code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# <img src="../static/images/joinnode.png" width="240">
#
# # JoinNode
#
# JoinNode have the opposite effect of [iterables](basic_iteration.ipynb). Where `iterables` split up the execution workflow into many different branches, a JoinNode merges them back into on node. For a more detailed explanation, check out [JoinNode, synchronize and itersource](http://nipype.readthedocs.io/en/latest/users/joinnode_and_itersource.html) from the main homepage.
# ## Simple example
#
# Let's consider the very simple example depicted at the top of this page:
# ```python
# from nipype import Node, JoinNode, Workflow
#
# # Specify fake input node A
# a = Node(interface=A(), name="a")
#
# # Iterate over fake node B's input 'in_file?
# b = Node(interface=B(), name="b")
# b.iterables = ('in_file', [file1, file2])
#
# # Pass results on to fake node C
# c = Node(interface=C(), name="c")
#
# # Join forked execution workflow in fake node D
# d = JoinNode(interface=D(),
# joinsource="b",
# joinfield="in_files",
# name="d")
#
# # Put everything into a workflow as usual
# workflow = Workflow(name="workflow")
# workflow.connect([(a, b, [('subject', 'subject')]),
# (b, c, [('out_file', 'in_file')])
# (c, d, [('out_file', 'in_files')])
# ])
# ```
# As you can see, setting up a ``JoinNode`` is rather simple. The only difference to a normal ``Node`` are the ``joinsource`` and the ``joinfield``. ``joinsource`` specifies from which node the information to join is coming and the ``joinfield`` specifies the input field of the JoinNode where the information to join will be entering the node.
# ## More realistic example
#
# Let's consider another example where we have one node that iterates over 3 different numbers and generates randome numbers. Another node joins those three different numbers (each coming from a separate branch of the workflow) into one list. To make the whole thing a bit more realistic, the second node will use the ``Function`` interface to do something with those numbers, before we spit them out again.
from nipype import JoinNode, Node, Workflow
from nipype.interfaces.utility import Function, IdentityInterface
# +
def get_data_from_id(id):
"""Generate a random number based on id"""
import numpy as np
return id + np.random.rand()
def merge_and_scale_data(data2):
"""Scale the input list by 1000"""
import numpy as np
return (np.array(data2) * 1000).tolist()
node1 = Node(Function(input_names=['id'],
output_names=['data1'],
function=get_data_from_id),
name='get_data')
node1.iterables = ('id', [1, 2, 3])
node2 = JoinNode(Function(input_names=['data2'],
output_names=['data_scaled'],
function=merge_and_scale_data),
name='scale_data',
joinsource=node1,
joinfield=['data2'])
# -
wf = Workflow(name='testjoin')
wf.connect(node1, 'data1', node2, 'data2')
eg = wf.run()
wf.write_graph(graph2use='exec')
from IPython.display import Image
Image(filename='graph_detailed.png')
# Now, let's look at the input and output of the joinnode:
res = [node for node in eg.nodes() if 'scale_data' in node.name][0].result
res.outputs
res.inputs
# ## Extending to multiple nodes
#
# We extend the workflow by using three nodes. Note that even this workflow, the joinsource corresponds to the node containing iterables and the joinfield corresponds to the input port of the JoinNode that aggregates the iterable branches. As before the graph below shows how the execution process is setup.
# +
def get_data_from_id(id):
import numpy as np
return id + np.random.rand()
def scale_data(data2):
import numpy as np
return data2
def replicate(data3, nreps=2):
return data3 * nreps
node1 = Node(Function(input_names=['id'],
output_names=['data1'],
function=get_data_from_id),
name='get_data')
node1.iterables = ('id', [1, 2, 3])
node2 = Node(Function(input_names=['data2'],
output_names=['data_scaled'],
function=scale_data),
name='scale_data')
node3 = JoinNode(Function(input_names=['data3'],
output_names=['data_repeated'],
function=replicate),
name='replicate_data',
joinsource=node1,
joinfield=['data3'])
# -
wf = Workflow(name='testjoin')
wf.connect(node1, 'data1', node2, 'data2')
wf.connect(node2, 'data_scaled', node3, 'data3')
eg = wf.run()
wf.write_graph(graph2use='exec')
Image(filename='graph_detailed.png')
# ### Exercise 1
#
# You have list of DOB of the subjects in a few various format : ``["10 February 1984", "March 5 1990", "April 2 1782", "June 6, 1988", "12 May 1992"]``, and you want to sort the list.
#
# You can use ``Node`` with ``iterables`` to extract day, month and year, and use [datetime.datetime](https://docs.python.org/2/library/datetime.html) to unify the format that can be compared, and ``JoinNode`` to sort the list.
# + solution2="hidden" solution2_first=true
# write your solution here
# + solution2="hidden"
# the list of all DOB
dob_subjects = ["10 February 1984", "March 5 1990", "April 2 1782", "June 6, 1988", "12 May 1992"]
# + solution2="hidden"
# let's start from creating Node with iterable to split all strings from the list
from nipype import Node, JoinNode, Function, Workflow
def split_dob(dob_string):
return dob_string.split()
split_node = Node(Function(input_names=["dob_string"],
output_names=["split_list"],
function=split_dob),
name="splitting")
#split_node.inputs.dob_string = "10 February 1984"
split_node.iterables = ("dob_string", dob_subjects)
# + solution2="hidden"
# and now let's work on the date format more, independently for every element
# sometimes the second element has an extra "," that we should remove
def remove_comma(str_list):
str_list[1] = str_list[1].replace(",", "")
return str_list
cleaning_node = Node(Function(input_names=["str_list"],
output_names=["str_list_clean"],
function=remove_comma),
name="cleaning")
# now we can extract year, month, day from our list and create ``datetime.datetim`` object
def datetime_format(date_list):
import datetime
# year is always the last
year = int(date_list[2])
#day and month can be in the first or second position
# we can use datetime.datetime.strptime to convert name of the month to integer
try:
day = int(date_list[0])
month = datetime.datetime.strptime(date_list[1], "%B").month
except(ValueError):
day = int(date_list[1])
month = datetime.datetime.strptime(date_list[0], "%B").month
# and create datetime.datetime format
return datetime.datetime(year, month, day)
datetime_node = Node(Function(input_names=["date_list"],
output_names=["datetime"],
function=datetime_format),
name="datetime")
# + solution2="hidden"
# now we are ready to create JoinNode and sort the list of DOB
def sorting_dob(datetime_list):
datetime_list.sort()
return datetime_list
sorting_node = JoinNode(Function(input_names=["datetime_list"],
output_names=["dob_sorted"],
function=sorting_dob),
joinsource=split_node, # this is the node that used iterables for x
joinfield=['datetime_list'],
name="sorting")
# + solution2="hidden"
# and we're ready to create workflow
ex1_wf = Workflow(name="sorting_dob")
ex1_wf.connect(split_node, "split_list", cleaning_node, "str_list")
ex1_wf.connect(cleaning_node, "str_list_clean", datetime_node, "date_list")
ex1_wf.connect(datetime_node, "datetime", sorting_node, "datetime_list")
# + solution2="hidden"
# you can check the graph
from IPython.display import Image
ex1_wf.write_graph(graph2use='exec')
Image(filename='graph_detailed.png')
# + solution2="hidden"
# and run the workflow
ex1_res = ex1_wf.run()
# + solution2="hidden"
# you can check list of all nodes
ex1_res.nodes()
# + solution2="hidden"
# and check the results from sorting_dob.sorting
list(ex1_res.nodes())[0].result.outputs
| notebooks/basic_joinnodes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PySpark
# language: ''
# name: pysparkkernel
# ---
# ## Tuning Model Parameters
#
# In this exercise, you will optimise the parameters for a classification model.
#
# ### Prepare the Data
#
# First, import the libraries you will need and prepare the training and test data:
# +
# Import Spark SQL and Spark ML libraries
from pyspark.sql.types import *
from pyspark.sql.functions import *
from pyspark.ml import Pipeline
from pyspark.ml.classification import LogisticRegression
from pyspark.ml.feature import VectorAssembler
from pyspark.ml.tuning import ParamGridBuilder, TrainValidationSplit
from pyspark.ml.evaluation import BinaryClassificationEvaluator
# Load the source data
csv = spark.read.csv('wasb:///data/flights.csv', inferSchema=True, header=True)
# Select features and label
data = csv.select("DayofMonth", "DayOfWeek", "OriginAirportID", "DestAirportID", "DepDelay", ((col("ArrDelay") > 15).cast("Int").alias("label")))
# Split the data
splits = data.randomSplit([0.7, 0.3])
train = splits[0]
test = splits[1].withColumnRenamed("label", "trueLabel")
# -
# ### Define the Pipeline
# Now define a pipeline that creates a feature vector and trains a classification model
# Define the pipeline
assembler = VectorAssembler(inputCols = ["DayofMonth", "DayOfWeek", "OriginAirportID", "DestAirportID", "DepDelay"], outputCol="features")
lr = LogisticRegression(labelCol="label", featuresCol="features")
pipeline = Pipeline(stages=[assembler, lr])
# ### Tune Parameters
# You can tune parameters to find the best model for your data. A simple way to do this is to use **TrainValidationSplit** to evaluate each combination of parameters defined in a **ParameterGrid** against a subset of the training data in order to find the best performing parameters.
# +
paramGrid = ParamGridBuilder().addGrid(lr.regParam, [0.3, 0.1, 0.01]).addGrid(lr.maxIter, [10, 5]).addGrid(lr.threshold, [0.35, 0.30]).build()
tvs = TrainValidationSplit(estimator=pipeline, evaluator=BinaryClassificationEvaluator(), estimatorParamMaps=paramGrid, trainRatio=0.8)
model = tvs.fit(train)
# -
# ### Test the Model
# Now you're ready to apply the model to the test data.
prediction = model.transform(test)
predicted = prediction.select("features", "prediction", "probability", "trueLabel")
predicted.show(100)
# ### Compute Confusion Matrix Metrics
# Classifiers are typically evaluated by creating a *confusion matrix*, which indicates the number of:
# - True Positives
# - True Negatives
# - False Positives
# - False Negatives
#
# From these core measures, other evaluation metrics such as *precision* and *recall* can be calculated.
tp = float(predicted.filter("prediction == 1.0 AND truelabel == 1").count())
fp = float(predicted.filter("prediction == 1.0 AND truelabel == 0").count())
tn = float(predicted.filter("prediction == 0.0 AND truelabel == 0").count())
fn = float(predicted.filter("prediction == 0.0 AND truelabel == 1").count())
metrics = spark.createDataFrame([
("TP", tp),
("FP", fp),
("TN", tn),
("FN", fn),
("Precision", tp / (tp + fp)),
("Recall", tp / (tp + fn))],["metric", "value"])
metrics.show()
# ### Review the Area Under ROC
# Another way to assess the performance of a classification model is to measure the area under a ROC curve for the model. the spark.ml library includes a **BinaryClassificationEvaluator** class that you can use to compute this.
evaluator = BinaryClassificationEvaluator(labelCol="trueLabel", rawPredictionCol="prediction", metricName="areaUnderROC")
aur = evaluator.evaluate(prediction)
print "AUR = ", aur
| notebooks/06. Parameter Tuning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib notebook
from asteria import config, detector
import astropy.units as u
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
mpl.rc('font', size=12)
# -
# ## Load Configuration
#
# This will load the source configuration from a file.
#
# For this to work, either the user needs to have done one of two things:
# 1. Run `python setup.py install` in the ASTERIA directory.
# 2. Run `python setup.py develop` and set the environment variable `ASTERIA` to point to the git source checkout.
#
# If these were not done, the initialization will fail because the paths will not be correctly resolved.
conf = config.load_config('../../data/config/default.yaml')
ic86 = detector.initialize(conf)
doms = ic86.doms_table()
doms
# +
fig = plt.figure(figsize=(6,6))
ax = fig.add_subplot(111, projection='3d')
# Plot the DeepCore DOMs
dc = doms['type'] == 'dc'
x, y, z = [doms[coord][dc] for coord in 'xyz']
ax.scatter(x, y, z, alpha=0.7)
# Plot the standard DOMs
i3 = doms['type'] == 'i3'
x, y, z = [doms[coord][i3] for coord in 'xyz']
ax.scatter(x, y, z, alpha=0.2)
ax.set(xlabel='x [m]', ylabel='y [m]', zlabel='z [m]')
ax.view_init(30, -40)
fig.tight_layout()
# -
| docs/nb/detector.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ARDUAIR: Procesamiento de datos PM10
# Se procesan los datos para obtener un archivo CSV con los promedios hora de las lecturas de pm10 de las fechas 17, 18 y 19 de abril de 2017
#
# Se filtran los datos extraños por encima de (lecturas por encima de 1000000) para este caso
#
# ## importacion de librerias
import pandas as pd
import numpy as np
import datetime as dt
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
pd.options.mode.chained_assignment = None
# +
#load
data=pd.read_csv('DATA.TXT',names=['year','month','day','hour','minute','second','hum','temp','pr','l','co','so2','no2','o3','pm10','pm25','void'])
#Dates to datetime
dates=data[['year','month','day','hour','minute','second']]
dates['year']=dates['year'].add(2000)
dates['minute']=dates['minute'].add(60)
data['datetime']=pd.to_datetime(dates)
#agregation
data=data[['datetime','pm10']]
plt.plot(data.datetime, data.pm10)
# -
# # filtration
#
# Se filtran los datos extraños por encima de (lecturas por encima de 1000000) para este caso
#
#
# +
#load
data=pd.read_csv('DATA.TXT',names=['year','month','day','hour','minute','second','hum','temp','pr','l','co','so2','no2','o3','pm10','pm25','void'])
data=data[data.pm10<1000000]
#Dates to datetime
dates=data[['year','month','day','hour','minute','second']]
dates['year']=dates['year'].add(2000)
dates['minute']=dates['minute'].add(60)
data['datetime']=pd.to_datetime(dates)
data=data[['datetime','pm10']]
#agregation
data1=data.groupby(pd.Grouper(key='datetime',freq='1h',axis=1)).mean()
data2=data.groupby(pd.Grouper(key='datetime',freq='2h',axis=1)).mean()
data3=data.groupby(pd.Grouper(key='datetime',freq='3h',axis=1)).mean()
# save
data1.to_csv('arduair_pm10_promedio_1h.csv')
data2.to_csv('arduair_pm10_promedio_2h.csv')
data3.to_csv('arduair_pm10_promedio_3h.csv')
# reset index
data=data.reset_index()
data1=data1.reset_index()
data2=data2.reset_index()
data2=data2.reset_index()
plt.plot(data1.datetime, data1.pm10)
# -
# ## ¿Que pasa si eliminamos los valores en 0.00?
#
# Realizando la correlacion con los datos generados por el Dusttrack,la correlacion es 0.3 comparada con 0.48 de los datos originales
# +
data_sin_ceros=data[data.pm10>0]
plt.plot(data_sin_ceros.datetime, data_sin_ceros.pm10)
data_sin_ceros_1h=data_sin_ceros.groupby(pd.Grouper(key='datetime',freq='1h',axis=1)).mean()
# -
| pm10_upb/arduair_data_processing_PM10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: mykernel
# language: python
# name: mykernel
# ---
# ## _*Quantum SVM (quantum kernel method)*_
#
# ### Introduction
#
# Please refer to [this file](https://github.com/Qiskit/qiskit-tutorials/blob/master/qiskit/aqua/artificial_intelligence/qsvm_kernel_classification.ipynb) for introduction.
#
# In this file, we show two ways for using the quantum kernel method: (1) the non-programming way and (2) the programming way.
#
# ### Part I: non-programming way.
# In the non-programming way, we config a json-like configuration, which defines how the svm instance is internally constructed. After the execution, it returns the json-like output, which carries the important information (e.g., the details of the svm instance) and the processed results.
from datasets import *
from qiskit import Aer
from qiskit_aqua.utils import split_dataset_to_data_and_labels, map_label_to_class_name
from qiskit_aqua import run_algorithm, QuantumInstance
from qiskit_aqua.input import SVMInput
from qiskit_aqua.components.feature_maps import SecondOrderExpansion
from qiskit_aqua.algorithms import QSVMKernel
# First we prepare the dataset, which is used for training, testing and the finally prediction.
#
# *Note: You can easily switch to a different dataset, such as the Breast Cancer dataset, by replacing 'ad_hoc_data' to 'Breast_cancer' below.*
# +
feature_dim = 2 # dimension of each data point
training_dataset_size = 20
testing_dataset_size = 10
random_seed = 10598
shots = 1024
sample_Total, training_input, test_input, class_labels = ad_hoc_data(training_size=training_dataset_size,
test_size=testing_dataset_size,
n=feature_dim, gap=0.3, PLOT_DATA=False)
datapoints, class_to_label = split_dataset_to_data_and_labels(test_input)
print(class_to_label)
# -
# Now we create the svm in the non-programming way.
# In the following json, we config:
# - the algorithm name
# - the feature map
params = {
'problem': {'name': 'svm_classification', 'random_seed': random_seed},
'algorithm': {
'name': 'QSVM.Kernel'
},
'backend': {'shots': shots},
'feature_map': {'name': 'SecondOrderExpansion', 'depth': 2, 'entanglement': 'linear'}
}
backend = Aer.get_backend('qasm_simulator')
algo_input = SVMInput(training_input, test_input, datapoints[0])
# With everything setup, we can now run the algorithm.
#
# The run method includes training, testing and predict on unlabeled data.
#
# For the testing, the result includes the success ratio.
#
# For the prediction, the result includes the predicted class names for each data.
#
# After that the trained model is also stored in the svm instance, you can use it for future prediction.
result = run_algorithm(params, algo_input, backend=backend)
# +
print("kernel matrix during the training:")
kernel_matrix = result['kernel_matrix_training']
img = plt.imshow(np.asmatrix(kernel_matrix),interpolation='nearest',origin='upper',cmap='bone_r')
plt.show()
print("testing success ratio: ", result['testing_accuracy'])
print("predicted classes:", result['predicted_classes'])
# -
# ### part II: Programming way.
# We construct the svm instance directly from the classes. The programming way offers the users better accessibility, e.g., the users can access the internal state of svm instance or invoke the methods of the instance. We will demonstrate this advantage soon.
# Now we create the svm in the programming way.
# - We build the svm instance by instantiating the class QSVMKernel.
# - We build the feature map instance (required by the svm instance) by instantiating the class SecondOrderExpansion.
backend = Aer.get_backend('qasm_simulator')
feature_map = SecondOrderExpansion(num_qubits=feature_dim, depth=2, entangler_map={0: [1]})
svm = QSVMKernel(feature_map, training_input, test_input, None)# the data for prediction can be feeded later.
svm.random_seed = random_seed
quantum_instance = QuantumInstance(backend, shots=shots, seed=random_seed, seed_mapper=random_seed)
result = svm.run(quantum_instance)
# Let us check the result.
# +
print("kernel matrix during the training:")
kernel_matrix = result['kernel_matrix_training']
img = plt.imshow(np.asmatrix(kernel_matrix),interpolation='nearest',origin='upper',cmap='bone_r')
plt.show()
print("testing success ratio: ", result['testing_accuracy'])
# -
# Different from the non-programming way, the programming way allows the users to invoke APIs upon the svm instance directly. In the following, we invoke the API "predict" upon the trained svm instance to predict the labels for the newly provided data input.
#
# Use the trained model to evaluate data directly, and we store a `label_to_class` and `class_to_label` for helping converting between label and class name
# +
predicted_labels = svm.predict(datapoints[0])
predicted_classes = map_label_to_class_name(predicted_labels, svm.label_to_class)
print("ground truth: {}".format(datapoints[1]))
print("preduction: {}".format(predicted_labels))
| community/aqua/artificial_intelligence/qsvm_kernel_directly.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # %latex
# # What's an activation function
# In artificial neural networks, the activation function of a node defines the output of that node given an input or set of inputs.
| deep_learning/activate_function/Introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
# <script>
# window.dataLayer = window.dataLayer || [];
# function gtag(){dataLayer.push(arguments);}
# gtag('js', new Date());
#
# gtag('config', 'UA-59152712-8');
# </script>
#
# # `GiRaFFE_NRPy`: Source Terms
#
# ## Author: <NAME>
#
# <a id='intro'></a>
#
# **Notebook Status:** <font color=green><b> Validated </b></font>
#
# **Validation Notes:** This code produces the expected results for generated functions.
#
# ## This module presents the functionality of [GiRaFFE_NRPy_Source_Terms.py](../../edit/in_progress/GiRaFFE_NRPy/GiRaFFE_NRPy_Source_Terms.py).
#
# ## Introduction:
# This writes and documents the C code that `GiRaFFE_NRPy` uses to compute the source terms for the right-hand sides of the evolution equations for the unstaggered prescription.
#
# The equations themselves are already coded up in other functions; however, for the $\tilde{S}_i$ source term, we will need derivatives of the metric. It will be most efficient and accurate to take them using the interpolated metric values that we will have calculated anyway; however, we will need to write our derivatives in a nonstandard way within NRPy+ in order to take advantage of this, writing our own code for memory access.
# <a id='toc'></a>
#
# # Table of Contents
# $$\label{toc}$$
#
# This notebook is organized as follows
#
# 1. [Step 1](#stilde_source): The $\tilde{S}_i$ source term
# 1. [Step 2](#code_validation): Code Validation against original C code
# 1. [Step 3](#latex_pdf_output): Output this notebook to $\LaTeX$-formatted PDF file
# +
# Step 0: Add NRPy's directory to the path
# https://stackoverflow.com/questions/16780014/import-file-from-parent-directory
import os,sys
nrpy_dir_path = os.path.join("..")
if nrpy_dir_path not in sys.path:
sys.path.append(nrpy_dir_path)
import cmdline_helper as cmd
outdir = os.path.join("GiRaFFE_NRPy","GiRaFFE_Ccode_validation","RHSs")
cmd.mkdir(outdir)
# -
# <a id='stilde_source'></a>
#
# ## Step 1: The $\tilde{S}_i$ source term \[Back to [top](#toc)\]
# $$\label{stilde_source}$$
#
# We start in the usual way - import the modules we need. We will also import the Levi-Civita symbol from `indexedexp.py` and use it to set the Levi-Civita tensor $\epsilon^{ijk} = [ijk]/\sqrt{\gamma}$.
# +
# Step 1: The StildeD RHS *source* term
from outputC import outputC, outCfunction # NRPy+: Core C code output module
import indexedexp as ixp # NRPy+: Symbolic indexed expression (e.g., tensors, vectors, etc.) support
import GRHD.equations as GRHD # NRPy+: Generate general relativistic hydrodynamics equations
import GRFFE.equations as GRFFE # NRPy+: Generate general relativistic force-free electrodynamics equations
thismodule = "GiRaFFE_NRPy_Source_Terms"
def generate_memory_access_code():
# There are several pieces of C code that we will write ourselves because we need to do things
# a little bit outside of what NRPy+ is built for.
# First, we will write general memory access. We will read in values from memory at a given point
# for each quantity we care about.
global general_access
general_access = ""
for var in ["GAMMADD00", "GAMMADD01", "GAMMADD02",
"GAMMADD11", "GAMMADD12", "GAMMADD22",
"BETAU0", "BETAU1", "BETAU2","ALPHA",
"BU0","BU1","BU2",
"VALENCIAVU0","VALENCIAVU1","VALENCIAVU2"]:
lhsvar = var.lower().replace("dd","DD").replace("u","U").replace("bU","BU").replace("valencia","Valencia")
# e.g.,
# const REAL gammaDD00dD0 = auxevol_gfs[IDX4S(GAMMA_FACEDD00GF,i0,i1,i2)];
general_access += "const REAL "+lhsvar+" = auxevol_gfs[IDX4S("+var+"GF,i0,i1,i2)];\n"
# This quick function returns a nearby point for memory access. We need this because derivatives are not local operations.
def idxp1(dirn):
if dirn==0:
return "i0+1,i1,i2"
if dirn==1:
return "i0,i1+1,i2"
if dirn==2:
return "i0,i1,i2+1"
# Next we evaluate needed derivatives of the metric, based on their values at cell faces
global metric_deriv_access
metric_deriv_access = []
for dirn in range(3):
metric_deriv_access.append("")
for var in ["GAMMA_FACEDDdD00", "GAMMA_FACEDDdD01", "GAMMA_FACEDDdD02",
"GAMMA_FACEDDdD11", "GAMMA_FACEDDdD12", "GAMMA_FACEDDdD22",
"BETA_FACEUdD0", "BETA_FACEUdD1", "BETA_FACEUdD2","ALPHA_FACEdD"]:
lhsvar = var.lower().replace("dddd","DDdD").replace("udd","UdD").replace("dd","dD").replace("u","U").replace("_face","")
rhsvar = var.replace("dD","")
# e.g.,
# const REAL gammaDDdD000 = (auxevol_gfs[IDX4S(GAMMA_FACEDD00GF,i0+1,i1,i2)]-auxevol_gfs[IDX4S(GAMMA_FACEDD00GF,i0,i1,i2)])/dxx0;
metric_deriv_access[dirn] += "const REAL "+lhsvar+str(dirn)+" = (auxevol_gfs[IDX4S("+rhsvar+"GF,"+idxp1(dirn)+")]-auxevol_gfs[IDX4S("+rhsvar+"GF,i0,i1,i2)])/dxx"+str(dirn)+";\n"
metric_deriv_access[dirn] += "REAL Stilde_rhsD"+str(dirn)+";\n"
# This creates the C code that writes to the Stilde_rhs direction specified.
global write_final_quantity
write_final_quantity = []
for dirn in range(3):
write_final_quantity.append("")
write_final_quantity[dirn] += "rhs_gfs[IDX4S(STILDED"+str(dirn)+"GF,i0,i1,i2)] += Stilde_rhsD"+str(dirn)+";"
def write_out_functions_for_StildeD_source_term(outdir,outCparams,gammaDD,betaU,alpha,ValenciavU,BU,sqrt4pi):
generate_memory_access_code()
# First, we declare some dummy tensors that we will use for the codegen.
gammaDDdD = ixp.declarerank3("gammaDDdD","sym01",DIM=3)
betaUdD = ixp.declarerank2("betaUdD","nosym",DIM=3)
alphadD = ixp.declarerank1("alphadD",DIM=3)
# We need to rerun a few of these functions with the reset lists to make sure these functions
# don't cheat by using analytic expressions
GRHD.compute_sqrtgammaDET(gammaDD)
GRHD.u4U_in_terms_of_ValenciavU__rescale_ValenciavU_by_applying_speed_limit(alpha, betaU, gammaDD, ValenciavU)
GRFFE.compute_smallb4U(gammaDD, betaU, alpha, GRHD.u4U_ito_ValenciavU, BU, sqrt4pi)
GRFFE.compute_smallbsquared(gammaDD, betaU, alpha, GRFFE.smallb4U)
GRFFE.compute_TEM4UU(gammaDD,betaU,alpha, GRFFE.smallb4U, GRFFE.smallbsquared,GRHD.u4U_ito_ValenciavU)
GRHD.compute_g4DD_zerotimederiv_dD(gammaDD,betaU,alpha, gammaDDdD,betaUdD,alphadD)
GRHD.compute_S_tilde_source_termD(alpha, GRHD.sqrtgammaDET,GRHD.g4DD_zerotimederiv_dD, GRFFE.TEM4UU)
for i in range(3):
desc = "Adds the source term to StildeD"+str(i)+"."
name = "calculate_StildeD"+str(i)+"_source_term"
outCfunction(
outfile = os.path.join(outdir,name+".h"), desc=desc, name=name,
params ="const paramstruct *params,const REAL *auxevol_gfs, REAL *rhs_gfs",
body = general_access \
+metric_deriv_access[i]\
+outputC(GRHD.S_tilde_source_termD[i],"Stilde_rhsD"+str(i),"returnstring",params=outCparams)\
+write_final_quantity[i],
loopopts ="InteriorPoints",
rel_path_to_Cparams=os.path.join("../"))
def add_to_Cfunction_dict__functions_for_StildeD_source_term(outCparams,gammaDD,betaU,alpha,ValenciavU,BU,sqrt4pi,
includes=None, rel_path_to_Cparams=os.path.join("../"),
path_from_rootsrcdir_to_this_Cfunc=os.path.join("RHSs/")):
generate_memory_access_code()
# First, we declare some dummy tensors that we will use for the codegen.
gammaDDdD = ixp.declarerank3("gammaDDdD","sym01",DIM=3)
betaUdD = ixp.declarerank2("betaUdD","nosym",DIM=3)
alphadD = ixp.declarerank1("alphadD",DIM=3)
# We need to rerun a few of these functions with the reset lists to make sure these functions
# don't cheat by using analytic expressions
GRHD.compute_sqrtgammaDET(gammaDD)
GRHD.u4U_in_terms_of_ValenciavU__rescale_ValenciavU_by_applying_speed_limit(alpha, betaU, gammaDD, ValenciavU)
GRFFE.compute_smallb4U(gammaDD, betaU, alpha, GRHD.u4U_ito_ValenciavU, BU, sqrt4pi)
GRFFE.compute_smallbsquared(gammaDD, betaU, alpha, GRFFE.smallb4U)
GRFFE.compute_TEM4UU(gammaDD,betaU,alpha, GRFFE.smallb4U, GRFFE.smallbsquared,GRHD.u4U_ito_ValenciavU)
GRHD.compute_g4DD_zerotimederiv_dD(gammaDD,betaU,alpha, gammaDDdD,betaUdD,alphadD)
GRHD.compute_S_tilde_source_termD(alpha, GRHD.sqrtgammaDET,GRHD.g4DD_zerotimederiv_dD, GRFFE.TEM4UU)
for i in range(3):
desc = "Adds the source term to StildeD"+str(i)+"."
name = "calculate_StildeD"+str(i)+"_source_term"
params ="const paramstruct *params,const REAL *auxevol_gfs, REAL *rhs_gfs",
body = general_access \
+metric_deriv_access[i]\
+outputC(GRHD.S_tilde_source_termD[i],"Stilde_rhsD"+str(i),"returnstring",params=outCparams)\
+write_final_quantity[i],
loopopts ="InteriorPoints",
add_to_Cfunction_dict(
includes=includes,
desc=desc,
name=name, params=params,
body=body, loopopts=loopopts,
rel_path_to_Cparams=rel_path_to_Cparams)
# -
# <a id='code_validation'></a>
#
# # Step 2: Code Validation against original C code \[Back to [top](#toc)\]
# $$\label{code_validation}$$
#
# To validate the code in this tutorial we check for agreement between the files
#
# 1. that were written in this tutorial and
# 1. those that are stored in `GiRaFFE_NRPy/GiRaFFE_Ccode_library` or generated by `GiRaFFE_NRPy_A2B.py`
#
# +
# Declare gridfunctions necessary to generate the C code:
gammaDD = ixp.register_gridfunctions_for_single_rank2("AUXEVOL","gammaDD","sym01",DIM=3)
betaU = ixp.register_gridfunctions_for_single_rank1("AUXEVOL","betaU",DIM=3)
alpha = gri.register_gridfunctions("AUXEVOL","alpha",DIM=3)
BU = ixp.register_gridfunctions_for_single_rank1("AUXEVOL","BU",DIM=3)
ValenciavU = ixp.register_gridfunctions_for_single_rank1("AUXEVOL","ValenciavU",DIM=3)
StildeD = ixp.register_gridfunctions_for_single_rank1("EVOL","StildeD",DIM=3)
# Declare this symbol:
sqrt4pi = par.Cparameters("REAL",thismodule,"sqrt4pi","sqrt(4.0*M_PI)")
# First, we generate the file using the functions written in this notebook:
outCparams = "outCverbose=False"
write_out_functions_for_StildeD_source_term(outdir,outCparams,gammaDD,betaU,alpha,ValenciavU,BU,sqrt4pi)
# Define the directory that we wish to validate against:
valdir = os.path.join("GiRaFFE_NRPy","GiRaFFE_Ccode_library","RHSs")
cmd.mkdir(valdir)
import GiRaFFE_NRPy.GiRaFFE_NRPy_Source_Terms as source
source.write_out_functions_for_StildeD_source_term(valdir,outCparams,gammaDD,betaU,alpha,ValenciavU,BU,sqrt4pi)
import difflib
import sys
print("Printing difference between original C code and this code...")
# Open the files to compare
files = ["calculate_StildeD0_source_term.h","calculate_StildeD1_source_term.h","calculate_StildeD2_source_term.h"]
for file in files:
print("Checking file " + file)
with open(os.path.join(valdir,file)) as file1, open(os.path.join(outdir,file)) as file2:
# Read the lines of each file
file1_lines = file1.readlines()
file2_lines = file2.readlines()
num_diffs = 0
for line in difflib.unified_diff(file1_lines, file2_lines, fromfile=os.path.join(valdir+file), tofile=os.path.join(outdir+file)):
sys.stdout.writelines(line)
num_diffs = num_diffs + 1
if num_diffs == 0:
print("No difference. TEST PASSED!")
else:
print("ERROR: Disagreement found with .py file. See differences above.")
sys.exit(1)
# -
# <a id='latex_pdf_output'></a>
#
# # Step 3: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
# $$\label{latex_pdf_output}$$
#
# The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
# [Tutorial-GiRaFFE_NRPy_C_code_library-Source_Terms](TTutorial-GiRaFFE_NRPy_C_code_library-Source_Terms.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
cmd.output_Jupyter_notebook_to_LaTeXed_PDF("Tutorial-GiRaFFE_NRPy-Source_Terms",location_of_template_file=os.path.join(".."))
| in_progress/Tutorial-GiRaFFE_NRPy-Source_Terms.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # How does the SFF method work?
# Vanderburg and Johnson (2014) introduced a method for "Self Flat Fielding" by tracking how the lightcurve changes with motion of the spacecraft:
#
# [A Technique for Extracting Highly Precise Photometry for the Two-Wheeled Kepler Mission](http://adsabs.harvard.edu/abs/2014PASP..126..948V)
#
# In this notebook we replicate the K2SFF method following the same example source, #60021426, as that in the publication. We aim to demystify the technique, which is extremely popular within the K2 community. We have focused on reproducibility, so that we achieve the same result at the publication.
#
# The Vanderburg & Johnson 2014 paper uses data from the Kepler two-wheel "Concept Engineering Test", predating campaign 0, and sometimes called campaign *"eng"* or abbreviated CET. This vestigal "campaign" lacks some of the standardization of later K2 campaigns--- it was much shorter, only about 9 days long, it lacks some of the standard quality flags, targets have non-traditional EPIC IDs, and other quirks.
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
from astropy.io import fits
import pandas as pd
# ## Retrieve the K2SFF data for ENG test source `60021426`
# First we will retrieve data and inspect the mask used in the paper.
path = 'http://archive.stsci.edu/hlsps/k2sff/cet/060000000/21426/hlsp_k2sff_k2_lightcurve_060021426-cet_kepler_v1_llc.fits'
vdb_fits = fits.open(path)
# The `BESTAPER` keyword explains which aperture was chosen as the "best" by Vanderburg & Johnson 2014. The FITS header for that slice contains the metadata needed to reproduce the mask.
keys = ['MASKTYPE', 'MASKINDE', 'NPIXSAP']
_ = [print(key, ' : ', vdb_fits['BESTAPER'].header[key]) for key in keys]
# We want the *exact same* mask as Vanderburg & Johnson 2014, but the publication version and MAST version differ!
#
# Publication version:
# 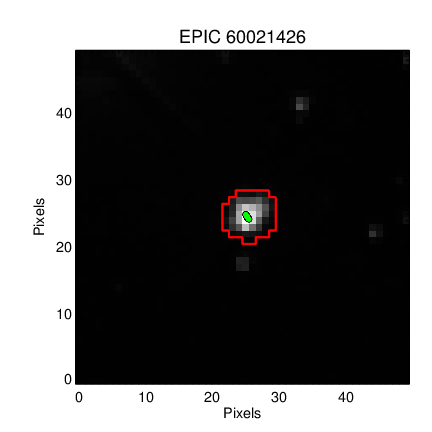
# MAST Version:
# 
# Aperture 7 should yield a bigger mask, more similar to what was used in the paper.
VDB_J_mask = vdb_fits['PRF_APER_TBL'].data[7,:, :] == True
VDB_J_mask.sum()
# Save the mask for easy use in our next notebook.
np.save('VDB_J_2014_mask.npy', VDB_J_mask)
# ## Manually reproduce with the Vanderburg-provided diagnostic data
# Retrieve the Vanderburg-provided diagnostic data for the Kepler ENG testing.
# Uncomment the line below to retrieve the data programmatically, or manually get the linked file in a browser and save it to this directory.
# +
# #! wget https://www.cfa.harvard.edu/~avanderb/k2/ep60021426alldiagnostics.csv
# -
df = pd.read_csv('ep60021426alldiagnostics.csv',index_col=False)
df.head()
# We can mean-subtract the provided $x-y$ centroids, assigning them column and row identifiers, then rotate the coordinates into their major and minor axes.
col = df[' X-centroid'].values
col = col - np.mean(col)
row = df[' Y-centroid'].values
row = row - np.mean(row)
def _get_eigen_vectors(centroid_col, centroid_row):
'''get the eigenvalues and eigenvectors given centroid x, y positions'''
centroids = np.array([centroid_col, centroid_row])
eig_val, eig_vec = np.linalg.eigh(np.cov(centroids))
return eig_val, eig_vec
def _rotate(eig_vec, centroid_col, centroid_row):
'''rotate the centroids into their predominant linear axis'''
centroids = np.array([centroid_col, centroid_row])
return np.dot(eig_vec, centroids)
eig_val, eig_vec = _get_eigen_vectors(col, row)
v1, v2 = eig_vec
# The major axis is the latter.
platescale = 4.0 # The Kepler plate scale; has units of arcseconds / pixel
plt.figure(figsize=(5, 6))
plt.plot(col * platescale, row * platescale, 'ko', ms=4)
plt.plot(col * platescale, row * platescale, 'ro', ms=1)
plt.xticks([-2, -1,0, 1, 2])
plt.yticks([-2, -1,0, 1, 2])
plt.xlabel('X position [arcseconds]')
plt.ylabel('Y position [arcseconds]')
plt.xlim(-2, 2)
plt.ylim(-2, 2)
plt.plot([0, v1[0]], [0, v1[1]], color='blue', lw=3)
plt.plot([0, v2[0]], [0, v2[1]], color='blue', lw=3);
# Following the form of **Figure 2** of Vanderburg & Johsnon 2014.
rot_colp, rot_rowp = _rotate(eig_vec, col, row) #units in pixels
# You can rotate into the new reference frame.
plt.figure(figsize=(5, 6))
plt.plot(rot_rowp * platescale, rot_colp * platescale, 'ko', ms=4)
plt.plot(rot_rowp * platescale, rot_colp * platescale, 'ro', ms=1)
plt.xticks([-2, -1,0, 1, 2])
plt.yticks([-2, -1,0, 1, 2])
plt.xlabel("X' position [arcseconds]")
plt.ylabel("Y' position [arcseconds]")
plt.xlim(-2, 2)
plt.ylim(-2, 2)
plt.plot([0, 1], [0, 0], color='blue')
plt.plot([0, 0], [0, 1], color='blue');
# We need to calculate the arclength using:
# \begin{equation}s= \int_{x'_0}^{x'_1}\sqrt{1+\left( \frac{dy'_p}{dx'}\right)^2} dx'\end{equation}
#
# where $x^\prime_0$ is the transformed $x$ coordinate of the point with the smallest $x^\prime$ position, and $y^\prime_p$ is the best--fit polynomial function.
# Fit a $5^{th}$ order polynomial to the rotated coordinates.
z = np.polyfit(rot_rowp, rot_colp, 5)
p5 = np.poly1d(z)
p5_deriv = p5.deriv()
x0_prime = np.min(rot_rowp)
xmax_prime = np.max(rot_rowp)
x_dense = np.linspace(x0_prime, xmax_prime, 2000)
plt.plot(rot_rowp, rot_colp, '.')
plt.plot(x_dense, p5(x_dense))
plt.ylabel('Position along minor axis (pixels)')
plt.xlabel('Position along major axis (pixels)')
plt.title('Performance of polynomial regression')
plt.ylim(-0.1, 0.1);
# We see evidence for a [bias-variance tradeoff](https://en.wikipedia.org/wiki/Bias%E2%80%93variance_tradeoff), suggesting some modest opportunity for improvement.
@np.vectorize
def arclength(x):
'''Input x1_prime, get out arclength'''
gi = x_dense <x
s_integrand = np.sqrt(1 + p5_deriv(x_dense[gi]) ** 2)
s = np.trapz(s_integrand, x=x_dense[gi])
return s
# Let's double check that we compute the same arclength as the published paper.
aspect_ratio = plt.figaspect(1)
plt.figure(figsize=aspect_ratio)
plt.plot(df[' arclength'], arclength(rot_rowp)*4.0, '.')
plt.xlabel('$s$ (Vanderburg & Johnson 2014)')
plt.ylabel('$s$ (This work)')
plt.plot([0, 4], [0, 4], 'k--');
# Yes, we compute arclength correctly.
# Now we apply a **high-pass filter** to the raw lightcurve data. We follow the original paper by using *BSplines* with 1.5 day breakpoints. You can also apply data exclusion at this stage.
from scipy.interpolate import BSpline
from scipy import interpolate
times, raw_fluxes = df['BJD - 2454833'].values, df[' Raw Flux'].values
# We find the weighted least square spline for a given set of knots, $t$. We supply interior knots as knots on the ends are added automatically, as stated in the `interpolate.splrep()` docstring.
interior_knots = np.arange(times[0]+1.5, times[0]+6, 1.5)
t,c,k = interpolate.splrep(times, raw_fluxes, s=0, task=-1, t=interior_knots)
bspl = BSpline(t,c,k)
plt.plot(times, raw_fluxes, '.')
plt.plot(times, bspl(times))
plt.xlabel('$t$ (days)')
plt.ylabel('Raw Flux');
# The Spline fit looks good, so we can normalize the flux by the long-term trend.
# Plot the normalized flux versus arclength to see the position-dependent flux.
fluxes = raw_fluxes/bspl(times)
# Mask the data by keeping only the good samples.
bi = df[' Thrusters On'].values == 1.0
gi = df[' Thrusters On'].values == 0.0
clean_fluxes = fluxes[gi]
al = arclength(rot_rowp[gi]) * platescale
sorted_inds = np.argsort(al)
# We will follow the paper by interpolating **flux versus arclength position** in 15 bins of means, which is a *piecewise linear fit*.
knots = np.array([np.min(al)]+
[np.median(splt) for splt in np.array_split(al[sorted_inds], 15)]+
[np.max(al)])
bin_means = np.array([clean_fluxes[sorted_inds][0]]+
[np.mean(splt) for splt in np.array_split(clean_fluxes[sorted_inds], 15)]+
[clean_fluxes[sorted_inds][-1]])
zz = np.polyfit(al, clean_fluxes,6)
sff = np.poly1d(zz)
al_dense = np.linspace(0, 4, 1000)
interp_func = interpolate.interp1d(knots, bin_means)
# +
plt.figure(figsize=(5, 6))
plt.plot(arclength(rot_rowp)*4.0, fluxes, 'ko', ms=4)
plt.plot(arclength(rot_rowp)*4.0, fluxes, 'o', color='#3498db', ms=3)
plt.plot(arclength(rot_rowp[bi])*4.0, fluxes[bi], 'o', color='r', ms=3)
plt.plot(np.sort(al), interp_func(np.sort(al)), '-', color='#e67e22')
plt.xticks([0, 1,2, 3, 4])
plt.minorticks_on()
plt.xlabel('Arclength [arcseconds]')
plt.ylabel('Relative Brightness')
plt.title('EPIC 60021426, Kp =10.3')
plt.xlim(0,4)
plt.ylim(0.997, 1.002);
# -
# Following **Figure 4** of Vanderburg & Johnson 2014.
# Apply the Self Flat Field (SFF) correction:
corr_fluxes = clean_fluxes / interp_func(al)
# +
plt.figure(figsize=(10,6))
dy = 0.004
plt.plot(df['BJD - 2454833'], df[' Raw Flux']+dy, 'ko', ms=4)
plt.plot(df['BJD - 2454833'], df[' Raw Flux']+dy, 'o', color='#3498db', ms=3)
plt.plot(df['BJD - 2454833'][bi], df[' Raw Flux'][bi]+dy, 'o', color='r', ms=3)
plt.plot(df['BJD - 2454833'][gi], corr_fluxes*bspl(times[gi]), 'o', color='k', ms = 4)
plt.plot(df['BJD - 2454833'][gi], corr_fluxes*bspl(times[gi]), 'o', color='#e67e22', ms = 3)
plt.xlabel('BJD - 2454833')
plt.ylabel('Relative Brightness')
plt.xlim(1862, 1870)
plt.ylim(0.994, 1.008);
# -
# Following **Figure 5** of Vanderburg & Johnson 2015.
# *The end.*
| docs/source/tutorials/04-replicate-vanderburg-2014-k2sff.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Optimal stopping
# > Or how I learned you can loose more than 100% value on options
#
# - toc: true
# - badges: true
# - comments: true
# - categories: [jupyter]
# - image: images/SPY_raw_bfly_prob.png
# Discrete time case
# Continuous time case
| _notebooks/2022-05-01-Optimal Stopping American Options.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Do you want to speed up the fitting of your machine learning algorithm? Scikit-learn offers quite a few ways to do this. One way is to train your model in parallel using the n_jobs parameter which exists for many scikit-learn models. A really simple way is to reduce the number of rows or columns in your data. The problem with this approach is that its hard to know which rows and especially which columns to remove. Principal component analysis, commonly known as PCA, is a technique that you can use to smartly reduce the dimensionality of your dataset while losing the least amount of information possible. In this video, I'll share with you the process of how you can use principal component analysis to speed up the fitting of a logistic regression model.
# ## Import Libraries
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.linear_model import LogisticRegression
# -
# ## Load the Dataset
# The dataset is a modified version of the MNIST dataset that contains 2000 labeled images of each digit 0 and 1. The images are 28 pixels by 28 pixels.
#
# Parameters | Number
# --- | ---
# Classes | 2 (digits 0 and 1)
# Samples per class | 2000 samples per class
# Samples total | 4000
# Dimensionality | 784 (28 x 28 images)
# Features | integers values from 0 to 255
#
# For convenience, I have arranged the data into csv file.
df = pd.read_csv('data/MNISTonly0_1.csv')
df.head()
# ## Visualize Each Digit
pixel_colnames = df.columns[:-1]
# Get all columns except the label column for the first image
image_values = df.loc[0, pixel_colnames].values
plt.figure(figsize=(8,4))
for index in range(0, 2):
plt.subplot(1, 2, 1 + index )
image_values = df.loc[index, pixel_colnames].values
image_label = df.loc[index, 'label']
plt.imshow(image_values.reshape(28,28), cmap ='gray')
plt.title('Label: ' + str(image_label), fontsize = 18)
# ## Splitting Data into Training and Test Sets
X_train, X_test, y_train, y_test = train_test_split(df[pixel_colnames], df['label'], random_state=0)
# ## Standardize the Data
# PCA and logisitic regression are sensitive to the scale of your features. You can standardize your data onto unit scale (mean = 0 and variance = 1) by using Scikit-Learn's `StandardScaler`.
# +
scaler = StandardScaler()
# Fit on training set only.
scaler.fit(X_train)
# Apply transform to both the training set and the test set.
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
# -
# Variable created for demonstational purposes in the notebook
scaledTrainImages = X_train.copy()
# ## PCA then Logistic Regression
# +
"""
n_components = .90 means that scikit-learn will choose the minimum number
of principal components such that 90% of the variance is retained.
"""
pca = PCA(n_components = .90)
# Fit PCA on training set only
pca.fit(X_train)
# Apply the mapping (transform) to both the training set and the test set.
X_train = pca.transform(X_train)
X_test = pca.transform(X_test)
# Logistic Regression
clf = LogisticRegression()
clf.fit(X_train, y_train)
print('Number of dimensions before PCA: ' + str(len(pixel_colnames)))
print('Number of dimensions after PCA: ' + str(pca.n_components_))
print('Classification accuracy: ' + str(clf.score(X_test, y_test)))
# -
# ## Relationship between Cumulative Explained Variance and Number of Principal Components
#
# Don't worry if you don't understand the code in this section. It is to show the level of redundancy present in multiple dimensions.
# +
# if n_components is not set, all components are kept (784 in this case)
pca = PCA()
pca.fit(scaledTrainImages)
# Summing explained variance
tot = sum(pca.explained_variance_)
var_exp = [(i/tot)*100 for i in sorted(pca.explained_variance_, reverse=True)]
# Cumulative explained variance
cum_var_exp = np.cumsum(var_exp)
# PLOT OUT THE EXPLAINED VARIANCES SUPERIMPOSED
fig, ax = plt.subplots(nrows = 1, ncols = 1, figsize = (10,7));
ax.tick_params(labelsize = 18)
ax.plot(range(1, 785), cum_var_exp, label='cumulative explained variance')
ax.set_ylabel('Cumulative Explained variance', fontsize = 16)
ax.set_xlabel('Principal components', fontsize = 16)
ax.axhline(y = 95, color='k', linestyle='--', label = '95% Explained Variance')
ax.axhline(y = 90, color='c', linestyle='--', label = '90% Explained Variance')
ax.axhline(y = 85, color='r', linestyle='--', label = '85% Explained Variance')
ax.legend(loc='best', markerscale = 1.0, fontsize = 12)
# -
# So that's it, PCA can be used to speed up the fitting of your algorithm.
| scikitlearn/Ex_Files_ML_SciKit_Learn/Exercise Files/03_04_PCA_to_Speed_Up_Machine_Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Identifying People in CDCS Metadata Descriptions
# Run named entity recognition (NER) on the metadata descriptions extracted from the CDCS' online archival catalog.
#
# The code in this Jupyter Notebook is part of a PhD project to create a gold standard dataset labeled for gender biased language, on which a classifier can be trained to identify gender bias in archival metadata descriptions.
#
# This project is focused on the English language and archival institutions in the United Kingdom.
#
# * Author: <NAME>
# * Date: November 2020 - February 2021
# * Project: PhD Case Study 1
# * Data Provider: [ArchivesSpace](https://archives.collections.ed.ac.uk/), Centre for Research Collections, University of Edinburgh
#
# ***
#
# **Table of Contents**
#
# [I. Corpus Statistics](#corpus-stats)
#
# [II. Named Entity Recognition with SpaCy](#ner)
#
# [III. Checking for Duplicate Descritions](#check-dups)
#
# ***
# +
import pandas as pd
import nltk
from nltk.tokenize import word_tokenize
from nltk.tokenize import sent_tokenize
# nltk.download('punkt')
from nltk.corpus import PlaintextCorpusReader
# nltk.download('averaged_perceptron_tagger')
from nltk.corpus import stopwords
# nltk.download('stopwords')
from nltk.tag import pos_tag
import string
import csv
import re
import spacy
from spacy import displacy
from collections import Counter
try:
import en_core_web_sm
except ImportError:
print("Downlading en_core_web_sm model")
import sys
# !{sys.executable} -m spacy download en_core_web_sm
else:
print("Already have en_core_web_sm")
import en_core_web_sm
nlp = en_core_web_sm.load()
# -
# <a id="corpus-stats"></a>
# ## I. Corpus Statisctics
descs = PlaintextCorpusReader("../AnnotationData/descriptions_by_fonds_split_with_ann/descriptions_by_fonds_split_with_ann/", ".+\.txt")
tokens = descs.words()
print(tokens[0:20])
sentences = descs.sents()
print(sentences[0:5])
# +
def corpusStatistics(plaintext_corpus_read_lists):
total_chars = 0
total_tokens = 0
total_sents = 0
total_files = 0
# fileids are the TXT file names in the nls-text-ladiesDebating folder:
for fileid in plaintext_corpus_read_lists.fileids():
total_chars += len(plaintext_corpus_read_lists.raw(fileid))
total_tokens += len(plaintext_corpus_read_lists.words(fileid))
total_sents += len(plaintext_corpus_read_lists.sents(fileid))
total_files += 1
print("Total Estimated...")
print(" Characters:", total_chars)
print(" Tokens:", total_tokens)
print(" Sentences:", total_sents)
print(" Files:", total_files)
corpusStatistics(descs)
# -
words = [t for t in tokens if t.isalpha()]
print("Total Estimated Words:",len(words))
to_exclude = set((stopwords.words("english")) + ["Title", "Identifier", "Scope",
"Contents", "Biograhical", "Historical", "Processing", "Information"]
)
words = [w for w in words if w not in to_exclude]
print("Total Words Excluding Metadata Field Names:",len(words))
# <a id="ner"></a>
# ## II. Name Entity Recognition with spaCy
# Run named entity recognition (NER) to estimate the names in the dataset and get a sense for the value in manually labeling names during the annotation process.
fileids = descs.fileids()
sentences = []
for fileid in fileids:
file = descs.raw(fileid)
sentences += nltk.sent_tokenize(file)
person_list = []
for s in sentences:
s_ne = nlp(s)
for entity in s_ne.ents:
if entity.label_ == 'PERSON':
person_list += [entity.text]
unique_persons = list(set(person_list))
print(len(unique_persons))
# + jupyter={"outputs_hidden": true}
unique_persons
# -
# Not perfect...some non-person entities labeled such as `Librarian` and `Diploma`. I'll add labeling person names to the annotation instructions!
# <a id="check-dups"></a>
# ## III. Checking for Duplicate Descriptions
# +
# descs = PlaintextCorpusReader("../AnnotationData/descriptions_by_fonds_split_with_ann/descriptions_by_fonds_split_with_ann/", ".+\.txt")
# fileids = descs.fileids()
# descs.raw(fileids[0])
# -
x = descs.raw(fileids[0])
x.split("\n")
descs_split = []
for fileid in fileids:
d = descs.raw(fileid).split("\n")
for section in d:
if len(section) > 0:
if "Identifier" not in section and "Title" not in section and "Scope and Contents" not in section and "Biographical / Historical" not in section and "Processing Information" not in section and "No information provided" not in section:
descs_split += [section]
print(descs_split[0:4])
len(descs_split)
len(set(descs_split))
print(len(descs_split)-len(set(descs_split)))
# It looks like there are 11,378 descriptions that are repeated in the corpus.
| Analysis_Preannotation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NER with bertreach
#
# > "Finetuning bertreach for NER"
#
# - toc: false
# - branch: master
# - hidden: true
# - badges: true
# - categories: [irish, ner, bert, bertreach]
# + [markdown] id="UldPY2vaxCOJ"
# This is a lightly edited version of [this notebook](https://github.com/huggingface/notebooks/blob/master/examples/token_classification.ipynb).
# + [markdown] id="X4cRE8IbIrIV"
# If you're opening this Notebook on colab, you will probably need to install 🤗 Transformers and 🤗 Datasets. Uncomment the following cell and run it.
# + id="MOsHUjgdIrIW"
# %%capture
# !pip install datasets transformers seqeval
# + [markdown] id="1IU9pa_DPSOk"
# If you're opening this notebook locally, make sure your environment has an install from the last version of those libraries.
#
# To be able to share your model with the community and generate results like the one shown in the picture below via the inference API, there are a few more steps to follow.
#
# First you have to store your authentication token from the Hugging Face website (sign up [here](https://huggingface.co/join) if you haven't already!) then execute the following cell and input your username and password:
# + [markdown] id="ARRVh3-ixvs9"
# (Huggingface notebooks skip this bit, but you need to set credential.helper before anything else works).
# + id="9RX-uPZLxurJ"
# !git config --global credential.helper store
# + colab={"base_uri": "https://localhost:8080/", "height": 283, "referenced_widgets": ["445ac7ed80f7436eaa1f81e8f71cb1e5", "<KEY>", "f7a501140914451d952b3f1528c907d7", "11eedcf6113342f8a5795f40686cb7cd", "efb271ceea364dde8e54a5397615c267", "ed03af8d393543be9abedf5f6ac070d5", "<KEY>", "<KEY>", "<KEY>", "5e16bebe96ad4b6fb4b141675fe6be4c", "<KEY>", "9594610636864c05a147ad11dfee7063", "418a605ac1d54f8f852c76504a941934", "d981126fe0a649b5aba5d4a7f2c3bbdf", "abc72496d5eb48a787ddd8f4f1ab21e8", "2cccaab9e726491592efc43d2296698c"]} id="npzw8gOYPSOl" outputId="f9c0bfaa-2a9e-4f0f-9746-bb2002a619b2"
from huggingface_hub import notebook_login
notebook_login()
# + [markdown] id="5-CthmJKPSOm"
# Then you need to install Git-LFS. Uncomment the following instructions:
# + colab={"base_uri": "https://localhost:8080/"} id="7YAk6M5KPSOn" outputId="154c5842-906c-4f27-e26b-aa7ad595cd4b"
# !apt install git-lfs
# + [markdown] id="CHYxLRR8PSOo"
# Make sure your version of Transformers is at least 4.11.0 since the functionality was introduced in that version:
# + colab={"base_uri": "https://localhost:8080/"} id="U7ZVbepsPSOp" outputId="8036e6e0-eb72-4b89-ca57-e3d60c0b7f0f"
import transformers
print(transformers.__version__)
# + [markdown] id="HFASsisvIrIb"
# You can find a script version of this notebook to fine-tune your model in a distributed fashion using multiple GPUs or TPUs [here](https://github.com/huggingface/transformers/tree/master/examples/token-classification).
# + [markdown] id="rEJBSTyZIrIb"
# # Fine-tuning a model on a token classification task
# + [markdown] id="w6vfS60cPSOt"
# In this notebook, we will see how to fine-tune one of the [🤗 Transformers](https://github.com/huggingface/transformers) model to a token classification task, which is the task of predicting a label for each token.
#
# 
#
# The most common token classification tasks are:
#
# - NER (Named-entity recognition) Classify the entities in the text (person, organization, location...).
# - POS (Part-of-speech tagging) Grammatically classify the tokens (noun, verb, adjective...)
# - Chunk (Chunking) Grammatically classify the tokens and group them into "chunks" that go together
#
# We will see how to easily load a dataset for these kinds of tasks and use the `Trainer` API to fine-tune a model on it.
# + [markdown] id="4RRkXuteIrIh"
# This notebook is built to run on any token classification task, with any model checkpoint from the [Model Hub](https://huggingface.co/models) as long as that model has a version with a token classification head and a fast tokenizer (check on [this table](https://huggingface.co/transformers/index.html#bigtable) if this is the case). It might just need some small adjustments if you decide to use a different dataset than the one used here. Depending on you model and the GPU you are using, you might need to adjust the batch size to avoid out-of-memory errors. Set those three parameters, then the rest of the notebook should run smoothly:
# + id="zVvslsfMIrIh"
task = "ner" # Should be one of "ner", "pos" or "chunk"
model_checkpoint = "jimregan/BERTreach"
batch_size = 16
# + [markdown] id="whPRbBNbIrIl"
# ## Loading the dataset
# + [markdown] id="W7QYTpxXIrIl"
# We will use the [🤗 Datasets](https://github.com/huggingface/datasets) library to download the data and get the metric we need to use for evaluation (to compare our model to the benchmark). This can be easily done with the functions `load_dataset` and `load_metric`.
# + id="IreSlFmlIrIm"
from datasets import load_dataset, load_metric
# + [markdown] id="CKx2zKs5IrIq"
# For our example here, we'll use the [CONLL 2003 dataset](https://www.aclweb.org/anthology/W03-0419.pdf). The notebook should work with any token classification dataset provided by the 🤗 Datasets library. If you're using your own dataset defined from a JSON or csv file (see the [Datasets documentation](https://huggingface.co/docs/datasets/loading_datasets.html#from-local-files) on how to load them), it might need some adjustments in the names of the columns used.
# + colab={"base_uri": "https://localhost:8080/", "height": 67, "referenced_widgets": ["50c7c01d13da42d6b264ea36684e5f1f", "<KEY>", "<KEY>", "d6c350910a3a48d485cfeab446eb60ff", "6c1e4ea54ae844a3a2d32a8b4a71e962", "e41938f14b3946cf9874de60a1cf5fa5", "174e47a8e6da4edca874390cab52707e", "f54f7775ee234038a4071ab8f1841186", "<KEY>", "61451ee6101842a98d05aca771754332", "d5c6208c774d4440894efb480abc1664"]} id="s_AY1ATSIrIq" outputId="f5442b3e-5c6a-409f-ce71-a5b46b894803"
datasets = load_dataset("wikiann", "ga")
# + [markdown] id="RzfPtOMoIrIu"
# The `datasets` object itself is [`DatasetDict`](https://huggingface.co/docs/datasets/package_reference/main_classes.html#datasetdict), which contains one key for the training, validation and test set.
# + colab={"base_uri": "https://localhost:8080/"} id="GWiVUF0jIrIv" outputId="4cda094b-9ed3-4a00-fc41-f5a163b3bff2"
datasets
# + [markdown] id="X_XUcknPPSO1"
# We can see the training, validation and test sets all have a column for the tokens (the input texts split into words) and one column of labels for each kind of task we introduced before.
# + [markdown] id="u3EtYfeHIrIz"
# To access an actual element, you need to select a split first, then give an index:
# + colab={"base_uri": "https://localhost:8080/"} id="X6HrpprwIrIz" outputId="24e54194-65b2-433a-dba2-7bf1ac7eb5bb"
datasets["train"][0]
# + [markdown] id="lppNSJaKPSO3"
# The labels are already coded as integer ids to be easily usable by our model, but the correspondence with the actual categories is stored in the `features` of the dataset:
# + colab={"base_uri": "https://localhost:8080/"} id="vetcKtTJPSO3" outputId="d06b4662-e8c2-4c6f-ea22-67cbc5c9a7a5"
datasets["train"].features[f"ner_tags"]
# + [markdown] id="URaPOdjRPSO5"
# So for the NER tags, 0 corresponds to 'O', 1 to 'B-PER' etc... On top of the 'O' (which means no special entity), there are four labels for NER here, each prefixed with 'B-' (for beginning) or 'I-' (for intermediate), that indicate if the token is the first one for the current group with the label or not:
# - 'PER' for person
# - 'ORG' for organization
# - 'LOC' for location
# - 'MISC' for miscellaneous
# + [markdown] id="dCCD_uaQPSO5"
# Since the labels are lists of `ClassLabel`, the actual names of the labels are nested in the `feature` attribute of the object above:
# + colab={"base_uri": "https://localhost:8080/"} id="Q9jfLo3rPSO6" outputId="a258d350-8279-498b-d94a-da31853b5dfb"
label_list = datasets["train"].features[f"{task}_tags"].feature.names
label_list
# + [markdown] id="WHUmphG3IrI3"
# To get a sense of what the data looks like, the following function will show some examples picked randomly in the dataset (automatically decoding the labels in passing).
# + id="i3j8APAoIrI3"
from datasets import ClassLabel, Sequence
import random
import pandas as pd
from IPython.display import display, HTML
def show_random_elements(dataset, num_examples=10):
assert num_examples <= len(dataset), "Can't pick more elements than there are in the dataset."
picks = []
for _ in range(num_examples):
pick = random.randint(0, len(dataset)-1)
while pick in picks:
pick = random.randint(0, len(dataset)-1)
picks.append(pick)
df = pd.DataFrame(dataset[picks])
for column, typ in dataset.features.items():
if isinstance(typ, ClassLabel):
df[column] = df[column].transform(lambda i: typ.names[i])
elif isinstance(typ, Sequence) and isinstance(typ.feature, ClassLabel):
df[column] = df[column].transform(lambda x: [typ.feature.names[i] for i in x])
display(HTML(df.to_html()))
# + colab={"base_uri": "https://localhost:8080/", "height": 432} id="SZy5tRB_IrI7" outputId="35f50cf0-4606-48be-97cc-54643321ccba"
show_random_elements(datasets["train"])
# + [markdown] id="n9qywopnIrJH"
# ## Preprocessing the data
# + [markdown] id="YVx71GdAIrJH"
# Before we can feed those texts to our model, we need to preprocess them. This is done by a 🤗 Transformers `Tokenizer` which will (as the name indicates) tokenize the inputs (including converting the tokens to their corresponding IDs in the pretrained vocabulary) and put it in a format the model expects, as well as generate the other inputs that model requires.
#
# To do all of this, we instantiate our tokenizer with the `AutoTokenizer.from_pretrained` method, which will ensure:
#
# - we get a tokenizer that corresponds to the model architecture we want to use,
# - we download the vocabulary used when pretraining this specific checkpoint.
#
# That vocabulary will be cached, so it's not downloaded again the next time we run the cell.
# + colab={"base_uri": "https://localhost:8080/"} id="eXNLu_-nIrJI" outputId="5981ce18-02da-4278-c75d-9939729142aa"
from transformers import RobertaTokenizerFast
tokenizer = RobertaTokenizerFast.from_pretrained(model_checkpoint, add_prefix_space=True)
# + [markdown] id="Vl6IidfdIrJK"
# The following assertion ensures that our tokenizer is a fast tokenizers (backed by Rust) from the 🤗 Tokenizers library. Those fast tokenizers are available for almost all models, and we will need some of the special features they have for our preprocessing.
# + id="pZaRFHN7PSO-"
import transformers
assert isinstance(tokenizer, transformers.PreTrainedTokenizerFast)
# + [markdown] id="cfiX-_BgPSO_"
# You can check which type of models have a fast tokenizer available and which don't on the [big table of models](https://huggingface.co/transformers/index.html#bigtable).
# + [markdown] id="rowT4iCLIrJK"
# You can directly call this tokenizer on one sentence:
# + colab={"base_uri": "https://localhost:8080/"} id="a5hBlsrHIrJL" outputId="2bd9771e-e59b-460c-a325-45333c61cee2"
tokenizer("Is abairt amháin é seo!")
# + [markdown] id="B9K1KIg4PSPA"
# Depending on the model you selected, you will see different keys in the dictionary returned by the cell above. They don't matter much for what we're doing here (just know they are required by the model we will instantiate later), you can learn more about them in [this tutorial](https://huggingface.co/transformers/preprocessing.html) if you're interested.
#
# If, as is the case here, your inputs have already been split into words, you should pass the list of words to your tokenzier with the argument `is_split_into_words=True`:
# + colab={"base_uri": "https://localhost:8080/"} id="qdxB67GzPSPA" outputId="3bedb26b-6e89-4885-a85e-8f3416478539"
tokenizer(["Hello", ",", "this", "is", "one", "sentence", "split", "into", "words", "."], is_split_into_words=True)
# + [markdown] id="XjXRHpJLPSPB"
# Note that transformers are often pretrained with subword tokenizers, meaning that even if your inputs have been split into words already, each of those words could be split again by the tokenizer. Let's look at an example of that:
# + colab={"base_uri": "https://localhost:8080/"} id="mi8IJrMFPSPB" outputId="cad513ef-3f29-4e5b-c134-4e290d067ef1"
example = datasets["train"][4]
print(example["tokens"])
# + colab={"base_uri": "https://localhost:8080/"} id="NKP2znNrPSPC" outputId="93a5dee3-b047-4231-974a-bf569d1cc0ea"
tokenized_input = tokenizer(example["tokens"], is_split_into_words=True)
tokens = tokenizer.convert_ids_to_tokens(tokenized_input["input_ids"])
print(tokens)
# + [markdown] id="aL91xiUePSPC"
# Here the words "Zwingmann" and "sheepmeat" have been split in three subtokens.
#
# This means that we need to do some processing on our labels as the input ids returned by the tokenizer are longer than the lists of labels our dataset contain, first because some special tokens might be added (we can a `[CLS]` and a `[SEP]` above) and then because of those possible splits of words in multiple tokens:
# + colab={"base_uri": "https://localhost:8080/"} id="3intBYzJPSPD" outputId="b5296afe-243a-4dc6-f0f2-14c4c81814fd"
len(example[f"{task}_tags"]), len(tokenized_input["input_ids"])
# + [markdown] id="YFEDGF_WPSPD"
# Thankfully, the tokenizer returns outputs that have a `word_ids` method which can help us.
# + colab={"base_uri": "https://localhost:8080/"} id="ZjPu1wxoPSPD" outputId="96a09327-cc87-478c-ee60-356f38207b03"
print(tokenized_input.word_ids())
# + [markdown] id="WqHQp3otPSPE"
# As we can see, it returns a list with the same number of elements as our processed input ids, mapping special tokens to `None` and all other tokens to their respective word. This way, we can align the labels with the processed input ids.
# + colab={"base_uri": "https://localhost:8080/"} id="S2bLHIshPSPE" outputId="b5aa29df-fb7b-4eaf-de0b-3e06b66f4847"
word_ids = tokenized_input.word_ids()
aligned_labels = [-100 if i is None else example[f"{task}_tags"][i] for i in word_ids]
print(len(aligned_labels), len(tokenized_input["input_ids"]))
# + [markdown] id="cj3LA4UXPSPE"
# Here we set the labels of all special tokens to -100 (the index that is ignored by PyTorch) and the labels of all other tokens to the label of the word they come from. Another strategy is to set the label only on the first token obtained from a given word, and give a label of -100 to the other subtokens from the same word. We propose the two strategies here, just change the value of the following flag:
# + id="lJkzdIlIPSPF"
label_all_tokens = True
# + [markdown] id="2C0hcmp9IrJQ"
# We're now ready to write the function that will preprocess our samples. We feed them to the `tokenizer` with the argument `truncation=True` (to truncate texts that are bigger than the maximum size allowed by the model) and `is_split_into_words=True` (as seen above). Then we align the labels with the token ids using the strategy we picked:
# + id="vc0BSBLIIrJQ"
def tokenize_and_align_labels(examples):
tokenized_inputs = tokenizer(examples["tokens"], truncation=True, is_split_into_words=True)
labels = []
for i, label in enumerate(examples[f"{task}_tags"]):
word_ids = tokenized_inputs.word_ids(batch_index=i)
previous_word_idx = None
label_ids = []
for word_idx in word_ids:
# Special tokens have a word id that is None. We set the label to -100 so they are automatically
# ignored in the loss function.
if word_idx is None:
label_ids.append(-100)
# We set the label for the first token of each word.
elif word_idx != previous_word_idx:
label_ids.append(label[word_idx])
# For the other tokens in a word, we set the label to either the current label or -100, depending on
# the label_all_tokens flag.
else:
label_ids.append(label[word_idx] if label_all_tokens else -100)
previous_word_idx = word_idx
labels.append(label_ids)
tokenized_inputs["labels"] = labels
return tokenized_inputs
# + [markdown] id="0lm8ozrJIrJR"
# This function works with one or several examples. In the case of several examples, the tokenizer will return a list of lists for each key:
# + colab={"base_uri": "https://localhost:8080/"} id="-b70jh26IrJS" outputId="4ffb9712-e409-43d1-991f-239bbfa46c51"
tokenize_and_align_labels(datasets['train'][:5])
# + [markdown] id="zS-6iXTkIrJT"
# To apply this function on all the sentences (or pairs of sentences) in our dataset, we just use the `map` method of our `dataset` object we created earlier. This will apply the function on all the elements of all the splits in `dataset`, so our training, validation and testing data will be preprocessed in one single command.
# + colab={"base_uri": "https://localhost:8080/", "height": 113, "referenced_widgets": ["ee50223163a0429b83391b426dbf4191", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "af435b9755a942caba61094ec781870a", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "265a7a3116f4435abadf5b8d5e8ab594", "8f947072da59454c88e18196dc9976d0", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "fa5a85c7a80642d79abf1fe8f8d8dca4", "c406925717dd42e480e0f6b3957e390f", "df92ae2e9102414aa589c3a1040aed9a", "<KEY>", "<KEY>", "6ea5e881272241569d79091f8478ab95", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "3bda313521644f9284681fc804a8848f"]} id="DDtsaJeVIrJT" outputId="abf1d75d-e60a-4e55-ced6-67c554a7d165"
tokenized_datasets = datasets.map(tokenize_and_align_labels, batched=True)
# + [markdown] id="voWiw8C7IrJV"
# Even better, the results are automatically cached by the 🤗 Datasets library to avoid spending time on this step the next time you run your notebook. The 🤗 Datasets library is normally smart enough to detect when the function you pass to map has changed (and thus requires to not use the cache data). For instance, it will properly detect if you change the task in the first cell and rerun the notebook. 🤗 Datasets warns you when it uses cached files, you can pass `load_from_cache_file=False` in the call to `map` to not use the cached files and force the preprocessing to be applied again.
#
# Note that we passed `batched=True` to encode the texts by batches together. This is to leverage the full benefit of the fast tokenizer we loaded earlier, which will use multi-threading to treat the texts in a batch concurrently.
# + [markdown] id="545PP3o8IrJV"
# ## Fine-tuning the model
# + [markdown] id="FBiW8UpKIrJW"
# Now that our data is ready, we can download the pretrained model and fine-tune it. Since all our tasks are about token classification, we use the `AutoModelForTokenClassification` class. Like with the tokenizer, the `from_pretrained` method will download and cache the model for us. The only thing we have to specify is the number of labels for our problem (which we can get from the features, as seen before):
# + colab={"base_uri": "https://localhost:8080/", "height": 1000, "referenced_widgets": ["9cd45896f45a432b8a213df2d36fee1a", "dd73302517d1485f9981fc477fa66e68", "aae9a23e300748709ebd309ec8d5bee4", "5a4a7bcec3524810b8ffb6f3421e5cf6", "9f474e56030c45daa6b359022b7841ff", "2dd373f51945427fa783df4faba16950", "0bb8e705b37949a0a286f9710f643342", "ab33d3a7034d480d97987d38916a12f5", "3d1450e3dab2478586bbbd6680583472", "82189c839ca9483da689c28a69604747", "90c96a4de80c48e39322f80b4ad8a1e0"]} id="TlqNaB8jIrJW" outputId="c26cf91c-1d56-42fb-f4bc-de6f233aa1a3"
from transformers import AutoModelForTokenClassification, TrainingArguments, Trainer, AutoConfig
config = AutoConfig.from_pretrained(model_checkpoint,
id2label={i: label for i, label in enumerate(label_list)},
label2id={label: i for i, label in enumerate(label_list)})
model = AutoModelForTokenClassification.from_pretrained(model_checkpoint, config=config)
# + [markdown] id="CczA5lJlIrJX"
# The warning is telling us we are throwing away some weights (the `vocab_transform` and `vocab_layer_norm` layers) and randomly initializing some other (the `pre_classifier` and `classifier` layers). This is absolutely normal in this case, because we are removing the head used to pretrain the model on a masked language modeling objective and replacing it with a new head for which we don't have pretrained weights, so the library warns us we should fine-tune this model before using it for inference, which is exactly what we are going to do.
# + [markdown] id="_N8urzhyIrJY"
# To instantiate a `Trainer`, we will need to define three more things. The most important is the [`TrainingArguments`](https://huggingface.co/transformers/main_classes/trainer.html#transformers.TrainingArguments), which is a class that contains all the attributes to customize the training. It requires one folder name, which will be used to save the checkpoints of the model, and all other arguments are optional:
# + colab={"base_uri": "https://localhost:8080/"} id="Bliy8zgjIrJY" outputId="f5c5761b-b0d0-4121-9789-a4925e433457"
model_name = model_checkpoint.split("/")[-1]
args = TrainingArguments(
f"BERTreach-finetuned-{task}",
evaluation_strategy = "epoch",
learning_rate=2e-5,
per_device_train_batch_size=batch_size,
per_device_eval_batch_size=batch_size,
num_train_epochs=5,
weight_decay=0.01,
push_to_hub=True,
)
# + [markdown] id="km3pGVdTIrJc"
# Here we set the evaluation to be done at the end of each epoch, tweak the learning rate, use the `batch_size` defined at the top of the notebook and customize the number of epochs for training, as well as the weight decay.
#
# The last argument to setup everything so we can push the model to the [Hub](https://huggingface.co/models) regularly during training. Remove it if you didn't follow the installation steps at the top of the notebook. If you want to save your model locally in a name that is different than the name of the repository it will be pushed, or if you want to push your model under an organization and not your name space, use the `hub_model_id` argument to set the repo name (it needs to be the full name, including your namespace: for instance `"sgugger/bert-finetuned-ner"` or `"huggingface/bert-finetuned-ner"`).
