code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
# <script>
# window.dataLayer = window.dataLayer || [];
# function gtag(){dataLayer.push(arguments);}
# gtag('js', new Date());
#
# gtag('config', 'UA-59152712-8');
# </script>
#
# # Tutorial: Gaussian pulse initial data for a massless scalar field in spherical-like coordinates
#
# ## Authors: <NAME> and <NAME>
#
# # This tutorial notebook explains how to obtain time-symmetric initial data for the problem of gravitational collapse of a massless scalar field. We will be following the approaches of [Akbarian & Choptuik (2015)](https://arxiv.org/pdf/1508.01614.pdf) and [Baumgarte (2018)](https://arxiv.org/pdf/1807.10342.pdf).
#
# **Notebook Status**: <font color='green'><b> Validated </b></font>
#
# **Validation Notes**: The initial data generated by the NRPy+ module corresponding to this tutorial notebook are used shown to satisfy Einstein's equations as expected [in this tutorial notebook](Tutorial-Start_to_Finish-BSSNCurvilinear-Setting_up_ScalarField_initial_data.ipynb).</font>
#
# ## Python module which performs the procedure described in this tutorial: [ScalarField/ScalarField_InitialData.py](../edit/ScalarField/ScalarField_InitialData.py)
#
# ## References
#
# * [Akbarian & Choptuik (2015)](https://arxiv.org/pdf/1508.01614.pdf) (Useful to understand the theoretical framework)
# * [Baumgarte (2018)](https://arxiv.org/pdf/1807.10342.pdf) (Useful to understand the theoretical framework)
# * [Baumgarte & Shapiro's Numerical Relativity](https://books.google.com.br/books/about/Numerical_Relativity.html?id=dxU1OEinvRUC&redir_esc=y): Section 6.2.2 (Useful to understand how to solve the Hamiltonian constraint)
#
# <a id='toc'></a>
#
# # Table of Contents
# $$\label{toc}$$
#
# 1. [Step 1](#initial_data) Setting up time-symmetric initial data
# 1. [Step 1.a](#id_time_symmetry) Time symmetry: $\tilde{K}_{ij}$, $\tilde K$, $\tilde\beta^{i}$, and $\tilde B^{i}$
# 1. [Step 1.b](#id_sf_ic) The scalar field initial condition: $\tilde{\varphi}$, $\tilde{\Phi}$, $\tilde{\Pi}$
# 1. [Step 1.c](#id_metric) The physical metric: $\tilde{\gamma}_{ij}$
# 1. [Step 1.c.i](#id_conformal_metric) The conformal metric $\bar\gamma_{ij}$
# 1. [Step 1.c.ii](#id_hamiltonian_constraint) Solving the Hamiltonian constraint
# 1. [Step 1.c.ii.1](#id_tridiagonal_matrix) The tridiagonal matrix: $A$
# 1. [Step 1.c.ii.2](#id_tridiagonal_rhs) The right-hand side of the linear system: $\vec{s}$
# 1. [Step 1.c.ii.3](#id_conformal_factor) The conformal factor: $\psi$
# 1. [Step 1.d](#id_lapse_function) The lapse function: $\tilde{\alpha}$
# 1. [Step 1.e](#id_output) Outputting the initial data to file
# 1. [Step 2](#id_interpolation_files) Interpolating the initial data file as needed
# 1. [Step 3](#id_sph_to_curvilinear) Converting Spherical initial data to Curvilinear initial data
# 1. [Step 4](#validation) Validation of this tutorial against the [ScalarField/ScalarField_InitialData.py](../edit/ScalarField/ScalarField_InitialData.py) module
# 1. [Step 5](#output_to_pdf) Output this module as $\LaTeX$-formatted PDF file
# <a id='initialize_nrpy'></a>
#
# # Step 0: Initialize Python/NRPy+ modules \[Back to [top](#toc)\]
# $$\label{initialize_nrpy}$$
# +
# Step 0: Load all needed Python/NRPy+ modules
import os,sys,shutil # Standard Python modules for multiplatform OS-level functions
import sympy as sp # SymPy: The Python computer algebra package upon which NRPy+ depends
import numpy as np # NumPy: A large collection of mathematical functions for Python
from scipy.sparse import spdiags # SciPy: Sparse, tri-diagonal matrix setup function
from scipy.sparse import csc_matrix # SciPy: Sparse matrix optimization function
from scipy.sparse.linalg import spsolve # SciPy: Solver of linear systems involving sparse matrices
import outputC as outC # NRPy+: Core C code output module
import reference_metric as rfm # NRPy+: Reference metric support
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
# Step 0.a: Create the output directory
Ccodesdir = "ScalarFieldID_validation"
shutil.rmtree(Ccodesdir,ignore_errors=True)
cmd.mkdir(Ccodesdir)
# -
# <a id='initial_data'></a>
#
# # Step 1: Setting up time-symmetric initial data \[Back to [top](#toc)\]
# $$\label{initial_data}$$
#
# In this section we will set up time symmetric initial data for the gravitational collapse of a massless scalar field, in spherical coordinates. Our discussion will follow closely section III.A of [Akbarian & Choptuik (2015)](https://arxiv.org/pdf/1508.01614.pdf) (henceforth A&C). We will be using a *uniform* radial sampling. All initial data quantities will be written with tildes over them, meaning that, for example, $\tilde{\alpha} \equiv \alpha(0,r)$.
#
# <a id='id_time_symmetry'></a>
#
# ## Step 1.a: Time symmetry: $\tilde{K}_{ij}$, $\tilde K$, $\tilde\beta^{i}$, and $\tilde B^{i}$ \[Back to [top](#toc)\]
# $$\label{id_time_symmetry}$$
#
# We are here considering a spherically symmetric problem, so that $f=f(t,r)$, for every function discussed in this tutorial. The demand for time-symmetric initial data then imples that
#
# \begin{align}
# \tilde K_{ij} &= 0\ ,\\
# \tilde K &= 0\ ,\\
# \tilde \beta^{i} &= 0\ ,\\
# \tilde B^{i} &= 0\ .
# \end{align}
#
# For the scalar field, $\varphi$, it also demands
#
# $$
# \partial_{t}\varphi(0,r) = 0\ ,
# $$
#
# which we discuss below.
#
# <a id='id_sf_ic'></a>
#
# ## Step 1.b: The scalar field initial condition: $\tilde{\varphi}$, $\tilde{\Phi}$, $\tilde{\Pi}$ \[Back to [top](#toc)\]
# $$\label{id_sf_ic}$$
#
# We will be implementing the following options for the initial profile of the scalar field
#
# $$
# \begin{aligned}
# \tilde{\varphi}_{\rm I} &= \varphi_{0}\exp\left(-\frac{r^{2}}{\sigma^{2}}\right)\ ,\\
# \tilde{\varphi}_{\rm II} &= \varphi_{0}r^{3}\exp\left[-\left(\frac{r-r_{0}}{\sigma}\right)^{2}\right]\ ,\\
# \tilde{\varphi}_{\rm III} &= \varphi_{0}\left\{1 - \tanh\left[\left(\frac{r-r_{0}}{\sigma}\right)^{2}\right]\right\}.
# \end{aligned}
# $$
#
# We introduce the two auxiliary fields
#
# $$
# \tilde\Phi\equiv\partial_{r}\tilde\varphi\quad \text{and}\quad \Pi\equiv-\frac{1}{\alpha}\left(\partial_{t}\varphi - \beta^{i}\partial_{i}\varphi\right)\ ,
# $$
#
# of which $\tilde\Phi$ will only be used as an auxiliary variable for setting the initial data, but $\Pi$ is a dynamical variable which will be evolved in time. Because we are setting time-symmetric initial data, $\partial_{t}\sf = 0 = \beta^{i}$, and thus $\tilde\Pi=0$.
# +
# Step 1: Setting up time-symmetric initial data
# Step 1.a: Define basic parameters
# Step 1.a.i: Domain size
RMAX = 50
# Step 1.a.ii: Number of gridpoints in the radial direction
NR = 30000
# Step 1.a.iii: Initial data family. Available options are:
# Gaussian_pulse, Gaussian_pulsev2, and Tanh_pulse
ID_Family = "Gaussian_pulsev2"
# Step 1.a.iv: Coordinate system. Available options are:
# Spherical and SinhSpherical
CoordSystem = "Spherical"
# Step 1.a.v: SinhSpherical parameters
sinhA = RMAX
sinhW = 0.1
# Step 1.b: Set the radial array
if CoordSystem == "Spherical":
r = np.linspace(0,RMAX,NR+1) # Set the r array
dr = np.zeros(NR)
for i in range(NR):
dr[i] = r[1]-r[0]
r = np.delete(r-dr[0]/2,0) # Shift the vector by -dr/2 and remove the negative entry
elif CoordSystem == "SinhSpherical":
if sinhA is None or sinhW is None:
print("Error: SinhSpherical coordinates require initialization of both sinhA and sinhW")
sys.exit(1)
else:
x = np.linspace(0,1.0,NR+1)
dx = 1.0/(NR+1)
x = np.delete(x-dx/2,0) # Shift the vector by -dx/2 and remove the negative entry
r = sinhA * np.sinh( x/sinhW ) / np.sinh( 1.0/sinhW )
dr = sinhA * np.cosh( x/sinhW ) / np.sinh( 1.0/sinhW ) * dx
else:
print("Error: Unknown coordinate system")
sys.exit(1)
# Step 1.c: Step size squared
dr2 = dr**2
# Step 1.d: Set SymPy variables for the initial condition
phi0,rr,rr0,sigma = sp.symbols("phi0 rr rr0 sigma",real=True)
# Step 1.e: Now set the initial profile of the scalar field
if ID_Family == "Gaussian_pulse":
phiID = phi0 * sp.exp( -r**2/sigma**2 )
elif ID_Family == "Gaussian_pulsev2":
phiID = phi0 * rr**3 * sp.exp( -(rr-rr0)**2/sigma**2 )
elif ID_Family == "Tanh_pulse":
phiID = phi0 * ( 1 - sp.tanh( (rr-rr0)**2/sigma**2 ) )
else:
print("Unkown initial data family: ",ID_Family)
print("Available options are: Gaussian_pulse, Gaussian_pulsev2, and Tanh_pulse")
sys.exit(1)
# Step 1.f: Compute Phi := \partial_{r}phi
PhiID = sp.diff(phiID,rr)
# Step 1.g: Generate NumPy functions for phi
# and Phi from the SymPy variables.
phi = sp.lambdify((phi0,rr,rr0,sigma),phiID)
Phi = sp.lambdify((phi0,rr,rr0,sigma),PhiID)
# Step 1.h: populating the varphi(0,r) array
phi0 = 0.1
r0 = 0
sigma = 1
ID_sf = phi(phi0,r,r0,sigma)
# -
# <a id='id_metric'></a>
#
# ## Step 1.c: The physical metric: $\tilde{\gamma}_{ij}$ \[Back to [top](#toc)\]
# $$\label{id_metric}$$
#
# <a id='id_conformal_metric'></a>
#
# ### Step 1.c.i: The conformal metric $\bar\gamma_{ij}$ \[Back to [top](#toc)\]
# $$\label{id_conformal_metric}$$
#
# To set up the physical metric initial data, $\tilde\gamma_{ij}$, we will start by considering the conformal transformation
#
# $$
# \gamma_{ij} = e^{4\phi}\bar\gamma_{ij}\ ,
# $$
#
# where $\bar\gamma_{ij}$ is the conformal metric and $e^{\phi}$ is the conformal factor. We then fix the initial value of $\bar\gamma_{ij}$ according to eqs. (32) and (43) of [A&C](https://arxiv.org/pdf/1508.01614.pdf)
#
# $$
# \bar\gamma_{ij} = \hat\gamma_{ij}\ ,
# $$
#
# where $\hat\gamma_{ij}$ is the *reference metric*, which is the flat metric in spherical symmetry
#
# $$
# \hat\gamma_{ij}
# =
# \begin{pmatrix}
# 1 & 0 & 0\\
# 0 & r^{2} & 0\\
# 0 & 0 & r^{2}\sin^{2}\theta
# \end{pmatrix}\ .
# $$
#
# To determine the physical metric, we must then determine the conformal factor $e^{\phi}$. This is done by solving the Hamiltonian constraint (cf. eq. (12) of [Baumgarte](https://arxiv.org/pdf/1807.10342.pdf))
#
# $$
# \hat\gamma^{ij}\hat D_{i}\hat D_{j}\psi = -2\pi\psi^{5}\rho\ ,
# $$
#
# where $\psi\equiv e^{\tilde\phi}$. For a massless scalar field, we know that
#
# $$
# T^{\mu\nu} = \partial^{\mu}\varphi\partial^{\nu}\varphi - \frac{1}{2}g^{\mu\nu}\left(\partial^{\lambda}\varphi\partial_{\lambda}\varphi\right)\ .
# $$
#
# where $g^{\mu\nu}$ is the inverse of the ADM 4-metric given by eq. (2.119) of [Baumgarte & Shapiro's Numerical Relativity](https://books.google.com.br/books/about/Numerical_Relativity.html?id=dxU1OEinvRUC&redir_esc=y),
#
# $$
# g^{\mu\nu}=\begin{pmatrix}
# -\alpha^{-2} & \alpha^{-2}\beta^{i}\\
# \alpha^{-2}\beta^{j} & \gamma^{ij} - \alpha^{-2}\beta^{i}\beta^{j}
# \end{pmatrix}\ .
# $$
#
# We know that (see Step 2 in [this tutorial module](Tutorial-ADM_Setting_up_massless_scalarfield_Tmunu.ipynb) for the details)
#
# \begin{align}
# \partial^{t}\varphi &= \alpha^{-1}\Pi\ ,\\
# \partial^{\lambda}\varphi\partial_{\lambda}\varphi &= -\Pi^{2} + \gamma^{ij}\partial_{i}\varphi\partial_{j}\varphi\ .
# \end{align}
#
# The tt-component of the energy-momentum tensor at the initial time is then given by (we will ommit the "tildes" below to avoid cluttering the equation, but keep in mind that all quantities are considered at $t=0$)
#
# \begin{align}
# T^{tt} &= \left(\partial^{t}\varphi\right)^{2} - \frac{1}{2} g^{tt}\left(\partial^{\lambda}\varphi\partial_{\lambda}\varphi\right)\nonumber\\
# &= \left(\frac{\Pi}{\alpha}\right)^{2} - \frac{1}{2}\left(-\frac{1}{\alpha^{2}}\right)\left(-\Pi^{2} + \gamma^{ij}\partial_{i}\varphi\partial_{j}\varphi\right)\nonumber\\
# &= \frac{\gamma^{ij}\partial_{i}\varphi\partial_{j}\varphi}{2\alpha^{2}}\nonumber\\
# &= \frac{e^{-4\phi}\bar\gamma^{ij}\partial_{i}\varphi\partial_{j}\varphi}{2\alpha^{2}}\nonumber\\
# &= \frac{e^{-4\phi}\hat\gamma^{ij}\partial_{i}\varphi\partial_{j}\varphi}{2\alpha^{2}}\nonumber\\
# &= \frac{e^{-4\phi}\hat\gamma^{rr}\partial_{r}\varphi\partial_{r}\varphi}{2\alpha^{2}}\nonumber\\
# &= \frac{e^{-4\phi}\Phi^{2}}{2\alpha^{2}}\nonumber\\
# \end{align}
#
# By remembering the definition of the normal vector $n_{\mu} = (-\alpha,0,0,0)$ (eq. (2.117) of [B&S](https://books.google.com.br/books/about/Numerical_Relativity.html?id=dxU1OEinvRUC&redir_esc=y)), we can then evaluate the energy density $\rho$ given by eq. (24) of [A&C](https://arxiv.org/pdf/1508.01614.pdf)
#
# $$
# \tilde\rho = \tilde n_{\mu}\tilde n_{\nu}\tilde T^{\mu\nu} = \frac{e^{-4\tilde\phi}}{2}\tilde\Phi^{2}\ .
# $$
#
# Plugging this result in the Hamiltonian constraint, remembering that $\psi\equiv e^{\tilde\phi}$, we have
#
# $$
# \partial^{2}_{r}\psi + \frac{2}{r}\partial_{r}\psi + \pi\psi\Phi^{2} = 0\ .
# $$
#
# This is a linear elliptic equation which will solve using the procedure described in detail in section 6.2.2 of [B&S](https://books.google.com.br/books/about/Numerical_Relativity.html?id=dxU1OEinvRUC&redir_esc=y).
#
# <a id='id_hamiltonian_constraint'></a>
#
# ### Step 1.c.ii: Solving the Hamiltonian constraint \[Back to [top](#toc)\]
# $$\label{id_hamiltonian_constraint}$$
#
# We will discretize the Hamiltonian constraint using [second-order accurate finite differences](https://en.wikipedia.org/wiki/Finite_difference_coefficient). We get
#
# $$
# \frac{\psi_{i+1} - 2\psi_{i} + \psi_{i-1}}{\Delta r^{2}} + \frac{2}{r_{i}}\left(\frac{\psi_{i+1}-\psi_{i-1}}{2\Delta r}\right) + \pi\psi_{i}\Phi^{2}_{i} = 0\ ,
# $$
#
# or, by multiplying the entire equation by $\Delta r^{2}$ and then grouping the coefficients of each $\psi_{j}$:
#
# $$
# \boxed{\left(1-\frac{\Delta r}{r_{i}}\right)\psi_{i-1}+\left(\pi\Delta r^{2}\Phi_{i}^{2}-2\right)\psi_{i} + \left(1+\frac{\Delta r}{r_{i}}\right)\psi_{i+1} = 0}\ .
# $$
#
# We choose to set up a grid that is cell-centered, with:
#
# $$
# r_{i} = \left(i-\frac{1}{2}\right)\Delta r\ ,
# $$
#
# so that $r_{0} = - \frac{\Delta r}{2}$. This is a two-point boundary value problem, which we solve using the same strategy as [A&C](https://arxiv.org/pdf/1508.01614.pdf), described in eqs. (48)-(50):
#
# \begin{align}
# \left.\partial_{r}\psi\right|_{r=0} &= 0\ ,\\
# \lim_{r\to\infty}\psi &= 1\ .
# \end{align}
#
# In terms of our grid structure, the first boundary condition (regularity at the origin) is written to second-order in $\Delta r$ as:
#
# $$
# \left.\partial_{r}\psi\right|_{r=0} = \frac{\psi_{1} - \psi_{0}}{\Delta r} = 0 \Rightarrow \psi_{0} = \psi_{1}\ .
# $$
#
# The second boundary condition (asymptotic flatness) can be interpreted as
#
# $$
# \psi_{N} = 1 + \frac{C}{r_{N}}\ (r_{N}\gg1)\ ,
# $$
#
# which then implies
#
# $$
# \partial_{r}\psi_{N} = -\frac{C}{r_{N}^{2}} = -\frac{1}{r_{N}}\left(\frac{C}{r_{N}}\right) = -\frac{1}{r_{N}}\left(\psi_{N} - 1\right) = \frac{1-\psi_{N}}{r_{N}}\ ,
# $$
#
# which can then be written as
#
# $$
# \frac{\psi_{N+1}-\psi_{N-1}}{2\Delta r} = \frac{1-\psi_{N}}{r_{N}}\Rightarrow \psi_{N+1} = \psi_{N-1} - \frac{2\Delta r}{r_{N}}\psi_{N} + \frac{2\Delta r}{r_{N}}\ .
# $$
#
# Substituting the boundary conditions at the boxed equations above, we end up with
#
# \begin{align}
# \left(\pi\Delta r^{2}\Phi^{2}_{1} - 1 - \frac{\Delta r}{r_{1}}\right)\psi_{1} + \left(1+\frac{\Delta r}{r_{1}}\right)\psi_{2} = 0\quad &(i=1)\ ,\\
# \left(1-\frac{\Delta r}{r_{i}}\right)\psi_{i-1}+\left(\pi\Delta r^{2}\Phi_{i}^{2}-2\right)\psi_{i} + \left(1+\frac{\Delta r}{r_{i}}\right)\psi_{i+1} = 0\quad &(1<i<N)\ ,\\
# 2\psi_{N-1} + \left[\pi\Delta r^{2}\Phi^{2}_{N} - 2 - \frac{2\Delta r}{r_{N}}\left(1+\frac{\Delta r}{r_{N}}\right)\right]\psi_{N} = - \frac{2\Delta r}{r_{N}}\left(1+\frac{\Delta r}{r_{N}}\right)\quad &(i=N)\ .
# \end{align}
#
# This results in the following tridiagonal system of linear equations
#
# $$
# A \cdot \vec{\psi} = \vec{s}\Rightarrow \vec{\psi} = A^{-1}\cdot\vec{s}\ ,
# $$
#
# where
#
# $$
# A=\begin{pmatrix}
# \left(\pi\Delta r^{2}\Phi^{2}_{1} - 1 - \frac{\Delta r}{r_{1}}\right) & \left(1+\frac{\Delta r}{r_{1}}\right) & 0 & 0 & 0 & 0 & 0\\
# \left(1-\frac{\Delta r}{r_{2}}\right) & \left(\pi\Delta r^{2}\Phi_{2}^{2}-2\right) & \left(1+\frac{\Delta r}{r_{2}}\right) & 0 & 0 & 0 & 0\\
# 0 & \ddots & \ddots & \ddots & 0 & 0 & 0\\
# 0 & 0 & \left(1-\frac{\Delta r}{r_{i}}\right) & \left(\pi\Delta r^{2}\Phi_{i}^{2}-2\right) & \left(1+\frac{\Delta r}{r_{i}}\right) & 0 & 0\\
# 0 & 0 & 0 & \ddots & \ddots & \ddots & 0\\
# 0 & 0 & 0 & 0 & \left(1-\frac{\Delta r}{r_{N-1}}\right) & \left(\pi\Delta r^{2}\Phi_{N-1}^{2}-2\right) & \left(1+\frac{\Delta r}{r_{N-1}}\right)\\
# 0 & 0 & 0 & 0 & 0 & 2 & \left[\pi\Delta r^{2}\Phi^{2}_{N} - 2 - \frac{2\Delta r}{r_{N}}\left(1+\frac{\Delta r}{r_{N}}\right)\right]
# \end{pmatrix}\ ,
# $$
#
# $$
# \vec{\psi} =
# \begin{pmatrix}
# \psi_{1}\\
# \psi_{2}\\
# \vdots\\
# \psi_{i}\\
# \vdots\\
# \psi_{N-1}\\
# \psi_{N}
# \end{pmatrix}\ ,
# $$
#
# and
#
# $$
# \vec{s} =
# \begin{pmatrix}
# 0\\
# 0\\
# \vdots\\
# 0\\
# \vdots\\
# 0\\
# -\frac{2\Delta r}{r_{N}}\left(1+\frac{\Delta r}{r_{N}}\right)
# \end{pmatrix}
# $$
#
# <a id='id_tridiagonal_matrix'></a>
#
# #### Step 1.c.ii.1: The tridiagonal matrix: $A$ \[Back to [top](#toc)\]
# $$\label{id_tridiagonal_matrix}$$
#
# We now start solving the tridiagonal linear system. We start by implementing the tridiagonal matrix $A$ defined above. We break down it down by implementing each diagonal into an array. We start by looking at the main diagonal:
#
# $$
# {\rm diag}_{\rm main}
# =
# \begin{pmatrix}
# \left(\pi\Delta r^{2}\Phi^{2}_{1} - 1 - \frac{\Delta r}{r_{1}}\right)\\
# \left(\pi\Delta r^{2}\Phi_{2}^{2}-2\right)\\
# \vdots\\
# \left(\pi\Delta r^{2}\Phi_{i}^{2}-2\right)\\
# \vdots\\
# \left(\pi\Delta r^{2}\Phi_{N-1}^{2}-2\right)\\
# \left[\pi\Delta r^{2}\Phi^{2}_{N} - 2 - \frac{2\Delta r}{r_{N}}\left(1+\frac{\Delta r}{r_{N}}\right)\right]\\
# \end{pmatrix}
# =
# \begin{pmatrix}
# \left(\pi\Delta r^{2}\Phi^{2}_{1} - 2\right)\\
# \left(\pi\Delta r^{2}\Phi_{2}^{2} - 2\right)\\
# \vdots\\
# \left(\pi\Delta r^{2}\Phi_{i}^{2} - 2\right)\\
# \vdots\\
# \left(\pi\Delta r^{2}\Phi_{N-1}^{2}-2\right)\\
# \left(\pi\Delta r^{2}\Phi^{2}_{N} - 2\right)\\
# \end{pmatrix}
# +
# \left.\begin{pmatrix}
# 1 - \frac{\Delta r}{r_{1}}\\
# 0\\
# \vdots\\
# 0\\
# \vdots\\
# 0\\
# - \frac{2\Delta r}{r_{N}}\left(1+\frac{\Delta r}{r_{N}}\right)
# \end{pmatrix}\quad \right\}\text{N elements}
# $$
# +
# Set the main diagonal
main_diag = np.pi * dr2 * Phi(phi0,r,r0,sigma)**2 - 2
# Update the first element of the main diagonal
main_diag[0] += 1 - dr[0]/r[0]
# Update the last element of the main diagonal
main_diag[NR-1] += - (2 * dr[NR-1] / r[NR-1])*(1 + dr[NR-1] / r[NR-1])
# -
# Then we look at the upper diagonal of the A matrix:
#
# $$
# {\rm diag}_{\rm upper}
# =
# \left.\begin{pmatrix}
# 1+\frac{\Delta r}{r_{1}}\\
# 1+\frac{\Delta r}{r_{2}}\\
# \vdots\\
# 1+\frac{\Delta r}{r_{i}}\\
# \vdots\\
# 1+\frac{\Delta r}{r_{N-2}}\\
# 1+\frac{\Delta r}{r_{N-1}}
# \end{pmatrix}\quad\right\}\text{N-1 elements}
# $$
# Set the upper diagonal, ignoring the last point in the r array
upper_diag = np.zeros(NR)
upper_diag[1:] = 1 + dr[:-1]/r[:-1]
# Finally, we look at the lower diagonal of the A matrix:
#
# $$
# {\rm diag}_{\rm lower}
# =
# \left.\begin{pmatrix}
# 1-\frac{\Delta r}{r_{2}}\\
# 1-\frac{\Delta r}{r_{3}}\\
# \vdots\\
# 1-\frac{\Delta r}{r_{i+1}}\\
# \vdots\\
# 1-\frac{\Delta r}{r_{N-1}}\\
# 2
# \end{pmatrix}\quad\right\}\text{N-1 elements}
# $$
# +
# Set the lower diagonal, start counting the r array at the second element
lower_diag = np.zeros(NR)
lower_diag[:-1] = 1 - dr[1:]/r[1:]
# Change the last term in the lower diagonal to its correct value
lower_diag[NR-2] = 2
# -
# Finally, we construct the tridiagonal matrix by adding the three diagonals, while shifting the upper and lower diagonals to the right and left, respectively. Because A is a sparse matrix, we will also use scipy to solve the linear system faster.
# !pip install scipy >/dev/null
# +
# Set the sparse matrix A by adding up the three diagonals
A = spdiags([main_diag,upper_diag,lower_diag],[0,1,-1],NR,NR)
# Then compress the sparse matrix A column wise, so that SciPy can invert it later
A = csc_matrix(A)
# -
# <a id='id_tridiagonal_rhs'></a>
#
# #### Step 1.c.ii.2 The right-hand side of the linear system: $\vec{s}$ \[Back to [top](#toc)\]
# $$\label{id_tridiagonal_rhs}$$
#
# We now focus our attention to the implementation of the $\vec{s}$ vector:
#
# $$
# \vec{s} =
# \begin{pmatrix}
# 0\\
# 0\\
# \vdots\\
# 0\\
# \vdots\\
# 0\\
# -\frac{2\Delta r}{r_{N}}\left(1+\frac{\Delta r}{r_{N}}\right)
# \end{pmatrix}
# $$
# +
# Set up the right-hand side of the linear system: s
s = np.zeros(NR)
# Update the last entry of the vector s
s[NR-1] = - (2 * dr[NR-1] / r[NR-1])*(1 + dr[NR-1] / r[NR-1])
# Compress the vector s column-wise
s = csc_matrix(s)
# -
# <a id='id_conformal_factor'></a>
#
# #### Step 1.c.ii.3 The conformal factor: $\psi$ \[Back to [top](#toc)\]
# $$\label{id_conformal_factor}$$
#
# We now use scipy to solve the sparse linear system of equations and determine the conformal factor $\psi$.
# Solve the sparse linear system using scipy
# https://docs.scipy.org/doc/scipy-0.14.0/reference/generated/scipy.sparse.linalg.spsolve.html
psi = spsolve(A, s.T)
# We then show useful plots of the conformal factor $\psi$ and of the *evolved conformal factors*
#
# \begin{align}
# \phi &= \log\psi\ ,\\
# W &= \psi^{-2}\ ,\\
# \chi &= \psi^{-4}\ .
# \end{align}
# +
import matplotlib.pyplot as plt
# Compute phi
phi = np.log(psi)
# Compute W
W = psi**(-2)
# Compute chi
chi = psi**(-4)
f = plt.figure(figsize=(12,8),dpi=100)
ax = f.add_subplot(221)
ax.set_title(r"Conformal factor $\psi(0,r)$")
ax.set_ylabel(r"$\psi(0,r)$")
ax.plot(r,psi,'k-')
ax.grid()
ax2 = f.add_subplot(222)
ax2.set_title(r"Evolved conformal factor $\phi(0,r)$")
ax2.set_ylabel(r"$\phi(0,r)$")
ax2.plot(r,phi,'r-')
ax2.grid()
ax3 = f.add_subplot(223)
ax3.set_title(r"Evolved conformal factor $W(0,r)$")
ax3.set_xlabel(r"$r$")
ax3.set_ylabel(r"$W(0,r)$")
ax3.plot(r,W,'b-')
ax3.grid()
ax4 = f.add_subplot(224)
ax4.set_title(r"Evolved conformal factor $\chi(0,r)$")
ax4.set_xlabel(r"$r$")
ax4.set_ylabel(r"$\chi(0,r)$")
ax4.plot(r,chi,'c-')
ax4.grid()
outfile = os.path.join(Ccodesdir,"cfs_scalarfield_id.png")
plt.savefig(outfile)
plt.close(f)
# Display the figure
from IPython.display import Image
Image(outfile)
# -
# <a id='id_lapse_function'></a>
#
# ## Step 1.d The lapse function: $\tilde\alpha$ \[Back to [top](#toc)\]
# $$\label{id_lapse_function}$$
#
# There are two common initial conditions for $\tilde\alpha$. The first one is eq. (44) of [A&C](https://arxiv.org/pdf/1508.01614.pdf), namely setting the lapse to unity
#
# $$
# \tilde\alpha = 1\ .
# $$
# Set the unity lapse initial condition
alpha_unity = np.ones(NR)
# The second one is discussed in the last paragraph of section II.B in [Baumgarte](https://arxiv.org/pdf/1807.10342.pdf), which is to set the "pre-collapsed lapse"
#
# $$
# \tilde\alpha = \psi^{-2}\ .
# $$
# Set the "pre-collapsed lapse" initial condition
alpha_precollapsed = psi**(-2)
# <a id='id_output'></a>
#
# ## Step 1.e Outputting the initial data to file \[Back to [top](#toc)\]
# $$\label{id_output}$$
# Check to see which version of Python is being used
# For a machine running the final release of Python 3.7.1,
# sys.version_info should return the tuple [3,7,1,'final',0]
if sys.version_info[0] == 3:
np.savetxt(os.path.join(Ccodesdir,"outputSFID_unity_lapse.txt"), list(zip( r, ID_sf, psi**4, alpha_unity )),
fmt="%.15e")
np.savetxt(os.path.join(Ccodesdir,"outputSFID_precollapsed_lapse.txt"), list(zip( r, ID_sf, psi**4, alpha_precollapsed )),
fmt="%.15e")
elif sys.version_info[0] == 2:
np.savetxt(os.path.join(Ccodesdir,"outputSFID_unity_lapse.txt"), zip( r, ID_sf, psi**4, alpha_unity ),
fmt="%.15e")
np.savetxt(os.path.join(Ccodesdir,"outputSFID_precollapsed_lapse.txt"), zip( r, ID_sf, psi**4, alpha_precollapsed ),
fmt="%.15e")
# <a id='id_interpolation_files'></a>
#
# # Step 2: Interpolating the initial data file as needed \[Back to [top](#toc)\]
# $$\label{id_interpolation_files}$$
#
# In order to use the initial data file properly, we must tell the program how to interpolate the values we just computed to the values of $r$ in our numerical grid. We do this by creating two C functions: one that interpolates the ADM quantities, $\left\{\gamma_{ij},K_{ij},\alpha,\beta^{i},B^{i}\right\}$, and one that interpolates the scalar field quantities, $\left\{\varphi,\Pi\right\}$. The two files written below use the scalarfield_interpolate_1D( ) function, which is defined in the [ScalarField/ScalarField_interp.h](../edit/ScalarField/ScalarField_interp.h) file. This function performs a Lagrange polynomial interpolation between the initial data file and the numerical grid used during the simulation.
def ID_scalarfield_ADM_quantities(Ccodesdir=".",new_way=False):
includes = ["NRPy_basic_defines.h", "NRPy_function_prototypes.h"]
desc = """(c) 2021 <NAME>
This function takes as input either (x,y,z) or (r,th,ph) and outputs
all ADM quantities in the Cartesian or Spherical basis, respectively.
"""
c_type = "void"
name = "ID_scalarfield_ADM_quantities"
params = """const REAL xyz_or_rthph[3],const ID_inputs other_inputs,
REAL *restrict gammaDD00,REAL *restrict gammaDD01,REAL *restrict gammaDD02,
REAL *restrict gammaDD11,REAL *restrict gammaDD12,REAL *restrict gammaDD22,
REAL *restrict KDD00,REAL *restrict KDD01,REAL *restrict KDD02,
REAL *restrict KDD11,REAL *restrict KDD12,REAL *restrict KDD22,
REAL *restrict alpha,
REAL *restrict betaU0,REAL *restrict betaU1,REAL *restrict betaU2,
REAL *restrict BU0,REAL *restrict BU1,REAL *restrict BU2"""
body = """
const REAL r = xyz_or_rthph[0];
const REAL th = xyz_or_rthph[1];
const REAL ph = xyz_or_rthph[2];
REAL sf_star,psi4_star,alpha_star;
scalarfield_interpolate_1D(r,
other_inputs.interp_stencil_size,
other_inputs.numlines_in_file,
other_inputs.r_arr,
other_inputs.sf_arr,
other_inputs.psi4_arr,
other_inputs.alpha_arr,
&sf_star,&psi4_star,&alpha_star);
// Update alpha
*alpha = alpha_star;
// gamma_{rr} = psi^4
*gammaDD00 = psi4_star;
// gamma_{thth} = psi^4 r^2
*gammaDD11 = psi4_star*r*r;
// gamma_{phph} = psi^4 r^2 sin^2(th)
*gammaDD22 = psi4_star*r*r*sin(th)*sin(th);
// All other quantities ARE ZERO:
*gammaDD01 = 0.0; *gammaDD02 = 0.0;
/**/ *gammaDD12 = 0.0;
*KDD00 = 0.0; *KDD01 = 0.0; *KDD02 = 0.0;
/**/ *KDD11 = 0.0; *KDD12 = 0.0;
/**/ *KDD22 = 0.0;
*betaU0 = 0.0; *betaU1 = 0.0; *betaU2 = 0.0;
*BU0 = 0.0; *BU1 = 0.0; *BU2 = 0.0;
"""
if new_way == True:
outC.add_to_Cfunction_dict(includes=includes,desc=desc,c_type=c_type,name=name,
params=params,body=body,enableCparameters=False)
else:
outfile = os.path.join(Ccodesdir,"ID_scalarfield_ADM_quantities-validation.h")
outC.outCfunction(outfile=outfile,
includes=None,desc=desc,c_type=c_type,name=name,
params=params,body=body,enableCparameters=False)
def ID_scalarfield_spherical(Ccodesdir=".",new_way=False):
includes = ["NRPy_basic_defines.h", "NRPy_function_prototypes.h"]
desc = """(c) 2021 <NAME>
This function takes as input either (x,y,z) or (r,th,ph) and outputs all
scalar field quantities in the Cartesian or Spherical basis, respectively.
"""
c_type = "void"
name = "ID_scalarfield_spherical"
params = "const REAL xyz_or_rthph[3],const ID_inputs other_inputs,REAL *restrict sf,REAL *restrict sfM"
body = """
const REAL r = xyz_or_rthph[0];
const REAL th = xyz_or_rthph[1];
const REAL ph = xyz_or_rthph[2];
REAL sf_star,psi4_star,alpha_star;
scalarfield_interpolate_1D(r,
other_inputs.interp_stencil_size,
other_inputs.numlines_in_file,
other_inputs.r_arr,
other_inputs.sf_arr,
other_inputs.psi4_arr,
other_inputs.alpha_arr,
&sf_star,&psi4_star,&alpha_star);
// Update varphi
*sf = sf_star;
// Update Pi
*sfM = 0;
"""
if new_way == True:
outC.add_to_Cfunction_dict(includes=includes,desc=desc,c_type=c_type,name=name,
params=params,body=body,enableCparameters=False)
else:
outfile = os.path.join(Ccodesdir,"ID_scalarfield_spherical-validation.h")
outC.outCfunction(outfile=outfile,
includes=None,desc=desc,c_type=c_type,name=name,
params=params,body=body,enableCparameters=False)
# <a id='id_sph_to_curvilinear'></a>
#
# # Step 3: Converting Spherical initial data to Curvilinear initial data \[Back to [top](#toc)\]
# $$\label{id_sph_to_curvilinear}$$
#
# In this tutorial module we have explained how to obtain spherically symmetric, time-symmetric initial data for the collapse of a massless scalar field in Spherical coordinates (see [Step 1](#initial_data)). We have also explained how to interpolate the initial data file to the numerical grid we will use during the simulation (see [Step 2](#id_interpolation_files)).
#
# NRPy+ is capable of generating the BSSN evolution equations in many different Curvilinear coordinates (for example SinhSpherical coordinates, which are of particular interest for this problem). Therefore, it is essential that we convert the Spherical initial data generated here to any Curvilinear system supported by NRPy+.
#
# We start by calling the reference_metric() function within the [reference_metric.py](../edit/reference_metric.py) NRPy+ module. This will set up a variety of useful quantities for us.
# Then the code below interpolate the values of the Spherical grid $\left\{r,\theta,\phi\right\}$ to the Curvilinear grid $\left\{{\rm xx0,xx1,xx2}\right\}$.
def ID_scalarfield_xx0xx1xx2_to_BSSN_xx0xx1xx2(Ccodesdir=".",pointer_to_ID_inputs=False,new_way=False):
rfm.reference_metric()
rthph = outC.outputC(rfm.xxSph[0:3],["rthph[0]", "rthph[1]", "rthph[2]"],
"returnstring", "includebraces=False,outCverbose=False,preindent=1")
includes = ["NRPy_basic_defines.h", "NRPy_function_prototypes.h"]
desc = """(c) 2021 <NAME>
This function takes as input either (x,y,z) or (r,th,ph) and outputs all
scalar field quantities in the Cartesian or Spherical basis, respectively.
"""
c_type = "void"
name = "ID_scalarfield_xx0xx1xx2_to_BSSN_xx0xx1xx2"
params = "const paramstruct *restrict params,const REAL xx0xx1xx2[3],\n"
if pointer_to_ID_inputs == True:
params += "ID_inputs *other_inputs,\n"
else:
params += "ID_inputs other_inputs,\n"
params += "REAL *restrict sf, REAL *restrict sfM"
body = """
const REAL xx0 = xx0xx1xx2[0];
const REAL xx1 = xx0xx1xx2[1];
const REAL xx2 = xx0xx1xx2[2];
REAL rthph[3];
"""+rthph+"""
ID_scalarfield_spherical(rthph,other_inputs,sf,sfM);
"""
if new_way == True:
outC.add_to_Cfunction_dict(includes=includes,desc=desc,c_type=c_type,name=name,
params=params,body=body)
else:
outfile = os.path.join(Ccodesdir,"ID_scalarfield_xx0xx1xx2_to_BSSN_xx0xx1xx2-validation.h")
outC.outCfunction(outfile=outfile,
includes=None,desc=desc,c_type=c_type,name=name,
params=params,body=body)
# Finally, we create the driver function which puts everything together using OpenMP.
def ID_scalarfield(Ccodesdir=".",new_way=False):
includes = ["NRPy_basic_defines.h", "NRPy_function_prototypes.h"]
desc = """(c) 2021 <NAME>
This is the scalar field initial data driver functiono.
"""
c_type = "void"
name = "ID_scalarfield"
params = """const paramstruct *restrict params,REAL *restrict xx[3],
ID_inputs other_inputs,REAL *restrict in_gfs"""
body = """
const int idx = IDX3S(i0,i1,i2);
const REAL xx0xx1xx2[3] = {xx0,xx1,xx2};
ID_scalarfield_xx0xx1xx2_to_BSSN_xx0xx1xx2(params,xx0xx1xx2,other_inputs,
&in_gfs[IDX4ptS(SFGF,idx)],
&in_gfs[IDX4ptS(SFMGF,idx)]);
"""
loopopts = "AllPoints,Read_xxs"
if new_way == True:
outC.add_to_Cfunction_dict(includes=includes,desc=desc,c_type=c_type,name=name,
params=params,body=body,loopopts=loopopts)
else:
outfile = os.path.join(Ccodesdir,"ID_scalarfield-validation.h")
outC.outCfunction(outfile=outfile,
includes=None,desc=desc,c_type=c_type,name=name,
params=params,body=body,loopopts=loopopts)
def NRPy_param_funcs_register_C_functions_and_NRPy_basic_defines(Ccodesdir=".",pointer_to_ID_inputs=False,new_way=False):
ID_scalarfield_ADM_quantities(Ccodesdir=Ccodesdir,new_way=new_way)
ID_scalarfield_spherical(Ccodesdir=Ccodesdir,new_way=new_way)
ID_scalarfield_xx0xx1xx2_to_BSSN_xx0xx1xx2(Ccodesdir=Ccodesdir,pointer_to_ID_inputs=pointer_to_ID_inputs,new_way=new_way)
ID_scalarfield(Ccodesdir=Ccodesdir,new_way=new_way)
NRPy_param_funcs_register_C_functions_and_NRPy_basic_defines(Ccodesdir=Ccodesdir)
# <a id='validation'></a>
#
# # Step 4: Validation of this tutorial against the [ScalarField/ScalarField_InitialData.py](../edit/ScalarField/ScalarField_InitialData.py) module \[Back to [top](#toc)\]
# $$\label{validation}$$
#
# First we load the [ScalarField/ScalarField_InitialData.py](../edit/ScalarField/ScalarField_InitialData.py) module and compute everything by using the scalarfield_initial_data( ) function, which should do exactly the same as we have done in this tutorial.
# +
# Import the ScalarField.ScalarField_InitialData NRPy module
import ScalarField.ScalarField_InitialData as sfid
# Output the unity lapse initial data file
outputname = os.path.join(Ccodesdir,"outputSFID_unity_lapse-validation.txt")
sfid.ScalarField_InitialData(outputname,ID_Family,
phi0,r0,sigma,NR,RMAX,CoordSystem=CoordSystem,
sinhA=sinhA,sinhW=sinhW,lapse_condition="Unity")
# Output the "pre-collapsed" lapse initial data file
outputname = os.path.join(Ccodesdir,"outputSFID_precollapsed_lapse-validation.txt")
sfid.ScalarField_InitialData(outputname,ID_Family,
phi0,r0,sigma,NR,RMAX,CoordSystem=CoordSystem,
sinhA=sinhA,sinhW=sinhW,lapse_condition="Pre-collapsed")
# Output C codes
sfid.NRPy_param_funcs_register_C_functions_and_NRPy_basic_defines(Ccodesdir=Ccodesdir)
# +
import filecmp
if filecmp.cmp(os.path.join(Ccodesdir,'outputSFID_unity_lapse.txt'),
os.path.join(Ccodesdir,'outputSFID_unity_lapse-validation.txt')) == False:
print("ERROR: Unity lapse initial data test FAILED!")
sys.exit(1)
else:
print(" Unity lapse initial data test: PASSED!")
if filecmp.cmp(os.path.join(Ccodesdir,'outputSFID_precollapsed_lapse.txt'),
os.path.join(Ccodesdir,'outputSFID_precollapsed_lapse-validation.txt')) == False:
print("ERROR: \"Pre-collapsed\" lapse initial data test FAILED!")
sys.exit(1)
else:
print(" \"Pre-collapsed\" lapse initial data test: PASSED!")
if filecmp.cmp(os.path.join(Ccodesdir,'ID_scalarfield_ADM_quantities.h'),
os.path.join(Ccodesdir,'ID_scalarfield_ADM_quantities-validation.h')) == False:
print("ERROR: ADM quantities interpolation file test FAILED!")
sys.exit(1)
else:
print(" ADM quantities interpolation file test: PASSED!")
if filecmp.cmp(os.path.join(Ccodesdir,'ID_scalarfield_spherical.h'),
os.path.join(Ccodesdir,'ID_scalarfield_spherical-validation.h')) == False:
print("ERROR: Scalar field interpolation file test FAILED!")
sys.exit(1)
else:
print(" Scalar field interpolation file test: PASSED!")
if filecmp.cmp(os.path.join(Ccodesdir,'ID_scalarfield_xx0xx1xx2_to_BSSN_xx0xx1xx2.h'),
os.path.join(Ccodesdir,'ID_scalarfield_xx0xx1xx2_to_BSSN_xx0xx1xx2-validation.h')) == False:
print("ERROR: Scalar field Spherical to Curvilinear test FAILED!")
sys.exit(1)
else:
print("Scalar field Spherical to Curvilinear test: PASSED!")
if filecmp.cmp(os.path.join(Ccodesdir,'ID_scalarfield.h'),
os.path.join(Ccodesdir,'ID_scalarfield-validation.h')) == False:
print("ERROR: Scalar field driver test: FAILED!")
sys.exit(1)
else:
print(" Scalar field driver test: PASSED!")
# -
# <a id='output_to_pdf'></a>
#
# # Step 5: Output this module as $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
# $$\label{output_to_pdf}$$
#
# The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
# [Tutorial-ADM_Initial_Data-ScalarField.pdf](Tutorial-ADM_Initial_Data-ScalarField.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
cmd.output_Jupyter_notebook_to_LaTeXed_PDF("Tutorial-ADM_Initial_Data-ScalarField")
| Tutorial-ADM_Initial_Data-ScalarField.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# In this empiric test we look at 'runs' in the random sequence. A run is a strict strictly monotonically increasing subsequence.
#
# sequence sub sequences runs run counts
# 1,2,9,8,5,3,6,7,0,4 -> 1,2,9|8|5|3,6,7|0,4 -> 3,1,1,3,2 -> 2,1,2,0,0,0
#
# Since adjacent run counts are not independent, we can not directly apply a chi-square test, but must use a special statistic
#
# V = 1/(n-6) \sum_{i=1}^{6}\sum_{j=1}^{6} (count[i] - n*B[i]) (count[j] - n*B[j]) a[i][j]
#
# With special matrices A and B, that he explains how to derrive.
#
# The statistic V should have a chi-square distribution with 6 degrees of freedom.
#
# 0.90 -> 10.645
# 0.95 -> 12.59
#
# ## Code
# +
import itertools
from collections import Counter
import random
def getPseudoRandomSequence(x0, k, m):
last = x0
while True:
last = k * last % m
yield last/m
def getFinitePseudorandomSequence(x0, k, m, n):
return itertools.islice(getPseudoRandomSequence(x0, k, m),n)
def getPythonRandom():
while True:
yield random.random()
def getOsRandom():
os_random = random.SystemRandom()
while True:
yield os_random.random()
# +
def getRuns(sequence, condition):
currentRun = []
for element in sequence:
if not currentRun or condition(currentRun[-1], element):
currentRun.append(element)
else:
yield currentRun
currentRun = [element]
yield currentRun
def getIncreasingRuns(sequence):
return getRuns(sequence, lambda a,b: a<b)
def getDecreasingRuns(sequence):
return getRuns(sequence, lambda a,b: a>b)
def getRunLenghts(runs):
for element in runs:
yield len(element)
def getIncreasingRunLenghts(sequence):
return getRunLenghts(getIncreasingRuns(sequence))
def getDecreasingRunLenghts(sequence):
return getRunLenghts(getDecreasingRuns(sequence))
def getRunLenghtCounts(runLenghts):
counter = dict(Counter(runLenghts))
counts = [counter[key] for key in sorted(counter.keys())]
counts[5:] = [sum(counts[5:])]
return counts
def getIncreasingRunLenghtCounts(sequence):
return getRunLenghtCounts(getRunLenghts(getIncreasingRuns(sequence)))
def getDecreasingRunLenghtCounts(sequence):
return getRunLenghtCounts(getRunLenghts(getDecreasingRuns(sequence)))
# -
A = [[ 4529.4, 9044.9, 13568.0, 18091.0, 22615.0, 27892.0],
[ 9044.9, 18097.0, 27139.0, 36187.0, 45234.0, 55789.0],
[ 13568.0, 27139.0, 40721.0, 54281.0, 67852.0, 83685.0],
[ 18091.0, 36187.0, 54281.0, 72414.0, 90470.0, 111580.0],
[ 22615.0, 45234.0, 67852.0, 90470.0, 113262.0, 139476.0],
[ 27892.0, 55789.0, 83685.0, 111580.0, 139476.0, 172860.0]]
B = [1/6, 5/24, 11/120, 19/720, 29/5040, 1/840]
def getStatisticV(count, n):
return 1/(n-6) * sum([(count[i]-n*B[i])*(count[j]-n*B[j])*A[i][j] for i in range(0,6) for j in range(0,6)])
# sequence = [1,2,9,8,5,3,6,7,0,4]
# sequence = [1,3,8,7,5,2,6,7,1,6]
x0 = 1001
k = 8192
m = 67101323
n = 10000
sequence = list(getFinitePseudorandomSequence(x0, k, m, n))
print(getStatisticV(getIncreasingRunLenghtCounts(sequence), n))
print(getStatisticV(getDecreasingRunLenghtCounts(sequence), n))
# ## Test Runs
#
# Values for generated sequences taken from:
# ```
# The Runs-Up and Runs-Down Tests By <NAME>
# Journal of the Royal Statistical Society. Series C (Applied Statistics), Vol. 30, No. 1 (1981), pp. 81-85```
s1 = getPseudoRandomSequence(x0=1001, k=8192, m= 67101323)
s2 = getPseudoRandomSequence(x0=1001, k=8192, m= 67099547)
s3 = getPseudoRandomSequence(x0=1001, k=32768, m= 16775723)
s4 = getPseudoRandomSequence(x0=1001, k=54751, m= 99707)
s5 = getPseudoRandomSequence(x0=1001, k=8, m= 67100963)
s6 = getPseudoRandomSequence(x0=1001, k=32, m= 7999787)
s7 = getPythonRandom()
s8 = getOsRandom()
size = [4000,10000,15000,20000,25000,30000,35000,40000]
size.extend(size)
size.extend(size)
size.sort()
# +
import plotly
def getCoordsForGraph(sequence, size):
# sequence = list(itertools.islice(sequence, max(size)))
# data = {
# 'up': [getStatisticV(getIncreasingRunLenghtCounts(sequence[:n]), n) for n in size],
# 'down': [getStatisticV(getDecreasingRunLenghtCounts(sequence[:n]), n) for n in size],
# }
# print(data)
graphs = {}
graphs['sequence'] = plotly.graph_objs.Scatter(
x = list(range(1, max(size)+1)),
y = list(itertools.islice(sequence, max(size))),
mode = 'markers',
marker = dict(size=1, line=dict(width=0), color='blue'),
name = 'sequence'
)
graphs['up'] = plotly.graph_objs.Scatter(
x = size,
y = [getStatisticV(getIncreasingRunLenghtCounts(itertools.islice(sequence, n)), n) for n in size],
mode = 'markers',
marker = dict(size=8, line=dict(width=1), color='red'),
name = 'runs up'
)
graphs['down'] = plotly.graph_objs.Scatter(
x = size,
y = [getStatisticV(getDecreasingRunLenghtCounts(itertools.islice(sequence, n)), n) for n in size],
mode = 'markers',
marker = dict(size=8, line=dict(width=1), color='blue'),
name = 'runs down'
)
return graphs
def plotRunTest(sequence, size, name, plot_sequence=False):
plot = getCoordsForGraph(sequence, size)
layout = plotly.graph_objs.Layout(
title=name,
xaxis = dict(
title='Size of sequence'
),
yaxis = dict(
title='Statistic V'
),
shapes = [
{ # horizontal line for .90
'type': 'line',
'x0': 0,
'y0': 10.645,
'x1': 45000,
'y1': 10.645,
'line': {
'color': 'orange',
'width': 3,
'dash': 'longdash'
}
},
{ # horizontal line for .95
'type': 'line',
'x0': 0,
'y0': 12.59,
'x1': 45000,
'y1': 12.59,
'line': {
'color': 'darkred',
'width': 3,
'dash': 'dash'
}
}
]
)
if plot_sequence:
plotly.plotly.plot(plotly.graph_objs.Figure(
data = [plot['sequence']],
layout = dict(
title = name
)
),
filename = 'sequence',
auto_open = True
)
plotly.plotly.plot(plotly.graph_objs.Figure(
data = [plot['up'], plot['down']],
layout = layout
),
filename = name,
auto_open = True
)
# -
plotRunTest(s1, size, 'Sequence1', True)
plotRunTest(s2, size, 'Sequence2', True)
plotRunTest(s3, size, 'Sequence3', True)
plotRunTest(s4, size, 'Sequence4', True)
plotRunTest(s5, size, 'Sequence5', True)
plotRunTest(s6, size, 'Sequence6', True)
plotRunTest(s7, size, 'Python Random', True)
plotRunTest(s8, size, 'OS Random', True)
| SS2018-Seminar-In-Computational-Engineering-The-Art-Of-Computer-Programming/Run Test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Convert practical salinity open boundary files to TEOS-10 reference salinity open boundary files
#
# 2016-03-12 : Couldn't get the ncatted to work (segmentation fault). Went back to the PrepareSimpleTS-Johnstone and added TEOS10 calculation at the bottom of the that (made new file from scratch)
import matplotlib.pyplot as plt
import netCDF4 as nc
import numpy as np
from salishsea_tools import nc_tools
from salishsea_tools import teos_tools
# %matplotlib inline
# Read in the original practical salinity file
practical_data = nc.Dataset('../../../NEMO-forcing/open_boundaries/north/SalishSea2_North_tra.nc')
practical_salinity = practical_data.variables['vosaline'][:]
print (practical_salinity.shape)
# Now we have to work around a bug in netcdf4 library. We want to change the attributes of the vosaline
# variable, but this crashes. So instead we go out to the command line and copy the file and change the vosaline attributes. Note ncatted is only available on some machines, e.g. Salish (these are for west file, last use of this notebook was North file)
#
# # cp ../../../NEMO-forcing/open_boundaries/west/SalishSea2_Masson_corrected.nc SalishSea_west_TEOS10.nc
# ncatted -O -a units,vosaline,m,c,'g/kg' SalishSea_west_TEOS10.nc
# ncatted -O -a long_name,vosaline,m,c,'Reference Salinity' SalishSea_west_TEOS10.nc
# ncatted -O -h -a history,global,a,c,'[2016-01-23] Converted to Reference Salinity' SalishSea_west_TEOS10.nc
# ncatted -O -h -a title,global,o,c,'Modified boundary conditions based on weekly climatolgy from <NAME> converted to TEOS' SalishSea_west_TEOS10.nc
# ncatted -O -h -a source,global,a,c,'https://bitbucket.org/salishsea/tools/src/tip/I_ForcingFiles/OBC/TEOSfromPracticalOBC.ipynb' SalishSea_west_TEOS10.nc
# ncatted -O -h -a comment,global,a,c,'Converted to Reference Salinity (TEOS-10)' SalishSea_west_TEOS10.nc
# ncatted -O -h -a references,global,a,c,'https://bitbucket.org/salishsea/nemo-forcing/src/tip/open_boundaries/west/SalishSea_west_TEOS10.nc' SalishSea_west_TEOS10.nc
TEOS_data = nc.Dataset('SalishSea_north_TEOS10.nc', 'r+')
print (nc_tools.show_variable_attrs(TEOS_data))
print (nc_tools.show_dataset_attrs(TEOS_data))
print (practical_salinity[1,20,0,40],teos_tools.psu_teos(practical_salinity[1,20,0,40]))
ref_sal = np.zeros_like(practical_salinity)
ref_sal = teos_tools.psu_teos(practical_salinity)
fig, ax = plt.subplots(1,3,figsize=(15,5))
i = 1; k=0; l1=0; l2=30
ax[0].pcolormesh(practical_salinity[i,:,k,l1:l2])
ax[0].set_title('Practical Salinity')
ax[1].pcolormesh(ref_sal[i,:,k,l1:l2])
ax[1].set_title('Reference Salinity')
ax[2].pcolormesh(ref_sal[i,:,k,l1:l2]/practical_salinity[i,:,k,l1:l2])
ax[2].set_title('Ratio')
print (ref_sal[i,10,k,15]/practical_salinity[i,10,k,15])
vosaline = TEOS_data.variables['vosaline']
vosaline[:] = ref_sal
TEOS_data.close()
| I_ForcingFiles/OBC/TEOSfromPracticalOBC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Stock Linear Correlation Analysis
# Correlation is a measure of the strength of linear between 2 or more stocks. However, when 2 stocks is highly correlated, they tend to move in teh same direction.
# + outputHidden=false inputHidden=false
# Library
import pandas as pd
import numpy as np
import math
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
warnings.filterwarnings("ignore")
from pandas_datareader import data as pdr
import yfinance as yf
yf.pdr_override()
# + outputHidden=false inputHidden=false
start = '2016-01-01'
end = '2019-01-01'
market = 'SPY'
symbol1 = 'AAPL'
symbol2 = 'MSFT'
bench = yf.download(market, start=start, end=end)['Adj Close']
stock1 = yf.download(symbol1, start=start, end=end)['Adj Close']
stock2 = yf.download(symbol2, start=start, end=end)['Adj Close']
# + outputHidden=false inputHidden=false
plt.figure(figsize=(12,8))
plt.scatter(stock1,stock2)
plt.xlabel(symbol1)
plt.ylabel(symbol2)
plt.title('Stock prices from ' + start + ' to ' + end)
# + outputHidden=false inputHidden=false
plt.figure(figsize=(12,8))
plt.scatter(stock1,bench)
plt.xlabel(symbol1)
plt.ylabel(market)
plt.title('Stock prices from ' + start + ' to ' + end)
# + outputHidden=false inputHidden=false
plt.figure(figsize=(12,8))
plt.scatter(stock2,bench)
plt.xlabel(symbol2)
plt.ylabel(market)
plt.title('Stock prices from ' + start + ' to ' + end)
# + outputHidden=false inputHidden=false
print("Correlation coefficients")
print(symbol1 + ' and ' + symbol2 + ':', np.corrcoef(stock1,stock2)[0,1])
print(symbol1 + ' and ' + market + ':', np.corrcoef(stock1,bench)[0,1])
print(market + ' and ' + symbol2 + ':', np.corrcoef(bench,stock2)[0,1])
# + outputHidden=false inputHidden=false
rolling_correlation = stock1.rolling(60).corr(stock2)
plt.figure(figsize=(12,8))
plt.plot(rolling_correlation)
plt.xlabel('Day')
plt.ylabel('60-day Rolling Correlation')
| Python_Stock/Stock_Linear_Correlation_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Working with scripts
#
# So far this week we've been running all our Python commands in the notebook, which is a kind of combination between a command-line Python interpreter and a text editor. This works well as a teaching environment (and also for writing up things like blogs or code demonstrations, since you can combine plain text with code and its output), but it's actually not commonly used for day-to-day programming work. More commonly, you will want to create dedicated Python files. (These files are often called 'scripts', or 'codes' - we will be using all of these terms interchangeably.) Some text editors like Spyder include a Python interpreter which allows you to run your Python files inside the editor itself, as we have been with the Jupyter notebooks. However, you may not always have this option and will need to learn how to work with Python scripts through the command line.
#
# For this lesson we will be working in a text editor. For this we recommend you use the one you were using yesterday with bash and git, but if you already have another editor you prefer to use for writing Python files, feel free to use that instead. Open your text editor now and open a new file. Call it something like `rcsc18-data-analysis.py` (descriptive filenames are usually a good idea). Here, the `.py` file extension indicates that this is a Python file - you should always include this in the names of your Python scripts.
#
# In this file, let's recreate some of the work we did in the last lesson, analysing data and detecting problems. Here's some of the code we wrote:
# +
import glob
import numpy as np
import matplotlib.pyplot as plt
def analyse(filename):
"""
Reads data from the specified file and plots the average, maximum and minimum along the first axis of the data.
Parameters
----------
filename : str
Name or path to a file containing data to be plotted. Data should be 2-dimensional and values should be
separated by commas.
Examples
--------
>>> analyse('/path/to/mydata.dat')
"""
data = np.loadtxt(fname=filename, delimiter=',')
fig = plt.figure(figsize=(10.0, 3.0))
axes1 = fig.add_subplot(1, 3, 1)
axes2 = fig.add_subplot(1, 3, 2)
axes3 = fig.add_subplot(1, 3, 3)
axes1.set_ylabel('average')
axes1.plot(np.mean(data, axis=0))
axes2.set_ylabel('max')
axes2.plot(np.max(data, axis=0))
axes3.set_ylabel('min')
axes3.plot(np.min(data, axis=0))
fig.tight_layout()
plt.show()
def detect_problems(filename):
"""
Tests data stored in the specified file for spurious or unexpected values.
Parameters
----------
filename : str
Name or path to a file containing data to tested. Data should be 2-dimensional and values should be
separated by commas.
Examples
--------
>>> analyse('/path/to/mydata.dat')
"""
data = numpy.loadtxt(fname=filename, delimiter=',')
if np.max(data, axis=0)[0] == 0 and np.max(data, axis=0)[20] == 20:
print('Suspicious looking maxima!')
elif np.sum(np.min(data, axis=0)) == 0:
print('Minima add up to zero!')
else:
print('Seems OK!')
filenames = sorted(glob.glob('inflammation*.csv')) # is equivalent to glob.glob(filname) -> filename.sort()
for f in filenames[:3]:
print(f)
analyse(f)
detect_problems(f)
# -
#
# <section class="callout panel panel-warning">
# <div class="panel-heading">
# <h2><span class="fa fa-thumb-tack"></span> Copying code</h2>
# </div>
#
#
# <div class="panel-body">
#
# <p>From time to time, you will end up being in a situation where you want to use code from somewhere else, perhaps from another script you've written, from some code like the above that you are being given, or from an online forum. When you do this we recommend that you copy and paste the code rather than retyping it all out yourself. This is often much faster, of course, but just as importantly it reduces the chance of mistyping something and introducing errors into the code (assuming it was correct to start with, of course). If you're working in a command-line editor you may find that the keyboard shortcut for pasting text is not the one you're used to, but if you can't find the appropriate keys, right-clicking the mouse and selecting 'Paste' should still work.</p>
# <p><strong>ALWAYS</strong> check carefully that you have permission to use whatever code you're copying before doing so. This will depend on where you're getting the code from, what you intend to use it for and what license it's published under, if any (more on licenses in a later lesson).</p>
#
# </div>
#
# </section>
#
# Copy the above into your new file and save it. Now, go to the command line (if you are editing your Python file in vim, nano or another command-line editor, either close it or open a new terminal and navigate to where your file is saved). We saw earlier that you can access the Python interpreter in the terminal with the `python` command, but this command can also take a filename as an argument, like this:
#
# ```bash
# python rcsc18-data-analysis.py
# ```
#
# Used like this, Python will read the contents of the file and start running the commands it contains from the top, until it gets to the bottom of the file or encounters an error. Run the command above now. You should get all the same output produced by that code when you ran it in the notebook, but now this output appears in the terminal instead.
#
# There are a number of advantages to running code from a script rather than in the notebook. First, with this approach we can now use all the tricks we learned in the [bash session](link to the bash session) - we can pass flags to the `python` command to change its behaviour, we can capture the output and pipe it to another command or a file, and so on. Also, if you find that you have to do some work on a remote server to which you only have command-line access (not an uncommon occurrence), you will have to be comfortable using Python in the terminal, since graphical interfaces like the notebook will not be available. Finally, and perhaps most importantly for the purposes of this summer school, plain-text formats such as `.py` files are much easier to track with git than notebooks are (because notebooks contain a lot of additional formatting as well as the text itself).
| 04-further-python/01-scripts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import requests
import gmaps
import matplotlib.pyplot as plt
from config import gKey
gmaps.configure(api_key = gKey)
apiURL = "https://maps.googleapis.com/maps/api/place/nearbysearch/json"
cityWeatherPath = "../WeatherPy/output/cities.csv"
cityWeather_df = pd.read_csv(cityWeatherPath)
# +
#humidity heatmap
humidity = cityWeather_df["Humidity:"]
coordinates = cityWeather_df[["Latitude:", "Longitude:"]]
gMap = gmaps.figure()
heatLayer = gmaps.heatmap_layer(coordinates, weights = humidity, dissipating = False, max_intensity = 300, point_radius = 5)
gMap.add_layer(heatLayer)
gMap
# +
#Narrow down cities
narrowedCityWeather_df = cityWeather_df.loc[(cityWeather_df["Max Temp:"].between(65, 75, inclusive = True) )
& (cityWeather_df["Cloud Coverage:"].between(20, 30, inclusive = False))
& (cityWeather_df["Wind:"] < 5)
].dropna()
narrowedCityWeather_df
# +
#create hotel pins for map
hotel_df = narrowedCityWeather_df[["City:","Country:", "Latitude:", "Longitude:"]]
hotel_df["Hotel Name"] = "initialize"
params = {"radius" : 5000,
"types" : "lodging",
"key" : gKey
}
for x, row in hotel_df.iterrows():
latitude = row["Latitude:"]
longitude = row["Longitude:"]
params["location"] = f'{latitude},{longitude}'
print("fetching lodging data")
response = requests.get(apiURL, params = params).json()
try:
hotel_df.loc[x, "Hotel Name"] = response["results"][0]["name"]
except:
print("broaden your horizons, no lodging available")
continue
hotel_df
# +
#template formatting is not a fan of colons apparently
hotel_df = hotel_df.rename(columns = {'City:': 'City', 'Country:' : 'Country'})
info_box_template = """ <dl>
<dt>Name</dt><dd>{Hotel Name}</dd>
<dt>City</dt><dd>{City:}</dd>
<dt>Country</dt><dd>{Country:}</dd>
</dl>
"""
hotelData = [info_box_template.format(**row) for x, row in hotel_df.iterrows()]
coordinates = hotel_df[["Latitude:", "Longitude:"]]
marker_layer = gmaps.marker_layer(coordinates, info_box_content = hotelData)
gMap.add_layer(marker_layer)
gMap
# -
| VacationPy/VacationPy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.0 64-bit (''.venv'': venv)'
# name: python3
# ---
# # Goal
#
# The goal of this notebook is to look at the power usage for rectangular matrices.
# +
# Data preprocessing
import os
FREQUENCY = 905250000 # Available Frequencies: 905250000 and 1377000000
DEVICE = "AGX"
path = "./data/"+DEVICE+"/non-square/"+str(FREQUENCY)+"/"
files = os.listdir(path)
data = []
for file_name in files:
temp = {
# Inputs
"device": "",
"datatype": "",
"matrix_rows": -1,
"matrix_cols": -1,
"tensor": None,
"gpu_frequency": -1,
# Results
"power_usage": [],
"flops": -1,
# Calculates Results
"avg_power": -1,
"flops_per_watt": -1
}
with open(path+file_name) as f:
temp['device'], temp['datatype'], temp['matrix_rows'], temp['matrix_cols'], temp['tensor'], temp['gpu_frequency'] = file_name.split('.')[0].split('-')
temp['matrix_rows'] = int(temp['matrix_rows'])
temp['matrix_cols'] = int(temp['matrix_cols'])
temp['tensor'] = True if temp['tensor'] == "tensor" else False
temp['gpu_frequency'] = int(temp['gpu_frequency'])
file_data = f.readlines()
_, temp['power_usage'] = zip(*[d.strip().split(",") for d in file_data[1:-1]])
temp['power_usage'] = list(map(float, temp['power_usage']))
temp['avg_power'] = sum(temp['power_usage'])/len(temp['power_usage'])
temp['flops'] = float(file_data[-1])
temp['flops_per_watt'] = temp['flops'] / temp['avg_power']
data.append(temp)
print(len(data), "files processed.")
# +
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib widget
# import ipywidgets as widgets
# from IPython.display import display
# plt.ion()
search = {
"device": DEVICE,
"datatype": "float",
"matrix_rows": -1, # 64-1024, step=64
# "matrix_cols": -1, # 8-2048, step=8
"tensor": True,
"gpu_frequency": FREQUENCY,
}
fig, ax = plt.subplots(1, 1, figsize=(10, 7))
ax.set_axisbelow(True)
ax.grid(axis='y')
ax.set_xticks(range(0, 2048+1, 128))
ax.set_title("Flops per Watt by Matrix Size")
ax.set_xlabel("Columns")
ax.set_ylabel("Flops per Watt")
# ax.set_prop_cycle('color', [plt.get_cmap('gist_rainbow')(1.*i/16) for i in range(16)])
for row in range(64, 1024+1, 64):
search['matrix_rows'] = row
results = [d for d in data if search.items() <= d.items() ]
x, y = zip(*sorted([(r['matrix_cols'], r['flops_per_watt']) for r in results], key=lambda d : d[0]))
ax.plot(x, y, label=str(row)+" rows", linestyle='-' if row > 682 else '--' if row > 341 else '-.')
ax.legend(loc='upper left')
plt.show()
| AGX Rectangular.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ### Visual Wake Words Model Trainer
#
# ##### Run as a Jupyter notebook on the GCP AI platform : TF 1.0 : GPU + CUDA
#
# ##### Note : there are explicit calls to /device:GPU:0 so running the training script on a CPU will exception
# ##### Simple test for GPU presence...
# +
import tensorflow as tf
with tf.device('/cpu:0'):
a_c = tf.constant([1.0, 2.0, 3.0, 4.0, 5.0, 6.0], shape=[2, 3], name='a-cpu')
b_c = tf.constant([1.0, 2.0, 3.0, 4.0, 5.0, 6.0], shape=[3, 2], name='b-cpu')
c_c = tf.matmul(a_c, b_c, name='c-cpu')
with tf.device('/gpu:0'):
a_g = tf.constant([1.0, 2.0, 3.0, 4.0, 5.0, 6.0], shape=[2, 3], name='a-gpu')
b_g = tf.constant([1.0, 2.0, 3.0, 4.0, 5.0, 6.0], shape=[3, 2], name='b-gpu')
c_g = tf.matmul(a_g, b_g, name='c-gpu')
with tf.Session(config=tf.ConfigProto(log_device_placement=True)) as sess:
print (sess.run(c_c))
print (sess.run(c_g))
print 'DONE!'
# -
# ##### Train the model after buidling the dataset
# ##### Note : this can take days on a GPU but useable accuracy should be reached after a few hours
# ! python models/research/slim/train_image_classifier.py \
# --train_dir=vww_96_grayscale \
# --dataset_name=visualwakewords \
# --dataset_split_name=train \
# --dataset_dir=./visualwakewords \
# --model_name=mobilenet_v1_025 \
# --preprocessing_name=mobilenet_v1 \
# --train_image_size=96 \
# --input_grayscale=True \
# --save_summaries_secs=300 \
# --learning_rate=0.045 \
# --label_smoothing=0.1 \
# --learning_rate_decay_factor=0.98 \
# --num_epochs_per_decay=2.5 \
# --moving_average_decay=0.9999 \
# --batch_size=96 \
# --max_number_of_steps=1000000
| tensorflow/lite/micro/examples/object_detection/vww_model_trainer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="xfP0Hl6TxW9x"
# (1) Le but de ce projet est de déterminer le *genre musical* d'une chanson en nous basant principalement sur ses paroles. Il s'agit donc d'un cas de classification, puisque l'on cherche la 'classe' (genre musicale) à laquelle appartient une chanson et non d'un régression (extrapolation de valeurs continues).
#
# Bien qu'en réalité le nombre de genres musicaux soit grand, voire très grand selon ce qu'on entend par 'genre', les chansons du jeu de données sont regroupées en 11 genres assez larges comme 'Hip Hop', 'Rock' etc., ce qui réduit le nombre de classes possibles et donc devrait rendre la classification plus facile.
# + colab={"base_uri": "https://localhost:8080/"} id="OBnRjebG1rU5" outputId="549f9b31-9481-43ec-a350-722f1bd458f0"
import itertools
# regular expressions
import re
# Pour la lecture du fichier csv
import requests, zipfile, io
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
import scipy
import nltk
from nltk.corpus import wordnet
from nltk.stem import WordNetLemmatizer, PorterStemmer
nltk.download('stopwords')
nltk.download('punkt')
nltk.download('wordnet')
nltk.download('averaged_perceptron_tagger')
from sklearn.model_selection import train_test_split as tts
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.feature_extraction.text import CountVectorizer
from sklearn import preprocessing, metrics, pipeline, tree, ensemble, naive_bayes
# Permet de décontracter les mots comme "you're" en "you are"
# !pip install contractions
import contractions
import time
# !pip install memory_profiler
# %load_ext memory_profiler
# Pour faire le tableau comparatif des temps d'éxécution
import plotly.graph_objects as go
# + [markdown] id="jgoKa5zWHbrn"
# # Lecture des données
# + id="Jc17QaH-1rU_" colab={"base_uri": "https://localhost:8080/", "height": 699} outputId="ac3231a4-1389-417f-cd62-d5002e9b7095"
# inspiré de https://stackoverflow.com/questions/9419162/download-returned-zip-file-from-url et https://stackoverflow.com/questions/18885175/read-a-zipped-file-as-a-pandas-dataframe
zip_file_url = "https://github.com/hiteshyalamanchili/SongGenreClassification/raw/master/dataset/english_cleaned_lyrics.zip"
r = requests.get(zip_file_url)
with zipfile.ZipFile(io.BytesIO(r.content)) as z:
with z.open("english_cleaned_lyrics.csv") as f:
df = pd.read_csv(f)
display(df)
display(df.info())
# + [markdown] id="9KqshqGkxWsQ"
# (2) La base de donnée utilisée pour le projet contient des centaines de milliers de chansons et pour chacune on a le titre, l'année de parution, l'artiste le genre et les paroles (transcription).
#
# Le nom de l'artiste n'est pas pertinent pour plusieurs raisons:
#
# D'abord, puisque les genres musicaux utilisés sont si larges, très peu d'artistes ont des titres de plusieurs genres différents, donc connaître le genre d'une chanson d'un artiste permettrait de connaître le genre de toutes les chansons de cet artiste.
#
# Ensuite, comme les noms d'artistes ont très rarement un lien direct avec leur genre musical, il ne serait pas vraiment possible de généraliser à partir des données d'entraînement puisque seuls les noms d'artistes connus par l'algorithme seraient utiles à la prédiction.
#
# On omettra aussi le titre, qui ne contient pas assez d'informations (trop uniques).
#
# Dans leur état brut, les paroles ne nous sont pas très utiles parce qu'elles contiennent trop d'information, n'ont pas vraiment de structure ou d'organisation et n'ont pas de standard de transcription. Les étapes de pré-traitement nécessaires seront décrites dans le bloc suivant.
# + [markdown] id="-KxqUuR01rVB"
# # Prétraitement
# + [markdown] id="5g2BNIEy1rVB"
# (3ab, 4c) On retire les deux premières colonnes, qui sont redondantes, ainsi que les colonnes `artist` (interprète) et `song` (titre). Du reste, nous avons 2 variables explicatives. `year` correspond à la date de sortie et `lyrics` aux paroles. Ce sont respectivement des variables quantitatives discrètes et qualitatives nominales. `lyrics` est la variable dont nous allons essayer d'exploiter au maximum le potentiel en utilisant des outils/statistiqus du traitement du langage naturel comme `Term Frequency` (fréquence des mots dans chaque texte), `TF-IDF` et `Bag of words`. La variable qu'on désire prédire est `genre`. Comme on l'a déjà mentionné, Puisqu'il s'agit d'un label, nous sommes en face d'un problème de classification. Les algorithmes utilisés pour y arriver seront `Decision Tree`, `Random Forest` et `Naive Bayes`. Puisque `Decision Tree` ne prend que des valeurs numériques en entrée, nous allons devoir encoder les variables qualitatives.
# + id="oadDxQMQ1rVB"
df.drop(list(df.columns[0:3]) + ["artist"], axis=1, inplace = True)
# + id="5E7GZPag1rVC" colab={"base_uri": "https://localhost:8080/", "height": 123} outputId="c9b81b50-4bb1-46a6-abdc-21e293390249"
display(df.isnull().sum())
df[df.isnull().any(axis=1)]
# + [markdown] id="llu7af831rVC"
# Nous avons aucune donnée manquante dans tout le tableau. Si on en avait eu, on aurait pu soit la laisser tomber, ou bien utiliser une imputation par la règle, puisqu'il suffit de chercher le titre sur le web pour trouver un attribut associé.
#
# Intéressons-nous maintenant au nombre d'entrées qui seront nécessaires pour le modèle.
# + id="epDpMGcj1rVC" colab={"base_uri": "https://localhost:8080/", "height": 256} outputId="973482ce-ce07-4ef1-d64e-56b5ff52d462"
display(df.value_counts(df['genre']))
# + [markdown] id="YSj3B5Ug1rVD"
# Le nombre de chansons des différents genres varie beaucoup, quelques uns en ont beaucoup plus que d'autres, particulièrement Rock.
#
# Si on entraîne un modèle sur ces données, on risquerait d'avoir un gros biais pour ces genres. On devrait donc rendre le set plus équilibré. En effet, on trouve à l'entrée de *Downsampling* du "Machine Learning Glossary" de *Google*:
#
# ```
# [...]
# For example, in a class-imbalanced dataset, models tend to learn a lot about
# the majority class and not enough about the minority class. Downsampling helps
# balance the amount of training on the majority and minority classes.
# ```
#
# Le jeu de données est trop volumineux pour créer le modèle. On va donc se limiter à environ 1/10 de celui-ci. Nous avons un total d'environ 218 000 entrées, ce qui veut dire qu'on utiliserait environ 21 000 entrées. Si nous voulons avoir une représentation égale de chaque genre alors il faudrait environ 21000/11 genres, donc environ 1900 chanson par genre. `Folks` est la catégorie en ayant le moins avec 1689 donc c'est le maximum que nous allons utiliser pour que les genres soient égalements représentés.
#
# **Note à posteriori**: en raison du temps de calcul important, on a réduit la taille à $800*11 = 8 800$ individus.
# + id="yuWbnqAw1rVD" colab={"base_uri": "https://localhost:8080/"} outputId="c6ff4120-c3cc-4b58-a3fd-8058857dd887"
n_samples = 800
df_sample = df.groupby('genre').sample(n_samples, random_state = 17)
#Test réduire nombre de classe
df_sample = df_sample[df_sample['genre'] != 'Folk']
df_sample = df_sample[df_sample['genre'] != 'Pop']
df_sample = df_sample[df_sample['genre'] != 'Rock']
df_sample = df_sample[df_sample['genre'] != 'Electronic']
print(df_sample.value_counts(df['genre']))
# + [markdown] id="AMrGSwk71rVE"
# # Tranformations du vecteur `lyrics` vers une représentation numérique
# + [markdown] id="Trsg18Yl1rVE"
# ## Partie commune
# Pour tous les algorithmes, il est nécessaire de fragmenter les phrases en liste de mots, de retirer les contractions, d'enlever les `stopwords` (mots qui ne sont pas significatifs, car ils sont trop communs), etc.
# + [markdown] id="VIvVQ37G1rVE"
# La première étape consiste à tout mettre en minuscule afin de standardiser et faciliter la tâche du traitement des mots.
# + id="72fLM2vF1rVE" colab={"base_uri": "https://localhost:8080/", "height": 424} outputId="f56217ec-8ea9-4227-d7e8-c93cd215df39"
strCols = df.select_dtypes(include='object')
df_sample[strCols.columns] = strCols.applymap(lambda string: string.lower())
display(df_sample)
#Pour simplifier la manipulation
target = df_sample["genre"]
# + [markdown] id="GegIPhMxtZ1m"
# Dans le fragment de code suivant, on *tokenize* les mots de chaque chanson, qui a préalablement été "corrigée" pour enlever les contractions.
# + id="3zJflMlL1rVF"
df_sample['tokenized'] = df_sample["lyrics"].apply(lambda l: nltk.word_tokenize(contractions.fix(l)))
# + [markdown] id="Z8rt0Qz_GtXw"
# Avant de poursuivre, nous allons en profiter pour ajouter une variable explicative: le nombre de mots dans une chanson. En effet, peut-être que certains genres de musiques sont généralement plus courts; on peut penser au country qui raconte des histoires, alors que le jazz est davantage orienté vers la mélodie.
# + id="JcGi5RhWH2AB" colab={"base_uri": "https://localhost:8080/", "height": 592} outputId="230b8f6b-2473-4662-ddd0-d52807e48f4f"
df_sample["length"] = df_sample["tokenized"].apply(len)
pivoted = df_sample.pivot_table(columns="genre", values="length", aggfunc="mean")
plt.figure(figsize=(20,10))
plt.bar(x = pivoted.columns, height=pivoted.loc["length"].values)
plt.show()
# + [markdown] id="ddEuwRlnPfuJ"
# Il semble effectivement avoir une différence. En particulier, cette variable est très discriminante pour le style `hip-hop`.
# + [markdown] id="6A6tO1ch1rVF"
# Nous avons maintenant une liste de mots non contractés. Maintenant, nous allons essayer de repérer des mots aberrants comme "aaaaaaaaaaaaaaaaaaaaaaaaaaaah".
# + id="xZ-ZEczM1rVF"
words_lengths = df_sample['tokenized'].apply(lambda array : list(map(len, array))).explode()
freq = words_lengths.value_counts(sort=False)
# + id="e77eYG7o1rVG" colab={"base_uri": "https://localhost:8080/", "height": 663} outputId="1fde01da-f515-4177-92cd-6a184cf55a8b"
plt.figure(figsize=(20,10))
plt.suptitle("Nombre d'occcurrences selon la longueurs des mots")
plt.subplot(1,2,1)
plt.bar(x = freq.index ,height = freq)
plt.subplot(1,2,2)
plt.boxplot(words_lengths)
plt.show()
# + id="W48Qk7KmU7e3" colab={"base_uri": "https://localhost:8080/"} outputId="67fca6f7-120d-4fea-fdf7-8f4c8d36670f"
print("coefficient d'asymétrie:",words_lengths.skew())
# + [markdown] id="rBBOG1PM1rVG"
# Nous avons une distribution légèrement asymétrique à gauche, et c'est probablement lié aux données aberrantes. D'ailleurs, on peut constater sur le boxplot que certains mots ont une longueur avoisinant les 50 lettres!
#
# + id="Vye4AO8tQSjX" colab={"base_uri": "https://localhost:8080/"} outputId="adc99dd7-caa3-41f4-859f-258c4319a4e4"
# inspiré de https://stackoverflow.com/questions/10072744/remove-repeating-characters-from-words
# itertools.groupby(string) retourne une liste de tuples dont le 2e élément est un itérateur contenant les caractères identiques groupés
# Donc si une lettre est présente plus de 2 fois, alors on retient le mot
stretched = df_sample['tokenized'].apply(lambda array : [word for word in array if any((len(list(s))>2 for _, s in itertools.groupby(word)))])
stretched = stretched.explode().value_counts()
print(stretched.head(50),"\n")
for word in stretched.index:
print(word)
# + [markdown] id="ytypbveyXOWD"
# On aurait pu utiliser la librairie spellchecker qui corrige les mots comme "haappyy" (qui est lui-même une réduction de "hhhaaaapppppyyyy" après avoir été réduit avec REGEX tel quel s'il contient plus de 2 lettres identiques consécutives, alors ce nombre de lettres est réduit à 2) en "happy". Par contre, à regarder la liste des mots ayant plus de 3 lettres identiques consécutives (résultat d'éxécution caché), ça semble assez négligeable pour ce que ça apporterait. Et en fait, 95% sont des "noise words", donc nous allons tout simplement les "filter out" avec le reste.
#
# Après ce nettoyage, on devrait avoir une courbe qui s'apparente davantage à la gaussienne. Par contre, même dans ce cas, puisqu'il s'agit de mots, il ne semble pas judicieux de couper tous les mots ayant une longueur telle que son écart-type est supérieur à un certain seuil. En prenant comme hypothèse qu'il s'agit de paroles de chansons et que les mots utilisés sont généralement courts, on pourrait décider, en tenant compte des graphiques obtenus, de fixer un seuil à environ 15 lettres, mais ça reseterait arbitraire, surtout si l'on tient compte de la Loi de Zipf qui stipule que la fréquence d'apparition des mots a une distribution similaire à la fonction $$\frac {c} {x}\rightarrow \left (\frac {\mbox{fréq_mot}_i} {\mbox{total}}\right )$$Si nous mettons cela en lien avec la théorie de l'information de <NAME>, alors il est probable que les mots les plus utilisés soient aussi les plus courts (l'encodage d'un concept sémantique dans un son ou un mot est généralement proportionnel à son poids sémantique; un mot comme "de" est sémantiquement pauvre, mais court en raison de sa fréquence, alors qu'un mot plus spécifique comme "parachutiste" contient plus d'information (racine, affixes, etc.)). De ce fait, les mots les plus intéressants sont probablement les plus longs car ils véhiculent davantage d'information spécifique. En prenant cela en considération, nous allons opter pour un intervalle [3, 20].
# + id="8x0kgW6i1rVG"
stopwords = nltk.corpus.stopwords.words('english')
df_sample['tokenized'] = df_sample['tokenized'].apply(lambda array : [word for word in array if all([word not in stopwords
, len(word) in range(3,20)
, word.isalpha()
, all((len(list(s))<3 for _, s in itertools.groupby(word)))])])
# + [markdown] id="qwy0QwpU1rVI"
# Ici on peut utiliser le *stemming* ou le *lemmatizing*, deux techniques similaires qui permettent de transformer un mot en son 'unité de base' ou sa racine, pour ignorer toutes les formes conjuguées possibles. Ainsi, on réduit le nombre total de tokens différents et on conserve le sens des mots, ce qui devrait aider la classification.
#
# Le *stemming* est entièrement 'algorithmique' et ne tient compte que de l'orthographe donc on peut obtenir des racines qui n'ont pas vraiment de sens, ou regrouper plusieurs mots qui n'ont pas de liens, comme 'universelle' et 'université' en 'univers'. Cependant, cette technique devrait être la plus rapide.
#
# Le *lemmatizing* tient compte des règles linguistiques et du sens même des mots, en ne retournant que des mots qui 'existent' ou sont dans le dictionnaire, donc on ne devrait pas avoir de mots qui n'ont pas de liens sous une même racine, et aucune racine n'ayant pas de sens. Cependant ce processus est plus long et couteux, mais permet plus de précision.
#
# On utilisera donc le *lemmatizing*.
# + id="ImeoH4Uh1rVI" colab={"base_uri": "https://localhost:8080/", "height": 424} outputId="6c98e911-fefa-42c7-9f7d-96ee5bb488ca"
lemmatizer = WordNetLemmatizer()
# inspiré de "https://stackoverflow.com/questions/15586721/wordnet-lemmatization-and-pos-tagging-in-python"
conversion_table = dict(zip(list("JVNR"), [wordnet.ADJ,wordnet.VERB, wordnet.NOUN, wordnet.ADV]))
tagged = nltk.tag.pos_tag_sents(df_sample["tokenized"])
df_sample['lemmatized'] = list(map(lambda sentence: [lemmatizer.lemmatize(word, conversion_table.get(cat[0], wordnet.NOUN)) for word, cat in sentence], tagged))
display(df_sample)
df_sample.drop(columns=['tokenized'], inplace=True)
# + [markdown] id="Arfh4zjgXvA1"
# `WordNetLemmatizer().lemmatize()`prend l'argument `pos` en paramètre. Il est requis pour déterminer à quelle groupe un mot appartient (nom, adjectif, verbe, adverbe). Ainsi, on n'a pas le choix d'utiliser la fonction `pos_tag_sents` qui permet d'associer à chaque mot le groupe auquel il appartient. Cette fonction prend beaucoup de temps à exécuter, par contre, elle reste plus rapide que la fonction `pos_tag`. En effet, dans la documentation de `pos_tag`:
# ```
# Use `pos_tag_sents()` for efficient tagging of more than one sentence.
# ```
#
# **Note**: on utilise seulement 4 catégories, car `pos` n'accepte que les 4 susmentionnées, donc les mots qui ont des catégories en dehors de celles-là sont réassignés à la catégorie `NOUN`.
