
with autograd.record():
y = f(x)
# + slideshow={"slide_type": "fragment"}
x, y
# + [markdown] slideshow={"slide_type": "notes"}
# Let’s invoke back propagation (backprop) by calling `y.backward()`. When $y$ has more than one entry, `y.backward()` is equivalent to `y.sum().backward()`.
# <!-- I'm not sure what this second part really means. I don't have enough context. TMI?-->
# + [markdown] slideshow={"slide_type": "slide"}
# Backward propagation of y
# + attributes={"classes": [], "id": "", "n": "8"} slideshow={"slide_type": "fragment"}
y.backward()
# + [markdown] slideshow={"slide_type": "notes"}
# Now, let’s see if this is the expected output. Note that $y=2x^2$ and $\frac{dy}{dx} = 4x$, which should be `[[4, 8],[12, 16]]`. Let's check the automatically computed results:
# + [markdown] slideshow={"slide_type": "fragment"}
# $y=2x^2$
#
# $\frac{dy}{dx} = 4x$
# + attributes={"classes": [], "id": "", "n": "9"} slideshow={"slide_type": "fragment"}
x, x.grad
# + [markdown] slideshow={"slide_type": "slide"}
# ## Using Python control flows
# + [markdown] slideshow={"slide_type": "notes"}
# Sometimes we want to write dynamic programs where the execution depends on some real-time values. MXNet will record the execution trace and compute the gradient as well.
#
# Consider the following function `f`: it doubles the inputs until it's `norm` reaches 1000. Then it selects one element depending on the sum of its elements.
# <!-- I wonder if there could be another less "mathy" demo of this -->
# + [markdown] slideshow={"slide_type": "fragment"}
# $Y=f(X)$
# - Take a vector `X` of two random numbers in [-1, 1]
# - `X` is multiplied by `2` until its norm reach `1000`
# - If `X`'s sum is positive, return 1st element
# - Otherwise 2nd
# + slideshow={"slide_type": "slide"}
def f(x):
x = x * 2
while x.norm().asscalar() < 1000:
x = x * 2
# If sum positive
# pick 1st
if x.sum().asscalar() >= 0:
y = x[0]
# else pick 2nd
else:
y = x[1]
return y
# + [markdown] slideshow={"slide_type": "notes"}
# We record the trace and feed in a random value:
# + slideshow={"slide_type": "slide"}
x = nd.random.uniform(-1, 1, shape=2)
x
# + slideshow={"slide_type": "fragment"}
x.attach_grad()
with autograd.record():
y = f(x)
y.backward()
# + [markdown] slideshow={"slide_type": "notes"}
# We know that `y` is a linear function of `x`, and `y` is chosen from `x`. Then the gradient with respect to `x` be will be either `[y/x[0], 0]` or `[0, y/x[1]]`, depending on which element from `x` we picked. Let's find the results:
# + [markdown] slideshow={"slide_type": "slide"}
# $y=k.x[0]$
#
# or
#
# $y=k.x[1]$,
#
# hence $\frac{dy}{dx} = \begin{vmatrix} 0 \\ k \end{vmatrix} $ or $ \begin{vmatrix} k \\ 0 \end{vmatrix}$
#
# with $k = 2^n$ where n is the number of times $x$ was multiplied by 2
# + slideshow={"slide_type": "fragment"}
x
# + slideshow={"slide_type": "fragment"}
x.grad
| part1_basics_mlp_cnn/lab/2_autograd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/srijan-singh/machine-learning/blob/main/ML%20Libraries/Metplotlib.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="yqFfk9Rrkcya"
import matplotlib.pyplot as plt
# + id="yUNO8p3jkrrF"
x = [i for i in range(10)]
# + id="HFl4BrYIk5eK" outputId="a9c8e7a2-b277-445f-9aab-f20f63fb1b6c" colab={"base_uri": "https://localhost:8080/", "height": 35}
print(x)
# + id="HSZ6PpH7k88O"
y = [2*i for i in range(10)]
# + id="kMYfWLRdlDC8" outputId="54b40495-49d1-41c5-b175-5fd2453b05b4" colab={"base_uri": "https://localhost:8080/", "height": 35}
print(y)
# + id="8FtbxTlVlbfG" outputId="2dd329dd-ec92-4326-8e2f-4a540c9a3cd7" colab={"base_uri": "https://localhost:8080/", "height": 301}
plt.xlabel("x-axis")
plt.ylabel("y-axis")
# + id="BDp2JfEOmEzI" outputId="568dbbd3-41a0-4072-ec42-8fcebe933e05" colab={"base_uri": "https://localhost:8080/", "height": 283}
plt.plot(x, y)
# + id="Y_cenhx7lFTP" outputId="e04fb396-eccf-448a-a359-f312fdc6a829" colab={"base_uri": "https://localhost:8080/", "height": 297}
plt.xlabel("x-axis")
plt.ylabel("y-axis")
plt.plot(x, y)
# + id="7cOVrjWImMSu" outputId="c1d761c2-c418-4540-d78d-c79dff962cd5" colab={"base_uri": "https://localhost:8080/", "height": 283}
plt.scatter(x, y)
| ML Libraries/Metplotlib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Line
#
# Let us draw a trajectory of `N` steps of an approximation to the [Wiener Process](https://en.wikipedia.org/wiki/Wiener_process) in three dimensions.
#
# Below, a blue thin line is a trajectory and the total displacement is shown with red thick line.
#
# Line takes data as an array of coordinates `[number_of_points,3]`.
import numpy as np
import k3d
plot = k3d.plot(camera_auto_fit=False)
N = 10000
traj = np.cumsum(np.random.randn(N,3).astype(np.float32),axis=0)
plt_line = k3d.line(traj, shader='mesh', width=0.5)
plt_line2 = k3d.line([traj[0],traj[-1]],shader='mesh', width=2.5, color=0xff0000)
plot += plt_line
plot += plt_line2
plot.display()
# The update of vertices can be easily done by updating attribute of an object. For example we can generate other realization of the Wiener process:
plot.camera_auto_fit = False
plot.grid_auto_fit = False
import time
for i in range(40):
traj = np.cumsum(np.random.randn(N,3).astype(np.float32),axis=0)
plt_line.vertices = traj
plt_line2.vertices = [traj[0],traj[-1]]
time.sleep(1)
| docs/source/basic_functionality/Line.ipynb |
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: spylon-kernel
// language: scala
// name: spylon-kernel
// ---
import org.apache.spark.SparkConf
import org.apache.spark.sql.{DataFrame, SparkSession}
import org.apache.spark.sql.functions.{datediff, hour, to_date, udf,lower,concat,lit, from_unixtime,format_number,when}
val df : DataFrame = spark.read
.option("header",true)
.option("inferSchema","true")
.csv("data/train_clean.csv")
// +
print(s"Nombre de ligne : ${df.count}")
println(s"Nombre de colonnes : ${df.columns.length}")
//df.show()
//df.printSchema()
val dfCasted : DataFrame = df
.withColumn("goal", df("goal").cast("Int"))
.withColumn("deadline" , df("deadline").cast("Int"))
.withColumn("state_changed_at", df("state_changed_at").cast("Int"))
.withColumn("created_at", df("created_at").cast("Int"))
.withColumn("launched_at", df("launched_at").cast("Int"))
.withColumn("backers_count", df("backers_count").cast("Int"))
.withColumn("final_status", df("final_status").cast("Int"))
// -
dfCasted
.select("goal", "backers_count", "final_status")
.describe()
.show
/*
dfCasted.groupBy("disable_communication").count.orderBy($"count".desc).show(20)
dfCasted.groupBy("country").count.orderBy($"count".desc).show(20)
dfCasted.groupBy("currency").count.orderBy($"count".desc).show(20)
dfCasted.select("deadline").dropDuplicates.show()
dfCasted.groupBy("state_changed_at").count.orderBy($"count".desc).show(20)
dfCasted.groupBy("backers_count").count.orderBy($"count".desc).show(20)
dfCasted.select("goal", "final_status").show(30)
dfCasted.groupBy("country", "currency").count.orderBy($"count".desc).show(50)
*/
val df2: DataFrame = dfCasted.drop("disable_communication")
val dfNoFutur : DataFrame = df2.drop("backers_count","state_changed_at")
df.filter($"country" === "False")
.groupBy("currency")
.count
.orderBy($"count".desc)
.show(50)
def cleanCountry(country: String, currency: String): String = {
if (country == "False" )
currency
else
country
}
def cleanCurrency(currency: String): String = {
if (currency != null && currency.length != 3)
null
else
currency
}
val cleanCountryUdf = udf(cleanCountry _)
val cleanCurrencyUdf = udf(cleanCurrency _)
val dfCountry: DataFrame = dfNoFutur
.withColumn("country2", cleanCountryUdf($"country", $"currency"))
.withColumn("currency2", cleanCurrencyUdf($"currency"))
.drop("country", "currency")
//dfCountry.groupBy("final_status").count.orderBy($"count".desc).show()
// Ici nous allons séléctionner que les campagens ayant un final-status à 0 ou 1.
// On pourrait toutefois tester en mettant toutes les autres valeurs à 0
// en considérant que les campagnes qui ne sont pas un Success sont un Fail.
val dfFinalStatus : DataFrame = dfCountry
.withColumn("final_status", when($"final_status"===0 || $"final_status"===1,$"final_status").otherwise(null))
.filter($"final_status".isNotNull)
//dfFinalStatus.groupBy("final_status").count.orderBy($"count".desc).show()
//dfFinalStatus.printSchema()
// dfFinalStatus.show()
val dfNbDays : DataFrame = dfFinalStatus
.withColumn("deadline2",from_unixtime($"deadline"))
.withColumn("launched_at2",from_unixtime($"launched_at"))
.withColumn("created_at2",from_unixtime($"created_at"))
.withColumn("days_campaign", datediff($"deadline2",$"launched_at2"))
.withColumn("hours_prepa", format_number(($"launched_at" - $"created_at")/3600,3))
.drop("launched_at","created_at","deadline","launched_at2","created_at2","deadline2")
//dfNbDays.show()
val dfText : DataFrame = dfNbDays
.withColumn("desc", lower($"desc"))
.withColumn("name", lower($"name"))
.withColumn("keywords", lower($"keywords"))
.withColumn("text",concat($"name",lit(" "),$"desc",lit(" "),$"keywords"))
.drop("name","desc","keywords")
//dfText.show()
//val cleanNullIntUdf = udf(cleanNullInt _)
//val cleanNullStringUdf = udf(cleanNullString _)
val dfCleanNull : DataFrame = dfText
.withColumn("days_campaign",when($"days_campaign".isNull,-1).otherwise($"days_campaign"))
.withColumn("goal",when($"goal"isNull, -1).otherwise($"goal"))
.withColumn("hours_prepa",when($"hours_prepa"isNull,-1).otherwise($"hours_prepa"))
.withColumn("country2",when($"country2"==="True","unknown").otherwise($"country2"))
.withColumn("currency2",when($"currency2".isNull,"unknown").otherwise($"currency2"))
// +
//Environ 22000 ligne avec des hours_prepa negatifs et que 37 en dessous de -10 370 en dessous de -5 => solution de les mettre tous à 0
print("preclean",dfCleanNull.where(($"hours_prepa" <0) ).count())
val dfCleaned : DataFrame = dfCleanNull
.withColumn("hours_prepa", when($"hours_prepa" < 0, -1).otherwise($"hours_prepa"))
print("ceci est un test post clean ",dfCleaned.filter(($"hours_prepa" <0) ).count())
// -
dfCleaned.where($"goal" < 0).count()
dfCleaned.where($"goal".isNull).count()
dfCleaned.where($"days_campaign" < 0).count()
dfCleaned.where($"hours_prepa".isNull).count()
// # TP 3
// ## Utilisation des données textuelles
import org.apache.spark.ml.feature.{IDF, Tokenizer, RegexTokenizer, StopWordsRemover, CountVectorizer, StringIndexer, OneHotEncoder, VectorAssembler}
import org.apache.spark.ml.{Pipeline, PipelineModel}
import org.apache.spark.ml.classification.LogisticRegression
val df : DataFrame = spark.read
.parquet("data/prepared_trainingset")
df.show()
df.select($"text".isNull).groupBy($"(text IS NULL)").count().show()
val tokenizer = new RegexTokenizer()
.setPattern("\\W+")
.setGaps(true)
.setInputCol("text")
.setOutputCol("tokens")
val stopWordsRemover = new StopWordsRemover()
.setInputCol("tokens")
.setOutputCol("tokensWOstopwords")
val cvModel = new CountVectorizer()
.setInputCol("tokensWOstopwords")
.setOutputCol("countedWord")
.setMinDF(2) //a word has to appear 2 times to be in the vocabulary
.fit(stopWordsRemover.transform(tokenizer.transform(df)))
val idf = new IDF()
.setInputCol("countedWord")
.setOutputCol("tfidf")
// ## Conversion des variables catégorielles en variables numériques
val indexerCountry = new StringIndexer()
.setInputCol("country2")
.setOutputCol("country_indexed")
val indexerCurrency = new StringIndexer()
.setInputCol("currency2")
.setOutputCol("currency_indexed")
val onehotencoderCountry = new OneHotEncoder()
.setInputCol("country_indexed")
.setOutputCol("country_onehot")
val onehotencoderCurrency = new OneHotEncoder()
.setInputCol("currency_indexed")
.setOutputCol("currency_onehot")
val assembler = new VectorAssembler()
.setInputCols(Array("tfidf", "days_campaign", "hours_prepa", "goal", "country_onehot", "currency_onehot"))
.setOutputCol("features")
val lr = new LogisticRegression()
.setElasticNetParam(0.0)
.setFitIntercept(true)
.setFeaturesCol("features")
.setLabelCol("final_status")
.setStandardization(true)
.setPredictionCol("predictions")
.setRawPredictionCol("raw_predictions")
.setThresholds(Array(0.7, 0.3))
.setTol(1.0e-6)
.setMaxIter(20)
val splits = df.randomSplit(Array(0.9, 0.1))
val training = splits(0).cache()
val test = splits(1)
val pipeline = new Pipeline()
.setStages(Array(tokenizer, stopWordsRemover,cvModel,idf, indexerCountry,indexerCurrency,
onehotencoderCountry, onehotencoderCurrency, assembler, lr))
val model = pipeline.fit(training)
df.show()
val predic = model.transform(test)
// + active=""
// predic.select("features","raw_predictions","probability","predictions","final_status").show()
// -
import org.apache.spark.ml.evaluation.MulticlassClassificationEvaluator
val evaluator = new MulticlassClassificationEvaluator()
.setLabelCol("final_status")
.setPredictionCol("predictions")
.setMetricName("f1")
val result = evaluator.evaluate(predic)
val sameModel = PipelineModel.load("spark-logistic-regression-model")
// +
val paramGrid = new ParamGridBuilder()
.addGrid(cvModel.minDF, Array(55.0,75.0,95.0))
.addGrid(lr.regParam, Array(10e-8, 10e-6, 10e-4, 10e-2))
.build()
val lrtv = TrainValidationSplit()
.setEstimator(pipeline)
.setEstimatorParamMaps(paramGrid)
.setEvaluator(evaluator)
val modelGrid = lrtv.fit(training)
// -
import org.apache.spark.ml.feature.{IDF, Tokenizer, RegexTokenizer, StopWordsRemover, CountVectorizer, StringIndexer, OneHotEncoder, VectorAssembler,CountVectorizerModel}
import org.apache.spark.ml.{Pipeline, PipelineModel}
import org.apache.spark.ml.classification.LogisticRegression
import org.apache.spark.ml.evaluation.MulticlassClassificationEvaluator
import org.apache.spark.ml.tuning.{ParamGridBuilder, TrainValidationSplit,TrainValidationSplitModel, CrossValidator, CrossValidatorModel}
// +
val paramGrid = new ParamGridBuilder()
.addGrid(cvModel.minDF, Array(55.0,75.0,95.0))
.addGrid(lr.regParam, Array(10e-8, 10e-6, 10e-4, 10e-2))
.build()
val lrcv = new CrossValidator()
.setEstimator(pipeline)
.setEstimatorParamMaps(paramGrid)
.setEvaluator(evaluator)
.setNumFolds(5)
val modelGridCV = lrcv.fit(training)
// -
| Scala notebook .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Map plots
#
# Plots on Mapbox maps are available only considering you have a Mapbox account and a Mapbox Access Token. After getting a mabox token it can be written set to pandapower as the following (where `'<token>'` needs to be replaced with provided mapbox token)
from pandapower.plotting.plotly.mapbox_plot import set_mapbox_token
set_mapbox_token('<token>')
# If network geodata are in Geographic coordinate system as latitude/longitude, a network can be plot on different maps.
# Moreover, if network geodata are not in latitude/longitude, but in some of the projections, it may be converted to lat/long by providing name of the projection (in the form '*epsg:projection_number*'according to http://spatialreference.org/ref/epsg/).
#
# Following example shows plot of the network mv_oberrhein, where network geodata are in Gauss-Kruger projection (zone 3).
# Since geodata are not in lat/long, plot using only `on_map=True` cannot be realized on a map:
# +
from pandapower.plotting.plotly import simple_plotly, pf_res_plotly, vlevel_plotly
from pandapower.networks import mv_oberrhein
net = mv_oberrhein()
# -
# The plot can be obtained if one knows specific projection and zone. In this case it is 3-degree Gauss-Kruger zone 3, which corresponds to [epsg:31467](http://spatialreference.org/ref/epsg/31467/):
net = mv_oberrhein()
simple_plotly(net, on_map=True, projection='epsg:31467')
# ## Transforming geo-data from a projection to lat/long
# There is a function available in pandapower which uses `pyproj` to transform geodata from a projection to WGS84 (lat/long). It transforms and replaces `net.bus_geodata` and `net.line_geodata` (if existing). An example for `mv_oberreihn`:
# +
net = mv_oberrhein()
print('before:\n', net.bus_geodata.head())
from pandapower.plotting.plotly.mapbox_plot import geo_data_to_latlong
geo_data_to_latlong(net, projection='epsg:31467')
print('\nafter:\n', net.bus_geodata.head())
# -
# ## Some more map plots...
# The following map styles are available:
# * 'streets'
# * 'bright'
# * 'light'
# * 'dark'
# * 'satellite'
simple_plotly(net, on_map=True,map_style='satellite')
net = mv_oberrhein()
simple_plotly(net, on_map=True, projection='epsg:31467', map_style='streets')
pf_res_plotly(net, on_map=True, map_style='dark')
# More tutorials about interactive plots using ploltly:
# * [built-in interactive plots](http://nbviewer.jupyter.org/github/e2nIEE/pandapower/blob/develop/tutorials/plotly_built-in.ipynb)
# * [custom interactive plots](http://nbviewer.jupyter.org/github/e2nIEE/pandapower/blob/develop/tutorials/plotly_traces.ipynb)
| tutorials/plotly_maps.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Boolean masks
import numpy as np
from numpy.random import default_rng
rg = default_rng(12345)
random_integers = rg.integers(low=1,high=5,size=100)
random_integers[:5]
is_equal_to_3 = random_integers == 3
is_equal_to_3[:5]
sum(is_equal_to_3)
random_integers[is_equal_to_3]
| Chapter01/Example_Boolean_Masks/Example_Boolean_masks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <font color='blue'>Data Science Academy - Python Fundamentos - Capítulo 2</font>
#
# ## Download: http://github.com/dsacademybr
# Versão da Linguagem Python
from platform import python_version
print('Versão da Linguagem Python Usada Neste Jupyter Notebook:', python_version())
# ## Exercícios Cap02
# +
# Exercício 1 - Imprima na tela os números de 1 a 10. Use uma lista para armazenar os números.
# -
lista = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10]
print(lista)
# +
# Exercício 2 - Crie uma lista de 5 objetos e imprima na tela
# -
lista = [1, 2, 3, 4, 5]
print(lista)
# +
# Exercício 3 - Crie duas strings e concatene as duas em uma terceira string
# -
s1 = "hello"
s2 = " world"
s3 = s1+s2
print(s3)
# +
# Exercício 4 - Crie uma tupla com os seguintes elementos: 1, 2, 2, 3, 4, 4, 4, 5 e depois utilize a função count do
# objeto tupla para verificar quantas vezes o número 4 aparece na tupla
# -
t = (1, 2, 2, 3, 4, 4, 4, 5)
t.count(4)
# +
# Exercício 5 - Crie um dicionário vazio e imprima na tela
# -
d = {}
print(d)
# +
# Exercício 6 - Crie um dicionário com 3 chaves e 3 valores e imprima na tela
# -
dict = {"key1": 1, "key2": 2, "key3": 3}
print(dict)
# +
# Exercício 7 - Adicione mais um elemento ao dicionário criado no exercício anterior e imprima na tela
# -
dict2 = {"key4":4}
dict.update(dict2)
print(dict)
# +
# Exercício 8 - Crie um dicionário com 3 chaves e 3 valores. Um dos valores deve ser uma lista de 2 elementos numéricos.
# Imprima o dicionário na tela.
# -
dict3 = {1: "key1", 2: [1, 2, 3], 3:"key3"}
print(dict3)
# +
# Exercício 9 - Crie uma lista de 4 elementos. O primeiro elemento deve ser uma string,
# o segundo uma tupla de 2 elementos, o terceiro um dcionário com 2 chaves e 2 valores e
# o quarto elemento um valor do tipo float.
# Imprima a lista na tela.
# -
lista = ["teste", (1, 2), {1: "um", 2: "dois"}, 2.4]
print(lista)
# Exercício 10 - Considere a string abaixo. Imprima na tela apenas os caracteres da posição 1 a 18.
frase = 'Cientista de Dados é o profissional mais sexy do século XXI'
print(frase[0:18])
# # Fim
# ### Obrigado
#
# ### Visite o Blog da Data Science Academy - <a href="http://blog.dsacademy.com.br">Blog DSA</a>
# Parabéns se você chegou até aqui. Use o voucher PYTHONDSA9642 para comprar qualquer curso ou Formação da DSA com 5% de desconto.
| Data Science Academy/PythonFundamentos/Cap02/Notebooks/DSA-Python-Cap02-Exercicios.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Classification with imbalanced class distributions is a major problem in machine learning. Researchers have given considerable attention to the applications in many real-world scenarios. Although several works have utilized the area under the receiver operating characteristic (ROC) curve to select potentially optimal classifiers in imbalanced classifications, limited studies have been devoted to finding the classification threshold for testing or unknown datasets. In general, the classification threshold is simply set to 0.5, which is usually unsuitable for an imbalanced classification. Here I showed, thresholds based on FPR-TPR, Precision-Recall-F1score, and Youden J-index in Python
# ## Loading Libraries
# +
# Loading libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib as mpl
from scipy import interp
from sklearn.preprocessing import scale
from sklearn.metrics import roc_auc_score, classification_report, accuracy_score, roc_curve, confusion_matrix, average_precision_score, precision_recall_curve
from sklearn.model_selection import cross_val_score, KFold, StratifiedKFold, train_test_split
from xgboost import XGBClassifier
import itertools
import glmnet
import xgboost as xgb
import seaborn as sns
sns.set_style("ticks")
mpl.rcParams['axes.linewidth'] = 3
mpl.rcParams['lines.linewidth'] = 2
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:95% !important; }</style>"))
import warnings
warnings.filterwarnings("ignore")
# %matplotlib inline
# -
# # Functions
# +
#*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-
# Function: _model
def _clf_train(X_train, y_train, X_test, y_test,
learning_rate = 0.05,
n_estimators = 100,
max_depth = 3,
min_child_weight = 5.0,
gamma = 1,
reg_alpha = 0.0,
reg_lambda = 1.0,
subsample = 0.9,
colsample_bytree = 0.9,
objective = "binary:logistic",
nthread = 4,
scale_pos_weight = 1.0,
seed = 1367,
random_state = 1367):
"""
an xgboost model for training
"""
clf = XGBClassifier(learning_rate = learning_rate,
n_estimators = n_estimators,
max_depth = max_depth,
min_child_weight = min_child_weight,
gamma = gamma,
reg_alpha = reg_alpha,
reg_lambda = reg_lambda,
subsample = subsample,
colsample_bytree = colsample_bytree,
objective = objective,
nthread = nthread,
scale_pos_weight = scale_pos_weight,
seed = seed,
random_state = random_state)
clf.fit(X_train, y_train, eval_metric = "auc", early_stopping_rounds = 20, verbose = True, eval_set = [(X_test, y_test)])
return clf
#*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-
# Function: Finding thresholds
def _threshold_finder(model, X, y_true):
"""
a function to find the optimal threshold for binary classification
model: a trained model object (such as xgboost, glmnet, ...)
X: the test set of features (pandas dataframe or numpy array)
y_true: the true class labels (list or array of 0's and 1's)
"""
y_predict_proba = model.predict_proba(X)[:, 1]
fpr, tpr, thresholds = roc_curve(y_true, y_predict_proba)
auc = roc_auc_score(y_true, y_predict_proba)
precision, recall, thresholds2 = precision_recall_curve(y_true, y_predict_proba)
class_names = [0, 1]
# Youden Threshold
youden_idx = np.argmax(np.abs(tpr - fpr))
youden_threshold = thresholds[youden_idx]
y_pred_youden = (y_predict_proba > youden_threshold).astype(int)
cnf_matrix = confusion_matrix(y_true, y_pred_youden)
np.set_printoptions(precision=2)
# Sensitivity-Specifity Threshold
sens_spec_threshold = thresholds[np.argmin(abs(tpr + fpr - 1))]
y_pred_sens_spec = (y_predict_proba > sens_spec_threshold).astype(int)
cnf_matrix2 = confusion_matrix(y_true, y_pred_sens_spec)
# precision-recall threshold
prec_rec_threshold = thresholds2[np.argmin(abs(precision-recall))]
y_pred_prec_rec = (y_predict_proba > prec_rec_threshold).astype(int)
cnf_matrix3 = confusion_matrix(y_true, y_pred_prec_rec)
f1 = []
for i in range(len(precision)):
f1.append(2 * (precision[i] * recall[i]) / (precision[i] + recall[i]))
queue_rate = []
for thr in thresholds2:
queue_rate.append((y_predict_proba >= thr).mean())
# plotting
plt.figure(figsize = (12, 5))
plt.subplot(1,2,1)
plt.plot(fpr, tpr, color = "red", label = F"AUC = {auc:.3f}")
plt.plot(fpr[youden_idx], tpr[youden_idx], marker = "o", color = "navy", ms =10, label =F"Youden Threshold = {youden_threshold:.2f}" )
plt.axvline(x = fpr[youden_idx], ymin = fpr[youden_idx], ymax = tpr[youden_idx], color = "navy", ls = "--")
plt.plot([0,1], [0,1] , color = "black", ls = "--")
plt.xlim([-0.01, 1.01])
plt.ylim([-0.01, 1.01])
plt.xlabel('1 - Specificity' , fontsize=12)
plt.ylabel('Sensitivity' , fontsize=12)
plt.tick_params(axis='both', which='major', labelsize=12)
plt.legend( prop={'size':12} , loc = 4)
plt.subplot(1,2,2)
_plot_confusion_matrix(cnf_matrix, classes=class_names, normalize = False, cmap=plt.cm.Reds, title = F"Youden Threshold = {youden_threshold:.3f}\nAccuracy = {accuracy_score(y_true, y_pred_youden)*100:.2f}%")
plt.show()
plt.figure(figsize = (12, 5))
plt.subplot(1,2,1)
plt.plot(thresholds, 1-fpr, label = "1 - Specificity")
plt.plot(thresholds, tpr, label = "Sensitivity")
plt.xlabel("Threshold", fontsize = 12)
plt.ylabel("Score", fontsize = 12)
plt.legend(loc = 0)
plt.xlim([0.025, thresholds[np.argmin(abs(tpr + fpr - 1))]+0.2])
plt.axvline(thresholds[np.argmin(abs(tpr + fpr - 1))], color="k", ls = "--")
plt.title(F"Threshold = {sens_spec_threshold:.3f}", fontsize = 12)
plt.subplot(1,2,2)
_plot_confusion_matrix(cnf_matrix2, classes=class_names, normalize = False, cmap=plt.cm.Reds, title = F"Sensitivity-Specificity Threshold = {sens_spec_threshold:.3f}\nAccuracy = {accuracy_score(y_true, y_pred_sens_spec)*100:.2f}%")
plt.show()
plt.figure(figsize = (12, 5))
plt.subplot(1,2,1)
plt.plot(thresholds2, precision[1:], label = "Precision")
plt.plot(thresholds2, recall[1:], label = "Recall")
plt.plot(thresholds2, f1[1:], label = "F1-Score")
plt.plot(thresholds2, queue_rate, label = "Queue Rate")
plt.legend(loc = 0)
plt.xlim([0.025, thresholds2[np.argmin(abs(precision-recall))] + 0.2])
plt.xlabel("Threshold", fontsize = 12)
plt.ylabel("Score", fontsize = 12)
plt.axvline(thresholds2[np.argmin(abs(precision-recall))], color="k", ls = "--")
plt.title(label = F"Threshold = {prec_rec_threshold:.3f}", fontsize = 12)
plt.subplot(1,2,2)
_plot_confusion_matrix(cnf_matrix3, classes=class_names, normalize = False, cmap=plt.cm.Reds, title = F"F1-Score Threshold = {prec_rec_threshold:.3f}\nAccuracy = {accuracy_score(y_true, y_pred_prec_rec)*100:.2f}%")
plt.show()
#*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-*-
# Function: Plotting Confusion Matrix
def _plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Greens):
from sklearn.metrics import precision_score, recall_score, roc_auc_score, accuracy_score, roc_curve, auc, confusion_matrix
import itertools
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title, fontsize = 14)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="black")
# horizontalalignment="center",
# ha="center", va="bottom",
plt.ylabel('True Class', fontsize = 14)
plt.xlabel('Predicted Class', fontsize = 14)
plt.tick_params(axis='both', which='major', labelsize=14)
plt.tight_layout()
# -
_threshold_finder(model = model, X = X_test, y_true = y_test)
# # Device Failure Data
# ### First, I loaded the data into a pandas dataframe to get some idea.
# readin the data into a dataframe
dateparser = lambda x: pd.datetime.strptime(x, "%Y-%m-%d")
df_raw = pd.read_csv("/home/amirhessam/Projects/Amazon/device_failure.csv",
parse_dates = ["date"],
date_parser = dateparser,
encoding = "cp1252")
print("Shape: {}".format(df_raw.shape))
print("Prevalence = {:.3f}%".format(df_raw["failure"].sum()/df_raw.shape[0] * 100))
df_raw.head()
# ### Preprocessing and builing X, Y training/testing sets.
# +
target = "failure"
to_drop = ["date", "device", "attribute8", "failure"]
y = df_raw[target].values
X = df_raw.drop(to_drop, axis = 1)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.20, shuffle = True, random_state = 1367, stratify = y)
print(F"Train Size = {X_train.shape}")
print(F"Test Size = {X_test.shape}")
# -
# ### Printing the first 5 rows of the sclaed features.
X_train.head()
# ### Train a model
model = _clf_train(X_train, y_train, X_test, y_test)
# ### Visualization of Thresholds
_threshold_finder(model = model, X = X_test, y_true = y_test)
# # MNIST DATA
from sklearn.datasets import fetch_openml
mnist = fetch_openml("mnist_784", version = 1)
mnist.keys()
X, y = mnist["data"], mnist["target"]
X.shape
plt.figure()
plt.imshow(X[1000].reshape(28, 28), cmap = "gray")
plt.show()
# ### As you know, minist contains 10 classes. So, we need to turn this multi-class data to a binary class.
X_train, X_test, y_train, y_test = X[:60000], X[60000:], y[:60000], y[60000:]
y_train_0 = np.where(y_train == "0", 1, 0)
y_test_0 = np.where(y_test == "0", 1, 0)
model = _clf_train(X_train, y_train_0, X_test, y_test_0)
_threshold_finder(model = model, X = X_test, y_true = y_test_0)
# +
print(__doc__)
import numpy as np
import matplotlib.pyplot as plt
from sklearn import svm, datasets
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix
from sklearn.utils.multiclass import unique_labels
# import some data to play with
iris = datasets.load_iris()
X = iris.data
y = iris.target
class_names = iris.target_names
# Split the data into a training set and a test set
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
# Run classifier, using a model that is too regularized (C too low) to see
# the impact on the results
classifier = svm.SVC(kernel='linear', C=0.01)
y_pred = classifier.fit(X_train, y_train).predict(X_test)
def plot_confusion_matrix(y_true, y_pred, classes,
normalize=False,
title=None,
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if not title:
if normalize:
title = 'Normalized confusion matrix'
else:
title = 'Confusion matrix, without normalization'
# Compute confusion matrix
cm = confusion_matrix(y_true, y_pred)
# Only use the labels that appear in the data
classes = classes[unique_labels(y_true, y_pred)]
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
print(cm)
fig, ax = plt.subplots()
im = ax.imshow(cm, interpolation='nearest', cmap=cmap)
ax.figure.colorbar(im, ax=ax)
# We want to show all ticks...
ax.set(xticks=np.arange(cm.shape[1]),
yticks=np.arange(cm.shape[0]),
# ... and label them with the respective list entries
xticklabels=classes, yticklabels=classes,
title=title,
ylabel='True label',
xlabel='Predicted label')
# Rotate the tick labels and set their alignment.
plt.setp(ax.get_xticklabels(), rotation=45, ha="right",
rotation_mode="anchor")
# Loop over data dimensions and create text annotations.
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i in range(cm.shape[0]):
for j in range(cm.shape[1]):
ax.text(j, i, format(cm[i, j], fmt),
ha="center", va="center",
color="white" if cm[i, j] > thresh else "black")
fig.tight_layout()
return ax
np.set_printoptions(precision=2)
# Plot non-normalized confusion matrix
plot_confusion_matrix(y_test, y_pred, classes=class_names,
title='Confusion matrix, without normalization')
# Plot normalized confusion matrix
plot_confusion_matrix(y_test, y_pred, classes=class_names, normalize=True,
title='Normalized confusion matrix')
plt.show()
# -
| notebooks/finding-thresholds.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# sbpy.data.Phys Example Notebook
# =================================
#
# [sbpy.data.Phys](https://sbpy.readthedocs.io/en/latest/api/sbpy.data.Phys.html#sbpy.data.Phys) provides functionality to query and store small-body physical properties.
# Querying Physical Properties from JPL SBDB
# --------------------------------------------
#
# Query the physical properties for a few asteroids:
# +
from sbpy.data import Phys
phys = Phys.from_sbdb(['433', 'Itokawa', '12893'])
print(phys.column_names)
# -
# We can calculate their volumes assuming spherical shapes:
import numpy as np
list(zip(phys['targetname'], 4/3*np.pi*phys['diameter']**3))
# Please keep in mind that physical properties information provided by SBDB is incomplete and scarce. Missing values are replaced with `nan`.
# Querying Molecular Data from `astroquery.jplspec`
# ==========================
#
# `sbpy.data.Phys` also contains a function to query molecular data that
# might be useful for various calculations such as production rate calculations.
# `sbpy.data.Phys.from_jplspec` queries the [JPL Molecular Spectroscopy Catalog](https://spec.jpl.nasa.gov/home.html) molecular properties, and stores the
# data in a `sbpy.data.Phys` object, offering the same functionality as all the
# other `sbpy.data` functions, including the use of `~astropy.units`. `sbpy.data.Phys.from_jplspec` also
# calculates the partition function at the desired temperature using log-space interpolation. For
# a briefing of how this is done look at the sbpy jplspec notebook: [jplspec notebook](jplspec.ipynb). The results
# from `sbpy.data.Phys.from_jplspec` include the following data:
#
# - Transition frequency
# - Temperature
# - Integrated line intensity at 300 K
# - Partition function at 300 K
# - Partition function at designated temperature
# - Upper state degeneracy
# - Upper level energy in Joules
# - Lower level energy in Joules
# - Degrees of freedom
#
# For the names of these fields and their alternatives, see [here](https://sbpy.readthedocs.io/en/latest/sbpy/data/fieldnames.html#id1)
import astropy.units as u
# +
temp_estimate = 47. * u.K # kinetic temperature
mol_tag = 28001 # JPLSpec unique identifier
transition_freq = (345.7 * u.GHz).to('MHz') # Transition frequency
mol_data = Phys.from_jplspec(temp_estimate, transition_freq, mol_tag) # build Phys object
# -
# Once the phys object has been created, one can access all the information given by astroquery.jplspec as well as the partition function at the desired temperature:
print(mol_data['elo_j']) # print energy of lower level
print(mol_data['degfr']) # print degrees of freedom
print(mol_data['partfn']) # print interpolated partition function at desired temp
# Regular expressions can also be used as molecule identifiers since astroquery.jplspec provides the functionality. It is important to understand regular expressions and how to use them in a way that gets the user exactly what they want. If the user is unfamiliar with regular expressions, using the JPL Spectral catalog unique identifier is the best course of action.
#
# Hint: in regular expressions, putting something between '^' and '\$' will match the literal text in between. This is useful to remember in order to avoid matching unnecessary terms. i.e. using 'co' will match 'co', 'co2', 'hco' so you would want to input '^co$' to avoid such nuances
# +
mol_tag = '^CO$'
mol_data = Phys.from_jplspec(temp_estimate, transition_freq, mol_tag) # build Phys object
print(mol_data['elo_j']) # print energy of lower level
print(mol_data['degfr']) # print degrees of freedom
print(mol_data['partfn']) # print interpolated partition function at desired temp
| notebooks/data/Phys.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "skip"}
# <table>
# <tr align=left><td><img align=left src="./images/CC-BY.png">
# <td>Text provided under a Creative Commons Attribution license, CC-BY. All code is made available under the FSF-approved MIT license. (c) <NAME></td>
# </table>
#
# Note: This material largely follows the text "Numerical Linear Algebra" by <NAME> Bau (SIAM, 1997) and is meant as a guide and supplement to the material presented there.
# + init_cell=true slideshow={"slide_type": "skip"}
# %matplotlib inline
# %precision 3
import numpy
import matplotlib.pyplot as plt
# + [markdown] slideshow={"slide_type": "slide"}
# # Numerical Linear Algebra
#
# Numerical methods for linear algebra problems lies at the heart of many numerical approaches and is something we will spend some time on. Roughly we can break down problems that we would like to solve into three general problems, solving a system of equations
#
# $$
# A \mathbf{x} = \mathbf{b}
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# Projection problems or Linear Least Squares
#
# $$ A^TA\mathbf{x} = A^T\mathbf{b}$$
# + [markdown] slideshow={"slide_type": "fragment"}
# and solving the eigenvalue problem
#
# $$A \mathbf{v} = \lambda \mathbf{v}.$$
# + [markdown] slideshow={"slide_type": "fragment"}
# We examine each of these problems separately and will evaluate some of the fundamental properties and methods for solving these problems. We will be careful in deciding how to evaluate the results of our calculations and try to gain some understanding of when and how they fail.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Factorizations
#
# For each fundamental problem, there are a range of algorithms to solve them, but all of these algorithms can often be succinctly described in terms of a matrix "factorization", the ability to write an arbitrary matrix $A$ as a product of matrices with special properties (e.g. triangular, diagonal, orthogonal) that make solving the general problem straightforward. For example
# + [markdown] slideshow={"slide_type": "fragment"}
# <table width="80%">
# <tr align="left"><th>Problem</th> <th align="center">Algorithms</th> <th align="center">"Factorizations"</th></tr>
# <tr align="left"><td>$$A \mathbf{x} = \mathbf{b}$$</td> <td align="left">Gaussian-Elimination, Gauss-Jordan Elimination</td> <td align="center">$PA=LU$, $A=ER$</td></tr>
# <tr align="center"><td>$$ A^TA\mathbf{x} = A^T\mathbf{b}$$</td> <td align="center">Orthogonalization algorithms (Gram-Schmidt, Modified GS, Householder, Givens)</td> <td align="center">$A=QR$</td></tr>
# <tr align="center"><td>$$A \mathbf{v} = \lambda \mathbf{v}$$</td> <td align="center">Various Iterative methods (Power, inverse power, $RQ$ with shifts$\ldots$</td> <td align="center">$A=S\Lambda S^{-1}$, $A=Q\Lambda Q^T$, $A=MJM^{-1}$</td></tr>
# <tr align="center"><td align="left">All of the above</td> <td align="center">Singular Value Decomposition (SVD)</td> <td align="center">$A = U\Sigma V^T$</td></tr>
# </table>
# + [markdown] slideshow={"slide_type": "fragment"}
# To develop all of these algorithms in depth is a course in itself, however, here we will highlight some of the key factorizations and algorithms used in modern computational linear algebra. In particular we will highlight issues of accuracy, stability and computational cost particularly in the limit of large systems.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Some Example Problems
#
# The number and power of the different tools made available from the study of linear algebra makes it an invaluable field of study. Before we dive in to numerical approximations we first consider some of the pivotal problems that numerical methods for linear algebra are used to address.
#
# For this discussion we will be using the common notation $m \times n$ to denote the dimensions of a matrix $A$. The $m$ refers to the number of rows and $n$ the number of columns. If a matrix is square, i.e. $m = n$, then we will use the notation that $A$ is $m \times m$.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Systems of Equations
#
# The first type of problem is to find the solution to a linear system of equations. If we have $m$ equations for $m$ unknowns it can be written in matrix/vector form,
#
# $$A \mathbf{x} = \mathbf{b}.$$
#
# For this example $A$ is an $m \times m$ matrix, denoted as being in $\mathbb{R}^{m\times m}$, and $\mathbf{x}$ and $\mathbf{b}$ are column vectors with $m$ entries, denoted as $\mathbb{R}^m$.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Example 1: Polynomial Fitting
# In our previous work on interpolation we found that the unique interpolating polynomial of order $n$ through the $n+1$ points $(x_i, y_i)$ can be found by solving a Linear system of equations
#
# $$
# V[\phi_j(\mathbf{x})]\mathbf{w} = \mathbf{y}
# $$
#
# where $V$ is a VanderMonde matrix whose columns are the basis functions $\phi_j(\mathbf{x})$ (e.g. the monomials) and $\mathbf{w}\in\mathbb{R}^{n+1}$ are the coefficients (weights) such that the interpolating polynomial is given uniquely by
# $$
# \cal{P}_n(x) = \sum_{j=0}^n w_j\phi_j(x)
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Examples 2: Solution of Non-linear equations by Newton's Method
#
# Given a non-linear system of equations
#
# $$\mathbf{F}(\mathbf{x})=\mathbf{0}$$
#
# Newton's method provides an iterative method to find roots where at every step of the iteration a linear system of equations
#
# $$
# J(\mathbf{x_k})\boldsymbol{\delta}_k = -\mathbf{F}(\mathbf{x_k})
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Examples 3: Solution of Boundary Value problems by Finite Differences or Finite Elements
#
# Given a general linear differential equation
#
# $${\cal L}u = f$$
#
# where ${\cal L}$ is a linear differential operator e.g.
#
# $$
# {\cal L} = \frac{d^2}{dx^2} + \alpha\frac{d}{dx} + \beta
# $$
#
# Finite difference or finite element discretizations usually reduce the continuous, infinite dimensional problem to a discrete linear problem of form
#
# $$
# A\mathbf{u} = \mathbf{f}
# $$
#
# Where $\mathbf{u}$ and $\mathbf{f}$ are discrete approximations to the solution function and right hand side and $A$ is a discrete approximation to ${\cal L}$. Moreover, for many discretizations, $A$ can be very sparse.
# + [markdown] hide_input=true slideshow={"slide_type": "subslide"}
# ### Linear least squares
#
# In a similar case as above, say we want to fit a particular function (could be a polynomial) to a given number of data points except in this case we have more data points than free parameters. In the case of polynomials this could be the same as saying we have $m$ data points but only want to fit a $n - 1$ order polynomial through the data where $n - 1 \leq m$. One of the common approaches to this problem is to minimize the "least-squares" error between the data and the resulting function:
# $$
# E = \left( \sum^m_{i=1} |y_i - f(x_i)|^2 \right )^{1/2}.
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# E.g. Consider fitting the function
# $$
# f(x) = w_1 + w_2 x + w_3 e^x
# $$
#
# to data that has random noise added to it.
# + [markdown] slideshow={"slide_type": "subslide"}
# If our function $f(x)$ can be written as a linear combination of basis functions
#
# $$
# f(x) = \sum_{j=1}^n w_j\phi_j(x)
# $$
#
# then the linear system through $m$ points becomes $A\mathbf{w} = \mathbf{y}$ where $A$ is a generalized $m\times n$ vandermonde matrix
#
# $$
# A = \begin{bmatrix}
# \phi_1(x_1) & \phi_2(x_1) & \cdots & \phi_n(x_1) \\
# \phi_1(x_2) & \phi_2(x_2) & \cdots & \phi_n(x_2) \\
# \vdots & \vdots & &\vdots \\
# \phi_1(x_m) & \phi_2(x_m) & \cdots & \phi_n(x_m) \\
# \end{bmatrix}
# $$
#
# Which will be over-determined for $m > n$. However the Least-squares solution for the weights $\mathbf{w}$ will satisfy
#
# $$A^T A \mathbf{w} = A^T \mathbf{y}$$
#
# we can guarantee that the error is minimized in the least-squares sense[<sup>1</sup>](#footnoteRegression). (Although we will also show that this is not the most numerically stable way to solve this problem)
#
#
# + [markdown] slideshow={"slide_type": "skip"}
# #### Define the data
# + slideshow={"slide_type": "skip"}
N = 20
x = numpy.linspace(-1.0, 1.0, N)
y = x + numpy.random.random((N))
# + [markdown] slideshow={"slide_type": "skip"}
# #### Define basis functions and Vandermonde matrix
# + slideshow={"slide_type": "skip"}
# define the basis functions
phi_1 = lambda x: numpy.ones(x.shape)
phi_2 = lambda x: x
phi_3 = lambda x: numpy.exp(x)
# Define various Vandermonde matrix based on our x-values
A = numpy.array([ phi_1(x), phi_2(x), phi_3(x)]).T
A.shape
# + slideshow={"slide_type": "skip"}
# Determine the weights of our linear function
# result in the smallest sum of the squares of the residual.
w = numpy.linalg.solve(numpy.dot(A.T, A), numpy.dot(A.T, y))
error = y - A.dot(w)
print('w = {}, ||e|| = {}'.format(w,numpy.linalg.norm(error)))
# + slideshow={"slide_type": "skip"}
# Do the same using numpy's lstsq routine (which solves a more numerically stable problem)
w = numpy.linalg.lstsq(A,y, rcond=None)[0]
error = y - A.dot(w)
print('w = {}, ||e|| = {}'.format(w,numpy.linalg.norm(error)))
# + hide_input=true slideshow={"slide_type": "subslide"}
# just repeat the calculations to draw the figure
# define data
N = 20
x = numpy.linspace(-1.0, 1.0, N)
y = x + numpy.random.random((N))
# define the basis functions
phi_1 = lambda x: numpy.ones(x.shape)
phi_2 = lambda x: x
phi_3 = lambda x: numpy.exp(x)
# Define various Vandermonde matrix based on our x-values
A = numpy.array([ phi_1(x), phi_2(x), phi_3(x)]).T
w = numpy.linalg.lstsq(A,y, rcond=None)[0]
# Plot it out, cuz pictures are fun!
fig = plt.figure(figsize=(8,6))
axes = fig.add_subplot(1, 1, 1)
f = A.dot(w)
axes.plot(x, y, 'ko')
axes.plot(x, f, 'r')
axes.set_title("Least Squares Fit to Data")
axes.set_xlabel("$x$")
axes.set_ylabel("$f(x)$ and $y_i$")
axes.grid()
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# ### Eigenproblems
#
# Eigenproblems come up in a variety of contexts and often are integral to many problem of scientific and engineering interest. It is such a powerful idea that it is not uncommon for us to take a problem and convert it into an eigenproblem.
#
# One of my favorite examples [The Tacoma Narrows Bridge Collapse](https://www.youtube.com/watch?v=XggxeuFDaDU)
#
# Or the original Google [Page-Rank](https://en.wikipedia.org/wiki/PageRank) algorithm which essentially finds the dominant eigenvector of an enormous sparse Markov Matrix.
# + [markdown] slideshow={"slide_type": "skip"}
# Here we introduce the idea and give some examples.
#
# As a review, if $A \in \mathbb{C}^{m\times m}$ (a square matrix with complex values), a non-zero vector $\mathbf{v}\in\mathbb{C}^m$ is an **eigenvector** of $A$ with a corresponding **eigenvalue** $\lambda \in \mathbb{C}$ if
#
# $$A \mathbf{v} = \lambda \mathbf{v}.$$
#
# One way to interpret the eigenproblem is that we are attempting to ascertain the "action" of the matrix $A$ on some subspace of $\mathbb{C}^m$ where this action acts like scalar multiplication. This subspace is called an **eigenspace**.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### General idea of EigenProblems
#
# Rewriting the standard Eigen problem $A\mathbf{v}=\lambda\mathbf{v}$ for $A \in \mathbb{C}^{m\times m}$, $\mathbf{v}\in\mathbb{C}^m$ as
#
# $$
# (A - \lambda I)\mathbf{v} = \mathbf{0}
# $$
#
# it becomes clear that for $\mathbf{v}$ to be non-trivial (i.e. $\neq \mathbf{0}$), requires that the matrix $(A-\lambda I)$ be singular,
#
# This is equivalent to finding all values of $\lambda$ such that $|A-\lambda I| = 0$ (the determinant of singular matrices is always zero). However, it can also be shown that
#
# $$
# | A-\lambda I| = P_m(\lambda)
# $$
#
# which is a $m$th order polynomial in $\lambda$. Thus $P_m(\lambda)=0$ implies the eigenvalues are the $m$ roots of $P$, and the **eigenspace** corresponding to $\lambda_i$ is just $N(A-\lambda_i I)$
#
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Solving EigenProblems
#
# The temptation (and what we usually teach in introductory linear algebra classes) is to find the roots of $P_m(\lambda)$ to get the eigenvalues, then find the null-space of $A-\lambda I$.
# + [markdown] slideshow={"slide_type": "fragment"}
# However that would be **wrong**. The best algorithms for finding Eigenvalues are completely unrelated to rootfinding as we shall see (and in fact, the way you find the roots of polynomials is to find the eigenvalues of a "companion matrix")
# + [markdown] slideshow={"slide_type": "subslide"}
# ### The Singular Value Decomposition
#
# One of the most beautiful, and useful, factorizations is the Singular Value Decomposition (or SVD),
#
# $$
# A = U\Sigma V^T
# $$
#
# where $A$ is a general $m\times n$ matrix, $U$ and $V$ are unitary (orthogonal) matrices such that $U^TU=I^{m\times m}$, $V^TV = I^{n\times n}$ and $\Sigma$ is a diagonal matrix with real, positive diagonal entries (the singular values)
#
# $$
# \sigma_1 \geq \sigma_2\geq \ldots \sigma_{r} > 0
# $$
# where $r$ is the rank of $A$
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Applications of the SVD
#
# The SVD combines all the aspects of basic linear algebra and is used in a large number of applications including
#
# * diagnosing ill-conditioned matrices
# * providing orthonormal bases for the 4-subspaces of $A$
# * Solving linear systems and linear least-squares problems
# * Solving ill-conditioned and singular linear systems (Pseudo-inverse)
# * Dimensional reduction in Data analysis (PCA, EOF, POD analysis)
# * and more...
#
# Because of its ubiquity in computational linear algebra and data science. we will spend a bit of time to understand the SVD and its applications (but not the specific algorithms). But first
# + [markdown] slideshow={"slide_type": "slide"}
# ## Fundamentals
#
# **Objectives**
# * Understand basic linear-algebraic operations and their **computational costs**
# * Understand Numpy implementation (and performance) for Linear Algebra
# * Understand Singular vs Invertible matrices
# * Understand Orthonormal vectors and $Q$ matrices
# * Understand vector and Matrix norms
# * Undertand the **Condition Number**
# + [markdown] slideshow={"slide_type": "slide"}
# ### Matrix-Vector Multiplication
#
# One of the most basic operations we can perform with matrices is to multiply them by a vector which maps a vector to a vector. There are multiple ways to interpret and compute the matrix-vector product $A \mathbf{x}$.
#
# #### index notation
# $$
# b_i = \sum^n_{j=1} a_{ij} x_j \quad \text{where}\quad i = 1, \ldots, m
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# #### row picture (dot products)
# We can also consider matrix-vector multiplication as a sequence of inner products (dot-products between the rows of $A$ and the vector $\mathbf{x}$.
# \begin{align}
# \mathbf{b} &= A \mathbf{x}, \\
# &=
# \begin{bmatrix} \mathbf{a}_1^T \mathbf{x} \\ \mathbf{a}_2^T \mathbf{x} \\ \vdots \\ \mathbf{a}_m^T \mathbf{x}\end{bmatrix}
# \end{align}
# where $\mathbf{a}_i^T$ is the $i$th **row** of $A$
# + [markdown] slideshow={"slide_type": "subslide"}
# #### column picture
#
# An alternative (and entirely equivalent way) to write the matrix-vector product is as a linear combination of the columns of $A$ where each column's weighting is $x_j$.
#
# $$
# \begin{align}
# \mathbf{b} &= A \mathbf{x}, \\
# &=
# \begin{bmatrix} & & & \\ & & & \\ \mathbf{a}_1 & \mathbf{a}_2 & \cdots & \mathbf{a}_n \\ & & & \\ & & & \end{bmatrix}
# \begin{bmatrix} x_1 \\ x_2 \\ \vdots \\ x_n \end{bmatrix}, \\
# &= x_1 \mathbf{a}_1 + x_2 \mathbf{a}_2 + \cdots + x_n \mathbf{a}_n.
# \end{align}
# $$
#
# This view will be useful later when we are trying to interpret various types of matrices.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Operation Counts
# No matter how you compute $A\mathbf{x}$, the total number of operations is the same (for a dense matrix $A$).
# The row view however is convenient for calculating the **Operation counts** required for $A\mathbf{x}$.
#
# If $A\in\mathbb{C}^{m\times n}$ and $\mathbf{x}\in\mathbb{C}^n$. Then just counting the number of multiplications involved to compute $A\mathbf{x}$ is $O(??)$
#
# + [markdown] slideshow={"slide_type": "subslide"}
# One important property of the matrix-vector product is that is a **linear** operation, also known as a **linear operator**. This means that the for any $\mathbf{x}, \mathbf{y} \in \mathbb{C}^n$ and any $c \in \mathbb{C}$ we know that
#
# 1. $A (\mathbf{x} + \mathbf{y}) = A\mathbf{x} + A\mathbf{y}$
# 1. $A\cdot (c\mathbf{x}) = c A \mathbf{x}$
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Example: Numerical matrix-vector multiply
#
# Write a matrix-vector multiply function and check it with the appropriate `numpy` routine. Also verify the linearity of the matrix-vector multiply.
# + slideshow={"slide_type": "subslide"}
# Mat-vec in index notation (the long way...don't do this)
def matrix_vector_product_index(A, x):
m, n = A.shape
b = numpy.zeros(m)
for i in range(m):
for j in range(n):
b[i] += A[i, j] * x[j]
return b
# Mat-vec by row picture (still don't do this)
def matrix_vector_product_row(A, x):
m, n = A.shape
b = numpy.zeros(m)
# loop on rows
for i in range(m):
b[i] = A[i, :].dot(x)
return b
# Mat-vec by column picture (still don't do this)
def matrix_vector_product_col(A, x):
m, n = A.shape
b = numpy.zeros(m)
# loop over columns
for j in range(n):
b += A[:, j] * x[j]
return b
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Check equivalence
#
# first set up some large random matrices and vectors and compare to numpy built in .dot function
# + slideshow={"slide_type": "-"}
m = 1000
n = 1000
A = numpy.random.uniform(size=(m,n))
x = numpy.random.uniform(size=(n))
y = numpy.random.uniform(size=(n))
c = numpy.random.uniform()
# + slideshow={"slide_type": "-"}
funcs = [ matrix_vector_product_index, matrix_vector_product_row, matrix_vector_product_col ]
for f in funcs:
b = f(A, x)
print('{}(A,x) = A.dot(x)? {}'.format(f.__name__, numpy.allclose(b, A.dot(x))))
# + [markdown] slideshow={"slide_type": "skip"}
# #### Check Linearity
# + slideshow={"slide_type": "skip"}
for f in funcs:
print('{}(A,x+y) = Ax + Ay is {}'.format(f.__name__,
numpy.allclose(f(A, (x + y)), f(A, x) + f(A, y))))
print('{}(A,cx) = cAx is {}\n'.format(f.__name__,
numpy.allclose(f(A, c * x), c*f(A, x))))
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Check Timing/performance
# + slideshow={"slide_type": "-"}
for f in funcs:
print(f.__name__,end='\t')
# %timeit f(A,x)
print('numpy.dot(A,x)',end='\t\t\t')
# %timeit A.dot(x)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Matrix-Matrix Multiplication
#
# The matrix product with another matrix $ C=AB$ is defined as
# $$
# c_{ij} = \sum^m_{k=1} a_{ik} b_{kj} = \mathbf{a}_i^T\mathbf{b}_j
# $$
#
# i.e. each component of $C$ is a dot-product between the $i$th row of $A$ and the $j$th column of $B$
# + [markdown] slideshow={"slide_type": "subslide"}
# As with matrix-vector multiplication, Matrix-matrix multiplication can be thought of multiple ways
#
# * $m\times p$ dot products (each with $n$ flops)
# * $A$ multiplying the columns of $B$
# $$
# C = AB = \begin{bmatrix}
# A\mathbf{b}_1 & A\mathbf{b}_2 & \ldots & A\mathbf{b}_p\\
# \end{bmatrix}
# $$
# * Linear combinations of the rows of $B$
# $$
# C = AB = \begin{bmatrix}
# \mathbf{a}_1^T B \\ \mathbf{a}_2^T B \\ \vdots \\ \mathbf{a}_m^T B\\
# \end{bmatrix}
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Questions
# * What are the dimensions of $A$ and $B$ so that the multiplication works?
# * What are the Operations Counts for Matrix-Matrix Multiplication?
# * Comment on the product $\mathbf{c}=(AB)\mathbf{x}$ vs. $\mathbf{d} = A(B\mathbf{x})$
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Example: Outer Product
#
# The product of two vectors $\mathbf{u} \in \mathbb{C}^m$ and $\mathbf{v} \in \mathbb{C}^n$ is a $m \times n$ matrix where the columns are the vector $u$ multiplied by the corresponding value of $v$:
# $$
# \begin{align}
# \mathbf{u} \mathbf{v}^T &=
# \begin{bmatrix} u_1 \\ u_2 \\ \vdots \\ u_m \end{bmatrix}
# \begin{bmatrix} v_1 & v_2 & \cdots & v_n \end{bmatrix}, \\
# & = \begin{bmatrix} v_1u_1 & \cdots & v_n u_1 \\ \vdots & & \vdots \\ v_1 u_m & \cdots & v_n u_m \end{bmatrix}.
# \end{align}
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# It is useful to think of these as operations on the column vectors, and an equivalent way to express this relationship is
# $$
# \begin{align}
# \mathbf{u} \mathbf{v}^T &=
# \begin{bmatrix} \\ \mathbf{u} \\ \\ \end{bmatrix}
# \begin{bmatrix} v_1 & v_2 & \cdots & v_n \end{bmatrix}, \\
# &=
# \begin{bmatrix} & & & \\ & & & \\ \mathbf{u}v_1 & \mathbf{u} v_2 & \cdots & \mathbf{u} v_n \\ & & & \\ & & & \end{bmatrix}, \\
# & = \begin{bmatrix} v_1u_1 & \cdots & v_n u_1 \\ \vdots & & \vdots \\ v_1 u_m & \cdots & v_n u_m \end{bmatrix}.
# \end{align}
# $$
#
# Or each column is just a scalar multiple of $\mathbf{u}$
# + [markdown] slideshow={"slide_type": "fragment"}
# Alternatively you can think of this as the rows are all just scalar multiples of $\mathbf{v}$
# + [markdown] slideshow={"slide_type": "subslide"}
# ### rank 1 updates
#
# We call any matrix of the form $\mathbf{u}\mathbf{v}^T$ a "rank one matrix" ( because its rank r=?). These sort of matrix operations are very common in numerical algorithms for orthogonalization, eigenvalues and the original page-rank algorithm for google. Again, the order of operations is critical.
# + [markdown] slideshow={"slide_type": "fragment"}
# Comment on the difference in values and operation counts between
#
# $$
# \mathbf{y} = (\mathbf{u}\mathbf{v}^T)\mathbf{x}
# $$
#
# and
# $$
# \tilde{\mathbf{y}} = \mathbf{u}(\mathbf{v}^T\mathbf{x})
# $$
# for $\mathbf{u}$, $\mathbf{v}$, $\mathbf{x}$, $\mathbf{y}$, $\tilde{\mathbf{y}}\in\mathbb{R}^n$,
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Check
#
# Set up some random vectors
# + slideshow={"slide_type": "-"}
m = 10000
n = 10000
u = numpy.random.rand(m)
v = numpy.random.rand(n)
x = numpy.random.rand(n)
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Check Equivalence of numbers
#
# show
# $$
# (\mathbf{u}\mathbf{v}^T)\mathbf{x} = \mathbf{u}(\mathbf{v}^T\mathbf{x})
# $$
# + slideshow={"slide_type": "-"}
b = numpy.outer(u,v).dot(x)
c = u.dot(v.dot(x))
d = u*v.dot(x)
print('Max Difference = {}'.format(numpy.max(numpy.abs(b-c))))
print('Max Difference = {}'.format(numpy.max(numpy.abs(b-d))))
print('Max Difference = {}'.format(numpy.max(numpy.abs(c-d))))
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Check Timing/performance
# + slideshow={"slide_type": "-"}
# timing
print('(uv^T).dot(x)',end='\t\t')
# %timeit numpy.outer(u,v).dot(x)
print('\nu*v.dot(x)',end='\t\t')
# %timeit u*v.dot(x)
# + [markdown] slideshow={"slide_type": "skip"}
# #### Example: Upper Triangular Multiplication
#
# Consider the multiplication of a matrix $A \in \mathbb{C}^{m\times n}$ and the **upper-triangular** matrix $R$ defined as the $n \times n$ matrix with entries $r_{ij} = 1$ for $i \leq j$ and $r_{ij} = 0$ for $i > j$. The product can be written as
# $$
# \begin{bmatrix} \\ \\ \mathbf{b}_1 & \cdots & \mathbf{b}_n \\ \\ \\ \end{bmatrix} = \begin{bmatrix} \\ \\ \mathbf{a}_1 & \cdots & \mathbf{a}_n \\ \\ \\ \end{bmatrix} \begin{bmatrix} 1 & \cdots & 1 \\ & \ddots & \vdots \\ & & 1 \end{bmatrix}.
# $$
#
# The columns of $B$ are then
# $$
# \mathbf{b}_j = A \mathbf{r}_j = \sum^j_{k=1} \mathbf{a}_k
# $$
# so that $\mathbf{b}_j$ is the sum of the first $j$ columns of $A$.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Example: Write Matrix-Matrix Multiplication
#
# Write a function that computes matrix-matrix multiplication and demonstrate the following properties:
# 1. $A (B + C) = AB + AC$ (for square matrices))
# 1. $A (cB) = c AB$ where $c \in \mathbb{C}$
# 1. $AB \neq BA$ in general
# + hide_input=false slideshow={"slide_type": "skip"}
def matrix_matrix_product(A, B):
C = numpy.zeros((A.shape[0], B.shape[1]))
for i in range(A.shape[0]):
for j in range(B.shape[1]):
for k in range(A.shape[1]):
C[i, j] += A[i, k] * B[k, j]
return C
m = 4
n = 4
p = 4
A = numpy.random.uniform(size=(m, n))
B = numpy.random.uniform(size=(n, p))
C = numpy.random.uniform(size=(m, p))
c = numpy.random.uniform()
print(numpy.allclose(matrix_matrix_product(A, B), numpy.dot(A, B)))
print(numpy.allclose(matrix_matrix_product(A, (B + C)), matrix_matrix_product(A, B) + matrix_matrix_product(A, C)))
print(numpy.allclose(matrix_matrix_product(A, c * B), c*matrix_matrix_product(A, B)))
print(numpy.allclose(matrix_matrix_product(A, B), matrix_matrix_product(B, A)))
# + [markdown] slideshow={"slide_type": "subslide"}
# #### NumPy Products
#
# NumPy and SciPy contain routines that are optimized to perform matrix-vector and matrix-matrix multiplication. Given two `ndarray`s you can take their product by using the `dot` function.
# + slideshow={"slide_type": "subslide"}
n = 10
# Matrix vector with identity
I = numpy.identity(n)
x = numpy.random.random(n)
print(x)
# + slideshow={"slide_type": "fragment"}
print('x = Ix is {}\n'.format(numpy.allclose(x, numpy.dot(I, x))))
print('x - I.dot(x) = {}\n'.format(x-I.dot(x)))
print('I*x = \n{}\n'.format(I*x))
print(I.dot(x))
# + slideshow={"slide_type": "skip"}
# Matrix vector product
m = 5
A = numpy.random.random((m, n))
print(numpy.dot(A, x))
# + slideshow={"slide_type": "skip"}
# Matrix matrix product
B = numpy.random.random((n, m))
print(numpy.dot(A, B))
print()
print(A.dot(B))
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Check non-commutative property of matrix-matrix multiplication
#
# It's easy to demonstrate the non-commutative nature of Matrix-Matrix multiplication with *almost* any two matrices $A$ and $B$. For a clear demonstration, consider the two matrices $D$ a diagonal matrix and $A$ a matrix of all 1s
# + slideshow={"slide_type": "-"}
N=5
D = numpy.diag(numpy.array(range(1,N+1)))
A = numpy.ones((N,N))
print(D,'\n')
print(A)
# + slideshow={"slide_type": "fragment"}
print(D.dot(A))
# + slideshow={"slide_type": "fragment"}
print(A.dot(D))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Range and Null-Space
#
# #### Range
# - The **range** of a matrix $A \in \mathbb R^{m \times n}$ (similar to any function), denoted as $\text{range}(A)$, is the set of vectors that can be expressed as $A x$ for $x \in \mathbb R^n$.
# - We can also then say that that $\text{range}(A)$ is the space **spanned** by the columns of $A$. In other words the **linearly independent** columns of $A$ provide a basis for $\text{range}(A)$, also called the **column space** of the matrix $A$.
# - $C(A)$ controls the **existence** of solutions to $A\mathbf{x}=\mathbf{b}$
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Null-Space
# - Similarly the **null-space** of a matrix $A$, denoted $\text{null}(A)$ is the set of vectors $\mathbf{x}$ that satisfy $A \mathbf{x} = \mathbf{0}$.
# - $N(A)$ controls the **uniqueness** of solutions to $A\mathbf{x}=\mathbf{b}$
# - A similar concept is the **rank** of the matrix $A$, denoted as $\text{rank}(A)$, is the dimension of the column space. A matrix $A$ is said to have **full-rank** if $\text{rank}(A) = \min(m, n)$. This property also implies that the matrix mapping is **one-to-one**.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Inverse
#
# A **non-singular** or **invertible** matrix is characterized as a square matrix with full-rank. This is related to why we know that the matrix is one-to-one, we can use it to transform a vector $x$ and using the inverse, denoted $A^{-1}$, we can map it back to the original matrix. The familiar definition of this is
# \begin{align*}
# A \mathbf{x} &= \mathbf{b}, \\
# A^{-1} A \mathbf{x} & = A^{-1} \mathbf{b}, \\
# x &=A^{-1} \mathbf{b}.
# \end{align*}
# Since $A$ has full rank, its columns form a basis for $\mathbb{R}^m$ and the vector $\mathbf{b}$ must be in the column space of $A$.
# + [markdown] slideshow={"slide_type": "subslide"}
# There are a number of important properties of an invertible matrix $A$. Here we list them as the following equivalent statements
# 1. $A$ has a *unique* inverse $A^{-1}$ such that $A^{-1}A=AA^{-1}=I$
# 1. $\text{rank}(A) = m$
# 1. $\text{range}(A) = \mathbb{C}^m$
# 1. $\text{null}(A) = \mathbf{0}$
# 1. 0 is not an eigenvalue of $A$
# 1. $\text{det}(A) \neq 0$
# + [markdown] slideshow={"slide_type": "skip"}
# #### Example: Properties of invertible matrices
#
# Show that given an invertible matrix that the rest of the properties hold. Make sure to search the `numpy` packages for relevant functions.
# + slideshow={"slide_type": "skip"}
m = 5
# generate a random m x m invertible matrix
for n in range(100):
A = numpy.random.uniform(size=(m, m))
if numpy.linalg.det(A) != 0:
break
print('A^{{-1}}*A = \n\n{}\n'.format(numpy.dot(numpy.linalg.inv(A), A)))
print('rank(A) = {}\n'.format(numpy.linalg.matrix_rank(A)))
print("N(A)= {}\n".format(numpy.linalg.solve(A, numpy.zeros(m))))
print("Eigenvalues = {}".format(numpy.linalg.eigvals(A)))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Orthogonal Vectors and Matrices
#
# Orthogonality is a very important concept in linear algebra that forms the basis of many of the modern methods used in numerical computations.
# + [markdown] slideshow={"slide_type": "subslide"}
# Two vectors are said to be *orthogonal* if their **inner-product** or **dot-product** defined as
# $$
# < \mathbf{x}, \mathbf{y} > \equiv (\mathbf{x}, \mathbf{y}) \equiv \mathbf{x}^T\mathbf{y} \equiv \mathbf{x} \cdot \mathbf{y} = \sum^m_{i=1} x_i y_i = 0
# $$
# Here we have shown the various notations you may run into (the inner-product is in-fact a general term for a similar operation for mathematical objects such as functions).
# + [markdown] slideshow={"slide_type": "subslide"}
# If $\langle \mathbf{x},\mathbf{y} \rangle = 0$ then we say $\mathbf{x}$ and $\mathbf{y}$ are orthogonal. The reason we use this terminology is that the inner-product of two vectors can also be written in terms of the angle between them where
#
# $$
# \cos \theta = \frac{\langle \mathbf{x}, \mathbf{y} \rangle}{||\mathbf{x}||_2~||\mathbf{y}||_2}
# $$
#
# and $||\mathbf{x}||_2$ is the Euclidean ($\ell^2$) norm of the vector $\mathbf{x}$, which we can interpret as the *length* of a vector.
# + [markdown] slideshow={"slide_type": "subslide"}
# We can write the 2-norm or length of a vector in terms of the inner-product as well as
# $$
# ||\mathbf{x}||_2^2 = \langle \mathbf{x}, \mathbf{x} \rangle = \mathbf{x}^T\mathbf{x} = \sum^m_{i=1} |x_i|^2.
# $$
#
# $$
# ||\mathbf{x}||_2 = \sqrt{\langle \mathbf{x}, \mathbf{x} \rangle}
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# The generalization of the inner-product to complex vector spaces is defined as
# $$
# \langle x, y \rangle = \sum^m_{i=1} x_i^* y_i
# $$
# where $x_i^*$ is the complex-conjugate of the value $x_i$.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Orthonormality
#
# Taking this idea one step further we can say a set of vectors $\mathbf{x} \in X$ are orthogonal to $\mathbf{y} \in Y$ if
#
# $$
# < \mathbf{x}, \mathbf{y} > = 0 \quad \forall \mathbf{x},\mathbf{y},
# $$
#
# If in addition
# $$||\mathbf{x}|| = 1,\, ||\mathbf{y}|| = 1\quad \forall \mathbf{x},\mathbf{y}
# $$
#
# then they are also called orthonormal.
#
# Note that we dropped the 2 as a subscript to the notation for the norm of a vector. Later we will explore other ways to define a norm of a vector other than the Euclidean norm defined above.
# + [markdown] slideshow={"slide_type": "subslide"}
# Another concept that is related to orthogonality is linear-independence. A set of vectors $\mathbf{x} \in X$ are **linearly independent** if $\forall \mathbf{x} \in X$ that each $\mathbf{x}$ cannot be written as a linear combination of the other vectors in the set $X$.
#
#
#
# An equivalent statement is that given a set of $n$ vectors $\mathbf{x}_i$, the only set of scalars $c_i$ that satisfies
# $$
# \sum_{i=1}^n c_i\mathbf{x}_i = \mathbf{0}
# $$
# is if $c_i=0$ for all $i\in[1,n]$
#
# + [markdown] slideshow={"slide_type": "fragment"}
# Show that if a set of vectors are orthonormal, they must be linearly independent.
# + [markdown] slideshow={"slide_type": "skip"}
# This can be related directly through the idea of projection. If we have a set of vectors $\mathbf{x} \in X$ we can project another vector $\mathbf{v}$ onto the vectors in $X$ by using the inner-product. This is especially powerful if we have a set of **orthogonal** vectors $X$, which are said to **span** a space (or provide a **basis** for a space), s.t. any vector in the space spanned by $X$ can be expressed as a linear combination of the basis vectors $X$
# $$
# \mathbf{v} = \sum^n_{i=1} \, \langle \mathbf{v}, \mathbf{x}_i \rangle \, \mathbf{x}_i.
# $$
# Note if $\mathbf{v} \in X$ that
# $$
# \langle \mathbf{v}, \mathbf{x}_i \rangle = 0 \quad \forall \mathbf{x}_i \in X \setminus \mathbf{v}.
# $$
#
# Looping back to matrices, the column space of a matrix is spanned by its linearly independent columns. Any vector $v$ in the column space can therefore be expressed via the equation above.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Unitary Matrices
#
# A special class of matrices are called **unitary** matrices when complex-valued and **orthogonal** when purely real-valued if the columns of the matrix are *orthonormal* to each other. Importantly this implies that for a unitary matrix $Q$ we know the following
#
# 1. $Q^*Q = I$
# 1. therefore if $Q$ is square, $Q^* = Q^{-1}$
#
# where $Q^*$ is called the **adjoint** (or Hermitian) of $Q$. The adjoint is defined as the transpose of the original matrix with the entries being the complex conjugate of each entry as the notation implies. Note, if $Q\in\mathbb{R}^{n\times n}$ then $Q^*=Q^T$
# + [markdown] slideshow={"slide_type": "subslide"}
# As an example if we have the matrix
# $$
# \begin{aligned}
# Q &= \begin{bmatrix} q_{11} & q_{12} \\ q_{21} & q_{22} \\ q_{31} & q_{32} \end{bmatrix} \quad \text{then} \\
# Q^* &= \begin{bmatrix} q^*_{11} & q^*_{21} & q^*_{31} \\ q^*_{12} & q^*_{22} & q^*_{32} \end{bmatrix}
# \end{aligned}
# $$
#
# The important part of being an unitary matrix is that the projection onto the column space of the matrix $Q$ preserves geometry in an Euclidean sense, i.e. preserves the Cartesian distance.
# + [markdown] hide_input=true slideshow={"slide_type": "slide"}
# # Vector and Matrix Norms and the condition number
#
# The following sections will lay out the definitions and computation required to calculate a range of vector and matrix norms which provide a measure of the "size" of an object or the distance in some vector space.
#
# In the context of Numerical Linear algebra, norms are essential for defining a key property of a matrix, the "condition number".
#
# In infinite precision, a square matrix is either invertible or singular, however, in finite precision, even an invertible matrix can behave poorly if it is 'ill-conditioned', i.e. it is almost singular.
#
# We need a quantitiative measure of how "near-singular" a matrix is
# + [markdown] hide_input=true slideshow={"slide_type": "subslide"}
# ### Why not the Determinant $|A|$?
#
# We know that if a matrix $A$ is singular, $|A|=0$. But what if $|A|$ is small?
#
# Turns out even a perfectly invertible, well-conditioned matrix can have arbitrarily small determinant.
# + [markdown] hide_input=true slideshow={"slide_type": "fragment"}
# Consider the two diagonal matrices
#
# $$
# I = \begin{bmatrix} 1 & & \\ & 1 & \\ & & 1\\ \end{bmatrix}\quad\text{and}\quad
# \epsilon I = \begin{bmatrix} \epsilon & & \\ & \epsilon & \\ & & \epsilon \\ \end{bmatrix}
# $$
#
# by definition, $|I|=1$, but $|\epsilon I| = ??$.
# + [markdown] hide_input=true slideshow={"slide_type": "fragment"}
# More generally if $I\in\mathbb{R}^{n\times n}$, $|\epsilon I| = ??$.
#
# Yet all these matrices are diagonal and easily inverted (i.e. $(\epsilon I )^{-1} = (1/\epsilon) I$). Thus we need something better (the condition number). But to get there requires developing important ideas about vector and matrix norms.
# + [markdown] hide_input=true slideshow={"slide_type": "slide"}
# ### Vector Norms
#
# Norms (and also measures) provide a means for measure the "size" or distance in a space. In general a norm is a function, denoted by $||\cdot||$, that maps $\mathbb{C}^m \rightarrow \mathbb{R}$. In other words we stick in a multi-valued object and get a single, real-valued number out the other end.
#
# All norms satisfy the properties:
#
# 1. $||\mathbf{x}|| \geq 0$
# 2. $||\mathbf{x}|| = 0$ only if $\mathbf{x} = \mathbf{0}$
# 3. $||\mathbf{x} + \mathbf{y}||\leq ||\mathbf{x}|| + ||\mathbf{y}||$ (triangle inequality)
# 4. $||c \mathbf{x}|| = |c|||\mathbf{x}||$ where $c \in \mathbb{C}$
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# There are a number of relevant norms that we can define beyond the Euclidean norm, also know as the 2-norm or $\ell_2$ norm:
#
# * $\ell_1$ norm:
# $$
# ||\mathbf{x}||_1 = \sum^m_{i=1} |x_i|,
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# * $\ell_2$ norm:
# $$
# ||\mathbf{x}||_2 = \left( \sum^m_{i=1} |x_i|^2 \right)^{1/2},
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# * $\ell_p$ norm:
# $$
# ||\mathbf{x}||_p = \left( \sum^m_{i=1} |x_i|^p \right)^{1/p}, \quad \quad 1 \leq p < \infty,
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# * $\ell_\infty$ norm:
# $$
# ||\mathbf{x}||_\infty = \max_{1\leq i \leq m} |x_i|,
# $$
# + [markdown] slideshow={"slide_type": "skip"}
# 1. weighted $\ell_p$ norm:
# $$
# ||\mathbf{x}||_{W_p} = \left( \sum^m_{i=1} |w_i x_i|^p \right)^{1/p}, \quad \quad 1 \leq p < \infty,
# $$
#
# These are also related to other norms denoted by capital letters ($L_2$ for instance). In this case we use the lower-case notation to denote finite or discrete versions of the infinite dimensional counterparts.
# + [markdown] slideshow={"slide_type": "skip"}
# #### Example: Comparisons Between Norms
#
# Compute the norms given some vector $\mathbf{x}$ and compare their values. Verify the properties of the norm for one of the norms.
# + slideshow={"slide_type": "skip"}
def pnorm(x, p):
""" return the vector p norm of a vector
parameters:
-----------
x: numpy array
vector
p: float or numpy.inf
value of p norm such that ||x||_p = (sum(|x_i|^p))^{1/p} for p< inf
for infinity norm return max(abs(x))
returns:
--------
pnorm: float
pnorm of x
"""
if p == numpy.inf:
norm = numpy.max(numpy.abs(x))
else:
norm = numpy.sum(numpy.abs(x)**p)**(1./p)
return norm
# + slideshow={"slide_type": "skip"}
m = 10
p = 4
x = numpy.random.uniform(size=m)
ell_1 = pnorm(x, 1)
ell_2 = pnorm(x, 2)
ell_p = pnorm(x, p)
ell_infty = pnorm(x, numpy.inf)
print('x = {}'.format(x))
print()
print("L_1 = {}\nL_2 = {}\nL_{} = {}\nL_inf = {}".format(ell_1, ell_2, p, ell_p, ell_infty))
# + slideshow={"slide_type": "skip"}
y = numpy.random.uniform(size=m)
print()
print("Properties of norms:")
print('x = {}'.format(x))
print('y = {}\n'.format(y))
p = 2
print('||x+y||_{p} = {nxy}\n||x||_{p} + ||y||_{p} = {nxny}'.format(
p=p,nxy=pnorm(x+y, p), nxny=pnorm(x, p) + pnorm(y, p)))
c = -0.1
print('||c x||_{} = {}'.format(p,pnorm(c * x, p)))
print('|c||x||_{} = {}'.format(p,abs(c) * pnorm(x, p)))
# + [markdown] slideshow={"slide_type": "slide"}
# #### Geometric Interpretation
#
# For every $p-$norm we can define a set of 'unit vectors' that is the set of all vectors in $\mathbb{R}^n$ with $||\mathbf{x}||_p = 1$.
#
# For example in $\mathbb{R}^2$, the unit spheres in the 1-,2- and $\infty$-norm look like
# + hide_input=true slideshow={"slide_type": "-"}
import matplotlib.patches as patches
# Note: that this code is a bit fragile to angles that go beyond pi
# due to the use of arccos.
head_width = 0.1
head_length = 1.5 * head_width
def draw_unit_vectors(axes, A, head_width=0.1):
head_length = 1.5 * head_width
image_e = numpy.empty(A.shape)
angle = numpy.empty(A.shape[0])
image_e[:, 0] = numpy.dot(A, numpy.array((1.0, 0.0)))
image_e[:, 1] = numpy.dot(A, numpy.array((0.0, 1.0)))
for i in range(A.shape[0]):
angle[i] = numpy.arccos(image_e[0, i] / numpy.linalg.norm(image_e[:, i], ord=2))
axes.arrow(0.0, 0.0, image_e[0, i] - head_length * numpy.cos(angle[i]),
image_e[1, i] - head_length * numpy.sin(angle[i]),
head_width=head_width, color='b', alpha=0.5)
# comparison of norms
# ============
# 1-norm
# Unit-ball
fig = plt.figure(figsize=(8,6))
#fig.suptitle("1-Norm: $||A||_1 = {}$".format(numpy.linalg.norm(A,ord=1)), fontsize=16)
theta=numpy.linspace(0., 2.*numpy.pi,100)
axes = fig.add_subplot(1, 1, 1, aspect='equal')
axes.plot((1.0, 0.0, -1.0, 0.0, 1.0), (0.0, 1.0, 0.0, -1.0, 0.0), 'r', label='$||\mathbf{x}||_1=1$')
axes.plot(numpy.cos(theta),numpy.sin(theta), 'g', label='$||\mathbf{x}||_2=1$')
axes.plot((1.0, -1.0, -1.0, 1.0, 1.0), (1.0, 1.0, -1.0, -1.0, 1.0), 'b', label='$||\mathbf{x}||_\infty=1$')
draw_unit_vectors(axes, numpy.eye(2))
axes.arrow(0.0, 0.0, 1.0 - head_length * numpy.cos(numpy.pi/4.),
1.0 - head_length * numpy.sin(numpy.pi/4.),
head_width=head_width, color='k', linestyle='--', alpha=0.5)
axes.set_title("Unit Ball in 1-,2-,and $\infty$ norm")
axes.set_xlim((-1.1, 1.1))
axes.set_ylim((-1.1, 1.1))
axes.grid(True)
#axes.legend([ '$||\mathbf{x}||_1=1$', '$||\mathbf{x}||_2=1$', '$||\mathbf{x}||_\infty=1$'], loc='best')
axes.legend(loc='upper center', bbox_to_anchor = (0.5,0.5))
plt.show()
# + [markdown] slideshow={"slide_type": "fragment"}
# And if $\mathbf{x} = \begin{bmatrix} 1 \\ 1 \\ \end{bmatrix}$ then $||\mathbf{x}||_1 = ??,~~$
# $||\mathbf{x}||_2 = ??,~~$ and $||\mathbf{x}||_\infty = ??$
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Induced Matrix Norms
#
# The most direct way to consider a matrix norm is one induced by a vector-norm. Given a vector norm, we can define a matrix p-norm as the smallest number $C$ that satisfies the inequality
# $$
# ||A \mathbf{x}||_{p} \leq C ||\mathbf{x}||_{p} \quad\forall\quad \mathbf{x}\in\mathbb{C}^n
# $$
# or as the supremum of the ratios so that
# $$
# C = ||A||_p = \sup_{\mathbf{x}\in\mathbb{C}^n ~ \mathbf{x}\neq\mathbf{0}} \frac{||A \mathbf{x}||_{p}}{||\mathbf{x}||_p}.
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# with no loss of generality, we can restrict $\mathbf{x}$ to the set of all unit vectors $||\mathbf{x}||_p=1$ and interpret the matrix p-norm as the maximum distortion of the "unit sphere"
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Properties of all Matrix Norms (induced and non-induced)
#
# In general matrix-norms have the following properties whether they are induced from a vector-norm or not:
# 1. $||A|| \geq 0$ and $||A|| = 0$ only if $A = 0$
# 1. $||A + B|| \leq ||A|| + ||B||$ (Triangle Inequality)
# 1. $||c A|| = |c| ||A||$
# + [markdown] slideshow={"slide_type": "fragment"}
# In addition, all induced p-norms satisfy the product rules
# 1. $||AB|| \leq ||A||\,||B||$
# 1. $||A\mathbf{x}|| \leq ||A||\,||\mathbf{x}||$
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Computation of induced matrix p-norms
#
# With a little work it can be shown that
#
# 1. $||A||_1$ = `numpy.linalg.norm(A, ord=1)` is the maximum 1-norm of the Columns of $A$
# 1. $||A||_2$ = `numpy.linalg.norm(A, ord=2) = max(numpy.linalg.svd(A)[1]` is the maximum Singular Value of $A$
# 1. $||A||_\infty$ = `numpy.linalg.norm(A, ord=numpy.inf) ` is the maximum 1-norm of the Rows of $A$ (or $||A^T||_1$)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Examples
#
# #### The Identity Matrix $||I||$
# $$
# I = \begin{bmatrix} 1 & & \\ & 1 & \\ & & 1\\ \end{bmatrix}
# $$
#
# $||I||_1=||I||_2=||I||_\infty = ??$
# + slideshow={"slide_type": "fragment"}
for ord in [1,2,numpy.inf]:
print('||I||_{} = {}'.format(ord,numpy.linalg.norm(I,ord=ord)))
# + [markdown] slideshow={"slide_type": "subslide"}
#
# #### Diagonal Matrices $||D||$
# $$
# D = \begin{bmatrix} d_1 & & \\ & d_2 & \\ & & d_3\\ \end{bmatrix} \quad\text{e.g.}\quad
# \begin{bmatrix} 2 & & \\ & 1 & \\ & & -3\\ \end{bmatrix}
# $$
#
# $||D||_1=|D||_2=||D||_\infty = ??$
# + slideshow={"slide_type": "fragment"}
D = numpy.diag([2,1,-3])
for ord in [1,2,numpy.inf]:
print('||D||_{} = {}'.format(ord,numpy.linalg.norm(D,ord=ord)))
# + [markdown] slideshow={"slide_type": "subslide"}
#
# #### Orthogonal Matrices $||Q||_2$ where $Q^*Q=I$
#
# A fundamental property of $Q$ matrices is that they do not change the length of vectors i.e.
# $$
# ||Q\mathbf{x}||_2^2 = \mathbf{x}^*Q^*Q\mathbf{x} = \mathbf{x}^*\mathbf{x} = ||\mathbf{x}||_2^2
# $$
#
# Therefore
# $$
# ||Q||_2 = \sup_{\mathbf{x}\in\mathbb{C}^n, ||\mathbf{x}||_2=1} ||Q\mathbf{x}||_2 = ??
# $$
# + [markdown] slideshow={"slide_type": "skip"}
# #### Example: Induced Matrix Norms
#
# Consider the matrix
# $$
# A = \begin{bmatrix} 1 & 2 \\ 0 & 2 \end{bmatrix}.
# $$
# Compute the induced-matrix norm of $A$ for the vector norms $\ell_2$ and $\ell_\infty$.
# + [markdown] slideshow={"slide_type": "skip"}
# $\ell^2$: For both of the requested norms the unit-length vectors $[1, 0]$ and $[0, 1]$ can be used to give an idea of what the norm might be and provide a lower bound.
#
# $$
# ||A||_2 = \sup_{x \in \mathbb{R}^n} \left( ||A \cdot [1, 0]^T||_2, ||A \cdot [0, 1]^T||_2 \right )
# $$
#
# computing each of the norms we have
#
# $$\begin{aligned}
# \begin{bmatrix} 1 & 2 \\ 0 & 2 \end{bmatrix} \cdot \begin{bmatrix} 1 \\ 0 \end{bmatrix} &= \begin{bmatrix} 1 \\ 0 \end{bmatrix} \\
# \begin{bmatrix} 1 & 2 \\ 0 & 2 \end{bmatrix} \cdot \begin{bmatrix} 0 \\ 1 \end{bmatrix} &= \begin{bmatrix} 2 \\ 2 \end{bmatrix}
# \end{aligned}$$
#
# which translates into the norms $||A \cdot [1, 0]^T||_2 = 1$ and $||A \cdot [0, 1]^T||_2 = 2 \sqrt{2}$. This implies that the $\ell_2$ induced matrix norm of $A$ is at least $||A||_{2} = 2 \sqrt{2} \approx 2.828427125$.
# + [markdown] slideshow={"slide_type": "skip"}
# The exact value of $||A||_2$ can be computed using the spectral radius defined as
# $$
# \rho(A) = \max_{i} |\lambda_i|,
# $$
# where $\lambda_i$ are the eigenvalues of $A$. With this we can compute the $\ell_2$ norm of $A$ as
# $$
# ||A||_2 = \sqrt{\rho(A^\ast A)}
# $$
#
# Computing the norm again here we find
# $$
# A^\ast A = \begin{bmatrix} 1 & 0 \\ 2 & 2 \end{bmatrix} \begin{bmatrix} 1 & 2 \\ 0 & 2 \end{bmatrix} = \begin{bmatrix} 1 & 2 \\ 2 & 8 \end{bmatrix}
# $$
# which has eigenvalues
# $$
# \lambda = \frac{1}{2}\left(9 \pm \sqrt{65}\right )
# $$
# so $||A||_2 \approx 2.9208096$.
# + [markdown] slideshow={"slide_type": "skip"}
# The actual induced 2-norm of a matrix can be derived using the Singular Value Decomposition (SVD) and is simply the largest singular value $\sigma_1$.
#
# **Proof**:
# Given that every Matrix $A\in\mathbb{C}^{m\times n}$ can be factored into its SVD (see notebook 10.1):
#
# $$
# A = U\Sigma V^*
# $$
#
# where $U\in\mathbb{C}^{m\times m}$ and $V\in\mathbb{C}^{n\times n}$ are unitary matrices with the property $U^*U=I$ and $V^*V=I$ (of their respective sizes) and $\Sigma$ is a real diagonal matrix of singular values $\sigma_1 \geq\sigma_2\geq...\sigma_n\geq 0$.
