code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chapter 7: Measuring Uncertainty With the Bootstrap
# +
##### Data and libraries
# Common libraries
import pandas as pd
import numpy as np
import statsmodels.formula.api as smf
from statsmodels.formula.api import ols
import seaborn as sns
# Chapter-specific libraries
import statsmodels.stats.outliers_influence as st_inf # For Cook's distance
import matplotlib.pyplot as plt
import statsmodels.api as sm # For QQ plot
### Generating the data
times = [2,2,3,5,6,9,10,47,61,413]
experience = [11,17,18,1,10,4,6,3,8,0]
data_df = pd.DataFrame(
{'times': times,
'experience': experience})
# -
# ## Intro to the Bootstrap: Polling Oneself Up
# +
# Building linear model
lin_mod = ols("times~1", data=data_df).fit()
print(lin_mod.summary())
est = lin_mod.params['Intercept']
se = lin_mod.bse['Intercept']
#Building normal confidence interval
LL = est-1.96*se #Lower limit
UL = est+1.96*se #Upper limit
print("LL = ", LL)
print("UL = ",UL)
# +
#Building bootstrap CI
mean_lst = []
B = 2000
N = len(data_df)
for i in range(B):
boot_df = data_df.sample(N, replace = True)
M = np.mean(boot_df.times)
mean_lst.append(M)
LL_b = np.quantile(mean_lst, 0.025)
UL_b = np.quantile(mean_lst, 0.975)
print("LL_b = ", LL_b)
print("UL_b = ",UL_b)
# -
#### Bootstrap for time promise
promise_lst = []
B = 2000
N = len(data_df)
for i in range(B):
boot_df = data_df.sample(N, replace = True)
above180 = len(boot_df[boot_df.times >= 180]) / N
promise_lst.append(above180)
LL_b = np.quantile(promise_lst, 0.025)
UL_b = np.quantile(promise_lst, 0.975)
print("LL_b = ", LL_b)
print("UL_b = ",UL_b)
# ## The Bootstrap for Regression Analysis
print(ols("times~experience", data=data_df).fit().summary())
# +
reg_lst = []
B = 4000
N = len(data_df)
for i in range(B):
boot_df = data_df.sample(N, replace = True)
lin_mod = ols("times~experience", data=boot_df).fit()
coeff = lin_mod.params['experience']
reg_lst.append(coeff)
LL_b = np.quantile(reg_lst, 0.025)
UL_b = np.quantile(reg_lst, 0.975)
sns.displot(reg_lst)
# -
pval = 2 * sum(1 for x in reg_lst if x > 0) / B
print(pval)
# ## When to use the Bootstrap
lin_mod = ols("times~experience", data=data_df).fit()
print(lin_mod.summary())
#Extract Cook's distance for influential points
CD = st_inf.OLSInfluence(lin_mod).summary_frame()['cooks_d']
CD[CD > 1]
#Density plot of residuals
res_df = lin_mod.resid
sns.kdeplot(res_df)
#QQ plot of residuals
fig = sm.qqplot(res_df, line='s')
plt.show()
# ## Optimizing the Bootstrap in R and Python
# +
#Creating unique numpy array for sampling
data_ar = data_df.to_numpy()
rng = np.random.default_rng()
np_lst = []
for i in range(B):
#Extracting the relevant columns from array
boot_ar = rng.choice(data_ar, size=N, replace=True)
X = boot_ar[:,1]
X = np.c_[X, np.ones(N)]
Y = boot_ar[:,0]
### LSTQ implementation
np_lst.append(np.linalg.lstsq(X, Y, rcond=-1)[0][0])
#Plotting histogram
sns.displot(np_lst)
LL_b_np = np.quantile(np_lst, 0.025)
UL_b_np = np.quantile(np_lst, 0.975)
| Chapter 7 - Measuring uncertainty with the Bootstrap/ch07.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: test
# language: python
# name: test
# ---
# +
# # !pip install jiwer -qq
# +
# # !pip install git+https://github.com/redapesolutions/suara-kami-community
# -
# %load_ext autoreload
# %autoreload 2
from sk import predict
from pathlib import Path
import jiwer
import pandas as pd
import numpy as np
# # Using predict function
# Predict function is a generic function that will handle all type of input.
#
# It is a high layer api for transcription, if want to do more flexible transcription can use SK class
# # Predict single or multiple file
fn = "/content/test/youtube/0228444ff34081eda587c0ca53712486.wav" # or ["/content/audio1.wav","/content/audio2.wav"]
out = predict(fn)
Path(fn).with_suffix(".txt").read_text()
out
# # Predict from single or multiple folder
fn = "/content/test/" # or ["/content/test","/content/data"]
out = predict(fn)
out.keys()
preds = out["texts"]
files = out["filenames"]
data = []
for i,j in zip(preds,files):
label = j.with_suffix('.txt').read_text()
data.append([j,label,i,jiwer.compute_measures(label,i)['wer'],jiwer.compute_measures([l for l in label],[ii for ii in i])['wer']])
df = pd.DataFrame(data)
df.columns = ["path","label","pred","wer","cer"]
df.head()
df["wer"].mean(),df["cer"].mean()
# # Using SK class
# Why use SK class? if want to transcribe multiple time predict will load the model and lm multiple time which is not efficient, with SK object you can call it multiple time without reloading anything again.
from sk import SK
asr = SK(model="conformer_small",decoder=None)
asr.transcribe_file(fn="/content/test/youtube/0228444ff34081eda587c0ca53712486.wav")
import librosa
asr.transcribe_array(array=librosa.load("/content/test/youtube/0228444ff34081eda587c0ca53712486.wav",16000)[0])
asr = SK(model="silero_en",decoder=None)
asr.transcribe_file(fn="/content/testset-imda/0.wav")
asr = SK(model="silero_en",decoder="en")
asr.transcribe_file(fn="/content/testset-imda/0.wav")
| tutorials/1.speech_to_text.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# # IRGAN: Generative Adversarial Nets for Information Retrival
# + [markdown] deletable=true editable=true
# This notebook provides the implimentations of [IRGAN](https://arxiv.org/pdf/1705.10513.pdf) for a learning to rank of a pairwise approach published on SIGIR 2017.
# + [markdown] deletable=true editable=true
# Copyright (C) 2017 <NAME>
#
# Permission is hereby granted, free of charge, to any person obtaining a copy of this software and associated documentation files (the "Software"), to deal in the Software without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Software, and to permit persons to whom the Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
# + deletable=true editable=true
import numpy as np
import tensorflow as tf
# + [markdown] deletable=true editable=true
# ## Dataset (MQ2008-semi)
#
# You should download the data set from [here](https://drive.google.com/drive/folders/0B-dulzPp3MmCM01kYlhhNGQ0djA?usp=sharing), and then put in MQ2008-semi/.
# + deletable=true editable=true
class MQ2008NoTargetException(Exception):
pass
class MQ2008:
def __init__(self, dataset_dir='MQ2008-semi'):
self.pool = {}
self.pool['train'] = self._load_data(dataset_dir + '/train.txt')
self.pool['test'] = self._load_data(dataset_dir + '/test.txt')
def get_queries(self, target='train'):
if target in self.pool.keys():
return list(self.pool[target].keys())
else:
raise MQ2008NoTargetException()
def get_docs(self, query, target='train'):
if target in self.pool.keys():
return list(self.pool[target][query].keys())
else:
raise MQ2008NoTargetException()
def get_features(self, query, doc, target='train'):
if target in self.pool.keys():
return self.pool[target][query][doc]['f']
else:
raise MQ2008NoTargetException()
def get_rank(self, query, doc, target='train'):
if target in self.pool.keys():
return self.pool[target][query][doc]['r']
else:
raise MQ2008NoTargetException()
def get_pos_queries(self, target='train'):
if target in self.pool.keys():
return list({query for query in self.get_queries(target=target)
for doc in self.get_docs(query, target=target) if self.get_rank(query, doc, target=target) > 0.0})
else:
raise MQ2008NoTargetException()
def get_pos_docs(self, query, target='train'):
if target in self.pool.keys():
return list({doc for doc in self.get_docs(query, target=target) if self.get_rank(query, doc, target=target) > 0.0})
else:
raise MQ2008NoTargetException()
# load docs and features for a query.
def _load_data(self, file, feature_size=46):
query_doc_feature = {}
with open(file) as f:
for line in f:
cols = line.strip().split()
rank = cols[0]
query = cols[1].split(':')[1]
doc = cols[-7]
feature = []
for i in range(2, 2 + feature_size):
feature.append(float(cols[i].split(':')[1]))
if query in query_doc_feature.keys():
query_doc_feature[query][doc] = {'r': float(rank), 'f': np.array(feature)}
else:
query_doc_feature[query] = {doc: {'r': float(rank), 'f': np.array(feature)}}
return query_doc_feature
# + [markdown] deletable=true editable=true
# ## Generator
# + [markdown] deletable=true editable=true
# Here we need to use `tf.variable_scope` for two reasons. Firstly, we're going to make sure all the variable names start with `generator`. Similarly, we'll prepend `discriminator` to the discriminator variables. This will help out later when we're training the separate networks.
#
# Here's more from [the TensorFlow documentation](https://www.tensorflow.org/programmers_guide/variable_scope#the_problem) to get another look at using `tf.variable_scope`.
# + deletable=true editable=true
class Generator:
def __init__(self, feature_size, hidden_size, keep_prob=1.0):
self.feature_size = feature_size
self.hidden_size = hidden_size
self.keep_prob = keep_prob
with tf.variable_scope('generator'):
# input placeholders
self.reward = tf.placeholder(tf.float32, [None], name='reward')
self.pred_data = tf.placeholder(tf.float32, [None, self.feature_size], name='pred_data')
self.sample_index = tf.placeholder(tf.int32, [None], name='sample_index')
########## score of RankNet ##########
# trainable variables
self.weight_1 = tf.Variable(tf.truncated_normal([self.feature_size, self.hidden_size], mean=0.0, stddev=0.1), name='weight_1')
self.bias_1 = tf.Variable(tf.zeros([self.hidden_size]), name='bias_1')
self.weight_2 = tf.Variable(tf.truncated_normal([self.hidden_size, 1], mean=0.0, stddev=0.1), name='weight_2')
self.bias_2 = tf.Variable(tf.zeros([1]), name='bias_2')
# layer 1 (hidden layer)
self.layer_1 = tf.nn.tanh(tf.nn.xw_plus_b(self.pred_data, self.weight_1, self.bias_1))
# dropout
self.dropout_1 = tf.nn.dropout(self.layer_1, self.keep_prob)
# layer 2 (output layer)
self.layer_2 = tf.nn.xw_plus_b(self.dropout_1, self.weight_2, self.bias_2)
#################################
# probability distribution
self.opt_prob =tf.gather(tf.reshape(tf.nn.softmax(tf.reshape(self.layer_2, [1, -1])), [-1]), self.sample_index)
# loss for optimization
self.opt_loss = -tf.reduce_mean(tf.log(self.opt_prob) * self.reward) # minus signe is needed for maximum.
# score for prediction
self.pred_score = tf.nn.xw_plus_b(self.layer_1, self.weight_2, self.bias_2)
self.pred_score = tf.reshape(self.pred_score, [-1])
# + [markdown] deletable=true editable=true
# ## Discriminator
#
# This is a pairwaise case implimentation.
# + deletable=true editable=true
class Discriminator:
def __init__(self, feature_size, hidden_size, keep_prob=1.0):
self.feature_size = feature_size
self.hidden_size = hidden_size
self.keep_prob = keep_prob
with tf.variable_scope('discriminator'):
# input placeholders
self.pos_data = tf.placeholder(tf.float32, [None, self.feature_size], name='pos_data')
self.neg_data = tf.placeholder(tf.float32, [None, self.feature_size], name='neg_data')
self.pred_data = tf.placeholder(tf.float32, [None, self.feature_size], name='pred_data')
########## score of RankNet ##########
## trainable variables
self.weight_1 = tf.Variable(tf.truncated_normal([self.feature_size, self.hidden_size], mean=0.0, stddev=0.1), name='weight_1')
self.bias_1 = tf.Variable(tf.zeros([self.hidden_size]), name='bias_1')
self.weight_2 = tf.Variable(tf.truncated_normal([self.hidden_size, 1], mean=0.0, stddev=0.1), name='weight_2')
self.bias_2 = tf.Variable(tf.zeros([1]), name='bias_2')
# layer 1 (hidden layer)
self.pos_layer_1 = tf.nn.tanh(tf.nn.xw_plus_b(self.pos_data, self.weight_1, self.bias_1))
self.neg_layer_1 = tf.nn.tanh(tf.nn.xw_plus_b(self.neg_data, self.weight_1, self.bias_1))
# dropout
self.pos_dropout_1 = tf.nn.dropout(self.pos_layer_1, self.keep_prob)
self.neg_dropout_1 = tf.nn.dropout(self.neg_layer_1, self.keep_prob)
# layer 2 (output layer)
self.pos_layer_2 = tf.nn.xw_plus_b(self.pos_dropout_1, self.weight_2, self.bias_2)
self.neg_layer_2 = tf.nn.xw_plus_b(self.neg_dropout_1, self.weight_2, self.bias_2)
#################################
# loss for optimization
self.opt_loss = -tf.reduce_mean(tf.log(tf.sigmoid(self.pos_layer_2 - self.neg_layer_2))) # minus signe is needed for miximum
# reward for generator
self.reward = tf.reshape(tf.log(1 + tf.exp(self.neg_layer_2 - self.pos_layer_2)), [-1])
# score for prediction
self.pred_score = tf.nn.xw_plus_b(tf.nn.tanh(tf.nn.xw_plus_b(self.pred_data, self.weight_1, self.bias_1)), self.weight_2, self.bias_2)
self.pred_score = tf.reshape(self.pred_score , [-1])
# + [markdown] deletable=true editable=true
# ## Optimizer
#
# We want to update the generator and discriminator variables separately. So we need to get the variables for each part build optimizers for the two parts. To get all the trainable variables, we use `tf.trainable_variables()`. This creates a list of all the variables we've defined in our graph.
#
# For the generator optimizer, we only want to generator variables. Our past selves were nice and used a variable scope to start all of our generator variable names with `generator`. So, we just need to iterate through the list from `tf.trainable_variables()` and keep variables to start with `generator`. Each variable object has an attribute `name` which holds the name of the variable as a string (`var.name == 'weights_0'` for instance).
#
# We can do something similar with the discriminator. All the variables in the discriminator start with `discriminator`.
#
# Then, in the optimizer we pass the variable lists to `var_list` in the `minimize` method. This tells the optimizer to only update the listed variables. Something like `tf.train.AdamOptimizer().minimize(loss, var_list=var_list)` will only train the variables in `var_list`.
# + deletable=true editable=true
class Optimizer:
def __init__(self, g, d, learning_rate):
# get the trainable_variables, split into generator and discriminator parts.
t_vars = tf.trainable_variables()
self.g_vars = [var for var in t_vars if var.name.startswith('generator')]
self.d_vars = [var for var in t_vars if var.name.startswith('discriminator')]
self.g_train_opt = tf.train.AdamOptimizer(learning_rate).minimize(g.opt_loss, var_list=self.g_vars)
self.d_train_opt = tf.train.AdamOptimizer(learning_rate).minimize(d.opt_loss, var_list=self.d_vars)
# + [markdown] deletable=true editable=true
# ## Dataset (MQ2008-semi extension)
# + [markdown] deletable=true editable=true
# If there are only two levels of relevance and for each "observed" relevant-irrelevant document pair (d_i, d_j) we sample an unlabelled document d_k to form the "generated" document pair (d_k, d_j), then it can be shown that the objective function of the IRGAN-pairwise minimax game Eq. (7) in the paper is bounded by the mathematical expectation of (f_phi(d_i, q) - f_phi(d_k, q)) / 2 which is independent of the irrelevant document d_j, via a straightforward application of Jensen's inequality on the logarithm function.
# + deletable=true editable=true
class Dataset(MQ2008):
def __init__(self, batch_size, dataset_dir='MQ2008-semi'):
MQ2008.__init__(self, dataset_dir=dataset_dir)
self.batch_size = batch_size
self.docs_pairs = []
def set_docs_pairs(self, sess, generator):
for query in dataset.get_pos_queries():
can_docs = dataset.get_docs(query)
can_features = [dataset.get_features(query, doc) for doc in can_docs]
can_score = sess.run(generator.pred_score, feed_dict={generator.pred_data: can_features})
# softmax for candidate
exp_rating = np.exp(can_score)
prob = exp_rating / np.sum(exp_rating)
pos_docs = dataset.get_pos_docs(query)
neg_docs = []
for i in range(len(pos_docs)):
while True:
doc = np.random.choice(can_docs, p=prob)
if doc not in pos_docs:
neg_docs.append(doc)
break
for i in range(len(pos_docs)):
self.docs_pairs.append((query, pos_docs[i], neg_docs[i]))
def get_batches(self):
size = len(self.docs_pairs)
cut_off = size // self.batch_size
for i in range(0, self.batch_size * cut_off, self.batch_size):
batch_pairs = self.docs_pairs[i:i+self.batch_size]
yield np.asarray([self.get_features(p[0], p[1]) for p in batch_pairs]), np.asarray([self.get_features(p[0], p[2]) for p in batch_pairs])
# + [markdown] deletable=true editable=true
# ## Training
# + [markdown] deletable=true editable=true
# There are some auxiliary functions.
# + deletable=true editable=true
def train_generator(sess, generator, discriminator, optimizer, dataset):
for query in dataset.get_pos_queries():
pos_docs = dataset.get_pos_docs(query)
can_docs = dataset.get_docs(query)
can_features = [dataset.get_features(query, doc) for doc in can_docs]
can_score = sess.run(generator.pred_score, feed_dict={generator.pred_data: can_features})
# softmax for all
exp_rating = np.exp(can_score)
prob = exp_rating / np.sum(exp_rating)
# sampling
neg_index = np.random.choice(np.arange(len(can_docs)), size=[len(pos_docs)], p=prob)
neg_docs = np.array(can_docs)[neg_index]
pos_features = [dataset.get_features(query, doc) for doc in pos_docs]
neg_features = [dataset.get_features(query, doc) for doc in neg_docs]
neg_reward = sess.run(discriminator.reward,
feed_dict={discriminator.pos_data: pos_features, discriminator.neg_data: neg_features})
_ = sess.run(optimizer.g_train_opt,
feed_dict={generator.pred_data: can_features, generator.sample_index: neg_index, generator.reward: neg_reward})
return sess.run(generator.opt_loss,
feed_dict={generator.pred_data: can_features, generator.sample_index: neg_index, generator.reward: neg_reward})
# + deletable=true editable=true
def train_discriminator(sess, generator, discriminator, optimizer, dataset):
dataset.set_docs_pairs(sess, generator)
for input_pos, input_neg in dataset.get_batches():
_ = sess.run(optimizer.d_train_opt,
feed_dict={discriminator.pos_data: input_pos, discriminator.neg_data: input_neg})
return sess.run(discriminator.opt_loss,
feed_dict={discriminator.pos_data: input_pos, discriminator.neg_data: input_neg})
# + deletable=true editable=true
def ndcg_at_k(sess, discriminator, dataset, k=5):
ndcg = 0.0
cnt = 0
for query in dataset.get_pos_queries(target='test'):
pos_docs = dataset.get_pos_docs(query, target='test')
pred_docs = dataset.get_docs(query, target='test')
if len(pred_docs) < k:
continue
pred_features = np.asarray([dataset.get_features(query, doc, target='test') for doc in pred_docs])
pred_score = sess.run(discriminator.pred_score, feed_dict={discriminator.pred_data: pred_features})
pred_doc_score = sorted(zip(pred_docs, pred_score), key=lambda x: x[1], reverse=True)
dcg = 0.0
for i in range(0, k):
doc, _ = pred_doc_score[i]
if doc in pos_docs:
dcg += (1 / np.log2(i + 2))
n = len(pos_docs) if len(pos_docs) < k else k
idcg = np.sum(np.ones(n) / np.log2(np.arange(2, n + 2)))
ndcg += (dcg / idcg)
cnt += 1
return ndcg / float(cnt)
# + [markdown] deletable=true editable=true
# There are hyperparameters.
# + deletable=true editable=true
# size of input vector
feature_size = 46
# size of latent vector
hidden_size = 46
# keep probability
keep_prod = 0.5
# learning_rate
learning_rate = 0.00001
# batch_size
batch_size = 8
# generator training epochs
epochs = 30
# + [markdown] deletable=true editable=true
# Some objects are created.
# + deletable=true editable=true
dataset = Dataset(batch_size)
# + deletable=true editable=true
generator = Generator(feature_size, hidden_size, keep_prod)
discriminator = Discriminator(feature_size, hidden_size, keep_prod)
optimizer = Optimizer(generator, discriminator, learning_rate)
# + [markdown] deletable=true editable=true
# A generator and discriminator are optimized with dataset.
# + deletable=true editable=true
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
for e in range(epochs):
g_loss = train_generator(sess, generator, discriminator, optimizer, dataset)
d_loss = train_discriminator(sess, generator, discriminator, optimizer, dataset)
ndcg_at_3 = ndcg_at_k(sess, discriminator, dataset, k=3)
ndcg_at_5 = ndcg_at_k(sess, discriminator, dataset, k=5)
ndcg_at_10 = ndcg_at_k(sess, discriminator, dataset, k=10)
print("Epoch {}/{}...".format(e+1, epochs),
"Generator Loss: {:.4f}".format(g_loss),
"Discriminator Loss: {:.4f}".format(d_loss),
"NDCG@3: {:.4f}".format(ndcg_at_3),
"NDCG@5: {:.4f}".format(ndcg_at_5),
"NDCG@10: {:.4f}".format(ndcg_at_10))
| mlmodels/model_rank/irgan_keras/IRGAN_Learning_to_Rank_Pairwise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear Regression
import matplotlib.pyplot as plt
import numpy as np
from sklearn import datasets, linear_model
from sklearn.metrics import mean_squared_error, r2_score
from sklearn.model_selection import train_test_split
# +
# load the diabetes dataset
diabetes = datasets.load_diabetes()
diabetes_x = diabetes.data
diabetes_y = diabetes.target
# train_test_split
diabetes_x_train, diabetes_x_test, diabetes_y_train, diabetes_y_test = train_test_split(diabetes_x, diabetes_y, test_size=0.2)
# build linear regression model
model = linear_model.LinearRegression()
# train the model using the training sets
model.fit(diabetes_x_train, diabetes_y_train)
# 把測試資料丟進模型predict y
diabetes_y_pred = model.predict(diabetes_x_test)
# mean squared error
print(f'Mean squared error: {mean_squared_error(diabetes_y_test, diabetes_y_pred)}')
# r2 score
print(f'Variance score: {r2_score(diabetes_y_test, diabetes_y_pred)}')
# -
# # Polynomial Regression
import numpy as np
import pandas as pd
from sklearn.linear_model import Ridge
from sklearn.preprocessing import PolynomialFeatures
from sklearn.pipeline import make_pipeline
from sklearn import preprocessing, linear_model
# +
def f(x):
return x**3-x*2+3
# creating 100 data
X = np.linspace(0,10,100)
np.random.shuffle(X)
X = X.reshape(100,1)
y = f(X)
poly = PolynomialFeatures(degree=2).fit(X)
X_poly = poly.transform(X)
# train-test split
x_train, x_test, y_train, y_test = train_test_split(X_poly, y, test_size=0.3, random_state=1)
# Training data normalization
scaler = preprocessing.StandardScaler().fit(x_train)
x_train = scaler.transform(x_train)
# build model
model = linear_model.LinearRegression()
model.fit(x_train, y_train)
# Testing data normalization
x_test = scaler.transform(x_test)
# predict y
y_pred = model.predict(x_test)
print(f'Mean squared error: {mean_squared_error(y_test, y_pred)}')
print(f'Variance score: {r2_score(y_test, y_pred)}')
# -
# # Logistic Regression
import numpy as np
from sklearn.linear_model import LogisticRegression
from sklearn import datasets, preprocessing
from sklearn.model_selection import train_test_split
# +
iris = datasets.load_iris()
x = iris.data
y = iris.target
# train-test split
x_train,x_test,y_train,y_test=train_test_split(x, y, test_size=0.3, random_state=1)
# Training data normalization
scaler = preprocessing.StandardScaler().fit(x_train)
x_train = scaler.transform(x_train)
# build model
model = LogisticRegression()
model.fit(x_train, y_train)
# Testing data normalization
x_test = scaler.transform(x_test)
# predict y
y_pred = model.predict(x_test)
# model直接計算y_pred的正確率
# accuracy = model.score(x_test, y_test)
from sklearn.metrics import mean_squared_error, r2_score, accuracy_score, confusion_matrix
accuracy = accuracy_score(y_test, y_pred)
print(f'Accuracy: {accuracy}')
# -
| Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from sklearn.datasets import load_boston
from sklearn.neural_network import MLPRegressor
from sklearn.preprocessing import MinMaxScaler
boston = load_boston()
X = MinMaxScaler().fit(boston.data).transform(boston.data)
Y = MinMaxScaler().fit(boston.target.reshape(-1,1)).transform(boston.target.reshape(-1,1))
adaline = MLPRegressor(hidden_layer_sizes=[],solver="sgd",alpha=0.1,batch_size=1,max_iter=5000,momentum=0,verbose=True)
adaline.fit(X,Y)
| Boston.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Domain Generation Algorithm (DGA) Detection
#
# ## Authors
# - <NAME> (NVIDIA) [<EMAIL>]
# - <NAME> (NVIDIA) [<EMAIL>]
#
# ## Development Notes
# * Developed using: RAPIDS v0.12.0 and CLX v0.12
# * Last tested using: RAPIDS v21.08 and CLX v21.08 on Aug 18, 2021
#
# ## Table of Contents
# * Introduction
# * Data Importing
# * Data Preprocessing
# * Training and Evaluation
# * Inference
# * Conclusion
#
# ## Introduction
# [Domain Generation Algorithms](https://en.wikipedia.org/wiki/Domain_generation_algorithm) (DGAs) are used to generate domain names that can be used by the malware to communicate with the command and control servers. IP addresses and static domain names can be easily blocked, and a DGA provides an easy method to generate a large number of domain names and rotate through them to circumvent traditional block lists. We will use a type of recurrent neural network called the [Gated Recurrent Unit](https://towardsdatascience.com/illustrated-guide-to-lstms-and-gru-s-a-step-by-step-explanation-44e9eb85bf21) (GRU) for this example. The [CLX](https://github.com/rapidsai/clx) and [RAPIDS](https://rapids.ai) libraries enable users train their models with up-to-date domain names representative of both benign and DGA generated strings. Using a CLX workflow, this capability could also be used in production. This notebook provides a view into the data science workflow to create a DGA detection implementation.
import os
import cudf
import torch
import s3fs
import logging
import numpy as np
from datetime import datetime
from sklearn.metrics import accuracy_score, average_precision_score
from clx.analytics.dga_detector import DGADetector
from clx.utils.data.dataloader import DataLoader
from clx.analytics.dga_dataset import DGADataset
from cuml.model_selection import train_test_split
# #### Enable console logging
logging.basicConfig(
level=logging.INFO,
format="%(asctime)s [%(levelname)s] %(message)s",
handlers=[
logging.StreamHandler()
]
)
# #### Download Input Dataset from S3
# +
INPUT_CSV = "benign_and_dga_domains.csv"
S3_BASE_PATH = "rapidsai-data/cyber/clx"
# -
# Read Benign and DGA dataset
if not os.path.exists(INPUT_CSV):
fs = s3fs.S3FileSystem(anon=True)
fs.get(S3_BASE_PATH + "/" + INPUT_CSV, INPUT_CSV)
# #### Load Input Dataset to GPU Dataframe
gdf = cudf.read_csv(INPUT_CSV)
train_data = gdf['domain']
labels = gdf['type']
# Because we have only benign and DGA (malicious) categoriesm, the number of domain types need to be set to 2 (`N_DOMAIN_TYPE=2`). Vocabulary size(`CHAR_VOCAB`) is set to 128 ASCII characters. The values below set for `HIDDEN_SIZE`, `N_LAYERS` of the network, and the `LR` (Learning Rate) give an optimum balance for the network size and performance. They might need be set via experiments when working with other datasets.
LR = 0.001
N_LAYERS = 3
CHAR_VOCAB = 128
HIDDEN_SIZE = 100
N_DOMAIN_TYPE = 2
# #### Instantiate DGA Detector
# Now that the data is ready, the datasets are created, and we've set the parameters for the model, we can use the DGADetector method built into CLX to create and train the model.
dd = DGADetector(lr=LR)
dd.init_model(n_layers=N_LAYERS, char_vocab=CHAR_VOCAB, hidden_size=HIDDEN_SIZE, n_domain_type=N_DOMAIN_TYPE)
EPOCHS = 25
TRAIN_SIZE = 0.7
BATCH_SIZE = 10000
MODELS_DIR = 'models'
# ### Training and Evaluation
# Now we train and evaluate the model.
# %%time
dd.train_model(train_data, labels, batch_size=BATCH_SIZE, epochs=EPOCHS, train_size=0.7)
# ### Save Model
# Save pretrained model to a given output location.
# +
if not os.path.exists(MODELS_DIR):
print("Creating directory '{}'".format(MODELS_DIR))
os.makedirs(MODELS_DIR)
now = datetime.now()
model_filename = "rnn_classifier_{}.bin".format(now.strftime("%Y-%m-%d_%H_%M_%S"))
model_filepath = os.path.join(MODELS_DIR, model_filename)
dd.save_checkpoint(model_filepath)
# -
# ### Inference
# Using the model generated above, we now score the test dataset against the model to determine if the domain is likely generated by a DGA or benign.
# +
dga_detector = DGADetector()
dga_detector.load_checkpoint(model_filepath)
domain_train, domain_test, type_train, type_test = train_test_split(gdf, "type", train_size=0.7)
test_df = cudf.DataFrame()
test_df["type"] = type_test.reset_index(drop=True)
test_df["domain"] = domain_test.reset_index(drop=True)
test_dataset = DGADataset(test_df, 100)
test_dataloader = DataLoader(test_dataset, batchsize=BATCH_SIZE)
pred_results = []
true_results = []
for chunk in test_dataloader.get_chunks():
pred_results.append(list(dga_detector.predict(chunk['domain']).values_host))
true_results.append(list(chunk['type'].values_host))
pred_results = np.concatenate(pred_results)
true_results = np.concatenate(true_results)
accuracy_score_result = accuracy_score(pred_results, true_results)
print('Model accuracy: %s'%(accuracy_score_result))
# +
average_precision = average_precision_score(true_results, pred_results)
print('Average precision score: {0:0.3f}'.format(average_precision))
# -
# ## Conclusion
# DGA detector in CLX enables users to train their models for detection and also use existing models. This capability could also be used in conjunction with log parsing efforts if the logs contain domain names. DGA detection done with CLX and RAPIDS keeps data in GPU memory, removing unnecessary copy/converts and providing a 4X speed advantage over CPU only implementations. This is esepcially true with large batch sizes.
| notebooks/dga_detection/DGA_Detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib notebook
import seaborn as sns
# -
plt.style.use("seaborn-colorblind")
data = pd.read_csv("C:/Users/VARUN/Desktop/AirPollution/Dataset/cpcb_dly_aq_gujarat-2014.csv")
data.head()
dates = ['-'.join(i.split('-')[1:]) for i in data['Sampling Date']]
data['Sampling Date'] = dates
for i in range(len(data['City/Town/Village/Area'])):
if data['City/Town/Village/Area'][i] != 'Ahmedabad':
data.drop(i, inplace = True)
data.head()
data = data.groupby("Sampling Date").mean()
data
data = data.groupby("Sampling Date").mean().drop("PM 2.5", axis=1)
data
# +
plt.figure(figsize=(9, 7))
plt.subplot(211)
plt.cla()
plotter = ['Jan', 'Feb', 'Mar', 'Apr', 'May', 'Jun','Jul', 'Aug', 'Sep',
'Oct', 'Nov', 'Dec']
x = np.arange(0,len(data["NO2"]),1)
ax = plt.gca()
ax.plot(x, data["NO2"])
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.set_xticks(x)
plt.xticks(rotation=45)
plt.subplots_adjust(bottom=0.2)
plt.legend([2014],loc=2)
plt.title("$\mathregular{NO_2}$ (Nitrogen Dioxide) levels in Ahmedabad")
ax.set_xticklabels(plotter)
plt.subplot(212)
plt.cla()
plotter = ['Jan', 'Feb', 'Mar', 'Apr', 'May', 'Jun','Jul', 'Aug', 'Sep',
'Oct', 'Nov', 'Dec']
x = np.arange(0,len(data["SO2"]),1)
ax = plt.gca()
ax.plot(x, data["SO2"])
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.set_xticks(x)
plt.xticks(rotation=45)
plt.title("$\mathregular{SO_2}$ (Sulphur Dioxide) levels in Ahmedabad")
ax.set_xticklabels(plotter)
plt.tight_layout()
#plt.show()
# -
| Air-Pollution-Levels-Exploratory-Data-Analysis-master/Ahmedabad.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PySpark
# language: ''
# name: pysparkkernel
# ---
# + id="62QKNkJ1Sqf_" colab={"referenced_widgets": [""]} outputId="dbfc58fa-f11e-4a92-f434-4c1b7714ee7e"
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName('Final Project').master('spark://spark-master:7077').getOrCreate()
#spark = SparkSession.builder.appName('Final Project').master('local(*)').getOrCreate()
# + id="IaGOGlr4Sqih" colab={"referenced_widgets": [""]} outputId="a55f5923-540b-43ce-a70a-37d97e2c7420"
import pyspark.sql.functions as fn
from pyspark.sql.types import StructType, StructField, StringType,IntegerType
import pandas as pd
import numpy as np
import pyspark.ml.evaluation as ev
from pyspark.ml import Pipeline
from pyspark.ml.feature import StringIndexer
import pyspark.sql.types as types
import pyspark.ml.feature as ft
import pyspark.ml.classification as cl
from pyspark.ml import PipelineModel
import pyspark.ml.tuning as tune
from pyspark.sql.functions import isnan, when, count, col
# + id="T-Q5l9Q1Sqit" colab={"referenced_widgets": [""]} outputId="357a251a-fa66-46ca-cba2-bddcedb9dacd"
T4 = spark.read.csv('s3://msbx5420-spr21/sherlock/T4_filtered.tsv', sep=r'\t', header=False,inferSchema = True).toDF('UserId','UUID','t4_version','CpuHertz','CPU_0','CPU_1','CPU_2','CPU_3','Total_CPU',
'TotalMemory_freeSize','TotalMemory_max_size','TotalMemory_total_size','TotalMemory_used_size',
'Traffic_MobileRxBytes','Traffic_MobileRxPackets','Traffic_MobileTxBytes',
'Traffic_MobileTxPackets','Traffic_TotalRxBytes','Traffic_TotalRxPackets','Traffic_TotalTxBytes',
'Traffic_TotalTxPackets','Traffic_TotalWifiRxBytes','Traffic_TotalWifiRxPackets',
'Traffic_TotalWifiTxBytes','Traffic_TotalWifiTxPackets','Traffic_timestamp','Battery_charge_type',
'Battery_current_avg','Battery_health','Battery_icon_small','Battery_invalid_charger',
'Battery_level','Battery_online','Battery_plugged','Battery_present','Battery_scale',
'Battery_status','Battery_technology','Battery_temperature','Battery_timestamp','Battery_voltage',
'MemTotal','MemFree','Buffers','Cached','SwapCached','Active','Inactive','Active_anon',
'Inactive_anon','Active_file','Inactive_file','Unevictable','Mlocked','HighTotal','HighFree',
'LowTotal','LowFree','SwapTotal','SwapFree','Dirty','Writeback','AnonPages','Mapped',
'Shmem','Slab','SReclaimable','SUnreclaim','KernelStack','PageTables','CommitLimit','Committed_AS',
'VmallocTotal','VmallocUsed','VmallocChunk','msmgpio_cpu0','msmgpio_sum_cpu123','wcd9xxx_cpu0',
'wcd9xxx_sum_cpu123','pn547_cpu0','pn547_sum_cpu123','cypress_touchkey_cpu0',
'cypress_touchkey_sum_cpu123','synaptics_rmi4_i2c_cpu0','synaptics_rmi4_i2c_sum_cpu123',
'sec_headset_detect_cpu0','sec_headset_detect_sum_cpu123','flip_cover_cpu0','flip_cover_sum_cpu123',
'home_key_cpu0','home_key_sum_cpu123','volume_down_cpu0','volume_down_sum_cpu123','volume_up_cpu0',
'volume_up_sum_cpu123','companion_cpu0','companion_sum_cpu123','SLIMBUS_cpu0','SLIMBUS_sum_cpu123',
'function_call_interrupts_cpu0','function_call_interrupts_sum_cpu123','cpu123_intr_prs','tot_user',
'tot_nice','tot_system','tot_idle','tot_iowait','tot_irq','tot_softirq','ctxt','btime','processes',
'procs_running','procs_blocked','connectedWifi_SSID','connectedWifi_Level', '117','118','119',
'120', '121','122','123','124','125','126','127','128','129','130')
T4 = T4.withColumn("CPU_0",fn.col("CPU_0").cast('double')).withColumn("CPU_1",fn.col("CPU_1").cast('double')).withColumn("CPU_2",fn.col("CPU_2").cast('double')).withColumn("CPU_3",fn.col("CPU_3").cast('double'))
T4 = T4.fillna(value=0,subset=['CPU_0','CPU_1','CPU_2','CPU_3'])
cols = ['TotalMemory_freeSize','TotalMemory_max_size','TotalMemory_total_size','TotalMemory_used_size',
'Traffic_MobileRxBytes','Traffic_MobileRxPackets','Traffic_MobileTxBytes',
'Traffic_MobileTxPackets','Traffic_TotalRxBytes','Traffic_TotalRxPackets','Traffic_TotalTxBytes',
'Traffic_TotalTxPackets','Traffic_TotalWifiRxBytes','Traffic_TotalWifiRxPackets',
'Traffic_TotalWifiTxBytes','Traffic_TotalWifiTxPackets','Traffic_timestamp','Battery_charge_type',
'Battery_current_avg','Battery_health','Battery_icon_small','Battery_invalid_charger',
'Battery_level','Battery_online','Battery_plugged','Battery_present','Battery_scale',
'Battery_status','Battery_technology','Battery_temperature','Battery_timestamp','Battery_voltage',
'MemTotal','MemFree','Buffers','Cached','SwapCached','Active','Inactive','Active_anon',
'Inactive_anon','Active_file','Inactive_file','Unevictable','Mlocked','HighTotal','HighFree',
'LowTotal','LowFree','SwapTotal','SwapFree','Dirty','Writeback','AnonPages','Mapped',
'Shmem','Slab','SReclaimable','SUnreclaim','KernelStack','PageTables','CommitLimit','Committed_AS',
'VmallocTotal','VmallocUsed','VmallocChunk','msmgpio_cpu0','msmgpio_sum_cpu123','wcd9xxx_cpu0',
'wcd9xxx_sum_cpu123','pn547_cpu0','pn547_sum_cpu123','cypress_touchkey_cpu0',
'cypress_touchkey_sum_cpu123','synaptics_rmi4_i2c_cpu0','synaptics_rmi4_i2c_sum_cpu123',
'sec_headset_detect_cpu0','sec_headset_detect_sum_cpu123','flip_cover_cpu0','flip_cover_sum_cpu123',
'home_key_cpu0','home_key_sum_cpu123','volume_down_cpu0','volume_down_sum_cpu123','volume_up_cpu0',
'volume_up_sum_cpu123','companion_cpu0','companion_sum_cpu123','SLIMBUS_cpu0','SLIMBUS_sum_cpu123',
'function_call_interrupts_cpu0','function_call_interrupts_sum_cpu123','cpu123_intr_prs','tot_user',
'tot_nice','tot_system','tot_idle','tot_iowait','tot_irq','tot_softirq','ctxt','btime','processes',
'procs_running','procs_blocked','connectedWifi_SSID','connectedWifi_Level', '117','118','119',
'120', '121','122','123','124','125','126','127','128','129','130']
T4 = T4.drop(*cols)
T4.printSchema()
# + id="fy-Kg_1sSqjG" colab={"referenced_widgets": [""]} outputId="5857f27e-3c52-4361-c0bc-01aa97b8e7b7"
Moriarty = spark.read.options(inferSchema = True).csv('s3://msbx5420-spr21/sherlock/Moriarty.csv', header=True)
Moriarty = Moriarty.withColumnRenamed("Version","Version_Moriarty")
# + id="EOqyXzZOSqjN" colab={"referenced_widgets": [""]} outputId="badbdcc8-cb03-4ff7-9555-b6c8365d8df5"
Moriarty = Moriarty.select("*").toPandas()
Moriarty['ActionType_binary'] = np.where(Moriarty['ActionType']== 'malicious', 1, 0)
Moriarty['target'] = np.where(Moriarty['SessionType']== 'malicious', 1, 0)
Moriarty = Moriarty.dropna()
Moriarty = spark.createDataFrame(Moriarty)
# + id="VHleJJz5SqjP" colab={"referenced_widgets": [""]} outputId="a9e84e01-591f-4fd9-afa6-49ec9239c934"
ScreenOn = spark.read.options(inferSchema = True).csv('s3://msbx5420-spr21/sherlock/ScreenOn.csv', header=True)
ScreenOn = ScreenOn.withColumn("Timestamp",fn.to_timestamp("Timestamp"))
ScreenOn = ScreenOn.withColumn("ScreenOn",fn.col("ScreenOn").cast("Integer"))
ScreenOn = ScreenOn.withColumnRenamed('Timestamp', 'Timestamp_ScreenOn').withColumnRenamed('UserID', 'UserId')
ScreenOn.printSchema()
# + [markdown] id="pwJYxv76SqjS"
# ## Join DF's
# + id="WtK7WBw6SqjU" colab={"referenced_widgets": [""]} outputId="b33dd194-cbb8-4c23-ff3d-4825f185a63b"
Moriarty.createOrReplaceTempView("Moriarty")
T4.createOrReplaceTempView("T4")
ScreenOn.createOrReplaceTempView("ScreenOn")
# + id="shHUK-WWSqjY" colab={"referenced_widgets": [""]} outputId="e3f17736-8375-4f66-b865-1583e4eb865e"
ms4 = Moriarty.join(ScreenOn, ['UserId','UUID'],'full')
ms4 = ms4.join(T4, ['UserId','UUID'],'full')
ms4.printSchema()
# + id="HkGpmGBnSqjc" colab={"referenced_widgets": [""]} outputId="10146590-fd65-4f36-c641-9cb87218db3b"
cols = ['Details','SessionType','ActionType','Version_Moriarty','SessionID','ActionType_binary','Timestamp_ScreenOn','t4_version','CpuHertz','Total_CPU']
ms4 = ms4.drop(*cols)
# + id="2Q1YUyRoSqje" colab={"referenced_widgets": [""]} outputId="8fa8fffd-560a-4ab0-f98a-0e824329b00e"
ms4 = ms4.fillna(value=0,subset=["target"])
ms4 = ms4.fillna(value='None',subset=["Action"])
# + id="OkGXyNBVSqjf" colab={"referenced_widgets": [""]} outputId="f621d2de-5580-4170-ca53-7fabfc9b9571"
# get the sort key
def getKey(item):
return item.UUID
# fill function
def fill(x):
out = []
last_val_so = None
last_val_0 = None
last_val_1 = None
last_val_2 = None
last_val_3 = None
for v in x:
if v["ScreenOn"] is None:
data = [v["UserId"], v["UUID"], v["Action"], v["target"], last_val_so, v["CPU_0"], v["CPU_1"], v["CPU_2"], v["CPU_3"]]
else:
data = [v["UserId"], v["UUID"], v["Action"], v["target"], v["ScreenOn"], v["CPU_0"], v["CPU_1"], v["CPU_2"], v["CPU_3"]]
last_val_so = v["ScreenOn"]
if v["CPU_0"] is None:
data = [v["UserId"], v["UUID"], v["Action"], v["target"], v["ScreenOn"], last_val_0, v["CPU_1"], v["CPU_2"], v["CPU_3"]]
else:
data = [v["UserId"], v["UUID"], v["Action"], v["target"], v["ScreenOn"], v["CPU_0"], v["CPU_1"], v["CPU_2"], v["CPU_3"]]
last_val_0 = v["CPU_0"]
if v["CPU_1"] is None:
data = [v["UserId"], v["UUID"], v["Action"], v["target"], v["ScreenOn"], v["CPU_0"], last_val_1, v["CPU_2"], v["CPU_3"]]
else:
data = [v["UserId"], v["UUID"], v["Action"], v["target"], v["ScreenOn"], v["CPU_0"], v["CPU_1"], v["CPU_2"], v["CPU_3"]]
last_val_1 = v["CPU_1"]
if v["CPU_2"] is None:
data = [v["UserId"], v["UUID"], v["Action"], v["target"], v["ScreenOn"], v["CPU_0"], v["CPU_1"], last_val_2, v["CPU_3"]]
else:
data = [v["UserId"], v["UUID"], v["Action"], v["target"], v["ScreenOn"], v["CPU_0"], v["CPU_1"], v["CPU_2"], v["CPU_3"]]
last_val_2 = v["CPU_2"]
if v["CPU_3"] is None:
data = [v["UserId"], v["UUID"], v["Action"], v["target"], v["ScreenOn"], v["CPU_0"], v["CPU_1"], v["CPU_2"], last_val_3]
else:
data = [v["UserId"], v["UUID"], v["Action"], v["target"], v["ScreenOn"], v["CPU_0"], v["CPU_1"], v["CPU_2"], v["CPU_3"]]
last_val_3 = v["CPU_3"]
out.append(data)
return out
# + id="HJ-S7MsASqjg" colab={"referenced_widgets": [""]} outputId="8ac4a6ee-7135-47ef-c9b9-1664d322f8ad"
# Partition the data
rdd = ms4.rdd.groupBy(lambda x: x.UserId).mapValues(list)
# Sort the data by date
rdd = rdd.mapValues(lambda x: sorted(x, key=getKey))
# fill missing value and flatten
rdd = rdd.mapValues(fill).flatMapValues(lambda x: x)
# discard the key
rdd = rdd.map(lambda v: v[1])
# + id="MJAydodqSqjs" colab={"referenced_widgets": [""]} outputId="fa700dce-ad0c-4393-f929-56bf575e28ba"
from pyspark.sql.types import StructType, StructField, StringType, DoubleType, IntegerType, LongType
schema = StructType([StructField("UserId", StringType(), True), StructField("UUID", LongType(), True), StructField("Action", StringType(), True), StructField("target", IntegerType(), True), StructField("ScreenOn", IntegerType(), True), StructField("CPU_0", DoubleType(), True), StructField("CPU_1", DoubleType(), True), StructField("CPU_2", DoubleType(), True), StructField("CPU_3", DoubleType(), True)])
ms4 = sqlContext.createDataFrame(rdd,schema)
ms4.printSchema()
# + id="E6BKWDI1Sqju" colab={"referenced_widgets": [""]} outputId="6f10671f-0bd9-4eb7-dba3-fad21bdc42d2"
ms4 = ms4.fillna(value=0,subset=["ScreenOn",'CPU_0','CPU_1','CPU_2','CPU_3'])
# + id="zkKJaydWSqjv" colab={"referenced_widgets": [""]} outputId="027ad550-7fa2-4749-c509-62f123f4833a"
ms4.write.parquet('s3://msbx5420-spr21/sherlock/ms4_update.parquet')
# + id="nDMXuZ3pSqjw" colab={"referenced_widgets": [""]} outputId="6046c5e2-86ca-482b-9d7f-91641d8991b8"
ms4 = spark.read.parquet('s3://msbx5420-spr21/sherlock/ms4_update.parquet', header=True, inferSchema=True)
ms4.printSchema()
# + [markdown] id="NlyaMpGzSqjx"
# ## Create Transformer
# + id="ltJwnK0hSqjy" colab={"referenced_widgets": [""]} outputId="22a63daf-42d6-4c41-b8f8-399b42efbc54"
ms4 = StringIndexer(inputCol='Action', outputCol="a_index").fit(ms4).transform(ms4)
# + id="rNzRtDZjSqj3" colab={"referenced_widgets": [""]} outputId="4144e7a8-506a-4432-e527-b227254eb6a5"
encoder = ft.OneHotEncoder(inputCol="a_index", outputCol='avec')
# + id="wl5Jx-rGSqj6" colab={"referenced_widgets": [""]} outputId="1e963f84-1c7a-4150-8878-efc5ad1e03d8"
featuresCreator = ft.VectorAssembler(inputCols=[col for col in ms4.columns[4:9]] + [encoder.getOutputCol()], outputCol='features')
# + [markdown] id="6pXE6ZUqSqj8"
# ## Create Estimator
# + id="XeuHVtnQSqj-" colab={"referenced_widgets": [""]} outputId="94ec0329-788b-49fb-afc6-81b744ccee72"
logistic = cl.LogisticRegression(maxIter=10, regParam=0.01, labelCol='target')
# + [markdown] id="uTuvdomRSqkA"
# ## Create Pipeline
# + id="g5BM-UvsSqkB" colab={"referenced_widgets": [""]} outputId="4e2c6374-9c30-4d6a-f2d9-e12480cdc319"
pipeline = Pipeline(stages=[encoder, featuresCreator, logistic])
# + [markdown] id="Sq_DhsgYSqkC"
# ## Fit Model
# + id="LKmiPNPLSqkD" colab={"referenced_widgets": [""]} outputId="236ef020-ac46-441c-a553-709488727a92"
ms4_train, ms4_test = ms4.randomSplit([0.7, 0.3], seed=200)
# + id="wiFcpZD7SqkE" colab={"referenced_widgets": [""]} outputId="93d0133d-fcb9-43bc-db6b-b0fb3b3a4b6a"
model = pipeline.fit(ms4_train)
test_model = model.transform(ms4_test)
test_model.take(5)
# + [markdown] id="4IC-FW7ySqkH"
# ## Model Performance
# + id="6oilOMeUSqkI" colab={"referenced_widgets": [""]} outputId="441c99ba-91e8-46f8-a48b-7d8eded3c1e2"
evaluator = ev.BinaryClassificationEvaluator(rawPredictionCol='probability', labelCol='target')
print(evaluator.evaluate(test_model, {evaluator.metricName: 'areaUnderROC'}))
print(evaluator.evaluate(test_model, {evaluator.metricName: 'areaUnderPR'}))
# + [markdown] id="8omOphLVSqkJ"
# ## Saving Model
# + id="M0XI1512SqkK" colab={"referenced_widgets": [""]} outputId="b9ae4c10-1bde-4277-859e-723dd53f299c"
pipeline_path = './Logistic_Pipeline'
pipeline.write().overwrite().save(pipeline_path)
model_path = './Logistic_Model'
model.write().overwrite().save(model_path)
# + [markdown] id="JN-lVpAxSqkq"
# ## Hyperparameter Tuning
# + id="DxPr_wWHSqkq" colab={"referenced_widgets": [""]} outputId="03b77485-b1eb-4af1-a5b1-bef20456cbfa"
logistic = cl.LogisticRegression(labelCol='target')
grid = tune.ParamGridBuilder().addGrid(logistic.maxIter, [2, 10, 50]).addGrid(logistic.regParam, [0.01, 0.05, 0.3]).build()
# + id="GFFrEF4DSqlB" colab={"referenced_widgets": [""]} outputId="4da94ddb-a124-4185-972a-5f14d6862b65"
evaluator = ev.BinaryClassificationEvaluator(rawPredictionCol='probability', labelCol='target')
# + id="zXnY__llSqlE" colab={"referenced_widgets": [""]} outputId="9c1da3a8-19b3-4d16-8881-6165dc685bb5"
cv = tune.CrossValidator(estimator=logistic, estimatorParamMaps=grid, evaluator=evaluator)
# + id="WPDupLgMSqlY" colab={"referenced_widgets": [""]} outputId="2bde3a45-1876-463c-a6ef-df080649f0e4"
pipeline = Pipeline(stages=[encoder, featuresCreator])
data_transformer = pipeline.fit(ms4_train)
# + id="EodYOp2mSqlZ" colab={"referenced_widgets": [""]} outputId="492993ab-f94e-4a6b-dc79-16fe344f907d"
cvModel = cv.fit(data_transformer.transform(ms4_train))
# + id="bPNJzgjPSqlb" colab={"referenced_widgets": [""]} outputId="a0ff7eeb-799b-42f9-d7dd-e2a6ad1e0641"
data_test = data_transformer.transform(ms4_test)
results = cvModel.transform(data_test)
print(evaluator.evaluate(results, {evaluator.metricName: 'areaUnderROC'}))
print(evaluator.evaluate(results, {evaluator.metricName: 'areaUnderPR'}))
# + id="3nQtOaWRSqlc" colab={"referenced_widgets": [""]} outputId="4b009992-75ad-4d0b-d8f8-992f51d05c32"
cvModel.getEstimatorParamMaps()
# + id="fnLG6HvWSqle" colab={"referenced_widgets": [""]} outputId="74a126c6-816a-45bd-b7fe-dc887fcdaa76"
cvModel.avgMetrics
# + id="-2u4PHzRSqlg" colab={"referenced_widgets": [""]} outputId="97b504fd-87ef-42ea-b9b8-ab43480dc8f6"
results = []
for params, metric in zip(cvModel.getEstimatorParamMaps(), cvModel.avgMetrics):
parameters = []
for key, paramValue in zip(params.keys(), params.values()):
parameters.append({key.name: paramValue})
results.append((parameters, metric))
sorted(results, key=lambda x: x[1], reverse=True)
# + [markdown] id="2-OJUMFBSqlk"
# ## Train-Validation Splitting
# + id="jpiEMl-lSqlk" colab={"referenced_widgets": [""]} outputId="7a0c83a7-9d57-417c-99ef-977e49594a81"
selector = ft.ChiSqSelector(numTopFeatures=5, featuresCol=featuresCreator.getOutputCol(), outputCol='selectedFeatures', labelCol='target')
logistic = cl.LogisticRegression(labelCol='target', featuresCol='selectedFeatures')
pipeline = Pipeline(stages=[encoder, featuresCreator, selector])
feature_transformer = pipeline.fit(ms4_train)
# + id="M0GCoc95Sqll" colab={"referenced_widgets": [""]} outputId="a0023aa6-0401-46d3-dc43-22c0b2a4cb10"
grid = tune.ParamGridBuilder().addGrid(logistic.maxIter, [2, 10, 50]).addGrid(logistic.regParam, [0.01, 0.05, 0.3]).build()
tvs = tune.TrainValidationSplit(estimator=logistic, estimatorParamMaps=grid, evaluator=evaluator, collectSubModels=True)
# + id="FaUndgROSqlo" colab={"referenced_widgets": [""]} outputId="c3df3db8-d4a4-453d-ad60-539710c440a1"
tvsModel = tvs.fit(feature_transformer.transform(ms4_train))
data_test = feature_transformer.transform(ms4_test)
results = tvsModel.transform(data_test)
print(evaluator.evaluate(results, {evaluator.metricName: 'areaUnderROC'}))
print(evaluator.evaluate(results, {evaluator.metricName: 'areaUnderPR'}))
# + id="SqGraa_kSqlp" colab={"referenced_widgets": [""]} outputId="1742cafa-0623-459f-81c7-f2bd68962842"
tvsModel.validationMetrics
# + id="wftXEOC6Sqlz" colab={"referenced_widgets": [""]} outputId="9e92f75b-e51e-4e03-a2bc-16b5dec0e0e7"
results = []
for params, metric in zip(tvsModel.getEstimatorParamMaps(), tvsModel.validationMetrics):
parameters = []
for key, paramValue in zip(params.keys(), params.values()):
parameters.append({key.name: paramValue})
results.append((parameters, metric))
sorted(results, key=lambda x: x[1], reverse=True)
# + [markdown] id="9dawkaWMSql0"
# ## Random Forest
# + id="yAvwmZa-Sql1" colab={"referenced_widgets": [""]} outputId="c9769c23-ba47-40e3-a942-6428cc7ccd2f"
ms4 = ms4.withColumn('target', fn.col('target').cast(types.DoubleType()))
ms4_train, ms4_test = ms4.randomSplit([0.7, 0.3], seed=200)
# + id="SqlSNVbrSql2" colab={"referenced_widgets": [""]} outputId="9fa8a708-5d6d-425f-f236-a10213c76f97"
classifier = cl.RandomForestClassifier(numTrees=5, maxDepth=5, labelCol='target')
pipeline = Pipeline(stages=[encoder, featuresCreator, classifier])
model = pipeline.fit(ms4_train)
test = model.transform(ms4_test)
# + id="NdZmVJEASql5" colab={"referenced_widgets": [""]} outputId="63ca2211-fe83-4dda-d183-33e7821fd689"
evaluator = ev.BinaryClassificationEvaluator(labelCol='target')
print(evaluator.evaluate(test, {evaluator.metricName: "areaUnderROC"}))
print(evaluator.evaluate(test, {evaluator.metricName: "areaUnderPR"}))
# + id="A82WxfSRSql7" colab={"referenced_widgets": [""]} outputId="375892c3-1fbe-4b2c-b61a-69c4ead6c0e6"
classifier = cl.DecisionTreeClassifier(maxDepth=5, labelCol='target')
pipeline = Pipeline(stages=[encoder, featuresCreator, classifier])
model = pipeline.fit(ms4_train)
test = model.transform(ms4_test)
evaluator = ev.BinaryClassificationEvaluator(labelCol='target')
print(evaluator.evaluate(test, {evaluator.metricName: "areaUnderROC"}))
print(evaluator.evaluate(test, {evaluator.metricName: "areaUnderPR"}))
| docs/SherLock vs. Moriarty Project/Final_MAIN_PySpark.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 作業 : (Kaggle)鐵達尼生存預測
# ***
# https://www.kaggle.com/c/titanic
# # [作業目標]
# - 試著模仿範例寫法, 在鐵達尼生存預測中, 觀察計數編碼與特徵雜湊的效果
# # [作業重點]
# - 仿造範例, 完成自己挑選特徵的群聚編碼 (In[2], Out[2])
# - 觀察群聚編碼, 搭配邏輯斯回歸, 看看有什麼影響 (In[5], Out[5], In[6], Out[6])
# # 作業1
# * 試著使用鐵達尼號的例子,創立兩種以上的群聚編碼特徵( mean、median、mode、max、min、count 均可 )
# +
# 做完特徵工程前的所有準備 (與前範例相同)
import pandas as pd
import numpy as np
import copy
from sklearn.preprocessing import LabelEncoder, MinMaxScaler
from sklearn.model_selection import cross_val_score
from sklearn.linear_model import LogisticRegression
data_path = 'data/'
df = pd.read_csv(data_path + 'titanic_train.csv')
train_Y = df['Survived']
df = df.drop(['PassengerId', 'Survived'] , axis=1)
df.head(100)
# -
# 取一個類別型欄位, 與一個數值型欄位, 做群聚編碼
"""
Your Code Here
"""
df['Embarked'] = df['Embarked'].fillna('None')
mean_df = df.groupby(['Embarked'])['Fare'].mean().reset_index()
mode_df = df.groupby(['Embarked'])['Fare'].apply(lambda x: x.mode()[0]).reset_index()
median_df = df.groupby(['Embarked'])['Fare'].median().reset_index()
max_df = df.groupby(['Embarked'])['Fare'].max().reset_index()
temp = pd.merge(mean_df, mode_df, how='left', on=['Embarked'])
temp = pd.merge(temp, median_df, how='left', on=['Embarked'])
temp = pd.merge(temp, max_df, how='left', on=['Embarked'])
temp.columns = ['Embarked', 'Embarked_Mean', 'Embarked_Mode', 'Embarked_Median', 'Embarked_Max']
temp.head()
# +
df = pd.merge(df, temp, how='left', on=['Embarked'])
#只取 int64, float64 兩種數值型欄位, 存於 num_features 中
num_features = []
for dtype, feature in zip(df.dtypes, df.columns):
if dtype == 'float64' or dtype == 'int64':
num_features.append(feature)
print(f'{len(num_features)} Numeric Features : {num_features}\n')
# 削減文字型欄位, 只剩數值型欄位
df = df[num_features]
df = df.fillna(-1)
MMEncoder = MinMaxScaler()
df.head()
# -
# # 作業2
# * 將上述的新特徵,合併原有的欄位做生存率預估,結果是否有改善?
# +
# 原始特徵 + 邏輯斯迴歸
"""
Your Code Here
"""
df_minus = df.drop(['Embarked_Mean', 'Embarked_Mode', 'Embarked_Median', 'Embarked_Max'] , axis=1)
# 原始特徵 + 線性迴歸
train_X = MMEncoder.fit_transform(df_minus)
estimator = LogisticRegression()
cross_val_score(estimator, train_X, train_Y, cv=5).mean()
# -
# 新特徵 + 邏輯斯迴歸
"""
Your Code Here
"""
train_X = MMEncoder.fit_transform(df)
cross_val_score(estimator, train_X, train_Y, cv=5).mean()
| homeworks/D027/Day_027_HW.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Improving performance
import pandas as pd
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
# Load the data
df = pd.read_csv('../data/new_titanic_features.csv')
# Create Features and Labels
X = df[['Male', 'Family',
'Pclass2_one', 'Pclass2_two', 'Pclass2_three',
'Embarked_C', 'Embarked_Q', 'Embarked_S',
'Age2', 'Fare3_Fare11to50', 'Fare3_Fare51+', 'Fare3_Fare<=10']]
y = df['Survived']
X.describe()
X.head(15)
from sklearn.model_selection import train_test_split
# Train test split
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=.2, random_state=0)
from sklearn.linear_model import LogisticRegression
model = LogisticRegression()
model.fit(X_train, y_train)
pred_train = model.predict(X_train)
pred_test = model.predict(X_test)
from sklearn.metrics import accuracy_score, confusion_matrix, classification_report
# Check the baseline accuracy against the actual distribution of survival rates:
y_train.value_counts() / len(y_train)
# We see that if we made a model that just predicted that everybody died, it would be correct 62% of the time. So our scoring below (test accuracy of 72%) is somewhat better, but there is room for improvement.
print('Train Accuracy: {:0.3}'.format(accuracy_score(y_train, pred_train)))
print('Test Accuracy: {:0.3}'.format(accuracy_score(y_test, pred_test)))
confusion_matrix(y_test, pred_test)
print(classification_report(y_test, pred_test))
# ## Feature importances (wrong! see exercise 1)
# These are the coefficients that multiply the features in the model, plus the bias.
#
# Larger coefficient magnitude generally means more important feature.
model.coef_
# Can get bias too:
#
model.intercept_
df['Fare'].plot(kind='hist', bins=100)
df.groupby(['Pclass','Survived'])['PassengerId'].count()
coeffs = pd.Series(model.coef_.ravel(), index=X.columns)
coeffs
coeffs.plot(kind='barh')
# Have to be careful interpreting these as importance of the features-- have to look at the distributions and magnitudes of each feature. The Age feature is not yet normalized and has a mean of 29, a max of 80, and a min of -5. Have to normalize to get a better picture of the contributions of these different features.
# ## Cross Validation
from sklearn.model_selection import cross_val_score, ShuffleSplit
# +
cv = ShuffleSplit(n_splits=5, test_size=.4, random_state=0)
scores = cross_val_score(model, X, y, cv=cv)
#scores = cross_val_score(model, X, y, cv=3) #would be exclusive train/test corss validation
scores
# -
'Crossval score: %0.3f +/- %0.3f ' % (scores.mean(), scores.std())
# ## Learning curve
from sklearn.model_selection import learning_curve
tsz = np.linspace(0.1, 1, 10)
train_sizes, train_scores, test_scores = learning_curve(model, X, y, train_sizes=tsz)
fig = plt.figure()
plt.plot(train_sizes, train_scores.mean(axis=1), 'ro-', label="Train Scores")
plt.plot(train_sizes, test_scores.mean(axis=1), 'go-', label="Test Scores")
plt.title('Learning Curve: Logistic Regression')
plt.ylim((0.5, 1.0))
plt.legend()
plt.draw()
plt.show()
# Adding more data appears to decrease model performance, so we need to make changes to the model to make an improvement.
# ### Exercise 1
#
# Try rescaling the Age feature with [`preprocessing.StandardScaler`](http://scikit-learn.org/stable/modules/preprocessing.html) so that it will have comparable size to the other features.
#
# - Do the model prediction change?
# - Does the performance of the model change?
# - Do the feature importances change?
# - How can you explain what you've observed?
from sklearn.preprocessing import StandardScaler
X_train['Age2'].plot(kind='hist', bins = 30)
scaler = StandardScaler()
scaler.fit(X_train[['Age2']])
# +
X_train_scaled = X_train.copy()
X_test_scaled = X_test.copy()
X_train_scaled['Age2'] = scaler.transform(X_train[['Age2']])
X_test_scaled['Age2'] = scaler.transform(X_test[['Age2']])
# -
X_train_scaled['Age2'].plot(kind='hist', bins = 30)
model = LogisticRegression()
model.fit(X_train_scaled, y_train)
pred_train = model.predict(X_train_scaled)
pred_test = model.predict(X_test_scaled)
# +
##not finished, return to check solution
# -
# ### Exercise 2
#
# Experiment with another classifier for example `DecisionTreeClassifier`, `RandomForestClassifier`, `SVC`, `MLPClassifier`, `SGDClassifier` or any other classifier of choice you can find here: http://scikit-learn.org/stable/auto_examples/classification/plot_classifier_comparison.html.
#
# - Train the model on both the scaled data and on the unscaled data
# - Compare the score for the scaled and unscaled data
# - how can you get the features importances for tree based models? Check [here](http://scikit-learn.org/stable/auto_examples/ensemble/plot_forest_importances.html) for some help.
# - Which classifiers are impacted by the age rescale? Why?
# +
from sklearn.ensemble import RandomForestClassifier
model = RandomForestClassifier()
model.fit(X_train, y_train)
pred_train = model.predict(X_train)
pred_test = model.predict(X_test)
coeffs = pd.Series(model.feature_importances_, index=X.columns)
coeffs.plot(kind='barh')
print('Train Accuracy: {:0.3}'.format(accuracy_score(y_train, pred_train)))
print('Test Accuracy: {:0.3}'.format(accuracy_score(y_test, pred_test)))
# -
# ### Exercise 3
#
# Pick your preferred classifier from Exercise 2 and search for the best hyperparameters. You can read about hyperparameter search [here](http://scikit-learn.org/stable/modules/grid_search.html)
#
# - Decide the range of hyperparameters you intend to explore
# - Try using [`GridSearchCV`](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.GridSearchCV.html#sklearn.model_selection.GridSearchCV) to perform brute force search
# - Try using [`RandomizedSearchCV`](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.RandomizedSearchCV.html#sklearn.model_selection.RandomizedSearchCV) for a random search
# - Once you've chosen the best classifier and the best hyperparameter set, redo the learning curve.
# Do you need more data or a better model?
| day_2/Lab_08_ML Improving performance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Variables, cadenas y listas
#
# ### 1. Grados celsius a fahrenheint. Dada una temperatura en grados celsius, determina su equivalente en grados Fahrenheint
# $$ F = 9/5C + 32 $$
print(F)
type(C)
type(F)
C = 106 # Grados centigrados
F = 9/5*C+32 # Grados fahrenheint
print(C, "°C equivale a", F,"°F") #Imprimir valores
print(str(C)+"° equivale a "+ str(F)) #Concatenar
centi = str(C)
type(centi)
int(centi)
C - 5
str(C) - 5
# ### 2. Convertir listas a cadenas y al revés
#
# Input: "Python es mi clase favorita"
#
# Output: ["Python", "es", "mi", "clase", "favorita"]
#
# https://docs.python.org/3/library/stdtypes.html
frase = "Python es mi clase favorita"
frase.split("s") #separar
lista_1 = frase.split()
type(lista_1)
lista_1
espacio = "+"
espacio.join(lista_1) #unir
# ### 3. Buscar palabras. Busca todas las repeticiones de la palabra "energía" en el siguiente texto:
# > Desde la crisis del petróleo en 1973, fue claro tanto para los países desarrollados como los que se encontraban en vías de desarrollo, el de realizar grandes esfuerzos científicos y tecnológicos encaminados principalmente en dos vertientes: el ahorro y uso eficiente de energía, y la diversificación de las fuentes de la energía, entre las que se encuentran, las energías renovables. En este contexto, la Universidad Nacional Autónoma de México (UNAM) tomando conciencia de estos esfuerzos, tuvo la visión de formar profesionales de alto nivel en las dos vertientes señaladas, por lo que en el año de 1984 creó la Maestría en Energética en la Facultad de Ingeniería de Ciudad Universitaria. Tres años más tarde se abrió la Maestría en Energía Solar (con dos opciones: Fototérmica y Fotovoltaica) en el antiguo Centro de Investigación en Energía (antes Laboratorio de Energía Solar) ahora Instituto de Energías Renovables en Temixco, Morelos. Desde sus inicios hasta 1998 se inscribieron en este programa un total de 82 alumnos de maestría, obteniéndose una eficiencia terminal del 55%. Posteriormente en 1999 y por acuerdo del Rector <NAME>, se adecuaron todos los posgrados de la UNAM, de tal forma que en este nuevo esquema, los posgrados existentes dejaron de pertenecer a una entidad determinada y se crearon nuevos programas en donde participan todas las entidades involucradas con un tema general (pero incluyendo campos de conocimiento más específicos). En este nuevo programa de posgrado se logró integrar a profesores de facultades e investigadores de centros e institutos, incluyendo el aprovechamiento de la infraestructura de cada entidad académica participante. De esta forma, se creó en ese mismo año 1999 el Programa de Maestría en Ingeniería (Energía), único en el País, con una orientación hacia la investigación científica y científica-práctica.
texto = "Desde la crisis del petróleo en 1973, fue claro tanto para los países desarrollados como los que se encontraban en vías de desarrollo, el de realizar grandes esfuerzos científicos y tecnológicos encaminados principalmente en dos vertientes: el ahorro y uso eficiente de energía, y la diversificación de las fuentes de la energía, entre las que se encuentran, las energías renovables. En este contexto, la Universidad Nacional Autónoma de México (UNAM) tomando conciencia de estos esfuerzos, tuvo la visión de formar profesionales de alto nivel en las dos vertientes señaladas, por lo que en el año de 1984 creó la Maestría en Energética en la Facultad de Ingeniería de Ciudad Universitaria. Tres años más tarde se abrió la Maestría en Energía Solar (con dos opciones: Fototérmica y Fotovoltaica) en el antiguo Centro de Investigación en Energía (antes Laboratorio de Energía Solar) ahora Instituto de Energías Renovables en Temixco, Morelos. Desde sus inicios hasta 1998 se inscribieron en este programa un total de 82 alumnos de maestría, obteniéndose una eficiencia terminal del 55%. Posteriormente en 1999 y por acuerdo del Rector <NAME>, se adecuaron todos los posgrados de la UNAM, de tal forma que en este nuevo esquema, los posgrados existentes dejaron de pertenecer a una entidad determinada y se crearon nuevos programas en donde participan todas las entidades involucradas con un tema general (pero incluyendo campos de conocimiento más específicos). En este nuevo programa de posgrado se logró integrar a profesores de facultades e investigadores de centros e institutos, incluyendo el aprovechamiento de la infraestructura de cada entidad académica participante. De esta forma, se creó en ese mismo año 1999 el Programa de Maestría en Ingeniería (Energía), único en el País, con una orientación hacia la investigación científica y científica-práctica."
palabra = "Energía"
texto[735:746]
texto.find(palabra)
texto.count(palabra)
palabra.lower()
texto_tmp = texto.lower()
palabra_tmp = palabra.lower()
texto_tmp.count(palabra_tmp)
# +
texto = "Desde la crisis del petróleo en 1973, fue claro tanto para los países desarrollados como los que se encontraban en vías de desarrollo, el de realizar grandes esfuerzos científicos y tecnológicos encaminados principalmente en dos vertientes: el ahorro y uso eficiente de energía, y la diversificación de las fuentes de la energía, entre las que se encuentran, las energías renovables. En este contexto, la Universidad Nacional Autónoma de México (UNAM) tomando conciencia de estos esfuerzos, tuvo la visión de formar profesionales de alto nivel en las dos vertientes señaladas, por lo que en el año de 1984 creó la Maestría en Energética en la Facultad de Ingeniería de Ciudad Universitaria. Tres años más tarde se abrió la Maestría en Energía Solar (con dos opciones: Fototérmica y Fotovoltaica) en el antiguo Centro de Investigación en Energía (antes Laboratorio de Energía Solar) ahora Instituto de Energías Renovables en Temixco, Morelos. Desde sus inicios hasta 1998 se inscribieron en este programa un total de 82 alumnos de maestría, obteniéndose una eficiencia terminal del 55%. Posteriormente en 1999 y por acuerdo del Rector <NAME>, se adecuaron todos los posgrados de la UNAM, de tal forma que en este nuevo esquema, los posgrados existentes dejaron de pertenecer a una entidad determinada y se crearon nuevos programas en donde participan todas las entidades involucradas con un tema general (pero incluyendo campos de conocimiento más específicos). En este nuevo programa de posgrado se logró integrar a profesores de facultades e investigadores de centros e institutos, incluyendo el aprovechamiento de la infraestructura de cada entidad académica participante. De esta forma, se creó en ese mismo año 1999 el Programa de Maestría en Ingeniería (Energía), único en el País, con una orientación hacia la investigación científica y científica-práctica."
palabra = "Energía"
texto_tmp = texto.lower()
palabra_tmp = palabra.lower()
count = texto_tmp.count(palabra_tmp)
count
# +
texto = "Desde la crisis del petróleo en 1973, fue claro tanto para los países desarrollados como los que se encontraban en vías de desarrollo, el de realizar grandes esfuerzos científicos y tecnológicos encaminados principalmente en dos vertientes: el ahorro y uso eficiente de energía, y la diversificación de las fuentes de la energía, entre las que se encuentran, las energías renovables. En este contexto, la Universidad Nacional Autónoma de México (UNAM) tomando conciencia de estos esfuerzos, tuvo la visión de formar profesionales de alto nivel en las dos vertientes señaladas, por lo que en el año de 1984 creó la Maestría en Energética en la Facultad de Ingeniería de Ciudad Universitaria. Tres años más tarde se abrió la Maestría en Energía Solar (con dos opciones: Fototérmica y Fotovoltaica) en el antiguo Centro de Investigación en Energía (antes Laboratorio de Energía Solar) ahora Instituto de Energías Renovables en Temixco, Morelos. Desde sus inicios hasta 1998 se inscribieron en este programa un total de 82 alumnos de maestría, obteniéndose una eficiencia terminal del 55%. Posteriormente en 1999 y por acuerdo del Rector <NAME>, se adecuaron todos los posgrados de la UNAM, de tal forma que en este nuevo esquema, los posgrados existentes dejaron de pertenecer a una entidad determinada y se crearon nuevos programas en donde participan todas las entidades involucradas con un tema general (pero incluyendo campos de conocimiento más específicos). En este nuevo programa de posgrado se logró integrar a profesores de facultades e investigadores de centros e institutos, incluyendo el aprovechamiento de la infraestructura de cada entidad académica participante. De esta forma, se creó en ese mismo año 1999 el Programa de Maestría en Ingeniería (Energía), único en el País, con una orientación hacia la investigación científica y científica-práctica."
texto.lower().count("Energía".lower())
# -
# ### 4. Ordenar números
# Crea un programa que lea tres enteros y los muestre en orden (de menor a mayor).
# Por ejemplo, para
# ```python
# a=14
# b=3
# c=54
# ```
# deberá mostrar
#
# Los números ordenados son: 3 14 54
a = 1400
b = 300
c = 0.54
print("Los numeros ordenados son",b, a, c)
# +
#min() = mn #No debe hacerse, palabra reservada
# +
mn = min(a,b,c)
mx = max(a,b,c)
md = a+b+c-mn-mx
print("Los numeros ordenados son",mn,md,mx)
# -
# ### 5. Elementos en una lista
# Agrega el elemento 7000 después de 6000 en la siguiente lista:
# ```python
# list1 = [10, 20, [300, 400, [5000, 6000], 500], 30, 40]
#
# list1 = [10, 20, [300, 400, [5000, 6000,7000], 500], 30, 40]
# ```
#
# __Las listas siempre empiezan en cero__
list1 = [10, 20, [300, 400, [5000, 6000], 500], 30, 40]
list1[0]
list1[1]
list1[2]
list1[2][0]
list1[2][0].append(7000)
list1
# ### Intercambiar variables
a = "variable 1"
b = "variable 2"
print(a,b)
# +
tmp = a
a = b
b = tmp
print(a,b)
# -
a = "variable 1"
b = "variable 2"
c = 100
# +
a, b = b, a
print(a, b)
# -
print(a,b,c)
# +
c,b,a = a,c,b
print(a,b,c)
# -
| notebooks/dia_02/dia02_ejercicios.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Module 2
#
# ## Video 7: Generating Lists
# **Python for the Energy Industry**
#
# It's common to make a list by conditionally appending inside a loop. See this example of making a list of even numbers:
# +
evens = []
for i in range(11):
if i % 2 == 0:
evens.append(i)
print(evens)
# -
# Here's another example, of finding the words in a list beginning with A:
#
# *Note: strings can be treated in many ways as a list of characters - see below how we access the first letter of word as word[0]*
# +
words = ['AARDVARK','APPLE','ARROW','BARN','CACTUS']
a_words = []
for word in words:
if word[0] == 'A':
a_words.append(word)
print(a_words)
# -
# This procedure is so common that there is a Python shorthand for it, called the 'list comprehension' which takes the form:
#
# ``` new_list = [item for item in old_list if condition] ```
#
# Here's how this looks for our even numbers example:
# +
evens = [i for i in range(11) if i % 2 == 0]
print(evens)
# -
# You can also apply a function onto the item before storing it in the list. We will cover functions in more detail later, but here is a simple example:
# +
squares = [i**2 for i in range(10) if i % 2 == 0]
print(squares)
# -
# ### Exercise
#
# Try to write the list comprehension corresponding to our A words example.
| docs/examples/academy/7. Generating Lists.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### MNIST
import numpy as np
from sklearn.datasets import fetch_mldata
mnist = fetch_mldata('MNIST original', data_home='../data')
mnist
X, y = mnist['data'], mnist['target']
X.shape
y.shape
X_train = np.array(X[:60000], dtype=float)
y_train = np.array(y[:60000], dtype=float)
X_test = np.array(X[60000:], dtype=float)
y_test = np.array(y[60000:], dtype=float)
X_train.shape
# ### 使用KNN
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier()
# %time knn.fit(X_train, y_train)
# %time knn.score(X_test, y_test)
# ### PCA降维度
from sklearn.decomposition import PCA
pca = PCA(0.9)
pca.fit(X_train)
X_train_reduction = pca.transform(X_train)
X_test_reduction = pca.transform(X_test)
X_train_reduction.shape
knn = KNeighborsClassifier()
# %time knn.fit(X_train_reduction, y_train)
# %time knn.score(X_test_reduction, y_test)
| ml/pca/PCA-MNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Clustering animals with attributes
# In this notebook, we'll get a feel for clustering by applying <font color="magenta">**k-means**</font> and <font color="magenta">**average linkage**</font> to cluster a collection of animals.
#
# The data set we'll work with is called `Animals with Attributes`. It contains information about 50 animals. For each, it has 85 real-valued features that capture various properties of the animal: where it lives, what it eats, and so on. You can download the data set from:
#
# http://attributes.kyb.tuebingen.mpg.de
#
# (Under "Downloads", choose the "base package".) Make sure the files `'classes.txt'` and `'predicate-matrix-continuous.txt'` are in the same directory as this notebook before you begin.
# ## 1. Load in the data set
# Standard includes as well as some modules for k-means and hierarchical clustering.
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
import scipy.cluster.hierarchy as sch
from sklearn.cluster import KMeans
# Load in the data set. The file `'classes.txt'` contains the names of the 50 animals. The file `'predicate-matrix-continuous.txt'` contains the data itself: 85 attributes per animal.
#
# Load in animal names
#
f = open('Animals_with_Attributes/classes.txt')
lines = f.readlines()
names = []
for line in lines:
(key,val) = line.split()
names.append(val)
f.close()
#
# Load attribute matrix
#
x = np.loadtxt('Animals_with_Attributes/predicate-matrix-continuous.txt')
len(names)
# ## 2. K-means clustering
# We now run Lloyd's algorithm to obtain a flat clustering of the data. In the code below, we ask for k=15 clusters, but you should experiment with other choices.
#
# We ask for random initialization, which means that different runs of the algorithm will potentially have different outcomes. It is worth running the algorithm several times to see how the results change.
# +
#
# Run Lloyd's algorithm
#
k = 10 # number of clusters
kmeans = KMeans(n_clusters=k, init='k-means++')
kmeans.fit(x)
#
# Get a list of the animals in each cluster
#
clusters = []
for j in range(0,k):
clusters.append([])
for i in range(0,50):
clusters[kmeans.labels_[i]].append(names[i])
# print(kmeans.labels_[i])
#
# Print out clusters
#
for j in range(0,k):
print (j+1, clusters[j])
# -
kmeans.inertia_
# +
#
# Run Lloyd's algorithm
#
k = 10 # number of clusters
#init='random'
kmeans = KMeans(n_clusters=k, init='random')
kmeans.fit(x)
#
# Get a list of the animals in each cluster
#
clusters_random = []
for j in range(0,k):
clusters_random.append([])
for i in range(0,50):
clusters_random[kmeans.labels_[i]].append(names[i])
# print(kmeans.labels_[i])
#
# Print out clusters
#
for j in range(0,k):
print (j+1, clusters_random[j])
# -
kmeans.inertia_
# ## 3. Hierarchical clustering
# We use the built-in hierarchical clustering module of `scipy` to apply Ward's method to our data. The resulting tree is often called a *dendrogram*, and the module also has a routine that can display one of these.
#
# Adjust figure size to accommodate the full tree
#
from pylab import rcParams
rcParams['figure.figsize'] = 5, 10
#
# Do hierarchical clustering using Ward's method
#
z = sch.linkage(x, method='ward')
#
# Display dendrogram
#
info = sch.dendrogram(z, orientation='left', labels=names, leaf_font_size=12)
# ## 4. Questions to ponder
# Here are some things to think about:
#
# <font color="magenta">Multiple runs of Lloyd's algorithm.</font>
# Lloyd's algorithm potentially returns a different solution each time it is run. Is there any reason to run it more than once? For instance, is there a sensible way of combining the information from several runs, of interpreting the similarities and differences?
#
# <font color="magenta">Sensitivity to the choice of features.</font>
# Both clustering methods are highly sensitive to the choice of features. How would you feel if the results changed dramatically when just one or two features were dropped?
#
# <font color="magenta">Criteria for success.</font>
# This is clearly an application in which we are hoping that clustering will discover 'natural groups' in the data. To what extent do the algorithms succeed at this? Are the clusters mostly reasonable? Can we, in general, hope that tha clustering will perfectly capture what we want? Under what conditions would we be pleased with the clustering?
#For 1st run
d1=dict()
for j in range(0,k):
d1[j+1]=clusters[j]
#For second run
d2=dict()
for j in range(0,k):
d2[j+1]=clusters_random[j]
#Finiding similarities and differences
same = set()
different = dict()
for key in set(d1.keys()) & set(d2.keys()):
if d1[key] == d2[key]:
same.add(key)
else:
if key not in different:
different[key] = []
different[key].extend([d1[key], d2[key]])
same
different
# #### Never do they have same clusters
S = np.array([[4.0, 2.0, -3.0],
[2.0, 9.0, 0],
[-3.0, 0, 9.0]])
def correlation_from_covariance(covariance):
v = np.sqrt(np.diag(covariance))
outer_v = np.outer(v, v)
correlation = covariance / outer_v
correlation[covariance == 0] = 0
print(outer_v)
return correlation
correlation = correlation_from_covariance(S)
correlation
u = np.array([1,1,0])
ut = np.transpose(u)
u_norm = np.multiply(np.linalg.norm(u),u)
ut_norm = np.multiply(np.linalg.norm(ut),ut)
S
1/2*np.matmul(ut,np.matmul(S,u))
## Priblem 11 (a,b)
a = np.array([1/2,1/2,1/2,1/2]); b = np.array([-1/2,1/2,-1/2,1/2]); X = np.array([2,4,2,6])
np.matmul(np.transpose(a),X) ; np.matmul(np.transpose(b),X)
a*np.matmul(np.transpose(a),X) + b*np.matmul(np.transpose(b),X)
# +
# Problem 12 (a,b)
M = np.array([[5., 1.],[1., 5.]]) ; u1 = np.array([1,1])
#(a) sqrt((6^2 + 6^2)/sqrt(2) = ... Basically we have to find the length of the Mu1 and divide by sqrt(2)
Mu1 = np.matmul(M,u1)
Mu1
#(b) sqrt((4^2 + 4^2))/sqrt(2) = 4
u2 = np.array([1,-1]) ; Mu2 = np.matmul(M,u2)
Mu2
# -
### Problem 14
## (a)
M1 = np.array([[5.0, -3.0, 0.0],
[-3.0, 5.0, 0],
[0.0, 0.0, 4.0]])
u3 = np.array([1/1.73,1/1.73,1/1.73])
np.matmul(np.transpose(u3),np.matmul(M1,u3))
M1
np.matmul(M1,np.array([0,1/1.41,1/1.41]))
np.linalg.eigvals(M1)
### problem 14(b)
w, v = np.linalg.eig(M1)
w #### Eigen Values
## where v[:0] and v[:,2] are eigen vectors
v[:,0]
# Problem 14(d)
X = np.array([1.41,-4.24,2])
np.matmul(X,v[:,0]) #### First eigen vector
np.matmul(X,v[:,2]) ### Second eigen vector
(np.matmul(X,v[:,0]))*v[:,0]
(np.matmul(X,v[:,2]))*v[:,2]
x = np.linalg.norm((np.matmul(X,v[:,0]))*v[:,0]) + np.linalg.norm((np.matmul(X,v[:,2]))*v[:,2])
x_hat = np.sqrt(np.square(1.41) + np.square(-4.24) + np.sqrt(2))
x
x - x_hat
np.linalg.norm((np.matmul(X,v[:,0]))*v[:,0])
A1 = ((np.matmul(X,v[:,2]))*v[:,2])
B1 = ((np.matmul(X,v[:,0]))*v[:,0])
# np.matmul(B1,v[:,0])
# np.matmul(A1,v[:,2]
A1*v[:,2]
B1*v[:,0]
np.matmul(v[:,0],X)*v[:,0] + np.matmul(v[:,1],X)*v[:,1]
np.sqrt(np.square(1.41 - 1.41) + np.square(-4.24 + 4.24) + np.square(0-2))
X1 = np.array([[1, -2.0],
[-2.0, 1.0]])
X1
np.matmul(np.array([-1/1.41,1/1.41]),np.array([1,2]))
| Assignment 8/animals-clustering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Convolutional Neural Networks
#
#
# In this notebook we are going to explore the [CIFAR-10](https://www.cs.toronto.edu/~kriz/cifar.html) dataset (you don't need to download this dataset, we are going to use keras to download this dataset). This is a great dataset to train models for visual recognition and to start to build some models in Convolutional Neural Networks (CNN). This dataset consists of 60,000 32x32 colour images in 10 classes, with 6,000 images per class. There are 50,000 training images and 10,000 test images
#
# As CNN's requires high-computational effort, we are going to use a reduced version of this training dataset. Given our time and computational resources restrictions, we are going to select 3 categories (airplane, horse and truck).
#
# In this notebook, we are going to build two different models in order to classify the objects. First, we are going to build Shallow Neural Network based just in a few Fully-Connected Layers (aka Multi-layer Perceptron) and we are going to understand why is not feasible to classify images with such networks. Then, we are going to build a CNN network to perform the same task and evaluate its performance.
#
# Again, in order to have a clean notebook, some functions are implemented in the file *utils.py* (e.g., plot_loss_and_accuracy).
#
# Summary:
# - [Downloading CIFAR-10 Dataset](#cifar)
# - [Data Pre-processing](#reduce)
# - [Reducing the Dataset](#red)
# - [Normalising the Dataset](#normalise)
# - [One-hot Encoding](#onehot)
# - [Building the Shallow Neural Network](#shallow)
# - [Training the Model](#train_shallow)
# - [Prediction and Performance Analysis](#performance_sh)
# - [Building the Convolutional Neural Network](#cnn)
# - [Training the Model](#train_cnn)
# - [Prediction and Performance Analysis](#performance_cnn)
#
# +
# Standard libraries
import numpy as np # written in C, is faster and robust library for numerical and matrix operations
import pandas as pd # data manipulation library, it is widely used for data analysis and relies on numpy library.
import matplotlib.pyplot as plt # for plotting
import seaborn as sns # Plot nicely =) . Importing seaborn modifies the default matplotlib color schemes and plot
# styles to improve readability and aesthetics.
# Auxiliar functions
from utils import *
# the following to lines will tell to the python kernel to always update the kernel for every utils.py
# modification, without the need of restarting the kernel.
# %load_ext autoreload
# %autoreload 2
# using the 'inline' backend, your matplotlib graphs will be included in your notebook, next to the code
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
# -
# ## Downloading CIFAR-10 Dataset
# <a id='cifar'></a>
#
# Keras provides several [datasets](https://keras.io/datasets/) for experimentation, this makes it easy to try new network architectures. In order to download the CIFAR-10 dataset, we need to import the library "[cifar10](https://keras.io/datasets/#cifar100-small-image-classification)" and call the method *load_data()".
# +
from keras.datasets import cifar10 # Implements the methods to dowload CIFAR-10 dataset
(x_train, y_train), (x_test, y_test) = cifar10.load_data() #this will download the dataset
# by defaul, the dataset was split in 50,000 images for training and 10,000 images for testing
# we are going to use this configuration
y_train = y_train.ravel() # Return a contiguous flattened y_train
y_test = y_test.ravel() #Return a contiguous flattened y_test
# -
# Let's visualise how the images looks like. To plot the images we are going to use the function **plot_images** (see *utils.py*)
# +
# from https://www.cs.toronto.edu/~kriz/cifar.html we can grab the class names
# 0 1 2 3 4 5 6 7 8 9
class_name = np.array(
['airplane', 'automobile','bird','cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck'])
#
plot_samples(x_train, y_train, class_name)
# -
# ## Data Pre-processing
# <a id='reduce'></a>
#
# As CNN's requires high-computational effort, we are going to use a reduced training dataset. Given our time and computational resources restrictions, we are going to select 3 categories (airplane, horse and truck) and for each category and select in total 1500 images.
#
# Once obtained the reduced version, we are going to normalise the images and generate the one-hot enconding representation of the labels.
# ### Reducing the Dataset
# <a id='red'></a>
# +
# Lets select just 3 classes to make this tutorial feasible
selected_idx = np.array([0, 7, 9])
n_images = 1500
y_train_idx = np.isin(y_train, selected_idx)
y_test_idx = np.isin(y_test, selected_idx)
y_train_red = y_train[y_train_idx][:n_images]
x_train_red = x_train[y_train_idx][:n_images]
y_test_red = y_test[y_test_idx][:n_images]
x_test_red = x_test[y_test_idx][:n_images]
# replacing the labels 0, 7 and 9 to 0, 1, 2 repectively.
y_train_red[y_train_red == selected_idx[0]] = 0
y_train_red[y_train_red == selected_idx[1]] = 1
y_train_red[y_train_red == selected_idx[2]] = 2
y_test_red[y_test_red == selected_idx[0]] = 0
y_test_red[y_test_red == selected_idx[1]] = 1
y_test_red[y_test_red == selected_idx[2]] = 2
# -
y_test_red[:4]
# visulising the images in the reduced dataset
plot_samples(x_train_red, y_train_red, class_name[selected_idx])
# **Question 1**: Is the reduced dataset imbalanced?
# **Question 2**: As you can see, the images have low resolution (32x32x3), how this can affect the model?
# ### Normalising the Dataset
# <a id='normalise'></a>
#
# Here we are going to normalise the dataset. In this task, we are going to divide each image by 255.0, as the images are represented as 'uint8' and we know that the range is from 0 to 255. By doing so, the range of the images will be between 0 and 1.
# Normalising the
x_train_red = x_train_red.astype('float32')
x_test_red = x_test_red.astype('float32')
x_train_red /= 255.0
x_test_red /= 255.0
# ### One-hot Encoding
# <a id='onehot'></a>
#
# The labels are encoded as integers (0, 1 and 2), as we are going to use a *softmax layer* as output for our models we need to convert the labels as binary matrix. For example, the label 0 (considering that we have just 3 classes) can be represented as [1 0 0], which is the class 0.
#
# One-hot enconding together with the sofmax function will give us an interesting interpretation of the output as a probability distribution over the classes.
#
# For this task, are going to use the function *[to_categorical](https://keras.io/utils/)*, which converts a class vector (integers) to binary class matrix.
# +
y_train_oh = keras.utils.to_categorical(y_train_red)
y_test_oh = keras.utils.to_categorical(y_test_red)
print('Label: ',y_train_red[0], ' one-hot: ', y_train_oh[0])
print('Label: ',y_train_red[810], ' one-hot: ', y_train_oh[810])
print('Label: ',y_test_red[20], ' one-hot: ', y_test_oh[20])
# -
# ## Building the Shallow Neural Network
# <a id='shallow'></a>
#
# Here we are going to build a Shallow Neural Network with 2 Fully Connected layers and one output layer. Basically, we are implemting a Multi-Layer Perceptron classifier.
#
# To build the model, we are going use the following components from Keras:
#
# - [Sequencial](https://keras.io/models/sequential/): allows us to create models layer-by-layer.
# - [Dense](https://keras.io/layers/core/): provides a regular fully-connected layer
# - [Dropout](https://keras.io/layers/core/#dropout): provides dropout regularisation
#
# Basically, we are going to define the sequence of our model by using _Sequential()_, which include the layers:
#
# ```python
# model = Sequential()
# model.add(Dense(...))
# ...
# ```
# once created the model we can configure the model for training by using the method [compile](https://keras.io/models/model/). Here we need to define the [loss](https://keras.io/losses/) function (mean squared error, categorical cross entropy, among others.), the [optimizer](https://keras.io/optimizers/) (Stochastic gradient descent, RMSprop, adam, among others) and the [metric](https://keras.io/metrics/) to define the evaluation metric to be used to evaluate the performance of the model in the training step, as follows:
#
# ```python
# model.compile(loss = "...",
# optimizer = "...")
# ```
#
# Also, we have the option to see a summary representation of the model by using thebfunction [summary](https://keras.io/models/about-keras-models/#about-keras-models). This function summarise the model and tell us the number of parameters that we need to tune.
from keras.models import Sequential # implements sequential function
from keras.layers import Dense # implements the fully connected layer
from keras.layers import Dropout # implements Dropout regularisation
from keras.layers import Flatten # implements Flatten function
# +
mlp = Sequential()
# Flatten will reshape the input an 1D array with dimension equal to 32 x 32 x 3 (3072)
# each pixel is an input for this model.
mlp.add(Flatten(input_shape=x_train_red.shape[1:])) #x_train.shape[1:] returns the shape
# First layer with 1024 neurons and relu as activation function
mlp.add(Dense(1024, activation='relu'))
mlp.add(Dropout(0.7)) # regularization with 70% of keep probability
# Second layer with 1024 neurons and relu as activation function
mlp.add(Dense(1024, activation='relu'))
mlp.add(Dropout(0.7))# regularization with 70% of keep probability
# Output layer with 3 neurons and sofmax as activation function
mlp.add(Dense(y_test_oh.shape[1], activation='softmax'))
# -
# Summarising the model
mlp.summary()
# +
# Compile:
# Optimiser: rmsprop
# Loss: categorical_crossentropy, as our problem is multi-label classification
# Metric: accuracy
mlp.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
# -
# ### Training the Model
# <a id=train_shallow></a>
#
# Once defined the model, we need to train it by using the function [fit](https://keras.io/models/model/). This function performs the optmisation step. Hence, we can define the following parameters such as:
#
# - batch size: defines the number of samples that will be propagated through the network
# - epochs: defines the number of times in which all the training set (x_train_scaled) are used once to update the weights
# - validation split: defines the percentage of training data to be used for validation
# - among others (click [here](https://keras.io/models/model/) for more information)
#
# This function return the _history_ of the training, that can be used for further performance analysis.
# training the model (this will take a few minutes)
history = mlp.fit(x_train_red,
y_train_oh,
batch_size = 256,
epochs = 100,
validation_split = 0.2,
verbose = 1)
# ### Prediction and Performance Analysis
# <a id='performance_sh'></a>
#
# Here we plot the 'loss' and the 'Accuracy' from the training step.
plot_loss_and_accuracy_am2(history=history)
# Let's evaluate the performance of this model under unseen data (x_test)
loss_value_mlp, acc_value_mlp = mlp.evaluate(x_test_red, y_test_oh, verbose=0)
print('Loss value: ', loss_value_mlp)
print('Acurracy value: ', acc_value_mlp)
# ## Building the Convolutional Neural Network
# <a id='cnn'></a>
#
# Here we are going to build a Convolutional Neural Network (CNN) for image classification. Given the time and computational resources limitations, we are going to build a very simple CNN, however, more complex and deep CNN's architectures such as VGG, Inception and ResNet are the state of the art in computer vision and they superpass the human performance in image classification tasks.
#
# To build the model, we are going use the following components from Keras:
#
# - [Sequencial](https://keras.io/models/sequential/): allows us to create models layer-by-layer.
# - [Dense](https://keras.io/layers/core/): provides a regular fully-connected layer
# - [Dropout](https://keras.io/layers/core/#dropout): provides dropout regularisation
# - [Conv2D](https://keras.io/layers/convolutional/): implement 2D convolution function
# - [BatchNormalization](https://keras.io/layers/normalization/): normalize the activations of the previous layer at each batch
# - [MaxPooling2D](https://keras.io/layers/pooling/): provides pooling operation for spatial data
#
# Basically, we are going to define the sequence of our model by using _Sequential()_, which include the layers:
#
# ```python
# model = Sequential()
# model.add(Conv2D(...))
# ...
# ```
# once created the model the training configuration is the same as [before](#shallow):
#
# ```python
# model.compile(loss = "...",
# optimizer = "...")
# ```
from keras.models import Sequential
from keras.layers import Dense, Flatten, Activation
from keras.layers import Dropout, Conv2D, MaxPooling2D, BatchNormalization
# +
model_cnn = Sequential()
# First layer:
# 2D convolution:
# Depth: 32
# Kernel shape: 3 x 3
# Stride: 1 (default)
# Activation layer: relu
# Padding: valid
# Input shape: 32 x 32 x 3 (3D representation, not Flatten as MLP)
# as you can see now the input is an image and not an flattened array
model_cnn.add(Conv2D(32, (3, 3), padding='valid', activation = 'relu',
input_shape=x_train_red.shape[1:]))
model_cnn.add(BatchNormalization())
model_cnn.add(MaxPooling2D(pool_size=(5,5))) # max pooling with kernel size 5x5
model_cnn.add(Dropout(0.7)) # 70% of keep probability
# Second layer:
# 2D convolution:
# Depth: 64
# Kernel shape: 3 x 3
# Stride: 1 (default)
# Activation layer: relu
# Padding: valid
model_cnn.add(Conv2D(64, (3, 3), padding='valid', activation = 'relu'))
model_cnn.add(BatchNormalization())
model_cnn.add(MaxPooling2D(pool_size=(2,2)))
model_cnn.add(Dropout(0.7))
# Flatten the output from the second layer to become the input of the Fully-connected
# layer (flattened representation as MLP)
model_cnn.add(Flatten())
# First fully-connected layer with 128 neurons and relu as activation function
model_cnn.add(Dense(128, activation = 'relu'))
# Output layer with 3 neurons and sofmax as activation function
model_cnn.add(Dense(y_test_oh.shape[1], activation='softmax'))
# -
# Summarising the model
model_cnn.summary()
# As you can see, the CNN model (53,059 parameters) has less parameters than the MLP model (4,199,427 parameters). So this model is less prone to overfit.