# + [markdown] id="dftyFJPyx14b"
# Enfin, On retransforme chaque liste de string en un seul string pour appliquer les 'Vectorizer'.
# + id="YqkwBQ6goApR"
def list_to_str(text):
return ' '.join(e for e in text)
df_sample['lemmas'] = df_sample['lemmatized'].apply(list_to_str)
df_sample.drop(columns=['lemmatized'], inplace=True)
# + [markdown] id="Xrj1S4R5n90Y"
# ## Bag of Words (BoW)
#
# Première représentation numérique des paroles. Pour chaque chanson on crée un vecteur dont les composantes sont les occurences dans la chansons de chaque mot du 'vocabulaire' du corpus qui est l'ensemble des chansons.
# + id="B0ZHUas2oA1Q"
#Création du bag of words avec CountVectorizer
count_vect = CountVectorizer(lowercase=False, ngram_range=(1,1))
counts = count_vect.fit_transform(df_sample['lemmas'])
# + [markdown] id="WZHFPjRrm2wx"
# ## Term Frequency (TF)
#
# Cette mesure indique à quel point chaque token (ou mot) est fréquent dans les textes, relativement à la longueur des textes. Si un token particulier représente un pourcentage élevé de tous les tokens d'une chanson, alors celui-ci est peut-être bien "représentatif" de la chanson, ou y est bien spécifique.
# + id="zWIV3kc-YRGi"
#Sommes des colonnes de la matrice counts, compte le nombre total d'occurences
#pour calculer les term frequencies.
sums = np.sum(counts, axis=0)
tfs = scipy.sparse.csr.csr_matrix(counts/sums)
# + [markdown] id="KcEBNR0jY96j"
# Pour une amélioration très importante du temps de fit et prediction du *Random Forest*, on va utiliser une version *sparse* de la matrice des *Term Frequencies*. On passe d'un temps de calcul, pour `.fit()` avec *Random forest*, de 1m30s à 30s.
# + [markdown] id="zZbCxOEtoDV3"
# ## Term Frequency - Inverse Document Frequency (TF-IDF).
#
#
# Cette mesure plus complexe utilise aussi le term frequency calculé précedemment, mais le multiplie ensuite par l' *inverse document frequency*, qui indique dans quelle fraction de tous les documents un terme se trouve.
#
# Ainsi, le tf-idf représente l'importance d'un token contenu dans un document (ici les paroles d'une chanson) relativement au corpus entier.
#
# On suppose que cette mesure devrait être meilleure que la term frequency seule pour la classification.
# + id="PY6ZbPSZaDq4" colab={"base_uri": "https://localhost:8080/"} outputId="005c01f1-7831-46d9-af3a-7e5dce21c7d9"
#Même logique que pour les tfs mais ici le vectorizer fais tous les calculs
tfidf_vectorizer = TfidfVectorizer(lowercase=False,ngram_range=(1,1))
tfidfs = tfidf_vectorizer.fit_transform(df_sample['lemmas'])
print(tfidf_vectorizer)
# + [markdown] id="Y91dFVVDSMNx"
# Par défaut, `norm="l2"`. C'est un traitement important à faire quand on a des features qui sont équivalents. Ainsi, les chanson plus courtes ne sont pas pénalisées vis-à-vis des chansons plus longues; le nombre de mots dans une chanson plus longue est plus grand que celui dans les chansons courtes.
#
# $$\|\vec x\|_2 = \sqrt{x_1^2+x_2^2+ \dots + x_n^2}$$
#
# Donc pour avoir $\|x\|_2 = 1$, il suffit de diviser $\vec x$ par $\|\vec x\|$.
# + [markdown] id="z_q5Wz1id5p7"
# # Préparation des classificateurs
# + [markdown] id="3z9gqP07CzZW"
# Pour *Naive Bayes*, on utilisera `MultinomialNB()` avec *Bag of Words* et *TF-IDF* puisque les vecteurs contiennent des valeurs entières (voir plus bas pour *TF-IDF*) et `GaussianNB()` pour la représentation numériques *Term Frequency* qui contient des valeurs continues.
# + id="NEGZ-8lal2WB"
dt_classifier = tree.DecisionTreeClassifier(random_state=0)
rf_classifier = ensemble.RandomForestClassifier(random_state=0)
gnb_classifier = naive_bayes.GaussianNB()
mnb_classifier = naive_bayes.MultinomialNB(alpha=0.5)
# + [markdown] id="IXEsmVmH0MMb"
# Pour `MultinomialNB`, le paramètre `alpha` est égal à 1.0 par défaut, mais il est suggérer de tester différentes valeurs. Après plusieurs essais, 0.5 semble donner dans les meilleurs résultats.
# + [markdown] id="ZR2VHgTHbjr7"
# # (4ab) Bag of Words
# + [markdown] id="CKpz77ESefdr"
# ## (3c) Séparation en datasets d'entrainement et de test
# + id="vIjPC7-bewxa"
X_train_bow, X_test_bow, y_train_bow, y_test_bow = tts(counts, target, test_size=0.2, random_state = 0)
# + [markdown] id="RbDXeeJwbvc3"
# ## Decision Tree
# + id="dVkmyG2UbjOa" colab={"base_uri": "https://localhost:8080/"} outputId="48e071a3-a420-4705-ea90-306c2b869202"
t1 = time.process_time()
# %memit
dt_classifier.fit(X_train_bow, y_train_bow)
t_bw_dt_fit = (time.process_time()-t1)
t2 = time.process_time()
# %memit
dt_pred_bow = dt_classifier.predict(X_test_bow)
t_bw_dt_pred = (time.process_time()-t2)
bw_dt_acc= dt_classifier.score(X_test_bow, y_test_bow).round(2)
# + [markdown] id="uDEe1PYzPBnE"
# Le *Decision Tree* n'est pas très bon en *NLP*, puisque peu de *labels* partagent les même *features*, ce qui rend difficile de trouver une bonne hiérarchie de noeuds.
# + [markdown] id="nxe8-ftGrTGx"
# ## Random Forest
# + id="FlMmXHmjtCj2" colab={"base_uri": "https://localhost:8080/"} outputId="65e10771-fe74-47b4-8e8c-20891006d42d"
t1 = time.process_time()
# %memit
rf_classifier.fit(X_train_bow, y_train_bow)
t_bw_rf_fit = time.process_time() - t1
t2 = time.process_time()
# %memit
rf_pred_bow = rf_classifier.predict(X_test_bow)
t_bw_rf_pred = time.process_time()-t2
bw_rf_acc = metrics.accuracy_score(y_test_bow, rf_pred_bow).round(3)
# + [markdown] id="ulNLXosThF5q"
# *On* remarque que l'accuracy s'est améliorée, mais l'apprentissage prend plus de temps en raison du fait qu'un Random Forest est une collection de Decision Tree. La mémoire utilisée est comparable à l'algorithme précédent.
# + [markdown] id="YplC1X2Hh301"
# ## Naive Bayes
# + id="OVEWBqNFj0yl"
t1 = time.process_time()
# #%memit
mnb_classifier.fit(X_train_bow, y_train_bow) # .toarray() car `dense numpy array is required` .toarray()
t_bw_nb_fit = time.process_time() -t1
# #%memit
mnb_pred_bow = mnb_classifier.predict(X_test_bow)#ici
t_bw_nb_pred = time.process_time()-(t1+t_bw_nb_fit)
bw_nb_acc = metrics.accuracy_score(y_test_bow, mnb_pred_bow).round(3)
# + colab={"base_uri": "https://localhost:8080/"} id="J8VLSuGjJHB5" outputId="588194de-e99c-4f16-d7e1-be5f76fbac46"
t_bw_nb_fit
# + [markdown] id="1hWFardwP1ZS"
# Meilleure performance obtenue jusqu'à présent. Le seul problème est le fait que l'algorithme utilise beaucoup de ressource (temps de calcul, mémoire).
# + [markdown] id="PsFNeVC2h9EF"
# # (4ab) Term Frequency
# + [markdown] id="yhNDAGLU3z1E"
# Pour la comparaison détaillée du reste des algorithmes, voir le tableau à la fin.
# + [markdown] id="Ktmq7H4UCHjs"
# ## (3c) Séparation en datasets d'entrainement et de test
# + id="EGgLlNtbiK9M"
X_train_tf, X_test_tf,y_train_tf, y_test_tf = tts(tfs, target, test_size =0.2, random_state=0)
# + [markdown] id="2uOlgsoQjOcp"
# ## Decision Tree
# + id="_66ycXW1jQef" colab={"base_uri": "https://localhost:8080/"} outputId="18930d38-0c01-465c-acff-4221946663e6"
t1 = time.process_time()
# %memit
dt_classifier.fit(X_train_tf, y_train_tf)
t_tf_dt_fit = time.process_time()-t1
t2 = time.process_time()
# %memit
dt_pred_tf = dt_classifier.predict(X_test_tf)
t_tf_dt_pred = time.process_time()-t2
tf_dt_acc =dt_classifier.score(X_test_tf, y_test_tf).round(3)
# + [markdown] id="r4sd-DoxiY11"
# ## Random Forest
# + id="nRzilb6BCLYw" colab={"base_uri": "https://localhost:8080/"} outputId="0fbc5845-4662-472f-bf1f-ab9c26c56e7a"
t1 = time.process_time()
# %memit
rf_classifier.fit(X_train_tf, y_train_tf)
t_tf_rf_fit = time.process_time()-t1
# %memit
rf_pred_tf = rf_classifier.predict(X_test_tf)
t_tf_rf_pred = time.process_time()-(t1+t_tf_rf_fit)
tf_rf_acc= metrics.accuracy_score(y_test_tf,rf_pred_tf).round(2)
# + [markdown] id="OD8EdlgUksXt"
# ## Naive Bayes
# + id="x22Q7J2JkuH3" colab={"base_uri": "https://localhost:8080/"} outputId="fb66a013-c429-4a11-8cd8-a57ba8632154"
t1 = time.process_time()
# %memit
gnb_classifier.fit(X_train_tf.toarray(), y_train_tf)
t_tf_nb_fit = time.process_time()-t1
# %memit
gnb_pred_tf = gnb_classifier.predict(X_test_tf.toarray())
t_tf_nb_pred = time.process_time()-(t1+t_tf_nb_fit)
tf_nb_acc = metrics.accuracy_score(y_test_tf, gnb_pred_tf).round(3)
# + [markdown] id="tooiiSeoBPpS"
# # (4ab) Term Frequency - Inverse Document Frequency
# + [markdown] id="q1xspNSb4Qsg"
# Avec cette méthode, les algorithmes utilisent beaucoup plus de mémoire.
# + [markdown] id="bjn9YE9Mixdy"
# ## (3c) Séparation en datasets d'entrainement et de test
# + id="OIDEayl1i2n1"
X_train_tfidf, X_test_tfidf, y_train_tfidf, y_test_tfidf = tts(tfidfs, target.tolist(), test_size =0.2, random_state = 0)
# + [markdown] id="esxRZ_u0kj0V"
# ## Decision Tree
# + id="5G2JmBHTkl_q" colab={"base_uri": "https://localhost:8080/"} outputId="90a09a0a-f282-4b2a-a91a-3dda5dd22d42"
t1 = time.process_time()
# %memit
dt_classifier.fit(X_train_tfidf, y_train_tfidf)
t_idf_dt_fit = time.process_time()-t1
# %memit
dt_pred_tfidf = dt_classifier.predict(X_test_tfidf)
t_idf_dt_pred = time.process_time()-(t1+t_idf_dt_fit)
idf_dt_acc = metrics.accuracy_score(y_test_tfidf,dt_pred_tfidf).round(3)
# + [markdown] id="Bl3o19K_kzMw"
# ## Random Forest
# + id="Vx1oJdGj9H1d" colab={"base_uri": "https://localhost:8080/"} outputId="7e2d7429-2167-420e-b3aa-c8677d9ae3eb"
t1 = time.process_time()
# %memit
rf_classifier.fit(X_train_tfidf, y_train_tfidf)
t_idf_rf_fit = time.process_time()-t1
# %memit
rf_pred_tfidf = rf_classifier.predict(X_test_tfidf)
t_idf_rf_pred = time.process_time()-(t1+t_idf_rf_fit)
idf_rf_acc = metrics.accuracy_score(y_test_tfidf, rf_pred_tfidf).round(3)
# + [markdown] id="68TXOb54k4OG"
# ## Naive Bayes
# + id="D5zupPpsi5BT" colab={"base_uri": "https://localhost:8080/"} outputId="bfc5d4c1-5898-4bbb-89ef-a343e97b9e98"
t1 = time.process_time()
# %memit
mnb_classifier.fit(X_train_tfidf.todense(), y_train_tfidf)
t_idf_nb_fit = time.process_time()-t1
# %memit
gnb_pred_tfidf = mnb_classifier.predict(X_test_tfidf.todense())
t_idf_nb_pred = time.process_time()-(t1+t_idf_nb_fit)
idf_nb_acc = metrics.accuracy_score(y_test_tfidf, gnb_pred_tfidf).round(3)
# + [markdown] id="Nd9GjEIAzVja"
# Techniquement, `GaussianNB` devrait être utilisé puisque le jeu de données contient des valeurs continues, par contre, selon la documentation de `MultinomialNB`:
# ```
# The multinomial Naive Bayes classifier is suitable for classification with
# discrete features (e.g., word counts for text classification). The
# multinomial distribution normally requires integer feature counts. However,
# in practice, fractional counts such as tf-idf may also work.
# ```
#
# Et en effet, le score obtenu est meilleur en utilisant `MultinomialNB`. On passe d'environ $0.24$ (et un temps d'exécution de 6.2s) à $0.323$ en 2.4s.
# + [markdown] id="XAfmsXKgfpOR"
# # Prise en charge des prédicteurs `year` et `length`
# Ces deux paramètres nous semblent intéressants, car il est connu que chaque époque a eu un genre musical surreprésenté en fonction des modes et des tendences. C'est d'ailleurs une donnée qui se vérifie empiriquement. Pour ce qui est de la longueur des paroles, nous en avons déjà parlé un peu plus haut.
#
# Les deux meilleures performance ont été obtenues avec **Naive Bayes** et **Random Forest**. On fera donc 2 modèles:
# - *Random Forest* avec *TF-IDF*;
# - *Naive Bayes* avec *BoW*.
# + [markdown] id="XhOOrFg-Tb3S"
# On doit normaliser pour ne pas qu'il y ait une prédominance de `year`et `length`.
# + id="_clB2vt25hNJ"
scaler = preprocessing.MinMaxScaler()
# + id="RBbN6YrntE_p"
X = tfidfs.toarray()
X = np.array(list(map(lambda r, y, l: np.append(r, [y,l]), X, df_sample["year"], df_sample["length"])))
X_train_tfidf_mod, X_test_tfidf_mod, y_train_tfidf_mod, y_test_tfidf_mod = tts(X, target, test_size =0.2, random_state = 0)
X_train_tfidf_mod[:,X_train_tfidf_mod.shape[1]-2:X_train_tfidf_mod.shape[1]] = scaler.fit_transform(
X_train_tfidf_mod[:,X_train_tfidf_mod.shape[1]-2:X_train_tfidf_mod.shape[1]])
X_test_tfidf_mod[:,X_test_tfidf_mod.shape[1]-2:X_test_tfidf_mod.shape[1]] = scaler.transform(
X_test_tfidf_mod[:,X_test_tfidf_mod.shape[1]-2:X_test_tfidf_mod.shape[1]])
t1 = time.process_time()
# #%memit
rf_classifier.fit(X_train_tfidf_mod, y_train_tfidf_mod)
t_idf_extra_fit = time.process_time()-t1
# #%memit
rf_pred_tfidf_mod = rf_classifier.predict(X_test_tfidf_mod)
t_idf_extra_pred = time.process_time()-(t1+t_idf_extra_fit)
accu_idf_extra = metrics.accuracy_score(y_test_tfidf_mod, rf_pred_tfidf_mod).round(3)
# + colab={"base_uri": "https://localhost:8080/"} id="64PM4BEPGhvz" outputId="fe6fe289-e457-4160-cf85-e02896d04100"
X = counts.toarray()
X = np.array(list(map(lambda r, y, l: np.append(r, [y,l]), X, df_sample["year"], df_sample["length"])),dtype='float64')
X_train_bow_mod, X_test_bow_mod, y_train_bow_mod, y_test_bow_mod = tts(X, target, test_size =0.2, random_state = 0)
X_train_bow_mod[:,X_train_bow_mod.shape[1]-2:X_train_bow_mod.shape[1]] = scaler.fit_transform(
X_train_bow_mod[:,X_train_bow_mod.shape[1]-2:X_train_bow_mod.shape[1]])
X_test_bow_mod[:,X_test_bow_mod.shape[1]-2:X_test_bow_mod.shape[1]] = scaler.transform(
X_test_bow_mod[:,X_test_bow_mod.shape[1]-2:X_test_bow_mod.shape[1]])
t1 = time.process_time()
mnb_classifier.fit(X_train_bow_mod, y_train_bow_mod)
t_bow_extra_fit = time.process_time()-t1
# %memit
mnb_pred_bow_mod = mnb_classifier.predict(X_test_bow_mod)
t_bow_extra_pred = time.process_time()-(t1+t_bow_extra_fit)
accu_bow_extra = metrics.accuracy_score(y_test_bow_mod, mnb_pred_bow_mod).round(3)
# + [markdown] id="x8npAJKgMO9_"
# Ce qui est remarquable, c'est qu'en ajoutant deux prédicteurs, le temps de calcul fond à 6 secondes. C'est un contraste important avec le modèle sans ces 2 variables (1m30s). Du reste, la performance est similaire.
# + [markdown] id="Y3W84ZCyggdF"
# En somme, les scores obtenus sont plutôt bas, même si les deux prédicteurs supplémentaires semblent aider. Toutefois, si l'on tient compte du fait qu'il y a 11 genres, alors une réponse complètement aléatoire devrait donner un score d'environ 9%. Or, dans les simulations que nous avons faites, on a pu atteindre une accuracy frôlant les 40%, ce qui est excellent compte tenu de la tâche. En effet, il semble beaucoup plus complexe de dire à quel genre une chanson appartient en fonction du vocabulaire qu'il contient en comparaison avec d'autre classifications, comme le sport ou la cuisine. C'est que l'ensemble du vocabulaire est beaucoup moins pointu (narrow) que pour catégoriser des recettes, par exemple. Des chansons ayant un champ lexical tournant autour de l'amour existent dans environ tous les styles et c'est d'ailleurs probablement l'un des thèmes les plus récurrent, mais c'est un point qui serait à vérifier. Quoi qu'il en soit, c'est une tâche qui resterait difficile à faire même pour un humain.
#
# Une chose à laquelle nous nous attendions pas est le fait que la taille d'échantillon pour développer les modèles a eu un impact très marginal sur la force du modèle. En effet, le fait de passer d'un jeu de données de $11*150$ entrées à $11*1689$ n'a fait varier l'accuracy favorablement que de 2-3 points de pourcentage. On aurait pu comparer sur un graphique l'accuracy en fonction de la taille de l'échantillon, mais le temps de calcul devient rapidement très long.
#
# Note bonus: nous avons fait l'essai de classifier le genre uniquement selon l'année et le nombre de mots dans la chanson avec *Random Forest* et nous avons obtenu un score de .2, ce qui est mieux que le hasard et ce qui est, en fait, aussi mieux que certains résultats obtenus précédemment!
# + [markdown] id="C0I_hAlDmJPf"
# # (5) Visualisation de la performance avec la matrice de confusion
# + [markdown] id="G3gTl9CSmWYi"
# Pour cette dernière partie, on va utiliser les meilleurs résultats que nous avons obtenus (*Naive Bayes* et *Random Forest* avec les prédicteurs supplémentaires).
# + id="rk__fuZKxQWk" colab={"base_uri": "https://localhost:8080/", "height": 540} outputId="2026c604-2d51-4709-d4b5-34b4ffd2379e"
cf_rf = metrics.confusion_matrix(y_test_tfidf_mod, rf_pred_tfidf_mod, normalize="true")
cf_nb = metrics.confusion_matrix(y_test_bow_mod, mnb_pred_bow_mod, normalize="true")
plt.figure(figsize=(20,8))
plt.suptitle("Matrices de confusion")
plt.subplot(1,2,1)
sns.heatmap(cf_rf.round(1), annot=True, xticklabels=target.unique(), yticklabels=target.unique(), cmap="Blues")
plt.title("Random Forest")
plt.subplot(1,2,2)
sns.heatmap(cf_nb.round(1), annot=True, xticklabels=target.unique(), yticklabels=target.unique(), cmap="Blues")
plt.title("Naive Bayes")
plt.show()
# + [markdown] id="-uOsxawi7myE"
# On voit que ce ne sont pas tout à fait les mêmes genres musicaux qui sont le mieux classés. En effet, *Random Forest* arrive à bien classifier les chansons appartenant à `other`, mais ce n'est pas le cas de *Naive Bayes*. Du reste, on peut dire que les deux genres ayant eu la meilleure classification sont `hip-hop` et `metal`. On avait d'ailleurs vu, dans le diagramme de distribution, que `length` était très discriminant pour le genre `hip-hop`, où le nombre de paroles est beaucoup plus grand, en moyenne, que le reste des genres. Pourtant, les algorithmes performent aussi bien sans cette variable.
# + [markdown] id="ASJGeC7fmPiY"
# Si l'on compare la matrice de confusion de gauche avec celle (ci-bas) obtenue par les chercheurs ayant travailler avec le même jeu de données pour développer un modèle de classification selon les paroles, on constate qu'il y a peu de similarités, mais cela est explicable par le fait qu'ils ont utilisé un algorithme de *deep learning*. D'ailleurs, ils ont pu obtenir une accuracy d'environ 62%, ce qui est pratiquement deux fois mieux que ce qu'on a pu obtenir avec le Random Forest.
#
# Quant aux genres les mieux classifiés avec *Deep Learning*, ce sont `Metal` et `Jazz`. Nous avons donc une catégorie en commun, mais pour les deux autres, elles ont été mal classifiées avec l'autre algorithme respectif.
#
# Enfin, les chansons qui ont été mal classifiées n'ont pas été toutes placées dans une catégorie "fourre-tout", à l'inverse des résultats obtenus avec le modèle de *deep learning*, où l'on peut voir que l'algorithme avait une préférence pour la catégorie `Indie`. En effet, on remarque qu'il y a des zéros pratiquement partout, hormis sur la diagonale principale et sur la dernière colonne, alors que dans le cas de nos modèle, les erreurs sont plus dispersées.
#
# La raison qui explique un taux de classification qui n'est pas de l'ordre du hasard se situe probablement dans le fait que l'algorithme classifie très bien certain genres, mais est médiocre pour le reste.
#
# Bref, on aurait probablement un meilleur taux de classification si l'on utilisait la mélodie plutôt que les paroles comme prédicteur.
# + [markdown] id="bYe3EdOds-Vk"
# 
# + [markdown] id="oyzkWFkF9bqn"
# $\boxed{\color{green}{\mbox{BONUS}}} \rightarrow$ *Just for fun*
# + colab={"base_uri": "https://localhost:8080/"} id="f5RSAZQOiHha" outputId="cf65aa7b-e266-4797-d030-9713084f4388"
from sklearn import svm
SVM = svm.SVC()
SVM.fit(X_train_tfidf,y_train_tfidf)
predictions_SVM = SVM.predict(X_test_tfidf)
print("SVM Accuracy Score -> ",metrics.accuracy_score(predictions_SVM, y_test_tfidf).round(3))
# + [markdown] id="y9yopc8qKHNV"
# # (4d) Run times
# + id="5eYdfWSos2cU"
#temps dexec du DT
times_dt_fit = [t_bw_dt_fit, t_tf_dt_fit, t_idf_dt_fit]
times_dt_pred = [t_bw_dt_pred,t_tf_dt_pred, t_idf_dt_pred]
#random forest
times_rf_fit = [t_bw_rf_fit, t_tf_rf_fit, t_idf_rf_fit]
times_rf_pred = [t_bw_rf_pred,t_tf_rf_pred, t_idf_rf_pred]
#naive bayes
times_nb_fit = [t_bw_nb_fit, t_tf_nb_fit, t_idf_nb_fit]
times_nb_pred = [t_bw_nb_pred,t_tf_nb_pred, t_idf_nb_pred]
#extras
times_extra_fit = [t_idf_extra_fit, t_bow_extra_fit ]
times_extra_pred = [t_idf_extra_pred, t_bow_extra_pred]
#accuracies
accu_dt = [bw_dt_acc, tf_dt_acc, idf_dt_acc]
accu_rf = [bw_rf_acc, tf_rf_acc, idf_rf_acc]
accu_nb = [bw_nb_acc, tf_nb_acc, idf_nb_acc]
accu_extra = [accu_idf_extra, accu_bow_extra]
accu_dt = [100*acc for acc in accu_dt]
accu_rf = [100*acc for acc in accu_rf]
accu_nb = [100*acc for acc in accu_nb]
accu_extra = [100*acc for acc in accu_extra]
# + id="UOpz8XXzOLFf"
algos = ['Decision tree BoW', 'Decision tree TF', 'Decision tree TFIDF',
'Random forest BoW','Random forest TF','Random forest TFIDF',
'Naive Bayes BoW','Naive Bayes TF','Naive Bayes TFIDF','RF+prédicteurs extras', 'Naive Bayes+prédicteurs extras']
times_fit = [round(elem, 3) for elem in times_dt_fit+times_rf_fit+times_nb_fit+times_extra_fit]
times_pred = [round(elem, 3) for elem in times_dt_pred+times_rf_pred+times_nb_pred+times_extra_pred]
accuracies = [round(elem, 3) for elem in accu_dt+accu_rf+accu_nb+accu_extra]
# + [markdown] id="eY4qSZ-K-dXm"
# ### Comparaison des temps d'entraînement et prédiction ainsi qu'accuracy des algorithmes.
# + id="3z01RFOzKIQT" colab={"base_uri": "https://localhost:8080/", "height": 542} outputId="cc70c193-714b-4792-d3cc-d458e873877f"
table = pd.DataFrame(data = np.array([algos,times_fit, times_pred, accuracies]).T).sort_values(by = 3, ascending=False).T
colonnes=['Algorithme','Temps fitting (s)', 'Temps prediction (s)', 'Accuracy (%)']
fig = go.Figure(data = [go.Table(
header=dict(values = colonnes,
align = 'center'),
cells=dict(values = table,
align = 'left')
)])
fig.show()
# + [markdown] id="0YxZB6PT5r-H"
# Comparaison des différents algorithmes utilisés et de leurs temps de fitting et prediction sur 800 échantillons par genre pour un total de 8800.
#
# Les temps de fitting sont ce qui change le plus, l'algorithme le plus lent (random forest sur TF-IDF avec prédicteurs extras) a pris près de 1000 fois plus de temps à s'exécuter que le plus rapide. Le temps de prédiction ne varie pas autant, variant de moins d'une seconde à environ 4 secondes.
#
# En termes d'*accuracy*, le meilleur algorithme a environ 15 points de pourcentage de plus que le pire, soit une augmentation de près de 75%.
#
# Les 'prédicteurs extras' utilisés deux fois sont `year` et `length`. Dans les deux cas, on voit une légère augmentation de l'accuracy par rapport au même algorithme sans ces prédicteurs. Dans le cas du *Naive Bayes*, l'augmentation est minime, soit 0.2 points de pourcentage (moins de 1% relatif) ce qui pourrait être entièrement attribuable au hasard. Le *random forest* gagne 2 points de pourcentage, soit une augmentation relative de 6%.
# + [markdown] id="zqSzVCarBNsR"
# Qu'on tienne compte ou pas de la représentation numérique utilisée pour les données, certaines tendances ressortent du tableau. Par exemple, le *decision tree* a systématiquement une moins bonne accuracy que les deux autres algorithmes. Il était attendu que le *random forest* obtienne de meilleurs résultats puisqu'il s'agit de plusieurs *decision trees*, mais aucune telle hypothèse avait été posée pour le *naive bayes*.
#
# D'ailleurs, le *naive bayes* semble généralement être aussi voire plus accurate que le *random forest* tout en étant beaucoup plus rapide. En effet, avec n=800, *naive bayes* a une accuracy relative environ 7% plus élevée que le *random forest* mais s'exécute en environ 1/30e du temps, une différence énorme.
#
# De plus, le temps d'exécution du *random forest* augmente beaucoup plus rapidement que celui du *naive bayes* lorsque l'on augmente le nombre de samples, sans nécessairement que l'accuracy augmente beaucoup. Ainsi, le *naive* bayes semble réellement le meilleur algorithme dans notre situation.
| projet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# test_fastKNN.m
# <NAME>
# script based on README
# Dataset taken from http://www.jiaaro.com/KNN-for-humans/
# -------------------------------------------------------
# | weight (g) | color | # seeds || Type of fruit |
# |==============|=========|===========||=================|
# | 303 | 3 | 1 || Banana |
# | 370 | 1 | 2 || Apple |
# | 298 | 3 | 1 || Banana |
# | 277 | 3 | 1 || Banana |
# | 377 | 4 | 2 || Apple |
# | 299 | 3 | 1 || Banana |
# | 382 | 1 | 2 || Apple |
# | 374 | 4 | 6 || Apple |
# | 303 | 4 | 1 || Banana |
# | 309 | 3 | 1 || Banana |
# | 359 | 1 | 2 || Apple |
# | 366 | 1 | 4 || Apple |
# | 311 | 3 | 1 || Banana |
# | 302 | 3 | 1 || Banana |
# | 373 | 4 | 4 || Apple |
# | 305 | 3 | 1 || Banana |
# | 371 | 3 | 6 || Apple |
# -------------------------------------------------------
import matplotlib.pyplot as plt
import numpy as np
import numpy.linalg as lin
import matplotlib.patches as mpatches
import scipy as sypy
from scipy import signal
from scipy import io
from scipy.stats import mode
from ipynb.fs.full.fastKNN import getDistance
from ipynb.fs.full.fastKNN import fastKNN
def normalize(x):
norm= (x- min(x))/max((x-min(x)))
return norm
# a simple mappin
fruit=('Banana','Apple')
color=('red', 'orange', 'yellow', 'green', 'blue', 'purple')
training_dataset = np.array([
# weight, color, # seeds, type
[303, 2, 1, 0],
[370, 0, 2, 1],
[298, 2, 1, 0],
[277, 2, 1, 0],
[377, 3, 2, 1],
[299, 2, 1, 0],
[382, 0, 2, 1],
[374, 3, 6, 1],
[303, 3, 1, 0],
[309, 2, 1, 0],
[359, 0, 2, 1],
[366, 0, 4, 1],
[311, 2, 1, 0],
[302, 2, 1, 0],
[373, 3, 4, 1],
[305, 2, 1, 0],
[371, 2, 6, 1]
],dtype=np.float32
)
validation_dataset =np.array([
[301, color.index('green'),1],
[346 ,color.index('yellow'), 4],
[290, color.index('red'), 2 ]
],dtype=np.float32
)
normalize_datasets=1;
[row,col]=np.shape(training_dataset)
if(normalize_datasets):
# normalize = @(x) (x - min(x)) / max((x - min(x))); % reduce by smallest value
for i in range(col-1):
training_dataset[::,i]=normalize(training_dataset[::,i]);
validation_dataset[::,i]=normalize(validation_dataset[::,i]);
[classified_type, k, index]=fastKNN(training_dataset,validation_dataset);
for i in range(0,len(classified_type)):
print(fruit[classified_type[i]])
| problem_sets/Nearest Neighbor/test_fastKNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Day and Night Image Classifier
#
# The day/night image dataset consists of 200 RGB color images in two categories: 100 day images and 100 night images.
#
# We'd like to build a classifier that can accurately label these images as day or night, and that relies on finding distinguishing features between the two types of images!
#
# Note: All images come from the [AMOS dataset](http://cs.uky.edu/~jacobs/datasets/amos/) (Archive of Many Outdoor Scenes).*
# +
import cv2
import helpers
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
# %matplotlib inline
# -
# ### Training and Testing Data
# The 200 day/night images are separated into training and testing datasets.
#
# * 60% of these images are training images, for you to use as you create a classifier.
# * 40% are test images, which will be used to test the accuracy of your classifier.
#
# First, we set some variables to keep track of some where our images are stored:
#
# image_dir_training: the directory where our training image data is stored
# image_dir_test: the directory where our test image data is stored
# Image data directories
image_dir_training = 'day_night_images/training/'
image_dir_test = 'day_night_images/test/'
# ### Load the datasets
#
# These first few lines of code will load the training day/night images and store all of them in a variable, `IMAGE_LIST`. This list contains the images and their associated label ("day" or "night").
#
# For example, the first image-label pair in `IMAGE_LIST` can be accessed by index:
# ``` IMAGE_LIST[0][:]```.
#
# Load training data
IMAGE_LIST = helpers.load_dataset(image_dir_training)
# ### 1. Visualize the input images
# +
# Select an image and its label by list index
image_index = 0
selected_image = IMAGE_LIST[image_index][0]
selected_label = IMAGE_LIST[image_index][1]
print('This image label is', selected_label, 'with shape: ', selected_image.shape)
plt.imshow(selected_image)
# -
# ### 2. Pre-process the Data
#
# After loading in each image, you have to standardize the input and output.
#
# #### Solution code
#
# You are encouraged to try to complete this code on your own, but if you are struggling or want to make sure your code is correct, there i solution code in the `helpers.py` file in this directory. You can look at that python file to see complete `standardize_input` and `encode` function code. For this day and night challenge, you can often jump one notebook ahead to see the solution code for a previous notebook!
#
# ---
# #### Input
#
# It's important to make all your images the same size so that they can be sent through the same pipeline of classification steps! Every input image should be in the same format, of the same size, and so on.
#
# #### TODO: Standardize the input images
#
# * Resize each image to the desired input size: 600x1100px (hxw).
def standardize_input(image):
standard_im = cv2.resize(image, (1100, 600))
return standard_im
# ### TODO: Standardize the output
#
# With each loaded image, you also need to specify the expected output. For this, use binary numerical values 0/1 = night/day.
def encode(label):
num = 0 if label == 'night' else 1
return num
# ### Construct STANDARDIZE_LIST
#
# This function takes in a lisst of image-label pairs and outputs a standardized list of resized images and numerical labels.
def standardize(image_list):
standard_list = []
for item in image_list:
image = item[0]
label = item[1]
standardized_im = standardize_input(image)
binary_label = encode(label)
standard_list.append((standardized_im, binary_label))
return standard_list
STANDARDIZED_LIST = standardize(IMAGE_LIST)
# ### Visualize the standardized data
# +
selected_image = STANDARDIZED_LIST[0][0]
selected_label = STANDARDIZED_LIST[0][1]
# Display image and label about it.
print('This image label is', selected_label, 'with shape: ', selected_image.shape)
plt.imshow(selected_image)
# -
# ### Feature Extraction
#
# Create a feature that represents the brightness in an image. We will be extracting the average brightness using HSV colorspace. Specifically, we'll use the V channel(a measure of brightness), add up the pixel values in the V channel, then divide that sum by the area of the image to get the average Value of the image.
# ### RGB to HSV conversion
#
# Below, a test image is converted from RGB to HSV colorspace and each component is displayed in an image.
# +
# Convert to HSV
hsv = cv2.cvtColor(selected_image, cv2.COLOR_RGB2HSV)
# Print image label
print('Label:', selected_label)
# HSV channels
h = hsv[:, :, 0]
s = hsv[:, :, 1]
v = hsv[:, :, 2]
# Plot the orignal image and three channels
f, (ax1, ax2, ax3, ax4) = plt.subplots(1, 4, figsize=(20, 10))
ax1.set_title('Standardized image')
ax1.imshow(selected_image)
ax2.set_title('H channel')
ax2.imshow(h, cmap='gray')
ax3.set_title('S channel')
ax3.imshow(s, cmap='gray')
ax4.set_title('V channel')
ax4.imshow(v, cmap='gray')
# -
# ### Find the average brightness using the V channel
def avg_brightness(image):
# Convert image to HSV
hsv = cv2.cvtColor(image, cv2.COLOR_RGB2HSV)
# Add up all the pixel values in the V channel
sum_brightness = np.sum(hsv[:, :, 2])
# Calculate the average brightness using the area of image
# and the sum calculated above.
area = 600 * 1100.
avg = sum_brightness / area
return avg
# +
test_im = STANDARDIZED_LIST[100][0]
test_label = STANDARDIZED_LIST[100][1]
avg = avg_brightness(test_im)
print('Avg brightness:', avg)
plt.imshow(test_im)
# -
# ### Build a complete classifier
def estimate_label(image):
avg = avg_brightness(image)
# set the value of a threshold that will separate day and night images
threshold = 100
predicted_label = 0 if avg < threshold else 1
return predicted_label
# ### Testing the classifier
# +
TEST_IMAGE_LIST = helpers.load_dataset(image_dir_test)
STANDARDIZED_TEST_LIST = helpers.standardize(TEST_IMAGE_LIST)
def get_misclassified_images(images):
misclassified_images_labels = []
for image in images:
im = image[0]
true_label = image[1]
predicted_label = estimate_label(im)
if (predicted_label != true_label):
misclassified_images_labels.append((im, predicted_label, true_label))
return misclassified_images_labels
MISCLASSIFIED = get_misclassified_images(STANDARDIZED_TEST_LIST)
accuracy = 1. - float(len(MISCLASSIFIED) / len(STANDARDIZED_TEST_LIST))
print('Accuracy:', accuracy)
# -
# ### Visualize the misclassified images
mis = MISCLASSIFIED[0][0]
print('Label:', MISCLASSIFIED[0][1])
plt.imshow(mis)
| DeepLearning/Udacity/Computer-Vision/Part 1- Introduction to Computer Vision/1_1_Image_Representation/6. Day and Night Image Classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Shivani-pawar26/Machine_learning/blob/main/Lab_5_linear_regression_with_csv.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="SPxhumkoU2dZ"
#
# + id="JNdVTigPR3MK"
# + [markdown] id="y4jDXvybU23q"
# The dataset for this example is available at:
#
# https://drive.google.com/open?id=1mVmGNx6cbfvRHC_DvF12ZL3wGLSHD9f_
# + colab={"base_uri": "https://localhost:8080/"} id="3ndONWeOWBUG" outputId="b99824bf-7e37-42d9-fdf8-16208ed7ddc4"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="mIXLV_bXU7Mo" outputId="248b21b9-8f62-4bf8-85b5-107c3aff6967"
import pandas as pd
dataset = pd.read_csv('/content/drive/MyDrive/Colab Notebooks/petrol_consumption.csv')
dataset
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="A5Z8NeGDXFwf" outputId="3066ea4c-09ba-42cb-eb2a-0cbecb1fbe66"
dataset.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="xztbw-ZnXJDc" outputId="19577538-8c0e-4a78-daaf-319b4d480fac"
dataset.describe()
# + [markdown] id="m-mbZELQXNk2"
# Preparing the Data
# The next step is to divide the data into attributes and labels as we did previously. However, unlike last time, this time around we are going to use column names for creating an attribute set and label. Execute the following script:
# + id="5O9kzesyXM9x"
X = dataset[['Petrol_tax', 'Average_income', 'Paved_Highways',
'Population_Driver_licence(%)']]
y = dataset['Petrol_Consumption']
# + id="JiOLMO3ZXT68"
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)
# + colab={"base_uri": "https://localhost:8080/"} id="WceCMWt_XWt0" outputId="d8178c94-a0e0-4bce-a318-2b0e882bf478"
from sklearn.linear_model import LinearRegression
regressor = LinearRegression()
regressor.fit(X_train, y_train)
# + id="qmhg_SI8X7SB"
y_pred = regressor.predict(X_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 363} id="DYzc90F0X-Pg" outputId="1e6711c0-e186-4e6e-e090-b1436b2c4b14"
df = pd.DataFrame({'Actual': y_test, 'Predicted': y_pred})
df
# + colab={"base_uri": "https://localhost:8080/"} id="0e2aXMF3YBym" outputId="2db85842-e367-4272-8db4-19a252bf8f83"
from sklearn import metrics
import numpy as np
print('Mean Absolute Error:', metrics.mean_absolute_error(y_test, y_pred))
print('Mean Squared Error:', metrics.mean_squared_error(y_test, y_pred))
print('Root Mean Squared Error:', np.sqrt(metrics.mean_squared_error(y_test, y_pred)))
| Lab_5_linear_regression_with_csv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Example script for ysprod
# ### This script demonstrates the usage of ysprod and ysprod181003550 as presented in arxiv:18100.3550
#
# #### Conventions:
#
# There is a conveniton choice for the sign of the complex part of the ringdown frequency that translates into a convention for the sign of the complex part of the inner-product.
#
#
# By default, $\omega_{\ell m n}$, sep_const = positive.leaver(dimensionless_spin,l,m,n=0,...)[0], outputs QNM frequencies with negative valued complex part. This corresponds to a multipolar convention in which there is an explicity minus sign in the definition of the time domain phase: $h_{\ell m n} = A_{\ell m n} \; exp(-im\omega_{\ell m n})$. However, when considering spherical harmonic moments (eg from NR), a common convention is that positive $m$ moments (ie "modes") have positive frequencies. In that context, we are interested in $h_{\ell m n} = A_{\ell m n} \; exp(im\omega*_{\ell m n})$, where $\omega*_{\ell m n}$ is the complex conjugate of $\omega_{\ell m n}$. As a result, it is necessary to consider the complex conjugate of output from ysprod when seeking to convert from spherical to spheroidal data.
#
#
# In short, if the your spherical harmonic mass-quadrupole (\ell,m)=(2,2), has positive frequencies, then you'll want conj( ysprod(...) ).
# ### Import usefuls
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# The Important Things
from positive import *
from nrutils.core.units import *
from matplotlib.pyplot import *
from numpy import *
# ### Define multipole indeces
# +
#
dimensioless_bh_spin = linspace(0,0.99,2e2)
# Define spherical indeces
L,M = 3,2
# Define spheroidal indeces
lmn = (2,2,0)
# -
# ### Calculate numerical inner-products
# Note that the conjugate is taken
# Calculate spherical-spheroidal
sigma = array([ ysprod( j, L, M, lmn ) for j in dimensioless_bh_spin ]).conj()
# ### Evaluate related model
# Note that the conjugate is taken
#
sigma_from_model = ysprod181003550( dimensioless_bh_spin, L, M, lmn ).conj()
# ### Plot for comparison
# +
#
figure( figsize=1.2*figaspect(0.618/2) )
#
subplot(1,2,1)
plot( dimensioless_bh_spin, abs(sigma), lw=4, color='k', alpha=0.2 )
plot( dimensioless_bh_spin, abs(sigma_from_model), ls='-' )
xlabel('$j$')
ylabel(r'$\sigma_{\bar{\ell}\bar{m}\ell m n}$')
#
subplot(1,2,2)
plot( sigma.real, sigma.imag, lw=4, color='k', alpha=0.2 )
plot( sigma_from_model.real, sigma_from_model.imag, ls='-' )
xlabel(r'$\mathrm{Re} \; \sigma_{\bar{\ell}\bar{m}\ell m n}$')
ylabel(r'$\mathrm{Im} \; \sigma_{\bar{\ell}\bar{m}\ell m n}$')
#
tight_layout(w_pad=2)
| examples/ysprod_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # INFO 3402 – Week 04: Tidying and Tables
#
# [<NAME>, Ph.D.](http://brianckeegan.com/)
# [Assistant Professor, Department of Information Science](https://www.colorado.edu/cmci/people/information-science/brian-c-keegan)
# University of Colorado Boulder
#
# Copyright and distributed under an [MIT License](https://opensource.org/licenses/MIT)
# +
import numpy as np
import pandas as pd
idx = pd.IndexSlice
pd.options.display.max_columns = 100
# -
# ## Background
#
# The COVID-19 pandemic caused disruptions throughout society at local, state, national, and global levels. I am curious about how the pandemic impacted transportation traffic in Colorado. I contacted the Airport Statistics office at Denver International and they shared their monthly [passenger traffic reports](https://www.flydenver.com/about/financials/passenger_traffic) from 1994 (when the airport opened) through 2021.
#
# ## EDA checklist
#
# 1. **Formulate your question** → see “Characteristics of a good question”
# 2. **Read in your data** → Is it properly formatted? Perform cleanup activities
# 3. **Check the packaging** → Make sure there are the right number of rows & columns, formats, etc.
# 4. **Look at the top and bottom of data** → Confirm that all observations are there
# 5. **Check the “n”s** → Identify “landmark” values and to check expectations (number of states, etc.)
# 6. **Validate against an external data source** → Right order of magnitude, expected distribution, etc.
# 7. **Make a plot** → Checking and creating expectations about the shape of data and appropriate analyses
# 8. **Try an easy solution** → What is the simplest test for your question?
# ### Formulate your question
#
# How did the COVID-19 shutdowns impact passenger and flight traffic at Denver International Airport?
# ### Read in data
# Use pandas's `read_csv` function to read in "den_passengers.csv"
passenger_df = pd.read_csv('passengers.csv')
passenger_df.head()
passenger_df.tail()
# ### Check the packaging
#
# The data should go from 1995 through late 2021.
#
# A simple but fragile way to do this is sort the "Month" column.
passenger_df['Month'].sort_values()
# Use [`pd.to_datetime`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.to_datetime.html) to convert the "Month" column into valid `datetime` or `Timestamp` objects. Then you can use functions/methods like `.min()` and `.max()`.
# +
passenger_df['Month'] = pd.to_datetime(passenger_df['Month'])
passenger_df['Month'].min(), passenger_df['Month'].max()
# -
# ### Clean up the data
# Make the "Month" column the index using the [`.set_index`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.set_index.html) method.
passenger_df.set_index('Month',inplace=True)
passenger_df = passenger_df.set_index('Month')
# Drop the column for "Cargo".
passenger_df.drop(columns=['Cargo'],inplace = True)
passenger_df.head()
# ### Validate against an external data source
# These statistics are collected and shared by the DIA authority. The U.S. Department of Transportation's [Bureau of Transportation Statistics](https://www.bts.gov/) also publishes [historical passenger statistics by airport](https://www.transtats.bts.gov/Data_Elements.aspx?Data=1) (select "Denver, CO: Denver International" from the drop-down menu).
#
# Compare the monthly passenger numbers from the BTS database to the DIA data. This can just be a manual/eyeball comparison of a few months of data sampled from different years. Are the numbers exactly the same? Are the trends similar? Are the values correlated?
#
# Or let's do the *overkill* version and flex some of our data analysis and cleaning skills. Read in this file, clean it up, join it with `passenger_df`, and measure the correlation.
# +
# Read in file
transstats_df = pd.read_excel(
'transtats_passengers.xlsx',
sheet_name=0,
header=1
)
# Filter out annual total rows
transstats_df = transstats_df[transstats_df['Month'] != "TOTAL"]
# Drop null rows
transstats_df.dropna(subset=['Month'],inplace=True)
# Inspect
transstats_df.tail()
# -
transstats_df.head()
# This next part is advanced and you're not expected to know it for the Weekly Quiz or Assignment, but expect to see it as an Extra Credit opportunity so experiment with it. It's a common enough data-cleaning task and it's better than anything on StackOverflow so I'll wager some of you will come back to it in the future!
#
# We want to combine the Year and Month columns into a column that can then be passed to `pd.to_datetime` to turn into `datetime` or `Timestamp` objects. We use string formatting with put the Year and Month values with some zero-padding.
# +
# Combine the Year and Month columns into a YYYY-MM string
string_dates = transstats_df[['Year','Month']].apply(lambda x:'{Year}-{Month:02d}'.format(**x),axis=1)
# Create a new column with these strings converted to datetime/Timestamps
transstats_df['Date'] = pd.to_datetime(string_dates)
# Drop the original Year and Month columns
transstats_df.drop(columns=['Year','Month'],inplace=True)
# Inspect
transstats_df.tail()
# -
# Join them together (for practice!).
passenger_df.head(2)
transstats_df.head(2)
# +
dia_bts_merge_df = pd.merge(
left = passenger_df,
right = transstats_df,
left_index = True,
right_on = 'Date',
how = 'right'
)
dia_bts_merge_df.tail()
# -
# Calculate the correlation between the monthly totals from each dataset. Not perfect, but more than high enough to trust!
dia_bts_merge_df[['Grand','TOTAL']].corr()#.iloc[0,1]
# ### Make a plot
#
# Visualize the monthly passenger data for each of the airlines. What are some general trends?
passenger_df['Grand'].plot(figsize=(20,10))
# We'll go over more of this in future weeks.
# +
# Make the plot
ax = passenger_df['Grand'].plot()
# Make vertical lines corresponding to major events
ax.axvline(pd.Timestamp('2001-09-01'),color='r',linestyle='--',linewidth=1,zorder=-1)
ax.axvline(pd.Timestamp('2020-03-01'),color='r',linestyle='--',linewidth=1,zorder=-1)
# Annotate with labels
ax.annotate('9/11 attacks',('2001-09-01',6.5e6),c='r')
ax.annotate('COVID-19 pandemic',('2020-03-01',6.5e6),c='r')
# -
# ### Try an easy solution
#
# How much did passenger traffic decrease in March 2020 compared to March 2019?
passenger_df.head(2)
# +
total_march_2019 = passenger_df.loc['2019-03-01','Grand']
total_march_2020 = passenger_df.loc['2020-03-01','Grand']
"{0:.1%}".format((total_march_2020 - total_march_2019)/total_march_2019)
# -
# ## Stacking and unstacking data
#
# If we have a DataFrame and want to convert its columns into indices, we use pandas's `.stack()` ([docs](https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.stack.html), [user guide](https://pandas.pydata.org/pandas-docs/stable/user_guide/reshaping.html#reshaping-by-stacking-and-unstacking)) method.
#
# 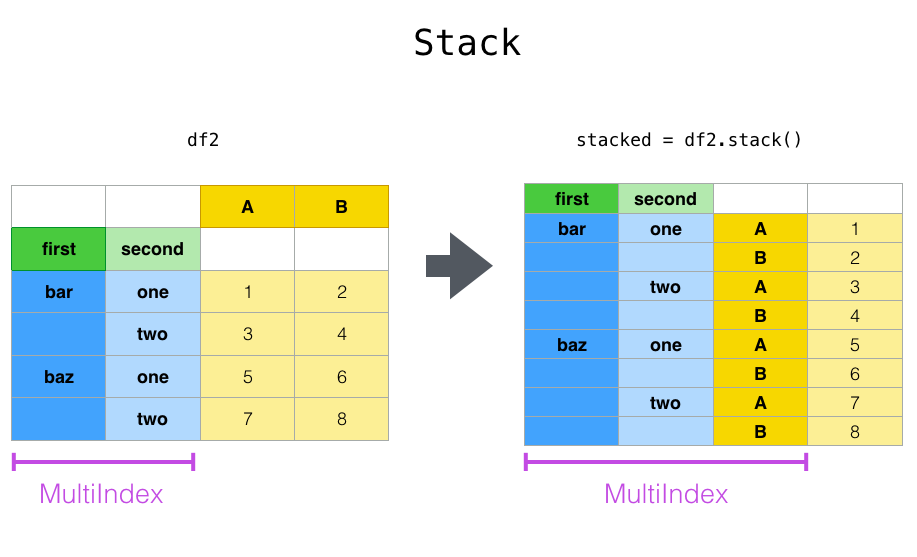
#
# We can reverse a stack or turn indices into columns with pandas's `.unstack()` method ([docs](https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.unstack.html), [user guide](https://pandas.pydata.org/pandas-docs/stable/user_guide/reshaping.html#reshaping-by-stacking-and-unstacking)).
#
# 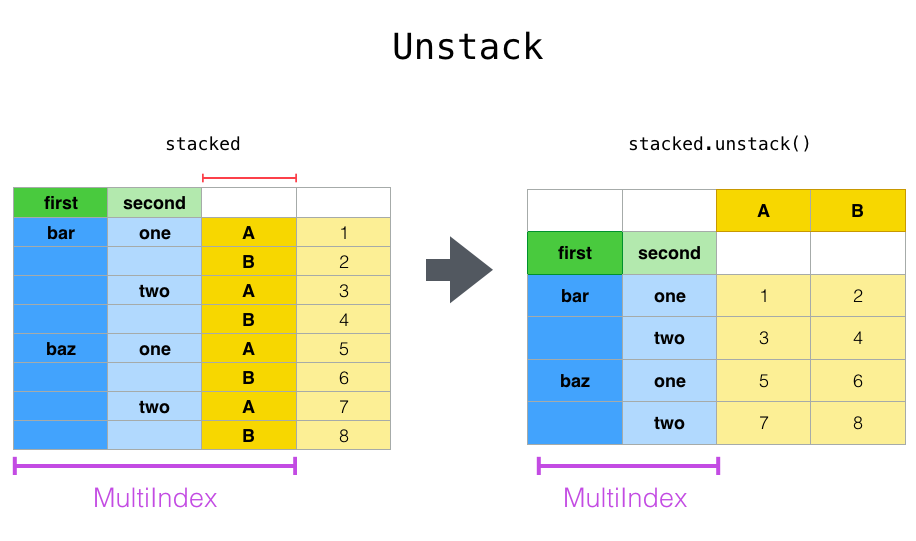
# ### Stacking data
#
# Start by stacking the `passenger_df` DataFrame. This returns a pandas Series with a MultiIndexed index with Month at the first level and the airline at the second level.
passenger_df.head()
# +
passenger_stack_df = passenger_df.stack()
passenger_stack_df.head(10)
# -
# We can turn this MultiIndexed Series into a long DataFrame with `.reset_index()`
# +
passenger_stack_df = passenger_stack_df.reset_index()
passenger_stack_df.head(2)
# -
# The column names after resetting aren't great, so rename them too.
# +
passenger_stack_df.rename(
columns = {
'level_1':'Airline',
0:'Passengers'}
,inplace=True
)
passenger_stack_df.head()
# -
# ### Unstacking data
#
# In the example above, we went from our original "wide" DataFrame, stacked all the columns into the index, reset the index, and renamed the columns to made "long" DataFrame with the same values, just a different shape.
#
# Now we want to reverse this process by setting some columns in our "long" DataFrame as an index and then unstacking the data to return it to a "wide" DataFrame.
#
# Start by creating a MultiIndex by passing a list of column names to the `.set_index()` method
passenger_stack_df.head(2)
# +
passenger_unstack_df = passenger_stack_df.set_index(['Month','Airline'])
passenger_unstack_df
# -
# Once you have the MultiIndex created, you can turn one level into columns by passing the column name (or level position) to the `.unstack` method.
# +
passenger_unstack_df = passenger_unstack_df.unstack('Airline')
passenger_unstack_df.head()
# -
# This first output is a little annoying because it created a MultiIndex on the columns. See the "Passengers" above the airline names? Accessing the Series for "United" requires passing `passenger_unstack_df[('Passengers','United')]`. It would be simpler if that "Passengers" level wasn't there!
#
# One option is to `.drop_level()` ([docs](https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.droplevel.html)) on the columns to get rid of it.
passenger_unstack_df.droplevel(0,axis='columns')
# The other option is to unpivot on a Series rather than a DataFrame. When we `.set_index()` above, we got a DataFrame with a single column back. Which seems similar to a Series, but not the same! We could access that one column's Series and unstack that to get a simpler unstacked DataFrame back.
# +
passenger_stack_s = passenger_stack_df.set_index(['Month','Airline']).loc[:,'Passengers']
passenger_stack_s
# -
# Then unstack this Series and we get a nice simple "wide" DataFrame. Again, you can pass either the name of a MultiIndex level or its position.
passenger_stack_s.unstack(1)
# ## Melting the data
# Use pandas's [`melt`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.melt.html) function on your cleaned `passengers_df` to make it tidier. The primary reason this data is not tidy is that the airline names are in columns even thought these are valuable variables. Note that you'll likely need to use `.reset_index()` on `passengers_df` to turn the "Month" index back into a column. Rename the columns so they're more interpretable too.
#
# Your melted data should look something like this:
#
# | Month | Airline | Count |
# | --- | --- | --- |
# | 1995-02-01 | American | 5833 |
# | 1995-03-01 | American | 228196 |
# | 1995-04-01 | American | 184229 |
# | 1995-05-01 | American | 187990 |
# | 1995-06-01 | American | 229348 |
# Reset the indext to get the "Month" as a column.
passenger_df.head()
passenger_df_reset = passenger_df.reset_index()
print(passenger_df_reset.shape)
passenger_df_reset.head()
# Note the `.shape` I printed out: 321 rows by 9 columns. After we melt, 8 of those 9 rows (the columns corresponding to each airline) should be rows instead. So we should have something like 321 * 8 rows of data.
321 * 8
# Perform the melt keeping the "Month" column as an identifier. The other columns will become values. Notice the shape of the resulting data and compare to our prediction.
passengers_melt_df = pd.melt(
frame = passenger_df_reset,
id_vars = ['Month'],
value_vars = ['American','Delta','Frontier','Other','Southwest','United','United Express','Grand']
)
print(passengers_melt_df.shape)
passengers_melt_df.head()
passengers_melt_df = pd.melt(
frame = passenger_df_reset,
id_vars = ['Month']
)
print(passengers_melt_df.shape)
passengers_melt_df.head()
# Rename the columns.
passengers_melt_df.columns = ['Month','Airline','Count']
passengers_melt_df.head()
# You could also pass a few more arguments to the `melt` function to do the renaming in the same step.
passengers_melt_df = pd.melt(
frame = passenger_df_reset,
id_vars = ['Month'],
var_name = 'Airline',
value_name = 'Count'
)
print(passengers_melt_df.shape)
passengers_melt_df.head()
# ### Unmelting data with a pivot
#
# A melted/"long" DataFrame can be "unmelted" with the `.pivot()` ([docs](https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.pivot.html)) method. Pass the column name(s) you want for the index, columns, and values as parameters.
passengers_melt_df.pivot(index='Month',columns='Airline',values='Count').head()
pd.DataFrame({'Totals':passenger_df.sum()}).style.format(thousands=',',precision=0)
passengers_melt_df.pivot(index='Airline',columns='Month',values='Count')
# ## Exercises
# ### Exercise 01: Filter the melted data (Week 01 review)
#
# Filter the `passengers_melt_df` data down to only "United". Identify the months with the most and fewest passengers.
passenger_df.shape, passengers_melt_df.shape
# +
c0 = passengers_melt_df['Airline'] == 'United'
united_melted_df = passengers_melt_df.loc[c0]
united_melted_df.head()
# -
united_melted_df.loc[united_melted_df['Count'].idxmax(),'Month']
united_melted_df['Count'].idxmin()
united_melted_df.loc[united_melted_df['Count'].idxmin(),'Month']
# ### Exercise 02: Aggregate the total passengers by airline (Week 02 review)
#
# Do a groupby-aggregation or pivot table on `passengers_melt_df` and report on the airlines' total passengers since 1994.
passenger_df.sum()
passengers_melt_df.head(2)
passengers_melt_df.groupby('Airline').agg({'Count':'sum'})
pd.pivot_table(
data = passengers_melt_df,
index = 'Airline',
values = 'Count',
aggfunc = 'sum'
)
# ### Exercise 03: Convert back to wide data
#
# We've practiced turning "wide" data into "tidy" data. Now undo it and turn the tidy data in `passengers_melt_df` back into "wide" data using either a pivot or an unstacking approach. Experiment with making the months into columns!
passengers_melt_df.head(2)
passengers_melt_df.pivot(
index = 'Airline',
columns = 'Month',
values = 'Count'
)
# ### Exercise 04: Load the CDC data and identify the ID and value columns
#
# Think back to the CDC data we used in Week 02. It has a similar patterns of "untidyness" with multiple columns of data that should be rows if you subscribe to the tidy data philosophy. Load the "CDC_deaths_2014_2022.csv" data, print the `.shape`, and inspect the first few rows.
cdc_df = pd.read_csv('../Week 02 - Aggregating and Summarizing/CDC_deaths_2014_2022.csv')
# cdc_df = pd.read_csv('CDC_deaths_2014_2022.csv')
cdc_df.head()
# Write out the column names that should be "id_vars" and/or the column names that should be "value_vars" for `pd.melt`. Alternatively, what columns would you make into an index to stack? Unlike the DIA air passenger data, the CDC data definitely more than one "id_vars" column.
# + active=""
# id_vars = ['State','Year','Week','Week date','All Cause','Natural Cause']
# -
# Based on the data's current shape and the number of "value_vars" columns you identified, approximately how many rows should the melted DataFrame be?
cdc_df.shape
21736*13
# ### Exercise 05: Tidy up the CDC data... then untidy it
#
# Use `melt` or `stack` to reshape the CDC data into a tidy format. Print the `.shape`. Inspect the the first few rows. How does this match with your expectations and predictions?
# +
cdc_melt_df = pd.melt(
frame = cdc_df,
id_vars = ['State','Year','Week','Week date','All Cause','Natural Cause'],
var_name = "Disease",
value_name = "Count"
)
print(cdc_melt_df.shape)
cdc_melt_df.head()
# +
id_cols = ['State','Year','Week','Week date','All Cause','Natural Cause']
cdc_stacked = cdc_df.set_index(id_cols).stack().reset_index()
cdc_stacked.rename(columns={'level_6':'Disease',0:'Count'},inplace=True)
print(cdc_stacked.shape)
cdc_stacked.head()
# -
# Perform a pivot or unstack on the CDC data you just melted to turn it back into a "wide" format.
cdc_stacked.set_index(id_cols+['Disease'])['Count'].unstack('Disease').reset_index()
# ## Appendix
# Clean up the "Passenger" sheet.
# +
# Read in "Passenger" sheet from Excel
# The first two rows are headers
# Make the first column the index
passenger_df = pd.read_excel(
io = 'dia_passenger_reports.xlsx',
sheet_name='Passenger',
header = [0,1],
index_col = 0
)
# Name the index
passenger_df.index.name = 'Month'
# Select the column names that have "Total" in them
total_cols = [col for col in passenger_df.columns if 'Total' in col[0]]
# Select all rows and only the Total columns
passenger_df = passenger_df.loc[:,total_cols]
# Drop the second level of headers
passenger_df = passenger_df.droplevel(1,axis=1)
# Remove "Total" from column names
passenger_df.columns = [col.split(' Total')[0] for col in passenger_df.columns]
# Remove the "Grand Total" row
passenger_df.drop('Grand Total',inplace=True)
# Write to CSV
passenger_df.to_csv('passengers.csv')
# Inspect
passenger_df.tail()
# -
# Clean up the "Flight" sheet
# +
# Read in "Flight" sheet from Excel
# The first two rows are headers
# Make the first column the index
flight_df = pd.read_excel(
io = 'dia_passenger_reports.xlsx',
sheet_name='Flight',
header = [0,1],
index_col = 0
)
# Name the index
flight_df.index.name = 'Month'
# Select the column names that have "Total" in them
total_cols = [col for col in flight_df.columns if 'Total' in col[0]]
# Select all rows and only the Total columns
flight_df = flight_df.loc[:,total_cols]
# Drop the second level of headers
flight_df = flight_df.droplevel(1,axis=1)
# Remove "Total" from column names
flight_df.columns = [col.split(' Total')[0] for col in flight_df.columns]
# Remove "Grand" column
flight_df.drop(columns=['Grand'],inplace=True)
# Remove the "Grand Total" row
flight_df.drop('Grand Total',inplace=True)
# Write to CSV
flight_df.to_csv('flights.csv')
# Inspect
flight_df.tail()
| Week 04 - Tidying and Tables/Week 04 - Lecture.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py3
# language: python
# name: py3
# ---
# +
import os
import numpy as np
import math
import matplotlib.pyplot as plt
import sentencepiece as spm
import torch
import torch.nn as nn
import torch.nn.functional as F
# -
import torch
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
vocab_file = '/home/henry/Documents/wrapper/transformer-evolution/transformer/kowiki.model'
vocab = spm.SentencePieceProcessor()
vocab.load(vocab_file)
# # 1. vocab
# - Word Piece Model
# - 단어와 subword unit으로 나눔
# - 단어의 시작을 특수문자; subword unit은 space로 나눔;
# +
# data_dir = '/home/henry/Documents/wrapper/transformer-evolution/ratings_train.txt'
# lines = []; outputs = [];
# with open(data_dir, "r") as f:
# for idx, line in enumerate(f):
# if idx == 0:
# continue
# if idx == 4:
# break
# temp = line.split()
# lines.append(' '.join(temp[1:-1]))
# outputs.append(temp[-1])
# -
lines = [
"겨울은 추워요.",
"감기 조심하세요."
]
inputs = []
for line in lines:
pieces = vocab.encode_as_pieces(line)
ids = vocab.encode_as_ids(line)
inputs.append(torch.tensor(ids))
print(pieces)
print(inputs)
inputs = torch.nn.utils.rnn.pad_sequence(inputs, batch_first=True, padding_value=0)
print(inputs, inputs.size())
# # 2. embedding
#
# ### 2.1. word embedding
# +
n_vocab = len(vocab)
d_hidn = 128
nn_emb = nn.Embedding(num_embeddings=n_vocab, embedding_dim=d_hidn)
# torch.Size([2, 8, 128])
input_embs = nn_emb(inputs) # 임베딩 레이어에 input
# -
# ### 2.2. position embedding
# - $PE(pos, 2i) = sin(\frac{pos}{1E+4^{2i / d_{model}}}),$
# - $PE(pos, 2i+1) = cos(\frac{pos}{1E+4^{2i / d_{model}}}),$
# - i: embedding vector에서 idx
# - pos: sentence에서 position
def get_sinusoid_encoding_table(n_seq, d_hidn):
"""
n_seq: seq_len
d_hidn: sinusoid table dim
"""
def _cal_angle(pos, i_hidn):
return pos / np.power(10000, 2 * (i_hidn // 2) / d_hidn)
def _get_posi_angle_vec(pos):
"""
sentence의 pos 하나(즉 단어 하나)에 대해 embedding; feature1, feature2, ...., feature128
즉, sequence의 각 position을 128차원으로 position embedding
예컨대, 첫번째 자리 pos embedding은 정해져 있으므로, 첫번째 자리에 어떤 단어가 들어오든,
해당 pos embedding이 word embedding에 더해짐
"""
return [_cal_angle(pos, i_hidn) for i_hidn in range(d_hidn)]
# seq_pos: Word의 sentence 내의 위치 = i
sinusoid_table = np.array([_get_posi_angle_vec(seq_pos) for seq_pos in range(n_seq)])
sinusoid_table[:, 0::2] = np.sin(sinusoid_table[:, 0::2])
sinusoid_table[:, 1::2] = np.cos(sinusoid_table[:, 1::2])
return sinusoid_table
# 64개 단어가 하나의 sentence; 128차원 posion임베딩;
n_seq = 64
pos_encoding = get_sinusoid_encoding_table(n_seq, d_hidn)
print(pos_encoding.shape)
# row: sentence에서 위치(pos)
# %matplotlib inline
plt.figure(figsize=(15,10))
plt.pcolormesh(pos_encoding, cmap='twilight_shifted')
plt.xlabel('embd_feature')
plt.xlim([0, d_hidn])
plt.ylabel('word_pos')
plt.colorbar()
plt.show()
# ### 2.3. pos embd + word embd
# - 의문: pos embd도 train 필요? Nope
# - batch_size & seq_len만큼 position mat 생성
# - position mat에서 padding 위치의 value는 0처리
# +
pos_encoding = torch.FloatTensor(pos_encoding)
nn_pos = nn.Embedding.from_pretrained(pos_encoding, freeze=True) # layer; lookup table
# >> Embedding(64, 128)
position = torch.arange(inputs.size(1), device=inputs.device, dtype=inputs.dtype)
# >> tensor([0, 1, 2, 3, 4, 5, 6, 7])
position = position.expand(inputs.size())
# >> tensor([[0, 1, 2, 3, 4, 5, 6, 7],
# [0, 1, 2, 3, 4, 5, 6, 7]])
position = position.contiguous() + 1 # pad=0이므로 1을 더해 pos에 0없앰
# >> tensor([[1, 2, 3, 4, 5, 6, 7, 8],
# [1, 2, 3, 4, 5, 6, 7, 8]])
pos_mask = inputs.eq(0) # find elems equal to 0 = pad
# tensor([[False, False, False, False, False, False, True, True],
# [False, False, False, False, False, False, False, False]])
position.masked_fill_(pos_mask, 0) # pad는 position 정보도 주지 않음 <- idx 0의 weight를 동일하게 부여
# tensor([[1, 2, 3, 4, 5, 6, 0, 0],
# [1, 2, 3, 4, 5, 6, 7, 8]])
pos_embs = nn_pos(position)
# pos_embs.shape
# >> torch.Size([2, 8, 128]) # batch_size, sequence_len, emb_dim
# +
print(inputs) # embeddimg idx coresponding to the word
print(position)
print(pos_embs.size())
input_vector = input_embs + pos_embs
print(input_vector.size())
# -
# # 3. Scale Dot Product Attention
# $$
# Attention(Q, K, V) = softmax_k(\dfrac{QK^T}{\sqrt{d_k}})V
# $$
# - 왜 scale?
# - d_k는 key vector의 dim.
# - depth가 깊다면, $QK^T$ 값이 커짐
# > "The dot-product attention is scaled by a factor of square root of the depth. This is done because for large values of depth, the dot product grows large in magnitude pushing the softmax function where it has small gradients resulting in a very hard softmax."
# (https://www.tensorflow.org/tutorials/text/transformer)
#
# - detail: 내적은 두 벡터의 크기 * 둘 간의 cosin angle
# - $cos \theta = \dfrac{A \dot B}{|A||B|}$
# - key dim으로 정규화하면 값이 너무 커지는 것을 방지할 수 있음(https://physics.stackexchange.com/questions/252086/dot-product-approaches-zero-as-the-magnitude-of-the-vectors-increase)
# - key, query, value?
# - 기존 attention 모델은, decoder time step t의 hidden state와 encoder의 모든 time step의 hidden state간의 score를 산출.
# - 그리고 그 score를 다시 encoder의 hidden state에 곱하여 weight를 주고, 합산하여 context vector를 산출.
# - 위 context vector는, time step t의 hidden state와 concat.
# - concat된 vector는 feedforward 통과하여 time step t의 출력값
# - 이를 transformer와 비교해보자면, score은 **value**(but value vector 따로 있음), decoder의 hidden state는 **query**, encoder의 hidden state는 **key**에 해당.
#
# > "Traditionally, the attention weights were the relevance of the encoder hidden states (values) in processing the decoder state (query) and were calculated based on the encoder hidden states (keys) and the decoder hidden state (query)."(https://mlexplained.com/2017/12/29/attention-is-all-you-need-explained/)
# - Encoder attention layer는 self attention이므로, Key, Value vector가 모두 자기 자신
Q = input_vector
K = input_vector
V = input_vector
# - attn_mask: key padding indice 마스킹
attn_mask = inputs.eq(0).unsqueeze(1).expand(Q.size(0), Q.size(1), Q.size(1))
# >> torch.Size([2, 8, 8])
# inputs.eq(0) <- 0과 동일한 indice
# inputs.eq(0).unsqueeze(1) <- dim에 1차원 추가
# # >> torch.Size([2, 1, 8])
# - Q.k^T
scores = torch.matmul(Q, K.transpose(-1, -2))
print(scores.size())
print(scores[0])
# ### 3.1. scale
# - $1/\sqrt{d_k}$
# - d_head? k, q, v vector에 weight mat을 곱한 뒤의 dim
d_head = 64
scores = scores.mul(1/d_head**0.5)
# ### 3.2 mask
# - score matrix
# - col: key
# - row: query
scores.masked_fill_(attn_mask, -1e9)
# mask padded area
# torch.Size([2, 8, 8])
# ### 3.3. softmax
# - https://stackoverflow.com/questions/49036993/pytorch-softmax-what-dimension-to-use
# - https://blog.csdn.net/qq_36097393/article/details/89319643
# - dim=-1이면 가장 높은 차원 기준 softmax
attn_prob = nn.Softmax(dim=-1)(scores) # dim 유지
print(attn_prob.size())
# ### 3.4. attn_prov * V
context = torch.matmul(attn_prob, V)
print(context.shape)
# # 4. Multi-Head attn
# - d_hidn = embedding layer 아웃풋의 dim
# - embedding dim
# - d_head = Q, K, V의 dim
# - embedding된 input값이
# - d_hidn > d_head: 연산량 감소하기 위함 eg. 128 > 64
Q = input_vector
K = input_vector
V = input_vector
Q.shape
# - input을 multi head로 복제
# +
batch_size = Q.size(0)
d_hidn = 128 # embedding dim
d_head = 64 # K, Q, V의 dim
n_head = 2
# linear feed forward layer
W_Q = nn.Linear(d_hidn, n_head * d_head)
W_K = nn.Linear(d_hidn, n_head * d_head)
W_V = nn.Linear(d_hidn, n_head * d_head)
# -
# ### 4.1. multi-head K, Q, V input
# bs, n_seq, n_head, d_head
# torch.Size([2, 8, 2, 64])
W_Q(Q).view(batch_size, -1, n_head, d_head).shape
# torch.Size([2, 2, 8, 64])
W_Q(Q).view(batch_size, -1, n_head, d_head).transpose(1,2).shape
# (bs, n_head, n_seq, d_head)
Qs = W_Q(Q).view(batch_size, -1, n_head, d_head).transpose(1, 2)
Ks = W_K(K).view(batch_size, -1, n_head, d_head).transpose(1, 2)
Vs = W_V(V).view(batch_size, -1, n_head, d_head).transpose(1, 2)
Vs.shape
# ### 4.2. multi-head attn mask
# torch.Size([2, 8, 8]) -> torch.size([2, 2, 8, 8])
attn_mask = attn_mask.unsqueeze(1).repeat(1, n_head, 1, 1)
# +
# torch.Size([2, 2, 8, 64]) * torch.Size([2, 2, 64, 8])
# output: torch.Size([2, 2, 8, 8])
scores = torch.matmul(Qs, Ks.transpose(-1, -2))
scores = scores.masked_fill(attn_mask, -np.inf)
# bs, n_head, [n_q_seq, n_k_seq]
# torch.Size([2, 2, 8, 8])
attn_prob = nn.Softmax(dim=-1)(scores)
# bs, n_head, n_q_seq, d_v
# torch.Size([2, 2, 8, 64])
context = torch.matmul(attn_prob, Vs)
# -
# ### 4.3. concat
# transpose: bs, n_q_seq, n_head, d_v
# view: bs, n_q_seq, n_head * d_v
# -> torch.Size([2, 8, 128])
context = context\
.transpose(1, 2)\
.contiguous()\
.view(batch_size, -1, n_head * d_head)
# # 5. output
# torch.Size([2, 8, 128])
# bs, [n_q_seq, n_hidn] <- embedding된 input size와 동일
enc_output = nn.Linear(n_head * d_head, d_hidn)(context)
# # 6. feedforward
# ### 6.1. 1st linear feedforward
conv1 = nn.Conv1d(in_channels=d_hidn, \
out_channels=d_hidn * 4, kernel_size=1)
# +
# bs, [n_hidn, n_q_seq]
# torch.Size([2, 128, 8])
enc_output.transpose(1,2).shape
# torch.Size([2, 512, 8])
enc_output = conv1(enc_output.transpose(1,2))
# -
# ### 6.2. activation
enc_output = F.gelu(enc_output)
# ### 6.3. 2nd linear feedforward
conv2 = nn.Conv1d(in_channels=d_hidn * 4, \
out_channels=d_hidn, \
kernel_size=1)
# torch.Size([2, 128, 8])
enc_output = conv2(enc_output)
# ---
| Transformer/transformer_implement.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Get the challenge:
from aocd import get_data
d = (get_data(day=2)).split('\n')
# +
import regex
count = 0
for p in d:
m = regex.match("(\d+)-(\d+) (\w+): (\w+)",p)
# m.group(1) is lower limit, 2 is upper limit, 3 is character, and 4 is password
if int(m.group(1)) <= m.group(4).count(m.group(3)) <= int(m.group(2)): count = count+1
print('Answer to Part 1: ' + str(count))
# +
count = 0
for p in d:
m = regex.match("(\d+)-(\d+) (\w+): (\w+)",p)
if bool(m.group(4)[int(m.group(1))-1] == m.group(3)) ^ bool(m.group(4)[int(m.group(2))-1] == m.group(3)) :
count = count+1
print('Answer to Part 2: ' + str(count))
| AOC 2020 - Day 2.ipynb |
# +
"""
Compares L1, L2, allSubsets, and OLS linear regression on the prostate data set
Author : <NAME> (@karalleyna)
Based on https://github.com/probml/pmtk3/blob/master/demos/prostateComparison.m
Sourced from https://github.com/empathy87/The-Elements-of-Statistical-Learning-Python-Notebooks/blob/master/examples/Prostate%20Cancer.ipynb
"""
try:
import probml_utils as pml
except ModuleNotFoundError:
# %pip install git+https://github.com/probml/probml-utils.git
import probml_utils as pml
import numpy as np
import matplotlib.pyplot as plt
try:
import pandas as pd
except ModuleNotFoundError:
# %pip install pandas
import pandas as pd
from itertools import combinations
try:
from sklearn.linear_model import Ridge
except ModuleNotFoundError:
# %pip install scikit-learn
from sklearn.linear_model import Ridge
from sklearn.linear_model import Lasso
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import GridSearchCV, KFold
from sklearn.metrics import mean_squared_error
from sklearn.preprocessing import StandardScaler
def get_features_and_label(dataset, is_training):
"""
Gets matrices representing features and target from the original data set
Parameters
----------
dataset: DataFrame
Original dataset
is_training : str
Label to show whether a data point is in training data or not.
* "T" -> Training data
* "F" -> Test data
Return
------
X : ndarray
Feature matrix
y : ndarray
Lpsa values of each data point.
"""
X = dataset.loc[dataset.train == is_training].drop("train", axis=1)
y = X.pop("lpsa").values
X = X.to_numpy()
return X, y
class OneStandardErrorRuleModel:
"""
Select the least complex model among one standard error of the best.
Attributes
----------
estimator :
A regression model to be parametrized.
params : dict
* Keys : The parameter of the model to be chosen by cross-validation.
* Values : The values for the parameter to be tried.
cv : Int
The number of folds for cross-validation.
"""
def __init__(self, estimator, params, cv=10):
self.estimator = estimator
self.cv = cv
self.params = params
self.random_state = 69438 # Seed of the pseudo random number generator
def fit(self, X, y):
grid_search = GridSearchCV(
self.estimator,
self.params,
cv=KFold(self.cv, shuffle=True, random_state=self.random_state),
scoring="neg_mean_squared_error",
return_train_score=True,
)
grid_search = grid_search.fit(X, y)
# Gets best estimator according to one standard error rule model
model_idx = self._get_best_estimator(grid_search.cv_results_)
self.refit(X, y, model_idx)
return self
def _get_best_estimator(self, cv_results):
cv_mean_errors = -cv_results["mean_test_score"] # Mean errors
cv_errors = -np.vstack([cv_results[f"split{i}_test_score"] for i in range(self.cv)]).T
cv_mean_errors_std = np.std(cv_errors, ddof=1, axis=1) / np.sqrt(self.cv) # Standard errors
# Finds smallest mean and standard error
cv_min_error, cv_min_error_std = self._get_cv_min_error(cv_mean_errors, cv_mean_errors_std)
error_threshold = cv_min_error + cv_min_error_std
# Finds the least complex model within one standard error of the best
model_idx = np.argmax(cv_mean_errors < error_threshold)
cv_mean_error_ = cv_mean_errors[model_idx]
cv_mean_errors_std_ = cv_mean_errors_std[model_idx]
return model_idx
def _get_cv_min_error(self, cv_mean_errors, cv_mean_errors_std):
# Gets the index of the model with minimum mean error
best_model_idx = np.argmin(cv_mean_errors)
cv_min_error = cv_mean_errors[best_model_idx]
cv_min_error_std = cv_mean_errors_std[best_model_idx]
return cv_min_error, cv_min_error_std
def refit(self, X, y, model_idx):
if self.params:
param_name = list(self.params.keys())[0]
self.estimator.set_params(**{param_name: self.params[param_name][model_idx]})
# Fits the selected model
self.estimator.fit(X, y)
def get_test_scores(self, y_test, y_pred):
y_test, y_pred = y_test.reshape((1, -1)), y_pred.reshape((1, -1))
errors = (y_test - y_pred) ** 2 # Least sqaure errors
error = np.mean(errors) # Mean least sqaure errors
error_std = np.std(errors, ddof=1) / np.sqrt(y_test.size) # Standard errors
return error, error_std
class BestSubsetRegression(LinearRegression):
"""
Linear regression based on the best features subset of fixed size.
Attributes
----------
subset_size : Int
The number of features in the subset.