# + [markdown] id="VKETFK5OPSPL"
# Then we will need a data collator that will batch our processed examples together while applying padding to make them all the same size (each pad will be padded to the length of its longest example). There is a data collator for this task in the Transformers library, that not only pads the inputs, but also the labels:
# + id="SB-5qhhdPSPL"
from transformers import DataCollatorForTokenClassification
data_collator = DataCollatorForTokenClassification(tokenizer)
# + [markdown] id="pFnZ8C9zPSPM"
# The last thing to define for our `Trainer` is how to compute the metrics from the predictions. Here we will load the [`seqeval`](https://github.com/chakki-works/seqeval) metric (which is commonly used to evaluate results on the CONLL dataset) via the Datasets library.
# + id="MehhnsFLPSPM"
metric = load_metric("seqeval")
# + [markdown] id="TXgIHJGrPSPM"
# This metric takes list of labels for the predictions and references:
# + colab={"base_uri": "https://localhost:8080/"} id="W1Pl8eDRPSPN" outputId="8a92dd56-fd90-426e-e0d9-d8adf4285e07"
labels = [label_list[i] for i in example[f"{task}_tags"]]
metric.compute(predictions=[labels], references=[labels])
# + [markdown] id="7sZOdRlRIrJd"
# So we will need to do a bit of post-processing on our predictions:
# - select the predicted index (with the maximum logit) for each token
# - convert it to its string label
# - ignore everywhere we set a label of -100
#
# The following function does all this post-processing on the result of `Trainer.evaluate` (which is a namedtuple containing predictions and labels) before applying the metric:
# + id="UmvbnJ9JIrJd"
import numpy as np
def compute_metrics(p):
predictions, labels = p
predictions = np.argmax(predictions, axis=2)
# Remove ignored index (special tokens)
true_predictions = [
[label_list[p] for (p, l) in zip(prediction, label) if l != -100]
for prediction, label in zip(predictions, labels)
]
true_labels = [
[label_list[l] for (p, l) in zip(prediction, label) if l != -100]
for prediction, label in zip(predictions, labels)
]
results = metric.compute(predictions=true_predictions, references=true_labels)
return {
"precision": results["overall_precision"],
"recall": results["overall_recall"],
"f1": results["overall_f1"],
"accuracy": results["overall_accuracy"],
}
# + [markdown] id="rXuFTAzDIrJe"
# Note that we drop the precision/recall/f1 computed for each category and only focus on the overall precision/recall/f1/accuracy.
#
# Then we just need to pass all of this along with our datasets to the `Trainer`:
# + colab={"base_uri": "https://localhost:8080/"} id="imY1oC3SIrJf" outputId="c73205ff-288a-420b-a0a2-acd27ee201bd"
trainer = Trainer(
model,
args,
train_dataset=tokenized_datasets["train"],
eval_dataset=tokenized_datasets["validation"],
data_collator=data_collator,
tokenizer=tokenizer,
compute_metrics=compute_metrics
)
# + [markdown] id="CdzABDVcIrJg"
# We can now finetune our model by just calling the `train` method:
# + colab={"base_uri": "https://localhost:8080/", "height": 842} id="6aEz13C_PSPP" outputId="b4d28ba3-379e-41b7-e237-e4b54a1009bf"
trainer.train()
# + [markdown] id="CKASz-2vIrJi"
# The `evaluate` method allows you to evaluate again on the evaluation dataset or on another dataset:
# + colab={"base_uri": "https://localhost:8080/", "height": 283} id="UOUcBkX8IrJi" outputId="75b87c23-3cfc-42dc-9b7d-8e8b6a2cef7a"
trainer.evaluate()
# + [markdown] id="1358wwuiPSPQ"
# To get the precision/recall/f1 computed for each category now that we have finished training, we can apply the same function as before on the result of the `predict` method:
# + colab={"base_uri": "https://localhost:8080/", "height": 404} id="_06GH4g0PSPQ" outputId="4a38c3ca-c1d4-45c1-f54e-899e2eb523b4"
predictions, labels, _ = trainer.predict(tokenized_datasets["validation"])
predictions = np.argmax(predictions, axis=2)
# Remove ignored index (special tokens)
true_predictions = [
[label_list[p] for (p, l) in zip(prediction, label) if l != -100]
for prediction, label in zip(predictions, labels)
]
true_labels = [
[label_list[l] for (p, l) in zip(prediction, label) if l != -100]
for prediction, label in zip(predictions, labels)
]
results = metric.compute(predictions=true_predictions, references=true_labels)
results
# + [markdown] id="rc5EPoMYPSPR"
# You can now upload the result of the training to the Hub, just execute this instruction:
# + colab={"base_uri": "https://localhost:8080/", "height": 413, "referenced_widgets": ["8ffb7f6f6b9b4609b5738ff62bc14e64", "f4465aba227f401e97c72b21497e1299", "a1f81973f254436894e767556a3d3a77", "4341e8385e0d48b9b3bd00a83074df14", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "fee9e8f7d44a4a8e8dee77b951bf119a", "d273e45038064c25ae7d196f74c06aa3", "<KEY>", "efc83217251e4fc283d30badf957d35a", "6be0fbacf9824cd0bd309e85ba542337", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "e269877f6a62482f98338466c27eb35c", "91b671a50c5c4a85925c0ec50db723ca", "de50243e93aa44f494a49e155dd2bf41", "<KEY>", "fda78f6340dc4b31a1dfa1997c89a668", "cd77ce81e2d646069a38cfeea2005e0b", "<KEY>", "30490d6ddada4768b113543e3ea4f224", "50cfd93cd58a47dfa9d57ff5f0ca9052", "<KEY>", "<KEY>", "6ed1124846a84b2fa8968c44ed6a7c55", "<KEY>", "13dee99469594071a750bd8512cb55ad", "<KEY>", "1111a433384743648f5aca0fbf49a910", "<KEY>", "3b2367739fe34503b38b73b2801e7def", "979d75386902447e842631d6bf032431", "76556862f0a5497ba6692323301cc6f8", "<KEY>", "0111982f12ca4949b4996f1a8e1ad17e", "<KEY>", "6c9e6f3130164dd3bdeecf603c46e13e", "023aa82684494291bdf18985e362a1c0", "<KEY>", "be39f5cdbabb4eb0aed79884e170e04d", "25fe968dbc1146ed81f087e2742d2e49", "a6c9e9a3635d42f08cbf43265835ab3e", "1c7879ac2a8d485db45ae840dbaf9067", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "341c8604e0074e33b2d43f539fa4fa7f", "79ad0dc00c6a41c7b42315898e246adc"]} id="BLlw2RnTPSPR" outputId="3d5d5f15-3bc1-422a-f691-b2a6420ce0dd"
trainer.push_to_hub()
# + [markdown] id="14VrOnlDPSPR"
# You can now share this model with all your friends, family, favorite pets: they can all load it with the identifier `"your-username/the-name-you-picked"` so for instance:
#
# ```python
# from transformers import AutoModelForTokenClassification
#
# model = AutoModelForTokenClassification.from_pretrained("sgugger/my-awesome-model")
# ```
# + id="FN7gcQ-MPSPS"
| _notebooks/2021-12-01-token_classification_bertreach.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Dropbox link for beans20190718.nwb: https://www.dropbox.com/s/srra33e0igaj5rm/beans20190718.nwb?dl=0
# + tags=[]
# %reload_ext autoreload
# %autoreload 2
# + tags=[]
# %env DJ_SUPPORT_FILEPATH_MANAGEMENT=TRUE
import os
data_dir = '/Users/loren/data/nwb'
# + tags=[]
from pathlib import Path
import datajoint as dj
# CONFIG FOR LOCAL DATABASE - CHANGE AS NEEDED
dj.config['database.host'] = 'localhost'
dj.config['database.user'] = 'root'
dj.config['database.password'] = '<PASSWORD>'
import nwb_datajoint as nd
data_dir = Path('/Users/loren/data/nwb') # CHANGE ME TO THE BASE DIRECTORY FOR DATA STORAGE ON YOUR SYSTEM
os.environ['NWB_DATAJOINT_BASE_DIR'] = str(data_dir)
os.environ['KACHERY_STORAGE_DIR'] = str(data_dir / 'kachery-storage')
os.environ['SPIKE_SORTING_STORAGE_DIR'] = str(data_dir / 'spikesorting')
# -
import warnings
warnings.simplefilter('ignore')
# + tags=[]
nd.insert_sessions(['beans20190718.nwb'])
# -
# #### Let's look at the core schema (schema = database table).
# First, Nwbfile:
nd.common.SampleCount()
# +
nd.common.Nwbfile()
# -
# Each NWB file defines a session which also also information about the subject, institution, etc.:
nd.common.Session()
# We can use the datajoint Diagram method to represent the relationship between Nwbfile and Session:
# Session is defined by a Nwbfile
dj.Diagram(nd.common.Nwbfile()) + dj.Diagram(nd.common.Session())
# The solid line indicates that the Nwbfile is the primary key for Session, so each Session has exactly one Nwbfile associated with it.
#
# The session also contains references to other schema, including Subject(), Institution, etc.
dj.Diagram(nd.common.Nwbfile()) + dj.Diagram(nd.common.Session()) + dj.Diagram(nd.common.Subject()) + dj.Diagram(nd.common.Institution()) + dj.Diagram(nd.common.Lab())
nd.common.Raw()
dj.Diagram(nd.common.Subject()) + dj.Diagram(nd.common.Session())+dj.Diagram(nd.common.Nwbfile())+dj.Diagram(nd.common.Raw())
dj.Diagram(nd.common.Session())+dj.Diagram(nd.common.Nwbfile())+dj.Diagram(nd.common.Raw())+dj.Diagram(nd.common.LFP())+dj.Diagram(nd.common.SpikeSortingParameters())
# + jupyter={"source_hidden": true}
a = (nd.common.IntervalList() & {'interval_list_name' : 'raw data valid times'}).fetch1('valid_times')
print(a)
# -
nd.common.VideoFile()
data = (nd.common.Raw() & {'nwb_file_name' : 'beans20190718.nwb'}).fetch_nwb()
data
# + jupyter={"source_hidden": true}
nd.common.SpikeSorting().drop()
# + jupyter={"source_hidden": true} tags=[]
nd.common.Session().delete()
nd.common.Nwbfile().drop()
# -
nd.common.Nwbfile().delete()
(nd.common.Raw() & {'nwb_file_name' : 'beans20190718.nwb'}).fetch1('interval_list_name')
nd.common.SpikeSorting().drop()
# +
s1 = (nd.common.IntervalList() & {'interval_list_name': '01_s1'}).fetch1()
s2 = (nd.common.IntervalList() & {'interval_list_name': '03_s2'}).fetch1()
r1 = (nd.common.IntervalList() & {'interval_list_name': '02_r1'}).fetch1()
r2 = (nd.common.IntervalList() & {'interval_list_name': '04_r2'}).fetch1()
# -
nd.common.IntervalList()
s1 = (nd.common.IntervalList() & {'interval_list_name' : '01_s1'}).fetch1('valid_times')
s1
t[0][1] = t[0][0] + 100
t = s1.copy()
nd.common.IntervalList.insert1({'nwb_file_name' : 'beans20190718.nwb', 'interval_list_name' : 'test', 'valid_times' : t})
nd.common.IntervalList()
a = (nd.common.IntervalList() & {'interval_list_name' : '01_s1'}).fetch1()
a
b = a.copy()
b['interval_list_name'] = 'test'
(nd.common.IntervalList & {'interval_list_name' : 'test'}).delete()
| notebooks/Populate_from_NWB_franklab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/wisenutgolead/wisenut_NER/blob/master/make_input_data_using_word_pos_entity.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="IyLeg3y589Ht" colab_type="text"
# json 데이터 가져오기
#
# + id="63ZQAVkG8osw" colab_type="code" colab={}
import json
import re
# json 파일 열기
# json 파일 경로는 알아서 수정
with open('/content/drive/My Drive/와이즈넛/RESULT/annotations/chosj/2013_reviewed_deleted_1.txt.adm.json', 'r', encoding='utf-8') as f:
json_data = json.load(f)
datas = json_data['data'] # 문장들만 가져오기
attributes = json_data['attributes']['entities']['items'] # 단어의 정보를 담고 있는 딕셔너리들을 가져온다
# attributes 순서를 전체 문장에서 나오는 순서로 바꾼다(start index 기준으로 오름차순 정렬)
attributes = sorted(attributes, key=lambda x : x['mentions'][0]['startOffset'])
# 함수
def get_start_index(i):
return attributes[i]['mentions'][0]['startOffset']
def get_end_index(i):
return attributes[i]['mentions'][0]['endOffset']
def get_entity(i):
return attributes[i]['type']
entities = [get_entity(i) for i in range(len(attributes))] # 문서에 있는 모든 entity(문서에 나오는 순서로 저장되어 있다)
entity_words = [datas[get_start_index(i):get_end_index(i)].strip().replace(' ', '') for i in range(len(attributes))] # entity인 단어
sentences = re.split('\r\n|\u3000', datas) # 정규표현식을 이용해 문서를 문장으로 나눈다
sentences = [x for x in sentences if len(x) != 0] # 길이가 0인 문장 제거
# + [markdown] id="wng29LVG9E2y" colab_type="text"
# mecab 설치</br></br>
# 코랩(리눅스)에서만 설치 가능. 윈도우 환경에서 설치하려면 [여기](https://cleancode-ws.tistory.com/97) 참고
# + id="IfjEyMob9ETZ" colab_type="code" colab={}
# !pip install konlpy
# + id="zZgH15Ns9M7o" colab_type="code" colab={}
# !bash <(curl -s https://raw.githubusercontent.com/konlpy/konlpy/master/scripts/mecab.sh)
# + [markdown] id="iCflrwrd9PBF" colab_type="text"
# mecab 으로 문장을 형태소로 분해
# + id="75fXjPU39O1h" colab_type="code" colab={}
from konlpy.tag import Mecab
mecab = Mecab()
mecab_sentences = [] # mecab으로 형태소 분석 된 문장을 저장
def get_pos(sentences):
for sentence in sentences:
mecab_sentences.append(mecab.pos(sentence))
get_pos(sentences)
# + [markdown] id="PkR2YrTg-WlJ" colab_type="text"
# 형태소로 분해된 단어를 기반으로 entity 태그 붙혀서 input data 만들기
# + id="JIjic59x9R9J" colab_type="code" colab={}
word_pos_entity_all = []
idx = 0 # entity로 취급되는 단어 리스트에서 단어를 가져오기 위해 사용하는 인덱스
for mecab_sentence in mecab_sentences: # 전체 문장
word_pos_entity = []
word_len = 0 # 형태소 분해된 단어의 길이
entity_len = 0 # entity 단어의 길이
for word, pos in mecab_sentence: # 문장
if word in entity_words[idx]: # 형태소가 entity 단어에 포함되면
# 형태소와 entity 단어의 길이를 잰다
word_len += len(word)
entity_len = len(entity_words[idx])
word_pos_entity.append([word, pos, 'I-'+entities[idx]])
# 형태소의 길이와 entity 단어의 길이가 같아지면 인덱스가 다음 entity 단어를 가리키게 +1을 해준다
if word_len == entity_len:
word_len = 0
entity_len = 0
idx += 1
# 형태소가 entity가 아닌 경우
else:
word_pos_entity.append([word, pos, 'O'])
word_pos_entity_all.append(word_pos_entity)
# + [markdown] id="ehTDU9XJ-gu7" colab_type="text"
# 만든 데이터 확인
# + id="HVEA1hib9U5K" colab_type="code" colab={}
word_pos_entity_all[0]
| make_input_data_using_word_pos_entity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.6 64-bit (''base'': conda)'
# name: python3
# ---
# ## Introduction
#
# Unfortunately, `Matplotlib` does not have a function to generate streamcharts out of the box. Instead, it's up to you to smooth the data and use a [`stacked area plot`](https://www.python-graph-gallery.com/stacked-area-plot/) to get a streamchart.
#
# This may be quite challenging in the beginning. But don't be afraid, we have this small section for you. Here, we're gonna try to explain step-by-step how to convert a regular stackplot into a beautiful and smooth streamchart.
#
#
# ## Basic stacked area chart
# The gallery has a [whole section](https://www.python-graph-gallery.com/stacked-area-plot/) on stacked area chart and it is probably a good idea to get familiar with this kind of chart first. Let's get started by creating the most basic stackplot in Matplotlib:
# +
# Libraries
import matplotlib.pyplot as plt
import numpy as np
from scipy import stats
x = np.arange(1990, 2020) # (N,) array-like
y = [np.random.randint(0, 5, size=30) for _ in range(5)] # (M, N) array-like
fig, ax = plt.subplots(figsize=(10, 7))
ax.stackplot(x, y);
# -
# And that's it, a simple stackplot. Not too much work, not a shocking result either. A message to take from here is that `.stackplot()` recycles the values of `x`. In other words, the same 30 values in `x` are used for the 5 sequences in `y`.
# ## Change the baseline
#
# The next step is to use the `baseline` argument, which controls how to stack the different areas. The default is `"zero"` and it means the baseline is a constant at zero and the result is the simple stackplot above. Other options are `sym`, `wiggle` and `weighted_wiggle`.
#
# Let's use `sym`, which means the baseline stack is going to be symmetric around the horizontal 0 line:
fig, ax = plt.subplots(figsize=(10, 7))
ax.stackplot(x, y, baseline="sym")
ax.axhline(0, color="black", ls="--");
# If you put the colors aside, you can see the shape below the horizontal line is a mirrored version of the shape on top. Instead of stacking the areas one on top of each other, this method stacks the individual areas in such a way that both the lower and upper limits of the filled area as a whole are at the same distance from the horizontal line.
#
#
# ## Smoothing the stacked area chart
#
# The result above puts us only a tiny step closer to what we aim to have today. Having the symmetry is nice, but what we want now is to make it **smoother**. To do so, we use a data smoothing technique.
#
# Smoothing is a process by which data points are averaged with their neighbors. For example, the value of `y` when `x` is 2000 is an average of the points around `y`. There are many data smoothing techniques. In this problem, we're going to use a Gaussian kernel smoothing.
#
# The **kernel** for smoothing defines the shape of the function that is used to take the average of the neighboring points. A Gaussian kernel is a kernel with the shape of a Gaussian curve. Here is a standard Gaussian with a mean of 0 and a standard deviation of 1:
grid = np.linspace(-3, 3, num=100)
plt.plot(grid, stats.norm.pdf(grid));
# The basic process of smoothing is very simple. It goes through the data point by point. For each data point, we generate a new value that is some function of the original value at that point and the surrounding data points. With Gaussian smoothing, we center a Gaussian curve at each point, assign weights to that point and the surrounding points according to the curve, and compute a weighted average of the points.
#
# How smooth is the Gaussian smoothing is controlled by the standard deviation of the Gaussian curve. For now, let's stick to the default standard deviation of 1.
#
# For each point `m` in the sequence `x`, put a Gaussian curve with standard deviation `sd`. In this function `x` is both the sequence and the grid at which the gaussian curve is evaluated. The `np.array()` wrap converts it into a `(len(x), len(x))` array. The next line normalizes the weights so they add up to 1 for each sequence in `y`. Finally, the function returns an array where the `x` values are a weighted average resulting from using a Gaussian smoothing.
def gaussian_smooth(x, y, sd):
weights = np.array([stats.norm.pdf(x, m, sd) for m in x])
weights = weights / weights.sum(1)
return (weights * y).sum(1)
fig, ax = plt.subplots(figsize=(10, 7))
y_smoothed = [gaussian_smooth(x, y_, 1) for y_ in y]
ax.stackplot(x, y_smoothed, baseline="sym");
# ## Use a grid to make it smoother
#
# It's definetely better than the previous result, but it's not the panacea either. The plot looks spiky yet. The problem is the Gaussian curve is evaluated at very few data points because we're using the values of `x`. In this case, it's possible to use a `grid` that spans the same range than `x`, but is much denser. The function and the plot then look as follows:
def gaussian_smooth(x, y, grid, sd):
weights = np.transpose([stats.norm.pdf(grid, m, sd) for m in x])
weights = weights / weights.sum(0)
return (weights * y).sum(1)
fig, ax = plt.subplots(figsize=(10, 7))
grid = np.linspace(1985, 2025, num=500)
y_smoothed = [gaussian_smooth(x, y_, grid, 1) for y_ in y]
ax.stackplot(grid, y_smoothed, baseline="sym");
# Yay! That looks really looks smooth now! No spiky areas anymore.
#
# Now, you may wonder how to control the degree of smoothness. Well, that's pretty simple actually. Remember the default standard deviation of 1? That can be changed. Smaller values will give more wiggly results, and bigger values will result in smoother ones. Let's better see an example:
# +
fig, ax = plt.subplots(1, 2, figsize=(12, 6))
# sd of 0.6
y_smoothed_1 = [gaussian_smooth(x, y_, grid, 0.6) for y_ in y]
# sd of 1.5
y_smoothed_2 = [gaussian_smooth(x, y_, grid, 1.5) for y_ in y]
ax[0].stackplot(grid, y_smoothed_1, baseline="sym")
ax[1].stackplot(grid, y_smoothed_2, baseline="sym");
# -
# ## Color customization
#
# And last but not least, let's see how to customize the colors of the filled areas. `stackplot` has a `colors` argument. Here you can pass a list of colors that will be used to color the different areas. If you pass less colors than the number of areas, they will be repeated.
COLORS = ["#D0D1E6", "#A6BDDB", "#74A9CF", "#2B8CBE", "#045A8D"]
fig, ax = plt.subplots(figsize=(10, 7))
# Colors in the `COLORS` list are assigned to individual areas from bottom to top.
ax.stackplot(grid, y_smoothed, colors=COLORS, baseline="sym");
# And that's it! We've seen howt to build a basic streamchart with Python and Matplotlib! You can now visit the [streamchart section](https://www.python-graph-gallery.com/streamchart/) of the gallery for more customized example!
| src/notebooks/streamchart-basic-matplotlib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Mburia/2nd-Place-Solution-UmojaHack_2021-Sendy/blob/main/UmojaHack_2021_Sendy_002.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="YUocpC9JwZRy"
# # Zindi UmojaHack Africa 2021 #2: Sendy - Delivery Rider Response Challenge (INTERMEDIATE) by UmojaHack Africa
# + [markdown] id="M3zp3rt_wsws"
# This notebook was created in Google Colab
# + colab={"base_uri": "https://localhost:8080/"} id="HaqyxSPsJmwc" outputId="658a3412-823e-4595-d901-0c69f7e8c69a"
# Importing Libraries
import requests
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
# Set user authentification path
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
# logging to avoid discovery_cache error
import logging
logging.getLogger('googleapiclient.discovery_cache').setLevel(logging.ERROR)
# Getting Id of data file and load data in Google Colab
# https://drive.google.com/drive/folders/16RYdhdDcj_glWKjXR7OnT4bfcSXUymYw?usp=sharing
file_list = drive.ListFile({'q': "'16RYdhdDcj_glWKjXR7OnT4bfcSXUymYw' in parents and trashed=false"}).GetList()
# View files in the Drive Folder
for file1 in file_list:
print('title: %s, id: %s' % (file1['title'], file1['id']))
# + id="r2Y00sMCMIwj"
# Loading the Data from the source i.e. csv
import os
from io import StringIO
# Now, we will be getting content of file by using id.
# Train file
train_file = drive.CreateFile({'id': '1V_4PpDw_HyxlJldsLVp_jD4MKJL3Z3TF'})
train_file.GetContentFile('Train.csv')
# Test file
test_file = drive.CreateFile({'id': '1IvFdJrFh5D7BCLdxIgkdc3TWDjmTMaOQ'})
test_file.GetContentFile('Test.csv')
# Rider file
rider_file = drive.CreateFile({'id': '1i8rR25xDMe73IxqGdj4CKL64yWxj4ypP'})
rider_file.GetContentFile('Riders.csv')
# Data URL's
train_url = 'https://drive.google.com/file/d/1V_4PpDw_HyxlJldsLVp_jD4MKJL3Z3TF/view?usp=sharing'
test_url = 'https://drive.google.com/file/d/1IvFdJrFh5D7BCLdxIgkdc3TWDjmTMaOQ/view?usp=sharing'
rider_url = 'https://drive.google.com/file/d/1i8rR25xDMe73IxqGdj4CKL64yWxj4ypP/view?usp=sharing'
# Function to read CSV file
def read_csv(url):
url = 'https://drive.google.com/uc?export=download&id=' + url.split('/')[-2]
csv_raw = requests.get(url).text
csv = StringIO(csv_raw)
return csv
# Load the data
import pandas as pd
df_train = pd.read_csv(read_csv(train_url))
df_test = pd.read_csv(read_csv(test_url))
df_riders = pd.read_csv(read_csv(rider_url))
# + id="CEISdfQjO7kV"
# Creating Copy to work with
train = df_train.copy()
test = df_test.copy()
riders = df_riders.copy()
# + colab={"base_uri": "https://localhost:8080/"} id="qOljWkRzQQTX" outputId="e731924e-1191-4b73-e99a-840832ed60c6"
# Determining the no. of records in our datasets
print('The train dataset has ', train.shape[1], ' columns and ', train.shape[0], ' rows!')
print('The test dataset has ', test.shape[1], ' columns and ', test.shape[0], ' rows!')
print('The test dataset has ', riders.shape[1], ' columns and ', riders.shape[0], ' rows!')
# + id="3dvoeaoaQghx"
# Separate target variable
target = train['target']
# + id="AGlxebDVSjnk"
# Drop target column then join Train and test
train.drop('target',axis = 1, inplace = True)
# Merge the train and test dataframe
frames = [train,test]
df = pd.concat(frames)
# + colab={"base_uri": "https://localhost:8080/"} id="MdR9eN7_clKG" outputId="1afdfd8b-51eb-407a-c4f2-473d0dc86c59"
# Checking unique values and number of rows asigned for each column in the dataset
for i in df.columns:
print('\n')
print(i,df[i].nunique())
print(i,df[i].unique())
# + [markdown] id="D6Io2tDPoq79"
# ## Feature Engineering
# + [markdown] id="Nbo7yFgw-ndk"
# ##### Rider Amount Class
# + colab={"base_uri": "https://localhost:8080/", "height": 592} id="ybgR3jTJBq5a" outputId="5005c0c6-7e5b-4ed9-f005-dfad7098d3d2"
# Let's view the distribution of the rider's payment for the dispatch
hist_amount = df.rider_amount.hist(bins=25,figsize=[15,10])
# + colab={"base_uri": "https://localhost:8080/", "height": 155} id="k0AROAE44w8a" outputId="45206b11-3da7-47f2-e7ae-814f28a815e4"
import numpy as np
# create a list of our conditions
conditions = [
(df['rider_amount'] <= 390),
(df['rider_amount'] > 390) & (df['rider_amount'] <= 650),
(df['rider_amount'] > 650) & (df['rider_amount'] <= 820),
(df['rider_amount'] > 800) & (df['rider_amount'] <= 1200)
]
# create a list of the values we want to assign for each condition
values = ['2', '4', '3', '1']
# create a new column and use np.select to assign values to it using our lists as arguments
df['rider_amount_range'] = np.select(conditions, values)
# display updated DataFrame
df.head(3)
# + [markdown] id="ghYjG_ZBDHLg"
# Day of month
# + colab={"base_uri": "https://localhost:8080/", "height": 340} id="LCpk_dUjDKG9" outputId="dac5c7f8-4e61-4abe-ed65-364349002e61"
# Dispatches frequency per day of month
df['dispatch_day'].value_counts().plot(kind='bar',figsize=[15,5])
# + colab={"base_uri": "https://localhost:8080/", "height": 155} id="Y0JnaIY9DgIE" outputId="4f48d807-20cd-4202-c438-6814aa116b69"
# create a list of our conditions
conditions = [
(df['dispatch_day'] == 1),
(df['dispatch_day'] > 1) & (df['dispatch_day'] <= 6),
(df['dispatch_day'] > 6) & (df['dispatch_day'] <= 8),
(df['dispatch_day'] >8)
]
# create a list of the values we want to assign for each condition
values = ['1', '2','3','4']
# create a new column and use np.select to assign values to it using our lists as arguments
df['dispatch_day_range'] = np.select(conditions, values)
# display updated DataFrame
df.head(3)
# + [markdown] id="zBmGlejzDEUT"
# Weekends
# + colab={"base_uri": "https://localhost:8080/", "height": 334} id="uDzCPnOtFw1e" outputId="460db8af-1e17-4b6a-d662-79cfafc61a6c"
# Dispatches frequency per weekday
df['dispatch_day_of_week'].value_counts().plot(kind='bar',figsize=[15,5])
# + colab={"base_uri": "https://localhost:8080/", "height": 155} id="Pt5hrt9MDZs9" outputId="32a1b221-723e-4621-968a-4b3ac85aa8f4"
# create a list of our conditions
conditions = [
(df['dispatch_day_of_week'] == 1),
(df['dispatch_day_of_week'] == 7)
]
# create a list of the values we want to assign for each condition
values = ['2', '1']
# create a new column and use np.select to assign values to it using our lists as arguments
df['weekend'] = np.select(conditions, values)
# display updated DataFrame
df.head(3)
# + [markdown] id="OWIkdougI8VJ"
# Busy Hours
# + colab={"base_uri": "https://localhost:8080/", "height": 340} id="0jGAN0_AI9wX" outputId="0c428593-39af-4d16-e037-a6ca57367310"
# Convert time column to datetime
df['dispatch_time'] = pd.to_datetime(df['dispatch_time'])
# Extract Hour Column
df['dispatch_hour'] = df['dispatch_time'].dt.hour
# Let's view the hourly distribution of dispatches
df['dispatch_hour'].value_counts().plot(kind='bar',figsize=[15,5])
# + id="7bhVXbdr_UBS"
# Let's get the cos and cosine of our Time Variable
time_column = ["dispatch_time"]
for col in time_column:
df[col.split('_')[0] + '_hour_sine'] = np.sin(2 * np.pi * df[col].dt.hour/24)
df[col.split('_')[0] + '_hour_cos'] = np.cos(2* np.pi * df[col].dt.hour/24)
# + colab={"base_uri": "https://localhost:8080/", "height": 160} id="Y0ak_At-L88L" outputId="074e19c6-fc22-4e5a-81a4-815b134cf6b9"
# create a list of our conditions
conditions = [
(df['dispatch_hour'] <= 6),
(df['dispatch_hour'] > 8) & (df['dispatch_hour'] <= 16),
(df['dispatch_hour'] > 16) & (df['dispatch_hour'] <= 20),
(df['dispatch_hour'] > 20)
]
# create a list of the values we want to assign for each condition
values = ['1', '4', '3', '2']
# create a new column and use np.select to assign values to it using our lists as arguments
df['busy_hours'] = np.select(conditions, values)
# display updated DataFrame
df.head(2)
# + [markdown] id="QYqsgKsT-jN_"
# Geo Distance
# + id="zgU-YABBOzRg"
# Calculate distance between coordinates
def distance_calc(lat1, lng1, lat2, lng2):
lat1, lng1, lat2, lng2 = map(np.radians, (lat1, lng1, lat2, lng2))
AVG_EARTH_RADIUS = 6371 # in km
lat = lat2 - lat1
lng = lng2 - lng1
d = np.sin(lat * 0.5) ** 2 + np.cos(lat1) * np.cos(lat2) * np.sin(lng * 0.5) ** 2
h = 2 * AVG_EARTH_RADIUS * np.arcsin(np.sqrt(d))
return h
# + id="LxuINM4SO1da"
import numpy as np
# Let's create variables for the distances between the locations of the riders, pickup points and dropoff points
df['rider_dropoff_dist'] = distance_calc(df.rider_lat, df.rider_long, df.drop_off_lat, df.drop_off_long)
df['rider_pickup_dist'] = distance_calc(df.rider_lat, df.rider_long, df.pickup_lat, df.pickup_long)
df['pickup_dropoff_dist'] = distance_calc(df.pickup_lat, df.pickup_long, df.drop_off_lat, df.drop_off_long)
# + [markdown] id="3esDBwInRX4E"
# ### Distance Ranges
# + [markdown] id="ME6ZOCMYUG9f"
# Pickup Drop_off ranges
# + colab={"base_uri": "https://localhost:8080/", "height": 392} id="Lkrg5bIUW4Jm" outputId="cf590c3a-3f6c-4908-9aa5-00af8a80fc11"
# We can now view the distribution of the distance between pickup and drop off locations
bin_values = np.arange(start=-0, stop=60, step=0.3)
df['pickup_dropoff_dist'].hist(bins=bin_values, figsize=[14,6])
# + id="x8VwJbL9SNoh" colab={"base_uri": "https://localhost:8080/", "height": 160} outputId="170b6908-68d7-4946-c87a-52261484e7b2"
# create a list of our conditions
conditions = [
(df['pickup_dropoff_dist'] <= 6),
(df['pickup_dropoff_dist'] > 6) & (df['pickup_dropoff_dist'] <= 16),
(df['pickup_dropoff_dist'] > 16) & (df['pickup_dropoff_dist'] <= 30),
(df['pickup_dropoff_dist'] > 30) & (df['pickup_dropoff_dist'] <= 50),
(df['pickup_dropoff_dist'] > 50)
]
# create a list of the values we want to assign for each condition
values = ['5','4', '3', '2', '1']
# create a new column and use np.select to assign values to it using our lists as arguments
df['pickup_dropoff_dist_range'] = np.select(conditions, values)
# display updated DataFrame
df.head(2)
# + [markdown] id="njkhyX66ULuF"
# Rider_Pickup_Ranges
# + colab={"base_uri": "https://localhost:8080/", "height": 392} id="ORwx5WhYZOa_" outputId="f9fc729b-f498-4cdf-d3a8-8c8cf724ceb8"
# Let's also look at distribution of the distance between the rider and the pickup location
bin_values = np.arange(start=-0, stop=4, step=0.1)
df['rider_pickup_dist'].hist(bins=bin_values, figsize=[14,6])
# + id="srO_8NMJSMmW" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="d65526b3-6536-4e98-9bad-141935d9ad3b"
# create a list of our conditions
conditions = [
(df['rider_pickup_dist'] <= 1),
(df['rider_pickup_dist'] > 1) & (df['rider_pickup_dist'] <= 1.5),
(df['rider_pickup_dist'] > 1.5) & (df['rider_pickup_dist'] <= 2.5),
(df['rider_pickup_dist'] > 2.5) & (df['rider_pickup_dist'] <= 3.7),
(df['rider_pickup_dist'] > 3.7)
]
# create a list of the values we want to assign for each condition
values = ['1', '2', '3', '4','5']
# create a new column and use np.select to assign values to it using our lists as arguments
df['rider_pickup_dist_range'] = np.select(conditions, values)
# display updated DataFrame
df.head(3)
# + [markdown] id="7SbdQYYGUQzT"
# Rider Drop_off_Ranges
# + colab={"base_uri": "https://localhost:8080/", "height": 392} id="ix7qTLHCWF6C" outputId="059181ab-f7f6-48a2-afc7-c0bfe3158a57"
# Let's also visualize the distance between the rider and drop off location
bin_values = np.arange(start=-0, stop=30, step=0.3)
df['rider_dropoff_dist'].hist(bins=bin_values, figsize=[14,6])
# + id="Q4-N75D3RWOt" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="8f8e0ef7-ac9c-494d-c1ae-306557a73a42"
# create a list of our conditions
conditions = [
(df['rider_dropoff_dist'] <= 5),
(df['rider_dropoff_dist'] > 5) & (df['rider_dropoff_dist'] <= 10),
(df['rider_dropoff_dist'] > 10) & (df['rider_dropoff_dist'] <= 18),
(df['rider_dropoff_dist'] > 18) & (df['rider_dropoff_dist'] <= 30),
(df['rider_dropoff_dist'] > 30)
]
# create a list of the values we want to assign for each condition
values = ['1', '2', '3', '4','5']
# create a new column and use np.select to assign values to it using our lists as arguments
df['rider_dropoff_dist_range'] = np.select(conditions, values)
# display updated DataFrame
df.head(3)
# + [markdown] id="j25FtQ-BNtbQ"
# Time and Distance
# + id="EXROX-OGNxp0" colab={"base_uri": "https://localhost:8080/", "height": 160} outputId="7a7e21bf-ee68-415e-f1df-f2e5624be3c1"
# create a list of our conditions
conditions = [
(df['rider_dropoff_dist_range'] == '5') & (df['busy_hours'] == '1'),
(df['rider_dropoff_dist_range'] == '4') & (df['busy_hours'] == '2'),
(df['rider_dropoff_dist_range'] == '3') & (df['busy_hours'] == '3'),
(df['rider_dropoff_dist_range'] == '2') & (df['busy_hours'] == '4'),
(df['rider_dropoff_dist_range'] == '1') & (df['busy_hours'] == '4'),
(df['rider_dropoff_dist_range'] == '2') & (df['busy_hours'] == '0')
]
# create a list of the values we want to assign for each condition
values = ['1', '2', '3', '4','6','5']
# create a new column and use np.select to assign values to it using our lists as arguments
df['time_distance_bias'] = np.select(conditions, values)
# display updated DataFrame
df.head(2)
# + [markdown] id="rUM3CoTdaHcu"
# Weekday vs Time_distance
# + colab={"base_uri": "https://localhost:8080/", "height": 160} id="PCT1_1t7aHDz" outputId="7f40a454-66c0-42d8-81c3-692bb9001b47"
# create a list of our conditions
conditions = [
(df['dispatch_day_of_week'] == '1') & (df['time_distance_bias'] == '5'), # Sunday early morning & average distance
(df['dispatch_day_of_week'] == '7') & (df['time_distance_bias'] == '3'), # Saturday late night & long distance
(df['dispatch_day_of_week'] == '7') & (df['time_distance_bias'] == '5'), # Saturday early morning average distance
(df['dispatch_day_of_week'] == '0') & (df['time_distance_bias'] == '3'), # Weekday late night & long distance
(df['dispatch_day_of_week'] == '0') & (df['time_distance_bias'] == '6') # Weekday daytime and average distance
]
# create a list of the values we want to assign for each condition
values = ['1', '2', '3', '4','5']
# create a new column and use np.select to assign values to it using our lists as arguments
df['weekend_time_distance_bias'] = np.select(conditions, values)
# display updated DataFrame
df.head(2)
# + [markdown] id="Jo6nn_a811UH"
# Money Earned VS time & Distance
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="eWfti7qK11oF" outputId="6e06e9b8-936f-40bf-bc63-538ef23a84b9"
# create a list of our conditions
conditions = [
(df['rider_amount_range'] == '4') & (df['busy_hours'] == '4'),
(df['rider_pickup_dist_range'] == '3') & (df['busy_hours'] == '2'),
(df['rider_pickup_dist_range'] == '1') & (df['rider_amount_range'] == '4'),
(df['rider_pickup_dist_range'] == '4') & (df['rider_amount_range'] == '4')
]
# create a list of the values we want to assign for each condition
values = ['4','2','6','1']
# create a new column and use np.select to assign values to it using our lists as arguments
df['money_distance_bias'] = np.select(conditions, values)
# display updated DataFrame
df.head(3)
# + [markdown] id="3j2Ys73j2gRS"
# Correcting Dtypes
# + id="mDA26qKr0HHY"
# let's convert some string columns to numeric
df[['rider_dropoff_dist_range', 'time_distance_bias', 'weekend_time_distance_bias','busy_hours','weekend','money_distance_bias']] = df[['rider_dropoff_dist_range', 'time_distance_bias', 'weekend_time_distance_bias','busy_hours','weekend','money_distance_bias']].apply(pd.to_numeric)
# + [markdown] id="RFjpENv-Vxmc"
# ## Model
# + colab={"base_uri": "https://localhost:8080/"} id="XJ6GViaTYmes" outputId="a7893c6f-6c92-4dfa-c868-ebdd94603ba8"
# Lets install catboost for creating a model
# !pip install catboost
# + id="waRxBsBYgN3r"
# Split train and test set
train = df.iloc[:179867, :]
test = df.iloc[179867:, :]
# + id="bcppiidAVHCG"
# Selecting the columns to be used as data for accuracy testing
cols = [col for col in train.columns if col not in ['dispatch_time']]
# Set the target and independent variables
X = train[cols]
y = target
# + id="UCuxJEyEaOK9"
test.drop(['dispatch_time'],axis = 1, inplace=True)
# + id="OuQJtltaYqQQ"
# Fitting the data to model
from catboost import CatBoostClassifier
# You can increase the iterations parameter but it will take longer to learn
model = CatBoostClassifier(iterations = 1000, early_stopping_rounds = 50)
# + colab={"base_uri": "https://localhost:8080/"} id="XoP0CgdhZecY" outputId="0ce1077d-b606-45d7-e497-7824765bcfcf"
# Here we fit our data then make predictions
import numpy as np
# Since Catboost accepts categorical features we'll make then identifiable to the model
categorical_features_indices = np.where(X.dtypes != np.float)[0]
# Fitting our data to the model
model.fit(X,y,cat_features=categorical_features_indices)
# Making Predictions
y_pred = model.predict(test)
model.score(X,y)
# + id="nYK5BtKjbGGd"
# Let's create a submission file to input our predictions
submissionFile= pd.DataFrame({"ID":df_test["ID"]})
submissionFile['target'] = y_pred
# + id="rr_FOnjedsMn"
# Saving our data in a CSV
submissionFile.to_csv("submit.csv",index=False)
| UmojaHack_2021_Sendy_002.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Deep Learning
#
#
# ### Neural Networks
# A neural network is composed of layers, each containing neurons or units as they are called nowadays, see image. The goal in Deep Learning is to create powerful models that learn features in the input data. A neural network is a function composition of layers.
#
from IPython.display import Image
Image(filename="figures/complete_cnn.jpeg")
# ## Convolutional Neural Networks
# In computer vision convolutional neural networks are used for example in image classification, object detection, image captioning and semantic segmentation among other things.
#
# The principal components in an CNN are:
# * convolution layers
# * max or average pooling
# * full connected layers
#
#
# ### Convolutional layer
Image(filename="figures/convolution.png")
import tensorflow as tf
from functools import partial
from tensorflow import keras
import numpy as np
tf.random.set_seed(42)
# Download data from keras. Use the cifar 10.
# + active=""
# (x_train, y_train), (x_test, y_test) = tf.keras.datasets.cifar10.load_data()
# +
fashion_mnist = keras.datasets.fashion_mnist
(x_train, y_train), (x_test, y_test) = fashion_mnist.load_data()
num_classes = 10
# Make sure images have shape (28, 28, 1)
x_train = np.expand_dims(x_train, -1)
x_test = np.expand_dims(x_test, -1)
# convert class vectors to binary class matrices
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
# -
x_train.shape, x_train[0, :, :].max(), x_train[0, :, :].min()
x_train, x_test = x_train/255.0, x_test/255.0
# # Create model
def exponential_decay(lr0, s):
def exponential_decay_fn(epoch):
return lr0 * 0.1 ** (epoch/s)
return exponential_decay_fn
def create_cnn(activation='relu',
padding='same',
input_shape=[28, 28, 1],
output_dim=10):
"""Create convolutional neural network"""
model = tf.keras.Sequential()
partial_cnn = partial(tf.keras.layers.Conv2D, activation=activation, padding=padding)
model.add(partial_cnn(64, 7, input_shape=input_shape))
model.add(tf.keras.layers.BatchNormalization())
model.add(tf.keras.layers.MaxPooling2D(2))
model.add(partial_cnn(128, 3))
model.add(tf.keras.layers.BatchNormalization())
model.add(partial_cnn(128, 3))
model.add(tf.keras.layers.BatchNormalization())
model.add(tf.keras.layers.MaxPooling2D(2))
model.add(partial_cnn(256, 3))
model.add(tf.keras.layers.BatchNormalization())
model.add(partial_cnn(256, 3))
model.add(tf.keras.layers.BatchNormalization())
model.add(tf.keras.layers.MaxPooling2D(2))
model.add(tf.keras.layers.Flatten())
model.add(tf.keras.layers.Dense(128, activation='relu'))
model.add(tf.keras.layers.Dropout(0.5))
model.add(tf.keras.layers.Dense(64, activation='relu'))
model.add(tf.keras.layers.Dropout(0.5))
model.add(tf.keras.layers.Dense(output_dim, activation='softmax'))
optimizer = tf.keras.optimizers.Nadam()
model.compile(optimizer=optimizer,
loss='categorical_crossentropy',
metrics=['accuracy'])
return model
model = create_cnn()
model.summary()
x_train.shape, y_train.shape
# # Train Model
# +
exponential_decay_fn = exponential_decay(lr0=0.1, s=10)
exp_schedule = tf.keras.callbacks.LearningRateScheduler(exponential_decay_fn)
early_stopping = tf.keras.callbacks.EarlyStopping(patience=10)
callbacks = [early_stopping,
exp_schedule]
history = model.fit(x_train, y_train,
validation_data=(x_test, y_test),
epochs=100,
batch_size=32,
# callbacks=callbacks
)
# -
import pandas as pd
import matplotlib.pyplot as plt
pd.DataFrame(history.history).plot(figsize=(8, 5), grid=True)
plt.gca().set_ylim(0, 1)
plt.show();
# ### Exercise
# Replace batch-normalization with the SELU activation function. Is the performance better?
# * Normalize the input
# * Use LeCun normal initialization
# * Make sure that the DNN contains only a sequence of dense layers
| notebooks/CNN on Fashion MNIST tf.keras Batch Normalization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <b>Calcule a integral dada</b>
# $7. e^{1 - x}dx$
# $u = 1 - x$
# $du = -dx$
# <b>Substituindo $1-x$ por $u$</b>
# $-\int e^udu$
# <b>Integrando $-\int e^udu$</b>
# $-\int e^u du = e^u + C $
# <b>Substituindo $u$ por $1-x$</b>
# $-\int e^u du = -e^{1-x} + C$
| Problemas 5.2/07.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/pragmatizt/DS-Unit-3-Sprint-1-Software-Engineering/blob/master/IRA_E_Unit2_Assignment_2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="ciMzs64ZfL8e" colab_type="code" colab={}
## This is the class I created that has several methods of finding nulls,
## extracting the X and y, and then doing a train_test_split on the data.
from sklearn.model_selection import train_test_split
class Cleaning_Data():
def __init__(self, df, target):
self.df = df
self.target = target
def find_nulls(self):
return self.df.isnull().sum().sort_values(ascending=True)
def extract_target(self):
return self.df.drop(self.target, axis = 1)
def extract(self):
return self.df[self.target]
def ira_t_t_split(self):
self.X = self.extract_target()
self.y = self.extract()
X_train, X_val, y_train, y_val = train_test_split(
self.X, self.y, train_size=0.5, test_size=0.5, random_state=42)
print("X_train:", X_train.shape)
print("X_val:", X_val.shape)
print("y_train:", y_train.shape)
print("y_val:", y_val.shape)
return (X_train, X_val, y_train, y_val)
| IRA_E_Unit2_Assignment_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Зависимости
import gym
import numpy
from collections import deque
import keras
from keras.models import Sequential, load_model
from keras.layers import Dense
from keras.optimizers import Adam
from random import sample
# Создание нейронной сети, которая для заданного состояния S вычисляет значения Q(s,a) для всех возможных a.
def create_model(state_size, num_actions) :
learning_rate = 0.01
# Структура сети
input_layer_size = state_size
hidden_layers_size = [10, 10, 10]
output_layer_size = num_actions
# Построение сети
model = Sequential()
model.add(Dense(hidden_layers_size[0], input_dim=input_layer_size, activation="relu"))
for layer in range(1, len(hidden_layers_size)) :
model.add(Dense(hidden_layers_size[layer], activation="relu"))
model.add(Dense(output_layer_size))
model.compile(loss="mean_squared_error", optimizer=Adam(lr=learning_rate))
return model
# Обучение сети с использованием ранее заполненной памяти
def fit_on_memory(model, memory, state_size):
# Размер батча для извлечения из памяти
batch_size = 16
# Коэффициент уменьшения (дисконта)
gamma = 0.95
# Случайное получение батча, чтобы обучаться на нём
batch = sample(memory, min(len(memory), batch_size))
# Извлечение значений из батча
actions, rewards, results = [], [], []
states = numpy.zeros((batch_size, state_size))
next_states = numpy.zeros((batch_size, state_size))
for i in range(batch_size) :
states[i] = batch[i][0]
actions.append(batch[i][1])
rewards.append(batch[i][2])
next_states[i] = batch[i][3]
results.append(batch[i][4])
# Вычисление значений Q для текущего и следующего состояний
q_values = model.predict(states)
q_next_values = model.predict(next_states)
# Коррекция значений Q
for t in range(batch_size) :
# Если мы на последнем шаге, то значение Q равно полученному подкреплению
if results[t] == True :
q_values[t][actions[t]] = rewards[t]
# Иначе мы учитываем дисконтированный максимум из следующих значений Q
else :
q_values[t][actions[t]] = rewards[t] + gamma * (numpy.amax(q_next_values[t]))
# Обучение сети
model.fit(states, q_values, batch_size=batch_size, verbose=0)
# Выполнение действий в окружающей среде, заполнение памяти и обучение на ней
def try_and_learn(env, model, model_name, num_runs, save_condition, logs = True) :
# Параметры памяти
maximum_memory_size = 500
memory = deque(maxlen = maximum_memory_size)
minimum_memory_size_for_learning = 50
# Параметры выбора действия
epsilon = 1.0
epsilon_min = 0.01
epsilon_decay = 0.9995
# Значение по умолчанию для отрицательного подкрепления, если итерация закончилась преждевременно
negative_reward = -100
state_size = env.observation_space.shape[0]
# В каждой итерации мы пытаемся обучить сеть, чтобы достичь заданный макисмум эффективности
for run in range(num_runs) :
env_state = env.reset()
state = numpy.reshape(env_state, [1, state_size])
done = False
steps_survived = 0
while not done :
# Сначала действие выбирается случайным образом,
# но если эпсилон-жадное правило позволяет, то действие выбирается в соответствии с прогнозом сети
action = env.action_space.sample()
random_pick = numpy.random.random()
if random_pick > epsilon :
action = numpy.argmax(model.predict(state))
# Если память достаточно велика, мы начинаем уменьшать эпсилон
if len(memory) >= minimum_memory_size_for_learning and epsilon > epsilon_min :
epsilon = max(epsilon * epsilon_decay, epsilon_min)
# Взаимодействие с окружающей средой
env_next_state, reward, done, info = env.step(action)
next_state = numpy.reshape(env_next_state, [1, state_size])
# Если итерация завершена, но сеть прожила меньше заданного максимума, то подкрепление отрицательное
if done and steps_survived != env._max_episode_steps - 1 :
reward = negative_reward
# Добавление записи в память
memory.append((state, action, reward, next_state, done))
# Переход в следующее состояние
state = next_state
steps_survived += 1
# Если итерация завершена, то выводятся некоторые логи
if done :
if logs : print("run: {}/{}, score: {}/{}, eps: {:.6}, len(memory): {}".format(run, num_runs, steps_survived, save_condition, epsilon, len(memory)))
# Если сеть прожила больше заданного значения, то она сохраняется в файл и обучение заканчивается
if steps_survived >= save_condition :
if logs : print("Saving the model as {}".format(model_name))
model.save(model_name)
return run
# Если память достаточно велика, начиаем обучение
if len(memory) >= minimum_memory_size_for_learning :
fit_on_memory(model, memory, env.observation_space.shape[0])
# Если код дошёл до этой строки, значит сеть не смогла обучиться
if logs : print("The model is not good enough")
return -1
# Тестирование поведения обученной сети
def test(env, model_name, num_runs, render = True, logs = True):
# Загрузка сети
model = load_model(model_name)
state_size = env.observation_space.shape[0]
sum_result = 0
complete_success = 0
for run in range(num_runs) :
env_state = env.reset()
state = numpy.reshape(env_state, [1, state_size])
done = False
steps_survived = 0
while not done :
if render :
env.render()
action = numpy.argmax(model.predict(state))
env_next_state, reward, done, info = env.step(action)
state = numpy.reshape(env_next_state, [1, state_size])
steps_survived += 1
if done :
if logs : print("run: {}/{}, score: {}".format(run, num_runs, steps_survived))
sum_result += steps_survived
if steps_survived == env._max_episode_steps :
complete_success += 1
break
return sum_result / num_runs, complete_success
# +
# Инициализация окружающей среды
env_name = 'CartPole-v1'
env = gym.make(env_name)
print(env.observation_space.shape[0])
print(env.action_space.n)
print(env._max_episode_steps)
# -
# Параметры обучения и тестирования
num_runs = 1000
save_condition = 500
num_test_runs = 10
# Инициализация и запуск сети
model_name = 'basic_model/basic.model'
model = create_model(env.observation_space.shape[0], env.action_space.n)
last_run = try_and_learn(env, model, model_name, num_runs, min(save_condition, env._max_episode_steps), True)
if last_run == -1 :
average_result = 0
complete_success = 0
else :
average_result, complete_success = test(env, model_name, num_test_runs, False, True)
print("LR: {}, AR: {}, CS: {}".format(last_run, average_result, complete_success))
# +
# Раскомментировать и использовать для остановки визуализации, если она была ранее запущена
# env.close()
| reinforcement_learning_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assessing
# Use the space below to explore `winequality-red.csv` and `winequality-white.csv` to answer the quiz questions below.
# +
import pandas as pd
# %matplotlib inline
df_red = pd.read_csv('winequality-red.csv',';')
df_white = pd.read_csv('winequality-white.csv',';')
# +
print('Red Wine Sample Info:\n')
df_red.info()
print('White Wine Sample Info:\n')
df_white.info()
print('# of Columns in Red Wine Sample is {}, '.format(len(df_red.columns)) +
'# in of Columns in White Wine Sample is {}'.format(len(df_white.columns)))
print(sum(df_white.duplicated()))
df_red_quality = df_red['quality']
print(df_red_quality.nunique())
df_white_quality = df_white['quality']
print(df_white_quality.nunique())
# -
| assessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Title: Pan Tropical Forest Strata
#
# ### Description
# The purpose for stratifying forest cover was to delineate regions (strata) associated with different carbon stock reference values in order to estimate aboveground carbon loss from tropical forest disturbance using sample-based forest loss area estimate based on forest cover loss map by Hansen et al. (2013). However, consistently characterized pan-tropical forest type maps are not available at the 30-m spatial resolution corresponding to the forest loss data. Characterizing forest cover based on complex multi-parameter definitions (e.g. "primary forests", "secondary forests", "woodlands") as we have performed at a national scale (Potapov et al 2012) is not easily achieved at a biome scale. Instead, we defined tropical forest strata using remotely sensed-derived structural characteristics of tree canopy (year 2000 percent tree canopy cover (Hansen et al 2013)), tree height (current study) and forest intactness (Potapov et al 2008). Stratification thresholds were developed to minimize within-strata AGC variance using a statistical regression tree approach with point-based GLAS carbon estimates (Baccini et al 2012) for the period 2003 - 2008 as the dependent variable.<br>
#
# Three pantropical regions - South America, Southeast Asia and central Africa.
#
# ### FLINT
# This dataset has been checked/pre-processed and is suitable for use in FLINT. Please adhere to individual dataset licence conditions and citations. Processed data can be accessed here: [Land Sector Datasets](https://datasets.mojaglobal.workers.dev/) https://datasets.mojaglobal.workers.dev/<br>
#
# ### Format
# <b>Extent: </b>Three pantropical regions - South America, Southeast Asia and central Africa (-180, -60, 180, 40) <br>
# <b>Resolution:</b> Raster 0.00025-degree<br>
# <b>File type:</b> geotiff (.tif)<br>
# <b>Cordinate system:</b> EPSG:4326 (WGS84)<br>
# <b>Temporal Resolution: </b>2003 - 2008<br>
# <b>Size:</b>6 GB+
#
# ### Original source
# Original data sourced from Global Land Analysis & Discovery - Pan Tropical Forest Strata: https://glad.umd.edu/dataset/pan-tropical-forest-strata Access 08/01/2020.<br>
# Format: geoTIFF, three regions Africa, SE Asia and Sth America. Pixel size: ~0.000278, Cordinate system: EPSG:4326 (WGS84)<br>
#
# ### Licence
# Creative Commons Attributions 3.0 CC-BY-3.0 as long as is properly cited where used. See [Additional Permissions](https://github.com/moja-global/Land_Sector_Datasets/tree/master/Additional-Permissions) for permission to use this data in FLINT.
#
# ### Citation
# Please, use the following credit when the provided data are cited: <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>. (2015) Aboveground carbon loss in natural and managed tropical forests from 2000 to 2012. Environmental Research Letters, 10(7), 074002.
#
# ### Metadata
# Numbers in the legend refer to forest strata: 1 - low cover; 2 - medium cover short; 3 - medium cover tall; 4 - dense cover short; 5 - dense cover short intact; 6 - dense cover tall; 7 - dense cover tall intact; 8: non-forest lands (treecover <25%); 9: water; 0: no data.
#
# ### Notes
# Cells were resampled to 0.00025 for use in FLINT padded to nearest degree. Additional resampling recommended from original files to reduce data loss - code provided below to batch process. Extent only cover the pan tropical zone.
#
# ### Processing
# Resample in gdal to 0.00025 keeping original regional extents. A merge might be possible but will produce large areas of NoData, and BIGTIFF will need to be enabled.
#Process (resample) resolution in gdal (osgeo4W shell)
for /R C:\inputfolder\ %f IN (*.tif) do gdalwarp.exe -co COMPRESS=DEFLATE -co PREDICTOR=2 -co ZLEVEL=9 -s_srs EPSG:4326 -tr 0.00025 0.00025 -tap -r near -of GTiff -t_srs EPSG:4326 -multi -overwrite -co BIGTIFF=YES %f C:\outputgfsad\%~nf.tif
| Data/LandCover/PanTropicalForestStrata.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pyml
# language: python
# name: pyml
# ---
import pandas as pd
import numpy as np
df_wine = pd.read_csv('wine.data', header=None)
df_wine.columns = ['Class label', 'Alcohol', 'Malic acid', 'Ash', 'Alcalinity of ash', 'Magnesium',
'Total phenols', 'Flavanoids', 'Nonflavanoid phenols', 'Proanthocyanins',
'Color intensity', 'Hue', 'OD280/OD315 of diluted wines', 'Proline']
df_wine = df_wine[df_wine['Class label'] != 1]
X = df_wine[['Alcohol', 'OD280/OD315 of diluted wines']].values
y = df_wine['Class label'].values
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
le = LabelEncoder()
y = le.fit_transform(y)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=1, stratify=y)
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import AdaBoostClassifier
tree = DecisionTreeClassifier(criterion='entropy', random_state=1, max_depth=1)
ada = AdaBoostClassifier(base_estimator=tree, n_estimators=500, learning_rate=0.1, random_state=1)
from sklearn.metrics import accuracy_score
tree = tree.fit(X_train, y_train)
y_train_pred = tree.predict(X_train)
y_test_pred = tree.predict(X_test)
tree_train = accuracy_score(y_train, y_train_pred)
tree_test = accuracy_score(y_test, y_test_pred)
print('Decision tree train/test accuracies %.3f/%.3f' % (tree_train, tree_test))
ada = ada.fit(X_train, y_train)
y_train_pred = ada.predict(X_train)
y_test_pred = ada.predict(X_test)
ada_train = accuracy_score(y_train, y_train_pred)
ada_test = accuracy_score(y_test, y_test_pred)
print('AdaBoost train/test accuracies %.3f/%.3f' % (ada_train, ada_test))
import matplotlib.pyplot as plt
x_min = X_train[:, 0].min() - 1
x_max = X_train[:, 0].max() + 1
y_min = X_train[:, 1].min() - 1
y_max = X_train[:, 1].max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, 0.1), np.arange(y_min, y_max, 0.1))
f, axarr = plt.subplots(nrows=1, ncols=2, sharex='col', sharey='row', figsize=(8, 3))
for idx, clf, tt in zip([0, 1], [tree, ada], ['Decision tree', 'AdaBoost']):
clf.fit(X_train, y_train)
Z = clf.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
axarr[idx].contourf(xx, yy, Z, alpha=0.3)
axarr[idx].scatter(X_train[y_train==0, 0], X_train[y_train==0, 1], c='blue', marker='^')
axarr[idx].scatter(X_train[y_train==1, 0], X_train[y_train==1, 1], c='green', marker='o')
axarr[idx].set_title(tt)
axarr[0].set_ylabel('Alcohol', fontsize=12)
plt.text(10.2, -1.2, s='OD280/OD315 of diluted wines', ha='center', va='center', fontsize=12)
plt.show()
| ch07/wine_adaboost.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.9 64-bit (''yolov3_huynhngocanh'': conda)'
# metadata:
# interpreter:
# hash: 2880e12141afd390ebaeda03ff70ed4419875b829829d7a048b3efdcfe3aa086
# name: python3
# ---
# Import libraries
# %matplotlib inline
from pycocotools.coco import COCO
import tensorflow
KERAS_BACKEND=tensorflow
from keras.models import load_model
# from utils.utils import *
# from utils.bbox import *
# from utils.image import load_image_pixels
from keras.preprocessing.image import load_img
from keras.preprocessing.image import img_to_array
import numpy as np
import pandas as pd
# import skimage.io as io
import matplotlib.pyplot as plt
import pylab
# import torchvision.transforms.functional as TF
# import PIL
import os
import json
from urllib.request import urlretrieve
pylab.rcParams['figure.figsize'] = (8.0, 10.0)
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3'
# Define image directory
projectDir=os.getcwd()
dataDir='..'
imageDir='{}/images/'.format(dataDir)
# +
class BoundBox:
def __init__(self, xmin, ymin, xmax, ymax, objness = None, classes = None):
self.xmin = xmin
self.ymin = ymin
self.xmax = xmax
self.ymax = ymax
self.objness = objness
self.classes = classes
self.label = -1
self.score = -1
def get_label(self):
if self.label == -1:
self.label = np.argmax(self.classes)
return self.label
def get_score(self):
if self.score == -1:
self.score = self.classes[self.get_label()]
return self.score
def _sigmoid(x):
return 1. / (1. + np.exp(-x))
def decode_netout(netout, anchors, obj_thresh, net_h, net_w):
grid_h, grid_w = netout.shape[:2]
nb_box = 3
netout = netout.reshape((grid_h, grid_w, nb_box, -1))
nb_class = netout.shape[-1] - 5
boxes = []
netout[..., :2] = _sigmoid(netout[..., :2])
netout[..., 4:] = _sigmoid(netout[..., 4:])
netout[..., 5:] = netout[..., 4][..., np.newaxis] * netout[..., 5:]
netout[..., 5:] *= netout[..., 5:] > obj_thresh
for i in range(grid_h*grid_w):
row = i // grid_w
col = i % grid_w
for b in range(nb_box):
# 4th element is objectness score
objectness = netout[int(row)][int(col)][b][4]
if(objectness.all() <= obj_thresh): continue
# first 4 elements are x, y, w, and h
x, y, w, h = netout[int(row)][int(col)][b][:4]
x = (col + x) / grid_w # center position, unit: image width
y = (row + y) / grid_h # center position, unit: image height
w = anchors[2 * b + 0] * np.exp(w) / net_w # unit: image width
h = anchors[2 * b + 1] * np.exp(h) / net_h # unit: image height
# last elements are class probabilities
classes = netout[int(row)][col][b][5:]
box = BoundBox(x-w/2, y-h/2, x+w/2, y+h/2, objectness, classes)
boxes.append(box)
return boxes
def correct_yolo_boxes(boxes, image_h, image_w, net_h, net_w):
new_w, new_h = net_w, net_h
for i in range(len(boxes)):
x_offset, x_scale = (net_w - new_w)/2./net_w, float(new_w)/net_w
y_offset, y_scale = (net_h - new_h)/2./net_h, float(new_h)/net_h
boxes[i].xmin = int((boxes[i].xmin - x_offset) / x_scale * image_w)
boxes[i].xmax = int((boxes[i].xmax - x_offset) / x_scale * image_w)
boxes[i].ymin = int((boxes[i].ymin - y_offset) / y_scale * image_h)
boxes[i].ymax = int((boxes[i].ymax - y_offset) / y_scale * image_h)
def _interval_overlap(interval_a, interval_b):
x1, x2 = interval_a
x3, x4 = interval_b
if x3 < x1:
if x4 < x1:
return 0
else:
return min(x2,x4) - x1
else:
if x2 < x3:
return 0
else:
return min(x2,x4) - x3
def bbox_iou(box1, box2):
intersect_w = _interval_overlap([box1.xmin, box1.xmax], [box2.xmin, box2.xmax])
intersect_h = _interval_overlap([box1.ymin, box1.ymax], [box2.ymin, box2.ymax])
intersect = intersect_w * intersect_h
w1, h1 = box1.xmax-box1.xmin, box1.ymax-box1.ymin
w2, h2 = box2.xmax-box2.xmin, box2.ymax-box2.ymin
union = w1*h1 + w2*h2 - intersect
return float(intersect) / union
def do_nms(boxes, nms_thresh):
if len(boxes) > 0:
nb_class = len(boxes[0].classes)
else:
return
for c in range(nb_class):
sorted_indices = np.argsort([-box.classes[c] for box in boxes])
for i in range(len(sorted_indices)):
index_i = sorted_indices[i]
if boxes[index_i].classes[c] == 0: continue
for j in range(i+1, len(sorted_indices)):
index_j = sorted_indices[j]
if bbox_iou(boxes[index_i], boxes[index_j]) >= nms_thresh:
boxes[index_j].classes[c] = 0
# load and prepare an image
def load_image_pixels(filename, shape):
# load the image to get its shape
image = load_img(filename)
width, height = image.size
# load the image with the required size
image = load_img(filename, target_size=shape)
# convert to numpy array
image = img_to_array(image)
# scale pixel values to [0, 1]
image = image.astype('float32')
image /= 255.0
# add a dimension so that we have one sample
image = np.expand_dims(image, 0)
return image, width, height
# get all of the results above a threshold
def get_boxes(boxes, labels, thresh):
v_boxes, v_labels, v_scores = list(), list(), list()
# enumerate all boxes
for box in boxes:
# enumerate all possible labels
for i in range(len(labels)):
# check if the threshold for this label is high enough
if box.classes[i] > thresh:
v_boxes.append(box)
v_labels.append(labels[i])
v_scores.append(box.classes[i]*100)
# don't break, many labels may trigger for one box
return v_boxes, v_labels, v_scores
# draw all results
def draw_boxes(filename, v_boxes, v_labels, v_scores):
# load the image
data = plt.imread(filename)
# plot the image
plt.imshow(data)
# get the context for drawing boxes
ax = plt.gca()
# plot each box
for i in range(len(v_boxes)):
box = v_boxes[i]
# get coordinates
y1, x1, y2, x2 = box.ymin, box.xmin, box.ymax, box.xmax
# calculate width and height of the box
width, height = x2 - x1, y2 - y1
# create the shape
rect = plt.Rectangle((x1, y1), width, height, fill=False, color='white')
# draw the box
ax.add_patch(rect)
# draw text and score in top left corner
label = "%s (%.3f)" % (v_labels[i], v_scores[i])
plt.text(x1, y1, label, color='white')
# show the plot
plt.show()
# + tags=[]
# load yolov3 model
model = load_model('../yolov3_model.h5')
# define the expected input shape for the model
input_w, input_h = 416, 416
# define the anchors
anchors = [[116,90, 156,198, 373,326], [30,61, 62,45, 59,119], [10,13, 16,30, 33,23]]
# define the probability threshold for detected objects
class_threshold = 0.6
# define the labels
labels = ["person", "bicycle", "car", "motorbike", "airplane", "bus", "train", "truck",
"boat", "traffic light", "fire hydrant", "stop sign", "parking meter", "bench",
"bird", "cat", "dog", "horse", "sheep", "cow", "elephant", "bear", "zebra", "giraffe",
"backpack", "umbrella", "handbag", "tie", "suitcase", "frisbee", "skis", "snowboard",
"sports ball", "kite", "baseball bat", "baseball glove", "skateboard", "surfboard",
"tennis racket", "bottle", "wine glass", "cup", "fork", "knife", "spoon", "bowl", "banana",
"apple", "sandwich", "orange", "broccoli", "carrot", "hot dog", "pizza", "donut", "cake",
"chair", "couch", "pottedplant", "bed", "diningtable", "toilet", "tvmonitor", "laptop", "mouse",
"remote", "keyboard", "cell phone", "microwave", "oven", "toaster", "sink", "refrigerator",
"book", "clock", "vase", "scissors", "teddy bear", "hair drier", "toothbrush"]
# -
file_path = "../images/000000206831.jpg"
image, image_w, image_h = load_image_pixels(file_path, (input_w, input_h))
yhat = model.predict(image)
print(yhat)
| paper_followup/test_yolov3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/pcsilcan/aed/blob/master/week14/14_graph_nice_adjacency_list.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="ne1-bfno45MN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="8e79da99-0e52-41bd-ce6b-aff428ab5f86"
# %%writefile graph.h
#ifndef __GRAPH_H__
#define __GRAPH_H__
#include <fstream>
#include <map>
#include <queue>
#include <string>
#include <vector>
using namespace std;
typedef pair<int, float> edge;
typedef vector<edge> vp;
typedef vector<vp> vvp;
/*
nodeNames:
"felipe": 0
"rosa" : 1
"jose" : 2
"ricky" : 3
"vicky" : 4
...
G:
0: (2, 1.0), (4, 1.0)
1: (3, 1.0)
2: (0, 1.0)
3: (1, 1.0), (4, 1.0)
4: (0, 1.0), (3, 1.0)
*/
class Graph {
vvp G;
bool directed;
map<string, int> name2i;
vector<string> i2name;
public:
Graph(bool directed=true) : directed(directed) {}
void addNode(string nodeName) {
name2i[nodeName] = G.size();
i2name.push_back(nodeName);
G.push_back(vp());
}
void addEdge(string nameU, string nameV, float w=1.0) {
int u = name2i[nameU];
int v = name2i[nameV];
G[u].push_back(edge(v, w));
if (!directed) {
G[v].push_back(edge(u, w));
}
}
map<string, string> bfs(string nameS) {
auto s = name2i[nameS];
auto n = G.size();
vector<bool> visited(n, false);
vector<int> path(n, -1);
queue<int> q;
visited[s] = true;
q.push(s);
while (!q.empty()) {
auto u = q.front();
for (auto edge : G[u]) {
auto v = edge.first;
if (!visited[v]) {
visited[v] = true;
path[v] = u;
q.push(v);
}
}
q.pop();
}
map<string, string> result;
for (int i = 0; i < n; ++i) {
result[i2name[i]] = path[i] == -1? "" : i2name[path[i]];
}
return result;
}
void saveDot(string fileName) {
ofstream dot(fileName);
string rel = directed? "->" : "--";
dot << (directed? "digraph" : "strict graph") << " G {\n";
for (auto kv : name2i) {
dot << " " << kv.second << "[label=\"" << kv.first << "\"];\n";
}
dot << endl;
for (auto kv : name2i) {
auto u = kv.second;
for (auto edge : G[u]) {
auto v = edge.first;
auto w = edge.second;
dot << " " << u << rel << v << ";\n";
}
}
dot << "}";
}
};
#endif
# + id="HB0amzdg-3Tr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="077f71b0-b20a-47dd-e5fd-7e72edf6f3c3"
# %%writefile graphTest.cpp
#include <iostream>
#include "graph.h"
using namespace std;
int main() {
Graph* G = new Graph(false);
G->addNode("felipe");
G->addNode("rosa");
G->addNode("jose");
G->addNode("ricky");
G->addNode("vicky");
G->addEdge("felipe", "jose");
G->addEdge("felipe", "vicky");
G->addEdge("rosa", "ricky");
G->addEdge("ricky", "vicky");
auto result = G->bfs("felipe");
for (auto kv : result) {
cout << kv.first << "\t: " << kv.second << endl;
}
G->saveDot("amigos.dot");
delete G;
return 0;
}
# + id="Z2rLayvs_0Gh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 370} outputId="1d0823b6-eb62-4ad1-fbb6-fbb39bb7478d" magic_args="bash" language="script"
# g++ -std=c++17 graphTest.cpp \
# && ./a.out \
# && cat amigos.dot
# + id="6uWnqn50ACLl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 176} outputId="5052681b-2224-493a-cd6f-9b07e34483f4"
import graphviz
graphviz.Source.from_file("amigos.dot")
# + id="bm_v3PaRAhu7" colab_type="code" colab={}
| week14/14_graph_nice_adjacency_list.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.0
# language: julia
# name: julia-1.5
# ---
# ## Linear compartment models with constant input
#
# We consider three series of linear compartment models: cyclic, catenary, and mamillary. For each of the models, we replace each reaction rate $a_{ij}$ with $b_{ij} + c_{ij}x_0$, where $b_{ij}$ and $c_{ij}$ are scalar parameters, and $x_0$ is a constant input. We report the results and runtimes.
using BenchmarkTools
using DataFrames
include("../experiments_bounds.jl")
include("linear_compartment.jl");
function run_model(func, min_n, max_n)
"""
func is a function for generating a model:
cycle, catenary, mammilary
min_n and max_n define the range of sizes to test
"""
results = []
runtimes = []
for n in min_n:max_n
ode = linear_compartment_model(func(n), [1])
x0, x1 = gens(ode.poly_ring)[1:2]
time = @belapsed bound_number_experiments($ode, [$x0, $x1])
bnd = bound_number_experiments(ode, [x0, x1])
push!(results, bnd)
push!(runtimes, time)
end
return df = DataFrame(n = min_n:max_n, results = results, runtimes = runtimes)
end;
run_model(cycle, 3, 15)
run_model(catenary, 3, 15)
run_model(mammilary, 3, 15)
| examples/LinearCompartment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # EXPLORATORY DATA ANALYSIS
#
# In this Notebook, we will be exploring a dataset from Kaggle on Telco Customer churn (https://www.kaggle.com/blastchar/telco-customer-churn).
#
# # OBJECTIVES
#
# We will be training a Support Vector Classifier and a RandomForest Classifier algorithms for predicting which customer will churn based on the given dataset.
# #### Columns of the data
# - CustomerID = Customer ID
# - Gender Customer = gender (female, male)
# - SeniorCitizen = Whether the customer is a senior citizen or not (1, 0)
# - Partner = Whether the customer has a partner or not (Yes, No)
# - Dependents = Whether the customer has dependents or not (Yes, No)
# - Tenure = Number of months the customer has stayed with the company
# - PhoneService = Whether the customer has a phone service or not (Yes, No)
# - MultipleLines = Whether the customer has multiple lines or not (Yes, No, No phone service)
# - InternetService = Customer’s internet service provider (DSL, Fiber optic, No)
# - OnlineSecurity = Whether the customer has online security or not (Yes, No, No internet service)
# - OnlineBackup = Whether the customer has online backup or not (Yes, No, No internet service)
# - DeviceProtection = Whether the customer has device protection or not (Yes, No, No internet service)
# - TechSupport = Whether the customer has tech support or not (Yes, No, No internet service)
# - StreamingTV = Whether the customer has streaming TV or not (Yes, No, No internet service)
# - StreamingMovies = Whether the customer has streaming movies or not (Yes, No, No internet service)
# - Contract = The contract term of the customer (Month-to-month, One year, Two year)
# - PaperlessBilling = Whether the customer has paperless billing or not (Yes, No)
# - PaymentMethod = The customer’s payment method (Electronic check, Mailed check, Bank transfer (automatic), Credit card (automatic))
# - MonthlyCharges = The amount charged to the customer monthly
# - TotalCharges = The total amount charged to the customer
# - Churn = Whether the customer churned or not (Yes or No)
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import sklearn
import seaborn as sb
# %run future_encoders
# Loading the Datasets.
# Let's have a look at the first 5 and the last 5 rows of the dataset
df = pd.read_csv("Telco-Customer-Churn.csv")
df.head()
df.tail()
df.info()
# From the info, We are glad there is no missing data in any of the columns. Each columns has 7,043 objects in each.
# #### What else can we say about the data
df.describe()
# From the above, **SeniorCitizen** column is a categorical variable presented in a (0,1). **Tenure and MonthlyCharges** are the numerical variables in the dataset.