# + [markdown] slideshow={"slide_type": "skip"}
# Then the 2-norm squared of a square matrix is
# $$
# ||A||^2_2 = \sup_{\mathbf{x} \in \mathbb{C}^n ~ ||\mathbf{x}||_2 = 1} ||A \mathbf{x}||_2^2 = \mathbf{x}^TA^*A\mathbf{x}
# $$
# but $A^*A = V\Sigma^2V^*$ so
#
# \begin{align}
# ||A \mathbf{x}||_2^2 &= \mathbf{x}^*V\Sigma^2V^*\mathbf{x} \\
# &= \mathbf{y}^*\Sigma^2\mathbf{y} \quad\mathrm{where}\quad \mathbf{y}=V^*\mathbf{x}\\
# &= \sum_{i=1}^n \sigma_i^2|y_i|^2\\
# &\leq \sigma_1^2\sum_{i=1}^n |y_i|^2 = \sigma_i^2||\mathbf{y}||_2\\
# \end{align}
#
# but if $||\mathbf{x}||_2 = 1$ (i.e. is a unit vector), then so is $\mathbf{y}$ because unitary matrices don't change the length of vectors. So it follows that
# $$
# ||A||_2 = \sigma_1
# $$
#
# + slideshow={"slide_type": "skip"}
A = numpy.array([[1, 2], [0, 2]])
#calculate the SVD(A)
U, S, Vt = numpy.linalg.svd(A)
print('Singular_values = {}'.format(S))
print('||A||_2 = {}'.format(S.max()))
print('||A||_2 = {}'.format(numpy.linalg.norm(A, ord=2)))
# more fun facts about the SVD
#print(U.T.dot(U))
#print(Vt.T.dot(Vt))
#print(A - numpy.dot(U,numpy.dot(numpy.diag(S),Vt)))
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Some Quick proofs
#
# The induced 1-norm is the max of the 1-norm of the **columns** of $A$
#
# Given
# $$
# A\mathbf{x} = x_1\mathbf{a}_1 + x_2\mathbf{a}_2 + \ldots + x_n\mathbf{a}_n
# $$
#
# where $||\mathbf{x}||_1 = 1$. Then
# + [markdown] slideshow={"slide_type": "fragment"}
# $$
# \begin{align}
# ||A \mathbf{x}||_1 &= || x_1\mathbf{a}_1 + x_2\mathbf{a}_2 + \ldots + x_n\mathbf{a}_n || \\
# &\leq |x_1|\,||\mathbf{a}_1||_1 + |x_2|\,||\mathbf{a}_2||_1 + \ldots + |x_n|\,||\mathbf{a}_n||_1 \quad\text{(triangle rule)}\\
# &\leq \max_{1\leq j\leq n} ||\mathbf{a}_j||_1\sum_{j=1}^n |x_j| = \max_{1\leq j\leq n} ||\mathbf{a}_j||_1 ||\mathbf{x}||_1\\
# &= \max_{1\leq j\leq n} ||\mathbf{a}_j||_1\\
# \end{align}
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# The induced 2-norm is the maximum singular value (short version...slightly scrappy)
#
# Given
# $$
# A = U\Sigma V^T
# $$
#
# where $U^TU = V^TV = I$ and $\Sigma$ is a diagonal matrix with diagonal entries $\sigma_1\geq\sigma_2\geq\ldots\geq\sigma_r>0$
# + [markdown] slideshow={"slide_type": "fragment"}
# then
# $$
# \begin{align}
# AV &= U\Sigma\\
# ||AV||_2 &= ||U\Sigma||_2\\
# ||A||_2||V||_2 &= ||U||_2||\Sigma||_2\\
# ||A||_2 &\leq ||\Sigma||_2 = \sigma_1 \\
# \end{align}
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# The induce $\infty$-norm is the max of the 1-norm of **rows** of $A$
#
# $$
# ||A \mathbf{x}||_\infty = \max_{1 \leq i \leq m} | \mathbf{a}^*_i \mathbf{x} | \leq \max_{1 \leq i \leq m} ||\mathbf{a}^*_i||_1
# $$
# because the largest unit vector on the unit sphere in the $\infty$ norm is a vector of 1's.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Example:
#
# $$
# A = \begin{bmatrix} 1 & 2 \\ 0 & 2 \end{bmatrix}.
# $$
#
# $$ ||A||_1 = 4, \quad ||A||_\infty = 3$$
#
# + slideshow={"slide_type": "fragment"}
# Calculate the 1-norm of A
A = numpy.array([ [ 1, 2], [ 0, 2]])
normA_1 = numpy.max(numpy.sum(numpy.abs(A), axis=0))
print('||A||_1 = {}'.format(normA_1))
print('||A||_1 = {}'.format(numpy.linalg.norm(A, ord=1)))
# + slideshow={"slide_type": "fragment"}
# calculate the 2 norm of A
normA_2 = numpy.max(numpy.linalg.svd(A, compute_uv=False))
print('||A||_2 = {}'.format(normA_2))
print('||A||_2 = {}'.format(numpy.linalg.norm(A, ord=2)))
print()
U,S, V = numpy.linalg.svd(A)
print(U.dot(numpy.diag(S).dot(V)))
# + slideshow={"slide_type": "fragment"}
# calculate the infinity norm of A
normA_inf = numpy.max(numpy.sum(numpy.abs(A), axis=1))
print('||A||_inf = {}'.format(normA_inf))
print('||A||_inf = {}'.format(numpy.linalg.norm(A, ord=numpy.inf)))
# + [markdown] slideshow={"slide_type": "subslide"}
#
# ### The Geometric Picture
#
# One of the most useful ways to think about matrix norms is as a transformation of a unit-ball to an ellipse. Depending on the norm in question, the norm will be some combination of the resulting ellipse.
#
# + [markdown] hide_input=false slideshow={"slide_type": "subslide"}
# #### 1-Norm: $A = \begin{bmatrix} 1 & 2 \\ 0 & 2 \\ \end{bmatrix}$
# + hide_input=true slideshow={"slide_type": "-"}
A = numpy.array([[1, 2], [0, 2]])
#============
# 1-norm
# Unit-ball
fig = plt.figure()
fig.set_figwidth(fig.get_figwidth() * 2)
fig.suptitle("1-Norm: $||A||_1 = {}$".format(numpy.linalg.norm(A,ord=1)), fontsize=16)
axes = fig.add_subplot(1, 2, 1, aspect='equal')
axes.plot((1.0, 0.0, -1.0, 0.0, 1.0), (0.0, 1.0, 0.0, -1.0, 0.0), 'r')
draw_unit_vectors(axes, numpy.eye(2))
axes.set_title("Unit Ball")
axes.set_xlim((-1.1, 1.1))
axes.set_ylim((-1.1, 1.1))
axes.grid(True)
# Image
axes = fig.add_subplot(1, 2, 2, aspect='equal')
axes.plot((1.0, 2.0, -1.0, -2.0, 1.0), (0.0, 2.0, 0.0, -2.0, 0.0), 'r')
draw_unit_vectors(axes, A, head_width=0.2)
axes.set_title("Images Under A")
axes.grid(True)
plt.show()
# + [markdown] slideshow={"slide_type": "subslide"}
# #### 2-Norm: $A = \begin{bmatrix} 1 & 2 \\ 0 & 2 \\ \end{bmatrix}$
# + hide_input=true slideshow={"slide_type": "-"}
# ============
# 2-norm
# Unit-ball
fig = plt.figure()
fig.suptitle("2-Norm: $||A||_2 = ${:3.4f}".format(numpy.linalg.norm(A,ord=2)),fontsize=16)
fig.set_figwidth(fig.get_figwidth() * 2)
axes = fig.add_subplot(1, 2, 1, aspect='equal')
axes.add_artist(plt.Circle((0.0, 0.0), 1.0, edgecolor='r', facecolor='none'))
draw_unit_vectors(axes, numpy.eye(2))
axes.set_title("Unit Ball")
axes.set_xlim((-1.1, 1.1))
axes.set_ylim((-1.1, 1.1))
axes.grid(True)
# Image
# Compute some geometry
u, s, v = numpy.linalg.svd(A)
theta = numpy.empty(A.shape[0])
ellipse_axes = numpy.empty(A.shape)
theta[0] = numpy.arccos(u[0][0]) / numpy.linalg.norm(u[0], ord=2)
theta[1] = theta[0] - numpy.pi / 2.0
for i in range(theta.shape[0]):
ellipse_axes[0, i] = s[i] * numpy.cos(theta[i])
ellipse_axes[1, i] = s[i] * numpy.sin(theta[i])
axes = fig.add_subplot(1, 2, 2, aspect='equal')
axes.add_artist(patches.Ellipse((0.0, 0.0), 2 * s[0], 2 * s[1], theta[0] * 180.0 / numpy.pi,
edgecolor='r', facecolor='none'))
for i in range(A.shape[0]):
axes.arrow(0.0, 0.0, ellipse_axes[0, i] - head_length * numpy.cos(theta[i]),
ellipse_axes[1, i] - head_length * numpy.sin(theta[i]),
head_width=head_width, color='k')
draw_unit_vectors(axes, A, head_width=0.2)
axes.set_title("Images Under A")
axes.set_xlim((-s[0] + 0.1, s[0] + 0.1))
axes.set_ylim((-s[0] + 0.1, s[0] + 0.1))
axes.grid(True)
plt.show()
# + [markdown] slideshow={"slide_type": "subslide"}
# #### $\infty$-Norm: $A = \begin{bmatrix} 1 & 2 \\ 0 & 2 \\ \end{bmatrix}$
# + hide_input=true slideshow={"slide_type": "-"}
# ============
# infty-norm
# Unit-ball
fig = plt.figure()
fig.suptitle("$\infty$-Norm: $||A||_\infty = {}$".format(numpy.linalg.norm(A,ord=numpy.inf)),fontsize=16)
fig.set_figwidth(fig.get_figwidth() * 2)
axes = fig.add_subplot(1, 2, 1, aspect='equal')
axes.plot((1.0, -1.0, -1.0, 1.0, 1.0), (1.0, 1.0, -1.0, -1.0, 1.0), 'r')
draw_unit_vectors(axes, numpy.eye(2))
axes.set_title("Unit Ball")
axes.set_xlim((-1.1, 1.1))
axes.set_ylim((-1.1, 1.1))
axes.grid(True)
# Image
# Geometry - Corners are A * ((1, 1), (1, -1), (-1, 1), (-1, -1))
# Symmetry implies we only need two. Here we just plot two
u = numpy.empty(A.shape)
u[:, 0] = numpy.dot(A, numpy.array((1.0, 1.0)))
u[:, 1] = numpy.dot(A, numpy.array((-1.0, 1.0)))
theta[0] = numpy.arccos(u[0, 0] / numpy.linalg.norm(u[:, 0], ord=2))
theta[1] = numpy.arccos(u[0, 1] / numpy.linalg.norm(u[:, 1], ord=2))
axes = fig.add_subplot(1, 2, 2, aspect='equal')
axes.plot((3, 1, -3, -1, 3), (2, 2, -2, -2, 2), 'r')
for i in range(A.shape[0]):
axes.arrow(0.0, 0.0, u[0, i] - head_length * numpy.cos(theta[i]),
u[1, i] - head_length * numpy.sin(theta[i]),
head_width=head_width, color='k')
draw_unit_vectors(axes, A, head_width=0.2)
axes.set_title("Images Under A")
axes.set_xlim((-4.1, 4.1))
axes.set_ylim((-3.1, 3.1))
axes.grid(True)
plt.show()
# + [markdown] slideshow={"slide_type": "skip"}
# #### Cauchy-Schwarz and Hölder Inequalities
#
# Computing matrix norms where $p \neq 1$ or $\infty$ is more difficult unfortunately. We have a couple of tools that can be useful however.
#
# - **Cauchy-Schwarz Inequality**: For the special case where $p=q=2$, for any vectors $\mathbf{x}$ and $\mathbf{y}$
# $$
# |\mathbf{x}^*\mathbf{y}| \leq ||\mathbf{x}||_2 ||\mathbf{y}||_2
# $$
# - **Hölder's Inequality**: Turns out this holds in general if given a $p$ and $q$ that satisfy $1/p + 1/q = 1$ with $1 \leq p, q \leq \infty$
#
# $$
# |\mathbf{x}^*\mathbf{y}| \leq ||\mathbf{x}||_p ||\mathbf{y}||_q.
# $$
#
# **Note**: this is essentially what we used in the proof of the $\infty-$norm with $p=1$ and $q=\infty$
# + [markdown] slideshow={"slide_type": "subslide"}
# The most widely used matrix norm not induced by a vector norm is the **Frobenius norm** defined by
# $$
# ||A||_F = \left( \sum^m_{i=1} \sum^n_{j=1} |A_{ij}|^2 \right)^{1/2}.
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Invariance under unitary multiplication
#
# One important property of the matrix 2-norm (and Frobenius norm) is that multiplication by a unitary matrix does not change the product (kind of like multiplication by 1). In general for any $A \in \mathbb{C}^{m\times n}$ and unitary matrix $Q \in \mathbb{C}^{m \times m}$ we have
# \begin{align*}
# ||Q A||_2 &= ||A||_2 \\ ||Q A||_F &= ||A||_F.
# \end{align*}
# + [markdown] slideshow={"slide_type": "skip"}
# <sup>1</sup><span id="footnoteRegression"> http://www.utstat.toronto.edu/~brunner/books/LinearModelsInStatistics.pdf</span>
# + [markdown] hide_input=true slideshow={"slide_type": "slide"}
# # The condition number
#
# Finally we have enough machinery to define the condition number $\kappa(A)$ (or often $\mathrm{cond}(A)$) which is simply
#
# $$
# \kappa(A) = ||A||\,||A^{-1}|| \in [1,\infty)
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# ### Examples for simple matrices
#
# * Identity Matrix
# $$\kappa(I) = ||I||\,||I^{-1}|| = ??$$
# + [markdown] slideshow={"slide_type": "fragment"}
# * Diagonal Matrix
# $$\kappa(D) = ||D||\,||D^{-1}|| = ??$$
# + [markdown] slideshow={"slide_type": "fragment"}
# * 2-norm condition number of a matrix
# $$\kappa_2(A) = ||A||_2\,||A^{-1}||_2 = \frac{\sigma_1}{\sigma_n}$$
#
# if $A$ is singular, $\kappa_2(A) = ??$
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Back to our initial example
#
# Let
# $$
# A = \epsilon I \in \mathbb{R}^{n\times n}
# $$
#
# * The Determinant $|A| =\epsilon^n$ (which can be arbitrarily small)
# * But
# $$
# \begin{align}
# \kappa(A) &= ||A||\,||A^{-1}|| \\
# &= ||\epsilon I||\,||(1/\epsilon)I|| \\
# & \leq |\epsilon||1/\epsilon|||I||^2 = 1\\
# \end{align}
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# More generally, it's easy to show that scaling of a matrix does not change its condition number
#
# $$
# \kappa(\alpha A) = ||\alpha A||\,||(1/\alpha)A^{-1}|| =\kappa(A)
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Example
#
# $$
# A = \begin{bmatrix} 1 & 2 \\ 1 +\epsilon & 2\\ \end{bmatrix}\quad \epsilon \geq \epsilon_{mach}
# $$
#
# then
# $$
# A^{-1} = \frac{-1}{2\epsilon} \begin{bmatrix} 2 & -2 \\ -(1 +\epsilon) & 1\\ \end{bmatrix}
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# So
# $$
# \kappa_1(A) = ||A||_1||A^{-1}||_1 = \frac{4}{2\epsilon}(3 + \epsilon)\sim \frac{6}{\epsilon}
# $$
#
# which is very ill-conditioned.
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ## The condition number and error analysis of $A\mathbf{x}=\mathbf{b}$
#
# The condition number is important in many parts of analysis of numerical linear algebra, but is easily illustrated in understanding the behavior of solutions of linear systems
#
# Assume that $\mathbf{x}$ is a solution to $A\mathbf{x}=\mathbf{b}$ and we want to understand how a small change in the RHS $\mathbf{b}$ propagates to errors in $\mathbf{x}$
# + [markdown] slideshow={"slide_type": "fragment"}
# Consider the perturbed problem
# $$
# A(\mathbf{x} +\Delta\mathbf{x}) = \mathbf{b} + \Delta\mathbf{b}
# $$
# which by linearity of Matrix vector multiplication, and that $A\mathbf{x}=\mathbf{b}$ implies that
#
# $$
# A\Delta\mathbf{x} = \Delta\mathbf{b}
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# or
# $$
# \Delta\mathbf{x} = A^{-1}\Delta\mathbf{b}
# $$
#
# + [markdown] slideshow={"slide_type": "subslide"}
# Given
# $$
# \Delta\mathbf{x} = A^{-1}\Delta\mathbf{b}
# $$
# Taking the norm of both sides implies
#
# $$
# ||\Delta\mathbf{x}|| \leq ||A^{-1}||\,||\Delta\mathbf{b}||
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# since $||A\mathbf{x}|| = ||\mathbf{b}||$, it follows that
#
# $$
# \frac{||\Delta\mathbf{x}||}{||A\mathbf{x}||} \leq ||A^{-1}||\,\frac{||\Delta\mathbf{b}||}{||\mathbf{b}||}
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# or since $||A\mathbf{x}||\leq||A||\,||\mathbf{x}||$, it follows that
#
# $$
# \frac{||\Delta\mathbf{x}||}{||A||\,||\mathbf{x}||} \leq ||A^{-1}||\,\frac{||\Delta\mathbf{b}||}{||\mathbf{b}||}
# $$
#
# or
#
# $$
# \frac{||\Delta\mathbf{x}||}{||\mathbf{x}||} \leq \kappa(A)\,\frac{||\Delta\mathbf{b}||}{||\mathbf{b}||}
# $$
#
# + [markdown] slideshow={"slide_type": "fragment"}
# or the relative error in the solution $\mathbf{x}$ depends on the relative error in the RHS $\mathbf{b}$ times the condition number.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Example
#
# Consider the problem
#
# $$
# \begin{bmatrix} 1 & 2 \\ 1+\alpha & 2\\ \end{bmatrix}\mathbf{x} = \begin{bmatrix} 1 \\ 1 \\ \end{bmatrix}
# $$
# for $\alpha = O(\epsilon_{mach})$
# + slideshow={"slide_type": "-"}
alpha = numpy.finfo(float).eps
A = numpy.array([ [ 1, 2], [1 + alpha, 2] ])
# first version
b = numpy.array( [ 1., 1.])
x = numpy.linalg.solve(A,b)
print('b = {}\nx = {}'.format(b,x))
# + slideshow={"slide_type": "fragment"}
# now perturb b by epsilon
bp = numpy.array( [ 1 , 1 - alpha])
xp = numpy.linalg.solve(A,bp)
print("b'={}\nx' = {}".format(bp,xp))
# + slideshow={"slide_type": "fragment"}
# and calculate relative error and condition number
err_b = numpy.linalg.norm(bp-b)/numpy.linalg.norm(b)
err_x = numpy.linalg.norm(xp-x)/numpy.linalg.norm(x)
condA = numpy.linalg.cond(A)
print('k(A) = {}'.format(condA))
print('err_b = {}, err_x = {}, err_x/err_b ={}'.format(err_b, err_x, err_x/err_b))
| 10_LA_intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.2
# language: julia
# name: julia-1.6
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # 2021-10-18 Iterative methods
#
# ## Last time
#
# * Sparse direct solvers
# * matrix orderings
# * impact on formulation
# * cost scaling
# * Discussion with <NAME>
#
# ## Today
#
# * Why iterative solvers
# * Stationary iterative methods
# * Preconditioning
# + cell_style="center" slideshow={"slide_type": "skip"}
using Plots
using LinearAlgebra
using SparseArrays
default(linewidth=4)
function advdiff_matrix(n; kappa=1, wind=[0, 0])
"Advection-diffusion with Dirichlet boundary conditions eliminated"
h = 2 / (n + 1)
rows = Vector{Int64}()
cols = Vector{Int64}()
vals = Vector{Float64}()
idx((i, j),) = (i-1)*n + j
in_domain((i, j),) = 1 <= i <= n && 1 <= j <= n
stencil_advect = [-wind[1], -wind[2], 0, wind[1], wind[2]] / h
stencil_diffuse = [-1, -1, 4, -1, -1] * kappa / h^2
stencil = stencil_advect + stencil_diffuse
for i in 1:n
for j in 1:n
neighbors = [(i-1, j), (i, j-1), (i, j), (i+1, j), (i, j+1)]
mask = in_domain.(neighbors)
append!(rows, idx.(repeat([(i,j)], 5))[mask])
append!(cols, idx.(neighbors)[mask])
append!(vals, stencil[mask])
end
end
sparse(rows, cols, vals)
end
function my_spy(A)
cmax = norm(vec(A), Inf)
s = max(1, ceil(120 / size(A, 1)))
spy(A, marker=(:square, s), c=:diverging_rainbow_bgymr_45_85_c67_n256, clims=(-cmax, cmax))
end
# + [markdown] slideshow={"slide_type": "slide"}
# # Why iterative solvers over direct solvers?
#
# * Less reliable, more leaky abstraction
# * More sensitive to problem formulation
# * Slower for small problems
# * Several different strategies, each with tuning knobs
# * Accuracy tolerances needed
# + [markdown] slideshow={"slide_type": "fragment"}
# ## $O(N)$ solvers available for many important problems
# * High-order discretization *can* be okay
# + [markdown] slideshow={"slide_type": "slide"}
# # Gradient descent
# + [markdown] cell_style="split" slideshow={"slide_type": "slide"}
# Suppose $A$ is a symmetric positive definite matrix and consider the scalar functional
#
# $$f(u) = \frac 1 2 u^T A u - b^T u . $$
#
# Then the gradient is
#
# $$\nabla_u f = A u - b .$$
# + cell_style="split"
x = LinRange(-4, 4, 40)
A = [1 0; 0 3]
b = [1, 1]
f(u) = .5 * u' * A * u - b' * u
contour(x, x, (u1, u2) -> f([u1, u2]), aspect_ratio=:equal)
# + [markdown] slideshow={"slide_type": "slide"}
# # Aside: Derivative of a dot product
#
# Let $f(\boldsymbol x) = \boldsymbol y^T \boldsymbol x = \sum_i y_i x_i$ and compute the derivative
#
# $$ \frac{\partial f}{\partial \boldsymbol x} = \begin{bmatrix} y_0 & y_1 & \dotsb \end{bmatrix} = \boldsymbol y^T . $$
#
# Note that $\boldsymbol y^T \boldsymbol x = \boldsymbol x^T \boldsymbol y$ and we have the product rule,
#
# $$ \frac{\partial \lVert \boldsymbol x \rVert^2}{\partial \boldsymbol x} = \frac{\partial \boldsymbol x^T \boldsymbol x}{\partial \boldsymbol x} = 2 \boldsymbol x^T . $$
#
# Also,
# $$ \frac{\partial \lVert \boldsymbol x - \boldsymbol y \rVert^2}{\partial \boldsymbol x} = \frac{\partial (\boldsymbol x - \boldsymbol y)^T (\boldsymbol x - \boldsymbol y)}{\partial \boldsymbol x} = 2 (\boldsymbol x - \boldsymbol y)^T .$$
# + [markdown] slideshow={"slide_type": "slide"}
# # Aside: Variational notation
#
# It's convenient to express derivatives in terms of how they act on an infinitessimal perturbation. So we might write
#
# $$ \delta f = \frac{\partial f}{\partial x} \delta x .$$
#
# (It's common to use $\delta x$ or $dx$ for these infinitesimals.) This makes inner products look like a normal product rule
#
# $$ \delta(\mathbf x^T \mathbf y) = (\delta \mathbf x)^T \mathbf y + \mathbf x^T (\delta \mathbf y). $$
#
# A powerful example of variational notation is differentiating a matrix inverse
#
# $$ 0 = \delta I = \delta(A^{-1} A) = (\delta A^{-1}) A + A^{-1} (\delta A) $$
# and thus
# $$ \delta A^{-1} = - A^{-1} (\delta A) A^{-1} $$
# + [markdown] slideshow={"slide_type": "slide"}
# # Try gradient descent
#
# $$ u_{k+1} = u_k - \omega \nabla_u f $$
# + cell_style="center"
function grad_descent(loss, grad, u0; omega=1e-3, tol=1e-5)
"""Minimize loss(c) via gradient descent with initial guess u0
using learning rate gamma. Declares convergence when gradient
is less than tol or after 500 steps.
"""
u = copy(u0)
uhist = [copy(u)]
lhist = [loss(u)]
for it in 1:500
g = grad(u)
u -= omega * g
push!(uhist, copy(u))
push!(lhist, loss(u))
if norm(g) < tol
break
end
end
(u, hcat(uhist...), lhist)
end
# + [markdown] slideshow={"slide_type": "slide"}
# # Visualize gradient descent
# + cell_style="split" slideshow={"slide_type": ""}
A = [1 1; 1 4]
loss(u) = .5 * u' * A * u
grad(u) = A * u
u, uhist, lhist = grad_descent(loss, grad, [.9, .9],
omega=.48)
plot(lhist, yscale=:log10)
# + cell_style="split"
plot(uhist[1, :], uhist[2, :], marker=:circle)
x = LinRange(-1, 1, 30)
contour!(x, x, (x,y) -> loss([x, y]))
# + [markdown] slideshow={"slide_type": "slide"}
# # Richardson iteration
#
# The simplest iterative method is [Richardson's method](https://en.wikipedia.org/wiki/Modified_Richardson_iteration), which solves $A u = b$ by the iteration
# $$ u_{k+1} = u_k + \omega (b - A u_k) $$
# where $\omega > 0$ is a damping parameter and $u_0$ is an initial guess (possibly the zero vector).
#
# * Algebraically equivalent to gradient descent when $A$ is SPD
# * Non-symmetric matrices are harder to visualize
# + [markdown] slideshow={"slide_type": "slide"}
# # Richardson convergence
#
# $$ u_{k+1} = u_k + \omega (b - A u_k) $$
# + [markdown] cell_style="split" slideshow={"slide_type": ""}
# If $b = A u_*$, this iteration is equivalent to
# \begin{align}
# u_{k+1} - u_* &= (u_k - u_*) - \omega A (u_k - u_*) \\
# &= (I - \omega A) (u_k - u_*) .
# \end{align}
# It is convenient for convergence analysis to identify the "error" $e_k = u_k - u_*$, in which this becomes
# $$ e_{k+1} = (I - \omega A) e_k $$
# or
# $$ e_k = (I - \omega A)^k e_0 $$
# in terms of the initial error. Evidently powers of the *iteration matrix* $I - \omega A$ tell the whole story.
# + [markdown] cell_style="split" slideshow={"slide_type": "fragment"}
# Suppose that the eigendecomposition
# $$ X \Lambda X^{-1} = I - \omega A $$
# exists. Then
# $$ (I - \omega A)^k = (X \Lambda X^{-1})^k = X \Lambda^k X^{-1} $$
# and the convergence (or divergence) rate depends only on the largest magnitude eigenvalue.
# This analysis isn't great for two reasons:
#
# 1. Not all matrices are diagonalizable.
# 2. The matrix $X$ may be very ill-conditioned.
# + [markdown] cell_style="center" slideshow={"slide_type": "slide"}
# # Aside: Schur decomposition
#
# We can repair these weaknesses by using the [Schur decomposition](https://en.wikipedia.org/wiki/Schur_decomposition)
# $$ Q R Q^h = I - \omega A $$
# where $R$ is right-triangular and $Q$ is unitary (i.e., orthogonal if real-valued; $Q^h$ is the Hermitian conjugate of $Q$).
# The Schur decomposition always exists and $Q$ has a condition number of 1.
#
# * Where are the eigenvalues in $R$?
#
# Evidently we must find $\omega$ to minimize the maximum eigenvalue of $I - \omega A$. We can do this if $A$ is well conditioned, but not in general.
# + [markdown] slideshow={"slide_type": "slide"}
# # Ill-conditioning
#
# ## Question: What is the condition number of the Laplacian on 100 evenly spaced points?
#
# * How does it scale under grid refinement?
# + cell_style="split"
A = advdiff_matrix(10)
cond(Matrix(A))
# + cell_style="split"
omega = .01
ev = eigvals(Matrix(I - omega * A))
scatter(real.(ev), imag.(ev))
# + [markdown] slideshow={"slide_type": "slide"}
# # Monic polynomials small on the spectrum
# + [markdown] cell_style="split" slideshow={"slide_type": "slide"}
# Equivalently to finding $\omega$ such that $\lVert I - \omega A \rVert$ is minimized, we may seek a monic polynomial $p(z) = 1 - \omega z$ that minimizes
#
# $$ \max_{\lambda \in \sigma(A)} \lvert p(\lambda) \rvert . $$
#
# This concept can be extended to higher degree polynomials, which is essentially what Krylov methods do (discovering the polynomial adaptively, weighted by the right hand side).
# + cell_style="split"
ev = eigvals(Matrix(A))
scatter(real.(ev), zero.(ev))
plot!(x -> 1 - omega * x)
# + [markdown] slideshow={"slide_type": "slide"}
# # Preconditioning
#
# Preconditioning is the act of creating an "affordable" operation "$P^{-1}$" such that $P^{-1} A$ (or $A P^{-1}$) is is well-conditoned or otherwise has a "nice" spectrum. We then solve the system
#
# $$ P^{-1} A x = P^{-1} b \quad \text{or}\quad A P^{-1} \underbrace{(P x)}_y = b $$
#
# in which case the convergence rate depends on the spectrum of the iteration matrix
# $$ I - \omega P^{-1} A . $$
#
# * The preconditioner must be applied on each iteration.
# * It is *not* merely about finding a good initial guess.
#
# There are two complementary techniques necessary for efficient iterative methods:
#
# * "accelerators" or Krylov methods, which use orthogonality to adaptively converge faster than Richardson
# * preconditioners that improve the spectrum of the preconditioned operator
#
# Although there is ongoing research in Krylov methods and they are immensely useful, I would say preconditioning is 90% of the game for practical applications, particularly as a research area.
| slides/2021-10-18-iterative.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **[Machine Learning Micro-Course Home Page](https://www.kaggle.com/learn/intro-to-machine-learning)**
#
# ---
#
# ## Recap
# You've built a model. In this exercise you will test how good your model is.
#
# Run the cell below to set up your coding environment where the previous exercise left off.
# +
# Code you have previously used to load data
import pandas as pd
from sklearn.tree import DecisionTreeRegressor
# Path of the file to read
iowa_file_path = '../input/home-data-for-ml-course/train.csv'
home_data = pd.read_csv(iowa_file_path)
y = home_data.SalePrice
feature_columns = ['LotArea', 'YearBuilt', '1stFlrSF', '2ndFlrSF', 'FullBath', 'BedroomAbvGr', 'TotRmsAbvGrd']
X = home_data[feature_columns]
# Specify Model
iowa_model = DecisionTreeRegressor()
# Fit Model
iowa_model.fit(X, y)
print("First in-sample predictions:", iowa_model.predict(X.head()))
print("Actual target values for those homes:", y.head().tolist())
# Set up code checking
from learntools.core import binder
binder.bind(globals())
from learntools.machine_learning.ex4 import *
print("Setup Complete")
# -
# # Exercises
#
# ## Step 1: Split Your Data
# Use the `train_test_split` function to split up your data.
#
# Give it the argument `random_state=1` so the `check` functions know what to expect when verifying your code.
#
# Recall, your features are loaded in the DataFrame **X** and your target is loaded in **y**.
#
# +
# Import the train_test_split function and uncomment
# from _ import _
# fill in and uncomment
# train_X, val_X, train_y, val_y = ____
step_1.check()
# -
# The lines below will show you a hint or the solution.
# step_1.hint()
# step_1.solution()
# ## Step 2: Specify and Fit the Model
#
# Create a `DecisionTreeRegressor` model and fit it to the relevant data.
# Set `random_state` to 1 again when creating the model.
# +
# You imported DecisionTreeRegressor in your last exercise
# and that code has been copied to the setup code above. So, no need to
# import it again
# Specify the model
iowa_model = ____
# Fit iowa_model with the training data.
____
step_2.check()
# +
# step_2.hint()
# step_2.solution()
# -
# ## Step 3: Make Predictions with Validation data
#
# +
# Predict with all validation observations
val_predictions = ____
step_3.check()
# +
# step_3.hint()
# step_3.solution()
# -
# Inspect your predictions and actual values from validation data.
# print the top few validation predictions
print(____)
# print the top few actual prices from validation data
print(____)
# What do you notice that is different from what you saw with in-sample predictions (which are printed after the top code cell in this page).
#
# Do you remember why validation predictions differ from in-sample (or training) predictions? This is an important idea from the last lesson.
#
# ## Step 4: Calculate the Mean Absolute Error in Validation Data
#
# +
from sklearn.metrics import mean_absolute_error
val_mae = ____
# uncomment following line to see the validation_mae
#print(val_mae)
step_4.check()
# +
# step_4.hint()
# step_4.solution()
# -
# Is that MAE good? There isn't a general rule for what values are good that applies across applications. But you'll see how to use (and improve) this number in the next step.
#
# # Keep Going
#
# You are ready for **[Underfitting and Overfitting](https://www.kaggle.com/dansbecker/underfitting-and-overfitting).**
#
# ---
# **[Machine Learning Micro-Course Home Page](https://www.kaggle.com/learn/intro-to-machine-learning)**
#
#
| notebooks/04_model_validation_excercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import time
#list of input files
dir_in='f:/data/tc_wakes/database/info/'
dir_out='f:/data/tc_wakes/database/sst/'
dir_mur = 'F:/data/sst/jpl_mur/v4.1/'
dir_flux = 'F:/data/model_data/oaflux/data_v3/daily/turbulence/'
dir_cmc = 'F:/data/sst/cmc/CMC0.2deg/v2/'
#################################################################################
import datetime as dt
import xarray as xr
from datetime import datetime
import matplotlib.pyplot as plt
import math
dir_ccmp='F:/data/sat_data/ccmp/v02.0/Y'
date_1858 = dt.datetime(1858,11,17,0,0,0) # start date is 11/17/1958
dx=0.25
dy=0.25
dx_offset = -179.875
dy_offset = -78.3750
storm_date = dt.datetime(2006,10,3)
syr=str(storm_date.year)
smon=str(storm_date.month)
sdym=str(storm_date.day)
sjdy=str(storm_date.timetuple().tm_yday)
print(sjdy)
filename='F:/data/sst/cmc/CMC0.2deg/v2/climatology/clim1993_2016' + sjdy.zfill(3) + '-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0a.nc'
ds_day=xr.open_dataset(filename,drop_variables=['analysis_error','sea_ice_fraction','sq_sst'])
ds_day = ds_day.rename({'analysed_sst':'analysed_sst_clim','mask':'mask_clim'}) #, inplace = True)
ds_day.close()
ds_day
# +
#make lat and lon of storm onto 25 km grid for below
import numpy
minlon, maxlon, minlat, maxlat = -123.,-95., 2., 37.
print(round((maxlat - minlat)/.25))
ydim_storm = int(round((maxlat - minlat)/.25))
new_lat_storm = np.linspace(minlat, maxlat, ydim_storm)
xdim_storm = int(round((maxlon - minlon)/.25))
new_lon_storm = np.linspace(minlon, maxlon, xdim_storm)
storm_date = dt.datetime(2006,10,3)
syr=str(storm_date.year)
smon=str(storm_date.month)
sdym=str(storm_date.day)
sjdy=str(storm_date.timetuple().tm_yday)
fname_tem=syr + smon.zfill(2) + sdym.zfill(2) + '120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
filename = dir_cmc + syr + '/' + sjdy.zfill(3) + '/' + fname_tem
ds_day=xr.open_dataset(filename,drop_variables=['analysis_error','sea_ice_fraction'])
ds_day.close()
ds_masked = ds_day.where(ds_day['mask'] == 1.)
ds_storm = ds_day.interp(lat = new_lat_storm,lon = new_lon_storm)
ds_storm_masked = ds_masked.interp(lat = new_lat_storm,lon = new_lon_storm)
# -
ds_storm
ds_storm_masked.mask.plot()
#dssubset = ds_day.sel(lat=slice(2,37),lon=slice(-123,-95))
#subset = subset.where(subset['mask_clim'] == 1.)
fig, (ax1) = plt.subplots(nrows=1, figsize=(6, 5.4))
im = ax1.imshow(ds_storm.analysed_sst[0,:,:], interpolation='bilinear',aspect='auto',origin='lower',vmin=280,vmax=300)
#extent=[x0, x1, y0, y1])
#ax1.set_title('SST')
#cbar = fig.colorbar(im, extend='both', shrink=0.9, ax=ax1)
#cbar.set_label('SST - SST climatology')
subset = ds_day.sel(lat=slice(2,37),lon=slice(-123,-95))
#subset = subset.where(subset['mask_clim'] == 1.)
fig, (ax1) = plt.subplots(nrows=1, figsize=(6, 5.4))
im = ax1.imshow(subset.analysed_sst_clim, interpolation='bilinear',
aspect='auto',
origin='lower',vmin=0,vmax=300)
#extent=[x0, x1, y0, y1])
ax1.set_title('SST')
cbar = fig.colorbar(im, extend='both', shrink=0.9, ax=ax1)
cbar.set_label('SST - SST climatology')
filename = 'F:/data/tc_wakes/database/sst/2006/055_combined_data.nc'
ds=xr.open_dataset(filename)
ds
ds.analysed_sst_clim[0,:,:].plot()
dir_ccmp='F:/data/sat_data/ccmp/v02.0/Y'
# lyr, idyjl = 2015,1
# storm_date = dt.datetime(2015,1,1)
syr, smon, sdym, sjdy=str(storm_date.year),str(storm_date.month),str(storm_date.day),str(storm_date.timetuple().tm_yday)
fname_tem='/CCMP_Wind_Analysis_' + syr + smon.zfill(2) + sdym.zfill(2) + '_V02.0_L3.0_RSS.nc'
ccmp_filename = dir_ccmp + syr + '/M' + smon.zfill(2) + fname_tem
ds=xr.open_dataset(ccmp_filename,drop_variables=['nobs'])
ds.close()
ds.uwnd[0,:,:].plot()
filename = dir_flux + 'lh_oaflux_' + syr + '.nc';
ds=xr.open_dataset(filename,drop_variables=['err'])
ds_day = ds.sel(time = storm_date.timetuple().tm_yday) #select day of year from annual file
ds_day.close()
print(ds_day)
ds_day.lhtfl.plot()
print(ds_day.lhtfl[0,0])
# +
for root, dirs, files in os.walk(dir_in, topdown=False):
if root[len(dir_in):len(dir_in)+1]=='.':
continue
# if root[len(dir_in):len(dir_in)+4]=='2002':
# continue
# for ii in range(12,13):
for name in files:
# name = files[ii]
# for name in files:
fname_in=os.path.join(root, name)
fname_out=dir_out + fname_in[31:39] + '_all_25km.nc'
inum_storm=int(fname_in[36:39])
iyr_storm=int(fname_in[31:35])
if iyr_storm<=2003: # or iyr_storm<2003:
continue
# if iyr_storm==2011 and inum_storm<15:
# continue
print(name,fname_in)
dsx = xr.open_dataset(fname_in)
lats = dsx.lat[0,:]
lons = dsx.lon[0,:] #lons goes from 0 to 360
lons = (lons + 180) % 360 - 180
dysince = dsx.time
dsx.close()
#make lat and lon of storm onto 25 km grid for below
lons = (((lons - .125)/.25+1).astype(int)-1)*.25+.125
lats = (((lats + 89.875)/.25+1).astype(int)-1)*.25-89.875
iwrap=0
minlon=min(lons.values)-10
maxlon=max(lons.values)+10
minlat=min(lats.values)-10
maxlat=max(lats.values)+10
print('here:',minlon,maxlon)
ydim_storm = round((maxlat - minlat)/.25).astype(int)
new_lat_storm = np.linspace(minlat, maxlat, ydim_storm)
if (minlon<-90 and maxlon>=90) or (minlon<-180 and maxlon<0): #this storm wraps keep everythig 0 to 360 then wrap data at very end
iwrap = 1
lons2 = np.mod(lons, 360)
minlon, maxlon = min(lons2.values)-10, max(lons2.values)+10
xdim_storm = round((maxlon - minlon)/.25).astype(int)
new_lon_storm = np.linspace(minlon, maxlon, xdim_storm)
else:
xdim_storm = round((maxlon - minlon)/.25).astype(int)
new_lon_storm = np.linspace(minlon, maxlon, xdim_storm)
print(iwrap,minlon,maxlon)
print(iwrap,xdim_storm, new_lon_storm[:5],new_lon_storm[-5:])
dims=lats.shape
tdim=dims[0]
tem_date=[0]*tdim #print(dysince.values)
for i in range(0,tdim):
tem_date[i]=date_1858+dt.timedelta(days=float(dysince[0,i].values)) #create new time array that can be queried for year etc
minjdy = min(tem_date).timetuple().tm_yday #create new time array that can be queried for year etc
minyear =min(tem_date).year #create new time array that can be queried for year etc
minmon =min(tem_date).month #create new time array that can be queried for year etc
minday =min(tem_date).day #create new time array that can be queried for year etc
maxjdy = max(tem_date).timetuple().tm_yday #create new time array that can be queried for year etc
maxyear =max(tem_date).year #create new time array that can be queried for year etc
print(minyear,minjdy,maxyear,maxjdy)
dif = max(tem_date)-min(tem_date)
tdim=int(dif.days)+30 #calculate ssts for 30 days after storm
for i in range(0,tdim):
storm_date = dt.datetime(minyear,minmon,minday)+dt.timedelta(days=i)+dt.timedelta(hours=12)
syr=str(storm_date.year)
smon=str(storm_date.month)
sdym=str(storm_date.day)
sjdy=str(storm_date.timetuple().tm_yday)
#sst data
fname_tem=syr + smon.zfill(2) + sdym.zfill(2) + '120000-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
filename = dir_cmc + syr + '/' + sjdy.zfill(3) + '/' + fname_tem
ds_day=xr.open_dataset(filename,drop_variables=['analysis_error','sea_ice_fraction'])
if iwrap==1: #data is -180 to 180 for sst, so need to bring to 0 to 360 when wrapped
ds_day.coords['lon'] = np.mod(ds_day['lon'], 360)
ds_day = ds_day.sortby(ds_day.lon)
ds_day.close()
ds_storm = ds_day.interp(lat = new_lat_storm,lon = new_lon_storm)
#ds_storm['time']=storm_date
if iwrap==1:
ds_storm.coords['lon'] = (ds_storm.coords['lon'] + 180) % 360 - 180
if i==0:
ds_storm_sst = ds_storm
else:
ds_storm_sst = xr.concat([ds_storm_sst,ds_storm],dim='time')
#sst climatology
if storm_date.timetuple().tm_yday==366:
sjdy = '365'
filename='F:/data/sst/cmc/CMC0.2deg/v2/climatology/clim1993_2016' + sjdy.zfill(3) + '-CMC-L4_GHRSST-SSTfnd-CMC0.2deg-GLOB-v02.0-fv02.0.nc'
ds_day=xr.open_dataset(filename,drop_variables=['analysis_error','sea_ice_fraction','sq_sst'])
ds_day = ds_day.rename({'analysed_sst':'analysed_sst_clim','mask':'mask_clim'}) #, inplace = True)
if iwrap==1: #data is -180 to 180 for sst, so need to bring to 0 to 360 when wrapped
ds_day.coords['lon'] = np.mod(ds_day['lon'], 360)
ds_day = ds_day.sortby(ds_day.lon)
ds_day.close()
ds_storm = ds_day.interp(lat = new_lat_storm,lon = new_lon_storm)
ds_storm = ds_storm.assign_coords(time=storm_date)
if iwrap==1:
ds_storm.coords['lon'] = (ds_storm.coords['lon'] + 180) % 360 - 180
if i==0:
ds_storm_sst_clim = ds_storm
else:
ds_storm_sst_clim = xr.concat([ds_storm_sst_clim,ds_storm],dim='time')
# -
# +
#OLD CODE using netcdf and calculating location directly rather than using xarray
###### dir_mur = 'F:/data/sst/jpl_mur/v4.1/'
for root, dirs, files in os.walk(dir_in, topdown=False):
# for ii in range(12,13):
for name in files:
# name = files[ii]
# for name in files:
fname_in=os.path.join(root, name)
fname_out=dir_out + fname_in[31:39] + '_all_25km.nc'
inum_storm=int(fname_in[36:39])
iyr_storm=int(fname_in[31:35])
if iyr_storm>2003 or iyr_storm<2003:
continue
# if iyr_storm==2011 and inum_storm<15:
# continue
print(name,fname_in)
dsx = xr.open_dataset(fname_in)
lats = dsx.lat[0,:]
lons = dsx.lon[0,:] #lons goes from 0 to 360
dysince = dsx.time
#minlon=min(lons[0,:].values)-10
#maxlon=max(lons[0,:].values)+10
#minlat=min(lats[0,:].values)-10
#maxlat=max(lats[0,:].values)+10
iwrap=0
minlon=min(lons.values)-10
maxlon=max(lons.values)+10
minlat=min(lats.values)-10
maxlat=max(lats.values)+10
if minlon<10 and maxlon>350: #wrapping around meridion need to cal new min/max lon
minlon=max(lons[lons<180].values)+10
maxlon=min(lons[lons>180].values)-10
iwrap=1 #set flag for wraparound
#here is a fix for when a storm goes from 350 across 360 to 1 2 longitude
# iwrap=0
# print('first and last!',lons[0,1].values,lons[0,-1].values)
# if abs(min(lons[0,:].values)-max(lons[0,:].values))>180:
# lons1=lons[0,:].values-10>180
# lons2=lons[0,:].values+10<180
# maxlon=min(lons[0,lons1].values-10)
# minlon=max(lons[0,lons2].values+10)
# print('wrapped',minlon,maxlon)
# iwrap=1
#wrap_lons = ((lons+180) % 360) - 180
#maxlon=max(wrap_lons[0,:].values)+10 #this will find the positive maximum
#minlon=min(wrap_lons[0,:].values)-10
#if minlon<0:
# maxlon=min(wrap_lons[0,:].values)-10+360
# minlon=max(wrap_lons[0,:].values)+10
print('min/max lon lat',minlon,maxlon,minlat,maxlat)
ix1=int(round((minlon-dx_offset)/dx))
ix2=int(round((maxlon-dx_offset)/dx))
iy1=int(round((minlat-dy_offset)/dy))
iy2=int(round((maxlat-dy_offset)/dy))
if iy2 > 628:
iy2=628
if iy1 < 1:
iy1=1
if ix1 < 0:
ix1 = ix1 + 1440
if ix2 < 0:
ix2 = ix2 + 1440
print(minlon,maxlon,minlat,maxlat)
xdim=ix2-ix1
if iwrap==1: #wraps around so make sure xdim reflects that
xdim=ix1-ix2+1440
ydim=iy2-iy1
dims=lats.shape
tdim=dims[0]
tem_date=[0]*tdim #print(dysince.values)
for i in range(0,tdim):
tem_date[i]=date_1858+dt.timedelta(days=float(dysince[0,i].values)) #create new time array that can be queried for year etc
minjdy = min(tem_date).timetuple().tm_yday #create new time array that can be queried for year etc
minyear =min(tem_date).year #create new time array that can be queried for year etc
maxjdy = max(tem_date).timetuple().tm_yday #create new time array that can be queried for year etc
maxyear =max(tem_date).year #create new time array that can be queried for year etc
print(minyear,minjdy,maxyear,maxjdy)
dif = max(tem_date)-min(tem_date)
tdim=int(dif.days)+30
print(tdim,ix1,ix2,iy1,iy2,xdim,ydim)
sst_out_sv= np.zeros([tdim,ydim,xdim], dtype="float")
sst_clim_out_sv= np.zeros([tdim,ydim,xdim], dtype="float")
wndu_out_sv= np.zeros([tdim,ydim,xdim], dtype="float")
wndv_out_sv= np.zeros([tdim,ydim,xdim], dtype="float")
wndu_clim_out_sv= np.zeros([tdim,ydim,xdim], dtype="float")
wndv_clim_out_sv= np.zeros([tdim,ydim,xdim], dtype="float")
print('sst_out_sv',sst_out_sv.shape)
for i in range(0,tdim):
storm_date = tem_date[0]+dt.timedelta(days=i)
#print(storm_date)
syr=str(storm_date.year)
smon=str(storm_date.month)
sdym=str(storm_date.day)
sjdy=str(storm_date.timetuple().tm_yday)
fname_tem=syr + smon.zfill(2) + sdym.zfill(2) + '090000-JPL-L4_GHRSST-SSTfnd-MUR-GLOB-v02.0-fv04.1.nc'
mur_filename = dir_mur + syr + '/' + sjdy.zfill(3) + '/' + fname_tem
fname_tem='/CCMP_Wind_Analysis_' + syr + smon.zfill(2) + sdym.zfill(2) + '_V02.0_L3.0_RSS.nc'
ccmp_filename = dir_ccmp + syr + '/M' + smon.zfill(2) + fname_tem
#flux data
lh_flux_filename = dir_flux + 'lh_oaflux_' + syr + '.nc';
sh_flux_filename = dir_flux + 'sh_oaflux_' + syr + '.nc';
ta_flux_filename = dir_flux + 'ta_oaflux_' + syr + '.nc';
qa_flux_filename = dir_flux + 'qa_oaflux_' + syr + '.nc';
# fname='F:\data\model_data\oaflux\data_v3\daily\radiation_1985-2009\sw_isccp_2004.nc';
# [nswrs]=ncread(fname,'nswrs',[1 1 idy],[360 180 1]);
if storm_date.timetuple().tm_yday==366:
sjdy = '365'
clim_filename='F:/data/sst/jpl_mur/v4.1/clim/clim2_' + sjdy.zfill(3) +'_2003_2013_MUR-GLOB-v02.0-fv04.1.nc'
ccmp_clim_filename='F:/data/sat_data/ccmp/v02.0/clim/ccmp_daily_clim_' + sjdy.zfill(3) +'.nc'
# print(ccmp_filename)
# print(ccmp_clim_filename)
#ccmp wind
nc_fid = Dataset(ccmp_filename, 'r')
nc_fid2 = Dataset(ccmp_clim_filename, 'r')
tem = nc_fid.variables['uwnd'][:,iy1:iy2,:] #read in data all longitude, limited latitude
tem = np.mean(tem,axis=0) #take average across all 6 hourly data fields
wndu = np.append(tem[:,ydim:],tem[:,:ydim], axis=1) #switch from 0-360 to -180 to 180 ydim is half of xdim
tem = nc_fid.variables['vwnd'][:,iy1:iy2,:]
tem = np.mean(tem,axis=0)
wndv = np.append(tem[:,ydim:],tem[:,:ydim], axis=1)
mlat_ccmp = nc_fid.variables['latitude'][iy1:iy2]
tem = nc_fid.variables['longitude'][:]
mlon_ccmp = np.append(tem[ydim:],tem[:ydim], axis=0)
mlon_save = mlon_ccmp[:]
mlon_ccmp = ((mlon_ccmp - 180) % 360) - 180 #make -180 to 180 rather than 0 360
tem = nc_fid2.variables['av_u'][iy1:iy2,:]
wndu_clim = np.append(tem[:,ydim:],tem[:,:ydim], axis=1)
tem = nc_fid2.variables['av_v'][iy1:iy2,:]
wndv_clim = np.append(tem[:,ydim:],tem[:,:ydim], axis=1)
nc_fid.close()
nc_fid2.close()
#flux data
ds = xr.open_dataset(lh_flux_filename)
ds_subset = ds.sel(time = idyjl)
ds_res = ds_subset.interp(latitude = new_lat,longitude = mlon_save)
# nc_fid4 = Dataset(sh_flux_filename, 'r')
# nc_fid5 = Dataset(ta_flux_filename, 'r')
# nc_fid6 = Dataset(qa_flux_filename, 'r')
#[lhf1]=ncread(fname,'lhtfl',[1 1 idy],[360 180 1]);
#[Tair1]=ncread(fname,'tmp2m',[1 1 idy],[360 180 1]);
#[Qair1]=ncread(fname,'hum2m',[1 1 idy],[360 180 1]);
#[shf1]=ncread(fname,'shtfl',[1 1 idy],[360 180 1]);
# if i==0:
# print('i=0',iy1,iy2,ix1,ix2,iy2-iy1,ix2-ix1)
if ix1<=1440 and ix2<=1440 and iwrap==0:
# if i==0:
# print('inside1',iy1,iy2,ix1,ix2)
wndu_out = wndu[:,ix1:ix2]
wndu_clim_out = wndu_clim[:,ix1:ix2]
wndv_out = wndv[:,ix1:ix2]
wndv_clim_out = wndv_clim[:,ix1:ix2]
if ix1>1440 and ix2>1440 and iwrap==0:
# if i==0:
# print('inside2',iy1,iy2,ix1,ix2)
wndu_out = wndu[:,ix1-1440:ix2-1440]
wndu_clim_out = wndu_clim[:,ix1-1440:ix2-1440]
wndv_out = wndv[:,ix1-1440:ix2-1440]
wndv_clim_out = wndv_clim[:,ix1-1440:ix2-1440]
if ix1<=1440 and ix2>1440 and iwrap==0:
# if i==0:
# print('inside3',iy1,iy2,ix1,ix2)
tem1 = wndu[:,ix1:]
tem2 = wndu[:,:ix2-1440]
wndu_out = np.append(tem1,tem2, axis=1)
tem1 = wndv[:,ix1:]
tem2 = wndv[:,:ix2-1440]
wndv_out = np.append(tem1,tem2, axis=1)
tem1 = wndu_clim[:,ix1:]
tem2 = wndu_clim[:,:ix2-1440]
wndu_clim_out = np.append(tem1,tem2, axis=1)
tem1 = wndv_clim[:,ix1:]
tem2 = wndv_clim[:,:ix2-1440]
wndv_clim_out = np.append(tem1,tem2, axis=1)
if ix1<=1440 and ix2>1440 and iwrap==1:
# if i==0:
# print('inside1',iy1,iy2,ix1,ix2)
wndu_out = wndu[:,ix2-1440:ix1]
wndu_clim_out = wndu_clim[:,ix2-1440:ix1]
wndv_out = wndv[:,ix2-1440:ix1]
wndv_clim_out = wndv_clim[:,ix2-1440:ix1]
wndu_out_sv[i,:,:]=wndu_out
wndv_out_sv[i,:,:]=wndv_out
wndu_clim_out_sv[i,:,:]=wndu_clim_out
wndv_clim_out_sv[i,:,:]=wndv_clim_out
#sst data
nc_fid = Dataset(mur_filename, 'r')
mlat = nc_fid.variables['lat'][1149:16849]
ilat_mur1 = np.argmin(abs(mlat-mlat_ccmp.min()))-12
#print('mlat first point:', mlat[ilat_mur1_tem])
ilat_mur2 = np.argmin(abs(mlat-mlat_ccmp.max()))+13
mlat = mlat[ilat_mur1:ilat_mur2]
sst = nc_fid.variables['analysed_sst'][0,ilat_mur1:ilat_mur2,:]
mlon = nc_fid.variables['lon'][:]
nc_fid.close()
nc_fid2 = Dataset(clim_filename, 'r')
sst_clim = nc_fid2.variables['sst'][ilat_mur1:ilat_mur2,:]
nc_fid2.close()
coarseness = 25
temp = mlon.reshape((mlon.shape[0] // coarseness, coarseness))
coarse_mlon = np.mean(temp, axis=(1), dtype=np.float64)
temp = mlat.reshape((mlat.shape[0] // coarseness, coarseness))
coarse_mlat = np.mean(temp, axis=(1), dtype=np.float64)
temp = sst.reshape((sst.shape[0] // coarseness, coarseness, sst.shape[1] // coarseness, coarseness))
coarse_sst = np.mean(temp, axis=(1,3), dtype=np.float64)
temp = sst_clim.reshape((sst_clim.shape[0] // coarseness, coarseness, sst_clim.shape[1] // coarseness, coarseness))
coarse_sst_clim = np.mean(temp, axis=(1,3), dtype=np.float64)
#need to recalculate iy1 and iy2 because of offset made earlier to read less of file
# iy1=np.argmin(abs(coarse_mlat-minlat))
# iy2=np.argmin(abs(coarse_mlat-maxlat))
# ydim=iy2-iy1
# print(coarse_mlat[0],coarse_mlat[-1])
# print(iy1,iy2,ydim,minlat,maxlat)
# if i==0:
# print('i=0',ix1,ix2,iy2-iy1,ix2-ix1)
if ix1<=1440 and ix2<=1440 and iwrap==0:
# if i==0:
# print('inside1',ix1,ix2)
sst_out = coarse_sst[:,ix1:ix2]
mlat_out = coarse_mlat[:]
mlon_out = coarse_mlon[ix1:ix2]
sst_clim_out = coarse_sst_clim[:,ix1:ix2]
if ix1>1440 and ix2>1440 and iwrap==0:
# if i==0:
# print('inside2',ix1,ix2)
sst_out = coarse_sst[:,ix1-1440:ix2-1440]
mlat_out = coarse_mlat[:]
mlon_out = coarse_mlon[ix1-1440:ix2-1440]
sst_clim_out = coarse_sst_clim[:,ix1-1440:ix2-1440]
if ix1<=1440 and ix2>1440 and iwrap==0:
# if i==0:
# print('inside3',ix1,ix2)
tem1 = coarse_sst[:,ix1:]
tem2 = coarse_sst[:,:ix2-1440]
sst_out = np.append(tem1,tem2, axis=1)
mlat_out = coarse_mlat[:]
mlon1 = coarse_mlon[ix1:]
mlon2 = coarse_mlon[:ix2-1440]
print(mlon1.shape,mlon2.shape)
mlon_out = np.append(mlon1,mlon2, axis=0)
tem1 = coarse_sst_clim[:,ix1:]
tem2 = coarse_sst_clim[:,:ix2-1440]
sst_clim_out = np.append(tem1,tem2, axis=1)
if ix1<=1440 and ix2>1440 and iwrap==1:
# if i==0:
# print('inside1',ix1,ix2)
sst_out = coarse_sst[:,ix2-1440:ix1]
mlat_out = coarse_mlat[:]
mlon_out = coarse_mlon[ix2-1440:ix1]
sst_clim_out = coarse_sst_clim[:,ix2-1440:ix1]
if i==0:
print('sst',sst_out.shape,'coarse',coarse_sst.shape,wndu_out.shape)
print('sst',sst_out.shape,'sst_sv',sst_out_sv.shape)
print('mlon',mlon_out.shape,'mlat',mlat_out.shape)
#sst_sv[i,:,:]=sst-sst_clim
sst_out_sv[i,:,:]=sst_out
sst_clim_out_sv[i,:,:]=sst_clim_out
ilen=len(fname_in)
dif_dys=[0]*tdim
for i in range(0,tdim):
dif_dys[i] = i
print('file out:',fname_out)
#f.close()
f = Dataset(fname_out,'w', format='NETCDF4')
tempgrp = f.createGroup('data')
tempgrp.setncattr_string('start time',str(tem_date[0]))
tempgrp.createDimension('t', tdim)
tempgrp.createDimension('y', ydim)
tempgrp.createDimension('x', xdim)
#tem_date[i]
sst_netcdf = tempgrp.createVariable('sst', 'f4', ('t', 'y', 'x'))
sst_clim_netcdf = tempgrp.createVariable('sst_clim', 'f4', ('t', 'y', 'x'))
wndu_netcdf = tempgrp.createVariable('wndu', 'f4', ('t', 'y', 'x'))
wndv_netcdf = tempgrp.createVariable('wndv', 'f4', ('t', 'y', 'x'))
wndu_clim_netcdf = tempgrp.createVariable('wndu_clim', 'f4', ('t', 'y', 'x'))
wndv_clim_netcdf = tempgrp.createVariable('wndv_clim', 'f4', ('t', 'y', 'x'))
longitude = tempgrp.createVariable('lon', 'f4', 'x')
latitude = tempgrp.createVariable('lat', 'f4', 'y')
time = tempgrp.createVariable('time', 'i4', 't')
sst_netcdf[:] = sst_out_sv
sst_clim_netcdf[:] = sst_clim_out_sv
wndu_netcdf[:] = wndu_out_sv
wndv_netcdf[:] = wndv_out_sv
wndu_clim_netcdf[:] = wndu_clim_out_sv
wndv_clim_netcdf[:] = wndv_clim_out_sv
latitude[:] = mlat_out
longitude[:] = mlon_out
time[:]=dif_dys
f.close()
# -
| .ipynb_checkpoints/testing collocated SST for low values-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# from https://github.com/socrata/dev.socrata.com/blob/39c6581986466edb5e7f72f5beea5ce69238f8de/snippets/pandas.py
import pandas as pd
from sodapy import Socrata
# Unauthenticated client only works with public data sets. Note 'None'
# in place of application token, and no username or password:
client = Socrata("data.cityofchicago.org", None)
# First 50000 results, returned as JSON from API
# Connverted to Python list of dictionaries by sodapy.
# Column names converted to snake case, special chars removed
# Dates and location formatted
results = client.get("me59-5fac", limit=50000)
# Convert to pandas DataFrame
complaints = pd.DataFrame.from_records(results)
# -
# download remaining (limit 50000 / call)
start = 50000
while results:
print(start)
results = client.get("me59-5fac", limit=50000, offset=start)
complaints = complaints.append(pd.DataFrame.from_records(results))
start += 50000
# Drop rows with missing data
complaints.dropna(subset=["latitude", "longitude", "creation_date"], inplace=True)
# Filter by status
complaints = complaints[complaints.status.isin(["Completed", "Open"])]
# Convert latitude & longitude to floats
complaints.latitude = complaints.latitude.astype(float)
complaints.longitude = complaints.longitude.astype(float)
# +
import os.path
root_path = os.path.dirname(os.getcwd())
# Save result
complaints.to_csv(os.path.join(root_path, "DATA/sanitation_complaints.csv"), index=False)
# -
| CODE/15_sanitation_download.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Fuzzy Logic for Python 3
#
# The doctests in the modules should give a good idea how to use things by themselves, while here are some examples how to use everything together.
# # Installation
# First things first: To install fuzzylogic, just enter `python -m pip install fuzzylogic` and you should be good to go!
# ### Functions and Sets
# Defining a domain with its range and resolution should be trivial since most real world instruments come with those specifications. However, defining the fuzzy sets within those domains is where the fun begins as only a human can tell whether something is "hot" or "not", right?
#
# Why the distinction? Functions only map values, nothing special there at all - which is good for testing and performance. Sets on the other hand implement logical operations that have special python syntax, which makes it easy to work with but a little more difficult to test and adds some performance overhead. So, sets are for abstraction and easy handling, functions for performance.
#
# ### Domains
# You can use (I do so regularly) fuzzy functions outside any specific fuzzy context. However, if you want to take advantage of the logic of fuzzy sets, plot stuff or defuzzyfy values, you need to use Domains. Domains and Sets are special in a way that they intrically rely on each other. This is enforced by how assignments work. Regular Domain attributes are the sets that were assigned to the domain. Also, if just a function is assigned it is automatically wrapped in a Set.
from matplotlib import pyplot
pyplot.rc("figure", figsize=(10, 10))
# +
from fuzzylogic.classes import Domain
from fuzzylogic.functions import R, S, alpha
T = Domain("test", 0, 30, res=0.1)
# -
T.up = R(1,10)
T.up.plot()
T.down = S(20, 29)
T.down.plot()
T.polygon = T.up & T.down
T.polygon.plot()
T.inv_polygon = ~T.polygon
T.inv_polygon.plot()
# let's show off a few interesting functions ;)
# +
from fuzzylogic.classes import Domain, Set
from fuzzylogic.functions import (sigmoid, gauss, trapezoid,
triangular_sigmoid, rectangular)
T = Domain("test", 0, 70, res=0.1)
T.sigmoid = sigmoid(1,1,20)
T.sigmoid.plot()
T.gauss = gauss(10, 0.01, c_m=0.9)
T.gauss.plot()
T.trapezoid = trapezoid(25, 30, 35, 40, c_m=0.9)
T.trapezoid.plot()
T.triangular_sigmoid = triangular_sigmoid(40, 70, c=55)
T.triangular_sigmoid.plot()
# -
# ### Domains
#
# After specifying the domain and assigning sets, calling a domain with a value returns a dict of memberships of the sets in that domain.
# +
from fuzzylogic.classes import Domain
from fuzzylogic.functions import alpha, triangular
from fuzzylogic.hedges import plus, minus, very
numbers = Domain("numbers", 0, 20, res=0.1)
close_to_10 = alpha(floor=0.2, ceiling=0.8, func=triangular(0, 20))
close_to_5 = triangular(1, 10)
numbers.foo = minus(close_to_5)
numbers.bar = very(close_to_10)
numbers.bar.plot()
numbers.foo.plot()
numbers.baz = numbers.foo + numbers.bar
numbers.baz.plot()
numbers(8)
# +
from fuzzylogic.classes import Domain
from fuzzylogic.functions import bounded_sigmoid
T = Domain("temperature", 0, 100, res=0.1)
T.cold = bounded_sigmoid(5,15, inverse=True)
T.cold.plot()
T.hot = bounded_sigmoid(20, 40)
T.hot.plot()
T.warm = ~T.hot & ~T.cold
T.warm.plot()
T(10)
# -
# Many times you end up with sets that never hit 1 like with sigmoids, triangular funcs that hit the border of the domain or after operations with other sets. Then it is often needed to normalize (define max(set) == 1). Note that Set.normalized() returns a set that (unlike other set ops) is already bound to the domain and given the name "normalized\_{set.name}". This can't be circumvented because normalizing is only defined on a given domain.
# +
from fuzzylogic.classes import Domain
from fuzzylogic.functions import alpha, trapezoid
N = Domain("numbers", 0, 6, res=0.01)
N.two_or_so = alpha(floor=0, ceiling=0.7, func=trapezoid(0, 1.9, 2.1, 4))
N.two_or_so.plot()
N.x = N.two_or_so.normalized()
N.x.plot()
# -
# ### Inference
#
# After measuring a RL value and mapping it to sets within a domain, it is normally needed to translate the result to another domain that corresponds to some sort of control mechanism. This translation or mapping is called inference and is rooted in the logical conclusion operation A => B, for example: If it rains then the street is wet.
# The street may be wet for a number of reasons, but if it rains it will be wet for sure. This **IF A THEN B** can also be written as
# ***(A AND B) OR NOT(A AND TRUE)***. This may look straight forward for boolean logic, but since we are not just dealing with True and False, there are a number of ways in fuzzy logic to actually implement this.
# Here is a simple but fully working example with all moving parts, demonstrating the use in the context of an HVAC system.
#
# It also demonstrates the three different ways to set up complex combinations of rules: you can either define each rule one by one and then combine them via the | operator, or you can put the rules into a list and use sum(..) to combine them into one in a single step, or you can define one big and complex rule right from the start. Which way best suits your needs depends on how complex each rule is and how/where you define them in your code and whether you need to use them in different places in different combinations.
# +
from fuzzylogic.classes import Domain, Set, Rule
from fuzzylogic.hedges import very
from fuzzylogic.functions import R, S
temp = Domain("Temperature", -80, 80)
hum = Domain("Humidity", 0, 100)
motor = Domain("Speed", 0, 2000)
temp.cold = S(0,20)
temp.hot = R(15,30)
hum.dry = S(20,50)
hum.wet = R(40,70)
motor.fast = R(1000,1500)
motor.slow = ~motor.fast
R1 = Rule({(temp.hot, hum.dry): motor.fast})
R2 = Rule({(temp.cold, hum.dry): very(motor.slow)})
R3 = Rule({(temp.hot, hum.wet): very(motor.fast)})
R4 = Rule({(temp.cold, hum.wet): motor.slow})
rules = Rule({(temp.hot, hum.dry): motor.fast,
(temp.cold, hum.dry): very(motor.slow),
(temp.hot, hum.wet): very(motor.fast),
(temp.cold, hum.wet): motor.slow,
})
rules == R1 | R2 | R3 | R4 == sum([R1, R2, R3, R4])
values = {hum: 45, temp: 22}
print(R1(values), R2(values), R3(values), R4(values), "=>", rules(values))
# -
# There are a few things to note in this example. Firstly, make sure to pass in the values as a single dictionary at the end, not as parameters.
# If a rule has zero weight - in this example a temp of 22 results in cold weighted with S(0,20) as 0 - the Rule returns None, which makes sure this condition is ignored in subsequent calculations. Also you might notice that the result is a value of 1633, which is way more than motor.fast with R(1000,1500) would suggest. However, since the domain motor is defined between 0 and 2000, the center of gravity method used for evaluation takes many of the values between 1500 and 2000 weighted with 1 into account, giving this slightly unexpected result.
# Oftentimes "real" logical conclusion isn't actually what you need, instead you might want to build something like a weighted decision tree, which is very easy to do.
# Let's see how to rate a meal as an example.
# Just keep in mind that rule functions take a dict of membership values as single argument once parametrized. So, in this case, the ratings dict either needs to be sanitized before passed into the rule or constructed from the beginning (which I've done here) in the appropriate way.
# +
from fuzzylogic.classes import Domain
from fuzzylogic.functions import bounded_linear
from fuzzylogic.rules import weighted_sum
rating = Domain("rating", 1, 10, res=0.1)
rating.norm = bounded_linear(1, 10)
weights = {"beverage": 0.3,
"atmosphere": 0.2,
"looks":0.2,
"taste": 0.3}
w_func = weighted_sum(weights=weights, target_d=rating)
# we've rated beverage etc. on a scale from 1 to 10 separately
ratings = {"beverage": rating.min(9),
"atmosphere": rating.min(5),
"looks": rating.min(4),
"taste": rating.min(8)}
w_func(ratings)
# -
# ## Sources
# * Fuzzy Logic and Control: Software and Hardware Applications, Volume 2
#
# By: <NAME>; <NAME>; <NAME> - University of New Mexico
# Publisher: Prentice Hall
# Pub. Date: June 07, 1993
#
# * Computational Intelligence - Fuzzy-Tutorial
#
# Prof. Dr. <NAME>
#
# * http://petro.tanrei.ca/fuzzylogic/fuzzy_negnevistky.html
# * http://kik.informatik.fh-dortmund.de/abschlussarbeiten/fuzzyControl/operatoren.html
# * Fundamentals of Fuzzy Logic Control – Fuzzy Sets, Fuzzy Rules and Defuzzifications
#
# <NAME> and <NAME>
#
# https://www.researchgate.net/profile/Ying_Bai/publication/225872318_Fundamentals_of_Fuzzy_Logic_Control_-_Fuzzy_Sets_Fuzzy_Rules_and_Defuzzifications/links/0fcfd5057a874858b1000000/Fundamentals-of-Fuzzy-Logic-Control-Fuzzy-Sets-Fuzzy-Rules-and-Defuzzifications.pdf
#
| docs/Showcase.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] lang="en"
# # Dog Autoencoder
# In this kernel, we learn about autoencoders. By understanding autoencoders, we will better understand GANs (Generative Adversarial Networks) and VAEs (Variational Autoencoders). We will also learn how to use an autoencoder to generate images of dogs.
#
# 
#
# Kaggle's "Generative Dog Images" competition asks us to generate dog images using **generative methods**. It is unclear whether we must use GANs. If we must use GANs, then this kernel's output is **not** a valid competition submission.
# # Load Data and Augment
# We will randomly crop the original 20,000 images and make 500,000 new training images.
# + [markdown] lang="ja"
# #Dog Autoencoder
# このカーネルでは、オートエンコーダについて学びます。オートエンコーダを理解することで、GAN(Generative Adversarial Networks)とVAE(変分オートエンコーダ)をよりよく理解することができます。また、オートエンコーダを使って犬の画像を生成する方法も学びます。
#
# 
#
# Kaggleの "Generative Dog Images"コンペティションでは、**生成方法を使って犬の画像を生成するよう求められています**。 GANを使用する必要があるかどうかは不明です。 GANを使わなければならないのなら、このカーネルの出力は**ではありません**有効な競合の投稿です。
# #データの読み込みと強化
# 元の20,000枚の画像をランダムに切り取り、50万枚の新しいトレーニング画像を作成します。
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _kg_hide-input=true _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
ComputeLB = False
import os, gc, zipfile
import numpy as np, pandas as pd
from PIL import Image
import matplotlib.pyplot as plt
if ComputeLB: PATH = '../input/generative-dog-images/all-dogs/all-dogs/'
else: PATH = '../input/all-dogs/all-dogs/'
IMAGES = os.listdir(PATH)
print('There are',len(IMAGES),'images. Here are 5 example filesnames:')
print(IMAGES[:5])
# + _kg_hide-input=true
os.mkdir('../tmp')
os.mkdir('../tmp/images')
# CREATE RANDOMLY CROPPED IMAGES
for i in range(500000):
img = Image.open(PATH + IMAGES[i%len(IMAGES)])
img = img.resize(( 100,int(img.size[1]/(img.size[0]/100) )), Image.ANTIALIAS)
w = img.size[0]; h = img.size[1]; a=0; b=0
if w>64: a = np.random.randint(0,w-64)
if h>64: b = np.random.randint(0,h-64)
img = img.crop((a, b, 64+a, 64+b))
img.save('../tmp/images/'+str(i)+'.png','PNG')
if i%100000==0: print('created',i,'cropped images')
print('created 500000 cropped images')
# + [markdown] lang="en"
# # Build Autoencoder
# This code is inspired by Keras' tutorial [here][1]
#
# [1]: https://blog.keras.io/building-autoencoders-in-keras.html
# + [markdown] lang="ja"
# #オートエンコーダをビルドする
# このコードはKerasのチュートリアルに触発されました[ここ] [1]
#
# [1]:https://blog.keras.io/building-autoencoders-in-keras.html
# +
from keras.models import Model
from keras.layers import Input, Dense, Conv2D, MaxPooling2D, UpSampling2D
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import LearningRateScheduler
BATCH_SIZE = 256; EPOCHS = 10
train_datagen = ImageDataGenerator(rescale=1./255)
train_batches = train_datagen.flow_from_directory('../tmp/',
target_size=(64,64), shuffle=True, class_mode='input', batch_size=BATCH_SIZE)
# +
# ENCODER
input_img = Input(shape=(64, 64, 3))
x = Conv2D(48, (3, 3), activation='relu', padding='same')(input_img)
x = MaxPooling2D((2, 2), padding='same')(x)
x = Conv2D(96, (3, 3), activation='relu', padding='same')(x)
x = MaxPooling2D((2, 2), padding='same')(x)
x = Conv2D(192, (3, 3), activation='relu', padding='same')(x)
x = MaxPooling2D((2, 2), padding='same')(x)
encoded = Conv2D(32, (1, 1), activation='relu', padding='same')(x)
# LATENT SPACE
latentSize = (8,8,32)
# DECODER
direct_input = Input(shape=latentSize)
x = Conv2D(192, (1, 1), activation='relu', padding='same')(direct_input)
x = UpSampling2D((2, 2))(x)
x = Conv2D(192, (3, 3), activation='relu', padding='same')(x)
x = UpSampling2D((2, 2))(x)
x = Conv2D(96, (3, 3), activation='relu', padding='same')(x)
x = UpSampling2D((2, 2))(x)
x = Conv2D(48, (3, 3), activation='relu', padding='same')(x)
decoded = Conv2D(3, (3, 3), activation='sigmoid', padding='same')(x)
# COMPILE
encoder = Model(input_img, encoded)
decoder = Model(direct_input, decoded)
autoencoder = Model(input_img, decoder(encoded))
autoencoder.compile(optimizer='Adam', loss='binary_crossentropy')
# -
# # Train Autoencoder
history = autoencoder.fit_generator(train_batches,
steps_per_epoch = train_batches.samples // BATCH_SIZE,
epochs = EPOCHS, verbose=2)
# + [markdown] lang="en"
# # View Reconstruction
# Our encoder works by mapping images from 12288 dimensional space (64 x 64 x 3) into 2048 dimensional space (latent image). This is a 6x compression rate. Our decoder works by mapping our latent image back into 12288 dimensional space. Below are examples. (Note that a decoder is like a GAN generator).
# + [markdown] lang="ja"
# #再構成を見る
# 私たちのエンコーダは、12288次元の空間(64 x 64 x 3)から2048次元の空間(潜像)に画像をマッピングすることによって機能します。これは6倍の圧縮率です。私たちのデコーダは、私たちの潜像を12288次元の空間にマッピングし直すことによって機能します。以下は例です。 (デコーダはGANジェネレータのようなものです)。
# + _kg_hide-input=true
images = next(iter(train_batches))[0]
for i in range(5):
plt.figure(figsize=(15,5))
plt.subplot(1,3,1)
# ORIGINAL IMAGE
orig = images[i,:,:,:].reshape((-1,64,64,3))
img = Image.fromarray( (255*orig).astype('uint8').reshape((64,64,3)))
plt.title('Original')
plt.imshow(img)
# LATENT IMAGE
latent_img = encoder.predict(orig)
mx = np.max( latent_img[0] )
mn = np.min( latent_img[0] )
latent_flat = ((latent_img[0] - mn) * 255/(mx - mn)).flatten(order='F')
img = Image.fromarray( latent_flat[:2025].astype('uint8').reshape((45,45)), mode='L')
plt.subplot(1,3,2)
plt.title('Latent')
plt.xlim((-10,55))
plt.ylim((-10,55))
plt.axis('off')
plt.imshow(img)
# RECONSTRUCTED IMAGE
decoded_imgs = decoder.predict(latent_img[0].reshape((-1,latentSize[0],latentSize[1],latentSize[2])))
img = Image.fromarray( (255*decoded_imgs[0]).astype('uint8').reshape((64,64,3)))
plt.subplot(1,3,3)
plt.title('Reconstructed')
plt.imshow(img)
plt.show()
# + [markdown] lang="en"
# # Latent Space and Dog Generation
# We choose the size of latent space. In our autoencoder, we chose for latent space to be 2048 dimensional (6x compression). If we map all 20,000 images into latent space, they would cluster inside a 2048 dimensional hyperellipsoid. That ellipsoid would represent dog images. (Cat images would form a different ellipsoid). Below I have plotted 256 of our dog images in latent space as blue dots and drew their ellipsoid. (Note that latent space is being projected onto 2D for display in this kernel). (Note because of ReLU, you may see clipping).
#
# If we would like to generate a new dog image, we can chose a new random point (different from existing training image dots) inside this ellipsoid and then decode it. For example, we could choose the 9 red points below and then convert them into dog images.
# + [markdown] lang="ja"
# #潜在空間と犬の世代
# 潜在空間のサイズを選択します。私たちのオートエンコーダでは、潜在空間が2048次元(6倍圧縮)になるように選択しました。 2万枚すべての画像を潜在空間にマッピングすると、それらは2048次元の超楕円体の中に集まります。その楕円は犬のイメージを表します。 (猫の画像は別の楕円体を形成します)。下の図では、256個の犬のイメージを青い点として潜在空間にプロットし、その楕円体を描きました。 (このカーネルでは、潜在空間が表示用に2Dに投影されていることに注意してください)。 (ReLUのため、クリッピングが発生する可能性があります)。
#
# 新しい犬の画像を生成したい場合は、この楕円の中に新しいランダムな点(既存のトレーニング画像のドットとは異なる)を選択してからデコードすることができます。たとえば、以下の9つの赤い点を選択してから犬の画像に変換できます。
# + _kg_hide-input=true
from matplotlib.patches import Ellipse
# PROJECT LATENT INTO 2D, AVOID DEAD RELU
latent_img = encoder.predict(images)
latent_img2 = latent_img.reshape((-1,latentSize[0]*latentSize[1]*latentSize[2]))
d = 0; s = 0
while s<0.1:
x = latent_img2[:,d]
s = np.std(x); d += 1
s = 0
while s<0.1:
y = latent_img2[:,d]
s = np.std(y); d += 1
# CALCULATE ELLIPSOID FROM 256 IMAGES
cov = np.cov(x, y)
lambda_, v = np.linalg.eig(cov)
lambda_ = np.sqrt(lambda_)
for j in [1,2,3]:
ell = Ellipse(xy=(np.mean(x), np.mean(y)), width=lambda_[0]*j*2,
height=lambda_[1]*j*2, angle=np.rad2deg(np.arccos(v[0, 0])))
ell.set_facecolor('None')
ell.set_edgecolor('black')
plt.gca().add_artist(ell)
# PLOT 256 IMAGES AS DOTS IN LATENT SPACE
plt.scatter(x,y)
d = np.random.multivariate_normal([np.mean(x),np.mean(y)],cov,9)
plt.scatter(d[:,0],d[:,1],color='red',s=100)
plt.title('Dog Images form an Ellipsoid in Latent Space')
plt.show()
# +
# CREATE 10000 CROPPED IMAGES
x = np.random.choice(np.arange(20000),10000)
images = np.zeros((10000,64,64,3))
for i in range(len(x)):
img = Image.open(PATH + IMAGES[x[i]])
img = img.resize((100,int(img.size[1]/(img.size[0]/100))), Image.ANTIALIAS)
img = img.crop((18, 0, 82, 64))
images[i,:,:,:] = np.asarray(img).astype('float32') / 255.
#if i%1000==0: print(i)
# CALCULATE ELLIPSOID FROM 10000 IMAGES
encoded_imgs = encoder.predict(images)
sz = latentSize[0] * latentSize[1] * latentSize[2]
encoded_imgs = encoded_imgs.reshape((-1,sz))
mm = np.mean(encoded_imgs,axis=0)
ss = np.cov(encoded_imgs,rowvar=False)
# GENERATE 9 RANDOM DOG IMAGES
generated = np.random.multivariate_normal(mm,ss,9)
generated = generated.reshape((-1,latentSize[0],latentSize[1],latentSize[2]))
# + _kg_hide-input=true
# PLOT 9 RANDOM DOG IMAGES
for k in range(3):
plt.figure(figsize=(15,5))
plt.subplot(1,3,1)
decoded_imgs = decoder.predict(generated[k*3].reshape((-1,latentSize[0],latentSize[1],latentSize[2])))
img = Image.fromarray( (255*decoded_imgs[0]).astype('uint8').reshape((64,64,3)))
plt.imshow(img)
plt.subplot(1,3,2)
decoded_imgs = decoder.predict(generated[k*3+1].reshape((-1,latentSize[0],latentSize[1],latentSize[2])))
img = Image.fromarray( (255*decoded_imgs[0]).astype('uint8').reshape((64,64,3)))
plt.imshow(img)
plt.subplot(1,3,3)
decoded_imgs = decoder.predict(generated[k*3+2].reshape((-1,latentSize[0],latentSize[1],latentSize[2])))
img = Image.fromarray( (255*decoded_imgs[0]).astype('uint8').reshape((64,64,3)))
plt.imshow(img)
plt.show()
# + [markdown] lang="en"
# # How is LB scored?
# What is the FID (Fréchet Inception Distance) metric? Now that we understand latent space, the FID metric is easy to understand. After we train our autoencoder, we can compute the latent hyper-ellipsoid of our dog training data. Then if someone generates dog images, we could feed them into our autoencoder and calculate the latent hyper-ellipsoid of their generated dog images. Then FID is basically the distance between the two latent hyper-ellipsoids squared.
#
# Let `e1` be the training dog ellipsoid and `e2` be the generated dog ellipsoid. Let `x0, x1, ..., x2047` be the 2048 features of latent space. Each vector `x_k` from training dataset `e1` has approx 20000 values, and `e2` has 10000. Then the distance between the ellipsoids is approx the sum of the difference of means (ellipse centers) and sum of the difference of standard deviations (ellipse widths) of each 2048 `x_k`, i.e. `(mean(x_k)_e1 - mean(x_k)_e2)` plus `(std(x_k)_e1 - std(x_k)_e2)` for each of the 2048 `k` values. Finally `FID = this sum^2`.
#
# In the image below, `FID approx = ||c2-c1||^2 + (h2-h1)^2 + (w2-w1)^2`. (Note this is exact when features are uncorrelated). MiFID adds a multiplier penalty if your dog images are too similar to the training data. `MiFID = FID * penalty`.
#
# 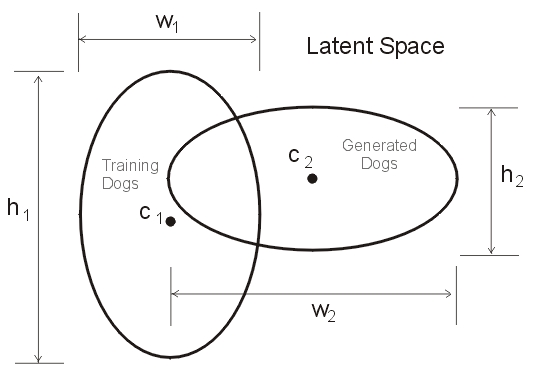
#
# # Generating Better Dog Images
# The generated dog images above are not very recognizable. Instead of choosing completely random points in latent space, we can choose points near existing training images. Let's try that.
# + [markdown] lang="ja"
# #LBの得点は?
# FID(フレシェインセプションディスタンス)メトリックとは何ですか?潜在空間を理解したので、FIDメトリックは理解しやすくなりました。オートエンコーダをトレーニングした後、犬の訓練データの潜在的な超楕円体を計算することができます。次に、誰かが犬の画像を生成した場合、それらをオートエンコーダに入力して、生成された犬の画像の潜在的な超楕円体を計算することができます。その場合、FIDは基本的に2つの潜在的な超楕円体間の距離の2乗です。
#
# 訓練犬の楕円体を 'e1'とし、生成された犬の楕円体を 'e2'とする。 `x0、x1、...、x2047`を潜在空間の2048個の特徴とする。訓練データセット「e1」からの各ベクトル「x <k>」は約20000個の値を有し、「e2」は10000個の値を有する。次いで、楕円間の距離は、平均の差の合計(楕円中心)との和の合計である。各2048 x x kの標準偏差(楕円幅)の差、すなわち(mean(x k)e1 - mean(x k)e2)プラス2048個の「k」値のそれぞれについて、「(std(x <k>)e1 − std(x <k)<2> e2)」となる。最後に `FID = this sum ^ 2`です。
#
# 下の図では、「FID約= undefinedc2-c1未定義^ 2 +(h2-h1)^ 2 +(w2-w1)^ 2」です。 (特徴が相関していない場合、これは正確です)。犬の画像がトレーニングデータと非常に類似している場合、MiFIDは乗数のペナルティを追加します。 `MiFID = FID *ペナルティ`。
#
# 
#
# #よりよい犬の画像を生成する
# 上記の生成された犬の画像はあまり認識できません。潜在空間内で完全にランダムな点を選択する代わりに、既存のトレーニング画像の近くの点を選択できます。やってみましょう。
# -
# DISTANCE TO MOVE AWAY FROM EXISTING TRAIN IMAGES
beta = 0.35
# GENERATE 9 RANDOM DOG IMAGES
generated = np.random.multivariate_normal(mm,ss,9)
generated = beta*generated + (1-beta)*encoded_imgs[:9]
# + _kg_hide-input=true
for k in range(3):
plt.figure(figsize=(15,5))
plt.subplot(1,3,1)
decoded_imgs = decoder.predict(generated[k*3].reshape((-1,latentSize[0],latentSize[1],latentSize[2])))
img = Image.fromarray( (255*decoded_imgs[0]).astype('uint8').reshape((64,64,3)))
plt.imshow(img)
plt.subplot(1,3,2)
decoded_imgs = decoder.predict(generated[k*3+1].reshape((-1,latentSize[0],latentSize[1],latentSize[2])))
img = Image.fromarray( (255*decoded_imgs[0]).astype('uint8').reshape((64,64,3)))
plt.imshow(img)
plt.subplot(1,3,3)
decoded_imgs = decoder.predict(generated[k*3+2].reshape((-1,latentSize[0],latentSize[1],latentSize[2])))
img = Image.fromarray( (255*decoded_imgs[0]).astype('uint8').reshape((64,64,3)))
plt.imshow(img)
plt.show()
# -
beta = 0.2
# GENERATE 10000 RANDOM DOG IMAGES FOR KAGGLE
generated = np.random.multivariate_normal(mm,ss,10000)
encoded_imgs = beta*generated + (1-beta)*encoded_imgs
decoded_imgs = decoder.predict(encoded_imgs.reshape((-1,latentSize[0],latentSize[1],latentSize[2])))
decoded_imgs.shape
# # Submit to Kaggle
# SAVE TO ZIP FILE NAMED IMAGES.ZIP
z = zipfile.PyZipFile('images.zip', mode='w')
for k in range(10000):
img = Image.fromarray( (255*decoded_imgs[k]).astype('uint8').reshape((64,64,3)))
f = str(k)+'.png'
img.save(f,'PNG'); z.write(f); os.remove(f)
#if k % 1000==0: print(k)
z.close()
# # Calculate LB Score
# If you wish to compute LB, you must add the LB metric dataset [here][1] to this kernel and change the boolean variable in the first cell block.
#
# [1]: https://www.kaggle.com/wendykan/dog-face-generation-competition-kid-metric-input
# + _kg_hide-input=true
from __future__ import absolute_import, division, print_function
import numpy as np
import os
import gzip, pickle
import tensorflow as tf
from scipy import linalg
import pathlib
import urllib
import warnings
from tqdm import tqdm
from PIL import Image
class KernelEvalException(Exception):
pass
model_params = {
'Inception': {
'name': 'Inception',
'imsize': 64,
'output_layer': 'Pretrained_Net/pool_3:0',
'input_layer': 'Pretrained_Net/ExpandDims:0',
'output_shape': 2048,
'cosine_distance_eps': 0.1
}
}
def create_model_graph(pth):
"""Creates a graph from saved GraphDef file."""
# Creates graph from saved graph_def.pb.
with tf.gfile.FastGFile( pth, 'rb') as f:
graph_def = tf.GraphDef()
graph_def.ParseFromString( f.read())
_ = tf.import_graph_def( graph_def, name='Pretrained_Net')
def _get_model_layer(sess, model_name):
# layername = 'Pretrained_Net/final_layer/Mean:0'
layername = model_params[model_name]['output_layer']
layer = sess.graph.get_tensor_by_name(layername)
ops = layer.graph.get_operations()
for op_idx, op in enumerate(ops):
for o in op.outputs:
shape = o.get_shape()
if shape._dims != []:
shape = [s.value for s in shape]
new_shape = []
for j, s in enumerate(shape):
if s == 1 and j == 0:
new_shape.append(None)
else:
new_shape.append(s)
o.__dict__['_shape_val'] = tf.TensorShape(new_shape)
return layer
def get_activations(images, sess, model_name, batch_size=50, verbose=False):
"""Calculates the activations of the pool_3 layer for all images.
Params:
-- images : Numpy array of dimension (n_images, hi, wi, 3). The values
must lie between 0 and 256.
-- sess : current session
-- batch_size : the images numpy array is split into batches with batch size
batch_size. A reasonable batch size depends on the disposable hardware.
-- verbose : If set to True and parameter out_step is given, the number of calculated
batches is reported.
Returns:
-- A numpy array of dimension (num images, 2048) that contains the
activations of the given tensor when feeding inception with the query tensor.
"""
inception_layer = _get_model_layer(sess, model_name)
n_images = images.shape[0]
if batch_size > n_images:
print("warning: batch size is bigger than the data size. setting batch size to data size")
batch_size = n_images
n_batches = n_images//batch_size + 1
pred_arr = np.empty((n_images,model_params[model_name]['output_shape']))
for i in tqdm(range(n_batches)):
if verbose:
print("\rPropagating batch %d/%d" % (i+1, n_batches), end="", flush=True)
start = i*batch_size
if start+batch_size < n_images:
end = start+batch_size
else:
end = n_images
batch = images[start:end]
pred = sess.run(inception_layer, {model_params[model_name]['input_layer']: batch})
pred_arr[start:end] = pred.reshape(-1,model_params[model_name]['output_shape'])
if verbose:
print(" done")
return pred_arr
# def calculate_memorization_distance(features1, features2):
# neigh = NearestNeighbors(n_neighbors=1, algorithm='kd_tree', metric='euclidean')
# neigh.fit(features2)
# d, _ = neigh.kneighbors(features1, return_distance=True)
# print('d.shape=',d.shape)
# return np.mean(d)
def normalize_rows(x: np.ndarray):
"""
function that normalizes each row of the matrix x to have unit length.
Args:
``x``: A numpy matrix of shape (n, m)
Returns:
``x``: The normalized (by row) numpy matrix.
"""
return np.nan_to_num(x/np.linalg.norm(x, ord=2, axis=1, keepdims=True))
def cosine_distance(features1, features2):
# print('rows of zeros in features1 = ',sum(np.sum(features1, axis=1) == 0))
# print('rows of zeros in features2 = ',sum(np.sum(features2, axis=1) == 0))
features1_nozero = features1[np.sum(features1, axis=1) != 0]
features2_nozero = features2[np.sum(features2, axis=1) != 0]
norm_f1 = normalize_rows(features1_nozero)
norm_f2 = normalize_rows(features2_nozero)
d = 1.0-np.abs(np.matmul(norm_f1, norm_f2.T))
print('d.shape=',d.shape)
print('np.min(d, axis=1).shape=',np.min(d, axis=1).shape)
mean_min_d = np.mean(np.min(d, axis=1))
print('distance=',mean_min_d)
return mean_min_d
def distance_thresholding(d, eps):
if d < eps:
return d
else:
return 1
def calculate_frechet_distance(mu1, sigma1, mu2, sigma2, eps=1e-6):
"""Numpy implementation of the Frechet Distance.
The Frechet distance between two multivariate Gaussians X_1 ~ N(mu_1, C_1)
and X_2 ~ N(mu_2, C_2) is
d^2 = ||mu_1 - mu_2||^2 + Tr(C_1 + C_2 - 2*sqrt(C_1*C_2)).
Stable version by <NAME>.
Params:
-- mu1 : Numpy array containing the activations of the pool_3 layer of the
inception net ( like returned by the function 'get_predictions')
for generated samples.
-- mu2 : The sample mean over activations of the pool_3 layer, precalcualted
on an representive data set.
-- sigma1: The covariance matrix over activations of the pool_3 layer for
generated samples.
-- sigma2: The covariance matrix over activations of the pool_3 layer,
precalcualted on an representive data set.
Returns:
-- : The Frechet Distance.