# +
# Compile:
# Optimiser: adam
# Loss: categorical_crossentropy, as our problem is multi-label classification
# Metric: accuracy
model_cnn.compile(optimizer='adam',
loss='categorical_crossentropy',
metrics=['accuracy'])
# -
# ### Training the Model
# <a id=train_cnn></a>
# this will take a few minutes
history_cnn = model_cnn.fit(x_train_red,
y_train_oh,
batch_size = 256,
epochs = 100,
validation_split = 0.2,
verbose = 1)
# ### Prediction and Performance Analysis
# <a id='performance_cnn'></a>
plot_loss_and_accuracy_am2(history=history_cnn)
# Let's evaluate the performance of this model under unseen data (x_test)
# +
model_cnn.evaluate(x_test_red,y_test_oh)
loss_value_cnn, acc_value_cnn = model_cnn.evaluate(x_test_red, y_test_oh, verbose=0)
print('Loss value: ', loss_value_cnn)
print('Acurracy value: ', acc_value_cnn)
# -
# **Task**: Discuss CNN and MLP results.
# **Your Turn**: Now we changed our mind, we found that detecting airplanes, horses and trucks is a bit boring :(. We would like to detect whether an image has a bird, a dog or a ship =)
#
# Implement a CNN to classify the images of the new reduced dataset.
# **Creating the dataset**
# +
# Lets select just 3 classes to make this tutorial feasible
selected_idx = np.array([2, 5, 8])
n_images = 1500
y_train_idx = np.isin(y_train, selected_idx)
y_test_idx = np.isin(y_test, selected_idx)
y_train_new = y_train[y_train_idx][:n_images]
x_train_new = x_train[y_train_idx][:n_images]
y_test_new = y_test[y_test_idx][:n_images]
x_test_new = x_test[y_test_idx][:n_images]
# replacing the labels 0, 7 and 9 to 0, 1, 2 repectively.
y_train_new[y_train_new == selected_idx[0]] = 0
y_train_new[y_train_new == selected_idx[1]] = 1
y_train_new[y_train_new == selected_idx[2]] = 2
y_test_new[y_test_new == selected_idx[0]] = 0
y_test_new[y_test_new == selected_idx[1]] = 1
y_test_new[y_test_new == selected_idx[2]] = 2
# visulising the images in the reduced dataset
plot_samples(x_train_new, y_train_new, class_name[selected_idx])
# -
# **Pre-processing the new dataset**
# +
# normalising the data
x_train_new = x_train_new.astype('float32')
x_test_new = x_test_new.astype('float32')
x_train_new /= 255.0
x_test_new /= 255.0
# creating the one-hot representation
y_train_oh_n = keras.utils.to_categorical(y_train_new)
y_test_oh_n = keras.utils.to_categorical(y_test_new)
print('Label: ',y_train_new[0], ' one-hot: ', y_train_oh_n[0])
print('Label: ',y_train_new[810], ' one-hot: ', y_train_oh_n[810])
print('Label: ',y_test_new[20], ' one-hot: ', y_test_oh_n[20])
# -
# **Step 1**: Create the CNN Model.
#
# For example, you can try (Danger, <NAME>! This model can overfits):
#
# ```python
# model_cnn_new = Sequential()
#
# model_cnn_new.add(Conv2D(32, (3, 3), padding='valid', activation = 'relu',
# input_shape=x_train_new.shape[1:]))
# model_cnn_new.add(BatchNormalization())
# model_cnn_new.add(MaxPooling2D(pool_size=(2,2)))
# model_cnn_new.add(Dropout(0.7))
#
# # You can stack several convolution layers before apply BatchNormalization, MaxPooling2D
# # and Dropout
# model_cnn_new.add(Conv2D(32, (3, 3), padding='valid', activation = 'relu',
# input_shape=x_train_new.shape[1:]))
# model_cnn_new.add(Conv2D(16, (3, 3), padding='valid', activation = 'relu'))
# model_cnn_new.add(Conv2D(64, (3, 3), padding='valid', activation = 'relu'))
# model_cnn_new.add(BatchNormalization())
# # You can also don't use max pooling... it is up to you
# #model_cnn_new.add(MaxPooling2D(pool_size=(2,2))) # this line can lead to negative dimension problem
# model_cnn_new.add(Dropout(0.7))
#
# model_cnn_new.add(Conv2D(32, (5, 5), padding='valid', activation = 'relu'))
# model_cnn_new.add(BatchNormalization())
# model_cnn_new.add(MaxPooling2D(pool_size=(2,2)))
# model_cnn_new.add(Dropout(0.7))
#
# model_cnn_new.add(Flatten())
# model_cnn_new.add(Dense(128, activation = 'relu'))
#
# model_cnn_new.add(Dense(y_test_oh_n.shape[1], activation='softmax'))
# ```
# **Step 2**: Summarise the model.
#
# For example, you can try:
#
# ```python
# model_cnn_new.summary()
# ```
# **Step 3**: Define optimiser (try 'rmsprop', 'sgd', 'adagrad' or 'adadelta' if you wich), loss and metric
#
# For example:
# ``` python
# model_cnn_new.compile(optimizer='adam',
# loss='categorical_crossentropy',
# metrics=['accuracy'])
# ```
# **Step 4**: Train the model, here you can define the number of epochs and batch_size that best fit for you model
#
# For example:
# ```python
# # this can take SEVERAL minutes or even hours.. days... if your model is quite deep
# history_cnn_new = model_cnn_new.fit(x_train_new,
# y_train_oh_n,
# batch_size = 256,
# epochs = 100,
# validation_split = 0.2,
# verbose = 1)
# ```
# **Step 4**: Evaluate the model performance by using the metric that you think is the best.
#
# For example:
# ```python
# model_cnn_new.evaluate(x_test_new,y_test_oh_n)
#
# loss_value_cnn_n, acc_value_cnn_n = model_cnn_new.evaluate(x_test_new, y_test_oh_n, verbose=0)
# print('Loss value: ', loss_value_cnn_n)
# print('Acurracy value: ', acc_value_cnn_n)
# ```
#
# Plot the loss and accuracy if you which.
| notebooks/week07-convolutional-neural-networks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PythonData
# language: python
# name: pythondata
# ---
# +
# Importing dependencies and libraries
import numpy as np
import json
import requests
import pandas as pd
import time
import matplotlib.pyplot as plt
import scipy.stats as st
import os
from config import api_key
from citipy import citipy
from IPython.display import clear_output
from datetime import datetime
# +
# Building the 500+ cities list
# Setting ranges to look for cities
latitude_range = np.random.randint(-90,91, 70) #change 10 to 70 when testing is done
longitude_range = np.random.randint(-180,181, 70) #change 10 to 70 when testing is done
# This loop will ensure that the sample size is enough
# At least 500 (will do 550 just to be sure) cities with data are needed,
# with a 5.8% of missing data 577 cities is the minimum sample size
test = False
while test == False:
cities_list = []
#nested for loop that will retrieve every city
for x in longitude_range:
for y in latitude_range:
city = citipy.nearest_city(x,y)
cities_list.append(city.city_name)
# Converting the resulting list to a dataframe, dropping the duplicates and reindexing
cities_list_df = pd.DataFrame({'Cities':cities_list})
cities_list_df = cities_list_df.drop_duplicates().reset_index(drop = True)
# Making sure that cities list meets minimum requirements
if len(cities_list) >= 577: #change conditional to 577 after testing
test = True
# Making sure that cities list isn't too big, some test runs returned an 850 cities list
if len(cities_list_df.index) >= 650:
cities_list_df = cities_list_df.sample(650).reset_index(drop = True)
print(f'Sample size is {len(cities_list_df.index)} list built succesfully')
cities_list_df.head(10)
# +
#API requests
# Lists initialization
latitude = []
longitude = []
temperature = []
humidity = []
cloudiness = []
wind = []
failed_cities = []
# Base URL
url = "http://api.openweathermap.org/data/2.5/weather?"
# Variable initialization for progress tracker
request_num = 0
succ_request = 0
failed_request = 0
# for loop to request data from all cities
for city in cities_list_df['Cities']:
# Build query URL
query_url = url + "appid=" + api_key + "&q=" + city + "&units=imperial"
# Request data for the current city
weather_response = requests.get(query_url)
weather_json = weather_response.json()
# Printing progress information
clear_output(wait=True)
request_num += 1
print(f'Processing request {request_num}/{len(cities_list_df.index)} City: {city}')
print(f'{succ_request} succesfull {failed_request} failed')
# Making sure the request was succesful
if weather_response.status_code == 200:
# Appending current data to lists
latitude.append(weather_json['coord']['lat'])
longitude.append(weather_json['coord']['lon'])
temperature.append(weather_json['main']['temp'])
humidity.append(weather_json['main']['humidity'])
cloudiness.append(weather_json['clouds']['all'])
wind.append(weather_json['wind']['speed'])
succ_request += 1
else:
failed_request += 1
# Saving the failed city to be able to delete it from the dataframe
failed_cities.append(city)
# Making sure that time between requests is enough so that API key doesn't get blocked
time.sleep(1.01)
# Print out final request results
clear_output(wait=True)
print('API requests finished')
print(f'{succ_request} succesfull {failed_request} failed')
# +
# Cleaning and building the results dataframe
clean_weather_df = cities_list_df.copy(deep=True)
# Deleting the cities with missing values
for city in failed_cities:
# Finding the row to delete
row_index = clean_weather_df.loc[clean_weather_df['Cities'] == city, :].index
clean_weather_df.drop(row_index, inplace = True, axis = 0)
# Appending the data retrieved
clean_weather_df['Latitude'] = latitude
clean_weather_df['Longitude'] = longitude
clean_weather_df['Temperature (°F)'] = temperature
clean_weather_df['Humidity'] = humidity
clean_weather_df['Cloudiness'] = cloudiness
clean_weather_df['Wind Speed (mph)']= wind
clean_weather_df = clean_weather_df.reset_index(drop = True)
#saving the information dataframe to a csv file
# Obtaining a timestamp
now = datetime.now()
timestamp = int(datetime.timestamp(now))
filename = 'weather_data_' + str(timestamp) + '.csv'
path = os.path.join('results_csv', filename)
clean_weather_df.to_csv(path)
clean_weather_df.head(10)
# +
# If you wish to run the code witout making the API calls:
# 1. Run the first cell
# 2. Skip until cell 5 (this one)
# 3. Uncomment the block below assign filename to the csv file, and run all the remaining cells
# filename = 'weather_data_test.csv'
# path = os.path.join('results_csv', filename)
# clean_weather_df = pd.read_csv(path)
# Plotting Temperature (F) vs. Latitude
# Plot parameters
# X axis is fixed to show all possible latitude values
xticks = np.arange(-90,100,10)
# Y axis will contain max/min temperature values rounded to the closest 10
min_temp = (round(clean_weather_df['Temperature (°F)'].min()/10, 0))*10 - 10
max_temp = (round(clean_weather_df['Temperature (°F)'].max()/10,0))*10 + 10
yticks = np.arange(min_temp, max_temp+10, 10)
# Plotting
clean_weather_df.plot(kind="scatter", x="Latitude", y="Temperature (°F)", grid=True, figsize=(10,10),
title=f"Temperature (°F) vs Latitude for {datetime.today().strftime('%m/%d/%Y')}", xticks= xticks, yticks= yticks)
# Adding an explanatory line for the graph
plt.annotate("Possible correlation between temperature and latitude", xy=(-60,-10), annotation_clip=False, fontsize=15)
# Saving to png
filename = 'temp_lat.png'
path = os.path.join('plot_images', filename)
plt.savefig(path)
# +
# Plotting Humidity (%) vs. Latitude
# Plot parameters
# X axis is fixed to show all possible latitude values
xticks = np.arange(-90,100,10)
# Y axis is a percentage will go from 0 to 100 always
yticks = np.arange(0,110,10)
clean_weather_df.plot(kind="scatter", x="Latitude", y="Humidity", grid=True, figsize=(10,10),
title=f"Humidity (%) vs Latitude for {datetime.today().strftime('%m/%d/%Y')}", xticks= xticks, yticks= yticks)
# Adding an explanatory line for the graph
plt.annotate("Possible correlation between humidity and latitude", xy=(-60,-10), annotation_clip=False, fontsize=15)
# Saving to png
filename = 'hum_lat.png'
path = os.path.join('plot_images', filename)
plt.savefig(path)
# +
# Plotting Cloudiness (%) vs. Latitude
# Plot parameters
# X axis is fixed to show all possible latitude values
xticks = np.arange(-90,100,10)
# Y axis is a percentage will go from 0 to 100, but cloudiness can go to 100%, will adjust the axis to 105
yticks = np.arange(0,105,10)
clean_weather_df.plot(kind="scatter", x="Latitude", y="Cloudiness", grid=True, figsize=(10,10),
title=f"Cloudiness vs Latitude for {datetime.today().strftime('%m/%d/%Y')}", xticks= xticks, yticks= yticks)
# Adding an explanatory line for the graph
plt.annotate("Possible correlation between cloudiness and latitude", xy=(-60,-15), annotation_clip=False, fontsize=15)
# Saving to png
filename = 'cloud_lat.png'
path = os.path.join('plot_images', filename)
plt.savefig(path)
# +
# Plotting Wind Speed (mph) vs. Latitude
# Plot parameters
# X axis is fixed to show all possible latitude values
xticks = np.arange(-90,100,10)
# Y axis will contain max/min wind speed values rounded to the closest 10
min_wind = (round(clean_weather_df['Wind Speed (mph)'].min()/10, 0))*10 - 10
max_wind = (round(clean_weather_df['Wind Speed (mph)'].max()/10,0))*10 + 10
yticks = np.arange(min_wind, max_wind, 10)
clean_weather_df.plot(kind="scatter", x="Latitude", y="Wind Speed (mph)", grid=True, figsize=(10,10),
title=f"Wind Speed (mph) vs Latitude for {datetime.today().strftime('%m/%d/%Y')}", xticks= xticks, yticks= yticks)
# Adding an explanatory line for the graph
plt.annotate("Possible correlation between wind speed and latitude", xy=(-60,-14), annotation_clip=False, fontsize=15)
# Saving to png
filename = 'wind_lat.png'
path = os.path.join('plot_images', filename)
plt.savefig(path)
# +
# Temperature (F) vs. Latitude broken by hemisphere
# Splitting the dataframe into north and south hemisphere
north_df = clean_weather_df.loc[clean_weather_df['Latitude'] >= 0]
south_df = clean_weather_df.loc[clean_weather_df['Latitude'] < 0]
## North hemisphere
# Plot parameters
# X axis is fixed to show all possible latitude values
xticks = np.arange(0,100,10)
# Y axis will contain max/min temp values rounded to the closest 10
min_temp = (round(clean_weather_df['Temperature (°F)'].min()/10, 0))*10 - 10
max_temp = (round(clean_weather_df['Temperature (°F)'].max()/10,0))*10 + 10
yticks = np.arange(min_temp, max_temp, 10)
# Setting x and y for line
x_values = north_df['Latitude']
y_values = north_df['Temperature (°F)']
(m, b, r, p, stderr) = st.stats.linregress(x_values, y_values)
# Building the line
line_eq = 'y = ' + str(round(m, 2)) + 'x+' + str(round (b, 2))
# Plotting
fig, axes =plt.subplots(1,2,figsize=(11,11))
axes[0].scatter(x=north_df["Latitude"], y=north_df["Temperature (°F)"])
axes[0].grid()
axes[0].annotate(line_eq + ', R=' + str(round(r**2, 2)),(40,65), fontsize=15,color="red")
axes[0].plot(x_values, m*x_values + b, color ='r')
axes[0].set_xticks(xticks)
axes[0].set_yticks(yticks)
axes[0].set_xlabel('Latitude')
axes[0].set_ylabel('Temperature (°F)')
axes[0].set_title('Northern Hemisphere')
## South hemisphere
# Plot parameters
# X axis is fixed to show all possible latitude values
xticks = np.arange(-90,10,10)
# Y axis will contain max/min temp values rounded to the closest 10
min_temp = (round(south_df['Temperature (°F)'].min()/10,0))*10 - 10
max_temp = (round(south_df['Temperature (°F)'].max()/10,0))*10 + 10
yticks = np.arange(min_temp, max_temp, 10)
# Setting x and y for line
x_values = south_df['Latitude']
y_values = south_df['Temperature (°F)']
(m, b, r, p, stderr) = st.stats.linregress(x_values, y_values)
# Building the line
line_eq = 'y = ' + str(round(m, 2)) + 'x+' + str(round (b, 2))
axes[1].scatter(x=south_df["Latitude"], y=south_df["Temperature (°F)"])
axes[1].grid()
axes[1].annotate(line_eq + ', R=' + str(round(r**2, 2)),(-80,75), fontsize=15,color="red")
axes[1].plot(x_values, m*x_values + b, color ='r')
axes[1].set_xticks(xticks)
axes[1].set_yticks(yticks)
axes[1].set_xlabel('Latitude')
axes[1].set_ylabel('Temperature (°F)')
axes[1].set_title('Southern Hemisphere')
plt.suptitle (f"Northern And Southern Hemisphere Correlation Between Temperature and Latitude for {datetime.today().strftime('%m/%d/%Y')}", fontsize = 15)
plt.tight_layout()
# Adding an explanatory line for the graph
plt.annotate("Looking at the scatter points, the regression line and the R value it is determined that the \n\
closer we get to latitude 0° the higher the temperature \n\
The correlation between the 2 variables is strong", xy=(-200,20), annotation_clip=False, fontsize=15)
# Saving to png
filename = 'hemi_temp_lat.png'
path = os.path.join('plot_images', filename)
plt.savefig(path)
# +
# Humidity (%) vs. Latitude broken by hemisphere
## North hemisphere
# Plot parameters
# X axis is fixed to show all possible latitude values
xticks = np.arange(0,100,10)
# Y axis will have 0-100% humidity
yticks = np.arange(0, 110, 10)
# Setting x and y for line
x_values = north_df['Latitude']
y_values = north_df['Humidity']
(m, b, r, p, stderr) = st.stats.linregress(x_values, y_values)
# Building the line
line_eq = 'y = ' + str(round(m, 2)) + 'x+' + str(round (b, 2))
# Plotting
fig, axes =plt.subplots(1,2,figsize=(11,11))
axes[0].scatter(x=north_df["Latitude"], y=north_df["Humidity"])
axes[0].grid()
axes[0].annotate(line_eq + ', R=' + str(round(r**2, 2)),(40,65), fontsize=15,color="red")
axes[0].plot(x_values, m*x_values + b, color ='r')
axes[0].set_xticks(xticks)
axes[0].set_yticks(yticks)
axes[0].set_xlabel('Latitude')
axes[0].set_ylabel('Humidity (%)')
axes[0].set_title('Northern Hemisphere')
## South hemisphere
# Plot parameters
# X axis is fixed to show all possible latitude values
xticks = np.arange(-90,10,10)
# Y axis will have 0-100% humidity
yticks = np.arange(0, 110, 10)
# Setting x and y for line
x_values = south_df['Latitude']
y_values = south_df['Humidity']
(m, b, r, p, stderr) = st.stats.linregress(x_values, y_values)
# Building the line
line_eq = 'y = ' + str(round(m, 2)) + 'x+' + str(round (b, 2))
axes[1].scatter(x=south_df["Latitude"], y=south_df["Humidity"])
axes[1].grid()
axes[1].annotate(line_eq + ', R=' + str(round(r**2, 2)),(-80,75), fontsize=15,color="red")
axes[1].plot(x_values, m*x_values + b, color ='r')
axes[1].set_xticks(xticks)
axes[1].set_yticks(yticks)
axes[1].set_xlabel('Latitude')
axes[1].set_ylabel('Humidity (%)')
axes[1].set_title('Southern Hemisphere')
plt.suptitle (f"Northern And Southern Hemisphere Correlation Between Humidity and Latitude for {datetime.today().strftime('%m/%d/%Y')}", fontsize = 15)
plt.tight_layout()
# Adding an explanatory line for the graph
plt.annotate("Looking at the scatter points, the regression line and the R value it is determined that\n\
latitude is not determinant in the humidity levels \n\
The correlation between the 2 variables is very weak", xy=(-200,-15), annotation_clip=False, fontsize=15)
# Saving to png
filename = 'hemi_humi_lat.png'
path = os.path.join('plot_images', filename)
plt.savefig(path)
# +
# Northern Hemisphere - Cloudiness (%) vs. Latitude broken by hemisphere
## North hemisphere
# Plot parameters
# X axis is fixed to show all possible latitude values
xticks = np.arange(0,100,10)
# Y axis will have 0-100% cloudiness
yticks = np.arange(0, 110, 10)
# Setting x and y for line
x_values = north_df['Latitude']
y_values = north_df['Cloudiness']
(m, b, r, p, stderr) = st.stats.linregress(x_values, y_values)
# Building the line
line_eq = 'y = ' + str(round(m, 2)) + 'x+' + str(round (b, 2))
# Plotting
fig, axes =plt.subplots(1,2,figsize=(11,11))
axes[0].scatter(x=north_df["Latitude"], y=north_df["Cloudiness"])
axes[0].grid()
axes[0].annotate(line_eq + ', R=' + str(round(r**2, 2)),(40,65), fontsize=15,color="red")
axes[0].plot(x_values, m*x_values + b, color ='r')
axes[0].set_xticks(xticks)
axes[0].set_yticks(yticks)
axes[0].set_xlabel('Latitude')
axes[0].set_ylabel('Cloudiness (%)')
axes[0].set_title('Northern Hemisphere')
## South hemisphere
# Plot parameters
# X axis is fixed to show all possible latitude values
xticks = np.arange(-90,10,10)
# Y axis will have 0-100% humidity
yticks = np.arange(0, 110, 10)
# Setting x and y for line
x_values = south_df['Latitude']
y_values = south_df['Cloudiness']
(m, b, r, p, stderr) = st.stats.linregress(x_values, y_values)
# Building the line
line_eq = 'y = ' + str(round(m, 2)) + 'x+' + str(round (b, 2))
axes[1].scatter(x=south_df["Latitude"], y=south_df["Cloudiness"])
axes[1].grid()
axes[1].annotate(line_eq + ', R=' + str(round(r**2, 2)),(-80,75), fontsize=15,color="red")
axes[1].plot(x_values, m*x_values + b, color ='r')
axes[1].set_xticks(xticks)
axes[1].set_yticks(yticks)
axes[1].set_xlabel('Latitude')
axes[1].set_ylabel('Cloudiness (%)')
axes[1].set_title('Southern Hemisphere')
plt.suptitle (f"Northern And Southern Hemisphere Correlation Between Cloudiness and Latitude for {datetime.today().strftime('%m/%d/%Y')}", fontsize = 15)
plt.tight_layout()
# Adding an explanatory line for the graph
plt.annotate("Looking at the scatter points, the regression line and the R value it is determined that\n\
latitude is not determinant in the cloudiness levels \n\
The correlation between the 2 variables is very weak", xy=(-200,-20), annotation_clip=False, fontsize=15)
# Saving to png
filename = 'hemi_cloud_lat.png'
path = os.path.join('plot_images', filename)
plt.savefig(path)
# +
# Northern Hemisphere - Wind Speed (mph) vs. Latitude broken by hemisphere
## North hemisphere
# Plot parameters
# X axis is fixed to show all possible latitude values
xticks = np.arange(0,100,10)
# Y axis will contain max/min wind speed values rounded to the closest 10
min_temp = (round(clean_weather_df['Wind Speed (mph)'].min()/10, 0))*10 - 10
max_temp = (round(clean_weather_df['Wind Speed (mph)'].max()/10,0))*10 + 10
yticks = np.arange(min_temp, max_temp, 10)
# Setting x and y for line
x_values = north_df['Latitude']
y_values = north_df['Wind Speed (mph)']
(m, b, r, p, stderr) = st.stats.linregress(x_values, y_values)
# Building the line
line_eq = 'y = ' + str(round(m, 2)) + 'x+' + str(round (b, 2))
# Plotting
fig, axes =plt.subplots(1,2,figsize=(11,11))
axes[0].scatter(x=north_df["Latitude"], y=north_df["Wind Speed (mph)"])
axes[0].grid()
axes[0].annotate(line_eq + ', R=' + str(round(r**2, 2)),(10,0), fontsize=15,color="red")
axes[0].plot(x_values, m*x_values + b, color ='r')
axes[0].set_xticks(xticks)
axes[0].set_yticks(yticks)
axes[0].set_xlabel('Latitude')
axes[0].set_ylabel('Wind Speed (mph)')
axes[0].set_title('Northern Hemisphere')
## South hemisphere
# Plot parameters
# X axis is fixed to show all possible latitude values
xticks = np.arange(-90,10,10)
# Y axis will contain max/min temp values rounded to the closest 10
min_temp = (round(south_df['Wind Speed (mph)'].min()/10,0))*10 - 10
max_temp = (round(south_df['Wind Speed (mph)'].max()/10,0))*10 + 10
yticks = np.arange(min_temp, max_temp, 10)
# Setting x and y for line
x_values = south_df['Latitude']
y_values = south_df['Wind Speed (mph)']
(m, b, r, p, stderr) = st.stats.linregress(x_values, y_values)
# Building the line
line_eq = 'y = ' + str(round(m, 2)) + 'x+' + str(round (b, 2))
axes[1].scatter(x=south_df["Latitude"], y=south_df["Wind Speed (mph)"])
axes[1].grid()
axes[1].annotate(line_eq + ', R=' + str(round(r**2, 2)),(-80,0), fontsize=15,color="red")
axes[1].plot(x_values, m*x_values + b, color ='r')
axes[1].set_xticks(xticks)
axes[1].set_yticks(yticks)
axes[1].set_xlabel('Latitude')
axes[1].set_ylabel('Wind Speed (mph)')
axes[1].set_title('Southern Hemisphere')
plt.suptitle (f"Northern And Southern Hemisphere Correlation Between Wind Speed and Latitude for {datetime.today().strftime('%m/%d/%Y')}", fontsize = 15)
plt.tight_layout()
# Adding an explanatory line for the graph
plt.annotate("Looking at the scatter points, the regression line and the R value it is determined that\n\
latitude is not determinant in the wind speed \n\
The correlation between the 2 variables is very weak", xy=(-200,-15), annotation_clip=False, fontsize=15)
# Saving to png
filename = 'hemi_wind_lat.png'
path = os.path.join('plot_images', filename)
plt.savefig(path)
# -
| WeatherPy/WeatherPy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# We have data about activity on the USA-Canada and USA-Mexico border.
#
# In this kernel, I have tried to do an exploratory analysis of the activity at the border. This kernel only contains the trends of activity at the borders over the years. Initially I have focused on the type of activity which involved people. Later on, I will proceed to do exploratory analysis of border activity regarding goods.
# Any suggestions regarding this kernels are welcome :-)
# Lets start with importing the data as below
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load in
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# Any results you write to the current directory are saved as output.
# -
# After that, lets import some libraries.
#Import Library
import matplotlib.pyplot as plt
plt.style.use('ggplot')
import seaborn as sns
from wordcloud import WordCloud
from sklearn.preprocessing import StandardScaler
# Data has been taken into a variable and primitive cleaning is done.
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
data=pd.read_csv('/kaggle/input/border-crossing-entry-data/Border_Crossing_Entry_Data.csv')
data['Date']=pd.to_datetime(data['Date'])
# -
# Lets see the levels of activity on the two borders
# +
state_cou=data.groupby(['Border']).size().reset_index(name='count')
data.groupby(['Border']).size().reset_index(name='count')
sns.set_context('talk')
sns.barplot(x=state_cou['Border'],y=state_cou['count'],palette='deep')
plt.xticks(rotation=90)
plt.ylabel('Activity at Entry Ports')
plt.title('Border vs Count')
plt.show()
# -
# Wordcloud can be created for so see the kinds of objects that are listed in the data which pass through the borders.
# +
abc=','
abc2=data.Measure.unique()
abc3=abc.join(abc2)
abc3=abc3.replace(" ","_")
wordcloud = WordCloud(width = 800, height = 800,
background_color ='white',
min_font_size = 10).generate(abc3)
plt.figure(figsize = (8, 8), facecolor = None)
plt.imshow(wordcloud)
plt.axis("off")
plt.tight_layout(pad = 0)
plt.show()
# -
# Below, two arrays have been created each listing mobiles that pass through border which can be differentiated by if they carry people or goods.
people=['Personal vehicles','Bus Passengers','Personal Vehicle Passengers','Train Passengers','Trains','Buses']
data[data.Measure.isin(people)]
goods=['Truck Containers Empty','Rail Containers Full','Rail Containers Empty','Truck Containers Full','Trucks']
#data[data.Measure.isin(goods)]
# Get the latest date from the data
data['Date'].max()
# Get the earliest date from the data
data['Date'].min()
# These are just used for checking and getting to know the data better.
#
#
# Group the data based on the Measure, Border and the date of activity as below
data_border_pass=data[data.Measure.isin(people)].groupby(['Border','Measure','Date'],as_index=False).sum()
#data_border_pass
# Firstly, I have taken data solely for US-Mexico Border and aggregated the activity based on the Measure.
#
# Port code has been dropped as it the columns was irrelevant for my scope of analysis. Additionally, the date has been mutated and only year from this columns has been extracted as i had to observe the trends of activities at the border over the years.
#
#data_border_pass[data_border_pass['Border']=='US-Mexico Border']['Border']
dat1=data_border_pass[data_border_pass['Border']=='US-Mexico Border']
dat1=dat1.drop(['Port Code'],axis=1)
dat1=dat1.groupby(['Measure','Date'],as_index=False).sum()
dat1['Year']=pd.DatetimeIndex(dat1['Date']).year
dat1.groupby(['Measure','Year'],as_index=False).sum()
# See the Unique measures which carry people
dat1['Measure'].unique()
# A python function can be created to plot bart graphs(which are mostly used in the analysis) to avoid redundant code.
def bar_plt(x_bar,y_bar,ylab,titl):
sns.set_context('talk')
sns.barplot(x=x_bar,y=y_bar,palette='deep')
plt.xticks(rotation=90)
plt.ylabel(ylab)
plt.title(titl)
# Lets see how the activity of Bus with passengers has been over the years.
dat_temp=dat1[dat1['Measure']=='Bus Passengers']
dat_temp=dat_temp.groupby(['Measure','Year'],as_index=False).sum()
bar_plt(dat_temp['Year'],dat_temp['Value']
,'Count of Activity'
,'Bus Passengers Activity US-MEX')
# It can be seen from the graph that the Passengers activity at the US-Mex border declined in year 2009 but bounced back up for a brief period of time only to fall again.
dat_temp=dat1[dat1['Measure']=='Buses']
dat_temp=dat_temp.groupby(['Measure','Year'],as_index=False).sum()
bar_plt(dat_temp['Year'],dat_temp['Value']
,'Count of Activity'
,'Buses Activity US-MEX')
# It can be seen from the graog that the frequency of buses on US-Mex border has been declining steadily since 2009. This might be as a result of reduced demand after 2009. This phenomenon of graphs tipping downwards in 2009 is a result of financial crisis of 2009 when economy was slowed down. You will see this phenomenon in the rest of the following graphs as well.
#
# In the following map we will add two graphs namely, Bus Passengers Activity and Buses activity across the US-MEX border to see the relation between these two trends more clearly.
# +
fig, ax1 = plt.subplots()
color = 'tab:red'
#ax1.set_xlabel('time (s)')
ax1.set_ylabel('Buses', color=color)
ax1.bar(dat1[dat1['Measure']=='Buses']['Year'], dat1[dat1['Measure']=='Buses']['Value'], color=color)
ax1.tick_params(axis='y', labelcolor=color)
ax2 = ax1.twinx() # instantiate a second axes that shares the same x-axis
color = 'tab:blue'
ax2.set_ylabel('Bus Passengers', color=color) # we already handled the x-label with ax1
ax2.plot(dat1[dat1['Measure']=='Bus Passengers']['Year'], dat1[dat1['Measure']=='Bus Passengers']['Value'], color=color)
ax2.tick_params(axis='y', labelcolor=color)
fig.tight_layout() # otherwise the right y-label is slightly clipped
plt.show()
# -
# Look at the train activity for US-MEX border
dat_temp=dat1[dat1['Measure']=='Trains']
dat_temp=dat_temp.groupby(['Measure','Year'],as_index=False).sum()
bar_plt(dat_temp['Year'],dat_temp['Value']
,'Count of Activity'
,'Trains Activity US-MEX')
# As expected, the trains activity at the border US-Mexico declined sharply in 2009 but increased every year after it. In 2017 the train frequency was greater than 2007, the year in which the economy was at best.
# This might be the reason that frequency of buses was reduced at the border, increased frequencies of trains might have reduced the demand of Buses. It will be interesting to see the passenger activity though trains after this.
dat_temp=dat1[dat1['Measure']=='Train Passengers']
dat_temp=dat_temp.groupby(['Measure','Year'],as_index=False).sum()
bar_plt(dat_temp['Year'],dat_temp['Value']
,'Count of Activity'
,'Trains Passengers Activity US-MEX')
# What a surprise! Even though the train frequency at the US-MEX border was increasing through 2009-2013, the passenger activity was almost the same throughout that period. It did catch up after 2013 but it wasnt on the same levels as before 2009.
# We will plot the same kind of graph we plotted for the Bus related activity for trains to see the relation and different between them.
# +
fig, ax1 = plt.subplots()
color = 'tab:red'
#ax1.set_xlabel('time (s)')
ax1.set_ylabel('Trains', color=color)
ax1.bar(dat1[dat1['Measure']=='Trains']['Year'], dat1[dat1['Measure']=='Trains']['Value'], color=color)
ax1.tick_params(axis='y', labelcolor=color)
ax2 = ax1.twinx() # instantiate a second axes that shares the same x-axis
color = 'tab:blue'
ax2.set_ylabel('Train Passengers', color=color) # we already handled the x-label with ax1
ax2.plot(dat1[dat1['Measure']=='Train Passengers']['Year'], dat1[dat1['Measure']=='Train Passengers']['Value'], color=color)
ax2.tick_params(axis='y', labelcolor=color)
fig.tight_layout() # otherwise the right y-label is slightly clipped
plt.show()
# -
# Lets see the activity at US-MEX border for Personal Vehicles
dat_temp=dat1[dat1.Measure.isin(['Personal Vehicle Passengers', 'Personal Vehicles'])]
dat_temp=dat_temp.groupby(['Measure','Year'],as_index=False).sum()
bar_plt(dat_temp['Year'],dat_temp['Value']
,'Count of Activity'
,'Personal Vehcles Activity US-MEX')
# The activity was at the lowest at 2010 but has been increasing steadily after that.
# Below graphs in which the above mentioned activities were taken and mean frequency for the each one them was taken. After that graph for plotted for the activities over the years with respect to the mean frequency. This was done in hope to create a Facet Grid for the activities. It was important to plot the count of activities with respecto to mean frequency to bring them all to the same scale which is very much needed for the facet grid.
scaler=StandardScaler()
dat_bus=dat1[dat1['Measure']=='Bus Passengers']
dat_bus=dat_bus.groupby(['Measure','Year'],as_index=False).sum()
dat_bus=pd.DataFrame(dat_bus)
scaler.fit(dat_bus.iloc[:,2].values.reshape(-1,1))
dat_bus['Value']=pd.DataFrame(scaler.transform(np.asarray(dat_bus['Value']).reshape(-1,1)))
bar_plt(dat_bus['Year'],dat_bus['Value'],'Count of Activity(wrt Mean Activity)','Bus Passenger Activity')
scaler1=StandardScaler()
dat_train=dat1[dat1['Measure']=='Train Passengers']
dat_train=dat_train.groupby(['Measure','Year'],as_index=False).sum()
dat_train=pd.DataFrame(dat_train)
scaler1.fit(dat_train.iloc[:,2].values.reshape(-1,1))
dat_train['Value']=pd.DataFrame(scaler1.transform(np.asarray(dat_train['Value']).reshape(-1,1)))
bar_plt(dat_train['Year'],dat_train['Value'],'Count of Activity(wrt Mean Activity)','Train Passengers Activity')
scaler2=StandardScaler()
dat_pass=dat1[dat1['Measure']=='Personal Vehicle Passengers']
dat_pass=dat_pass.groupby(['Measure','Year'],as_index=False).sum()
dat_pass=pd.DataFrame(dat_pass).reset_index()
scaler2.fit(dat_pass.iloc[:,3].values.reshape(-1,1))
dat_pass['Value']=pd.DataFrame(scaler2.transform(np.asarray(dat_pass['Value']).reshape(-1,1)))
bar_plt(dat_pass['Year'],dat_pass['Value'],'Count of Activity(wrt Mean Activity)','Personal Vehicle Passengers Activity')
scaler3=StandardScaler()
dat_tra=dat1[dat1['Measure']=='Trains']
dat_tra=dat_tra.groupby(['Measure','Year'],as_index=False).sum()
dat_tra=pd.DataFrame(dat_tra).reset_index()
scaler2.fit(dat_tra.iloc[:,3].values.reshape(-1,1))
dat_tra['Value']=pd.DataFrame(scaler2.transform(np.asarray(dat_tra['Value']).reshape(-1,1)))
bar_plt(dat_tra['Year'],dat_tra['Value'],'Count of Activity(wrt Mean Activity)','Trains Activity')
# Following is an attempt to create a facetgrid for the above mentioned activities but somehow the layout is off. So any suggestions to make it better are welcome.
dat_append=dat_bus.append(dat_train).append(dat_pass).append(dat_tra)
sns.set_context('talk')
g=sns.FacetGrid(dat_append,col="Measure",col_wrap=2)
g.map(sns.barplot,'Year','Value',palette='deep')
g.fig.tight_layout()
# In following graphs the similar trends for US-Canada Border are plotted.
dat2=data_border_pass[data_border_pass['Border']=='US-Canada Border']
dat2=dat2.drop(['Port Code'],axis=1)
dat2=dat2.groupby(['Measure','Date'],as_index=False).sum()
dat2['Year']=pd.DatetimeIndex(dat1['Date']).year
dat2.groupby(['Measure','Year'],as_index=False).sum()
dat2['Measure'].unique()
bar_plt(dat2[dat2['Measure']=='Bus Passengers']['Year'],dat2[dat2['Measure']=='Bus Passengers']['Value'],
'Count of Activity'
,'Bus Passengers Activity US-CAN')
bar_plt(dat2[dat2['Measure']=='Buses']['Year'],dat2[dat2['Measure']=='Buses']['Value'],
'Count of Activity'
,'Buses Activity US-CAN')
bar_plt(dat2[dat2['Measure']=='Train Passengers']['Year'],dat2[dat2['Measure']=='Train Passengers']['Value'],
'Count of Activity'
,'Train Passengers Activity US-CAN')
bar_plt(dat2[dat2['Measure']=='Trains']['Year'],dat2[dat2['Measure']=='Trains']['Value'],
'Count of Activity'
,'Trains Activity US-CAN')
bar_plt(dat2[dat2['Measure']=='Personal Vehicle Passengers']['Year'],dat2[dat2['Measure']=='Personal Vehicle Passengers']['Value'],
'Count of Activity'
,'Personal Passengers vehicles Activity US-CAN')
| Border_Crossing_Entry_Data.csv/.ipynb_checkpoints/usa-border-activity-exploratory-analysis-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Processing data in chunks (1)
# Sometimes, data sources can be so large in size that storing the entire dataset in memory becomes too resource-intensive. In this exercise, you will process the first 1000 rows of a file line by line, to create a dictionary of the counts of how many times each country appears in a column in the dataset.
#
# The csv file 'world_dev_ind.csv' is in your current directory for your use. To begin, you need to open a connection to this file using what is known as a context manager. For example, the command with open('datacamp.csv') as datacamp binds the csv file 'datacamp.csv' as datacamp in the context manager. Here, the with statement is the context manager, and its purpose is to ensure that resources are efficiently allocated when opening a connection to a file.
#
# If you'd like to learn more about context managers, refer to the DataCamp course on Importing Data in Python.
#
# ### Instructions
#
# - Use open() to bind the csv file 'world_dev_ind.csv' as file in the context manager.
# - omplete the for loop so that it iterates 1000 times to perform the loop body and process only the first 1000 rows of data of the file.
#
# +
# Open a connection to the file
with open('world_ind_pop_data.csv') as file:
# Skip the column names
file.readline()
# Initialize an empty dictionary: counts_dict
counts_dict = {}
# Process only the first 1000 rows
for j in range(1000):
# Split the current line into a list: line
line = file.readline().split(',')
# Get the value for the first column: first_col
first_col = line[0]
# If the column value is in the dict, increment its value
if first_col in counts_dict.keys():
counts_dict[first_col] += 1
# Else, add to the dict and set value to 1
else:
counts_dict[first_col] = 1
# Print the resulting dictionary
print(counts_dict)
# -
# readlines splits the entire file into lines and is equivalent to file.read().split('\n'), but is a bit more efficient. Your example,
# for line in file: values = line.split()
# splits each line by its spaces, building a list of words in the line. value is overwritten on each iteration so unless you save values somewhere, only parts of the file are in-memory at a single time.
| Python Data Science Toolbox -Part 2/Bringing it all together/05. Processing data in chunks (1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.11 64-bit (''steel_segmentation'': conda)'
# name: python3
# ---
# +
# default_exp losses
# all_slow
# -
# # Loss functions
#
# > Various loss functions in PyTorch
# hide
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# export
import torch
import torch.nn as nn
import torch.nn.functional as F
from fastai.torch_core import TensorBase
from fastai.losses import *
from fastai.callback.core import Callback
from fastai.vision.all import *
import numpy as np
from torch.nn.modules.loss import _Loss
import segmentation_models_pytorch as smp
from steel_segmentation.utils import get_train_df
from steel_segmentation.transforms import SteelDataBlock, SteelDataLoaders
path = Path("../data")
train_pivot = get_train_df(path=path, pivot=True)
block = SteelDataBlock(path)
dls = SteelDataLoaders(block, train_pivot, bs=8)
xb, yb = dls.one_batch()
print(xb.shape, xb.device)
print(yb.shape, yb.device)
device = "cuda" if torch.cuda.is_available() else "cpu"
device
# +
model = smp.Unet("resnet18", classes=4).to(device)
logits = model(xb)
probs = torch.sigmoid(logits)
preds = ( probs > 0.5).float()
# -
# export
#reference: https://github.com/asanakoy/kaggle_carvana_segmentation/blob/master/asanakoy/losses.py
class SoftDiceLoss(nn.Module):
def __init__(self):
super().__init__()
def forward(self, logits, labels):
probs = torch.sigmoid(logits)
num = labels.size(0)
m1 = probs.view(num, -1)
m2 = labels.view(num, -1)
intersection = (m1 * m2)
score = 2. * (intersection.sum(1) + 1) / (m1.sum(1) + m2.sum(1) + 1)
score = 1 - score.sum() / num
return score
criterion = SoftDiceLoss()
criterion(logits.detach().cpu(), yb)
# export
#reference: https://github.com/zdaiot/Kaggle-Steel-Defect-Detection
class WeightedSoftDiceLoss(nn.Module):
def __init__(self, size_average=True, weight=[0.2, 0.8]):
super().__init__()
self.size_average = size_average
self.weight = torch.FloatTensor(weight)
def forward(self, logit_pixel, truth_pixel):
batch_size = len(logit_pixel)
logit = logit_pixel.view(batch_size, -1)
truth = truth_pixel.view(batch_size, -1)
assert(logit.shape == truth.shape)
loss = self.soft_dice_criterion(logit, truth)
if self.size_average:
loss = loss.mean()
return loss
def soft_dice_criterion(self, logit, truth):
batch_size = len(logit)
probability = torch.sigmoid(logit)
p = probability.view(batch_size, -1)
t = truth.view(batch_size, -1)
w = truth.detach()
self.weight = self.weight.type_as(logit)
w = w * (self.weight[1] - self.weight[0]) + self.weight[0]
p = w * (p*2 - 1) #convert to [0,1] --> [-1, 1]
t = w * (t*2 - 1)
intersection = (p * t).sum(-1)
union = (p * p).sum(-1) + (t * t).sum(-1)
dice = 1 - 2 * intersection/union
loss = dice
return loss
criterion = WeightedSoftDiceLoss()
criterion(logits.detach().cpu(), yb)
# export
#reference: https://github.com/zdaiot/Kaggle-Steel-Defect-Detection
class SoftBCEDiceLoss(nn.Module):
def __init__(self, bce_pos_weight, size_average=True, dice_weights=[0.2, 0.8], loss_weights=[0.7, 0.3]):
super().__init__()
self.size_average = 'mean' if size_average else 'none'
self.loss_weights = loss_weights
self.bce_loss = nn.BCEWithLogitsLoss(
reduction=self.size_average,
pos_weight=torch.tensor(bce_pos_weight)
)
self.softdiceloss = WeightedSoftDiceLoss(
size_average=self.size_average,
weight=dice_weights
)
def forward(self, input, target):
input, target = TensorBase(input).float(), TensorBase(target).float()
bce_loss = self.bce_loss(input, target)
soft_dice_loss = self.softdiceloss(input, target)
loss = self.loss_weights[0] * bce_loss + self.loss_weights[1] * soft_dice_loss
return loss
criterion = SoftBCEDiceLoss(bce_pos_weight=1.5)
criterion(logits.detach().cpu(), yb)
# export
#reference: https://github.com/zdaiot/Kaggle-Steel-Defect-Detection
class MultiClassesSoftBCEDiceLoss(nn.Module):
def __init__(self, classes_num=4, size_average=True, dice_weights=[0.2, 0.8], bce_pos_weights=[2.0,2.0,1.0,1.5], loss_weights=[0.7, 0.3], thresh=0.5):
super().__init__()
self.thresh = thresh
self.classes_num = classes_num
self.soft_bce_dice_losses = [
SoftBCEDiceLoss(bce_pos_weight=pos_weight, size_average=size_average, dice_weights=dice_weights, loss_weights=loss_weights)
for pos_weight in bce_pos_weights
]
def forward(self, input, target):
"""
Args:
input: tensor, [batch_size, classes_num, height, width]
target: tensor, [batch_size, classes_num, height, width]
"""
loss = 0
for class_index in range(self.classes_num):
input_single_class = input[:, class_index, :, :]
target_singlt_class = target[:, class_index, :, :]
single_class_loss = self.soft_bce_dice_losses[class_index](input_single_class, target_singlt_class)
loss += single_class_loss
loss /= self.classes_num
return loss
def decodes(self, x):
return (x>self.thresh).float().argmax(dim=1)
def activation(self, x):
return torch.sigmoid(x)
criterion = MultiClassesSoftBCEDiceLoss()
loss = criterion(logits.detach().cpu(), yb)
loss
criterion.decodes(logits.detach().cpu())
criterion.activation(logits.detach().cpu()).shape
# For the Tensorboard callback we need this Learner Callback to handle the step after the prediction.
# export
class LossEnabler(Callback):
"""Cast predictions and labels to TensorBase to compute the smp.losses"""
def after_pred(self):
if len(self.learn.yb) > 0:
yb = self.learn.yb[0]
self.learn.yb = (TensorBase(yb), )
self.learn.pred = TensorBase(self.learn.pred)
dls.valid.bs
| nbs/04_loss.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + Collapsed="false"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.model_selection import train_test_split
from xgboost import XGBRegressor
# %pylab inline
# %matplotlib inline
# + Collapsed="false"
data=pd.read_csv('train.csv')
# + Collapsed="false"
test=pd.read_csv('test.csv')
# + Collapsed="false"
variaveis=[]
for i in test:
variaveis.append(i)
variaveis.append('NU_NOTA_MT')
# + Collapsed="false"
xdata=data[variaveis]
# + Collapsed="false"
total = xdata.isnull().sum().sort_values(ascending=False)
percent = (xdata.isnull().sum()/xdata.isnull().count()).sort_values(ascending=False)
missing_data = pd.concat([total, percent], axis=1, keys=['Total', 'Percent'])
missing_data.head(20)
# + Collapsed="false"
xdata['NU_NOTA_MT'].describe()
# + Collapsed="false"
xdata.plot.scatter(x='NU_NOTA_CN', y='NU_NOTA_MT')
xdata.plot.scatter(x='NU_NOTA_CH', y='NU_NOTA_MT')
xdata.plot.scatter(x='NU_NOTA_LC', y='NU_NOTA_MT')
xdata.plot.scatter(x='NU_NOTA_REDACAO', y='NU_NOTA_MT')
# + [markdown] Collapsed="false"
# Vendo as correlações
# + Collapsed="false"
aux = xdata.copy()
aux2 = xdata.copy()
aux = aux.loc[:, xdata.columns]
aux['NU_NOTA_MT'] = aux2.NU_NOTA_MT
c = aux.corr()
c.NU_NOTA_MT.sort_values()
# + Collapsed="false"
new_vector_training = [
'NU_NOTA_COMP1',
'NU_NOTA_COMP2',
'NU_NOTA_COMP4',
'NU_NOTA_COMP5',
'NU_NOTA_COMP3',
'NU_NOTA_REDACAO',
'NU_NOTA_LC',
'NU_NOTA_CH',
'NU_NOTA_CN',
'NU_NOTA_MT'
]
new_vector_test = [
'NU_INSCRICAO',
'NU_NOTA_COMP1',
'NU_NOTA_COMP2',
'NU_NOTA_COMP4',
'NU_NOTA_COMP5',
'NU_NOTA_COMP3',
'NU_NOTA_REDACAO',
'NU_NOTA_LC',
'NU_NOTA_CH',
'NU_NOTA_CN'
]
# + Collapsed="false"
enemTrain_data = xdata.copy()
enemTrain_data = enemTrain_data.loc[:, new_vector_training]
enemTrain_data.dropna(subset=['NU_NOTA_MT'], inplace=True)
enemTrain_data.head()
# + Collapsed="false"
y = enemTrain_data.NU_NOTA_MT
X = enemTrain_data.drop(['NU_NOTA_MT'], axis=1)
# + Collapsed="false"
enem_validation_data = test.copy()
enem_validation_data_1 = enem_validation_data.loc[:, new_vector_test]
enem_validation_data_2 = enem_validation_data.loc[:, new_vector_test]
# + Collapsed="false"
enem_train_X, enem_validation_X, enem_train_y, enem_validation_y = train_test_split(X, y, random_state = 0)
# + Collapsed="false"
model = XGBRegressor(n_estimators=200, learning_rate=0.1)
model.fit(enem_train_X, enem_train_y, early_stopping_rounds=5, eval_set=[(enem_validation_X, enem_validation_y)], verbose=False)
# + Collapsed="false"
enem_validation_data_1.drop(['NU_INSCRICAO'], axis=1, inplace=True)
# + Collapsed="false"
predicted_nota = model.predict(enem_validation_data_1)
result_df = pd.DataFrame({'NU_INSCRICAO': enem_validation_data_2['NU_INSCRICAO'], 'NU_NOTA_MT': predicted_nota})
# + Collapsed="false"
result_df.head()
# + Collapsed="false"
result_df.isnull().any().any()
# + Collapsed="false"
result_df_final = result_df.loc[: , ['NU_INSCRICAO', 'NU_NOTA_MT']]
result_df.to_csv('answer.csv', index=False)
# + Collapsed="false"
| enem-2/Enem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from tqdm import tqdm
from database.strategy import Strategy
from database.market import Market
import warnings
warnings.simplefilter(action='ignore', category=Warning)
import matplotlib.pyplot as plt
from datetime import datetime, timedelta
import math
import numpy as np
import statistics
from sklearn.preprocessing import normalize
strat_db = Strategy("unity")
market = Market()
suffix = "tsmc_full"
market.connect()
sp5 = market.retrieve_data("sp500")
market.close()
strat_db.connect()
tse = strat_db.retrieve_data("{}_epochs".format(suffix))
strat_db.close()
bench = pd.read_csv("GSPC.csv")
list(tse.columns)
strat_db.connect()
analysis = []
start = datetime(2018,1,1)
end = datetime(2021,1,1)
seats = 4
cash = 100
for epoch in tqdm(tse["epoch"].unique()):
epoch_dict = tse[tse["epoch"]==epoch].iloc[0]
trades = strat_db.retrieve_data("{}_{}".format(suffix,epoch))
trades = trades[(trades["date"]>=start) & (trades["sell_date"] <= end)]
if trades.index.size < 1:
print(epoch,trades.index.size)
continue
else:
index = 0
captured_seats = 0
while captured_seats <= seats or index <= 20:
initial = float(cash/(seats + 1))
seat_trades = trades[trades["seat"] == index]
seat_trades.sort_values("sell_date",inplace=True)
if index > 20:
break
if captured_seats > seats:
break
if seat_trades.index.size > 0:
for delta in seat_trades["sell_delta"]:
initial = initial * (1 + delta)
stg = seat_trades.groupby("seat").mean().reset_index()[["adjclose","sell_price","seat","sell_delta"]]
for col in list(tse.columns):
if col == "__id":
continue
else:
stg[col] = epoch_dict[col]
# stg["pv"] = seat_trades.tail(1)["pv"].item()
stg["initial_pv"] = initial
stg["trades"] = seat_trades.index.size
stg["seat"] = captured_seats
stg["winrate"] = seat_trades[seat_trades["sell_delta"] > 0].index.size / seat_trades.index.size
initial = float(cash/(seats + 1))
analysis.append(stg)
index += 1
captured_seats += 1
continue
else:
index += 1
continue
strat_db.close()
a = pd.concat(analysis)
metric = "initial_pv"
index_cols = list(tse.columns)
index_cols.remove("_id")
tsmc = ""
apt = a.pivot_table(index=index_cols,columns="seat",values=metric)
if metric == "winrate":
apt[metric] = [statistics.mean([row[1][i] for i in range(seats) if not np.isnan(row[1][i])]) for row in apt.iterrows()]
else:
apt.fillna(cash/(seats+1),inplace=True)
apt[metric] = [sum([row[1][i] for i in range(seats+1) if i in apt.columns]) for row in apt.iterrows()]
apt["perf"] = 1 + (apt[metric] - cash) / cash
apt.reset_index(inplace=True)
apt.sort_values(metric,ascending=False).head(50)
if tsmc == "industry":
bf = []
for industry in apt["industry"].unique():
industry_apt = apt[apt["industry"] == industry].sort_values(metric,ascending=False).head(1)
bf.append(industry_apt)
bff = pd.concat(bf)
bff.sort_values(metric,ascending=False)
strat_db.connect()
epoch = apt.sort_values(metric,ascending=False).reset_index().iloc[0]["epoch"]
analysis = []
trades = strat_db.retrieve_data("{}_{}".format(suffix,int(epoch)))
trades = trades[(trades["date"]>=start) & (trades["sell_date"] <= end)]
strat_db.close()
print("trades:",len(trades))
top_epoch = tse[tse["epoch"]==epoch]
for col in top_epoch.columns:
print(col,":",top_epoch[col].item())
trades.sort_values("date")
trades["date"] = pd.to_datetime(trades["date"])
trades["sell_date"] = pd.to_datetime(trades["sell_date"])
dta = "date"
trades["week"] = [x.week for x in trades[dta]]
trades["quarter"] = [x.quarter for x in trades[dta]]
trades["dayofWeek"] = [x.weekday() for x in trades[dta]]
trades["month"] = [x.month for x in trades[dta]]
trades["year"] = [x.year for x in trades[dta]]
trades = trades[trades["sell_date"] <= end]
bench
prices = trades
daily = prices.groupby(["year","dayofWeek"]).mean().reset_index()
quarterly = prices.groupby(["year","quarter"]).mean().reset_index()
monthly = prices.groupby(["year","month"]).mean().reset_index()
weekly = prices.groupby(["year","week"]).mean().reset_index()
trades
timelines= []
cs = 0
index = 0
index = 0
print(seats)
while cs <= seats or index <= 20:
timeline = []
initial = float(cash/(seats + 1))
st = prices[prices["seat"] == index]
st.sort_values("sell_date",inplace=True)
if index > 20:
break
if cs > seats:
break
if st.index.size > 0:
for row in st.iterrows():
# for day in range((row[1]["sell_date"] - row[1]["date"]).days):
# timeline.append({"date":row[1]["date"] + timedelta(days=day),"seat":cs,"pv":initial})
initial = initial * (1 + row[1]["sell_delta"])
timeline.append({"date":row[1]["sell_date"],"seat":cs,"pv":initial})
timelines.append(pd.DataFrame(timeline))
initial = float(100/(seats + 1))
index+=1
cs+=1
continue
else:
index += 1
continue
ts = pd.concat(timelines)
tspivot = ts.pivot_table(index="date",columns="seat",values="pv")
tspivot.iloc[0].fillna(100/(seats+1),inplace=True)
cleaned = []
for i in range(tspivot.index.size):
row = tspivot.iloc[i]
for col in tspivot.columns:
if col == "date":
continue
else:
col_val = row[col].item()
if math.isnan(col_val):
row[col] = tspivot.iloc[i-1][col].item()
else:
continue
cleaned.append(row)
tspivot = pd.DataFrame(cleaned)
for i in range(seats+1):
if i not in tspivot.columns:
tspivot[i] = 100/(seats +1)
# tspivot = ts.pivot_table(index="date",columns="seat",values="pv")
tspivot["pv"] = [sum([row[1][i] for i in range(seats+1) if i in tspivot.columns]) for row in tspivot.iterrows()]
tspivot.reset_index(inplace=True)
tspivot
pvs = [100]
current_pv = 100
for i in range(1,tspivot.index.size):
replacement = current_pv/seats
current_date = tspivot.iloc[i].fillna(replacement)
current_pv =sum([current_date[c] for c in range(seats+1) if c in tspivot.columns])
pvs.append(current_pv)
tspivot["pv"] = pvs
bench.rename(columns={"Date":"index","Adj Close":"bench"},inplace=True)
bench["index"]= pd.to_datetime(bench["index"])
bench = bench[(bench["index"]>=start) & (bench["index"]<=end)]
bench["bench"] = (bench["bench"] - bench["bench"].iloc[0]) / bench["bench"].iloc[0]
tspivot["pv"] = (tspivot["pv"] - 100) / 100
tspivot = tspivot.merge(bench,on="index",how="left")
# +
fig, ax1 = plt.subplots()
color = 'tab:red'
ax1.set_xlabel('Date')
ax1.set_ylabel('price', color=color)
ax1.plot(tspivot["index"], tspivot["pv"], color=color)
ax1.tick_params(axis='y', labelcolor=color)
ax2 = ax1.twinx() # instantiate a second axes that shares the same x-axis
color = 'tab:blue'
# ax2.set_ylabel('bench', color=color) # we already handled the x-label with ax1
ax1.plot(bench["index"],bench["bench"], color=color)
# ax2.tick_params(axis='y', labelcolor=color)
fig.tight_layout() # otherwise the right y-label is slightly clipped
plt.figure(figsize=(32,18))
fig.tight_layout()
# -
normalized = []
for year in daily["year"].unique():
subset = daily[daily["year"] == year]
subset["sell_delta"] = normalize([subset["sell_delta"]])[0]
normalized.append(subset)
grouped = pd.concat(normalized).groupby("dayofWeek").mean().reset_index()
plt.plot(grouped["sell_delta"])
normalized = []
for year in quarterly["year"].unique():
try:
subset = quarterly[quarterly["year"] == year]
subset["sell_delta"] = normalize([subset["sell_delta"]])[0]
normalized.append(subset)
except:
continue
grouped = pd.concat(normalized).groupby("quarter").mean().reset_index()
plt.plot(grouped["sell_delta"])
normalized = []
for year in monthly["year"].unique():
try:
subset = monthly[monthly["year"] == year]
subset["sell_delta"] = normalize([subset["sell_delta"]])[0]
normalized.append(subset)
except:
continue
grouped = pd.concat(normalized).groupby("month").mean().reset_index()
plt.plot(grouped["sell_delta"])
normalized = []
for year in weekly["year"].unique():
try:
subset = weekly[weekly["year"] == year]
subset["sell_delta"] = normalize([subset["sell_delta"]])[0]
normalized.append(subset)
except:
continue
grouped = pd.concat(normalized).groupby("week").mean().reset_index()
plt.plot(grouped["sell_delta"])
ntrades = []
trades.sort_values("date",inplace=True)
for year in trades["year"].unique():
subset = trades[(trades["year"] == year)]
for week in subset["week"].unique():
ntrades.append(len(subset[(subset["week"] == week)]))
weekly["ntrades"]= ntrades
sp5.columns
trades["trade"] = 1
sp5["ticker"] = sp5["Symbol"]
ticker_yearly = trades.groupby(["year","ticker"]).sum().reset_index().merge(sp5, on="ticker",how="left")
yp = []
for year in ticker_yearly["year"].unique():
yp.append(ticker_yearly[ticker_yearly["year"] == year].sort_values("sell_delta",ascending=False))
yp_final = pd.concat(yp)
yp_final["aggregate_gain"] = [(1+(row[1]["sell_delta"] / row[1]["trade"])) ** row[1]["trade"] - 1 for row in yp_final.iterrows()]
yp_final.sort_values("aggregate_gain",ascending=False)[["year","Security","ticker","GICS Sector","sell_delta","trade","aggregate_gain"]].head(50)
ticker_analysis = trades.groupby("ticker").sum().reset_index().merge(sp5, on="ticker",how="left")
ticker_analysis["aggregate_gain"] = [(1+(row[1]["sell_delta"] / row[1]["trade"])) ** row[1]["trade"] - 1 for row in ticker_analysis.iterrows()]
ticker_analysis["average_gain"] = [(row[1]["sell_delta"] / row[1]["trade"]) for row in ticker_analysis.iterrows()]
ticker_analysis.sort_values("aggregate_gain",ascending=False)[["Security","ticker","sell_delta","trade","average_gain"]].head(50)
ticker_analysis = trades.merge(sp5, on="ticker",how="left").groupby("GICS Sector").sum().reset_index()
ticker_analysis["aggregate_gain"] = [(1+(row[1]["sell_delta"] / row[1]["trade"])) ** row[1]["trade"] - 1 for row in ticker_analysis.iterrows()]
ticker_analysis["average_gain"] = [(row[1]["sell_delta"] / row[1]["trade"]) for row in ticker_analysis.iterrows()]
ticker_analysis.sort_values("sell_delta",ascending=False)[["GICS Sector","sell_delta","trade","average_gain"]].head(50)
trades.sort_values("sell_delta",ascending=True).head(50).columns
| unity_backtest_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.6 64-bit
# metadata:
# interpreter:
# hash: a062be9dc8f0b4e802ee41733a00058b9f58e2e67b7919a84fe2c611795dfe04
# name: python3
# ---
# # Aggregation and Grouping
#
# An essential peice of analysis of large data is efficient summarization: computing aggregations like `sum()`, `mean()`, `median()`, `min()` and `max()`, in which a single number gives insight into the nature of a potentially large dataset. In this section, we'll explore aggreagtions in Pandas, from simple operations akin to what we've seen on NumPy arrays, to more sophisticated operations based on the concept of a `groupby`.
#
# For convenience, we'll use the same `display`magic function that we've seen in previous sections:
# +
import numpy as np
import pandas as pd
class display(object):
"""Display HTML representation of multiple objects"""
template = """<div style="float: left; padding: 10px;">
<p style='font-family:"Courier New", Courier, monospace'>{0}</p>{1}
</div>"""
def __init__(self, *args):
self.args = args
def _repr_html_(self):
return '\n'.join(self.template.format(a, eval(a)._repr_html_())
for a in self.args)
def __repr__(self):
return '\n\n'.join(a + '\n' + repr(eval(a))
for a in self.args)
# -
# ## Planets Data
#
# Here we wil luse the Planets dataset, available via the seaborn package. It gives information on planets that astronomers have discovered around other stars (known *as extrasolar planets* or *exoplanets* for short).
import seaborn as sns
planets = sns.load_dataset('planets')
planets.shape
planets.head()
# This has some details on the $1000+$ extrasolar planets discovered up to 2014.
# ## Simple Aggregation in Pandas
#
# Earlier, we explored some of the data aggregations available for NumPy arrays. As with a one-dimensional NumPy array, for a Pandas `Series` the aggregates return a single value:
rng = np.random.RandomState(42)
ser = pd.Series(rng.rand(5))
ser
ser.sum()
ser.mean()
# For a `DataFrame,` by default the aggregates return results within each column:
df = pd.DataFrame({'A': rng.rand(5),
'B': rng.rand(5)})
df
df.mean()
df.mean(axis=1)
# Pandas `Series` and `DataFrame`s include all of the common aggregates mentioned earlier; in addition, there is a convenience method `describe()` that computes several common aggregates for each column and returns the result. Let's use this on the Planets data, for now dropping rows with missing values:
planets.dropna().describe()
# This can be a useful way to begin understanding the overall properties of a dataset. For example, we see in the `year` column that although exoplanets were discovered as far back as 1989, half fo all known exoplanets were not discovered until 2010 or after. This is largely thanks to the *Kepler* mission, which is a space-based telescope specifically designed for finding eclipsing planets around other stars.
#
# The following table summarizes other built-in Pandas aggregations:
#
# ```
# Aggregation Description
# count() Total number of items
# first(), last() First and last item
# mean(), median() Mean and median
# min(), max() Minimum and maximum
# std(), var() Standard deviation and variance
# mad() Mean absolute deviation
# prod() Product of all items
# sum() Sum of all items
#
# ```
#
# These are all methods of `DataFrame` and `Series` objects.
#
# To go deeper into the data, however, simple aggregates are often not enough. The next level of data summarization is the `groupby` operation, which allows you to quickly and efficiently compute aggregates on subsets of data.
# ## GroupBy: Split, Apply, Combine
#
# Simple aggregations can give you a flavor of your dataset, but often we would prefer to aggregate conditionally on some label or index: this is implemented in the so-called `groupby` operation. The name "groub by" comes from a command in the SQL database language, but it is perhaps more illuminative to think of it in the terms first coniend by <NAME> of Rstats fame: `split, apply, combine`.
# ### Split, apply, combine
#
# This makes clear what `groupby` accomplished:
#
# - The *split* step involves breaking up and grouping a `DataFrame` depending on the value of the specified key.
# - The *apply* step involves computing some function, usually an aggregate, transformation, or filtering, whithin the individual groups.
# - The *combine* step merges the results of these operations into an output array.
#
# While this could certainly be done manually using some combination of the masking, aggregation, and mergin commands covered earlier, an important realization is that the *intermediate splits do not need to be explicitly intantiated*. Rather, the `GroupBy` can (often) do this in a single pass over the data, updating the sum, mean, count, min, or other aggregate for each group along the way. The power of the `GroupBy` is that is abstracts away these steps: the user need not think about *how* the computation is done under the hood, but rahter thinks about the *operation as a whole*.
#
# As a concrete example, let's take a look at using Pandas for the computation shown in this diagram. We'll star tby creating the input `DataFrame`:
df = pd.DataFrame({'key': list('ABCABC'),
'data': range(6)}, columns=['key', 'data'])
df
# The most basic split-apply-combine operation can be computed with the `groupby()` method of `DataFrame`s, passing the name of the desired key column:
df.groupby('key')
# The `sum()` method is just one possibility here; you can apply virtually any common Pandas or NumPy aggregation function, as well as virtually any valid `DataFrame` operation, as we will see in the following discussion.
# ### The GroupBy object
#
# The `GroupBy` object is a very flexible abstraction. In many ways, you can simply treat it as if it's a collection of `DataFrame`s, and it does the difficult things under the hood. Let's see some examples using the Planets data.
# ### Column indexing
#
# The `GroupBy` object supports column indexing in the same way as the `DataFrame`, and returns a modified `GroupBy` object.
planets.groupby('method')
planets.groupby('method')['orbital_period']
# Here we've selected a particular `Series` group from the original `DataFrame` group by reference to its column name. As with the `GroupBy` object, no computation is done until we call some aggregate on the object:
planets.groupby('method')['orbital_period'].median()
# That gives an idea of the general scale of orbital periods (in days) that each method is sensitive to.
# #### Iteration over groups
#
# The `GroupBy` object supports direct iteration over the groups, returning each group as a `Series` or `DataFrame`:
for (method, group) in planets.groupby('method'):
print("{0:30s} shape={1}".format(method, group.shape))
# This can be useful for doing certain things manually, though it is often much faster to sue the built-in `apply` functionality, which we will discuss momentarily.
# #### Dispatch methods
#
# Thorugh some Python class magic, any method not explicitly implemented by the `GroupBy` object will be passed through and called on the groups, whether they are `DataFrame` or `Series` objects. For example, you can sue the `describe()` method of `DataFrame`s to perform a set of aggreagtions that describe each grop in the data:
planets.groupby('method')['year'].describe().unstack()
# Looking at this table help us to better understand the data: for example, the vast majority of planets have been discovered by the Radial Velocity and transit methods, though the latter only became common (due to new, more accurate telescopes) in the last decade. The newest methods seem to be Transit Timing Variation on and Orbital Brightness Modulation, which were not used to discover a new planet until 2011.
#
# This is just one example of the utiliy of dispatch methods. Notice that they are applied to *each individual group*, and the results are the combined within `GroupBy` and returned. Again, any valid `DataFrame / Series` method can be used on the corresponding `GroupBy` object, which allows for some very flexible and powerful operations.
# ### Aggregate, filter, transform, apply
#
# The preceding discussion focused on aggregation of the combine operation, but there are more options available. In particular, `GroupBy` objects have `aggregate()`, `filter()`, `transform()`, and `apply()` methods that efficiently implementa a variety of useful operations before combining the grouped data.
#
# For the purpose of the following subsections, we'll use this `DataFrame`:
rng = np.random.RandomState(0)
df = pd.DataFrame({'key': list('ABCABC'),
'data1': range(6),
'data2': rng.randint(0, 16, 6)},
columns= ['key', 'data1', 'data2'])
df
# #### Aggregation
#
# We're now familiar with `GroupBy` aggregations with `sum()`, `median()`, and the like, but the `aggregate()` method allows for even more flexibility. It can take a string, a function, or alist thereof, and compute all the aggregates at once. Here is a quick example combining all these:
df.groupby('key').aggregate(['min', np.median, max])
# Another useful pattern is to pass a dictionary mapping column names to operations to be applied on that column:
df.groupby('key').aggregate({'data1': 'min', 'data2': 'max'})
# #### Filtering
#
# A filtering operation allows you to drop data based on the group properties. For example , we might want to keep all gorups in which the standard deviation is larger than some critical value:
# +
def filter_func(x):
return x['data2'].std() > 5
display('df', "df.groupby('key').std()", "df.groupby('key').filter(filter_func)")
# -
# The filter function should return a Boolean value specifying whether the gropu passed the filtering. There because group A does not have a standard deviation greater than 4, it is dropped from the result.
# #### Transformation
#
# While aggregation must return a reduced version of the data, transformation can return some transformed version of the full data to recombine. For such a transformation, the output is the same shape as the input. A common example is to center the data by subtracting the group-wise mean:
df.groupby('key').transform(lambda x: x- x.mean())
# #### The apply() method
#
# The `apply()` method lets you apply an arbitrary function to the group results. The function should take a `DataFrame`, and return either a Pandas object (e.g, `DataFrame`, `Series`) or a scalar; the combine opration will be tailored to the type of output returned.
#
# For example, here is an `apply()` that normalizes the first column by the sum of the second:
# +
def norm_by_data2(x):
# x is a DataFrame of group values
x['data1'] /= x['data2'].sum()
return x
display('df', "df.groupby('key').apply(norm_by_data2)")
# -
# `apply()` within a `Groupby` is quite flexible: the only criterion is that the function takes a `DataFrame` and returns a Pandas object or scalar; what you do in the middle is up to you.
# ### Specifying the split key
#
# In the simple examples presented before, we split the `DataFrame` on a single column name. This is just one of many options by which the groups can be defined, and we'll go through some other options for group specification here.
# #### A list, array, series, or index providing the grouping keys
#
# They key can be any series or list with a length matching that of the `DataFrame`. For example:
L = [0, 1, 0, 1, 2, 0]
display('df', 'df.groupby(L).sum()')
# Of course, this means there's another, more verbose way of accomplishing the `df.grouby('key')` from before:
display('df', "df.groupby(df['key']).sum()")
# #### A dictionary or series mapping index to group
#
# Another method is to provide a dictionary that maps index values to the group keys:
df2 = df.set_index('key')
mapping = {'A': 'vowel', 'B': 'consonant', 'C': 'consonant'}
display('df2', 'df2.groupby(mapping).sum()')
# #### Any Python function
#
# Similar to mapping, you can pass any Python function that will input the index value and output the group:
display('df2', 'df2.groupby(str.lower).mean()')
# #### A list of valid keys
#
# Further, any of the preceding key choices can be combined to group on a multi-index:
df2.groupby([str.lower, mapping]).mean()
# ### Grouping example
#
# As an example of this, in a couple lines of Python code we can put all these togehter and count discovered planets by method and by decade:
# +
decade = 10 * (planets['year'] // 10)
decade = decade.astype(str) + 's'
decade.name = 'decade'
decade
planets.groupby(['method', decade])['number'].sum().unstack().fillna(0)
# -
# This shows the power of combining many of the operations we've discussed up to this point when looking at realistic datasets. We immediately gain a coarse understanding of then and how planets have been discovered over the past several decades!
#
# Here I would suggest digging into these few lines of code, and evaluating the individual steps to make sure you understand exactly what they are doing to the result. It's certainly a somewhat complicated example, but understanding these pices will give you the means to similarly explore your own data.
| pandas/aggregation_and_grouping.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .js
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Javascript (Node.js)
// language: javascript
// name: javascript
// ---
// # Plotly examples
// ## [line-and-scatter](https://plotly.com/javascript/line-and-scatter/)
var ds = require('dstools');
// +
var trace1 = {
x: [1, 2, 3, 4],
y: [10, 15, 13, 17],
mode: 'markers',
type: 'scatter'
};
var trace2 = {
x: [2, 3, 4, 5],
y: [16, 5, 11, 9],
mode: 'lines',
type: 'scatter'
};
var trace3 = {
x: [1, 2, 3, 4],
y: [12, 9, 15, 12],
mode: 'lines+markers',
type: 'scatter'
};
var data = [trace1, trace2, trace3];
var layout = {};
ds.Collection().plotly(data, layout).show();
// +
var trace1 = {
x: [1, 2, 3, 4, 5],
y: [1, 6, 3, 6, 1],
mode: 'markers+text',
type: 'scatter',
name: 'Team A',
text: ['A-1', 'A-2', 'A-3', 'A-4', 'A-5'],
textposition: 'top center',
textfont: {
family: 'Raleway, sans-serif'
},
marker: { size: 12 }
};
var trace2 = {
x: [1.5, 2.5, 3.5, 4.5, 5.5],
y: [4, 1, 7, 1, 4],
mode: 'markers+text',
type: 'scatter',
name: 'Team B',
text: ['B-a', 'B-b', 'B-c', 'B-d', 'B-e'],
textfont : {
family:'Times New Roman'
},
textposition: 'bottom center',
marker: { size: 12 }
};
var data = [ trace1, trace2 ];
var layout = {
xaxis: {
range: [ 0.75, 5.25 ]
},
yaxis: {
range: [0, 8]
},
legend: {
y: 0.5,
yref: 'paper',
font: {
family: 'Arial, sans-serif',
size: 20,
color: 'grey',
}
},
title:'Data Labels on the Plot'
};
var data = [trace1, trace2, trace3];
ds.Collection().plotly(data, layout).show();
// -
// ## [Pie Charts in JavaScript](https://plotly.com/javascript/pie-charts)
// ## [basic-pie-chart](https://plotly.com/javascript/pie-charts/#basic-pie-chart)
// +
var data = [{
values: [19, 26, 55],
labels: ['Residential', 'Non-Residential', 'Utility'],
type: 'pie'
}];
var layout = {
height: 400,
width: 500
};
ds.Collection().plotly(data, layout).show();
// -
// ### [Donut Chart](https://plotly.com/javascript/pie-charts/#donut-chart)
// +
var data = [{
values: [16, 15, 12, 6, 5, 4, 42],
labels: ['US', 'China', 'European Union', 'Russian Federation', 'Brazil', 'India', 'Rest of World' ],
domain: {column: 0},
name: 'GHG Emissions',
hoverinfo: 'label+percent+name',
hole: .4,
type: 'pie'
},{
values: [27, 11, 25, 8, 1, 3, 25],
labels: ['US', 'China', 'European Union', 'Russian Federation', 'Brazil', 'India', 'Rest of World' ],
text: 'CO2',
textposition: 'inside',
domain: {column: 1},
name: 'CO2 Emissions',
hoverinfo: 'label+percent+name',
hole: .4,
type: 'pie'
}];
var layout = {
title: 'Global Emissions 1990-2011',
annotations: [
{
font: {
size: 20
},
showarrow: false,
text: 'GHG',
x: 0.17,
y: 0.5
},
{
font: {
size: 20
},
showarrow: false,
text: 'CO2',
x: 0.82,
y: 0.5
}
],
height: 400,
width: 600,
showlegend: false,
grid: {rows: 1, columns: 2}
};
ds.Collection().plotly(data, layout).show();
// -
// ## [bar-charts](https://plotly.com/javascript/bar-charts/)
// +
var trace1 = {
x: ['giraffes', 'orangutans', 'monkeys'],
y: [20, 14, 23],
name: '<NAME>',
type: 'bar'
};
var trace2 = {
x: ['giraffes', 'orangutans', 'monkeys'],
y: [12, 18, 29],
name: '<NAME>',
type: 'bar'
};
var data = [trace1, trace2];
var layout = {barmode: 'group'};
ds.Collection().plotly(data, layout).show();
| Plotly-NodeJS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:tensorflow35]
# language: python
# name: conda-env-tensorflow35-py
# ---
# +
from keras import backend as K
from ImageLoader import TextImageLoader
from CRNN import *
import os
import matplotlib.pyplot as plt
import numpy as np
from parameter import *
from utils import *
import json
# -
# './IIIT5K/train/' is used for testing.<br>
# Because there is more data in the test image file, the test file iamges was used for training.
# +
val_img_file_path = './IIIT5K/train/'
val_label_file_path = './IIIT5K/train_label/'
val_img_generator_params = {
'img_path' : val_img_file_path,
'label_path' : val_label_file_path,
'img_w' : params['img_w'],
'img_h' : params['img_h'],
'batch_size' : 4,
'downsample_factor' : params['downsample_factor'],
'max_text_len' : params['max_text_len'],
'do_shuffle' : params['do_shuffle']
}
# -
val_dataloader = TextImageLoader(**val_img_generator_params)
# # TextImageLoader use
gener = val_dataloader.generator()
inputs, outputs = next(gener)
inputs.keys()
outputs.keys()
img1 = inputs['the_input'][0]
plt.imshow(np.squeeze(img1))
# ### the_input is image data value
inputs['the_input'].shape
inputs['the_labels'].shape
# ### Convert letters to numeric labels
inputs['the_labels']
# ### input length for LSTM's input
inputs['input_length']
# ### length of text
inputs['label_length']
crnn_params = {
'img_w' : params['img_w'],
'img_h' : params['img_h'],
'num_classes' : params['num_classes'],
'max_text_len' : params['max_text_len'],
'drop_rate' : params['drop_rate'],
'weight_decay' : params['weight_decay'],
'learning_rate' : 1e-3,
'training' : True
}
crnn_model = CRNN(**crnn_params)
try:
crnn_model.saved_model_use('./save_model/crnn/crnn_weights.hdf5')
except:
raise Exception("No weight!")
test_dir =val_img_file_path
test_imgs = os.listdir(val_img_file_path)
test_labels = os.listdir(val_label_file_path)
# # Test images
# +
total = 0
acc = 0
for k, test_img in enumerate(test_imgs):
net_out_value = crnn_model.predict(test_dir + test_img)
pred_texts= decode_label(net_out_value)
jstring = open(val_label_file_path + os.path.splitext(test_img)[0] + '.json', "r").read()
jstring = json.loads(jstring)
chars = jstring['image_label']
if pred_texts.lower() == chars.lower():
acc += 1
total += 1
print(k,'/',len(test_imgs)," ACC : ", acc / total)
print('===')
print('Predicted: %s / True: %s' % (pred_texts.lower(), chars.lower()))
print('===')
print("TOTAL ACC : ", acc / total)
# -
# ### result
# +
test_img = val_dataloader.total_img_list[0]
img_ex = val_dataloader.get_image(test_dir + test_img, val_dataloader.img_w, val_dataloader.img_h)
plt.imshow(img_ex, cmap='gray')
plt.title('Grayscale Image')
plt.show()
# +
net_out_value = crnn_model.predict(test_dir + test_img)
pred_texts= decode_label(net_out_value)
jstring = open(val_label_file_path + os.path.splitext(test_img)[0] + '.json', "r").read()
jstring = json.loads(jstring)
chars = jstring['image_label']
# -
plt.imshow(img_ex, cmap='gray')
plt.title('Predicted: %s / True: %s' % (pred_texts.lower(), chars.lower()))
plt.show()
# ### show multi results
for i in range(4):
test_img = val_dataloader.total_img_list[i]
img_ex = val_dataloader.get_image(test_dir + test_img, val_dataloader.img_w, val_dataloader.img_h)
net_out_value = crnn_model.predict(test_dir + test_img)
pred_texts= decode_label(net_out_value)
jstring = open(val_label_file_path + os.path.splitext(test_img)[0] + '.json', "r").read()
jstring = json.loads(jstring)
chars = jstring['image_label']
plt.imshow(img_ex, cmap='gray')
plt.title('Predicted: %s / True: %s' % (pred_texts.lower(), chars.lower()))
plt.show()
| using_CRNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from selenium import webdriver
from bs4 import BeautifulSoup
from urllib import parse
from urllib import request
import time
import os
from io import BytesIO
import pandas as pd
import win32clipboard
from PIL import Image
import pyperclip
import win32com.client as client
def send_to_clipboard(clip_type, data):
win32clipboard.OpenClipboard()
win32clipboard.EmptyClipboard()
win32clipboard.SetClipboardData(clip_type, data)
win32clipboard.CloseClipboard()
hwp = client.gencache.EnsureDispatch("HWPFrame.HwpObject")
hwp.RegisterModule('FilePathCheckDLL', 'Filepath')
hwp.Open(os.getcwd()+'/datas/conti.hwp')
hwp.HAction.Run("SelectAll")
i = 1
for link in links :
if i % 2 :
res = request.urlopen(link).read()
image = Image.open(BytesIO(res))
image = image.resize([700,900])
output = BytesIO()
image.convert("RGB").save(output, "BMP")
data = output.getvalue()[14:]
send_to_clipboard(win32clipboard.CF_DIB, data)
hwp.HAction.Run("Paste")
i += 1
i = 1
for link in links :
if i % 2 == 0 :
res = request.urlopen(link).read()
image = Image.open(BytesIO(res))
image = image.resize([700,900])
output = BytesIO()
image.convert("RGB").save(output, "BMP")
data = output.getvalue()[14:]
send_to_clipboard(win32clipboard.CF_DIB, data)
hwp.HAction.Run("Paste")
i += 1
# +
querys = pd.read_csv('datas/querys.txt', header=None)
driver = webdriver.Chrome()
links = []
for query in querys.values:
query = query[0]
enc = parse.quote(query + ' 악보')
url = f'https://www.google.com/search?q={enc}&tbm=isch&'
driver.get(url)
images = driver.find_elements_by_tag_name("img")
for image in images:
if list(image.size.values()) != [50, 50] :
image.click()
break
time.sleep(1)
link = driver.find_elements_by_css_selector('div.v4dQwb')[0]
links.append(link.find_element_by_class_name('n3VNCb').get_attribute('src'))
driver.quit()
# -
links
link = driver.find_elements_by_css_selector('div.v4dQwb')[0]
link.find_element_by_class_name('n3VNCb').get_attribute('src')
a
images = driver.find_elements_by_tag_name("img")
len(images)
images = driver.find_elements_by_tag_name("img")
url = 'https://www.google.com/search?q=%EC%8B%9C%EC%84%A0+%EC%95%85%EB%B3%B4&tbm=isch&#imgrc=qj6m0eEkeSSV7M'
url = driver.current_url
url
response = requests.get(url)
dom = BeautifulSoup(response.content, "html.parser")
dom.select('c-wiz > div > div > div > div > div')
| selenium.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.image as mpimg
import matplotlib.pyplot as plt
import numpy as np
import cv2
import glob
from skimage.feature import hog
from skimage import color, exposure
# %matplotlib inline
# images are divided up into vehicles and non-vehicles
car_images_udacity = glob.glob(r'C:\Users\mohar\Desktop\Vinod\vehicles\*\*.png')
notcar_images_udacity = glob.glob(r'C:\Users\mohar\Desktop\Vinod\non-vehicles\*\*.png')
cars = []
notcars = []
for image in car_images_udacity:
cars.append(image)
for image in notcar_images_udacity:
notcars.append(image)
# Define a function to return some characteristics of the dataset
def data_look(car_list, notcar_list):
data_dict = {}
# Define a key in data_dict "n_cars" and store the number of car images
data_dict["n_cars"] = len(car_list)
# Define a key "n_notcars" and store the number of notcar images
data_dict["n_notcars"] = len(notcar_list)
# Read in a test image, either car or notcar
test_img = mpimg.imread(car_list[0])
# Define a key "image_shape" and store the test image shape 3-tuple
data_dict["image_shape"] = test_img.shape
# Define a key "data_type" and store the data type of the test image.
data_dict["data_type"] = test_img.dtype
# Return data_dict
return data_dict
data_info = data_look(cars, notcars)
print('Your function returned a count of',
data_info["n_cars"], ' cars and',
data_info["n_notcars"], ' non-cars')
print('of size: ',data_info["image_shape"], ' and data type:',
data_info["data_type"])
car_images_crowdai = glob.glob(r'C:\Users\mohar\Desktop\Vinod\resized\*.jpg')
# load image path randomly from resized images to augment car class
np.random.seed(0) # generate same random numbers everytime to avoid issues due to variation in training data later
for i in range(len(notcars) - len(cars)):
cars.append(car_images_crowdai[np.random.randint(0, len(car_images_crowdai))])
data_info = data_look(cars, notcars)
print('Your function returned a count of',
data_info["n_cars"], ' cars and',
data_info["n_notcars"], ' non-cars')
print('of size: ',data_info["image_shape"], ' and data type:',
data_info["data_type"])
# -
print(cars[200])
print(cars[-100])
# +
rgb_png_mpimg = mpimg.imread(cars[200]) # reads png color pixels in RGB between 0 and 1, dtype = float32
rgb_jpg_mpimg = mpimg.imread(cars[-100]) # reads jpg color pixels in RGB between 0 and 255, dtype = uint8
bgr_png_cv2 = cv2.imread(cars[200]) # reads png color pixels in BGR between 0 and 255, dtype = uint8
bgr_jpg_cv2 = cv2.imread(cars[-100]) # reads jpg color pixels in BGR between 0 and 255, dtype = uint8
rgb_png_cv2 = bgr_png_cv2[...,::-1]
rgb_jpg_cv2 = bgr_jpg_cv2[...,::-1]
print(cv2.cvtColor(rgb_jpg_mpimg, cv2.COLOR_RGB2LUV)[0][0][0])
print(np.dtype(cv2.cvtColor(rgb_jpg_mpimg, cv2.COLOR_RGB2LUV)[0][0][0]))
| Term1-Computer-Vision-and-Deep-Learning/Project5-Vehicle-Detection-and-Tracking/Explore mpimg & cv2 imread.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ___
#
# <a href='http://www.pieriandata.com'><img src='../Pierian_Data_Logo.png'/></a>
# ___
# <center><em>Copyright <NAME></em></center>
# <center><em>For more information, visit us at <a href='http://www.pieriandata.com'>www.pieriandata.com</a></em></center>
# # RNN Example for Time Series
# + jupyter={"outputs_hidden": true}
import pandas as pd
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
# -
# ## Data
# https://fred.stlouisfed.org/series/S4248SM144NCEN
# + jupyter={"outputs_hidden": true}
df = pd.read_csv('../Data/Alcohol_Sales.csv',index_col='DATE',parse_dates=True)
df.index.freq = 'MS'
# -
df.head()
# + jupyter={"outputs_hidden": true}
df.columns = ['Sales']
# -
df.plot(figsize=(12,8))
# + jupyter={"outputs_hidden": true}
from statsmodels.tsa.seasonal import seasonal_decompose
# -
results = seasonal_decompose(df['Sales'])
results.observed.plot(figsize=(12,2))
results.trend.plot(figsize=(12,2))
results.seasonal.plot(figsize=(12,2))
results.resid.plot(figsize=(12,2))
# ## Train Test Split
len(df)
325-12
# + jupyter={"outputs_hidden": true}
train = df.iloc[:313]
test = df.iloc[313:]
# -
len(test)
# ## Scale Data
# + jupyter={"outputs_hidden": true}
from sklearn.preprocessing import MinMaxScaler
# + jupyter={"outputs_hidden": true}
scaler = MinMaxScaler()
# -
# IGNORE WARNING ITS JUST CONVERTING TO FLOATS
# WE ONLY FIT TO TRAININ DATA, OTHERWISE WE ARE CHEATING ASSUMING INFO ABOUT TEST SET
scaler.fit(train)
# + jupyter={"outputs_hidden": true}
scaled_train = scaler.transform(train)
scaled_test = scaler.transform(test)
# -
# # Time Series Generator
#
# This class takes in a sequence of data-points gathered at
# equal intervals, along with time series parameters such as
# stride, length of history, etc., to produce batches for
# training/validation.
#
# #### Arguments
# data: Indexable generator (such as list or Numpy array)
# containing consecutive data points (timesteps).
# The data should be at 2D, and axis 0 is expected
# to be the time dimension.
# targets: Targets corresponding to timesteps in `data`.
# It should have same length as `data`.
# length: Length of the output sequences (in number of timesteps).
# sampling_rate: Period between successive individual timesteps
# within sequences. For rate `r`, timesteps
# `data[i]`, `data[i-r]`, ... `data[i - length]`
# are used for create a sample sequence.
# stride: Period between successive output sequences.
# For stride `s`, consecutive output samples would
# be centered around `data[i]`, `data[i+s]`, `data[i+2*s]`, etc.
# start_index: Data points earlier than `start_index` will not be used
# in the output sequences. This is useful to reserve part of the
# data for test or validation.
# end_index: Data points later than `end_index` will not be used
# in the output sequences. This is useful to reserve part of the
# data for test or validation.
# shuffle: Whether to shuffle output samples,
# or instead draw them in chronological order.
# reverse: Boolean: if `true`, timesteps in each output sample will be
# in reverse chronological order.
# batch_size: Number of timeseries samples in each batch
# (except maybe the last one).
from keras.preprocessing.sequence import TimeseriesGenerator
# + jupyter={"outputs_hidden": true}
scaled_train
# -
# define generator
n_input = 2
n_features = 1
generator = TimeseriesGenerator(scaled_train, scaled_train, length=n_input, batch_size=1)
len(scaled_train)
len(generator) # n_input = 2
# + jupyter={"outputs_hidden": true}
scaled_train
# -
# What does the first batch look like?
X,y = generator[0]
print(f'Given the Array: \n{X.flatten()}')
print(f'Predict this y: \n {y}')
# Let's redefine to get 12 months back and then predict the next month out
n_input = 12
generator = TimeseriesGenerator(scaled_train, scaled_train, length=n_input, batch_size=1)
# What does the first batch look like?
X,y = generator[0]
print(f'Given the Array: \n{X.flatten()}')
print(f'Predict this y: \n {y}')
# ### Create the Model
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import LSTM
# + jupyter={"outputs_hidden": true}
# define model
model = Sequential()
model.add(LSTM(100, activation='relu', input_shape=(n_input, n_features)))
model.add(Dense(1))
model.compile(optimizer='adam', loss='mse')
# -
model.summary()
# + jupyter={"outputs_hidden": true}
# fit model
model.fit_generator(generator,epochs=50)
# -
model.history.history.keys()
loss_per_epoch = model.history.history['loss']
plt.plot(range(len(loss_per_epoch)),loss_per_epoch)
# ## Evaluate on Test Data
first_eval_batch = scaled_train[-12:]
first_eval_batch
first_eval_batch = first_eval_batch.reshape((1, n_input, n_features))
model.predict(first_eval_batch)
scaled_test[0]
# Now let's put this logic in a for loop to predict into the future for the entire test range.
#
# ----
# + jupyter={"outputs_hidden": true}
test_predictions = []
first_eval_batch = scaled_train[-n_input:]
current_batch = first_eval_batch.reshape((1, n_input, n_features))
# -
current_batch.shape
current_batch
np.append(current_batch[:,1:,:],[[[99]]],axis=1)
# **NOTE: PAY CLOSE ATTENTION HERE TO WHAT IS BEING OUTPUTED AND IN WHAT DIMENSIONS. ADD YOUR OWN PRINT() STATEMENTS TO SEE WHAT IS TRULY GOING ON!!**
# +
# Holds the predections
test_predictions = []
# last n_input points from the training dataset
first_eval_batch = scaled_train[-n_input:]
# Reshaping it to the format RNN expects
current_batch = first_eval_batch.reshape((1, n_input, n_features))
# How far to predict?
for i in range(len(test)):
# get prediction 1 time stamp ahead ([0] is for grabbing just the number instead of [array])
current_pred = model.predict(current_batch)[0]
# store prediction
test_predictions.append(current_pred)
# update batch to now include prediction and drop first value
current_batch = np.append(current_batch[:,1:,:],[[current_pred]],axis=1)
# -
test_predictions
scaled_test
# ## Inverse Transformations and Compare
true_predictions = scaler.inverse_transform(test_predictions)
true_predictions
test
# IGNORE WARNINGS
test['Predictions'] = true_predictions
test
test.plot(figsize=(12,8))
# # Saving and Loading Models
model.save('my_rnn_model.h5')
# ## load a model
from keras.models import load_model
new_model = load_model('my_rnn_model.h5')
new_model.summary()
| tsa/jose/TSA_COURSE_NOTEBOOKS/07-Deep-Learning-Models/01-RNN-Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import seaborn as sns
import warnings
warnings.filterwarnings('ignore')
matriz = pd.read_csv("matrizajustada.csv")
matriz.drop(columns="Unnamed: 0",inplace=True)
matriz.head()
# +
#sns.pairplot(matriz)
# -
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
from sklearn.neural_network import MLPClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
from sklearn import svm
from sklearn.naive_bayes import BernoulliNB
from sklearn.ensemble import GradientBoostingClassifier
import itertools
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix
# %matplotlib inline
from sklearn.metrics import classification_report
import seaborn as sns
X = matriz.drop("practica", axis = 1).values
y = matriz["practica"].values
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
clf = MLPClassifier()
clf.fit(X_train, y_train)
y_predicted = clf.predict(X_test)
len(matriz)
print(classification_report(y_predicted, y_test))
#["respons","gestion","equip","ingeni","inform","industrial","offic","excel","diseñ","analisis"]
X_test_manual = [1,0,0,0,1,0,0,0,1,0]
y_predicted_manual = clf.predict([X_test_manual])
y_predicted_manual[0]
| ModeloPredictivo2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # AI-LAB SESSION 5: Deep Reinforcement Learning
#
# In this lesson we will use the CartPole environment and we will see how to create and work with a neural network using Kears on top of Tensorflow.
#
# ## CartPole
# The environment used is **CartPole** (taken from the book of Sutton and Barto as visible in the figure)
#
# 
#
# A pole is attached by an un-actuated joint to a cart, which moves along a frictionless track. The system is controlled by applying a force of +1 or -1 to the cart. The pendulum starts upright, and the goal is to prevent it from falling over. A reward of +1 is provided for every timestep that the pole remains upright. The episode ends when the pole is more than 15 degrees from vertical, or the cart moves more than 2.4 units from the center.
# +
import os, sys, tensorflow.keras, random, numpy
module_path = os.path.abspath(os.path.join('../tools'))
if module_path not in sys.path:
sys.path.append(module_path)
import gym, envs
from utils.ai_lab_functions import *
from timeit import default_timer as timer
from tqdm import tqdm as tqdm
from collections import deque
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout
from tensorflow.keras.optimizers import Adam
# -
# The **state** of environment is represented as a tuple of 4 values:
# - *Cart Position* range from -4.9 to 4.8
# - *Cart Velocity* range from -inf to +inf
# - *Pole Angle* range from -24 deg to 24 deg
# - *Pole Velocity* range from -inf to +inf
#
# The **actions** allowed in the environment are 2:
# - *action 0*: push cart to left
# - *action 1*: push cart to right
#
# The **reward** is 1 for every step taken, including the termination step.
# +
env = gym.make("CartPole-v1")
state = env.reset()
print("STARTING STATE: {}".format(state))
print("\tCart Position: {}\n\tCart Velocity {}\n\tPole Angle {} \n\tPole Velocity {}".format(state[0], state[1], state[2], state[3]))
print("\nPOSSIBLE ACTIONS: ", env.action_space.n)
# -
# Finally, we still have the standard functionalities of a Gym environment:
# - step(action): the agent performs action from the current state. Returns a tuple (new_state, reward, done, info) where:
# - new_state: is the new state reached as a consequence of the agent's last action
# - reward: the reward obtained by the agent in this step
# - done: True if the episode is terminal, False otherwise
# - info: not used, you can safely discard it
#
# - reset(): the environment is reset and the agent goes back to the starting position. Returns the initial state id
# ## Neural Network with Kears
# **Keras** is an open-source neural-network library written in Python. It is capable of running on top of TensorFlow, Microsoft Cognitive Toolkit, R, Theano, or PlaidML. Designed to enable fast experimentation with deep neural networks, it focuses on being user-friendly, modular, and extensible.