"""
def __init__(self, subset_size=1):
LinearRegression.__init__(self)
self.subset_size = subset_size
def fit(self, X, y):
best_combination, best_mse = None, np.inf
best_intercept_, best_coef_ = None, None
# Tries all combinations of subset_size
for combination in combinations(range(X.shape[1]), self.subset_size):
X_subset = X[:, combination]
LinearRegression.fit(self, X_subset, y)
mse = mean_squared_error(y, self.predict(X_subset))
# Updates the best combination if it gives better result than the current best
if best_mse > mse:
best_combination, best_mse = combination, mse
best_intercept_, best_coef_ = self.intercept_, self.coef_
LinearRegression.fit(self, X, y)
# Sets intercept and parameters
self.intercept_ = best_intercept_
self.coef_[:] = 0
self.coef_[list(best_combination)] = best_coef_
return self
path = "https://raw.githubusercontent.com/probml/probml-data/main/data/prostate/prostate.csv"
X = pd.read_csv(path, sep="\t").iloc[:, 1:]
X_train, y_train = get_features_and_label(X, "T")
X_test, y_test = get_features_and_label(X, "F")
# Standardizes training and test data
scaler = StandardScaler().fit(X.loc[:, "lcavol":"pgg45"])
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
n_models, cv, n_alphas = 4, 10, 30
_, n_features = X_train.shape
alpha_lasso, alpha_ridge = [0.680, 0.380, 0.209, 0.100, 0.044, 0.027, 0.012, 0.001], [
436,
165,
82,
44,
27,
12,
4,
1e-05,
]
linear_regression = OneStandardErrorRuleModel(LinearRegression(), {}).fit(X_train, y_train)
bs_regression = OneStandardErrorRuleModel(BestSubsetRegression(), {"subset_size": list(range(1, 9))}).fit(
X_train, y_train
)
ridge_regression = OneStandardErrorRuleModel(Ridge(), {"alpha": alpha_ridge}).fit(X_train, y_train)
lasso_regression = OneStandardErrorRuleModel(Lasso(), {"alpha": alpha_lasso}).fit(X_train, y_train)
regressions = [linear_regression, bs_regression, ridge_regression, lasso_regression]
residuals = np.zeros((X_test.shape[0], n_models)) # (num of test data) x num of models
table = np.zeros(
(n_features + 3, n_models)
) # (num of features + 1(mean error) + 1(std error) + 1(bias coef)) x num of models
for i in range(n_models):
table[:, i] = regressions[i].estimator.intercept_ # bias
table[1 : n_features + 1, i] = regressions[i].estimator.coef_
y_pred = regressions[i].estimator.predict(X_test)
table[n_features + 1 :, i] = np.r_[regressions[i].get_test_scores(y_test, y_pred)]
residuals[:, i] = np.abs(y_test - y_pred)
xlabels = ["Term", "LS", "Best Subset", "Ridge", "Lasso"] # column headers
row_labels = np.r_[
[["Intercept"]], X.columns[:-2].to_numpy().reshape(-1, 1), [["Test Error"], ["Std Error"]]
] # row headers
row_values = np.c_[row_labels, np.round(table, 3)]
fig = plt.figure(figsize=(10, 4))
ax = plt.gca()
fig.patch.set_visible(False)
ax.axis("off")
ax.axis("tight")
table = ax.table(cellText=row_values, colLabels=xlabels, loc="center", cellLoc="center")
table.set_fontsize(20)
table.scale(1.5, 1.5)
fig.tight_layout()
pml.savefig("prostate-subsets-coef.pdf")
plt.show()
plt.figure()
plt.boxplot(residuals)
plt.xticks(np.arange(n_models) + 1, xlabels[1:])
pml.savefig("prostate-subsets-CV.pdf")
plt.show()
| notebooks/book1/11/prostate_comparison.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
from transformers import pipeline
import time
# EleutherAI/gpt-neo-1.3B
start = time.time()
generator = pipeline('text-generation', model='EleutherAI/gpt-neo-1.3B')
end = time.time()
print(end - start)
start = time.time()
result = generator("The meaning of life ", max_length=100, do_sample=True, temperature=0.9)
end = time.time()
print(end - start)
print(result[0]['generated_text'])
| part4/python/.ipynb_checkpoints/GPT-3-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# #!pip install natsort
# -
import cv2
import pytesseract
# %matplotlib inline
from os.path import join,exists, basename, dirname, splitext
from glob import glob
from natsort import natsorted
import cv2
import matplotlib.pyplot as plt
import json
import numpy as np
import pandas as pd
# +
import os
fn = "f1.5-VIDEO_2018-03-1609.33.31__ProjectPetrobras-SP12__Dive702_C1"
ext= ".png"
csv_name = fn+".csv" # MUDAR AQUI
# csv_name = "f1.3-.csv" # MUDAR AQUI
img_name = fn + ext
# img_name = "f1.3-.png"
PATH_FOLDER_CSVS = r"\\ica-094\share\nilton\formatos" # MUDAR AQUI
# PATH_FOLDER_CSVS = r"\\ica-095\tables\manntis\formatos" # MUDAR AQUI
path_csv = os.path.join(PATH_FOLDER_CSVS , csv_name)
print(path_csv)
# -
# %matplotlib inline
def cut_images(path, xmin, ymin, xmax, ymax, label):
im = cv2.imread(path)
im = cv2.cvtColor(im, cv2.COLOR_BGR2RGB)
im_crop = im[ymin:ymin+np.abs(ymax - ymin), xmin:xmin+np.abs(xmax - xmin)]
return im_crop
#fig = plt.figure(figsize=(5, 3))
#fig.suptitle("{} {} {} {} {} :".format(xmin, ymin, xmax, ymax, label), fontsize=20)
#plt.imshow(im_crop)
pytesseract.pytesseract.tesseract_cmd = r'C:\Program Files\Tesseract-OCR\tesseract.exe'
# %matplotlib inline
def plot_images(path, xmin, ymin, xmax, ymax, label):
im = cv2.imread(path)
im = cv2.cvtColor(im, cv2.COLOR_BGR2RGB)
im_crop = im[ymin:ymin+np.abs(ymax - ymin), xmin:xmin+np.abs(xmax - xmin)]
fig = plt.figure(figsize=(5, 3))
fig.suptitle("{} {} {} {} {} :".format(xmin, ymin, xmax, ymax, label), fontsize=20)
plt.imshow(im_crop)
PATH_FOLDER_CSVS+"\\" +fn+ext
# +
df = pd.read_csv(path_csv)
for index, row in df.iterrows():
# path_img = join(PATH_FOLDER_CSVS,img_name)
img = cut_images(PATH_FOLDER_CSVS+"\\" +fn+ext, row['xmin'], row['ymin'], row['xmax'], row['ymax'], row['label'])
# img = cut_images(r'\\ica-094\share\nilton\formatos\f1.3-.png', row['xmin'], row['ymin'], row['xmax'], row['ymax'], row['label'])
# Adding custom options
custom_config = r'--oem 3 --psm 6'
print(pytesseract.image_to_string(img, config=custom_config))
#break
# -
print(type(pytesseract.image_to_string(img, config=custom_config)))
# +
df = pd.read_csv(path_csv)
for index, row in df.iterrows():
path_img = join(PATH_FOLDER_CSVS,img_name)
plot_images(path_img, row['xmin'], row['ymin'], row['xmax'], row['ymax'], row['label'])
img = cut_images(PATH_FOLDER_CSVS+"\\" +fn+ext, row['xmin'], row['ymin'], row['xmax'], row['ymax'], row['label'])
# img = cut_images(r'\\ica-094\share\nilton\formatos\f1.3-.png', row['xmin'], row['ymin'], row['xmax'], row['ymax'], row['label'])
# Adding custom options
custom_config = r'--oem 3 --psm 6'
# print(pytesseract.image_to_string(img, config=custom_config))
df.loc[index,('text')]=pytesseract.image_to_string(img, config=custom_config)
#break
# -
# +
df.to_csv(f"{PATH_FOLDER_CSVS}\\csv_novo\\{fn}.csv",index=False, header=True)
# df.to_csv(PATH_FOLDER_CSVS+"\\csv_novo\\"+fn+".csv",index=False, header=True)
# df.to_csv(r'\\ica-094\share\nilton\formatos\csv_novo\f1.3-copy.csv',index=False, header=True)
# -
df.head()
path_img
# +
# img = cut_images(r'\\ica-095\tables\manntis\formatos\f1.2-.jpg', 139, 41, 281, 71, 'label_E')
# # img = cv2.imread('image.jpg')
# # Adding custom options
# custom_config = r'--oem 3 --psm 6'
# pytesseract.image_to_string(img, config=custom_config)
# -
| plto_csv2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.2 64-bit
# language: python
# name: python3
# ---
# +
import numpy as np
a = np.zeros(3)
a
type(a)
a.shape
a.shape = (3, 1)
a
a = np.ones(10)
a
a = np.linspace(2, 10, 5)
a
a = np.array([[1,2,34,4], [1,2,3,4,5,6],[2,1,33,4,55]])
a.shape
#see information about a
# ?a
#produce random value
np.random.seed(0)
z1 = np.random.randint(10, size=6)
z1
#get last element of array
z1[-1]
# +
#working on photo
from skimage import io
photo = io.imread('W0Sww.jpg')
type(photo)
photo.shape
import matplotlib.pyplot as plt
plt.imshow(photo)
# -
plt.imshow(photo[::-1])
plt.imshow(photo[:, ::-1])
plt.imshow(photo[50:100, 40:80])
plt.imshow(photo[::2, ::2])
photo_sin = np.sin(photo)
photo_sin
print(np.sum(photo))
print(np.prod(photo))
print(np.mean(photo))
print(np.std(photo))
print(np.var(photo))
print(np.argmin(photo))
print(np.argmax(photo))
z = np.array([1, 2, 3, 4, 5])
z>3
z[z>2]
photo_masked = np.where(photo > 100, 255, 0)
plt.imshow(photo_masked)
plt.imshow(photo[:, :, 0].T)
np.sort([4, 3, 2])
| my intereseted short codes/numpy/how_numpy_works.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="rznSDgbvGggG"
# ## Tic-Tac-Toe Agent
#
# In this notebook, you will learn to build an RL agent (using Q-learning) that learns to play Numerical Tic-Tac-Toe with odd numbers. The environment is playing randomly with the agent, i.e. its strategy is to put an even number randomly in an empty cell. The following is the layout of the notebook:
# - Defining epsilon-greedy strategy
# - Tracking state-action pairs for convergence
# - Define hyperparameters for the Q-learning algorithm
# - Generating episode and applying Q-update equation
# - Checking convergence in Q-values
# + [markdown] colab_type="text" id="8eDb8PxBGggH"
# #### Importing libraries
# Write the code to import Tic-Tac-Toe class from the environment file
# + colab={} colab_type="code" id="6SFNYceFGggJ"
## from <TC_Env> import <TicTacToe> - import your class from environment file
from TCGame_Env import TicTacToe ## Importing TicTacToe class from 'TCGame_Env.py' file
import collections
import numpy as np
import random
import pickle
import time
from matplotlib import pyplot as plt
# %matplotlib inline
## Define environment as env
env= TicTacToe() #Instantiation
# + colab={} colab_type="code" id="wYLQyopEG8nz"
# Function to convert state array into a string to store it as keys in the dictionary
# states in Q-dictionary will be of form: x-4-5-3-8-x-x-x-x
# x | 4 | 5
# ----------
# 3 | 8 | x
# ----------
# x | x | x
def Q_state(state):
return ('-'.join(str(e) for e in state)).replace('nan','x')
# + colab={} colab_type="code" id="ZebMOoiVHBBr"
## Defining a function which will return valid (all possible actions) actions corresponding to a state
## Important to avoid errors during deployment.
def valid_actions(state):
valid_Actions = []
valid_Actions = [i for i in env.action_space(state)[0]] ###### -------please call your environment as env
return valid_Actions
# + colab={} colab_type="code" id="IRciPUkYHDWf"
## Defining a function which will add new Q-values to the Q-dictionary.(Initialization of Q-values at the start: Only for Agent)
def add_to_dict(state):
state1 = Q_state(state)
if state1 not in Q_dict.keys():
valid_act = valid_actions(state)
for action in valid_act:
Q_dict[state1][action]=0
# -
## Printing the default tic tac toe board positions
Q_state(env.state)
## Printing all the valid actions (all possible agent actions available at start)
valid_actions(env.state)
##Number of all possible valid agent actions at start
len(valid_actions(env.state))
# + [markdown] colab_type="text" id="fNNi_EfHGggM"
# #### Epsilon-greedy strategy - Write your code here
#
# (you can build your epsilon-decay function similar to the one given at the end of the notebook)
# + colab={} colab_type="code" id="m0lMfqiJGggN"
## Defining epsilon-greedy policy. You can choose any function epsilon-decay strategy
def epsilon_greedy_policy(state, time):
epsilon = min_epsilon + (max_epsilon - min_epsilon) * np.exp(-decay_rate*time) ##Based on the given epsilon-decay example
z = np.random.random() ##Takes random value [0,1)
if z > epsilon:
state1 = Q_state(state)
action = max(Q_dict[state1],key=Q_dict[state1].get) #Exploitation: this gets the action corresponding to max Q-value for the current state
else:
possible_actions = [i for i in env.action_space(state)[0]]
action = possible_actions[np.random.choice(range(len(possible_actions)))] #Exploration: randomly choosing an action from all possible actions for the agent
return action
# + [markdown] colab_type="text" id="H2kyQHOMGggR"
# #### Tracking the state-action pairs for checking convergence - write your code here
# + colab={} colab_type="code" id="qcxZ29vdGggS"
## Initialise Q_dictionary as 'Q_dict' and States_tracked as 'States_track' (for convergence)
Q_dict = collections.defaultdict(dict)
States_track =collections.defaultdict(dict)
# + colab={} colab_type="code" id="vs73iv8fHOxV"
## Initialising states to be tracked
def initialise_tracking_states():
sample_state_action_Qvalues = [('x-x-x-x-x-x-x-x-x',(2,5)), ('x-x-9-x-x-6-x-x-x',(7,7)),
('x-1-6-x-x-x-x-x-x',(0,9)),('x-1-x-x-x-x-2-x-x',(2,3))] #Select some 4 Q-values
for q_val in sample_state_action_Qvalues:
state = q_val[0]
action = q_val[1]
States_track[state][action] = []
# -
initialise_tracking_states()
States_track
## Listing states in 'State_track'
States_track.keys()
## Action/s within the state
States_track['x-x-x-x-x-x-x-x-x'].keys()
# + colab={} colab_type="code" id="dAbwJDMVHpwl"
## Defining a function to save the Q-dictionary as a pickle file
def save_obj(obj, name ):
with open(name + '.pkl', 'wb') as f:
pickle.dump(obj, f, pickle.HIGHEST_PROTOCOL)
# + colab={} colab_type="code" id="6Pyj7nMVHsBi"
def save_tracking_states():
for state in States_track.keys():
for action in States_track[state].keys():
if state in Q_dict and action in Q_dict[state]:
States_track[state][action].append(Q_dict[state][action])
# + [markdown] colab_type="text" id="-iPt--E9GggV"
# #### Define hyperparameters ---write your code here
# + colab={} colab_type="code" id="G0_f5czFGggW"
## Defining hyperparameters for the training
EPISODES = 4000000 # total no. of episodes
LR = 0.01 # learning rate
GAMMA = 0.9 # discount factor
max_epsilon = 1.0 # Greed: 100%
min_epsilon = 0.001 # Min_Greed: 0.1%
decay_rate = 0.000002 # epsilon decay rate
threshold = 2500 # no. of episodes after which states_tracked will be saved
policy_threshold = 10000 # no of episodes after which Q dictionary/table will be saved
# + [markdown] colab_type="text" id="Md6twJ7wGggh"
# ### Q-update loop ---write your code here
# + colab={} colab_type="code" id="ldCgQuDNGggj"
import time
start_time = time.time()
for episode in range(EPISODES):
##### Start writing your code from the next line
env = TicTacToe() # call the environment
curr_state = env.state
terminal = False
add_to_dict(curr_state) # adding the current state to dictionary
while terminal != True:
curr_state1 = Q_state(curr_state)
curr_action = epsilon_greedy_policy(curr_state, episode) # applying epislon-greedy policy method
next_state, reward, terminal = env.step(curr_state, curr_action) # getting next_state, reward, terminal flag
next_state_temp = Q_state(next_state)
add_to_dict(next_state)
# Updating rules
if terminal != True:
max_next = max(Q_dict[next_state_temp],key=Q_dict[next_state_temp].get)
#this gets the action corresponding to max q-value of next state
Q_dict[curr_state1][curr_action] += LR * ((reward + (GAMMA*(Q_dict[next_state_temp][max_next])))
- Q_dict[curr_state1][curr_action] )
else:
Q_dict[curr_state1][curr_action] += LR * ((reward - Q_dict[curr_state1][curr_action]))
# navigating to next state
curr_state = next_state
#states tracking
if ((episode+1)%threshold)==0:
save_tracking_states()
save_obj(States_track,'States_tracking')
print(episode) ## Tracking episode at step increment of threshold value
if ((episode+1)% policy_threshold) == 0: # Every 10000 episodes, the Q-dict will be saved
save_obj(Q_dict,'Policy_Q_dict')
elapsed_time = time.time() - start_time
save_obj(States_track,'States_tracked') #States_tracked in 'States_tracked' file
save_obj(Q_dict,'Policy') #Q_table saved in 'Policy' file
# + [markdown] colab_type="text" id="t6eMFbb8Ggg2"
# #### Check the Q-dictionary
# + colab={} colab_type="code" id="fr9d2fcVGgg4"
Q_dict
# + colab={} colab_type="code" id="F1tnDJWkGgg9"
len(Q_dict)
# + colab={} colab_type="code" id="cFgUqfcQGghB"
# Try checking for one of the states - that which action your agent thinks is the best -----This will not be evaluated
#Random State Key
random_state_key= np.random.choice(list(Q_dict.keys()))
random_state_key
# +
## Dictionary of all state-action pairs and their respective Q_values for the 'random_state_key' (i.e. any random state)
Q_val_rand = Q_dict[random_state_key]
## Creating tuples with action pairs and Q-values for 'random_state_key' (descending order)
sorted_tuples= sorted(Q_val_rand.items(), key= lambda item: item[1], reverse= True)
## Created sorted dictionary of action and Q-value pairs using sorted_tuples
sorted_dict= {}
sorted_dict= {k:v for k,v in sorted_tuples} #Sorted dictionary (reverse: descending order of values)
print(sorted_dict)
# +
##Selecting the best action for the state 'random_state_key' (Finding the action with max Q value for 'random_state_key')
import itertools
#Best action for the random_state_key
print(f"The best action for state '{random_state_key}' is:")
list(dict(itertools.islice(sorted_dict.items(), 1)).keys())[0]
# + [markdown] colab_type="text" id="KGPZEQDFGghG"
# #### Check the states tracked for Q-values convergence
# (non-evaluative)
# + colab={} colab_type="code" id="9s1Tvz8HGghH"
## Write the code for plotting the graphs for state-action pairs tracked
plt.figure(0, figsize=(20,15))
#Graph 1: Convergence Plot for state-action pair: ['x-x-x-x-x-x-x-x-x'][(2,5)]
x_axis = np.asarray(range(0, len(States_track['x-x-x-x-x-x-x-x-x'][(2,5)])))
plt.subplot(221)
plt.plot(x_axis,np.asarray(States_track['x-x-x-x-x-x-x-x-x'][(2,5)]))
plt.title("Convergence plot for: ['x-x-x-x-x-x-x-x-x'][(2,5)]", fontsize=15, fontweight='bold')
plt.ylabel("Q_values", fontsize=13, fontstyle='italic')
plt.xlabel("No. of Episodes (step increment of threshold value)", fontsize=13, fontstyle='italic')
plt.grid(True)
plt.show
#Graph 2: Convergence Plot for state-action pair: ['x-x-9-x-x-6-x-x-x'][(7,7)]
x_axis = np.asarray(range(0, len(States_track['x-x-9-x-x-6-x-x-x'][(7,7)])))
plt.subplot(222)
plt.plot(x_axis,np.asarray(States_track['x-x-9-x-x-6-x-x-x'][(7,7)]))
plt.title("Convergence plot for: ['x-x-9-x-x-6-x-x-x'][(7,7)]", fontsize=15, fontweight='bold')
plt.ylabel("Q_values", fontsize=13, fontstyle='italic')
plt.xlabel("No. of Episodes (step increment of threshold value)", fontsize=13, fontstyle='italic')
plt.grid(True)
plt.show
#Graph 3: Convergence Plot for state-action pair: ['x-1-6-x-x-x-x-x-x'][(0,9)]
x_axis = np.asarray(range(0, len(States_track['x-1-6-x-x-x-x-x-x'][(0,9)])))
plt.subplot(223)
plt.plot(x_axis,np.asarray(States_track['x-1-6-x-x-x-x-x-x'][(0,9)]))
plt.title("Convergence plot for: ['x-1-6-x-x-x-x-x-x'][(0,9)]", fontsize=15, fontweight='bold')
plt.ylabel("Q_values", fontsize=13, fontstyle='italic')
plt.xlabel("No. of Episodes (step increment of threshold value)", fontsize=13, fontstyle='italic')
plt.grid(True)
plt.show
#Graph 4: Convergence Plot for state-action pair: ['x-1-x-x-x-x-2-x-x'][(2,3)]
x_axis = np.asarray(range(0, len(States_track['x-1-x-x-x-x-2-x-x'][(2,3)])))
plt.subplot(224)
plt.plot(x_axis,np.asarray(States_track['x-1-x-x-x-x-2-x-x'][(2,3)]))
plt.title("Convergence plot for: ['x-1-x-x-x-x-2-x-x'][(2,3)]", fontsize=15, fontweight='bold')
plt.ylabel("Q_values", fontsize=13, fontstyle='italic')
plt.xlabel("No. of Episodes (step increment of threshold value)", fontsize=13, fontstyle='italic')
plt.grid(True)
plt.show
# + [markdown] colab_type="text" id="b2Opp8_NITkC"
# ### Epsilon - Decay Check
# + colab={} colab_type="code" id="gQ_D_JsuGghR"
max_epsilon = 1.0 #Greedy 100%
min_epsilon = 0.001 #Min_epsilon: 0.1%
time = np.arange(0,4000000) # 4e+6 episodes
epsilon = []
for i in range(0,4000000):
epsilon.append(min_epsilon + (max_epsilon - min_epsilon) * np.exp(-0.000002*i))
plt.plot(time, epsilon)
plt.grid(True)
plt.show()
# + colab={} colab_type="code" id="59BRf43IJiQ1"
| TicTacToe_Agent.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Q-learning Method
# ## Prepare the packages
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
import sys
if "../" not in sys.path:
sys.path.append("../")
from collections import defaultdict
import gym
from el_agent import ELAgent
from frozen_lake_util import show_q_value
# ## Define Q-learning Agent
class QLearningAgent(ELAgent):
def __init__(self, epsilon=0.1):
super().__init__(epsilon)
def learn(self, env, episode_count=1000, gamma=0.9,
learning_rate=0.1, render=False, report_interval=50):
self.init_log()
self.Q = defaultdict(lambda: [0] * len(actions))
actions = list(range(env.action_space.n))
for e in range(episode_count):
s = env.reset()
done = False
while not done:
if render:
env.render()
a = self.policy(s, actions)
n_state, reward, done, info = env.step(a)
gain = reward + gamma * max(self.Q[n_state])
estimated = self.Q[s][a]
self.Q[s][a] += learning_rate * (gain - estimated)
s = n_state
else:
self.log(reward)
if e != 0 and e % report_interval == 0:
self.show_reward_log(episode=e)
# ## Train Agent
def train():
agent = QLearningAgent()
env = gym.make("FrozenLakeEasy-v0")
agent.learn(env, episode_count=500)
show_q_value(agent.Q)
agent.show_reward_log()
agent = train()
| EL/notebooks/Q-learning.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.7.2
# language: julia
# name: julia-1.7
# ---
# # Evolution via method of lines
# +
using FFTW
using Plots
using OrdinaryDiffEq
using LinearAlgebra: norm, mul!
using Test
using BenchmarkTools
using Random
@info "Threads: $(FFTW.nthreads())"
# -
# ## Operators
function get_operators(N, κ₀)
# Differentiation in spectral space
Dx_hat = im * κ₀ * [
ifelse(k1 ≤ div(N, 2) + 1, k1 - 1, k1 - 1 - N) for k2 in 1:div(N, 2)+1, k1 in 1:N
]
Dy_hat = im * κ₀ * [k2 - 1 for k2 in 1:div(N, 2)+1, k1 in 1:N]
Delta_hat = - κ₀^2 * [
ifelse(k1 ≤ div(N, 2) + 1, (k1 - 1)^2 + (k2 - 1)^2, (k1 - 1 - N)^2 + (k2 - 1)^2)
for k2 in 1:div(N, 2)+1, k1 in 1:N
]
# For the Basdevant formulation
DxsqDysq_hat = Dx_hat.^2 .- Dy_hat.^2
Dxy_hat = Dx_hat .* Dy_hat
# Recovering of the velocity field from the vorticity
Hu_hat = - Dy_hat ./ Delta_hat
Hu_hat[1, 1] = 0.0
Hv_hat = Dx_hat ./ Delta_hat
Hv_hat[1, 1] = 0.0
return Dx_hat, Dy_hat, Delta_hat, Hu_hat, Hv_hat, DxsqDysq_hat, Dxy_hat
end
# Methods to generate a scalar field from a list of wavenumbers and amplitudes
# +
function field_from_modes(L, N, modes::Matrix{<:Integer}, amps::Matrix{<:Real})
κ₀ = 2π/L
x = y = (L/N):(L/N):L
field = sum(
[
2κ₀^2 * (k[1]^2 + k[2]^2) * (
a[1] * cos.(κ₀ * (k[1] * one.(y) * x' + k[2] * y * one.(x)'))
- a[2] * sin.(κ₀ * (k[1] * one.(y) * x' + k[2] * y * one.(x)'))
)
for (k, a) in zip(eachrow(modes), eachrow(amps))
]
)
return field
end
function field_from_modes(rng::AbstractRNG, L, N, num_modes::Int)
modes = rand(rng, 1:div(N,10), num_modes, 2)
amps = rand(rng, num_modes, 2)
field = field_from_modes(L, N, modes, amps)
return field
end
field_from_modes(L, N, num_modes::Int) = field_from_modes(Xoshiro(), L, N, num_modes)
# -
# Differential equations
function nsepervort_hat_rhs!(dvorhattdt, vort_hat, params, t)
operators, vars, auxs, plans = params
Dx_hat, Dy_hat, Delta_hat, Hu_hat, Hv_hat,
DxsqDysq_hat, Dxy_hat = operators
ν, g_hat, N, Nsub = vars
u_hat, v_hat, u, v, uv, v2u2, uv_hat, v2u2_hat = auxs
plan, plan_inv = plans
vort_hat[div(Nsub,2) + 1:end, :] .= 0.0im
vort_hat[:, div(Nsub,2) + 1:div(N,2) + div(Nsub,2)] .= 0.0im
u_hat .= Hu_hat .* vort_hat
v_hat .= Hv_hat .* vort_hat
mul!(u, plan_inv, u_hat)
mul!(v, plan_inv, v_hat)
uv .= u .* v
v2u2 .= v.^2 .- u.^2
mul!(uv_hat, plan, uv)
mul!(v2u2_hat, plan, v2u2)
dvorhattdt .= g_hat .+ ν .* Delta_hat .* vort_hat .- DxsqDysq_hat .* uv_hat .- Dxy_hat .* v2u2_hat
# dealiasing
dvorhattdt[div(Nsub,2) + 1:end, :] .= 0.0im
dvorhattdt[:, div(Nsub,2) + 1:div(N,2) + div(Nsub,2)] .= 0.0im
return dvorhattdt
end
# ## The spatial domain and its discretization
L = 2π
κ₀ = 2π/L
N = 128
Nsub = 84
x = y = (L/N):(L/N):L
# ## Test convergence to one-mode steady state
# +
ν = 1.0e-0 # viscosity
Dx_hat, Dy_hat, Delta_hat, Hu_hat, Hv_hat, DxsqDysq_hat, Dxy_hat = get_operators(N, κ₀)
vort_init = field_from_modes(L, N, 4)
vort_init_hat = rfft(vort_init)
g_steady = field_from_modes(L, N, 1)
g_steady_hat = rfft(g_steady)
vort_steady_hat = - g_steady_hat ./ Delta_hat
vort_steady_hat[1, 1] = 0.0im
vort_steady = irfft(vort_steady_hat, N)
vort_hat = copy(vort_init_hat)
vort = irfft(vort_hat, N)
u_hat = similar(vort_hat)
v_hat = similar(vort_hat)
u = similar(vort)
v = similar(vort)
uv = similar(vort)
v2u2 = similar(vort)
uv_hat = similar(vort_hat)
v2u2_hat = similar(vort_hat)
plan = plan_rfft(vort, flags=FFTW.MEASURE)
plan_inv = plan_irfft(vort_hat, N, flags=FFTW.MEASURE)
operators = Dx_hat, Dy_hat, Delta_hat, Hu_hat, Hv_hat, DxsqDysq_hat, Dxy_hat
vars = ν, g_steady_hat, N, Nsub
auxs = u_hat, v_hat, u, v, uv, v2u2, uv_hat, v2u2_hat
plans = plan, plan_inv
params = (
operators,
vars,
auxs,
plans
)
nothing
# +
tspan = (0.0, 10.0)
prob = ODEProblem(nsepervort_hat_rhs!, vort_init_hat, tspan, params)
nothing
# +
OrdinaryDiffEq.ForwardDiff.can_dual(::Type{ComplexF64}) = true
sol = solve(prob, Vern9()) # QNDF() etc.
sol.retcode
# -
# Distância entre a solução estacionária e a solução ao final do intervalo de tempo.
norm(sol.u[end] - vort_steady_hat) * L/N /N
vort = irfft(sol.u[end], N)
nothing
# No espaço físico:
norm(vort .- vort_steady) * L / N
# Na norma do máximo:
maximum(abs, vort .- vort_steady)
heatmap(x, y, vort, xlabel="x", ylabel="y", title="vorticity", titlefont=12)
surface(x, y, vort, xlabel="x", ylabel="y", title="vorticity", titlefont=12)
surface(x, y, vort .- vort_steady, xlabel="x", ylabel="y", title="vorticity", titlefont=12)
# We can check how the $L^2$ distance of the vorticity to the steady state evolves.
error = [norm(sol.u[n] .- vort_steady_hat) * L/N / N for n in eachindex(sol.u)]
nothing
# +
plt1 = plot(sol.t, error, title="Enstrophy convergence", titlefont = 10, xaxis="time", yaxis = "enstrophy", label=false)
plt2 = plot(sol.t[1:div(end, 100)], error[1:div(end, 100)], title="Enstrophy convergence", titlefont = 10, xaxis="time", yaxis = "enstrophy", label=false)
plt3 = plot(sol.t[div(end,100):div(end, 50)], error[div(end,100):div(end, 50)], title="Enstrophy convergence", titlefont = 10, xaxis="time", yaxis = "enstrophy", label=false)
plt4 = plot(sol.t[end-1000:end], error[end-1000:end], title="Enstrophy convergence", titlefont = 10, xaxis="time", yaxis = "enstrophy", label=false)
plot(plt1, plt2, plt3, plt4, layout = 4)
| generated/literated/tests_mol.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.2 64-bit (''venv'': venv)'
# name: python3
# ---
import pandas as pd
data = pd.read_excel("./nakladnaya.xls", header=None, skiprows=2)
data.dropna(how="all", inplace=True)
# data.dropna(how="all", inplace=True, axis=1)
data
data.iloc[0, 4][2:9]
data.iloc[6:8, :].dropna(axis=1, how='all')
data.iloc[6:8, [1, 2, 6, 9, 11, 12]]
data.iloc[6:8, [0, 2, 6, 9, 11, 12]]
data.iloc[7:8, :].dropna(axis=1, how='all')
table = data.iloc[6:8, :].dropna(axis=1, how='any')
table
table.to_excel("table.xls", index=False)
table.to_excel(writer, index=False, sheet_name='Таблица')
# writer.save()
| unit_3/python_13.8.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Optimization
# language: python
# name: optimai
# ---
from z3 import *
from itertools import combinations
from typing import Sequence
# Read instance file:
# +
input_filename = '../../Instances/15x15.txt'
w, h, n, DX, DY = None, None, None, None, None
with open(input_filename, 'r') as f_in:
lines = f_in.read().splitlines()
split = lines[0].split(' ')
w = int(split[0])
h = int(split[1])
n = int(lines[1])
DX = []
DY = []
for i in range(int(n)):
split = lines[i + 2].split(' ')
DX.append(int(split[0]))
DY.append(int(split[1]))
# -
# Solver:
solver = Solver()
# Model:
XY = [(Int(f'XY_{i}_0'), Int(f'XY_{i}_1')) for i in range(n)]
XY
# Constraints:
# Non-overlapping constraint
for (i, j) in combinations(range(n), 2):
solver.add(Or(XY[i][0] + DX[i] <= XY[j][0],
XY[j][0] + DX[j] <= XY[i][0],
XY[i][1] + DY[i] <= XY[j][1],
XY[j][1] + DY[j] <= XY[i][1]))
# Boundaries consistency constraint
for i in range(n):
solver.add(XY[i][0] >=0)
solver.add(XY[i][1] >= 0)
solver.add(XY[i][0] + DX[i] <= w)
solver.add(XY[i][1] + DY[i] <= h)
# +
# Cumulative constraint
def cumulative(solver, S: Sequence, D: Sequence, R: Sequence, C: int):
# Iterate over the durations
for u in D:
solver.add(
Sum(
[If(And(S[i] <= u, u < S[i] + D[i]), R[i], 0) for i in range(n)]
) <= C)
# Implied constraints
cumulative(solver,
S=list(map(lambda t: t[0], XY)), # take x coordinates
D=DX,
R=DY,
C=h)
cumulative(solver,
S=list(map(lambda t: t[1], XY)), # take y coordinates
D=DY,
R=DX,
C=w)
# -
# %%time
solver.check()
# From Z3 model solution to file:
# +
model = solver.model()
xy = [(model[XY[i][0]], model[XY[i][1]]) for i in range(n)]
xy
# -
output_filename = '../../pwp_utilities/15x15_sol.txt'
with open(output_filename, 'w') as f_out:
f_out.write('{} {}\n'.format(w, h))
f_out.write('{}\n'.format(n))
for i in range(n):
f_out.write('{} {}\t{} {}\n'.format(DX[i], DY[i], xy[i][0], xy[i][1]))
| SMT/src/pwp-final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
import warnings
warnings.filterwarnings("ignore")
# + [markdown] pycharm={"name": "#%% md\n"}
# class: center, middle
#
# # DEMOPS: Roll decay
# ## Status 2020-02-27
# ---
#
# ### The work has been focused on:
# --
#
# ### Building a roll decay DB
# * SQL database with MDL model tests 2007-2020</li>
#
# --
#
# ### Measure Rolldamping
# * System identification of roll decay test
# * Four methods have been tested
#
# --
#
# ### Building roll damping DB
# * System identification of roll decay DB
#
# -
# ---
# name: DB
# ## Building a roll decay DB
# ---
# template: DB
# <img src="mdl_db.png" alt="drawing" height="550"/>
#
# +
from sqlalchemy import create_engine
import data
from mdldb.mdl_db import MDLDataBase
from mdldb.tables import Base, Model, LoadingCondition, Run, RolldecayLinear, RolldecayDirect, RolldecayNorwegian
import pandas as pd
from sympy.physics.vector.printing import vpprint, vlatex
from IPython.display import display, Math, Latex, Markdown
engine = create_engine('sqlite:///' + data.mdl_db_path)
db = MDLDataBase(engine=engine)
df = pd.read_sql_table(table_name='run', con=engine, index_col='id')
# -
# ---
# template: DB
#
# ### MDL DB
s = """The database from MDL currently contains *%i* tests conducted between **%i** and **%i**.
""" % (len(df),
df['date'].min().year,
df['date'].max().year,)
Markdown(s)
# --
# +
exclude = ['smeg','prepare','rodergrundvinkel','rerun','unknown','none standard','teckenkoll']
emphase = ['roll decay']
test_type_s=''
for test_type in df['test_type'].unique():
if test_type in exclude:
continue
if test_type in emphase:
output = '**%s**' % test_type
else:
output = test_type
test_type_s+=' %s,' % output
s = """<u>Test types:</u>%s
""" % (test_type_s)
Markdown(s)
# -
# --
number_of_rolldecay = (df['test_type']=='roll decay').sum()
s = """**%i** of the tests are roll decay tests """ % number_of_rolldecay
Markdown(s)
# + pycharm={"is_executing": false, "name": "#%%\n"}
import sympy as sp
from rolldecayestimators.symbols import *
import rolldecayestimators.equations as equations
import rolldecayestimators.direct_estimator as direct_estimator
import rolldecayestimators.direct_linear_estimator as direct_linear_estimator
import rolldecayestimators.direct_estimator_improved as direct_estimator_improved
from latex_helpers import pylatex_extenders
import latex_helpers
import os.path
from rigidbodysimulator.substitute_dynamic_symbols import substitute_dynamic_symbols, find_name, find_derivative_name, lambdify, find_derivatives
import dill
# -
# ---
# name: equations
# ## Measure Rolldamping
# ---
# template: equations
#
# ### General equation for roll decay motion:
Math(vlatex(equations.roll_equation_general))
# --
#
# During a roll decay test external moment is zero:
#
# --
Math(vlatex(equations.roll_decay_equation_general))
# --
#
# ### Linear model
#
# Linearizing the stiffness (ship static stability)
#
# --
#
latex = vlatex(equations.linear_stiffness_equation)
Math(latex)
# --
#
# Linear roll decay equation:
Math(vlatex(equations.roll_decay_linear_equation))
# ---
# template: equations
#
# Damping term $ B $ can be linear:
Math(vlatex(sp.Eq(B,zeta)))
# --
#
# or quadratic:
Math(vlatex(sp.Eq(B,zeta+d*sp.Abs(phi_dot))))
# ---
# name: nonlinearity
# ## Damping nonlinearity
# ---
# template: nonlinearity
# The linear model is sometimes too simple:
# <img src="nonlinearity.png" alt="drawing" width=800/>
#
# ---
# template: nonlinearity
s="""Adding quadratic damping: $%s$""" % vlatex(sp.Eq(B,zeta+d*sp.Abs(phi_dot)))
Markdown(s)
# <img src="nonlinearity_quadratic.png" alt="drawing" width=800/>
#
# ---
# template: nonlinearity
# <img src="nonlinearity_sweep.png" alt="drawing" height=450 width=800/>
#
# --
#
# * linear model has low score when including large angles
#
# --
# * linear model damping $\zeta$ increase for large angles
#
# --
# * for quadratic model $d$ increase instead
#
#
# ---
# name: varying-stiffness
# ## Varying stiffness
#
# ---
# template: varying-stiffness
# The linear stiffness assumption is not valid for all ships at large roll angles:
#
# ---
# template: varying-stiffness
Math(vlatex(equations.linear_stiffness_equation))
# <img src="varying_stiffness.png" alt="drawing" width=800/>
# ---
# template: varying-stiffness
#
# ### Natural frequency $ \omega_0 $ for each oscillation
# <img src="varying_stiffness2.png" alt="drawing" width=800/>
#
# ---
# template: varying-stiffness
#
# ### Adding a quadratic term to the stiffness:
Math(vlatex(equations.quadratic_stiffness_equation))
# <img src="varying_stiffness3.png" alt="drawing" width=800/>
# ---
# name: building-roll-damping-DB
# ## Building roll damping DB
# ---
# +
plots = ['zeta','d','mean_damping','omega0','score']
s = ''
for plot in plots:
code = """
template: building-roll-damping-DB
.right[]
---
""" % plot
s+=code
Markdown(s)
# -
# ## Next steps
#
# --
#
# ### Regression on roll damping DB
# --
#
# ### Ikeda method to predict roll damping
# --
#
# ### Gather more meta data
# * Bilge keels
# * Inertia
# * Hull form (probably takes too much time)
#
# --
#
# ### Start writing paper
#
#
#
# ---
# class: center, middle
# ## End
| docs/presentation1/presentation1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://www.kaggle.com/code/tanavbajaj/anime-recommend-notebook?scriptVersionId=93199932" target="_blank"><img align="left" alt="Kaggle" title="Open in Kaggle" src="https://kaggle.com/static/images/open-in-kaggle.svg"></a>
# + [markdown] papermill={"duration": 0.011674, "end_time": "2022-04-16T22:47:49.548772", "exception": false, "start_time": "2022-04-16T22:47:49.537098", "status": "completed"} tags=[]
# 
# # Insert Anime Name
# + papermill={"duration": 0.023818, "end_time": "2022-04-16T22:47:49.584199", "exception": false, "start_time": "2022-04-16T22:47:49.560381", "status": "completed"} tags=[]
anime_name='Death Note'
# + [markdown] papermill={"duration": 0.010354, "end_time": "2022-04-16T22:47:49.605511", "exception": false, "start_time": "2022-04-16T22:47:49.595157", "status": "completed"} tags=[]
# # Importing Libraries
# + papermill={"duration": 0.017517, "end_time": "2022-04-16T22:47:49.633597", "exception": false, "start_time": "2022-04-16T22:47:49.61608", "status": "completed"} tags=[]
import numpy as np
import warnings
import os
import pandas as pd
warnings.filterwarnings('ignore')
# + [markdown] papermill={"duration": 0.010727, "end_time": "2022-04-16T22:47:49.655094", "exception": false, "start_time": "2022-04-16T22:47:49.644367", "status": "completed"} tags=[]
# # Reading Dataset
# + papermill={"duration": 3.119099, "end_time": "2022-04-16T22:47:52.784844", "exception": false, "start_time": "2022-04-16T22:47:49.665745", "status": "completed"} tags=[]
anime = pd.read_csv('../input/anime-recommendations-database/anime.csv')
anime_rating = pd.read_csv('../input/anime-recommendations-database/rating.csv')
# + [markdown] papermill={"duration": 0.010798, "end_time": "2022-04-16T22:47:52.807276", "exception": false, "start_time": "2022-04-16T22:47:52.796478", "status": "completed"} tags=[]
# # Cleaning Data
# + papermill={"duration": 38.203534, "end_time": "2022-04-16T22:48:31.021914", "exception": false, "start_time": "2022-04-16T22:47:52.81838", "status": "completed"} tags=[]
ratings = pd.DataFrame(anime_rating.groupby('anime_id')['rating'].mean())
ratings['num of ratings'] = pd.DataFrame(
anime_rating.groupby('anime_id')['rating'].count())
# anime_rating.drop(['genre', 'type', 'episodes'],
# axis=1, inplace=True)
anime2 = anime_rating.pivot_table(
index='user_id', columns='anime_id', values='rating')
anime2.fillna(0, inplace=True)
# + papermill={"duration": 0.023347, "end_time": "2022-04-16T22:48:31.056927", "exception": false, "start_time": "2022-04-16T22:48:31.03358", "status": "completed"} tags=[]
anime_ids= anime[anime['name']==anime_name]
anime_id=anime_ids['anime_id']
anime_user_ratings = anime2[int(anime_id)]
# + [markdown] papermill={"duration": 0.011441, "end_time": "2022-04-16T22:48:31.081017", "exception": false, "start_time": "2022-04-16T22:48:31.069576", "status": "completed"} tags=[]
# # Pairwise Correlation
# + papermill={"duration": 11.20835, "end_time": "2022-04-16T22:48:42.300904", "exception": false, "start_time": "2022-04-16T22:48:31.092554", "status": "completed"} tags=[]
similar_anime = anime2.corrwith(anime_user_ratings, method='pearson')
corr_anime = pd.DataFrame(similar_anime, columns=['Correlation'])
# + [markdown] papermill={"duration": 0.011406, "end_time": "2022-04-16T22:48:42.323917", "exception": false, "start_time": "2022-04-16T22:48:42.312511", "status": "completed"} tags=[]
# # Print Output
# + papermill={"duration": 0.024156, "end_time": "2022-04-16T22:48:42.359476", "exception": false, "start_time": "2022-04-16T22:48:42.33532", "status": "completed"} tags=[]
x = corr_anime.sort_values('Correlation', ascending=False).index
anime_names= anime[anime['anime_id']==x[1]]
anime_y=anime_names['name']
print(str(anime_y))
| anime-recommend-notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import jax.numpy as jnp
poorHigh_w = jnp.array([ 5. , 25.489805 , 21.933598 , 19.946459 , 18.448935 ,
17.45715 , 16.52748 , 15.581397 , 15.200653 , 15.140821 ,
15.242023 , 15.012157 , 14.901801 , 14.826152 , 14.718977 ,
14.471284 , 14.245392 , 14.148019 , 13.90585 , 13.748291 ,
13.730789 , 13.6384 , 13.615738 , 13.678733 , 13.951984 ,
14.444705 , 15.235136 , 16.139376 , 17.287508 , 18.990013 ,
21.015928 , 23.95729 , 25.575556 , 27.094559 , 28.418629 ,
28.952412 , 29.83769 , 31.076324 , 32.366013 , 33.850327 ,
36.271427 , 38.8551 , 42.12923 , 44.86371 , 46.725567 ,
47.317757 , 40.52259 , 34.561634 , 29.594255 , 25.075418 ,
21.5741 , 18.79675 , 17.09875 , 15.85105 , 15.2683735,
14.994113 , 14.885412 , 14.888294 , 14.549519 , 13.525763 ,
11.5109005])
poorLow_w = jnp.array([ 5. , 25.489805, 21.337265, 20.165863, 21.798952,
22.987448, 24.736362, 25.127794, 26.073416, 28.507818,
29.035753, 30.520132, 30.57796 , 31.633507, 33.25373 ,
33.95855 , 34.59395 , 34.78158 , 37.150833, 39.25521 ,
41.129295, 41.7405 , 44.94166 , 45.37275 , 45.924587,
46.66275 , 47.932114, 50.2065 , 50.939945, 51.17362 ,
55.16559 , 61.201077, 64.03338 , 68.46635 , 72.37863 ,
74.07547 , 75.993774, 78.803276, 82.673836, 85.39994 ,
87.995384, 88.03068 , 93.32808 , 94.61199 , 97.61955 ,
96.57511 , 91.99988 , 83.67492 , 79.224365, 72.62946 ,
66.08186 , 62.183556, 56.496174, 51.15098 , 44.65614 ,
41.01253 , 36.966286, 32.43728 , 28.903612, 24.055302,
18.436434])
richHigh_w = jnp.array([ 5. , 44.169 , 39.188175 , 37.95716 ,
37.479176 , 38.057766 , 39.403027 , 41.0048 ,
43.630054 , 46.80984 , 51.52312 , 56.437943 ,
60.986717 , 63.368095 , 68.02622 , 75.10258 ,
78.859886 , 79.52833 , 83.09218 , 81.95796 ,
85.69704 , 85.47596 , 88.454575 , 91.17749 ,
95.12584 , 95.58575 , 99.408165 , 102.66816 ,
103.02038 , 107.9051 , 107.72032 , 111.8956 ,
116.7536 , 118.76333 , 121.8654 , 123.99123 ,
124.62809 , 122.46186 , 119.743484 , 115.05782 ,
110.030136 , 106.00019 , 102.1959 , 96.18025 ,
86.68821 , 77.530266 , 63.242558 , 50.330845 ,
39.58192 , 31.540968 , 24.79612 , 19.6335 ,
15.757279 , 12.767186 , 11.137127 , 10.1711855,
9.303571 , 8.688891 , 8.259497 , 7.59842 ,
17.715118 ])
richLow_w = jnp.array([ 5. , 44.169 , 42.858627 , 43.603024 ,
49.520683 , 53.506363 , 59.840458 , 64.150024 ,
68.95748 , 73.39583 , 78.67845 , 84.01723 ,
81.94015 , 81.26681 , 88.00493 , 84.772964 ,
90.30113 , 91.04693 , 92.51674 , 97.758766 ,
100.51319 , 103.45857 , 104.626564 , 106.56832 ,
110.73649 , 111.43963 , 117.13121 , 119.765076 ,
123.79866 , 122.705536 , 124.00465 , 131.17126 ,
134.83432 , 136.05646 , 135.94238 , 130.88701 ,
127.932274 , 126.37201 , 120.503525 , 114.96932 ,
109.82672 , 107.14074 , 102.168816 , 93.78776 ,
83.32689 , 71.685265 , 60.297577 , 48.106426 ,
38.676346 , 31.609468 , 24.474401 , 19.16532 ,
14.775312 , 11.450411 , 9.281223 , 7.759332 ,
7.2246366, 6.4535017, 5.716505 , 5.1355343,
9.695507 ])
# +
# %pylab inline
matplotlib.rcParams['figure.figsize'] = [16, 8]
plt.rcParams.update({'font.size': 15})
plt.title("wealth level at different age periods")
plt.plot(poorHigh_w, label = "poor high cost")
plt.plot(poorLow_w, label = "poor low cost")
plt.plot(richHigh_w, label = "rich high cost")
plt.plot(richLow_w, label = "rich low cost")
plt.legend()
# -
| 20220122/shutDownRetirement/table.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 7.5.1
# language: ''
# name: sagemath
# ---
#import jdk2py_MD as my
from jdk2py_MD import *
p,q,r,s,t,u,w=var("p,q,r,s,t,u,w")
#Ejercicio 2.a
tabla=[["p","q","r","s:(p xor q)", "lado izquierdo t:(s xor r)", "u:(q xor r)", "lado derecho v:(p xor q)", "t iff v"]]
for p in [1,0]:
for q in [1,0]:
for r in [1,0]:
#lado izquierdo
s=xor(p,q)
t=xor(s,r)
#lado derecho
u=xor(q,r)
v=xor(p,u)
renglon = [p,q,r,s,t,u,v,iff(t,v)]
tabla.append(renglon)
show(table(tabla, header_row=True))
#Ejercicio 2.b
tabla=[["p","q","r","s=imply(q,r)", "lado izquierdo t=xor(p,s)", "u=xor(p,q)", "v=xor(p,r)", "lado derecho w=imply(u,v)", "t iff w"]]
for p in [1,0]:
for q in [1,0]:
for r in [1,0]:
#lado izquierdo
s=imply(q,r)
t=xor(p,s)
#lado derecho
u=xor(p,q)
v=xor(p,r)
w=imply(u,v)
renglon = [p,q,r,s,t,u,v,w,iff(t,w)]
tabla.append(renglon)
show(table(tabla, header_row=True))
#Ejercicio 4.b
tabla=[["p","q","r","s = p and r", "t = q and r", "u = p and not r", "v = q and not r", "w = iff(s,t)", "x = iff(u,v)",
"y = w and x", "z = iff(p,q)", "imply(y,z)"]]
for p in [1,0]:
for q in [1,0]:
for r in [1,0]:
s = p and r
t = q and r
u = p and not r
v = q and not r
w = iff(s,t)
x = iff(u,v)
y = w and x
z = iff(p,q)
renglon=[p,q,r,s,t,u,v,w,x,y,z, imply(y,z)]
tabla.append(renglon)
show(table(tabla, header_row=True))
#Ejercicio 5.a
tabla=[["p","q","r","s=xor(p,q)", "t=xor(p,r)", "u=iff(s,t)", "v=iff(q,r)", "imply(u,v)"]]
for p in [1,0]:
for q in [1,0]:
for r in [1,0]:
#condición suficiente
s=xor(p,q)
t=xor(p,r)
u=iff(s,t)
#condición necesaria
v=iff(q,r)
renglon=[p,q,r,s,t,u,v,imply(u,v)]
tabla.append(renglon)
show(table(tabla, header_row=True))
#Ejercicio 5.b
tabla=[["p","q","r","s=xor(q,r)", "t=(p and s)", "u=(p and q)", "v=(p and r)", "w=xor(u,v)", "iff(t,w)"]]
for p in [1,0]:
for q in [1,0]:
for r in [1,0]:
#condición suficiente
s=xor(q,r)
t=(p and s)
u=(p and q)
#condición necesaria
v=(p and r)
w=xor(u,v)
renglon=[p,q,r,s,t,u,v,w,iff(t,w)]
tabla.append(renglon)
show(table(tabla, header_row=True))
| latex/MD_SMC/MD01_Primer_Parcial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
from nb_005 import *
# # STL-10
# ## Basic data aug
PATH = Path('data/stl10')
data_mean, data_std = map(tensor, ([0.485, 0.456, 0.406], [0.229, 0.224, 0.225]))
data_norm,data_denorm = normalize_funcs(data_mean,data_std)
train_ds = FilesDataset.from_folder(PATH/'train')
valid_ds = FilesDataset.from_folder(PATH/'valid')
x=Image(valid_ds[0][0])
x.show()
x.shape
size=96
tfms = get_transforms(do_flip=True, max_rotate=10, max_lighting=0.2, max_warp=0.15, max_zoom=1.2)
# tfms = get_transforms(do_flip=True, max_rotate=10, max_lighting=0.2)
tds = transform_datasets(train_ds, valid_ds, tfms, size=size)#, padding_mode='zeros')
data = DataBunch(*tds, bs=32, num_workers=8, tfms=data_norm)
# +
(x,y) = next(iter(data.valid_dl))
_,axs = plt.subplots(4,4,figsize=(12,12))
for i,ax in enumerate(axs.flatten()): show_image(data_denorm(x[i].cpu()), ax)
# +
(x,y) = next(iter(data.train_dl))
_,axs = plt.subplots(4,4,figsize=(12,12))
for i,ax in enumerate(axs.flatten()): show_image(data_denorm(x[i].cpu()), ax)
# -
_,axs = plt.subplots(4,4,figsize=(12,12))
for i,ax in enumerate(axs.flat): show_image(tds[0][1][0], ax)
# ## Train
from torchvision.models import resnet18, resnet34, resnet50
arch = resnet50
lr = 5e-3
# +
def _set_mom(m, mom):
if isinstance(m, bn_types): m.momentum=mom
def set_mom(m, mom): m.apply(lambda x: _set_mom(x, mom))
def set_bn_train(l, b):
if isinstance(l, bn_types):
for p in l.parameters(): p.requires_grad = b
# +
learn = ConvLearner(data, arch, 2 , wd=1e-1 #, train_bn=False #, callback_fns=[BnFreeze]
# ,dps=[0.01,0.02])
, opt_fn=partial(optim.SGD, momentum=0.9))
learn.metrics = [accuracy]
learn.split(lambda m: (m[0][6], m[1]))
learn.freeze()
# -
lr_find(learn)
learn.recorder.plot()
# +
# learn.model.apply(partial(set_bn_train, b=True));
# +
# set_mom(learn.model[0], 0.01)
# -
lrs = np.array([lr/9, lr/3, lr])
# +
# TODO min_lr/max_lr
# -
learn.fit(6, lrs)
learn.save('0')
# ## Gradual unfreezing
learn.load('0')
learn.unfreeze()
learn.fit(3, lrs)
learn.fit(10, lrs)
learn.save('1')
learn.recorder.plot_losses()
# ## Fin
import pandas as pd
csv = pd.read_csv(PATH/'default.csv')
is_valid = csv['2']=='valid'
valid_df,train_df = csv[is_valid],csv[~is_valid]
len(valid_df),len(train_df)
len(valid_ds)
# +
train_fns,train_lbls,valid_fns,valid_lbls = map(np.array,
(train_df['0'],train_df['1'],valid_df['0'],valid_df['1']))
train_fns = [PATH/o for o in train_fns]
valid_fns = [PATH/o for o in valid_fns]
train_ds = FilesDataset(train_fns,train_lbls)
valid_ds = FilesDataset(valid_fns,valid_lbls, classes=train_ds.classes)
# -
| dev_nb/stl-10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="-wFUjZTmcG5a"
# # Library loading and restoring of CNN weights
#
# First we load the library and restore the weights.
# + colab={"base_uri": "https://localhost:8080/", "height": 405} colab_type="code" executionInfo={"elapsed": 97482, "status": "error", "timestamp": 1600625422711, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiEI-GAxszORnjA0WqbcKrAT2dFjghG7ikD4zjctA=s64", "userId": "05156771066106099172"}, "user_tz": -120} id="0xLkmPMibmpd" outputId="37401fce-48c8-42cb-eadc-cc4f7e2c035e"
# TensorFlow and tf.keras
import tensorflow as tf
from tensorflow import keras
import numpy as np
import matplotlib.pyplot as plt
print(tf.__version__)
from google.colab import drive
drive.mount('/content/drive')
model = tf.keras.models.load_model('/content/drive/My Drive/modul_2_cnn/model')
# + [markdown] colab_type="text" id="jE7Uqzd8fkzm"
# # Testing
# We have already classified the nine correctly in module 2. Let's check if this still works.
# + colab={"base_uri": "https://localhost:8080/", "height": 298} colab_type="code" executionInfo={"elapsed": 1289, "status": "ok", "timestamp": 1600017978980, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiEI-GAxszORnjA0WqbcKrAT2dFjghG7ikD4zjctA=s64", "userId": "05156771066106099172"}, "user_tz": -120} id="EJv19cyoeu9r" outputId="934296d7-79f4-4540-bf84-a3b6578458c5"
# We test for the first time on our old data to test if the loading worked.
mnist = keras.datasets.mnist
(_, _), (test_images, test_labels) = mnist.load_data()
test_images = np.expand_dims(test_images / 255.0, -1)
# Load a 0 from the testing set
indicies_of_0 = (np.where(test_labels == 0))[0]
image_with_0 = test_images[indicies_of_0[0]]
plt.figure()
plt.imshow(image_with_0[:,:,0], cmap=plt.cm.binary)
plt.title("This is a 0.")
plt.show()
from scipy.special import softmax
logits_of_zero = model.predict(np.expand_dims(image_with_0, 0))
probabilities_of_zero = softmax(logits_of_zero)[0]
detected_class_of_zero = np.argmax(probabilities_of_zero)
print('The NN classified the 0 as ', detected_class_of_zero, ' with a probability of ', probabilities_of_zero[detected_class_of_zero])
# + [markdown] colab_type="text" id="95Nf2F41f25S"
# # Own digit
# Can the network also recognize our own digit? For this we have to load the digit first, visualize it and see if the NN recognizes it :-)
# + colab={"base_uri": "https://localhost:8080/", "height": 334} colab_type="code" executionInfo={"elapsed": 1265, "status": "ok", "timestamp": 1600017988164, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiEI-GAxszORnjA0WqbcKrAT2dFjghG7ikD4zjctA=s64", "userId": "05156771066106099172"}, "user_tz": -120} id="gRO4xoDEgDPI" outputId="426de91c-82fb-4a48-d015-fb7442f7a818"
my_image = plt.imread('/content/drive/My Drive/zwei.png')
print('Das Bild hat folgende Dimensionen ', my_image.shape) # png images are stored as RGBA --> but we requrie a grayscale image with 28x28x1
# take mean values of image for grayscale
my_gray_image = np.mean(my_image[:,:,:3], axis=-1) # 28x28
my_gray_image = np.expand_dims(my_gray_image, axis=-1)
print('The grayscale image has following dimenson ', my_gray_image.shape)
plt.figure()
plt.imshow(my_gray_image[:,:,0], cmap=plt.cm.binary)
plt.title("My Digit")
plt.show()
logits = model.predict(np.expand_dims(my_gray_image, axis=0))
probabilities = softmax(logits)[0]
detected_class = np.argmax(probabilities)
print('My image is classified as ', detected_class, ' with a probability of ', probabilities[detected_class])
# + [markdown] colab_type="text" id="StpMtRLiQ3xs"
# # Task Data Formats
#
#
# * How important is it to show the network the same data it already saw during training?
# * What does RGB mean and what is a gray scale image?
| english_version/modul_3_reuse_cnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <center><h1> <font color="#2d97c4"> Check that Data is Packed correctly, all sessions are accounted for, and no data is currapted or missed aligned </font> </h1> </center>
#since noteboke doesn't work in jupiterlabs %matplotlib notebook
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
import os
os.chdir('/home/dana_z/ssd_2TB/6OHDA')
#import mpld3
#mpld3.enable_notebook()
import numpy as np
import scipy as sci
from scipy import signal
from matplotlib import pyplot as plt
from matplotlib import gridspec
import matplotlib.colors as Mcolors
import matplotlib.cm as cmx
import sys
import h5py
from IO import *
from utils import *
from plotUtils import *
from ColorSchems import colorPallet as CP
import pptx
from pptx import Presentation
from pptx.util import Inches
from io import BytesIO
import re
import warnings
import pandas as pd
import sqlalchemy as db
import gc
Files = ['FinalData_6OHDA.h5','FinalData_6OHDA_H.h5','FinalData_6OHDA_H_skip.h5','FinalData_6OHDA_skip.h5']
# <b> Make sure all sessions are in the struct </b>
Sess = {}
for dataFile in Files:
Sess[dataFile] = getSessionList(dataFile)
print(dataFile,':',len(Sess[dataFile]))
# <b> <span style="color:red;"> Missing sessions:</span> </b> <br />
# <strike>1 -unknown<br />
# 1253_baselineS <br />
# 1793_day34L <br />
# 4539_BaselineA <br />
# 7909_BaselineA2 <br />
# ---------------------------------------------------------------------
# <b> check which session miss partial data, how many TD tomato cells are in each session, and how many skipped cell in each session </b>
df = pd.DataFrame(columns = ['File','Session','missing_traces',
'missing_mvmt','missing_lfp','numRed','num_skip','creType'])
# +
data = []
for dataFile in Files:
sessions = Sess[dataFile]
skiped = dataFile.find('skip')!= -1
lfps = getData(dataFile,['lfp'])
lfps = list(lfps.keys())
mvmt = getData(dataFile,['mvmt'])
mvmt = list(mvmt.keys())
dff = getData(dataFile,['trace'])
dff = list(dff.keys())
for s in sessions:
d ={'File':dataFile,'Session':s}
m = s[0:4]
d['numRed'] = getNumRed(dataFile,m,s[5:])
d['missing_traces'] = s not in dff
if skiped and not d['missing_traces']:
d['numSkip'] = np.sum(getSkipList(dataFile,m,s[5:]))
d['creType'] = getCreType(dataFile,m)
d['missing_lfp'] = not s in lfps
d['missing_mvmt'] = not s in mvmt
data.append(d)
df = pd.DataFrame(data)
# +
user = 'auto_processing'
password = '<PASSWORD>'
engine = db.create_engine('mysql+pymysql://'+user+':'+password+'@localhost/preProcess')
df.to_sql('PackedData',engine,index =False,if_exists= 'replace')
# -
df.groupby('File').sum()
df['mouse'] = df.apply(lambda row: row.Session[0:4],axis=1)
pd.pivot_table(df[(df.missing_lfp>0) | (df.missing_mvmt>0) | (df.missing_traces>0)] ,
values='numSkip', index=['File','Session','missing_lfp','missing_mvmt','missing_traces'], aggfunc=np.sum)
# <b> 1208_day12 currupted LFP session, <br>
# 2976_day4 only 30s of LFP recorded </b>
pd.pivot_table(df[(df.File =='FinalData_6OHDA_H_skip.h5') & (df.numSkip>0)] ,
values='numSkip', index=['File', 'mouse'], columns=['creType'], aggfunc=np.sum)
pd.pivot_table(df[(df.File =='FinalData_6OHDA_H_skip.h5') & (df.numSkip>0) & (df.mouse == '7909')] ,
values='numSkip', index=['File', 'Session'], columns=['creType'], aggfunc=np.sum)
# <b> look at all traces that are marked as TD-tomato + skip </b>
# +
# global presentation
Oprs = Presentation()
title_layout = Oprs.slide_layouts[5]
title_slide_layout = Oprs.slide_layouts[0]
slide = Oprs.slides.add_slide(title_slide_layout)
slide.shapes.title.text = 'Skipped TD-tomato cells - per mouse'
# position, size, and colors:
lf= {'left':0.00, 'top':1.20, 'height':5.80, 'width':10.00}
rawArgs = {'left':Inches(lf['left']),'top':Inches(lf['top']), 'height':Inches(lf['height']), 'width':Inches(lf['width'])}
miceList = getMiceList(Files[2])
for m in miceList:
data = getData(Files[2],['trace'],period ='Pre', red=True, mice=m)
days = np.zeros(len(data))
ind = 0
for sess in data:
#store max min mean median
if sess[5] == 'B':
day = 0
else:
day = int(re.findall(r'\d+',sess[5:])[0])
days[ind] = day
ind= ind+1
a = np.argsort(days)
dKeys = list(data.keys())
for aa in range(0,len(data)):
sess = dKeys[a[aa]]
dff = data[sess]['trace']['dff']
numred = data[sess]['trace']['numred']
skiped = getSkipList(Files[2],m,sess[5:])
skiped = skiped[:numred]
if np.sum(skiped) == 0:
continue
else:
slide = Oprs.slides.add_slide(title_layout)
slide.shapes.title.text = sess
dff = dff[skiped.astype('bool'),:]
dt = 1/data[sess]['trace']['FS']
fig, ax = plt.subplots(1,1,figsize=(lf['width'],lf['height']))
rosterPlot(ax, dff,dt,specing = np.max(dff), Color = None)
pic = plt2pptx(slide, fig, **rawArgs)
fig.clf()
plt.close(fig)
Oprs.save('ppts/skiiped_TDtomato.pptx')
# -
# <b> Make sure lfp is not currupted (spectron looks reasonable) </b> <br />
# Store all sessions in ppt - so can look at each session individually later
# +
# global presentation handling:
Oprs = Presentation() # store overall (1 slide/mouse)
Iprs = Presentation() # store individual sessions
title_layout = Oprs.slide_layouts[5]
title_slide_layout = Oprs.slide_layouts[0]
slide = Oprs.slides.add_slide(title_slide_layout)
slide.shapes.title.text = 'lfp summary - per mouse'
slide = Iprs.slides.add_slide(title_slide_layout)
slide.shapes.title.text = 'lfp summary - per session'
# global color scheme and positions:
lf= {'left':0.64, 'top':1.85, 'height':2.07, 'width':8.25}
sf= {'left':0.64, 'top':4.4, 'height':2.07, 'width':8.25}
cf = {'left':1.35, 'top':1.46, 'height':5.58, 'width':7.14}
rawArgs = {'left':Inches(lf['left']),'top':Inches(lf['top']), 'height':Inches(lf['height']), 'width':Inches(lf['width'])}
specArgs = {'left':Inches(lf['left']),'top':Inches(sf['top']), 'height':Inches(lf['height']), 'width':Inches(lf['width'])}
sumArgs = {'left':Inches(cf['left']),'top':Inches(cf['top']), 'height':Inches(cf['height']), 'width':Inches(cf['width'])}
# create the color maps
cNorm = Mcolors.Normalize(vmin=1, vmax=35)
cm = plt.get_cmap('YlOrRd')
cMap = cmx.ScalarMappable(norm=cNorm, cmap = cm)
miceList = getMiceList(Files[0])
for m in miceList:
data = getData(Files[0],['lfp'],period ='Pre', mice=m)
figt, axt = plt.subplots(1,1,figsize=(cf['width'],cf['height']))
figt.set_size_inches(cf['width'],cf['height'],forward=True)
days = np.zeros(len(data))
ind = 0
for sess in data:
#store max min mean median
if sess[5] == 'B':
day = 0
else:
day = int(re.findall(r'\d+',sess[5:])[0])
days[ind] = day
ind= ind+1
a = np.argsort(days)
dKeys = list(data.keys())
for aa in range(0,len(data)):
# try:
sess = dKeys[a[aa]]
slide = Iprs.slides.add_slide(title_layout)
slide.shapes.title.text = sess
lfp = data[sess]['lfp']['lfp']
Fs = data[sess]['lfp']['FS']
# plot raw lfp:
fig, ax = plt.subplots(1,1,figsize=(lf['width'],lf['height']))
ax.plot(lfp)
fig.set_size_inches(lf['width'],lf['height'], forward=True)
pic = plt2pptx(slide, fig, **rawArgs)
fig.clf()
plt.close(fig)
# plot spectogram:
f, t, Sxx = signal.spectrogram(lfp[:,0],Fs,window=('hamming'),nperseg=140*8,noverlap =120*8,nfft=1200*8)
Pxx = 10*np.log10(np.abs(Sxx))
Pxx[np.isinf(Pxx)] = 0
tlfp = np.linspace(0,lfp.size*(1/Fs),lfp.size)
fig, ax = plt.subplots(1,1,figsize=(lf['width'],lf['height']))
fig.set_size_inches(lf['width'],lf['height'], forward=True)
ind = np.searchsorted(f,100)
ax.pcolormesh(t,f[:ind],Pxx[:ind,:],vmin=-170,vmax=-70, cmap='jet')
ax.set_ylim((5,100))
pic = plt2pptx(slide, fig, **specArgs)
fig.clf()
plt.close(fig)
# plot spectrom in the right color on fig_t
if sess[5] == 'B':
day = 0
colorVal = 'green'
else:
day = int(re.findall(r'\d+',sess[5:])[0])
colorVal = cMap.to_rgba(day)
Power = np.sum(Sxx[:ind,:],1)
totPower = np.sum(Power)
if totPower == 0:
totPower = 1
M = Power/totPower
axt.plot(f[:ind],M,color = colorVal, label = str(day))
axt.set_xlim((5,100))
del f
gc.collect()
# except:
# print(m,sess)
# continue
slide = Oprs.slides.add_slide(title_layout)
slide.shapes.title.text = m
handles,labels = axt.get_legend_handles_labels()
axt.legend(handles, labels, loc='upper right')
pic = plt2pptx(slide, figt, **sumArgs)
figt.clf()
plt.close(figt)
Iprs.save('ppts/lfp_individual_'+m+'.pptx')
Oprs.save('ppts/lfp_Overall_Mice.pptx')
# -
# <b> <span style="color:red;">Currapted lfp sessions:</span> </b> <br />
# <strike> 1208_day12 --> all session </strike> session removed <br />
# 1236_day30A --> ~30s-180s <br />
# 1236_day35L --> ~30s-300s <br />
#
# <b> <span style="color:red;">Short/missing lfp sessions:</span> </b> <br />
# 2976_day4 <br />
#
# <b> <span style="color:red;">Excessive outliers in lfp:</span> </b> <br />
# 2981_day15A - many <br />
# 8803_day10 - 1 outlier <br />
# 8815_day19L - 1 outlier <br />
# +
# define presentation params:
prs = Presentation()
prs.slide_width = Inches(11)
title_layout = prs.slide_layouts[5]
title_slide_layout = prs.slide_layouts[0]
slide = prs.slides.add_slide(title_slide_layout)
slide.shapes.title.text = 'Mvmt onset'
# define figure params:
lf = {'left':0.30, 'top':1.30, 'height':5.80, 'width':10.10}
fArgs = {'left':Inches(lf['left']),'top':Inches(lf['top']), 'height':Inches(lf['height']), 'width':Inches(lf['width'])}
Colors = CP('mvmtType')
th = 2
hi = 9
hiWin=40
thWin=30
shift=3
# get mice list:
miceList = getMiceList(Files[0])
# prepare data storage for segments:
# make plot and save as ppt
for m in miceList:
data = getData(Files[0],['speed'],period ='Pre', mice=m)
days = np.zeros(len(data))
ind = 0
# sort by session for my own OCD
for sess in data:
if sess[5] == 'B':
day = 0
else:
day = int(re.findall(r'\d+',sess[5:])[0])
days[ind] = day
ind= ind+1
a = np.argsort(days)
dKeys = list(data.keys())
# calculte high speed period, do 3 sessions per plot, and stor in ppt
ind = 0;
for aa in range(0,len(data)):
sess = dKeys[a[aa]]
speed = data[sess]['speed']['speed']
speed = speed.T
smoothSpeed = smooth(speed,20)
dt = 1/data[sess]['speed']['Fs']
if ind%3==0:
fig, ax = plt.subplots(3,1,figsize=(lf['width'],lf['height']),
gridspec_kw = {'top':0.995,'bottom':0.008,'wspace':0.1})
fig.set_size_inches(lf['width'],lf['height'],forward=True)
fig.subplots_adjust(left=0.03, right=0.99)
slide = prs.slides.add_slide(title_layout)
slide.shapes.title.text = m + 'params: th='+ str(th) + ' hi='+ str(hi)
try:
sOnset = FindMvmtOnset(speed,th,hi,hiWin,thWin, shift)
t = np.linspace(0,len(speed)*dt,len(speed))
ax[ind%3].plot(t,speed)
ax[ind%3].plot(t,smoothSpeed, color='black')
ax[ind%3].plot(t[sOnset],smoothSpeed[sOnset],'X',color='firebrick')
ax[ind%3].set_xlim(0,600)
except:
print('error')
ax[ind%3].set_title(sess)
if ind%3==2 or aa ==len(data)-1:
pic = plt2pptx(slide, fig, **fArgs)
plt.close(fig)
ind = ind+1
prs.save('ppts/SpeedOnset_final.pptx')
# -
# <b> <font color="#2d97c4"> Check Movement onset and high/low speed </font> </b> <br />
# since mvmt onset not ideal for all mice, change policty to reflect parameters based on mice baseline sessions<br />
# <br />
#
# Recipee: <br />
# <b> 1) </b> look at 3 baseline session for mouse and detrmine speed statistics <br />
# <b> 2) </b> from 1, automatically chooce params for speed onset <br />
# <b> 3) </b> use params from 2 to find mvmt onset for all session and store in hdf5 dataset <br />
# <b> 4) </b> implement an i/o function that load speed onset for session <br />
# +
# visualize Baseline speed range, quantiles, mean, median, std for all mice.
miceList = getMiceList(Files[0])
speedData = []
for m in miceList:
data = getData(Files[0],['speed'],period ='Pre',mice=m, day = lambda x: x==0)
for sess in data:
speed = data[sess]['speed']['speed']
d ={'Session':sess[5:], 'Mouse':m,'min':np.min(speed),'max':np.max(speed),
'std':np.std(speed),'mean':np.mean(speed)}
Q = np.quantile(speed,[.25,.5,.75])
d['Q25'],d['Q50'],d['Q75'] =np.quantile(speed,[.25,.5,.75])
d['Q25_std'] = np.std(speed[np.where(speed<=d['Q25'])])
speedData.append(d)
df = pd.DataFrame(speedData)
df.to_sql('SpeedData',engine,index =False,if_exists= 'replace')
# -
miceList = getMiceList(Files[0])
speedOnsetPars = {}
for m in miceList:
data = getData(Files[0],['speed'],period ='Pre',mice=m, day = lambda x: x==0)
maxSpeed = [];
for sess in data:
speed = smooth(data[sess]['speed']['speed'],20)
maxSpeed.append(np.max(speed))
hi = np.mean(maxSpeed)/4
print(m,' hi: ',hi)
speedOnsetPars[m] = hi
# +
# define presentation params:
prs = Presentation()
prs.slide_width = Inches(11)
title_layout = prs.slide_layouts[5]
title_slide_layout = prs.slide_layouts[0]
slide = prs.slides.add_slide(title_slide_layout)
slide.shapes.title.text = 'Mvmt onset'
# define figure params:
lf = {'left':0.30, 'top':1.30, 'height':5.80, 'width':20.10}
fArgs = {'left':Inches(lf['left']),'top':Inches(lf['top']), 'height':Inches(lf['height']), 'width':Inches(lf['width'])}
Colors = CP('mvmtType')
hiWin=20
thWin=40
th_strong = 1
shift=2
# get mice list:
miceList = getMiceList(Files[0])
# prepare data storage for segments:
# make plot and save as ppt
for m in miceList:
data = getData(Files[0],['speed'],period ='Pre', mice=m)
days = np.zeros(len(data))
ind = 0
# sort by session for my own OCD
for sess in data:
if sess[5] == 'B':
day = 0
else:
day = int(re.findall(r'\d+',sess[5:])[0])
days[ind] = day
ind= ind+1
a = np.argsort(days)
dKeys = list(data.keys())
# calculte high speed period, do 3 sessions per plot, and stor in ppt
ind = 0;
hi = speedOnsetPars[m]
th_weak = np.min([3.3, hi/2.5])
for aa in range(0,len(data)):
sess = dKeys[a[aa]]
speed = data[sess]['speed']['speed']
speed = speed.T
smoothSpeed = smooth(speed,20)
dt = 1/data[sess]['speed']['Fs']
if ind%3==0:
fig, ax = plt.subplots(3,1,figsize=(lf['width'],lf['height']),
gridspec_kw = {'top':0.995,'bottom':0.008,'wspace':0.1})
fig.set_size_inches(lf['width'],lf['height'],forward=True)
fig.subplots_adjust(left=0.03, right=0.99)
slide = prs.slides.add_slide(title_layout)
slide.shapes.title.text = m + 'params: th_weak='+ str(round(th_weak,2)) + ' hi='+ str(round(hi,2))
try:
sOnset = FindMvmtOnset2(speed, th_weak,th_strong ,hi,hiWin,thWin,shift)
t = np.linspace(0,len(speed)*dt,len(speed))
ax[ind%3].plot(t,speed)
ax[ind%3].plot(t,smoothSpeed, color='black')
ax[ind%3].plot(t[sOnset],smoothSpeed[sOnset],'X',color='firebrick')
ax[ind%3].set_xlim(0,600)
except:
print('error')
ax[ind%3].set_title(sess)
if ind%3==2 or aa ==len(data)-1:
pic = plt2pptx(slide, fig, **fArgs)
plt.close(fig)
ind = ind+1
prs.save('ppts/SpeedOnset_Final3.pptx')
# -
# <b> Document final decision on mvmt onset algo, <br />
# pack into an hdf5 file, <br />
# and write an I/O function to load them.
#
# hiWin=20 <br />
# thWin=40 <br />
# th_strong = 1 <br />
# shift=2 <br />
#
# hi = np.mean(maxSpeed)/4 where max speed is smooth with 1s rolling window for all 3 baseline sessions <br />
# th_weak = np.min([3.3, hi/2.5])
# +
# create hdf5 file with all the mice and mvmt Onset
f = h5py.File('OnsetsAndPeriods.hdf5','a')
hiWin=20
thWin=40
th_strong = 1
shift=2
miceList = speedOnsetPars.keys()
for m in miceList:
grp = f.create_group(m)
sgrp = grp.create_group('mvmtOnset_params')
sgrp.attrs['hi'] = speedOnsetPars[m]
sgrp.attrs['th_weak'] = np.min([3.3, speedOnsetPars[m]/2.5])
sgrp.attrs['hiWin'] = hiWin
sgrp.attrs['thWin'] = thWin
sgrp.attrs['th_strong'] = th_strong
sgrp.attrs['shift'] = shift
data_pre = getData(Files[0],['speed'],period ='Pre', mice=m)
data_post = getData(Files[0],['speed'],period ='Post', mice=m)
days = np.zeros(len(data_pre))
ind = 0
# sort by session for my own OCD
for sess in data_pre:
if sess[5] == 'B':
day = 0
else:
day = int(re.findall(r'\d+',sess[5:])[0])
days[ind] = day
ind= ind+1
a = np.argsort(days)
dKeys = list(data_pre.keys())
# calculte high speed period, do 3 sessions per plot, and stor in ppt
ind = 0;
hi = sgrp.attrs['hi']
th_weak = sgrp.attrs['th_weak']
for aa in range(0,len(data_pre)):
sess = dKeys[a[aa]]
speed = data_pre[sess]['speed']['speed']
speed = speed.T
sOnset = FindMvmtOnset2(speed, th_weak,th_strong ,hi,hiWin,thWin,shift)
subgrp = grp.create_group(sess)
ssubgrp = subgrp.create_group('Pre')
ssubgrp['mvmtOnset'] = sOnset
if sess in data_post.keys():
speed = data_post[sess]['speed']['speed']
speed = speed.T
sOnset = FindMvmtOnset2(speed, th_weak,th_strong ,hi,hiWin,thWin,shift)
spsubgrp = subgrp.create_group('Post')
spsubgrp['mvmtOnset'] = sOnset
f.close()
# +
def getOnsetOrPeriod(m,s,period,OPtype,fileName='OnsetsAndPeriods.hdf5'):
# takes in a file name and return all the mice that exsits in file
with h5py.File(fileName,'r') as hf:
if m+'/'+s+'/'+period+'/'+ OPtype not in hf:
print(m+'/'+s+'/'+period+'/'+ OPtype +' NOT in FILE')
return []
else:
return hf[m][s][period][OPtype].value
# -
# <b> double check that speed ONSET was saved properly, IO function works, and th make sense for post infusion data </b>
# +
# define presentation params:
prs = Presentation()
prs.slide_width = Inches(11)
title_layout = prs.slide_layouts[5]
title_slide_layout = prs.slide_layouts[0]
slide = prs.slides.add_slide(title_slide_layout)
slide.shapes.title.text = 'Mvmt onset'
# define figure params:
lf = {'left':0.30, 'top':1.30, 'height':5.80, 'width':20.10}
fArgs = {'left':Inches(lf['left']),'top':Inches(lf['top']), 'height':Inches(lf['height']), 'width':Inches(lf['width'])}
Colors = CP('mvmtType')
# get mice list:
miceList = getMiceList(Files[0])
# prepare data storage for segments:
# make plot and save as ppt
for m in miceList:
data = getData(Files[0],['speed'],period ='Pre', mice=m)
data_post = getData(Files[0],['speed'],period ='Post', mice=m)
days = np.zeros(len(data))
ind = 0
# sort by session for my own OCD
for sess in data:
if sess[5] == 'B':
day = 0
else:
day = int(re.findall(r'\d+',sess[5:])[0])
days[ind] = day
ind= ind+1
a = np.argsort(days)
dKeys = list(data.keys())
# calculte high speed period, do 3 sessions per plot, and stor in ppt
ind = 0;
for aa in range(0,len(data)):
sess = dKeys[a[aa]]
speed = data[sess]['speed']['speed']
speed = speed.T
smoothSpeed = smooth(speed,20)
dt = 1/data[sess]['speed']['Fs']
if ind%3==0:
fig, ax = plt.subplots(3,1,figsize=(lf['width'],lf['height']),
gridspec_kw = {'top':0.995,'bottom':0.008,'wspace':0.1})
fig.set_size_inches(lf['width'],lf['height'],forward=True)
fig.subplots_adjust(left=0.03, right=0.99)
slide = prs.slides.add_slide(title_layout)
slide.shapes.title.text = m #+ 'params: th_weak='+ str(round(th_weak,2)) + ' hi='+ str(round(hi,2))
try:
sOnset = getOnsetOrPeriod(m,sess,'Pre','mvmtOnset')
t = np.linspace(0,len(speed)*dt,len(speed))
ax[ind%3].plot(t,speed)
ax[ind%3].plot(t,smoothSpeed, color='black')
ax[ind%3].plot(t[sOnset],smoothSpeed[sOnset],'X',color='firebrick')
ax[ind%3].set_xlim(0,600)
ax[ind%3].set_title(sess)
if sess in data_post.keys():
pic = plt2pptx(slide, fig, **fArgs)
plt.close(fig)
fig, ax = plt.subplots(3,1,figsize=(lf['width'],lf['height']),
gridspec_kw = {'top':0.995,'bottom':0.008,'wspace':0.1})
fig.set_size_inches(lf['width'],lf['height'],forward=True)
fig.subplots_adjust(left=0.03, right=0.99)
slide = prs.slides.add_slide(title_layout)
slide.shapes.title.text = m +sess +'_post'#+ 'params: th_weak='+ str(round(th_weak,2)) + ' hi='+ st
sOnset = getOnsetOrPeriod(m,sess,'Post','mvmtOnset')
speed = data_post[sess]['speed']['speed']
speed = speed.T
smoothSpeed = smooth(speed,20)
t = np.linspace(0,len(speed)*dt,len(speed))
for ind in range(3):
if ind ==2:
ax[ind%3].plot(t[ind*600:],speed[ind*600:])
else:
ax[ind%3].plot(t[ind*600:(ind+1)*600],speed[ind*600:(ind+1)*600])
ax[ind%3].plot(t,smoothSpeed, color='black')
ax[ind%3].plot(t[sOnset],smoothSpeed[sOnset],'X',color='firebrick')
pic = plt2pptx(slide, fig, **fArgs)
plt.close(fig)
except:
print('error')
if ind%3==2 or aa ==len(data)-1:
pic = plt2pptx(slide, fig, **fArgs)
plt.close(fig)
ind = ind+1
prs.save('ppts/SpeedOnset_check.pptx')
# -
| .ipynb_checkpoints/Data_Check-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#python3 t2_process.py --tof ts1_16x320_inv --ntile 16 --tsize 80
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
import os, gc, subprocess, time, sys, shutil, argparse
import scipy
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import matplotlib
import matplotlib.pyplot as plt
from PIL import Image
import cv2
from tqdm import tqdm
#from tqdm.notebook import tqdm
import skimage.io
from skimage.transform import resize, rescale
from math import sqrt
import statistics
sys.path.append("..")
# -
import PIL.Image
#PIL.Image.MAX_IMAGE_PIXELS = 400000000
PIL.Image.MAX_IMAGE_PIXELS = None
# +
# #%matplotlib inline
# -
from panda_bvv_config import *
# ## 1. Inputs ##
Zoom = 1
input_folder = train_path
input_mask_folder = mask_size1
# +
ap = argparse.ArgumentParser()
ap.add_argument('--tof', help="folder name to save tile", dest = 'tof',\
type=str)
args = vars(ap.parse_args())
# -
#data-from folder:
to_folder = args["tof"]
new_proc_folder = os.path.join(base_path, to_folder)
# wrapping inputs:
if not os.path.exists(new_proc_folder):
print("[INFO] 'creating {}' directory".format(new_proc_folder))
os.makedirs(new_proc_folder)
# ## 2. Standard functions ##
#wrapper for image processing function
def process_all_images(proc_func):
def wrapper(zoom = Zoom,
input_biopsy_folder = input_folder,
output_data_path = new_proc_folder,
df_name = train_labels,
tif_file = False,
with_mask = False,
input_mask_folder = input_mask_folder,
**kw):
print(input_biopsy_folder)
print(new_proc_folder)
with tqdm(total=df_name.shape[0]) as pbar:
for i, row in enumerate(tqdm(df_name.iterrows())):
img_num = row[1]['image_id']
provider = row[1]['data_provider']
mask = None
if tif_file:
try:
biopsy = skimage.io.MultiImage(os.path.join(input_biopsy_folder,\
img_num + '.tiff'))[zoom]
except:
print('Failed to read tiff:', img_num)
else:
try:
biopsy = skimage.io.imread(os.path.join(input_biopsy_folder, img_num + '.png'))
shape = biopsy.shape
if shape[0]<shape[1]:
biopsy = np.rot90(biopsy)
shape = biopsy.shape
except:
print(f'can not proceed with {img_num}')
if with_mask:
try:
mask = skimage.io.imread(os.path.join(input_mask_folder, img_num + '_mask.png'))
shape = mask.shape
if shape[0]<shape[1]:
mask = np.rot90(mask)
shape = mask.shape
except:
print('Failed to process mask:', img_num)
try:
data_new = proc_func(biopsy, mask, provider, **kw)
cv2.imwrite(os.path.join(output_data_path, img_num + '.png'), data_new)
except Exception as ee:
print('Processing mistake:\n', ee, '\n', img_num)
try:
del biopsy, mask, data_new, mask_new
except:
pass
pbar.update(1)
gc.collect()
return
return wrapper
# ## 3. Processing functions ##
@process_all_images
def tiff_to_png_size1(img, mask, provider, **kw):
return img
# ## 4. Process input files for training purpose ##
tiff_to_png_size1(tif_file = True,
df_name = train_labels)
# import os
# module_name = 't2_extract_size1'
#
# os.system('jupyter nbconvert --to python ' + module_name + '.ipynb')
# with open(module_name + '.py', 'r') as f:
# lines = f.readlines()
# with open(module_name + '.py', 'w') as f:
# for line in lines:
# if 'nbconvert --to python' in line:
# break
# else:
# f.write(line)
| cloud/t2_extract_size1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import warnings
warnings.filterwarnings('ignore')
# ### Data Description
desc_df = pd.read_csv('dataset/LCDataDictionary.csv')
pd.set_option("display.max_colwidth",-1)
desc_df[:60]
desc_df[60:]
# ### Loading Dataset
loan_df = pd.read_csv('dataset/lending_club_loans.csv', skiprows=1, low_memory=False)
loan_df.head()
loan_df.shape
# # Data Cleansing
# ### Missing Values
clean_df = loan_df.copy()
clean_df.head()
# +
# checking total null values in each columns
for cols in clean_df:
print(clean_df[cols].isna().sum())
# +
# columns, where null values are more than 38000
null_cols = []
for cols in clean_df:
if clean_df[cols].isna().sum() > 38000:
null_cols.append(cols)
# -
len(null_cols)
clean_df[null_cols]
clean_df.drop(columns=null_cols, axis=1, inplace=True)
clean_df.shape
clean_df.isna().sum()
clean_df['emp_title'].value_counts()
clean_df['emp_title'].isna().sum()
clean_df['emp_title'].fillna('Other', inplace=True)
clean_df['emp_title'].value_counts()
clean_df['emp_length'].value_counts()
clean_df['emp_length'].isna().sum()
clean_df['emp_length'].fillna('< 1 year', inplace=True)
clean_df['emp_length'].value_counts()
clean_df['desc']
clean_df['desc'].isna().sum()
clean_df['mths_since_last_delinq'].isna().sum()
clean_df['mths_since_last_delinq'].isna().sum()/clean_df.shape[0]
clean_df.drop(columns=['desc','url','mths_since_last_delinq'], axis=1, inplace=True)
clean_df.isna().sum()
clean_df['pub_rec_bankruptcies'].value_counts()
clean_df['pub_rec_bankruptcies'].fillna(0, inplace=True)
clean_df.isna().sum()
clean_df['collections_12_mths_ex_med'].value_counts()
clean_df['chargeoff_within_12_mths'].value_counts()
clean_df['tax_liens'].value_counts()
clean_df.drop(columns=['collections_12_mths_ex_med','chargeoff_within_12_mths','tax_liens'], axis=1, inplace=True)
clean_df.isna().sum()
clean_df['last_pymnt_d'].value_counts()
clean_df.dropna(subset=['last_pymnt_d','last_credit_pull_d'], axis=0, inplace=True)
clean_df.shape
clean_df.isna().sum()
clean_df['title'].value_counts()
clean_df['title'].fillna('Other', inplace=True)
clean_df.isna().sum()
clean_df['delinq_2yrs'].isna().sum()
clean_df['delinq_2yrs'].value_counts()
clean_df['delinq_2yrs'].fillna(0, inplace=True)
clean_df['earliest_cr_line']
clean_df.drop(columns=['earliest_cr_line'], axis=1, inplace=True)
clean_df['revol_util'].value_counts()
clean_df['revol_util'].fillna('0%', inplace=True)
clean_df.isna().sum()
clean_df.dropna(axis=0, inplace=True)
clean_df.reset_index(drop=True, inplace=True)
clean_df.head()
# ## Data Cleaning
new_clean_df = clean_df.copy()
new_clean_df.shape
new_clean_df.info()
# ### member_id
new_clean_df['member_id'] = new_clean_df['member_id'].astype('int')
new_clean_df.head()
# ### term
new_clean_df['term'].value_counts()
new_clean_df['term'].replace({' 36 months':36, ' 60 months':60}, inplace=True)
new_clean_df['term_months'] = new_clean_df['term'].copy()
new_clean_df.drop(columns=['term'], axis=1, inplace=True)
new_clean_df['term_months'].dtype
new_clean_df.head()
# ### int_rate
import re
def nums(x):
num = re.compile(r'[\d.]+').findall(x)
return num[0]
new_clean_df['int_rate_%'] = new_clean_df['int_rate'].apply(nums)
new_clean_df['int_rate_%'] = pd.to_numeric(new_clean_df['int_rate_%'])
new_clean_df['int_rate_%'].dtype
new_clean_df.drop(columns=['int_rate'], axis=1, inplace=True)
new_clean_df.head()
# ### emp_length
new_clean_df['emp_length'].value_counts()
new_clean_df['emp_length_year'] = new_clean_df['emp_length'].replace({'< 1 year':'0'})
new_clean_df['emp_length_year'] = new_clean_df['emp_length_year'].apply(nums)
new_clean_df['emp_length_year'] = new_clean_df['emp_length_year'].astype('int64')
# ### revol_util
new_clean_df['revol_util']
new_clean_df['revol_util_%'] = new_clean_df['revol_util'].apply(nums)
new_clean_df['revol_util_%'] = new_clean_df['revol_util_%'].astype('float64')
new_clean_df.drop(columns=['emp_length','revol_util'], axis=1, inplace=True)
new_clean_df.head()
# ### total_pymnt
new_clean_df['total_pymnt']
new_clean_df['total_pymnt'] = new_clean_df['total_pymnt'].apply(lambda x:np.round(x, decimals=2))
new_clean_df['total_pymnt']
new_clean_df.head()
new_clean_df.info()
new_clean_df.shape
cols = new_clean_df.columns.to_list()
new_clean_df[cols[:26]].describe()
new_clean_df[cols[26:]].describe()
# ### Saving Data
new_clean_df.to_csv('dataset/new_lending_club_loans.csv', index=None)
| data-cleansing-lending_club_loans.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# In this exercise, you will use your new knowledge to propose a solution to a real-world scenario. To succeed, you will need to import data into Python, answer questions using the data, and generate **histograms** and **density plots** to understand patterns in the data.
#
# ## Scenario
#
# You'll work with a real-world dataset containing information collected from microscopic images of breast cancer tumors, similar to the image below.
#
# 
#
# Each tumor has been labeled as either [**benign**](https://en.wikipedia.org/wiki/Benign_tumor) (_noncancerous_) or **malignant** (_cancerous_).
#
# To learn more about how this kind of data is used to create intelligent algorithms to classify tumors in medical settings, **watch the short video [at this link](https://www.youtube.com/watch?v=9Mz84cwVmS0)**!
#
#
# ## Setup
#
# Run the next cell to import and configure the Python libraries that you need to complete the exercise.
import pandas as pd
pd.plotting.register_matplotlib_converters()
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
print("Setup Complete")
# The questions below will give you feedback on your work. Run the following cell to set up our feedback system.
# Set up code checking
import os
if not os.path.exists("../input/cancer_b.csv"):
os.symlink("../input/data-for-datavis/cancer_b.csv", "../input/cancer_b.csv")
os.symlink("../input/data-for-datavis/cancer_m.csv", "../input/cancer_m.csv")
from learntools.core import binder
binder.bind(globals())
from learntools.data_viz_to_coder.ex5 import *
print("Setup Complete")
# ## Step 1: Load the data
#
# In this step, you will load two data files.
# - Load the data file corresponding to **benign** tumors into a DataFrame called `cancer_b_data`. The corresponding filepath is `cancer_b_filepath`. Use the `"Id"` column to label the rows.
# - Load the data file corresponding to **malignant** tumors into a DataFrame called `cancer_m_data`. The corresponding filepath is `cancer_m_filepath`. Use the `"Id"` column to label the rows.
# +
# Paths of the files to read
cancer_b_filepath = "../input/cancer_b.csv"
cancer_m_filepath = "../input/cancer_m.csv"
# Fill in the line below to read the (benign) file into a variable cancer_b_data
cancer_b_data = ____
# Fill in the line below to read the (malignant) file into a variable cancer_m_data
cancer_m_data = ____
# Run the line below with no changes to check that you've loaded the data correctly
step_1.check()
# -
# #%%RM_IF(PROD)%%
cancer_b_data = pd.read_csv(cancer_b_filepath, index_col="Id")
cancer_m_data = pd.read_csv(cancer_m_filepath, index_col="Id")
step_1.assert_check_passed()
# Lines below will give you a hint or solution code
#_COMMENT_IF(PROD)_
step_1.hint()
#_COMMENT_IF(PROD)_
step_1.solution()
# ## Step 2: Review the data
#
# Use a Python command to print the first 5 rows of the data for benign tumors.
# Print the first five rows of the (benign) data
____ # Your code here
# Use a Python command to print the first 5 rows of the data for malignant tumors.
# Print the first five rows of the (malignant) data
____ # Your code here
# In the datasets, each row corresponds to a different image. Each dataset has 31 different columns, corresponding to:
# - 1 column (`'Diagnosis'`) that classifies tumors as either benign (which appears in the dataset as **`B`**) or malignant (__`M`__), and
# - 30 columns containing different measurements collected from the images.
#
# Use the first 5 rows of the data (for benign and malignant tumors) to answer the questions below.
# +
# Fill in the line below: In the first five rows of the data for benign tumors, what is the
# largest value for 'Perimeter (mean)'?
max_perim = ____
# Fill in the line below: What is the value for 'Radius (mean)' for the tumor with Id 842517?
mean_radius = ____
# Check your answers
step_2.check()
# -
# #%%RM_IF(PROD)%%
max_perim = 87.46
mean_radius = 20.57
step_2.assert_check_passed()
# Lines below will give you a hint or solution code
#_COMMENT_IF(PROD)_
step_2.hint()
#_COMMENT_IF(PROD)_
step_2.solution()
# ## Step 3: Investigating differences
# #### Part A
#
# Use the code cell below to create two histograms that show the distribution in values for `'Area (mean)'` for both benign and malignant tumors. (_To permit easy comparison, create a single figure containing both histograms in the code cell below._)
# +
# Histograms for benign and maligant tumors
____ # Your code here (benign tumors)
____ # Your code here (malignant tumors)
# Check your answer
step_3.a.check()
# -
# #%%RM_IF(PROD)%%
sns.distplot(a=cancer_b_data['Area (mean)'], label="Benign", kde=False)
sns.distplot(a=cancer_m_data['Area (mean)'], label="Malignant", kde=False)
plt.legend()
step_3.a.assert_check_passed()
# Lines below will give you a hint or solution code
#_COMMENT_IF(PROD)_
step_3.a.hint()
#_COMMENT_IF(PROD)_
step_3.a.solution_plot()
# #### Part B
#
# A researcher approaches you for help with identifying how the `'Area (mean)'` column can be used to understand the difference between benign and malignant tumors. Based on the histograms above,
# - Do malignant tumors have higher or lower values for `'Area (mean)'` (relative to benign tumors), on average?
# - Which tumor type seems to have a larger range of potential values?
#_COMMENT_IF(PROD)_
step_3.b.hint()
#_COMMENT_IF(PROD)_
step_3.b.solution()
# ## Step 4: A very useful column
#
# #### Part A
#
# Use the code cell below to create two KDE plots that show the distribution in values for `'Radius (worst)'` for both benign and malignant tumors. (_To permit easy comparison, create a single figure containing both KDE plots in the code cell below._)
# +
# KDE plots for benign and malignant tumors
____ # Your code here (benign tumors)
____ # Your code here (malignant tumors)
# Check your answer
step_4.a.check()
# -
# #%%RM_IF(PROD)%%
sns.kdeplot(data=cancer_b_data['Radius (worst)'], shade=True, label="Benign")
sns.kdeplot(data=cancer_m_data['Radius (worst)'], shade=True, label="Malignant")
step_4.a.assert_check_passed()
# #%%RM_IF(PROD)%%
sns.kdeplot(data=cancer_b_data['Radius (worst)'], shade=True, label="Benign")
step_4.a.assert_check_failed()
# Lines below will give you a hint or solution code
#_COMMENT_IF(PROD)_
step_4.a.hint()
#_COMMENT_IF(PROD)_
step_4.a.solution_plot()
# #### Part B
#
# A hospital has recently started using an algorithm that can diagnose tumors with high accuracy. Given a tumor with a value for `'Radius (worst)'` of 25, do you think the algorithm is more likely to classify the tumor as benign or malignant?