"""
mu1 = np.atleast_1d(mu1)
mu2 = np.atleast_1d(mu2)
sigma1 = np.atleast_2d(sigma1)
sigma2 = np.atleast_2d(sigma2)
assert mu1.shape == mu2.shape, "Training and test mean vectors have different lengths"
assert sigma1.shape == sigma2.shape, "Training and test covariances have different dimensions"
diff = mu1 - mu2
# product might be almost singular
covmean, _ = linalg.sqrtm(sigma1.dot(sigma2), disp=False)
if not np.isfinite(covmean).all():
msg = "fid calculation produces singular product; adding %s to diagonal of cov estimates" % eps
warnings.warn(msg)
offset = np.eye(sigma1.shape[0]) * eps
# covmean = linalg.sqrtm((sigma1 + offset).dot(sigma2 + offset))
covmean = linalg.sqrtm((sigma1 + offset).dot(sigma2 + offset))
# numerical error might give slight imaginary component
if np.iscomplexobj(covmean):
if not np.allclose(np.diagonal(covmean).imag, 0, atol=1e-3):
m = np.max(np.abs(covmean.imag))
raise ValueError("Imaginary component {}".format(m))
covmean = covmean.real
# covmean = tf.linalg.sqrtm(tf.linalg.matmul(sigma1,sigma2))
print('covmean.shape=',covmean.shape)
# tr_covmean = tf.linalg.trace(covmean)
tr_covmean = np.trace(covmean)
return diff.dot(diff) + np.trace(sigma1) + np.trace(sigma2) - 2 * tr_covmean
# return diff.dot(diff) + tf.linalg.trace(sigma1) + tf.linalg.trace(sigma2) - 2 * tr_covmean
#-------------------------------------------------------------------------------
def calculate_activation_statistics(images, sess, model_name, batch_size=50, verbose=False):
"""Calculation of the statistics used by the FID.
Params:
-- images : Numpy array of dimension (n_images, hi, wi, 3). The values
must lie between 0 and 255.
-- sess : current session
-- batch_size : the images numpy array is split into batches with batch size
batch_size. A reasonable batch size depends on the available hardware.
-- verbose : If set to True and parameter out_step is given, the number of calculated
batches is reported.
Returns:
-- mu : The mean over samples of the activations of the pool_3 layer of
the incption model.
-- sigma : The covariance matrix of the activations of the pool_3 layer of
the incption model.
"""
act = get_activations(images, sess, model_name, batch_size, verbose)
mu = np.mean(act, axis=0)
sigma = np.cov(act, rowvar=False)
return mu, sigma, act
def _handle_path_memorization(path, sess, model_name, is_checksize, is_check_png):
path = pathlib.Path(path)
files = list(path.glob('*.jpg')) + list(path.glob('*.png'))
imsize = model_params[model_name]['imsize']
# In production we don't resize input images. This is just for demo purpose.
x = np.array([np.array(img_read_checks(fn, imsize, is_checksize, imsize, is_check_png)) for fn in files])
m, s, features = calculate_activation_statistics(x, sess, model_name)
del x #clean up memory
return m, s, features
# check for image size
def img_read_checks(filename, resize_to, is_checksize=False, check_imsize = 64, is_check_png = False):
im = Image.open(str(filename))
if is_checksize and im.size != (check_imsize,check_imsize):
raise KernelEvalException('The images are not of size '+str(check_imsize))
if is_check_png and im.format != 'PNG':
raise KernelEvalException('Only PNG images should be submitted.')
if resize_to is None:
return im
else:
return im.resize((resize_to,resize_to),Image.ANTIALIAS)
def calculate_kid_given_paths(paths, model_name, model_path, feature_path=None, mm=[], ss=[], ff=[]):
''' Calculates the KID of two paths. '''
tf.reset_default_graph()
create_model_graph(str(model_path))
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
m1, s1, features1 = _handle_path_memorization(paths[0], sess, model_name, is_checksize = True, is_check_png = True)
if len(mm) != 0:
m2 = mm
s2 = ss
features2 = ff
elif feature_path is None:
m2, s2, features2 = _handle_path_memorization(paths[1], sess, model_name, is_checksize = False, is_check_png = False)
else:
with np.load(feature_path) as f:
m2, s2, features2 = f['m'], f['s'], f['features']
print('m1,m2 shape=',(m1.shape,m2.shape),'s1,s2=',(s1.shape,s2.shape))
print('starting calculating FID')
fid_value = calculate_frechet_distance(m1, s1, m2, s2)
print('done with FID, starting distance calculation')
distance = cosine_distance(features1, features2)
return fid_value, distance, m2, s2, features2
# + _kg_hide-input=true
if ComputeLB:
# FREE MEMORY
del decoded_imgs, encoded_imgs, images, encoder, decoder, autoencoder, generated
x = gc.collect()
# UNCOMPRESS OUR IMGAES
with zipfile.ZipFile("../working/images.zip","r") as z:
z.extractall("../tmp/images2/")
# COMPUTE LB SCORE
m2 = []; s2 =[]; f2 = []
user_images_unzipped_path = '../tmp/images2/'
images_path = [user_images_unzipped_path,'../input/generative-dog-images/all-dogs/all-dogs/']
public_path = '../input/dog-face-generation-competition-kid-metric-input/classify_image_graph_def.pb'
fid_epsilon = 10e-15
fid_value_public, distance_public, m2, s2, f2 = calculate_kid_given_paths(images_path, 'Inception', public_path, mm=m2, ss=s2, ff=f2)
distance_public = distance_thresholding(distance_public, model_params['Inception']['cosine_distance_eps'])
print("FID_public: ", fid_value_public, "distance_public: ", distance_public, "multiplied_public: ",
fid_value_public /(distance_public + fid_epsilon))
# REMOVE FILES TO PREVENT KERNEL ERROR OF TOO MANY FILES
# ! rm -r ../tmp
| kernel/dog-autoencoder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Reading CSV, CCTV Status of Seoul City
# ## Loading Data
# [공공데이터포털](https://data.go.kr)
#
# Recommend getting Comma Seperated Values
# ## Getting District CCTV Informations
# https://www.data.go.kr/dataset/3073216/fileData.do
# ## Load Pandas
import pandas as pd
CCTV = pd.read_csv("seoulYearlyCCTV.csv", encoding="utf-8")
CCTV.head(5)
# ## Check Data Columns
CCTV.columns
# ## Change Column Name
CCTV.rename(columns={CCTV.columns[0]: "구"}, inplace=True)
CCTV.head(5)
# ## Load Excel File - Load Population
# ### Install xlrd
# !pip install xlrd || pip3 install xlrd
# ### Loading Excel
population = pd.read_excel("seoulPopulation.xls")
population.head(5)
# ### Loading with Options
# +
population = pd.read_excel(
"seoulPopulation.xls", # FileName
header = 2, # row number of header
usecols = "B, D, G, J, N", # only load these columns (comma seperated)
encoding = "utf-8" # encoding type
)
population.head()
# -
# ## Change Column Name, Again
population.rename(
columns={
population.columns[1]: "전체인구",
population.columns[2]: "한국인",
population.columns[3]: "외국인",
population.columns[4]: "고령자"
},
inplace=True # 덮어쓰기
)
population.head()
# ## Analyze CCTV Data
CCTV.head(5)
# ### Sorting By Values
# = ORDER BY
CCTV.sort_values(by=['소계']).head(5)
CCTV.sort_values(by=['소계'],ascending=False).head(5)
# ### Add Column
CCTV['최근증가율'] = ((CCTV['2016년'] + CCTV['2015년'] + CCTV['2014년']) / CCTV['2013년도 이전']) * 100
CCTV.sort_values(by=['최근증가율'], ascending=False).head(5)
# ## Analyze Population Data
population.head(5)
wa = population.where(population['자치구'] == "합계").dropna().index
population.drop(wa, inplace=True)
population.head(5)
# Check Data is Unique
population['자치구'].unique()
population['외국인비율'] = population['외국인'] / population['전체인구'] * 100
population['고령자비율'] = population['고령자'] / population['전체인구'] * 100
population.head(5)
population.sort_values(by='전체인구',ascending=False).head(5)
population.sort_values(by='외국인',ascending=False).head(5)
population.sort_values(by='외국인비율',ascending=False).head(5)
population.sort_values(by='고령자',ascending=False).head(5)
population.sort_values(by='고령자비율',ascending=False).head(5)
# ## Merge DataFrames
population.rename(
columns={
population.columns[0]:"구"
},
inplace=True
)
population.head(5)
data = pd.merge(population, CCTV, how='inner', on="구")
data.head(5)
data.set_index("구", inplace=True)
data.head(5)
del data['2013년도 이전']
del data['2014년']
del data['2015년']
del data['2016년']
data.head()
# ## Draw Graph using Matplotlib
# ### Setting Fonts
# +
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
import platform
winPath = "c:/Windows/Fonts/malgun.ttf"
linuxFont = ""
from matplotlib import font_manager, rc
if platform.system() == 'Darwin':
rc('font', family="AppleGothic")
elif platform.system() == 'Windows':
font_name = font_manager.FontProperties(fname=winPath).get_name()
rc('font', family=font_name)
elif platform.system() == 'Linux':
print("Setting Font as", linuxFont)
rc('font', family=linuxFont)
else:
print("Woops! Failed to detect platform.system()")
# -
# ### Drawing Heatmap
# +
target_col = ['외국인비율', '고령자비율']
data_sorted = data.sort_values(by='고령자비율', ascending=False)
plt.figure(figsize=(10,10))
sns.heatmap(data_sorted[target_col], annot=True, fmt='f', linewidths=.5, cmap='RdPu')
plt.title('외국인 및 고령자 비율')
plt.show()
# -
# ### Correlation coefficient
#
# If absolute of correlation coefficient is...
# 0 <= corrcoef < 0.2: Not Related.
# 0.2 <= corrcoef < 0.5: Slightly Related.
# 0.5 <= corrcoef < 1.0: Very Related.
# #### Import Numpy
import numpy as np
# #### Getting Correlation Coefficient
np.corrcoef(data['고령자비율'], data['소계'])
# 0.26: Very Slightly Related (Negative)
np.corrcoef(data['외국인비율'], data['소계'])
# 0.05: Not Related
np.corrcoef(data['전체인구'], data['소계'])
# 0.24: Very Slightly Related
# ## Analyze CCTV and Populations
data.head()
# ### Import matplotlib
import matplotlib.pyplot as plt
# #### Setting Fonts
# +
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
import platform
winPath = "c:/Windows/Fonts/malgun.ttf"
linuxFont = ""
from matplotlib import font_manager, rc
if platform.system() == 'Darwin':
rc('font', family="AppleGothic")
elif platform.system() == 'Windows':
font_name = font_manager.FontProperties(fname=winPath).get_name()
rc('font', family=font_name)
elif platform.system() == 'Linux':
print("Setting Font as", linuxFont)
rc('font', family=linuxFont)
else:
print("Woops! Failed to detect platform.system()")
# -
# ### Drawing Graph
plt.figure()
data['소계'].plot(kind='barh',grid=True, figsize=(10,10))
plt.show()
plt.figure()
data['소계'].sort_values().plot(kind='barh',grid=True, figsize=(10,10))
plt.show()
plt.figure(figsize=(10,6))
plt.scatter(data['전체인구'],data['소계'], s=50)
plt.xlabel('인구수')
plt.ylabel('CCTV')
plt.grid()
plt.show()
# ### Run Polyfit
#
# polyfitted data must be **correlated**
fp1 = np.polyfit(data['전체인구'], data['소계'], 1)
fp1
f1 = np.poly1d(fp1)
fx = np.linspace(100000, 700000, 100)
plt.figure(figsize=(10,10))
plt.scatter(data['전체인구'], data['소계'], s=50)
plt.plot(fx, f1(fx), ls='dashed', lw=3, color='g')
plt.xlabel('인구수')
plt.ylabel('CCTV')
plt.grid()
plt.show()
# ## CCTV amount and Error Data in Graph
# +
fp1 = np.polyfit(data['전체인구'], data['소계'], 1)
f1 = np.poly1d(fp1)
fx = np.linspace(100000, 700000, 100)
data['오차'] = np.abs(data['소계'] - f1(data['전체인구']))
df_sort = data.sort_values(by='오차', ascending=False)
df_sort.head()
# +
plt.figure(figsize=(14,10))
plt.scatter(data['전체인구'], data['소계'],
c=data['오차'], s=50)
plt.plot(fx, f1(fx), ls='dashed', lw=3, color='g')
for n in range(10):
plt.text(df_sort['전체인구'][n]*1.02, df_sort['소계'][n]*0.98,
df_sort.index[n], fontsize=15)
plt.xlabel('전체인구')
plt.ylabel('인구당비율')
plt.colorbar()
plt.grid()
plt.show()
# -
| 08.Data Analysis/Data Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Finite Elements in 50 lines
#
# Taken from the Matlab code ([see this link](https://www.particleincell.com/2012/matlab-fem/)).
from IPython.core.display import HTML
css_file = 'https://raw.githubusercontent.com/ngcm/training-public/master/ipython_notebook_styles/ngcmstyle.css'
HTML(url=css_file)
# %matplotlib inline
import numpy
from matplotlib import pyplot
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
# Create a square grid
m = 11; n = 11
x = numpy.linspace(0, 1, m); y = numpy.linspace(0, 1, n)
X, Y = numpy.meshgrid(x, y)
p = numpy.vstack((numpy.ravel(X), numpy.ravel(Y))).T # locations of all nodes
triangles = numpy.array([[0, 1, m+1], [0, m+1, m]]) # First two triangles
triangles = numpy.kron(triangles, numpy.ones((m-1, 1), dtype=numpy.int)) + \
numpy.kron(numpy.ones(triangles.shape, dtype=numpy.int).T, numpy.arange(m-1)).T # Replicate vertically
triangles = numpy.kron(triangles, numpy.ones((n-1, 1), dtype=numpy.int)) + \
numpy.kron(numpy.ones(triangles.shape, dtype=numpy.int).T, m*numpy.arange(n-1)).T # All triangles
b = numpy.hstack((numpy.arange(m), numpy.arange(m, m*n, m), \
numpy.arange(2*m-1, m*n, m), numpy.arange(m*n-m+1, m*n-1))) # Boundary nodes
N = p.shape[0] # Number of nodes
T = triangles.shape[0] # Number of triangles
K = numpy.zeros((N, N)) # Should make this sparse
F = numpy.zeros((N, 1))
# Loop over the triangles; assemble the stiffness matrix and load vector
for nodes in triangles:
Pe = numpy.hstack((numpy.ones((3, 1)), p[nodes, :]))
Area = abs(numpy.linalg.det(Pe)) / 2
C = numpy.linalg.inv(Pe)
grad = C[1:3, :]
Ke = Area * numpy.dot(grad.T, grad)
Fe = Area / 3
nodes2 = numpy.outer(numpy.ones((3,1), dtype=numpy.int), nodes)
K[nodes2, nodes2.T] += Ke
F[nodes] += Fe
# Set Dirichlet boundary conditions
K[b, :] = 0; K[:, b] = 0; F[b] = 0
b2 = numpy.outer(numpy.ones(b.shape, dtype=numpy.int), b)
K[b2, b2.T] = numpy.eye(len(b))
# Solve
U = numpy.linalg.solve(K, F)
# Plot result
fig=pyplot.figure(figsize=(12,8))
ax = fig.add_subplot(111, projection='3d')
ax.plot_trisurf(p[:,0], p[:,1], U[:,0], triangles=triangles, cmap=cm.jet)
ax.view_init(elev=90,azim=0);ax.set_xlabel(r"$x$");ax.set_ylabel(r"$y$");
# Plot result
fig=pyplot.figure(figsize=(12,8))
ax=fig.add_subplot(111)
ax.tripcolor(p[:,0], p[:,1], U[:,0], triangles=triangles, cmap=cm.viridis, shading='gouraud')
ax.set_xlabel(r"$x$");ax.set_ylabel(r"$y$");
| FEEG6016 Simulation and Modelling/07a-Finite-Elements-Lab-1a-50-lines.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Fatigue failure model
# <a href="../handbook/reliability_prediction/structural_models_equations.html#modelling-of-failures-due-to-fatigue" class="back-forward-button">Go to handbook</a>
# ## Model description
#
# Fatigue is a failure mechanism incurred by cyclic loading, leading to the initiation and extension of cracks, which degrade the strength of materials and structures. We consider here case of high-cycle fatigue failure, i.e., failures that occurs after a modelled component is exposed to large numbers of load cycles. The limit state function for this type of failure can be written as:
#
# $$
# g\left( D_{cr}, A, \text{SSF}, \Theta \right) = D_{cr} - \Theta \cdot 10^{- A} {\text{SSF}}^{B} \sum_{j = 1}^{k}{N_{j} {S_{eq,j}}^{B}},
# $$
#
# This expression contains a set of variables that we consider uncertain ($D_{cr}, A, \text{SSF}, \Theta$) and parameters that we consider to be known with a sufficiently high accuracy ($B, \{S_{eq,j}, N_j\}_{j=1,\cdots,N}$). A summary of these variables and their meaning is given in {numref}`fatigue_model_inputs_table`.
#
# ```{list-table} Input variables for reliability analysis
# :header-rows: 1
# :widths: 15 45 20 20
# :name: fatigue_model_inputs_table
#
# * - Name
# - Description
# - Unit
# - Type
# * - $D_{cr}$/D_cr
# - Threshold for accumulated damage
# - $-$
# - uncertain
# * - $A$/A
# - S/N curve slope
# - $\log(N/m^2)^{-1}$
# - uncertain
# * - $B$/B
# - S/N curve intercept
# - $-$
# - deterministic
# * - SSF
# - Global stress scaling factor
# - $-$
# - uncertain
# * - $S$/S
# - Load collective distribution
# - $N/m^2$
# - deterministic
# * - $N$/N
# - Number of load cycles
# - $-$
# - deterministic
# * - $\Theta$/Theta
# - Model uncertainty
# - $-$
# - uncertain
# ```
#
# ### Load collective
#
# The load collective is the set of load events that the component was subjected to during its lifetime. The collective is denoted here by $\{S_{eq,j}, N_j\}_{j=1,\cdots,N}$. To simplify usage of this interactive tool, the user can specify a distribution from which the load collective is sampled and a number of total load cycles $N$. The generated distribution is then shown in a plot after the analysis.
#
# ## Interactive reliability prediction
#
# This page offers an interactive reliability prediction that lets the user specify the properties of all variables listed in {numref}`fatigue_model_inputs_table`. The value of **deterministic variables** can be selected with a slider. **Uncertain variables** are characterized by:
# - _Distribution_ denoted by "Dist" and can be choosen from a set of parametric probability distributions;
# - _Mean_ value denoted by "E" and can be selected with a slider;
# - _Coefficient of variation_ denoted by "C.o.V." and can be selected with a slider.
#
# ```{note}
# To run the interactive reliability prediction on this page, click the {fa}`rocket` --> {guilabel}`Live Code` button on the top of the page. Wait a few seconds until the Kernel has loaded and run the cell below with {guilabel}`Run`.
# ```
# ```{admonition} Under construction
# :class: todo
# The numerical values and bounds for the input variables are not finalized yet. Therefore, computed failure probabilities might not be representative of the considered components.
# ```
# + tags=["thebe-init", "remove-output"]
from nrpmint.booktools import fatigue_failure
# start the web user-interface
fatigue_failure.web_ui()
# -
| 02_NRPMHandbook/mechanical/models/fatigue_failure.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + provenance=[{"end_time": "Unknown", "execution_time": "Unknown", "outputs": [], "source": "", "start_time": "Unknown"}]
import main
import config
main.print_cakes(config.cake_list)
| Module_error.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="-BpEYICMx35f"
# # "Numpy 기본"
# > "numpy 기본 코드 실습(한글)"
#
# - toc: true
# - branch: master
# - badges: true
# - comments: true
# - author: YeEun
# - categories: [jupyter, python]
# + [markdown] id="RoJBQ174e18V"
# **도구 - 넘파이(NumPy)**
#
# *넘파이(NumPy)는 파이썬의 과학 컴퓨팅을 위한 기본 라이브러리입니다. 넘파이의 핵심은 강력한 N-차원 배열 객체입니다. 또한 선형 대수, 푸리에(Fourier) 변환, 유사 난수 생성과 같은 유용한 함수들도 제공합니다."
# + [markdown] id="0PJnnb8Ge18W"
# <table align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/rickiepark/handson-ml2/blob/master/tools_numpy.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />구글 코랩에서 실행하기</a>
# </td>
# </table>
# + [markdown] id="kZv9j5lye18X"
# # 배열 생성
# + [markdown] id="Z1NRiZrNe18X"
# `numpy`를 임포트해 보죠. 대부분의 사람들이 `np`로 알리아싱하여 임포트합니다:
# + id="cLHSwTL-e18X"
import numpy as np
# + [markdown] id="ohmdO906e18X"
# ## `np.zeros`
# + [markdown] id="9tqkXgL4e18X"
# `zeros` 함수는 0으로 채워진 배열을 만듭니다:
# + colab={"base_uri": "https://localhost:8080/"} id="_5-A5CIoe18Y" outputId="aec96a78-4c8f-4d0d-87a9-f1873044d189"
np.zeros(5)
# + [markdown] id="VpJ57KNXe18Y"
# 2D 배열(즉, 행렬)을 만들려면 원하는 행과 열의 크기를 튜플로 전달합니다. 예를 들어 다음은 $3 \times 4$ 크기의 행렬입니다:
# + colab={"base_uri": "https://localhost:8080/"} id="iS2ZVhqje18Y" outputId="2f058107-6de9-4f5c-8a43-176b948e563f"
np.zeros((3,4))
# + [markdown] id="8Rp3lMcie18Y"
# ## 용어
#
# * 넘파이에서 각 차원을 **축**(axis) 이라고 합니다
# * 축의 개수를 **랭크**(rank) 라고 합니다.
# * 예를 들어, 위의 $3 \times 4$ 행렬은 랭크 2인 배열입니다(즉 2차원입니다).
# * 첫 번째 축의 길이는 3이고 두 번째 축의 길이는 4입니다.
# * 배열의 축 길이를 배열의 **크기**(shape)라고 합니다.
# * 예를 들어, 위 행렬의 크기는 `(3, 4)`입니다.
# * 랭크는 크기의 길이와 같습니다.
# * 배열의 **사이즈**(size)는 전체 원소의 개수입니다. 축의 길이를 모두 곱해서 구할 수 있습니다(가령, $3 \times 4=12$).
# + colab={"base_uri": "https://localhost:8080/"} id="g5Dy159He18Y" outputId="e41ffa56-23fd-46fc-8bd8-f420428de652"
a = np.zeros((3,4))
a
# + colab={"base_uri": "https://localhost:8080/"} id="tSXp4dISe18Y" outputId="e88d5ff1-2ebd-40a4-f59d-7e21d06f60dc"
a.shape
# + colab={"base_uri": "https://localhost:8080/"} id="aMY1UjlTe18Y" outputId="78e761e0-7ec2-448b-968c-b4288c11f0bd"
a.ndim # len(a.shape)와 같습니다
# + colab={"base_uri": "https://localhost:8080/"} id="FVI25cs6e18Z" outputId="0887d837-63e7-46ae-fd7b-125d84826e11"
a.size
# + [markdown] id="HkPN3ijSe18Z"
# ## N-차원 배열
# 임의의 랭크 수를 가진 N-차원 배열을 만들 수 있습니다. 예를 들어, 다음은 크기가 `(2,3,4)`인 3D 배열(랭크=3)입니다:
# + colab={"base_uri": "https://localhost:8080/"} id="olP3UVoVe18Z" outputId="1c1ef080-44ca-4c53-9086-15c19d8b694c"
np.zeros((2,2,5))
# + [markdown] id="js_UNm5pe18Z"
# ## 배열 타입
# 넘파이 배열의 타입은 `ndarray`입니다:
# + colab={"base_uri": "https://localhost:8080/"} id="QO6hVeSze18Z" outputId="7709877a-ec3b-4d0f-daa9-92d98c2033ce"
type(np.zeros((3,4)))
# + [markdown] id="33G8lxzCe18Z"
# ## `np.ones`
# `ndarray`를 만들 수 있는 넘파이 함수가 많습니다.
#
# 다음은 1로 채워진 $3 \times 4$ 크기의 행렬입니다:
# + colab={"base_uri": "https://localhost:8080/"} id="n72gORzye18Z" outputId="95ab7176-1a8c-498f-feea-fba89092b7ff"
np.ones((3,4))
# + [markdown] id="Xbp1lBMde18Z"
# ## `np.full`
# 주어진 값으로 지정된 크기의 배열을 초기화합니다. 다음은 `π`로 채워진 $3 \times 4$ 크기의 행렬입니다.
# + id="YVzE5gANe18Z" outputId="e02e1c63-24eb-4e2b-91ba-da85281e801b"
np.full((3,4), np.pi)
# + [markdown] id="lGM5v1BKe18Z"
# ## `np.empty`
# 초기화되지 않은 $2 \times 3$ 크기의 배열을 만듭니다(배열의 내용은 예측이 불가능하며 메모리 상황에 따라 달라집니다):
# + id="LT_g08kte18a" outputId="72f17e9f-22bc-4e5c-aa6c-08d15d8f5a50"
np.empty((2,3))
# + [markdown] id="CronQ6fTe18a"
# ## np.array
# `array` 함수는 파이썬 리스트를 사용하여 `ndarray`를 초기화합니다:
# + id="qVLkDuC5e18a" outputId="83226f2a-ed8c-4a0d-da21-eedb4a23874c"
np.array([[1,2,3,4], [10, 20, 30, 40]])
# + [markdown] id="dIlBGhTke18a"
# ## `np.arange`
# 파이썬의 기본 `range` 함수와 비슷한 넘파이 `arange` 함수를 사용하여 `ndarray`를 만들 수 있습니다:
# + id="GNdq6rY1e18a" outputId="6cb6f07c-52af-44c7-d41a-072458f621e3"
np.arange(1, 5)
# + [markdown] id="MBr2EIUFe18a"
# 부동 소수도 가능합니다:
# + id="1WExGjLfe18a" outputId="0bd63be9-aeb8-443c-cf26-51c097226b53"
np.arange(1.0, 5.0)
# + [markdown] id="5bX9HW6Xe18b"
# 파이썬의 기본 `range` 함수처럼 건너 뛰는 정도를 지정할 수 있습니다:
# + id="nqPAWSN8e18b" outputId="58916f4f-38d3-44d0-a6b7-9ab9df05b62b"
np.arange(1, 5, 0.5)
# + [markdown] id="shM-hIqze18b"
# 부동 소수를 사용하면 원소의 개수가 일정하지 않을 수 있습니다. 예를 들면 다음과 같습니다:
# + id="ZS_9tMhQe18b" outputId="d8f753a3-2a20-453e-f69c-8d6c189e12a8"
print(np.arange(0, 5/3, 1/3)) # 부동 소수 오차 때문에, 최댓값은 4/3 또는 5/3이 됩니다.
print(np.arange(0, 5/3, 0.333333333))
print(np.arange(0, 5/3, 0.333333334))
# + [markdown] id="jXHF_Tise18b"
# ## `np.linspace`
# 이런 이유로 부동 소수를 사용할 땐 `arange` 대신에 `linspace` 함수를 사용하는 것이 좋습니다. `linspace` 함수는 지정된 개수만큼 두 값 사이를 나눈 배열을 반환합니다(`arange`와는 다르게 최댓값이 **포함**됩니다):
# + id="JDcNhQ-_e18b" outputId="eb2865c7-c39e-4095-d500-e237f83b7c92"
print(np.linspace(0, 5/3, 6))
# + [markdown] id="r1c0sm27e18b"
# ## `np.rand`와 `np.randn`
# 넘파이의 `random` 모듈에는 `ndarray`를 랜덤한 값으로 초기화할 수 있는 함수들이 많이 있습니다.
# 예를 들어, 다음은 (균등 분포인) 0과 1사이의 랜덤한 부동 소수로 $3 \times 4$ 행렬을 초기화합니다:
# + id="J4Mv5Dxke18b" outputId="5212a761-a650-4fff-fad3-fe639639c552"
np.random.rand(3,4)
# + [markdown] id="S0l7NAWbe18b"
# 다음은 평균이 0이고 분산이 1인 일변량 [정규 분포](https://ko.wikipedia.org/wiki/%EC%A0%95%EA%B7%9C_%EB%B6%84%ED%8F%AC)(가우시안 분포)에서 샘플링한 랜덤한 부동 소수를 담은 $3 \times 4$ 행렬입니다:
# + id="RMK3OjKXe18c" outputId="4a7a6149-d80d-44de-b991-c3b174811822"
np.random.randn(3,4)
# + [markdown] id="H8wUWW6Le18c"
# 이 분포의 모양을 알려면 맷플롯립을 사용해 그려보는 것이 좋습니다(더 자세한 것은 [맷플롯립 튜토리얼](tools_matplotlib.ipynb)을 참고하세요):
# + id="npjk6A9oe18c"
# %matplotlib inline
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="JSc7Zphfe18c" outputId="df9dd86c-41d4-41c9-b249-f90e905cd3a7"
plt.hist(np.random.rand(100000), density=True, bins=100, histtype="step", color="blue", label="rand")
plt.hist(np.random.randn(100000), density=True, bins=100, histtype="step", color="red", label="randn")
plt.axis([-2.5, 2.5, 0, 1.1])
plt.legend(loc = "upper left")
plt.title("Random distributions")
plt.xlabel("Value")
plt.ylabel("Density")
plt.show()
# + [markdown] id="VNILctSme18c"
# ## np.fromfunction
# 함수를 사용하여 `ndarray`를 초기화할 수도 있습니다:
# + id="1PPxhCvKe18c" outputId="4520cb28-963e-4d1a-db31-01e02f59ca76"
def my_function(z, y, x):
return x + 10 * y + 100 * z
np.fromfunction(my_function, (3, 2, 10))
# + [markdown] id="U-dUAGbVe18c"
# 넘파이는 먼저 크기가 `(3, 2, 10)`인 세 개의 `ndarray`(차원마다 하나씩)를 만듭니다. 각 배열은 축을 따라 좌표 값과 같은 값을 가집니다. 예를 들어, `z` 축에 있는 배열의 모든 원소는 z-축의 값과 같습니다:
#
# [[[ 0. 0. 0. 0. 0. 0. 0. 0. 0. 0.]
# [ 0. 0. 0. 0. 0. 0. 0. 0. 0. 0.]]
#
# [[ 1. 1. 1. 1. 1. 1. 1. 1. 1. 1.]
# [ 1. 1. 1. 1. 1. 1. 1. 1. 1. 1.]]
#
# [[ 2. 2. 2. 2. 2. 2. 2. 2. 2. 2.]
# [ 2. 2. 2. 2. 2. 2. 2. 2. 2. 2.]]]
#
# 위의 식 `x + 10 * y + 100 * z`에서 `x`, `y`, `z`는 사실 `ndarray`입니다(배열의 산술 연산에 대해서는 아래에서 설명합니다). 중요한 점은 함수 `my_function`이 원소마다 호출되는 것이 아니고 딱 **한 번** 호출된다는 점입니다. 그래서 매우 효율적으로 초기화할 수 있습니다.
# + [markdown] id="PHJTX0h2e18c"
# # 배열 데이터
# ## `dtype`
# 넘파이의 `ndarray`는 모든 원소가 동일한 타입(보통 숫자)을 가지기 때문에 효율적입니다. `dtype` 속성으로 쉽게 데이터 타입을 확인할 수 있습니다:
# + colab={"base_uri": "https://localhost:8080/"} id="BntmDU3me18c" outputId="a6d39949-ff4a-404f-9c5d-5156f0e8be88"
c = np.arange(1, 5)
print(c.dtype, c)
# + colab={"base_uri": "https://localhost:8080/"} id="V0qVWqW1e18d" outputId="f0233337-0c11-4579-9a77-0559015c9d62"
c = np.arange(1.0, 5.0)
print(c.dtype, c)
# + [markdown] id="FpIP98kue18d"
# 넘파이가 데이터 타입을 결정하도록 내버려 두는 대신 `dtype` 매개변수를 사용해서 배열을 만들 때 명시적으로 지정할 수 있습니다:
# + colab={"base_uri": "https://localhost:8080/"} id="ELWDvG9Ae18d" outputId="5fa42da8-c6d7-4ea9-b2a9-18cd4c630f2d"
d = np.arange(1, 5, dtype=np.complex64)
print(d.dtype, d)
# + [markdown] id="LYE1YHZQe18d"
# 가능한 데이터 타입은 `int8`, `int16`, `int32`, `int64`, `uint8`|`16`|`32`|`64`, `float16`|`32`|`64`, `complex64`|`128`가 있습니다. 전체 리스트는 [온라인 문서](http://docs.scipy.org/doc/numpy/user/basics.types.html)를 참고하세요.
#
# ## `itemsize`
# `itemsize` 속성은 각 아이템의 크기(바이트)를 반환합니다:
# + id="xOtAWh7Ee18d" outputId="e62be970-e82f-43d2-efce-17f5ccb56cf1"
e = np.arange(1, 5, dtype=np.complex64)
e.itemsize
# + [markdown] id="CLNopDire18d"
# ## `data` 버퍼
# 배열의 데이터는 1차원 바이트 버퍼로 메모리에 저장됩니다. `data` 속성을 사용해 참조할 수 있습니다(사용할 일은 거의 없겠지만요).
# + id="Xy4Uwjcze18e" outputId="30adfdc5-ae0d-421e-80d2-6745d53c78dc"
f = np.array([[1,2],[1000, 2000]], dtype=np.int32)
f.data
# + [markdown] id="xxI2ePQte18e"
# 파이썬 2에서는 `f.data`가 버퍼이고 파이썬 3에서는 memoryview입니다.
# + id="eDJLlHISe18e" outputId="be1942c3-a904-47e6-f9e1-245139eec99f"
if (hasattr(f.data, "tobytes")):
data_bytes = f.data.tobytes() # python 3
else:
data_bytes = memoryview(f.data).tobytes() # python 2
data_bytes
# + [markdown] id="tkK5xTVce18e"
# 여러 개의 `ndarray`가 데이터 버퍼를 공유할 수 있습니다. 하나를 수정하면 다른 것도 바뀝니다. 잠시 후에 예를 살펴 보겠습니다.
# + [markdown] id="y8N0o_53e18e"
# # 배열 크기 변경
#
# ## 자신을 변경
#
# `ndarray`의 `shape` 속성을 지정하면 간단히 크기를 바꿀 수 있습니다. 배열의 원소 개수는 동일하게 유지됩니다.
# + id="tUusVP4Ee18e" outputId="d871639e-ad6e-4218-8413-4a7d18bf968b"
g = np.arange(24)
print(g)
print("랭크:", g.ndim)
# + id="DGI6k-ePe18e" outputId="28447575-824f-4b5a-c0c0-e8fe9fabbf42"
g.shape = (6, 4)
print(g)
print("랭크:", g.ndim)
# + id="ZYYm1gVUe18e" outputId="adb8faea-34ee-4b89-c459-b126556b37b2"
g.shape = (2, 3, 4)
print(g)
print("랭크:", g.ndim)
# + [markdown] id="9XoR2crwe18f"
# ## `reshape`
#
# `reshape` 함수는 동일한 데이터를 가리키는 새로운 `ndarray` 객체를 반환합니다. 한 배열을 수정하면 다른 것도 함께 바뀝니다.
# + id="eY-c8dr0e18f" outputId="d6ef5ee8-7598-4959-a54f-dea4032f7e25"
g2 = g.reshape(4,6)
print(g2)
print("랭크:", g2.ndim)
# + [markdown] id="5UxABFO6e18f"
# 행 1, 열 2의 원소를 999로 설정합니다(인덱싱 방식은 아래를 참고하세요).
# + id="q9pTNitre18f" outputId="b6406ebb-49f1-4593-be6f-da52227ee1e0"
g2[1, 2] = 999
g2
# + [markdown] id="ViCL0brVe18f"
# 이에 상응하는 `g`의 원소도 수정됩니다.
# + id="FB8GNr99e18f" outputId="84d77aad-f774-4478-f314-59eabe5a8d27"
g
# + [markdown] id="uMMXIC2pe18f"
# ## `ravel`
#
# 마지막으로 `ravel` 함수는 동일한 데이터를 가리키는 새로운 1차원 `ndarray`를 반환합니다:
# + id="jfGnxvIhe18f" outputId="c53be7f6-8225-466e-8dee-eeef7eb1be3c"
g.ravel()
# + [markdown] id="2y0Wg_ZGe18f"
# # 산술 연산
#
# 일반적인 산술 연산자(`+`, `-`, `*`, `/`, `//`, `**` 등)는 모두 `ndarray`와 사용할 수 있습니다. 이 연산자는 원소별로 적용됩니다:
# + id="wTVKZhAme18f" outputId="81bc44c0-dea0-4022-dd5d-9164bf246997"
a = np.array([14, 23, 32, 41])
b = np.array([5, 4, 3, 2])
print("a + b =", a + b)
print("a - b =", a - b)
print("a * b =", a * b)
print("a / b =", a / b)
print("a // b =", a // b)
print("a % b =", a % b)
print("a ** b =", a ** b)
# + [markdown] id="kpMeyQyae18g"
# 여기 곱셈은 행렬 곱셈이 아닙니다. 행렬 연산은 아래에서 설명합니다.
#
# 배열의 크기는 같아야 합니다. 그렇지 않으면 넘파이가 브로드캐스팅 규칙을 적용합니다.
# + [markdown] id="hY8Mk8Vye18g"
# # 브로드캐스팅
# + [markdown] id="CefC9OGoe18g"
# 일반적으로 넘파이는 동일한 크기의 배열을 기대합니다. 그렇지 않은 상황에는 브로드캐시틍 규칙을 적용합니다:
#
# ## 규칙 1
#
# 배열의 랭크가 동일하지 않으면 랭크가 맞을 때까지 랭크가 작은 배열 앞에 1을 추가합니다.
# + id="33JIuE1Ve18g" outputId="51db0f5f-2476-4525-eebb-c2e1918b977f"
h = np.arange(5).reshape(1, 1, 5)
h
# + [markdown] id="kGwQkU9Qe18g"
# 여기에 `(1,1,5)` 크기의 3D 배열에 `(5,)` 크기의 1D 배열을 더해 보죠. 브로드캐스팅의 규칙 1이 적용됩니다!
# + id="Yfzln3pbe18g" outputId="fa396519-4eab-4a69-fc7c-5c5a465fa137"
h + [10, 20, 30, 40, 50] # 다음과 동일합니다: h + [[[10, 20, 30, 40, 50]]]
# + [markdown] id="05djVE0Je18g"
# ## 규칙 2
#
# 특정 차원이 1인 배열은 그 차원에서 크기가 가장 큰 배열의 크기에 맞춰 동작합니다. 배열의 원소가 차원을 따라 반복됩니다.
# + id="GwqhHlP9e18g" outputId="35bb7e32-f3e9-49cb-c127-259e783071ab"
k = np.arange(6).reshape(2, 3)
k
# + [markdown] id="fYePtBoIe18g"
# `(2,3)` 크기의 2D `ndarray`에 `(2,1)` 크기의 2D 배열을 더해 보죠. 넘파이는 브로드캐스팅 규칙 2를 적용합니다:
# + id="4bU_7EtKe18g" outputId="ed8b04f1-ba93-4ea1-8210-1e5aed1ab121"
k + [[100], [200]] # 다음과 같습니다: k + [[100, 100, 100], [200, 200, 200]]
# + [markdown] id="muwkgFQJe18h"
# 규칙 1과 2를 합치면 다음과 같이 동작합니다:
# + id="9EM-Dfake18h" outputId="9e66a1e4-36d3-44d6-88e8-8c95796a302e"
k + [100, 200, 300] # 규칙 1 적용: [[100, 200, 300]], 규칙 2 적용: [[100, 200, 300], [100, 200, 300]]
# + [markdown] id="PCjsT87ye18h"
# 또 매우 간단히 다음 처럼 해도 됩니다:
# + id="3TIhMyJMe18h" outputId="ad1e05f2-4931-4b71-a2b2-bcb872487970"
k + 1000 # 다음과 같습니다: k + [[1000, 1000, 1000], [1000, 1000, 1000]]
# + [markdown] id="vRH8YKCde18h"
# ## 규칙 3
#
# 규칙 1 & 2을 적용했을 때 모든 배열의 크기가 맞아야 합니다.
# + id="Q6gMlLdTe18h" outputId="35d328c0-237a-4401-9465-8f4e8895a340"
try:
k + [33, 44]
except ValueError as e:
print(e)
# + [markdown] id="m1xKfOI0e18h"
# 브로드캐스팅 규칙은 산술 연산 뿐만 아니라 넘파이 연산에서 많이 사용됩니다. 아래에서 더 보도록 하죠. 브로드캐스팅에 관한 더 자세한 정보는 [온라인 문서](https://docs.scipy.org/doc/numpy-dev/user/basics.broadcasting.html)를 참고하세요.
# + [markdown] id="RazGg4iFe18h"
# ## 업캐스팅
#
# `dtype`이 다른 배열을 합칠 때 넘파이는 (실제 값에 상관없이) 모든 값을 다룰 수 있는 타입으로 업캐스팅합니다.
# + id="-QpkGg3Re18h" outputId="3c80e710-b5c8-4ff0-b904-fa959224f6dc"
k1 = np.arange(0, 5, dtype=np.uint8)
print(k1.dtype, k1)
# + id="7bv6CdyTe18h" outputId="997402a6-a5ff-4a38-a24f-8a49bfe19b9f"
k2 = k1 + np.array([5, 6, 7, 8, 9], dtype=np.int8)
print(k2.dtype, k2)
# + [markdown] id="cqJAQ7Uje18i"
# 모든 `int8`과 `uint8` 값(-128에서 255까지)을 표현하기 위해 `int16`이 필요합니다. 이 코드에서는 `uint8`이면 충분하지만 업캐스팅되었습니다.
# + id="RbLcnjDhe18i" outputId="fb5e63a2-5b57-407a-f082-c66bd6405525"
k3 = k1 + 1.5
print(k3.dtype, k3)
# + [markdown] id="kW-yEKcWe18i"
# # 조건 연산자
# + [markdown] id="807ng-0se18i"
# 조건 연산자도 원소별로 적용됩니다:
# + id="_2De9VRje18i" outputId="0546d38b-d344-404c-e56f-4a4728c6ecad"
m = np.array([20, -5, 30, 40])
m < [15, 16, 35, 36]
# + [markdown] id="C-BPYqKde18i"
# 브로드캐스팅을 사용합니다:
# + id="FheSPsIIe18i" outputId="7946709f-be50-40d5-f7a4-a51de862d1a0"
m < 25 # m < [25, 25, 25, 25] 와 동일
# + [markdown] id="wwPTkGfne18i"
# 불리언 인덱싱과 함께 사용하면 아주 유용합니다(아래에서 설명하겠습니다).
# + id="NY5Qm7jne18i" outputId="96c66f25-da2e-4e4f-8f34-8042bb730624"
m[m < 25]
# + [markdown] id="KagoY9cqe18i"
# # 수학 함수와 통계 함수
# + [markdown] id="J7y3L3VSe18j"
# `ndarray`에서 사용할 수 있는 수학 함수와 통계 함수가 많습니다.
#
# ## `ndarray` 메서드
#
# 일부 함수는 `ndarray` 메서드로 제공됩니다. 예를 들면:
# + id="LIix_Jl6e18j" outputId="78a268db-f453-4f4e-e7e3-899603d5837f"
a = np.array([[-2.5, 3.1, 7], [10, 11, 12]])
print(a)
print("평균 =", a.mean())
# + [markdown] id="Ynv9rv5ye18j"
# 이 명령은 크기에 상관없이 `ndarray`에 있는 모든 원소의 평균을 계산합니다.
#
# 다음은 유용한 `ndarray` 메서드입니다:
# + id="_-kMH4Rue18j" outputId="d281ca87-22ec-4e3f-80f7-814df264f4c5"
for func in (a.min, a.max, a.sum, a.prod, a.std, a.var):
print(func.__name__, "=", func())
# + [markdown] id="6lwQHGJIe18j"
# 이 함수들은 선택적으로 매개변수 `axis`를 사용합니다. 지정된 축을 따라 원소에 연산을 적용하는데 사용합니다. 예를 들면:
# + id="3in-Q2mLe18j" outputId="d37c2d4b-21bf-4df9-f91c-43c5a7e56aa1"
c=np.arange(24).reshape(2,3,4)
c
# + id="F_AYCAo1e18j" outputId="501ae060-7889-44ad-d447-ca70d73671f6"
c.sum(axis=0) # 첫 번째 축을 따라 더함, 결과는 3x4 배열
# + id="SRf1rNPye18j" outputId="09e9394f-a745-4539-9ca5-c0d1b3f85e46"
c.sum(axis=1) # 두 번째 축을 따라 더함, 결과는 2x4 배열
# + [markdown] id="vfxUKNuze18j"
# 여러 축에 대해서 더할 수도 있습니다:
# + id="9Qiv59VYe18k" outputId="9c915533-f5f6-4014-cb95-7cf9a914f181"
c.sum(axis=(0,2)) # 첫 번째 축과 세 번째 축을 따라 더함, 결과는 (3,) 배열
# + id="odwrqqGse18k" outputId="dfa7489b-c662-4191-8621-d8c69610e6e4"
0+1+2+3 + 12+13+14+15, 4+5+6+7 + 16+17+18+19, 8+9+10+11 + 20+21+22+23
# + [markdown] id="myi5rRlle18k"
# ## 일반 함수
#
# 넘파이는 일반 함수(universal function) 또는 **ufunc**라고 부르는 원소별 함수를 제공합니다. 예를 들면 `square` 함수는 원본 `ndarray`를 복사하여 각 원소를 제곱한 새로운 `ndarray` 객체를 반환합니다:
# + id="zV_XOyCGe18k" outputId="fe881a00-5bea-4830-c0d3-2a513d594908"
a = np.array([[-2.5, 3.1, 7], [10, 11, 12]])
np.square(a)
# + [markdown] id="SMUb2Dz5e18k"
# 다음은 유용한 단항 일반 함수들입니다:
# + id="pLVfUQlee18k" outputId="9251ff04-934e-4f1e-f8e6-41f9e7629d8c"
print("원본 ndarray")
print(a)
for func in (np.abs, np.sqrt, np.exp, np.log, np.sign, np.ceil, np.modf, np.isnan, np.cos):
print("\n", func.__name__)
print(func(a))
# + [markdown] id="P6WLVBN7e18k"
# ## 이항 일반 함수
#
# 두 개의 `ndarray`에 원소별로 적용되는 이항 함수도 많습니다. 두 배열이 동일한 크기가 아니면 브로드캐스팅 규칙이 적용됩니다:
# + id="l27etEMve18k" outputId="00c6fd50-49e7-4012-c814-fee289631bea"
a = np.array([1, -2, 3, 4])
b = np.array([2, 8, -1, 7])
np.add(a, b) # a + b 와 동일
# + id="HCIGD_dOe18l" outputId="ff8b3dee-1717-4a51-c059-e028845c1f45"
np.greater(a, b) # a > b 와 동일
# + id="n-Q9AFLye18l" outputId="6cbce538-05ef-414c-c83d-af0f7d77d8e1"
np.maximum(a, b)
# + id="iY8GnA1qe18l" outputId="266a7127-8f76-455f-8abc-3efc9d4c05a5"
np.copysign(a, b)
# + [markdown] id="fFfmH5nle18l"
# # 배열 인덱싱
#
# ## 1차원 배열
#
# 1차원 넘파이 배열은 보통의 파이썬 배열과 비슷하게 사용할 수 있습니다:
# + id="7WPXxqVAe18l" outputId="16e9e96d-6c49-43e1-a7e3-68043b91576e"
a = np.array([1, 5, 3, 19, 13, 7, 3])
a[3]
# + id="KHFwjJboe18l" outputId="e14765e6-b642-48cc-82f8-e4de880d5c6f"
a[2:5]
# + id="m00dyOz4e18l" outputId="5064d90d-d1d0-4cfc-d9c1-bce410032d45"
a[2:-1]
# + id="6r4_FOPde18l" outputId="4ce2d3b8-8409-4bd1-85b6-dc23abf9c605"
a[:2]
# + id="tik9SCLte18l" outputId="422ac250-c82c-4daf-dc00-3b36330d97df"
a[2::2]
# + id="BZ-T87vOe18m" outputId="5e4c6c40-8ac2-4de5-db80-ef948010bb8f"
a[::-1]
# + [markdown] id="QZNzwoFwe18m"
# 물론 원소를 수정할 수 있죠:
# + id="vZDX4AjHe18m" outputId="3563fedc-683e-4090-c66f-e8d0d3a9da64"
a[3]=999
a
# + [markdown] id="ZkyUjKcMe18m"
# 슬라이싱을 사용해 `ndarray`를 수정할 수 있습니다:
# + id="aNJvUbxce18m" outputId="1a3d3da0-eeb9-4310-bf12-74fad4432e37"
a[2:5] = [997, 998, 999]
a
# + [markdown] id="P3DbyrLoe18m"
# ## 보통의 파이썬 배열과 차이점
#
# 보통의 파이썬 배열과 대조적으로 `ndarray` 슬라이싱에 하나의 값을 할당하면 슬라이싱 전체에 복사됩니다. 위에서 언급한 브로드캐스팅 덕택입니다.
# + id="tFSrPtaMe18m" outputId="bc23bb17-aeb6-4a0d-e3ed-e110e0897939"
a[2:5] = -1
a
# + [markdown] id="7bCrkb59e18m"
# 또한 이런 식으로 `ndarray` 크기를 늘리거나 줄일 수 없습니다:
# + id="32c-sDzBe18m" outputId="bf19e35a-8626-4906-a071-454e073751e7"
try:
a[2:5] = [1,2,3,4,5,6] # 너무 길어요
except ValueError as e:
print(e)
# + [markdown] id="a0gf3yrAe18n"
# 원소를 삭제할 수도 없습니다:
# + id="lk1Zgme_e18n" outputId="74dfa441-83dd-4baf-8ad8-6c7cbac4c9f6"
try:
del a[2:5]
except ValueError as e:
print(e)
# + [markdown] id="kYBdIw-De18n"
# 중요한 점은 `ndarray`의 슬라이싱은 같은 데이터 버퍼를 바라보는 뷰(view)입니다. 슬라이싱된 객체를 수정하면 실제 원본 `ndarray`가 수정됩니다!
# + id="yBDgH25-e18n" outputId="17888b83-8342-4303-bbe1-49af75cb7ca3"
a_slice = a[2:6]
a_slice[1] = 1000
a # 원본 배열이 수정됩니다!
# + id="B4dpMTBje18n" outputId="19c0deb3-12a7-40e8-815d-86e4db026aae"
a[3] = 2000
a_slice # 비슷하게 원본 배열을 수정하면 슬라이싱 객체에도 반영됩니다!
# + [markdown] id="90EqRFpre18n"
# 데이터를 복사하려면 `copy` 메서드를 사용해야 합니다:
# + id="7FgDTA6Xe18n" outputId="4d705b67-9553-4c8d-90ec-5b308a4ddd72"
another_slice = a[2:6].copy()
another_slice[1] = 3000
a # 원본 배열이 수정되지 않습니다
# + id="upX5iWIee18n" outputId="f83b75bb-699f-4ac3-e320-b09fdd22c55f"
a[3] = 4000
another_slice # 마찬가지로 원본 배열을 수정해도 복사된 배열은 바뀌지 않습니다
# + [markdown] id="CjHv573Je18o"
# ## 다차원 배열
#
# 다차원 배열은 비슷한 방식으로 각 축을 따라 인덱싱 또는 슬라이싱해서 사용합니다. 콤마로 구분합니다:
# + id="aAHgLjTme18o" outputId="e39742bd-6dda-4d63-c15a-278eed212274"
b = np.arange(48).reshape(4, 12)
b
# + id="2lIRE6dGe18o" outputId="833aaeef-0b6c-4921-afff-51370d85fb19"
b[1, 2] # 행 1, 열 2
# + id="KZ3AY0BJe18o" outputId="64962f41-214f-41f1-8426-0a4c7a0b3f30"
b[1, :] # 행 1, 모든 열
# + id="I5d9TK93e18o" outputId="b98f8dfa-e307-4125-d23d-44941d49e7e0"
b[:, 1] # 모든 행, 열 1
# + [markdown] id="GoovJkWie18o"
# **주의**: 다음 두 표현에는 미묘한 차이가 있습니다:
# + id="QlhhxNp8e18o" outputId="00a6aba1-8566-4768-c2d6-8cc43fd65aea"
b[1, :]
# + id="s5s8uM2ge18o" outputId="ba0a4654-7a98-4504-e90d-df6a46767c92"
b[1:2, :]
# + [markdown] id="2UU8DSuJe18o"
# 첫 번째 표현식은 `(12,)` 크기인 1D 배열로 행이 하나입니다. 두 번째는 `(1, 12)` 크기인 2D 배열로 같은 행을 반환합니다.
# + [markdown] id="tkqKX2JGe18p"
# ## 팬시 인덱싱(Fancy indexing)
#
# 관심 대상의 인덱스 리스트를 지정할 수도 있습니다. 이를 팬시 인덱싱이라고 부릅니다.
# + id="tYJcpYdRe18p" outputId="df0a6cc1-98f2-4218-f6da-1c998e6c29bf"
b[(0,2), 2:5] # 행 0과 2, 열 2에서 4(5-1)까지
# + id="BKXr4leXe18p" outputId="4affd0e7-becc-439d-e19f-fea5e3851897"
b[:, (-1, 2, -1)] # 모든 행, 열 -1 (마지막), 2와 -1 (다시 반대 방향으로)
# + [markdown] id="cTM2oZU_e18p"
# 여러 개의 인덱스 리스트를 지정하면 인덱스에 맞는 값이 포함된 1D `ndarray`를 반환됩니다.
# + id="-uREynwFe18p" outputId="85292774-dc6e-4b94-adac-84512b222bd5"
b[(-1, 2, -1, 2), (5, 9, 1, 9)] # returns a 1D array with b[-1, 5], b[2, 9], b[-1, 1] and b[2, 9] (again)
# + [markdown] id="e5EXK7gpe18p"
# ## 고차원
#
# 고차원에서도 동일한 방식이 적용됩니다. 몇 가지 예를 살펴 보겠습니다:
# + id="ZTnnUVz5e18p" outputId="053bb96f-ba56-46fe-beb7-694cb8f281e1"
c = b.reshape(4,2,6)
c
# + id="Y4DKvZhLe18p" outputId="95c32c48-ae97-4a9f-b628-f8e8fc9510fd"
c[2, 1, 4] # 행렬 2, 행 1, 열 4
# + id="1mfj_5whe18p" outputId="790cb87a-5f85-4ba7-8a9a-897755f95e4f"
c[2, :, 3] # 행렬 2, 모든 행, 열 3
# + [markdown] id="kVTY5Hx_e18q"
# 어떤 축에 대한 인덱스를 지정하지 않으면 이 축의 모든 원소가 반환됩니다:
# + id="TMowyixXe18q" outputId="5046e5ce-16b3-4152-f90b-97763961de17"
c[2, 1] # 행렬 2, 행 1, 모든 열이 반환됩니다. c[2, 1, :]와 동일합니다.
# + [markdown] id="GYaas-SIe18q"
# ## 생략 부호 (`...`)
#
# 생략 부호(`...`)를 쓰면 모든 지정하지 않은 축의 원소를 포함합니다.
# + id="2-76HqTme18q" outputId="cfc05bbb-65b5-47c7-c331-14ea926af8d7"
c[2, ...] # 행렬 2, 모든 행, 모든 열. c[2, :, :]와 동일
# + id="r0xrjZuHe18q" outputId="803b9c44-e2d6-4740-e155-457fbeb9503f"
c[2, 1, ...] # 행렬 2, 행 1, 모든 열. c[2, 1, :]와 동일
# + id="GDamNZlAe18q" outputId="ded0f0d4-c509-463a-dfa7-fb701191fc95"
c[2, ..., 3] # 행렬 2, 모든 행, 열 3. c[2, :, 3]와 동일
# + id="ep_ZLfjge18q" outputId="876a3bd4-740a-43e8-9848-f68281585e3b"
c[..., 3] # 모든 행렬, 모든 행, 열 3. c[:, :, 3]와 동일
# + [markdown] id="AQHnSY4ye18q"
# ## 불리언 인덱싱
#
# 불리언 값을 가진 `ndarray`를 사용해 축의 인덱스를 지정할 수 있습니다.
# + id="jxA7H4L4e18q" outputId="51067425-5ae4-4a1f-cfa1-3f2aa4218514"
b = np.arange(48).reshape(4, 12)
b
# + id="Eip-K1ape18r" outputId="44596b0f-20d1-4a9a-c37f-602a2d9e2eae"
rows_on = np.array([True, False, True, False])
b[rows_on, :] # 행 0과 2, 모든 열. b[(0, 2), :]와 동일
# + id="I4ZgDP1be18r" outputId="de2cb1ae-e888-4161-87c2-6141266e15ba"
cols_on = np.array([False, True, False] * 4)
b[:, cols_on] # 모든 행, 열 1, 4, 7, 10
# + [markdown] id="Mn05nb4re18r"
# ## `np.ix_`
#
# 여러 축에 걸쳐서는 불리언 인덱싱을 사용할 수 없고 `ix_` 함수를 사용합니다:
# + id="yz71pbWRe18r" outputId="a53bbc0d-b0df-4584-c5bc-85e587a04b6b"
b[np.ix_(rows_on, cols_on)]
# + id="MwNA7dnwe18r" outputId="c875a197-5121-43b3-e44b-7966be5e2140"
np.ix_(rows_on, cols_on)
# + [markdown] id="PSOIE22oe18s"
# `ndarray`와 같은 크기의 불리언 배열을 사용하면 해당 위치가 `True`인 모든 원소를 담은 1D 배열이 반환됩니다. 일반적으로 조건 연산자와 함께 사용합니다:
# + id="aZuxVpTme18s" outputId="4ced80b4-0be9-46f6-c328-428270f90cd8"
b[b % 3 == 1]
# + [markdown] id="H-xKX1mwe18s"
# # 반복
#
# `ndarray`를 반복하는 것은 일반적인 파이썬 배열을 반복한는 것과 매우 유사합니다. 다차원 배열을 반복하면 첫 번째 축에 대해서 수행됩니다.
# + id="QjtG4sb8e18s" outputId="9ee9dd9c-3b24-4a01-9e29-d14e9604a0e9"
c = np.arange(24).reshape(2, 3, 4) # 3D 배열 (두 개의 3x4 행렬로 구성됨)
c
# + id="DnW3K-U7e18s" outputId="a63639b2-1b15-4f45-9783-5505fac015d4"
for m in c:
print("아이템:")
print(m)
# + id="o3M47xuhe18s" outputId="76901c19-1616-405d-aea6-fbcf758f05fd"
for i in range(len(c)): # len(c) == c.shape[0]
print("아이템:")
print(c[i])
# + [markdown] id="pKJv5DKLe18s"
# `ndarray`에 있는 모든 원소를 반복하려면 `flat` 속성을 사용합니다:
# + id="kHoBLoLMe18s" outputId="107d506c-4343-4d71-f2f0-063a127d89b3"
for i in c.flat:
print("아이템:", i)
# + [markdown] id="_9gt9F8se18s"
# # 배열 쌓기
#
# 종종 다른 배열을 쌓아야 할 때가 있습니다. 넘파이는 이를 위해 몇 개의 함수를 제공합니다. 먼저 배열 몇 개를 만들어 보죠.
# + colab={"base_uri": "https://localhost:8080/"} id="_wMUy0VKe18t" outputId="2553528f-b921-49f0-afc7-6c6643138897"
q1 = np.full((3,4), 1.0)
q1
# + colab={"base_uri": "https://localhost:8080/"} id="LfwIPIN3e18t" outputId="3d999f66-d406-47a4-d4fa-a9cc4acc48ae"
q2 = np.full((4,4), 2.0)
q2
# + colab={"base_uri": "https://localhost:8080/"} id="6R3OwBrie18t" outputId="8378d656-91f1-4ff8-d4f4-bf1007bd49eb"
q3 = np.full((3,4), 3.0)
q3
# + [markdown] id="YbMso3gce18t"
# ## `vstack`
#
# `vstack` 함수를 사용하여 수직으로 쌓아보죠:
# + colab={"base_uri": "https://localhost:8080/"} id="DnCmoigke18t" outputId="1f874162-3d05-495e-800c-0a8f2b5ed5ea"
q4 = np.vstack((q1, q2, q3))
q4
# + colab={"base_uri": "https://localhost:8080/"} id="OFRpKbEle18u" outputId="d7612bc4-afbc-4817-a8ca-f99aa99db121"
q4.shape
# + [markdown] id="j85Z1XGAe18u"
# q1, q2, q3가 모두 같은 크기이므로 가능합니다(수직으로 쌓기 때문에 수직 축은 크기가 달라도 됩니다).
#
# ## `hstack`
#
# `hstack`을 사용해 수평으로도 쌓을 수 있습니다:
# + colab={"base_uri": "https://localhost:8080/"} id="JQa34LmHe18u" outputId="95ecfa89-5a2b-49cc-a539-0af18d51d30b"
q5 = np.hstack((q1, q3))
q5
# + colab={"base_uri": "https://localhost:8080/"} id="5Uee9Znve18u" outputId="7b7de53a-c8d4-4e9e-bfb7-7797a485724a"
q5.shape
# + [markdown] id="aZOEWaVTe18u"
# q1과 q3가 모두 3개의 행을 가지고 있기 때문에 가능합니다. q2는 4개의 행을 가지고 있기 때문에 q1, q3와 수평으로 쌓을 수 없습니다:
# + id="oE8kyqILe18u" outputId="de029567-4fe6-4266-e69a-b4eaecb45126"
try:
q5 = np.hstack((q1, q2, q3))
except ValueError as e:
print(e)
# + [markdown] id="GrlZVno5e18v"
# ## `concatenate`
#
# `concatenate` 함수는 지정한 축으로도 배열을 쌓습니다.
# + colab={"base_uri": "https://localhost:8080/"} id="Apns-G9Ue18v" outputId="6a52022b-74d7-4aad-f05f-d3c2749d40c9"
q7 = np.concatenate((q1, q2, q3), axis=0) # vstack과 동일
q7
# + colab={"base_uri": "https://localhost:8080/"} id="4BfGXHtHe18v" outputId="2b73c8a3-d0d7-4d2d-9934-33bd8ee5589f"
q7.shape
# + [markdown] id="2Me1pSQ6e18v"
# 예상했겠지만 `hstack`은 `axis=1`으로 `concatenate`를 호출하는 것과 같습니다.
# + [markdown] id="3DAW9LMoe18v"
# ## `stack`
#
# `stack` 함수는 새로운 축을 따라 배열을 쌓습니다. 모든 배열은 같은 크기를 가져야 합니다.
# + id="FqRaLFR4e18v" outputId="3ffea131-aab1-4a15-a889-9a440b84db54"
q8 = np.stack((q1, q3))
q8
# + id="34-Kd1Zue18v" outputId="c3b55166-6569-4875-db18-96f05bf9fd26"
q8.shape
# + [markdown] id="hcQsm2oae18v"
# # 배열 분할
#
# 분할은 쌓기의 반대입니다. 예를 들어 `vsplit` 함수는 행렬을 수직으로 분할합니다.
#
# 먼저 6x4 행렬을 만들어 보죠:
# + id="rXqp6ziYe18v" outputId="e01f6c9a-51f0-4a03-e822-236c2a79683b"
r = np.arange(24).reshape(6,4)
r
# + [markdown] id="DeGSFbhLe18w"
# 수직으로 동일한 크기로 나누어 보겠습니다:
# + id="dzDdSqZye18w" outputId="32a80bb3-8baf-45ae-9f61-afd8def1f5d0"
r1, r2, r3 = np.vsplit(r, 3)
r1
# + id="8w-w_Vepe18w" outputId="d1d2f8ca-d850-4fc8-a38a-aee35e74a30f"
r2
# + id="Q-sdKxXBe18x" outputId="571c8430-69ef-4bb6-fbf2-8450449314e8"
r3
# + [markdown] id="k72ED90ze18x"
# `split` 함수는 주어진 축을 따라 배열을 분할합니다. `vsplit`는 `axis=0`으로 `split`를 호출하는 것과 같습니다. `hsplit` 함수는 `axis=1`로 `split`를 호출하는 것과 같습니다:
# + id="7TuO7b-He18x" outputId="848f8013-73a5-4a9a-c892-3e23ef368fb8"
r4, r5 = np.hsplit(r, 2)
r4
# + id="UuZI-P3ce18x" outputId="4f7a3641-3541-4f4a-89a8-a3682ff3048c"
r5
# + [markdown] id="v4cQffUce18x"
# # 배열 전치
#
# `transpose` 메서드는 주어진 순서대로 축을 뒤바꾸어 `ndarray` 데이터에 대한 새로운 뷰를 만듭니다.
#
# 예를 위해 3D 배열을 만들어 보죠:
# + id="uGnmx0nte18x" outputId="b0df506f-766b-4ff0-b13a-12d4e616db78"
t = np.arange(24).reshape(4,2,3)
t
# + [markdown] id="gaYsRN8We18x"
# `0, 1, 2`(깊이, 높이, 너비) 축을 `1, 2, 0` (깊이→너비, 높이→깊이, 너비→높이) 순서로 바꾼 `ndarray`를 만들어 보겠습니다:
# + id="h8IQrU9ge18x" outputId="2f2c404d-f84b-46ca-e9cc-8f2d6005970b"
t1 = t.transpose((1,2,0))
t1
# + id="Uhk1ezmme18x" outputId="81ad764b-67b4-46ea-8dd7-85515824f672"
t1.shape
# + [markdown] id="qmTBt5Xve18y"
# `transpose` 기본값은 차원의 순서를 역전시킵니다:
# + id="xfqz8Pwre18y" outputId="2b53e787-346a-45fc-e517-a49a2abc518b"
t2 = t.transpose() # t.transpose((2, 1, 0))와 동일
t2
# + id="VJN_4ldye18y" outputId="8cced2a2-98eb-44ad-8545-3d867650de89"
t2.shape
# + [markdown] id="C0tO-R44e18y"
# 넘파이는 두 축을 바꾸는 `swapaxes` 함수를 제공합니다. 예를 들어 깊이와 높이를 뒤바꾸어 `t`의 새로운 뷰를 만들어 보죠:
# + id="4KXFTH1Ue18y" outputId="e7052bb4-1691-4bdd-a871-2757a677f6e7"
t3 = t.swapaxes(0,1) # t.transpose((1, 0, 2))와 동일
t3
# + id="a27daqefe18y" outputId="8f8948d3-6644-4ad3-90a8-d16c11335817"
t3.shape
# + [markdown] id="yFPU2UrHe18y"
# # 선형 대수학
#
# 넘파이 2D 배열을 사용하면 파이썬에서 행렬을 효율적으로 표현할 수 있습니다. 주요 행렬 연산을 간단히 둘러 보겠습니다. 선형 대수학, 벡터와 행렬에 관한 자세한 내용은 [Linear Algebra tutorial](math_linear_algebra.ipynb)를 참고하세요.
#
# ## 행렬 전치
#
# `T` 속성은 랭크가 2보다 크거나 같을 때 `transpose()`를 호출하는 것과 같습니다:
# + id="Mfm70xD_e18y" outputId="a8f842ed-85ac-43c9-fdcf-769b9e7098d0"
m1 = np.arange(10).reshape(2,5)
m1
# + id="n4-oaJHVe18z" outputId="f66a552e-3b73-40cc-a77c-1e5a6b297a6a"
m1.T
# + [markdown] id="Agm8-__ne18z"
# `T` 속성은 랭크가 0이거나 1인 배열에는 아무런 영향을 미치지 않습니다:
# + id="3bogGCzxe18z" outputId="64eec463-9bf3-4778-ee7d-5b98f802a50d"
m2 = np.arange(5)
m2
# + id="OrANJqIpe18z" outputId="f81ab50e-583c-42a7-e907-e68fd05159c4"
m2.T
# + [markdown] id="J1QJqhSUe18z"
# 먼저 1D 배열을 하나의 행이 있는 행렬(2D)로 바꾼다음 전치를 수행할 수 있습니다:
# + id="gqhmd0kve18z" outputId="4f6cb619-2366-49b2-84e2-371c69007731"
m2r = m2.reshape(1,5)
m2r
# + id="swfXpv62e18z" outputId="02dc93f2-e082-4223-d0a8-a5bb75011600"
m2r.T
# + [markdown] id="2-22jYule18z"
# ## 행렬 곱셈
#
# 두 개의 행렬을 만들어 `dot` 메서드로 행렬 [곱셈](https://ko.wikipedia.org/wiki/%ED%96%89%EB%A0%AC_%EA%B3%B1%EC%85%88)을 실행해 보죠.
# + id="f04nuoY6e180" outputId="dc0e7eba-b97c-482d-cc9c-39c75340f7ac"
n1 = np.arange(10).reshape(2, 5)
n1
# + id="itq-hJkze180" outputId="a7797da8-a3a2-4182-a046-4f3fdac26002"
n2 = np.arange(15).reshape(5,3)
n2
# + id="zU2DxAlJe180" outputId="46d63a06-6b14-4a33-e077-f614bc3113fe"
n1.dot(n2)
# + [markdown] id="ZYShi4GNe180"
# **주의**: 앞서 언급한 것처럼 `n1*n2`는 행렬 곱셈이 아니라 원소별 곱셈(또는 [아다마르 곱](https://ko.wikipedia.org/wiki/%EC%95%84%EB%8B%A4%EB%A7%88%EB%A5%B4_%EA%B3%B1)이라 부릅니다)입니다.
# + [markdown] id="m8YX1nuVe180"
# ## 역행렬과 유사 역행렬
#
# `numpy.linalg` 모듈 안에 많은 선형 대수 함수들이 있습니다. 특히 `inv` 함수는 정방 행렬의 역행렬을 계산합니다:
# + id="cw6aeVP2e180" outputId="17ed696f-2d67-4de0-cd9d-679bcbf86e80"
import numpy.linalg as linalg
m3 = np.array([[1,2,3],[5,7,11],[21,29,31]])
m3
# + id="gozSHiL6e181" outputId="3cc8de7e-cd99-4254-ca36-899660222f3e"
linalg.inv(m3)
# + [markdown] id="DtMzlPJqe181"
# `pinv` 함수를 사용하여 [유사 역행렬](https://en.wikipedia.org/wiki/Moore%E2%80%93Penrose_pseudoinverse)을 계산할 수도 있습니다:
# + id="hGwTj0cke181" outputId="b5e88378-f3c3-4317-9909-022be1c5d0dd"
linalg.pinv(m3)
# + [markdown] id="WxsbsLXCe181"
# ## 단위 행렬
#
# 행렬과 그 행렬의 역행렬을 곱하면 단위 행렬이 됩니다(작은 소숫점 오차가 있습니다):
# + id="8q6q3OB0e181" outputId="d6488784-2b7d-462f-9d6f-c68cddb3e94c"
m3.dot(linalg.inv(m3))
# + [markdown] id="FOaxxlTZe181"
# `eye` 함수는 NxN 크기의 단위 행렬을 만듭니다:
# + id="SktwewJ9e181" outputId="5c85c6b1-9bea-4bb3-c0cc-5eab89b496dd"
np.eye(3)
# + [markdown] id="999Fqh2ee181"
# ## QR 분해
#
# `qr` 함수는 행렬을 [QR 분해](https://en.wikipedia.org/wiki/QR_decomposition)합니다:
# + id="u9B-umaze181" outputId="bde2973e-8be9-4a15-f353-71dd0309d53e"
q, r = linalg.qr(m3)
q
# + id="CyuFQfOMe182" outputId="54547e10-7a7f-4720-e9af-d4e020ac0881"
r
# + id="WR02Ldn1e182" outputId="ae1f5927-5f79-4094-f71e-03d81b46bb9b"
q.dot(r) # q.r는 m3와 같습니다
# + [markdown] id="HWQjHhtce182"
# ## 행렬식
#
# `det` 함수는 [행렬식](https://en.wikipedia.org/wiki/Determinant)을 계산합니다:
# + id="xeFYRyTJe182" outputId="4fe5038b-cfa9-4fd1-be2e-246900b9b004"
linalg.det(m3) # 행렬식 계산
# + [markdown] id="Qow8U8rNe182"
# ## 고윳값과 고유벡터
#
# `eig` 함수는 정방 행렬의 [고윳값과 고유벡터](https://en.wikipedia.org/wiki/Eigenvalues_and_eigenvectors)를 계산합니다:
# + id="pe7tS7sVe182" outputId="4595af27-d671-48ac-c501-45cd12b86916"
eigenvalues, eigenvectors = linalg.eig(m3)
eigenvalues # λ
# + id="kCnbXmZme182" outputId="35ad562b-8b6a-48a0-9c07-eb028852d9d4"
eigenvectors # v
# + id="HtUftah9e182" outputId="94b626d4-c88d-4f4f-d81f-b573104f7f27"
m3.dot(eigenvectors) - eigenvalues * eigenvectors # m3.v - λ*v = 0
# + [markdown] id="tgc2QJiye182"
# ## 특잇값 분해
#
# `svd` 함수는 행렬을 입력으로 받아 그 행렬의 [특잇값 분해](https://en.wikipedia.org/wiki/Singular_value_decomposition)를 반환합니다:
# + id="WlOalB-3e182" outputId="7b8a941a-7ac5-4357-eabc-7318517b9ee8"
m4 = np.array([[1,0,0,0,2], [0,0,3,0,0], [0,0,0,0,0], [0,2,0,0,0]])
m4
# + id="n6NUhkP6e183" outputId="4f30eef0-f987-44e6-8499-329997206fc9"
U, S_diag, V = linalg.svd(m4)
U
# + id="PikS0qP3e183" outputId="3ea8c6c5-224e-4179-ca8d-4ec516984cf0"
S_diag
# + [markdown] id="3V4rnjaJe183"
# `svd` 함수는 Σ의 대각 원소 값만 반환합니다. 전체 Σ 행렬은 다음과 같이 만듭니다:
# + id="K5FGxeWSe183" outputId="2ca18a39-67ab-45b4-aca6-88bdc42e9860"
S = np.zeros((4, 5))
S[np.diag_indices(4)] = S_diag
S # Σ
# + id="j6tHlysse183" outputId="724119cd-20ba-48c9-9495-a4647212b688"
V
# + id="eK_qUUaHe183" outputId="0ea1751e-05c5-410c-dd8f-702b53e8da15"
U.dot(S).dot(V) # U.Σ.V == m4
# + [markdown] id="DaYdGDcYe184"
# ## 대각원소와 대각합
# + id="LTKq5Gnqe184" outputId="44b5e88c-d3c6-4af2-9a63-289057d9af70"
np.diag(m3) # m3의 대각 원소입니다(왼쪽 위에서 오른쪽 아래)
# + id="tieRNOfne184" outputId="5fec009c-503c-4ddc-8d08-a75dc8392323"
np.trace(m3) # np.diag(m3).sum()와 같습니다
# + [markdown] id="DbDtaqiOe184"
# ## 선형 방정식 풀기
# + [markdown] id="zXycDUd2e184"
# `solve` 함수는 다음과 같은 선형 방정식을 풉니다:
#
# * $2x + 6y = 6$
# * $5x + 3y = -9$
# + id="ZLT6-qxGe184" outputId="25c0b5f9-709e-4d73-eecf-5c450ccc6ed3"
coeffs = np.array([[2, 6], [5, 3]])
depvars = np.array([6, -9])
solution = linalg.solve(coeffs, depvars)
solution
# + [markdown] id="VDY6tt_1e184"
# solution을 확인해 보죠:
# + id="lZ0MyYele184" outputId="8536d5b6-8147-4329-aa5b-8b938ca2c656"
coeffs.dot(solution), depvars # 네 같네요
# + [markdown] id="57LNnFCIe184"
# 좋습니다! 다른 방식으로도 solution을 확인해 보죠:
# + id="Q90KVIO1e185" outputId="0ce8ee47-80bb-47f5-dafb-90c11ef483d8"
np.allclose(coeffs.dot(solution), depvars)
# + [markdown] id="98NehFb9e185"
# # 벡터화
#
# 한 번에 하나씩 개별 배열 원소에 대해 연산을 실행하는 대신 배열 연산을 사용하면 훨씬 효율적인 코드를 만들 수 있습니다. 이를 벡터화라고 합니다. 이를 사용하여 넘파이의 최적화된 성능을 활용할 수 있습니다.
#
# 예를 들어, $sin(xy/40.5)$ 식을 기반으로 768x1024 크기 배열을 생성하려고 합니다. 중첩 반복문 안에 파이썬의 math 함수를 사용하는 것은 **나쁜** 방법입니다:
# + id="Y87wd2Foe185"
import math
data = np.empty((768, 1024))
for y in range(768):
for x in range(1024):
data[y, x] = math.sin(x*y/40.5) # 매우 비효율적입니다!
# + [markdown] id="Shmp3lcce185"
# 작동은 하지만 순수한 파이썬 코드로 반복문이 진행되기 때문에 아주 비효율적입니다. 이 알고리즘을 벡터화해 보죠. 먼저 넘파이 `meshgrid` 함수로 좌표 벡터를 사용해 행렬을 만듭니다.
# + id="4sx7m6q8e185" outputId="4f29411c-0f4f-414b-8a20-6c8aabebf188"
x_coords = np.arange(0, 1024) # [0, 1, 2, ..., 1023]
y_coords = np.arange(0, 768) # [0, 1, 2, ..., 767]
X, Y = np.meshgrid(x_coords, y_coords)
X
# + id="DT263XJ7e185" outputId="14470c46-b941-4aba-f823-18f67cd43d04"
Y
# + [markdown] id="66oJw5gCe185"
# 여기서 볼 수 있듯이 `X`와 `Y` 모두 768x1024 배열입니다. `X`에 있는 모든 값은 수평 좌표에 해당합니다. `Y`에 있는 모든 값은 수직 좌표에 해당합니다.
#
# 이제 간단히 배열 연산을 사용해 계산할 수 있습니다:
# + id="3Ul0Rm1we185"
data = np.sin(X*Y/40.5)
# + [markdown] id="hktPLWYXe185"
# 맷플롯립의 `imshow` 함수를 사용해 이 데이터를 그려보죠([matplotlib tutorial](tools_matplotlib.ipynb)을 참조하세요).
# + id="V_6f0Ok3e185" outputId="49ebca9f-e97f-4cc0-82c1-5ea0f5419257"
import matplotlib.pyplot as plt
import matplotlib.cm as cm
fig = plt.figure(1, figsize=(7, 6))
plt.imshow(data, cmap=cm.hot)
plt.show()
# + [markdown] id="eqo8cB5Be186"
# # 저장과 로딩
#
# 넘파이는 `ndarray`를 바이너리 또는 텍스트 포맷으로 손쉽게 저장하고 로드할 수 있습니다.
#
# ## 바이너리 `.npy` 포맷
#
# 랜덤 배열을 만들고 저장해 보죠.
# + id="Is97P60le186" outputId="f4af9a53-3eb1-4698-eb47-6b783953557c"
a = np.random.rand(2,3)
a
# + id="opu3vhLce186"
np.save("my_array", a)
# + [markdown] id="5HutN5dhe186"
# 끝입니다! 파일 이름의 확장자를 지정하지 않았기 때문에 넘파이는 자동으로 `.npy`를 붙입니다. 파일 내용을 확인해 보겠습니다:
# + id="0C9U-FXAe186" outputId="fee57006-650a-44d6-df5d-0dea3e772a86"
with open("my_array.npy", "rb") as f:
content = f.read()
content
# + [markdown] id="6z0USMaPe186"
# 이 파일을 넘파이 배열로 로드하려면 `load` 함수를 사용합니다:
# + id="fsrz9PSbe186" outputId="66437fc6-5d10-43cf-a6c5-cf42cbe065ca"
a_loaded = np.load("my_array.npy")
a_loaded
# + [markdown] id="S5LF3FoDe186"
# ## 텍스트 포맷
#
# 배열을 텍스트 포맷으로 저장해 보죠:
# + id="smuuC8MCe186"
np.savetxt("my_array.csv", a)
# + [markdown] id="-nCCfjGme187"
# 파일 내용을 확인해 보겠습니다:
# + id="YOIiv-0We187" outputId="ffd121d9-4887-4548-feb0-f52adc4a01ff"
with open("my_array.csv", "rt") as f:
print(f.read())
# + [markdown] id="dKKyh2C9e189"
# 이 파일은 탭으로 구분된 CSV 파일입니다. 다른 구분자를 지정할 수도 있습니다:
# + id="juoMnvj6e189"
np.savetxt("my_array.csv", a, delimiter=",")
# + [markdown] id="ay6LGthDe189"
# 이 파일을 로드하려면 `loadtxt` 함수를 사용합니다:
# + id="qvOpC5pce189" outputId="621021f4-c475-4e0b-a4b7-23fbe53b45ce"
a_loaded = np.loadtxt("my_array.csv", delimiter=",")
a_loaded
# + [markdown] id="i_w7PfvEe189"
# ## 압축된 `.npz` 포맷
#
# 여러 개의 배열을 압축된 한 파일로 저장하는 것도 가능합니다:
# + id="AQ-R_KRle189" outputId="cc93ffa4-082e-4be7-9298-1cb87c425faf"
b = np.arange(24, dtype=np.uint8).reshape(2, 3, 4)
b
# + id="04bYxTrse189"
np.savez("my_arrays", my_a=a, my_b=b)
# + [markdown] id="GpSfxRzTe18-"
# 파일 내용을 확인해 보죠. `.npz` 파일 확장자가 자동으로 추가되었습니다.
# + id="TeH7X0vqe18-" outputId="6d2a8c13-f2d7-481e-9df9-0f126301e7e9"
with open("my_arrays.npz", "rb") as f:
content = f.read()
repr(content)[:180] + "[...]"
# + [markdown] id="8K3mOEd4e18-"
# 다음과 같이 이 파일을 로드할 수 있습니다:
# + id="skFzRE2Ye18-" outputId="6d9d4538-923d-4db8-e260-1996b6e0eb99"
my_arrays = np.load("my_arrays.npz")
my_arrays
# + [markdown] id="nxxkXXA-e18-"
# 게으른 로딩을 수행하는 딕셔너리와 유사한 객체입니다:
# + id="tS3-o9Sge18-" outputId="8ef04620-9fb0-46fa-a93e-0a34fa5ea955"
my_arrays.keys()
# + id="BGhMtxg6e18-" outputId="33fc035e-6c4d-43e4-e6d2-f71d948d062c"
my_arrays["my_a"]
# + [markdown] id="nm8nVLATe18-"
# # 그 다음은?
#
# 넘파이 기본 요소를 모두 배웠지만 훨씬 더 많은 기능이 있습니다. 이를 배우는 가장 좋은 방법은 넘파이를 직접 실습해 보고 훌륭한 [넘파이 문서](http://docs.scipy.org/doc/numpy/reference/index.html)에서 필요한 함수와 기능을 찾아 보세요.
| _notebooks/2022-03-11-numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Move Function
#
# Now that you know how a robot uses sensor measurements to update its idea of its own location, let's see how we can incorporate motion into this location. In this notebook, let's go over the steps a robot takes to help localize itself from an initial, uniform distribution to sensing, moving and updating that distribution.
#
# We include the `sense` function that you've seen, which updates an initial distribution based on whether a robot senses a grid color: red or green.
#
# Next, you're tasked with writing a function `move` that incorporates motion into the distribution. As seen below, **one motion `U= 1` to the right, causes all values in a distribution to shift one grid cell to the right.**
#
# <img src='images/motion_1.png' width=50% height=50% />
#
# First let's include our usual resource imports and display function.
# importing resources
import matplotlib.pyplot as plt
import numpy as np
# A helper function for visualizing a distribution.
def display_map(grid, bar_width=1):
if(len(grid) > 0):
x_labels = range(len(grid))
plt.bar(x_labels, height=grid, width=bar_width, color='b')
plt.xlabel('Grid Cell')
plt.ylabel('Probability')
plt.ylim(0, 1) # range of 0-1 for probability values
plt.title('Probability of the robot being at each cell in the grid')
plt.xticks(np.arange(min(x_labels), max(x_labels)+1, 1))
plt.show()
else:
print('Grid is empty')
# You are given the initial variables and the complete `sense` function, below.
# +
# given initial variables
p=[0, 1, 0, 0, 0]
# the color of each grid cell in the 1D world
world=['green', 'red', 'red', 'green', 'green']
# Z, the sensor reading ('red' or 'green')
Z = 'red'
pHit = 0.6
pMiss = 0.2
# You are given the complete sense function
def sense(p, Z):
''' Takes in a current probability distribution, p, and a sensor reading, Z.
Returns a *normalized* distribution after the sensor measurement has been made, q.
This should be accurate whether Z is 'red' or 'green'. '''
q=[]
# loop through all grid cells
for i in range(len(p)):
# check if the sensor reading is equal to the color of the grid cell
# if so, hit = 1
# if not, hit = 0
hit = (Z == world[i])
q.append(p[i] * (hit * pHit + (1-hit) * pMiss))
# sum up all the components
s = sum(q)
# divide all elements of q by the sum to normalize
for i in range(len(p)):
q[i] = q[i] / s
return q
# Commented out code for measurements
# for k in range(len(measurements)):
# p = sense(p, measurements)
# -
# ### QUIZ: Program a function that returns a new distribution q, shifted to the right by the motion (U) units.
#
# This function should shift a distribution with the motion, U. Keep in mind that this world is cyclic and that if U=0, q should be the same as the given p. You should see all the values in `p` are moved to the right by 1, for U=1.
# +
## TODO: Complete this move function so that it shifts a probability distribution, p
## by a given motion, U
def move(p, U):
q=[]
# Your code here
for i in range (len(p)):
q.append(p[i-U %len(p)])
return q
p = move(p,1)
print(p)
display_map(p)
# -
| Lessons&CourseWorks/3.ObjectTracking&Localization/2.RobotLocalization/6.MoveFunction/1. Move Function, exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <center> Santiment Hands On Blockchain Analytics Tutorial </center>
# # <center><b> Pumps&Dumps on Centralized Exchanges </center>
# <NAME> <EMAIL>
# ## Welcome to Santiment Hands On Tutorial!
# Next hour we will be speaking about:
# - How in Santiment we think about Ethereum data
# - What is the difference between Centralized and Decentralized exchanges
# - Pumps on centralized Exchanges
# - Conclusions and thoughts
# ## <b> How we think about Ethereum data
# <b> Challenges </b>
# - Dealing with LARGE amount of data
# - Crunching different blockchain types (UTXO and account-based)
# - Maintaining real-time systems
# - Providing fault-tolerance
# Data Engineering starts with quering blockchain fullnode and ends in the database with analyzable data format.
# How to export the Data: https://github.com/santiment/eth-exporter
# Think about Ethereum data from transactions perspective (<b>transaction</b> = unit of data)
#
# <b>Transfer Transaction:</b>
# - Timestamp (datetime or blockNumber)
# - Sender
# - Receiver
# - Sended amount
# - Token (ETH, ERC20, etc)
# - TransactionHash
# ## <b> Centralized and Decentralized Exchanges
# <b>Decentralized Exchanges (DEX):</b>
# - is a contract stored onchain and executed by EVM (in a case of Ethereum)
# - to bet you need to send the transaction
# - every trade is stored in the blockchain
# - do not owe users' funds
# - 100% transparent
# - may have or may not have UI
#
# DEXes:
# - IDEX
# - Kyber
# - Oasis etc..
#
#
# *may not work with ETH but with WETH or KyberETH
# <b>Сentralized Exchanges (CEX):</b>
# - is a real-world entity
# - owe custodian addresses (hot/cold wallets)
# - trades happen on the exchange engine and are not stored in the blockchain
# - do owe users' funds
# - non transparent
# - intercation via UI
#
# CEXes:
# - [Binance](https://etherscan.io/address/0x3f5ce5fbfe3e9af3971dd833d26ba9b5c936f0be)
# - [Bitfinex](https://etherscan.io/address/0x876eabf441b2ee5b5b0554fd502a8e0600950cfa)
# - [Huobi](https://etherscan.io/address/0xd8a83b72377476d0a66683cde20a8aad0b628713)
# - [Coinbase](https://etherscan.io/address/0x89135c5ea509a1395287ddcdeb1ec307aed78c15) etc..
# <img src="exchanges.png">
# <b> Deposit and Withdrawal addresses</b>
# https://etherscan.io/address/0xc96d9e6361d344781eae1314b306cfbd73696606
# ## Setup
# ### [Sanpy](https://github.com/santiment/sanpy)
# +
# # ! pip3 install sanpy
# +
# # ! pip3 install fbprophet
# +
import pandas as pd
import numpy as np
import datetime
import matplotlib.pyplot as plt
import san
from san.backtest import Backtest
from fbprophet import Prophet
from utils import fancy_plot, get_san_metric
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
# -
from keys import KEY
san.ApiConfig.api_key = KEY
# +
from_date = '2017-01-01'
to_date = '2019-11-01'
asset = 'ethereum'
interval = '1d'
# create batch object
batch = san.Batch()
# create a request
batch.get(
f'daily_active_addresses/{asset}',
from_date=from_date,
to_date=to_date,
interval=interval
)
batch.get(
f'daily_active_deposits/{asset}',
from_date=from_date,
to_date=to_date,
interval=interval
)
batch.get(
f'prices/{asset}',
from_date=from_date,
to_date=to_date,
interval=interval
)
# execute the request
[daa, dad, price] = batch.execute()
# merge dataframes
data = daa.rename(columns={'value':'activeAddresses'}).join(dad).join(price['priceUsd'])
# take a look
data.head()
# -
# Historical glimpse
fancy_plot(data.rolling(3).mean(), ['activeAddresses', 'activeDeposits'], 'priceUsd')
# ## <b> Pumps on Centralized Exchanges
# - no way to see what's inside CEX
# - but it's possible to observe the activity of CEX' wallets
#
# **Assumption:**
# There's an abnormal onchain activity before the pump.
#
# **Metric to observe**: exchange flow balance
# +
from_date = '2018-06-01' # skip bullrun
to_date = '2019-11-01'
asset = 'santiment'
ex_flow = get_san_metric(from_date, to_date, 'exchange_funds_flow', asset, interval='1d', iterate_over_days=700)
price = get_san_metric(from_date, to_date, 'prices', asset, interval='1h', iterate_over_days=120)
data = price.join(ex_flow).fillna(0)
data.head()
# -
fancy_plot(data, ['priceUsd'], 'inOutDifference')
# <b> How to define abnormal activity? </b>
# - Statistical way (Granger test, Event study, etc)
# - Backtest it!
# In order to create backtesting strategy you have to define 2 things:
# - when to BUY (1)
# - when to SELL (2)
#
# **1)** In order to specify **"entry point"**: create a signal on top of the metric (abnormal exchange flow => buy!)
#
# How to define an "abnormal" behaviour of the metric?
# - simple method (>= rolling mean value; outstanding values)
# - Facebook approach (actual value is outside the predicted confidence interval)
# - [Google approach](https://drive.google.com/file/d/1TyfMYV_hUmHbAKdY6iSnH_LqGkid3Srt/view)
#
#
# **2)** (tricky one) In order to specify **exit point**:
# - naive way (price thresholds)
# - signalling (create signals on top of other metrics)
# - ML approach (predict tops, haven't try it yet!)
# - your ideas? :)
# ## Detect anomalies: [Prophet](https://github.com/facebook/prophet/tree/master/python)
# <img src="prophet.png">
ex_flow_train = get_san_metric(
start='2018-01-01',
end='2019-11-01',
metric='exchange_funds_flow',
asset='santiment',
interval='1d',
iterate_over_days=700
)
# Transform data to prophet-readable format
ex_flow_train = ex_flow_train.reset_index()
ex_flow_train.columns = ['ds', 'y']
# +
# Create and fit the model
model = Prophet(seasonality_mode='additive')
model.fit(ex_flow_train[ex_flow_train['ds'] < ex_flow.index[0]])
# +
# Make predictions for the next 30 days
future = model.make_future_dataframe(periods=30)
pred = model.predict(future)
# -
pred.head()
# +
# try different confidence intervals
gap = (datetime.datetime.strptime(to_date, '%Y-%m-%d') - datetime.datetime.strptime(from_date, '%Y-%m-%d')).days
predicted = pd.DataFrame(None)
for month in range(gap // 30 + 1):
model = Prophet(seasonality_mode='additive', interval_width=0.25)
model.fit(ex_flow_train[ex_flow_train['ds'] < ex_flow.index[month * 30]])
future = model.make_future_dataframe(periods=30)
pred = model.predict(future.iloc[-30:])
predicted = predicted.append(
pred[['ds', 'yhat_lower', 'yhat_upper']]
)
predicted = predicted.rename(columns={'ds': 'datetime'}).set_index('datetime')
# -
data_predicted = data.join(predicted, how='left')
fancy_plot(data_predicted.ffill(), ['inOutDifference', 'yhat_lower', 'yhat_upper'])
# ### Simple backtest
# Lets use **sanpy backtest** tool.
#
# Sanpy backtest is a super-simple backtest tool that aims to backtest a given strategy on a given asset's price.
#
# It takes vector of bools for every datepoint. "True" indicates buying or hodling asset on a given time interval and "False" stands for selling.