#
# 
#
# With kears you can easly create a neural network with the **Sequential** module. Before training a neural netowrk you must compile it, selecting the loss function and the optimizer, in our experiment we will use the *mean_squared_error* for the loss function and the *adam* optimizer, that is a standard configuration for a DQN problem.
# +
input_layer = 3
layer_size = 5
output_layer = 2
model = Sequential()
model.add(Dense(layer_size, input_dim=input_layer, activation="relu")) #input layer + hidden layer #1
model.add(Dense(layer_size, activation="relu")) #hidden layer #2
model.add(Dense(layer_size, activation="relu")) #hidden layer #3
model.add(Dense(layer_size, activation="relu")) #hidden layer #4
model.add(Dense(layer_size, activation="relu")) #hidden layer #5
model.add(Dense(output_layer, activation="linear")) #output layer
model.compile(loss="mean_squared_error", optimizer='adam') #loss function and optimzer definition
# -
# In Keras you can compute the output of a network with the **predict** function, that requires as input the values of the input layer nodes and returns the corresponding values of the output layer.
input_network = [random.uniform(0, 1), random.uniform(0, 1), random.uniform(0, 1)]
output_network = model.predict(np.array([input_network]))
print("Input Network: {}".format(input_network))
print("Network Prediction: {}".format(output_network[0]))
# To train a network in Keras we must use the function **fit**, that take as input:
# - *input*: the input of the network that we are interested to train
# - *expected_output*: the output that we consider correct
# - *epochs*: the number of iteration for the backpropagation (in DQN this value is always 1).
# +
input_network = [random.uniform(0, 1), random.uniform(0, 1), random.uniform(0, 1)]
expected_output = [0, 0]
print("Prediction 'before' training:")
print(model.predict(np.array([input_network])))
model.fit(np.array([input_network]), np.array([expected_output]), epochs=1000, verbose=0)
print("\nPrediction 'after' training:")
print(model.predict(np.array([input_network])))
# -
# Finally, remember that for all the methods (*fit*, *predict*, ...) keras requires as input a numpy array of array, for example you must convert your state in the correct **shape**. Kears will return, in the same way, an array of array, so to extract the corresponding ouutput layer you must select the first element.
state = np.array([0, 0, 0])
# model.predict(input_network) will give you a shape error
state = state.reshape(1, 3)
print("Prediction:", model.predict(state)[0])
# ## Assignment: Q-Learning
#
# Your first assignement is to implement all the functions necessary for a deep q-learning algorithm. In particular you must implement the following functions: *create_model*, *train_model* and *DQN*.
#
# #### Hint:
# For the experience replay buffer you can use the python data structure *dequeue*, defining the maximum length allowed. With the *random.sample(replay_buffer, size)* function you can sample *size* element from the queue:
# +
replay_buffer = deque(maxlen=10000)
for _ in range(100): replay_buffer.append(random.uniform(0, 1))
samples = random.sample(replay_buffer, 3)
print("Get 3 elements from replay_buffer:", samples)
# -
def create_model(input_size, output_size, hidden_layer_size, hidden_layer_number):
"""
Create the neural netowrk model with the given parameters
Args:
input_size: the number of nodes for the input layer
output_size: the number of nodes for the output layer
hidden_layer_size: the number of nodes for each hidden layer
hidden_layer_number: the number of hidden layers
Returns:
model: the corresponding neural network
"""
#
# YOUR CODE HERE ...
#
return model
def train_model(model, memory, batch_size, gamma=0.99):
"""
Performs the value iteration algorithm for a specific environment
Args:
model: the neural network model to train
memory: the memory array on wich perform the training
batch_size: the size of the batch sampled from the memory
gamma: gamma value, the discount factor for the Bellman equation
"""
#
# YOUR CODE HERE ...
#
return model
def DQN(environment, neural_network, trials, goal_score, batch_size, epsilon_decay=0.9995):
"""
Performs the Q-Learning algorithm for a specific environment on a specific neural netowrk model
Args:
environment: OpenAI Gym environment
neural_network: the neural netowrk to train
trials: the number of iterations for the training phase
goal_score: the minimum score to consider 'solved' the problem
batch_size: the size of the batch sampled from the memory
epsilon_decay: the dacay value of epsilon for the eps-greedy exploration
Returns:
score_queue: 1-d dimensional array of the reward obtained at each trial step
"""
epsilon = 1.0; epsilon_min = 0.01
score = 0; score_queue = []
#
# YOUR CODE HERE ...
#
for trial in range(trials):
#
# YOUR CODE HERE ...
#
if(score > 130 or score_queue == []): break
print("Episode: {:7.0f}, Score: {:3.0f}, EPS: {:3.2f}".format(trial, score_queue[-1], epsilon))
return neural_network, score_queue
# ## Execution
# The following code executes the DQN and plots the reward function, the execution could require up to 10 minutes on some computer. A more efficent version of the code can be found [here](https://github.com/d-corsi/BasicRL).
# Correct results for comparison can be found here below. Notice that since the executions are stochastic the charts could differ: the important thing is the global trend and the final convergence to a visible reward improvement.
env = gym.make("CartPole-v1")
neural_network = create_model(4, 2, 32, 2)
neural_network, score = DQN(env, neural_network, trials=1000, goal_score=130, batch_size=64)
if score != []:
rewser = []
window = 10
score = rolling(np.array(score), window)
rewser.append({"x": np.arange(1, len(score) + 1), "y": score, "ls": "-", "label": "DQN"})
plot(rewser, "Rewards", "Episodes", "Rewards")
# **Standard DQN on CartPole results:**
# <img src="images/results-dqn.png" width="600">
| lesson_6/lesson_6_problem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Shallow Water Equations
import numpy as np
import matplotlib.pyplot as plt
% matplotlib inline
from matplotlib import animation
from IPython.display import HTML
# ## Newtonian, 2d, single layer
# +
# define grids
nx = 2000
nt = 2000
xmin = 0.
xmax = 10.
x = np.linspace(xmin, xmax, num=nx-2, endpoint=False)
hu = np.zeros((nx, nt+1))
h = np.zeros((nx, nt+1))
# Start off with an initial water hill
h[1:-1,0] = 1 + 0.4 * np.exp(-(x-5)**2*2) #0.01 * np.sin(0.25 * np.pi + 2. * np.pi * x / (xmax - xmin)) + 1.
h[0,0] = h[-2,0]
h[-1,0] = h[1,0]
#hu[:,0] = 0.1 * h[:,0]
dx = x[1] - x[0]
dt = 0.5 * dx
# set gravity
g = 1.0
# -
plt.plot(x,h[1:-1,0], lw=2)
plt.show()
# evolution using second-order Lax-Wendroff
def evolve(n):
# Jacobean
J = np.zeros((nx-2, 2, 2))
J[:,0,1] = 1.
J[:,1,0] = -(hu[1:-1,n]/h[1:-1,n])**2 + g * h[1:-1,n]
J[:,1,1] = 2 * hu[1:-1,n] / h[1:-1,n]
J2 = np.array([np.dot(J[i,:,:], J[i,:,:]) for i in range(nx-2)])
h[1:-1,n+1] = h[1:-1,n] - 0.5 * dt/dx * (J[:,0,0] * (h[2:,n] - h[:-2,n]) + J[:,0,1] * (hu[2:,n] - hu[:-2,n])) +\
0.5 * (dt/dx)**2 * (J2[:,0,0] * (h[:-2,n] - 2 * h[1:-1,n] + h[2:,n]) + \
J2[:,0,1] * (hu[:-2,n] - 2 * hu[1:-1,n] + hu[2:,n]))
hu[1:-1,n+1] = hu[1:-1,n] - 0.5 * dt/dx * (J[:,1,0] * (h[2:,n] - h[:-2,n]) + J[:,1,1] * (hu[2:,n] - hu[:-2,n]))+ \
0.5 * (dt/dx)**2 * (J2[:,1,0] * (h[:-2,n] - 2 * h[1:-1,n] + h[2:,n]) + \
J2[:,1,1] * (hu[:-2,n] - 2 * hu[1:-1,n] + hu[2:,n]))
# do boundaries - periodic
h[0,n+1] = h[-2,n+1]
h[-1,n+1] = h[1,n+1]
hu[0,n+1] = hu[-2,n+1]
hu[-1,n+1] = hu[1,n+1]
for i in range(nt):
evolve(i)
plt.plot(x,h[1:-1,800], lw=2)
plt.show()
# +
fig = plt.figure()
ax = plt.axes(xlim=(0,10), ylim=(1.,1.4))
line = ax.plot([],[], lw=2)[0]
def animate(i):
line.set_data(x, h[1:-1,i*10])
#return line
anim = animation.FuncAnimation(fig, animate, frames=130, interval=25)#, blit=True)
# -
HTML(anim.to_html5_video())
# ## Newtonian, 2d, multilayer, no heat transfer
# +
# define grids
nx = 2000
nt = 2000
nlayers = 2
xmin = 0.
xmax = 10.
x = np.linspace(xmin, xmax, num=nx-2, endpoint=False)
rho = np.ones(nlayers)
hu = np.zeros((nlayers, nx, nt+1))
h = np.zeros((nlayers, nx, nt+1))
# Start off with an initial water hill
h[0,1:-1,0] = 1 + 0.4 * np.exp(-(x-2)**2*2) #0.01 * np.sin(0.25 * np.pi + 2. * np.pi * x / (xmax - xmin)) + 1.
h[1,1:-1,0] = 0.8 + 0.2 * np.exp(-(x-7)**2*2)
h[:,0,0] = h[:,-2,0]
h[:,-1,0] = h[:,1,0]
#hu[:,0] = 0.1 * h[:,0]
# going to try setting the top fluid to be heavier
rho[0] = 3
dx = x[1] - x[0]
dt = 0.5 * dx
# set gravity
g = 1.0
# -
plt.plot(x,h[0,1:-1,0],x,h[1,1:-1,0], lw=2)
plt.show()
# evolution using second-order Lax-Wendroff
def evolve(n):
for j in range(nlayers):
# Jacobian
J = np.zeros((nx-2, 2, 2))
J[:,0,1] = 1.
J[:,1,0] = -(hu[j,1:-1,n]/h[j,1:-1,n])**2 + g * h[j,1:-1,n]
J[:,1,1] = 2 * hu[j,1:-1,n] / h[j,1:-1,n]
J2 = np.array([np.dot(J[i,:,:], J[i,:,:]) for i in range(nx-2)])
# calculate source term
sum_hs = np.zeros(nx)
for i in range(j):
sum_hs += rho[i] / rho[j] * h[i,:,n]
for i in range(j+1,nlayers):
sum_hs += h[i,:,n]
dx_sumhs = 0.5/dx * (sum_hs[2:] - sum_hs[:-2])# - 0.5/dx**2 * (sum_hs[2:] - 2 * sum_hs[1:-1] + sum_hs[:-2])
beta = g * dx_sumhs
h[j,1:-1,n+1] = h[j,1:-1,n] - 0.5 * dt/dx * (J[:,0,0] * (h[j,2:,n] - h[j,:-2,n]) + \
J[:,0,1] * (hu[j,2:,n] - hu[j,:-2,n])) +\
0.5 * (dt/dx)**2 * (J2[:,0,0] * (h[j,:-2,n] - 2 * h[j,1:-1,n] + h[j,2:,n]) + \
J2[:,0,1] * (hu[j,:-2,n] - 2 * hu[j,1:-1,n] + hu[j,2:,n]))
hu[j,1:-1,n+1] = (1 - dt * beta + 0.5 * dt**2 * beta**2) * hu[j,1:-1,n] - \
0.5 * dt/dx * (1 - dt*beta) * (J[:,1,0] * (h[j,2:,n] - h[j,:-2,n]) + \
J[:,1,1] * (hu[j,2:,n] - hu[j,:-2,n]))+ \
0.5 * (dt/dx)**2 * (J2[:,1,0] * (h[j,:-2,n] - 2 * h[j,1:-1,n] + h[j,2:,n]) + \
J2[:,1,1] * (hu[j,:-2,n] - 2 * hu[j,1:-1,n] + hu[j,2:,n]))
# do boundaries - periodic
h[:,0,n+1] = h[:,-2,n+1]
h[:,-1,n+1] = h[:,1,n+1]
hu[:,0,n+1] = hu[:,-2,n+1]
hu[:,-1,n+1] = hu[:,1,n+1]
for i in range(nt):
evolve(i)
plt.plot(x,h[0,1:-1,1000],x,h[1,1:-1,1000], lw=2)
plt.show()
# +
fig = plt.figure()
ax = plt.axes(xlim=(0,10), ylim=(0.7,1.4))
line = ax.plot([],[], lw=2)[0]
line2 = ax.plot([],[], lw=2)[0]
def animate(i):
line.set_data(x, h[0,1:-1,i*10])
line2.set_data(x, h[1,1:-1,i*10])
anim = animation.FuncAnimation(fig, animate, frames=130, interval=25)#, init_func=init)
# -
HTML(anim.to_html5_video())
# ## Newtonian, 2d, multilayer, heat transfer
# +
# define grids
nx = 2000
nt = 2000
nlayers = 2
xmin = 0.
xmax = 10.
x = np.linspace(xmin, xmax, num=nx-2, endpoint=False)
rho = np.ones(nlayers)
hu = np.zeros((nlayers, nx, nt+1))
h = np.zeros((nlayers, nx, nt+1))
# Start off with an initial water hill
h[0,1:-1,0] = 1 + 0.4 * np.exp(-(x-2)**2*2) #0.01 * np.sin(0.25 * np.pi + 2. * np.pi * x / (xmax - xmin)) + 1.
h[1,1:-1,0] = 0.8 + 0.2 * np.exp(-(x-7)**2*2)
h[:,0,0] = h[:,-2,0]
h[:,-1,0] = h[:,1,0]
#hu[:,0] = 0.1 * h[:,0]
# going to try setting the top fluid to be heavier
rho[0] = 3
dx = x[1] - x[0]
dt = 0.5 * dx
# set gravity
g = 1.0
# heating rate - flux of material from lower to upper layer. Set Q of top layer as 0.
Q = np.zeros(nlayers)
Q[1] = 0.01
# -
plt.plot(x,h[0,1:-1,0],x,h[1,1:-1,0], lw=2)
plt.show()
# evolution using second-order Lax-Wendroff
def evolve(n):
for j in range(nlayers):
# Jacobean
J = np.zeros((nx-2, 2, 2))
J[:,0,1] = 1.
J[:,1,0] = -(hu[j,1:-1,n]/h[j,1:-1,n])**2 + g * h[j,1:-1,n]
J[:,1,1] = 2 * hu[j,1:-1,n] / h[j,1:-1,n]
J2 = np.array([np.dot(J[i,:,:], J[i,:,:]) for i in range(nx-2)])
# calculate source terms
sum_hs = np.zeros(nx)
sum_qs = 0
if j < (nlayers - 1): # i.e. it has another layer beneath it
sum_qs += abs((Q[j+1] - Q[j]))
deltaQv = max(0,Q[j] - Q[j+1]) * (hu[j,:,n] - hu[j+1,:,n]) / h[j,:,n]
if j > 0: # i.e. has another layer above it
sum_qs += -rho[j-1]/rho[j] * (Q[j] - Q[j-1])
deltaQv = rho[j-1]/rho[j] * (Q[j] - Q[j-1]) * (hu[j,:,n] - hu[j-1,:,n]) / h[j,:,n]
for i in range(j):
sum_hs += rho[i] / rho[j] * h[i,:,n]
for i in range(j+1,nlayers):
sum_hs += h[i,:,n]
dx_sumhs = 0.5/dx * (sum_hs[2:] - sum_hs[:-2])# - 0.5/dx**2 * (sum_hs[2:] - 2 * sum_hs[1:-1] + sum_hs[:-2])
beta = g * dx_sumhs
h[j,1:-1,n+1] = h[j,1:-1,n] - 0.5 * dt/dx * (J[:,0,0] * (h[j,2:,n] - h[j,:-2,n]) + \
J[:,0,1] * (hu[j,2:,n] - hu[j,:-2,n])) +\
0.5 * (dt/dx)**2 * (J2[:,0,0] * (h[j,:-2,n] - 2 * h[j,1:-1,n] + h[j,2:,n]) + \
J2[:,0,1] * (hu[j,:-2,n] - 2 * hu[j,1:-1,n] + hu[j,2:,n])) +\
dt * sum_qs
hu[j,1:-1,n+1] = (1 - dt * beta + 0.5 * dt**2 * beta**2) * hu[j,1:-1,n] - \
0.5 * dt/dx * (1 - dt*beta) * (J[:,1,0] * (h[j,2:,n] - h[j,:-2,n]) + \
J[:,1,1] * (hu[j,2:,n] - hu[j,:-2,n]))+ \
0.5 * (dt/dx)**2 * (J2[:,1,0] * (h[j,:-2,n] - 2 * h[j,1:-1,n] + h[j,2:,n]) + \
J2[:,1,1] * (hu[j,:-2,n] - 2 * hu[j,1:-1,n] + hu[j,2:,n])) -\
dt * deltaQv[1:-1]
# do boundaries - periodic
h[:,0,n+1] = h[:,-2,n+1]
h[:,-1,n+1] = h[:,1,n+1]
hu[:,0,n+1] = hu[:,-2,n+1]
hu[:,-1,n+1] = hu[:,1,n+1]
for i in range(nt):
evolve(i)
# +
fig = plt.figure()
ax = plt.axes(xlim=(0,10), ylim=(0.7,1.4))
line = ax.plot([],[], lw=2)[0]
line2 = ax.plot([],[], lw=2)[0]
def animate(i):
line.set_data(x, h[0,1:-1,i*10])
line2.set_data(x, h[1,1:-1,i*10])
anim = animation.FuncAnimation(fig, animate, frames=130, interval=25)#, init_func=init)
# -
HTML(anim.to_html5_video())
# ## Newtonian, 3d
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
# +
# define grids
nx = 100
ny = 100
nt = 50
nlayers = 2
xmin = 0.
xmax = 10.
ymin = 0.
ymax = 10.
rho = np.ones(nlayers)
# going to try setting the top fluid to be heavier
rho[0] = 1.5
# set gravity
g = 1.0
# heating rate - flux of material from lower to upper layer. Set Q of top layer as 0.
Q = np.zeros(nlayers)
#Q[1] = 0.05
# Coriolis parameter - two times the angular frequency
f = 0.1
class U:
def __init__(self, nlayers, nx, ny, nt, xmin, xmax, ymin, ymax, rho, g, Q, f, periodic=True):
self.nlayers = nlayers
self.nx = nx
self.ny = ny
self.nt = nt
self.U = np.zeros((3, nlayers, nx, ny, nt+1))
self.x = np.linspace(xmin, xmax, num=nx-2, endpoint=False)
self.y = np.linspace(ymin, ymax, num=ny-2, endpoint=False)
self.rho = rho
self.dx = self.x[1] - self.x[0]
self.dy = self.y[1] - self.y[0]
self.dt = 0.5 * min(self.dx, self.dy)
self.g = g
self.Q = Q
self.f = f
self.periodic = periodic
def h(self,indices):
return self.U[(0,) + tuple(indices)]
def hu(self,indices):
return self.U[(1,) + tuple(indices)]
def hv(self,indices):
return self.U[(2,) + tuple(indices)]
def initial_data(self, h0=None, hu0=None, hv0=None):
"""
Set the initial data
"""
if h0 is not None:
self.U[0,:,:,:,0] = h0
if hu0 is not None:
self.U[1,:,:,:,0] = hu0
if hv0 is not None:
self.U[2,:,:,:,0] = hv0
# enforce bcs
self.bcs(0)
def Uj(self, layer, t):
return self.U[:,layer,:,:,t]
def bcs(self, t):
if self.periodic:
self.U[:,:,0,:,t] = self.U[:,:,-2,:,t]
self.U[:,:,:,0,t] = self.U[:,:,:,-2,t]
self.U[:,:,-1,:,t] = self.U[:,:,1,:,t]
self.U[:,:,:,-1,t] = self.U[:,:,:,1,t]
else: #outflow
self.U[:,:,0,:,t] = self.U[:,:,1,:,t]
self.U[:,:,:,0,t] = self.U[:,:,:,1,t]
self.U[:,:,-1,:,t] = self.U[:,:,-2,:,t]
self.U[:,:,:,-1,t] = self.U[:,:,:,-2,t]
U = U(nlayers, nx, ny, nt, xmin, xmax, ymin, ymax, rho, g, Q, f, periodic=False)
#q = np.zeros((nlayers, 3, nx, ny, nt+1))
#hu = np.zeros((nlayers, nx, ny, nt+1))
#h = np.zeros((nlayers, nx, ny, nt+1))
# Start off with an initial water hill
h0 = np.zeros_like(U.U[0,:,:,:,0])
h0[0,1:-1,1:-1] = 1 + 0.4 * np.exp(-((U.x[:,np.newaxis]-2)**2 + (U.y[np.newaxis,:]-2)**2)*2) #0.01 * np.sin(0.25 * np.pi + 2. * np.pi * x / (xmax - xmin)) + 1.
h0[1,1:-1,1:-1] = 0.8 + 0.2 * np.exp(-((U.x[:,np.newaxis]-7)**2 + (U.y[np.newaxis,:]-7)**2)*2)
#hu[:,0] = 0.1 * h[:,0]
# set coriolis source to be outside of grid, beyond bottom left corner
hu0 = np.zeros_like(U.U[1,:,:,:,0])
hu0[:,1:-1,1:-1] = -0.5 * f * (U.x[np.newaxis,:,np.newaxis]+1)
hu0[np.isnan(hu0)] = 0
hu0 *= h0
hv0 = np.zeros_like(U.U[2,:,:,:,0])
hv0[:,1:-1,1:-1] = 0.5 * f * (U.y[np.newaxis,np.newaxis,:]+1)
hv0[np.isnan(hv0)] = 0
hv0 *= h0
U.initial_data(h0=h0, hu0=hu0, hv0=hv0)
# -
# print initial velocity field
X, Y = np.meshgrid(U.x,U.y)
magv = np.sqrt(U.U[1,0,1:-1:5,1:-1:5,0]**2 + U.U[2,0,1:-1:5,1:-1:5,0]**2) / U.U[0,0,1:-1:5,1:-1:5,0]
#print(magv)
plot = plt.figure()
plt.quiver(X[::5,::5], Y[::5,::5], U.U[1,0,1:-1:5,1:-1:5,0], U.U[2,0,1:-1:5,1:-1:5,0], magv, cmap=cm.viridis)
plt.show()
# +
X, Y = np.meshgrid(U.x,U.y)
fig = plt.figure(figsize=(12,10))
ax = fig.gca(projection='3d')
ax.set_xlim(0,10)
ax.set_ylim(0,10)
ax.set_zlim(0.7,1.4)
ax.plot_surface(X,Y,U.U[0,1,1:-1,1:-1,0], rstride=1, cstride=2, lw=0, cmap=cm.viridis, antialiased=True)
ax.plot_wireframe(X,Y,U.U[0,0,1:-1,1:-1,0], rstride=2, cstride=2, lw=0.1, cmap=cm.viridis, antialiased=True)
#plt.plot(x,h[0,1:-1,0],x,h[1,1:-1,0], lw=2)
plt.show()
# -
# evolution using second-order Lax-Wendroff
def evolve(n):
for j in range(nlayers):
# Jacobians
def Jx(x,y):
h = U.U[0,j, x, y, n]
u = U.U[1,j, x, y, n] / U.U[0,j, x, y, n]
v = U.U[2,j, x, y, n] / U.U[0,j, x, y, n]
return np.array([[0, 1, 0], [-u**2 + U.g * h, 2*u, 0], [-u * v, v, u]]).T
def Jy(x,y):
h = U.U[0,j, x, y, n]
u = U.U[1,j, x, y, n] / U.U[0,j, x, y, n]
v = U.U[2,j, x, y, n] / U.U[0,j, x, y, n]
return np.array([[0, 0, 1], [-u*v, v, u], [-v**2 + U.g*h, 0, 2*v]]).T
A = np.array([[Jx(i,k) for k in range(1,U.ny-1)] for i in range(1,U.nx-1)])
B = np.array([[Jy(i,k) for k in range(1,U.ny-1)] for i in range(1,U.nx-1)])
A2 = np.array([[np.dot(A[i,k,:,:], A[i,k,:,:]) for k in range(U.ny-2)] for i in range(U.nx-2)])
B2 = np.array([[np.dot(B[i,k,:,:], B[i,k,:,:]) for k in range(U.ny-2)] for i in range(U.nx-2)])
AB = np.array([[np.dot(A[i,k,:,:], B[i,k,:,:]) for k in range(U.ny-2)] for i in range(U.nx-2)])
# to simplify indexing to 3 indices rather than 5
qn = U.U[:,j,:,:,n]
U.U[:,j,1:-1,1:-1,n+1] = qn[:,1:-1,1:-1] - \
0.5 * U.dt/U.dx * np.einsum('ijkl,lij->kij', A[:,:,:,:], (qn[:,2:,1:-1] - qn[:,:-2,1:-1])) -\
0.5 * U.dt/U.dy * np.einsum('ijkl,lij->kij', B[:,:,:,:], (qn[:,1:-1,2:] - qn[:,1:-1,:-2])) +\
0.5 * (U.dt/U.dx)**2 * np.einsum('ijkl,lij->kij', A2[:,:,:,:], (qn[:,2:,1:-1] - 2*qn[:,1:-1,1:-1] + qn[:,:-2,1:-1])) +\
0.5 * (U.dt/U.dy)**2 * np.einsum('ijkl,lij->kij', B2[:,:,:,:], (qn[:,1:-1,2:] - 2*qn[:,1:-1,1:-1] + qn[:,1:-1,:-2])) -\
0.25 * U.dt**2/(U.dx*U.dy) * np.einsum('ijkl,lij->kij', AB[:,:,:,:], (qn[:,2:,2:] - qn[:,2:,:-2] -\
qn[:,:-2,2:] + qn[:,:-2,:-2]))
# do boundaries
U.bcs(n+1)
U_half = U.U[:,:,:,:,n+1]
for j in range(nlayers):
# calculate source terms
# a more sophisticated scheme for the source terms is needed, but that's a real headache so shall ignore for now
# and just use operator splitting
h = U_half[0,:,:,:]
hu = U_half[1,:,:,:]
hv = U_half[2,:,:,:]
sum_hs = np.zeros((U.nx,U.ny))
sum_qs = 0
if j < (nlayers - 1): # i.e. it has another layer beneath it
sum_qs += ((U.Q[j+1] - U.Q[j]))
deltaQu = (U.Q[j] - U.Q[j+1]) * (hu[j,:,:] - hu[j+1,:,:]) / h[j,:,:]
deltaQv = (U.Q[j] - U.Q[j+1]) * (hv[j,:,:] - hv[j+1,:,:]) / h[j,:,:]
if j > 0: # i.e. has another layer above it
sum_qs += -U.rho[j-1]/U.rho[j] * (U.Q[j] - U.Q[j-1])
deltaQu = U.rho[j-1]/U.rho[j] * (U.Q[j] - U.Q[j-1]) * (hu[j,:,:] - hu[j-1,:,:]) / h[j,:,:]
deltaQv = U.rho[j-1]/U.rho[j] * (U.Q[j] - U.Q[j-1]) * (hv[j,:,:] - hv[j-1,:,:]) / h[j,:,:]
for i in range(j):
sum_hs += U.rho[i] / U.rho[j] * h[i,:,:]
for i in range(j+1,nlayers):
sum_hs += h[i,:,:]
dx_sumhs = 0.5/U.dx * (sum_hs[2:,1:-1] - sum_hs[:-2,1:-1])
dy_sumhs = 0.5/U.dy * (sum_hs[1:-1,2:] - sum_hs[1:-1,:-2])
# h
U.U[0,j,1:-1,1:-1,n+1] += U.dt * (sum_qs)
# hu
U.U[1,j,1:-1,1:-1,n+1] += U.dt * (-deltaQu[1:-1,1:-1] - U.g * dx_sumhs) * h[j,1:-1,1:-1] + \
U.dt * U.f * hv[j,1:-1,1:-1]
# hv
U.U[2,j,1:-1,1:-1,n+1] += U.dt * (-deltaQv[1:-1,1:-1] - U.g * dy_sumhs) * h[j,1:-1,1:-1] - \
U.dt * U.f * hu[j,1:-1,1:-1]
# do boundaries
U.bcs(n+1)
for i in range(nt):
evolve(i)
# +
X, Y = np.meshgrid(U.x,U.y)
fig = plt.figure(figsize=(12,10))
ax = fig.gca(projection='3d')
ax.set_xlim(0,10)
ax.set_ylim(0,10)
ax.set_zlim(0.7,1.4)
n = 1
ax.plot_surface(X,Y,U.U[0,1,1:-1,1:-1,n], rstride=1, cstride=2, lw=0, cmap=cm.viridis, antialiased=True)
ax.plot_wireframe(X,Y,U.U[0,0,1:-1,1:-1,n], rstride=2, cstride=2, lw=0.1, cmap=cm.viridis, antialiased=True)
#plt.plot(x,h[0,1:-1,0],x,h[1,1:-1,0], lw=2)
plt.show()
# +
fig = plt.figure(figsize=(12,10))
ax = fig.gca(projection='3d')
#ax = plt.axes(xlim=(0,10), zlim=(0.7,1.4))
#surface_1 = ax.plot_surface([],[],[], rstride=1, cstride=2, lw=0, cmap=cm.viridis, antialiased=True)[0]
#surface_2 = ax.plot_wireframe([],[],[], rstride=2, cstride=2, lw=0.1, cmap=cm.viridis, antialiased=True)[0]
#line = ax.plot([],[], lw=2)[0]
#line2 = ax.plot([],[], lw=2)[0]
surface_1 = ax.plot_surface(X,Y,U.U[0,1,1:-1,1:-1,0], rstride=1, cstride=2, lw=0, cmap=cm.viridis, antialiased=True)
surface_2 = ax.plot_wireframe(X,Y,U.U[0,0,1:-1,1:-1,0], rstride=2, cstride=2, lw=0.1, cmap=cm.viridis, antialiased=True)
def init():
surface_1 = ax.plot_surface(X,Y,U.U[0,1,1:-1,1:-1,0], rstride=1, cstride=2, lw=0, cmap=cm.viridis, antialiased=True)
surface_2 = ax.plot_wireframe(X,Y,U.U[0,0,1:-1,1:-1,0], rstride=2, cstride=2, lw=0.1, cmap=cm.viridis, antialiased=True)
def animate(i):
ax.clear()
ax.set_xlim(0,10)
ax.set_ylim(0,10)
ax.set_zlim(0.7,1.4)
ax.plot_surface(X,Y,U.U[0,1,1:-1,1:-1,i], rstride=1, cstride=2, lw=0, cmap=cm.viridis, antialiased=True)
ax.plot_wireframe(X,Y,U.U[0,0,1:-1,1:-1,i], rstride=2, cstride=2, lw=0.1, cmap=cm.viridis, antialiased=True)
anim = animation.FuncAnimation(fig, animate, frames=30, interval=200)#, init_func=init)
# -
HTML(anim.to_html5_video())
# ## GR, 2d
# +
# define grids
nx = 500
nt = 1000
nlayers = 2
# moved boundaries away as there was some nasty feedback occuring here
xmin = -5.
xmax = 15.
alpha = 0.9
beta = 0.
gamma = 1 / alpha**2
class U:
def __init__(self, nlayers, nx, nt, xmin, xmax, rho, alpha, beta, gamma, periodic=True):
self.nlayers = nlayers
self.nx = nx
self.nt = nt
self.U = np.zeros((2, nlayers, nx, nt+1))
self.x = np.linspace(xmin, xmax, num=nx-2, endpoint=False)
self.rho = rho
self.dx = self.x[1] - self.x[0]
self.dt = 0.1 * self.dx
# metric stuff
self.alpha = alpha
self.beta = beta
self.gamma = gamma # gamma down
self.gamma_up = 1/gamma
self.periodic = periodic
def D(self,indices):
return self.U[(0,) + tuple(indices)]
def Sx(self,indices):
return self.U[(1,) + tuple(indices)]
def initial_data(self, D0=None, Sx0=None):
"""
Set the initial data
"""
if D0 is not None:
self.U[0,:,:,0] = D0
if Sx0 is not None:
self.U[1,:,:,0] = Sx0
# enforce bcs
self.bcs(0)
def Uj(self, layer, t):
return self.U[:,layer,:,t]
def bcs(self, t):
if self.periodic:
self.U[:,:,0,t] = self.U[:,:,-2,t]
self.U[:,:,-1,t] = self.U[:,:,1,t]
else: # outflow
self.U[:,:,0,t] = self.U[:,:,1,t]
self.U[:,:,-1,t] = self.U[:,:,-2,t]
rho = np.ones(nlayers)
U = U(nlayers, nx, nt, xmin, xmax, rho, alpha, beta, gamma, periodic=False)
# Start off with an initial water hill
D0 = np.zeros_like(U.U[0,:,:,0])
D0[0,1:-1] = 1 + 0.2 * np.exp(-(U.x-2)**2*2)
D0[1,1:-1] = 0.8 + 0.1 * np.exp(-(U.x-7)**2*2)
U.initial_data(D0=D0)
# -
plt.plot(U.x,U.U[0,0,1:-1,0],U.x,U.U[0,1,1:-1,0], lw=2)
plt.show()
def evolve(n):
for j in range(nlayers):
# Jacobians
def Jx(x):
# do naive c2p for now
W = np.sqrt(U.U[1, j, x, n] * U.U[1, j, x, n] * U.gamma_up / U.U[0, j, x, n]**2 + 1)
ph = U.U[0, j, x, n] / W #
u = U.U[1, j, x, n] / (U.U[0, j, x, n] * W) # u_down
qx = u * U.gamma_up - U.beta/U.alpha
xi = 1 / (1 + u**2 * W**2)
return np.array([[qx - u*xi, xi / W], [ph*(1 + 0.5*xi*u**2*W**2)/W, qx - 0.5*ph*xi*u]]).T
A = np.array([Jx(i) for i in range(1,U.nx-1)])
A2 = np.array([np.dot(A[i,:], A[i,:]) for i in range(U.nx-2)])
# to simplify indexing to 2 indices rather than 4
qn = U.U[:,j,:,n]
U.U[:,j,1:-1,n+1] = qn[:,1:-1] - \
0.5 * U.dt/U.dx * np.einsum('ikl,li->ki', A[:,:,:], (qn[:,2:] - qn[:,:-2])) -\
0.5 * (U.dt/U.dx)**2 * np.einsum('ikl,li->ki', A2[:,:,:], (qn[:,2:] - 2*qn[:,1:-1] + qn[:,:-2]))
# do boundaries
U.bcs(n+1)
"""
Going to not include any source terms for now
U_half = U.U[:,:,:,n+1]
for j in range(nlayers):
# calculate source terms
# a more sophisticated scheme for the source terms is needed, but that's a real headache so shall ignore for now
# and just use operator splitting
h = U_half[0,:,:]
hu = U_half[1,:,:]
hv = U_half[2,:,:]
sum_hs = np.zeros(U.nx)
sum_qs = 0
if j < (nlayers - 1): # i.e. it has another layer beneath it
sum_qs += -U.rho[j+1]/U.rho[j] * abs((U.Q[j+1] - U.Q[j]))
deltaQu = U.rho[j+1]/U.rho[j] * max(0,U.Q[j] - U.Q[j+1]) * (hu[j,:] - hu[j+1,:]) / h[j,:]
if j > 0: # i.e. has another layer above it
sum_qs += abs(U.Q[j] - U.Q[j-1])
deltaQu = max(0, U.Q[j] - U.Q[j-1]) * (hu[j,:] - hu[j-1,:]) / h[j,:]
for i in range(j):
sum_hs += U.rho[i] / U.rho[j] * h[i,:]
for i in range(j+1,nlayers):
sum_hs += h[i,:,:]
dx_sumhs = 0.5/U.dx * (sum_hs[2:] - sum_hs[:-2])
# h
U.U[0,j,1:-1,n+1] += U.dt * (sum_qs)
# hu
U.U[1,j,1:-1,n+1] += U.dt * (-deltaQu[1:-1] - U.g * dx_sumhs) * h[j,1:-1]
# do boundaries
U.bcs(n+1)
"""
for i in range(nt):
evolve(i)
plt.plot(U.x,U.U[0, 0, 1:-1, 0],U.x,U.U[0, 1, 1:-1, 0], lw=2)
plt.show()
# +
fig = plt.figure()
ax = plt.axes(xlim=(0,10), ylim=(0.7,1.4))
line = ax.plot([],[], lw=2)[0]
line2 = ax.plot([],[], lw=2)[0]
def animate(i):
line.set_data(U.x, U.U[0, 0, 1:-1,i*10])
line2.set_data(U.x, U.U[0, 1, 1:-1,i*10])
anim = animation.FuncAnimation(fig, animate, frames=100, interval=60)#, init_func=init)
# -
HTML(anim.to_html5_video())
# ## GR 3d
from numpy.linalg import inv, det
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
# +
# define grids
nx = 100
ny = 100
nt = 200
nlayers = 2
xmin = -5.
xmax = 15.
ymin = -5.
ymax = 15.
alpha = 1.0
beta = [0., 0.]
gamma = 1 / alpha**2 * np.eye(2)
# heating rate - flux of material from lower to upper layer. Set Q of top layer as 0.
Q = np.zeros((nlayers, nx, ny))
Q[:,1:-1,1:-1] = 0.2 * np.exp(-((U.x[np.newaxis,:,np.newaxis]-5)**2 + (U.y[np.newaxis,np.newaxis,:]-5)**2)*2)
Q[0,:,:] = -Q[0,:,:]
rho = np.ones(nlayers)
# going to try setting the top fluid to be heavier
#rho[0] = 1.5
class U:
def __init__(self, nlayers, nx, ny, nt, xmin, xmax, ymin, ymax, rho, Q, alpha, beta, gamma, periodic=True):
self.nlayers = nlayers
self.nx = nx
self.ny = ny
self.nt = nt
self.U = np.zeros((3, nlayers, nx, ny, nt+1))
self.x = np.linspace(xmin, xmax, num=nx-2, endpoint=False)
self.y = np.linspace(ymin, ymax, num=ny-2, endpoint=False)
self.rho = rho
self.dx = self.x[1] - self.x[0]
self.dy = self.y[1] - self.y[0]
self.dt = 0.1 * min(self.dx, self.dy)
self.Q = Q
# metric stuff
self.alpha = alpha
self.beta = beta
self.gamma = gamma # gamma down
self.gamma_up = inv(gamma)
self.periodic = periodic
def D(self,indices):
return self.U[(0,) + tuple(indices)]
def Sx(self,indices):
return self.U[(1,) + tuple(indices)]
def Sy(self,indices):
return self.U[(2,) + tuple(indices)]
def initial_data(self, D0=None, Sx0=None, Sy0=None):
"""
Set the initial data
"""
if D0 is not None:
self.U[0,:,:,:,0] = D0
if Sx0 is not None:
self.U[1,:,:,:,0] = Sx0
if Sy0 is not None:
self.U[2,:,:,:,0] = Sy0
# enforce bcs
self.bcs(0)
def Uj(self, layer, t):
return self.U[:,layer,:,:,t]
def bcs(self, t):
if self.periodic:
self.U[:,:,0,:,t] = self.U[:,:,-2,:,t]
self.U[:,:,:,0,t] = self.U[:,:,:,-2,t]
self.U[:,:,-1,:,t] = self.U[:,:,1,:,t]
self.U[:,:,:,-1,t] = self.U[:,:,:,1,t]
else: #outflow
self.U[:,:,0,:,t] = self.U[:,:,1,:,t]
self.U[:,:,:,0,t] = self.U[:,:,:,1,t]
self.U[:,:,-1,:,t] = self.U[:,:,-2,:,t]
self.U[:,:,:,-1,t] = self.U[:,:,:,-2,t]
rho = np.ones(nlayers)
# +
U = U(nlayers, nx, ny, nt, xmin, xmax, ymin, ymax, rho, Q, alpha, beta, gamma, periodic=False)
# Start off with an initial water hill
D0 = np.zeros_like(U.U[0,:,:,:,0])
D0[0,1:-1,1:-1] = 1 + 0.4 * np.exp(-((U.x[:,np.newaxis]-2)**2 + (U.y[np.newaxis,:]-2)**2)*2)
D0[1,1:-1,1:-1] = 0.8 + 0.2 * np.exp(-((U.x[:,np.newaxis]-7)**2 + (U.y[np.newaxis,:]-7)**2)*2)
U.initial_data(D0=D0)
# +
X, Y = np.meshgrid(U.x,U.y)
fig = plt.figure(figsize=(12,10))
ax = fig.gca(projection='3d')
ax.set_xlim(-5,15)
ax.set_ylim(-5,15)
ax.set_zlim(0.7,1.4)
ax.plot_surface(X,Y,U.U[0,1,1:-1,1:-1,0], rstride=1, cstride=2, lw=0, cmap=cm.viridis, antialiased=True)
ax.plot_wireframe(X,Y,U.U[0,0,1:-1,1:-1,0], rstride=2, cstride=2, lw=0.1, cmap=cm.viridis, antialiased=True)
#plt.plot(x,h[0,1:-1,0],x,h[1,1:-1,0], lw=2)
plt.show()
# -
# evolution using second-order Lax-Wendroff
# note: have assumed metric is constant so can move outside of derivatives
def evolve(U, n):
for j in range(nlayers):
# Jacobians
def Jx(x,y):
# do naive c2p for now
W = np.sqrt((U.U[1, j, x, y, n]**2 * U.gamma_up[0,0] +
2 * U.U[1, j, x, y, n] * U.U[2, j, x, y, n] * U.gamma_up[0,1] +
U.U[2, j, x, y, n]**2 * U.gamma_up[1,1]) / U.U[0, j, x, y, n]**2 + 1)
ph = U.U[0, j, x, y, n] / W #
u = U.U[1, j, x, y, n] / (U.U[0, j, x, y, n] * W) # u_down
v = U.U[2, j, x, y, n] / (U.U[0, j, x, y, n] * W) # v_down
qx = u * U.gamma_up[0,0] + v * U.gamma_up[0,1] - U.beta[0]/U.alpha
chi = 1 / (1 - u**2 * W**2 - v**2 * W**2)
Jx = np.zeros((3,3))
Jx[0,0] = qx/chi - u
Jx[0,1] = (1 + v**2*W**2)/W
Jx[0,2] = -W * u * v
Jx[1,0] = -2*W**3*u*qx*(u**2 + v**2) + ph*(1/W - W*u**2)
Jx[1,1] = qx * (1+W**2*u**2 + W**2*v**2) + 0.5*ph*u*(v**2*W**2-1)
Jx[1,2] = -v*ph*(1+0.5*W**2*u**2)
Jx[2,0] = -W*v*(2*W**2*qx*(u**2+v**2) + 0.5*ph*u)
Jx[2,1] = 0.5*ph*v*(1+v**2*W**2)
Jx[2,2] = qx*(1+W**2*u**2+W**2*v**2) - 0.5*ph*W**2*u*v**2
return chi * Jx
def Jy(x,y):
# do naive c2p for now
W = np.sqrt((U.U[1, j, x, y, n]**2 * U.gamma_up[0,0] +
2 * U.U[1, j, x, y, n] * U.U[2, j, x, y, n] * U.gamma_up[0,1] +
U.U[2, j, x, y, n]**2 * U.gamma_up[1,1]) / U.U[0, j, x, y, n]**2 + 1)
ph = U.U[0, j, x, y, n] / W #
u = U.U[1, j, x, y, n] / (U.U[0, j, x, y, n] * W) # u_down
v = U.U[2, j, x, y, n] / (U.U[0, j, x, y, n] * W) # v_down
qy = v * U.gamma_up[1,1] + u * U.gamma_up[0,1] - U.beta[1]/U.alpha
chi = 1 / (1 - u**2 * W**2 - v**2 * W**2)
Jy = np.zeros((3,3))
Jy[0,0] = qy/chi - v
Jy[0,1] = -W * u * v
Jy[0,2] = (1 + u**2*W**2)/W
Jy[1,0] = -W*u*(2*W**2*qy*(u**2+v**2) + 0.5*ph*v)
Jy[1,1] = qy*(1+W**2*u**2+W**2*v**2) - 0.5*ph*W**2*u**2*v
Jy[1,2] = 0.5*ph*u*(1+u**2*W**2)
Jy[2,0] = -2*W**3*v*qy*(u**2 + v**2) + ph*(1/W - W*v**2)
Jy[2,1] = -u*ph*(1+0.5*W**2*v**2)
Jy[2,2] = qy * (1+W**2*u**2 + W**2*v**2) + 0.5*ph*v*(u**2*W**2-1)
return chi * Jy
A = np.array([[Jx(i,k) for k in range(1,U.ny-1)] for i in range(1,U.nx-1)])
B = np.array([[Jy(i,k) for k in range(1,U.ny-1)] for i in range(1,U.nx-1)])
A2 = np.array([[np.dot(A[i,k,:,:], A[i,k,:,:]) for k in range(U.ny-2)] for i in range(U.nx-2)])
B2 = np.array([[np.dot(B[i,k,:,:], B[i,k,:,:]) for k in range(U.ny-2)] for i in range(U.nx-2)])
AB = np.array([[np.dot(A[i,k,:,:], B[i,k,:,:]) for k in range(U.ny-2)] for i in range(U.nx-2)])
# to simplify indexing to 3 indices rather than 5
qn = U.U[:,j,:,:,n]
U.U[:,j,1:-1,1:-1,n+1] = qn[:,1:-1,1:-1] + U.alpha * (\
-0.5 * U.dt/U.dx * np.einsum('ijkl,lij->kij', A[:,:,:,:], (qn[:,2:,1:-1] - qn[:,:-2,1:-1])) -\
0.5 * U.dt/U.dy * np.einsum('ijkl,lij->kij', B[:,:,:,:], (qn[:,1:-1,2:] - qn[:,1:-1,:-2])) +\
0.5 * (U.dt/U.dx)**2 * np.einsum('ijkl,lij->kij', A2[:,:,:,:], (qn[:,2:,1:-1] - 2*qn[:,1:-1,1:-1] + qn[:,:-2,1:-1])) +\
0.5 * (U.dt/U.dy)**2 * np.einsum('ijkl,lij->kij', B2[:,:,:,:], (qn[:,1:-1,2:] - 2*qn[:,1:-1,1:-1] + qn[:,1:-1,:-2])) -\
0.25 * U.dt**2/(U.dx*U.dy) * np.einsum('ijkl,lij->kij', AB[:,:,:,:], (qn[:,2:,2:] - qn[:,2:,:-2] -\
qn[:,:-2,2:] + qn[:,:-2,:-2])))
# do boundaries
U.bcs(n+1)
U_half = U.U[:,:,:,:,n+1]
W = np.sqrt((U_half[1,:,:,:]**2 * U.gamma_up[0,0] +
2 * U_half[1,:,:,:] * U_half[2,:,:,:] * U.gamma_up[0,1] +
U_half[2,:,:,:]**2 * U.gamma_up[1,1]) / U_half[0,:,:,:]**2 + 1)
ph = U_half[0,:,:,:] / W
Sx = U_half[1,:,:,:]
Sy = U_half[2,:,:,:]
for j in range(nlayers):
# calculate source terms
# a more sophisticated scheme for the source terms is needed, but that's a real headache so shall ignore for now
# and just use operator splitting
sum_phs = np.zeros((U.nx,U.ny))
sum_qs = 0
if j < (nlayers - 1): # i.e. it has another layer beneath it
sum_qs += ((U.Q[j+1,1:-1,1:-1] - U.Q[j,1:-1,1:-1]))
deltaQx = (U.Q[j,:,:] - U.Q[j+1,:,:]) * (Sx[j,:,:] - Sx[j+1,:,:]) / ph[j,:,:]
deltaQy = (U.Q[j,:,:] - U.Q[j+1,:,:]) * (Sy[j,:,:] - Sy[j+1,:,:]) / ph[j,:,:]
if j > 0: # i.e. has another layer above it
sum_qs += -U.rho[j-1]/U.rho[j] * (U.Q[j,1:-1,1:-1] - U.Q[j-1,1:-1,1:-1])
deltaQx = U.rho[j-1]/U.rho[j] * (U.Q[j,:,:] - U.Q[j-1,:,:]) * (Sx[j,:,:] - Sx[j-1,:,:]) / ph[j,:,:]
deltaQy = U.rho[j01]/U.rho[j] * (U.Q[j,:,:] - U.Q[j-1,:,:]) * (Sy[j,:,:] - Sy[j-1,:,:]) / ph[j,:,:]
for i in range(j):
sum_phs += U.rho[i] / U.rho[j] * ph[i,:,:]
for i in range(j+1,nlayers):
sum_phs += ph[i,:,:]
dx_sum_phs = 0.5/U.dx * (sum_phs[2:,1:-1] - sum_phs[:-2,1:-1])
dy_sum_phs = 0.5/U.dy * (sum_phs[1:-1,2:] - sum_phs[1:-1,:-2])
# h
U.U[0,j,1:-1,1:-1,n+1] += U.alpha * U.dt * (sum_qs)
# hu
U.U[1,j,1:-1,1:-1,n+1] += U.alpha * U.dt * (-deltaQx[1:-1,1:-1] - dx_sum_phs) * ph[j,1:-1,1:-1]
# hv
U.U[2,j,1:-1,1:-1,n+1] += U.alpha * U.dt * (-deltaQy[1:-1,1:-1] - dy_sum_phs) * ph[j,1:-1,1:-1]
# do boundaries
U.bcs(n+1)
for i in range(nt):
evolve(U, i)
# +
X, Y = np.meshgrid(U.x,U.y)
fig = plt.figure(figsize=(12,10))
ax = fig.gca(projection='3d')
ax.set_xlim(0,10)
ax.set_ylim(0,10)
ax.set_zlim(0.7,1.4)
n = 1
ax.plot_surface(X[23:-23,23:-23],Y[23:-23,23:-23],U.U[0,1,24:-24,24:-24,n], rstride=1, cstride=2, lw=0, cmap=cm.viridis, antialiased=True)
ax.plot_wireframe(X[23:-23,23:-23],Y[23:-23,23:-23],U.U[0,0,24:-24,24:-24,n], rstride=2, cstride=2, lw=0.1, cmap=cm.viridis, antialiased=True)
#plt.plot(x,h[0,1:-1,0],x,h[1,1:-1,0], lw=2)
plt.show()
# +
fig = plt.figure(figsize=(12,10))
ax = fig.gca(projection='3d')
#ax = plt.axes(xlim=(0,10), zlim=(0.7,1.4))
#surface_1 = ax.plot_surface([],[],[], rstride=1, cstride=2, lw=0, cmap=cm.viridis, antialiased=True)[0]
#surface_2 = ax.plot_wireframe([],[],[], rstride=2, cstride=2, lw=0.1, cmap=cm.viridis, antialiased=True)[0]
#line = ax.plot([],[], lw=2)[0]
#line2 = ax.plot([],[], lw=2)[0]
surface_1 = ax.plot_surface(X[20:-20,20:-20],Y[20:-20,20:-20],U.U[0,1,21:-21,21:-21,0], rstride=1, cstride=2, lw=0, cmap=cm.viridis, antialiased=True)
surface_2 = ax.plot_wireframe(X[20:-20,20:-20],Y[20:-20,20:-20],U.U[0,0,21:-21,21:-21,0], rstride=2, cstride=2, lw=0.1, cmap=cm.viridis, antialiased=True)
def init():
surface_1 = ax.plot_surface(X[20:-20,20:-20],Y[20:-20,20:-20],U.U[0,1,21:-21,21:-21,0], rstride=1, cstride=2, lw=0, cmap=cm.viridis, antialiased=True)
surface_2 = ax.plot_wireframe(X[20:-20,20:-20],Y[20:-20,20:-20],U.U[0,0,21:-21,21:-21,0], rstride=2, cstride=2, lw=0.1, cmap=cm.viridis, antialiased=True)
def animate(i):
ax.clear()
ax.set_xlim(-0.5,10.5)
ax.set_ylim(-0.5,10.5)
ax.set_zlim(0.7,1.4)
ax.plot_surface(X[20:-20,20:-20],Y[20:-20,20:-20],U.U[0,1,21:-21,21:-21,i], rstride=1, cstride=2, lw=0, cmap=cm.viridis, antialiased=True)
ax.plot_wireframe(X[20:-20,20:-20],Y[20:-20,20:-20],U.U[0,0,21:-21,21:-21,i], rstride=2, cstride=2, lw=0.1, cmap=cm.viridis, antialiased=True)
anim = animation.FuncAnimation(fig, animate, frames=200, interval=80)#, init_func=init)
# -
HTML(anim.to_html5_video())
# Note this looks kind of weird, but I think it is correct - the interaction between the layers means that after initially sinking down, the centre of the hill moves back up as it is pulled/pushed back up by the wave it has induced in the other layer.
# +
# Start off with an initial water hill moving in x-direction
nt = 400
beta = [0.1, -0.5]
U2 = U(nlayers, nx, ny, nt, xmin, xmax, ymin, ymax, rho, Q, alpha, beta, gamma, periodic=False)
D0 = np.zeros_like(U2.U[0,:,:,:,0])
Sy0 = np.zeros_like(D0)
D0[:,1:-1,1:-1] = 1 + 0.4 * np.exp(-(U2.y[np.newaxis,np.newaxis,:]-1)**2*2)
Sy0[:,1:-1,1:-1] = D0[:,1:-1,1:-1] * 0.3
U2.initial_data(D0=D0, Sy0=Sy0)
# -
for i in range(nt):
evolve(U2, i)
# +
X, Y = np.meshgrid(U2.x,U2.y)
fig = plt.figure(figsize=(12,10))
ax = fig.gca(projection='3d')
ax.set_xlim(0,10)
ax.set_ylim(0,10)
ax.set_zlim(0.7,1.4)
n = 1
ax.plot_surface(X[23:-23,23:-23],Y[23:-23,23:-23],U2.U[0,1,24:-24,24:-24,n], rstride=1, cstride=2, lw=0, cmap=cm.viridis, antialiased=True)
ax.plot_wireframe(X[23:-23,23:-23],Y[23:-23,23:-23],U2.U[0,0,24:-24,24:-24,n], rstride=2, cstride=2, lw=0.1, cmap=cm.viridis, antialiased=True)
#plt.plot(x,h[0,1:-1,0],x,h[1,1:-1,0], lw=2)
plt.show()
# +
fig = plt.figure(figsize=(12,10))
ax = fig.gca(projection='3d')
#ax = plt.axes(xlim=(0,10), zlim=(0.7,1.4))
#surface_1 = ax.plot_surface([],[],[], rstride=1, cstride=2, lw=0, cmap=cm.viridis, antialiased=True)[0]
#surface_2 = ax.plot_wireframe([],[],[], rstride=2, cstride=2, lw=0.1, cmap=cm.viridis, antialiased=True)[0]
#line = ax.plot([],[], lw=2)[0]
#line2 = ax.plot([],[], lw=2)[0]
surface_1 = ax.plot_surface(X[20:-20,20:-20],Y[20:-20,20:-20],U2.U[0,1,21:-21,21:-21,0], rstride=1, cstride=2, lw=0, cmap=cm.viridis, antialiased=True)
surface_2 = ax.plot_wireframe(X[20:-20,20:-20],Y[20:-20,20:-20],U2.U[0,0,21:-21,21:-21,0], rstride=2, cstride=2, lw=0.1, cmap=cm.viridis, antialiased=True)
def init():
surface_1 = ax.plot_surface(X[20:-20,20:-20],Y[20:-20,20:-20],U2.U[0,1,21:-21,21:-21,0], rstride=1, cstride=2, lw=0, cmap=cm.viridis, antialiased=True)
surface_2 = ax.plot_wireframe(X[20:-20,20:-20],Y[20:-20,20:-20],U2.U[0,0,21:-21,21:-21,0], rstride=2, cstride=2, lw=0.1, cmap=cm.viridis, antialiased=True)
def animate(i):
ax.clear()
ax.set_xlim(-0.5,10.5)
ax.set_ylim(-0.5,10.5)
ax.set_zlim(0.7,1.4)
ax.plot_surface(X[20:-20,20:-20],Y[20:-20,20:-20],U2.U[0,1,21:-21,21:-21,i], rstride=1, cstride=2, lw=0, cmap=cm.viridis, antialiased=True)
ax.plot_wireframe(X[20:-20,20:-20],Y[20:-20,20:-20],U2.U[0,0,21:-21,21:-21,i], rstride=2, cstride=2, lw=0.1, cmap=cm.viridis, antialiased=True)
anim = animation.FuncAnimation(fig, animate, frames=400, interval=80)#, init_func=init)
# -
HTML(anim.to_html5_video())
# For the tsunami, the wave travels much faster for a negative y-shift vector component than for a positive one (as you would expect).
| Shallow_Water_Equations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="oP6bsAuXgNS-"
# #**ETS** **CODE**
# + id="7I8Nr9tGLtUQ"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import os
# %matplotlib inline
# + id="AGgl9y2QLtUQ"
os.chdir('/Users/pradeepjaswani/Documents/python_trading/Python-for-Finance-Repo-master/08-Time-Series-Analysis')
# + id="BpT6kH8yLtUR"
airline1 = pd.read_csv('airline_passengers.csv',index_col= 'Month')
airline1.dropna(inplace=True)
# + id="VtmX6VzoLtUR"
airline1.index=pd.to_datetime(airline1.index)
# + id="CL3ae5T3LtUR" outputId="523898b4-d04e-4b8a-b022-c99985a579e2"
airline1.head()
# + id="lYK6SqLuLtUR" outputId="733dde3c-40b9-4467-eb8d-31473a404e22"
airline1.plot()
# + id="hQsxKuLiLtUR"
from statsmodels.tsa.seasonal import seasonal_decompose
# + id="fCQyqQx8LtUR"
result=seasonal_decompose(airline1['Thousands of Passengers'],model='multiplicative')
# there are two tyoes of models in this there is an additive and multiplicative model in order to choose the
# correct one you may need to guess and check here, from the look here you can say its a linear model, its also
# a little exponential therefore, here the multiplicative model is better usedd.
# + id="TrLFtD8XLtUR" outputId="b43f49e6-bac5-4718-efe4-bd459cc8264d"
result.seasonal.plot()
#just the seasonal component of the data
# + id="PqKiLs21LtUS" outputId="98cd6df0-800d-4c45-9a85-7348b30c9921"
result.trend.plot()
#switching to additive model will give the same trend graph,
# in our case additive or multiplicative did not make any diff because the original grapph is similar to
# linear g or expo g
# + id="jHxK8F29LtUS" outputId="ef548f49-d6d7-4afd-c342-aeae1953cd0a"
result.plot()
# + id="Y5hDiXcbf8BK"
| Decomposing_error_trend_and_Seasonality.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import time
# @param dfFile: pandas.DataFrame ('nba_preprocessed.csv')
# @param dateStart, dateEnd: str in the format of 'YYYY-MM-DD'
# @param attriToDrop: list[str]
# @return X, Y: pandas.DataFrame
# featureExtraction() outputs X, Y for model training.
# Game date can be assigned
# Attribute to be dropped can be assigned
def featureExtraction(dfFile, dateStart='1000-01-01', dateEnd='2999-12-31', attriToDrop=None):
df = pd.read_csv(dfFile)
# Date selection
df = df.loc[lambda df: (df.Date_A > dateStart) & (df.Date_A < dateEnd), :].reset_index(drop=True)
# Get label Y
Y = df[['W/L_A']]
Y = Y.rename(columns={'W/L_A': 'Label'})
# Get attributes X
colToDrop = ['Team_A', 'Date_A', 'W/L_A', 'Score_A', 'Opponent_A', 'Team_B', 'Date_B', 'W/L_B', 'Home/Away_B', 'Score_B', 'Opponent_B']
colToDrop += attriToDrop if attriToDrop else []
X = df.drop(columns = colToDrop)
return X, Y
# +
from sklearn.datasets import load_breast_cancer
from sklearn.model_selection import StratifiedKFold
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import GridSearchCV
from matplotlib import pyplot as plt
from sklearn.svm import SVC
import numpy as np
import operator
# %matplotlib inline
def CrossValidationGridSearchNested(X_data, Y_data, num_trials, fold_num, est_classifcation, tuned_param, scoring):
max_score = -1
best_estimator = est_classifcation
for i in range(num_trials):
inner_cv = StratifiedKFold(n_splits=fold_num, random_state=i, shuffle=True)
outer_cv = StratifiedKFold(n_splits=fold_num, random_state=i+1, shuffle=True)
# Non_nested parameter search and scoring
clf = GridSearchCV(estimator=est_classifcation, param_grid=tuned_param, cv=inner_cv, scoring=scoring)
clf.fit(X_data, Y_data)
# CV with parameter optimization
param_score = cross_val_score(clf.best_estimator_, X=X_data, y=Y_data, cv=outer_cv, scoring=scoring).mean()
if(param_score > max_score):
max_score = param_score
best_estimator = clf.best_estimator_
progress = (i+1)/num_trials*100
print(f'> progress = {progress}%')
return (max_score, best_estimator)
# -
dfFile = '../NBA_Ino_part/nba_preprocessed.csv'
dateStart = '2015-10-01'
dateEnd = '2018-04-30'
# X, Y = featureExtraction(dfFile, dateStart, dateEnd)
X, Y = featureExtraction(dfFile, attriToDrop=['PTS_A', 'PTS_B'], dateStart=dateStart, dateEnd=dateEnd)
X_val = X.values
Y_val = Y.values
# +
import lightgbm as lgb
from lightgbm import LGBMClassifier
from time import time
params = {'boosting_type': 'gbdt',
'max_depth' : -1,
'objective': 'binary',
'nthread': 5, # Updated from nthread
'num_leaves': 64,
'learning_rate': 0.05,
'max_bin': 512,
'subsample_for_bin': 200,
'subsample': 1,
'subsample_freq': 1,
'colsample_bytree': 0.8,
'reg_alpha': 5,
'reg_lambda': 10,
'min_split_gain': 0.5,
'min_child_weight': 1,
'min_child_samples': 5,
'scale_pos_weight': 1,
'num_class' : 1,
'metric' : 'binary_error',
'cat_smooth' : 10,
'objective' : 'binary',
'metric' : 'auc'}
# Set the parameters by cross-validation
tuned_parameters = {
'learning_rate': [2, 1.5, 1, 0.5, 0.25, 0.1, 0.05, 0.01, 0.005, 0.001],
'n_estimators': range(100, 1001, 100),
"max_depth": [3, 5, 8, 9, 11, 12, 13],
#'num_leaves': range(30, 100, 30),
'boosting_type' : ['gbdt'],
#'random_state' : [501], # Updated from 'seed'
'colsample_bytree' : [0.1, 0.65, 0.8, 1],
'subsample' : [0.1, 0.5, 0.75, 1]
#'reg_alpha' : [1, 1.2],
#'reg_lambda' : [1, 1.2, 1.4],
}
# Number of random trials
NUM_TRIALS = 1
# We will use a Support Vector Classifier with "rbf" kernel
#lgbm = LGBMClassifier()
lgbm = LGBMClassifier()
start = time()
(max_score, lgbm_best_estimator) = CrossValidationGridSearchNested(X_val, Y_val.ravel(), NUM_TRIALS, 10, lgbm, tuned_parameters, 'roc_auc')
lgbm_best_parameter = lgbm_best_estimator.get_params()
print("CrossValidationGridSearchNested of LightGradientBoostingClassifier wih NUM_TRIALS = %2.0d took %.2f seconds."%(NUM_TRIALS, (time() - start)))
print(f'\nmax_score = {max_score}\n')
print(f'\nbest_estimator = {lgbm_best_estimator}\n')
print(f'\nbest_parameter = {lgbm_best_parameter}\n')
# -
| LightGBM_Grid_Search_CV_3years.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import turtle
def polygon(inicial_angle,n,size):
angle = 360/n
turtle.left(inicial_angle)
for i in range(n):
turtle.forward(size)
turtle.left(angle)
# +
ang=10
n=9
size=40
step=80
turtle.hideturtle()
turtle.pensize(0.1)
turtle.bgcolor("black")
turtle.pencolor('red')
for i in range(int(360/ang)):
polygon(ang,n,size)
turtle.penup()
turtle.forward(step)
turtle.pendown()
turtle.pencolor('orange')
for i in range(int(360/ang)):
polygon(ang,n,size)
turtle.penup()
turtle.right(120)
turtle.forward(step)
turtle.pendown()
turtle.pencolor('yellow')
for i in range(int(360/ang)):
polygon(ang,n,size)
turtle.done()
# -
| Fun Rosettes (1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example Model Servers with Seldon
# ## Setup Seldon Core
#
# Use the setup notebook to [Setup Cluster](seldon_core_setup.ipynb#Setup-Cluster) with [Ambassador Ingress](seldon_core_setup.ipynb#Ambassador) and [Seldon Core](seldon_core_setup.ipynb#Install-Seldon-Core). Instructions [also online](./seldon_core_setup.html).
# ## Serve SKlearn Iris Model
# !pygmentize ../servers/sklearnserver/samples/iris.yaml
# !kubectl apply -f ../servers/sklearnserver/samples/iris.yaml
# !kubectl rollout status deploy/iris-default-4903e3c
from seldon_core.seldon_client import SeldonClient
sc = SeldonClient(deployment_name="sklearn",namespace="seldon")
r = sc.predict(gateway="ambassador",transport="rest",shape=(1,4))
print(r)
# !kubectl delete -f ../servers/sklearnserver/samples/iris.yaml
# ## Serve XGBoost Iris Model
# !pygmentize ../servers/xgboostserver/samples/iris.yaml
# !kubectl apply -f ../servers/xgboostserver/samples/iris.yaml
# !kubectl rollout status deploy/iris-default-af1783b
from seldon_core.seldon_client import SeldonClient
sc = SeldonClient(deployment_name="xgboost",namespace="seldon")
r = sc.predict(gateway="ambassador",transport="rest",shape=(1,4))
print(r)
# !kubectl delete -f ../servers/xgboostserver/samples/iris.yaml
# ## Serve Tensorflow MNIST Model
#
# **Will only work on a GCP Kubernetes Cluster**
# !pygmentize ../servers/tfserving/samples/mnist_rest.yaml
# !kubectl apply -f ../servers/tfserving/samples/mnist_rest.yaml
# !kubectl rollout status deploy/mnist-default-725903e
from seldon_core.seldon_client import SeldonClient
sc = SeldonClient(deployment_name="tfserving",namespace="seldon")
r = sc.predict(gateway="ambassador",transport="rest",shape=(1,784))
print(r)
# !kubectl delete -f ../servers/tfserving/samples/mnist_rest.yaml
# ## Serve Tensorflow MNIST Model with Tensorflow protocol
#
# !pygmentize ../servers/tfserving/samples/halfplustwo_rest.yaml
# !kubectl apply -f ../servers/tfserving/samples/halfplustwo_rest.yaml
# !kubectl rollout status deploy/hpt-default-a79d958
# !curl -d '{"instances": [1.0, 2.0, 5.0]}' \
# -X POST http://localhost:8003/seldon/seldon/hpt/v1/models/halfplustwo/:predict \
# -H "Content-Type: application/json"
# !kubectl delete -f ../servers/tfserving/samples/halfplustwo_rest.yaml
# ## Serve MLFlow Elasticnet Wines Model
# !pygmentize ../servers/mlflowserver/samples/elasticnet_wine.yaml
# !kubectl apply -f ../servers/mlflowserver/samples/elasticnet_wine.yaml
# !kubectl rollout status deploy/wines-default-8c791aa
from seldon_core.seldon_client import SeldonClient
sc = SeldonClient(deployment_name="mlflow",namespace="seldon")
r = sc.predict(gateway="ambassador",transport="rest",shape=(1,11))
print(r)
# !kubectl delete -f ../servers/mlflowserver/samples/elasticnet_wine.yaml
| notebooks/server_examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# We want to make sure not just the code we open-sourced, but also goes to dataset, so everyone can validate.
#
# You can check in [Malaya-Dataset](https://github.com/huseinzol05/Malaya-Dataset) for our open dataset.
# ## [Article](https://github.com/huseinzol05/Malaya-Dataset/blob/master/articles)
#
# Total size: 3.1 MB
#
# 1. Filem
# 2. Kerajaan
# 3. Pembelajaran
# 4. Pendidikan
# 5. Sekolah
# ## [Dependency](https://github.com/huseinzol05/Malaya-Dataset/blob/master/dependency)
# ## [Dictionary, 24550 unique words](https://github.com/huseinzol05/Malaya-Dataset/blob/master/dictionary)
# ## [Emotion](https://github.com/huseinzol05/Malaya-Dataset/blob/master/emotion)
#
# Total size: 8.5 MB
#
# 1. Anger
# 2. Fear
# 3. Joy
# 4. Love
# 5. Sadness
# 6. Surprise
# ## [Gender](https://github.com/huseinzol05/Malaya-Dataset/blob/master/gender)
#
# 1. Unknown
# 2. Male
# 3. Female
# 4. Brand
# ## [Irony](https://github.com/huseinzol05/Malaya-Dataset/blob/master/irony)
#
# Total size: 100 KB
#
# 1. Positive
# 2. Negative
# ## [Entities, JSON](https://github.com/huseinzol05/Malaya-Dataset/blob/master/entities)
#
# Total size: 1.1 MB
#
# 1. OTHER - Other
# 2. law - law, regulation, related law documents, documents, etc
# 3. location - location, place
# 4. organization - organization, company, government, facilities, etc
# 5. person - person, group of people, believes, etc
# 6. quantity - numbers, quantity
# 7. time - date, day, time, etc
# 8. event - unique event happened, etc
# ## [Karangan sekolah](https://github.com/huseinzol05/Malaya-Dataset/blob/master/karangan-sekolah)
#
# Total size: 221 KB
# ## [Language-detection, Wikipedia](https://github.com/huseinzol05/Malaya-Dataset/blob/master/language-detection)
# ## [News, crawled](https://github.com/huseinzol05/Malaya-Dataset/blob/master/news)
#
# Total size: 28.9 MB
#
# <details><summary>Complete list (51 news)</summary>
#
# 1. Cuti sekolah
# 2. isu 1MDB
# 3. isu agama
# 4. isu agong
# 5. isu agrikultur
# 6. isu air
# 7. isu anwar ibrahim
# 8. isu artis
# 9. isu astro
# 10. isu bahasa melayu
# 11. isu barisan nasional
# 12. isu cikgu
# 13. isu cukai
# 14. isu cyberjaya
# 15. isu dunia
# 16. isu ekonomi
# 17. isu gst
# 18. isu harakah
# 19. isu harga
# 20. isu icerd
# 21. isu imigren
# 22. isu kapitalis
# 23. isu kerajaan
# 24. isu kesihatan
# 25. isu kuala lumpur
# 26. isu lgbt
# 27. isu mahathir
# 28. isu makanan
# 29. isu malaysia airlines
# 30. isu malaysia
# 31. isu minyak
# 32. isu isu najib razak
# 33. isu pelajar
# 34. isu pelakon
# 35. isu pembangkang
# 36. isu perkauman
# 37. isu permainan
# 38. isu pertanian
# 39. isu politik
# 40. isu rosmah
# 41. isu sabah
# 42. isu sarawak
# 43. isu sosial media
# 44. isu sultan melayu
# 45. isu teknologi
# 46. isu TM
# 47. isu ubat
# 48. isu universiti
# 49. isu wan azizah
# 50. peluang pekerjaan
# 51. perkahwinan
#
# </details>
# ## [Sentiment News](https://github.com/huseinzol05/Malaya-Dataset/blob/master/news-sentiment)
#
# Total size: 496 KB
#
# 1. Positive
# 2. Negative
# ## [Sentiment Twitter](https://github.com/huseinzol05/Malaya-Dataset/blob/master/twitter-sentiment)
#
# Total size: 27.4 MB
#
# 1. Positive
# 2. Negative
# ## [Sentiment Multidomain](https://github.com/huseinzol05/Malaya-Dataset/blob/master/multidomain-sentiment)
#
# 159 KB
#
# 1. Amazon review, Positive and Negative
# 2. IMDB review, Positive and Negative
# 3. Yelp review, Positive and Negative
# ## [Part-of-Speech](https://github.com/huseinzol05/Malaya-Dataset/blob/master/part-of-speech)
#
# Total size: 3.1 MB
#
# 1. ADJ - Adjective, kata sifat
# 2. ADP - Adposition
# 3. ADV - Adverb, kata keterangan
# 4. ADX - Auxiliary verb, kata kerja tambahan
# 5. CCONJ - Coordinating conjuction, kata hubung
# 6. DET - Determiner, kata penentu
# 7. NOUN - Noun, kata nama
# 8. NUM - Number, nombor
# 9. PART - Particle
# 10. PRON - Pronoun, kata ganti
# 11. PROPN - Proper noun, kata ganti nama khas
# 12. SCONJ - Subordinating conjunction
# 13. SYM - Symbol
# 14. VERB - Verb, kata kerja
# 15. X - Other
# ## [Polarity](https://github.com/huseinzol05/Malaya-Dataset/blob/master/polarity)
#
# Total size: 1.3 MB
#
# 1. Positive
# 2. Negative
# ## [Political landscape](https://github.com/huseinzol05/Malaya-Dataset/blob/master/political-landscape)
#
# Total size: 2 MB
#
# 1. Kerajaan
# 2. Pembangkang
# ## [Sarcastic news-headline](https://github.com/huseinzol05/Malaya-Dataset/blob/master/sarcastic-news-headline)
#
# 1. Positive
# 2. Negative
# ## [Stemmer](https://github.com/huseinzol05/Malaya-Dataset/blob/master/stemmer)
#
# Total size: 6.5 MB
#
# 1. News stemming
# 2. Wikipedia stemming
# ## [Subjectivity](https://github.com/huseinzol05/Malaya-Dataset/blob/master/subjectivity)
#
# Total size: 1.4 MB
#
# 1. Positive
# 2. Negative
# ## [Subtitle](https://github.com/huseinzol05/Malaya-Dataset/blob/master/subtitle)
#
# Total size: 1.5 MB
# ## Suggestion
#
# 1. Always apply text augmentation, like swapping based words using synonyms or thesaurus. I still waiting respond from third-party to open source Bahasa thesaurus.
# ## Citation
#
# 1. Please citate the repository if use these corpus.
# 2. Please at least email us first before distributing these data. Remember all these hard workings we want to give it for free.
# 3. What do you see just the data, but nobody can see how much we spent our cost to make it public.
# ## Donation
#
# 1. We want to make sure downloaders got the best bandwidth and top speed, we host everything on S3, **please consider a donation** to prevent top-speed shutdown or broken link!
# 2. **Husein** really need money to stay survive, he is still a human. **7053174643, CIMB Click, <NAME>**
| dataset/dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # COVID-19 calculation
# Quick calculation based on problem posed by @3blue1brown on Twitter : https://twitter.com/3blue1brown/status/1333121058824613889?s=20
#
# > Say where you live, 1 in 1,000 actively have covid-19. You feel fatigued and have a slight sore throat, so you take a test, get a positive result.
# >
# > You learn the test has a 1% false positives, and 10% false negatives. What's your best guess for your chances of having covid-19?
#
# Let $C$ be the probability that a random member of the population has covid-19, $S$ be the probability that you are symptomatic, and $T$ be the probability that you return a positive test. We want to calculate
#
# \begin{align*}
# P(C|ST) &= \frac{P(C)P(S|C)P(T|C)}{P(C)P(S|C)P(T|C) + P(\bar{C})P(S|\bar{C})P(T|\bar{C})}
# \end{align*}
#
# We know $P(C) = 0.001$, $P(T|C) = 0.9$, and $P(T|\bar{C}) = 0.01$, so we can investigate the dependence on the two unknowns $P(S|C)$ and $P(S|\bar{C})$.
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import cm
# %matplotlib inline
# +
C = 1e-3
nC = 1 - C
T_C = 0.9
T_nC = 0.01;
S_C, S_nC = np.mgrid[0.0001:1:100j, 0.0001:1:100j]
C_ST = (T_C*S_C*C) / (C*S_C*T_C + nC*S_nC*T_nC)
# -
fig = plt.figure(figsize=(8, 10))
ax = fig.gca(projection='3d')
surf = ax.plot_surface(S_C, S_nC, C_ST, cmap = cm.hot)
ax.view_init(29, 45)
plt.xlabel('P(symptomatic | covid)')
plt.ylabel('P(symptomatic | not covid)')
plt.title('P(covid | symptomatic and positive test)')
| COVID-19 calculation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Demo Notebook
#
# This notebook shall walk through the minimal basic functionality of this package, echoing the `README`.
# import
from legomena import Corpus, HeapsModel, InfSeriesModel, LogModel
# ## Data Sources
#
# This package may be driven by any data source, but the author has tested two: the [Natural Language ToolKit](https://www.nltk.org/) and the [Standard Project Gutenberg Corpus](https://arxiv.org/abs/1812.08092). The former being the gold standard of python NLP applications, but having a rather measly 18-book gutenberg corpus. The latter containing the full 55,000+ book gutenberg corpus, already tokenized and counted. NOTE: The overlap of the two datasets do _not_ agree in their exact type/token counts, their methodology differing, but this package takes type/token counts as raw data and is therefore methodology-agnostic.
# +
# moby dick from NLTK
import nltk
nltk.download("gutenberg")
from nltk.corpus import gutenberg
words = gutenberg.words("melville-moby_dick.txt")
corpus = Corpus(words)
assert corpus.M, corpus.N == (260819, 19317)
# moby dick from SPGC
# NOTE: download and unzip https://zenodo.org/record/2422561/files/SPGC-counts-2018-07-18.zip into DATA_FOLDER
DATA_FOLDER = "../data"
import pandas as pd
fname = "%s/SPGC-counts-2018-07-18/PG2701_counts.txt" % DATA_FOLDER
with open(fname) as f:
df = pd.read_csv(f, delimiter="\t", header=None, names=["word", "freq"])
f.close()
wfd = {str(row.word): int(row.freq) for row in df.itertuples()}
corpus = Corpus(wfd)
assert corpus.M, corpus.N == (210258, 16402)
# a corpus is just a dictionary
corpus
# -
# ## Basic Usage
#
# +
# basic properties
corpus.tokens # list of tokens
corpus.types # list of types
corpus.fdist # word frequency distribution dataframe
corpus.WFD # alias for corpus.fdist
corpus.M # number of tokens
corpus.N # number of types
corpus.k # n-legomena vector
corpus.k[1] # n-legomena count (n=1 -> number of hapaxes)
corpus.hapax # list of hapax legomena, alias for corpus.nlegomena(1)
corpus.dis # list of dis legomena, alias for corpus.nlegomena(2)
corpus.tris # list of tris legomena, alias for corpus.nlegomena(3)
corpus.tetrakis # list of tetrakis legomena, alias for corpus.nlegomena(4)
corpus.pentakis # list of pentakis legomena, alias for corpus.nlegomena(5)
# advanced properties
corpus.options # tuple of optional settings
corpus.resolution # number of samples to take to calculate TTR curve
corpus.dimension # n-legomena vector length to pre-compute (max 6)
corpus.seed # random number seed for sampling TTR data
corpus.TTR # type-token ratio dataframe
# basic functions
corpus.nlegomena(n=1) # list of types occurring exactly n times
corpus.sample(m=999) # samples m tokens from corpus *without replacement*
corpus.sample(x=0.5) # samples proportion x of corpus *without replacement*
# -
# ## Type-Token Models
#
# There are a variety of models in the literature predicting number of types as a function of tokens, the most well-known being [Heap's Law](https://en.wikipedia.org/wiki/Heaps%27_law). Here are a few implemented, overlaid by the `Corpus` class.
# +
# three models
hmodel = HeapsModel() # Heap's Law
imodel = InfSeriesModel(corpus) # Infinite Series Model [1]
lmodel = LogModel() # Logarithmic Model [1]
# model fitting
m_tokens = corpus.TTR.m_tokens
n_types = corpus.TTR.n_types
lmodel.fit(m_tokens, n_types)
predictions = lmodel.fit_predict(m_tokens, n_types)
# model parameters
lmodel.params
# model predictions
predictions = lmodel.predict(m_tokens)
# log model only
dim = corpus.dimension
predicted_k = lmodel.predict_k(m_tokens, dim)
# -
# ## Model Comparison
# +
# visualize
import matplotlib.pyplot as plt
# heaps model
hpredictions = hmodel.fit_predict(m_tokens, n_types)
plt.scatter(m_tokens, n_types, label="observed")
plt.plot(m_tokens, hpredictions, label="predicted", color="red")
plt.title("Heap's Law")
plt.xlabel("tokens (corpus size)")
plt.ylabel("types (lexicon size)")
plt.show()
# logarithmic model
plt.scatter(m_tokens, n_types, label="observed")
plt.plot(m_tokens, predictions, label="predicted", color="red")
plt.title("Logarithmic Model")
plt.xlabel("tokens (corpus size)")
plt.ylabel("types (lexicon size)")
plt.show()
| notebooks/demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/jenish2/032_Jenish_Dholariya/blob/main/Lab_03_Predict_Play_Or_No.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="k7TQDbv5jZQ3" outputId="e10b4358-bcae-45aa-b1ad-9da1796b836d" colab={"base_uri": "https://localhost:8080/"}
from google.colab import drive
drive.mount("/content/drive")
# + id="SSsrxT7js7rY"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn import datasets, preprocessing
from sklearn.naive_bayes import GaussianNB, MultinomialNB
# + id="r0THsKTAtXJ9"
dataset = pd.read_csv("/content/drive/MyDrive/Semester7/ML/Lab03/Dataset1.csv")
# + id="CHXw4UlitaQ6" outputId="3380e617-632f-4bce-a269-88091289a5cc" colab={"base_uri": "https://localhost:8080/"}
# labelEncoder Object
label_encoder = preprocessing.LabelEncoder()
Y_rows=None
for data_heading in dataset:
if data_heading!="Play":
print(f"\n\nHeading :- {data_heading}")
#print(list(dataset[data_heading]))
dummy = pd.get_dummies(dataset[data_heading])
#print("\n\nDummy :\n",dummy)
dataset = dataset.drop([data_heading],axis=1)
dataset = pd.concat([dataset,dummy],axis=1)
#print("\n\nFinal Data :\n",dataset)
else:
Y_rows = label_encoder.fit_transform(dataset[data_heading])
dataset = dataset.drop([data_heading],axis=1)
# + id="A5sIMMI8t1Q1" outputId="ca0a004d-27f2-480f-a439-a44de00b82d9" colab={"base_uri": "https://localhost:8080/"}
print(dataset,Y_rows)
# + id="gGfgAmjZt4pV"
from sklearn.model_selection import train_test_split
X_train, X_test, Y_train, Y_test = train_test_split(dataset, Y_rows, test_size = 0.30, random_state = 32)
# + id="oO0Jk7Ypt9jh" outputId="61a0a1e5-41a3-4cd2-e558-603471076a08" colab={"base_uri": "https://localhost:8080/"}
# create model
model = MultinomialNB()
model.fit(X_train, Y_train)
# Predict Y from X_text
Y_predicted = model.predict(X_test)
print(X_test)
print(Y_predicted)
# + id="dGlnMnLct_9J" outputId="b01467a9-73e4-4d08-bfea-42fc912540ab" colab={"base_uri": "https://localhost:8080/"}
from sklearn import metrics
print(f"Accuracy is :- {metrics.accuracy_score(Y_test, Y_predicted)}")
# print precision and recall
from sklearn.metrics import precision_score
from sklearn.metrics import recall_score
precision = precision_score(Y_test, Y_predicted)
recall = recall_score(Y_test, Y_predicted)
print(f"precision :- {precision}")
print(f"recall :- {recall}")
# + id="PmXItWkGuDsp" outputId="b20cae03-f813-4845-af79-b768b6b5fb53" colab={"base_uri": "https://localhost:8080/"}
# Excersice
# Task1
# Temp = "Hot" and Weather = "overcast"
# 1 1 0
#Outlook(O,R,S)=0 1 0, Temp(C,H,M)=0 1 0, Humidity(High,Low,Normal)=0 0 1, Wind(F,T)=1,0, Play=0
output = model.predict([[0,1,0, 0,1,0 ,0,0,1 ,1,0]])
print(f"final prediction :- {output}")
#Overcast , High, Normal, False
output = model.predict([[1,0,0 ,0,1,0 ,0,0,1 ,1,0]])
print(f"final prediction :- {output}")
# Excersice
# Task1
# Overcast, Mild, Normal, True
output = model.predict([[1,0,0, 0,0,1 ,0,0,1 ,0,1]])
print(f"final prediction :- {output}")
# + id="amrewXDeuIvQ"
| Lab_03_Predict_Play_Or_No.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc="true"
# # Table of Contents
# <p><div class="lev1 toc-item"><a href="#Basic-usage" data-toc-modified-id="Basic-usage-1"><span class="toc-item-num">1 </span>Basic usage</a></div><div class="lev2 toc-item"><a href="#Some-methods" data-toc-modified-id="Some-methods-11"><span class="toc-item-num">1.1 </span>Some methods</a></div><div class="lev2 toc-item"><a href="#String-input" data-toc-modified-id="String-input-12"><span class="toc-item-num">1.2 </span>String input</a></div><div class="lev2 toc-item"><a href="#Instantiation-without-source-file" data-toc-modified-id="Instantiation-without-source-file-13"><span class="toc-item-num">1.3 </span>Instantiation without source file</a></div>
# -
# # Basic usage
from Py3SaxCCLIWrapper import SaxWrapper as SW
# Example XML file
with open('/Users/max/test.xml', 'r') as test:
print(test.read())
# Instantiate the object with an XML file to work on
PMS = SW.PyMiniSaxon(file='/Users/max/test.xml')
# XPath method applied on 'test.xml'
PMS.XPath('//b[@id="x"]')
# Count method applied on 'test.xml'
PMS.count('//b[@id="x"]')
# ## Some methods
# Example XML file
with open('/Users/max/test.xsl', 'r') as test:
print(test.read())
# XSLT method applied on 'test.xml'
PMS.XSLT('/Users/max/test.xsl')
# XPath method applied on string input
fromString = PMS.XSLT('/Users/max/test.xsl')
PMS.XPath('//b-extended/s',fromString=fromString)
PMS.findAllNodes('//b')
PMS.findAllTextNodesOf('//b')
# ## String input
fromString = """<a>
<b id="x">Eintrag <s>mit einem</s> X</b>
<b id="z">Eintrag mit Z</b>
<b id="x">Zweiter Eintrag mit X</b>
</a>"""
print(fromString)
# XSLT method applied on string input
PMS.XSLT('/Users/max/test.xsl', fromString=fromString)
# ## Instantiation without source file
# Instantiation of 'empty' object without specific XML file to work on
PMS_without_source_file = SW.PyMiniSaxon()
# XPath method on string input
PMS_without_source_file.XPath('//b[@id="x"]', fromString)
| .ipynb_checkpoints/Example-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Customer Churn Decision Tree
#
# Note: on top of `numpy`, `pandas`, `matplotlib`, you need two more packages for this assignment: `scikit-learn` (for building decision tree and other machine learning models) and
# `pydotplus` (for visualizing the decision trees), you want to add those packages to your `requirements.txt` file and install them before running this notebook.
# load packages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# ## Question 1
# - what's the shape of the dataframe created from the csv file?
# - how many features and what are their datatype?
# - are there any null values in the dataset?
# - create a histogram for the target feature and calculate the churn rate
# - if you want to make a "naive" prediction on customer churn rate, what's the rule?
# load the csv file
churn = pd.read_csv('customer-churn-example-simple.csv')
# create a deep copy of the df
df_train = churn.copy()
# check the basic info for the df
print(df_train.info())
print(df_train.shape)
# get first five rows.
df_train.head()
# change churn to string from boolean for histogram and decision tree modeling
df_train['churn'] = df_train['churn'].apply(str)
df_train.head()
# create a histogram for churn
df_train['churn'].hist()
# calculate the churn rate
a = df_train['churn'].value_counts()
print(a)
print(f'the churn rate is {a[1]/a.sum():.1%}')
# * There are total 3333 rows, and 7 columns.
# * There are 6 features. Their data types are object, integer, and float.
# * There is no null values in the dataset.
# * The churn rate is 14.5%.
# * If I want to make a navie prediction, I would say there is less likely will churn the customers.
# ## Task
# Build a decision tree:
#
# - use only 'account length', 'total day calls', 'customer service calls' features
# - use entropy as the criterion and set the max depth for the tree to 4
# - visualize the tree as shown in the sample code provided in the class.
# - explain the first node (root) of your generated tree (example each line in the node - what they mean)
# - make predictions for three customers based on the sample input found in the comments below and explain your predictions - do they make sense?
# - by looking at the decision tree image and change one value for customer 2 to flip the prediction and explain why
# build your DT using 'account length', 'total day calls', 'customer service calls' features
X = df_train[[ 'account length', 'total day calls', 'customer service calls' ]]
y = df_train['churn']
print(X.shape)
print(y.shape)
# train a DT model
from sklearn.tree import DecisionTreeClassifier
tree_clf = DecisionTreeClassifier()
tree_clf.fit(X, y)
# load the packages for visulizing the tree
from IPython.display import Image
from sklearn import tree
import pydotplus
# get feature and class names for visulization
print(X.columns.values.tolist())
print(y.unique().tolist())
cls_names = ['Not churn' if i == 0 else 'churn' for i in y.unique().tolist ()] # convert to string for class names
cls_names
tree_clf = DecisionTreeClassifier(criterion='entropy', max_depth=3)
tree_clf.fit(X, y)
# Create DOT data and visualize the tree
dot_data = tree.export_graphviz(tree_clf, feature_names=X.columns.values.tolist(),
class_names=cls_names, )
# Draw graph
graph = pydotplus.graph_from_dot_data(dot_data)
# Show graph
Image(graph.create_png())
# make predictions for the following customers
# customer 1 who open the account 25 days ago with total day calls of 100 and called customer support 5 times
# customer 2 who open the account 125 days ago with total day calls of 150 and called customer support 0 times
# customer 3 who open the account 600 days ago with total day calls of 1150 and called customer support 8 times
customer1 = tree_clf.predict([[25, 100, 5]])
customer2 = tree_clf.predict([[125, 150,0]])
customer3 = tree_clf.predict([[600, 1150,8]])
print(customer1,customer2,customer3)
| churn-dt-basics-starter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
class Vertex:
def __init__(self, name):
self.name = name
def getName(self):
return self.name
def __str__(self):
return self.name
class Edge:
def __init__(self, src, dst, weight = 0):
self.src = src
self.dst = dst
self.weight = weight
def getSrc(self):
return self.src
def getDst(self):
return self.dst
def getWeight(self):
return self.weight
def __str__(self):
return str(self.src) + "->" + str(self.dst)
class Digraph:
def __init__(self):
self.vertices = set([])
self.edges = {}
def addVertex(self, vertex):
if vertex.getName() in self.vertices:
raise ValueError('Duplicate Vertex')
else:
self.vertices.add(vertex)
self.edges[vertex] = []
def addEdge(self, edge):
src = edge.getSrc()
dst = edge.getDst()
if not(src in self.vertices and dst in self.vertices):
raise ValueError('Vertex not in graph')
self.edges[src].append(dst)
def childrenOf(self, vertex):
return self.edges[vertex]
def hasVertex(self, vertex):
return vertex in self.vertices
def __str__(self):
res = ''
for k in self.edges:
for d in self.edges[k]:
res = res + str(k) + '->' + str(d) + '\n'
return res[:-1]
class Graph(Digraph):
def addEdge(self, edge):
Digraph.addEdge(self, edge)
rev = Edge(edge.getDestination(), edge.getSource())
Digraph.addEdge(self, rev)
# +
def test1(kind):
vertices = []
for name in range(10):
vertices.append(Vertex(str(name)))
g = kind()
for n in vertices:
g.addVertex(n)
g.addEdge(Edge(vertices[0],vertices[1]))
g.addEdge(Edge(vertices[1],vertices[2]))
g.addEdge(Edge(vertices[2],vertices[3]))
g.addEdge(Edge(vertices[3],vertices[4]))
g.addEdge(Edge(vertices[3],vertices[5]))
g.addEdge(Edge(vertices[0],vertices[2]))
g.addEdge(Edge(vertices[1],vertices[1]))
g.addEdge(Edge(vertices[1],vertices[0]))
g.addEdge(Edge(vertices[4],vertices[0]))
print('The graph:')
print(g)
test1(Digraph)
##test1(Graph)
# -
| python/Graph Theory/Adjacency Lists.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook generates random synthetic fMRI data and a random behavioral regressor, and performs a standard univariate analysis to find correlations between the two. It is meant to demonstrate how easy it is to find seemingly impressive correlations with fMRI data when multiple tests are not properly controlled for.
#
# In order to run this code, you must first install the standard Scientific Python stack (e.g. using [anaconda](https://www.continuum.io/downloads)) along with following additional dependencies:
# * [nibabel](http://nipy.org/nibabel/)
# * [nilearn](http://nilearn.github.io)
# * [statsmodels](http://statsmodels.sourceforge.net)
# * [nipype](http://nipype.readthedocs.io/en/latest/)
#
# In addition, this notebook assumes that [FSL](http://fsl.fmrib.ox.ac.uk/fsl/fslwiki/) is installed and that the FSLDIR environment variable is defined.
#
# +
import numpy
import nibabel
import os
import nilearn.plotting
import matplotlib.pyplot as plt
from statsmodels.regression.linear_model import OLS
import nipype.interfaces.fsl as fsl
import scipy.stats
if not 'FSLDIR' in os.environ.keys():
raise Exception('This notebook requires that FSL is installed and the FSLDIR environment variable is set')
# %matplotlib inline
# -
# Set up default parameters. We use 28 subjects, which is the median sample size of the set of fMRI studies published in 2015 that were estimated from Neurosynth in the paper. We use a heuristic correction for multiple comparisons of p<0.001 and 10 voxels, like that show by Eklund et al. (2016, PNAS) to result in Type I error rates of 0.6-0.9.
pthresh=0.001 # cluster forming threshold
cthresh=10 # cluster extent threshold
nsubs=28 # number of subjects
# In order to recreate the figure from the paper exactly, we need to fix the random seed so that it will generate exactly the same random data. If you wish to generate new data, then set the recreate_paper_figure variable to False and rerun the notebook.
#
# +
recreate_paper_figure=False
if recreate_paper_figure:
seed=6636
else:
seed=numpy.ceil(numpy.random.rand()*100000).astype('int')
print(seed)
numpy.random.seed(seed)
# -
# Use the standard MNI152 2mm brain mask as the mask for the generated data
#
#
maskimg=os.path.join(os.getenv('FSLDIR'),'data/standard/MNI152_T1_2mm_brain_mask.nii.gz')
mask=nibabel.load(maskimg)
maskdata=mask.get_data()
maskvox=numpy.where(maskdata>0)
print('Mask includes %d voxels'%len(maskvox[0]))
# Generate a dataset for each subject. fMRI data within the mask are generated using a Gaussian distribution (mean=1000, standard deviation=100). Behavioral data are generated using a Gaussian distribution (mean=100, standard deviation=1).