#
# Let's learn more about our dataset
df["Churn"].value_counts()
# From above, we can see that 1,869 customers churned out of the 7,043 in the dataset and this is our target variables.
# We will have to convert this to Hot encoders and represent the data in (1,0) for the customers that churned and the ones that didn.t.
df["MultipleLines"].value_counts()
df["Contract"].value_counts()
df["PaymentMethod"].value_counts()
df["tenure"].value_counts()
df["OnlineSecurity"].value_counts()
# We have 18 columns with categorical variables, and we will be building a transformation pipeline to turn it to OneHotEncoder for machine learning.
df.hist(bins=50, figsize=(20,15))
plt.show()
corr_matrix = df.corr()
corr_matrix["tenure"].sort_values(ascending=False)
# Importing Dataframes to handle categorical variable.
# +
from sklearn.base import BaseEstimator, TransformerMixin
# A class to select numerical or categorical columns
# since Scikit-Learn doesn't handle DataFrames yet
class DataFrameSelector(BaseEstimator, TransformerMixin):
def __init__(self, attribute_names):
self.attribute_names = attribute_names
def fit(self, X, y=None):
return self
def transform(self, X):
return X[self.attribute_names]
# -
# ## Data cleaning for Machine Learning Algorithms.
# - We will be converting our categorical values to OneHotEncoders for machine learning.
# - We will employ some data variables from Ageron's Hands on Machine learning'
# From Hands on Machine learning by Ageron
class MostFrequentImputer(BaseEstimator, TransformerMixin):
def fit(self, X, y=None):
self.most_frequent_ = pd.Series([X[c].value_counts().index[0] for c in X],
index=X.columns)
return self
def transform(self, X, y=None):
return X.fillna(self.most_frequent_)
# Before we proceed too far, let us split our data into test and train sets
from sklearn.model_selection import train_test_split
train_set, test_set = train_test_split(df, test_size=0.2, random_state=42)
# Building Pipelines for the categorical attributes and using OneHotEncoder to convert them to One-hot Vector
# +
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import Imputer
cat_pipeline = Pipeline([
("select_cat", DataFrameSelector(["gender", "SeniorCitizen", "Partner", "Dependents",
"PhoneService", "MultipleLines", "InternetService", "OnlineSecurity",
"OnlineBackup", "DeviceProtection", "TechSupport", "StreamingTV", "StreamingMovies",
"Contract", "PaperlessBilling", "PaymentMethod", "TotalCharges"])),
("imputer", MostFrequentImputer()),
("cat_encoder", OneHotEncoder(sparse=False))
])
# -
cat_pipeline.fit_transform(train_set)
# Building numerical attributes pipeline
# +
imputer = Imputer(strategy="median")
num_pipeline = Pipeline([
("select_numeric", DataFrameSelector(["tenure", "MonthlyCharges"])),
("imputer", Imputer(strategy="median"))
])
# -
num_pipeline.fit_transform(train_set)
# Joiniing the two pielines together
from sklearn.pipeline import FeatureUnion
preprocess_pipeline = FeatureUnion(transformer_list=[
("num_pipeline", num_pipeline),
("cat_pipeline", cat_pipeline),
])
# We have preprocessed our datasets using the pipelines
X_train = preprocess_pipeline.fit_transform(train_set)
X_train
# Gettiing our Labes
y_train = train_set["Churn"]
# TRAINING A CLASSIFIER MODEL
# +
from sklearn.svm import SVC
svm_clf = SVC()
svm_clf.fit(X_train, y_train)
# -
# Great Our SVC model hs been trained....let's see how accurate are it's predictions
# +
from sklearn.model_selection import cross_val_score
svm_scores = cross_val_score(svm_clf, X_train, y_train, cv=10)
svm_scores.mean()
# -
# We can also look at another model and see how well they will perform.
# +
from sklearn.ensemble import RandomForestClassifier
forest_clf = RandomForestClassifier(random_state=42)
forest_scores = cross_val_score(forest_clf, X_train, y_train, cv=10)
forest_scores.mean()
# -
corr_matrix = np.corrcoef(X_train)
#corr_matrix["churn"].sort_values(ascending=False)
# #### Conclusively, we can see that the SVC model did better in this dataset compared to the RandomForestClassifier. Don't forget that you can contribute to this Notebook by training this dataset with Neural nets and deep learning, would be most glad to welcome all you inputs.
| Predicting_customer_retentions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/deshanahan/DATA-602-Homework/blob/main/LogisticRegression_SVM_Classification.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="wJ283-QsmxHK"
import pandas as pd
import numpy as np
from sklearn.datasets import load_breast_cancer
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.svm import SVC
from sklearn.metrics import classification_report, confusion_matrix, roc_curve, auc
from sklearn.linear_model import LogisticRegressionCV
from matplotlib import pyplot as plt
# %matplotlib inline
# + colab={"base_uri": "https://localhost:8080/"} id="zZNb4CACp1MB" outputId="c87461a1-38fd-4c23-ea78-6afda10ad645"
breast_cancer = load_breast_cancer()
features = breast_cancer.feature_names
labels = breast_cancer.target_names
breast_cancer_data = breast_cancer.data
print(f'features: {features} \n')
print(f'labels: {labels} \n')
print(f'Shape: {breast_cancer_data.shape}')
# + colab={"base_uri": "https://localhost:8080/"} id="AcOdRnT5vU_w" outputId="2f9e2d77-1cf4-4bbb-faad-c8a5fd059556"
print(breast_cancer.DESCR)
# + colab={"base_uri": "https://localhost:8080/", "height": 249} id="3Ird5_bXukAZ" outputId="e612a126-a9c9-411a-d543-f7881fe2c64e"
breast_cancer_df = pd.DataFrame(breast_cancer_data)
breast_cancer_df.columns = features
breast_cancer_df.head(5)
# + colab={"base_uri": "https://localhost:8080/", "height": 455} id="gk_Ne_pywWZP" outputId="f201b75d-ddba-4e4b-ff81-d7a7cd868d00"
breast_cancer_df = breast_cancer_df.assign(target=pd.Series(breast_cancer.target))
breast_cancer_df
# + [markdown] id="NdLY10d7thiM"
# Since recall shows the classifier's ability to show find all positive samples, we can see that the support vector machine model correctly predicted 97% of the malignant breast masses and 98% of the benign breast masses.
# + id="QoUXnVb01v4b" colab={"base_uri": "https://localhost:8080/"} outputId="2befd409-2e89-4971-ab75-89d836f4b289"
X = breast_cancer_df.drop('target', axis = 1)
y = breast_cancer_df['target']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.3, random_state = 42)
svm = SVC()
scaling = StandardScaler().fit(X_train)
X_train = scaling.transform(X_train)
X_test = scaling.transform(X_test)
svm_train = svm.fit(X_train, y_train)
svm_predict = svm_train.predict(X_test)
print(f'Classification Report: \n\n {classification_report(y_test, svm_predict)}')
# + colab={"base_uri": "https://localhost:8080/"} id="GKsl8JEdxw0E" outputId="f9c3f730-fb4c-4da4-c347-1829b3f52850"
lr = LogisticRegressionCV()
lr_train = lr.fit(X_train, y_train)
lr_predict = lr_train.predict(X_test)
print(f'Classification Report: \n\n {classification_report(y_test, lr_predict)}\n')
print(f'\nConfusion Matrix: \n\n {confusion_matrix(y_test, lr_predict)}')
# + colab={"base_uri": "https://localhost:8080/", "height": 638} id="GeDP7gBawL6X" outputId="d79b17d6-ac32-43b8-b8f8-91d6ad8fc0da"
Y_scores = lr.decision_function(X_test)
fpr, tpr, thresholds = roc_curve(lr_predict, Y_scores)
plt.figure(figsize=(15, 15))
plt.plot(fpr, tpr, color='red', label="Logistic Regression (AUC: %.2f)"
% auc(fpr, tpr))
plt.plot([0, 1], [0, 1], color = 'blue', linestyle = '--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.01])
plt.title = ('ROC Curve')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.legend(loc = 'lower right')
plt.show()
| LogisticRegression_SVM_Classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6.9 64-bit
# name: python3
# ---
# +
import matplotlib.pyplot as plt
import numpy as np
import sys
import pprint
sys.path.append('..')
from facerec.facerec import *
pprint.pprint(sys.path)
# -
from PIL import Image
im = Image.open('../data/lena.jpg')
im
# +
def read_img(file: str) -> np.ndarray:
return np.array(Image.open(file))
img = read_img('../data/lena.jpg')
print(img.shape)
pil_img = Image.fromarray(img)
# pil_img.save('../data/temp/lena_save_pillow.jpg')
pil_img
# +
# def rgb2hsv(rgb: np.ndarray):
rgb = img
input_shape = rgb.shape
rgb = rgb.reshape(-1, 3)
r, g, b = rgb[:, 0], rgb[:, 1], rgb[:, 2]
maxc = np.maximum(np.maximum(r, g), b)
minc = np.minimum(np.minimum(r, g), b)
v = 100 * maxc // 255
deltac = maxc - minc
s = 100 * deltac // maxc
s[maxc == 0] = 0
h = np.empty_like(v)
deltac[deltac == 0] = 1 # ゼロ除算を避けるために代入。後で当該の要素は上書きされる。
h[maxc == r] = 60 * (g[maxc == r] - b[maxc == r]) / deltac[maxc == r]
h[maxc == g] = 60 * (b[maxc == g] - r[maxc == g]) / deltac[maxc == g] + 120
h[maxc == b] = 60 * (r[maxc == b] - g[maxc == b]) / deltac[maxc == b] + 240
h[minc == maxc] = 0
hsv_img = np.dstack([h, s, v]).reshape(input_shape)
hsv_img
# +
hsv = hsv_img
input_shape = hsv.shape
hsv = hsv.reshape(-1, 3)
h, s, v = hsv[:, 0], hsv[:, 1], hsv[:, 2]
maxc = np.round(255 * v / 100).astype(int)
minc = np.round(maxc - maxc * s / 100).astype(int)
i = h // 60
deltac = maxc - minc
maxc, minc = maxc.reshape(-1, 1), minc.reshape(-1, 1)
deltac, h = deltac.reshape(-1, 1), h.reshape(-1, 1)
x1 = deltac * np.abs(120 - h) // 60 + minc
x2 = deltac * np.abs(240 - h) // 60 + minc
rgb = np.zeros_like(hsv)
rgb[i == 0] = np.hstack([maxc, deltac * h // 60 + minc, minc])[i == 0]
rgb[i == 1] = np.hstack([x1, maxc, minc])[i == 1]
rgb[i == 2] = np.hstack([minc, maxc, x1])[i == 2]
rgb[i == 3] = np.hstack([minc, x2, maxc])[i == 3]
rgb[i == 4] = np.hstack([x2, minc, maxc])[i == 4]
rgb[(i == 5) | (i == 6)] = np.hstack([maxc, minc, deltac * (360 - h) // 60] + minc)[(i == 5) | (i == 6)]
rgb[s == 0] = np.hstack([maxc, maxc, maxc])[s == 0]
rgb.reshape(input_shape)
# +
h, w, _ = img.shape
# print(img.shape)
num_split = 2
split_hists = np.empty((num_split ** 2, 360)) # num_split^2行360列の行列を確保
count = 0
for h_count in range(num_split):
pos_h = h // num_split * h_count
for w_count in range(num_split):
pos_w = w // num_split * w_count
im_trim = trim(img, pos_w, pos_h, w // num_split, h // num_split)
print(pos_w, pos_h, w // num_split, h // num_split)
hist, _ = np.histogram(rgb2hsv(im_trim)[:, :, 0].ravel(), range=(0, 360), bins=360, density=True)
split_hists[count, :] = hist
count += 1
plt.imshow(im_trim)
plt.show()
print(split_hists.shape)
| notebooks/sandbox2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### <span style="color:green"> Exercise: Make a Function </span>
#
# Define a greeting function. It should take a name as input, and print a greeting.
# Then use the function to greet first Alice, and then Bob. The output should be:
#
# Hello Alice
# Hello Bob
#
# #### <span style="color:blue"> Solution: Make a Function </span>
# The important thing to note here is that is `name` is merely a placeholder (a variable).
#
# +
def greet(name):
print("Hello", name)
greet('Alice')
greet('Bob')
# -
# ### <span style="color:green"> Exercise: Returning Values </span>
#
# Define a function that returns the last element of a list.
# Then use the function on the list below:
#
# #### <span style="color:blue"> Solution: Returning Values </span>
# There are several ways to do this, but the simplest way to access the last element of a list is to index by -1.
# +
def get_last(sequence):
last = sequence[-1]
return last
items = ["first", "second", "third"]
print(get_last(items))
# -
# ### <span style="color:green"> Case Law Exercise: functions to get information </span>
#
# Here, we will continue working with our JSON data.
# Define a function that takes a case as an argument, and returns the name of the court.
#
# The code below applies this function to the cases in a loop. Make sure you use the function name that is called in the loop.
#
# You might need to browse the [data](https://api.case.law/v1/cases/?jurisdiction=ill&full_case=true&decision_date_min=2011-01-01&page_size=1").
#
# #### <span style="color:blue"> Solution: functions to get information </span>
#
# Remember that `case` is a nested dictionary. To find the court name, you first have to access `court` from the dictionary.
# +
import requests
import json
def get_court_name(case):
'''Returns the name of the court for this case'''
court = case["court"]
court_name = court["name"]
return court_name
URL = "https://api.case.law/v1/cases/?full_case=true&decision_date_min=2011-01-01&page_size=3"
data = requests.get(URL).json()
cases = data["results"]
for case in cases:
print(get_court_name(case))
# -
# ### <span style="color:green">Library Data Exercise: Function to get the Title </span>
#
# Here, we will continue working with our JSON data.
# Define a function that takes a book as an argument and returns the book title.
#
# The code below applies this function to the books in a loop. Make sure you use the function name that is called in the loop.
#
# You might need to browse the [data](https://api.nb.no/catalog/v1/items?digitalAccessibleOnly=true&size=3&filter=mediatype:bøker&q=Bing,Jon).
#
# #### <span style="color:blue">Solution: Function to get the Title </span>
#
# Remember that `book` is a nested dictionary. To find the title, you first have to access `metadata` from the dictionary.
# +
import requests
import json
def get_title(book):
'Returns the title of the given book'
metadata = book['metadata']
title = metadata['title']
return title
URL = "https://api.nb.no/catalog/v1/items?digitalAccessibleOnly=true&size=3&filter=mediatype:bøker&q=Bing,Jon"
data = requests.get(URL).json()
embedded = data['_embedded']
items = embedded['items']
for item in items:
print(get_title(item))
# -
# All done!
| solutions/solutions_06.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=["remove_cell"]
# # 複数量子ビットともつれ状態
# -
# 単一の量子ビットは興味深いものではありますが、これらは個別では計算的に優位性をもたらすものではありません。ここでは複数の量子ビットに関する表現方法と、これらの量子ビットがどのように相互作用することができるかについて見ていきます。これまでに、2次元ベクトルを使用して1つの量子ビットに関する状態を表現することができることを学びました。今回は複数量子ビットの状態についてどのように表現することができるかについて見ていくことにします。
# ## 目次
# 1. [複数量子ビット状態の表現](#represent)
# 1.1 [練習問題](#ex1)
# 2. [複数量子ビット状態ベクトル上の単一量子ビットゲート](#single-qubit-gates)
# 2.1 [練習問題](#ex2)
# 3. [複数量子ビットゲート](#multi-qubit-gates)
# 3.1 [CNOTゲート](#cnot)
# 3.2 [もつれ状態](#entangled)
# 3.3 [練習問題](#ex3)
#
#
# ## 1. 複数量子ビット状態の表現 <a id="represent"></a>
#
# 単一の量子ビットは2つの状態を持つことができ、その状態は2つの複素振幅を持つことを見てきました。同様に、2量子ビットは4つの状態を持ちます。
#
# `00` `01` `10` `11`
#
# さらに、2量子ビットの状態を説明するには4つの複素振幅が必要です。これらの振幅を以下のように4次元ベクトルで表します。
#
# $$ |a\rangle = a_{00}|00\rangle + a_{01}|01\rangle + a_{10}|10\rangle + a_{11}|11\rangle = \begin{bmatrix} a_{00} \\ a_{01} \\ a_{10} \\ a_{11} \end{bmatrix} $$
#
# 観測に関するルールについても単一量子ビットの時と同様に機能します。
#
# $$ p(|00\rangle) = |\langle 00 | a \rangle |^2 = |a_{00}|^2$$
#
# また、規格化条件などについても同様です。
#
# $$ |a_{00}|^2 + |a_{01}|^2 + |a_{10}|^2 + |a_{11}|^2 = 1$$
#
# 2つの分離された量子ビットがあった場合、テンソル積を使用することで2量子ビットの状態を説明することができます。
#
# $$ |a\rangle = \begin{bmatrix} a_0 \\ a_1 \end{bmatrix}, \quad |b\rangle = \begin{bmatrix} b_0 \\ b_1 \end{bmatrix} $$
#
# $$
# |ba\rangle = |b\rangle \otimes |a\rangle = \begin{bmatrix} b_0 \times \begin{bmatrix} a_0 \\ a_1 \end{bmatrix} \\ b_1 \times \begin{bmatrix} a_0 \\ a_1 \end{bmatrix} \end{bmatrix} = \begin{bmatrix} b_0 a_0 \\ b_0 a_1 \\ b_1 a_0 \\ b_1 a_1 \end{bmatrix}
# $$
#
# さらに、同じルールに則り、テンソル積を使用して任意の数の量子ビット状態について説明することができます。3量子ビットの場合の例は以下の通りです。
#
# $$
# |cba\rangle = \begin{bmatrix} c_0 b_0 a_0 \\ c_0 b_0 a_1 \\ c_0 b_1 a_0 \\ c_0 b_1 a_1 \\
# c_1 b_0 a_0 \\ c_1 b_0 a_1 \\ c_1 b_1 a_0 \\ c_1 b_1 a_1 \\
# \end{bmatrix}
# $$
#
# $n$個の量子ビットがある場合、$2^n$の複素振幅について記録する必要があります。これらのベクトルは量子ビットの数と共に指数関数的に大きくなります。これが多数の量子ビットを用いた量子コンピューターをシミュレートすることが大変困難である理由です。現代のラップトップは約20量子ビットのシミュレートは簡単にできますが、100量子ビットになると最も大きいスーパーコンピューターでも手に負えません。
#
# 回路の例を見てみましょう。
# + tags=["thebelab-init"]
from qiskit import *
from math import pi
import numpy as np
from qiskit.visualization import plot_bloch_multivector, plot_histogram
# -
qc = QuantumCircuit(3)
# Apply H-gate to each qubit:
for qubit in range(3):
qc.h(qubit)
# See the circuit:
qc.draw()
# 各量子ビットは$|+\rangle$の状態をとっており、ベクトルは以下のとおりとなります。
#
# $$
# |{+++}\rangle = \frac{1}{\sqrt{8}}\begin{bmatrix} 1 \\ 1 \\ 1 \\ 1 \\
# 1 \\ 1 \\ 1 \\ 1 \\
# \end{bmatrix}
# $$
# +
# Let's see the result
backend = Aer.get_backend('statevector_simulator')
final_state = execute(qc,backend).result().get_statevector()
# In Jupyter Notebooks we can display this nicely using Latex.
# If not using Jupyter Notebooks you may need to remove the
# array_to_latex function and use print(final_state) instead.
from qiskit_textbook.tools import array_to_latex
array_to_latex(final_state, pretext="\\text{Statevector} = ")
# -
# 期待した結果が得られたことが確認できます。
#
# ### 1.1 練習問題:<a id="ex1"></a>
#
# 1. 次の量子ビットのテンソル積の(状態)ベクトルを書け。
# a) $|0\rangle|1\rangle$
# b) $|0\rangle|+\rangle$
# c) $|+\rangle|1\rangle$
# d) $|-\rangle|+\rangle$
# 2. 状態を2つの別々の量子ビットとして記述せよ。
# $|\psi\rangle = \tfrac{1}{\sqrt{2}}|00\rangle + \tfrac{i}{\sqrt{2}}|01\rangle $
#
#
# ## 2. 複数量子ビット状態ベクトル上の単一量子ビットゲート<a id="single-qubit-gates"></a>
#
# Xゲートは以下の行列で表現されます。
#
# $$
# X = \begin{bmatrix} 0 & 1 \\ 1 & 0 \end{bmatrix}
# $$
#
# さらに、Xゲートは$|0\rangle$の状態に以下のように作用します。
#
# $$
# X|0\rangle = \begin{bmatrix} 0 & 1 \\ 1 & 0 \end{bmatrix}\begin{bmatrix} 1 \\ 0 \end{bmatrix} = \begin{bmatrix} 0 \\ 1\end{bmatrix}
# $$
#
# しかしながら、Xゲートが複数量子ビットベクトル内の量子ビットに対してどのような作用をするのかははっきりしません。幸運にもルールはかなり単純です。複数量子ビットの状態ベクトルを計算するのにテンソル積を使用したように、テンソル積を使用してこれらの状態ベクトルに作用する行列を計算することができます。例えば、以下の回路においては、
qc = QuantumCircuit(2)
qc.h(0)
qc.x(1)
qc.draw()
# HとXによる同時演算はそれらのテンソル積を用いて表現可能です。
#
# $$
# X|q_1\rangle \otimes H|q_0\rangle = (X\otimes H)|q_1 q_0\rangle
# $$
#
# 演算は以下の通りとなり、
#
# $$
# X\otimes H = \begin{bmatrix} 0 & 1 \\ 1 & 0 \end{bmatrix} \otimes \tfrac{1}{\sqrt{2}}\begin{bmatrix} 1 & 1 \\ 1 & -1 \end{bmatrix} = \frac{1}{\sqrt{2}}
# \begin{bmatrix} 0 \times \begin{bmatrix} 1 & 1 \\ 1 & -1 \end{bmatrix}
# & 1 \times \begin{bmatrix} 1 & 1 \\ 1 & -1 \end{bmatrix}
# \\
# 1 \times \begin{bmatrix} 1 & 1 \\ 1 & -1 \end{bmatrix}
# & 0 \times \begin{bmatrix} 1 & 1 \\ 1 & -1 \end{bmatrix}
# \end{bmatrix} = \frac{1}{\sqrt{2}}
# \begin{bmatrix} 0 & 0 & 1 & 1 \\
# 0 & 0 & 1 & -1 \\
# 1 & 1 & 0 & 0 \\
# 1 & -1 & 0 & 0 \\
# \end{bmatrix}
# $$
#
# 4次元状態ベクトル$|q_1 q_0\rangle$に適用することができます。表記がかなり煩雑になるため、以下のとおり簡略化した表記法がよく用いられます。
#
# $$
# X\otimes H =
# \begin{bmatrix} 0 & H \\
# H & 0\\
# \end{bmatrix}
# $$
#
# 手で計算する代わりにQiskitの`unitary_simulator`を使用すると計算をしてくれます。unitary simulatorは、回路にある全てのゲートを乗算して量子回路全体を表現する一つのユニタリー行列にしてくれます。
backend = Aer.get_backend('unitary_simulator')
unitary = execute(qc,backend).result().get_unitary()
# 結果を見てみます:
# In Jupyter Notebooks we can display this nicely using Latex.
# If not using Jupyter Notebooks you may need to remove the
# array_to_latex function and use print(unitary) instead.
from qiskit_textbook.tools import array_to_latex
array_to_latex(unitary, pretext="\\text{Circuit = }\n")
# 1量子ビットのみにゲートを適用する場合(以下の回路など)、単位行列を使用してテンソル積を実行します。例:
#
# $$ X \otimes I $$
qc = QuantumCircuit(2)
qc.x(1)
qc.draw()
# Simulate the unitary
backend = Aer.get_backend('unitary_simulator')
unitary = execute(qc,backend).result().get_unitary()
# Display the results:
array_to_latex(unitary, pretext="\\text{Circuit = } ")
# Qiskitが以下の通りテンソル積を計算したのが見て取れます。
# $$
# X \otimes I =
# \begin{bmatrix} 0 & I \\
# I & 0\\
# \end{bmatrix} =
# \begin{bmatrix} 0 & 0 & 1 & 0 \\
# 0 & 0 & 0 & 1 \\
# 1 & 0 & 0 & 0 \\
# 0 & 1 & 0 & 0 \\
# \end{bmatrix}
# $$
#
# ### 2.1 練習問題: <a id="ex2"></a>
#
# 1. 次の一連のゲートにより作成される単一量子ビットユニタリー ($U$) を計算せよ。$U = XZH$ 結果の確認にはQisikitのunitary simulatorを使用すること。
# 2. 上記の回路についてゲートを変更してみよ。テンソル積を計算し、unitary simulatorを用いて答え合わせをせよ。
#
# **Note:** 本、ソフトウェア、ウェブサイトによって量子ビットの順序が異なる。これは、同じ回路のテンソル積は全く異なるもののように見えることとなり、頭痛の種になることがある。他の情報源を参照する場合は注意すること。
# ## 3. 複数量子ビットゲート <a id="multi-qubit-gates"></a>
#
# 複数量子ビットの状態について表現方法を知ることができたため、複数量子ビットがそれぞれどのように相互作用するかについて学ぶ準備ができました。重要な2量子ビットゲートはCNOTゲートです。
#
# ### 3.1 CNOTゲート <a id="cnot"></a>
#
# このゲートは[計算の原子](../ch-states/atoms-computation.html)の章に出てきました。このゲートは条件付きゲートであり、1つ目の量子ビット(コントロール)が$|1\rangle$の場合に2つ目の量子ビット(ターゲット)にXゲートを適用する、というものです。このゲートは`q0`をコントロール、`q1`をターゲットとして以下のような回路として描画できます。
qc = QuantumCircuit(2)
# Apply CNOT
qc.cx(0,1)
# See the circuit:
qc.draw()
# 量子ビットが$|0\rangle$と$|1\rangle$の重ね合わせでない場合、このゲートはとても単純で直感的にわかりやすいです。古典の真理値表を使用できます。
#
# | Input (t,c) | Output (t,c) |
# |:-----------:|:------------:|
# | 00 | 00 |
# | 01 | 11 |
# | 10 | 10 |
# | 11 | 01 |
#
# 4次元状態ベクトルに従うと、どちらの量子ビットがコントロールでどちらがターゲットかにより、以下の2つの行列のいずれかになります。
#
# $$
# \text{CNOT} = \begin{bmatrix} 1 & 0 & 0 & 0 \\
# 0 & 0 & 0 & 1 \\
# 0 & 0 & 1 & 0 \\
# 0 & 1 & 0 & 0 \\
# \end{bmatrix}, \quad
# \text{CNOT} = \begin{bmatrix} 1 & 0 & 0 & 0 \\
# 0 & 1 & 0 & 0 \\
# 0 & 0 & 0 & 1 \\
# 0 & 0 & 1 & 0 \\
# \end{bmatrix}
# $$
#
# 本、シミュレーター、論文によって量子ビットの順番は異なります。今回のケースでは、左の行列が上記回路のCNOTに相当します。この行列は状態ベクトルの$|01\rangle$と$|11\rangle$の振幅を交換します。
#
# $$
# |a\rangle = \begin{bmatrix} a_{00} \\ a_{01} \\ a_{10} \\ a_{11} \end{bmatrix}, \quad \text{CNOT}|a\rangle = \begin{bmatrix} a_{00} \\ a_{11} \\ a_{10} \\ a_{01} \end{bmatrix} \begin{matrix} \\ \leftarrow \\ \\ \leftarrow \end{matrix}
# $$
#
# これまで古典的な状態に対する作用について見てきましたが、今度は重ね合わせ状態の量子ビットに対しての作用を見ていきましょう。1量子ビットを$|+\rangle$の状態にします。
qc = QuantumCircuit(2)
# Apply H-gate to the first:
qc.h(0)
qc.draw()
# Let's see the result:
backend = Aer.get_backend('statevector_simulator')
final_state = execute(qc,backend).result().get_statevector()
# Print the statevector neatly:
array_to_latex(final_state, pretext="\\text{Statevector = }")
# これは期待通りに$|0\rangle \otimes |{+}\rangle = |0{+}\rangle$の状態を作ります。
#
# $$
# |0{+}\rangle = \tfrac{1}{\sqrt{2}}(|00\rangle + |01\rangle)
# $$
#
# CNOTゲートを適用するとどうなるか見てみましょう。
qc = QuantumCircuit(2)
# Apply H-gate to the first:
qc.h(0)
# Apply a CNOT:
qc.cx(0,1)
qc.draw()
# Let's see the result:
backend = Aer.get_backend('statevector_simulator')
final_state = execute(qc,backend).result().get_statevector()
# Print the statevector neatly:
array_to_latex(final_state, pretext="\\text{Statevector = }")
# 以下の状態が得られます。
#
# $$
# \text{CNOT}|0{+}\rangle = \tfrac{1}{\sqrt{2}}(|00\rangle + |11\rangle)
# $$
#
# これはもつれ状態になっており、大変興味深いです。もつれ状態については次節に進みましょう。
# ### 3.2 もつれ状態 <a id="entangled"></a>
#
# 前節では以下の状態を作ることができることを見ました。:
#
# $$
# \tfrac{1}{\sqrt{2}}(|00\rangle + |11\rangle)
# $$
#
# これはベル状態として知られています。この状態は、50%の確率で$|00\rangle$の状態が観測され、50%の確率で$|11\rangle$の状態が観測されます。最も興味深いことに、$|01\rangle$または$|10\rangle$が観測される確率は**0%** です。このことは、Qiskitで確認することができます。
results = execute(qc,backend).result().get_counts()
plot_histogram(results)
# この結合された状態は、2つの別々の量子ビット状態として記述することはできません。このことには興味深い示唆があります。量子ビットが重ね合わせ状態にあるにも関わらず、1つの量子ビットを観測するともう1つの量子ビットの状態が得られると同時に重ね合わせ状態がなくなります。例として、一番上の量子ビットを観測して$|1\rangle$の状態が得られたとすると、2つの量子ビットの状態は以下のように変化します。
#
# $$
# \tfrac{1}{\sqrt{2}}(|00\rangle + |11\rangle) \quad \xrightarrow[]{\text{measure}} \quad |11\rangle
# $$
#
# 2つの量子ビットを数光年の距離に離したとしても、1つの量子ビットに関する観測はもう1つの量子ビットに対して影響を及ぼしているようにみえます。この[不気味な遠隔作用](https://en.wikipedia.org/wiki/Quantum_nonlocality) は20世紀初期に非常に多くの物理学者を悩ませました。
#
# 測定結果はランダムであり、一方の量子ビットの観測結果は、もう一方の量子ビットの操作の影響を受け**ない**ことに注意することが重要です。 このため、共量子もつれ状態を使用して通信をする方法は**ありません**。 これは、通信不可能定理[1]として知られています。
# ### 3.3 練習問題: <a id="ex3"></a>
# 1. 以下のベル状態を作り出す量子回路を作成せよ。 $\tfrac{1}{\sqrt{2}}(|01\rangle + |10\rangle)$.
# 状態ベクトルシミュレーターを使用して結果を検証せよ。
#
#
# 2. 設問1にて作成した回路は状態$|00\rangle$を$\tfrac{1}{\sqrt{2}}(|01\rangle + |10\rangle)$に変換するものである。この回路のユニタリー行列をQiskitシミュレータを使用して計算せよ。このユニタリー行列が正しい変換を行えることを検証せよ。
# ## 4. 参考文献
#
# [1] <NAME>, <NAME>, _Quantum Information and Relativity Theory,_ 2004, https://arxiv.org/abs/quant-ph/0212023
import qiskit
qiskit.__qiskit_version__
| i18n/locales/ja/ch-gates/multiple-qubits-entangled-states.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# ## What is this file?
# This file is a storehouse for our work that did not make it into our final submitted model. We approached the Random Acts of Pizza challenge from many angles and you'll find several of those approaches below. We hope it will provide additional context around how we thought through the challenge and some of the things we learned through this process.
#
# -------------------------------------------------------
# +
# For figures to show inline
# %matplotlib inline
## Import Libraries ##
import json
from pprint import pprint
from pandas.io.json import json_normalize
import pandas as pd
# General libraries.
import re
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
import mlxtend
import scipy
import datetime as dt
from itertools import product
# SK-learn library for importing the newsgroup data.
from sklearn.datasets import fetch_20newsgroups
# SK-learn libraries for feature extraction from text.
from sklearn.feature_extraction.text import *
# SK-learn libraries for pre/processing data
from sklearn import preprocessing
# NLTK for text processing, analyzing tools
from nltk.stem.porter import PorterStemmer
from nltk.stem import WordNetLemmatizer
from nltk.sentiment.util import *
from sklearn.decomposition import TruncatedSVD
from sklearn.decomposition import PCA
# SK-lear library for feature selection
from sklearn.feature_selection import VarianceThreshold
from sklearn.feature_selection import SelectKBest
from sklearn.feature_selection import chi2
from sklearn.feature_selection import SelectFromModel
from sklearn.feature_selection import mutual_info_classif
from sklearn.feature_selection import SelectPercentile
from vaderSentiment.vaderSentiment import SentimentIntensityAnalyzer
# SK-learn libraries for learning
from sklearn.pipeline import Pipeline
from sklearn.neighbors import KNeighborsClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.naive_bayes import BernoulliNB
from sklearn.svm import SVC
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import AdaBoostClassifier
from mlxtend.classifier import EnsembleVoteClassifier
# SK-learn libraries for evaluation
from sklearn import metrics
from sklearn.metrics import confusion_matrix
from sklearn.metrics import classification_report
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import roc_curve, roc_auc_score, recall_score
# +
## Get Data ##
# Reference for data: https://www.kaggle.com/c/random-acts-of-pizza/data
# Pull in the training and test data
with open('data/train.json', encoding='utf-8') as data_file:
trainData = json.loads(data_file.read())
with open('data/test.json', encoding='utf-8') as data_file:
testData = json.loads(data_file.read())
# create a dev data set
devData = trainData[0:1000]
trainData = trainData[1000:]
# show how the data looks in its original format
#pprint("data in json format:")
#pprint(trainData[1])
# create a normalized view
allTData = json_normalize(trainData)
print("\nSize of the normalized Data:", allTData.shape)
print("\nnormalized data columns:", list(allTData))
allDData = json_normalize(devData)
# -
# ### Section 1: Setting Up & Processing Data
# +
## Create subsets of data for analysis ###
# Create a flat dataset without the subreddits list
flatData = allTData.drop('requester_subreddits_at_request', 1)
# Create a separate dataset with just subreddits, indexed on request id
# We can creata a count vector on the words, run Naive Bayes against it,
# and add the probabilities to our flat dataset
subredTData = allTData[['request_id','requester_subreddits_at_request']]
subredTData.set_index('request_id', inplace=True)
subredDData= allDData[['request_id','requester_subreddits_at_request']]
subredDData.set_index('request_id', inplace=True)
# our training labels
trainLabel = allTData['requester_received_pizza']
devLabel = allDData['requester_received_pizza']
# What do these look like?
#print(list(flatData))
print(subredTData.shape)
#print(subredTData['requester_subreddits_at_request'][1])
# Create a corpus of subreddits to vectorize
trainCorpus = []
rTCorpus = []
rDCorpus = []
for index in range(len(subredTData)):
trainCorpus.append(' '.join(subredTData['requester_subreddits_at_request'][index]))
rTCorpus.append(' '.join(subredTData['requester_subreddits_at_request'][index]))
devCorpus = []
for index in range(len(subredDData)):
devCorpus.append(' '.join(subredDData['requester_subreddits_at_request'][index]))
rDCorpus.append(' '.join(subredDData['requester_subreddits_at_request'][index]))
# Baseline infofrom mlxtend.plotting import plot_decision_regions
print("\nPercent of people who got pizza:", round(sum(trainLabel)/len(trainLabel),3))
plt.figure(1,figsize=(10,4))
plt.subplot(121)
plt.hist(allTData['requester_received_pizza'])
plt.title("Distribtution of pizza's received in training data")
plt.subplot(122)
plt.hist(allDData['requester_received_pizza'])
plt.title("Distribtution of pizza's received in dev data")
# +
# Useful functions for analysis
def roc_curve1(y_true, y_pred_prob):
"""This function plots the ROC curve
Inputs: y_true, correct label
y_pred_prob, predicted probabilities
"""
fpr, tpr, thr = roc_curve(y_true, y_pred_prob)
plt.figure()
plt.plot(fpr,tpr)
plt.xlabel("False positive rate")
plt.ylabel("True positive rate")
plt.title("ROC Curve")
plt.show()
def score_rep(y_true, y_pred, desc):
"""Function to print out comprehensive report for classification test
Inputs: y_true, correct label
y_pred, predicted label from model
desc, description of model
Output: classification report
"""
print(desc)
print("-"*75)
print("Accuracy: ", metrics.accuracy_score(y_true, y_pred))
print("Area under curve of ROC: ", metrics.roc_auc_score(y_true, y_pred))
print("Classification report:\n")
print(metrics.classification_report(y_true, y_pred))
print("-"*75)
# -
# ### vaderSentiment Analysis
# +
# Quick learning exercise to figure out how
# to get vaderSentiment to work
from vaderSentiment.vaderSentiment import SentimentIntensityAnalyzer
analyser = SentimentIntensityAnalyzer()
test = "It was one of the worst movies I've seen, despite good reviews."
def print_sentiment_scores(sentence):
snt = analyser.polarity_scores(sentence)
print("{:-<40} {}".format(sentence, str(snt)))
print(snt['compound'])
print_sentiment_scores("It was one of the worst movies I've seen, despite good reviews.")
#sentences = "VADER is smart, handsome, and funny."
#print_sentiment_scores(sentences)
sentences = ["VADER is smart, handsome, and funny.", "VADER is silly, ugly, and rude!"]
for sentence in sentences:
print("\n")
print(sentence)
vs = print_sentiment_scores(sentence)
# +
# Setting up for Titles
title = allTData[['request_title', 'requester_received_pizza']].copy()
title.columns = ['Title', 'Got Pizza']
print("\n")
print("Titles and Pizza Success\n")
print(title.head(10))
pizza_title = title.groupby(['Got Pizza'])
print("\n")
# +
#Train Data Sentiment Analysis
import pandas as pd
titles = allTData['request_title']
df = pd.DataFrame(data = allTData)
#combing request title and Y variable
df = df[['request_title', 'requester_received_pizza']]
scores = []
for title in titles:
scores.append(analyser.polarity_scores(title)['compound'])
df["Vader Scores"] = scores
df = df.drop('request_title', axis = 1)
# print the y variable and sentiment score
print(df)
# +
#Dev Data Sentiment Analysis
import pandas as pd
titles = allDData['request_title']
df_d = pd.DataFrame(data = allDData)
df_d = df_d[['request_title', 'requester_received_pizza']]
scores_d = []
for title in titles:
scores_d.append(analyser.polarity_scores(title)['compound'])
df_d["Vader Scores"] = scores_d
df_d = df_d.drop('request_title', axis = 1)
print(df_d)
# +
# sentiment logistic regression accuracy
import pandas as pd
tTitles = allTData['request_title']
dTitles = allDData['request_title']
titleTSentiment = []
titleDSentiment = []
for title in tTitles:
snt = analyser.polarity_scores(title)
compoundScore = snt['compound']
titleTSentiment.append(compoundScore)
titleTSentiment = pd.DataFrame(titleTSentiment)
for title in dTitles:
snt = analyser.polarity_scores(title)
compoundScore = snt['compound']
titleDSentiment.append(compoundScore)
titleDSentiment = pd.DataFrame(titleDSentiment)
C = 100
modelLogit = LogisticRegression(penalty = 'l2', C = C)
trainLabel = allTData['requester_received_pizza']
devLabel = allDData['requester_received_pizza']
modelLogit.fit(titleTSentiment,trainLabel)
score_rep(devLabel,modelLogit.predict(titleDSentiment),'Logistic Regression, C = 0.001')
# -
# # Reducing vocabulary (doesn't work)
#
# +
# Keep this random seed here to make comparison easier.
np.random.seed(101)
### STUDENT START ###
# Countvectorizer options: turns on lower case, strip accents, and stop-words
vectorizer = CountVectorizer(min_df=2, max_df=0.95, lowercase=True, stop_words='english',
strip_accents='unicode', ngram_range=(1,4))
# Simple Pre-Processing Function
def data_preprocessor(s):
"""
Note: this function pre-processors data:
(1) removes non-alpha characters
(2) converts digits to 'number'
(3) regularizes spaces (although CountVectorizer ignores this unless they are part of words)
(4) reduces word size to n
"""
s = [re.sub(r'[?|$|.|!|@|\n|(|)|<|>|_|-|,|\']',r' ',s) for s in s] # strip out non-alpha numeric char, replace with space
s = [re.sub(r'\d+',r'number ',s) for s in s] # convert digits to number
s = [re.sub(r' +',r' ',s) for s in s] # convert multiple spaces to single space
# This sets word size to n=8
num = 8
def size_word(s):
temp = []
for s in s:
x = s.split()
z = [elem[:num] for elem in x]
z = ' '.join(z)
temp.append(z)
return temp
# Using NLTK 3.0
#stemmer = PorterStemmer()
lemmanizer = WordNetLemmatizer()
def set_word(s):
temp = []
for s in s:
#x = stemmer.stem(s)
z = lemmanizer.lemmatize(s,pos='v')
z = ''.join(z)
temp.append(z)
return temp
s = size_word(s)
s = set_word(s)
return s
def LR1(C):
"""
Function estimates an LR with l1 regularization and counts number of nonzero weights
Returns coefficient array
"""
# Preprocess data
train_data_v = vectorizer.fit_transform(data_preprocessor(trainCorpus))
dev_data_v = vectorizer.transform(data_preprocessor(devCorpus))
# Run the LR regression, l1 regularization, solving using liblinear [note: l1 doesn't work with multinomial]
clf = LogisticRegression(penalty='l1', C=C)
clf.fit(train_data_v,trainLabel)
test_predicted_labels = clf.predict(dev_data_v)
print ('\nLogistic Regression f1-score with C = %6.3f:' %C )
print (metrics.f1_score(devLabel,test_predicted_labels))
print ('Number of non-zero elements: %d' %(np.count_nonzero(clf.coef_)))
return clf.coef_
def LR2(C,lvocab):
"""
Calls LR with l2 for given vocab
Returns vocab size and accuracy
"""
# Define new vectorizer with vocab = vocab
vectorizer1 = CountVectorizer(lowercase=True, strip_accents='unicode',
stop_words='english',vocabulary=lvocab)
# Preprocess data using new vectorizer
train_data_v1 = vectorizer1.fit_transform(data_preprocessor(trainCorpus))
dev_data_v1 = vectorizer1.transform(data_preprocessor(devCorpus))
# Run the LR regression, l2 regularization, solving using sag
clf1 = LogisticRegression(penalty='l2', tol=0.01, C=C)
clf1.fit(train_data_v1,trainLabel)
test_predicted_labels = clf1.predict(dev_data_v1)
print ('\nLogistic Regression (using l2) f1-score with C = %6.3f:' %C )
print (metrics.f1_score(devLabel,test_predicted_labels))
score_rep(devLabel,test_predicted_labels,'Logistic Regression')
return (len(lvocab),metrics.f1_score(devLabel,test_predicted_labels) )
def create_vocab_list(s):
"""
inputs - clf.coef_
output - list of vocabulary
creates a list of vocabulary corresponding to non-zero features
"""
def build_vocab (s):
temp = []
for i in range (len(s)):
temp.append(s[i])
return temp
def build_vocab_list(s):
temp = []
for i in range(1):
y = np.nonzero(s[i])
y = list(y[0])
temp = temp + build_vocab(y)
temp = list(set(temp))
return temp
vocab = build_vocab_list(s)
x = vectorizer.get_feature_names()
temp = []
for vocab in vocab:
temp.append(x[vocab])
return temp
# Main program
C = [1e-1, 1] #2, 5, 10, 20, 50, 100, 200, 500, 1000 ] # Run over various C
a, b = [], []
for C in C:
z = LR1(C) # Call this function to estimate LR with L1, z is the matrix of coef
lvocab = create_vocab_list(z) # Call this function to create vocab list where coef not equal zero
print ('Vocabulary size: %d' %len(lvocab))
x, y = LR2(C,lvocab) # Call new LR estimate with L2
a.append(x)
b.append(y)
# Plot vocabulary size vs accuracy
plt.plot(a,b)
plt.xlabel('Vocabulary')
plt.ylabel('Accuracy (F1 score)')
plt.show()
# -
# # Using l1 to choose features (doesn't work)
# +
# get the best regularization
regStrength = [0.0001, 0.001, 0.01, 0.1, 0.5, 1.0, 2.0, 6.0, 10.0]
vectorizer_p = TfidfVectorizer(min_df=2, max_df=0.95, lowercase=True, stop_words='english',
strip_accents='unicode', ngram_range=(1,2))
tVector_p = vectorizer_p.fit_transform(data_preprocessor(trainCorpus))
dVector_p = vectorizer_p.transform(data_preprocessor(devCorpus))
for c in regStrength:
modelLogit = LogisticRegression(penalty='l1', C=c)
modelLogit.fit(tVector_p, trainLabel)
logitScore = round(modelLogit.score(dVector_p, devLabel), 4)
print("For C = ", c, "Logistic regression accuracy:", logitScore)
score_rep(devLabel,modelLogit.predict(dVector_p),'Logistic Regression, C = 0.01')
# although the best score comes from c=.001, the bet F1-score
# comes from c=.5, and this gives better weight options
modelLogit = LogisticRegression(penalty='l1', C=.5, tol = .1)
modelLogit.fit(tVector_p, trainLabel)
score_rep(devLabel,modelLogit.predict(dVector_p),'Logistic Regression')
roc_curve1(devLabel, modelLogit.predict_proba(dVector_p)[:,0])
print(max(modelLogit.coef_[0]))
numWeights = 5
sortIndex = np.argsort(modelLogit.coef_)
iLen = len(sortIndex[0])
print("\nTop", numWeights, "Weighted Features:")
for index in range((iLen - numWeights) , iLen):
lookup = sortIndex[0][index]
print(lookup)
weight = modelLogit.coef_[0][lookup]
print(vectorizer.get_feature_names()[sortIndex[0][index]], weight)
# -
# # PCA/LDA to reduce dimension
#
# +
pca = TruncatedSVD(n_components=600)
vectorizer = CountVectorizer(min_df=2, max_df=0.95, lowercase=True, stop_words='english',
strip_accents='unicode', ngram_range=(1,1))
tVector = vectorizer.fit_transform(data_preprocessor(trainCorpus))
dVector = vectorizer.transform(data_preprocessor(devCorpus))
#print(tVector.shape)
tVector_s = pca.fit(tVector)
dVector_s = pca.fit(dVector)
plt.figure(1, figsize=(4, 3))
plt.clf()
plt.axes([.2, .2, .7, .7])
plt.plot(np.cumsum(pca.explained_variance_ratio_), linewidth=2)
plt.axis('tight')
plt.xlabel('n_components')
plt.ylabel('explained_variance_')
plt.show()
RF = RandomForestClassifier(n_estimators=600)
RF.fit(tVector, trainLabel)
score_rep(devLabel, RF.predict(dVector),'Random Forest')
roc_curve1(devLabel, RF.predict_proba(dVector)[:,0])
alpha = 0.01
clf = BernoulliNB(alpha=alpha)
clf.fit(tVector, trainLabel)
score_rep(devLabel, clf.predict(dVector),'Naive Bayes, alpha = 0.01')
roc_curve1(devLabel, clf.predict_proba(dVector)[:,0])
C = 100 #(For now)
modelLogit = LogisticRegression(penalty='l2', C=C)
modelLogit.fit(tVector,trainLabel)
score_rep(devLabel,modelLogit.predict(dVector),'Logistic Regression, C = 0.01')
roc_curve1(devLabel, modelLogit.predict_proba(dVector)[:,0])
# -
| Learning_Notebook_Compiled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip3 install numpy
import numpy as np
l1 = [1, 2, 3]
x = np.array(l1, np.int16)
x = np.array(l1, dtype = np.int16)
x
type(x)
x[0]
x[2]
x[1]
x[-1]
x[4]
x1 = np.array([[1, 2, 3],
[4, 5, 6]],
np.int8)
x1
x1[0, 0]
x1[0, 1]
x1[0, 2]
x1[:, 0]
x1[:, 1]
x1[:, 2]
x1[0, :]
x1[1, :]
x1[2, :]
x1 = np.array([[[1, 2, 3], [4, 5, 6]],
[[0, -1, -2], [-3, -4, -5]]],
np.int8)
x1
x1 [ 0, 0, 0]
x1 [ -1, 0, 0]
x1 [ -1, -1, -1]
# # Properties of Ndarrays
x2 = np.array([[[1, 2, 3], [4, 5, 6]],
[[0, -1, -2], [-3, -4, -5]],
[[0, 0, 1], [-1, -3, 4]]],
np.int16)
x2
x2.ndim
x2.shape
x2.dtype
x2.size
x2.nbytes
x2.T
# # NumPy constants
np.inf
np.NAN
np.NINF
np.NZERO
np.PZERO
np.e
np.pi
np.euler_gamma
# # Slicing
a1 = np.array([1, 2, 3, 4, 5, 6, 7])
a1.dtype
a1.nbytes
a1[1:5]
a1[3:]
a1[:3]
a1[-4:-1]
a1[1:6:2]
| Section03/01_Getting_Started.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.3
# language: julia
# name: julia-1.5
# ---
include("../load.jl")
using Plots
using LaTeXStrings
using JuMP
# Creating the GlobalModel
m = JuMP.Model(with_optimizer(SOLVER_SILENT))
# @variable(m, -20 <= x[1:2] <= 20)
# set_lower_bound(x[2], 0)
# set_upper_bound(x[2], 10)
@variable(m, 1 <= x[1:2] <= 5)
@variable(m, y)
gm = GlobalModel(model = m)
# add_nonlinear_constraint(gm, :(x -> x[1]^2 + x[2]^2), dependent_var = y)
# add_nonlinear_constraint(gm, :(x -> log(x[1]^2 + x[2]^2)), dependent_var = y)
# add_nonlinear_constraint(gm, :(x -> exp(- 1/6*(x[1]^2) - 1/10*(x[2]^2))), dependent_var = y)
# add_nonlinear_constraint(gm, :(x -> x[1] + x[2]), dependent_var = y)
# add_nonlinear_constraint(gm, :(x -> x[1]^4*x[2]^2 + x[1]^2*x[2]^4 - 3*x[1]^2*x[2]^2 + 1), dependent_var = y)
add_nonlinear_constraint(gm, :(x -> max(0, x[1] - 2 - 0.1*x[1]*x[2]^2, x[2] - 2)), dependent_var = y)
bbr = gm.bbls[end]
set_param(bbr, :n_samples, 500)
uniform_sample_and_eval!(gm, lh_iterations=20)
classify_curvature(bbr)
# Curvature plot
bbr = gm.bbls[1]
plt3d = Plots.plot()
for val in [-1, 0, 1]
idxs = findall(i -> i == val, bbr.curvatures)
Plots.scatter!(bbr.X[idxs,"x[2]"], bbr.X[idxs,"x[1]"], bbr.Y[idxs], markersize = 2, label = val, camera = (10,50))
end
xlabel!(L"$x_2$"); ylabel!(L"$x_1$"); title!(L"$y$")
display(plt3d)
# We can train trees over the curvatures
lnr = base_classifier()
kwargs = Dict()
nl = OCTHaGOn.learn_from_data!(bbr.X, bbr.curvatures .> 0, lnr; fit_classifier_kwargs(; kwargs...)...)
# Plotting convexity
preds = IAI.predict(nl, bbr.X);
one_preds = findall(x -> x == 1, preds)
zero_preds = findall(x -> x == 0, preds)
Plots.scatter(bbr.X[one_preds,"x[2]"], bbr.X[one_preds,"x[1]"], bbr.Y[one_preds],
markersize = 2, seriescolor=:blue, camera = (10,50))
Plots.scatter!(bbr.X[zero_preds,"x[2]"], bbr.X[zero_preds,"x[1]"], bbr.Y[zero_preds],
markersize = 2, seriescolor=:red, camera = (10,50))
# How about univariate example?
m = JuMP.Model()
@variable(m, -5 <= x <= 10)
@variable(m, y)
gm = GlobalModel(model = m)
# add_nonlinear_constraint(gm, :(x -> max.(-6*x .-6,0.5x,0.2x.^5+0.5x)), dependent_var = y)
# add_nonlinear_constraint(gm, :(x -> x^1.2*sin(x)), dependent_var = y)
add_nonlinear_constraint(gm, :(x -> min((x-1)^2, (x+2)^2)), dependent_var = y)
uniform_sample_and_eval!(gm)
classify_curvature(gm.bbls[1])
bbr = gm.bbls[1]
plt2d = Plots.plot()
for val in [-1, 0, 1]
idxs = findall(i -> i == val, bbr.curvatures)
Plots.scatter!(bbr.X[idxs,"x"], bbr.Y[idxs], markersize = 4, label = val)
end
display(plt2d)
learn_constraint!(bbr)
bbr.learners[end]
| test/demos/curvature_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Qt DesignerによるGUI制御
# Qt DesignerでGUIを作成し、GUIへの入力をコード中で取得する方法について解説します。
# ## GUIの作成
# Qt DesignerはAnacondaに含まれ、コマンドプロンプト上で`designer`とタイプすると起動できます。中央に現れるダイアログでMain Windowを選択して作成ボタンをクリック。
# 
# 左側のWidgetが並んでいる所からPush ButtonとLine Editをドラッグして次のような簡単なGUIを作成します。
#
# 
#
#
# ドラッグしたオブジェクトは上の画像の赤矢印で示したobjectNameで管理されます。
# Ctrl+sでファイルをセーブします。ここではこのnotebookが置かれているフォルダに'`sample.ui`'という名前で保存されたとします。
# ## pythonコードにコンパイル
# 以下のようにして'`sample.ui`'を'`sample.py`'という名前のpythonファイルに変換します。
from PyQt5 import uic
with open('sample.ui','r') as source:
with open('sample.py','w') as compiled:
uic.compileUi(source,compiled)
# フォルダ内に`sample.py`というファイルが出来たはずです。
# ## GUI中の入力ボックスの値を取得
# ### GUIのインスタンス化、表示
# まずはGUIを表示してみます。
# +
import sample #コンパイルによってできたpythonファイルをimport
#jupyter上でpyqt5を使うためのマジックコマンド 普通は要らない
# %gui qt
from PyQt5.QtWidgets import QMainWindow,QApplication
app=QApplication([]) #ウィジットのイベントループを起動 これによってウィジットに触ったりできるようになる
mw=QMainWindow() #メインウィンドウを生成
#メインウィンドウをQt Designerで作ったものと同一に整える役目のオブジェクト
#インスタンス変数として作成したWidgetを保持する役目も担う
ui=sample.Ui_MainWindow()
ui.setupUi(mw) #uiにメインウィンドウを渡して整えてもらう
mw.show() #メインウィンドウを表示
# -
# ### 値の取得
# uiがインスタンス変数としてウィンドウを保持していて入力ボックスはlineEditという名前がついている。
# よってそのメソッドを呼び出すことで入力ボックスに入力した文字列を取得することができる。
# 入力ボックスに何か入力して次のステートメントを実行して下さい。
ui.lineEdit.text()
| QtDisgner.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pickle
import pandas as pd
# +
file_name = 'C:\\Users\\vgoncalvese\\Desktop\\Python\\Estudos_Zika\\zika-data\\data\\Unificado\\IBGE_Mun'
infile = open(file_name,'rb')
new_dict = pickle.load(infile)
infile.close()
#dados do IBGE data\\IBGE
abre_IBGE_MunTot = new_dict['IBGE_Mun']
# +
#resumo por estado estado parsed\\brazil
abre_dados_estadosTot = pickle.load( open( "C:\\Users\\vgoncalvese\\Desktop\\Python\\Estudos_Zika\\zika-data\\data\\Unificado\\dados_estadosTot.p", "rb" ) )
abre_dados_resumoTot = pickle.load( open( "C:\\Users\\vgoncalvese\\Desktop\\Python\\Estudos_Zika\\zika-data\\data\\Unificado\\dados_resumoTot.p", "rb" ) )
# -
#dados CDC data\\parsed\\cdc
abre_dadosCDCTot = pickle.load( open("C:\\Users\\vgoncalvese\\Desktop\\Python\\Estudos_Zika\\zika-data\\data\\Unificado\\dadosCDCTot.p", "rb" ))
#dados da Colombia data\\parsed\\colombia
abre_dados_regionTot = pickle.load( open( "C:\\Users\\vgoncalvese\\Desktop\\Python\\Estudos_Zika\\zika-data\\data\\Unificado\\dados_regionTot.p", "rb" ) )
abre_dados_departmentTot = pickle.load( open( "C:\\Users\\vgoncalvese\\Desktop\\Python\\Estudos_Zika\\zika-data\\data\\Unificado\\dados_departmentTot.p", "rb" ) )
abre_dados_sivigila_codeTot = pickle.load( open( "C:\\Users\\vgoncalvese\\Desktop\\Python\\Estudos_Zika\\zika-data\\data\\Unificado\\dados_sivigila_codeTot.p", "rb" ) )
#dados ECDE data\\parsed\\ecdc
abre_dadosECDETot = pickle.load( open( "C:\\Users\\vgoncalvese\\Desktop\\Python\\Estudos_Zika\\zika-data\\data\\Unificado\\dadosECDETot.p", "rb" ) )
#abre paho WHO data\\raw\\paho-who
abre_dados_pahoWhoTot = pickle.load( open("C:\\Users\\vgoncalvese\\Desktop\\Python\\Estudos_Zika\\zika-data\\data\\Unificado\\dados_pahoWhoTot.p", "rb" ))
abre_dadosTSVTot = pickle.load( open("C:\\Users\\vgoncalvese\\Desktop\\Python\\Estudos_Zika\\zika-data\\data\\Unificado\\dadosTSVTot.p", "rb" ))
abre_dados_pahoWhoTot
| data/Unificado/Bases_Unificado.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # 2D Model Inference on a 3D Volume
# Usecase: A 2D Model, such as, a 2D segmentation U-Net operates on 2D input which can be slices from a 3D volume (for example, a CT scan).
#
# After editing sliding window inferer as described in this tutorial, it can handle the entire flow as shown:
# 
#
# The input is a *3D Volume*, a *2D model* and the output is a *3D volume* with 2D slice predictions aggregated.
#
#
# [](https://colab.research.google.com/github/Project-MONAI/tutorials/blob/main/modules/2d_inference_3d_volume.ipynb)
#
# Install monai
# !python -c "import monai" || pip install -q "monai-weekly"
# Import libs
from monai.inferers import SlidingWindowInferer
import torch
from typing import Callable, Any
from monai.networks.nets import UNet
# ## Overiding SlidingWindowInferer
# The simplest way to achieve this functionality is to create a class `YourSlidingWindowInferer` that inherits from `SlidingWindowInferer` in `monai.inferers`
class YourSlidingWindowInferer(SlidingWindowInferer):
def __init__(self, spatial_dim: int = 0, *args, **kwargs):
# Set dim to slice the volume across, for example, `0` could slide over axial slices,
# `1` over coronal slices
# and `2` over sagittal slices.
self.spatial_dim = spatial_dim
super().__init__(*args, **kwargs)
def __call__(
self,
inputs: torch.Tensor,
network: Callable[..., torch.Tensor],
slice_axis: int = 0,
*args: Any,
**kwargs: Any,
) -> torch.Tensor:
assert (
self.spatial_dim < 3
), "`spatial_dim` can only be `[D, H, W]` with `0, 1, 2` respectively"
# Check if roi size (eg. 2D roi) and input volume sizes (3D input) mismatch
if len(self.roi_size) != len(inputs.shape[2:]):
# If they mismatch and roi_size is 2D add another dimension to roi size
if len(self.roi_size) == 2:
self.roi_size = list(self.roi_size)
self.roi_size.insert(self.spatial_dim, 1)
else:
raise RuntimeError(
"Currently, only 2D `roi_size` is supported, cannot broadcast to volume. "
)
return super().__call__(inputs, lambda x: self.network_wrapper(network, x))
def network_wrapper(self, network, x, *args, **kwargs):
"""
Wrapper handles cases where inference needs to be done using
2D models over 3D volume inputs.
"""
# If depth dim is 1 in [D, H, W] roi size, then the input is 2D and needs
# be handled accordingly
if self.roi_size[self.spatial_dim] == 1:
# Pass 4D input [N, C, H, W]/[N, C, D, W]/[N, C, D, H] to the model as it is 2D.
x = x.squeeze(dim=self.spatial_dim + 2)
out = network(x, *args, **kwargs)
# Unsqueeze the network output so it is [N, C, D, H, W] as expected by
# the default SlidingWindowInferer class
return out.unsqueeze(dim=self.spatial_dim + 2)
else:
return network(x, *args, **kwargs)
# ## Testing added functionality
# Let's use the `YourSlidingWindowInferer` in a dummy example to execute the workflow described above.
# +
# Create a 2D UNet with randomly initialized weights for testing purposes
# 3 layer network with down/upsampling by a factor of 2 at each layer with 2-convolution residual units
net = UNet(
spatial_dims=2,
in_channels=1,
out_channels=1,
channels=(4, 8, 16),
strides=(2, 2),
num_res_units=2,
)
# Initialize a dummy 3D tensor volume with shape (N,C,D,H,W)
input_volume = torch.ones(1, 1, 64, 256, 256)
# Create an instance of YourSlidingWindowInferer with roi_size as the 256x256 (HxW) and sliding over D axis
axial_inferer = YourSlidingWindowInferer(roi_size=(256, 256), sw_batch_size=1, cval=-1)
output = axial_inferer(input_volume, net)
# Output is a 3D volume with 2D slices aggregated
print("Axial Inferer Output Shape: ", output.shape)
# Create an instance of YourSlidingWindowInferer with roi_size as the 64x256 (DxW) and sliding over H axis
coronal_inferer = YourSlidingWindowInferer(
roi_size=(64, 256),
sw_batch_size=1,
spatial_dim=1, # Spatial dim to slice along is added here
cval=-1,
)
output = coronal_inferer(input_volume, net)
# Output is a 3D volume with 2D slices aggregated
print("Coronal Inferer Output Shape: ", output.shape)
| modules/2d_inference_3d_volume.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import plotly.express as px
import pickle
from sklearn.naive_bayes import GaussianNB
with open('credit.pkl', 'rb') as f:
X_credit_treinamento, X_credit_teste, y_credit_treinamento, y_credit_teste = pickle.load(f)
X_credit_treinamento.shape, y_credit_treinamento.shape
X_credit_teste.shape, y_credit_teste.shape
naive_credit_data = GaussianNB()
naive_credit_data.fit(X_credit_treinamento, y_credit_treinamento)
previsoes = naive_credit_data.predict(X_credit_teste)
previsoes
y_credit_teste
from sklearn.metrics import accuracy_score, confusion_matrix
accuracy_score(y_credit_teste, previsoes)
confusion_matrix(y_credit_teste, previsoes)
# +
#from yellowbrick.classifier import ConfusionMatrix - Precisa baixar o yellowbrick
# -
| estudo/aprendizagem-bayesiana/.ipynb_checkpoints/treinamento-credit-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json
json_path = '/home/edgar/cocosplit/hasty_merged.json'
with open(json_path) as json_file:
json_data = json.load(json_file)
cat_list = json_data['categories']
cat_list
# + tags=[]
json_ann = json_data['annotations']
new_ann = []
for annotation in json_ann:
cat_id = annotation['category_id']
if cat_id == 7:
annotation['category_id'] = 2
annotation['iscrowd'] = 1
elif cat_id == 8:
annotation['category_id'] = 6
annotation['iscrowd'] = 1
new_ann.append(annotation)
# json['annotations'] = new_ann
# print("target : " + nv[clssInd[cat_id]])
# -
print(len(new_ann))
print(len(json_ann))
json_data['annotations'] = new_ann
with open('hasty_merged.json', 'w') as outfile:
json.dump(json_data, outfile)
len(json_data['annotations'])
| PythonAPI/hasty_iscrowd.ipynb |
# # 📃 Solution for Exercise M7.01
#
# This notebook aims at building baseline classifiers, which we'll use to
# compare our predictive model. Besides, we will check the differences with
# the baselines that we saw in regression.
#
# We will use the adult census dataset, using only the numerical features.
# +
import pandas as pd
adult_census = pd.read_csv("../datasets/adult-census-numeric-all.csv")
data, target = adult_census.drop(columns="class"), adult_census["class"]
# -
# First, define a `ShuffleSplit` cross-validation strategy taking half of the
# sample as a testing at each round.
# +
# solution
from sklearn.model_selection import ShuffleSplit
cv = ShuffleSplit(n_splits=10, test_size=0.5, random_state=0)
# -
# Next, create a machine learning pipeline composed of a transformer to
# standardize the data followed by a logistic regression.
# +
# solution
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegression
classifier = make_pipeline(StandardScaler(), LogisticRegression())
# -
# Get the test score by using the model, the data, and the cross-validation
# strategy that you defined above.
# +
# solution
from sklearn.model_selection import cross_validate
result_classifier = cross_validate(classifier, data, target, cv=cv, n_jobs=2)
test_score_classifier = pd.Series(
result_classifier["test_score"], name="Classifier score")
# -
# Using the `sklearn.model_selection.permutation_test_score` function,
# check the chance level of the previous model.
# +
# solution
from sklearn.model_selection import permutation_test_score
score, permutation_score, pvalue = permutation_test_score(
classifier, data, target, cv=cv, n_jobs=2, n_permutations=10)
test_score_permutation = pd.Series(permutation_score, name="Permuted score")
# -
# Finally, compute the test score of a dummy classifier which would predict
# the most frequent class from the training set. You can look at the
# `sklearn.dummy.DummyClassifier` class.
# +
# solution
from sklearn.dummy import DummyClassifier
dummy = DummyClassifier(strategy="most_frequent")
result_dummy = cross_validate(dummy, data, target, cv=cv, n_jobs=2)
test_score_dummy = pd.Series(result_dummy["test_score"], name="Dummy score")
# -
# Now that we collected the results from the baselines and the model, plot
# the distributions of the different test scores.
# We concatenate the different test score in the same pandas dataframe.
# solution
final_test_scores = pd.concat(
[test_score_classifier, test_score_permutation, test_score_dummy],
axis=1,
)
# Next, plot the distributions of the test scores.
# +
# solution
import matplotlib.pyplot as plt
final_test_scores.plot.hist(bins=50, density=True, edgecolor="black")
plt.legend(bbox_to_anchor=(1.05, 0.8), loc="upper left")
plt.xlabel("Accuracy (%)")
_ = plt.title("Distribution of the test scores")
# + [markdown] tags=["solution"]
# We observe that the dummy classifier with the strategy `most_frequent` is
# equivalent to the permutation score. We can also conclude that our model
# is better than the other baseline.
# -
# Change the strategy of the dummy classifier to `stratified`, compute the
# results and plot the distribution together with the other results. Explain
# why the results get worse.
# solution
dummy = DummyClassifier(strategy="stratified")
result_dummy_stratify = cross_validate(dummy, data, target, cv=cv, n_jobs=2)
test_score_dummy_stratify = pd.Series(
result_dummy_stratify["test_score"], name="Dummy 'stratify' score")
# + tags=["solution"]
final_test_scores = pd.concat(
[
test_score_classifier, test_score_permutation,
test_score_dummy, test_score_dummy_stratify,
],
axis=1,
)
# + tags=["solution"]
final_test_scores.plot.hist(bins=50, density=True, edgecolor="black")
plt.legend(bbox_to_anchor=(1.05, 0.8), loc="upper left")
plt.xlabel("Accuracy (%)")
_ = plt.title("Distribution of the test scores")
# + [markdown] tags=["solution"]
# We see that using `strategy="stratified"`, the results are much worse than
# with the `most_frequent` strategy. Since the classes are imbalanced,
# predicting the most frequent involves that we will be right for the
# proportion of this class (~75% of the samples). However, by using the
# `stratified` strategy, wrong predictions will be made even for the most
# frequent class, hence we obtain a lower accuracy.
| notebooks/cross_validation_sol_02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ---
# title: "Apply Operations Over Items In A List"
# author: "<NAME>"
# date: 2017-12-20T11:53:49-07:00
# description: "Apply Operations Over Items In A List"
# type: technical_note
# draft: true
# ---
| docs/python/basics/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import praw
import re
import datetime as dt
import seaborn as sns
import requests
import json
import sys
import time
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras import regularizers
# -
wsb = pd.read_pickle("wsb_cleaned.pkl")
wsb.columns
| Sandbox/Notebooks/ExploratoryDataAnalysis/SentimentAnalysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.11 64-bit (''sdia-python'': conda)'
# name: python3
# ---
# # Practical session 3 - Practice with numpy
#
# Course: [SDIA-Python](https://github.com/guilgautier/sdia-python)
#
# Date: 10/06/2021
#
# Instructor: [<NAME>](https://guilgautier.github.io/)
#
# Students (pair):
# - [Student 1]([link](https://github.com/username1))
# - [Student 2]([link](https://github.com/username2))
# +
# %load_ext autoreload
# %autoreload 2
import numpy as np
from sdia_python.lab2.utils import get_random_number_generator
# -
my_array = np.array([0])
print(type(my_array))
dir(np.ndarray)
# Propose at leat 2 ways to create an integer vector of size 100 made of 1s
A = np.ones(100)
print(A)
A = np.full(100,1)
print(A)
# Create a vector with values ranging from 10 to 49
U = np.arange(10,50)
print(U)
# Propose a way to construct the vector $(0.0, 0.2, 0.4, 0.6, 0.8)$
U = np.linspace(0,0.8,5)
print(U)
U = np.arange(0,1,0.2)
print(U)
# Convert a float array into an integer array in place
U=np.array([5.,6.,2.5,3.8,3.14,9.58])
print(U)
V=U.astype(int)
print(V)
# Given a boolean array
#
# - return the indices where
# - negate the array inplace?
# +
U = np.array([True, False, False, True, True, True, False, True])
print(U)
print(np.argwhere(U))
print(np.where(U == False,True,False))
# -
# Given 2 vectors $u, v$, propose at least
#
# - 2 ways to compute the inner product $v^{\top} u$ (here they must have the same size)
# - 2 ways to compute the outer product matrix $u v^{\top}$
# - 2 ways to compute the outer sum matrix "$M = u + v^{\top}$", where $M_{ij} = u_i + v_j$
# +
U=np.array([1,2,3,4,5,6])
V=np.array([7,8,9,10,11,12])
print(np.dot(U,V))
print(np.inner(U,V))
tU = np.reshape(U,(-1,1))
tV = np.reshape(V,(-1,1))
print(np.outer(U,V))
print(np.dot(tU,tV.T))
M = tU.T + tV
print(M)
M=np.add.outer(U,V)
print(M)
# -
# Given the following matrix
#
# $$
# M =
# \begin{pmatrix}
# 0 & 1 & 2 \\
# 3 & 4 & 5 \\
# 6 & 7 & 8 \\
# \end{pmatrix}
# $$
#
# - Create $M$ using as a list of lists and access the element in the middle
# - Propose at least 2 ways to create $M$ using numpy and access the element in the middle
# - Swap its first and second row
# - Propose at least 3 ways to extract the submatrix $\begin{pmatrix}4 & 5 \\7 & 8 \\\end{pmatrix}$
# - Propose at least 2 ways to extract the diagonal of $M$
# - Propose at least 2 ways to compute $M^3$
# - Compute $v^{\top} M$, resp. $M N$ for a vector, resp. a matrix of your choice.
# - Propose 2 ways to "vectorize" the matrix, i.e., transform it into
# - $(0, 1, 2, 3, 4, 5, 6, 7, 8)$
# - $(0, 3, 6, 1, 4, 7, 2, 5, 8)$
# - Consider $v = (1, 2 , 3)$, compute the
# - row-wise multiplication of $M$ by $v$ ($M_{i\cdot}$ is multiplied by $v_i$)
# - column-wise multiplication of $M$ by $v$ ($M_{\cdot j}$ is multiplied by $v_i$)
# +
M=[ [0,1,2],
[3,4,5],
[6,7,8] ]
print(M[1][1])
M=np.array(M)
print(M)
print(M[1][1])
M=np.arange(9).reshape(3,3)
print(M)
print(M[1,1])
#M[[1,0]] = M[[0,1]]
#print(M)
#print(M[1:,1:])
#print(M[np.ix_([1,2],[1,2])])
#print(np.array([M[1][1:],M[2][1:]]))
print(np.diag(M))
#print([M[i][i] for i in range(len(M))])
print(M@M@M)
#print(np.dot(np.dot(M,M),M))
V=np.array(range(3))
print(np.dot(V,M))
N=np.array(range(12)).reshape(3,4)
print(M@N)
print(M.reshape(1,-1))
print((M.T).reshape(1,-1))
V = np.array([1,2,3])
print(V*M[:,None])
print((V*(M.T)[:,None]).T)
# -
# Write a function `is_symmetric` that checks whether a given n x n matrix is symmetric, and provide an example
# +
def is_symmetric(M):
assert len(M) == len(M[0])
return (M.T == M).all()
print(is_symmetric(M))
N = np.array([[1,2,3],
[2,4,5],
[3,5,8]])
print(is_symmetric(N))
# -
# ## Random
#
# REQUIREMENT: USE THE FUNCTION `get_random_number_generator` as previously used in Lab 2.
from sdia_python.lab2.utils import get_random_number_generator
# Consider the Bernoulli(0.4) distribution
#
# - Propose at least 2 ways to generate n=1000 samples from it
# - Compute the empirical mean and variance
# +
#2 ways to generate samples of a Bernoulli distribution
rng = get_random_number_generator(100)
a = rng.binomial(1, 0.4, 1000)
b = (rng.uniform(0,1,1000)<0.4).astype(int)
print (a,b)
# -
#empirical mean and variance
print (np.mean(a))
print (np.mean(b))
print (np.var(a))
print (np.var(b))
# Consider a random matrix of size $50 \times 100$, filled with i.i.d. standard Gaussian variables, compute
#
# - the absolute value of each entry
# - the sum of each row
# - the sum of each colomn
# - the (euclidean) norm of each row
# - the (euclidean) norm of each column
#generate a matrix of size 50×100 , filled with i.i.d. standard Gaussian variables
c = rng.normal(size = (50,100))
print (c)
#absolute value
print (abs(c))
#
#sum of each row
print (np.sum(c, axis=1))
#sum of each column
print (np.sum(c, axis=0))
#euclidian norm of each row
print(np.linalg.norm(c, axis = 1))
#euclidian norm of each column
print(np.linalg.norm(c, axis = 0))
| notebooks/lab3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Hash map function choice
#
# One of the features of most hash table keys is that they will be *strings*. These will not be immediately useful for making an integer that can be changed to a hash table address.
#
# For example, the common example of a _modulus_ function to select the slot in the table requires an integer.
#
# A hash map function must be used to change a string key into an integer?.
#
# One possible way is the ord() function which takes a character and returns its numbering in the Unicode lookup table data.
# run this cell to see ord values for 'a', 'b' and 'z'
for letter in 'a', 'b', 'z':
ord_letter = ord(letter)
print(f'ord({letter})= {ord_letter}')
# +
# now your turn - What is the ord values of 'A', 'Z', '$', '\n'
# -
# So one commonly suggested way to produce an integer key from a string is to add together the ord() values of its characters.