# Hourly price returns
data_predicted["returns"] = data_predicted['priceUsd'].pct_change().fillna(0)
# Created Strategy
data_predicted['signal'] = (data_predicted['inOutDifference'] < data_predicted['yhat_lower']) | (data_predicted['inOutDifference'] > data_predicted['yhat_upper'])
fancy_plot(
data_predicted,
['priceUsd'],
'inOutDifference',
signals=data_predicted[data_predicted['signal']==True].index
)
# +
# Create a simple strategy creation function.
# The idea is to take signals generated from the metric and
# create the "hodling vector" from it.
# Relies on simple stop-loss strategy.
def create_strategy(
df:pd.DataFrame,
price_name:str,
signal_name:str,
gain_to_exit:float,
loose_to_exit:float,
max_intervals_to_hodl=np.inf
):
strategy = df[signal_name].copy()
state = {'price':None, 'hodl_days': None}
for t in range(len(strategy)):
if strategy[t] and not state['price']: # if signal and asset is not already bought
# buy asset
state['price'] = df[price_name][t]
state['hodl_intervals'] = 0
elif state['price']: # if asset is already bought
if df[price_name][t] > (1 + gain_to_exit) * state['price'] or \
df[price_name][t] < (1 - loose_to_exit) * state['price'] or \
state['hodl_intervals'] > max_intervals_to_hodl:
# sell asset
state['price'], state['hodl_intervals'] = None, None
strategy[t] = False
else:
# keep hodling
strategy[t] = True
state['hodl_intervals'] += 1
return strategy
# -
# Hodling the asset (benchmark) gives **-75.76%**
# A kind of grid-search for finding best params:
#
# Running the code takes ~10min so I've precomputed results.
# +
# # %%time
# price_name = 'priceUsd'
# signals_name = 'signal'
# backtest_results = []
# for gain_to_exit in np.arange(0.01, 1.02, 0.05):
# for loose_to_exit in np.arange(0.01, 1.02, 0.05):
# data_predicted['trades'] = create_strategy(data_predicted, price_name, signals_name, gain_to_exit, loose_to_exit)
# bt = Backtest(data_predicted.returns.dropna(), data_predicted.trades)
# backtest_results.append({
# 'gain_to_exit': round(gain_to_exit, 3),
# 'loose_to_exit': round(loose_to_exit, 3),
# 'returns': bt.get_return(decimals=4)
# })
# backtest_results = pd.DataFrame(backtest_results)
# -
#backtest_results.to_csv('backtest_results_prophet.csv', index=False)
backtest_results = pd.read_csv('backtest_results_prophet.csv')
# +
# Create a heatmap of strategy outcomes:
backtest_results_df = backtest_results[['gain_to_exit', 'loose_to_exit', 'returns']]\
.set_index(['gain_to_exit', 'loose_to_exit'])\
.unstack('loose_to_exit')
backtest_results_df.columns = backtest_results_df.columns.droplevel()
plt.rcParams['figure.figsize'] = (14.0, 12.0)
plt.pcolor(backtest_results_df, cmap="seismic", edgecolors='k', linewidths=0.3)
plt.yticks(np.arange(0.5, len(backtest_results_df.index), 1), backtest_results_df.index)
plt.xticks(np.arange(0.5, len(backtest_results_df.columns), 1), backtest_results_df.columns, rotation='vertical')
plt.colorbar();
# -
# Let's estimate average strategy results Vs benchmark (hodling)
fig, ax1 = plt.subplots(figsize=(10,6))
ax1.plot(
[round(i,2) for i in np.arange(0,1.01,0.05)],
[pd.DataFrame(backtest_results)['returns'].quantile(i) for i in np.arange(0,1.01,0.05)],
label='strategy perfomance dist'
)
ax1.hlines(bt.get_return_benchmark(), 0, 1, label='hodling')
plt.grid()
plt.legend();
# ## Detect anomalies: naive way
# +
# Create signals
ex_flow_train['quantile90'] = ex_flow_train['y'].rolling(90, center=False).quantile(0.9)
ex_flow_train['signal'] = ex_flow_train.apply(
lambda row: True if row['y'] >= row['quantile90'] and row['quantile90'] != np.nan else False,
axis=1
)
data_simplestr = data.join(ex_flow_train.set_index('ds')[['signal']], how='left')
data_simplestr['signal'] = data_simplestr['signal'].fillna(False)
data_simplestr["returns"] = data_simplestr['priceUsd'].pct_change().fillna(0)
# -
fancy_plot(
data_simplestr,
['priceUsd'],
'inOutDifference',
signals=data_simplestr[data_simplestr['signal']==True].index
)
# +
# # %%time
# price_name = 'priceUsd'
# signals_name = 'signal'
# backtest_results = []
# for gain_to_exit in np.arange(0.01, 1.02, 0.05):
# for loose_to_exit in np.arange(0.01, 1.02, 0.05):
# data_simplestr['trades'] = create_strategy(data_simplestr, price_name, signals_name, gain_to_exit, loose_to_exit)
# bt = Backtest(data_simplestr.returns.dropna(), data_simplestr.trades)
# backtest_results.append({
# 'gain_to_exit': round(gain_to_exit, 3),
# 'loose_to_exit': round(loose_to_exit, 3),
# 'returns': bt.get_return(decimals=4)
# })
# backtest_results = pd.DataFrame(backtest_results)
# +
# Again use precomputed outcomes
#backtest_results.to_csv('backtest_results_simplestr90.csv', index=False)
backtest_results = pd.read_csv('backtest_results_simplestr90.csv')
# +
backtest_results_df = backtest_results[['gain_to_exit', 'loose_to_exit', 'returns']]\
.set_index(['gain_to_exit', 'loose_to_exit'])\
.unstack('loose_to_exit')
backtest_results_df.columns = backtest_results_df.columns.droplevel()
plt.rcParams['figure.figsize'] = (14.0, 12.0)
plt.pcolor(backtest_results_df, cmap="seismic", edgecolors='k', linewidths=0.3)
plt.yticks(np.arange(0.5, len(backtest_results_df.index), 1), backtest_results_df.index)
plt.xticks(np.arange(0.5, len(backtest_results_df.columns), 1), backtest_results_df.columns, rotation='vertical')
plt.colorbar();
# -
fig, ax1 = plt.subplots(figsize=(10,6))
ax1.plot(
[round(i,2) for i in np.arange(0,1.01,0.05)],
[pd.DataFrame(backtest_results)['returns'].quantile(i) for i in np.arange(0,1.01,0.05)],
label='strategy perfomance dist'
)
ax1.hlines(bt.get_return_benchmark(), 0, 1, label='hodling')
plt.grid()
plt.legend();
# ## OHLC pricing data: use hourly highest prices
# This section uses hourly highest price data as pricesource. The assumption is that you're able to
# spot exit points perfectly. To be more honest to other approaches lets also assume that the asset is bought
# in the worst moment.
# +
# Prepare the data
ohlc = get_san_metric(
start='2018-06-01',
end='2019-11-01',
metric='ohlc',
asset='santiment',
interval='1h',
iterate_over_days=120
)
high_price = ohlc[['highPriceUsd']]
high_price.ix[high_price[high_price.highPriceUsd == 0].index, 'highPriceUsd'] = 0.187765 # Fix missing point
high_price['returns'] = high_price['highPriceUsd'].pct_change().fillna(0)
high_price = high_price.join(ex_flow_train.set_index('ds')[['y', 'signal']], how='left').fillna(False)
# +
# # %%time
# price_name = 'highPriceUsd'
# signals_name = 'signal'
# backtest_results = []
# for gain_to_exit in np.arange(0.01, 1.02, 0.05):
# for loose_to_exit in np.arange(0.01, 1.02, 0.05):
# high_price['trades'] = create_strategy(high_price, price_name, signals_name, gain_to_exit, loose_to_exit)
# bt = Backtest(high_price.returns.dropna(), high_price.trades)
# backtest_results.append({
# 'gain_to_exit': round(gain_to_exit, 3),
# 'loose_to_exit': round(loose_to_exit, 3),
# 'returns': bt.get_return(decimals=4)
# })
# backtest_results = pd.DataFrame(backtest_results)
# +
# Again use precomputed outcomes
#backtest_results.to_csv('backtest_results_high_price_simplestr.csv', index=False)
backtest_results = pd.read_csv('backtest_results_high_price_simplestr.csv')
# +
backtest_results_df = backtest_results[['gain_to_exit', 'loose_to_exit', 'returns']]\
.set_index(['gain_to_exit', 'loose_to_exit'])\
.unstack('loose_to_exit')
backtest_results_df.columns = backtest_results_df.columns.droplevel()
plt.rcParams['figure.figsize'] = (14.0, 12.0)
plt.pcolor(backtest_results_df, cmap="seismic", edgecolors='k', linewidths=0.3)
plt.yticks(np.arange(0.5, len(backtest_results_df.index), 1), backtest_results_df.index)
plt.xticks(np.arange(0.5, len(backtest_results_df.columns), 1), backtest_results_df.columns, rotation='vertical')
plt.colorbar();
# +
# Overall results
fig, ax1 = plt.subplots(figsize=(10,6))
ax1.plot(
[round(i,2) for i in np.arange(0,1.01,0.05)],
[pd.DataFrame(backtest_results)['returns'].quantile(i) for i in np.arange(0,1.01,0.05)],
label='strategy perfomance dist'
)
ax1.hlines(bt.get_return_benchmark(), 0, 1, label='hodling')
plt.grid()
plt.legend();
# +
# data_simplestr['trades'] = create_strategy(data_simplestr, 'priceUsd', 'signal', 0.81, 0.01)
# bt = Backtest(data_simplestr.returns.dropna(), data_simplestr.trades)
# bt.summary()
# bt.plot_backtest(viz='hodl')
# +
# Play yourself :)
# df: high_price # Strategy with hourly high price and naive signals detection
# df: data_simplestr # Strategy with hourly average price and naive signals detection
# df: data_predicted # Strategy with hourly average price and predicted by prophet anomalies
# -
# ## <b> TLDR and open questions
# - Its possible to foresee upcoming pumps using centralized exchanges activity
#
# - Naive way of detecting anomalies works better than prophet predicted anomalies (TODO: experiment with confidence intervals).
#
# - It's important not only to foresee pumps but only to find right "exit" moment. (TODO: experiment with finding tops using other metrics (like soical volume, active addresses, etc)).
#
# - Than more exchange addresses is in the sample the better strategies work. (DISCUSSION: identifying CEX wallets onchain).
#
# - DISCUSSION: with deposit addresses it's possible to tract different wallets of the user. Any connections between pump-makers?
# Questions? Ideas?
# Find me here:
# <EMAIL>
# (WhatsApp, Telegram): +375256830154
# or join us on **santiment.net** or [our discord channel](https://discord.gg/q3m9vUg)!
| ICDM_2019/ICDM_no_outputs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# ## PHYS 105A: Introduction to Scientific Computing
#
# # The C programming language
#
# <NAME>
# + slideshow={"slide_type": "slide"}
# %%time
# What we did last time... how long does this piece of code take to run?
import random as rnd
from matplotlib import pyplot as plt
def randomwalk(n_steps=1000):
X = [0] # initial position
for t in range(n_steps):
last = X[-1] # last position
r = rnd.randint(0,1) # we generate 0 or 1 randomly
if r == 0: # depending on r, we step left or right
curr = last + 1
else:
curr = last - 1
X.append(curr) # append the current position to the list X
return X # return the result
n_trials = 100
T = range(1, 1000+1)
D = []
for t in T: # M = 0; M += m ...; M /= N => mean of m
M = 0
for trial in range(n_trials):
X = randomwalk(t)
M += abs(X[-1])
M /= n_trials
D.append(M)
# + slideshow={"slide_type": "slide"}
plt.loglog(T, D)
plt.plot(T, [t**0.5 for t in T]) # y = sqrt(x)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Performance
#
# * When we sovled the random walk and Brownian motion problems last time, or compute $\pi$ and try to study its convergnece rate, our computers didn't give us the answer immediate.
#
# * This was a pretty simple problem... why did it take so long?
#
# * One of the reasons is that we use python.
#
# * Python, being an interpreted lagnauge, is slow in nature.
#
# * Because one of the goals of this course is to see many many tools---so try to implement the same problem in a compiled langauge!
# + [markdown] slideshow={"slide_type": "slide"}
# ## Which Language to Choose?
#
# * In the second lecture, we had a quick introduction to the Unix/Linux operating systems. And we saw these two gentlemen: <NAME> and <NAME>.
#
# * In those early day, one thing that made Unix stood out from its competitors was that it was developed in a "high level programming language" C. <NAME> is actually the inventor of C.
#
# * The popularity and powerfulness of Unix and C go hand-in-hand.
#
# * Even C was invented in 1972 (49 years ago!), it is still the [*MOST POPULAR* programming language](https://www.tiobe.com/tiobe-index/) today!
#
# * It's very inferential and its syntax shows up in many modern laguages such as C++, Go, Rust, etc.
#
# 
# + [markdown] slideshow={"slide_type": "slide"}
# # The C Programming Language
#
# 
# + [markdown] slideshow={"slide_type": "slide"}
# ## Compiler
#
# * First thing to know, C is a compiled language. You need a *compiler* to use it.
#
# * A copmiler is a program that translate your program from human readable source codes into machine readable executables.
#
# * On Unix/Linux/Mac, the most popular choices are `gcc` and `clang`.
#
# * On Windows, you may use:
# * C/C++ for Visual Studio Code: https://code.visualstudio.com/docs/languages/cpp
# * Visual Studio Community Edition: https://visualstudio.microsoft.com/downloads/
# * Tiny C Compiler: https://bellard.org/tcc/
# * GCC: https://gcc.gnu.org/install/binaries.html
# + [markdown] slideshow={"slide_type": "slide"}
# ## Hello World in C
#
# * The following is the text inside a plan text file with ".c" extension, e.g., "hw.c"
#
# #include <stdio.h> /* I/O is not built-in to the language; include standard library */
# int main() /* a program always uses `main` as an entry point */
# {
# printf("Hello World!\n"); /* use "format-print" for output */
# return 0; /* the standard is to return 0 from main if no error */
# }
#
# * Running "hw.c" does not work. You need to first compile it:
#
# gcc hw.c -o hw.exe
#
# the above command would compile the "hw.c" program into the executable "hw.exe". The ".exe" extension is optional on Unix/Linux.
#
# * Now you can run the program by:
#
# ./hw.exe
#
# which outputs "Hello World!"
# + [markdown] slideshow={"slide_type": "slide"}
# ## Python vs C
#
# Python | C
# :-- | :--
# `import package` | `#include <package.h>` and then link with `gcc ... -lpackage`
# Dynamic type | `int`, `double`, `char []`
# `def func(): ...` | `double func(double x) {...}`
# Must use indentation | Indentation optional; use `{...}` for block
# New line for new statement | `;` to end a statement
# `print(x)` | `printf("%d\n", x);`
# `return x` | `return x;`
# `if x == 1: ... else: ...` | `if (x == 1) {...} else {...}`
# `for i in range(10): ...` | `for (int i = 0; i < 10; ++i) {...}`
# `/` vs `//` for divisions | `/` behavior depends on type
# `**` for power | `#include <math.h>` and then use `pow(x, y` for power; need to link with `gcc ... -lm`
# `# ...` for comments | `/* ... */` or `//...` for comments
# + slideshow={"slide_type": "slide"} active=""
# Python | C
# ------------------------------------ | -------------------------------------------
# import random as rnd | #include <stdlib.h>
# from matplotlib import pyplot as plt | #include <stdio.h>
# |
# n_trials = 100 | int n_trials = 100;
# |
# def randomwalk(n_steps=1000): | int randomwalk(int n_steps) {
# X = [0] # initial position | int X = 0;
# |
# for t in range(n_steps): | for (int t = 0; t < n_steps; ++t) {
# last = X[-1] | int last = X;
# | int curr;
# r = rnd.randint(0,1) | int r = rand();
# if r == 0: | if (r < RAND_MAX/2)
# curr = last + 1 | curr = last + 1;
# else: | else
# curr = last - 1 | curr = last - 1;
# X.append(curr) | X = curr;
# | }
# return X | return X;
# | }
# |
# T = range(1, 1000+1) |
# D = [] | int main() {
# for t in T: | for (int t = 1; t < 1001; ++t) {
# M = 0 | double M = 0;
# for trial in range(n_trials): | for (int trial = 0; trial < n_trials; ++trial) {
# X = randomwalk(t) | int X = randomwalk(t);
# M += abs(X[-1]) | M += abs(X);
# | }
# M /= n_trials | M /= n_trials;
# D.append(M) | printf("%d %g\n", t, M);
# | }
# | return 0;
# | }
# + [markdown] slideshow={"slide_type": "slide"}
# * If we save the C code in "rw.c", we may compile it in a terminal
# gcc rw.c -o rw.exe
#
# * We can now run it as
# ./rw.exe
#
# * This will give us a long list of numbers. We may "redirect" it to a text file. We may also time it.
# time ./rw.exe > rw.txt
#
# * We may even ask the compiler to work harder and optimize the code for us!
# gcc rw.c -O3 -o rw.exe
# + slideshow={"slide_type": "slide"}
# Let's also plot the result
from matplotlib import pyplot as plt
T, D = [], []
with open('output.txt', 'r') as f:
for line in f:
t, d = line.split()
T.append(int(t))
D.append(float(d))
plt.loglog(T, D)
plt.plot(T, [t**0.5 for t in T]) # y = sqrt(x)
# + [markdown] slideshow={"slide_type": "slide"}
# * On my laptop:
# * The python code took 50 sec to compute this.
# * The C code took 0.25 sec to compute the same thing!
#
# * The C code is 200x faster than our python code!!!
#
# * Now imagine you have a scientific simulation that takes a week to run on C. It will take four years to run if you use pure python!
#
# * Know your problem; know your tools; and then decide what to use.
#
# * We will learn how to do this fast in python next time.
| 05/C.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib notebook
import matplotlib
import seaborn as sb
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
# Jupyter Specifics
# %matplotlib inline
from IPython.display import display, HTML
from ipywidgets.widgets import interact, interactive, IntSlider, FloatSlider, Layout, ToggleButton, ToggleButtons, fixed
display(HTML("<style>.container { width:100% !important; }</style>"))
style = {'description_width': '100px'}
slider_layout = Layout(width='99%')
from time import time
import pickle as pk
# -
# # Base data setup from databases
# new module data_config imported by data.py as well as Cluster.py
import data_config
data_config.report_correct = True
from data import *
continents
# continents + Middle_East
for cc in countries_owid:
print(cc,continent[cc])
set([continent[cc] for cc in countries_owid])
countries_in_continent
# ## Data save
#
# Execute this section once to produce file `data_all_base.pk` in directory pks.
# +
# miscnms = ['clusdata_all','cases','datasets','contact_dic','age_group_dic']
miscnms = ['acute_dict','age_group_dic','base','contact_dic','final_date','gdp_per_capita_owid',
'icu_dict','owid_file','stringency_owid','tests_owid','translate_age','translate_contact']
deathnms = [x for x in dir() if 'deaths' in x]
casenms = [x for x in dir() if 'cases' in x if not callable(eval(x))]
covidnms = [x for x in dir() if 'covid' in x]
popnms = [x for x in dir() if 'population' in x]
testnms = [x for x in dir() if 'testing' in x if not callable(eval(x))]
recovnms = [x for x in dir() if 'recovered' in x]
confnms = [x for x in dir() if 'confirmed' in x]
countrynms = [x for x in dir() if 'countr' in x and not callable(eval(x))]
countrynms = [x for x in dir() if 'countr' in x and (isinstance(eval(x),dict) or isinstance(eval(x),list) or isinstance(eval(x),tuple))]
continentnms = [x for x in dir() if 'continent' in x]
allnmsdb = countrynms + covidnms + miscnms + deathnms + confnms + recovnms + casenms + popnms + testnms + ['allnmsdb']
data_all = {nm:eval(nm) for nm in allnmsdb}
start = time()
pk.dump(data_all,open('./pks/data_all_base.pk','wb'))
print('elapsed: ',time()-start)
# -
print(allnmsdb)
# the remaining list from dir() below should only contain temporary variables and system modules
[x for x in dir() if '__' not in x and not callable(eval(x)) and not x in allnmsdb and not x.startswith('_') and not x.endswith('nms')]
# ## Data Load
#
# Use this code to read in the data, e.g. at the top of another notebook, as an alternative to loading data.py
# +
# read in data
start=time()
print('reading in data...')
with open('./pks/data_all_base.pk','rb') as fp:
foo = pk.load(fp)
print('elapsed: ',time()-start)
# make each element of the dictionary a global variable named with key:
for x in foo:
stmp = x+"= foo['"+x+"']"
exec(stmp)
# -
| Notebooks/covid-19-caution/Data_base.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %pylab inline
import numpy as np
import torch
import os
from torch import nn
from torch import optim
from torch.nn import functional as F
from torch import autograd
from torch.autograd import Variable
import nibabel as nib
from torch.utils.data.dataset import Dataset
from torch.utils.data import dataloader
from nilearn import plotting
from ADNI_dataset import *
from BRATS_dataset import *
from ATLAS_dataset import *
from Model_alphaGAN import *
# # Configuration
# +
BATCH_SIZE=4
gpu = True
workers = 4
LAMBDA= 10
_eps = 1e-15
Use_BRATS = False
Use_ATLAS = False
#setting latent variable sizes
latent_dim = 1000
# +
trainset = ADNIdataset(augmentation=False)
train_loader = torch.utils.data.DataLoader(trainset,batch_size=BATCH_SIZE,
shuffle=True,num_workers=workers)
if Use_BRATS:
#imgtype -> 'flair' or 't2' or 't1ce'
trainset = BRATSdataset(train=True, imgtype = 'flair',augmentation=False)
train_loader = torch.utils.data.DataLoader(trainset,batch_size=BATCH_SIZE,
shuffle=True,num_workers=workers)
if Use_ATLAS:
trainset = ATLASdataset(augmentation=True)
train_loader = torch.utils.data.DataLoader(trainset,batch_size=BATCH_SIZE,
shuffle=True,num_workers=workers)
# -
def inf_train_gen(data_loader):
while True:
for _,images in enumerate(data_loader):
yield images
# +
G = Generator(noise = latent_dim)
CD = Code_Discriminator(code_size = latent_dim ,num_units = 4096)
D = Discriminator(is_dis=True)
E = Discriminator(out_class = latent_dim ,is_dis=False)
G.cuda()
D.cuda()
CD.cuda()
E.cuda()
# -
g_optimizer = optim.Adam(G.parameters(), lr=0.0002)
d_optimizer = optim.Adam(D.parameters(), lr=0.0002)
e_optimizer = optim.Adam(E.parameters(), lr = 0.0002)
cd_optimizer = optim.Adam(CD.parameters(), lr = 0.0002)
def calc_gradient_penalty(model, x, x_gen, w=10):
"""WGAN-GP gradient penalty"""
assert x.size()==x_gen.size(), "real and sampled sizes do not match"
alpha_size = tuple((len(x), *(1,)*(x.dim()-1)))
alpha_t = torch.cuda.FloatTensor if x.is_cuda else torch.Tensor
alpha = alpha_t(*alpha_size).uniform_()
x_hat = x.data*alpha + x_gen.data*(1-alpha)
x_hat = Variable(x_hat, requires_grad=True)
def eps_norm(x):
x = x.view(len(x), -1)
return (x*x+_eps).sum(-1).sqrt()
def bi_penalty(x):
return (x-1)**2
grad_xhat = torch.autograd.grad(model(x_hat).sum(), x_hat, create_graph=True, only_inputs=True)[0]
penalty = w*bi_penalty(eps_norm(grad_xhat)).mean()
return penalty
# # Training
# +
real_y = Variable(torch.ones((BATCH_SIZE, 1)).cuda(async=True))
fake_y = Variable(torch.zeros((BATCH_SIZE, 1)).cuda(async=True))
criterion_bce = nn.BCELoss()
criterion_l1 = nn.L1Loss()
criterion_mse = nn.MSELoss()
# -
gen_load = inf_train_gen(train_loader)
MAX_ITER = 200000
for iteration in range(MAX_ITER):
###############################################
# Train Encoder - Generator
###############################################
for p in D.parameters(): # reset requires_grad
p.requires_grad = False
for p in CD.parameters(): # reset requires_grad
p.requires_grad = False
for p in E.parameters(): # reset requires_grad
p.requires_grad = True
for p in G.parameters(): # reset requires_grad
p.requires_grad = True
g_optimizer.zero_grad()
e_optimizer.zero_grad()
for iters in range(1):
real_images = gen_load.__next__()
real_images = Variable(real_images,volatile=True).cuda(async=True)
_batch_size = real_images.size(0)
z_hat = E(real_images).view(_batch_size,-1)
z_rand = Variable(torch.randn((_batch_size,latent_dim)),requires_grad=False).cuda()
x_hat = G(z_hat)
x_rand = G(z_rand)
l1_loss = 10 * criterion_l1(x_hat, real_images)
c_loss = criterion_bce(CD(z_hat), real_y[:_batch_size])
d_real_loss = criterion_bce(D(x_hat), real_y[:_batch_size])
d_fake_loss = criterion_bce(D(x_rand), real_y[:_batch_size])
loss1 = l1_loss + c_loss + d_real_loss + d_fake_loss
loss1.backward(retain_graph=True)
e_optimizer.step()
g_optimizer.step()
g_optimizer.step()
###############################################
# Train D
###############################################
for p in D.parameters():
p.requires_grad = True
for p in CD.parameters():
p.requires_grad = False
for p in E.parameters():
p.requires_grad = False
for p in G.parameters():
p.requires_grad = False
for iters in range(1):
d_optimizer.zero_grad()
z_rand = Variable(torch.randn((_batch_size,latent_dim)),volatile=True).cuda()
z_hat = E(real_images).view(_batch_size,-1)
x_hat = G(z_hat)
x_rand = G(z_rand)
x_loss2 = 2.0 * criterion_bce(D(real_images), real_y[:_batch_size])+criterion_bce(D(x_hat), fake_y[:_batch_size])
z_loss2 = criterion_bce(D(x_rand), fake_y[:_batch_size])
loss2 = x_loss2 + z_loss2
if iters<4:
loss2.backward(retain_graph=True)
else:
loss2.backward(retain_graph=True)
d_optimizer.step()
###############################################
# Train CD
###############################################
for p in D.parameters(): # reset requires_grad
p.requires_grad = False
for p in CD.parameters(): # reset requires_grad
p.requires_grad = True
for p in E.parameters(): # reset requires_grad
p.requires_grad = False
for p in G.parameters(): # reset requires_grad
p.requires_grad = False
for iters in range(1):
cd_optimizer.zero_grad()
z_hat = E(real_images).view(_batch_size,-1)
x_loss3 = criterion_bce(CD(z_hat), fake_y[:_batch_size])
z_rand = Variable(torch.randn((_batch_size,latent_dim)),volatile=True).cuda()
z_loss3 = criterion_bce(CD(z_rand), real_y[:_batch_size])
loss3 = x_loss3 + z_loss3
loss3.backward(retain_graph=True)
cd_optimizer.step()
###############################################
# Visualization
###############################################
if iteration % 50 == 0:
print('[{}/{}]'.format(iteration,50000),
'D: {:<8.3}'.format(loss2.data[0].cpu().numpy()),
'En_Ge: {:<8.3}'.format(loss1.data[0].cpu().numpy()),
'Code: {:<8.3}'.format(loss3.data[0].cpu().numpy()),
)
featmask = np.squeeze((0.5*real_images[0]+0.5).data.cpu().numpy())
featmask = nib.Nifti1Image(featmask,affine = np.eye(4))
plotting.plot_img(featmask,title="Real")
plotting.show()
featmask = np.squeeze((0.5*x_hat[0]+0.5).data.cpu().numpy())
featmask = nib.Nifti1Image(featmask,affine = np.eye(4))
plotting.plot_img(featmask,title="DEC")
plotting.show()
featmask = np.squeeze((0.5*x_rand[0]+0.5).data.cpu().numpy())
featmask = nib.Nifti1Image(featmask,affine = np.eye(4))
plotting.plot_img(featmask,title="Rand")
plotting.show()
if (iteration+1)%500 ==0:
torch.save(G.state_dict(),'./checkpoint/G_noW_iter'+str(iteration+1)+'.pth')
torch.save(D.state_dict(),'./checkpoint/D_noW_iter'+str(iteration+1)+'.pth')
torch.save(E.state_dict(),'./checkpoint/E_noW_iter'+str(iteration+1)+'.pth')
torch.save(CD.state_dict(),'./checkpoint/CD_noW_iter'+str(iteration+1)+'.pth')
| Alpha_GAN_ADNI_train.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
os.chdir('..')
# +
from ssd_model import SSD300
model = SSD300((300, 300, 3), 21)
print(model.layers[1].get_weights()[0][:,:,:,5])
# -
model.load_weights('ssd300_voc_weights_fixed.hdf5', by_name=True)
print(model.layers[1].get_weights()[0][:,:,:,5])
# +
from ssd_utils import load_weights
#load_weights(model, 'ssd300_voc_weights_fixed.hdf5', ['conv1_1', 'conv1_2'])
load_weights(model, 'ssd300_voc_weights_fixed.hdf5')
print(model.layers[1].get_weights()[0][:,:,:,5])
# -
| misc/load_weights.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="miySfvLseXMj" executionInfo={"status": "ok", "timestamp": 1626916762900, "user_tz": 240, "elapsed": 396, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgimKXlswsRONzCsgtK97IzHh9zcvgsGKFppYo=s64", "userId": "10197604116353826977"}}
import pandas as pd
import numpy as np
import requests
url = 'https://raw.githubusercontent.com/jbrownlee/Datasets/master/airline-passengers.csv'
res = requests.get(url, allow_redirects=True)
with open('airline-passengers.csv','wb') as file:
file.write(res.content)
# + colab={"base_uri": "https://localhost:8080/", "height": 232} id="GL1He3xpGeuJ" executionInfo={"status": "ok", "timestamp": 1626916763274, "user_tz": 240, "elapsed": 4, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgimKXlswsRONzCsgtK97IzHh9zcvgsGKFppYo=s64", "userId": "10197604116353826977"}} outputId="8bfedf92-d662-46dd-d984-f7bbea6510f9"
df=pd.read_csv('/content/airline-passengers.csv',index_col='Month' ,parse_dates=True)
df=df.dropna()
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 352} id="HDthD4U8PfAu" executionInfo={"status": "ok", "timestamp": 1626916763714, "user_tz": 240, "elapsed": 443, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgimKXlswsRONzCsgtK97IzHh9zcvgsGKFppYo=s64", "userId": "10197604116353826977"}} outputId="2726a2c5-3da0-4e27-e391-a2a8405b8313"
df['Passengers'].plot(figsize=(15,5))
# + colab={"base_uri": "https://localhost:8080/"} id="XyXdFfNAPfl_" executionInfo={"status": "ok", "timestamp": 1626916764054, "user_tz": 240, "elapsed": 343, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgimKXlswsRONzCsgtK97IzHh9zcvgsGKFppYo=s64", "userId": "10197604116353826977"}} outputId="f0f8348d-59b3-4f2d-9498-2b38e9ea43eb"
from statsmodels.tsa.stattools import adfuller
def ad_test(dataset):
dftest = adfuller(dataset, autolag = 'AIC')
print("1. ADF : ",dftest[0])
print("2. P-Value : ", dftest[1])
print("3. Num Of Lags : ", dftest[2])
print("4. Num Of Observations Used For ADF Regression:", dftest[3])
print("5. Critical Values :")
for key, val in dftest[4].items():
print("\t",key, ": ", val)
ad_test(df['Passengers'])
# + colab={"base_uri": "https://localhost:8080/"} id="eJj-gtZMaduw" executionInfo={"status": "ok", "timestamp": 1626916767270, "user_tz": 240, "elapsed": 3217, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgimKXlswsRONzCsgtK97IzHh9zcvgsGKFppYo=s64", "userId": "10197604116353826977"}} outputId="cc1a04ad-e619-497e-9418-e4c7f7e81dc7"
# !pip install pmdarima
# + id="rxZODP64bGU6" executionInfo={"status": "ok", "timestamp": 1626916767271, "user_tz": 240, "elapsed": 3, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgimKXlswsRONzCsgtK97IzHh9zcvgsGKFppYo=s64", "userId": "10197604116353826977"}}
from pandas.util._decorators import (
deprecate_kwarg, Appender, Substitution, cache_readonly
)
# + colab={"base_uri": "https://localhost:8080/"} id="i89btLy4aRYF" executionInfo={"status": "ok", "timestamp": 1626916777799, "user_tz": 240, "elapsed": 10530, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgimKXlswsRONzCsgtK97IzHh9zcvgsGKFppYo=s64", "userId": "10197604116353826977"}} outputId="285b060d-9980-4cce-f442-ccc3de550da9"
from pmdarima import auto_arima
stepwise_fit = auto_arima(df['Passengers'], trace=True,
suppress_warnings=True)
# + colab={"base_uri": "https://localhost:8080/"} id="sBP1yzKHPlSV" executionInfo={"status": "ok", "timestamp": 1626916777799, "user_tz": 240, "elapsed": 10, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgimKXlswsRONzCsgtK97IzHh9zcvgsGKFppYo=s64", "userId": "10197604116353826977"}} outputId="eb6d21f8-6f1f-4a2d-fdfd-bffb86c683c3"
print(df.shape)
train=df.iloc[:-10]
test=df.iloc[-10:]
print(train.shape,test.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="ZB8hJRN3PnfZ" executionInfo={"status": "ok", "timestamp": 1626916779385, "user_tz": 240, "elapsed": 1595, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgimKXlswsRONzCsgtK97IzHh9zcvgsGKFppYo=s64", "userId": "10197604116353826977"}} outputId="660c8aaa-3590-4240-8fa8-969a389e0af5"
from statsmodels.tsa.arima_model import ARIMA
model=ARIMA(train['Passengers'],order=(4,1,3))
model=model.fit()
model.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 292} id="fCjhl2jSPpKa" executionInfo={"status": "ok", "timestamp": 1626916779747, "user_tz": 240, "elapsed": 366, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgimKXlswsRONzCsgtK97IzHh9zcvgsGKFppYo=s64", "userId": "10197604116353826977"}} outputId="a8ff4e95-2947-4bb3-c43a-ade67610fa35"
start=len(train)
end=len(train)+len(test)-1
pred=model.predict(start=start,end=end,typ='levels').rename('ARIMA predictions')
pred.plot(legend=True)
# + colab={"base_uri": "https://localhost:8080/"} id="Dgq6Oe5OPsmo" executionInfo={"status": "ok", "timestamp": 1626916779748, "user_tz": 240, "elapsed": 6, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgimKXlswsRONzCsgtK97IzHh9zcvgsGKFppYo=s64", "userId": "10197604116353826977"}} outputId="60e16387-27f8-411d-bfed-6bb4755ec2d8"
from sklearn.metrics import mean_squared_error
from math import sqrt
test['Passengers'].mean()
rmse=sqrt(mean_squared_error(pred,test['Passengers']))
print(rmse)
# + id="bTsY-T20XYsF" executionInfo={"status": "ok", "timestamp": 1626916779748, "user_tz": 240, "elapsed": 4, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgimKXlswsRONzCsgtK97IzHh9zcvgsGKFppYo=s64", "userId": "10197604116353826977"}}
| ARIMA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import os
import pandas as pd
import numpy as np
from numpy.random import RandomState
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.model_selection import GridSearchCV
from sklearn.pipeline import Pipeline
from sklearn import utils
from sklearn.metrics import accuracy_score
from sklearn.metrics import f1_score
from sklearn.cluster import KMeans
from sklearn.preprocessing import StandardScaler
from sklearn.cluster import SpectralClustering
from pprint import pprint
from time import time
import logging
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
from xgboost import XGBClassifier
from sklearn.linear_model import SGDClassifier
from sklearn.svm import LinearSVC
from sklearn.naive_bayes import CategoricalNB
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import SVC
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import ExtraTreesClassifier
from sklearn.ensemble import AdaBoostClassifier
from sklearn.ensemble import GradientBoostingClassifier
# Define randomState constant to make the experiments replicable
seed = RandomState(0)
utils.check_random_state(seed)
# Read training set
features = ['title', 'concatenated_tags', 'query', 'search_page', 'price', 'weight', 'seller_id', 'minimum_quantity', 'category']
products = pd.read_csv(os.environ['DATASET_PATH'], usecols=features)
# products = pd.read_csv('e:/work/intelligent-systems-project/data/sample_products.csv', usecols=features)
# Clean products null and NaN occurrences. Remove only 60 lines from 38000 in total.
products = products.dropna() # apaga apenas 60 linhas
products = products.reset_index()
# Read test set
products_test = pd.read_csv('/usr/src/data/test_products.csv', usecols=features)
# products_test = pd.read_csv('e:/work/intelligent-systems-project/data/test_products.csv', usecols=features)
# Display progress logs on stdout
logging.basicConfig(level=logging.INFO, format="%(asctime)s %(levelname)s %(message)s")
# Common functions
def evaluate_test(predicted):
target = products_test['category']
categories = ['Lembrancinhas', 'Decoração', 'Bebê', 'Papel e Cia', 'Outros', 'Bijuterias e Jóias']
accuracy = accuracy_score(target, predicted)
f1_score_micro = f1_score(products_test['category'], predicted, average='micro', labels=categories)
print(" test: %f (Accuracy: %f, F1_score(micro): %f)" % ( (accuracy+f1_score_micro)/2, accuracy, f1_score_micro) )
def concatCols(dataset, textColumnList):
SEP = " "
INPUT_COLUMNS_WITH_SEP = ",sep,".join(textColumnList).split(",")
return dataset.assign(sep=SEP)[INPUT_COLUMNS_WITH_SEP].sum(axis=1)
print("%d documents" % len(products))
print("%d categories" % len(products['category'].value_counts()))
print()
print('Setup complete!')
# -
# # 1. Initial training data verification
# - verify loaded columns data types OK
# - verify presence of nulls and NaN OK
# - verify target categories, is it balanced? Not, it is not balanced.
# ```
# Lembrancinhas 17524
# Decoração 8723
# Bebê 6930
# Papel e Cia 2750
# Outros 1133
# Bijuterias e Jóias 940
# ```
# # 2. Setting an experiment baseline
#
# GridSearchCV class provides a cross-validation method to search for hyperparameters. It will be used as base for next exploration. The experiment baseline uses a **5-fold cross-validation**, **no hyperparameters** exploration and **accuracy** and **f1_score** metrics.
#
# ### Pipeline
# Utilizes a **CountVectorizer** feature extraction steio (bag-of-words) in **'title' column**. Passing data to a **SGDClassifier**.
# Based on [Sample pipeline for text feature extraction and evaluation](https://scikit-learn.org/0.15/auto_examples/grid_search_text_feature_extraction.html)
#
# ### Scores:
# ```
# pipeline: ['vect', 'clf']
# Fitting 5 folds for each of 1 candidates, totalling 5 fits
# done in 2.861s
#
# train: 0.907940 (mean of Accuracy and F1_score(micro))
# val: 0.874750 (mean of Accuracy and F1_score(micro))
# test: 0.896000 (Accuracy: 0.896000, F1_score(micro): 0.896000)
# ```
# +
def exp1(textColumnsList):
X_train = concatCols(products, textColumnsList)
X_test = concatCols(products_test, textColumnsList)
Y_train = products['category']
scoring = {'Accuracy': 'accuracy', 'F1_score': 'f1_micro' }
pipeline = Pipeline(
[
("vect", CountVectorizer()),
("clf", SGDClassifier(random_state=0)),
]
)
parameters = {}
if __name__ == "__main__":
# multiprocessing requires the fork to happen in a __main__ protected block
# find the best parameters for both the feature extraction and the classifier
grid_search = GridSearchCV(pipeline, parameters, n_jobs=-1, verbose=4, scoring=scoring, refit="Accuracy", return_train_score=True)
print("pipeline:", [name for name, _ in pipeline.steps])
t0 = time()
grid_search.fit(X_train, Y_train)
print("done in %0.3fs" % (time() - t0))
print()
print("train: %f (mean of Accuracy and F1_score(micro))" % grid_search.cv_results_['mean_train_Accuracy']) # mean of Accuracy and F1_score(micro)
print(" val: %f (mean of Accuracy and F1_score(micro))" % grid_search.cv_results_['mean_test_Accuracy']) # mean of Accuracy and F1_score(micro)
predicted = grid_search.best_estimator_.predict(X_test)
evaluate_test(predicted)
exp1(['title'])
# -
# # 3. Getting a feeling about the other text columns
#
# May be there is a better text column than 'title' to use.
# May be I could combine all of them.
# The objetive of this experiement is to get a feeling if the others text columns could be useful.
#
# ### Results:
# Using a combination of columns **title**, **concatenated_tags** and **query** seems to contribute to improve results.
# ```
# <| ['title', 'concatenated_tags', 'query'] |>
# pipeline: ['vect', 'clf']
# Fitting 5 folds for each of 1 candidates, totalling 5 fits
# done in 5.334s
#
# train: 0.937078 (mean of Accuracy and F1_score(micro))
# val: 0.896310 (mean of Accuracy and F1_score(micro))
# test: 0.930000 (Accuracy: 0.930000, F1_score(micro): 0.930000)
#
#
# Crossvalidation: from 0.874750 to 0.896310.
# Test: from 0.896000 to 0.930000.
# Test accuracy: from 0.896000 to 0.930000.
# Test F1_score(micro): from 0.896000 to 0.930000
# ```
#
#
# +
combinationsToTest = [
['title'],
['concatenated_tags'],
['query'],
['title', 'concatenated_tags'],
['title', 'query'],
['concatenated_tags', 'query'],
['title', 'concatenated_tags', 'query']
]
for k in combinationsToTest:
print('<|',k,'|>')
exp1(k)
print('-----------------------------------------------------------------\n')
# -
# # 4. Exploring some parameters for CountVectorizer
#
# I think the combined text column from previous experiment (title, tags, query) problably will result in some duplicated words. I order to avoid misinterpreting duplicate words as more valuable words I will try adding parameter binary: True . When enabled, if True, all non zero counts are set to 1. When enabling it, I expect getting the same score in baseline and better score when using combined text column.
#
# I will use this experiment to explore other configurations of this module. Some parameters ideas came from [Sample pipeline for text feature extraction and evaluation](https://scikit-learn.org/0.15/auto_examples/grid_search_text_feature_extraction.html).
#
# ```
# strip_accents{None, ‘unicode’}
# stop_words{None, stop_portugues, stop_portugues_small }
# binary{False, True}
# max_features
# max_df: (0.5, 0.75, 1.0),
# max_features: (None, 5000, 10000, 50000),
# ngram_range: ((1, 1), (1, 2)), # unigrams or bigrams
# ```
#
# ### Results:
# The varatiation is small, but it seems that we should use binary: True
#
# ```
# Performing grid search...
# pipeline: ['vect', 'clf']
# parameters:
# {'vect__binary': (False, True),
# 'vect__max_df': (0.5, 0.75, 1.0),
# 'vect__max_features': (None, 5000, 10000, 50000),
# 'vect__ngram_range': ((...), (...)),
# 'vect__stop_words': (None, [...], [...], [...]),
# 'vect__strip_accents': (None, 'unicode')}
# Fitting 5 folds for each of 384 candidates, totalling 1920 fits
# done in 2147.774s
#
# Best parameters set:
# vect__binary: True
# vect__max_df: 0.5
# vect__max_features: None
# vect__ngram_range: (1, 2)
# vect__stop_words: ['de', 'do', 'dos', 'com', 'em', 'o', 'e', 'para', 'em']
# vect__strip_accents: None
# val: 0.924776 (mean of Accuracy and F1_score(micro))
# test: 0.956000 (Accuracy: 0.956000, F1_score(micro): 0.956000)
#
#
# train: 0.937078 (mean of Accuracy and F1_score(micro))
# val: 0.896310 (mean of Accuracy and F1_score(micro))
# test: 0.930000 (Accuracy: 0.930000, F1_score(micro): 0.930000)
#
# Crossvalidation: from 0.896310 to 0.924776.
# Test: from 0.930000 to 0.956000.
# Test accuracy: from 0.930000 to 0.956000.
# Test F1_score(micro): 0.930000 to 0.956000
# ```
#
#
# +
def exp2(textColumnsList):
X_train = concatCols(products, textColumnsList)
X_test = concatCols(products_test, textColumnsList)
Y_train = products['category']
scoring = {'Accuracy': 'accuracy', 'F1_score': 'f1_micro' }
pipeline = Pipeline(
[
("vect", CountVectorizer()),
("clf", SGDClassifier(random_state=0)),
]
)
stop_portugues_fromInternet = ['de', 'a', 'o', 'que', 'e', 'do', 'da', 'em', 'um', 'para', 'e', 'com', 'nao', 'uma', 'os', 'no', 'se', 'na', 'por', 'mais', 'as', 'dos', 'como', 'mas', 'foi', 'ao', 'ele', 'das', 'tem', 'a', 'seu', 'sua', 'ou', 'ser', 'quando', 'muito', 'ha', 'nos', 'ja', 'esta', 'eu', 'tambem', 'so', 'pelo', 'pela', 'ate', 'isso', 'ela', 'entre', 'era', 'depois', 'sem', 'mesmo', 'aos', 'ter', 'seus', 'quem', 'nas', 'me', 'esse', 'eles', 'estao', 'voce', 'tinha', 'foram', 'essa', 'num', 'nem', 'suas', 'meu', 'as', 'minha', 'tem', 'numa', 'pelos', 'elas', 'havia', 'seja', 'qual', 'sera', 'nos', 'tenho', 'lhe', 'deles', 'essas', 'esses', 'pelas', 'este', 'fosse', 'dele', 'tu', 'te', 'voces', 'vos', 'lhes', 'meus', 'minhas', 'teu', 'tua', 'teus', 'tuas', 'nosso', 'nossa', 'nossos', 'nossas', 'dela', 'delas', 'esta', 'estes', 'estas', 'aquele', 'aquela', 'aqueles', 'aquelas', 'isto', 'aquilo', 'estou', 'esta', 'estamos', 'estao', 'estive', 'esteve', 'estivemos', 'estiveram', 'estava', 'estavamos', 'estavam', 'estivera', 'estiveramos', 'esteja', 'estejamos', 'estejam', 'estivesse', 'estivessemos', 'estivessem', 'estiver', 'estivermos', 'estiverem', 'hei', 'ha', 'havemos', 'hao', 'houve', 'houvemos', 'houveram', 'houvera', 'houveramos', 'haja', 'hajamos', 'hajam', 'houvesse', 'houvessemos', 'houvessem', 'houver', 'houvermos', 'houverem', 'houverei', 'houvera', 'houveremos', 'houverao', 'houveria', 'houveriamos', 'houveriam', 'sou', 'somos', 'sao', 'era', 'eramos', 'eram', 'fui', 'foi', 'fomos', 'foram', 'fora', 'foramos', 'seja', 'sejamos', 'sejam', 'fosse', 'fossemos', 'fossem', 'for', 'formos', 'forem', 'serei', 'sera', 'seremos', 'serao', 'seria', 'seriamos', 'seriam', 'tenho', 'tem', 'temos', 'tem', 'tinha', 'tinhamos', 'tinham', 'tive', 'teve', 'tivemos', 'tiveram', 'tivera', 'tiveramos', 'tenha', 'tenhamos', 'tenham', 'tivesse', 'tivessemos', 'tivessem', 'tiver', 'tivermos', 'tiverem', 'terei', 'tera', 'teremos', 'terao', 'teria', 'teriamos', 'teriam']
stop_portguese_fromElo7 = ['cod', 'mod', 'de', 'a', 'o', 'que', 'e', 'do', 'da', 'em', 'um', 'para', 'com', 'uma', 'os', 'no', 'se', 'na', 'por', 'mais', 'as', 'dos', 'como', 'mas', 'ao', 'ele', 'das', 'a', 'seu', 'sua', 'ou', 'quando', 'muito', 'nos', 'ja', 'eu', 'tambem', 'so', 'pelo', 'pela', 'ate', 'isso', 'ela', 'entre', 'depois', 'sem', 'mesmo', 'aos', 'seus', 'quem', 'nas', 'me', 'esse', 'eles', 'voce', 'essa', 'num', 'suas', 'meu', 'as', 'minha', 'numa', 'pelos', 'elas', 'qual', 'nos', 'lhe', 'deles', 'essas', 'esses', 'pelas', 'este', 'dele', 'tu', 'vos', 'lhes', 'meus', 'minhas', 'teu', 'tua', 'teus', 'tuas', 'nosso', 'nossa', 'nossos', 'nossas', 'dela', 'delas', 'esta', 'estes', 'estas', 'aquele', 'aquela', 'aqueles', 'aquelas', 'isto', 'aquilo', 'estou', 'esta', 'estamos', 'estao', 'estive', 'esteve', 'estivemos', 'estiveram', 'estava', 'estavamos', 'estavam', 'estivera', 'estiveramos', 'esteja', 'estejamos', 'estejam', 'estivesse', 'estivessemos', 'estivessem', 'estiver', 'estivermos', 'estiverem', 'hei', 'ha', 'havemos', 'hao', 'houve', 'houvemos', 'houveram', 'houvera', 'houveramos', 'haja', 'hajamos', 'hajam', 'houvesse', 'houvessemos', 'houvessem', 'houver', 'houvermos', 'houverem', 'houverei', 'houvera', 'houveremos', 'houverao', 'houveria', 'houveriamos', 'houveriam', 'sou', 'somos', 'sao', 'era', 'eramos', 'eram', 'fui', 'foi', 'fomos', 'foram', 'fora', 'foramos', 'seja', 'sejamos', 'sejam', 'fosse', 'fossemos', 'fossem', 'for', 'formos', 'forem', 'serei', 'sera', 'seremos', 'serao', 'seria', 'seriamos', 'seriam', 'tenho', 'tem', 'temos', 'tem', 'tinha', 'tinhamos', 'tinham', 'tive', 'teve', 'tivemos', 'tiveram', 'tivera', 'tiveramos', 'tenha', 'tenhamos', 'tenham', 'tivesse', 'tivessemos', 'tivessem', 'tiver', 'tivermos', 'tiverem', 'terei', 'tera', 'teremos', 'terao', 'teria', 'teriamos', 'teriam', 'te', 'voces']
stop_portuquese_fromWordclouds = ['de', 'do', 'dos', 'com', 'em', 'o', 'e', 'para', 'em']
parameters = {
'vect__max_df': (0.5, 0.75, 1.0),
'vect__max_features': (None, 5000, 10000, 50000),
'vect__ngram_range': ((1, 1), (1, 2)), # unigrams or bigrams
'vect__strip_accents': (None, 'unicode'),
'vect__binary': (False, True),
'vect__stop_words': (None, stop_portugues_fromInternet, stop_portguese_fromElo7, stop_portuquese_fromWordclouds),
}
if __name__ == "__main__":
# multiprocessing requires the fork to happen in a __main__ protected block
# find the best parameters for both the feature extraction and the classifier
grid_search = GridSearchCV(pipeline, parameters, n_jobs=-1, verbose=1, scoring=scoring, refit="Accuracy", return_train_score=True)
print("Performing grid search...")
print("pipeline:", [name for name, _ in pipeline.steps])
print("parameters:")
pprint(parameters, depth=2, compact=True)
t0 = time()
grid_search.fit(X_train, Y_train)
print("done in %0.3fs" % (time() - t0))
print()
print("Best parameters set:")
best_parameters = grid_search.best_estimator_.get_params()
for param_name in sorted(parameters.keys()):
print("\t%s: %r" % (param_name, best_parameters[param_name]))
print(" val: %f (mean of Accuracy and F1_score(micro))" % grid_search.best_score_)
predicted = grid_search.best_estimator_.predict(X_test)
evaluate_test(predicted)
exp2(['title', 'concatenated_tags', 'query'])
# -
# # 5. Exploring word clouds
#
# I would like to take look in text columns.
# May be it can give me a hint for a better [stop_word](https://en.wikipedia.org/wiki/Stop_word) list.
#
# ### Results:
# Title and query columns we could see clearly some stop_words: de, do, dos, com, em, o, e, para, em
# On the other hand, in **concatenated_tags** I cannot see any stop_words.
#
# ```
# stop_portuquese_fromWordclouds = ['de', 'do', 'dos', 'com', 'em', 'o', 'e', 'para', 'em']
# ```
#
# After comming back to previous experiment to test the found stop_word list, this simpler stop_words was selected.
#
# +
import random
from wordcloud import WordCloud, ImageColorGenerator
def grey_color_func(word, font_size, position, orientation, random_state=None, **kwargs):
return "hsl(0, 0%%, %d%%)" % random.randint(10, 50)
def generate_cloud(wordcloud, lines, title):
wordcloud.generate(''.join(lines))
wordcloud.recolor(color_func=grey_color_func)
plt.title(title, fontdict={'fontsize': 16})
plt.imshow(wordcloud)
plt.axis("off")
plt.show()
# Raw word cloud (with stop words)
raw_wordcloud = WordCloud(random_state=42, width=720, height=368, max_words=8000, background_color='white', stopwords='')
# Cleant word cloud (without stop words)
clean_wordcloud = WordCloud(random_state=42, width=720, height=368, max_words=8000, background_color='white')
# All documents (with/without stop words)
data = products['title'].tolist()
generate_cloud(raw_wordcloud, data, "title - most relevant words, with stop words")
generate_cloud(clean_wordcloud, data, "title - most relevant words, without stop words")
data = products['concatenated_tags'].tolist()
generate_cloud(raw_wordcloud, data, "concatenated_tags - most relevant words, with stop words")
generate_cloud(clean_wordcloud, data, "concatenated_tags - most relevant words, without stop words")
data = products['query'].tolist()
generate_cloud(raw_wordcloud, data, "query - most relevant words, with stop words")
generate_cloud(clean_wordcloud, data, "query - most relevant words, without stop words")
# -
# # 6. Trying a term-frenquency to improve feature extraction
#
# The sample [pipeline](https://scikit-learn.org/stable/auto_examples/model_selection/grid_search_text_feature_extraction.html) that I am using metions a TfidfTransformer.
# Tf means term-frequency while tf-idf means term-frequency times inverse document-frequency.
# Even not thinking that word relative frequency can improve the results, as it do not need much time, I think it worth testing.
#
# ### Results:
# Adding TfidfTransformer() did not improve the pipeline.
# +
def exp3(textColumnsList):
X_train = concatCols(products, textColumnsList)
X_test = concatCols(products_test, textColumnsList)
Y_train = products['category']
scoring = {'Accuracy': 'accuracy', 'F1_score': 'f1_micro' }
stop_portugues_fromInternet = ['de', 'a', 'o', 'que', 'e', 'do', 'da', 'em', 'um', 'para', 'e', 'com', 'nao', 'uma', 'os', 'no', 'se', 'na', 'por', 'mais', 'as', 'dos', 'como', 'mas', 'foi', 'ao', 'ele', 'das', 'tem', 'a', 'seu', 'sua', 'ou', 'ser', 'quando', 'muito', 'ha', 'nos', 'ja', 'esta', 'eu', 'tambem', 'so', 'pelo', 'pela', 'ate', 'isso', 'ela', 'entre', 'era', 'depois', 'sem', 'mesmo', 'aos', 'ter', 'seus', 'quem', 'nas', 'me', 'esse', 'eles', 'estao', 'voce', 'tinha', 'foram', 'essa', 'num', 'nem', 'suas', 'meu', 'as', 'minha', 'tem', 'numa', 'pelos', 'elas', 'havia', 'seja', 'qual', 'sera', 'nos', 'tenho', 'lhe', 'deles', 'essas', 'esses', 'pelas', 'este', 'fosse', 'dele', 'tu', 'te', 'voces', 'vos', 'lhes', 'meus', 'minhas', 'teu', 'tua', 'teus', 'tuas', 'nosso', 'nossa', 'nossos', 'nossas', 'dela', 'delas', 'esta', 'estes', 'estas', 'aquele', 'aquela', 'aqueles', 'aquelas', 'isto', 'aquilo', 'estou', 'esta', 'estamos', 'estao', 'estive', 'esteve', 'estivemos', 'estiveram', 'estava', 'estavamos', 'estavam', 'estivera', 'estiveramos', 'esteja', 'estejamos', 'estejam', 'estivesse', 'estivessemos', 'estivessem', 'estiver', 'estivermos', 'estiverem', 'hei', 'ha', 'havemos', 'hao', 'houve', 'houvemos', 'houveram', 'houvera', 'houveramos', 'haja', 'hajamos', 'hajam', 'houvesse', 'houvessemos', 'houvessem', 'houver', 'houvermos', 'houverem', 'houverei', 'houvera', 'houveremos', 'houverao', 'houveria', 'houveriamos', 'houveriam', 'sou', 'somos', 'sao', 'era', 'eramos', 'eram', 'fui', 'foi', 'fomos', 'foram', 'fora', 'foramos', 'seja', 'sejamos', 'sejam', 'fosse', 'fossemos', 'fossem', 'for', 'formos', 'forem', 'serei', 'sera', 'seremos', 'serao', 'seria', 'seriamos', 'seriam', 'tenho', 'tem', 'temos', 'tem', 'tinha', 'tinhamos', 'tinham', 'tive', 'teve', 'tivemos', 'tiveram', 'tivera', 'tiveramos', 'tenha', 'tenhamos', 'tenham', 'tivesse', 'tivessemos', 'tivessem', 'tiver', 'tivermos', 'tiverem', 'terei', 'tera', 'teremos', 'terao', 'teria', 'teriamos', 'teriam']
pipeline = Pipeline(
[
("vect", CountVectorizer(binary=True, max_df=0.5, max_features=None, ngram_range=(1, 2), strip_accents=None, stop_words=stop_portugues_fromInternet )),
('tfidf', TfidfTransformer()),
("clf", SGDClassifier(random_state=0)),
]
)
parameters = {
'tfidf__use_idf': (True, False),
'tfidf__norm': ('l1', 'l2'),
}
if __name__ == "__main__":
# multiprocessing requires the fork to happen in a __main__ protected block
# find the best parameters for both the feature extraction and the classifier
grid_search = GridSearchCV(pipeline, parameters, n_jobs=-1, verbose=1, scoring=scoring, refit="Accuracy", return_train_score=True)
print("Performing grid search...")
print("pipeline:", [name for name, _ in pipeline.steps])
print("parameters:")
pprint(parameters, depth=2, compact=True)
t0 = time()
grid_search.fit(X_train, Y_train)
print("done in %0.3fs" % (time() - t0))
print()
print("Best parameters set:")
best_parameters = grid_search.best_estimator_.get_params()
for param_name in sorted(parameters.keys()):
print("\t%s: %r" % (param_name, best_parameters[param_name]))
print(" val: %f (mean of Accuracy and F1_score(micro))" % grid_search.best_score_)
predicted = grid_search.best_estimator_.predict(X_test)
evaluate_test(predicted)
exp3(['title', 'concatenated_tags', 'query'])
# -
# # 7. Another strategy for using 'query' column
#
# The **query** column is a open field that brings text inserted by users to find the product.
# There is a auxiliary columns called **search_page** that represents the page number where the product appeared.
# I wondering if queries that find the product in first pages are better to feed the classifier.
# This experiment will filter training dataset by search_page and follow any impact in performance.
#
# ### Results
# I tried two strategies.
# 1. Add rows only if search_page <= limit. Throw out some rows.
# 2. Use all rows, but combine in text columns only if search_page <= limit
#
# The results do not show any clear improvement.
#
# +
def exp4(textColumnsList, products, products_test):
X_train = concatCols(products, textColumnsList)
X_test = concatCols(products_test, textColumnsList)
Y_train = products['category']
scoring = {'Accuracy': 'accuracy', 'F1_score': 'f1_micro' }
stop_portuquese_fromWordclouds = ['de', 'do', 'dos', 'com', 'em', 'o', 'e', 'para', 'em']
pipeline = Pipeline(
[
("vect", CountVectorizer(binary=True, max_df=0.5, max_features=None, ngram_range=(1, 2), strip_accents=None, stop_words=stop_portuquese_fromWordclouds )),
("clf", SGDClassifier(random_state=0)),
]
)
parameters = {
}
if __name__ == "__main__":
# multiprocessing requires the fork to happen in a __main__ protected block
# find the best parameters for both the feature extraction and the classifier
grid_search = GridSearchCV(pipeline, parameters, n_jobs=-1, verbose=1, scoring=scoring, refit="Accuracy", return_train_score=True)
print("Performing grid search...")
print("pipeline:", [name for name, _ in pipeline.steps])
print("parameters:")
pprint(parameters, depth=2, compact=True)
t0 = time()
grid_search.fit(X_train, Y_train)
print("done in %0.3fs" % (time() - t0))
print()
print("Best parameters set:")
best_parameters = grid_search.best_estimator_.get_params()
for param_name in sorted(parameters.keys()):
print("\t%s: %r" % (param_name, best_parameters[param_name]))
print(" val: %f (mean of Accuracy and F1_score(micro))" % grid_search.best_score_)
predicted = grid_search.best_estimator_.predict(X_test)
evaluate_test(predicted)
# 1. Add rows only if search_page <= limit. Throw out some rows.
# search_pageLimitToEvaluate = range(1,6)
# for limit in search_pageLimitToEvaluate:
# print(limit)
# products_train_exp4 = products[products['search_page'] <= limit]
# products_test_exp4 = products_test
# print('search_page <=', str(limit))
# exp4(['title', 'concatenated_tags', 'query'], products_train_exp4, products_test_exp4)
# 2. Use all rows, but combine in text columns only if search_page <= limit
# def create_custom_text_exp4 (row, limit):
# if(row['search_page'] <= limit):
# return row['title'] + ' ' + row['concatenated_tags'] + ' ' + row['query']
# else:
# return row['title'] + ' ' + row['concatenated_tags']
# search_pageLimitToEvaluate = range(1,6)
# for limit in search_pageLimitToEvaluate:
# print(limit)
# products_train_exp4 = products
# products_test_exp4 = products_test
# products_train_exp4['text'] = products_train_exp4.apply (lambda row: create_custom_text_exp4(row, limit), axis=1)
# products_test_exp4['text'] = products_test_exp4.apply (lambda row: create_custom_text_exp4(row, limit), axis=1)
# exp4(['text'], products_train_exp4, products_test_exp4)
# -
# # 8. Balancing dataset
#
# The dataset is clearly unbalanced. Even the current F1_score being high (0.956). May be it is possible to improve the classifier performance.
#
# On the other hand, the test set appears to have the same unbalanced distribution of categories. Perhaps the teacher did this to make the work easier.
#
#
# ### Results:
# - 1. Random-under sampling degraded overall performance. Seems to be an under-fitting. Score crossvaldation and test are small.
# - 2. Random over-sampling. Seems to be an over-fitting. Score crossvaldation >> Score test.
# - 3. Random under-sampling of 'Lembrancinhas' to 'Decoracao' count. Random over-sampling others categories to 'Decoracao' count. Seems to be an over-fitting. Score crossvaldation >> Score test.
#
# No improvement in overall performance.
# Curent train and test distribuition
plt.subplot(1, 2, 1)
products['category'].value_counts().plot(kind='bar', title='Train (target)');
plt.subplot(1, 2, 2)
products_test['category'].value_counts().plot(kind='bar', title='Test (target)');
# +
# Class count
count_class_lembrancinhas, count_class_decoracao, count_class_bebe, count_class_papelecia, count_class_outros, count_class_bijuterias = products['category'].value_counts()
# Divide by class
df_class_lembrancinhas = products[products['category'] == 'Lembrancinhas']
df_class_decoracao = products[products['category'] == 'Decoração']
df_class_bebe = products[products['category'] == 'Bebê']
df_class_papelecia = products[products['category'] == 'Papel e Cia']
df_class_outros = products[products['category'] == 'Outros']
df_class_bijuterias = products[products['category'] == 'Bijuterias e Jóias']
# 1. Random under-sampling
df_class_lembrancinhas_under = df_class_lembrancinhas.sample(count_class_bijuterias)
df_class_decoracao_under = df_class_decoracao.sample(count_class_bijuterias)
df_class_bebe_under = df_class_bebe.sample(count_class_bijuterias)
df_class_papelecia_under = df_class_papelecia.sample(count_class_bijuterias)
df_class_outros_under = df_class_outros.sample(count_class_bijuterias)
products_train_under = pd.concat([df_class_lembrancinhas_under, df_class_decoracao_under, df_class_bebe_under, df_class_papelecia_under, df_class_outros_under, df_class_bijuterias], axis=0)
# sns.displot(products_under, x="category").set(title='Random under-sampling')
plt.subplot(1, 3, 1)
products_train_under['category'].value_counts().plot(kind='bar', title='Under-sampling (target)');
# Running experiment
print('# 1. Random under-sampling')
exp4(['title', 'concatenated_tags', 'query'], products_train_under, products_test)
# 2. Random over-sampling
df_class_decoracao_over = df_class_decoracao.sample(count_class_lembrancinhas, replace=True)
df_class_bebe_over = df_class_bebe.sample(count_class_lembrancinhas, replace=True)
df_class_papelecia_over = df_class_papelecia.sample(count_class_lembrancinhas, replace=True)
df_class_outros_over = df_class_outros.sample(count_class_lembrancinhas, replace=True)
df_class_bijuterias_over = df_class_bijuterias.sample(count_class_lembrancinhas, replace=True)
products_train_over = pd.concat([df_class_lembrancinhas, df_class_decoracao_over, df_class_bebe_over, df_class_papelecia_over, df_class_outros_over, df_class_bijuterias_over], axis=0)
products_train_over.reset_index()
plt.subplot(1, 3, 2)
products_train_over['category'].value_counts().plot(kind='bar', title='Over-sampling (target)');
print('# 2. Random over-sampling')
exp4(['title', 'concatenated_tags', 'query'], products_train_over, products_test)
# 3. Random under-sampling of 'Lembrancinhas' to 'Decoracao' count. andom over-sampling other other categories to 'Decoracao' count.
df_class_lembrancinhas_under = df_class_lembrancinhas.sample(count_class_decoracao)
df_class_bebe_over = df_class_bebe.sample(count_class_decoracao, replace=True)
df_class_papelecia_over = df_class_papelecia.sample(count_class_decoracao, replace=True)
df_class_outros_over = df_class_outros.sample(count_class_decoracao, replace=True)
df_class_bijuterias_over = df_class_bijuterias.sample(count_class_decoracao, replace=True)
products_train_under_over = pd.concat([df_class_lembrancinhas_under, df_class_decoracao, df_class_bebe_over, df_class_papelecia_over, df_class_outros_over, df_class_bijuterias_over], axis=0)
products_train_under_over.reset_index()
plt.subplot(1, 3, 3)
products_train_under_over['category'].value_counts().plot(kind='bar', title='Over-sampling (target)');
print("'# 3. Random under-sampling of 'Lembrancinhas' to 'Decoracao' count. andom over-sampling other other categories to 'Decoracao' count.")
exp4(['title', 'concatenated_tags', 'query'], products_train_under_over, products_test)
# -
# # 9. Trying to creating text features from numeric columns (manual intervals)
#
# The price, weight and minimum quantity columns seem to provide useful information. As the classifier is receiving the bag-of-words directly, I'm not sure how to integrate them into the model.
#
# I will try to create text columns with distinct words to mark ranges of values from these columns.
# For example: priceLow, priceMedium and priceHigh to indicate value ranges for the price column.
#
# Tried setting ranges manually based on the average values for each of the output categories.
#
# ### Results:
# ```
# val: 0.926990 (mean of Accuracy and F1_score(micro))
# test: 0.952000 (Accuracy: 0.952000, F1_score(micro): 0.952000)
#
# crossvalidation: from 0.924776 to 0.926990.
# test: from 0.956000 to 0.952000 (small degradation)
# test accuracy: from 0.956000 to 0.952000 (degradation)
# test F1_score(micro): (0.956000) to 0.952000 (degradation)
# ```
# **No improvement.**
#
# +
#1. Creating features.
def label_price (row):
if row['price'] <= 30:
return 'priceLow'
elif row['price'] > 30 and row['price'] < 100:
return 'priceMedium'
else:
return 'priceHigh'
def label_weight (row):
if row['weight'] <= 100:
return 'weightLow'
elif row['weight'] > 100 and row['weight'] < 300:
return 'weightMedium'
else:
return 'weightHigh'
def label_min_qt (row):
if row['minimum_quantity'] <= 4:
return 'mqLow'
elif row['minimum_quantity'] > 4 and row['minimum_quantity'] <= 16:
return 'mqMedium'
else:
return 'mqHigh'
products_train_exp5 = products
products_test_exp5 = products_test
products_train_exp5['priceInterval'] = products_train_exp5.apply (lambda row: label_price(row), axis=1)
products_train_exp5['weightInterval'] = products_train_exp5.apply (lambda row: label_weight(row), axis=1)
products_train_exp5['mqInterval'] = products_train_exp5.apply (lambda row: label_min_qt(row), axis=1)
products_test_exp5['priceInterval'] = products_test_exp5.apply (lambda row: label_price(row), axis=1)
products_test_exp5['weightInterval'] = products_test_exp5.apply (lambda row: label_weight(row), axis=1)
products_test_exp5['mqInterval'] = products_test_exp5.apply (lambda row: label_min_qt(row), axis=1)
exp4(['title', 'concatenated_tags', 'query', 'priceInterval', 'weightInterval', 'mqInterval'], products_train_exp5, products_test_exp5)
# -
# # 10. Trying to creating text features from numeric columns (automatic intervals)
#
# I didn't see an improvement in the previous experiment. Maybe it's because I didn't choose good intervals. I will try to define groups from these numeric columns using the k-means algorithm.
#
# ### Results:
# For **n-cluster = 23** we have got a good improvement.
# ```
# val: 0.926542 (mean of Accuracy and F1_score(micro))
# test: 0.962000 (Accuracy: 0.962000, F1_score(micro): 0.962000)
#
#
# crossvalidation: from 0.924776 to 0.926542.
# test: from 0.956000 to 0.962000
# test accuracy: from 0.956000 to 0.962000
# test F1_score(micro): from 0.956000 to 0.962000
# ```
#
#
#
# +
#2. Using a clustering algorhitm to define betters labels
def create_feature_kmeansPriceWeightMinimumQuantity(products_train, products_test, ncluster):
kMeansPipeline = Pipeline(
[
("scaler", StandardScaler()),
("kmeans", KMeans(n_clusters=ncluster, random_state=0)),
]
)
kmeansArray_train = kMeansPipeline.fit_predict(products_train[['price', 'weight', 'minimum_quantity']].dropna())
kmeansArray_test = kMeansPipeline.predict(products_test[['price', 'weight', 'minimum_quantity']])
kmeansSeries_train = pd.Series(kmeansArray_train, name="kmeans")
kmeansSeries_test = pd.Series(kmeansArray_test, name="kmeans")
products_train_exp6 = pd.concat([products_train, kmeansSeries_train], axis=1)
products_train_exp6['kmeansPriceWeightMinimumQuantity'] = 'grupo' + products_train_exp6['kmeans'].astype(str)
products_test_exp6 = pd.concat([products_test, kmeansSeries_test], axis=1)
products_test_exp6['kmeansPriceWeightMinimumQuantity'] = 'grupo' + products_test_exp6['kmeans'].astype(str)
return(products_train_exp6, products_test_exp6)
def exp6(nclusterList, products_train, products_test):
for ncluster in nclusterList:
print(ncluster)
products_train_exp6, products_test_exp6 = create_feature_kmeansPriceWeightMinimumQuantity(products_train, products_test, ncluster)
print('<| n-cluster:', ncluster, '|>')
exp4(['title', 'concatenated_tags', 'query', 'kmeansPriceWeightMinimumQuantity'], products_train_exp6, products_test_exp6)
# exp6(range(3, 50, 1), products, products_test) #best nclust=23
# exp6([23], products, products_test) #best nclust=23
# exp6([8], products, products_test)
#trying to explore [15,21] ncluster interval
# exp5(range(15, 21, 1), products, products_test)
# exp5([2], products, products_test)
# -
# # 11. Trying others sckit-learn classifiers
#
# The current classifier is using the SGDClassifier.
# I would like to try other models.
# The [Choosing the right estimator page]( https://scikit-learn.org/stable/tutorial/machine_learning_map/index.html) made me curious to explore.
#
# ### Results
# I did not manage to improve the current performance.
# As the train time started getting longer I decided, for now, to stay with SGDClassifier.
# +
modelsToTest = [
# SGDClassifier(random_state=0),
# LinearSVC(random_state=0),
# KNeighborsClassifier(),
# SVC(random_state=0),
# RandomForestClassifier(random_state=0),
# ExtraTreesClassifier(random_state=0),
# AdaBoostClassifier(random_state=0),
# GradientBoostingClassifier(random_state=0)
]
def createPipeline(classifier):
stop_portuquese_fromWordclouds = ['de', 'do', 'dos', 'com', 'em', 'o', 'e', 'para', 'em']
return Pipeline(
[
("vect", CountVectorizer(binary=True, max_df=0.5, max_features=None, ngram_range=(1, 2), strip_accents=None, stop_words=stop_portuquese_fromWordclouds )),
("clf", classifier),
]
)
products_train_exp7, products_test_exp7 = create_feature_kmeansPriceWeightMinimumQuantity(products, products_test, ncluster=23)
def exp7(textColumnsList, products, products_test, pipeline):
X_train = concatCols(products, textColumnsList)
X_test = concatCols(products_test, textColumnsList)
Y_train = products['category']
scoring = {'Accuracy': 'accuracy', 'F1_score': 'f1_micro' }
parameters = {
}
if __name__ == "__main__":
# multiprocessing requires the fork to happen in a __main__ protected block
# find the best parameters for both the feature extraction and the classifier
grid_search = GridSearchCV(pipeline, parameters, n_jobs=-1, verbose=1, scoring=scoring, refit="Accuracy", return_train_score=True)
print("Performing grid search...")
print("pipeline:", [name for name, _ in pipeline.steps])
print("parameters:")
pprint(parameters, depth=2, compact=True)
t0 = time()
grid_search.fit(X_train, Y_train)
print("done in %0.3fs" % (time() - t0))
print()
print("Best parameters set:")
best_parameters = grid_search.best_estimator_.get_params()
for param_name in sorted(parameters.keys()):
print("\t%s: %r" % (param_name, best_parameters[param_name]))
print(" val: %f (mean of Accuracy and F1_score(micro))" % grid_search.best_score_)
predicted = grid_search.best_estimator_.predict(X_test)
evaluate_test(predicted)
for m in modelsToTest:
print('<|', m, '>|')
pipeline = createPipeline(m)
exp7(['title', 'concatenated_tags', 'query', 'kmeansPriceWeightMinimumQuantity'], products_train_exp7, products_test_exp7, pipeline)
print('-----------------------------------------------------------------------------')
# -
# # 12. Trying XGBoost Classifier
#
# I would to see if I could get a better result with this classifier.
# There is some incompabilites between sckit-learn sparse matrix data structure and what this classifier receives.
# When I tried using the current CountVectorizer ngram_range=(1, 2) parameter, the program exiting saying that I dont have 36Gib needed.
# So I am trying a slight simpler version without this parameter.
# Last time took more than 40minutes. 🤞
#
# ### Result
# **No improvement**
# +
products_train_exp8, products_test_exp8 = create_feature_kmeansPriceWeightMinimumQuantity(products, products_test, ncluster=23)
def exp8(textColumnsList, products, products_test):
X_train = concatCols(products, ['title', 'concatenated_tags', 'query', 'kmeansPriceWeightMinimumQuantity'])
X_test = concatCols(products_test, ['title', 'concatenated_tags', 'query', 'kmeansPriceWeightMinimumQuantity'])
Y_train = products['category']
stop_portuquese_fromWordclouds = ['de', 'do', 'dos', 'com', 'em', 'o', 'e', 'para', 'em']
pPipeline(
[
("vect", CountVectorizer(binary=True, max_df=0.5, max_features=None, ngram_range=(1, 2), strip_accents=None, stop_words=stop_portuquese_fromWordclouds )),
("clf", classifier),
]
)
X = X_train
y = Y_train
# countvectorizer = CountVectorizer(binary=True, max_df=0.5, max_features=None, ngram_range=(1, 2), strip_accents=None, stop_words=stop_portuquese_fromWordclouds)
countvectorizer = CountVectorizer(binary=True, max_df=0.5, max_features=None, strip_accents=None, stop_words=stop_portuquese_fromWordclouds)
# countvectorizer = CountVectorizer()
Xt = countvectorizer.fit_transform(X)
# Convert from csr_matrix to sparse DataFrame
Xt_df_sparse = pd.DataFrame.sparse.from_spmatrix(Xt)
print("DF density: {}".format(Xt_df_sparse.sparse.density))
classifier = XGBClassifier(random_state=0)
classifier.fit(Xt_df_sparse, y)
Xt_t = countvectorizer.transform(X_test)
# Convert from csr_matrix to sparse DataFrame
Xt_df_sparse2 = pd.DataFrame.sparse.from_spmatrix(Xt_t)
predicted = classifier.predict(Xt_df_sparse2)
evaluate_test(predicted)
exp8(['title', 'concatenated_tags', 'query', 'kmeansPriceWeightMinimumQuantity'], products_train_exp8, products_test_exp8)
# -
# # 12. Trying to optimize SGDClassifier hyperparameters
#
# Parameters to search
# ```
# 'clf__alpha': (0.1, 0.3, 0.01, 0.03, 0.001, 0.003, 0.0001, 0.0003, 0.00001, 0.000001),
# 'clf__penalty': ('l2', 'l1', 'elasticnet'),
# 'clf__max_iter': (1000, 2000, 5000),
# 'clf__n_iter_no_change': (5, 10, 20),
# ```
#
#
# ### Results
# **No improvement**
# +
products_train_exp9, products_test_exp9 = create_feature_kmeansPriceWeightMinimumQuantity(products, products_test, ncluster=23)
def exp9(textColumnsList, products, products_test):
X_train = concatCols(products, textColumnsList)
X_test = concatCols(products_test, textColumnsList)
Y_train = products['category']
scoring = {'Accuracy': 'accuracy', 'F1_score': 'f1_micro' }
stop_portuquese_fromWordclouds = ['de', 'do', 'dos', 'com', 'em', 'o', 'e', 'para', 'em']
pipeline = Pipeline(
[
("vect", CountVectorizer(binary=True, max_df=0.5,
max_features=None,
ngram_range=(1, 2),
strip_accents=None,
stop_words=stop_portuquese_fromWordclouds )),
("clf", SGDClassifier(random_state=0)),
]
)
parameters = {
# 'clf__alpha': (0.1, 0.3, 0.01, 0.03, 0.001, 0.003, 0.0001, 0.0003, 0.00001, 0.000001),
# 'clf__penalty': ('l2', 'l1', 'elasticnet'),
# 'clf__max_iter': (1000, 2000, 5000),
# 'clf__n_iter_no_change': (5, 10, 20),
# 'clf__learning_rate': ('optimal', 'adaptive', 'constant')
'clf__loss': ('hinge', 'log', 'modified_huber', 'squared_hinge', 'perceptron', 'huber', 'epsilon_insensitive', 'squared_epsilon_insensitive')
}
if __name__ == "__main__":
# multiprocessing requires the fork to happen in a __main__ protected block
# find the best parameters for both the feature extraction and the classifier
grid_search = GridSearchCV(pipeline, parameters, n_jobs=-1, verbose=1, scoring=scoring, refit="Accuracy", return_train_score=True)
print("Performing grid search...")
print("pipeline:", [name for name, _ in pipeline.steps])
print("parameters:")
pprint(parameters, depth=2, compact=True)
t0 = time()
grid_search.fit(X_train, Y_train)
print("done in %0.3fs" % (time() - t0))
print()
print("Best parameters set:")
best_parameters = grid_search.best_estimator_.get_params()
for param_name in sorted(parameters.keys()):
print("\t%s: %r" % (param_name, best_parameters[param_name]))
print(" val: %f (mean of Accuracy and F1_score(micro))" % grid_search.best_score_)
predicted = grid_search.best_estimator_.predict(X_test)
evaluate_test(predicted)
exp9(['title', 'concatenated_tags', 'query', 'kmeansPriceWeightMinimumQuantity'], products_train_exp9, products_test_exp9)
# -
# # Exporting data for next exercise
#
products_test[['title', 'concatenated_tags', 'query', 'price', 'weight', 'minimum_quantity']].to_json(path_or_buf="/usr/src/data/test_products.json", orient='table', index=False)
| training/experiments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/priyanshgupta1998/Programming-/blob/master/python/hacker_earth_mocktest.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="io_Q5dkm2IOv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5e89a684-c4e4-49ca-df1f-443a143ea0bb"
liss = []
def binary(x):
if(x>1):
binary(x//2)
liss.append(x%2)
return liss
print(binary(3))
# + id="d-cjFGaK1zoe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="dbb4ca7e-e85e-466a-8a59-d44e8223ef02"
s = '1123'
s +=str(4)
s
# + id="irED736f1zs0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="44b160aa-6aaf-495b-fcc7-d15a54746a93"
def convert(s):
new = ""
for x in s:
new += x
return new
s = ['g', 'e', 'e', 'k', 's', 'f', 'o', 'r', 'g', 'e', 'e', 'k', 's']
print(convert(s))
# + id="y9-nO4xd1zzi" colab_type="code" colab={}
# + id="7vr_HJCr1z3j" colab_type="code" colab={}
# + id="iLQ10Hfa1z7b" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="f2b54c85-cda3-4dcf-c81f-e0450dd591bc"
def binary(x ,liss):
if(x>1):
binary(x//2 , liss )
liss.append(str(x%2))
return liss
num = list(map(int , input().split()))
for i in num:
count = 0
for j in range(1 , i+1):
liss = []
bini = binary(j , liss)
#print(j , bini)
new = ""
for k in bini:
new += k
if(new == new[::-1]):
count+=1
print(count , end =" ")
# + id="vFOg8AUf1zyF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4b2360da-9933-4df8-8182-102143252fe3"
# '@Priyansh Gupta'
from itertools import permutations
liss = []
# n1 = int(input())
# n2 = int(input())
# n3 = int(input())
# n4 = int(input())
# print(n1,n2,n3,n4)
inp = list(map(int, input().split()))
#print(inp)
perm = permutations(inp)
for i in list(perm):
k = list(i)
c = 0
for j in range(len(k)-1):
if(k[j] !=k[j+1]):
c+=1
if(c == len(k) -1):
new =""
for s in k:
new += str(s)
liss.append(new)
v = (10**9 + 7)
print(len(set(liss))%v)
# + id="sygFAwKuA5OL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="9ddbf247-2e9c-4b65-d6b3-57e732f73b81"
inp = list(map(int, input().split()))
print(inp)
# + id="cy2RHTmGA5Mq" colab_type="code" colab={}
# + id="HVSql05iA5LG" colab_type="code" colab={}
# + id="Dh09ffLZA5JN" colab_type="code" colab={}
# + id="Ykgyqeu7A5He" colab_type="code" colab={}
# + id="tZtHCBDIA5Fy" colab_type="code" colab={}
# + id="7S7Sb2aTA5D2" colab_type="code" colab={}
# + id="lK_6aBaBA5CD" colab_type="code" colab={}
# + id="wKXy-pQjA4_o" colab_type="code" colab={}
# + id="SIhItTFcA48r" colab_type="code" colab={}
# + id="I86o3ZfX1zvy" colab_type="code" colab={}
| python/hacker_earth_mocktest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# __Split__: This will split the array.
#
# There are 5 types:
#
# 1.Vertical split.
#
# 2.Horizonal split.
#
# 3.Depth_split.
#
# 4.array_split.
#
# 5.split.
import numpy as np
a = np.arange(1, 10).reshape(3, 3)
print(a)
b = np.vsplit(a, 3)
print(b)
c = np.arange(1, 17).reshape(4, 4)
print(c)
d = np.vsplit(c, 2)
print(d)
e = np.vsplit(c, np.array([3]))
#example for horizontal split.
import numpy as np
a = np.arange(16).reshape(4, 4)
print(a)
b = np.hsplit(a, 2)
print(b)
#example for dsplit.
import numpy as np
a = np.arange(36).reshape(3, 3, 2, 2)
print(a)
b = np.dsplit(a, 2)
print(b)
#example for array split
import numpy as np
a = np.arange(26)
b = np.array_split(a, 2)
print(a)
print(b)
#example for split.
import numpy as np
a = np.arange(16)
b = np.split(a, 4)
print(a)
print(b)
| Split operations Numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Logistic Regression
# Modules
import torch
import torch.nn as nn
import torchvision
import torchvision.transforms as transforms
# Hyper-parameters
input_size = 784
num_classes = 10
num_epochs = 5
batch_size = 100
learning_rate = 0.001
# MNIST dataset (images and labels)
# +
train_dataset = torchvision.datasets.MNIST(root='../../data',
train=True,
transform=transforms.ToTensor(),
download=True)
test_dataset = torchvision.datasets.MNIST(root='../../data',
train=False,
transform=transforms.ToTensor())
# -
# Data loader (input pipeline)
# +
train_loader = torch.utils.data.DataLoader(dataset=train_dataset,
batch_size=batch_size,
shuffle=True)
test_loader = torch.utils.data.DataLoader(dataset=test_dataset,
batch_size=batch_size,
shuffle=False)
# -
# Logistic regression model
model = nn.Linear(input_size, num_classes)
# Loss and optimizer
#
# `nn.CrossEntropyLoss()` computes softmax internally
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(model.parameters(), lr=learning_rate)
# Train the model
total_step = len(train_loader)
for epoch in range(num_epochs):
for i, (images, labels) in enumerate(train_loader):
# Reshape images to (batch_size, input_size)
images = images.reshape(-1, 28*28)
# Forward pass
outputs = model(images)
loss = criterion(outputs, labels)
# Backward and optimize
optimizer.zero_grad()
loss.backward()
optimizer.step()
if (i+1) % 100 == 0:
print ('Epoch [{}/{}], Step [{}/{}], Loss: {:.4f}'
.format(epoch+1, num_epochs, i+1, total_step, loss.item()))
# Test the model
#
# In test phase, we don't need to compute gradients (for memory efficiency)
with torch.no_grad():
correct = 0
total = 0
for images, labels in test_loader:
images = images.reshape(-1, 28*28)
outputs = model(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum()
print('Accuracy of the model on the 10000 test images: {} %'.format(100 * correct / total))
# Save the model checkpoint
torch.save(model.state_dict(), 'model.ckpt')
| tutorials_jupyter/01-basics/logistic_regression/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="TUO9qXMa17F-"
# # Galaxy Classification with Vision Transformer - Hidden States Extraction
# + [markdown] id="S1y8Ta0p2BUj"
# **Hugging Face's Vision Transformer implementation**
#
# - https://huggingface.co/docs/transformers/model_doc/vit
# - https://github.com/huggingface/transformers/blob/v4.14.1/src/transformers/models/vit/modeling_vit.py
#
# **GalaxyZoo 2 Data**
#
# - https://www.kaggle.com/c/galaxy-zoo-the-galaxy-challenge
#
#
# **ViT papers**
#
# - https://arxiv.org/abs/2010.11929
#
# - https://arxiv.org/abs/2110.01024
#
# - https://arxiv.org/abs/2005.00928
#
#
# **Coding references**
#
# - https://pytorch.org/tutorials/beginner/transfer_learning_tutorial.html
#
# - https://pytorch.org/tutorials/beginner/data_loading_tutorial.html
#
# - https://github.com/jeonsworld/ViT-pytorch/blob/main/visualize_attention_map.ipynb
#
# + [markdown] id="xbvsPMIF2NM-"
# ## Check GPU
# + colab={"base_uri": "https://localhost:8080/"} id="cR4qPHGPzlQZ" outputId="4711a6c8-83e1-437b-9fc7-c08a51bd7950"
# !nvidia-smi -L
# + [markdown] id="s9p47BSZcqDV"
# ## Install HuggingFace Vision Transformer (ViT)
# + id="CwQbrI_zc36J"
# !pip install transformers
# + [markdown] id="zjFnClt8c8sZ"
# ## Import Libraries
# + colab={"base_uri": "https://localhost:8080/"} id="xktvwYkoc7ND" outputId="9f3bff5e-f50a-4b80-d050-a9ea53d20e21"
import cv2
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from PIL import Image
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, accuracy_score, recall_score, f1_score
import seaborn as sns
import torch
import torch.nn as nn
from torch.utils.data import Dataset, DataLoader
from torchvision import models, transforms, utils
import torch.optim as optim
from torch.optim import lr_scheduler
import time
import os
import zipfile
from copy import deepcopy
# %matplotlib inline
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print(device)
# vision transformer
from transformers import ViTModel, ViTConfig, ViTFeatureExtractor, ViTForImageClassification
# + colab={"base_uri": "https://localhost:8080/"} id="fiMSKRqsdFwZ" outputId="1291c3f4-db2b-4679-c70f-55059407b729"
torch.manual_seed(0)
# + [markdown] id="4t4jV38S2VLk"
# ## Unzip images from Google Drive
#
# + id="Drc6IgZJ0JQ1"
## The unzipping function
def unzipping(zip_file_name, dest_dir):
"""
Arguments
zip_file_name: (str) the file name of the zip file
dest_dir: (str) the name of the destination folder
"""
# check if the destination folder exists. Create one if not.
#if not os.path.exists(dest_dir):
# os.makedirs(dest_dir)
# unzipping
zip_f = zipfile.ZipFile(zip_file_name, 'r')
zip_f.extractall(path=dest_dir)
zip_f.close()
# + id="mTPNOjVOmEYN"
## directory on Google Drive
datasets_dir = 'drive/MyDrive/gz2_datasets'
## Unzip training, validation, and test images
unzipping(os.path.join(datasets_dir, 'images_train.zip'), './')
unzipping(os.path.join(datasets_dir, 'images_valid.zip'), './')
unzipping(os.path.join(datasets_dir, 'images_test.zip'), './')
# + [markdown] id="P_oRGGTk2aUM"
# ## Custom GalaxyZoo Dataset
# + id="Kw-QIg3w0NNZ"
## Custom Galaxy Zoo 2 Dataset
class GalaxyZooDataset(Dataset):
"""Galaxy Zoo Dataset"""
def __init__(self, csv_file, images_dir, transform=None):
"""
Args:
csv_file (string): path to the label csv
images_dir (string): path to the dir containing all images
transform (callable, optional): transform to apply
"""
self.labels_df = pd.read_csv(csv_file)
self.labels_df = self.labels_df[['galaxyID', 'label1']].copy()
self.images_dir = images_dir
self.transform = transform
def __len__(self):
"""
Returns the size of the dataset
"""
return len(self.labels_df)
def __getitem__(self, idx):
"""
Get the idx-th sample.
Outputs the image (channel first) and the true label
"""
if torch.is_tensor(idx):
idx = idx.tolist()
# galaxy ID
galaxyid = self.labels_df.iloc[idx, 0].astype(str)
# path of the image
image_path = os.path.join(self.images_dir, galaxyid + '.jpg')
# read the image
image = Image.open(image_path)
# apply transform (optional)
if self.transform is not None:
image = self.transform(image)
# read the true label
label = int(self.labels_df.iloc[idx, 1])
return image, label, int(galaxyid)
# + [markdown] id="TA13w0dJdYtM"
# ## Custom Data Transforms (for HuggingFace ViT)
# + id="Jv9zPo1Gdb2l"
def create_data_transforms(input_size=224):
"""
Create Pytorch data transforms for the GalaxyZoo datasets
Args:
input_size: final size of the image. Default=224.