#
#
# +
imgmean=1000 # mean activation within mask
imgstd=100 # standard deviation of noise within mask
behavmean=100 # mean of behavioral regressor
behavstd=1 # standard deviation of behavioral regressor
data=numpy.zeros((maskdata.shape + (nsubs,)))
for i in range(nsubs):
tmp=numpy.zeros(maskdata.shape)
tmp[maskvox]=numpy.random.randn(len(maskvox[0]))*imgstd+imgmean
data[:,:,:,i]=tmp
newimg=nibabel.Nifti1Image(data,mask.get_affine(),mask.get_header())
newimg.to_filename('fakedata.nii.gz')
regressor=numpy.random.randn(nsubs,1)*behavstd+behavmean
numpy.savetxt('regressor.txt',regressor)
# -
# Spatially smooth data using a 6 mm FWHM Gaussian kernel
# +
smoothing_fwhm=6 # FWHM in millimeters
smooth=fsl.IsotropicSmooth(fwhm=smoothing_fwhm,
in_file='fakedata.nii.gz',
out_file='fakedata_smooth.nii.gz')
smooth.run()
# -
# Use FSL's GLM tool to run a regression at each voxel
glm = fsl.GLM(in_file='fakedata_smooth.nii.gz',
design='regressor.txt',
out_t_name='regressor_tstat.nii.gz',
demean=True)
glm.run()
# Use FSL's cluster tool to identify clusters of activation that exceed the specified cluster-forming threshold
tcut=scipy.stats.t.ppf(1-pthresh,nsubs-1)
cl = fsl.Cluster()
cl.inputs.threshold = tcut
cl.inputs.in_file = 'regressor_tstat.nii.gz'
cl.inputs.out_index_file='tstat_cluster_index.nii.gz'
results=cl.run()
# Generate a plot showing the brain-behavior relation from the top cluster
# +
clusterimg=nibabel.load(cl.inputs.out_index_file)
clusterdata=clusterimg.get_data()
indices=numpy.unique(clusterdata)
clustersize=numpy.zeros(len(indices))
clustermean=numpy.zeros((len(indices),nsubs))
indvox={}
for c in range(1,len(indices)):
indvox[c]=numpy.where(clusterdata==c)
clustersize[c]=len(indvox[c][0])
for i in range(nsubs):
tmp=data[:,:,:,i]
clustermean[c,i]=numpy.mean(tmp[indvox[c]])
corr=numpy.corrcoef(regressor.T,clustermean[-1])
print('Found %d clusters exceeding p<%0.3f and %d voxel extent threshold'%(c,pthresh,cthresh))
print('Largest cluster: correlation=%0.3f, extent = %d voxels'%(corr[0,1],len(indvox[c][0])))
# set cluster to show - 0 is the largest, 1 the second largest, and so on
cluster_to_show=0
# translate this variable into the index of indvox
cluster_to_show_idx=len(indices)-cluster_to_show-1
# plot the (circular) relation between fMRI signal and
# behavioral regressor in the chosen cluster
plt.scatter(regressor.T,clustermean[cluster_to_show_idx])
plt.title('Correlation = %0.3f'%corr[0,1],fontsize=14)
plt.xlabel('Fake behavioral regressor',fontsize=18)
plt.ylabel('Fake fMRI data',fontsize=18)
m, b = numpy.polyfit(regressor[:,0], clustermean[cluster_to_show_idx], 1)
axes = plt.gca()
X_plot = numpy.linspace(axes.get_xlim()[0],axes.get_xlim()[1],100)
plt.plot(X_plot, m*X_plot + b, '-')
plt.savefig('scatter.png',dpi=600)
# -
# Generate a thresholded statistics image for display
tstat=nibabel.load('regressor_tstat.nii.gz').get_data()
thresh_t=clusterdata.copy()
cutoff=numpy.min(numpy.where(clustersize>cthresh))
thresh_t[thresh_t<cutoff]=0
thresh_t=thresh_t*tstat
thresh_t_img=nibabel.Nifti1Image(thresh_t,mask.get_affine(),mask.get_header())
# Generate a figure showing the location of the selected activation focus.
mid=len(indvox[cluster_to_show_idx][0])/2
coords=numpy.array([indvox[cluster_to_show_idx][0][mid],
indvox[cluster_to_show_idx][1][mid],
indvox[cluster_to_show_idx][2][mid],1]).T
mni=mask.get_qform().dot(coords)
nilearn.plotting.plot_stat_map(thresh_t_img,
os.path.join(os.getenv('FSLDIR'),'data/standard/MNI152_T1_2mm_brain.nii.gz'),
threshold=cl.inputs.threshold,
cut_coords=mni[:3])
plt.savefig('slices.png',dpi=600)
| analysis/multiplecomparisons/Correlation_simulation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Fluxonium
# ## Initialization (load modules etc.)
# + init_cell=true
# %matplotlib inline
# %config InlineBackend.figure_format = 'svg'
import itertools
import scqubits as qubit
import matplotlib
matplotlib.rcParams['text.usetex'] = False
matplotlib.rcParams.update({'font.size': 22})
import matplotlib.pyplot as plt
import numpy as np
import scipy.linalg as la
import scipy.constants as const
from qutip import *
from scqubits.utils.spectrum_utils import get_matrixelement_table, matrix_element
from scqubits.utils.sweep_plotting import bare_spectrum, dressed_spectrum, difference_spectrum
from scqubits import HilbertSpace, InteractionTerm, ParameterSweep
# +
J_to_GHz = 1. / (2. * np.pi * const.hbar * (10**9)) # (1/(2pi hbar 10^9))
GHz_to_J = 1./J_to_GHz
resonator = qubit.Oscillator(
E_osc = 5.7286,
truncated_dim=5
)
fluxonium = qubit.Fluxonium(
EJ = 3.395,
EL = 0.132,
EC = 0.479,
flux = 0.5,
cutoff = 110,
truncated_dim = 60
)
hilbertspc = HilbertSpace([fluxonium, resonator])
adag = resonator.creation_operator()
a = resonator.annihilation_operator()
int_term = InteractionTerm(
g_strength = 0.076,
subsys1 = fluxonium,
op1 = fluxonium.n_operator(),
subsys2 = resonator,
op2 = a+adag
)
interaction_list = [int_term]
hilbertspc.interaction_list = interaction_list
dressed_hamiltonian = hilbertspc.get_hamiltonian()
evals, evecs = dressed_hamiltonian.eigenstates(eigvals=10)
# +
param_name = 'flux' # name of varying external parameter
param_vals = np.linspace(0.25, 0.5, 100) # parameter values
subsys_update_list = [fluxonium] # list of HilbertSpace subsystems which are affected by parameter changes
def update_hilbertspace(param_val): # function that shows how Hilbert space components are updated
fluxonium.flux = param_val
sweep = ParameterSweep(
param_name=param_name,
param_vals=param_vals,
evals_count=50,
hilbertspace=hilbertspc,
subsys_update_list=subsys_update_list,
update_hilbertspace=update_hilbertspace
)
# -
dressed_spectrum(sweep, xlim=(0.3,0.4),ylim=(0,7))
# +
temp=0.05
def nthermal(omega):
return(1./(np.exp(omega*GHz_to_J/(const.k*temp))-1.))
def Pthermalprob(n, resonator):
return((1-np.exp(-resonator.E_osc*GHz_to_J/(const.k*temp)))
*np.exp(-n*resonator.E_osc*GHz_to_J/(const.k*temp)))
def gamma_purcell_sweep_old(i, j, sweep, kappa, k, flux_val):
#i -> j
evals_dressed, evecs_dressed = sweep.dressed_specdata.energy_table[k], sweep.dressed_specdata.state_table[k]
fluxonium.flux = flux_val
evals_fluxonium, evecs_fluxonium = fluxonium.eigensys(evals_count=20)
gammap = 0.0
if (evals_fluxonium[i] > evals_fluxonium[j]):
#n1 ranges over initial states of the resonator
for n1 in range(2):
#n2 over final
for n2 in range(2):
indexin1 = sweep.lookup.dressed_index((i,n1), param_index=k)
indexjn2 = sweep.lookup.dressed_index((j,n2), param_index=k)
gammap += (Pthermalprob(n1, resonator)
*np.abs(matrix_element(evecs_dressed[indexjn2],
sweep.hilbertspace.identity_wrap(resonator.annihilation_operator(),
resonator),evecs_dressed[indexin1]))**2)
omegaij = evals_fluxonium[i]-evals_fluxonium[j]
return(10**9 * kappa * (nthermal(omegaij) + 1) * gammap)
else:
#n1 ranges over initial states of the resonator
for n1 in range(2):
#n2 over final
for n2 in range(2):
indexin1 = sweep.lookup.dressed_index((i,n1), param_index=k)
indexjn2 = sweep.lookup.dressed_index((j,n2), param_index=k)
gammap += (Pthermalprob(n1, resonator)
*np.abs(matrix_element(evecs_dressed[indexjn2],
sweep.hilbertspace.identity_wrap(resonator.creation_operator(),
resonator),evecs_dressed[indexin1]))**2)
omegaij = evals_fluxonium[j]-evals_fluxonium[i]
return(10**9 * kappa * (nthermal(omegaij)) * gammap)
# -
def gamma_purcell_sweep(i, j, sweep, kappa, k, flux_val):
#i -> j
evals_dressed, evecs_dressed = sweep.dressed_specdata.energy_table[k], sweep.dressed_specdata.state_table[k]
fluxonium.flux = flux_val
evals_fluxonium, evecs_fluxonium = fluxonium.eigensys(evals_count=20)
gammap = 0.0
if (evals_fluxonium[i] > evals_fluxonium[j]):
for n1 in range(2):
for n2 in range(2):
indexin1 = sweep.lookup.dressed_index((i,n1), param_index=k)
indexjn2 = sweep.lookup.dressed_index((j,n2), param_index=k)
omegaij = np.abs(evals_dressed[indexin1]-evals_dressed[indexjn2])
gammap += (Pthermalprob(n1, resonator) * (nthermal(omegaij) + 1)
*np.abs(matrix_element(evecs_dressed[indexjn2],
hilbertspc.identity_wrap(resonator.annihilation_operator(),
resonator),evecs_dressed[indexin1]))**2)
return(10**9 * kappa * gammap)
else:
for n1 in range(2):
for n2 in range(2):
indexin1 = sweep.lookup.dressed_index((i,n1), param_index=k)
indexjn2 = sweep.lookup.dressed_index((j,n2), param_index=k)
omegaij = np.abs(evals_dressed[indexin1]-evals_dressed[indexjn2])
gammap += (Pthermalprob(n1, resonator) * (nthermal(omegaij))
*np.abs(matrix_element(evecs_dressed[indexjn2],
hilbertspc.identity_wrap(resonator.creation_operator(),
resonator),evecs_dressed[indexin1]))**2)
return(10**9 * kappa * gammap)
gamma_purcell_list = np.zeros(len(param_vals))
gamma_purcell_0up = np.zeros_like(gamma_purcell_list)
gamma_purcell_1up = np.zeros_like(gamma_purcell_list)
for k, flux_val in enumerate(param_vals):
gamma_purcell_list[k] = (gamma_purcell_sweep(0,1,sweep,0.00165, k, flux_val)
+ gamma_purcell_sweep(1,0,sweep,0.00165, k, flux_val))
#j ranges over higher fluxonium levels
gamma_purcell_0up[k] = np.sum([gamma_purcell_sweep(0,j,sweep,0.00165,k,flux_val) for j in range(1,15)])
gamma_purcell_1up[k] = np.sum([gamma_purcell_sweep(1,j,sweep,0.00165,k,flux_val) for j in range(2,15)])
blist = np.linspace(0.25, 0.5, 100)
hbard2kb = const.hbar/const.k/2
spect = fluxonium.get_spectrum_vs_paramvals('flux', blist, evals_count=2, subtract_ground=True)
spect_table = spect.energy_table.T[1]*2*np.pi*1e9
wq = spect_table
levls = 8
philist = []
nlist = []
sinphilist = []
for ii in blist:
fluxonium.flux = ii
phi = abs((fluxonium.matrixelement_table('phi_operator', evals_count=levls)[0, 1]))**2
n = abs((fluxonium.matrixelement_table('n_operator', evals_count=levls)[0, 1]))**2
# sinphi = abs((fluxonium.matrixelement_table('sinphi_operator', evals_count=levls)[0, 1]))**2
philist.append(phi)
nlist.append(n)
# sinphilist.append(sinphi)
philist = np.array(philist)
nlist = np.array(nlist)
# sinphilist = np.array(sinphilist)
# +
T_cap = 0.042
EC = 0.479e9*const.h
Gamma_cap = const.hbar*wq*wq/np.tanh(const.hbar*wq/const.k/T_cap/2)*philist/(8*EC/(9e-6))
# fig, ax = plt.subplots(1,1, figsize=(10,5))
# ax.set_yscale('log')
# ax.plot(blist,1/Gamma_cap*1e6)
print(1/Gamma_cap[-1]*1e6)
# +
T_ind = 0.042
EL = 0.132e9*const.h
Gamma_ind = 2*EL/const.hbar/np.tanh(const.hbar*wq/const.k/T_ind/2)*philist/(6e8)
# fig, ax = plt.subplots(1,1, figsize=(12,5))
# ax.set_yscale('log')
# ax.plot(blist,1/Gamma_ind*1e6)
print(1/Gamma_ind[-1]*1e6)
# +
t1_cap = 1/Gamma_cap
t1_ind = 1/Gamma_ind
T1total = 1/(Gamma_cap+Gamma_ind+gamma_purcell_list + gamma_purcell_0up + gamma_purcell_1up)
T1purcell = 1./(gamma_purcell_list + gamma_purcell_0up + gamma_purcell_1up)
T1purcell01 = 1./gamma_purcell_list
T1purcell0up = 1./gamma_purcell_0up
T1purcell1up = 1./gamma_purcell_1up
#T1purcellperturb = 1./gamma_purcell_perturb_list
fig=plt.figure(figsize=(12,5))
plt.yscale("log")
plt.plot(param_vals, T1total, 'k-')
plt.plot(param_vals, T1purcell, 'b:')
# plt.plot(param_vals, T1purcell01, color="yellow")
# plt.plot(param_vals, T1purcell0up, color="red")
# plt.plot(param_vals, T1purcell1up, color="green")
# plt.plot(param_vals, t1_charge, color="orange")
plt.plot(param_vals, t1_cap, "r--")
plt.plot(param_vals, t1_ind, "g-.")
#plt.plot(param_vals, T1purcellperturb, color="purple")
errlist = 10**(-6)*np.array([78.034, 70.57, 20.71, 20.93, 12.92, 66.93, 137.06, 319.19,
194.84, 146.87, 170.584, 374.582, 322.533, 125.10, 105.987, 58.612, 22.295, 11.29])
t1_data = 10**(-6)*np.array([1597.923, 1627.93, 301.86, 269.03, 476.33, 1783.19, 2131.76, 2634.50,
4364.68, 2587.82, 1661.915, 1794.468, 2173.88, 1188.83,
1576.493, 965.183, 560.251, 310.88])
t1_flux = np.array([0.26, 0.28, 0.32, 0.34, 0.36, 0.38, 0.4, 0.42, 0.44, 0.46, 0.465,
0.47, 0.475, 0.48, 0.484, 0.488, 0.492, 0.5])
plt.errorbar(t1_flux,t1_data,errlist,fmt=' ',ecolor='k')
plt.plot(t1_flux,t1_data,'ko')
# plt.legend(("T1purcell","T1purcell01","T1purcell0up","T1purcell1up","T1total","t1_charge","t1_cap","t1_ind","data"), fontsize=12)
plt.legend(("T1total","T1purcell","t1_cap","t1_ind","data"), fontsize=12)
plt.ylim(10**(-4),2.2*10**(-2))
plt.xlim(0.245,0.505)
# plt.xlabel("flux")
# plt.ylabel("s")
plt.show()
# fig.savefig("T1vsFlux.pdf")
# -
| nb/Fluxonium-essentials-Helin_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/zaidalyafeai/Notebooks/blob/master/Deep_GCN_Spam.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="UjoTbUQVnCz8" colab_type="code" colab={}
# !pip install --upgrade torch-scatter
# !pip install --upgrade torch-sparse
# !pip install --upgrade torch-cluster
# !pip install --upgrade torch-spline-conv
# !pip install torch-geometric
# + [markdown] id="xjY9vtO9MgoL" colab_type="text"
# 
# + [markdown] id="P3UffAf8M2Gw" colab_type="text"
# # Intorduction
# + [markdown] id="_4_eVOI2M4Uo" colab_type="text"
# PyTorch Geometric [PyG](https://github.com/rusty1s/pytorch_geometric) is a geometric deep learning (GDN) extension library for PyTorch. In general GDN is used to generalize deep learning for non-Ecludian data. For the most part, CNN doesn't work very good for 3D shapes, point clouds and graph structures. Moreover, many real life datasets are inherently non-ecludian like social communicatin datasets, molecular structures, network traffic . etc ...
#
# Graph convolutional networks (GCN) come to the rescue to generalize CNNs to work for non-ecludian datasets. The basic architecture is illustrated below
#
# 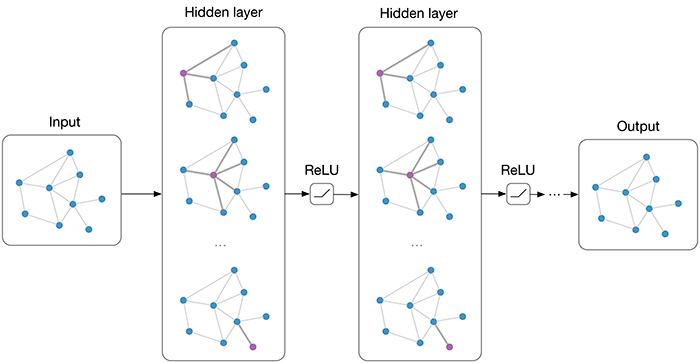
#
# where the input is a graph $G = (V,E)$ represented as
#
# * Feature repsentation for each node $N \times D$ where N is the number of nodes in the graph and $D$ is the number of features per node.
# * A matrix repsentation of the graph in the form $2\times L$ where $L$ is the number of edges in the graph. Each column in the matrix represents an edge between two nodes.
# * Edge attributes of the form $L \times R$ where R is the number of features per each edge.
#
# The output is of form $N \times F$ where $F$ is the number of features per each node in the graph.
#
#
#
# + id="YeA0slcJnQik" colab_type="code" colab={}
import numpy as np
import os.path as osp
import torch
import torch.nn.functional as F
from torch_geometric.nn import SplineConv
from torch_geometric.data import Data
from random import shuffle, randint
import networkx as nx
import matplotlib.pyplot as plt
import random
# + [markdown] id="6pQ-c3ftL_gp" colab_type="text"
# # Dataset
#
# We will simulate a spammer vs non-spammer graph network. Given a node which represents a client that can send emails to different node (another client).
#
# Spammers have some similarities
#
# * More likely to send lots of emails (more edges)
# * More likely to send lots of data through email (we will represent an edge feature is the number of bytes where the value [0, 1] where 1 represents more bytes sent)
# * Each spammer has an associated trust value which is given by the server. If the node is more likely to be a spammer then the value will be closer to 1.
#
# Non-spammers have the opposite features. In the next code snippet will try to simulate all of these features through randomization
#
#
# + id="MhlVjcdM7l6H" colab_type="code" colab={}
labels = []
N = 1000
nodes = range(0, N)
node_features = []
edge_features = []
for node in nodes:
#spammer
if random.random() > 0.5:
#more likely to have many connections with a maximum of 1/5 of the nodes in the graph
nb_nbrs = int(random.random() * (N/5))
#more likely to have sent many bytes
node_features.append((random.random()+1) / 2.)
#more likely to have a high trust value
edge_features += [(random.random()+2)/3.] * nb_nbrs
#associate a label
labels.append(1)
#non-spammer
else:
#at most connected to 10 nbrs
nb_nbrs = int(random.random() * 10 + 1)
#associate more bytes and random bytes
node_features.append(random.random())
edge_features += [random.random()] * nb_nbrs
labels.append(0)
#connect to some random nodes
nbrs = np.random.choice(nodes, size = nb_nbrs)
nbrs = nbrs.reshape((1, nb_nbrs))
#add the edges of nbrs
node_edges = np.concatenate([np.ones((1, nb_nbrs), dtype = np.int32) * node, nbrs], axis = 0)
#add the overall edges
if node == 0:
edges = node_edges
else:
edges = np.concatenate([edges, node_edges], axis = 1)
# + [markdown] id="qvfuQZv5lcM8" colab_type="text"
# Create a data structure
# + id="W1tyghgVFinu" colab_type="code" outputId="2c970876-76f9-4ef7-c8a5-<PASSWORD>" colab={"base_uri": "https://localhost:8080/", "height": 34}
x = torch.tensor(np.expand_dims(node_features, 1), dtype=torch.float)
y = torch.tensor(labels, dtype=torch.long)
edge_index = torch.tensor(edges, dtype=torch.long)
edge_attr = torch.tensor(np.expand_dims(edge_features, 1), dtype=torch.float)
data = Data(x = x, edge_index=edge_index, y =y, edge_attr=edge_attr )
print(data)
# + [markdown] id="bGcoGWzKlkHy" colab_type="text"
# We will create a trian/test mask where we split the data into training and test. This is necessary because during optimizing the loss when training we don't want to include the nodes part of the testing process
# + id="WRwBaYmyoLDX" colab_type="code" colab={}
data.train_mask = torch.zeros(data.num_nodes, dtype=torch.uint8)
data.train_mask[:int(0.8 * data.num_nodes)] = 1 #train only on the 80% nodes
data.test_mask = torch.zeros(data.num_nodes, dtype=torch.uint8) #test on 20 % nodes
data.test_mask[- int(0.2 * data.num_nodes):] = 1
# + [markdown] id="H2YFmL6kl5Dh" colab_type="text"
# # Deep GCN
#
# We will use [SplineConv](https://arxiv.org/abs/1711.08920) layer for the convolution. We will illsue exponential ReLU as an activation function and dropout for regulaization
# + id="MTlX4IBkoOnm" colab_type="code" colab={}
class Net(torch.nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = SplineConv(1, 16, dim=1, kernel_size=5)
self.conv2 = SplineConv(16, 32, dim=1, kernel_size=5)
self.conv3 = SplineConv(32, 64, dim=1, kernel_size=7)
self.conv4 = SplineConv(64, 128, dim=1, kernel_size=7)
self.conv5 = SplineConv(128, 128, dim=1, kernel_size=11)
self.conv6 = SplineConv(128, 2, dim=1, kernel_size=11)
def forward(self):
x, edge_index, edge_attr = data.x, data.edge_index, data.edge_attr
x = F.elu(self.conv1(x, edge_index, edge_attr))
x = self.conv2(x, edge_index, edge_attr)
x = F.elu(self.conv3(x, edge_index, edge_attr))
x = self.conv4(x, edge_index, edge_attr)
x = F.elu(self.conv5(x, edge_index, edge_attr))
x = self.conv6(x, edge_index, edge_attr)
x = F.dropout(x, training = self.training)
return F.log_softmax(x, dim=1)
# + [markdown] id="pULYL97tmYel" colab_type="text"
# # Optimization
#
# We will use nll_loss which can be used for classification of arbitrary classes
# + id="Hhabp4QvoP6V" colab_type="code" colab={}
def evaluate_loss(mode = 'train'):
#use masking for loss evaluation
if mode == 'train':
loss = F.nll_loss(model()[data.train_mask], data.y[data.train_mask])
else:
loss = F.nll_loss(model()[data.test_mask], data.y[data.test_mask])
return loss
def train():
#training
model.train()
optimizer.zero_grad()
loss = evaluate_loss()
loss.backward()
optimizer.step()
return loss.detach().cpu().numpy()
def test():
#testing
model.eval()
logits, accs = model(), []
loss = evaluate_loss(mode = 'test').detach().cpu().numpy()
for _, mask in data('train_mask', 'test_mask'):
pred = logits[mask].max(1)[1]
acc = pred.eq(data.y[mask]).sum().item() / mask.sum().item()
accs.append(acc)
return [loss] + accs
# + [markdown] id="y0XicLqpmqwR" colab_type="text"
# # Setup the model
# We will create the model and setup training using adam optimizer
# + id="sDvcl5eLoRb3" colab_type="code" colab={}
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
model, data = Net().to(device), data.to(device)
optimizer = torch.optim.Adam(model.parameters(), lr=1e-3)
# + [markdown] id="FyyfCGZimtX2" colab_type="text"
# # Training and Testing
# + id="qsslw_68oS52" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 3400} outputId="aa47e6c7-8985-4409-bfe5-805c0e3df3ae"
losses = []
for epoch in range(1, 200):
train_loss = train()
log = 'Epoch: {:03d}, train_loss: {:.3f}, test_loss:{:.3f}, train_acc: {:.2f}, test_acc: {:.2f}'
test_loss = test()[0]
losses.append([train_loss,test_loss])
print(log.format(epoch, train_loss, *test()))
# + [markdown] id="adWu02_enxNp" colab_type="text"
# # References
# [1] https://github.com/rusty1s/pytorch_geometric
#
# [2] https://rusty1s.github.io/pytorch_geometric/build/html/notes/introduction.html
#
# [3] https://tkipf.github.io/graph-convolutional-networks/
| Deep_GCN_Spam.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Axle-Bucamp/projet_robot_UI_WebAPP/blob/master/Stock_Market_V3_Keras_optimal_version.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="INiu-C68u19s"
# %%capture
import sys
import pandas as pd
import numpy as np
# parrameters
import warnings
warnings.filterwarnings('ignore')
# Choose your ticker
tickerSymbol = ["BTC-CAD","ETH-CAD","BTC-USD","CAD=X","CADEUR=X","CADUSD=X","XAD5.MI","GOOG","ARGX","^DJI",
"DOGE-USD","DOGE-EUR","EUR","USD","BTC","^CMC200",
"ETH-USD","ATOM-USD","ADA-USD","^GDAXI","^GSPC","^FCHI","ALGO-USD",
"ADA-EUR","ADA-CAD","SOL-CAD","^HSI",
"SOL-EUR","ETH-EUR","BTC-EUR"]
# Choose date range - format should be 'YYYY-MM-DD'
startDate = '2008-01-01' # as strings
endDate = '2021-12-31' # as strings
# Define key model parameters
# Set days out to predict
shifts = [14]
# Set a training percentage
train_pct = .85
# Plotting dimensions
w = 16 # width
h = 4 # height
# + id="6k_NgEcl_tQ2"
# + [markdown] id="utNHYCmOk95Y"
# ["BTC-USD","BTC-EUR","CAD=X","CADEUR=X","CADUSD=X","XAD5.MI","GOOG","ARGX","^DJI",
# "DOGE-USD","DOGE-EUR","EUR","USD","BTC","^CMC200",
# "ETH-USD","ATOM","ADA-USD",
# "ADA-EUR","ADA-CAD","SOL-CAD",
# "SOL-EUR","ETH-EUR","ETH-CAD","BTC-CAD"]
# + id="nVpoCeG-vjNi"
# %%capture
# Check if local computer has the library yfinance. If not, install. Then Import it.
# !python -m pip install yfinance # Check if the machine has yfinance, if not, download yfinance
import yfinance as yf # Import library to access Yahoo finance stock data
# Add financial information and indicators
# !python -m pip install ta # Download ta
from ta import add_all_ta_features # Library that does financial technical analysis
# Install fastai to use the date function
# !python -m pip install fastai # Download fastai
import fastai.tabular # Library that does date factors
# !python -m pip install plotly # Download plotly
import plotly.graph_objs as go # Import the graph ojbects
# Import sklearn modules that will help with modeling building
# !python -m pip install sklearn # Download sklearn
from sklearn.metrics import mean_squared_error # Install error metrics
from sklearn.linear_model import LinearRegression # Install linear regression model
from sklearn.preprocessing import StandardScaler # to scale for ann
# Go through each shift....
import pickle
from joblib import dump, load
from google.colab import files
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
from sklearn.preprocessing import MinMaxScaler
# + id="Vm3eZB_Zvnjp" colab={"base_uri": "https://localhost:8080/", "height": 991} outputId="c7f1ffe9-dbb1-4e66-b49b-b8cfcba50df2"
# Create ticker yfinance object
data = []
i = 0
for t in tickerSymbol:
print('----------------------- iteration :' + str(i))
tickerData = yf.Ticker(t)
# Create historic data dataframe and fetch the data for the dates given.
df = tickerData.history(start = startDate, end = endDate)
# to date format so we can get some juicy details
date_change = '%Y-%m-%d'
df['Date'] = df.index
df['Date'] = pd.to_datetime(df['Date'], format = date_change)
Dates = df['Date']
# add financial details
df = add_all_ta_features(df, "Open", "High", "Low", "Close", "Volume", fillna=True)
# Define the date parts
fastai.tabular.add_datepart(df,'Date', drop = 'True')
# Ensure the correct format
df['Date'] = pd.to_datetime(df.index.values, format = date_change)
# Add the date parts
fastai.tabular.add_cyclic_datepart(df, 'Date', drop = 'True')
data.append(df)
i +=1
# Print statement showing the download is done
# Show what the first 5 rows of the data frame
# Note the dataframe has:
# - Date (YYY-MM-DD) as an index
# - Open (price the stock started as)
# - High (highest price stock reached that day)
# - Low (lowest price stock reached that day)
# - Close (price the stock ended the day as)
# - Volume (how many shares were traded that day)
# - Dividends (any earnings shared to shareholders)
# - Stock Splits (any stock price changes)
# Create ticker yfinance object
print('-----------------------')
print('Done!')
data[0].head()
# + id="fP4HYv1Zyrcf"
# Ensure column types are correct
def CorrectColumnTypes(df):
# Input: dataframe
# ouptut: dataframe (with column types changed)
# Numbers
for col in df.columns[1:80]:
df[col] = df[col].astype('float')
for col in df.columns[-10:]:
df[col] = df[col].astype('float')
# Categories
for col in df.columns[80:-10]:
df[col] = df[col].astype('category').cat.codes
# Corriger le type,
# Soit transformer les cat en one hot soit
# Soit cat code sur tout le dataset
return df
# + id="rTPTWeaVwMTq"
# Create the lags
def CreateLags(df,lag_size):
# inputs: dataframe , size of the lag (int)
# ouptut: dataframe ( with extra lag column), shift size (int)
# add lag
shiftdays = lag_size
shift = -shiftdays
df['Close_lag'] = df['Close'].shift(shift)
return df, shift
# + id="th3vDEtWyvlv"
# Split the testing and training data
def SplitData(df, train_pct, shift):
# inputs: dataframe , training_pct (float between 0 and 1), size of the lag (int)
# ouptut: x train dataframe, y train data frame, x test dataframe, y test dataframe, train data frame, test dataframe
train_pt = int(len(df)*train_pct)
train = df.iloc[:train_pt,:]
test = df.iloc[train_pt:,:]
x_train = train.iloc[:shift,1:-1]
y_train = train['Close_lag'][:shift]
x_test = test.iloc[:shift,1:-1]
y_test = test['Close'][:shift]
return x_train, y_train, x_test, y_test, train, test
# + id="vB4Zrm2SyxHf"
# Function to make the plots
def PlotModelResults_Plotly(train, test, pred, ticker, w, h, shift_days,name,i):
# inputs: train dataframe, test dataframe, predicted value (list), ticker ('string'), width (int), height (int), shift size (int), name (string)
# output: None
# Create lines of the training actual, testing actual, prediction
D1 = go.Scatter(x=train.index,y=train['Close'],name = 'Train Actual') # Training actuals
D2 = go.Scatter(x=test.index[:shift],y=test['Close'],name = 'Test Actual') # Testing actuals
D3 = go.Scatter(x=test.index[:shift],y=pred,name = 'Our Prediction') # Testing predction
# Combine in an object
line = {'data': [D1,D2,D3],
'layout': {
'xaxis' :{'title': 'Date'},
'yaxis' :{'title': '$'},
'title' : name + ' - ' + tickerSymbol[i] + ' - ' + str(shift_days)
}}
# Send object to a figure
fig = go.Figure(line)
# Show figure
fig.show()
# + id="vzeII7gb0VAm"
# Regreesion Function
from sklearn import linear_model
from keras.callbacks import EarlyStopping,ModelCheckpoint,ReduceLROnPlateau
# + id="d7-fcVc00hMP"
def CalcProfit(test_df,pred,j):
pd.set_option('mode.chained_assignment', None)
test_df['pred'] = np.nan
test_df['pred'].iloc[:-j] = pred
test_df['change'] = test_df['Close_lag'] - test_df['Close']
test_df['change_pred'] = test_df['pred'] - test_df['Close']
test_df['MadeMoney'] = np.where(test_df['change_pred']/test_df['change'] > 0, 1, -1)
test_df['profit'] = np.abs(test['change']) * test_df['MadeMoney']
profit_dollars = test['profit'].sum()
print('Would have made: $ ' + str(round(profit_dollars,1)))
profit_days = len(test_df[test_df['MadeMoney'] == 1])
print('Percentage of good trading days: ' + str( round(profit_days/(len(test_df)-j),2)) )
return test_df, profit_dollars
# + id="J3ZTUMMs0ix9" colab={"base_uri": "https://localhost:8080/", "height": 435} outputId="4edeccb0-159f-4e7f-c8a6-4562d13923d5"
df.head()
# + id="K8zVs0L4eDxW"
def pgcd(a,b):
"""pgcd(a,b): calcul du 'Plus Grand Commun Diviseur' entre les 2 nombres entiers a et b"""
if b==0:
return a
else:
r=a%b
return pgcd(b,r)
# + id="fybpGmgG6jKw" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="c10fa96c-01af-40ea-e15c-ceb727f78e12"
from keras.utils.vis_utils import plot_model
#################################################################
inputs = keras.Input(shape=(20,110 ), name="digits")
norm = layers.BatchNormalization()(inputs)
Conv1 = layers.Conv1D(filters=64, kernel_size=10, activation='relu')(norm)
Conv2 = layers.Conv1D(filters=64, kernel_size=5, activation='relu')(Conv1)
Conv3 = layers.Conv1D(filters=64, kernel_size=3, activation='relu')(Conv2)
norm2 = layers.BatchNormalization()(Conv3)
Max2 = layers.MaxPooling1D(2)(norm2)
Flaterning = layers.Flatten()(Max2)
Dense1 = layers.Dense(64, activation="linear")(Flaterning)
Dense2 = layers.Dense(64, activation="linear")(Dense1)
Dense3 = layers.Dense(64, activation="linear")(Dense2)
Dense4 = layers.Dense(64, activation="linear")(Dense3)
outputs = layers.Dense(1, activation="linear", name="predictions")(Dense4)
#################################################################
model = keras.Model(inputs=inputs, outputs=outputs)
model.compile(
optimizer=keras.optimizers.Adam(
learning_rate=0.01), # Optimizer
# Loss function to minimize
loss=keras.losses.MeanSquaredError())
earlyStopping = EarlyStopping(monitor='val_loss', patience=15, verbose=0, mode='min')
checkpointer=ModelCheckpoint("weight", monitor='val_loss', verbose=0, save_best_only=True, save_weights_only=True, mode='auto', period=1),
reduce_lr_loss = ReduceLROnPlateau(monitor='val_loss', factor=0.1, patience=10, verbose=1, epsilon=1e-4, mode='min')
# summary du modèle
print(model.summary())
# graph
plot_model(model, to_file='multilayer_perceptron_graph.png')
# + id="rPkjptrH0k39" colab={"base_uri": "https://localhost:8080/"} outputId="2c1672f2-94bc-4048-a633-13252b0091c5"
i = 0
ShiftModelList = []
for j in shifts:
regList = []
i = 0
print('#####################################################################')
print(' days out: ' + str(j))
print()
print("prediction for values: " + tickerSymbol[i])
print('---------------------------------')
df = pd.concat(data)
df_lag, shift = CreateLags(df,j)
df_lag = CorrectColumnTypes(df_lag)
x_train, y_train, x_test, y_test, train, test = SplitData(df, train_pct, shift)
# Linear Regression
print("Linear Regression")
#################################################################
# inputs: x train data, y train data, x test data, y test data (all dataframe's)
# output: the predicted values for the test data (list)
# utiliser le model keras, normaliser les données
#################################################################
#for ltsm and convolutionnal network
Ymax = np.max(np.array(y_train))
pgcd= 20
if not x_train.shape[0]%pgcd == 0 :
x_train = x_train[:- (x_train.shape[0]%pgcd)]
y_train = y_train[:- (y_train.shape[0]%pgcd)]
if not x_test.shape[0]%pgcd == 0 :
x_test = x_test[:- (x_test.shape[0]%pgcd)]
y_test = y_test[:- (y_test.shape[0]%pgcd)]
x_train = np.array(x_train).reshape((int(x_train.shape[0]/pgcd), pgcd, x_train.shape[1]))
x_test = np.array(x_test).reshape((int(x_test.shape[0]/pgcd), pgcd, 110))
ya = []
for ii in range(len(y_train)):
if ii%20 == 0:
ya.append(y_train[ii])
y_train = np.array(ya) /Ymax
ya = []
for ii in range(len(y_test)):
if ii%20 == 0:
ya.append(y_test[ii])
y_test = np.array(ya) / Ymax
#for ltsm and convolutionnal network
#__________________________________________________________________#
model.fit(x_train,y_train,batch_size=50,epochs=100, shuffle=True,validation_data=(x_test,y_test), callbacks=[earlyStopping, reduce_lr_loss,checkpointer] )
lr_pred = model.predict( x_test).reshape(-1) * Ymax
#PlotModelResults_Plotly(train, test, lr_pred, tickerSymbol, w, h, j, 'Deep Learning',i)
regList.append(model)
i+=1
print('--------------------------------')
ShiftModelList.append(regList)
print('#####################################################################')
# + id="em9kHkH7qiKQ" colab={"base_uri": "https://localhost:8080/"} outputId="34eba35b-5238-4c58-e63e-80dcd4ab1eec"
print(lr_pred/Ymax)
print(y_test)
# + id="cf_SiOu_Jk-H" colab={"base_uri": "https://localhost:8080/", "height": 542} outputId="47cd9a05-2af9-4ddf-9148-ebec19922aad"
D2 = go.Scatter(x=np.array(range(lr_pred.shape[0])),y=y_test*Ymax,name = 'Test Actual') # Testing actuals
D3 = go.Scatter(x=np.array(range(lr_pred.shape[0])),y=lr_pred,name = 'Our Prediction') # Testing predction
# Combine in an object
line = {'data': [D2,D3],
'layout': {
'xaxis' :{'title': 'Date'},
'yaxis' :{'title': '$'},
'title' : "Deep Learning" + ' - ' + "Test " + ' - Day shift = ' + str(shifts)
}}
# Send object to a figure
fig = go.Figure(line)
# Show figure
fig.show()
| Stock_Market_V3_Keras_optimal_version.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
pd.options.display.float_format = '{:.5f}'.format
import numpy as np
np.set_printoptions(precision=4)
np.set_printoptions(suppress=True)
import warnings
warnings.filterwarnings("ignore")
import os.path
def path_base(base_name):
current_dir = os.path.abspath(os.path.join(os.getcwd(), os.pardir))
print(current_dir)
data_dir = current_dir.replace('notebook','data')
print(data_dir)
data_base = data_dir + '\\' + base_name
print(data_base)
return data_base
base = pd.read_csv(path_base('db_censo.csv'))
base.shape
base.head(3)
base.describe()
# # Previsores e classes
previsores = base.iloc[:,0:14].values
classe = base.iloc[:,14].values
# # Transformação de categóricos para numéricos
from sklearn.preprocessing import LabelEncoder, OneHotEncoder
from sklearn.compose import ColumnTransformer
labelencoder_previsores = LabelEncoder()
previsores[:, 1] = labelencoder_previsores.fit_transform(previsores[:, 1])
previsores[:, 3] = labelencoder_previsores.fit_transform(previsores[:, 3])
previsores[:, 5] = labelencoder_previsores.fit_transform(previsores[:, 5])
previsores[:, 6] = labelencoder_previsores.fit_transform(previsores[:, 6])
previsores[:, 7] = labelencoder_previsores.fit_transform(previsores[:, 7])
previsores[:, 8] = labelencoder_previsores.fit_transform(previsores[:, 8])
previsores[:, 9] = labelencoder_previsores.fit_transform(previsores[:, 9])
previsores[:, 13] = labelencoder_previsores.fit_transform(previsores[:, 13])
onehotencorder = ColumnTransformer(transformers=[("OneHot", OneHotEncoder(), [1,3,5,6,7,8,9,13])],remainder='passthrough')
previsores = onehotencorder.fit_transform(previsores).toarray()
previsores.shape
# type(previsores)
# # Deixar as variáveis na mesma escala
# Importante em algoritimos que usam distancia Euclidiana como o Kmeans
labelencorder_classe = LabelEncoder()
classe = labelencorder_classe.fit_transform(classe)
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
previsores = scaler.fit_transform(previsores)
# # Divisão em base de treinos e testes
from sklearn.model_selection import train_test_split
previsores_treinamento, previsores_teste, classe_treinamento, classe_teste = train_test_split(previsores, classe, test_size=0.15, random_state=0)
print(previsores_treinamento.shape)
print(classe_treinamento.shape)
print(previsores_teste.shape)
print(classe_teste.shape)
#importação da biblioteca
#criacao do classificador
from sklearn.ensemble import RandomForestClassifier
classificador = RandomForestClassifier(criterion='entropy',n_estimators=50,random_state=0)
classificador.fit(previsores_treinamento,classe_treinamento)
previsoes = classificador.predict(previsores_teste)
from sklearn.metrics import confusion_matrix,accuracy_score
precisao = accuracy_score(classe_teste,previsoes)
matriz = confusion_matrix(classe_teste,previsoes)
precisao
matriz
| notebook/_Tree/RandomForest_Censo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.cluster import KMeans
from sklearn.svm import SVC
from sklearn.metrics import roc_auc_score, roc_curve
from sklearn import preprocessing
from sklearn.linear_model import LogisticRegression
import warnings
import numpy as np
from collections import OrderedDict
from lob_data_utils import lob, db_result
from lob_data_utils.svm_calculation import lob_svm
sns.set_style('whitegrid')
warnings.filterwarnings('ignore')
# -
data_length = 15000
stock = '9265'
d, d_cv, d_test = lob.load_prepared_data(
stock, data_dir='../queue_imbalance/data/prepared', cv=True, length=data_length)
d.head()
d_test.head()
# +
## Data visualization
# -
lob.plot_density_imbalance_vs_mid(d, 0, len(d))
# ## Logistic
log_clf = lob.logistic_regression(d, 0, len(d))
pred = log_clf.predict(d_test['queue_imbalance'].values.reshape(-1, 1))
lob.plot_roc(d_test, log_clf, stock=stock, label='')
# ## SVM
# +
gammas = [0.0005, 0.005, 1, 5, 50, 500, 5000]
cs = [0.0005, 0.005, 1, 5.0, 50, 500, 1000]
coef0s = [0, 0.0005, 0.005, 1, 5, 50, 500, 5000]
try:
df_svm_res = pd.read_csv('res_svm_{}_{}.csv'.format(stock, data_length))
print('Results read from file')
except FileNotFoundError:
print('Results file does not exist yet')
df_svm_res = pd.DataFrame(columns=['svm', 'c', 'gamma', 'coef0', 'roc_cv_score', 'roc_train_score'])
s = stock
svm_results = []
for c in cs:
for g in gammas:
for coef0 in coef0s:
if np.any(df_svm_res[df_svm_res['c']
== c][df_svm_res['gamma'] == g][df_svm_res['coef0']
== coef0][df_svm_res['svm'] =='sigmoid']):
continue
svm = lob_svm.SVMSigmoid(s, d, c=c, coef0=coef0, gamma=g, data_length=data_length)
cv_score = svm.predict(d_cv, 'cv', check=False)
train_score = svm.predict(d, 'train', check=False)
svm_results.append({'svm': 'sigmoid', 'c': c, 'coef0': coef0, 'gamma': g,
'roc_cv_score': cv_score, 'roc_train_score': train_score})
if np.any(df_svm_res[df_svm_res['c'] == c][df_svm_res['gamma'] == g][df_svm_res['svm'] =='rbf']):
continue
svm = lob_svm.SVMRbf(s, d, c=c, gamma=g, data_length=data_length)
cv_score = svm.predict(d_cv, 'cv', check=False)
train_score = svm.predict(d, 'train', check=False)
svm_results.append({'svm': 'rbf', 'c': c, 'gamma': g,
'roc_cv_score': cv_score, 'roc_train_score': train_score})
if np.any(df_svm_res[df_svm_res['c'] == c][df_svm_res['svm'] =='linear']):
continue
svm = lob_svm.SVMLinear(s, d, c=c, data_length=data_length)
cv_score = svm.predict(d_cv, 'cv', check=False)
train_score = svm.predict(d, 'train', check=False)
svm_results.append({'svm': 'linear', 'c': c, 'roc_cv_score': cv_score, 'roc_train_score': train_score})
pd.DataFrame(svm_results).to_csv('res_svm_{}_{}.csv'.format(stock, data_length))
for svm_result in svm_results:
df_svm_res = df_svm_res.append(svm_result, ignore_index=True)
df_svm_res.drop(columns=[c for c in df_svm_res.columns if 'Unnamed:' in c], inplace=True)
df_svm_res.sort_values(by='roc_cv_score', ascending=False)
df_svm_res.head()
# -
df_svm_res.to_csv('res_svm_{}_{}.csv'.format(stock, data_length))
df_svm_res.sort_values(by='roc_cv_score')
# ## GDF
# +
K = 50
def gdf_svm_classification(df, K, C=1000, gamma=1):
clf = SVC(kernel='rbf', C=C, gamma=gamma)
gdf_columns = ['gdf_' + str(i) for i in range(0, K)]
X = df.loc[:, gdf_columns]
y = df['mid_price_indicator'].values.reshape(-1, 1)
y[0] = 0
clf.fit(X, y)
return clf
length = data_length
rr = [0.01, 0.05, 0.1, 0.5, 1]
ss = [0.01, 0.05, 0.1, 0.5, 1]
results = []
try:
df_gdf_res = pd.read_csv('res_gdf_svm_{}_{}.csv'.format(stock,data_length))
print('Results read from file')
except FileNotFoundError:
print('Results file does not exist yet')
df_gdf_res = pd.DataFrame(columns=['svm', 'c', 'gamma', 'roc_cv_score', 'roc_train_score',
'K', 'r', 's'])
for r in rr:
for s in ss:
filename = 'gdf_{}_len{}_r{}_s{}_K{}'.format(stock, length, r, s, K)
dfs, dfs_cv, dfs_test = lob.load_prepared_data(
filename, data_dir='../gaussian_filter/data_gdf/', cv=True, length=length)
gdf_columns = ['gdf_' + str(i) for i in range(0, K)]
for C in [1, 10, 100, 1000, 10000]:
for gamma in [1, 10, 100, 1000, 10000]:
res = {}
res['c'] = C
res['gamma'] = gamma
res['r'] = r
res['s'] = s
res['stock'] = stock
res['K'] = K
res['svm'] = 'rbf'
if np.any(df_gdf_res[df_gdf_res['c']
== C][df_gdf_res['gamma']
== gamma][df_gdf_res['r']
== r][df_gdf_res['s']
== s][df_gdf_res['K']
== K][df_gdf_res['svm'] == 'rbf']):
continue
clf = gdf_svm_classification(dfs, K, C=C, gamma=gamma)
predictions = clf.predict(dfs.loc[:, gdf_columns])
try:
roc_train = roc_auc_score(predictions, dfs['mid_price_indicator'])
res['roc_train_score'] = roc_train
except Exception as e:
print(e, r, s, C, gamma)
predictions = clf.predict(dfs_cv.loc[:, gdf_columns])
try:
roc_cv = roc_auc_score(predictions, dfs_cv['mid_price_indicator'])
res['roc_cv_score'] = roc_cv
except Exception as e:
print(e, r, s, C, gamma)
results.append(res)
pd.DataFrame(results).to_csv('res_gdf_svm_{}_{}.csv'.format(stock, data_length))
for result in results:
df_gdf_res = df_gdf_res.append(result, ignore_index=True)
df_gdf_res.drop(columns=[c for c in df_gdf_res.columns if 'Unnamed:' in c], inplace=True)
# -
df_gdf_res.to_csv('res_gdf_svm_{}_{}.csv'.format(stock, data_length))
a = df_gdf_res[df_gdf_res['r'] == 0.1].sort_values(by='roc_cv_score', ascending=False).sort_values(by='s')
a[['s', 'roc_train_score', 'roc_cv_score']].plot(kind='bar', figsize=(16,16))
# ## GDF with logistic reg
# +
K = 50
def gdf_log_classification(df, K, C=1000):
gdf_columns = ['gdf_' + str(i) for i in range(0, K)]
clf = LogisticRegression(C=C)
X = df.loc[:, gdf_columns]
y = df['mid_price_indicator'].values.reshape(-1, 1)
y[0] = 0
clf.fit(X, y)
return clf
length = data_length
rr = [0.01, 0.05, 0.1, 0.5, 1]
ss = [0.01, 0.05, 0.1, 0.5, 1]
results = []
try:
df_gdf_log_res = pd.read_csv('res_gdf_log_{}_{}.csv'.format(stock, data_length))
print('Results read from file')
except FileNotFoundError:
print('Results file does not exist yet')
df_gdf_log_res = pd.DataFrame(columns=['c', 'roc_cv_score', 'roc_train_score', 'K', 'r', 's'])
for r in rr:
for s in ss:
filename = 'gdf_{}_len{}_r{}_s{}_K{}'.format(stock, length, r, s, K)
dfs, dfs_cv, dfs_test = lob.load_prepared_data(
filename, data_dir='../gaussian_filter/data_gdf/', cv=True, length=length)
gdf_columns = ['gdf_' + str(i) for i in range(0, K)]
for C in [1, 10, 100, 1000, 10000]:
res = {}
res['c'] = C
res['r'] = r
res['s'] = s
res['stock'] = stock
res['K'] = K
if np.any(df_gdf_log_res[df_gdf_log_res['c']
== C][df_gdf_log_res['r'] == r][df_gdf_log_res['s'] == s][df_gdf_log_res['K'] == K]):
continue
clf = gdf_log_classification(dfs, K, C=C)
predictions = clf.predict(dfs.loc[:, gdf_columns])
try:
roc_train = roc_auc_score(predictions, dfs['mid_price_indicator'])
res['roc_train_score'] = roc_train
except Exception as e:
print(e, r, s, C, gamma)
predictions = clf.predict(dfs_cv.loc[:, gdf_columns])
try:
roc_cv = roc_auc_score(predictions, dfs_cv['mid_price_indicator'])
res['roc_cv_score'] = roc_cv
except Exception as e:
print(e, r, s, C, gamma)
results.append(res)
pd.DataFrame(results).to_csv('res_gdf_log_{}_{}.csv'.format(stock, data_length))
for result in results:
df_gdf_log_res = df_gdf_log_res.append(result, ignore_index=True)
df_gdf_log_res.drop(columns=[c for c in df_gdf_log_res.columns if 'Unnamed:' in c], inplace=True)
# -
df_gdf_log_res.to_csv('res_gdf_log_{}_{}.csv'.format(stock, data_length))
df_gdf_log_res.sort_values(by='roc_cv_score', ascending=False).head(5)
df_gdf_res.sort_values(by='roc_cv_score', ascending=False).head(5)
# ## Results on test
best_gdf_res = df_gdf_res.sort_values(by='roc_cv_score', ascending=False).iloc[0]
best_gdf_log_res = df_gdf_log_res.sort_values(by='roc_cv_score', ascending=False).iloc[0]
best_svm_sig_res = df_svm_res[df_svm_res['svm'] == 'sigmoid'].sort_values(
by='roc_cv_score', ascending=False).iloc[0]
best_svm_rbf_res = df_svm_res[df_svm_res['svm'] == 'rbf'].sort_values(
by='roc_cv_score', ascending=False).iloc[0]
best_svm_lin_res = df_svm_res[df_svm_res['svm'] == 'linear'].sort_values(
by='roc_cv_score', ascending=False).iloc[0]
res_dict = OrderedDict({
'gdf_svm': best_gdf_res,
'gdf_log': best_gdf_log_res,
'svm_rbf': best_svm_rbf_res,
'svm_lin': best_svm_lin_res,
'svm_sig': best_svm_sig_res,
})
plt.bar(list(range(len(res_dict.keys()))), [v['roc_cv_score'] for v in res_dict.values()],)
d = plt.xticks(list(range(len(res_dict.keys()))), list(res_dict.keys()))
plt.title('CV score')
for k, v in res_dict.items():
print(k, v['roc_cv_score'])
plt.bar(list(range(len(res_dict.keys()))), [v['roc_train_score'] for v in res_dict.values()],)
d = plt.xticks(list(range(len(res_dict.keys()))), list(res_dict.keys()))
plt.title('train score')
for k, v in res_dict.items():
print(k, v['roc_train_score'])
list(res_dict.values())[0]
filename = 'gdf_{}_len{}_r{}_s{}_K{}'.format(stock, length, best_gdf_res['r'], best_gdf_res['s'],
int(best_gdf_res['K']))
dfs, dfs_cv, dfs_test = lob.load_prepared_data(
filename, data_dir='../gaussian_filter/data_gdf/', cv=True, length=length)
svm_gdf_clf = gdf_svm_classification(dfs, K, C=best_gdf_res['c'], gamma=best_gdf_res['gamma'])
gdf_columns = ['gdf_' + str(i) for i in range(0, K)]
pred_test = svm_gdf_clf.predict(dfs_test.loc[:, gdf_columns])
roc_test = roc_auc_score(pred_test, dfs_test['mid_price_indicator'])
best_gdf_res['roc_test_score'] = roc_test
roc_test
filename = 'gdf_{}_len{}_r{}_s{}_K{}'.format(
stock, length, best_gdf_log_res['r'], best_gdf_log_res['s'], int(best_gdf_log_res['K']))
dfs, dfs_cv, dfs_test = lob.load_prepared_data(
filename, data_dir='../gaussian_filter/data_gdf/', cv=True, length=length)
svm_gdf_clf = gdf_log_classification(dfs, K, C=best_gdf_res['c'])
gdf_columns = ['gdf_' + str(i) for i in range(0, K)]
pred_test = svm_gdf_clf.predict(dfs_test.loc[:, gdf_columns])
roc_test = roc_auc_score(pred_test, dfs_test['mid_price_indicator'])
best_gdf_log_res['roc_test_score'] = roc_test
roc_test
# +
d, d_cv, d_test = lob.load_prepared_data(stock,
data_dir='../queue_imbalance/data/prepared', cv=True, length=10000)
svm = lob_svm.SVMRbf(stock, d, c=best_svm_rbf_res['c'], gamma=best_svm_rbf_res['gamma'], data_length=data_length)
roc_test = svm.predict(d_test, 'test', check=False)
best_svm_rbf_res['roc_test_score'] = roc_test
svm = lob_svm.SVMSigmoid(stock, d, c=best_svm_sig_res['c'],
gamma=best_svm_sig_res['gamma'], coef0=best_svm_sig_res['coef0'])
roc_test = svm.predict(d_test, 'test', check=False)
best_svm_sig_res['roc_test_score'] = roc_test
svm = lob_svm.SVMLinear(stock, d, c=best_svm_lin_res['c'])
roc_test = svm.predict(d_test, 'test', check=False)
best_svm_lin_res['roc_test_score'] = roc_test
# +
plt.bar(list(range(len(res_dict.keys()))), [v['roc_train_score'] for v in res_dict.values()])
plt.bar(list(range(len(res_dict.keys()))), [v['roc_test_score'] for v in res_dict.values()])
d = plt.xticks(list(range(len(res_dict.keys()))), list(res_dict.keys()))
plt.title('test score')
for k, v in res_dict.items():
print(k, v['roc_test_score'])
# +
res = []
for k, v in res_dict.items():
dd = v.to_dict()
dd['type'] = k
res.append(dd)
df_res = pd.DataFrame(res)
df_res[['roc_train_score', 'roc_cv_score', 'roc_test_score']].plot(kind='bar', figsize=(8, 8))
d = plt.xticks(list(range(len(res_dict.keys()))), list(res_dict.keys()))
plt.legend(loc='upper right')
# -
| overview_val10/overview_9265.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Foundations of Computational Economics #13
#
# by <NAME>, ANU
#
# <img src="_static/img/dag3logo.png" style="width:256px;">
# + [markdown] slideshow={"slide_type": "fragment"}
# ## Two very important algorithms for solving equations
#
# <img src="_static/img/lab.png" style="width:64px;">
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src="_static/img/youtube.png" style="width:65px;">
#
# [https://youtu.be/gUdYS5PmvWo](https://youtu.be/gUdYS5PmvWo)
#
# Description: Bisections and Newton-Raphson methods. Solving equations of one variable. Accuracy of solution. Rates of convergence.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Classic algorithm for equation solving
#
# 1. Bisection method
# 1. Newton-Raphson method
#
#
# Solve equations of the form $ f(x) = 0 $
#
# Focus on the scalar case today.
# + [markdown] slideshow={"slide_type": "slide"}
# #### Bisection method for solving equations
#
# Solve equation $ f(x)=0 $, conditional on $ x \in [a,b] \subset \mathbb{R} $ such that $ f(a)f(b)<0 $
#
# Algorithm: similar to binary search, but in **continuous space**.
# + hide-output=false slideshow={"slide_type": "slide"}
def bisection(f,a=0,b=1,tol=1e-6,maxiter=100,callback=None):
'''Bisection method for solving equation f(x)=0
on the interval [a,b], with given tolerance and number of iterations.
Callback function is invoked at each iteration if given.
'''
pass
# + hide-output=false slideshow={"slide_type": "slide"}
f = lambda x: -4*x**3+5*x+1
a,b = -3,-.5 # upper and lower limits
x = bisection(f,a,b)
print('Solution is x=%1.3f, f(x)=%1.12f' % (x,f(x)))
# + hide-output=false slideshow={"slide_type": "slide"}
# make nice plot
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
plt.rcParams['figure.figsize'] = [12, 8]
xd = np.linspace(a,b,1000) # x grid
plt.plot(xd,f(xd),c='red') # plot the function
plt.plot([a,b],[0,0],c='black') # plot zero line
ylim=[f(a),min(f(b),0)]
plt.plot([a,a],ylim,c='grey') # plot lower bound
plt.plot([b,b],ylim,c='grey') # plot upper bound
def plot_step(x,**kwargs):
plot_step.counter += 1
plt.plot([x,x],ylim,c='grey')
plot_step.counter = 0 # new public attribute
bisection(f,a,b,callback=plot_step)
print('Converged in %d steps'%plot_step.counter)
plt.show()
# + hide-output=false slideshow={"slide_type": "slide"}
def bisection(f,a=0,b=1,tol=1e-6,maxiter=100,callback=None):
'''Bisection method for solving equation f(x)=0
on the interval [a,b], with given tolerance and number of iterations.
Callback function is invoked at each iteration if given.
'''
if f(a)*f(b)>0:
raise ValueError('Function has the same sign at the bounds')
for i in range(maxiter):
err = abs(b-a)
if err<tol: break
x = (a+b)/2
a,b = (x,b) if f(a)*f(x)>0 else (a,x)
if callback != None: callback(err=err,x=x,iter=i)
else:
raise RuntimeError('Failed to converge in %d iterations'%maxiter)
return x
# + [markdown] slideshow={"slide_type": "slide"}
# #### Newton-Raphson (Newton) method
#
# General form $ f(x)=0 $
#
# - Equation solving
# - Finding maximum/minimum based on FOC, then $ f(x)=Q'(x) $
#
#
# Algorithm:
# 1. Start with some good guess $ x_0 $ not too far from the solution
# 2. Newton step: $ x_{i+1} = x_i - \frac{f(x_i)}{f'(x_i)} $
# 3. Iterate until convergence in some metric
# + [markdown] slideshow={"slide_type": "slide"}
# #### Derivation for Newton method using Taylor series expansion
#
# $$
# f(x) = \sum_{k=0}^{\infty} \frac{f^{(k)}(x_0)}{k!} (x-x_0)^k
# $$
#
# Take first two terms, assume $ f(x) $ is solution, and let
# $ x_0=x_i $ and $ x=x_{i+1} $
#
# $$
# 0 = f(x) = f(x_i) + f'(x_i) (x_{i+1}-x_i) \quad \Rightarrow \quad x_{i+1} = x_i - \frac{f(x_i)}{f'(x_i)}
# $$
# + hide-output=false slideshow={"slide_type": "slide"}
def newton(fun,grad,x0,tol=1e-6,maxiter=100,callback=None):
'''Newton method for solving equation f(x)=0
with given tolerance and number of iterations.
Callback function is invoked at each iteration if given.
'''
pass
# + hide-output=false slideshow={"slide_type": "slide"}
f = lambda x: -4*x**3+5*x+1
g = lambda x: -12*x**2+5
x = newton(f,g,x0=-2.5,maxiter=7)
print('Solution is x=%1.3f, f(x)=%1.12f' % (x,f(x)))
# + hide-output=false slideshow={"slide_type": "slide"}
# make nice seriest of plots
a,b = -3,-.5 # upper and lower limits
xd = np.linspace(a,b,1000) # x grid
def plot_step(x0,x1,iter,**kwargs):
plot_step.counter += 1
if iter<10:
plt.plot(xd,f(xd),c='red') # plot the function
plt.plot([a,b],[0,0],c='black') # plot zero line
ylim = [min(f(b),0),f(a)]
plt.plot([x0,x0],ylim,c='grey') # plot x0
l = lambda z: g(x0)*(z - x1)
plt.plot(xd,l(xd),c='green') # plot the function
plt.ylim(bottom=10*f(b))
plt.title('Iteration %d'%(iter+1))
plt.show()
plot_step.counter = 0 # new public attribute
newton(f,g,x0=-2.5,callback=plot_step)
print('Converged in %d steps'%plot_step.counter)
# + hide-output=false slideshow={"slide_type": "slide"}
def newton(fun,grad,x0,tol=1e-6,maxiter=100,callback=None):
'''Newton method for solving equation f(x)=0
with given tolerance and number of iterations.
Callback function is invoked at each iteration if given.
'''
for i in range(maxiter):
x1 = x0 - fun(x0)/grad(x0)
err = abs(x1-x0)
if callback != None: callback(err=err,x0=x0,x1=x1,iter=i)
if err<tol: break
x0 = x1
else:
raise RuntimeError('Failed to converge in %d iterations'%maxiter)
return (x0+x1)/2
# + [markdown] slideshow={"slide_type": "slide"}
# #### Rate of convergence of the two methods
#
# - How fast does a solution method converge on the root of the equation?
# - Rate of convergence = the rate of decrease of the bias (difference between current guess and the solution)
# - Can be approximated by the rate of decrease of the error in the stopping criterion
#
#
# Bisections: **linear convergence**
#
# Newton: **quadratic convergence**
# + hide-output=false slideshow={"slide_type": "slide"}
def print_err(iter,err,**kwargs):
x = kwargs['x'] if 'x' in kwargs.keys() else kwargs['x0']
print('{:4d}: x = {:17.14f} err = {:8.6e}'.format(iter,x,err))
print('Newton method')
newton(f,g,x0=-2.5,callback=print_err,tol=1e-10)
print('Bisection method')
bisection(f,a=-3,b=-0.5,callback=print_err,tol=1e-10)
# + [markdown] slideshow={"slide_type": "slide"}
# #### Measuring complexity of Newton and bisection methods
#
# - What is the size of input $ n $?
# - Desired precision of the solution!
# - Thus, attention to the errors in the solution as algorithm proceeds
# - Rate of convergence is part of the computational complexity of the algorithms
# + [markdown] slideshow={"slide_type": "slide"}
# #### Computational complexity of Newton method
#
# - Calculating a root of a function f(x) **with n-digit precision**
# - Provided that a good initial approximation is known
# - Is $ O((logn)F(n)) $, where $ F(n) $ is the cost of
# - calculating $ f(x)/f'(x) $ with $ n $-digit precision
# + [markdown] slideshow={"slide_type": "slide"}
# ### Further learning resources
#
# - On computational complexity of Newton method
# [https://m.tau.ac.il/~tsirel/dump/Static/knowino.org/wiki/Newton’s_method.html#Computational_complexity](https://m.tau.ac.il/~tsirel/dump/Static/knowino.org/wiki/Newton's_method.html#Computational_complexity)
# - “Improved convergence and complexity analysis of Newton’s method for solving equations”
# [https://www.tandfonline.com/doi/abs/10.1080/00207160601173431?journalCode=gcom20](https://www.tandfonline.com/doi/abs/10.1080/00207160601173431?journalCode=gcom20)
| 13_bisections_newton.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Programmatic Access to Genome Nexus
# This notebook gives some examples in Python for programmatic access to http://genomenexus.org. You can run these examples after installing Jupyter. Easiest way for using Jupyter is installing the Python 3 version of anaconda: https://www.anaconda.com/download/. After having that you can install Jupyter with:
# ```
# conda install jupyter
# ```
# For these exampels we also require the Swagger API client reader [Bravado](https://github.com/Yelp/bravado). Unfortunately not yet available in anaconda, but you can get it through `pip`:
# ```
# conda install pip
# pip install bravado
# ```
# Let's try connecting to the Genome Nexus API now:
from bravado.client import SwaggerClient
client = SwaggerClient.from_url('https://www.genomenexus.org/v2/api-docs',
config={"validate_requests":False,"validate_responses":False})
print(client)
dir(client)
for a in dir(client):
client.__setattr__(a[:-len('-controller')], client.__getattr__(a))
variant = client.annotation.fetchVariantAnnotationGET(variant='17:g.41242962_41242963insGA').result()
dir(variant)
tc1 = variant.transcript_consequences[0]
dir(tc1)
print(tc1)
# ## Connect with cBioPortal API
# [cBioPortal](http://www.cbioportal.org) also uses Swagger for their API.
import seaborn as sns
# %matplotlib inline
sns.set_style("white")
sns.set_context('talk')
import matplotlib.pyplot as plt
cbioportal = SwaggerClient.from_url('https://www.cbioportal.org/api/api-docs',
config={"validate_requests":False,"validate_responses":False})
print(cbioportal)
for a in dir(cbioportal):
cbioportal.__setattr__(a.replace(' ', '_').lower(), cbioportal.__getattr__(a))
dir(cbioportal)
muts = cbioportal.mutations.getMutationsInMolecularProfileBySampleListIdUsingGET(
molecularProfileId="msk_impact_2017_mutations", # {study_id}_mutations gives default mutations profile for study
sampleListId="msk_impact_2017_all", # {study_id}_all includes all samples
projection="DETAILED" # include gene info
).result()
import pandas as pd
mdf = pd.DataFrame([dict(m.__dict__['_Model__dict'],
**m.__dict__['_Model__dict']['gene'].__dict__['_Model__dict']) for m in muts])
mdf.groupby('uniqueSampleKey').studyId.count().plot(kind='hist', bins=400, xlim=(0,30))
plt.xlabel('Number of mutations in sample')
plt.ylabel('Number of samples')
plt.title('Number of mutations across samples in MSK-IMPACT (2017)')
sns.despine(trim=True)
mdf.variantType.astype(str).value_counts().plot(kind='bar')
plt.title('Types of mutations in MSK-IMPACT (2017)')
sns.despine(trim=False)
# ## Annotate cBioPortal mutations with Genome Nexus
# For convenience sake we're using only SNVs here. Eventually there will be an endpoint to help convert pos, ref, alt to the hgvs notation.
snvs = mdf[(mdf.variantType == 'SNP') & (mdf.variantAllele != '-') & (mdf.referenceAllele != '-')].copy()
# need query string like 9:g.22125503G>C
snvs['hgvs_for_gn'] = snvs.chromosome.astype(str) + ":g." + snvs.startPosition.astype(str) + snvs.referenceAllele + '>' + snvs.variantAllele
assert(snvs['hgvs_for_gn'].isnull().sum() == 0)
# +
import time
qvariants = list(set(snvs.hgvs_for_gn))
gn_results = []
chunk_size = 500
print("Querying {} variants".format(len(qvariants)))
for n, qvar in enumerate([qvariants[i:i + chunk_size] for i in range(0, len(qvariants), chunk_size)]):
try:
gn_results += client.annotation.fetchVariantAnnotationPOST(variants=qvar,fields=['hotspots']).result()
print("Querying [{}, {}]: Success".format(n*chunk_size, min(len(qvariants), n*chunk_size+chunk_size)))
except Exception as e:
print("Querying [{}, {}]: Failed".format(n*chunk_size, min(len(qvariants), n*chunk_size+chunk_size)))
pass
time.sleep(1) # add a delay, to not overload server
# -
gn_dict = {v.id:v for v in gn_results}
# +
def is_sift_high(variant):
return variant in gn_dict and \
len(list(filter(lambda x: x.sift_prediction == 'deleterious', gn_dict[variant].transcript_consequences))) > 0
def is_polyphen_high(variant):
return variant in gn_dict and \
len(list(filter(lambda x: x.polyphen_prediction == 'probably_damaging', gn_dict[variant].transcript_consequences))) > 0
# -
# ## Check overlap SIFT/PolyPhen-2
snvs['is_sift_high'] = snvs.hgvs_for_gn.apply(is_sift_high)
snvs['is_polyphen_high'] = snvs.hgvs_for_gn.apply(is_polyphen_high)
# +
from matplotlib_venn import venn2
venn2(subsets=((snvs.is_sift_high & (~snvs.is_polyphen_high)).sum(),
(snvs.is_polyphen_high & (~snvs.is_sift_high)).sum(),
(snvs.is_polyphen_high & snvs.is_sift_high).sum()), set_labels=["SIFT","PolyPhen-2"])
plt.title("Variants as predicted to have a high impact in MSK-IMPACT (2017)")
| notebooks/genome_nexus_python_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # OOP Syntax Exercise - Part 2
#
# Now that you've had some practice instantiating objects, it's time to write your own class from scratch. This lesson has two parts. In the first part, you'll write a Pants class. This class is similar to the shirt class with a couple of changes. Then you'll practice instantiating Pants objects
#
# In the second part, you'll write another class called SalesPerson. You'll also instantiate objects for the SalesPerson.
#
# For this exercise, you can do all of your work in this Jupyter notebook. You will not need to import the class because all of your code will be in this Jupyter notebook.
#
# Answers are also provided. If you click on the Jupyter icon, you can open a folder called 2.OOP_syntax_pants_practice, which contains this Jupyter notebook ('exercise.ipynb') and a file called answer.py.
# # Pants class
#
# Write a Pants class with the following characteristics:
# * the class name should be Pants
# * the class attributes should include
# * color
# * waist_size
# * length
# * price
# * the class should have an init function that initializes all of the attributes
# * the class should have two methods
# * change_price() a method to change the price attribute
# * discount() to calculate a discount
# +
### TODO:
# - code a Pants class with the following attributes
# - color (string) eg 'red', 'yellow', 'orange'
# - waist_size (integer) eg 8, 9, 10, 32, 33, 34
# - length (integer) eg 27, 28, 29, 30, 31
# - price (float) eg 9.28
### TODO: Declare the Pants Class
### TODO: write an __init__ function to initialize the attributes
class Pants:
def __init__(self, pants_color, pants_waist_size, pants_length, pants_price):
self.color = pants_color
self.waist_size = pants_waist_size
self.length = pants_length
self.price = pants_price
### TODO: write a change_price method:
# Args:
# new_price (float): the new price of the shirt
# Returns:
# None
def change_price(self, new_price):
self.price = new_price
### TODO: write a discount method:
# Args:
# discount (float): a decimal value for the discount.
# For example 0.05 for a 5% discount.
#
# Returns:
# float: the discounted price
def discount(self, discount):
return self.price * (1 - discount)
# -
# # Run the code cell below to check results
#
# If you run the next code cell and get an error, then revise your code until the code cell doesn't output anything.
# +
def check_results():
pants = Pants('red', 35, 36, 15.12)
assert pants.color == 'red'
assert pants.waist_size == 35
assert pants.length == 36
assert pants.price == 15.12
pants.change_price(10) == 10
assert pants.price == 10
assert pants.discount(.1) == 9
print('You made it to the end of the check. Nice job!')
check_results()
# -
# # SalesPerson class
#
# The Pants class and Shirt class are quite similar. Here is an exercise to give you more practice writing a class. **This exercise is trickier than the previous exercises.**
#
# Write a SalesPerson class with the following characteristics:
# * the class name should be SalesPerson
# * the class attributes should include
# * first_name
# * last_name
# * employee_id
# * salary
# * pants_sold
# * total_sales
# * the class should have an init function that initializes all of the attributes
# * the class should have four methods
# * sell_pants() a method to change the price attribute
# * calculate_sales() a method to calculate the sales
# * display_sales() a method to print out all the pants sold with nice formatting
# * calculate_commission() a method to calculate the salesperson commission based on total sales and a percentage
# +
### TODO:
# Code a SalesPerson class with the following attributes
# - first_name (string), the first name of the salesperson
# - last_name (string), the last name of the salesperson
# - employee_id (int), the employee ID number like 5681923
# - salary (float), the monthly salary of the employee
# - pants_sold (list of Pants objects),
# pants that the salesperson has sold
# - total_sales (float), sum of sales of pants sold
### TODO: Declare the SalesPerson Class
### TODO: write an __init__ function to initialize the attributes
### Input Args for the __init__ function:
# first_name (str)
# last_name (str)
# employee_id (int)
# . salary (float)
#
# You can initialize pants_sold as an empty list
# You can initialize total_sales to zero.
#
###
class SalesPerson:
def __init__(self, sp_first_name, sp_last_name, sp_employee_id, sp_salary):
self.first_name = sp_first_name
self.last_name = sp_last_name
self.employee_id = sp_employee_id
self.salary = sp_salary
self.pants_sold = []
self.total_sales = 0
### TODO: write a sell_pants method:
#
# This method receives a Pants object and appends
# the object to the pants_sold attribute list
#
# Args:
# pants (Pants object): a pants object
# Returns:
# None
def sell_pants(self, pants_object):
self.pants_sold.append(pants_object)
### TODO: write a display_sales method:
#
# This method has no input or outputs. When this method
# is called, the code iterates through the pants_sold list
# and prints out the characteristics of each pair of pants
# line by line. The print out should look something like this
#
# color: blue, waist_size: 34, length: 34, price: 10
# color: red, waist_size: 36, length: 30, price: 14.15
#
#
#
###
def display_sales(self):
for pants in self.pants_sold:
print(f'color: {pants.color}, waist_size: {pants.waist_size}, length: {pants.length}, price: {pants.price}')
### TODO: write a calculate_sales method:
# This method calculates the total sales for the sales person.
# The method should iterate through the pants_sold attribute list
# and sum the prices of the pants sold. The sum should be stored
# in the total_sales attribute and then return the total.
#
# Args:
# None
# Returns:
# float: total sales
#
###
def calculate_sales(self):
total = 0
for pants in self.pants_sold:
total += pants.price
self.total_sales = total
return total
### TODO: write a calculate_commission method:
#
# The salesperson receives a commission based on the total
# sales of pants. The method receives a percentage, and then
# calculate the total sales of pants based on the price,
# and then returns the commission as (percentage * total sales)
#
# Args:
# percentage (float): comission percentage as a decimal
#
# Returns:
# float: total commission
#
#
###
def calculate_commission(self, percentage):
sales_total = self.calculate_sales()
return sales_total * percentage
# -
# # Run the code cell below to check results
#
# If you run the next code cell and get an error, then revise your code until the code cell doesn't output anything.
# +
def check_results():
pants_one = Pants('red', 35, 36, 15.12)
pants_two = Pants('blue', 40, 38, 24.12)
pants_three = Pants('tan', 28, 30, 8.12)
salesperson = SalesPerson('Amy', 'Gonzalez', 2581923, 40000)
assert salesperson.first_name == 'Amy'
assert salesperson.last_name == 'Gonzalez'
assert salesperson.employee_id == 2581923
assert salesperson.salary == 40000
assert salesperson.pants_sold == []
assert salesperson.total_sales == 0
salesperson.sell_pants(pants_one)
salesperson.pants_sold[0] == pants_one.color
salesperson.sell_pants(pants_two)
salesperson.sell_pants(pants_three)
assert len(salesperson.pants_sold) == 3
assert round(salesperson.calculate_sales(),2) == 47.36
assert round(salesperson.calculate_commission(.1),2) == 4.74
print('Great job, you made it to the end of the code checks!')
check_results()
# -
# ### Check display_sales() method
#
# If you run the code cell below, you should get output similar to this:
#
# ```python
# color: red, waist_size: 35, length: 36, price: 15.12
# color: blue, waist_size: 40, length: 38, price: 24.12
# color: tan, waist_size: 28, length: 30, price: 8.12
# ```
# +
pants_one = Pants('red', 35, 36, 15.12)
pants_two = Pants('blue', 40, 38, 24.12)
pants_three = Pants('tan', 28, 30, 8.12)
salesperson = SalesPerson('Amy', 'Gonzalez', 2581923, 40000)
salesperson.sell_pants(pants_one)
salesperson.sell_pants(pants_two)
salesperson.sell_pants(pants_three)
salesperson.display_sales()
# -
# # Solution
#
# As a reminder, answers are also provided. If you click on the Jupyter icon, you can open a folder called 2.OOP_syntax_pants_practice, which contains this Jupyter notebook and a file called answer.py.
| exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# # Styles
#
# This shows the ``StyleTool`` and ``StyleTextTool`` to interactively change the style of GeoJSON and GeoData and similar layers.
# +
import requests
from ipyleaflet import basemaps, GeoJSON, LayersControl, Map
from ipywidgets import Layout
from leafmaptools.utils import bounds
from leafmaptools.styles import StyleTool, StyleTextTool
# + tags=[]
cc = "BRA"
url = f"https://raw.githubusercontent.com/johan/world.geo.json/master/countries/{cc}.geo.json"
data = requests.get(url).json()
gj = GeoJSON(data=data, name=cc)
# + tags=[]
bm = basemaps.OpenStreetMap.Mapnik
layout = Layout(width="800px", height="400px")
m = Map(center=[0, 0], zoom=3, basemap=bm, layout=layout)
m += LayersControl()
m += gj
m.fit_bounds(bounds(data))
st_stroke = StyleTool(a_map=m, layer=gj, transparent=True, position="bottomright")
st_fill = StyleTool(a_map=m, layer=gj, kind="fill", transparent=True, position="bottomright")
m
# -
bm = basemaps.OpenStreetMap.Mapnik
layout = Layout(width="800px", height="400px")
m = Map(center=[0, 0], zoom=3, basemap=bm, layout=layout)
m += LayersControl()
style={
"color": "blue", "fillColor": "red", "weight": 2,
"opacity": 1, "fillOpacity": 0.5, "dashArray": "10,20,5"
}
gj = GeoJSON(data=data, name=cc, style=style)
m += gj
m.fit_bounds(bounds(data))
st_stroke = StyleTextTool(a_map=m, layer=gj, kind="stroke", position="topright")
m
| examples/notebooks/styles.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import errno
import json
import os
import cv2
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import scipy.misc
from scipy.ndimage import rotate
from scipy.stats import bernoulli
# %matplotlib inline
# -
# Some useful constants
DRIVING_LOG_FILE = os.getcwd() + os.sep + os.pardir + '/datasets/data/driving_log.csv'
IMG_PATH = os.getcwd() + os.sep + os.pardir
IMG_PATH
def crop(image, top_percent, bottom_percent):
"""
Crops an image according to the given parameters
:param image: source image
:param top_percent:
The percentage of the original image will be cropped from the top of the image
:param bottom_percent:
The percentage of the original image will be cropped from the bottom of the image
:return:
The cropped image
"""
assert 0 <= top_percent < 0.5, 'top_percent should be between 0.0 and 0.5'
assert 0 <= bottom_percent < 0.5, 'top_percent should be between 0.0 and 0.5'
top = int(np.ceil(image.shape[0] * top_percent))
bottom = image.shape[0] - int(np.ceil(image.shape[0] * bottom_percent))
return image[top:bottom, :]
img = cv2.imread(IMG_PATH + '/datasets/data/IMG/100' + '.jpg')
# img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
plt.imshow(img)
plt.show()
cropped_img = crop(img, 0.1, 0.2)
plt.imshow(cropped_img)
plt.show()
def random_flip(image, steering_angle, flipping_prob=0.5):
"""
Based on the outcome of an coin flip, the image will be flipped.
If flipping is applied, the steering angle will be negated.
:param image: Source image
:param steering_angle: Original steering angle
:return: Both flipped image and new steering angle
"""
head = bernoulli.rvs(flipping_prob)
if head:
return np.fliplr(image), -1 * steering_angle
else:
return image, steering_angle
flipped_img, s = random_flip(cropped_img, 0.1, 1)
plt.imshow(flipped_img)
plt.show()
print(s)
def random_gamma(image):
"""
Random gamma correction is used as an alternative method changing the brightness of
training images.
http://www.pyimagesearch.com/2015/10/05/opencv-gamma-correction/
:param image:
Source image
:return:
New image generated by applying gamma correction to the source image
"""
gamma = np.random.uniform(0.4, 1.5)
inv_gamma = 1.0 / gamma
table = np.array([((i / 255.0) ** inv_gamma) * 255
for i in np.arange(0, 256)]).astype("uint8")
# apply gamma correction using the lookup table
return cv2.LUT(image, table)
gamma_img = random_gamma(flipped_img)
plt.imshow(gamma_img)
plt.show()
def random_shear(image, steering_angle, shear_range=50):
"""
Source: https://medium.com/@ksakmann/behavioral-cloning-make-a-car-drive-like-yourself-dc6021152713#.7k8vfppvk
:param image:
Source image on which the shear operation will be applied
:param steering_angle:
The steering angle of the image
:param shear_range:
Random shear between [-shear_range, shear_range + 1] will be applied
:return:
The image generated by applying random shear on the source image
"""
rows, cols, ch = image.shape
dx = np.random.randint(-shear_range, shear_range + 1)
random_point = [cols / 2 + dx, rows / 2]
pts1 = np.float32([[0, rows], [cols, rows], [cols / 2, rows / 2]])
pts2 = np.float32([[0, rows], [cols, rows], random_point])
dsteering = dx / (rows / 2) * 360 / (2 * np.pi * 25.0) / 6.0
M = cv2.getAffineTransform(pts1, pts2)
image = cv2.warpAffine(image, M, (cols, rows), borderMode=1)
steering_angle += dsteering
return image, steering_angle
sheared_img, s = random_shear(gamma_img, -0.3)
plt.imshow(sheared_img)
print(s)
plt.show()
resize_dim=(64, 64)
resized_img = scipy.misc.imresize(sheared_img, resize_dim)
plt.imshow(resized_img)
plt.show()
# +
import pandas as pd
data_frame = pd.read_csv(DRIVING_LOG_FILE, usecols=[0,1,2])
data_frame.describe(include='all')
# -
data_frame.hist(column='steering')
# ### Observations
# Data heavily biased towards left. Now I'd flip through all the images with a left steering angle, flip it and
# save it to the same folder and also add the respective steering angles to the csv file.
import csv
counter = 8309
with open(DRIVING_LOG_FILE, newline='') as f:
reader = csv.reader(f)
firstLine = True
for row in reader:
if firstLine:
firstLine = False
continue
steering = row[-1]
if(float(steering) < 0.0 and float(steering) > -0.2) and counter < 12400:
img = cv2.imread(IMG_PATH + '/datasets/' + row[0])
flipped_img, flipped_steering = random_flip(img, float(steering), 1)
save_path = IMG_PATH + '/datasets/data/IMG/' + str(counter) + '.jpg'
cv2.imwrite(save_path, flipped_img)
cv2.destroyAllWindows()
with open(DRIVING_LOG_FILE, "a") as fp:
wr = csv.writer(fp, dialect='excel')
row = ["data/IMG/" + str(counter) + ".jpg", float(0), flipped_steering]
wr.writerow(row)
counter += 1
print(save_path)
data_frame = pd.read_csv(DRIVING_LOG_FILE, usecols=[0,1,2])
data_frame.describe(include='all')
data_frame.hist(column='steering')
# Dataset augmentation with random shearing since the trained model still predicts too high steering angle values
import csv
counter = 5308
with open('data/driving_log.csv', newline='') as f:
reader = csv.reader(f)
firstRow = True
for row in reader:
if firstRow:
firstRow = False
continue
steering = row[-1]
if((float(steering) > -0.05 and float(steering) < 0.05 and float(steering)!=0)):
img = cv2.imread(row[0])
sheared_img, sheared_steering = random_shear(img, float(steering))
save_path = IMG_PATH + str(counter) + '.jpg'
cv2.imwrite(save_path, sheared_img)
cv2.destroyAllWindows()
with open('data/augmented.csv', "a") as fp:
wr = csv.writer(fp, dialect='excel')
row = ["data/IMG/" + str(counter) + ".jpg", float(0), str(sheared_steering)]
wr.writerow(row)
counter += 1
data_frame = pd.read_csv('data/driving_log.csv', usecols=[0,1,2])
data_frame.describe(include='all')
data_frame.hist(column='steering')
| iPythonNb/Preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import time
from matplotlib import pyplot as plt
from functools import lru_cache
# recursive
def init_fib(n):
assert n >= 0
return n if n <= 1 else init_fib(n - 1) + init_fib(n - 2)
res1a = init_fib(10)
# cache decorator
def memo(f):
cache = {}
def memo_fib(n):
if n not in cache:
cache[n] = f(n)
return cache[n]
return memo_fib
def fib1(n):
assert n >= 0
return n if n <= 1 else fib1(n - 1) + fib1(n - 2)
# recursion + cache
fib1 = memo(fib1)
res2a, res2b = fib1(10), fib1(80)
# recursion + python cache
@lru_cache(maxsize=None)
def lru_fib(n):
assert n >= 0
return n if n <= 1 else lru_fib(n - 1) + lru_fib(n - 2)
res3a, res3b = lru_fib(10), lru_fib(80)
# iterative
def iter_fib(n):
assert n >= 0
f0, f1 = 0, 1
for _ in range(n - 1):
f0, f1 = f1, f0 + f1
return f1
res4a, res4b = iter_fib(10), iter_fib(80)
#tests
assert res1a == res2a == res3a == res4a
assert res2b == res3b == res4b
def timed(f, *args, n_iter=100):
acc = float('inf')
for _ in range(n_iter):
t0 = time.perf_counter()
f(*args)
t1 = time.perf_counter()
acc = min(acc, t1 - t0)
return acc
print(f'Time for recursive fib is %.10f sec' % timed(init_fib, 22))
print(f'Time for cached fib is %.10f sec' % timed(lru_fib, 22))
print(f'Time for iterative fib is %.10f sec' % timed(iter_fib, 22))
# +
def compare(fs, args):
for f in fs:
plt.plot(args, [timed(f, arg) for arg in args], label=f.__name__)
plt.legend()
plt.grid()
compare([init_fib, lru_fib, iter_fib], list(range(22)))
# -
compare([fib1, lru_fib, iter_fib], list(range(500)))
compare([lru_fib, fib1], list(range(10**4)))
| algorithms/warmup/.ipynb_checkpoints/fibonacci-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Dependencies
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="mDT8S9C9CYtr"
# Tensorflow
import tensorflow as tf
print('Tested with TensorFLow 1.2.0')
print('Your TensorFlow version:', tf.__version__)
# Feeding function for enqueue data
from tensorflow.python.estimator.inputs.queues import feeding_functions as ff
# Rnn common functions
from tensorflow.contrib.learn.python.learn.estimators import rnn_common
# Model builder
from tensorflow.python.estimator import model_fn as model_fn_lib
# Run an experiment
from tensorflow.contrib.learn.python.learn import learn_runner
# Helpers for data processing
import pandas as pd
import numpy as np
import argparse
import random
# -
# ## Loading Data
#
# First, we want to create our word vectors. For simplicity, we're going to be using a pretrained model.
#
# As one of the biggest players in the ML game, Google was able to train a Word2Vec model on a massive Google News dataset that contained over 100 billion different words! From that model, Google [was able to create 3 million word vectors](https://code.google.com/archive/p/word2vec/#Pre-trained_word_and_phrase_vectors), each with a dimensionality of 300.
#
# In an ideal scenario, we'd use those vectors, but since the word vectors matrix is quite large (3.6 GB!), we'll be using a much more manageable matrix that is trained using [GloVe](http://nlp.stanford.edu/projects/glove/), a similar word vector generation model. The matrix will contain 400,000 word vectors, each with a dimensionality of 50.
#
# We're going to be importing two different data structures, one will be a Python list with the 400,000 words, and one will be a 400,000 x 50 dimensional embedding matrix that holds all of the word vector values.
# +
# data from: http://ai.stanford.edu/~amaas/data/sentiment/
TRAIN_INPUT = 'data/train.csv'
TEST_INPUT = 'data/test.csv'
# data manually generated
MY_TEST_INPUT = 'data/mytest.csv'
# wordtovec
# https://nlp.stanford.edu/projects/glove/
# the matrix will contain 400,000 word vectors, each with a dimensionality of 50.
word_list = np.load('word_list.npy')
word_list = word_list.tolist() # originally loaded as numpy array
word_list = [word.decode('UTF-8') for word in word_list] # encode words as UTF-8
print('Loaded the word list, length:', len(word_list))
word_vector = np.load('word_vector.npy')
print ('Loaded the word vector, shape:', word_vector.shape)
# -
# We can also search our word list for a word like "baseball", and then access its corresponding vector through the embedding matrix.
baseball_index = word_list.index('baseball')
print('Example: baseball')
print(word_vector[baseball_index])
# Now that we have our vectors, our first step is taking an input sentence and then constructing the its vector representation. Let's say that we have the input sentence "I thought the movie was incredible and inspiring". In order to get the word vectors, we can use Tensorflow's embedding lookup function. This function takes in two arguments, one for the embedding matrix (the wordVectors matrix in our case), and one for the ids of each of the words. The ids vector can be thought of as the integerized representation of the training set. This is basically just the row index of each of the words. Let's look at a quick example to make this concrete.
# +
max_seq_length = 10 # maximum length of sentence
num_dims = 300 # dimensions for each word vector
first_sentence = np.zeros((max_seq_length), dtype='int32')
first_sentence[0] = word_list.index("i")
first_sentence[1] = word_list.index("thought")
first_sentence[2] = word_list.index("the")
first_sentence[3] = word_list.index("movie")
first_sentence[4] = word_list.index("was")
first_sentence[5] = word_list.index("incredible")
first_sentence[6] = word_list.index("and")
first_sentence[7] = word_list.index("inspiring")
# first_sentence[8] = 0
# first_sentence[9] = 0
print(first_sentence.shape)
print(first_sentence) # shows the row index for each word
# -
# ###TODO### Insert image
# The 10 x 50 output should contain the 50 dimensional word vectors for each of the 10 words in the sequence.
with tf.Session() as sess:
print(tf.nn.embedding_lookup(word_vector, first_sentence).eval().shape)
# Before creating the ids matrix for the whole training set, let’s first take some time to visualize the type of data that we have. This will help us determine the best value for setting our maximum sequence length. In the previous example, we used a max length of 10, but this value is largely dependent on the inputs you have.
# The training set we're going to use is the Imdb movie review dataset. This set has 25,000 movie reviews, with 12,500 positive reviews and 12,500 negative reviews. Each of the reviews is stored in a txt file that we need to parse through. The positive reviews are stored in one directory and the negative reviews are stored in another. The following piece of code will determine total and average number of words in each review.
# +
from os import listdir
from os.path import isfile, join
positiveFiles = ['positiveReviews/' + f for f in listdir('positiveReviews/') if isfile(join('positiveReviews/', f))]
negativeFiles = ['negativeReviews/' + f for f in listdir('negativeReviews/') if isfile(join('negativeReviews/', f))]
numWords = []
for pf in positiveFiles:
with open(pf, "r", encoding='utf-8') as f:
line=f.readline()
counter = len(line.split())
numWords.append(counter)
print('Positive files finished')
for nf in negativeFiles:
with open(nf, "r", encoding='utf-8') as f:
line=f.readline()
counter = len(line.split())
numWords.append(counter)
print('Negative files finished')
numFiles = len(numWords)
print('The total number of files is', numFiles)
print('The total number of words in the files is', sum(numWords))
print('The average number of words in the files is', sum(numWords)/len(numWords))
# -
# We can also use the Matplot library to visualize this data in a histogram format.
import matplotlib.pyplot as plt
# %matplotlib inline
plt.hist(numWords, 50)
plt.xlabel('Sequence Length')
plt.ylabel('Frequency')
plt.axis([0, 1200, 0, 8000])
plt.show()
# From the histogram as well as the average number of words per file, we can safely say that most reviews will fall under 250 words, which is the max sequence length value we will set.
max_seq_len = 250
# ### Data
ids_matrix = np.load('ids_matrix.npy').tolist()
# ## Parameters
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="UrAyWt23AtCM"
# Parameters for training
STEPS = 100000
BATCH_SIZE = 32
# Parameters for data processing
REVIEW_KEY = 'review'
SEQUENCE_LENGTH_KEY = 'sequence_length'
# -
# ## Separating train and test data
#
# The training set we're going to use is the Imdb movie review dataset. This set has 25,000 movie reviews, with 12,500 positive reviews and 12,500 negative reviews.
#
# Let's first give a positive label [1, 0] to the first 12500 reviews, and a negative label [0, 1] to the other reviews.
# +
POSITIVE_REVIEWS = 12500
# copying sequences
data_sequences = [np.asarray(v, dtype=np.int32) for v in ids_matrix]
# generating labels
data_labels = [[1, 0] if i < POSITIVE_REVIEWS else [0, 1] for i in range(len(ids_matrix))]
# also creating a length column, this will be used by the Dynamic RNN
# see more about it here: https://www.tensorflow.org/api_docs/python/tf/nn/dynamic_rnn
data_length = [max_seq_len for i in range(len(ids_matrix))]
# -
# Then, let's shuffle the data and use 90% of the reviews for training and the other 10% for testing.
# +
data = list(zip(data_sequences, data_labels))
random.shuffle(data) # shuffle
data = np.asarray(data)
print(data.shape)
# separating train and test data
limit = int(len(data) * 0.9)
train_data = data[:limit]
test_data = data[limit:]
# -
# ### Verifying if the train and test data have enough positive and negative examples
# +
LABEL_INDEX = 1
def _number_of_pos_labels(df):
pos_labels = 0
for value in df:
if value[LABEL_INDEX] == [1, 0]:
pos_labels += 1
return pos_labels
pos_labels_train = _number_of_pos_labels(train_data)
total_labels_train = len(train_data)
pos_labels_test = _number_of_pos_labels(test_data)
total_labels_test = len(test_data)
print('Total number of positive labels:', pos_labels_train + pos_labels_test)
print('Proportion of positive labels on the Train data:', pos_labels_train/total_labels_train)
print('Proportion of positive labels on the Test data:', pos_labels_test/total_labels_test)
# -
# ## Input functions
def get_input_fn(df, batch_size, num_epochs=1, shuffle=True):
def input_fn():
# https://github.com/tensorflow/tensorflow/tree/master/tensorflow/contrib/data
sequences = np.asarray([v for v in df[:,0]], dtype=np.int32)
labels = np.asarray([v for v in df[:,1]], dtype=np.int32)
length = np.asarray(df[:,2], dtype=np.int32)
#
dataset = (
tf.contrib.data.Dataset.from_tensor_slices((sequences, labels, length)) # reading data from memory
.repeat(num_epochs) # repeat dataset the number of epochs
.batch(batch_size)
)
# for our "manual" test we don't want to shuffle the data
if shuffle:
dataset = dataset.shuffle(buffer_size=100000)
# create iterator
review, label, length = dataset.make_one_shot_iterator().get_next()
features = {
REVIEW_KEY: review,
SEQUENCE_LENGTH_KEY: length,
}
return features, label
return input_fn
# +
features, label = get_input_fn(train_data, 2)()
with tf.Session() as sess:
items = sess.run(features)
print(items[REVIEW_KEY])
print
items = sess.run(features)
print(items[REVIEW_KEY])
print
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="m5UJyvW5P0Sy"
train_input_fn = get_input_fn(train_data, BATCH_SIZE, None)
test_input_fn = get_input_fn(test_data, BATCH_SIZE)
# -
# ## Creating the Estimator model
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="VxXAUrYN7TvR"
def get_model_fn(rnn_cell_sizes,
label_dimension,
dnn_layer_sizes=[],
optimizer='SGD',
learning_rate=0.01,
embed_dim=128):
def model_fn(features, labels, mode):
review = features[REVIEW_KEY]
# Creating dense representation for the sentences
# and then converting it to embeding representation
data = tf.nn.embedding_lookup(word_vector, review)
# Each RNN layer will consist of a LSTM cell
#rnn_layers = [tf.nn.rnn_cell.LSTMCell(size) for size in rnn_cell_sizes]
lstm_cell = tf.nn.rnn_cell.LSTMCell(64)
lstm_cell = tf.nn.rnn_cell.DropoutWrapper(cell=lstm_cell, output_keep_prob=0.25)
# Construct the layers
#multi_rnn_cell = tf.nn.rnn_cell.MultiRNNCell(rnn_layers)
# Runs the RNN model dynamically
# more about it at:
# https://www.tensorflow.org/api_docs/python/tf/nn/dynamic_rnn
outputs, final_state = tf.nn.dynamic_rnn(cell=lstm_cell,
inputs=data,
dtype=tf.float32)
# Slice to keep only the last cell of the RNN
last_activations = rnn_common.select_last_activations(outputs,
sequence_length)
# Final dense layer for prediction
predictions = tf.layers.dense(last_activations, label_dimension)
predictions_softmax = tf.nn.softmax(predictions)
loss = None
train_op = None
preds_op = {
'prediction': predictions_softmax,
'label': labels
}
eval_op = {
"accuracy": tf.metrics.accuracy(
tf.argmax(input=predictions_softmax, axis=1),
tf.argmax(input=labels, axis=1))
}
if mode != tf.estimator.ModeKeys.PREDICT:
loss = tf.losses.softmax_cross_entropy(labels, predictions)
if mode == tf.estimator.ModeKeys.TRAIN:
train_op = tf.contrib.layers.optimize_loss(
loss,
tf.contrib.framework.get_global_step(),
optimizer=optimizer,
learning_rate=learning_rate)
return tf.contrib.learn.ModelFnOps(mode,
predictions=predictions_softmax,
loss=loss,
train_op=train_op,
eval_metric_ops=eval_op)
return model_fn
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="gUHR3Mzc7Tvb"
model_fn = get_model_fn(rnn_cell_sizes=[64], # size of the hidden layers
label_dimension=2, # since are just 2 classes
dnn_layer_sizes=[128, 64], # size of units in the dense layers on top of the RNN
optimizer='Adam',
learning_rate=0.001,
embed_dim=512)
estimator = tf.contrib.learn.Estimator(model_fn=model_fn, model_dir='tensorboard18/')
# -
# ## Create and Run Experiment
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="DUZEKQrdGgZE"
# create experiment
def generate_experiment_fn():
"""
Create an experiment function given hyperparameters.
Returns:
A function (output_dir) -> Experiment where output_dir is a string
representing the location of summaries, checkpoints, and exports.
this function is used by learn_runner to create an Experiment which
executes model code provided in the form of an Estimator and
input functions.
All listed arguments in the outer function are used to create an
Estimator, and input functions (training, evaluation, serving).
Unlisted args are passed through to Experiment.
"""
def _experiment_fn(output_dir):
return tf.contrib.learn.Experiment(
estimator,
train_input_fn=train_input_fn,
eval_input_fn=test_input_fn,
train_steps=STEPS
)
return _experiment_fn
# -
# run experiment
learn_runner.run(generate_experiment_fn(), '/tmp/outputdir')
# ## Making Predictions
# +
preds = estimator.predict(input_fn=my_test_input_fn, as_iterable=True)
sentences = _get_csv_column(MY_TEST_INPUT, 'review')
print()
for p, s in zip(preds, sentences):
print('sentence:', s)
print('bad review:', p[0], 'good review:', p[1])
print('-' * 10)
# -
| code_samples/RNN/sentiment_analysis/.ipynb_checkpoints/SentimentAnalysis-Copy1-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Jupyter Notebook to compute performance metrics from the segmentation predictions.