#_COMMENT_IF(PROD)_
step_4.b.hint()
#_COMMENT_IF(PROD)_
step_4.b.solution()
# ## Keep going
#
# Review all that you've learned and explore how to further customize your plots in the **[next tutorial](#$NEXT_NOTEBOOK_URL$)**!
| notebooks/data_viz_to_coder/raw/ex5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# The activations live in the neural network (nn) library) in TensorFlow. Besides using built-in activation functions, we can also design our own using tensorFlow operations. We can import the predefined activation functions (from tensorflow import nn) or be explicit and write nn in our function calls. Here, we'll choose to be explicit with each function call.
# ## 1. ReLU
import tensorflow as tf
print(tf.nn.relu([-3., 3., 10.]))
# ## 2. ReLU 6
print(tf.nn.relu6([-3., 3., 10.]))
# ## 3. Sigmoid
print(tf.nn.sigmoid([-1., 0., 1.]))
# ## 4. Hyper tangent
print(tf.nn.tanh([-1., 0., 1.]))
# ## 5. Softsign function
print(tf.nn.softsign([-1., 0., 1.]))
# ## 6. Softplus function
print(tf.nn.softplus([-1., 0., 1.]))
# ## 7. ELU
print(tf.nn.elu([-1., 0., 1]))
# ## 8. Further resources...
# More activations can be found on the Keras activation page: https://www.tensorflow.org/api_docs/python/tf/keras/activations
# ## There's more...
# We can easily create custom activations such as Swish, which is x\*sigmoid(x), which can be used as a more performing replacement for ReLU activations in image and tabular data problems:
# +
def swish(x):
return x * tf.nn.sigmoid(x)
print(swish([-1., 0., 1.]))
| Chapter 1 - Getting Started with TensorFlow 2.x/Implementing activation functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <center><h2>Prediction of Online News Popularity</h2></center>
#
# <center><h3>Midterm Report for DATA1030 Fall 2021 at Brown University</h3></center>
#
# <center><h3>Supervised by Dr. <NAME></h3></center>
#
# <center><a href="https://github.com/spacegoat1/news_article_popularity">Project on Github</a></center>
# ## 1. Introduction
#
# Over the past couple of decades, with online news platforms increasingly winning out over physical newspapers, media organizations are relying more and more on analytics and machine learning to understand their reader base, moderate interactions and analyze which content is more likely to generate traffic. The pandemic has only accelerated this effort, with advertising revenues falling and platforms having the rely on subscriptions to generate revenues.
#
# In this context, being able to predict what content is likely to resonate with readers is very important not only for media organizations, but also for activists and political organizations, and as highlighted in recent news, tech companies as well. Such analyses can use information which is available after the fact, which can make the prediction task easier, or only use features accessible before publication. This approach yields lower predition scores, but can be significantly more useful as a signal to a content provider if it can indicate where to focus efforts in order to optimize content, or to a platform that publishes such content so that it can focus its moderation efforts on articles which are likely to go viral.
#
# This projects attempts to explore the performance of various machine learning models in predicting the likelihood of a news article becoming popular, as measured by how often it is shared on social media. The dataset used comes from the UCI Machine Learning Repository, covering a set of articles published on the online platform Mashable over a period of 2 years, from January 7 2013 to January 7 2015. The data contain 39,644 records, each corresponding to an article published on Mashable. Each records has 61 attributes - 2 of which are non-predictive, and 1 of which is the target variable (no. of shares) and the remaining 58 are predictive features. While the number of shares is a continuous variable, in this project a threshold is set, above which an article is labelled 'popular', thus converting the task into a classification problem.
#
# This project uses the paper published by <NAME>, <NAME> and <NAME> in **TBD** as its primary reference, which introduced and published the associated dataset. The authors of the article focus on predicting classifying article popularity before publication, and then make build a system to make recommendations to optimize the likelihood of popularity. They used 47 features and applied various models, eventually settling on a Random Forest classifier with an accuracy of 67% as the best model. Further work by **Zhang et. al.** expands upon this by using PCA for feature selection, but their difference in thresholds and classification (they use low/medium/high popularity tiers) is not comparable to the original paper. **Ren et. al.** use Mutual Information and Fisher Criterion for feature selection, and improve the accuracy of the random forest classifier to 69%. This project attempts to replicate their findings.
# ## 2. Exploratory Data Analysis
#
# The dataset includes features of various types, which can be categorized as below:
# **FIG_0 GOES HERE - Tabular info **
#
# Figure 2 shows a timeline of how popular articles of each type of topic were in aggregate over each month in the 2 year timeline of the dataset.
#
# A major consideration is to convert the target variable from a continuous variable into a target variable by setting a threshold value, above which the article is considered to be popular.
# **FIG_2 GOES HERE**
# The histogram shows that the number of shares has a very long tailed distribution with a small proportion of articles getting a very large number of shares. For the purposes of the project, we set a threshold value that roughly splits the dataset into 2 balanced classes - a class of unpopular articles and a class of popular articles. Consistent with the literature, we find that a threshold value of 1,400 does a good job.
#
#
# The features also tend to exhibit long tailed distributions. While none of them individually stand out as predictive of popularity, a scatter matrix of different types of features shows some promise for combinations of features of different types in explaining popularity. We have
# ## 3. Methods
#
# ### 3.1 Data Splitting
# The dataset consists of records of online articles published by Mashable. Each row corresponds to a single article. There are a total of 39,644 articles.
#
# The target variable is 'popular' - this is a binary variable where 0 corresponds to not-popular and 1 corresponds to popular. This variable is derived from the 'shares' variable present in the original dataset. Shares is the number of shares for a given article, and we set a threshold = 1400, above which an article is considered popular.
#
# The features list various characteristics about the articles, including some NLP based metrics, the number of tokens in the title, in the content, the number of images and videos, the topic and the day of the week on which the article was published. There are 58 features.
#
# Since there is no group structure or time-series structure to the data, the data can be considered IID.
#
# The point of this project was to be able to predict how popular an article would be *before publication*, and make recommendations to increase popularity.
#
# Each of the feature variables is available to us prior to publication in a production environment, so we can safely split the dataset into the standard train:validation:test split. Here, a split of 70:20:10, which gives us 27751, 7929, 3964 articles for the train, validation and test sets respectively.
# ### 3.2 Data Preprocessing
#
# On the continuous features, the MinMaxEncoder for features relating to polarity, Latent Dirichlet Allocation (LDA) features, polarity related features and no. of tokens in title, average token length and number of keywords. None of the other continuous features are well bounded and exhibit fat tailed distributions. Hence, they are encoded with a StandardScaler.
#
# From the categorical features, I use OneHotEncoder on data_channel, since that tells us what the topic of the article is, and it is not possibly to order topics. Similarly, I use OneHot encoding on the is_weekend feature.
#
# The last remaining categorical feature is the day_of_week feature, for which I use the OrdinalEncoder since the days of the week occur in a sequence.
# ## 4. References
# [1_INSERT LINK HERE] <NAME>, <NAME>, and <NAME>, *“A Proactive Intelligent Decision Support System for Predicting the Popularity of Online News”* EPIA 2015, pp. 535–546, 2015.
#
# [2_INSERT LINK HERE] <NAME>, <NAME>, Stanford University, *“Predicting and Evaluating the Popularity of Online News”*, Machine Learning Project Work Report, 2015, pp. 1-5.
#
# **Github repository**: https://github.com/spacegoat1/news_article_popularity
| src/0_midterm_report.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %run preset.ipynb
def display_image(organism_image):
clear_output(wait=True)
display(organism_image)
epochs_number = 3000
population = get_initial_population(POPULATION)
population.sort(key=lambda x: x.fitness())
epoch = 0
best = population[0]
add_legend(best, epoch)
epochs = []
fitnesses = []
for i in range(epochs_number):
epoch += 1
population = selection(population, selection_range())
if epoch % 20 == 0:
current_best = population[0]
image_with_text = add_legend(current_best, epoch)
im = concatenate_images(image_with_text, reference_image)
save_image(im, epoch)
print("number of genes:", len(current_best.genes))
display_image(im)
if epoch % 50 == 0:
epochs.append(epoch)
fitnesses.append(current_best.fitness())
change_mutation_probabilities(best, current_best)
best = population[0]
epochs = np.array(epochs)
fitnesses = np.array(fitnesses)
import matplotlib.pyplot as plt
fig=plt.figure(figsize=(14, 12), dpi= 80, facecolor='w', edgecolor='k')
plt.plot(epochs, fitnesses)
plt.show()
| genetic__v1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
gpus = tf.config.experimental.list_physical_devices('GPU')
if gpus:
try:
# Currently, memory growth needs to be the same across GPUs
for gpu in gpus:
tf.config.experimental.set_memory_growth(gpu, True)
logical_gpus = tf.config.experimental.list_logical_devices('GPU')
print(len(gpus), "Physical GPUs,", len(logical_gpus), "Logical GPUs")
except RuntimeError as e:
# Memory growth must be set before GPUs have been initialized
print(e)
import matplotlib.pyplot as plt
# %matplotlib inline
from tqdm.notebook import tqdm
import numpy as np
import time
import glob
from pyquaternion import Quaternion
from scipy.spatial.transform import Rotation as Rot
import seaborn as sns
sns.set_theme()
tf.config.list_physical_devices('GPU')
# +
from collections import defaultdict
def get_R(angles):
''' Get rotation matrix from three rotation angles (radians). right-handed.
Args:
angles: [3,]. x, y, z angles
Returns:
R: [3, 3]. rotation matrix.
'''
x, y, z = angles.astype(np.float32)
# x
Rx = np.array([[1, 0, 0],
[0, np.cos(x), -np.sin(x)],
[0, np.sin(x), np.cos(x)]])
# y
Ry = np.array([[np.cos(y), 0, np.sin(y)],
[0, 1, 0],
[-np.sin(y), 0, np.cos(y)]])
# z
Rz = np.array([[np.cos(z), -np.sin(z), 0],
[np.sin(z), np.cos(z), 0],
[0, 0, 1]])
R = Rz.dot(Ry.dot(Rx))
return R
INIT_AXES = np.array([[1,0,0], [0,1,0], [0,0,1]]).astype(np.float32)
def get_data(min_angle_rad=-np.pi, max_angle_rad=np.pi, nb_examples=30000, split=0.1):
data = defaultdict(lambda: [])
for i in tqdm(range(nb_examples)):
angles = np.random.uniform(min_angle_rad, max_angle_rad, size=3)
R = get_R(angles)
q = Quaternion(matrix=R, rtol=1e-05, atol=1e-05).elements.astype(np.float32)
data['R'].append(R)
data['angles'].append(angles)
# full quaternion
data['q'].append(q)
# quaternion constraint to one hemisphere
data['qh'].append(-q if q[0] < 0 else q)
data['rotated_axes'].append(R.dot(INIT_AXES.T).T)
for key in data.keys():
data[key] = np.array(data[key])
return data
# +
class NP_Q:
"""
Set of Numpy based functions to work with quaternions
"""
@classmethod
def norm(cls, q):
return np.sqrt(np.dot(q,q))
@classmethod
def mult(cls, p,q):
s = p[0]*q[0] - np.dot(p[1:], q[1:])
v = p[0]*q[1:] + q[0]*p[1:] + np.cross(p[1:], q[1:])
return np.append([s], v)
@classmethod
def conjugate(cls, q):
return np.array([q[0], -q[1], -q[2], -q[3]])
@classmethod
def inverse(cls, q):
return cls.conjugate(q) / np.dot(q,q)
@classmethod
def log(cls, q):
v = q[1:]
a = q[0]
x = a/cls.norm(q)
real_part = np.log(cls.norm(q))
vec_part = v/np.linalg.norm(v, axis=-1) * np.arccos(x)
return np.append([real_part], vec_part)
@classmethod
def geodesic_dist(cls, q1, q2):
x = cls.mult(cls.inverse(q1), q2)
return cls.norm(cls.log(x))
@classmethod
def angle_dist(cls, q1,q2):
x = 2*(np.dot(q1,q2))**2 - 1
return np.arccos(x) / np.pi * 180
class TF_Q:
"""
Set of Tensorflow based functions to work with quaternions
"""
@classmethod
def conjugate(cls, q):
mult = tf.constant(np.array([1,-1,-1,-1])[np.newaxis], dtype=np.float32)
return q*mult
@classmethod
def inverse(cls, q):
return cls.conjugate(q) / tf.reduce_sum(q*q, axis=-1, keepdims=True)
@classmethod
def log(cls, q):
v = q[:, 1:]
a = q[:, :1]
q_norm = tf.norm(q, axis=-1, keepdims=True)
x = a / q_norm
eps = np.finfo(np.float32).eps * 8.0
x *= (1.0 - eps)
vec_part = tf.nn.l2_normalize(v, axis=-1) * tf.math.acos(x)
real_part = tf.math.log(q_norm)
return tf.concat([real_part, vec_part], axis=-1)
@classmethod
def mult(cls, quaternion1, quaternion2):
w1, x1, y1, z1 = tf.unstack(quaternion1, axis=-1)
w2, x2, y2, z2 = tf.unstack(quaternion2, axis=-1)
x = x1 * w2 + y1 * z2 - z1 * y2 + w1 * x2
y = -x1 * z2 + y1 * w2 + z1 * x2 + w1 * y2
z = x1 * y2 - y1 * x2 + z1 * w2 + w1 * z2
w = -x1 * x2 - y1 * y2 - z1 * z2 + w1 * w2
return tf.stack((w, x, y, z), axis=-1)
@classmethod
def geodesic_dist(cls, q1, q2):
x = cls.mult(cls.inverse(q1), q2)
x = tf.norm(cls.log(x), axis=-1)
return x
@classmethod
def angle_dist(cls, q1, q2):
x = tf.reduce_sum(q1*q2, axis=-1)
eps = np.finfo(np.float32).eps * 8.0
x *= (1.0 - eps)
x = 2*tf.math.acos(x)
return tf.reduce_mean(x)
@classmethod
def rotate_by_q(cls, point, quaternion):
point = tf.pad(tensor=point, paddings=[[0,0], [1,0]], mode="CONSTANT")
point = cls.mult(quaternion, point)
point = cls.mult(point, cls.conjugate(quaternion))
_, xyz = tf.split(point, (1, 3), axis=-1)
return xyz
def quat_antipodal_loss(y_true, y_pred):
dist1 = tf.reduce_mean(tf.abs(y_true-y_pred), axis=-1)
dist2 = tf.reduce_mean(tf.abs(y_true+y_pred), axis=-1)
loss = tf.where(dist1<dist2, dist1, dist2)
return tf.reduce_mean(loss)
def euler_angles_loss(y_true, y_pred):
dist1 = tf.abs(y_true - y_pred)
dist2 = tf.abs(2*np.pi + y_true - y_pred)
dist3 = tf.abs(-2*np.pi + y_true - y_pred)
loss = tf.where(dist1<dist2, dist1, dist2)
loss = tf.where(loss<dist3, loss, dist3)
return tf.reduce_mean(loss)
def mean_angle_btw_vectors(v1, v2):
dot_product = tf.reduce_sum(v1*v2, axis=-1)
cos_a = dot_product / (tf.norm(v1, axis=-1) * tf.norm(v2, axis=-1))
eps = 1e-8
cos_a = tf.clip_by_value(cos_a, -1 + eps, 1 - eps)
angle_dist = tf.math.acos(cos_a) / np.pi * 180.0
return tf.reduce_mean(angle_dist)
class QuatNet(tf.keras.Model):
def __init__(self):
super(QuatNet, self).__init__()
self.block = tf.keras.Sequential([
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.Dense(4, activation=None)
])
def get_rotated(self, q_pred):
q_pred = tf.nn.l2_normalize(q_pred, axis=-1)
init_v = tf.constant(INIT_AXES[np.newaxis], dtype=tf.float32)
init_v = tf.tile(init_v, multiples=[tf.shape(q_pred)[0], 1, 1])
x_axis = TF_Q.rotate_by_q(init_v[:,0], q_pred)
y_axis = TF_Q.rotate_by_q(init_v[:,1], q_pred)
z_axis = TF_Q.rotate_by_q(init_v[:,2], q_pred)
y_pred = tf.stack([x_axis, y_axis, z_axis], axis=1)
return y_pred
def call(self, inputs, training=False):
x = tf.reshape(inputs, (-1, 9))
x = self.block(x)
x = tf.nn.l2_normalize(x, axis=-1)
self.add_metric(mean_angle_btw_vectors(inputs, self.get_rotated(x)),
name='mean_angular_distance', aggregation='mean')
return x
class EulerNet(tf.keras.Model):
def __init__(self):
super(EulerNet, self).__init__()
self.block = tf.keras.Sequential([
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.Dense(3, activation=None)
])
def angle2matrix(self, angles):
''' get rotation matrix from three rotation angles (radians). right-handed.
Args:
angles: [3,]. x, y, z angles
x: pitch. positive for looking down.
y: yaw. positive for looking left.
z: roll. positive for tilting head right.
Returns:
R: [3, 3]. rotation matrix.
'''
x, y, z = angles[0], angles[1], angles[2]
# x
Rx = tf.stack([1.0, 0.0, 0.0,
0.0, tf.math.cos(x), -tf.math.sin(x),
0.0, tf.math.sin(x), tf.math.cos(x)])
Rx = tf.reshape(Rx, (3,3))
# y
Ry = tf.stack([tf.math.cos(y), 0, tf.math.sin(y),
0, 1, 0,
-tf.math.sin(y), 0, tf.math.cos(y)])
Ry = tf.reshape(Ry, (3,3))
# z
Rz = tf.stack([tf.math.cos(z), -tf.math.sin(z), 0,
tf.math.sin(z), tf.math.cos(z), 0,
0, 0, 1])
Rz = tf.reshape(Rz, (3,3))
R = tf.matmul(Rz,tf.matmul(Ry, Rx))
return R
def get_rotated(self, pred_angles):
init_v = tf.constant(INIT_AXES, dtype=tf.float32)
Rs = tf.map_fn(self.angle2matrix, pred_angles)
y_pred = tf.transpose(tf.matmul(Rs, tf.transpose(init_v)), [0, 2, 1])
return y_pred
def call(self, inputs, training=False):
x = tf.reshape(inputs, (-1, 9))
x = self.block(x)
self.add_metric(mean_angle_btw_vectors(inputs, self.get_rotated(x)),
name='mean_angular_distance', aggregation='mean')
return x
class Net6D(tf.keras.Model):
def __init__(self, ):
super(Net6D, self).__init__()
self.block = tf.keras.Sequential([
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.Dense(6, activation=None)
])
def dot(self, a, b):
return tf.reduce_sum(a*b, axis=-1, keepdims=True)
def get_rotation_matrix(self, x):
c1 = x[:,:3]
c2 = x[:,3:]
c3 = tf.linalg.cross(c1, c2)
return tf.stack([c1,c2,c3], axis=2)
def get_rotated(self, repr_6d):
init_v = tf.constant(INIT_AXES, dtype=tf.float32)
Rs = self.get_rotation_matrix(repr_6d)
y_pred = tf.transpose(tf.matmul(Rs, tf.transpose(init_v)), [0, 2, 1])
return y_pred
def call(self, inputs, training=False):
x = tf.reshape(inputs, (-1, 9))
x = self.block(x)
c1 = tf.nn.l2_normalize(x[:, :3], axis=-1)
c2 = tf.nn.l2_normalize(x[:, 3:] - self.dot(c1,x[:, 3:])*c1, axis=-1)
x = tf.concat([c1,c2], axis=-1)
self.add_metric(mean_angle_btw_vectors(inputs, self.get_rotated(x)),
name='mean_angular_distance', aggregation='mean')
return x
# +
import matplotlib.ticker as mticker
from scipy.ndimage import gaussian_filter1d
def plot_histories(histories, title='Validation performance',
legends = ['Euler angles', 'Quaternions', '6D representation'],
colors = ['red', 'green', 'blue'], ax=None):
if ax is None:
fig, ax = plt.subplots(figsize=(8,5))
else:
fig=None
for idx, hist in enumerate(histories):
metrics = hist.history['val_mean_angular_distance']
metrics = gaussian_filter1d(metrics, 2)
ax.plot(range(1, len(hist.epoch)+1), metrics, color=colors[idx])
ax.set(xlabel='Epoch', ylabel='Mean angle error (deg)',
title=title)
yticks = ax.get_yticks().tolist()
ax.yaxis.set_major_locator(mticker.FixedLocator(yticks))
ylabels = ['%.1f°' % y for y in yticks]
_ = ax.set_yticklabels(ylabels)
_ = ax.set_xticks(range(1, len(hist.epoch)+1, 2))
ax.legend(legends)
return fig
def train_models(data, nb_epochs, verbose=0, batch_size=64):
euler_net = EulerNet()
euler_net.compile(optimizer='adam', loss = euler_angles_loss)
print('Training EulerNet')
euler_hist = euler_net.fit(x=data['rotated_axes'], y=data['angles'], batch_size=batch_size,
validation_split=0.2, epochs=nb_epochs, verbose=verbose)
quat_net = QuatNet()
quat_net.compile(optimizer='adam', loss = quat_antipodal_loss)
print('Training QuatNet')
quat_hist = quat_net.fit(x=data['rotated_axes'], y=data['q'], batch_size=batch_size,
validation_split=0.2, epochs=nb_epochs, verbose=verbose)
repr6D = data['R'][:, :, :2].transpose(0,2,1).reshape(-1, 6)
net6D = Net6D()
net6D.compile(optimizer='adam', loss = 'mae')
print('Training Repr6DNet')
history_6d = net6D.fit(x=data['rotated_axes'], y=repr6D, batch_size=batch_size,
validation_split=0.2, epochs=nb_epochs, verbose=verbose)
return [euler_hist, quat_hist, history_6d]
# -
data = get_data(-np.pi/4, np.pi/4, 30000)
histories = train_models(data, nb_epochs=20, verbose=0, batch_size=64)
fig1 = plot_histories(histories, 'Models performances with rotations up to ± 45°')
fig1.savefig('assets/plot45.png')
data = get_data(-np.pi/2, np.pi/2, 30000)
histories2 = train_models(data, nb_epochs=20, verbose=0, batch_size=64)
fig2 = plot_histories(histories2, 'Models performances with rotations up to ± 90°')
fig2.savefig('assets/plot90.png')
data = get_data(-np.pi, np.pi, 30000)
histories3 = train_models(data, nb_epochs=20, verbose=0, batch_size=64)
fig3 = plot_histories(histories3[1:], 'Models performances with full-range rotations up to ± 180°',
legends = ['Quaternions', '6D representation'],
colors= ['green', 'blue'])
fig3.savefig('assets/plot180_1.png')
fig4 = plot_histories(histories3[:1], 'Models performances with full-range rotations up to ± 180°',
legends = ['Euler angles'],
colors= ['red'])
fig4.savefig('assets/plot180_2.png')
# +
import matplotlib.gridspec as gridspec
gs = gridspec.GridSpec(2, 4)
gs.update(wspace=1.0)
plt.figure(figsize=(13,13))
ax = plt.subplot(gs[0, :])
fig3 = plot_histories(histories3[1:], '',
legends = ['Quaternions', '6D representation'],
colors= ['green', 'blue'], ax=ax)
ax = plt.subplot(gs[1, :2])
plot_histories(histories3,
'',
ax = ax)
ax = plt.subplot(gs[1, 2:])
plot_histories(histories3[:1], '',
legends = ['Euler angles'],
colors= ['red'],
ax = ax)
_= plt.suptitle('Models performances with full-range rotations up to ± 180°',
fontsize=16, y=0.9)
plt.savefig('assets/plot180.png')
| rotations_performance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Overfitting vs. Underfitting
#
# Exploring a fundamental problem in modeling. This notebook will look at a simple example showing the problem of overfitting and underfitting as well as how to address it via cross-validation. This example is based on the [scikit-learn exercise](http://scikit-learn.org/stable/auto_examples/model_selection/plot_underfitting_overfitting.html).
# ## Imports
#
# We will use numpy and pandas, two of the most common libraries for data manipulation. We use scikit-learn, a popular machine learning library, for creating and evaluating the models. Matplotlib is used for model visualization.
# +
# Numpy and pandas as usual
import numpy as np
import pandas as pd
# Scikit-Learn for fitting models
from sklearn.preprocessing import PolynomialFeatures
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import cross_val_score
from sklearn.metrics import mean_squared_error
# For plotting in the notebook
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
# Default parameters for plots
matplotlib.rcParams['font.size'] = 12
matplotlib.rcParams['figure.titlesize'] = 16
matplotlib.rcParams['figure.figsize'] = [9, 7]
# -
# ## Generate a relationship
#
# First, we need a "true" relationship. We define a curve, in this case a sine curve to serve as our process that generates the data. As the real-world is never perfectly celan however, we also need to add some noise into the observations. This is done by adding a small random number to each value.
# +
# Set the random seed for reproducible results
np.random.seed(42)
# "True" generating function representing a process in real life
def true_gen(x):
y = np.sin(1.2 * x * np.pi)
return(y)
# x values and y value with a small amount of random noise
x = np.sort(np.random.rand(120))
y = true_gen(x) + 0.1 * np.random.randn(len(x))
# -
# ## Training and Testing Sets
# +
# np.random.choice(list(range(120)), size = 120, replace=False)
# +
# np.random.choice?
# +
# Random indices for creating training and testing sets
random_ind = np.random.choice(list(range(120)), size = 120, replace=False)
xt = x[random_ind]
yt = y[random_ind]
# Training and testing observations
train = xt[:int(0.7 * len(x))]
test = xt[int(0.7 * len(x)):]
y_train = yt[:int(0.7 * len(y))]
y_test = yt[int(0.7 * len(y)):]
# Model the true curve
x_linspace = np.linspace(0, 1, 1000)
y_true = true_gen(x_linspace)
# -
# Visualize observations and true curve
plt.plot(train, y_train, 'ko', label = 'Train');
plt.plot(test, y_test, 'ro', label = 'Test')
plt.plot(x_linspace, y_true, 'b-', linewidth = 2, label = 'True Function')
plt.legend()
plt.xlabel('x'); plt.ylabel('y'); plt.title('Data');
# # Polynomial Model
#
# We want to try and capture the data using a polynomial function. A polynomial is defined by the degree, or the highest power to for the x-values. A line has a degree of 1 because it is of the form $y = b_1*x + b_0$ where $b_1$ is the slope and $b_0$ is the intercept. A third degree polynomial would have the form $y = b_3 * x^3 + b_2 * x^2 + b_1 * x + b_0$ and so on. The higher the degree of the polynomial, the more flexible the model. A more flexible model is prone to overfitting because it can can "bend" to follow the training data.
#
# The following function creates a polynomial with the specified number of degrees and plots the results. We can use these results to determine the optimal degrees to achieve the right balance between over and underfitting.
# +
def fit_poly(train, y_train, test, y_test, degrees, plot='train', return_scores=False):
# Create a polynomial transformation of features
features = PolynomialFeatures(degree=degrees, include_bias=False)
# Reshape training features for use in scikit-learn and transform features
train = train.reshape((-1, 1))
train_trans = features.fit_transform(train)
# Create the linear regression model and train
model = LinearRegression()
model.fit(train_trans, y_train)
# Calculate the cross validation score
cross_valid = cross_val_score(model, train_trans, y_train, scoring='neg_mean_squared_error', cv = 5)
# Training predictions and error
train_predictions = model.predict(train_trans)
training_error = mean_squared_error(y_train, train_predictions)
# Format test features
test = test.reshape((-1, 1))
test_trans = features.fit_transform(test)
# Test set predictions and error
test_predictions = model.predict(test_trans)
testing_error = mean_squared_error(y_test, test_predictions)
# Find the model curve and the true curve
x_curve = np.linspace(0, 1, 100)
x_curve = x_curve.reshape((-1, 1))
x_curve_trans = features.fit_transform(x_curve)
# Model curve
model_curve = model.predict(x_curve_trans)
# True curve
y_true_curve = true_gen(x_curve[:, 0])
# Plot observations, true function, and model predicted function
if plot == 'train':
plt.plot(train[:, 0], y_train, 'ko', label = 'Observations')
plt.plot(x_curve[:, 0], y_true_curve, linewidth = 4, label = 'True Function')
plt.plot(x_curve[:, 0], model_curve, linewidth = 4, label = 'Model Function')
plt.xlabel('x'); plt.ylabel('y')
plt.legend()
plt.ylim(-1, 1.5); plt.xlim(0, 1)
plt.title('{} Degree Model on Training Data'.format(degrees))
plt.show()
elif plot == 'test':
# Plot the test observations and test predictions
plt.plot(test, y_test, 'o', label = 'Test Observations')
plt.plot(x_curve[:, 0], y_true_curve, 'b-', linewidth = 2, label = 'True Function')
plt.plot(test, test_predictions, 'ro', label = 'Test Predictions')
plt.ylim(-1, 1.5); plt.xlim(0, 1)
plt.legend(), plt.xlabel('x'), plt.ylabel('y'); plt.title('{} Degree Model on Testing Data'.format(degrees)), plt.show();
# Return the metrics
if return_scores:
return training_error, testing_error, -np.mean(cross_valid)
# -
# ## Try Model with Different Degrees
# ### Degrees = 1 -> Underfitting
#
# In this case, a linear model cannot accurate learn the relationship between x and y and will underfit the data.
fit_poly(train, y_train, test, y_test, degrees = 1, plot='train')
fit_poly(train, y_train, test, y_test, degrees = 1, plot='test')
# ## Degrees = 25 -> Overfitting
#
# We can go in the completely opposite direction and create a model that overfits the data. This model has too much flexibility and learns the training data too closely. As the training data has some amount of noise, it will end up capturing that noise and will be misled by that noise when it tries to make predictions on the test data.
fit_poly(train, y_train, test, y_test, plot='train', degrees = 25)
fit_poly(train, y_train, test, y_test, degrees=25, plot='test')
# ## Degrees = 5 -> Balanced Model
#
# Now that we have seen the two extremes, we can take a look at a model that does a good job of both accounting for the data while not following it too closely.
fit_poly(train, y_train, test, y_test, plot='train', degrees = 5)
fit_poly(train, y_train, test, y_test, degrees=5, plot='test')
# # Cross Validation
#
# To pick the optimal model, we need to use a validation set. Cross validation is even better than a single validation set because it uses numerous validation sets created from the training data. In this case, we are using 5 different validation sets. The model that performs best on the cross validation is usually the optimal model because it has shown that it can learn the relationships while not overfitting.
# +
# Range of model degrees to evaluate
degrees = [int(x) for x in np.linspace(1, 40, 40)]
# Results dataframe
results = pd.DataFrame(0, columns = ['train_error', 'test_error', 'cross_valid'], index = degrees)
# Try each value of degrees for the model and record results
for degree in degrees:
degree_results = fit_poly(train, y_train, test, y_test, degree, plot=False, return_scores=True)
results.ix[degree, 'train_error'] = degree_results[0]
results.ix[degree, 'test_error'] = degree_results[1]
results.ix[degree, 'cross_valid'] = degree_results[2]
# -
print('10 Lowest Cross Validation Errors\n')
train_eval = results.sort_values('cross_valid').reset_index(level=0).rename(columns={'index': 'degrees'})
train_eval.ix[:,['degrees', 'cross_valid']] .head(10)
plt.plot(results.index, results['cross_valid'], 'go-', ms=6)
plt.xlabel('Degrees'); plt.ylabel('Cross Validation Error'); plt.title('Cross Validation Results');
plt.ylim(0, 0.2);
print('Minimum Cross Validation Error occurs at {} degrees.\n'.format(int(np.argmin(results['cross_valid']))))
# # Final Model
#
# The model with the lowest cross validation error had four degrees. Therefore, we will use a 4th degree polynomial for the final model.
fit_poly(train, y_train, test, y_test, degrees=4, plot='train')
fit_poly(train, y_train, test, y_test, degrees=4, plot='test')
# ## Evaluate Models
#
# The next step is to examine the scores for the models. We will use a range of values to see how the performance on the training and testing set compares. A model with much lower errors on the training data than the testing data is overfit. A model with high error on the training data (which will lead to high testing error as well) is underfitting because it does not even learn the training data.
# #### Quantitative Comparison
print('10 Lowest Training Errors\n')
train_eval = results.sort_values('train_error').reset_index(level=0).rename(columns={'index': 'degrees'})
train_eval.ix[:,['degrees', 'train_error']] .head(10)
print('10 Lowest Testing Errors\n')
train_eval = results.sort_values('test_error').reset_index(level=0).rename(columns={'index': 'degrees'})
train_eval.ix[:,['degrees', 'test_error']] .head(10)
# #### Visual Comparison
# +
plt.plot(results.index, results['train_error'], 'b-o', ms=6, label = 'Training Error')
plt.plot(results.index, results['test_error'], 'r-*', ms=6, label = 'Testing Error')
plt.legend(loc=2); plt.xlabel('Degrees'); plt.ylabel('Mean Squared Error'); plt.title('Training and Testing Curves');
plt.ylim(0, 0.05); plt.show()
print('\nMinimum Training Error occurs at {} degrees.'.format(int(np.argmin(results['train_error']))))
print('Minimum Testing Error occurs at {} degrees.\n'.format(int(np.argmin(results['test_error']))))
# -
| over_vs_under/Over vs Under Fitting Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Compute GitHub Stats
# +
# NOTE: The RuntimeWarnings (if any) are harmless. See ContinuumIO/anaconda-issues#6678.
from pandas.io import gbq
import pandas as pd
import numpy as np
from importlib import reload
import itertools
# +
import getpass
import subprocess
# Configuration Variables. Modify as desired.
PROJECT = subprocess.check_output(["gcloud", "config", "get-value", "project"]).strip().decode()
# -
# %matplotlib
# ## Setup Authorization
#
# If you are using a service account run
# # %%bash
#
# # Activate Service Account provided by Kubeflow.
# gcloud auth activate-service-account --key-file=${GOOGLE_APPLICATION_CREDENTIALS}
#
# If you are running using user credentials
#
# gcloud auth application-default login
import datetime
datetime.datetime.now().month
# +
months = []
for year in ["2018"]:
for month in range(1, 13):
months.append("\"{0}{1:02}\"".format(year, month))
for year in ["2019"]:
for month in range(1, datetime.datetime.now().month + 1):
months.append("\"{0}{1:02}\"".format(year, month))
months
# -
# ## Unique PR Creators
# +
query = """
SELECT
DATE(created_at) AS pr_date,
actor.id,
actor.login,
JSON_EXTRACT(payload, '$.pull_request.user.id') as user_id,
JSON_EXTRACT(payload, '$.pull_request.id') as pr_id,
JSON_EXTRACT(payload, '$.pull_request.merged') as merged
FROM `githubarchive.month.*`
WHERE
_TABLE_SUFFIX IN ({0})
AND type = 'PullRequestEvent'
AND org.login = 'kubeflow'
AND JSON_EXTRACT(payload, '$.action') IN ('"closed"')
""".format(",".join(months))
all_prs=gbq.read_gbq(str(query), dialect='standard', project_id=PROJECT)
# -
# Filter PRs to merged PRs
v=all_prs["merged"].values == 'true'
merged_all_prs = all_prs.iloc[v]
p=pd.Series(data=merged_all_prs["user_id"].values,index=merged_all_prs["pr_date"])
p=p.sort_index()
# +
# Some solutions here: https://stackoverflow.com/questions/46470743/how-to-efficiently-compute-a-rolling-unique-count-in-a-pandas-time-series
# Need to figure out how to do a time based window
creators = p.rolling('28d').apply(lambda arr: pd.Series(arr).nunique())
# -
import matplotlib
from matplotlib import pylab
matplotlib.rcParams.update({'font.size': 22})
hf = pylab.figure()
hf.set_size_inches(18.5, 10.5)
pylab.plot(creators, linewidth=5)
ha = pylab.gca()
ha.set_title("Unique PR Authors (Last 28 Days)")
ha.set_xlabel("Date")
ha.set_ylabel("# Authors")
# ### Number Prs
# +
pr_impulse=pd.Series(data=merged_all_prs["pr_id"].values,index=merged_all_prs["pr_date"])
pr_impulse=pr_impulse.sort_index()
unique_prs = pr_impulse.rolling('28d').apply(lambda arr: pd.Series(arr).nunique())
# +
hf = pylab.figure()
hf.set_size_inches(18.5, 10.5)
pylab.plot(unique_prs, linewidth=5)
ha = pylab.gca()
ha.set_title("Unique PRs(Last 28 Days)")
ha.set_xlabel("Date")
ha.set_ylabel("# PRs")
# -
# ## Release stats per release (quarter)
#
# * Compute stats about a release
# * We do this based on time
# * You can see a sample of the payload at [https://api.github.com/repos/kubeflow/pipelines/pulls/1038](https://api.github.com/repos/kubeflow/pipelines/pulls/1038)
# +
release_months = []
year = 2019
for month in range(8, 11):
release_months.append("\"{0}{1:02}\"".format(year, month))
query = """
SELECT
DATE(created_at) AS pr_date,
actor.id,
actor.login,
JSON_EXTRACT(payload, '$.pull_request.merged') as merged,
JSON_EXTRACT(payload, '$.pull_request.id') as pr_id,
JSON_EXTRACT(payload, '$.pull_request.url') as pr_url,
JSON_EXTRACT(payload, '$.pull_request.user.id') as user_id
FROM `githubarchive.month.*`
WHERE
_TABLE_SUFFIX IN ({0})
AND type = 'PullRequestEvent'
AND org.login = 'kubeflow'
AND JSON_EXTRACT(payload, '$.action') IN ('"closed"')
""".format(",".join(release_months))
prs=gbq.read_gbq(str(query), dialect='standard', project_id=PROJECT)
# -
# Filter PRs to merged PRs
v=prs["merged"].values == 'true'
merged_prs = prs.iloc[v]
# +
unique_pr_logins = prs["user_id"].unique()
unique_prs = prs["pr_id"].unique()
merged_unique_logins = merged_prs["user_id"].unique()
merged_unique_prs = merged_prs["pr_id"].unique()
print("Number of unique pr authors (merged & unmerged) {0}".format(unique_pr_logins.shape))
print("Number of unique prs (merged & unmerged) {0}".format(unique_prs.shape))
print("Number of unique pr authors (merged) {0}".format(merged_unique_logins.shape))
print("Number of unique prs (merged) {0}".format(merged_unique_prs.shape))
# -
# ## Get a list of distinct actions
#
# * Here's a list of events in the [api](https://developer.github.com/v4/union/pullrequesttimelineitems/)
# * It looks like these are different from the ones in the github archive
# +
query = """
SELECT
distinct JSON_EXTRACT(payload, '$.action')
FROM `githubarchive.month.*`
WHERE
_TABLE_SUFFIX IN ({0})
""".format(",".join(months))
actions=gbq.read_gbq(str(query), dialect='standard', project_id=PROJECT)
# -
actions
# ## New Issues Last 28 Days
# +
query = """
SELECT
DATE(created_at) AS issue_date,
actor.id,
actor.login,
JSON_EXTRACT(payload, '$.pull_request.id') as issue_id,
JSON_EXTRACT(payload, '$.pull_request.url') as issue_url
FROM `githubarchive.month.*`
WHERE
_TABLE_SUFFIX IN ({0})
AND type = 'IssuesEvent'
AND org.login = 'kubeflow'
AND JSON_EXTRACT(payload, '$.action') IN ('"opened"')
""".format(",".join(months))
issues=gbq.read_gbq(str(query), dialect='standard', project_id=PROJECT)
# -
issue_counts=issues["issue_date"].value_counts()
issue_counts=issue_counts.sort_index()
rolling_issue_count = issue_counts.rolling('28d').sum()
# +
import matplotlib
from matplotlib import pylab
matplotlib.rcParams.update({'font.size': 22})
hf = pylab.figure()
hf.set_size_inches(18.5, 10.5)
pylab.plot(rolling_issue_count, linewidth=5)
ha = pylab.gca()
ha.set_title("New Kubeflow Issues (28 Days)")
ha.set_xlabel("Date")
ha.set_ylabel("# Of Issues")
# -
# ## GetSomeSampleIssue Events
# +
query = """
SELECT
*
FROM `githubarchive.month.*`
WHERE
_TABLE_SUFFIX IN ({0})
AND type = 'IssuesEvent'
AND org.login = 'kubeflow'
limit 20
""".format(",".join(months))
events=gbq.read_gbq(str(query), dialect='standard', project_id=PROJECT)
# -
events
# ## Get some sample pull request events
#
# * Want to inspect the data
#
# +
query = """
SELECT
*
FROM `githubarchive.month.*`
WHERE
_TABLE_SUFFIX IN ({0})
AND type = 'PullRequestEvent'
AND org.login = 'kubeflow'
limit 20
""".format(",".join(months))
events=gbq.read_gbq(str(query), dialect='standard', project_id=PROJECT)
# -
import pprint
import json
data = json.loads(events["payload"].values[3])
pprint.pprint(data)
data["pull_request"]["id"]
# ## Get Distinct Types
# +
query = """
SELECT
distinct type
FROM `githubarchive.month.*`
WHERE
_TABLE_SUFFIX IN ({0})
AND org.login = 'kubeflow'
limit 20
""".format(",".join(months))
events=gbq.read_gbq(str(query), dialect='standard', project_id=PROJECT)
# -
events
| scripts/github_stats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="rOHm30CGBYrF" colab_type="text"
# # Intro to Pandas
# 
#
# (logo by [<NAME>](https://github.com/pandas-dev/pandas/blob/master/web/pandas/static/img/pandas.svg))
#
# The Pandas Library is built on top of Numpy and is designed to make working with data fast and easy. Like Numpy, the library includes data structures and functions to manipulate that data.
#
# As we learned in the Numpy Notebook, we need to load in the library before we can use it.
# + id="4Shdx3b2BYrJ" colab_type="code" colab={}
from pandas import Series, DataFrame
import numpy as np
import pandas as pd
# + [markdown] id="ayjxD1jPBYrU" colab_type="text"
# Let's dissect the code above.
#
# The `Series` and `DataFrame` datatypes are commonly used so we import them directly with
#
# ```
# from pandas import Series, DataFrame
# ```
#
# For all other datatypes and functions in the library the `pd` prefix is commonly used so we import that with
#
# ```
# import pandas as pd
# ```
#
# ## Series
# A series is a 1d array-like object
#
# Let's consider the heights (in cm) of Japan's Women's Basketball Team at the 2016 Olympics.
#
# We can create a series in a number of ways.
#
# ##### directly from a Python list
# + id="buEkXx7zBYrX" colab_type="code" colab={}
japan = [173, 182, 185, 176, 183, 165, 191, 177, 165, 161, 175, 189]
athletesHeight = Series(japan)
# + [markdown] id="BBDKw9nwBYre" colab_type="text"
# We could also have done
# + id="nXGdbz4iBYrg" colab_type="code" colab={}
athletesHeight = Series([173, 182, 185, 176, 183, 165, 191, 177, 165, 161, 175, 189])
# + [markdown] id="pbEi2WLiBYrn" colab_type="text"
# In either case we can see the value of the Series athletesHeight
# + id="kSnYEZSrBYro" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 238} outputId="1df95f40-f9df-4f6a-80ae-feac4db495c9"
athletesHeight
# + [markdown] id="ZKcpGn_GBYrw" colab_type="text"
# As like arrays you are familiar with, the left number is the index and the right the value. And we can find the value at a particular index by the usual:
# + id="-IOxdmlrBYrw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="2b1a3fd7-7731-4a13-c5da-71ea7d80b83f"
athletesHeight[3]
# + [markdown] id="kpt6L0RcBYr3" colab_type="text"
# #### specifying indices
# Instead of the index 0, 1, 2, 3 ... you can specify your own index values. For example, we can label them 'Aiko'. 'Ayumi', "Fujiko" ... etc.
# + id="LQytLmIpBYr5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 238} outputId="bf14ef3d-9fd8-4e2e-f963-158bd19218c9"
athletes2 = Series(japan, index = ['Aiko', 'Ayumi', 'Fujiko', 'Hiroko', 'Itsumi', 'Junko', 'Kanae','Kotori', 'Mieko', 'Momoko', 'Nao', 'Rei'])
athletes2
# + [markdown] id="4uRQqLKoBYr-" colab_type="text"
# The names we see are not another column of the data. We can see the shape of athletes2 by:
# + id="dwCyF-Q4BYsA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9c5d27af-4493-4969-b180-b1edf129597e"
athletes2.shape
# + [markdown] id="TzALSffSBYsE" colab_type="text"
# This shows that athletes2 is a one dimensional matrix and that dimension has a length of 12. So the names we see are not values in a column but rather the indices.
#
#
# Let's use the index to get the Height of Junko:
# + id="PqkdaplvBYsG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="303da61e-a2a3-44f0-ad78-28c0560f70f3"
athletes2['Junko']
# + [markdown] id="wWTcb0mKBYsM" colab_type="text"
# How would you get the height of Nao?
#
# + id="t8HCm7CpBYsO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="754b55c3-8d27-452c-b988-5cbc211d1283"
athletes2['Nao']
# + [markdown] id="ej0jZxKABYsT" colab_type="text"
#
# ## DataFrame
# DataFrames are the most important data structure of Pandas and are simply
# a table or spreadsheet like structure. A DataFrame represents a table like:
#
# make | mpg | cylinders | HP | 0-60
# :---- | :---: | :---: | :---: | :---:
# Fiat | 38 | 4 | 157 | 6.9
# Ford F150 | 19 | 6 | 386 | 6.3
# Mazda 3 | 37 | 4 | 155 | 7.5
# Ford Escape | 27 | 4 | 245 | 7.1
# Kia Soul | 31 | 4 | 164 | 8.5
#
# A common way to create a DataFrame is to use a python dictionary as follows:
# + id="Sfch6JTxBYsT" colab_type="code" colab={}
cars = {'make': ['Fiat 500', 'Ford F-150', 'Mazda 3', 'Ford Escape', 'Kia Soul'],
'mpg': [38, 19, 37, 27, 31],
'cylinders': [4, 6, 4, 4, 4],
'HP': [157, 386, 155, 245, 164],
'0-60': [6.9, 6.3, 7.5, 7.1, 8.5]}
# + [markdown] id="Ybao1y-xBYsa" colab_type="text"
# Just to check that you are paying attention, what is the code to show the MPG of a Ford F-150? (so `cars['make'][3]` will print out something, what will print out the MPG of a Ford F-150?
# + id="m8PlHKsSBYsc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5e8af6c7-52b1-4bca-ca21-c6d1029fff01"
cars['mpg'][1]
# + [markdown] id="QDadkvC1BYsl" colab_type="text"
# and now we will create a DataFrame from the `cars` Python dictionary:
# + id="y1by6yLNBYsn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="7eeb7ddc-d588-4cae-b46f-504fda763f00"
df = DataFrame(cars)
df
# + [markdown] id="R1nYmp9JBYsr" colab_type="text"
# Prior to my life with Pandas, I would represent a table like the above one as:
# + id="LlmwxG5uBYss" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="bae051d4-f39e-4efe-dc77-0f1e54247eb4"
prePandas = [{'make': 'Fiat', 'mpg': 38, 'cylinders': 4, 'HP': 157, '0-60': 6.9},
{'make': 'Ford F-150', 'mpg': 19, 'cylinders': 6, 'HP': 386, '0-60': 6.3},
{'make': 'Mazda 3', 'mpg': 37, 'cylinders': 4, 'HP': 155, '0-60': 7.5},
{'make': 'Ford Escape', 'mpg': 27, 'cylinders': 4, 'HP': 245, '0-60': 7.1},
{'make': 'Kia Soul', 'mpg': 31, 'cylinders': 4, 'HP': 164, '0-60':8.5}]
prePandas[0]['make']
# + [markdown] id="BysFhLwZBYsy" colab_type="text"
# In the prePandas scheme the data is organized first by rows. That seemed logical to me since each row represents an object and is how we organize data in an SQL database. In the Pandas representation the data is organized by columns.
#
# Perhaps you noticed that the columns are displayed in alphabetically sorted order. If we want to specify a column order we can do so:
#
#
#
# + id="V2HrblcvBYsy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="ffe55429-e15a-446c-8804-ac25eb5d7206"
df2 = DataFrame(cars, columns=['make', 'mpg', 'cylinders', 'HP', '0-60' ])
df2
# + [markdown] id="Cg5IGn6dBYs2" colab_type="text"
# We can also create a DataFrame from a set of Series objects:
# + id="dTkSqtk-BYs3" colab_type="code" colab={}
cars2 = {'make': Series(['Fiat 500', 'Ford F-150', 'Mazda 3', 'Ford Escape', 'Kia Soul']),
'mpg': Series([38, 19, 37, 27, 31]),
'cylinders': Series([4, 6, 4, 4, 4]),
'HP': [157, 386, 155, 245, 164],
'0-60': Series([6.9, 6.3, 7.5, 7.1, 8.5])}
df3 = DataFrame(cars2)
# + [markdown] id="J5VHjZOLBYs7" colab_type="text"
# <h3 style="color:red">Q1. You Try</h3>
# <span style="color:red">Make a DataFrame representing the data in the following table:</span>
#
# Athlete | Sport | Height | Weight
# ----: | :---: | :---: | :---:
# <NAME> | Gymnastics | 62 | 115
# <NAME> | Basketball | 74 | 190
# <NAME> | Basketball | 72 | 163
# <NAME> | Track | 68 | 120
# <NAME> | Gymnastics | 57 | 104
# Madison Kocian | Gymnastics |62 | 101
# <NAME> |Track | 64 | 99
#
# + id="xLXFcB3EBYs8" colab_type="code" colab={}
# + colab_type="code" id="BNS8yap-n98l" colab={"base_uri": "https://localhost:8080/", "height": 153} outputId="d6ec26be-bf5e-4475-d569-17f64b97ede5"
wolympics = {'Athlete': ['<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>','Madison Kocian','<NAME>'],
'Sport': ['Gymnastics', 'Basketball', 'Basketball', 'Track', 'Gymnastics','Gymnastics','Track'],
'Height': [62, 74, 72, 68, 57,62,64],
'Weight': [115, 190, 163, 120, 104,101,99]}
dfwoplymics = DataFrame(wolympics)
print(dfwoplymics)
# + [markdown] id="_Hq-pLoJBYtB" colab_type="text"
# ## reading data from different sources.
# ### csv file:
# We can create a DataFrame from a CSV file (comma separated values), which is a common format for datasets.
#
#
# We use the `pd.read_csv` function to do so. `pd.read_csv` can read a csv file from either your local machine or the web. Let's start with the web.
#
# #### Reading a CSV file from the web.
# To read a file from the web, we simply provide a URL:
#
# + id="4T00ki7xBYtC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 917} outputId="4514c2a8-79e2-4bfb-890b-d09502164c49"
df4 = pd.read_csv('https://raw.githubusercontent.com/zacharski/ml-class/master/data/athletes.csv')
df4
# + [markdown] id="lbAXN6KLBYtF" colab_type="text"
# Sometimes the csv file has a header row as was the case in the example above. That file starts
#
# Name,Sport,Height,Weight
# <NAME>,Gymnastics,54,66
# Brittainey Raven,Basketball,72,162
# <NAME>,Basketball,78,204
# <NAME>,Gymnastics,49,90
#
#
#
# + id="N-YchHxaBYtF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 510} outputId="c7861db4-a7f7-40be-8a76-90e5402ade6c"
# !curl https://raw.githubusercontent.com/zacharski/ml-class/master/data/athletes.csv
# + [markdown] id="j89s6lQBBYtJ" colab_type="text"
# As you can see we can preface any Unix command with a bang (!) to have it execute in our Notebook. This is amazingly handy:
#
#
# + id="_HqSIgfsBYtK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1a3200f7-93c1-40e9-e07e-f56cca86876a"
# !ls
# + id="WehfdIjuBYtS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3e021610-7ec4-4bf6-b24a-150931525220"
# !pwd
# + [markdown] id="mVP4EZe3BYtg" colab_type="text"
# Back to Pandas.
#
# Sometimes the csv file does not have a header row. So for example, a csv file might have the contents:
#
# <NAME>,Gymnastics,54,66
# Brittainey Raven,Basketball,72,162
# <NAME>,Basketball,78,204
# <NAME>,Gymnastics,49,90
# In that case you specify the names of the columns using the names attribute:
# + id="T4NBPcb7BYti" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 917} outputId="48eacfd6-f684-4a05-87e1-8333656033bc"
athletes = pd.read_csv('https://raw.githubusercontent.com/zacharski/ml-class/master/data/athletesNoHeader.csv', names=['Name', 'Sport', 'Height', 'Weight'])
athletes
# + [markdown] id="eqztYqxbBYtn" colab_type="text"
# ### Reading a CSV file the local machine
# First, let's get that file onto our local machine:
# + id="52FhxXNTBYto" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="e6524ba8-b726-428c-f3b2-18840bcd4d81"
# !curl https://raw.githubusercontent.com/zacharski/ml-class/master/data/athletes.csv > localAthletes.csv
# + [markdown] id="NxjuNiI5BYtw" colab_type="text"
# Hmm. That didn't work. Can you fix the error and rerun that cell?
#
# ---
# Now we can specify the local file using `pd=read_csv`
# + id="2tJRCQgHBYtx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 917} outputId="fac3b0f5-158d-43b8-bd94-361bf1e37202"
d6 = pd.read_csv('localAthletes.csv')
d6
# + [markdown] id="svCc4jujBYt0" colab_type="text"
# Suppose we want that file in a data directory. Let's go ahead and create the directory and move the file there.
# + id="5QnO2pSDBYt0" colab_type="code" colab={}
# !mkdir data
# !mv localAthletes.csv data
# + [markdown] id="U8AGLyzGBYt7" colab_type="text"
# Now when we load the file we need to give more of a path:
# + id="1inZQEpNBYt8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 917} outputId="dced661d-3e97-40dc-b640-0d8a6cb7a72e"
d6 = pd.read_csv('data/localAthletes.csv')
d6
# + [markdown] id="7zbmMttPBYt_" colab_type="text"
# ## Missing Data
#
# In real machine learning tasks, we often encounter missing values.
#
# For example, suppose we didn't know Brittainey Raven's height. In that case our CSV file would start
#
# Name,Sport,Height,Weight
# <NAME>,Gymnastics,54,66
# Brittainey Raven,Basketball,,162
# <NAME>,Basketball,78,204
#
# with the double comma on the Brittainey line representing the missing data. When we read that file.
#
#
# + id="QKCKxI55BYuB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 917} outputId="80a84a04-f1cc-47cd-e97a-0cab6f458ffc"
df7 = pd.read_csv('https://raw.githubusercontent.com/zacharski/ml-class/master/data/athletesMissingValue.csv')
df7
# + [markdown] id="hTUtGnofBYuF" colab_type="text"
# We see that Brittainey Raven's height is now the floating point value NaN meaning Not a Number. This NaN value is used even in columns that do not contain floating point values. For example, in row 20 above, <NAME> has NaN in the Sport column. <NAME>, the developer of Pandas calls NaN a *sentinel* value that is easily detected and indicates a missing value.
#
# #### Adding a missing value by hand
# Suppose we didn't know the number of cylinders of a Ford F150:
#
# make | mpg | cylinders | HP | 0-60 |
# ---- | :---: | :---: | :---: | :---:
# Fiat | 38 | 4 | 157 | 6.9
# Ford F150 | 19 | - | 386 | 6.3
# Mazda 3 | 37 | 4 | 155 | 7.5
# Ford Escape | 27 | 4 | 245 | 7.1
# Kia Soul | 31 | 4 | 164 | 8.5
#
# In that case we can create a dataframe like:
#
# + id="eXOjWtaIBYuG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="3a944698-9807-4678-f546-6d7a8e49b623"
cars3 = {'make': Series(['Fiat 500', 'Ford F-150', 'Mazda 3', 'Ford Escape', 'Kia Soul']),
'mpg': Series([38, 19, 37, 27, 31]),
'cylinders': Series([4,np.nan, 4, 4, 4]),
'HP': [157, 386, 155, 245, 164],
'0-60': Series([6.9, 6.3, 7.5, 7.1, 8.5])}
carz = DataFrame(cars3)
carz
# + [markdown] id="tQOtc-GMBYuI" colab_type="text"
# where `np.nan` is Numpy's NaN. We can also use Python's `None`:
#
# + id="ouQ-AzBgBYuJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="d53b9876-4bef-48d6-aabb-b825d325ea40"
cars3 = {'make': Series(['Fiat 500', 'Ford F-150', 'Mazda 3', 'Ford Escape', 'Kia Soul']),
'mpg': Series([38, 19, 37, 27, 31]),
'cylinders': Series([4,None, 4, 4, 4]),
'HP': [157, 386, 155, 245, 164],
'0-60': Series([6.9, 6.3, 7.5, 7.1, 8.5])}
carz = DataFrame(cars3)
carz
# + [markdown] id="_3G-zz4HBYuL" colab_type="text"
# In addition to reading CSV files, there are many other ways of reading in data including from SQL databases, mongoDB, and webpages. See the Pandas documentation for details.
#
# <h3 style="color:red">Q2. Pima Indians Diabetes Dataset</h3>
# It is time to look at a new dataset, the Pima Indians Diabetes Data Set developed by the
# United States National Institute of Diabetes and Digestive and Kidney Diseases.
#
# The majority of the Pima people live on a reservation in Arizona.
#
# 
#
# Astonishingly, over 30% of Pima people develop diabetes. In contrast, the diabetes rate in
# the United States is around 12% and in China it is 4.2%. The country with the lowest rate of diabetes is Benin at 1.5%.
#
# Each instance in the dataset represents information about a Pima woman over the age of 21
# and belonged to one of two classes: a person who developed diabetes within five years, or a
# person that did not. There are eight attributes in addition to the column representing whether or not they developed diabetes:
#
#
# 1. The number of times the woman was pregnant
# 2. Plasma glucose concentration a 2 hours in an oral glucose tolerance test
# 3. Diastolic blood pressure (mm Hg)
# 4. Triceps skin fold thickness (mm)
# 5. 2-Hour serum insulin (mu U/ml)
# 6. Body mass index (weight in kg/(height in m)^2)
# 7. Diabetes pedigree function
# 8. Age
# 9. Whether they got diabetes or not (0 = no, 1 = yes)
#
#
# <span style="color:red">Please create a dataframe from the csv file at </span>
#
# https://raw.githubusercontent.com/zacharski/ml-class/master/data/pima-indians-diabetes.csv
#
# <span style="color:red">This file does not have a header row</span>
# + id="3Vhs4cAHBYuM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 419} outputId="96550c18-6da4-4d89-f4d1-ac2337128aae"
pimaDf = pd.read_csv('https://raw.githubusercontent.com/zacharski/ml-class/master/data/pima-indians-diabetes.csv', names = ['# of Times Pregnant','Plasma Glucose Concentration','Diastolic blood pressure(mm Hg)','Triceps skin fold thickness(mm)','2-Hour serum insulin(mu U/ml)','BMI','Diabetes Pedigree Function','Age','Has Diabetes'])
pimaDf
# + id="kBEUL1fKBYuV" colab_type="code" colab={}
## Your code here
# + [markdown] id="Sb2tJ0MeBYuX" colab_type="text"
# ## Accessing data in a DataFrame
# We can get a column of a DataFrame by using the column name:
# + id="nyBYvc0xBYua" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 510} outputId="a7b08965-4fc2-4c59-e7ae-588236cef56c"
athletes['Weight']
# + [markdown] id="fjiYttr2BYuc" colab_type="text"
# and we can get multiple columns by passing a list of column names
# + id="NXu6SNDxBYud" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 917} outputId="b5a02724-2bfe-4723-ac77-7407a3e18fd8"
athletes[['Weight', 'Height']]
# + [markdown] id="hZD1R7neBYug" colab_type="text"
# <h3 style="color:red">Q3. Pima Indians Diabetes Dataframe</h3>
#
# <span style="color:red">Suppose we want to create a new dataframe `pima2` from the original one but with only the columns</span>
#
#
#
# 1. Plasma glucose concentration a 2 hours in an oral glucose tolerance test
# 2. 2-Hour serum insulin (mu U/ml)
# 3. Body mass index (weight in kg/(height in m)^2)
# 4. Diabetes pedigree function
# 5. Age
# 6. Whether they got diabetes or not (0 = no, 1 = yes)
#
#
# <span style="color:red">How would you do so?</span>
#
# + id="eX-KjlBgBYuh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 419} outputId="8edeb5d4-e489-4890-9512-0e8895662445"
pimaDf2 = pimaDf[['Plasma Glucose Concentration', 'Diastolic blood pressure(mm Hg)' , '2-Hour serum insulin(mu U/ml)', 'BMI', 'Diabetes Pedigree Function', 'Age', 'Has Diabetes']]
pimaDf2
# + [markdown] id="yxc6oEG_BYuk" colab_type="text"
# ### Returning rows
#
# To get a row we can use the `loc` function
# + id="87BmJ489BYuk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="2bc6c901-e701-436f-e360-a3d4f6723ad3"
athletes.loc[0]
# + [markdown] id="jlZ-LijJBYup" colab_type="text"
# We can also get rows that match a specific criterion. For example:
# + id="2ThbM6EXBYuq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 328} outputId="36d2ff4f-3c4b-43ee-b508-bb12d14dfcd6"
basketballPlayers = athletes.loc[athletes['Sport'] == 'Basketball']
basketballPlayers
# + id="p8gnOaIzBYus" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 173} outputId="cff8f618-589a-42d2-d021-be46a6ab9c6e"
tallBasketballPlayers = athletes.loc[(athletes['Sport'] == 'Basketball') & (athletes['Height'] > 72)]
tallBasketballPlayers
# + [markdown] id="GbXCxt3ABYuw" colab_type="text"
# <span style="color:red">How would we create a new DataFrame that has all the athletes who weight under 100 pounds?</span>
# + id="_-8EAGO_BYux" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 359} outputId="99c4c4e0-8075-4162-bb5e-cb94d4663e3b"
lightPlayers = athletes.loc[(athletes['Weight'] <= 99)]
lightPlayers
# + [markdown] id="4ZrZ7nmCBYu0" colab_type="text"
# We can add columns to a DataFrame.
# For example the formula for Body Mass Index is
#
# $$BMI = \frac{weightInPounds}{heightInInches^2} \times 703 $$
# + id="DCTsKkZmBYu1" colab_type="code" colab={}
athletes['bmi'] = 703 *athletes['Weight'] / (athletes['Height']**2)
# + id="GMDaGprfBYvC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 917} outputId="f945c7ba-048f-464b-f62a-f296462fa36c"
athletes
# + [markdown] id="IwIL2ipMBYvH" colab_type="text"
# ## Descriptive Statistics on DataFrames
# One handy function is `describe`
# + id="QDIyjdJQBYvH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="654cd62a-1193-4c7a-cafb-6676c83f0fe3"
athletes.describe()
# + [markdown] id="eCGJn_uhBYvM" colab_type="text"
# Alternatively, I could ask for a specific statistic:
#
# + id="9NmHuQvDBYvM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="062574d8-b161-478b-f117-a2983d3f3a59"
athletes['Weight'].mean()
# + id="0G3ovVieBYvO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4a758142-64ea-4941-f0b5-77081f025008"
athletes.loc[athletes['Sport'] == 'Gymnastics' ]['Weight'].mean()
# + id="ee_yFp9QBYvR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5616bc61-3e16-4aa7-b091-8f6e2ff31b32"
athletes.loc[athletes['Sport'] == 'Basketball' ]['Weight'].mean()
# + [markdown] id="WOD7TqJABYvU" colab_type="text"
# <h3 style="color:red">Q4. Pima Indians</h3>
# <span style="color:red">I would like to fill out this little table:</span>
#
# x | Avg. BMI | Avg. Diabetes Pedigree | Avg. times pregnant | Avg. Plasma glucose |
# --- | :---: | :---: | :---: | :---: |
# Has Diabetes | | | |
# Doesn't have Diabetes | | | |
#
# <span style="color:red">Can you get this information (where 'Avg' refers to the mean)? So, for example, the first cell is the average Body Mass Index of people who have diabetes.</span> You don't need to write code to generate a table, I just want to see the values that would be put in such a table.
#
# + id="09rgXgMsBYvV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 272} outputId="f347727b-5dc5-4c14-94d5-de329cdefb71"
pimaStats = pimaDf[['BMI','# of Times Pregnant','Diabetes Pedigree Function','Plasma Glucose Concentration','Has Diabetes']]
pimaNoDiab = pimaStats.loc[pimaStats['Has Diabetes']==0]
print("Doesn't have Diabetes")
print(pimaNoDiab.mean())
print("")
pimaDiab = pimaStats.loc[pimaStats['Has Diabetes']==1]
print("Has Diabetes")
print(pimaDiab.mean())
# + [markdown] id="Uz-YSxY7BYvd" colab_type="text"
# ### Music Ratings
#
# Suppose I have customers of my vinyl record shop rate different artists
#
# |Customer | <NAME> | <NAME> | <NAME> | <NAME> | Ariana Grande |
# |:-----------|:------:|:------:|:---------:|:------:|:--------:|
# |Jake|5|-|5|2|2|
# |Clara|2|-|-|4|5|
# |Kelsey|5|5|5|2|-|
# |Angelica|2|3|-|5|5|
# |Jordyn|2|1|-|5|-|
#
# <h3 style="color:red">Q4. Ratings</h3>
# <span style="color:red">Create a dataframe called `ratings` from the CSV file</span>
#
# https://raw.githubusercontent.com/zacharski/ml-class/master/data/ratings.csv
# + id="gM1pDU5KBYvd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="6b5094f1-371b-49b8-d365-ee4091658ca6"
ratings = pd.read_csv('https://raw.githubusercontent.com/zacharski/ml-class/master/data/ratings.csv')
ratings
# + [markdown] id="KffLIWIYBYvg" colab_type="text"
# We can get the mean rating of each artist by:
#
# + id="ZX_wt4MjBYvh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="369cad8a-0fc3-486d-a281-ed6f2135429e"
ratings.mean()
# + [markdown] id="bloPERWIBYvj" colab_type="text"
# Many descriptive statistics functions take an optional parameter `axis` that tells which axis to reduce over. If we want the mean ratings for each **customer** instead of each artist we can do:
# + id="wrKDQg0qBYvj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="2c31e107-30ec-4293-fc6e-f1a11b9b1285"
ratings.mean(axis=1)
# + [markdown] id="peReLdGlBYvn" colab_type="text"
# Well, that was sort of unhelpful. We know that person 2 has a mean of 4.25, but it would be nice to use names as the indices for the rows instead of numbers. So right now we have the rows labeled 0, 1, 2, 3 ... but we would like them labeled *Jake, Clara, Kelsey* We can do so by creating a new dataframe:
# + id="UJ3oNSl9BYvn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="13b60bf7-4a61-49b3-fa61-2e8f383ef2a7"
ratings2 = ratings.set_index('Customer')
ratings2.mean(axis=1)
# + [markdown] id="gY4YT6HRBYvr" colab_type="text"
# *Note that while this looks like it erroneously creates an additional row called Customer, that is just the way the table is displayed*
#
# Sweet! So `axis=1` means reduce by rows and `axis=0` means reduce by columns:
# + id="gnY9qeJyBYvr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="d169fee7-db41-4233-872a-e02117b8aa0f"
ratings2.mean(axis=0)
# + [markdown] id="lIkf5cYEBYvu" colab_type="text"
# ## list of descriptive statistics
# (from the book *Python for Data Analysis*)
#
# Method | Description
# :-- | :--
# `count` | Number of non-NaN values
# `describe` | A set of common summary statistics
# `min, max` | compute minimum and maximum values
# `argmin, argmax` | compute index locations of minimum and maximum values
# `sum` | Sum the values
# `mean` | Mean of values
# `median` | Median of values
# `std` | Sample standard deviation
#
# So, for example, the lowest rating for each artist:
#
# + id="5Nbj-bNcBYvu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="2b2145ac-31ef-410b-8eb0-5eba82808080"
ratings2.min()
# + [markdown] id="gcGAKGBtBYvx" colab_type="text"
# <h3 style="color:red">Q5. Ratings 2</h3>
# <span style="color:red">What are the median ratings for each customer?</span>
# + id="pIQoahVvBYvx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="d4a42ead-922f-48e4-f35f-3e5046c80eaa"
ratings2.median()
# + [markdown] id="DUdMzVOhBYvz" colab_type="text"
# ## sorting
# To sort by the index we can use the `sort_index` method
# + id="ShRskIdQBYv0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 948} outputId="474b051c-fd4c-45eb-b636-c14670e9998b"
athlete = pd.read_csv('https://raw.githubusercontent.com/zacharski/ml-class/master/data/athletesNoHeader.csv', names=['Name', 'Sport', 'Height', 'Weight'])
a = athlete.set_index('Name')
a.sort_index()
# + [markdown] id="MfIpNd-3BYv2" colab_type="text"
# We can sort in reverse by
# + id="0DZd5uT8BYv3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 948} outputId="683d2688-834a-452b-b1b3-469c2e64d4d9"
a.sort_index(ascending=False)
# + [markdown] id="sNgx2SQABYv5" colab_type="text"
# We can use the `sort_values` method with the `by` argument to sort by a particular column:
# + id="4-1dISGFBYv6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 948} outputId="d58ba9d0-8797-47c0-839e-0baa762be8a0"
a.sort_values(by='Sport')
# + id="jTeclUQcBYv7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 948} outputId="d8511016-aa37-41cd-d7f1-64c5a1d1f616"
a.sort_values(by=['Height', 'Weight'])
# + [markdown] id="QCKyY3QdBYv-" colab_type="text"
# In that last example, we first sorted by height and then by weight.
#
# # The nearest neighbor Music example in Pandas
#
# Let's go back to our Numpy task that dealt with the following music data:
#
#
#
# | Guest | <NAME> | <NAME> | <NAME> | <NAME> | <NAME>|
# |---|---|---|---|---|---|
# | Ann | 4 | 5 | 2 | 1 | 3 |
# | Ben | 3 | 1 | 5 | 4 | 2|
# | Jordyn | 5 | 5 | 2 | 2 | 3|
# | Sam | 4 | 1 | 4 | 4 | 1|
# | Hyunseo | 1 | 1 | 5 | 4 | 1 |
# | Lauren | 3 | 1 | 5 | 5 | 5 |
# | Ahmed | 4 | 5 | 3 | 3 | 1 |
#
#
# I want to find out who is the most similar to Mikaela and who is most similar to Brandon, who rated the artists:
#
#
# | Guest | <NAME> | <NAME> | <NAME> | <NAME> | <NAME>|
# |---|---|---|---|---|---|
# | Mikaela | 3 | 2 | 4 | 5 |4 |
# | Brandon | 4 | 5 | 1 | 2 |3 |
#
# My first step is to read in the data and convert the guest name column as the index:
# + id="XVxJT250BYv_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="597f5315-42e4-4b85-e066-444fa2fe603e"
f = pd.read_csv('https://raw.githubusercontent.com/zacharski/ml-class/master/data/ratings2.csv')
fr = f.set_index('Guest')
fr
# + [markdown] id="l-VEUQSTBYwB" colab_type="text"
# Now let's load in Mikaela and Brandon in a different DataFrame
# + id="unWiJdL0BYwC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 142} outputId="d2d0e936-621b-4db0-8272-39e1ed3f867c"
mb = pd.read_csv('https://raw.githubusercontent.com/zacharski/ml-class/master/data/ratings3.csv')
mb.set_index('Guest', inplace=True)
mb
# + [markdown] id="Y5dkkbnYBYwD" colab_type="text"
# Let's work through the example step-by-step.
#
# First, can you set a variable named `mikaela` to equal the Mikaela row of `mb`?
# So something that starts with
#
# mikaela = mb???????????
# + id="TL3Fg2l7BYwE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="4ccfa52e-b06c-4b6c-9fd6-06a70d23d1ab"
mikaela = mb.iloc[0]
mikaela
# + [markdown] id="bsLYIYKyBYwG" colab_type="text"
# First, let's subtract Mikaela's ratings from the other people's:
# + id="ZPWWPpvWBYwH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="25bdf983-ee17-4166-89d7-7f799f51765f"
fr2 =fr.sub(mikaela)
fr2
# + [markdown] id="z9D9DKPMBYwJ" colab_type="text"
#
#
# Next, we will get the absolute values
# + id="SzyjucquBYwN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="64167ef6-8be6-468f-9db9-43e31eb56ab7"
fr3 = fr2.apply(np.abs)
fr3
# + [markdown] id="IGoGFuVcBYwP" colab_type="text"
# and sum the values in each row to compute the distances
# + id="dbZC0v-LBYwQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="bf6300c3-9543-4cad-dc2d-a61dba3346c3"
fr4 = fr3.sum(axis = 1)
fr4
# + [markdown] id="a4fPqYxsBYwS" colab_type="text"
# Finally, let's sort by the distance and get the three closest guests:
# + id="zPAq3KQVBYwS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="22d1a36e-477f-429f-a7ce-def9bd3d4a21"
fr4.sort_values().head(3)
# + [markdown] id="WClW5IxaBYwV" colab_type="text"
# So Lauren is the person most similar to Mikaela, followed by Ben and Sam.
#
# <h3 style="color:red">Q6. Find Closest</h3>
#
# <span style="color:red">Can you write a function nearestNeighbor that takes 2 arguments: a customers dataframe, and an array representing one customers ratings and returns the name and distance of the closest customer (using Euclidean Distance)?</span>
#
#
# + id="Yee8PYcnBYwV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="21cfb156-ddc8-4d07-e71a-28be47008621"
ahmed = [4, 5, 3, 3, 1]
def nearestNeighbor(ratings, customer):
subtract = ratings.sub(customer)
sub2 = subtract.apply(np.square)
summed = sub2.sum(axis =1)
summedDist = summed.apply(np.sqrt)
result = summedDist.sort_values().head(1)
return result
print(nearestNeighbor(fr, mikaela))
# TO DO
brandon = mb.iloc[1]
print(nearestNeighbor(fr, brandon))
# + [markdown] id="WTiUkvLZBYwZ" colab_type="text"
# <h3 style="color:red">Q7. Find Closest using Euclidean Distance</h3>
#
# <span style="color:red">Can you write a new function nearestNeighbor that takes 3 arguments</span>
#
# * ratings - an DataFrame representing all our customer ratings
# * customer - a DataFrame representing one customer's rating
# * metric - the value is either the string 'manhattan' or 'euclidean' representing which metric to use to compute the distance.
# a customers dataframe, and an array representing one customers ratings
#
# It returns the name and distance of the closest customer (using Euclidean Distance)?
#
# + id="3OTryWC9cUf4" colab_type="code" colab={}
def nearestNeighbor2(ratings, customer,metric):
if metric == 'euclidean':
subtract = ratings.sub(customer)
sub2 = subtract.apply(np.square)
summed = sub2.sum(axis =1)
summedDist = summed.apply(np.sqrt)
result = summedDist.sort_values().head(1)
elif metric == 'manhattan':
subtract = ratings.sub(customer)
sub2 = subtract.apply(np.abs)
summed = sub2.sum(axis =1)
result = summed.sort_values().head(1)
else:
print("The given metric cannot be calculated")
return result
# + id="2ZTTjzDtBYwc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="0d8d47c8-db13-45a2-daaf-922c28a3adb5"
print(nearestNeighbor2(fr, mikaela, metric='euclidean'))
# TO DO
# brandon = ...
print(nearestNeighbor2(fr, mb.loc['Brandon'], metric='euclidean'))
print(nearestNeighbor2(fr, mikaela, metric='manhattan'))
| labs/Copy_of_pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Sorry, this notebook has been moved to the new location:
#
# [https://github.com/atoti/notebooks/blob/master/notebooks/customer-churn/main.ipynb]([https://github.com/atoti/notebooks/blob/master/notebooks/customer-churn/main.ipynb])
| Communications/customer-churn/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **<NAME>**
# *1st June 2020*
#
# # Adding masked array
#
# ## Fix for periodic boundary conditions
# +
"""
Adding masked array (fix jumps)
<NAME>
2020-05-27
"""
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from celluloid import Camera # easy animation module
from IPython.display import HTML # to display in notebook
# PARAMETERS
r0 = [0, 0, 0] # initial position
v0 = [1, 2, 0] # initial velocity
B = [5, 0.02, 0.03] # magnetic field
E = [0.1, 0.5, 0] # electric field
q, m = 2, 0.7 # charge, mass
h = 0.05 # step size
end = 5 # t-value to stop integration
size = [5,0.3,1] # simulation dimensions
def lorentz(vel): # returns acceleration
return (q/m)*(E+np.cross(vel, B))
# RUNGE-KUTTA INTEGRATOR
def rk4(func, init1, init2, h, end):
"""
Takes the RHS of a 2nd-order ODE with initial conditions,
step size and end point, and integrates using the 4th-order
Runge-Kutta algorithm. Returns solution in an array.
r'' = f(t, r, v) where v = r'
func: the function to be integrated
init1: value of r at t=0
init2: value of v at t=0
h: step size
end: t-value to stop integrating
"""
steps = int(end/h) # number of steps
r = np.zeros((3, steps)) # empty matrix for solution
v = np.zeros((3, steps))
r[:,0] = init1 # inserting initial value
v[:,0] = init2
for i in range(0, steps-1):
k1r = h * v[:,i]
k1v = h * func(v[:,i])
k2r = h * (v[:,i] + 0.5*k1v)
k2v = h * func(v[:,i] + 0.5*k1v)
k3r = h * (v[:,i] + 0.5*k2v)
k3v = h * func(v[:,i] + 0.5*k2v)
k4r = h * (v[:,i] + k3v)
k4v = h * func(v[:,i] + k3v)
new_r = r[:,i] + (k1r + 2*k2r + 2*k3r + k4r) / 6
new_v = v[:,i] + (k1v + 2*k2v + 2*k3v + k4v) / 6
new_r[0] = new_r[0] % size[0]
new_r[1] = new_r[1] % size[1]
new_r[2] = new_r[2] % size[2]
r[:,i+1] = new_r
v[:,i+1] = new_v
return r
r = rk4(lorentz, r0, v0, h, end)
r_abdif = np.abs(np.diff(r, axis=1))
r_masked = np.ma.masked_where(r_abdif >= 0.02, r[:,1:])
fig = plt.figure()
camera = Camera(fig) # set up the 'camera' on the figure
ax = Axes3D(fig)
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.set_zlabel('z')
ax.set_xlim3d(np.amin(r[0]), np.amax(r[0])) # auto-scale axes
ax.set_ylim3d(np.amin(r[1]), np.amax(r[1]))
ax.set_zlim3d(np.amin(r[2]), np.amax(r[2]))
for i in range(r.shape[1]):
# plot data up to ith column and take snapshot
ax.plot3D(r_masked[0, :i], r_masked[1, :i], r_masked[2, :i], color='blue')
camera.snap()
animation = camera.animate(interval=30, blit=True)
HTML(animation.to_html5_video())
| dev-test/5a_masked_array.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Object Detection Notebook
# start with the first cells. In case you do not have `tensorflow-models/research` in your `PYTHONPATH` the import of `labinet` modules will fail.
# %load_ext autoreload
# %autoreload 1
# +
import os
import cv2
import sys
from matplotlib import pyplot as plt
from matplotlib.pyplot import imshow
from PIL import Image
import tensorflow as tf
import numpy as np
try:
from labinet.io_util import load_label_map
from labinet.io_util import load_model
import labinet.object_detect
import labinet.box
except ImportError:
# This part is only required to run the notebook
# directory when the module itself is not installed.
#
# If you have the module installed, just use "import labinet..."
import os
import inspect
# the .travis.yml is coded so that we execute tests from within test subdir. Relative to 'test' the .py is found in ../source/dev
cmd_subfolder = os.path.realpath(os.path.abspath(os.path.join(os.path.split(inspect.getfile(inspect.currentframe()))[0], "..")))
print(f"cmd_subfolder={cmd_subfolder}")
if cmd_subfolder not in sys.path:
sys.path.insert(0, cmd_subfolder)
from labinet.io_util import load_label_map
from labinet.io_util import load_model
import labinet.object_detect
import labinet.box
# %aimport labinet.io_util
# %aimport labinet.object_detect
# %aimport labinet.box
# %matplotlib inline
# -
# # Just a Test to access to camera
# you may skip the next 2 boxes to 'Object Detection via Webcam'
# access the cam (may fail, but ensure your cam is plugged, and we have a video device)
video = cv2.VideoCapture(0)
# Try capturing one frame
ret, frame = video.read()
if frame is None:
print("Error - did you connect your webcam?")
else:
cv2.imshow('object detection', cv2.resize(frame, (640,480)))
if cv2.waitKey(25) == ord('q'):
print(key_pressed)
pass
# it might happen that a window pops up (with content or without) that you have to close manually (force quit, 'X')
# or just execute the next cell
# close the window again
cv2.destroyAllWindows()
video.release()
# ## Object Detection via Webcam
# Actually it is a loop to capture picture via webcam and to Object detection
#
# Paths to model etc pp
MODEL_NAME = 'inference_graph' # the result from Step 6 Export Inference Graph
# Size, in inches, of the output images.
IMAGE_SIZE = (480, 640)
CWD_PATH = os.path.join(os.getcwd(),'..') # should become gitbase
NUM_CLASSES = 1
# model
PATH_TO_MODEL = os.path.join(CWD_PATH, MODEL_NAME, 'frozen_inference_graph.pb')
# label map
LABEL_MAP = os.path.join(CWD_PATH, 'data', 'object-detection.pbtxt')
# load model and labels
detection_graph = load_model(PATH_TO_MODEL)
categories, category_index = load_label_map(LABEL_MAP, NUM_CLASSES)
# prepare tensor dict for inference
tensor_dict = labinet.object_detect.get_tensor_dict_with_masks(IMAGE_SIZE[1], IMAGE_SIZE[0], detection_graph)
image_tensor = tensor_dict['image_tensor']
config = tf.ConfigProto()
config.gpu_options.allow_growth=True
# Prepare the Cam!
video = cv2.VideoCapture(0)
ret, frame = video.read()
if frame is None:
print("Error - did you connect your webcam?")