Outputs:
train_transform: transform for the training data
valid_transform: transform for the validation data
test_transform: transform for the testing data
"""
# transforms for training data
train_transform = transforms.Compose([transforms.CenterCrop(input_size),
transforms.RandomRotation(90),
transforms.RandomHorizontalFlip(),
transforms.RandomVerticalFlip(),
transforms.RandomResizedCrop(input_size, scale=(0.8, 1.0), ratio=(0.99, 1.01)),
transforms.ToTensor(),
transforms.Normalize([0.5, 0.5, 0.5], [0.5, 0.5, 0.5])])
# transforms for validation data
valid_transform = transforms.Compose([transforms.CenterCrop(input_size),
transforms.ToTensor(),
transforms.Normalize([0.5, 0.5, 0.5], [0.5, 0.5, 0.5])])
# transforms for test data
test_transform = transforms.Compose([transforms.CenterCrop(input_size),
transforms.ToTensor(),
transforms.Normalize([0.5, 0.5, 0.5], [0.5, 0.5, 0.5])])
return train_transform, valid_transform, test_transform
# + [markdown] id="6RoYRv692wSk"
# ## Create Dataloaders
# + id="tX-8evCxvDPP" colab={"base_uri": "https://localhost:8080/"} outputId="351cc505-ab7c-4afc-8eab-592a23ab4b5a"
"""
Data Loader
"""
# the batch size
BATCH_SIZE = 32
# create transforms
train_transform, valid_transform, test_transform = create_data_transforms(input_size=224)
# create datasets
data_train = GalaxyZooDataset('gz2_train.csv', 'images_train', train_transform)
data_valid = GalaxyZooDataset('gz2_valid.csv', 'images_valid', valid_transform)
data_test = GalaxyZooDataset('gz2_test.csv', 'images_test', test_transform)
# dataloaders
train_loader = DataLoader(data_train, batch_size=BATCH_SIZE, shuffle=True)
valid_loader = DataLoader(data_valid, batch_size=BATCH_SIZE, shuffle=True)
test_loader = DataLoader(data_test, batch_size=BATCH_SIZE, shuffle=True)
# check the sizes
print("**Dataloaders**")
print("Number of training data: {} ({} batches)".format(len(data_train), len(train_loader)))
print("Number of validation data: {} ({} batches)".format(len(data_valid), len(valid_loader)))
print("Number of test data: {} ({} batches)".format(len(data_test), len(test_loader)))
print("===============================")
# + [markdown] id="tPGd44F6d0Zc"
# ## Evaluation Function (HuggingFace ViT)
# + id="tOk2QcRUd2hi"
def evaluate_model(model, loader, device):
"""
Evaluate (no gradient tracking) the model on a dataset.
Args:
model: the model to be tested
loader: a custom dataloader
device: gpu or cpu
Outputs:
prints out confusion matrix and several metrics (accuracy, recall, f1)
"""
# move to gpu
model = model.to(device)
# evaluation
model.eval()
# empty lists for results
y_true = []
y_pred = []
y_label = []
for images, labels, galaxy_id in loader:
images = images.to(device)
labels = labels.long().to(device)
with torch.no_grad():
outputs = model(images)
pred_logits = outputs.logits
_, pred_classes = torch.max(pred_logits.detach(), dim=1)
y_true += torch.squeeze(labels.cpu()).tolist()
y_pred += torch.squeeze(pred_classes).tolist()
y_label += torch.squeeze(galaxy_id.cpu()).tolist()
# create a DataFrame with columns 'GalaxyID', 'class', 'predicted labels'
predict_df = pd.DataFrame(data={'GalaxyID': y_label, 'class': y_true, 'pred': y_pred})
# galaxy classes
gxy_labels = ['Round Elliptical',
'In-between Elliptical',
'Cigar-shaped Elliptical',
'Edge-on Spiral',
'Barred Spiral',
'Unbarred Spiral',
'Irregular',
'Merger']
# confusion matrix
cm = confusion_matrix(y_true, y_pred, normalize='true')
cm_df = pd.DataFrame(cm, index=gxy_labels, columns=gxy_labels)
# accuracy of each class
for c in range(8):
print("Class {}: accuracy = {:.4f} ({})".format(c, cm[c,c]/sum(cm[c,:]), gxy_labels[c]))
print("================")
# accuracy
acc = accuracy_score(y_true, y_pred)
print("Total Accuracy = {:.4f}\n".format(acc))
# recall
recall = recall_score(y_true, y_pred, average='macro')
print("Recall = {:.4f}\n".format(recall))
# f1 score
F1 = f1_score(y_true, y_pred, average='macro')
print("F1 score = {:.4f}\n".format(F1))
# plot confusion matrix
sns.set(font_scale=1.6)
fig = plt.figure(figsize=(10, 10))
sns.heatmap(cm_df, annot=True, fmt=".1%", cmap="YlGnBu", cbar=False, annot_kws={"size": 16})
plt.show()
# + [markdown] id="rHtDpWQceCD5"
# ## Create Model and Load Weights (HuggingFace ViT_Base)
# + colab={"base_uri": "https://localhost:8080/", "height": 116, "referenced_widgets": ["a6e842a935aa442e83ccca0ab5a3fa1b", "880b745f39c443b98e63fed5a6fefe87", "f1d030b7f5f44a67bfc3b4de9a58dbb7", "2ae69e85a0e24a5398d5dbb993a27664", "dd2e22fb789b4ebe8d258d678ce88699", "6df86f6e686a4681a67f93b56d452f30", "c5d2650de5ff42b69078b0b5b49dff03", "64b430a4b9a14b32a699091b54a24b07", "121a9479b87140b69a394f135ed8786d", "d9d0e4f31b1645369f0ee2a25cbc4424", "60e41cf752ed436b868f00a34a5ee6a2", "5d534d32b87d4ebc9347b1552cb1a24b", "5338bd17722844448bdce579a09dccc0", "<KEY>", "<KEY>", "71268eb785d04ecdae1c872643ca2601", "e53816c6ccb340edb8c3e1c9a3ab0a6d", "<KEY>", "e1c434616da84eaab8444bc8be63aeea", "<KEY>", "<KEY>", "0c62241acdf44ce88c58f6db04a12d3f"]} id="wpfnnJ1ZeFih" outputId="6967a695-3593-4230-89eb-741127bbbd4b"
"""
Create ViT
"""
## Vision Transformer
model = ViTForImageClassification.from_pretrained('google/vit-base-patch16-224')
# change the last layer
model.classifier = nn.Linear(in_features=768, out_features=8, bias=True)
print("Number of trainable parameters: {}".format(sum(param.numel() for param in model.parameters() if param.requires_grad)))
"""
Load Pretrained Weights (Current Best Model: 010822B)
"""
pth_filename = 'gz2_hug_vit_010822B.pth'
model.load_state_dict(torch.load(os.path.join('drive/MyDrive/gz2_datasets', pth_filename)))
# + [markdown] id="UXdoC_mCjmGr"
# ### Check Performance
#
# Make sure the correct weights were loaded. The testing accuracy should be **85.00%**.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="Idy5x59cjwDR" outputId="4509d9e9-af73-4ace-be73-fc0d61d891c3"
evaluate_model(model, test_loader, device)
# + [markdown] id="rQtHlf9UYXP-"
# ## Model Predictions
# + colab={"base_uri": "https://localhost:8080/", "height": 224} id="fzR6w9PaYaZN" outputId="eab11140-07f2-4bd3-f449-fd57a3b7d3d3"
"""
Load Prediction CSV (Current Best Model: 010822B)
"""
df = pd.read_csv("gz2_hug_vit_010822B_predictions.csv")
print(df.shape)
df.head()
# + [markdown] id="PNxMp21We1Cr"
# ## Hidden States Extraction
# + id="Guc7vk8Bez41"
def vit_hidden_states(model, layer, data_loader, df, device, dest_dir=None):
"""
Extract the hidden_states of ViT
Args:
model: vit model
layer: the layer to extract the hidden states min=0, max=11
data_loader: train/validation/test data loader
df: the dataframe containing the true/predicted labels
device: gpu or cpu
dest_dir: the destination dir to save the maps. If None, images of the
first batch will be displayed and no files will be save.
default: None
"""
# move to gpu
model = model.to(device)
# evaluation
model.eval()
## create a dict containing 'GalaxyID', 'y_true', 'y_pred', 'h_0', 'h_1', ..., 'h_768' (hidden_states_of_CLS)
# we will later convert this dict into a dataframe
hidden_states_dict = dict()
# create fields in the dicts
hidden_states_dict["GalaxyID"] = []
hidden_states_dict["y_true"] = []
hidden_states_dict["y_pred"] = []
for hs in range(768):
hidden_states_dict["h_{}".format(hs)] = []
# iterate batches
for images, _, galaxy_id in data_loader:
images = images.to(device)
with torch.no_grad():
## outputs has 3 items:
# outputs[0] is a tensor, the last layer output, with shape (size_batch, n_seq, hidden_dim)
# outputs[1] is a tuple with length=n_layers, each with shape (size_batch, n_seq, hidden_dim)
# outputs[2] is a tuple with length=n_layers, each with shape (size_batch, n_seq, hidden_dim, hidden_dim)
outputs = model.vit(images, output_attentions=True, output_hidden_states=True, return_dict=model.config.use_return_dict)
# log the data row by row
# iterate galaxies
for g in range(outputs[0].shape[0]):
# galaxy id
gid = galaxy_id.cpu().numpy()[g]
hidden_states_dict["GalaxyID"].append(gid)
# true label
true_label = df[df['GalaxyID']==gid]['class'].values[0]
hidden_states_dict["y_true"].append(true_label)
# predicted label
pred_label = df[df['GalaxyID']==gid]['pred'].values[0]
hidden_states_dict["y_pred"].append(pred_label)
# iterate hidden states
for hs in range(768):
# log the hs-th hidden state of CLS
hidden_states_dict["h_{}".format(hs)].append(outputs[1][layer][g][0].cpu().numpy()[hs])
# -- (all images read) --
# convert dict into a dataframe
hidden_states_df = pd.DataFrame.from_dict(hidden_states_dict)
# sort the Galaxy ID
hidden_states_df = hidden_states_df.sort_values(by=['GalaxyID'])
print("Layer {} done".format(layer))
# save to dir
if dest_dir:
csv_path = os.path.join(dest_dir, "HiddenStates_Layer{}.csv".format(layer))
hidden_states_df.to_csv(csv_path, index=False)
return None
else:
return hidden_states_df
# + [markdown] id="WG2Zl6mhC3rt"
# ### Test Code
# + id="p6X3VhNYnUJi"
res = vit_hidden_states(model, 11, test_loader, df, device, dest_dir=None)
res.head()
# + [markdown] id="NqdJ894cft-K"
# ## Extract hidden states from all layers (from layer=0 to layer=11)
# + id="ZCxlWv9nnhOg" colab={"base_uri": "https://localhost:8080/"} outputId="5007221c-21cc-4049-91af-f073fc6cfa13"
for layer in range(12):
# extract hidden features
_ = vit_hidden_states(model, layer, test_loader, df, device, dest_dir='drive/MyDrive/gz2_datasets/features/vit_base_hidden_states_all_layers')
# + id="N5Hs55O1rSer"
| gz2_hidden_states_ViT_base.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import requests
from bs4 import BeautifulSoup
import pandas as pd
import numpy as np
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
import time
import os
athlete_no = '6483989' # <NAME>
#athlete_no = '3013148' # <NAME>
athlete_no = '5824213' # <NAME>
url = 'https://www.athletic.net/TrackAndField/Athlete.aspx?AID='+athlete_no+'#!/L0'
response = requests.get(url)
# +
chromedriver = "/usr/local/bin/chromedriver"
os.environ["webdriver.chrome.driver"] = chromedriver
driver = webdriver.Chrome(chromedriver)
driver.get("https://www.athletic.net")
driver.get("https://www.athletic.net/account/login/?ReturnUrl=%2Fdefault.aspx")
# %load_ext dotenv
# %dotenv
EMAIL = '<EMAIL>'
#PASSWORD = os.environ.get('PASSWORD')
PASSWORD = '<PASSWORD>'
username_form = driver.find_element_by_name("email")
username_form.send_keys(EMAIL) # enter email
username_form.send_keys(Keys.RETURN)
password_form=driver.find_element_by_name("password") # note another approach
password_form.send_keys(PASSWORD) # enter password
password_form.send_keys(Keys.RETURN)
# -
# ## Using Pandas
# +
header = {
"User-Agent": "Mozilla/5.0 (X11; Linux x86_64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/50.0.2661.75 Safari/537.36",
"X-Requested-With": "XMLHttpRequest"
}
r = requests.get(url, headers=header)
dfs = pd.read_html(r.text)
# +
for df in dfs:
if df[0][0] == '400 Meters':
all_years = df
print(all_years)
# -
dfs[2]
# ### Pull data for several athletes
#
# Loop over a list of athletes from a csv file and read their 400 Meters numbers
header = {
"User-Agent": "Mozilla/5.0 (X11; Linux x86_64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/50.0.2661.75 Safari/537.36",
"X-Requested-With": "XMLHttpRequest"
}
# given a time string, return a float for the time in seconds
def cleantime(time_str):
time_str = time_str.replace('PR','')
time_str = time_str.replace('*','')
time_str = time_str.replace('c','')
time_str = time_str.replace('h','')
time_str = time_str.strip()
if ':' in time_str:
t = time_str.split(':')
time = float(t[0])*60 + float(t[1])
elif time_str.isdigit():
time = float(time_str)
else: time = np.nan
return(time)
# +
#file = '/Users/dana/metis/git/Metis_Bootcamp/Luther_Project/athletes.csv'
#file = '/Users/dana/metis/git/Metis_Bootcamp/Luther_Project/athlete_list.csv'
#file = '/Users/dana/metis/git/Metis_Bootcamp/Luther_Project/athlete_list_boy1600.csv'
file = '/Users/dana/metis/git/Metis_Bootcamp/Luther_Project/athlete_list_girl400.csv'
athlete_list = pd.read_csv(file)
# -
# Given a list of athletes, scrap their PR times in the 400Meter race
# and add to dataframe
all_data = athlete_list[['ID','Name','School_ID','School','District']]
all_data = all_data.set_index(['ID'])
all_data['9_PR'] = np.nan
all_data['10_PR'] = np.nan
all_data['11_PR'] = np.nan
all_data['12_PR'] = np.nan
all_data['Grad_Yr'] = 0
all_data.Grad_Yr = all_data.Grad_Yr.astype(int)
all_data.head()
all_data.shape
all_data1 = all_data[0:2000]
all_data2 = all_data[2000:4000]
all_data3 = all_data[4000:6000]
all_data4 = all_data[6000:8000]
#all_data5 = all_data[8000:10000]
# +
all_data = all_data4
Event = '400 Meters'
#Event = '1600 Meters'
n = 0
for ID, row in all_data.iterrows():
ID_str = str(ID)
url = 'https://www.athletic.net/TrackAndField/Athlete.aspx?AID='+ID_str+'#!/L0'
r = requests.get(url, headers=header)
dfs = pd.read_html(r.text)
for df in dfs:
if df[0][0] == Event:
n += 1
if n%100 == 0: print(n)
df.drop(df.index[0:1],inplace=True)
for index, row in df.iterrows():
if 'Indoor' not in str(row[0]):
# print(f"grade {int(row[1])}, time {row[2]}")
#row[1]= str(row[1]).replace('-','')
g = str(row[1])
g = g.replace('-','')
g = g.replace('.0','')
#print(ID_str,g)
if (g == '9' or g == '10' or g == '11' or g == '12'):
#grade = str(int(row[1]))
grade = g
else:
grade = '0'
time = cleantime(str(row[2]))
# print(grade,time)
all_data.at[ID,grade+"_PR"] = time
# find the graduation year
if g == '12':
gy = str(row[0])
gy = gy.replace(' Outdoor','')
all_data.at[ID,'Grad_Yr'] = int(gy)
#print(int(gy))
# -
all_data4 = all_data
all_data1
all_data = pd.concat([all_data1,all_data2,all_data3,all_data4])
#all_data = pd.concat([all_data1,all_data2])
all_data.columns
#all_data.drop(['8_PR'], axis=1,inplace=True)
#all_data.drop(['7_PR'], axis=1,inplace=True)
all_data.drop(['0_PR'], axis=1,inplace=True)
all_data.dropna(inplace=True)
all_data.dropna().shape
all_data.to_csv("allDistrict_girl400.csv")
all_data.shape
all_data.head()
| 2-Luther_Project/.ipynb_checkpoints/Project_Luther_athletes-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import dagstermill
import pandas as pd
import random
import matplotlib.pyplot as plt
from dbt_example.resources import postgres
from dagster import ModeDefinition
# + tags=["parameters"]
run_results = {}
# -
context = dagstermill.get_context(
mode_def=ModeDefinition(resource_defs={'db': postgres}),
run_config={
'resources': {
'db': {'config': {'db_url': 'postgresql://dbt_example:dbt_example@localhost:5432/dbt_example'}}
}
}
)
with context.resources.db.connect() as conn:
normalized_cereals = pd.read_sql('select * from normalized_cereals', con=conn)
normalized_cereals.head()
# +
colormap = {'H':'orangered', 'C':'cornflowerblue'}
xs = normalized_cereals['normalized_calories'].apply(lambda x: x + random.random() * 3 - 1.5)
ys = normalized_cereals['normalized_protein'].apply(lambda x: x + random.random()/10 - .05)
colors = normalized_cereals['type'].apply(lambda x: colormap[x])
labels = normalized_cereals['name']
fig, ax = plt.subplots(figsize=(15,15))
ax.scatter(x=xs, y=ys, c=colors, alpha=0.5)
ax.set_xlabel("calories")
ax.set_label("protein")
for x, y, label in zip(xs, ys, labels):
plt.annotate(
label,
(x,y),
textcoords="offset points",
xytext=(3, 10),
ha='left',
size=7
)
plt.show()
plot_path = 'cereal_analysis_{run_id}.pdf'.format(run_id=context.run_id)
fig.savefig(plot_path, bbox_inches='tight')
# -
dagstermill.yield_result(plot_path)
| examples/dbt_example/dbt_example/notebooks/Analyze Cereals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### WeatherPy
# ----
#
# #### This analysis will use Python scripting to visualize the weather of 600+ cities around the world with varying distances from the equator. The script will make external calls to the OpenWeatherMap API.
#
# ### Outcomes: Build a series of visualizations (scatter plots) to show the following relationships:
#
# #### Temperature (F) vs. Latitude
# #### Humidity (%) vs. Latitude
# #### Cloudiness (%) vs. Latitude
# #### Wind Speed (mph) vs. Latitude
#
# ### Framework :
#
# #### Python Requests
# #### APIs
# ### JSON Traversals
#
# ### Analysis:
#
# #### The weather becomes significantly warmer the closer we get to the equator (0 Degrees Latitude). On 4/13/2020, the southern hemisphere tends to be warmer than the northern hemisphere.
#
#
# #### Humidity in cities within a range of +/- 19 degrees of the equator appear to be consistently more humid. That said, the relationship between Latitude and Humidity is weak. High and Low humidity appear randomly in both hemispheres outside of the +/- 19 degree range.
#
#
# #### There is no strong relationship between latitude and cloudiness.
#
#
# #### There is no strong relationship between latitude and wind speed. However, in the northern hemisphere, there are a few cities that have above average (20 mph) windspeeds.
#
#
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import requests
import time
from scipy.stats import linregress
# Import API key
from api_keys import weather_api_key
#print (weather_api_key)
# Incorporated citipy to determine city based on latitude and longitude
from citipy import citipy
# Output File (CSV)
output_data_file = "output_data/cities.csv"
# Range of latitudes and longitudes
lat_range = (-90, 90)
lng_range = (-180, 180)
# +
# Create Northern and Southern Hemisphere DataFrames
# -
# ## Generate Cities List
# +
# List for holding lat_lngs and cities
lat_lngs = []
cities = []
# Create a set of random lat and lng combinations
lats = np.random.uniform(lat_range[0], lat_range[1], size=1500)
lngs = np.random.uniform(lng_range[0], lng_range[1], size=1500)
lat_lngs = zip(lats, lngs)
# Identify nearest city for each lat, lng combination
for lat_lng in lat_lngs:
city = citipy.nearest_city(lat_lng[0], lat_lng[1]).city_name
# If the city is unique, then add it to a our cities list
if city not in cities:
cities.append(city)
# Print the city count to confirm sufficient count
len(cities)
#cities
# -
# ### Perform API Calls
# * Perform a weather check on each city using a series of successive API calls.
# * Include a print log of each city as it'sbeing processed (with the city number and city name).
#
# +
#OpenWeather API URL...
url = "http://api.openweathermap.org/data/2.5/weather?"
units = "imperial"
# set up the query URL and make the request
weatherAPI_url = url + "appid=" + weather_api_key + "&units=" + units + "&q="
weatherAPI_response = requests.get(weatherAPI_url + cities[2])
#if weatherAPI_response.status_code == 200:
# print("yes")
print("Beginning Data Retrival")
print("------------------------------")
group_counter = 1
rec_counter = 1
weather_data = {"City":[],"Lat":[],"Lng":[],"Max Temp":[],"Humidity":[],"Cloudiness":[],
"Wind Speed":[],"Country":[],"Date":[]}
for city in cities:
weatherAPI_response = requests.get(weatherAPI_url + city)
weatherAPI_response_json = weatherAPI_response.json()
if weatherAPI_response.status_code == 200:
weather_data["City"].append(city)
weather_data["Cloudiness"].append(weatherAPI_response_json['clouds']['all'])
weather_data["Country"].append(weatherAPI_response_json['sys']['country'])
weather_data["Date"].append(weatherAPI_response_json['dt'])
weather_data["Humidity"].append(weatherAPI_response_json['main']['humidity'])
weather_data["Lat"].append(weatherAPI_response_json['coord']['lat'])
weather_data["Lng"].append(weatherAPI_response_json['coord']['lon'])
weather_data["Max Temp"].append(weatherAPI_response_json['main']['temp_max'])
weather_data["Wind Speed"].append(weatherAPI_response_json['wind']['speed'])
if rec_counter <= 50:
print(f"Processing Record {rec_counter} of Set {group_counter} | {city}")
rec_counter += 1
else:
rec_counter = 0
group_counter += 1
print(f"Processing Record {rec_counter} of Set {group_counter} | {city}")
rec_counter += 1
else:
print("City not found. Skipping...")
print("-------------------------")
print("Data Retrieval Complete")
print("-------------------------")
# -
# ### Convert Raw Data to DataFrame
# * Export the city data into a .csv.
# * Display the DataFrame
weather_df = pd.DataFrame(weather_data)
weather_df.head()
# Exporting the city data into csv
weather_df.to_csv('weather_data.csv', encoding='utf-8', index=False)
# ## Inspect the data and remove the cities where the humidity > 100%.
# ----
# Skip this step if there are no cities that have humidity > 100%.
over100_df = weather_df[weather_df['Humidity'] > 100]
print (over100_df)
# +
# Get the indices of cities that have humidity over 100%.
# N/A
# +
# Make a new DataFrame equal to the city data to drop all humidity outliers by index.
# Passing "inplace=False" will make a copy of the city_data DataFrame, which we call "clean_city_data".
# N/A -
#weather_df.drop(weather_df[weather_df['Humidity'] > 100].index, inplace = False)
# +
# Extract relevant fields from the data frame
# Export the City_Data into a csv
# -
# ## Plotting the Data
# * Use proper labeling of the plots using plot titles (including date of analysis) and axes labels.
# * Save the plotted figures as .pngs.
# ## Latitude vs. Temperature Plot
# +
plt.scatter(weather_df["Lat"],weather_df["Max Temp"], marker="o", alpha=0.75, edgecolor="black")
plt.title("City Latitude vs. Max Temperature (04/13/20)")
plt.xlabel("Latitude")
plt.ylabel("Max Temperature (F)")
plt.ylim(0,105)
plt.grid(True)
plt.savefig("LatVsTemp.png")
plt.show()
# The weather becomes significantly warmer the closer we get to the equator (0 Degrees Latitude).
# On 4/13/2020, the southern hemisphere tends to be warmer than the northern hemisphere.
# -
# ## Latitude vs. Humidity Plot
# +
plt.scatter(weather_df["Lat"],weather_df["Humidity"], marker="o", alpha=0.75, edgecolor="black")
plt.title("City Latitude vs. Humidity (04/13/20)")
plt.xlabel("Latitude")
plt.ylabel("Humidity (%)")
plt.ylim(0,105)
plt.grid(True)
plt.savefig("LatVsHum.png")
plt.show()
# Humidity in cities within a range of +/- 19 degrees of the equator appear to be consistently more humid. That said, the
# relationship between Latitude and Humidity is weak. High and Low humidity appear randomly in both hemispheres outside of the
# # +/- 19 degree range.
# -
# ## Latitude vs. Cloudiness Plot
# +
plt.scatter(weather_df["Lat"],weather_df["Cloudiness"], marker="o", alpha=0.75, edgecolor="black")
plt.title("City Latitude vs. Cloudiness (04/13/20)")
plt.xlabel("Latitude")
plt.ylabel("Cloudiness")
plt.ylim(0,105)
plt.grid(True)
plt.savefig("LatVsCloud.png")
plt.show()
# There is no strong relationship between latitude and cloudiness.
# -
# ## Latitude vs. Wind Speed Plot
# +
plt.scatter(weather_df["Lat"],weather_df["Wind Speed"], marker="o", alpha=0.75, edgecolor="black")
plt.title("City Latitude vs. Wind Speed (04/13/20)")
plt.xlabel("Latitude")
plt.ylabel("Wind Speed")
plt.ylim(0,50)
plt.grid(True)
plt.savefig("LatVsWndspd.png")
plt.show()
# There is no strong relationship between latitude and wind speed. However, in the northern hemisphere,
# there are a few cities that have above average (20 mph) windspeeds.
# -
# ## Linear Regression
# +
# OPTIONAL: Create a function to create Linear Regression plots
# -
# Create Northern and Southern Hemisphere DataFrames
northern_df = weather_df.loc[pd.to_numeric(weather_df["Lat"]).astype(float) > 0, :]
southern_df = weather_df.loc[pd.to_numeric(weather_df["Lat"]).astype(float) < 0, :]
# #### Northern Hemisphere - Max Temp vs. Latitude Linear Regression
x_values = pd.to_numeric(northern_df['Lat']).astype(float)
y_values = pd.to_numeric(northern_df['Max Temp']).astype(float)
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
print(f"Regression line equation is: {line_eq}")
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('Latitude')
plt.ylabel('Max Temperature (F)')
plt.title('Northern Hemisphere - Max Temp vs. Latitude Linear Regression')
print(f"The r-squared is: {rvalue}")
print("Strong relationship with Temp and Latitude: Temperature decreases the further north (- lat) you go from the equator.")
plt.show()
# #### Southern Hemisphere - Max Temp vs. Latitude Linear Regression
x_values = pd.to_numeric(southern_df['Lat']).astype(float)
y_values = pd.to_numeric(southern_df['Max Temp']).astype(float)
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
print(f"Regression line equation is: {line_eq}")
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('Latitude')
plt.ylabel('Max Temperature (F)')
plt.title('Southern Hemisphere - Max Temp vs. Latitude Linear Regression')
print(f"The r-squared is: {rvalue}")
print("The regression displays a positive correlation.")
print("Strong relationship with Temp and Latitude: Temperature increases the further south (+lat) you go toward the equator.")
plt.show()
# #### Northern Hemisphere - Humidity (%) vs. Latitude Linear Regression
x_values = pd.to_numeric(northern_df['Lat']).astype(float)
y_values = pd.to_numeric(northern_df['Humidity']).astype(float)
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
print(f"Regression line equation is: {line_eq}")
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('Latitude')
plt.ylabel('Humidity (%)')
plt.title('Northern Hemisphere - Humidity (%) vs. Latitude Linear Regression')
print(f"The r-squared is: {rvalue}")
print("This regression suggests a weak correlation between the Latitude and Humidity.")
plt.show()
# #### Southern Hemisphere - Humidity (%) vs. Latitude Linear Regression
x_values = pd.to_numeric(southern_df['Lat']).astype(float)
y_values = pd.to_numeric(southern_df['Humidity']).astype(float)
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
print(f"Regression line equation is: {line_eq}")
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('Latitude')
plt.ylabel('Humidity (%)')
plt.title('Southern Hemisphere - Humidity (%) vs. Latitude Linear Regression')
print(f"The r-squared is: {rvalue}")
print("This regression suggests a weak correlation between the Latitude and Humidity.")
plt.show()
# #### Northern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression
x_values = pd.to_numeric(northern_df['Lat']).astype(float)
y_values = pd.to_numeric(northern_df['Cloudiness']).astype(float)
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
print(f"Regression line equation is: {line_eq}")
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('Latitude')
plt.ylabel('Cloudiness (%)')
plt.title('Northern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression')
print(f"The r-squared is: {rvalue}")
print("This regression suggests a weak correlation between the Latitude and Cloudiness.")
plt.show()
# #### Southern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression
x_values = pd.to_numeric(southern_df['Lat']).astype(float)
y_values = pd.to_numeric(southern_df['Cloudiness']).astype(float)
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
print(f"Regression line equation is: {line_eq}")
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('Latitude')
plt.ylabel('Cloudiness (%)')
plt.title('Southern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression')
print(f"The r-squared is: {rvalue}")
print("This regression suggests a weak correlation between the Latitude and Cloudiness.")
plt.show()
# #### Northern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression
x_values = pd.to_numeric(northern_df['Lat']).astype(float)
y_values = pd.to_numeric(northern_df['Wind Speed']).astype(float)
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
print(f"Regression line equation is: {line_eq}")
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('Latitude')
plt.ylabel('Wind Speed (mph)')
plt.title('Northern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression')
print(f"The r-squared is: {rvalue}")
print("This regression suggests a weak correlation between the Latitude and Wind Speed.")
plt.show()
# #### Southern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression
x_values = pd.to_numeric(southern_df['Lat']).astype(float)
y_values = pd.to_numeric(southern_df['Wind Speed']).astype(float)
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
print(f"Regression line equation is: {line_eq}")
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('Latitude')
plt.ylabel('Wind Speed (mph)')
plt.title('Southern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression')
print(f"The r-squared is: {rvalue}")
print("This regression suggests a weak correlation between the Latitude and Wind Speed.")
plt.show()
| WeatherPy/WeatherPy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
flag = 0
count = 0
filename ='classroom_unit2000.ply'
filename = filename[:-4]
with open(filename + '.ply','r') as input_file:
with open(filename + '_pc.ply','a+') as output_file:
lines = input_file.readlines()
for line in lines:
count += 1
if 'end_header' in line:
flag = 1
output_file.write(line)
continue
if flag == 0:
if ('red' in line) or ('green' in line) or ('blue' in line):
continue
else:
output_file.write(line)
continue
if count % 2 == 0:
output_file.write(line)
# -
| demo_files/removeRGB.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
pip install instaloader
import instaloader
# +
ig = instaloader.Instaloader()
dp = input("Enter Insta Username:")
ig.download_profile(dp, profile_pic_only = True)
# -
| igphotoprofile.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import warnings
warnings.filterwarnings('ignore')
# # Prepare the Dataset
dataset_name = 'MNIST'
# +
import numpy as np
from keras.datasets import mnist
(X, y), (X_test, y_test) = mnist.load_data()
X = np.concatenate((X, X_test))
y = np.concatenate((y, y_test))
imgs = X
del X_test
del y_test
print('Dataset size {}'.format(X.shape))
# -
# # BayesianAE CNN
# %load_ext autoreload
# %autoreload 2
from utils.constants import Models as models
from models.AE import AE
ae = AE(model_type=models.BayAE, dataset_name=dataset_name,hidden_dim=500, plot=True, isConv=True)
ae.fit(X,y)
from utils.plots import plot_samples, merge
from skimage.transform import resize
import matplotlib.pyplot as plt
for _ in range(5):
samples = ae.reconst_samples_out_data()
scale = 10
im = merge(samples, (10,10))
fig_width = int(im.shape[0] * scale)
fig_height = int(im.shape[1] * scale)
im = resize(im, (fig_width, fig_height), anti_aliasing=True)
plt.figure(dpi=150)
plt.imshow(im)
plt.axis('off')
| MNIST-BayesianAE-CNN.ipynb |
# ---
# layout : jupyter
# title : 파이썬 - 제너레이터(Generator)에 대하여
# category : Code Snippet
# tags : python
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# ***
# [http://schoolofweb.net/blog/posts/파이썬-제너레이터-generator/](http://schoolofweb.net/blog/posts/파이썬-제너레이터-generator/) 를 보고 정리하였습니다.
#
# ***
# + [markdown] pycharm={"name": "#%% md\n"}
# 제너레이터란, 반복자(iterator)와 같은 루프의 작용을 컨트롤하기 위해 쓰여지는 특별한 함수 또는 루틴을 말합니다.
# 제너레이터는 배열이나 리스트를 리턴하는 함수와 비슷하지만 차이점은 한번에 모든 값을 만들어서 배열이나 리스트에 담은 다음 리턴하는 것이 아니라, yield구문을 이용해 한번 호출될 때마다 하나의 값만을 리턴합니다.
#
# 이러한 이유로 일반 반복자에 비해 적은 메모리를 필요로 합니다.
#
# 일반함수 같은 경우 코드의 첫줄부터 마지막줄 return, exception등을 만날때까지 실행된 후 호출자에게 리턴됩니다. 그리고 함수 내부의 모든 로컬 변수는 메모리에서 사라집니다.
#
# 그러나, 영원히 사라져버리는 함수가 아니라 하나의 일을 마치면 자기가 했던 일을 기억하고 대기했다가 다시 호출되면 전의 일을 이어서 하는 함수가 필요하기 시작했습니다. 그것이 바로 **제너레이터**입니다.
# + pycharm={"name": "#%%\n"}
def square_numbers(nums):
result = []
for i in nums:
result.append(i+1)
return result
my_nums = square_numbers([1,2,3,4,5])
print(my_nums)
# + [markdown] pycharm={"name": "#%% md\n"}
# 위의 코드를 generator로 만들면 아래와 같습니다.
#
#
# + pycharm={"name": "#%%\n"}
def square_numbers(nums):
for i in nums:
yield i*1
my_nums = square_numbers([1,2,3,4,5])
print(my_nums)
# + [markdown] pycharm={"name": "#%% md\n"}
# 결과를 보면 generator라는 오브젝트가 리턴됩니다. 자신이 리턴할 모든 값을 메모리에 저장하지 않기 때문입니다. 따라서 제너레이터가 만들어졌고, 이 제너레이터를 호출할 때마다 하나의 값만을 차례로 계산해서 리턴합니다.
# 더이상 전달할 값이 없으면 stopiteration이 발생합니다.
# + pycharm={"name": "#%%\n"}
print(next(my_nums))
print(next(my_nums))
print(next(my_nums))
print(next(my_nums))
print(next(my_nums))
# + [markdown] pycharm={"name": "#%% md\n"}
# next메서드 말고 for문을 통해서도 호출이 가능합니다.
#
# + pycharm={"name": "#%%\n"}
def square_numbers(nums):
for i in nums:
yield i*i
mynums = square_numbers([1,2,3,4,5])
for num in mynums:
print(num)
# + [markdown] pycharm={"name": "#%% md\n"}
# 아래와 같이 축약형태로 generator를 만들 수도 있습니다.
# + pycharm={"name": "#%%\n"}
mynums = (x*x for x in [1,2,3,4,5])
for num in mynums:
print(num)
# + [markdown] pycharm={"name": "#%% md\n"}
# 아래는 generator 내 데이터를 한번에 보고 싶을 때 입니다.
# 그런데 아래와 같이 사용하면 generator의 메모리 절약 장점이 사라지게 됩니다. 주의하기!!
# + pycharm={"name": "#%%\n"}
mynums = (x*x for x in [1,2,3,4,5])
print(mynums)
print(list(mynums))
# + [markdown] pycharm={"name": "#%% md\n"}
# 메모리 장점과 시간절약의 장점이 있다는 것을 아래 코드를 통해 확인해보겠습니다.
#
#
# + pycharm={"name": "#%%\n"}
from __future__ import division
import os
import psutil
import random
import time
names = ['최용호', '지길정', '진영욱', '김세훈', '오세훈', '김민우']
majors = ['컴퓨터 공학', '국문학', '영문학', '수학', '정치']
process = psutil.Process(os.getpid())
mem_before = process.memory_info().rss / 1024 / 1024
def people_list(num_people):
result = []
for i in range(num_people):
person = {
'id': i,
'name': random.choice(names),
'major': random.choice(majors)
}
result.append(person)
return result
def people_generator(num_people):
for i in range(num_people):
person = {
'id': i,
'name': random.choice(names),
'major': random.choice(majors)
}
yield person
t1 = time.clock()
people_list = people_list(1000000)
mem_after = process.memory_info().rss / 1024 / 1024
t2 = time.clock()
total_time = t2-t1
print('시작 전 메모리 사용량: {} MB'.format(mem_before))
print('종료 후 메모리 사용량: {} MB'.format(mem_after))
print('총 소요된 시간: {:.6f} 초'.format(total_time))
# + pycharm={"name": "#%%\n"}
mem_before = process.memory_info().rss / 1024 / 1024
t11 = time.clock()
people = people_generator(1000000)
mem_after2 = process.memory_info().rss / 1024 / 1024
t12 = time.clock()
total_time2 =t12 - t11
print('시작 전 메모리 사용량: {} MB'.format(mem_before))
print('종료 후 메모리 사용량: {} MB'.format(mem_after2))
print('총 소요된 시간: {:.6f} 초'.format(total_time2))
# + pycharm={"name": "#%%\n"}
| _ipynb/2021-01-21-python-generator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sweetpand/Algorithms/blob/master/sliding.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="m1bMItSIpmkf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9bb9d6b1-fad0-4a2b-b377-dad4d338e29b"
def solution(arr):
res=count = 0
flag = False
for i in arr:
if i == 'R':
flag = True
res += count
count = 0
else:
if flag:
count += 1
return res
print(solution('WRRWWR'))
# + id="wyKid8vSr9kD" colab_type="code" colab={}
from typing import List
# + id="KOoT1bhpsEzz" colab_type="code" colab={}
nums = [1,3,-1,-3,5,3,6,7]
# + id="Dv37yIbosS2m" colab_type="code" colab={}
k = 3
# + id="acPHYsWTrO5C" colab_type="code" colab={}
def maxSlidingWindow(self, nums: List[int], k: int) -> List[int]:
queue = collections.deque()
res = []
for i in range(len(nums)):
if i >= k and i - k == queue[0]:
queue.popleft()
while queue and nums[i] > nums[queue[-1]]:
queue.pop()
queue.append(i)
if i >= k - 1:
res.append(nums[queue[0]])
return res
| sliding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# +
import pickle
import jax
import jax.numpy as jnp
import timecast as tc
import tqdm
# -
# # Download all data
# !mkdir -p data
# !gsutil -m cp -r gs://skgaip/data/flood data
basins = pickle.load(open("data/flood/meta.pkl", "rb"))["basins"]
basin_to_yhats_LSTM = pickle.load(open("data/flood/tigerforecast/lstm.pkl", "rb"))
# # Define optimizers
class SGD:
def __init__(self,
loss_fn=lambda pred, true: jnp.square(pred - true).mean(),
learning_rate=0.0001,
project_threshold={}):
self.loss_fn = loss_fn
self.learning_rate = learning_rate
self.project_threshold = project_threshold
def update(self, module, params, x, y):
grad = jax.jit(jax.grad(lambda module, x, y: self.loss_fn(module(x), y)))(module, x, y)
new_params = {k:w - self.learning_rate * grad.params[k] for (k, w) in params.items()}
for k, param in new_params.items():
norm = jnp.linalg.norm(new_params[k])
new_params[k] = jax.lax.cond(norm > self.project_threshold[k],
new_params[k],
lambda x : (self.project_threshold[k]/norm) * x,
new_params[k],
lambda x : x)
return new_params
class MultiplicativeWeights:
def __init__(self, eta=0.008):
self.eta = eta
self.grad = jax.jit(jax.grad(lambda W, preds, y: jnp.square(jnp.dot(W, preds) - y).sum()))
def update(self, module, params, x, y):
grad = self.grad(params, x, y)
new_params = params * jnp.exp(-1 * self.eta * grad)
return new_params / new_params.sum()
# # Define modules
class AR(tc.Module):
def __init__(self, input_dim=32, output_dim=1, history_len=270):
self.kernel = jnp.zeros((history_len, input_dim, output_dim))
self.bias = jnp.zeros((output_dim, 1))
self.T = 0
def __call__(self, x):
self.T += 1
return jnp.tensordot(self.kernel, x, ([0,1],[0,1])) + self.bias
class GradientBoosting(tc.Module):
def __init__(self, N, input_dim=32, output_dim=1, history_len=270):
for i in range(N):
self.add_module(AR(input_dim=input_dim, output_dim=output_dim, history_len=history_len))
self.W = jnp.ones(N) / N
def __call__(self, x):
pred, preds = 0, []
for i, (name, submodule) in enumerate(self.modules.items()):
pred_i = submodule(x).squeeze()
preds.append(pred_i)
pred += self.W[i] * pred_i
return preds
# # Initialize optimizers
# +
bias_threshold = 1e-4
eta = 0.008
SGDs = [SGD(
learning_rate=learning_rate,
project_threshold={
"kernel": kernel_threshold,
"bias": bias_threshold
})
for kernel_threshold, learning_rate in [
(0.03, 2e-5),
(0.05, 2e-5),
(0.07, 2e-5),
(0.09, 2e-5),
]]
MW = MultiplicativeWeights(eta=eta)
# -
# # Predict basin
# +
def predict(basin):
N = len(SGDs)
model = GradientBoosting(N)
Y_LSTM = jnp.array(basin_to_yhats_LSTM[basin])
X = pickle.load(open("data/flood/test/{}.pkl".format(basin), "rb"))
Y = pickle.load(open("data/flood/qobs/{}.pkl".format(basin), "rb"))
def loop(model, xy):
x, y = xy
preds = jnp.asarray(model(x))
pred = 0
for i, (name, module) in enumerate(model.modules.items()):
module.params = SGDs[i].update(module, module.params, x, y - pred)
pred += model.W[i] * preds[i]
model.W = MW.update(model, model.W, preds, y)
return model, pred
Y_RESID = Y - Y_LSTM
model, Y_BOOST = jax.lax.scan(loop, model, (X, Y_RESID))
# for x, y in zip(X, Y_RESID):
# model, y_hat = loop(model, (x, y))
Y_BOOST = jnp.asarray(Y_BOOST).squeeze()
loss = ((Y - (Y_LSTM + Y_BOOST)) ** 2).mean()
return loss
# -
# # Run!
losses = {}
for basin in tqdm.tqdm(basins):
losses[basin] = predict(basin)
| notebooks/2020-07-12 flood in timecast.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
import numpy as np
from scipy.optimize import minimize
def rosen(x):
#"""The Rosenbrock function"""
return sum(100.0*(x[1:]-x[:-1]**2.0)**2.0 + (1-x[:-1])**2.0)
print(sys.argv)
x = np.array([float(i) for i in sys.argv[3:]]) #[float(i) for i in sys.argv[3:]]
x = np.array([1.1,2.2,3.3]) #[float(i) for i in sys.argv[3:]]
f = open('obj.txt', 'w')
#print('rosen '+ str(rosen(x)))
f.write('rosen '+ str(rosen(x)))
# -
# # rosenbrock_demo.py
# #
# #Python 2.7.3
# #Matplotlib 1.1.1rc
#
# #Code also works fine with Anaconda3 / matplotlib version 1.4.0!
# #Code also works fine with CPython 3.4.2 + Scipy-stack-14.8.27.win32-py3.4 from
# # <NAME>'s unofficial libraries:
# # http://www.lfd.uci.edu/~gohlke/pythonlibs/
#
# #Works fine with Python3 on Ubuntu 14.04 after adding SciPy stack:
# # sudo apt-get install python3-numpy python3-scipy python3-matplotlib
#
#
#
# #This function is not used, but illustrates the Rosenbrock function with
# # 2 parameters. The actual Rosenbrock function is embedded in the code below
# # using array operations so that it is calculated over a meshgrid of many
# # points to produce an array of many Z values.
# #Rosenbrock function of 2 variables:
# def rosenbrock(x,y):
# return (1-x)**2 + 100* ((y-x**2))**2
#
#
# from mpl_toolkits.mplot3d import Axes3D
# from matplotlib import cm
# from matplotlib.ticker import LinearLocator, FormatStrFormatter
# import matplotlib.pyplot as plot
# import numpy as np
#
# fig = plot.figure()
# ax = fig.gca(projection='3d')
#
# s = 0.05 # Try s=1, 0.25, 0.1, or 0.05
# X = np.arange(-2, 2.+s, s) #Could use linspace instead if dividing
# Y = np.arange(-2, 3.+s, s) #evenly instead of stepping...
#
# #Create the mesh grid(s) for all X/Y combos.
# X, Y = np.meshgrid(X, Y)
#
# #Rosenbrock function w/ two parameters using numpy Arrays
# Z = (1.-X)**2 + 100.*(Y-X*X)**2
#
# surf = ax.plot_surface(X, Y, Z, rstride=1, cstride=1, cmap=cm.coolwarm,
# linewidth=0, antialiased=False) #Try coolwarm vs jet
#
#
# ax.zaxis.set_major_locator(LinearLocator(10))
# ax.zaxis.set_major_formatter(FormatStrFormatter('%.02f'))
#
# fig.colorbar(surf, shrink=0.5, aspect=5)
#
# #Displays the figure, handles user interface, returns when user closes window
# plot.show()
#
| others/RosenBrock.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp core
# -
# # core
#
# > core stuff.
# hide
from nbdev.showdoc import *
# +
# export
from apiclient.discovery import build
from oauth2client.service_account import ServiceAccountCredentials
import os
from dotenv import load_dotenv
import pandas as pd
load_dotenv()
SCOPES = ['https://www.googleapis.com/auth/analytics.readonly']
KEY_FILE_LOCATION = os.getenv('GOOGLE_APPLICATION_CREDENTIALS')
if KEY_FILE_LOCATION:
credentials = ServiceAccountCredentials.from_json_keyfile_name(KEY_FILE_LOCATION, SCOPES)
else:
credentials = None
# +
# export
def ga_to_df(start_date: str, end_date: str, metrics: list, dimensions: list, filters: list = None, view_id = None) -> pd.DataFrame:
analytics = build('analyticsreporting', 'v4', credentials=credentials)
if not view_id:
view_id = os.getenv('GOOGLE_ANALYTICS_VIEW_ID')
qry = {
'dateRanges': [{'startDate': start_date, 'endDate': end_date}],
'metrics': [{'expression': m} for m in metrics],
'dimensions': [{'name': m} for m in dimensions],
}
if filters:
qry['dimensionFilterClauses'] = [
{'operator': f[0], 'filters': [{'dimensionName': f[1][0], 'operator': f[1][1], 'expressions': f[1][2]}]}
for f in filters
]
qry['viewId'] = view_id
request_list = [qry]
response = analytics.reports().batchGet(body={'reportRequests': request_list}).execute()
data = []
for report in response.get('reports'):
dimensions = report.get('columnHeader').get('dimensions')
metrics = [x['name'] for x in report.get('columnHeader').get('metricHeader').get('metricHeaderEntries')]
for row in report.get('data').get('rows'):
dimensions_data = row.get('dimensions', [])
metrics_data = [x['values'] for x in row.get('metrics', [])][0]
metrics_data = [float(x) for x in metrics_data]
colnames = dimensions + metrics
data_row = dimensions_data + metrics_data
data.append(data_row)
df = pd.DataFrame(data, columns=colnames)
return df
# +
# test
# make a example query
start_date = '7daysAgo'
end_date = 'today'
metrics = ['ga:users', 'ga:sessions', 'ga:pageviews']
dimensions = ['ga:date', 'ga:country']
filters = [['AND', ['ga:country', 'IN_LIST', ['United States', 'Ireland']]]]
df = ga_to_df(start_date, end_date, metrics, dimensions, filters)
#print(df)
# check have realistic looking data
assert df['ga:sessions'].sum() >= 0
assert df['ga:pageviews'].sum() >= 0
assert df['ga:pageviews'].sum() >= df['ga:sessions'].sum()
# -
| 00_core.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## PageRank notes ##
#
# Based on your many questions and puzzled looks, we've put together this set of notes on the PageRank algorithm, as we presented it in class.
#
# If these notes still don't make sense, you may also find the following references to be useful:
# * Cleve Moler's notes on PageRank -- Moler's explanation is similar to what we discussed in class and what appears in these notes, but they are _not_ identical as these notes as explained below: [link](https://www.mathworks.com/moler/exm/chapters/pagerank.pdf)
# * The original PageRank paper by Page et al.: [link](http://ilpubs.stanford.edu:8090/422/)
# ### High-level Overview ###
#
# **Motivating problem.** The original PageRank algorithm addresses the following problem. Imagine a user who is looking for information on the web, having only some search terms, e.g., "`free donuts.`" There may be many pages that match these terms; in what order should the search engine return matching pages to the user?
# **A search framework.** One idea is to adopt a two-phase framework for search. One phase is offline, meaning it happens _before_ any searches take place. The other phase is online, meaning it happens _at search time_, when the user's search terms are known.
#
# The offline phase _precomputes_ a global ranking of all web pages, given some snapshot of the web. The online phase uses the search terms to filter the ranked list, and it returns those pages in the global rank order.
# **The high-level PageRank idea.** The PageRank algorithm is the offline phase. It is based on a probabilistic model of how a hypothetical user might surf the web, in the absence of any specific query.
#
# Here is a somewhat informal description of the model; the details follow in the next section.
#
# Suppose there are $n$ web pages, represented by a _vertex set_ $V = \{1, 2, \ldots, n\}$. The pages link to one another; let $E$ denote the _edge set_, which is a set of pairs $(i, j)$ indicating that page $i$ points to page $j$. We will sometimes write this link relationship as $i \rightarrow j$. This representation is also known as a [_directed graph_](https://en.wikipedia.org/wiki/Directed_graph) representation.
#
# Next, consider a "random web surfer." This surfer visits web pages one at a time, according to the following stochastic process.
#
# 1. At each time step $t \geq 0$, the surfer visits a page. Further assume that the surfer's choice of page at time $t+1$ depends only on the page visited at time $t$. (This assumption makes this process a [_discrete-time Markov chain_](https://en.wikipedia.org/wiki/Markov_chain).)
# 2. Initially, at time $t=0$, the web surfer starts on a random page.
# 3. Suppose the surfer is on page $i$ at time $t$. With probability $\alpha$, the surfer will decide to follow a link going from $i$ to some new page $j$.
# 4. At time $t$, the surfer might instead decide, with probability $1-\alpha$, to _jump_ to some page $j$. Page $j$ is _not necessarily_ directly connected to $i$.
#
# As time proceeds, the surfer jumps from page to page, sometimes hitting a page it has already visited, and sometimes jumping to an entirely different part of the web graph. Now imagine that the surfer surfs for an _infinitely long time_; what is the probability that the surfer will be on any given page? If this probability distribution can be calculated, then the PageRank idea is to use the distribution to rank the web pages: the most likely page for the surfer to land on is the top-ranked page, the next most likely page is the second-ranked page, and so on.
# ### Mathematical Details ###
#
# To fully specify the process outlined above, we need to pin down a few more details. We will do so using probabilistic statements, which it then turns out we will be able to write down succinctly in the language of linear algebra.
# **Connectivity matrix.** Let's start by representing the graph by a matrix $G \equiv (g_{ij})$, where the $(i, j)$ entry, $g_{ij}$, is 1 if there is a link $i \rightarrow j$, and 0 otherwise.
# **Probability state vector.** Next, let $x(t)$ be a (column) vector that represents the probabilities that the surfer is on each page at time $t$. That is,
#
# $x(t) \equiv \left(\begin{array}{c} x_1(t) \\ x_2(t) \\ \vdots \\ x_n(t) \end{array}\right)$,
#
# where $x_i(t)$ is the probability that the surfer is on page $i$ at time $t$. Since the surfer must always be on some page, these probabilities must sum to 1: $\sum_{i=1}^{n} x_i(t) = 1$.
# **Surfing process.** At time $t=0$, suppose that the surfer is equally likely to start on any page. Then, $x_i(0) = \frac{1}{n}$.
#
# Now suppose that the surfer is on page $i$ at time $t$. What page will the surfer visit at time $t+1$? Recall that there are two scenarios in our high-level model: follow an out-link or jump to another page.
#
# _Scenario 1._ If the surfer decides to follow an out-link, which one will it choose?
#
# Let's assume the surfer picks an outgoing link uniformly at random. That is, if page $i$ has $d_i$ outgoing links, then let the probability of choosing an out-link be $\frac{1}{d_i}$. The value $d_i$ is also called the _out-degree_ of vertex $i$. It may be computed by summing each row of $G$, i.e., $d_i \equiv \sum_{j=1}^{n} g_{ij}$.
#
# Thus, given the decision to follow an out-link starting from page $i$, the probability of moving to page $j$ is $p_{ij} \equiv \frac{g_{ij}}{d_i}$.
#
# What if page $i$ _has no_ outgoing edges? There are several ways to handle this case. The simple one we will adopt is to _force_ it to have a self-edge, $i \rightarrow i$. In other words, the surfer has decided to follow an out-link but has nowhere to go; therefore, it stays put on page $i$. Mathematically, the self-edge means $d_i = g_{ii} = 1$ and $p_{ii} = 1$.
#
# > Aside: This way of handling pages without outgoing edges differs from the way they are treated both in the original PageRank scheme and in Moler's notes. The original PageRank scheme simply removed these pages. By contrast, Moler assumes that when there is no outgoing edge, then the surfer jumps to any random page, just like the $1-\alpha$ case. In other words, Moler would set all $g_{ij} = 1$ for all $j$ when $i$ has no outgoing links.
#
# Given all of the $g_{ij}$, including self-edges when needed, we can express all of these quantities in matrix notation:
#
# $G \equiv \left( g_{ij} \right),
# \qquad D \equiv \left(\begin{array}{ccc} d_1 && \\ & \ddots & \\ && d_n \end{array}\right),
# \qquad P \equiv \left( p_{ij} \right) = D^{-1}G$
#
# The matrix $P$ is sometimes called a [probability transition matrix](https://en.wikipedia.org/wiki/Stochastic_matrix). From the definitions above, you should convince yourself that every row of the matrix $P \equiv (p_{ij})$ sums to 1.
#
# _Scenario 2._ If instead the surfer decides to jump to a random page, which one will it choose?
#
# Again, let's assume the surfer jumps uniformly at random to any one of the $n$ pages, which includes itself and any outgoing links from the current page. Then, the probability of choosing any other page $j$ would be just $\frac{1}{n}$.
# **Putting it all together.** We now have all the details we need to compute the probability of ending up on a page $i$ at time $t+1$ starting from some page $j$ at time $t$. This probability is, as a scalar formula,
#
# $x_i(t+1) = \left[\alpha \cdot \sum_{j=1}^{n} p_{ji} x_j(t)\right] + \left[(1-\alpha) \cdot \frac{1}{n}\right].$
#
# We can also write this more compactly in matrix notation. Let $u$ be a vector whose entries are all equal to 1. Then the above formula for all $i$ is the same as,
#
# $x(t+1) = \alpha P^T x(t) + \frac{1 - \alpha}{n} u$.
#
# From the definition of $P$, note that $P^T = G^T D^{-1}$, which is the notation we used in class and which we wrote some code to compute.
#
# Thus, the PageRank algorithm is the following: run the preceding iteration, starting at $t=0$, until $x(t)$ appears to stabilize (i.e., has reached steady-state) or, as is typically done, until some maximum number of time steps has been reached.
# ### Convergence? ###
#
# At least one detail remains: how do we know that the state vector $x(t)$ will _ever_ reach a steady-state? Could the probabilities oscillate, diverge, or exhibit chaotic behavior?
#
# The analysis to prove that a steady-state exists is somewhat involved, but the gist is the following. The formula to compute $x(t+1)$ from $x(t)$ can also be written as follows:
#
# $x(t+1) = \hat{A} x(t)$,
#
# where
#
# $\hat{A} \equiv \alpha P^T + \frac{1-\alpha}{n} uu^T$. (You can convince yourself of this fact by first observing that $u^Tx(t)=1$.)
#
# Thus, when we ask whether this process reaches steady-state, then we are effectively asking whether there is a vector $x$ such that $x = \hat{A} x$.
#
# Like $P^T$ itself, the matrix $\hat{A}$ has the following properties, which you can verify:
#
# * Its entries all lie between 0 and 1.
# * The columns sum to 1.
#
# From these facts, one may apply a theorem from linear algebra called the _Perron-Frobenius theorem_ and conclude that $x = \hat{A} x$ has a solution that is _both_ non-zero _and_ unique to within some scaling factor. (See the notes by Moler.)
| pagerank-notes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Model Calibration
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import sklearn
plt.rcParams['font.size'] = 16
plt.rcParams['figure.figsize'] = [12, 8]
plt.rcParams['savefig.bbox'] = 'tight'
plt.rcParams["savefig.dpi"] = 300
sklearn.set_config(display='diagram')
# -
def plot_calibration_curve(y_true, y_prob, n_bins=5, ax=None, hist=True, normalize=False):
prob_true, prob_pred = calibration_curve(y_true, y_prob, n_bins=n_bins, normalize=normalize)
if ax is None:
ax = plt.gca()
if hist:
ax.hist(y_prob, weights=np.ones_like(y_prob) / len(y_prob), alpha=.4,
bins=np.maximum(10, n_bins))
ax.plot([0, 1], [0, 1], ':', c='k')
curve = ax.plot(prob_pred, prob_true, marker="o")
ax.set_xlabel("predicted probability")
ax.set_ylabel("fraction of positive samples")
ax.set(aspect='equal')
return curve
# ## Create dummy dataset
# +
from sklearn.datasets import make_classification
from sklearn.model_selection import train_test_split
X, y = make_classification(n_samples=10000, n_features=20,
n_informative=2, n_redundant=2)
train_samples = 100 # Samples used for training the models
X_train = X[:train_samples]
X_test = X[train_samples:]
y_train = y[:train_samples]
y_test = y[train_samples:]
# -
# ### Train linear model
# +
from sklearn.linear_model import LogisticRegression
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
lr = make_pipeline(StandardScaler(), LogisticRegression(random_state=42))
lr.fit(X_train, y_train)
# -
# ## Calibration curve
from sklearn.calibration import calibration_curve
lr_proba = lr.predict_proba(X_test)
# +
prob_true, prod_pred = calibration_curve(y_test, lr_proba[:, 1], n_bins=5)
print(prob_true)
print(prod_pred)
# -
plot_calibration_curve(y_test, lr_proba[:, 1]);
from sklearn.metrics import brier_score_loss
lr_brier = brier_score_loss(y_test, lr_proba[:, 1])
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(20, 8))
plot_calibration_curve(y_test, lr_proba[:, 1], n_bins=5, ax=ax1)
ax1.set_title("n_bins=5")
plot_calibration_curve(y_test, lr_proba[:, 1], n_bins=10, ax=ax2)
ax2.set_title("n_bins=10")
plot_calibration_curve(y_test, lr_proba[:, 1], n_bins=30, ax=ax3)
ax3.set_title("n_bins=30")
# ### Train Random Forest
from sklearn.ensemble import RandomForestClassifier
rf = RandomForestClassifier(random_state=0)
rf.fit(X_train, y_train)
rf_proba = rf.predict_proba(X_test)
rf_brier = brier_score_loss(y_test, rf_proba[:, 1])
rf_brier
# ### Train Single Tree
from sklearn.tree import DecisionTreeClassifier
tree = DecisionTreeClassifier(random_state=0)
tree.fit(X_train, y_train)
tree_proba = tree.predict_proba(X_test)
tree_brier = brier_score_loss(y_test, tree_proba[:, 1])
tree_brier
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(20, 8))
plot_calibration_curve(y_test, lr_proba[:, 1], n_bins=10, ax=ax1)
ax1.set_title(f"LogisticRegression: {lr_brier:0.4f}")
plot_calibration_curve(y_test, tree_proba[:, 1], n_bins=10, ax=ax2)
ax2.set_title(f"DecisionTreeClassifier: {tree_brier:0.4f}")
plot_calibration_curve(y_test, rf_proba[:, 1], n_bins=10, ax=ax3)
ax3.set_title(f"RandomForestClassifier: {rf_brier:0.4f}");
# ## Exercise 1
#
# 1. Train a `sklearn.naive_bayes.GaussianNB` on the training set.
# 2. Compute the brier score loss on the test set for the `GuassianNB`.
# 3. Plot the calibration curve with `n_bins=10`.
# +
# # %load solutions/02-ex01-solutions.py
# -
# ## Calibration
from sklearn.calibration import CalibratedClassifierCV
rf = RandomForestClassifier(random_state=0)
cal_rf = CalibratedClassifierCV(rf, method="isotonic")
cal_rf.fit(X_train, y_train)
cal_rf_proba = cal_rf.predict_proba(X_test)
cal_rf_brier = brier_score_loss(y_test, cal_rf_proba[:, 1])
fig, (ax1, ax2) = plt.subplots(1, 2)
plot_calibration_curve(y_test, rf_proba[:, 1], ax=ax1, n_bins=10)
ax1.set_title(f"forest no calibration: {rf_brier:0.4f}")
plot_calibration_curve(y_test, cal_rf_proba[:, 1], ax=ax2, n_bins=10)
ax2.set_title(f"calibrated: {cal_rf_brier:0.4f}");
# ### Calibrating the linear model
lr = make_pipeline(StandardScaler(), LogisticRegression(random_state=42))
cal_lr = CalibratedClassifierCV(lr, method='isotonic')
cal_lr.fit(X_train, y_train)
# +
cal_lr_proba = cal_lr.predict_proba(X_test)
cal_lr_brier = brier_score_loss(y_test, cal_lr_proba[:, 1])
# -
fig, (ax1, ax2) = plt.subplots(1, 2)
plot_calibration_curve(y_test, lr_proba[:, 1], ax=ax1, n_bins=10)
ax1.set_title(f"no calibration: {lr_brier:0.4f}")
plot_calibration_curve(y_test, cal_lr_proba[:, 1], ax=ax2, n_bins=10)
ax2.set_title(f"calibrated: {cal_lr_brier:0.4f}");
# ## Exercise 2
#
# 0. Finish Exercise 1 for training an uncalibrated `GaussianNB`.
# 1. Calibrate the `sklearn.naive_bayes.GaussianNB` on the training set.
# 2. Compute the brier score loss on the test set.
# 3. Plot the calibration curve with `n_bins=10`.
# 4. Did the calibration improve with `CalibratedClassifierCV`?
# +
# # %load solutions/02-ex02-solutions.py
| notebooks/02-model-calibration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
"""Python WPS execute"""
from owslib.wps import WebProcessingService, monitorExecution
from os import system
import owslib
owslib.__version__ # version must be >=0.10.3
#wps = WebProcessingService(url="http://birdhouse-lsce.extra.cea.fr:8093/wps", verbose=False)
wps = WebProcessingService(url="http://localhost:8093/wps", verbose=False)
print wps.identification.title
for process in wps.processes:
print '%s : \t %s' % (process.identifier, process.abstract)
# +
files = []
for i in range(1,16):
files.append('file:///home/estimr2/EUCLEIA/indices/RX5day/DJF/RX5day_DJF_HadGEM3-A-N216_historical_r1i1p%s_19600101-20131230.nc' % (i))
# -
files
# +
from os.path import join
execute = wps.execute(
identifier="robustness",
inputs=[
("resource",files[0]),
("resource",files[1]),
("resource",files[2]),
("resource",files[3]),
("resource",files[4]),
("resource",files[5]),
("resource",files[6]),
("resource",files[7]),
("resource",files[8]),
("resource",files[9]),
("resource",files[10]),
("resource",files[11]),
("resource",files[12]),
("resource",files[13]),
("resource",files[14])
])
monitorExecution(execute, sleepSecs=5)
print execute.getStatus()
for o in execute.processOutputs:
print o.reference
# +
graphic = execute.processOutputs[0].reference
from IPython.display import Image
from IPython.core.display import HTML
Image(url= graphic )
# -
| notebooks/WPS_ensembleRobustness.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Uncertainty Calibration
#
# Posterior uncertainty of the probabilistic linear solver can be calibrated by building a regression model for the Rayleigh coefficient $$\ln R(A, s) = \ln \frac{s^\top A s}{s^\top s}$$ at convergence after $k \ll n$ iterations.
# +
# Make inline plots vector graphics instead of raster graphics
# %matplotlib inline
from IPython.display import set_matplotlib_formats
set_matplotlib_formats('pdf', 'svg')
# Matplotlib settings
import matplotlib
import matplotlib.pyplot as plt
matplotlib.rc('font', **{'family': 'serif', 'serif': ['Computer Modern']})
plt.rcParams['text.usetex'] = True
plt.rcParams['text.latex.preamble'] = r'\usepackage{amsfonts} \usepackage{amsmath} \usepackage{bm}'
import warnings
warnings.filterwarnings("ignore")
# -
# ## Test Problem
#
# Generate a kernel matrix from toy data.
# +
import numpy as np
import scipy.linalg
import GPy
# Gram matrix test problem
np.random.seed(0)
# Toy data
n = 1000
x_min, x_max = (-10.0, 10.0)
X = np.random.uniform(x_min, x_max, (n, 1))
Y = np.cos(X ** 2) + np.random.normal(size=(n, 1)) * 0.05
# Kernel
kernel = GPy.kern.Matern32(input_dim=1, variance=1, lengthscale=1)
# Kernel Gram Matrix
K = kernel.K(X=X, X2=X)
sigma = 10 ** -6 * n
A = K + sigma * np.eye(K.shape[0])
# Sample solution
x_true = np.random.normal(size=(n,))
# Right hand side
b = A @ x_true
# Condition number
print(np.linalg.cond(A))
# -
# ## Solution via Probabilistic Linear Solver
#
# Solve a linear system involving the generated kernel matrix using the probabilistic linear solver.
# +
from probnum.linops import Identity, SymmetricKronecker
from probnum.random_variables import Normal
from probnum.linalg import problinsolve
# Callback function to get matrix projections
S = [] # search directions
Y = [] # observations
alpha = [] # step lengths
def callback_fun(xk, Ak, Ainvk, sk, yk, alphak, resid):
S.append(sk)
Y.append(yk)
alpha.append(np.squeeze(alphak))
# Solve with probabilistic linear solver
xhat, Ahat, Ainvhat, info_pls = problinsolve(
A=A, b=b, callback=callback_fun, calibration="gpkern", maxiter=n
)
print(info_pls)
# +
# Transform callback quantities
S = np.squeeze(np.array(S)).T
Y = np.squeeze(np.array(Y)).T
# Compute eigenvalues
eigs = np.real_if_close(np.sort(np.linalg.eigvals(A))[::-1])
# Condition number of problem
np.linalg.cond(A)
# -
# ## Rayleigh Coefficients and Regression Model
#
# Train a GP regression model on the Rayleigh coefficients collected by the algorithm.
# +
# Compute matrix projections
a_proj = np.einsum("nk,nk->k", S, Y) / np.einsum("nk,nk->k", S, S)
iters = np.arange(0, len(a_proj))
# GP mean function via Weyl's result on spectra of Gram matrices: ln(sigma(n)) ~= theta_0 - theta_1 ln(n)
lnmap = GPy.core.Mapping(1, 1)
lnmap.f = lambda n: np.log(n + 10 ** -16)
lnmap.update_gradients = lambda a, b: None
mf = GPy.mappings.Additive(
GPy.mappings.Constant(1, 1, value=0),
GPy.mappings.Compound(lnmap, GPy.mappings.Linear(1, 1)),
)
# Log-Rayleigh quotient modelled with a Gaussian process
k = GPy.kern.RBF(input_dim=1, lengthscale=1, variance=1)
# -
print(mf)
m = GPy.models.GPRegression(
iters[:, None] + 1, np.log(a_proj)[:, None], kernel=k, mean_function=mf
)
m.optimize(messages=1, ipython_notebook=True)
# Compute scale
logR_pred = m.predict(np.arange(info_pls["iter"] + 1, n + 1)[:, None])[0]
phi = np.exp(np.mean(logR_pred))
psi = np.exp(np.mean(-logR_pred))
print(f"Calibration scales phi = {phi} and psi = {psi}.")
# ## Predict and Plot
# +
# Figure and setup
fig, axes = plt.subplots(nrows=1, ncols=1, figsize=(4, 2), sharex=True, squeeze=False)
pred_iters = np.arange(1, info_pls["iter"] + 41)
# Predict log Rayleigh quotient
GP_pred = m.predict(pred_iters[:, None])
a_pred_GPmean = GP_pred[0].ravel() # + beta0
a_pred_GP2std = 2 * np.sqrt(GP_pred[1].ravel())
# Plot
axes[0, 0].axhline(
y=np.max(eigs),
color="gray",
alpha=0.5,
linestyle="--",
label="$[\\lambda_{\\min}(\\bm{A}), \\lambda_{\\max}(\\bm{A})]$",
)
axes[0, 0].axhline(y=np.min(eigs), color="gray", alpha=0.5, linestyle="--")
axes[0, 0].fill_between(
x=pred_iters - 1,
y1=np.exp(a_pred_GPmean - a_pred_GP2std),
y2=np.exp(a_pred_GPmean + a_pred_GP2std),
alpha=0.3,
)
axes[0, 0].plot(
pred_iters - 1,
np.exp(a_pred_GPmean),
label="GP posterior $p(\\ln R_i \\mid \\bm{Y}, \\bm{S})$",
)
axes[0, 0].plot(iters, a_proj, ".", label="Rayleigh quotient $R(\\bm{A}, \\bm{s}_{i})$")
axes[0, 0].axhline(
y=phi,
xmin=iters[-1] / pred_iters[-1],
xmax=0.955,
label="Uncertainty scale $\\phi=\psi^{-1}$",
color="red",
linestyle="-",
marker="o",
markevery=1.0,
)
axes[0, 0].set_xlabel("iteration $i$")
axes[0, 0].set_yscale("log")
plt.legend(fontsize=10, labelspacing=0.1)
plt.tight_layout(pad=0)
# Save to file
fig.savefig("../../figures/rayleigh_quotient.pdf", bbox_inches="tight", pad_inches=0.0)
# -
print(mf)
# ## Uncertainty Calibration
#
# We measure the effect of the calibration procedure by computing the log-ratio between the numerical uncertainty and the true error in euclidean norm.
# +
import scipy.sparse.linalg
# Solve with probabilistic linear solver
xhat, Ahat, Ainvhat, info_pls = problinsolve(A=A, b=b, maxiter=n, calibration="gpkern")
# -
print(info_pls)
sigma_trace = info_pls["trace_sol_cov"]
print("\sqrt(trace(Sigma)) = " + str(np.sqrt(sigma_trace)))
l2_error = np.linalg.norm(x_true.ravel() - xhat.mean.ravel(), ord=2)
print(f"l2-error = {l2_error}")
print(f"test statistic w = {0.5 * np.log(sigma_trace) - np.log(l2_error)}")
| experiments/notebooks/uncertainty_calibration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="QIsJs0mcEK_h"
# ### Connect to Kaggle to download dataset
# + id="LaYA5u8XC2vM" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 90} outputId="30cc1f9d-fcfc-4081-8b4e-314bab86057c"
from google.colab import files
files.upload()
# + id="xLCqepLDEoif"
# !pip install -q kaggle
# + id="VQYDhSE0EvMO"
# !mkdir -p ~/.kaggle
# !cp kaggle.json ~/.kaggle/
# !chmod 600 ~/.kaggle/kaggle.json
# + [markdown] id="maBC_ngQE50m"
# ### Setup Paths and download dataset
# + colab={"base_uri": "https://localhost:8080/"} id="4Em1RG02E7g1" outputId="3c74c385-a2a7-474f-94fe-9eaa436c4a75"
# !mkdir traffic_sign_dataset
# %cd traffic_sign_dataset
# + colab={"base_uri": "https://localhost:8080/"} id="bqbUgnIRFCot" outputId="8056a53b-ec25-46b7-b1b3-5ae8f92ae475"
# !kaggle datasets list -s gtsrb-german-traffic-sign
# + colab={"base_uri": "https://localhost:8080/"} id="4RNK3T2pFNeF" outputId="823fba90-595e-4394-9180-02de469a8517"
# !kaggle datasets download meowmeowmeowmeowmeow/gtsrb-german-traffic-sign
# %cd ..
# + colab={"base_uri": "https://localhost:8080/"} id="4zoVD1YbFTk7" outputId="92e60ec9-05d7-4486-b29e-40df07ba0153"
# !unzip traffic_sign_dataset/gtsrb-german-traffic-sign.zip -d traffic_sign_dataset
# !rm traffic_sign_dataset/gtsrb-german-traffic-sign.zip
# !rm -rf traffic_sign_dataset/Meta
# !rm -rf traffic_sign_dataset/meta
# !rm -rf traffic_sign_dataset/test
# !rm -rf traffic_sign_dataset/train
# !rm traffic_sign_dataset/Meta.csv
# + [markdown] id="r7o25gk7F81b"
# ### Import Libraries
# + id="i5ZWejsGF7F0"
import os
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.image import imread
import seaborn as sns
import random
from PIL import Image
from sklearn.model_selection import train_test_split
from tensorflow.keras.utils import to_categorical
import tensorflow as tf
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense,Flatten, Dropout, Conv2D, MaxPool2D
# + [markdown] id="cDyNa8jYGgLY"
# ### Image Processing
# + colab={"base_uri": "https://localhost:8080/", "height": 874} id="39Y29pezGZ16" outputId="a01bbfc2-c1b7-4211-a633-cda9858cea1f"
plt.figure(figsize=(12,12))
path = 'traffic_sign_dataset/Test'
for i in range(1,17):
plt.subplot(4,4,i)
plt.tight_layout()
random_image = imread(path +'/'+ random.choice(sorted(os.listdir(path))))
plt.imshow(random_image)
plt.xlabel(random_image.shape[1], fontsize = 10)
plt.ylabel(random_image.shape[0], fontsize = 10)
# + id="sR7Bj1RDHKkA"
dim1 = []
dim2 = []
for i in range(0,43):
labels = 'traffic_sign_dataset/Train' + '/{0}'.format(i)
image_path = os.listdir(labels)
for x in image_path:
img = imread(labels + '/' + x)
dim1.append(img.shape[0])
dim2.append(img.shape[1])
# + colab={"base_uri": "https://localhost:8080/"} id="NMDYb3UPHrH-" outputId="e792b707-aa3f-4d25-a1ce-c49808390838"
print('Dimension 1 Mean: ',np.mean(dim1))
print('Dimension 2 Mean: ',np.mean(dim2))
# + id="MtSnM8k3H25F"
images = []
label_id = []
for i in range(43):
labels = 'traffic_sign_dataset/Train' + '/{0}'.format(i)
image_path = os.listdir(labels)
for x in image_path:
img = Image.open(labels + '/' + x)
img = img.resize((50,50))
img = np.array(img)
images.append(img)
label_id.append(i)
# + id="BMJUU1fFIhUk"
images = np.array(images)
images = images/255
# + colab={"base_uri": "https://localhost:8080/"} id="KYKKMJ73ImIm" outputId="4f2b5a14-bd7e-41ee-91cd-5258ecd6cd25"
label_id = np.array(label_id)
label_id.shape
# + colab={"base_uri": "https://localhost:8080/"} id="NCIT_YekIrSr" outputId="3c01354a-9534-4084-f2b9-c530576c11ea"
images.shape
# + colab={"base_uri": "https://localhost:8080/"} id="4TYyGLJVItWk" outputId="98fe9a18-e741-47d7-99f7-3534c31893ff"
label_counts = pd.DataFrame(label_id).value_counts()
label_counts.head()
# + [markdown] id="8616v5frI4fS"
# ### Splitting the data
# + id="1dsNb7yQI57w"
x_train, x_val, y_train, y_val = train_test_split(images, label_id, test_size=0.2, random_state=42)
# + id="6_xgHB-VJDlP"
y_train_cat = to_categorical(y_train)
y_val_cat = to_categorical(y_val)
# + [markdown] id="cn_9i1p1JPni"
# ### Build the model
# + id="HYZay0A8JOVD"
model = Sequential()
model.add(Conv2D(filters=64, kernel_size=(3,3), input_shape=x_train.shape[1:], activation='relu', padding='same'))
model.add(MaxPool2D(pool_size=(2,2)))
model.add(Dropout(0.5))
model.add(Conv2D(filters=64, kernel_size=(3,3), activation='relu'))
model.add(MaxPool2D(pool_size=(2,2)))
model.add(Dropout(0.5))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(43, activation='softmax'))
# + [markdown] id="rYLsP44ZKDvh"
# ### Train the model
# + colab={"base_uri": "https://localhost:8080/"} id="BMTRWVyRKEGn" outputId="be32faf4-45f7-466d-8175-1cf68f67f0db"
model.compile(loss='sparse_categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
model.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="JAGVJ9okKYin" outputId="420ddaed-b95f-4424-b043-066441057b98"
model.fit(x_train, y_train, epochs=50, batch_size=128, validation_data=(x_val, y_val), verbose=2)
# + [markdown] id="r_hdctEyLj75"
# ### Plot the history
# + colab={"base_uri": "https://localhost:8080/", "height": 530} id="4fycoX7QLmrr" outputId="988f888f-e5fb-46cc-b770-13203da14dbc"
evaluation = pd.DataFrame(model.history.history)
evaluation[['accuracy', 'val_accuracy']].plot()
evaluation[['loss', 'val_loss']].plot()
# + [markdown] id="1L7ZEF4xMwOw"
# ### Testing the model
# + id="rRg1l3kZMwB4"
test_path = 'traffic_sign_dataset/Test'
# !rm traffic_sign_dataset/Test/GT-final_test.csv
# + id="RK96dwTBM4qP"
def scaling(test_images, test_path):
images = []
image_path = test_images
for x in image_path:
img = Image.open(test_path + '/' + x)
img = img.resize((50,50))
img = np.array(img)
images.append(img)
images = np.array(images)
images = images / 255
return images
# + id="daN_YttHNU9e"
test_images = scaling(sorted(os.listdir(test_path)), test_path)
# + colab={"base_uri": "https://localhost:8080/"} id="J5GD68m-OG31" outputId="4417d8f2-fce6-4cc3-a6ea-5960fa492199"
test = pd.read_csv('traffic_sign_dataset/Test.csv')
y_test = test['ClassId'].values
y_test
# + colab={"base_uri": "https://localhost:8080/"} id="FQ89MXsBORzU" outputId="d9f5355c-f714-4a59-f857-d9bfe1a5293e"
y_pred = np.argmax(model.predict(test_images), axis=-1)
y_pred
# + id="-dIddURnOboz"
all_labels = ['Speed limit (20km/h)','Speed limit (30km/h)','Speed limit (50km/h)','Speed limit (60km/h)',
'Speed limit (70km/h)','Speed limit (80km/h)','End of speed limit (80km/h)','Speed limit (100km/h)',
'Speed limit (120km/h)','No passing','No passing for vechiles over 3.5 metric tons',
'Right-of-way at the next intersection','Priority road','Yield','Stop','No vechiles',
'Vechiles over 3.5 metric tons prohibited','No entry','General caution','Dangerous curve to the left',
'Dangerous curve to the right','Double curve','Bumpy road','Slippery road','Road narrows on the right',
'Road work','Traffic signals','Pedestrians','Children crossing','Bicycles crossing','Beware of ice/snow',
'Wild animals crossing','End of all speed and passing limits','Turn right ahead','Turn left ahead',
'Ahead only','Go straight or right','Go straight or left','Keep right','Keep left','Roundabout mandatory',
'End of no passing','End of no passing by vechiles over 3.5 metric']
# + colab={"base_uri": "https://localhost:8080/", "height": 62} id="uot8e7x_Ov8G" outputId="5fb8690d-2882-4cf7-f850-2d48ec2d380e"
img = Image.open(test_path + '/00001.png')
img
# + colab={"base_uri": "https://localhost:8080/"} id="0JYnij85O63L" outputId="85e28c96-dca5-4a4d-d1f2-089d7088c006"
print('Original label: ', all_labels[y_test[1]])
print('Predicted label: ', all_labels[y_pred[1]])
| 23 - Traffic Sign Classification/23 - Traffic_Sign_Classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import numpy as np
import matplotlib.pyplot as plt
import math
# ### One polynomial and a sigmoid function are defined
# +
poly_f = lambda x: x**3 + 2.*x**2
poly_Df = lambda x: 3.*x**2 + 4.*x
sigmoid = lambda x: 1/(1 + np.exp(-x))
sigmoid_Df = lambda x: sigmoid(x)*(1-sigmoid(x))
# -
# ### Newton's method
def newton(f,Df,x0,epsilon,max_iter):
xn = x0
for n in range(0,max_iter):
fxn = f(xn)
if abs(fxn) < epsilon:
print('Found solution at', xn, 'after',n,'iterations.')
return None
Dfxn = Df(xn)
if Dfxn == 0:
print('Zero derivative. No solution found.')
return None
xn = xn - fxn/Dfxn
print('Exceeded maximum iterations. No solution found.')
return None
# ### Gradient descent
def gradient_descent(gradient, start, learn_rate, n_iter=50, tolerance=1e-06):
iters = 0
vector = start
for _ in range(n_iter):
diff = -learn_rate * gradient(vector)
if np.all(np.abs(diff) <= tolerance):
break
vector += diff
iters += 1
print('Found that the local minima at', vector, 'after', iters,'iterations.')
# ### Newton's is applied to the polynomial function
gradient_descent(poly_Df, 2, 0.01, 10000)
# ### Gradient descent and Newton's is applied to polynomial function
newton(poly_f, poly_Df, 2, 0.01, 100)
# ### Gradient descent and Newton's is applied to sigmoid function
newton(sigmoid, sigmoid_Df, 1, 0.01, 1000)
gradient_descent(sigmoid_Df, 1, 0.1, 100000, 0.001)
| ametel01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pipeline
# language: python
# name: pipeline
# ---
import pickle
model = pickle.load(open('/home/lxu/Desktop/node2vec_graph_p1q2/1/word2vec_model.pickle','rb'))
model.wv.most_similar('Disease::DOID:10652')
model.wv.save_word2vec_format('embedding vectors')
r = open('embedding vectors','r')
print(len(r.readlines()))
all =[]
for line in r:
#line = line.split()
all.append(line)
all
word_vectors = model.wv
word_vectors['Disease::DOID:332']
word_vectors
# +
import numpy as np
import matplotlib.pyplot as plt
from sklearn.cluster import KMeans
from sklearn.datasets import make_blobs
# -
| notebooks/cluster nodes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Multimode gmm
# Created by: <NAME> (<EMAIL>)
#
# Description: gaussian mixture model implementation on tensorflow, the objective is a multimode gaussian distribution.
#
# This file serves as a visual test for the gmm extension modules
# +
import tensorflow as tf
import numpy as np
import collections
from twodlearn.tf_lib.datasets.generic_dataset import Datasets
from twodlearn.tf_lib.GMM import *
# %matplotlib notebook
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
# -
# ## 1. Create dataset
# +
def gmm_sampling(mu, sigma, w, n_samples=1):
# generates random vectors, sampled from a gausian mixture model
#
# - mu: 3d tensor containing the means for the gaussians.
# the "depth" dimention (3rd) is used to index the
# gaussians. [ kernel_id, dim]
# - sigma: 3d tensor containing the covariance matrix of the
# gaussians. [ kernel_id, dim] for diagonal matrices
# - w: vector in form of a 3d tensor containing the weights
# for each one of the gaussians, they have to sum one.
# [kernel_id]
n_kernels = mu.shape[0]
n_dim = mu.shape[1]
# random sample the kernel from which the output samples are going
# to be drawn
kernel_id= np.argmax(np.random.multinomial(1, w, size=[n_samples]), axis=1 )
out = np.zeros([n_samples, n_dim])
for i in range(n_samples):
out[i,:]= np.random.multivariate_normal(mu[kernel_id[i],:], np.diag(sigma[kernel_id[i],:])) # if diagonal
return out;
# +
n_samples = 1000
n_kernels_r = 2
n_dim = 2
mu_r = np.array([[1, 1], [10, 10]])
sigma_r = np.array([[0.5, 0.5], [2, 2]])
w_r = [1/n_kernels_r]*n_kernels_r # has to sum to one
# random sample from reference distribution
aux_x = gmm_sampling(mu_r, sigma_r, w_r, n_samples);
# build the dataset
data = Datasets(aux_x)
data.normalize()
print('Data shape: ', data.train.x.shape)
# plot
plt.plot(data.train.x[:,0], data.train.x[:,1], 'o')
# -
# ## 2. Model definition
sess = tf.InteractiveSession()
# +
n_kernels = 2
gmm_x = tf.placeholder( tf.float32, shape=(n_samples, n_dim))
gmm_w= tf.Variable(tf.truncated_normal( [1, n_kernels + 2*n_kernels*n_dim ], stddev=1.0),
name= 'gmm_weights'
)
gmm_params = GmmParamsLayer(n_dim, n_kernels, diagonal= True)
gmm_model = GmmModel(n_dim, n_kernels, diagonal= True)
# +
# transform parameters to 'valid' parameters
mu_r, sigma_r, w_r = gmm_params.evaluate(gmm_w)
# evaluate the gaussian mixture model using the 'valid' parameters
gmm_out = gmm_model.evaluate(gmm_x, mu_r, sigma_r, w_r)
# compute the loss of the gmm
gmm_loss = tf.reduce_sum(-tf.log(gmm_out + 0.00000001))
# Optimizer.
optimizer = tf.train.AdamOptimizer(0.05).minimize(gmm_loss) #0.001
# +
print('mu shape:', mu_r.get_shape())
print('sigma shape:', sigma_r.get_shape())
print('w shape:', w_r.get_shape())
print('out shape:', gmm_out.get_shape())
print('loss shape:', gmm_loss.get_shape())
# -
# ## 3. Training
# +
num_steps = 300 #1000
n_logging = 10
n_test_logg = 10
tf.initialize_all_variables().run()
print('Initialized')
mean_loss= 0
train_accuracy= 0
for step in range(num_steps):
_, l = sess.run([optimizer, gmm_loss],feed_dict={gmm_x : data.train.x})
mean_loss += l
if step%n_logging == 0:
print(step, ' | loss:', mean_loss/n_logging)
mean_loss= 0
# -
# ## 4. Plot
# +
n_plot= [20, 20];
# for testing
n_test= n_plot[0]*n_plot[1];
gmm_x_test = tf.placeholder( tf.float32, shape=(n_test, n_dim))
# evaluate the gaussian mixture model using the 'valid' parameters
gmm_out_test = gmm_model.evaluate(gmm_x_test, mu_r, sigma_r, w_r)
# +
xv, yv = np.meshgrid(np.linspace(-2, 2, n_plot[1]), np.linspace(-2, 2, n_plot[0]))
x_plot = np.stack([xv.transpose(), yv.transpose()], 2)
x_plot = np.reshape(x_plot, [-1,2])
[p_plot, mu_out, sigma_out, w_out,] = sess.run([gmm_out_test, mu_r, sigma_r, w_r],feed_dict= {gmm_x_test : x_plot})
print('mu:', mu_out, 'sigma:', sigma_out, 'w:', w_out)
p_plot = np.reshape(p_plot, [n_plot[1], n_plot[0]])
print(p_plot.shape)
# plot
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.plot_wireframe(xv, yv, p_plot)
# -
| twodlearn/Examples/gmm_multimode.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:jupyter-r]
# language: python
# name: conda-env-jupyter-r-py
# ---
# ### Summarize machine learning results and plot in R
#
# An experiment was done to compare convolutional neural networks (CNNs) with traditional machine learning techniques for a large image classification task. This notebook reads the results, computes some summary statistics, and plots the summary.
#
# R's ggplot2 library is used for plotting.
import warnings; warnings.simplefilter('ignore')
# %load_ext rpy2.ipython
# +
import numpy as np
import pandas as pd
import os
DIR = '/home/joefutrelle/data/ifcb-cnn'
assert os.path.exists(DIR)
# -
# Here we define functions for reading the [confusion matrices](https://en.wikipedia.org/wiki/Confusion_matrix) and computing the [F1 score](https://en.wikipedia.org/wiki/F1_score) for each classifer for each class.