# ## Imports
# +
from pathlib import Path
import os
import json
from shutil import copy
# Scientific modules import
import numpy as np
from skimage import io
import matplotlib.pyplot as plt
# AxonDeepSeg import modules
from AxonDeepSeg.apply_model import axon_segmentation
from AxonDeepSeg.testing.segmentation_scoring import Metrics_calculator
import AxonDeepSeg.ads_utils as ads
from config import axonmyelin_suffix
# %matplotlib inline
# -
# ## Use case: compute axon and myelin validation metrics on a sample image
ads_path = Path(os.path.abspath('')).resolve().parent
path_img = Path(os.path.join(ads_path,'AxonDeepSeg','models','model_seg_rat_axon-myelin_sem','data_test')) / 'image.png'
# +
# Set paths
img = ads.imread(path_img)
path_folder, file_name = os.path.split(path_img)
model_name = 'model_seg_rat_axon-myelin_sem'
path_model = Path(os.path.join(ads_path,'AxonDeepSeg','models',model_name))
# Groundtruth image
mask = ads.imread(os.path.join(path_folder,'mask.png'))
gt_axon = mask > 200
gt_myelin = np.logical_and(mask >= 50, mask <= 200)
# Prediction image (output of AxonDeepSeg)
pred = ads.imread(os.path.join(path_folder,'image' + str(axonmyelin_suffix)))
pred_axon = pred > 200
pred_myelin = np.logical_and(pred >= 50, pred <= 200)
# -
fig, axes = plt.subplots(1,2, figsize=(13,10))
ax1, ax2 = axes[0], axes[1]
ax1.set_title('Prediction')
ax1.imshow(pred,cmap='gray')
ax2.set_title('Groundtruth')
ax2.imshow(mask,cmap='gray')
plt.show();
# ## Compute validation metrics for axon segmentation
# +
# Create object to manage segmentation metrics for axon mask
axon_metrics = Metrics_calculator(pred_axon, gt_axon)
# Compute and display results (pixelwise metrics):
print('Pixelwise Sensitivity:', axon_metrics.pw_sensitivity())
print('Pixelwise Accuracy:',axon_metrics.pw_accuracy())
print('Pixelwise Dice:',axon_metrics.pw_dice())
print('Pixelwise F1 score:',axon_metrics.pw_F1_score())
print('Pixelwise False negative rate:',axon_metrics.pw_FN_rate())
print('Pixelwise False positive rate:',axon_metrics.pw_FP_rate())
print('Pixelwise Jaccard index:',axon_metrics.pw_jaccard())
print('Pixelwise Precision:',axon_metrics.pw_precision())
print('Pixelwise Specificity:',axon_metrics.pw_specificity())
axon_metrics_array = np.array([axon_metrics.pw_sensitivity(),axon_metrics.pw_specificity(),axon_metrics.pw_precision(),
axon_metrics.pw_accuracy(), axon_metrics.pw_F1_score(), axon_metrics.pw_dice(), axon_metrics.pw_jaccard()])
# Compute and display results (elementwise metrics):
dice_output = axon_metrics.ew_dice('all')
print('Elementwise Dice:',axon_metrics.ew_dice('short'))
# -
# ## Compute validation metrics for myelin segmentation
# +
# Create object to manage segmentation metrics for myelin mask
myelin_metrics = Metrics_calculator(pred_myelin, gt_myelin)
# Compute and display results (pixelwise metrics):
print('Pixelwise Sensitivity:', myelin_metrics.pw_sensitivity())
print('Pixelwise Accuracy:',myelin_metrics.pw_accuracy())
print('Pixelwise Dice:',myelin_metrics.pw_dice())
print('Pixelwise F1 score:',myelin_metrics.pw_F1_score())
print('Pixelwise False negative rate:',myelin_metrics.pw_FN_rate())
print('Pixelwise False positive rate:',myelin_metrics.pw_FP_rate())
print('Pixelwise Jaccard index:',myelin_metrics.pw_jaccard())
print('Pixelwise Precision:',myelin_metrics.pw_precision())
print('Pixelwise Specificity:',myelin_metrics.pw_specificity())
myelin_metrics_array = np.array([myelin_metrics.pw_sensitivity(),myelin_metrics.pw_specificity(),myelin_metrics.pw_precision(),
myelin_metrics.pw_accuracy(), myelin_metrics.pw_F1_score(), myelin_metrics.pw_dice(),
myelin_metrics.pw_jaccard()])
# -
# ## Visualization of pixelwise metrics for both axon and myelin
# +
n_groups = 7
fig, ax = plt.subplots()
index = np.arange(n_groups)
ax.bar(index-0.1, axon_metrics_array, width=0.2, color='green',label='axon')
ax.bar(index+0.1, myelin_metrics_array, width=0.2, color='orange',label='myelin')
ax.set_ylabel('Values')
ax.set_title('Pixelwise segmentation metrics for axon and myelin')
ax.set_xticks(index)
ax.set_xticklabels(('Sens.', 'Spec.', 'Prec.', 'Accur.', 'F1 score','Dice','Jaccard'), minor=False, rotation=0)
ax.legend(loc='best')
ax.grid(True)
ax.set_ylim(0.5, 1)
plt.tight_layout()
plt.show()
# -
# ## Visualization of elementwise Dice metric for axon objects
# +
fig, ax = plt.subplots()
ax.boxplot(dice_output, 0, '',vert=False)
ax.set_title('Elementwise Dice of axon objects')
ax.grid(True)
# A way to remove the y labels
ax.set_yticklabels([]);
| notebooks/01-performance_metrics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="2ZlEIEW_REmg" colab_type="text"
# ## Exercicios - Troca de faces por reposicionamento da face
# Neste exercicio iremos inverter os rostos de duas imagens por meio da detecção
# da posiçaõ da face.
#
#
# Iremos utilizar pare esses exercicios:
#
#
# 1) shape_predictor_68_face_landmarks.dat - Detecta os 68 principais pontos de referencia da face
#
#
# 2) ffmpeg - conversor video/imagem
#
# 3) python - codigo do programa
#
# ---
#
#
# 
#
# ---
#
#
# Com esse exercicio você será capaz de fazer 2 operações:
#
# 1) Inverter rosto em imagens;
#
# 2) Iverter rosto em videos;
# + [markdown] id="Fxd9XzjLVrkf" colab_type="text"
# # Instalar programas
# 1) install ffmpeg
# 2)
# + id="-aF1lRA6Vsok" colab_type="code" outputId="029a5572-655c-4041-a520-b5a047ce8741" colab={"base_uri": "https://localhost:8080/", "height": 102}
# !apt install ffmpeg
# + id="xxNfTag2RA1M" colab_type="code" outputId="410ba436-e4bf-4d21-e09b-87868aae06de" colab={"base_uri": "https://localhost:8080/", "height": 377}
# !wget https://github.com/AKSHAYUBHAT/TensorFace/blob/master/openface/models/dlib/shape_predictor_68_face_landmarks.dat?raw=true
# + [markdown] id="P4chfpztjihU" colab_type="text"
# # EXERCICIO 1 - Troca de rosto em imagem
#
# -Substitua os nomes "IMAGE1" e "IMAGE2" pelo nome das imagens que você deseja
# + id="GdfDOyPRRERb" colab_type="code" outputId="1a26afa1-1357-4c2b-ec25-9d8f6a6e333d" colab={"base_uri": "https://localhost:8080/", "height": 34}
import cv2
import dlib
import numpy
import sys
IMAGE1='./sa.jpeg'
IMAGE2='./b.jpg'
PREDICTOR_PATH = "shape_predictor_68_face_landmarks.dat?raw=true"
SCALE_FACTOR = 1
FEATHER_AMOUNT = 9
FACE_POINTS = list(range(17, 68))
MOUTH_POINTS = list(range(48, 61))
RIGHT_BROW_POINTS = list(range(17, 22))
LEFT_BROW_POINTS = list(range(22, 27))
RIGHT_EYE_POINTS = list(range(36, 42))
LEFT_EYE_POINTS = list(range(42, 48))
NOSE_POINTS = list(range(27, 35))
JAW_POINTS = list(range(0, 17))
# Pontos usados para alinhar as imagens.
ALIGN_POINTS = (LEFT_BROW_POINTS + RIGHT_EYE_POINTS + LEFT_EYE_POINTS +
RIGHT_BROW_POINTS + NOSE_POINTS + MOUTH_POINTS+JAW_POINTS)
# Aponta da segunda imagem para sobrepor na primeira. O casco convexo de cada O elemento
# será sobreposto.
OVERLAY_POINTS = [
LEFT_EYE_POINTS + RIGHT_EYE_POINTS + LEFT_BROW_POINTS + RIGHT_BROW_POINTS,
NOSE_POINTS + MOUTH_POINTS,JAW_POINTS
]
# Quantidade de desfoque a ser usada durante a correção de cores, como uma fração da
# distância pupilar.
COLOUR_CORRECT_BLUR_FRAC = .8
detector = dlib.get_frontal_face_detector()
predictor = dlib.shape_predictor(PREDICTOR_PATH)
class TooManyFaces(Exception):
pass
class NoFaces(Exception):
pass
def get_landmarks(im):
rects = detector(im, 1)
if len(rects) > 1:
raise TooManyFaces
if len(rects) == 0:
raise NoFaces
return numpy.matrix([[p.x, p.y] for p in predictor(im, rects[0]).parts()])
def annotate_landmarks(im, landmarks):
im = im.copy()
for idx, point in enumerate(landmarks):
pos = (point[0, 0], point[0, 1])
cv2.putText(im, str(idx), pos,
fontFace=cv2.FONT_HERSHEY_SCRIPT_SIMPLEX,
fontScale=0.4,
color=(0, 0, 255))
cv2.circle(im, pos, 3, color=(0, 255, 255))
return im
def draw_convex_hull(im, points, color):
points = cv2.convexHull(points)
cv2.fillConvexPoly(im, points, color=color)
def get_face_mask(im, landmarks):
im = numpy.zeros(im.shape[:2], dtype=numpy.float64)
for group in OVERLAY_POINTS:
draw_convex_hull(im,
landmarks[group],
color=1)
im = numpy.array([im, im, im]).transpose((1, 2, 0))
im = (cv2.GaussianBlur(im, (FEATHER_AMOUNT, FEATHER_AMOUNT), 0) > 0) * 1.0
im = cv2.GaussianBlur(im, (FEATHER_AMOUNT, FEATHER_AMOUNT), 0)
return im
def transformation_from_points(points1, points2):
"""
Return an affine transformation [s * R | T] such that:
sum ||s*R*p1,i + T - p2,i||^2
is minimized.
"""
# Resolva o problema subtraindo os centróides, escalando pelo
# desvio padrão e, em seguida, usando o SVD para calcular a rotação. Vejo
# para mais detalhes:
# https://en.wikipedia.org/wiki/Orthogonal_Procrustes_problem
points1 = points1.astype(numpy.float64)
points2 = points2.astype(numpy.float64)
c1 = numpy.mean(points1, axis=0)
c2 = numpy.mean(points2, axis=0)
points1 -= c1
points2 -= c2
s1 = numpy.std(points1)
s2 = numpy.std(points2)
points1 /= s1
points2 /= s2
U, S, Vt = numpy.linalg.svd(points1.T * points2)
# OR que buscamos é, de fato, a transposição da dada por U * Vt.
# é porque a formulação acima assume que a matriz segue à direita
# (com vetores de linha) onde, como nossa solução requer que a matriz esteja na
# esquerda (com vetores de coluna).
R = (U * Vt).T
return numpy.vstack([numpy.hstack(((s2 / s1) * R,
c2.T - (s2 / s1) * R * c1.T)),
numpy.matrix([0., 0., 1.])])
def read_im_and_landmarks(fname):
im = cv2.imread(fname, cv2.IMREAD_COLOR)
im = cv2.resize(im, (im.shape[1] * SCALE_FACTOR,
im.shape[0] * SCALE_FACTOR))
s = get_landmarks(im)
return im, s
def warp_im(im, M, dshape):
output_im = numpy.zeros(dshape, dtype=im.dtype)
cv2.warpAffine(im,
M[:2],
(dshape[1], dshape[0]),
dst=output_im,
borderMode=cv2.BORDER_TRANSPARENT,
flags=cv2.WARP_INVERSE_MAP)
return output_im
def correct_colours(im1, im2, landmarks1):
blur_amount = COLOUR_CORRECT_BLUR_FRAC * numpy.linalg.norm(
numpy.mean(landmarks1[LEFT_EYE_POINTS], axis=0) -
numpy.mean(landmarks1[RIGHT_EYE_POINTS], axis=0))
blur_amount = int(blur_amount)
if blur_amount % 2 == 0:
blur_amount += 1
im1_blur = cv2.GaussianBlur(im1, (blur_amount, blur_amount), 0)
im2_blur = cv2.GaussianBlur(im2, (blur_amount, blur_amount), 0)
# Evitar divizão por zero.
im2_blur += (128 * (im2_blur <= 1.0)).astype(im2_blur.dtype)
return (im2.astype(numpy.float64) * im1_blur.astype(numpy.float64) /
im2_blur.astype(numpy.float64))
im1, landmarks1 = read_im_and_landmarks(IMAGE1)
im2, landmarks2 = read_im_and_landmarks(IMAGE2)
M = transformation_from_points(landmarks1[ALIGN_POINTS],
landmarks2[ALIGN_POINTS])
mask = get_face_mask(im2, landmarks2)
warped_mask = warp_im(mask, M, im1.shape)
combined_mask = numpy.max([get_face_mask(im1, landmarks1), warped_mask],
axis=0)
warped_im2 = warp_im(im2, M, im1.shape)
warped_corrected_im2 = correct_colours(im1, warped_im2, landmarks1)
output_im = im1 * (1.0 - combined_mask) + warped_corrected_im2 * combined_mask
cv2.imwrite('./resultado.jpg', output_im)
# + [markdown] id="AVYdQ83Gl63O" colab_type="text"
#
# # EXERCICIO 2 - Troca de rosto em video
# Substitua os nomes "VIDEO1" e "VIDEO2" pelo nome dos videos que você deseja
# + id="HeFYftDkRDka" colab_type="code" outputId="297c5606-e79a-4731-81b7-20417c740a13" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# #!/usr/bin/python
import cv2
import dlib
import numpy
import os
import sys
VIDEO1='download.mp4'
VIDEO2='pica.mp4'
TEMPO=10
try:
os.system('mkdir corpo')
os.system('mkdir res')
os.system('mkdir face')
pass
except expression as identifier:
pass
os.system('ffmpeg -i %s -vf fps=25 -t %s ./corpo/%s.jpeg' %(VIDEO1,TEMPO,'%d'))
os.system('ffmpeg -i %s -vf fps=25 -t %s ./face/%s.jpeg' %(VIDEO2,TEMPO,'%d'))
PREDICTOR_PATH = "shape_predictor_68_face_landmarks.dat?raw=true"
SCALE_FACTOR = 1
FEATHER_AMOUNT = 11
FACE_POINTS = list(range(17, 68))
MOUTH_POINTS = list(range(48, 61))
RIGHT_BROW_POINTS = list(range(17, 22))
LEFT_BROW_POINTS = list(range(22, 27))
RIGHT_EYE_POINTS = list(range(36, 42))
LEFT_EYE_POINTS = list(range(42, 48))
NOSE_POINTS = list(range(27, 35))
JAW_POINTS = list(range(0, 17))
ALIGN_POINTS = (LEFT_BROW_POINTS + RIGHT_EYE_POINTS + LEFT_EYE_POINTS +
RIGHT_BROW_POINTS + NOSE_POINTS + MOUTH_POINTS)
OVERLAY_POINTS = [
LEFT_EYE_POINTS + RIGHT_EYE_POINTS + LEFT_BROW_POINTS + RIGHT_BROW_POINTS,
NOSE_POINTS + MOUTH_POINTS
]
COLOUR_CORRECT_BLUR_FRAC = .6
detector = dlib.get_frontal_face_detector()
predictor = dlib.shape_predictor(PREDICTOR_PATH)
class TooManyFaces(Exception):
pass
class NoFaces(Exception):
pass
def get_landmarks(im):
rects = detector(im, 1)
if len(rects) > 1:
raise TooManyFaces
if len(rects) == 0:
raise NoFaces
return numpy.matrix([[p.x, p.y] for p in predictor(im, rects[0]).parts()])
def annotate_landmarks(im, landmarks):
im = im.copy()
for idx, point in enumerate(landmarks):
pos = (point[0, 0], point[0, 1])
cv2.putText(im, str(idx), pos,
fontFace=cv2.FONT_HERSHEY_SCRIPT_SIMPLEX,
fontScale=0.4,
color=(0, 0, 255))
cv2.circle(im, pos, 3, color=(0, 255, 255))
return im
def draw_convex_hull(im, points, color):
points = cv2.convexHull(points)
cv2.fillConvexPoly(im, points, color=color)
def get_face_mask(im, landmarks):
im = numpy.zeros(im.shape[:2], dtype=numpy.float64)
for group in OVERLAY_POINTS:
draw_convex_hull(im,
landmarks[group],
color=1)
im = numpy.array([im, im, im]).transpose((1, 2, 0))
im = (cv2.GaussianBlur(im, (FEATHER_AMOUNT, FEATHER_AMOUNT), 0) > 0) * 1.0
im = cv2.GaussianBlur(im, (FEATHER_AMOUNT, FEATHER_AMOUNT), 0)
return im
def transformation_from_points(points1, points2):
"""
Return an affine transformation [s * R | T] such that:
sum ||s*R*p1,i + T - p2,i||^2
is minimized.
"""
points1 = points1.astype(numpy.float64)
points2 = points2.astype(numpy.float64)
c1 = numpy.mean(points1, axis=0)
c2 = numpy.mean(points2, axis=0)
points1 -= c1
points2 -= c2
s1 = numpy.std(points1)
s2 = numpy.std(points2)
points1 /= s1
points2 /= s2
U, S, Vt = numpy.linalg.svd(points1.T * points2)
R = (U * Vt).T
return numpy.vstack([numpy.hstack(((s2 / s1) * R,
c2.T - (s2 / s1) * R * c1.T)),
numpy.matrix([0., 0., 1.])])
def read_im_and_landmarks(fname):
im = cv2.imread(fname, cv2.IMREAD_COLOR)
im = cv2.resize(im, (im.shape[1] * SCALE_FACTOR,
im.shape[0] * SCALE_FACTOR))
s = get_landmarks(im)
return im, s
def warp_im(im, M, dshape):
output_im = numpy.zeros(dshape, dtype=im.dtype)
cv2.warpAffine(im,
M[:2],
(dshape[1], dshape[0]),
dst=output_im,
borderMode=cv2.BORDER_TRANSPARENT,
flags=cv2.WARP_INVERSE_MAP)
return output_im
def correct_colours(im1, im2, landmarks1):
blur_amount = COLOUR_CORRECT_BLUR_FRAC * numpy.linalg.norm(
numpy.mean(landmarks1[LEFT_EYE_POINTS], axis=0) -
numpy.mean(landmarks1[RIGHT_EYE_POINTS], axis=0))
blur_amount = int(blur_amount)
if blur_amount % 2 == 0:
blur_amount += 1
im1_blur = cv2.GaussianBlur(im1, (blur_amount, blur_amount), 0)
im2_blur = cv2.GaussianBlur(im2, (blur_amount, blur_amount), 0)
# Avoid divide-by-zero errors.
im2_blur += (128 * (im2_blur <= 1.0)).astype(im2_blur.dtype)
return (im2.astype(numpy.float64) * im1_blur.astype(numpy.float64) /
im2_blur.astype(numpy.float64))
def run1(f1,f2,f3):
im1, landmarks1 = read_im_and_landmarks(f1)
im2, landmarks2 = read_im_and_landmarks(f2)
M = transformation_from_points(landmarks1[ALIGN_POINTS],
landmarks2[ALIGN_POINTS])
mask = get_face_mask(im2, landmarks2)
warped_mask = warp_im(mask, M, im1.shape)
combined_mask = numpy.max([get_face_mask(im1, landmarks1), warped_mask],
axis=0)
warped_im2 = warp_im(im2, M, im1.shape)
warped_corrected_im2 = correct_colours(im1, warped_im2, landmarks1)
output_im = im1 * (1.0 - combined_mask) + warped_corrected_im2 * combined_mask
cv2.imwrite(f3, output_im)
a=range(25*TEMPO)
worked=1
for x in a:
print(x)
try:
run1('./corpo/%s.jpeg' % x,'./face/%s.jpeg' % x,'./res/%s.jpeg' % x )
worked=x
except:
run1('./corpo/%s.jpeg' % worked,'./face/%s.jpeg' % worked,'./res/%s.jpeg' % x )
pass
os.system('ffmpeg -y -r 25 -i res/%s.jpeg -i %s -acodec copy -c:v copy -c:a aac -map 0:v:0 -map 1:a:0 -t %s out.mp4' %('%d',VIDEO2,TEMPO))
| coreswap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # [Fast RG - quickly sampling a sparse matrix with low rank expectation](https://arxiv.org/pdf/1703.02998.pdf)
#
# R Code: https://github.com/RoheLab/fastRG/
# Python original implementation: https://github.com/yunjhongwu/matrix-routines/blob/master/fastRG.py
#
# Key algorithm:
#
#
# ```
# Require: X ∈ Rn×Kx, S ∈ RKx×Ky, and Y ∈ Rd×Ky with all matrices containing non-negative entries.
# Compute diagonal matrix CX ∈ RKx×Kx with CX = diag(∑i Xi1 , ... , ∑i XiKx).
# Compute diagonal matrix CY ∈ RKy×Ky with CY = diag(∑i Yi1 , ... , ∑i YiKy).
# Define X˜ = XC−1X, S˜ = CX SCY , and Y˜ = YC−1Y.
# Sample the number of edges m ∼ Poisson(∑u,v S˜uv).
# for ` = 1 : m do
# Sample U ∈ {1, ..., Kx},V ∈ {1, ..., Ky} with P(U = u,V = v) ∝ S˜uv.
# Sample I ∈ {1, ..., n} with P(I = i) = X˜iU .
# Sample J ∈ {1, ..., d} with P(J = j) = Y˜jV .
# Add edge (I, J) to the graph, allowing for multiple edges (I, J).
# end for
# ```
#
# +
import numpy as np
import scipy.sparse as scs
import matplotlib.pyplot as plt
import seaborn as sns
# -
N, D = 10, 7
K1, K2 = 5, 4
# +
def create_sparse_matrix(N,K):
cols = np.random.randint(K, size=(N,)) # membership
rows = np.arange(N, dtype=int)
data = np.ones_like(rows)
X = scs.csc_matrix((data, (rows, cols)), shape=(N,K))
return X
def plot_matrix(X, name="X", ax=None):
if ax is None:
fig, ax = plt.subplots(1,1)
if not isinstance(X, np.ndarray):
X = X.toarray()
sns.heatmap(
X, cmap="Reds",
vmin=0, vmax=1,
linewidths=.5,
square=True, cbar=False,
ax=ax
)
ax.set_title(name)
# -
X = create_sparse_matrix(N,K1)
Y = create_sparse_matrix(D,K2)
S = create_sparse_matrix(K1,K2)
X, Y, S
A = X.dot(S).dot(Y.T)
A
fig, ax = plt.subplots(1,4)
plot_matrix(X, name="X", ax=ax[0])
plot_matrix(S, name="S", ax=ax[1])
plot_matrix(Y, name="Y", ax=ax[2])
plot_matrix(A, name="A", ax=ax[3])
fig.tight_layout()
# +
class FastRG(object):
def __init__(self, X, Y, S):
self.X_prime, self.Y_prime, self.S_prime = FastRG.compute_primes(X, Y, S)
@classmethod
def compute_C(cls, X):
CX = np.asarray(X.sum(axis=0)).sum(axis=0)#.ravel()
return CX
@classmethod
def compute_primes(cls, X, Y, S):
CX = FastRG.compute_C(X)
CY = FastRG.compute_C(Y)
X_prime = X / CX
Y_prime = Y / CY
S_prime = CX[:, np.newaxis] * S.toarray() * CY # CX.dot(S.toarray()).dot(CY)
return X_prime, Y_prime, S_prime
@classmethod
def sample_block_edges(cls, S_prime, m):
UV_counts = np.random.multinomial(m, S_prime.ravel()/S_prime.sum()).reshape(S_prime.shape)
UV_counts_indices = np.indices(UV_counts.shape) # row, col
U, V = UV_counts_indices[:, UV_counts > 0]
UV_counts_nz = UV_counts[U, V]
return U, V, UV_counts_nz
@classmethod
def sample_edges(cls, X_prime, Y_prime, U, V, UV_counts_nz, m):
sampled_edges = set()
sampled_edges_ids = dict()
for i,(u,v,c) in enumerate(zip(U, V, UV_counts_nz)):
print(f"Sampling using u={u}, v={v}, c={c}")
uv_edges = set()
remaining_edges = c - len(uv_edges)
while remaining_edges > 0:
i_samples = np.random.choice(np.arange(N), p=np.asarray(X_prime[:, u]).ravel(), size=c)
j_samples = np.random.choice(np.arange(D), p=np.asarray(Y_prime[:, v]).ravel(), size=c)
# print(i_samples, j_samples)
new_edges = set(zip(i_samples, j_samples)) - uv_edges
if len(new_edges) >= remaining_edges:
new_edges = list(new_edges)[:remaining_edges]
uv_edges.update(new_edges)
uv_edges = uv_edges - sampled_edges
remaining_edges = c - len(uv_edges)
break
# print(f"\tremaining_edges={remaining_edges}: {new_edges}")
sampled_edges_ids.update({e: i for e in uv_edges - sampled_edges})
sampled_edges.update(uv_edges)
print(f"Sampled {len(sampled_edges)} edges, needed: {m}")
sampled_rows, sampled_cols = map(np.array, zip(*list(sampled_edges)))
data = [sampled_edges_ids[(u,v)] for u,v in zip(sampled_rows, sampled_cols)]
# data = np.ones_like(sampled_rows)
return data, sampled_rows, sampled_cols
def sample(self):
X_prime, Y_prime, S_prime = self.X_prime, self.Y_prime, self.S_prime
N, K1 = X_prime.shape
D, K2 = Y_prime.shape
m = np.random.poisson(S_prime.sum())
U, V, UV_counts_nz = FastRG.sample_block_edges(S_prime, m)
data, sampled_rows, sampled_cols = FastRG.sample_edges(X_prime, Y_prime, U, V, UV_counts_nz, m)
# UV_counts = np.random.multinomial(m, S_prime.ravel()/S_prime.sum()).reshape(K1,K2)
# UV_counts_indices = np.indices(UV_counts.shape) # row, col
# U, V = UV_counts_indices[:, UV_counts > 0]
# UV_counts_nz = UV_counts[U, V]
sample_G = scs.csc_matrix((data, (sampled_rows, sampled_cols)), shape=(N,D))
return sample_G
# -
sampler = FastRG(X, Y, S)
sampled_A = sampler.sample()
fig, ax = plt.subplots(1,4)
plot_matrix(sampler.X_prime, name="X", ax=ax[0])
plot_matrix(sampler.Y_prime, name="Y", ax=ax[2])
plot_matrix(sampler.S_prime, name="S", ax=ax[1])
plot_matrix(sampled_A, name="A", ax=ax[3])
fig.tight_layout()
# +
from scipy.sparse import csc_matrix
def howManyEdges(X, S, Y=None):
if Y is None:
Y = X
Cx = np.sum(X, axis=0)
Cy = np.sum(Y, axis=0)
em = Cx @ np.sum(S * Cy, axis=1)
avDeg = em / X.shape[0]
return em, avDeg
def fastRG(X, S, Y=None, avgDeg=None, simple=None, PoissonEdges=True,
directed=False, selfLoops=False, returnEdgeList=False,
returnParameters=False):
"""
<NAME>., <NAME>., <NAME>., & <NAME>. (2017). A note on quickly
sampling a sparse matrix with low rank expectation. arXiv preprint
arXiv:1703.02998.
Implementation of fastRG in R
https://github.com/karlrohe/fastRG
"""
if Y is not None and Y.size > 0:
directed = True
selfLoops = True
simple = False
returnY = True
else:
Y = X
returnY = False
if np.any(X < 0) or np.any(S < 0) or np.any(Y < 0):
return None
if simple is not None and simple:
selfLoops = False
directed = False
PoissonEdges = False
n, K1 = X.shape
d, K2 = Y.shape
if avgDeg is not None:
_, eDbar = howManyEdges(X, S, Y)
S *= avgDeg / eDbar
if not directed:
S = (S + S.T) * 0.25
Cx = np.sum(X, axis=0)
Cy = np.sum(Y, axis=0)
Xt = (X * (1 / Cx)).T
Yt = (Y * (1 / Cy)).T
St = Cx[:, None] * S * Cy
m = np.random.poisson(np.sum(St))
if m == 0:
A = csc_matrix((n, d))
return (A, X, S, Y if returnY else None) if returnParameters else A
tabUV = np.random.multinomial(m, pvals=St.ravel() * (1 / np.sum(St))).reshape((K1, K2))
elist = np.empty((2, m))
eitmp = np.empty(m)
blockDegreesU = np.sum(tabUV, axis=1)
tickerU = np.insert(np.cumsum(blockDegreesU), 0, 0)
for u in range(K1):
if blockDegreesU[u] > 0:
elist[0, tickerU[u]:tickerU[u+1]] = np.random.choice(np.arange(n), size=blockDegreesU[u],
replace=True, p=Xt[u])
blockDegreesV = np.sum(tabUV, axis=0)
tickerV = np.insert(np.cumsum(blockDegreesV), 0, 0)
for v in range(K2):
if blockDegreesV[v] > 0:
eitmp[tickerV[v]:tickerV[v+1]] = np.random.choice(np.arange(d), size=blockDegreesV[v],
replace=True, p=Yt[v])
ticker = 0
for u in range(K1):
for v in range(K2):
if tabUV[u,v] > 0:
elist[1, ticker:ticker + tabUV[u,v]] = eitmp[tickerV[v]:tickerV[v] + tabUV[u,v]]
ticker += tabUV[u, v]
elist = elist.T
if not selfLoops:
elist = elist[np.where(elist[:, 0] != elist[:, 1])]
if not directed:
if n != d:
raise Exception("{0} != {1}: Undirected network requests n == d".format(n, d))
elist = np.concatenate((elist, elist[:, ::-1]))
if not PoissonEdges:
e = np.ascontiguousarray(elist)
e_unique = np.unique(e.view([('', np.int), ('', np.int)]))
elist = e_unique.view(np.int).reshape((e_unique.shape[0], 2))
if returnEdgeList:
return elist
else:
A = csc_matrix((np.ones(elist.shape[0], dtype=np.int),
(elist[:, 0], elist[:, 1])),
shape=(n, d), dtype=np.int)
return (A, X, S, Y if returnY else None) if returnParameters else A
# -
np.any(X.toarray() < 0)
np.any(X < 0).sum()
sampled_A1 = fastRG(X.toarray(), S.toarray(), Y.toarray())
fig, ax = plt.subplots(1,5)
plot_matrix(sampler.X_prime, name="X", ax=ax[0])
plot_matrix(sampler.Y_prime, name="Y", ax=ax[2])
plot_matrix(sampler.S_prime, name="S", ax=ax[1])
plot_matrix(sampled_A, name="A", ax=ax[3])
plot_matrix(sampled_A1, name="A1", ax=ax[4])
fig.tight_layout()
# fig, ax = plt.subplots(1,5)
ax = [None, None, None, None, None]
plot_matrix(sampler.X_prime, name="$\widetilde{X}$", ax=ax[0])
plot_matrix(sampler.Y_prime, name="$\widetilde{Y}$", ax=ax[2])
plot_matrix(sampler.S_prime, name="$\widetilde{S}$", ax=ax[1])
plot_matrix(sampled_A, name="$\widetilde{A}$", ax=ax[3])
plot_matrix(sampled_A1, name="$\widetilde{A_{FastRG}}$", ax=ax[4])
# fig.tight_layout()
sampled_A.todense()
sns.heatmap(
sampled_A.todense(), cmap="viridis",
# vmin=0, vmax=1,
linewidths=.5,
square=True, cbar=False,
# ax=ax
)
# ax.set_title(name)|
| FastRG - Python implementation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Chapter 1 Tutorial
#
# You can use NetworkX to construct and draw graphs that are undirected or directed, with weighted or unweighted edges. An array of functions to analyze graphs is available. This tutorial takes you through a few basic examples and exercises.
#
# Note that many exercises are followed by a block with some `assert` statements. These assertions may be preceded by some setup code. They are provided to give you feedback that you are on the right path -- receiving an `AssertionError` probably means you've done something wrong.
#
# ## Official documentation for version used in this tutorial
#
# https://networkx.github.io/documentation/networkx-2.2/
#
# ## Official tutorial for version used in this tutorial
#
# https://networkx.github.io/documentation/networkx-2.2/tutorial.html
# # The `import` statement
#
# Recall that `import` statements go at the top of your code, telling Python to load an external module. In this case we want to load NetworkX, but give it a short alias `nx` since we'll have to type it repeatedly, hence the `as` statement.
#
# Lines starting with the `%` character are not Python code, they are "magic" directives for Jupyter notebook. The `%matplotlib inline` magic tells Jupyter Notebook to draw graphics inline i.e. in the notebook. This magic should be used right after the import statement.
import networkx as nx
# %matplotlib inline
# Let's check the installed version of NetworkX. Version 2 is incompatible with v1, so we want to make sure we're not using an out of date package.
nx.__version__
# # Creating and drawing undirected graphs
# +
# a "plain" graph is undirected
G = nx.Graph()
# give each a node a 'name', which is a letter in this case.
G.add_node('a')
# the add_nodes_from method allows adding nodes from a sequence, in this case a list
nodes_to_add = ['b', 'c', 'd']
G.add_nodes_from(nodes_to_add)
# add edge from 'a' to 'b'
# since this graph is undirected, the order doesn't matter here
G.add_edge('a', 'b')
# just like add_nodes_from, we can add edges from a sequence
# edges should be specified as 2-tuples
edges_to_add = [('a', 'c'), ('b', 'c'), ('c', 'd')]
G.add_edges_from(edges_to_add)
# draw the graph
nx.draw(G, with_labels=True)
# -
# There are many optional arguments to the draw function to customize the appearance.
nx.draw(G,
with_labels=True,
node_color='blue',
node_size=1600,
font_color='white',
font_size=16,
)
# # A note on naming conventions
#
# Usually in Python, variables are named in `snake_case`, i.e. lowercase with underscores separating words. Classes are conventionally named in `CamelCase`, i.e. with the first letter of each word capitalized.
#
# Obviously NetworkX doesn't use this convention, often using single capital letters for the names of graphs. This is an example of convention leaking from the world of discrete mathematics. Since most of the documentation you will find online uses this convention, we will follow it as well.
# # Graph methods
#
# The graph object has some properties and methods giving data about the whole graph.
# List all of the nodes
G.nodes()
# List all of the edges
G.edges()
# NodeView and EdgeView objects have iterators, so we can use them in `for` loops:
for node in G.nodes:
print(node)
for edge in G.edges:
print(edge)
# Note that the edges are given as 2-tuples, the same way we entered them.
#
# We can get the number of nodes and edges in a graph using the `number_of_` methods.
G.number_of_nodes()
G.number_of_edges()
# Some graph methods take an edge or node as argument. These provide the graph properties of the given edge or node. For example, the `.neighbors()` method gives the nodes linked to the given node:
# list of neighbors of node 'b'
G.neighbors('b')
# For performance reasons, many graph methods return iterators instead of lists. They are convenient to loop over:
for neighbor in G.neighbors('b'):
print(neighbor)
# and you can always use the `list` constructor to make a list from an iterator:
list(G.neighbors('b'))
# # NetworkX functions vs. Graph methods
#
# The previous data are available via graph *methods*, *i.e.* they are called from the graph object:
#
# G.<method_name>(<arguments>)
#
# While several of the most-used NetworkX functions are provided as methods, many more of them are module functions and are called like this:
#
# nx.<function_name>(G, <arguments>)
#
# that is, with the graph provided as the first, and maybe only, argument. Here are a couple of examples of NetworkX module functions that provide information about a graph:
nx.is_tree(G)
nx.is_connected(G)
# # Node and edge existence
#
# To check if a node is present in a graph, you can use the `has_node()` method:
G.has_node('a')
G.has_node('x')
# Additionally, the loop syntax used above: `for n in G.nodes` suggests another way we can check if a node is in a graph:
'd' in G.nodes
# Likewise we can check if two nodes are connected by an edge:
G.has_edge('a', 'b')
G.has_edge('a', 'd')
('c', 'd') in G.edges
# # Node degree
#
# One of the most important questions we can ask about a node in a graph is how many other nodes it connects to. Using the `.neighbors()` method from above, we could formulate this question as so:
len(list(G.neighbors('a')))
# but this is such a common task that NetworkX provides us a graph method to do this in a much clearer way:
G.degree('a')
# # EXERCISE 1
# Often in the context of trees, a node with degree 1 is called a *leaf*. Write a function named `get_leaves` that takes a graph as an argument, loops through the nodes, and returns a list of nodes with degree 1.
def get_leaves(G):
return [node for node in G.nodes if G.degree(node) == 1]
G = nx.Graph()
G.add_edges_from([
('a', 'b'),
('a', 'd'),
('c', 'd'),
])
assert set(get_leaves(G)) == {'c', 'b'}
# # Aside: comprehensions
#
# Often we have one sequence of values and we want to generate a new sequence by applying an operation to each item in the first. List comprehensions and generator expressions are compact ways to do this.
#
# List comprehensions are specified inside square brackets, and immediately produce a list of the result.
items = ['spider', 'y', 'banana']
[item.upper() for item in items]
# In the context of NetworkX, this is often used to do something with the node or edge lists:
print(G.nodes())
print([G.degree(n) for n in G.nodes()])
# Generator expressions are slightly different as they are evaluated [lazily](https://en.wikipedia.org/wiki/Lazy_evaluation). These are specified using round braces, and if they are being expressed as a function argument, they can be specified without any braces. These are most often used in the context of aggregations like the `max` function:
g = (len(item) for item in items)
list(g)
max(len(item) for item in items)
sorted(item.upper() for item in items)
# # Node names
#
# The node names don't have to be single characters -- they can be strings or integers or any immutable object, and the types can be mixed. The example below uses strings and integers for names.
# +
G = nx.Graph()
G.add_nodes_from(['cat','dog','virus',13])
G.add_edge('cat','dog')
nx.draw(G, with_labels=True, font_color='white', node_size=1000)
# -
# # Adjacency lists
#
# One compact way to represent a graph is an adjacency list. This is most useful for unweighted graphs, directed or undirected. In an adjacency list, each line contains some number of node names. The first node name is the "source" and each other node name on the line is a "target". For instance, given the following adjacency list:
# ```
# a d e
# b c
# c
# d
# e
# ```
# the edges are as follows:
# ```
# (a, d)
# (a, e)
# (b, c)
# ```
# The nodes on their own line exist so that we are sure to include any singleton nodes. Note that if our graph is undirected, we only need to specify one direction for each edge. Importantly, whether the graph is directed or undirected is often not contained in the file itself -- you have to infer it. This is one limitation of the format.
#
# In the `datasets` directory, there is a file called `friends.adjlist`. It's a plain text file, so you can open it on your computer or in GitHub, but here are its contents:
print(open('../datasets/friends.adjlist').read())
# NetworkX provides a way to read a graph from an adjacency list: `nx.read_adjlist()`. We will name this graph SG, for social graph.
SG = nx.read_adjlist('../datasets/friends.adjlist')
# We know how to draw this graph:
nx.draw(SG, node_size=2000, node_color='lightblue', with_labels=True)
# And we know how to get information such as the number of friends linked from a node:
SG.degree('Alice')
# # EXERCISE 2
#
# Write a function max_degree that takes a graph as its argument, and returns a 2-tuple with the name and degree of the node with highest degree.
def max_degree(G):
m = ['', 0]
for node in G.nodes:
deg = G.degree(node)
if deg > m[1]:
m[0] = node
m[1] = deg
return tuple(m)
SG = nx.read_adjlist('../datasets/friends.adjlist')
assert max_degree(SG) == ('Claire', 4)
# # EXERCISE 3
#
# Write a function `mutual_friends` that takes a graph and two nodes as arguments, and returns a list (or set) of nodes that are linked to both given nodes. For example, in the graph `SG` drawn above,
#
# mutual_friends(SG, 'Alice', 'Claire') == ['Frank']
#
# an empty list or set should be returned in the case where two nodes have no mutual friends, e.g. George and Bob in `SG` drawn above.
def mutual_friends(G, node_1, node_2):
a = set(G.neighbors(node_1))
a.intersection_update(set(G.neighbors(node_2)))
return list(a)
SG = nx.read_adjlist('../datasets/friends.adjlist')
assert mutual_friends(SG, 'Alice', 'Claire') == ['Frank']
assert mutual_friends(SG, 'George', 'Bob') == []
assert sorted(mutual_friends(SG, 'Claire', 'George')) == ['Dennis', 'Frank']
# # Directed graphs
#
# Unless otherwise specified, we assume graph edges are undirected -- they are symmetric and go both ways. But some relationships, e.g. predator-prey relationships, are asymmetric and best represented as directed graphs. NetworkX provides the `DiGraph` class for directed graphs.
# +
D = nx.DiGraph()
D.add_edges_from([(1,2),(2,3),(3,2),(3,4),(3,5),(4,5),(4,6),(5,6),(6,4),(4,2)])
nx.draw(D, with_labels=True)
# -
# Note the asymmetry in graph methods dealing with edges such as `has_edge()`:
D.has_edge(1,2)
D.has_edge(2,1)
# Instead of the symmetric relationship "neighbors", nodes in directed graphs have predecessors ("in-neighbors") and successors ("out-neighbors"):
# +
print('Successors of 2:', list(D.successors(2)))
print('Predecessors of 2:', list(D.predecessors(2)))
# -
# Directed graphs have in-degree and out-degree, giving the number of edges pointing to and from the given node, respectively:
D.in_degree(2)
D.out_degree(2)
# ### Caveat
#
# Since NetworkX 2, the `.degree()` method on a directed graph gives the total degree: in-degree plus out-degree. However, in a bit of confusing nomenclature, the `neighbors` method is a synonym for `successors`, giving only the edges originating from the given node. This makes sense if you consider `neighbors` to be all the nodes reachable from the given node by following links, but it's easy to make the mistake of writing `.neighbors()` in your code when you really want both predecessors and successors.
D.degree(2)
print('Successors of 2:', list(D.successors(2)))
print('"Neighbors" of 2:', list(D.neighbors(2)))
| tutorials/Chapter 1 Tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="aK_EtWRb7rr8"
# %%capture
# !pip install cvxopt
import numpy as np
from matplotlib import pyplot as plt
from scipy.stats import chi2
from sklearn.neighbors import KernelDensity
from sklearn.mixture import GaussianMixture
from sklearn.metrics import roc_auc_score
from sklearn.neighbors import NearestNeighbors
from sklearn.ensemble import IsolationForest
import cvxopt
# + [markdown] id="-7rm8vG58BV0"
# ---
# ## 演習問題1 (25点)
#
# 以下の一次元の正規分布の最尤推定問題を考える。
#
# $\hat{\mu}, \hat{\sigma}^2 = \arg\max_{\mu, \sigma^2} \log \prod_{n=1}^N \frac{1}{\sqrt{2 \pi \sigma^2}}\exp\left(-\frac{1}{2 \sigma^2}(x^{(n)} - \mu)^2\right)$
#
# このとき、以下を証明せよ。
#
# * $\hat{\mu} = \frac{1}{N} \sum_{n=1}^N x^{(n)}$
#
# * $\hat{\sigma}^2 = \frac{1}{N} \sum_{n=1}^N \left(x^{(n)} - \hat{\mu}\right)^2$
#
# + [markdown] id="xeDGgTTb-uAR"
# ### 【回答】完了
#
# $\hat{\mu}, \hat{\sigma}^2 = \arg\max_{\mu, \sigma^2} \log \prod_{n=1}^N \frac{1}{\sqrt{2 \pi \sigma^2}}\exp\left(-\frac{1}{2 \sigma^2}(x^{(n)} - \mu)^2\right)$ となるような $\hat{\mu}, \hat{\sigma}^2$ は,それぞれに関して一階微分した値が $0$ となるときである.すなわち,尤度関数 $L \left(\mu, \sigma^2 \right) = \log \prod_{n=1}^N \frac{1}{\sqrt{2 \pi \sigma^2}}\exp\left(-\frac{1}{2 \sigma^2}(x^{(n)} - \mu)^2\right)$ とおくと,それぞれ $\frac{\partial L}{\partial \mu} = 0$ , $\frac{\partial L}{\partial \sigma^2} = 0$ となる $\mu, \sigma^2$ が平均$\hat{\mu}$と分散$\hat{\sigma}^2$の最尤推定量である.
#
# ここで $N$ 個の標本は互いに独立であるから,尤度関数 $L \left(\mu, \sigma^2 \right)$ を以下のように式変形する.
#
# $$
# \begin{aligned}
# L \left(\mu, \sigma^2 \right) &= \log \prod_{n=1}^N \frac{1}{\sqrt{2 \pi \sigma^2}}\exp\left(-\frac{1}{2 \sigma^2}(x^{(n)} - \mu)^2\right) \\
# &= \sum_{n=1}^N \log \left(\frac{1}{\sqrt{2 \pi \sigma^2}}\exp\left(-\frac{1}{2 \sigma^2}(x^{(n)} - \mu)^2\right) \right) \\
# &= N \log \left(\frac{1}{\sqrt{2 \pi \sigma^2}} \right) + \sum_{n=1}^N \log\left(\exp\left(-\frac{1}{2 \sigma^2}(x^{(n)} - \mu)^2\right)\right) \\
# &= N \log \left(\frac{1}{\sqrt{2 \pi \sigma^2}} \right) + \left(-\frac{1}{2 \sigma^2}(x^{(1)} - \mu)^2\right) + \left(-\frac{1}{2 \sigma^2}(x^{(2)} - \mu)^2\right) + \cdots + \left(-\frac{1}{2 \sigma^2}(x^{(N)} - \mu)^2\right) \\
# &= N \log \left(\frac{1}{\sqrt{2 \pi \sigma^2}} \right) -\frac{1}{2 \sigma^2}\left( (x^{(1)} - \mu)^2 + (x^{(2)} - \mu)^2 + \cdots + (x^{(N)} - \mu)^2 \right)
# \end{aligned}
# $$
#
# よって,$\left.\frac{\partial L}{\partial \mu}\right|_{\mu=\hat{\mu}}=0$, $\left.\frac{\partial L}{\partial \sigma^2}\right|_{\mu=\hat{\mu},\sigma^2=\hat{\sigma}^2}=0$ は以下のような解を得る.
#
# $$
# \begin{aligned}
# \left.\frac{\partial L}{\partial \mu}\right|_{\mu=\hat{\mu}} &= -\frac{1}{2 \sigma^2}\left(2(x^{(1)} - \hat{\mu})(-1) + 2(x^{(2)} - \hat{\mu})(-1) + \cdots + 2(x^{(N)} - \hat{\mu})(-1)\right) \\
# &= \frac{1}{\sigma^2}\left((x^{(1)} - \hat{\mu}) + (x^{(2)} - \hat{\mu}) + \cdots + (x^{(N)} - \hat{\mu})\right) \\
# &= \frac{1}{\sigma^2}\left(x^{(1)} + x^{(2)} + \cdots + x^{(N)}\right) - \frac{N}{\sigma^2}\hat{\mu} = 0
# \end{aligned}
# $$
#
# $$
# \begin{aligned}
# \therefore \quad \hat{\mu} = \frac{x^{(1)} + x^{(2)} + \cdots + x^{(N)}}{N} = \frac{1}{N}\sum_{n=1}^N x^{(n)}
# \end{aligned}
# $$
#
# $$
# \begin{aligned}
# \left.\frac{\partial L}{\partial \sigma^2}\right|_{\mu=\hat{\mu},\sigma^2=\hat{\sigma}^2} &= \left.\frac{\partial}{\partial\sigma^2}\left(N\log(2\pi\sigma^2)^{-\frac{1}{2}}-\frac{(x^{(1)}-\hat{\mu})^2+(x^{(2)}-\hat{\mu})^2+\cdots+(x^{(N)}-\hat{\mu})^2}{2}(\sigma^2)^{-1}\right)\right|_{\hat{\sigma}^2=\hat{\hat{\sigma}}^2} \\
# &= -\frac{1}{2}N\frac{2\pi}{2\pi\hat{\sigma}^2}-\frac{(x^{(1)}-\hat{\mu})^2+(x^{(2)}-\hat{\mu})^2+\cdots+(x^{(N)}-\hat{\mu})^2}{2}(-1)(\hat{\sigma}^2)^{-2} \\
# &= -\frac{N}{2\hat{\sigma}^2}+\frac{(x^{(1)}-\hat{\mu})^2+(x^{(2)}-\hat{\mu})^2+\cdots+(x^{(N)}-\hat{\mu})^2}{2}(\hat{\sigma}^2)^{-2} = 0
# \end{aligned}
# $$
#
# $$
# \begin{aligned}
# \therefore \quad \hat{\sigma}^2 &= \frac{(x^{(1)}-\hat{\mu})^2+(x^{(2)}-\hat{\mu})^2+\cdots+(x^{(N)}-\hat{\mu})^2}{N} = \frac{1}{N}\sum_{n=1}^N\left(x^{(n)} - \hat{\mu}\right)^2 \\
# &&\quad\blacksquare\end{aligned}
# $$
#
# + [markdown] id="gIKS9LD_8PUl"
# ---
# ## 演習問題2 [発展] (10点)
#
# 指数型分布族に属する分布を一つ選んで、その分布について以下をまとめよ。
# * どのようなデータを生成する分布か?
# * 実数、整数、ベクトル、行列など
# * どのような性質のデータを表現するのに使われるか
# * 密度関数の式は?
# * 最尤推定の式は?
# * 証明も含む
# + [markdown] id="ov4tLphD_THu"
# ### 【回答】完了
#
# **二項分布**
#
# - どのようなデータを生成する分布か?
# - 確率変数(事象) $X\in\{0, 1\}$ において 各独立試行の確率 $\mathbf{P}(X)=\left\{\begin{aligned}\theta\quad & (X=1) \\1-\theta\quad & (X=0)\end{aligned}\right|0 \leq \theta \leq 1$ を繰り返すことをベルヌーイ試行という.互いに独立したベルヌーイ試行を $n$ 回繰り返したときに,ある事象 $X\in\{0,1\}$ が何回起こるかの確率分布である.
# - したがって,実数 $\begin{bmatrix}0,1\end{bmatrix}$ の確率を生成する分布であり,「コインを投げたときに表が出るか裏が出るか」のように何かを行ったときに起こる結果(事象)が2つしかないベルヌーイ試行を繰り返しある事象が従う確率のデータを表現するのに使われる.
# - 密度関数の式は?
#
# 全試行回数を $n$ ,事象 $X$ の起こる回数を $x$ ,あるパラメータ $/theta$ に従う確率密度関数をf(x;\theta)とすると,下記のように表せる.
# $$
# f(x;\theta)=\left(\begin{aligned}n\\x\end{aligned}\right)\theta^x(1-\theta)^{n-x}={}_n\mathrm{C}_x\theta^x(1-\theta)^{n-x}
# $$
#
# - 最尤推定の式は?
#
# 尤度関数 $L(\theta;x)={}_n\mathrm{C}_x\theta^x(1-\theta)^{n-x}$ が最大となるようなパラメータ $\theta=\hat\theta$ を求める.計算の簡単化のために $L(\theta;x)$ と $\log L(\theta;x)$ が単調な関係であることを利用し,尤度関数の対数の最大化に問題を置き換える.
# $$
# \begin{aligned}
# \hat\theta&=\arg\max_\theta\log L(\theta;x) \\
# &=\arg\max_\theta\log \left({}_n\mathrm{C}_x\theta^x(1-\theta)^{n-x}\right) \\
# &=\arg\max_\theta\left(\log{}_n\mathrm{C}_x+\log\theta^x+\log(1-\theta)^{n-x}\right) \\
# &=\arg\max_\theta\left(\log{}_n\mathrm{C}_x+x\log\theta+(n-x)\log(1-\theta)\right)
# \end{aligned}
# $$
# $l(\theta)=\log L(\theta;x)$ とおくと $\left.\frac{\partial l}{\partial\theta}\right|_{\theta=\hat\theta} = 0$ となるような $\theta=\hat\theta$ が対数尤度関数を最大化する最尤推定量である.
# $$
# \begin{aligned}
# \left.\frac{\partial l}{\partial\theta}\right|_{\theta=\hat\theta} &= \left.\frac{\partial}{\partial\theta}\right|_{\theta=\hat\theta}\log{}_n\mathrm{C}_x+\left.\frac{\partial}{\partial\theta}\right|_{\theta=\hat\theta}x\log\theta+\left.\frac{\partial}{\partial\theta}\right|_{\theta=\hat\theta}(n-x)\log(1-\theta) \\
# &=\frac{x}{\hat\theta}-\frac{n-x}{1-\hat\theta} = 0
# \end{aligned}
# $$
# $$
# \begin{aligned}
# \therefore \quad \frac{x}{\hat\theta}&=\frac{n-x}{1-\hat\theta} \\
# x(1-\hat\theta)&=\hat\theta(n-x) \\
# x-x\hat\theta&=n\hat\theta-x\hat\theta \\
# \hat\theta&=\frac{x}{n} \\
# &&\quad\blacksquare\end{aligned}
# $$
#
# + [markdown] id="j63caQu6899m"
# ---
# ## 演習問題3 (25点)
#
# 正規分布に従わないデータでは、Hotteling’s $T^2$で計算される閾値𝛿と実際の誤判定との割合とが一致しないことを実験的に確認せよ。
# * 正規分布以外の分布からデータを生成して閾値𝛿と実際の誤判定割合とを比較せよ。
# * どのようなデータ・設定で実験をしたかをきちんと記すこと。
# + [markdown] id="zRttYQOB_bSj"
# ### 【回答】完了
#
# 間違え確率を $\epsilon=0.05$ ,閾値 $\delta=3.841458820694124\because\epsilon=1-\int_0^\delta\chi^2(u;1,1) .$ と設定した.一様分布からデータを生成した.閾値 $\delta$ と実際の誤判定割合とを比較すると,N数増加に伴い誤判定割合は単調減少し割合0に収束した.間違え確率 $\epsilon$ に従うことはなかった.
#
# + id="pjP9-RD2_Wpm" colab={"base_uri": "https://localhost:8080/", "height": 441} executionInfo={"status": "ok", "timestamp": 1608905173491, "user_tz": -540, "elapsed": 5525, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "03431199727064994008"}} outputId="13b70b05-bfe6-4b77-a39e-cec71d060bc5"
# 間違え確率と閾値
eps = 0.05
delta = chi2.ppf(1-eps, 1)
# 学習データの個数を変えながら実際に間違えた割合を計算
n_train = np.logspace(1, 3, 11)
c = []
for i, n in enumerate(n_train):
n = int(n)
ci = []
# 100回データを生成して評価
for seed in range(100):
# データ生成
np.random.seed(seed+n)
x_train = np.random.randn(n) # 標準正規分布
x_test = np.random.randn(1000) # テストデータは1000個で固定
# 平均、標準偏差の推定
m = np.mean(x_train)
s = np.std(x_train)
# 指標の計算
a = (x_test - m)**2 / s**2
# 誤りの個数をカウント
ci.append(np.mean(a >= delta))
c.append([np.mean(ci), np.std(ci)])
c = np.array(c)
# 結果を描画
myfig = plt.figure(figsize=(12,6))
ax = plt.subplot(1, 2, 1)
ax.set_xscale('log')
plt.plot(n_train, c[:, 0], 'b-')
plt.plot(n_train, [0.05]*n_train.size, 'k--')
plt.xlabel('# of samples N', fontsize=18)
plt.ylabel('Average Miss-Detection Rate', fontsize=18)
plt.legend(['rate', 'eps'], fontsize=15)
plt.title('the “standard normal” distribution', fontsize=18)
myfig.tight_layout()
# plt.show()
# 間違え確率と閾値
eps = 0.05
delta = chi2.ppf(1-eps, 1)
# 学習データの個数を変えながら実際に間違えた割合を計算
n_train = np.logspace(1, 3, 11)
c = []
for i, n in enumerate(n_train):
n = int(n)
ci = []
# 100回データを生成して評価
for seed in range(100):
# データ生成
np.random.seed(seed+n)
x_train = np.random.rand(n) # 一様分布[0,1)
x_test = np.random.rand(1000) # テストデータは1000個で固定
# x_train = np.random.beta(2, 2, n) # beta分布
# x_test = np.random.beta(2, 2, 1000) # テストデータは1000個で固定
# x_train = np.random.binomial(n, 0.5) # 二項分布
# x_test = np.random.binomial(1000, 0.5) # テストデータは1000個で固定
# x_train = np.random.chisquare(5, n) # χ^2分布
# x_test = np.random.chisquare(5, 1000) # テストデータは1000個で固定
# 平均、標準偏差の推定
m = np.mean(x_train)
s = np.std(x_train)
# 指標の計算
a = (x_test - m)**2 / s**2
# 誤りの個数をカウント
ci.append(np.mean(a >= delta))
c.append([np.mean(ci), np.std(ci)])
c = np.array(c)
# 結果を描画
ax = plt.subplot(1, 2, 2)
ax.set_xscale('log')
plt.plot(n_train, c[:, 0], 'b-')
plt.plot(n_train, [0.05]*n_train.size, 'k--')
plt.xlabel('# of samples N', fontsize=18)
plt.ylabel('Average Miss-Detection Rate', fontsize=18)
plt.legend(['rate', 'eps'], fontsize=15)
plt.title('a uniform distribution', fontsize=18)
# plt.title('a beta distribution', fontsize=18)
# plt.title('a binomial distribution', fontsize=18)
# plt.title('a chi-square distribution', fontsize=18)
myfig.tight_layout()
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="uJ_J9P0EMsxV" executionInfo={"status": "ok", "timestamp": 1608905173494, "user_tz": -540, "elapsed": 5517, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "03431199727064994008"}} outputId="c1b17bc4-3f24-456f-8163-5bcafc9f3181"
print(delta)
# + [markdown] id="1VAkcXL69ekq"
# ---
# ## 演習問題4 [発展](10点)
#
# 以下の$d$次元の正規分布の最尤推定問題を考える。
#
# $\hat{\mu}, \hat{\Sigma} = \arg\max_{\mu, \Sigma} \log \prod_{n=1}^N \frac{1}{(2 \pi)^{d/2} \sqrt{\det \Sigma}}\exp\left(-\frac{1}{2}(x^{(n)} - \mu)^\top \Sigma^{-1} (x^{(n)} - \mu)\right)$
#
# このとき、以下を証明せよ。
#
# * $\hat{\mu} = \frac{1}{N} \sum_{n=1}^N x^{(n)}$
#
# * $\hat{\Sigma} = \frac{1}{N} \sum_{n=1}^N \left(x^{(n)} - \hat{\mu}\right) \left(x^{(n)} - \hat{\mu}\right)^\top$
# + [markdown] id="Yu3W1NKSAUa3"
# ### 【回答】完了
#
# $\hat{\mu}, \hat{\Sigma} = \arg\max_{\mu, \Sigma} \log \prod_{n=1}^N \frac{1}{(2 \pi)^{d/2} \sqrt{\det \Sigma}}\exp\left(-\frac{1}{2}(x^{(n)} - \mu)^\top \Sigma^{-1} (x^{(n)} - \mu)\right)$ となるような $\hat{\mu}, \hat{\Sigma}$ は,それぞれに関して一階微分した値が $0$ となるときである.すなわち,尤度関数 $L \left(\mu, \Sigma \right) = \log \prod_{n=1}^N \frac{1}{\sqrt{2 \pi \Sigma}}\exp\left(-\frac{1}{2 \Sigma}(x^{(n)} - \mu)\right)$ とおくと,それぞれ $\frac{\partial L}{\partial \mu} = 0$ , $\frac{\partial L}{\partial \Sigma} = 0$ となる $\mu, \Sigma$ が平均$\hat{\mu}$と分散$\hat{\Sigma}$の最尤推定量である.
#
# ここで $N$ 個の標本は互いに独立であるから,尤度関数 $L \left(\mu, \Sigma \right)$ を以下のように式変形する.
#
# $$
# \begin{aligned}
# L \left(\mu, \Sigma \right) &= \log \prod_{n=1}^N \frac{1}{(2 \pi)^{d/2} \sqrt{\det \Sigma}}\exp\left(-\frac{1}{2}(x^{(n)} - \mu)^\top \Sigma^{-1} (x^{(n)} - \mu)\right) \\
# &= \sum_{n=1}^N \log \left(\frac{1}{(2 \pi)^{d/2} \sqrt{\det \Sigma}}\exp\left(-\frac{1}{2}(x^{(n)} - \mu)^\top \Sigma^{-1} (x^{(n)} - \mu)\right)\right) \\
# &= N \log \left(\frac{1}{(2 \pi)^{d/2} \sqrt{\det \Sigma}} \right) + \sum_{n=1}^N \log\left(\exp\left(-\frac{1}{2}(x^{(n)} - \mu)^\top \Sigma^{-1} (x^{(n)} - \mu)\right)\right) \\
# &= N \log \left((2 \pi)^{-\frac{d}{2}}\det\Sigma^{-\frac{1}{2}}\right) + \left(-\frac{1}{2}\Sigma^{-1}(x^{(1)} - \mu)^\top(x^{(1)} - \mu)\right) + \left(-\frac{1}{2}\Sigma^{-1}(x^{(2)} - \mu)^\top(x^{(2)} - \mu)\right) + \cdots + \left(-\frac{1}{2}\Sigma^{-1}(x^{(N)} - \mu)^\top(x^{(N)} - \mu)\right) \\
# &= -\frac{Nd}{2}\log\left(2 \pi\right)-\frac{N}{2}\log\left(\det\Sigma\right) -\frac{1}{2}\Sigma^{-1}\left( (x^{(1)} - \mu)^\top(x^{(1)} - \mu) + (x^{(2)} - \mu)^\top(x^{(2)} - \mu) + \cdots + (x^{(N)} - \mu)^\top(x^{(N)} - \mu) \right) \\
# \end{aligned}
# $$
#
# よって,$\left.\frac{\partial L}{\partial \mu}\right|_{\mu=\hat{\mu}}=0$, $\left.\frac{\partial L}{\partial \Sigma}\right|_{\mu=\hat{\mu},\Sigma=\hat{\Sigma}}=0$ は以下のような解を得る.
#
# $$
# \begin{aligned}
# \left.\frac{\partial L}{\partial \mu}\right|_{\mu=\hat{\mu}} &= -\frac{1}{2}\Sigma^{-1}\left(2(x^{(1)} - \hat{\mu})(-1) + 2(x^{(2)} - \hat{\mu})(-1) + \cdots + 2(x^{(N)} - \hat{\mu})(-1)\right) \\
# &= \Sigma^{-1}\left((x^{(1)} - \hat{\mu}) + (x^{(2)} - \hat{\mu}) + \cdots + (x^{(N)} - \hat{\mu})\right) \\
# &= \Sigma^{-1}\left(x^{(1)} + x^{(2)} + \cdots + x^{(N)}\right) - N\Sigma^{-1}\hat{\mu} = 0
# \end{aligned}
# $$
#
# $$
# \begin{aligned}
# \therefore \quad \hat{\mu} = \frac{x^{(1)} + x^{(2)} + \cdots + x^{(N)}}{N} = \frac{1}{N}\sum_{n=1}^N x^{(n)}
# \end{aligned}
# $$
#
# $$
# \begin{aligned}
# \left.\frac{\partial L}{\partial \Sigma}\right|_{\mu=\hat{\mu},\Sigma=\hat{\Sigma}} &= -\frac{N}{2}\frac{1}{\det\hat\Sigma}\left.\frac{\partial\det\Sigma}{\partial\Sigma}\right|_{\Sigma=\hat\Sigma}-\frac{(x^{(1)} - \hat\mu)^\top(x^{(1)} - \hat\mu) + (x^{(2)} - \hat\mu)^\top(x^{(2)} - \hat\mu) + \cdots + (x^{(N)} - \hat\mu)^\top(x^{(N)} - \hat\mu)}{2}(-1)\hat\Sigma^{-1}\hat\Sigma^{-1} \\
# &= -\frac{N}{2}\frac{1}{\det\hat\Sigma}\det\hat\Sigma\left(\hat\Sigma^{-1}\right)^\top + \frac{(x^{(1)} - \hat\mu)^\top(x^{(1)} - \hat\mu) + (x^{(2)} - \hat\mu)^\top(x^{(2)} - \hat\mu) + \cdots + (x^{(N)} - \hat\mu)^\top(x^{(N)} - \hat\mu)}{2}\left(\hat\Sigma^{-1}\right)^2 \\
# &= -\frac{N}{2}\left(\hat\Sigma^{-1}\right)^\top+\frac{(x^{(1)} - \hat\mu)^\top(x^{(1)} - \hat\mu) + (x^{(2)} - \hat\mu)^\top(x^{(2)} - \hat\mu) + \cdots + (x^{(N)} - \hat\mu)^\top(x^{(N)} - \hat\mu)}{2}\left(\hat{\Sigma}^{-1}\right)^2 \\
# &= -\frac{N}{2}\hat\Sigma^{-1}+\frac{(x^{(1)} - \hat\mu)^\top(x^{(1)} - \hat\mu) + (x^{(2)} - \hat\mu)^\top(x^{(2)} - \hat\mu) + \cdots + (x^{(N)} - \hat\mu)^\top(x^{(N)} - \hat\mu)}{2}\left(\hat{\Sigma}^{-1}\right)^2 \quad\because\hat\Sigma^{-1}は正定値(対称行列)\\
# &= 0
# \end{aligned}
# $$
#
# $$
# \begin{aligned}
# \therefore \quad \hat{\Sigma} &= \frac{(x^{(1)} - \hat\mu)^\top(x^{(1)} - \hat\mu) + (x^{(2)} - \hat\mu)^\top(x^{(2)} - \hat\mu) + \cdots + (x^{(N)} - \hat\mu)^\top(x^{(N)} - \hat\mu)}{N} \\
# &= \frac{1}{N}\sum_{n=1}^N\left(x^{(n)} - \hat\mu\right)\left(x^{(n)} - \hat\mu\right)^\top \\
# &&\quad\blacksquare\end{aligned}
# $$
#
# + [markdown] id="6lCOijd49_k9"
# ---
# ## 演習問題5 (20点)
#
# * カーネル密度推定を使って外れ値検知をし、結果をAUROCで評価せよ。
# * 混合正規分布を使って外れ値検知をし、結果をAUROCで評価せよ。
# + [markdown] id="hXQDUZSPAdF_"
# ### 【回答】未完
# (原:回答時にはこのコメントを消すこと)
#
# 以下のコードに追記する。
#
# 回答をここに記入して、ページをPDF出力しての提出も可。
# + id="IjmWKXTIA2eE"
# 乱数シードの固定
np.random.seed(0)
# 分布のパラメータ
m1 = np.zeros(2) # 正規分布1の平均
S1 = np.identity(2) # 正規分布1の分散共分散行列
m2 = np.array([6, 4]) # 正規分布2の平均
S2 = np.array([[1, -0.5], [-0.5, 1]]) # 正規分布2の分散共分散行列
ma = np.array([-3, -3]) # 異常データの正規分布の平均
Sa = np.array([[1, 0.5], [0.5, 1]]) # 異常データの正規分布の分散共分散行列
# データ:学習用
x = np.random.multivariate_normal(m1, S1, 120)
x = np.r_[x, np.random.multivariate_normal(m2, S2, 80)]
# データ:テスト用
y = np.random.multivariate_normal(m1, S1, 200)
y = np.r_[y, np.random.multivariate_normal(m2, S2, 80)]
y = np.r_[y, np.random.multivariate_normal(ma, Sa, 20)] # 外れ値20個を追加
# カーネル密度推定
kde = KernelDensity(kernel='gaussian', bandwidth=0.5).fit(x)
# 混合正規分布推定
gmm = GaussianMixture(n_components=2).fit(x)
# # Following the codes I added.
# print(y.shape)
# print(kde.score_samples(y).shape)
# print(gmm.score_samples(y).shape)
# auroc_kde = roc_auc_score(y, -kde.score_samples(y))
# auroc_gmm = roc_auc_score(y, -gmm.score_samples(y))
# print("AUROC")
# print("KDE : {}".format(auroc_kde))
# print("GMM : {}".format(auroc_gmm))
# + [markdown] id="gzq8cF4B-TyX"
# ---
# ## 演習問題6 (60点)
#
# 正規分布、カーネル密度推定、混合正規分布、LOF、SVDD、Isolation Forestを使った外れ値検知それぞれについて結果を比較せよ。それぞれの手法について得意なデータ・不得意なデータの例を述べよ。
# * 色々なデータでそれぞれの方法を比較し、どのような場合にどの方法が高い精度で外れ値が検知できるかを検証せよ。
# * 例 正規分布の場合は? 一様分布の場合は? 分布の山が複数ある場合は?
# * 次元が1の場合は? 次元が10の場合は? 次元が100の場合は?
# + [markdown] id="bl9lM3dC-hK6"
# ### 【回答】完了
#
# それぞれ正規分布,一様分布,2つの正規分布の重ね合わせの3つの分布に対する各外れ値検知手法の結果を,次元数ごと(10次元,1次元,100次元)に以下に示した.
#
# **10 Dimension**
#
# | |Normal |Uniform |Two Normal|
# |:-|:-:|:-:|:-:|
# |Gauss |0.969956 |0.956502 |0.817280|
# |KDE |0.956619 |0.956543 |0.843427|
# |GMM |0.969675 |0.955018 |0.921648|
# |LOF |0.924411 |0.951527 |0.864953|
# |SVDD |0.969447 |0.957893 |0.677201|
# |IF |0.965383 |0.952174 |0.882929|
#
# **1 Dimension**
#
# | |Normal |Uniform |Two Normal|
# |:-|:-:|:-:|:-:|
# |Gauss |0.441611 |0.861795 |0.473761|
# |KDE |0.448975 |0.861655 |0.444264|
# |GMM |0.455618 |0.847756 |0.421002|
# |LOF |0.535485 |0.833851 |0.523922|
# |SVDD |0.441867 |0.862890 |0.510697|
# |IF |0.419854 |0.851878 |0.432352|
#
# **100 Dimension**
#
# | |Normal |Uniform |Two Normal|
# |:-|:-:|:-:|:-:|
# |Gauss |1.0 |0.934353 |1.0|
# |KDE |1.0 |0.756759 |1.0|
# |GMM |1.0 |0.929496 |1.0|
# |LOF |1.0 |0.920240 |1.0|
# |SVDD |1.0 |0.939838 |1.0|
# |IF |1.0 |0.904891 |1.0|
#
# 以上の結果を以下に言語化した.
#
# - 正規分布に対する外れ値検知精度は,各手法総じて高い.10次元ではLOF以外95%超えである.
# - 一様分布に対する外れ値検知精度は,各手法総じて高い.10次元では95%超えである.
# - 2つの正規分布の重ね合わせに対する外れ値検知精度は,GMMの精度が92.2%と最も高くSVDDの精度が67.7%と最も低かった.(GMMは混合正規分布を用いたものであるから自明の理であると考えられる.)
#
# 次元数の外れ値検知精度への影響を以下に記した.
#
# - 1次元の正規分布と2つの正規分布の重ね合わせに対してはLOFの精度が最も高かった.
# - 100次元では,正規分布と2つの正規分布の重ね合わせに対しては各手法すべて精度100%と最大値であった.
# - 次元数を変化させても一様分布に対する各精度に大きな変化は見受けられなかった.
#
# 以上の結果から,次のようにまとめた.
#
# - 次元数が低い(本試行では1次元)場合,LOFが外れ値検知に有効である.
# - 次元数が適度な(本試行では10次元)場合,GMMが外れ値検知に有効である.
# - 次元数が高い(本試行では100次元)場合,各手法が有効である.
# - 正規分布に従わない分布(本試行では一様分布)の場合,各手法で一定の効果を得られる.
#
# + id="1j48F8sDmNY5" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608905173495, "user_tz": -540, "elapsed": 5504, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "03431199727064994008"}} outputId="191a8f1d-d436-4bbc-dcb9-615fbd01a1d9"
# データの生成(例1)正規分布
def data(dim=2, num_tr=200, num_te=2000, seed=0):
# 乱数シードの固定
np.random.seed(seed)
# 分布のパラメータ
m1 = np.zeros(dim) # 正常データ(正規分布)の平均
S1 = np.identity(dim) # 正常データ(正規分布)の分散共分散行列
m2 = np.random.rand(dim) # 異常データ(正規分布)の平均
L = np.random.rand(dim, dim)
S2 = L.dot(L.T)
scale = 1 / np.sqrt(np.diag(S2))
S2 = S2 / np.outer(scale, scale) # 異常データ(正規分布)の分散共分散行列
# データ:学習用
x = np.random.multivariate_normal(m1, S1, int(1.0*num_tr))
x = np.r_[x, np.random.multivariate_normal(m2, S2, int(0.0*num_tr))]
# データ:テスト用
y = np.random.multivariate_normal(m1, S1, int(0.95*num_te))
y = np.r_[y, np.random.multivariate_normal(m2, S2, int(0.05*num_te))]
h = np.r_[np.zeros(int(0.95*num_te)), np.ones(int(0.05*num_te))]
return x, y, h
x, y, h = data()
print(x.shape,y.shape,h.shape) # (200, 2) (2000, 2) (2000,)
# + id="XoqRxZMdmPUn"
def solveQP(P, q, C):
m = P.shape[0]
P = cvxopt.matrix(P)
q = cvxopt.matrix(q)
A = cvxopt.matrix(np.ones((1,m)))
b = cvxopt.matrix(np.array([1.0]))
G = cvxopt.matrix(np.r_[-np.identity(m), np.identity(m)])
h = cvxopt.matrix(np.r_[np.zeros(m), C*np.ones(m)])
cvxopt.solvers.options['show_progress'] = False
sol = cvxopt.solvers.qp(P,q,A=A,b=b,G=G,h=h)
ans = np.transpose(np.array(sol['x']))
return ans[0]
# + id="B4zzkhN8B8BV"
class Gauss:
def fit(self, x):
self.mu = np.mean(x, axis=0)
self.sigma = np.cov(x.T, bias=True)
return self
def anomaly_score(self, x):
m = x - self.mu[np.newaxis, :]
if self.sigma.size > 1:
p = np.linalg.solve(self.sigma, m.T).T
else:
p = self.sigma * m
return np.sum(p * m, axis=1)
class LOF:
def fit(self, x):
self.x = x.copy()
self.nbrs = NearestNeighbors(n_neighbors=2, algorithm='ball_tree').fit(x)
return self
def anomaly_score(self, x):
n = x.shape[0]
dist, ind = self.nbrs.kneighbors(x)
a = []
for i in range(n):
dist_sub, _ = self.nbrs.kneighbors(self.x[ind[i, 0], :][np.newaxis, :])
a.append(dist[i, 0] / dist_sub[0, 1])
return np.array(a)
class SVDD:
def __init__(self, C=0.1, tol=1e-6):
self.C = C
self.tol = tol
def fit(self, x):
# 最適化問題を解く
K = x.dot(x.T)
P = 2 * K
q = - np.diag(K)
self.beta = solveQP(P, q, self.C)
# 球の中心bと半径Rを求める
idx_boundary = np.where((self.beta >= self.tol) * (self.beta <= self.C - self.tol))[0]
self.b = x.T.dot(self.beta)
if self.b.size > 1:
self.R = np.linalg.norm(x[idx_boundary, :] - self.b, axis=1).max()
else:
self.R = x.max() - self.b
# print("x_max : {}, b : {}, R : {}".format(x.max(),self.b,self.R))
return self
def anomaly_score(self, x):
# 球からの"はみ出し"を計算する
return np.linalg.norm(x - self.b, axis=1) - self.R
# + id="jIhodlduFmQX"
def evaluate(x, y, h):
methods = [Gauss(), KernelDensity(kernel='gaussian', bandwidth=0.3), GaussianMixture(n_components=2), LOF(), SVDD(C=0.1), IsolationForest()]
evals = [lambda v: methods[0].anomaly_score(v),
lambda v: -methods[1].score_samples(v),
lambda v: -methods[2].score_samples(v),
lambda v: methods[3].anomaly_score(v),
lambda v: methods[4].anomaly_score(v),
lambda v: -methods[5].decision_function(v)]
auc = []
for m, e in zip(methods, evals):
m = m.fit(x)
p = e(y)
auc.append(roc_auc_score(h, p))
return np.array(auc)
# + id="CDQyn820aRLE"
import pandas as pd
emptyData = {
'Normal': [0.0, 0.0, 0.0, 0.0, 0.0, 0.0],
'Uniform': [9., 9., 9., 9., 9., 9.],
'Two Normal': [9., 9., 9., 9., 9., 9.]
}
method_names = ['Gauss', 'KDE', 'GMM', 'LOF', 'SVDD', 'IF']
df10Dim = pd.DataFrame(emptyData, index = method_names)
df1Dim = pd.DataFrame(emptyData, index = method_names)
df100Dim = pd.DataFrame(emptyData, index = method_names)
# + id="9FFn2Z1NPF1z" colab={"base_uri": "https://localhost:8080/", "height": 202, "referenced_widgets": ["31c64cadda6141efaec34bf48625b40d", "5257aaaecd0d45d8b05ff9f1eda9425d", "54eb324dacc84030aed39b985bd4df26", "ad9d3e893d2a48b2bd43b7b5fca1bb09", "<KEY>", "aa03630a8cc04c27b5ba2482c6570621", "5405f346103e4a7ba939cace1be0b165", "93c1691f1a2a467bb63e14c93e4537b7"]} executionInfo={"status": "ok", "timestamp": 1608905188392, "user_tz": -540, "elapsed": 20369, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "03431199727064994008"}} outputId="00ad835a-da5d-42dd-8be7-cde8466e9d08"
from tqdm.notebook import tqdm
method_names = ['Gauss', 'KDE', 'GMM', 'LOF', 'SVDD', 'IF']
auc = []
for seed in tqdm(range(10)):
x, y, h = data(dim=10, num_tr=500, num_te=2000, seed=seed)
a = evaluate(x, y, h)
auc.append(a)
auc = np.array(auc)
auc_avg = np.mean(auc, axis=0)
print('The Normal Distibution, 10 Dimensions')
print('Average AUROC')
for n, a in zip(method_names, auc_avg):
print('%s: %.3f' % (n, a))
df10Dim['Normal'][n] = a
# + colab={"base_uri": "https://localhost:8080/", "height": 202, "referenced_widgets": ["ec9400bc498d42a0a2d7fcec4ff5444f", "911300d7dd4745828b201ee03f44ca00", "222e4f897b6e40e5ad734cdc19681969", "5b2f198132fd4dffb18afe637405cadb", "<KEY>", "d06372c3754f4b2e980cb9074ac2214d", "da79681f79ee4f9ab72f77c8a33aba01", "b6cd43a5a6b740109c7dc5fae0cc04df"]} id="J3EvTauLAbo7" executionInfo={"status": "ok", "timestamp": 1608905202267, "user_tz": -540, "elapsed": 34235, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "03431199727064994008"}} outputId="eadd99b3-82a0-4622-dbeb-6b384e1e9425"
from tqdm.notebook import tqdm
method_names = ['Gauss', 'KDE', 'GMM', 'LOF', 'SVDD', 'IF']
auc = []
for seed in tqdm(range(10)):
x, y, h = data(dim=1, num_tr=500, num_te=2000, seed=seed)
a = evaluate(x, y, h)
auc.append(a)
auc = np.array(auc)
auc_avg = np.mean(auc, axis=0)
print('The Normal Distibution, 1 Dimensions')
print('Average AUROC')
for n, a in zip(method_names, auc_avg):
print('%s: %.3f' % (n, a))
df1Dim['Normal'][n] = a
# + colab={"base_uri": "https://localhost:8080/", "height": 202, "referenced_widgets": ["7335b6ffddea4dc187eaf25396202e2b", "5c3aef70927948eea40f9849a3f099fb", "<KEY>", "0309268d30f4474181db096992dd01ce", "<KEY>", "3b5cb788cfe74e76822a270b697f78b6", "<KEY>", "a08ce52c1f644552a6d6fb846107c411"]} id="sNYjj-8wAbVJ" executionInfo={"status": "ok", "timestamp": 1608905224546, "user_tz": -540, "elapsed": 56505, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "03431199727064994008"}} outputId="496f1df9-c77c-45c7-b715-eebcde90e001"
from tqdm.notebook import tqdm
method_names = ['Gauss', 'KDE', 'GMM', 'LOF', 'SVDD', 'IF']
auc = []
for seed in tqdm(range(10)):
x, y, h = data(dim=100, num_tr=500, num_te=2000, seed=seed)
a = evaluate(x, y, h)
auc.append(a)
auc = np.array(auc)
auc_avg = np.mean(auc, axis=0)
print('The Normal Distibution, 100 Dimensions')
print('Average AUROC')
for n, a in zip(method_names, auc_avg):
print('%s: %.3f' % (n, a))
df100Dim['Normal'][n] = a
# + id="zSQE9ACgPVZJ" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608905224547, "user_tz": -540, "elapsed": 56497, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "03431199727064994008"}} outputId="da51a41d-b123-4d26-8e71-481491c6bf60"
# データの生成(例2)一様分布
def data_uniform(dim=2, num_tr=200, num_te=2000, seed=0):
# 乱数シードの固定
np.random.seed(seed)
# データ:学習用
x = np.random.rand(int(1.0*num_tr),dim)
x = np.r_[x, np.random.rand(int(0.0*num_tr),dim)]
# データ:テスト用
y = np.random.rand(int(0.95*num_te),dim)
y = np.r_[y, np.random.rand(int(0.05*num_te),dim)+np.random.rand()]
h = np.r_[np.zeros(int(0.95*num_te)), np.ones(int(0.05*num_te))]
return x, y, h
x, y, h = data_uniform()
print(x.shape,y.shape,h.shape) # (200, 2) (2000, 2) (2000,)
# + colab={"base_uri": "https://localhost:8080/", "height": 202, "referenced_widgets": ["48ea1ab2a8a649cea832c98d093fbcbb", "7a6f0a8b98594b96949156711e31eed5", "<KEY>", "82eca5c95588457d935f245571edba76", "<KEY>", "<KEY>", "<KEY>", "98b9bbea127846319af2d2818e22ac79"]} id="QySWCsCdVy1F" executionInfo={"status": "ok", "timestamp": 1608905240415, "user_tz": -540, "elapsed": 72356, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "03431199727064994008"}} outputId="3ae2cf88-c996-4562-f981-5dfc847515d0"
from tqdm.notebook import tqdm
method_names = ['Gauss', 'KDE', 'GMM', 'LOF', 'SVDD', 'IF']
auc = []
for seed in tqdm(range(10)):
x, y, h = data_uniform(dim=10, num_tr=500, num_te=2000, seed=seed)
a = evaluate(x, y, h)
auc.append(a)
auc = np.array(auc)
auc_avg = np.mean(auc, axis=0)
print('The Uniform Distibution, 10 Dimensions')
print('Average AUROC')
for n, a in zip(method_names, auc_avg):
print('%s: %.3f' % (n, a))
df10Dim['Uniform'][n] = a
# + colab={"base_uri": "https://localhost:8080/", "height": 202, "referenced_widgets": ["2c0875b842784cae9329d4ca18a459ff", "fc1a5a1f53c842ac9420dd6ce02f9680", "62364f0bff3c4e4797163f3bdc1c96d6", "d71b1ee389b9437b9ba3c12ab9ba808a", "3a8867546fa14d968f99a982891a9734", "fdb3d131cf3e4cb6ba60a63fbcfadc57", "c0840923f4804bd69853c929eb7cf53c", "c0664bda0ac94a36acbe1f89d6f226ec"]} id="bRh1HHU4Aiw_" executionInfo={"status": "ok", "timestamp": 1608905255362, "user_tz": -540, "elapsed": 87293, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "03431199727064994008"}} outputId="26735bfb-9582-40d4-a4ff-ef78f17b91ef"
from tqdm.notebook import tqdm
method_names = ['Gauss', 'KDE', 'GMM', 'LOF', 'SVDD', 'IF']
auc = []
for seed in tqdm(range(10)):
x, y, h = data_uniform(dim=1, num_tr=500, num_te=2000, seed=seed)
a = evaluate(x, y, h)
auc.append(a)
auc = np.array(auc)
auc_avg = np.mean(auc, axis=0)
print('The Uniform Distibution, 1 Dimensions')
print('Average AUROC')
for n, a in zip(method_names, auc_avg):
print('%s: %.3f' % (n, a))
df1Dim['Uniform'][n] = a
# + colab={"base_uri": "https://localhost:8080/", "height": 202, "referenced_widgets": ["868fabe747bc4107841125834f2cd29b", "1898c3a108a84f809d51835608a9c661", "1fe0623571d2412fab3c22a3ee1dd0bd", "5061c153fee14bcab52b872d1a5cee68", "b90856ced408450680702e848a8ec951", "<KEY>", "8f2de568775141988fbdf5bbe2d65000", "69a1b46988e6429eb549cdbdee6b3c83"]} id="XDhpBA2RAip1" executionInfo={"status": "ok", "timestamp": 1608905279022, "user_tz": -540, "elapsed": 110942, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "03431199727064994008"}} outputId="6ec118f8-ed20-4bb0-d220-7347b1b21046"
from tqdm.notebook import tqdm
method_names = ['Gauss', 'KDE', 'GMM', 'LOF', 'SVDD', 'IF']
auc = []
for seed in tqdm(range(10)):
x, y, h = data_uniform(dim=100, num_tr=500, num_te=2000, seed=seed)
a = evaluate(x, y, h)
auc.append(a)
auc = np.array(auc)
auc_avg = np.mean(auc, axis=0)
print('The Uniform Distibution, 100 Dimensions')
print('Average AUROC')
for n, a in zip(method_names, auc_avg):
print('%s: %.3f' % (n, a))
df100Dim['Uniform'][n] = a
# + colab={"base_uri": "https://localhost:8080/"} id="DUmNJgLRtGpN" executionInfo={"status": "ok", "timestamp": 1608905279023, "user_tz": -540, "elapsed": 110934, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "03431199727064994008"}} outputId="9add2133-2b44-4956-ed33-426fc2b2db5f"
# データの生成(例3)2つの正規分布の重ね合わせ
def data_multi(dim=2, num_tr=200, num_te=2000, seed=0):
# 乱数シードの固定
np.random.seed(seed)
# 分布のパラメータ
m0 = np.zeros(dim) # 正常データ0(正規分布)の平均
S0 = np.identity(dim) # 正常データ0(正規分布)の分散共分散行列
m1 = np.ones(dim)*4 # 正常データ1(正規分布)の平均
S1 = np.identity(dim)*2 # 正常データ1(正規分布)の分散共分散行列
m2 = np.random.rand(dim) # 異常データ(正規分布)の平均
L = np.random.rand(dim, dim)
S2 = L.dot(L.T)
scale = 1 / np.sqrt(np.diag(S2))
S2 = S2 / np.outer(scale, scale) # 異常データ(正規分布)の分散共分散行列
# データ:学習用
x = np.random.multivariate_normal(m0, S0, int(0.4*num_tr))
x = np.r_[x, np.random.multivariate_normal(m1, S1, int(0.6*num_tr))]
x = np.r_[x, np.random.multivariate_normal(m2, S2, int(0.0*num_tr))]
# データ:テスト用
y = np.random.multivariate_normal(m0, S0, int(0.38*num_te))
y = np.r_[y, np.random.multivariate_normal(m1, S1, int(0.57*num_te))]
y = np.r_[y, np.random.multivariate_normal(m2, S2, int(0.05*num_te))]
h = np.r_[np.zeros(int(0.95*num_te)), np.ones(int(0.05*num_te))]
return x, y, h
x, y, h = data_multi()
print(x.shape,y.shape,h.shape) # (200, 2) (2000, 2) (2000,)
# + colab={"base_uri": "https://localhost:8080/", "height": 202, "referenced_widgets": ["427cffe96bb34149b95a3f0eb84e41ae", "1372e86d94df450db249ec648b1a8bb4", "<KEY>", "4d7f5eac654a40bdb8e1fba538ff7dd1", "<KEY>", "3bb71c256e4d49dcbb32347e82cee802", "8f46ec554195462f80ee9c97ff2b9b8a", "0d998892306e46d5ada19aff81d65274"]} id="jm3RLqhSxXdl" executionInfo={"status": "ok", "timestamp": 1608905292332, "user_tz": -540, "elapsed": 124234, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "03431199727064994008"}} outputId="f34d7182-2ac1-41c4-d94f-8b2518e7dd4d"
from tqdm.notebook import tqdm
method_names = ['Gauss', 'KDE', 'GMM', 'LOF', 'SVDD', 'IF']
auc = []
for seed in tqdm(range(10)):
x, y, h = data_multi(dim=10, num_tr=500, num_te=2000, seed=seed)
a = evaluate(x, y, h)
auc.append(a)
auc = np.array(auc)
auc_avg = np.mean(auc, axis=0)
print('The Two Normal Distibution, 10 Dimensions')
print('Average AUROC')
for n, a in zip(method_names, auc_avg):
print('%s: %.3f' % (n, a))
df10Dim['Two Normal'][n] = a
# + colab={"base_uri": "https://localhost:8080/", "height": 202, "referenced_widgets": ["329bfb3b97bb4572851028a360c147da", "eeeffc3baf4443f78300462f14b41faf", "b1c46932b87043d6a0a6ab76f0c1f8c1", "8efbd4fbbae94864900d5d34ce3fb27d", "446ed29e27b04d8786a927849c1e5e43", "<KEY>", "1fd2d52882b34048ae12825afbba5a3c", "46b785349a094ef89ec3a4e532bfef37"]} id="VBEbst0IAnUp" executionInfo={"status": "ok", "timestamp": 1608905306199, "user_tz": -540, "elapsed": 138088, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "03431199727064994008"}} outputId="9dc780e4-1676-494c-f235-f0074a86d9ae"
from tqdm.notebook import tqdm
method_names = ['Gauss', 'KDE', 'GMM', 'LOF', 'SVDD', 'IF']
auc = []
for seed in tqdm(range(10)):
x, y, h = data_multi(dim=1, num_tr=500, num_te=2000, seed=seed)
a = evaluate(x, y, h)
auc.append(a)
auc = np.array(auc)
auc_avg = np.mean(auc, axis=0)
print('The Two Normal Distibution, 1 Dimensions')
print('Average AUROC')
for n, a in zip(method_names, auc_avg):
print('%s: %.3f' % (n, a))
df1Dim['Two Normal'][n] = a
# + colab={"base_uri": "https://localhost:8080/", "height": 202, "referenced_widgets": ["b714df6e9fb44cf7a28d1de8cfb15083", "744f4674eaec4612959d64b2dabd1189", "75f66cce0fb44f4a967f3dcbc3573dab", "fe921185de2c4d43a29cf3b5d9d47214", "565b826752144cf38e7173da9dbe6b82", "<KEY>", "1458e2a57fc048679894ed6b2157d24d", "a6247dc7b6804643b019b09c723d49e7"]} id="VBLTx3SYAnPF" executionInfo={"status": "ok", "timestamp": 1608905327247, "user_tz": -540, "elapsed": 159127, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "03431199727064994008"}} outputId="e34aa25f-bb4e-4adc-cb56-e58311d26be5"
from tqdm.notebook import tqdm
method_names = ['Gauss', 'KDE', 'GMM', 'LOF', 'SVDD', 'IF']
auc = []
for seed in tqdm(range(10)):
x, y, h = data_multi(dim=100, num_tr=500, num_te=2000, seed=seed)
a = evaluate(x, y, h)
auc.append(a)
auc = np.array(auc)
auc_avg = np.mean(auc, axis=0)
print('The Two Normal Distibution, 100 Dimensions')
print('Average AUROC')
for n, a in zip(method_names, auc_avg):
print('%s: %.3f' % (n, a))
df100Dim['Two Normal'][n] = a
# + colab={"base_uri": "https://localhost:8080/"} id="bnpPXwUBTRyQ" executionInfo={"status": "ok", "timestamp": 1608905327248, "user_tz": -540, "elapsed": 159118, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "03431199727064994008"}} outputId="3a2d3278-a99f-4131-8d50-ac0c658fc485"
print(" 1 Dimension")
print(df1Dim)
print(" 10 Dimension")
print(df10Dim)
print("100 Dimension")
print(df100Dim)
| AnomalyDetection/kh_CDS_AnomalyDetection_Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
import tensorflow_hub as hub
import tensorflow_datasets as tfds
tfds.disable_progress_bar()
from official.modeling import tf_utils
from official import nlp
from official.nlp import bert
# Load the required submodules
import official.nlp.optimization
import official.nlp.bert.bert_models
import official.nlp.bert.configs
import official.nlp.bert.run_classifier
import official.nlp.bert.tokenization
import official.nlp.data.classifier_data_lib
import official.nlp.modeling.losses
import official.nlp.modeling.models
import official.nlp.modeling.networks
# Set CPU as available physical device
tf.config.set_visible_devices([], 'GPU')
# https://www.tensorflow.org/official_models/fine_tuning_bert
# -
gs_folder_bert = "C:/Users/Ryan/Desktop/projects/ConspiracyBERT/config"
tf.io.gfile.listdir(gs_folder_bert)
# +
glue, info = tfds.load('glue/mrpc', with_info=True, batch_size=-1)
# glue['train'][list(glue['train'].keys())]
# glue_train = glue['train']
# for key, value in glue_train.items():
# print(f"{key:9s}: {value[0].numpy()}")
# +
# Set up tokenizer to generate Tensorflow dataset
tokenizer = bert.tokenization.FullTokenizer(
vocab_file=os.path.join(gs_folder_bert, "vocab.txt"),
do_lower_case=True)
print("Vocab size:", len(tokenizer.vocab))
tokens = tokenizer.tokenize("Hello TensorFlow!")
print(tokens)
ids = tokenizer.convert_tokens_to_ids(tokens)
print(ids)
tokenizer.convert_tokens_to_ids(['[CLS]', '[SEP]'])
# +
def encode_sentence(s, tokenizer):
tokens = list(tokenizer.tokenize(s))
tokens.append('[SEP]')
return tokenizer.convert_tokens_to_ids(tokens)
def bert_encode(glue_dict, tokenizer):
num_examples = len(glue_dict["sentence1"])
sentence1 = tf.ragged.constant([
encode_sentence(s, tokenizer)
for s in np.array(glue_dict["sentence1"])])
sentence2 = tf.ragged.constant([
encode_sentence(s, tokenizer)
for s in np.array(glue_dict["sentence2"])])
cls = [tokenizer.convert_tokens_to_ids(['[CLS]'])]*sentence1.shape[0]
input_word_ids = tf.concat([cls, sentence1, sentence2], axis=-1)
input_mask = tf.ones_like(input_word_ids).to_tensor()
type_cls = tf.zeros_like(cls)
type_s1 = tf.zeros_like(sentence1)
type_s2 = tf.ones_like(sentence2)
input_type_ids = tf.concat(
[type_cls, type_s1, type_s2], axis=-1).to_tensor()
inputs = {
'input_word_ids': input_word_ids.to_tensor(),
'input_mask': input_mask,
'input_type_ids': input_type_ids}
return inputs
# +
glue_train = bert_encode(glue['train'], tokenizer)
glue_train_labels = glue['train']['label']
glue_validation = bert_encode(glue['validation'], tokenizer)
glue_validation_labels = glue['validation']['label']
glue_test = bert_encode(glue['test'], tokenizer)
glue_test_labels = glue['test']['label']
# +
for key, value in glue_train.items():
print(f'{key:15s} shape: {value.shape}')
print(f'glue_train_labels shape: {glue_train_labels.shape}')
# +
import json
bert_config_file = os.path.join(gs_folder_bert, "bert_config.json")
config_dict = json.loads(tf.io.gfile.GFile(bert_config_file).read())
bert_config = bert.configs.BertConfig.from_dict(config_dict)
config_dict
# -
bert_classifier, bert_encoder = bert.bert_models.classifier_model(
bert_config, num_labels=2)
# +
glue_batch = {key: val[:10] for key, val in glue_train.items()}
bert_classifier(
glue_batch, training=True
).numpy()
# -
checkpoint = tf.train.Checkpoint(model=bert_encoder)
checkpoint.restore(
os.path.join(gs_folder_bert, 'bert_model.ckpt')).assert_consumed()
# +
# Set up epochs and steps
epochs = 3
batch_size = 32
eval_batch_size = 32
train_data_size = len(glue_train_labels)
steps_per_epoch = int(train_data_size / batch_size)
num_train_steps = steps_per_epoch * epochs
warmup_steps = int(epochs * train_data_size * 0.1 / batch_size)
# creates an optimizer with learning rate schedule
optimizer = nlp.optimization.create_optimizer(
2e-5, num_train_steps=num_train_steps, num_warmup_steps=warmup_steps)
# -
glue_train['input_word_ids'][1]
# +
metrics = [tf.keras.metrics.SparseCategoricalAccuracy('accuracy', dtype=tf.float32)]
loss = tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True)
bert_classifier.compile(
optimizer=optimizer,
loss=loss,
metrics=metrics)
bert_classifier.fit(
glue_train, glue_train_labels,
validation_data=(glue_validation, glue_validation_labels),
batch_size=32,
epochs=epochs)
# -
| scripts/bert_pretrain.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Quickstart: Using the Speech Service from Python
#
# This sample shows how to use the Speech Service through the Speech SDK for Python. It illustrates how the SDK can be used to synthesize speech to speaker output.
#
# See the [accompanying article](https://docs.microsoft.com/azure/cognitive-services/speech-service/quickstart-text-to-speech-python) on the SDK documentation page for step-by-step instructions.
#
# ## Prerequisites
#
# Before you get started, here's a list of prerequisites:
#
# * A subscription key for the Speech service. See [Try the speech service for free](https://docs.microsoft.com/azure/cognitive-services/speech-service/get-started).
# * Python 3.5 or later needs to be installed. Downloads are available [here](https://www.python.org/downloads/).
# * The Python Speech SDK package is available for Windows (x64 or x86) and Linux (x64; Ubuntu 16.04 or Ubuntu 18.04).
# * On Ubuntu 16.04 or 18.04, run the following commands for the installation of required packages:
# ```sh
# sudo apt-get update
# sudo apt-get install libssl1.0.0 libasound2
# ```
# * On Debian 9, run the following commands for the installation of required packages:
# ```sh
# sudo apt-get update
# sudo apt-get install libssl1.0.2 libasound2
# ```
# * On Windows you need the [Microsoft Visual C++ Redistributable for Visual Studio 2017](https://support.microsoft.com/help/2977003/the-latest-supported-visual-c-downloads) for your platform.
#
# ## Get the Speech SDK Python Package
#
# **By downloading the Microsoft Cognitive Services Speech SDK, you acknowledge its license, see [Speech SDK license agreement](https://aka.ms/csspeech/license201809).**
#
# The Cognitive Services Speech SDK Python package can be installed from [pyPI](https://pypi.org/) using this command:
#
# ```sh
# pip install azure-cognitiveservices-speech
# ```
#
# ## Speech Synthesis Using the Speech SDK
# First, set up some general items. Import the Speech SDK Python:
import azure.cognitiveservices.speech as speechsdk
# Set up the subscription info for the Speech Service:
speech_key, service_region = "YourSubscriptionKey", "YourServiceRegion"
# Create an instance of a speech config with specified subscription key and service region.
# Replace with your own subscription key and service region (e.g., "westus").
speech_config = speechsdk.SpeechConfig(subscription=speech_key, region=service_region)
# Create a synthesizer with the given settings. Since no explicit audio config is specified, the default speaker will be used (make sure the audio settings are correct).
speech_synthesizer = speechsdk.SpeechSynthesizer(speech_config=speech_config)
# Receives a text from user input.
print("Type some text that you want to speak...")
text = input()
# Synthesizes the received text to speech. The synthesized speech is expected to be heard on the speaker with below line executed.
result = speech_synthesizer.speak_text_async(text).get()
# Checks the synthesis result.
if result.reason == speechsdk.ResultReason.SynthesizingAudioCompleted:
print("Speech synthesized to speaker for text [{}]".format(text))
elif result.reason == speechsdk.ResultReason.Canceled:
cancellation_details = result.cancellation_details
print("Speech synthesis canceled: {}".format(cancellation_details.reason))
if cancellation_details.reason == speechsdk.CancellationReason.Error:
if cancellation_details.error_details:
print("Error details: {}".format(cancellation_details.error_details))
print("Did you update the subscription info?")
| quickstart/python/text-to-speech/quickstart.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Welcome to OpenVINO(TM) Hands-on Lab Session 3 - Advanced Features
#
# In this tutorial, we will look at some of the advanced featues of Intel(R) Distribution of OpenVINO(TM) to help you get performance upgrades, using multiple target devices at the same time, creating custom layers and configuring execution of layers manually on different hardware devices.