# ## Detection Loop
# just execute. Press 'q' to quit.
#
# ### common issues
# - camera not found (error on cv2.VideoCaputure()) or video.fread()
# - ensure you have something in `ls /dev/video*`
# - kernel dies with `Connection refused` ... `cannot open display`
# - ensure $DISPLAY is set
# - you might need to disable x-control (`xhost +`) / or set correct XAuthorities
# +
with tf.Session(graph=detection_graph, config=config) as sess:
# capture
while(True):
ret, frame = video.read()
#print(f"Captured frame.shape={frame.shape} - type(frame)={type(frame)}")
image_np_exp = np.expand_dims(frame, axis=0)
#print(f'np-frame.shape={image_np_exp.shape}')
# inference
output_dict = sess.run(tensor_dict, feed_dict={image_tensor: image_np_exp})
labinet.object_detect.convert_output_dict(output_dict)
# visualize boxes
image_with_boxes = labinet.object_detect.visualize_boxes_after_detection(frame, output_dict, category_index)
# show image
cv2.imshow('Detection Running...', cv2.resize(image_with_boxes,(IMAGE_SIZE[1],IMAGE_SIZE[0])))
#cv2.waitKey(25)
if cv2.waitKey(25) == ord('q'):
break
# -
video.release()
cv2.destroyAllWindows()
| notebook/object_detect_cam.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"is_executing": false}
from pyspark.sql import SparkSession
import pyspark.sql.functions as func
from pyspark.sql.types import StructType, StructField, StringType, LongType, DateType, DoubleType, BooleanType, \
ArrayType
# + pycharm={"name": "#%%\n", "is_executing": false}
spark = SparkSession.builder.appName("YelpHelp")\
.master("local")\
.config("spark.executor.memory", "16g")\
.config("spark.driver.memory", "16g")\
.getOrCreate()
schema2 = StructType([
StructField("business_id", StringType(), True),
StructField("date", StringType(), True)
])
dataset2 = spark.read.json("../yelp_dataset/yelp_academic_dataset_checkin.json", schema=schema2)
# + pycharm={"name": "#%%\n", "is_executing": false}
import datetime
def convert(x):
x = x.split(', ')
return x
def convert2(x):
x = [str(datetime.datetime.strptime(i, '%Y-%m-%d %H:%M:%S').date()) for i in x]
return x
dataset2 = dataset2.rdd.map(lambda x: (x[0], convert(x[1])))
dataset2 = dataset2.map(lambda x: (x[0], convert2(x[1])))
dataset2 = dataset2.toDF()
# + pycharm={"name": "#%%\n", "is_executing": false}
df_exploded = dataset2.withColumn('Checkin', func.explode('_2'))
df_exploded.show(5)
# + pycharm={"name": "#%%\n", "is_executing": false}
df_exploded = df_exploded.select(func.col("_1").alias("business_id"), func.col("Checkin").alias("checkin").cast(DateType()))
# + pycharm={"name": "#%%\n", "is_executing": false}
df_exploded.show(5)
# + pycharm={"name": "#%%\n", "is_executing": false}
df_exploded = df_exploded.withColumn('year', func.year("checkin")).repartition(10, "year")
df_exploded.write.partitionBy("year").json("../YelpDatasetYearly/CheckIns/yelp_academic_dataset_checkin")
# + pycharm={"name": "#%%\n"}
| Analysis/checkin.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import h5py
from sklearn.metrics import accuracy_score
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import confusion_matrix
from sklearn.metrics import precision_score, recall_score
from sklearn.metrics import cohen_kappa_score
from sklearn import preprocessing
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import PolynomialFeatures
from sklearn.model_selection import train_test_split
from sklearn.svm import LinearSVC
from sklearn.svm import SVC
from sklearn.tree import DecisionTreeClassifier
from sklearn.tree import export_graphviz
from sklearn.ensemble import BaggingClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import make_scorer
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.metrics import log_loss
from sklearn.model_selection import cross_val_score
import statsmodels.api as sm
import itertools
from scipy.stats import mode
import random
import matplotlib.pyplot as plt
plt.style.use('classic')
# %matplotlib inline
import seaborn as sns
sns.set()
import xgboost as xgb
import pandas as pd
from scipy import signal
from intervals import FloatInterval
# -
dataPath="C:\\Users\\i053131\\Desktop\\Epilepsie\\Dreem\\data\\raw\\"
h5filename="train.h5"
h5file= dataPath + h5filename
h5 = h5py.File(h5file, "r")
eeg_1 = pd.DataFrame(h5['eeg_1'][:])
eeg_2 = pd.DataFrame(h5['eeg_2'][:])
eeg_3 = pd.DataFrame(h5['eeg_3'][:])
eeg_4 = pd.DataFrame(h5['eeg_4'][:])
h5['eeg_1'][:]
fs=125
freq1, pxx_den1 = signal.periodogram(eeg_1, fs, axis=1)
PSD1= pd.DataFrame(data=pxx_den1, columns= freq1)
plt.semilogy(f, PSD1.iloc[0,:])
plt.show()
# +
from scipy import signal
data = eeg_1.iloc[1,:]
sf = 125 #hz
# Define window length (4 seconds)
win = 4 * sf
freqs, psd = signal.welch(data, sf, nperseg=win, scaling='density')
print(freqs.shape)
print(psd.shape)
# Plot the power spectrum
sns.set(font_scale=1.2, style='white')
plt.figure(figsize=(8, 4))
plt.plot(freqs, psd, color='k', lw=2)
#plt.plot(psd, color='k', lw=2)
plt.xlabel('Frequency (Hz)')
plt.ylabel('Power spectral density (V^2 / Hz)')
plt.ylim([0, psd.max() * 1.1])
plt.title("Welch's periodogram")
plt.xlim([0, 20])
sns.despine()
# -
i=0
sig = eeg_1.iloc[i,:]
f, Pxx_den = signal.periodogram(sig, fs)
plt.semilogy(f, Pxx_den)
plt.show()
b = f==fA
b.all()
print(Pxx_denA.shape)
print(len(Pxx_den))
print(len(Pxx_denA[0,]))
#print()
bb =Pxx_den== Pxx_denA[0,]
bb.all()
print(len(freq1))
print(PSD1.shape[1])
def giveIndex(freq, interval):
index= []
for f in freq:
if f in interval:
index.append(f)
return index
slow = FloatInterval.from_string('[0.5, 2.0)')
giveIndex(freq1, slow)
def generate_columns_names(L=['Slow', 'Delta', 'Theta', 'Alpha', 'Beta', 'Gamma', "Energy"]):
r = "["
for i in L:
if (i == L[-1]) :
r = r + "'" + str(i) + "']"
else:
r = r + "'" + str(i) + "',"
return eval(r)
generate_columns_names()
def periodigram_by_eeg_bandwidth(EEG, fs=125,
columnsName=['Slow', 'Delta', 'Theta', 'Alpha', 'Beta', 'Gamma', 'Energy']):
freqs, pxx_den1 = signal.periodogram(EEG, fs, axis=1)
Pgram= pd.DataFrame(data=pxx_den1, columns= freqs)
slow = FloatInterval.from_string('[0.5, 2.0)')
delta = FloatInterval.from_string('[2, 4.0)')
theta = FloatInterval.from_string('[4.0, 8.0)')
alpha = FloatInterval.from_string('[8.0, 16.0)')
beta = FloatInterval.from_string('[16.0, 32.0)')
gamma = FloatInterval.from_string('[32.0, 100.0)')
##above100Hz = FloatInterval.from_string('[100.0,)')
# no signal above100Hz
islow = giveIndex(freqs, slow)
idelta = giveIndex(freqs, delta)
itheta = giveIndex(freqs, theta)
ialpha = giveIndex(freqs, alpha)
ibeta = giveIndex(freqs, beta)
igamma = giveIndex(freqs, gamma)
columns = generate_columns_names(columnsName)
result = pd.DataFrame(columns = columns)
result.iloc[:, 0]= Pgram.loc[:,islow].sum(axis=1)
result.iloc[:, 1]= Pgram.loc[:,idelta].sum(axis=1)
result.iloc[:, 2]= Pgram.loc[:,itheta].sum(axis=1)
result.iloc[:, 3]= Pgram.loc[:,ialpha].sum(axis=1)
result.iloc[:, 4]= Pgram.loc[:,ibeta].sum(axis=1)
result.iloc[:, 5]= Pgram.loc[:,igamma].sum(axis=1)
result.iloc[:, 6]= Pgram.sum(axis=1)
#result["Slow"]= Pgram.loc[:,islow].sum(axis=1)
# result["Delta"]= Pgram.loc[:,idelta].sum(axis=1)
# result["Theta"]= Pgram.loc[:,itheta].sum(axis=1)
# result["Alpha"]= Pgram.loc[:,ialpha].sum(axis=1)
# result["Beta"]= Pgram.loc[:,ibeta].sum(axis=1)
# result["Gamma"]= Pgram.loc[:,igamma].sum(axis=1)
# result["Energy"] = Pgram.sum(axis=1)
return result
# %%time
fs=125
freq1, pxx_den1 = signal.periodogram(eeg_1, fs, axis=1)
PSD1= pd.DataFrame(data=pxx_den1, columns= freq1)
# %%time
e1= periodigram_by_eeg_bandwidth(eeg_1)
#e1.head()
e1.head()
# +
# %%time
def make_df():
dataPath="C:\\Users\\i053131\\Desktop\\Epilepsie\\Dreem\\data\\raw\\"
h5filename="train.h5"
h5file= dataPath + h5filename
h5 = h5py.File(h5file, "r")
eeg_1 = pd.DataFrame(h5['eeg_1'][:])
eeg_2 = pd.DataFrame(h5['eeg_2'][:])
eeg_3 = pd.DataFrame(h5['eeg_3'][:])
eeg_4 = pd.DataFrame(h5['eeg_4'][:])
eeg1 = periodigram_by_eeg_bandwidth(eeg_1,
columnsName=['eeg1_Slow', 'eeg1_Delta', 'eeg1_Theta', 'eeg1_Alpha', 'eeg1_Beta', 'eeg1_Gamma', 'eeg1_energy'])
eeg2 = periodigram_by_eeg_bandwidth(eeg_2,
columnsName=['eeg2_Slow','eeg2_Delta', 'eeg2_Theta', 'eeg2_Alpha', 'eeg2_Beta', 'eeg2_Gamma', 'eeg2_energy'])
eeg3 = periodigram_by_eeg_bandwidth(eeg_3,
columnsName=['eeg3_Slow','eeg3_Delta', 'eeg3_Theta', 'eeg3_Alpha', 'eeg3_Beta', 'eeg3_Gamma', 'eeg3_energy'])
eeg4 = periodigram_by_eeg_bandwidth(eeg_4,
columnsName=['eeg4_Slow','eeg4_Delta', 'eeg4_Theta', 'eeg4_Alpha', 'eeg4_Beta', 'eeg4_Gamma', 'eeg4_energy'])
eeg = pd.concat([eeg1, eeg2, eeg3, eeg4], axis=1, sort=False)
naif = pd.read_excel('C:\\Users\\i053131\\Desktop\\Epilepsie\\Dreem\\data\\interim\\featuresTrain.xlsx')
#eegO = pd.read_excel('C:\\Users\\i053131\\Desktop\\Epilepsie\\Dreem\\data\\interim\\spectrogram_eeg_features30Train.xlsx')
acc = pd.read_excel('C:\\Users\\i053131\\Desktop\\Epilepsie\\Dreem\\data\\interim\\acc_featuresTrain.xlsx')
pulse = pd.read_csv('C:\\Users\\i053131\\Desktop\\Epilepsie\\Dreem\\data\\interim\\pulse_featuresTrain.csv')
pulse = pulse.iloc[:, 1:]
pulse = pulse.drop(columns = ['max_r', 'min_r', 'max_ir', 'min_ir']) #remove features that are double
pulse = pulse.drop(columns =['BPMlessthan30_ir', 'BPMlessthan30_r']) # remove features with no importance
#eeg.drop(columns=["eeg1_Above100Hz0", "eeg2_Above100Hz0", "eeg3_Above100Hz0", "eeg4_Above100Hz0"], inplace=True)
#pulseAmp = pd.read_csv('C:\\Users\\i053131\\Desktop\\Epilepsie\\Dreem\\data\\interim\\pulse_ampTrain.csv')
#pulseAmp = pulseAmp.iloc[:, 1:]
df = pd.concat([eeg, pulse, naif], axis=1)
df.drop(columns=['std_eeg_1', 'std_eeg_2', 'std_eeg_3', 'std_eeg_4', 'RMSSD_r', 'IBI_r','RMSSD_ir', 'IBI_ir'],
inplace=True)
return df
df= make_df()
# +
#df = pd.concat([eeg, acc, pulse, naif], axis=1)
#df = pd.concat([eeg, pulse, naif], axis=1)
#df.drop(columns=['std_eeg_1', 'std_eeg_2', 'std_eeg_3', 'std_eeg_4', 'RMSSD_r', 'IBI_r','RMSSD_ir', 'IBI_ir'], inplace=True)
#df.drop(columns=['eeg4_energy', 'eeg3_energy', 'MAD_r', 'max_eeg_3'],inplace= True)
# drop less important features
#print("OK")
#df = pd.concat([eeg, naif], axis=1)
#df = eeg.copy()
#df["Y"] = naif.iloc[:, -1]
df.shape
# -
train = df.iloc[0:int(df.shape[0]*0.8), :]
test = df.iloc[int(df.shape[0]*0.8):, :]
train.shape[1]-1
eeg.shape[1]+ naif.shape[1]-1+ pulse.shape[1]
# Reference one 1 epoch (previous)
# - log loss = 0.7206771579062425
# - kappa = 0.6518430793293312
# - accuracy = 0.74366872005475
#
# (new) eeg + naif
# - log loss = 0.6990058512874828
# - kappa = 0.6549201342792562
#
# (new) eeg + pulse + naif
# - log loss = 0.7045746186176681
# - kappa = 0.6579936897373033
#
# (new) eeg + pulse + naif, dropping ['std_eeg_1', 'std_eeg_2', 'std_eeg_3', 'std_eeg_4', 'RMSSD_r', 'IBI_r','RMSSD_ir', 'IBI_ir']
# - log loss = 0.6960332634858694
# - kappa = 0.6587795975356928
#
# ??:
# -log loss = 0.7001175747835894
# - kappa = 0.6586589724834191
#
# dropping in addition ['eeg4_energy', 'eeg3_energy', 'MAD_r', 'max_eeg_3']
# - log loss = 0.7020145307848867
# - kappa = 0.6479186069682229
#
# (new) eeg + all
# - log loss = 0.695721407589258
# - kappa = 0.6574798196459137
#
# acc and pulse bring very little. it is in fact less good with Acc than without
#
#
# +
# %%time
errors = []
Lk = []
La = []
X = train.iloc[:,:-1]
y = train.iloc[:,-1]
print(y.unique())
X_test = test.iloc[:,:-1]
y_true = test.iloc[:,-1]
xbc = xgb.XGBClassifier(n_estimators = 170, random_state=42, learning_rate= 0.1, max_depth= 8, subsample= 0.7,
n_jobs=-2)
xbc.fit(X, y)
ll = log_loss(y_true, xbc.predict_proba(X_test))
errors.append(ll)
y_pred = xbc.predict(X_test)
k=cohen_kappa_score(y_true, y_pred)
a= accuracy_score(y_true, y_pred)
print("considering only one epoch")
print("log loss = ", ll)
print("kappa = ", k)
print("accuracy = ", a)
Lk.append(k)
La.append(a)
# -
fig, ax = plt.subplots(figsize=(20,20))
xgb.plot_importance(xbc,ax=ax, max_num_features=110)
# +
importances = xbc.feature_importances_
feature_importances = pd.DataFrame(importances, index = X.columns,
columns=['importance']).sort_values('importance', ascending=True)
feature_importances.head(8)
# -
low_imp =feature_importances.head(20).index
low_imp
# +
# %%time
errors = []
Lk = []
La = []
dfi =df.copy()
for col in low_imp:
print(col)
dfi.drop(columns=[col], inplace=True)
train = dfi.iloc[0:int(df.shape[0]*0.8), :]
test = dfi.iloc[int(df.shape[0]*0.8):, :]
X = train.iloc[:,:-1]
y = train.iloc[:,-1]
print(y.unique())
X_test = test.iloc[:,:-1]
y_true = test.iloc[:,-1]
xbc = xgb.XGBClassifier(n_estimators = 170, random_state=42, learning_rate= 0.1, max_depth= 8, subsample= 0.7,
n_jobs=-2)
xbc.fit(X, y)
ll = log_loss(y_true, xbc.predict_proba(X_test))
errors.append(ll)
y_pred = xbc.predict(X_test)
k=cohen_kappa_score(y_true, y_pred)
a= accuracy_score(y_true, y_pred)
print("dropping", col)
print("log loss = ", ll)
print("kappa = ", k)
print("accuracy = ", a)
Lk.append(k)
La.append(a)
# -
# without extra drop reference
# - log loss = 0.6960332634858694
# - kappa = 0.6587795975356928
d=16
s = 0
r = low_imp[s:d]
plt.plot(r, errors[s:d], label = "log loss")
plt.legend(loc='lower right')
plt.show()
#plt.plot(r, La, label = "accuracy")
plt.plot(r, Lk[s:d], label = "kappa")
plt.legend(loc='lower right')
plt.show()
# ## with several epochs
#
#
def reshape_n(df, n=5):
if ((df.shape[0] <n) or (n%2==0)):
print("Input error!")
return df
r = df.shape[0]
c = df.shape[1]
newColumns = []
for i in range(0, n):
for colName in df.columns:
newColumns.append(str(colName) + str(i+1))
result = pd.DataFrame(columns = newColumns, data = np.zeros((r, c*n)))
for i in range(n//2, r-n//2):
for j in range(0, n):
k = j-n//2
result.iloc[i,c*j:c*(j+1)]= df.iloc[i+k, :].values
return result
# +
eeg1 = periodigram_by_eeg_bandwidth(eeg_1,
columnsName=['eeg1_Slow', 'eeg1_Delta', 'eeg1_Theta', 'eeg1_Alpha', 'eeg1_Beta', 'eeg1_Gamma', 'eeg1_energy'])
eeg2 = periodigram_by_eeg_bandwidth(eeg_2,
columnsName=['eeg2_Slow','eeg2_Delta', 'eeg2_Theta', 'eeg2_Alpha', 'eeg2_Beta', 'eeg2_Gamma', 'eeg2_energy'])
eeg3 = periodigram_by_eeg_bandwidth(eeg_3,
columnsName=['eeg3_Slow','eeg3_Delta', 'eeg3_Theta', 'eeg3_Alpha', 'eeg3_Beta', 'eeg3_Gamma', 'eeg3_energy'])
eeg4 = periodigram_by_eeg_bandwidth(eeg_4,
columnsName=['eeg4_Slow','eeg4_Delta', 'eeg4_Theta', 'eeg4_Alpha', 'eeg4_Beta', 'eeg4_Gamma', 'eeg4_energy'])
eeg = pd.concat([eeg1, eeg2, eeg3, eeg4], axis=1, sort=False)
naif = pd.read_excel('C:\\Users\\i053131\\Desktop\\Epilepsie\\Dreem\\data\\interim\\featuresTrain.xlsx')
#eegO = pd.read_excel('C:\\Users\\i053131\\Desktop\\Epilepsie\\Dreem\\data\\interim\\spectrogram_eeg_features30Train.xlsx')
acc = pd.read_excel('C:\\Users\\i053131\\Desktop\\Epilepsie\\Dreem\\data\\interim\\acc_featuresTrain.xlsx')
pulse = pd.read_csv('C:\\Users\\i053131\\Desktop\\Epilepsie\\Dreem\\data\\interim\\pulse_featuresTrain.csv')
pulse = pulse.iloc[:, 1:]
pulse = pulse.drop(columns = ['max_r', 'min_r', 'max_ir', 'min_ir']) #remove features that are double
pulse = pulse.drop(columns =['BPMlessthan30_ir', 'BPMlessthan30_r']) # remove features with no importance
df = pd.concat([eeg, pulse, naif], axis=1)
df.drop(columns=['std_eeg_1', 'std_eeg_2', 'std_eeg_3', 'std_eeg_4', 'RMSSD_r', 'IBI_r','RMSSD_ir', 'IBI_ir'],
inplace=True)
# +
errors = []
Lk = []
La = []
train = df.iloc[0:int(df.shape[0]*0.8), :]
test = df.iloc[int(df.shape[0]*0.8):, :]
X = train.iloc[:,:-1]
y = train.iloc[:,-1]
X_test = test.iloc[:,:-1]
y_true = test.iloc[:,-1]
#xbc = xgb.XGBClassifier(n_estimators = 180, random_state=42, learning_rate= 0.1, max_depth= 8, subsample= 0.7, n_jobs=-2)
xbc = xgb.XGBClassifier(n_estimators = 180, random_state=42, learning_rate= 0.1, max_depth= 8, subsample= 0.7,
n_jobs=-2, reg_lambda=5)
xbc.fit(X, y)
ll = log_loss(y_true, xbc.predict_proba(X_test))
errors.append(ll)
y_pred = xbc.predict(X_test)
k=cohen_kappa_score(y_true, y_pred)
a= accuracy_score(y_true, y_pred)
print("considering only one epoch")
print("log loss = ", ll)
print("kappa = ", k)
print("accuracy = ", a)
Lk.append(k)
La.append(a)
r = [3, 5, 7, 9, 11, 13]
for i in r:
train5 = reshape_n(train.iloc[:,:-1], i)
train5["Y"]= train.iloc[:,-1]
for j in range(0, i//2):
train5.drop([train5.shape[0]-(i+1)], inplace=True)
test5 = reshape_n(test.iloc[:,:-1], i)
test5["Y"]= test.iloc[:,-1].values
for j in range(0, i//2):
test5.drop([test5.shape[0]-(i+1)], inplace=True)
X = train5.iloc[:,:-1]
y = train5.iloc[:,-1]
X_test = test5.iloc[:,:-1]
y_true = test5.iloc[:,-1]
xbc = xgb.XGBClassifier(n_estimators = 180, random_state=42, learning_rate= 0.1, max_depth= 8, subsample= 0.7,
n_jobs=-2, reg_lambda=5)
xbc.fit(X, y)
ll = log_loss(y_true, xbc.predict_proba(X_test))
errors.append(ll)
y_pred = xbc.predict(X_test)
k=cohen_kappa_score(y_true, y_pred)
a= accuracy_score(y_true, y_pred)
print("considering ", i, " epochs")
print("log loss = ", ll)
print("kappa = ", k)
print("accuracy = ", a)
Lk.append(k)
La.append(a)
# +
r =[1, 3, 5, 7, 9, 11, 13]
plt.plot(r, errors, label = "log loss")
plt.legend(loc='lower right')
plt.show()
plt.plot(r, Lk, label = "kappa")
plt.legend(loc='lower right')
plt.show()
# -
train.shape
# Let's go for N=5
# +
errors = []
Lk = []
La = []
train = df.iloc[0:int(df.shape[0]*0.8), :]
test = df.iloc[int(df.shape[0]*0.8):, :]
train5 = reshape_n(train.iloc[:,:-1], 5)
train5["Y"]= train.iloc[:,-1]
for j in range(0, i//2):
train5.drop([train5.shape[0]-(i+1)], inplace=True)
test5 = reshape_n(test.iloc[:,:-1], 5)
test5["Y"]= test.iloc[:,-1].values
for j in range(0, i//2):
test5.drop([test5.shape[0]-(i+1)], inplace=True)
X = train5.iloc[:,:-1]
y = train5.iloc[:,-1]
X_test = test5.iloc[:,:-1]
y_true = test5.iloc[:,-1]
r = [3, 4, 5, 6, 7, 8, 9, 10]
for i in r:
xbc = xgb.XGBClassifier(n_estimators = 180, random_state=42, learning_rate= 0.1, max_depth= i, subsample= 0.7,
n_jobs=-2, reg_lambda=5)
xbc.fit(X, y)
ll = log_loss(y_true, xbc.predict_proba(X_test))
errors.append(ll)
y_pred = xbc.predict(X_test)
k=cohen_kappa_score(y_true, y_pred)
a= accuracy_score(y_true, y_pred)b
print("considering ", i, " epochs")
print("log loss = ", ll)
print("kappa = ", k)
print("accuracy = ", a)
Lk.append(k)
La.append(a)
# +
plt.plot(r, errors, label = "log loss")
plt.legend(loc='lower right')
plt.show()
plt.plot(r, Lk, label = "kappa")
plt.legend(loc='lower right')
plt.show()
# +
errors = []
Lk = []
La = []
train = df.iloc[0:int(df.shape[0]*0.8), :]
test = df.iloc[int(df.shape[0]*0.8):, :]
train5 = reshape_n(train.iloc[:,:-1], 5)
train5["Y"]= train.iloc[:,-1]
for j in range(0, i//2):
train5.drop([train5.shape[0]-(i+1)], inplace=True)
test5 = reshape_n(test.iloc[:,:-1], 5)
test5["Y"]= test.iloc[:,-1].values
for j in range(0, i//2):
test5.drop([test5.shape[0]-(i+1)], inplace=True)
X = train5.iloc[:,:-1]
y = train5.iloc[:,-1]
X_test = test5.iloc[:,:-1]
y_true = test5.iloc[:,-1]
r = range(50, 210, 10)
for i in r:
xbc = xgb.XGBClassifier(n_estimators = i, random_state=42, learning_rate= 0.1, max_depth= 5, subsample= 0.7,
n_jobs=-2, reg_lambda=5)
xbc.fit(X, y)
ll = log_loss(y_true, xbc.predict_proba(X_test))
errors.append(ll)
y_pred = xbc.predict(X_test)
k=cohen_kappa_score(y_true, y_pred)
a= accuracy_score(y_true, y_pred)
print("considering ", i, " epochs")
print("log loss = ", ll)
print("kappa = ", k)
print("accuracy = ", a)
Lk.append(k)
La.append(a)
# +
plt.plot(r, errors, label = "log loss")
plt.legend(loc='lower right')
plt.show()
plt.plot(r, Lk, label = "kappa")
plt.legend(loc='lower right')
plt.show()
# +
errors = []
Lk = []
La = []
train = df.iloc[0:int(df.shape[0]*0.8), :]
test = df.iloc[int(df.shape[0]*0.8):, :]
train5 = reshape_n(train.iloc[:,:-1], 5)
train5["Y"]= train.iloc[:,-1]
for j in range(0, i//2):
train5.drop([train5.shape[0]-(i+1)], inplace=True)
test5 = reshape_n(test.iloc[:,:-1], 5)
test5["Y"]= test.iloc[:,-1].values
for j in range(0, i//2):
test5.drop([test5.shape[0]-(i+1)], inplace=True)
X = train5.iloc[:,:-1]
y = train5.iloc[:,-1]
X_test = test5.iloc[:,:-1]
y_true = test5.iloc[:,-1]
r = range(10)
for i in r:
xbc = xgb.XGBClassifier(n_estimators = 160, random_state=42, learning_rate= 0.1, max_depth= 5, subsample= 0.7,
n_jobs=-2, reg_lambda=i)
xbc.fit(X, y)
ll = log_loss(y_true, xbc.predict_proba(X_test))
errors.append(ll)
y_pred = xbc.predict(X_test)
k=cohen_kappa_score(y_true, y_pred)
a= accuracy_score(y_true, y_pred)
print("considering ", i, " epochs")
print("log loss = ", ll)
print("kappa = ", k)
print("accuracy = ", a)
Lk.append(k)
La.append(a)
# +
plt.plot(r, errors, label = "log loss")
plt.legend(loc='lower right')
plt.show()
plt.plot(r, Lk, label = "kappa")
plt.legend(loc='lower right')
plt.show()
# +
# %%time
errors = []
Lk = []
La = []
train = df.iloc[0:int(df.shape[0]*0.8), :]
test = df.iloc[int(df.shape[0]*0.8):, :]
train5 = reshape_n(train.iloc[:,:-1], 5)
train5["Y"]= train.iloc[:,-1]
for j in range(0, i//2):
train5.drop([train5.shape[0]-(i+1)], inplace=True)
test5 = reshape_n(test.iloc[:,:-1], 5)
test5["Y"]= test.iloc[:,-1].values
for j in range(0, i//2):
test5.drop([test5.shape[0]-(i+1)], inplace=True)
X = train5.iloc[:,:-1]
y = train5.iloc[:,-1]
X_test = test5.iloc[:,:-1]
y_true = test5.iloc[:,-1]
# +
ck_score = make_scorer(cohen_kappa_score, greater_is_better=True)
scoring = {'neg_log_loss': 'neg_log_loss', "kappa": ck_score}
xbc = xgb.XGBClassifier(n_estimators = 2, random_state=42, learning_rate= 0.1, max_depth= 5, subsample= 0.7,
n_jobs=-2, reg_lambda=5, eval_set=[(X, y), (X_test, y_true)],verbose=True,
#eval_metric='logloss'
eval_metric=['merror']
)
xbc.fit(X, y)
ll = log_loss(y_true, xbc.predict_proba(X_test))
errors.append(ll)
y_pred = xbc.predict(X_test)
k=cohen_kappa_score(y_true, y_pred)
a= accuracy_score(y_true, y_pred)
print("considering ", i, " epochs")
print("log loss = ", ll)
print("kappa = ", k)
print("accuracy = ", a)
Lk.append(k)
La.append(a)
# -
er = xbc.evals_result()
#evals_result = clf.evals_result()
# +
# %%time
errors = []
Lk = []
La = []
train = df.iloc[0:int(df.shape[0]*0.8), :]
test = df.iloc[int(df.shape[0]*0.8):, :]
train5 = reshape_n(train.iloc[:,:-1], 5)
train5["Y"]= train.iloc[:,-1]
for j in range(0, i//2):
train5.drop([train5.shape[0]-(i+1)], inplace=True)
test5 = reshape_n(test.iloc[:,:-1], 5)
test5["Y"]= test.iloc[:,-1].values
for j in range(0, i//2):
test5.drop([test5.shape[0]-(i+1)], inplace=True)
X = train5.iloc[:,:-1]
y = train5.iloc[:,-1]
X_test = test5.iloc[:,:-1]
y_true = test5.iloc[:,-1]
r = range(100, 1000, 50)
for i in r:
xbc = xgb.XGBClassifier(n_estimators = i, random_state=42, learning_rate= 0.1, max_depth= 5, subsample= 0.7,
n_jobs=-2, reg_lambda=5)
xbc.fit(X, y)
ll = log_loss(y_true, xbc.predict_proba(X_test))
errors.append(ll)
y_pred = xbc.predict(X_test)
k=cohen_kappa_score(y_true, y_pred)
a= accuracy_score(y_true, y_pred)
print("considering ", i, " epochs")
print("log loss = ", ll)
print("kappa = ", k)
print("accuracy = ", a)
Lk.append(k)
La.append(a)
# +
plt.plot(r, errors, label = "log loss")
plt.legend(loc='lower right')
plt.show()
plt.plot(r, Lk, label = "kappa")
plt.legend(loc='lower right')
plt.show()
# +
# %%time
errors = []
Lk = []
La = []
df = make_df()
train = df.iloc[0:int(df.shape[0]*0.8), :]
test = df.iloc[int(df.shape[0]*0.8):, :]
train5 = reshape_n(train.iloc[:,:-1], 5)
train5["Y"]= train.iloc[:,-1]
for j in range(0, i//2):
train5.drop([train5.shape[0]-(i+1)], inplace=True)
test5 = reshape_n(test.iloc[:,:-1], 5)
test5["Y"]= test.iloc[:,-1].values
for j in range(0, i//2):
test5.drop([test5.shape[0]-(i+1)], inplace=True)
X = train5.iloc[:,:-1]
y = train5.iloc[:,-1]
X_test = test5.iloc[:,:-1]
y_true = test5.iloc[:,-1]
r = range(250, 280, 10)
#350 ref kappa = 0.7273361761827176
#
for i in r:
xbc = xgb.XGBClassifier(n_estimators = i, random_state=42, learning_rate= 0.1, max_depth= 5, subsample= 0.7,
n_jobs=-2, reg_lambda=5)
xbc.fit(X, y)
ll = log_loss(y_true, xbc.predict_proba(X_test))
errors.append(ll)
y_pred = xbc.predict(X_test)
k=cohen_kappa_score(y_true, y_pred)
a= accuracy_score(y_true, y_pred)
print("considering ", i, " epochs")
print("log loss = ", ll)
print("kappa = ", k)
print("accuracy = ", a)
Lk.append(k)
La.append(a)
# +
plt.plot(r, errors, label = "log loss")
plt.legend(loc='lower right')
plt.show()
plt.plot(r, Lk, label = "kappa")
plt.legend(loc='lower right')
plt.show()
# -
X.shape
| notebooks/periodigram.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # PC lab 13: PyTorch & Convolutional Neural Networks
# ----
# <img src="img/school_chop.jpg" style="width:100%">
# ## Introduction
# Convolutional neural networks caused a major step forward in the performance of image recognition. These networks are mostly identical to standard neural networks, in which features are first learned through multiple (layers of) convolutions. Obtained features are subsequently used as the input for a standard neural network, often performing a classification problem.
# ### Convolution
# A convolution is the iteration of a kernel with size $ M \times N $ over a given input $ \textbf{X} $, performing a 2D linear combination of the weights $ W $ of the kernel with the overlapping area of the input. For a normal convolution with single striding and no padding, the output $ y_{ij} $ is equal to:
#
# $$ y_{ij} = \sum_{a=0}^{m-1} \sum_{b=0}^{n-1} W_{ab} x_{(i+a)(j+b)} $$
#
# During a convolution, the kernels slides over the input image to obtain a new image of outputs. The stride of a kernel defines the horizontal and vertical stepsize during iteration. Input data can be padded with multiple layers of a zero-filled border, increasing the output dimensions.
# **convolution step with M,N = 3; stride = 2 and padding layer of 1**
# <img src="img/kern_mult.gif">
# **several other examples. An extended explanation on all types of convolutions can be found [here](https://towardsdatascience.com/types-of-convolutions-in-deep-learning-717013397f4d)**
# <img src="img/conv_ex_all.gif">
# ## Convolutional neural network
# A convolutional neural network usually processes the image with **multiple sequential convolutions**. For each layer, the kernel is evaluated for **all channels** of the input data. It is important to understand that the output depth is correlated to the amount of different kernels, or features, every node has been initialized with. The kernel, although often depicted as only evaluating one layer, actually takes **the sum of all layers (channels)** to obtain an output.
# <img src="img/conv_ex_3.gif" style="width:100%">
# <img src="img/conv_schema.jpg" style="width:100%">
# A classic example of a convolutional neural network applies an activation function (e.g. **ReLU**) on the output of every convolutional layer, after which the activation signals are **maximum pooled**. Maximum pooling can reduces the amount of parameters present in a neural network, which reduces overfitting and computational burden. Maximum pooling is also initialized with specific arguments such as kernel size, stride and padding.
#
# <img src="img/max_pool.png">
# The last layers of every convolutional neural netwerk always consist of several fully connected layers. The mathematical description of these layers were discussed in the previous PC lab. One can interpret the convolutional layers as the section of the netwerk in which features training is done (edges, contours, contrasts,...). These are used as inputs for the fully connected neural netwerk, which combines these features to train the classifier.
# **The softmax function** takes the n-dimensional output of the model and rescales these values to probabilities that sum up to one. It is typically used as the output layer for multiclass classification.
#
# $$ \sigma(\hat{y})_j = \frac{e^{\hat{y}_j}}{\sum_{k=1}^{K} e^{\hat{y}_k}} $$
# <hr>
# <img src="img/pytorch.png" style="width:40%;float:left">
# PyTorch is a python package that provides two high-level features:
#
# - Tensor computation (like numpy) with **strong GPU acceleration**
# - **Deep Neural Networks** built on a tape-based autograd system
#
# <div class="alert alert-warning">
# <h2>Installing PyTorch</h2>
# <p>Run from terminal. In the same environment as scikit learn has been installed.</p>
# <p>install on <b>LINUX</b>:</p>
# <code>conda install pytorch-cpu torchvision-cpu -c pytorch</code>
# <p>install on <b>WINDOWS</b>:</p>
# <code>conda install pytorch-cpu -c pytorch
# pip3 install torchvision</code>
# <p>install on <b>MAC</b>:</p>
# <code>conda install pytorch torchvision -c pytorch</code>
# </div>
import numpy as np
import torch
# Torch can be approached in a similar way as numpy, where data is mainly exchanged through **Tensor** objects instead of **numpy** objects.
torch.zeros(4,3)
#
# Switching between numpy arrays and tensors is easy:
torch.zeros(4,3).numpy()
torch.from_numpy(np.zeros((4,3)))
# <div class="alert alert-warning">
# <h2>INFO</h2>
# <p>PyTorch has many functions and objects that behave similar to the ones found in the numpy module. </p>
# <ul>
# <li><code>torch.zeros()</code></li>
# <li><code>torch.ones()</code></li>
# <li><code>torch.arange()</code></li>
# <li><code>torch.Tensor.min()</code></li>
# <li><code>torch.Tensor.shape</code></li>
# <li>...</li>
# </ul>
# </div>
#
# ----
# <div class="alert alert-info">
# <h1>Structure of the exercises.</h1>
# <p>During this PC-lab you will be introduced with an examplory workflow when training a predictive model for image recognition. More specifically, we will create a <b>convolutional neural network</b> to <b>recognize vehicles from animals</b> using the <b>CIFAR-10</b> dataset in <b>PyTorch</b>. Most of the code has been written out already, leaving time to build further upon the existing codebase to improve results or add interesting features.
#
# </div>
#
# ----
# ## 1. Data loading and preprocessing
# <img src="img/CIFAR10.jpg" width='25%' align="right">
#
# # CIFAR-10
#
# - 60000 $32 \times 32$ colour images
# - 50000 training images + 10000 validation images
# - 10 classes $\rightarrow$ 6000 images per class
#
#
# Collected for **MSc thesis**
#
# <ul>
# <a href="learning-features-2009-TR.pdf">Learning Multiple Layers of Features from Tiny Images</a>, <NAME>, 2009.
# </ul>
# The **CIFAR-10** is next to **MNIST** an established computer-vision dataset which has been extensively used to evaluate the performance of a wide variety machine learning techniques. A list featuring some papers using CIFAR-10 can be found [here](http://rodrigob.github.io/are_we_there_yet/build/classification_datasets_results.html#CIFAR-10). To load the CIFAR-10 dataset into our environment, we can use the pytorch module.
# +
import torch
import numpy as np
import torchvision
import torchvision.transforms as transforms
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True,
download=True, transform=transform)
validationset = torchvision.datasets.CIFAR10(root='./data', train=False,
download=True, transform=transform)
validation_loader = torch.utils.data.DataLoader(validationset, batch_size=32,
shuffle=False, num_workers=2)
train_loader = torch.utils.data.DataLoader(trainset, batch_size=32,
shuffle=True, num_workers=2)
classes = ('plane', 'car', 'bird', 'cat',
'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
binary_classes = np.array(['vehicle', 'animal'])
# -
# ## 2. Building the network
# Neural networks are initialized through the use of class objects. Many of the functionalities necessary to create [**all types of neural networks**](http://www.asimovinstitute.org/neural-network-zoo/) have [**already been implemented**](http://pytorch.org/docs/master/nn.html). The following code creates a neural network with two convolutions and two fully connected layers. The network is identical to the neural net shown at the beginning of the notebook (with the car displayed), albeit with the addition of one fully connected layer.
# <div class="alert alert-success">
# <h2>Exercise:</h2>
# <p>Calculate the input nodes of the the first fully connected layer <code>self.fc1</code>. This is the first argument of the function <code>nn.Linear()</code>. The batch size does not influence this value. Notice that <code>x.view(-1, ...)</code> assigns the different batches to the first dimension.</p>
# </div>
# +
import torch
from torch.autograd import Variable
import torch.nn as nn
import torch.nn.functional as F
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
# 3 input image channels, 16 output channels, 5x5 square convolution
self.conv1 = nn.Conv2d(3, 16, (5,5))
# 16 input image channels, 32 output channels, 5x5 square convolution
self.conv2 = nn.Conv2d(16, 32, (5,5))
# --- SOLUTION
self.fc1 = nn.Linear(32 * 5 * 5, 64)
# --- SOLUTION
self.fc2 = nn.Linear(64, 2)
def forward(self, x):
# Max pooling over a (2, 2) window + RELU
x = F.max_pool2d(F.relu(self.conv1(x)), (2, 2))
# Max pooling over a (2, 2) window + RELU
x = F.max_pool2d(F.relu(self.conv2(x)), 2)
# Flatten array
# --- SOLUTION
x = x.view(-1, 32*5*5)
# --- SOLUTION
x = F.relu(self.fc1(x))
x = self.fc2(x)
return x
# -
# Notice the similarities between the creation of this neural network and the one created in the previous lab. To be able to call a variant of this network, we can choose to add parameter values to `def __init__(self, .....)`, such as kernel sizes, or the total amount of nodes in each fully connected layer.
# ### Backpropagation
# Pytorch offers an easy way to obtain gradients, where no explicit calculation of the derivatives is required. This is done through `autograd.Variable`.
# <div class="alert alert-warning">
# <p><code>autograd.Variable</code> is the central class of the package. It wraps a <code>Tensor</code>, and supports nearly all operations defined on it. Once you finish your computation you can call <code>.backward()</code> and have all the gradients computed automatically.</p>
# <p><code>Variable</code> and <code>Function</code> are interconnected and build up an acyclic graph, which encodes a complete history of computation. Each variable has a <code>.grad_fn</code> attribute that references a <code>Function</code> that has created the <code>Variable</code></p>
# </div>
x = Variable(torch.ones(2, 2), requires_grad=True)
print(x)
y = x + 2
z = y**4
out = z.mean()
out
x.grad
out.backward()
x.grad
# Switching between a Variable and Tensor is also possible
x = x.data
print(x)
x = Variable(x)
print(x)
# ## 3. Training the model
# ### Helper functions
#
# The next step is to train the model. The train_loader and validation_loader object is used to obtain data batches of fixed size. The `fit()` function trains the model for a specified amount of epochs while storing data obtained during training with the `logger()` class. `convert_to_binary_class()` is a custom function defined to convert the dataset into a binary class problem (vehicles <-> animals). This is merely done to make training faster. To train with the original classes, simply comment out the line `y_batch = convert_to_binary_class(y_batch)` twice, as found in the `fit()` function. Furthermore, the model has to be changed to have 10 outputs.
# +
def convert_to_binary_class(y_batch):
labels = []
for label in y_batch:
if label in [0,1,8,9]:
labels.append(0)
else:
labels.append(1)
return torch.LongTensor(labels)
def fit(model, train_loader, criterion, optimizer, log, epochs=20):
epoch = 0 # set starting epoch
while epoch<epochs:
print("\nepoch {}".format(epoch))
epoch +=1
model.train()
for i, data in enumerate(train_loader): # iterate randomized batches
optimizer.zero_grad()
X_batch, y_batch = data
y_batch = convert_to_binary_class(y_batch)
X_batch, y_batch = Variable(X_batch), Variable(y_batch)
optimizer.zero_grad()
y_hat = model.forward(X_batch)
loss = criterion(y_hat, y_batch)
loss.backward() # Calculate gradient
optimizer.step() # Update weights using defined optimizer
log.log_metrics(y_batch.data.numpy(), y_hat.data.numpy(), loss.item())
if (i%100 == 1):
log.output_metrics()
# Repeat this process for the validation dataset
model.eval()
for i, data in enumerate(validation_loader, 0):
X_batch, y_batch = data
y_batch = convert_to_binary_class(y_batch)
X_batch, y_batch = Variable(X_batch), Variable(y_batch)
y_hat = model.forward(X_batch)
loss = criterion(y_hat, y_batch)
log.log_metrics(y_batch.data.numpy(), y_hat.data.numpy(), loss.item(), validation=True)
log.output_metrics(validation=True)
# -
# <div class="alert alert-warning">
# <h2><code>class logger()</code> </h2>
# <p><code>logger()</code> has been implemented as a convenient way to store model metrics throughout the training process. An object of this class can be created before training and is used for calculating, storing, printing and plotting model metrics. The object does only store the metrics as defined at initialization. Feel free to add yor own metrics to the class.</p>
# </div>
# +
from sklearn.metrics import roc_auc_score
import matplotlib.pyplot as plt
# %matplotlib inline
class logger(object):
def __init__(self, metrics, max_i):
self.i = [0,0]
self.max_i = max_i
self.log_loss, self.log_auc, self.log_acc = False, False, False
self.metrics = {"train":{}, "validation":{}}
if "loss" in metrics:
self.log_loss = True
self.metrics["train"].update({"loss":[0]})
self.metrics["validation"].update({"loss":[0]})
if "AUC" in metrics:
self.log_auc = True
self.metrics["train"].update({"auc":[0]})
self.metrics["validation"].update({"auc":[0]})
if "acc" in metrics:
self.log_acc = True
self.metrics["train"].update({"acc":[0]})
self.metrics["validation"].update({"acc":[0]})
def log_metrics(self, y_true, y_hat, loss, validation=False):
if validation:
sw = 1
sw_str = "validation"
else:
sw = 0
sw_str = "train"
self.i[sw] += 1
if self.log_loss:
update = (self.metrics[sw_str]["loss"][-1]*(self.i[sw]-1)+loss)/self.i[sw]
self.metrics[sw_str]["loss"].append(update)
if self.log_auc:
auc = roc_auc_score(y_true, y_hat[:,1])
update = (self.metrics[sw_str]["auc"][-1]*(self.i[sw]-1)+auc)/self.i[sw]
self.metrics[sw_str]["auc"].append(update)
if self.log_acc:
acc = sum(y_hat.argmax(axis=1) == y_true)/len(y_true)
update = (self.metrics[sw_str]["acc"][-1]*(self.i[sw]-1)+acc)/self.i[sw]
self.metrics[sw_str]["acc"].append(update)
def output_metrics(self, validation=False):
data = "validation" if validation else "train"
if validation:
print_str = "\n{:<10s}:\t100.0%".format(data)
else:
print_str = "\r{:<10s}:".format(data)
print_str += "\t{:4.2f}%".format((self.i[0]%self.max_i)/self.max_i*100)
for k, v in self.metrics[data].items():
print_str += "\t{}: {:5.3f}".format(k, v[-1])
print(print_str, end = "")
def plot_metrics(self):
fig, axes = plt.subplots(len(self.metrics["train"]),
2, figsize=(12,6*len(self.metrics["train"])))
for i, dict_0 in enumerate(self.metrics.items()):
for j, dict_1 in enumerate(dict_0[1].items()):
axes[j,i].plot(range(len(dict_1[1][1:])),dict_1[1][1:])
axes[j,i].set_title("{} {}".format(dict_0[0], dict_1[0]))
# -
# ### Training
# Now that we have loaded our data, defined our convolutional neural network, and created all the functions necessary for training, the actual training process can start. For this instance, we will use the cross entropy loss to optimize our model. Notice how `nn.CrossEntropyLoss()` incorporaties the softmax function on the inputs. [Adam](http://sebastianruder.com/optimizing-gradient-descent/index.html#adam) is used to determine the step size using the gradient of the loss with respect to the weights. Adam is currently often considered the best option for this task.
# +
import torch.optim as optim
from sklearn.metrics import confusion_matrix
model = Net()
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters(), lr=0.001)
log = logger(metrics=["loss","acc", "AUC"], max_i =len(train_loader))
fit(model, train_loader, criterion, optimizer, log, epochs=20)
# -
# ## 4. Evaluating the model
# ### Metrics
# After training we can plot saved metrics using the `logger.plot_metrics()` function.
log.plot_metrics()
# <div class="alert alert-success">
# <h2>Exercise:</h2>
# <p><b>Evaluate</b> the metrics after 15-20 epochs. Should the model train longer?</p>
# </div>
# <div class="alert alert-success">
# <h2>Exercise:</h2>
# <p><b>Try</b> different optimizer functions and compare the different loss curves?</p>
# </div>
# ### Confusion matrix
# <div class="alert alert-success">
# <h2>Exercise:</h2>
# <p><b>Write out </b> some code to obtain the confusion matrix of the predictions on the validation data.</p>
# </div>
# +
# We first load all data in two arrays
y_hat_all = []
y_true_all = []
for i, data in enumerate(testloader, 0):
X_batch, y_batch = data
y_batch = convert_to_binary_class(y_batch)
X_batch, y_batch = Variable(X_batch), Variable(y_batch)
y_hat = model.forward(X_batch)
y_hat_all.append(y_hat)
y_true_all.append(y_batch)
y_hat_all = torch.cat(y_hat_all)
y_true_all = torch.cat(y_true_all)
print(y_hat_all[:10], y_true_all[:10])
# +
### solution
from sklearn.metrics import confusion_matrix
def confusion_matrix_from_variables(y_hat, y_true):
y_true, y_hat = y_true.data.numpy(), y_hat.data.numpy()
y_hat = np.argmax(y_hat, axis=1)
matrix = confusion_matrix(y_true, y_hat)
return matrix
### solution
# -
confusion_matrix_from_variables(y_hat_all, y_true_all)
# ### Sample predictions
def imshow(img):
img = img / 2 + 0.5 # unnormalize
npimg = img.numpy()
fig, ax = plt.subplots(1,1,figsize=(20,20))
plt.imshow(np.transpose(npimg, (1, 2, 0)),)
dataiter = iter(testloader)
images, labels = dataiter.next()
imshow(torchvision.utils.make_grid(images[:8]),)
outputs = model.forward(Variable(images))
predicted = torch.max(outputs.data, 1)[1]
binary_classes[predicted.numpy()[:8]]
# ### Convolutional weights
# +
import torch
from matplotlib import pyplot as plt
def plot_kernels(tensor, num_cols=6):
if not tensor.ndim==4:
raise Exception("assumes a 4D tensor")
num_kernels = tensor.shape[0]
num_rows = 1+ num_kernels // num_cols
fig = plt.figure(figsize=(num_cols,2*num_rows))
for i in range(tensor.shape[0]):
ax1 = fig.add_subplot(2*num_rows,num_cols,i+1+tensor.shape[0])
ax1.imshow(np.mean(tensor[i],0))
ax1.axis('off')
ax1.set_xticklabels([])
ax1.set_yticklabels([])
plt.subplots_adjust(wspace=0.1, hspace=0.1)
plt.show()
filters = model.modules
body_model = [i for i in model.children()]
conv1_weights = body_model[0].weight.data.numpy()
plot_kernels(conv1_weights)
# -
# <hr>
# <div class="alert alert-success">
# <h2>Exercise:</h2>
# <p><b>Optimize</b> the model in any way you can. You can change to the architecture of the model, vary the hyperparameters and add any of the many optimization techniques found in literature. </p>
# </div>
| predmod/lab12/PClab013_CNN_solved.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# # Manually load and inspect Ylm_depict data
# +
# Low-level import
from numpy import array,loadtxt,linspace,zeros,exp,ones,unwrap,angle,pi
# Setup ipython environment
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# Import useful things from kerr
from kerr.formula.ksm2_cw import CW as cwfit
from kerr.formula.ksm2_sc import SC as scfit
from kerr.pttools import leaver_workfunction as lvrwork
from kerr import leaver,rgb
from kerr.models import mmrdns
#
from nrutils import scsearch,gwylm,gwf
# Setup plotting backend
import matplotlib as mpl
mpl.rcParams['lines.linewidth'] = 0.8
mpl.rcParams['font.family'] = 'serif'
mpl.rcParams['font.size'] = 12
mpl.rcParams['axes.labelsize'] = 20
from matplotlib.pyplot import *
# -
T0 = 10
ll,mm = 3,2
q = 2
data_file_string = '/Users/book/GARREG/Spectroscopy/Ylm_Depictions/NonPrecessing/MULTI_DATA_6/T0_%i_nmax2_Mmin97ll_Mmin_r75_qref1.50__p1_17-Mar-2014gnx_/Data_Sets/HRq-series/D9_q%1.1f_a0.0_m160/DEPICTION_INFO::NODD_INPUT_ll%i_mm%i_r75.asc'%(T0,q,ll,mm)
data = loadtxt(data_file_string)
# Collect raw fit data for later convenience
rfdata = {}
for k,row in enumerate(data):
#
ll,mm,q,m1,m2,x1,x2,jf,Mf,qid,rew,imw,rewfit,imwfit,reA,imA,reAmean,imAmean,minA,maxA,T1,dT,match,rmse,reB,imB,reBmean,imBmean,minB,maxB = row
ll,mm = int(ll),int(mm)
A = reA+1j*imA
cw = rew + 1j*imw
try:
l,m,n,p = mmrdns.calc_z(qid)
except:
l,m,n,p,l2,m2,n2,p2 = mmrdns.calc_z(qid)
rfdata[(l,m,n,p)] = {}
rfdata[(l,m,n,p)]['ll'],rfdata[(l,m,n,p)]['mm'],rfdata[(l,m,n,p)]['A'],rfdata[(l,m,n,p)]['cw'] = ll,mm,A,cw
print angle( rfdata[(mm,mm,0,1)]['A'] * rfdata[(ll,mm,0,1)]['A'].conj() )
print angle( mmrdns.Afit(mm,mm,0,mmrdns.q2eta(q)) * mmrdns.Afit(ll,mm,0,mmrdns.q2eta(q)).conj() )
# +
#
def rawfit(t):
y = zeros( t.shape, dtype=complex )
for k,row in enumerate(data):
#
ll,mm,q,m1,m2,x1,x2,jf,Mf,qid,rew,imw,rewfit,imwfit,reA,imA,reAmean,imAmean,minA,maxA,T1,dT,match,rmse,reB,imB,reBmean,imBmean,minB,maxB = row
ll,mm = int(ll),int(mm)
A = reA+1j*imA
cw = rew + 1j*imw
try:
l,m,n,p = mmrdns.calc_z(qid)
except:
l,m,n,p,l2,m2,n2,p2 = mmrdns.calc_z(qid)
# NOTE that the amplitudes are for Psi4 here
if True: # (l,m,n,p) in [ (2,2,0,1) ,(2,2,1,1) ] :
y += A*exp( 1j*cw*(t-T0) )
#
a = gwf( array( [t,y.real,-y.imag] ).T )
#
return a,q
_,q = rawfit( linspace(T0,50) )
#
A = scsearch( keyword='sxs', q=q, nonspinning=True,verbose=True )[0]
#
imrnr = gwylm( A, lm=([ll,mm],[2,2]), verbose=True, dt=0.5 )
nr = imrnr.ringdown(T0=T0)
y,_ = rawfit( nr.lm[(ll,mm)]['psi4'].t )
#
eta = mmrdns.q2eta(q)
h = mmrdns.meval_spherical_mode(ll,mm,eta,kind='psi4',gwfout=True)(nr.ylm[0].t)
h.align(nr.lm[(ll,mm)]['psi4'],method='average-phase',mask=nr.ylm[0].t<60)
y.align(nr.lm[(ll,mm)]['psi4'],method='average-phase',mask=nr.ylm[0].t<60)
nr.lm[(ll,mm)]['psi4'].plot()
y.plot()
h.plot()
fig = figure( figsize=2*array([5,3]) )
gca().set_yscale("log", nonposy='clip')
plot( nr.ylm[0].t, nr.lm[(ll,mm)]['psi4'].amp, color=0.5*ones((3,)), label=None )
plot( nr.ylm[0].t, y.amp, '--k', label=None )
plot( nr.ylm[0].t, h.amp, 'k', alpha=0.2, linewidth=6, label=None )
plot( nr.ylm[0].t, nr.lm[(ll,mm)]['psi4'].plus, color=0.5*ones((3,)), label='NR' )
plot( nr.ylm[0].t, y.plus, '--k', label='RAW-FIT' )
plot( nr.ylm[0].t, h.plus, 'k', alpha=0.2, linewidth=6,label='MMRDNS' )
# plot( nr.ylm[0].t, nr.lm[(ll,mm)]['psi4'].cross, color=0.5*ones((3,)), label='NR', alpha=0.8 )
# plot( nr.ylm[0].t, y.cross, '--k', label='RAW-FIT', alpha=0.8 )
# plot( nr.ylm[0].t, h.cross, 'k', alpha=0.1, linewidth=6, label='MMRDNS' )
ylim( [max(nr.lm[(ll,mm)]['psi4'].amp)*1e-5,1.2*max(nr.lm[(ll,mm)]['psi4'].amp)] )
xlim( [T0,150] )
xlabel(r'$(t-{t^{\mathrm{Peak}}}_{\mathrm{Lum.}})/M$')
ylabel(r'${rM}\psi_{%i%i}$'%(ll,mm))
legend(frameon=False)
title( nr.label )
savefig('mmrdns_psi4_comparison_%s_ll%imm%i.pdf'%(nr.label.replace('-','_'),ll,mm))
# gca().set_yscale("log", nonposy='clip')
# -
# ### NOTES
#
#
nr.simdir
| notes/ns/notebooks/manual_psi4_data_comparison.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Modelling DNN
# # Module loading
# +
import datetime, time, os
import numpy as np
import pandas as pd
import tensorflow as tf
import seaborn as sns
import matplotlib.pyplot as plt
plt.style.use('seaborn')
from tensorflow.keras import layers
from tensorflow.keras import regularizers
from tensorflow.keras.metrics import RootMeanSquaredError
print('Using TensorFlow version: %s' % tf.__version__)
RSEED = 42
# +
# #!pip install -q git+https://github.com/tensorflow/docs
import tensorflow_docs as tfdocs
import tensorflow_docs.modeling
import tensorflow_docs.plots
# -
# ## Data loading
# +
# Load data
df = pd.read_csv("data/flight_data.csv")
X = df.copy()
y = X.pop("target")
df.head()
df_sub = pd.read_csv("data/flight_submission.csv", index_col=0)
submission = pd.read_csv("data/ID.csv", index_col=0)
X_sub = df_sub
# -
# Data Preprocessing
# +
from sklearn.pipeline import Pipeline
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import StandardScaler, MinMaxScaler, OneHotEncoder, OrdinalEncoder
categories_country_ARR = [np.asarray(df.groupby("country_ARR").mean().sort_values(by="target").index)]
categories_country_DEP = [np.asarray(df.groupby("country_DEP").mean().sort_values(by="target").index)]
categories_DEPSTN = [np.asarray(df.groupby("DEPSTN").mean().sort_values(by="target").index)]
categories_ARRSTN = [np.asarray(df.groupby("ARRSTN").mean().sort_values(by="target").index)]
categories_FLTID = [np.asarray(df.groupby("FLTID").mean().sort_values(by="target").index)]
categories_AC = [np.asarray(df.groupby("AC").mean().sort_values(by="target").index)]
# Preprocessor pipelines
num_cols = ['distance', 'domestic', 'dep_hour', 'dep_weekday',
'duration_min', "dep_day", "arr_hour"]
cat_cols = ["STATUS", "operator"]
# Preprocessor for numerical features
num_pipeline = Pipeline([
#('num_scaler', StandardScaler()),
('num_scaler', MinMaxScaler())
])
# Preprocessor for categorical features
cat_pipeline = Pipeline([
('cat_encoder', OneHotEncoder(handle_unknown='ignore'))
])
# Put together preprocessor pipeline
preprocessor = ColumnTransformer([
('num', num_pipeline, num_cols),
('cat', cat_pipeline, cat_cols),
('cat_AC', OrdinalEncoder(categories=categories_AC, handle_unknown="use_encoded_value",
unknown_value=(len(categories_AC[0])+1)), ["AC"]),
('cat_FLTID', OrdinalEncoder(categories=categories_FLTID, handle_unknown="use_encoded_value",
unknown_value=(len(categories_FLTID[0])+1)), ["FLTID"]),
('cat_ARRSTN', OrdinalEncoder(categories=categories_ARRSTN, handle_unknown="use_encoded_value",
unknown_value=(len(categories_ARRSTN[0])+1)), ["ARRSTN"]),
('cat_DEPSTN', OrdinalEncoder(categories=categories_DEPSTN, handle_unknown="use_encoded_value",
unknown_value=(len(categories_DEPSTN[0])+1)), ["DEPSTN"]),
('cat_country_ARR', OrdinalEncoder(categories=categories_country_ARR, handle_unknown="use_encoded_value",
unknown_value=(len(categories_country_ARR[0])+1)), ["country_ARR"]),
('cat_country_DEP', OrdinalEncoder(categories=categories_country_DEP, handle_unknown="use_encoded_value",
unknown_value=(len(categories_country_DEP[0])+1)), ["country_DEP"])
])
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=RSEED, test_size=0.2)
X_sub = df_sub
X_train = preprocessor.fit_transform(X_train)
X_test = preprocessor.transform(X_test)
X_sub = preprocessor.transform(X_sub)
print(X_train.shape, X_test.shape)
# -
# ## Training
# ### Build, compile and fit the model
# Clear recent logs
# !rm -rf my_logs/
# Define path for new directory
root_logdir = os.path.join(os.curdir, "my_logs")
# Define function for creating a new folder for each run
def get_run_logdir():
run_id = time.strftime('run_%d_%m_%Y-%H_%M_%S')
return os.path.join(root_logdir, run_id)
# Store directory path for current run
run_logdir = get_run_logdir()
# Define function for creating callbacks
def get_callbacks(name):
return [
tfdocs.modeling.EpochDots(),
#tf.keras.callbacks.EarlyStopping(monitor='val_binary_crossentropy', patience=200),
tf.keras.callbacks.TensorBoard(run_logdir+name, histogram_freq=1),
]
def model_compile_and_fit(model, name, optimizer=None, max_epochs=30):
BATCH_SIZE = 500
LEARNING_RATE = 0.001 # Get optimizer
if optimizer == None:
lr_schedule = tf.keras.optimizers.schedules.InverseTimeDecay(
LEARNING_RATE,
decay_steps=len(X_train) // BATCH_SIZE,
decay_rate=1,
staircase=False)
optimizer = tf.keras.optimizers.Adam(learning_rate=LEARNING_RATE , name='Adam')
# model.compile
with tf.device('/cpu:0'):
model.compile(optimizer=optimizer,
loss='mse',
metrics=[RootMeanSquaredError()])
print(model.summary())
# model.fit
history[name] = model.fit(X_train,
y_train,
validation_split=0.2,
verbose=1,
#steps_per_epoch=len(X_train) // BATCH_SIZE,
batch_size=BATCH_SIZE,
epochs=max_epochs,
callbacks=get_callbacks(name),
)
return history
# ### Train the model
# +
DROPOUT_RATE = 0.0
REGULARIZATION = 0.01
with tf.device('/cpu:0'):
model = tf.keras.Sequential([
tf.keras.layers.Dense(37,kernel_initializer = 'uniform', activation='relu',input_dim = X_train.shape[1], kernel_regularizer=regularizers.l2(REGULARIZATION)),
tf.keras.layers.Dense(32,kernel_initializer = 'uniform', activation='relu', kernel_regularizer=regularizers.l2(REGULARIZATION)),
tf.keras.layers.Dropout(DROPOUT_RATE),
tf.keras.layers.Dense(32,kernel_initializer = 'uniform', activation='relu', kernel_regularizer=regularizers.l2(REGULARIZATION)),
tf.keras.layers.Dropout(DROPOUT_RATE),
tf.keras.layers.Dense(1,kernel_initializer = 'uniform')
])
# compile and fit model
history = {}
history = model_compile_and_fit(model, 'next_try', max_epochs=50)
# plot history
history_plotter = tfdocs.plots.HistoryPlotter(metric = 'root_mean_squared_error', smoothing_std=10)
history_plotter.plot(history)
# +
#import seaborn as sns
y_pred = model.predict(X_test).flatten()
residuals = y_pred - np.asarray(y_test)
sns.scatterplot(y_pred, residuals, alpha=0.2)
plt.xlabel("Predicted Delay (min)")
plt.ylabel("Delay (min)")
#plt.xlim(-5,250)
plt.ylim(-700,300)
# -
loss, rmse = model.evaluate(X_test, y_test)
print(rmse)
# +
sub_target = model.predict(X_sub)
submission["target"] = sub_target
submission.to_csv("data/submission.csv", index=False)
max(sub_target)
| ModellingDNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <p style="align: center;"><img src="https://static.tildacdn.com/tild6636-3531-4239-b465-376364646465/Deep_Learning_School.png", width=300, height=300></p>
#
# <h3 style="text-align: center;"><b>Физтех-Школа Прикладной математики и информатики (ФПМИ) МФТИ</b></h3>
# <h2 style="text-align: center;"><b>Библиотека <a href="http://pandas.pydata.org/">pandas</a></b></h2>
# Библиотека `pandas` активно используется в современном data science для работы с данными, которые могут быть представлены в виде таблиц (а это очень, очень большая часть данных)
# `pandas` есть в пакете Anaconda, но если вдруг у Вас её по каким-то причинам нет, то можно установить, раскомментировав следующую команду:
# +
# #!pip install pandas
# -
import pandas as pd
# Нужны данные. Про AppStore слышали? :)
# ### Скачать данные: https://www.kaggle.com/ramamet4/app-store-apple-data-set-10k-apps
# Нас интересует файл `AppStore.csv`.
# Кстати, `.csv` (Comma Separated Values) - наверное, самый частый формат данных в современном data science. По сути, это то же самое, что и `.xls` (и `.xlsx`), то есть таблица.
# Итак, *чтение файла с данными*:
data = pd.read_csv('./app-store-apple-data-set-10k-apps/AppleStore.csv')
# Посмотрим, что же такое переменная `data`:
data
type(data)
# Тип == `pandas.core.frame.DataFrame`, обычно говорят просто "датафрейм", то есть "кусок данных".
# Можно вывести информацию по датафрейму:
data.info()
# И более статистически значимую информацию:
data.describe()
# Итак, наши данные -- это какая-то **информация про приложения в AppStore**. Давайте посмотрим, что тут есть:
# все столбцы датафрейма (названия)
data.columns
# ---
# #### Задание 0
# О чём данные? (судя по столбцам)
# ---
# Все значения датафрейма:
# одна строка == описание одного объекта, в данном случае -- приложения из AppStore
data.values
data.values[0]
data.shape
# -- то есть 7197 строк и 17 столбцов.
# Важно уметь обращаться к определённой строке или столбцу dataframe'а. Индексирование очень похоже на numpy, однако есть свои тонкости:
# * Получить весь столбец (в даном случае столбец 'track_name'):
data['track_name']
type(data['track_name'])
data['track_name'][0]
# `pandas.core.series.Series` -- это тип подвыборки датафрейма. С этим типом работают реже, но стоит знать, что он есть.
# * Получить определённую строку:
data.iloc[10]
# Более продвинутое и полезное индексирование:
data.iloc[5, 10]
# -- `data.iloc[i, j]`, где `i` -- номер строки, `j` -- номер столбца
# Почти все типы индексирования:
# * к строкам по числовому индексу, к столбцам -- по имени
data.loc[[0, 1, 2], ['track_name', 'id']]
# * к строкам и к столбцам по числовому индексу
data.iloc[[1, 3, 5], [2, 1]]
# * поддерживает и то, и то, но **запрещён**:
data.ix[[2,1,0], ['track_name', 'id']]
data.ix[[0,1,2], [2, 1]]
# Точно также дело обстоит со **срезами**:
data.iloc[5:100, 0:5]
# Но обычно нужно уметь отвечать на более осмысленные вопросы, например:
# * сколько приложений имеют рейтинг юзеров >= 4?
len(data[data['user_rating'] >= 4])
# * какие есть валюты в столбце currency?
np.unique(data['currency'])
# * сколько приложений жанра Games? А тех, которые Games и рейтинг >= 4?
# 1
len(data[data['prime_genre'] == 'Games'])
# 2
len(data[(data['user_rating'] >= 4) & (data['prime_genre'] == 'Games')])
# Оператор "ИЛИ" аналогично &, только он через |
len(data[(data['user_rating'] >= 4) | (data['prime_genre'] == 'Games')])
# ---
# #### Задание 1
# Получите ответ на следующие вопросы (с помощью pandas):
#
# 1). Сколько всего жанров и какие они?
# 2). Сколько суммарно байт весят все приложения жанра Finance?
# 3). Сколько всего бесплатных приложений, у которых рейтинг юзеров больше трёх?
# 4). Есть ли приложения, которые стоят больше 100 долларов?
# +
# Ваш код здесь
# -
# ---
# Часто надо сделать что-то такое:
import numpy as np
np.array(data['size_bytes'])
# То есть взять столбец из таблицы и засунуть его в np.array(). Это можно сделать и просто обратившись к .values, ведь его тип и есть np.array:
type(data['size_bytes'].values)
# Больше про pandas можно найти по этом полезным ссылкам:
# * Официальные туториалы: http://pandas.pydata.org/pandas-docs/stable/tutorials.html
# * Статья на Хабре от OpenDataScience сообщества: https://habr.com/company/ods/blog/322626/
# * Подробный гайд: https://media.readthedocs.org/pdf/pandasguide/latest/pandasguide.pdf
# Главное в работе с новыми библиотеками -- не бояться тыкать в разные функции, смотреть типы возвращаемых объектов и активно пользоваться Яндексом, а ещё лучше понимать всё из docstring'а (`Shift+Tab` при нахождении курсора внутри скобок функции).
| dlschool/week3/pandas/[seminar]pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # First thing is to write the data in a dataframe
# +
import pandas as pd
df_red = pd.read_csv('winequality-red.csv', sep = ';') # Telling python that data is separated by ';' (default is ',')
df_white = pd.read_csv('winequality-white.csv', sep = ';')
# Create a new column with type of wine (red or white)
df_red['wine_type'] = 'red'
df_white['wine_type'] = 'white'
# Merge the two dataframes
df_wine = pd.concat([df_red, df_white])
# -
df_wine.info()
# What are the names of columns?
df_wine.columns
df_wine
# # Description (simple statistics on each column)
# Select some columns with quantitative data to run some descriptive statistics
wine_quant = ['fixed acidity', 'citric acid', 'residual sugar',
'chlorides', 'free sulfur dioxide', 'total sulfur dioxide', 'density',
'pH', 'sulphates', 'alcohol']
round(df_red[wine_quant].describe(),2)
reds = round(df_red[wine_quant].describe(),2)
whites = round(df_white[wine_quant].describe(),2)
pd.concat([reds,whites], axis=1, keys=['Red stats', 'White stats'])
# Let's try again some descriptive data, including wine quality
wine_quant = ['residual sugar', 'total sulfur dioxide', 'sulphates', 'alcohol', 'quality']
reds = round(df_red[wine_quant].describe(),2)
whites = round(df_white[wine_quant].describe(),2)
pd.concat([reds,whites], axis=1, keys=['Red stats', 'White stats'])
# # Univariate Analysis: Just look at each column in better details using histograms
df_wine.hist()
df_wine.hist(bins = 20, color = 'red', grid = False)
df_wine.hist(bins = 20, color = 'red', grid = False)
# +
import matplotlib.pyplot as plt
df_wine.hist(bins = 20, color = 'red', grid = False)
plt.tight_layout(rect=(0, 0, 2, 3))
# -
df_wine.hist(bins = 20, color = 'red', grid = False)
plt.tight_layout(rect=(0, 0, 1.2, 1.2))
wt = plt.suptitle('Red Wine Univariate Plots', x=0.65, y=1.25, fontsize=14)
# +
import matplotlib.pyplot as plt
df_wine.hist(bins=15, color='red', edgecolor='black', linewidth=1.0, xlabelsize=8, ylabelsize=8)
plt.tight_layout(rect=(0, 0, 1.2, 1.2))
wt = plt.suptitle('Red Wine Univariate Plots', color = 'blue', x=0.65, y=1.25, fontsize=14)
df_white.hist(bins=15, color='yellow', edgecolor='black', linewidth=1.0, xlabelsize=8, ylabelsize=8)
plt.tight_layout(rect=(0, 0, 1.2, 1.2))
wt = plt.suptitle('White Wine Univariate Plots', color = 'blue', x=0.65, y=1.25, fontsize=14)
df_wine.hist(bins=15, color='green', edgecolor='black', linewidth=1.0, xlabelsize=8, ylabelsize=8)
plt.tight_layout(rect=(0, 0, 1.2, 1.2))
wt = plt.suptitle('All Wine Univariate Plots', color = 'blue', x=0.65, y=1.25, fontsize=14)
# -
# # Multivariate Analysis: Analysis of the relationship between different variables.
round(df_wine.corr(),2)
print(round(df_wine.corr(),2))
x = df_wine.iloc[:,1].tolist()
y = df_wine.iloc[:,2].tolist()
fig, axs = plt.subplots(1,2)
axs[0].hist(x)
axs[1].hist(y)
# +
# Apparently the library seaborn helps producing nice graphs!!!
import seaborn as sns
x = df_wine.iloc[:,1].tolist()
sns.distplot(x)
# -
# # Step up: correlations
df_wine.corr()
f, ax = plt.subplots(figsize=(10, 5))
corr = df_wine.corr()
sns.heatmap(round(corr,2), annot=True, ax=ax,cmap = "coolwarm", fmt='.2f', linewidths=.05)
t = f.suptitle('Wine Attributes Correlation Heatmap', fontsize=12)
wine_quant = ['sulphates', 'alcohol', 'quality', 'wine_type']
sns.pairplot(df_wine[wine_quant], hue='wine_type', height = 2.4, palette={"red": "#FF9999", "white": "#FFE888"}, plot_kws=dict(edgecolor="black", linewidth=0.5))
| NotesAndTutorials/WineQuality.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from sklearn.datasets import make_blobs
import pandas as pd
from bokeh.plotting import figure, show, output_file, save, reset_output
from bokeh.resources import CDN
from bokeh.embed import file_html
from bokeh.io import output_notebook
from bokeh.models import HoverTool
# +
##################
# Scatter Plot #
##################
N = 500
data, labels = make_blobs(n_samples=N, n_features=3, cluster_std=1.0, centers=2)
data = pd.DataFrame(data)
x = data[0]
y = data[1]
radii = np.random.random(size=N) * 0.2
x_color = np.random.random(size=N) * 100
y_color = np.random.random(size=N) * 100
colors = [
"#%02x%02x%02x" % (int(r), int(g), 150) for r, g in zip(50+2*x_color, 30+2*y_color)
]
p = figure(tools="pan", sizing_mode="stretch_both", toolbar_location=None)
p.grid.visible = False
p.scatter(x, y, radius=radii,
fill_color=colors, fill_alpha=0.6,
line_color=None)
p.add_tools(HoverTool(
tooltips=[("User", "$index"),
("Flow Index", "@radius"),
("X, Y", "(@x, @y)")],
mode="mouse",
point_policy="follow_mouse"
))
#output_notebook()
#show(p)
output_file("scatter_plot.html", mode="inline")
save(p)
# +
############
# Hexbin #
############
data, labels = make_blobs(n_samples=500, n_features=2, cluster_std=1.0)
data = pd.DataFrame(data)
x = data[0]
y = data[1]
p = figure(match_aspect=True, toolbar_location=None,
tools="wheel_zoom,reset", background_fill_color='#440154', sizing_mode="stretch_both")
p.grid.visible = False
r, bins = p.hexbin(x, y, size=0.5, hover_color="pink", hover_alpha=0.8)
p.circle(x, y, color="white", size=1)
p.add_tools(HoverTool(
tooltips=[("count", "@c"), ("(q,r)", "(@q, @r)")],
mode="mouse", point_policy="follow_mouse", renderers=[r]
))
#output_notebook()
#show(p)
output_file("hexbin_plot_neu.html", mode="inline")
save(p)
| Visualizations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# 107062566 黃鈺程 CVLab, Delta 722
# Dependencies: numpy, pandas, matplotlib, seaborn, sklearn=0.20
# +
# %matplotlib inline
# %config InlineBackend.figure_format = 'svg'
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import OneHotEncoder
from sklearn.impute import SimpleImputer
from sklearn.pipeline import Pipeline
from sklearn.compose import ColumnTransformer
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import SVC
# -
df = pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/'
'mushroom/agaricus-lepiota.data', header=None, engine='python')
column_name = ['classes','cap-shape', 'cap-surface','cap-color','bruises?','odor',
'gill-attachment','gill-spacing','gill-size','gill-color',
'stalk-shape','stalk-root','stalk-surface-above-ring',
'stalk-surface-below-ring','stalk-color-above-ring',
'stalk-color-below-ring','veil-type','veil-color','ring-number',
'ring-type','spore-print-color','population','habitat']
df.columns = column_name
X = df.drop(columns=['classes'], axis=1)
y = df['classes']
xt, xv, yt, yv = train_test_split(X, y, test_size=0.2, random_state=0)
# +
transformer = Pipeline(steps=[
('imputer', SimpleImputer(missing_values='?', strategy='most_frequent')),
('onehot', OneHotEncoder(handle_unknown='ignore'))])
preprocessor = ColumnTransformer(
transformers=[('cat', transformer, df.columns[1:])])
# -
# KNN
clf_knn = Pipeline(steps=[
('preprocess', preprocessor),
('classifier', KNeighborsClassifier())
]).fit(xt, yt)
acc = clf_knn.score(xv, yv)
print(acc)
# SVC
clf_svc = Pipeline(steps=[
('preprocess', preprocessor),
('classifier', SVC(gamma='scale'))
]).fit(xt, yt)
acc = clf_svc.score(xv, yv)
print(acc)
# # Report
#
# ※ 需要 sklearn 0.20 以上來跑我的程式
#
# 好險,差點做成了 train, test 資料一起 preprocessing,這樣會造成不嚴謹的程式。
# 所幸有發現,然後改成了稅用 Pipeline 的方式實作,處理 NA 真是麻煩~
#
# 另外提供助教參考,並不需要使用 LabelEncoder 將資料轉成數值資料。
# 使用 sklearn 0.20 的 SimpleImputer 即可簡單的做預處理~
# [官方範例](http://scikit-learn.org/stable/auto_examples/compose/plot_column_transformer_mixed_types.html#sphx-glr-auto-examples-compose-plot-column-transformer-mixed-types-py)
#
# 結果很喜人:
# 1. KNN: 1.0
# 2. SVC: 1.0
| lab07.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import torch
from spotlight.factorization.explicit import ExplicitFactorizationModel
from spotlight.interactions import Interactions
from spotlight.cross_validation import random_train_test_split
from spotlight.datasets.movielens import get_movielens_dataset
from spotlight.evaluation import rmse_score
dataset = get_movielens_dataset(variant='100K')
# -
model = ExplicitFactorizationModel(loss='regression',
embedding_dim=20, # latent dimensionality
n_iter=10, # number of epochs of training
batch_size=1024, # minibatch size
l2=1e-9, # strength of L2 regularization
learning_rate=1e-3,
use_cuda=torch.cuda.is_available())
# +
train, test = random_train_test_split(dataset, random_state=np.random.RandomState(42))
print('Split into \n {} and \n {}.'.format(train, test))
# -
type(train.user_ids[0])
type(train.item_ids[0])
type(train.ratings[0])
model.fit(train, verbose=True)
# +
train_rmse = rmse_score(model, train)
test_rmse = rmse_score(model, test)
print('Train RMSE {:.3f}, test RMSE {:.3f}'.format(train_rmse, test_rmse))
# -
def get_datasets():
"""
Returns
-------
Interactions: :class:`spotlight.interactions.Interactions`
instance of the interactions class
"""
def get_part(num, part):
print("Reading {} part {}".format(part[1:], num))
URL_PREFIX = 'ml-10M100K/r' + str(num)
data = np.genfromtxt(URL_PREFIX + part, delimiter='::', dtype=(int, int, float))
users, items, ratings = np.zeros(len(data), dtype=int), \
np.zeros(len(data), dtype=int), \
np.zeros(len(data), dtype=np.float32)
for i, inst in enumerate(data):
users[i], items[i], ratings[i] = inst[0], inst[1], inst[2]
return users, items, ratings
train_extension = '.train'
test_extension = '.test'
trains = [Interactions(*get_part(r, train_extension)) for r in range(1, 6)]
tests = [Interactions(*get_part(r, test_extension)) for r in range(1, 6)]
return trains, tests
train_datasets, test_datasets = get_datasets()
lams = [0.1, 0.01, 0.001]
for lam in lams:
print("L2 Regularization value - {}\n".format(lam))
model = ExplicitFactorizationModel(loss='regression',
embedding_dim=20, # latent dimensionality
n_iter=10, # number of epochs of training
batch_size=1024, # minibatch size
l2=lam, # strength of L2 regularization
learning_rate=1e-3,
use_cuda=torch.cuda.is_available())
errors = []
for train, test in zip(train_datasets, test_datasets):
model.fit(train, verbose=True)
errors.append(rmse_score(model, test))
print("Cross-validation RMSE is {}".format(np.mean(errors)))
| Recommendation_Engines/Spotlight.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 循环
# - 循环是一种控制语句块重复执行的结构
# - while 适用于广度遍历
# - for 开发中经常使用
# +
#堆 :先进后出
#队列:先进先出
#广度遍历:用于抓取信息 要用正则匹配
# -
# ## while 循环
# - 当一个条件保持真的时候while循环重复执行语句
# - while 循环一定要有结束条件,否则很容易进入死循环
# - while 循环的语法是:
#
# while loop-contunuation-conndition:
#
# Statement
import os
while 1:
os.system('say 天气真好!') #死循环
#先定义结束值
sum_ = 0
i = 1
while i <10:
sum_ = sum_ + i
i = i + 1
print(sum_)
# ## 示例:
# sum = 0
#
# i = 1
#
# while i <10:
#
# sum = sum + i
# i = i + 1
# ## 错误示例:
# sum = 0
#
# i = 1
#
# while i <10:
#
# sum = sum + i
#
# i = i + 1
# - 一旦进入死循环可按 Ctrl + c 停止
# ## EP:
# 
# 
#死循环 i=1 % 2 等于1 if条件不成立
i = 1
while 1 < 10:
if i % 2 == 0:
print(i)
i = 1
while i< 10:
if i % 2 == 0:
print(i)
i += 1
i = 0
while i< 10:
if i % 2 == 0:
print(i)
i += 1
#错误在于没有结束条件
count = 0
while count<100:
print(count)
#count -= 1 永远小于100 一直循环
count = 0
while count<100:
print(count)
count -= 1
#没有缩进
# # 验证码
# - 随机产生四个字母的验证码,如果正确,输出验证码正确。如果错误,产生新的验证码,用户重新输入。
# - 验证码只能输入三次,如果三次都错,返回“别爬了,我们小网站没什么好爬的”
# - 密码登录,如果三次错误,账号被锁定
#
#利用ASCII码,
import random
i = 0
a = ['a','b','c','d','e','f','g','h','i','j','k','l','m','n','o','p','q','r','s','t','u','v','w','x','y','z']
while i<3:
num1 = chr(random.randint(97,122))
num2 = chr(random.randint(97,122))
num3 = chr(random.randint(97,122))
num4 = chr(random.randint(97,122))
zong = num1+num2+num3+num4
print('验证码是:'+zong)
zong = input('输入验证码:')
if zong == num1+num2+num3+num4:
print('验证码正确',zong)
break
else:
print('验证码错误,请重新输入',zong)
i +=1
else:
print('验证超过次数')
# ## 尝试死循环
# ## 实例研究:猜数字
# - 你将要编写一个能够随机生成一个0到10之间的且包括两者的数字程序,这个程序
# - 提示用户连续地输入数字直到正确,且提示用户输入的数字是过高还是过低
# ## 使用哨兵值来控制循环
# - 哨兵值来表明输入的结束
# - 
# ## 警告
# 
# ## for 循环
# - Python的for 循环通过一个序列中的每个值来进行迭代
# - range(a,b,k), a,b,k 必须为整数
# - a: start
# - b: end
# - k: step
# - 注意for 是循环一切可迭代对象,而不是只能使用range
for iterm in range(0,10,2):
print(iterm)
for i in (1,2,3):
print(i)
dir('abc') #查看对象是否迭代
#用while循环实现遍历 abcdefg
i = 0
str_ = ['a','b','c']
while i< 3:
print(str_[i])
i += 1
# # 在Python里面一切皆对象
# ## EP:
# - 
#5.8
i = 0
sum_ = 0
while i < 1001:
sum_ = sum_ + i
i+=1
print(sum_)
#5.9
sum_ = 0
for i in range(1,10000):
sum_ += i
if sum_ > 10000:
break
print(sum_)
sum_ = 0
for count in range(5):
number = eval(input('>>'))
sum_ += number
print('sum_ is',sum_)
print('count is',count)
# ## 嵌套循环
# - 一个循环可以嵌套另一个循环
# - 每次循环外层时,内层循环都会被刷新重新完成循环
# - 也就是说,大循环执行一次,小循环会全部执行一次
# - 注意:
# > - 多层循环非常耗时
# - 最多使用3层循环
#大循环执行一次,小循环全部执行完,大循环才会进入下一次执行
for i in range(10):
for j in range(5):
print(i,j)
# ## EP:
# - 使用多层循环完成9X9乘法表
# - 显示50以内所有的素数
for i in range(1,10):
for j in range(1,i+1): #才能取到i
print('{}x{}={}'.format(j,i,i*j),end=' ')
print() #强制换行
for i in range(2,51):
for j in range(2,i):
if i % j == 0:
break
else:
print(i)
# ## 关键字 break 和 continue
# - break 跳出循环,终止循环
# - continue 跳出此次循环,继续执行
# ## 注意
# 
# 
# # Homework
# - 1
# 
i = 0
j = 0
sum_ = 0
num = eval(input('输入一个整数:'))
while num != 0:
if num > 0:
i += 1
sum_ = sum_ + i
else:
j += 1
sum_ = sum_ - j
num = eval(input('输入一个整数:'))
print('正数:',i)
print('负数:',j)
print('和:',sum_)
print('平均值:',sum_ / (i+j))
# - 2
# 
# - 3
# 
i = 0
j = 0
sum_ = 0
num = eval(input('输入一个整数:'))
while num != 0:
if num > 0:
i += 1
sum_ = sum_ + i
else:
j += 1
sum_ = sum_ - j
num = eval(input('输入一个整数:'))
print('正数:',i)
print('负数:',j)
print('和:',sum_)
print('平均值:',sum_ / (i+j))
# - 4
# 
count = 0
for i in range(100,1000):
if (i % 5 == 0) and (i % 6 == 0):
print(i,end=' ')
count += 1
if (count % 10 ==0):
print(end='\n')
# - 5
# 
n = 0
max =0
while n**2 <= 12000:
if n**3 <= 12000:
if n>=max:
max=n
n = n + 1
print(n,max)
# - 6
# 
# - 7
# 
sum1 = 0
sum2 = 0
for i in range(1,50001):
sum2 += 1 / i
print('从左到右的结果为:',sum2)
j = 50000
while j != 0:
sum1 += 1/j
j -=1
print('从右到左的结果为:',sum1)
# - 8
# 
sum_ = 0
i = 1
for i in range(1,98):
if i % 2 ==1:
j = i / (i+2)
sum_ = sum_ + j
print('数列和为:',sum_)
# - 9
# 
num = eval(input('输入一个数:'))
pi = 0
for i in range(1,num+1): #为了能取到num
pi = pi+ 4 * (((-1)**(i+1))/(2*i-1))
print('近似π的值为:',pi)
# - 10
# 
# 2**n -1 先判断质数
#(2的n次方-1)*(2的n-1次方i = 0
for i in range(1,10000):
n = 0
for j in range(1,i):
if i % j == 0:
n += j
if n == i:
print(i,'是完全数')
# - 11
# 
count = 0
for i in range(1,8):
for j in range(1,i):
if i == j:
break
else:
print(i,j,end=',')
#print('{}'.format(i,j),end=',')
count += 1
print()
print('可能的组合数为:',count)
# - 12
# 
| 9.13.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## MA
# +
import menpo.io as mio
import numpy as np
import tensorflow as tf
from sklearn import svm
import matplotlib.pyplot as plt
from tensorflow.examples.tutorials.mnist import input_data
# %matplotlib inline
# -
# ### data preparation
stock_slices = mio.import_pickle('/homes/yz4009/wd/gitdev/MarketAnalysor/DataAnalysis/data/090_030_slices.pkl')
len(stock_slices)
data = []
for s in stock_slices:
for k in s.keys():
data += s[k]
# +
def label_y(trend):
cl = [0, 0, 1]
if trend > 0.3:
cl = [1, 0, 0]
elif trend < -0.3:
cl = [0, 1, 0]
return cl
data_X = [d['data'] for d in data]
data_Y = [label_y(d['trend']) for d in data]
# +
data_X = np.array(data_X)
data_Y = np.array(data_Y)
data_X[np.isnan(data_X)] = 0
# -
n_data = len(data)
n_train = int(n_data * 0.8)
n_valid = int(n_data * 0.1)
train_X = data_X[:n_train]
train_Y = data_Y[:n_train]
valid_X = data_X[n_train:n_train+n_valid]
valid_Y = data_Y[n_train:n_train+n_valid]
test_X = data_X[n_train+n_valid:]
test_Y = data_Y[n_train+n_valid:]
# ### SVM Prediction
# + active=""
# acc2 = []
# for C in range(90000, 900000, 10000):
# print C
# clf = svm.SVC(C=C, tol=0.0001)
# clf.fit(train_X, train_Y)
#
# predict_Y = clf.predict(valid_X)
#
# acc2.append(float(np.sum(predict_Y == valid_Y) ) / len(valid_Y))
# + active=""
# plt.plot(acc2)
# -
# ### Tensor Flow Prediction
train_X.shape
np.isnan(train_Y).any()
train_Y.shape
# +
batch_size = 100
sess = tf.InteractiveSession()
x = tf.placeholder(tf.float32, [None, 89])
y_ = tf.placeholder(tf.float32, [None, 3])
W1 = tf.Variable(tf.random_uniform([89, 3], -1.0, 1.0))
b1 = tf.Variable(tf.random_uniform([3], -1.0, 1.0))
pred = tf.nn.softmax(tf.matmul(x, W1) + b1)
keep_prob = tf.placeholder(tf.float32)
y = tf.nn.dropout(pred, keep_prob)
cross_entropy = -tf.reduce_sum(y_*tf.log(tf.clip_by_value(y,1e-10,1.0)))
train_step = tf.train.GradientDescentOptimizer(0.01).minimize(cross_entropy)
correct_prediction = tf.equal(tf.argmax(y,1), tf.argmax(y_,1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
sess.run(tf.initialize_all_variables())
for i in xrange(train_X.shape[0] / batch_size):
batch_xs = train_X[i * batch_size: i * batch_size + batch_size]
batch_ys = train_Y[i * batch_size: i * batch_size + batch_size]
if i % batch_size == 0:
train_accuracy = accuracy.eval(feed_dict={
x:batch_xs, y_: batch_ys, keep_prob: 1.0})
print("step %d, training accuracy %g"%(i, train_accuracy))
train_step.run(feed_dict={x: batch_xs, y_: batch_ys, keep_prob: 0.5})
print(sess.run(accuracy, feed_dict={x: test_X, y_: test_Y, keep_prob: 1.0}))
# -
ret = y.eval(feed_dict={x: test_X})
np.sum((ret[:,0] - ret[:,1]) < 0)
# #### demo
# +
import tensorflow as tf
import numpy as np
# Create 100 phony x, y data points in NumPy, y = x * 0.1 + 0.3
x_data = np.random.rand(100).astype(np.float32)
y_data = x_data * 0.1 + 0.3
# Try to find values for W and b that compute y_data = W * x_data + b
# (We know that W should be 0.1 and b 0.3, but Tensorflow will
# figure that out for us.)
W = tf.Variable(tf.zeros([1]))
b = tf.Variable(tf.zeros([1]))
y = W * x_data + b
# Minimize the mean squared errors.
loss = tf.reduce_mean(tf.square(y - y_data))
optimizer = tf.train.GradientDescentOptimizer(0.5)
train = optimizer.minimize(loss)
# Before starting, initialize the variables. We will 'run' this first.
init = tf.initialize_all_variables()
# Launch the graph.
sess = tf.Session()
sess.run(init)
# Fit the line.
for step in xrange(401):
sess.run(train)
if step % 20 == 0:
print(step, sess.run(W), sess.run(b))
# Learns best fit is W: [0.1], b: [0.3]
# -
# #### mnist
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("MNIST_data/", one_hot=True)
# +
sess = tf.InteractiveSession()
x = tf.placeholder(tf.float32, [None, 784])
W = tf.Variable(tf.zeros([784, 10]))
b = tf.Variable(tf.zeros([10]))
y = tf.nn.softmax(tf.matmul(x, W) + b)
y_ = tf.placeholder(tf.float32, [None, 10])
cross_entropy = -tf.reduce_sum(y_*tf.log(y))
train_step = tf.train.GradientDescentOptimizer(0.01).minimize(cross_entropy)
init = tf.initialize_all_variables()
sess.run(init)
for i in range(1000):
batch_xs, batch_ys = mnist.train.next_batch(100)
train_step.run(feed_dict={x: batch_xs, y_: batch_ys})
correct_prediction = tf.equal(tf.argmax(y,1), tf.argmax(y_,1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
print(sess.run(accuracy, feed_dict={x: mnist.test.images, y_: mnist.test.labels}))
# -
# #### deep mnist
# +
def weight_variable(shape):
initial = tf.truncated_normal(shape, stddev=0.1)
return tf.Variable(initial)
def bias_variable(shape):
initial = tf.constant(0.1, shape=shape)
return tf.Variable(initial)
def conv2d(x, W):
return tf.nn.conv2d(x, W, strides=[1, 1, 1, 1], padding='SAME')
def max_pool_2x2(x):
return tf.nn.max_pool(x, ksize=[1, 2, 2, 1],
strides=[1, 2, 2, 1], padding='SAME')
# -
mnist = input_data.read_data_sets("MNIST_data/", one_hot=True)
sess = tf.InteractiveSession()
x = tf.placeholder(tf.float32, [None, 784])
y_ = tf.placeholder(tf.float32, [None, 10])
W_conv1 = weight_variable([5, 5, 1, 32])
b_conv1 = bias_variable([32])
x_image = tf.reshape(x, [-1,28,28,1])
h_conv1 = tf.nn.relu(conv2d(x_image, W_conv1) + b_conv1)
h_pool1 = max_pool_2x2(h_conv1)
# +
W_conv2 = weight_variable([5, 5, 32, 64])
b_conv2 = bias_variable([64])
h_conv2 = tf.nn.relu(conv2d(h_pool1, W_conv2) + b_conv2)
h_pool2 = max_pool_2x2(h_conv2)
# +
W_fc1 = weight_variable([7 * 7 * 64, 1024])
b_fc1 = bias_variable([1024])
h_pool2_flat = tf.reshape(h_pool2, [-1, 7*7*64])
h_fc1 = tf.nn.relu(tf.matmul(h_pool2_flat, W_fc1) + b_fc1)
# -
keep_prob = tf.placeholder(tf.float32)
h_fc1_drop = tf.nn.dropout(h_fc1, keep_prob)
# +
W_fc2 = weight_variable([1024, 10])
b_fc2 = bias_variable([10])
y_conv=tf.nn.softmax(tf.matmul(h_fc1_drop, W_fc2) + b_fc2)
# +
cross_entropy = -tf.reduce_sum(y_*tf.log(y_conv))
train_step = tf.train.AdamOptimizer(1e-4).minimize(cross_entropy)
correct_prediction = tf.equal(tf.argmax(y_conv,1), tf.argmax(y_,1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
sess.run(tf.initialize_all_variables())
for i in range(20000):
batch = mnist.train.next_batch(50)
if i%100 == 0:
train_accuracy = accuracy.eval(feed_dict={
x:batch[0], y_: batch[1], keep_prob: 1.0})
print("step %d, training accuracy %g"%(i, train_accuracy))
train_step.run(feed_dict={x: batch[0], y_: batch[1], keep_prob: 0.5})
print("test accuracy %g"%accuracy.eval(feed_dict={
x: mnist.test.images, y_: mnist.test.labels, keep_prob: 1.0}))
# -
result = y_conv.eval(feed_dict={
x: mnist.test.images[:1], y_: range(10), keep_prob: 1.0})
result[0]
| Market Analysor.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # Título
#
# ```
# Digite:
# # Título
# ```
# ## Subtítulo
#
# ```
# Digite:
# ## Subtítulo
# ```
# Texto normal
# ## Listas
# Não numerada:
# - Item 1;
# - Item 2;
# - Item 3;
#
# ```
# Digite:
# - Item 1;
# - Item 2;
# - Item 3;
# ```
#
# Numerada:
# 1. Item a;
# 2. Item b;
# 1. Subitem a1
# 2. Subitem a2
# 3. Item c;
#
# ```
# Digite:
# 1. Item a;
# 2. Item b;
# 1. Subitem a1
# 2. Subitem a2
# 3. Item c;
#
# ```
# ## Destaques
#
# Colocar texto **em negrito** ou *em itálico* é muito fácil.
# ```
# Digite:
# Colocar texto **em negrito** ou *em itálico* é muito fácil.
# ```
# ## Imagens
# Vamos inserir uma imagem:
# 
#
# ```
# Digite:
# 
#
# ```
# ## Links internos ou externos
# [Google](https://www.google.com)
#
# [Imagem](./logo-curso.png)
#
# ```
# Digite:
# [Google](https://www.google.com)
#
# [Imagem](./logo-curso.png)
#
# ```
# ## Texto pré-formatado (comandos)
#
# ```
# Texto pré formatado comum.
# ```
# Digite 3 caracteres acento grave, pule uma linha e escreva o texto. Feche com outros e caracteres acento grave.
#
# Em R:
# ```R
# df <- mtcars
# head(df)
# ```
# Digite 3 caracteres acento grave, seguido do nome da linguagem, neste caso "R", pule uma linha e escreva o texto. Feche com outros e caracteres acento grave.