# +
def read_cm(kind, n_classes=49, features_version=3):
fn = 'cm_{}_{}_v{}.csv'.format(kind, n_classes, features_version)
path = os.path.join(DIR, fn)
return pd.read_csv(path, index_col=0)
def read_cnn_cm(ft):
path = os.path.join(DIR, 'cm_cnn_49_{}.csv'.format(ft))
return pd.read_csv(path, index_col=0)
def summarize_confusion_matrix(cm):
total = cm.sum(axis=1)
true_positive = np.diag(cm)
false_positive = total - true_positive
false_negative = cm.sum(axis=0) - true_positive
true_negative = total.sum() - true_positive - false_negative - false_positive
precision = true_positive / (true_positive + false_positive)
recall = true_positive / (true_positive + false_negative)
f1 = 2 * (precision * recall) / (precision + recall)
pr = pd.DataFrame(dict(precision=precision, recall=recall, f1=f1),
columns=['precision','recall','f1'])
return pr
# -
# Five methods were evaluated. Summarize the confusion matrix of each one:
# +
features_version = 3
n_classes = len(read_cnn_cm('full').index)
rf_pr = summarize_confusion_matrix(read_cm('rf',n_classes,features_version))
nn_pr = summarize_confusion_matrix(read_cm('nn',n_classes,features_version))
svm_pr = summarize_confusion_matrix(read_cm('svm',n_classes,features_version))
cnn_full_pr = summarize_confusion_matrix(read_cnn_cm('full'))
cnn_transfer_pr = summarize_confusion_matrix(read_cnn_cm('transfer'))
rf_label = 'Random Forest'
mlp_label = 'MLP Neural Network'
svm_label = 'Support Vector Machines'
cnn_full_label = 'CNN (full)'
cnn_transfer_label = 'CNN (transfer)'
# -
# Reorganize the resulting summary into a Pandas dataframe where there is a row for each method/class pair, holding the F1 score
# +
# make a tidy dataframe where one column is method and one column is for each class's f1 score
from functools import reduce
dfs = []
for method, pr in [(rf_label, rf_pr),
(mlp_label, nn_pr),
(svm_label, svm_pr),
(cnn_full_label, cnn_full_pr),
(cnn_transfer_label, cnn_transfer_pr)]:
dfs.append(pd.DataFrame({'method': method, 'f1': pr['f1'] }))
df = reduce(lambda a,b: a.append(b), dfs)
df.head()
# -
# Now use R to make a violin plot of the results
# + magic_args="-i df -w 750 -h 400" language="R"
#
# library(ggplot2)
#
# # Create a Violin plot
# ggplot(df, aes(x = method, y = f1, fill = method)) +
# geom_violin() +
# labs(x = "Method", y = "F1 scores")
| 6. Summarize machine learning results.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %load_ext autoreload
# %autoreload 2
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import os
import sys
flopypth = os.path.join('..', '..', '..', 'flopy')
if flopypth not in sys.path:
sys.path.append(flopypth)
import flopy.modflow as mf
import flopy.mt3d as mt
import flopy.seawat as swt
import flopy.utils as fu
name = 'island'
mf_model = mf.Modflow(modelname = name)
L = 1000.
nlay = 20
nrow = 1
ncol = 100
delr = L / ncol
H = 20.
delz = H / nlay
top = np.ones((1, ncol)) * delz / 2.
top[0, 10:-10] += delz / 2.
botm = np.linspace(-delz / 2., -H + delz / 2., nlay)
perlen = 7300
dis = mf.ModflowDis(mf_model, nlay, nrow, ncol, delr = delr, delc = 1,
top = top, botm = botm, laycbd = 0, perlen = perlen,
nstp = 730, steady = False)
ibound = np.ones((nlay, nrow, ncol))
ibound[0, 0, :10] = -1
ibound[0, 0, 90:] = -1
bas = mf.ModflowBas(mf_model, ibound = ibound, strt = 0.0)
lpf = mf.ModflowLpf(mf_model, hk = 100.0)
rech = np.zeros((nrow, ncol))
rech[0, (ibound[0, 0, :] == 1)] = 2e-3
rch = mf.ModflowRch(mf_model, rech = rech)
pcg = mf.ModflowPcg(mf_model)
oc = mf.ModflowOc(mf_model)
mf_model.write_input()
#mf_model.run_model2()
mt_model = mt.Mt3dms(name, 'nam_mt3dms', mf_model)
adv = mt.Mt3dAdv(mt_model, mixelm = 0, percel = 1.)
sconc_1 = np.ones([nlay, nrow, ncol]) * 35.7
timprs = np.arange(1, perlen, 100)
btn = mt.Mt3dBtn(mt_model, ncomp=1, sconc=sconc_1, prsity = 0.25, timprs = timprs)
dsp = mt.Mt3dDsp(mt_model, al=0.1, dmcoef=5e-10*3600.*24.)
gcg = mt.Mt3dGcg(mt_model, cclose = 1e-8)
# +
crch = {0: 0.001}
sp_data = []
for c in range(10):
dd = np.array([0, 0, c, 0, 1])
sp_data.append(dd)
for c in range(90, 100):
dd = np.array([0, 0, c, 0, 1])
sp_data.append(dd)
stress_period_data = {0:sp_data}
ssm = mt.Mt3dSsm(mt_model, crch=crch, stress_period_data=stress_period_data)
mt_model.write_input()
# -
swt_model = swt.Seawat(name, 'nam_swt', mf_model, mt_model, exe_name='swt_v4') # Coupled to modflow model mf and mt3dms model mt
vdf = swt.SeawatVdf(swt_model, firstdt = 0.1, denseslp = .7)
swt_model.write_input()
swt_model.run_model(silent=True)
# +
conc = fu.UcnFile('MT3D001.UCN')
c = conc.get_data(totim = 7300)
y, x, z = dis.get_node_coordinates()
X, Z = np.meshgrid(x, z[:, 0, 0])
plt.contourf(X, Z, c[:, 0, :])
# -
| examples/Testing/island_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## SFR package example
# Demonstrates functionality of Flopy SFR module using the example documented by [Prudic and others (2004)](http://pubs.er.usgs.gov/publication/ofr20041042):
#
# #### Problem description:
#
# * Grid dimensions: 1 Layer, 15 Rows, 10 Columns
# * Stress periods: 1 steady
# * Flow package: LPF
# * Stress packages: SFR, GHB, EVT, RCH
# * Solver: SIP
#
# <img src="./img/Prudic2004_fig6.png" width="400" height="500"/>
# +
import sys
import platform
import os
import numpy as np
import glob
import shutil
import matplotlib as mpl
import matplotlib.pyplot as plt
import pandas as pd
# run installed version of flopy or add local path
try:
import flopy
except:
fpth = os.path.abspath(os.path.join('..', '..'))
sys.path.append(fpth)
import flopy
import flopy.utils.binaryfile as bf
from flopy.utils.sfroutputfile import SfrFile
mpl.rcParams['figure.figsize'] = (11, 8.5)
print(sys.version)
print('numpy version: {}'.format(np.__version__))
print('pandas version: {}'.format(pd.__version__))
print('matplotlib version: {}'.format(mpl.__version__))
print('flopy version: {}'.format(flopy.__version__))
# -
#Set name of MODFLOW exe
# assumes executable is in users path statement
exe_name = 'mf2005'
if platform.system() == 'Windows':
exe_name += '.exe'
# # #### copy over the example files to the working directory
path = 'data'
if os.path.isfile(path):
os.remove(path)
elif os.path.isdir(path):
shutil.rmtree(path)
os.mkdir(path)
gpth = os.path.join('..', 'data', 'mf2005_test', 'test1ss.*')
for f in glob.glob(gpth):
shutil.copy(f, path)
gpth = os.path.join('..', 'data', 'mf2005_test', 'test1tr.*')
for f in glob.glob(gpth):
shutil.copy(f, path)
# ### Load example dataset, skipping the SFR package
m = flopy.modflow.Modflow.load('test1ss.nam', version='mf2005', exe_name=exe_name,
model_ws=path, load_only=['ghb', 'evt', 'rch', 'dis', 'bas6', 'oc', 'sip', 'lpf'])
oc = m.oc
oc.stress_period_data
# ### Read pre-prepared reach and segment data into numpy recarrays using numpy.genfromtxt()
# Reach data (Item 2 in the SFR input instructions), are input and stored in a numpy record array
# http://docs.scipy.org/doc/numpy/reference/generated/numpy.recarray.html
# This allows for reach data to be indexed by their variable names, as described in the SFR input instructions.
#
# For more information on Item 2, see the Online Guide to MODFLOW:
# <http://water.usgs.gov/nrp/gwsoftware/modflow2000/MFDOC/index.html?sfr.htm>
rpth = os.path.join('..', 'data', 'sfr_examples', 'test1ss_reach_data.csv')
reach_data = np.genfromtxt(rpth, delimiter=',', names=True)
reach_data
# ### Segment Data structure
# Segment data are input and stored in a dictionary of record arrays, which
spth = os.path.join('..', 'data', 'sfr_examples', 'test1ss_segment_data.csv')
ss_segment_data = np.genfromtxt(spth, delimiter=',', names=True)
segment_data = {0: ss_segment_data}
segment_data[0][0:1]['width1']
# ### define dataset 6e (channel flow data) for segment 1
# dataset 6e is stored in a nested dictionary keyed by stress period and segment,
# with a list of the following lists defined for each segment with icalc == 4
# FLOWTAB(1) FLOWTAB(2) ... FLOWTAB(NSTRPTS)
# DPTHTAB(1) DPTHTAB(2) ... DPTHTAB(NSTRPTS)
# WDTHTAB(1) WDTHTAB(2) ... WDTHTAB(NSTRPTS)
channel_flow_data = {0: {1: [[0.5, 1.0, 2.0, 4.0, 7.0, 10.0, 20.0, 30.0, 50.0, 75.0, 100.0],
[0.25, 0.4, 0.55, 0.7, 0.8, 0.9, 1.1, 1.25, 1.4, 1.7, 2.6],
[3.0, 3.5, 4.2, 5.3, 7.0, 8.5, 12.0, 14.0, 17.0, 20.0, 22.0]]}}
# ### define dataset 6d (channel geometry data) for segments 7 and 8
# dataset 6d is stored in a nested dictionary keyed by stress period and segment,
# with a list of the following lists defined for each segment with icalc == 4
# FLOWTAB(1) FLOWTAB(2) ... FLOWTAB(NSTRPTS)
# DPTHTAB(1) DPTHTAB(2) ... DPTHTAB(NSTRPTS)
# WDTHTAB(1) WDTHTAB(2) ... WDTHTAB(NSTRPTS)
channel_geometry_data = {0: {7: [[0.0, 10.0, 80.0, 100.0, 150.0, 170.0, 240.0, 250.0],
[20.0, 13.0, 10.0, 2.0, 0.0, 10.0, 13.0, 20.0]],
8: [[0.0, 10.0, 80.0, 100.0, 150.0, 170.0, 240.0, 250.0],
[25.0, 17.0, 13.0, 4.0, 0.0, 10.0, 16.0, 20.0]]}}
# ### Define SFR package variables
nstrm = len(reach_data) # number of reaches
nss = len(segment_data[0]) # number of segments
nsfrpar = 0 # number of parameters (not supported)
nparseg = 0
const = 1.486 # constant for manning's equation, units of cfs
dleak = 0.0001 # closure tolerance for stream stage computation
ipakcb = 53 # flag for writing SFR output to cell-by-cell budget (on unit 53)
istcb2 = 81 # flag for writing SFR output to text file
dataset_5 = {0: [nss, 0, 0]} # dataset 5 (see online guide)
# ### Instantiate SFR package
# Input arguments generally follow the variable names defined in the Online Guide to MODFLOW
sfr = flopy.modflow.ModflowSfr2(m, nstrm=nstrm, nss=nss, const=const, dleak=dleak, ipakcb=ipakcb, istcb2=istcb2,
reach_data=reach_data,
segment_data=segment_data,
channel_geometry_data=channel_geometry_data,
channel_flow_data=channel_flow_data,
dataset_5=dataset_5, unit_number=15)
sfr.reach_data[0:1]
# ### Plot the SFR segments
# any column in the reach_data array can be plotted using the ```key``` argument
sfr.plot(key='iseg');
# ### Check the SFR dataset for errors
chk = sfr.check()
m.external_fnames = [os.path.split(f)[1] for f in m.external_fnames]
m.external_fnames
m.write_input()
m.run_model()
# ### Load SFR formated water balance output into pandas dataframe using the `SfrFile` class
# * requires the **pandas** library
sfr_outfile = os.path.join('..', 'data', 'sfr_examples', 'test1ss.flw')
sfrout = SfrFile(sfr_outfile)
df = sfrout.get_dataframe()
df.head()
# #### Plot streamflow and stream/aquifer interactions for a segment
inds = df.segment == 3
print(df.reach[inds].astype(str))
#ax = df.ix[inds, ['Qin', 'Qaquifer', 'Qout']].plot(x=df.reach[inds])
ax = df.loc[inds, ['reach', 'Qin', 'Qaquifer', 'Qout']].plot(x='reach')
ax.set_ylabel('Flow, in cubic feet per second')
ax.set_xlabel('SFR reach');
# ### Look at stage, model top, and streambed top
streambed_top = m.sfr.segment_data[0][m.sfr.segment_data[0].nseg == 3][['elevup', 'elevdn']][0]
streambed_top
df['model_top'] = m.dis.top.array[df.row.values - 1, df.column.values -1]
fig, ax = plt.subplots()
plt.plot([1, 6], list(streambed_top), label='streambed top')
#ax = df.loc[inds, ['stage', 'model_top']].plot(ax=ax, x=df.reach[inds])
ax = df.loc[inds, ['reach', 'stage', 'model_top']].plot(ax=ax, x='reach')
ax.set_ylabel('Elevation, in feet')
plt.legend();
# ### Get SFR leakage results from cell budget file
bpth = os.path.join('data', 'test1ss.cbc')
cbbobj = bf.CellBudgetFile(bpth)
cbbobj.list_records()
sfrleak = cbbobj.get_data(text=' STREAM LEAKAGE')[0]
sfrleak[sfrleak == 0] = np.nan # remove zero values
# ### Plot leakage in plan view
im = plt.imshow(sfrleak[0], interpolation='none', cmap='coolwarm', vmin = -3, vmax=3)
cb = plt.colorbar(im, label='SFR Leakage, in cubic feet per second');
# ### Plot total streamflow
sfrQ = sfrleak[0].copy()
sfrQ[sfrQ == 0] = np.nan
sfrQ[df.row.values-1, df.column.values-1] = df[['Qin', 'Qout']].mean(axis=1).values
im = plt.imshow(sfrQ, interpolation='none')
plt.colorbar(im, label='Streamflow, in cubic feet per second');
# ## Reading transient SFR formatted output
# the `SfrFile` class handles this the same way
#
# files for the transient version of the above example were already copied to the `data` folder in the third cell above
# first run the transient model to get the output:
# ```
# >mf2005 test1tr.nam
# ```
flopy.run_model(exe_name, 'test1tr.nam', model_ws=path, silent=True)
sfrout_tr = SfrFile(os.path.join('data', 'test1tr.flw'))
dftr = sfrout_tr.get_dataframe()
dftr.head()
# ### plot a hydrograph
# plot `Qout` (simulated streamflow) and `Qaquifer` (simulated stream leakage) through time
fig, axes = plt.subplots(2, 1, sharex=True)
dftr8 = dftr.loc[(dftr.segment == 8) & (dftr.reach == 5)]
dftr8.Qout.plot(ax=axes[0])
axes[0].set_ylabel('Simulated streamflow, cfs')
dftr8.Qaquifer.plot(ax=axes[1])
axes[1].set_ylabel('Leakage to aquifer, cfs');
| examples/Notebooks/flopy3_sfrpackage_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data exploration
# In this notebook we are going to study the data that will be use to develop benchmarking tools for binary classification algorithms.
#
# The data was developed for the [LHC Olympics 2020](https://lhco2020.github.io/homepage/). They published four different dataset of multijets events with the possibility of Beyond the Standar Model (BSM) physics. The data used for the project consist of the [R&D dataset](https://zenodo.org/record/2629073#.XKdewGXlRg0) and the dataset and masterkey for the [Black Box 1](https://zenodo.org/record/4536624)(BB1).
#
# ## The data
#
# Both the R&D and BB1 datasets have data for the same event but are generated with a slightly different configuration on the Monte Carlo simulation:
#
# The datasets consists on events from quantum chromodynamic (QCD), our background, and events of the type $Z'\rightarrow XY$ with $X\rightarrow q\bar{q}$ and $Y\rightarrow q\bar{q}$, our signal. The masses for the BSM particles on the R&D dataset are *Z'*, *X* and *Y* ara 3.5 TeV, 500 GeV and 100 GeV, respectively. The Feynman diagram is as follow:
#
# 
#
# For the R&D dataset there are 1M background events and 100.000 signal events.
#
# The BB1 dataset has 834 signal events from the total 1M events.
#
# ### .h5 files
# Each row of the file is an event with:
# - at least one anti-kT jet with R=1.0
# - pseudorapidity $|\eta|<2,5$
# - and transverse momentum $pT> 1,2$ TeV.
#
# For each event there is a list of all the hadrons as follow:
#
# |Evento|$pT$ | $\eta$ | $\phi$ | $pT$ | $\eta$ | $\phi$ | $\cdots$ | signal |
# |------|-----|--------|--------|------|--------|--------|--------|--------|
# | 1 | $\cdots$| $\cdots$| $\cdots$| $\cdots$| $\cdots$| $\cdots$| $\cdots$| 1.0|
# | 2 | $\cdots$| $\cdots$| $\cdots$| $\cdots$| $\cdots$| $\cdots$| $\cdots$| 0.0|
# | $\vdots$| $\cdots$| $\cdots$| $\cdots$| $\cdots$| $\cdots$| $\cdots$| $\cdots$| 0.0|
#
# The last column is the information about background (0) or signal (1). The events are zero-padded up to 700 hadrons.
#
# For more information about the data visit the [LHC Olympics 2020 website](https://lhco2020.github.io/homepage/)
#
# ## R&D dataset
#
# This dataset is mainly to setup and train the algorithms:
# Importing the main libraries
import pandas as pd
import h5py
import numpy as np
import matplotlib.pyplot as plt
# Depending on the memory of the computer,
# you'll be able to read more or less data
df = pd.read_hdf("../../events_anomalydetection.h5", stop=100000)
# We know that for this dataset the percentage of signal is $100.000/1.100.000 = 9.09 \%$. Let's check the percentage for the data that was read on the cell above:
# +
# Last column = 1 is signal
dfsig = df[df.iloc[:,-1]==1]
ratio_sig = dfsig.shape[0]/df.shape[0]
print('The percentage of signal is {:.2%}'.format(ratio_sig))
# -
# We have a good representation of signal to continue with the analysis.
#
# ### Raw data
# The data is as explained before:
df.head()
# The events shouldn't have the same amount of hadrons:
# +
# Eliminating the signal to avoid counting it as hadrons
ns_df = df.iloc[:,:-1]
# We count the values of pT that are not zero
df['n_hadrons'] = ns_df.iloc[:,::3].gt(0).sum(axis='columns')
for event in range(5):
print("Number of hadrons in event {}: {}".format(event, df.loc[event,'n_hadrons']))
# -
# By plotting distributions we can get a better idea of the data.
#
# Starting by plotting the distribution of the number of hadron separating signal from background. For that we will be using functions on the <code>plotools</code> module on the <code>benchtools</code> package.
# +
from benchtools.src.plotools import bkg_sig_hist
bkg_sig_hist(df, variable='n_hadrons', label=2100) # label=2100 because that's the column for signal or background
plt.show()
# -
# We see that the number of hadrons for background events is more broad with a lower peak around 150 hadrons. However, for the signal we have a narrower peak closer to 100 hadrons, meaning that signal events tends to have less hadrons.
#
# Let's separate $pT$, $\eta$ and $\phi$ in their own dataframes, so we can plot the distributions:
# +
# Each 3 values is pT, each 3 starting from 1 is eta, each 3 starting from 2 is phi
# And replacing the zeros with NaN so we can calculate the mean with .mean()
pt_df = ns_df.iloc[:,::3].replace(0, np.NaN)
eta_df = ns_df.iloc[:,1::3].replace(0, np.NaN)
phi_df = ns_df.iloc[:,2::3].replace(0, np.NaN)
# Calculating the mean value for each
df['mean_pT'] = pt_df.mean(axis=1)
df['mean_eta']= eta_df.mean(axis=1)
df['mean_phi']= phi_df.mean(axis=1)
# -
# Plotting this values:
for variable in ['mean_pT', 'mean_eta', 'mean_phi']:
bkg_sig_hist(df, variable, label=2100)
plt.show()
# We found **a difference between the mean value of pT for background and signal**. For phi and eta the data is distributed normally for both the signal and the background.
#
# Maybe the distribution for the more energetic hadron tell us something:
# +
df['pT_max'] = pt_df.max(axis=1)
bkg_sig_hist(df, variable='pT_max', label=2100)
plt.show()
# -
# There are some differences: the peak for the signal is higher, but let's see if we can get more information by clustering the jets.
# ### Clustered data
# For clustering the events we are using <code>pyjet</code>.
#
# I'll use functions from <code>benchtools.src.clustering</code> to get some important features about the events as: $pT$, *mass*, $\eta$, $\phi$, *energy*, *subjettiness* ($\tau_{21}$) and *number of constituyents* for the two principal jets, their *angular distance* ($\delta R$), the *invariant mass*($m_{jj}$), the *number of hadrons for the event* and the *label*.
#
# First, I'm gonna use <code>build_features</code> to iterate over the file to generate the features:
from benchtools.src.clustering import build_features
build_features("../../events_anomalydetection.h5", 3, "data_exploration_RD")
# Using <code>read_multifiles</code> we get the files as one dataframe:
from benchtools.src.datatools import read_multifiles
df = read_multifiles("data_exploration_RD", 3)
df.head()
# ### Distributions
# We can graph the distribution of the variables to see if there is a difference for the background and the signal:
# Getting the name of the variables
variables = df.columns.tolist()
variables
# Starting with the transversal momentum for the most energetic jet (or jet 1) of each event
bkg_sig_hist(df, variable='pT_j1', label='label')
plt.show()
# We see that the transversal momentum *pT* for background events decays smoothly, indicating that the most energetic jets from these events tends to have *pT* lower than 1500 GeV. For signal events the most energetic jets haver larger *pT*, between 1500 and 2000 GeV.
# The mass of jet 1
bkg_sig_hist(df, variable='m_j1', label='label')
plt.show()
# Again, for background events the distribution of mass decays, bein most of the jets less massive. However, for the signal events we see two peaks: at 100 and 500 GeV. These are the masses of the *X* and *Y* particles for the event $Z'\rightarrow XY$
# The energy of jet 1
bkg_sig_hist(df, variable='E_j1', label='label')
plt.show()
# Again, the background decays smoothly and most of the jets have lower energies. For the jets on the signal we see a peak at around 2000 GeV, with a different shape, indicating that jets of the signal tend to have higher energies than those of the background.
# The subjettiness variable of jet 1
bkg_sig_hist(df, variable='tau_21_j1', label='label')
plt.show()
# We also see a different distribution for these variables. According to the definition of subjettiness we are looking for the probability of having subjets. This variable is lower when the jets have two subjets in it. Hence, the signal is grouped towards smaller values than the background, meaning that the jets of the signal are more consistent with having two jets.
# Graphing eta and phi
for variable in ['eta_j1', 'phi_j1', 'nhadrons_j1']:
bkg_sig_hist(df, variable=variable, label='label')
# The variable $\eta$ and the number of hadrons have differences but aren't as evident.
#
# For $\eta$ we see that is more central on the signal events. The number of hadrons are narrower for the signal. On the variable $\phi$ there is not a marked difference between signal and background.
#
# Now, for the relation between the two most energetic jets of each event:
# The invariant mass
bkg_sig_hist(df, variable='m_jj', label='label')
plt.show()
# The invariant mass also decays for background events. However, for the signal events we see a narrow peak at 3500 GeV which is the mass for *Z* in $Z'\rightarrow XY$
# The distance between jets
bkg_sig_hist(df, variable='deltaR_j12', label='label')
plt.show()
# The signal distribution has a narrower and higher peak. However, the distributions are similar.
# And the number of hadrons for the events
bkg_sig_hist(df, variable='n_hadrons', label='label')
plt.show()
# Here we see that the number of hadrons for signal events tends to be lower thant for background events. Also, for signal the distribution is narrower.
# ## Correlations
# We want to see how the variables relate to each other
# +
# We want to have the correlations for signal and background in separate dataframes
df_bkg = df[df['label']==0].drop('label', axis=1)
df_sig = df[df['label']==1].drop('label', axis=1)
df_bkg_corr = df_bkg.corr()
df_sig_corr = df_sig.corr()
# To graph the correlation map:
for name, corr in [('background', df_bkg_corr), ('signal', df_sig_corr)]:
# Define the figure and size
fig = plt.figure(figsize=[7,6])
# Plot axis
ax = fig.add_subplot(111)
# Color bar for the mapping
cax = ax.matshow(corr,cmap='coolwarm', vmin=-1, vmax=1)
fig.colorbar(cax)
# Axis
ticks = np.arange(0,len(corr.columns),1)
ax.set_xticks(ticks)
plt.xticks(rotation=90)
ax.set_yticks(ticks)
ax.set_xticklabels(corr.columns)
ax.set_yticklabels(corr.columns)
# Title
plt.title('Correlation: {}'.format(name))
plt.show()
# -
# We can plot some of the relations that looks interesting to the project using the <code>bkg_sig_scatter</code> function:
# +
from benchtools.src.plotools import bkg_sig_scatter
relations = [('pT_j1', 'm_j1'), ('pT_j1', 'm_jj'), ('pT_j1', 'pT_j2'), ('pT_j1', 'deltaR_j12'), ('nhadrons_j1', 'm_j1'), ('nhadrons_j1', 'm_j2'),
('nhadrons_j1', 'nhadrons_j2'), ('eta_j1', 'E_j1'), ('phi_j1', 'phi_j2')]
for x,y in relations:
bkg_sig_scatter(df, x, y)
plt.show()
# -
# All the distributions and correlation plots are in [this notebook](2.0-all-distribution-correlation-plots.ipynb)
# ## BB1 dataset
# For this dataset, the masses for the BSM particles are *Z'*, *X* and *Y* ara 3.823 TeV, 732 GeV and 378 GeV, respectively.
#
# We are going to run the exact same code as before, but with the BB1 data:
# Depending on the memory of the computer,
# you'll be able to read more or less data
df = pd.read_hdf("../../events_anomalydetection.h5", stop=100000)
| notebooks/.ipynb_checkpoints/1.0-data-exploration-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
class BinaryTreeNode:
def __init__(self, data):
self.data = data
self.left = None
self.right = None
# -
import queue
def takeinputlevelwise():
q = queue.Queue()
print("enter root")
rootdata = int(input())
if rootdata == -1:
return None
root = BinaryTreeNode(rootdata)
q.put(root)
while (not(q.empty())):
curr_node = q.get()
print("enter left child of" ,curr_node.data)
leftnode = int(input())
if leftnode != -1:
leftchild = BinaryTreeNode(leftnode)
curr_node.left = leftchild
q.put(leftchild)
print("enter right child of" ,curr_node.data)
rightnode = int(input())
if rightnode != -1:
righttchild = BinaryTreeNode(rightnode)
curr_node.right = righttchild
q.put(righttchild)
return root
def nodetorootpath(root, s):
if root == None:
return None
if root.data == s:
l = list()
l.append(root.data)
return l
leftoutput = nodetorootpath(root.left, s)
if leftoutput != None:
leftoutput.append(root.data)
return leftoutput
rightoutput = nodetorootpath(root.right, s)
if rightoutput != None:
rightoutput.append(root.data)
return rightoutput
else:
return None
root = takeinputlevelwise()
l = nodetorootpath(root, 5)
for i in l:
print(i)
| node_to_root_path.py.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: policy-toolkit
# language: python
# name: policy-toolkit
# ---
# # 1 Introduction
# #### Author: <NAME>
#
# Visualize the geographic extent and density of candidate articles identified by keyword subsetting of document titles. Should be run after `2-download-fulltext`.
# ## 1.0 Package imports
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import geopandas as gpd
# %matplotlib inline
import os
import itertools
# # 2 Visualizations
#
# ## 2.0 Function definitions
def load_year(country: str, year: int) -> pd.DataFrame:
path = '../data/{}/metadata/{}/'.format(country, str(year))
lats, longs = [], []
types = []
admin = gpd.read_file('../data/reference/{}.geojson'.format(country))
for file in os.listdir(path):
if ".csv" in file:
df = pd.read_csv(path + file)
lat = list(df['ActionGeo_Lat'])
long = list(df['ActionGeo_Long'])
lats.extend(lat)
longs.extend(long)
types.extend(list(df['QuadClass']))
coords = pd.DataFrame({'Latitude': np.array(lats).flatten(),
'Longitude': np.array(longs).flatten(),
'Type': np.array(types).flatten()})
coords['latlong'] = coords["Latitude"].astype(str) + coords["Longitude"].astype(str)
coords = coords.groupby(['latlong', 'Latitude', 'Longitude']).count().reset_index()
coords = gpd.GeoDataFrame(coords, geometry=gpd.points_from_xy(coords.Longitude,
coords.Latitude))
coords['Type'] = coords['Type'].astype(float)
return admin, coords
import pickle as pkl
def load_positives(country: str, year: int, month: int) -> pd.DataFrame:
lats, longs = [], []
types = []
classifications = pd.read_csv(f'../data/{country}/output/{str(year)}/{str(month).zfill(2)}.csv')
positive_idx = list(classifications['Index'][classifications['Conflict'] == True])
for idx in positive_idx:
path = f'../data/{country}/json/{str(year)}/{str(month).zfill(2)}/{str(idx).zfill(5)}.pkl'
with open(path, 'rb') as pickle_file:
data = pkl.load(pickle_file)
lat = data['actions'][0]['ActionGeo_Lat']
long = data['actions'][0]['ActionGeo_Long']
print(data['actions'])
lats.append(lat)
longs.append(long)
types.append(data['actions'][0]['QuadClass'])
coords = pd.DataFrame({'Latitude': np.array(lats).flatten(),
'Longitude': np.array(longs).flatten(),
'Type': np.array(types).flatten()})
coords['latlong'] = coords["Latitude"].astype(str) + coords["Longitude"].astype(str)
coords = coords.groupby(['latlong', 'Latitude', 'Longitude']).count().reset_index()
coords = gpd.GeoDataFrame(coords, geometry=gpd.points_from_xy(coords.Longitude,
coords.Latitude))
coords['Type'] = coords['Type'].astype(float)
return coords
# ## 2.1 Brazil
# +
f, (ax1, ax2, ax3) = plt.subplots(ncols = 3, figsize=(20, 12))
brazil, coords = load_year('brazil', 2017)
ax1 = brazil.plot(color='white', edgecolor='black', ax = ax1)
ax1.set_title("Brazil - 2017 - {}".format(str(coords['Type'].sum())))
coords.plot(ax=ax1, markersize = (3 + coords['Type'] / 30))
brazil, coords = load_year('brazil', 2018)
ax2 = brazil.plot(color='white', edgecolor='black', ax = ax2)
ax2.set_title("Brazil - 2018 - {}".format(str(coords['Type'].sum())))
coords.plot(ax=ax2, markersize = (3 + coords['Type'] / 30))
brazil, coords = load_year('brazil', 2019)
ax3 = brazil.plot(color='white', edgecolor='black', ax = ax3)
ax3.set_title("Brazil - 2019 - {}".format(str(coords['Type'].sum())))
coords.plot(ax=ax3, markersize = (3 + coords['Type'] / 30))
# +
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = (13,10)
for year in (2017, 2018, 2019):
for month in range(12, 13):
brazil, coords1 = load_year('brazil', year)
coords = load_positives('brazil', year, month)
pl1 = brazil.plot(color='white', edgecolor='black')
pl1.set_title(f"{str(year)}/{str(month).zfill(2)} - {str(coords['Type'].sum())[:-2]} events")
coords.plot(ax=pl1, markersize = (15 + coords['Type'] * 2))
plt.savefig(f'../data/brazil/figs/{str(year)}-{str(month).zfill(2)}.png')
#pl.show()
# -
import imageio
images = []
filenames = [x for x in os.listdir("../data/brazil/figs/") if '.png' in x]
filenames = sorted(filenames)
print(filenames)
for filename in filenames:
print(filename)
images.append(imageio.imread('../data/brazil/figs/' + filename))
imageio.mimsave('../data/brazil/figs/movie.gif', images, duration=0.25)
# ## 2.2 Indonesia
# +
f, (ax1, ax2, ax3) = plt.subplots(nrows = 3, figsize=(12, 17))
indonesia, coords = load_year('indonesia', 2017)
ax1 = indonesia.plot(color='white', edgecolor='black', ax = ax1)
ax1.set_title("Indonesia - 2017 - {}".format(str(coords['Type'].sum())))
ax1.set_xlim(90, 145)
ax1.set_ylim(-12, 8)
coords.plot(ax=ax1, markersize = (3 + coords['Type'] / 300))
indonesia, coords = load_year('indonesia', 2018)
ax2 = indonesia.plot(color='white', edgecolor='black', ax = ax2)
ax2.set_title("Indonesia - 2018 - {}".format(str(coords['Type'].sum())))
ax2.set_xlim(90, 145)
ax2.set_ylim(-12, 8)
coords.plot(ax=ax2, markersize = (3 + coords['Type'] / 300))
indonesia, coords = load_year('indonesia', 2019)
ax3 = indonesia.plot(color='white', edgecolor='black', ax = ax3)
ax3.set_title("Indonesia - 2019 - {}".format(str(coords['Type'].sum())))
ax3.set_xlim(90, 145)
ax3.set_ylim(-12, 8)
coords.plot(ax=ax3, markersize = (3 + coords['Type'] / 300))
# +
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = (13,5)
for year in (2017, 2018, 2019):
for month in range(1, 12):
indonesia, coords1 = load_year('indonesia', year)
coords = load_positives('indonesia', year, month)
pl = indonesia.plot(color='white', edgecolor='black')
pl.set_title(f"{str(year)}/{str(month).zfill(2)} - {str(coords['Type'].sum())[:-2]} events")
pl.set_xlim(90, 145)
pl.set_ylim(-12, 8)
coords.plot(ax=pl, markersize = (10 + coords['Type']))
plt.savefig(f'../data/indonesia/figs/{str(year)}-{str(month).zfill(2)}.png')
#pl.show()
# -
import imageio
images = []
filenames = [x for x in os.listdir("../data/indonesia/figs/") if '.png' in x]
filenames = sorted(filenames)
print(filenames)
for filename in filenames:
print(filename)
images.append(imageio.imread('../data/indonesia/figs/' + filename))
imageio.mimsave('../data/indonesia/figs/movie.gif', images, duration=0.25)
# # 2.3 Mexico
# +
f, (ax1, ax2, ax3) = plt.subplots(ncols = 3, figsize=(20, 12))
mexico, coords = load_year('mexico', 2017)
ax1 = mexico.plot(color='white', edgecolor='black', ax = ax1)
ax1.set_title("Mexico - 2017 - {}".format(str(coords['Type'].sum())))
ax1.set_ylim(13, 35)
ax1.set_xlim(-120, -85)
coords.plot(ax=ax1, markersize = (3 + coords['Type'] / 150))
mexico, coords = load_year('mexico', 2018)
ax2 = mexico.plot(color='white', edgecolor='black', ax = ax2)
ax2.set_title("Mexico - 2018 - {}".format(str(coords['Type'].sum())))
ax2.set_ylim(13, 35)
ax2.set_xlim(-120, -85)
coords.plot(ax=ax2, markersize = (3 + coords['Type'] / 150))
mexico, coords = load_year('mexico', 2019)
ax3 = brazil.plot(color='white', edgecolor='black', ax = ax3)
ax3.set_title("Mexico - 2019 - {}".format(str(coords['Type'].sum())))
ax3.set_ylim(13, 35)
ax3.set_xlim(-120, -85)
coords.plot(ax=ax3, markersize = (3 + coords['Type'] / 150))
| notebooks/2-visualize-fulltext.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import requests
import pandas as pd
import json
from pandas.io.json import json_normalize
import sqlite3 as lite
import numpy as np
from sqlalchemy import create_engine
sqlite_file = 'sqlite://///Users/Kruthika/Projects/DDL/04-team3/census.db'
engine = create_engine(sqlite_file)
from pandas.io import sql
home_work=pd.read_sql_table('home_work', engine)
#df=home_work.iloc[751:]
df=home_work
# +
from time import sleep
count=0
for index, row in df.iterrows():
#print row['home_lat'], row['home_lon'], row['work_lat'], row['work_lon']
source_lat = row['home_lat']
source_lon = row['home_lon']
dest_lat= row['work_lat']
dest_lon= row['work_lon']
headers = {
'key':'<KEY>'
}
params = {
# Request parameters
#'origin': source_lat+','+source_lon,
#'destination':dest_lat+','+dest_lon,
'origin': dest_lat+','+dest_lon,
'destination':source_lat+','+source_lon,
'alternatives':'false',
'mode':'car',
'traffic_model':'optimistic',
'departure_time':'now'
}
r = requests.get('https://maps.googleapis.com/maps/api/directions/json?', params=params, headers=headers)
if len(r.json()['routes']) != 0:
if len(json_normalize(r.json()['routes'][0]['legs'][0])) != 0:
df=json_normalize(r.json()['routes'][0]['legs'][0])
df1=df.drop(['steps','traffic_speed_entry','via_waypoint'], axis=1)
df1['unique_id']=source_lat+'_'+source_lon+'_'+dest_lat+'_'+dest_lon
df1.to_sql('car_info1', engine,if_exists='append')
else:
continue
else:
continue
count += 1
print 'Latitude and longitude count value - ',count
print 'Source and destination pair - ', source_lat, source_lon, dest_lat, dest_lon
print "Successful data collection!"
# -
| python_code/.ipynb_checkpoints/googleApiCars-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PaddlePaddle 2.1.0 (Python 3.5)
# language: python
# name: py35-paddle1.2.0
# ---
# # **iFLYTEK叶菜病虫害图像识别挑战赛**
#
# ## **赛事链接:** http://challenge.xfyun.cn/topic/info?type=pests-diseases
#
# ## **目前排名:**
#
#
# ## **一、赛事背景**
# 农作物病虫害严重制约着农业生产,因为农作物病虫害种类多、密度大,极易造成农作物大量减产。同时由于传统人眼识别病虫害的方法速度较慢、准确度较低,会导致农药的滥用,破坏自然环境。如今随着精准农业和智慧农业概念的兴起和发展,利用信息技术辅助农业生产,实现对农作物病虫害的智能识别和检测,以减少不必要的农药喷施,对保护生态系统均衡,保障农作物安全生产,提高农作物的质量方面,有着十分重要的促进作用。
#
# ## **二、赛事任务**
# 最为有效的病虫害识别方法是图片识别,本次大赛提供了大量农民在田间地头拍摄的叶菜的病虫害图片,参赛选手需基于提供的样本构建模型,实现叶菜的病虫害图像识别,即为图片分类。
#
# ## **三、评审规则**
#
# 评估指标
# 本模型依据提交的结果文件,采用准确率进行评价。
#
# 
#
# ## **四、数据说明**
#
# 1.数据说明
#
#
# 本次比赛为参赛选手提供了叶菜的病虫害图像数据:包括图像及其所属病虫害标签。数据主体为农民在不同环境条件下拍摄的叶菜农作物图像,每张图像的主体突出度,背景复杂程度、光照条件,图像清晰度均存在一定差别。图片已按类别放在不同文件夹内,文件夹名称即为图片的category_id。
#
# 1:用药不当
#
#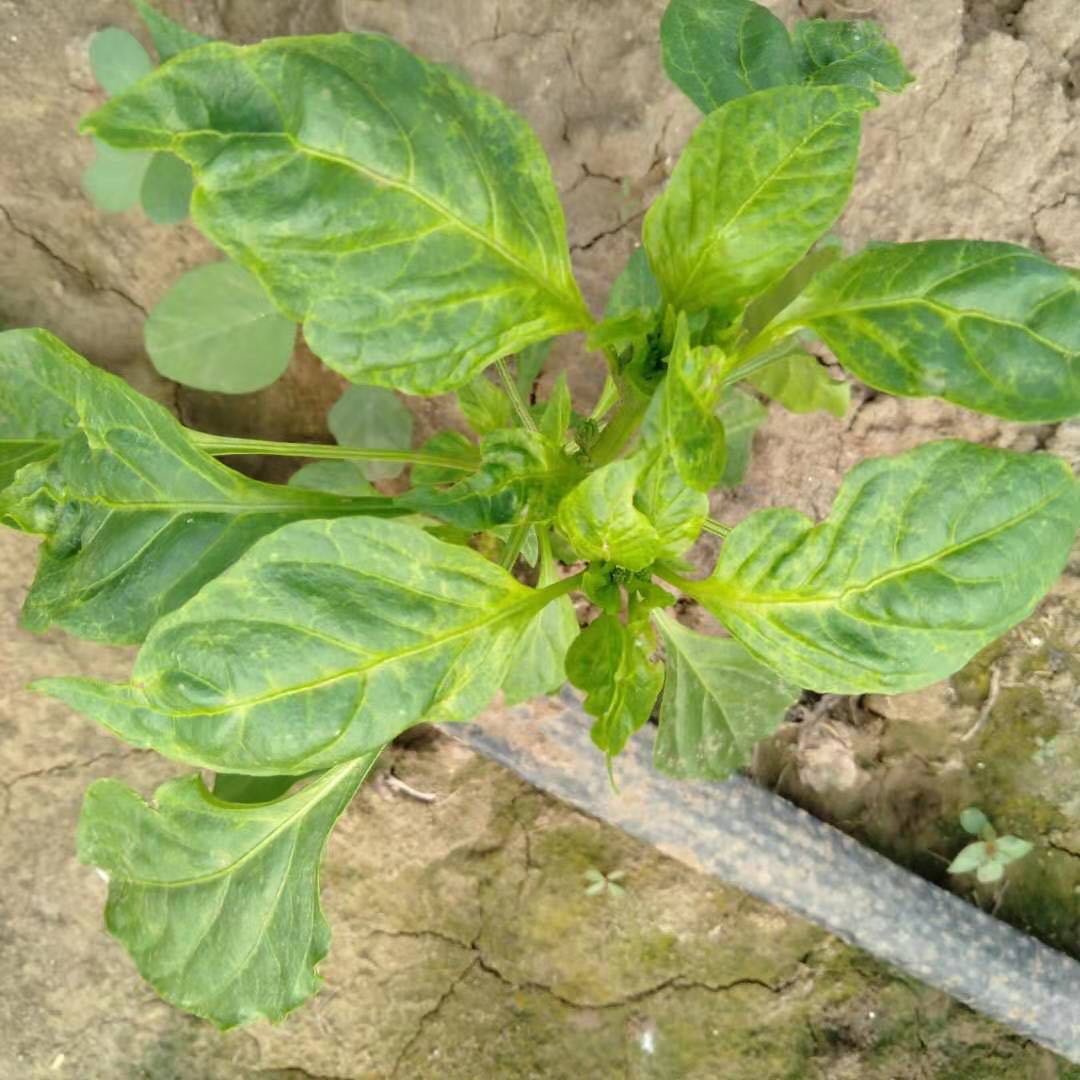 
#
#
# 2:疫病
#
#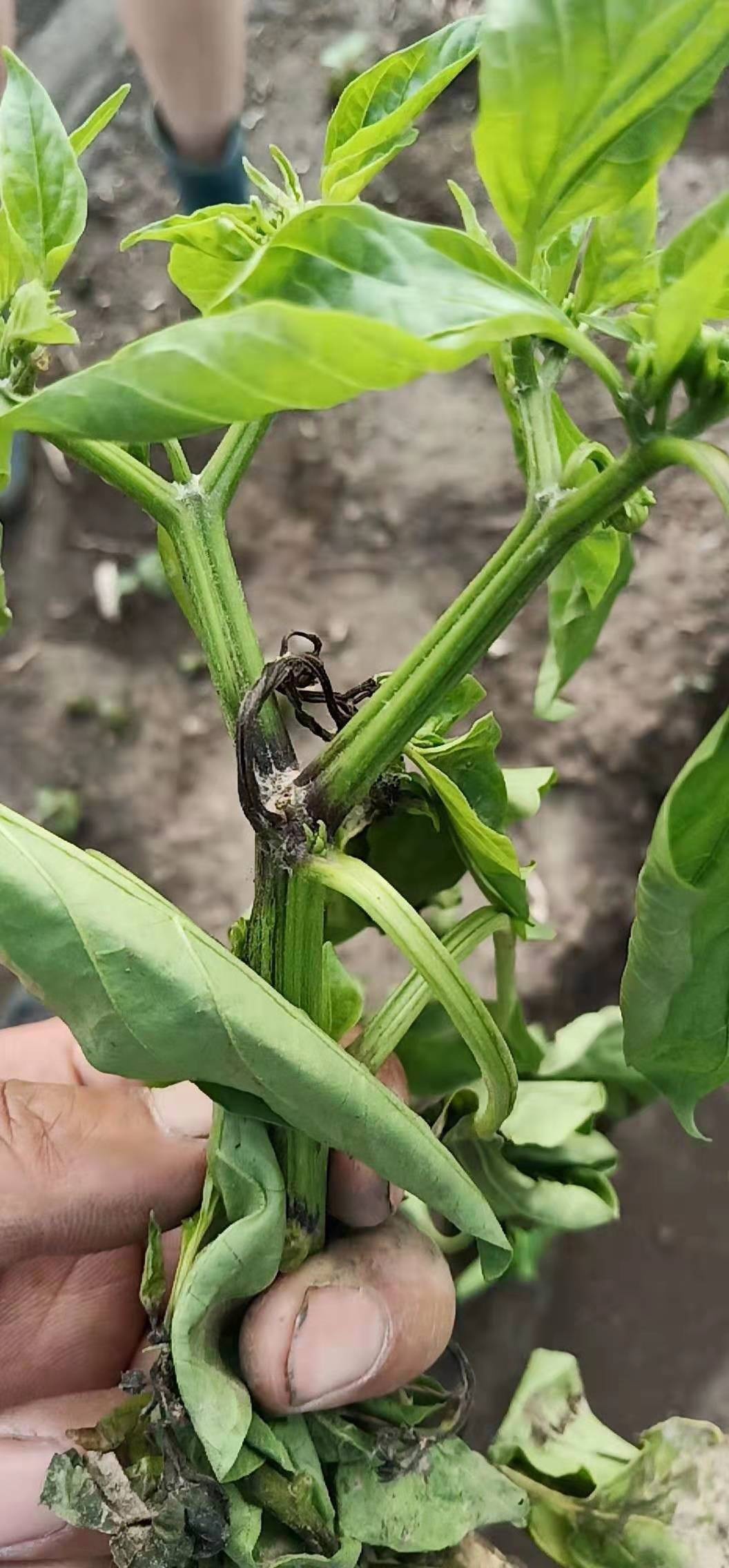 
#
#
# 3:炭疽病
#
# 
#
#
#
# 本次比赛为参赛选手提供的数据分为训练集、测试集、提交样例三类文件:
#
# 训练集:包含1000多张。
#
# 测试集:包含300张,图片文件的名称即为image_id。
#
# 提交样例:表头为image_id和category_id的CSV文件,选手提交数据时需要将测试集的图片id与模型预测的类别id按样例格式填入CSV中,进行提交。
#
# # **解题思路:**
#
# 1.首先做一些数据的统计以及数据探索
#
# 2.然后进行选择合适的模型进行构造baseline
#
# 3.准确无误输出结果并对其进行提交成功
#
# 4.后期对模型进行修改
#
# 5.对模型进行融合
#
# # **一、解压数据**
#
# 首先解压压缩包,然后把里面的train.csv文件复制出来,然后把所有类别的图片都放在train这个文件夹下面
# !unzip data/data98942/leaf.zip
# !cp trian1/train2/train.csv -d ./
# !cp trian1/train2/1/* -d ./train/
# !cp trian1/train2/2/* -d ./train/
# !cp trian1/train2/3/* -d ./train/
# ## **数据EDA**
#
# 探索性数据分析(Exploratory Data Analysis,简称EDA),是指对已有的数据(原始数据)进行分析探索,通过作图、制表、方程拟合、计算特征量等手段探索数据的结构和规律的一种数据分析方法。一般来说,我们最初接触到数据的时候往往是毫无头绪的,不知道如何下手,这时候探索性数据分析就非常有效。
#
# 对于图像分类任务,我们通常首先应该统计出每个类别的数量,查看训练集的数据分布情况。通过数据分布情况分析赛题,形成解题思路。(洞察数据的本质很重要。)
import pandas as pd
df = pd.read_csv('train.csv')
d=df['label'].hist().get_figure()
d.savefig('1.jpg')
# 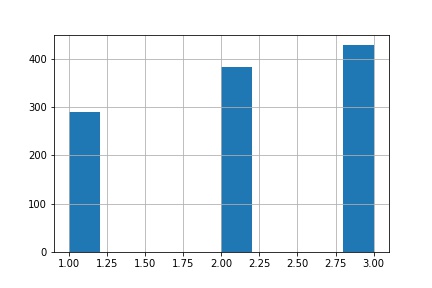
#
# # **二、按比例划分数据集**
#
# **用pandas模块读取train.csv信息,即图像的名称以及对应的标签**
#
# **再按照8:2的比列生成训练集和验证集,并且保存到work/文件夹下**
# +
import pandas as pd
import codecs
import os
from PIL import Image
import numpy as np
import random
df = pd.read_csv('train.csv')
from sklearn.utils import shuffle
df = shuffle(df)#随机打乱顺序
all_file_dir = 'work'
train_file = codecs.open(os.path.join(all_file_dir, "train_list.txt"), 'w')#设置训练集的地址
eval_file = codecs.open(os.path.join(all_file_dir, "eval_list.txt"), 'w')#设置验证集的地址
# df = df.values
# random.shuffle(df)
print(df)
# df.sample(frac=1)
image_path_list = df['image'].values
label_list = df['label'].values
# 划分训练集和校验集
all_size = len(image_path_list)
train_size = int(all_size * 0.8)#设置训练集数目
train_image_path_list = image_path_list[:train_size]
train_label_list = label_list[:train_size]
val_image_path_list = image_path_list[train_size:]
val_label_list = label_list[train_size:]
image_path_pre = 'train'
for file,label_id in zip(train_image_path_list, train_label_list):
# print(file)
# print(label_id)
try:
img = Image.open(os.path.join(image_path_pre, file))
# train_file.write("{0}\0{1}\n".format(os.path.join(image_path_pre, file), label_id))
train_file.write("{0}{1}{2}\n".format(file,' ', label_id))
# eval_file.write("{0}\t{1}\n".format(os.path.join(image_path_pre, file), label_id))
except Exception as e:
pass
# 存在一些文件打不开,此处需要稍作清洗
# print('error!')
for file,label_id in zip(val_image_path_list, val_label_list):
# print(file)
# print(label_id)
try:
img = Image.open(os.path.join(image_path_pre, file))
# train_file.write("{0}\t{1}\n".format(os.path.join(image_path_pre, file), label_id))
eval_file.write("{0}{1}{2}\n".format(file,' ', label_id))
except Exception as e:
# pass
# 存在一些文件打不开,此处需要稍作清洗
# print('error!')
train_file.close()
# #需要稍作清洗
# print('error!')
train_file.close()
eval_file.close()
# -
# 训练集文件
#
# 
#
#
# 验证集文件
#
# 
#
# # **三、解压PaddleClas-release-2.1包**
# !unzip data/data98136/PaddleClas-release-2.1.zip
# ## **下载ResNet50_vd_pretrained预训练权重**
# !mkdir pretrained
# !cd pretrained && wget https://paddle-imagenet-models-name.bj.bcebos.com/dygraph/ResNet50_vd_pretrained.pdparams
# # **四、执行训练**
#
# 本次使用的是使用的ResNet50_vd模块,路径放在configs/ResNet/ResNet50_vd.yaml,下面试对这个配置文件进行说明要更改的地方:
#
# 
#
# pretrained_model: 加载预训练模型,"/home/aistudio/pretrained/ResNet50_vd_pretrained"
#
# model_save_dir: 存放地址,"/home/aistudio/output/"
#
# classes_num: 3,要识别的类别总数
#
# total_images: 1103,照片总数=训练集+验证集
#
# save_interval: 5 隔多少epochs保存一次权重
#
# validate: True 开启验证
#
# valid_interval: 1 每个多少epochs验证一次模型
#
#
# 使用mix数据增强
#
# use_mix: True
#
# ls_epsilon: 0.1
#
# 学习率调整
#
# LEARNING_RATE:
# function: 'Cosine'
#
# params:
# lr: 0.001
#
# 优化器
#
# OPTIMIZER:
#
# function: 'Momentum'
#
# params:
# momentum: 0.9
#
# regularizer:
# function: 'L2'
# factor: 0.000070
#
#
# ...........................
# 
#
# **训练过程**进行数据增强的过程,这里使用了
#
# DecodeImage()
#
# RandCropImage(随机裁剪)
#
# RandFlipImage(随机翻转)
#
# NormalizeImage(归一化)
#
# 还有Mix(照片拼接)
#
# 数据增强还有许多办法,大家可以多多尝试,比如加入随机噪声,对比度的调整等。
#
# **验证过程**则使用了
#
# DecodeImage()
#
# ResizeImage(重新设置照片大小)
#
# RandCropImage(随机裁剪)
#
# NormalizeImage(归一化)
# cd PaddleClas-release-2.1
# !python tools/train.py -c configs/ResNet/ResNet50_vd.yaml
# 2epoch精度就已经0.44344了,准确率还是可以的,后面我就没去再训练了,只是给大家做了一个演示。
# 
#
# ## **Tips:**
#
# 后期训练策略可以是用精度更高的网络文件配置,并且对它进行相对应的调整
#
# 例如调整学习率Lr,以及schedule_LR,还有符合模型的数据增强
#
# 最后可以进行模型融合或者结果融合等。
#
# # **五、模型评估**
# !python -u tools/eval.py -c configs/ResNet/ResNet50_vd.yaml \
# -o weights=output/ResNet50_vd/best_model/ppcls.pdparams
# # **六、进行预测并生成结果文件**
#
# 结果文件为submisson.csv,生成后便可提交。
# !python PaddleClas-release-2.1/tools/infer/infer.py \
# -i test \
# --model ResNet50_vd \
# --pretrained_model "output/ResNet50_vd/best_model/ppcls" \
# --load_static_weights False \
# --class_num=3
# 这里我对PaddleClas-release-2.1/tools/infer/infer.py进行了一点修改
#
# 因为直接用infer.py输出的结果是这种格式 File:{}, Top-{} result: class id(s): {}, score(s): {}。
#
# 我们可以看到我们要提交的文件格式是image_id以及category_id,也就是对应这里的File以及class id。
#
# 所以我在85行插入了如下代码,就是把读取到的文件信息以及其标签信息进行存放,然后利用pandas模块把它制作成表格,形成可提交文件。
#
# 
#
# 
#
# ## **生成结果文件**
#
#
# 
#
#
#
# # **总结**
#
# 针对图像分类比赛,大家在选定了一个baseline之后可以尝试各种技巧,包括学习率调整策略,模型调参等等。关于图像分类竞赛的一些技巧,大家可以去网上搜索相关的trick。
#
# 建议
#
# 刚开始的小白,比赛优先使用简单的模型(如ResNet50),快速跑完整个训练和预测流程。
#
# 要有一定毅力和耐心,不怕失败,比赛过程往往会遇到很多预想不到的问题。另外数据扩增方法一定要反复尝试,会很大程度上影响模型精度。
#
# 后期上分可使用模型融合
# # **关于更多关于PaddleClas的信息请参考下面的链接地址。**
#
# PaddleClas教程文档地址:https://github.com/PaddlePaddle/PaddleClas
#
# PaddleClas Github地址:https://github.com/PaddlePaddle/PaddleClas
#
# # **关于作者**
#
# 感兴趣的方向为:目标检测,图像分类等
#
# AIstudio主页: 我在AI Studio上获得白银等级,点亮3个徽章,来互关呀~ https://aistudio.baidu.com/aistudio/personalcenter/thirdview/474269
#
# Github主页: https://github.com/Niki173
#
# 欢迎大家有问题留言交流学习,共同进步成长。
| 2152212.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="knOigRU1UJ9Y"
# # Estimating Workplace Location
#
# This notebook illustrates how to re-estimate ActivitySim's auto ownership model. The steps in the process are:
# - Run ActivitySim in estimation mode to read household travel survey files, run the ActivitySim submodels to write estimation data bundles (EDB) that contains the model utility specifications, coefficients, chooser data, and alternatives data for each submodel.
# - Read and transform the relevant EDB into the format required by the model estimation package [larch](https://larch.newman.me) and then re-estimate the model coefficients. No changes to the model specification will be made.
# - Update the ActivitySim model coefficients and re-run the model in simulation mode.
#
# The basic estimation workflow is shown below and explained in the next steps.
#
# 
# -
# # Load libraries
import larch # !conda install larch #for estimation
import larch.util.activitysim
import pandas as pd
import numpy as np
import yaml
import larch.util.excel
import os
# # Required Inputs
#
# In addition to a working ActivitySim model setup, estimation mode requires an ActivitySim format household travel survey. An ActivitySim format household travel survey is very similar to ActivitySim's simulation model tables:
#
# - households
# - persons
# - tours
# - joint_tour_participants
# - trips (not yet implemented)
#
# Examples of the ActivitySim format household travel survey are included in the [example_estimation data folders](https://github.com/RSGInc/activitysim/tree/develop/activitysim/examples/example_estimation). The user is responsible for formatting their household travel survey into the appropriate format.
#
# After creating an ActivitySim format household travel survey, the `scripts/infer.py` script is run to append additional calculated fields. An example of an additional calculated field is the `household:joint_tour_frequency`, which is calculated based on the `tours` and `joint_tour_participants` tables.
#
# The input survey files are below.
# ### Survey households
pd.read_csv("../data_sf/survey_data/override_households.csv")
# ### Survey persons
pd.read_csv("../data_sf/survey_data/override_persons.csv")
# ### Survey tours
pd.read_csv("../data_sf/survey_data/override_tours.csv")
# ### Survey joint tour participants
pd.read_csv("../data_sf/survey_data/survey_joint_tour_participants.csv")
# # Example Setup if Needed
#
# To avoid duplication of inputs, especially model settings and expressions, the `example_estimation` depends on the `example`. The following commands create an example setup for use. The location of these example setups (i.e. the folders) are important because the paths are referenced in this notebook. The commands below download the skims.omx for the SF county example from the [activitysim resources repository](https://github.com/RSGInc/activitysim_resources).
# !activitysim create -e example_estimation_sf -d test
# # Run the Estimation Example
#
# The next step is to run the model with an `estimation.yaml` settings file with the following settings in order to output the EDB for all submodels:
#
# ```
# enable=True
#
# bundles:
# - school_location
# - workplace_location
# - auto_ownership
# - free_parking
# - cdap
# - mandatory_tour_frequency
# - mandatory_tour_scheduling
# - joint_tour_frequency
# - joint_tour_composition
# - joint_tour_participation
# - joint_tour_destination
# - joint_tour_scheduling
# - non_mandatory_tour_frequency
# - non_mandatory_tour_destination
# - non_mandatory_tour_scheduling
# - tour_mode_choice
# - atwork_subtour_frequency
# - atwork_subtour_destination
# - atwork_subtour_scheduling
# - atwork_subtour_mode_choice
# survey_tables:
# households:
# file_name: survey_data/override_households.csv
# index_col: household_id
# persons:
# file_name: survey_data/override_persons.csv
# index_col: person_id
# tours:
# file_name: survey_data/override_tours.csv
# joint_tour_participants:
# file_name: survey_data/override_joint_tour_participants.csv
# ```
#
# This enables the estimation mode functionality, identifies which models to run and their output estimation data bundles (EDBs), and the input survey tables, which include the override settings for each model choice.
#
# With this setup, the model will output an EBD with the following tables for this submodel:
# - model settings - workplace_location_model_settings.yaml
# - coefficients - workplace_location_coefficients.csv
# - utilities specification - workplace_location_SPEC.csv
# - land use data - workplace_location_landuse.csv
# - size terms - workplace_location_size_terms.csv
# - alternatives values - workplace_location_alternatives_combined.csv
# - chooser data - workplace_location_choosers_combined.csv
#
# The following code runs the software in estimation mode, inheriting the settings from the simulation setup and using the San Francisco county data setup. It produces the EDB for all submodels but runs all the model steps identified in the inherited settings file.
# %cd test
# !activitysim run -c configs_estimation/configs -c configs -o output -d data_sf
# # Read EDB
#
# The next step is to read the EDB, including the coefficients, model settings, utilities specification, and chooser and alternative data.
# +
edb_directory = "output/estimation_data_bundle/workplace_location/"
def read_csv(filename, **kwargs):
return pd.read_csv(os.path.join(edb_directory, filename), **kwargs)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="cBqPPkBpnaUZ" outputId="bd780019-c200-4cf6-844a-991c4d026480"
coefficients = read_csv("workplace_location_coefficients.csv", index_col='coefficient_name')
spec = read_csv("workplace_location_SPEC.csv")
alt_values = read_csv("workplace_location_alternatives_combined.csv")
chooser_data = read_csv("workplace_location_choosers_combined.csv")
landuse = read_csv("workplace_location_landuse.csv", index_col='TAZ')
size_spec = read_csv("workplace_location_size_terms.csv")
# -
# ### Zone size term specification
work_size_spec = size_spec \
.query("model_selector == 'workplace'") \
.drop(columns='model_selector') \
.set_index('segment')
work_size_spec = work_size_spec.loc[:,work_size_spec.max()>0]
work_size_spec
# ### Zone size term coefficients
size_coef = work_size_spec.stack().reset_index()
size_coef.index = size_coef.iloc[:,0] +"_"+ size_coef.iloc[:,1]
size_coef = size_coef.loc[size_coef.iloc[:,2]>0]
size_coef['constrain'] = 'F'
one_each = size_coef.groupby('segment').first().reset_index()
size_coef.loc[one_each.iloc[:,0] +"_"+ one_each.iloc[:,1], 'constrain'] = 'T'
size_coef = size_coef.iloc[:,2:]
size_coef.columns = ['value','constrain']
size_coef.index.name = 'coefficient_name'
size_coef['value'] = np.log(size_coef['value'])
size_coef
# ### Model settings
settings = yaml.load(
open(os.path.join(edb_directory,"workplace_location_model_settings.yaml"),"r"),
Loader=yaml.SafeLoader,
)
settings
# ### Coefficients
coefficients
# ### Utility specification
spec
# ### Remove shadow pricing and pre-existing size expression for re-estimation
spec = spec\
.set_index('Label')\
.drop(index=['util_size_variable', 'util_utility_adjustment'])\
.reset_index()
# ### Alternatives data
alt_values
# ### Chooser data
chooser_data
# # Data Processing and Estimation Setup
#
# The next step is to transform the EDB for larch for model re-estimation.
from larch import P, X
# ### Utility specifications
m = larch.Model()
m.utility_ca = larch.util.activitysim.linear_utility_from_spec(
spec, x_col='Label', p_col='coefficient',
ignore_x=('local_dist',),
)
print(m.utility_ca)
m.quantity_ca = sum(
P(f"{i}_{q}") * X(q) * X(f"income_segment=={settings['SEGMENT_IDS'][i]}")
for i in work_size_spec.index
for q in work_size_spec.columns
)
larch.util.activitysim.apply_coefficients(coefficients, m)
larch.util.activitysim.apply_coefficients(size_coef, m, minimum=-6, maximum=6)
# ### Coefficients
m.pf
x_co = chooser_data.set_index('person_id').rename(columns={'TAZ':'HOMETAZ'})
x_ca = larch.util.activitysim.cv_to_ca(
alt_values.set_index(['person_id', 'variable'])
)
# ### Remove choosers with invalid observed choice
workplace_tazs = landuse[landuse['TOTEMP'] > 0].index
x_co = x_co[x_co['override_choice'].isin(workplace_tazs)]
x_ca = x_ca[x_ca.index.get_level_values('person_id').isin(x_co.index)]
x_ca_1 = pd.merge(x_ca, landuse, on='TAZ', how='left')
x_ca_1.index = x_ca.index
# ### Availability
av = x_ca_1['util_no_attractions'].apply(lambda x: False if x == 1 else True)
d = larch.DataFrames(
co=x_co,
ca=x_ca_1,
av=av,
)
m.dataservice = d
# ### Survey choice
m.choice_co_code = 'override_choice'
# # Estimate
#
# With the model setup for estimation, the next step is to estimate the model coefficients. Make sure to use a sufficiently large enough household sample and set of zones to avoid an over-specified model, which does not have a numerically stable likelihood maximizing solution. Larch has two built-in estimation methods: BHHH and SLSQP. BHHH is the default and typically runs faster, but does not follow constraints on parameters. SLSQP is safer, but slower, and may need additional iterations.
# m.estimate(method='SLSQP', options={'maxiter':1000})
m.estimate(method='BHHH', options={'maxiter':1000})
# ### Estimated coefficients
m.parameter_summary()
# + [markdown] colab_type="text" id="TojXWivZsx7M"
# # Output Estimation Results
# -
est_names = [j for j in coefficients.index if j in m.pf.index]
coefficients.loc[est_names,'value'] = m.pf.loc[est_names, 'value']
os.makedirs(os.path.join(edb_directory,'estimated'), exist_ok=True)
# ### Write the re-estimated coefficients file
coefficients.reset_index().to_csv(
os.path.join(edb_directory,'estimated',"workplace_location_coefficients_revised.csv"),
index=False,
)
# ### Write the model estimation report, including coefficient t-statistic and log likelihood
m.to_xlsx(
os.path.join(edb_directory,'estimated',"workplace_location_model_estimation.xlsx"), data_statistics=False
)
# Write size coefficients into size_spec
for c in work_size_spec.columns:
for i in work_size_spec.index:
param_name = f"{i}_{c}"
j = (size_spec['segment'] == i) & (size_spec['model_selector'] == 'workplace')
size_spec.loc[j,c] = np.exp(m.get_value(param_name))
# Rescale each row to total 1, not mathematically needed
# but to maintain a consistent approach from existing ASim
size_spec.iloc[:,2:] = (size_spec.iloc[:,2:].div(size_spec.iloc[:,2:].sum(1), axis=0))
# ### Write updated size coefficients
size_spec.to_csv(
os.path.join(edb_directory,'estimated',"workplace_location_size_terms.csv"),
index=False,
)
# # Next Steps
#
# The final step is to either manually or automatically copy the `workplace_location_coefficients_revised.csv` file and `workplace_location_size_terms.csv` file to the configs folder, rename them to `workplace_location_coeffs.csv` and `destination_choice_size_terms.csv`, and run ActivitySim in simulation mode.
pd.read_csv(os.path.join(edb_directory,'estimated',"workplace_location_coefficients_revised.csv"))
pd.read_csv(os.path.join(edb_directory,'estimated',"workplace_location_size_terms.csv"))
| activitysim/examples/example_estimation/notebooks/estimating_workplace_location.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Stateful Elasticsearch Feedback Workflow for Metrics Server
# In this example we will add statistical performance metrics capabilities by leveraging the Seldon metrics server with persistence through the elasticsearch setup.
#
# Dependencies
# * Seldon Core installed
# * Ingress provider (Istio or Ambassador)
# * Install [Elasticsearch for the Seldon Core Logging](https://docs.seldon.io/projects/seldon-core/en/latest/analytics/logging.html)
# * KNative eventing v0.18.3
# * KNative serving v0.18.1 (optional)
#
# See the centralized logging example (also in the examples directory) for how to set these up.
#
# Easiest way is to run `examples/centralized-logging/full-kind-setup.sh` and then:
# `helm delete seldon-core-loadtesting`
# `seldon-single-model`
#
# Then port-forward to that ingress on localhost:8080 in a separate terminal either of (istio suggested):
#
# Ambassador:
#
# kubectl port-forward $(kubectl get pods -n seldon -l app.kubernetes.io/name=ambassador -o jsonpath='{.items[0].metadata.name}') -n seldon 8080:8080
#
# Istio:
#
# kubectl port-forward -n istio-system svc/istio-ingressgateway 8080:80
#
#
#
# %%writefile requirements-dev.txt
elasticsearch==7.9.1
# !pip install -r requirements-dev.txt
# !kubectl create namespace seldon || echo "namespace already created"
# !kubectl create namespace seldon-logs || echo "namespace already created"
# !kubectl config set-context $(kubectl config current-context) --namespace=seldon
# !mkdir -p config
# Setting up Knative eventing routing for request logger
# + language="bash"
# kubectl apply -f - <<EOF
# apiVersion: eventing.knative.dev/v1
# kind: Broker
# metadata:
# name: default
# namespace: seldon-logs
# EOF
# -
# Verify broker is up
# + language="bash"
# kubectl -n seldon-logs get broker default -o jsonpath='{.status.address.url}'
# -
# Adding payload request logger component for redirection of logs
# + language="bash"
# kubectl apply -f - << END
# apiVersion: eventing.knative.dev/v1
# kind: Trigger
# metadata:
# name: seldon-request-logger-trigger
# namespace: seldon-logs
# spec:
# broker: default
# subscriber:
# ref:
# apiVersion: serving.knative.dev/v1
# kind: Service
# name: seldon-request-logger
# END
#
# kubectl apply -f - << END
# apiVersion: serving.knative.dev/v1
# kind: Service
# metadata:
# name: seldon-request-logger
# namespace: seldon-logs
# metadata:
# labels:
# fluentd: "true"
# spec:
# template:
# metadata:
# annotations:
# autoscaling.knative.dev/minScale: "1"
# spec:
# containers:
# - image: docker.io/seldonio/seldon-request-logger:1.5.1
# imagePullPolicy: Always
# env:
# - name: ELASTICSEARCH_HOST
# value: "elasticsearch-opendistro-es-client-service.seldon-logs.svc.cluster.local"
# - name: ELASTICSEARCH_PORT
# value: "9200"
# - name: ELASTICSEARCH_PROTOCOL
# value: "https"
# - name: ELASTICSEARCH_USER
# value: "admin"
# - name: ELASTICSEARCH_PASS
# value: "admin"
# END
# -
# ### Create a simple model
# We create a multiclass classification model - iris classifier.
#
# The iris classifier takes an input array, and returns the prediction of the 4 classes.
#
# The prediction can be done as numeric or as a probability array.
# + language="bash"
# kubectl apply -f - << END
# apiVersion: machinelearning.seldon.io/v1
# kind: SeldonDeployment
# metadata:
# name: multiclass-model
# spec:
# predictors:
# - graph:
# children: []
# implementation: SKLEARN_SERVER
# modelUri: gs://seldon-models/sklearn/iris
# name: classifier
# logger:
# url: http://broker-ingress.knative-eventing.svc.cluster.local/seldon-logs/default
# mode: all
# name: default
# replicas: 1
# END
# -
# !kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=multiclass-model -o jsonpath='{.items[0].metadata.name}')
# ### Send Prediction Request
import requests
url = "http://localhost:8080/seldon/seldon/multiclass-model/api/v1.0"
pred_req_1 = {"data":{"ndarray":[[1,2,3,4]]}}
pred_resp_1 = requests.post(f"{url}/predictions", json=pred_req_1)
print(pred_resp_1.json())
assert(len(pred_resp_1.json()["data"]["ndarray"][0])==3)
# ### Check data in Elasticsearch
# We'll be able to check the elasticsearch through the service or the pods in our cluster.
#
# To do this we'll have to port-forward to elastic in another window. e.g.
#
# kubectl port-forward -n seldon-logs svc/elasticsearch-opendistro-es-client-service 9200
#
# Verify by going to https://admin:admin@localhost:9200/_cat/indices
from elasticsearch import Elasticsearch
es = Elasticsearch(['https://admin:admin@localhost:9200'],verify_certs=False)
# See the indices that have been created
es.indices.get_alias("*")
# Look at the data that is stored in the elasticsearch index
res = es.search(index="inference-log-seldon-seldon-multiclass-model-default", body={"query": {"match_all": {}}})
print("Logged Request:")
print(res["hits"]["hits"][0]["_source"]["request"])
print("\nLogged Response:")
print(res["hits"]["hits"][0]["_source"]["response"])
# ### Send feedback
#
# We can now send the correction, or the truth value of the prediction.
#
# For this we'll need to send the UUID of the feedback request to ensure it's added to the correct index.
# +
puid_seldon_1 = pred_resp_1.headers.get("seldon-puid")
print(puid_seldon_1)
# -
# We can also be able to add extra metadata, such as the user providing the feedback, date, time, etc.
feedback_tags_1 = {
"user": "<NAME>",
"date": "11/07/2020"
}
# And finally we can put together the feedback request.
feedback_req_1 = {
"reward": 0,
"truth": {
'data': {
'names': ['t:0', 't:1', 't:2'],
'ndarray': [[0, 0, 1]]
},
"meta": {
"tags": feedback_tags_1
}
}
}
# And send the feedback request
feedback_resp_1 = requests.post(f"{url}/feedback", json=feedback_req_1, headers={"seldon-puid": puid_seldon_1})
print(feedback_resp_1)
# Check that feedback has been received and stored in the Elasticsearch index
# +
res = es.search(index="inference-log-seldon-seldon-multiclass-model-default", body={"query": {"match_all": {}}})
print(res["hits"]["hits"][-1]["_source"]["feedback"])
# -
# ### Deploying Metrics Server
#
# Now we'll be able to see how the metrics server makes use of this infrastructure patterns to provide real time performance metrics.
# %%writefile config/multiclass-deployment.yaml
apiVersion: apps/v1
kind: Deployment
metadata:
name: seldon-multiclass-model-metrics
namespace: seldon-logs
labels:
app: seldon-multiclass-model-metrics
spec:
replicas: 1
selector:
matchLabels:
app: seldon-multiclass-model-metrics
template:
metadata:
annotations:
prometheus.io/path: /v1/metrics
prometheus.io/scrape: "true"
labels:
app: seldon-multiclass-model-metrics
spec:
securityContext:
runAsUser: 8888
containers:
- name: user-container
image: seldonio/alibi-detect-server:1.7.0-dev
imagePullPolicy: IfNotPresent
args:
- --model_name
- multiclassserver
- --http_port
- '8080'
- --protocol
- seldonfeedback.http
- --storage_uri
- "adserver.cm_models.multiclass_one_hot.MulticlassOneHot"
- --reply_url
- http://message-dumper.default
- --event_type
- io.seldon.serving.feedback.metrics
- --event_source
- io.seldon.serving.feedback
- --elasticsearch_uri
- https://admin:admin@elasticsearch-opendistro-es-client-service.seldon-logs:9200
- MetricsServer
env:
- name: "SELDON_DEPLOYMENT_ID"
value: "multiclass-model"
- name: "PREDICTIVE_UNIT_ID"
value: "classifier"
- name: "PREDICTIVE_UNIT_IMAGE"
value: "alibi-detect-server:1.7.0-dev"
- name: "PREDICTOR_ID"
value: "default"
ports:
- containerPort: 8080
name: metrics
protocol: TCP
---
apiVersion: v1
kind: Service
metadata:
name: seldon-multiclass-model-metrics
namespace: seldon-logs
labels:
app: seldon-multiclass-model-metrics
spec:
selector:
app: seldon-multiclass-model-metrics
ports:
- protocol: TCP
port: 80
targetPort: 8080
# !kubectl apply -f config/multiclass-deployment.yaml
# !kubectl rollout status -n seldon-logs deploy/seldon-multiclass-model-metrics
# ### Trigger for metrics server
#
# The trigger will be created in the seldon-logs namespace as that is where the initial trigger will be sent to.
# + language="bash"
#
# kubectl apply -f - << END
# apiVersion: eventing.knative.dev/v1
# kind: Trigger
# metadata:
# name: multiclass-model-metrics-trigger
# namespace: seldon-logs
# spec:
# broker: default
# filter:
# attributes:
# inferenceservicename: multiclass-model
# type: io.seldon.serving.feedback
# subscriber:
# ref:
# apiVersion: v1
# kind: Service
# name: seldon-multiclass-model-metrics
# END
# -
import time
time.sleep(20)
# ### (Alternative) create kservice
#
# If you want to create a kservice, and you've installed knative eventing and knative serving, you can use the instructions below.
#
# The value of the file `config/multiclass-service.yaml` would be:
# ```
# apiVersion: serving.knative.dev/v1alpha1
# kind: Service
# metadata:
# name: seldon-multiclass-model-metrics
# namespace: seldon-logs
# spec:
# template:
# metadata:
# annotations:
# prometheus.io/path: /v1/metrics
# prometheus.io/scrape: "true"
# autoscaling.knative.dev/minScale: "1"
# spec:
# containers:
# - image: "seldonio/alibi-detect-server:1.7.0-dev"
# args:
# - --model_name
# - multiclassserver
# - --http_port
# - '8080'
# - --protocol
# - seldonfeedback.http
# - --storage_uri
# - "adserver.cm_models.multiclass_one_hot.MulticlassOneHot"
# - --reply_url
# - http://message-dumper.default
# - --event_type
# - io.seldon.serving.feedback.metrics
# - --event_source
# - io.seldon.serving.feedback
# - MetricsServer
# env:
# - name: "SELDON_DEPLOYMENT_ID"
# value: "multiclass-model"
# - name: "PREDICTIVE_UNIT_ID"
# value: "classifier"
# - name: "PREDICTIVE_UNIT_IMAGE"
# value: "alibi-detect-server:1.7.0-dev"
# - name: "PREDICTOR_ID"
# value: "default"
# ports:
# - containerPort: 8080
# name: metrics
# protocol: TCP
# securityContext:
# runAsUser: 8888
# ```
#
# You can run the kservice with the command below:
# ```
# kubectl apply -f config/multiclass-service.yaml
# ```
# And then check with:
#
# ```
# kubectl get kservice
# ```
#
# You'll then have to create the trigger, first by creating the broker:
#
# ```bash
# # %%bash
# kubectl apply -f - <<EOF
# apiVersion: eventing.knative.dev/v1
# kind: Broker
# metadata:
# name: default
# namespace: seldon-logs
# EOF
# ```
#
# And then the trigger contents:
#
# ```
# apiVersion: eventing.knative.dev/v1
# kind: Trigger
# metadata:
# name: multiclass-model-metrics-trigger
# namespace: seldon-logs
# spec:
# broker: default
# filter:
# attributes:
# inferenceservicename: multiclass-model
# type: io.seldon.serving.feedback
# subscriber:
# uri: http://seldon-multiclass-model-metrics.seldon-logs:80
# ```
#
# And you can run it with:
# ```
# kubectl apply -f config/trigger.yaml
# ```
# ### Confirm empty metrics
# !kubectl run --quiet=true -it --rm curl --image=radial/busyboxplus:curl --restart=Never -- \
# curl -v -X GET "http://seldon-multiclass-model-metrics.seldon-logs.svc.cluster.local:80/v1/metrics"
# ### Send Feedback
feedback_resp_1 = requests.post(f"{url}/feedback", json=feedback_req_1, headers={"seldon-puid": puid_seldon_1})
print(feedback_resp_1)
# ### Confirm metrics available
# !kubectl run --quiet=true -it --rm curl --image=radial/busyboxplus:curl --restart=Never -- \
# curl -v -X GET "http://seldon-multiclass-model-metrics.seldon-logs.svc.cluster.local:80/v1/metrics"
# ## Forward Grafana Dashboard
# Now we should be able to access the grafana dashboard for the model. You can access it by port-forwarding the grafana dashboard with:
# ```
# kubectl port-forward -n seldon-system svc/seldon-core-analytics-grafana 7000:80
# ```
#
# And you can load the following dashboard via the import tool:
{"annotations": {"list": [{"builtIn": 1,"datasource": "-- Grafana --","enable": true,"hide": true,"iconColor": "rgba(0, 211, 255, 1)","name": "Annotations & Alerts","type": "dashboard"}]},"editable": true,"gnetId": null,"graphTooltip": 0,"id": 4,"links": [],"panels": [{"datasource": "prometheus","description": "","fieldConfig": {"defaults": {"custom": {},"mappings": [],"thresholds": {"mode": "absolute","steps": [{"color": "green","value": null},{"color": "red","value": 80}]}},"overrides": []},"gridPos": {"h": 5,"w": 8,"x": 0,"y": 0},"id": 5,"options": {"colorMode": "background","graphMode": "area","justifyMode": "auto","orientation": "auto","reduceOptions": {"calcs": ["mean"],"fields": "","values": false}},"pluginVersion": "7.0.3","targets": [{"expr": "((sum(seldon_metric_true_positive_total) by (seldon_deployment_name) + sum(seldon_metric_true_negative_total) by (seldon_deployment_name)) / (sum(seldon_metric_true_positive_total) by (seldon_deployment_name) + sum(seldon_metric_false_positive_total) by (seldon_deployment_name) + sum(seldon_metric_false_negative_total) by (seldon_deployment_name))) ","format": "time_series","instant": false,"interval": "","legendFormat": "{{class}} - {{seldon_app}}","refId": "A"}],"timeFrom": null,"timeShift": null,"title": "ACCURACY","transparent": true,"type": "stat"},{"datasource": "prometheus","description": "","fieldConfig": {"defaults": {"custom": {},"mappings": [],"thresholds": {"mode": "absolute","steps": [{"color": "green","value": null},{"color": "red","value": 80}]}},"overrides": []},"gridPos": {"h": 5,"w": 8,"x": 8,"y": 0},"id": 6,"options": {"colorMode": "value","graphMode": "area","justifyMode": "auto","orientation": "auto","reduceOptions": {"calcs": ["mean"],"fields": "","values": false}},"pluginVersion": "7.0.3","targets": [{"expr": "(sum(seldon_metric_true_positive_total) by (seldon_deployment_name) / (sum(seldon_metric_true_positive_total) by (seldon_deployment_name) + sum(seldon_metric_false_positive_total) by (seldon_deployment_name))) ","instant": false,"interval": "","legendFormat": "{{class}} - {{seldon_deployment_name}}","refId": "A"}],"timeFrom": null,"timeShift": null,"title": "PRECISION","transparent": true,"type": "stat"},{"datasource": "prometheus","description": "","fieldConfig": {"defaults": {"custom": {},"mappings": [],"thresholds": {"mode": "absolute","steps": [{"color": "green","value": null},{"color": "red","value": 80}]}},"overrides": []},"gridPos": {"h": 5,"w": 8,"x": 16,"y": 0},"id": 7,"options": {"colorMode": "value","graphMode": "area","justifyMode": "auto","orientation": "auto","reduceOptions": {"calcs": ["mean"],"fields": "","values": false}},"pluginVersion": "7.0.3","targets": [{"expr": "(sum(seldon_metric_true_positive_total) by (seldon_deployment_name) / (sum(seldon_metric_true_positive_total) by (seldon_deployment_name) + sum(seldon_metric_false_negative_total) by (seldon_deployment_name))) ","instant": false,"interval": "","legendFormat": "{{class}} - {{seldon_app}}","refId": "A"}],"timeFrom": null,"timeShift": null,"title": "Recall","transparent": true,"type": "stat"},{"aliasColors": {},"bars": false,"dashLength": 10,"dashes": false,"datasource": "prometheus","description": "","fieldConfig": {"defaults": {"custom": {}},"overrides": []},"fill": 1,"fillGradient": 0,"gridPos": {"h": 6,"w": 24,"x": 0,"y": 5},"hiddenSeries": false,"id": 3,"legend": {"avg": false,"current": false,"max": false,"min": false,"show": true,"total": false,"values": false},"lines": true,"linewidth": 1,"nullPointMode": "null","options": {"dataLinks": []},"percentage": false,"pointradius": 2,"points": true,"renderer": "flot","seriesOverrides": [],"spaceLength": 10,"stack": false,"steppedLine": false,"targets": [{"expr": "((sum(seldon_metric_true_positive_total) by (class, seldon_deployment_name) + sum(seldon_metric_true_negative_total) by (class, seldon_deployment_name)) / (sum(seldon_metric_true_positive_total) by (class, seldon_deployment_name) + sum(seldon_metric_false_positive_total) by (class, seldon_deployment_name) + sum(seldon_metric_false_negative_total) by (class, seldon_deployment_name))) ","interval": "","legendFormat": "{{class}} - {{seldon_deployment_name}}","refId": "A"}],"thresholds": [],"timeFrom": null,"timeRegions": [],"timeShift": null,"title": "Real Time Model ACCURACY by Class","tooltip": {"shared": true,"sort": 0,"value_type": "individual"},"transparent": true,"type": "graph","xaxis": {"buckets": null,"mode": "time","name": null,"show": true,"values": []},"yaxes": [{"decimals": null,"format": "short","label": null,"logBase": 1,"max": "1","min": "0","show": true},{"format": "short","label": null,"logBase": 1,"max": null,"min": null,"show": true}],"yaxis": {"align": false,"alignLevel": null}},{"aliasColors": {},"bars": false,"dashLength": 10,"dashes": false,"datasource": "prometheus","description": "","fieldConfig": {"defaults": {"custom": {}},"overrides": []},"fill": 1,"fillGradient": 0,"gridPos": {"h": 7,"w": 12,"x": 0,"y": 11},"hiddenSeries": false,"id": 2,"legend": {"avg": false,"current": false,"max": false,"min": false,"show": true,"total": false,"values": false},"lines": true,"linewidth": 1,"nullPointMode": "null","options": {"dataLinks": []},"percentage": false,"pointradius": 2,"points": true,"renderer": "flot","seriesOverrides": [],"spaceLength": 10,"stack": false,"steppedLine": false,"targets": [{"expr": "((sum(seldon_metric_true_positive_total) by (class, seldon_deployment_name)) / (sum(seldon_metric_true_positive_total) by (class, seldon_deployment_name) + sum(seldon_metric_false_positive_total) by (class, seldon_deployment_name))) ","interval": "","legendFormat": "{{class}} - {{seldon_deployment_name}}","refId": "A"}],"thresholds": [],"timeFrom": null,"timeRegions": [],"timeShift": null,"title": "Real Time Model PRECISION by Class","tooltip": {"shared": true,"sort": 0,"value_type": "individual"},"type": "graph","xaxis": {"buckets": null,"mode": "time","name": null,"show": true,"values": []},"yaxes": [{"decimals": null,"format": "short","label": null,"logBase": 1,"max": "1","min": "0","show": true},{"format": "short","label": null,"logBase": 1,"max": null,"min": null,"show": true}],"yaxis": {"align": false,"alignLevel": null}},{"aliasColors": {},"bars": false,"dashLength": 10,"dashes": false,"datasource": "prometheus","description": "","fieldConfig": {"defaults": {"custom": {}},"overrides": []},"fill": 1,"fillGradient": 0,"gridPos": {"h": 7,"w": 12,"x": 12,"y": 11},"hiddenSeries": false,"id": 4,"legend": {"avg": false,"current": false,"max": false,"min": false,"show": true,"total": false,"values": false},"lines": true,"linewidth": 1,"nullPointMode": "null","options": {"dataLinks": []},"percentage": false,"pointradius": 2,"points": true,"renderer": "flot","seriesOverrides": [],"spaceLength": 10,"stack": false,"steppedLine": false,"targets": [{"expr": "((sum(seldon_metric_true_negative_total) by (class, seldon_deployment_name)) / (sum(seldon_metric_true_positive_total) by (class, seldon_deployment_name) + sum(seldon_metric_false_negative_total) by (class, seldon_deployment_name))) ","interval": "","legendFormat": "{{class}} - {{seldon_deployment_name}}","refId": "A"}],"thresholds": [],"timeFrom": null,"timeRegions": [],"timeShift": null,"title": "Real Time Model RECALL by Class","tooltip": {"shared": true,"sort": 0,"value_type": "individual"},"type": "graph","xaxis": {"buckets": null,"mode": "time","name": null,"show": true,"values": []},"yaxes": [{"decimals": null,"format": "short","label": null,"logBase": 1,"max": "1","min": "0","show": true},{"format": "short","label": null,"logBase": 1,"max": null,"min": null,"show": true}],"yaxis": {"align": false,"alignLevel": null}}],"refresh": false,"schemaVersion": 25,"style": "dark","tags": [],"templating": {"list": []},"time": {"from": "2020-11-03T12:06:17.311Z","to": "2020-11-03T13:06:17.315Z"},"timepicker": {"refresh_intervals": ["10s","30s","1m","5m","15m","30m","1h","2h","1d"]},"timezone": "","title": "Real Time Statistical Performance","uid": "St9vqHnGk","version": 1}
# !pip install sklearn
# ?datasets.load_iris
# ### Download prediction and feedback data
# +
from sklearn.model_selection import train_test_split
from sklearn import datasets
import numpy as np
X_test, y_test = datasets.load_iris(return_X_y=True)
# convert y to one hot
y_test = np.eye(y_test.max() + 1)[y_test]
print(y_test[:2])
# -
# ### Send Prediction Requests
# +
import time
puids = []
for x in X_test:
pred_req = {"data":{"ndarray":[x.tolist()]}}
pred_resp = requests.post(f"{url}/predictions", json=pred_req)
puid_seldon = pred_resp.headers.get("seldon-puid")
puids.append(puid_seldon)
time.sleep(0.1)
# -
puids[:2]
for puid, y in zip(puids, y_test):
data = {
"truth": {
'data': {
'names': ['t:0', 't:1', 't:2'],
'ndarray': [y.tolist()]
}
}
}
requests.post(f"{url}/feedback", json=data, headers={"seldon-puid": puid})
time.sleep(0.5)
# ### Send randomized metrics to visualise degrade performance
np.random.shuffle(y_test)
for puid, y in zip(puids, y_test):
data = {
"truth": {
'data': {
'names': ['t:0', 't:1', 't:2'],
'ndarray': [y.tolist()]
}
}
}
requests.post(f"{url}/feedback", json=data, headers={"seldon-puid": puid})
time.sleep(0.5)
# ### VIsualise metrics
#
# Now you should be able to see real time performance in the dashboard for "Accuracy", "Precision" and "Recall". If it doesn't appear, try editing the dashboard and reapplying the time range. You can also double-check prometheus.
#
# Furthermore you should also be able to get further insights from the data that is stored in Elasticsearch.
| examples/feedback/feedback-metrics-server/README.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:autosk07] *
# language: python
# name: conda-env-autosk07-py
# ---
# +
import os
import math
import random
import pandas as pd
import numpy as np
import seaborn as sns
import autosklearn.regression as autoreg
from autosklearn.experimental.askl2 import AutoSklearn2Classifier
from autosklearn.metrics import make_scorer
from sklearn.model_selection import TimeSeriesSplit
from pandas import datetime
from matplotlib import pyplot as plt
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
from sklearn.metrics import mean_absolute_error
from sklearn.metrics import r2_score
from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
from statsmodels.tsa.stattools import acf, pacf
import sys
sys.path.insert(1, '../')
from functions import *
from plot import *
# -
from sklearn.model_selection import BaseCrossValidator
class BlockingTimeSeriesSplit(BaseCrossValidator):
def __init__(self, n_splits):
self.n_splits = n_splits
def get_n_splits(self, X, y, groups):
return self.n_splits
def split(self, X, y=None, groups=None):
n_samples = len(X)
k_fold_size = n_samples // self.n_splits
indices = np.arange(n_samples)
margin = 0
for i in range(self.n_splits):
start = i * k_fold_size
stop = start + k_fold_size
mid = int(0.8 * (stop - start)) + start
yield indices[start: mid], indices[mid + margin: stop]
# +
def quantile_loss(solution, prediction, quantile):
e = solution - prediction
return np.mean(np.maximum(e*quantile, e*(quantile - 1)))
qloss_05= make_scorer(
name="quantile_loss_05",
score_func=quantile_loss,
optimum=1,
greater_is_better=False,
needs_proba=False,
needs_threshold=False,
quantile=0.05,
)
qloss_95= make_scorer(
name="quantile_loss_95",
score_func=quantile_loss,
optimum=1,
greater_is_better=False,
needs_proba=False,
needs_threshold=False,
quantile=0.95,
)
qloss_10= make_scorer(
name="quantile_loss_10",
score_func=quantile_loss,
optimum=1,
greater_is_better=False,
needs_proba=False,
needs_threshold=False,
quantile=0.10,
)
qloss_90= make_scorer(
name="quantile_loss_90",
score_func=quantile_loss,
optimum=1,
greater_is_better=False,
needs_proba=False,
needs_threshold=False,
quantile=0.90,
)
qloss_025= make_scorer(
name="quantile_loss_025",
score_func=quantile_loss,
optimum=1,
greater_is_better=False,
needs_proba=False,
needs_threshold=False,
quantile=0.025,
)
qloss_975= make_scorer(
name="quantile_loss_975",
score_func=quantile_loss,
optimum=1,
greater_is_better=True,
needs_proba=False,
needs_threshold=False,
quantile=0.975,
)
qloss_5 = make_scorer(
name="quantile_loss_5",
score_func=quantile_loss,
optimum=1,
greater_is_better=True,
needs_proba=False,
needs_threshold=False,
quantile=0.5,
)
# -
train = pd.read_csv("../data/train_solar.csv", index_col = 'timestamp')
test = pd.read_csv("../data/test_solar.csv", index_col = 'timestamp')
plt.plot(train.energy.to_numpy())
plt.plot(test.energy.to_numpy())
from tsfresh.feature_extraction.feature_calculators import agg_autocorrelation
agg_autocorrelation(train.energy, param=[{"f_agg":'mean',
"maxlag":300}])
train = extract_dmhq(train)
test = extract_dmhq(test)
# +
from sklearn.model_selection import train_test_split
## Shuffle by every two days
data = pd.concat([train, test],axis = 0)
data = extract_dmhq(data)
cols = ['yd']
ftrain, ttrain = feature_target_construct(data, 'energy', 300, 192, 0, 192, cols, 4,
wd_on = False, d_on = False,
m_on = False, h_on = False, q_on = False)
data = pd.concat([ftrain, ttrain], axis = 1)
# now define random split groups
groups = [data for _, data in data.groupby('yd1(t+0)')]
random.shuffle(groups,)
for i, df in enumerate(groups):
data['yd1(t+0)'] = i+1
shuffled = pd.concat(groups).reset_index(drop=True)
ftrain = shuffled.iloc[:,1:201]
ttrain = shuffled.iloc[:,201:]
## split in 7/3 manner
train_X, test_X, train_Y, test_Y = train_test_split(ftrain, ttrain, train_size = 0.7, shuffle = False)
# +
cols = ['cloudCover','uvIndex']
ftrain, ttrain = feature_target_construct(train, 'energy', 300, 192, 0, 192, cols, 4,
wd_on = True, d_on = False,
m_on = False, h_on = True, q_on = False)
ftest, ttest = feature_target_construct(test, 'energy', 300, 192, 0, 192, cols, 4,
wd_on = True, d_on = False,
m_on = False, h_on = True, q_on = False)
# -
from sklearn.linear_model import Ridge
rr = Ridge(alpha=10000)
rr.fit(ftrain, ttrain)
ypred = rr.predict(ftest)
yhat = rr.predict(ftrain)
get_eval(ttrain, yhat)
get_eval(ttest, ypred)
from sklearn.linear_model import Ridge
rr = Ridge(alpha=10000)
rr.fit(train_X, train_Y)
ypred = rr.predict(test_X)
yhat = rr.predict(train_X)
get_eval(train_Y, yhat)
get_eval(test_Y, ypred)
# +
# pacf_val, conf = pacf(train.energy, nlags = 700, alpha = 0.05)
# indices = np.argwhere((np.abs(pacf_val) > 0.025))
# -
fig, ax = plt.subplots(nrows=1, ncols=1, figsize = (15,7))
plt.ylim(-0.1,0.1)
plot_pacf(train.energy, lags = 500, ax = ax)
plt.show()
# ### tsFresh featuer construction
train = pd.read_csv("../data/train_solar.csv", )
test = pd.read_csv("../data/test_solar.csv", )
ftrain, ttrain = tf_construct(train, 'energy', 200, 192)
ftest, ttest = tf_construct(test, 'energy', 200, 192)
# +
from tsfresh.utilities.dataframe_functions import make_forecasting_frame
from tsfresh import extract_features
# ftrain_tf = pd.DataFrame()
# for i in range(5000):
# df_time, y = make_forecasting_frame(ftrain.iloc[i,:].to_numpy(), kind = 'time', max_timeshift=300, rolling_direction=1)
# df_time.drop(['time','kind'],axis = 1, inplace = True)
# df_time = df_time.loc[df_time['id'] == 'id=id,timeshift=299']
# extracted_features = extract_features(df_time, column_id='id').dropna(axis = 1)
# ftrain_tf = ftrain_tf.append(extracted_features)
ftest_tf = pd.DataFrame()
for i in range(ftest.shape[0]):
df_time, y = make_forecasting_frame(ftest.iloc[i,:].to_numpy(), kind = 'time', max_timeshift=300, rolling_direction=1)
df_time.drop(['time','kind'],axis = 1, inplace = True)
df_time = df_time.loc[df_time['id'] == 'id=id,timeshift=299']
extracted_features = extract_features(df_time, column_id='id').dropna(axis = 1)
ftest_tf = ftest_tf.append(extracted_features)
# -
ftrain_tf.to_csv("tftrain_solar300_n5000.csv", index= 'id')
ftest_tf.to_csv("tftest_solar300_p1000.csv", index= 'id')
ftrain_tf.shape, ftest_tf.shape
# ## Lag features
# +
from sklearn.model_selection import TimeSeriesSplit, KFold
regl = autoreg.AutoSklearnRegressor(time_left_for_this_task=180000,
per_run_time_limit=18000,
initial_configurations_via_metalearning=0,
ensemble_size=50,
ensemble_nbest=25,
ensemble_memory_limit=5120,
seed=219, ml_memory_limit=10092,
include_estimators=None,
exclude_estimators=['gaussian_process'],
include_preprocessors=None,
exclude_preprocessors=None,
resampling_strategy = TimeSeriesSplit,
resampling_strategy_arguments={'folds':5,
'shuffle': False},
tmp_folder=None,
output_folder=None,
delete_tmp_folder_after_terminate=False,
delete_output_folder_after_terminate=False,
shared_mode=False,
n_jobs = 2,
disable_evaluator_output=False,
get_smac_object_callback=None,
smac_scenario_args=None,
logging_config=None,
metadata_directory=None)
regl.fit(ftrain, ttrain)
# -
plt.plot(test_Y.to_numpy()[100,])
plt.plot(ypred[100,], 'ro')
plt.figure(figsize=(15,7))
plt.plot(ypred[192,:], 'b.')
plt.plot(ttest.to_numpy()[192,:],'r-')
from pickle import dump
## pickle the model
dump(regl, open('200lag_10hr','wb'))
## load model
from pickle import load
with open('solar_200lag_20hr_ns', 'rb') as pickcle_file:
model = load(pickcle_file)
model.__class__
# %time model.refit(ftrain, ttrain)
model.get_models_with_weights()
from sklearn.pipeline import Pipeline
transform_pip = Pipeline([('data_pre', model.get_models_with_weights()[0][1][0]),
('feature_pre', model.get_models_with_weights()[0][1][1])])
pftrain = transform_pip.transform(ftrain)
pftest = transform_pip.transform(ftest)
model.get_models_with_weights()[0][1][2].choice.estimator.estimators_[0].predict(pftest)
model.get_models_with_weights()[0][1].predict(ftrain)
import sklearn.ensemble
from sklearn.pipeline import Pipeline
model_list = model.get_models_with_weights()
weight_list = []
predict_list = []
for i in range(len(model_list)):
weight_list.append(model_list[i][0])
prediction_list = []
# if contain forest based estimator, construct prediction list for
if issubclass(type(model_list[i][1][2].choice.estimator), \
sklearn.ensemble._forest.BaseForest):
for est in model_list[i][1][2].choice.estimator.estimators_:
pip = Pipeline([('data_pre', model_list[i][1][0]),
('feature_pre', model_list[i][1][1])])
pftest = pip.transform(ftest)
prediction_list.append(est.predict(pftest))
predict_list.append(prediction_list)
else:
predict_list.append(model_list[i][1].predict(ftest))
ub = 0
lb = 0
for i, weight in enumerate(weight_list):
# check if the prediction is list, then construct ci
if issubclass(type(predict_list[i]), list):
ub += weight * np.quantile(predict_list[i], 0.9, axis = 0)[0,:]
lb += weight * np.quantile(predict_list[i], 0.1, axis = 0)[0,:]
else:
ub += weight*predict_list[i]
lb += weight*predict_list[i]
ypred = model.predict(ftest)
plt.plot(ub, 'g--')
plt.plot(lb, 'g--')
plt.plot(ypred[0,:], 'b-')
plt.plot(ttest.to_numpy()[0,:],'r.')