#
# Jupyter notebook is a browser based IDE and allows partial code executions and text based inputs as markdown at each cell.
#
# Please follow the notations for each instruction in the following sections:
#
# - If tutorial wants you run a certain command on terminal you will see the text as below. You should open a terminal or continue on the opened one as instructed.
#
# **Commands To Run on Terminal**
# ***
# ```bash
# python3
# ```
# ***
#
# In Ubuntu, when you copy text, easiest way to paste code with keyboard is to press:
#
# **SHIFT + INS** buttons
#
# - If there are text looks like code; it is mainly the output of an example run of a sample code, please don't copy those back into terminal.
#
# **Example Terminal Output**
# ***
# ``` output ```
# ***
#
# - If you see python code inside the cell like below, take focus to cell by clicking to it then press:
#
# **SHIFT + ENTER**
#
# or Click on `>| Run` run button above.
# Below example is a code block, which you can run inside this browser session and see the output directly below the cell. You don't need to copy the code to terminal or any other IDE.
# +
import numpy as np
arr1 = np.zeros(5)
print('Array 1 ', arr1)
arr2 = np.ones(5)
print('Array 2', arr2)
# Array Product
print(arr1 * arr2)
# -
# # Agenda
#
# ## Part 1 - Inference Engine - Heterogeneous Plugin
# At this section we will run Inference Engine Heterogeneous Plugin code samples inside Jupyter Notebook to see how Heterogeneous Plugin runs on heterogeneous hardware environments.
#
# ## Part 2 - Inference Engine - Layer Affinity
# This part is a follow-up of Heterogeneous Plugin which shows how manual configuration be applied to DL layer assignment to HW devices.
#
# ## Part 3 - Inference Engine - Performance Counters
# Another feature of Inference Engine will be seen at this part to see performance metrics of layer executions.
#
#
# ## Part 4 - Custom Layers
# Custom layers are NN (Neural Network) layers that are not explicitly supported by a given framework. This tutorial demonstrates how to run inference on topologies featuring custom layers allowing you to plug in your own implementation for existing or completely new layers.
# # Part 1: Intel(R) Distribution of OpenVINO(TM) : Heterogenous Plugin
#
# At this session, we would like to go over more detailed explanation of OpenVINO(TM) Toolkit Inference Engine's Heterogeneous API which helps to run inference on heterogeneous platforms.
#
# If you are running on a platform with Intel CPU and GPU, you can control what layers of Deep Learning Model would be running on the selected hardware platform.
#
# Heterogeneous API has been developed first with fallback principle, it means whenever a deep learning layer not implemented for target platform it should fallback to CPU, which can execute all layers.
#
# Not all layer's implementations are complete for each platform and certain layers can execute a lot faster on different platforms where some can't. As such, there has been great progress over the heterogeneous API use cases for Inference to analyze the performance bottlenecks and improve the inference process.
#
# At this section, we want to showcase its use on object detection scenario and get detailed analysis of executions of layers on devices a quick course of Heterogeneous Plugin of Intel(R) Distribution of OpenVINO(TM) Toolkit.
# ## 1 - Load Libraries & Implement Helper Methods
#
# As in all previous sections, we again use `createNetwork`, `loadNetwork` and `prerocessImage` method before implementing inference with heterogeneous plugin.
#
# #### Change Focus to Below Cell and Press (SHIFT + ENTER) or Click on Run Button
# +
# Let's Import Required Libraries first
import sys
import os
import time
import cv2 as cv
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# Import OpenVINO
from openvino.inference_engine import IENetwork, IEPlugin
# For labeling the image after inference.
from utils.out_process import placeBoxes
# Define Methods
def createNetwork(model_xml, model_bin, plugin):
# Importing network weights from IR models.
net = IENetwork(model=model_xml, weights=model_bin)
return net
def loadNetwork(plugin, net, num_requests=2):
# Loading IR model to the plugin.
exec_net = plugin.load(network=net, num_requests=num_requests)
# Getting the input and outputs of the network
input_blob = next(iter(net.inputs))
out_blob = next(iter(net.outputs))
return exec_net,input_blob,out_blob
def preprocessImage(img_path, net, input_blob):
# Reading the frame from a jpeg file
frame = cv.imread(img_path)
# Reshaping data
n, c, h, w = net.inputs[input_blob].shape
in_frame = cv.resize(frame, (w, h))
in_frame = in_frame.transpose((2, 0, 1)) # Change data layout from HWC to CHW
return in_frame.reshape((n, c, h, w)),frame
print('SUCCESS')
# -
# At this part, we are implementing a new inference method which is working with heterogeneous plugin.
#
# There are couple more steps we have added to inference process.
#
# - First we check if there is a CPU device on the list of devices, we load cpu extensions library.
#
# - Then, we load the network as usual process.
#
# - `set_config` function configures the plugin fallback devices and their order.
#
# Heterogeneous Plugin used with `HETERO:` prefix, which is followed with a list of devices, according to their prioritization.
#
# - After network load, we send the network to plugin to set affinity of layers according to priority list we give to plugin. When affinity sets the layer's target device to be executed.
#
# - Additionally we have performance counters or certain outputs we can later analyze the network. `.set_config({"HETERO_DUMP_GRAPH_DOT": "YES"})` prints .dot output which has a detailed graph representation of model.
#
# - Finally, `get_perf_counts` method prints the detailed execution times of layers on the devices.
#
# Let's run the next cell and make `runInference` method ready.
# ## 2 - Implement New `runInference` Method
#
# In this implementation we will do as defined in previous part. If you examine the code you will see the steps as following:
#
# - Load Plugin, set `device` parameter with 'HETERO:GPU,CPU`
#
# - Then parse plugin string to check if any extention is required.
#
# - Create network
#
# - Set Configurations:
# - `TARGET_FALLBACK`
#
# - `HETERO_DUMP_GRAPH_DOT` to output graph output.
#
# - Generate Executable Network and Run Inference
#
# - At the end of the script, if enabled prints `performance counters` values.
#
# #### Change Focus to Below Cell and Press (SHIFT + ENTER) or Click on Run Button
# +
# At this stage we implement our inference method to run with Heterogeneous plugin.
def runInference(hetero_device = 'HETERO:GPU,CPU',
model_xml='/home/intel/openvino_models/models/object_detection/common/mobilenet-ssd/FP32/mobilenet-ssd.xml',
model_bin='/home/intel/openvino_models/models/object_detection/common/mobilenet-ssd/FP32/mobilenet-ssd.bin',
image_file='images/car.png',
performance_counters = False,
dot_graph = True,
confidence_threshold=0.6):
# Plugin initialization for specified device. We will be targeting CPU initially.
plugin = IEPlugin(device=hetero_device, plugin_dirs='/opt/intel/openvino/deployment_tools/inference_engine/lib/intel64')
# Heterogeneous Plugin is provided as
#HETERO:GPU,CPU or similar.
vals = hetero_device.split(':')
targets = vals[1].split(',')
# Let's check if heterogeneous plugin has CPU target so we can add cpu extensions
# Loading additional exension libraries for the CPU
if hetero_device == 'CPU' or ('CPU' in targets):
extension_list=['/home/intel/inference_engine_samples_build/intel64/Release/lib/libcpu_extension.so']
for extension in extension_list:
plugin.add_cpu_extension(extension)
net = createNetwork(model_xml, model_bin, plugin)
# Let's check if the target is heterogeneous then we add fallback device priorities
#If we set TARGET_FALLBACK configuration, we want IE to decide what device to fallback.
plugin.set_initial_affinity(net)
plugin.set_config({"TARGET_FALLBACK": hetero_device})
if dot_graph:
# Just for a show case, remove
plugin.set_config({"HETERO_DUMP_GRAPH_DOT": "YES"})
exec_net, input_blob, out_blob = loadNetwork(plugin, net)
in_frame,original_frame = preprocessImage(image_file, net, input_blob)
my_request_id = 0
# Starting the inference in async mode, which starts the inference in parallel
inference_start = time.time()
exec_net.start_async(request_id=my_request_id, inputs={input_blob: in_frame})
# ... You can do additional processing or latency masking while we wait ...
# Blocking wait for a particular request_id
if exec_net.requests[my_request_id].wait(-1) == 0:
# getting the result of the network
res = exec_net.requests[my_request_id].outputs[out_blob]
inference_end = time.time()
# Processing the output result and adding labels on the image. Implementation is not shown in the
# this notebook; you can find it in object_detection_demo_ssd_async.py
initial_w = original_frame.shape[1]
initial_h = original_frame.shape[0]
frame = placeBoxes(res, None, confidence_threshold, original_frame, initial_w, initial_h, False, my_request_id, ((inference_end - inference_start)))
# We use pyplot because it plays nicer with Jupyter Notebooks
fig = plt.figure(dpi=300)
ax = fig.add_subplot(111)
ax.imshow(cv.cvtColor(frame, cv.COLOR_BGR2RGB), interpolation='none')
plt.axis("off")
plt.show()
else:
print("There was an error with the request")
if performance_counters:
perf_counts = exec_net.requests[0].get_perf_counts()
print("Performance counters:")
for layer, stats in perf_counts.items():
print(layer, ': ', stats)
return (plugin, net, exec_net)
print('SUCCESS')
# -
# In next cell, I would like to use Heterogeneous plugin to run object detection sample.
#
# With `HETERO:GPU,CPU,MYRIAD` we indicate to our plugin to prioritize `GPU > CPU > MYRIAD` for layers to be executed.
#
# For MobileNet-SSD example, almost all layers can run on GPU except `PriorBox` Caffe layer. Therefore, it will use GPU for all the `Convolution` layers and CPU for PriorBox layer.
#
# If, we have been used `HETERO:GPU,MYRIAD,CPU` , `PriorBox` layer would be running on MYRIAD since it has also support for it.
#
# Note that, we use FP16 because, in case CPU target is being used, IE helps to convert FP16 layers to FP32 automatically.
# ## 3 - Run Inference
#
# #### Change Focus to Below Cell and Press (SHIFT + ENTER) or Click on Run Button
# +
# Run code with : (SHIFT + ENTER) or Press Run Button #
hetero_objects = runInference(hetero_device = 'HETERO:GPU,CPU',
model_xml='/home/intel/openvino_models/models/object_detection/common/mobilenet-ssd/FP16/mobilenet-ssd.xml',
model_bin='/home/intel/openvino_models/models/object_detection/common/mobilenet-ssd/FP16/mobilenet-ssd.bin',
image_file='images/car.png',
performance_counters = False)
# -
# Running principles of Hetergeneous plugin is simple but if you want to take all the advantages of process and manually change the running layers and their target layers you should use layer affinity feature the change the execution target device.
# # Part 2: Layer Affinity
#
# Affinity by meaning likeness so you can image like a linkage of layer to target device.
#
# In this part we will look at how we can use this feature with Python API.
#
# This is a simple output of below code showing layer's affinity.
#
# ```
# Type: Input Device: CPU
# Type: ScaleShift Device: GPU
# Type: Convolution Device: GPU
# ```
# ## 1 - Print Layer Affinities of Previous Section.
#
# Below part reads IE network object and prints the network layer values.
#
# **Note**: Please make sure you already run the previous part.
#
# #### Change Focus to Below Cell and Press (SHIFT + ENTER) or Click on Run Button
# +
net = hetero_objects[1]
for l in net.layers.values():
print('Type: ', l.type, 'Device: ', l.affinity)
# -
# `{"HETERO_DUMP_GRAPH_DOT": "YES"}` configuraion let's us to print the network visualisation in .dot graph format. You can navigate to current directory and run it as below from a new terminal.
#
#
# ```bash
# # !xdot hetero_affinity_MobileNet-SSD.dot
# ```
#
# 
# ## 2 - Change Priority in Heterogeneous Plugin
#
# Let's use CPU First
#
# #### Change Focus to Below Cell and Press (SHIFT + ENTER) or Click on Run Button
hetero_objects = runInference(hetero_device = 'HETERO:CPU,GPU',
model_xml='/home/intel/openvino_models/models/object_detection/common/mobilenet-ssd/FP16/mobilenet-ssd.xml',
model_bin='/home/intel/openvino_models/models/object_detection/common/mobilenet-ssd/FP16/mobilenet-ssd.bin',
image_file='images/car.png',
performance_counters = False)
# ## 3 - Let's Check Layer Affinities
#
# You will notice that all layers assigned to CPU now.
#
# #### Change Focus to Below Cell and Press (SHIFT + ENTER) or Click on Run Button
# +
net = hetero_objects[1]
for l in net.layers.values():
print('Type: ', l.type, 'Device: ', l.affinity)
# -
# ## 4 - Let's Try with Myriad
#
#
# #### Change Focus to Below Cell and Press (SHIFT + ENTER) or Click on Run Button
# +
hetero_objects = runInference(hetero_device = 'HETERO:MYRIAD,GPU,CPU',
model_xml='/home/intel/openvino_models/models/object_detection/common/mobilenet-ssd/FP16/mobilenet-ssd.xml',
model_bin='/home/intel/openvino_models/models/object_detection/common/mobilenet-ssd/FP16/mobilenet-ssd.bin',
image_file='images/car.png',
performance_counters = False)
net = hetero_objects[1]
for l in net.layers.values():
print('Type: ', l.type, 'Device: ', l.affinity)
# -
# # Part 3: Performance Counters
#
# Inference Engine implements additional feature to let you know detailed performance report with performance counters. You can access them using Executable Network.
#
# Below code will report the layer name and its execution information.
#
# **NOTE**: Layet Affinity is only be used with Heterogeneous plugin since other plugins already determines the layer's execution target devices, performance counters can be used with all plugins.
#
# #### Change Focus to Below Cell and Press (SHIFT + ENTER) or Click on Run Button
# +
# Get the exec net
exec_net = hetero_objects[2]
# Print the performance counteres
perf_counts = exec_net.requests[0].get_perf_counts()
print("Performance counters:")
for layer, stats in perf_counts.items():
print(layer, ': ', stats)
# -
hetero_objects = runInference(hetero_device = 'HETERO:CPU,GPU',
model_xml='/home/intel/openvino_models/models/object_detection/common/mobilenet-ssd/FP16/mobilenet-ssd.xml',
model_bin='/home/intel/openvino_models/models/object_detection/common/mobilenet-ssd/FP16/mobilenet-ssd.bin',
image_file='images/car.png',
performance_counters = False)
# +
# Get the exec net
exec_net = hetero_objects[2]
# Print the performance counteres
perf_counts = exec_net.requests[0].get_perf_counts()
print("Performance counters:")
for layer, stats in perf_counts.items():
print(layer, ': ', stats)
# -
# # Part 4 - Custom Layers
#
# **NOTE**: This is a more advanced use case of OpenVINO(TM) Toolkit you may skip this part if you didn't cover the previous sections.
#
# This is a replication of https://github.com/intel-iot-devkit/smart-video-workshop/tree/master/custom-layer workshop.
#
# ### Custom Layers
#
# Custom layers are NN (Neural Network) layers that are not explicitly supported by a given framework. This tutorial demonstrates how to run inference on topologies featuring custom layers allowing you to plug in your own implementation for existing or completely new layers.
#
# The list of known layers is different for any particular framework. To see the layers supported by the Intel® Distribution of OpenVINO™ toolkit, refer to the Documentation: https://docs.openvinotoolkit.org/latest/_docs_MO_DG_Deep_Learning_Model_Optimizer_DevGuide.html#intermediate-representation-notation-catalog
#
#
# If your topology contains layers that are not in the list of known layers, the Model Optimizer considers them to be custom.
#
# The Model Optimizer searches for each layer of the input model in the list of known layers before building the model's internal representation, optimizing the model and producing the Intermediate Representation.
#
# Custom Layers implementation workflow in the Intel® Distribution of OpenVINO™ toolkit
# When implementing the custom layer in the Intel® Distribution of OpenVINO™ toolkit for your pre-trained model, you will need to add extensions in both the Model Optimizer and the Inference Engine. The following figure shows the work flow for the custom layer implementation.
#
# 
# ### Example custom layer: Hyperbolic Cosine (cosh) function
# We showcase custom layer implementation using a simple function; hyperbolic cosine (cosh). It's mathematically represented as:
#
# 
# ### Extension Generator
# This tool generates extension source files with stubs for the core functions. To get the workable extension, you will add your implementation of these functions to the generated files.
# ## 1 - Install Dependencies
#
# We need to execute all the commands through terminal so before going further, let's open a new terminal `CTRL + ALT + T`
#
# Let's first install the dependencies, copy the below code to terminal to setup `cogapp` package
#
# ```bash
# sudo pip3 install cogapp
# ```
# ## 2 - Create TF Model
#
# Let's navigate to source folder where the resources are placed.
#
# ```bash
# # cd /home/intel/smart-video-workshop/custom-layer/create_tf_model
# ```
#
# Then, let's build the model.
#
# We create a simple model with a custom cosh layer. The weights are random and untrained, however sufficient for demonstrating Custom Layer conversion.
#
# ```bash
# # mkdir -p tf_model
#
# chmod +x build_cosh_model.py
#
# ./build_cosh_model.py tf_model
# ```
#
# You will see that model file is generated into `tf_model` folder.
#
# ```
# Model saved in path: tf_model/model.ckpt
# ```
# ## 3 - Generate template files using the Extension Generator:
# We're using `/home/intel/smart-video-workshop/custom-layer/extgen_output/` as the target extension path:
#
# This will create templates that will be partially replaced by Python* and C++ code for executing the layer.
#
# ```bash
# # mkdir -p /home/intel/smart-video-workshop/custom-layer/extgen_output/
#
# python3 /opt/intel/openvino/deployment_tools/tools/extension_generator/extgen.py new --mo-tf-ext --mo-op --ie-cpu-ext --ie-gpu-ext --output_dir=/home/intel/smart-video-workshop/custom-layer/extgen_output/
#
# ```
#
# Answer the Model Optimizer extension generator questions as follows:
#
# ```bash
# Please enter layer name:
# [Cosh]
#
# Do you want to automatically parse all parameters from model file...
# [n]
#
# Please enter all parameters in format...
# When you finish please enter 'q'
# [q]
#
# Do you want to change any answer (y/n) ?
# [n]
#
# Please enter operation name:
# [Cosh]
#
# Please input all attributes that should be output in IR...
# ...
# When you finish enter 'q'
# [q]
#
# Please input all internal attributes for your operation...
# ...
# When you finish enter 'q'
# [q]
#
# Does your operation change shape? (y/n)
# [n]
#
# Do you want to change any answer (y/n) ?
# [n]
#
# Please enter operation name:
# [Cosh]
#
# Please enter all parameters in format...
# ...
# When you finish please enter 'q'
# [q]
#
# Do you want to change any answer (y/n) ?
# [n]
#
# Please enter operation name:
# [Cosh]
#
# Please enter all parameters in format...
# ...
# When you finish please enter 'q'
# [q]
#
# Do you want to change any answer (y/n) ?
# [n]
#
# ```
#
# Output will be like below:
#
# ```bash
# Stub file for TensorFlow Model Optimizer extractor is in /home/intel/smart-video-workshop/custom-layer/extgen_output/user_mo_extensions/front/tf folder
# Stub file for Model Optimizer operation is in /home/intel/smart-video-workshop/custom-layer/extgen_output/./user_mo_extensions/ops folder
# Stub files for Inference Engine CPU extension are in /home/intel/smart-video-workshop/custom-layer/extgen_output/./user_ie_extensions/cpu folder
# Stub files for Inference Engine GPU extension are in /home/intel/smart-video-workshop/custom-layer/extgen_output/./user_ie_extensions/gpu folder
# ```
# ## 4 - Register custom layer for the Model Optimizer
#
# Add Custom (cosh) Python Layers: Copy to the Model Optimizer Ops Directory:
#
# This allows the Model Optimizer to find the Python implementation of cosh.
#
# ```bash
# sudo cp /home/intel/smart-video-workshop/custom-layer/cosh.py /opt/intel/openvino/deployment_tools/model_optimizer/mo/ops/
# ```
# ## 5 - Generate IR with custom layer using Model Optimizer
# We run the Model Optimizer for TensorFlow to convert and optimize the new model for the Intel® Distribution of OpenVINO™ toolkit. We explicitly set the batch to 1 because the model has an input dim of "-1". TensorFlow allows "-1" as a variable indicating "to be filled in later", but the Model Optimizer requires explicit information for the optimization process. The output is the full name of the final output layer.
#
# ```bash
# # cd tf_model
#
# # mkdir -p /home/intel/smart-video-workshop/custom-layer/cl_ext_cosh
#
# python3 /opt/intel/openvino/deployment_tools/model_optimizer/mo_tf.py --input_meta_graph model.ckpt.meta --batch 1 --output "ModCosh/Activation_8/softmax_output" --extensions /home/intel/smart-video-workshop/custom-layer/extgen_output/user_mo_extensions --output_dir /home/intel/smart-video-workshop/custom-layer/create_tf_model/tf_model
# ```
# ## 6 - Inference Engine custom layer implementation for the Intel® CPU
#
# ### Copy CPU and GPU source code to the Model Optimizer extensions directory:
#
# This will be used for building a back-end library for applications that implement cosh.
#
# ```bash
# # cp /home/intel/smart-video-workshop/custom-layer/ext_cosh.cpp /home/intel/smart-video-workshop/custom-layer/extgen_output/user_ie_extensions/cpu/
# ```
#
# ### Compile the C++ extension library:
# Here we're building the back-end C++ library to be used by the Inference Engine for executing the cosh layer.
#
# **NOTE**: Make sure you changed line in CMakeLists.txt
#
# ***
# /opt/intel/openvino_2019.1.094/deployment_tools/inference_engine/external/tbb/include
# ***
#
# to
#
# ***
# /opt/intel/openvino/deployment_tools/inference_engine/external/tbb/include
# ***
#
# ```bash
# # cd /home/intel/smart-video-workshop/custom-layer/extgen_output/user_ie_extensions/cpu
#
# # cp /home/intel/smart-video-workshop/custom-layer/CMakeLists.txt .
#
# # mkdir -p build && cd build
#
# cmake ..
#
# make -j$(nproc)
#
# # cp libcosh_cpu_extension.so /home/intel/smart-video-workshop/custom-layer/cl_ext_cosh
# ```
# ## 7 - Run Sample Code
#
# #### Using a C++ Sample:
#
# ```bash
# /home/intel/inference_engine_samples_build/intel64/Release/classification_sample -i /opt/intel/openvino/deployment_tools/demo/car.png -m /home/intel/smart-video-workshop/custom-layer/create_tf_model/tf_model/model.ckpt.xml -d CPU -l /home/intel/smart-video-workshop/custom-layer/cl_ext_cosh/libcosh_cpu_extension.so
# ```
#
# #### Using a Python Sample:
#
# Prep: Install the OpenCV library and copy an appropriate sample to your home directory for ease of use:
#
# Try running the Python Sample without including the cosh extension library.
#
# You should see the error describing unsupported Cosh operation.
#
# ```bash
# python3 /opt/intel/openvino/deployment_tools/inference_engine/samples/python_samples/classification_sample/classification_sample.py -i /opt/intel/openvino/deployment_tools/demo/car.png -m /home/intel/smart-video-workshop/custom-layer/create_tf_model/tf_model/model.ckpt.xml -d CPU
# ```
#
# Error Output:
# ***
# ```
# [ INFO ] Loading network files:
# /home/intel/smart-video-workshop/custom-layer/create_tf_model/tf_model/model.ckpt.xml
# /home/intel/smart-video-workshop/custom-layer/create_tf_model/tf_model/model.ckpt.bin
# [ ERROR ] Following layers are not supported by the plugin for specified device CPU:
# ModCosh/cosh/Cosh, ModCosh/cosh_1/Cosh, ModCosh/cosh_2/Cosh
# [ ERROR ] Please try to specify cpu extensions library path in sample's command line parameters using -l or --cpu_extension command line argument
# ```
# ***
#
# - Now run the command with the cosh extension library:
#
# ```
# python3 /opt/intel/openvino/deployment_tools/inference_engine/samples/python_samples/classification_sample/classification_sample.py -i /opt/intel/openvino/deployment_tools/demo/car.png -m /home/intel/smart-video-workshop/custom-layer/create_tf_model/tf_model/model.ckpt.xml -l /home/intel/smart-video-workshop/custom-layer/cl_ext_cosh/libcosh_cpu_extension.so -d CPU
# ```
| Advanced Features - Lab 3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _uuid="e6762caaf637987d3f2579619c4614613e15eea7"
# # Overview
# + [markdown] _uuid="1fa3354f5ee23651516374c4f94f7c91c1c93ceb"
# ### Source:
# + [markdown] _uuid="9ffa26cd951a015a4bb5fc2c8446143ec60c0791"
# The data set was contributed to the benchmark collection by <NAME>, now at the Salk Institute and the University of California at San Deigo. The data set was developed in collaboration with <NAME> of Allied-Signal Aerospace Technology Center.
# + [markdown] _uuid="54004cf55686364a5d8c6df4178248c95cc47373"
# ### Data Set Information:
# + [markdown] _uuid="98c9c9bc1322b488c2feafc411ec799b967d5e5d"
# The file "sonar.mines" contains 111 patterns obtained by bouncing sonar signals off a metal cylinder at various angles and under various conditions. The file "sonar.rocks" contains 97 patterns obtained from rocks under similar conditions. The transmitted sonar signal is a frequency-modulated chirp, rising in frequency. The data set contains signals obtained from a variety of different aspect angles, spanning 90 degrees for the cylinder and 180 degrees for the rock.
#
# Each pattern is a set of 60 numbers in the range 0.0 to 1.0. Each number represents the energy within a particular frequency band, integrated over a certain period of time. The integration aperture for higher frequencies occur later in time, since these frequencies are transmitted later during the chirp.
#
# The label associated with each record contains the letter "R" if the object is a rock and "M" if it is a mine (metal cylinder). The numbers in the labels are in increasing order of aspect angle, but they do not encode the angle directly.
# + [markdown] _uuid="730982b69911757be257adc47d9ea4f35525f150"
# # Let's Work
# + _uuid="fb5718ddac4afecf1f50dad3fe5b1df8ad3261f9"
import numpy as np
import pandas as pd
import tensorflow as tf
import matplotlib.pyplot as plt
from sklearn.utils import shuffle
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
# + _uuid="055afd2a314bf560551026679beea96171b9cdfd"
# Reading the dataset
data = pd.read_csv('../input/sonar.all-data.csv')
# + _uuid="a641b43852e3c33743d7e77f829a19f9c6cbe112"
data.info()
# + _uuid="ec675b9c83aa11ca370954b0ce2843294459cbd7"
data.head()
# + _uuid="743e1fd2ad1e2f42a18c4bfc8148b7cabc82c8af"
print("Total Columns : ", len(data.columns))
# + _uuid="d5f1b3fe0937f1a1fe5eb4ec0f82b492bdb73a38"
# Total Labels
data[data.columns[60]].value_counts()
# + [markdown] _uuid="5dfa749d1c89ac19425c112017dcc0b4b07e82b1"
# ## Input and Output Data
# + _uuid="10f202bae631d576ed5a00d2f67c2cf7d2b5468e"
X = data[data.columns[0:60]].values
y = data[data.columns[60]].values
# + _uuid="61348f5edd37af2d7c00dc2975d4feb1f11680dd"
print(X.shape)
# + [markdown] _uuid="3db6944905099158a98de576045974035909ee9e"
# ## Using OneHotEncoder
# + _uuid="6a424d9bc94030678d81bbcf407966e853c13610"
def one_hot_encode(labels):
n_labels = len(labels)
n_unique_labels = len(np.unique(labels))
one_hot_encode = np.zeros((n_labels,n_unique_labels))
one_hot_encode[np.arange(n_labels), labels] = 1
return one_hot_encode
# + _uuid="2708d86d288d7f43bebf232f66cf4f8ab25a9d6d"
encoder = LabelEncoder()
encoder.fit(y)
y = encoder.transform(y)
Y = one_hot_encode(y)
# + _uuid="754adc8a675eaaf8aa58cabbf964a47c9380ad03"
encoder = LabelEncoder()
encoder.fit(y)
y = encoder.transform(y)
Y = one_hot_encode(y)
# + _uuid="c669eb185639c4806901fab4117b0dedf8255b3f"
Y[0]
# + [markdown] _uuid="a2a585eda4ec77d051f0dace36527ca8e44873fd"
# ## Training and Testing Data
# + [markdown] _uuid="ca66249b0a8cb1a9044cc1f095a6bdd9d0e7284b"
# As our output column has labels arranged together, so we first shuffle our data
# + _uuid="40f9063fa501cb6843a778b3271f989701d3d85b"
X,Y = shuffle (X, Y, random_state = 0)
# + _uuid="2c1218eab7e14994547d337e983b353c172732cf"
train_x, test_x, train_y, test_y = train_test_split(X, Y, test_size = 0.20, random_state = 0)
# + _uuid="28fbb261521299ced4eeb7d19aac231021254fe5"
print(train_x.shape)
print(train_y.shape)
print(test_x.shape)
# + [markdown] _uuid="ffbc717690308e03828a27fcca6fe3d4060a6425"
# # Neural Net Model
# + _uuid="431207ef89f54c994747f235509ede70d2468333"
learning_rate = 0.2
training_epochs = 500
# + _uuid="1fb8a1b65aa57fd498ff29ff678bfaebb5972fd4"
n_dim = X.shape[1]
print("n_dim = ", n_dim)
n_class = 2
# + _uuid="f4c51a86c00fa6dca8e83b6a6e48e82cdf7e18e4"
cost_history = np.empty(shape=[1],dtype=float)
# + [markdown] _uuid="535f43907edba899e77c384eebcc1228837ba94a"
# ## 4 layer Dense Neural Networks
# + _uuid="30399ae283fce05843a50403dd8d019a7acb280c"
n_hidden_1 = 60
n_hidden_2 = 60
n_hidden_3 = 60
n_hidden_4 = 60
# + _uuid="dd5cfc25d7025c329b024fa8bbb8eda133ef76d6"
x = tf.placeholder(tf.float32, [None, n_dim])
W = tf.Variable(tf.zeros([n_dim, n_class]))
b = tf.Variable(tf.zeros([n_class]))
y_ = tf.placeholder(tf.float32, [None, n_class])
# + _uuid="028a956d95ebc7a4b75b5e22ba69027be1684a39"
# Define the model
def multilayer_perceptron(x, weights, biases):
# Hidden layer with RELU activationsd
layer_1 = tf.add(tf.matmul(x, weights['h1']), biases['b1'])
layer_1 = tf.nn.relu(layer_1)
# Hidden layer with sigmoid activation
layer_2 = tf.add(tf.matmul(layer_1, weights['h2']), biases['b2'])
layer_2 = tf.nn.relu(layer_2)
# Hidden layer with sigmoid activation
layer_3 = tf.add(tf.matmul(layer_2, weights['h3']), biases['b3'])
layer_3 = tf.nn.relu(layer_3)
# Hidden layer with RELU activation
layer_4 = tf.add(tf.matmul(layer_3, weights['h4']), biases['b4'])
layer_4 = tf.nn.sigmoid(layer_4)
# Output layer with linear activation
out_layer = tf.matmul(layer_4, weights['out']) + biases['out']
return out_layer
# + _uuid="e4cd92bbfbe5744ca4202cffd5bd0c315697a8ae"
# Define the weights for each layers
weights = {
'h1': tf.Variable(tf.truncated_normal([n_dim, n_hidden_1])),
'h2': tf.Variable(tf.truncated_normal([n_hidden_1, n_hidden_2])),
'h3': tf.Variable(tf.truncated_normal([n_hidden_2, n_hidden_3])),
'h4': tf.Variable(tf.truncated_normal([n_hidden_3, n_hidden_4])),
'out': tf.Variable(tf.truncated_normal([n_hidden_4, n_class]))
}
# + _uuid="07c8bcdca73d935812ebde69ac9dd26d7ac07e46"
# Define the bias for each layers
biases = {
'b1': tf.Variable(tf.truncated_normal([n_hidden_1])),
'b2': tf.Variable(tf.truncated_normal([n_hidden_2])),
'b3': tf.Variable(tf.truncated_normal([n_hidden_3])),
'b4': tf.Variable(tf.truncated_normal([n_hidden_4])),
'out': tf.Variable(tf.truncated_normal([n_class]))
}
# + _uuid="cba1956b103c78a18ab92682b50388304732f3aa"
init = tf.global_variables_initializer()
# + _uuid="e8c69dacb69373569cd2fb6645e6a3acceeb3a27"
# Calling model
y = multilayer_perceptron(x, weights, biases)
# + [markdown] _uuid="3615d7746463e7788e0b35b419243fcad4f32f7a"
# ## Cost Function
# + _uuid="9868feb339d262ce11def8049f64e8284ec18851"
cost_function = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=y, labels=y_))
training_step = tf.train.GradientDescentOptimizer(learning_rate).minimize(cost_function)
# + [markdown] _uuid="0d81298a77d91dc40cd181d5ca2ed417bb9eafb3"
# ## Initializing Session
# + _uuid="e835863bb77585b4402ae0d4cfeb5ebcafeb7733"
sess = tf.Session()
sess.run(init)
# + _uuid="c95cf9403f05d190644e919f4f8826e15b9234af"
mse_history = []
accuracy_history = []
# + [markdown] _uuid="74b5235de56df19131fedf0d9e81e5c97ff3b3c8"
# ## Training Model
# + _uuid="18da48c40a479e4e10d419635ca95bcdd1c9d256"
for epoch in range(training_epochs):
sess.run(training_step, feed_dict = {x: train_x, y_: train_y})
cost = sess.run(cost_function, feed_dict={x: train_x, y_: train_y})
cost_history = np.append(cost_history, cost)
correct_prediction = tf.equal(tf.argmax(y, 1), tf.argmax(y_, 1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
pred_y = sess.run(y, feed_dict = {x: test_x})
mse = tf.reduce_mean(tf.square(pred_y - test_y))
mse_ = sess.run(mse)
mse_history.append(mse_)
accuracy = (sess.run(accuracy, feed_dict={x: train_x, y_: train_y}))
accuracy_history.append(accuracy)
if epoch % 50 == 0:
print('epoch : ', epoch, ' ; ', 'cost: ', cost, " ; MSE: ", mse_, "- Train Accuracy: ", accuracy )
# + [markdown] _uuid="fd88d915b19295d9473c558a153854ff14b535c3"
# ## Cost Graph
# + _uuid="baeed0f965b6305229d19dd1d33b5868546d5e63"
plt.plot(range(len(cost_history)), cost_history)
plt.xlabel('Epochs ')
plt.ylabel('Cost_History ')
plt.title("Cost per Epoch Graph")
plt.xlim(0, training_epochs + 10)
plt.ylim(0, np.max(cost_history))
plt.rcParams['figure.figsize'] = [12, 5]
plt.show()
# + [markdown] _uuid="e9346a4944350612a3b5024cde022664ebbe924a"
# # Accuracy Graph
# + _uuid="c1b2558c6399a96024e81433e189958e3d096220"
plt.plot(range(len(accuracy_history)), accuracy_history)
plt.xlabel('Epochs ')
plt.ylabel('Accuracy_History ')
plt.title("Cost per Epoch Graph")
plt.xlim(0, training_epochs + 10)
plt.ylim(0.4, np.max(accuracy_history)+0.2)
plt.rcParams['figure.figsize'] = [12, 5]
plt.show()
# + [markdown] _uuid="30f290fdb3dfded6064b22eaa2b8faa349fa2c53"
# ### Accuracy
# + _uuid="e4ab3a56af8547b673083d2f3b7fcdd4faf5d528"
correct_prediction = tf.equal(tf.argmax(y, 1), tf.argmax(y_, 1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
print("Test Accuracy: ", (sess.run(accuracy, feed_dict={x: test_x, y_: test_y})))
# + [markdown] _uuid="4dc032f179594ed509a2653bef45e0d08f8d1cff"
# ### Mean Square Error
# + _uuid="ee9ea1a1eb66df918bc2ad91a08fa5baa0c5b88a"
pred_y = sess.run(y, feed_dict={x: test_x})
mse = tf.reduce_mean(tf.square(pred_y - test_y))
print("MSE: %.4f" % sess.run(mse))
# + [markdown] _uuid="023c9a9a91bddc76b10f68c8fc703713c3efd7a1"
# .
# + [markdown] _uuid="5bc84c0dc209674b79d485cea3e5a1cc986b9cf2"
# Thanks for having a look. If u like the Deep Neural Network approach, Pls give a LIKE to my kernel :)
| courses/udacity_deep_learning/4-layer-dense-neural-net-using-tensorflow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/qinyufm/deeplearning-models/blob/master/LSTM.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="aRDUGd13kUFu"
# # LSTM
#
# + id="-uTOYW1P0fR3" colab_type="code" outputId="d7168878-266b-47d8-f20a-e5701fa16b8e" colab={"base_uri": "https://localhost:8080/", "height": 81}
import math
import pandas_datareader as web
import numpy as np
import pandas as pd
from sklearn.preprocessing import MinMaxScaler
from keras.models import Sequential
from keras.layers import Dense, LSTM
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
# + [markdown] colab_type="text" id="LhDiAMaaAnQF"
# ## Get the data
# + id="Yb7LgH6N04pS" colab_type="code" outputId="38acd0b3-24be-4af1-db16-a207f43c118e" colab={"base_uri": "https://localhost:8080/", "height": 431}
df = web.DataReader('MRK', data_source='yahoo', start='2018-01-01', end='2020-02-11')
#Show the data
df
# + id="Lus8P7pk1LZB" colab_type="code" outputId="b84b9933-a4a8-454b-bf70-2c36e803151f" colab={"base_uri": "https://localhost:8080/", "height": 684}
#Visualize the closing price history
plt.figure(figsize=(16,8))
plt.title('Close Price History')
plt.plot(df['Close'])
plt.xlabel('Date', fontsize=18)
plt.ylabel('Close Price USD ($)', fontsize=18)
plt.show()
# + [markdown] id="i6XCsV7J6j8i" colab_type="text"
# ## Filter the data by close
# + id="CGSe5NYG1RVf" colab_type="code" outputId="9f955feb-c59f-40a7-df95-2ee1180bcd25" colab={"base_uri": "https://localhost:8080/", "height": 34}
#Create a new dataframe with only the 'Close column
data = df.filter(['Close'])
#Convert the dataframe to a numpy array
dataset = data.values
#Get the number of rows to train the model on
training_data_len = math.ceil( len(dataset) * .8 )
training_data_len
# + [markdown] id="lCj8E53H6uUj" colab_type="text"
# ### Scale the data
# + id="E-f060Qw1hTE" colab_type="code" colab={}
#Scale the data
scaler = MinMaxScaler(feature_range=(0,1))
scaled_data = scaler.fit_transform(dataset)
# + [markdown] id="G-QLWY0k6yIy" colab_type="text"
# ### Prepare training
# + id="l2_28Nq21kQY" colab_type="code" outputId="42622e6d-c2d3-4740-b001-de05fd0476d5" colab={"base_uri": "https://localhost:8080/", "height": 703}
#Create the training data set
#Create the scaled training data set
train_data = scaled_data[0:training_data_len , :]
#Split the data into x_train and y_train data sets
x_train = []
y_train = []
for i in range(60, len(train_data)):
x_train.append(train_data[i-60:i, 0])
y_train.append(train_data[i, 0])
if i<= 61:
print(x_train)
print(y_train)
#print()
# + [markdown] id="ly1z-l9l68wo" colab_type="text"
# #### convert to numpy array
# + id="9d0rSDKg1kUe" colab_type="code" outputId="b2d47de0-6a49-4894-f7b4-b292e80659d8" colab={"base_uri": "https://localhost:8080/", "height": 34}
#Convert the x_train and y_train to numpy arrays
x_train, y_train = np.array(x_train), np.array(y_train)
#Reshape the data
x_train = np.reshape(x_train, (x_train.shape[0], x_train.shape[1], 1))
x_train.shape
# + [markdown] id="Wc-HiY3P7BV6" colab_type="text"
# ## Build the model
# + id="Fmt5GLPE1uzE" colab_type="code" outputId="77db655b-e5ee-4337-95ef-4b7e794bc034" colab={"base_uri": "https://localhost:8080/", "height": 158}
#Build the LSTM model
model = Sequential()
model.add(LSTM(50, return_sequences=True, input_shape= (x_train.shape[1], 1)))
model.add(LSTM(50, return_sequences= False))
model.add(Dense(25))
model.add(Dense(1))
#Compile the model
model.compile(optimizer='adam', loss='mean_squared_error')
# + [markdown] id="yhlrfacD7Gph" colab_type="text"
# ### Train the model
# + id="f79vMxr611jG" colab_type="code" outputId="34331d96-30ea-4ede-c33f-180ca35223b1" colab={"base_uri": "https://localhost:8080/", "height": 386}
#Train the model
model.fit(x_train, y_train, batch_size=1, epochs=1)
#Create the testing data set
#Create a new array containing scaled values from index 1543 to 2002
test_data = scaled_data[training_data_len - 60: , :]
#Create the data sets x_test and y_test
x_test = []
y_test = dataset[training_data_len:, :]
for i in range(60, len(test_data)):
x_test.append(test_data[i-60:i, 0])
#Convert the data to a numpy array
x_test = np.array(x_test)
#Reshape the data
x_test = np.reshape(x_test, (x_test.shape[0], x_test.shape[1], 1 ))
# + [markdown] id="uDt_FmML7J26" colab_type="text"
# ### Prediction and RMSE
# + id="OdBcYarP15nm" colab_type="code" outputId="3c54b1bd-8723-452d-d908-54318f8f2381" colab={"base_uri": "https://localhost:8080/", "height": 34}
#Get the models predicted price values
predictions = model.predict(x_test)
predictions = scaler.inverse_transform(predictions)
#Get the root mean squared error (RMSE)
rmse=np.sqrt(np.mean(((predictions- y_test)**2)))
rmse
# + [markdown] id="gFiOkI1P7PCl" colab_type="text"
# ### Visualization
# + id="_nl8Jxb21-Po" colab_type="code" outputId="2cfbbb1c-4bcf-4831-b86a-4de084390398" colab={"base_uri": "https://localhost:8080/", "height": 1000}
#Plot the data
train = data[:training_data_len]
valid = data[training_data_len:]
valid['Predictions'] = predictions
#Visualize the data
plt.figure(figsize=(16,8))
plt.title('Model')
plt.xlabel('Date', fontsize=18)
plt.ylabel('Close Price USD ($)', fontsize=18)
plt.plot(train['Close'])
plt.plot(valid[['Close', 'Predictions']])
plt.legend(['Train', 'Val', 'Predictions'], loc='lower right')
plt.show()
#Show the valid and predicted prices
valid
# + id="SNtoebiz2IxX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 721} outputId="03f59835-d6c0-4458-b1f0-2426f150b988"
#Get the quote
apple_quote = web.DataReader('TSN', data_source='yahoo', start='2018-01-01', end='2020-02-11')
#Create a new dataframe
new_df = apple_quote.filter(['Close'])
#Get teh last 60 day closing price values and convert the dataframe to an array
last_60_days = new_df[-60:].values
#Scale the data to be values between 0 and 1
last_60_days_scaled = scaler.transform(last_60_days)
#Create an empty list
X_test = []
#Append teh past 60 days
X_test.append(last_60_days_scaled)
#Convert the X_test data set to a numpy array
X_test = np.array(X_test)
#Reshape the data
X_test = np.reshape(X_test, (X_test.shape[0], X_test.shape[1], 1))
#Get the predicted scaled price
pred_price = model.predict(X_test)
#undo the scaling
pred_price = scaler.inverse_transform(pred_price)
print(pred_price)
#Get the quote
apple_quote2 = web.DataReader('TSN', data_source='yahoo', start='2019-12-18', end='2020-02-11')
print(apple_quote2['Close'])
# + [markdown] id="X7ot1FeG7kO_" colab_type="text"
# # All together prediction
# + id="OqlrrmQm7qVV" colab_type="code" colab={}
df = web.DataReader('MRK', data_source='yahoo', start='2018-01-01', end='2020-02-11')
| LSTM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/thlystrd/LinearAlgebra_2ndsem/blob/main/Activity_1_Python_Fundamentals.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="DjROMxalGij2"
# # Welcome to Python Fundamentals
# In this module, we are going to establish or review our skills in Python programming. In this notebook we are going to cover:
# * Variables and Data Types
# * Operations
# * Input and Output Operations
# * Logic Control
# * Iterables
# * Functions
# + [markdown] id="Efqy4fb9rLQu"
# ## Variable and Data Types
# + [markdown] id="tK5liyY9uFWt"
# - codes used for identifying and calling out certain data types such as the following:
# - **Integer** - whole numbers.
# - **Float** - numbers that contain decimals.
# - **String** - charaters that are placed inside quotation marks.
# + id="KbCzjBh7ro3c" colab={"base_uri": "https://localhost:8080/"} outputId="a44f2691-88e1-4098-f44f-e8eee72b0984"
uwu = 11
aha,ahha = 3, -11
eme, emme, emme = 4, 5, 6
uwu
# + id="a-VnW9LXuFpz" colab={"base_uri": "https://localhost:8080/"} outputId="4a7d2b5f-e511-40f3-9cbd-6aad4405dc5a"
type(uwu)
# + id="gbyeF7IguZsX" colab={"base_uri": "https://localhost:8080/"} outputId="5e1bec15-5d6d-456c-8c71-1c47e2712da0"
ehe = 1.0
type(ehe)
# + id="vh2WkX9iuvxm" colab={"base_uri": "https://localhost:8080/"} outputId="4aaf395f-e408-46f4-f3e8-9bda4fa44bdb"
uwu = float(uwu)
type (uwu)
# + id="jU0HWIKgv5FI" colab={"base_uri": "https://localhost:8080/"} outputId="194d520b-dce6-4fde-96d2-4838feadaa5f"
k,a,r = 3 , 11.11 , 'twenty'
type (r)
# + id="EuWtlqW-wwvM" colab={"base_uri": "https://localhost:8080/"} outputId="85bbc15e-1caa-4192-efd4-230e9dd75910"
k_int = int(k)
k_int
# + [markdown] id="P5omo7qnxlAH"
# ## Operations
# + [markdown] id="kO3k4Hqnx2us"
# ###Arithmetic
# + [markdown] id="uv5_V4TkuXFQ"
# - assigning numbers (integer or float) to each variable and using these variables to perform the following basic operations:
# - **Addition** - wherein the sum of variables is equal to the sum of their values.
# - **Subtraction** - wherein the difference of variables is equal to the difference of their designated values.
# - **Multiplication** - wherein the product of variables is equal to the product of their designated values.
# - **Division** - wherein the quotient of variables is equal to the quotient of their designated values.
# - **Floor Division** - Division wherein the quotient containing a decimal point is the rounded up to the closest integer.
# - **Exponentiation** - Wherein a variable is exponentiated.
# - **Modulo** - Division operation wherein only the remainder is returned. [1]
# + id="UesuSc2cx78c"
a , b , c , d = -3 , -11 , 20 , 20
# + id="lMMOMDMZyR8U" colab={"base_uri": "https://localhost:8080/"} outputId="c3d3d0ee-a887-4e0a-f7ad-8273a89e69c2"
### Addition
S = a+b
S
# + id="ulomfj_MzKwl" colab={"base_uri": "https://localhost:8080/"} outputId="1b37e276-cf12-4e8c-9036-0c8784c6f45c"
### Subtraction
D = b - d
D
# + id="0_rvjPmYzeje" colab={"base_uri": "https://localhost:8080/"} outputId="783df498-b5d9-4252-8614-436e48cf3add"
### Multiplicaion
P = a * d
P
# + id="3Xtf2t-rzwn2" colab={"base_uri": "https://localhost:8080/"} outputId="48144c51-9d7a-455f-9fd3-ef0a5d1a65ad"
### Division
Q = a / c
Q
# + id="ltn1c6fT0hY0" colab={"base_uri": "https://localhost:8080/"} outputId="868cc1eb-ceac-4d0e-ed6f-b39830092675"
### Floor Division
Fq = d // c
Fq
# + id="r1FwIJCE1eq9" colab={"base_uri": "https://localhost:8080/"} outputId="b122e1c8-294f-464a-a1ac-d701a45a7066"
### Exponentation
E = a**c
E
# + id="u_mZ5iSZ2gx8" colab={"base_uri": "https://localhost:8080/"} outputId="7ee47556-402b-4f87-dffd-fcbcc6f6475b"
### Modulo
mod = a%c
mod
# + [markdown] id="LvTSSpZl3T0U"
# ### Assignment Operations
# + [markdown] id="2tw84Fquub_o"
# - assigning numbers (integer or float) to each variable and using these variables to perform the following operations:
# - **Increment or Addition Assignment Operator** - wherein each time the code runs the value of G increases by d.
# - **Decrement or Subtraction Assignment Operator** - wherein each time the code runs the value of H decreases by a.
# - **Multiplication Assignment Operator** - wherein each time the code runs the value of J would be multiplied to a value of 2.
# - **Exponent Assignment Operator** - wherein each time the code runs the value of K would be raised to a value of 3.
# + id="A8gXjv513eTX"
G , H , J , K = 6 , 12 , 20 , 2
# + id="ZDspYGS73ym0" colab={"base_uri": "https://localhost:8080/"} outputId="94ed4528-4cbe-4c4d-fcdc-9b56eae3d1f5"
G += a
G
# + id="TUGmxchv4nSJ" colab={"base_uri": "https://localhost:8080/"} outputId="ac8cdee7-5ceb-4bd1-fddb-9d46fadfe6c3"
H -= b
H
# + id="Cdo1Nc0z5Mcw" colab={"base_uri": "https://localhost:8080/"} outputId="1deaca66-34ae-4cf2-e788-eb46e91e715c"
J *= c
J
# + id="LIBxB40X5e07" colab={"base_uri": "https://localhost:8080/"} outputId="8e79c5c4-dcb9-49cb-ed52-06395a2ffe9b"
K **= d
K
# + [markdown] id="9NNI0SrE5vCO"
# ### Comparators
# + [markdown] id="bCbG0aZQugj_"
# - assigning and integer, float, or string to each variable and using these variables to perform the following comparison operators to determine if the condition would prove to be either true or false:
# - **Equality** - when two operands are equal then the condition is true, if not then false.
# - **Non-Equality** - when two operands are not equal then the condition is true, if not then false.
# - **Inequality**
# - *Greater than* - when the left operand is greater than the right then the condition is true, if not then false.
# - *Less than* - when the left operand is less than the right then the condition is true, if not then false.
# - *Greater than or Equal to* - when the left operand is greater than or equal to the right then the condition is true, if not then false.
# - *Less than or Equal to* - when the left operand is less than or equal to the right then the condition is true, if not then false.
# + id="HlsgxbKU6FB7"
res_1, res_2, res_3 = 3, 6, 19
true_val = 3.0
# + id="Sf2xemIe6l-d" colab={"base_uri": "https://localhost:8080/"} outputId="b0254edf-3b98-4dea-8426-9e9e9d76837a"
## Equality
res_2 == true_val
# + id="BZJRONgb7JTW" colab={"base_uri": "https://localhost:8080/"} outputId="01ba64cf-4e83-4a7e-c1cd-445d0dda2d84"
## Non-equality
res_2 != true_val
# + id="XC5Q6-lz7eZk" colab={"base_uri": "https://localhost:8080/"} outputId="bf672003-a62a-454d-9949-f3718af81924"
## Inequality
t1= res_1 > res_2
t2= res_1 < res_2/2
t3= res_1 >= res_2/2
t4= res_1 <= res_2
t3
# + [markdown] id="WdnR2J4o9IbK"
# ### Logical
# + [markdown] id="NN9bFBBBumrq"
# **Boolean value** - This is a built-in data type in Python that is mostly used in logical operations to represent an expressions truth value.
#
# - assigning and integer, float, string, or boolean value [2] to each variable and using these variables to perform the following logical operators:
# - **Equality** - used to compare two operands to check if they are equal. This would show true if both operands are equal and false if not.
# - **Is** - used to compare two operands to check if they are the same identity [3]. This would show true if both operands are the same and false if not.
# - **Conjunction** - uses "and" in between two operands. This would show true if both operands are true and false if not.
# - **Disjunction** - uses "or" in between two operands. This would show true if atleast one of the operands is true and false if none are true.
# - **Negation** - uses "not" at the beginning of an operand. This would show true if the operand is false and false if the operand is true.
# + id="Tfr_546_9M6G" colab={"base_uri": "https://localhost:8080/"} outputId="e227b695-30fb-41ca-ce1b-52444ff5ea60"
res_1 == true_val
# + colab={"base_uri": "https://localhost:8080/"} id="5o5HxrEnzbDn" outputId="89b8fc35-f8f7-42ae-9a1d-2374964315dd"
res_2 == true_val
# + id="n40cQf1kCBd_" colab={"base_uri": "https://localhost:8080/"} outputId="69777e43-0b05-4062-b6f4-ebd9b3b49068"
res_1 is true_val
# + id="DG2ej-FnCIla" colab={"base_uri": "https://localhost:8080/"} outputId="a5be592b-4f24-4c3b-905b-348297b6ada9"
res_1 is not true_val
# + id="_uqCma7XCWAq" colab={"base_uri": "https://localhost:8080/"} outputId="83132c14-22ff-425c-ed84-0a60bddcd5e6"
p,q = True, False
conj = p and q
conj
# + colab={"base_uri": "https://localhost:8080/"} id="Fm-7HbbG0284" outputId="2da9454b-feef-4cdc-a5b4-e476cfb441a5"
p,q = True, True
conj = p and q
conj
# + id="Uwj7FfyBCofC" colab={"base_uri": "https://localhost:8080/"} outputId="b01834b9-174a-441c-f38d-a1950eeb6943"
p,j = True, True
disj = p or q
disj
# + colab={"base_uri": "https://localhost:8080/"} id="eV4hAUji1a-q" outputId="ae1be4d3-59c9-4754-d324-965ea2d93fbd"
p,j = True, False
disj = p or q
disj
# + colab={"base_uri": "https://localhost:8080/"} id="AYlWgJtt1eYe" outputId="5ae6506b-0dc8-45bc-cb1a-16afb1f78164"
p,j = False, False
disj = p or q
disj
# + id="Em-oci37EpiM" colab={"base_uri": "https://localhost:8080/"} outputId="8b9d3d8d-6104-4a1e-fb40-d5b6b78a92a5"
p, q = True , False
e = not (q)
e
# + id="Uuvhi9pzFZ2S" colab={"base_uri": "https://localhost:8080/"} outputId="194a3308-dbd5-477d-af87-fb47ac80f7be"
p,q = True, False
xor = (not p and q) or (p and not q)
xor
# + [markdown] id="9JicNDeUHVbq"
# ### I/O
# + [markdown] id="smlhxN59uwcp"
# **I/O** - stands for Input/Output and it is informaly known as IO or io. I/O shows a string that serves as an output based on what you input. [4]
#
#
#
# > Commands or Declarations that are used under Input/Output are the following:
#
#
# > - **Print** - easiest way to produce an output whrein you can type a statement or different statements that are to be separated by a comma. Converts statements into strings and produces an output.
# - **Counter** - a declaration where you can set the starting number for the command "cnt". An increment will result to an increase to the set value of the counter.
# - **Input** - displays a prompt that will ask you to input a text or a value. Depending on your code, it can display the text or value right away or it can use your text or value for the string that will be printed as the output.
#
# + id="YKQac77cHcjy" colab={"base_uri": "https://localhost:8080/"} outputId="e3e25eac-029a-4410-fdfc-b3b9707165e1"
print ("Hello Master")
# + id="yA_YKw02Hqy6"
cnt=1
# + id="_dkLWiAmHwCN" colab={"base_uri": "https://localhost:8080/"} outputId="59efa67c-f3fd-4d3a-df2f-12c20e6211e9"
string = "Hello World"
print(string, ",Current run count is:" , cnt)
cnt += 1
# + id="8mBTbP1YJhyf" colab={"base_uri": "https://localhost:8080/"} outputId="3e3dacd8-fbf5-4a34-8700-344241bbc1e2"
print(f"{string}, Current count is: {cnt}")
# + id="s4yg9UKOLEVp" colab={"base_uri": "https://localhost:8080/"} outputId="99609f93-5fdc-4ad5-b015-d066acc518f3"
sem_grade = 95.3333333
name = "<NAME>"
print("Hello {} , your semestral grade is: {}".format(name, sem_grade))
# + id="pm-q-S0KXoB3" colab={"base_uri": "https://localhost:8080/"} outputId="cb4293d0-310b-4c46-e540-e4e2db1b2831"
w_pg, w_mg, w_fg = 0.3, 0.3, 0.4
print("The weights of your semestral grades are; \
\n\t{: .2%} for Prelims\
\n\t{: .2%} for Midterms, and\
\n\t{: .2%} for Finals.".format(w_pg, w_mg, w_fg))
# + id="mHP2x8xnkfV6" colab={"base_uri": "https://localhost:8080/", "height": 53} outputId="c8048674-aa9e-41c1-be1c-80600a9b2de6"
x = input("enter a number: ")
x
# + id="KX2gd4tnksVh" colab={"base_uri": "https://localhost:8080/"} outputId="25e1ab80-676c-44df-f56e-46888cb0fcc1"
name = input("Name: ")
pg = input("Enter prelim grade: ")
mg = input("Enter midterm grade: ")
fg = input("Enter finals grade: ")
sem_grade = 95
print("Hello {}, your semestral grade is: {}" .format(name, sem_grade))
# + [markdown] id="K-JfbSMvnXt1"
# ## Looping Statements
# + [markdown] id="NRrzSnKXngMC"
# ### While
# + [markdown] id="AhQOZ4_Ku154"
# **While Loop** - continually executes until it reaches the value that is given in the command. While loop can also be used in statements given that the condition is true and it will stop the loop once the condition is false. [5]
# + id="-zJddTNZnhzd" colab={"base_uri": "https://localhost:8080/"} outputId="b17bec26-da2a-4f4f-a002-23b694d94a2c"
## while loops
i, j = 12, 19
while(i<=j):
print(f"{i}\t|\t{j}")
i+=1
# + [markdown] id="z5cTAbTHpy_M"
# ### For
# + [markdown] id="WMItmYsju3dy"
# **For Loop** - executes a sequence of statement or value. It is frequently used to display a sequence of lists, sets, strings, or even songs. [5] When it comes to values, it will display up to what value is declared for the range of your for loop.
# + id="s7yXFK9EokES" colab={"base_uri": "https://localhost:8080/"} outputId="b59ff47c-bb16-4ddb-c4bf-7cbbf3558b75"
# for(int i=0; i<10; i++){
# print(i)
# } eto pwede pang Java and c++
i=0
for i in range(10):
print(i)
# + id="RqGIF9bap18t" colab={"base_uri": "https://localhost:8080/"} outputId="7a3fb327-1319-48b8-fb4b-9aa5dc8d1c8b"
playlist = ["We don't talk about Bruno", "Family Madrigal"]
print('Now playing: \n')
for song in playlist:
print(song)
# + [markdown] id="RWOtpVUJqocr"
# ## Flow Control
# + [markdown] id="YJWsZLtVqqxa"
# ### Condition Statements
# + [markdown] id="mML3c3GOu9be"
# **Condition Statements** - used for different decision-making situations. It's action depends whether a Boolean value [2] is true or false.
#
#
# > The three condition statements are as follows:
# - **If Statement** - executed once the given condition is met or true.
# - **Else If Statement** - executed once 'if statement' did not meet the given condition or is false.
# - **Else Statement** - executed once the condition did not meet neither 'if statement' or 'else if statement'. In other words, else statement will be executed once the two above are false.
#
#
#
#
# + id="CMelRgtTquEj" colab={"base_uri": "https://localhost:8080/"} outputId="f1f6765c-b99f-43b7-c818-9d369871a361"
numeral1, numeral2 = 11, 10+2
if(numeral1 == numeral2):
print("Yzel")
elif(numeral1>numeral2):
print("Oreo")
else:
print("Mocha")
# + [markdown] id="PB0Hi8fgrXMa"
# ## Functions
# + [markdown] id="7DvYnZh-vBjy"
# **Functions** - statement that performs a task. It lessens the job for you can use the same code infinitely just by using different inputs. This command only runs when it is called. With functions, you can input statement to be runned or use it for computation.
#
#
# > We can declare a function through:
# - **Define** - or "def" creates a function once it is followed by a parenthesis containing the statement or parameter of your choice. The function will then be called once you input the name of the function. [6]
#
#
# + id="z9XQuewnrY-a"
def delete_user (userid):
print("Succesfully deleted user:{}".format(userid))
def delete_all_users ():
print("Successfully deleted all users")
# + id="EOecfq10sXmy" colab={"base_uri": "https://localhost:8080/"} outputId="f20c6998-dedf-4723-f5e9-03878d9871c9"
userid = "Purple"
delete_user("Purple")
delete_all_users()
# + id="87xgsNSfsme8"
def add(addend1, addend2):
return addend1 + addend2
def power_of_base4(exponent):
return 4**exponent
# + id="ssnN925vtBGy" colab={"base_uri": "https://localhost:8080/"} outputId="ff02d2d9-257f-41f6-fd61-7049920f4022"
# addend1, addend2 = 34, 35
# add(addend1, addend2)
exponent = 2
power_of_base4(exponent)
# + [markdown] id="-aPeNYXScZpO"
# ##Grade Calculator
# + id="wPuWCSTiccco" colab={"base_uri": "https://localhost:8080/"} outputId="1070b44b-d01f-4f8b-cb27-a4b793b62871"
name = input("Name: ")
course = input("Course:")
prelim = float(input("Prelim Grade: "))
midterm = float(input("Midterm Grade: "))
finals = float(input("Finals Grade: "))
sem_grade = prelim*.30+midterm*.30+finals*.40
print("Hello {} of {}! Your Semestral Grade is {}. Congratulations!" .format(name, course, sem_grade))
if (sem_grade > 70.00) :
print("\U0001F600")
elif(sem_grade == 70.00) :
print("\U0001F606")
else:
print("\U0001F62D")
| Activity_1_Python_Fundamentals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''bulgarian-constitutional-court-decisions'':
# pipenv)'
# metadata:
# interpreter:
# hash: 9e35bad0b5aa7a87571eaa6bfdcd491c11984b99cb58639a3f82d97f7d8977e2
# name: python3
# ---
# # Baseline Models
#
# - Using three baseline models for text classification, training a model to identify political language in court documents. The three classification algorithms used are:
# 1. Logistic regression
# 2. Naive Bayes
# 3. Support Vector Classification
#
# ## Next steps
#
# - Data needs more sentences labelled as political, as data is imbalanced and models observe few political sentences
# - Need to optimize hyperparameters to improve model performance
# +
import glob
import os
import pickle
import re
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from nltk.corpus import stopwords
from nltk.stem.porter import PorterStemmer
from sklearn import naive_bayes, svm
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.linear_model import LogisticRegressionCV
from sklearn.metrics import classification_report, f1_score, plot_roc_curve
from sklearn.model_selection import train_test_split
from yellowbrick.classifier import PrecisionRecallCurve, roc_auc
# +
# load all json files in data directory
files = glob.glob("data/json/*.json")
data = []
# for loop for processing files and adding doc id
for json in files:
frame = pd.read_json(json)
# get file name as string
# create column identifying dfs as doc_id
# split string (remove .json from file name)
frame["doc_id"] = os.path.splitext(os.path.basename(json))[0]
data.append(frame)
# concatenate all data frames
df = pd.concat(data, ignore_index=True)
# -
# ## Cleaning Text
# +
# turn label_id into binary variable
# numpy where function states:
# where label_id is not 4, change to 0, otherwise change to 1
df["label_id"] = np.where(df["label_id"] != 4, 0, 1)
# -
def preprocessing(text):
text = re.sub("<[^>]*>", "", text)
text = re.sub(r"[^\w\s]", "", text)
stop_words = set(stopwords.words("english"))
words = [word for word in text.lower().split() if not word in stop_words]
text = " ".join(words)
return text
df["text"] = df["text"].apply(preprocessing)
# +
ps = PorterStemmer()
def token_ps(text):
return [ps.stem(word) for word in text.split()]
# -
# ## Logistic Regression
#
# Logistic regression is a binary classifier that uses the weighted combination of the input features and models them using a sigmoid function, which gives outputs between 0 and 1. Logistic regression models the probability that _y_, the dependent variable, belongs to a particular class.
#
# - The logistic regression model is based on the values created from a vectorizer algorithm called tf-idf, which stands for term-frequency inverse document frequency.
# - tf-idf measures the originality of the word by comparing how often it appears in a doc with the number of docs the word appears in. The frequency of the words in a doc (compared against other docs) measures the importance of that word in the wider corpus.
# - The logistic regression below is computed by building a vector of word values based on the importance of each word, before using the word vectors to identify the characteristics of the political label to predict which sentences will be political.
# transforming text into vectors
tfidf = TfidfVectorizer(lowercase=False, use_idf=True, norm="l2", smooth_idf=True)
# compute tfidf values for all words in 'text' column of df
X = tfidf.fit_transform(df["text"])
y = df.label_id.values
# splitting data into train and test splits in order to test predictive accuracy
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.4, random_state=0, shuffle=True
)
# +
# computes and then fits logistic regression that implements cross-validation as a part of the process
# cv = number of cross validation folds
log_reg = LogisticRegressionCV(cv=8, scoring="accuracy", n_jobs=-1)
# fit model
log_reg.fit(X_train, y_train)
# model accuracy
log_predictions = log_reg.predict(X_test)
# -
# defining a function that prints model prediction accuracy
def model_accuracy(name, preds):
print("---{} Test Set Results---".format(name))
print("Weighted F1 Average: {}".format(f1_score(y_test, preds, average="weighted")))
# precision = % predicted accurately
# recall = % positives identified
# f1-score = weighted harmonic mean of precision & recall
# weighted f-1 avg used for comparing classification models
print(classification_report(y_test, preds))
model_accuracy("Logit", log_predictions)
# +
# compute and plot roc curves
# auc = measure of "seperability", roc = probability curve
# the closer the curves are to the top left-hand corner, the better the model
roc_auc(log_reg, X_train, y_train, X_test=X_test, y_test=y_test, classes=[0, 1])
# -
# ## Naive Bayes
#
# The Naive Bayes algorithm classifies classes using Bayes' theorem with the assumption of conditional independence between every pair of features, given the value of the class. It is a popular baseline in text classification because it offers several benefits:
#
# - Naive Bayes classifiers are known to perform well on real-world data tasks.
# - They require a small amount of training data to estimate the necessary parameters.
# - Implementation is quick and simple.
# - It is typically quite effective at text classification.
# +
# fit the training dataset on the naive bayes classifier
nb = naive_bayes.ComplementNB()
nb.fit(X_train, y_train)
# model accuracy
nb_predictions = nb.predict(X_test)
model_accuracy("Naive Bayes", nb_predictions)
# -
roc_auc(nb, X_train, y_train, X_test=X_test, y_test=y_test, classes=[0, 1])
# ## Support Vector Classification (SVC)
#
# Support Vector Classifiers find the linear hyperplane (known as the decision boundary) that splits the dataset into classes with the maximum margin between the hyperplane and the nearest data-points (the support vectors). The support vectors are samples that would alter the position of the separating hyerplane if removed, meaning that those samples are the most influential in defining the placement of the decision boundary. By maximizing the functional margin, this reduces how sensitive the decision boundary is to noise, making it more generalizable.
#
# SVCs are effective for classification problems, especially classifying text, for the following reasons:
#
# - They are flexible and fast.
# - They are effective in high-dimensional spaces.
# - They don't require much training data to produce accurate results.
# +
# fit the training dataset on the Support vector classifier
svc = svm.SVC(C=1, kernel="linear", class_weight="balanced")
svc.fit(X_train, y_train)
# model accuracy
svc_predictions = svc.predict(X_test)
model_accuracy("SVC", svc_predictions)
# -
# error plotting yb method, using sklearn instead
## TODO - solve error and plot using more detailed yb method
plot_roc_curve(svc, X_test, y_test)
# ## Pickling Models (for Future Use)
# +
# saving tfidf
pickle.dump(tfidf, open("tfidf.pickle", "wb"))
# saving models
pickle.dump(log_reg, open("log_reg.pickle", "wb"))
pickle.dump(nb, open("nb.pickle", "wb"))
pickle.dump(svc, open("svc.pickle", "wb"))
# -
# ## Comparing Model Performance When Dealing With Imbalanced Data
#
# While ROC Curves are useful, they are sensitive to imbalanced data, as is the case here. One alternative that helps tell us a little more about the data, and is effective when dealing with imbalanced data, is precision-recall curves.
#
# ### Precision-Recall Curve (PRC)
#
# A Precision-Recall Curve is a measure of prediction success. The precision and recall values are calculated from multiple confusion matrices for different probability cut-offs. For example, at a cut-off of x, every case with a probability of greater than x is given a positive label (in this context, every sentence with a probability of greater than x is labeled as political). Precision and recall are calculated at multiple cut-off points, which produces a range of values for which the curve can be plotted.
#
# - PRC effectively plots the tradeoff between the precision (% predicted accurately) and recall (% positives identified).
# - Precision is the proportion of sentences that the model identifies as political that are actually politcal (also know as the Positive Predictive Value (PPV)). Recall is the proportion of politcal sentences that were successfully predicted as political.
# - Precision = True Positives/(True Positives + False Positives)
# - Recall = True Positives/(True Positives + False Negatives)
# - PRC is sensitive to data distribution, is effective for imbalanced data, and responds to positive cases more than negative, which makes it a good fit for this dataset (because there are only a small number of positive cases) and this question (because positives are more important).
# - The Precision-Recall Curves below also plot average precision, which is the weighted average of precision achieved at each cut-off.
# - Average precision represents the area under the curve, and can be considered as a measure of the overall model performance.
# - The goal is to maximize both precision and recall, which would mean that the model is predicting accurately for the majority of classes it selects. When this is the case, the area under the curve would be greater.
# +
# precision-recall curve
# define models
models = [log_reg, nb, svc]
# set plot parameters
_, axes = plt.subplots(ncols=3, figsize=(15, 7))
# for loop calling a PRC curve on each model
for idx, ax in enumerate(axes.flatten()):
viz = PrecisionRecallCurve(models[idx], ax=ax, show=False)
viz.fit(X_train, y_train)
viz.score(X_test, y_test)
viz.finalize()
plt.show()
| models/baseline_models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
from sympy import *
init_printing()
import numpy as np
import sys
import math
# # Diffusion Monte Carlo propagators
# Most of the equations taken from Chapter 24 ("Projector quantum Monte Carlo") in "Interacting Electrons" (2016) by <NAME>, <NAME>, and <NAME>.
# # Trotter breakup
T_op = Symbol('That') # Kinetic energy operator
V_op = Symbol('Vhat') # Potential energy operator
tau = Symbol('tau') # Projection time
n = Symbol('n',isinteger=True) # Number of timestep divisions
dt = Symbol(r'\Delta\tau') # Time for individual timestep
# Eq. 24.7
Eq(exp(-tau *(T_op + V_op)),
Limit(exp(-dt*T_op) * exp(-dt*V_op),n,oo))
# ## In coordinate space, no importance sampling
R = Symbol('R')
Rp = Symbol("R'")
Rpp = Symbol("R''")
ET = Symbol("E_T") # Trial energy
N = Symbol('N',isinteger=True) # number of particles
V = Symbol('V') # potential energy
bracket = lambda a,b,c : Symbol(r'\left\langle{%s}\left|{%s}\right|{%s}\right\rangle'%(latex(a),latex(b),latex(c)))
# Kinetic energy - Eq. 24.8
Eq(bracket(R, exp(-dt*T_op), Rpp),
(2 *pi*dt)**(-3*N/2) * exp(-(R-Rpp)**2/(2*dt)))
# Potential energy - Eq. 24.9
Eq(bracket(Rpp, exp(-dt*(V_op-ET)),Rp),
exp(-dt*V(Rp))*DiracDelta(Rpp-Rp))
# ## In coordinate space, with importance sampling
F = Symbol('F_i')
psiG = Symbol('Psi_G',commutative=False)
EL = Symbol("E_L") # Local energy
H_op = Symbol("Hhat",commutative=False)
gradient = lambda x: Symbol(r'\nabla{%s}'%latex(x))
gradient_with_index = lambda x,i : Symbol(r'\nabla_{%s}{%s}'%(latex(i),latex(x)))
# Quantum force
Eq(F(R), 2*gradient_with_index(log(psiG),Symbol('i')))
# Local energy
Eq(EL(R), psiG**-1 * H_op * psiG)
drift_diffusion = exp(-(Rp-R-S.Half*dt*F(R))**2/(2*dt))
drift_diffusion
branching = exp(-dt*(EL(R)-ET))
branching
prefactor = (2*pi*dt)**(-3*N/2)
prefactor
# Eq. 24.18
prefactor*drift_diffusion*branching
# ## Sampling the drift-diffusion term
chi = Symbol('chi') # gaussian random sample with zero mean and variance delta tau
r = Symbol('r')
rp = Symbol("r'")
# Sample new positions with this formula (Eq 23.13)
# Question - how to detemine sampling formula from evolution equation/distribution above?
sample_drift_diffusion = Eq(rp, r + dt * F + chi)
sample_drift_diffusion
# ## Scaling the drift
#
# In QMCPACK, the drift term is scaled. (From <NAME>, <NAME>, <NAME> "A diffusion Monte Carlo algorithm with very small time-step errors" JCP 99, 2865 (1993) doi: 10.1063/1.465195 )
Fmag = Symbol('Fmag^2')
epsilon = Symbol('epsilon')
drift_scale = Piecewise( (tau,Fmag < epsilon ),
((sqrt(1 + 2*Fmag*tau)-1)/Fmag, True))
drift_scale
scaled_drift = F*drift_scale
scaled_drift
# # Values for Testing
class SymPrinter(printing.lambdarepr.NumPyPrinter):
def _print_Symbol(self, expr):
if expr.name == r'\Delta\tau':
return 'dt'
return expr.name
# +
# RNG corresponding to src/ParticleBase/RandomSeqGenerator.h
def gaussian_rng_list(n):
input_rng = [0.5]*(n+1)
slightly_less_than_one = 1.0 - sys.float_info.epsilon
vals = []
for i in range(0,n,2):
temp1 = math.sqrt(-2.0 * math.log(1.0- slightly_less_than_one*input_rng[i]))
temp2 = 2*math.pi*input_rng[i+1]
vals.append(temp1*math.cos(temp2))
vals.append(temp2*math.sin(temp2))
if n%2 == 1:
temp1 = math.sqrt(-2.0 * math.log(1.0- slightly_less_than_one*input_rng[n-1]))
temp2 = 2*math.pi*input_rng[n]
vals.append(temp1*math.cos(temp2))
return vals
chi_vals = np.array(gaussian_rng_list(6)).reshape((2,3))
chi_vals
# -
r_vals = np.array( [ [1.0, 0.0, 0.0],
[0.0, 0.0, 1.0]])
tau_val = 0.1
scaled_chi_vals = chi_vals * math.sqrt(tau_val)
drift_diffuse_func = lambdify((r, F, chi, dt),sample_drift_diffusion.rhs, printer=SymPrinter)
scaled_drift_func = lambdify((tau, Fmag, F), scaled_drift.subs(epsilon, sys.float_info.epsilon) )
# For a constant wavefunction, gradient is zero
for r_val, chi_val in zip(r_vals, scaled_chi_vals):
rp_val = np.zeros(3)
rp_val = drift_diffuse_func(r_val, np.zeros(3), chi_val, tau_val)
print rp_val
# For a linear wavefunction, gradient is constant
grad_coeff = np.array([ 1.0, 2.0, 3.0])
for r_val, chi_val in zip(r_vals, scaled_chi_vals):
rp_val = np.zeros(3)
# Scaled drift is already multiplied by dt, accomodate by setting dt param to 1.0
rp_val = drift_diffuse_func(r_val, scaled_drift_func(tau_val, np.dot(grad_coeff, grad_coeff),grad_coeff), chi_val, 1.0)
print ['%.15g'%v for v in rp_val]
# Compute scaled drift
drift_scale.subs({epsilon:sys.float_info.epsilon, tau:tau_val, Fmag:np.dot(grad_coeff, grad_coeff)})
| Diffusion/DMC_propagator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Биномиальный критерий для доли
import numpy as np
from scipy import stats
# %pylab inline
# ## Shaken, not stirred
# <NAME> говорит, что предпочитает мартини взболтанным, но не смешанным. Проведём слепой тест (blind test): $n$ раз предложим ему пару напитков и выясним, какой из двух он предпочитает. Получаем:
# * **выборка:** бинарный вектор длины $n$, где 1 — <NAME> предпочел взболтанный напиток, 0 — смешанный;
# * **гипотеза $H_0$:** Дж<NAME> не различает 2 вида напитков и выбирает наугад;
# * **статистика $T$:** количество единиц в выборке.
# Если нулевая гипотеза справедлива и <NAME> действительно выбирает наугад, то мы можем с одинаковой вероятностью получить любой из $2^n$ бинарных векторов длины $n$.
#
# Мы могли бы перебрать все такие векторы, посчитать на каждом значение статистики $T$ и получить таким образом её нулевое распределение. Но в данном случае этот этап можно пропустить: мы имеем дело с выборкой, состоящей из 0 и 1, то есть, из распределения Бернулли $Ber(p)$. Нулевая гипотеза выбора наугад соответствует значению $p=\frac1{2}$, то есть, в каждом эксперименте вероятность выбора взболтанного мартини равна $\frac1{2}$. Сумма $n$ одинаково распределённых бернуллиевских случайных величин с параметром $p$ имеет биномиальное распределение $Bin(n, p)$. Следовательно, нулевое распределение статистики $T$ — $Bin\left(n, \frac1{2}\right)$.
#
# Пусть $n=16.$
n = 16
F_H0 = stats.binom(n, 0.5)
# +
x = np.linspace(0,16,17)
pylab.bar(x, F_H0.pmf(x), align = 'center')
xlim(-0.5, 16.5)
pylab.show()
# -
# ## Односторонняя альтернатива
# **гипотеза $H_1$:** Джеймс Бонд предпочитает взболтанный мартини.
#
# При такой альтернативе более вероятны большие значения статистики; при расчёте достигаемого уровня значимости будем суммировать высоту столбиков в правом хвосте распределения.
pylab.bar(x, F_H0.pmf(x), align = 'center')
pylab.bar(np.linspace(12,16,5), F_H0.pmf(np.linspace(12,16,5)), align = 'center', color='red')
xlim(-0.5, 16.5)
pylab.show()
stats.binom_test(12, 16, 0.5, alternative = 'greater')
pylab.bar(x, F_H0.pmf(x), align = 'center')
pylab.bar(np.linspace(11,16,6), F_H0.pmf(np.linspace(11,16,6)), align = 'center', color='red')
xlim(-0.5, 16.5)
pylab.show()
stats.binom_test(11, 16, 0.5, alternative = 'greater')
# ## Двусторонняя альтернатива
# **гипотеза $H_1$:** <NAME> предпочитает какой-то определённый вид мартини.
#
# При такой альтернативе более вероятны очень большие и очень маленькие значения статистики; при расчёте достигаемого уровня значимости будем суммировать высоту столбиков в правом и левом хвостах распределения.
pylab.bar(x, F_H0.pmf(x), align = 'center')
pylab.bar(np.linspace(12,16,5), F_H0.pmf(np.linspace(12,16,5)), align = 'center', color='red')
pylab.bar(np.linspace(0,4,5), F_H0.pmf(np.linspace(0,4,5)), align = 'center', color='red')
xlim(-0.5, 16.5)
pylab.show()
stats.binom_test(12, 16, 0.5, alternative = 'two-sided')
pylab.bar(x, F_H0.pmf(x), align = 'center')
pylab.bar(np.linspace(13,16,4), F_H0.pmf(np.linspace(13,16,4)), align = 'center', color='red')
pylab.bar(np.linspace(0,3,4), F_H0.pmf(np.linspace(0,3,4)), align = 'center', color='red')
xlim(-0.5, 16.5)
pylab.show()
stats.binom_test(13, 16, 0.5, alternative = 'two-sided')
| 4 Stats for data analysis/Lectures notebooks/5 binomial test for proportion/stat.binomial_test_with_plots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
X = np.array([[-1, -1], [-2, -1], [-3, -2], [1, 1], [2, 1], [3, 2]]) # Different points
Y = np.array([1, 1, 1, 2, 2, 2]) # Labels for each point
from sklearn.naive_bayes import GaussianNB
clf = GaussianNB()
clf.fit(X, Y) # X features Y label
print(clf.predict([[-0.8, -1]]))
clf_pf = GaussianNB()
clf_pf.partial_fit(X, Y, np.unique(Y))
print(clf_pf.predict([[-0.8, -1]]))
| Gaussian NB Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# + code_folding=[]
import numpy as np
import pandas as pd
SOURCE_PATH = "crawled_data/"
DEST_PATH = "anonomized_data/"
def anonomizePage(page, post_map):
print(page)
df = pd.read_csv(SOURCE_PATH + "%d-users.csv.gz" % page)
df['fb_id-a'] = df[col].map({
r['index']: i
for i, r in df['fb_id'].value_counts().reset_index().iterrows()
})
df['post_id-a'] = df['post_id'].map(post_map)
(df.dropna().sort_values(['fb_id-a']).to_csv(
DEST_PATH + "%d.csv" % page,
float_format="%d",
columns=['post_id-a', 'fb_id-a', 'type'],
header=['post', 'user', 'type'],
index=False))
def anonomizeInteractions(interactions):
post_map = interactions['post_id'].drop_duplicates().reset_index(drop=True)
post_map = pd.Series(post_map.index.values, index=post_map)
interactions['post_id'] = interactions['post_id'].map(post_map)
interactions.to_csv(DEST_PATH + '00__combinedPageInteractions.csv',
float_format="%d", index=False)
return post_map
# +
interactions = pd.read_csv(SOURCE_PATH + 'combinedPageInteractions.csv.bz2', parse_dates=[7])
post_map = anonomizeInteractions(interactions)
pages = interactions.page_id.unique()
print(len(pages))
for page in pages:
print('Anonomizes {}'.format(page))
anonomizePage(page, post_map)
| Anonomizer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import h5py, os
import numpy as np
import matplotlib.pyplot as plt
from imp import reload
import pandas as pd
from dca import analysis, plotting, data_util, style
# -
reload(data_util)
M1 = data_util.load_sabes_data('neural/indy_20160627_01.mat')
reload(data_util)
HC = data_util.load_kording_paper_data('neural/example_data_hc.pickle')
reload(data_util)
weather = data_util.load_weather_data('weather/temperature.csv')
reload(data_util)
file = 'accel/sub_19.csv'
accel = data_util.load_accel_data(file)
HC['neural'].shape
# +
T_pi_vals = np.arange(1, 11)
dim_vals = np.array([5, 10, 15, 25])
accel_dims = np.array([3, 4, 5, 6])
temp_dims = np.array([3, 4, 5, 6])
hc_dims = np.array([10, 15, 25, 30])
offsets = np.array([0, 5, 10, 15])
accel_offsets = np.array([0, 3, 6, 9])
win = 3
n_cv = 5
n_init = 5
# +
reload(analysis)
reload(plotting)
M1_results = analysis.run_analysis(M1['M1'], M1['cursor'], T_pi_vals, dim_vals=dim_vals, offset_vals=offsets,
num_cv_folds=n_cv, decoding_window=win, n_init=n_init, verbose=True)
HC_results = analysis.run_analysis(HC['neural'], HC['loc'], T_pi_vals, dim_vals=hc_dims, offset_vals=offsets,
num_cv_folds=n_cv, decoding_window=win, n_init=n_init, verbose=True)
weather_results = analysis.run_analysis(weather, weather, T_pi_vals, dim_vals=dim_vals, offset_vals=offsets,
num_cv_folds=n_cv, decoding_window=win, n_init=n_init, verbose=True)
accel_results = analysis.run_analysis(accel, accel, T_pi_vals, dim_vals=accel_dims, offset_vals=accel_offsets,
num_cv_folds=n_cv, decoding_window=win, n_init=n_init, verbose=True)
# -
np.savez('real_data.npz', M1_results=M1_results, HC_results=HC_results,
weather_results=weather_results, accel_results=accel_results)
d = np.load('real_data.npz')
M1_results = d['M1_results']
HC_results = d['HC_results']
weather_results = d['weather_results']
accel_results = d['accel_results']
# +
reload(plotting)
fig, axes = plotting.decoding_fix_axes(fig_width=5.5, fig_height=2.7,
wpad_mid=.075, hpad_mid=.1,
wpad_left=.09, wpad_right=.025,
hpad_bot=.12)
loc='lower right'
bbox_to_anchor=(1.18, -.05)
plotting.scatter_r2_vals(M1_results, 4, dim_vals, offsets, T_pi_vals, min_val=0, max_val=.5,
ax=axes[0], title='M1', timestep_units='ms', timestep=50, loc=loc,
bbox_to_anchor=bbox_to_anchor)
plotting.plot_r2_vs_T(M1_results, T_pi_vals, dim_vals, offsets, offset_idx=1, min_max_val=.08,
ax=axes[4], legend=False, timestep_units='ms', timestep=50)
plotting.scatter_r2_vals(HC_results, 4, hc_dims, offsets, T_pi_vals, min_val=0, max_val=.15,
ax=axes[1], ylabel=False, title='Hippocampus',
timestep_units='ms', timestep=50, loc=loc,
bbox_to_anchor=bbox_to_anchor)
plotting.plot_r2_vs_T(HC_results, T_pi_vals, hc_dims, offsets, offset_idx=1, min_max_val=.04,
ax=axes[5], legend=False, ylabel=False, timestep_units='ms', timestep=50)
plotting.scatter_r2_vals(weather_results, 4, temp_dims, offsets, T_pi_vals, min_val=.6, max_val=1.,
ax=axes[2], ylabel=False, title='Temperature',
timestep_units='days', timestep=1, legend=True, loc=loc,
bbox_to_anchor=bbox_to_anchor)
plotting.plot_r2_vs_T(weather_results, T_pi_vals, temp_dims, offsets, offset_idx=0, min_max_val=.05,
ax=axes[6], legend=False, ylabel=False, timestep_units='day', timestep=1)
plotting.scatter_r2_vals(accel_results, 4, accel_dims, accel_offsets, T_pi_vals, min_val=.2, max_val=.9,
ax=axes[3], ylabel=False, title='Accelerometer',
timestep_units='ms', timestep=20,loc=loc,
bbox_to_anchor=bbox_to_anchor)
plotting.plot_r2_vs_T(accel_results, T_pi_vals, accel_dims, accel_offsets, offset_idx=1, min_max_val=.4,
ax=axes[7], legend=False, ylabel=False, timestep_units='ms', timestep=20)
#plt.savefig(os.path.join(os.environ['HOME'], 'Downloads/real_data.pdf'), dpi=300)
#plt.savefig(os.path.join(os.environ['HOME'], 'Downloads/real_data.png'), dpi=300)
# -
# # SFA R^2
# +
reload(plotting)
fig, axes = plotting.decoding_fix_axes(fig_width=5.5, fig_height=2.7,
wpad_mid=.075, hpad_mid=.11,
wpad_left=.09, wpad_right=.025,
hpad_bot=.12)
loc='lower right'
bbox_to_anchor=(1.18, -.05)
plotting.plot_absolute_r2_vs_T(M1_results, T_pi_vals, dim_vals, offsets, offset_idx=1, min_max_val=.5,
ax=axes[0], legend=False, timestep_units='ms', timestep=50)
plotting.plot_absolute_r2_vs_T(M1_results, T_pi_vals, dim_vals, offsets, offset_idx=1, min_max_val=.5,
ax=axes[4], legend=False, timestep_units='ms', timestep=50, dca=False)
plotting.plot_absolute_r2_vs_T(HC_results, T_pi_vals, hc_dims, offsets, offset_idx=1, min_max_val=.15,
ax=axes[1], legend=False, ylabel=False, timestep_units='ms', timestep=50)
plotting.plot_absolute_r2_vs_T(HC_results, T_pi_vals, hc_dims, offsets, offset_idx=1, min_max_val=.15,
ax=axes[5], legend=False, ylabel=False, timestep_units='ms', timestep=50, dca=False)
plotting.plot_absolute_r2_vs_T(weather_results, T_pi_vals, temp_dims, offsets, offset_idx=0, min_max_val=1.1,
ax=axes[2], legend=False, ylabel=False, timestep_units='day', timestep=1)
plotting.plot_absolute_r2_vs_T(weather_results, T_pi_vals, temp_dims, offsets, offset_idx=0, min_max_val=1.1,
ax=axes[6], legend=False, ylabel=False, timestep_units='day', timestep=1, dca=False)
plotting.plot_absolute_r2_vs_T(accel_results, T_pi_vals, accel_dims, accel_offsets, offset_idx=1, min_max_val=1.,
ax=axes[3], legend=False, ylabel=False, timestep_units='ms', timestep=20)
plotting.plot_absolute_r2_vs_T(accel_results, T_pi_vals, accel_dims, accel_offsets, offset_idx=1, min_max_val=1.,
ax=axes[7], legend=False, ylabel=False, timestep_units='ms', timestep=20, dca=False)
plt.savefig(os.path.join(os.environ['HOME'], 'Downloads/real_data_absolute_r2.pdf'), dpi=300)
#plt.savefig(os.path.join(os.environ['HOME'], 'Downloads/real_data.png'), dpi=300)
# -
# # Neural Forcasting
# +
T_pi_vals = np.arange(1, 11)
m1_dims = np.array([10, 20, 30, 40])
dims = np.array([5, 10, 15, 25])
offsets = np.array([0, 2, 4, 8])
win = 3
n_cv = 5
n_init = 5
# +
reload(analysis)
reload(plotting)
M1_forecast_results = analysis.run_analysis(M1['M1'], M1['M1'], T_pi_vals, dim_vals=m1_dims, offset_vals=offsets,
num_cv_folds=n_cv, decoding_window=win, n_init=n_init, verbose=True)
HC_forecast_results = analysis.run_analysis(HC['neural'], HC['neural'], T_pi_vals, dim_vals=dims,
offset_vals=offsets, num_cv_folds=n_cv, decoding_window=win,
n_init=n_init, verbose=True)
# -
np.savez('neural_forecast.npz', M1_forecast_results=M1_forecast_results, HC_forecast_results=HC_forecast_results)
# +
reload(plotting)
fig, axes = plotting.decoding_fix_axes2(fig_width=3.5, fig_height=3.5,
wpad_mid=.15, hpad_mid=.1,
wpad_left=.2, wpad_right=.05,
hpad_bot=.12)
loc='upper left'
bbox_to_anchor=(.525, .4)
plotting.scatter_r2_vals(M1_forecast_results, 4, m1_dims, offsets, T_pi_vals, min_val=0, max_val=.65,
ax=axes[0], title='M1', timestep_units='ms', timestep=50, loc=loc,
bbox_to_anchor=bbox_to_anchor)
plotting.plot_pi_vs_T(M1_forecast_results, T_pi_vals, m1_dims, offsets, offset_idx=2, min_max_val=2e-3,
ax=axes[2], legend=False, timestep_units='ms', timestep=50)
loc='upper left'
bbox_to_anchor=(-.1, .8)
plotting.scatter_r2_vals(HC_forecast_results, 4, dims, offsets, T_pi_vals, min_val=0, max_val=1.,
ax=axes[1], ylabel=False, title='Hippocampus',
timestep_units='ms', timestep=50, loc=loc,
bbox_to_anchor=bbox_to_anchor)
plotting.plot_pi_vs_T(HC_forecast_results, T_pi_vals, dims, offsets, offset_idx=3, min_max_val=.01,
ax=axes[3], legend=False, ylabel=False, timestep_units='ms', timestep=50)
plt.savefig(os.path.join(os.environ['HOME'], 'Downloads/real_data_neural_forecast.pdf'), dpi=300)
#plt.savefig(os.path.join(os.environ['HOME'], 'Downloads/real_data.png'), dpi=300)
# -
| notebooks/real_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GRpc接口服务
#
# GRpc正如其名,是一种RPC.它实际上和RESTful接口在功能上是相近的,本质都是一种请求响应模式的服务.只是作为一个RPC,GRpc一般描述动作而非资源,并且它可以返回的不光是一个数据,而是一组流数据.
#
# GRpc是一种跨语言的Rpc,它建立在http2上使用protobuf作为结构化数据的序列化工具.
#
# 它有4种形式:
#
# + 请求-响应
# + 请求-流响应
# + 流请求-响应
# + 流请求-流响应
#
# 其基本使用方式是:
#
# 1. 服务端与客户端开发者协商创建一个protobuf文件用于定义rpc的形式和方法名以及不同方法传输数据的schema
# 2. 编译protobuf文件至服务端客户端的实现语言
# 3. 服务端实现protobuf文件中定义的方法
# 4. 客户端调用protobuf文件中定义的方法
#
# 在python中我们使用[protobuf](https://pypi.org/project/protobuf/)和[grpcio](https://pypi.org/project/grpcio/)来编译protobuf文件.
# ## 请求-响应
#
# 这个例子[C0](https://github.com/TutorialForPython/python-io/tree/master/%E6%8E%A5%E5%8F%A3%E6%9C%8D%E5%8A%A1/RPC%E6%9C%8D%E5%8A%A1/code/GRpc%E6%8E%A5%E5%8F%A3%E6%9C%8D%E5%8A%A1/C0)我们来实现一个简单的服务--输入一个数,输出这个数的平方
#
# ### 创建一个protobuf文件
#
# 创建protobuf文件的语法可以看[protobuf的语法指南](https://developers.google.com/protocol-buffers/docs/proto3)
#
# 我们将函数命名为Square,每次传传入的数据是一个double型的数,传回的也是一个double型的数.
#
# ```proto
# syntax = "proto3";
# package squarerpc_service;
#
# service SquareService {
# rpc square (Message) returns (Message){}
# }
#
# message Message {
# double message = 1;
# }
# ```
#
# ### 将这个proto文件编译为python模块
#
# 要将proto文件编译为python模块我们需要工具[protoc](https://github.com/protocolbuffers/protobuf/releases)和[grpcio-tools](https://pypi.org/project/grpcio-tools/)
#
# 安装好这两个后我们可以使用如下命令将目标protobuf文件编译为
#
# ```shell
# python -m grpc_tools.protoc -I=$proto_dir \
# --python_out=$target_dir \
# --grpc_python_out=$target_dir \
# $proto_file
# ```
#
# ### 服务端实现定义的方法
#
# python的grpc服务端是使用线程实现的,这也就意味着它无法承受高并发.但这通常不是rpc关注的问题,rpc一般都是要通过起多个实例做负载均衡的,同时这也要求了我们的rpc要做到无状态.
#
# + server.py
#
# ```python
# # #!/usr/bin/env python
# import time
# from concurrent import futures
# import grpc
# from data_pb2_grpc import SquareServiceServicer, add_SquareServiceServicer_to_server
# from data_pb2 import Message
#
# HOST = "0.0.0.0"
# PORT = 5000
# ONE_DAY_IN_SECONDS = 60 * 60 * 24
#
# class SquareServic(SquareServiceServicer):
# def square(self, request, context):
# return Message(message=request.message**2)
#
# def main():
# grpcServer = grpc.server(futures.ThreadPoolExecutor(max_workers=10))
# add_SquareServiceServicer_to_server(SquareServic(), grpcServer)
# print(f'"msg":"grpc start @ grpc://{HOST}:{PORT}"')
# grpcServer.add_insecure_port(f"{HOST}:{PORT}")
# grpcServer.start()
# try:
# while True:
# time.sleep(ONE_DAY_IN_SECONDS)
# except KeyboardInterrupt:
# grpcServer.stop(0)
# except Exception as e:
# grpcServer.stop(0)
# raise
#
# if __name__ == "__main__":
# main()
# ```
#
# grpc服务端写法步骤:
#
# 1. 继承我们定义的`service`名字加`Servicer`的抽象类,并实现其中的方法,姑且叫他`RPC`类
# 2. 使用`grpc.server(executor[,maximum_concurrent_rpcs])`创建一个服务器,executor必须使用标准库的`futures.ThreadPoolExecutor`,通过它来确定最大使用多少个worker.`maximum_concurrent_rpcs`则是用于设置最大同时处理多少个请求,当请求超过`maximum_concurrent_rpcs`的数值,name后来的请求就会被拒绝
# 3. 使用`add_[我们定义的service名字]Servicer_to_server(RPC类的实例,grpcServer)`
# 4. `grpcServer.add_insecure_port(f"{HOST}:{PORT}")`绑定ip和端口
# 5. `grpcServer.start()`启动服务,需要注意的是`grpcServer.start()`是启动新线程实现的,需要阻塞主线程以防止退出,因此需要有一个死循环在主线程.
#
# #### 借助多进程提高cpu利用率
#
# python写rpc往往是处理计算密集型任务,但GIL让rpc无法高效的利用cpu.我们来重新实现下上面的服务,让计算的部分由多进程实现
#
# ```python
# Executor = futures.ProcessPoolExecutor(max_workers=3)
#
# class SquareServic(SquareServiceServicer):
# def square(self, request, context):
# f = Executor.submit(square,request.message)
# futures.as_completed(f)
# result = f.result()
# return Message(message=result)
#
# ...
#
# def main():
# try:
# while True:
# time.sleep(ONE_DAY_IN_SECONDS)
# except KeyboardInterrupt:
# grpcServer.stop(0)
# Executor.shutdown()
# except Exception as e:
# grpcServer.stop(0)
# Executor.shutdown()
# raise
# ```
#
# ### 客户端实现方式
#
# python作为客户端同样比较常见.毕竟更多的时候我们是要调用别人写的服务
#
# 客户端需要做的是
#
# 1. 连接上服务器
# 2. 构造一个Stub的实例
# 3. 调用stub实例上的对应方法并获得结果
#
# #### 同步客户端
#
# 官方默认使用的是同步写法,比较直观
#
# + cli_sync.py
#
# ```python
# # #!/usr/bin/env python
# import grpc
# from data_pb2_grpc import SquareServiceStub
# from data_pb2 import Message
# url = "localhost:5000"
# channel = grpc.insecure_channel(url)
# client = SquareServiceStub(channel=channel)
# result = client.square(Message(message=12.3))
# print(result)
# ```
#
# #### 异步客户端
#
# grpc的客户端有两种请求有`.future()`方法可以返回`grpc.Future`,它的接口和`asyncio.Future`十分类似,但却不满足`awaitable`协议.感谢[aiogrpc](https://github.com/hubo1016/aiogrpc)为我们做了一个包装器,用它我们就可以使用协程语法了
#
# + cli_async.py
#
# ```python
# # #!/usr/bin/env python
# import asyncio
# from aiogrpc import insecure_channel
# from data_pb2_grpc import SquareServiceStub
# from data_pb2 import Message
#
# url = "localhost:5000"
#
# async def query():
# async with insecure_channel(url) as conn:
# client = SquareServiceStub(channel=conn)
# result = await client.square(Message(message=12.3))
# print(result)
#
#
# def main():
# loop = asyncio.get_event_loop()
# loop.run_until_complete(query())
#
#
# if __name__ == "__main__":
# main()
#
# ```
#
# ### 添加ssl支持
#
# 由于python常用于做原型开发,所以很多时候它需要独立完成部署而不能借助其他工具,那ssl支持就是一个必须要考虑的问题了
#
# grpc原生支持ssl只需要:
#
# + 服务端修改
#
# ```python
# grpcServer.add_insecure_port
# ```
#
# 改为
#
# ```python
# with open('crt/example.key', 'rb') as f:
# private_key = f.read()
# with open('crt/example.crt', 'rb') as f:
# certificate_chain = f.read()
# server_credentials = grpc.ssl_server_credentials(
# ((private_key, certificate_chain,),))
# grpcServer.add_secure_port(f"{HOST}:{PORT}", server_credentials)
# ```
#
# + 客户端修改
#
# ```python
# conn = grpc.insecure_channel(url)
# ```
#
# 改为
#
# ```python
# with open('crt/example.crt', 'rb') as f:
# trusted_certs = f.read()
# credentials = grpc.ssl_channel_credentials(root_certificates=trusted_certs)
# channel = grpc.secure_channel(url, credentials)
# ```
#
# ## 请求-流响应
#
# 这种需求比较常见,有点类似python中的range函数,它生成的是一个流而非一个数组,它会一次一条的按顺序将数据发送回请求的客户端.
#
# 这个例子[C1](https://github.com/TutorialForPython/python-io/tree/master/%E6%8E%A5%E5%8F%A3%E6%9C%8D%E5%8A%A1/RPC%E6%9C%8D%E5%8A%A1/code/GRpc%E6%8E%A5%E5%8F%A3%E6%9C%8D%E5%8A%A1/C1)实现了给出一个正整数,它会返回从0开始到它为止的每个整数的平方.