# run this cell to define the function str_hash_simple
def str_hash_simple(mystring, hash_table_size=27):
_sum = 0
for _char in mystring:
_sum += ord(_char)
return _sum%hash_table_size
# run this cell to see the str_hash_simple value of "AUG"
str_hash_simple('AUG')
# One obvious issue with a simple sum is that of course it does not depend on order. This would create a lot of collisions especially with a limited alphabet.
# +
# now your turn - write code to get the str_hash_simple() values of 'GUA' and 'AGU' - why?
# -
# A way to get around this is to weight each character position. For example using the position of the character in the string. This is available via the Python [enumerate function](https://www.programiz.com/python-programming/methods/built-in/enumerate).
# run this cell to define str_hash_02
def str_hash_02(mystring, hash_table_size=27):
_sum = 0
for index, char in enumerate(mystring, start=1):
_sum += ord(char) * index
return _sum%hash_table_size
# +
# now your turn - write code to get the str_hash_02 values of 'GUA' and 'AGU'
# -
# One other simple approach is to use binary numbers. These are numbers in the base 2 so the binary '10' is equivalent to the decimal number 2. If this is unfamilar to you please see https://www.ducksters.com/kidsmath/binary_numbers_basics.php
#
# To convert decimal numbers to binary strings in python it is easiest to use the format function:
# run this cell to see how to convert decimals to binary in python
for value in range(17):
binary_string = format(value, 'b')
print(f'decimal integer {value} in binary is {binary_string}')
# So an alternative hash map is convert the ordinals into binary but then to simply concatenate the digits rather than using addition. The result can be converted back to a base 10 integer key using the int() function with a base '2' option.
#
# So `GUA` would be
# * `71` `85` `65` in decimal or
# * `1000111` `1010101` `1000001` in binary.
# * Concatenating the 3 binary strings gives `100011110101011000001`
# * which is converted back to `1174209` by the [Python `int` function](https://www.geeksforgeeks.org/python-int-function/).
# * The final result would be `1174209%27` = `6` using the default `hash_table_size=27`
# run this cell to see 1174209%27 from worked example
1174209%27
# run this cell to define str_hash_03 function
def str_hash_03(mystring, hash_table_size=27):
concat = ''.join(format(ord(c), 'b') for c in mystring)
return int(concat,2)%hash_table_size
# +
# now your turn write Python to confirm that str_hash_03('GUA') is 6
# +
# Write Python to find out what is the str_hash_03 for 'AGU'
# does this differ from the value for 'GUA'
# -
# # Optional advanced question
#
# Another (possibley neater) way of using conversion to binary with [Python Bitwise Operators](https://www.tutorialspoint.com/python/bitwise_operators_example.htm). Can you come up with a function that uses this? Does your function lead to different hash bins for `GUA` and `AGU`
| demo_string_key_to_integer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (TensorFlow 2.3 Python 3.7 CPU Optimized)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:us-east-1:081325390199:image/tensorflow-2.3-cpu-py37-ubuntu18.04-v1
# ---
# # Amazon SageMaker Model Monitor
# This notebook shows how to:
# * Host a machine learning model in Amazon SageMaker and capture inference requests, results, and metadata
# * Analyze a training dataset to generate baseline constraints
# * Monitor a live endpoint for violations against constraints
#
# ---
# ## Background
#
# Amazon SageMaker provides every developer and data scientist with the ability to build, train, and deploy machine learning models quickly. Amazon SageMaker is a fully-managed service that encompasses the entire machine learning workflow. You can label and prepare your data, choose an algorithm, train a model, and then tune and optimize it for deployment. You can deploy your models to production with Amazon SageMaker to make predictions and lower costs than was previously possible.
#
# In addition, Amazon SageMaker enables you to capture the input, output and metadata for invocations of the models that you deploy. It also enables you to analyze the data and monitor its quality. In this notebook, you learn how Amazon SageMaker enables these capabilities.
#
# ---
# ## Setup
#
# To get started, make sure you have these prerequisites completed.
#
# * Specify an AWS Region to host your model.
# * An IAM role ARN exists that is used to give Amazon SageMaker access to your data in Amazon Simple Storage Service (Amazon S3). See the documentation for how to fine tune the permissions needed.
# * Create an S3 bucket used to store the data used to train your model, any additional model data, and the data captured from model invocations. For demonstration purposes, you are using the same bucket for these. In reality, you might want to separate them with different security policies.
# +
import boto3
import os
import sagemaker
from sagemaker import get_execution_role
region = boto3.Session().region_name
role = get_execution_role()
sess = sagemaker.session.Session()
bucket = sess.default_bucket()
prefix = 'tf-2-workflow'
s3_capture_upload_path = 's3://{}/{}/monitoring/datacapture'.format(bucket, prefix)
reports_prefix = '{}/reports'.format(prefix)
s3_report_path = 's3://{}/{}'.format(bucket,reports_prefix)
print("Capture path: {}".format(s3_capture_upload_path))
print("Report path: {}".format(s3_report_path))
# -
# # PART A: Capturing real-time inference data from Amazon SageMaker endpoints
# Create an endpoint to showcase the data capture capability in action.
#
# ### Deploy the model to Amazon SageMaker
# Start with deploying the trained TensorFlow model from lab 03.
# +
import boto3
def get_latest_training_job_name(base_job_name):
client = boto3.client('sagemaker')
response = client.list_training_jobs(NameContains=base_job_name, SortBy='CreationTime',
SortOrder='Descending', StatusEquals='Completed')
if len(response['TrainingJobSummaries']) > 0 :
return response['TrainingJobSummaries'][0]['TrainingJobName']
else:
raise Exception('Training job not found.')
def get_training_job_s3_model_artifacts(job_name):
client = boto3.client('sagemaker')
response = client.describe_training_job(TrainingJobName=job_name)
s3_model_artifacts = response['ModelArtifacts']['S3ModelArtifacts']
return s3_model_artifacts
latest_training_job_name = get_latest_training_job_name('tf-2-workflow')
print(latest_training_job_name)
model_path = get_training_job_s3_model_artifacts(latest_training_job_name)
print(model_path)
# -
# Here, you create the model object with the image and model data.
# +
from sagemaker.tensorflow.model import TensorFlowModel
tensorflow_model = TensorFlowModel(
model_data = model_path,
role = role,
framework_version = '2.3.1'
)
# +
from time import gmtime, strftime
from sagemaker.model_monitor import DataCaptureConfig
endpoint_name = 'tf-2-workflow-endpoint-' + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
print(endpoint_name)
predictor = tensorflow_model.deploy(
initial_instance_count=1,
instance_type='ml.m5.xlarge',
endpoint_name=endpoint_name,
data_capture_config=DataCaptureConfig(
enable_capture=True,
sampling_percentage=100,
destination_s3_uri=s3_capture_upload_path
)
)
# -
# ### Prepare dataset
#
# Next, we'll import the dataset. The dataset itself is small and relatively issue-free. For example, there are no missing values, a common problem for many other datasets. Accordingly, preprocessing just involves normalizing the data.
# +
import numpy as np
from tensorflow.python.keras.datasets import boston_housing
from sklearn.preprocessing import StandardScaler
(x_train, y_train), (x_test, y_test) = boston_housing.load_data()
scaler = StandardScaler()
scaler.fit(x_train)
x_train = scaler.transform(x_train)
x_test = scaler.transform(x_test)
# -
# ## Invoke the deployed model
#
# You can now send data to this endpoint to get inferences in real time. Because you enabled the data capture in the previous steps, the request and response payload, along with some additional metadata, is saved in the Amazon Simple Storage Service (Amazon S3) location you have specified in the DataCaptureConfig.
# This step invokes the endpoint with included sample data for about 3 minutes. Data is captured based on the sampling percentage specified and the capture continues until the data capture option is turned off.
# +
# %%time
import time
print("Sending test traffic to the endpoint {}. \nPlease wait...".format(endpoint_name))
flat_list =[]
for item in x_test:
result = predictor.predict(item)['predictions']
flat_list.append(float('%.1f'%(np.array(result))))
time.sleep(1.8)
print("Done!")
print('predictions: \t{}'.format(np.array(flat_list)))
# -
# ## View captured data
#
# Now list the data capture files stored in Amazon S3. You should expect to see different files from different time periods organized based on the hour in which the invocation occurred. The format of the Amazon S3 path is:
#
# `s3://{destination-bucket-prefix}/{endpoint-name}/{variant-name}/yyyy/mm/dd/hh/filename.jsonl`
#
# <b>Note that the delivery of capture data to Amazon S3 can require a couple of minutes so next cell might error. If this happens, please retry after a minute.</b>
s3_client = boto3.Session().client('s3')
result = s3_client.list_objects(Bucket=bucket, Prefix='tf-2-workflow/monitoring/datacapture/')
capture_files = [capture_file.get("Key") for capture_file in result.get('Contents')]
print("Found Capture Files:")
print("\n ".join(capture_files))
# Next, view the contents of a single capture file. Here you should see all the data captured in an Amazon SageMaker specific JSON-line formatted file. Take a quick peek at the first few lines in the captured file.
# +
def get_obj_body(obj_key):
return s3_client.get_object(Bucket=bucket, Key=obj_key).get('Body').read().decode("utf-8")
capture_file = get_obj_body(capture_files[-1])
print(capture_file[:2000])
# -
# Finally, the contents of a single line is present below in a formatted JSON file so that you can observe a little better.
import json
print(json.dumps(json.loads(capture_file.split('\n')[0]), indent=2))
# As you can see, each inference request is captured in one line in the jsonl file. The line contains both the input and output merged together. In the example, you provided the ContentType as `text/csv` which is reflected in the `observedContentType` value. Also, you expose the encoding that you used to encode the input and output payloads in the capture format with the `encoding` value.
#
# To recap, you observed how you can enable capturing the input or output payloads to an endpoint with a new parameter. You have also observed what the captured format looks like in Amazon S3. Next, continue to explore how Amazon SageMaker helps with monitoring the data collected in Amazon S3.
# # PART B: Model Monitor - Baselining and continuous monitoring
# In addition to collecting the data, Amazon SageMaker provides the capability for you to monitor and evaluate the data observed by the endpoints. For this:
# 1. Create a baseline with which you compare the realtime traffic.
# 1. Once a baseline is ready, setup a schedule to continously evaluate and compare against the baseline.
# ## 1. Constraint suggestion with baseline/training dataset
# The training dataset with which you trained the model is usually a good baseline dataset. Note that the training dataset data schema and the inference dataset schema should exactly match (i.e. the number and order of the features).
#
# From the training dataset you can ask Amazon SageMaker to suggest a set of baseline `constraints` and generate descriptive `statistics` to explore the data. For this example, upload the training dataset that was used to train the pre-trained model included in this example. If you already have it in Amazon S3, you can directly point to it.
# ### Prepare training dataset with headers
# +
import pandas as pd
dt = pd.DataFrame(data = x_train,
columns = ["CRIM", "ZN", "INDUS", "CHAS","NOX","RM","AGE","DIS","RAD","TAX","PTRATIO","B","LSTAT"])
dt.to_csv("training-dataset-with-header.csv", index = False)
# +
# # copy over the training dataset to Amazon S3 (if you already have it in Amazon S3, you could reuse it)
baseline_prefix = prefix + '/baselining'
baseline_data_prefix = baseline_prefix + '/data'
baseline_results_prefix = baseline_prefix + '/results'
baseline_data_uri = 's3://{}/{}'.format(bucket,baseline_data_prefix)
baseline_results_uri = 's3://{}/{}'.format(bucket, baseline_results_prefix)
print('Baseline data uri: {}'.format(baseline_data_uri))
print('Baseline results uri: {}'.format(baseline_results_uri))
# -
training_data_file = open("training-dataset-with-header.csv", 'rb')
s3_key = os.path.join(baseline_prefix, 'data', 'training-dataset-with-header.csv')
boto3.Session().resource('s3').Bucket(bucket).Object(s3_key).upload_fileobj(training_data_file)
# ### Create a baselining job with training dataset
# Now that you have the training data ready in Amazon S3, start a job to `suggest` constraints. `DefaultModelMonitor.suggest_baseline(..)` starts a `ProcessingJob` using an Amazon SageMaker provided Model Monitor container to generate the constraints.
# +
from sagemaker.model_monitor import DefaultModelMonitor
from sagemaker.model_monitor.dataset_format import DatasetFormat
my_default_monitor = DefaultModelMonitor(
role=role,
instance_count=1,
instance_type='ml.m5.xlarge',
volume_size_in_gb=20,
max_runtime_in_seconds=3600,
)
my_default_monitor.suggest_baseline(
baseline_dataset=baseline_data_uri+'/training-dataset-with-header.csv',
dataset_format=DatasetFormat.csv(header=True),
output_s3_uri=baseline_results_uri,
wait=True
)
# -
# ### Explore the generated constraints and statistics
s3_client = boto3.Session().client('s3')
result = s3_client.list_objects(Bucket=bucket, Prefix=baseline_results_prefix)
report_files = [report_file.get("Key") for report_file in result.get('Contents')]
print("Found Files:")
print("\n ".join(report_files))
# +
import pandas as pd
baseline_job = my_default_monitor.latest_baselining_job
schema_df = pd.io.json.json_normalize(baseline_job.baseline_statistics().body_dict["features"])
schema_df.head(10)
# -
constraints_df = pd.io.json.json_normalize(baseline_job.suggested_constraints().body_dict["features"])
constraints_df.head(10)
# ## 2. Analyzing collected data for data quality issues
# ### Create a schedule
# You can create a model monitoring schedule for the endpoint created earlier. Use the baseline resources (constraints and statistics) to compare against the realtime traffic.
# From the analysis above, you saw how the captured data is saved - that is the standard input and output format for Tensorflow models. But Model Monitor is framework-agnostic, and expects a specific format [explained in the docs](https://docs.aws.amazon.com/sagemaker/latest/dg/model-monitor-pre-and-post-processing.html#model-monitor-pre-processing-script):
# - Input
# - Flattened JSON `{"feature0": <value>, "feature1": <value>...}`
# - Tabular `"<value>, <value>..."`
# - Output:
# - Flattened JSON `{"prediction0": <value>, "prediction1": <value>...}`
# - Tabular `"<value>, <value>..."`
#
# We need to transform the input records to comply with this requirement. Model Monitor offers _pre-processing scripts_ in Python to transform the input. The cell below has the script that will work for our case.
# +
# %%writefile preprocessing.py
import json
def preprocess_handler(inference_record):
input_data = json.loads(inference_record.endpoint_input.data)
input_data = {f"feature{i}": val for i, val in enumerate(input_data)}
output_data = json.loads(inference_record.endpoint_output.data)["predictions"][0][0]
output_data = {"prediction0": output_data}
return{**input_data}
# -
# We'll upload this script to an s3 destination and pass it as the `record_preprocessor_script` parameter to the `create_monitoring_schedule` call.
preprocessor_s3_dest_path = f"s3://{bucket}/{prefix}/artifacts/modelmonitor"
preprocessor_s3_dest = sagemaker.s3.S3Uploader.upload("preprocessing.py", preprocessor_s3_dest_path)
print(preprocessor_s3_dest)
# +
from sagemaker.model_monitor import CronExpressionGenerator
from time import gmtime, strftime
mon_schedule_name = 'DEMO-tf-2-workflow-model-monitor-schedule-' + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
my_default_monitor.create_monitoring_schedule(
monitor_schedule_name=mon_schedule_name,
endpoint_input=predictor.endpoint,
record_preprocessor_script=preprocessor_s3_dest,
output_s3_uri=s3_report_path,
statistics=my_default_monitor.baseline_statistics(),
constraints=my_default_monitor.suggested_constraints(),
schedule_cron_expression=CronExpressionGenerator.hourly(),
enable_cloudwatch_metrics=True,
)
# -
# ### Generating violations artificially
#
# In order to get some result relevant to monitoring analysis, you can try and generate artificially some inferences with feature values causing specific violations, and then invoke the endpoint with this data
#
# Looking at our RM and AGE features:
#
# - RM - average number of rooms per dwelling
# - AGE - proportion of owner-occupied units built prior to 1940
#
# Let's simulate a situation where the average number of rooms and proportion of owner-occupied units built are both -10.
df_with_violations = pd.read_csv("training-dataset-with-header.csv")
df_with_violations["RM"] = -10
df_with_violations["AGE"] = -10
df_with_violations
# ### Start generating some artificial traffic
# The cell below starts a thread to send some traffic to the endpoint. Note that you need to stop the kernel to terminate this thread. If there is no traffic, the monitoring jobs are marked as `Failed` since there is no data to process.
# +
from threading import Thread
from time import sleep
import time
def invoke_endpoint():
for item in df_with_violations.to_numpy():
result = predictor.predict(item)['predictions']
time.sleep(0.5)
def invoke_endpoint_forever():
while True:
invoke_endpoint()
thread = Thread(target = invoke_endpoint_forever)
thread.start()
# Note that you need to stop the kernel to stop the invocations
# -
# ### Describe and inspect the schedule
# Once you describe, observe that the MonitoringScheduleStatus changes to Scheduled.
desc_schedule_result = my_default_monitor.describe_schedule()
print('Schedule status: {}'.format(desc_schedule_result['MonitoringScheduleStatus']))
# ### List executions
# The schedule starts jobs at the previously specified intervals. Here, you list the latest five executions. Note that if you are kicking this off after creating the hourly schedule, you might find the executions empty. You might have to wait until you cross the hour boundary (in UTC) to see executions kick off. The code below has the logic for waiting.
#
# Note: Even for an hourly schedule, Amazon SageMaker has a buffer period of 20 minutes to schedule your execution. You might see your execution start in anywhere from zero to ~20 minutes from the hour boundary. This is expected and done for load balancing in the backend.
# +
mon_executions = my_default_monitor.list_executions()
print("We created a hourly schedule above and it will kick off executions ON the hour (plus 0 - 20 min buffer.\nWe will have to wait till we hit the hour...")
while len(mon_executions) == 0:
print("Waiting for the 1st execution to happen...")
time.sleep(60)
mon_executions = my_default_monitor.list_executions()
# -
# ### Inspect a specific execution (latest execution)
# In the previous cell, you picked up the latest completed or failed scheduled execution. Here are the possible terminal states and what each of them mean:
# * Completed - This means the monitoring execution completed and no issues were found in the violations report.
# * CompletedWithViolations - This means the execution completed, but constraint violations were detected.
# * Failed - The monitoring execution failed, maybe due to client error (perhaps incorrect role premissions) or infrastructure issues. Further examination of FailureReason and ExitMessage is necessary to identify what exactly happened.
# * Stopped - job exceeded max runtime or was manually stopped.
# +
latest_execution = mon_executions[-1] # latest execution's index is -1, second to last is -2 and so on..
#time.sleep(60)
latest_execution.wait(logs=False)
print("Latest execution status: {}".format(latest_execution.describe()['ProcessingJobStatus']))
print("Latest execution result: {}".format(latest_execution.describe()['ExitMessage']))
latest_job = latest_execution.describe()
if (latest_job['ProcessingJobStatus'] != 'Completed'):
print("====STOP==== \n No completed executions to inspect further. Please wait till an execution completes or investigate previously reported failures.")
# -
report_uri=latest_execution.output.destination
print('Report Uri: {}'.format(report_uri))
# ### List the generated reports
# +
from urllib.parse import urlparse
s3uri = urlparse(report_uri)
report_bucket = s3uri.netloc
report_key = s3uri.path.lstrip('/')
print('Report bucket: {}'.format(report_bucket))
print('Report key: {}'.format(report_key))
s3_client = boto3.Session().client('s3')
result = s3_client.list_objects(Bucket=report_bucket, Prefix=report_key)
report_files = [report_file.get("Key") for report_file in result.get('Contents')]
print("Found Report Files:")
print("\n ".join(report_files))
# -
# ### Violations report
# If there are any violations compared to the baseline, they will be listed here.
violations = my_default_monitor.latest_monitoring_constraint_violations()
pd.set_option('display.max_colwidth', -1)
constraints_df = pd.io.json.json_normalize(violations.body_dict["violations"])
constraints_df.head(10)
# ### Triggering execution manually
#
# In order to trigger the execution manually, we first get all paths to data capture, baseline statistics, baseline constraints, etc.
# Then, we use a utility fuction, defined in <a href="./monitoringjob_utils.py">monitoringjob_utils.py</a>, to run the processing job.
# +
result = s3_client.list_objects(Bucket=bucket, Prefix='tf-2-workflow/monitoring/datacapture/')
capture_files = ['s3://{0}/{1}'.format(bucket, capture_file.get("Key")) for capture_file in result.get('Contents')]
print("Capture Files: ")
print("\n".join(capture_files))
data_capture_path = capture_files[len(capture_files) - 1][: capture_files[len(capture_files) - 1].rfind('/')]
statistics_path = baseline_results_uri + '/statistics.json'
constraints_path = baseline_results_uri + '/constraints.json'
print(data_capture_path)
print(preprocessor_s3_dest)
print(statistics_path)
print(constraints_path)
print(s3_report_path)
# +
from monitoringjob_utils import run_model_monitor_job_processor
processor = run_model_monitor_job_processor(region, 'ml.m5.xlarge',
role,
data_capture_path,
statistics_path,
constraints_path,
s3_report_path,
preprocessor_path=preprocessor_s3_dest)
# -
# ### Inspect the execution
# +
import boto3
def get_latest_model_monitor_processing_job_name(base_job_name):
client = boto3.client('sagemaker')
response = client.list_processing_jobs(NameContains=base_job_name, SortBy='CreationTime',
SortOrder='Descending', StatusEquals='Completed')
if len(response['ProcessingJobSummaries']) > 0 :
return response['ProcessingJobSummaries'][0]['ProcessingJobName']
else:
raise Exception('Processing job not found.')
def get_model_monitor_processing_job_s3_report(job_name):
client = boto3.client('sagemaker')
response = client.describe_processing_job(ProcessingJobName=job_name)
s3_report_path = response['ProcessingOutputConfig']['Outputs'][0]['S3Output']['S3Uri']
return s3_report_path
latest_model_monitor_processing_job_name = get_latest_model_monitor_processing_job_name('sagemaker-model-monitor-analyzer')
print(latest_model_monitor_processing_job_name)
report_path = get_model_monitor_processing_job_s3_report(latest_model_monitor_processing_job_name)
print(report_path)
# -
pd.set_option('display.max_colwidth', -1)
df = pd.read_json('{}/constraint_violations.json'.format(report_path))
df
# ## Delete the resources
#
# You can keep your endpoint running to continue capturing data. If you do not plan to collect more data or use this endpoint further, you should delete the endpoint to avoid incurring additional charges. Note that deleting your endpoint does not delete the data that was captured during the model invocations. That data persists in Amazon S3 until you delete it yourself.
#
# But before that, you need to delete the schedule first.
my_default_monitor.delete_monitoring_schedule()
time.sleep(120) # actually wait for the deletion
predictor.delete_endpoint()
| labs/05_model_monitor/05_model_monitor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Uploading Data Set
# +
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Applied-Modeling/master/data/'
# If you're working locally:
else:
DATA_PATH = '../data/'
# Ignore this Numpy warning when using Plotly Express:
# FutureWarning: Method .ptp is deprecated and will be removed in a future version. Use numpy.ptp instead.
import warnings
warnings.filterwarnings(action='ignore', category=FutureWarning, module='numpy')
# +
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.model_selection import train_test_split, GridSearchCV, RandomizedSearchCV
from sklearn.ensemble import RandomForestClassifier
from category_encoders import OneHotEncoder, OrdinalEncoder
from sklearn.impute import SimpleImputer
import category_encoders as ce
from sklearn.pipeline import Pipeline
from sklearn.feature_selection import SelectKBest
from sklearn.preprocessing import StandardScaler, FunctionTransformer
from sklearn.metrics import classification_report
from sklearn.pipeline import make_pipeline
from sklearn.metrics import accuracy_score
from xgboost import XGBClassifier
from sklearn.linear_model import LinearRegression, LogisticRegression
df = pd.read_csv('/Users/bradbrauser/Desktop/Data Science/MoviesOnStreamingPlatforms_updated.csv')
# -
df.shape
# # Which column in your tabular dataset will you predict, and how is your target distributed?
#
# The dataset has two rating features - IMDb and Rotten Tomatoes.
#
# IMDb is great for seeing what general audiences think of a movie. If you don’t care what the critics say and want to see what people like yourself think of a movie, then you should use IMDb. Just be aware that fans often skew the vote with 10-star ratings, which may inflate scores somewhat.
#
# Rotten Tomatoes offers the best overall picture of whether a movie is worth seeing at a glance. If you only trust the opinions of top critics and just want to know if a movie is at least decent, you should use Rotten Tomatoes. While the Fresh/Rotten binary can oversimplify the often complex opinions of critics, it should still help you weed out lousy films.
#
# My goal with this project is more in line with IMDb, as even though scores may be skewed a bit by fans of the movies, I still want to know what the public thinks, because it seems that more often than not critics do not always line up with the public opinion.
# +
def wrangle(df, thresh=500):
df = df.copy()
# Setting Title as index
df.set_index(df['Title'], inplace = True)
# Since Rotten Tomatoes feature has over 11,000 missing ratings, I'm going to just drop the Rotten Tomatoes column
df = df.drop(['Rotten Tomatoes'], axis = 1)
# Dropping rows if nulls exist in IMDb column
df.dropna(subset=['IMDb'], how='all')
# Creating new target column
df['Worth Watching?'] = df['IMDb'] >= 6.6
# Creating conditions for grading scale based on Rating column
# condition = [(df['IMDb'] >= 9.0),
# (df['IMDb'] >= 8.0) & (df['IMDb'] < 9.0),
# (df['IMDb'] >= 7.0) & (df['IMDb'] < 8.0),
# (df['IMDb'] >= 6.0) & (df['IMDb'] < 7.0),
# (df['IMDb'] >= 0) & (df['IMDb'] < 6.0)]
# # Creating grading scale
# values = ['A', 'B', 'C', 'D', 'E']
# # Creating new Rating colums
# df['Rating'] = np.select(condition, values)
# # Replacing values in Age column
# df['Age'] = df['Age'].replace(to_replace ="13+", value = 'PG')
# df['Age'] = df['Age'].replace(to_replace ="18+", value = 'R')
# df['Age'] = df['Age'].replace(to_replace ="7+", value = 'G')
# df['Age'] = df['Age'].replace(to_replace ="all", value = 'G')
# df['Age'] = df['Age'].replace(to_replace ="16+", value = 'PG-13')
# # Rename Age to MPAA Rating
# df = df.rename(columns = {'Age': 'MPAA Rating'})
# Creating individual genre columns
df['Action'] = df['Genres'].str.contains('Action')
df['Adventure'] = df['Genres'].str.contains('Adventure')
df['Animation'] = df['Genres'].str.contains('Animation')
df['Biography'] = df['Genres'].str.contains('Biography')
df['Comedy'] = df['Genres'].str.contains('Comedy')
df['Crime'] = df['Genres'].str.contains('Crime')
df['Documentary'] = df['Genres'].str.contains('Documentary')
df['Drama'] = df['Genres'].str.contains('Drama')
df['Family'] = df['Genres'].str.contains('Family')
df['Fantasy'] = df['Genres'].str.contains('Fantasy')
df['Film Noir'] = df['Genres'].str.contains('Film Noir')
df['History'] = df['Genres'].str.contains('History')
df['Horror'] = df['Genres'].str.contains('Horror')
df['Music'] = df['Genres'].str.contains('Music')
df['Musical'] = df['Genres'].str.contains('Musical')
df['Mystery'] = df['Genres'].str.contains('Mystery')
df['Romance'] = df['Genres'].str.contains('Romance')
df['Sci-Fi'] = df['Genres'].str.contains('Sci-Fi')
df['Short Film'] = df['Genres'].str.contains('Short Film')
df['Sport'] = df['Genres'].str.contains('Sport')
df['Superhero'] = df['Genres'].str.contains('Superhero')
df['Thriller'] = df['Genres'].str.contains('Thriller')
df['War'] = df['Genres'].str.contains('War')
df['Western'] = df['Genres'].str.contains('Western')
# Dropping unnecessary values
df.drop(['Genres', 'Unnamed: 0', 'ID', 'Type', 'Title', 'IMDb'], axis=1, inplace=True)
# Dropping other nulls
df = df.dropna()
# Turning boolean values into binary
df = df*1
# Split label and feature matrix
y = df['Worth Watching?']
df.drop(['Worth Watching?'], axis=1, inplace=True)
return df, y
# Wrangling
X, y = wrangle(df)
# +
from sklearn.model_selection import train_test_split, GridSearchCV, RandomizedSearchCV
# Wrangling
X, y = wrangle(df)
# -
y.value_counts(normalize = True)
# +
# Train test split on years movies were released
cutoff = 2010
X_train = X[X['Year'] < cutoff]
y_train = y.loc[X_train.index]
X_val = X[X['Year'] > cutoff]
y_val = y.loc[X_val.index]
# # Baseline
y_train.value_counts(normalize = True)
# -
import matplotlib.pyplot as plt
plt.rcParams['figure.dpi'] = 72
# +
# Random Forest Model
from sklearn.model_selection import GridSearchCV, RandomizedSearchCV
tree_model = make_pipeline(
ce.OneHotEncoder(cols=['Age', 'Directors', 'Country', 'Language',
'Action', 'Adventure', 'Animation',
'Biography', 'Comedy', 'Crime',
'Documentary', 'Drama', 'Family',
'Fantasy', 'Film Noir', 'History',
'Horror', 'Music', 'Musical', 'Mystery',
'Romance', 'Sci-Fi', 'Short Film', 'Sport',
'Superhero', 'Thriller', 'War',
'Western']),
SimpleImputer(),
StandardScaler(),
RandomForestClassifier(criterion='entropy',
max_depth=15,
n_estimators=149,
min_samples_leaf=1,
random_state=42,
min_samples_split = 40))
# Pipeline(steps=[('onehotencoder',
# OneHotEncoder(cols=['Age', 'Directors', 'Country', 'Language',
# 'Action', 'Adventure', 'Animation',
# 'Biography', 'Comedy', 'Crime',
# 'Documentary', 'Drama', 'Family',
# 'Fantasy', 'Film Noir', 'History',
# 'Horror', 'Music', 'Musical', 'Mystery',
# 'Romance', 'Sci-Fi', 'Short Film', 'Sport',
# 'Superhero', 'Thriller', 'War',
# 'Western'])),
# ('simpleimputer', SimpleImputer()),
# ('standardscaler', StandardScaler()),
# ('randomforestclassifier',
# RandomForestClassifier(criterion='entropy', max_depth=15,
# n_estimators=149, random_state=42))])
# Fitting the model
tree_model.fit(X_train, y_train)
print('Training Accuracy:', tree_model.score(X_train, y_train))
print('Validation Accuracy:', tree_model.score(X_val, y_val))
# +
# Logistic Model
log_model = Pipeline([
('oe', OrdinalEncoder()),
('imputer', SimpleImputer()),
('classifier', LogisticRegression())
])
log_model.fit(X_train, y_train);
print('Train accuracy:', log_model.score(X_train, y_train))
print('Val accuracy:', log_model.score(X_val, y_val))
# -
# # Visualizations
# +
from sklearn.metrics import plot_confusion_matrix, classification_report
import matplotlib.pyplot as plt
plt.rcParams['figure.dpi'] = 100
plot_confusion_matrix(tree_model, X_val, y_val, values_format='.0f', xticks_rotation='vertical')
# +
import numpy as np
feature = 'Netflix'
print(X_val[feature].head())
print()
print(X_val[feature].value_counts())
X_val_permuted = X_val.copy()
X_val_permuted[feature] = np.random.permutation(X_val_permuted[feature])
acc = tree_model.score(X_val, y_val)
acc_permuted = tree_model.score(X_val_permuted, y_val)
print(f'Validation accuracy with {feature}:', acc)
print(f'Validation accuracy with {feature} permuted:', acc_permuted)
print(f'Permutation importance:', acc - acc_permuted)
# +
import numpy as np
feature = 'Netflix'
print(X_val[feature].head())
print()
print(X_val[feature].value_counts())
X_val_permuted = X_val.copy()
X_val_permuted[feature] = np.random.permutation(X_val_permuted[feature])
acc = log_model.score(X_val, y_val)
acc_permuted = log_model.score(X_val_permuted, y_val)
print(f'Validation accuracy with {feature}:', acc)
print(f'Validation accuracy with {feature} permuted:', acc_permuted)
print(f'Permutation importance:', acc - acc_permuted)
# -
y_train.head()
# +
# Ignore warnings
transformers = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(strategy='median')
)
X_train_transformed = transformers.fit_transform(X_train)
X_val_transformed = transformers.transform(X_val)
model = RandomForestClassifier(n_estimators=100, random_state=42, n_jobs=-1)
model.fit(X_train_transformed, y_train)
# +
import eli5
from eli5.sklearn import PermutationImportance
permuter = PermutationImportance(
model,
scoring='accuracy',
n_iter=5,
random_state=42
)
permuter.fit(X_val_transformed, y_val)
eli5.show_weights(
permuter,
top=None,
feature_names=X_val.columns.tolist()
)
# +
# Model 6
model6 = make_pipeline(
OrdinalEncoder(),
SimpleImputer(strategy='median'),
StandardScaler(),
RandomForestClassifier(
min_samples_split=4,
max_depth=15,
n_estimators= 200,
n_jobs=1)
)
param_distributions = {
'randomforestclassifier__max_depth' : (11, 12, 13, 14, 15),
'randomforestclassifier__min_samples_split': (2, 4, 6, 8, 10),
}
search = RandomizedSearchCV(
tree_model,
param_distributions=param_distributions,
n_iter=40,
cv=7,
scoring='accuracy',
verbose = 30,
return_train_score=True,
n_jobs=4,
)
search.fit(X_train, y_train)
print('Cross-validation Best Score:', search.best_score_)
print('Best Estimator:', search.best_params_)
print('Best Model:', search.best_estimator_)
# -
Pipeline(steps=[('onehotencoder',
OneHotEncoder(cols=['Age', 'Directors', 'Country', 'Language',
'Action', 'Adventure', 'Animation',
'Biography', 'Comedy', 'Crime',
'Documentary', 'Drama', 'Family',
'Fantasy', 'Film Noir', 'History',
'Horror', 'Music', 'Musical', 'Mystery',
'Romance', 'Sci-Fi', 'Short Film', 'Sport',
'Superhero', 'Thriller', 'War',
'Western'])),
('simpleimputer', SimpleImputer()),
('standardscaler', StandardScaler()),
('randomforestclassifier',
RandomForestClassifier(criterion='entropy', max_depth=15,
n_estimators=149, random_state=42))])
# +
from sklearn.ensemble import RandomForestRegressor
from scipy.stats import randint, uniform
model7 = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(),
StandardScaler(),
RandomForestRegressor(random_state=42)
)
param_distributions = {
'targetencoder__min_samples_leaf': randint(1, 1000),
'simpleimputer__strategy': ['mean', 'median'],
'randomforestregressor__n_estimators': 15,
'randomforestregressor__max_depth': 14,
'randomforestregressor__max_features': 0.3763983510221083,
}
search.fit(X_train, y_train);
print('Best hyperparameters', search.best_params_)
print('Cross-validation MAE', -search.best_score_)
# -
search.best_estimator_
# Partial Dependence Plot
import matplotlib.pyplot as plt
plt.rcParams['figure.dpi'] = 72
# # Partial Dependence Plot
# +
from sklearn.metrics import r2_score
from xgboost import XGBRegressor
gb = make_pipeline(
ce.OrdinalEncoder(),
XGBRegressor(n_estimators=200, objective='reg:squarederror', n_jobs=-1)
)
gb.fit(X_train, y_train)
y_pred = gb.predict(X_val)
print('Gradient Boosting R^2', r2_score(y_val, y_pred))
# -
import matplotlib.pyplot as plt
plt.rcParams['figure.dpi'] = 72
X_val.isnull().sum()
import matplotlib.pyplot as plt
plt.rcParams['figure.dpi'] = 72
# +
from pdpbox.pdp import pdp_isolate, pdp_plot
feature = 'Year'
isolated = pdp_isolate(
model=tree_model,
dataset=X_val,
model_features=X_val.columns,
feature=feature
)
# -
pdp_plot(isolated, feature_name=feature, plot_lines=True);
# +
from pdpbox.pdp import pdp_interact, pdp_interact_plot
features = ['Runtime', 'Year']
interaction = pdp_interact(
model=tree_model,
dataset=X_val,
model_features=X_val.columns,
features=features
)
pdp_interact_plot(interaction, plot_type='grid', feature_names=features);
# -
row = X_val.iloc[[0]]
row
y_val.iloc[[0]]
model3.predict(row)
# !pip install shap
# +
import shap
explainer = shap.TreeExplainer(model3)
shap_values = explainer.shap_values(row)
shap.initjs()
shap.force_plot(
base_value=explainer.expected_value,
shap_values=shap_values,
features=row
)
# -
| module1-define-ml-problems/Brad_Brauser_DS17_Unit_2_Build_Week_Project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Handson
# language: python
# name: handson_ml
# ---
# +
#default_exp alexnet
# -
# export
from pwc_gabor_layer.core import GaborLayer, SigmaRegularizer
import tensorflow_datasets as tfds
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow import keras
import numpy as np
# hide
# %load_ext autoreload
# %autoreload 2
# ## Build a GaborBlock
# exports
class GaborBlock(keras.Model):
def __init__(self, filters_gabor, filters_conv, kernel_size,
learn_orientations=False, strides=(1, 1),
padding='SAME', sigma_regularizer=None, **kwargs):
super().__init__(**kwargs)
self.gabor_layer = GaborLayer(filters=filters_gabor,
kernel_size=kernel_size,
sigma_regularizer=sigma_regularizer,
learn_orientations=learn_orientations,
use_bias=False,
orientations=8,
activation='relu',
strides=strides,
padding=padding)
self.conv_layer = keras.layers.Conv2D(filters=filters_conv,
kernel_size=(1, 1),
use_bias=False,
activation='relu')
def call(self, x, training=False):
x = self.gabor_layer(x)
x = self.conv_layer(x)
return x
# hide
# Testing gabor block implementation
GB = GaborBlock(3, 3, (11, 11))
img = np.ones((1, 256, 256, 3)).astype(np.float32)
out = np.array(GB(img))
# # Implementing AlexNet in Keras
# exports
class AlexNet(keras.Model):
def __init__(self, num_classes=10, input_channels=3,
kernels1=None, kernels2=None, kernels3=None,
learn_orientations=False, **kwargs):
super().__init__(*kwargs)
self.learn_orientations = learn_orientations
self.num_classes = num_classes
self.input_channels = input_channels
self.conv1 = self.add_conv_layer(kernels1, 96, (11, 11), strides=4)
self.conv2 = self.add_conv_layer(kernels2, 256, (5, 5), strides=1)
self.conv3 = self.add_conv_layer(kernels3, 384, (3, 3), strides=1)
# Define further conv layers:
self.conv4 = self.add_conv_layer(None, 384, (3, 3), strides=1)
self.conv5 = self.add_conv_layer(None, 256, (3, 3), strides=1)
# Define output (dense connections)
self.linear1 = keras.layers.Dense(512, activation='relu')
self.linear2 = keras.layers.Dense(512, activation='relu')
self.linear3 = keras.layers.Dense(self.num_classes, activation='sigmoid')
# Supporting layers
self.maxpool = keras.layers.MaxPooling2D()
self.dropout = keras.layers.Dropout(0.5)
self.flatten = keras.layers.Flatten()
def call(self, x, training=False):
# block 1
x = self.conv1(x)
x = self.maxpool(x)
# block 2
x = self.conv2(x)
x = self.maxpool(x)
# block 3
x = self.conv3(x)
x = self.conv4(x)
x = self.conv5(x)
x = self.maxpool(x)
# classifier
x = self.flatten(x)
x = self.linear1(x)
if training:
x = self.dropout(x)
x = self.linear2(x)
if training:
x = self.dropout(x)
x = self.linear3(x)
return x
def add_conv_layer(self, gabor, filters_conv, kernel_size, strides):
if gabor:
block = GaborBlock(filters_gabor=gabor,
filters_conv=filters_conv,
kernel_size=kernel_size,
strides=strides,
learn_orientations=self.learn_orientations)
else:
block = keras.layers.Conv2D(filters=filters_conv,
kernel_size=kernel_size,
strides=strides,
activation='relu',
padding='same')
return block
# +
# hide
# Create some test examples, just to be sure
img = np.ones((1, 224, 224, 3)).astype(np.float32)
y = np.zeros((1, 10), dtype=int)
y[0, 0] = 1
# Test alexnet with gabor layers
ANG = AlexNet(kernels1=3, kernels2=3, kernels3=3)
out = np.array(ANG(img))
ANG.compile(optimizer='sgd', loss='categorical_crossentropy')
ANG.fit(x=img, y=y)
# +
# hide
# Test alexnet with conv layer
ANC = AlexNet()
out = np.array(ANC(img))
ANC.compile(optimizer='sgd', loss='categorical_crossentropy')
ANC.fit(x=img, y=y)
| 01_alexnet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Overview
#
# Specify a data folder. Run all the cleaning functions.
# Using the cleaning routine, you need to determine the start and end of activity manually.
# Once clean, we can process the EDA data and extract phasic and SMNA data.
folder = "irb/722"
SESSION_ID = "CHI2019"
# # Cleaning Functions
# Run all these cells.
# +
import json, os, csv, platform, datetime, time
from glob import glob
import numpy as np
import pandas as pd
from pprint import pprint
# from matplotlib import pyplot as pl
DATA_ROOT = "irb"
E4_MANIFEST = {
'phasic': {
'name': "Phasic GSR",
'description': "Event-specific EDA"
},
'smna':{
'name': "SMNA",
'description': "sparse SMNA driver of phasic component"
},
'textchunk': {
'name': "Text Chunk",
'description': "Values correspond to typing behaviors. # of consecutive characters typed."
},
'temp': {
'name': "Temperature",
'description': "Data from temperature sensor expressed degrees on the Celsius (°C) scale.",
'unit': "celsius"
},
'tags':{
'name': "Tags",
'description': "Event mark times. Each row corresponds to a physical button press on the device; the same time as the status LED is first illuminated. The time is expressed as a unix timestamp in UTC and it is synchronized with initial time of the session indicated in the related data files from the corresponding session."
},
'acc':{
'description': "Data from 3-axis accelerometer sensor. The accelerometer is configured to measure acceleration in the range [-2g, 2g]. Therefore the unit in this file is 1/64g. Data from x, y, and z axis are respectively in first, second, and third column.",
'name': "3-Axis Accelerometer",
'unit': "1/64g"
},
'eda':{
'description': "Data from the electrodermal activity sensor expressed as microsiemens (μS).",
'name':"Electrodermal Activity",
'unit': "μS"
},
'bvp':{
'name': "Blood Volume Pulse (BVP) from PPG",
'description': "Data from photoplethysmograph.",
},
'ibi':{
'name': "IBI",
'description': "Time between individuals heart beats extracted from the BVP signal. No sample rate is needed for this file. The first column is the time (respect to the initial time) of the detected inter-beat interval expressed in seconds (s). The second column is the duration in seconds (s) of the detected inter-beat interval (i.e., the distance in seconds from the previous beat)."
},
'hr':{
'name': "Heart rate",
'description': "Average heart rate extracted from the BVP signal.The first row is the initial time of the session expressed as unix timestamp in UTC. The second row is the sample rate expressed in Hz."
},
'kinnunen':{
'name': "Kinnunen codes",
'description': "From Kinnunen et al.; self-efficacy assessments x emotions; used as a closed-coding system. Addition of a failure stage."
}
}
def save_jsonfile(name, data):
with open(name, 'w') as outfile:
json.dump(data, outfile)
print("File saved!", name)
def gen_save_file(folder, feature):
user = os.path.basename(folder)
return os.path.join(folder, feature + "_" + user + ".json")
def get_file(folder, prefix):
user = os.path.basename(folder)
files = glob(folder + "/"+prefix+"*.json")
if len(files) == 0:
print("File not found", prefix, 'in', folder)
return None
else:
with open(files[0], 'r+') as f:
contents = json.load(f)
return contents, files[0]
def adjust_data(data, t, Fs):
metadata,f = get_file(folder, "sessionmetadata")
# ADJUST Y AND T RANGE
start = metadata["session_start"] - t
end = metadata["session_end"] - t
t0 = start * Fs
t0 = start * Fs if start > 0 else 0
tf = end * Fs - 1 if end < len(data) else len(data)
data = data[t0:tf]
return data
# -
# MAKING A SESSION METADATA JSON
def session_metadata(folder):
data = {}
data['session_start'] = session_start
data['session_end'] = session_end
data['elapsed_time'] = session_end - session_start
log = ["ACC_START", time.ctime(eda_start), "SESSION_START", time.ctime(session_start), "SESSION_END", time.ctime(session_end), "ELAPSED", str(end-start)]
data['description'] = "\n".join(log)
savefile = gen_save_file(folder, "sessionmetadata")
save_jsonfile(savefile, data)
def find_session(folder, start):
contents, filename = get_file(folder, "acc")
fig, ax = pl.subplots()
eda_start = None
eda_Fs = None
data = contents["data"]
eda_start = contents["timestamp"]
eda_Fs = contents["sampling_rate"]
# PLOTTING ACCELERATION VALUES
data = np.array(data, dtype='f')
ax.plot(data[:, 0], 'r')
ax.plot(data[:, 1], 'g')
ax.plot(data[:, 2], 'b')
# ADDING TAG EVENTS
contents, f = get_file(folder, "tags")
data = contents["data"]
for tag in data:
elapsed_time = tag - eda_start
x = elapsed_time * eda_Fs
ax.axvline(x=x)
# ADD LOG EVENTS
contents, f = get_file(folder, "log")
data = contents["data"]
timestamps = [d['time'] for d in data]
for tag in timestamps:
elapsed_time = tag - eda_start
x = elapsed_time * eda_Fs
ax.axvline(x=x, color='y')
def onclick(event):
global edaFs, eda_start
ix, iy = event.xdata, event.ydata
if start:
global session_start
session_start = eda_start + (ix/eda_Fs)
else:
global session_end
session_end = eda_start + (ix/eda_Fs)
return False
def handle_close(evt):
print('Closed Figure!')
print("START", time.ctime(eda_start))
cid = fig.canvas.mpl_connect('close_event', handle_close)
cid = fig.canvas.mpl_connect('button_press_event', onclick)
return eda_start, eda_Fs
# # Cleaning Routine
#
# 1. Run session_start and click on the start of the activity.
# 2. Run session_end and click on the start of the activity.
# 3. Verify start/end
# 4. Generate sessionmetadata file.
# 5. Run EDA analysis.
# +
# %matplotlib osx
import matplotlib
import numpy as np
import matplotlib.pyplot as pl
# import pylab as pl
#FIND SESSION START
session_start = None
eda_start, edaFs = find_session(folder, True)
# -
# FIND SESSION END
session_end = None
eda_start, edaFs = find_session(folder, False)
# Soldering
contents, filename = get_file(folder, "a-x")
session_end = contents["end_timestamp"]
session_start = contents["timestamp"]
# +
# REVIEW INFORMATION
end = datetime.datetime.fromtimestamp(session_end)
start = datetime.datetime.fromtimestamp(session_start)
print(time.ctime(eda_start))
print(time.ctime(session_start))
print(time.ctime(session_end))
print("ELAPSED", str(end-start))
# +
# # HAPPY?
session_metadata(folder)
# -
| codewords/2_StudyDataCleaning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="VqEpGyyyGE1Z" tags=["pdf-title"]
# ## Solving the linear regression problem with gradient descent
#
# Today we rewise the linear regression algorithm and it's gradient solution.
#
# Your main goal will be to __derive and implement the gradient of MSE, MAE, L1 and L2 regularization terms__ respectively in general __vector form__ (when both single observation $\mathbf{x}_i$ and corresponding target value $\mathbf{y}_i$ are vectors).
#
# This techniques will be useful later in Deep Learning module of our course as well.
#
# We will work with [Boston housing prices dataset](https://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_boston.html) subset, which have been preprocessed for your convenience.
# + colab={} colab_type="code" id="bQPlCJYI8eWB"
'''
If you are using Google Colab, uncomment the next lines to download `loss_and_derivatives.py` and `boston_subset.json`
You can open and change downloaded `.py` files in Colab using the "Files" sidebar on the left.
'''
# # !wget https://raw.githubusercontent.com/girafe-ai/ml-mipt/basic_f20/homeworks_basic/assignment0_02_Lin_reg/loss_and_derivatives.py
# # !wget https://raw.githubusercontent.com/girafe-ai/ml-mipt/basic_f20/homeworks_basic/assignment0_02_Lin_reg/boston_subset.json
# + colab={} colab_type="code" id="8lQUR89nGE1f"
# Run some setup code for this notebook.
import random
import numpy as np
import matplotlib.pyplot as plt
# Some more magic so that the notebook will reload external python modules;
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
# %load_ext autoreload
# %autoreload 2
# + colab={} colab_type="code" id="OGf3ShTNGE1q"
import json
with open('boston_subset.json', 'r') as iofile:
dataset = json.load(iofile)
feature_matrix = np.array(dataset['data'])
targets = np.array(dataset['target'])
# + [markdown] colab_type="text" id="BIUU1cOZGE10"
# ## Warming up: matrix differentiation
# _You will meet these questions later in Labs as well, so we highly recommend to answer them right here._
#
# Credits: this theoretical part is copied from [YSDA Practical_DL course](https://github.com/yandexdataschool/Practical_DL/tree/spring2019/homework01) homework01.
# + [markdown] colab_type="text" id="CvrZt_xNGE12"
# Since it easy to google every task please please please try to understand what's going on. The "just answer" thing will not be counted, make sure to present derivation of your solution. It is absolutely OK if you will find an answer on web then just exercise in $\LaTeX$ copying it into here.
# + [markdown] colab_type="text" id="ty4m156yGE15"
# Useful links:
# [1](http://www.machinelearning.ru/wiki/images/2/2a/Matrix-Gauss.pdf)
# [2](http://www.atmos.washington.edu/~dennis/MatrixCalculus.pdf)
# [3](http://cal.cs.illinois.edu/~johannes/research/matrix%20calculus.pdf)
# [4](http://research.microsoft.com/en-us/um/people/cmbishop/prml/index.htm)
# + [markdown] colab_type="text" id="k8StFOCFGE17"
# #### Inline question 1
# $$
# y = x^Tx, \quad x \in \mathbb{R}^N
# $$
#
# $$
# \frac{dy}{dx} = \frac{dx^T E x}{dx} = (E + E^T) x = 2 x
# $$
# #### E - identity matrix
# + [markdown] colab_type="text" id="qtnNCP4JGE19"
# #### Inline question 2
# $$ y = tr(AB) \quad A,B \in \mathbb{R}^{N \times N} $$
#
# $$
# \frac{dy}{dA} = B^T
# $$
# #### proved by computing trace and derivation by definition
# + [markdown] colab_type="text" id="JWfcC7_dGE2A"
# #### Inline question 3
# $$
# y = x^TAc , \quad A\in \mathbb{R}^{N \times N}, x\in \mathbb{R}^{N}, c\in \mathbb{R}^{N}
# $$
#
# $$
# \frac{dy}{dx} = Ac
# $$
#
# $$
# \frac{dy}{dA} = xc^T
# $$
#
# Hint for the latter (one of the ways): use *ex. 2* result and the fact
# $$
# tr(ABC) = tr (CAB)
# $$
# #### proved by computation and derivation by definition
# + [markdown] colab_type="text" id="WbBc_5FhGE2B"
# ## Loss functions and derivatives implementation
# You will need to implement the methods from `loss_and_derivatives.py` to go further.
# __In this assignment we ignore the bias term__, so the linear model takes simple form of
# $$
# \hat{\mathbf{y}} = XW
# $$
# where no extra column of 1s is added to the $X$ matrix.
#
# Implement the loss functions, regularization terms and their derivatives with reference to (w.r.t.) weight matrix.
#
# __Once again, you can assume that linear model is not required for bias term for now. The dataset is preprocessed for this case.__
# + [markdown] colab_type="text" id="l-CX9dTLGE1y"
# Autoreload is a great stuff, but sometimes it does not work as intended. The code below aims to fix that. __Do not forget to save your changes in the `.py` file before reloading the desired functions.__
# -
w = np.random.rand(3, 2)
X = np.random.rand(5, 3)
y = np.random.rand(5, 2)
np.transpose(X).dot(X.dot(w) - y)
w = np.array([3, -2])
np.sign(w)
# + colab={} colab_type="code" id="dtELlRTOGE2E" tags=["pdf-ignore"]
# This dirty hack might help if the autoreload has failed for some reason
try:
del LossAndDerivatives
except:
pass
from loss_and_derivatives import LossAndDerivatives
# + [markdown] colab_type="text" id="XAqMI3N98eXK"
# Mention, that in this case we compute the __MSE__ and __MAE__ for vector __y__. In the reference implementation we are averaging the error along the __y__ dimentionality as well.
#
# E.g. for residuals vector $[1., 1., 1., 1.]$ the averaged error value will be $\frac{1}{4}(1. + 1. + 1. + 1.)$
#
# This may be needed to get the desired mutliplier for loss functions derivatives. You also can refer to the `.mse` method implementation, which is already available in the `loss_and_derivatives.py`.
# + colab={} colab_type="code" id="71VCxUwHGE2L"
w = np.array([1., 1.])
x_n, y_n = feature_matrix, targets
# + [markdown] colab_type="text" id="sMN81aYyGE2T"
# Here come several asserts to check yourself:
# + colab={} colab_type="code" id="KKUYnPWuGE2V"
w = np.array([1., 1.])
x_n, y_n = feature_matrix, targets
# Repeating data to make everything multi-dimentional
w = np.vstack([w[None, :] + 0.27, w[None, :] + 0.22, w[None, :] + 0.45, w[None, :] + 0.1]).T
y_n = np.hstack([y_n[:, None], 2*y_n[:, None], 3*y_n[:, None], 4*y_n[:, None]])
# + colab={} colab_type="code" id="UtkO4hWYGE2c"
reference_mse_derivative = np.array([
[ 7.32890068, 12.88731311, 18.82128365, 23.97731238],
[ 9.55674399, 17.05397661, 24.98807528, 32.01723714]
])
reference_l2_reg_derivative = np.array([
[2.54, 2.44, 2.9 , 2.2 ],
[2.54, 2.44, 2.9 , 2.2 ]
])
assert np.allclose(
reference_mse_derivative,
LossAndDerivatives.mse_derivative(x_n, y_n, w), rtol=1e-3
), 'Something wrong with MSE derivative'
assert np.allclose(
reference_l2_reg_derivative,
LossAndDerivatives.l2_reg_derivative(w), rtol=1e-3
), 'Something wrong with L2 reg derivative'
print(
'MSE derivative:\n{} \n\nL2 reg derivative:\n{}'.format(
LossAndDerivatives.mse_derivative(x_n, y_n, w),
LossAndDerivatives.l2_reg_derivative(w))
)
# + colab={} colab_type="code" id="sKX6zLQa8eXo"
reference_mae_derivative = np.array([
[0.19708867, 0.19621798, 0.19621798, 0.19572906],
[0.25574138, 0.25524507, 0.25524507, 0.25406404]
])
reference_l1_reg_derivative = np.array([
[1., 1., 1., 1.],
[1., 1., 1., 1.]
])
assert np.allclose(
reference_mae_derivative,
LossAndDerivatives.mae_derivative(x_n, y_n, w), rtol=1e-3
), 'Something wrong with MAE derivative'
assert np.allclose(
reference_l1_reg_derivative,
LossAndDerivatives.l1_reg_derivative(w), rtol=1e-3
), 'Something wrong with L1 reg derivative'
print(
'MAE derivative:\n{} \n\nL1 reg derivative:\n{}'.format(
LossAndDerivatives.mae_derivative(x_n, y_n, w),
LossAndDerivatives.l1_reg_derivative(w))
)
# + [markdown] colab_type="text" id="kJcSPj8UGE20"
# ### Gradient descent on the real data
# Here comes small loop with gradient descent algorithm. We compute the gradient over the whole dataset.
# + colab={} colab_type="code" id="On6aSWuIGE21"
def get_w_by_grad(X, Y, w_0, loss_mode='mse', reg_mode=None, lr=0.05, n_steps=100, reg_coeff=0.05):
if loss_mode == 'mse':
loss_function = LossAndDerivatives.mse
loss_derivative = LossAndDerivatives.mse_derivative
elif loss_mode == 'mae':
loss_function = LossAndDerivatives.mae
loss_derivative = LossAndDerivatives.mae_derivative
else:
raise ValueError('Unknown loss function. Available loss functions: `mse`, `mae`')
if reg_mode is None:
reg_function = LossAndDerivatives.no_reg
reg_derivative = LossAndDerivatives.no_reg_derivative # lambda w: np.zeros_like(w)
elif reg_mode == 'l2':
reg_function = LossAndDerivatives.l2_reg
reg_derivative = LossAndDerivatives.l2_reg_derivative
elif reg_mode == 'l1':
reg_function = LossAndDerivatives.l1_reg
reg_derivative = LossAndDerivatives.l1_reg_derivative
else:
raise ValueError('Unknown regularization mode. Available modes: `l1`, `l2`, None')
w = w_0.copy()
for i in range(n_steps):
empirical_risk = loss_function(X, Y, w) + reg_coeff * reg_function(w)
gradient = loss_derivative(X, Y, w) + reg_coeff * reg_derivative(w)
gradient_norm = np.linalg.norm(gradient)
if gradient_norm > 5.:
gradient = gradient / gradient_norm * 5.
w -= lr * gradient
if i % 25 == 0:
print('Step={}, loss={},\ngradient values={}\n'.format(i, empirical_risk, gradient))
return w
# + [markdown] colab_type="text" id="ZZxVwyGJ8eX6"
# Let's check how it works.
# + colab={} colab_type="code" id="A1pyDIyqGE25"
# Initial weight matrix
w = np.ones((2,1), dtype=float)
y_n = targets[:, None]
# + colab={} colab_type="code" id="erTRQiAFGE29"
w_grad = get_w_by_grad(x_n, y_n, w, loss_mode='mse', reg_mode='l2', n_steps=250)
# + [markdown] colab_type="text" id="N4BxZybN8eYW"
# ### Comparing with `sklearn`
# Finally, let's compare our model with `sklearn` implementation.
# + colab={} colab_type="code" id="PThE47Xx8eYX"
from sklearn.linear_model import Ridge
# + colab={} colab_type="code" id="soPE4M5R8eYn"
lr = Ridge(alpha=0.05)
lr.fit(x_n, y_n)
print('sklearn linear regression implementation delivers MSE = {}'.format(np.mean((lr.predict(x_n) - y_n)**2)))
# + colab={} colab_type="code" id="Gse1m4nyGE3C"
plt.scatter(x_n[:, -1], y_n[:, -1])
plt.scatter(x_n[:, -1], x_n.dot(w_grad)[:, -1], color='orange', label='Handwritten linear regression', linewidth=5)
plt.scatter(x_n[:, -1], lr.predict(x_n), color='cyan', label='sklearn Ridge')
plt.legend()
plt.show()
# + [markdown] colab_type="text" id="jVb1wnsd8eY6"
# While the solutions may look like a bit different, remember, that handwritten linear regression was unable to fit the bias term, it was equal to $0$ by default.
# + [markdown] colab_type="text" id="6GgeWdBmGE3H"
# ### Submit your work
# To submit your work you need to log into Yandex contest (link will be provided later) and upload the `loss_and_derivatives.py` file for the corresponding problem.
| assignment_02_Lin_reg/assignment0_02_linear_regression_and_gradient_descent.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# In this notebook we will see the basics of how to use MiniSom.
#
# Let's start importing MiniSom:
from minisom import MiniSom
# MiniSom relies on the Python ecosystem to import and preprocess the data. For this example we will load the <a href="https://archive.ics.uci.edu/ml/datasets/seeds">seeds</a> dataset dataset using pandas:
# +
import pandas as pd
import numpy as np
data = pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/00236/seeds_dataset.txt',
names=['area', 'perimeter', 'compactness', 'length_kernel', 'width_kernel',
'asymmetry_coefficient', 'length_kernel_groove', 'target'],
sep='\t+', engine='python')
target = data['target'].values
label_names = {1:'Kama', 2:'Rosa', 3:'Canadian'}
data = data[data.columns[:-1]]
# data normalization
data = (data - np.mean(data, axis=0)) / np.std(data, axis=0)
data = data.values
# -
# We can initialize and train MiniSom as follows:
# +
# Initialization and training
n_neurons = 9
m_neurons = 9
som = MiniSom(n_neurons, m_neurons, data.shape[1], sigma=1.5, learning_rate=.5,
neighborhood_function='gaussian', random_seed=0)
som.pca_weights_init(data)
som.train(data, 1000, verbose=True) # random training
# -
# To visualize the result of the training we can plot the distance map (U-Matrix) using a pseudocolor where the neurons of the maps are displayed as an array of cells and the color represents the (weights) distance from the neighbour neurons. On top of the pseudo color we can add markers that repesent the samples mapped in the specific cells:
# +
import matplotlib.pyplot as plt
# %matplotlib inline
plt.figure(figsize=(9, 9))
plt.pcolor(som.distance_map().T, cmap='bone_r') # plotting the distance map as background
plt.colorbar()
# Plotting the response for each pattern in the iris dataset
# different colors and markers for each label
markers = ['o', 's', 'D']
colors = ['C0', 'C1', 'C2']
for cnt, xx in enumerate(data):
w = som.winner(xx) # getting the winner
# palce a marker on the winning position for the sample xx
plt.plot(w[0]+.5, w[1]+.5, markers[target[cnt]-1], markerfacecolor='None',
markeredgecolor=colors[target[cnt]-1], markersize=12, markeredgewidth=2)
plt.show()
# -
# To have an overview of how the samples are distributed across the map a scatter chart can be used where each dot represents the coordinates of the winning neuron. A random offset is added to avoid overlaps between points within the same cell.
# +
w_x, w_y = zip(*[som.winner(d) for d in data])
w_x = np.array(w_x)
w_y = np.array(w_y)
plt.figure(figsize=(10, 9))
plt.pcolor(som.distance_map().T, cmap='bone_r', alpha=.2)
plt.colorbar()
for c in np.unique(target):
idx_target = target==c
plt.scatter(w_x[idx_target]+.5+(np.random.rand(np.sum(idx_target))-.5)*.8,
w_y[idx_target]+.5+(np.random.rand(np.sum(idx_target))-.5)*.8,
s=50, c=colors[c-1], label=label_names[c])
plt.legend(loc='upper right')
plt.grid()
plt.savefig('resulting_images/som_seed.png')
plt.show()
# -
# To have an idea of which neurons of the map are activated more often we can create another pseudocolor plot that reflects the activation frequencies:
plt.figure(figsize=(7, 7))
frequencies = som.activation_response(data)
plt.pcolor(frequencies.T, cmap='Blues')
plt.colorbar()
plt.show()
# Wehn dealing with a supervised problem, one can visualize the proportion of samples per class falling in a specific neuron using a pie chart per neuron:
# +
import matplotlib.gridspec as gridspec
labels_map = som.labels_map(data, [label_names[t] for t in target])
fig = plt.figure(figsize=(9, 9))
the_grid = gridspec.GridSpec(n_neurons, m_neurons, fig)
for position in labels_map.keys():
label_fracs = [labels_map[position][l] for l in label_names.values()]
plt.subplot(the_grid[n_neurons-1-position[1],
position[0]], aspect=1)
patches, texts = plt.pie(label_fracs)
plt.legend(patches, label_names.values(), bbox_to_anchor=(3.5, 6.5), ncol=3)
plt.savefig('resulting_images/som_seed_pies.png')
plt.show()
# -
# To understand how the training evolves we can plot the quantization and topographic error of the SOM at each step. This is particularly important when estimating the number of iterations to run:
# +
som = MiniSom(10, 20, data.shape[1], sigma=3., learning_rate=.7,
neighborhood_function='gaussian', random_seed=10)
max_iter = 1000
q_error = []
t_error = []
for i in range(max_iter):
rand_i = np.random.randint(len(data))
som.update(data[rand_i], som.winner(data[rand_i]), i, max_iter)
q_error.append(som.quantization_error(data))
t_error.append(som.topographic_error(data))
plt.plot(np.arange(max_iter), q_error, label='quantization error')
plt.plot(np.arange(max_iter), t_error, label='topographic error')
plt.ylabel('quantization error')
plt.xlabel('iteration index')
plt.legend()
plt.show()
# -
# Notice that in the snippet above weto run each learning iteration in a for loop and saved the errors in separated lists.
| examples/BasicUsage.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Routing optical and RF ports
#
# Optical and high speed RF ports have an orientation that routes need to follow to avoid sharp turns that produce reflections.
# +
import gdsfactory as gf
gf.config.set_plot_options(show_subports=False)
gf.CONF.plotter = "matplotlib"
# -
c = gf.Component()
mmi1 = c << gf.components.mmi1x2()
mmi2 = c << gf.components.mmi1x2()
mmi2.move((100, 50))
c
# ## get_route
#
# `get_route` returns a Manhattan route between 2 ports
# +
# gf.routing.get_route?
# -
c = gf.Component("sample_connect")
mmi1 = c << gf.components.mmi1x2()
mmi2 = c << gf.components.mmi1x2()
mmi2.move((100, 50))
route = gf.routing.get_route(mmi1.ports["o2"], mmi2.ports["o1"])
c.add(route.references)
c
route
# **Problem**: get_route with obstacles
#
# sometimes there are obstacles that connect strip does not see!
c = gf.Component("sample_problem")
mmi1 = c << gf.components.mmi1x2()
mmi2 = c << gf.components.mmi1x2()
mmi2.move((110, 50))
x = c << gf.components.cross(length=20)
x.move((135, 20))
route = gf.routing.get_route(mmi1.ports["o2"], mmi2.ports["o2"])
c.add(route.references)
c
# **Solutions:**
#
# - specify the route waypoints
# - specify the route steps
# +
c = gf.Component("sample_avoid_obstacle")
mmi1 = c << gf.components.mmi1x2()
mmi2 = c << gf.components.mmi1x2()
mmi2.move((110, 50))
x = c << gf.components.cross(length=20)
x.move((135, 20))
x0 = mmi1.ports["o3"].x
y0 = mmi1.ports["o3"].y
x2 = mmi2.ports["o3"].x
y2 = mmi2.ports["o3"].y
route = gf.routing.get_route_from_waypoints(
[(x0, y0), (x2 + 40, y0), (x2 + 40, y2), (x2, y2)]
)
c.add(route.references)
c
# -
route.length
route.ports
route.references
# Lets say that we want to extrude the waveguide using a different waveguide crosssection, for example using a different layer
# +
import gdsfactory as gf
c = gf.Component("sample_connect")
mmi1 = c << gf.components.mmi1x2()
mmi2 = c << gf.components.mmi1x2()
mmi2.move((100, 50))
route = gf.routing.get_route(
mmi1.ports["o3"], mmi2.ports["o1"], cross_section=gf.cross_section.metal1
)
c.add(route.references)
c
# -
# ## auto_widen
#
# To reduce loss and phase errors you can also auto-widen waveguide routes straight sections that are longer than a certain length.