# ## lag + tsfresh
ftrain_lts = np.concatenate([ftrain.to_numpy(), ftrain_tf.to_numpy()], axis = 1)
ftest_lts = np.concatenate([ftest.to_numpy(), ftest_tf.to_numpy()], axis = 1)
# +
reg_lts = autoreg.AutoSklearnRegressor(time_left_for_this_task=1000,
per_run_time_limit=300,
initial_configurations_via_metalearning=0,
ensemble_size=0,
ensemble_nbest=0,
ensemble_memory_limit=5120,
seed=921, ml_memory_limit=6144,
include_estimators=None,
exclude_estimators='gaussian_process',
include_preprocessors=None,
exclude_preprocessors=None,
resampling_strategy=TimeSeriesSplit,
resampling_strategy_arguments={'folds': 5},
tmp_folder=None,
output_folder=None,
delete_tmp_folder_after_terminate=False,
delete_output_folder_after_terminate=False,
shared_mode=False,
n_jobs = 6,
disable_evaluator_output=False,
get_smac_object_callback=None,
smac_scenario_args=None,
logging_config=None,
metadata_directory=None)
reg_lts.fit(ftrain_lts, ttrain)
# -
reg_lts.cv_results_
print(reg_lts.get_models_with_weights())
print(reg_lts.sprint_statistics())
reg_lts.refit(ftrain_lts, ttrain)
ypred_lts = reg_lts.predict(ftest_lts[:250,:])
yhat_lts = reg_lts.predict(ftrain_lts)
get_eval(ttest.iloc[:250,:], ypred_lts)
get_eval(ttrain, yhat_lts)
# +
from sklearn.metrics import r2_score
r2c_list_test = []
for i in range(192):
o_y = np.transpose(ttest.iloc[i,:].to_numpy().reshape(1,-1))
p_y = np.transpose(ypred_lts[i,:].reshape(1,-1))
r2 = r2_score(o_y, p_y)
r2c_list_test.append(r2)
fig = plt.figure(figsize = (16, 7))
plt.ylim(-0.5,0.9)
plt.plot(r2c_list_test, 'b-')
# -
plot_conf_std_static(ttrain, yhat, ttest, ypred_t, 200,1.96)
verf_ci_std_static(1.96, ttrain, yhat, ttest.iloc[:800,:], ypred_t[:800,:])
plot_conf_static(ttrain, yhat, ttest, ypred_t, 1000, 0.05)
verf_ci_static(0.05, ttrain, yhat, ttest, ypred_t)
| autosklearn/halfhourly_autosklearn_solar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <center><font size="+4">Programming and Data Analytics 1 2021/2022</font></center>
# <center><font size="+2">Sant'Anna School of Advanced Studies, Pisa, Italy</font></center>
# <center><img src="https://github.com/EMbeDS-education/StatsAndComputing20212022/raw/main/PDA/jupyter/jupyterNotebooks/images/SSSA.png" width="700" alt="EMbeDS"></center>
#
# <center><font size="+2">Course responsible</font></center>
# <center><font size="+2"><NAME> <EMAIL></font></center>
#
# <center><font size="+2">Co-lecturer </font></center>
# <center><font size="+2"><NAME> <EMAIL></font></center>
#
# ---
# + [markdown] id="PzTk_3lR7k5D"
# <center><font size="+4">Assignments for</font></center>
# <center><font size="+4">Lecture 1: Course Introduction</font></center>
#
# ---
# + cellView="form" colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 317, "status": "ok", "timestamp": 1622905634426, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgLeN60kpJtdBKQxyRrU7GwxagDOGnFA3G3Z8BwMA=s64", "userId": "01113523768495748338"}, "user_tz": -120} id="p666zbf89eCD" outputId="ad62730a-3d73-4074-fabc-eb9cee9067f8"
#@title <mark>RUN, BUT DO NOT MODIFY</mark>
# !curl -O https://raw.githubusercontent.com/EMbeDS-education/StatsAndComputing20212022/main/PDA/jupyter/jupyterNotebooks/assignments/auto_testing.py
# %reload_ext autoreload
# %autoreload 2
from auto_testing import *
# + [markdown] id="Om177ojl7k5J"
# # Assignment 01.01: Ciao, mondo!
# ## Statement
#
# Write a program that prints the Italian translation of `Hello, world!`.
#
# That is: `Ciao, mondo!`
#
# ## Example input
#
# ```
#
# ```
#
# ## Example output
#
# ```
# Ciao, mondo!
# ```
#
# ## Theory
#
# Well, there is not much theory to use here.
#
# In the example, you can see how to print 'Hello, world!'.
# + [markdown] id="FMZE1vmMbRLz"
# ## Write your solution here
#
#
# * Do not change the first line (`def ...():`)
# * Maintain the given indentation
# * Run the cell once you are done to load your solution
# * You can run some tests by yourself by decommenting the last line
#
#
# + executionInfo={"elapsed": 460, "status": "ok", "timestamp": 1622905638233, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgLeN60kpJtdBKQxyRrU7GwxagDOGnFA3G3Z8BwMA=s64", "userId": "01113523768495748338"}, "user_tz": -120} id="Yg4jeBnA7k5K"
def asgn01_01Hello_world():
# This program prints 'Hello, world!':
print('Hello, world!')
# Can you change it so that it prints the same,
#but in Italian?
#You can test independently your solution by executing the following line
#asgn01_01Hello_world()
# + [markdown] id="_lqTangP8R46"
# ## Run the following cells to perform the provided tests
# + cellView="form" colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 11, "status": "ok", "timestamp": 1622905639328, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgLeN60kpJtdBKQxyRrU7GwxagDOGnFA3G3Z8BwMA=s64", "userId": "01113523768495748338"}, "user_tz": -120} id="fwX9u8aW7k5L" outputId="241d2a05-543c-4496-a256-ad961b54f9d7"
#@title TEST 1
inputs=[]
expected_outputs=["Ciao, mondo!"]
run_and_test(inputs,expected_outputs,asgn01_01Hello_world)
# + [markdown] id="1JMDln-vIKRB"
# # Assignment 01.02: Hello, name1 name2
# ## Statement
#
# Write a program that reads two names (one per line), and prints
#
# `'Hello, name1 name2'`, where `name1` and `name2` are the two read names.
#
#
# ## Example input
#
# ```
# Andrea
# ```
#
# ```
# Daniele
# ```
#
# ## Example output
#
# ```
# Hello, <NAME>
# ```
#
# ## Theory
#
# In the notebook shown in class, we have seen
#
# - how to read a name from console and print `Hello, name`
# - how to print more parts/sentences using a single `print()`
#
# You have to combine these two features.
# + [markdown] id="IROCYUDQb0t7"
# ## Write your solution here
#
#
# * Do not change the first line (`def ...():`)
# * Maintain the given indentation
# * Run the cell once you are done to load your solution
# * You can run some tests by yourself by decommenting the last line
#
#
# + executionInfo={"elapsed": 255, "status": "ok", "timestamp": 1622905774690, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgLeN60kpJtdBKQxyRrU7GwxagDOGnFA3G3Z8BwMA=s64", "userId": "01113523768495748338"}, "user_tz": -120} id="QtPMkR0aJRYv"
def asgn01_02Hello_name1_name2():
# This program reads one name, name1, and then prints
# 'Hello, name1':
name1=input()
print('Hello,', name1)
# Can you change it so that it reads two names,
# name1 and name2, and then prints prints
# 'Hello, name1 name2'?
#You can test independently your solution by executing the following line
#asgn01_02Hello_name1_name2()
# + [markdown] id="HcoyoBpScQ7f"
# ## Run the following cells to perform the provided tests
# + cellView="form" colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 322, "status": "ok", "timestamp": 1622905777520, "user": {"displayName": "<NAME>", "photoUrl": "https://<KEY>", "userId": "01113523768495748338"}, "user_tz": -120} id="bMb_4CIDtvsp" outputId="5728d180-3d56-408b-8766-71de61594959"
#@title RUN and TEST ALL
#@markdown 1. TEST <NAME>
#@markdown 2. TEST <NAME>
#@markdown 3. TEST <NAME>
inputs=[['Renzo','Lucia'],['Andrea','Daniele'],['Daniele','Andrea']]
expected_outputs=[ ["Hello, <NAME>"],["Hello, <NAME>"],["Hello, <NAME>"]]
for k in range(len(inputs)):
print('-'*60)
run_and_test(inputs[k],expected_outputs[k],asgn01_02Hello_name1_name2)
# + [markdown] id="CvRjQW_ITQ8T"
# # Assignment 01.03: Hello, name1 and name2
# ## Statement
#
# Write a program that reads two names (one per line), and prints
#
# `'Hello, name1 and name2'`, where `name1` and `name2` are the two read names.
#
#
# ## Example input
#
# ```
# Andrea
# ```
#
# ```
# Daniele
# ```
#
# ## Example output
#
# ```
# Hello, Andrea and Daniele
# ```
#
# ## Theory
#
# In the notebook shown in class, we have seen
#
# - how to read one name from console and print `Hello, name`
# - how to print more parts/sentences using a single `print()`
#
# You have to combine these two features.
#
# Note that with respect to assignment 01.02 you have to add an 'and' between the two names.
# + [markdown] id="Lr-UYAK6b18z"
# ## Write your solution here
#
#
# * Do not change the first line (`def ...():`)
# * Maintain the given indentation
# * Run the cell once you are done to load your solution
# * You can run some tests by yourself by decommenting the last line
#
#
# + executionInfo={"elapsed": 256, "status": "ok", "timestamp": 1622905844723, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgLeN60kpJtdBKQxyRrU7GwxagDOGnFA3G3Z8BwMA=s64", "userId": "01113523768495748338"}, "user_tz": -120} id="ghJWGiEPUD9v"
def asgn01_03Hello_name1_and_name2():
# This program reads one name, name1, and then prints 'Hello, name1':
name1=input()
print('Hello,', name1)
# Can you change it so that it reads two names,
# name1 and name2, and then prints prints
# 'Hello, name1 and name2'?
#You can test independently your solution by executing the following line
#asgn01_03Hello_name1_and_name2()
# + [markdown] id="XJA5H0ULcVAM"
# ## Run the following cells to perform the provided tests
# + cellView="form" colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 249, "status": "ok", "timestamp": 1622905865939, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgLeN60kpJtdBKQxyRrU7GwxagDOGnFA3G3Z8BwMA=s64", "userId": "01113523768495748338"}, "user_tz": -120} id="TXNx80MMuVLI" outputId="74dac658-922e-4378-d216-1faf9fe6e2da"
#@title RUN and TEST ALL
#@markdown 1. TEST <NAME>
#@markdown 2. TEST <NAME>
#@markdown 3. TEST <NAME>
inputs=[['Renzo','Lucia'],['Andrea','Daniele'],['Daniele','Andrea']]
expected_outputs=[ ["Hello, Renzo and Lucia"],["Hello, Andrea and Daniele"],["Hello, Daniele and Andrea"]]
for k in range(len(inputs)):
print('-'*60)
run_and_test(inputs[k],expected_outputs[k],asgn01_03Hello_name1_and_name2)
| PDA/jupyter/jupyterNotebooks/assignments/.ipynb_checkpoints/01ConsoleIOandVariables_Assignments-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Computation of cutting planes: example 1
# # The set-up
import numpy as np
import pandas as pd
import accpm
# %load_ext autoreload
# %autoreload 1
# %aimport accpm
# $\DeclareMathOperator{\domain}{dom}
# \newcommand{\transpose}{\text{T}}
# \newcommand{\vec}[1]{\begin{pmatrix}#1\end{pmatrix}}$
# # Example
# To test the computation of cutting planes we consider the unconstrained convex optimization problem
# \begin{align*}
# &\text{minimize} \quad f_\text{obj}(x_0, x_1) = (x_0 - 5)^2 + (x_1 - 5)^2,
# \end{align*}
# and also the same problem with inequality constraints convex. That is, the problem
# \begin{align*}
# &\text{minimize} \quad f_\text{obj}(x_0, x_1) = (x_0 - 5)^2 + (x_1 - 5)^2 \\
# &\phantom{\text{minimize}} \quad f_0(x_0, x_1) =
# a_0^\transpose x - b_0 = \vec{1\\0}^\transpose \vec{x_0\\x_1} - 20 = x_0 - 20 \leq 0\\
# &\phantom{\text{minimize}} \quad f_1(x_0, x_1) =
# a_1^\transpose x - b_1 = \vec{-1\\0}^\transpose \vec{x_0\\x_1} = -x_0 \leq 0\\
# &\phantom{\text{minimize}} \quad f_2(x_0, x_1) =
# a_2^\transpose x - b_2 = \vec{0\\1}^\transpose \vec{x_0\\x_1} - 20 = x_1 - 20 \leq 0 \\
# &\phantom{\text{minimize}} \quad f_3(x_0, x_1) =
# a_3^\transpose x - b_3 = \vec{0\\-1}^\transpose \vec{x_0\\x_1} = -x_1 \leq 0.
# \end{align*}
# In both cases it is clear that the solution is $x^\star = (x_1^\star, x_2^\star) = (5, 5)$.
#
# The ACCPM requires the gradients of the objective function and constraint functions, which are
# \begin{align*}
# &\nabla f_\text{obj}(x_0, x_1) = \vec{2(x_0 - 5)\\2(x_1 - 5)}, \\
# &\nabla f_0(x_0, x_1) = \vec{1\\0}, \quad \nabla f_1(x_0, x_1) = \vec{-1\\0}, \\
# &\nabla f_2(x_0, x_1) = \vec{0\\1}, \quad \nabla f_3(x_0, x_1) = \vec{0\\-1}.
# \end{align*}
# We implement these functions as follows:
# +
def funcobj(x):
return (x[0]-5)**2 + (x[1]-5)**2
def func0(x):
return x[0] - 20
def func1(x):
return -x[0]
def func2(x):
return x[1] - 20
def func3(x):
return -x[1]
def grad_funcobj(x):
return np.array([2*(x[0] - 5), 2*(x[1] - 5)])
def grad_func0(x):
return np.array([1, 0])
def grad_func1(x):
return np.array([-1, 0])
def grad_func2(x):
return np.array([0, 1])
def grad_func3(x):
return np.array([0, -1])
# -
# Here we analytically compute the initial few iterations for the unconstrained problem. The ACCPM requires that the initial polygon $\mathcal{P}_0$ (here I've abused terminology and by the initial polygon $\mathcal{P}_0$ I actually mean the system of linear inequalities $Ax \leq b$) contain at least some of the points we are interested in. For the purposes of this example we take
# \begin{align*}
# A = \vec{a_0^\transpose\\a_1^\transpose\\a_2^\transpose\\a_3^\transpose}, b = \vec{20\\0\\20\\0}.
# \end{align*}
#
# Now, we start with $k=0$.
# Now, $x^{(0)}_{ac}$ is the solution of the minimization problem
# \begin{equation*}
# \min_{\domain \phi} \phi(x) = - \sum_{i=0}^{3}{\log{(b_i - a_i^\transpose x)}}.
# \end{equation*}
# So, we solve the problem
# \begin{align*}
# &\phantom{iff}\nabla \phi(x) = \sum_{i=0}^{3
# } \frac{1}{b_i - a_i^\transpose x}a_i = 0 \\
# &\iff \frac{1}{20-x_0}\begin{bmatrix}1\\0\end{bmatrix} + \frac{1}{x_0}\begin{bmatrix}-1\\0\end{bmatrix} + \frac{1}{20-x_1}\begin{bmatrix}0\\1\end{bmatrix} + \frac{1}{x_1}\begin{bmatrix}0\\-1\end{bmatrix} = 0 \\
# &\iff \frac{1}{20-x_0} - \frac{1}{x_0} = 0, \frac{1}{20-x_1} - \frac{1}{x_1} = 0 \\
# &\iff x_0 = \frac{20}{2} = 10, x_1 = \frac{20}{2} = 10,
# \end{align*}
# and conclude $x^{(0)}_{ac} = (10, 10)$. We then query the oracle at $x^{(0)}_{ac}$. (Here, $f_\text{best} = f_\text{obj}(10, 10) = 50$ since this is the $0$-th iteration.) As there are no inequality constraints we have
# \begin{align*}
# &a_4 = \nabla f_\text{obj}(10, 10) = \vec{10\\10}, \\
# &b_4 = \nabla f_\text{obj}(10, 10)^\transpose \vec{10\\10} = \vec{10\\10}^\transpose \vec{10\\10} = 200,
# \end{align*}
# which we normalize to get
# \begin{align*}
# &a_4 = \frac{1}{\sqrt{100^2 + 100^2}} \nabla f_\text{obj}(10, 10)
# = \vec{\frac{1}{\sqrt{2}} \\ \frac{1}{\sqrt{2}} } \approx \vec{0.7071 \\ 0.7071}, \\
# &b_4 = \frac{1}{\sqrt{100^2 + 100^2}} \nabla f_\text{obj}(10, 10)^\transpose \vec{10\\10} = \vec{10\\10}^\transpose \vec{10\\10} = \frac{20}{\sqrt{2}} = 10\sqrt{2} \approx 14.1421,
# \end{align*}
# and therefore update
# \begin{align*}
# A = \vec{a_0^\transpose\\a_1^\transpose\\a_2^\transpose\\a_3^\transpose\\ \frac{1}{\sqrt{2}} \;\; \frac{1}{\sqrt{2}}}, b = \vec{20\\0\\20\\0\\10\sqrt{2}}, k = 1.
# \end{align*}
# Now, $x^{(1)}_{ac}$ is the solution of the minimization problem
# \begin{equation*}
# \min_{\domain \phi} \phi(x) = - \sum_{i=0}^{4}{\log{(b_i - a_i^\transpose x)}}.
# \end{equation*}
# So, we solve the problem
# \begin{align*}
# &\phantom{iff}\nabla \phi(x) = \sum_{i=0}^{4
# } \frac{1}{b_i - a_i^\transpose x}a_i = 0 \\
# &\iff \frac{1}{20-x_0}\vec{1\\0} + \frac{1}{x_0}\vec{-1\\0} + \frac{1}{20-x_1}\vec{0\\1} + \frac{1}{x_1}\vec{0\\-1} + \frac{\sqrt{2}}{20-x_0-x_1} \vec{\frac{1}{\sqrt{2}}\\\frac{1}{\sqrt{2}}} = 0\\
# &\iff \frac{1}{20-x_0} - \frac{1}{x_0} + \frac{1}{20 - x_0- x_1}= 0, \frac{1}{20-x_1} - \frac{1}{x_1} + \frac{1}{20 - x_0- x_1} = 0 \\
# &\iff x_0 = x_1 = 2(5 \pm \sqrt{5}) \approx 14.4721 \text{ or } 5.52786,
# \end{align*}
# and take $x^{(1)}_{ac} = (2(5-\sqrt{5}), 2(5-\sqrt{5})) \approx (5.52786, 5.52786)$. We then query the
# oracle at $x^{(1)}_{ac}$. Here
# $f_\text{obj}(x^{(1)}_{ac}) = f_\text{obj}(2(5-\sqrt{5}), 2(5-\sqrt{5})) = 90 - 40\sqrt{5} \approx 0.557281 \leq f_\text{best} = 50$ so we update
# $f_\text{best} = 90 - 40\sqrt{5} \approx 0.557281$ and therefore put (and normalize)
# \begin{align*}
# &a_5 = \frac{1}{\sqrt{2(10-4\sqrt{5})^2}} \nabla f_\text{obj}(x^{(1)}_{ac}) =
# \frac{1}{\sqrt{2(10-4\sqrt{5})^2}} \nabla f_\text{obj}\vec{2(5-\sqrt{5}) \\ 2(5-\sqrt{5})} =
# \frac{1}{\sqrt{2(10-4\sqrt{5})^2}} \vec{10-4\sqrt{5}\\10-4\sqrt{5}}
# = \vec{\frac{1}{\sqrt{2}}\\\frac{1}{\sqrt{2}}}\approx \vec{0.7071 \\ 0.7071}, \\
# &b_5 = \frac{1}{\sqrt{2(10-4\sqrt{5})^2}} \nabla f_\text{obj}(x^{(1)}_{ac})^\transpose \vec{2(5-\sqrt{5}) \\ 2(5-\sqrt{5})} =
# \frac{1}{\sqrt{2}} \vec{1\\1}^\transpose \vec{2(5-\sqrt{5}) \\ 2(5-\sqrt{5})} = 2\sqrt{2} (5 - \sqrt{5})
# \approx 7.8176,
# \end{align*}
# updating $A$ and $b$ also. Iteration $k = 1$ is concluded by incrementing $k$.
#
# We see that this computation will only get more complicated. Therefore, in our implementation we test the computed analytic center by simply computing the norm of the gradient of the log barrier at the analytic center and see if it sufficiently small.
# We first test the unconstrained version of the problem.
A = np.array([[1, 0],[-1,0],[0,1],[0,-1]])
b = np.array([20, 0, 20, 0])
accpm.accpm(A, b, funcobj, grad_funcobj, alpha=0.01, beta=0.7,
start=1, tol=10e-3, maxiter = 200, testing=1)
# Next we test a trivial version of the inequality constrained problem where the inequality constraints and linear inequality constraints are the same
A = np.array([[1, 0],[-1,0],[0,1],[0,-1]])
b = np.array([20, 0, 20, 0])
accpm.accpm(A, b, funcobj, grad_funcobj,
(func0, func1, func2, func3), (grad_func0, grad_func1, grad_func2, grad_func3),
alpha=0.01, beta=0.7, start=1, tol=10e-3, maxiter=200, testing=True)
# Next we test a version of the inequality constrained problem where the initial polygon lies within the feasible region given by the inequality constraints.
A = np.array([[1, 0],[-1,0],[0,1],[0,-1]])
b = np.array([5, 0, 5, 0])
accpm.accpm(A, b, funcobj, grad_funcobj,
(func0, func1, func2, func3), (grad_func0, grad_func1, grad_func2, grad_func3),
alpha=0.01, beta=0.7, start=1, tol=10e-3, maxiter=200, testing=True)
# We now test a version of the inequality constrained problem where the initial polyhedron contains the feasible region but also regions that are infeasible.
A = np.array([[1, 0],[-1,0],[0,1],[0,-1]])
b = np.array([30, 0, 30, 0])
accpm.accpm(A, b, funcobj, grad_funcobj,
(func0, func1, func2, func3), (grad_func0, grad_func1, grad_func2, grad_func3),
alpha=0.01, beta=0.7, start=1, tol=10e-3, maxiter=200, testing=True)
# We test a version of the inequality constrained problem where the initial polyhedron is moderately large.
A = np.array([[1, 0],[-1,0],[0,1],[0,-1]])
b = np.array([100, 0, 100, 0])
accpm.accpm(A, b, funcobj, grad_funcobj,
(func0, func1, func2, func3), (grad_func0, grad_func1, grad_func2, grad_func3),
alpha=0.01, beta=0.7, start=1, tol=10e-3, maxiter=200, testing=True)
# Finally we test a version of the inequality constrained problem where the initial polyhedron is very large. We observe that the algorithm does not converge.
#
# It is possible more iterations, or pruning of the redundant inequalities, would allow for convergence, but in practice, it is unlikely this situation will arise.
A = np.array([[1, 0],[-1,0],[0,1],[0,-1]])
b = np.array([1000, 0, 1000, 0])
accpm.accpm(A, b, funcobj, grad_funcobj,
(func0, func1, func2, func3), (grad_func0, grad_func1, grad_func2, grad_func3),
alpha=0.01, beta=0.7, start=1, tol=10e-3, maxiter=200, testing=True)
A = np.array([[1, 0],[-1,0],[0,1],[0,-1]])
b = np.array([1000, 0, 1000, 0])
accpm.accpm(A, b, funcobj, grad_funcobj,
(func0, func1, func2, func3), (grad_func0, grad_func1, grad_func2, grad_func3),
alpha=0.01, beta=0.7, start=1, tol=10e-3, maxiter=500, testing=True)
| projects/david/lab/test_accpm_toy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.0 64-bit (''ensembletech'': virtualenv)'
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import statistics as stts
from sklearn.tree import DecisionTreeClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
from sklearn.ensemble import VotingClassifier
# +
data = pd.read_csv('../data/data_cleaned.csv')
X, y = data.drop('Survived', axis=1), data.pop('Survived')
X_train, X_test, y_train, y_test = train_test_split(X,y, random_state = 9 , stratify = y)
# +
DT = DecisionTreeClassifier()
KNN = KNeighborsClassifier()
LR = LogisticRegression(solver='lbfgs', max_iter=1000)
DT.fit(X_train, y_train)
KNN.fit(X_train, y_train)
LR.fit(X_train, y_train)
y_pred1 = DT.predict(X_test)
y_pred2 = KNN.predict(X_test)
y_pred3 = LR.predict(X_test)
# -
final_pred = (0.25*y_pred1 + 0.25*y_pred2 + 0.5*y_pred3)
final_pred = np.round(final_pred)
accuracy_score(y_test, final_pred)
| basic/weighted_average.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Working with the Bag-of-Words representation
#
# The [bow module](api.rst#tmtoolkit-bow) in tmtoolkit contains several functions for working with Bag-of-Words (BoW) representations of documents. It's divided into two sub-modules: [bow.bow_stats](api.rst#module-tmtoolkit.bow.bow_stats) and [bow.dtm](api.rst#module-tmtoolkit.bow.dtm). The former implements several statistics and transformations for BoW representations, the latter contains functions to create and convert sparse or dense document-term matrices (DTMs).
#
# Most of the functions in both sub-modules accept and/or return sparse DTMs. The [previous chapter](preprocessing.ipynb) contained a section about what sparse DTMs are and [how they can be generated with tmtoolkit](preprocessing.ipynb#Generating-a-sparse-document-term-matrix-(DTM)).
# ## An example document-term matrix
#
# Before we start with the [bow.dtm](api.rst#module-tmtoolkit.bow.dtm) module, we will generate a sparse DTM from a small example corpus.
# +
import random
random.seed(20191113) # to make the sampling reproducible
import numpy as np
np.set_printoptions(precision=5)
from tmtoolkit.corpus import Corpus
corpus = Corpus.from_builtin_corpus('english-NewsArticles').sample(5)
# -
# Let's have a look at a sample document:
print(corpus['NewsArticles-2058'][:227])
# We employ a preprocessing pipeline that removes a lot of information from our original data in order to obtain a very condensed DTM.
# +
from tmtoolkit.preprocess import TMPreproc
preproc = TMPreproc(corpus)
preproc.pos_tag() \
.lemmatize() \
.filter_for_pos('N') \
.tokens_to_lowercase() \
.remove_special_chars_in_tokens() \
.clean_tokens(remove_shorter_than=2) \
.remove_common_tokens(5, absolute=True) # remove tokens that occur in all documents
preproc.tokens_datatable
# -
preproc.n_docs, preproc.vocabulary_size
# We fetch the document labels and vocabulary and convert them to NumPy arrays, because such arrays allow advanced indexing methods such as boolean indexing.
doc_labels = np.array(preproc.doc_labels)
doc_labels
vocab = np.array(preproc.vocabulary)
vocab[:10] # only showing the first 10 tokens here
# Finally, we fetch the sparse DTM:
dtm = preproc.dtm
dtm
# We now have a sparse DTM `dtm`, an array of document labels `doc_labels` that represent the rows of the DTM and an array of vocabulary tokens `vocab` that represent the columns of the DTM. We will use this data for the remainder of the chapter.
# ## The `bow.dtm` module
#
# This module is quite small. There are two functions to convert a DTM to a [datatable](https://github.com/h2oai/datatable/) or [DataFrame](https://pandas.pydata.org/): [dtm_to_datatable()](api.rst#tmtoolkit.bow.dtm.dtm_to_datatable) and [dtm_to_dataframe()](api.rst#tmtoolkit.bow.dtm.dtm_to_dataframe). Note that the generated datatable or DataFrame is *dense*, i.e. it uses up (much) more memory than the input DTM.
#
# Let's generate a datatable via [dtm_to_datatable()](api.rst#tmtoolkit.bow.dtm.dtm_to_datatable) from our DTM, the document labels and the vocabulary:
# +
from tmtoolkit.bow.dtm import dtm_to_datatable
dtm_to_datatable(dtm, doc_labels, vocab)
# -
# We can see that a row `_doc` with the document labels was created and that the vocabulary tokens become the column names. [dtm_to_dataframe()](api.rst#tmtoolkit.bow.dtm.dtm_to_dataframe) works the same way.
#
# You can combine tmtoolkit with [Gensim](https://radimrehurek.com/gensim/). The `bow.dtm` module provides several functions to convert data between both packages:
#
# - [dtm_and_vocab_to_gensim_corpus_and_dict()](api.rst#tmtoolkit.bow.dtm.dtm_and_vocab_to_gensim_corpus_and_dict): converts a (sparse) DTM and a vocabulary list to a *Gensim Corpus* and *Gensim Dictionary*
# - [dtm_to_gensim_corpus()](api.rst#tmtoolkit.bow.dtm.dtm_to_gensim_corpus): convert a (sparse) DTM only to a *Gensim Corpus*
# - [gensim_corpus_to_dtm()](api.rst#tmtoolkit.bow.dtm.gensim_corpus_to_dtm): converts a *Gensim Corpus* object to a sparse DTM in COO format
# ## The `bow.bow_stats` module
#
# This module provides several statistics and transformations for sparse or dense DTMs.
#
# ### Document lengths, document and term frequencies, token co-occurrences
#
# Let's start with the [doc_lengths()](api.rst#tmtoolkit.bow.bow_stats.doc_lengths) function, which simply gives the number of tokens per document (i.e. the row-wise sum of the DTM):
# +
from tmtoolkit.bow.bow_stats import doc_lengths
doc_lengths(dtm)
# -
# The returned array is aligned to the document labels `doc_labels` so we can see that the last document, "NewsArticles-3665", is the one with the most tokens. Or to do it computationally:
doc_labels[doc_lengths(dtm).argmax()]
# While [doc_lengths()](api.rst#tmtoolkit.bow.bow_stats.doc_lengths) gives the row-wise sum across the DTM, [term_frequencies()](api.rst#tmtoolkit.bow.bow_stats.term_frequencies) gives the column-wise sum. This means it returns an array of the length of the vocabulary's size where each entry in that array reflects the number of occurrences of the respective vocabulary token (aka term).
#
# Let's calculate that measure, get its maximum and the vocabulary token(s) for that maximum value:
# +
from tmtoolkit.bow.bow_stats import term_frequencies
term_freq = term_frequencies(dtm)
(term_freq.max(), vocab[term_freq == term_freq.max()])
# -
# It's also possible to calculate the proportional frequency, i.e. normalize the counts by the overall number of tokens via `proportions=True`:
term_prop = term_frequencies(dtm, proportions=True)
vocab[term_prop >= 0.01]
# The function [doc_frequencies()](api.rst#tmtoolkit.bow.bow_stats.doc_frequencies) returns how often each token in the vocabulary occurs at least *n* times per document. You can control *n* per parameter `min_val` which is set to `1` by default. The returned array is aligned with the vocabulary. Here, we calculate the document frequency with the default value `min_val=1`, extract the maximum document frequency and see which of the tokens in the `vocab` array reach the maximum document frequency:
# +
from tmtoolkit.bow.bow_stats import doc_frequencies
df = doc_frequencies(dtm)
max_df = df.max()
max_df, vocab[df == max_df]
# -
# It turns out that the maximum document frequency is 4 and only the token "minister" reaches that document frequency. This means only "minister" is mentioned across 4 documents at least once (because `min_val` is `1`). Remember that during preprocessing, we removed all tokens that occur across *all* five documents, hence there can't be a vocabulary token with a document frequency of 5.
#
# Let's see which vocabulary tokens occur within a single document at least 10 times:
df = doc_frequencies(dtm, min_val=10)
vocab[df > 0]
# We can also calculate the *co-document frequency* or *token co-occurrence* matrix via [codoc_frequencies()](api.rst#tmtoolkit.bow.bow_stats.codoc_frequencies). This measures how often each pair of vocabulary tokens occurs at least *n* times together in the same document. Again, you can control *n* per parameter `min_val` which is set to `1` by default. The result is a sparse matrix of shape *vocabulary size* by *vocabulary size*. The columns and rows give the pairs of tokens from the vocabulary.
#
# Let's generate a co-document frequency matrix and convert it to a dense representation, because our further operations don't support sparse matrices.
#
# A co-document frequency matrix is symmetric along the diagonal, because co-occurrence between a pair `(token1, token2)` is always the same as between `(token2, token1)`. We want to filter out the duplicate pairs and for that use [np.triu()](https://docs.scipy.org/doc/numpy/reference/generated/numpy.triu.html) to take only the upper triangle of the matrix, i.e. set all values in the lower triangle including the matrix diagonal to zero (`k=1` does this):
# +
from tmtoolkit.bow.bow_stats import codoc_frequencies
codoc_mat = codoc_frequencies(dtm).todense()
codoc_upper = np.triu(codoc_mat, k=1)
codoc_upper
# -
# Now we create a list that contains the pairs of tokens that occur together in at least two documents (`codoc_upper > 1`) together with their co-document frequency:
interesting_pairs = [(vocab[t1], vocab[t2], codoc_upper[t1, t2])
for t1, t2 in zip(*np.where(codoc_upper > 1))]
interesting_pairs[:10] # showing only the first ten pairs
# ### Generate sorted lists and datatables according to term frequency
#
# When working with DTMs, it's often helpful to rank terms per document according to their frequency. This is what [sorted_terms()](api.rst#tmtoolkit.bow.bow_stats.sorted_terms) does for you. It further allows to specify the sorting order (the default is descending order via `ascending=False`) and several limits:
#
# - `lo_thresh` for the minimum term frequency
# - `hi_thresh` for the maximum term frequency
# - `top_n` for the maximum number of terms per document
#
# Let's display the top three tokens per document by frequency:
# +
from tmtoolkit.bow.bow_stats import sorted_terms
sorted_terms(dtm, vocab, top_n=3)
# -
# The output is a list for each document (this means the output is aligned with the document labels `doc_labels`), with three pairs of `(token, frequency)` each. It's also possible to get this data as datatable via [sorted_terms_datatable()](api.rst#tmtoolkit.bow.bow_stats.sorted_terms_datatable), which gives a better overview and also includes labels for the documents. It accepts the same parameters for sorting and limitting the results:
# +
from tmtoolkit.bow.bow_stats import sorted_terms_datatable
sorted_terms_datatable(dtm, vocab, doc_labels, top_n=3)
# -
sorted_terms_datatable(dtm, vocab, doc_labels, lo_thresh=5)
# ### Term frequency–inverse document frequency transformation (tf-idf)
#
# [Term frequency–inverse document frequency transformation (tf-idf)](https://en.wikipedia.org/wiki/Tf%E2%80%93idf) is a matrix transformation that is often applied to DTMs in order to reflect the importance of a token to a document. The `bow_stats` module provides the function [tfidf()](api.rst#tmtoolkit.bow.bow_stats.tfidf) for this. When the input is a sparse matrix, and the calculation supports operating on sparce matrices, the output will also be a sparse matrix, which means that the tf-idf transformation is implemented in a very memory-efficient way.
#
# Let's apply tf-idf to our DTM using the default way:
# +
from tmtoolkit.bow.bow_stats import tfidf
tfidf_mat = tfidf(dtm)
tfidf_mat
# -
# We can see that the output is a sparse matrix. Let's have a look at its values:
tfidf_mat.todense()
# Of course we can also pass this matrix to `sorted_terms_datatable()` and observe that some rankings have changed in comparison to the untransformed DTM:
sorted_terms_datatable(tfidf_mat, vocab, doc_labels, top_n=3)
# The tf-idf matrix is calculated from a DTM $D$ as $\text{tf}(D) \text{idf}(D)$.
#
#
# There are different variants for how to calculate the term frequency $\text{tf}(D)$ and the inverse document frequency $\textit{idf(D)}$. tmtoolkit contains several functions that implement some of these variants. For $\text{tf()}$ these are:
#
# - [tf_binary()](api.rst#tmtoolkit.bow.bow_stats.tf_binary): binary term frequency matrix (matrix contains 1 whenever a term occurred in a document, else 0)
# - [tf_proportions()](api.rst#tmtoolkit.bow.bow_stats.tf_proportions): proportional term frequency matrix (term counts are normalized by document length)
# - [tf_log()](api.rst#tmtoolkit.bow.bow_stats.tf_log): log-normalized term frequency matrix (by default $\log(1 + D)$)
# - [tf_double_norm()](api.rst#tmtoolkit.bow.bow_stats.tf_double_norm): double-normalized term frequency matrix
# $K + (1-K) \cdot \frac{D}{\text{rowmax}(D)}$, where $\text{rowmax}(D)$ is a vector containing the maximum term count per document
#
# As you can see, all the term frequency functions are prefixed with a `tf_`. There are also two variants for $\text{idf()}$:
#
# - [idf()](api.rst#tmtoolkit.bow.bow_stats.idf): calculates $\log(\frac{a + N}{b + \text{df}(D)})$ where $a$ and $b$ are smoothing constants, $N$ is the number of documents and $\text{df}(D)$ calculates the [document frequency](#Document-lengths,-document-and-term-frequencies,-token-co-occurrences)
# - [idf_probabilistic()](api.rst#tmtoolkit.bow.bow_stats.idf_probabilistic): calculates $\log(a + \frac{N - \text{df}(D)}{\text{df}(D)})$
#
# The term frequency functions always return a sparse matrix if possible and if the input is sparse. Let's try out two term frequency functions:
# +
from tmtoolkit.bow.bow_stats import tf_binary, tf_proportions
tf_binary(dtm).todense()
# -
tf_proportions(dtm).todense()
# Just like the [document frequency](#Document-lengths,-document-and-term-frequencies,-token-co-occurrences) function [doc_frequencies()](api.rst#tmtoolkit.bow.bow_stats.doc_frequencies), the inverse document frequency functions also return a vector with the same length as the vocabulary. Let's use these functions and have a look at the inverse document frequency of certain tokens:
# +
from tmtoolkit.bow.bow_stats import idf, idf_probabilistic
idf_vec = idf(dtm)
list(zip(vocab, idf_vec))[:10]
# +
probidf_vec = idf_probabilistic(dtm)
list(zip(vocab, probidf_vec))[:10]
# -
# Note that due to our very small sample, there's not much variation in the inverse document frequency values.
#
# By default, [tfidf()](api.rst#tmtoolkit.bow.bow_stats.tfidf) uses `tf_proportions()` and `idf()` to calculate the tf-idf matrix. You can plug in other functions to get other variants of tf-idf:
# +
from tmtoolkit.bow.bow_stats import tf_double_norm
# we also set a "K" parameter for "tf_double_norm"
tfidf_mat2 = tfidf(dtm, tf_func=tf_double_norm,
idf_func=idf_probabilistic, K=0.25)
tfidf_mat2
# -
sorted_terms_datatable(tfidf_mat2, vocab, doc_labels, top_n=3)
# Once we have generated a DTM, we can use it for topic modeling. The [next chapter](topic_modeling.ipynb) will show how tmtoolkit can be used to evaluate the quality of your model, export essential information from it and visualize the results.
| doc/source/bow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # s{t} = [X{t}, Y{t}, X{t−1}, Y{t−1},..., X{t−7}, Y{t−7}, C].
# The input features s{t} are processed by a residual tower that consists of a single convolutional block
# followed by either 19 or 39 residual blocks.
#
# The convolutional block applies the following modules:
# (1) A convolution of 256 filters of kernel size 3 × 3 with stride 1
# (2) Batch normalization
# (3) A rectifier nonlinearity
#
# Each residual block applies the following modules sequentially to its input:
# (1) A convolution of 256 filters of kernel size 3 × 3 with stride 1
# (2) Batch normalization
# (3) A rectifier nonlinearity
# (4) A convolution of 256 filters of kernel size 3 × 3 with stride 1
# (5) Batch normalization
# (6) A skip connection that adds the input to the block
# (7) A rectifier nonlinearity
#
# The output of the residual tower is passed into two separate ‘heads’ for computing the policy and value.
#
# The policy head applies the following modules:
# (1) A convolution of 2 filters of kernel size 1 × 1 with stride 1
# (2) Batch normalization
# (3) A rectifier nonlinearity
# (4) A fully connected linear layer that outputs a vector of size 19*19 + 1 = 362, corresponding to logit probabilities for all intersections and the pass move
#
# The value head applies the following modules:
# (1) A convolution of 1 filter of kernel size 1 × 1 with stride 1
# (2) Batch normalization
# (3) A rectifier nonlinearity
# (4) A fully connected linear layer to a hidden layer of size 256
# (5) A rectifier nonlinearity
# (6) A fully connected linear layer to a scalar
# (7) A tanh nonlinearity outputting a scalar in the range [−1, 1]
#
# The overall network depth, in the 20 or 40 block network, is 39 or 79 parameterized layers, respectively,
# for the residual tower, plus an additional 2 layers for the policy head and 3 layers for the value head.
# """
#
# +
import keras
from keras.layers import Activation, BatchNormalization
from keras.layers import Conv2D, Dense, Flatten, Input
from keras.models import Model
import numpy as np
import time
class smallNN:
def __init__(self):
#self.board_input = Input(shape=TrojanGoPlane.shape(), name='board_input')
self.board_input = Input(shape=(7, 5, 5), name='board_input')
def nn_model(self):
pb = self.board_input
for i in range(4): # <1>
pb = Conv2D(64, (3, 3), # <1>
padding='same', # <1>
data_format='channels_first', # <1>
activation='relu')(pb) # <1>
policy_conv = \
Conv2D(2, (1, 1), # <2>
data_format='channels_first', # <2>
activation='relu')(pb) # <2>
policy_flat = Flatten()(policy_conv) # <2>
policy_output = \
Dense(26,
activation='softmax')(policy_flat) # <2>
value_conv = \
Conv2D(1, (1, 1), # <3>
data_format='channels_first', # <3>
activation='relu')(pb) # <3>
value_flat = Flatten()(value_conv) # <3>
value_hidden = Dense(256, activation='relu')(value_flat) # <3>
value_output = Dense(1, activation='tanh')(value_hidden) # <3>
model = Model(
inputs=[self.board_input],
outputs=[policy_output, value_output])
return model
class trojanGoZero:
def __init__(self, num_resnet_block=19):
#self.board_input = Input(shape=TrojanGoPlane.shape(), name='board_input')
self.board_input = Input(shape=(7, 5, 5), name='board_input')
self.num_resnet_block = num_resnet_block
self.num_filters = 256
def resNetBlock(self, x,filters,pool=False):
res = x
if pool:
x = MaxPooling2D(pool_size=(2, 2))(x)
res = Conv2D(filters=filters,kernel_size=[1,1],strides=(2,2),padding="same", data_format='channels_first')(res)
#out = BatchNormalization()(x)
#out = Activation("relu")(out)
out = Conv2D(filters=filters, kernel_size=[3, 3], strides=[1, 1], padding="same", data_format='channels_first')(x)
out = BatchNormalization()(out)
out = Activation("relu")(out)
out = Conv2D(filters=filters, kernel_size=[3, 3], strides=[1, 1], padding="same", data_format='channels_first')(out)
out = BatchNormalization()(out)
out = keras.layers.add([res,out])
out = Activation("relu")(out)
return out
def nn_model(self, input_shape):
#Input feature of 17*19*19 or 7*5*5 as board_input or board_images
board_images = Input(input_shape)
#board_images = self.board_input
#CNN-1 with Batch Normalization and rectifier nonlinearity.
cnn1 = Conv2D(filters=256, kernel_size=[3, 3], strides=[1, 1], padding="same", data_format='channels_first')(board_images)
cnn1_batch = BatchNormalization()(cnn1)
cnn1_act = Activation("relu")(cnn1_batch)
self_in = cnn1_act
#Now build 19 or 39 ResNet block networks depends on "num_resnet_block" variable.
for i in range(self.num_resnet_block):
self_out = self.resNetBlock(self_in, self.num_filters)
self_in = self_out
out = self_out
policy_conv = \
Conv2D(2, (1, 1), # <2>
data_format='channels_first', # <2>
activation='relu')(out) # <2>
policy_conv_bn = BatchNormalization()(policy_conv)
policy_flat = Flatten()(policy_conv_bn) # <2>
policy_output = \
Dense(26,
activation='softmax')(policy_flat) # <2>
value_conv = \
Conv2D(1, (1, 1), # <3>
data_format='channels_first', # <3>
activation='relu')(out) # <3>
value_conv_bn = BatchNormalization()(value_conv)
value_flat = Flatten()(value_conv_bn) # <3>
value_hidden = Dense(256, activation='relu')(value_flat) # <3>
value_output = Dense(1, activation='tanh')(value_hidden) # <3>
model = Model(
inputs=[board_images],
outputs=[policy_output, value_output])
return model
# +
net = trojanGoZero()
input_shape = (7,5,5)
model = net.nn_model(input_shape)
print(model.summary())
net_small = smallNN()
model_small = net_small.nn_model()
print(model_small.summary())
# +
model_input = []
for _ in range(100):
board_tensor = np.random.randint(0, 3, size=(7, 5, 5))
model_input.append(board_tensor)
model_input = np.array(model_input)
action_target = []
for _ in range (100):
search_prob = np.random.randn(26)
#search_prob_flat = search_prob.reshape(25,)
action_target.append(search_prob)
action_target = np.array(action_target)
value_target = np.random.rand(100)
value_target = np.array(value_target)
# +
#pickle the data
import pickle
def storeData():
start = time.time()
# database
db = {}
db['model_input'] = model_input
db['action_target'] = action_target
db['value_target'] = value_target
# Its important to use binary mode
dbfile = open('examplePickle', 'ab')
# source, destination
pickle.dump(db, dbfile)
dbfile.close()
finish = time.time()
print("Write Time taken :", finish - start)
def loadData():
start = time.time()
# for reading also binary mode is important
dbfile = open('examplePickle', 'rb')
db = pickle.load(dbfile)
"""
for keys in db:
print(keys, '=>', db[keys])
"""
dbfile.close()
finish = time.time()
print("Read Time taken :", finish - start)
storeData()
loadData()
# +
#use HDF5 to store and load
import h5py
class ExperienceBuffer:
def __init__(self, model_input, action_target, value_target):
self.model_input = model_input
self.action_target = action_target
self.value_target = value_target
def serialize(self, h5file):
h5file.create_group('experience')
h5file['experience'].create_dataset('model_input', data=self.model_input)
h5file['experience'].create_dataset('action_target', data=self.action_target)
h5file['experience'].create_dataset('value_target', data=self.value_target)
def load_experience(self, h5file):
return ExperienceBuffer(model_input=np.array(h5file['experience']['model_input']),
action_target=np.array(h5file['experience']['action_target']),
value_target=np.array(h5file['experience']['value_target'])
)
start = time.time()
with h5py.File('test.hdf5', 'w') as exp_outf:
ExperienceBuffer(model_input, action_target, value_target).serialize(exp_outf)
finish = time.time()
print("Write Time taken :", finish - start)
start = time.time()
with h5py.File('test.hdf5', 'r') as exp_input:
experience_buffer = ExperienceBuffer(model_input, action_target, value_target).load_experience(exp_input)
finish = time.time()
print("Read Time taken :", finish - start)
# +
from keras.optimizers import SGD
model.compile(SGD(lr=0.01), loss=['categorical_crossentropy', 'mse'])
model_small.compile(SGD(lr=0.01), loss=['categorical_crossentropy', 'mse'])
# +
import time
start = time.time()
model.fit(model_input, [action_target, value_target], batch_size=64, epochs=1)
finish = time.time()
print("Time taken : ", finish - start)
start = time.time()
model_small.fit(model_input, [action_target, value_target], batch_size=64, epochs=1)
finish = time.time()
print("Time taken : ", finish - start)
# +
X = model_input[0]
X = np.expand_dims(X, axis=0)
print(X.shape)
prediction = model.predict(X)
print(prediction)
print("*"*60)
prediction = model_small.predict(X)
print(prediction)
# -
index = np.argmax(prediction[0])
rows = int(index/5)
cols = index%5
print("Move : ", (rows, cols))
print("Win chance :", prediction[1])
import tensorflow as tf
filepath = 'model'
#save the model
model.save('model')
"""
import tensorflow as tf
#load the model
loaded_model = tf.keras.models.load_model('model')
"""
| code/algos/nn/AGZ.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Develop your model
# +
import os
from six.moves import xrange # pylint: disable=redefined-builtin
import tensorflow as tf
from tensorflow.examples.tutorials.mnist import input_data
from tensorflow.examples.tutorials.mnist import mnist
INPUT_DATA_DIR = '/tmp/tensorflow/mnist/input_data/'
MAX_STEPS = 1000
BATCH_SIZE = 100
LEARNING_RATE = 0.3
HIDDEN_1 = 128
HIDDEN_2 = 32
# HACK: Ideally we would want to have a unique subpath for each instance of the job, but since we can't
# we are instead appending HOSTNAME to the logdir
LOG_DIR = os.path.join(os.getenv('TEST_TMPDIR', '/tmp'),
'tensorflow/mnist/logs/fully_connected_feed/', os.getenv('HOSTNAME', ''))
class TensorflowModel():
def train(self, **kwargs):
tf.logging.set_verbosity(tf.logging.ERROR)
self.data_sets = input_data.read_data_sets(INPUT_DATA_DIR)
self.images_placeholder = tf.placeholder(
tf.float32, shape=(BATCH_SIZE, mnist.IMAGE_PIXELS))
self.labels_placeholder = tf.placeholder(tf.int32, shape=(BATCH_SIZE))
logits = mnist.inference(self.images_placeholder,
HIDDEN_1,
HIDDEN_2)
self.loss = mnist.loss(logits, self.labels_placeholder)
self.train_op = mnist.training(self.loss, LEARNING_RATE)
self.summary = tf.summary.merge_all()
init = tf.global_variables_initializer()
self.sess = tf.Session()
self.summary_writer = tf.summary.FileWriter(LOG_DIR, self.sess.graph)
self.sess.run(init)
data_set = self.data_sets.train
for step in xrange(MAX_STEPS):
images_feed, labels_feed = data_set.next_batch(BATCH_SIZE, False)
feed_dict = {
self.images_placeholder: images_feed,
self.labels_placeholder: labels_feed,
}
_, loss_value = self.sess.run([self.train_op, self.loss],
feed_dict=feed_dict)
if step % 100 == 0:
print("At step {}, loss = {}".format(step, loss_value))
summary_str = self.sess.run(self.summary, feed_dict=feed_dict)
self.summary_writer.add_summary(summary_str, step)
self.summary_writer.flush()
# -
# ## Train your model inside notebook
local_train = TensorflowModel()
local_train.train()
# ## Remote train
# > Note: `__file__` won't work inside notebook, please don't execute follow block. Instead, we create a python file `distributed_training.py` to run.
#
#
# ```py
# if __name__ == '__main__':
# if os.getenv('FAIRING_RUNTIME', None) is None:
# from kubeflow import fairing
# AWS_ACCOUNT_ID=fairing.cloud.aws.guess_account_id()
# AWS_REGION='us-west-2'
# DOCKER_REGISTRY = '{}.dkr.ecr.{}.amazonaws.com'.format(AWS_ACCOUNT_ID, AWS_REGION)
#
# fairing.config.set_preprocessor('python', input_files=[__file__])
# fairing.config.set_builder(name='append', registry=DOCKER_REGISTRY,
# base_image='tensorflow/tensorflow:1.14.0-py3')
# fairing.config.set_deployer(
# name='tfjob', worker_count=1, ps_count=1)
# fairing.config.run()
# else:
# remote_train = TensorflowModel()
# remote_train.train()
# ```
# ## Run this comamnd to create a Tensoflow Flow Job (TFJob)
#
# You can check file `distributed_traing.py` to see details.
# !python ./distributed_training.py
| notebooks/02_Fairing/02_03_fairing_distributed_training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: clouds113_kernel
# language: python
# name: clouds113_kernel
# ---
# ### Tests concerning the speed of training
# +
# Ran with 800GB (750GB should also be fine)
import sys
import numpy as np
import time
import pandas as pd
import matplotlib.pyplot as plt
import os
import copy
import gc
#Import sklearn before tensorflow (static Thread-local storage)
from sklearn.preprocessing import StandardScaler
import tensorflow as tf
from tensorflow.keras.models import load_model
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout
from tensorflow.keras.regularizers import l1_l2
from tensorflow.keras import backend as K
from tensorflow.keras.layers import Activation
# For Leaky_ReLU:
from tensorflow import nn
t0 = time.time()
path = '/pf/b/b309170'
path_figures = path + '/workspace_icon-ml/cloud_cover_parameterization/grid_cell_based_QUBICC_R02B05/figures'
path_model = path + '/workspace_icon-ml/cloud_cover_parameterization/grid_cell_based_QUBICC_R02B05/saved_models'
path_data = path + '/my_work/icon-ml_data/cloud_cover_parameterization/grid_cell_based_QUBICC_R02B05/based_on_var_interpolated_data'
# Add path with my_classes to sys.path
sys.path.insert(0, path + '/workspace_icon-ml/cloud_cover_parameterization/')
# Reloading custom file to incorporate changes dynamically
import importlib
import my_classes
importlib.reload(my_classes)
from my_classes import read_mean_and_std
from my_classes import TimeOut
# Minutes per fold
timeout = 2120
# For logging purposes
days = 'all_days'
# Maximum amount of epochs for each model
epochs = 30
# Set seed for reproducibility
seed = 10
tf.random.set_seed(seed)
# For store_mean_model_biases
VERT_LAYERS = 31
gpus = tf.config.experimental.list_physical_devices('GPU')
# tf.config.experimental.set_visible_devices(gpus[3], 'GPU')
# -
# Prevents crashes of the code
physical_devices = tf.config.list_physical_devices('GPU')
tf.config.set_visible_devices(physical_devices[0], 'GPU')
# Allow the growth of memory Tensorflow allocates (limits memory usage overall)
for gpu in gpus:
tf.config.experimental.set_memory_growth(gpu, True)
scaler = StandardScaler()
# ### Load the data
# +
# input_narval = np.load(path_data + '/cloud_cover_input_narval.npy')
# input_qubicc = np.load(path_data + '/cloud_cover_input_qubicc.npy')
# output_narval = np.load(path_data + '/cloud_cover_output_narval.npy')
# output_qubicc = np.load(path_data + '/cloud_cover_output_qubicc.npy')
# -
input_data = np.concatenate((np.load(path_data + '/cloud_cover_input_narval.npy'),
np.load(path_data + '/cloud_cover_input_qubicc.npy')), axis=0)
output_data = np.concatenate((np.load(path_data + '/cloud_cover_output_narval.npy'),
np.load(path_data + '/cloud_cover_output_qubicc.npy')), axis=0)
samples_narval = np.load(path_data + '/cloud_cover_output_narval.npy').shape[0]
(samples_total, no_of_features) = input_data.shape
(samples_total, no_of_features)
# *Temporal cross-validation*
#
# Split into 2-weeks increments (when working with 3 months of data). It's 25 day increments with 5 months of data. <br>
# 1.: Validate on increments 1 and 4 <br>
# 2.: Validate on increments 2 and 5 <br>
# 3.: Validate on increments 3 and 6
#
# --> 2/3 training data, 1/3 validation data
# +
training_folds = []
validation_folds = []
two_week_incr = samples_total//6
for i in range(3):
# Note that this is a temporal split since time was the first dimension in the original tensor
first_incr = np.arange(samples_total//6*i, samples_total//6*(i+1))
second_incr = np.arange(samples_total//6*(i+3), samples_total//6*(i+4))
validation_folds.append(np.append(first_incr, second_incr))
training_folds.append(np.arange(samples_total))
training_folds[i] = np.delete(training_folds[i], validation_folds[i])
# -
# ### Define the model
# Activation function for the last layer
def lrelu(x):
return nn.leaky_relu(x, alpha=0.01)
# +
# Create the model
model = Sequential()
# First hidden layer
model.add(Dense(units=256, activation=lrelu, input_dim=no_of_features,
kernel_regularizer=l1_l2(l1=0.000162, l2=0.007437)))
# Second hidden layer
model.add(Dense(units=256, activation=lrelu, kernel_regularizer=l1_l2(l1=0.000162, l2=0.007437)))
model.add(Dropout(0.184124)) # We drop 18% of the hidden nodes
# Output layer
model.add(Dense(1, activation='linear', kernel_regularizer=l1_l2(l1=0.000162, l2=0.007437)))
# -
# Preliminary baselines
# +
# # This would be the loss of a NN which outputs zeros everywhere
# np.mean(np.array(output_data)**2)
# +
# # This would be the loss of a NN which outputs the best constant value everywhere
# constant_mean = np.mean(np.array(output_data))
# np.mean((np.array(output_data) - constant_mean)**2)
# +
# # Freeing up memory (~46 GB). Memory usage after this cell: 251 GB
# del input_narval, input_qubicc, output_narval, output_qubicc
# gc.collect()
# + [markdown] jupyter={"outputs_hidden": true}
# ### 3-fold cross-validation
# +
# By decreasing timeout we make sure every fold gets the same amount of time
# After all, data-loading took some time (Have 3 folds, 60 seconds/minute)
# timeout = timeout - 1/3*1/60*(time.time() - t0)
timeout = timeout - 1/60*(time.time() - t0)
t0 = time.time()
#We loop through the folds
for i in range(1,2):
filename = 'cross_validation_cell_based_fold_%d'%(i+1)
#Standardize according to the fold
scaler.fit(input_data[training_folds[i]])
#Load the data for the respective fold and convert it to tf data
input_train = scaler.transform(input_data[training_folds[i]])
input_valid = scaler.transform(input_data[validation_folds[i]])
output_train = output_data[training_folds[i]]
output_valid = output_data[validation_folds[i]]
# Clear memory (Reduces memory requirement to 151 GB)
del input_data, output_data, first_incr, second_incr, validation_folds, training_folds
gc.collect()
# # Column-based: batchsize of 128
# # Possibly better to use .apply(tf.data.experimental.copy_to_device("/gpu:0")) before prefetch
# train_ds = tf.data.Dataset.zip((tf.data.Dataset.from_tensor_slices(input_train),
# tf.data.Dataset.from_tensor_slices(output_train))) \
# .shuffle(1000, seed=seed) \
# .batch(batch_size=1028, drop_remainder=True) \
# .prefetch(1)
# # Clear memory
# del input_train, output_train
# gc.collect()
# # No need to add prefetch.
# # tf data with batch_size=10**5 makes the validation evaluation 10 times faster
# valid_ds = tf.data.Dataset.zip((tf.data.Dataset.from_tensor_slices(input_valid),
# tf.data.Dataset.from_tensor_slices(output_valid))) \
# .batch(batch_size=10**5, drop_remainder=True)
# # Clear memory (Reduces memory requirement to 151 GB)
# del input_valid, output_valid
# gc.collect()
#Feed the model
model.compile(
optimizer=tf.keras.optimizers.Adam(learning_rate=0.008726, epsilon=0.1),
loss=tf.keras.losses.MeanSquaredError()
)
# #Train the model
# # time_callback = TimeOut(t0, timeout*(i+1))
# time_callback = TimeOut(t0, timeout)
# history = model.fit(train_ds, epochs=epochs, verbose=2, validation_data=valid_ds,
# callbacks=[time_callback])
# # history = model.fit(train_ds, epochs=epochs, validation_data=valid_ds, callbacks=[time_callback])
# #Save the model
# #Serialize model to YAML
# model_yaml = model.to_yaml()
# with open(os.path.join(path_model, filename+".yaml"), "w") as yaml_file:
# yaml_file.write(model_yaml)
# #Serialize model and weights to a single HDF5-file
# model.save(os.path.join(path_model, filename+'.h5'), "w")
# print('Saved model to disk')
# #Plot the training history
# if len(history.history['loss']) > len(history.history['val_loss']):
# del history.history['loss'][-1]
# pd.DataFrame(history.history).plot(figsize=(8,5))
# plt.grid(True)
# plt.ylabel('Mean Squared Error')
# plt.xlabel('Number of epochs')
# plt.savefig(os.path.join(path_figures, filename+'.pdf'))
# with open(os.path.join(path_model, filename+'.txt'), 'a') as file:
# file.write('Results from the %d-th fold\n'%(i+1))
# file.write('Training epochs: %d\n'%(len(history.history['val_loss'])))
# file.write('Weights restored from epoch: %d\n\n'%(1+np.argmin(history.history['val_loss'])))
# -
# ### Training with 800 Mio samples (three-hourly QUBICC data and stricter class balancing)
# +
# Column-based: batchsize of 128
# Possibly better to use .apply(tf.data.experimental.copy_to_device("/gpu:0")) before prefetch
train_ds = tf.data.Dataset.zip((tf.data.Dataset.from_tensor_slices(input_train),
tf.data.Dataset.from_tensor_slices(output_train))) \
.shuffle(100000, seed=seed) \
.batch(batch_size=1028, drop_remainder=True) \
.prefetch(1)
# shuffle 1000 and bs 2056 increases...
# shuffle 1000 and bs 1028 increases... but it starts to decrease after a while!
# -
history = model.fit(train_ds, epochs=5, verbose=1)
# +
# Column-based: batchsize of 128
# Possibly better to use .apply(tf.data.experimental.copy_to_device("/gpu:0")) before prefetch
train_ds = tf.data.Dataset.zip((tf.data.Dataset.from_tensor_slices(input_train),
tf.data.Dataset.from_tensor_slices(output_train))) \
.shuffle(1000000, seed=seed) \
.batch(batch_size=1028, drop_remainder=True) \
.prefetch(1)
# shuffle 1000 and bs 2056 increases...
# shuffle 1000 and bs 1028 increases... but it starts to decrease after a while!
# -
history = model.fit(train_ds, epochs=5, verbose=1)
# +
# Column-based: batchsize of 128
# Possibly better to use .apply(tf.data.experimental.copy_to_device("/gpu:0")) before prefetch
train_ds = tf.data.Dataset.zip((tf.data.Dataset.from_tensor_slices(input_train),
tf.data.Dataset.from_tensor_slices(output_train))) \
.shuffle(100000, seed=seed) \
.batch(batch_size=512, drop_remainder=True) \
.prefetch(1)
# shuffle 1000 and bs 2056 increases...
# shuffle 1000 and bs 1028 increases... but it starts to decrease after a while!
# -
history = model.fit(train_ds, epochs=5, verbose=1)
# ### Training with two-hourly QUBICC data is quite infeasible:
# Batch size of 1028 is faster than one of 2048!
# So batch size of 1028 is best. A shuffle buffer of 100000 is best.
# +
# Column-based: batchsize of 128
# Possibly better to use .apply(tf.data.experimental.copy_to_device("/gpu:0")) before prefetch
train_ds = tf.data.Dataset.zip((tf.data.Dataset.from_tensor_slices(input_train),
tf.data.Dataset.from_tensor_slices(output_train))) \
.shuffle(100000, seed=seed) \
.batch(batch_size=1028, drop_remainder=True) \
.prefetch(1)
# shuffle 1000 and bs 2056 increases...
# shuffle 1000 and bs 1028 increases... but it starts to decrease after a while!
# + jupyter={"outputs_hidden": true}
history = model.fit(train_ds, epochs=5, verbose=1)
# +
# Column-based: batchsize of 128
# Possibly better to use .apply(tf.data.experimental.copy_to_device("/gpu:0")) before prefetch
train_ds = tf.data.Dataset.zip((tf.data.Dataset.from_tensor_slices(input_train),
tf.data.Dataset.from_tensor_slices(output_train))) \
.shuffle(100000, seed=200) \
.batch(batch_size=1028, drop_remainder=True) \
.prefetch(1)
# shuffle 1000 and bs 2056 increases...
# shuffle 1000 and bs 1028 increases... but it starts to decrease after a while!
# -
history = model.fit(train_ds, epochs=5, verbose=1)
# +
# Shuffle after every epoch
train_ds = tf.data.Dataset.zip((tf.data.Dataset.from_tensor_slices(input_train),
tf.data.Dataset.from_tensor_slices(output_train))) \
.shuffle(100000, seed=seed, reshuffle_each_iteration=True) \
.batch(batch_size=1028, drop_remainder=True) \
.prefetch(1)
# shuffle 1000 and bs 2056 increases...
# shuffle 1000 and bs 1028 increases... but it starts to decrease after a while!
# -
history = model.fit(train_ds, epochs=5, verbose=1)
# +
# Shuffle after every epoch
train_ds = tf.data.Dataset.zip((tf.data.Dataset.from_tensor_slices(input_train),
tf.data.Dataset.from_tensor_slices(output_train))) \
.shuffle(100000, seed=200, reshuffle_each_iteration=True) \
.batch(batch_size=1028, drop_remainder=True) \
.prefetch(1)
# shuffle 1000 and bs 2056 increases...
# shuffle 1000 and bs 1028 increases... but it starts to decrease after a while!
# -
history = model.fit(train_ds, epochs=5, verbose=1)
# +
# Column-based: batchsize of 1028
# Possibly better to use .apply(tf.data.experimental.copy_to_device("/gpu:0")) before prefetch
train_ds = tf.data.Dataset.zip((tf.data.Dataset.from_tensor_slices(input_train),
tf.data.Dataset.from_tensor_slices(output_train))) \
.shuffle(1000, seed=seed) \
.batch(batch_size=1028, drop_remainder=True) \
.prefetch(1)
# shuffle 1000 and bs 2056 increases...
# shuffle 1000 and bs 1028 increases... but it starts to decrease after a while!
# -
history = model.fit(train_ds, epochs=1, verbose=1)
# Column-based: batchsize of 128
# Possibly better to use .apply(tf.data.experimental.copy_to_device("/gpu:0")) before prefetch
train_ds = tf.data.Dataset.zip((tf.data.Dataset.from_tensor_slices(input_train),
tf.data.Dataset.from_tensor_slices(output_train))) \
.shuffle(10, seed=seed) \
.batch(batch_size=1028, drop_remainder=True) \
.prefetch(1)
history = model.fit(train_ds, epochs=1, verbose=1)
# Column-based: batchsize of 128
# Possibly better to use .apply(tf.data.experimental.copy_to_device("/gpu:0")) before prefetch
train_ds = tf.data.Dataset.zip((tf.data.Dataset.from_tensor_slices(input_train),
tf.data.Dataset.from_tensor_slices(output_train))) \
.shuffle(1000, seed=seed) \
.batch(batch_size=2048, drop_remainder=True) \
.prefetch(1)
history = model.fit(train_ds, epochs=1, verbose=1)
# +
# Column-based: batchsize of 128
# Possibly better to use .apply(tf.data.experimental.copy_to_device("/gpu:0")) before prefetch
train_ds = tf.data.Dataset.zip((tf.data.Dataset.from_tensor_slices(input_train),
tf.data.Dataset.from_tensor_slices(output_train))) \
.shuffle(10000, seed=seed) \
.batch(batch_size=1028, drop_remainder=True) \
.prefetch(1)
# shuffle 1000 and bs 2056 increases...
# shuffle 1000 and bs 1028 increases... but it starts to decrease after a while!
# -
history = model.fit(train_ds, epochs=1, verbose=1)
# +
# Column-based: batchsize of 128
# Possibly better to use .apply(tf.data.experimental.copy_to_device("/gpu:0")) before prefetch
train_ds = tf.data.Dataset.zip((tf.data.Dataset.from_tensor_slices(input_train),
tf.data.Dataset.from_tensor_slices(output_train))) \
.shuffle(100000, seed=seed) \
.batch(batch_size=1028, drop_remainder=True) \
.prefetch(1)
# shuffle 1000 and bs 2056 increases...
# shuffle 1000 and bs 1028 increases... but it starts to decrease after a while!
# -
history = model.fit(train_ds, epochs=1, verbose=1)
# +
# Different seed
# Possibly better to use .apply(tf.data.experimental.copy_to_device("/gpu:0")) before prefetch
train_ds = tf.data.Dataset.zip((tf.data.Dataset.from_tensor_slices(input_train),
tf.data.Dataset.from_tensor_slices(output_train))) \
.shuffle(100000, seed=100) \
.batch(batch_size=1028, drop_remainder=True) \
.prefetch(1)
# shuffle 1000 and bs 2056 increases...
# shuffle 1000 and bs 1028 increases... but it starts to decrease after a while!
# -
history = model.fit(train_ds, epochs=1, verbose=1)
# Try to shuffle after batching
train_ds = tf.data.Dataset.zip((tf.data.Dataset.from_tensor_slices(input_train),
tf.data.Dataset.from_tensor_slices(output_train))) \
.batch(batch_size=1028, drop_remainder=True) \
.shuffle(100000, seed=seed) \
.prefetch(1)
history = model.fit(train_ds, epochs=1, verbose=1)
# +
# Shuffle dataset with numpy and train afterwards
permuted_indices = np.random.permutation(np.arange(input_train.shape[0]))
train_ds = tf.data.Dataset.zip((tf.data.Dataset.from_tensor_slices(input_train[permuted_indices]),
tf.data.Dataset.from_tensor_slices(output_train[permuted_indices]))) \
.batch(batch_size=1028, drop_remainder=True) \
.prefetch(1)
# -
history = model.fit(train_ds, epochs=1, verbose=1)
# +
# Is a batchsize of 512 really much slower than one of 1028? Yes
# Column-based: batchsize of 128
# Possibly better to use .apply(tf.data.experimental.copy_to_device("/gpu:0")) before prefetch
train_ds = tf.data.Dataset.zip((tf.data.Dataset.from_tensor_slices(input_train),
tf.data.Dataset.from_tensor_slices(output_train))) \
.shuffle(10, seed=seed) \
.batch(batch_size=512, drop_remainder=True) \
.prefetch(1)
# + jupyter={"outputs_hidden": true}
history = model.fit(train_ds, epochs=1, verbose=1)
# +
# Column-based: batchsize of 128
# Possibly better to use .apply(tf.data.experimental.copy_to_device("/gpu:0")) before prefetch
train_ds = tf.data.Dataset.zip((tf.data.Dataset.from_tensor_slices(input_train),
tf.data.Dataset.from_tensor_slices(output_train))) \
.shuffle(100000, seed=seed) \
.batch(batch_size=1028, drop_remainder=True) \
.prefetch(1)
# shuffle 1000 and bs 2056 increases...
# shuffle 1000 and bs 1028 increases... but it starts to decrease after a while!
# -
# Try multiple epochs
history = model.fit(train_ds, epochs=3, verbose=1)
# Try multiple epochs
history = model.fit(train_ds, epochs=6, verbose=1)
| q1_cell_based_qubicc_r2b5/source_code/tests/hourly_vs_two_hourly_vs_three_hourly.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # Review (복습)
# - Machine Learning : (True) Function Approxmation (함수 근사)
# - Based on Data.
# - f(x) = y
# - 함수가 x와 y의 관계를 설명해줄 수 있는지
# - 전체 함수에서 f를 근사하라고 하는 건 너무 광범위 하다.
# - 특수한 함수들의 모임에 대해서 생각을 했다.
# - 제한시켜주는 공간에서 함
# - RF
# - Tree
# - 신경망 <br>
# - 실제 f는 어딘가에 살고 있을 것이다.
# - 그러면 어떤 집합을 선택해야
# - 그걸 다루는 게 통계적 이론이다. Statistical Learning Theory
# - Cross Validation
#
# - obj : f를 찾아라
# - 1) 제한을 시킨다 = Model을 선택한다.
# - ex) y = ax + b
# - 2) 제한시킨 함수들의 공간에서 하나를 뽑아낸다.
# - a_hat, b_hat을 추정한다.
# - 여러 가지가 있을 수 있다.
# - 그중에서 a, b를 효율적으로 추정하기 위해서 __'경사하강법'__을 사용한다.
# - 3) 평가를 내란디.
# - 제한을 시켰는데, 그 가정이 얼마나 타당했는가?
# - 타당했다면, 그 함수가 좋아야 한다. (실제 f랑 비슷해야 한다.)
# - 1단계의 가정이 얼마나 타당했는지 평가를 한다.
# - Cross Validation(자주 사용), R-Squared
#
# - Cross Validation의 목적
# - 1) 가장 좋은 가정을 선택 (Which Model?)
# - CV는 Train set을 사용한다.
#
# - Test-Set의 목적
# - 1) 실제로 현실에서 얼마나 좋을까? 검증
# - CV로 검증을 한 다음에 최종적으로 딱! 한번 Test
#
# - CV의 방법
# - 1) Validation set approach
# - 2) k-fold cross validation
# - 3) LooCV
#
# - Case 1
# - Train set을 가지고, 3000번을 training했는데
# - 모델 A(3000개의 모델 중 선택)
# - training error = 0, validation error = 0.1
# - Test error = 100
# - 이런 상황을 해결하는 방법은 데이터를 진짜 많이 모으면 해결이 될 것이다.
# - 아니라고 생각해서 실험을 10만 번을 더해서, 모델 Z를 뽑음
# - 모델 Z(10만 개의 모델 중 선택)
# - tr = 0.01
# - val = 0.3
# - test error = 10
# - 쓰면 안 된다. 비교하는데 test set을 썼기에, 오염됐다.
# - (철학적으로 봤을 때) test error 높아질 것이다.
# - (실제 세계에서는 그렇게 높아지지 않겠지만...)
# - 근데 현업에서는 어느 정도 허용, 논문에서는 불허
#
# - Case 2
# - case 1을 다시 생각해보기
# - 실험을 3000번 했는데, test error가 높았다.
# - test set과 training set을 섞었다.
#
# - test err = generalization err
# - statistics vs. machine learning
# # 1. Load Dataset
# - 비식별화
# - input의 의미를 알 수는 없다.
# - task : target을 예측
# - columns(4993) > data row (4459)
# - 컬럼이 데이터 보다 더 많은 특이한 경우
# - <E
# +
data <- read.csv("../../../../Downloads/Kaggle/Santander Value Prediction Challenge//train.csv")
str(data)
# -
# # 2. Missing Data
# - 빼거나
# - 채우거나
#
# - [김재광](http://mathsci.kaist.ac.kr/newsletter/article-tags/%EA%B9%80%EC%9E%AC%EA%B4%91/)
sum(is.na(data))
# # 3. Separate Data
# +
set.seed(1)
train <- sample(1:nrow(data), 0.7 * nrow(data))
set.seed(1)
valid <- sample(setdiff(1:nrow(data), train), 0.15 * nrow(data))
test <- setdiff(1:nrow(data), c(train, valid))
# -
length(train)
length(valid)
length(test)
# ## RMSLE 1
# - The evaluation metric for this competition is Root Mean Squared Logarithmic Error.
RMSLE <- function(pred, real){
err <- sqrt(mean((log(pred + 1) - log(real + 1))^2))
return (err)
}
# ## RMSLE 2
# - The evaluation metric for this competition is Root Mean Squared Logarithmic Error.
RMSLE <- function(pred, real){
err <- sqrt(mean((log(pmax(0, pred) + 1) - log(real + 1))^2))
return (err)
}
# ## Random Forest
# - ranger package
# - predict할 때, 특이한 방법이 있다.
# - 마지막에 $표시를 붙임
library(ranger)
Model <- ranger(target~. -ID, data= data[train, ], mtry = 300) # target을 제외한 컬럼을 사용하는데 그 중에서도 ID 컬럼은 예측하는데 필요 없다.
Model
pred <- predict(Model, data[valid, ])$predictions
RMSLE(pred, data$target[valid])
# +
params <- c(500, 2000, 4000)
for (i in 1:5){
Model <- ranger(target~. -ID,
data= data[train, ],
mtry = params[i])
pred <- predict(Model, data[valid, ])$predictions
err <- RMSLE(pred, data$target[valid])
cat("mtry가", params[i], "일 때,", "error는", err)
}
# -
# - mtry = 70, err = 1.742246
# - mtry = 300, err = 1.731769
# - mtry = , err =
# - mtry = , err =
# - mtry = , err =
# ## R-squared
# - 에러가 높은지 낮은지를 객관적으로 알 수 있는 방법
mean(data$target[train])
# ## "가장" 간단한 모델
# - y = a
# - y가 x에 전혀 영향을 받지 않는 상수(Constant)다.
# - mean이 나온다.
RMSLE(mean(data$target[train]), data$target[valid])
# - 어떤 모델이라도 위의 수치보다 작아야 한다.
# - 평균 던져서 만들 모델이 2.1 이니까 <Br>
# <Br>
# - 객관적으로 알려면?
# - Leaderboard를 보면 Score를 알 수 있다.
#
# ## Boosting
# ## Regularization
# - 1) 모델의 Capacity를 낮춰준다.
# - 2) 우리가 사용한 모델의 Variance를 낮춰준다.
# - 3) Tranin set에 민감한 정도를 낮춰준다.
# - obj : overfitting을 막기 위한 테크닉
# - 테크닉이 엄청나게 많다.
# - L1, L2 Regularization
# - 가장 고전적인 방식이자, 지금도 많이 쓰는 방식<br>
# <br>
# - 모델의 capacity가 높다는 건 나쁜 소리는 아니다.
# - capacity가 높은 모형을 그대로 쓸 건데,
# - overfitting이 안 됐으면 좋겠다.<br>
# <br>
# - cost 함수를 바꿔버리면 된다.
# - 1) cost = (Model이 Training set을 얼마나 설명 못하는가) ... "설"
# - 2) cost = (Model이 얼마나 복잡한가?) ... "복"
# - 3) Penelizaed Method
# - 1만 사용하면 overfitting이 될 가능성이 높고, 2만 사용하면 상수가 된다.
# - 1, 2에 가중치를 준 다음에 더한다
# - 모델이 복잡하면 패널티를 주겠다.<br>
# <br>
# - hyperparmeter
# - 가장 적당한 알파와 베타를 선택하는 모형
# - __lambda__
# - 모델의 복잡도(capacity)를 얼마나 제한시켜줄 거냐<br>
# <br>
# - 설명력을 측정하는 방법
# - (y - y_hat)^2<br>
# <br>
# - 복잡도를 측정하는 방법
# - 0하고 가까운 값이면 좋다. 복잡도가 내려감!
# - 1) 절대값 ... L1 Regulralization ... Lasso
# - x값 앞에 있는 절대값들을 다 더한다.<br>
# <br>
# - 2) 제곱값 ... L2 Regulralization ... Ridge
# - 제곱한다.
#
#
# +
# install.packages("glmnetUtils")
# -
library(glmnetUtils)
# +
Model <- glmnet(formula = target~. -ID,
data = data[train, ],
lambda = 0.01) # lambda를 작게 잡으면.. ()
pred <- predict(Model, data[valid, ])
RMSLE(pred, data$target[valid])
# -
hist(pred)
# - Nan값이 나와서 (1시간 30분)
# - RMSLE를 다시 변경
# +
Model <- glmnet(formula = target~. -ID,
data = data[train, ],
lambda = 10) # lambda를 작게 잡으면.. ()
pred <- predict(Model, data[valid, ])
RMSLE(pred, data$target[valid])
# -
# - 평균쓰는 것(2.1) 보다 겁나 높게 나옴(9.16) ... (1시간 35분)
# +
Model <- glmnet(formula = target~. -ID,
data = data[train, ],
lambda = 500000) # lambda를 작게 잡으면.. ()
pred <- predict(Model, data[valid, ])
RMSLE(pred, data$target[valid])
# -
# - Regularization의 위력
hist(pred)
# +
k <- (1:10) * 100000
for (i in 1:100){
Model <- glmnet(formula = target~. -ID,
data = data[train, ],
lambda = k[i]) # lambda를 작게 잡으면.. ()
pred <- predict(Model, data[valid, ])
cat(k[i], "일 때 오차는" , RMSLE(pred, data$target[valid]), "\n")
}
# +
Model <- glmnet(formula = target~. -ID,
data = data[train, ],
lambda = 300000,
alpha = 0.5) # lambda를 작게 잡으면.. ()
pred <- predict(Model, data[valid, ])
RMSLE(pred, data$target[valid])
# -
# ## Boost
# - [lightgbm](https://github.com/Microsoft/LightGBM)
# - [catboost](https://tech.yandex.com/catboost/doc/dg/concepts/r-installation-docpage/)
#
# ## Xgboost
# install.packages("xgboost")
library(xgboost)
# +
X <- model.matrix(target~. -ID, data = data)
Y <- data$target
Model <- xgboost(data = X[train, ],
label = Y[train],
params = list(eta = 0.3,
gamma = 0.1),
nrounds = 100
)
pred <- predict(Model, X[valid, ])
RMSLE(pred, Y[valid])
# -
min(pred)
hist(pred)
# ### xgb.train
?xgb.train
# +
dtrain <- xgb.DMatrix(X[train, ], label = Y[train])
dvalid <- xgb.DMatrix(X[valid, ], label = Y[valid])
Model <- xgb.train(data = dtrain,
params = list(eta = 0.3),
watchlist = list(train = dtrain,
eval = dvalid),
nrounds = 100)
# -
# #### 뭘 뽑아낼 수 있는지 파악하기
attributes(Model)
attributes(Model$evaluation_log)
plot(Model$evaluation_log)
plot(Model$evaluation_log$train_rmse, type = 'l')
plot(Model$evaluation_log$eval_rmse, type = 'l', lwd = 5)
# 2시간 25분
Model <- xgb.train(data = dtrain,
params = list(eta = 0.3),
watchlist = list(train = dtrain,
eval = dvalid),
nrounds = 99999999,
early_stopping_rounds = 20)
pred <- predict(Model, X[valid, ])
RMSLE(pred, Y[valid])
# +
hyper <- list(eta = 0.01,
gamma = 0.1,
subsample = 0.6,
colsample_bytree = 0.5)
Model <- xgb.train(data = dtrain,
params = hyper,
watchlist = list(train = dtrain,
eval = dvalid),
nrounds = 99999999,
early_stopping_rounds = 20)
# -
pred <- predict(Model, X[valid, ])
RMSLE(pred, Y[valid])
# +
hyper <- list(objective = "reg:linear",
booster = "gbtree",
eval_metric = "rmse",
nthread = 4,
eta = 0.01,
max_depth = 20,
min_child_weight = 62,
gamma = 5,
subsample = 0.9,
colsample_bytree = 0.3,
colsample_bylevel = 0.305,
alpha = 0,
lambda = 0.8832217
)
Model <- xgb.train(data = dtrain,
params = hyper,
watchlist = list(train = dtrain,
eval = dvalid),
nrounds = 99999999,
early_stopping_rounds = 30)
# -
pred <- predict(Model, X[valid, ])
RMSLE(pred, Y[valid])
# +
hyper <- list(objective = "reg:linear",
booster = "gbtree",
eval_metric = "rmse",
nthread = 4,
eta = 0.01,
max_depth = 20,
min_child_weight = 62,
gamma = 5,
subsample = 0.9,
colsample_bytree = 0.3,
colsample_bylevel = 0.305,
alpha = 0,
lambda = 0.8832217
)
Model <- xgb.train(data = dtrain,
params = hyper,
watchlist = list(train = dtrain,
eval = dvalid),
nrounds = 99999999,
early_stopping_rounds = 400)
# -
pred <- predict(Model, X[valid, ])
RMSLE(pred, Y[valid])
# +
for(i in 1:100){
hyper <- list(objective = "reg:linear",
booster = "gbtree",
eval_metric = "rmse",
nthread = 4,
eta = sample(seq(0.01, 0.1, length = 100), 1),
max_depth = sample(1:30, 1),
min_child_weight = 62,
gamma = 5,
subsample = 0.9,
colsample_bytree = 0.3,
colsample_bylevel = 0.305,
alpha = 0,
lambda = 0.8832217
)
Model <- xgb.train(data = dtrain,
params = hyper,
watchlist = list(train = dtrain,
eval = dvalid),
nrounds = 99999999,
early_stopping_rounds = 30)
pred <- predict(Model, X[valid, ])
RMSLE(pred, Y[valid])
data.frame(hyper), Model$evaluation_log
}
# -
| study/machine-learning/r/ML-6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment No - 1
# # Question !
# ### Write a Program to identify sub list [1,1,5] is there in the given list in the same order,If Yes print "it's a Match" if No then print "it's Gone" in function.