#
# ### 修改protobuf文件
#
# ```proto
# service SquareService {
# rpc rangeSquare (Message) returns (stream Message){}
# }
# ```
#
# ### 修改服务端实现
#
# 服务一端的流是一年`yield`关键字推送
#
# ```python
# ...
# class SquareServic(SquareServiceServicer):
# def rangeSquare(self, request, context):
# print(request.message)
# for i in range(int(request.message+1)):
# yield Message(message=i**2)
# ```
#
# ### 修改客户端实现
#
# 我们在客户端可以直接用for循环读取返回的流
#
# #### 同步客户端
#
# ```python
# ...
# for i in client.rangeSquare(Message(message=12)):
# print(i.message)
# ```
#
# #### 异步客户端
#
# ```python
# ...
# async def query():
# async with insecure_channel(url) as conn:
# client = SquareServiceStub(channel=conn)
# async for response in client.rangeSquare(Message(message=12)):
# print(response.message)
# ...
# ```
#
# ## 流请求-响应
#
# 这种需求不是很多见,可能用的比较多的是收集一串数据后统一进行处理吧,流只是可以确保是同一个客户端发过来的而已.
#
# 这个例子[C2](https://github.com/TutorialForPython/python-io/tree/master/%E6%8E%A5%E5%8F%A3%E6%9C%8D%E5%8A%A1/RPC%E6%9C%8D%E5%8A%A1/code/GRpc%E6%8E%A5%E5%8F%A3%E6%9C%8D%E5%8A%A1/C2)实现了传过来一串数,之后返回他们的平方和
#
# ### 修改protobuf文件
#
# ```proto
# service SquareService {
# rpc sumSquare (stream Message) returns (Message){}
# }
# ```
#
# ### 修改服务端实现
#
# 以流为请求的服务端第二个参数为一个`iterator`,因此我们可以使用for循环来获取其中的内容
#
# ```python
# ...
# class SquareServic(SquareServiceServicer):
# def sumSquare(self, request_iterator, context):
# result = 0
# for i in request_iterator:
# result += i.message**2
# return Message(message=result)
# ```
#
# ### 修改客户端实现
#
# 我们在客户端可以直接用for循环读取返回的流
#
# #### 同步客户端
#
# 同步客户端通过将一个`iterator`作为参数来调用以流为请求的服务端
#
# ```python
# ...
# result = client.sumSquare(Message(message=i) for i in range(12))
# ```
#
# #### 异步客户端
#
# 异步客户端则接收一个异步iterator作为参数,我们可以使用`aitertools`来生成或者处理异步迭代器,如果请求流比较复杂,我们也可以创建一个异步生成器,异步语法可以看[我的这篇文章](https://tutorialforpython.github.io/%E8%AF%AD%E6%B3%95%E7%AF%87/%E6%B5%81%E7%A8%8B%E6%8E%A7%E5%88%B6/%E9%98%BB%E5%A1%9E%E5%BC%82%E6%AD%A5%E4%B8%8E%E5%8D%8F%E7%A8%8B.html#%E5%BC%82%E6%AD%A5%E8%BF%AD%E4%BB%A3%E5%99%A8%E5%92%8Casync-for)
#
# ```python
# from aitertools import AsyncIterWrapper
# ...
# response = await client.sumSquare(AsyncIterWrapper(Message(message=i) for i in range(12)))
# ...
# ```
#
#
# ## 流请求-流响应
#
# 将上面两种方式结合起来,就是我们的第四种方式,请求为一个流,响应也是流.这两个流可以是相互交叉的也可以是请求完后再返回一个流.他们在写pb文件时是相同的写法
#
# ```proto
# service SquareService {
# rpc streamrangeSquare (stream Message) returns (stream Message){}
# }
# ```
#
#
# ### 请求流完成后返回流
#
#
# 这个例子[C3](https://github.com/TutorialForPython/python-server/tree/master/code/GRpc%E6%8E%A5%E5%8F%A3%E6%9C%8D%E5%8A%A1/C3)实现了传过来一串数,之后以流的形式返回这组数每个的平方.
#
# #### 修改服务端实现
#
# 服务端获得的请求是一个`iterator`,而返回的流则是通过yield语法推出的.
#
# ```python
# ...
# class SquareServic(SquareServiceServicer):
# def streamrangeSquare(self, request_iterator, context):
# result = []
# for i in request_iterator:
# result.append(i.message**2)
# for j in result:
# yield Message(message=j)
# ```
#
# #### 修改同步客户端
#
# 同步客户端请求的参数是一个`iterator`,返回的也是一个`iterator`
#
# ```python
# ...
# for result in client.streamrangeSquare(Message(message=i) for i in range(12)):
# print(result)
# ```
#
# #### 修改异步客户端
#
# 异步客户端请求的参数是一个`async iterator`,返回的也是一个`async iterator`
#
# ```python
# async with insecure_channel(url) as conn:
# client = SquareServiceStub(channel=conn)
# async for response in client.streamrangeSquare(AsyncIterWrapper(Message(message=i) for i in range(12))):
# print(response.message)
# ...
# ```
#
# ### 请求流中返回流
#
# 这个例子[C4](https://github.com/TutorialForPython/python-io/tree/master/%E6%8E%A5%E5%8F%A3%E6%9C%8D%E5%8A%A1/RPC%E6%9C%8D%E5%8A%A1/code/GRpc%E6%8E%A5%E5%8F%A3%E6%9C%8D%E5%8A%A1/C4)实现了传过来一串数,过程中每传来一个数就返回它的平方
#
# #### 修改服务端实现
#
# 这种其实只需要修改服务端即可,在每获得一个数据后就yield出去结果就行了
#
# ```python
# ...
# class SquareServic(SquareServiceServicer):
# def streamrangeSquare(self, request_iterator, context):
# for i in request_iterator:
# yield Message(message=i.message**2)
# ```
#
# ## 总结
#
# python的GRpc接口充分利用了python语法中的iterator协议,因此无论是服务端客户端都可以写出相当简短的服务.调用起来也最像本地的模块.
| 接口服务/RPC服务/GRpc接口服务.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# ## Array Manipulation in Numpy
#
# - Lets define the following 2D array where the rows are subjects (people) the first column is people weights and the second column is their heights
# +
import numpy as np
#[weight, height] of each people
X = np.array(
[[73.847017017515, 241.893563180437],
[68.7819040458903, 162.3104725213],
[74.1101053917849, 212.7408555565],
[71.7309784033377, 220.042470303077],
[69.8817958611153, 206.349800623871],
[67.2530156878065, 152.212155757083],
[68.7850812516616, 183.927888604031],
[68.3485155115879, 167.971110489509],
[67.018949662883, 175.92944039571],
[63.4564939783664, 156.399676387112]]
)
# -
X.shape
# ## the above values means we have 10 subjects (10 samples) each have two features (weight and height for each subject)
# ## or X has 10 rows and 2 columns
# + [markdown] slideshow={"slide_type": "slide"}
# ## Activity: What is the mean of weights
# + slideshow={"slide_type": "subslide"}
print(X[:, 0])
# + slideshow={"slide_type": "-"}
np.mean(X[:, 0]) #row 1 is weight
# + [markdown] slideshow={"slide_type": "slide"}
# ## Activity: What is the mean of heights
# + slideshow={"slide_type": "subslide"}
print(X[:, 1]) #row 1 is heights
# -
np.mean(X[:, 1])
# + [markdown] slideshow={"slide_type": "slide"}
# ## Activity: what is the mean of height for the first 5 people
# + slideshow={"slide_type": "subslide"}
print(X[0:5, 1]) #first 5 elements of row 1
# -
np.mean(X[0:5, 1])
# + [markdown] slideshow={"slide_type": "slide"}
# ## Activity: what is the weight of the 9th subject?
# + slideshow={"slide_type": "subslide"}
print(X[8, 0]) #height of 9th person
# + [markdown] slideshow={"slide_type": "slide"}
# ## Activity: Subtract the mean from corresponding columns
# + slideshow={"slide_type": "subslide"}
np.mean(X, axis=0) #average of columns, meaning the weight column's mean and height column's mean
#axis=0 gets the mean column wise
#axis=1 gets the mean of row wise; which is meaningless
# -
X - np.mean(X, axis=0) #each element will get subtracted from the row's mean
X - np.mean(X, axis=1)
# This is meaningless for our dataset
np.mean(X, axis=1)
np.mean(X[0, :])
np.mean(X[1, :])
# ## Column-wise addition, row-wise addition
# + slideshow={"slide_type": "slide"}
import numpy as np
A = np.array([[1,2],[3,4]])
print(A)
print(np.sum(A)) # Compute sum of all elements; prints "10"
print(np.sum(A, axis=0)) # Compute sum of each column; prints "[4 6]"
print(np.sum(A, axis=1)) # Compute sum of each row; prints "[3 7]"
# -
# ## Broadcasting in Numpy
# +
import numpy as np
A = np.array([[1,2,3], [4,5,6], [7,8,9], [10, 11, 12]])
print(A.shape)
v = np.array([1, 0, 1])
# no need to do A + np.ones((4, 1))*v
A + v
# -
B= A.reshape(2, 6)
# ## Resources
#
# - http://cs231n.github.io/python-numpy-tutorial/
| classwork/array_manipulation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Encrypted Deep Learning in PyTorch
import torch
import torch.nn as nn
from torch import optim
import torch.nn.functional as F
# Toy Dataset
data = torch.tensor([[0,1], [0,1], [1,0], [1,1.]], requires_grad=True)
targets = torch.tensor([[0], [0], [1], [1.]], requires_grad=True)
# +
class Model(nn.Module):
def __init__(self):
super().__init__()
self.fc1 = nn.Linear(2, 20)
self.fc2 = nn.Linear(20, 1)
def forward(self, x):
x = F.relu(self.fc1(x))
x = self.fc2(x)
return x
model = Model()
# -
optimizer = optim.SGD(model.parameters(), lr=0.05)
for e in range(10):
optimizer.zero_grad()
output = model(data)
loss = ((output - targets) ** 2).sum()
loss.backward()
optimizer.step()
print(loss.data)
preds = model(data)
preds
# ### Using Encryption
import syft as sy
hook = sy.TorchHook(torch)
bob = sy.VirtualWorker(hook, id='bob').add_worker(sy.local_worker)
alice = sy.VirtualWorker(hook, id='alice').add_worker(sy.local_worker)
secure_worker = sy.VirtualWorker(hook, id='sec_worker').add_worker(sy.local_worker)
encrypted_model = model.fix_precision().share(bob, alice, crypto_provider=secure_worker)
list(encrypted_model.parameters())
encrypted_data = data.fix_precision().share(bob, alice, crypto_provider=secure_worker)
encrypted_data
encrypted_prediction = encrypted_model(encrypted_data)
decrypted_prediction = encrypted_prediction.get().float_precision()
decrypted_prediction
| day-4/Encrypted Deep Learning with Pytorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Training an Object Detector (ResNet)
# !pip install gluoncv
import gluoncv as gcv
import mxnet as mx
# ## Preparing the Training Dataset
# +
import os
class DetectionDataset(gcv.data.VOCDetection):
CLASSES = ['cocacola', 'cocacola-zero', 'juice', 'noodles', 'hand'] # , 'cocacola-shadow', 'cocacola-zero-shadow', 'juice-shadow', 'noodles-shadow'
def __init__(self, root):
self._im_shapes = {}
self._root = os.path.expanduser(root)
self._transform = None
self._items = [(self._root, x.strip('.xml')) for x in os.listdir(self._root) if x.endswith('.xml')]
self._anno_path = os.path.join('{}', '{}.xml')
self._image_path = os.path.join('{}', '{}.jpg')
self.index_map = dict(zip(self.classes, range(self.num_class)))
self._label_cache = self._preload_labels()
def __str__(self):
detail = self._root
return self.__class__.__name__ + '(' + detail + ')'
@property
def classes(self):
return self.CLASSES
@property
def num_class(self):
return len(self.classes)
train_dataset = DetectionDataset('../images/v2')
print('class_names:', train_dataset.classes)
print('num_images:', len(train_dataset))
# -
# ## Visualizing an Image
# +
from matplotlib import pyplot as plt
from gluoncv.utils import viz
sample = train_dataset[0]
train_image = sample[0]
train_label = sample[1]
ax = viz.plot_bbox(
train_image.asnumpy(),
train_label[:, :4],
labels=train_label[:, 4:5],
class_names=train_dataset.classes)
plt.show()
# for i in range(len(train_dataset)):
# sample = train_dataset[i]
# train_image = sample[0]
# train_label = sample[1]
# ax = viz.plot_bbox(
# train_image.asnumpy(),
# train_label[:, :4],
# labels=train_label[:, 4:5],
# class_names=train_dataset.classes)
# plt.show()
# -
# ## Model Training
# ### Training Functions
import time
from datetime import datetime
from mxnet import autograd
from gluoncv.data.batchify import Tuple, Stack, Pad
from gluoncv.data.transforms.presets.ssd import SSDDefaultTrainTransform
# +
def train_model(train_dataset, epochs=50):
ctx = mx.gpu(0)
net = gcv.model_zoo.get_model('ssd_512_resnet50_v1_custom',
classes=train_dataset.classes,
transfer='coco')
net.collect_params().reset_ctx(ctx)
width, height = 512, 512 # suppose we use 512 as base training size
gcv.utils.random.seed(233)
batch_size = 16 # 32 for p3.2xlarge, 16 for p2.2xlarge
num_workers = 4
with autograd.train_mode():
_, _, anchors = net(mx.nd.zeros((1, 3, height, width), ctx))
anchors = anchors.as_in_context(mx.cpu())
train_transform = SSDDefaultTrainTransform(width, height, anchors)
batchify_fn = Tuple(Stack(), Stack(), Stack())
train_loader = mx.gluon.data.DataLoader(
train_dataset.transform(train_transform),
batch_size,
shuffle=True,
batchify_fn=batchify_fn,
last_batch='rollover',
num_workers=num_workers)
mbox_loss = gcv.loss.SSDMultiBoxLoss()
ce_metric = mx.metric.Loss('CrossEntropy')
smoothl1_metric = mx.metric.Loss('SmoothL1')
for k, v in net.collect_params().items():
if 'convpredictor' not in k:
# freeze upper layers
v.grad_req = 'null'
trainer = mx.gluon.Trainer(
net.collect_params(), 'sgd',
{'learning_rate': 0.001, 'wd': 0.0005, 'momentum': 0.9})
net.hybridize(static_alloc=True, static_shape=True)
for epoch in range(epochs):
tic = time.time()
btic = time.time()
for i, batch in enumerate(train_loader):
data = mx.gluon.utils.split_and_load(batch[0], ctx_list=[ctx], batch_axis=0)
cls_targets = mx.gluon.utils.split_and_load(batch[1], ctx_list=[ctx], batch_axis=0)
box_targets = mx.gluon.utils.split_and_load(batch[2], ctx_list=[ctx], batch_axis=0)
with autograd.record():
cls_preds = []
box_preds = []
for x in data:
cls_pred, box_pred, _ = net(x)
cls_preds.append(cls_pred)
box_preds.append(box_pred)
sum_loss, cls_loss, box_loss = mbox_loss(
cls_preds, box_preds, cls_targets, box_targets)
autograd.backward(sum_loss)
# since we have already normalized the loss, we don't want to normalize
# by batch-size anymore
trainer.step(1)
ce_metric.update(0, [l * batch_size for l in cls_loss])
smoothl1_metric.update(0, [l * batch_size for l in box_loss])
name1, loss1 = ce_metric.get()
name2, loss2 = smoothl1_metric.get()
print('[Epoch {}][Batch {}], Speed: {:.3f} samples/sec, {}={:.3f}, {}={:.3f}'.format(
epoch, i, batch_size/(time.time()-btic), name1, loss1, name2, loss2))
btic = time.time()
return net
# -
# ### Training an Object Detector
epochs = 300
net = train_model(train_dataset, epochs=epochs)
save_file = 'object_detector_epoch{}_{}.params'.format(
epochs, datetime.now().strftime("%m_%d_%Y_%H_%M_%S"))
net.save_parameters(save_file)
print('Saved model to disk: ' + save_file)
| src/training_object_detector.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 23570, "status": "ok", "timestamp": 1623923672121, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhSasqlkSGouVaJ4_6HwZYs3ccarzD6XSfX3qG7=s64", "userId": "11001256407696526704"}, "user_tz": -330} id="A215vhb3e024" outputId="7723da73-2255-4576-a2f0-da255dec0243"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 469, "status": "ok", "timestamp": 1623923690856, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhSasqlkSGouVaJ4_6HwZYs3ccarzD6XSfX3qG7=s64", "userId": "11001256407696526704"}, "user_tz": -330} id="s9sKpxn7ilgz" outputId="5a7d514a-0cb9-4a1f-fd4e-4c46d5322641"
# !ls "/content/drive/My Drive/bert/"
# + id="-p4NujezMUTX"
# # !pip install transformers
# # !pip install dataset
# + id="hFzTo2zCZoEj"
# # !pip install simpletransformers
# + id="KlDh5wCeiOYs"
# Config Details
import transformers
import torch
from tqdm import tqdm
import torch.nn as nn
import pandas as pd
import numpy as np
import joblib
import torch
from sklearn import preprocessing
from sklearn import model_selection
from transformers import AdamW
from transformers import get_linear_schedule_with_warmup
import dataset
MAX_LEN = 128
TRAIN_BATCH_SIZE = 32
VALID_BATCH_SIZE = 8
EPOCHS = 8
#BASE_MODEL_PATH = "microsoft/BiomedNLP-PubMedBERT-base-uncased-abstract-fulltext"
BASE_MODEL_PATH = "bert-base-uncased"
#TOKENIZER_PATH = "microsoft/BiomedNLP-PubMedBERT-base-uncased-abstract-fulltext"
# TOKENIZER_PATH = "/content/drive/MyDrive/bert-fine-tuned_version_2/tokenizer"
MODEL_PATH ="/content/drive/MyDrive/bert/train_model_save/model.bin"
TRAINING_FILE = "/content/drive/My Drive/bert/dataset/BIO_taggingdata_ALL_Spacy_TRAIN.csv"
TOKENIZER = transformers.BertTokenizer.from_pretrained(
BASE_MODEL_PATH,
do_lower_case=True
)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} executionInfo={"elapsed": 1555, "status": "error", "timestamp": 1623923841999, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhSasqlkSGouVaJ4_6HwZYs3ccarzD6XSfX3qG7=s64", "userId": "11001256407696526704"}, "user_tz": -330} id="215QQOJ-ZYxj" outputId="7a6d5be5-6cab-410e-ff57-9fdc3e419603"
# model = torch.load('/content/drive/MyDrive/bert/model_2000_manually_ROW_WISE')
# + id="6xG0tmIIMf2S"
# #!cp model.bin /content/drive/MyDrive/model_ner_ft_2.bin
# #!cp meta.bin /content/drive/MyDrive/meta_ft_2.bin
# + id="OERDUU2OuRbH"
# # !pip install transformers
# # !pip install dataset
# + [markdown] id="rl-ukxNl79pA"
# ## Entity Dataset, Entity Model & Training Function
# + id="gwb7zkgxe7Ly"
class EntityDataset:
def __init__(self, texts, tags):
# texts: [["hi", ",", "my", "name", "is", "abhishek"], ["hello".....]]
# pos/tags: [[1 2 3 4 1 5], [....].....]]
self.texts = texts
#self.pos = pos
self.tags = tags
def __len__(self):
return len(self.texts)
def __getitem__(self, item):
text = self.texts[item]
#pos = self.pos[item]
tags = self.tags[item]
ids = []
#target_pos = []
target_tag =[]
for i, s in enumerate(text):
inputs = TOKENIZER.encode(
s,
add_special_tokens=False
)
# abhishek: ab ##hi ##sh ##ek
input_len = len(inputs)
ids.extend(inputs)
#target_pos.extend([pos[i]] * input_len)
target_tag.extend([tags[i]] * input_len)
ids = ids[:MAX_LEN - 2]
# for CLS/SEP tokens
#target_pos = target_pos[:config.MAX_LEN - 2]
target_tag = target_tag[:MAX_LEN - 2]
ids = [2] + ids + [3]
#target_pos = [0] + target_pos + [0]
target_tag = [0] + target_tag + [0]
mask = [1] * len(ids)
token_type_ids = [0] * len(ids)
padding_len = MAX_LEN - len(ids)
ids = ids + ([0] * padding_len)
mask = mask + ([0] * padding_len)
token_type_ids = token_type_ids + ([0] * padding_len)
#target_pos = target_pos + ([0] * padding_len)
target_tag = target_tag + ([0] * padding_len)
return {
"ids": torch.tensor(ids, dtype=torch.long),
"mask": torch.tensor(mask, dtype=torch.long),
"token_type_ids": torch.tensor(token_type_ids, dtype=torch.long),
"target_tag": torch.tensor(target_tag, dtype=torch.long),
}
# + id="HL0pe_7gh-Re"
def train_fn(data_loader, model, optimizer, device, scheduler):
model.train()
final_loss = 0
for data in tqdm(data_loader, total=len(data_loader)):
for k, v in data.items():
data[k] = v.to(device)
optimizer.zero_grad()
_, loss = model(**data)
loss.backward()
optimizer.step()
scheduler.step()
final_loss += loss.item()
return final_loss / len(data_loader)
def eval_fn(data_loader, model, device):
model.eval()
final_loss = 0
for data in tqdm(data_loader, total=len(data_loader)):
for k, v in data.items():
data[k] = v.to(device)
_, loss = model(**data)
final_loss += loss.item()
return final_loss / len(data_loader)
# + id="fAdjBsTbh_Tl"
def loss_fn(output, target, mask, num_labels):
lfn = nn.CrossEntropyLoss()
active_loss = mask.view(-1) == 1
active_logits = output.view(-1, num_labels)
active_labels = torch.where(
active_loss,
target.view(-1),
torch.tensor(lfn.ignore_index).type_as(target)
)
loss = lfn(active_logits, active_labels)
return loss
class EntityModel(nn.Module):
def __init__(self, num_tag):
super(EntityModel, self).__init__()
self.num_tag = num_tag
self.bert = transformers.BertModel.from_pretrained(BASE_MODEL_PATH,return_dict=False)
self.bert_drop_1 = nn.Dropout(0.3)
#self.bert_drop_2 = nn.Dropout(0.3)
self.out_tag = nn.Linear(768, self.num_tag)
def forward(self, ids, mask, token_type_ids, target_tag):
o1, _ = self.bert(ids, attention_mask=mask, token_type_ids=token_type_ids)
bo_tag = self.bert_drop_1(o1)
#bo_pos = self.bert_drop_2(o1)
tag = self.out_tag(bo_tag)
#pos = self.out_pos(bo_pos)
loss_tag = loss_fn(tag, target_tag, mask, self.num_tag)
#loss_pos = loss_fn(pos, target_pos, mask, self.num_pos)
#loss = (loss_tag + loss_pos) / 2
loss = loss_tag
return tag, loss
# + id="xOsb2233iLEK"
def process_data(data_path):
df = pd.read_csv(data_path, encoding="latin-1")
df=df.replace(r'^\s*$', np.nan, regex=True)
df.loc[:, "Sentence #"] = df["Sentence #"].fillna(method="ffill")
print('Number of empty values are ', df["Word"].isna().sum())
df["Word"].fillna("None", inplace = True)
df.loc[:, "Tag"] = df["Tag"].fillna(method="ffill")
print("Check 1")
enc_pos = preprocessing.LabelEncoder()
enc_tag = preprocessing.LabelEncoder()
print("Check 2")
#df.loc[:, "POS"] = enc_pos.fit_transform(df["POS"])
df.loc[:, "Tag"] = enc_tag.fit_transform(df["Tag"])
print("Check 3")
sentences = df.groupby("Sentence #")["Word"].apply(list).values
print("Check 4")
#pos = df.groupby("Sentence #")["POS"].apply(list).values
pos = []
tag = df.groupby("Sentence #")["Tag"].apply(list).values
print("Check 5")
return sentences, pos, tag, enc_pos, enc_tag
# + [markdown] id="xRjpiJmR8FZh"
# ## Preprocess Dataset
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 3217, "status": "ok", "timestamp": 1623853464217, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhSasqlkSGouVaJ4_6HwZYs3ccarzD6XSfX3qG7=s64", "userId": "11001256407696526704"}, "user_tz": -330} id="8VxDg-i_vs2B" outputId="9aead34c-5b46-41e5-8553-ab635ad20d34"
sentences, pos, tag, enc_pos, enc_tag = process_data(TRAINING_FILE)
print("\nsentences:\n", sentences)
print("\n pos:\n", pos)
print("\n tag:\n", tag)
print("\n enc_pos:\n", enc_pos)
print("\n enc_tag:\n", enc_tag)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 7, "status": "ok", "timestamp": 1623853533822, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhSasqlkSGouVaJ4_6HwZYs3ccarzD6XSfX3qG7=s64", "userId": "11001256407696526704"}, "user_tz": -330} id="IWTURzDFwgLj" outputId="df63138a-7fa2-4227-ec44-e3a441bcf3ed"
meta_data = {
"enc_pos": enc_pos,
"enc_tag": enc_tag
}
joblib.dump(meta_data, "meta.bin")
# + id="Ln16zDgHwow0"
num_tag = len(list(enc_tag.classes_))
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 5, "status": "ok", "timestamp": 1623853537027, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhSasqlkSGouVaJ4_6HwZYs3ccarzD6XSfX3qG7=s64", "userId": "11001256407696526704"}, "user_tz": -330} id="Dl8ePtskExoP" outputId="efe2f93d-c59f-475c-fa65-08fc7fbbd6a2"
print(num_tag)
# + id="31pUpP3MwvWy"
(train_sentences, test_sentences, train_tag,test_tag) = model_selection.train_test_split(sentences, tag, random_state=42, test_size=0.2)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 2218, "status": "ok", "timestamp": 1623790231448, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhSasqlkSGouVaJ4_6HwZYs3ccarzD6XSfX3qG7=s64", "userId": "11001256407696526704"}, "user_tz": -330} id="y8yQtgykxAeZ" outputId="1c34fdfb-893f-43e3-eb2a-62ef68e63b2e"
train_dataset = EntityDataset(
texts=train_sentences, tags=train_tag
)
train_data_loader = torch.utils.data.DataLoader(
train_dataset, batch_size=TRAIN_BATCH_SIZE, num_workers=2
)
valid_dataset = EntityDataset(
texts=test_sentences, tags=test_tag
)
valid_data_loader = torch.utils.data.DataLoader(
valid_dataset, batch_size=VALID_BATCH_SIZE, num_workers=1
)
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
model = EntityModel(num_tag=num_tag)
model.to(device)
# + [markdown] id="ZTURpxw68KAW"
# ## Training
# + id="UEh1a4YO9WdF"
#import os
#os.mkdir(MODEL_PATH)
# + id="6SqP7nAxmx-9"
#data
# + id="eBh2OQSAl92n"
# for data in tqdm(train_data_loader, total=len(train_data_loader)):
# print('Next Data Loader')
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 0, "status": "ok", "timestamp": 1623790584065, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhSasqlkSGouVaJ4_6HwZYs3ccarzD6XSfX3qG7=s64", "userId": "11001256407696526704"}, "user_tz": -330} id="pPtan55B0Lni" outputId="e86d3cf0-be77-465c-d064-6fdd7ab8f7f3"
param_optimizer = list(model.named_parameters())
no_decay = ["bias", "LayerNorm.bias", "LayerNorm.weight"]
optimizer_parameters = [
{
"params": [
p for n, p in param_optimizer if not any(nd in n for nd in no_decay)
],
"weight_decay": 0.001,
},
{
"params": [
p for n, p in param_optimizer if any(nd in n for nd in no_decay)
],
"weight_decay": 0.0,
},
]
num_train_steps = int(len(train_sentences) / TRAIN_BATCH_SIZE * EPOCHS)
optimizer = AdamW(optimizer_parameters, lr=3e-5)
scheduler = get_linear_schedule_with_warmup(
optimizer, num_warmup_steps=0, num_training_steps=num_train_steps
)
best_loss = np.inf
for epoch in range(EPOCHS):
train_loss = train_fn(train_data_loader, model, optimizer, device, scheduler)
test_loss = eval_fn(valid_data_loader, model, device)
print(f"Train Loss = {train_loss} Valid Loss = {test_loss}")
if test_loss < best_loss:
torch.save(model.state_dict(), MODEL_PATH)
best_loss = test_loss
# + [markdown] id="fYjT0RRv8Tj-"
# ## Prediction
# + colab={"base_uri": "https://localhost:8080/", "height": 351} executionInfo={"elapsed": 379, "status": "error", "timestamp": 1623853484536, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhSasqlkSGouVaJ4_6HwZYs3ccarzD6XSfX3qG7=s64", "userId": "11001256407696526704"}, "user_tz": -330} id="UAIbaxjH7pmh" outputId="530b4744-b5de-4b0f-cc14-2cb5220b9b32"
meta_data = joblib.load("meta.bin")
enc_tag = meta_data["enc_tag"]
#num_pos = len(list(enc_pos.classes_))
num_tag = len(list(enc_tag.classes_))
# sentence = "seasoned backend developer. entrepreneur. open source contributor. scalable, highly-available web development: python (django), ruby (rails, sinatra), node.js, go, react js/native, angular, java. web backend scalability and performance tuning: new relic, ruby-prof, cprofile. queue-based solutions: kue, resque/sidekiq, celery, jms, rabbitmq. mobile: swift, objective-c, restkit/afnetworking, coredata, corelocation, gcd, sentestingkit, android studio, play service, retrofit. deep learning: convolutional neural network. test-driven-development: rspec, cucumber, python unittest, junit, jasmine. continuous integration/delivery: travis, jenkins, capistrano, vagrant, git, subversion, rake, maven, ant, buildout, make/gnu make. cloud/container: aws, azure, docker, docker-compose, dcos, kubernetes. devops: mesos, chef, puppet, mcollective, pxe, ipmi, nagios, zabbix. scripting: bash, python, ruby, perl. open source projects that enjoy 200+ stars on github and 100+k downloads on sourceforge., seasoned backend developer and entrepreneur.\n\nspecialties: \nscalable, highly-available web development: java, ruby (rails, sinatra), python (django), node.js.\nqueue-based solutions: resque/sidekiq, celery, jms, rabbitmq.\nweb backend performance tuning.\nios: coredata, corelocation, gcd, restkit/afnetworking, sentestingkit, swift.\ntest-driven-development: rspec, cucumber, python unittest, junit.\ncontinuous integration/delivery: jenkins, chef, capistrano, vagrant, git, subversion, rake, maven, ant, buildout, make/gnu make.\ninfrastructure-as-a-service: openstack nova, aws ec2.\nplatform-as-a-service: cloudfoundry, heroku, rightscale, enstratus, scalr, juju.\ncloud storage: hadoop hdfs, aws s3, openstack swift, mongodb.\nagile methodologies: scrum, fdd (feature-driven-development).\ndevops: chef, puppet, mcollective, pxe, ipmi, nagios, zabbix.\nscripting: bash, python, ruby, perl., programmer. entrepreneur at banian labs, pdh - networking/network engineering, pdh - network planner/provisioning, vp engineering at rhumbix"
# tokenized_sentence = TOKENIZER.encode(sentence)
# sentence = sentence.split()
# print(sentence)
# print(tokenized_sentence)
# test_dataset = EntityDataset(
# texts=[sentence],
# tags=[[0] * len(sentence)]
# )
# device = torch.device("cuda")
# pred_model = EntityModel(num_tag=num_tag)
# pred_model.load_state_dict(torch.load(MODEL_PATH))
# pred_model.to(device)
# with torch.no_grad():
# data = test_dataset[0]
# for k, v in data.items():
# data[k] = v.to(device).unsqueeze(0)
# tag, _ = pred_model(**data)
# print(
# enc_tag.inverse_transform(
# tag.argmax(2).cpu().numpy().reshape(-1)
# )[:len(tokenized_sentence)]
# )
# # print(
# # enc_pos.inverse_transform(
# # pos.argmax(2).cpu().numpy().reshape(-1)
# # )[:len(tokenized_sentence)]
# # )
# + id="P6P1VHARLOz-"
device = torch.device("cuda")
# #
# train_dataset = EntityDataset(
# texts=train_sentences, tags=train_tag
# )
# train_data_loader = torch.utils.data.DataLoader(
# train_dataset, batch_size=TRAIN_BATCH_SIZE, num_workers=2
# )
# valid_dataset = EntityDataset(
# texts=test_sentences, tags=test_tag
# )
# valid_data_loader = torch.utils.data.DataLoader(
# valid_dataset, batch_size=VALID_BATCH_SIZE, num_workers=1
# )
# device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# model = EntityModel(num_tag=num_tag)
# model.to(device)
# #
for i in df["summaries"]:
sentence = i
tokenized_sentence = TOKENIZER.encode(sentence)
sentence = sentence.split()
print("******** sentence ********", sentence)
print("******** tokenised sentence ********", tokenized_sentence)
test_dataset = EntityDataset(
texts=[sentence],
tags=[[0] * len(sentence)]
)
pred_model = EntityModel(num_tag=num_tag)
pred_model.load_state_dict(torch.load(MODEL_PATH))
pred_model.to(device)
with torch.no_grad():
data = test_dataset[0]
for k, v in data.items():
data[k] = v.to(device).unsqueeze(0)
tag, _ = pred_model(**data)
preds = enc_tag.inverse_transform(
tag.argmax(2).cpu().numpy().reshape(-1)
)[:len(tokenized_sentence)]
for elem,cat in zip(tokenized_sentence , preds):
print(TOKENIZER.decode([elem]), '=====>',cat)
# sentence = "seasoned backend developer. entrepreneur. open source contributor. scalable, highly-available web development: python (django), ruby (rails, sinatra), node.js, go, react js/native, angular, java. web backend scalability and performance tuning: new relic, ruby-prof, cprofile. queue-based solutions: kue, resque/sidekiq, celery, jms, rabbitmq. mobile: swift, objective-c, restkit/afnetworking, coredata, corelocation, gcd, sentestingkit, android studio, play service, retrofit. deep learning: convolutional neural network. test-driven-development: rspec, cucumber, python unittest, junit, jasmine. continuous integration/delivery: travis, jenkins, capistrano, vagrant, git, subversion, rake, maven, ant, buildout, make/gnu make. cloud/container: aws, azure, docker, docker-compose, dcos, kubernetes. devops: mesos, chef, puppet, mcollective, pxe, ipmi, nagios, zabbix. scripting: bash, python, ruby, perl. open source projects that enjoy 200+ stars on github and 100+k downloads on sourceforge., seasoned backend developer and entrepreneur.\n\nspecialties: \nscalable, highly-available web development: java, ruby (rails, sinatra), python (django), node.js.\nqueue-based solutions: resque/sidekiq, celery, jms, rabbitmq.\nweb backend performance tuning.\nios: coredata, corelocation, gcd, restkit/afnetworking, sentestingkit, swift.\ntest-driven-development: rspec, cucumber, python unittest, junit.\ncontinuous integration/delivery: jenkins, chef, capistrano, vagrant, git, subversion, rake, maven, ant, buildout, make/gnu make.\ninfrastructure-as-a-service: openstack nova, aws ec2.\nplatform-as-a-service: cloudfoundry, heroku, rightscale, enstratus, scalr, juju.\ncloud storage: hadoop hdfs, aws s3, openstack swift, mongodb.\nagile methodologies: scrum, fdd (feature-driven-development).\ndevops: chef, puppet, mcollective, pxe, ipmi, nagios, zabbix.\nscripting: bash, python, ruby, perl., programmer. entrepreneur at banian labs, pdh - networking/network engineering, pdh - network planner/provisioning, vp engineering at rhumbix"
# tokenized_sentence = TOKENIZER.encode(sentence)
# sentence = sentence.split()
# print(sentence)
# print(tokenized_sentence)
# test_dataset = EntityDataset(
# texts=[sentence],
# tags=[[0] * len(sentence)]
# )
# device = torch.device("cuda")
# pred_model = EntityModel(num_tag=num_tag)
# pred_model.load_state_dict(torch.load(MODEL_PATH))
# pred_model.to(device)
# with torch.no_grad():
# data = test_dataset[0]
# for k, v in data.items():
# data[k] = v.to(device).unsqueeze(0)
# tag, _ = pred_model(**data)
# print(
# enc_tag.inverse_transform(
# tag.argmax(2).cpu().numpy().reshape(-1)
# )[:len(tokenized_sentence)]
# )
# # print(
# # enc_pos.inverse_transform(
# # pos.argmax(2).cpu().numpy().reshape(-1)
# # )[:len(tokenized_sentence)]
# # )
# + id="cyBRv7AxQSHA"
preds = enc_tag.inverse_transform(
tag.argmax(2).cpu().numpy().reshape(-1)
)[:len(tokenized_sentence)]
# + colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"elapsed": 21, "status": "ok", "timestamp": 1623790891871, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhSasqlkSGouVaJ4_6HwZYs3ccarzD6XSfX3qG7=s64", "userId": "11001256407696526704"}, "user_tz": -330} id="VZS6PPMUStoC" outputId="76e9cd02-3514-4824-d265-6446765852d7"
TOKENIZER.decode(30000)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 7, "status": "ok", "timestamp": 1623790892858, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhSasqlkSGouVaJ4_6HwZYs3ccarzD6XSfX3qG7=s64", "userId": "11001256407696526704"}, "user_tz": -330} id="zS_gbgI7RmuQ" outputId="71cf2dcb-fd61-49bd-b2ec-1c11c92b8856"
for elem,cat in zip(tokenized_sentence , preds):
print(TOKENIZER.decode([elem]), '=====>',cat)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 422, "status": "ok", "timestamp": 1623790895886, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhSasqlkSGouVaJ4_6HwZYs3ccarzD6XSfX3qG7=s64", "userId": "11001256407696526704"}, "user_tz": -330} id="CuoxJB45QKSq" outputId="fd39af4b-f621-4584-b8e9-e2920688aed4"
tag.argmax(2)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 7, "status": "ok", "timestamp": 1623790897816, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhSasqlkSGouVaJ4_6HwZYs3ccarzD6XSfX3qG7=s64", "userId": "11001256407696526704"}, "user_tz": -330} id="h5oM8jVDP_n1" outputId="c42a5cc3-ebca-40ee-9355-2477693261c2"
tokenized_sentence
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 13, "status": "ok", "timestamp": 1623790898307, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhSasqlkSGouVaJ4_6HwZYs3ccarzD6XSfX3qG7=s64", "userId": "11001256407696526704"}, "user_tz": -330} id="bq1B3p7XQB8C" outputId="b49d51e2-d98c-4efe-8875-78dd8fee9875"
enc_tag
# + colab={"base_uri": "https://localhost:8080/", "height": 171} executionInfo={"elapsed": 11, "status": "ok", "timestamp": 1623790898308, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhSasqlkSGouVaJ4_6HwZYs3ccarzD6XSfX3qG7=s64", "userId": "11001256407696526704"}, "user_tz": -330} id="wwN0iNDePPNL" outputId="a9e40a96-1fcd-42ab-ee6b-e08c43bdece4"
TOKENIZER.decode(tokenized_sentence)
# + id="5dw-jR1ovpCi"
# if __name__ == "__main__":
# sentences, pos, tag, enc_pos, enc_tag = process_data(config.TRAINING_FILE)
# meta_data = {
# "enc_pos": enc_pos,
# "enc_tag": enc_tag
# }
# joblib.dump(meta_data, "meta.bin")
# num_pos = len(list(enc_pos.classes_))
# num_tag = len(list(enc_tag.classes_))
# (
# train_sentences,
# test_sentences,
# train_pos,
# test_pos,
# train_tag,
# test_tag
# ) = model_selection.train_test_split(sentences, pos, tag, random_state=42, test_size=0.1)
# train_dataset = dataset.EntityDataset(
# texts=train_sentences, pos=train_pos, tags=train_tag
# )
# train_data_loader = torch.utils.data.DataLoader(
# train_dataset, batch_size=config.TRAIN_BATCH_SIZE, num_workers=4
# )
# valid_dataset = dataset.EntityDataset(
# texts=test_sentences, pos=test_pos, tags=test_tag
# )
# valid_data_loader = torch.utils.data.DataLoader(
# valid_dataset, batch_size=config.VALID_BATCH_SIZE, num_workers=1
# )
# device = torch.device("cuda")
# model = EntityModel(num_tag=num_tag, num_pos=num_pos)
# model.to(device)
# param_optimizer = list(model.named_parameters())
# no_decay = ["bias", "LayerNorm.bias", "LayerNorm.weight"]
# optimizer_parameters = [
# {
# "params": [
# p for n, p in param_optimizer if not any(nd in n for nd in no_decay)
# ],
# "weight_decay": 0.001,
# },
# {
# "params": [
# p for n, p in param_optimizer if any(nd in n for nd in no_decay)
# ],
# "weight_decay": 0.0,
# },
# ]
# num_train_steps = int(len(train_sentences) / config.TRAIN_BATCH_SIZE * config.EPOCHS)
# optimizer = AdamW(optimizer_parameters, lr=3e-5)
# scheduler = get_linear_schedule_with_warmup(
# optimizer, num_warmup_steps=0, num_training_steps=num_train_steps
# )
# best_loss = np.inf
# for epoch in range(config.EPOCHS):
# train_loss = engine.train_fn(train_data_loader, model, optimizer, device, scheduler)
# test_loss = engine.eval_fn(valid_data_loader, model, device)
# print(f"Train Loss = {train_loss} Valid Loss = {test_loss}")
# if test_loss < best_loss:
# torch.save(model.state_dict(), config.MODEL_PATH)
# best_loss = test_loss
# + id="0uFYI8VP3qrq"
| GPU Notebooks/BERT_Scratch/Train_NER_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: keras-gpu
# language: python
# name: keras-gpu
# ---
# # 6장. 케라스2로 구현하는 AE(오토인코더)
# ## 6.2 완전 연결 계층을 이용한 AE 구현
# 필기체 숫자 즉 MNIST를 이용해 AE을 구현
# set to use CPU
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '-1'
# ### 6.2.1 완전 연결 계층 AE 모델링
# 1. AE를 위한 딥러닝 모델을 만듭니다.
from keras import layers, models
class AE(models.Model):
def __init__(self, x_nodes=784, z_dim=36):
x_shape = (x_nodes,)
x = layers.Input(shape=x_shape)
z = layers.Dense(z_dim, activation='relu')(x)
y = layers.Dense(x_nodes, activation='sigmoid')(z)
super().__init__(x, y)
self.x = x
self.z = z
self.z_dim = z_dim
# Encoder, Decoder ??
self.compile(optimizer='adadelta', loss='binary_crossentropy', metrics=['accuracy'])
def Encoder(self):
return models.Model(self.x, self.z)
def Decoder(self):
z_shape = (self.z_dim,)
z = layers.Input(shape=z_shape)
y_layer = self.layers[-1]
y = y_layer(z)
return models.Model(z, y)
# ### 6.2.2 데이터 준비
# 2. 사용할 MNIST 데이터를 케라스의 서브패키지로부터 불러옵니다
# +
from keras.datasets import mnist
import numpy as np
(X_train, _), (X_test, _) = mnist.load_data()
X_train = X_train.astype('float32') / 255.
X_test = X_test.astype('float32') / 255.
X_train = X_train.reshape((len(X_train), np.prod(X_train.shape[1:])))
X_test = X_test.reshape((len(X_test), np.prod(X_test.shape[1:])))
print(X_train.shape)
print(X_test.shape)
# -
# ### 6.2.3 학습 효과 분석
# 3. 학습 효과를 그래프로 분석하기 위해 필요한 함수들과 패키지를 불러옵니다
from keraspp.skeras import plot_loss, plot_acc
import matplotlib.pyplot as plt
# ### 6.2.4 완전 연결 계층 AE 동작 확인
# 4. 완전 연결 계층 AE가 어떻게 동작했는지 확인합시다
def show_ae(autoencoder):
encoder = autoencoder.Encoder()
decoder = autoencoder.Decoder()
encoded_imgs = encoder.predict(X_test)
decoded_imgs = decoder.predict(encoded_imgs)
n = 10
plt.figure(figsize=(20, 6))
for i in range(n):
ax = plt.subplot(3, n, i + 1)
plt.imshow(X_test[i].reshape(28, 28))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
ax = plt.subplot(3, n, i + 1 + n)
plt.stem(encoded_imgs[i].reshape(-1))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
ax = plt.subplot(3, n, i + 1 + n + n)
plt.imshow(decoded_imgs[i].reshape(28, 28))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()
# ### 6.2.5 학습 및 성능 평가
# 5. 학습과 성능 평가에 사용할 main() 함수를 만듭니다.
def main():
x_nodes = 784
z_dim = 36
autoencoder = AE(x_nodes, z_dim)
history = autoencoder.fit(X_train, X_train,
epochs=10,
batch_size=256,
shuffle=True,
validation_data=(X_test, X_test))
plot_acc(history, 'Accuracy')
plt.show()
plot_loss(history, 'Loss')
plt.show()
show_ae(autoencoder)
plt.show()
main()
# ---
# ### 6.2.6 전체 코드
# +
###########################
# AE 모델링
###########################
from keras import layers, models # (Input, Dense), (Model)
class AE(models.Model):
def __init__(self, x_nodes=784, z_dim=36):
x_shape = (x_nodes,)
x = layers.Input(shape=x_shape)
z = layers.Dense(z_dim, activation='relu')(x)
y = layers.Dense(x_nodes, activation='sigmoid')(z)
super().__init__(x, y)
self.x = x
self.z = z
self.z_dim = z_dim
# Encoder, Decoder ??
self.compile(optimizer='adadelta', loss='binary_crossentropy', metrics=['accuracy'])
def Encoder(self):
return models.Model(self.x, self.z)
def Decoder(self):
z_shape = (self.z_dim,)
z = layers.Input(shape=z_shape)
y_layer = self.layers[-1]
y = y_layer(z)
return models.Model(z, y)
###########################
# 데이터 준비
###########################
from keras.datasets import mnist
import numpy as np
(X_train, _), (X_test, _) = mnist.load_data()
X_train = X_train.astype('float32') / 255.
X_test = X_test.astype('float32') / 255.
X_train = X_train.reshape((len(X_train), np.prod(X_train.shape[1:])))
X_test = X_test.reshape((len(X_test), np.prod(X_test.shape[1:])))
print(X_train.shape)
print(X_test.shape)
###########################
# 학습 효과 분석
###########################
from keraspp.skeras import plot_loss, plot_acc
import matplotlib.pyplot as plt
###########################
# AE 동작 확인
###########################
def show_ae(autoencoder):
encoder = autoencoder.Encoder()
decoder = autoencoder.Decoder()
encoded_imgs = encoder.predict(X_test)
decoded_imgs = decoder.predict(encoded_imgs)
n = 10
plt.figure(figsize=(20, 6))
for i in range(n):
ax = plt.subplot(3, n, i + 1)
plt.imshow(X_test[i].reshape(28, 28))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
ax = plt.subplot(3, n, i + 1 + n)
plt.stem(encoded_imgs[i].reshape(-1))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
ax = plt.subplot(3, n, i + 1 + n + n)
plt.imshow(decoded_imgs[i].reshape(28, 28))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()
###########################
# 학습
###########################
def main():
x_nodes = 784
z_dim = 36
autoencoder = AE(x_nodes, z_dim)
history = autoencoder.fit(X_train, X_train,
epochs=10,
batch_size=256,
shuffle=True,
validation_data=(X_test, X_test))
plot_acc(history, '(a) 학습 경과에 따른 정확도 변화 추이')
plt.show()
plot_loss(history, '(b) 학습 경과에 따른 손실값 변화 추이')
plt.show()
show_ae(autoencoder)
plt.show()
main()
# -
| cpu_only/nb_ex6_1_ae_fc_mnist-cpu.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # This script is built for implementation of FunkSVD
# +
# #!/usr/bin/python
# -*- coding: UTF-8 -*-
import random
import pandas as pd
import numpy as np
from collections import Counter
seed=15
random.seed(seed)
np.random.seed(seed=seed)
# -
# ## Load Data
df = pd.read_csv("../data/movies/ratings_small.csv")
df = df.drop(["timestamp"], axis=1)
df = df[~df["rating"].isna()]
df.head()
# ## Find Unique Users and Items. Then, Map to Pre-Defined IDs
# +
uniqueUsers = df["userId"].unique()
uniqueItems = df["movieId"].unique()
userNum = len(uniqueUsers)
itemNum = len(uniqueItems)
print(f"Total {userNum} users and {itemNum} items")
# Build mapper for users and items
userMapper = dict(zip(uniqueUsers, range(userNum)))
itemMapper = dict(zip(uniqueItems, range(itemNum)))
df["userId"] = df["userId"].apply(lambda x: userMapper[x])
df["movieId"] = df["movieId"].apply(lambda x: itemMapper[x])
# -
# ## Split Data
# +
validRatingMatrix = df.sample(frac=0.4, random_state=seed)
trainRatingMatrix = df.drop(index=validRatingMatrix.index)
print(f"Train dataframe has shape: {trainRatingMatrix.shape}")
print(f"Valid dataframe has shape: {validRatingMatrix.shape}")
trainRatingMatrix = trainRatingMatrix.to_numpy()
validRatingMatrix = validRatingMatrix.to_numpy()
# -
# ## Shuffle
# +
indices = np.arange(trainRatingMatrix.shape[0])
np.random.shuffle(indices)
trainRatingMatrix = trainRatingMatrix[indices]
# -
# ## Normalize
# +
trainRatings = trainRatingMatrix[:, 2].astype(np.float32)
trainUserItems = trainRatingMatrix[:, :2].astype(np.int)
validRatings = validRatingMatrix[:, 2].astype(np.float32)
validUserItems = validRatingMatrix[:, :2].astype(np.int)
ratingMean, ratingStd = np.mean(trainRatings), np.std(trainRatings)
print(f"Ratings have mean={ratingMean:.3f}, std={ratingStd:.3f}")
trainRatings -= ratingMean
trainRatings /= ratingStd
validRatings -= ratingMean
validRatings /= ratingStd
# -
# ## Hyperparameters
# +
# Set for training
featureNum = 10
lr = 8e-3
decayRate = 0.9
epochs = 50
bestMse = 1e6
# Set for regularization
regRate = 1e-2
# Set for display
showInterval = 10
# -
# ## Train and Validate
# +
userMatrix = np.random.randn(userNum, featureNum).astype(np.float32)
itemMatrix = np.random.randn(itemNum, featureNum).astype(np.float32)
userBias = np.zeros(userNum).astype(np.float32)
itemBias = np.zeros(itemNum).astype(np.float32)
trainMses, trainMaes = [0] * epochs, [0] * epochs
validMses, validMaes = [0] * epochs, [0] * epochs
for epoch in range(epochs):
if epoch > 0 and epoch % 20 == 0:
lr *= decayRate
print(f"***Set learning rate to {lr:.4f}")
# ==========
# Train
# ==========
for ((u, i), r) in zip(trainUserItems, trainRatings):
pred = np.dot(userMatrix[u, :], itemMatrix[i, :]) + userBias[u] + itemBias[i]
diff = r - pred
trainMses[epoch] += diff ** 2
trainMaes[epoch] += abs(diff)
for k in range(featureNum):
userMatrix[u, k] += lr * (diff * itemMatrix[i, k] - regRate * userMatrix[u, k])
itemMatrix[i, k] += lr * (diff * userMatrix[u, k] - regRate * itemMatrix[i, k])
userBias[u] += lr * (diff - regRate * userBias[u])
itemBias[i] += lr * (diff - regRate * itemBias[i])
# ==========
# Validation
# ==========
for ((u, i), r) in zip(validUserItems, validRatings):
pred = np.dot(userMatrix[u, :], itemMatrix[i, :]) + userBias[u] + itemBias[i]
diff = r - pred
validMses[epoch] += diff ** 2
validMaes[epoch] += abs(diff)
trainMses[epoch] /= trainRatings.shape[0]
validMses[epoch] /= trainRatings.shape[0]
trainMaes[epoch] /= validRatings.shape[0]
validMaes[epoch] /= validRatings.shape[0]
# If model is the best, save matrices
isbest = validMses[epoch] < bestMse
bestMse = min(validMses[epoch], bestMse)
if isbest:
with open("funksvd/userMatrix.npy", "wb") as f:
np.save(f, userMatrix)
with open("funksvd/itemMatrix.npy", "wb") as f:
np.save(f, itemMatrix)
print(f"Save the best model at epoch={epoch+1}, mse={validMses[epoch]:.3f}, mae={validMaes[epoch]:.3f}")
# Display stats
if (epoch+1) % showInterval == 0:
print(f"\n{'-' * 50}")
print(f"(Epoch {epoch+1})")
print(f"MSE: train:{trainMses[epoch]:.3f}, valid:{validMses[epoch]:.3f}")
print(f"MAE: train:{trainMaes[epoch]:.3f}, valid:{validMaes[epoch]:.3f}")
print(f"{'-' * 50}\n")
# -
# ## Plot MSE and MAE
# +
# %matplotlib inline
import matplotlib.pyplot as plt
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(18,10))
fig.subplots_adjust(hspace=0.5)
ax1.set_title("MSE")
ax1.plot(range(epochs), trainMses, "x-", color="r", label="Train")
ax1.plot(range(epochs), validMses, "x-", color="b", label="Valid")
ax1.set(xlabel="Epoch", ylabel="Loss")
ax1.legend(loc="upper right")
ax2.set_title("MAE")
ax2.plot(range(epochs), trainMaes, "x-", color="r", label="Train")
ax2.plot(range(epochs), validMaes, "x-", color="b", label="Valid")
ax2.set(xlabel="Epoch", ylabel="Loss")
ax2.legend(loc="upper right")
plt.savefig("funksvd/loss.png")
plt.show()
# -
| funksvd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Varying initial concentrations and parameters
# A version of this notebook may be run online via Google Colab at https://tinyurl.com/rxd-initial-and-parameters
# (make a copy or open in playground mode).
# %matplotlib inline
# Often we will want to see how the choice of initial conditions affects the dynamics. We can do this by setting the initial attribute of an `rxd.Species` and rerunning.
# For example, suppose at a single point we have the bistable dynamics introduced in the first part of this tutorial. That is, $u'=-u(1-u)(\alpha - u)$. (Here we use $u$ instead of a specific molecule name to indicate that we are not describing any particular molecule's kinetics, but rather modeling a class of phenomena.)
# This time, we'll use an `rxd.Parameter` for $\alpha$ instead of a constant. This offers two advantages: (1) this allows $\alpha$ to vary spatially, and (2) this allows us to change the parameter values and rerun without changing the reaction/rate specification.
# We start by defining the model and setting up the recordings:
# +
from neuron import h, rxd
from neuron.units import mV, ms, mM
from matplotlib import pyplot as plt
h.load_file('stdrun.hoc')
soma = h.Section(name='soma')
cyt = rxd.Region([soma], name='cyt', nrn_region='i')
u = rxd.Species(cyt, name='u')
α = rxd.Parameter(cyt, value=0.3 * mM)
rate = rxd.Rate(u, -u * (1 * mM - u) * (α - u))
h.finitialize(-65 * mV)
t = h.Vector().record(h._ref_t)
y = h.Vector().record(soma(0.5)._ref_ui)
# -
# Now we define a function that sets the initial value for ca, runs the simulation, and plots the curves:
def plot_it(ca_init):
u.initial = ca_init
h.finitialize(-65 * mV)
h.continuerun(10 * ms)
plt.plot(t, y)
# And now let's run it a few times and plot it:
# +
for u0 in range(-4 * mM, 15 * mM, 2 * mM):
plot_it(u0 * 0.1)
plt.xlabel('t (ms)')
plt.ylabel('[u] (mM)')
plt.show()
# -
# Here we see the stable fixed points at $0$ and $1$ and how all other trajectories (except for the one starting exactly at $\alpha$, not shown) tend towards one of those values.
# <hr/>
# Alternatively, we might want to know how the kinetics change if we start at a fixed value (say 0.5) and vary $\alpha$ instead. We can do this in the same way as above by setting the `value` property of $\alpha$:
def plot_α(α_init):
u.initial = 0.5 * mM
α.value = α_init
h.finitialize(-65 * mV)
h.continuerun(10 * ms)
plt.gca().plot(t, y, label=f'α = {α_init} mM')
# Here we have added a `label` argument to the plot and explicitly specify the `gca()` axes to allow displaying a legend; see the <a href="https://matplotlib.org/api/_as_gen/matplotlib.pyplot.legend.html">matplotlib.pyplot.legend</a> documentation for more.
#
# Now we run it:
# +
for my_α in [0 * mM, 0.2 * mM, 0.4 * mM, 0.6 * mM, 0.8 * mM, 1 * mM]:
plot_α(my_α)
plt.gca().legend()
plt.xlabel('t (ms)')
plt.ylabel('[u] (mM)')
plt.show()
| docs/rxd-tutorials/Changing initial conditions and parameters.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # <NAME> - A53213478
# Code for 250A - HW1
#
# 10-10-2017
# # Hangman
__author__ = "<NAME>"
import string
import operator
from collections import defaultdict
# ## Reads data
r = open('hw1_word_counts_05.txt','r')
r.seek(0)
corpus = r.readlines()
# ## Computes prior probabilities
cosplit = [i.strip().split() for i in corpus]
word_count = defaultdict(int)
for i,j in cosplit:
word_count[i]=int(j)
word_probabilities = word_count.copy()
sumcounts = sum(word_count.values())
for i in word_probabilities:
word_probabilities[i]=word_probabilities[i]*1.0/sumcounts
print "Total #unique words : ",len(word_count),"\nTotal words : ",sumcounts
sorted_dict = sorted(word_count.items(), key=operator.itemgetter(1))
print "14 least frequent 5-letter words along with their counts:"
for i in sorted_dict[:14]:
print i[0],i[1]
print "\n15 most frequent 5-letter words along with their counts :"
for i in sorted_dict[-15:]:
print i[0],i[1]
# The above results do make sense, since we see that the least frequent words are mostly typos or proper nouns, and the most frequent words are numbers and pronouns, which is exactly what we can expect in a large corpus of words.
# ## Creates place-holder for included and excluded words
def create_given():
for word in word_count :
for i in range(5) :
if ((correct[i]!= word[i]) and (correct[i]!=' ')):
exclude[word] = word_count[word]
break
for word in word_count :
dele = 0
for a in range(5) :
for b in range(len(inc[a])) :
if inc[a][b]==word[a] :
exclude[word] = word_count[word]
dele = 1
break
if dele == 1:
dele = 0
break
for z in word_count :
if z not in exclude :
given[z] = word_count[z]
# ## Filtering out unwanted words for ease in probability calculation
def create_incorrect():
for x in range(len(incorrect)):
for y in range(5) :
if y in inc:
inc[y].append(incorrect[x])
else :
inc[y] = [incorrect[x]]
for m in range(len(correct)) :
for n in range(5) :
if m!=n and correct[m]!=correct[n] :
if n in inc:
if correct[m]!= ' ':
inc[n].append(correct[m])
else :
if correct[m]!= ' ':
inc[n] = [correct[m]]
# ## Next guess probability computation
def best_next_guess():
prob_evi_given_word = {}
for word in word_count :
if word in given :
prob_evi_given_word[word] = 1
else :
prob_evi_given_word[word] = 0
denominator_1 = sum((prob_evi_given_word[word] * word_probabilities[word]) for word in word_count)
prob_word_given_evi = {}
for word in word_count :
prob_word_given_evi[word] = float((prob_evi_given_word[word]*word_probabilities[word])/denominator_1)
alphabet = {'A':0, 'B':0, 'C':0, 'D':0, 'E':0, 'F':0, 'G':0, 'H':0, 'I':0, 'J':0, 'K':0, 'L':0, 'M':0, 'N':0, 'O':0, 'P':0, 'Q':0, 'R':0, 'S':0, 'T':0, 'U':0, 'V':0, 'W':0, 'X':0, 'Y':0, 'Z':0}
for word in word_count:
for letter in alphabet :
if letter in word :
alphabet[letter]+=prob_word_given_evi[word]
for x in correct :
if x in alphabet :
del alphabet[x]
alphabet_sort = sorted(alphabet.items(), key=operator.itemgetter(1))
print("Next Best Guess is ",alphabet_sort[-1][0])
print("Probability of that is ",float(alphabet_sort[-1][1]))
# # Results for 1.9(b)
for i in range(0,9):
inc = {0:[],1:[],2:[],3:[],4:[]}
exclude,given = {},{}
correct = []
incorrect=[]
while len(correct)!=5 :
correct = raw_input ('\n Enter correct characters (Enter 5 characters and use Space to denote if character not filled till now :)')
if len(correct)!=5 :
print("Correct Character Length not 5, Please try again")
incorrect = raw_input('Enter incorrect characters : ')
create_incorrect()
create_given()
best_next_guess()
| CSE 250A/hw1/Code/CSE 250A - HW1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''.venv'': venv)'
# name: python3
# ---
# # Update Model Parameters API Examples
#
# This demonstrates how to use the [update_model_parameters](https://siliconlabs.github.io/mltk/docs/python_api/operations.html#update_model_parameters) API.
#
# Refer to the [Model Parameters](https://siliconlabs.github.io/mltk/docs/guides/model_parameters.html) guide for more details.
#
# __NOTES:__
#
# - Click here: [](https://colab.research.google.com/github/siliconlabs/mltk/blob/master/mltk/examples/update_params.ipynb) to run this example interactively in your browser
# - Refer to the [Notebook Examples Guide](https://siliconlabs.github.io/mltk/docs/guides/notebook_examples_guide.html) for how to run this example locally in VSCode
# ## Install MLTK Python Package
# Install the MLTK Python package (if necessary)
# !pip install --upgrade silabs-mltk
# ## Import Python Packages
# Import the necessary MLTK APIs
from mltk.core import update_model_parameters, summarize_model
# ## Example 1: Update model specification
#
# The most common use case of the [update_model_parameters](https://siliconlabs.github.io/mltk/docs/python_api/operations.html#update_model_parameters) API is:
# 1. Fully [train](https://siliconlabs.github.io/mltk/docs/guides/model_training.html) a model
# 2. Later modify the [model specification](https://siliconlabs.github.io/mltk/docs/guides/model_specification.html) script with additional parameters
# 3. Run the [update_model_parameters](https://siliconlabs.github.io/mltk/docs/python_api/operations.html#update_model_parameters) API to update the `.tflite` model file in the [model archive](https://siliconlabs.github.io/mltk/docs/guides/model_archive.html).
#
# In this example, it's assumed that the [MltkModel.model_parameters](https://siliconlabs.github.io/mltk/docs/python_api/core/mltk_model.html#mltk.core.MltkModel.model_parameters) settings in
# the [tflite_micro_speech](https://siliconlabs.github.io/mltk/docs/python_api/models/tflite_micro/tflite_micro_speech.html) model specification script have been modified _after_ the model have been
# trained.
#
# ```python
# my_model.model_parameters['average_window_duration_ms'] = 1000
# my_model.model_parameters['detection_threshold'] = 185
# my_model.model_parameters['suppression_ms'] = 1500
# my_model.model_parameters['minimum_count'] = 3
# my_model.model_parameters['volume_db'] = 5.0
# my_model.model_parameters['latency_ms'] = 0
# my_model.model_parameters['log_level'] = 'info'
# ```
#
# After this API completes, the `tflite_micro_speech.mltk.zip` model archive is updated with a new `tflite_micro_speech.tflite` model file.
# Note that _only_ the parameters in the `.tflite`'s metadata section are modified. The model weights and layers are untouched.
# +
# Update the model parameters
update_model_parameters('tflite_micro_speech')
# Generate a summary of the updated model with new parameters
print(summarize_model('tflite_micro_speech'))
# -
# ## Example 2: Update with argument
#
# The [update_model_parameters](https://siliconlabs.github.io/mltk/docs/python_api/operations.html#update_model_parameters) API also works with `.tflite` model files generated outside of the MLTK.
# In this mode, model parameters can be supplied via API argument, e.g.:
# +
import os
import tempfile
import urllib
import shutil
# Use .tflite mode found here:
# https://github.com/mlcommons/tiny/tree/master/benchmark/training/keyword_spotting/trained_models
# NOTE: Update this URL to point to your model if necessary
TFLITE_MODEL_URL = 'https://github.com/mlcommons/tiny/raw/master/benchmark/training/keyword_spotting/trained_models/kws_ref_model.tflite'
# Download the .tflite file and save to the temp dir
external_tflite_path = os.path.normpath(f'{tempfile.gettempdir()}/kws_ref_model.tflite')
with open(external_tflite_path, 'wb') as dst:
with urllib.request.urlopen(TFLITE_MODEL_URL) as src:
shutil.copyfileobj(src, dst)
# +
# Set the parameters in a Python dictionary
parameters = {
"volume": 10.0,
"log_level": "debug",
"threshold": 43
}
# Update the model parameters
update_model_parameters(external_tflite_path, params=parameters)
# Generate a summary of the updated model with new parameters
print(summarize_model(external_tflite_path))
# -
# ## Example 3: Return TfliteModel instance
#
# Rather than update the given model, the [update_model_parameters](https://siliconlabs.github.io/mltk/docs/python_api/operations.html#update-model-parameters) API can also return a [TfliteModel](https://siliconlabs.github.io/mltk/docs/python_api/core/tflite_model.html) instance with the updated parameters.
#
# This is done by specifying the `output='tflite_model'` API argument:
# +
# Set the parameters in a Python dictionary
parameters = {
"volume": 10.0,
"log_level": "debug",
"threshold": 43
}
# Generate a TfliteModel instance with the given parameters
# NOTE: The input external_tflite_path file is NOT modified
tflite_model = update_model_parameters(external_tflite_path, params=parameters, output='tflite_model')
# Generate a summary of the returned TfliteModel instance
print(summarize_model(tflite_model))
| mltk/examples/update_params.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Biogeme Basics: Probit Model
import pandas as pd
import numpy as np
import biogeme.database as db
import biogeme.biogeme as bio
import biogeme.models as models
import biogeme.optimization as opt
from biogeme.expressions import Beta, DefineVariable, bioNormalCdf, log, Elem
import seaborn as sns
import matplotlib.pyplot as plt
# **Import Swissmetro data**
pandas = pd.read_csv("../../Data/6-Discrete Choice Models/swissmetro.dat",sep='\t')
database = db.Database("data/swissmetro", pandas)
# **Use collumn names as variables**
globals().update(database.variables)
# **Exclude some unwanted entries**
#
# Rmove observations where Swissmetro was chosen (CHOICE == 2) and also remove observations where one of the two alternatives is not available
CAR_AV_SP = DefineVariable('CAR_AV_SP',CAR_AV * (SP != 0),database)
TRAIN_AV_SP = DefineVariable('TRAIN_AV_SP',TRAIN_AV * (SP != 0),database)
exclude = (TRAIN_AV_SP == 0) + (CAR_AV_SP == 0) + ( CHOICE == 2 ) + (( PURPOSE != 1 ) * ( PURPOSE != 3 ) + ( CHOICE == 0 )) > 0
database.remove(exclude)
# **Define some dummy variables**
SM_COST = SM_CO * ( GA == 0 )
TRAIN_COST = TRAIN_CO * ( GA == 0 )
# **Rescale some data**
TRAIN_TT_SCALED = DefineVariable('TRAIN_TT_SCALED', TRAIN_TT / 100.0, database)
TRAIN_COST_SCALED = DefineVariable('TRAIN_COST_SCALED', TRAIN_COST / 100, database)
SM_TT_SCALED = DefineVariable('SM_TT_SCALED', SM_TT / 100.0 , database)
SM_COST_SCALED = DefineVariable('SM_COST_SCALED', SM_COST / 100 , database)
CAR_TT_SCALED = DefineVariable('CAR_TT_SCALED', CAR_TT / 100 , database)
CAR_CO_SCALED = DefineVariable('CAR_CO_SCALED', CAR_CO / 100 , database)
# **Create parameters to be estimated**
#
# `Beta`
# 1. name of parameter
# 2. default value for the parameter
# 3. lower bound
# 4. upper bound
# 5. flag indicating if parameter is to be estimated
ASC_CAR = Beta('ASC_CAR',0,None,None,0)
ASC_TRAIN = Beta('ASC_TRAIN',0,None,None,1)
B_TIME = Beta('B_TIME',0,None,None,0)
B_COST = Beta('B_COST',0,None,None,0)
# **Define the utility functions**
# \begin{align}
# V_1 & = \beta_{time}X_{Train_{TT}} + \beta_{cost}X_{Train_{cost}}\\
# V_3 & = \beta_{Car} + \beta_{time}X_{Car_{TT}} + \beta_{cost}X_{Car_{cost}}\\
# \end{align}
V1 = B_TIME * TRAIN_TT_SCALED + \
B_COST * TRAIN_COST_SCALED
V3 = ASC_CAR + \
B_TIME * CAR_TT_SCALED + \
B_COST * CAR_CO_SCALED
# **Associate utility functions with alternatives and associate availability of alternatives**
#
# Create a python dictionary with all utility functions
#
# Create a python dictionary with availability of choices
P = {1: bioNormalCdf(V1-V3),
3: bioNormalCdf(V3-V1)}
# **Define the model**
logprob = log(Elem(P, CHOICE))
# **Define the Biogeme object**
#
# * Give the database with all variables
# * Give the log likelihood model
# +
biogeme = bio.BIOGEME(database, logprob)
biogeme.modelName = "swissmetro_probit_basic"
# -
# **Estimate the model**
#
# 1. A `.html` can be generated with a report of the results and can be opened with a browser
# 2. A `.pickle` file can also be generaetd with a snapshot with the results. This file can then be used in other scripts
# +
biogeme.generateHtml = True
biogeme.generatePickle = False
results = biogeme.estimate()
print(f"HTML file: {results.data.htmlFileName}")
print(f"Pickle file: {results.data.pickleFileName }")
# -
# **Print results**
betas = results.getBetaValues()
for k,v in betas.items():
print(f"{k:10}=\t{v:.3g}")
# **Get the general statistics**
# +
gs = results.getGeneralStatistics()
for k,v in gs.items():
print("{}= {}".format(k.ljust(45),v[0]))
| Code/8.1-MultinomialLogitAndProbitModels/05-probit-basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# name: python3
# ---
# This notebook is almost a copy of the single-block notebook except analyzing different experiment results.
# For more information, please refer to the single-block notebook.
#
# Microbenchmark corresponds to
# Figures 6(a,b) 7(a,b) 8(a,b) 9(a,b) 10(a,b) 16(a,b) 17(a,b) 18(a,b) in our paper.
# This notebook reproduces the graphs correspond to Figures 8(a,b) 9(a,b) 10(a,b) 18(a,b).
# download and unzip dataset
# !mkdir dataset_release
# !wget --directory-prefix=dataset_release "https://github.com/columbia/PrivateKube/releases/download/v1.0/multi_block.tar.gz"
# !tar -xzf ./dataset_release/multi_block.tar.gz --directory ./dataset_release
# + pycharm={"name": "#%%\n"}
import pandas as pd
from dpsched.analysis.data_collection import workspace2dataframe
from dpsched.analysis.plot import plot_granted_tasks, plot_delay_cdf
from dpsched.utils.configs import DpPolicyType
import matplotlib.pylab as plt
plt.rcParams['figure.facecolor'] = 'white'
# + pycharm={"name": "#%%\n"}
workspace_file = "workspace_06-25-04H-34-55" # remote multi
workspace_dir = "./dataset_release/%s" % workspace_file
table = workspace2dataframe(workspace_dir)
table.columns
# + [markdown] pycharm={"name": "#%% md\n"}
# Following graph reproduces
# - fig 8(a) DPF behavior on multiple blocks. - Number of pipelines allocated.
# - fig 9(a) DPF and DPF-T behavior on multiple blocks. - Number of pipelines allocated.
# - fig 10(a) Traditional vs. Rényi DP, multiple blocks. - Number of pipelines allocated.
# - fig 18(a) Rényi DPF and DPF-T behaviors on multiple blocks. - Number of pipelines allocated.
#
# Note that fig 9(a) 10(a) 18(a) aim to compare different policies.
# Therefore, they group curves in ways different than what we present here.
# + pycharm={"name": "#%% figure1\n"}
title = "Number of completed pipelines as a function of N/T"
table1 = table.loc[(table['epsilon_mice_fraction'] == 75)]
## extend FCFS data ponits into horizontal lines in different subplots.
fcfs_padding_list = []
fcfs_padding = table1[table1.policy == DpPolicyType.DP_POLICY_FCFS.value]
for N_or_T_based in ('N', 'T'):
for is_rdp in (True, False):
left = fcfs_padding[(fcfs_padding.is_rdp==is_rdp)].copy()
left.N_or_T_ = -1
left.N_or_T_based = N_or_T_based
fcfs_padding_list.append(left)
right = left.copy()
max_N_or_T = max(table1[(table1.N_or_T_based == N_or_T_based) & (table1.is_rdp ==is_rdp) ].N_or_T_)
right.N_or_T_ = max_N_or_T
right.N_or_T_based = N_or_T_based
# if not is_rdp:
# print(N_or_T_based)
# print(max_N_or_T)
fcfs_padding_list.append(right)
table1_1 = pd.concat([table1] + fcfs_padding_list, axis=0)
plot_granted_tasks(save_file_name="figure1.pdf", table=table1_1, title=title, xaxis_col='N_or_T_',
yaxis_col='granted_tasks_total')
# + [markdown] pycharm={"name": "#%% md\n"}
# Following graph reproduces
# - fig 8(b) DPF behavior on multiple blocks. - Number of pipelines allocated.
# - fig 9(b) DPF and DPF-T behavior on multiple blocks. - Number of pipelines allocated.
# - fig 10(b) Traditional vs. Rényi DP, multiple blocks. - Number of pipelines allocated.
# - fig 18(b) Rényi DPF and DPF-T behaviors on multiple blocks. - Number of pipelines allocated.
#
# Note that fig 9(b) 10(b) 18(b) aim to compare different policies.
# Therefore, they group curves in ways different than what we present here.
# + pycharm={"name": "#%% figure2\n"}
title = 'CDFs of scheduling delay for different values of N/T'
should_modify_alloc_duration = True # immediate rejection's delay is treated as timeout
should_plot_granted = False
should_exclude_late_task = False # exclude tasks arrived late
delay_lst_column = 'dp_allocation_duration_list'
plot_delay_cdf(table1, delay_lst_column, 'N_or_T_', 'figure2.pdf', should_exclude_late_task,
should_modify_timeout_duration=True, task_timeout=300, plot_title=title)
# + [markdown] pycharm={"name": "#%% md\n"}
# Our paper doesn't contain following figures.
# Basically, they are multiple-block version of fig 7(a) 17(a) in our paper.
# + pycharm={"name": "#%% figure3\n"}
title = 'Number of tasks completed as function of epsilon-mice percentage '
# fix the N/T parameter for multiple block workload
fixed_N_or_T_dp = 125
fixed_N_or_T_rdp = 25399
is_fixed_n = lambda x: x in (-1, fixed_N_or_T_dp, fixed_N_or_T_rdp)
table2 = table.loc[table.N_.apply(is_fixed_n)]
fcfs_padding_list = []
fcfs_padding = table2[table2.policy == DpPolicyType.DP_POLICY_FCFS.value]
# left = fcfs_padding.copy()
# left.N_or_T_based = 'N'
for N_or_T_based in ('N', 'T'):
pad_nt = fcfs_padding.copy()
pad_nt.N_or_T_based = N_or_T_based
fcfs_padding_list.append(pad_nt)
table2_1 = pd.concat([table2] + fcfs_padding_list, axis=0)
plot_granted_tasks(save_file_name="figure3.pdf", table=table2_1, title=title, xaxis_col='epsilon_mice_fraction',
yaxis_col='granted_tasks_total')
# + [markdown] pycharm={"name": "#%% md\n"}
# Our paper doesn't contain following figures.
# Basically, they are multiple-block version of fig 7(b) 17(b) in our paper.
# + pycharm={"name": "#%% figure4\n"}
title = 'CDFs of scheduling delay for different fraction of epsilon mice'
plot_delay_cdf(table2, delay_lst_column, 'epsilon_mice_fraction', 'figure4.pdf', should_exclude_late_task,
should_modify_timeout_duration=True, task_timeout=300, plot_title=title)
| evaluation/microbenchmark/microbenchmark_figures_multiple_block.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
with open('input.txt', 'r') as f:
luggage_rules = dict(
rule.strip('.').split(' bags contain ')
for rule in f.read().splitlines()
)
# +
def find_bags_containing(search_colors, found_colors=set()):
if search_colors:
next_search_colors = set([
k
for k, v in luggage_rules.items()
for c in search_colors
if c in v
]) - found_colors
return find_bags_containing(
next_search_colors,
found_colors.union(next_search_colors)
)
else:
return found_colors
bags_containing_shiny_gold = find_bags_containing(['shiny gold'])
print(f"Answer part one: {len(bags_containing_shiny_gold)}")
# -
luggage_rules = {
k: v.replace(' bags', '').replace(' bag', '').split(', ')
for k, v in luggage_rules.items()
}
# +
def count_bags_contained_in(key, factor=1):
total_nb_bags = 0
if luggage_rules[key] == ['no other']:
return 0
else:
for rule_part in luggage_rules[key]:
nb_bags = int(rule_part[0])
color = rule_part[2:]
total_nb_bags += nb_bags * (count_bags_contained_in(color, nb_bags) + 1)
return total_nb_bags
print(f"Answer part two: {count_bags_contained_in('shiny gold')}")
| day_07/Day 7 - Handy Haversacks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/RodrigoTenorio86/Machine_Learning__Classificacao_para_Motoristas/blob/master/Machine_Learning__Classificacao_para_Motoristas.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Yh1GDGy4wMQR" colab_type="text"
# # Machine Learning: para classifica qualidade de motoristas
# + id="3m42gu5HwU0O" colab_type="code" colab={}
import pandas as pd
from sklearn.naive_bayes import MultinomialNB
# + [markdown] id="25_n4UV2w05L" colab_type="text"
# ## importação da base de dados com historicos dos cliente.
# + [markdown] id="yKRa2uDBAwtg" colab_type="text"
# ## Importação da base de dados com 10 costumes dos motoristas como: Andar com combustível na reserva, Passar em lombadas ou valetas na diagonal, Encostar as rodas na guia, Girar o volante com o veículo parado ou com as rodas coladas na guia, Descansar o pé na embreagem, Passar em áreas alagadas, Descer a serra desengrenado, Dar arrancadas e reduzidas intensas, Usar óleo vencido e Andar com o carro desalinhado.(0=> Não comete e 1=>comete)
# + id="Y2gvSLPW3eEC" colab_type="code" colab={}
dados=pd.read_excel('/content/dados.xlsx',index_col=0,header=0 )
# + id="vH26_BbS33ld" colab_type="code" outputId="1bc94dae-88c1-4d5f-8045-27544b3b429d" colab={"base_uri": "https://localhost:8080/", "height": 272}
dados.head
# + id="ZF-FNr9iWeHg" colab_type="code" colab={}
dados_motoristas_treinando=[]
# + [markdown] id="gMLEyX4XxFdv" colab_type="text"
# ## conversão de DataFrame do Pandas para lista.
# + id="ULUx_XzRWjm-" colab_type="code" colab={}
dados_motoristas_treinando= dados.values.tolist()
# + id="hLbyIl41Wpk8" colab_type="code" outputId="a8a27212-a464-401b-cb8d-c6eb5a6b6425" colab={"base_uri": "https://localhost:8080/", "height": 34}
type(dados_motoristas_treinando)
# + id="qPCye_whWwyF" colab_type="code" outputId="a8abf5fb-5214-40b6-b06a-12c8fb1773a7" colab={"base_uri": "https://localhost:8080/", "height": 54}
print(dados_motoristas_treinando)
# + id="SqcBXRhzfXR3" colab_type="code" outputId="648c902e-d130-4626-bbac-0981eff09b2b" colab={"base_uri": "https://localhost:8080/", "height": 34}
len(dados_motoristas_treinando)
# + [markdown] id="CwqqDCQTxT_U" colab_type="text"
# ## Criação das marcas de classificação dos clientes da base de treinamento (0 => bons Motorista e 1 => maus Motoristas)
# + id="ohIYaPA9I9hK" colab_type="code" colab={}
marcacoes_treinando = []
# + id="QpxiK8vQG6Oy" colab_type="code" colab={}
for d in range(109):
if(d<55):
marcacoes_treinando.append(0)
elif(d >= 55):
marcacoes_treinando.append(1)
# + id="hzV40wEbI2-Z" colab_type="code" outputId="c38977fc-4721-4675-9a12-3fb01e086722" colab={"base_uri": "https://localhost:8080/", "height": 34}
len(marcacoes_treinando)
# + [markdown] id="YYehO6g1069h" colab_type="text"
# ## Criação de 10 novos clientes com perfis variados para base de teste do algoritmo
# + id="MeEdx7MHLgqA" colab_type="code" colab={}
motorista_1=[1,0,0,0,0,0,0,0,0,1]
motorista_2=[0,0,0,0,0,0,0,0,0,0]
motorista_3=[0,0,0,0,0,0,0,1,1,1]
motorista_4=[1,1,1,1,1,1,1,1,1,1]
motorista_5=[0,0,1,1,1,1,1,1,1,1]
motorista_6=[1,1,1,0,1,1,1,1,1,1]
motorista_7=[0,0,0,0,0,0,0,1,1,1]
motorista_8=[1,1,1,1,1,1,1,1,1,1]
motorista_9=[0,0,0,1,1,1,1,1,1,1]
motorista_10=[1,1,0,0,1,1,1,1,1,1]
# + id="wLmdWjh_SkBD" colab_type="code" colab={}
dados_motoristas_teste = [motorista_1, motorista_2, motorista_3,motorista_4, motorista_5,motorista_6,motorista_7,motorista_8,motorista_9,motorista_10]
# + id="9dTsAxGeycyz" colab_type="code" colab={}
resultado_do_teste = [0,0,0,1,1,1,0,1,1,1]
# + [markdown] id="B8ReYGI51KRn" colab_type="text"
# ## Desenvolvimento do algoritmo para classificação dos clientes entre bom ou mal motoristas
# + id="zbzhOM9P-J9G" colab_type="code" colab={}
modelo = MultinomialNB()
# + id="khAnh9YBB2-c" colab_type="code" outputId="ff5de2c8-4983-47ff-e9d8-ee378e896438" colab={"base_uri": "https://localhost:8080/", "height": 34}
modelo.fit(dados_motoristas_treinando,marcacoes_treinando)
# + [markdown] id="v04LxENi1ovw" colab_type="text"
# ## Predição dos clientes de testes(0 => bons motoristas e 1=> maus motoristas)
# + id="uDDhu9d-Mmt3" colab_type="code" outputId="93111c46-a3c6-415c-b4d4-9a4ce26148e7" colab={"base_uri": "https://localhost:8080/", "height": 34}
print(modelo.predict(dados_motoristas_teste))
# + [markdown] id="3BVu6i4sB0GO" colab_type="text"
# ## Calcular porcentagem de Acertos do Algoritmo
# + id="kAKb3QxSvHS1" colab_type="code" outputId="109bdf1d-2b2a-4d6e-8bfa-160917d96bb3" colab={"base_uri": "https://localhost:8080/", "height": 34}
resultado_predicao= modelo.predict(dados_motoristas_teste)
total_de_acertos = len( True == (resultado_predicao == resultado_do_teste) )
total_de_elemento_treinamento = len(dados_motoristas_treinando)
total_de_elemento_testa = len(dados_motoristas_teste)
porcentagem_acerto = 100.0 * total_de_acertos / total_de_elemento_testa
print('A taxa de Acerto do Algoritmo esta em {}.'.format(porcentagem_acerto))
# + id="_THtp0Hakn9-" colab_type="code" colab={}
#Array
par = [i for i in range(100) if i % 2 == 0 ]
| Machine_Learning__Classificacao_para_Motoristas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear Regression(线性回归)
# https://www.statsmodels.org/stable/regression.html
#
# 适用于自变量X和因变量Y为线性关系,具体来说,画出散点图可以用一条直线来近似拟合。一般线性模型要求观测值之间相互独立、残差(因变量)服从正态分布、残差(因变量)方差齐性
# 统计模型被假定为 Y=Xβ+μ, μ∼N(0,Σ)
# **Examples**
# +
#加载模块和数据
import numpy as np
import statsmodels.api as sm
spector_data = sm.datasets.spector.load()
spector_data.exog = sm.add_constant(spector_data.exog,prepend = False)
#训练并输出OLS模型
mod = sm.OLS(spector_data.endog,spector_data.exog)
res = mod.fit()
print(res.summary())
# -
# 
# 
# # Generalized Linear(广义线性回归)
# http://www.statsmodels.org/stable/glm.html#module-reference
#
# 是为了克服线性回归模型的缺点出现的,是线性回归模型的推广。首先自变量可以是离散的,也可以是连续的。离散的可以是0-1变量,也可以是多种取值的变量。广义线性模型又取消了对残差(因变量)服从正态分布的要求。残差不一定要服从正态分布,可以服从二项、泊松、负二项、正态、伽马、逆高斯等分布,这些分布被统称为指数分布族。
# 与线性回归模型相比较,有以下推广:
#
# 随机误差项不一定服从正态分布,可以服从二项、泊松、负二项、正态、伽马、逆高斯等分布,这些分布被统称为指数分布族。
# 引入link函数g(⋅)。因变量和自变量通过联接函数产生影响。根据不同的数据,可以自由选择不同的模型。大家比较熟悉的Logit模型就是使用Logit联接、随机误差项服从二项分布得到模型。
# The statistical model for each observation ii is assumed to be
# 
# **Examples**
# +
import statsmodels.api as sm
data = sm.datasets.scotland.load()
data.exog = sm.add_constant(data.exog)
gamma_model = sm.GLM(data.endog,data.exog,family = sm.families.Gamma())
gamma_results = gamma_model.fit()
print(gamma_results.summary())
# -
# 
# 
# 
# # Generalized Estimating Equations(广义估计方程)
# http://www.statsmodels.org/stable/gee.html
#
# 实际工作中一些资料由于部分观察值含有非独立或相关的信息,不能用传统的一般线性(或广义线性进行分析),故而发展出了广义估计方程。如纵向数据,重复测量数据,整群抽样设计资料,聚集性资料或是多次层次系统结构资料。
# **Examples**
# +
import statsmodels.api as sm
import statsmodels.formula.api as smf
data = sm.datasets.get_rdataset('epil', package='MASS').data
fam = sm.families.Poisson()
ind = sm.cov_struct.Exchangeable()
mod = smf.gee("y ~ age + trt + base", "subject", data,
cov_struct=ind, family=fam)
res = mod.fit()
print(res.summary())
# -
# 
# 
# # Robust Linear Models(稳健的线性模型)
# http://www.statsmodels.org/stable/rlm.html
#
# 稳健回归(robust regression)是将稳健估计方法用于回归模型,以拟合大部分数据存在的结构,同时可识别出潜在可能的离群点、强影响点或与模型假设相偏离的结构。当误差服从正态分布时,其估计几乎和最小二乘估计一样好,而最小二乘估计条件不满足时,其结果优于最小二乘估计。
# +
#加载模块和数据
import statsmodels.api as sm
data = sm.datasets.stackloss.load()
data.exog = sm.add_constant(data.exog)
#训练模型并输出结果
rlm_model = sm.RLM(data.endog, data.exog, M=sm.robust.norms.HuberT())
rlm_results = rlm_model.fit()
print(rlm_results.params)
# -
# 
# # Linear Mixed Effects Models(线性混合效应模型)
# http://www.statsmodels.org/stable/mixed_linear.html
#
# 在线性模型中加入随机效应项,消了观测值之间相互独立和残差(因变量)方差齐性的要求。
import statsmodels.api as sm
import statsmodels.formula.api as smf
data = sm.datasets.get_rdataset("dietox", "geepack").data
md = smf.mixedlm("Weight ~ Time", data, groups=data["Pig"])
mdf = md.fit()
print(mdf.summary())
# 
# # Regression with Discrete Dependent Variable(具有离散因变量的回归)
# http://www.statsmodels.org/stable/discretemod.html
spector_data = sm.datasets.spector.load()
spector_data.exog = sm.add_constant(spector_data.exog)
logit_mod = sm.Logit(spector_data.endog, spector_data.exog)
logit_res = logit_mod.fit()
print(logit_res.summary())
# 
# DiscreteModel是所有离散回归模型的超类。估算结果作为其中一个子类的实例返回 DiscreteResults。模型的每个类别(二进制,计数和多项)都有其自己的中级模型和结果类。这个中间类主要是为了方便实现由DiscreteModel和 定义的方法和属性DiscreteResults。
# 
# # Generalized Linear Mixed Effects Models(广义线性混合效应模型)
# http://www.statsmodels.org/stable/mixed_glm.html
#
# 广义线性混合效应模型是混合效应模型的推广
# 
| 初稿/Regression(回归).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "skip"}
# # Analysis <a class="tocSkip">
#
# For homework: [profs.info.uaic.ro/~adria/teach/courses/pcd//homework/PCD_Homework1.pdf](https://profs.info.uaic.ro/~adria/teach/courses/pcd//homework/PCD_Homework1.pdf)
# Used dataset: [kaggle.com/wcukierski/enron-email-dataset/home](https://www.kaggle.com/wcukierski/enron-email-dataset/home)
#
#
# [wiki.python.org/moin/UdpCommunication](https://wiki.python.org/moin/UdpCommunication)
# [stackoverflow.com/questions/22819214/udp-message-too-long
# ](https://stackoverflow.com/questions/22819214/udp-message-too-long)
#
#
# + [markdown] slideshow={"slide_type": "skip"}
# ## Background <a class="tocSkip">
#
# Solution architecture: <drawio fig>
#
# Perform the following scenarios like a battery of tests
# # + UDP - single-node
# # + UDP - single-node, multithread
# # + UDP - single-node, multicast
#
# # + TCP - single-node
# # + TCP - single-node, multithread
#
# # + use [RabbitMQ](https://www.rabbitmq.com/) or [ZeroMQ](http://zeromq.org/)
# + https://github.com/booksbyus/zguide/tree/master/examples/Python
#
# https://stackoverflow.com/questions/39246259/tcp-threaded-python
# https://stackoverflow.com/questions/28840624/forking-server-in-python
# https://stackoverflow.com/questions/32654551/tcp-vs-udp-socket-latency-benchmark
#
#
# Extrapolate to utils, scenario files
# #client sends END, server switches to next case
#
#
# udp-streaming - https://www.google.com/search?safe=off&ei=_051XO-aO42EmwW4i5mAAw&q=udp+streaming+python&oq=udp+streaming+python&gs_l=psy-ab.3..0i19j0i22i30i19l2.8985.10124..10325...0.0..0.164.1007.0j7......0....1..gws-wiz.......0i71j0i8i13i30i19.ITQZ5dy4ifs
#
# https://stackoverflow.com/questions/15909064/python-implementation-for-stop-and-wait-algorithm
#
#
#
# airflow
# https://stackoverflow.com/questions/43386003/airflow-inside-docker-running-a-docker-container
# http://localhost:8080/admin/
# http://michal.karzynski.pl/blog/2017/03/19/developing-workflows-with-apache-airflow/
# https://airflow.apache.org/tutorial.html
# http://www.golangbootcamp.com/book/concurrency
#
#
# + [markdown] slideshow={"slide_type": "skip"}
# # Imports
# + [markdown] slideshow={"slide_type": "skip"}
# ## Import deps
# + slideshow={"slide_type": "skip"}
import os
import pandas as pd
import utils
import plotly.graph_objs as go
import plotly.figure_factory as ff
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
init_notebook_mode(connected=True)
# + [markdown] slideshow={"slide_type": "skip"}
# ## Import data
# + slideshow={"slide_type": "skip"} language="bash"
# ls -l data/ | grep csv
# + slideshow={"slide_type": "skip"}
root, dirs, files = os.walk('data/').__next__()
metadata_files = [file for file in files if file.startswith('metadata')]
print(metadata_files)
# + slideshow={"slide_type": "skip"}
lst_df = []
for file in metadata_files:
lst_df.append(pd.read_csv(root + file))
df = pd.concat(lst_df)
df.reset_index(inplace=True)
df.Time = pd.to_timedelta(df.Time)
df['SizeBytes'] = df['Size(MB)'].apply(utils.parseSize)
df = df[['Protocol', 'Type', 'Location',
'Setup', 'Mode', 'Rounds', 'Time',
'Count', 'SizeBytes', 'Size(MB)']]
df = df.sort_values('Rounds').sort_values('Protocol')
df.reset_index(inplace=True, drop=True)
df.info()
df.head(df.shape[0])
# + [markdown] slideshow={"slide_type": "skip"}
# # Deltas
# -
# ## Delta UDP
delta_udp_time = []
delta_udp_count = []
delta_udp_bytes = []
delta_udp_mb = []
# + slideshow={"slide_type": "skip"}
delta_udp = df[df.Protocol == 'UDP']
delta_udp
# -
(delta_udp.iloc[1].Time - delta_udp.iloc[0].Time)# .seconds
# + slideshow={"slide_type": "skip"}
# delta_udp_time = delta_udp[delta_udp.Type == 'server'].Time - delta_udp[delta_udp.Type == 'client'].Time
delta_udp_time.append((delta_udp.iloc[1].Time - delta_udp.iloc[0].Time))
delta_udp_time.append((delta_udp.iloc[3].Time - delta_udp.iloc[2].Time))
delta_udp_time.append((delta_udp.iloc[4].Time - delta_udp.iloc[5].Time)) # diff
delta_udp_time
# +
y = delta_udp_time
trace = [go.Box(
y=y,
jitter=0.3,
pointpos=-1.8,
boxpoints = 'suspectedoutliers',
marker = dict(
color = 'rgb(8,81,156)',
outliercolor = 'rgba(219, 64, 82, 0.6)',
line = dict(
outliercolor = 'rgba(219, 64, 82, 0.6)',
outlierwidth = 2)
),
# name = name + ' - suspected outliers'
)]
iplot(trace)
# + slideshow={"slide_type": "skip"}
# delta_udp_size = delta_udp[delta_udp.Type == 'client'].SizeBytes - delta_udp[delta_udp.Type == 'server'].SizeBytes
delta_udp_bytes.append((delta_udp.iloc[0].SizeBytes - delta_udp.iloc[1].SizeBytes))
delta_udp_bytes.append((delta_udp.iloc[2].SizeBytes - delta_udp.iloc[3].SizeBytes))
delta_udp_bytes.append((delta_udp.iloc[4].SizeBytes - delta_udp.iloc[5].SizeBytes))
print(delta_udp_bytes)
delta_udp_mb.append(utils.convert_size(delta_udp_bytes[0]))
delta_udp_mb.append(utils.convert_size(delta_udp_bytes[1]))
delta_udp_mb.append(utils.convert_size(delta_udp_bytes[2]))
print(delta_udp_mb)
# +
y = delta_udp_bytes
trace = [go.Box(
y=y,
jitter=0.3,
pointpos=-1.8,
boxpoints = 'suspectedoutliers',
marker = dict(
color = 'rgb(8,81,156)',
outliercolor = 'rgba(219, 64, 82, 0.6)',
line = dict(
outliercolor = 'rgba(219, 64, 82, 0.6)',
outlierwidth = 2)
),
# name = name + ' - suspected outliers'
)]
iplot(trace)
# + slideshow={"slide_type": "skip"}
delta_udp_count.append((delta_udp.iloc[0].Count - delta_udp.iloc[1].Count))
delta_udp_count.append((delta_udp.iloc[2].Count - delta_udp.iloc[3].Count))
delta_udp_count.append((delta_udp.iloc[4].Count - delta_udp.iloc[5].Count))
print(delta_udp_count)
# +
y = delta_udp_count
trace = [go.Box(
y=y,
jitter=0.3,
pointpos=-1.8,
boxpoints = 'suspectedoutliers',
marker = dict(
color = 'rgb(8,81,156)',
outliercolor = 'rgba(219, 64, 82, 0.6)',
line = dict(
outliercolor = 'rgba(219, 64, 82, 0.6)',
outlierwidth = 2)
),
# name = name + ' - suspected outliers'
)]
iplot(trace)
# + [markdown] slideshow={"slide_type": "skip"}
# ## Delta ZeroMQ
# -
delta_zmq_time = []
delta_zmq_count = []
delta_zmq_bytes = []
delta_zmq_mb = []
# + slideshow={"slide_type": "skip"}
delta_zmq = df[df.Protocol == 'ZeroMQ']
delta_zmq
# + slideshow={"slide_type": "skip"}
# delta_zmq_time = delta_zmq[delta_zmq.Type == 'server'].Time - delta_zmq[delta_zmq.Type == 'client'].Time
delta_zmq_time.append((delta_zmq.iloc[1].Time - delta_zmq.iloc[0].Time))
delta_zmq_time.append((delta_zmq.iloc[2].Time - delta_zmq.iloc[3].Time)) # diff
delta_zmq_time.append((delta_zmq.iloc[4].Time - delta_zmq.iloc[5].Time))
delta_zmq_time
# + slideshow={"slide_type": "skip"}
# delta_zmq_size = delta_udp[delta_udp.Type == 'client'].SizeBytes - delta_udp[delta_udp.Type == 'server'].SizeBytes
delta_zmq_bytes.append((delta_zmq.iloc[0].SizeBytes - delta_zmq.iloc[1].SizeBytes))
delta_zmq_bytes.append((delta_zmq.iloc[2].SizeBytes - delta_zmq.iloc[3].SizeBytes))
delta_zmq_bytes.append((delta_zmq.iloc[4].SizeBytes - delta_zmq.iloc[5].SizeBytes))
print(delta_zmq_bytes)
delta_zmq_mb.append(utils.convert_size(delta_zmq_bytes[0]))
delta_zmq_mb.append(utils.convert_size(delta_zmq_bytes[1]))
delta_zmq_mb.append(utils.convert_size(delta_zmq_bytes[2]))
print(delta_udp_mb)
# + slideshow={"slide_type": "skip"}
delta_zmq_count.append((delta_zmq.iloc[0].Count - delta_zmq.iloc[1].Count))
delta_zmq_count.append((delta_zmq.iloc[2].Count - delta_zmq.iloc[3].Count))
delta_zmq_count.append((delta_zmq.iloc[4].Count - delta_zmq.iloc[5].Count))
print(delta_udp_count)
# + slideshow={"slide_type": "skip"}
# + slideshow={"slide_type": "skip"}
# + slideshow={"slide_type": "skip"}
# + [markdown] slideshow={"slide_type": "skip"}
# ---
| data-engineering/labs-concurrent-distributed-programming/Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import os
import json
import pathlib
import re
from collections import defaultdict
DATA = str(pathlib.Path('./').resolve()) + '/data'
punct = re.compile("[:.,l!?-]+")
paths = None
papers = []
for root, dirs, files in os.walk('./data'):
paths = [ os.path.join(DATA, flname) for flname in files ]
for path in paths:
with open(path, 'r', encoding='utf-8') as fh:
papers.extend(json.load(fh))
abstracts = [paper["abstract"] for paper in papers]
lengs = list(set([len(abs1) for abs1 in abstracts]))
sorted(lengs)
print(lengs[0], lengs[-1])
abstracts_tok = [ re.sub(punct, "", abs1.replace("\n", " ")).lower().split(" ") for abs1 in abstracts]
cnt = defaultdict(int)
for abs1 in abstracts_tok:
for tok in abs1:
cnt[tok] += 1
# -
res = dict()
for key, val in cnt.items():
if val < 100 and val > 50 and key.isalpha() and len(key) > 3:
res[len(key)] = val
res
with open("forwiz.json", "w") as fh:
fh.write(json.dumps(list(res.items())))
| lesson2/Tokenization-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from keras.preprocessing import sequence
from keras.models import Sequential
from keras.layers import Dense, Dropout, Activation
from keras.layers import Embedding
from keras.layers import LSTM
from keras.callbacks import EarlyStopping
from keras.datasets import imdb
# -
n_words = 1000
(X_train, y_train), (X_test, y_test) = imdb.load_data(num_words=n_words)
print('Train seq: {}'.format(len(X_train)))
print('Test seq: {}'.format(len(X_train)))
# +
print('Train example: \n{}'.format(X_train[0]))
print('\nTest example: \n{}'.format(X_test[0]))
# Note: the data is already preprocessed (words are mapped to vectors)
# -
# Pad sequences with max_len
max_len = 200
X_train = sequence.pad_sequences(X_train, maxlen=max_len)
X_test = sequence.pad_sequences(X_test, maxlen=max_len)
# +
# Define network architecture and compile
model = Sequential()
model.add(Embedding(n_words, 50, input_length=max_len))
model.add(Dropout(0.2))
model.add(LSTM(100, dropout=0.2, recurrent_dropout=0.2))
model.add(Dense(250, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(1, activation='sigmoid'))
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
model.summary()
# -
callbacks = [EarlyStopping(monitor='val_acc', patience=3)]
# +
batch_size = 128
n_epochs = 100
model.fit(X_train, y_train, batch_size=batch_size, epochs=n_epochs, validation_split=0.2, callbacks=callbacks)
# +
print('Accuracy on test set: {}'.format(model.evaluate(X_test, y_test)[1]))
# Accuracy on test set: 0.82884
| Chapter04/Chapter 4 - Adding Long Short-Term Memory (LSTM).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# 说明:
# 给定 N,想象一个凸 N 边多边形,其顶点按顺时针顺序依次标记为 A[0], A[i], ..., A[N-1]。
# 假设您将多边形剖分为 N-2 个三角形。
# 对于每个三角形,该三角形的值是顶点标记的乘积,三角剖分的分数是进行三角剖分后所有 N-2 个三角形的值之和。
# 返回多边形进行三角剖分后可以得到的最低分。
#
#
# 示例 2:
# 输入:[3,7,4,5]
# 输出:144
# 解释:有两种三角剖分,可能得分分别为:3*7*5 + 4*5*7 = 245,或 3*4*5 + 3*4*7 = 144。最低分数为 144。
#
# 示例 3:
# 输入:[1,3,1,4,1,5]
# 输出:13
# 解释:最低分数三角剖分的得分情况为 1*1*3 + 1*1*4 + 1*1*5 + 1*1*1 = 13。
#
# 提示:
# 1、3 <= A.length <= 50
# 2、1 <= A[i] <= 100
# -
# <img src='https://assets.leetcode.com/uploads/2019/05/01/minimum-score-triangulation-of-polygon-1.png'>
# + active=""
# 示例 1:
# 输入:[1,2,3]
# 输出:6
# 解释:多边形已经三角化,唯一三角形的分数为 6。
# -
class Solution:
def minScoreTriangulation(self, A):
N = len(A)
dp = [[float('inf')] * N for _ in range(N)]
for i in range(N-1):
dp[i][i+1] = 0
for dis in range(3, N + 1):
for i in range(N - dis + 1):
j = i + dis - 1
for k in range(i + 1, j):
dp[i][j] = min(dp[i][j], dp[i][k] + A[i] * A[j] * A[k] + dp[k][j])
return dp[0][-1]
solution = Solution()
solution.minScoreTriangulation([3,7,4,5])
| Dynamic Programming/1029/1039. Minimum Score Triangulation of Polygon.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Ricnelle/CPEN-21A-ECE-2-2/blob/main/Operations_and_Expressions.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="p_UuDPMo8IbA"
# #Operations and Expressions
# + [markdown] id="3d9s2wLFq50-"
# ##Boolean Operators
# + colab={"base_uri": "https://localhost:8080/"} id="bcHNPmR5rDF2" outputId="6609d1aa-3024-4198-c187-81493c1d14db"
print(10>9)
print(10==9)
print (10<9)
a=10
b=9
print(a>b)
# + colab={"base_uri": "https://localhost:8080/"} id="hurYnPt4t7Ty" outputId="9762f28e-eac3-4695-c7ed-71d4c00e28c2"
print(bool("Hello"))
print(bool(15))
print(bool(False))
print(bool(1))
print(bool(0))
print(bool(None))
print(bool({}))
print(bool([]))
# + colab={"base_uri": "https://localhost:8080/"} id="E5M_hFuTu1k1" outputId="f7d35e51-a2ea-44d5-cbe1-1df8fc69791e"
def myFunction():return True
print(myFunction())
# + colab={"base_uri": "https://localhost:8080/"} id="ZlaF80LovX8W" outputId="1b4b9125-2964-4f40-883c-bea825c365f3"
def myFunction():return True
if myFunction():
print("Yes")
else:
print("No")
# + colab={"base_uri": "https://localhost:8080/"} id="Q1ImIsgVv9aS" outputId="e2ec0fef-4598-4f5a-cf11-a2fbe8023b81"
print(10>9)
a=6
b=7
print(a==b)
print(6==6)
print(a!=a)
# + [markdown] id="i1MbZPOzw5xs"
# ##Python Operators
# + colab={"base_uri": "https://localhost:8080/"} id="R6T_tzzxw89C" outputId="305ed7f1-540c-4b35-833f-bd6d90e3423d"
print(10+5)
print(10-5)
print(10*5)
print(10/5)
print(10%5)
print(10//3)
print(10**2)
# + [markdown] id="HOF0S7vmydMS"
# ##Bitwise Operators
# + colab={"base_uri": "https://localhost:8080/"} id="k05uySxDyMYh" outputId="3b33e409-249c-4a26-8f5d-e354c50561aa"
a=60 #0011 1100
b=13 #0000 1101
print(a&b)
print(a^b)
print(~a)
print(a<<2)#1111 0000
print(a>>2)#0000 1111
# + [markdown] id="Spl6uz7K0zxP"
# ##Assignment Operator
# + colab={"base_uri": "https://localhost:8080/"} id="T9rdc2ha04nk" outputId="8b8d189c-5baf-40e1-eb09-f837073f5b3b"
x=2
x+=3
print(x) #same as x=x+3
# + [markdown] id="h8UK859h2iOT"
# ##Logical Operators
# + colab={"base_uri": "https://localhost:8080/"} id="0ZKjD6_62law" outputId="7f9b64f1-ad18-4454-8ca3-51296ff7d9c3"
a=5
b=6
print(a>b and a==a)
a<b or b==a
# + [markdown] id="HJ_Nzmot4ERh"
# ##Identity Operator
# + colab={"base_uri": "https://localhost:8080/"} id="Xrt2BpHs4HpD" outputId="e3baabd5-347c-4165-ec98-afe1db888fc2"
print(a is b)
a is not b
| Operations_and_Expressions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch as tc
from scipy.sparse.linalg import eigsh
from BasicFun import eigs_AD
# +
# 自动微分求解实对称矩阵的本征值与本征向量
# 构建随机实对称矩阵
dim = 6
M = tc.randn(dim, dim)
M = M + M.t()
print('利用scipy中的本征值分解求解最大本征值与本征向量')
lm0, v0 = eigsh(M.numpy(), k=1, which='LA')
print('矩阵的最大本征值为:')
print(lm0[0])
print('矩阵的最大本征向量为:')
print(v0.reshape(-1, ))
print('\n利用自动微分求解最大本征值与本征向量')
lm1, v1 = eigs_AD(M)
print('矩阵的最大本征值为:')
print(lm1.item())
print('矩阵的最大本征向量为:')
print(v1.data.to('cpu').numpy().reshape(-1, ))
| sec4_8_eigAD.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## <span style="color:Blue"> Categorical Feature Selection using Chi-Square Test of Independence - Part 2
#
# Youtube Video Explanation : https://youtu.be/2PbfYO7fu3I
# ### <span style="color:green"> Categorical Feature Selection using <span style="color:red"> sklearn <span style="color:green"> Library <span style="color:red"> chi2 <span style="color:green"> and <span style="color:red"> SelectKbest <span style="color:green"> function
# #### Youtube Video Explanation :
# - The Chi-Square test of independence is used to determine if there is a significant relationship between two categorical (nominal) variables.