# +
import gdsfactory as gf
c = gf.Component("sample_connect")
mmi1 = c << gf.components.mmi1x2()
mmi2 = c << gf.components.mmi1x2()
mmi2.move((200, 50))
route = gf.routing.get_route(
mmi1.ports["o3"],
mmi2.ports["o1"],
cross_section=gf.cross_section.strip,
auto_widen=True,
width_wide=2,
auto_widen_minimum_length=100,
)
c.add(route.references)
c
# -
# ## get_route_from_waypoints
#
# Sometimes you need to set up a route with custom waypoints. `get_route_from_waypoints` is a manual version of `get_route`
# +
import gdsfactory as gf
c = gf.Component("waypoints_sample")
w = gf.components.straight()
left = c << w
right = c << w
right.move((100, 80))
obstacle = gf.components.rectangle(size=(100, 10))
obstacle1 = c << obstacle
obstacle2 = c << obstacle
obstacle1.ymin = 40
obstacle2.xmin = 25
p0x, p0y = left.ports["o2"].midpoint
p1x, p1y = right.ports["o2"].midpoint
o = 10 # vertical offset to overcome bottom obstacle
ytop = 20
routes = gf.routing.get_route_from_waypoints(
[
(p0x, p0y),
(p0x + o, p0y),
(p0x + o, ytop),
(p1x + o, ytop),
(p1x + o, p1y),
(p1x, p1y),
],
)
c.add(routes.references)
c
# -
# ## get_route_from_steps
#
# As you can see waypoints can only change one point (x or y) at a time, making the waypoint definition a bit redundant.
#
# You can also use a `get_route_from_steps` which is a more concise route definition, that supports defining only the new steps `x` or `y` together with increments `dx` or `dy`
#
# `get_route_from_steps` is a manual version of `get_route` and a more concise and convenient version of `get_route_from_waypoints`
# +
import gdsfactory as gf
c = gf.Component("get_route_from_steps")
w = gf.components.straight()
left = c << w
right = c << w
right.move((100, 80))
obstacle = gf.components.rectangle(size=(100, 10))
obstacle1 = c << obstacle
obstacle2 = c << obstacle
obstacle1.ymin = 40
obstacle2.xmin = 25
port1 = left.ports["o2"]
port2 = right.ports["o2"]
routes = gf.routing.get_route_from_steps(
port1=port1,
port2=port2,
steps=[
{"x": 20, "y": 0},
{"x": 20, "y": 20},
{"x": 120, "y": 20},
{"x": 120, "y": 80},
],
)
c.add(routes.references)
c
# +
import gdsfactory as gf
c = gf.Component("get_route_from_steps_shorter_syntax")
w = gf.components.straight()
left = c << w
right = c << w
right.move((100, 80))
obstacle = gf.components.rectangle(size=(100, 10))
obstacle1 = c << obstacle
obstacle2 = c << obstacle
obstacle1.ymin = 40
obstacle2.xmin = 25
port1 = left.ports["o2"]
port2 = right.ports["o2"]
routes = gf.routing.get_route_from_steps(
port1=port1,
port2=port2,
steps=[
{"x": 20},
{"y": 20},
{"x": 120},
{"y": 80},
],
)
c.add(routes.references)
c
# -
# ## get_bundle
#
# **Problem**
#
# See the route collisions When connecting groups of ports using `get_route` manhattan single-route router
# +
import gdsfactory as gf
xs_top = [0, 10, 20, 40, 50, 80]
pitch = 127
N = len(xs_top)
xs_bottom = [(i - N / 2) * pitch for i in range(N)]
layer = (1, 0)
top_ports = [
gf.Port(f"top_{i}", (xs_top[i], 0), 0.5, 270, layer=layer) for i in range(N)
]
bottom_ports = [
gf.Port(f"bottom_{i}", (xs_bottom[i], -100), 0.5, 90, layer=layer) for i in range(N)
]
c = gf.Component(name="connect_bundle")
for p1, p2 in zip(top_ports, bottom_ports):
route = gf.routing.get_route(p1, p2)
c.add(route.references)
c
# -
# **solution**
#
# `get_bundle` provides you with river routing capabilities, that you can use to route bundles of ports without collisions
# +
c = gf.Component(name="connect_bundle")
routes = gf.routing.get_bundle(top_ports, bottom_ports)
for route in routes:
c.add(route.references)
c
# +
import gdsfactory as gf
ys_right = [0, 10, 20, 40, 50, 80]
pitch = 127.0
N = len(ys_right)
ys_left = [(i - N / 2) * pitch for i in range(N)]
layer = (1, 0)
right_ports = [
gf.Port(f"R_{i}", (0, ys_right[i]), width=0.5, orientation=180, layer=layer)
for i in range(N)
]
left_ports = [
gf.Port(
f"L_{i}".format(i), (-200, ys_left[i]), width=0.5, orientation=0, layer=layer
)
for i in range(N)
]
# you can also mess up the port order and it will sort them by default
left_ports.reverse()
c = gf.Component(name="connect_bundle2")
routes = gf.routing.get_bundle(
left_ports, right_ports, sort_ports=True, start_straight_length=100
)
for route in routes:
c.add(route.references)
c
# +
xs_top = [0, 10, 20, 40, 50, 80]
pitch = 127.0
N = len(xs_top)
xs_bottom = [(i - N / 2) * pitch for i in range(N)]
layer = (1, 0)
top_ports = [
gf.Port(
f"top_{i}", midpoint=(xs_top[i], 0), width=0.5, orientation=270, layer=layer
)
for i in range(N)
]
bot_ports = [
gf.Port(
f"bot_{i}",
midpoint=(xs_bottom[i], -300),
width=0.5,
orientation=90,
layer=layer,
)
for i in range(N)
]
c = gf.Component(name="connect_bundle")
routes = gf.routing.get_bundle(
top_ports, bot_ports, separation=5.0, end_straight_length=100
)
for route in routes:
c.add(route.references)
c
# -
# `get_bundle` can also route bundles through corners
# +
import gdsfactory as gf
from gdsfactory.cell import cell
from gdsfactory.component import Component
from gdsfactory.port import Port
@cell
def test_connect_corner(N=6, config="A"):
d = 10.0
sep = 5.0
top_cell = gf.Component(name="connect_corner")
layer = (1, 0)
if config in ["A", "B"]:
a = 100.0
ports_A_TR = [
Port(
f"A_TR_{i}",
midpoint=(d, a / 2 + i * sep),
width=0.5,
orientation=0,
layer=layer,
)
for i in range(N)
]
ports_A_TL = [
Port(
f"A_TL_{i}",
midpoint=(-d, a / 2 + i * sep),
width=0.5,
orientation=180,
layer=layer,
)
for i in range(N)
]
ports_A_BR = [
Port(
f"A_BR_{i}",
midpoint=(d, -a / 2 - i * sep),
width=0.5,
orientation=0,
layer=layer,
)
for i in range(N)
]
ports_A_BL = [
Port(
f"A_BL_{i}",
midpoint=(-d, -a / 2 - i * sep),
width=0.5,
orientation=180,
layer=layer,
)
for i in range(N)
]
ports_A = [ports_A_TR, ports_A_TL, ports_A_BR, ports_A_BL]
ports_B_TR = [
Port(
f"B_TR_{i}",
midpoint=(a / 2 + i * sep, d),
width=0.5,
orientation=90,
layer=layer,
)
for i in range(N)
]
ports_B_TL = [
Port(
f"B_TL_{i}",
midpoint=(-a / 2 - i * sep, d),
width=0.5,
orientation=90,
layer=layer,
)
for i in range(N)
]
ports_B_BR = [
Port(
f"B_BR_{i}",
midpoint=(a / 2 + i * sep, -d),
width=0.5,
orientation=270,
layer=layer,
)
for i in range(N)
]
ports_B_BL = [
Port(
f"B_BL_{i}",
midpoint=(-a / 2 - i * sep, -d),
width=0.5,
orientation=270,
layer=layer,
)
for i in range(N)
]
ports_B = [ports_B_TR, ports_B_TL, ports_B_BR, ports_B_BL]
elif config in ["C", "D"]:
a = N * sep + 2 * d
ports_A_TR = [
Port(
f"A_TR_{i}",
midpoint=(a, d + i * sep),
width=0.5,
orientation=0,
layer=layer,
)
for i in range(N)
]
ports_A_TL = [
Port(
f"A_TL_{i}",
midpoint=(-a, d + i * sep),
width=0.5,
orientation=180,
layer=layer,
)
for i in range(N)
]
ports_A_BR = [
Port(
f"A_BR_{i}",
midpoint=(a, -d - i * sep),
width=0.5,
orientation=0,
layer=layer,
)
for i in range(N)
]
ports_A_BL = [
Port(
f"A_BL_{i}",
midpoint=(-a, -d - i * sep),
width=0.5,
orientation=180,
layer=layer,
)
for i in range(N)
]
ports_A = [ports_A_TR, ports_A_TL, ports_A_BR, ports_A_BL]
ports_B_TR = [
Port(
f"B_TR_{i}",
midpoint=(d + i * sep, a),
width=0.5,
orientation=90,
layer=layer,
)
for i in range(N)
]
ports_B_TL = [
Port(
f"B_TL_{i}",
midpoint=(-d - i * sep, a),
width=0.5,
orientation=90,
layer=layer,
)
for i in range(N)
]
ports_B_BR = [
Port(
f"B_BR_{i}",
midpoint=(d + i * sep, -a),
width=0.5,
orientation=270,
layer=layer,
)
for i in range(N)
]
ports_B_BL = [
Port(
f"B_BL_{i}",
midpoint=(-d - i * sep, -a),
width=0.5,
orientation=270,
layer=layer,
)
for i in range(N)
]
ports_B = [ports_B_TR, ports_B_TL, ports_B_BR, ports_B_BL]
if config in ["A", "C"]:
for ports1, ports2 in zip(ports_A, ports_B):
routes = gf.routing.get_bundle(ports1, ports2, layer=(2, 0), radius=5)
for route in routes:
top_cell.add(route.references)
elif config in ["B", "D"]:
for ports1, ports2 in zip(ports_A, ports_B):
routes = gf.routing.get_bundle(ports2, ports1, layer=(2, 0), radius=5)
for route in routes:
top_cell.add(route.references)
return top_cell
c = test_connect_corner(config="A")
c
# -
c = test_connect_corner(config="C")
c
# +
@cell
def test_connect_bundle_udirect(dy=200, angle=270, layer=(1, 0)):
xs1 = [-100, -90, -80, -55, -35, 24, 0] + [200, 210, 240]
axis = "X" if angle in [0, 180] else "Y"
pitch = 10.0
N = len(xs1)
xs2 = [70 + i * pitch for i in range(N)]
if axis == "X":
ports1 = [
Port(f"top_{i}", (0, xs1[i]), 0.5, angle, layer=layer) for i in range(N)
]
ports2 = [
Port(f"bottom_{i}", (dy, xs2[i]), 0.5, angle, layer=layer) for i in range(N)
]
else:
ports1 = [
Port(f"top_{i}", (xs1[i], 0), 0.5, angle, layer=layer) for i in range(N)
]
ports2 = [
Port(f"bottom_{i}", (xs2[i], dy), 0.5, angle, layer=layer) for i in range(N)
]
top_cell = Component(name="connect_bundle_udirect")
routes = gf.routing.get_bundle(ports1, ports2, radius=10.0)
for route in routes:
top_cell.add(route.references)
return top_cell
c = test_connect_bundle_udirect()
c
# +
@cell
def test_connect_bundle_u_indirect(dy=-200, angle=180, layer=(1, 0)):
xs1 = [-100, -90, -80, -55, -35] + [200, 210, 240]
axis = "X" if angle in [0, 180] else "Y"
pitch = 10.0
N = len(xs1)
xs2 = [50 + i * pitch for i in range(N)]
a1 = angle
a2 = a1 + 180
if axis == "X":
ports1 = [Port(f"top_{i}", (0, xs1[i]), 0.5, a1, layer=layer) for i in range(N)]
ports2 = [
Port(f"bot_{i}", (dy, xs2[i]), 0.5, a2, layer=layer) for i in range(N)
]
else:
ports1 = [Port(f"top_{i}", (xs1[i], 0), 0.5, a1, layer=layer) for i in range(N)]
ports2 = [
Port(f"bot_{i}", (xs2[i], dy), 0.5, a2, layer=layer) for i in range(N)
]
top_cell = Component("connect_bundle_u_indirect")
routes = gf.routing.get_bundle(
ports1,
ports2,
bend=gf.components.bend_euler,
radius=5,
)
for route in routes:
top_cell.add(route.references)
return top_cell
c = test_connect_bundle_u_indirect(angle=0)
c
# +
import gdsfactory as gf
@gf.cell
def test_north_to_south(layer=(1, 0)):
dy = 200.0
xs1 = [-500, -300, -100, -90, -80, -55, -35, 200, 210, 240, 500, 650]
pitch = 10.0
N = len(xs1)
xs2 = [-20 + i * pitch for i in range(N // 2)]
xs2 += [400 + i * pitch for i in range(N // 2)]
a1 = 90
a2 = a1 + 180
ports1 = [gf.Port(f"top_{i}", (xs1[i], 0), 0.5, a1, layer=layer) for i in range(N)]
ports2 = [gf.Port(f"bot_{i}", (xs2[i], dy), 0.5, a2, layer=layer) for i in range(N)]
c = gf.Component()
routes = gf.routing.get_bundle(ports1, ports2, auto_widen=False)
for route in routes:
c.add(route.references)
return c
c = test_north_to_south()
c
# +
def demo_connect_bundle():
"""combines all the connect_bundle tests"""
y = 400.0
x = 500
y0 = 900
dy = 200.0
c = gf.Component("connect_bundle")
for j, s in enumerate([-1, 1]):
for i, angle in enumerate([0, 90, 180, 270]):
ci = test_connect_bundle_u_indirect(dy=s * dy, angle=angle)
ref = ci.ref(position=(i * x, j * y))
c.add(ref)
ci = test_connect_bundle_udirect(dy=s * dy, angle=angle)
ref = ci.ref(position=(i * x, j * y + y0))
c.add(ref)
for i, config in enumerate(["A", "B", "C", "D"]):
ci = test_connect_corner(config=config)
ref = ci.ref(position=(i * x, 1700))
c.add(ref)
return c
c = demo_connect_bundle()
c
# +
import gdsfactory as gf
c = gf.Component("route_bend_5um")
c1 = c << gf.components.mmi2x2()
c2 = c << gf.components.mmi2x2()
c2.move((100, 50))
routes = gf.routing.get_bundle(
[c1.ports["o4"], c1.ports["o3"]], [c2.ports["o1"], c2.ports["o2"]], radius=5
)
for route in routes:
c.add(route.references)
c
# +
import gdsfactory as gf
c = gf.Component("electrical")
c1 = c << gf.components.pad()
c2 = c << gf.components.pad()
c2.move((200, 100))
routes = gf.routing.get_bundle(
[c1.ports["e3"]], [c2.ports["e1"]], cross_section=gf.cross_section.metal1
)
for route in routes:
c.add(route.references)
c
# +
c = gf.Component("get_bundle_with_ubends_bend_from_top")
pad_array = gf.components.pad_array()
c1 = c << pad_array
c2 = c << pad_array
c2.rotate(90)
c2.movex(1000)
c2.ymax = -200
routes_bend180 = gf.routing.get_routes_bend180(
ports=c2.get_ports_list(),
radius=75 / 2,
cross_section=gf.cross_section.metal1,
bend_port1="e1",
bend_port2="e2",
)
c.add(routes_bend180.references)
routes = gf.routing.get_bundle(
c1.get_ports_list(), routes_bend180.ports, cross_section=gf.cross_section.metal1
)
for route in routes:
c.add(route.references)
c
# +
c = gf.Component("get_bundle_with_ubends_bend_from_bottom")
pad_array = gf.components.pad_array()
c1 = c << pad_array
c2 = c << pad_array
c2.rotate(90)
c2.movex(1000)
c2.ymax = -200
routes_bend180 = gf.routing.get_routes_bend180(
ports=c2.get_ports_list(),
radius=75 / 2,
cross_section=gf.cross_section.metal1,
bend_port1="e2",
bend_port2="e1",
)
c.add(routes_bend180.references)
routes = gf.routing.get_bundle(
c1.get_ports_list(), routes_bend180.ports, cross_section=gf.cross_section.metal1
)
for route in routes:
c.add(route.references)
c
# -
# **Problem**
#
# Sometimes 90 degrees routes do not have enough space for a Manhattan route
# +
import gdsfactory as gf
c = gf.Component("route_fail_1")
c1 = c << gf.components.nxn(east=3, ysize=20)
c2 = c << gf.components.nxn(west=3)
c2.move((80, 0))
c
# +
import gdsfactory as gf
c = gf.Component("route_fail_1")
c1 = c << gf.components.nxn(east=3, ysize=20)
c2 = c << gf.components.nxn(west=3)
c2.move((80, 0))
routes = gf.routing.get_bundle(
c1.get_ports_list(orientation=0),
c2.get_ports_list(orientation=180),
auto_widen=False,
)
for route in routes:
c.add(route.references)
c
# +
c = gf.Component("route_fail_2")
pitch = 2.0
ys_left = [0, 10, 20]
N = len(ys_left)
ys_right = [(i - N / 2) * pitch for i in range(N)]
layer = (1, 0)
right_ports = [
gf.Port(f"R_{i}", (0, ys_right[i]), 0.5, 180, layer=layer) for i in range(N)
]
left_ports = [
gf.Port(f"L_{i}", (-50, ys_left[i]), 0.5, 0, layer=layer) for i in range(N)
]
left_ports.reverse()
routes = gf.routing.get_bundle(right_ports, left_ports, radius=5)
for route in routes:
c.add(route.references)
c
# -
# **Solution**
#
# Add Sbend routes using `get_bundle_sbend`
# +
import gdsfactory as gf
c = gf.Component("route_solution_1_get_bundle_sbend")
c1 = c << gf.components.nxn(east=3, ysize=20)
c2 = c << gf.components.nxn(west=3)
c2.move((80, 0))
routes = gf.routing.get_bundle_sbend(
c1.get_ports_list(orientation=0), c2.get_ports_list(orientation=180)
)
c.add(routes.references)
c
# -
routes
c = gf.Component("route_solution_2_get_bundle_sbend")
route = gf.routing.get_bundle_sbend(right_ports, left_ports)
c.add(route.references)
# ## get_bundle_from_waypoints
#
# While `get_bundle` routes bundles of ports automatically, you can also use `get_bundle_from_waypoints` to manually specify the route waypoints.
#
# You can think of `get_bundle_from_waypoints` as a manual version of `get_bundle`
#
# +
import numpy as np
import gdsfactory as gf
@gf.cell
def test_connect_bundle_waypoints(layer=(1, 0)):
"""Connect bundle of ports with bundle of routes following a list of waypoints."""
ys1 = np.array([0, 5, 10, 15, 30, 40, 50, 60]) + 0.0
ys2 = np.array([0, 10, 20, 30, 70, 90, 110, 120]) + 500.0
N = ys1.size
ports1 = [
gf.Port(
name=f"A_{i}", midpoint=(0, ys1[i]), width=0.5, orientation=0, layer=layer
)
for i in range(N)
]
ports2 = [
gf.Port(
name=f"B_{i}",
midpoint=(500, ys2[i]),
width=0.5,
orientation=180,
layer=layer,
)
for i in range(N)
]
p0 = ports1[0].position
c = gf.Component("B")
c.add_ports(ports1)
c.add_ports(ports2)
waypoints = [
p0 + (200, 0),
p0 + (200, -200),
p0 + (400, -200),
(p0[0] + 400, ports2[0].y),
]
routes = gf.routing.get_bundle_from_waypoints(ports1, ports2, waypoints)
lengths = {}
for i, route in enumerate(routes):
c.add(route.references)
lengths[i] = route.length
return c
cell = test_connect_bundle_waypoints()
cell
# +
import numpy as np
import gdsfactory as gf
c = gf.Component()
r = c << gf.components.array(
component=gf.components.straight, rows=2, columns=1, spacing=(0, 20)
)
r.movex(60)
r.movey(40)
lt = c << gf.components.straight(length=15)
lb = c << gf.components.straight(length=5)
lt.movey(5)
ports1 = lt.get_ports_list(orientation=0) + lb.get_ports_list(orientation=0)
ports2 = r.get_ports_list(orientation=180)
dx = 20
p0 = ports1[0].midpoint + (dx, 0)
p1 = (ports1[0].midpoint[0] + dx, ports2[0].midpoint[1])
waypoints = (p0, p1)
routes = gf.routing.get_bundle_from_waypoints(ports1, ports2, waypoints=waypoints)
for route in routes:
c.add(route.references)
c
# -
# ## get_bundle_from_steps
# +
import gdsfactory as gf
c = gf.Component("get_route_from_steps_sample")
w = gf.components.array(
gf.partial(gf.components.straight, layer=(2, 0)),
rows=3,
columns=1,
spacing=(0, 50),
)
left = c << w
right = c << w
right.move((200, 100))
p1 = left.get_ports_list(orientation=0)
p2 = right.get_ports_list(orientation=180)
routes = gf.routing.get_bundle_from_steps(
p1,
p2,
steps=[{"x": 150}],
)
for route in routes:
c.add(route.references)
c
# -
# ## get_bundle_path_length_match
#
# Sometimes you need to set up a route a bundle of ports that need to keep the same lengths
# +
import gdsfactory as gf
c = gf.Component("path_length_match_sample")
dy = 2000.0
xs1 = [-500, -300, -100, -90, -80, -55, -35, 200, 210, 240, 500, 650]
pitch = 100.0
N = len(xs1)
xs2 = [-20 + i * pitch for i in range(N)]
a1 = 90
a2 = a1 + 180
layer = (1, 0)
ports1 = [gf.Port(f"top_{i}", (xs1[i], 0), 0.5, a1, layer=layer) for i in range(N)]
ports2 = [gf.Port(f"bot_{i}", (xs2[i], dy), 0.5, a2, layer=layer) for i in range(N)]
routes = gf.routing.get_bundle_path_length_match(ports1, ports2)
for route in routes:
c.add(route.references)
print(route.length)
c
# -
# ### Add extra length
#
# You can also add some extra length to all the routes
# +
import gdsfactory as gf
c = gf.Component("path_length_match_sample")
dy = 2000.0
xs1 = [-500, -300, -100, -90, -80, -55, -35, 200, 210, 240, 500, 650]
pitch = 100.0
N = len(xs1)
xs2 = [-20 + i * pitch for i in range(N)]
a1 = 90
a2 = a1 + 180
layer = (1, 0)
ports1 = [gf.Port(f"top_{i}", (xs1[i], 0), 0.5, a1, layer=layer) for i in range(N)]
ports2 = [gf.Port(f"bot_{i}", (xs2[i], dy), 0.5, a2, layer=layer) for i in range(N)]
routes = gf.routing.get_bundle_path_length_match(ports1, ports2, extra_length=44)
for route in routes:
c.add(route.references)
print(route.length)
c
# -
# ### increase number of loops
#
# You can also increase the number of loops
# +
c = gf.Component("path_length_match_sample")
dy = 2000.0
xs1 = [-500, -300, -100, -90, -80, -55, -35, 200, 210, 240, 500, 650]
pitch = 200.0
N = len(xs1)
xs2 = [-20 + i * pitch for i in range(N)]
a1 = 90
a2 = a1 + 180
layer = (1, 0)
ports1 = [gf.Port(f"top_{i}", (xs1[i], 0), 0.5, a1, layer=layer) for i in range(N)]
ports2 = [gf.Port(f"bot_{i}", (xs2[i], dy), 0.5, a2, layer=layer) for i in range(N)]
routes = gf.routing.get_bundle_path_length_match(
ports1, ports2, nb_loops=2, auto_widen=False
)
for route in routes:
c.add(route.references)
print(route.length)
c
# +
# Problem, sometimes when you do path length matching you need to increase the separation
import gdsfactory as gf
c = gf.Component()
c1 = c << gf.components.straight_array(spacing=90)
c2 = c << gf.components.straight_array(spacing=5)
c2.movex(200)
c1.y = 0
c2.y = 0
routes = gf.routing.get_bundle_path_length_match(
c1.get_ports_list(orientation=0),
c2.get_ports_list(orientation=180),
end_straight_length=0,
start_straight_length=0,
separation=30,
radius=5,
)
for route in routes:
c.add(route.references)
c
# +
# Solution: increase separation
import gdsfactory as gf
c = gf.Component()
c1 = c << gf.components.straight_array(spacing=90)
c2 = c << gf.components.straight_array(spacing=5)
c2.movex(200)
c1.y = 0
c2.y = 0
routes = gf.routing.get_bundle_path_length_match(
c1.get_ports_list(orientation=0),
c2.get_ports_list(orientation=180),
end_straight_length=0,
start_straight_length=0,
separation=80, # increased
radius=5,
)
for route in routes:
c.add(route.references)
c
# -
# ## Route to IO (Pads, grating couplers ...)
#
#
# ### Route to electrical pads
# +
import gdsfactory as gf
mzi = gf.components.straight_heater_metal(length=30)
mzi
# +
import gdsfactory as gf
mzi = gf.components.mzi_phase_shifter(
length_x=30, straight_x_top=gf.components.straight_heater_metal_90_90
)
mzi_te = gf.routing.add_electrical_pads_top(component=mzi, layer=(41, 0))
mzi_te
# +
import gdsfactory as gf
hr = gf.components.straight_heater_metal()
cc = gf.routing.add_electrical_pads_shortest(component=hr, layer=(41, 0))
cc
# +
# Problem: Sometimes the shortest path does not work well
import gdsfactory as gf
c = gf.components.mzi_phase_shifter_top_heater_metal(length_x=70)
cc = gf.routing.add_electrical_pads_shortest(component=c, layer=(41, 0))
cc
# +
# Solution: you can use define the pads separate and route metal lines to them
c = gf.Component("mzi_with_pads")
c1 = c << gf.components.mzi_phase_shifter_top_heater_metal(length_x=70)
c2 = c << gf.components.pad_array(columns=2)
c2.ymin = c1.ymax + 20
c2.x = 0
c1.x = 0
c
# +
c = gf.Component("mzi_with_pads")
c1 = c << gf.components.mzi_phase_shifter(
straight_x_top=gf.components.straight_heater_metal_90_90, length_x=70 # 150
)
c2 = c << gf.components.pad_array(columns=2, orientation=270)
c2.ymin = c1.ymax + 30
c2.x = 0
c1.x = 0
ports1 = c1.get_ports_list(port_type="electrical")
ports2 = c2.get_ports_list()
routes = gf.routing.get_bundle(
ports1=ports1,
ports2=ports2,
cross_section=gf.cross_section.metal1,
width=10,
bend=gf.components.wire_corner,
)
for route in routes:
c.add(route.references)
c
# -
# ### Route to Fiber Array
#
# Routing allows you to define routes to optical or electrical IO (grating couplers or electrical pads)
# + attributes={"classes": [], "id": "", "n": "1"}
import numpy as np
import gdsfactory as gf
from gdsfactory import LAYER
from gdsfactory import Port
@gf.cell
def big_device(w=400.0, h=400.0, N=16, port_pitch=15.0, layer=LAYER.WG, wg_width=0.5):
"""big component with N ports on each side"""
component = gf.Component()
p0 = np.array((0, 0))
dx = w / 2
dy = h / 2
points = [[dx, dy], [dx, -dy], [-dx, -dy], [-dx, dy]]
component.add_polygon(points, layer=layer)
port_params = {"layer": layer, "width": wg_width}
for i in range(N):
port = Port(
name=f"W{i}",
midpoint=p0 + (-dx, (i - N / 2) * port_pitch),
orientation=180,
**port_params,
)
component.add_port(port)
for i in range(N):
port = Port(
name=f"E{i}",
midpoint=p0 + (dx, (i - N / 2) * port_pitch),
orientation=0,
**port_params,
)
component.add_port(port)
for i in range(N):
port = Port(
name=f"N{i}",
midpoint=p0 + ((i - N / 2) * port_pitch, dy),
orientation=90,
**port_params,
)
component.add_port(port)
for i in range(N):
port = Port(
name=f"S{i}",
midpoint=p0 + ((i - N / 2) * port_pitch, -dy),
orientation=-90,
**port_params,
)
component.add_port(port)
return component
component = big_device(N=10)
c = gf.routing.add_fiber_array(component=component, radius=10.0, fanout_length=60.0)
c
# +
import gdsfactory as gf
c = gf.components.ring_double(width=0.8)
cc = gf.routing.add_fiber_array(component=c, taper_length=150)
cc
# -
cc.pprint()
# You can also mix and match `TE` and `TM` grating couplers
# +
c = gf.components.mzi_phase_shifter()
gcte = gf.components.grating_coupler_te
gctm = gf.components.grating_coupler_tm
cc = gf.routing.add_fiber_array(
component=c,
optical_routing_type=2,
grating_coupler=[gctm, gcte, gctm, gcte],
radius=20,
)
cc
# -
# ### Route to fiber single
# +
import gdsfactory as gf
c = gf.components.ring_single()
cc = gf.routing.add_fiber_single(component=c)
cc
# +
import gdsfactory as gf
c = gf.components.ring_single()
cc = gf.routing.add_fiber_single(component=c, with_loopback=False)
cc
# -
c = gf.components.mmi2x2()
cc = gf.routing.add_fiber_single(component=c, with_loopback=False)
cc
c = gf.components.mmi1x2()
cc = gf.routing.add_fiber_single(component=c, with_loopback=False, fiber_spacing=150)
cc
c = gf.components.mmi1x2()
cc = gf.routing.add_fiber_single(component=c, with_loopback=False, fiber_spacing=50)
cc
c = gf.components.crossing()
cc = gf.routing.add_fiber_single(component=c, with_loopback=False)
cc
c = gf.components.cross(length=200, width=2, port_type="optical")
cc = gf.routing.add_fiber_single(component=c, with_loopback=False)
cc
c = gf.components.spiral()
cc = gf.routing.add_fiber_single(component=c, with_loopback=False)
cc
| docs/notebooks/04_routing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import scipy.stats as sts
import seaborn as sns
sns.set()
# %matplotlib inline
# -
# # 01. Smooth function optimization
#
# Рассмотрим все ту же функцию из задания по линейной алгебре:
# $ f(x) = \sin{\frac{x}{5}} * e^{\frac{x}{10}} + 5 * e^{-\frac{x}{2}} $
# , но теперь уже на промежутке `[1, 30]`.
#
# В первом задании будем искать минимум этой функции на заданном промежутке с помощью `scipy.optimize`. Разумеется, в дальнейшем вы будете использовать методы оптимизации для более сложных функций, а `f(x)` мы рассмотрим как удобный учебный пример.
# Напишите на Питоне функцию, вычисляющую значение `f(x)` по известному `x`. Будьте внимательны: не забывайте про то, что по умолчанию в питоне целые числа делятся нацело, и о том, что функции `sin` и `exp` нужно импортировать из модуля `math`.
# +
from math import sin, exp, sqrt
def f(x):
return sin(x / 5) * exp(x / 10) + 5 * exp(-x / 2)
f(10)
# +
xs = np.arange(41, 60, 0.1)
ys = np.array([f(x) for x in xs])
plt.plot(xs, ys)
# -
# Изучите примеры использования `scipy.optimize.minimize` в документации `Scipy` (см. "Материалы").
#
# Попробуйте найти минимум, используя стандартные параметры в функции `scipy.optimize.minimize` (т.е. задав только функцию и начальное приближение). Попробуйте менять начальное приближение и изучить, меняется ли результат.
from scipy.optimize import minimize, rosen, rosen_der, differential_evolution
x0 = 60
minimize(f, x0)
# поиграемся с розенброком
x0 = [1., 10.]
minimize(rosen, x0, method='BFGS')
# ___
#
# ## Submission #1
#
# Укажите в `scipy.optimize.minimize` в качестве метода `BFGS` (один из самых точных в большинстве случаев градиентных методов оптимизации), запустите из начального приближения $ x = 2 $. Градиент функции при этом указывать не нужно – он будет оценен численно. Полученное значение функции в точке минимума - ваш первый ответ по заданию 1, его надо записать с точностью до 2 знака после запятой.
# Теперь измените начальное приближение на x=30. Значение функции в точке минимума - ваш второй ответ по заданию 1, его надо записать через пробел после первого, с точностью до 2 знака после запятой.
# Стоит обдумать полученный результат. Почему ответ отличается в зависимости от начального приближения? Если нарисовать график функции (например, как это делалось в видео, где мы знакомились с Numpy, Scipy и Matplotlib), можно увидеть, в какие именно минимумы мы попали. В самом деле, градиентные методы обычно не решают задачу глобальной оптимизации, поэтому результаты работы ожидаемые и вполне корректные.
# +
# 1. x0 = 2
x0 = 2
res1 = minimize(f, x0, method='BFGS')
# 2. x0 = 30
x0 = 30
res2 = minimize(f, x0, method='BFGS')
with open('out/06. submission1.txt', 'w') as f_out:
output = '{0:.2f} {1:.2f}'.format(res1.fun, res2.fun)
print(output)
f_out.write(output)
# -
# # 02. Глобальная оптимизация
#
# Теперь попробуем применить к той же функции $ f(x) $ метод глобальной оптимизации — дифференциальную эволюцию.
# Изучите документацию и примеры использования функции `scipy.optimize.differential_evolution`.
#
# Обратите внимание, что границы значений аргументов функции представляют собой список кортежей (list, в который помещены объекты типа tuple). Даже если у вас функция одного аргумента, возьмите границы его значений в квадратные скобки, чтобы передавать в этом параметре список из одного кортежа, т.к. в реализации `scipy.optimize.differential_evolution` длина этого списка используется чтобы определить количество аргументов функции.
#
# Запустите поиск минимума функции f(x) с помощью дифференциальной эволюции на промежутке [1, 30]. Полученное значение функции в точке минимума - ответ в задаче 2. Запишите его с точностью до второго знака после запятой. В этой задаче ответ - только одно число.
# Заметьте, дифференциальная эволюция справилась с задачей поиска глобального минимума на отрезке, т.к. по своему устройству она предполагает борьбу с попаданием в локальные минимумы.
#
# Сравните количество итераций, потребовавшихся BFGS для нахождения минимума при хорошем начальном приближении, с количеством итераций, потребовавшихся дифференциальной эволюции. При повторных запусках дифференциальной эволюции количество итераций будет меняться, но в этом примере, скорее всего, оно всегда будет сравнимым с количеством итераций BFGS. Однако в дифференциальной эволюции за одну итерацию требуется выполнить гораздо больше действий, чем в BFGS. Например, можно обратить внимание на количество вычислений значения функции (nfev) и увидеть, что у BFGS оно значительно меньше. Кроме того, время работы дифференциальной эволюции очень быстро растет с увеличением числа аргументов функции.
res = differential_evolution(f, [(1, 30)])
res
# ___
#
# ## Submission #2
# +
res = differential_evolution(f, [(1, 30)])
with open('out/06. submission2.txt', 'w') as f_out:
output = '{0:.2f}'.format(res.fun)
print(output)
f_out.write(output)
# -
# # 03. Минимизация негладкой функции
#
# Теперь рассмотрим функцию $ h(x) = int(f(x)) $ на том же отрезке `[1, 30]`, т.е. теперь каждое значение $ f(x) $ приводится к типу int и функция принимает только целые значения.
#
# Такая функция будет негладкой и даже разрывной, а ее график будет иметь ступенчатый вид. Убедитесь в этом, построив график $ h(x) $ с помощью `matplotlib`.
# +
def h(x):
return int(f(x))
xs = np.arange(0, 70, 1)
ys = [h(x) for x in xs]
plt.plot(xs, ys)
# -
minimize(h, 40.3)
# Попробуйте найти минимум функции $ h(x) $ с помощью BFGS, взяв в качестве начального приближения $ x = 30 $. Получившееся значение функции – ваш первый ответ в этой задаче.
res_bfgs = minimize(h, 30)
res_bfgs
# Теперь попробуйте найти минимум $ h(x) $ на отрезке `[1, 30]` с помощью дифференциальной эволюции. Значение функции $ h(x) $ в точке минимума – это ваш второй ответ в этом задании. Запишите его через пробел после предыдущего.
res_diff_evol = differential_evolution(h, [(1, 30)])
res_diff_evol
# Обратите внимание на то, что полученные ответы различаются. Это ожидаемый результат, ведь BFGS использует градиент (в одномерном случае – производную) и явно не пригоден для минимизации рассмотренной нами разрывной функции. Попробуйте понять, почему минимум, найденный BFGS, именно такой (возможно в этом вам поможет выбор разных начальных приближений).
#
# Выполнив это задание, вы увидели на практике, чем поиск минимума функции отличается от глобальной оптимизации, и когда может быть полезно применить вместо градиентного метода оптимизации метод, не использующий градиент. Кроме того, вы попрактиковались в использовании библиотеки SciPy для решения оптимизационных задач, и теперь знаете, насколько это просто и удобно.
# ___
#
# ## Submission #3
with open('out/06. submission3.txt', 'w') as f_out:
output = '{0:.2f} {1:.2f}'.format(res_bfgs.fun, res_diff_evol.fun)
print(output)
f_out.write(output)
# ___
#
# Дальше играюсь с визуализацией ф-ии розенброка
# +
lb = -10
rb = 10
step = 0.2
gen_xs = np.arange(lb, rb, step)
xs = np.meshgrid(np.arange(-1, 1, 0.1), np.arange(-10, 10, 0.1))
ys = rosen(xs)
print(xs[0].shape, xs[1].shape, ys.shape)
# -
plt.contour(xs[0], xs[1], ys, 30)
# +
lb = 0
rb = 4
step = 0.3
gen_xs = np.arange(lb, rb, step)
#xs = np.meshgrid(gen_xs, gen_xs)
#ys = (xs[0]**2 + xs[1]**2)**0.5
xs = np.meshgrid(np.arange(-2, 2, 0.1), np.arange(-10, 10, 0.1))
ys = rosen(xs)
print(xs[0].shape, xs[1].shape, ys.shape)
cmap = sns.cubehelix_palette(light=1, as_cmap=True)
plt.contour(xs[0], xs[1], ys, 30, cmap=cmap)
#plt.plot(xs[0], xs[1], marker='.', color='k', linestyle='none', alpha=0.1)
plt.show()
# +
from mpl_toolkits.mplot3d import Axes3D
fig = plt.figure()
ax = fig.gca(projection='3d')
surf = ax.plot_surface(xs[0], xs[1], ys, cmap=cmap, linewidth=0, antialiased=False)
plt.show()
# +
x0 = [1.3, 0.7, 0.8, 1.9, 1.2]
res = minimize(rosen, x0, method='Nelder-Mead', tol=1e-6)
res.x
# -
| 01_math_and_python/06_optimizations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Caching Data
#
# Spark offers the possibility to cache data, which means that it tries to keep (intermediate) results either in memory or on disk. This can be very helpful in iterative algorithms or interactive analysis, where you want to prevent that the same processing steps are performed over and over again.
#
# ### Approach to Caching
# Instead of performing timings of individual executions, we use the `explain()` method again to see how output changes with cached intermediate results.
#
# ### Weather Example
# We will again use the weather example to understand how caching works.
# ## Create or Reuse Spark Session
# +
from pyspark.sql import SparkSession
if not 'spark' in locals():
spark = SparkSession.builder \
.master("local[*]") \
.config("spark.driver.memory","24G") \
.getOrCreate()
spark
# -
spark.conf.set("spark.sql.adaptive.enabled", False)
spark.conf.set("spark.sql.autoBroadcastJoinThreshold", -1)
# # 1. Load Data
#
# First we load the weather data, which consists of the measurement data and some station metadata.
storageLocation = "s3://dimajix-training/data/weather"
# ## 1.1 Load Measurements
#
# Measurements are stored in multiple directories (one per year)
# +
import pyspark.sql.functions as f
# Union all years together
raw_weather = spark.read.text(storageLocation + "/2003").withColumn("year", f.lit(2003))
# -
# ### Extract Measurements
#
# Measurements were stored in a proprietary text based format, with some values at fixed positions. We need to extract these values with a simple `SELECT` statement.
# +
weather = raw_weather.select(
f.col("year"),
f.substring(f.col("value"),5,6).alias("usaf"),
f.substring(f.col("value"),11,5).alias("wban"),
f.substring(f.col("value"),16,8).alias("date"),
f.substring(f.col("value"),24,4).alias("time"),
f.substring(f.col("value"),42,5).alias("report_type"),
f.substring(f.col("value"),61,3).alias("wind_direction"),
f.substring(f.col("value"),64,1).alias("wind_direction_qual"),
f.substring(f.col("value"),65,1).alias("wind_observation"),
(f.substring(f.col("value"),66,4).cast("float") / f.lit(10.0)).alias("wind_speed"),
f.substring(f.col("value"),70,1).alias("wind_speed_qual"),
(f.substring(f.col("value"),88,5).cast("float") / f.lit(10.0)).alias("air_temperature"),
f.substring(f.col("value"),93,1).alias("air_temperature_qual")
)
weather.limit(10).toPandas()
# -
# ## 1.2 Load Station Metadata
#
# We also need to load the weather station meta data containing information about the geo location, country etc of individual weather stations.
# +
stations = spark.read \
.option("header", True) \
.csv(storageLocation + "/isd-history")
# Display first 10 records
stations.limit(10).toPandas()
# -
# # 2 Caching Data
#
# For analysing the impact of cachign data, we will use a slightly simplified variant of the weather analysis (only temperature will be aggregated). We will change the execution by caching intermediate results and watch how execution plans change.
# ## 2.1 Original Execution Plan
#
# First let's have the execution plans of the original query as our reference.
joined_weather = weather.join(stations, (weather.usaf == stations.USAF) & (weather.wban == stations.WBAN))
aggregates = joined_weather.groupBy(joined_weather.CTRY, joined_weather.year).agg(
f.min(f.when(joined_weather.air_temperature_qual == f.lit(1), joined_weather.air_temperature)).alias('min_temp'),
f.max(f.when(joined_weather.air_temperature_qual == f.lit(1), joined_weather.air_temperature)).alias('max_temp')
)
# +
result = joined_weather.join(f.broadcast(aggregates), ["ctry", "year"])
# YOUR CODE HERE
# -
# ## 2.2 Caching Weather
#
# First let us simply cache the joined input DataFrame.
# +
# YOUR CODE HERE
# -
# ### Forcing physical caching
#
# The `cache()` method again works lazily and only marks the DataFrame to be cached. The physical cache itself will only take place once the elements are evaluated. A common and easy way to enforce this is to call a `count()` on the to-be cached DataFrame.
# +
# YOUR CODE HERE
# -
# ### Execution Plan with Cache
#
# Now let us have a look at the execution plan with the cache for the `weather` DataFrame enabled.
# +
aggregates = joined_weather.groupBy(joined_weather.CTRY, joined_weather.year).agg(
f.min(f.when(joined_weather.air_temperature_qual == f.lit(1), joined_weather.air_temperature)).alias('min_temp'),
f.max(f.when(joined_weather.air_temperature_qual == f.lit(1), joined_weather.air_temperature)).alias('max_temp')
)
result = joined_weather.join(f.broadcast(aggregates), ["ctry", "year"])
# YOUR CODE HERE
# -
result.limit(10).toPandas()
# ### Remarks
#
# Although the data is already cached, the execution plan still contains all steps. But the caching step won't be executed any more (since data is already cached), it is only mentioned here for completenss of the plan. We will see in the web interface.
#
# The cache itself is presented as two steps in the execution plan:
# * Creating the cache (InMemoryRelation)
# * Using the cache (InMemoryTableScan)
#
# If you look closely at the execution plans and compare these to the original uncached plan, you will notice that certain optimizations are not performed any more:
# * Cache contains ALL columns of the weather DataFrame, although only a subset is required.
# * Filter operation of JOIN is performed part of caching.
#
# Caching is an optimization barrier. This means that Spark can only optimize plans before building the cache and plans after using the cache. No optimization is possible that spans building and using the cache. The idea simply is that the DataFrame should be cached exactly how it was specified without any column truncating or record filtering in place which appears after the cache.
# ## 2.2 Uncaching Data
#
# Caches occupy resources (memory and/or disk). Once you do not need the cache any more, you'd probably like to free up the resources again. This is easily possible with the `unpersist()` method.
# +
# YOUR CODE HERE
# -
# ### Exeuction plan after unpersist
#
# Now we'd expect to have the original execution plan again. But for some reason (bug?) we don't get that any more:
# +
result = joined_weather.groupBy(joined_weather.CTRY, joined_weather.year).agg(
f.min(f.when(joined_weather.air_temperature_qual == f.lit(1), joined_weather.air_temperature)).alias('min_temp'),
f.max(f.when(joined_weather.air_temperature_qual == f.lit(1), joined_weather.air_temperature)).alias('max_temp')
)
result.explain(False)
# -
# ### Remarks
#
# As you see in the execution plan, the cache has been removed now and the plan equals to the original one before we started caching data.
# # 3 Cache Levels
#
# Spark supports different levels of cache (memory, disk and a combination). These can be specified explicitly if you use `persist()` instead of `cache()`. Cache actually is a shortcut for `persist(MEMORY_AND_DISK)`.
# +
from pyspark.storagelevel import StorageLevel
joined_weather.persist(StorageLevel.MEMORY_ONLY)
joined_weather.persist(StorageLevel.DISK_ONLY)
joined_weather.persist(StorageLevel.MEMORY_AND_DISK)
joined_weather.persist(StorageLevel.MEMORY_ONLY_2)
joined_weather.persist(StorageLevel.DISK_ONLY_2)
joined_weather.persist(StorageLevel.MEMORY_AND_DISK_2)
# -
# ### Cache level explanation
#
# * `MEMORY_ONLY` - stores all records directly in memory
# * `DISK_ONLY` - stores all records serialized on disk
# * `MEMORY_AND_DISK` - stores all records first in memory and spills onto disk when no space is left in memory
# * `..._2` - stores caches on two nodes instead of one for additional redundancy
# # 4 Caching within a Single Query
#
# Caching only helps in very rare cases within a single query, one case being if a DataFrame is used multiple times (for example in a `UNION` operation). But even then, things don't always work out nicely. Let's start with a small example, where the `weather` DataFrame is used twice with a simple modification.
# +
# Remove any caches
weather.unpersist()
result = # YOUR CODE HERE
result.explain()
# -
result.count()
# ### Adding a Cache
# Now let's use some caching to prevent Spark from reading the input twice:
# +
weather.cache()
result = weather.union(
weather.withColumn("air_temperature", 2*weather["air_temperature"])
)
result.explain()
# -
result.count()
# ## 4.1 Shuffle Reuse
#
# In some constellations Spark automatically detects that it can reuse the output of a shuffle operation. In this case, caching won't help much and even risks hurting the performance. But this Spark logic does not catch many cases.
# +
# Remove any caches
weather.unpersist()
joined_weather.unpersist()
result = # YOUR CODE HERE
result.explain()
# -
result.count()
# # 5 Don'ts
#
# Although reading from a cache can be faster than reprocessing data from scratch, especially if that involves reading original data from slow IO devices (S3) or complex operations (joins), some caution should be taken. Caching is not free, not only is it a optimization barrier, it also occupies resources (memory and disk) and definately slows down the first query that has to build the cache.
#
# In order to limit the physical resources (RAM and disk), you should reduce the amount to cache to the bare minimum and even exclude simple calculations from the cache. For example if we included conversions to mph and °F in our weather data as precalculated measurements, it would be a wise idea to exclude these simple calculations from the cache, since they would only blow up the overall volume while these conversions are simple and cheap to calculate even after reading from the cache (plus they can be removed by the optimizer when they are not needed in a specific query)
# +
# Remove any previous caches
weather.unpersist()
weather_intl = # YOUR CODE HERE
# DON'T !
# YOUR CODE HERE
# +
# Remove any previous caches
weather_intl.unpersist()
# Prefer caching the smaller input data set and perform trivial calculations after caching
# YOUR CODE HERE
weather_intl = weather.withColumn("air_temperature_fahrenheit", weather["air_temperature"]*9.0/5.0+32) \
.withColumn("wind_speed_mph", weather["wind_speed"]*2.236936)
# -
| pyspark-advanced/jupyter-caching/Caching Data - Skeleton.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:scikit-hep-tutorials]
# language: python
# name: conda-env-scikit-hep-tutorials-py
# ---
import pandas as pd
import boost_histogram as bh
import numpy as np
skhep = pd.read_csv("scikit-hep-20180101-20250101.csv", usecols=["timestamp", "file_project", "details_python"], parse_dates=["timestamp"], dtype={"file_project": "category", "details_python": "category"})
skhep.dropna(inplace=True) # Drop 11 NA's for Python version
# ## Classic method
def compute_pandas(skhep, *projects: str, year: int):
start=f"{year}-01-01"
end=f"{year}-12-31"
results = {}
for project in projects:
val = skhep[skhep.file_project == project]
val = val[(val.timestamp > start) & (val.timestamp < end)]
vers = val.details_python.str[0]
results[project] = (sum(vers == '2'), sum(vers == '3'))
return results
# %%time
results2020 = compute_pandas(skhep, *set(skhep.file_project), year=2020)
results2019 = compute_pandas(skhep, *set(skhep.file_project), year=2019)
results2018 = compute_pandas(skhep, *set(skhep.file_project), year=2018)
# ## Boost Histogram method
def compute_bh(hist, *projects: str, year: int):
results = {}
for project in projects:
ver2 = hist[bh.loc(project), bh.loc(year), bh.loc(2)]
ver3 = hist[bh.loc(project), bh.loc(year), bh.loc(3)]
results[project] = (ver2, ver3)
return results
hist = bh.Histogram(
bh.axis.StrCategory(list(set(skhep.file_project))),
bh.axis.Integer(2018, 2021, underflow=False, overflow=False),
bh.axis.Integer(2, 4, underflow=False, overflow=False),
storage=bh.storage.Int64()
)
# %%time
_ = hist.fill(np.asarray(skhep.file_project, dtype=str),
np.asarray(skhep.timestamp.dt.year),
np.asarray(skhep.details_python.str[0].astype(int)))
# %%time
results2020 = compute_bh(hist, *set(skhep.file_project), year=2020)
results2019 = compute_bh(hist, *set(skhep.file_project), year=2019)
results2018 = compute_bh(hist, *set(skhep.file_project), year=2018)
# ## Make table (from either result set)
def show(*args):
results = args[0]
keys = filter(lambda x: sum(results[x]) > 100, results)
keys = sorted(keys, key=lambda x: results[x][0]/sum(results[x]))
for project in keys:
print(f"| {project:20} |", end="")
for extra in args:
res = ""
if project in extra:
v2, v3 = extra[project]
tot = (v2 + v3) / 1000
if tot >= 0.1:
frac = v2 / (v2 + v3)
x = 1 if tot < 10 else 0
y = 1 if frac < 0.1 else 0
res = f"{frac:5.{y}%} of {tot:4.{x}f}k"
print(f" {res:14} |", end="")
print("")
print("| Package name | 2020 (partial) | 2019 | 2018 |")
print("|----------------------|----------------|----------------|----------------|")
show(results2020, results2019, results2018)
| Python2vs3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # In Class Notebook, Week 03
# You can always paste the URL of this notebook (https://github.com/UIUC-iSchool-DataViz/is445_spring2022/blob/master/week03/inClass_week03.ipynb ) into the nbviewer interface for a plain-text rendering:
#
# https://kokes.github.io/nbviewer.js/viewer.html
# I am typing a thing here!
# import usual libraries
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
from PIL import Image
# Fix today's Stictch link!
img = Image.open("/Users/jillnaiman/Downloads/stitch_reworked.png")
#Image.open("/Users/jillnaiman/Downloads/stitch_reworked.png").convert('RGB')
type(img)
imgdata = np.array(img)
type(imgdata)
#imgdata
imgdata.shape
plt.imshow(imgdata)
np.unique(imgdata)
# unwrap my image data into pixel color values (RGBA), R=red channel, G=green channel, B=blue, A=alpha (opacity)
imgdata.reshape(-1,imgdata.shape[2]) #-1 is saying figure it out reshape function, based on that I want data in 4's
# +
# imgdata.reshape?
# +
# np.reshape?
# -
imgdata.shape[2]
np.unique(imgdata.reshape(-1,imgdata.shape[2]),axis=0) # so look for unique values along all pixels
# +
# np.unique?
# -
imgdata.reshape(-1,imgdata.shape[2]).shape
plt.imshow(imgdata)
plt.show()
fig, ax = plt.subplots()
ax
fig, ax = plt.subplots(figsize=(5,5)) # width, height
ax.imshow(imgdata)
plt.show()
fig, ax = plt.subplots(figsize=(5,5))
ax.set_facecolor('gray')
plt.show()
fig, ax = plt.subplots(figsize=(5,5))
ax.set_facecolor('gray')
ax.imshow(imgdata)
plt.show()
red_good_mask = imgdata[:,:,0] == 255
red_good_mask
np.unique(imgdata.reshape(-1,imgdata.shape[2]),axis=0) # so look for unique values along all pixels
# +
red_good_mask = imgdata[:,:,0] == 255
green_good_mask = imgdata[:,:,1] == 255
blues_good_mask = imgdata[:,:,2] == 255
alpha_good_mask = imgdata[:,:,3] == 255
pixel_good_mask = red_good_mask & green_good_mask & blues_good_mask & alpha_good_mask
# -
good_pixels = imgdata[pixel_good_mask]
good_pixels
# how many "good" pixels?
ngood = len(good_pixels)
ngood
imgdata_good = imgdata.copy() # copy of original image
imgdata_good[~pixel_good_mask] = 0 # all other pixels outside of "good" mask
fig, ax = plt.subplots(figsize=(5,5))
ax.set_facecolor('gray')
ax.imshow(imgdata_good)
plt.show()
np.unique(imgdata.reshape(-1,imgdata.shape[2]),axis=0) # so look for unique values along all pixels
# So some of the errors were missing "=" and also missing () around each interior boolean statement.
pixel_mask_bad = (imgdata[:,:,0]==126) & (imgdata[:,:,1]==22) & (imgdata[:,:,2]==33) & (imgdata[:,:,3]==255)
nbad = len(imgdata[pixel_mask_bad])
nbad
imgdata_bad = imgdata.copy()
imgdata_bad[~pixel_mask_bad] = 0
fig, ax = plt.subplots(figsize=(5,5))
ax.set_facecolor('gray')
ax.imshow(imgdata_bad)
plt.show()
total = ngood+nbad
badness = nbad/total
goodness = ngood/total
print(badness,goodness)
# ## RGB decomposition for the homework
fig, ax = plt.subplots(figsize=(10,5))
ax.hist(imgdata[:,:,0]) # how many pixels have each of the 3 different channel values
plt.show()
imgdata[:,:,0].shape
fig, ax = plt.subplots(figsize=(10,5))
ax.hist(imgdata[:,:,0].flatten()) # how many pixels have each of the 3 different channel values
plt.show()
imgdata[:,:,0].flatten()
fig, ax = plt.subplots(figsize=(10,5))
ax.hist(imgdata[:,:,0].flatten(),bins=256) # how many pixels have each of the 3 different channel values
plt.show()
# +
# ax.hist?
# +
fig, ax = plt.subplots(figsize=(10,5))
ax.hist(imgdata[:,:,0].flatten(),bins=256) # how many pixels have each of the 3 different channel values
ax.set_xlabel('Channel Bins (R)')
ax.set_ylabel('Number of pixels in each bin')
plt.show()
# +
fig, ax = plt.subplots(figsize=(15,3))
ax.hist(imgdata[:,:,0].flatten(),bins=256) # 0=Red channel
ax.hist(imgdata[:,:,1].flatten(),bins=256) # 1=Green channel
ax.set_xlabel('Channel Bins (R)')
ax.set_ylabel('Number of pixels in each bin')
plt.show()
# +
fig, ax = plt.subplots(figsize=(15,3))
ax.hist(imgdata[:,:,0].flatten(),bins=256,color='red') # 0=Red channel
ax.hist(imgdata[:,:,1].flatten(),bins=256,color='green') # 1=Green channel
ax.set_xlabel('Channel Bins (R)')
ax.set_ylabel('Number of pixels in each bin')
plt.show()
# -
fig, ax = plt.subplots(3,1, figsize=(15,5)) # now we have 3 rows and 1 column
ax
ax[0]
# +
fig, ax = plt.subplots(3,1, figsize=(15,5)) # now we have 3 rows and 1 column
ax[0].hist(imgdata[:,:,0].flatten(),bins=256,color='red') # 0=Red channel
ax[1].hist(imgdata[:,:,1].flatten(),bins=256,color='green') #1=Green channel
ax[2].hist(imgdata[:,:,2].flatten(),bins=256,color='blue') #2=Blue channel
plt.show()
# +
fig, ax = plt.subplots(4,1, figsize=(15,5)) # now we have 3 rows and 1 column
ax[0].hist(imgdata[:,:,0].flatten(),bins=256,color='red') # 0=Red channel
ax[1].hist(imgdata[:,:,1].flatten(),bins=256,color='green') #1=Green channel
ax[2].hist(imgdata[:,:,2].flatten(),bins=256,color='blue') #2=Blue channel
ax[3].hist(imgdata[:,:,3].flatten(),bins=256,color='black') # 3=Alpha channel
plt.show()
# +
fig, ax = plt.subplots(4,1, figsize=(15,12)) # now we have 3 rows and 1 column
ax[0].hist(imgdata[:,:,0].flatten(),bins=256,color='red') # 0=Red channel
ax[1].hist(imgdata[:,:,1].flatten(),bins=256,color='green') #1=Green channel
ax[2].hist(imgdata[:,:,2].flatten(),bins=256,color='blue') #2=Blue channel
ax[3].hist(imgdata[:,:,3].flatten(),bins=256,color='black') # 3=Alpha channel
# label x-axis
ax[0].set_xlabel('Red channel bins')
ax[1].set_xlabel('Green channel bins')
ax[2].set_xlabel('Blue channel bins')
ax[3].set_xlabel('Alpha channel bins')
ax[0].set_ylabel('# of pixels')
ax[1].set_ylabel('# of pixels')
ax[2].set_ylabel("# of pixels")
ax[3].set_ylabel('# of pixels')
plt.show()
# +
fig, ax = plt.subplots(4,1, figsize=(15,12)) # now we have 3 rows and 1 column
ax[0].hist(imgdata[:,:,0].flatten(),bins=256,color='red') # 0=Red channel
ax[1].hist(imgdata[:,:,1].flatten(),bins=256,color='green') #1=Green channel
ax[2].hist(imgdata[:,:,2].flatten(),bins=256,color='blue') #2=Blue channel
ax[3].hist(imgdata[:,:,3].flatten(),bins=256,color='black') # 3=Alpha channel
# label x-axis
ax[0].set_xlabel('Red channel bins')
ax[1].set_xlabel('Green channel bins')
ax[2].set_xlabel('Blue channel bins')
ax[3].set_xlabel('Alpha channel bins')
for i in range(4): ax[i].set_ylabel('# of pixels')
plt.show()
# +
fig, ax = plt.subplots(5,1, figsize=(15,15)) # now we have 3 rows and 1 column
ax[0].hist(imgdata[:,:,0].flatten(),bins=256,color='red') # 0=Red channel
ax[1].hist(imgdata[:,:,1].flatten(),bins=256,color='green') #1=Green channel
ax[2].hist(imgdata[:,:,2].flatten(),bins=256,color='blue') #2=Blue channel
ax[3].hist(imgdata[:,:,3].flatten(),bins=256,color='black') # 3=Alpha channel
# label x-axis
ax[0].set_xlabel('Red channel bins')
ax[1].set_xlabel('Green channel bins')
ax[2].set_xlabel('Blue channel bins')
ax[3].set_xlabel('Alpha channel bins')
for i in range(4): ax[i].set_ylabel('# of pixels')
ax[4].imshow(imgdata)
plt.show()
# -
# ## Some fancy layout ideas
# +
fig = plt.figure(figsize=(15,8))
gs = fig.add_gridspec(3,2) # three rows (3 color histograms), 2 columns (histograms on the left, fig on right)
ax1 = fig.add_subplot(gs[0,0]) # upper left, plot red
ax1.hist(imgdata[:,:,0].flatten(),bins=256, color='red')
ax1.set_xlabel('R channel')
ax2 = fig.add_subplot(gs[1,0]) # middle row (1), first column (0) -- green
ax2.hist(imgdata[:,:,1].flatten(),bins=256,color="green")
ax2.set_xlabel('G channel')
ax3 = fig.add_subplot(gs[2,0]) # last row (2), first column (0) -- blue
ax3.hist(imgdata[:,:,2].flatten(),bins=256,color="blue")
ax3.set_xlabel('B channel')
# fancy part -- add in the image axes for all of the rows, and the 2nd column
ax4 = fig.add_subplot(gs[:,1]) # all rows (:), 2nd column (1) for the image
ax4.imshow(imgdata)
plt.show()
# -
# Current mystery -- why is the 0 bin so tiny?
# +
fig = plt.figure(figsize=(15,8))
gs = fig.add_gridspec(3,2) # three rows (3 color histograms), 2 columns (histograms on the left, fig on right)
nbins = 257
ax1 = fig.add_subplot(gs[0,0]) # upper left, plot red
ax1.hist(imgdata[:,:,0].flatten(),bins=nbins, color='red')
ax1.set_xlabel('R channel')
ax2 = fig.add_subplot(gs[1,0]) # middle row (1), first column (0) -- green
ax2.hist(imgdata[:,:,1].flatten(),bins=nbins,color="green")
ax2.set_xlabel('G channel')
ax3 = fig.add_subplot(gs[2,0]) # last row (2), first column (0) -- blue
ax3.hist(imgdata[:,:,2].flatten(),bins=nbins,color="blue")
ax3.set_xlabel('B channel')
# fancy part -- add in the image axes for all of the rows, and the 2nd column
ax4 = fig.add_subplot(gs[:,1]) # all rows (:), 2nd column (1) for the image
ax4.imshow(imgdata)
plt.show()
# +
# fig.add_gridspec?
# +
#fig.add_gridspec(nrows, ncols,)
| week03/.ipynb_checkpoints/inClass_week03-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="pHIJVqHsh4An"
# # GPT-J-6B Inference Demo
#
# <a href="http://colab.research.google.com/github/kingoflolz/mesh-transformer-jax/blob/master/colab_demo.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
#
# This notebook demonstrates how to run the [GPT-J-6B model](https://github.com/kingoflolz/mesh-transformer-jax/#GPT-J-6B). See the link for more details about the model, including evaluation metrics and credits.
# + [markdown] id="8CMw_dSQKfhT"
# ## Install Dependencies
#
# First we download the model and install some dependencies. This step takes at least 5 minutes (possibly longer depending on server load).
#
# !!! **Make sure you are using a TPU runtime!** !!!
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="n7xAFw-LOYfe" outputId="7bd7fd83-a40c-41a2-8a14-9a11fc2f150c"
# !apt install zstd
# the "slim" version contain only bf16 weights and no optimizer parameters, which minimizes bandwidth and memory
# !time wget -c https://the-eye.eu/public/AI/GPT-J-6B/step_383500_slim.tar.zstd
# !time tar -I zstd -xf step_383500_slim.tar.zstd
# !git clone https://github.com/kingoflolz/mesh-transformer-jax.git
# !pip install -r mesh-transformer-jax/requirements.txt
# jax 0.2.12 is required due to a regression with xmap in 0.2.13
# !pip install mesh-transformer-jax/ jax==0.2.12
# + [markdown] id="aO1UXepF-0Uq"
# ## Setup Model
#
# + id="ex0qJgaueZtJ"
import os
import requests
from jax.config import config
colab_tpu_addr = os.environ['COLAB_TPU_ADDR'].split(':')[0]
url = f'http://{colab_tpu_addr}:8475/requestversion/tpu_driver0.1_dev20210607'
requests.post(url)
# The following is required to use TPU Driver as JAX's backend.
config.FLAGS.jax_xla_backend = "tpu_driver"
config.FLAGS.jax_backend_target = "grpc://" + os.environ['COLAB_TPU_ADDR']
# + [markdown] id="NIgUVdFLe4A8"
# Sometimes the next step errors for some reason, just run it again ¯\\\_(ツ)\_/¯
# + id="-A5IGYSaeze3"
import time
import jax
from jax.experimental import maps
import numpy as np
import optax
import transformers
from mesh_transformer.checkpoint import read_ckpt
from mesh_transformer.sampling import nucleaus_sample
from mesh_transformer.transformer_shard import CausalTransformer
# + id="QAgKq-X2kmba" colab={"base_uri": "https://localhost:8080/", "height": 167, "referenced_widgets": ["5b8a31b3b4034116af0a81d897c3123b", "835ee2070c05443c9fa3ae6fece3fb44", "9e6646f7066341ef9619f5f7e6f7c8b8", "4f803e3d56b74e18997b216bf9d65337", "<KEY>", "<KEY>", "61b0857802184b51abf77039fad681d7", "<KEY>", "<KEY>", "<KEY>", "3ba9fe91731f4ff380cda2183ad72c4d", "<KEY>", "<KEY>", "fb1263f5a3da42e6acab0e9a66551b32", "3426927332884df4b1ec5544ed479c45", "<KEY>", "7197de8a39ab4e4186eabd061c730cf5", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "74672c43909444c592955afd0d727a28", "67f7c3bc59384dc7812d7130c7cde450", "bb7410ff9cb4484da73302993427b90a"]} outputId="041117df-a315-4b95-9caa-26cb870ff3df"
params = {
"layers": 28,
"d_model": 4096,
"n_heads": 16,
"n_vocab": 50400,
"norm": "layernorm",
"pe": "rotary",
"pe_rotary_dims": 64,
"seq": 2048,
"cores_per_replica": 8,
"per_replica_batch": 1,
}
per_replica_batch = params["per_replica_batch"]
cores_per_replica = params["cores_per_replica"]
seq = params["seq"]
params["sampler"] = nucleaus_sample
# here we "remove" the optimizer parameters from the model (as we don't need them for inference)
params["optimizer"] = optax.scale(0)
mesh_shape = (jax.device_count() // cores_per_replica, cores_per_replica)
devices = np.array(jax.devices()).reshape(mesh_shape)
maps.thread_resources.env = maps.ResourceEnv(maps.Mesh(devices, ('dp', 'mp')))
tokenizer = transformers.GPT2TokenizerFast.from_pretrained('gpt2')
# + [markdown] id="yFgRkUgfiNdA"
# Here we create the network and load the parameters from the downloaded files. Expect this to take around 5 minutes.
# + colab={"base_uri": "https://localhost:8080/"} id="lwNETD2Uk8nu" outputId="a659285b-5b46-4ddf-d65c-fbdad0594b86"
total_batch = per_replica_batch * jax.device_count() // cores_per_replica
network = CausalTransformer(params)
network.state = read_ckpt(network.state, "step_383500/", devices.shape[1])
network.state = network.move_xmap(network.state, np.zeros(cores_per_replica))
# + [markdown] id="A-eT7Sw6if4J"
# ## Run Model
#
# Finally, we are ready to infer with the model! The first sample takes around a minute due to compilation, but after that it should only take about 10 seconds per sample.
#
# Feel free to mess with the different sampling parameters (top_p and temp), as well as the length of the generations (gen_len, causes a recompile when changed).
#
# You can also change other things like per_replica_batch in the previous cells to change how many generations are done in parallel. A larger batch has higher latency but higher throughput when measured in tokens generated/s. This is useful for doing things like best-of-n cherry picking.
#
# *Tip for best results: Make sure your prompt does not have any trailing spaces, which tend to confuse the model due to the BPE tokenization used during training.*
# + id="V85FvxdRAGdQ"
# allow text wrapping in generated output: https://stackoverflow.com/a/61401455
from IPython.display import HTML, display
def set_css():
display(HTML('''
<style>
pre {
white-space: pre-wrap;
}
</style>
'''))
get_ipython().events.register('pre_run_cell', set_css)
# + colab={"base_uri": "https://localhost:8080/"} id="ZVzs2TYlvYeX" outputId="80c8b1e5-0d1c-4799-d682-4a5be0c038a1"
def infer(context, top_p=0.9, temp=1.0, gen_len=512):
tokens = tokenizer.encode(context)
provided_ctx = len(tokens)
pad_amount = seq - provided_ctx
padded_tokens = np.pad(tokens, ((pad_amount, 0),)).astype(np.uint32)
batched_tokens = np.array([padded_tokens] * total_batch)
length = np.ones(total_batch, dtype=np.uint32) * len(tokens)
start = time.time()
output = network.generate(batched_tokens, length, gen_len, {"top_p": np.ones(total_batch) * top_p, "temp": np.ones(total_batch) * temp})
samples = []
decoded_tokens = output[1][0]
for o in decoded_tokens[:, :, 0]:
samples.append(f"\033[1m{context}\033[0m{tokenizer.decode(o)}")
print(f"completion done in {time.time() - start:06}s")
return samples
print(infer("EleutherAI is")[0])
# + colab={"base_uri": "https://localhost:8080/"} id="nvlAK6RbCJYg" outputId="c0525611-ecc8-422e-d8e3-24b3a2159083"
#@title { form-width: "300px" }
top_p = 0.9 #@param {type:"slider", min:0, max:1, step:0.1}
temp = 1 #@param {type:"slider", min:0, max:1, step:0.1}
context = """In a shocking finding, scientist discovered a herd of unicorns living in a remote, previously unexplored valley, in the Andes Mountains. Even more surprising to the researchers was the fact that the unicorns spoke perfect English."""
print(infer(top_p=top_p, temp=temp, gen_len=512, context=context)[0])
| colab_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Programacion Orientada a Objetos
# La Programación Orientada a Objetos viene de una filosofía o forma de pensar que es la Orientación a Objetos y esto surge a partir de los problemas que necesitamos plasmar en código.
#
# Es analizar un problema en forma de objetos para después llevarlo a código, eso es la Orientación a Objetos.
#
# Un paradigma es una teoría que suministra la base y modelo para resolver problemas. La paradigma de Programación Orientada a Objetos se compone de 4 elementos:
# * Clases: Agrupacion de objetos
# * Atributos: Caracteristicas del objeto (Propiedades)
# * Métodos: Acciones del objeto o acciones que podemos realizar con los atributos (funciones)
# * Objetos:
# La programación orientada a objetos tiene cuatro características principales:<br>
#
# <b>Encapsulamiento:</b>
# * Quiere decir que oculta datos mediante código.<br>
#
# <b>Abstracción:</b>
# * Es como se pueden representar los objetos en modo de código.<br>
#
# <b>Herencia:</b>
# * Es donde una clase nueva se crea a partir de una clase existente.<br>
#
# <b>Polimorfismo:</b>
# * Se refiere a la propiedad por la que es posible enviar mensajes sintácticamente iguales a objetos de tipos distintos.
# <img src=https://static.platzi.com/media/user_upload/Captura-274ae063-55d1-4944-9280-f1eda1ca04e9.jpg>
# ## Objetivos
# * Entender el funcionamiento de la POO.
# * Entender como medir la eficiencia temporal y espacial de nuestros algoritmos
# * Entender como y porque debemos graficar.
# * Aprender a resolver problemas de búsqueda, ordenación y optimización.
# La clave para entender la programación orientada a objetos es pensar en objetos
# como agrupaciones de datos y los métodos que operan en dichos datos.
#
# Por ejemplo, podemos representar a una persona con propiedades como nombre,
# edad, género, etc. y los comportamientos de dicha persona como caminar, cantar,
# comer, etc. De la misma manera podemos representar unos audífonos con propiedades
# como su marca, tamaño, color, etc. y sus comportamientos como reproducir música,
# pausar y avanzar a la siguiente canción.
#
# Puesto de otra manera, la programación orientada a objetos nos permite modelar
# cosas reales y concretas del mundo y sus relaciones con otros objetos.
# ## Diagramas de Modelado
# <b>OMT Object Modeling Techniques</b> año `1991`: <br>
# * Es una metodología para el análisis orientado a objetos.
# * Se encuentra descontinuado
#
# <b>UML - Unified Modeling Language</b>: año `1997` <br>
# * Lenguaje de Modelado Unificado.
# * Tomó las bases y técnicas de OMT unificándolas.
# * Tenemos más opciones de diagramas como lo son Clases, Casos de Uso, Objetos, Actividades, Iteración, Estados, Implementación.
# ## Class
# Las estructuras primitivas con las que hemos trabajado hasta ahora nos permiten
# definir cosas sencillas, como:
# * el costo de algo,
# * el nombre de un usuario
# * las veces que debe correr un bucle, etc.
#
# Sin embargo, existen ocasiones cuando necesitamos definir estructuras más complejas, por ejemplo un hotel.
# Podríamos utilizar dos listas: una para definir los cuartos y una segunda para definir
# si el cuarto se encuentra ocupado o no.
class Hotel:
pass
Hotel
print(type(2))
print(type('Hotel'))
print(type(Hotel))
print(Hotel)
# Difiero un poco con la definición que das de Clase. Porque en si, La clase contiene o está formada por atributos y métodos, pero en sí, es un modelo, un molde que representa de forma abstracta un objeto del mundo real. Y lo otro es que podemos tener en nuestro código clases sin atributos pero con metodos, y clases con solo atributos sin métodos.
#
# Por ejemplo:
#
# Clases sin metodos pero con atributos: Son clases usadas por otras. Puede una clase que almacene valores constantes, como PI, un valor de IVA etc.
#
# Clases con métodos pero sin atributos Son clases que puede ser usadas como utilitarias para realizar calculos o manejar algun string. Por ejemplo: Calcular distancia entre dos coordenadas, calcular valor total a pagar dados el subtotal, descuento, iva, oferta o un simple metodos para valdiar si un objeto tiene o no ciertos atributos y valores.
# +
class Hotel:
def __init__(self, numero_maximo_de_huespedes, lugares_de_estacionamiento):
self.numero_maximo_de_huespedes = numero_maximo_de_huespedes
self.lugares_de_estacionamiento = lugares_de_estacionamiento
self.huespedes = 0
hotel_hvca = Hotel(numero_maximo_de_huespedes=50, lugares_de_estacionamiento=20)
print(hotel_hvca.numero_maximo_de_huespedes)
# -
# +
class Hotel:
def anadir_huespedes(self, cantidad_de_huespedes):
self.huespedes += cantidad_de_huespedes
def checkout(self, cantidad_de_huespedes):
self.huespedes -= cantidad_de_huespedes
def ocupacion_total(self):
return self.huespedes
hotel = Hotel(50, 20)
hotel.anadir_huespedes(3)
hotel.checkout(1)
hotel.ocupacion_total()
# +
class Coordenada:
def __init__(self, x, y):
self.x = x
self.y = y
def distancia(self, otra_coordendada):
x_diff = (self.x - otra_coordendada.x)**2
y_diff = (self.y - otra_coordendada.y)**2
return (x_diff + y_diff)**0.5
if __name__ == '__main__':
coord_1 = Coordenada(3, 30)
coord_2 = Coordenada(4, 8)
#print(coord_1.distancia(coord_2))
print(isinstance(3, Coordenada))
# -
# +
class Hotel:
def __init__(self, numero_maximo_de_huespedes, lugares_de_estacionamiento):
self.numero_maximo_de_huespedes = numero_maximo_de_huespedes
self.lugares_de_estacionamiento = lugares_de_estacionamiento
self.huespedes = 0
def anadir_huespedes(self, cantidad_de_huespedes):# --> Operaciones entro de clase Hotel
self.huespedes += cantidad_de_huespedes # Aumentar huespedes
def checkout(self, cantidad_de_huespedes):# --> Operaciones entro de clase Hotel
self.huespedes -= cantidad_de_huespedes # Disminuir huespeder
def ocupacion_total(self):# --> Operaciones entro de clase Hotel
return self.huespedes # Calcula huespedes actuales
hotel = Hotel(numero_maximo_de_huespedes=50, lugares_de_estacionamiento=20)
print(f'Numero de huespedes maximo')
print(hotel.numero_maximo_de_huespedes)
print(f'Numero de estacionamiento maximo')
print(hotel.lugares_de_estacionamiento)
# hotel = Hotel(50, 20) Se puede modificar la cantidad maxima de huespedes dado el caso.
## --- Flujo de informacion --- ##
print('Ingresan 7 huespedes, Marzo 24')
print(f'Huespedes checkin: 7')
hotel.anadir_huespedes(7) # --> Operaciones dentro de class Hotel, hacer llamado con hotel. IMPORTANTE
# verificar y entender cuales son las funciones existentes dentro de class Hotel
# para saber que operaciones pueden hacer en esta.
print('Salen 3 huespedes, Abril 2')
print(f'Huespedes checkout:', 3 )
hotel.checkout(3)
print(f'Huespedes actuales:', hotel.huespedes)
# -
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from io import BytesIO
data = "1, 2, 3\n4, 5, 6"
np.genfromtxt(BytesIO(data.encode('utf-8')), delimiter=",")
data = " 1 2 3\n 4 5 67\n890123 4"
np.genfromtxt(BytesIO(data.encode('utf-8')), delimiter=3)
data = "123456789\n 4 7 9\n 4567 9"
np.genfromtxt(BytesIO(data.encode('utf-8')), delimiter=(4, 3, 2))
data = "1, abc , 2\n 3, xxx, 4"
np.genfromtxt(BytesIO(data.encode('utf-8')), delimiter=',', dtype='|S5', autostrip=True)
data = """#
# Skip me !
# Skip me too !
1, 2
3, 4
5, 6 #This is the third line of the data
7, 8
# And here comes the last line
9, 0
"""
np.genfromtxt(BytesIO(data.encode('utf-8')), comments="#", delimiter=",")
data = '\n'.join(str(i) for i in range(10))
np.genfromtxt(BytesIO(data.encode('utf-8')),)
np.genfromtxt(BytesIO(data.encode('utf-8')), skip_header=3, skip_footer=5)
data = "1 2 3\n4 5 6"
np.genfromtxt(BytesIO(data.encode('utf-8')), names="a, b, c", usecols=("a", "c"))
np.genfromtxt(BytesIO(data.encode('utf-8')), names="a, b, c", usecols=("a, c"))
data = BytesIO("1 2 3\n 4 5 6".encode('utf-8'))
np.genfromtxt(data, dtype=[(_, int) for _ in "abc"])
data = BytesIO("1 2 3\n 4 5 6".encode('utf-8'))
np.genfromtxt(data, names="A, B, C")
data = BytesIO("So it goes\n#a b c\n1 2 3\n 4 5 6".encode('utf-8'))
np.genfromtxt(data, skip_header=1, names=True)
data = BytesIO("1 2 3\n 4 5 6".encode('utf-8'))
ndtype = [('a', int), ('b', int), ('c', int)]
names = ['A', 'B', 'C']
np.genfromtxt(data, names=names, dtype=ndtype)
data = BytesIO("1 2 3\n 4 5 6".encode('utf-8'))
np.genfromtxt(data, dtype=(int, float, int))
data = BytesIO("1 2 3\n 4 5 6".encode('utf-8'))
np.genfromtxt(data, dtype=(int, float, int), names='a')
data = BytesIO("1 2 3\n 4 5 6".encode('utf-8'))
np.genfromtxt(data, dtype=(int, float, int), defaultfmt='var_%02i')
convertfunc = lambda x: float(x.strip('%'))/100.
data = '1, 2.3%, 45.\n6, 78.9%, 0'
np.genfromtxt(BytesIO(data.encode('utf-8')), delimiter=',', names=names)
# +
# np.genfromtxt(BytesIO(data), delimiter=',', names=names, converters={1: convertfunc})
# -
np.genfromtxt(BytesIO(data), delimiter=',', names=names, converters={'p': convertfunc})
data = "1, , 3\n 4, 5, 6"
convert = lambda x: float(x.strip() or -999)
np.genfromtxt(BytesIO(data.encode('utf-8')), delimiter=',', converters={1: convert})
data = "N/A, 2, 3\n4, ,???"
kwargs = dict(delimiter=",",
dtype=int,
names="a,b,c",
missing_values={0:"N/A", 'b':" ", 2:"???"},
filling_values={0:0, 'b':0, 2:-999})
np.genfromtxt(BytesIO(data.encode('utf-8')), **kwargs)
| np_io.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
#default_exp adapters.prophet
# -
# # Adapters for Prophet
# +
#export
import sys
from copy import deepcopy
import numpy as np
import statsmodels.api as sm
import pandas as pd
from statsforecast.arima import AutoARIMA
if sys.version_info.minor != 6 or (sys.platform not in ['win32', 'cygwin']):
try:
from prophet import Prophet
except ModuleNotFoundError as e:
msg = (
'{e}. To use prophet adapters you have to install '
'prophet. Please run `pip install prophet`. '
'Note that it is recommended to install prophet '
'using conda environments due to dependencies.'
)
raise ModuleNotFoundError(msg) from e
elif sys.version_info.minor == 6 and (sys.platform in ['win32', 'cygwin']):
try:
from fbprophet import Prophet
except ModuleNotFoundError as e:
msg = (
'{e}. To use prophet adapters you have to install '
'fbprophet. Please run `pip install fbprophet`. '
'Note that it is recommended to install prophet '
'using conda environments due to dependencies.'
)
raise ModuleNotFoundError(msg) from e
# -
# ## Arima
#export
class AutoARIMAProphet(Prophet):
"""Returns best ARIMA model using external variables created
by the Prophet interface.
This class receives as parameters the same as prophet.Prophet
and statsforecast.arima.AutoARIMA.
If your pipeline uses Prophet you can simply replace Prophet
with AutoARIMAProphet and you'll be using AutoARIMA instead
of Prophet.
"""
def __init__(self,
growth='linear',
changepoints=None,
n_changepoints=25,
changepoint_range=0.8,
yearly_seasonality='auto',
weekly_seasonality='auto',
daily_seasonality='auto',
holidays=None,
seasonality_mode='additive',
seasonality_prior_scale=10.0,
holidays_prior_scale=10.0,
changepoint_prior_scale=0.05,
mcmc_samples=0,
interval_width=0.80,
uncertainty_samples=1000,
stan_backend=None,
d=None,
D=None,
max_p=5,
max_q=5,
max_P= 2,
max_Q= 2,
max_order= 5,
max_d=2,
max_D=1,
start_p=2,
start_q=2,
start_P=1,
start_Q=1,
stationary=False,
seasonal=True,
ic='aicc',
stepwise=True,
nmodels=94,
trace=False,
approximation=False,
method=None,
truncate=None,
test='kpss',
test_kwargs=None,
seasonal_test='seas',
seasonal_test_kwargs=None,
allowdrift=False,
allowmean=False,
blambda=None,
biasadj=False,
parallel=False,
num_cores=2,
period=1):
Prophet.__init__(self,
growth,
changepoints,
n_changepoints,
changepoint_range,
yearly_seasonality,
weekly_seasonality,
daily_seasonality,
holidays,
seasonality_mode,
seasonality_prior_scale,
holidays_prior_scale,
changepoint_prior_scale,
mcmc_samples,
interval_width,
uncertainty_samples,
stan_backend)
self.arima = AutoARIMA(d=d,
D=D,
max_p=max_p,
max_q=max_q,
max_P=max_P,
max_Q=max_Q,
max_order=max_order,
max_d=max_d,
max_D=max_D,
start_p=start_p,
start_q=start_q,
start_P=start_P,
start_Q=start_Q,
stationary=stationary,
seasonal=seasonal,
ic=ic,
stepwise=stepwise,
nmodels=nmodels,
trace=trace,
approximation=approximation,
method=method,
truncate=truncate,
test=test,
test_kwargs=test_kwargs,
seasonal_test=seasonal_test,
seasonal_test_kwargs=seasonal_test_kwargs,
allowdrift=allowdrift,
allowmean=allowmean,
blambda=blambda,
biasadj=biasadj,
parallel=parallel,
num_cores=num_cores,
period=period)
def fit(self, df, disable_seasonal_features=True, **kwargs):
"""Fit the AutoARIMAProphet model.
Parameters
----------
df: pd.DataFrame containing the history. Must have columns ds (date
type) and y, the time series.
disable_seasonal_features: bool
Wheter disable seasonal features generated by Prophet.
kwargs: Additional arguments.
Returns
-------
The AutoARIMAProphet object.
"""
if self.history is not None:
raise Exception('Prophet object can only be fit once. '
'Instantiate a new object.')
if ('ds' not in df) or ('y' not in df):
raise ValueError(
'Dataframe must have columns "ds" and "y" with the dates and '
'values respectively.'