# # Answer !
# +
# Program to check the list contains elements of another list
# List1
List1 = [2,3,6,7,1,3,1,6,7,5,8]
# List2
List2 = [1,1,5]
check = all(item in List1 for item in List2)
if check is True:
print("it's a Match".format(List1, List2))
else :
print("it's Gone")
# +
# Program to check the list contains elements of another list
# List1
List1 = [2,3,1,6,7,3,1,6,7,8]
# List2
List2 = [1,1,5]
check = all(item in List1 for item in List2)
if check is True:
print("it's a Match".format(List1, List2))
else :
print("it's Gone")
# -
# # Thank You !
# # Assignment No - 2
# # Question !
# ### Make a Function for Prime Numbers and use filter to filter out all the Prime Numbers from 1 - 2500
# +
lower = int(input("Enter lower range: "))
upper = int(input("Enter upper range: "))
for num in range(lower,upper + 1):
if num > 1:
for i in range(2,num):
if (num % i) == 0:
break
else:
print(num)
# -
# # Thank You !
# # Assignment No - 3
# # Question !
# ### Make a Lamba Function Capitalizing the whole Sentence Passed during Arguments
#
# ### and Map all the sentences in the List,with the Lambda Function.
message = "Hey this is Ashwin"
parts = message.split(" ")
capitalized_parts = [p.capitalize() for p in parts]
capitalized_message = " ".join(capitalized_parts)
message = "Hey this is Ashwin"
capitalized_message = " ".join([
word.capitalize()
for word in message.split(" ")
])
print(capitalized_message)
# # Thank You !
| Day-5 Assignment No-1&2&3 05.09.2020.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="r336aHzrjifv"
import numpy as np
from numpy.random import randint
from numpy.random import choice
from math import pow
from fractions import Fraction
# + id="pemm_gcrNrKQ"
def GibbsSampler(Dna, k, t, N):
motifs = SelectRandomKmers(Dna, k, t)
best_motifs = motifs
for j in range(N):
i = randint(t)
motifs.pop(i)
profile = GenerateProfile(motifs, k)
motif_i = ProfileRandomlyGeneratedKmer(Dna[i], profile, k)
motifs.insert(i, motif_i)
if Score(motifs, k) < Score(best_motifs, k):
best_motifs = motifs
return best_motifs
# + id="rni-u3cXjrdQ"
def SelectRandomKmers(Dna, k, t):
random_kmers = []
for dna_string in Dna:
random_kmers.append(FindAllKmers(dna_string, k)[randint(0, len(dna_string) - k + 1)]) #upper bound not included
return random_kmers
# + id="s5c1USNsNPXv"
def GenerateProfile(motifs, k):
profile = np.zeros((4,k))
motifs_array = DnaToArray(motifs)
for i in range(k):
frequency_dict = {'A':0, 'C':0, 'G':0, 'T':0}
for j in range(len(np.unique(motifs_array[:,i], return_counts=True)[0])):
frequency_dict[np.unique(motifs_array[:,i], return_counts=True)[0][j]] = np.unique(motifs_array[:,i], return_counts=True)[1][j]
profile[:,i] = np.array(list(frequency_dict.values()))
profile = profile + 1 #Laplace's rule of succession
profile = profile / (len(motifs_array) + 4)
return profile
# + id="-HDYUBBlNkTx"
def KmerProbability(profile, kmer):
probability = 1
for nucleotide in enumerate(kmer):
probability = probability * profile[NucleotideIndex(nucleotide[1])][nucleotide[0]]
return probability
# + id="f2Fq4UUbX1pY"
def ProfileRandomlyGeneratedKmer(deleted_dna_string, profile, k):
kmers = FindAllKmers(deleted_dna_string, k)
kmers_probabilities = []
initial_kmers_probabilities_sum = 0
nominators_list = []
denominators_list = []
elementary_events_list = []
for kmer in kmers:
kmers_probabilities.append(KmerProbability(profile, kmer))
initial_kmers_probabilities_sum = sum(kmers_probabilities)
for i in range(len(kmers_probabilities)):
kmers_probabilities[i] = kmers_probabilities[i] / initial_kmers_probabilities_sum
nominators_list.append(int(str(Fraction(kmers_probabilities[i]).limit_denominator())[0]))
denominators_list.append(int(str(Fraction(kmers_probabilities[i]).limit_denominator())[str(Fraction(kmers_probabilities[i]).limit_denominator()).find('/') + 1:len(str(Fraction(kmers_probabilities[i]).limit_denominator()))]))
for i in range(len(nominators_list)):
nominators_list[i] = int(nominators_list[i] * (np.lcm.reduce(denominators_list)/denominators_list[i]))
for i in range(len(nominators_list)):
for j in range(nominators_list[i]):
elementary_events_list.append(i)
return kmers[choice(elementary_events_list)]
# + id="jLgHmkIHNhvZ"
def NucleotideIndex(nucleotide):
if nucleotide == 'A':
return 0
elif nucleotide == 'C':
return 1
elif nucleotide == 'G':
return 2
else:
return 3
# + id="xYq5JqVANVs5"
def IndexNucleotide(index):
if index == 0:
return 'A'
elif index == 1:
return 'C'
elif index == 2:
return 'G'
else:
return 'T'
# + id="YlI3_BgmNXW0"
def FindAllKmers(dna_string, k):
kmers_list = []
for kmer_end_index in range(k - 1, len(dna_string)):
kmers_list.append(dna_string[kmer_end_index + 1 - k:kmer_end_index + 1])
return kmers_list
# + id="CY58FfytNYyI"
def Score(motifs, k):
score = 0
profile = GenerateProfile(motifs, k)
consensus = Consensus(profile, k)
for motif in motifs:
score = score + HammingDistance(consensus, motif)
return score
# + id="fZZTG19VNZY4"
def Consensus(profile, k):
consensus = []
for i in range(k):
consensus.append(IndexNucleotide(np.argmax(profile[:,i])))
return ''.join(consensus)
# + id="Qu_JBSBjNazw"
def HammingDistance(string1, string2):
counter = 0
if len(string1) > len(string2):
for i in range(len(string2)):
if string1[i] != string2[i]:
counter = counter + 1
counter = counter + (len(string1) - len(string2))
else:
for i in range(len(string1)):
if string1[i] != string2[i]:
counter = counter + 1
counter = counter + (len(string2) - len(string1))
return counter
# + id="CHpoNjfwNcNd"
def DnaToArray(Dna):
dna_array = np.zeros((len(Dna), len(Dna[0])), dtype='str')
for dna_string in enumerate(Dna):
dna_array[dna_string[0],:] = np.asarray(list(dna_string[1]), dtype='str')
return dna_array
# + id="PWEV1OfT-Kjf"
Dna = ['CGCCCCTCTCGGGGGTGTTCAGTAAACGGCCA', 'GGGCGAGGTATGTGTAAGTGCCAAGGTGCCAG', 'TAGTACCGAGACCGAAAGAAGTATACAGGCGT', 'TAGATCAAGTTTCAGGTGCACGTCGGTGAACC', 'AATCCACCAGCTCCACGTGCAATGTTGGCCTA']
# + colab={"base_uri": "https://localhost:8080/"} id="XmJMnFdU8EHk" outputId="6bb5350b-d60c-4f1a-812f-420eac1eaafa"
Dna
# + id="M1H7hMw1-Uk_"
k = 8
# + id="AYsirg5t-Y3m"
t = 5
# + id="wKZHRSA3-ZxV"
N = 100
# + colab={"base_uri": "https://localhost:8080/"} id="Dpd0DeTj-dJy" outputId="d43d5619-4511-40dd-884c-ad7bfe647706"
GibbsSampler(Dna, k, t, N)
# + id="lGHnGExG9gch"
k = 15
# + id="DNuVE1GA9kqM"
t = 20
# + id="dvtAhR439mij"
N = 2000
# + id="bqg_2gGm9odE"
with open('/content/rosalind_ba2g.txt') as task_file:
Dna = [line.rstrip() for line in task_file]
# + colab={"base_uri": "https://localhost:8080/"} id="Q1BBbE6g9w09" outputId="ac0fa379-9e17-4b6b-8a90-b730ee1d2a41"
Dna
# + id="bwmmmL-i9yYF"
GibbsSampler(Dna, k, t, N)
| BA2G.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### Feature Crosses
# ##### In this exercise we learn about feature crosses and create code to use feature crosses and experiment with different ways to represent features
#
# ##### We also learn to use different features of tensor flow and pands like
# * Use tf.feature_column methods to represent features in different ways.
# * Represent features as bins.
# * Cross bins to create a feature cross.
# <b> We make use of the same californa housing dataset for our coding purpose </b>
# %tensorflow_version 2.x
# +
import numpy as np
import pandas as pd
import tensorflow as tf
from tensorflow import feature_column
from tensorflow.keras import layers
from matplotlib import pyplot as plt
# The following lines adjust the granularity of reporting.
pd.options.display.max_rows = 10
pd.options.display.float_format = "{:.1f}".format
tf.keras.backend.set_floatx('float32')
# -
# <b> We do the same loading, scaling and shuffling of data before we use the dataset for training the model for better accuracy
# and also as a means to prepare the data.</b>
# +
# Load the dataset
train_df = pd.read_csv("https://download.mlcc.google.com/mledu-datasets/california_housing_train.csv")
test_df = pd.read_csv("https://download.mlcc.google.com/mledu-datasets/california_housing_test.csv")
# Scale the labels
scale_factor = 1000.0
# Scale the training set's label.
train_df["median_house_value"] /= scale_factor
# Scale the test set's label
test_df["median_house_value"] /= scale_factor
# Shuffle the examples
train_df = train_df.reindex(np.random.permutation(train_df.index))
# -
# <b> We will try to represent the Latitude and Longitude values as floating point values
# To create feature colums we call</b>
# * tf.feature_column to represent a single feature, single feature cross or a single synthetic feature. To represent a feature as a numeric floating point number we can use tf.feature_column.numeric_column, to represent as a bucket or bins use tf.feature_column.bucketized_column
# * Add the columns into a python list
#
# +
# Create a python list
feature_columns = []
# numerical feature to represent latiutde
latitude = tf.feature_column.numeric_column("latitude")
feature_columns.append(latitude)
# numerical feature represent longitude.
longitude = tf.feature_column.numeric_column("longitude")
# add the features into the python list
feature_columns.append(longitude)
# convert the features into a layer which will be part of the model.
fp_feature_layer = layers.DenseFeatures(feature_columns)
# -
# #### Lets define the functions used to create the model
#
# * create_model - defines the Tensorflow to build a linear regression model and use the fp_feature_layer to represent the model's feature
# * train_model - trains the model which the specified features.
# * plot_the_loss_curve - generates a loss curve
# +
#define the functions
def create_model(my_learning_rate, feature_layer):
#Create and compile a simple linear regression model.
model = tf.keras.models.Sequential()
# Add the layer containing the feature columns to the model.
model.add(feature_layer)
# Add one linear layer to the model to yield a simple linear regressor.
model.add(tf.keras.layers.Dense(units=1, input_shape=(1,)))
# Construct the layers into a model that TensorFlow can execute.
model.compile(optimizer=tf.keras.optimizers.RMSprop(lr=my_learning_rate),
loss="mean_squared_error",
metrics=[tf.keras.metrics.RootMeanSquaredError()])
return model
def train_model(model, dataset, epochs, batch_size, label_name):
# we feed the dataset into the model here so it can train it.
features = {name:np.array(value) for name, value in dataset.items()}
label = np.array(features.pop(label_name))
history = model.fit(x=features, y=label, batch_size=batch_size,
epochs=epochs, shuffle=True)
# store the list of epochs
epochs = history.epoch
# Isolate the mean absolute error for each epoch.
hist = pd.DataFrame(history.history)
rmse = hist["root_mean_squared_error"]
return epochs, rmse
def plot_the_loss_curve(epochs, rmse):
# Plot the loss curve against the epoch.
plt.figure()
plt.xlabel("Epoch")
plt.ylabel("Root Mean Squared Error")
plt.plot(epochs, rmse, label="Loss")
plt.legend()
plt.ylim([rmse.min()*0.94, rmse.max()* 1.05])
plt.show()
# -
# ##### We now train the model with the floating point representations that was created earlier.
# +
# define the hyperparameters.
learning_rate = 0.05
epochs = 30
batch_size = 100
label_name = 'median_house_value'
# compile the model's topography.
my_model = create_model(learning_rate, fp_feature_layer)
# Train the model.
epochs, rmse = train_model(my_model, train_df, epochs, batch_size, label_name)
plot_the_loss_curve(epochs, rmse)
test_features = {name:np.array(value) for name, value in test_df.items()}
test_label = np.array(test_features.pop(label_name))
my_model.evaluate(x=test_features, y=test_label, batch_size=batch_size)
# -
# ##### Lets represent the Latitude and Longitude values as buckets.
#
# <b> We can create the latitude and longitude values as buckets or bins. Each bin represents all the neighbourhoods within a single degree. for example neighbourhoods within 34.4 to 34.8 are in a single bucket but neighbourhoods in 34.4 to 35.2 are in different buckets.
# The model will learn a new weight for each bucket.
# We create 10 buckets each for Latitude and Longitude </b>
# +
resolution_in_degrees = 1.0
# Create a python list to hold the features
feature_columns = []
# Create a bucket feature for latitude.
latitude_as_a_numeric_column = tf.feature_column.numeric_column("latitude")
latitude_boundaries = list(np.arange(int(min(train_df['latitude'])),
int(max(train_df['latitude'])),
resolution_in_degrees))
latitude = tf.feature_column.bucketized_column(latitude_as_a_numeric_column,
latitude_boundaries)
feature_columns.append(latitude)
# Create a bucket feature column for longitude.
longitude_as_a_numeric_column = tf.feature_column.numeric_column("longitude")
longitude_boundaries = list(np.arange(int(min(train_df['longitude'])),
int(max(train_df['longitude'])),
resolution_in_degrees))
longitude = tf.feature_column.bucketized_column(longitude_as_a_numeric_column,
longitude_boundaries)
feature_columns.append(longitude)
# Convert the list of feature columns into a layer which is part of the model.
buckets_feature_layer = layers.DenseFeatures(feature_columns)
# +
# Train the model with the bucketized representations
# define the hyperparameters.
learning_rate = 0.04
epochs = 35
# Build the model passing in the buckets_feature_layer.
my_model = create_model(learning_rate, buckets_feature_layer)
# Train the model.
epochs, rmse = train_model(my_model, train_df, epochs, batch_size, label_name)
plot_the_loss_curve(epochs, rmse)
my_model.evaluate(x=test_features, y=test_label, batch_size=batch_size)
# -
# #### After you run the model we notice the bucket representation does better than the floating point representation of features.
# ##### But we can do better by using feature crosses, ie, by using a feature cross of latitude and longitude and creating a single feature cross, as in real life scenarios the location exists in two dimensions of latitude and longitude it makes sense to use the latitude and longitude as a feature cross
# +
# following code demonstrates the feature cross.
resolution_in_degrees = 1.0
# python list to hold generated feature column.
feature_columns = []
# bucket feature column for latitude.
latitude_as_a_numeric_column = tf.feature_column.numeric_column("latitude")
latitude_boundaries = list(np.arange(int(min(train_df['latitude'])), int(max(train_df['latitude'])), resolution_in_degrees))
latitude = tf.feature_column.bucketized_column(latitude_as_a_numeric_column, latitude_boundaries)
# feature column for longitude.
longitude_as_a_numeric_column = tf.feature_column.numeric_column("longitude")
longitude_boundaries = list(np.arange(int(min(train_df['longitude'])), int(max(train_df['longitude'])), resolution_in_degrees))
longitude = tf.feature_column.bucketized_column(longitude_as_a_numeric_column, longitude_boundaries)
# feature cross of latitude and longitude.
latitude_x_longitude = tf.feature_column.crossed_column([latitude, longitude], hash_bucket_size=125)
crossed_feature = tf.feature_column.indicator_column(latitude_x_longitude)
feature_columns.append(crossed_feature)
# Convert the list of feature columns into a layer to feed to the model.
feature_cross_feature_layer = layers.DenseFeatures(feature_columns)
# +
# lets run the model with the feature crossed columns.
# define the hyperparameters.
learning_rate = 0.04
epochs = 35
# Build the model passing in the feature_cross_feature_layer:
my_model = create_model(learning_rate, feature_cross_feature_layer)
# Train the model on the training set.
epochs, rmse = train_model(my_model, train_df, epochs, batch_size, label_name)
plot_the_loss_curve(epochs, rmse)
my_model.evaluate(x=test_features, y=test_label, batch_size=batch_size)
# -
| featurecrosses.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
library(tidyverse)
source("plotting_common.r")
# + jupyter={"outputs_hidden": true}
d = bind_rows(lapply(Sys.glob("../city2ba/*_*_*.csv"),
function(x){ read_csv(x) %>%
mutate(bal=x, type=ifelse(grepl("forward",x), "forward", "both"))
}))
d_agg = d %>% group_by(solver, cameras, type) %>% summarize(solve_time=sum(solve_time), setup_time=sum(setup_time))
# -
ggplot(d_agg, aes(x=cameras, y=solve_time, color=type, linetype=solver)) + geom_line()
| notebooks/connectivity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp GasConsumptionForecasting
# -
# # # Gas Consumption Forecasting
#
# > Extended learning predictive model for gas consumption pattern in order to decide the strategies of North Khorasan Gas Company
# > University of Bojnurd and NKH Gas Company
#hide
from nbdev.showdoc import *
| 00_core.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import csv
import re
positive_count, negative_count=0,0
positive_corpus,negative_corpus=[],[]
total=0
with open('IMDB Dataset.csv','r') as csv_file:
csv_reader=csv.reader(csv_file)
for line in csv_reader:
total+=1
if total==1394: #something wrong with the other line
break
else:
if line[1]=='positive':
positive_count+=1
positive_corpus.append(line[0])
elif line[1]=='negative':
negative_count+=1
negative_corpus.append(line[0])
print(positive_count,negative_count)
def totalWords(corpus):
words=0
for i in corpus:
words+=len(i.split(" "))
return words
totalWords(positive_corpus),totalWords(negative_corpus)
totalWordspositive=totalWords(positive_corpus)
totalWordsnegative=totalWords(negative_corpus)
total_corpus_count=positive_count+negative_count
probabilities=dict()
probabilities["positive"]=positive_count/total_corpus_count
probabilities["negative"]=negative_count/total_corpus_count
probabilities
# +
ref_words=[]
with open('positive.csv') as pos:
pos_reader=csv.reader(pos)
for line in pos_reader:
ref_words.append(line[0])
with open('negative.csv') as neg:
neg_reader=csv.reader(neg)
for line in neg_reader:
ref_words.append(line[0])
# -
np.array(ref_words)
# +
def parameter_estimation(probabilities,ref_words,totalWordspositive,totalWordsnegative):
#probabilities refers to a dictionary of given probabilities
#ref words (in the name itself)
#Compute P(Word | pos) and P(Word | neg)
for word in ref_words:
count=0
for texts in positive_corpus:
count+=len(re.findall(word,texts))
probabilities[word+" | positive"]=(count+1)/totalWordspositive
if count==0:
probabilities[word+" | positive"]=1/totalWordspositive
count=0
for texts in negative_corpus:
count+=len(re.findall(word,texts))
probabilities[word+" | negative"]=(count+1)/totalWordsnegative
if count==0:
probabilities[word+" | negative"]=1/totalWordsnegative
return probabilities
prob_dict=parameter_estimation(probabilities,ref_words,totalWordspositive,totalWordsnegative)
# -
for prob in prob_dict:
if prob_dict[prob]>1:
print(prob_dict[prob])
print(prob)
if prob_dict[prob]==0.0:
print(prob_dict[prob])
print(prob)
prob_dict
def NaiveBayesMultinomial(prob_dict,new_text,ref_words):
new_text=new_text.split(" ")
positive_score,negative_score=prob_dict["positive"],prob_dict["negative"]
for everyword in new_text:
if everyword in ref_words:
positive_score*=prob_dict[everyword+" | positive"]
negative_score*=prob_dict[everyword+" | negative"]
if negative_score > positive_score:
print("It is negative")
elif positive_score>negative_score:
print("It is positive")
else:
print("It is a moderate comment")
print(positive_score,negative_score)
NaiveBayesMultinomial(prob_dict,"I love this movie to death. OMG I cannot wait to watch it again soon !",ref_words)
NaiveBayesMultinomial(prob_dict,"This movie is the tragically the not-so good and thus the worst shit I have ever had to feast my eyes on",ref_words)
def Gaussian(mean,variance,x):
return (1/np.sqrt(2*np.pi*variance))*np.exp(-0.5*((x-mean)/np.sqrt(variance))**2)
male,female=[],[]
with open('human_data.csv') as pos:
pos_reader=csv.reader(pos)
count=-1
for line in pos_reader:
count+=1
if line[0].lower()=="male":
male.append([line[1],line[2]])
elif line[0].lower()=="female":
female.append([line[1],line[2]])
male=np.array(male,dtype="float")
female=np.array(female,dtype="float")
means=np.array([np.mean(male[:,0]),np.mean(male[:,1]),np.mean(female[:,0]),np.mean(female[:,1])])
stdev=np.array([np.std(male[:,0]),np.std(male[:,1]),np.std(female[:,0]),np.std(female[:,1])])
print(means)
print(stdev)
parameters=np.array([means,stdev])
parameters.shape
def NaiveBayesGaussian(parameters,new_data):
likelihood_vector=Gaussian(means,stdev**2,new_data)
logs=np.log(likelihood_vector)
male=np.sum(logs[:2],keepdims=True)
female=np.sum(logs[2:],keepdims=True)
print("Male Score: ", male)
print("Female Score:", female)
if male>female:
print("Subject with {} kg weight and {} cm height is most likely a male".format(new_data[0],new_data[1]))
return 1
else:
print("Subject with {} kg weight and {} cm height is most likely a female".format(new_data[0],new_data[1]))
return 0
NaiveBayesGaussian(parameters,np.array([88,152,88,152]))
# +
test_data=np.zeros((337,3))
with open('test_data_human_dim.csv') as tester:
reader=csv.reader(tester)
count=0
for line in reader:
if float(line[2])<20.0:
continue
else:
test_data[count,0:3]=line[1],line[0],line[3]
count+=1
print(count)
test_data
# -
y_bar=test_data[:,2].reshape(337,1)
y_bar
def create_array(test_data):
new_array=np.zeros((337,4))
count=0
for row in test_data:
new_array[count,:]=row[0],row[1],row[0],row[1]
count+=1
return new_array
formatted_data=create_array(test_data)
formatted_data
y_predicted=np.zeros((337,1))
row_number=0
for row in formatted_data:
y_predicted[row_number,:][0]=NaiveBayesGaussian(parameters,row)
if y_predicted[row_number,:]==1:
print("male found")
row_number+=1
#print(y_predicted)
NaiveBayesGaussian(parameters,np.array([62.99,163.83,62.99,163.83]))
difference_vector=y_bar-y_predicted
difference_vector.shape
count=0
for i in range(0,337):
if difference_vector[i,:][0]!=0:
count+=1
print(count)
percentage_error=count*100/337
percentage_error
| Naive Bayes Classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.2 ('FAIKR_venv')
# language: python
# name: python3
# ---
import pandas as pd
from matplotlib import pyplot as plt
import seaborn as sns
from scipy.stats import chi2_contingency
import numpy as np
from IPython.display import display
df = pd.read_csv('dataset/survey_results_public.csv', sep=',')
display(df.info())
display(df.describe())
# +
columns=['Hobby','OpenSource','Country','Employment',\
'FormalEducation','UndergradMajor','JobSatisfaction','ConvertedSalary']
data=df.get(columns).dropna()
data=data.rename(columns={'ConvertedSalary':'Salary'})
data.head()
# +
js_dict = {
'Extremely dissatisfied':0,
'Moderately dissatisfied':1,
'Slightly dissatisfied':2,
'Neither satisfied nor dissatisfied':3,
'Slightly satisfied':4,
'Moderately satisfied':5,
'Extremely satisfied':6,
}
data['JobSatisfaction']=data['JobSatisfaction'].map(js_dict)
# -
data = data.replace('Bachelor’s degree (BA, BS, B.Eng., etc.)',"Bachelor")
data = data.replace('Master’s degree (MA, MS, M.Eng., MBA, etc.)',"Master")
data = data.replace('Other doctoral degree (Ph.D, Ed.D., etc.)',"Ph.D")
data = data[data['FormalEducation'].isin(['Bachelor','Master','Ph.D'])]
countries=data.groupby('Country').size().sort_values()[-3:].index.tolist()
data = data[data['Country'].isin(countries)]
plt.figure(figsize=(10,8))
data.Salary=data.Salary.map(lambda x: '0-250.000' if x<=250000 else '250.000-1.000.000' if x<=1000000 else '1.000.000+')
data.Salary.hist()
# +
stem=['A natural science (ex. biology, chemistry, physics)',
'Computer science, computer engineering, or software engineering',
'Web development or web design',
'Another engineering discipline (ex. civil, electrical, mechanical)',
'Mathematics or statistics',
'Information systems, information technology, or system administration',
]
not_stem=[ 'A social science (ex. anthropology, psychology, political science)',
'A humanities discipline (ex. literature, history, philosophy)',
'A business discipline (ex. accounting, finance, marketing)',
'Fine arts or performing arts (ex. graphic design, music, studio art)',
'A health science (ex. nursing, pharmacy, radiology)',
]
data=data[data['UndergradMajor'].isin(stem+not_stem)]
data.UndergradMajor=data.UndergradMajor.map(lambda x: 'STEM' if x in stem else 'NOT_STEM')
#'I never declared a major' WHERE DO THEY GO?
# -
display(data.head())
for col in data.columns:
print(col,data[col].unique())
# +
def chi2_contingency_mat(data,alpha=None):
s=data.columns.size
a = 0
b = 0
mat=np.zeros((s,s))
for i in data.columns:
for j in data.columns:
contigency_pct = pd.crosstab(data[i], data[j])
c, p, dof, expected = chi2_contingency(contigency_pct)
mat[a][b]=p
b=b+1
a = a+1
b=0
if alpha:
mat[mat>=alpha]=1
mat[mat<alpha]=0
return mat
# -
chi2Mat=chi2_contingency_mat(data,0.000005)
plt.figure(figsize=(10,8))
sns.heatmap(chi2Mat,annot=True,xticklabels=data.columns, yticklabels=data.columns)
def getEdges(mat,names=[]):
result=[]
l,_=mat.shape
for i in range(l):
for j in range(i+1,l):
if mat[i,j]==0:
if len(names)>0:
result.append((names[i],names[j]))
else: result.append((i,j))
return result
edges=getEdges(chi2Mat,names=data.columns)
edges
# +
from pgmpy.models import BayesianNetwork
from pgmpy.estimators import ParameterEstimator,MaximumLikelihoodEstimator
model= BayesianNetwork(edges)
pe = ParameterEstimator(model, data)
model.fit(data, estimator=MaximumLikelihoodEstimator)
for cpd in model.get_cpds():
print(cpd)
model.get_independencies()
# -
model.check_model()
| demo_V2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Extrinsic Calibration from 2D-2D Correspondences
#
# ### Goal
#
# Given a set of 2D-2D correspondences between two camera images of known intrinsic parameters $K$, this notebook estimates their relative pose $R, t$.
#
# * Input
# * $x_1$: 2D points in Cam1 image
# * $x_2$: 2D points in Cam2 image
# * $K_1$: Cam1 intrinsic parameter
# * $K_2$: Cam2 intrinsic parameter
# * Output
# * $R, t$: Relative pose satisfying $X_2 = R X_1 + t$ (i.e., Cam1 is the world coordinate system)
#
#
# Note:
# 1. If the two cameras share a single intrinsic parameter (e.g., structure-from-motion, or physically-identical cameras),
# use `cv2.findEssentialMat()` and `cv2.recoverPose()`.
#
# ## Libraries
# +
# %matplotlib notebook
import sys, os, cv2
import numpy as np
from glob import glob
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
module_path = os.path.abspath(os.path.join('..'))
if module_path not in sys.path:
sys.path.append(module_path)
from pycalib.plot import plotCamera
from pycalib.calib import lookat
# -
# ## Synthetic data
#
#
# +
# 3D points
# X_gt = (np.random.rand(16, 3) - 0.5)*5 + [0, 0, 10] # random points centered at [0, 0, 10]
X_gt = np.array(np.meshgrid(np.linspace(-2, 2, 5), np.linspace(-2, 2, 5), np.linspace(9, 11, 3))).reshape((3, -1)).T # 3D grid points
# Camera poses
R1_gt = np.eye(3)
t1_gt = np.zeros(3).reshape((3,1))
R2_gt, t2_gt = lookat(np.array([1,0,0]), np.array([0,0,10]), np.array([0,1,0]))
rvec1_gt = cv2.Rodrigues(R1_gt)[0]
rvec2_gt = cv2.Rodrigues(R2_gt)[0]
# Camera intrinsics
K1 = np.array([[600, 0, 320], [0, 600, 240], [0, 0, 1]]).astype(np.float) # VGA camera
K2 = np.array([[800, 0, 640], [0, 800, 360], [0, 0, 1]]).astype(np.float) # 720p camera
# 2D corresponding points
x1 = cv2.projectPoints(X_gt.reshape((-1, 1, 3)), rvec1_gt, t1_gt, K1, None)[0].reshape((-1, 2))
x2 = cv2.projectPoints(X_gt.reshape((-1, 1, 3)), rvec2_gt, t2_gt, K2, None)[0].reshape((-1, 2))
# Verify triangulation
Y = cv2.triangulatePoints(K1 @ np.hstack((R1_gt, t1_gt)), K2 @ np.hstack((R2_gt, t2_gt)), x1.T, x2.T)
Y = Y[:3] / Y[3,:]
assert np.allclose(0, X_gt - Y.T)
# Verify z > 0 at each camera
assert np.all(X_gt[:, 2] > 0)
assert np.all((R2_gt @ X_gt.T + t2_gt)[2, :] > 0)
# -
# ## Essential matrix
#
# * Input: $K_1$, $K_2$, $x_1$, $x_2$
# * Output: $E$ satisfying $ (K_2^{-1}\tilde{x}_2)^\top E (K_1^{-1} \tilde{x}_1) = 0$
# +
# x1, x2 -> F
F, _ = cv2.findFundamentalMat(x1, x2, cv2.FM_8POINT)
x1h = cv2.convertPointsToHomogeneous(x1).reshape((-1,3)).T
x2h = cv2.convertPointsToHomogeneous(x2).reshape((-1,3)).T
n1 = np.linalg.inv(K1) @ x1h
n2 = np.linalg.inv(K2) @ x2h
# check F error
F_err = x2h.T @ F @ x1h
F_err = np.mean(F_err**2)
print(f'F error = {F_err}')
# F, K1, K2 -> E
# x2.T F x1 = 0 <-> n2.T E n1 = 0 where n1 = K1^-1 x1
E = K2.T @ F @ K1
E = E / np.linalg.norm(E)
# check E error
E_err = np.mean((n2.T @ E @ n1)**2)
print(f'E error = {E_err}') # should be identical to F_err
# -
# ## Decompose $R, t$ from $E$
#
# Given an essentinal matrix $E$, we have four possible solutions
# * $R, t$,
# * $R, -t$,
# * $R', t$, and
# * $R', -t$.
#
# We can find the collect one by checking if the triangulated 3D points appear in front of the both cameras.
#
#
# +
R2a, R2b, t2 = cv2.decomposeEssentialMat(E)
R1 = np.eye(3)
t1 = np.zeros((3,1))
def z_count(R1, t1, R2, t2, K1, K2, x1, x2):
"""
Count number of points appeared in front of the cameras
"""
P1 = K1 @ np.hstack((R1, t1))
P2 = K2 @ np.hstack((R2, t2))
Xh1 = cv2.triangulatePoints(P1, P2, x1, x2)
Xh1 /= Xh1[3,:]
z1 = np.sum(Xh1[2,:]>0) # num of positive z points in Cam1 coordinate system
Xh2 = R2 @ Xh1[:3,:] + t2
z2 = np.sum(Xh2[2,:]>0) # num of positive z points in Cam2 coordinate system
return (z1 + z2), Xh1[:3,:]
zmax = -1
for R2x, t2x in [[R2a, t2], [R2a, -t2], [R2b, t2], [R2b, -t2]]:
z, Xx = z_count(R1, t1, R2x, t2x, K1, K2, x1.T, x2.T)
if zmax < z:
zmax = z
R2_est = R2x
t2_est = t2x
X_est = Xx
print('R2 = ', R2_est)
print('t2 = ', t2_est)
print('R2_gt = ', R2_gt) # ground truth
print('t2_gt = ', t2_gt) # ground truth
# -
# ## Plot
# +
fig = plt.figure()
ax = Axes3D(fig)
ax.set_xlim(-10, 10)
ax.set_ylim(-10, 10)
ax.set_zlim(-1, 10)
plotCamera(ax, R1, t1, "b", 1)
plotCamera(ax, R2_est.T, - R2_est.T @ t2_est, "g", 1)
ax.plot(X_est[0,:], X_est[1,:], X_est[2,:], ".")
fig.show()
# -
# ## Exercises
#
# 1. Add Gaussian noise (e.g. $\mu=0, \sigma=1 \mathrm{px}$) to the 2D observations `x1` and `x2`, and see how the results are degraded.
# 2. Add outliers to `x1` and `x2`.
# 3. Introduce distortion correction.
# * Assume distorted observations and distortion coefficients are given. Rectify the observations as a preprocessing.
| ipynb/excalib_2d.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Exercício Aula 3: Erros (Parte 1)
import math
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# %matplotlib inline
# ## Serão usadas duas séries matemáticas para calcular e^-5 e após, e^-15.
# ## Abaixo estão calculados os erros para x = 5 e para x = 15, com 20 termos para cada cálculo.
# - Quantos termos são necessários para que a primeira série tenha o mesmo erro real que a segunda série tem com 20 termos?
# - Se x = 15, qual o erro se obtém para cada série com o uso de 20 termos?
# - Quanto é o maior erro de x = 15 em cada série?
# - Quantos termos se deve usar para x = 15 ter o mesmo erro que para x = 5?
# O valor real que queremos calcular corresponde a:
# ## Valor real:
e_real = math.e**(-5)
print(e_real)
# # Primeiro caso: x = 5.
# ## Segunda Série
# <img src="segunda.png">
euler = []
x = 5
e = 0
for n in range(20):
e += ((x**n)/(math.factorial(n)))
euler.append(1/e)
print(euler)
# # Valor final: 0.00673794
# # Erro relativo aproximado:
erros = []
for i in range(1,20):
erros.append(math.fabs(100*((euler[i] - euler[i-1])/euler[i])))
print(erros)
# # Valor final, erro relativo aproximado: 0.000105648 %
# # Erro relativo real:
erros2 = []
for i in range(20):
erros2.append(math.fabs(100*((e_real - euler[i])/e_real)))
print(erros2)
# # Valor final, erro relativo real: 3.45213 * 10^(-5) %
tabela = pd.DataFrame(erros2)
tabela.index = [i for i in range(20)]
tabela.columns = ['Erro Real da Segunda Série']
tabela
# ## Primeira Série
# <img src="primeira.png">
euler2 = []
x = 5
e2 = 0
for n in range(20):
e2 += ((-1)**n)*((x**n)/(math.factorial(n)))
euler2.append(e2)
print(euler2)
# # Valor final: 0.00670634
# ## Erro relativo aproximado:
erros3 = []
for i in range(1,20):
erros3.append(math.fabs(100*((euler2[i] - euler2[i-1])/euler2[i])))
print(erros3)
# # Valor final, erro relativo aproximado: 2.33802 %
# ## Erro relativo real:
erros4 = []
for i in range(20):
erros4.append(math.fabs(100*((e_real - euler2[i])/e_real)))
print(erros4)
# # Valor final, erro relativo real: 0.469073 %
# Tabela:
tabela2 = pd.DataFrame(erros4)
tabela2.index = [i for i in range(20)]
tabela2.columns = ['Erro Real, Primeira Série']
tabela2
# # Segundo caso: x = -15.
# ## Valor real:
e_real = math.e**(-15)
print(e_real)
# ## Segunda série
# <img src="segunda.png">
euler = []
x = 15
e = 0
for n in range(20):
e += ((x**n)/(math.factorial(n)))
euler.append(1/e)
print(euler)
# # Valor final: 3.49515 * 10^(-7)
# # Erro relativo aproximado:
erros = []
for i in range(1,20):
erros.append(math.fabs(100*((euler[i] - euler[i-1])/euler[i])))
print(erros)
# # Valor final, erro relativo aproximado: 6.80280 %
# # Erro relativo real:
erros2 = []
for i in range(20):
erros2.append(math.fabs(100*((e_real - euler[i])/e_real)))
print(erros2)
# # Valor final, erro relativo real: 14.2571 %
tabela = pd.DataFrame(erros2)
tabela.index = [i for i in range(20)]
tabela.columns = ['Erro Real da Segunda Série']
tabela
# ## Primeira série
# <img src="primeira.png">
euler2 = []
x = 15
e2 = 0
for n in range(20):
e2 += ((-1)**n)*((x**n)/(math.factorial(n)))
euler2.append(e2)
print(euler2)
# # Valor final: -79065.7
# # Erro relativo aproximado:
euler2 = []
x = 15
e2 = 0
for n in range(20):
e2 += ((-1)**n)*((x**n)/(math.factorial(n)))
euler2.append(e2)
print(euler2)
# # Valor final, erro relativo aproximado: -79065.7 %
# # Erro relativo real:
erros3 = []
for i in range(1,20):
erros3.append(math.fabs(100*((euler2[i] - euler2[i-1])/euler2[i])))
print(erros3)
# # Valor final, erro relativo real: 230.489 %
erros4 = []
for i in range(20):
erros4.append(math.fabs(100*((e_real - euler2[i])/e_real)))
print(erros4)
tabela2 = pd.DataFrame(erros4)
tabela2.index = [i for i in range(20)]
tabela2.columns = ['Erro Real, Primeira Série']
tabela2
# ## Itens
# O maior erro é o erro relativo aproximado: -79065.7 %, para a primeira série e o erro relativo real: 14.2571 %, para a segunda série.
# ## Comentário
#
# Ambas as séries funcionam muito melhor quando x = 5, ou seja, apresentam os erros menores, de ambos os tipos, para x = 5, o que faz sentido, visto que como os cálculos envolvem potências, os erros de arredondamento são menores quanto menor for o número o qual se está multiplicando por ele mesmo repetidas vezes.
# Na solução de problemas de engenharia, é necessário sempre escolher o método mais apropriado para minimizar os erros.
# Observando a tabela, é possível concluir que a primeira série precisa de 27 iterações para que seu erro seja menor que o erro
# do vigésimo termo da segunda série.
# Assim, a resposta da segunda pergunta é que o número de termos é 27 (pois zero é um termo), no vigésimo sexto termo o erro se torna
# menor que 0,000035.
# Conforme observado na leitura do capítulo 3, já era esperado que o erro da primeira série fosse maior, pois a subtração
# de números parecidos acarreta mais erros que a soma de números parecidos.
# Sendo assim, a segunda série é mais adequada do ponto de vista dos erros, o que pode ser atestado pelo fato de ela ter apresentado erros menores em ambos os casos estudados no exercício (x = 5 e x = 15).
| exercicio_aula_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.4.3
# language: julia
# name: julia-0.4
# ---
# +
using SymPy
xi = symbols("x_i", real=true)
yi = symbols("y_i", real=true)
xip1 = symbols("x_{i+1}", real=true)
yip1 = symbols("y_{i+1}", real=true)
xim1 = symbols("x_{i-1}", real=true)
yim1 = symbols("y_{i-1}", real=true)
# Definition of surface area gradient (computed by hand) at node i
lp = sqrt((xi-xip1)^2+(yi-yip1)^2) # length of next edge
lm = sqrt((xi-xim1)^2+(yi-yim1)^2) # length of previous edge
fix = (xip1-xi)/lp - (xi-xim1)/lm # force x component
fiy = (yip1-yi)/lp - (yi-yim1)/lm # force y component
# Compute Hessian Stencil
Dixx = simplify(diff(fix,xi))
Dixy = simplify(diff(fiy,xi))
Diyx = simplify(diff(fix,yi)) # expect to be the same as Dixy
Diyy = simplify(diff(fiy,yi))
Dip1xx = simplify(diff(fix,xip1))
Dip1xy = simplify(diff(fiy,xip1))
Dip1yx = simplify(diff(fix,yip1)) # expect to be the same as Dip1xy
Dip1yy = simplify(diff(fiy,yip1))
assert(Diyx == Dixy)
assert(Dip1xy == Dip1yx)
# +
using LaTeXStrings
# prepare results for display
D(i,x,y) = "\\frac{\\partial f_{$(i)}^$(x)}{\\partial $(y)_{$(i)}}"
res = [
"\$ $(D("i","x","x")) = $(SymPy.latex(Dixx))\$",
"\$ $(D("i","x","y")) = $(D("i","y","x")) = $(SymPy.latex(Dixy))\$",
"\$ $(D("i","y","y")) = $(SymPy.latex(Diyy))\$",
"\$ $(D("i+1","x","x")) = $(SymPy.latex(Dip1xx))\$",
"\$ $(D("i+1","x","y")) = $(D("i+1","y","x")) = $(SymPy.latex(Dip1xy))\$",
"\$ $(D("i+1","y","y")) = $(SymPy.latex(Dip1yy))\$",
]
# display results
for str in res
display(LaTeXString(str))
end
# -
# Express the above derivatives in terms of edge vector components
eix = symbols("{e^x_i}", real=true)
eiy = symbols("{e^y_i}", real=true)
ein = symbols("{\\|{e_{i}}\\|}", positive=true)
eim1x = symbols("{e^x_{i-1}}", real=true)
eim1y = symbols("{e^y_{i-1}}", real=true)
eim1n = symbols("{\\|{e_{i-1}}\\|}", positive=true)
function sim(f)
f = subs(subs(subs(f,xip1-xi,eix),yip1-yi,eiy),eix^2+eiy^2,ein^2)
f = subs(subs(subs(f,xi-xim1,eim1x),yi-yim1,eim1y),eim1x^2+eim1y^2,eim1n^2)
simplify(f)
end
res = [
"\$ $(D("i","x","x")) = $(SymPy.latex(sim(Dixx)))\$",
"\$ $(D("i","x","y")) = $(D("i","y","x")) = $(SymPy.latex(sim(Dixy)))\$",
"\$ $(D("i","y","y")) = $(SymPy.latex(sim(Diyy)))\$",
"\$ $(D("i+1","x","x")) = $(SymPy.latex(sim(Dip1xx)))\$",
"\$ $(D("i+1","x","y")) = $(D("i+1","y","x")) = $(SymPy.latex(sim(Dip1xy)))\$",
"\$ $(D("i+1","y","y")) = $(SymPy.latex(sim(Dip1yy)))\$",
]
for str in res
display(LaTeXString(str))
end
| hessian-of-surface-area.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dependencies
#
# This demo uses three main libs (aside from jupyter notebooks):
#
# - Czml3: a lib made by <NAME>, to generate Cesium data from Python in CZML format ([more info](https://github.com/poliastro/czml3))
# - Telluric: a lib from Satellogic, to work with geospatial data ([docs](https://telluric.readthedocs.io/en/latest/))
# - Orbit-predictor: a lib from Satellogic, to work with orbital data ([more info](https://github.com/satellogic/orbit-predictor))
# !pip install czml3 telluric orbit-predictor
# +
from datetime import timedelta
from orbit_predictor.locations import Location
from orbit_predictor.sources import get_predictor_from_tle_lines
from telluric import GeoVector, GeoFeature, FeatureCollection
from czml3.widget import CZMLWidget
# this will be ported to CZML3 in the future, but for now, it's part of this demo repo :)
from cesium_helpers import czml_from_feature_collection
# -
# # Prepare the data for the visualization
#
# We will display three things:
#
# - The area of a country (Cyprus)
# - A point marking its capital (Nicosia)
# - And the path of a satellite passing over the capital (Maryam, one of Satellogic's satellites)
# ### Cyprus geodata
# +
# nicosia coordinates
nicosia_lat = 35.166667
nicosia_lon = 33.366667
cyprus = GeoVector.from_geojson("../satellogic/model_based_planner/mbp/data/countries/Cyprus.geojson")
nicosia = GeoVector.point(nicosia_lon, nicosia_lat)
# -
# ### Satellite path data
# +
# first we need a predictor, to be able to calculate the satellite path
# this is a predictor for Maryam, one of the satellites from Satellogic's constellation
# (the number lines here are a TLE, a format that describes orbits)
predictor = get_predictor_from_tle_lines([
"1 43204U 18015K 19358.42624529 .00000968 00000-0 42378-4 0 9999",
"2 43204 97.4185 123.8187 0017139 4.8687 80.8213 15.24342402104681",
])
# then we use the predictor to find a pass over Nicosia
pass_over_nicosia = predictor.get_next_pass(Location("Nicosia", nicosia_lat, nicosia_lon, 0),
max_elevation_gt=80)
# and we calculate the satellite path during the pass over Nicosia
# start at the "acquisition of signal", the moment the satellite starts seeing Nicosia
# and end at the "loss of signal", when the satellite stops seeing Nicosia
current_date = pass_over_nicosia.aos
max_date = pass_over_nicosia.los
# get the position (as lon, lat) of the satellite every 10 seconds
path_positions = []
while current_date <= max_date:
position = predictor.get_position(current_date).position_llh
path_positions.append((position[1], position[0]))
current_date += timedelta(seconds=10)
# build a line with the points
satellite_path = GeoVector.line(path_positions)
# -
# # Visualize the data
features_on_map = FeatureCollection([
# show Cyprus in slightly transparent purple
GeoFeature(cyprus, {"color": [150, 0, 150, 100]}),
# show Nicosia in green
GeoFeature(nicosia, {"color": [0, 250, 0]}),
# show the satellite path in yellow
GeoFeature(satellite_path, {"color": [250, 250, 0]}),
])
# ### (rotate and zoom on the globe to see Cyprus)
czml = czml_from_feature_collection(features_on_map, "Visualization")
CZMLWidget(czml)
| cesium-demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Gensim tests
import logging
from gensim import corpora, models, similarities
logging.basicConfig(format='%(asctime)s : %(levelname)s : %(message)s', level=logging.INFO)
# ## Preliminaries
corpus = [[(0, 1.0), (1, 1.0), (2, 1.0)],
[(2, 1.0), (3, 1.0), (4, 1.0), (5, 1.0), (6, 1.0), (8, 1.0)],
[(1, 1.0), (3, 1.0), (4, 1.0), (7, 1.0)],
[(0, 1.0), (4, 2.0), (7, 1.0)],
[(3, 1.0), (5, 1.0), (6, 1.0)],
[(9, 1.0)],
[(9, 1.0), (10, 1.0)],
[(9, 1.0), (10, 1.0), (11, 1.0)],
[(8, 1.0), (10, 1.0), (11, 1.0)]]
tfidf = models.TfidfModel(corpus)
vec = [(0, 1), (4, 1)]
print(tfidf[vec])
index = similarities.SparseMatrixSimilarity(tfidf[corpus], num_features=12)
sims = index[tfidf[vec]]
print(list(enumerate(sims)))
# ## Tutorial 1 : Corpora and Vector Spaces
documents = ["Human machine interface for lab abc computer applications",
"A survey of user opinion of computer system response time",
"The EPS user interface management system",
"System and human system engineering testing of EPS",
"Relation of user perceived response time to error measurement",
"The generation of random binary unordered trees",
"The intersection graph of paths in trees",
"Graph minors IV Widths of trees and well quasi ordering",
"Graph minors A survey"]
# +
from collections import defaultdict
from pprint import pprint # pretty-printer
# remove common words and tokenize
stoplist = set('for a of the and to in'.split())
texts = [[word for word in document.lower().split() if word not in stoplist]
for document in documents]
# remove words that appear only once
frequency = defaultdict(int)
for text in texts:
for token in text:
frequency[token] += 1
texts = [[token for token in text if frequency[token] > 1]
for text in texts]
pprint(texts)
# -
dictionary = corpora.Dictionary(texts)
dictionary.save('../tmp/deerwester.dict') # store the dictionary, for future reference
print(dictionary)
print(dictionary.token2id)
new_doc = "Human computer interaction"
new_vec = dictionary.doc2bow(new_doc.lower().split())
print(new_vec) # the word "interaction" does not appear in the dictionary and is ignored
corpus = [dictionary.doc2bow(text) for text in texts]
corpora.MmCorpus.serialize('../tmp/deerwester.mm', corpus) # store to disk, for later use
pprint(corpus)
class MyCorpus(object):
def __iter__(self):
for line in open('../data/mycorpus.txt'):
# assume there's one document per line, tokens separated by whitespace
yield dictionary.doc2bow(line.lower().split())
corpus_memory_friendly = MyCorpus() # doesn't load the corpus into memory!
print(corpus_memory_friendly)
for vector in corpus_memory_friendly: # load one vector into memory at a time
print(vector)
# +
from six import iteritems
# collect statistics about all tokens
dictionary = corpora.Dictionary(line.lower().split() for line in open('../data/mycorpus.txt'))
# remove stop words and words that appear only once
stop_ids = [dictionary.token2id[stopword] for stopword in stoplist
if stopword in dictionary.token2id]
once_ids = [tokenid for tokenid, docfreq in iteritems(dictionary.dfs) if docfreq == 1]
dictionary.filter_tokens(stop_ids + once_ids) # remove stop words and words that appear only once
dictionary.compactify() # remove gaps in id sequence after words that were removed
print(dictionary)
# +
# create a toy corpus of 2 documents, as a plain Python list
corpus = [[(1, 0.5)], []] # make one document empty, for the heck of it
corpora.MmCorpus.serialize('../tmp/corpus.mm', corpus)
# -
corpus = corpora.MmCorpus('../tmp/corpus.mm')
# +
print(corpus)
# one way of printing a corpus: load it entirely into memory
print(list(corpus))
# another way of doing it: print one document at a time, making use of the streaming interface
#for doc in corpus:
# print(doc)
# -
# ## Tutorial 2 : Topics and Transformations
# +
import os
from gensim import corpora, models, similarities
if (os.path.exists("../tmp/deerwester.dict")):
dictionary = corpora.Dictionary.load('../tmp/deerwester.dict')
corpus = corpora.MmCorpus('../tmp/deerwester.mm')
print("Used files generated from first tutorial")
else:
print("Please run first tutorial to generate data set")
# -
tfidf = models.TfidfModel(corpus) # step 1 -- initialize a model
doc_bow = [(0, 1), (1, 1)]
print(tfidf[doc_bow]) # step 2 -- use the model to transform vectors
corpus_tfidf = tfidf[corpus]
for doc in corpus_tfidf:
print(doc)
lsi = models.LsiModel(corpus_tfidf, id2word=dictionary, num_topics=2) # initialize an LSI transformation
corpus_lsi = lsi[corpus_tfidf] # create a double wrapper over the original corpus: bow->tfidf->fold-in-lsi
for doc in corpus_lsi: # both bow->tfidf and tfidf->lsi transformations are actually executed here, on the fly
print(doc)
lsi.save('../tmp/model.lsi') # same for tfidf, lda, ...
lsi = models.LsiModel.load('../tmp/model.lsi')
# ## Tutorial 3 : Similarity Queries
from gensim import corpora, models, similarities
dictionary = corpora.Dictionary.load('../tmp/deerwester.dict')
corpus = corpora.MmCorpus('../tmp/deerwester.mm') # comes from the first tutorial, "From strings to vectors"
print(corpus)
lsi = models.LsiModel(corpus, id2word=dictionary, num_topics=2)
doc = "Human computer interaction"
vec_bow = dictionary.doc2bow(doc.lower().split())
vec_lsi = lsi[vec_bow] # convert the query to LSI space
print(vec_lsi)
index = similarities.MatrixSimilarity(lsi[corpus]) # transform corpus to LSI space and index it
index.save('../tmp/deerwester.index')
index = similarities.MatrixSimilarity.load('../tmp/deerwester.index')
sims = index[vec_lsi] # perform a similarity query against the corpus
print(list(enumerate(sims))) # print (document_number, document_similarity) 2-tuples
sims = sorted(enumerate(sims), key=lambda item: -item[1])
print(sims) # print sorted (document number, similarity score) 2-tuples
| src/test_gensim.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/gcastelao/TFG-Economia_GonzaloCastelao/blob/main/TFG.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="Ll-Phpc__rGv"
from tkinter import *
from tkinter import messagebox
from tkinter import ttk
from tkinter.ttk import Label
import numpy_financial as npf
#---------------------------------- Funcionalidades ------------------------------------
def salirAplicacion():
valor=messagebox.askquestion("Salir", "¿Deseas salir de la aplicación?")
if valor=="yes":
root.destroy()
def infoLicencia():
info=messagebox.showinfo("Licencia", "Esta aplicación ha sido diseñada por"
" <NAME> dentro de la realización del TFG de Economía.")
def infoAcerca():
infoAyuda=messagebox.showinfo("Ayuda", "Este programa se ha desarrollado para"
" demostrar la importancia de la reinversión de los flujos intermedios."
" Es decir, la influencia del riesgo de reinversión." + "\n" + "En el caso"
" español, los Bonos del Estado a 10 años, que son los que se están analizando"
" en el programa, implican en todos los casos pago de cupones intermedios por"
" lo que este riesgo aparece siempre." + "\n" + "Lo que se pretende con este"
" programa es ver cuáles son los efectos de dicho riesgo tomando diferentes"
" precios, cupones y tasas de reinversión.")
def LimpiaResultados():
textoResultados['text'] = ""
#------------------------------------------ Estilo --------------------------------------
root=Tk()
root.title("TFG Economía")
barraMenu=Menu(root)
root.config(menu=barraMenu, width=300, height=300)
salirMenu=Menu(barraMenu, tearoff=0)
salirMenu.add_command(label="Salir", command= salirAplicacion)
ayudaMenu=Menu(barraMenu, tearoff=0)
ayudaMenu.add_command(label="Licencia", command=infoLicencia)
ayudaMenu.add_command(label="Acerca de...", command= infoAcerca)
barraMenu.add_cascade(label="Salir", menu=salirMenu)
barraMenu.add_cascade(label="Ayuda", menu=ayudaMenu)
#---------------------------------- Comienzo de campos ------------------------------------
miFrame=Frame(root)
miFrame.pack()
miId=StringVar() #con esto puedo recuperar info de un entry
miPrecio=StringVar()
miCupón=StringVar()
cuadroPrecio=ttk.Combobox(miFrame, values=["950", "1000", "1500"], state="readonly")
cuadroPrecio.grid(row=0, column=1, padx=10, pady=10)
cuadroPrecio.current(0)
cuadroCupon=ttk.Combobox(miFrame, values=["0.5%", "1%", "2%"], state="readonly")
cuadroCupon.grid(row=1, column=1, padx=10, pady=10)
cuadroCupon.current(0)
#---------------------------------- Comienzo de label ------------------------------------
PrecioLabel=Label(miFrame, text="Elija el precio: ")
PrecioLabel.grid(row=0, column=0, sticky="e", padx=10, pady=10)
CupónLabel=Label(miFrame, text="Elija el cupón: ")
CupónLabel.grid(row=1, column=0, sticky="e", padx=10, pady=10)
comentariosLabel=Label(miFrame, text="Resultados: ")
comentariosLabel.grid(row=3, column=0, sticky="e", padx=10, pady=10)
#-------------------------------- RESULTADOS -----------------------------------------------
def montante1(cf1,r1,n1):
m1 = 0 # montante
for i in range(1, n1+1):
m1 += cf1[i]*(1+r1)**(n1-i)
return m1
def CalculaResultados():
precio1 = int(cuadroPrecio.get())
cupon = cuadroCupon.get()
nominal1 = 1000
n1 = 10 # años
resultados = ""
# Extracción de los valores del cupón en pantalla
if cupon == "0.5%":
cupon1 = float("0.005")
elif cupon == "1%":
cupon1 = float("0.01")
elif cupon == "2%":
cupon1 = float("0.02")
# Flujos de caja del bono
cf1 = [0]*(n1+1)
for i in range(1,n1+1):
cf1[i] = cupon1 * nominal1
cf1[n1] += nominal1
cf1[0] = -precio1
resultados = resultados + f"Cuando el precio es {precio1} y el cupón es del {cupon1:.2%}, la TIR del bono es {npf.irr(cf1):.2%}" + "\n"
tasas_reinversion = [-0.02,-0.01,0,0.01,0.02]
for t in tasas_reinversion:
m1 = montante1(cf1,t,n1)
resultados = resultados + "\n"
resultados = resultados + f"El montante reinvirtiendo al tanto del {t:.0%} es {m1:,.2f} €" + "\n"
resultados = resultados + f"La rentabilidad del inversor reinvirtiendo al tanto {t:.0%} es {(m1/precio1)**(1/n1)-1:.2%}" + "\n"
tasa_supuesta1 = [npf.irr(cf1)]
for t in tasa_supuesta1:
m1 = montante1(cf1,t,n1)
resultados= resultados + "\n"
resultados = resultados + f"Para obtener la rentabilidad supuesta, el inversor debe conseguir reinvertir al {t:.2%} obteniendo un montante de {m1:,.2f} €" + "\n"
resultados = resultados + f"En este caso, la rentabilidad del inversor sería efectivamente de {(m1/precio1)**(1/n1)-1:.2%}"
textoResultados['text'] = resultados
textoResultados = Label(miFrame, text="")
textoResultados.grid(row=3, column=1, padx=10, pady=10)
#---------------------------------- Comienzo de botones ------------------------------------
botonCalcular=Button(miFrame, text="Calcular", command=CalculaResultados)
botonCalcular.grid(row=2, column=0, sticky="e", padx=10, pady=10)
botonLimpiar=Button(miFrame, text="Limpiar", command=LimpiaResultados)
botonLimpiar.grid(row=2, column=1, sticky="e", padx=10, pady=10)
root.mainloop()
| TFG.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %autosave 10
# **First, create a dataframe of *cleaned_data.csv*. Then, use scikit-learn's `train_test_split` to make a new set of training and testing data. This time, instead of `EDUCATION`, use `LIMIT_BAL`: the account's credit limit.**
# + eid="662ef"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn import metrics
# %matplotlib inline
from sklearn.linear_model import LogisticRegression
import matplotlib as mpl
mpl.rcParams['figure.dpi'] = 400
df = pd.read_csv('cleaned_data.csv')
df.head()
# -
df['default payment next month'].mean()
df.groupby('default payment next month')['ID'].count()
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(df['LIMIT_BAL'].values.reshape(-1,1), df['default payment next month'].values,test_size=0.2, random_state=24)
# _________________________________________________________________________
# **Next, train a logistic regression model using the training data from your split.**
#
example_lr = LogisticRegression(C=0.1, class_weight=None, dual=False, fit_intercept=True,intercept_scaling=1, max_iter=100, multi_class='auto',n_jobs=None, penalty='l2', random_state=None, solver='liblinear',tol=0.0001, verbose=0, warm_start=False)
example_lr.fit(X_train, y_train)
# _________________________________________________________________________
# **Then, create the array of predicted probabilities for the testing data.**
#
y_pred_proba = example_lr.predict_proba(X_test)
# _________________________________________________________________________
# **Next, calculate the ROC AUC using the predicted probabilities and the true labels of the testing data. Compare this to the ROC AUC from using the `EDUCATION` feature.**
#
metrics.roc_auc_score(y_test, y_pred_proba[:,1])
fpr, tpr, thresholds = metrics.roc_curve(y_test, y_pred_proba[:,1])
plt.plot(fpr, tpr, '*-')
plt.plot([0, 1], [0, 1], 'r--')
plt.legend(['Logistic regression', 'Random chance'])
plt.xlabel('FPR')
plt.ylabel('TPR')
plt.title('ROC curve for logistic regression with LIMIT_BAL feature')
# _________________________________________________________________________
# **Then, calculate the data for the precision-recall curve on the testing data using scikit-learn functionality.**
#
precision, recall, thresh = metrics.precision_recall_curve(y_test, y_pred_proba[:,1])
# _________________________________________________________________________
# **Plot the precision-recall curve using matplotlib.**
#
plt.plot(recall, precision, '-x')
plt.xlabel('Recall')
plt.ylabel('Precision')
plt.title('Precision and recall for the logistic regression with LIMIT_BAL')
plt.xlim([0, 1])
plt.ylim([0, 1])
# _________________________________________________________________________
# **Now, use scikit-learn to calculate the area under the precision-recall curve.**
#
metrics.auc(recall, precision)
y_train_pred_proba = example_lr.predict_proba(X_train)
# _________________________________________________________________________
# **Finally, recalculate the ROC AUC, except this time do it for the training data.**
# + eid="d61c3"
metrics.roc_auc_score(y_train, y_train_pred_proba[:,1])
# -
# How is this different, conceptually and quantitatively, from your earlier calculation?
#
| Mini-Project-2/Project 3/.ipynb_checkpoints/Performing_Logistic_Regression-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#import os module and csv file
import os
import csv
budget_data_csv_path = ("budget_data.csv")
months = []
net_total = []
monthly_change = []
with open (budget_data_csv_path) as csvfile:
csvreader = csv.reader(csvfile, delimiter=",")
first_row = next(csvreader)
# append to the lists
for row in csvreader:
months.append(row[0])
net_total.append(int(row[1]))
month = len(months)
total = sum(net_total)
#create for loop to get changes in profits/losses
for i in range(len(net_total)-1):
monthly_change.append(net_total[i+1]-net_total[i])
average_change = sum(monthly_change)/len(monthly_change)
# find the greatest increase/decrease in profits/losses and the dates to go along with above
greatest_increase = max(monthly_change)
greatest_decrease = min(monthly_change)
# Print Financial Analysis results
print("Financial Analysis")
print("----------------------------")
print("Total Months: " + (str(month)))
print("Total: " + "$" + str(total))
print("Average Change: " + "$" + str(round(average_change, 2)))
print("Greatest Increase in Profits: " + str((months[monthly_change.index(greatest_increase)+1])) + " (" + "$" + str((greatest_increase)) + ")")
print("Greatest Decrease in Profits: " + str((months[monthly_change.index(greatest_decrease)+1])) + " (" + "$" + str((greatest_decrease)) + ")")
data_output = os.path.join("budget_data.csv")
output = open("data_output.txt", "w")
with open(data_output.txt, "w") as file:
writer = csv.writer(csvfile)
writer.writerow("Financial Analysis")
# -
| PyBank/Resources/.ipynb_checkpoints/main-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: papers
# language: python
# name: papers
# ---
# # Dev: Papers IDs collector
#
# - arXiv API
# - cleantext
#
# ### References
#
# - [arXiv.org](https://arxiv.org/)
# - [arXiv API-Homepage](https://pypi.org/project/arxiv/)
# - [arXiv API-Documentation](http://lukasschwab.me/arxiv.py/index.html)
# - [arXiv API-Documentation-Custom queries](https://arxiv.org/help/api/user-manual#query_details)
import pandas as pd
import numpy as np
# ## parsing using beautifulsoup
import requests
from bs4 import BeautifulSoup
from datetime import datetime, date
url = 'https://arxiv.org/search/advanced?advanced=&terms-0-operator=AND&terms-0-term=&terms-0-field=title&classification-computer_science=y&classification-physics_archives=all&classification-include_cross_list=include&date-filter_by=all_dates&date-year=&date-from_date=&date-to_date=&date-date_type=submitted_date&abstracts=hide&size=200&order=-announced_date_first&start=0'
reqs = requests.get(url)
soup = BeautifulSoup(reqs.text, 'lxml')
print("\nFind and print all li tags:\n")
for tag in soup.find_all("li", {"class": "arxiv-result"})[:2]:
for tag_a in tag.find_all("a")[:1]:
print("{0}: {1}".format(tag_a.name, tag_a.text.replace('arXiv', '').rstrip().lstrip()))
for tag_d in tag.find_all("div", {"class": "tags is-inline-block"})[:1]:
print("{0}: {1}".format(tag_d.name, [it.rstrip().lstrip() for it in tag_d.text.split('\n') if it != '']))
tag_p = tag.find_all("p")[3]
sdate = tag_p.text.split(';')[0].replace('Submitted', '').rstrip().lstrip()
dt = datetime.strptime(sdate, '%d %B, %Y')
print("{0}: {1}".format(tag_p.name, date(dt.year, dt.month, dt.day)))
print('\n')
# ## arXiv API
import arxiv
search = arxiv.Search(id_list=["2109.04744"])
paper = next(search.results())
print(paper.title)
print([author.name for author in paper.authors])
print(paper.categories)
print(paper.primary_category)
print(paper.summary)
print(paper.pdf_url)
print(paper.published)
print(paper.updated)
import arxiv
search = arxiv.Search(query="cat:stat.ML")
paper = next(search.results())
print(paper.title)
print([author.name for author in paper.authors])
print(paper.categories)
print(paper.primary_category)
print(paper.summary)
print(paper.pdf_url)
print(paper.published)
print(paper.updated)
print(paper.get_short_id())
import arxiv
search = arxiv.Search(query="automl")
paper = next(search.results())
print(paper.title)
print([author.name for author in paper.authors])
print(paper.categories)
print(paper.primary_category)
print(paper.summary)
print(paper.pdf_url)
print(paper.published)
print(paper.updated)
print(paper.get_short_id())
import arxiv
search = arxiv.Search(query="cat:cs.LG", max_results=10, sort_by=arxiv.SortCriterion.SubmittedDate)
for paper in search.results():
print(paper.get_short_id(), paper.published)
# ## cleantext tool
import cleantext
txt = paper.summary
txt_cleaned = cleantext.clean_words(txt,
all= False, # Execute all cleaning operations
extra_spaces=True , # Remove extra white space
stemming=False , # Stem the words
stopwords=True ,# Remove stop words
lowercase=True ,# Convert to lowercase
numbers=True ,# Remove all digits
punct=True ,# Remove all punctuations
stp_lang='english' # Language for stop words
)
txt_cleaned[:5]
cleantext.clean(txt,
all= False, # Execute all cleaning operations
extra_spaces=True , # Remove extra white space
stemming=False , # Stem the words
stopwords=True ,# Remove stop words
lowercase=True ,# Convert to lowercase
numbers=True ,# Remove all digits
punct=True ,# Remove all punctuations
stp_lang='english' # Language for stop words
)
| notebooks/0-dev-review_tools.ipynb |
# ---
# jupyter:
# jupytext:
# cell_metadata_filter: -all
# formats: ipynb
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction #
#
# Once you've identified a set of features with some potential, it's time to start developing them. In this lesson, you'll learn a number of common transformations you can do entirely in Pandas. If you're feeling rusty, we've got a great [course on Pandas](https://www.kaggle.com/learn/pandas).
#
# We'll use four datasets in this lesson having a range of feature types: [*US Traffic Accidents*](https://www.kaggle.com/sobhanmoosavi/us-accidents), [*1985 Automobiles*](https://www.kaggle.com/toramky/automobile-dataset), [*Concrete Formulations*](https://www.kaggle.com/sinamhd9/concrete-comprehensive-strength), and [*Customer Lifetime Value*](https://www.kaggle.com/pankajjsh06/ibm-watson-marketing-customer-value-data). The following hidden cell loads them up.
# +
#$HIDE_INPUT$
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
plt.style.use("seaborn-whitegrid")
plt.rc("figure", autolayout=True)
plt.rc(
"axes",
labelweight="bold",
labelsize="large",
titleweight="bold",
titlesize=14,
titlepad=10,
)
accidents = pd.read_csv("../input/fe-course-data/accidents.csv")
autos = pd.read_csv("../input/fe-course-data/autos.csv")
concrete = pd.read_csv("../input/fe-course-data/concrete.csv")
customer = pd.read_csv("../input/fe-course-data/customer.csv")
# -
# <blockquote style="margin-right:auto; margin-left:auto; background-color: #ebf9ff; padding: 1em; margin:24px;">
# <strong>Tips on Discovering New Features</strong>
# <ul>
# <li>Understand the features. Refer to your dataset's <em>data documentation</em>, if available.
# <li>Research the problem domain to acquire <strong>domain knowledge</strong>. If your problem is predicting house prices, do some research on real-estate for instance. Wikipedia can be a good starting point, but books and <a href="https://scholar.google.com/">journal articles</a> will often have the best information.
# <li>Study previous work. <a href="https://www.kaggle.com/sudalairajkumar/winning-solutions-of-kaggle-competitions">Solution write-ups</a> from past Kaggle competitions are a great resource.
# <li>Use data visualization. Visualization can reveal pathologies in the distribution of a feature or complicated relationships that could be simplified. Be sure to visualize your dataset as you work through the feature engineering process.
# <ul>
# </blockquote>
#
# # Mathematical Transforms #
#
# Relationships among numerical features are often expressed through mathematical formulas, which you'll frequently come across as part of your domain research. In Pandas, you can apply arithmetic operations to columns just as if they were ordinary numbers.
#
# In the *Automobile* dataset are features describing a car's engine. Research yields a variety of formulas for creating potentially useful new features. The "stroke ratio", for instance, is a measure of how efficient an engine is versus how performant:
# +
autos["stroke_ratio"] = autos.stroke / autos.bore
autos[["stroke", "bore", "stroke_ratio"]].head()
# -
# The more complicated a combination is, the more difficult it will be for a model to learn, like this formula for an engine's "displacement", a measure of its power:
autos["displacement"] = (
np.pi * ((0.5 * autos.bore) ** 2) * autos.stroke * autos.num_of_cylinders
)
# Data visualization can suggest transformations, often a "reshaping" of a feature through powers or logarithms. The distribution of `WindSpeed` in *US Accidents* is highly skewed, for instance. In this case the logarithm is effective at normalizing it:
# +
# If the feature has 0.0 values, use np.log1p (log(1+x)) instead of np.log
accidents["LogWindSpeed"] = accidents.WindSpeed.apply(np.log1p)
# Plot a comparison
fig, axs = plt.subplots(1, 2, figsize=(8, 4))
sns.kdeplot(accidents.WindSpeed, shade=True, ax=axs[0])
sns.kdeplot(accidents.LogWindSpeed, shade=True, ax=axs[1]);
# -
# Check out our [lesson on normalization](https://www.kaggle.com/alexisbcook/scaling-and-normalization) in [*Data Cleaning*](https://www.kaggle.com/learn/data-cleaning) where you'll also learn about the *Box-Cox transformation*, a very general kind of normalizer.
#
# # Counts #
#
# Features describing the presence or absence of something often come in sets, the set of risk factors for a disease, say. You can aggregate such features by creating a **count**.
#
# These features will be *binary* (`1` for Present, `0` for Absent) or *boolean* (`True` or `False`). In Python, booleans can be added up just as if they were integers.
#
# In *Traffic Accidents* are several features indicating whether some roadway object was near the accident. This will create a count of the total number of roadway features nearby using the `sum` method:
# +
roadway_features = ["Amenity", "Bump", "Crossing", "GiveWay",
"Junction", "NoExit", "Railway", "Roundabout", "Station", "Stop",
"TrafficCalming", "TrafficSignal"]
accidents["RoadwayFeatures"] = accidents[roadway_features].sum(axis=1)
accidents[roadway_features + ["RoadwayFeatures"]].head(10)
# -
# You could also use a dataframe's built-in methods to *create* boolean values. In the *Concrete* dataset are the amounts of components in a concrete formulation. Many formulations lack one or more components (that is, the component has a value of 0). This will count how many components are in a formulation with the dataframe's built-in greater-than `gt` method:
# +
components = [ "Cement", "BlastFurnaceSlag", "FlyAsh", "Water",
"Superplasticizer", "CoarseAggregate", "FineAggregate"]
concrete["Components"] = concrete[components].gt(0).sum(axis=1)
concrete[components + ["Components"]].head(10)
# -
# # Building-Up and Breaking-Down Features #
#
# Often you'll have complex strings that can usefully be broken into simpler pieces. Some common examples:
# - ID numbers: `'123-45-6789'`
# - Phone numbers: `'(999) 555-0123'`
# - Street addresses: `'8241 Kaggle Ln., Goose City, NV'`
# - Internet addresses: `'http://www.kaggle.com`
# - Product codes: `'0 36000 29145 2'`
# - Dates and times: `'Mon Sep 30 07:06:05 2013'`
#
# Features like these will often have some kind of structure that you can make use of. US phone numbers, for instance, have an area code (the `'(999)'` part) that tells you the location of the caller. As always, some research can pay off here.
#
# The `str` accessor lets you apply string methods like `split` directly to columns. The *Customer Lifetime Value* dataset contains features describing customers of an insurance company. From the `Policy` feature, we could separate the `Type` from the `Level` of coverage:
# +
customer[["Type", "Level"]] = ( # Create two new features
customer["Policy"] # from the Policy feature
.str # through the string accessor
.split(" ", expand=True) # by splitting on " "
# and expanding the result into separate columns
)
customer[["Policy", "Type", "Level"]].head(10)
# -
# You could also join simple features into a composed feature if you had reason to believe there was some interaction in the combination:
autos["make_and_style"] = autos["make"] + "_" + autos["body_style"]
autos[["make", "body_style", "make_and_style"]].head()
# <blockquote style="margin-right:auto; margin-left:auto; background-color: #ebf9ff; padding: 1em; margin:24px;">
# <strong>Elsewhere on Kaggle Learn</strong><br>
# There are a few other kinds of data we haven't talked about here that are especially rich in information. Fortunately, we've got you covered!
# <ul>
# <li> For <strong>dates and times</strong>, see <a href="https://www.kaggle.com/alexisbcook/parsing-dates">Parsing Dates</a> from our Data Cleaning course.
# <li> For <strong>latitudes and longitudes</strong>, see our <a href="https://www.kaggle.com/learn/geospatial-analysis">Geospatial Analysis</a> course.
# <li> For <strong>text</strong>, try <a href="https://www.kaggle.com/learn/natural-language-processing">Natural Language Processing</a>.
# </ul>
# </blockquote>
# # Group Transforms #
#
# Finally we have **Group transforms**, which aggregate information across multiple rows grouped by some category. With a group transform you can create features like: "the average income of a person's state of residence," or "the proportion of movies released on a weekday, by genre." If you had discovered a category interaction, a group transform over that categry could be something good to investigate.
#
# Using an aggregation function, a group transform combines two features: a categorical feature that provides the grouping and another feature whose values you wish to aggregate. For an "average income by state", you would choose `State` for the grouping feature, `mean` for the aggregation function, and `Income` for the aggregated feature. To compute this in Pandas, we use the `groupby` and `transform` methods:
# +
customer["AverageIncome"] = (
customer.groupby("State") # for each state
["Income"] # select the income
.transform("mean") # and compute its mean
)
customer[["State", "Income", "AverageIncome"]].head(10)
# -
# The `mean` function is a built-in dataframe method, which means we can pass it as a string to `transform`. Other handy methods include `max`, `min`, `median`, `var`, `std`, and `count`. Here's how you could calculate the frequency with which each state occurs in the dataset:
#
# +
customer["StateFreq"] = (
customer.groupby("State")
["State"]
.transform("count")
/ customer.State.count()
)
customer[["State", "StateFreq"]].head(10)
# -
# You could use a transform like this to create a "frequency encoding" for a categorical feature.
#
# If you're using training and validation splits, to preserve their independence, it's best to create a grouped feature using only the training set and then join it to the validation set. We can use the validation set's `merge` method after creating a unique set of values with `drop_duplicates` on the training set:
# +
# Create splits
df_train = customer.sample(frac=0.5)
df_valid = customer.drop(df_train.index)
# Create the average claim amount by coverage type, on the training set
df_train["AverageClaim"] = df_train.groupby("Coverage")["ClaimAmount"].transform("mean")
# Merge the values into the validation set
df_valid = df_valid.merge(
df_train[["Coverage", "AverageClaim"]].drop_duplicates(),
on="Coverage",
how="left",
)
df_valid[["Coverage", "AverageClaim"]].head(10)
# -
# <blockquote style="margin-right:auto; margin-left:auto; background-color: #ebf9ff; padding: 1em; margin:24px;">
# <strong>Tips on Creating Features</strong><br>
# It's good to keep in mind your model's own strengths and weaknesses when creating features. Here are some guidelines:
# <ul>
# <li> Linear models learn sums and differences naturally, but can't learn anything more complex.
# <li> Ratios seem to be difficult for most models to learn. Ratio combinations often lead to some easy performance gains.
# <li> Linear models and neural nets generally do better with normalized features. Neural nets especially need features scaled to values not too far from 0. Tree-based models (like random forests and XGBoost) can sometimes benefit from normalization, but usually much less so.
# <li> Tree models can learn to approximate almost any combination of features, but when a combination is especially important they can still benefit from having it explicitly created, especially when data is limited.
# <li> Counts are especially helpful for tree models, since these models don't have a natural way of aggregating information across many features at once.
# </ul>
# </blockquote>
# # Your Turn #
#
# [**Combine and transform features**](#$NEXT_NOTEBOOK_URL$) from *Ames* and improve your model's performance.
| notebooks/feature_engineering_new/raw/tut3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
from matplotlib import pyplot
import numpy
# Grids in AMUSE have a similar functionality as particle sets, but grids are inmutable in the number of grid points in any direction. You cannot add a gridpoint like you add a particle in a particle set, instead you define the number of points in each direction on creation of the grid and amuse will create the grid points.
from amuse.lab import *
# Let's start by creating a simple 3 by 4 grid.
grid = Grid(3,4)
print grid
# A grid created in this way does not define any positions for the gridpoints, you can make a more useful grid with the ``create`` function. The ``create`` function needs a list of the number of points in each direction and the total distance covered by the grid in each direction. The created grid will cover the space from 0 to the given total distance in any direction
grid = Grid.create((3,4),(1|units.m, 2|units.m))
print grid
print grid.x
# The returned positions are the centers of the gridpoints, we can make a small plot to show where the grid centers are positioned.
grid = Grid.create((3,4),(1|units.m, 2|units.m))
pyplot.scatter(
grid.x.value_in(units.m),
grid.y.value_in(units.m)
)
pyplot.xlim(0,1)
pyplot.ylim(0,2)
# You can index a grid in two ways, direct indexing on the grid or indexing on an attribute. Direct indexing on a grid can be more efficient as no data is retrieved until you actually requenst an attribute. If you first request an attribute and then do the indexing, all data for the attribute is retrieved and returned first as a vector quantity (or numpy array), next a subselection is made using the indexing routines. Both method should return the same quantities.
print grid.position[0][0]
print grid[0][0].position
# Grids can be moved around if needed
grid.position -= [0.5,1.0] | units.m
pyplot.scatter(
grid.x.value_in(units.m),
grid.y.value_in(units.m)
)
pyplot.xlim(-0.5,0.5)
pyplot.ylim(-1,1)
| doc/interactive_tutorial/08-Grids.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
# Licensed under the MIT License.
# # How to Setup a Schedule for a Published Pipeline
# In this notebook, we will show you how you can run an already published pipeline on a schedule.
# ## Prerequisites and AML Basics
# Make sure you go through the configuration Notebook located at https://github.com/Azure/MachineLearningNotebooks first if you haven't. This sets you up with a working config file that has information on your workspace, subscription id, etc.
#
# ### Initialization Steps
# +
import azureml.core
from azureml.core import Workspace
# Check core SDK version number
print("SDK version:", azureml.core.VERSION)
ws = Workspace.from_config()
print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\n')
# -
# ### Compute Targets
# #### Retrieve an already attached Azure Machine Learning Compute
# +
from azureml.core import Run, Experiment, Datastore
from azureml.widgets import RunDetails
# -
from azureml.core.compute import AmlCompute, ComputeTarget
aml_compute_target = "aml-compute"
try:
aml_compute = AmlCompute(ws, aml_compute_target)
print("Found existing compute target: {}".format(aml_compute_target))
except:
print("Creating new compute target: {}".format(aml_compute_target))
provisioning_config = AmlCompute.provisioning_configuration(vm_size = "STANDARD_D2_V2",
min_nodes = 1,
max_nodes = 4)
aml_compute = ComputeTarget.create(ws, aml_compute_target, provisioning_config)
aml_compute.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)
# ## Build and Publish Pipeline
# Build a simple pipeline, publish it and add a schedule to run it.
# ### Define a pipeline step
# Define a single step pipeline for demonstration purpose.
# +
from azureml.pipeline.steps import PythonScriptStep
# project folder
project_folder = 'scripts'
trainStep = PythonScriptStep(
name="Training_Step",
script_name="train.py",
compute_target=aml_compute_target,
source_directory=project_folder
)
print("TrainStep created")
# -
# ### Build the pipeline
# +
from azureml.pipeline.core import Pipeline
pipeline1 = Pipeline(workspace=ws, steps=[trainStep])
print ("Pipeline is built")
pipeline1.validate()
# -
# ### Publish the pipeline
# +
from datetime import datetime
timenow = datetime.now().strftime('%m-%d-%Y-%H-%M')
pipeline_name = timenow + "-Pipeline"
print(pipeline_name)
published_pipeline1 = pipeline1.publish(
name=pipeline_name,
description=pipeline_name)
print("Newly published pipeline id: {}".format(published_pipeline1.id))
# -
# ## Schedule Operations
# Schedule operations require id of a published pipeline. You can get all published pipelines and do Schedule operations on them, or if you already know the id of the published pipeline, you can use it directly as well.
# ### Get published pipeline ID
# +
from azureml.pipeline.core import PublishedPipeline
# You could retrieve all pipelines that are published, or
# just get the published pipeline object that you have the ID for.
# Get all published pipeline objects in the workspace
all_pub_pipelines = PublishedPipeline.get_all(ws)
# We will iterate through the list of published pipelines and
# use the last ID in the list for Schelue operations:
print("Published pipelines found in the workspace:")
for pub_pipeline in all_pub_pipelines:
print(pub_pipeline.id)
pub_pipeline_id = pub_pipeline.id
print("Published pipeline id to be used for Schedule operations: {}".format(pub_pipeline_id))
# -
# ### Create a schedule for the pipeline using a recurrence
# This schedule will run on a specified recurrence interval.
# +
from azureml.pipeline.core.schedule import ScheduleRecurrence, Schedule
recurrence = ScheduleRecurrence(frequency="Day", interval=2, hours=[22], minutes=[30]) # Runs every other day at 10:30pm
schedule = Schedule.create(workspace=ws, name="My_Schedule",
pipeline_id=pub_pipeline_id,
experiment_name='Schedule_Run',
recurrence=recurrence,
wait_for_provisioning=True,
description="Schedule Run")
# You may want to make sure that the schedule is provisioned properly
# before making any further changes to the schedule
print("Created schedule with id: {}".format(schedule.id))
# -
# Note: Set the `wait_for_provisioning` flag to False if you do not want to wait for the call to provision the schedule in the backend.
# ### Get all schedules for a given pipeline
# Once you have the published pipeline ID, then you can get all schedules for that pipeline.
# +
schedules = Schedule.get_all(ws, pipeline_id=pub_pipeline_id)
# We will iterate through the list of schedules and
# use the last ID in the list for further operations:
print("Found these schedules for the pipeline id {}:".format(pub_pipeline_id))
for schedule in schedules:
print(schedule.id)
schedule_id = schedule.id
print("Schedule id to be used for schedule operations: {}".format(schedule_id))
# -
# ### Get all schedules in your workspace
# You can also iterate through all schedules in your workspace if needed.
# Use active_only=False to get all schedules including disabled schedules
schedules = Schedule.get_all(ws, active_only=True)
print("Your workspace has the following schedules set up:")
for schedule in schedules:
print("{} (Published pipeline: {}".format(schedule.id, schedule.pipeline_id))
# ### Get the schedule
fetched_schedule = Schedule.get(ws, schedule_id)
print("Using schedule with id: {}".format(fetched_schedule.id))
# ### Disable the schedule
# Set the wait_for_provisioning flag to False if you do not want to wait
# for the call to provision the schedule in the backend.
fetched_schedule.disable(wait_for_provisioning=True)
fetched_schedule = Schedule.get(ws, schedule_id)
print("Disabled schedule {}. New status is: {}".format(fetched_schedule.id, fetched_schedule.status))
# ### Reenable the schedule
# Set the wait_for_provisioning flag to False if you do not want to wait
# for the call to provision the schedule in the backend.
fetched_schedule.enable(wait_for_provisioning=True)
fetched_schedule = Schedule.get(ws, schedule_id)
print("Enabled schedule {}. New status is: {}".format(fetched_schedule.id, fetched_schedule.status))
# ### Change recurrence of the schedule
# +
# Set the wait_for_provisioning flag to False if you do not want to wait
# for the call to provision the schedule in the backend.
recurrence = ScheduleRecurrence(frequency="Hour", interval=2) # Runs every two hours
fetched_schedule = Schedule.get(ws, schedule_id)
fetched_schedule.update(name="My_Updated_Schedule",
description="Updated_Schedule_Run",
status='Active',
wait_for_provisioning=True,
recurrence=recurrence)
fetched_schedule = Schedule.get(ws, fetched_schedule.id)
print("Updated schedule:", fetched_schedule.id,
"\nNew name:", fetched_schedule.name,
"\nNew frequency:", fetched_schedule.recurrence.frequency,
"\nNew status:", fetched_schedule.status)
# -
# ### Create a schedule for the pipeline using a Datastore
# This schedule will run when additions or modifications are made to Blobs in the Datastore container.
# Note: Only Blob Datastores are supported.
# +
from azureml.core.datastore import Datastore
datastore = Datastore(workspace=ws, name="workspaceblobstore")
schedule = Schedule.create(workspace=ws, name="My_Schedule",
pipeline_id=pub_pipeline_id,
experiment_name='Schedule_Run',
datastore=datastore,
wait_for_provisioning=True,
description="Schedule Run")
# You may want to make sure that the schedule is provisioned properly
# before making any further changes to the schedule
print("Created schedule with id: {}".format(schedule.id))
# -
# Set the wait_for_provisioning flag to False if you do not want to wait
# for the call to provision the schedule in the backend.
schedule.disable(wait_for_provisioning=True)
schedule = Schedule.get(ws, schedule_id)
print("Disabled schedule {}. New status is: {}".format(schedule.id, schedule.status))
| how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-setup-schedule-for-a-published-pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 2.483251, "end_time": "2021-10-18T07:24:03.625418", "exception": false, "start_time": "2021-10-18T07:24:01.142167", "status": "completed"} tags=[]
import pandas as pd
import numpy as np
import lightgbm as lgb
import pickle
import os
import gc
from sklearn.preprocessing import MinMaxScaler
from matplotlib import pyplot as plt
import seaborn as sns
import json
from utilities import (
RANDOM_STATE, TARGET_COL, N_FOLD, FOLD_STRAT_NAME,
)
INPUT_PATH = '../input/tabular-playground-series-oct-2021'
PATH_NOTEBOOK = '../input/preprocess'
PATH_PARAM = '../input/optuna-lightgbm'
# + papermill={"duration": 11.916999, "end_time": "2021-10-18T07:24:15.559778", "exception": false, "start_time": "2021-10-18T07:24:03.642779", "status": "completed"} tags=[]
train = pd.read_pickle(
os.path.join(PATH_NOTEBOOK, 'train_unscaled.pkl')
)
# + papermill={"duration": 0.029446, "end_time": "2021-10-18T07:24:15.604698", "exception": false, "start_time": "2021-10-18T07:24:15.575252", "status": "completed"} tags=[]
with open(os.path.join(PATH_NOTEBOOK, 'feature_dic.pkl'), 'rb') as file:
feature_dic = pickle.load(file)
# + papermill={"duration": 0.180911, "end_time": "2021-10-18T07:24:15.801904", "exception": false, "start_time": "2021-10-18T07:24:15.620993", "status": "completed"} tags=[]
#CONSTANT
FEATURE = feature_dic['feature']
CAT_COL = feature_dic['categorical']
NUMERIC_COL = feature_dic['numerical']
FOLD_LIST = range(train[FOLD_STRAT_NAME].nunique())
gc.collect()
# + [markdown] papermill={"duration": 0.01508, "end_time": "2021-10-18T07:24:15.832244", "exception": false, "start_time": "2021-10-18T07:24:15.817164", "status": "completed"} tags=[]
# # Set parameter
# + papermill={"duration": 0.03646, "end_time": "2021-10-18T07:24:15.883691", "exception": false, "start_time": "2021-10-18T07:24:15.847231", "status": "completed"} tags=[]
params = {
'objective': 'binary',
'boosting_type': 'gbdt',
'metric': 'auc',
'learning_rate': 0.01,
'random_state': RANDOM_STATE,
'verbose': -1,
'n_jobs': -1,
}
#import lgb optimized parameter (file_name is wrong :()
with open(os.path.join(PATH_PARAM, 'final_xgb_param.pkl'), 'rb') as file:
param_lgb_final = pickle.load(file)
params.update(param_lgb_final)
#pretty print
print(json.dumps(params, indent=4))
# + [markdown] papermill={"duration": 0.014812, "end_time": "2021-10-18T07:24:15.913840", "exception": false, "start_time": "2021-10-18T07:24:15.899028", "status": "completed"} tags=[]
# # TRAIN
# + papermill={"duration": 18703.497985, "end_time": "2021-10-18T12:35:59.427297", "exception": false, "start_time": "2021-10-18T07:24:15.929312", "status": "completed"} tags=[]
score = 0
model_list = []
prediction_df = pd.DataFrame(
{
'fold': train[FOLD_STRAT_NAME],
'prediction': np.zeros((train.shape[0]))
}
)
for i, fold_ in enumerate(FOLD_LIST):
mask_train = (train[FOLD_STRAT_NAME] != fold_)
mask_test = (train[FOLD_STRAT_NAME] == fold_)
train_x, train_y = train.loc[mask_train, FEATURE], train.loc[mask_train, TARGET_COL]
test_x, test_y = train.loc[mask_test, FEATURE], train.loc[mask_test, TARGET_COL]
model = lgb.train(
params,
lgb.Dataset(train_x, label=train_y,categorical_feature=CAT_COL), 100000,
valid_sets = lgb.Dataset(test_x, label=test_y,categorical_feature=CAT_COL),
valid_names ='validation', verbose_eval=100, early_stopping_rounds = 100,
)
#oof prediction
prediction_df.loc[mask_test, 'prediction'] = model.predict(test_x)
#evaluate score and save model for importance/prediction
score_fold = model.best_score['validation']['auc']
score += score_fold/N_FOLD
model_list.append(model)
print('\nFold: {}; Auc: {:.5f}\n'.format(fold_, score_fold))
print('-'*50)
print('\n\n\n')
gc.collect()
print('CV-Auc: {:.5f}\n'.format(score))
# + [markdown] papermill={"duration": 0.282265, "end_time": "2021-10-18T12:35:59.988398", "exception": false, "start_time": "2021-10-18T12:35:59.706133", "status": "completed"} tags=[]
# # Feature importance
# + papermill={"duration": 0.3654, "end_time": "2021-10-18T12:36:00.634125", "exception": false, "start_time": "2021-10-18T12:36:00.268725", "status": "completed"} tags=[]
feature_importances = pd.DataFrame()
feature_importances['feature'] = FEATURE
for fold_, model in enumerate(model_list):
feature_importances['fold_{}'.format(fold_ + 1)] = model.feature_importance(importance_type='gain')
# + papermill={"duration": 1.180057, "end_time": "2021-10-18T12:36:02.097075", "exception": false, "start_time": "2021-10-18T12:36:00.917018", "status": "completed"} tags=[]
scaler = MinMaxScaler(feature_range=(0, 100))
average_importance = feature_importances.drop('feature', axis = 1).mean(axis=1)
feature_importances['average'] = scaler.fit_transform(X=pd.DataFrame(average_importance))
feature_importances = feature_importances.sort_values(by='average', ascending=False)
feature_importances[['feature', 'average']].to_csv('feature_importances.csv',index=False)
fig = plt.figure(figsize=(12,8))
sns.barplot(data=feature_importances.head(50), x='average', y='feature');
plt.title(f'50 TOP feature importance over {N_FOLD} average')
# + [markdown] papermill={"duration": 0.281575, "end_time": "2021-10-18T12:36:02.659140", "exception": false, "start_time": "2021-10-18T12:36:02.377565", "status": "completed"} tags=[]
# # TEST Blending
# + papermill={"duration": 0.595424, "end_time": "2021-10-18T12:36:03.540539", "exception": false, "start_time": "2021-10-18T12:36:02.945115", "status": "completed"} tags=[]
del train
gc.collect()
# + papermill={"duration": 7.64711, "end_time": "2021-10-18T12:36:11.477326", "exception": false, "start_time": "2021-10-18T12:36:03.830216", "status": "completed"} tags=[]
test = pd.read_pickle(
os.path.join(PATH_NOTEBOOK, 'test_unscaled.pkl')
)
# + papermill={"duration": 2367.178632, "end_time": "2021-10-18T13:15:38.940798", "exception": false, "start_time": "2021-10-18T12:36:11.762166", "status": "completed"} tags=[]
pred_test = np.zeros(test.shape[0])
for fold_, model in enumerate(model_list):
pred_test += model.predict(test)/N_FOLD
# + [markdown] papermill={"duration": 0.281107, "end_time": "2021-10-18T13:15:39.508723", "exception": false, "start_time": "2021-10-18T13:15:39.227616", "status": "completed"} tags=[]
# # SAVE RESULT
# + papermill={"duration": 2.406977, "end_time": "2021-10-18T13:15:42.196806", "exception": false, "start_time": "2021-10-18T13:15:39.789829", "status": "completed"} tags=[]
submission = pd.read_csv(os.path.join(INPUT_PATH, 'sample_submission.csv'))
submission['target'] = pred_test
submission.to_csv('submission.csv', index = False)
# + papermill={"duration": 3.795932, "end_time": "2021-10-18T13:15:46.273261", "exception": false, "start_time": "2021-10-18T13:15:42.477329", "status": "completed"} tags=[]
prediction_df.to_csv('oof_prediction', index = False)
| Tabular Playground Series - Oct 2021/tuned_model/lightgbm-tuned.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Плотная упаковка равных сфер (2D-проекции)
import numpy as np
import scipy.linalg as la
import shapely.geometry as geo
import matplotlib.pyplot as plt
import matplotlib.patches as pch
from matplotlib import cm
# #### Гранецентрированная Кубическая (ГЦК)
# По оси симметрии 4-го порядка
# +
fig = plt.figure(figsize=(8,8),dpi=100)
ax = fig.add_subplot(projection='rectilinear')
ax.set_aspect(1)
ax.set_xbound(0,9)
ax.set_ybound(0,9)
ax.grid()
r = 0.5
for iz in np.arange(0,8):
for iy in np.arange(0,8-iz):
for ix in np.arange(0,8-iz):
x = 2*r*ix + r*iz + 1
y = 2*r*iy + r*iz + 1
c = cm.Paired.colors[iz]
p = pch.Circle((x,y), r, alpha=0.95, color=c, ec='gray', lw=0.5)
ax.add_patch(p)
plt.draw()
# -
# #### Гранецентрированная Кубическая (ГЦК)
# По оси симметрии 3-го порядка
# +
fig = plt.figure(figsize=(8,8), dpi=100)
ax = fig.add_subplot(projection='rectilinear')
ax.set_aspect(1)
ax.set_xbound(0,9)
ax.set_ybound(0,9)
ax.grid()
r = 0.5
for iz in np.arange(0,8):
for iy in np.arange(0,8-iz):
for ix in np.arange(0,8-iz-iy):
x = 2*r*ix + r*iy + r*iz + 1
y = np.sqrt(3)*r*iy + (np.sqrt(3)/3)*r*iz + 1
c = cm.Paired.colors[iz]
p = pch.Circle((x,y), r, alpha=0.95, color=c, ec='gray', lw=0.5)
ax.add_patch(p)
plt.draw()
# -
# #### Гексагональная Плотноупакованная (ГПУ)
# +
fig = plt.figure(figsize=(8,8), dpi=100)
ax = fig.add_subplot(projection='rectilinear')
ax.set_aspect(1)
ax.set_xbound(0,10)
ax.set_ybound(0,9)
ax.set_xticks(np.arange(0,11,1))
ax.set_yticks(np.arange(0,10,1))
ax.grid()
r = 0.5
def mk_x_coord(ix, iy, iz):
return 2*r*ix + r*((iy+iz)%2) + 1
def mk_y_coord(ix, iy, iz):
return np.sqrt(3)*r*iy + np.sqrt(3)/3*r*(iz%2) + 1
def create_regular_polygon(xy,v,r=1,o=0):
angles = np.arange(0, 2*np.pi, 2*np.pi/v)
p = [[xy[0]+r*np.cos(a+o),xy[1]+r*np.sin(a+o)] for a in angles]
return geo.Polygon(p)
centroid = np.array([mk_x_coord(4,4,0),mk_y_coord(4,4,0)])
def is_visible(x,y,iz):
l = 2*r*((9-iz)//2) + 1e-10
border = create_regular_polygon(centroid, 6, l)
return border.covers(geo.Point(x,y))
for iz in np.arange(0,9):
for iy in np.arange(0,9):
for ix in np.arange(0,9):
x = mk_x_coord(ix,iy,iz)
y = mk_y_coord(ix,iy,iz)
c = cm.Paired.colors[iz]
if is_visible(x,y,iz):
p = pch.Circle((x,y), r, alpha=0.95, color=c, ec='gray', lw=0.5)
ax.add_patch(p)
plt.draw()
| close-sphere-packing-2d.ipynb |