# - It means the Chi-Square Test of Independence is a hypothesis testing test with 2 hypotheses present; the Null Hypothesis and the Alternative Hypothesis.
# - The hypothesis is written below.
# - Null Hypothesis (H0): There is no relationship between the variables
# - Alternative Hypothesis (H1): There is a relationship between variables
#
# - Just like any statistical testing,
# - Lets we choose our p-value = 0.05 : Choose a significance level (e.g. SL = 0.05 with a 95% confidence).
# - if p-value test result is more than 0.05 , it means that test result will lie in acceptance region and we will accept the null hypothesis
# - if p-value test result is less than 0.05 , it means that test result will lie in rejection(critical) region and we will reject the null hypothesis and will accept the alternate hypothesis.
#
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import math
from sklearn.feature_selection import chi2,SelectKBest
#Load the dataset #https://www.kaggle.com/burak3ergun/loan-data-set
df_loan = pd.read_csv("https://raw.githubusercontent.com/atulpatelDS/Data_Files/master/Loan_Dataset/loan_data_set.csv")
df_loan.head()
df_loan.info()
# Remove all null value
df_loan.dropna(inplace=True)
# drop the uninformative column("Loan_ID")
df_loan.drop(labels=["Loan_ID"],axis=1,inplace=True)
df_loan.reset_index(drop=True,inplace=True)
df_loan["Credit_History"]=df_loan["Credit_History"].apply(lambda x: "N" if x == 0 else "Y")
df_loan1 = df_loan.copy()
cat_cols = df_loan.select_dtypes(include= "object").columns
cat_cols
cat_col = df_loan.select_dtypes(include= "object").drop('Loan_Status', axis = 1).columns
cat_col
# Convert object to category
df_loan[cat_cols] = df_loan[cat_cols].apply(lambda x:x.astype("category"))
# encoding
df_loan[cat_cols]=df_loan[cat_cols].apply(lambda x: x.cat.codes)
df_loan.info()
df_loan.head()
X = df_loan[cat_col]
y = df_loan["Loan_Status"]
# Lets use the sklearn chi2 fnction
cs = SelectKBest(score_func=chi2,k=6)
cs.fit(X,y)
feature_score = pd.DataFrame({"Score":cs.scores_,"P_Value":cs.pvalues_},index=X.columns)
feature_score.nlargest(n=6,columns="Score")
# ### <span style="color:red"> Lets use the the getdummy encoding and use the chi2
df_loan1.head()
cat_col = df_loan1.select_dtypes(include= "object").columns
cat_col = cat_col.drop("Loan_Status")
cat_col
df_dummy = pd.get_dummies(df_loan1[cat_col])
df_dummy.head()
df_loan2 = pd.concat([df_dummy,df_loan1],axis=1)
df_loan2 = df_loan2.drop(columns=cat_col,axis=1)
df_loan2.head()
df_loan2["Loan_Status"] = df_loan2["Loan_Status"].replace({"N":0,"Y":1})
# +
#df_loan2['Loan_Status'] = df_loan2['Loan_Status'].apply(lambda x: 0 if x == 'N' else 1)
# -
cat_col
df_loan2.info()
X_loan = df_loan2.iloc[:,0:-1]
y_loan = df_loan2["Loan_Status"]
X_loan.shape,y_loan.shape
# Apply sklearn ch2
chc=SelectKBest(score_func=chi2,k=21)
chc.fit(X_loan,y_loan)
features_score_pvalue=pd.DataFrame({"Score":chc.scores_,"P_Value":np.round(chc.pvalues_,3)},index=X_loan.columns)
features_score_pvalue.nlargest(n=21,columns="Score")
# ### <span style="color:red"> Lets apply the Logistic Regression Model without feature selection
df_loan1.head()
df_loan1.info()
# As we are using the dummy encoding so to avoid multicollinearity for Logistic Regression we will delete the drop_first while doing encoding
df_dummy1 = pd.get_dummies(df_loan1,columns=df_loan1.select_dtypes(
exclude=["float64","int64"]).drop("Loan_Status",axis=1).columns,drop_first=True)
# +
#Change the Loan_Status into numerical value
# -
df_dummy1["Loan_Status"] = df_dummy1["Loan_Status"].apply(lambda x:0 if x=="N" else 1)
df_dummy1.head()
df_dummy1.Loan_Status.value_counts()
X_loan = df_dummy1.drop("Loan_Status",axis=1)
y_loan = df_dummy1["Loan_Status"]
print(X_loan.shape)
print(y_loan.shape)
print(type(X_loan))
print(type(y_loan))
# Split train and test
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test = train_test_split(X_loan,y_loan,test_size = 0.30,random_state = 42)
# Creating the prediction model
from sklearn.linear_model import LogisticRegression
log_model = LogisticRegression()
log_model.fit(X_train,y_train)
# Evaluate the metrics
from sklearn.metrics import classification_report,confusion_matrix,roc_curve,auc,accuracy_score
pred = log_model.predict(X_test)
print("Accuracy_bae_model :",accuracy_score(y_test,pred))
print("Classification Report :")
print(classification_report(y_test,pred))
# Plot the ROC-AUC plot
import warnings
warnings.filterwarnings("ignore", category=UserWarning)
import matplotlib.pyplot as plt
pred_prob = log_model.predict_log_proba(X_test)[:,1]
fpr,tpr,threshold = roc_curve(y_test,pred_prob)
roc_auc = auc(fpr,tpr)
plt.figure(figsize=(5,3))
plt.title("ROC")
plt.plot(fpr,tpr,"b",Label="AUC:{}".format(round(roc_auc,2)))
plt.legend(loc=2)
plt.plot([0,1],[0,1],"r--")
plt.xlim([0,1])
plt.ylim([0,1])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
plt.show()
# ### <span style="color:red"> Lets apply the Logistic Regression Model with feature selection using Chi-Square Test of Independence.
df_loan1.head()
# Takes only Categorical variables and do encoding
df_dummy2 = pd.get_dummies(df_loan1[cat_col],drop_first=True)
df_dummy2.head()
df_dummy2 = pd.concat([df_dummy2,df_loan1["Loan_Status"]],axis=1)
df_dummy2.head()
df_dummy2["Loan_Status"] = df_dummy2["Loan_Status"].apply(lambda x:0 if x=="N" else 1)
df_dummy2.info()
X_dummy = df_dummy2.iloc[:,0:-1]
y_dummy = df_dummy2["Loan_Status"]
# Apply sklearn ch2
ch2c=SelectKBest(score_func=chi2,k=10)
ch2c.fit(X_dummy,y_dummy)
features_score_pvalue=pd.DataFrame({"Score":ch2c.scores_,"P_Value":np.round(ch2c.pvalues_,3)},index=X_dummy.columns)
features_score_pvalue.nlargest(n=10,columns="Score")
features_score_pvalue = features_score_pvalue[features_score_pvalue["P_Value"]<0.50]
selected_cat_cols = features_score_pvalue.index.values.tolist()
selected_cat_cols
num_cols = df_loan1.select_dtypes(include=["float64","int64"]).columns.to_list()
num_cols
selected_cols = selected_cat_cols + num_cols
#selected_cols = num_cols
#selected_cols = selected_cat_cols
selected_cols
df_dummy1.head()
# Lets
X_select = df_dummy1[selected_cols]
y_select = df_dummy1["Loan_Status"]
# Split the dataset in train and test
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import auc,classification_report,accuracy_score
Xs_train,Xs_test,ys_train,ys_test = train_test_split(X_select,y_select,test_size = 0.3,random_state = 42)
## Apply Logistic Regression
log_reg = LogisticRegression()
log_reg.fit(Xs_train,ys_train)
pred_reg = log_reg.predict(Xs_test)
print("Accuracy_bae_model :",accuracy_score(ys_test,pred_reg))
print("Classification Report :")
print(classification_report(ys_test,pred_reg))
| Feature_Engineering/Categorical Feature Selection using sklearn Library chi2 and SelectKbest function Tutorial 13.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# # Ch `02`: Concept `07`
# + [markdown] deletable=true editable=true
# ## Loading variables
# + [markdown] deletable=true editable=true
# Concept 06 was about saving variables. This one's about loading what you saved. Start by creating an interactive session:
# + deletable=true editable=true
import tensorflow as tf
sess = tf.InteractiveSession()
# + [markdown] deletable=true editable=true
# Create a boolean vector called `spikes` of the same dimensions as before:
# + deletable=true editable=true
spikes = tf.Variable([False]*8, name='spikes')
# + [markdown] deletable=true editable=true
# Restored the variable data from disk, serve warm, and enjoy:
# + deletable=true editable=true
saver = tf.train.Saver()
try:
saver.restore(sess, 'spikes.ckpt')
print(spikes.eval())
except:
print('file not found')
# + [markdown] deletable=true editable=true
# Show's over, goodnight:
# + deletable=true editable=true
sess.close()
| Concept07_loading_variables.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Full name: <NAME>
# ## R#: 321654987
# ## HEX: 0x132c10cb
# ## Title of the notebook
# ## Date: 9/3/2020
# # Laboratory 5 Functions
# Functions are simply pre-written code fragments that perform a certain task.
# In older procedural languages functions and subroutines are similar, but a function returns a value whereas
# a subroutine operates on data.
# The difference is subtle but important.
#
# More recent thinking has functions being able to operate on data (they always could) and the value returned may be simply an exit code.
# An analogy are the functions in *MS Excel*.
# To add numbers, we can use the sum(range) function and type `=sum(A1:A5)` instead of typing `=A1+A2+A3+A4+A5`
#
# ## Calling the Function
# We call a function simply by typing the name of the function or by using the dot notation.
# Whether we can use the dot notation or not depends on how the function is written, whether it is part of a class, and how it is imported into a program.
#
# Some functions expect us to pass data to them to perform their tasks.
# These data are known as parameters( older terminology is arguments, or argument list) and we pass them to the function by enclosing their values in parenthesis ( ) separated by commas.
#
# For instance, the `print()` function for displaying text on the screen is \called" by typing `print('Hello World')` where print is the name of the function and the literal (a string) 'Hello World' is the argument.
#
# ## Program flow
# A function, whether built-in, or added must be defined *before* it is called, otherwise the script will fail. Certain built-in functions "self define" upon start (such as `print()` and `type()` and we need not worry about those funtions). The diagram below illustrates the requesite flow control for functions that need to be defined before use.
#
# 
#
# An example below will illustrate, change the cell to code and run it, you should get an error.
# Then fix the indicated line (remove the leading "#" in the import math ... line) and rerun, should get a functioning script.
# + jupyter={"outputs_hidden": false}
# reset the notebook using a magic function in JupyterLab
# %reset -f
# An example, run once as is then activate indicated line, run again - what happens?
x= 4.
sqrt_by_arithmetic = x**0.5
print('Using arithmetic square root of ', x, ' is ',sqrt_by_arithmetic )
import math # import the math package ## activate and rerun
sqrt_by_math = math.sqrt(x) # note the dot notation
print('Using math package square root of ', x,' is ',sqrt_by_arithmetic)
# -
# An alternate way to load just the sqrt() function is shown below, either way is fine.
# + jupyter={"outputs_hidden": false}
# reset the notebook using a magic function in JupyterLab
# %reset -f
# An example, run once as is then activate indicated line, run again - what happens?
x= 4.
sqrt_by_arithmetic = x**0.5
print('Using arithmetic square root of ', x, ' is ',sqrt_by_arithmetic )
from math import sqrt # import sqrt from the math package ## activate and rerun
sqrt_by_math = sqrt(x) # note the notation
print('Using math package square root of ', x,' is ',sqrt_by_arithmetic)
# -
# ## Built-In in Primitive Python (Base install)
#
# The base Python functions and types built into it that are always available, the figure below lists those functions.
#
# 
#
# Notice all have the structure of `function_name()`, except `__import__()` which has a constructor type structure, and is not intended for routine use. We will learn about constructors later.
#
# ## Added-In using External Packages/Modules and Libaries (e.g. math)
#
# Python is also distributed with a large number of external functions.
# These functions are saved
# in files known as modules.
# To use the built-in codes in Python modules, we have to import
# them into our programs first. We do that by using the import keyword.
# There are three
# ways to import:
# 1. Import the entire module by writing import moduleName; For instance, to import the random module, we write import random. To use the randrange() function in the random module, we write random.randrange( 1, 10);28
# 2. Import and rename the module by writing import random as r (where r is any name of your choice). Now to use the randrange() function, you simply write r.randrange(1, 10); and
# 3. Import specific functions from the module by writing from moduleName import name1[,name2[, ... nameN]]. For instance, to import the randrange() function from the random module, we write from random import randrange. To import multiple functions, we separate them with a comma. To import the randrange() and randint() functions, we write from random import randrange, randint. To use the function now, we do not have to use the dot notation anymore. Just write randrange( 1, 10).
# + jupyter={"outputs_hidden": false}
# Example 1 of import
# %reset -f
import random
low = 1 ; high = 10
random.randrange(low,high) #generate random number in range low to high
# + jupyter={"outputs_hidden": false}
# Example 2 of import
# %reset -f
import random as r
low = 1 ; high = 10
r.randrange(low,high)
# + jupyter={"outputs_hidden": false}
# Example 3 of import
# %reset -f
from random import randrange
low = 1 ; high = 10
randrange(low,high)
# -
# The modules that come with Python are extensive and listed at
# https://docs.python.org/3/py-modindex.html.
# There are also other modules that can be downloaded and used
# (just like user defined modules below).
# In these labs we are building primitive codes to learn how to code and how to create algorithms.
# For many practical cases you will want to load a well-tested package to accomplish the tasks.
#
# That exercise is saved for the end of the document.
#
# ## User-Built
# We can define our own functions in Python and reuse them throughout the program.
# The syntax for defining a function is:
#
# def functionName( argument ):
# code detailing what the function should do
# note the colon above and indentation
# ...
# ...
# return [expression]
#
# The keyword `def` tells the program that the indented code from the next line onwards is
# part of the function.
# The keyword `return `tells the program to return an answer from the
# function.
# There can be multiple return statements in a function.
# Once the function executes
# a return statement, the program exits the function and continues with *its* next executable
# statement.
# If the function does not need to return any value, you can omit the return
# statement.
#
# Functions can be pretty elaborate; they can search for things in a list, determine variable
# types, open and close files, read and write to files.
#
# To get started we will build a few really
# simple mathematical functions; we will need this skill in the future anyway, especially in
# scientific programming contexts.
# ### User-built within a Code Block
# For our first function we will code $$f(x) = x\sqrt{1 + x}$$ into a function named `dusty()`.
#
# When you run the next cell, all it does is prototype the function (defines it), nothing happens until we use the function.
# + jupyter={"outputs_hidden": false}
def dusty(x) :
temp = x * ((1.0+x)**(0.5)) # don't need the math package
return temp
# the function should make the evaluation
# store in the local variable temp
# return contents of temp
# + jupyter={"outputs_hidden": false}
# + cocalc={"outputs": {"0": {"name": "input", "opts": {"password": false, "prompt": "enter a numeric value"}, "output_type": "stream", "value": "0"}}} jupyter={"outputs_hidden": false}
# wrapper to run the dusty function
yes = 0
while yes == 0:
xvalue = input('enter a numeric value')
try:
xvalue = float(xvalue)
yes = 1
except:
print('enter a bloody number! Try again \n')
# call the function, get value , write output
yvalue = dusty(xvalue)
print('f(',xvalue,') = ',yvalue) # and we are done
# -
# ## Example
#
# Create the AVERAGE function for three values and test it for these values:
# - 3,4,5
# - 10,100,1000
# - -5,15,5
# + jupyter={"outputs_hidden": false}
def AVERAGE3(x,y,z) : #define the function "AVERAGE3"
Ave = (x+y+z)/3 #computes the average
return Ave
# + jupyter={"outputs_hidden": false}
print(AVERAGE3(3,4,5))
print(AVERAGE3(10,100,1000))
print(AVERAGE3(-5,15,5))
# -
# ## Example
#
# Create the FC function to convert Fahrenhiet to Celsius and test it for these values:
# - 32
# - 15
# - 100
#
# *hint: Formula-(°F − 32) × 5/9 = °C
# + jupyter={"outputs_hidden": false}
def FC(x) : #define the function "AVERAGE3"
C = (x - 32)*5/9
return C
# + jupyter={"outputs_hidden": false}
print(FC(32))
print(FC(15))
print(FC(100))
# -
# ## Exercise 1
#
# Create the function $$f(x) = e^x - 10 cos(x) - 100$$ as a function (i.e. use the `def` keyword)
#
# def name(parameters) :
# operations on parameters
# ...
# ...
# return (value, or null)
#
# Then apply your function to the value.
#
# Use your function to complete the table below:
#
# | x | f(x) |
# |---:|---:|
# | 0.0 | |
# | 1.50 | |
# | 2.00 | |
# | 2.25 | |
# | 3.0 | |
# | 4.25 | |
#
# + jupyter={"outputs_hidden": false}
from math import exp # package that contains exp function
from math import cos # package that contains cosine functionسفق
def gollum(x) : #define the function "gollum"
precious = exp(x) - 10*cos(x) -100 #as stated by the question
return precious
# + jupyter={"outputs_hidden": false}
from prettytable import PrettyTable #package that contains the PrettyTable function
t = PrettyTable(['x', 'y']) #Define an empty table
x = [0.0,1.5,2.0,2.25,3.0,4.25] #the list of x values according to the table
for i in range(0,6,1): #a counter to go through the list x
t.add_row([x[i], gollum(x[i])]) #for each x value, fill one row in table t with the value of x and the value of gollum function of that x
print(t)
# -
# ## Variable Scope
# An important concept when defining a function is the concept of variable scope.
# Variables defined inside a function are treated differently from variables defined outside.
# Firstly, any variable declared within a function is only accessible within the function.
# These are known as local variables.
#
# In the `dusty()` function, the variables `x` and `temp` are local to the function.
# Any variable declared outside a function in a main program is known as a program variable
# and is accessible anywhere in the program.
#
# In the example, the variables `xvalue` and `yvalue` are program variables (global to the program; if they are addressed within a function, they could be operated on.)
# Generally we want to protect the program variables from the function unless the intent is to change their values.
# The way the function is written in the example, the function cannot damage `xvalue` or `yvalue`.
#
# If a local variable shares the same name as a program variable, any code inside the function is
# accessing the local variable. Any code outside is accessing the program variable
# ### As Separate Module/File
#
# In this section we will invent the `neko()` function, export it to a file, so we can reuse it in later notebooks without having to retype or cut-and-paste. The `neko()` function evaluates:
#
# $$f(x) = x\sqrt{|(1 + x)|}$$
#
# Its the same as the dusty() function, except operates on the absolute value in the wadical.
#
# 1. Create a text file named "mylibrary.txt"
# 2. Copy the neko() function script below into that file.
#
# def neko(input_argument) :
# import math #ok to import into a function
# local_variable = input_argument * math.sqrt(abs(1.0+input_argument))
# return local_variable
#
#
# 4. rename mylibrary.txt to mylibrary.py
# 5. modify the wrapper script to use the neko function as an external module
# + cocalc={"outputs": {"0": {"name": "input", "opts": {"password": false, "prompt": "enter a numeric value"}, "output_type": "stream", "value": "2.2"}}} jupyter={"outputs_hidden": false}
# wrapper to run the neko function
import mylibrary
yes = 0
while yes == 0:
xvalue = input('enter a numeric value')
try:
xvalue = float(xvalue)
yes = 1
except:
print('enter a bloody number! Try again \n')
# call the function, get value , write output
yvalue = mylibrary.neko(xvalue)
print('f(',xvalue,') = ',yvalue) # and we are done
# -
# In JupyterHub environments, you may discover that changes you make to your external python file are not reflected when you re-run your script; you need to restart the kernel to get the changes to actually update. The figure below depicts the notebook, external file relatonship
#
# 
#
#
# * Future version - explain absolute path
# ## Rudimentary Graphics
#
# Graphing values is part of the broader field of data visualization, which has two main
# goals:
#
# 1. To explore data, and
# 2. To communicate data.
#
# In this subsection we will concentrate on introducing skills to start exploring data and to
# produce meaningful visualizations we can use throughout the rest of this notebook.
# Data visualization is a rich field of study that fills entire books.
# The reason to start visualization here instead of elsewhere is that with functions plotting
# is a natural activity and we have to import the matplotlib module to make the plots.
#
# The example below is code adapted from Grus (2015) that illustrates simple generic
# plots. I added a single line (label the x-axis), and corrected some transcription
# errors (not the original author's mistake, just the consequence of how the API handled the
# cut-and-paste), but otherwise the code is unchanged.
# + jupyter={"outputs_hidden": false}
# python script to illustrate plotting
# CODE BELOW IS ADAPTED FROM:
# <NAME> (2015-04-14). Data Science from Scratch: First Principles with Python
# (Kindle Locations 1190-1191). O'Reilly Media. Kindle Edition.
#
from matplotlib import pyplot as plt # import the plotting library from matplotlibplt.show()
years = [1950, 1960, 1970, 1980, 1990, 2000, 2010] # define one list for years
gdp = [300.2, 543.3, 1075.9, 2862.5, 5979.6, 10289.7, 14958.3] # and another one for Gross Domestic Product (GDP)
plt.plot( years, gdp, color ='green', marker ='o', linestyle ='solid') # create a line chart, years on x-axis, gdp on y-axis
# what if "^", "P", "*" for marker?
# what if "red" for color?
# what if "dashdot", '--' for linestyle?
plt.title("Nominal GDP")# add a title
plt.ylabel("Billions of $")# add a label to the x and y-axes
plt.xlabel("Year")
plt.show() # display the plot
# -
# Now lets put the plotting script into a function so we can make line charts of any two numeric lists
# + jupyter={"outputs_hidden": false}
def plotAline(list1,list2,strx,stry,strtitle): # plot list1 on x, list2 on y, xlabel, ylabel, title
from matplotlib import pyplot as plt # import the plotting library from matplotlibplt.show()
plt.plot( list1, list2, color ='green', marker ='o', linestyle ='solid') # create a line chart, years on x-axis, gdp on y-axis
plt.title(strtitle)# add a title
plt.ylabel(stry)# add a label to the x and y-axes
plt.xlabel(strx)
plt.show() # display the plot
return #null return
# + jupyter={"outputs_hidden": false}
# wrapper
years = [1950, 1960, 1970, 1980, 1990, 2000, 2010] # define two lists years and gdp
gdp = [300.2, 543.3, 1075.9, 2862.5, 5979.6, 10289.7, 14958.3]
print(type(years[0]))
print(type(gdp[0]))
plotAline(years,gdp,"Year","Billions of $","Nominal GDP")
# -
# ## Example
# Use the plotting script and create a function that draws a straight line between two points.
# + jupyter={"outputs_hidden": false}
def Line():
from matplotlib import pyplot as plt # import the plotting library from matplotlibplt.show()
x1 = input('Please enter x value for point 1')
y1 = input('Please enter y value for point 1')
x2 = input('Please enter x value for point 2')
y2 = input('Please enter y value for point 2')
xlist = [x1,x2]
ylist = [y1,y2]
plt.plot( xlist, ylist, color ='orange', marker ='*', linestyle ='solid')
#plt.title(strtitle)# add a title
plt.ylabel("Y-axis")# add a label to the x and y-axes
plt.xlabel("X-axis")
plt.show() # display the plot
return #null return
# + cocalc={"outputs": {"0": {"name": "input", "opts": {"password": false, "prompt": "Please enter x value for point 1"}, "output_type": "stream", "value": "1"}, "1": {"name": "input", "opts": {"password": false, "prompt": "Please enter y value for point 1"}, "output_type": "stream"}}} jupyter={"outputs_hidden": false}
Line()
# -
# ## Example- Lets have some fun!
# Copy the wrapper script for the `plotAline()` function, and modify the copy to create a plot of
# $$ x = 16sin^3(t) $$
# $$ y = 13cos(t) - 5cos(2t) - 2cos(3t) - cos(4t) $$
# for t raging from [0,2$\Pi$] (inclusive).
#
# Label the plot and the plot axes.
#
# + jupyter={"outputs_hidden": false}
from matplotlib import pyplot as plt # import the plotting library from matplotlibplt.show()
import numpy as np # import NumPy: for large, multi-dimensional arrays and matrices, along with high-level mathematical functions to operate on these arrays.
pi = np.pi #pi value from the np package
t= np.linspace(0,2*pi,360)# the NumPy function np.linspace is similar to the range()
x = 16*np.sin(t)**3
y = 13*np.cos(t) - 5*np.cos(2*t) - 2*np.cos(3*t) - np.cos(4*t)
plt.plot( x, y, color ='purple', marker ='.', linestyle ='solid')
plt.ylabel("Y-axis")# add a label to the x and y-axes
plt.xlabel("X-axis")
plt.axis('equal') #sets equal axis ratios
plt.title("A Hopeless Romantic's Curve")# add a title
plt.show() # display the plot
# -
# ## Exercise 2
# Copy the wrapper script for the `plotAline()` function, and modify the copy to create a plot of
# $$ y = x^2 $$
# for x raging from 0 to 9 (inclusive) in steps of 1.
#
# Label the plot and the plot axes.
#
# + jupyter={"outputs_hidden": false}
xxx = [] # null list
yyy = [] # null list
for i in range(0,10):
xxx.append(1.0*i) #float i and append to the list
for i in range(0,10):
yyy.append(xxx[i]**2)
plotAline(xxx,yyy,"X","Y","Plot of Y = X^2 ")
# + jupyter={"outputs_hidden": false}
| 1-Lessons/Lesson05/dev_src/Lab5/src/fp-src/Lab5-dev_Fp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib
#matplotlib.use('Agg')
# %matplotlib tk
# %autosave 180
# %load_ext autoreload
# %autoreload 2
import matplotlib.pyplot as plt
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
import matplotlib.cm as cm
from matplotlib import gridspec
import parmap
import numpy as np
import pandas as pd
import os
import shutil
import cv2
import scipy.io as sio
# from Specgram import Specgram
import csv
import glob
from numba import jit
from sklearn.svm import SVC # "Support vector classifier"
from utility_DLC import *
from tqdm import tqdm
# manually add labels from DLC
labels = ['left_paw',
'right_paw',
'nose',
'jaw',
'right_ear',
'tongue',
'lever']
colors = ['blue','orange','green','red','magenta','brown']
# +
########################################
########################################
########################################
def get_satiety_scatter_plots(animal_id,
plotting=False):
A = Analyze()
#A.root_dir = '/media/cat/1TB/yuki/'
A.root_dir = '/media/cat/4TBSSD/yuki/'
A.lockout = 1
A.animal_names = ['IA1','IA2','IA3','IJ1','IJ2','AQ2']
A.animal_ids = np.arange(len(A.animal_names))
#
import scipy.stats
width1 = 1
width2 = 10
linewidth = 3
offset1 = 200
offset2 = 10
offset3 = 250
fontsize = 16
dotsize = 2
alpha = .15
bins = np.arange(0,1300,width1)
pvals = [0.05, 0.01, 0.001, 0.0001, 0.00001]
cmap = matplotlib.cm.get_cmap('magma')
#
animal_name = A.animal_names[animal_id]
#for animal_id in ['AQ2']:
print ("animal_nam: ", animal_name)
fname = A.root_dir + animal_name + '/tif_files/satiety_consumption_all_sessions.npz'
#print (fname)
d = np.load(fname,
allow_pickle=True)
s02 = d['s02']
s04 = d['s04']
#############################################
#############################################
#############################################
if plotting:
fig=plt.figure(figsize=(10,10))
img1 = []
img2 = []
all_sums1 = []
all_sums2 = []
vsums1 = []
vsums2 = []
#
all1 = []
all2 = []
for k in range(len(s04)):
temp = s04[k]
hist = np.histogram(temp, bins = bins)
temp2 = s02[k]
hist2 = np.histogram(temp2, bins = bins)
all1.append(temp)
all2.append(temp2)
#
s1 = hist[0].sum()
if s1==0:
h1 = np.zeros(hist[0].shape[0])#+0.01
else:
h1 = hist[0]/np.max(hist[0])
#
s2 = hist2[0].sum()
if s2==0:
h2 = np.zeros(hist2[0].shape[0])#+0.01
else:
h2 = hist2[0]/np.max(hist2[0])
#
if s1 ==0 and s2==0:
continue
#
all_sums1.append(len(temp))
all_sums2.append(len(temp2))
if True:
sigma = 3
h1 = scipy.ndimage.gaussian_filter1d(h1, sigma)
h2 = scipy.ndimage.gaussian_filter1d(h2, sigma)
#
vsums1.extend(temp)
vsums2.extend(temp2)
#
img1.append(h1)
img2.append(h2)
all_sums1 = np.array(all_sums1)
img1 = np.vstack(img1)
rewarded_scatter = all_sums1.copy()
###################################################
############ CODE 04 RASTER/IMG PLOT ##############
###################################################
if plotting:
ax=plt.subplot(121)
plt.imshow(img1,
aspect='auto',
cmap='Blues',
interpolation='none')
plt.xticks([])
###################################################
############ CODE 04 VERTICAL HISTOGRAM ###########
###################################################
rewarded_vertical_histogram1 = all_sums1.copy()
if plotting:
plt.plot(1300-all_sums1/np.max(all_sums1)*offset1+ offset3,
np.arange(len(all_sums1)),
linewidth = linewidth,
c='blue',
alpha=.5)
plt.xlim(0,1300+offset3)
###################################################
############ CODE 04 SCATTER PLOT #################
###################################################
vsums1 = np.array(vsums1)
vsums1 = vsums1.flatten()
#print ("vsums1: ", vsums1.shape)
#
bins2 = np.arange(0,1300,width2)
y = np.histogram(vsums1, bins=bins2)
vsums1 = np.float32(y[0])
#
#vsums1 = vsums1/np.nanmax(vsums1)*offset2+len(all_sums1)
x = np.arange(y[1][0],y[1][-1],width2)
y = (vsums1 - np.min(vsums1))/(np.max(vsums1)-np.min(vsums1))*10+len(all_sums1)
#
if plotting:
plt.scatter(x,
y,
linewidth = linewidth,
s=dotsize,
c='blue',
alpha=alpha)
#############################################
############ CODE 04 SCATTER FIT ############
#############################################
coef = np.polyfit(x,y,1)
poly1d_fn = np.poly1d(coef)
if plotting:
plt.plot(x, poly1d_fn(x), '--', c='blue',
linewidth=linewidth)
rewarded_scatter = [x,y]
rewarded_fit = [x, poly1d_fn(x).copy()]
#
slope, intercept, r_value, p_value1, std_err = scipy.stats.linregress(x,y)
#print (r_value, p_value)
title = ''
for k in range(len(pvals)):
if p_value1 < pvals[k]:
title = title+ "*"
#plt.title("Rewarded Pulls "+title,fontsize=fontsize)
if plotting:
plt.title(title,fontsize=fontsize)
plt.ylim(-0.5, len(all_sums1)+offset2+1)
###################################################
################# CODE 02 RASTER IMAGE ############
###################################################
if plotting:
ax=plt.subplot(122)
img2 = np.vstack(img2)
#print ("img2: ", img2.shape)
if plotting:
plt.imshow(img2, aspect='auto',
cmap='Reds',
interpolation='none')
plt.xticks([])
plt.yticks([])
###################################################
################# CODE 02 VERTICAL HIST ###########
###################################################
all_sums2 = np.array(all_sums2)
rewarded_vertical_histogram2 = all_sums2.copy()
if plotting:
plt.plot(1300-all_sums2/np.max(all_sums2)*offset1+ offset3,
np.arange(len(all_sums2)),
linewidth = linewidth,
c='red')
plt.xlim(0,1300+offset3)
###################################################
############ CODE 02 SCATTER ######################
###################################################
vsums2 = np.array(vsums2).flatten()
y = np.histogram(vsums2, bins=bins2)
vsums2 = np.float32(y[0])
#
x = np.arange(y[1][0],y[1][-1],width2)
#y = vsums2+len(all_sums2)+1+offset2
y = (vsums2 - np.min(vsums2))/(np.max(vsums2)-np.min(vsums2))*10+len(all_sums2)
if plotting:
plt.scatter(x,
y,
linewidth = linewidth,
s=dotsize,
c='red',
alpha=alpha)
# FIT
slope, intercept, r_value, p_value2, std_err = scipy.stats.linregress(x,y)
#print (r_value, p_value)
title = ''
for k in range(len(pvals)):
if p_value2 < pvals[k]:
title = title+ "*"
#plt.title("UnRewarded Pulls "+title,fontsize=fontsize)
if plotting:
plt.title(title,fontsize=fontsize)
coef = np.polyfit(x,y,1)
poly1d_fn = np.poly1d(coef)
rewarded_scatter2 = [x,y]
rewarded_fit2 = [x, poly1d_fn(x).copy()]
#
if plotting:
plt.plot(x, poly1d_fn(x), '--', c='red',
linewidth=linewidth)
#
plt.ylim(-0.5, len(all_sums1)+offset2+1)
##############################################
if plotting:
if True:
plt.savefig('/home/cat/'+str(animal_name)+'.png', dpi=300)
plt.close()
else:
plt.show()
all1 = np.array(all1)
all2 = np.array(all2)
return (all1, all2, rewarded_scatter,
rewarded_vertical_histogram1, rewarded_vertical_histogram2,
rewarded_fit, rewarded_scatter, p_value1,
rewarded_fit2, rewarded_scatter2, p_value2)
########################################
########################################
########################################
def generate_satiety_data(A):
#
for animal_id in A.animal_ids:
A.animal_id = animal_id
#
fname_out = os.path.join(A.root_dir, A.animal_id,'tif_files/satiety_consumption_all_sessions.npz')
if os.path.exists(fname_out)==False:
#
A = get_ordered_fnames_stand_alone(A)
#
s04 = []
s02 = []
for session in tqdm(A.sessions):
root_dir = os.path.join(A.root_dir, A.animal_id,'tif_files',session,session)
abstimes, abspositions, abscodes = load_lever_data(root_dir)
#
if len(abstimes)>0:
(starts_04,
starts_04_idx,
starts_02,
starts_02_idx) = find_code04_starts2(abscodes,
abstimes,
abspositions,
A.lockout)
#
s04.append(starts_04)
s02.append(starts_02)
else:
s04.append([])
s02.append([])
np.savez(fname_out,
s04=s04,
s02=s02)
########################################
########################################
########################################
def plot_vertical_histograms(data_type, plot_legend):
plotting = False
animal_names = ['M1','M2','M3','M4','M5','M6']
animal_ids = np.arange(6)
linewidth=5
fig = plt.figure(figsize=(12,12))
#
for animal_id in animal_ids:
(all1, all2, rewarded_scatter,
rewarded_vertical_histogram1, rewarded_vertical_histogram2,
rewarded_fit, rewarded_scatter, p_value1,
rewarded_fit2, rewarded_scatter2, p_value2) = get_satiety_scatter_plots(animal_id, plotting)
if data_type=='rewarded':
y = rewarded_vertical_histogram1
else:
y = rewarded_vertical_histogram2
y = y/np.max(y)
x = np.arange(y.shape[0])
x = x/np.max(x)
plt.plot(x, y+animal_id*1.5,#'--',
linewidth = linewidth,
label=animal_names[animal_id])
plt.plot([0,1],[animal_id*1.5,animal_id*1.5],'--', c='black',alpha=.5)
plt.xticks([])
plt.yticks([])
plt.xlim(0,1)
plt.ylim(0,7*1.5)
if plot_legend:
plt.legend(fontsize=8,ncol=3, loc='upper right')
#
if True:
plt.savefig('/home/cat/vertical.png',dpi=300)
plt.close()
else:
plt.show()
########################################
########################################
########################################
def plot_horizontal_scatter(data_type,
plot_legend,
animal_id):
#plotting = False
animal_names = ['M1','M2','M3','M4','M5','M6']
#animal_names = animal_names[animal_id]
if data_type == 'rewarded':
color = 'blue'
else:
color = 'red'
#
linewidth=5
scatter_size = 10
scale = 15
#fig = plt.figure() #figsize=(3,10))
#
animal_ids = np.arange(6)
animal_ids = [animal_id]
#animal_ids = [0]
for animal_id in animal_ids:
(all1, all2, rewarded_scatter,
rewarded_vertical_histogram1, rewarded_vertical_histogram2,
rewarded_fit, rewarded_scatter, p_value1,
rewarded_fit2, rewarded_scatter2, p_value2) = get_satiety_scatter_plots(animal_id, plotting)
if data_type=='rewarded':
d1 = rewarded_fit
d2 = rewarded_scatter
p_value = p_value1
else:
d1 = rewarded_fit2
d2 = rewarded_scatter2
p_value = p_value2
pvals = [0.05, 0.01, 0.001, 0.0001, 0.00001]
title = ''
for k in range(len(pvals)):
if p_value < pvals[k]:
title = title+ "*"
########################################
########### PLOT FIT ###################
########################################
d1[1] = (d1[1]-np.min(d2[1]))#/(np.max(d1[1])-np.min(d1[1]))
plt.plot(d1[0],d1[1], #+animal_id*scale,
label=animal_names[animal_id]+ " "+title,
c=color,
linewidth=linewidth)
########################################
########### PLOT SCATTER ###############
########################################
d2[1] = (d2[1]-np.min(d2[1]))#/(np.max(d2[1])-np.min(d2[1]))
plt.scatter(d2[0],d2[1], #+animal_id*scale,
linestyle='--',
alpha=.2,
# edgecolor=colors[animal_id],
# facecolor = colors[animal_id],
s=scatter_size,
c=color,
linewidth=linewidth)
plt.plot([0,1300] , [np.mean(d2[1]), np.mean(d2[1])],
'--', c='black',linewidth=2,
alpha=.5)
#
plt.xticks([])
plt.yticks([])
plt.xlim(0,1300)
plt.title(title, fontsize=14, y=.9)
#plt.ylim(0,(animal_id+2)*scale)
if plot_legend:
plt.legend(fontsize=3,ncol=2, loc='upper right')
def compute_trends_intra_session(data_type,
animal_id,
plot_legend,
plot_data,
plot_images):
animal_names = ['M1','M2','M3','M4','M5','M6']
#
linewidth=4
scatter_size = 2
scale = 15
if plot_data:
fig = plt.figure(figsize=(10,5))
#
#animal_ids = np.arange(6)
#animal_ids = [2]
#
#plt.subplot(2,3,animal_id+1)
(all1, all2, rewarded_scatter,
rewarded_vertical_histogram1, rewarded_vertical_histogram2,
rewarded_fit, rewarded_scatter, p_value1,
rewarded_fit2, rewarded_scatter2, p_value2) = get_satiety_scatter_plots(animal_id, plotting)
if data_type=='rewarded':
d1 = rewarded_fit
d2 = rewarded_scatter
p_value = p_value1
all_ = all1
else:
d1 = rewarded_fit2
d2 = rewarded_scatter2
p_value = p_value2
all_ = all2
print ("lend all ", len(all_))
# LOOP OVER WINDOWS OF DATA:
window = 10
width2 = 10
ctr=0
for k in range(0,len(all1)-window, 1):
ctr+=1
if data_type == 'rewarded':
colors = plt.cm.Blues
cmap=plt.cm.get_cmap('Blues')
else:
colors = plt.cm.Reds
cmap=plt.cm.get_cmap('Reds')
colors=cmap(np.arange(len(all_)-window+10)/(len(all_)-window+10))
coefs = []
n_trials = []
for k in range(0,len(all_)-window, 1):
data = np.hstack(all_[k:k+window])
n_trials.append(data.shape[0])
#
bins2 = np.arange(0,1300,width2)
y = np.histogram(data, bins=bins2)
xx = y[1][1:]
yy = y[0]
# print (xx.shape, yy.shape)
# FIT
coef = np.polyfit(xx,yy,1)
coefs.append(coef)
poly1d_fn = np.poly1d(coef)
#
yy = poly1d_fn(xx)
yy = yy-yy[0]
if plot_data:
plt.plot(xx, yy,
c=colors[k],
linewidth=linewidth)
if plot_data:
plt.plot([0,1300],[0,0],'--',c='black',
linewidth=2,
alpha=.5)
#
plt.title(animal_names[animal_id])
#plt.ylim(-3, 4.5)
plt.xlim(0, 1300)
plt.xticks([])
plt.yticks([])
# if plot_legend:
# plt.legend(fontsize=3,ncol=2, loc='upper right')
#
cax = fig.add_axes([0.95, 0.2, 0.02, 0.6])
cb = matplotlib.colorbar.ColorbarBase(cax, cmap=cmap)
plt.suptitle(data_type)
if False:
plt.savefig('/home/cat/vertical.png',dpi=300)
plt.close()
else:
plt.show()
coefs = np.array(coefs)
return (coefs, n_trials)
# +
################################################################
############ COMPUTE SATIETY SCATTER DATA AND SAVE #############
################################################################
#
A = Analyze()
A.root_dir = '/media/cat/4TBSSD/yuki/'
A.lockout = 1
A.animal_ids = ['IA1','IA2','IA3','IJ1','IJ2','AQ2']
generate_satiety_data(A)
# +
######################################################
###### GENERATE AND VISUALIZE SATIETY DATA ###########
######################################################
# FIG 1 E,F ...
animal_id = 0
plotting = True
(all1, all2, rewarded_scatter,
rewarded_vertical_histogram1, rewarded_vertical_histogram2,
rewarded_fit, rewarded_scatter, p_value1,
rewarded_fit2, rewarded_scatter2, p_value2) = get_satiety_scatter_plots(animal_id, plotting)
#
# +
########################################################
######## PLOT ALL VERTICAL HISTOGRAM PLOTS #############
########################################################
# FIG 1 E,F ...
#
data_type= 'rewarded' # options: 'rewarded', 'unrewarded'
plot_legend = False
plot_vertical_histograms(data_type, plot_legend)
# +
########################################################
######## SHOW ALL HORIZONTAL HISTOGRAMS AND FITS #######
########################################################
# FIG 1 E,F ...
data_type= 'rewarded' # options: 'rewarded', 'unrewarded'
plot_legend = False
animal_id = 1
animal_ids = np.arange(6)
fig = plt.figure(figsize=(1,10))
for animal_id in animal_ids:
ax=plt.subplot(6,1,animal_id+1)
plot_horizontal_scatter(data_type,
plot_legend,
animal_id)
#
if True:
plt.savefig('/home/cat/horizontal.png',dpi=100)
plt.close()
else:
plt.show()
# +
###################################################################
######## SHOW TIME TRENDS IN HORIZONTAL HISTOGRAMS AND FITS #######
###################################################################
# FIG 1 E,F ...
#
data_type= 'rewarded' # options: 'rewarded', 'unrewarded'
plot_legend = True
plot_data = True
plot_images = False
animal_id = 5
coefs, n_trials = compute_trends_intra_session(data_type,
animal_id,
plot_legend,
plot_data,
plot_images)
# +
################################################
################################################
################################################
data_type= 'rewarded' # options: 'rewarded', 'unrewarded'
plot_legend = True
plot_data = False
fig = plt.figure(figsize=(1,10))
for animal_id in animal_ids:
coefs, n_trials = compute_trends_intra_session(data_type,
animal_id,
plot_legend,
plot_data,
plot_images)
n_trials = np.array(n_trials)
################################################
################################################
################################################
ax=plt.subplot(6,1,animal_id+1)
x = np.arange(coefs.shape[0])
plt.scatter(x,
coefs[:,0],
c='red')
plt.plot([x[0],x[-1]],[0,0],'--', c='black')
plt.xlim(x[0],x[-1])
plt.xticks([])
plt.yticks([])
#
if True:
plt.savefig('/home/cat/trends.png',dpi=100)
plt.close()
else:
plt.show()
# -
# +
d = np.load('/media/cat/4TBSSD/yuki/IA1/tif_files/IA1pm_Feb1_30Hz/IA1pm_Feb1_30Hz_3secNoMove_movements.npz',allow_pickle=True)
quiet = d['all_quiescent']
print (quiet[:10])
print ('')
features = d['feature_quiescent']
for k in range(len(features)):
print (k,features[k][:10])
print ('')
#all_movs = d['all_feature_movements']
#print (all_movs)
# +
#############################################
######### MAKE A MOVIE WITH DLC TRACES ######
#############################################
#
fname_traces = ''
traces = np.load(fname_traces)
print (traces.shape)
#
fname_video = ''
#
start = 478*15
end = start+20*15
make_video_dlc(traces,
movements,
fname_video,
start,
end)
# +
#########################################
##### GET MOVEMENT INITIATION ARRAYS ####
#########################################
window = 4.0 # LOCKOUT WINDOW REQUESTED
plotting = False
fname_traces = '/media/cat/4TBSSD/yuki/IA1/tif_files/IA1pm_Feb1_30Hz/IA1pm_Feb1_30HzDLC_resnet50_yuki_leverJul21shuffle1_38000.npy'
root_dir = '/media/cat/4TBSSD/yuki/IA1/'
(starts_arrays_lockout, starts_arrays_all, traces_filtered) = get_starts_arrays2(
window,
fname_traces,
labels,
plotting)
# -
starts_arrays_all= np.load('/home/cat/movements.npy')
# +
# extract quiet periods
length_quiet_period = 3 # seconds
quiet_periods = find_quiet_periods_specific_length_and_first_feature(starts_arrays_all,
length_quiet_period
)
print (quiet_periods)
#
# +
# FIND DISTRIBUTION OF QUIESCENT PERIODS
# window=3
# plotting=False
# _, durations = quiet_periods_histogram(starts_arrays, plotting)
#print ("Durations; ",durations)
fig=plt.figure()
y = np.histogram(durations, bins=np.arange(0,20,1))
#plt.bar(y[1][:-1]+0.5,y[0],0.9)
plt.plot(y[1][:-1]+0.5,y[0])
plt.xlim(0,20)
#print (np.cumsum(y[0][10:]))
#plt.plot(y[1][10:-1],np.cumsum(y[0][10:]),linewidth=5,c='blue')
#plt.ylim(0,20)
plt.xticks(fontsize=40)
plt.ylabel("# of periods", fontsize=40)
plt.xlabel("Duration of quiescent period (sec)",fontsize=40)
plt.yticks(fontsize=40)
plt.semilogy()
plt.show()
# +
# #temp_cov = np.cov(data, data)
# temp_cov= np.matmul(data[None].T, data[None])
# sig = np.sqrt(np.diag(temp_cov))
# temp_cov = temp_cov/(sig[:,None]*sig[None])
# -
plt.imshow(temp_cov)
plt.show()
data = np.load('/home/cat/Downloads/2018-11-12-1_vision_spike_train(2).npy',allow_pickle=True)
print (data)
data= np.load('/home/cat/Downloads/2018-12-10-4_vision_spike_train(2).npy',allow_pickle=True)
print (data.shape)
print (data)
data = np.load('/media/cat/4TBSSD/yuki/IA1/tif_files/IA1pm_Feb1_30Hz/IA1pm_Feb1_30Hz_left_paw_trial_ROItimeCourses_15sec_pca_0.95.npy')
print (data.shape)
| Fig1_lever_pull_analysis/Fig1_E,H_histogram_satiety_signals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import json
import pandas as pd
import math
# %matplotlib inline
# +
# reading the JSON data using json.load()
file = './data/GoogleLocation.json'
with open(file) as train_file:
data_json = json.load(train_file)
df = pd.DataFrame({'raw': data_json['locations']})
# +
# Extract the base data
df['timestampMs'] = df.apply(lambda row: int(row['raw']['timestampMs']), axis=1)
df['latitudeE7'] = df.apply(lambda row: row['raw']['latitudeE7'], axis=1)
df['longitudeE7'] = df.apply(lambda row: row['raw']['longitudeE7'], axis=1)
df['accuracy'] = df.apply(lambda row: row['raw']['accuracy'], axis=1)
# Derivatives
df['date'] = pd.to_datetime(df['timestampMs'], unit='ms')
df['year'] = df.apply(lambda row: row['date'].year, axis=1)
df['month'] = df.apply(lambda row: row['date'].month, axis=1)
df['day'] = df.apply(lambda row: row['date'].day, axis=1)
df['hour'] = df.apply(lambda row: row['date'].hour, axis=1)
# -
df_part = pd.DataFrame(df[(df['accuracy'] < 100) & (df['year']==2018) & (df['month']==7) & (df['day']==17)])
# +
grouped = df.groupby(['year','month','day','hour']).agg({'latitudeE7': ['median', 'var', 'count'], 'longitudeE7': 'var'})
# # +epsilon for stability
grouped['lat_var_log'] = grouped.apply(lambda row: math.log(row['latitudeE7']['var']+0.0001), axis=1)
grouped
# -
grouped[grouped['latitudeE7']['count'] == 1]['latitudeE7']['median'].hist(bins=1000)
grouped.hist(['lat_var_log'], bins=100, figsize=(16, 10))
# +
df_part['dx'] = df_part.longitudeE7.diff()
df_part['dy'] = df_part.latitudeE7.diff()
df_part['dot_prod'] = df_part.dx * df_part.dx.shift() + df_part.dy * df_part.dy.shift()
df_part['movement'] = df_part['dot_prod'].rolling(5, center=True).mean()
df_part.head(60)
# -
df_part.plot(x='timestampMs', y='movement')
# +
df_part['gps_var'] = df_part['latitudeE7'].rolling(10, center=True).var()
df_part.plot(x='timestampMs', y='gps_var')
# -
df_part.plot(x='timestampMs', y='latitudeE7')
df_part.plot(x='timestampMs', y='longitudeE7')
| gps/GoogleLocation4-Segmentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: StyleGAN
# language: python
# name: stylegan
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Load-Network" data-toc-modified-id="Load-Network-1"><span class="toc-item-num">1 </span>Load Network</a></span></li><li><span><a href="#Style-Mixing" data-toc-modified-id="Style-Mixing-2"><span class="toc-item-num">2 </span>Style Mixing</a></span></li><li><span><a href="#Latents-Transition/Morphing" data-toc-modified-id="Latents-Transition/Morphing-3"><span class="toc-item-num">3 </span>Latents Transition/Morphing</a></span></li><li><span><a href="#Explore-PSI" data-toc-modified-id="Explore-PSI-4"><span class="toc-item-num">4 </span>Explore PSI</a></span></li><li><span><a href="#Explore-Latents-Indexes" data-toc-modified-id="Explore-Latents-Indexes-5"><span class="toc-item-num">5 </span>Explore Latents Indexes</a></span></li></ul></div>
# -
# Playground for experiments with StyleGANv2 latents.
# Includes interactive style mixing, latents interpolation or morphing and latents tweaking.
# +
from pathlib import Path
import matplotlib.pyplot as plt
import numpy as np
import sys
import os
from datetime import datetime
from tqdm import tqdm
import imageio
from ipywidgets import interact, interact_manual
from IPython.display import display
import warnings
warnings.filterwarnings('ignore', category=FutureWarning)
# ffmpeg installation location, for creating videos
plt.rcParams['animation.ffmpeg_path'] = str(Path.home() / "Documents/dev_tools/ffmpeg-20190623-ffa64a4-win64-static/bin/ffmpeg.exe")
# %load_ext autoreload
# %autoreload 2
# StyleGAN Utils
from stylegan_utils import load_network, gen_image_fun, synth_image_fun, create_video
from stylegan_utils import load_latents, get_ipywidget_elements, load_directions
# StyleGAN2 Repo
sys.path.append(os.path.join(os.pardir, 'stylegan2encoder'))
import projector
import training.dataset
# Data Science Utils
sys.path.append(os.path.join(os.pardir, 'data-science-learning'))
from ds_utils import generative_utils
# -
res_dir = Path.home() / 'Documents/generated_data/stylegan'
# # Load Network
# +
MODELS_DIR = Path("C:/Users/User/Documents/models/stylegan2")
MODEL_NAME = 'drawing2_1024'
SNAPSHOT_NAME = 'network-snapshot-002048'
Gs, Gs_kwargs, noise_vars = load_network(str(MODELS_DIR / MODEL_NAME / SNAPSHOT_NAME) + '.pkl')
Z_SIZE = Gs.input_shape[1:][0]
IMG_SIZE = Gs.output_shape[2:]
IMG_SIZE
# -
# # Style Mixing
# +
# used when saving the currently displayed image
current_displayed_latents = None
current_displayed_img = None
# save directory
save_dir = res_dir / 'projection' / MODEL_NAME / SNAPSHOT_NAME / "picked"
save_dir.mkdir(parents=True, exist_ok=True)
# -
def mix_latents(latents_1, latents_2, layers_idxs, alpha=1.):
latents_1 = load_latents(latents_1)
latents_2 = load_latents(latents_2)
assert latents_1.shape == latents_2.shape
# crossover option, from latents_1 to latents_2
mixed_latents = latents_2.copy()
mixed_latents[layers_idxs] = latents_1[layers_idxs] * alpha + mixed_latents[layers_idxs] * (1.-alpha)
return mixed_latents
# +
# util to get names of various latents
target_dir_name = ''
data_dir = res_dir / 'projection' / MODEL_NAME / SNAPSHOT_NAME / target_dir_name
entries = [p.name for p in data_dir.glob("*") if p.is_dir()]
entries.remove('tfrecords')
entries2 = np.arange(30)
# load directions
directions_dir = MODELS_DIR / MODEL_NAME / 'ganspace/directions_01'
directions = load_directions(directions_dir, is_ganspace=True)
print(directions.keys())
# +
# %matplotlib notebook
# Setup plot image
fig, ax = plt.subplots(figsize=(9, 9))
fig.subplots_adjust(left=0, right=1, bottom=0, top=1, hspace=0, wspace=0)
plt.axis('off')
im = ax.imshow(gen_image_fun(Gs, np.random.randn(1, Z_SIZE), Gs_kwargs, noise_vars))
def on_button_clicked(b):
global current_displayed_latents, current_displayed_img
timestamp = datetime.now().strftime("%Y%m%d_%H%M%S")
plt.imsave(save_dir / (timestamp + '.png'), current_displayed_img)
np.save(save_dir / (timestamp + '.npy'), current_displayed_latents)
def i_style_mixing(entry1, entry2, layers_idxs, alpha, directions_coeffs, directions_layers):
latents_1 =res_dir / 'projection' / MODEL_NAME / SNAPSHOT_NAME / target_dir_name / f'{entry1}' / 'image_latents1000.npy'
latents_2 = res_dir / 'projection' / MODEL_NAME / SNAPSHOT_NAME / 'rand_gen/psi05' / f'{entry2}.npy'
layers_idxs=np.arange(layers_idxs[0], layers_idxs[1])
# compute mixed latents
mixed_latents = mix_latents(latents_2, latents_1, layers_idxs=layers_idxs, alpha=alpha)
# add directions
for direction_name, coeff in directions_coeffs.items():
if coeff != 0.:
d_layers_idxs = directions_layers[direction_name]
mixed_latents[d_layers_idxs] = mixed_latents[d_layers_idxs] + (coeff * directions[direction_name])
# generate image
gen_image = synth_image_fun(Gs, mixed_latents[np.newaxis, :, :], Gs_kwargs, randomize_noise=True)
# store in case we want to export results from widget
global current_displayed_latents, current_displayed_img
current_displayed_latents = mix_latents
current_displayed_img = gen_image
im.set_data(gen_image)
# ipywdigets setup
display_element = get_ipywidget_elements(i_style_mixing, on_button_clicked, entries, entries2, list(directions.keys()))
display(display_element)
# -
# # Latents Transition/Morphing
#PLOT_IMG_SHAPE = (512, 512, 3)
PLOT_IMG_SHAPE = (IMG_SIZE[0], IMG_SIZE[1], 3)
# +
render_dir = res_dir / MODEL_NAME / SNAPSHOT_NAME / "explore_latent"
nb_samples = 2
nb_transition_frames = 450
nb_frames = min(450, (nb_samples-1)*nb_transition_frames)
psi=1
# run animation
for i in range(0, 2):
# setup the passed latents
z_s = np.random.randn(nb_samples, Z_SIZE)
#latents = Gs.components.mapping.run(z_s, None)
passed_latents=z_s
animate_latent_transition(latent_vectors=passed_latents,
#gen_image_fun=synth_image_fun,
gen_image_fun=lambda latents : gen_image_fun(Gs, latents, Gs_kwargs, truncation_psi=psi),
gen_latent_fun=lambda z_s, i: gen_latent_linear(passed_latents, i, nb_transition_frames),
img_size=PLOT_IMG_SHAPE,
nb_frames=nb_frames,
render_dir=render_dir / "transitions")
# -
# # Explore PSI
#PLOT_IMG_SHAPE = (512, 512, 3)
PLOT_IMG_SHAPE = (IMG_SIZE[0], IMG_SIZE[1], 3)
# +
render_dir = res_dir / MODEL_NAME / SNAPSHOT_NAME / 'explore_latent'
nb_samples = 20
nb_transition_frames = 24
nb_frames = min(450, (nb_samples-1)*nb_transition_frames)
# setup the passed latents
z_s = np.random.randn(nb_samples, Z_SIZE)
#latents = Gs.components.mapping.run(z_s, None)
passed_latents = z_s
# run animation
#[2., 1.5, 1., 0.7, 0.5, 0., -0.5, -0.7, -1., -1.5, -2.]
for psi in np.linspace(-0.5, 1.5, 9):
animate_latent_transition(latent_vectors=passed_latents,
#gen_image_fun=synth_image_fun,
gen_image_fun=lambda latents : gen_image_fun(Gs, latents, Gs_kwargs, truncation_psi=psi),
gen_latent_fun=lambda z_s, i: gen_latent_linear(passed_latents, i, nb_transition_frames),
img_size=PLOT_IMG_SHAPE,
nb_frames=nb_frames,
render_dir=render_dir / 'psi',
file_prefix='psi{}'.format(str(psi).replace('.', '_')[:5]))
# -
# # Explore Latents Indexes
#PLOT_IMG_SHAPE = (512, 512, 3)
PLOT_IMG_SHAPE = (IMG_SIZE[0], IMG_SIZE[1], 3)
# +
render_dir = res_dir / MODEL_NAME / SNAPSHOT_NAME / "explore_latent"
nb_transition_frames = 48
# random list of z vectors
#rand_idx = np.random.randint(len(X_train))
z_start = np.random.randn(1, Z_SIZE)
#dlatents = Gs.components.mapping.run(z_start, None, dlatent_broadcast=None)
#vals = np.linspace(-2., 2., nb_transition_frames)
nb_styles = dlatents.shape[0]
stylelatent_vals= np.random.randn(nb_transition_frames, Z_SIZE) + np.linspace(-1., 1., nb_transition_frames)[:, np.newaxis]
for z_idx in range(nb_styles):
animate_latent_transition(latent_vectors=dlatents[0],
gen_image_fun=synth_image_fun,
gen_latent_fun=lambda z_s, i: gen_latent_style_idx(dlatents[0], i, z_idx, stylelatent_vals),
img_size=PLOT_IMG_SHAPE,
nb_frames=nb_transition_frames,
render_dir=render_dir / 'latent_indexes')
| deep learning/StyleGAN/StyleGAN - Latents Exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/livjab/DS-Unit-2-Sprint-3-Classification-Validation/blob/master/module2-baselines-validation/LS_DS_232_Baselines_Validation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="HCMrksCzGM3A" colab_type="text"
# _Lambda School Data Science — Classification & Validation_
#
# # Baselines & Validation
#
# Objectives
# - Train/Validate/Test split
# - Cross-Validation
# - Begin with baselines
# + [markdown] id="6huxBtHAGM3C" colab_type="text"
# ## Weather data — mean baseline
#
# Let's try baselines for regression.
#
# You can [get Past Weather by Zip Code from Climate.gov](https://www.climate.gov/maps-data/dataset/past-weather-zip-code-data-table). I downloaded the data for my town: Normal, Illinois.
# + id="7nDTI_FXGM3D" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 284} outputId="58e2b6a2-5e76-4cdb-ccc7-2286071316a1"
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
url = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Sprint-3-Classification-Validation/master/module2-baselines-validation/weather-normal-il.csv'
weather = pd.read_csv(url, parse_dates=['DATE']).set_index('DATE')
weather['2014-05':'2019-05'].plot(y='TMAX')
plt.title('Daily high temperature in Normal, IL');
# + [markdown] id="Czcg3EilGM3I" colab_type="text"
# Over the years, across the seasons, the average daily high temperature in my town is about 63 degrees.
# + id="wumIOBBbGM3J" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="cd28d685-df43-416f-9a90-6af94b54eb1b"
weather['TMAX'].mean()
# + [markdown] id="wfH9kFUTGM3M" colab_type="text"
# Remember from [the preread:](https://github.com/LambdaSchool/DS-Unit-2-Sprint-3-Classification-Validation/blob/master/module2-baselines-validation/model-validation-preread.md#what-does-baseline-mean) "A baseline for regression can be the mean of the training labels."
# + [markdown] id="8MSQhnFoGM3N" colab_type="text"
# If I predicted that every day, the high will be 63 degrees, I'd be off by about 19 degrees on average.
# + id="YIBMxks0GM3O" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7ba1c9d3-00b6-4b27-8088-ed3d312f63ad"
from sklearn.metrics import mean_absolute_error
predicted = [weather['TMAX'].mean()] * len(weather)
mean_absolute_error(weather['TMAX'], predicted)
# + [markdown] id="S2PC63ZKGM3R" colab_type="text"
# But, we can get a better baseline here: "A baseline for time-series regressions can be the value from the previous timestep."
#
# *Data Science for Business* explains,
#
# > Weather forecasters have two simple—but not simplistic—baseline models that they compare against. ***One (persistence) predicts that the weather tomorrow is going to be whatever it was today.*** The other (climatology) predicts whatever the average historical weather has been on this day from prior years. Each model performs considerably better than random guessing, and both are so easy to compute that they make natural baselines of comparison. Any new, more complex model must beat these.
# + [markdown] id="5i4_fxUJGM3S" colab_type="text"
# Let's predict that the weather tomorrow is going to be whatever it was today. Which is another way of saying that the weather today is going to be whatever it was yesterday.
#
# We can engineer this feature with one line of code, using the pandas [`shift`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.shift.html) function.
#
# This new baseline is off by less than 6 degress on average.
# + id="PPHul3UwGM3T" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0d3e4ea6-7a8a-4f6d-ac8f-de6671be0dfc"
weather['TMAX_yesterday'] = weather.TMAX.shift(1)
weather = weather.dropna() # Drops the first date, because it doesn't have a "yesterday"
mean_absolute_error(weather.TMAX, weather.TMAX_yesterday)
# + [markdown] id="BA2RkRY1GM3Y" colab_type="text"
# I applied this same concept for [my first submission to the Kaggle Instacart competition.](https://github.com/rrherr/springboard/blob/master/Kaggle%20Instacart%20first%20submission.ipynb)
# + [markdown] id="gWKmWS-AGM3a" colab_type="text"
# ## Bank Marketing — majority class baseline
#
# https://archive.ics.uci.edu/ml/datasets/Bank+Marketing
#
# >The data is related with direct marketing campaigns of a Portuguese banking institution. The marketing campaigns were based on phone calls. Often, more than one contact to the same client was required, in order to access if the product (bank term deposit) would be ('yes') or not ('no') subscribed.
#
# >Output variable (desired target):
# >y - has the client subscribed a term deposit? (binary: 'yes','no')
#
# >bank-additional-full.csv with all examples (41188) and 20 inputs, ordered by date (from May 2008 to November 2010)
# + [markdown] id="ES1i3sxxGM3b" colab_type="text"
# Get and read the data
# + id="NkFqWqrEGM3b" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 212} outputId="c39e1cb2-e59d-48b7-bd27-d0fde4e7ebbe"
# !wget https://archive.ics.uci.edu/ml/machine-learning-databases/00222/bank-additional.zip
# + id="bdBVnxx0GM3e" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 176} outputId="93020022-5e64-4083-ec7b-e23ab77ca625"
# !unzip bank-additional.zip
# + id="RUQXwNSCGM3i" colab_type="code" colab={}
bank = pd.read_csv('bank-additional/bank-additional-full.csv', sep=';')
# + [markdown] id="lp3DfcAqGM3l" colab_type="text"
# Assign to X and y
# + id="7PexeJi3GM3o" colab_type="code" colab={}
X = bank.drop(columns='y')
y = bank['y'] == 'yes'
# + [markdown] id="iVQ43mtTGM3r" colab_type="text"
# ## 3-way split: Train / Validation / Test
# + [markdown] id="O6svyVR_GM3s" colab_type="text"
# We know how to do a _two-way split_, with the [**`sklearn.model_selection.train_test_split`**](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html) function:
# + id="xA_9kM3aGM3t" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2, random_state=42, stratify=y)
# + [markdown] id="tRzRuqvYGM3w" colab_type="text"
# How can we get from a two-way split, to a three-way split?
#
# We can use the same function again, to split the training data into training and validation data.
# + id="XJyWQUTKGM3x" colab_type="code" colab={}
X_train, X_val, y_train, y_val = train_test_split(
X_train, y_train, test_size=0.3, random_state=42, stratify=y_train)
# + id="ySjb5S2oGM37" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d68daa24-28d4-46ce-b4a1-74497fa5b1c6"
X_train.shape, X_val.shape, X_test.shape, y_train.shape, y_val.shape, y_test.shape
# + [markdown] id="XcHmFrkeGM39" colab_type="text"
# ## Majority class baseline
# + [markdown] id="_0BpMVAsGM3-" colab_type="text"
# Determine the majority class:
# + id="YXridtnLGM3-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 70} outputId="a3091479-39a7-4ff6-cce8-fd1af4b0591f"
y_train.value_counts(normalize=True)
# + [markdown] id="R6vVd2cfGM4B" colab_type="text"
# What if we guessed the majority class for every prediction?
# + id="hoLJhQ_GGM4E" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="dfb3650a-6d57-4914-a020-17de28d89d3e"
majority_class = y_train.mode()[0]
y_pred = [majority_class] * len(y_val)
len(y_pred)
# + [markdown] id="VX7yCkhlGM4G" colab_type="text"
# #### [`sklearn.metrics.accuracy_score`](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.accuracy_score.html)
#
# Baseline accuracy by guessing the majority class for every prediction:
# + id="DH3_4YhaGM4I" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1c3912c7-8717-481a-dcea-0e451ce9619c"
from sklearn.metrics import accuracy_score
accuracy_score(y_val, y_pred)
# + [markdown] id="x6kGhw28GM4Q" colab_type="text"
# #### [`sklearn.metrics.roc_auc_score`](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.roc_auc_score.html)
#
# Baseline "ROC AUC" score by guessing the majority class for every prediction:
# + id="LuqP1dMZGM4Q" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c5cca619-e1dc-4b75-ac97-04f7552545e3"
from sklearn.metrics import roc_auc_score
roc_auc_score(y_val, y_pred)
# + [markdown] id="MhsIdQR5GM4T" colab_type="text"
# ## Fast first models
# + [markdown] id="Z57MQif-GM4U" colab_type="text"
# ### Ignore rows/columns with nulls
# + [markdown] id="g9lXM_WZGM4V" colab_type="text"
# Does this dataset have nulls?
# + id="3JRjghtXGM4Z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 390} outputId="01570be6-1cd8-4ca1-e61f-73c2dc2f1701"
X_train.isnull().sum()
# + [markdown] id="4JHNbRTPGM4c" colab_type="text"
# ### Ignore nonnumeric features
# + [markdown] id="v9R3mAqtGM4d" colab_type="text"
# Here are the numeric features:
# + id="VfvsIWzWGM4d" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 307} outputId="581cf7d7-9052-4a2b-9c50-3602337ea85c"
X_train.describe(include="number")
# + [markdown] id="4-LlZOX9GM4g" colab_type="text"
# Here are the nonnumeric features:
# + id="aWowR1o0GM4g" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 167} outputId="af277682-c73f-4861-a884-f1575be0dcc9"
X_train.describe(exclude="number")
# + [markdown] id="Z_Do2cJ5GM4i" colab_type="text"
# Just select the nonnumeric features:
# + id="UAT2xPw3GM4j" colab_type="code" colab={}
X_train_numeric = X_train.select_dtypes('number')
X_val_numeric = X_val.select_dtypes('number')
# + [markdown] id="KgQtDoeqGM4k" colab_type="text"
# ### Shallow trees are good for fast, first baselines, and to look for "leakage"
# + [markdown] id="saMVqiZ4GM4l" colab_type="text"
# #### Shallow trees
# + [markdown] id="H3KSMxR6GM4m" colab_type="text"
# After naive baselines, *Data Science for Business* suggests ["decision stumps."](https://en.wikipedia.org/wiki/Decision_stump)
#
# > A slightly more complex alternative is a model that only considers a very small amount of feature information. ...
#
# > One example is to build a "decision stump"—a decision tree with only one internal node, the root node. A tree limited to one internal node simply means that the tree induction selects the single most informative feature to make a decision. In a well-known paper in machine learning, [<NAME> (1993)](https://link.springer.com/article/10.1023/A:1022631118932) showed that ***decision stumps often produce quite good baseline performance*** ...
#
# > A decision stump is an example of the strategy of ***choosing the single most informative piece of information*** available and basing all decisions on it. In some cases most of the leverage may be coming from a single feature, and this method assesses whether and to what extent this is the case.
#
# To fit a "decision stump" we could use a [`DecisionTreeClassifier`](http://scikit-learn.org/stable/modules/generated/sklearn.tree.DecisionTreeClassifier.html) model with parameter `max_depth=1`.
#
# In this case, we'll let our tree grow a little deeper, and use the parameter `max_depth=2`
#
# In the previous code cell, we selected only the numeric features, to avoid data wrangling and save time. For now, we'll use only the numeric features.
# + [markdown] id="4r76l2YmGM4m" colab_type="text"
# #### Looking for leakage
# + [markdown] id="3YyAqMHQGM4n" colab_type="text"
# [<NAME> recommends,](https://www.quora.com/What-are-some-best-practices-for-training-machine-learning-models/answer/Xavier-Amatriain)
#
# "Make sure your training features do not contain data from the “future” (aka time traveling). While this might be easy and obvious in some cases, it can get tricky. ... If your test metric becomes really good all of the sudden, ask yourself what you might be doing wrong. Chances are you are time travelling or overfitting in some way."
# + [markdown] id="QJeSg13-GM4o" colab_type="text"
# We can test this with the [UCI repository's Bank Marketing dataset](https://archive.ics.uci.edu/ml/datasets/Bank+Marketing). It has a feature which leaks information from the future and should be dropped:
#
# >11 - duration: last contact duration, in seconds (numeric). Important note: this attribute highly affects the output target (e.g., if duration=0 then y='no'). Yet, the duration is not known before a call is performed. Also, after the end of the call y is obviously known. Thus, this input ... should be discarded if the intention is to have a realistic predictive model.
# + [markdown] id="BB-9RVozGM4r" colab_type="text"
# #### Let's train a shallow tree basline
#
# ... without dropping the leaky `duration` feature.
# + id="5Q6Q5N0tGM4s" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="85e546a4-4340-4ad5-9913-64b3df96a71d"
from sklearn.tree import DecisionTreeClassifier
tree = DecisionTreeClassifier(max_depth=2)
tree.fit(X_train_numeric, y_train)
y_pred_proba = tree.predict_proba(X_val_numeric)[:,1]
roc_auc_score(y_val, y_pred_proba)
# + [markdown] id="IDO85GjfGM40" colab_type="text"
# Then we can visualize the tree to see which feature(s) were the "most informative":
# + id="S3Hd9Yl7GM41" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 378} outputId="35086f93-b0b7-4a3b-8d91-7abdbcc30287"
import graphviz
from sklearn.tree import export_graphviz
dot_data = export_graphviz(tree, out_file=None, feature_names=X_train_numeric.columns,
class_names=['No', 'Yes'], filled=True, impurity=False, proportion=True)
graphviz.Source(dot_data)
# + [markdown] id="xN4fP9npGM43" colab_type="text"
# This baseline has a ROC AUC score above 0.85, and it uses the `duration` feature, as well as `nr.employed`, a "social and economic context attribute" for "number of employees - quarterly indicator."
# + [markdown] id="eGQMRJN3GM44" colab_type="text"
# #### Let's drop the `duration` feature
# + id="gqVLIZu5GM45" colab_type="code" colab={}
X_train = X_train.drop(columns='duration')
X_val = X_val.drop(columns='duration')
X_test = X_test.drop(columns='duration')
# + id="h8yotNWiAu7W" colab_type="code" colab={}
X_train_numeric = X_train.select_dtypes('number')
X_val_numeric = X_val.select_dtypes('number')
# + [markdown] id="qdrygI0WGM47" colab_type="text"
# When the `duration` feature is dropped, then the ROC AUC score drops. Which is what we expect, it's not a bad thing in this situation!
# + id="GsGYtpIbGM48" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9c3b8e75-c1f7-4d2a-fee5-9b2579fa11ba"
tree = DecisionTreeClassifier(max_depth=2)
tree.fit(X_train_numeric, y_train)
y_pred_proba = tree.predict_proba(X_val_numeric)[:,1]
roc_auc_score(y_val, y_pred_proba)
# + id="5-_fZYrqGM4_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 378} outputId="a91efc11-4a49-4801-e34b-55b10b0586bb"
dot_data = export_graphviz(tree, out_file=None, feature_names=X_train_numeric.columns,
class_names=['No', 'Yes'], filled=True, impurity=False, proportion=True)
graphviz.Source(dot_data)
# + [markdown] id="t8OWKwPEGM5F" colab_type="text"
# ### Logistic Regression
#
# Logistic Regression is another great option for fast, first baselines!
# + id="4-Ko1Bu0GM5G" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b75c2ffe-f8a2-42f0-be3f-fbe5cca14918"
from sklearn.linear_model import LogisticRegression
model = LogisticRegression(solver="lbfgs", max_iter=1000)
model.fit(X_train_numeric, y_train)
model.score(X_val_numeric, y_val)
# + id="uiFPAiKtB0uu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e6fb2383-333e-4510-d0d2-9c7b29052a90"
y_pred_proba = model.predict_proba(X_val_numeric)[:,1]
roc_auc_score(y_val, y_pred_proba)
# + [markdown] id="rm7HaJyyGM5I" colab_type="text"
# ### With Scaler
# https://scikit-learn.org/stable/modules/preprocessing.html
# + id="SR_WV3o2GM5I" colab_type="code" colab={}
import warnings
from sklearn.exceptions import DataConversionWarning
warnings.filterwarnings(action='ignore', category=DataConversionWarning)
# + id="naRpWyKOGM5L" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d8cc708e-e6bb-4fa0-91d6-ead306521678"
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train_numeric)
X_val_scaled = scaler.transform(X_val_numeric)
model = LogisticRegression(solver='lbfgs', max_iter=1000)
model.fit(X_train_scaled, y_train)
y_pred_proba = model.predict_proba(X_val_scaled)[:,1]
roc_auc_score(y_val, y_pred_proba)
# + [markdown] id="4XZJXwO1GM5O" colab_type="text"
# ### Same, as a pipeline
# + id="o9nMEgrjGM5O" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="bd0ba394-31e6-4609-a71f-1fb8cc5e8019"
from sklearn.pipeline import make_pipeline
pipeline = make_pipeline(StandardScaler(), LogisticRegression(solver="lbfgs", max_iter=1000))
pipeline.fit(X_train_scaled, y_train)
y_pred_proba = pipeline.predict_proba(X_val_scaled)[:,1]
roc_auc_score(y_val, y_pred_proba)
# + [markdown] id="fouCvV8SGM5Q" colab_type="text"
# ### Encode "low cardinality" categoricals
# + [markdown] id="xuFKp5y9GM5R" colab_type="text"
# [Cardinality](https://simple.wikipedia.org/wiki/Cardinality) means the number of unique values that a feature has:
# > In mathematics, the cardinality of a set means the number of its elements. For example, the set A = {2, 4, 6} contains 3 elements, and therefore A has a cardinality of 3.
#
# One-hot encoding adds a dimension for each unique value of each categorical feature. So, it may not be a good choice for "high cardinality" categoricals that have dozens, hundreds, or thousands of unique values.
#
# In this dataset, all the categoricals seem to be "low cardinality", so we can use one-hot encoding.
# + id="t71R5HsnGM5T" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 212} outputId="6e155359-fe04-4f90-bc41-5b0852d80cad"
X_train.select_dtypes(exclude="number").nunique()
# + [markdown] id="MmFH52rRGM5e" colab_type="text"
# #### Install the [Category Encoders](https://github.com/scikit-learn-contrib/categorical-encoding) library
#
# If you're running on Google Colab:
#
# ```
# # # !pip install category_encoders
# ```
#
# If you're running locally with Anaconda:
#
# ```
# # # !conda install -c conda-forge category_encoders
# ```
# + id="eQf0JF9XGM5f" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 214} outputId="e1c3fdea-4511-4296-b8bd-3c9d54671903"
# !pip install category_encoders
# + id="5QyOoQcTHI3m" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="397d88dc-9e29-42c5-abad-0c8c84fc0cc8"
import category_encoders as ce
pipeline = make_pipeline(ce.OneHotEncoder(use_cat_names=True),
StandardScaler(),
LogisticRegression(solver="lbfgs", max_iter=1000))
pipeline.fit(X_train, y_train)
y_pred_proba = pipeline.predict_proba(X_val)[:,1]
roc_auc_score(y_val, y_pred_proba)
# + id="sreMeIsBILrm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d2e17075-0b8a-4c92-8306-c1be1f3e5051"
pipeline = make_pipeline(ce.OneHotEncoder(use_cat_names=True),
StandardScaler(),
DecisionTreeClassifier(max_depth=5))
pipeline.fit(X_train, y_train)
y_pred_proba = pipeline.predict_proba(X_val)[:,1]
roc_auc_score(y_val, y_pred_proba)
# + [markdown] id="fxp4HJKtGM5i" colab_type="text"
# # Baseline with cross-validation + independent test set
# A complete example, as an alternative to Train/Validate/Test
#
#
# #### scikit-learn documentation
# - [`sklearn.model_selection.cross_val_score`](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.cross_val_score.html)
# - [ The `scoring` parameter: defining model evaluation rules](https://scikit-learn.org/stable/modules/model_evaluation.html#the-scoring-parameter-defining-model-evaluation-rules)
# + id="Pt6ksKU_GM5n" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 87} outputId="252152d8-2aef-4801-a0af-bc719ab6e72d"
# Imports
# %matplotlib inline
import warnings
import category_encoders as ce
import matplotlib.pyplot as plt
import pandas as pd
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import cross_val_score
from sklearn.pipeline import make_pipeline
from sklearn.exceptions import DataConversionWarning
from sklearn.preprocessing import StandardScaler
warnings.filterwarnings(action='ignore', category=DataConversionWarning)
# Load data
bank = pd.read_csv('bank-additional/bank-additional-full.csv', sep=';')
# Assign to X, y
X = bank.drop(columns='y')
y = bank['y'] == 'yes'
# Drop leaky & random features
X = X.drop(columns='duration')
# Split Train, Test
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2, random_state=42, stratify=y)
# Make pipeline
pipeline = make_pipeline(
ce.OneHotEncoder(use_cat_names=True),
StandardScaler(),
LogisticRegression(solver='lbfgs', max_iter=1000)
)
# Cross-validate with training data
scores = cross_val_score(pipeline, X_train, y_train, scoring='roc_auc', cv=10, n_jobs=-1, verbose=10)
# + [markdown] id="EW9dhEtJGM5t" colab_type="text"
# This is the baseline score that more sophisticated models must beat.
# + id="09RaXaPLGM5u" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 70} outputId="9a8f0926-0597-4094-9e2e-386f14ddbf66"
print('Cross-Validation ROC AUC scores:', scores)
print('Average:', scores.mean())
# + [markdown] id="G_3dJBYgGM5x" colab_type="text"
# Is more effort justified? It depends. The blogpost ["Always start with a stupid model"](https://blog.insightdatascience.com/always-start-with-a-stupid-model-no-exceptions-3a22314b9aaa) explains,
#
# > Here is a very common story: a team wants to implement a model to predict something like the probability of a user clicking an ad. They start with a logistic regression and quickly (after some minor tuning) reach 90% accuracy.
#
# > From there, the question is: Should the team focus on getting the accuracy up to 95%, or should they solve other problems 90% of the way?
#
# > ***If a baseline does well, then you’ve saved yourself the headache of setting up a more complex model. If it does poorly, the kind of mistakes it makes are very instructive*** ...
#
# So what else can we learn from this baseline?
#
# ["Always start with a stupid model"](https://blog.insightdatascience.com/always-start-with-a-stupid-model-no-exceptions-3a22314b9aaa) suggests to look at
#
# > **What type of signal your model picks up on.** Most baselines will allow you to extract ***feature importances***, revealing which aspects of the input are most predictive. Analyzing feature importance is a great way to realize how your model is making decisions, and what it might be missing.
#
# We can do that:
# + id="m3mdIpO2GM5y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1694} outputId="5e1ce69a-fd81-41f2-b0d7-b0daab932d39"
# (Re)fit on training data
pipeline.fit(X_train, y_train)
# Visualize coefficients
plt.figure(figsize=(10,30))
plt.title('Coefficients')
coefficients = pipeline.named_steps['logisticregression'].coef_[0]
feature_names = pipeline.named_steps['onehotencoder'].transform(X_train).columns
pd.Series(coefficients, feature_names).sort_values().plot.barh(color='gray');
# + [markdown] id="aPz9DCuiGM50" colab_type="text"
# [The post](https://blog.insightdatascience.com/always-start-with-a-stupid-model-no-exceptions-3a22314b9aaa) also recommends we consider,
#
# > **What signal your model is missing.** If there is a certain aspect of the data that seems intuitively important but that your model is ignoring, ***a good next step is to engineer a feature*** or pick a different model that could better leverage this particular aspect of your data.
# + [markdown] id="v6E27-wBGM52" colab_type="text"
# ### Look at your data (you still need to do it!)
#
# Cautionary tales
# - [Exploring the ChestXray14 dataset: problems](https://lukeoakdenrayner.wordpress.com/2017/12/18/the-chestxray14-dataset-problems/)
# - [The Quartz guide to bad data](https://github.com/Quartz/bad-data-guide)
#
# Incomplete list of issues to address
# - Categoricals (text, dates/times, high cardinality)
# - Feature Engineering (extraction, interaction, transformations)
# - Missing Values
# - Outliers
# + [markdown] id="m5cI9rAaGM52" colab_type="text"
# # ASSIGNMENT options
#
# - **Replicate the lesson code.** [Do it "the hard way" or with the "Benjamin Franklin method."](https://docs.google.com/document/d/1ubOw9B3Hfip27hF2ZFnW3a3z9xAgrUDRReOEo-FHCVs/edit)
# - Apply the lesson to other datasets you've worked with before, and compare results.
# - Iterate and improve your **Bank Marketing** model. Engineer new features.
# - Get **weather** data for your own area and calculate both baselines. _"One (persistence) predicts that the weather tomorrow is going to be whatever it was today. The other (climatology) predicts whatever the average historical weather has been on this day from prior years."_ What is the mean absolute error for each baseline? What if you average the two together?
# - [This example from scikit-learn documentation](https://scikit-learn.org/stable/auto_examples/compose/plot_column_transformer_mixed_types.html) demonstrates its improved `OneHotEncoder` and new `ColumnTransformer` objects, which can replace functionality from [third-party libraries](https://github.com/scikit-learn-contrib) like category_encoders and sklearn-pandas. Adapt this example, which uses Titanic data, to work with Bank Marketing or another dataset.
# - When would this notebook's pipelines fail? How could you fix them? Add more [preprocessing](https://scikit-learn.org/stable/modules/preprocessing.html) and [imputation](https://scikit-learn.org/stable/modules/impute.html) to your [pipelines](https://scikit-learn.org/stable/modules/compose.html) with scikit-learn.
| module2-baselines-validation/LS_DS_232_Baselines_Validation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # First Neurons
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
# ** Set Random Seeds for results reproducibility **
np.random.seed(101)
tf.set_random_seed(101)
# ** Data Setup **
# Setting Up some Random Data for Demonstration Purposes
rand_a = np.random.uniform(low = 0,
high = 100,
size = (5,5))
rand_a
rand_b = np.random.uniform(low = 0,
high = 100,
size = (5,1))
rand_b
# ### Placeholders
# Initialize placeholders as of type float32
a = tf.placeholder(tf.float32)
b = tf.placeholder(tf.float32)
# ### Operations
add_op = a + b # tf.add(a,b)
mult_op = a * b #tf.multiply(a,b)
# ### Running Sessions to create Graphs with Feed Dictionaries
with tf.Session() as sess:
add_result = sess.run(add_op,
feed_dict={a : rand_a,
b : rand_b})
print(add_result)
print('\n')
mult_result = sess.run(mult_op,
feed_dict={a : rand_a,
b : rand_b})
print(mult_result)
# ________________________
#
# ________________________
# ## Example Neural Network
n_features = 10
n_dense_neurons = 3
# Placeholder for x
x = tf.placeholder(tf.float32, (None, n_features))
# +
# Variables for w and b
b = tf.Variable(tf.zeros([n_dense_neurons]))
W = tf.Variable(tf.random_normal([n_features, n_dense_neurons]))
# -
b.get_shape()
W.get_shape()
# ** Operation Activation Function **
xW = tf.matmul(x, W)
z = tf.add(xW,b)
# tf.nn.relu() or tf.tanh()
a = tf.sigmoid(z)
# ** Variable Intializer! **
init = tf.global_variables_initializer()
with tf.Session() as sess:
# Run session with the initializer
sess.run(init)
# Result
layer_out = sess.run(a,feed_dict={x : np.random.random([1, n_features])})
print(layer_out)
# We still need to finish off this process with optimization! Let's learn how to do this next.
#
# _____
# ## Full Network Example
#
# Let's work on a regression example, we are trying to solve a very simple equation:
#
# y = mx + b
#
# y will be the y_labels and x is the x_data. We are trying to figure out the slope and the intercept for the line that best fits our data!
# ### Artifical Data (Some Made Up Regression Data)
x_data = np.linspace(0, 10, 10) + np.random.uniform(-1.5, 1.5, 10)
x_data
y_label = np.linspace(0, 10, 10) + np.random.uniform(-1.5, 1.5, 10)
# %matplotlib inline
plt.figure(figsize = (10, 10))
plt.plot(x_data,y_label,'*')
# ** Variables **
np.random.rand(2)
m = tf.Variable(0.442)
b = tf.Variable(0.877)
# ### Cost Function
error = 0
for x, y in zip(x_data, y_label):
# Predicted value
y_hat = m * x + b
# Mean Squared Error (MSE)
error += (y - y_hat) ** 2
# ### Optimizer
optimizer = tf.train.GradientDescentOptimizer(learning_rate = 0.001)
train = optimizer.minimize(error)
# ### Initialize Variables
init = tf.global_variables_initializer()
# ### Create Session and Run!
epochs = 1000
with tf.Session() as sess:
sess.run(init)
for i in range(epochs):
sess.run(train)
# Print the error for every 50th epoch.
if (i % 50 == 0):
print("Epoch {}: {}".format(i, sess.run(error)))
# Get the m and b values.
final_slope , final_intercept = sess.run([m, b])
final_slope
final_intercept
# ### Evaluate Results
# +
x_test = np.linspace(-1, 11, 10)
y_pred_plot = final_slope * x_test + final_intercept
plt.plot(x_test,y_pred_plot,'r')
plt.plot(x_data,y_label,'*')
# -
# # Great Job!
| 18-05-28-Complete-Guide-to-Tensorflow-for-Deep-Learning-with-Python/02-Tensorflow-Basics/03-TF-Neural-Network.ipynb |