)
history = df[df['y'].notnull()].copy()
if history.shape[0] < 2:
raise ValueError('Dataframe has less than 2 non-NaN rows.')
self.history_dates = pd.to_datetime(pd.Series(df['ds'].unique(), name='ds')).sort_values()
history = self.setup_dataframe(history, initialize_scales=True)
self.history = history
self.set_auto_seasonalities()
seasonal_features, prior_scales, component_cols, modes = (
self.make_all_seasonality_features(history))
self.train_component_cols = component_cols
self.component_modes = modes
self.fit_kwargs = deepcopy(kwargs)
if disable_seasonal_features:
seas = tuple(self.seasonalities.keys())
seasonal_features = seasonal_features.loc[:,~seasonal_features.columns.str.startswith(seas)]
self.xreg_cols = seasonal_features.columns
y = history['y'].values
X = seasonal_features.values if not seasonal_features.empty else None
self.arima = self.arima.fit(y=y, X=X)
return self
def predict(self, df=None):
"""Predict using the AutoARIMAProphet model.
Parameters
----------
df: pd.DataFrame with dates for predictions (column ds.
If not provided, predictions are
made on the history.
Returns
-------
A pd.DataFrame with the forecast components.
"""
if self.history is None:
raise Exception('Model has not been fit.')
if df is None:
df = self.history.copy()
else:
if df.shape[0] == 0:
raise ValueError('Dataframe has no rows.')
df = self.setup_dataframe(df.copy())
seasonal_features = self.make_all_seasonality_features(df)[0].loc[:, self.xreg_cols]
ds_forecast = set(df['ds'])
h = len(ds_forecast - set(self.history['ds']))
if h > 0:
X = seasonal_features.values[-h:] if not seasonal_features.empty else None
df2 = self.arima.predict(h=h,
X=X,
level=int(100*self.interval_width))
else:
df2 = pd.DataFrame()
if len(ds_forecast) > h:
in_sample = self.arima.predict_in_sample(level=int(100*self.interval_width))
df2 = pd.concat([in_sample, df2]).reset_index(drop=True)
yhat = df2.pop('mean')
df2.columns = ['yhat_lower', 'yhat_upper']
df2.insert(0, 'yhat', yhat)
df2.insert(0, 'ds', df['ds'])
return df2
# ### Peyton Manning example
df = pd.read_csv('https://raw.githubusercontent.com/facebook/prophet/main/examples/example_wp_log_peyton_manning.csv')
# + [markdown] tags=[]
# #### Without additional info
# -
# Usually, a Prophet pipeline without external regressors looks like this.
m = Prophet()
m.fit(df)
future = m.make_future_dataframe(365)
forecast = m.predict(future)
fig = m.plot(forecast)
# With the class `AutoARIMAProphet` you can simply replace `Prophet` and you'll be training an `auto_arima` model without changing the pipeline.
# %%capture
m = AutoARIMAProphet()
m.fit(df)
future = m.make_future_dataframe(365)
forecast = m.predict(future)
fig = m.plot(forecast)
# ### With exogenous regressors provided by prophet
# Usually `Prophet` pipelines include the usage of external regressors such as holidays.
playoffs = pd.DataFrame({
'holiday': 'playoff',
'ds': pd.to_datetime(['2008-01-13', '2009-01-03', '2010-01-16',
'2010-01-24', '2010-02-07', '2011-01-08',
'2013-01-12', '2014-01-12', '2014-01-19',
'2014-02-02', '2015-01-11', '2016-01-17',
'2016-01-24', '2016-02-07']),
'lower_window': 0,
'upper_window': 1,
})
superbowls = pd.DataFrame({
'holiday': 'superbowl',
'ds': pd.to_datetime(['2010-02-07', '2014-02-02', '2016-02-07']),
'lower_window': 0,
'upper_window': 1,
})
holidays = pd.concat((playoffs, superbowls))
# %%capture
m = Prophet(holidays=holidays)
m.add_country_holidays(country_name='US')
m.fit(df)
future = m.make_future_dataframe(365)
forecast = m.predict(future)
fig = m.plot(forecast)
# The class `AutoARIMAProphet` allows you to handle these scenarios to fit an `auto_arima` model with exogenous variables.
# %%capture
m = AutoARIMAProphet(holidays=holidays)
m.add_country_holidays(country_name='US')
m.fit(df)
future = m.make_future_dataframe(365)
forecast = m.predict(future)
fig = m.plot(forecast)
| nbs/adapters.prophet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %matplotlib inline
# %autoreload 2
from IPython import display
# +
import torch
import sys
from torch import nn, optim
from torch.autograd.variable import Variable
from torchvision import transforms, datasets
USE_CUDA = torch.cuda.is_available()
device = torch.device("cuda" if USE_CUDA else "cpu")
print(USE_CUDA)
# -
# this is a utility file developed by the author
from utils import Logger
# the following will download and proces the MNIST dataset: http://yann.lecun.com/exdb/mnist/
def mnist_data():
compose = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((.5, .5, .5), (.5, .5, .5))
])
out_dir = './dataset'
return datasets.MNIST(root=out_dir, train=True, transform=compose, download=True)
# Load data
data = mnist_data()
# Create loader with data, so that we can iterate over it
data_loader = torch.utils.data.DataLoader(data, batch_size=100, shuffle=True)
# Num batches
num_batches = len(data_loader)
class DiscriminatorNet(torch.nn.Module):
"""
A three hidden-layer discriminative neural network
"""
def __init__(self):
super(DiscriminatorNet, self).__init__()
n_features = 784 # the input size for each image is 28x28=784
n_out = 1 # one dropout layer to prevent overfitting
self.hidden0 = nn.Sequential(
nn.Linear(n_features, 1024),
nn.LeakyReLU(0.2),
nn.Dropout(0.3)
)
self.hidden1 = nn.Sequential(
nn.Linear(1024, 512),
nn.LeakyReLU(0.2),
nn.Dropout(0.3)
)
self.hidden2 = nn.Sequential(
nn.Linear(512, 256),
nn.LeakyReLU(0.2),
nn.Dropout(0.3)
)
self.out = nn.Sequential(
torch.nn.Linear(256, n_out),
torch.nn.Sigmoid()
)
def forward(self, x):
x = self.hidden0(x)
x = self.hidden1(x)
x = self.hidden2(x)
x = self.out(x)
return x
# +
# convert a flattened image into its 2-dimensional representation, and another one that does the opposite
def images_to_vectors(images):
return images.view(images.size(0), 784)
def vectors_to_images(vectors):
return vectors.view(vectors.size(0), 1, 28, 28)
# -
class GeneratorNet(torch.nn.Module):
"""a three hidden-layer generative neural netwrok"""
def __init__(self):
super(GeneratorNet, self).__init__()
n_features = 100
n_out = 784
self.hidden0 = nn.Sequential(
nn.Linear(n_features, 256),
nn.LeakyReLU(0.2)
)
self.hidden1 = nn.Sequential(
nn.Linear(256, 512),
nn.LeakyReLU(0.2)
)
self.hidden2 = nn.Sequential(
nn.Linear(512, 1024),
nn.LeakyReLU(0.2)
)
self.out = nn.Sequential(
nn.Linear(1024, n_out),
nn.Tanh()
)
def forward(self, x):
x = self.hidden0(x)
x = self.hidden1(x)
x = self.hidden2(x)
x = self.out(x)
return x
def noise(size):
"""generates a 1-d vector of gaussian sampled random values"""
n = Variable(torch.randn(size, 100))
return n.to(device)
discriminator = DiscriminatorNet().to(device)
generator = GeneratorNet().to(device)
# +
# optimizers
d_optimizer = optim.Adam(discriminator.parameters(), lr=0.0002)
g_optimizer = optim.Adam(generator.parameters(), lr=0.0002)
# Loss Function
loss = nn.BCELoss()
# +
def ones_target(size):
'''
Tensor containing ones, with shape = size
'''
data = Variable(torch.ones(size, 1))
return data.to(device)
def zeros_target(size):
'''
Tensor containing zeros, with shape = size
'''
data = Variable(torch.zeros(size, 1))
return data.to(device)
# -
def train_discriminator(optimizer, real_data, fake_data):
N = real_data.size(0)
# reset gradients
optimizer.zero_grad()
# 1.1 train on real data
prediction_real = discriminator(real_data)
# calculate error and backpropagate
error_real = loss(prediction_real, ones_target(N))
error_real.backward()
# 1.2 train on fake data
prediction_fake = discriminator(fake_data)
# calculate error and backpropagate
error_fake = loss(prediction_fake, zeros_target(N))
error_fake.backward()
#1.3 update weights with gradients
optimizer.step()
# return error and predictions for real and fake inputs
return error_real + error_fake, prediction_real, prediction_fake
def train_generator(optimizer, fake_data):
N = fake_data.size(0)
# reset gradients
optimizer.zero_grad()
# sample noise and generate fake data
prediction = discriminator(fake_data)
# calculate error and backpropagate
error = loss(prediction, ones_target(N))
error.backward()
# update weights with gradients
optimizer.step()
# return error
return error
num_test_samples = 16
test_noise = noise(num_test_samples)
# +
# create logger instance
logger = Logger(model_name='VGAN', data_name='MNIST')
# Total number of epochs to train
num_epochs = 2
for epoch in range(num_epochs):
for n_batch, (real_batch,_) in enumerate(data_loader):
real_batch = real_batch.to(device)
N = real_batch.size(0)
# train discriminator
readl_data = Variable(images_to_vectors(real_batch))
# generating fake data and detach so gradients are not calculated for generator
fake_data = generator(noise(N)).detach()
# train
d_error, d_pred_real, d_pred_fake = \
train_discriminator(d_optimizer, readl_data, fake_data)
# train generator
#generate fake data
fake_data = generator(noise(N))
# train g
g_error = train_generator(g_optimizer, fake_data)
# log batch error
logger.log(d_error, g_error, epoch, n_batch, num_batches)
# display progress every few batches
if (n_batch) % 100 == 0:
test_images = vectors_to_images(generator(test_noise))
test_images = test_images.data.cpu()
logger.log_images(
test_images, num_test_samples,
epoch, n_batch, num_batches
)
logger.display_status(
epoch, num_epochs, n_batch, num_batches,
d_error, g_error, d_pred_real, d_pred_fake
)
# -
| pytorch_gan_intro/gan_intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sv_utils
import torch
import numpy as np
import pylab as plt
from matplotlib import pyplot
from matplotlib.pyplot import figure
from os.path import dirname, realpath
from singleNeuron import preSpikes, nextSpikes, STDPLIFDensePopulation
# # Filter Visualizer
# El proposito de esta notebook es el poder graficar el peso de los filtros de los distintos layers
# ## Visualizacion de los filtros
# ### Filtros 1er Layer (Sin delay)
# +
# Load and set img size
filters = np.load('../cleanDSNN/results/weight_0.npy')
figure(figsize=(18,18))
print(filters.shape)
n_filters, ix = filters.shape[3], 1
for i in range(n_filters):
# get the filter
f = filters[:, :, :, i]
f = np.squeeze(f)
# specify subplot and turn of axis
ax = pyplot.subplot(n_filters, 4, ix)
ax.set_xticks([])
ax.set_yticks([])
# plot filter channel in grayscale
pyplot.imshow(f, cmap='gray')
ix += 1
# show the figure
pyplot.show()
# -
# ### Filtros 2do Layer (Sin delay)
# +
# Load and set img size
filters = np.load('../cleanDSNN/results/weight_2.npy')
figure(figsize=(18,18))
print(filters.shape)
channels, n_filters, ix = filters.shape[2], filters.shape[3], 1
for i in range(n_filters):
# get the filter
f = filters[:, :, :, i]
f = np.squeeze(f)
for j in range(channels):
# specify subplot and turn of axis
ax = pyplot.subplot(n_filters, channels, ix)
ax.set_xticks([])
ax.set_yticks([])
# plot filter channel in grayscale
pyplot.imshow(f[:, :, j], cmap='gray')
ix += 1
# show the figure
pyplot.show()
# -
# ### Filtros 3er Layer (Sin delay)
# +
# Load and set img size
filters = np.load('../cleanDSNN/results/weight_4.npy')
figure(figsize=(15,15))
print(filters.shape)
channels, n_filters, ix = filters.shape[2], filters.shape[3], 1
for i in range(n_filters):
# get the filter
f = filters[:, :, :, i]
f = np.squeeze(f)
for j in range(channels):
# specify subplot and turn of axis
ax = pyplot.subplot(n_filters, channels, ix)
ax.set_xticks([])
ax.set_yticks([])
# plot filter channel in grayscale
pyplot.imshow(f[:, :, j], cmap='gray')
ix += 1
# show the figure
pyplot.show()
# -
# ### Filtros 1er Layer (Con delay)
# +
# Load and set img size
filters = np.load('../cleanDSNN/results/delayed_weight_0.npy')
figure(figsize=(18,18))
print(filters.shape)
n_filters, ix = filters.shape[3], 1
for i in range(n_filters):
# get the filter
f = filters[:, :, :, i]
f = np.squeeze(f)
# specify subplot and turn of axis
ax = pyplot.subplot(n_filters, 4, ix)
ax.set_xticks([])
ax.set_yticks([])
# plot filter channel in grayscale
pyplot.imshow(f, cmap='gray')
ix += 1
# show the figure
pyplot.show()
# -
# ### Filtros 2do Layer (Con delay)
# +
# Load and set img size
filters = np.load('../cleanDSNN/results/delayed_weight_2.npy')
figure(figsize=(18,18))
print(filters.shape)
channels, n_filters, ix = filters.shape[2], filters.shape[3], 1
for i in range(n_filters):
# get the filter
f = filters[:, :, :, i]
f = np.squeeze(f)
for j in range(channels):
# specify subplot and turn of axis
ax = pyplot.subplot(n_filters, channels, ix)
ax.set_xticks([])
ax.set_yticks([])
# plot filter channel in grayscale
pyplot.imshow(f[:, :, j], cmap='gray')
ix += 1
# show the figure
pyplot.show()
# -
| cleanDSNN/results/old_results/all_nums_less_freq/with_decay/FiltersVisualizer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# DEPENDENCIES AND SETUP
import pandas as pd
# specifying csv path
csv_path_ride = "./Resources/ride_data.csv"
csv_path_city = "./Resources/city_data.csv"
# read_csv reads the csv file
read_csv_ride = pd.read_csv(csv_path_ride)
read_csv_city = pd.read_csv(csv_path_city)
# creating data frame
ride_df = pd.DataFrame(read_csv_ride)
ride_df
ride_df.head()
city_df = pd.DataFrame(read_csv_city)
city_df
city_df.head()
# converting data frame to html file called "ride_data_table.html"
# set index as false
ride_df.to_html('ride_data_table.html',index=False)
# converting data frame to html file called "city_data_table.html"
# set index as false
city_df.to_html('city_data_table.html',index=False)
| BONUS/pyber_csv_files_to_html.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using Db2 11.5.4 RESTful Programming with Data Virtualization
# The following notebook is a brief example of how to use the Db2 11.5.4 RESTful Endpoint service to extend the capabilies of Cloud Pak for Data Virtualization.
#
# You can extend your Cloud Pak for Data system so that application programmers can create Representational State Transfer (REST) endpoints that can be used to interact with the Data Virtualization Service.
#
# Each endpoint is associated with a single SQL statement. Authenticated users of web, mobile, or cloud applications can use these REST endpoints from any REST HTTP client without having to install any Db2 drivers.
#
# The Db2 REST server accepts an HTTP request, processes the request body, and returns results in JavaScript Object Notation (JSON).
#
# The Db2 REST server is pre-installed and running on Docker on server7 (10.1.1.12) in the Demonstration cluster. As a programmer you can communicate with the service on port 50050. Your welcome note includes the external port you can use to interact with the Db2 RESTful Endpoint service directly.
#
# You can find more information about this service at: https://www.ibm.com/support/producthub/db2/docs/content/SSEPGG_11.5.0/com.ibm.db2.luw.admin.rest.doc/doc/c_rest.html.
# ## Finding the Db2 RESTful Endpoint Service API Documentation
# If you are running this notebook from a browser running inside the Cloud Pak for Data cluster, click: http://10.1.1.12:50050/docs
# If you are running this from a browser from your own desktop, check your welcome note for the address of the Db2 RESTful Service at port 50050.
# ## Import the required programming libraries
# The requests library is the minimum required by Python to construct RESTful service calls. The Pandas library is used to format and manipulate JSON result sets as tables.
import requests
import pandas as pd
# ## Create the Header File required for getting an authetication token
# The RESTful call to the Db2 RESTful Endpoint service is contructed and transmitted as JSON. The first part of the JSON structure is the headers that define the content tyoe of the request.
headers = {
"content-type": "application/json"
}
# ## RESTful Host
# The next part defines where the request is sent to. It provides the location of the RESTful service for our calls.
Db2RESTful = "http://10.1.1.12:50050"
# ## API Authentication Service
# Each service has its own path in the RESTful call. For authentication we need to point to the `v1/auth` service.
API_Auth = "/v1/auth"
# ## Authentication
# To authenticate to the RESTful service you must provide the connection information for the database along with the userid and password that you are using to authenticate with. You can also provide an expiry time so that the access token that gets returned will be invalidated after that time period.
body = {
"dbParms": {
"dbHost": "10.1.1.1",
"dbName": "bigsql",
"dbPort": 30797,
"isSSLConnection": False,
"username": "user999",
"password": "<PASSWORD>"
},
"expiryTime": "300m"
}
# ## API Service
# When communicating with the RESTful service, you must provide the name of the service that you want to interact with. In this case the authentication service is */v1/auth*.
try:
response = requests.post("{}{}".format(Db2RESTful,API_Auth), headers=headers, json=body)
except Exception as e:
print("Unable to call RESTful service. Error={}".format(repr(e)))
# A response code of 200 means that the authentication worked properly, otherwise the error that was generated is printed.
print(response)
# The response includes a connection token that is reused throughout the rest of this lab. It ensures secure a connection without requiring that you reenter a userid and password with each request.
if (response.status_code == 200):
token = response.json()["token"]
print("Token: {}".format(token))
else:
print(response.json()["errors"])
# ## Reusing the token in the standard header
# The standard header for all subsequent calls will use this format. It includes the access token.
headers = {
"authorization": f"{token}",
"content-type": "application/json"
}
# ## Executing an SQL Statement
# Executing SQL requires a different service endpoint. In this case we will use "/services/execsql"
API_execsql = "/v1/services/execsql"
# In this example, the code requests that the RESTful function waits until the command is complete.
body = {
"isQuery": True,
"sqlStatement": "SELECT * FROM NETEZZA.STOCK_SYMBOLS WHERE SYMBOL = 'CAT'",
"sync": True
}
try:
response = requests.post("{}{}".format(Db2RESTful,API_execsql), headers=headers, json=body)
except Exception as e:
print("Unable to call RESTful service. Error={}".format(repr(e)))
print(response)
print(response.json())
# Retrieve the results. The Dataframe class converts the json result set into a table. Dataframes can be used to further manipulate results in Python.
display(pd.DataFrame(response.json()['resultSet']))
# ## Use Parameters in a SQL Statement
# Simple parameter passing is also available through the execsql service. In this case we are passing the Stock symbol "CAT" into the query to retrieve the full stock name. Try substituting different symbols and run the REST call again. Symbols like PG, DIS, or MMM.
body = {
"isQuery": True,
"parameters" : {
"1" : "CAT"
},
"sqlStatement": "SELECT * FROM NETEZZA.STOCK_SYMBOLS WHERE SYMBOL = ?",
"sync": True
}
try:
response = requests.post("{}{}".format(Db2RESTful,API_execsql), headers=headers, json=body)
except Exception as e:
print("Unable to call RESTful service. Error={}".format(repr(e)))
response.json()["resultSet"]
display(pd.DataFrame(response.json()['resultSet']))
# ## Generate a Call and don't wait for the results
# Turn sync off to require us to poll for the results. This is especially useful for long running queries.
body = {
"isQuery": True,
"sqlStatement": "SELECT * FROM NETEZZA.STOCK_SYMBOLS WHERE SYMBOL = 'CAT'",
"sync": False
}
try:
response = requests.post("{}{}".format(Db2RESTful,API_execsql), headers=headers, json=body)
except Exception as e:
print("Unable to call RESTful service. Error={}".format(repr(e)))
print(response)
# Retrieve the job id to retrieve the results later.
job_id = response.json()["id"]
print(job_id)
# ## Retrieve Result set using Job ID
# The service API needs to be appended with the Job ID.
API_get = "/v1/services/"
# We can limit the number of rows that we return at a time. Setting the limit to zero means all of the rows are to be returned.
body = {
"limit": 0
}
# Get the results.
try:
response = requests.get("{}{}{}".format(Db2RESTful,API_get,job_id), headers=headers, json=body)
except Exception as e:
print("Unable to call RESTful service. Error={}".format(repr(e)))
# Retrieve the results.
print(response.json()["resultSet"])
display(pd.DataFrame(response.json()["resultSet"]))
# ## Create a Unique RESTful Service
# The most common way of interacting with the service is to fully encapsulate an SQL statement, including any parameters, in a unique RESTful service. This creates a secure separation between the database service and the RESTful programming service. It also allows you to create versions of the same service to make maintenance and evolution of programming models simple and predictable.
API_makerest = "/v1/services"
# Define the SQL that we want in the RESTful call.
body = {"isQuery": True,
"parameters": [
{
"datatype": "VARCHAR(4)",
"name": "@SYMBOL"
}
],
"schema": "STOCK",
"serviceDescription": "Get full name given symbol",
"serviceName": "getstock",
"sqlStatement": "SELECT * FROM NETEZZA.STOCK_SYMBOLS WHERE SYMBOL = @SYMBOL",
"version": "1.0"
}
try:
response = requests.post("{}{}".format(Db2RESTful,API_makerest), headers=headers, json=body)
except Exception as e:
print("Unable to call RESTful service. Error={}".format(repr(e)))
if (response.status_code == 201):
print("Service Created")
else:
print(response.json()['errors'])
# ## Call the new RESTful Service
# Now you can call the RESTful service. In this case we will pass the stock symbol CAT. But like in the previous example you can try rerunning the service call with different stock symbols.
API_runrest = "/v1/services/getstock/1.0"
body = {
"parameters": {
"@SYMBOL": "CAT"
},
"sync": True
}
try:
response = requests.post("{}{}".format(Db2RESTful,API_runrest), headers=headers, json=body)
except Exception as e:
print("Unable to call RESTful service. Error={}".format(repr(e)))
# A response of 200 indicates a successful service call.
print(response)
print(response.json())
# You can now retrieve the result set, convert it into a Dataframe and display the table.
print(response.json())
display(pd.DataFrame(response.json()['resultSet']))
# ## Retreive Service Details
# You can query each service to see its details, including authoritization, input parameters and output results.
API_listrest = "/v1/services/getstock/1.0"
try:
response = requests.get("{}{}".format(Db2RESTful,API_listrest), headers=headers)
except Exception as e:
print("Unable to call RESTful service. Error={}".format(repr(e)))
print(response.json())
# +
print("Service Details:")
print("Service Name: " + response.json()['serviceName'])
print("Service Version: " + response.json()['version'])
print("Service Description: " + response.json()['serviceDescription'])
print("Service Creator: " + response.json()['serviceCreator'])
print("Service Updater: " + response.json()['serviceUpdater'])
print('Users:')
display(pd.DataFrame(response.json()['grantees']['users']))
print('Groups:')
display(pd.DataFrame(response.json()['grantees']['groups']))
print('Roles:')
display(pd.DataFrame(response.json()['grantees']['roles']))
print('')
print('Input Parameters:')
display(pd.DataFrame(response.json()['inputParameters']))
print('Result Set Fields:')
display(pd.DataFrame(response.json()['resultSetFields']))
# -
# ## List Available Services
# You can also list all the user defined services you have access to
API_listrest = "/v1/services"
try:
response = requests.get("{}{}".format(Db2RESTful,API_listrest), headers=headers)
except Exception as e:
print("Unable to call RESTful service. Error={}".format(repr(e)))
display(pd.DataFrame(response.json()['Db2Services']))
# ## Delete a Service
# A single call is also available to delete a service
API_deleteService = "/v1/services"
Service = "/getstock"
Version = "/1.0"
try:
response = requests.delete("{}{}{}{}".format(Db2RESTful,API_deleteService,Service,Version), headers=headers)
except Exception as e:
print("Unable to call RESTful service. Error={}".format(repr(e)))
print (response)
# ## Get Service Logs
# You can easily download service logs. However you must be authorized as the principal administration user to do so.
API_listrest = "/v1/logs"
try:
response = requests.get("{}{}".format(Db2RESTful,API_listrest), headers=headers)
except Exception as e:
print("Unable to call RESTful service. Error={}".format(repr(e)))
if (response.status_code == 200):
myFile = response.content
open('/tmp/logs.zip', 'wb').write(myFile)
print("Downloaded",len(myFile),"bytes.")
else:
print(response.json())
| RESTEndpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="HgljXEAJEcFq"
# <!-- ---
# title: Getting Started
# date: 2021-07-27
# downloads: true
# sidebar: true
# tags:
# - PyTorch-Ignite
# --- -->
#
# # Getting Started
# + [markdown] id="P9VjKOirEcFu"
# Welcome to **PyTorch-Ignite**’s quick start guide that covers the
# essentials of getting a project up and running while walking through
# basic concepts of Ignite. In just a few lines of code, you can get your
# model trained and validated. The complete code can be found at the end
# of this guide.
#
# <!--more-->
# + [markdown] id="1QNvbg3SEcFw"
# ## Prerequisites
#
# This tutorial assumes you are familiar with the:
#
# 1. Basics of Python and deep learning
# 2. Structure of PyTorch code
# + [markdown] id="XTHzzYyoEcFy"
# ## Installation
#
# From `pip`
#
# ``` shell
# pip install pytorch-ignite
# ```
#
# From `conda`
#
# ``` shell
# conda install ignite -c pytorch
# ```
#
# See [here](https://pytorch-ignite.ai/how-to-guides/installation/) for other installation
# options.
# + [markdown] id="DcnSr5sGEcFz"
# ## Code
#
# Import the following:
# + id="Saizk3heEcFz"
import torch
from torch import nn
from torch.utils.data import DataLoader
from torchvision.datasets import MNIST
from torchvision.models import resnet18
from torchvision.transforms import Compose, Normalize, ToTensor
from ignite.engine import Engine, Events, create_supervised_trainer, create_supervised_evaluator
from ignite.metrics import Accuracy, Loss
from ignite.handlers import ModelCheckpoint
from ignite.contrib.handlers import TensorboardLogger, global_step_from_engine
# + [markdown] id="ecMYtJF7OvgT"
# Speed things up by setting [device](https://pytorch.org/docs/stable/tensor_attributes.html#torch.torch.device) to `cuda` if available else `cpu`.
# + id="sdjDKcFhOuQn"
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# + [markdown] id="4r_PUH1yEcF1"
# Define a class of your model or use the predefined ResNet18 model (modified for MNIST) below, instantiate it and move it to device:
# + id="dVSVAT0OEcF1"
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
# Changed the output layer to output 10 classes instead of 1000 classes
self.model = resnet18(num_classes=10)
# Changed the input layer to take grayscale images for MNIST instaed of RGB images
self.model.conv1 = nn.Conv2d(
1, 64, kernel_size=3, padding=1, bias=False
)
def forward(self, x):
return self.model(x)
model = Net().to(device)
# + [markdown] id="DDIW2zedEcF3"
# Now let us define the training and validation datasets (as
# [torch.utils.data.DataLoader](https://pytorch.org/docs/stable/data.html#torch.utils.data.DataLoader))
# and store them in `train_loader` and `val_loader` respectively. We have
# used the [MNIST](https://pytorch.org/vision/stable/datasets.html#mnist)
# dataset for ease of understanding.
#
# + id="PFNgx_-TEcF4"
data_transform = Compose([ToTensor(), Normalize((0.1307,), (0.3081,))])
train_loader = DataLoader(
MNIST(download=True, root=".", transform=data_transform, train=True), batch_size=128, shuffle=True
)
val_loader = DataLoader(
MNIST(download=True, root=".", transform=data_transform, train=False), batch_size=256, shuffle=False
)
# + [markdown] id="VC9BUtWXEcF6"
# Finally, we will specify the optimizer and the loss function:
# + id="6VkGmtVZEcF7"
optimizer = torch.optim.RMSprop(model.parameters(), lr=0.005)
criterion = nn.CrossEntropyLoss()
# + [markdown] id="cb-ak9gEEcF7"
# And we’re done with setting up the important parts of the project.
# PyTorch-Ignite will handle all other boilerplate code as we will see
# below. Next we have to define a trainer engine by passing our model,
# optimizer and loss function to
# [`create_supervised_trainer`](https://pytorch.org/ignite/generated/ignite.engine.create_supervised_trainer.html),
# and two evaluator engines by passing Ignite’s out-of-the-box
# [metrics](https://pytorch.org/ignite/metrics.html#complete-list-of-metrics)
# and the model to
# [`create_supervised_evaluator`](https://pytorch.org/ignite/generated/ignite.engine.create_supervised_evaluator.html#create-supervised-evaluator). We have defined separate evaluator engines for training and validation because they will serve different functions as we will see later in this tutorial:
# + id="NufcPqJaEcF8"
trainer = create_supervised_trainer(model, optimizer, criterion, device)
val_metrics = {
"accuracy": Accuracy(),
"loss": Loss(criterion)
}
train_evaluator = create_supervised_evaluator(model, metrics=val_metrics, device=device)
val_evaluator = create_supervised_evaluator(model, metrics=val_metrics, device=device)
# + [markdown] id="S7YThetiEcF8"
# The objects `trainer`, `train_evaluator` and `val_evaluator` are all instances of
# [`Engine`](https://pytorch.org/ignite/generated/ignite.engine.engine.Engine.html#ignite.engine.engine.Engine) - the main component of Ignite, which is essentially an abstraction over
# the training or validation loop.
#
# If you need more control over your training and validation loops, you
# can create custom `trainer`, `train_evaluator` and `val_evaluator` objects by wrapping the step
# logic in `Engine` :
#
# ```python
# def train_step(engine, batch):
# model.train()
# optimizer.zero_grad()
# x, y = batch[0].to(device), batch[1].to(device)
# y_pred = model(x)
# loss = criterion(y_pred, y)
# loss.backward()
# optimizer.step()
# return loss.item()
#
# trainer = Engine(train_step)
#
# def validation_step(engine, batch):
# model.eval()
# with torch.no_grad():
# x, y = batch[0].to(device), batch[1].to(device)
# y_pred = model(x)
# return y_pred, y
#
# train_evaluator = Engine(validation_step)
# val_evaluator = Engine(validation_step)
#
# # Attach metrics to the evaluators
# for name, metric in val_metrics.items():
# metric.attach(train_evaluator, name)
#
# for name, metric in val_metrics.items():
# metric.attach(val_evaluator, name)
# ```
# + [markdown] id="Sw90sOK9EcF9"
# We can customize the code further by adding all kinds of event handlers.
# `Engine` allows adding handlers on various events that are triggered
# during the run. When an event is triggered, attached handlers
# (functions) are executed. Thus, for logging purposes we add a function
# to be executed at the end of every `log_interval`-th iteration:
# + id="YGm_-loUEcF9"
# How many batches to wait before logging training status
log_interval = 100
# + id="V3xpFBI6EcF9"
@trainer.on(Events.ITERATION_COMPLETED(every=log_interval))
def log_training_loss(engine):
print(f"Epoch[{engine.state.epoch}], Iter[{engine.state.iteration}] Loss: {engine.state.output:.2f}")
# + [markdown] id="O6uwwXO8EcF-"
# or equivalently without the decorator but attaching the handler function
# to the `trainer` via
# [`add_event_handler`](https://pytorch.org/ignite/generated/ignite.engine.engine.Engine.html#ignite.engine.engine.Engine.add_event_handler)
#
# ``` python
# def log_training_loss(engine):
# print(f"Epoch[{engine.state.epoch}], Iter[{engine.state.iteration}] Loss: {engine.state.output:.2f}")
#
# trainer.add_event_handler(Events.ITERATION_COMPLETED, log_training_loss)
# ```
# + [markdown] id="quQzbAv6EcF-"
# After an epoch ends during training, we can compute the training and
# validation metrics by running `train_evaluator` on `train_loader` and `val_evaluator` on
# `val_loader` respectively. Hence we will attach two additional handlers to `trainer`
# when an epoch completes:
# + id="eCE552PFEcF_"
@trainer.on(Events.EPOCH_COMPLETED)
def log_training_results(trainer):
train_evaluator.run(train_loader)
metrics = train_evaluator.state.metrics
print(f"Training Results - Epoch[{trainer.state.epoch}] Avg accuracy: {metrics['accuracy']:.2f} Avg loss: {metrics['loss']:.2f}")
@trainer.on(Events.EPOCH_COMPLETED)
def log_validation_results(trainer):
val_evaluator.run(val_loader)
metrics = val_evaluator.state.metrics
print(f"Validation Results - Epoch[{trainer.state.epoch}] Avg accuracy: {metrics['accuracy']:.2f} Avg loss: {metrics['loss']:.2f}")
# + [markdown] id="7JRLbhiw903w"
# We can use [`ModelCheckpoint()`](https://pytorch.org/ignite/generated/ignite.handlers.checkpoint.ModelCheckpoint.html#modelcheckpoint) as shown below to save the `n_saved` best models determined by a metric (here accuracy) after each epoch is completed. We attach `model_checkpoint` to `val_evaluator` because we want the two models with the highest accuracies on the validation dataset rather than the training dataset. This is why we defined two separate evaluators (`val_evaluator` and `train_evaluator`) before.
# + id="W6Zd7vKn1LLO"
# Score function to return current value of any metric we defined above in val_metrics
def score_function(engine):
return engine.state.metrics["accuracy"]
# Checkpoint to store n_saved best models wrt score function
model_checkpoint = ModelCheckpoint(
"checkpoint",
n_saved=2,
filename_prefix="best",
score_function=score_function,
score_name="accuracy",
global_step_transform=global_step_from_engine(trainer), # helps fetch the trainer's state
)
# Save the model after every epoch of val_evaluator is completed
val_evaluator.add_event_handler(Events.COMPLETED, model_checkpoint, {"model": model})
# + [markdown] id="KAB0QtrgiTLK"
# We will use [`TensorboardLogger()`](https://pytorch.org/ignite/generated/ignite.contrib.handlers.tensorboard_logger.html#ignite.contrib.handlers.tensorboard_logger.TensorboardLogger) to log trainer's loss, and training and validation metrics separately.
# + id="Rdt6AE6oeh6k"
# Define a Tensorboard logger
tb_logger = TensorboardLogger(log_dir="tb-logger")
# Attach handler to plot trainer's loss every 100 iterations
tb_logger.attach_output_handler(
trainer,
event_name=Events.ITERATION_COMPLETED(every=100),
tag="training",
output_transform=lambda loss: {"batch_loss": loss},
)
# Attach handler for plotting both evaluators' metrics after every epoch completes
for tag, evaluator in [("training", train_evaluator), ("validation", val_evaluator)]:
tb_logger.attach_output_handler(
evaluator,
event_name=Events.EPOCH_COMPLETED,
tag=tag,
metric_names="all",
global_step_transform=global_step_from_engine(trainer),
)
# + [markdown] id="Aq0qwiZrEcF_"
# Finally, we start the engine on the training dataset and run it for 5
# epochs:
# + colab={"base_uri": "https://localhost:8080/"} id="qnmTh4FeEcGA" outputId="f444d98f-8f45-44ea-bd82-9cecb6971bbe"
trainer.run(train_loader, max_epochs=5)
# + id="ZXhL1-vDgBeT"
# Let's close the logger and inspect our results
tb_logger.close()
# %load_ext tensorboard
# %tensorboard --logdir=.
# + colab={"base_uri": "https://localhost:8080/"} id="_xj4NMjdArYh" outputId="3291ca3b-809a-4ed4-d657-0b83eeb45bc5"
# At last we can view our best models
# !ls checkpoints
# + [markdown] id="wJ9k2coEEcGD"
# ## Next Steps
#
# 1. Check out [tutorials](https://pytorch-ignite.ai/tutorials) if you want to continue
# learning more about PyTorch-Ignite.
# 2. Head over to [how-to guides](https://pytorch-ignite.ai/how-to-guides) if you’re looking
# for a specific solution.
# 3. If you want to set-up a PyTorch-Ignite project, visit [Code
# Generator](https://code-generator.pytorch-ignite.ai/) to get a variety of
# easily customizable templates and out-of-the-box features.
# + [markdown] tags=[] id="vya75pqVEcGE"
# ## Complete Code
#
# ``` python
# import torch
# from torch import nn
# from torch.utils.data import DataLoader
# from torchvision.datasets import MNIST
# from torchvision.models import resnet18
# from torchvision.transforms import Compose, Normalize, ToTensor
#
# from ignite.engine import Engine, Events, create_supervised_trainer, create_supervised_evaluator
# from ignite.metrics import Accuracy, Loss
# from ignite.contrib.handlers import TensorboardLogger, global_step_from_engine
#
# device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
#
#
# class Net(nn.Module):
#
# def __init__(self):
# super(Net, self).__init__()
#
# self.model = resnet18(num_classes=10)
#
# self.model.conv1 = self.model.conv1 = nn.Conv2d(
# 1, 64, kernel_size=3, padding=1, bias=False
# )
#
# def forward(self, x):
# return self.model(x)
#
#
# model = Net().to(device)
#
# data_transform = Compose([ToTensor(), Normalize((0.1307,), (0.3081,))])
#
# train_loader = DataLoader(
# MNIST(download=True, root=".", transform=data_transform, train=True), batch_size=128, shuffle=True
# )
#
# val_loader = DataLoader(
# MNIST(download=True, root=".", transform=data_transform, train=False), batch_size=256, shuffle=False
# )
#
# optimizer = torch.optim.RMSprop(model.parameters(), lr=0.005)
# criterion = nn.CrossEntropyLoss()
#
# trainer = create_supervised_trainer(model, optimizer, criterion, device)
#
# val_metrics = {
# "accuracy": Accuracy(),
# "loss": Loss(criterion)
# }
#
# train_evaluator = create_supervised_evaluator(model, metrics=val_metrics, device=device)
# val_evaluator = create_supervised_evaluator(model, metrics=val_metrics, device=device)
#
# log_interval = 100
#
# @trainer.on(Events.ITERATION_COMPLETED(every=log_interval))
# def log_training_loss(engine):
# print(f"Epoch[{engine.state.epoch}], Iter[{engine.state.iteration}] Loss: {engine.state.output:.2f}")
#
# @trainer.on(Events.EPOCH_COMPLETED)
# def log_training_results(trainer):
# train_evaluator.run(train_loader)
# metrics = train_evaluator.state.metrics
# print(f"Training Results - Epoch[{trainer.state.epoch}] Avg accuracy: {metrics['accuracy']:.2f} Avg loss: {metrics['loss']:.2f}")
#
#
# @trainer.on(Events.EPOCH_COMPLETED)
# def log_validation_results(trainer):
# val_evaluator.run(val_loader)
# metrics = val_evaluator.state.metrics
# print(f"Validation Results - Epoch[{trainer.state.epoch}] Avg accuracy: {metrics['accuracy']:.2f} Avg loss: {metrics['loss']:.2f}")
#
#
# def score_function(engine):
# return engine.state.metrics["accuracy"]
#
#
# model_checkpoint = ModelCheckpoint(
# "checkpoint",
# n_saved=2,
# filename_prefix="best",
# score_function=score_function,
# score_name="accuracy",
# global_step_transform=global_step_from_engine(trainer),
# )
#
# val_evaluator.add_event_handler(Events.COMPLETED, model_checkpoint, {"model": model})
#
# tb_logger = TensorboardLogger(log_dir="tb-logger")
#
# tb_logger.attach_output_handler(
# trainer,
# event_name=Events.ITERATION_COMPLETED(every=100),
# tag="training",
# output_transform=lambda loss: {"batch_loss": loss},
# )
#
# for tag, evaluator in [("training", train_evaluator), ("validation", val_evaluator)]:
# tb_logger.attach_output_handler(
# evaluator,
# event_name=Events.EPOCH_COMPLETED,
# tag=tag,
# metric_names="all",
# global_step_transform=global_step_from_engine(trainer),
# )
#
# trainer.run(train_loader, max_epochs=5)
#
# tb_logger.close()
# ```
| tutorials/getting-started.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Implement an Accelerometer
# In this notebook you will define your own `get_derivative_from_data` function and use it to differentiate position data ONCE to get velocity information and then again to get acceleration information.
#
# In part 1 I will demonstrate what this process looks like and then in part 2 you'll implement the function yourself.
# -----
# ## Part 1 - Reminder and Demonstration
# +
# run this cell for required imports
from helpers import process_data
from helpers import get_derivative_from_data as solution_derivative
from matplotlib import pyplot as plt
# +
# load the parallel park data
PARALLEL_PARK_DATA = process_data("parallel_park.pickle")
# get the relevant columns
timestamps = [row[0] for row in PARALLEL_PARK_DATA]
displacements = [row[1] for row in PARALLEL_PARK_DATA]
# calculate first derivative
speeds = solution_derivative(displacements, timestamps)
# plot
plt.title("Position and Velocity vs Time")
plt.xlabel("Time (seconds)")
plt.ylabel("Position (blue) and Speed (orange)")
plt.scatter(timestamps, displacements)
plt.scatter(timestamps[1:], speeds)
plt.show()
# -
# But you just saw that acceleration is the derivative of velocity... which means we can use the same derivative function to calculate acceleration!
# +
# calculate SECOND derivative
accelerations = solution_derivative(speeds, timestamps[1:])
# plot (note the slicing of timestamps from 2 --> end)
plt.scatter(timestamps[2:], accelerations)
plt.show()
# -
# As you can see, this parallel park motion consisted of four segments with different (but constant) acceleration. We can plot all three quantities at once like this:
#
#
plt.title("x(t), v(t), a(t)")
plt.xlabel("Time (seconds)")
plt.ylabel("x (blue), v (orange), a (green)")
plt.scatter(timestamps, displacements)
plt.scatter(timestamps[1:], speeds)
plt.scatter(timestamps[2:], accelerations)
plt.show()
# ----
# ## Part 2 - Implement it yourself!
def get_derivative_from_data(position_data, time_data):
if len(position_data) != len(time_data):
raise(ValueError, "Data sets must have same length")
# 2. Prepare empty list of speeds
speeds = []
# 3. Get first values for position and time
previous_position = position_data[0]
previous_time = time_data[0]
# 4. Begin loop through all data EXCEPT first entry
for i in range(1, len(position_data)):
# 5. get position and time data for this timestamp
position = position_data[i]
time = time_data[i]
# 6. Calculate delta_x and delta_t
delta_x = position - previous_position
delta_t = time - previous_time
# 7. Speed is slope. Calculate it and append to list
speed = delta_x / delta_t
speeds.append(speed)
# 8. Update values for next iteration of the loop.
previous_position = position
previous_time = time
return speeds
# +
# Testing part 1 - visual testing of first derivative
# compare this output to the corresponding graph above.
speeds = get_derivative_from_data(displacements, timestamps)
plt.title("Position and Velocity vs Time")
plt.xlabel("Time (seconds)")
plt.ylabel("Position (blue) and Speed (orange)")
plt.scatter(timestamps, displacements)
plt.scatter(timestamps[1:], speeds)
plt.show()
# +
# Testing part 2 - visual testing of second derivative
# compare this output to the corresponding graph above.
speeds = get_derivative_from_data(displacements, timestamps)
accelerations = get_derivative_from_data(speeds, timestamps[1:])
plt.title("x(t), v(t), a(t)")
plt.xlabel("Time (seconds)")
plt.ylabel("x (blue), v (orange), a (green)")
plt.scatter(timestamps, displacements)
plt.scatter(timestamps[1:], speeds)
plt.scatter(timestamps[2:], accelerations)
plt.show()
| 3_4_Vehicle_Motion/.ipynb_checkpoints/Implement an Accelerometer-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Merge two UTM zones
#
# ```
# #
# # Copyright (c) Sinergise, 2019 -- 2021.
# #
# # This file belongs to subproject "field-delineation" of project NIVA (www.niva4cap.eu).
# # All rights reserved.
# #
# # This source code is licensed under the MIT license found in the LICENSE
# # file in the root directory of this source tree.
# #
# ```
# This notebook implements the methods needed to merge two UTM zones into a single output geopackage.
#
# The procedure outline is:
# * define geometries for two UTM zones and their overlap
# * load the two single-UTM-zone vector predictions
# * split them into parts: non-overlapping (completely within UTM zone) and overlapping
# * merge the overlaps by:
# * transform them to single CRS (WGS84)
# * spatial join of the overlapping geodataframes from the two zones
# * finding geometries that do not overlap (and keeping them)
# * unary_union-ize the polygons that intersect and merge them to the geometries from previous step
# * transform everything to resulting (common) CRS
# * clean up the results (remove geometries with area larger than X * largest GSAA polygon from 2020
# * simplify geometries
# %load_ext autoreload
# %autoreload 2
# +
from pathlib import Path
import geopandas as gpd
import matplotlib.pyplot as plt
from shapely.ops import unary_union
from fd.vectorisation import MergeUTMsConfig, utm_zone_merging
# -
# ### The two UTM zones and their overlap
# +
INPUT_DATA_DIR = Path('../../input-data/')
grid_definition = gpd.read_file(INPUT_DATA_DIR/'cyl-grid-definition.gpkg')
grid_definition.head()
# -
grid_definition['crs'] = grid_definition['name'].apply(lambda name: f'326{name[:2]}')
grid_definition.head()
crs_names = sorted(grid_definition['crs'].unique())
crs_names
utm_geoms = [grid_definition[grid_definition['crs']==crs_name].geometry.unary_union
for crs_name in crs_names]
overlap = utm_geoms[0].intersection(utm_geoms[1]).buffer(-0.0001)
tiled_overlap = grid_definition[grid_definition.intersects(overlap)].unary_union.buffer(-0.0001)
zones = gpd.GeoDataFrame(geometry=[g for g in grid_definition[~grid_definition.intersects(tiled_overlap)].buffer(0.00001).unary_union],
crs=grid_definition.crs)
zones['crs'] = zones.geometry.apply(lambda g: grid_definition[grid_definition.intersects(g)]['crs'].unique()[0])
# +
# a dataframe holding the overlap geometry
# useful because it is much simpler to transform between CRSs
overlap_df = gpd.GeoDataFrame(geometry=[tiled_overlap], crs=grid_definition.crs)
# -
fig, ax = plt.subplots(figsize=(15,10))
zones.plot('crs', ax=ax)
overlap_df.boundary.plot(ax=ax, color='r')
# +
MAX_GSAA_AREA = 19824325 # m2, derived from reference data
merging_config = MergeUTMsConfig(
bucket_name='bucket-name',
aws_access_key_id='',
aws_secret_access_key='',
aws_region='eu-central-1',
time_intervals=['APRIL'],
utms=crs_names,
contours_dir='/home/ubuntu/cyl-contours', # where partial vectors are stored
resulting_crs='epsg:2062', # CRS of resulting geometries
max_area=1.3*MAX_GSAA_AREA, # Specify max area in m2 of parcels to keep
simplify_tolerance=2.5, # This is in teh resulting CRS, careful about unit of measure
n_workers=20
)
# +
import logging
import sys
logging.basicConfig(format='%(asctime)s | %(levelname)s : %(message)s',
level=logging.INFO, stream=sys.stdout)
# -
# Depending on the number of polygons in the overlap area, and on their size, this step can take quite a long time (appr 4 hours per time interval).
utm_zone_merging(merging_config, overlap_df, zones, parallel=False)
| notebooks/12-utm-zone-merging.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Evaluating docker image libraries embedding : library seq2seq
#
# This notebook evaluates the quality of a library embedding by trying to estimate installed native packages from python packages.
#
# It expects the swivel embedding to be accessible in a `./swivel_data` folder, and the result of dockerhub_crawler to be accessible in a `./packages` folder.
#
# The model is a keras LSTM, primarly to test the sequence pipeline, I wanted to use a tranformer network but did not have the time to achieve it.
import pandas as pd
em = pd.read_csv("./swivel_input/row_embedding.tsv", sep="\t", index_col=0, header=None)
em.dropna()
em.head()
from bokeh.io import output_notebook
output_notebook()
# packages_list = !find packages -type f
import json
def get_images_libs():
for path in packages_list:
package_name = path[13:]
with open(path) as json_file:
try:
package = json.load(json_file)
native_libs = [natpack[0] for natpack in package["packages"]["native"]]
python_libs = [natpack[0] for natpack in package["packages"]["python3"]]
node_libs = [natpack[0] for natpack in package["packages"]["node"]]
image = {
"name": package['image'],
"distro": package['distribution'],
"size": package["size"],
"python": python_libs,
"node": node_libs,
"native": native_libs,
}
yield image
except Exception as e:
#print(e)
pass
from tqdm import tqdm
images_libs = [image for image in tqdm(get_images_libs())]
len(images_libs)
# libs = !cat ./swivel_input/row_vocab.txt
def train_test_split(images_libs, ratio=0.2):
test_libs = set(libs[-int(len(libs)*ratio):])
train_images = []
test_images = []
for image in images_libs:
if (set(image['python']) | set(image['node']) | set(image['native']) ) & test_libs:
test_images.append(image)
else:
train_images.append(image)
return train_images, test_images
train, test = train_test_split(images_libs)
print(len(train), len(test))
import numpy as np
def assign_library_vectors(images):
count = 0
for image in tqdm(images):
count += 1
try:
vectors = [em.loc[lib].values for lib in image['python']]
image['python_embedding'] = np.array(vectors)
vectors = [em.loc[lib].values for lib in image['node']]
image['node_embedding'] = np.array(vectors)
vectors = [em.loc[lib].values for lib in image['native']]
image['native_embedding'] = np.array(vectors)
except Exception as e:
pass
assign_library_vectors(train)
assign_library_vectors(test)
test = pd.DataFrame(test)
train = pd.DataFrame(train)
test = test.dropna()
train = train.dropna()
test_save = test
train_save = train
train.head()
train[train.apply(lambda x: len(x['python']) != 0 , axis=1)].info()
#test = test[test.apply(lambda x: len(x['python']) != 0, axis=1)]
#train.head()
print(train.info(), test.info())
# Creating Features and targets
from keras.preprocessing.sequence import pad_sequences
# Getting numpy arrays from pandas
X_train = train['python_embedding'].values
y_train = train['native_embedding'].values
X_test = test['python_embedding'].values
y_test = test['native_embedding'].values
# padding and stacking sequences to obtain one 3D (image, library, lib_embedding_component) array
X_train = np.stack(pad_sequences(X_train, maxlen=100, dtype="object", padding='post', truncating='post', value=np.zeros(300,)))
y_train = np.stack(pad_sequences(y_train, maxlen=100, dtype="object", padding='post', truncating='post', value=np.zeros(300,)))
X_test = np.stack(pad_sequences(X_test, maxlen=100, dtype="object", padding='post', truncating='post', value=np.zeros(300,)))
y_test = np.stack(pad_sequences(y_test, maxlen=100, dtype="object", padding='post', truncating='post', value=np.zeros(300,)))
X = np.append(X_train, X_test, axis=0)
y = np.append(y_train, y_test, axis=0)
print(X.shape, y.shape)
# +
decoder_target_data = np.zeros(
(X.shape[0], X.shape[1], X.shape[2]),
dtype='float32')
for i, (input_libs, target_libs) in enumerate(zip(X, y)):
for t, lib in enumerate(target_libs):
# decoder_target_data is ahead of decoder_input_data by one timestep
if t > 0:
# decoder_target_data will be ahead by one timestep
# and will not include the start character.
decoder_target_data[i, t - 1, :] = lib
decoder_target_data.shape
# +
from keras.models import Model
from keras.layers import Input, Dense, Activation, LSTM
from keras import optimizers
latent_dim = 30
encoder_inputs = Input(shape=(None, 300))
encoder = LSTM(latent_dim, return_state=True)
encoder_outputs, state_h, state_c = encoder(encoder_inputs)
encoder_states = [state_h, state_c]
decoder_inputs = Input(shape=(None, 300))
decoder_lstm = LSTM(latent_dim, return_sequences=True, return_state=True)
decoder_outputs, _, _ = decoder_lstm(decoder_inputs, initial_state=encoder_states)
decoder_dense = Dense(300, activation='sigmoid')
decoder_outputs = decoder_dense(decoder_outputs)
model = Model([encoder_inputs, decoder_inputs], decoder_outputs)
model.compile(optimizer='rmsprop', loss='categorical_crossentropy')
model.summary()
# -
batch_size = 4
epochs = 50
history = model.fit([X, y], decoder_target_data,
batch_size=batch_size,
epochs=epochs,
validation_split=0.2)
# Save model
model.save('s2s.h5')
# +
# Define sampling models
encoder_model = Model(encoder_inputs, encoder_states)
decoder_state_input_h = Input(shape=(latent_dim,))
decoder_state_input_c = Input(shape=(latent_dim,))
decoder_states_inputs = [decoder_state_input_h, decoder_state_input_c]
decoder_outputs, state_h, state_c = decoder_lstm(
decoder_inputs, initial_state=decoder_states_inputs)
decoder_states = [state_h, state_c]
decoder_outputs = decoder_dense(decoder_outputs)
decoder_model = Model(
[decoder_inputs] + decoder_states_inputs,
[decoder_outputs] + decoder_states)
# -
from matplotlib import pyplot as plt
# %matplotlib inline
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model Loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
| DockerHubPackages/research/notebooks/lib_embedding_eval_lib2lib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import pandas as pd
import pytz
import matplotlib
matplotlib.use("Agg")
import matplotlib.pyplot as plt
import pyarrow.parquet as pq
from plotnine import *
import calendar
import numpy as np
import databricks.koalas as ks
import seaborn as sns
from datetime import datetime
# +
data_root = "path to machine metric dataset"
loads = [
('load1', 'node_load1'),
('load5', 'node_load5'),
('load15', 'node_load15'),
('load1ML', 'node_load1')
]
gpu_nodes = {
"r28n1", "r28n2", "r28n3", "r28n4", "r28n5",
"r29n1", "r29n2", "r29n3", "r29n4", "r29n5",
"r30n1", "r30n2", "r30n3", "r30n4", "r30n5", "r30n6", "r30n7",
"r31n1", "r31n2", "r31n3", "r31n4", "r31n5", "r31n6"
"r32n1", "r32n2", "r32n3", "r32n4", "r32n5", "r32n6", "r32n7",
"r33n2", "r33n3", "r33n5", "r33n6",
"r34n1", "r34n2", "r34n3", "r34n4", "r34n5", "r34n6", "r34n7",
"r35n1", "r35n2", "r35n3", "r35n4", "r35n5",
"r36n1", "r36n2", "r36n3", "r36n4", "r36n5",
"r38n1", "r38n2", "r38n3", "r38n4", "r38n5",
}
# -
df = pd.read_parquet(data_root + 'node_load1')
df = df.stack()
df.index.names = ['time', 'node']
df = df.rename("load1").to_frame()
df
# + pycharm={"name": "#%%\n"}
color = ['lightcoral', 'steelblue', 'yellowgreen', 'orchid']
#marker = ['o', '^', 's', '+']
hatch = ['', '/', '\\', '+']
fig, ax = plt.subplots(figsize=(11,5))
index = 0
barWidth = 0.2
offset = [-0.3, -0.1, 0.1, 0.3]
df = None
for load, folder_name in loads:
cach_file = os.path.join("./cache", f"loads_diurnal_hourly_cache_{load}.npy")
# os.remove(cach_file) # Trash the cache?
if not os.path.isfile(cach_file):
df = pd.read_parquet(data_root + folder_name)
if load == "load1ML": # Only keep ML nodes
df = df[set(df.columns).intersection(gpu_nodes)]
# Pivot all columns so that it becomes a multi-index of (time, node).
df = df.stack()
# Set the names of the multi-index
df.index.names = ['time', 'node']
# Change the series name to the load name and then make it a dataframe
df = df.rename(load).to_frame()
# Drop all rows that do not feature at least one value >= 0
df = df.loc[(df >= 0).any(axis=1)]
df.reset_index(inplace=True)
df["dt"] = pd.to_datetime(df['time'], utc=True, unit="s")
# Convert everything into localized Amsterdam time and then drop the timezone info again
# dropping it is required to save the parquet file.
df["dt"] = df["dt"].dt.tz_convert(pytz.timezone('Europe/Amsterdam')).dt.tz_localize(None)
# Get hour of day and day columns to plot
df["hour_of_day"] = df["dt"].dt.hour
yerr_vals = df.groupby("hour_of_day")[load].std()
df = df.groupby("hour_of_day").mean()
x_vals = np.arange(len(df[load])) + offset[index]
y_vals = df[load]
with open(cach_file, 'wb') as cache_file:
np.save(cache_file, x_vals)
np.save(cache_file, y_vals)
np.save(cache_file, yerr_vals)
else:
with open(cach_file, 'rb') as cache_file:
x_vals = np.load(cache_file)
y_vals = np.load(cache_file)
yerr_vals = np.load(cache_file)
negative_direction_values = np.zeros(len(yerr_vals)) # We create a 2d array to make sure matplotlib does not create downwards errorbars
label = load
if label == "load1ML":
label = "load1 ML"
ax.bar(x_vals, y_vals, yerr=[negative_direction_values, yerr_vals], edgecolor='black', color=color[index], hatch=hatch[index], label=label, width=barWidth, capsize=3)
index += 1
# Add the GPU nodes to it
ax.set_xlim(left=-1, right=24)
ax.set_ylim(bottom=0, top=100)
ax.set_xlabel("Hour of Day", fontsize=20)
ax.set_ylabel("Load", fontsize=20)
ax.tick_params(axis='both', which='major', labelsize=18)
ax.tick_params(axis='both', which='minor', labelsize=16)
ax.legend(ncol=len(color), prop={"size": 14}, bbox_to_anchor=(0.5, 1.15), loc=9)
fig.tight_layout()
date_time = datetime.now().strftime("%Y-%m-%d-%H-%M-%S")
fig.savefig(f"loads_diurnal_hourly_{date_time}.pdf")
del fig
del ax
if df is not None: del df
# +
color = ['lightcoral', 'steelblue', 'yellowgreen']
#marker = ['o', '^', 's']
hatch = ['', '/', '\\']
fig, ax = plt.subplots(figsize=(11,5))
index = 0
offset = [-0.3, 0, 0.3]
df = None
for load, folder_name in loads:
cach_file = os.path.join("./cache", f"loads_daily_cache_{load}.npy")
if not os.path.isfile(cach_file):
df = pd.read_parquet(data_root + folder_name)
# Pivot all columns so that it becomes a multi-index of (time, node).
df = df.stack()
# Set the names of the multi-index
df.index.names = ['time', 'node']
# Change the series name to the load name and then make it a dataframe
df = df.rename(load).to_frame()
# Drop all rows that do not feature at least one value >= 0
df = df[(df >= 0).any(axis=1)]
df.reset_index(inplace=True)
df["dt"] = pd.to_datetime(df['time'], utc=True, unit="s")
# Convert everything into localized Amsterdam time and then drop the timezone info again
# dropping it is required to save the parquet file.
df["dt"] = df["dt"].dt.tz_convert(pytz.timezone('Europe/Amsterdam')).dt.tz_localize(None)
# Get hour of day and day columns to plot
# df["hour_of_day"] = df["dt"].dt.hour
df["day"] = df["dt"].apply(lambda x : x.weekday())
yerr_vals = df.groupby("day")[load].std()
df = df.groupby("day").mean()
x_vals = np.arange(len(df[load])) + offset[index]
y_vals = df[load]
with open(cach_file, 'wb') as cache_file:
np.save(cache_file, x_vals)
np.save(cache_file, y_vals)
np.save(cache_file, yerr_vals)
else:
with open(cach_file, 'rb') as cache_file:
x_vals = np.load(cache_file)
y_vals = np.load(cache_file)
yerr_vals = np.load(cache_file)
negative_direction_values = np.zeros(len(yerr_vals)) # We create a 2d array to make sure matplotlib does not create downwards errorbars
ax.bar(x_vals, y_vals, yerr=[negative_direction_values, yerr_vals], edgecolor='black', color=color[index], hatch=hatch[index], label=load, width=barWidth, capsize=3)
index += 1
ax.set_xlim(left=-1)
ax.set_ylim(bottom=0, top=100)
ax.set_xticks(list(np.arange(7)))
ax.set_xticklabels(['Mon', 'Tue', 'Wed', 'Thu', 'Fri', 'Sat', 'Sun'])
ax.set_xlabel("Day of Week", fontsize=20)
ax.set_ylabel("Load", fontsize=20)
ax.tick_params(axis='both', which='major', labelsize=18)
ax.tick_params(axis='both', which='minor', labelsize=16)
ax.legend(ncol=len(color), prop={"size": 14}, bbox_to_anchor=(0.5, 1.15), loc=9)
fig.tight_layout()
date_time = datetime.now().strftime("%Y-%m-%d-%H-%M-%S")
fig.savefig(f"loads_diurnal_daily_{date_time}.pdf")
del fig
del ax
if df: del df
# + pycharm={"name": "#%%\n"}
print(df["dt"].min(), df["dt"].max())
print(len(df), len(df.columns))
# -
with open('./cache/loads_diurnal_hourly_cache_load1ML.npy', 'rb') as cache_file:
x_vals = np.load(cache_file)
y_vals = np.load(cache_file)
yerr_vals = np.load(cache_file)
print(y_vals.min(), y_vals.max())
# + pycharm={"name": "#%%\n"}
# First, compute the average for the same hours per node, then the mean across all nodes
load1_per_node_per_hour = df.groupby("hour_of_day").mean()
load1_per_hour = load1_per_node_per_hour.mean(axis=1).reset_index()
load1_per_hour.columns = ["hour_of_day", "load"]
print(load1_per_hour.head())
# + pycharm={"name": "#%%\n"}
# This cell plots the average across all nodes.
plt = ggplot(load1_per_hour) +\
theme_light(base_size=16) +\
theme(legend_title=element_text(size=0, alpha=0),
legend_box_spacing=0.1,
legend_box_margin=0,
legend_margin=0,
legend_position=(0.51, 0.7),
legend_direction="horizontal",
legend_key=element_blank(),
legend_background=element_rect(fill=(0,0,0,0))) +\
guides(color=guide_legend(ncol=3)) +\
geom_line(aes(x="hour_of_day", y="load")) +\
geom_point(aes(x="hour_of_day", y="load"), size=3) +\
ylim(0,None) +\
xlab("Hour of day") +\
ylab("Avg. load1 across all nodes")
plt.save("load1_per_hour.pdf")
plt
# + pycharm={"name": "#%%\n"}
# First, compute the average for the same hours per node, then the mean across all nodes
load1_per_node_per_day = df.groupby("day").mean()
load1_per_day = load1_per_node_per_day.mean(axis=1).reset_index()
load1_per_day.columns = ["day", "load"]
# + pycharm={"name": "#%%\n"}
plt = ggplot(load1_per_day) +\
theme_light(base_size=16) +\
theme(legend_title=element_text(size=0, alpha=0),
legend_box_spacing=0.1,
legend_box_margin=0,
legend_margin=0,
legend_position=(0.51, 0.7),
legend_direction="horizontal",
legend_key=element_blank(),
legend_background=element_rect(fill=(0,0,0,0))) +\
guides(color=guide_legend(ncol=3)) +\
geom_line(aes(x="day", y="load")) +\
geom_point(aes(x="day", y="load"), size=3) +\
ylim(0,None) +\
xlab("Day in Week (0=Monday, 6=Sunday)") +\
ylab("Avg. load1 across all nodes")
plt.save("load1_per_day.pdf")
plt
# +
# # Make bins of 15 minutes using resample and then create a sliding window so that for every 15 minutes we get the mean load.
# bin_df = df.copy()
# bin_df.index = pd.to_datetime(bin_df.index, unit="s")
# # Bin per 15 minute and create a sliding window of 1 hour.
# # We take the right timestamp of the bin as this is the current time when measuring the mean.
# bin_df = bin_df.resample("15min", label='right').mean().rolling('1h').mean()
# bin_df = bin_df.dropna(how="all") # Remove all rows with only NaN values
# # IMPORTANT: as we took all right labels of each bin, the hour_of_day and day themselves are now incorrect,
# # as all timestamps effectively shifted by 15 minutens. We need to recompute them.
# bin_df["dt"] = pd.to_datetime(bin_df.index, unit="s") # No need to convert time timezones again, this was already done!
# bin_df["hour_of_day"] = bin_df["dt"].dt.hour
# bin_df["day"] = bin_df["dt"].apply(lambda x : x.weekday())
# bin_df
# +
# load1_per_node_per_hour = bin_df.groupby("hour_of_day").mean()
# load1_per_hour = load1_per_node_per_hour.mean(axis=1).reset_index()
# load1_per_hour.columns = ["hour_of_day", "load"]
# +
# This cell plots the average across all nodes per 15m using a rollowing window of 1 hour
# plt = ggplot(load1_per_hour) +\
# theme_light(base_size=16) +\
# theme(legend_title=element_text(size=0, alpha=0),
# legend_box_spacing=0.1,
# legend_box_margin=0,
# legend_margin=0,
# legend_position=(0.51, 0.7),
# legend_direction="horizontal",
# legend_key=element_blank(),
# legend_background=element_rect(fill=(0,0,0,0))) +\
# guides(color=guide_legend(ncol=3)) +\
# geom_line(aes(x="hour_of_day", y="load")) +\
# geom_point(aes(x="hour_of_day", y="load"), size=3) +\
# ylim(0,None) +\
# xlab("Hour of day") +\
# ylab("Avg. load1 across all nodes")
plt = ggplot(load1_per_hour, aes(x="hour_of_day", y="load")) +\
theme_light(base_size=16) +\
theme(axis_text_x = element_text(angle = 45)) +\
geom_bar(stat = "identity") +\
ylim(0,None) +\
xlab("Hour of day") +\
ylab("Avg. load1 across all nodes")
plt.save("load1_per_hour_of_day_diurnal_15min_bin_1h_window.pdf")
plt
# -
# First, compute the average for the same hours per node, then the mean across all nodes
load1_per_node_per_day = bin_df.groupby("day").mean()
load1_per_day = load1_per_node_per_day.mean(axis=1).reset_index()
load1_per_day.columns = ["day", "load"]
# +
plt = ggplot(load1_per_day, aes(x="day", y="load")) +\
theme_light(base_size=16) +\
theme(axis_text_x = element_text(angle = 45)) +\
geom_bar(stat = "identity") +\
ylim(0,None) +\
xlab("Day in Week") +\
ylab("Avg. load1 across all nodes") +\
scale_x_continuous(breaks=list(range(0,7)), labels=list(calendar.day_name))
plt.save("load1_per_day_of_week_diurnal_15min_bin_1h_window.pdf")
plt
# +
# Create a series of hour of day -> all values to plot in a violin/boxplot.
def get_values(rows):
print(rows.columns)
hour = rows['hour_of_day'].iloc[0]
rows.drop('hour_of_day', axis=1, inplace=True)
arr = rows.to_numpy()
return arr[arr >= 0].ravel()
ndf = df.groupby('hour_of_day').apply(get_values)
ndf
# +
# Create per hour a violin boxplot plot
plt = ggplot(hour_of_day_df, aes(x="hour_of_day", y="values")) +\
geom_violin(width=0.2) +\
geom_boxplot(width=0.1, color="grey", alpha=0.2) +\
scale_fill_cmap(discrete = True) +\
theme_light() +\
theme(
legend_position="none",
plot_title = element_text(size=11)
) +\
ggtitle("A Violin wrapping a boxplot") +\
xlab("")
plt
| daily_weekly_trend_load.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Random Values in Tensors
import tensorflow as tf
# #### a) Normal distribution
# +
# random_normal method signature
# tf. random.normal(shape, mean = 0, stddev =2, dtype=tf.float32, seed=None, name=None)
# -
tf.random.normal(shape=(6, 5))
tf.random.normal(shape=(2, 4), mean=8, stddev=3.2)
# ### b) Uniform Distribution
# +
# random_uniform Method signature
# tf.random.uniform(shape, minval = 0, maxval= None, dtype=tf.float32, seed=None, name=None)
# -
tf.random.uniform(shape=(4, 2))
tf.random.uniform(shape=(3, 3), minval = 0, maxval= 10, seed=30)
#
# #### c) Seeding
#
# + active=""
#
# Seeding allows us to get the same values from random operations
# -
tf.random.set_seed(2)
a = tf.random.uniform(shape=(3, 3), minval = 0, maxval= 10,dtype=tf.int32)
b = tf.random.uniform(shape=(3, 3), minval = 0, maxval= 10,dtype=tf.int32)
print(f"a: {a}\n")
print(f'b: {b}')
# +
# Seed with same value (2)
tf.random.set_seed(2)
# -
a1 = tf.random.uniform(shape=(3, 3), minval = 0, maxval= 10,dtype=tf.int32)
b1 = tf.random.uniform(shape=(3, 3), minval = 0, maxval= 10,dtype=tf.int32)
print(f"a1: {a1}\n")
print(f'b1: {b1}')
| notebooks/1 intro_to_tensorflow/1.7 random.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # 1D harmonic oscillator physics-informed neural network (PINN)
#
# This notebook is based on <NAME>'s blog post ["So, what is a physics-informed neural network?"](https://benmoseley.blog/my-research/so-what-is-a-physics-informed-neural-network/) and his original code which can be found [here](https://github.com/benmoseley/harmonic-oscillator-pinn). Please refer to Ben's blog post and code if you're interested.
#
# This notebook differs from the original in that it
# - is based on **tensorflow 2** (instead of PyTorch) for neural network implementation
# - adds the option to **learn model parameters** while training the neural network
#
# We will use a simple physical example problem to explain how PINNs work, how to implement them, and will demonstrate their benefit compared to regular neural nets as function approximators.
#
# The code is partially based on other PINN implementations which can be found at [Github by pierremtb](https://github.com/pierremtb/PINNs-TF2.0), [Github by maziarraissi](https://github.com/maziarraissi/PINNs), and [Github by janblechschmidt](https://github.com/janblechschmidt/PDEsByNNs).
# ## Problem description
#
# The example problem we solve here is the 1D damped harmonic oscillator:
# $$
# m \dfrac{d^2 x}{d t^2} + \mu \dfrac{d x}{d t} + kx = 0~,
# $$
# with initial conditions
# $$
# x(0) = 1~~,~~\dfrac{d x}{d t} = 0~.
# $$
# For the under-damped state, i.e. when
# $$
# \delta < \omega_0~,~~~~~\mathrm{with}~~\delta = \dfrac{\mu}{2m}~,~\omega_0 = \sqrt{\dfrac{k}{m}}~.
# $$
# we can find the following exact solution:
# $$
# x(t) = e^{-\delta t}(2 A \cos(\phi + \omega t))~,~~~~~\mathrm{with}~~\omega=\sqrt{\omega_0^2 - \delta^2}~.
# $$
#
# Please refer to the following blog post for more information: https://beltoforion.de/en/harmonic_oscillator/.
# ## Workflow
#
# - First, we will generate some **training data** along part of the exact solution
# - then, we will implement a **regular neural network** and train it to approximate the function between input $t$ and output $x(t)$; we will examine the performance of the model prediction for a set of unseen data points $t$, around the training data points and in areas without training data
# - lastly, we will implement a **physics-informed neural network**, enforcing the ordinary differential equation at a number of collocation points over the time interval of interest; we will compare the predictive performance of the PINN to that of the regular NN.
# +
# import necessary python packages
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
from time import time
import tensorflow.keras as keras
from tensorflow.keras.layers import Input
from tensorflow.keras.layers import Dense
from tensorflow.keras.models import Model
from tensorflow.keras.optimizers import Adam
# Set data type
DTYPE='float32'
tf.keras.backend.set_floatx(DTYPE)
# -
# ### 1. Generate training data
#
# We define a function representing the analytical solution to the underdamped harmonic oscillator problem given above.
def oscillator(d, w0, t):
"""Defines the analytical solution to the 1D underdamped harmonic oscillator problem.
Equations taken from: https://beltoforion.de/en/harmonic_oscillator/"""
assert d < w0
w = np.sqrt(w0**2-d**2)
phi = np.arctan(-d/w)
A = 1/(2*np.cos(phi))
cosine = np.cos(phi+w*t)
sine = np.sin(phi+w*t)
exp = np.exp(-d*t)
y = exp*2*A*cosine
return y
# We pick an observation period of $T=1$ and compute the analytical solution $x(t)$ in this timeframe $t\in [0,1]$ for fixed model parameters $\delta = 2.0$ and $\omega_0 = 20.0$.
# We select a total of 10 data points $(t,x(t))$ in the first half of the observation period as our training data for the neural network.
# +
d, w0 = 2.0, 20.0
# get the analytical solution over the full domain
t = np.linspace(0,1,500)
x = oscillator(d, w0, t)
# slice out a small number of points from the left half of the domain: training data
t_data = t[0:200:20]
x_data = x[0:200:20]
# plot analytical solution and training data
plt.figure()
plt.plot(t, x, label="Exact solution")
plt.scatter(t_data, x_data, color="tab:orange", label="Training data")
plt.xlabel('t')
plt.ylabel('x(t)')
plt.legend()
plt.show()
# -
# ### 2. Regular neural network
#
# First, we convert the training data arrays into the correct format and shape for working with tensorflow
# +
# convert arrays to tf tensors
t_data_tf = tf.convert_to_tensor(t_data, dtype=DTYPE)
x_data_tf = tf.convert_to_tensor(x_data, dtype=DTYPE)
# reshape to correct format
T_data = tf.reshape(t_data_tf[:], shape=(t_data.shape[0],1))
X_data = tf.reshape(x_data_tf[:], shape=(x_data.shape[0],1))
# -
# Now we define a standard neural network architecture, using `T_data` as input and train the neural net to predict `X_data`, by minimizing the MSE between `X_model` and `X_data` using gradient descent. Note that input and output of the network are both one-dimensional. We input one time value and receive one x-value as output.
# +
## NN architecture
inputs = Input(shape=(1,))
hidden = Dense(32, activation='tanh')(inputs)
hidden = Dense(32, activation='tanh')(hidden)
out = Dense(1, activation='linear')(hidden)
model_NN = Model(inputs=inputs, outputs=out)
model_NN.compile(Adam(learning_rate=1e-3), loss=tf.keras.losses.MeanSquaredError())
# Describe model
model_NN.summary()
# +
# train regular NN
from time import time
start = time()
history = model_NN.fit(T_data, X_data, epochs=1500, verbose=0,batch_size=10)
print('time taken : ',np.round(time() - start,3))
# +
# plot loss over epochs
plt.plot(history.history['loss'])
plt.legend(['MSE loss'])
plt.xlabel('$n_{epoch}$',fontsize=18)
plt.ylabel('$\\phi^{n_{epoch}}$',fontsize=18)
plt.title('loss for regular NN')
plt.show()
# -
# ### Result: regular NN
#
# We use the model to predict $x(t)$ over the whole observation domain $t\in[0,1]$.
# The trained neural network does well at approximating the analytical solution in the vicinity of the training data points in $t\in[0,0.4]$. All data points and the curvature of the trajectory are well matched.
#
# However, the neural network does not capture the analytical trajectory in the time regimen where we don't have training data points $t\in[0.4,1.0]$. There are no constraints on the solution here that the neural network could learn from.