#
| book-R/markdown.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# https://cloud.google.com/dataproc/docs/tutorials/bigquery-sparkml
# -
# # Data Preparation
# +
# #!/usr/bin/env python
# Copyright 2018 Google Inc. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
def run_natality_tutorial(override_values={}):
# [START bigquery_query_natality_tutorial]
"""Create a Google BigQuery linear regression input table.
In the code below, the following actions are taken:
* A new dataset is created "natality_regression."
* A query is run against the public dataset,
bigquery-public-data.samples.natality, selecting only the data of
interest to the regression, the output of which is stored in a new
"regression_input" table.
* The output table is moved over the wire to the user's default project via
the built-in BigQuery Connector for Spark that bridges BigQuery and
Cloud Dataproc.
"""
from google.cloud import bigquery
# Create a new Google BigQuery client using Google Cloud Platform project
# defaults.
client = bigquery.Client()
# Prepare a reference to a new dataset for storing the query results.
dataset_id = 'natality_regression'
# [END bigquery_query_natality_tutorial]
# To facilitate testing, we replace values with alternatives
# provided by the testing harness.
dataset_id = override_values.get("dataset_id", dataset_id)
# [START bigquery_query_natality_tutorial]
dataset = bigquery.Dataset(client.dataset(dataset_id))
# Create the new BigQuery dataset.
dataset = client.create_dataset(dataset)
# In the new BigQuery dataset, create a reference to a new table for
# storing the query results.
table_ref = dataset.table('regression_input')
# Configure the query job.
job_config = bigquery.QueryJobConfig()
# Set the destination table to the table reference created above.
job_config.destination = table_ref
# Set up a query in Standard SQL, which is the default for the BigQuery
# Python client library.
# The query selects the fields of interest.
query = """
SELECT
weight_pounds, mother_age, father_age, gestation_weeks,
weight_gain_pounds, apgar_5min
FROM
`bigquery-public-data.samples.natality`
WHERE
weight_pounds IS NOT NULL
AND mother_age IS NOT NULL
AND father_age IS NOT NULL
AND gestation_weeks IS NOT NULL
AND weight_gain_pounds IS NOT NULL
AND apgar_5min IS NOT NULL
"""
# Run the query.
query_job = client.query(query, job_config=job_config)
query_job.result() # Waits for the query to finish
# [END bigquery_query_natality_tutorial]
if __name__ == '__main__':
run_natality_tutorial()
# -
| natality-regression/1.1_natality_data-preparation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import lsst.sims.maf.plots as plots
import lsst.sims.maf.slicers as slicers
import lsst.sims.maf.metrics as metrics
import lsst.sims.maf.metricBundles as metricBundles
import lsst.sims.maf.db as db
# +
# hack to get the path right
import sys
sys.path.append('..')
from ztf_maf.slicers import HealpixZTFSlicer
from ztf_maf.plots import ZTFBaseSkyMap
# +
slicer = HealpixZTFSlicer(nside=64, lonCol='fieldRA', latCol='fieldDec')
# if we have no field overlaps (single pointing grid), a FieldSlicer will be faster
#slicer = slicers.OpsimFieldSlicer()
# modify to use ZTF field shapes
#slicer.plotFuncs = [ZTFBaseSkyMap, plots.OpsimHistogram]
metricList = []
m1 = metrics.IntraNightGapsMetric()
metricList.append(m1)
m2 = metrics.InterNightGapsMetric()
metricList.append(m2)
m3 = metrics.CountMetric(col='fieldRA')
metricList.append(m3)
filters = ['r','g']
programs = [1,2,3]
summaryMetrics = [metrics.MinMetric(), metrics.MeanMetric(), metrics.MaxMetric(),
metrics.MedianMetric(), metrics.RmsMetric(),
metrics.PercentileMetric(percentile=25), metrics.PercentileMetric(percentile=75)]
bDict={}
for i,metric in enumerate(metricList):
# for j, filt in enumerate(filters):
# sqlconstraint = "filter = '%s'"%(filt)
for j, prog in enumerate(programs):
sqlconstraint = "propID = '%s'"%(prog)
# sqlconstraint=""
bDict[sqlconstraint+metric.name] = metricBundles.MetricBundle(metric, slicer, sqlconstraint,
summaryMetrics=summaryMetrics,plotDict={'radius': np.radians(3.689)})
# +
outDir = 'ZTF_test'
ztfDB = db.OpsimDatabase('../sims/test_schedule_v3.db',
defaultdbTables={'Summary':['Summary','obsHistID']})
ztfDB.raColName = 'fieldRA'
ztfDB.decColName = 'fieldDec'
resultsDb = db.ResultsDb(outDir=outDir)
bgroup = metricBundles.MetricBundleGroup(bDict, ztfDB, outDir=outDir, resultsDb=resultsDb)
bgroup.runAll()
bgroup.plotAll(closefigs=False)
# -
for key in sorted(bDict):
bDict[key].computeSummaryStats(resultsDb=resultsDb)
print key, bDict[key].summaryValues
| notebooks/ZTF_MAF_intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# external libraries
import pandas as pd
import numpy as np
from collections import Counter
from ast import literal_eval
import time
import sys
from shutil import copyfile
from sklearn.metrics import accuracy_score, f1_score
# tensorflow and keras
import keras.optimizers
from keras.datasets import imdb
from keras.models import Model, Sequential
from keras.layers import Input, Dense, Concatenate, Bidirectional, Reshape
from keras.layers import GRU, CuDNNGRU, CuDNNLSTM
from keras.layers.embeddings import Embedding
from keras.constraints import maxnorm
from keras.regularizers import L1L2
from keras.preprocessing import sequence
from keras.callbacks import EarlyStopping
from keras.callbacks import TensorBoard
from keras.backend import tile
import keras.backend as K
from keras.layers import Lambda
# fix random seed for reproducibility - only works for CPU version of tensorflow
np.random.seed(42)
sentences_df = pd.read_csv('../../../../data/processed/tok_sentence_baby_reviews_spell.csv')
reviews_df = pd.read_csv('../../../../data/processed/tok_baby_reviews.csv')
df = reviews_df.merge(sentences_df, on='uuid')
print("\nFiles read, converting tokens to lists.")
for col in ['summary_tokens', 'review_tokens', 'sentence_tokens']:
df[col] = df[col].map(literal_eval)
df.head()
# +
### Preprocessing
# declare the padding and unknown symbols
pad_mask_int = 0
pad_mask_sym = '==pad_mask=='
unknown_int = 1
unknown_sym = '==unknown_sym=='
# vocabulary set
vocab_counter = Counter()
for doc in df['sentence_tokens']:
vocab_counter.update(doc)
min_times_word_used = 2 # if at least 2 then the model will be prepared for unknown words in test and validation sets
print(len(vocab_counter), "tokens before discarding those that appear less than {} times.".format(min_times_word_used))
for key in list(vocab_counter.keys()):
if vocab_counter[key] < min_times_word_used:
vocab_counter.pop(key)
print(len(vocab_counter), "tokens after discarding those that appear less than {} times.".format(min_times_word_used))
vocab_set = set(vocab_counter.keys())
# vocabulary list and int map
vocab_list = [pad_mask_sym, unknown_sym] + sorted(vocab_set)
vocab_map = {word: index for index, word in enumerate(vocab_list)}
# label set
label_set = set(df['sentiment'].unique())
# label list and int map
label_list = sorted(label_set)
label_map = {word: index for index, word in enumerate(label_list)}
# polarity feature set
polarity_set = set(df['polarity'].unique())
# polarity list and int map
polarity_list = sorted(polarity_set)
polarity_map = {word: index for index, word in enumerate(polarity_list)}
# group feature set
group_set = set(df['group_id'].unique())
# group list and int map
group_list = sorted(group_set)
group_map = {word: index for index, word in enumerate(group_list)}
# +
# pretrained embeddings are from https://nlp.stanford.edu/projects/glove/
# start by loading in the embedding matrix
# load the whole embedding into memory
print("\nReading big ol' word embeddings")
count = 0
embeddings_index_1 = dict()
with open('../../../../data/external/glove.twitter.27B.50d.txt') as f:
for line in f:
values = line.split()
word = values[0]
try:
coefs = np.asarray(values[1:], dtype='float32')
except:
print(values)
embeddings_index_1[word] = coefs
print('Loaded %s word vectors.' % len(embeddings_index_1))
#embeddings_index_2 = dict()
#with open('../../../data/external/glove.twitter.27B.100d.txt') as f:
# for line in f:
# values = line.split()
# word = values[0]
# try:
# coefs = np.asarray(values[1:], dtype='float32')
# except:
# print(values)
# embeddings_index_2[word] = coefs
#print('Loaded %s word vectors.' % len(embeddings_index_2))
embedding_dim_1 = 50
embedding_dim_2 = 0
embedding_dim = embedding_dim_1 + embedding_dim_2
# create a weight matrix for words in training docs
if embedding_dim_2 > 0:
embedding_matrix = np.zeros((len(vocab_list), embedding_dim))
for i, word in enumerate(vocab_list):
embedding_vector_1 = embeddings_index_1.get(word)
embedding_vector_2 = embeddings_index_2.get(word)
if embedding_vector_1 is not None and embedding_vector_2 is not None:
embedding_matrix[i] = np.concatenate((embedding_vector_1, embedding_vector_2))
elif embedding_vector_1 is None and embedding_vector_2 is not None:
embedding_matrix[i] = np.concatenate((np.zeros(embedding_dim_1), embedding_vector_2))
elif embedding_vector_1 is not None and embedding_vector_2 is None:
embedding_matrix[i] = np.concatenate((embedding_vector_1, np.zeros(embedding_dim_2)))
else:
print(word)
count += 1 # maybe we should use fuzzywuzzy to get vector of nearest word? Instead of all zeros
else:
embedding_matrix = np.zeros((len(vocab_list), embedding_dim))
for i, word in enumerate(vocab_list):
embedding_vector = embeddings_index_1.get(word)
if embedding_vector is not None:
embedding_matrix[i] = embedding_vector
else:
print(word)
count += 1 # maybe we should use fuzzywuzzy to get vector of nearest word? Instead of all zeros
print(count)
# +
from scipy import sparse
from typing import List, Set, Dict, Tuple, Optional
import numpy as np
def create_one_hot(labels, label_dict: dict):
"""
Args:
labels: array of labels, e.g. NumPy array or Pandas Series
label_dict: dict of label indices
Return:
one_hot_numpy: sparse CSR 2d array of one-hot vectors
"""
one_hot_numpy = sparse.dok_matrix((len(labels), len(label_dict)), dtype=np.int8)
for i, label in enumerate(labels):
one_hot_numpy[i, label_dict[label]] = 1
return sparse.csr_matrix(one_hot_numpy)
def undo_one_hot(pred, label_list: list) -> List[List[str]]:
"""
Args:
pred: NumPy array of one-hot predicted classes
label_list: a list of the label strings
Return:
label_pred: a list of predicted labels
"""
label_pred = [label_list[np.argmax(row)] for row in pred]
return label_pred
# this could probably be done awesomely fast as NumPy vectorised but it works
def word_index(los: List[List[str]], vocab_dict: Dict[str, int], unknown: int, reverse: bool=False) -> List[List[int]]:
"""
Replaces words with integers from a vocabulary dictionary or else with the integer for unknown
Args:
los: list of lists of split sentences
pad_to: how big to make the padded list
unknown: the integer to put in for unknown tokens (either because they were pruned or not seen in training set)
reverse: reverse the order of tokens in the sub-list
Returns:
new_los: list of lists of split sentences where each token is replaced by an integer
Examples:
>>> print(word_index([['one', 'two', 'three'], ['one', 'two']], {'one': 1, 'two': 2, 'three': 3}, unknown=4))
[[1, 2, 3], [1, 2]]
>>> print(word_index([['one', 'two', 'three'], ['one', 'two']], {'one': 1, 'two': 2, 'three': 3}, unknown=4, reverse=True))
[[3, 2, 1], [2, 1]]
"""
new_los = []
if reverse:
for sentence in los:
new_los.append([vocab_dict[word] if word in vocab_dict else unknown for word in sentence][::-1])
else:
for sentence in los:
new_los.append([vocab_dict[word] if word in vocab_dict else unknown for word in sentence])
return new_los
# +
# create one-hot sparse matrix of labels
y = create_one_hot(df['sentiment'], label_map).todense()
# create one-hot of review polarity
polarity = create_one_hot(df['polarity'], polarity_map)[:, 0].todense()
# create one-hot of group number
group = create_one_hot(df['group_id'], group_map).todense()
# replace strings with ints (tokenization is done on the Series fed to word_index())
sentences = word_index(df['sentence_tokens'], vocab_map, unknown_int, reverse=False)
# pad / truncate
from keras.preprocessing.sequence import pad_sequences
sentence_len = max(map(len, list(df['sentence_tokens'])))
sentences = pad_sequences(sequences=sentences,
maxlen=sentence_len,
dtype='int32',
padding='pre',
value=pad_mask_int)
#group = pad_sequences(sequences=group,
# maxlen=embedding_dim,
# dtype='int32',
# padding='pre',
# value=pad_mask_int)
# -
print(sentences[:2])
print(polarity[:2])
print(group[:2])
print(y[:2])
# +
NAME = 'sentences-ablation-group-9-{}'.format(time.strftime('%y%m%d_%H%M', time.localtime(time.time())))
for g in range(6,9):
training_mask = np.logical_or(df['group_id'] != g, df['group_id'] != 9)
validation_mask = df['group_id'] == g
input_s = Input(shape=(sentence_len,), dtype='int32', name='input_s')
input_p = Input(shape=(1,), dtype='float32', name='input_p')
input_g = Input(shape=(len(group_list),), dtype='float32', name='input_g')
embedding_vector_length = embedding_dim
GRU_nodes_sentences = 8
emb = Embedding(len(vocab_list), embedding_vector_length, mask_zero=True,
weights=[embedding_matrix], trainable=False)
emb_s = emb(input_s)
gru_s = GRU(GRU_nodes_sentences,
kernel_initializer='glorot_uniform',
recurrent_initializer='orthogonal',
bias_initializer='zeros',
kernel_regularizer=None,
recurrent_regularizer=None,
bias_regularizer=L1L2(l1=0.1, l2=0.0),
activity_regularizer=L1L2(l1=1e-07, l2=0.0),
kernel_constraint=maxnorm(3),
recurrent_constraint=maxnorm(3),
bias_constraint=None,
return_sequences=False,
return_state=False,
go_backwards=False,
stateful=False,
dropout=0.3)(emb_s)
concat_1 = Concatenate()([gru_s, input_p, input_g]) #
output = Dense(len(label_set), activation='softmax')(gru_s)
model = Model([input_s, input_p, input_g], output) # , ,
nadam = keras.optimizers.nadam(lr=0.001)
model.compile(loss='categorical_crossentropy', optimizer=nadam, metrics=['accuracy'])
print(model.summary())
# es = EarlyStopping(monitor='val_loss', mode='min', verbose=1, patience=10)
tensorboard = TensorBoard(log_dir = './tb_logs/{}'.format('group_'+str(g)+'_'+NAME))
hist1 = model.fit(x=[sentences[training_mask], polarity[training_mask], group[training_mask]], # ,
y=y[training_mask],
validation_data=([sentences[validation_mask],
polarity[validation_mask],
group[validation_mask]], #
y[validation_mask]),
epochs=50, batch_size=64, callbacks=[tensorboard])
pred = model.predict([sentences[validation_mask],
polarity[validation_mask],
group[validation_mask]]) #
pred = undo_one_hot(pred, label_list)
true_sentiment = df.loc[validation_mask, 'sentiment']
f1_micro = f1_score(true_sentiment, pred, average='micro')
f1_macro = f1_score(true_sentiment, pred, average='macro')
accu = accuracy_score(true_sentiment, pred)
metrics_string = """
Group {}
Sklearn
f1 micro {}
f1 macro is {}
Accuracy {}
TF
{}
""".format(g, f1_micro, f1_macro, accu, [key + " " + str(hist1.history[key][-1]) for key in hist1.history.keys()])
print(metrics_string)
with open(NAME+'.txt', mode='a') as fp:
fp.write(metrics_string)
copyfile('sentence_predictions.ipynb', './tb_logs/{}.ipynb'.format(NAME)) # sys.argv[0] for .py files
# -
f1_score(pred, true_sentiment, average='macro')
true_sentiment = df.loc[np.logical_not(group_mask), 'sentiment']
pred = undo_one_hot(pred, label_list)
[key + " " + str(hist1.history[key][-1]) for key in hist1.history.keys()]
| notebooks/Part 3/sentence_predictions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: mydsp
# language: python
# name: mydsp
# ---
# [<NAME>](https://orcid.org/0000-0001-7225-9992),
# Professorship Signal Theory and Digital Signal Processing,
# [Institute of Communications Engineering (INT)](https://www.int.uni-rostock.de/),
# Faculty of Computer Science and Electrical Engineering (IEF),
# [University of Rostock, Germany](https://www.uni-rostock.de/en/)
#
# # Tutorial Signals and Systems (Signal- und Systemtheorie)
#
# Summer Semester 2021 (Bachelor Course #24015)
#
# - lecture: https://github.com/spatialaudio/signals-and-systems-lecture
# - tutorial: https://github.com/spatialaudio/signals-and-systems-exercises
#
# WIP...
# The project is currently under heavy development while adding new material for the summer semester 2021
#
# Feel free to contact lecturer [<EMAIL>](https://orcid.org/0000-0002-3010-0294)
#
# ## Fourier Series Left Time Shift <-> Phase Mod
# +
import numpy as np
import matplotlib.pyplot as plt
def my_sinc(x): # we rather use definition sinc(x) = sin(x)/x, thus:
return np.sinc(x/np.pi)
# +
Th_des = [1, 0.2]
om = np.linspace(-100, 100, 1000)
plt.figure(figsize=(10, 8))
plt.subplot(2,1,1)
for idx, Th in enumerate(Th_des):
A = 1/Th # such that sinc amplitude is always 1
# Fourier transform for single rect pulse
Xsinc = A*Th * my_sinc(om*Th/2)
Xsinc_phase = Xsinc*np.exp(+1j*om*Th/2)
plt.plot(om, Xsinc, 'C7', lw=1)
plt.plot(om, np.abs(Xsinc_phase), label=r'$T_h$=%1.0e s' % Th, lw=5-idx)
plt.legend()
plt.title(r'Fourier transform of single rectangular impulse with $A=1/T_h$ left-shifted by $\tau=T_h/2$')
plt.ylabel(r'magnitude $|X(\mathrm{j}\omega)|$')
plt.xlim(om[0], om[-1])
plt.grid(True)
plt.subplot(2,1,2)
for idx, Th in enumerate(Th_des):
Xsinc = A*Th * my_sinc(om*Th/2)
Xsinc_phase = Xsinc*np.exp(+1j*om*Th/2)
plt.plot(om, np.angle(Xsinc_phase), label=r'$T_h$=%1.0e s' % Th, lw=5-idx)
plt.legend()
plt.xlabel(r'$\omega$ / (rad/s)')
plt.ylabel(r'phase $\angle X(\mathrm{j}\omega)$')
plt.xlim(om[0], om[-1])
plt.ylim(-4, +4)
plt.grid(True)
plt.savefig('1CFE5FE3A1.pdf')
# -
# ## Copyright
#
# This tutorial is provided as Open Educational Resource (OER), to be found at
# https://github.com/spatialaudio/signals-and-systems-exercises
# accompanying the OER lecture
# https://github.com/spatialaudio/signals-and-systems-lecture.
# Both are licensed under a) the Creative Commons Attribution 4.0 International
# License for text and graphics and b) the MIT License for source code.
# Please attribute material from the tutorial as *<NAME>,
# Continuous- and Discrete-Time Signals and Systems - A Tutorial Featuring
# Computational Examples, University of Rostock* with
# ``main file, github URL, commit number and/or version tag, year``.
| ft/FourierTransformation_1CFE5FE3A1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/yukinaga/ai_programming/blob/main/lecture_06/02_cpu_gpu.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="PL-WhmHvI9aI"
# # CPUとGPU
# 実際にディープラーニングを行い、CPUとGPUのパフォーマンスを比較します。
# + [markdown] id="UHe07R4lfzqg"
# ## ●CIFAR-10
# CIFARは、6万枚の画像にそれに写っているものが何なのかを表すラベルをつけたデータセットです。
# 今回は、こちらのデータセットを使ってニューラルネットワークを訓練します。
# 以下のコードでは、CIFAR-10を読み込み、ランダムな25枚の画像を表示します。
# + id="tvE2Zb3catc3"
import numpy as np
import matplotlib.pyplot as plt
import tensorflow.keras
from tensorflow.keras.datasets import cifar10
(x_train, t_train), (x_test, t_test) = cifar10.load_data()
print("Image size:", x_train[0].shape)
cifar10_labels = np.array(["airplane", "automobile", "bird", "cat", "deer",
"dog", "frog", "horse", "ship", "truck"])
n_image = 25
rand_idx = np.random.randint(0, len(x_train), n_image)
plt.figure(figsize=(10,10)) # 画像の表示サイズ
for i in range(n_image):
cifar_img=plt.subplot(5,5,i+1)
plt.imshow(x_train[rand_idx[i]])
label = cifar10_labels[t_train[rand_idx[i]]]
plt.title(label)
plt.tick_params(labelbottom=False, labelleft=False, bottom=False, left=False) # ラベルとメモリを非表示に
plt.show()
# + [markdown] id="VY2aX1CaZnK-"
# ## ●パフォーマンス比較
# 以下はフレームワークKerasを使って実装した、典型的な畳み込みニューラルネットワークのコードです。
# ニューラルネットワークが5万枚の画像を学習します。
# CPUとGPUで、学習に要する時間を比較しましょう。
# デフォルトではCPUが使用されますが、編集→ノートブックの設定のハードウェアアクセラレーターでGPUを選択することでGPUが使用されるようになります。
# 実行時間は、実行結果が表示される領域の左のアイコンにカーソルを合わせると確認することができます。
# + id="ciwFomv_50Hm"
import numpy as np
import matplotlib.pyplot as plt
from tensorflow.keras.datasets import cifar10
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout, Activation, Flatten
from tensorflow.keras.layers import Conv2D, MaxPooling2D
from tensorflow.keras.optimizers import Adam
(x_train, t_train), (x_test, t_test) = cifar10.load_data()
batch_size = 32
epochs = 1
n_class = 10
t_train = to_categorical(t_train, n_class)
t_test = to_categorical(t_test, n_class)
model = Sequential()
model.add(Conv2D(32, (3, 3), padding='same', input_shape=x_train.shape[1:]))
model.add(Activation('relu'))
model.add(Conv2D(32, (3, 3)))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(64, (3, 3), padding='same'))
model.add(Activation('relu'))
model.add(Conv2D(64, (3, 3)))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten())
model.add(Dense(256))
model.add(Activation('relu'))
model.add(Dropout(0.5))
model.add(Dense(n_class))
model.add(Activation('softmax'))
model.compile(optimizer=Adam(), loss='categorical_crossentropy', metrics=['accuracy'])
x_train = x_train / 255
x_test = x_test / 255
model.fit(x_train, t_train, epochs=epochs, batch_size=batch_size, validation_data=(x_test, t_test))
# + [markdown] id="xMBacVeeib9F"
# CPUの場合、コードの実行時間は約240秒、GPUの場合は約43秒でした。
# このように、GPUを利用することで学習に要する時間を大幅に短縮することができます。
| lecture_06/02_cpu_gpu.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # VacationPy
# ----
#
# #### Note
# * Keep an eye on your API usage. Use https://developers.google.com/maps/reporting/gmp-reporting as reference for how to monitor your usage and billing.
#
# * Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
import gmaps
import json
import requests
from api_keys import api_key
from api_keys import g_key
import random
import pandas as pd
import numpy as np
import json
import matplotlib.pyplot as plt
from scipy.stats import linregress
import os
# Access maps with unique API key
gmaps.configure(api_key=g_key)
# ### Store Part I results into DataFrame
# * Load the csv exported in Part I to a DataFrame
# +
#cities_pd = pd.read_csv("worldcities.csv")
cities_pd = pd.read_csv("cities.csv")
cities_pd.head(100)
# -
# ### Humidity Heatmap
# * Configure gmaps.
# * Use the Lat and Lng as locations and Humidity as the weight.
# * Add Heatmap layer to map.
# +
url = "http://api.openweathermap.org/data/2.5/weather?"
#cities = cities_pd["city_ascii"]
#api.openweathermap.org/data/2.5/weather?lat={lat}&lon={lon}&appid={your api key}
cities = cities_pd["City"]
cntry = cities_pd["Country"]
lat = cities_pd["Lat"]
lng = cities_pd["Lng"]
temper = cities_pd["Max Temp"]
hum = cities_pd["Humidity"]
cloud = cities_pd["Cloudiness"]
speed = cities_pd["Wind Speed"]
nor_lat = []
nor_hum = []
nor_temper = []
nor_cloud = []
nor_speed = []
sou_lat = []
sou_hum = []
sou_temper = []
sou_cloud = []
sou_speed = []
units = "metric"
impl = "imperial"
query_url = f"{url}appid={api_key}&units={impl}&q="
# +
# Get the indices of cities that have humidity over 100%.
# Make a new DataFrame equal to the city data to drop all humidity outliers by index.
# Passing "inplace=False" will make a copy of the city_data DataFrame, which we call "clean_city_data".
#by default all humidity are less than 100
for index, row in cities_pd.iterrows():
try:
if (row["Lat"] >= 0 ):
nor_lat.append(row['Lat'])
nor_temper.append(row['Max Temp'])
nor_hum.append(row['Humidity'])
nor_speed.append(row['Wind Speed'])
nor_cloud.append(row['Cloudiness'])
else:
sou_lat.append(row['Lat'])
sou_temper.append(row['Max Temp'])
sou_hum.append(row['Humidity'])
sou_speed.append(row['Wind Speed'])
sou_cloud.append(row['Cloudiness'])
except:
pass
weather_dict = {
"lat": lat,
"lng": lng,
"temper": temper,
"cloud": cloud,
"speed": speed,
"hum": hum
}
weather_data = pd.DataFrame(weather_dict)
weather_data.to_csv('cities_with_temper.csv', index=True)
# +
# Plot Heatmap
locations = weather_data[["lat", "lng"]]
humidty = weather_data["hum"]
fig = gmaps.figure()
# Create heat layer
heat_layer = gmaps.heatmap_layer(locations, weights=humidty,
dissipating=False, max_intensity=100,
point_radius=1)
# Add layer
fig.add_layer(heat_layer)
# Display figure
fig
# -
# ### Create new DataFrame fitting weather criteria
# * Narrow down the cities to fit weather conditions.
# * Drop any rows will null values.
# +
#perfect weather conditions
per_data = np.arange(len(weather_data))
percnt=0
npercnt=0
perfect_weather_dict = {
"lat": [],
"lng": [],
"temper": [],
"cloud": [],
"speed": [],
"hum": []
}
not_perfect_weather_dict = {
"lat": [],
"lng": [],
"temper": [],
"cloud": [],
"speed": [],
"hum": []
}
per_coordinates = []
not_per_coordinates = []
for x in per_data:
latlng = ()
if weather_data["temper"][x] < 80 and weather_data["temper"][x] > 70 and weather_data["speed"][x] < 10 and weather_data["cloud"][x] == 0:
perfect_weather_dict["lat"].append(weather_data["lat"][x])
perfect_weather_dict["lng"].append(weather_data["lng"][x])
perfect_weather_dict["temper"].append(weather_data["temper"][x])
perfect_weather_dict["cloud"].append(weather_data["cloud"][x])
perfect_weather_dict["speed"].append(weather_data["speed"][x])
perfect_weather_dict["hum"].append(weather_data["hum"][x])
latlng=weather_data["lat"][x],weather_data["lng"][x]
per_coordinates.append(latlng)
percnt=percnt+1
else:
not_perfect_weather_dict["lat"].append(weather_data["lat"][x])
not_perfect_weather_dict["lng"].append(weather_data["lng"][x])
not_perfect_weather_dict["temper"].append(weather_data["temper"][x])
not_perfect_weather_dict["cloud"].append(weather_data["cloud"][x])
not_perfect_weather_dict["speed"].append(weather_data["speed"][x])
not_perfect_weather_dict["hum"].append(weather_data["hum"][x])
latlng=weather_data["lat"][x],weather_data["lng"][x]
not_per_coordinates.append(latlng)
npercnt=npercnt+1
perfect_weather_data = pd.DataFrame(perfect_weather_dict)
not_perfect_weather_data = pd.DataFrame(not_perfect_weather_dict)
#not_perfect_weather_data.head()
#len(not_perfect_weather_data)
# Customize the size of the figure
figure_layout = {
'width': '700px',
'height': '500px',
'border': '1px solid black',
'padding': '1px',
'margin': '0 auto 0 auto'
}
fig = gmaps.figure(layout=figure_layout)
# Assign the marker layer to a variable
markers = gmaps.marker_layer(per_coordinates)
# Add the layer to the map
fig.add_layer(markers)
fig
# +
#not perfect weather conditions
figure_layout = {
'width': '700px',
'height': '500px',
'border': '1px solid black',
'padding': '1px',
'margin': '0 auto 0 auto'
}
fig = gmaps.figure(layout=figure_layout)
# Assign the marker layer to a variable
markers = gmaps.marker_layer(not_per_coordinates)
# Add the layer to the map
fig.add_layer(markers)
fig
# -
# ### Hotel Map
# * Store into variable named `hotel_df`.
# * Add a "Hotel Name" column to the DataFrame.
# * Set parameters to search for hotels with 5000 meters.
# * Hit the Google Places API for each city's coordinates.
# * Store the first Hotel result into the DataFrame.
# * Plot markers on top of the heatmap.
# +
base_url = "https://maps.googleapis.com/maps/api/place/nearbysearch/json?"
params = {
"key": g_key,
"radius": "50"
}
# set up additional columns to hold information
perfect_weather_data['name'] = ""
perfect_weather_data['address'] = ""
perfect_weather_data['country_code'] = ""
dataset= []
locations = perfect_weather_data[["lat", "lng"]]
# use iterrows to iterate through pandas dataframe
for index, row in perfect_weather_data.iterrows():
base_url = "https://maps.googleapis.com/maps/api/place/nearbysearch/json?"
# get restaurant type from df
#restr_type = row['hotel']
# add keyword to params dict
params['keyword'] = "hotel"
params['location'] = row["lat"],row["lng"]
# assemble url and make API request
#print(f"Retrieving Results for Index {index}: {restr_type}.")
base_url= base_url+"location="+str(row['lat'])+","+str(row['lng'])+"&radius=5000&keyword=hotel&key="+g_key
response = requests.get(base_url).json()
# extract results
results = response['results']
# dataset.append(results)
try:
perfect_weather_data.loc[index, 'name'] = results[0]['name']
perfect_weather_data.loc[index, 'address'] = results[0]['vicinity']
perfect_weather_data.loc[index, 'country_code'] = results[0]['plus_code']['compound_code']
except:
pass
#hotels_nearby = perfect_weather_data["name"].tolist()
#vicinity_nearby = perfect_weather_data["address"].tolist()
#country_nearby = perfect_weather_data["country_code"].tolist()
x_axis = np.arange(len(perfect_weather_data))
fig = gmaps.figure(layout=figure_layout)
markers = gmaps.marker_layer(locations,
info_box_content=[f'<dl><dt>Hotel name :</dt><dd> {perfect_weather_data["name"][ind]} </dd><dt> Address :</dt><dd> {perfect_weather_data["address"][ind]} </dd><dt> Country code : </dt><dd>{perfect_weather_data["country_code"][ind]} </dd></dl>' for ind in x_axis])
fig.add_layer(markers)
humidty = perfect_weather_data["hum"]
heat_layer = gmaps.heatmap_layer(locations, weights=humidty,
dissipating=False, max_intensity=100,
point_radius=1)
# Add layer
fig.add_layer(heat_layer)
fig
# -
# +
# NOTE: Do not change any of the code in this cell
# Using the template add the hotel marks to the heatmap
#info_box_template = """
##<dt>Name</dt><dd>{Hotel Name}</dd>
#<dt>City</dt><dd>{City}</dd>
#<dt>Country</dt><dd>{Country}</dd>
#</dl>
#"""
# Store the DataFrame Row
# NOTE: be sure to update with your DataFrame name
#hotel_info = [info_box_template.format(**row) for index, row in hotel_df.iterrows()]
#locations = hotel_df[["Lat", "Lng"]]
# +
# Add marker layer ontop of heat map
# Display figure
#hotels_nearby = perfect_weather_data["name"].tolist()
#vicinity_nearby = perfect_weather_data["address"].tolist()
#country_nearby = perfect_weather_data["country_code"].tolist()
x_axis = np.arange(len(perfect_weather_data))
fig = gmaps.figure(layout=figure_layout)
markers = gmaps.marker_layer(locations,
info_box_content=[f'<dl><dt>Hotel name :</dt><dd> {perfect_weather_data["name"][ind]} </dd><dt> Address :</dt><dd> {perfect_weather_data["address"][ind]} </dd><dt> Country code : </dt><dd>{perfect_weather_data["country_code"][ind]} </dd></dl>' for ind in x_axis])
fig.add_layer(markers)
humidty = perfect_weather_data["hum"]
heat_layer = gmaps.heatmap_layer(locations, weights=humidty,
dissipating=False, max_intensity=100,
point_radius=1)
# Add layer
fig.add_layer(heat_layer)
fig
# -
| VacationPy/VacationPy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# name: python3
# ---
# +
from datasets import load_from_disk
wiki_datasets = load_from_disk("/opt/ml/data/wiki_preprocessed_droped")
wiki_datasets.load_elasticsearch_index("text", host="localhost", port="9200", es_index_name="wikipedia_contexts")
# -
query = "이순신 장군은 언제 태어났는가?"
scores, retrieved_examples = wiki_datasets.get_nearest_examples("text", query, k=3)
scores, retrieved_examples
# 인덱스 제거하기
# !curl -XDELETE localhost:9200/wikipedia_contexts
| install/elasticsearch/elasticsearch_load.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# This app demonstrates how to use a variety of different [plotting libraries and data types](https://github.com/pyviz/panel/issues/2) with Panel. Here let's set up four different plotting libraries controlled by a couple of widgets, for <NAME>'s [gapminder](https://demo.bokehplots.com/apps/gapminder) example.
#
# **Live App URL**: https://gapminders.pyviz.demo.anaconda.com
#
# The app is defined as a notebook ipynb file and can also be viewed on MyBinder.org:
#
# [](https://mybinder.org/v2/gh/panel-demos/gapminder/master?urlpath=/gapminder.ipynb)
#
# <style>
# .iframe-container {
# overflow: hidden;
# padding-top: 56.25%;
# position: relative;
# background-size: 80%;
# background: url(https://raw.githubusercontent.com/panel-demos/gapminder/master/assets/dashboard.png) center center no-repeat;
# }
#
# .iframe-container iframe {
# border: 0;
# height: 100%;
# left: 0;
# position: absolute;
# top: 0;
# width: 100%;
# }
# </style>
#
# <div class="iframe-container">
# <iframe src="https://gapminders.pyviz.demo.anaconda.com" width="100%" frameborder="0" onload="this.parentNode.style.background = 'none'"></iframe>
# </div>
| examples/gallery/demos/gapminders.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# - Import SEIRs+ module
from seirsplus.models import *
# - Load required libraries
import networkx
# %matplotlib inline
# - Create graph
numNodes = 10000
baseGraph = networkx.barabasi_albert_graph(n=numNodes, m=9)
# - define probabilistic distributions
G_normal = custom_exponential_graph(baseGraph, scale=100)
# Social distancing interactions:
G_distancing = custom_exponential_graph(baseGraph, scale=10)
# Quarantine interactions:
G_quarantine = custom_exponential_graph(baseGraph, scale=5)
# - create model
# Params: G Network adjacency matrix (numpy array) or Networkx graph object.
# beta Rate of transmission (exposure)
# sigma Rate of infection (upon exposure)
# gamma Rate of recovery (upon infection)
# xi Rate of re-susceptibility (upon recovery)
# mu_I Rate of infection-related death
# mu_0 Rate of baseline death
# nu Rate of baseline birth
# p Probability of interaction outside adjacent nodes
#
# Q Quarantine adjacency matrix (numpy array) or Networkx graph object.
# beta_D Rate of transmission (exposure) for individuals with detected infections
# sigma_D Rate of infection (upon exposure) for individuals with detected infections
# gamma_D Rate of recovery (upon infection) for individuals with detected infections
# mu_D Rate of infection-related death for individuals with detected infections
# theta_E Rate of baseline testing for exposed individuals
# theta_I Rate of baseline testing for infectious individuals
# phi_E Rate of contact tracing testing for exposed individuals
# phi_I Rate of contact tracing testing for infectious individuals
# psi_E Probability of positive test results for exposed individuals
# psi_I Probability of positive test results for exposed individuals
# q Probability of quarantined individuals interaction outside adjacent nodes
#
# initE Init number of exposed individuals
# initI Init number of infectious individuals
# initD_E Init number of detected infectious individuals
# initD_I Init number of detected infectious individuals
# initR Init number of recovered individuals
# initF Init number of infection-related fatalities
# (all remaining nodes initialized susceptible)
model = SEIRSNetworkModel(G=G_normal, beta=0.155, sigma=1/5.2, gamma=1/12.39, mu_I=0.0004, p=0.5,
Q=G_quarantine, beta_D=0.155, sigma_D=1/5.2, gamma_D=1/12.39, mu_D=0.0004,
theta_E=0.02, theta_I=0.02, phi_E=0.2, phi_I=0.2, psi_E=1.0, psi_I=1.0, q=0.5,
initI=10)
# - checkpoints: change simulation parameters during the simulation
# (e.g., quarantine)
checkpoints = {'t': [20, 100], 'G': [G_distancing, G_normal],
'p': [0.1, 0.5], 'theta_E': [0.02, 0.02], 'theta_I': [0.02, 0.02], 'phi_E': [0.2, 0.2]}
# - Compute the model
time_span = 200 # days
model.run(T=time_span, checkpoints=checkpoints)
# - Make a nice plot
model.figure_infections()
| ntb/SEIRsplus_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# create index and document
# +
import sys, os, json
sys.path.append(sys.path.append(os.path.join(os.getcwd(), '..')))
from datamart.query_manager import QueryManager
from datamart.index_builder import IndexBuilder
es_index = "datamart_tmp" # es index for your metadata, make sure you change to your own
index_builder = IndexBuilder()
# -
tmp_description_dir = "tmp" # dataset schema json file
tmp_description = "tmp/tmp.json" # dir of all dataset schema json files
tmp_out = "tmp/tmp.metadata" # output of metadata
# Indexing single dataset
this_metadata = index_builder.indexing(description_path=tmp_description,
es_index=es_index,
data_path=None,
query_data_for_indexing=True,
save_to_file=tmp_out,
save_to_file_mode="w",
delete_old_es_index=True)
# Take a look at the last metadata generated
print(json.dumps(this_metadata, indent=2))
# Bulk indexing multiple dataset
index_builder.bulk_indexing(description_dir=tmp_description_dir,
es_index=es_index,
data_dir=None,
query_data_for_indexing=True,
save_to_file=tmp_out,
save_to_file_mode="w",
delete_old_es_index=True)
# Index is built in elasticsearch, check it here: http://dsbox02.isi.edu:9200/_cat/indices?v
#
# Try some query through Kibana: http://dsbox02.isi.edu:5601/app/kibana#/dev_tools/console?_g=()
# Output metadata is written to `save_to_file` in indexing
| test/indexing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="3zHDt_FdV04L" outputId="4<PASSWORD>"
import numpy as np
import torch
from torch import distributions
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.utils.data import Dataset
from torchvision import datasets
import torchvision.transforms as T
from torchvision.utils import make_grid, save_image
import time
from PIL import Image
from tqdm import tqdm
from matplotlib import pyplot as plt
import os
# %matplotlib inline
# %pip install pytorch-ignite
# %pip install --pre pytorch-ignite
# %pip install torchsummary
from ignite.metrics import FID
from torchsummary import summary
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
to_pil_image = T.ToPILImage()
# -
# # Definition of VAE with Gaussian Prior and Flow-Based Prior (RealNVP)
# + id="4DflX6CyV04Q"
class Encoder(nn.Module):
def __init__(self, input_size, hidden_size = 300, latent_size = 100):
super(Encoder, self).__init__()
self.encoder = nn.Sequential(
nn.Linear(input_size, hidden_size),
nn.LeakyReLU(),
nn.Linear(hidden_size, hidden_size),
nn.LeakyReLU(),
nn.Linear(hidden_size, 2 * latent_size)
)
def sample(self, mu, sigma):
eps = torch.randn_like(sigma)
return mu + sigma * eps
def forward(self, x):
h = self.encoder(x)
mu, log_var = torch.chunk(h, 2, dim=1)
sigma = torch.exp(0.5*log_var)
z = self.sample(mu, sigma)
return z, mu, sigma
class Decoder(nn.Module):
def __init__(self, output_size, hidden_size = 300, latent_size = 100):
super(Decoder, self).__init__()
self.decoder = nn.Sequential(
nn.Linear(latent_size, hidden_size),
nn.LeakyReLU(),
nn.Linear(hidden_size, hidden_size),
nn.LeakyReLU(),
nn.Linear(hidden_size, output_size),
nn.Sigmoid()
)
def forward(self, x):
return self.decoder(x)
class VAE(nn.Module):
def __init__(self, input_size):
super(VAE, self).__init__()
self.encoder = Encoder(input_size)
self.decoder = Decoder(input_size)
def forward(self, x):
z, mu, sigma = self.encoder(x)
return self.decoder(z), z, mu, sigma
class GaussianPrior(nn.Module):
def __init__(self, latent_size=100):
super(GaussianPrior, self).__init__()
self.latent_size = latent_size
def sample(self, batch_size):
z = torch.randn((batch_size, self.latent_size))
return z
def log_prob(self, z):
PI = torch.from_numpy(np.asarray(np.pi)).to(device)
return -0.5 * torch.log(2. * PI) - 0.5 * z**2.
class FlowPrior(nn.Module):
def __init__(self, nets, nett, num_flows, D=2):
super(FlowPrior, self).__init__()
self.D = D
self.t = torch.nn.ModuleList([nett() for _ in range(num_flows)])
self.s = torch.nn.ModuleList([nets() for _ in range(num_flows)])
self.num_flows = num_flows
def coupling(self, x, index, forward=True):
(xa, xb) = torch.chunk(x, 2, 1)
s = self.s[index](xa)
t = self.t[index](xa)
if forward:
#yb = f^{-1}(x)
yb = (xb - t) * torch.exp(-s)
else:
#xb = f(y)
yb = torch.exp(s) * xb + t
return torch.cat((xa, yb), 1), s
def permute(self, x):
return x.flip(1)
def f(self, x):
log_det_J, z = x.new_zeros(x.shape[0]), x
for i in range(self.num_flows):
z, s = self.coupling(z, i, forward=True)
z = self.permute(z)
log_det_J = log_det_J - s.sum(dim=1)
return z, log_det_J
def f_inv(self, z):
x = z
for i in reversed(range(self.num_flows)):
x = self.permute(x)
x, _ = self.coupling(x, i, forward=False)
return x
def sample(self, batch_size):
z = torch.randn(batch_size, self.D)
x = self.f_inv(z)
return x.view(-1, self.D)
def log_prob(self, x):
z, log_det_J = self.f(x)
PI = torch.from_numpy(np.asarray(np.pi))
log_standard_normal = -0.5 * torch.log(2. * PI) - 0.5 * z**2.
log_p = (log_standard_normal + log_det_J.unsqueeze(1))
return -log_p
class ELBO():
def __init__(self, prior):
self.prior = prior
self.reconstruction_error = nn.BCELoss(reduction='none')
def kullback_Leibler_divergence(self, z, mu, sigma):
q = torch.distributions.Normal(mu, sigma)
log_qz = q.log_prob(z)
log_pz = self.prior.log_prob(z)
kl = (log_qz - log_pz).sum(-1)
return kl
def __call__(self, inputs, outputs, z, mu, sigma):
re = self.reconstruction_error(outputs, inputs).sum(-1)
kl = self.kullback_Leibler_divergence(z, mu, sigma)
elbo = (re + kl)
return elbo.mean()
# + id="w99av-UjV04R"
def train(net, prior, train_data, val_data, img_dim, batch_size=10, learning_rate=0.0001, epochs=20, nr_test_samples=64, img_dir='None'):
optimizer = optim.Adam(net.parameters(), lr=learning_rate)
criterion = ELBO(prior)
train_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size, shuffle=True)
val_loader = torch.utils.data.DataLoader(val_data, batch_size=batch_size, shuffle=True)
running_loss = []
running_val_loss = []
# m = FID()
for epoch in range(epochs):
print("Epoch:" , epoch+1)
for i, data in tqdm(enumerate(train_loader), total=int(len(train_data)/train_loader.batch_size)):
optimizer.zero_grad()
inputs, _ = data
inputs = inputs.to(device)
# Forward
outputs, z, mu, sigma = net(inputs)
# Backward
loss = criterion(inputs, outputs, z, mu, sigma)
loss.backward()
optimizer.step()
running_loss.append(loss.item())
sample = prior.sample(nr_test_samples)
generated_img = net.decoder(sample).view(nr_test_samples,img_dim,32,32)
generated_img = make_grid(generated_img)
im = Image.fromarray(np.array(to_pil_image(generated_img)))
im.save(f"{img_dir}/epoch_{epoch}.jpeg")
with torch.no_grad():
for inputs, _ in val_loader:
inputs = inputs.to(device)
outputs, z, mu, sigma = net(inputs)
loss = criterion(inputs, outputs, z, mu, sigma)
running_val_loss.append(loss)
# m.update(outputs, inputs)
print(f'Train Loss: {np.mean(running_loss[-len(train_data):])} | Validation Loss: {np.mean(running_val_loss[-len(val_data):])}')
# print(m.compute())
return running_loss, running_val_loss
# + id="CTzOx7eNV04T"
def plot_interpolated(net, img_dim, n=10):
w = 32
img = np.zeros((n*w, n*w, img_dim))
s, e1, e2 = prior.sample(3)
for i, y in enumerate(np.linspace(0, 1, n)):
for j, x in enumerate(np.linspace(0, 1, n)):
z = s + (e1-s) * x + (e2-s) * y
generated_image = net.decoder(z.view(-1,100)).view(img_dim, 32,32)
img[(n-1-i)*w:(n-1-i+1)*w, j*w:(j+1)*w,:] = generated_image.detach().numpy().transpose((1,2,0))
plt.imshow(img)
# -
# ## Standard VAE on MNIST
# + id="XR3NsVMnV04N"
mnist_train = datasets.MNIST(root='data', train=True, download=True, transform=T.Compose([T.Resize(32), T.ToTensor(), T.Lambda(lambda x: torch.flatten(x))]))
# + id="TFmCywVrZyzS"
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
# -
input_size = 32*32
batch_size=64
net1 = VAE(input_size).to(device)
prior = GaussianPrior()
print("Number of trainable parameters in VAE:", count_parameters(net1))
print("Number of trainable parameters in Prior:", count_parameters(prior))
train_set, val_set = torch.utils.data.random_split(mnist_train, [55000, 5000])
train_loss, val_loss = train(net1, prior, train_set, val_set, 1, batch_size=batch_size, epochs=50, img_dir='VAE/MNIST')
# +
train_loss = np.array(train_loss).reshape(-1, int(55000/64)+1).mean(axis=1)
val_loss = np.array(val_loss).reshape(-1, int(5000/64)+1).mean(axis=1)
plt.plot(train_loss, label='Training')
plt.plot(val_loss, label='Validation')
plt.xlabel('Epoch')
plt.ylabel('Loss')
plt.title('Loss for VAE on MNIST')
plt.legend()
plt.show()
# -
plot_interpolated(net1, 1)
# ## Standard VAE on SVHN
# + id="1GMzFPLUV04O"
svhn_train = datasets.SVHN(root='data', download=True, transform=T.Compose([T.ToTensor(), T.Lambda(lambda x: torch.flatten(x))]))
# -
input_size = 32*32*3
net2 = VAE(input_size).to(device)
prior = GaussianPrior()
print("Number of trainable parameters in VAE:", count_parameters(net2))
print("Number of trainable parameters in Prior:", count_parameters(prior))
train_set, val_set = torch.utils.data.random_split(svhn_train, [65000, 8257])
train_loss, val_loss = train(net2, prior, train_set, val_set, 3, batch_size=batch_size, epochs=50, img_dir='VAE/SVHN')
# +
train_loss = np.array(train_loss).reshape(-1, int(65000/64)+1).mean(axis=1)
val_loss = np.array(val_loss).reshape(-1, int(8257/64)+1).mean(axis=1)
plt.plot(train_loss, label='Training')
plt.plot(val_loss, label='Validation')
plt.xlabel('Epoch')
plt.ylabel('Loss')
plt.title('Loss for VAE on SVHN')
plt.legend()
plt.show()
# -
plot_interpolated(net2, 3)
# + [markdown] id="6Sr8RbqlV04S"
# ## RealNVP on MNIST
# + id="A10yvuBFaCX3"
num_flows = 3
L = 100
M = 300
nets = lambda: nn.Sequential(nn.Linear(L // 2, M), nn.LeakyReLU(),
nn.Linear(M, M), nn.LeakyReLU(),
nn.Linear(M, L // 2), nn.Tanh())
nett = lambda: nn.Sequential(nn.Linear(L // 2, M), nn.LeakyReLU(),
nn.Linear(M, M), nn.LeakyReLU(),
nn.Linear(M, L // 2))
prior = RealNVP(nets, nett, num_flows=num_flows, D=L)
batch_size = 64
# + colab={"base_uri": "https://localhost:8080/", "height": 511} id="lLVdqzngV04S" jupyter={"outputs_hidden": true} outputId="2e249f04-ed94-466d-e60b-9630c6f8dc48"
input_size = 32*32
net3 = VAE(input_size).to(device)
print("Number of trainable parameters in VAE:", count_parameters(net3))
print("Number of trainable parameters in Prior:", count_parameters(prior))
train_set, val_set = torch.utils.data.random_split(mnist_train, [55000, 5000])
train_loss, val_loss = train(net3, prior, train_set, val_set, 1, batch_size=batch_size, epochs=50, img_dir='RealNVP/MNIST')
# + id="Q3PtcdOHV04S"
train_loss = np.array(train_loss).reshape(-1, int(55000/64)+1).mean(axis=1)
val_loss = np.array(val_loss).reshape(-1, int(5000/64)+1).mean(axis=1)
plt.plot(train_loss, label='Training')
plt.plot(val_loss, label='Validation')
plt.xlabel('Epoch')
plt.ylabel('Loss')
plt.title('Loss for RealNVP on MNIST')
plt.legend()
plt.show()
# + id="nMxRxeouV04T"
plot_interpolated(net3, 1)
# + [markdown] id="Pk-S6eLia6Mh"
# # RealNVP on SVHN
# + id="m0JoctcQa9BK"
input_size = 32*32*3
net4 = VAE(input_size).to(device)
print("Number of trainable parameters in VAE:", count_parameters(net4))
print("Number of trainable parameters in Prior:", count_parameters(prior))
train_set_svhn, val_set_svhn = torch.utils.data.random_split(svhn_train, [65000, 8257])
train_loss_svhn, val_loss_svhn = train(net4, prior, train_set_svhn, val_set_svhn, 3, batch_size=batch_size, epochs=50, img_dir='RealNVP/SVHN')
# + id="QjzrVDvmV04U"
train_loss_epoch = np.array(train_loss_svhn).reshape(-1, int(60000/64)+1).mean(axis=1)
val_loss_epoch = np.array(val_loss_svhn).reshape(-1, int(13257/64)+1).mean(axis=1)
plt.plot(train_loss_epoch, label='Training')
plt.plot(val_loss_epoch, label='Validation')
plt.xlabel('Epoch')
plt.ylabel('Loss')
plt.title('Loss for RealNVP on SVHN')
plt.legend()
plt.show()
# + id="bsn5zMCnV04U"
plot_interpolated(net4, 3)
# -
| Project 4/VAE_Priors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + slideshow={"slide_type": "skip"}
# !jupyter nbconvert eesardocs.ipynb --to slides --post serve
# + slideshow={"slide_type": "skip"}
import warnings
# these are innocuous but irritating
warnings.filterwarnings("ignore", message="numpy.dtype size changed")
warnings.filterwarnings("ignore", message="numpy.ufunc size changed")
# + [markdown] slideshow={"slide_type": "slide"}
# # Change Detection with Sentinel-1 PolSAR imagery on the GEE
#
# ### <NAME>
# <EMAIL>
# ### <NAME>, <NAME>
# Jülich Forschungszentrum, Germany
# ### <NAME>, <NAME>, <NAME>
# Technical University of Denmark
# ### September 2018
# + [markdown] slideshow={"slide_type": "slide"}
# ## Software Installation
#
# Pull and/or run the container with
#
# docker run -d -p 443:8888 --name=eesar mort/eesardocker
#
# or, if you are on a Raspberry Pi,
#
# docker run -d -p 443:8888 --name=eesar mort/rpi-eesardocker
#
# Point your browser to http://localhost:443 to see the Jupyter notebook home page.
#
# Open the Notebook
#
# interface.ipynb
#
# Stop the container with
#
# docker stop eesar
#
# Re-start with
#
# docker start eesar
# + [markdown] slideshow={"slide_type": "slide"}
# ### The GEE Sentinel-1 Archive
#
# https://explorer.earthengine.google.com/#detail/COPERNICUS%2FS1_GRD
# + [markdown] slideshow={"slide_type": "slide"}
# ## Background
#
# ### Vector and matrix representations
#
# A fully polarimetric SAR measures a
# $2\times 2$ _scattering matrix_ $S$ at each resolution cell on the ground.
# The scattering matrix relates the incident and the backscattered
# electric fields $E^i$ and $E^b$ according to
#
# $$
# \pmatrix{E_h^b \cr E_v^b}
# =\pmatrix{S_{hh} & S_{hv}\cr S_{vh} & S_{vv}}\pmatrix{E_h^i \cr E_v^i}.
# $$
#
# The per-pixel polarimetric information in the scattering matrix $S$, under the assumption
# of reciprocity ($S_{hv} = S_{vh}$), can then be expressed as a three-component complex vector
#
# $$
# s = \pmatrix{S_{hh}\cr \sqrt{2}S_{hv}\cr S_{vv}},
# $$
#
# The total intensity is referred to as the _span_ and is the complex inner product of the vector $s$,
#
# $$
# {\rm span} = s^\top s = |S_{hh}|^2 + 2|S_{hv}|^2 + |S_{vv}|^2.
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# The polarimetric signal is can also be represented by taking the complex outer product of $s$ with itself:
#
# $$
# C = s s^\top = \pmatrix{ |S_{hh}|^2 & \sqrt{2}S_{hh}S_{hv}^* & S_{hh}S_{vv}^* \cr
# \sqrt{2}S_{hv}S_{hh}^* & 2|S_{hv}|^2 & \sqrt{2}S_{hv}S_{vv}^* \cr
# S_{vv}S_{hh}^* & \sqrt{2}S_{vv}S_{hv}^* & |S_{vv}|^2 }.
# $$
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Multi-looking
#
# The matrix $C$ can be averaged over the number of looks (number of adjacent cells used to average out the effect of speckle) to give an estimate of the __covariance matrix__ of each multi-look pixel:
#
# $$
# \bar{C} ={1\over m}\sum_{\nu=1}^m s(\nu) s(\nu)^\top = \langle s s^\top \rangle
# = \pmatrix{ \langle |S_{hh}|^2\rangle & \langle\sqrt{2}S_{hh}S_{hv}^*\rangle & \langle S_{hh}S_{vv}^*\rangle \cr
# \langle\sqrt{2} S_{hv}S_{hh}^*\rangle & \langle 2|S_{hv}|^2\rangle & \langle\sqrt{2}S_{hv}S_{vv}^*\rangle \cr
# \langle S_{vv}S_{hh}^*\rangle & \langle\sqrt{2}S_{vv}S_{hv}^*\rangle & \langle |S_{vv}|^2\rangle },
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# ### Dual polarimetric imagery
#
# The Sentinel-1 sensors operate in reduced, power-saving polarization modes, emitting only one polarization and receiving two (dual polarization) or one (single polarization).
#
# For vertical transmission and horizontal and vertical reception,
#
# $$
# \bar{C} = \pmatrix{ \langle |S_{vv}|^2\rangle & \langle S_{vv}S_{vh}^*\rangle \cr
# \langle S_{vh}S_{vv}^*\rangle & \langle |S_{vh}|^2\rangle },
# $$
#
# The GEE archives only the diagonal (intensity) matrix elements, so we work in fact with
#
# $$
# \bar{C} = \pmatrix{ \langle |S_{vv}|^2\rangle & 0 \cr
# 0 & \langle |S_{vh}|^2\rangle },
# $$
# + [markdown] slideshow={"slide_type": "slide"}
#
# ### Change detection, bitemporal imagery
#
# The probability distribution of $\bar C$ is completely determined by the parameter $\Sigma$ (the covariance matrix) and by the __equivalent number of looks__ ENL.
#
# Given two measurements of polarized backscatter, one can set up an hypothesis test in order to decide whether or not a change has occurred.
#
# $$H_0: \Sigma_1 = \Sigma_2$$
#
# i.e., the two observations were sampled from the same distribution and no change has occurred
#
# $$H_1: \Sigma_1\ne\Sigma_2$$
#
# in other words, there was a change.
#
# Since the distributions are known, a test statistic can be formulated which allows one to decide to a desired degree of significance whether or not to reject the null hypothesis.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Change detection, multitemporal imagery
#
# In the case of $k > 2$ observations this procedure can be generalized to test a null hypothesis that all of the $k$ pixels are characterized by the same $\Sigma$, against the alternative that at least one of the $\Sigma_i$, $i=1\dots k$, are different, i.e., that at least one change has taken place.
#
#
# Furthermore this so-called __omnibus test procedure__ can be factored into a sequence of tests involving hypotheses of the form:
#
# $\Sigma_1 = \Sigma_2$ against $\Sigma_1 \ne \Sigma_2$,
#
# $\Sigma_1 = \Sigma_2 = \Sigma_3$ against $\Sigma_1 = \Sigma_2 \ne \Sigma_3$,
#
# and so forth.
# + [markdown] slideshow={"slide_type": "slide"}
# Denoting the test statistics $R^\ell_j,\ \ell = 1\dots k-1,\ j=\ell+1\dots k$, for a series of, say, $k=5$ images, we have the following tests to consider
#
# $$
# \matrix{
# \ell/j &2 &3 &4 &5\cr
# 1 & R^1_2 & R^1_3 & R^1_4 & R^1_5 \cr
# 2 & & R^2_3 & R^2_4 & R^2_5 \cr
# 3 & & & R^3_4 & R^3_5 \cr
# 4 & & & & R^4_5 }
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# ## The GEE interface
#
# The interface is programmed against the GEE Python API and uses Jupyter widgets to generate the desired Sentinel-1 times series for processing.
#
# Results (changes maps) can be previewed on-the-fly and then exported to the GEE Code Editor for visualization and animation.
# + slideshow={"slide_type": "slide"}
from auxil.eeSar_seq import run
run()
# -
| src/eesardocs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### In this notebook the following points has been discussed:
# <ol>
# <li>Some points releated to Skewness</li>
# <li>Is skewness bad</li>
# <li>How to remove skewness?</li>
# <li>How to calculate skewness in Pandas</li>
# </ol>
# ## 1. Some points related to Skewness
# <p>
# 1. When the value of the skewness is negative, the tail of the distribution is longer towards the left hand side of the curve.<Br>
# 2. When the value of the skewness is positive, the tail of the distribution is longer towards the right hand side of the curve.<br>
# 3. For a normal distribution the mean, median and mode coincide.<br>
# 4. In case of Positive skewness the <b> Mean >Median > Mode </b><br>
# 5. In case of Negative Skewness the <b> Mean < Median < Mode</b><br>
# 6. In positive skew most of the data is less than the mean<br>
# 7. In negative skew most of the data is greater than mean<br>
# <img src="https://miro.medium.com/max/600/1*nj-Ch3AUFmkd0JUSOW_bTQ.jpeg"/>
# </p>
# ## 2. Is skewness bad?
# <p>Many models assume normal distribution; i.e., data are symmetric about the mean. The normal distribution has a skewness of zero. But in reality, data points may not be perfectly symmetric. So, an understanding of the skewness of the dataset indicates whether deviations from the mean are going to be positive or negative.</p>
# ## 3. How to remove skewness
# <p>In order to remove skewness we apply transformations on to our data. One such transformation is the Box cox Transform</p>
# ## 4. How to calculate Skewness in Pandas
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
delivery=pd.read_csv('deliveries.csv')
delivery.head()
country=pd.read_csv('Life Expectancy Data.csv')
country.head()
# ### 4a. Positive Skew Example
# Career runs of all the batsman
runs=delivery.groupby('batsman')['batsman_runs'].sum()
print(runs.skew())
sns.kdeplot(runs)
#print(runs)
x=delivery['fielder'].value_counts()
x.skew()
sns.kdeplot(x)
# ### 4b. Negative Skew Example
print(country['Life expectancy '].skew())
sns.kdeplot(country['Life expectancy '])
# ### So, when is the skewness too much?
#
# <p>
# The rule of thumb seems to be:<br><br>
# 1. If the skewness is between -0.5 and 0.5, the data are fairly symmetrical.<br>
# 2. If the skewness is between -1 and -0.5(negatively skewed) or between 0.5 and 1(positively skewed), the data are moderately skewed.<br>
# 3. If the skewness is less than -1(negatively skewed) or greater than 1(positively skewed), the data are highly skewed.</p>
# <h1>About Me </h1>
#
# <center>
# <img src="https://media-exp1.licdn.com/dms/image/C5103AQEgE5y5PWLcbA/profile-displayphoto-shrink_800_800/0/1582881994900?e=1650499200&v=beta&t=<KEY>" style="width:80px; height:80px; border-radius:40px" alt="picture"/>
# <h3 style="margin-top:5px" ><center><NAME></center></h3>
# <p style="margin-top:-5px;" ><center><b>Hi!</b> I'm <NAME>, a guy who is truly in love with creating Ideas and developing Innovations with help of Technology. I'm looking for an opportunity to pursue a challenging career and be a part of progressive organization that gives a scope to enhance my knowledge and utilizing my skills to convert Ideas into innovations towards the growth of the organization.</center></p>
# <div style="margin-top:0px">
# <p><center><b>Connect with me.</b></center></p>
# <div style="display:inline-block;margin-right:5px">
# <a href="https://www.linkedin.com/in/iamshivprakash" style="text-decoration:none; margin-right:5px"><img src="https://cdn-icons-png.flaticon.com/512/174/174857.png" style="width:25px; height:25px"></a>
# </div>
# <div style="display:inline-block;margin-right:5px">
# <a href="https://www.github.com/iamshivprakash" style="text-decoration:none; margin-right:5px"><img src="https://cdn-icons-png.flaticon.com/512/733/733609.png" style="width:25px; height:25px"></a>
# </div>
# <div style="display:inline-block;margin-right:5px">
# <a href="https://www.instagram.com/shi_pra_" style="text-decoration:none; margin-right:5px"><img src="https://cdn-icons-png.flaticon.com/512/174/174855.png" style="width:25px; height:25px"></a>
# </div>
# <div style="display:inline-block;margin-right:5px">
# <a href="https://www.kaggle.com/shivprakash21" style="text-decoration:none; margin-right:5px"><img src="https://storage.googleapis.com/kaggle-avatars/images/default-thumb.png" style="width:30px; height:30px"></a>
# </div>
# </div>
# </center>
#
# <h2>Change Log</h2>
#
# <div>
# <table>
# <thead>
# <tr>
# <th>Sno</th>
# <th>Version</th>
# <th>Changed By</th>
# <th>Date(DD/MM/YYYY)</th>
# <th>Remarks</th>
# </tr>
# </thead>
# <tbody>
# <tr>
# <td>1</td>
# <td>1.0</td>
# <td><NAME></td>
# <td>05/01/2022</td>
# <td>Notebook Created</td>
# </tr>
# <tr>
# <td>2</td>
# <td>1.1</td>
# <td><NAME></td>
# <td>09/01/2022</td>
# <td>Adeed some of the parameters of <code>read_csv()</code> function</td>
# </tr>
# </tbody>
# </table>
# </div>
# <hr>
# <p><center>Last Modified by <b><NAME></b> on <i>9th January 2022</i></center></p>
#
| statistics/Measure of Asymmetry - Skewness.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
from pylab import *
# %matplotlib inline
# +
import sys
caffe_root = '../' # this file should be run from {caffe_root}/examples (otherwise change this line)
sys.path.insert(0, caffe_root + 'python')
import caffe
# -
# run scripts from caffe root
import os
os.chdir(caffe_root)
# Download data
# !data/mnist/get_mnist.sh
# Prepare data
# !examples/mnist/create_mnist.sh
# back to examples
os.chdir('examples')
# +
from caffe import layers as L, params as P
def lenet(lmdb, batch_size):
# our version of LeNet: a series of linear and simple nonlinear transformations
n = caffe.NetSpec()
n.data, n.label = L.Data(batch_size=batch_size, backend=P.Data.LMDB, source=lmdb,
transform_param=dict(scale=1./255), ntop=2)
n.conv1 = L.Convolution(n.data, kernel_size=5, num_output=20, weight_filler=dict(type='xavier'))
n.pool1 = L.Pooling(n.conv1, kernel_size=2, stride=2, pool=P.Pooling.MAX)
n.conv2 = L.Convolution(n.pool1, kernel_size=5, num_output=10, weight_filler=dict(type='xavier'))
n.loss = L.SoftmaxWithLoss(n.conv2, n.label)
return n.to_proto()
with open('mnist/lenet_auto_train.prototxt', 'w') as f:
f.write(str(lenet('mnist/mnist_train_lmdb', 64)))
with open('mnist/lenet_auto_test.prototxt', 'w') as f:
f.write(str(lenet('mnist/mnist_test_lmdb', 100)))
# -
def lenet_gen(lmdb, batch_size):
# our version of LeNet: a series of linear and simple nonlinear transformations
n = caffe.NetSpec()
n.data, n.label = L.Data(batch_size=batch_size, backend=P.Data.LMDB, source=lmdb,
transform_param=dict(scale=1./255), ntop=2)
n.conv1 = L.Convolution(n.data, kernel_size=5, num_output=20, weight_filler=dict(type='xavier'))
n.pool1 = L.Pooling(n.conv1, kernel_size=2, stride=2, pool=P.Pooling.MAX)
n.conv2 = L.Convolution(n.pool1, kernel_size=5, num_output=10, weight_filler=dict(type='xavier'))
n.loss = L.SoftmaxWithLoss(n.conv2, n.label)
return n
# !cat mnist/lenet_auto_train.prototxt
# !cat mnist/lenet_auto_solver.prototxt
# +
caffe.set_mode_cpu()
### load the solver and create train and test nets
solver = None # ignore this workaround for lmdb data (can't instantiate two solvers on the same data)
solver = caffe.SGDSolver('mnist/lenet_auto_solver.prototxt')
# -
| examples/Lenet without inner product.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="3E96e1UKQ8uR"
# # MoViNet Tutorial
#
# This notebook provides basic example code to create, build, and run [MoViNets (Mobile Video Networks)](https://arxiv.org/pdf/2103.11511.pdf). Models use TF Keras and support inference in TF 1 and TF 2. Pretrained models are provided by [TensorFlow Hub](https://tfhub.dev/google/collections/movinet/), trained on [Kinetics 600](https://deepmind.com/research/open-source/kinetics) for video action classification.
# + [markdown] id="8_oLnvJy7kz5"
# ## Setup
#
# It is recommended to run the models using GPUs or TPUs.
#
# To select a GPU/TPU in Colab, select `Runtime > Change runtime type > Hardware accelerator` dropdown in the top menu.
#
# ### Install the TensorFlow Model Garden pip package
#
# - tf-models-official is the stable Model Garden package. Note that it may not include the latest changes in the tensorflow_models github repo.
# - To include latest changes, you may install tf-models-nightly, which is the nightly Model Garden package created daily automatically.
# pip will install all models and dependencies automatically.
#
# Install the [mediapy](https://github.com/google/mediapy) package for visualizing images/videos.
# + id="s3khsunT7kWa"
# !pip install -q tf-models-nightly tfds-nightly
# !command -v ffmpeg >/dev/null || (apt update && apt install -y ffmpeg)
# !pip install -q mediapy
# + id="dI_1csl6Q-gH"
import os
from six.moves import urllib
import matplotlib.pyplot as plt
import mediapy as media
import numpy as np
from PIL import Image
import tensorflow as tf
import tensorflow_datasets as tfds
import tensorflow_hub as hub
from official.vision.beta.configs import video_classification
from official.projects.movinet.configs import movinet as movinet_configs
from official.projects.movinet.modeling import movinet
from official.projects.movinet.modeling import movinet_layers
from official.projects.movinet.modeling import movinet_model
# + [markdown] id="6g0tuFvf71S9"
# ## Example Usage with TensorFlow Hub
#
# Load MoViNet-A2-Base from TensorFlow Hub, as part of the [MoViNet collection](https://tfhub.dev/google/collections/movinet/).
#
# The following code will:
#
# - Load a MoViNet KerasLayer from [tfhub.dev](https://tfhub.dev).
# - Wrap the layer in a [Keras Model](https://www.tensorflow.org/api_docs/python/tf/keras/Model).
# - Load an example image, and reshape it to a single frame video.
# - Classify the video
# + id="nTUdhlRJzl2o"
movinet_a2_hub_url = 'https://tfhub.dev/tensorflow/movinet/a2/base/kinetics-600/classification/1'
inputs = tf.keras.layers.Input(
shape=[None, None, None, 3],
dtype=tf.float32)
encoder = hub.KerasLayer(movinet_a2_hub_url, trainable=True)
# Important: To use tf.nn.conv3d on CPU, we must compile with tf.function.
encoder.call = tf.function(encoder.call, experimental_compile=True)
# [batch_size, 600]
outputs = encoder(dict(image=inputs))
model = tf.keras.Model(inputs, outputs)
# + [markdown] id="7kU1_pL10l0B"
# To provide a simple example video for classification, we can load a static image and reshape it to produce a video with a single frame.
# + id="Iy0rKRrT723_"
image_url = 'https://upload.wikimedia.org/wikipedia/commons/8/84/Ski_Famille_-_Family_Ski_Holidays.jpg'
image_height = 224
image_width = 224
with urllib.request.urlopen(image_url) as f:
image = Image.open(f).resize((image_height, image_width))
video = tf.reshape(np.array(image), [1, 1, image_height, image_width, 3])
video = tf.cast(video, tf.float32) / 255.
image
# + [markdown] id="Yf6EefHuWfxC"
# Run the model and output the predicted label. Expected output should be skiing (labels 464-467). E.g., 465 = "skiing crosscountry".
#
# See [here](https://gist.github.com/willprice/f19da185c9c5f32847134b87c1960769#file-kinetics_600_labels-csv) for a full list of all labels.
# + id="OOpEKuqH8sH7"
output = model(video)
output_label_index = tf.argmax(output, -1)[0].numpy()
print(output_label_index)
# + [markdown] id="_s-7bEoa3f8g"
# ## Example Usage with the TensorFlow Model Garden
#
# Fine-tune MoViNet-A0-Base on [UCF-101](https://www.crcv.ucf.edu/research/data-sets/ucf101/).
#
# The following code will:
#
# - Load the UCF-101 dataset with [TensorFlow Datasets](https://www.tensorflow.org/datasets/catalog/ucf101).
# - Create a [`tf.data.Dataset`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset) pipeline for training and evaluation.
# - Display some example videos from the dataset.
# - Build a MoViNet model and load pretrained weights.
# - Fine-tune the final classifier layers on UCF-101.
# + [markdown] id="o7unW4WVr580"
# ### Load the UCF-101 Dataset with TensorFlow Datasets
#
# Calling `download_and_prepare()` will automatically download the dataset. After downloading, this cell will output information about the dataset.
# + id="FxM1vNYp_YAM"
dataset_name = 'ucf101'
builder = tfds.builder(dataset_name)
config = tfds.download.DownloadConfig(verify_ssl=False)
builder.download_and_prepare(download_config=config)
# + executionInfo={"elapsed": 2957, "status": "ok", "timestamp": 1619748263684, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 360} id="boQHbcfDhXpJ" outputId="eabc3307-d6bf-4f29-cc5a-c8dc6360701b"
num_classes = builder.info.features['label'].num_classes
num_examples = {
name: split.num_examples
for name, split in builder.info.splits.items()
}
print('Number of classes:', num_classes)
print('Number of examples for train:', num_examples['train'])
print('Number of examples for test:', num_examples['test'])
print()
builder.info
# + [markdown] id="BsJJgnBBqDKZ"
# Build the training and evaluation datasets.
# + id="9cO_BCu9le3r"
batch_size = 8
num_frames = 8
frame_stride = 10
resolution = 172
def format_features(features):
video = features['video']
video = video[:, ::frame_stride]
video = video[:, :num_frames]
video = tf.reshape(video, [-1, video.shape[2], video.shape[3], 3])
video = tf.image.resize(video, (resolution, resolution))
video = tf.reshape(video, [-1, num_frames, resolution, resolution, 3])
video = tf.cast(video, tf.float32) / 255.
label = tf.one_hot(features['label'], num_classes)
return (video, label)
train_dataset = builder.as_dataset(
split='train',
batch_size=batch_size,
shuffle_files=True)
train_dataset = train_dataset.map(
format_features,
num_parallel_calls=tf.data.AUTOTUNE)
train_dataset = train_dataset.repeat()
train_dataset = train_dataset.prefetch(2)
test_dataset = builder.as_dataset(
split='test',
batch_size=batch_size)
test_dataset = test_dataset.map(
format_features,
num_parallel_calls=tf.data.AUTOTUNE,
deterministic=True)
test_dataset = test_dataset.prefetch(2)
# + [markdown] id="rToX7_Ymgh57"
# Display some example videos from the dataset.
# + id="KG8Z7rUj06of"
videos, labels = next(iter(train_dataset))
media.show_videos(videos.numpy(), codec='gif', fps=5)
# + [markdown] id="R3RHeuHdsd_3"
# ### Build MoViNet-A0-Base and Load Pretrained Weights
# + [markdown] id="JXVQOP9Rqk0I"
# Here we create a MoViNet model using the open source code provided in [tensorflow/models](https://github.com/tensorflow/models) and load the pretrained weights. Here we freeze the all layers except the final classifier head to speed up fine-tuning.
# + id="JpfxpeGSsbzJ"
model_id = 'a0'
tf.keras.backend.clear_session()
backbone = movinet.Movinet(
model_id=model_id)
model = movinet_model.MovinetClassifier(
backbone=backbone,
num_classes=600)
model.build([batch_size, num_frames, resolution, resolution, 3])
# Load pretrained weights from TF Hub
movinet_hub_url = f'https://tfhub.dev/tensorflow/movinet/{model_id}/base/kinetics-600/classification/1'
movinet_hub_model = hub.KerasLayer(movinet_hub_url, trainable=True)
pretrained_weights = {w.name: w for w in movinet_hub_model.weights}
model_weights = {w.name: w for w in model.weights}
for name in pretrained_weights:
model_weights[name].assign(pretrained_weights[name])
# Wrap the backbone with a new classifier to create a new classifier head
# with num_classes outputs
model = movinet_model.MovinetClassifier(
backbone=backbone,
num_classes=num_classes)
model.build([batch_size, num_frames, resolution, resolution, 3])
# Freeze all layers except for the final classifier head
for layer in model.layers[:-1]:
layer.trainable = False
model.layers[-1].trainable = True
# + [markdown] id="ucntdu2xqgXB"
# Configure fine-tuning with training/evaluation steps, loss object, metrics, learning rate, optimizer, and callbacks.
#
# Here we use 3 epochs. Training for more epochs should improve accuracy.
# + id="WUYTw48BouTu"
num_epochs = 3
train_steps = num_examples['train'] // batch_size
total_train_steps = train_steps * num_epochs
test_steps = num_examples['test'] // batch_size
loss_obj = tf.keras.losses.CategoricalCrossentropy(
from_logits=True,
label_smoothing=0.1)
metrics = [
tf.keras.metrics.TopKCategoricalAccuracy(
k=1, name='top_1', dtype=tf.float32),
tf.keras.metrics.TopKCategoricalAccuracy(
k=5, name='top_5', dtype=tf.float32),
]
initial_learning_rate = 0.01
learning_rate = tf.keras.optimizers.schedules.CosineDecay(
initial_learning_rate, decay_steps=total_train_steps,
)
optimizer = tf.keras.optimizers.RMSprop(
learning_rate, rho=0.9, momentum=0.9, epsilon=1.0, clipnorm=1.0)
model.compile(loss=loss_obj, optimizer=optimizer, metrics=metrics)
callbacks = [
tf.keras.callbacks.TensorBoard(),
]
# + [markdown] id="0IyAOOlcpHna"
# Run the fine-tuning with Keras compile/fit. After fine-tuning the model, we should be able to achieve >70% accuracy on the test set.
# + executionInfo={"elapsed": 982253, "status": "ok", "timestamp": 1619750139919, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 360} id="Zecc_K3lga8I" outputId="e4c5c61e-aa08-47db-c04c-42dea3efb545"
results = model.fit(
train_dataset,
validation_data=test_dataset,
epochs=num_epochs,
steps_per_epoch=train_steps,
validation_steps=test_steps,
callbacks=callbacks,
validation_freq=1,
verbose=1)
# + [markdown] id="XuH8XflmpU9d"
# We can also view the training and evaluation progress in TensorBoard.
# + id="9fZhzhRJRd2J"
# %reload_ext tensorboard
# %tensorboard --logdir logs --port 0
| official/projects/movinet/movinet_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **This is a preliminary ML script to code Naive Bayes and Logistic Regression models for our project.**
#
# Naive Bayes Overview:
# "...if we make very naive assumptions about the generative model for each label, we can find a rough approximation of the generative model for each class, and then proceed with the Bayesian classification. Different types of naive Bayes classifiers rest on different naive assumptions about the data..." (<NAME>. Python Data Science Handbook. O'Reilly Media, Inc.: 2016.
#
# Description of Target(replace with .gov source in final notebook, this is enough to get started though):
# https://regulatorysol.com/action-taken-action-taken-date/
# +
# %matplotlib inline
import os
import json
import time
import pickle
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns; sns.set()
# -
#delete this in a future version of the notebook
root = ''
# **Import 2017 sample of 25,000 observations.** Note import warning:"Columns (29,30,39,40) have mixed types. Specify dtype option on import or set low_memory=False."
# Fetch the data if required
filepath = os.path.abspath(os.path.join( "..", "fixtures", "hmda2017sample.csv"))
DATA = pd.read_csv(filepath)
DATA.describe(include='all')
# **Write the initial script using subset of features which are already int or float, plus the target** Future version of script will address full set of features, and will move away from use of the lambda function for readability.
DATA['action_taken'] = DATA.action_taken_name.apply(lambda x: 1 if x in ['Loan purchased by the institution', 'Loan originated'] else 0)
pd.crosstab(DATA['action_taken_name'],DATA['action_taken'], margins=True)
DATA = DATA[['tract_to_msamd_income',
'population',
'minority_population',
'number_of_owner_occupied_units',
'number_of_1_to_4_family_units',
'loan_amount_000s',
'hud_median_family_income',
'applicant_income_000s',
'action_taken']]
DATA.info()
#TO DO: fix column [0]
tofilepath = os.path.abspath(os.path.join( "..", "fixtures", "hmda2017sample_test.csv"))
DATA.to_csv(tofilepath)
# +
FEATURES = [
'tract_to_msamd_income',
'population',
'minority_population',
'number_of_owner_occupied_units',
'number_of_1_to_4_family_units',
'loan_amount_000s',
'hud_median_family_income',
'applicant_income_000s',
'action_taken'
]
ACTION_TAKEN_MAP = {
1: "originated or purchased",
0: "other"
}
# +
# Determine the shape of the data
print("{} instances with {} features\n".format(*DATA.shape))
# Determine the frequency of each class
print(pd.crosstab(index=DATA['action_taken'], columns="count"))
# -
# **Stage the data for ML algorithms.** Need to determine whether we can keep y as binary or if it in fact has to be labeled for Scikit-Learn, Yellowbrick et al to work.
# +
from sklearn.preprocessing import LabelEncoder
# Extract our X and y data
X = DATA[FEATURES[:-1]]
y = DATA['action_taken']
# Encode our target variable
encoder = LabelEncoder().fit(y)
y = encoder.transform(y)
print(X.shape, y.shape)
# -
# Create a scatter matrix of the dataframe features
from pandas.plotting import scatter_matrix
scatter_matrix(X, alpha=0.2, figsize=(12, 12), diagonal='kde')
plt.show()
# ## Data Extraction
#
# One way that we can structure our data for easy management is to save files on disk. The Scikit-Learn datasets are already structured this way, and when loaded into a `Bunch` (a class imported from the `datasets` module of Scikit-Learn) we can expose a data API that is very familiar to how we've trained on our toy datasets in the past. A `Bunch` object exposes some important properties:
#
# - **data**: array of shape `n_samples` * `n_features`
# - **target**: array of length `n_samples`
# - **feature_names**: names of the features
# - **target_names**: names of the targets
# - **filenames**: names of the files that were loaded
# - **DESCR**: contents of the readme
#
# **Note**: This does not preclude database storage of the data, in fact - a database can be easily extended to load the same `Bunch` API. Simply store the README and features in a dataset description table and load it from there. The filenames property will be redundant, but you could store a SQL statement that shows the data load.
#
# In order to manage our data set _on disk_, we'll structure our data as follows:
from sklearn.datasets.base import Bunch
# **NOTE: np.genfromtxt and objects for extracting the target from the data have syntactical issues**
# +
def load_data(root=root):
# Construct the `Bunch` for the HMDA dataset
filenames = {
'meta': os.path.join(root, 'fixtures','hmdameta.json'),
'rdme': os.path.join(root, 'fixtures','hmdareadme.txt'),
'data': os.path.join(root, 'fixtures','hmda2017sample_test.csv'),
}
# Load the meta data from the meta json
with open(filenames['meta'], 'r') as f:
meta = json.load(f)
target_names = meta['target_names']
feature_names = meta['feature_names']
# Load the description from the README.
with open(filenames['rdme'], 'r') as f:
DESCR = f.read()
# Load the dataset from the text file.
dataset = np.genfromtxt(filenames['data'], delimiter = ",", names = True)
# Extract the target from the data
data = dataset[:, 0:7]
target = dataset[:, -1]
# Create the bunch object
return Bunch(
data=data,
target=target,
filenames=filenames,
target_names=target_names,
feature_names=feature_names,
DESCR=DESCR
)
# Save the dataset as a variable we can use.
dataset = load_data()
print(dataset.data.shape)
print(dataset.target.shape)
# -
# ## Classification
#
# Now that we have a dataset `Bunch` loaded and ready, we can begin the classification process. Let's attempt to build a classifier with kNN, SVM, and Random Forest classifiers.
# +
from sklearn import metrics
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import OneHotEncoder
from sklearn.model_selection import KFold
from sklearn.naive_bayes import GaussianNB
from sklearn.linear_model import LogisticRegression
# -
def fit_and_evaluate(dataset, model, label, **kwargs):
"""
Because of the Scikit-Learn API, we can create a function to
do all of the fit and evaluate work on our behalf!
"""
start = time.time() # Start the clock!
scores = {'precision':[], 'recall':[], 'accuracy':[], 'f1':[]}
kf = KFold(n_splits = 12, shuffle=True)
for train, test in kf.split(dataset.data):
X_train, X_test = dataset.data[train], dataset.data[test]
y_train, y_test = dataset.target[train], dataset.target[test]
estimator = model(**kwargs)
estimator.fit(X_train, y_train)
expected = y_test
predicted = estimator.predict(X_test)
# Append our scores to the tracker
scores['precision'].append(metrics.precision_score(expected, predicted, average="weighted"))
scores['recall'].append(metrics.recall_score(expected, predicted, average="weighted"))
scores['accuracy'].append(metrics.accuracy_score(expected, predicted))
scores['f1'].append(metrics.f1_score(expected, predicted, average="weighted"))
# Report
print("Build and Validation of {} took {:0.3f} seconds".format(label, time.time()-start))
print("Validation scores are as follows:\n")
print(pd.DataFrame(scores).mean())
# Write official estimator to disk
estimator = model(**kwargs)
estimator.fit(dataset.data, dataset.target)
outpath = label.lower().replace(" ", "-") + ".pickle"
with open(outpath, 'wb') as f:
pickle.dump(estimator, f)
print("\nFitted model written to:\n{}".format(os.path.abspath(outpath)))
# Perform Gaussian Naive Bayes
# need to try this out and extend to MultinomialNB, introducing Pipeline
fit_and_evaluate(dataset, GaussianNB, "Gaussian Naive Bayes",)
# Perform Logistic Regression
fit_and_evaluate(dataset, LogisticRegression, "Logistic Regression", )
| notebooks/01e_InitialMLTemplate_ak.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SELECTORES: XPATH Y CSS
# Cuando se requiere realizar la extracción de datos web, la tarea más común es la extracción de esta data a partir del árbol HTML de la página web. Para poder realizar ello se requieren de los <b>selectores, los cuales se encargarán de obtener las partes del documento HTML que querramos recuperar</b>
# ## 1. XPATH SELECTOR
# XPath es un lenguaje de <b>selección de nodos en documentos XML</b> (Lenguaje de Marcas Extensible), los cuales son usados también con documentos HTML
#
# #### Utilizando selectores XPATH
from scrapy.selector import Selector
html_doc='''
<html>
<head>
<base href='http://example.com/' />
<title>Example website</title>
</head>
<body>
<div id='images'>
<a href='image1.html'>Name: My image 1 <br /><img src='image1_thumb.jpg' /></a>
<a href='image2.html'>Name: My image 2 <br /><img src='image2_thumb.jpg' /></a>
<a href='image3.html'>Name: My image 3 <br /><img src='image3_thumb.jpg' /></a>
<a href='image4.html'>Name: My image 4 <br /><img src='image4_thumb.jpg' /></a>
<a href='image5.html'>Name: My image 5 <br /><img src='image5_thumb.jpg' /></a>
</div>
</body>
</html>
'''
#pasando el codigo html de la página
response=Selector(text=html_doc)
#obteniendo el título de la página web
response.xpath('//head/title/text()').get()
#obteniendo de los elementos a
response.xpath('//div/a/text()').getall()
#obteniendo de los elementos href dentro de a
response.xpath('//div/a/@href').getall()
# ## 2. CSS SELECTOR
# CSS (Cascading Style Sheets) es un <b>lenguaje para describir la representación de documentos HTML y XML </b>en pantalla, papel, voz, etc. CSS utiliza selectores para vincular propiedades de estilo a elementos en el documento web. En líneas generales la selección de elementos CSS se realiza a partir de la selección de los atributos contenidos en el documento web.