# +
# examine posterior prediction
# create test-values over whole domain t=0...1
t_pred = np.linspace(0,1,500)
# evaluate model prediction for t_pred
x_pred = model_NN.predict(t_pred)
# plot prediction
fig, ax2 = plt.subplots(figsize=(700/72,500/72))
ax2.set_title('Regular NN')
ax2.scatter(t_data,x_data,s=300,color="tab:orange", alpha=1.0,marker='.') #training data points
ax2.plot(t,x,color="steelblue",linewidth=3.0)
ax2.plot(t_pred,x_pred,color="black",linewidth=3.0,linestyle="--")
ax2.set_ylabel('x(t)',fontsize=14)
ax2.set_xlabel('t',fontsize=14)
ax2.legend(('training data','exact solution','model prediction'), loc='center right',fontsize=14)
# -
# ### 3. Physics-informed neural network
#
# In order to improve the network's ability to approximate the solution, we will ensure that the solution is consistent with the undelying differential equation defining the oscillator problem
#
# $$
# m \dfrac{d^2 x}{d t^2} + \mu \dfrac{d x}{d t} + kx = 0~,
# $$
#
# In addition to minimizing the difference between `X_data` and `X_model` at the training data points, we now also want to minimize the residual of the ODE (assuming $m=1$)
#
# $$
# r = \dfrac{d^2 x}{d t^2} + \mu \dfrac{d x}{d t} + kx,
# $$
#
# at a number of collocation points along the whole observation period.
# We used $\delta = 2.0$ and $\omega_0 = 20.0$ to generate our analytical solution, so the corresponding parameters for the ODE are (assuming $m=1$)
#
# $$
# \mu = 2 \delta = 4.0~,~k = \omega_0^2~=~400.0~.
# $$
#
# Let's first define the collocation points at which we want to enforce the ODE and convert them into tensorflow format:
# +
t_physics = np.linspace(0,1,30)# sample locations over the problem domain: collocation points
no_physics_points = t_physics.shape[0]
# convert to tensor and correct shape
t_physics_tf = tf.convert_to_tensor(t_physics, dtype=DTYPE)
T_r = tf.reshape(t_physics_tf[:], shape=(t_physics.shape[0],1))
# -
# ### PINN architecture
#
# Next, we will define a class "PINNIdentificationNet" which we can call later to create an instance of a fully connected neural network architecture of desired complexity.
#
# Often, we know the physical laws behind a problem but we don't know the exact model parameters and want to learn them from the data.
#
# To this end, tensorflow allows to define trainable "variables" ([tf.Variable](https://www.tensorflow.org/api_docs/python/tf/Variable)) which will be trained together with the weights and biases of the neural net, but are not connected to the model input. In fact, we don't want them to be connected to the model input, because that would imply our model parameters are functions of time.
#
# We define one trainable variable for each model parameter $\mu$ and $k$, set an initial guess and set the argument `trainable=True`. If you don't want to learn the model parameters from the data, set `trainable=False` and the variable will be fixed to the set initial value.
# Define model architecture
class PINNIdentificationNet(tf.keras.Model):
""" Set basic architecture of the PINN model."""
def __init__(self,
output_dim=1,
num_hidden_layers=2,
num_neurons_per_layer=32,
activation='tanh',
kernel_initializer='glorot_normal',
**kwargs):
super().__init__(**kwargs)
self.num_hidden_layers = num_hidden_layers
self.output_dim = output_dim
# Define NN architecture
self.hidden = [tf.keras.layers.Dense(num_neurons_per_layer,
activation=tf.keras.activations.get(activation),
kernel_initializer=kernel_initializer)
for _ in range(self.num_hidden_layers)]
self.out = tf.keras.layers.Dense(output_dim)
# Initialize variable for mu, k: if trainable=True, will be trained along NN
self.mu = tf.Variable(1.0, trainable=True, dtype=DTYPE)
self.mu_list = []
self.k = tf.Variable(300.0, trainable=True, dtype=DTYPE)
self.k_list = []
def call(self, T):
"""Forward-pass through neural network."""
Z = self.hidden[0](T)
for i in range(1,self.num_hidden_layers):
Z = self.hidden[i](Z)
return self.out(Z)
# ### PINN solver
#
# We define a custom solver that allows us to compute the ODE residual associated loss over all collocation points at each training step and add it to the loss associated with the difference between model prediction and training data.
#
# The function `get_r` computes the first and second derivatives of the model output with respect to the model inputs at the collocation points. The automatic differentiation features of tensorflow ([tf.GradientTape](https://www.tensorflow.org/api_docs/python/tf/GradientTape)) make this very easy.
#
# We then feed these derivatives to the function `fun_r` which returns the ODE residual at the collocation points based on the derivatives and the trainable model parameters `self.model.mu` and `self.model.k`.
#
# The function `loss_fn` sums up the computed residual loss at the collocation points and the mean squared error between model prediction and data at the training data points.
#
# `get_grad` and `solve_with_TFoptimizer` take care of the gradient descent method for updating the weights, biases, and trainable variables of the network.
#
# The remaining functions are helper functions for tracking loss history and creating results plots.
class PINNSolver_ID():
def __init__(self, model, T_r):
self.model = model
# Store collocation points
self.t = T_r[:,0:1]
# Initialize history of losses and global iteration counter
self.hist = []
self.iter = 0
def get_r(self):
with tf.GradientTape(persistent=True) as tape:
# Watch variables representing t during this GradientTape
tape.watch(self.t)
# Compute current values u(t)
u = self.model(self.t[:,0:1])
u_t = tape.gradient(u, self.t)
u_tt = tape.gradient(u_t, self.t)
del tape
return self.fun_r(u, u_t, u_tt)
def fun_r(self, u, u_t, u_tt):
return u_tt + self.model.mu*u_t + self.model.k*u
def loss_fn(self, T, u):
# Compute phi_r: loss coming from ODE residual
r = self.get_r()
phi_r = (1e-4)*tf.reduce_mean(tf.square(r))
# Initialize loss
loss = phi_r
# Add loss coming from difference between model prediction and training data
for i in range(len(T)):
u_pred = self.model(T[i:i+1,0:1])
loss += tf.reduce_mean(tf.square(u[i] - u_pred))
return loss
def get_grad(self, T, u):
with tf.GradientTape(persistent=True) as tape:
# This tape is for derivatives with
# respect to trainable variables
tape.watch(self.model.trainable_variables)
loss = self.loss_fn(T, u)
g = tape.gradient(loss, self.model.trainable_variables)
del tape
return loss, g
def solve_with_TFoptimizer(self, optimizer, T, u, N=10000):
"""This method performs a gradient descent type optimization."""
@tf.function
def train_step():
loss, grad_theta = self.get_grad(T, u)
# Perform gradient descent step
optimizer.apply_gradients(zip(grad_theta, self.model.trainable_variables))
return loss
for i in range(N):
loss = train_step()
self.current_loss = loss.numpy()
self.callback()
def callback(self, tr=None):
mu = self.model.mu.numpy()
self.model.mu_list.append(mu)
k = self.model.k.numpy()
self.model.k_list.append(k)
if self.iter % 100 == 0:
print('It {:05d}: loss = {:10.8e} mu = {:10.8e} k = {:10.8e}'.format(self.iter, self.current_loss, mu, k))
self.hist.append(self.current_loss)
self.iter+=1
def plot_solution(self, **kwargs):
n = 500
t_pred = np.reshape(np.linspace(0,1,n),(n,1))
x_pred = self.model(t_pred)
# plot prediction
fig, ax2 = plt.subplots(figsize=(700/72,500/72))
ax2.set_title('PINN')
ax2.scatter(t_data,x_data,s=300,color="tab:orange", alpha=1.0,marker='.') #observed data points
ax2.plot(t,x,color="steelblue",linewidth=3.0)
ax2.plot(t_pred,x_pred,color="black",linewidth=3.0,linestyle="--")
ax2.legend(('training data','exact solution','model prediction'), loc='upper right',fontsize=14)
ax2.set_xlabel('t',fontsize=14)
ax2.set_ylabel('x(t)',fontsize=14)
return ax2
def plot_loss_history(self, ax=None):
if not ax:
fig = plt.figure(figsize=(700/72,500/72))
ax = fig.add_subplot(111)
ax.semilogy(range(len(self.hist)), self.hist,'k-')
ax.set_xlabel('$n_{epoch}$',fontsize=18)
ax.set_ylabel('$\\phi^{n_{epoch}}$',fontsize=18)
return ax
def plot_loss_and_param(self, axs=None):
color_mu = 'tab:blue'
color_k = 'tab:red'
fig = plt.figure(figsize=(1200/72,800/72))
gs = fig.add_gridspec(2, 2)
ax1 = plt.subplot(gs[0, 0:2])
ax1 = self.plot_loss_history(ax1)
ax2 = plt.subplot(gs[1, 0])
ax2.plot(range(len(self.hist)), self.model.mu_list,'-',color=color_mu)
ax2.set_ylabel('$\\mu^{n_{epoch}}$', color=color_mu, fontsize=18)
ax2.set_xlabel('$n_{epoch}$',fontsize=18)
ax3 = plt.subplot(gs[1, 1])
ax3.plot(range(len(self.hist)), self.model.k_list,'-',color=color_k)
ax3.set_ylabel('$k^{n_{epoch}}$', color=color_k, fontsize=18)
ax3.set_xlabel('$n_{epoch}$',fontsize=18)
return (ax1,ax2,ax3)
# ### Run PINN
#
# Now, we just need to instantiate the model and solver and call the `solve_with_TFoptimizer` method to train the network. We decided to use a basic Adam optimizer with a fixed learing rate here.
# +
# Initialize model
model = PINNIdentificationNet()
model.build(input_shape=(None,1))
# Initialize PINN solver
solver = PINNSolver_ID(model, T_r)
# Start timer
t0 = time()
lr = 1e-2
optim = tf.keras.optimizers.Adam(learning_rate=lr)
solver.solve_with_TFoptimizer(optim, T_data, X_data, N=20000)
# Print computation time
print('\nComputation time: {} seconds'.format(time()-t0))
# -
# ### Results PINN
#
# Now let's take a look at how the PINN approximates the function. This time, the solution is well matched even in the time interval where we don't have any training data.
#
# We also observe that our guesses for $\mu$ and $k$ slowly approach the correct values of $\mu = 4.0$ and $k = 400.0$ over the course of the training epochs. Yay!
# plot loss history and model prediction
ax = solver.plot_solution();
axs = solver.plot_loss_and_param();
| oscillator_PINN_tf2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ### Dependencies
from lab5_solution import Lab5Solution
# ### Task Solutions
# +
#Initialization
#Format:
#city city_ascii lat lng country iso2 iso3 admin_name capital population
file = "worldcities-free-100.csv"
solution = Lab5Solution(file)
#Solution task 1
task = "task1"
#Solution task 1
task = "task2"
#Instead of having several small mappings the transformation is done using a single function or mapping
#task = "Simple_Mapping"
#Create RDF triples
if task == "task1":
solution.Task1() #Fresh entity URIs
elif task == "task2":
solution.Task2() #Reusing URIs from DBPedia
else:
solution.SimpleUniqueMapping() #Simple and unique mapping/transformation
#Graph with only data
solution.saveGraph(file.replace(".csv", "-"+task)+".ttl")
#OWL 2 RL reasoning
#We will see reasoning next week. Not strictly necessary for this
solution.performReasoning("../ontology_lab5.ttl") ##ttl format
#Graph with ontology triples and entailed triples
solution.saveGraph(file.replace(".csv", "-"+task)+"-reasoning.ttl")
#SPARQL results into CSV
solution.performSPARQLQuery(file.replace(".csv", "-"+task)+"-query-results.csv")
# -
| lab5/solution/lab5_solution_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Goal:
#
# * Modeling a theoretical diffusive boundary layer (DBL).
# * A DBL may be contributing to 'smearing' observed in 16S rRNA MiSeq data from real experiments.
# # Init
# +
import os
import numpy as np
from scipy.integrate import quad
# %load_ext rpy2.ipython
workDir = '/home/nick/notebook/SIPSim/dev/theory/'
# + language="R"
# library(readxl)
# library(dplyr)
# library(tidyr)
# library(ggplot2)
# library(rootSolve)
# -
if not os.path.isdir(workDir):
os.makedirs(workDir)
# %cd $workDir
# # Setting parameters
# + language="R"
# # tube characteristics (cm)
# tube_diam = 1.3
# tube_height = 4.8
# tube_round_bottom_height = 0.65
# tube_capacity__ml = 4.7
# tube_composition = 'polypropylene'
#
# # rotor (cm)
# rotor_id = 'TLA-110'
# r_min = 2.6
# r_ave = 3.72
# r_max = 4.85
# frac_tube_angle = 90
#
# # cfg run
# ## rpm of run
# rpm = 55000
# ## angular velocity (w^2)
# angular_velocity = 17545933.74
# ## average particle density
# ave_gradient_density = 1.70
# ## beta^o
# BetaO = 1.14e9 # CsCl at density of 1.70
# ## position of particle at equilibrium
# particle_at_eq = 3.78
# ## max 13C shift
# max_13C_shift_in_BD = 0.036
# ## min BD (that we care about)
# min_GC = 13.5
# min_BD = min_GC/100.0 * 0.098 + 1.66
# ## max BD (that we care about)
# max_GC = 80
# max_BD = max_GC / 100.0 * 0.098 + 1.66 # 80.0% G+C
# max_BD = max_BD + max_13C_shift_in_BD
#
# # diffusive boundary layer (DBL)
# DBL_size_range__micron = c(10,100)
#
#
# # misc
# fraction_vol__cm3 = 0.1
# + language="R"
# # rotor angle
# ## sin(x) = opp / hypo
# ## x = sin**-1(opp/hypo)
#
# rad2deg = function(rad) {
# return((180 * rad) / pi)
# }
# deg2rad = function(deg){
# return(deg * pi / 180)
# }
#
#
# x = r_max - r_min
# hyp = tube_height
# rotor_tube_angle = rad2deg(asin(x / hyp))
# cat("Tube angle from axis of rotation:", rotor_tube_angle, "\n")
# + language="R"
# rad2deg = function(rad) {
# return((180 * rad) / pi)
# }
# deg2rad = function(deg){
# return(deg * pi / 180)
# }
#
#
# x = r_max - r_min
# hyp = tube_height
# rotor_tube_angle = rad2deg(asin(x / hyp))
# cat("Tube angle from axis of rotation:", rotor_tube_angle, "\n")
# + language="R"
# # calc tube angle from tube params
# calc_tube_angle = function(r_min, r_max, tube_height){
# x = r_max - r_min
# hyp = tube_height
# rotor_angle = rad2deg(asin(x / hyp))
# return(rotor_angle)
# }
#
# # test
# ## angled tube
# ret = calc_tube_angle(r_min, r_max, tube_height)
# print(ret)
# ## vertical tube
# r_min_v = 7.47
# r_max_v = 8.79
# ret = calc_tube_angle(r_min_v, r_max_v, tube_height)
# print(ret)
# + language="R"
# # isoconcentration point
# ## Formula 6.7 in Birnine and Rickwood 1978
# I = sqrt((r_min**2 + r_min * r_max + r_max**2)/3)
#
# cat('Isoconcentration point:', I, '(cm)\n')
# -
# # ratio of DBL size : fraction size as a function of DBL size
# ## Rough approximation
# + language="R"
#
# DBL_rel_size = function(DBL_size, tube_diam, frac_size){
# # sizes in cm
# tube_radius = tube_diam / 2
# frac_vol = pi * tube_radius**2 * frac_size
# nonDBL_vol = pi * (tube_radius - DBL_size)**2 * frac_size
# DBL_vol = frac_vol - nonDBL_vol
# DBL_to_frac = DBL_vol / frac_vol * 100
# return(DBL_to_frac)
# }
#
# # in cm
# frac_size = 0.01
# tube_diam = 1.3
# #DBL_size = 0.01
# DBL_sizes = seq(0, 0.07, 0.005)
#
# DBL_perc = sapply(DBL_sizes, DBL_rel_size, tube_diam=tube_diam, frac_size=frac_size)
#
# df = data.frame('DBL_size' = DBL_sizes, 'DBL_perc' = DBL_perc)
#
# ggplot(df, aes(DBL_size, DBL_perc)) +
# geom_point() +
# geom_line() +
# labs(x='DBL size (cm)', y='% tube volume that is DBL') +
# theme_bw() +
# theme(
# text = element_text(size=16)
# )
# -
# #### Notes
#
# * Assuming cfg tube is just a cylinder
# # Determining DBL from fragment G+C content
#
# * fragment GC -->
# * BD (diffusive boundary layer) -->
# * angled tube position of DBL -->
# * vertical tube position range of DBL (min, mid, max)
# # Functions for calculating DBL
# ## GC to BD
# + language="R"
# GC2BD = function(GC){
# # GC = percent G+C
# GC / 100.0 * 0.098 + 1.66
# }
#
# # test
# GC = seq(0, 100, 10)
# sapply(GC, GC2BD)
# -
# ## BD to distance from the axis of rotation
# \begin{align}
# x = \sqrt{( ({\rho}-p_m) \frac{2B^{\circ}}{w^2}) + r_c^2}
# \end{align}
# + language="R"
# BD2distFromAxis = function(BD, D, BetaO, w2, I){
# # converting BD to distance from axis of rotation
# # BD = density at a given radius
# # w^2 = angular velocity
# # \beta^o = beta coef
# # I = isocencentration point (cm)
# # D = average density of gradient
# sqrt(((BD-D)*2*BetaO/w2) + I^2)
# }
#
# # test
# min_BD_r = BD2distFromAxis(min_BD, ave_gradient_density, BetaO, angular_velocity, I)
# max_BD_r = BD2distFromAxis(max_BD, ave_gradient_density, BetaO, angular_velocity, I)
#
# cat('radius range for BD-min to BD-max: ', min_BD_r, 'to', max_BD_r, '\n')
# -
# ## distance from axis of rotation to tube height of BD 'band'
#
# * The band is angled in the tube, so the BD band in the gradient (angled tube) will touch the wall of the tube at a min/max height of h1 and h2. This function determines those tube height values.
# \begin{align}
# y_t =
# \end{align}
#
# * x = a distance from the axis of rotation
# * r = radius of cfg tube
# * D = max tube distance from axis of rotation
# * A = angle of tube to axis of rotation (degrees)
# + language="R"
#
# distFromAxis2angledTubePos = function(x, r, D, A){
# # converting distance from axis of rotation to cfg tube position (min & max of tube height)
# # x = a distance from the axis of rotation
# # r = radius of cfg tube
# # D = max tube distance from axis of rotation
# # A = angle of tube to axis of rotation (degrees)
#
# # Equation for finding the lower point of the band
# if(x >= D-(r*aspace::cos_d(A))-r) {
# d = x-(D-r)
# a = A-aspace::asin_d(d/r)
# LowH = r-r*aspace::cos_d(a)
# #print(LowH) ## This band will be in the rounded part
# }else{
# d = D-(r*aspace::cos_d(A))-r-x
# hc = d/aspace::sin_d(A)
# LowH = r+hc
# # print(LowH) ## This band will be in the cylinder part
# }
#
# # Equation for finding the upper band
# if(x > D-(r-r*aspace::cos_d(A))) {
# d = x-(D-r)
# a = (A)-(180-aspace::asin_d(d/r))
# HighH = r-r*aspace::cos_d(a)
# #print(HighH) ## This band will be in the rounded part
# }else{
# d = D-(r-r*aspace::cos_d(A))-x
# hc = d/aspace::sin_d(A)
# HighH = r+hc
# #print(HighH) ## This band will be in the cylinder part
# }
#
# return(c(LowH, HighH))
# }
#
#
# # test
# r = 0.65 # radius of tube (cm)
# D = 4.85 # distance from axis of rotation to furthest part of tube (cm)
# A = 27.95 # angle of tube to axis of rotation (degrees)
# x = 3.5 # some distance from axis of rotation (from equation)
#
# pos = distFromAxis2angledTubePos(x, r, D, A)
# pos %>% print
# delta = pos[2] - pos[1]
# delta %>% print
# -
# ### Python version
# +
sin_d = lambda d : np.sin(np.deg2rad(d))
cos_d = lambda d : np.cos(np.deg2rad(d))
asin_d = lambda x : np.arcsin(x) * 180/np.pi #np.arcsin(np.deg2rad(d))
acos_d = lambda x : np.arccos(x) * 180/np.pi #np.arccos(np.deg2rad(d))
def axisDist2angledTubePos(x, tube_radius, r_max, A):
if np.isnan(x):
return (x, x)
if(x >= r_max - (tube_radius * cos_d(A)) - tube_radius):
# band in rounded bottom of cfg tube
d = x - (r_max - tube_radius)
a = A - asin_d(d / tube_radius)
LowH = tube_radius - tube_radius * cos_d(a)
#print LowH
else:
# band in cylinder of cfg tube
d = r_max - (tube_radius * cos_d(A)) - tube_radius - x
h_c = d/sin_d(A)
LowH = tube_radius + h_c
# print LowH
if(x > r_max - (tube_radius - tube_radius * cos_d(A))):
# Equation for finding the upper band
d = x - (r_max - tube_radius)
a = A - (180 - asin_d(d/tube_radius))
HighH = tube_radius - tube_radius * cos_d(a)
#print HighH
else:
# This band will be in the cylinder part
d = r_max - (tube_radius - tube_radius * cos_d(A)) - x
h_c = d/sin_d(A)
HighH = tube_radius + h_c
#print(HighH)
return(LowH, HighH)
# test
r = 0.65 # radius of tube (cm)
D = 4.85 # distance from axis of rotation to furthest part of tube (cm)
A = 27.95 # angle of tube to axis of rotation (degrees)
x = 3.5 # some distance from axis of rotation (from equation)
ret = axisDist2angledTubePos(x, r, D, A)
print(ret)
delta = ret[1] - ret[0]
print(delta)
# -
# ## Converting distance from axis of rotation to angled tube volume
# ### Python
# +
def _SphVol(t, r, p2, R12):
# helper function for axisDist2angledTubeVol
v1 = t*((2*r)-t)/2
v2 = 2*np.pi*((p2-t)/R12)
v3 = np.sin(2*np.pi*((p2-t)/R12))
return v1 * (v2 - v3)
def _CylWedVol(t, r, b, h):
# helper function for axisDist2angledTubeVol
return 2*(h*(t-r+b)/ b) * np.sqrt(r**2-t**2)
def axisDist2angledTubeVol(x, r, D, A):
"""Convert distance from axis of rotation to volume of gradient
where the BD is >= to the provided BD.
Parameters
----------
x : float
distance from axis of rotation (cm)
r : float
cfg tube radius (cm)
D : float
max distance from axis of rotation (cm)
A : float
cdf tube angle in rotor (degrees)
Returns
-------
volume (ml) occupied by gradient heavier or as heavy as at that point.
Note: nan returned if x = nan
"""
# return nan if nan provided
if np.isnan(x):
return x
a = np.deg2rad(A)
p1 = r-(r*np.cos(a))
p2 = r+(r*np.cos(a))
R12 = p2-p1
d = D-x
D1 = D-p1
D2 = D-p2
if x < D2:
if a == 0:
z = 1
else:
z = np.sin(a)
h1 = (D2-x)/z
h2 = (D1-x)/z
volume1 = (2/3.0)*np.pi*r**3
volume2 = (0.5)*np.pi*r**2*(h1+h2)
volume = volume1+volume2
elif D1 >= x >= D2:
volume1 = (1/3.0)*np.pi*p1**2*(3*r-p1)
volume2 = quad(_SphVol, p1, d, args=(r, p2, R12))
b = (d-p1)/np.cos(a)
if a == 0:
h = b
else:
h = b/np.tan(a)
volume3 = quad(_CylWedVol, r-b, r, args=(r, b, h))
volume = volume1+volume2[0]+volume3[0]
elif D >= x > D1:
volume = (1/3.0)*np.pi*d**2*(3*r-d)
elif x > D:
volume = np.nan
else:
volume = np.nan
# status
if np.isnan(volume):
lmsg = 'axisDist2angledTubeVol: nan returned for x value: {}\n'
sys.stderr.write(lmsg.format(x))
return volume
# test
## fixed-angle rotor
r = 0.65 # radius of tube (cm)
D = 4.85 # distance from axis of rotation to furthest part of tube
A = 27.95 # angle of tube to axis of rotation (degrees)
x = 3.5 # some distance from axis of rotation (from equation)
ret = axisDist2angledTubeVol(x, r, D, A)
print(ret)
## vertical rotor
#x = 7.66
x = 8.5
r = 0.65
D = 8.79
A = 0
ret = axisDist2angledTubeVol(x, r, D, A)
print(ret)
# -
# ## Converting tube volume to vertical tube height
# ### Python
# +
# converting cylinder volume to height
def cylVol2height(v, r):
# v = volume (ml)
# r = tube radius (cm)
h = v / (np.pi * r**2)
return h
# test
cylVol2height(0.1, 0.65)
# +
# converting sphere cap volume to sphere height
from scipy import optimize
def sphereCapVol2height(v, r):
# v = volume (ml)
# r = tube radius (cm)
# h**3 - 3*r*h**2 + (3v / pi) = 0
f = lambda x : x**3 - 3*r*x**2 + 3*v/np.pi
try:
root = optimize.brentq(f, 0, r*2, maxiter=1000)
except ValueError:
msg = 'WARNING: not roots for volume {}\n'
sys.stderr.write(msg.format(v))
root = np.nan
return(root)
# test
sphereCapVol2heightV = np.vectorize(sphereCapVol2height)
heights = np.arange(0, 0.65**2, 0.1)
sphereCapVol2heightV(heights, 0.65)
# +
# convert liquid volume in vertical cfg tube to tube height
def tubeVol2height(v, r):
# v = volume (ml)
# r = tube radius (cm)
sphere_cap_vol = (4/3 * np.pi * r**3)/2
if v <= sphere_cap_vol:
# height does not extend to cylinder
h = sphereCapVol2height(v, r)
else:
# height = sphere_cap + cylinder
sphere_cap_height = sphereCapVol2height(sphere_cap_vol, r)
h = sphere_cap_height + cylVol2height(v - sphere_cap_vol, r)
return(h)
# test
vol = 0.1 # 100 ul
vols = np.arange(0, 4+vol, vol)
tubeVol2heightV = np.vectorize(tubeVol2height)
tubeVol2heightV(vols, r=0.65)
# -
# # Test run of SIPSim DBL
# ## Angled rotor
runDir = '/home/nick/notebook/SIPSim/t/genome100/'
# !cd $runDir; \
# SIPSim DBL \
# --np 4 \
# ampFrag_skewN90-25-n5-nS_dif_kde.pkl \
# > ampFrag_skewN90-25-n5-nS_dif_DBL_kde.pkl
# + magic_args="-w 600 -h 450" language="R"
# inFile = '/home/nick/notebook/SIPSim/t/genome100/DBL_index.txt'
# df = read.delim(inFile, sep='\t') %>%
# gather(pos, vert_grad_BD, vert_gradient_BD_low, vert_gradient_BD_high)
#
# # example
# df.ex = data.frame('DBL_BD' = c(1.675, 1.769), 'vert_grad_BD' = c(1.75, 1.75))
#
# # plot
# p.TLA = ggplot(df, aes(DBL_BD, vert_grad_BD, color=pos, group=DBL_BD)) +
# geom_line(color='black', size=1) +
# geom_point(data=df.ex, color='red', size=4) +
# geom_line(data=df.ex, aes(group=vert_grad_BD), color='red', linetype='dashed', size=1.2) +
# #geom_vline(xintercept=1.774, linetype='dashed', alpha=0.5, color='blue') + # theoretical max fragment BD
# #scale_y_reverse(limits=c(1.85, 1.50)) +
# scale_y_reverse() +
# scale_x_continuous(limits=c(1.63, 1.77)) +
# labs(x='BD of DBL',
# y='BD of vertical gradient\n(during fractionation)',
# title='TLA-110, Beckman fixed-angle rotor') +
# theme_bw() +
# theme(
# text = element_text(size=16)
# )
# p.TLA
# + magic_args="-i workDir" language="R"
# # saving figure
# F = file.path(workDir, 'DBL_TLA110.pdf')
# ggsave(F, p.TLA, width=6, height=4.5)
# cat('File written:', F, '\n')
# -
# #### Notes
#
# * The dashed line provides an example of the 'true' BD of fragments contained in the DBL at the gradient density of 1.7 when the gradient is in the vertically oriented during fractionation.
# ## Vertical rotor
#
# * VTi 65.2, Beckman rotor
# * Refs:
# * >http://www.nature.com/ismej/journal/v1/n6/full/ismej200765a.html
# * ><NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, et al. (2007). DNA stable-isotope probing. Nat Protocols 2: 860–866.
# * params:
# * tube width = 1.3 cm
# * tube height = 5.1 cm
# * tube volume = 5.1 ml
# * r_min = 7.47 cm
# * r_max = 8.79 cm
# * final density = 1.725
# * speed = 177000 g_av (42500 rpm)
# * angular velocity = $((2 * 3.14159 * R)/60)^2$ = 19807714
# * time = 40 hr
runDir = '/home/nick/notebook/SIPSim/t/genome100/'
# !cd $runDir; \
# SIPSim DBL \
# -D 1.725 \
# -w 19807714 \
# --tube_height 5.1 \
# --r_min 7.47 \
# --r_max 8.79 \
# --vertical \
# --np 4 \
# ampFrag_skewN90-25-n5-nS_dif_kde.pkl \
# > ampFrag_skewN90-25-n5-nS_dif_DBL_kde_VERT.pkl
# + magic_args="-w 600" language="R"
# inFile = '/home/nick/notebook/SIPSim/t/genome100/DBL_index.txt'
# df = read.delim(inFile, sep='\t') %>%
# gather(pos, vert_grad_BD, vert_gradient_BD_low, vert_gradient_BD_high)
#
# # example
# df.ex = data.frame('DBL_BD' = c(1.638, 1.769), 'vert_grad_BD' = c(1.75, 1.75))
#
# # plot
# p.VTi = ggplot(df, aes(DBL_BD, vert_grad_BD, color=pos, group=DBL_BD)) +
# geom_line(color='black', size=1) +
# geom_point(data=df.ex, color='red', size=4) +
# geom_line(data=df.ex, aes(group=vert_grad_BD), color='red', linetype='dashed', size=1.2) +
# #scale_y_reverse(limits=c(1.85, 1.50)) +
# scale_y_reverse() +
# scale_x_continuous(limits=c(1.63, 1.77)) +
# labs(x='BD of DBL', y='BD of vertical gradient\n(during fractionation)',
# title='VTi 65.2, Beckman vertical rotor') +
# theme_bw() +
# theme(
# text = element_text(size=16)
# )
# p.VTi
# + magic_args="-i workDir" language="R"
# # saving figure
# F = file.path(workDir, 'DBL_VTi65.2.pdf')
# ggsave(F, p.VTi, width=6, height=4.5)
# cat('File written:', F, '\n')
# -
# #### Notes
#
# * The dashed line provides an example of the 'true' BD of fragments contained in the DBL at the gradient density of 1.7 when the gradient is in the vertically oriented during fractionation.
# * WARNING: the DBL simulation makes the simplifying assumption of a 2d tube object and finds the vertical distance that a band spans in the tube, which sets the span of DBL contamination in a fixed-angle rotor. However, for vertical tubes, the DBL would probably be more accurately modeled from a 3d representation of the tube.
# * Regardless, there would be substantially more DBL 'smearing' with a vertical rotor than a fixed-angle rotor.
# ***
# # Misc
# ## DNA diffusion
#
# * sedimentation coefficient of DNA (S)
# * $S = 2.8 + (0.00834 * M^{0.479})$
# * where
# * M = molecular weight of DNA
# * OR $S = 2.8 + (0.00834 * (L*666)^{0.479})$
# * where
# * L = length of DNA
# * Svedberg's equation
# * $s/D = \frac{M(1-\bar{V}p)}{RT}$
# * where
# * s = sedimentation coefficient
# * D = diffusion coefficient
# * M = molecular weight
# * $\bar{V} = 1/\rho_p$
# * $\rho_p$ = density of the sphere
# * p = density of the liquid
# * R = universal gas constant
# * T = absolute temperature
# * Finding diffusion coefficient of DNA in CsCl ($\mu m^2 / s$)
# * $D = \frac{RT}{M(1-\bar{V}p)}*s$
# * where
# * R = 8.3144598 (J mol^-1 K^-1)
# * T = 293.15 (K)
# * p = 1.7 (Buckley lab gradients)
# * $\bar{V} = 1/\rho_p$
# * $\rho_p$ = 1.99
# * $s = 2.8 + (0.00834 * (L*666)^{0.479})$
# * L = DNA length (bp)
#
# + magic_args="-h 300" language="R"
#
# length2MW = function(L){ L * 666 }
#
# length2sedCoef = function(L){
# 2.8 + (0.00834 * (L*666)**0.479)
# }
#
# MW2diffuseCoef = function(L, p, R=8.3144598, T=293.15){
# V = 1/1.99
# M = length2MW(L)
# s = length2sedCoef(L)
# (R*T)/(M*(1-V*p)) * s
# }
#
# # test
# L = seq(100, 50000, 100)
# p = 1.7
# D = sapply(L, MW2diffuseCoef, p=p)
# df = data.frame('L' = L, 'D' = D)
#
#
# # plotting
# ggplot(df, aes(L, D)) +
# geom_point() +
# geom_line(alpha=0.5) +
# theme_bw() +
# theme(
# text = element_text(size=16)
# )
# -
# ## Calculating diffusion from DBL
#
# * Einstein-Smoluchowski relation
# * $t = \frac{z^2}{0.9 * D}$
# * where
# * t = time (sec)
# * z = mean deviation of molecules from starting position
# * D = diffusion coefficient (cm^2 s^-1)
# * rewritten: $z = \sqrt{0.9*D*t}$
# + language="R"
#
# # converting D to cm^2/s
# df$D_cm = df$D * 1e-5
#
# # time periods (sec)
# t = seq(1, 300, 10)
#
# # calculating z (cm)
# ES = function(D, t){
# sqrt(0.9 * D * t)
# }
# df2 = expand.grid(df$D_cm, t)
# colnames(df2) = c('D_cm', 't')
# df2$z = mapply(ES, df2$D_cm, df2$t)
# tmp = expand.grid(df$L, t)
#
# # adding variable
# df2$L = tmp$Var1
# df2$t_min = df2$t / 60
# df2$z_uM = df2$z / 1e-5
#
# ## plotting
# ggplot(df2, aes(t_min, z_uM, color=L, group=L)) +
# #geom_point(size=1.5) +
# geom_line() +
# labs(x='Time (minutes)',
# y='mean deviation of molecules\nfrom starting position (uM)') +
# scale_color_continuous('DNA fragment\nlength (bp)') +
# theme_bw() +
# theme(
# text = element_text(size=16)
# )
# + magic_args="-w 800" language="R"
# ## plotting
# ggplot(df2, aes(L, z_uM, color=t_min, group=t_min)) +
# #geom_point(size=1.5) +
# geom_line() +
# labs(x='DNA fragment length (bp)',
# y='mean deviation of molecules\nfrom starting position (uM)') +
# scale_color_continuous('Time\n(minutes)') +
# theme_bw() +
# theme(
# text = element_text(size=16)
# )
| ipynb/theory/diff_bound_layer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.9 64-bit (''bert'': virtualenv)'
# name: python36964bitbertvirtualenv5824b8894d5b4b1c8bf62ae2f0b6d2a4
# ---
import pandas as pd
import numpy as np
from analysis_utils import *
PAREDAO = "paredao15"
CAND1_PATH = "data/paredao15/ivy.csv"
CAND2_PATH = "data/paredao15/thelma.csv"
CAND3_PATH = "data/paredao15/rafa.csv"
DATE = 4
IGNORE_HASHTAGS = ["#bbb20", "#redebbb", "#bbb2020"]
candidate1_df = pd.read_csv(CAND1_PATH)
candidate2_df = pd.read_csv(CAND2_PATH)
candidate3_df = pd.read_csv(CAND3_PATH)
cand1 = candidate1_df[["tweet", "sentiment", "date", "likes_count", "retweets_count", "hashtags"]]
cand2 = candidate2_df[["tweet", "sentiment", "date", "likes_count", "retweets_count", "hashtags"]]
cand3 = candidate3_df[["tweet", "sentiment", "date", "likes_count", "retweets_count", "hashtags"]]
# # Ivy (Eliminada)
cand1["sentiment"].hist()
# # Thelma
cand2["sentiment"].hist()
# # Rafa
cand3["sentiment"].hist()
# # Quantidades absolutas
candidates = {"ivy": cand1, "thelma": cand2, "rafa": cand3}
qtds_df = get_raw_quantities(candidates)
qtds_df
qtds_df.plot.bar(rot=45, color=['green', 'gray', 'red'])
# # Porcentagens em relação aos total de tweets de cada candidato
pcts_df = get_pct_by_candidate(candidates)
pcts_df
pcts_df.plot.bar(rot=45, color=['green', 'gray', 'red'])
# # Porcentagens em relação ao total de tweets por categoria
qtds_df_copy = qtds_df.copy()
qtds_df["positivos"] /= qtds_df["positivos"].sum()
qtds_df["neutros"] /= qtds_df["neutros"].sum()
qtds_df["negativos"] /= qtds_df["negativos"].sum()
qtds_df
qtds_df.plot.bar(rot=45, color=['green', 'gray', 'red'])
# # Tweets por dia
names = list(candidates.keys())
tweets_by_day_df = get_tweets_by_day(candidates[names[0]], names[0])
for name in names[1:]:
current = get_tweets_by_day(candidates[name], name)
tweets_by_day_df = tweets_by_day_df.append(current)
tweets_by_day_df.transpose().plot()
# # Análise de hashtags
# +
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = (20,10)
unique_df = get_unique_hashtags(list(candidates.values()))
unique_df.drop(index=IGNORE_HASHTAGS, inplace=True)
unique_df.sort_values(by="quantidade", ascending=False).head(30).plot.bar(rot=45)
# -
alias = {name: name for name in candidates.keys()}
fica_fora_df = get_fica_fora_quantities(unique_df, alias)
fica_fora_df
# # Seleção de atributos
atributes_df = qtds_df_copy.join(pcts_df, rsuffix="_individual_pct")
atributes_df = atributes_df.join(qtds_df, rsuffix="_global_pct")
atributes_df = atributes_df.join(tweets_by_day_df)
atributes_df = atributes_df.join(fica_fora_df)
raw_participantes_info = get_participantes_info()[DATE]
print("Seguidores atualizados em:", raw_participantes_info["date"])
participantes_info = raw_participantes_info["infos"]
paredoes_info = get_paredoes_info()
followers = [participantes_info[participante]["seguidores"] for participante in atributes_df.index]
likes = [get_likes_count(candidates[participante]) for participante in atributes_df.index]
retweets = [get_retweets_count(candidates[participante]) for participante in atributes_df.index]
paredao_info = paredoes_info[PAREDAO]["candidatos"]
results_info = {candidate["nome"]: candidate["porcentagem"]/100 for candidate in paredao_info}
rejection = [results_info[participante] for participante in atributes_df.index]
atributes_df["likes"] = likes
atributes_df["retweets"] = retweets
atributes_df["seguidores"] = followers
atributes_df["rejeicao"] = rejection
atributes_df
atributes_df.to_csv("data/{}/paredao_atributes.csv".format(PAREDAO))
| analysis/paredao15.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import json
from copy import deepcopy, copy
import geopandas as gpd
from covidcaremap.constants import *
from covidcaremap.data import processed_data_path, published_data_path
# -
# # Generate CovidCareMap facility data
#
# The CovidCareMap (CCM) facility data describes a facility's capacity in 3 different scenarios: Conventional, Contingency and Crisis.
#
# ## Note: This is a work in progress.
#
# Methods taken from [1].
#
# [1] https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4636910/#R20
with open(published_data_path('us_healthcare_capacity-facility-CovidCareMap.geojson')) as f:
geojson = json.loads(f.read())
# Show the fields we'll be working with.
geojson['features'][1]
# ## Methods
#
# We use the methods in [1], but instead of starting with staffing numbers and working towards calculating "# of patients treatable", we work backwards and determine staffing requirements based on the # of patients being treated by a hospital that is saturated based on the ICU bed available determined by the Level of Care scenario.
#
# 
# *Figure from [1]*
#
# ### Scenario ICU Beds Available
#
# For each scenario, we estimate the total number of beds at the facility that will be used as an ICU bed (whether it is a true ICU bed or not), and use an occupancy rate to determine a surge percentage. This differs from the methodology in [1] as they use a surge percentage of 10% and 20% in their estimates. Here we use the equation `100% - {Occupancy Rate %}` to estimate the number of beds available for use in a PHE. Similar to [1], we tansfer this estimate to staffing needs. If no occupancy data is available for the facility, the 10% figure from [1] is used.
#
# The total ICU beds considered vary by the level of care scenario (see below).
#
# ### Ventilators Needed
#
# We assume each licensed ICU bed has a ventilator. We therefore calcululate the number of ventilators needed by the total ICU beds considered for the scenario minus the total licensed ICU beds.
#
# We estimate the total licensed ICU beds based on the ratio of total licensed beds to total staffed beds applied to the known staffed ICU beds. **Note:** This methodology should be improved. One idea is to collect as much known licensed ICU bed counts from state data sources as possible and create estimates there from the known staffed ICU bed counts.
#
# ### Staff ratios
#
# The patient-per-staff ratio and shifts-per-day are directly taken from [1].
#
# ### Parameters by Scenario
#
# #### Conventional
#
# Total IU beds are determined using the ICU Staffed beds counts in the data.
# Available percentage is calculated using the ICU Occupance Rate information in the data.
#
# #### Contingency
#
# Total ICU beds are estimated by using the ratio of the US-wide counts in [1]. Availability is calculated using the same rates as the Conventional scenario.
#
# #### Crisis
#
# Total ICU beds are determined to be total licensed beds - every bed in the hospital is converted to an ICU bed. This would require more mechanical ventilators than the facility
# is assumed to have, which will be reflected in the estimated ventilators required for
# this scenario. The availability rate is calculated as the weighted average of the
# ICU occupancy rate and total occupancy rate duing Business as Usual (BAU), with the weights
# being based on the ratio of staffed ICU beds to total staffed beds and staffed non-ICU beds
# to total staffed beds.
#
# +
# The number of shifts each staff type works per day. Taken from [1]
SHIFTS_PER_DAY = 2
# From national counts in [1]
CONTINGENCY_BED_RATIO = 261790 / 81790.0
DEBUG = False
def pdebug(msg):
if DEBUG:
print(msg)
def printprops(props):
print(json.dumps(props, indent=4))
def get_staffed_icu_beds(props):
return props[CCM_STAFFED_ICU_BEDS_COLUMN]
def get_total_staffed_beds(props):
return props[CCM_STAFFED_BEDS_COLUMN]
def get_licensed_beds(props, check_against_staffed=True):
return props[CCM_LICENSED_BEDS_COLUMN]
def get_bed_occupancy_rate(props):
return props[CCM_BED_OCCUPANCY_COLUMN]
def get_icu_bed_occupancy_rate(props):
return props[CCM_ICU_BED_OCCUPANCY_COLUMN]
def construct_scenario_data(props):
pdebug(json.dumps(props, indent=4))
staffed_icu_beds = get_staffed_icu_beds(props)
total_staffed_beds = get_total_staffed_beds(props)
icu_occupancy_rate = get_icu_bed_occupancy_rate(props)
total_occupancy_rate = get_bed_occupancy_rate(props)
# If occupancy rate data is not available, use 20%
if total_occupancy_rate == 0.0:
total_occupancy_rate = 0.2
if icu_occupancy_rate == 0.0:
icu_occupancy_rate = 0.2
licensed_beds = get_licensed_beds(props)
pdebug('staffed beds {}'.format(total_staffed_beds))
pdebug('ICU OCC {}'.format(icu_occupancy_rate))
pdebug('OCC {}'.format(total_occupancy_rate))
contingency_icu_beds = staffed_icu_beds * CONTINGENCY_BED_RATIO
# Weighted average based on staffed bed counts and occupancy rates
try:
crisis_occupancy_rate = \
((staffed_icu_beds / total_staffed_beds) * icu_occupancy_rate) + \
((1 - (staffed_icu_beds / total_staffed_beds)) * total_occupancy_rate)
except:
print(json.dumps(props, indent=4))
print('Total staffed beds: {}'.format(total_staffed_beds))
print('total_occupancy_rate: {}'.format(total_occupancy_rate))
raise
pdebug('CRISIS OCC {}'.format(crisis_occupancy_rate))
return {
CONVENTIONAL: {
BEDS: {
'available percentage': (1 - icu_occupancy_rate),
'total': staffed_icu_beds,
},
PHYSICIANS: {
'available percentage': (1 - icu_occupancy_rate),
'patients per': [10, 15],
},
RESP_THERP: {
'available percentage': (1 - icu_occupancy_rate),
'patients per': [4, 6],
},
NURSE: {
'available percentage': (1 - icu_occupancy_rate),
'patients per': [1, 1],
}
},
CONTINGENCY: {
BEDS: {
'available percentage': (1 - icu_occupancy_rate),
'total': contingency_icu_beds,
},
PHYSICIANS: {
'available percentage': (1 - icu_occupancy_rate),
'patients per': [10, 15],
},
RESP_THERP: {
'available percentage': (1 - icu_occupancy_rate),
'patients per': [7, 9],
},
NURSE: {
'available percentage': (1 - icu_occupancy_rate),
'patients per': [2, 2],
}
},
CRISIS: {
BEDS: {
'available percentage': (1 - crisis_occupancy_rate),
'total': licensed_beds,
},
PHYSICIANS: {
'available percentage': (1 - crisis_occupancy_rate),
'patients per': [24, 24],
},
RESP_THERP: {
'available percentage': (1 - crisis_occupancy_rate),
'patients per': [4, 6],
},
NURSE: {
'available percentage': (1 - crisis_occupancy_rate),
'patients per': [1, 1],
}
}
}
# +
new_gj = deepcopy(geojson)
for feature in new_gj['features']:
props = feature['properties']
new_props = copy(props)
scenario_data = construct_scenario_data(props)
# Number of staffed ICU beds durig 'Business As Usual' (BAU)
staffed_icu_beds_BAU = get_staffed_icu_beds(props)
licensed_beds = get_licensed_beds(props)
# Total number of staffed beds during BAU
staffed_beds_BAU = get_total_staffed_beds(props)
# Estimate the total licensed ICU beds based on total/staffed ratio
licensed_to_staffed_ratio = (licensed_beds / staffed_beds_BAU) * staffed_beds_BAU
## TODO: Replace this with something better (as per note in Methods)
estimated_licensed_icu_beds = licensed_to_staffed_ratio * staffed_icu_beds_BAU
for scenario_name, scenario in scenario_data.items():
icu_beds_total = scenario[BEDS]['total']
icu_beds = icu_beds_total * scenario[BEDS]['available percentage']
new_props[construct_beds_field_name(scenario_name)] = round(icu_beds)
# Ventilator need
new_props[construct_vents_field_name(scenario_name)] = \
round(max(0, icu_beds - estimated_licensed_icu_beds))
for staff_need in STAFF:
patients_per = scenario[staff_need]['patients per']
available_percentage = scenario[staff_need]['available percentage']
if icu_beds <= 0:
min_need, max_need = 0, 0
else:
min_need, max_need = None, None
for patients in patients_per:
need_count = (icu_beds * available_percentage) / patients
need_count /= SHIFTS_PER_DAY
if min_need is None or need_count < min_need:
min_need = need_count
if max_need is None or max_need < need_count:
max_need = need_count
new_props[construct_staff_field_name(
scenario_name, staff_need, BOUND_LOWER)] = round(max(1, min_need))
new_props[construct_staff_field_name(
scenario_name, staff_need, BOUND_UPPER)] = round(max(1, max_need))
feature['properties'] = new_props
print(json.dumps(new_gj['features'][1], indent=4))
# -
# ### Write out files
# +
geojson_path = processed_data_path('CareModel_data-facility-CovidCareMap.geojson')
csv_path = processed_data_path('CareModel_data-facility-CovidCareMap.csv')
open(geojson_path, 'w').write(json.dumps(new_gj, indent=2))
# +
# Write out CSV
def get_lon(row):
return row['geometry'].x
def get_lat(row):
return row['geometry'].y
final_gdf = gpd.read_file(geojson_path)
final_gdf['Latitude'] = final_gdf.apply(get_lat, axis=1)
final_gdf['Longitude'] = final_gdf.apply(get_lon, axis=1)
final_df = final_gdf[CCM_CSV_COLUMNS + CAREMODEL_CAPACITY_COLUMNS()]
final_df.to_csv(csv_path, index=False)
# -
| notebooks/processing/Generate_CCM_CareModel_Facility_Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="FXlaqYYxwXqy" colab_type="text"
# This file contains about
#
#
# 1. User input
# 2. Conditional Statements (if_elif_else)
# 3. Comments
#
#
# + [markdown] id="1AATA_FDyJbp" colab_type="text"
# ### **Asking user-input**
# + id="rxmucLEfvr7z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 87} executionInfo={"status": "ok", "timestamp": 1594888572417, "user_tz": -390, "elapsed": 4936, "user": {"displayName": "<NAME>, AI, R & DS", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggh4U2DIE3BiNSBSVPM5e6QQKnQOTyp7A5mGHY=s64", "userId": "14938682159450874323"}} outputId="8c3f0c77-65f3-414b-8a2d-e2d7e0a47253"
# input( )
print('Do you know any class that teaches python basics?')
class_name = input()
print('I\'m currenlt learning python basics at', class_name)
# + [markdown] id="RdZeL-O8yPpm" colab_type="text"
# ### **Conditional statement (If_else_elif)**
# + id="nKtxnAwyxB0R" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} executionInfo={"status": "ok", "timestamp": 1594888675879, "user_tz": -390, "elapsed": 1084, "user": {"displayName": "<NAME>, AI, R & DS", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggh4U2DIE3BiNSBSVPM5e6QQKnQOTyp7A5mGHY=s64", "userId": "14938682159450874323"}} outputId="4d68f636-4a54-4b04-b93d-0dc0a31d50c4"
# if(condition)
age = 17
if age < 18:
print('You are not allowed to buy alcohol and beer')
# + id="9u8aoPypxd0u" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} executionInfo={"status": "ok", "timestamp": 1594888757223, "user_tz": -390, "elapsed": 927, "user": {"displayName": "<NAME>, AI, R & DS", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggh4U2DIE3BiNSBSVPM5e6QQKnQOTyp7A5mGHY=s64", "userId": "14938682159450874323"}} outputId="fcde6dc1-0358-4fa6-a838-07c8952f4943"
# elif(condition)
if age < 18:
print('You are not allowed to buy alcohol and beer')
elif age >=18:
print('You are allowed to buy but don\'t drink too much')
elif age == 80:
print('Too old to drink!')
# + id="KJsy3fSIxxvH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} executionInfo={"status": "ok", "timestamp": 1594888840639, "user_tz": -390, "elapsed": 726, "user": {"displayName": "AVAIRDS AV, AI, R & DS", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggh4U2DIE3BiNSBSVPM5e6QQKnQOTyp7A5mGHY=s64", "userId": "14938682159450874323"}} outputId="14a815c4-4fa4-4022-c4f5-ae03fd56f027"
# else
name = 'Albert'
if name is 'Johnny':
print('Welcome from home Johnny')
elif name is 'Tony':
print('Welcome from home Tony')
else:
print('Sorry I don\'t know you')
# + [markdown] id="VJ-2EVIyyXsv" colab_type="text"
# ### **Comments**
# + id="wYaqY8uvyGJg" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1594888969899, "user_tz": -390, "elapsed": 1066, "user": {"displayName": "AVAIRDS AV, AI, R & DS", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggh4U2DIE3BiNSBSVPM5e6QQKnQOTyp7A5mGHY=s64", "userId": "14938682159450874323"}}
# This is a single line comment
# This is a
# multiline comment
count = 4 # this variable is used to count the total number
# + id="0BCBCYUoylo3" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1594889017707, "user_tz": -390, "elapsed": 1137, "user": {"displayName": "AVAIRDS AV, AI, R & DS", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggh4U2DIE3BiNSBSVPM5e6QQKnQOTyp7A5mGHY=s64", "userId": "14938682159450874323"}}
# Using docstrings
def test():
"""
This function is used t add two numbers
"""
a = 10
b = 20
z = a + b
return z
# + id="KDOjqzlDysYh" colab_type="code" colab={}
| coding-exercises-avairds/week2/part1-conditional-statements-lists/userinput-if-elif-else-comments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Testing ways to speed up octree creation
# +
# %load_ext autoreload
# %autoreload 2
from octreeStreamOrg import octreeStream
# -
import datetime
o = octreeStream('/Users/ageller/VISUALIZATIONS/FIREdata/m12i_res7100/snapdir_600/snapshot_600.0.hdf5',
h5PartKey = 'PartType0', keyList = ['Coordinates', 'Density', 'Velocities'],
NNodeMax = 10000, NMemoryMax = 5e4, Nmax=1e5, verbose=1, minWidth=1e-4,
path='/Users/ageller/VISUALIZATIONS/octree_threejs_python/WebGL_octreePartition/src/data/junk/octreeNodes/Gas')
begin_time = datetime.datetime.now()
o.compileOctree()
end_time = datetime.datetime.now()
print('normal execution time : ',end_time - begin_time)
# ## Test writing files without pandas
#
# *This is offering a speed up. I will use this version as the default for now.*
import sys
sys.path.insert(1, '/Users/ageller/VISUALIZATIONS/octree_threejs_python/python_octreeStream/noPandasTest/')
# %load_ext autoreload
# %autoreload 2
from octreeStreamNoPandas import octreeStream as octreeStreamNoPandas
oP1 = octreeStreamNoPandas('/Users/ageller/VISUALIZATIONS/FIREdata/m12i_res7100/snapdir_600/snapshot_600.0.hdf5',
h5PartKey = 'PartType0', keyList = ['Coordinates', 'Density', 'Velocities'],
NNodeMax = 10000, NMemoryMax = 5e4, Nmax=1e5, verbose=1, minWidth=1e-4,
cleanDir = True,
path='/Users/ageller/VISUALIZATIONS/octree_threejs_python/WebGL_octreePartition/src/data/junk/octreeNodes/Gas')
begin_timeP1 = datetime.datetime.now()
oP1.compileOctree()
end_timeP1 = datetime.datetime.now()
print('no pandas execution time : ',end_timeP1 - begin_timeP1)
# ## Try with multiprocessing
#
# *Starting from the no pandas version.*
#
# *This is somehow WAAAYYY slower!!! (even with multiepl cores in use.) I guess it's because of all the shared memory?*
import sys
import datetime
sys.path.insert(1, '/Users/ageller/VISUALIZATIONS/octree_threejs_python/python_octreeStream/multiprocessingTest/')
# %load_ext autoreload
# %autoreload 2
from octreeStreamMultiprocessing import octreeStream as octreeStreamMultiprocessing
oM1 = octreeStreamMultiprocessing('/Users/ageller/VISUALIZATIONS/FIREdata/m12i_res7100/snapdir_600/snapshot_600.0.hdf5',
h5PartKey = 'PartType0', keyList = ['Coordinates', 'Density', 'Velocities'],
NNodeMax = 10000, NMemoryMax = 5e4, Nmax=1e5, verbose=1, minWidth=1e-4,
cleanDir = True,
path='/Users/ageller/VISUALIZATIONS/octree_threejs_python/WebGL_octreePartition/src/data/junk/octreeNodes/Gas')
begin_timeM1 = datetime.datetime.now()
oM1.compileOctree()
end_timeM1 = datetime.datetime.now()
print('multiprocessing execution time : ',end_timeM1 - begin_timeM1)
# ## Trying cythonize (not working)
#
# https://cython.readthedocs.io/en/latest/src/tutorial/cython_tutorial.html
#
# Had to install Visual Studio's C++ compiler:
# https://visualstudio.microsoft.com/visual-cpp-build-tools/
#
#
# ```
# $ cp noPandaTest/octreeStreamNoPandas.py cythonTest/octreeStreamCython1.pyx
# ```
#
# And created a setup.py file. Then
#
# ```
# $ python setup.py build_ext --inplace
#
# ```
#
# (I will probably want to try to rewrite the code with cython syntax ... but let's see how fast this test is.)
#
import sys
sys.path.insert(1, '/Users/ageller/VISUALIZATIONS/octree_threejs_python/python_octreeStream/cythonTest/')
# #%load_ext autoreload
# #%autoreload 2
from octreeStreamCython1 import octreeStream as octreeStreamCython1
oC1 = octreeStreamCython1('/Users/ageller/VISUALIZATIONS/FIREdata/m12i_res7100/snapdir_600/snapshot_600.0.hdf5',
h5PartKey = 'PartType0', keyList = ['Coordinates', 'Density', 'Velocities'],
NNodeMax = 10000, NMemoryMax = 5e4, Nmax=1e5, verbose=1, minWidth=1e-4,
cleanDir = True,
path='/Users/ageller/VISUALIZATIONS/octree_threejs_python/WebGL_octreePartition/src/data/junk/octreeNodes/Gas')
begin_timeC1 = datetime.datetime.now()
oC1.compileOctree()
end_timeC1 = datetime.datetime.now()
print('cythonize execution time : ',end_timeC1 - begin_timeC1)
# ## Trying with numba (not working)
#
# https://numba.pydata.org/numba-doc/dev/user/5minguide.html
#
#
# ```
# $ cp octreeStream.py numbaTest/octreeStreamNumba1.py
# ```
#
# added
# ```
# @jit(nopython=True)
# ```
# throughout the code
#
import datetime
import sys
sys.path.insert(1, '/Users/ageller/VISUALIZATIONS/octree_threejs_python/python_octreeStream/numbaTest/')
# %load_ext autoreload
# %autoreload 2
from octreeStreamNumba1 import octreeStream as octreeStreamNumba1
oN1 = octreeStreamNumba1('/Users/ageller/VISUALIZATIONS/FIREdata/m12i_res7100/snapdir_600/snapshot_600.0.hdf5',
h5PartKey = 'PartType0', keyList = ['Coordinates', 'Density', 'Velocities'],
NNodeMax = 10000, NMemoryMax = 5e6, Nmax=1e5, verbose=1, minWidth=1e-4,
path='/Users/ageller/VISUALIZATIONS/octree_threejs_python/WebGL_octreePartition/src/data/junk/octreeNodes/Gas')
begin_timeN1 = datetime.datetime.now()
oN1.compileOctree()
end_timeN1 = datetime.datetime.now()
print('numba execution time : ',end_timeN1 - begin_timeN1)
| octreeStreamSpeedup_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Basic Mechanics
# -------------------------
#
# Let's get an overivew of how authentication/etc work with Planet's library and set the groundwork for how we're going to interact with the Planet Data API.
#
# Note that we're trying to stay relatively language agnostic, so we're still going to build things up mostly from scratch. We'll make use of Python's standard library functionality and we'll use the `requests` library to simplify communicating with the api a bit. We're not going to focus heavily on Planet's Python client library or other high-level ways of accessing the data. Our goal here is to explore and understand the underlying REST api.
#
# To start with, let's make a request against the top-level endpoint for the Planet data api. It returns some information about the api and doesn't require authentication (we'll cover that soon).
#
# +
# Just for nicer display of response dicts -- This is a standard library module
from pprint import pprint
# Not part of the standard library, but makes life much easier when working with http APIs
import requests
# Let's do this properly and raise an exception if we don't get a 200 (ok) response code
response = requests.get('https://api.planet.com/data/v1')
response.raise_for_status()
body = response.json()
# Nicer display than a bare print
pprint(body)
# -
# Authentication
# ----------------------
#
# Planet's APIs handle authentication primarily through API keys. It's a bit more secure than using your username and password, as the key only has access to our api (and not, say, deleting your account).
#
# You can find your api key by logging in to https://www.planet.com/account You should already have had an account created for you as part of the workshop signup. Your account will have access to a variety of data in California for the next month.
#
# The python client reads from the `PL_API_KEY` environment variable if it exists. We'll read the api key from that env variable rather than hard-coding it in these exercises. Let's go ahead and set that now:
#
# +
import os
# If you're following along with this notebook, you can enter your API Key on the following line, and uncomment it:
#os.environ['PL_API_KEY'] = "YOUR API KEY HERE"
# Setup the API Key from the `PL_API_KEY` environment variable
PLANET_API_KEY = os.getenv('PL_API_KEY')
# -
# To start with, let's look at what happens when we don't authenticate:
# +
# We'll explain this url later -- This is just a demo of something you can't access without auth
scene_url = 'https://api.planet.com/data/v1/item-types/PSScene4Band/items/20191010_183406_0f28'
response = requests.get(scene_url)
response.raise_for_status()
# -
# You can use your api key to authenticate through basic http authentication:
# Basic auth for our API expects api key as the username and no password
response = requests.get(scene_url, auth=(PLANET_API_KEY, ''))
response.raise_for_status()
# We can also use a session in the `requests` library that will store our auth and use it for all requests:
# +
session = requests.Session()
session.auth = (PLANET_API_KEY, '')
response = session.get(scene_url)
response.raise_for_status()
# -
# Rate Limiting and Retries
# --------------------------------------
#
# Planet's services limit the number of requests you can make in a short amount of time to avoid unintentional and intentional denial of service attacks. A 429 response code indicates that the service won't deliver results until you slow down.
#
# As an example, let's do something that will trigger a "slow down" response (429):
# +
# We'll use multiple threads to really hammer things... Otherwise it's unlikely to trigger.
from concurrent.futures import ThreadPoolExecutor
def make_request(ignored):
requests.get(scene_url, auth=(PLANET_API_KEY, '')).raise_for_status()
nthreads = 8
with ThreadPoolExecutor(nthreads) as executor:
for _ in executor.map(make_request, range(100)):
pass
# -
# We ask that you follow an exponential backoff pattern when this occurs -- E.g. on the first occurence, wait `n` seconds, on the second `n**2` seconds, on the third `n**3`, etc.
#
# I'm sure a lot of you have probably implemented similar retry functionality or have a library you often use to do so. There are also methods for this in our python client.
#
# However, to keep with the spirit of staying relatively low-level and using widely available tools, we'll set up the `requests` library to do this for us.
# +
import requests
from urllib3.util.retry import Retry
from requests.adapters import HTTPAdapter
session = requests.Session()
session.auth = (PLANET_API_KEY, '')
retries = Retry(total=5,
backoff_factor=0.2, # Exponential backoff scaled by 0.2
status_forcelist=[429]) # In practice, you may want other codes there too, e.g. 500s...
session.mount('https://', HTTPAdapter(max_retries=retries))
# -
# And just to demonstrate that we can now hammer the service in parallel and properly back off and retry when it asks us to:
# +
def make_request(ignored):
# Note that we're using the session we set up above
session.get(scene_url, auth=(PLANET_API_KEY, '')).raise_for_status()
nthreads = 8
with ThreadPoolExecutor(nthreads) as executor:
for _ in executor.map(make_request, range(100)):
pass
# -
# Setup for Future Exercises
# ----------------------------------------
#
# Taking what we've just walked through, we'll use the following as a bit of boilerplate to set up our later exercises. For actual use cases, you can see how this could form the start of an `APIClient` class. We're going to keep things as mininimal as possible in this tutorial, though.
# +
import os
from pprint import pprint
from urllib3.util.retry import Retry
import requests
from requests.adapters import HTTPAdapter
PLANET_API_URL = 'https://api.planet.com/data/v1'
def setup_session(api_key=None):
"""
Initialize a requests.Session that handles Planet api key auth and retries.
:param str api_key:
A Planet api key. Will be read from the PL_API_KEY env var if not specified.
:returns requests.Session session:
A Session instance optimized for use with Planet's api.
"""
if api_key is None:
api_key = os.getenv('PL_API_KEY')
session = requests.Session()
session.auth = (api_key, '')
retries = Retry(total=5,
backoff_factor=0.2,
status_forcelist=[429])
session.mount('https://', HTTPAdapter(max_retries=retries))
return session
session = setup_session() # Or pass in an api key if the environment variable isn't set
| notebooks/1_BasicMechanics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# importing necessary libraries
from keras.layers import Conv2D, UpSampling2D, InputLayer, Conv2DTranspose
from keras.layers import Activation, Dense, Dropout, Flatten
from keras.layers.normalization import BatchNormalization
from keras.models import Sequential
from keras.preprocessing.image import ImageDataGenerator, array_to_img, img_to_array, load_img
from skimage.color import rgb2lab, lab2rgb, rgb2gray, xyz2lab
from skimage.io import imsave
import numpy as np
import os
import random
import tensorflow as tf
# Building the CNN model
model = Sequential()
model.add(InputLayer(input_shape=(None, None, 1)))
model.add(Conv2D(8, (3, 3), activation='relu', padding='same', strides=2))
model.add(Conv2D(8, (3, 3), activation='relu', padding='same'))
model.add(Conv2D(16, (3, 3), activation='relu', padding='same'))
model.add(Conv2D(16, (3, 3), activation='relu', padding='same', strides=2))
model.add(Conv2D(32, (3, 3), activation='relu', padding='same'))
model.add(Conv2D(32, (3, 3), activation='relu', padding='same', strides=2))
model.add(UpSampling2D((2, 2)))
model.add(Conv2D(32, (3, 3), activation='relu', padding='same'))
model.add(UpSampling2D((2, 2)))
model.add(Conv2D(16, (3, 3), activation='relu', padding='same'))
model.add(UpSampling2D((2, 2)))
model.add(Conv2D(2, (3, 3), activation='tanh', padding='same'))
# Finish model
model.compile(optimizer='rmsprop',loss='mse')
def process_image(image_id):
image = img_to_array(load_img('dataset/'+image_id+'.jpg'))
image = np.array(image, dtype=float)
# Process the RGB image to LAB
X = rgb2lab(1.0/255*image)[:,:,0]
Y = rgb2lab(1.0/255*image)[:,:,1:]
Y /= 128
X = X.reshape(1, 400, 400, 1)
Y = Y.reshape(1, 400, 400, 2)
return X, Y
# +
# Get images
choices = ['woman', 'man', 'swim']
for train_image in choices:
train_X, train_Y = process_image(train_image)
# Train on each pixel of one image
model.fit(x=train_X, y=train_Y, batch_size=1, epochs=1000)
for test_image in choices:
test_X, test_Y = process_image(test_image)
print(model.evaluate(test_X, test_Y, batch_size=1))
output = model.predict(test_X)
output *= 128
# Output colorizations
cur = np.zeros((400, 400, 3))
cur[:,:,0] = test_X[0][:,:,0]
cur[:,:,1:] = output[0]
imsave('results/normal_train_'+train_image+'_test_'+test_image+'.png', lab2rgb(cur))
imsave('results/gray_train_'+train_image+'_test_'+test_image+'.png', rgb2gray(lab2rgb(cur)))
# -
| alpha/alpha_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <b>Calcule a integral dada</b>
# $\int_{0}^{5} (3x + 2)dx $
# $\frac{3x^2}{2} + 2x, 0 \leq X \leq 5$
# $\frac{3\cdot(5)^2}{2} + 2\cdot(5)- \frac{3 \cdot(0)^2}{2} + 2 \cdot 0$
# $\frac{75}{2} + 10$
# $\frac{95}{2}$
| Problemas 5.3/03.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:root] *
# language: python
# name: conda-root-py
# ---
# This is a sanity check, as well as usage demo, for the following estimators:
# - MISED = Mean Integrated Square Error for Derivatives (i.e. Density Derivative Estimation)
# - LSDDR = Least Squares Density Derivative Ratio (Estimation)
# +
# Imports
import numpy as np
from scipy import stats
import matplotlib.pyplot as plt
# %matplotlib inline
# -
from MISEDpython.MISED import MISED_full as densderiv
from LSDDRpython.LSDDR_full import LSDDR_full as densderivratio
# ### MISED
# +
# set the distribution
n = 1000
mu = 0.3
sigma = 0.7
# generate the data
t = np.linspace(-2, 2, 500)
X = np.random.normal(mu, sigma, n)
# Exact pdf
pt = stats.norm.pdf(t, loc=mu, scale=sigma)
# Exact density derivative
# (Formula is simple because of normal distribution!)
ddt = pt * (mu - t) / sigma**2
# Exact pdf at sample points
px = stats.norm.pdf(X, loc=mu, scale=sigma)
# -
# Compute kernel approximation
kerdens = densderiv(X)
dhh = kerdens.compute_density_deriv(t)
print(dhh.shape)
# +
# draw the first figure: densities and samples
fig = plt.figure(figsize=(16,5))
ax1 = fig.add_subplot(1,2,1)
hl, = ax1.plot(t, pt, linewidth=1, color='r', zorder=1)
ax1.scatter(X, px, c=u'r', marker='v', s=5, zorder=2)
ax1.legend([hl], ['p(x)'])
plt.xlabel('x')
# draw the second figure: true and estimate density derivative
ax2 = fig.add_subplot(1,2,2)
hl3, = ax2.plot(t, ddt, linewidth=2, color='k')
hl4, = ax2.plot(t, dhh[0], linewidth=2, color='c')
ax2.legend([hl3, hl4], ['d(x)', 'd_{est}(x)' ])
plt.xlabel('x')
plt.show()
# -
# ### LSDDR
# +
# set the distribution
n = 1000
mu = 0.5
sigma = 0.7
# generate the data
t = np.linspace(-1, 2, 500)
X = np.random.normal(mu, sigma, n)
# Exact pdf
pt = stats.norm.pdf(t, loc=mu, scale=sigma)
# Exact density derivative ratio
# (Formula is simple because of normal distribution!)
ddt = (mu - t) / sigma**2
# Exact pdf at sample points
px = stats.norm.pdf(X, loc=mu, scale=sigma)
# -
# Compute kernel approximation
kerdens = densderivratio(X)
dhh = kerdens.compute_density_deriv_ratio(t)
print(dhh.shape)
# +
# draw the first figure: densities and samples
fig = plt.figure(figsize=(16,5))
ax1 = fig.add_subplot(1,2,1)
hl, = ax1.plot(t, pt, linewidth=1, color='r', zorder=1)
ax1.scatter(X, px, c=u'r', marker='v', s=5, zorder=2)
ax1.legend([hl], ['p(x)'])
plt.xlabel('x')
# draw the second figure: true and estimate density derivative ratio
ax2 = fig.add_subplot(1,2,2)
hl3, = ax2.plot(t, ddt, linewidth=2, color='k')
hl4, = ax2.plot(t, dhh[0], linewidth=2, color='c')
ax2.legend([hl3, hl4], ['d(x)', 'd_{est}(x)' ])
plt.xlabel('x')
plt.show()
# -
| Test_kernel_methods.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Temperature effects
#
# Try varying the temperature of the simulation, through values of 3, 30, and 300 K, and watch the scattering profile for the presence of Bragg peaks in the data when the system is cold enough to freeze.
# Compare this to the radial distribution functions discussed previously.
# +
import numpy as np
def debye(qvalues, xposition, yposition, box_length):
"""
Calculates the scattering profile from the
simulation
positions.
Parameters
----------
qvalues: float, array-like
The q-vectors over which the scattering
should be calculated
xposition: float, array-like
The positions of the particles in the x-axis
yposition: float, array-like
The positions of the particles in the y-axis
box_length: float
The length of the simulation square
Returns
-------
intensity: float, array-like
The calculated scattered intensity
"""
intensity = np.zeros_like(qvalues)
for e, q in enumerate(qvalues):
for m in range(0, xposition.size-1):
for n in range(m+1, xposition.size):
xdist = xposition[n] - xposition[m]
xdist = xdist % box_length
ydist = yposition[n] - yposition[m]
ydist = ydist % box_length
r_mn = np.sqrt(np.square(xdist) + np.square(ydist))
intensity[e] += 1 * 1 * np.sin(
r_mn * q) / (r_mn * q)
if intensity[e] < 0:
intensity[e] = 0
return intensity
from pylj import md, sample
def md_simulation(number_of_particles, temperature,
box_length, number_of_steps,
sample_frequency):
"""
Runs a molecular dynamics simulation in using the pylj
molecular dynamics engine.
Parameters
----------
number_of_particles: int
The number of particles in the simulation
temperature: float
The temperature for the initialisation and
thermostating
box_length: float
The length of the simulation square
number_of_steps: int
The number of molecular dynamics steps to run
sample_frequency:
How regularly the visualisation should be updated
Returns
-------
pylj.util.System
The complete system information from pylj
"""
# %matplotlib notebook
system = md.initialise(number_of_particles, temperature,
box_length, 'square')
sample_system = sample.CellPlus(system,
'q/m$^{-1}$', 'I(q)')
system.time = 0
for i in range(0, number_of_steps):
system.integrate(md.velocity_verlet)
system.md_sample()
system.heat_bath(temperature)
system.time += system.timestep_length
system.step += 1
if system.step % sample_frequency == 0:
min_q = 2. * np.pi / box_length
qs = np.linspace(min_q, 10e10, 120)[20:]
inten = debye(qs, system.particles['xposition'],
system.particles['yposition'],
box_length)
sample_system.update(system, qs, inten)
return system
system = md_simulation(10, 3, 15, 5000, 10)
| run_local/21_temp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # MadMiner particle physics tutorial
#
# # Part 3a: Training a likelihood ratio estimator
#
# <NAME>, <NAME>, <NAME>, and <NAME> 2018-2019
# In part 3a of this tutorial we will finally train a neural network to estimate likelihood ratios. We assume that you have run part 1 and 2a of this tutorial. If, instead of 2a, you have run part 2b, you just have to load a different filename later.
# ## Preparations
# +
from __future__ import absolute_import, division, print_function, unicode_literals
import logging
import numpy as np
import matplotlib
from matplotlib import pyplot as plt
# %matplotlib inline
# +
# MadMiner output
logging.basicConfig(
format='%(asctime)-5.5s %(name)-20.20s %(levelname)-7.7s %(message)s',
datefmt='%H:%M',
level=logging.INFO
)
# Output of all other modules (e.g. matplotlib)
for key in logging.Logger.manager.loggerDict:
if "madminer" not in key:
logging.getLogger(key).setLevel(logging.WARNING)
# -
from madminer import SampleAugmenter, sampling, ParameterizedRatioEstimator
# ## 1. Make (unweighted) training and test samples with augmented data
# At this point, we have all the information we need from the simulations. But the data is not quite ready to be used for machine learning. The `madminer.sampling` class `SampleAugmenter` will take care of the remaining book-keeping steps before we can train our estimators:
#
# First, it unweights the samples, i.e. for a given parameter vector `theta` (or a distribution `p(theta)`) it picks events `x` such that their distribution follows `p(x|theta)`. The selected samples will all come from the event file we have so far, but their frequency is changed -- some events will appear multiple times, some will disappear.
#
# Second, `SampleAugmenter` calculates all the augmented data ("gold") that is the key to our new inference methods. Depending on the specific technique, these are the joint likelihood ratio and / or the joint score. It saves all these pieces of information for the selected events in a set of numpy files that can easily be used in any machine learning framework.
sampler = SampleAugmenter('data/lhe_data_shuffled.h5')
# sampler = SampleAugmenter('data/delphes_data_shuffled.h5')
# The `SampleAugmenter` class defines five different high-level functions to generate train or test samples:
# - `sample_train_plain()`, which only saves observations x, for instance for histograms or ABC;
# - `sample_train_local()` for methods like SALLY and SALLINO, which will be demonstrated in the second part of the tutorial;
# - `sample_train_density()` for neural density estimation techniques like MAF or SCANDAL;
# - `sample_train_ratio()` for techniques like CARL, ROLR, CASCAL, and RASCAL, when only theta0 is parameterized;
# - `sample_train_more_ratios()` for the same techniques, but with both theta0 and theta1 parameterized;
# - `sample_test()` for the evaluation of any method.
#
# For the arguments `theta`, `theta0`, or `theta1`, you can (and should!) use the helper functions `benchmark()`, `benchmarks()`, `morphing_point()`, `morphing_points()`, and `random_morphing_points()`, all defined in the `madminer.sampling` module.
#
# Here we'll train a likelihood ratio estimator with the ALICES method, so we focus on the `extract_samples_train_ratio()` function. We'll sample the numerator hypothesis in the likelihood ratio with 1000 points drawn from a Gaussian prior, and fix the denominator hypothesis to the SM.