# #### Utilizando selectores CSS
from scrapy.selector import Selector
#página web de prueba
html_doc='''
<html>
<head>
<base href='http://example.com/' />
<title>Example website</title>
</head>
<body>
<div id='images'>
<a href='image1.html'>Name: My image 1 <br /><img src='image1_thumb.jpg' /></a>
<a href='image2.html'>Name: My image 2 <br /><img src='image2_thumb.jpg' /></a>
<a href='image3.html'>Name: My image 3 <br /><img src='image3_thumb.jpg' /></a>
<a href='image4.html'>Name: My image 4 <br /><img src='image4_thumb.jpg' /></a>
<a href='image5.html'>Name: My image 5 <br /><img src='image5_thumb.jpg' /></a>
</div>
</body>
</html>
'''
response=Selector(text=html_doc)
response.css('title::text').get()
response.css('div#images > a::text').getall()
response.css('img::attr(src)').getall()
response.css('div#images > a::attr(href)').getall()
| Modulo1/.ipynb_checkpoints/2.SELECTORES_XPATH_CSS-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### Review: 2D arrays
#
# * boolean indexing
# * other ways to create 2d array
# +
ages = np.array([22, 10, 34, 71, 14, 50, 61, 15]) # age in years
heights = np.array([1.7, 1.0, 1.8, 1.9, 0.9, 1.5, 1.5, 1.0]) # height in m
# How can we get:
# the average height of people older than 20 ?
greater_than_20 = ages > 20
# this works, by "filtering out" the people younger than 20, and leaving
# zeros in their place
# but taking the average of these wouldn't work, because the zeros would mess it up
filtered = heights * greater_than_20
# to get the average from this:
number_of_people_older_than_20 = np.sum(greater_than_20)
mean_height_of_people_over_20 = np.sum(filtered) / number_of_people_older_than_20
# -
print(greater_than_20)
# +
# how can we ignore the zeros in filtered array?
ages = np.array([22, 10, 34, 71, 14, 50, 61, 15]) # age in years
heights = np.array([1.7, 1.0, 1.8, 1.9, 0.9, 1.5, 1.5, 1.0]) # height in m
greater_than_20 = ages > 20
filtered = heights * greater_than_20
for i in range(len(filtered)):
if filtered[i] == 0:
filtered[i] = None
print(filtered)
# nan : "not a number"
print(np.mean(filtered))
# to get around that:
print(np.nanmean(filtered))
# +
# an easier approach:
ages = np.array([22, 10, 34, 71, 14, 50, 61, 15]) # age in years
heights = np.array([1.7, 1.0, 1.8, 1.9, 0.9, 1.5, 1.5, 1.0]) # height in m
greater_than_20 = ages > 20
heights[greater_than_20]
# normally we would index with numbers inside
# here are doing something new: indexing with an array of True/False's
# +
bools = np.array([True, True, False])
heights[bools]
# +
heights[ages>20] # the "ages>20" part of this creates a boolean array
# we then use that boolean array to index the elements of "heights"
heights_over_1point5 = heights[heights > 1.5]
print(heights)
print(heights_over_1point5)
# +
# how about 2d arrays and boolean indexing?
arr = np.array([[1,2,3],[4,5,6],[7,8,9],[10,11,12]])
# how can we select the 0th, 1st, and 3rd rows at once
indexer = np.array([True, True, False, True])
arr[indexer]
arr[indexer,:] # colon means all items in that axis
#print(arr[0])
print(arr[indexer])
# +
# can we nest a boolean index inside our regular number-based indexing?
# for example, can we index the first row, only where it is greater than 4?
print(arr[1, arr[1]>4])
# but you would normally do this in 2 steps to be clearer:
row1 = arr[1]
#print(row1)
answer = row1[row1>4]
print(answer)
# -
# we didn't talk about this approach explicitly
arr = np.arange(200).reshape([20,10])
print(arr)
# +
arr[arr>84]
# notice that this resulted in a 1D array of all the values over 84
# which makes sense because 85 was in the middle of a row,
# so we (and numpy) would not know how to perserve the 2D nature of it
#
# because this scenario can happen, numpy will always flatten
# the result when indexing like this
# -
arr[arr>=100]
print(arr)
# +
x = np.array([1,2,3,4,5])
x[2] = 0
print(x)
# +
arr[arr>=100] = 0
print(arr)
# +
arr[arr<50] = arr[arr<50] * 2
print(arr)
# -
# <hr/>
# ### matplotlib
# +
# if any of this is confusing, just ignore it for now:
import matplotlib # not needed for the line below with pyplot to work
# this is not a true python command and is just telling python
# how to show our graphs/plots when we make them today
# %matplotlib tk
##
# this is the important line for giving us the module and code we need today
import matplotlib.pyplot as pl
# a lot of people use `plt` instead of `pl`
# so you might see that in online help
# +
# subplots
# this is new: when a function returns 2 variables
# you can collect them like this
fig,ax = pl.subplots()
# +
# first plot, interactive
x = [0,1,2]
y = [1,3,2]
ax.plot(x, y)
# -
ax.plot(x, y, linewidth=3, color='red')
# +
ax.set_yticks([0, 2, 4])
ax.set_ylabel('My y axis here', labelpad=30, fontsize=40)
# labelpad determine how far from the axis the label shows up
# +
x = [0,1,2]
y = [3,1,4]
ax.plot(x, y, linewidth=3)
# +
# if we want another new figure to work on
fig2,ax2 = pl.subplots()
# to plot on that one, use ax2.plot...
# +
# line
# +
# lims, ticks, labels
ax.set_ylim([1,2])
# +
# scatter
# +
# histogram
# +
# errorbar
# +
# bar
# +
# matplotlib gallery
# +
# imshow
# -
# +
lines = ax.plot([1,3,2])
print(lines)
line = lines[0]
line.set_linewidth(100)
# +
# leaving this here to ponder
# we did not teach it
my_settings = dict(linewidth=5, color='cornflowerblue')
ax.plot(x, y, **my_settings)
# -
# +
img = pl.imread('/Users/ben/Desktop/cat.jpg')
np.shape(img)
#img.shape # same thing as np.shape(img)
fig,ax = pl.subplots()
ax.imshow(img)
# -
# <hr/>
# ## Questions, review, and exercises
# <hr/>
# #### Problem 1
# __(a)__ Load in the image called `'sunset.jpg'` using the `pl.imread` function.
sunset = pl.imread('sunset.jpg')
# __(b)__ Inspect the shape of the image data. What does each axis of the array correspond to?
# __(c)__ Use `matplotlib` to display the image on some axes in a figure.
sunset.shape
# +
fig,ax = pl.subplots()
sunset_upsidedown = sunset[::-1]
sunset_flipped = sunset[:, ::-1]
ax.imshow(sunset_flipped)
# -
# __(d)__ Which is brighter on average: the top half or bottom half of the image? Answer this quantitatively using `numpy`.
#
# *Hint: in most images, small values correspond to darkness, and large values correspond to brightness.*
# +
fig,ax = pl.subplots()
ax.imshow(sunset, cmap='gist_gray')
# -
# * y axis ?
# * flip
# * black and white ?
# +
# pl.imshow?
# -
# <hr/>
# #### Problem 2
# __(a)__ Load in the dataset called `neural_data.npy`.
data = np.load('neural_data.npy')
# __(b)__ These data come from some neurons recorded in the lab. Every neuron was recorded for 30 seconds.
#
# The values represent the instantaneous firing rate of the recorded neurons, in units of Hz. (You can think of this as a measure of how active the neuron is.)
#
# Each row of the array contains the data from one recorded neuron.
#
# Each column contains a time point from the 30 seconds of the recording.
#
# How many neurons did we record?
# +
n_neurons = data.shape[0]
print(n_neurons)
# -
# __(c)__ Given that each recording was exactly 30 seconds long, what is the sampling rate of the recording? In other words, how many samples did we record in each second?
# +
data.shape
n_samples = data.shape[1]
duration = 30 # seconds
samples_per_second = n_samples / duration
print(samples_per_second) # aka sampling rate, `fs`
# units = Hz = samples per second
# -
# __(d)__ Plot the firing rate of each neuron, with each neuron being represented by a single trace with a different color.
data.shape
# +
fig,ax = pl.subplots()
n_datapoints = data.shape[1]
time = np.linspace(0,30,n_datapoints)
# we know that our data consist of 30 seconds of equally spaced
# samples, and there are n_datapoints samples
# using a for loop on a 2d array
# what do I get in each iteration?
for neuron in data:
ax.plot(time, neuron)
# answer: each item in a 2d array is a row
# for example, my_array[2] is the third row
# therefore, each iteration of this for loop
# gives us one row of the array
# which in this case, corresponds to the data from one neuron
ax.set_xlabel('Time (s)', fontsize='large')
ax.set_ylabel('Instantaneous firing rate (Hz)', fontsize='large')
# +
my_list = [1,2,3,4,5]
for item in my_list:
print(item)
# +
my_list = [1,2,3,4,5]
i = 0
while i<len(my_list):
print(my_list[i])
i = i + 1
# -
ax.plot
# __(e)__ Plot the average firing rate across neurons.
#
# (This should be a 30-second-long trace representing the average firing rate at each time, where you averaged over all the neurons.)
# +
fig,ax = pl.subplots()
average_over_neurons = np.mean(data, axis=0)
ax.plot(average_over_neurons)
# -
# __(f)__ At some unknown point, a stimulus was applied to the neurons. Looking at your plot, can you guess when that was?
#
# __(g)__ Compute the highest firing rate observed for each neuron.
# +
highest_rate_per_neuron = np.max(data, axis=1)
print(highest_rate_per_neuron)
# -
# Does your answer match what you observe in your plot?
# __(h)__ For each neuron, determine the time at which the highest firing rate was observed.
# +
index_of_max_value = np.argmax(data, axis=1)
print(index_of_max_value)
time_of_max_value = index_of_max_value / 100
print(time_of_max_value)
# or, another approach:
time = np.linspace(0,30,n_datapoints)
first_neuron_index_max = index_of_max_value[0]
time_of_max_value = time[index_of_max_value]
print(time_of_max_value)
# +
index_of_max_value = np.argmax(data, axis=1)
print(index_of_max_value)
time = np.linspace(0,30,n_datapoints)
time_of_max_value = time[index_of_max_value]
print(time_of_max_value)
# +
# types of indexing in an array
my_array = np.array(['a','b','c','d','e','f','g','h','i'])
# using a single integer
print(my_array[3])
# using a slice
print(my_array[3:7])
# using boolean
indexers = np.array([True,True,True,False,False,False,True,False,True])
print(my_array[indexers])
# using an array of integers
indexers = np.array([0,5,6,7,8])
print(my_array[indexers])
# -
# # loading excel files
# +
import pandas as pd
my_data = pd.read_excel('/Users/ben/Desktop/my_data.xlsx', header=2)
# -
print(my_data)
type(my_data)
# +
# this converts the fancy pandas data structure
# to a numpy array of data
my_new_data = my_data.values
# -
type(my_new_data)
#print(my_new_data)
print(my_new_data.shape)
csv_data = pd.read_csv('/Users/ben/Desktop/multiTimeline.csv')
my_data = csv_data.values
print(my_data.shape)
# +
fig,ax = pl.subplots()
ax.plot(my_new_data[:,3])
# -
my_new_data[:,2]
# +
for i in np.arange(1000):
print(i)
# +
# an example of how one might make a progress bar
import time
for i in range(100):
print('|'+'#'*i+' '*(100-i-1)+f'|{i+1}%', end='')
time.sleep(0.05)
print('\r', end='')
# -
| Week1_Python/Day5/Week1Day5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
import cv2
import numpy as np
from skimage.morphology import skeletonize, remove_small_objects
#cropping the image, h-77 subtracts the title at the bottom
original_img = cv2.imread('giraffe.png',0)
(h,w)=np.shape(original_img)
img = original_img[0:h, 0:w]
plt.imshow(img, cmap = 'gray')
#plt.xticks([]), plt.yticks([]) # to hide tick values on X and Y axis
# -
#plotting histogram
plt.hist(img.ravel(),256,[0,256])
plt.show()
#equalizing the histogram
equ = cv2.equalizeHist(img)
plt.hist(equ.ravel(),256,[0,256])
plt.show()
plt.imshow(equ, cmap='gray')
plt.xticks([]), plt.yticks([])
#blurring
ablur = cv2.blur(equ,(10, 10))
plt.imshow(ablur, cmap='gray', interpolation='sinc')
plt.xticks([]), plt.yticks([])
#blurring original
oblur = cv2.blur(img,(10, 10))
plt.imshow(oblur, cmap='gray', interpolation='sinc')
plt.xticks([]), plt.yticks([])
a_mthres = cv2.adaptiveThreshold(ablur,1,cv2.ADAPTIVE_THRESH_MEAN_C,\
cv2.THRESH_BINARY,41,4)
plt.imshow(a_mthres, interpolation= 'sinc')
o_mthres = cv2.adaptiveThreshold(oblur,1,cv2.ADAPTIVE_THRESH_MEAN_C,\
cv2.THRESH_BINARY,41,4)
plt.imshow(o_mthres, interpolation= 'sinc')
#This section blurs, thresholds, and skeletonizes the image
#It uses an average blur and an adaptive (local) mean threshold
#Remove_small_objects gets rid of any unconnected dots or lines
a_mthres = cv2.adaptiveThreshold(ablur,1,cv2.ADAPTIVE_THRESH_MEAN_C,\
cv2.THRESH_BINARY,41,4)
adaptiveskeleton = skeletonize(a_mthres)
plt.imshow(adaptiveskeleton,interpolation='sinc')
plt.xticks([]), plt.yticks([])
#without histogram equalization
o_mthres = cv2.adaptiveThreshold(oblur,1,cv2.ADAPTIVE_THRESH_MEAN_C,\
cv2.THRESH_BINARY,41,4)
oskel = skeletonize(o_mthres)
plt.imshow(oskel,interpolation='sinc')
plt.xticks([]), plt.yticks([])
difference=oskel-adaptiveskeleton
plt.imshow(difference, interpolation='sinc')
#just thresholding the original image
a_mthres = cv2.adaptiveThreshold(ablur,1,cv2.ADAPTIVE_THRESH_MEAN_C,\
cv2.THRESH_BINARY,41,4)
# +
adaptiveskeleton = skeletonize(a_mthres)
plt.imshow(adaptiveskeleton,interpolation='sinc')
plt.xticks([]), plt.yticks([])
# -
#cleans the skeleton by removing unnecessary unconnected dots/lines
cleanskel=remove_small_objects(adaptiveskeleton, min_size=70, connectivity=3)
plt.imshow(cleanskel,interpolation='sinc')
plt.xticks([]), plt.yticks([])
# +
#the original image values (integers) become floats
#Dividing by 255 normalizes the floats; now they are all between 0 and 1
floatimg=img.astype(float)
normimg=floatimg/255
#skeleton is a float, rgbpic is a RGB image with the dimensions of the cropped diatom image
#the values of R,G, and B are defined later
floatskel=cleanskel.astype(float)
(x,y)=np.shape(img)
rgbpic=np.zeros((x,y,3))
# -
# +
#dilating the lines of the skeleton to a 3x3 square of pixels (lines are bolder)
kernel=np.ones((1,1))
skel_dilation=cv2.dilate(floatskel,kernel,iterations=1)
#defining r, g, and b values for the overlay
#if the skeleton is 1, then it plots the skeleton.
#if the skeleton is 0, it plots the grayscale image of the diatom
rgbpic[:,:,0]=skel_dilation*(1-normimg)+normimg
rgbpic[:,:,1]=(1-skel_dilation)*normimg
rgbpic[:,:,2]=(1-skel_dilation)*normimg
#plotting the overlay
plt.imshow(rgbpic, interpolation='sinc')
plt.xticks([]), plt.yticks([])
# -
# this section finds the number of neighbors to each point in the skeleton
# line ends are points that have only one neighbor
# we want to get rid of these
# keep looping over this section until the number of line ends equals zero
nLineEnds = 1
while (nLineEnds>0):
#blur the skeleton with 3x3; this gives the average value in a 3x3 portion of the skeleton
# 9 times the average - 1 should be the number of neighbors
skelblur = cv2.blur(floatskel,(3,3))
neighbors=((9*skelblur)-1)
#making sure edges always have values
#any skeleton pixels within 'a' pixels from the edge will not be eroded
edges=np.zeros((x,y))
a=15
edges[0:a,:]=np.ones((a,y))
edges[x-a:x,:]=np.ones((a,y))
edges[:,0:a]=np.ones((x,a))
edges[:,y-a:y]=np.ones((x,a))
#removing the end pixel if a skeleton pixel has 1 neighbor
neighbors=(neighbors+5*edges)*floatskel
line_ends = (neighbors==1)
float_line_ends = line_ends.astype(float)
floatskel=floatskel-line_ends
nLineEnds=np.sum(line_ends)
#plt.imshow(neighbors, interpolation='sinc')
| work/CNMS/giraffe for figure 7 and 18 overlay.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Preprocessing
# import relevant statistical packages
import numpy as np
import pandas as pd
from sklearn.linear_model import LinearRegression
# import relevant data visualisation packages
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# ### 11.a. Generating a response $Y$ with two predictors $X1$ and $X2$, with $n$=100
X1 = np.random.normal(size=100)
X2 = np.random.normal(size=100)
epsilon = np.random.normal(scale=2.5, size=100)
Y = -12.5 + 35.7*X1 - 23.4*X2 + epsilon
# ### 11.b. Generating $\hat{\beta}$
beta1 = 25.7
# ### 11.c. Keeping $\hat{\beta}$ and fitting the model, $Y - \hat{\beta_1}X_1 = \beta_0 + \beta_2X_2 + \epsilon$
a2 = Y - beta1*X1
lm2 = LinearRegression(fit_intercept=True).fit(X2.reshape(-1, 1), a2)
beta2 = lm2.coef_[0]
# ### 11.d. Keeping $\hat{\beta_2}$ fixed and fitting the model, $Y - \hat{\beta_2}X_2 = \beta_0 + \beta_1X_1 + \epsilon$
a1 = Y - beta2*X2
lm1 = LinearRegression(fit_intercept=True).fit(X1.reshape(-1, 1), a1)
beta1 = lm1.coef_[0]
lm1.intercept_, lm2.intercept_
beta1, beta2
# ### 11.d. Estimating $\hat{\beta_0}$, $\hat{\beta_1}$ and $\hat{\beta_2}$
n = 1000
beta0 = np.zeros(n)
beta1 = np.zeros(n)
beta2 = np.zeros(n)
beta1[0] = 25.7
for k in range(0, n):
a2 = Y - beta1[k]*X1
lm2 = LinearRegression(fit_intercept=True).fit(X2.reshape(-1, 1), a2)
beta2[k] = lm2.coef_[0]
a1 = Y - beta2[k]*X2
lm1 = LinearRegression(fit_intercept=True).fit(X1.reshape(-1, 1), a1)
if k<n-1:
beta1[k+1] = lm1.coef_[0]
beta0[k] = lm1.intercept_
betadf = pd.concat([pd.DataFrame(beta0), pd.DataFrame(beta1), pd.DataFrame(beta2)], axis=1)
betadf.columns = ['beta0', 'beta1', 'beta2']
betadf.head()
plt.xkcd()
plt.figure(figsize=(25, 10))
plt.plot(betadf)
# **The coefficients attaing their least square values quite quickly.**
# ### 11.f. Performing multiple linear regression tp predict $Y$ using $X1$ and $X2$
df = pd.concat([pd.DataFrame(Y), pd.DataFrame(X1), pd.DataFrame(X2)], axis=1)
df.columns = ['Y', 'X1', 'X2']
df.head()
lmreg = LinearRegression().fit(df[['X1', 'X2']], df['Y'])
coef = pd.DataFrame([lmreg.coef_])
intercept = pd.DataFrame([lmreg.intercept_])
allval = pd.concat([intercept, coef], axis=1)
allval.columns = ['beta0', 'beta1', 'beta2']
allval
plt.xkcd()
plt.figure(figsize=(25, 10))
[plt.axhline(k , 0, 1000, lw=2, linestyle='--', c='r') for k in [-12.474106, 35.896213, -23.297124]]
plt.plot(betadf)
# **So, we can see that *backfitting* provides similar estimates for coefficient values as that of performing multiple linear regression.**
# ### 11.g. Number of backfitting iterations required to obtain a "good" approximation to the multiple regression coefficient estimates
# **The first value was close enough. So, just one backfitting iteration sufficed. However, it's always a good idea to have multiple iterations to get a good estimation.**
| Chapter 7/Applied Exercises/11.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''udemyPro_py388'': venv)'
# name: python3
# ---
def squaring_list(input_list):
squared_list = [val**2 for val in input_list]
return input_list, squared_list, True
# +
my_list = [val for val in range(10)]
return_tuple = squaring_list(my_list)
print(type(return_tuple), return_tuple)
# Tuple unpacking: all values has to be unpacked
my_list, squared_list, func_success = return_tuple
# or if don't want all:
my_list, *other_return_value = return_tuple
print("myList:", my_list)
print("other:", other_return_value, type(other_return_value))
# +
# Tuple packing
x = 2
y = 3
z = 3
# Klammern werden nicht benötigt, sieht aber besser aus
t = x, y, z
print(t, type(t))
# -
| Python/zzz_training_challenge/UdemyPythonPro/Chapter4_Iterables/Tuples/packing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Spatial model
# +
import random
import os
import numpy as np
import json
import corner
import random
import os
import pandas as pd
import geopandas as gp
import datetime
import scipy
import matplotlib
import matplotlib.colors as colors
import matplotlib.dates as mdates
import math
import xarray as xr
import emcee
import matplotlib.pyplot as plt
from covid19model.optimization import objective_fcns
from covid19model.models import models
from covid19model.models.utils import name2nis
from covid19model.data import google, sciensano, polymod, model_parameters
from covid19model.visualization.output import population_status, infected
from covid19model.visualization.optimization import plot_fit, traceplot
# OPTIONAL: Load the "autoreload" extension so that package code can change
# %load_ext autoreload
# OPTIONAL: always reload modules so that as you change code in src, it gets loaded
# %autoreload 2
# -
# ### Check name2nis functionality
name2nis('<NAME>')
name2nis('<NAME>')
name2nis('<NAME>')
name2nis(5)
# ### Extract population size (initN)
initN_df=pd.read_csv('../../data/interim/census_2011/initN.csv', index_col=[0])
initN=initN_df.iloc[:,2:].values
initN_df.head()
# ### Load Polymod interaction matrices
dummy_initN, Nc_home, Nc_work, Nc_schools, Nc_transport, Nc_leisure, Nc_others, Nc_total = polymod.get_interaction_matrices()
# ### Initialize model
# Load the parameters using `get_COVID19_SEIRD_parameters()`.
params = model_parameters.get_COVID19_SEIRD_parameters(spatial=True)
# Add the delayed ramp parameters to the parameter dictionary.
params.update({'l': 5,
'tau': 5})
# Define a cluster of 5 infectees in one or two arrondissements
E = np.zeros(initN.shape)
E[np.where(initN_df.index.values==name2nis('arrondissement ieper'))[0][0],:] = 2
E[np.where(initN_df.index.values==name2nis('arrondissement tongeren'))[0][0],:] = 2
# Define the initial condition: one exposed inidividual in every age category
initial_states = {'S': initN, 'E': E}
# Load the compliance model
from covid19model.models.compliance import ramp_2
# Initialize the model
model = models.COVID19_SEIRD_sto_spatial(initial_states, params, compliance=ramp_2, discrete=True)
# ### Change beta to a higher value
model.parameters['beta'] = 0.5
# ### Define a checkpoints dictionary and perform some simulations
chk = {
'time': ['20-04-2020'],
'Nc': [0.3*(1.0*Nc_home + 0.3*Nc_work + 0.4*Nc_transport)]
}
fig,ax=plt.subplots()
for i in range(5):
out=model.sim('21-09-2020',excess_time=50,checkpoints=chk)
sumNIS=out.sum(dim="place").sum(dim="Nc")
plt.plot(out["time"].values[50:200],sumNIS["H_in"].values[50:200],alpha=0.05,color='green')
out
# ## Visualisation on a map
# ### Read the arrondissements shape file
# Read shape file
gemeentes = gp.read_file("../../data/raw/geopandas/BE/BELGIUM__Municipalities.shp")
# select the columns that you with to use for the dissolve and that will be retained
gemeentes_new = gemeentes[['arrond','geometry']]
# dissolve the state boundary by region
arrondissementen = gemeentes_new.dissolve(by='arrond')
# ### Perform a single simulation
out=model.sim(250,checkpoints=chk)
sumNIS=out.sum(dim="place").sum(dim="Nc")
plt.plot(out["time"],sumNIS["M"],alpha=0.15,color='green')
# +
output_path = '../../results/maps'
# create the plot
fig, ax = plt.subplots(figsize = (12,12))
start = 0
for day in range(start,len(out['time'].values)):
# Get data
data2plot = out.sum(dim="Nc")['M'][:,day].values/initN_df['total'].values*100
arrondissementen['data'] = data2plot
# Visualize data
if day == start:
fig = arrondissementen.plot(column = 'data', ax=ax, cmap='plasma',
norm=colors.LogNorm(vmin=0.001, vmax=1), legend=True, edgecolor = 'k')
else:
fig = arrondissementen.plot(column = 'data', ax=ax, cmap='plasma',
norm=colors.LogNorm(vmin=0.001, vmax=1), legend=False, edgecolor = 'k')
# Disable axis
ax.set_axis_off()
# This will save the figure as a high-res png in the output path. you can also save as svg if you prefer.
chart = fig.get_figure()
chart.savefig('maps/'+str(day)+'_B.jpg',dpi=100)
plt.close()
# -
fig,ax=plt.subplots()
fig.set_size_inches(16, 4)
ax.plot(out['time'][50:200],out.sel(place=name2nis('arrondissement antwerpen')).sum(dim='Nc')['M'][50:200],color='blue',alpha=0.40)
ax.plot(out['time'][50:200],out.sel(place=name2nis('arrondissement brussel-hoofdstad')).sum(dim='Nc')['M'][50:200],color='red',alpha=0.40)
ax.plot(out['time'][50:200],out.sel(place=name2nis('arrondissement luik')).sum(dim='Nc')['M'][50:200],color='black',alpha=0.40)
ax.legend(['Antwerp','Brussels','Luik'])
ax.axvline(130,color='black',linestyle='dashed')
ax.set_title('Daily hospitalizations $(H_{in})$')
fig.savefig('daily_hosp.jpg',dpi=300)
| notebooks/scratch/0.1-twallema-spatial-stochastic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''venv'': venv)'
# name: python3
# ---
# # Manipulando variavéis
x = 10
y = 'Olá'
x, y
# #### Trocando os valores
z = x
x = y
y = z
# #### Veja que para fazer a troca de valores entre as variavéis foi preciso declacrar z = x, caso contrário os valores de "x" e "y" seriam os mesmos
x, y
# #### Forma mais pratica
x, y = y, x
x,y
| introducao/jupyter-notebook/manipulando_variaveis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Reduction Operators
# This section shows you how to use operators from `reductions` module.
# 1. Start with a simple pipeline based on `ExternalSource`. Input has two samples per batch. Shape of both samples is (3, 3). First contains consecutive numbers, second contains consecutive even numbers. This will be useful to visualize possible reductions.
# +
import nvidia.dali.fn as fn
import nvidia.dali.types as types
import nvidia.dali.backend as backend
from nvidia.dali.pipeline import Pipeline
import numpy as np
batch_size = 2
def get_batch():
return [np.reshape(np.arange(9), (3, 3)) * (i+1) for i in range(batch_size)]
def run_and_print(pipe):
pipe.build()
output = pipe.run()
for i, out in enumerate(output):
if type(out) == backend.TensorListGPU:
out = out.as_cpu()
output_array = out.as_array()
print('Output {}:\n{} \n'.format(i, output_array))
pipe = Pipeline(batch_size=batch_size, num_threads=4, device_id=0)
with pipe:
input = fn.external_source(source=get_batch, dtype=types.INT64)
pipe.set_outputs(input)
run_and_print(pipe)
# -
# 2. Add some reductions to the pipeline above. Begin with the `Max` operator.
# +
pipe = Pipeline(batch_size=batch_size, num_threads=4, device_id=0)
with pipe:
input = fn.external_source(source=get_batch, dtype=types.INT64)
max = fn.reductions.max(input)
pipe.set_outputs(max)
run_and_print(pipe)
# -
# As you can see, it returned the biggest value from each sample.
#
# 3. Perform other reductions like `Min` or `Sum`.
# +
pipe = Pipeline(batch_size=batch_size, num_threads=4, device_id=0)
with pipe:
input = fn.external_source(source=get_batch, dtype=types.INT64)
min = fn.reductions.min(input)
sum = fn.reductions.sum(input)
pipe.set_outputs(min, sum)
run_and_print(pipe)
# -
# In the code samples above we see reductions performed for all elements of each sample.
#
# 4. Reductions can be performed along an arbitrary set of axes. To control this behavior you can use `axes` argument.
# +
pipe = Pipeline(batch_size=batch_size, num_threads=4, device_id=0)
with pipe:
input = fn.external_source(source=get_batch, dtype=types.INT64)
min_axis_0 = fn.reductions.min(input, axes=0)
min_axis_1 = fn.reductions.min(input, axes=1)
pipe.set_outputs(min_axis_0, min_axis_1)
run_and_print(pipe)
# -
# `Min` reduction was performed along axis 0 and 1, and it returned minimum element per column and per row respectively.
#
# To make it easier, reductions support `axis_names` argument. It allows to pass axis names rather than indices. Names are matched based on the layout of the input. You need to provide layout argument in `ExternalSource`.
# +
pipe = Pipeline(batch_size=batch_size, num_threads=4, device_id=0)
with pipe:
input = fn.external_source(source=get_batch, layout='AB', dtype=types.INT64)
min_axis_0 = fn.reductions.min(input, axis_names='A')
min_axis_1 = fn.reductions.min(input, axis_names='B')
pipe.set_outputs(min_axis_0, min_axis_1)
run_and_print(pipe)
# -
# **Note**: Passing all axes will result in a full reduction, while passing empty axes will result in no reduction. This is true for both indices and layouts.
# +
pipe = Pipeline(batch_size=batch_size, num_threads=4, device_id=0)
with pipe:
input = fn.external_source(source=get_batch, layout='AB', dtype=types.INT64)
min_axes_full = fn.reductions.min(input, axes=(0, 1))
min_axes_empty = fn.reductions.min(input, axes=())
min_layout_full = fn.reductions.min(input, axis_names='AB')
min_layout_empty = fn.reductions.min(input, axis_names='')
pipe.set_outputs(
min_axes_full, min_axes_empty,
min_layout_full, min_layout_empty)
run_and_print(pipe)
# -
# 5. For inputs with higher dimensionality you can pass any combination of the axes.
def get_batch():
return [np.reshape(np.arange(8, dtype=np.int32), (2, 2, 2)) * (i+1) for i in range(batch_size)]
# +
pipe = Pipeline(batch_size=batch_size, num_threads=4, device_id=0)
with pipe:
input = fn.external_source(source=get_batch, layout='ABC', dtype=types.INT32)
min_axes_empty = fn.reductions.min(input, axes=())
min_axes_0_1 = fn.reductions.min(input, axes=(0, 1))
min_layout_A_C = fn.reductions.min(input, axis_names='AC')
pipe.set_outputs(min_axes_empty, min_axes_0_1, min_layout_A_C)
run_and_print(pipe)
# -
# 6. There are reductions that require additional inputs. `StdDev` and `Variance` rely on the externally provided mean, and it can be calculated with the `Mean` reduction operator.
# +
pipe = Pipeline(batch_size=batch_size, num_threads=4, device_id=0)
with pipe:
input = fn.external_source(source=get_batch, dtype=types.INT32)
mean = fn.reductions.mean(input)
std_dev = fn.reductions.std_dev(input, mean)
variance = fn.reductions.variance(input, mean)
pipe.set_outputs(mean, std_dev, variance)
run_and_print(pipe)
# -
# 7. By default, reductions remove unnecessary dimensions. This behaviour can be controlled with the `keep_dims` argument.
# +
pipe = Pipeline(batch_size=batch_size, num_threads=4, device_id=0)
with pipe:
input = fn.external_source(source=get_batch, dtype=types.INT32)
mean = fn.reductions.mean(input)
std_dev = fn.reductions.std_dev(input, mean, keep_dims=True)
variance = fn.reductions.variance(input, mean)
pipe.set_outputs(mean, std_dev, variance)
run_and_print(pipe)
# -
# In the code sample above applying reductions resulted in changing the output type.
#
# 8. The argument `dtype` can be used to specify the desired output data type.
# +
pipe = Pipeline(batch_size=batch_size, num_threads=4, device_id=0)
with pipe:
input = fn.external_source(source=get_batch, dtype=types.INT32)
sum_int_64 = fn.reductions.sum(input, dtype=types.INT64)
sum_float = fn.reductions.sum(input, dtype=types.FLOAT)
pipe.set_outputs(sum_int_64, sum_float)
run_and_print(pipe)
# -
# **Note**: Not all data types combinations are supported. The default behaviour varies from operator to operator. A general rule is for the output type to be able to accommodate the result depending on the input type. For example, for the input type `INT32` the default output type of a sum is `INT32` and the default output type of a mean is `FLOAT`.
#
# 9. All reductions can be offloaded to the GPU. GPU variants work the same way as their CPU counterparts. Below we show a code sample containing all reductions offloaded to the GPU with various parameters.
# +
pipe = Pipeline(batch_size=batch_size, num_threads=4, device_id=0)
with pipe:
input = fn.external_source(source=get_batch, layout='ABC', dtype=types.INT32)
min = fn.reductions.min(input.gpu(), axis_names='AC', keep_dims=True)
max = fn.reductions.max(input.gpu(), keep_dims=True)
sum = fn.reductions.sum(input.gpu(), dtype=types.INT64)
mean = fn.reductions.mean(input.gpu(), axes=0)
mean_square = fn.reductions.mean_square(input.gpu())
rms = fn.reductions.rms(input.gpu(), axes=(), dtype=types.FLOAT)
std_dev = fn.reductions.std_dev(input.gpu(), mean, axes=0)
variance = fn.reductions.variance(input.gpu(), mean.gpu(), axes=0, keep_dims=True)
pipe.set_outputs(
min, max, sum, mean, mean_square, rms, std_dev, variance)
run_and_print(pipe)
# -
| docs/examples/general/reductions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import s3fs
import xarray as xr
import numpy as np
import cmocean
import matplotlib.pyplot as plt
# +
# open the data with s3fs from the databucket
fsg = s3fs.S3FileSystem(anon=False,
client_kwargs={
'endpoint_url': 'https://karen.uiogeo-apps.sigma2.no'
})
data_path1 = 's3://data/CREG12.L75-REF08_mesh_zgr.zarr'
data_path2 = 's3://data/vel_dataIII.zarr/vel_dataIII.zarr'
remote_files1 = fsg.glob(data_path1)
remote_files2 = fsg.glob(data_path2)
store1 = s3fs.S3Map(root=data_path1, s3=fsg, check=False)
store2 = s3fs.S3Map(root=data_path2, s3=fsg, check=False)
dzz = xr.open_zarr(store=store1, consolidated=True)
dv = xr.open_zarr(store=store2, consolidated=True)
# -
dv
u = dv.vozocrtx.mean(dim='time_counter')
v = dv.vomecrty.mean(dim='time_counter')
u.shape
# +
# Quiver
x = np.linspace(0, 1200, 1200)
dx = 20
y = np.linspace(0, 900, 900)
dy = 20
X, Y = np.meshgrid(y, x)
# +
# chooseing a level to look at
level = 35
depth = dv.depth[level].values
print(depth)
y_grid = np.array(Y.T[::dx,::dy])
print(y_grid.shape)
x_grid = np.array(X.T[::dx,::dy])
u_grid = np.array(u[level,::dx,::dy].values)#endring
print(u_grid.shape)
v_grid = np.array(v[level,::dx,::dy].values)
vel = (u**2 + v**2)**0.5
vel_grid = np.array(vel[level,::dx,::dy].values)
# +
# plotting the quiver
import warnings
warnings.filterwarnings('ignore')
fig, ax = plt.subplots(1, 1, figsize=(10, 8))
cmap_o = plt.get_cmap(cmocean.cm.speed)
ax.contour(dv.mbathy.isel(t=0), colors='grey', levels=8, linewidths=0.4)
norm = plt.Normalize(vmin=0.,vmax=10.,clip=False)
c = ax.quiver(y_grid,
x_grid,
u_grid/vel_grid,
v_grid/vel_grid,
vel_grid*100, #*100 to get cm/s as the mooring velcity are in
scale = 40,
width=0.003,
#headwidth=2,
#headlength=1,
cmap = cmap_o,
norm=norm)
fig.colorbar(c,orientation='vertical', pad = 0.04, aspect=50, extendrect=True)
plt.title(f'Quiverplot of time averaged current at '+ str(depth) + 'm depth')
plt.savefig('quiver_333.pdf', dpi = 300)
# +
# chooseing a level to look at
level = 0
depth = dv.depth[level].values
print(depth)
y_grid = np.array(Y.T[::dx,::dy])
print(y_grid.shape)
x_grid = np.array(X.T[::dx,::dy])
u_grid = np.array(u[level,::dx,::dy].values)#endring
print(u_grid.shape)
v_grid = np.array(v[level,::dx,::dy].values)
vel = (u**2 + v**2)**0.5
vel_grid = np.array(vel[level,::dx,::dy].values)
# +
import warnings
warnings.filterwarnings('ignore')
fig, ax = plt.subplots(1, 1, figsize=(10, 8))
cmap_o = plt.get_cmap(cmocean.cm.speed)
ax.contour(dv.mbathy.isel(t=0), colors='grey', levels=8, linewidths=0.4)
norm = plt.Normalize(vmin=0.,vmax=10.,clip=False)
c = ax.quiver(y_grid,
x_grid,
u_grid/vel_grid,
v_grid/vel_grid,
vel_grid*100, #*100 to get cm/s as the mooring velcity are in
scale = 40,
width=0.003,
#headwidth=2,
#headlength=1,
cmap = cmap_o,
norm=norm)
fig.colorbar(c,orientation='vertical', pad = 0.04, aspect=50, extendrect=True)
plt.title(f'Quiverplot of time averaged current at '+ str(depth) + 'm depth')
plt.savefig('quiver_0.pdf', dpi = 300)
# -
| quiver.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from matplotlib.font_manager import FontProperties
from collections import Counter
from tqdm.auto import tqdm
import sys
import gc
# plt.style.use('fivethirtyeight')
# print(plt.style.available)
# -
# # Data
train = pd.read_csv('inputs/train_set.csv',sep='\t')
test = pd.read_csv('inputs/test_a.csv',sep='\t')
train.head()
# # EDA
# ### 文本长度
# 文本长度分布与统计
train['len'] = train['text'].apply(lambda x: len(x.split(' ')))
test['len'] = test['text'].apply(lambda x: len(x.split(' ')))
train['len'].describe(),test['len'].describe()
# **Task2-1:**
# - train中句子(篇章)包含字符的平均个数为907.20;
# - test中句子(篇章)包含字符的平均个数为909.84
plt.figure(figsize=(10,5))
fig1 = sns.distplot(train['len'], bins=100,color='blue')
fig2 = sns.distplot(test['len'], bins=100,color='yellow')
# plt.xlim([0, max(max(train['len_text']), max(test['len_text']))])
plt.xlabel("length of sample")
plt.ylabel("prob of sample")
plt.legend(['train_len','test_len'])
# ### 同分布验证
import scipy
scipy.stats.ks_2samp(train['len'], test['len'])
# P值为0.52,比指定的显著水平(假设为5%)大,我们认为二者同分布。
# ### 标签分布
# +
# from matplotlib.font_manager import _rebuild
# _rebuild() #reload一下
# +
# zhfont = FontProperties(fname="inputs/msyhl.ttc", size=14)
# zhfont = FontProperties(fname="simsun.ttc", size=14)
label_to_index_dict={'科技': 0, '股票': 1, '体育': 2, '娱乐': 3, '时政': 4,
'社会': 5, '教育': 6, '财经': 7, '家居': 8, '游戏': 9,
'房产': 10, '时尚': 11, '彩票': 12, '星座': 13}
index_to_label_dict = {v:k for k,v in label_to_index_dict.items()}
# -
import matplotlib
matplotlib.rcParams['font.sans-serif'] = ['SimHei']
matplotlib.rcParams['font.family'] ='sans-serif'
# +
# Task2-3 类别分布规律:类别分布严重不平衡,会对预测造成较大影响
plt.bar(x=range(0,14),height=np.bincount(train['label']))
# plt.xlabel("label")
# plt.ylabel("number of sample")
# sns.histplot(train,x='label',binwidth=1)
plt.xticks(range(14),list(index_to_label_dict.values()), rotation=60)
plt.show()
# -
plt.figure(figsize=(15,10))
ax = sns.catplot(x='label', y='len', data=train, kind='strip')
plt.xticks(range(14), list(index_to_label_dict.values()), rotation=60);
# Task2-2 统计数据集中不同类别下句子平均字符的个数
group = train.groupby('label').agg({'len':np.mean})
group.rename({'len':'mean_len'})
group
# 类别不平衡,且各类别的文本长度存在差异
log_len_dist = np.log(1+train['len'])
log_len_test_dist = np.log(1+test['len'])
plt.figure(figsize=(15,5))
ax = sns.distplot(log_len_dist)
ax = sns.distplot(log_len_test_dist, color='yellow')
plt.xlabel("log length of sample")
plt.ylabel("prob of log")
plt.legend(['train_len','test_len'])
# 从log图上看,基本同分布
# ks检验是否为正态分布
_, lognormal_ks_pvalue = scipy.stats.kstest(rvs=log_len_dist, cdf='norm')
lognormal_ks_pvalue
del log_len_dist, log_len_test_dist
gc.collect()
# p<5%, 拒绝原假设 -> 不是正态分布
# - pvalue:p 值 越大,越支持原假设,一般会和指定显著水平 5% 比较,大于 5%,支持原假设;【支持原假设无法否定原假设,不代表原假设绝对正确】
#
# 对元数据做box-cox变换,再做一次检验
trans_data, lam = scipy.stats.boxcox(train['len']+1)
scipy.stats.normaltest(trans_data)
# p<5%, 拒绝原假设(正态分布)
# ### 其他特征
# - text-split,将text字段分词
# - len,每条新闻长度
# - first_char,新闻第一个字符
# - last_char,新闻最后一个字符
# - most_freq,新闻最常出现的字符
# %%time
train['text_split'] = train['text'].apply(lambda x:x.split())
# train['len'] = train['text'].apply(lambda x:len(x.split()))
train['first_char'] = train['text_split'].apply(lambda x:x[0])
train['last_char'] = train['text_split'].apply(lambda x:x[-1])
train['most_freq'] = train['text_split'].apply(lambda x:np.argmax(np.bincount(x)))
train.head()
train.to_pickle('inputs/train_11.pkl')
globals().keys()
# +
# dir()
# -
# 类别信息表
# - count,该类别新闻个数
# - len_mean,该类别新闻平均长度
# - len_std,该类别新闻长度标准差
# - len_min,该类别新闻长度最小值
# - len_max,该类别新闻长度最大值
# - freq_fc,该类别新闻最常出现的第一个字符
# - freq_lc,该类别新闻最常出现的最后一个字符
# - freq_freq,该类别新闻最常出现的字符
train_info = pd.DataFrame(columns=['count','len_mean','len_std','len_min','len_max',
'freq_fc','freq_lc','freq_freq'])
for name, group in train.groupby('label'):
count = len(group) # 该类别新闻数
len_mean = np.mean(group['len']) # 该类别长度平均值
len_std = np.std(group['len']) # 长度标准差
len_min = np.min(group['len']) # 最短的新闻长度
len_max = np.max(group['len']) # 最长的新闻长度
freq_fc = np.argmax(np.bincount(group['first_char'])) # 最频繁出现的首词
freq_lc = np.argmax(np.bincount(group['last_char'])) # 最频繁出现的末词
freq_freq = np.argmax(np.bincount(group['most_freq'])) # 该类别最频繁出现的词
# freq_freq_5 =
train_info.loc[name] = [count,len_mean,len_std,len_min,len_max,freq_fc,freq_lc,freq_freq]
train_info
# +
# import gc
# # del
# gc.collect()
# -
# ### 字符分布
# +
# %%time
from collections import Counter
all_lines = ' '.join(list(train['text']))
word_count = Counter(all_lines.split(" "))
word_count = sorted(word_count.items(), key=lambda d:d[1], reverse = True)
print(len(word_count)) # 字典长度
print(word_count[0]) # 字典最高频词
print(word_count[-1]) # 字典最低频词
# -
text_of_labels = {}
for i in tqdm(range(14)):
text = ' '.join(list(train[train['label']==i]['text']))
text_of_labels[i] = text
# exec('text_of_label_{}={}'.format(i,i))
# %%time
print('Top 5 frequent words and frequency info: ')
for i in tqdm(range(len(text_of_labels))):
label_word_count = Counter(text_of_labels[i].split(' '))
label_word_count = sorted(label_word_count.items(), key=lambda d:d[1], reverse = True)
print('label {}: {}'.format(i,label_word_count[:5]))
del label_word_count, text_of_labels
gc.collect()
# 根据字符频率反推出标点符号
# %%time
train['text_unique'] = train['text'].apply(lambda x: ' '.join(list(set(x.split(' '))))) # set->只保留不重复元素
all_lines_unique = ' '.join(list(train['text_unique']))
word_count_unique = Counter(all_lines_unique.split(" "))
word_count_unique = sorted(word_count_unique.items(), key=lambda d:int(d[1]), reverse = True)
# 打印整个训练集中覆盖率前5的词
for i in range(5):
print("{} occurs {} times, {}%".format(word_count_unique[i][0],
word_count_unique[i][1],
(word_count_unique[i][1]/200000)*100))
# +
# set('1 2 3 1 2 4'.split(' '))
# -
# ### WordCloud
# +
# from wordcloud import WordCloud
# import imageio
# def gen_img(texts, img_file):
# data = ' '.join(text for text in texts)
# image_coloring = imageio.imread(img_file)
# wc = WordCloud(
# background_color='white',
# mask=image_coloring,
# # width=100, height=100,
# margin=1,
# # scale=32, # 越大越清楚,输出图片分辨率越大
# # font_path='WC_inputs/fonts/FZQKBYSJW.TTF', # 方正清刻宋
# font_path='msyh.ttc' # 微软雅黑
# )
# wc.generate(data)
# plt.figure()
# plt.imshow(wc, interpolation="bilinear")
# plt.axis("off")
# plt.show()
# wc.to_file(img_file.split('.')[0] + '_wc1.png')
# +
# gen_img(all_lines[:80000000], 'WC_inputs/figs/edge.png')
# -
| EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import lightgbm as lgb
import numpy as np
from sklearn.metrics import confusion_matrix, classification_report, f1_score
df_train_origin = pd.read_csv('../data/train.csv')
df_test_origin = pd.read_csv('../data/test_leak.csv')
df_train_vecs = pd.read_csv('../fact/train_vecs.csv')
df_test_vecs = pd.read_csv('../fact/test_vecs.csv')
# -
df_train = pd.merge(df_train_origin ,df_train_vecs, on='id')
df_test = pd.merge(df_test_origin ,df_test_vecs, on='id')
# +
def get_c_with_prefix(train, prefix):
return [column for column in train.columns.tolist() if prefix == column[:len(prefix)]]
c_vecs = get_c_with_prefix(df_train, 'vecs')
X_train, X_valid = df_train[c_vecs], df_test[c_vecs]
y_train, y_valid = df_train.target.values, df_test.target.values
lgb_train = lgb.Dataset(X_train, y_train)
lgb_valid = lgb.Dataset(X_valid, y_valid, reference=lgb_train)
def lgb_f1_score(y_hat, data):
y_true = data.get_label()
y_hat = np.round(y_hat) # scikits f1 doesn't like probabilities
return 'f1', f1_score(y_true, y_hat, average=None)[0], True
lgbm_params = {
'objective': 'binary',
'metric': 'binary_logloss',
'verbosity': 4,
'boosting_type': 'gbdt',
'learning_rate': 0.1,
'lambda_l1': 3.642434329823594,
'lambda_l2': 1.0401748765492007e-08,
'num_leaves': 172,
'feature_fraction': 0.8251431673667773,
'bagging_fraction': 0.9755605959841563,
'bagging_freq': 2,
'min_child_samples': 5,
'random_state': 68
}
model = lgb.train(
lgbm_params,
lgb_train,
valid_sets=lgb_valid,
verbose_eval=True,
feval=lgb_f1_score,
num_boost_round=300,
)
y_pred = model.predict(X_valid, num_iteration=model.best_iteration)
y_pred_cls = y_pred >= 0.5
print(f1_score(y_valid, y_pred_cls, average=None)[0])
# -
pd.DataFrame({'id': df_test.id, 'target': y_pred_cls.astype(int)}).to_csv('../output/submit.csv', index=None)
| notebook/bert.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Making Pandas DataFrames from API Requests
# In this example, we will use the U.S. Geological Survey's API to grab a JSON object of earthquake data and convert it to a `pandas.DataFrame`.
#
# USGS API: https://earthquake.usgs.gov/fdsnws/event/1/
# ### Get Data from API
# +
import datetime as dt
import pandas as pd
import requests
yesterday = dt.date.today() - dt.timedelta(days=1)
api = 'https://earthquake.usgs.gov/fdsnws/event/1/query'
payload = {
'format': 'geojson',
'starttime': yesterday - dt.timedelta(days=30),
'endtime': yesterday
}
response = requests.get(api, params=payload)
# let's make sure the request was OK
response.status_code
# -
# Response of 200 means OK, so we can pull the data out of the result. Since we asked the API for a JSON payload, we can extract it from the response with the `json()` method.
#
# ### Isolate the Data from the JSON Response
# We need to check the structures of the response data to know where our data is.
earthquake_json = response.json()
earthquake_json.keys()
# The USGS API provides information about our request in the `metadata` key. Note that your result will be different, regardless of the date range you chose, because the API includes a timestamp for when the data was pulled:
earthquake_json['metadata']
# Each element in the JSON array `features` is a row of data for our dataframe.
type(earthquake_json['features'])
# Your data will be different depending on the date you run this.
earthquake_json['features'][0]
# ### Convert to DataFrame
# We need to grab the `properties` section out of every entry in the `features` JSON array to create our dataframe.
earthquake_properties_data = [
quake['properties'] for quake in earthquake_json['features']
]
df = pd.DataFrame(earthquake_properties_data)
df.head()
# ### (Optional) Write Data to CSV
df.to_csv('earthquakes.csv', index=False)
# <hr>
# <div>
# <a href="./2-creating_dataframes.ipynb">
# <button style="float: left;">← Previous Notebook</button>
# </a>
# <a href="./4-inspecting_dataframes.ipynb">
# <button style="float: right;">Next Notebook →</button>
# </a>
# </div>
# <br>
# <hr>
| ch_02/3-making_dataframes_from_api_requests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # OPTICS
# This Code template is for the Cluster analysis using a simple OPTICS (Ordering Points To Identify Cluster Structure) Clustering algorithm.It draws inspiration from the DBSCAN clustering algorithm. The Code Template includes 2D and 3D cluster visualization of the Clusters.
# ### Required Packages
# !pip install plotly
import operator
import warnings
import itertools
import numpy as np
import pandas as pd
import seaborn as sns
import plotly.express as px
import matplotlib.pyplot as plt
from mpl_toolkits import mplot3d
import plotly.graph_objects as go
from sklearn.cluster import OPTICS
warnings.filterwarnings("ignore")
# ### Initialization
#
# Filepath of CSV file
file_path = ""
# List of features which are required for model training
features=[]
# ### Data Fetching
#
# Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
#
# We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
df=pd.read_csv(file_path)
df.head()
# ### Feature Selections
#
# It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
#
# We will assign all the required input features to X.
X = df[features]
# ### Data Preprocessing
#
# Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
#
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
# Calling preprocessing functions on the feature and target set.
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
X.head()
# ### Model
#
# The OPTICS algorithm shares many similarities with the DBSCAN algorithm, and can be considered a generalization of DBSCAN that relaxes the eps requirement from a single value to a value range. The key difference between DBSCAN and OPTICS is that the OPTICS algorithm builds a reachability graph, which assigns each sample both a reachability_ distance, and a spot within the cluster ordering_ attribute; these two attributes are assigned when the model is fitted, and are used to determine cluster membership.
# [More detail](https://scikit-learn.org/stable/modules/clustering.html#optics)
#
# #### Tuning Parameters
#
# **min_samples**: The number of samples in a neighborhood for a point to be considered as a core point.
#
# **max_eps**: The maximum distance between two samples for one to be considered as in the neighborhood of the other.
#
# **metric**: Metric to use for distance computation. Any metric from scikit-learn or scipy.spatial.distance can be used.
#
# **p**: Parameter for the Minkowski metric from pairwise_distances.
#
# **cluster_method**: The extraction method used to extract clusters using the calculated reachability and ordering.
#
# **eps**: The maximum distance between two samples for one to be considered as in the neighborhood of the other.
#
# **xi**: Determines the minimum steepness on the reachability plot that constitutes a cluster boundary.
#
# **min_cluster_size**: Minimum number of samples in an OPTICS cluster, expressed as an absolute number or a fraction of the number of samples (rounded to be at least 2).
#
# **algorithm**: Algorithm used to compute the nearest neighbors
#
# [For more detail on API](https://scikit-learn.org/stable/modules/generated/sklearn.cluster.OPTICS.html)
y_pred = OPTICS(max_eps=20,min_cluster_size=30,n_jobs=-1).fit_predict(X)
# ### Cluster Analysis
#
# First, we add the cluster labels from the trained model into the copy of the data frame for cluster analysis/visualization.
ClusterDF = X.copy()
ClusterDF['ClusterID'] = y_pred
ClusterDF.head()
# #### Cluster Records
# The below bar graphs show the number of data points in each available cluster.
ClusterDF['ClusterID'].value_counts().plot(kind='bar')
# #### Cluster Plots
# Below written functions get utilized to plot 2-Dimensional and 3-Dimensional cluster plots on the available set of features in the dataset. Plots include different available clusters along with cluster centroid.
# +
def Plot2DCluster(X_Cols,df):
for i in list(itertools.combinations(X_Cols, 2)):
plt.rcParams["figure.figsize"] = (8,6)
xi,yi=df.columns.get_loc(i[0]),df.columns.get_loc(i[1])
for j in df['ClusterID'].unique():
DFC=df[df.ClusterID==j]
plt.scatter(DFC[i[0]],DFC[i[1]],cmap=plt.cm.Accent,label=j)
plt.xlabel(i[0])
plt.ylabel(i[1])
plt.legend()
plt.show()
def Plot3DCluster(X_Cols,df):
for i in list(itertools.combinations(X_Cols, 3)):
xi,yi,zi=df.columns.get_loc(i[0]),df.columns.get_loc(i[1]),df.columns.get_loc(i[2])
fig,ax = plt.figure(figsize = (16, 10)),plt.axes(projection ="3d")
ax.grid(b = True, color ='grey',linestyle ='-.',linewidth = 0.3,alpha = 0.2)
for j in df['ClusterID'].unique():
DFC=df[df.ClusterID==j]
ax.scatter3D(DFC[i[0]],DFC[i[1]],DFC[i[2]],alpha = 0.8,cmap=plt.cm.Accent,label=j)
ax.set_xlabel(i[0])
ax.set_ylabel(i[1])
ax.set_zlabel(i[2])
plt.legend()
plt.show()
def Plotly3D(X_Cols,df):
for i in list(itertools.combinations(X_Cols,3)):
xi,yi,zi=df.columns.get_loc(i[0]),df.columns.get_loc(i[1]),df.columns.get_loc(i[2])
fig2=px.scatter_3d(df, x=i[0], y=i[1],z=i[2],color=df['ClusterID'])
fig2.show()
# -
sns.set_style("whitegrid")
sns.set_context("talk")
plt.rcParams["lines.markeredgewidth"] = 1
sns.pairplot(data=ClusterDF, hue='ClusterID', palette='Dark2', height=5)
Plot2DCluster(X.columns,ClusterDF)
Plot3DCluster(X.columns,ClusterDF)
Plotly3D(X.columns,ClusterDF)
# #### [Created by <NAME>](https://github.com/Thilakraj1998)
| Clustering/OPTICS/OPTICS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/pathstream-curriculum/Stats-Scratch/blob/master/Probability_Distributions.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="TIfH-uGXJlaC" colab_type="text"
# # Python exercises for probability distributions
#
# ## Learning objectives
# 1. Simulate a binary experiment.
# 2. Run a series of binary experiment simulations and compare the result with the probability associated with a specific number of successful outcomes computed using the binomial distribution.
# 2. Simulate an experiment to find the probability associated with the number of trials to reach the first success and compare with the result computed using the geometric distribution.
# 4. Simulate the probability of recording a given number of events in an interval or region and compare the result with the probability computed directly using the Poisson distribution.
# + [markdown] id="Mpexpq6CXk-k" colab_type="text"
# **Step 0:** Import Python packages
# + id="QvH2euPsEryv" colab_type="code" colab={}
# Import required packages
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
from scipy import stats
# Set up seaborn dark view
sns.set()
# + [markdown] id="4ik2DwRRqf3a" colab_type="text"
# ### **Learning objective 1:** Simulate a binary experiment.
# **Step 1:** Define function that conducts a single Bernouilli trial.
# + id="T1_rIZ4UMnbH" colab_type="code" outputId="29acfabe-1d11-45d2-f382-ccf2a5e28e0e" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Define a function called "bernouilli_trial" that takes a variable "prob_of_success" indicating the probability of a successful outcome.
def bernouilli_trial(prob_of_success=0.5):
# Choose a random number between 0 and 1
result = np.random.random_sample()
# If the result is less than or equal to the probability of success, declare a successful outcome
if result <= prob_of_success:
outcome = 'success'
# Otherwise declare a failure
else:
outcome = 'failure'
# Return the outcome
return outcome
# Run the new function to test
bernouilli_trial()
# + [markdown] id="GgFTZ-MDeOHW" colab_type="text"
# **Step 2:** Run multiple Bernouilli trials and record the result
# + id="HnUDYeBhOtDI" colab_type="code" outputId="403448d1-c465-4323-cb89-0c81434464c3" colab={"base_uri": "https://localhost:8080/", "height": 185}
# Define a function to run multiple Bernouilli trials
def binary_experiment(trials=10, p_of_success=0.5):
# Initialize a list to store all the outcomes
outcomes = []
# Run through a loop to perform each Bernouilli trial and record each outcome
for i in range(trials):
outcome = bernouilli_trial(prob_of_success=p_of_success)
outcomes.append(outcome)
# Return the resulting list of outcomes
return outcomes
# Run the new function to test
binary_experiment()
# + [markdown] id="kN7mAuCILVjh" colab_type="text"
# ### **Learning objective 2:** Run a series of binary experiment simulations and compare the result with the probability associated with a specific number of successful outcomes computed using the binomial distribution.
# **Step 1:** Write a function to run the binary experiment multiple times to simulate a binomial distribution.
# + id="fIPpO0KvA7BH" colab_type="code" outputId="7de6c3a2-72e5-4f79-9eb2-acb61361f84a" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Define a new function to run multiple binary experiments
def multiple_experiments(experiments=10, trials=100, success=0.5):
# Initialize a list to store the number of successful outcomes in each experiment
n_success = []
# Run through a loop to conduct each experiment and record the outcome
for i in range(experiments):
outcomes = binary_experiment(trials=trials, p_of_success=success)
n_success.append(len([s for s in outcomes if s == 'success']))
# Return the list of the number of successful outcomes in each experiment
return n_success
# Run the new function to test
multiple_experiments()
# + [markdown] id="fpQKY0PRLob2" colab_type="text"
# **Step 2:** Plot up a histogram of the result of running multiple binary experiments and compare with the theoretical result computed using the binomial distribution.
# + id="_UlDgR7JOv7S" colab_type="code" outputId="19fd60c1-6f7e-4a4d-83c2-cc84d3609128" colab={"base_uri": "https://localhost:8080/", "height": 268}
# Define the number of trials, probability of success, and number of experiments to run
num_trials = 100
p_success = 0.9
num_experiments = 1000
# Run binary experiments using the parameters defined above
n_success = multiple_experiments(experiments=num_experiments,
trials=num_trials,
success=p_success)
# Generate x-values to use in calculating a theoretical geometric distribution
x = np.arange(np.min(n_success), np.max(n_success)+1)
# Use the x-values to define bins to be used for a histogram of experimental data
bin_edges = x - 0.5
# Compute and plot a histogram of experimental results (blue bars)
plt.hist(n_success, bins=bin_edges, label='Experimental Result')
# Plot the theoretical result from a binomial distribution (green dots)
plt.plot(x, stats.binom.pmf(x, num_trials, p_success)*num_experiments, 'go', ms=8, label='Binomial Dist.')
plt.legend()
plt.show()
# + [markdown] id="-Aggp7pUV_x4" colab_type="text"
# **Step 3:** Compare the result of running multiple binary experiments with the theoretical result computed using the binomial distribution for a specified range of values.
# + id="DzicQsJGV-8h" colab_type="code" outputId="5b78a295-c59f-4632-f27c-d22486ba4e5e" colab={"base_uri": "https://localhost:8080/", "height": 318}
# Write a function to compare an experimental result for some number of successes
# with the theoretical result from a binomial distribution
def compare_n_successes(n_to_compare=50, comparison='equal', experiments=1000, trials=100, success=0.5):
# Run binary experiments
n_success = multiple_experiments(experiments=experiments,
trials=trials,
success=success)
# Generate x-values to use in calculating a theoretical geometric distribution
x = np.arange(np.min(n_success), np.max(n_success)+1)
# Use the x-values to define bins to be used for a histogram of experimental data
bin_edges = np.append(x-0.5, x[-1]+0.5)
# Compute and plot a histogram of experimental results (blue bars)
hist = plt.hist(n_success, bins=bin_edges)
# Plot the theoretical result from a binomial distribution (green dots)
theoretical_result = stats.binom.pmf(x, trials, success)
plt.plot(x, theoretical_result*experiments, 'go', ms=8, label='Binomial Dist.')
# Check if the number requested for comparison exists in the array of x-values
if n_to_compare not in list(x):
print('The number of successes for comparison is not within the experimental results.')
print(f'Try again with one of these numbers:{x}')
return
# Check to see if the type of comparison requested is valid
compare_options = ['equal', 'less than or equal', 'greater than or equal']
if comparison not in compare_options:
print(f'{comparison} is not an option for comparison')
print(f'Try again with one of these:{compare_options}')
return
# Extract the array of experimental counts
experimental_counts = hist[0]
# Extract the indices for comparison
if comparison == 'equal':
ind = (x == n_to_compare).nonzero()[0]
if comparison == 'less than or equal':
ind = (x <= n_to_compare).nonzero()[0]
if comparison == 'greater than or equal':
ind = (x >= n_to_compare).nonzero()[0]
# Compute the fraction of counts (probability) for that value(s) in the experiment
experimental_probability = np.sum(experimental_counts[ind])/np.sum(experimental_counts)
theoretical_probability = np.sum(theoretical_result[ind])
# Plot the experimental and theoretical comparison
plt.bar(x[ind], experimental_counts[ind], color='r', label='Experimental')
plt.plot(x[ind], theoretical_result[ind]*experiments, color='magenta', marker='o', ms=8, ls='None', label='Theoretical')
plt.legend()
plt.title('Binary Experiment Simulation vs. Binomial Distribution')
plt.show()
# Compute the theoretical probability of that exact result using the binomial distribution
print(f'The experimental probability of n successes {comparison} to {n_to_compare} is {experimental_probability:.3f}')
print(f'The theoretical probability of n successes {comparison} to {n_to_compare} is {theoretical_probability:.3f}')
return
# Run the new function to test
compare_n_successes()
# + [markdown] id="Mxconkbe45y0" colab_type="text"
# **Step 4:** Try running the comparison for different input parameters.
# + id="f9C_HOBMa95L" colab_type="code" outputId="10ffabc8-159b-46bc-9ab9-7f5047356d20" colab={"base_uri": "https://localhost:8080/", "height": 318}
# Define parameters to use in the comparison of simulated vs. theoretical number of successes
n = 12
compare = 'greater than or equal'
n_exp = 1000
n_trials = 100
p_success = 0.1
# Run the experiment
compare_n_successes(n_to_compare=n,
comparison=compare,
experiments=n_exp,
trials=n_trials,
success=p_success)
# + [markdown] id="0a-wjV_BMjtY" colab_type="text"
# ### **Learning objective 3:** Simulate an experiment to find the probability associated with the number of trials to reach the first success and compare with the result computed using the geometric distribution.
# **Step 1:** Write a function to simulate a geometric experiment that records the number of trials to reach the first success.
# + id="v3E3F-sLJ1VR" colab_type="code" outputId="23cb9fd8-d276-4ef2-bfd8-a534dbeb7ce8" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Define a new experiment that simply counts the number of trials to reach the first success
def geometric_experiment(success=0.05):
# Initialize a count variable
count = 1
# Loop through conducting Bernouilli trials until reaching success
while True:
outcome = bernouilli_trial(prob_of_success=success)
# If the outcome is a success, break out of the loop and return the count
if outcome == 'success':
break
# Otherwise add one to the count
else:
count += 1
# Return the final count
return count
# Run the new function to test
geometric_experiment()
# + [markdown] id="niImIxPYNLfe" colab_type="text"
# **Step 2:** Write a function to run multiple geometric experiments and record the results of each.
# + id="fu3W4hJ0IOSd" colab_type="code" outputId="1f963e70-106d-4cd2-a950-e086a087b0cb" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Define a function to perform multiple geometric experiments
def multiple_geometric(experiments=10, success=0.1):
# Initialize a list to hold the count of number of trials to reach success
count_to_success = []
# Loop through running each experiment and recording the result
for i in range(experiments):
count_to_success.append(geometric_experiment(success=success))
# Return the results
return count_to_success
# Run the new function to test
multiple_geometric()
# + [markdown] id="h17aEgDYNXdu" colab_type="text"
# **Step 3:** Plot up a histogram of the result of running multiple geometric experiments and compare with the theoretical result computed using the geometric distribution.
# + id="u9UwOrMfOZzS" colab_type="code" outputId="4e06f399-9a68-48d0-aab3-b61028736843" colab={"base_uri": "https://localhost:8080/", "height": 268}
# Define parameters for the number of experiments to run and probability of success
n_experiments = 1000
p_success = 0.3
# Run geometric experiments with these parameters
count_to_success = multiple_geometric(experiments=n_experiments,
success=p_success)
# Generate x-values to use in calculating a theoretical geometric distribution
x = np.arange(1, np.max(count_to_success)+1)
# Use the x-values to define bins to be used for a histogram of experimental data
bin_edges = x - 0.5
# Compute and plot a histogram of experimental results (blue bars)
plt.hist(count_to_success, bins=bin_edges, label='Experimental Result')
# Plot the theoretical result from a geometric distribution (green dots)
plt.plot(x, stats.geom.pmf(x, p_success)*n_experiments, 'go', ms=8, label='Geometric Dist.')
plt.legend()
plt.show()
# + [markdown] id="BAD2oyG_ybpV" colab_type="text"
# **Step 4:** Compare the result of running multiple binary experiments with the theoretical result computed using the geometric distribution for a specified range of values.
# + id="fTyWcWT1ycQ0" colab_type="code" outputId="5abfd1b5-8a84-4cd0-bbd5-88754c41cd0d" colab={"base_uri": "https://localhost:8080/", "height": 318}
# Write a function to compare an experimental result for some number of successes
# with the theoretical result from a binomial distribution
def compare_trials_to_success(n_to_compare=3, comparison='equal', experiments=1000, success=0.3):
# Run geometric experiments
count_to_success = multiple_geometric(experiments=experiments,
success=success)
# Generate x-values to use in calculating a theoretical geometric distribution
x = np.arange(1, np.max(count_to_success)+1)
# Use the x-values to define bins to be used for a histogram of experimental data
bin_edges = np.append(x-0.5, x[-1]+0.5)
# Compute and plot a histogram of experimental results (blue bars)
hist = plt.hist(count_to_success, bins=bin_edges)
# Plot the theoretical result from a binomial distribution (green dots)
theoretical_result = stats.geom.pmf(x, success)
plt.plot(x, theoretical_result*experiments, 'go', ms=8, label='Geometric Dist.')
# Check if the number requested for comparison exists in the array of x-values
if n_to_compare not in list(x):
print('The number of successes for comparison is not within the experimental results.')
print(f'Try again with one of these numbers:{x}')
return
# Check to see if the type of comparison requested is valid
compare_options = ['equal', 'less than or equal', 'greater than or equal']
if comparison not in compare_options:
print(f'{comparison} is not an option for comparison')
print(f'Try again with one of these:{compare_options}')
return
# Extract the array of experimental counts
experimental_counts = hist[0]
# Extract the indices for comparison
if comparison == 'equal':
ind = (x == n_to_compare).nonzero()[0]
if comparison == 'less than or equal':
ind = (x <= n_to_compare).nonzero()[0]
if comparison == 'greater than or equal':
ind = (x >= n_to_compare).nonzero()[0]
# Compute the fraction of counts (probability) for that value(s) in the experiment
experimental_probability = np.sum(experimental_counts[ind])/np.sum(experimental_counts)
theoretical_probability = np.sum(theoretical_result[ind])
# Plot the experimental and theoretical comparison
plt.bar(x[ind], experimental_counts[ind], color='r', label='Experimental')
plt.plot(x[ind], theoretical_result[ind]*experiments, color='magenta', marker='o', ms=8, ls='None', label='Theoretical')
plt.legend()
plt.title('Simulation vs. Geometric Distribution')
plt.show()
# Compute the theoretical probability of that exact result using the binomial distribution
print(f'The experimental probability of n trials to success {comparison} to {n_to_compare} is {experimental_probability:.3f}')
print(f'The theoretical probability of n trials to success {comparison} to {n_to_compare} is {theoretical_probability:.3f}')
# Run the new function to test
compare_trials_to_success()
# + [markdown] id="oKqXJS9I328s" colab_type="text"
# **Step 5:** Try running the comparison for different input parameters.
# + id="Jz2Hi_S9z5p0" colab_type="code" outputId="bc331d8e-cae2-4971-f008-793ddaef48b3" colab={"base_uri": "https://localhost:8080/", "height": 318}
# Define parameters to use in the comparison of simulated and theoretical number of trials to reach first success
n = 5
compare = 'greater than or equal'
n_exp = 1000
p_success = 0.3
# Run the comparison
compare_trials_to_success(n_to_compare=n,
comparison=compare,
experiments=n_exp,
success=p_success)
# + [markdown] id="ENjWJb84Nnht" colab_type="text"
# ### **Learning objective 4:** Simulate the probability of recording a given number of events in an interval or region and compare the result with the probability computed directly using the Poisson distribution.
# **Step 1:** Write a function to simulate a Poisson experiment that records the number of successes over an interval.
# + id="xW7XJehRPG1E" colab_type="code" outputId="625af9d8-ef8b-4f21-abc7-9980fd983b36" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Define a function to count the number of successful trials in an interval (region)
# In terms of parameter definitions, let's say the event rate lambda is defined for some "baseline" interval or region, then:
### "subinterval_size" is the fraction of that baseline interval over which to conduct each trial
### "n_subintervals" is the number of subintervals (trials) to conduct
### "poisson_lambda" is the average number of successes or event rate over that baseline interval
def poisson_experiment(subinterval_size=1/60, n_subintervals=60, poisson_lambda=10):
# Calculate the probability of success for any given subinterval
# by multiplying the average number of successes per interval by the number of subintervals
prob_per_subinterval = poisson_lambda*subinterval_size
# Run a binary experiment recording outcomes for each trial
outcomes = binary_experiment(trials=n_subintervals,
p_of_success=prob_per_subinterval)
# Count the number of successful outcomes
n_success = len([s for s in outcomes if s == 'success'])
# Return the result
return n_success
# Run the new function to test
poisson_experiment()
# + [markdown] id="GIiLI1oJOVx0" colab_type="text"
# **Step 2:** Write a function to run multiple Poisson experiments and record the results of each.
# + id="jlQE4R-hPsIe" colab_type="code" outputId="8dbae75f-a70a-40aa-b808-55370825e11e" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Define a function to run multiple Poisson experiments
def multiple_poisson(experiments=10, subinterval_size=1/60, n_subintervals=60, poisson_lambda=10):
# Initialize a list to hold the result of each experiment
n_successes = []
# Loop through running each experiment and recording the result
for i in range(experiments):
n_success = poisson_experiment(subinterval_size=subinterval_size,
n_subintervals=n_subintervals,
poisson_lambda=poisson_lambda)
# Append each result to the list
n_successes.append(n_success)
# Return the result
return n_successes
# Run the new function to test
multiple_poisson()
# + [markdown] id="VMKhbcCAOcj3" colab_type="text"
# **Step 3:** Plot up a histogram of the result of running multiple Poisson experiments and compare with the theoretical result computed using the Poisson distribution.
# + id="txbJI0gLT_tn" colab_type="code" outputId="127ca49d-f45c-4eaa-eae3-272ad61c76b5" colab={"base_uri": "https://localhost:8080/", "height": 268}
# Define parameters for the number of experiments to run, number of subintervals and successes per interval
n_experiments=1000
sub_size=1/60
n_subs=60
p_lambda=3
# Run the experiments and record the result
n_successes = multiple_poisson(experiments=n_experiments,
subinterval_size=sub_size,
n_subintervals=n_subs,
poisson_lambda=p_lambda)
# Generate x-values to use in calculating a theoretical Poisson distribution
x = np.arange(np.min(n_successes), np.max(n_successes)+1)
# Use the x-values to define bins to be used for a histogram of experimental data
bin_edges = x - 0.5
# Compute and plot a histogram of experimental results (blue bars)
plt.hist(n_successes, bins=bin_edges, label='Experimental Result')
# Plot the theoretical result from a Poisson distribution (green dots)
plt.plot(x, stats.poisson.pmf(x, n_subs*sub_size*p_lambda)*n_experiments, 'go', ms=8, label='Poisson Dist.')
plt.legend()
plt.show()
# + [markdown] id="bLX-9I0A4aYT" colab_type="text"
# **Step 4:** Compare the result of running multiple Poisson experiments with the theoretical result computed using the Poisson distribution for a specified range of values.
# + id="krdBk3V3R9ZX" colab_type="code" outputId="6f44ceed-5bfb-4b55-f1b8-868a8260ce88" colab={"base_uri": "https://localhost:8080/", "height": 318}
# Write a function to compare an experimental result for some number of successes
# with the theoretical result from a binomial distribution
def compare_n_over_interval(n_to_compare=10, comparison='equal', experiments=1000,
subinterval_size=1/60,
n_subintervals=60,
poisson_lambda=10):
# Run the experiments
n_successes = multiple_poisson(experiments=experiments,
subinterval_size=subinterval_size,
n_subintervals=n_subintervals,
poisson_lambda=poisson_lambda)
# Generate x-values to use in calculating a theoretical geometric distribution
x = np.arange(np.min(n_successes), np.max(n_successes)+1)
# Use the x-values to define bins to be used for a histogram of experimental data
bin_edges = np.append(x-0.5, x[-1]+0.5)
# Compute and plot a histogram of experimental results (blue bars)
hist = plt.hist(n_successes, bins=bin_edges)
# Plot the theoretical result from a binomial distribution (green dots)
theoretical_result = stats.poisson.pmf(x, n_subintervals*subinterval_size*poisson_lambda)
plt.plot(x, theoretical_result*experiments, 'go', ms=8, label='Poisson Dist.')
# Check if the number requested for comparison exists in the array of x-values
if n_to_compare not in list(x):
print('The number of successes for comparison is not within the experimental results.')
print(f'Try again with one of these numbers:{x}')
return
# Check to see if the type of comparison requested is valid
compare_options = ['equal', 'less than or equal', 'greater than or equal']
if comparison not in compare_options:
print(f'{comparison} is not an option for comparison')
print(f'Try again with one of these:{compare_options}')
return
# Extract the array of experimental counts
experimental_counts = hist[0]
# Extract the indices for comparison
if comparison == 'equal':
ind = (x == n_to_compare).nonzero()[0]
if comparison == 'less than or equal':
ind = (x <= n_to_compare).nonzero()[0]
if comparison == 'greater than or equal':
ind = (x >= n_to_compare).nonzero()[0]
# Compute the fraction of counts (probability) for that value(s) in the experiment
experimental_probability = np.sum(experimental_counts[ind])/np.sum(experimental_counts)
theoretical_probability = np.sum(theoretical_result[ind])
# Plot the experimental and theoretical comparison
plt.bar(x[ind], experimental_counts[ind], color='r', label='Experimental')
plt.plot(x[ind], theoretical_result[ind]*experiments, color='magenta', marker='o', ms=8, ls='None', label='Theoretical')
plt.legend()
plt.title('Simulation vs. Poisson Distribution')
plt.show()
# Compute the theoretical probability of that exact result using the binomial distribution
print(f'The experimental probability of n events in the interval being {comparison} to {n_to_compare} is {experimental_probability:.3f}')
print(f'The theoretical probability of n events in the interval being {comparison} to {n_to_compare} is {theoretical_probability:.3f}')
return
# Run the new function to test
compare_n_over_interval()
# + [markdown] id="XS-gA7ns4oYx" colab_type="text"
# **Step 5:** Try running the comparison for different input parameters.
# + id="TakhPV2Z2ah1" colab_type="code" outputId="87fa10af-0cc9-4fdb-d3f4-8a31395d0346" colab={"base_uri": "https://localhost:8080/", "height": 318}
# Define parameters to use in the comparison of simulated and theoretical number of events per interval
n = 2
compare = 'less than or equal'
n_exp = 1000
sub_size = 1/60
n_subs = 60
p_lambda = 3
# Run the comparison
compare_n_over_interval(n_to_compare=n,
comparison=compare,
experiments=n_exp,
subinterval_size=sub_size,
n_subintervals=n_subs,
poisson_lambda=p_lambda)
# + id="Uhm5IUS43HHW" colab_type="code" colab={}
| Probability_Distributions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import glob
import numpy as np
import pandas as pd
import vaex
from astropy.coordinates import SkyCoord
from astropy.time import Time
from m33_utils import open_and_select
# +
df = pd.read_csv('../header_info.csv', index_col=0)
df['FILT'] = df['FILT_DET'].str.split('-').str[1]
coo = SkyCoord(df[['RAC','DECC']].values, frame='icrs', unit='deg')
df['COO_STR'] = coo.to_string('hmsdms', format='latex', precision=0)
df['RA_STR'] = df['COO_STR'].str.split().str[0]
df['DEC_STR'] = df['COO_STR'].str.split().str[1]
t = Time(df.EXPSTART, format='mjd')
t.format = 'iso'
t.precision = 0
df['EXPDATE'] = t.value
df['EXPT'] = df['EXPTIME'].round(2)
df['ORIENT'] = df['ORIENTAT'].round(4)
cols = ['TARGNAME','RA_STR','DEC_STR','EXPDATE','EXPT','INSTRUME','APERTURE','FILT','ORIENT']
table_obs = df[cols].sort_values(by=['TARGNAME','EXPDATE'])
table_obs.to_csv('tables_full/table_obs.csv', index=False)
with open('tables/table_obs.tex', 'w') as f:
f.write(table_obs[table_obs.TARGNAME.str.startswith('M33-B01-F01')
].to_latex(index=False, escape=False))
# +
ds = vaex.open('../legacy_phot/M33_full_matched.hdf5')
filters = ['F275W','F336W','F475W','F814W','F110W','F160W']
for f in filters:
gst_tf = ds[f'{f}_GST_FLAG'].astype(str).str.replace('0','F').str.replace('1','T')
ds.add_virtual_column(f'{f}_GST', gst_tf)
cols = ['RA', 'DEC']
cols += [f'{f}_{q}' for f in filters for q in ['VEGA','SNR','GST']]
ds_phot = ds[cols].fillna(99.999)
for c in ds_phot.get_column_names(regex='.*?((VEGA)|(SNR))'):
ds_phot[c] = ds_phot[c].apply(lambda x: np.round(x, 3))
ds_phot.export_csv('tables_full/table_phot.csv')
# df_phot = pd.read_csv('tables/table_phot.csv', index_col=False)
df_phot = ds_phot[:10].to_pandas_df()
df_phot.rename(lambda x: x.split('_')[0] if x.endswith('VEGA') else x.split('_')[-1].replace('SNR','S/N'),
axis='columns', inplace=True)
with open('tables/table_phot.tex', 'w') as f:
f.write(df_phot.to_latex(index=False))
# ds.close_files()
# +
# img1:img30-wfc3/ir, img31:img70-wfc3/uvis, img71:-acs/wfc,
df_par = pd.read_csv('../M33_B03_SW/M33_B03_SW_3.param', sep='=', comment='#',
index_col=0, names=['Parameter','Value'])
df_par.index = df_par.index.str.strip()
df_par['Value'] = df_par['Value'].astype(str).str.strip()
df_par.rename(lambda x: x.replace('img1_', 'IR_'), axis='rows', inplace=True)
df_par.rename(lambda x: x.replace('img31_', 'UVIS_'), axis='rows', inplace=True)
df_par.rename(lambda x: x.replace('img71_', 'WFC_'), axis='rows', inplace=True)
df_par.drop(df_par.filter(regex='img.*?_', axis='rows').index, inplace=True)
df_par.drop(df_par.filter(regex='_file', axis='rows').index, inplace=True)
df_par.drop(['Nimg', 'xytfile', 'xytpsf', 'psfstars', 'UsePhot', 'photsec'], inplace=True)
df_par = df_par.assign(Detector='All')
df_par.loc[df_par.filter(regex='(^IR_)|(^WFC3IR)', axis='rows').index, 'Detector'] = 'IR'
df_par.loc[df_par.filter(regex='(^UVIS_)|(^WFC3UVIS)', axis='rows').index, 'Detector'] = 'UVIS'
df_par.loc[df_par.filter(regex='(^WFC_)|(^ACS)', axis='rows').index, 'Detector'] = 'WFC'
df_par.loc['WFC3useCTE', 'Detector'] = 'UVIS/IR'
df_par.reset_index(inplace=True)
df_par.loc[15, ['Detector', 'Parameter']] = ['All', 'apsky']
df_par.drop([16,17], inplace=True)
df_par['Parameter'] = df_par['Parameter'].str.replace('IR_','').str.replace('UVIS_','').str.replace('WFC_','')
with open('tables/table_par.tex', 'w') as f:
f.write(df_par[['Detector','Parameter','Value']].to_latex(index=False))
# +
def read_artstars(infile):
ds = open_and_select(infile)
ds['DPOS'] = ds['((X_OUT-X_IN)**2 + (Y_OUT-Y_IN)**2)**0.5']
for f in ds.get_column_names(regex='F.*?W_IN'):
filt = f.split('_')[0]
ds.add_column(f'{filt}_DELTA', ds[f'{filt}_VEGA-{filt}_IN'].evaluate())
selection = f'({filt}_GST) & (DPOS < 2) & (abs({filt}_DELTA) < 1)'
ds.select(selection, name=f'{filt}_detected')
is_detected = ds.evaluate_selection_mask(f'{filt}_detected').astype(int)
ds.add_column(f'{filt}_DET', is_detected)
return ds
ds = read_artstars(glob.glob('../artstars/M33_B0?_??_fake_final.hdf5')).sort(by='RA')
filters = ['F275W','F336W','F475W','F814W','F110W','F160W']
for f in filters:
gst_tf = pd.Series(ds.evaluate_selection_mask(f'{f}_GST').astype(int)
).astype(str).str.replace('0','F').str.replace('1','T')
ds.add_column(f'{f}_DT', gst_tf.values)
cols = ['RA', 'DEC']
cols += [f'{f}_{q}' for f in filters for q in ['IN', 'DELTA', 'SNR', 'DT']]
ds_ast = ds[cols].fillna(99.999)
for c in ds_ast.get_column_names(regex='.*?((VEGA)|(SNR)|(DELTA)|(IN))'):
ds_ast[c] = ds_ast[c].apply(lambda x: np.round(x, 3))
ds_ast.export_csv('tables_full/table_asts.csv')
df_ast = ds_ast[:10].to_pandas_df()
def rename(x):
if x.endswith('IN'):
name = x.split('_')[0] + ' in'
elif x.endswith('DELTA'):
name = 'Out - in'
elif x.endswith('DT'):
name = 'GST'
else:
name = x.split('_')[-1].replace('SNR','S/N')
return name
df_ast.rename(rename, axis='columns', inplace=True)
with open('tables/table_asts.tex', 'w') as f:
f.write(df_ast.to_latex(index=False))
ds.close_files()
# -
| Make tex tables.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Numba 0.47.0 Release Demo
# =======================
#
# This notebook contains a demonstration of new features present in the 0.47.0 release of Numba. Whilst release notes are produced as part of the [`CHANGE_LOG`](<UPDATE ME>), there's nothing like seeing code in action! This release contains a large number of exciting new features!
#
# Demonstrations of new features include:
# * [Bounds checking](#Bounds-checking)
# * [Dynamic function definition](#Dynamic-function-definition)
# * [Support for `map`, `filter` and `reduce`](#Support-for-map,-filter,-reduce)
# * [Support for `list.sort()` and `sorted` with a `key`](#Support-for-list.sort()/sorted-with-key)
# * [Initial support for `try/except`](#Initial-support-for-basic-try/except)
# * [Iteration over mixed type containers](#Iterating-over-mixed-type-containers)
# * [New NumPy function support](#Newly-supported-NumPy-functions/features)
# * [New unicode string features](#New-unicode-string-features)
#
# First, import the necessary from Numba and NumPy...
from numba import jit, njit, config, __version__, errors
from numba.errors import NumbaPendingDeprecationWarning
import warnings
# we're going to ignore a couple of deprecation warnings
warnings.simplefilter('ignore', category=NumbaPendingDeprecationWarning)
from numba.extending import overload
config.SHOW_HELP = 0
import numba
import numpy as np
assert tuple(int(x) for x in __version__.split('.')[:2]) >= (0, 47)
# Bounds checking
# ==============
#
# The long awaited support for bounds checking has been added in this release, the associated documentation is available [here](https://numba.pydata.org/numba-doc/latest/reference/jit-compilation.html). Here's a demonstration:
# +
config.FULL_TRACEBACKS = 1
@njit(boundscheck=True)
def OOB_access(x):
sz = len(x)
a = x[0] # fine, first element of x
a += x[sz - 1] # fine, last element of x
a += x[sz] # oops, out of bounds!
try:
OOB_access(np.ones(10))
except IndexError as e:
print(type(e), e)
# -
# The setting of `config.FULL_TRACEBACKS` ([environment variable equivalent](https://numba.pydata.org/numba-doc/latest/reference/envvars.html#envvar-NUMBA_BOUNDSCHECK)) forces the printing of the index, axis and dimension size to the terminal (assuming a terminal was used to invoke python). For example, the terminal that launched this notebook now has:
# ```
# debug: IndexError: index 10 is out of bounds for axis 0 with size 10
# ```
# on it. A future release will enhance this feature to include the out of bounds access information in the error message.
# Dynamic function definition
# =======================
#
# The 0.47.0 release adds the following new capability to Numba: dynamic function generation. Essentially functions (closures) defined in a JIT decorated function can now "escape" the function they are defined in and be used as arguments in subsequent function calls. For example:
# +
# takes a function and calls it with argument arg, multiplies the result by 7
@njit
def consumer(function, arg):
return function(arg) * 7
_GLOBAL = 5
@njit
def generator_func():
_FREEVAR = 10
def escapee(x): # closure, 'a' is a local, '_FREEVAR' is a freevar, '_GLOBAL' is global
a = 9
return x * _FREEVAR + a * _GLOBAL
# data argument for the consumer call
x = np.arange(5)
# escapee function is passed to the consumer function along with its argument
return consumer(escapee, x)
generator_func()
# -
# Support for `map`, `filter`, `reduce`
# ============================
#
# The ability to create dynamic functions lead to being able to write support for `map`, `filter` and `reduce`. This makes it possible to write more "pythonic" code in Numba :-)
# +
import operator
from functools import reduce
from numba.typed import List
@njit
def demo_map_filter_reduce():
# This will be used in map
def mul_n(x, multiplier):
return x * multiplier
# This will be used in filter
V = 20
def greater_than_V(x):
return x > V # captures V from freevars
# this will be used in reduce
reduce_lambda = lambda x, y: (x * 2) + y
a = [x ** 2 for x in range(10)]
n = len(a)
return reduce(reduce_lambda, filter(greater_than_V, map(mul_n, a, range(n))))
demo_map_filter_reduce()
# -
# Support for `list.sort()`/`sorted` with key
# =================================
#
# A further extension born from the ability to create dynamic functions was being able to support the ``key`` argument to `list.sort` and `sorted`, a quick demonstration:
# +
@njit
def demo_sort_sorted(chars):
def key(x):
return x.upper()
x = chars[:]
x.sort()
print("sorted:", ''.join(x))
x = chars[:]
x.sort(reverse=True)
print("sorted backwards:", ''.join(x))
x = chars[:]
x.sort(key=key)
print("sorted key=x.upper():", ''.join(x))
print("sorted(), reversed", ''.join(sorted(x, reverse=True)))
def numba_order(x):
return 'NUMBA🐍numba⚡'.index(x)
x = chars[:]
x.sort(key=numba_order)
print("sorted key=numba_order:", ''.join(x))
# let's sort a list of characters
input_list = ['m','M','a','N','n','u','⚡','🐍','B','b','U','A']
demo_sort_sorted(input_list)
# -
# Initial support for basic try/except
# ============================
#
# Numba 0.47.0 has some basic support for the use of `try`/`except` in JIT compiled functions. This is a long awaited feature that has been requested many times. Support is limited at present to two use cases [docs](https://numba.pydata.org/numba-doc/latest/reference/pysupported.html#try-except).
# +
@njit
def demo_try_bare_except(a, b):
try:
c = a / b
return c
except:
print("caught exception")
return -1
print("ok input:", demo_try_bare_except(5., 10.))
print("div by zero input:", demo_try_bare_except(5, 0))
# -
# The class `Exception` can also be caught, let's mix this with the new bounds checking support:
# +
@njit(boundscheck=True)
def demo_try_except_exception(array, index):
try:
return array[index]
except Exception:
print("caught exception")
return -1
x = np.ones(5)
print("ok input:", demo_try_except_exception(x, 0))
print("OOB access:", demo_try_except_exception(x, 10))
# -
# User defined exception classes also work:
# +
class UserDefinedException(Exception):
def __init__(self, some_arg):
self._some_arg = some_arg
@njit(boundscheck=True)
def demo_try_except_ude():
try:
raise UserDefinedException(123)
except Exception:
return "caught UDE!"
print(demo_try_except_ude())
# -
# Iterating over mixed type containers
# =============================
#
# As users of Numba are very aware, Numba has to be able to work out the type of all the variables in a function to be able to compile it (function must be statically typable!). Prior to Numba 0.47.0 tuples of heterogeneous type could not be iterated over as the type of the induction variable in a loop could not be statically computed and further the loop body contents would have a different set of types of each type in the tuple. For example, this doesn't work:
# +
from numba import literal_unroll
@njit
def does_not_work():
tup = (1, 'a', 2j)
for i in tup:
print(i) # Numba cannot work out type of `i`, it changes each loop iteration
print("Typing problem")
try:
does_not_work()
except errors.TypingError as e:
print(e)
# -
# In Numba 0.47.0 a new function, `numba.literal_unroll`, is introduced. The function itself does nothing much, it's just a token to tell the Numba compiler that the argument needs special treatment for use as an iterable. When this function is applied in situations like in the following, the body of the loop is "versioned" based on the types in the tuple such that Numba can actually statically work out the types for each iteration and compilation will succeed. Here's a working version of the above failing example:
# +
# use special function `numba.literal_unroll`
@njit
def works():
tup = (1, 'a', 2j)
for i in literal_unroll(tup):
print(i) # literal_unroll tells the compiler to version the loop body based on type.
print("Apply literal_unroll():")
works()
# -
# A more involved example might be a tuple of locally defined functions (which are all different types by virtue of the Numba type system) that are iterated over:
# +
@njit
def fruit_cookbook():
def get_apples(x):
return ['apple' for _ in range(x * 3)]
def get_oranges(x):
return ['orange' for _ in range(x * 4)]
def get_bananas(x):
return ['banana' for _ in range(x * 2)]
ingredients = (get_apples, get_oranges, get_bananas)
def fruit_salad(scale):
shopping_list = []
for ingredient in literal_unroll(ingredients):
shopping_list.extend(ingredient(scale))
return shopping_list
print(fruit_salad(2))
fruit_cookbook()
# -
# Finally, because Numba has string and integer literal support, it's possible to dispatch on these values at compile time and version the loop body with a value based specialisations:
# +
from numba import types
# function stub to overload
def dt(value):
pass
@overload(dt, inline='always')
def ol_dt(li):
# dispatch based on a string literal
if isinstance(li, types.StringLiteral):
value = li.literal_value
if value == "apple":
def impl(li):
return 1
elif value == "orange":
def impl(li):
return 2
elif value == "banana":
def impl(li):
return 3
return impl
# dispatch based on an integer literal
elif isinstance(li, types.IntegerLiteral):
value = li.literal_value
if value == 0xca11ab1e:
def impl(li):
# close over the dispatcher :)
return 0x5ca1ab1e + value
return impl
@njit
def unroll_and_dispatch_on_literal():
acc = 0
for t in literal_unroll(('apple', 'orange', 'banana', 0xca11ab1e)):
acc += dt(t)
return acc
print(unroll_and_dispatch_on_literal())
# -
# It's hoped that in a future version of Numba the token function `literal_unroll` will not be needed and loop body versioning opportunities will be automatically identified.
# Newly supported NumPy functions/features
# =====
# This release contains a number of newly supported NumPy functions, all written by contributors from the Numba community:
#
# * `np.arange` now supports the `dtype` keyword argument.
#
# * Also now supported are:
# * `np.lcm`
# * `np.gcd`
#
# A quick demo of the above:
# +
@njit
def demo_numpy():
a = np.arange(5, dtype=np.uint8)
b = np.lcm(a, 2)
c = np.gcd(a, 3)
return a, b, c
demo_numpy()
# -
# New unicode string features
# =======================
#
# A large number of unicode string features/enhancements were added in 0.47.0, namely:
#
# * `str.index()`
# * `str.rindex()`
# * `start/end` parameters for `str.find()`
# * `str.rpartition()`
# * `str.lower()`
#
# and a lot of querying functions:
# * `str.isalnum()`
# * `str.isalpha()`
# * `str.isascii()`
# * `str.isidentifier()`
# * `str.islower()`
# * `str.isprintable()`
# * `str.isspace()`
# * `str.istitle()`
#
# +
@njit
def demo_string_enhancements(arg):
print("index:", arg.index("🐍")) # index of snake
print("rindex:", arg.rindex("🐍")) # rindex of snake
print("find:", arg.find("🐍", start=2, end=6)) # find snake with start+end
print("rpartition:", arg.rpartition("🐍")) # rpartition snake
print("lower:", arg.lower()) # lower snake
print("isalnum:", 'abc123'.isalnum(), '🐍'.isalnum())
print("isalpha:", 'abc'.isalpha(), '123'.isalpha())
print("isascii:", 'abc'.isascii(), '🐍'.isascii())
print("isidentifier:", '1'.isidentifier(), 'var'.isidentifier())
print("islower:", 'SHOUT'.islower(), 'whisper'.islower())
print("isprintable:", '\x07'.isprintable(), 'BEL'.isprintable())
print("isspace:", ' '.isspace(), '_'.isspace())
print("istitle:", "Titlestring".istitle(), "notTitlestring".istitle())
arg = "N🐍u🐍M🐍b🐍A⚡"
demo_string_enhancements(arg)
| notebooks/Numba_047_release_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Semantic Segmentation Demo
#
# This is a notebook for running the benchmark semantic segmentation network from the the [ADE20K MIT Scene Parsing Benchchmark](http://sceneparsing.csail.mit.edu/).
#
# The code for this notebook is available here
# https://github.com/CSAILVision/semantic-segmentation-pytorch/tree/master/notebooks
#
# It can be run on Colab at this URL https://colab.research.google.com/github/CSAILVision/semantic-segmentation-pytorch/blob/master/notebooks/DemoSegmenter.ipynb
# ### Environment Setup
#
# First, download the code and pretrained models if we are on colab.
'''
%%bash
# Colab-specific setup
!(stat -t /usr/local/lib/*/dist-packages/google/colab > /dev/null 2>&1) && exit
pip install yacs 2>&1 >> install.log
git init 2>&1 >> install.log
git remote add origin https://github.com/CSAILVision/semantic-segmentation-pytorch.git 2>> install.log
git pull origin master 2>&1 >> install.log
DOWNLOAD_ONLY=1 ./demo_test.sh 2>> install.log
'''
print()
# ## Imports and utility functions
#
# We need pytorch, numpy, and the code for the segmentation model. And some utilities for visualizing the data.
# +
import cv2
# System libs
import os, csv, torch, numpy, scipy.io, PIL.Image, torchvision.transforms
# Our libs
from mit_semseg.models import ModelBuilder, SegmentationModule
from mit_semseg.utils import colorEncode, unique
colors = scipy.io.loadmat('data/color150.mat')['colors']
names = {}
with open('data/object150_info.csv') as f:
reader = csv.reader(f)
next(reader)
for row in reader:
names[int(row[0])] = row[5].split(";")[0]
def visualize_result(img, pred, index=None, show=True):
# filter prediction class if requested
if index is not None:
pred = pred.copy()
pred[pred != index] = -1
print(f'{names[index+1]}:')
# colorize prediction
pred_color = colorEncode(pred, colors).astype(numpy.uint8)
# aggregate images and save
im_vis = numpy.concatenate((img, pred_color), axis=1)
if show==True:
display(PIL.Image.fromarray(im_vis))
else:
return pred_color, im_vis
# -
# ## Loading the segmentation model
#
# Here we load a pretrained segmentation model. Like any pytorch model, we can call it like a function, or examine the parameters in all the layers.
#
# After loading, we put it on the GPU. And since we are doing inference, not training, we put the model in eval mode.
# +
# Network Builders
net_encoder = ModelBuilder.build_encoder(
arch='resnet50dilated',
fc_dim=2048,
weights='ckpt/ade20k-resnet50dilated-ppm_deepsup/encoder_epoch_20.pth')
net_decoder = ModelBuilder.build_decoder(
arch='ppm_deepsup',
fc_dim=2048,
num_class=150,
weights='ckpt/ade20k-resnet50dilated-ppm_deepsup/decoder_epoch_20.pth',
use_softmax=True)
crit = torch.nn.NLLLoss(ignore_index=-1)
segmentation_module = SegmentationModule(net_encoder, net_decoder, crit)
segmentation_module.eval()
segmentation_module.cuda()
# -
# ## Load test data
#
# Now we load and normalize a single test image. Here we use the commonplace convention of normalizing the image to a scale for which the RGB values of a large photo dataset would have zero mean and unit standard deviation. (These numbers come from the imagenet dataset.) With this normalization, the limiiting ranges of RGB values are within about (-2.2 to +2.7).
def process_img(path=None, frame=None):
# Load and normalize one image as a singleton tensor batch
pil_to_tensor = torchvision.transforms.Compose([
torchvision.transforms.ToTensor(),
torchvision.transforms.Normalize(
mean=[0.485, 0.456, 0.406], # These are RGB mean+std values
std=[0.229, 0.224, 0.225]) # across a large photo dataset.
])
# pil_image = PIL.Image.open('../ADE_val_00001519.jpg').convert('RGB')
if path!=None:
pil_image = PIL.Image.open(path).convert('RGB')
else:
pil_image = PIL.Image.fromarray(frame)
img_original = numpy.array(pil_image)
img_data = pil_to_tensor(pil_image)
singleton_batch = {'img_data': img_data[None].cuda()}
# singleton_batch = {'img_data': img_data[None]}
output_size = img_data.shape[1:]
return img_original, singleton_batch, output_size
# img_original, singleton_batch, output_size = process_img('../ADE_val_00001519.jpg')
img_original, singleton_batch, output_size = process_img("/home/zyang/Downloads/car_detection_sample1.png")
display(PIL.Image.fromarray(img_original))
# ## Transparent_overlays function
# +
import cv2
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
def transparent_overlays(image, annotation, alpha=0.5):
img1 = image.copy()
img2 = annotation.copy()
# I want to put logo on top-left corner, So I create a ROI
rows,cols,channels = img2.shape
roi = img1[0:rows, 0:cols ]
# Now create a mask of logo and create its inverse mask also
img2gray = cv2.cvtColor(img2,cv2.COLOR_BGR2GRAY)
ret, mask = cv2.threshold(img2gray, 10, 255, cv2.THRESH_BINARY)
mask_inv = cv2.bitwise_not(mask)
# Now black-out the area of logo in ROI
# img1_bg = cv2.bitwise_and(roi,roi,mask = mask_inv)
# Take only region of logo from logo image.
img2_fg = cv2.bitwise_and(img2,img2,mask = mask)
# Put logo in ROI and modify the main image
# dst = cv2.add(img1_bg, img2_fg)
dst = cv2.addWeighted(image.copy(), 1-alpha, img2_fg, alpha, 0)
img1[0:rows, 0:cols ] = dst
return dst
# -
# ## Run the Model
#
# Finally we just pass the test image to the segmentation model.
#
# The segmentation model is coded as a function that takes a dictionary as input, because it wants to know both the input batch image data as well as the desired output segmentation resolution. We ask for full resolution output.
#
# Then we use the previously-defined visualize_result function to render the segmentation map.
def predict_img(segmentation_module, singleton_batch, output_size):
# Run the segmentation at the highest resolution.
with torch.no_grad():
scores = segmentation_module(singleton_batch, segSize=output_size)
# Get the predicted scores for each pixel
_, pred = torch.max(scores, dim=1)
pred = pred.cpu()[0].numpy()
return pred
pred = predict_img(segmentation_module, singleton_batch, output_size)
pred_color, im_vis = visualize_result(img_original, pred, show=False)
display(PIL.Image.fromarray(im_vis))
dst = transparent_overlays(img_original, pred_color)
display(PIL.Image.fromarray(dst))
# ## Append color palette
#
# To see which colors are which, here we visualize individual classes, one at a time.
# Top classes in answer
predicted_classes = numpy.bincount(pred.flatten()).argsort()[::-1]
for c in predicted_classes[:15]:
# visualize_result(img_original, pred, c)
pass
# +
pred = np.int32(pred)
pixs = pred.size
uniques, counts = np.unique(pred, return_counts=True)
#print("Predictions in [{}]:".format(info))
for idx in np.argsort(counts)[::-1]:
name = names[uniques[idx] + 1]
ratio = counts[idx] / pixs * 100
if ratio > 0.1:
print("{} {}: {:.2f}% {}".format(uniques[idx]+1, name, ratio, colors[uniques[idx]]))
# -
def get_color_palette(pred, bar_height):
pred = np.int32(pred)
pixs = pred.size
top_left_y = 0
bottom_right_y = 30
uniques, counts = np.unique(pred, return_counts=True)
# Create a black image
# bar_height = im_vis.shape[0]
img = np.zeros((bar_height,250,3), np.uint8)
for idx in np.argsort(counts)[::-1]:
color_index = uniques[idx]
name = names[color_index + 1]
ratio = counts[idx] / pixs * 100
if ratio > 0.1:
print("{} {}: {:.2f}% {}".format(color_index+1, name, ratio, colors[color_index]))
img = cv2.rectangle(img, (0,top_left_y), (250,bottom_right_y),
(int(colors[color_index][0]),int(colors[color_index][1]),int(colors[color_index][2])), -1)
img = cv2.putText(img, "{}: {:.3f}%".format(name, ratio), (0,top_left_y+20), 5, 1, (255,255,255), 2, cv2.LINE_AA)
top_left_y+=30
bottom_right_y+=30
return img
img = get_color_palette(pred, im_vis.shape[0])
display(PIL.Image.fromarray(img))
# img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
# display(PIL.Image.fromarray(img))
pred_color_palette = numpy.concatenate((pred_color, img), axis=1)
pred_color_palette_dst = numpy.concatenate((dst, img), axis=1)
pred_color_palette_all = numpy.concatenate((im_vis, img), axis=1)
display(PIL.Image.fromarray(pred_color))
type(pred_color)
cv2.imwrite("pred_color.png",cv2.cvtColor(pred_color, cv2.COLOR_RGB2BGR))
display(PIL.Image.fromarray(pred_color_palette))
display(PIL.Image.fromarray(pred_color_palette_all))
display(PIL.Image.fromarray(pred_color_palette_dst))
| notebooks/DemoSegmenter_image.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # 使用兼容 NetworkX 的 API 进行图操作
#
# GraphScope 支持使用兼容 NetworkX 的 API 进行图操作。
# 本次教程参考了 [tutorial in NetworkX](https://networkx.org/documentation/stable/tutorial.html) 的组织方式来介绍这些 API。
#
# +
# Install graphscope package if you are NOT in the Playground
# !pip3 install graphscope
# +
# Import the graphscope and graphscope networkx module.
import graphscope
import graphscope.nx as nx
graphscope.set_option(show_log=True) # enable logging
# -
# ## 创建图
#
# 创建一个空图,只需要简单地创建一个 Graph 对象。
G = nx.Graph()
# ## 点
#
# 图 `G` 可以通过多种方式进行扩充。 在 graphscope.nx 中, 支持一些可哈希的 Python object 作为图的点, 其中包括 int,str,float,tuple,bool 对象。
# 首先,我们从空图和简单的图操作开始,如下所示,你可以一次增加一个顶点,
G.add_node(1)
# 也可以从任何[可迭代](https://docs.python.org/3/glossary.html#term-iterable)的容器中增加顶点,例如一个列表
G.add_nodes_from([2, 3])
#
#
# 你也可以通过格式为`(node, node_attribute_dict)`的二元组的容器, 将点的属性和点一起添加,如下所示:
#
# 点属性我们将在后面进行讨论。
#
G.add_nodes_from([
(4, {"color": "red"}),
(5, {"color": "green"}),
])
# 一个图的节点也可以直接添加到另一个图中:
H = nx.path_graph(10)
G.add_nodes_from(H)
# 经过上面的操作后,现在图 `G` 中包含了图 `H` 的节点。
list(G.nodes)
list(G.nodes.data()) # shows the node attributes
# ## 边
#
# 图 `G` 也可以一次增加一条边来进行扩充,
G.add_edge(1, 2)
e = (2, 3)
G.add_edge(*e) # unpack edge tuple*
list(G.edges)
# 或者通过一次增加包含多条边的list,
G.add_edges_from([(1, 2), (1, 3)])
list(G.edges)
#
#
# 或者通过增加任意 `ebunch` 的边。 *ebunch* 表示任意一个可迭代的边元组的容器。一个边元组可以是一个只包含首尾两个顶点的二元组,例如 `(1, 3)` ,或者一个包含顶点和边属性字典的三元组,例如 `(2, 3, {'weight': 3.1415})`。
#
# 边属性我们会在后面进行讨论
G.add_edges_from([(2, 3, {'weight': 3.1415})])
list(G.edges.data()) # shows the edge arrtibutes
G.add_edges_from(H.edges)
list(G.edges)
#
# 用户也可以通过 `.update(nodes, edges)` 同时增加点和边
G.update(edges=[(10, 11), (11, 12)], nodes=[10, 11, 12])
list(G.nodes)
list(G.edges)
#
#
# 当增加已存在的点或边时,这些点和边会被忽略,不会产生报错。如下所示,在去除掉所有的点和边之后,
G.clear()
#
# 这里我们增加点和边,graphscope.nx 会忽略掉已经存在的点和边。
G.add_edges_from([(1, 2), (1, 3)])
G.add_node(1)
G.add_edge(1, 2)
G.add_node("spam") # adds node "spam"
G.add_nodes_from("spam") # adds 4 nodes: 's', 'p', 'a', 'm'
G.add_edge(3, 'm')
#
# 目前图 `G` 共包含8个顶点和3条边,可以使用如下所示方法进行查看:
G.number_of_nodes()
G.number_of_edges()
#
#
# ## 查看图的元素
#
# 我们可以查看图的顶点和边。可以使用四种基本的图属性来查看图元素:`G.nodes`,`G.edges`,`G.adj` 和 `G.degree`。这些属性都是 `set-like` 的视图,分别表示图中点,边,点邻居和度数。这些接口提供了一个只读的关于图结构的视图。这些视图也可以像字典一样,用户可以查看点和边的属性,然后通过方法 `.items()`,`.data('span')` 遍历数据属性。
#
# 用户可以指定使用一个特定的容器类型,而不是一个视图。这里我们使用了lists,然而sets, dicts, tuples和其他容器可能在其他情况下更合适。
list(G.nodes)
list(G.edges)
list(G.adj[1]) # or list(G.neighbors(1))
G.degree[1] # the number of edges incident to 1
#
# 用户可以使用一个 *nbunch* 来查看一个点子集的边和度。一个 *nbunch* 可以是 `None` (表示全部节点),一个节点或者一个可迭代的顶点容器。
G.edges([2, 'm'])
G.degree([2, 3])
#
#
# ## 删除图元素
#
# 用户可以使用类似增加节点和边的方式来从图中删除顶点和边。
# 相关方法
# `Graph.remove_node()`,
# `Graph.remove_nodes_from()`,
# `Graph.remove_edge()`
# 和
# `Graph.remove_edges_from()`, 例如
G.remove_node(2)
G.remove_nodes_from("spam")
list(G.nodes)
list(G.edges)
G.remove_edge(1, 3)
G.remove_edges_from([(1, 2), (2, 3)])
list(G.edges)
#
#
# ## 使用图构造函数来构建图
#
# 图对象并不一定需要以增量的方式构建 - 用户可以直接将图数据传给 Graph/DiGraph 的构造函数来构建图对象。
# 当通过实例化一个图类来创建一个图结构时,用户可以使用多种格式来指定图数据, 如下所示。
#
G.add_edge(1, 2)
H = nx.DiGraph(G) # create a DiGraph using the connections from G
list(H.edges())
edgelist = [(0, 1), (1, 2), (2, 3)]
H = nx.Graph(edgelist)
list(H.edges)
#
# ## 访问边和邻居
#
# 除了通过 `Graph.edges` 和 `Graph.adj` 视图外,用户也可以通过下标来访问边和顶点的邻居;
G = nx.Graph([(1, 2, {"color": "yellow"})])
G[1] # same as G.adj[1]
G[1][2]
G.edges[1, 2]
# 当边已经存在时,可以通过下标来获取或设置边的属性:
G.add_edge(1, 3)
G[1][3]['color'] = "blue"
G.edges[1, 3]
G.edges[1, 2]['color'] = "red"
G.edges[1, 2]
#
# 用户可以通过 `G.adjacency()`, 或 `G.adj.items()` 快速地查看所有点的 `(节点,邻居)`对。如下所示:
#
# 注意当图是无向图时,每条边会在遍历时出现两次。
FG = nx.Graph()
FG.add_weighted_edges_from([(1, 2, 0.125), (1, 3, 0.75), (2, 4, 1.2), (3, 4, 0.375)])
for n, nbrs in FG.adj.items():
for nbr, eattr in nbrs.items():
wt = eattr['weight']
if wt < 0.5: print(f"({n}, {nbr}, {wt:.3})")
# 如下所示,可以方便地访问所有边和边的属性。
for (u, v, wt) in FG.edges.data('weight'):
if wt < 0.5:
print(f"({u}, {v}, {wt:.3})")
#
# ## 添加图属性,顶点属性和边属性
#
# 属性如权重、标签、颜色等可以被attach到图、点或者边上。
#
# 每个图、节点和边都可以保存 key/value 属性,默认属性是空。属性可以通过 `add_edge`, `add_node` 或直接对属性字典进行操作来增加或修改属性。
#
# ### 图属性
#
# 在创建新图的时候定义图属性
G = nx.Graph(day="Friday")
G.graph
# 或者在创建后修改图属性
G.graph['day'] = "Monday"
G.graph
#
# ### 节点属性
#
# 可以使用 `add_node()`, `add_nodes_from()`, or `G.nodes` 等方法增加节点属性。
G.add_node(1, time='5pm')
G.add_nodes_from([3], time='2pm')
G.nodes[1]
G.nodes[1]['room'] = 714
G.nodes.data()
#
# 注意 向 `G.nodes` 增加一个节点并不会真正增加节点到图中,如果需要增加新节点,应该使用 `G.add_node()`. 边的使用同样如此。
#
#
# ### 边属性
#
#
# 可以通过 `add_edge()`, `add_edges_from()` 或下标来增加或修改边属性。
G.add_edge(1, 2, weight=4.7 )
G.add_edges_from([(3, 4), (4, 5)], color='red')
G.add_edges_from([(1, 2, {'color': 'blue'}), (2, 3, {'weight': 8})])
G[1][2]['weight'] = 4.7
G.edges[3, 4]['weight'] = 4.2
G.edges.data()
# 特殊的属性如 `weight` 的值应该是数值型,因为一些需要带权重的边的算法会使用到这一属性。
#
#
# ## 抽取子图和边子图
#
# graphscope.nx 支持通过传入一个点集或边集来抽取一个 `deepcopy` 的子图。
#
G = nx.path_graph(10)
# induce a subgraph by nodes
H = G.subgraph([0, 1, 2])
list(H.nodes)
list(H.edges)
# induce a edge subgraph by edges
K = G.edge_subgraph([(1, 2), (3, 4)])
list(K.nodes)
list(K.edges)
#
# 需要注意的是,这里抽取子图与NetworkX的实现有一些区别,NetworkX返回的是一个子图的视图,但 graphscope.nx 返回的子图是一个独立于原始图的子图或边子图。
#
# ## 图的拷贝
#
# 用户可以使用 `to_directed` 方法来获取一个图的有向表示。
DG = G.to_directed() # here would return a "deepcopy" directed representation of G.
list(DG.edges)
# or with
DGv = G.to_directed(as_view=True) # return a view.
list(DGv.edges)
# or with
DG = nx.DiGraph(G) # return a "deepcopy" of directed representation of G.
list(DG.edges)
# 或者通过 `copy` 方法得到一个图的拷贝。
H = G.copy() # return a view of copy
list(H.edges)
# or with
H = G.copy(as_view=False) # return a "deepcopy" copy
list(H.edges)
# or with
H = nx.Graph(G) # return a "deepcopy" copy
list(H.edges)
# 注意,graphscope.nx 不支持浅拷贝。
#
#
# ## 有向图
#
# `DiGraph` 类提供了额外的方法和属性来指定有向边,如:`DiGraph.out_edges`, `DiGraph.in_degree`,
# `DiGraph.predecessors()`, `DiGraph.successors()` etc.
#
# 为了让算法方便地在两种图类型上运行,有向版本的 `neighbors` 等同于 `successors()` ,`degree` 返回 `in_degree` 和 `out_degree` 的和。
DG = nx.DiGraph()
DG.add_weighted_edges_from([(1, 2, 0.5), (3, 1, 0.75)])
DG.out_degree(1, weight='weight')
DG.degree(1, weight='weight')
list(DG.successors(1))
list(DG.neighbors(1))
list(DG.predecessors(1))
#
#
# 在 graphscope.nx 中,存在有些算法仅能用于有向图的分析,而另一些算法仅能用于无向图的分析。如果你想将一个有向图转化为无向图,你可以使用 `Graph.to_undirected()`:
H = DG.to_undirected() # return a "deepcopy" of undirected represetation of DG.
list(H.edges)
# or with
H = nx.Graph(DG) # create an undirected graph H from a directed graph G
list(H.edges)
#
# DiGraph 也可以通过 `DiGraph.reverse()` 来反转边。
K = DG.reverse() # retrun a "deepcopy" of reversed copy.
list(K.edges)
# or with
K = DG.reverse(copy=False) # return a view of reversed copy.
list(K.edges)
#
# ## 图分析
# 图 `G` 的结构可以通过使用各式各样的图理论函数进行分析,例如:
G = nx.Graph()
G.add_edges_from([(1, 2), (1, 3)])
G.add_node(4)
sorted(d for n, d in G.degree())
nx.builtin.clustering(G)
#
# 在 graphscope.nx 中,我们支持用于图分析的内置算法,算法的详细内容可以参考 [builtin algorithm](https://graphscope.io/docs/reference/networkx/builtin.html)
#
#
# ## 通过 GraphScope graph object来创建图
#
# 除了通过networkx的方式创建图之外,我们也可以使用标准的 `GraphScope` 的方式创建图,这一部分将会在下一个教程中进行介绍,下面我们展示一个简单的示例:
# +
# we load a GraphScope graph with load_ldbc
from graphscope.dataset import load_ldbc
graph = load_ldbc(directed=False)
# create graph with the GraphScope graph object
G = nx.Graph(graph)
# -
#
#
# ## 将图转化为graphscope.graph
#
#
# 正如同 graphscope.nx Graph 可以从 GraphScope graph 转化而来,graphscope.nx Graph也可以转化为 GraphScope graph. 例如:
nodes = [(0, {"foo": 0}), (1, {"foo": 1}), (2, {"foo": 2})]
edges = [(0, 1, {"weight": 0}), (0, 2, {"weight": 1}), (1, 2, {"weight": 2})]
G = nx.Graph()
G.update(edges, nodes)
g = graphscope.g(G)
| tutorials/zh/2_graph_manipulations_with_networkx_compatible_apis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={} tags=[]
# <img width="10%" alt="Naas" src="https://landen.imgix.net/jtci2pxwjczr/assets/5ice39g4.png?w=160"/>
# + [markdown] papermill={} tags=[]
# # Hubspot - Create contacts from linkedin post likes
# <a href="https://app.naas.ai/user-redirect/naas/downloader?url=https://raw.githubusercontent.com/jupyter-naas/awesome-notebooks/master/Hubspot/Hubspot_create_contacts_from_linkedin_post_likes.ipynb" target="_parent"><img src="https://naasai-public.s3.eu-west-3.amazonaws.com/open_in_naas.svg"/></a>
# + [markdown] papermill={} tags=[]
# **Tags:** #hubspot #crm #sales #contact #naas_drivers #linkedin #post #contact
# + [markdown] papermill={} tags=[]
# ## Input
# + [markdown] papermill={} tags=[]
# ### Install librairies
# + papermill={} tags=[]
from naas_drivers import linkedin, hubspot
import naas
import requests
# + [markdown] papermill={} tags=[]
# ### Setup your secrets
# Uncomment lines to setup.<br>
# <a href='https://docs.naas.ai/features/secret'>How to setup your secret keys ?</a>
# + [markdown] papermill={} tags=[]
# #### → For Hubspot
# + papermill={} tags=[]
#naas.secret.add('HUBSPOT_API_KEY', secret="7696655f-3aff-4524-84f1-5c988abdab9f")
# + [markdown] papermill={} tags=[]
# #### → For LinkedIn
# + [markdown] papermill={} tags=[]
# <a href='https://www.notion.so/LinkedIn-driver-Get-your-cookies-d20a8e7e508e42af8a5b52e33f3dba75'>How to get your cookies ?</a>
# + papermill={} tags=[]
#naas.secret.add('LINKEDIN_LI_AT', secret="***")# EXAMPLE AQFAzQN_PLPR4wAAAXc-FCKmgiMit5FLdY1af3-2
#naas.secret.add('LINKEDIN_JSESSIONID', secret= "ajax:***") # EXAMPLE ajax:8379907400220387585
# + [markdown] papermill={} tags=[]
# ### Configure secrets to connect to APIs
# + papermill={} tags=[]
#Hubspot API
auth_token = naas.secret.get('HUBSPOT_API_KEY')
# + papermill={} tags=[]
# LinkedIn API
LI_AT = naas.secret.get('LINKEDIN_LI_AT')
JSESSIONID = naas.secret.get('LINKEDIN_JSESSIONID')
# + [markdown] papermill={} tags=[]
# ### Enter post URL
# + papermill={} tags=[]
POST_URL = "----"
# + [markdown] papermill={} tags=[]
# ### Get post likes
# + papermill={} tags=[]
df_posts = linkedin.connect(LI_AT, JSESSIONID).post.get_likes(POST_URL)
# Display the number of likes
print("Number of likes: ", df_posts.PROFILE_URN.count())
# + papermill={} tags=[]
# Show dataframe with list of profiles from likes
df_posts
# + [markdown] papermill={} tags=[]
# ## Model
# + [markdown] papermill={} tags=[]
# ### Create contacts from LinkedIn post likes
# + papermill={} tags=[]
def create_contacts_from_post(df,
c_profile_urn="PROFILE_URN",
c_firstname="FIRSTNAME",
c_lastname="LASTNAME",
c_occupation="OCCUPATION"):
for _, row in df.iterrows():
profile_urn = row[c_profile_urn]
firstname = row[c_firstname]
lastname = row[c_lastname]
occupation = row[c_occupation]
linkedinbio = f"https://www.linkedin.com/in/{profile_urn}"
email = None
phone = None
# contact
try:
contact = linkedin.connect(LI_AT, JSESSIONID).profile.get_contact(linkedinbio)
email = contact.loc[0, "EMAIL"]
phone = contact.loc[0, "PHONENUMBER"]
except:
print("No contact info")
# With send method
data = {"properties":
{
"linkedinbio": linkedinbio,
"firstname": firstname,
"lastname": lastname,
"jobtitle": occupation,
"email": email,
"phone": phone,
}
}
print(data)
hubspot.connect(auth_token).contacts.send(data)
# + [markdown] papermill={} tags=[]
# ## Output
# + [markdown] papermill={} tags=[]
# ### Display result
# + papermill={} tags=[]
create_contacts_from_post(df_posts)
| Hubspot/Hubspot_create_contacts_from_linkedin_post_likes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # "Topic 03.2: Text Representation(PART-2)"
#
# > "Representing raw text to suitable numeric format"
#
# - toc: true
# - branch: master
# - badges: true
# - comments: true
# - categories: [basic-nlp]
# - image: images/logo/logo.png
# - hide: False
# - sticky_rank: 6
#
# + [markdown] id="az70e-5kcYVn"
#
# In the previous topic we have covered the distributional apporach(which have high dimension vector to represent words and also sparse in nature) but in this post we will cover the Distributed apporach(which have low dimension vecotr and are dense in nature) and how to create word embedding using pretrained model.
#
#
# ## Distributed Representation
# To overcome the issue of high-dimensional representation and sparse vector to represent word, Distributed Representation help in these issue and therefore they have gained a lot of momentum in the past six to seven days.
# Different distributed representation are
#
# ### Word Embedding
# Word Embeddings are the texts converted into numbers. Embeddings translate large sparse vectors into a lower-dimensional space that preserves semantic relationships.
# Word embeddings is a technique where individual words of a domain or language are represented as real-valued vectors in a lower dimensional space and placing vectors of semantically similar items close to each other. This way words that have similar meaning have similar distances in the vector space as shown below.
#
# *“king is to queen as man is to woman” encoded in the vector space as well as verb Tense and Country and their capitals are encoded in low dimensional space preserving the semantic relationships.*
#
#
# <img src = "my_icons/topic_3.2.a.png">
#
# Word2vec is an algorithm invented at Google for training word embeddings. word2vec relies on the distributional hypothesis. The distributional hypothesis states that words which, often have the same neighboring words tend to be semantically similar. This helps to map semantically similar words to geometrically close embedding vectors.
#
#
# Now the question arises that how we will create word embedding?
#
# Well we can also use pre-trained word embedding arcitecture or we can also train our own word embedding.
#
# #### Pre-trained word embeddings
# * **What is pre-trained word embeddings?**
#
# Pretrained Word Embeddings are the embeddings learned in one task that are used for solving another similar task.
#
# These embeddings are trained on large datasets, saved, and then used for solving other tasks. That’s why pretrained word embeddings are a form of Transfer Learning.
#
#
#
# * **Why do we need Pretrained Word Embeddings?**
#
# Pretrained word embeddings capture the semantic and syntactic meaning of a word as they are trained on large datasets. They are capable of boosting the performance of a Natural Language Processing (NLP) model. These word embeddings come in handy during hackathons and of course, in real-world problems as well.
#
#
#
# * **But why should we not learn our own embeddings?**
#
# Well, learning word embeddings from scratch is a challenging problem due to two primary reasons:
#
# * Sparsity of training data
# * Large number of trainable parameters
#
#
# With pretrained embedding you just need to download the embeddings and use it to get the vectors for the word you want.Such embeddings can be
# thought of as a large collection of key-value pairs, where keys are the
# words in the vocabulary and values are their corresponding word
# vectors. Some of the most popular pre-trained embeddings are
# Word2vec by Google, GloVe by Stanford, and fasttext
# embeddings by Facebook, to name a few. Further, they’re
# available for various dimensions like d = 25, 50, 100, 200, 300, 600.
#
#
# >Here is the code where we will find the words that are semantically most similar to the word "beautiful".
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="4kZBIQE_t_7M" outputId="56244fed-641f-4171-bbfd-fe41ef7eedfa"
#We will use the Google News vectors embeddings.
#Downdloading Google News vectors embeddings.
# !wget -P /tmp/input/ -c "https://s3.amazonaws.com/dl4j-distribution/GoogleNews-vectors-negative300.bin.gz"
# + colab={"base_uri": "https://localhost:8080/"} id="ZHc9-ZJXceyT" outputId="8a210eca-9d0c-427a-e356-f91fc2ac15c1"
from gensim.models import Word2Vec, KeyedVectors
pretrainedpath = '/tmp/input/GoogleNews-vectors-negative300.bin.gz'
w2v_model = KeyedVectors.load_word2vec_format(pretrainedpath, binary=True) #load the model
print("done loading word2vec")
print("Numver of words in vocablulary: ",len(w2v_model.vocab)) #Number of words in the vocabulary.
# + colab={"base_uri": "https://localhost:8080/"} id="EEmtmt7fwgva" outputId="7f3b3b7c-83b3-4f91-9875-0ac04976b08c"
#Let us examine the model by knowing what the most similar words are, for a given word!
w2v_model.most_similar('beautiful')
# + [markdown] id="VyR7YiSOxDZV"
# **Note that if we search for a word that is not present in the Word2vec
# model (e.g., “practicalnlp”), we’ll see a “key not found” error. Hence,
# as a good coding practice, it’s always advised to first check if the
# word is present in the model’s vocabulary before attempting to
# retrieve its vector.**
# + colab={"base_uri": "https://localhost:8080/", "height": 323} id="akecvo_SxRRZ" outputId="08d932cd-bb73-4b80-9dfa-e9f9cd78b881"
#What if I am looking for a word that is not in this vocabulary?
w2v_model['practicalnlp']
# + [markdown] id="aVwB9XXzxtsV"
# >If you’re new to embeddings, always start by using pre-trained word
# embeddings in your project. Understand their pros and cons, then start thinking
# of building your own embeddings. Using pre-trained embeddings will quickly
# give you a strong baseline for the task at hand.
#
#
#
# In the next blog post we will cover the **Training our own emdeddings models**.
#
#
# + [markdown] id="uDwWnExk1bhY"
# #### TRAINING OUR OWN EMBEDDINGS
# For training our own word embeddings we’ll look at two architectural variants that were propossed in Word2Vec
# * Continuous bag of words(CBOW)
# * SkipGram
#
#
# ##### **Continuous Bag of Words**
#
# CBOW tries to learn a language model that tries to predict the “center” word from the words in its context. Let’s understand this using our toy corpus(the quick brown fox jumped over the lazy dog). If we take the word “jumps” as the center word, then its context is formed by words in its vicinity. If we take the context size of 2, then for our example, the context is given by brown, fox, over, the. CBOW uses the context words to predict the target word—jumped. CBOW tries to do this
# for every word in the corpus; i.e., it takes every word in the corpus as the target word and tries to predict the target word from its
# corresponding context words.
#
# <img src = "my_icons/topic_3.2.b.png">
#
# **Understanding CBOW architecture**
#
# <img src = "my_icons/topic_3.2.c.png">
#
# consider the training corpus having the following sentences:
#
# *“the dog saw a cat”, “the dog chased the cat”, “the cat climbed a tree”*
#
# The corpus vocabulary has eight words. Once ordered alphabetically, each word can be referenced by its index. For this example, our neural network will have eight input neurons and eight output neurons. Let us assume that we decide to use three neurons in the hidden layer. This means that WI and WO will be 8×3 and 3×8 matrices, respectively. Before training begins, these matrices are initialized to small random values as is usual in neural network training. Just for the illustration sake, let us assume WI and WO to be initialized to the following values:
#
# <img src = "my_icons/topic_3.2.d.png">
#
#
# Suppose we want the network to learn relationship between the words “cat” and “climbed”. That is, the network should show a high probability for “climbed” when “cat” is inputted to the network. In word embedding terminology, the word “cat” is referred as the context word and the word “climbed” is referred as the target word. In this case, the input vector X will be [0 1 0 0 0 0 0 0]. Notice that only the second component of the vector is 1. This is because the input word is “cat” which is holding number two position in sorted list of corpus words. Given that the target word is “climbed”, the target vector will look like [0 0 0 1 0 0 0 0 ]t.
#
# With the input vector representing “cat”, the output at the hidden layer neurons can be computed as
#
# **Ht = XtWI = [-0.490796 -0.229903 0.065460]**
#
# It should not surprise us that the vector H of hidden neuron outputs mimics the weights of the second row of WI matrix because of 1-out-of-V representation. So the function of the input to hidden layer connections is basically to copy the input word vector to hidden layer. Carrying out similar manipulations for hidden to output layer, the activation vector for output layer neurons can be written as
#
# **HtWO = [0.100934 -0.309331 -0.122361 -0.151399 0.143463 -0.051262 -0.079686 0.112928]**
#
#
# now we will use the formula
# <img src = "my_icons/topic_3.2.e.png">
#
#
# Thus, the probabilities for eight words in the corpus are:
#
# **[0.143073 0.094925 0.114441 0.111166 0.149289 0.122874 0.119431 0.144800]**
#
# The probability in bold is for the chosen target word “climbed”. Given the target vector is [0 0 0 1 0 0 0 0 ]
#
# The above description and architecture is meant for learning relationships between pair of words. In the continuous bag of words model, context is represented by multiple words for a given target words. For example, we could use “cat” and “tree” as context words for “climbed” as the target word. This calls for a modification to the neural network architecture. The modification, shown below, consists of replicating the input to hidden layer connections C times, the number of context words, and adding a divide by C operation in the hidden layer neurons.
#
# **[An alert reader pointed that the figure below might lead some readers to think that CBOW learning uses several input matrices. It is not so. It is the same matrix, WI, that is receiving multiple input vectors representing different context words]**
#
# <img src = "my_icons/topic_3.2.f.png">
# -
# I can understand that things can be little hazy at first.But if you read this one more time it will be crystal clear.
#
# In the next blog i will cover skip-gram and other text representation technique.
# + [markdown] id="U0CZy26Y1pCB"
#
# {{ 'Notes are compiled from [ Practical Natural Language Processing: A Comprehensive Guide to Building Real-World NLP Systems](https://www.oreilly.com/library/view/practical-natural-language/9781492054047/), [Medium](https://towardsdatascience.com/word-embeddings-for-nlp-5b72991e01d4),[CBOW and Skip-gram](https://iksinc.online/tag/continuous-bag-of-words-cbow/#:~:text=In%20the%20continuous%20bag%20of,to%20the%20neural%20network%20architecture.) and [Code from github repo](https://github.com/practical-nlp/practical-nlp/tree/master/Ch3)' | fndetail: 1 }}
# {{ 'If you face any problem or have any feedback/suggestions feel free to comment.' | fndetail: 2 }}
| _notebooks/2021_02_1_Text_Representation(PART_2).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.10 64-bit (''base'': conda)'
# name: python3
# ---
# # Figure 1. Accuracy over training epoch
# ## Import libraries
import pandas as pd
import altair as alt
from helper import RawData, apply_font_size
# ## Import part I data
raw = RawData("../../data/data_part1_1250.csv")
# ## Function for plotting figure 1
def fig1(sim_df:pd.DataFrame, font_size:int=18) -> alt.Chart:
"""Plot accuracy by condition and word type over epoch"""
df = sim_df.copy()
df["condition"] = df.cond.apply(lambda x: "NW" if x in ("NW_AMB", "NW_UN") else x)
df = df.groupby(["epoch", "condition"]).mean().reset_index()
plot = alt.Chart(df).mark_line().encode(
x=alt.X("epoch:Q", title="Sample (Mil.)"),
y=alt.Y("score:Q", title="Accuracy", scale=alt.Scale(domain=(0, 1))),
color=alt.Color(
"condition:N",
legend=alt.Legend(orient="bottom-right"),
title="Stimulus",
),
strokeDash=alt.condition(
alt.datum.condition == "NW", alt.value([5, 5]), alt.value([0])
)
).properties(width=400, height=300)
return apply_font_size(plot, font_size)
# ## Plotting figure 1 from part I raw data
fig1(raw.df)
# Figure 1. Accuracy over training epoch (in fractions of a million samples) including high-frequency consistent words (HF_CON), high-frequency inconsistent words (HF_INC), low-frequency consistent words (LF_CON), low-frequency inconsistent words (LF_INC), as well as overall nonwords (NW).
| code/python/figure1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from numpy import *
from itertools import combinations
dataSet = [[1, 3, 4], [2, 3, 5], [1, 2, 3, 5], [2, 5]]
def createC1(dataSet):
C1 = []
for transaction in dataSet:
for item in transaction:
if not [item] in C1:
C1.append([item])
C1.sort()
return list(C1)
def scanD(D, Ck, minSupport):
ssCnt = {}
for tid in D:
for can in Ck:
if set(can).issubset(tid):
if not tuple(can) in ssCnt: ssCnt[tuple(can)]=1
else: ssCnt[tuple(can)] += 1
numItems = float(len(D))
retList = []
supportData = {}
for key in ssCnt:
support = ssCnt[key]/numItems
if support >= minSupport:
retList.insert(0,key)
supportData[key] = support
return retList, supportData
def checkSubsetFrequency(candidate, Lk, k):
if k>1:
subsets = list(combinations(candidate, k))
else:
return True
for elem in subsets:
if not elem in Lk:
return False
return True
def aprioriGen(Lk, k): #creates Ck
resList = [] #result set
candidatesK = []
lk = sorted(set([item for t in Lk for item in t])) #get and sort elements from frozenset
candidatesK = list(combinations(lk, k))
for can in candidatesK:
if checkSubsetFrequency(can, Lk, k-1):
resList.append(can)
return resList
def apriori(dataSet, minSupport):
C1 = createC1(dataSet)
D = list(dataSet)
L1, supportData = scanD(D, C1, minSupport)
L = [L1]
k = 2
while (len(L[k-2]) > 0):
Ck = aprioriGen(L[k-2], k)
Lk, supK = scanD(D, Ck, minSupport)#scan DB to get Lk
supportData.update(supK)
L.append(Lk)
k += 1
#remove empty last itemset from L
if L[-1] == []:
L.pop()
return L, supportData
L, suppData = apriori(dataSet, 0.5)
L
def generateRules(L, supportData, minConf): #supportData is a dict coming from scanD
for i in range(1, len(L)): #only get the sets with two or more items
for item in L[i]: #for each item in a level
for j in range(1, i+1): # i+1 equal to length of an item
lhsList = list(combinations(item, j))
for lhs in lhsList:
rhs = set(item).difference(lhs)
conf = supportData[item]/supportData[lhs]
if conf >= minConf:
print(list(lhs), " ==> ", list(rhs), " [", supportData[item], ", ", conf,"]", sep="")
generateRules(L, suppData, 0.5)
| Apriori.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
s1 = input("Enter string 1 : ")
s2 = input("Enter string 2 : ")
print(s1[1::2] + s2[::2])
# +
s1 = input("Enter string 1 : ")
s2 = input("Enter string 2 : ")
if len(s1) < len(s2):
min_length = len(s1)
else:
min_length = len(s2)
"".join([s1[i] if s1[i].isupper() else s2[i].upper() for i in range(0, min_length)])
# +
l = input("Enter length : ")
b = input("Enter breadth : ")
h = input("Enter height : ")
N = input("Enter N : ")
b = {(i, j, k) : abs(i) + abs(j) + abs(k)
for i in [0, int(l)]
for j in [0, int(b)]
for k in [0, int(h)]
if abs(i) + abs(j) + abs(k) < int(N)}
print(b)
# +
a = [[1]]
b = [[1, 2, 3],
[4, 5, 6],
[7, 8, 9]]
c = [[1, 2, 3, 4, 5],
[6, 7, 8, 9, 10],
[11, 12, 13, 14, 15],
[16, 17, 18, 19, 20],
[21, 22, 23, 24, 25]]
def spiral(A):
n = len(A)
for i in range(0, n // 2 + 1):
for j in range(i, n - i):
print(A[i][j], end=', ')
for j in range(i + 1, n - i):
print(A[j][n - i - 1], end=', ')
for j in range(n - i - 2, i - 1, -1):
print(A[n - i - 1][j], end=', ')
for j in range(n - i - 2, i, -1):
print(A[j][i], end=', ')
print()
spiral(a)
spiral(b)
spiral(c)
| Week 2/Practice Set 1(Solutions).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
library(keras)
use_condaenv("r-tensorflow")
X = read.csv(url("https://www.dropbox.com/s/h4hgr256iicmws5/X5k.txt?dl=1"));
Y = read.csv(url("https://www.dropbox.com/s/i3jxoprwn8iwfg1/Y5k.txt?dl=1"));
sz=dim(X)
dim(Y)
sz
X_std <- scale(X)
mu = colMeans(x_std) # faster version of apply(scaled.dat, 2, mean)
sd = apply(x_std, 2, sd)
mu[1:2]
sd[1:2]
smp_size <- floor(0.75 * nrow(x_std))
set.seed(123)
train_ind <- sample(seq_len(nrow(x_std)), size = smp_size)
X_train <- x_std[train_ind, ]
X_val <- x_std[-train_ind, ]
Y_train <- Y[train_ind,1]
Y_val <- Y[-train_ind,1]
num_class = max(Y)+1
label = to_categorical(Y, num_class)
label_train <- to_categorical(Y_train, num_class)
label_val <- to_categorical(Y_val, num_class)
model <- keras_model_sequential()
model %>%
layer_dense(units = 64, activation = "relu", input_shape = c(sz[2])) %>%
layer_dropout(rate = 0.4) %>%
layer_dense(units = 64, activation = "relu") %>%
layer_dropout(rate = 0.3) %>%
layer_dense(units = num_class, activation = "softmax")
model %>% compile(
loss = "categorical_crossentropy",
optimizer = optimizer_rmsprop(),
metrics = c("accuracy")
)
summary(model)
history <- model %>% fit(
X_train, label_train,
epochs = 20, batch_size = 128,
validation_split = 0.2
)
plot(history)
model %>% evaluate(X_val, label_val,verbose = 1)
model %>% predict_classes(x_test)
| src/dl_airbnb_5k_r.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn import linear_model, ensemble, svm, tree, neural_network
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import MinMaxScaler, StandardScaler
from sklearn.metrics import mean_squared_error, mean_absolute_error, make_scorer
from sklearn.model_selection import cross_val_score, train_test_split, GridSearchCV
from sklearn.feature_selection import f_classif
from sklearn.feature_selection import SelectKBest
import warnings
warnings.filterwarnings("ignore")
# -
result = {}
# +
hhids=[26, 59, 77, 86, 93, 94, 101, 114, 171, 187]
hhids=[86]
for hhid in hhids:
X=[]
result[hhid] = []
print('Start :: Process on household {}...'.format(hhid))
df = pd.read_csv('data_filled2/processed_hhdata_{}_2.csv'.format(hhid), index_col=0)
features = ['temperature', 'cloud_cover','wind_speed','month','hour']
Y = list(df.AC)[500:]
Y = np.array(Y)
print(Y.shape)
print(Y[0])
#get X
for index, row in df.iterrows():
if index>=500:
rowlist=row[features]
rowlist = rowlist.tolist()
X.append(rowlist)
# rowlist.append(df.use.iloc[index-1])
# rowlist.append(df.use.iloc[index-24])
# rowlist.append(df.use.iloc[index-168])
for i in range(1,169):
rowlist.append(df.use.iloc[index-i])
X = np.array(X)
print(X.shape)
print(X[0])
print(len(X[0]))
X_train, X_test, Y_train, Y_test = train_test_split(X, Y,
test_size=0.10,
random_state=666)
# Create an SelectKBest object to select features with two best ANOVA F-Values
fvalue_selector = SelectKBest(f_classif, k=9)
# Apply the SelectKBest object to the features and target
X_kbest = fvalue_selector.fit_transform(X, Y)
# Show results
print(X_kbest[0])
cols = fvalue_selector.get_support(indices=True)
print(cols)
print('Original number of features:', X.shape[1])
print('Reduced number of features:', X_kbest.shape[1])
| data_processing/ghi_model_score.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Geometric Multigrid
# <NAME>
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Introduction
# We follow the analysis from "A Multigrid Tutorial" by <NAME>. This presentation will focus on the use of the Multigrid Method as a solver for the 1D Poisson problem, though the concepts and ideas can be applied to problems of higher dimension, or as a preconditioner instead of a solver.
# + jupyter={"source_hidden": true} slideshow={"slide_type": "skip"}
from prettytable import PrettyTable
from IPython.display import HTML
import time
import numpy as np
from numpy.linalg import norm, solve
import scipy.sparse as sp
import scipy.sparse.linalg as splu
import matplotlib.pyplot as plt
from numpy.random import MT19937, RandomState, SeedSequence
rs = RandomState(MT19937(SeedSequence(123456789)))
# We will be profiling code later, this extension allows line-by-line profiling
# %load_ext line_profiler
# Unified mvmult user interface for both scipy.sparse and numpy matrices.
# In scipy.sparse, mvmult is done using the overloaded * operator, e.g., A*x.
# In numpy, mvmult is done using the dot() function, e.g., dot(A,x).
# This function chooses which to use based on whether A is stored as
# a sparse matrix.
def mvmult(A, x):
if sp.issparse(A):
return A*x
else:
return np.dot(A,x)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Analyzing the Jacobi Method
# + [markdown] slideshow={"slide_type": "slide"}
# ### Problem Set Up
# First, we create the A matrix. For simplicity and easy of understanding, we will choose the 1D poisson problem with Dirichlet Bondary Conditions.
#
# The matrix for this problem is given by
# $$
# A = \frac{1}{h^2}
# \begin{bmatrix}
# 2 & -1 & & & & \\
# -1 & 2 & -1 & & & \\
# & -1 & 2 & -1 & & \\
# & & \ddots & \ddots & \ddots & \\
# & & & -1 & 2 & -1 \\
# & & & & -1 & 2
# \end{bmatrix}
# $$
# so the problem can be written as $Ax = b$. If $N$ is the number of subintervals we divide the domain into, then this matrix is $N-1 \times N-1$.
#
# For the sake of demonstration, we let the right-hand side vector $b$ correspond to the case of Dirichlet boundary conditions where both ends are held at $0$. We choose this because the true solution is therefore the zero vector so the error corresponds exactly with the current iterate.
# + slideshow={"slide_type": "skip"}
def GeneratePoisson(N):
# Generate A matrix for poisson problem with N subintervals (N-1 unknowns)
# Result will be a N-1 x N-1 matrix
row = N*N*np.array([-1, 2, -1], dtype=np.double)
A = sp.diags(row, [-1, 0, 1], shape=(N-1, N-1), dtype=np.double, format="csr")
return A
N = 512
A = GeneratePoisson(N)
b = np.zeros((N-1,1), dtype=np.double)
# + [markdown] slideshow={"slide_type": "subslide"}
# We also generate some initial guesses (which are also initial errors) which are sine waves with varying frequencies.
# + slideshow={"slide_type": "fragment"}
x = np.linspace(0,1,N+1)
waveNumbers = [1, 3, 10, 20, 50, 100]
#waveNumbers = [1, 10, 50, 100]
xinitial = [np.sin(w*np.pi*x) for w in waveNumbers]
fig, axs = plt.subplots(len(waveNumbers),1)
for (i,p) in enumerate(xinitial):
axs[i].plot(p)
# + [markdown] slideshow={"slide_type": "skip"}
# ### Create a Jacobi Function
# + slideshow={"slide_type": "skip"}
def Jacobi(x, A, b, numiters=1):
for i in range(numiters):
# Need a copy of x since we'll be updating x in place
xold = x.copy()
# Loop through the rows
for i in range(len(xold)):
# Calculate the indices of the CSR data array that hold the row
rowindices = range(A.indptr[i],A.indptr[i+1])
r = 0.
for j in rowindices:
col = A.indices[j]
if col != i:
r += A.data[j]*xold[col]
else:
d = A.data[j]
# Update x
x[i] = (b[i] - r)/d
return x
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Running Jacobi
# Now let's run 100 Jacobi iterations on each of the initial conditions, tracking the error at each iteration.
# + slideshow={"slide_type": "skip"}
numJacobiIters = 100
errors = [np.zeros((numJacobiIters+1,1), dtype=np.double) for i in range(len(waveNumbers))]
initialErrorNorms = [np.linalg.norm(w) for w in xinitial]
x_working = xinitial.copy()
for j in range(numJacobiIters+1):
for (i, w) in enumerate(x_working):
if j == 0:
errors[i][j] = np.linalg.norm(w[1:-1])/initialErrorNorms[i]
else:
errors[i][j] = np.linalg.norm(Jacobi(w[1:-1], A, b, 1))/initialErrorNorms[i]
# + slideshow={"slide_type": "fragment"}
for (i,w) in enumerate(errors):
plt.plot(w, label = waveNumbers[i])
plt.legend(loc="right")
plt.xlabel("Iteration")
plt.ylabel("Error")
plt.show()
# + [markdown] slideshow={"slide_type": "subslide"}
# We can also look at our iterates now:
# + slideshow={"slide_type": "fragment"}
fig, axs = plt.subplots(len(waveNumbers),1)
for (i,p) in enumerate(x_working):
axs[i].plot(p)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Why Multigrid Works
#
# Here we see the key to understanding the effectiveness of multigrid: The Jacobi iteration scheme is much better at eliminating high frequency error than low frequency error.
# -
# ### Other Iteration Methods
#
# While the preceeding discussion uses Jacobi iteration, all relaxation-type iterations methods like Gauss-Seidel and variations of Jacobi and Gauss-Seidel also present this property. In multigrid terminology, these are all generally referred to as relaxations.
# ### How Do We Use This?
#
# The matrix system does not have any explicit information about the phyiscal structure of the problem. Hence, if we take a longer vector with low frequency error, and remove half of the points:
y_fine = np.sin(3*np.pi*x)
x_coarse = np.array([x[i] for i in range(len(x)) if i % 2 == 0])
y_coarse = np.sin(3*np.pi*x_coarse)
fig, axs = plt.subplots(2, 1)
axs[0].plot(y_fine)
axs[1].plot(y_coarse)
plt.xlim(right=N)
plt.show()
# we can see that we have the same number of oscillations, but in half the number of nodes. For Jacobi, which has no information about the entries represent, the shorter vector has higher frequency error than the longer vector so it would be more effective on this new shorter vector.
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## In Practice
# Consider a discretized PDE problem on a grid (which we'll denote $\Omega^h$) where $h$ represents the spacing between nodes. As the name suggests, for the multigrid method we'll be using multiple grids, each with a different spacing of nodes. From here on, we'll be using superscript to denote which grid a quantity is on.
#
# Our discretized problem is written as $A^h x^h = b^h$. We'll start with $k$ Jacobi iterations. Since we don't expect our current iteration $x^h$ to be the exact solution, let's assume the exact solution is of the form $x^* = x^h + e^h$. This gives us an equation of the form:
# \begin{equation*}
# A^h(x^h + e^h) = b^h
# \end{equation*}
# Rearranging this equation gives
# \begin{equation*}
# A^h e^h = b^h - A^hx^h = r^h
# \end{equation*}
# So if we calculate $r^h$ and solve $A^h e^h = r^h$ for $e^h$, then we could find the exact solution as $x^* = x^h + e^h$.
#
# So how do we find or (more accurately) approximate $e^h$? Running more Jacobi iterations at this level has already shown to be less effective since the high frequency error has already been removed. Only the lower frequency error remains. Instead, we will move the problem down to a coarser grid, $\Omega^{2h}$. In the coarser grid, the low frequency error changes to higher frequency error and Jacobi can be more effective.
#
# That is, we want to solve $A^{2h}e^{2h} = r^{2h}$, where $A^{2h}$, $e^{2h}$, and $r^{2h}$ are the \"coarse grid versions\" of $A^h$, $e^h$, and $r^h$. We will discuss how to find these later.
#
# This coarser grid problem is smaller (by a factor of 4) so it will take less computational effort. We can either solve the system exactly or use another method to approximate the solution to the coarse grid system. We then \"transfer\" this back into the $\Omega^h$ grid and it becomes an approximation to $e^h$. We then calculate $x^h + e^h$ to get a better approximation for $x^h$. In doing so, the transfer may have introduced more high frequency error, so we typically complete more Jacobi iterations at the fine level to remove these. This process leverages the change of grids to use Jacobi iteration more effectively.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Moving Between Grids
# We need a way to transform the problem, and all the quantities involved, between the various grids in order to leverage such a scheme. We will examine the simplest case for geometric multigrid in 1D. Assume that the coarse grid has grid spacing which is twice as large as the finer grid (that is, the coarse grid is the fine grid with every other node removed). This is almost universal practice since evidence does not seem to indicate there is any advantage to choosing a different ratio.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Restriction Operator
# First, let's talk about an operator for transforming from the fine grid to the coarse grid. This is typically referred to as the **restriction** operator. We will denote is as $I_h^{2h}$ and there are a few reasonable options for how to build this operator. One option is to simply remove half of the grid points. This option could work, but tends to remove too much information. The more recommended option is **full weighting**. With this operator, we produce coarse grid vectors according to the rule $I_{h}^{2h} x^{h} = x^{2h}$ where
# \begin{align*}
# x_{j}^{2h} &= \frac{1}{4} \left( x_{2j-1}^{h} + 2x_{2j} + x_{j+1}^{h} \right)
# \end{align*}
# For example, if we have 8 subintervals in our fine grid (that is 7 interior nodes), and 4 subintervals in our coarse grid (with 3 interior nodes), then we have the following:
# $$
# I_{h}^{2h} x^{h} = \frac{1}{4}
# \begin{bmatrix}
# 1 & 2 & 1 & & & & \\
# & & 1 & 2 & 1 & & \\
# & & & & 1 & 2 & 1 \\
# \end{bmatrix}
# \begin{bmatrix}
# x_1 \\ x_2 \\ x_3 \\ x_4 \\ x_5 \\ x_6 \\ x_7
# \end{bmatrix}_{h}
# = \begin{bmatrix}
# x_1 \\ x_2 \\ x_3
# \end{bmatrix}_{2h} = x^{2h}
# $$
# We can also represent this operator with a "stencil" (a common representation for weighted averages of geometric nodes) given by
# $$
# \frac{1}{4} \begin{bmatrix} 1 & 2 & 1 \end{bmatrix}
# $$
#
# This opreator also has another advantage that we'll mention later.
# -
# Let's build a function to generate the Full Weighting operator for transforming
# a vector of length 2^n-1 to a vector of length 2^(n-1) - 1
def BuildFullWeighting(N):
# We assume N = 2^n-1 is the number of unknown in the fine mesh
# We calculate the number of unknowns in the coarse mesh
# Note: this is doing a cast to integer which discards any decimal components
Nc = int(N/2)
# We will build it in coordinate format, though it with a little more effort,
# it could be efficiently built in CSR format as well.
#
# There are 3*Nc non-zeros in the matrix
row_indicies = np.zeros(3*Nc, dtype="int")
col_indicies = np.zeros(3*Nc, dtype="int")
data = np.zeros(3*Nc)
for i in range(Nc):
row_indicies[3*i:3*i+3] = [i, i, i]
col_indicies[3*i:3*i+3] = [2*i, 2*i + 1, 2*i + 2]
data[3*i:3*i+3] = [0.25, 0.5, 0.25]
# Build the matrix
I_FW = sp.coo_matrix((data, (row_indicies,col_indicies))).tocsr()
return I_FW
# Let's look at how this operator acts on a short vector:
# + slideshow={"slide_type": "subslide"}
# Set up the fine grid first
x_fine = np.linspace(0,1,9)
y_fine = np.sin(3*np.pi*x_fine)
# Now the coarse grid
x_coarse = np.linspace(0,1,5)
y_coarse = np.zeros((5,1))
# Create the restriction matrix using full weighting
I_restrict = BuildFullWeighting(7)
# Interpolate from the coarse y values into the finer y-values
y_coarse[1:4,0] = mvmult(I_restrict,y_fine[1:8])
# Plot the coarse grid quantities as a blue line and the fine grid as red circles
fig, axs = plt.subplots(2,1)
axs[0].plot(x_fine, y_fine, '-', x_fine, y_fine,'ro')
axs[1].plot(x_coarse,y_coarse,'-',x_coarse, y_coarse, 'ro')
# + [markdown] slideshow={"slide_type": "subslide"}
# While, for simplicity in presentation, we will not be presenting a 2D problem, the stencil for the full weighting operator in this case is given:
# $$
# \frac{1}{16}
# \begin{bmatrix}
# 1 & 2 & 1 \\
# 2 & 4 & 2 \\
# 1 & 2 & 1
# \end{bmatrix}
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# ### Interpolation Operator
# Now, let's discuss what's typically referred to as the **interpolation** or **prolongation** operator. It takes vectors in a coarse grid and interpolates them into a finer grid. We will denote it as $I_{2h}^h$ and it produces fine grid vectors according to the rule $I_{2h}^h x^{2h} = x^{h}$ where
# \begin{align*}
# x_{2j}^h &= x_j^{2h} \\
# x_{2j+1}^h &= \frac{1}{2} \left( x_j^{2h} + x_{j+1}^{2h} \right)
# \end{align*}
# In other words, for the shared grid points, we simply let the values coinside and for the additional fine grid points, we use the average of the surrounding coarse grid points. We can describe this transformation with a matrix operator. For example, if we have 8 subintervals in our fine grid (that is 7 interior nodes), and 4 subintervals in our coarse grid (with 3 interior nodes), then we have the following:
# $$
# I_{2h}^h x^{2h} = \frac{1}{2}
# \begin{bmatrix}
# 1 & & \\
# 2 & & \\
# 1 & 1 & \\
# & 2 & \\
# & 1 & 1 \\
# & & 2 \\
# & & 1
# \end{bmatrix}
# \begin{bmatrix}
# x_1 \\ x_2 \\ x_3 \\
# \end{bmatrix}_{2h}
# = \begin{bmatrix}
# x_1 \\ x_2 \\ x_3 \\ x_4 \\ x_5 \\ x_6 \\ x_7
# \end{bmatrix}_h = x^h
# $$
# Another advantage of using the full weighting operator for the restriction operator is that $I_{2h}^h = c(I_h^{2h})^T$. This is an important property for a lot of the theory of multigrid.
# We can also depict this graphically.
# + slideshow={"slide_type": "subslide"}
# Set up the coarse grid first
x_coarse = np.linspace(0,1,5)
y_coarse = np.sin(3*np.pi*x_coarse)
# Now the fine grid
x_fine = np.linspace(0,1,9)
y_fine = np.zeros((9,1))
# Create the prolongation matrix - it's the transpose of the restriction operator created earlier
I_prolong = 2*I_restrict.T
# Interpolate from the coarse y values into the finer y-values
y_fine[1:8,0] = mvmult(I_prolong, y_coarse[1:4])
# Plot the coarse grid quantities as a blue line and the fine grid as red circles
fig, axs = plt.subplots(2,1)
axs[0].plot(x_coarse,y_coarse,'-',x_coarse, y_coarse, 'ro')
axs[1].plot(x_fine, y_fine, '-', x_fine, y_fine,'ro')
# + [markdown] slideshow={"slide_type": "slide"}
# ### Galerkin Projection
# The last piece we need to discuss is the creation of the coarse grid version of the $A^{2h}$ matrix.
# One option is to generate a discretization for the coarse grid as well as the fine grid. This
# technique can be cumbersome, and adds to the effort involved in utilizing the multigrid method.
# Another option is to use the **Galerkin Projection**:
# $$
# A^{2h} = I_h^{2h} A^h I_{2h}^h
# $$
#
# As it turns out, if we use full weighting on the 1D problem, is projection will be the same as the
# projection created by a discretization of the coarse grid. Let's demonstrate this, step-by-step.
#
# First, let $e_j^{2h}$ denote the vector on the coarse grid with a 1 in the $j$th entry, and zeros
# elsewhere. Then $A^{2h}e_j^{2h}$ will be the $j$th column of $A^{2h}$. We will calculate this column
# in steps:
# $$
# I_{2h}^{h}e_j^{2h} =
# \frac{1}{2}
# \begin{bmatrix}
# 1 & & & \\
# 2 & & & \\
# 1 & 1 & & \\
# & 2 & & \\
# & 1 & 1 & \\
# & & 2 & \\
# & & 1 & \ddots \\
# & & & \ddots \\
# & & & \ddots
# \end{bmatrix}
# \begin{bmatrix}
# 0 \\ \vdots \\ 0 \\ 1 \\ 0 \\ \vdots \\ 0
# \end{bmatrix}
# =
# \begin{bmatrix}
# 0 \\ \vdots \\ 0 \\ \frac{1}{2} \\ 1 \\ \frac{1}{2} \\ 0 \\ \vdots \\ 0
# \end{bmatrix}
# $$
# Notice, this vector now lies in the fine grid so we can now apply the fine grid operator $A^h$ to
# this vector:
# $$
# A^h I_{2h}^h e_j^{2h} =
# \frac{1}{h^2}
# \begin{bmatrix}
# 2 & -1 & & & \\
# -1 & 2 & -1 & & \\
# & -1 & 2 & -1 & \\
# & & \ddots & \ddots & \ddots
# \end{bmatrix}
# \begin{bmatrix}
# 0 \\ \vdots \\ 0 \\ \frac{1}{2} \\ 1 \\ \frac{1}{2} \\ 0 \\ \vdots \\ 0
# \end{bmatrix}
# =
# \begin{bmatrix}
# 0 \\ \vdots \\ 0 \\ \frac{-1}{2h^2} \\ \frac{1}{h^2} - \frac{1}{h^2} \\ \frac{-1}{2h^2} + \frac{2}{h^2}
# - \frac{1}{2h^2} \\ \frac{-1}{h^2} + \frac{1}{h^2} \\ \frac{-1}{2h^2} \\ 0 \\ \vdots \\ 0
# \end{bmatrix}
# = \begin{bmatrix}
# 0 \\ \vdots \\ 0 \\ \frac{-1}{2h^2} \\ 0 \\ \frac{1}{h^2} \\ 0 \\ \frac{-1}{2h^2} \\ 0 \\ \vdots \\ 0
# \end{bmatrix}
# $$
# Finally, we apply the restriction operator to this vector to obtain a vector in the course grid space:
# $$
# I_{h}^{2h} A^{h} I_{2h}^h e_j^{2h} =
# \begin{bmatrix}
# \frac{1}{4} & \frac{1}{2} & \frac{1}{4} & & & & & & \\
# & & \frac{1}{4} & \frac{1}{2} & \frac{1}{4} & & & & \\
# & & & & \frac{1}{4} & \frac{1}{2} & \frac{1}{4} & & \\
# & & & & & & \ddots & \ddots & \ddots \\
# \end{bmatrix}
# \begin{bmatrix}
# 0 \\ \vdots \\ 0 \\ \frac{-1}{2h^2} \\ 0 \\ \frac{1}{h^2} \\ 0 \\ \frac{-1}{2h^2} \\ 0 \\ \vdots \\ 0
# \end{bmatrix}
# =
# \begin{bmatrix}
# 0 \\ \vdots \\ 0 \\ \frac{-1}{4h^2} \\ \frac{1}{2h^2} \\ \frac{-1}{4h^2} \\ 0 \\ \vdots \\ 0
# \end{bmatrix}
# =
# \begin{bmatrix}
# 0 \\ \vdots \\ 0 \\ \frac{-1}{(2h)^2} \\ \frac{2}{(2h)^2} \\ \frac{-1}{(2h)^2} \\ 0 \\ \vdots \\ 0
# \end{bmatrix}
# $$
# Notice that this is exactly the same column we obtain from creating discretization on the coarse grid. This projection will not be the same as the coarse grid discretization in a 2D problem or if full weighting is not used. Nevertheless, it is a common practice and has been shown to produce good results. It also has the advantage that it requires no extra effort on the part of the user, it can simply be another step in the algorithm.
# + [markdown] slideshow={"slide_type": "slide"}
# ## A Formal Two-Grid Cycle
# (in Briggs, this is called a Coarse Grid Correction Scheme)
#
# 1. Relax $\nu_1$ times on $A^h x^h = b^h$ on $\Omega^h$ with initial guess $x^h$
# 2. Compute $r^{2h} = I_h^{2h}(b^h - A^h x^h)$.
# 3. Solve $A^{2h} e^{2h} = r^{2h}$ on $\Omega^{2h}$
# 4. Correct fine grid approximation: $x^h \leftarrow x^h + I_{2h}^h e^{2h}$
# 5. Relax $\nu_2$ times on $A^h x^h = b^h$ on $\Omega^h$ with initial guess $x^h$
# + slideshow={"slide_type": "subslide"}
# Import CG code
import os
import sys
module_path = os.path.abspath(os.path.join('../CGProgrammingProblem'))
if module_path not in sys.path:
sys.path.append(module_path)
from PCG import PCG
def TwoGridScheme(A_fine, b, numPreRelax, numPostRelax, numiters=1):
# For simplicity, we assume A_fine is (2^n-1) by (2^n-1) for some n
# We will also assume that A is SPD so that we can use CG to solve the coarse system
# Build the restriction and prolongation operators
# They can be re-used if we run more than 1 iteration
I_Restrict = BuildFullWeighting(A_fine.shape[0])
I_Prolong = 2*I_Restrict.T
# Use an initial guess of zero
x = np.zeros_like(b)
# The coarse A only needs to be calculated once, using Galerkin Projection
A_coarse = I_Restrict.dot(A_fine.dot(I_Prolong))
# We could run this scheme more than once if more accuracy is required
for i in range(numiters):
# First we relax on the fine grid:
x = Jacobi(x, A_fine, b, numiters=numPreRelax)
# Now compute the restricted residual
r_coarse = mvmult(I_Restrict, b - mvmult(A_fine, x))
# Now we solve the coarse problem Ae = r, which is
# cheaper than solving the fine grid problem due to the decrease in size
# We use the initial guess of zero
(conv, _, e_coarse, _, _) = PCG(A_coarse, r_coarse, maxiter=100000)
if not conv:
raise RuntimeError("PCG did not converge on the coarse_grid")
# Correct the fine-grid x with the prolongated residual
x += mvmult(I_Prolong, e_coarse)
# The above Prolongation could be introducing additional high frequency errors
# So we relax again to get rid of them
x = Jacobi(x, A_fine, b, numiters=numPostRelax)
return x
# + [markdown] slideshow={"slide_type": "subslide"}
# Let's test it out. We will generate `A_fine` to be $(2^{16}-1)\times(2^{16}-1)$. We use the method of manufactured solutions to generate $b$ from a random true solution $x^*$ and start with $x_0 = 0$ for an initial guess. We will run 5 iterations of Jacobi before and after the transfer to the coarse grid.
# + slideshow={"slide_type": "subslide"}
N = 2**16
A_fine = GeneratePoisson(N)
xTrue = rs.rand(N-1)
b = mvmult(A_fine, xTrue)
x = np.zeros_like(b)
results = PrettyTable()
results.field_names = ["Algorithm", "Num. Iterations", "Rel. Error", "Time (sec)"]
results.align = "l"
# Run Jacobi - 1000 iterations
startT = time.time()
x_Jac = Jacobi(np.zeros_like(b), A_fine, b, 100)
endT = time.time()
relError_Jac = norm(x_Jac - xTrue)/norm(xTrue)
results.add_row(["Jacobi", 100, relError_Jac, endT-startT ])
# Run TwoGrid - 1 pre and 1 post relax, 1 iteration
startT = time.time()
x_TG = TwoGridScheme(A_fine, b, 1, 1, 1)
endT = time.time()
relError = norm(x_TG - xTrue)/norm(xTrue)
results.add_row(["Two Grid (1 pre, 1 post)", 1, relError, endT-startT])
# Run TwoGrid - 1 pre and 1 post relax, 3 iteration
startT = time.time()
x_TG = TwoGridScheme(A_fine, b, 1, 1, 3)
endT = time.time()
relError = norm(x_TG - xTrue)/norm(xTrue)
results.add_row(["Two Grid (1 pre, 1 post)", 3, relError, endT-startT])
# Run TwoGrid - 3 pre and 3 post relax, 1 iteration
startT = time.time()
x_TG = TwoGridScheme(A_fine, b, 3, 3, 1)
endT = time.time()
relError = norm(x_TG - xTrue)/norm(xTrue)
results.add_row(["Two Grid (3 pre, 3 post)", 1, relError, endT-startT])
print(results)
display(HTML(results.get_html_string()))
# + [markdown] slideshow={"slide_type": "subslide"}
# While these numbers look impressive, you can't read too much into it since we are actually cheating a little bit by using Conjugate Gradient on the coarse level. There is some hope, however, looking at the last two rows. The run that completes more relaxation and few CG solves acheives the same error in half the time. Let's see how long it takes CG to solve the fine grid problem to the same relative residual and try using more relaxations for the Two Grid method.
# + slideshow={"slide_type": "subslide"}
# Run TwoGrid - 5 pre and 5 post relax, 1 iteration
startT = time.time()
x_TG = TwoGridScheme(A_fine, b, 5, 5, 1)
endT = time.time()
relError = norm(x_TG - xTrue)/norm(xTrue)
results.add_row(["Two Grid (5 pre, 5 post)", 1, relError, endT-startT])
# Run CG to the similar relative error
startT = time.time()
(conv, iters_CG, x_CG, _, _) = PCG(A_fine, b, maxiter=100000, tau=1.8e-9)
endT = time.time()
relError = norm(x_CG - xTrue)
results.add_row(["CG", iters_CG, relError, endT-startT])
display(HTML(results.get_html_string()))
# + [markdown] slideshow={"slide_type": "subslide"}
# Looking at these results, we see that we get the same error, but about a 2x speedup if we use CG on the coarse grid then just use Jacobi on the fine grid to refine that solution. Still, there's more we can do to make this better. If you look at step 3 in the algorithm above, you'll notice we do a linear system solve on the coarse grid, but this solve is also of the form $Ax = b$. That means we could apply this process __recursively__ which is where the real power of Multigrid appears.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Typical Multigrid Cycles
# There are several ways to create a recursive multigrid pattern. The most common by far is known as the V-Cycle.
# ### V-Cycle
# In the V-Cycle, we run the above Two-Grid Cycle, but replace the solve on the coarse grid with a recursive call another Two-Grid Cycle, where we move to yet another coarser mesh. This is called a V-Cycle because if you trace the work done on the various grid levels, you get a picture in the shape of a V:
#
# <img src="Graphics/V-Cycle-Graphic.png" alt="V-Cycle" style="width: 300px;"/>
#
# We can get this cycle by modifying the `TwoGridScheme` function above:
# + slideshow={"slide_type": "slide"}
def VCycle(A_fine, b, numPreRelax, numPostRelax, coarsest_N, numiters=1, x=None):
# For simplicity, we assume A_fine is (2^n-1) by (2^n-1)
# and A_coarse is (2^(n-1)-1) by (2^(n-1)-1) for some n
#
# We will also assume that A is SPD so that we can use CG to solve the coarse system
#
# It should be noted that this implementation is not best to use if numiters is not 1
# since we are not caching the calculated A, I_restrict, I_prolong matrices
# here we re-calculate them during each V, doing much extra computation
# Build the restriction and prolongation operators
# They can be re-used if we run more than 1 iteration
N = A_fine.shape[0]
I_Restrict = BuildFullWeighting(N)
I_Prolong = 2*I_Restrict.T
# start with the initial guess of zero if one isn't given
if x is None:
x = np.zeros_like(b)
# Calculate the coarse mesh
A_coarse = I_Restrict.dot(A_fine.dot(I_Prolong))
N_coarse = A_coarse.shape[0]
# We could run more than once if more accuracy is required
for i in range(numiters):
# First we relax on the fine grid:
x = Jacobi(x, A_fine, b, numiters=numPreRelax)
# Now compute the restricted residual
r_coarse = mvmult(I_Restrict, b - mvmult(A_fine, x))
# If not on the "bottom of the V", we call recursively
if N_coarse > coarsest_N:
# We start with an initial guess of zero, only 1 iteration to get the V-Cycle
e_coarse = VCycle(A_coarse, r_coarse, numPreRelax, numPostRelax, coarsest_N, 1)
else: # If on the bottom of the V, we solve the coarsest matrix exactly
(conv, _, e_coarse, _, _) = PCG(A_coarse, r_coarse, maxiter=100000)
if not conv:
raise RuntimeError("PCG did not converge on the coarse_grid")
# Correct the fine-grid x with the prolongated residual
x += mvmult(I_Prolong, e_coarse)
# The above Prolongation could be introducing additional high frequency errors
# So we relax again to get rid of them
x = Jacobi(x, A_fine, b, numiters=numPostRelax)
return x
# + slideshow={"slide_type": "subslide"}
# Run VCycle
startT = time.time()
x_VCyc = VCycle(A_fine, b, 3, 3, 128, numiters=1)
endT = time.time()
relError = norm(x_VCyc - xTrue)/norm(xTrue)
results.add_row(["V-Cycle (3 pre, 3 post, 127x127 coarse)", 1, relError, endT-startT])
# Run VCycle
startT = time.time()
x_VCyc = VCycle(A_fine, b, 3, 3, 128, numiters=3)
endT = time.time()
relError = norm(x_VCyc - xTrue)/norm(xTrue)
results.add_row(["V-Cycle (3 pre, 3 post, 127x127 coarse)", 3, relError, endT-startT])
# Run VCycle
startT = time.time()
x_VCyc = VCycle(A_fine, b, 5, 5, 128, numiters=1)
endT = time.time()
relError = norm(x_VCyc - xTrue)/norm(xTrue)
results.add_row(["V-Cycle (5 pre, 5 post, 127x127 coarse)", 1, relError, endT-startT])
display(HTML(results.get_html_string()))
# + [markdown] slideshow={"slide_type": "subslide"}
# This looks like a good improvement, they are the fastest single run so far and achieve about the same error as the other runs. During these runs, I observed that the CPU usage in my multi-core CPU is higher for the some of the computation and gets lower for the coarser meshes. This makes sense, those matrices are smaller and hence, take less computation. This however means that the size of the coarsest grid should make a difference. If the course grid is too small, the CPU is under-utilized, and if the coarse grid is too large, CG will take longer than moving to a coarser grid. Let's see if we can find a more optimal coarse-grid size.
#
# We run trials of 1 V-Cycle with 5 pre and 5 post relaxations for differing coarse matrix sizes:
# + slideshow={"slide_type": "subslide"}
coarseGridSize_results = PrettyTable()
coarseGridSize_results.field_names = ["Coarse Matrix Size", "Rel Error", "Time (sec)"]
coarseGridSize_results.align = "l"
relErrors = np.ones(14)
timings = np.zeros(14)
for exp in range(2,16):
startT = time.time()
x_VCyc = VCycle(A_fine, b, 5, 5, 2**exp, numiters=1)
endT = time.time()
relErrors[exp-2] = norm(x_VCyc - xTrue)/norm(xTrue)
timings[exp-2] = endT-startT
coarseGridSize_results.add_row([f'{2**exp - 1}x{2**exp-1}', relErrors[exp-2], timings[exp-2]])
display(HTML(coarseGridSize_results.get_html_string()))
# + [markdown] slideshow={"slide_type": "fragment"}
# It appears that a $8191\times 8191$ matrix is the most efficient coarse grid size for this computer. Any larger and the CG method takes too long, either due to cache size, number of cache misses, or simply the number of iterations CG needs to converge for the coarse problem (due to the increased condition number). Let's add this run to our table to see all the results together:
# + slideshow={"slide_type": "subslide"}
results.add_row(["V-Cycle (5 pre, 5 post, 8191x8191 coarse)", 1, relErrors[11], timings[11]])
display(HTML(results.get_html_string()))
# + [markdown] slideshow={"slide_type": "subslide"}
# Finally, let's run some extra iterations to see what the convergence looks like. Since the code below interupts the V-Cycle function after every V-Cycle, we are doing extra work and so the timings are not representative, hence we won't calculate them.
# + slideshow={"slide_type": "subslide"}
maxIters = 30
coarseGridSize = 2**13
numRelax = 5
# container to hold the errors
relError = np.ones(maxIters+1)
# Provide an initial guess
x_VCyc = np.zeros_like(b)
for i in range(1,maxIters+1):
x_VCyc = VCycle(A_fine, b, numRelax, numRelax, coarseGridSize, numiters=1, x=x_VCyc)
relError[i] = norm(x_VCyc - xTrue)/norm(xTrue)
plt.plot(relError)
# -
# Notice the large decrease in error from the first iteration (80% reduction). This is one of the primary reasons why one iteration of multigrid is widely used as a preconditioner.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Other Multigrid Cycles
# While the V-Cycle is the most popular, there are other proposed cycles as well. One possible extension is to recursively run two consecutive V-Cycles:
#
# <img src="Graphics/W-Cycle-Graphic.png" alt="W-Cycle" style="width: 400px;"/>
#
# This is typically called a **W-Cycle**. You can of course extend this to running more than 2 consecutive V-Cycles, Briggs's book calls these **$\mu$-Cycles** (where $\mu$ refers to the number of consecutive V-Cycles completed recursively).
#
# Finally, there is the **Full Multigrid Cycle**:
#
# <img src="Graphics/FMV-Cycle-Graphic.png" alt="FMV-Cycle" style="width: 400px;"/>
#
# The idea behind the full multigrid cycle is to first solve on the coarse grid, getting a good starting guess for the next finer grid. Then run a V-Cycle on that grid to get a good starting point for the next finer grid, and continue that process until the finest grid is reached.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Tuning Multigrid
# There are several "knobs to turn" to tune Multigrid Methods:
# - Several different relaxation schemes have been shown to be effective: weighted Jacobi, Red-Black Jacobi, Gauss-Seidel, Red-Black Gauss Seidel, SOR, Block Jacobi, Block Gauss-Seidel
# - Method for solving the coarsest grid problem can be chosen
# - The number of relaxations can have some effect in the convergence of the method, typically 3-5 are used
# - Which type of cycle to use: the most common is the V-Cycle, but the W and Full Multigrid Cycle are also common, $\mu$-cycles with $\mu \geq 3$ are rarely seen
# -
# # Cons of Multigrid
# While the multigrid method has been shown to be effective in terms of computational time, it does cost more in terms of memory. This is due to the fact that all grids need to be in storage as once. The cost here is migated however since the dimensions of the coarse matrices decreases exponentially.
#
# Another negative aspect of multrigrid is the fact that it is not as effective on smaller matrices. For example, straight CG is often faster than multigrid for smaller matrix sizes, where CG does not have to complete as many iterations.
# + [markdown] slideshow={"slide_type": "slide"}
# # Algebraic Multigrid
# While geometric multigrid is useful for gaining intuition into multigrid methods, it's not often used in practice. It's tougher to design the restriction and prolongation operators for non-uniform meshes where the number of bordering nodes is variable. It's also less useful for systems with more than one state variable, since only the physical dimensions can be made coarser. Instead, we will use the same idea to develop a multigrid method that doesn't explicity depend on the mesh, but instead depends on the coefficient matrix.
#
# If we look at our matrix
# $$
# A = \frac{1}{h^2}
# \begin{bmatrix}
# 2 & -1 & & & & \\
# -1 & 2 & -1 & & & \\
# & -1 & 2 & -1 & & \\
# & & \ddots & \ddots & \ddots & \\
# & & & -1 & 2 & -1 \\
# & & & & -1 & 2
# \end{bmatrix}
# $$
# we can interpret it in the following way: an entry's magnitude in the matrix corresponds to its level of contribution in calculating the element on the diagonal. For example, row 2 has $-1/h^2$, $2/h^2$, and $-1/h^2$ in the first three columns. This signifies that only $x_1, x_2, x_3$ directly contribute to the node $x_2$, with the value $x_2$ contributing more than $x_1$ and $x_3$. In algebriac multigrid, we use this idea of "significance" to determine which unknowns can be "merged" to obtain a coarse matrix. This process will also create prolongation and restriction operators which only depend on the coefficient matrix and not on the geometric structure of the physical problem. Algebraic multigrid can therefore be programmed in a more general way and can more easily extended to more problems. This property also contributes to its usefulness as a preconditioner since it takes less setup and quickly gives modest accuracy.
| MultigridPresentation/Multigrid Presentation.ipynb |