#
# Note the keyword `sample_only_from_closest_benchmark=True`, which makes sure that for each parameter point we only use the events that were originally (in MG) generated from the closest benchmark. This reduces the statistical fluctuations in the outcome quite a bit.
x, theta0, theta1, y, r_xz, t_xz, n_effective = sampler.sample_train_ratio(
theta0=sampling.benchmarks(['sm', '50', 'neg_50', '200', 'neg_200', '500', 'neg_500']),
theta1=sampling.benchmark('sm'),
n_samples=500000,
folder='./data/samples',
filename='train_ratio',
sample_only_from_closest_benchmark=True,
return_individual_n_effective=True,
)
# For the evaluation we'll need a test sample:
_ = sampler.sample_test(
theta=sampling.benchmark('sm'),
n_samples=1000,
folder='./data/samples',
filename='test'
)
_,_,neff=sampler.sample_train_plain(
#theta=sampling.morphing_point([0,0.5]),
theta=sampling.benchmarks(['50', 'neg_50', '200', 'neg_200', '500', 'neg_500']),
n_samples=10000,
)
# You might notice the information about the "eeffective number of samples" in the output. This is defined as `1 / max_events(weights)`; the smaller it is, the bigger the statistical fluctuations from too large weights. Let's plot this over the parameter space:
# +
cmin, cmax = 10., 10000.
cut = (y.flatten()==0)
fig = plt.figure(figsize=(5,4))
sc = plt.scatter(theta0[cut][:,0], theta0[cut][:,1], c=n_effective[cut],
s=30., cmap='viridis',
norm=matplotlib.colors.LogNorm(vmin=cmin, vmax=cmax),
marker='o')
cb = plt.colorbar(sc)
cb.set_label('Effective number of samples')
plt.xlim(-1.0,1.0)
plt.ylim(-1.0,1.0)
plt.tight_layout()
plt.show()
# -
# ## 2. Plot cross section over parameter space
# This is not strictly necessary, but we can also plot the cross section as a function of parameter space:
# +
thetas_benchmarks, xsecs_benchmarks, xsec_errors_benchmarks = sampler.cross_sections(
theta=sampling.benchmarks(list(sampler.benchmarks.keys()))
)
thetas_morphing, xsecs_morphing, xsec_errors_morphing = sampler.cross_sections(
theta=sampling.random_morphing_points(1000, [('gaussian', 0., 1.), ('gaussian', 0., 1.)])
)
# +
cmin, cmax = 0., 2.5 * np.mean(xsecs_morphing)
fig = plt.figure(figsize=(5,4))
sc = plt.scatter(thetas_morphing[:,0], thetas_morphing[:,1], c=xsecs_morphing,
s=40., cmap='viridis', vmin=cmin, vmax=cmax,
marker='o')
plt.scatter(thetas_benchmarks[:,0], thetas_benchmarks[:,1], c=xsecs_benchmarks,
s=200., cmap='viridis', vmin=cmin, vmax=cmax, lw=2., edgecolor='black',
marker='s')
cb = plt.colorbar(sc)
cb.set_label('xsec [pb]')
plt.xlim(-3.,3.)
plt.ylim(-3.,3.)
plt.tight_layout()
plt.show()
# -
# What you see here is a morphing algorithm in action. We only asked MadGraph to calculate event weights (differential cross sections, or basically squared matrix elements) at six fixed parameter points (shown here as squares with black edges). But with our knowledge about the structure of the process we can interpolate any observable to any parameter point without loss (except that statistical uncertainties might increase)!
# ## 3. Train likelihood ratio estimator
# It's now time to build the neural network that estimates the likelihood ratio. The central object for this is the `madminer.ml.ParameterizedRatioEstimator` class. It defines functions that train, save, load, and evaluate the estimators.
#
# In the initialization, the keywords `n_hidden` and `activation` define the architecture of the (fully connected) neural network:
from madminer.ml.morphing_aware import MorphingAwareRatioEstimator
#estimator = ParameterizedRatioEstimator(
estimator = MorphingAwareRatioEstimator(
morphing_setup_filename='data/setup.h5',
n_hidden=(60,60),
activation="tanh",
)
# To train this model we will minimize the ALICES loss function described in ["Likelihood-free inference with an improved cross-entropy estimator"](https://arxiv.org/abs/1808.00973). Many alternatives, including RASCAL, are described in ["Constraining Effective Field Theories With Machine Learning"](https://arxiv.org/abs/1805.00013) and ["A Guide to Constraining Effective Field Theories With Machine Learning"](https://arxiv.org/abs/1805.00020). There is also SCANDAL introduced in ["Mining gold from implicit models to improve likelihood-free inference"](https://arxiv.org/abs/1805.12244).
# +
estimator.train(
method='carl',
theta='data/samples/theta0_train_ratio.npy',
x='data/samples/x_train_ratio.npy',
y='data/samples/y_train_ratio.npy',
r_xz='data/samples/r_xz_train_ratio.npy',
t_xz='data/samples/t_xz_train_ratio.npy',
alpha=10,
n_epochs=1000,
#scale_parameters=True,
)
estimator.save('models/carl')
# -
# ### 50 epochs Robustness Check
# +
estimator.train(
method='carl',
theta='data/samples/theta0_train_ratio.npy',
x='data/samples/x_train_ratio.npy',
y='data/samples/y_train_ratio.npy',
r_xz='data/samples/r_xz_train_ratio.npy',
t_xz='data/samples/t_xz_train_ratio.npy',
alpha=10,
n_epochs=50,
#scale_parameters=True,
)
estimator.save('models/carl50-1')
# +
#estimator = ParameterizedRatioEstimator(
estimator = MorphingAwareRatioEstimator(
morphing_setup_filename='data/setup.h5',
n_hidden=(60,60),
activation="tanh",
)
estimator.train(
method='carl',
theta='data/samples/theta0_train_ratio.npy',
x='data/samples/x_train_ratio.npy',
y='data/samples/y_train_ratio.npy',
r_xz='data/samples/r_xz_train_ratio.npy',
t_xz='data/samples/t_xz_train_ratio.npy',
alpha=10,
n_epochs=50,
#scale_parameters=True,
)
estimator.save('models/carl50-2')
# +
#estimator = ParameterizedRatioEstimator(
estimator = MorphingAwareRatioEstimator(
morphing_setup_filename='data/setup.h5',
n_hidden=(60,60),
activation="tanh",
)
estimator.train(
method='carl',
theta='data/samples/theta0_train_ratio.npy',
x='data/samples/x_train_ratio.npy',
y='data/samples/y_train_ratio.npy',
r_xz='data/samples/r_xz_train_ratio.npy',
t_xz='data/samples/t_xz_train_ratio.npy',
alpha=10,
n_epochs=50,
#scale_parameters=True,
)
estimator.save('models/carl50-3')
# -
# ### 10 epochs Reproduce
# +
#estimator = ParameterizedRatioEstimator(
estimator = MorphingAwareRatioEstimator(
morphing_setup_filename='data/setup.h5',
n_hidden=(60,60),
activation="tanh",
)
estimator.train(
method='carl',
theta='data/samples/theta0_train_ratio.npy',
x='data/samples/x_train_ratio.npy',
y='data/samples/y_train_ratio.npy',
r_xz='data/samples/r_xz_train_ratio.npy',
t_xz='data/samples/t_xz_train_ratio.npy',
alpha=10,
n_epochs=10,
#scale_parameters=True,
)
estimator.save('models/carl10')
# +
#estimator = ParameterizedRatioEstimator(
estimator = MorphingAwareRatioEstimator(
morphing_setup_filename='data/setup.h5',
n_hidden=(60,60),
activation="tanh",
)
estimator.train(
method='carl',
theta='data/samples/theta0_train_ratio.npy',
x='data/samples/x_train_ratio.npy',
y='data/samples/y_train_ratio.npy',
r_xz='data/samples/r_xz_train_ratio.npy',
t_xz='data/samples/t_xz_train_ratio.npy',
alpha=10,
n_epochs=10,
#scale_parameters=True,
)
estimator.save('models/carl10-1')
# +
#estimator = ParameterizedRatioEstimator(
estimator = MorphingAwareRatioEstimator(
morphing_setup_filename='data/setup.h5',
n_hidden=(60,60),
activation="tanh",
)
estimator.train(
method='carl',
theta='data/samples/theta0_train_ratio.npy',
x='data/samples/x_train_ratio.npy',
y='data/samples/y_train_ratio.npy',
r_xz='data/samples/r_xz_train_ratio.npy',
t_xz='data/samples/t_xz_train_ratio.npy',
alpha=10,
n_epochs=10,
#scale_parameters=True,
)
estimator.save('models/carl10-2')
# +
#estimator = ParameterizedRatioEstimator(
estimator = MorphingAwareRatioEstimator(
morphing_setup_filename='data/setup.h5',
n_hidden=(60,60),
activation="tanh",
)
estimator.train(
method='carl',
theta='data/samples/theta0_train_ratio.npy',
x='data/samples/x_train_ratio.npy',
y='data/samples/y_train_ratio.npy',
r_xz='data/samples/r_xz_train_ratio.npy',
t_xz='data/samples/t_xz_train_ratio.npy',
alpha=10,
n_epochs=10,
#scale_parameters=True,
)
estimator.save('models/carl10-3')
# -
# ### Bigger Batch size
# +
#estimator = ParameterizedRatioEstimator(
estimator = MorphingAwareRatioEstimator(
morphing_setup_filename='data/setup.h5',
n_hidden=(60,60),
activation="tanh",
)
estimator.train(
method='carl',
theta='data/samples/theta0_train_ratio.npy',
x='data/samples/x_train_ratio.npy',
y='data/samples/y_train_ratio.npy',
r_xz='data/samples/r_xz_train_ratio.npy',
t_xz='data/samples/t_xz_train_ratio.npy',
alpha=10,
n_epochs=10,
batch_size=int(6000000/4),
)
estimator.save('models/carl10-bigbatch')
# +
#estimator = ParameterizedRatioEstimator(
estimator = MorphingAwareRatioEstimator(
morphing_setup_filename='data/setup.h5',
n_hidden=(60,60),
activation="tanh",
)
estimator.train(
method='carl',
theta='data/samples/theta0_train_ratio.npy',
x='data/samples/x_train_ratio.npy',
y='data/samples/y_train_ratio.npy',
r_xz='data/samples/r_xz_train_ratio.npy',
t_xz='data/samples/t_xz_train_ratio.npy',
alpha=10,
n_epochs=50,
batch_size=int(6000000/4),
)
estimator.save('models/carl50-bigbatch')
# +
#estimator = ParameterizedRatioEstimator(
estimator = MorphingAwareRatioEstimator(
morphing_setup_filename='data/setup.h5',
n_hidden=(60,60),
activation="tanh",
)
estimator.train(
method='carl',
theta='data/samples/theta0_train_ratio.npy',
x='data/samples/x_train_ratio.npy',
y='data/samples/y_train_ratio.npy',
r_xz='data/samples/r_xz_train_ratio.npy',
t_xz='data/samples/t_xz_train_ratio.npy',
alpha=10,
n_epochs=1000,
batch_size=int(6000000/4),
)
estimator.save('models/carl1000-bigbatch')
# -
# Let's for fun also train a model that only used `pt_j1` as input observable, which can be specified using the option `features` when defining the `ParameterizedRatioEstimator`
# +
#estimator_pt = ParameterizedRatioEstimator(
estimator_pt = MorphingAwareRatioEstimator(
morphing_setup_filename='data/setup.h5',
n_hidden=(40,40),
activation="tanh",
features=[0],
)
estimator_pt.train(
method='carl',
theta='data/samples/theta0_train_ratio.npy',
x='data/samples/x_train_ratio.npy',
y='data/samples/y_train_ratio.npy',
r_xz='data/samples/r_xz_train_ratio.npy',
t_xz='data/samples/t_xz_train_ratio.npy',
alpha=8,
n_epochs=1000,
#scale_parameters=True,
)
estimator_pt.save('models/carl_pt')
# -
# ## 4. Evaluate likelihood ratio estimator
# `estimator.evaluate_log_likelihood_ratio(theta,x)` estimated the log likelihood ratio and the score for all combination between the given phase-space points `x` and parameters `theta`. That is, if given 100 events `x` and a grid of 25 `theta` points, it will return 25\*100 estimates for the log likelihood ratio and 25\*100 estimates for the score, both indexed by `[i_theta,i_x]`.
# +
theta_each = np.linspace(-1,1,25)
theta0, theta1 = np.meshgrid(theta_each, theta_each)
theta_grid = np.vstack((theta0.flatten(), theta1.flatten())).T
np.save('data/samples/theta_grid.npy', theta_grid)
theta_denom = np.array([[0.,0.]])
np.save('data/samples/theta_ref.npy', theta_denom)
# +
estimator.load('models/carl')
log_r_hat, _ = estimator.evaluate_log_likelihood_ratio(
theta='data/samples/theta_grid.npy',
x='data/samples/x_test.npy',
evaluate_score=False
)
# -
# Let's look at the result:
# +
bin_size = theta_each[1] - theta_each[0]
edges = np.linspace(theta_each[0] - bin_size/2, theta_each[-1] + bin_size/2, len(theta_each)+1)
fig = plt.figure(figsize=(6,5))
ax = plt.gca()
expected_llr = np.mean(log_r_hat,axis=1)
best_fit = theta_grid[np.argmin(-2.*expected_llr)]
cmin, cmax = np.min(-2*expected_llr), np.max(-2*expected_llr)
pcm = ax.pcolormesh(edges, edges, -2. * expected_llr.reshape((25,25)),
norm=matplotlib.colors.Normalize(vmin=cmin, vmax=cmax),
cmap='viridis_r')
cbar = fig.colorbar(pcm, ax=ax, extend='both')
plt.scatter(best_fit[0], best_fit[1], s=80., color='black', marker='*')
plt.xlabel(r'$\theta_0$')
plt.ylabel(r'$\theta_1$')
cbar.set_label(r'$\mathbb{E}_x [ -2\, \log \,\hat{r}(x | \theta, \theta_{SM}) ]$')
plt.tight_layout()
plt.show()
# -
# Note that in this tutorial our sample size was very small, and the network might not really have a chance to converge to the correct likelihood ratio function. So don't worry if you find a minimum that is not at the right point (the SM, i.e. the origin in this plot). Feel free to dial up the event numbers in the run card as well as the training samples and see what happens then!
log_r_hat.shape
np.load('data/samples/x_train_ratio.npy').shape
theta = np.load('data/samples/theta0_train_ratio.npy')
theta.shape
| examples/tutorial_particle_physics/3a_likelihood_ratio.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# **Chapter 11 – Deep Learning**
# + [markdown] deletable=true editable=true
# _This notebook contains all the sample code and solutions to the exercices in chapter 11._
# + [markdown] deletable=true editable=true
# # Setup
# + [markdown] deletable=true editable=true
# First, let's make sure this notebook works well in both python 2 and 3, import a few common modules, ensure MatplotLib plots figures inline and prepare a function to save the figures:
# + deletable=true editable=true
# To support both python 2 and python 3
from __future__ import division, print_function, unicode_literals
# Common imports
import numpy as np
import numpy.random as rnd
import os
# to make this notebook's output stable across runs
rnd.seed(42)
# To plot pretty figures
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
plt.rcParams['axes.labelsize'] = 14
plt.rcParams['xtick.labelsize'] = 12
plt.rcParams['ytick.labelsize'] = 12
# Where to save the figures
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "deep"
def save_fig(fig_id, tight_layout=True):
path = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID, fig_id + ".png")
print("Saving figure", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format='png', dpi=300)
# + [markdown] deletable=true editable=true
# # Activation functions
# + deletable=true editable=true
def logit(z):
return 1 / (1 + np.exp(-z))
# + deletable=true editable=true
z = np.linspace(-5, 5, 200)
plt.plot([-5, 5], [0, 0], 'k-')
plt.plot([-5, 5], [1, 1], 'k--')
plt.plot([0, 0], [-0.2, 1.2], 'k-')
plt.plot([-5, 5], [-3/4, 7/4], 'g--')
plt.plot(z, logit(z), "b-", linewidth=2)
props = dict(facecolor='black', shrink=0.1)
plt.annotate('Saturating', xytext=(3.5, 0.7), xy=(5, 1), arrowprops=props, fontsize=14, ha="center")
plt.annotate('Saturating', xytext=(-3.5, 0.3), xy=(-5, 0), arrowprops=props, fontsize=14, ha="center")
plt.annotate('Linear', xytext=(2, 0.2), xy=(0, 0.5), arrowprops=props, fontsize=14, ha="center")
plt.grid(True)
plt.title("Sigmoid activation function", fontsize=14)
plt.axis([-5, 5, -0.2, 1.2])
save_fig("sigmoid_saturation_plot")
plt.show()
# + deletable=true editable=true
def leaky_relu(z, alpha=0.01):
return np.maximum(alpha*z, z)
# + deletable=true editable=true
plt.plot(z, leaky_relu(z, 0.05), "b-", linewidth=2)
plt.plot([-5, 5], [0, 0], 'k-')
plt.plot([0, 0], [-0.5, 4.2], 'k-')
plt.grid(True)
props = dict(facecolor='black', shrink=0.1)
plt.annotate('Leak', xytext=(-3.5, 0.5), xy=(-5, -0.2), arrowprops=props, fontsize=14, ha="center")
plt.title("Leaky ReLU activation function", fontsize=14)
plt.axis([-5, 5, -0.5, 4.2])
save_fig("leaky_relu_plot")
plt.show()
# + deletable=true editable=true
def elu(z, alpha=1):
return np.where(z<0, alpha*(np.exp(z)-1), z)
# + deletable=true editable=true
plt.plot(z, elu(z), "b-", linewidth=2)
plt.plot([-5, 5], [0, 0], 'k-')
plt.plot([-5, 5], [-1, -1], 'k--')
plt.plot([0, 0], [-2.2, 3.2], 'k-')
plt.grid(True)
props = dict(facecolor='black', shrink=0.1)
plt.title(r"ELU activation function ($\alpha=1$)", fontsize=14)
plt.axis([-5, 5, -2.2, 3.2])
save_fig("elu_plot")
plt.show()
# + deletable=true editable=true
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("/tmp/data/")
# + deletable=true editable=true
def leaky_relu(z, name=None):
return tf.maximum(0.01 * z, z, name=name)
# + deletable=true editable=true
import tensorflow as tf
# + deletable=true editable=true
from IPython.display import clear_output, Image, display, HTML
def strip_consts(graph_def, max_const_size=32):
"""Strip large constant values from graph_def."""
strip_def = tf.GraphDef()
for n0 in graph_def.node:
n = strip_def.node.add()
n.MergeFrom(n0)
if n.op == 'Const':
tensor = n.attr['value'].tensor
size = len(tensor.tensor_content)
if size > max_const_size:
tensor.tensor_content = b"<stripped %d bytes>"%size
return strip_def
def show_graph(graph_def, max_const_size=32):
"""Visualize TensorFlow graph."""
if hasattr(graph_def, 'as_graph_def'):
graph_def = graph_def.as_graph_def()
strip_def = strip_consts(graph_def, max_const_size=max_const_size)
code = """
<script>
function load() {{
document.getElementById("{id}").pbtxt = {data};
}}
</script>
<link rel="import" href="https://tensorboard.appspot.com/tf-graph-basic.build.html" onload=load()>
<div style="height:600px">
<tf-graph-basic id="{id}"></tf-graph-basic>
</div>
""".format(data=repr(str(strip_def)), id='graph'+str(np.random.rand()))
iframe = """
<iframe seamless style="width:1200px;height:620px;border:0" srcdoc="{}"></iframe>
""".format(code.replace('"', '"'))
display(HTML(iframe))
# + [markdown] deletable=true editable=true
# Note: the book uses `tensorflow.contrib.layers.fully_connected()` rather than `tf.layers.dense()` (which did not exist when this chapter was written). It is now preferable to use `tf.layers.dense()`, because anything in the contrib module may change or be deleted without notice. The `dense()` function is almost identical to the `fully_connected()` function. The main differences relevant to this chapter are:
# * several parameters are renamed: `scope` becomes `name`, `activation_fn` becomes `activation` (and similarly the `_fn` suffix is removed from other parameters such as `normalizer_fn`), `weights_initializer` becomes `kernel_initializer`, etc.
# * the default `activation` is now `None` rather than `tf.nn.relu`.
# * it does not support `tensorflow.contrib.framework.arg_scope()` (introduced later in chapter 11).
# * it does not support regularizer params (introduced later in chapter 11).
# + deletable=true editable=true
tf.reset_default_graph()
n_inputs = 28*28 # MNIST
n_hidden1 = 300
n_hidden2 = 100
n_outputs = 10
learning_rate = 0.01
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int64, shape=(None), name="y")
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=leaky_relu, name="hidden1")
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=leaky_relu, name="hidden2")
logits = tf.layers.dense(hidden2, n_outputs, name="outputs")
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("train"):
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32))
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# + deletable=true editable=true
n_epochs = 20
batch_size = 100
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for iteration in range(len(mnist.test.labels)//batch_size):
X_batch, y_batch = mnist.train.next_batch(batch_size)
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
acc_train = accuracy.eval(feed_dict={X: X_batch, y: y_batch})
acc_test = accuracy.eval(feed_dict={X: mnist.test.images, y: mnist.test.labels})
print(epoch, "Train accuracy:", acc_train, "Test accuracy:", acc_test)
save_path = saver.save(sess, "my_model_final.ckpt")
# + [markdown] deletable=true editable=true
# # Batch Normalization
# + [markdown] deletable=true editable=true
# Note: the book uses `tensorflow.contrib.layers.batch_norm()` rather than `tf.layers.batch_normalization()` (which did not exist when this chapter was written). It is now preferable to use `tf.layers.batch_normalization()`, because anything in the contrib module may change or be deleted without notice. Instead of using the `batch_norm()` function as a regularizer parameter to the `fully_connected()` function, we now use `batch_normalization()` and we explicitly create a distinct layer. The parameters are a bit different, in particular:
# * `decay` is renamed to `momentum`,
# * `is_training` is renamed to `training`,
# * `updates_collections` is removed: the update operations needed by batch normalization are added to the `UPDATE_OPS` collection and you need to explicity run these operations during training (see the execution phase below),
# * we don't need to specify `scale=True`, as that is the default.
#
# Also note that in order to run batch norm just _before_ each hidden layer's activation function, we apply the ELU activation function manually, right after the batch norm layer.
#
# Note: since the `tf.layers.dense()` function is incompatible with `tf.contrib.layers.arg_scope()` (which is used in the book), we now use python's `functools.partial()` function instead. It makes it easy to create a `my_dense_layer()` function that just calls `tf.layers.dense()` with the desired parameters automatically set (unless they are overridden when calling `my_dense_layer()`). As you can see, the code remains very similar.
# + deletable=true editable=true
tf.reset_default_graph()
from functools import partial
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300
n_hidden2 = 100
n_outputs = 10
learning_rate = 0.01
momentum = 0.25
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int64, shape=(None), name="y")
is_training = tf.placeholder(tf.bool, shape=(), name='is_training')
with tf.name_scope("dnn"):
he_init = tf.contrib.layers.variance_scaling_initializer()
my_batch_norm_layer = partial(
tf.layers.batch_normalization,
training=is_training,
momentum=0.9)
my_dense_layer = partial(
tf.layers.dense,
kernel_initializer=he_init)
hidden1 = my_dense_layer(X, n_hidden1, name="hidden1")
bn1 = tf.nn.elu(my_batch_norm_layer(hidden1))
hidden2 = my_dense_layer(bn1, n_hidden2, name="hidden2")
bn2 = tf.nn.elu(my_batch_norm_layer(hidden2))
logits_before_bn = my_dense_layer(bn2, n_outputs, activation=None, name="outputs")
logits = my_batch_norm_layer(logits_before_bn)
extra_update_ops = tf.get_collection(tf.GraphKeys.UPDATE_OPS)
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("train"):
optimizer = tf.train.MomentumOptimizer(learning_rate, momentum)
training_op = optimizer.minimize(loss)
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32))
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# + [markdown] deletable=true editable=true
# Note: since we are using `tf.layers.batch_normalization()` rather than `tf.contrib.layers.batch_norm()` (as in the book), we need to explicitly run the extra update operations needed by batch normalization (`sess.run([training_op, extra_update_ops],...`).
# + deletable=true editable=true
n_epochs = 20
batch_size = 200
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for iteration in range(len(mnist.test.labels)//batch_size):
X_batch, y_batch = mnist.train.next_batch(batch_size)
sess.run([training_op, extra_update_ops], feed_dict={is_training: True, X: X_batch, y: y_batch})
acc_train = accuracy.eval(feed_dict={is_training: False, X: X_batch, y: y_batch})
acc_test = accuracy.eval(feed_dict={is_training: False, X: mnist.test.images, y: mnist.test.labels})
print(epoch, "Train accuracy:", acc_train, "Test accuracy:", acc_test)
save_path = saver.save(sess, "my_model_final.ckpt")
# + [markdown] deletable=true editable=true
# Now the same model with $\ell_1$ regularization:
# + deletable=true editable=true
tf.reset_default_graph()
from functools import partial
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int64, shape=(None), name="y")
is_training = tf.placeholder(tf.bool, shape=(), name='is_training')
with tf.name_scope("dnn"):
he_init = tf.contrib.layers.variance_scaling_initializer()
my_batch_norm_layer = partial(
tf.layers.batch_normalization,
training=is_training,
momentum=0.9)
my_dense_layer = partial(
tf.layers.dense,
kernel_initializer=he_init,
kernel_regularizer=tf.contrib.layers.l1_regularizer(0.01))
hidden1 = my_dense_layer(X, n_hidden1, name="hidden1")
bn1 = tf.nn.elu(my_batch_norm_layer(hidden1))
hidden2 = my_dense_layer(bn1, n_hidden2, name="hidden2")
bn2 = tf.nn.elu(my_batch_norm_layer(hidden2))
logits_before_bn = my_dense_layer(bn2, n_outputs, activation=None, name="outputs")
logits = my_batch_norm_layer(logits_before_bn)
extra_update_ops = tf.get_collection(tf.GraphKeys.UPDATE_OPS)
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
reg_losses = tf.get_collection(tf.GraphKeys.REGULARIZATION_LOSSES)
base_loss = tf.reduce_mean(xentropy, name="base_loss")
loss = tf.add_n([base_loss] + reg_losses, name="loss")
with tf.name_scope("train"):
optimizer = tf.train.MomentumOptimizer(learning_rate, momentum)
training_op = optimizer.minimize(loss)
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32))
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# + deletable=true editable=true
n_epochs = 20
batch_size = 200
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for iteration in range(len(mnist.test.labels)//batch_size):
X_batch, y_batch = mnist.train.next_batch(batch_size)
sess.run([training_op, extra_update_ops], feed_dict={is_training: True, X: X_batch, y: y_batch})
acc_train = accuracy.eval(feed_dict={is_training: False, X: X_batch, y: y_batch})
acc_test = accuracy.eval(feed_dict={is_training: False, X: mnist.test.images, y: mnist.test.labels})
print(epoch, "Train accuracy:", acc_train, "Test accuracy:", acc_test)
save_path = saver.save(sess, "my_model_final.ckpt")
# + deletable=true editable=true
[v.name for v in tf.global_variables()]
# + [markdown] deletable=true editable=true
# Note: the weights variable created by the `tf.layers.dense()` function is called `"kernel"` (instead of `"weights"` when using the `tf.contrib.layers.fully_connected()`, as in the book):
# + deletable=true editable=true
with tf.variable_scope("", default_name="", reuse=True): # root scope
weights1 = tf.get_variable("hidden1/kernel")
weights2 = tf.get_variable("hidden2/kernel")
# + deletable=true editable=true
tf.reset_default_graph()
x = tf.constant([0., 0., 3., 4., 30., 40., 300., 400.], shape=(4, 2))
c = tf.clip_by_norm(x, clip_norm=10)
c0 = tf.clip_by_norm(x, clip_norm=350, axes=0)
c1 = tf.clip_by_norm(x, clip_norm=10, axes=1)
with tf.Session() as sess:
xv = x.eval()
cv = c.eval()
c0v = c0.eval()
c1v = c1.eval()
print(xv)
# + deletable=true editable=true
print(cv)
# + deletable=true editable=true
print(np.linalg.norm(cv))
# + deletable=true editable=true
print(c0v)
# + deletable=true editable=true
print(np.linalg.norm(c0v, axis=0))
# + deletable=true editable=true
print(c1v)
# + deletable=true editable=true
print(np.linalg.norm(c1v, axis=1))
# + deletable=true editable=true
tf.reset_default_graph()
from functools import partial
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int64, shape=(None), name="y")
is_training = tf.placeholder(tf.bool, shape=(), name='is_training')
def max_norm_regularizer(threshold, axes=1, name="max_norm", collection="max_norm"):
def max_norm(weights):
clip_weights = tf.assign(weights, tf.clip_by_norm(weights, clip_norm=threshold, axes=axes), name=name)
tf.add_to_collection(collection, clip_weights)
return None # there is no regularization loss term
return max_norm
with tf.name_scope("dnn"):
my_dense_layer = partial(
tf.layers.dense,
activation=tf.nn.relu,
kernel_regularizer=max_norm_regularizer(1.5))
hidden1 = my_dense_layer(X, n_hidden1, name="hidden1")
hidden2 = my_dense_layer(hidden1, n_hidden2, name="hidden2")
logits = my_dense_layer(hidden2, n_outputs, activation=None, name="outputs")
clip_all_weights = tf.get_collection("max_norm")
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("train"):
optimizer = tf.train.MomentumOptimizer(learning_rate, momentum)
threshold = 1.0
grads_and_vars = optimizer.compute_gradients(loss)
capped_gvs = [(tf.clip_by_value(grad, -threshold, threshold), var)
for grad, var in grads_and_vars]
training_op = optimizer.apply_gradients(capped_gvs)
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32))
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# + deletable=true editable=true
n_epochs = 20
batch_size = 50
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for iteration in range(len(mnist.test.labels)//batch_size):
X_batch, y_batch = mnist.train.next_batch(batch_size)
sess.run(training_op, feed_dict={is_training: True, X: X_batch, y: y_batch})
sess.run(clip_all_weights)
acc_train = accuracy.eval(feed_dict={is_training: False, X: X_batch, y: y_batch})
acc_test = accuracy.eval(feed_dict={is_training: False, X: mnist.test.images, y: mnist.test.labels})
print(epoch, "Train accuracy:", acc_train, "Test accuracy:", acc_test)
save_path = saver.save(sess, "my_model_final.ckpt")
# + deletable=true editable=true
show_graph(tf.get_default_graph())
# + [markdown] deletable=true editable=true
# Note: the book uses `tf.contrib.layers.dropout()` rather than `tf.layers.dropout()` (which did not exist when this chapter was written). It is now preferable to use `tf.layers.dropout()`, because anything in the contrib module may change or be deleted without notice. The `tf.layers.dropout()` function is almost identical to the `tf.contrib.layers.dropout()` function, except for a few minor differences. Most importantly:
# * you must specify the dropout rate (`rate`) rather than the keep probability (`keep_prob`), where `rate` is simply equal to `1 - keep_prob`,
# * the `is_training` parameter is renamed to `training`.
# + deletable=true editable=true
from functools import partial
tf.reset_default_graph()
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int64, shape=(None), name="y")
is_training = tf.placeholder(tf.bool, shape=(), name='is_training')
initial_learning_rate = 0.1
decay_steps = 10000
decay_rate = 1/10
global_step = tf.Variable(0, trainable=False)
learning_rate = tf.train.exponential_decay(initial_learning_rate, global_step,
decay_steps, decay_rate)
dropout_rate = 0.5
with tf.name_scope("dnn"):
he_init = tf.contrib.layers.variance_scaling_initializer()
my_dense_layer = partial(
tf.layers.dense,
activation=tf.nn.elu,
kernel_initializer=he_init)
X_drop = tf.layers.dropout(X, dropout_rate, training=is_training)
hidden1 = my_dense_layer(X_drop, n_hidden1, name="hidden1")
hidden1_drop = tf.layers.dropout(hidden1, dropout_rate, training=is_training)
hidden2 = my_dense_layer(hidden1_drop, n_hidden2, name="hidden2")
hidden2_drop = tf.layers.dropout(hidden2, dropout_rate, training=is_training)
logits = my_dense_layer(hidden2_drop, n_outputs, activation=None, name="outputs")
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("train"):
optimizer = tf.train.MomentumOptimizer(learning_rate, momentum)
training_op = optimizer.minimize(loss, global_step=global_step)
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32))
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# + deletable=true editable=true
n_epochs = 20
batch_size = 50
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for iteration in range(len(mnist.test.labels)//batch_size):
X_batch, y_batch = mnist.train.next_batch(batch_size)
sess.run(training_op, feed_dict={is_training: True, X: X_batch, y: y_batch})
acc_train = accuracy.eval(feed_dict={is_training: False, X: X_batch, y: y_batch})
acc_test = accuracy.eval(feed_dict={is_training: False, X: mnist.test.images, y: mnist.test.labels})
print(epoch, "Train accuracy:", acc_train, "Test accuracy:", acc_test)
save_path = saver.save(sess, "my_model_final.ckpt")
# + deletable=true editable=true
train_vars = tf.get_collection(tf.GraphKeys.TRAINABLE_VARIABLES,
scope="hidden[2]|outputs")
# + deletable=true editable=true
training_op2 = optimizer.minimize(loss, var_list=train_vars)
# + deletable=true editable=true
for i in tf.global_variables():
print(i.name)
# + deletable=true editable=true
for i in tf.get_collection(tf.GraphKeys.TRAINABLE_VARIABLES):
print(i.name)
# + deletable=true editable=true
for i in train_vars:
print(i.name)
# + [markdown] deletable=true editable=true
# # Exercise solutions
# + [markdown] deletable=true editable=true
# **Coming soon**
# + deletable=true editable=true
| 11_deep_learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import tensorflow as tf
import os
root_dir = '/Users/abhishekverma/Desktop/python-env/TrafficCongestion/trafficdb/'
file = os.path.join(root_dir,'info.txt')
column_names=['filename', 'date', 'timestamp', 'direction', 'day/night', 'weather', 'start_frame','num_frames', 'class', 'notes']
df = pd.read_csv(file, sep='\\s+', names=column_names, skiprows=[0,])
df.head()
df['direction'].unique()
df['day/night'].unique()
df['num_frames'].unique()
df['class'].unique()
df['direction'].unique()
| day1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="IhsOv_14X1x1" outputId="59d45d53-fe54-42a4-d46d-db4e56e4155d"
from google.colab import drive
drive.mount('/content/drive')
# + id="MYZWrmrhYQHT"
import plotly
import plotly.express as px
import pandas as pd
import numpy as np
import os
import re
# + colab={"base_uri": "https://localhost:8080/"} id="SiKoqDFnsYub" outputId="a56992b2-1807-4b1c-f4c5-4205156444f0"
# %cd /content/drive/MyDrive/Colab Notebooks/data-analysis
# + [markdown] id="lgZR5oz7qp1C"
# ## Dataset preprocessing
# + [markdown] id="iMuImfDDwtRz"
# ### Kaggle
# + colab={"base_uri": "https://localhost:8080/", "height": 678} id="t7ICpEV_xCda" outputId="10b095a2-181e-4ed8-8e0f-d6f0b0281515"
kaggle_path = get_csv_from_gdrive(KAGGLE_URL)
kaggle_multiple_choice = pd.read_csv("data/multiple_choice_responses.csv", low_memory=False)
kaggle_multiple_choice.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 661} id="yeQ2OgTixffL" outputId="010336e4-7670-4662-be4f-444cf7e7ce9c"
kaggle = kaggle_multiple_choice.iloc[1:,:]
kaggle.head()
# + [markdown] id="Oo3vpbNPxSd4"
# Separate questions from answers
# + colab={"base_uri": "https://localhost:8080/"} id="RhMvefd1xVB0" outputId="28d1843e-c11a-413e-84fa-f8b8d7d38741"
kaggle_questions = kaggle_multiple_choice.iloc[0,:]
kaggle_questions.head()
# + [markdown] id="3ZyRgzC6zchO"
# #### Data cleaning
# + [markdown] id="GpwM33V3xfKg"
# Remove surveys that take less than 3 minutes or more than 600 minutes (These surveys are not so valuable)
# + id="ol6HLxGCy7TN"
less_3_minutes = kaggle[round(kaggle.iloc[:,0].astype(int) / 60) <= 3].index
kaggle = kaggle.drop(less_3_minutes, axis=0)
more_600_minutes = kaggle[round(kaggle.iloc[:,0].astype(int) / 60) >= 600].index
kaggle = kaggle.drop(more_600_minutes, axis=0)
# + [markdown] id="HAW1vgsIzKYL"
# Remove respondents who are not employed or project/product managers
# + id="U2IES-bazNG-"
students_and_others = kaggle[(kaggle.Q5 == 'Student') | (kaggle.Q5 == 'Other') | (kaggle.Q5 == 'Not employed') | (kaggle.Q5 == 'Product/Project Manager')].index
kaggle = kaggle.drop(list(students_and_others), axis=0)
# + [markdown] id="1tOJ6vdJzRue"
# Remove those who didn't disclose compensation (Q10 is `NaN`)
# + id="40Sij-SmzW__"
kaggle.dropna(subset=['Q10'], inplace=True)
# + [markdown] id="isdFRIPOzYod"
# #### Grouping
# + colab={"base_uri": "https://localhost:8080/"} id="Rx_l4aGjziaj" outputId="1e34c445-916f-410a-e0ab-2e10c9270cc2"
kaggle.Q5.value_counts()
# + [markdown] id="gkbIP0NLzj5L"
# Group DBA and Data engineer together as one job title
# + colab={"base_uri": "https://localhost:8080/"} id="uNM6PFsEzpMR" outputId="e41aede6-7a86-4703-b4b0-0283b581d8eb"
kaggle.Q5 = kaggle.Q5.replace('DBA/Database Engineer', 'Data Engineer/DBA')
kaggle.Q5 = kaggle.Q5.replace('Data Engineer', 'Data Engineer/DBA')
kaggle.Q5.value_counts()
# + [markdown] id="6rUiHmFyzqaB"
# Groupping statistician and Research Scientist:
# + colab={"base_uri": "https://localhost:8080/"} id="D3diXCbHzxHt" outputId="c086faf1-6fb9-4829-915b-1d6a9b0c9ca2"
kaggle.Q5 = kaggle.Q5.replace('Statistician', 'Statistician/Research Scientist')
kaggle.Q5 = kaggle.Q5.replace('Research Scientist', 'Statistician/Research Scientist')
kaggle.Q5.value_counts()
# + [markdown] id="iwkPBbs4zyRv"
# #### Process country names
# + id="aMxIJlWE0TQv"
kaggle.Q3 = kaggle.Q3.replace('United Kingdom of Great Britain and Northern Ireland', 'United Kingdom')
kaggle.Q3 = kaggle.Q3.replace('United States of America', 'United States')
# + [markdown] id="Rf1tSQjv0UXD"
# #### Process programming languages (Q18)
# + colab={"base_uri": "https://localhost:8080/"} id="_47Y_QZv0aJx" outputId="7bbb8bdf-8513-4773-ce20-08b554abf77f"
kaggle['ProgLang'] = kaggle[['Q18_Part_1', 'Q18_Part_2', 'Q18_Part_3', 'Q18_Part_4', 'Q18_Part_5',
'Q18_Part_6', 'Q18_Part_7', 'Q18_Part_8', 'Q18_Part_9', 'Q18_Part_10',
'Q18_Part_11', 'Q18_Part_12']].values.tolist()
kaggle.ProgLang.head()
# + colab={"base_uri": "https://localhost:8080/"} id="-YuBxp470ahO" outputId="96667e65-84f6-4264-bc0c-75cf4a47d160"
# remove nulls
kaggle.ProgLang = kaggle.ProgLang.apply(lambda x: [item for item in x if not pd.isnull(item)])
kaggle.ProgLang.head()
# + [markdown] id="TeZH4h6f0eHS"
# Calculates the quantity of different Programming Languages
# + colab={"base_uri": "https://localhost:8080/"} id="4dTB6ThZ0iZI" outputId="5b273a70-d138-432a-f47d-ebdf5b36851e"
kaggle['QtyProgLang'] = kaggle.ProgLang.apply(lambda x: len(x))
# If Quantity > 6 then it will be 6
kaggle.loc[kaggle.QtyProgLang > 6, 'QtyProgLang'] = 6
kaggle.QtyProgLang.head()
# + [markdown] id="Rnjd_grG0jrv"
# #### Process cloud platforms (Q29)
# + id="SY9KyLqm0sGn"
kaggle['CloudPlatf']= kaggle[['Q29_Part_1', 'Q29_Part_2', 'Q29_Part_3', 'Q29_Part_4',
'Q29_Part_5', 'Q29_Part_6', 'Q29_Part_7', 'Q29_Part_8',
'Q29_Part_9', 'Q29_Part_10', 'Q29_Part_11', 'Q29_Part_12']].values.tolist()
# remove nulls
kaggle.CloudPlatf = kaggle.CloudPlatf.apply(lambda x: [item.strip().lower() for item in x if not pd.isnull(item)])
# Calculates the quantity
kaggle['QtyCloudPlatf'] = kaggle.CloudPlatf.apply(lambda x: len(x))
kaggle.loc[kaggle.QtyCloudPlatf > 6, 'QtyCloudPlatf'] = 6
# + [markdown] id="MN0U1qBb0sf2"
# #### Process databases (Q34)
# + id="znT3BRSp0vWc"
kaggle['Databases']= kaggle[['Q34_Part_1', 'Q34_Part_2', 'Q34_Part_3', 'Q34_Part_4', 'Q34_Part_5', 'Q34_Part_6',
'Q34_Part_7', 'Q34_Part_8', 'Q34_Part_9', 'Q34_Part_10', 'Q34_Part_11', 'Q34_Part_12']].values.tolist()
kaggle.Databases = kaggle.Databases.apply(lambda x: [item.strip().lower() for item in x if not pd.isnull(item)])
kaggle['QtyDatabases'] = kaggle.Databases.apply(lambda x: len(x))
kaggle.loc[kaggle.QtyDatabases > 6, 'QtyDatabases'] = 6
# + [markdown] id="HLsJxieJ0xUO"
# #### Rename some columns
# + id="baiXh3iI0zZ1"
kaggle.columns = kaggle.columns.str.replace('Q15', 'TimeWritingCode')
kaggle.columns = kaggle.columns.str.replace('Q10', 'Salary')
kaggle.columns = kaggle.columns.str.replace('Q1', 'Age')
kaggle.columns = kaggle.columns.str.replace('Q5', 'JobTitle')
kaggle.columns = kaggle.columns.str.replace('Q3', 'Country')
kaggle.columns = kaggle.columns.str.replace('Q6', 'CompanySize')
# + [markdown] id="cE_hicOn01YN"
# #### Data transformation
#
# Transform some columns into categories
# + id="JXgPABa20_9l"
# Transform TimeWritingCode column into category
time_writting_code = ['I have never written code', '< 1 years', '1-2 years', '3-5 years',
'5-10 years', '10-20 years', '20+ years']
cat_dtype = pd.api.types.CategoricalDtype(categories=time_writting_code, ordered=True)
kaggle.TimeWritingCode = kaggle.TimeWritingCode.astype(cat_dtype)
# + id="1GKOJwhA1C_h"
# Transform CompanySize into category
company_size = ['0-49 employees', '50-249 employees', '250-999 employees', '1000-9,999 employees', '> 10,000 employees']
cat_dtype = pd.api.types.CategoricalDtype(categories=company_size, ordered=True)
kaggle.CompanySize = kaggle.CompanySize.astype(cat_dtype)
# + id="MOPSVm4E1LcV"
# Transform JobTitle into category
job_titles = ['Business Analyst', 'Data Analyst', 'Data Scientist',
'Data Engineer/DBA', 'Software Engineer', 'Statistician/Research Scientist']
cat_dtype = pd.api.types.CategoricalDtype(categories=job_titles, ordered=True)
kaggle.JobTitle = kaggle.JobTitle.astype(cat_dtype)
# + [markdown] id="IXNdpK0G1ZqM"
# Add count column to make groupby easier
# + id="XwhDUtj31Vdn"
kaggle['Count'] = 1
# + [markdown] id="VTgRwCa41ZKI"
# Transform range of salaries into numerical value
# + id="otvcqYxa1fWP"
compensation = kaggle.Salary.str.replace(r'(?:(?!\d|\-).)*', '').str.replace('500000', '500-500000').str.split('-')
kaggle.Salary = compensation.apply(lambda x: (int(x[0]) * 1000 + int(x[1]))/ 2) / 1000 # calculated in thousand dollars
# + [markdown] id="okVeCdRh1i5z"
# Transform range of ages into numerical value
# + id="YEtct3Ds1nWm"
age = kaggle.Age.str.replace(r'(?:(?!\d|\-).)*', '').str.replace('70', '70-80').str.split('-')
kaggle.Age = (age.apply(lambda x: (int(x[0]) + int(x[1]))/ 2)).astype(int)
# + [markdown] id="JFZMDMrV1pE4"
# #### Filter only the columns we're interested in
# + colab={"base_uri": "https://localhost:8080/", "height": 580} id="A09ZcWdZ1vXJ" outputId="e8ae5ab5-778f-459a-e822-bc36361c9d3f"
kaggle = kaggle[['Age', 'Country', 'JobTitle', 'CompanySize', 'Salary', 'TimeWritingCode', 'ProgLang',
'QtyProgLang', 'CloudPlatf', 'QtyCloudPlatf', 'Databases', 'QtyDatabases', 'Count']]
kaggle.head(10)
# + id="KzhoKuDE1xW4"
kaggle.to_csv("data/multiple_choice_responses_preprocessed.csv")
# + colab={"base_uri": "https://localhost:8080/", "height": 357} id="UJcDZr2yjYqW" outputId="65f7e38c-a5d7-4056-da72-2bcbb5ca03c9"
pd.read_csv("https://gist.githubusercontent.com/EckoTan0804/7ba61515d185c6558f77504044b485bb/raw/4caac4c296138e0d40aa22c90ae38d712ba0531d/multiple_choice_responses_preprocessed.csv").head()
# + [markdown] id="-ni0Tp_UwfR0"
# ### Glassdoor
# + colab={"base_uri": "https://localhost:8080/", "height": 355} id="pJjwgSRplgcI" outputId="01e68e34-3f84-4569-a27f-efcaa2dd598d"
glassdoor_full = pd.read_csv('data/glassdoor.csv')
glassdoor_full.head()
# + [markdown] id="FypMtBW_9kYX"
# #### Select and rename columns
# + [markdown] id="HZwMsGt4tHLY"
# We select only the columns we're interested in: `header.jotTitle`, `job.description`, `map.country`
# + id="ddSmkNFhvm3M"
COLUMNS = ["header.jobTitle", "job.description", "map.country"]
glassdoor = glassdoor_full[COLUMNS].copy()
# + [markdown] id="YOCxowI_v6x2"
# Rename selected columns to meaningful names:
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="Ff1LqT1CwARF" outputId="253e3887-ab42-4372-a49d-66f465ef0daa"
glassdoor.columns = ["JobTitle", "JobDescription", "Country"]
glassdoor.head()
# + [markdown] id="yoqVgj0v9nz_"
# #### Processing `Country` column
# + [markdown] id="wIgDUeviwJYT"
# Drop `NaN` countries:
# + id="8DtrNuaYwPvh"
glassdoor.dropna(subset=["Country"], inplace=True)
# + [markdown] id="APV45QYWwVRz"
# The value in `Country` column is not consistent: some country names are written in full, while others are in 2 digit codes.
# + colab={"base_uri": "https://localhost:8080/"} id="ayJieWBj9wEv" outputId="aaea5f78-9ee8-4c21-ac2b-e2388e61b598"
glassdoor["Country"]
# + [markdown] id="17PUtNtt9y3X"
# We fix this problem using `country_names_2_digit_codes.csv`
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="dgkE3Ep8-Ezv" outputId="02831831-75a8-4811-c201-832218e7887a"
country_codes = pd.read_csv("data/country_names_2_digit_codes.csv")
country_codes.head()
# + [markdown] id="m08-Ohns-SiN"
# Merge both datasets by 2 digit code, and fill `NaN`s with full country name:
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="QmHuusIJ-MYX" outputId="7ff1ef0f-bb23-4625-eda0-388fd0cefefe"
glassdoor = pd.merge(glassdoor, country_codes, left_on='Country', right_on='Code', how='left')
glassdoor.head()
# + [markdown] id="dtYm0DRX-Yn7"
# Then replace the 2 digit codes with full name:
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="TxwJ8Slt-dRx" outputId="25d6ac4a-ffee-4f43-d6af-f237a3659421"
glassdoor["Country"] = glassdoor["Name"].fillna(glassdoor["Country"])
glassdoor.head()
# + [markdown] id="N-dtsJuE-vIX"
# Lastly, drop `Name` and `Code` columns:
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="1FKOjJcr--uR" outputId="18f88b5b-d0bd-427b-9701-5a0212f880ae"
glassdoor = glassdoor.drop(["Name", "Code"], axis=1)
glassdoor.head()
# + [markdown] id="IdN-OKrK_3zr"
# Remove some countries that didn't match official nomenclature
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="R9epzeAm_GM1" outputId="88fed3bf-2356-42b7-f26d-f6350a0044f4"
glassdoor = pd.merge(glassdoor, country_codes, left_on='Country', right_on='Name', how='left')
glassdoor.head()
# + id="4D6nrpqo_rU-"
glassdoor.dropna(subset=['Name'], inplace=True)
glassdoor = glassdoor.drop(['Name', 'Code'], axis=1)
# + [markdown] id="svrS-_0L_14Z"
# #### Processing `JobTitle`
#
# + [markdown] id="wcdUgr34ApwL"
# Job titles we're interested in:
# + id="Ib96m3uAADXf"
JOB_TITLES = [
'data scientist',
'software engineer',
'data analyst',
'research scientist',
'business analyst',
'data engineer',
'statistician',
'dba',
'database engineer',
'machine learning engineer'
]
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="7Z28h09IAIiI" outputId="54251825-861d-4c3b-e466-582607856231"
import re
import numpy as np
job_masks = [glassdoor.JobTitle.str.contains(job_title, flags=re.IGNORECASE, regex=True) for job_title in JOB_TITLES]
combined_mask = np.vstack(job_masks).any(axis=0)
glassdoor = glassdoor[combined_mask].reset_index(drop=True)
glassdoor.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="DHK6EWFQA8oJ" outputId="3c6659b8-91f4-4fca-e937-94c92b97a454"
job_titles_regex = '|'.join(JOB_TITLES)
glassdoor.JobTitle = glassdoor.JobTitle.str.findall(job_titles_regex, flags=re.IGNORECASE)
glassdoor.JobTitle = glassdoor.JobTitle.str[0]
glassdoor.JobTitle = glassdoor.JobTitle.str.title()
glassdoor.head()
# + [markdown] id="Rplu-jmPB4Ll"
# Rename some job titles:
# + colab={"base_uri": "https://localhost:8080/"} id="TJuVZcqgBuvQ" outputId="d209cdac-8d70-434c-9901-7f2ca43b5a7d"
glassdoor.JobTitle = glassdoor.JobTitle.replace('Dba', 'Data Engineer/DBA')
glassdoor.JobTitle = glassdoor.JobTitle.replace('Database Engineer', 'Data Engineer/DBA')
glassdoor.JobTitle = glassdoor.JobTitle.replace('Data Engineer', 'Data Engineer/DBA')
# group Statistician + Research Scientist
glassdoor.JobTitle = glassdoor.JobTitle.replace('Statistician', 'Statistician/Research Scientist')
glassdoor.JobTitle = glassdoor.JobTitle.replace('Research Scientist', 'Statistician/Research Scientist')
glassdoor.JobTitle.value_counts()
# + [markdown] id="5vOYYX7qB9Su"
# Then we tranform `JobTitle` column into category:
# + id="0R4ZwkQdCHQ-"
job_titles = ['Business Analyst', 'Data Analyst', 'Data Scientist',
'Data Engineer/DBA', 'Software Engineer', 'Statistician/Research Scientist']
cat_dtype = pd.api.types.CategoricalDtype(categories=job_titles, ordered=True)
glassdoor.JobTitle = glassdoor.JobTitle.astype(cat_dtype)
# + id="-1h8IyS3CZ2T"
# Add column to make groupby easier
glassdoor['Count'] = 1
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="Jt_7l5a4CQZK" outputId="30a40a5e-0133-497c-e87d-d848677fae9a"
glassdoor.head()
# + [markdown] id="jk6-JfM7CfdD"
# #### Processing `JobDescription`
# + [markdown] id="WF1JrHEmCl12"
# Make `JobDescription` lower case:
# + id="16jmIQ67Jatv"
glassdoor.JobDescription = glassdoor.JobDescription.str.lower()
# + [markdown] id="VP4vPZ9hJgwf"
# Find metions to cloud platforms and databases in `JobDescription`:
# + id="J6HYRt6XJ9u3"
cloud_platforms = {
'Alibaba': ' Alibaba Cloud ',
'Amazon Web Services': ' Amazon Web Services (AWS) ',
'AWS': ' Amazon Web Services (AWS) ',
'Google Cloud Platform': ' Google Cloud Platform (GCP) ',
'GCP': ' Google Cloud Platform (GCP) ',
'Google Cloud': ' Google Cloud Platform (GCP) ',
'IBM': ' IBM Cloud ',
'Azure': ' Microsoft Azure ',
'Oracle': ' Oracle Cloud ',
'Red Hat': ' Red Hat Cloud ',
'SAP': ' SAP Cloud ',
'Salesforce': ' Salesforce Cloud ',
'VMware': ' VMware Cloud '
}
# Replacing terms into Job Description
for find, repl in cloud_platforms.items():
glassdoor.JobDescription = glassdoor.JobDescription.str.replace(find.lower(), repl.lower())
# + id="DEhh8fPZKAbu"
databases ={
'dynamodb': ' aws dynamodb ',
'dynamo': ' aws dynamodb ',
' rds ': ' aws relational database service ',
'relational database service': ' aws relational database service ',
'azure sql': ' azure sql database ',
'google cloud sql': ' google cloud sql ',
'microsoft access': ' microsoft access ',
'sql server': ' microsoft sql server ',
'my sql': ' mysql ',
'oracle db': ' oracle database ',
'postgres': ' postgressql ',
'postgre': ' postgressql ',
'postgre sql': ' postgressql ',
'sqlite': 'sqlite '
}
for find, repl in databases.items():
glassdoor.JobDescription = glassdoor.JobDescription.str.replace(find.lower(), repl.lower())
# + id="xynmoXRjcZBZ"
glassdoor.to_csv("data/glassdoor_preprocessed.csv")
# + [markdown] id="pGC7MoWjKDI7"
# ## Data visualization
# + [markdown] id="Dtjn5iQtqVsb"
# ### Proportion for each job title
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="T01LBn2RLn_c" outputId="9a932cc0-a22b-45b9-cdfe-d653cd26b665"
glassdoor.head()
# + colab={"base_uri": "https://localhost:8080/"} id="juiQVfJLajD2" outputId="02400e1a-399c-4a7d-c784-82d13b5129f1"
glassdoor_de = glassdoor[glassdoor.Country == "Germany"].groupby(["JobTitle"], as_index=False).Count.sum().Count.tolist()
glassdoor_de = (np.array(glassdoor_de) / sum(glassdoor_de) * 100).tolist()
glassdoor_de
# + colab={"base_uri": "https://localhost:8080/"} id="vhqPB3ZqeQgw" outputId="bd0a7241-5643-492c-b26c-6e450d0cf347"
glassdoor_fr = glassdoor[glassdoor.Country == "France"].groupby(["JobTitle"], as_index=False).Count.sum().Count.tolist()
glassdoor_fr = (np.array(glassdoor_fr) / sum(glassdoor_fr) * 100).tolist()
glassdoor_fr
# + id="U6Yzx4GMuw6g"
df_de = pd.DataFrame(dict(r=glassdoor_de, theta=job_titles))
df_fr = pd.DataFrame(dict(r=glassdoor_fr, theta=job_titles))
# + id="LtVrQ6dOiV5z"
import plotly.graph_objects as go
class PolarPlot():
def __init__(self):
self.figure = go.Figure() # instatiates plotly figure
self.range = (0, 0) # define the initial range of polar plots
self.theta = ['Business Analyst', 'Data Analyst', 'Data Scientist', 'Data Engineer/DBA',
'Software Engineer', 'Statistician/Research Scientist', 'Business Analyst'] # Those are the Theta values for our plot
def update_common_layout(self):
"""
Updates general layout characteristics
"""
self.figure.update_layout(
showlegend = True,
legend_itemclick = 'toggleothers',
legend_itemdoubleclick = 'toggle',
width = 800,
height = 500
)
def update_commom_polar_layout(self):
"""
Updates polar layout characteristics
"""
self.figure.update_layout(
# polar_bgcolor='white', # White background is always better
# polar_radialaxis_visible=True, # we want to show the axis
# polar_radialaxis_showticklabels=True, # we want to show the axis titles
# polar_radialaxis_tickfont_color='darkgrey', # grey to the axis label (Software Engineer, Data Scientist, etc)
# polar_angularaxis_color='grey', # Always grey for all elements that are not important
# polar_angularaxis_showline=False, # hide lines that are not necessary
# polar_radialaxis_showline=False, # hide lines that are not necessary
# polar_radialaxis_layer='below traces', # show the axis bellow all traces
# polar_radialaxis_gridcolor='#F2F2F2', # grey to not draw attention
polar_radialaxis_range=self.range # gets the range attribute, that is calculated in another method
)
def add_data(self, data, country, hover_template='%{r:0.0f}%'):
"""
Adds a trace to the figure following the same standard for each trace
"""
data.append(data[0]) # add the first element to the end of the list, this will "close" the polar chart
self.figure.add_trace(
go.Scatterpolar(
r=data,
theta=self.theta,
mode='lines',
name=country,
hoverinfo='name+r',
hovertemplate=hover_template,
showlegend=True,
line_shape='spline',
line_smoothing=0.8,
# line_width=1.6
)
)
self.update_range(data) # Calls the method that will update the max range
def update_range(self, data):
"""
Updates the range to be 110% of maximum value of all traces
"""
max_range = max(data) * 1.1
self.range = (0, max_range) if max_range > self.range[1] else self.range # updates the range attribute
def show(self):
"""
Update layouts and shows the figure
"""
self.update_common_layout()
self.update_commom_polar_layout()
self.figure.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 517} id="3XMrcPOQiwCg" outputId="137f7b78-061c-4316-f4d8-c82e4e1225e1"
pp = PolarPlot()
# example plot
pp.add_data(glassdoor_de, "Germany")
pp.add_data(glassdoor_fr, "France")
pp.show()
# + id="zGDMbrZjSFy3"
def remove_trace(fig, name):
fig.data = tuple(trace for trace in fig.data if trace["name"] != name)
# + colab={"base_uri": "https://localhost:8080/", "height": 517} id="uFMOO08iS3iV" outputId="1ad0abce-525a-41e9-dc34-77375e8ada29"
remove_trace(pp.figure, "France")
pp.show()
# + [markdown] id="oTHrevYAldSa"
# ### Salaries
# + id="WfFWZ-3btrpd"
class LinePlot():
def __init__(self):
self.figure = go.Figure()
self.range = (0, 100)
def update_axis_title(self, x, y):
self.figure.update_layout(
xaxis_title_text=x,
yaxis_title_text=y,
)
def update_layout(self):
"""
Creates a clean layout for ploting, adjusting multiple settings
"""
self.figure.update_layout(
showlegend=True,
legend_font_color='gray',
legend_itemclick='toggleothers',
legend_itemdoubleclick='toggle',
width = 800,
height=500,
# plot_bgcolor='white',
xaxis_title_font_color='grey',
xaxis_color='grey',
yaxis_title_font_color='grey',
yaxis_color='grey',
)
def add_data(self, x_names, y_data, trace_name, hover_template):
"""
Adds a trace to the figure following the same standard for each trace
"""
self.figure.add_trace(
go.Scatter(
x=x_names,
y=y_data,
mode='lines',
name=trace_name,
hoverinfo='name+y',
hovertemplate=hover_template,
line_shape='spline',
line_smoothing=0.8,
line_width=1.6
)
)
def show(self):
self.update_layout()
self.figure.show()
# + id="MFyFuqg1t7m_"
def plot_lines(line_plot, data, traces, x_names, agg_column, group_column, trace_column, hover_template):
"""
Creates aggregation to plot
"""
for trace_name in traces:
data_filtered = data[data[trace_column] == trace_name]
plot_data = data_filtered.groupby([group_column], as_index=False).agg({agg_column: ['mean', 'count']})
plot_data = plot_data[agg_column]['mean'].tolist()
line_plot.add_data(x_names, plot_data, trace_name, hover_template=hover_template)
# + [markdown] id="08R4B17v4lia"
# #### Time of coding
# + colab={"base_uri": "https://localhost:8080/", "height": 517} id="bKHZVyz0t87B" outputId="16b1d2d3-eabd-4193-8cc3-d6978732f3f9"
JOB_TITLES = ['Business Analyst', 'Data Analyst', 'Data Scientist', 'Data Engineer/DBA',
'Software Engineer', 'Statistician/Research Scientist']
x_names = ['0 years', '< 1 years', '1-2 years', '3-5 years', '5-10 years', '10-20 years', '20+ years']
time_of_coding_line_plot = LinePlot()
plot_lines(
time_of_coding_line_plot,
data=kaggle,
traces=JOB_TITLES,
x_names=x_names,
agg_column='Salary',
group_column='TimeWritingCode',
trace_column='JobTitle',
hover_template='U$%{y:,.2r}'
)
xaxis_title='Time of writing code'
yaxis_title='Average Salary (USD per Year)'
time_of_coding_line_plot.update_axis_title(xaxis_title, yaxis_title)
time_of_coding_line_plot.show()
# + [markdown] id="H8q7v65UuauG"
# #### Company size
# + colab={"base_uri": "https://localhost:8080/", "height": 517} id="upAPzW4o4t78" outputId="6f5d1d79-e99e-497e-d935-cb94dbcab197"
salary_line_plot = LinePlot()
COMPANY_SIZE = [
'0-49 employees',
'50-249 employees',
'250-999 employees',
'1000-9,999 employees',
'> 10,000 employees'
]
plot_lines(
salary_line_plot,
data=kaggle,
traces=JOB_TITLES,
x_names=COMPANY_SIZE,
agg_column='Salary',
group_column='CompanySize',
trace_column='JobTitle',
hover_template='U$%{y:,.2r}'
)
xaxis_title='Company size'
yaxis_title='Average Salary (USD per Year)'
salary_line_plot.update_axis_title(xaxis_title, yaxis_title)
salary_line_plot.show()
# + [markdown] id="1KU2KFfw5Hd0"
# ### Job skills
# + id="0eswEtE27IUy"
def plot_polar(polar_plot, data, traces, x_names, agg_column, group_column, trace_column, hover_template):
data_cp = data.copy()
for trace_name in traces:
if agg_column in ('JobDescription', 'CloudPlatf'):
data_cp['TempCol'] = data_cp[agg_column].apply(lambda x: trace_name.lower() in x)
else:
data_cp['TempCol'] = data_cp[agg_column].apply(lambda x: trace_name in x)
plot_data = data_cp.groupby([group_column], as_index=False).agg({'TempCol': ['sum', 'count']})
plot_data['TempColPct'] = plot_data['TempCol']['sum'] / plot_data['TempCol']['count'] * 100
plot_data = plot_data.TempColPct.tolist()
polar_plot.add_data(plot_data, trace_name, hover_template)
# + [markdown] id="IPM6RA_O8T31"
# #### Popularity of different programming languages in Kaggle
# + colab={"base_uri": "https://localhost:8080/", "height": 517} id="Gfm1RYHb7b4O" outputId="3e6b542f-1e62-4c93-8cb4-c5c899294539"
job_skills_polar_plot = PolarPlot()
SKILLS = ['Bash', 'C', 'C++', 'Java', 'Javascript', 'MATLAB',
'Other', 'Python', 'R', 'SQL', 'TypeScript']
plot_polar(
job_skills_polar_plot,
data=kaggle,
traces=SKILLS,
x_names=JOB_TITLES,
agg_column='ProgLang',
group_column='JobTitle',
trace_column='ProgLang',
hover_template='%{r:0.0f}%'
)
job_skills_polar_plot.figure.update_layout(
polar_radialaxis_tickvals=[25, 50, 75],
polar_radialaxis_ticktext=['25%', '50%', '75%'],
polar_radialaxis_tickmode='array',
)
job_skills_polar_plot.show()
# + [markdown] id="RQcw4DqN8CFx"
# #### Popularity of different programming languages in Job Description
# + colab={"base_uri": "https://localhost:8080/", "height": 517} id="zcMne7ns_O3x" outputId="d7d527d4-d405-4ab8-9f7e-374cf222a694"
job_desc_polar_plot = PolarPlot()
plot_polar(
job_desc_polar_plot,
data=glassdoor,
traces=SKILLS,
x_names=JOB_TITLES,
agg_column='JobDescription',
group_column='JobTitle',
trace_column='JobDescription',
hover_template='%{r:0.0f}%'
)
job_desc_polar_plot.figure.update_layout(
polar_radialaxis_tickvals=[25, 50, 75],
polar_radialaxis_ticktext=['25%', '50%', '75%'],
polar_radialaxis_tickmode='array',
)
job_desc_polar_plot.show()
# + [markdown] id="DZR2zcHa_a9c"
# ### How many programming languages should you learn?
# + colab={"base_uri": "https://localhost:8080/", "height": 517} id="661uqWpr_soj" outputId="f3530ed6-5f6c-4a3d-bca5-b3d040cfa504"
prog_language_line_plot = LinePlot()
traces = list(set(kaggle.TimeWritingCode.tolist()))
# traces = []
x_names = ['{} languages'.format(x) for x in range(7)]
plot_lines(
prog_language_line_plot,
data=kaggle,
traces=traces,
x_names=x_names,
agg_column='Salary',
group_column='QtyProgLang',
trace_column='TimeWritingCode',
hover_template='U$%{y:,.2r}'
)
# Adding Averarage
plot_data = kaggle.groupby(['QtyProgLang'], as_index=False).agg({'Salary': 'mean'})
plot_data = plot_data.Salary.tolist()
prog_language_line_plot.add_data(x_names, plot_data, 'Average', hover_template='U$%{y:,.2r}')
xaxis_title='Quantity of programming languages'
yaxis_title='Average Salary (USD per Year)'
prog_language_line_plot.update_axis_title(xaxis_title, yaxis_title)
prog_language_line_plot.show()
| talent_worth.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MAX318955 Temperature Sensor
# To run this script, select `Run` → `Run All Cells` from the menu.
# +
## Don't modify this line! :)
config = {}
## MQTT - our client id
config['mqtt_id'] = 'MCP3204-Reader'
## MQTT - broker ip / hostname
config['mqtt_broker'] = 'iot.fh-muenster.de'
## MQTT - broker username / password
config['mqtt_broker_user'] = 'MQTT_USER'
config['mqtt_broker_pass'] = '<PASSWORD>'
## MQTT - empty sset of topics
config['mqtt_topics'] = set([])
## MQTT - subscribed topics
config['mqtt_topics'].add('sensor/3C:71:BF:AA:7B:18')
#config['mqtt_topics'].add('sensor/60:01:94:4A:A8:95')
## Recording - stop the program if the defined amount of samples has been recorded (0 = unlimited)
config['record_sample_limit'] = 0
## Recording - stop the program if the defined amount of seconds has passed (0 = unlimited)
config['record_duration_limit'] = 10
## Display - upper limit of samples displayed in the bokeh graph
config['bokeh_stream_rollover'] = 250
## Export formats (True/False)
config['export_json'] = False
config['export_csv'] = True
config['export_excel'] = False
# + slideshow={"slide_type": "slide"} tags=["hide_input"]
import MCP3204
MCP3204.record_and_display(config)
# -
import MCP3204
| Python/MCP3204/MAX31855.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from pathlib import Path as pt
import matplotlib.pyplot as plt
from lmfit.models import GaussianModel
import uncertainties as uc
filename = pt("./12_09_19-2.thz")
filename.exists()
# +
# Opening file and reading its content
with open(filename, "r") as fileContents:
file = fileContents.readlines()
file = file[1:]
#############################################
# Resonce ON counts value
resOn = []
for line in file:
if line.startswith("#"): break
line = line.split("\n")[0].split("\t")[:-1]
resOn.append(line)
resOn = resOn[1:]
#############################################
# Resonce OFF counts value
resOff = []
start = False
for line in file:
if line.startswith("# freq"):
start = True
continue
if start:
if line.startswith("#"): break
line = line.split("\n")[0].split("\t")[:-1]
resOff.append(line)
resOff = resOff[1:]
#############################################
resOn = np.array(resOn, dtype=np.float)
resOff = np.array(resOff, dtype=np.float)
#############################################
freq = resOn.T[0]
depletion = (resOff.T[1:] - resOn.T[1:])/resOff.T[1:]
depletion_counts = depletion.T.mean(axis=1)
depletion_error = depletion.T.std(axis=1)
#############################################
depletion_counts = depletion_counts*100
# +
model = GaussianModel()
guess = model.guess(depletion_counts, x=freq)
guess_values = guess.valuesdict()
fit = model.fit(depletion_counts, x=freq, amplitude = guess_values['amplitude'],
center = guess_values['center'],
sigma = guess_values['sigma'],
fwhm = guess_values['fwhm'],
height = guess_values['height'])
fit_data = fit.best_fit
line_freq_fit = fit.best_values['center']
##########################################################################################
FWHM = lambda sigma: 2*np.sqrt(2*np.log(2))*sigma
Amplitude = lambda a, sigma: a/(sigma*np.sqrt(2*np.pi))
fwhm = FWHM(fit.best_values['sigma'])
amplitude = Amplitude(fit.best_values['amplitude'], fit.best_values['sigma'])
half_max = amplitude/2
sigma = fit.best_values['sigma']
##########################################################################################
fig, ax = plt.subplots(figsize=(12, 6), dpi=150)
#ax.errorbar(freq, depletion_counts, yerr=depletion_error, ecolor ="k")
ax.plot(freq, depletion_counts, ".", label="Data (~30${\mu}$W)")
ax.plot(freq, fit_data, label="Fitted")
ax.vlines(line_freq_fit, 0, amplitude, label=f'{line_freq_fit:.7f} GHz')
ax.hlines(half_max, line_freq_fit-fwhm/2, line_freq_fit+fwhm/2, color="C2", label=f"FWHM: {fwhm*1e6:.1f}KHz")
ax.grid()
ax.set(title="CD$^{+}$: j=1-0 line", xlabel="Frequency (GHz)", ylabel="Depletion (%)")
ax.ticklabel_format(useOffset=False)
ax.legend()
plt.savefig('./CD+_j1-0_line')
plt.show()
plt.close()
# -
fit.fit_report()
| testing_module/.ipynb_checkpoints/thz_code-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <table> <tr>
# <td style="background-color:#ffffff;">
# <a href="http://qworld.lu.lv" target="_blank"><img src="..\images\qworld.jpg" width="25%" align="left"> </a></td>
# <td style="background-color:#ffffff;vertical-align:bottom;text-align:right;">
# prepared by <a href="http://abu.lu.lv" target="_blank"><NAME></a> (<a href="http://qworld.lu.lv/index.php/qlatvia/" target="_blank">QLatvia</a>)
# </td>
# </tr></table>
# <table width="100%"><tr><td style="color:#bbbbbb;background-color:#ffffff;font-size:11px;font-style:italic;text-align:right;">This cell contains some macros. If there is a problem with displaying mathematical formulas, please run this cell to load these macros. </td></tr></table>
# $ \newcommand{\bra}[1]{\langle #1|} $
# $ \newcommand{\ket}[1]{|#1\rangle} $
# $ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
# $ \newcommand{\dot}[2]{ #1 \cdot #2} $
# $ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
# $ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
# $ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
# $ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
# $ \newcommand{\mypar}[1]{\left( #1 \right)} $
# $ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
# $ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
# $ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
# $ \newcommand{\onehalf}{\frac{1}{2}} $
# $ \newcommand{\donehalf}{\dfrac{1}{2}} $
# $ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
# $ \newcommand{\vzero}{\myvector{1\\0}} $
# $ \newcommand{\vone}{\myvector{0\\1}} $
# $ \newcommand{\stateplus}{\myvector{ \sqrttwo \\ \sqrttwo } } $
# $ \newcommand{\stateminus}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
# $ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
# $ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
# $ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
# $ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
# $ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
# $ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
# $ \newcommand{\pstate}[1]{ \lceil \mspace{-1mu} #1 \mspace{-1.5mu} \rfloor } $
# $ \newcommand{\bstate}[1]{ [ \mspace{-1mu} #1 \mspace{-1.5mu} ] } $
# <h2>Project | Simulating a Real-Valued Qubit</h2>
#
# We play with a single qubit living on the unit circle.
#
# - _All angles are in radian._
# - _Please do not use any quantum programming library or any scientific python library such as `NumPy`._
# - _You can use python module `math` and python interface `matplotlib.pyplot`._
# ### Create a python class called `SingleQubit(theta=0)`
#
# When an instance is created, the quantum state of qubit is set to the specified angle _theta_. If no angle is specified by the user, then the initial state of the qubit will be $\ket{0}$.
# ### The basic methods
#
# 1. `read_state()`: Return the current quantum state of the qubit.
#
# 1. `rotation(theta)`: Update the current quantum state by applying the rotation operator with the angle _theta_.
#
# 1. `reflection(theta)`: Update the current quantum state by applying the reflection operator having a reflection axis with angle _theta_.
#
# 1. `draw_state()`: Draw the current quantum state as an arrow on the unit circle.
#
# 1. `draw_all_states()`: Draw all the quantum states visited on the unit circle starting from the initial one by indicating their visited orders.
#
# 1. `reflect_and_draw(theta)`: Draw the unit circle, the current quantum state, the reflection line of the reflection operator, apply the method `reflection(theta)` to the current quantum state, and draw the new quantum state.
#
# 1. `prob()`: Return the probabilities of observing state 0 and state 1 if the qubit is measured at this moment.
#
# 1. `measure(number_of_shots)`: Simulate the measurement operator _number_of_shots_ times based on the current quantum state, and then return the number(s) of the observed 0’s and 1’s (remark that only a single state is observed after each measurement, i.e., either 0 or 1).
# ### The extra methods
#
# 9. `change_basis(theta)`: Return an error if the basis will remain the same. Change the basis of qubit with the unit vectors $ \ket{v_1} $ and $ \ket{v_2} $ such that
# - the angles of $ \ket{v_1} $ and $ \ket{v_2} $ are $ \theta $ and $ \theta + \frac{\pi}{2} $ in the standard basis, respectively, and
# - $\ket{v_1}$ becomes the new $ \ket{0} $ and $ \ket{v_2} $ becomes the new $ \ket{1} $; and then,
# update all visited quantum states until now respectively, i.e, each quantum state in the standard basis should be shifted by $ -\theta $.
#
# 2. `draw_state_in_both_basis()`: Return an error if the basis has not changed. Draw two separete unit circles where the basis are the current basis and the previous basis, and then draw the current quantum state as an arrow on each unit circle.
#
# 3. `prob_in_both_basis()`: Return an error if the basis has not changed. Return the probabilities of observing state 0 and state 1 in the current and previous bases.
#
# 4. `take_back_basis_change()`: Do nothing if there is no basis change. Return back to the previous basis by deleting the latest call of `change_basis()` from the basis history. On the other hand, any quantum state visited after the latest call of `change_basis()` must be kept.
| projects/Project_Simulating_a_RealValued_Qubit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PyCharm (open_cv_study)
# language: python
# name: pycharm-2c40972a
# ---
# # Data Manipulation
#
# It is impossible to get anything done if we cannot manipulate data. Generally, there are two important things we need to do with data: (i) acquire it and (ii) process it once it is inside the computer. There is no point in acquiring data if we do not even know how to store it, so let's get our hands dirty first by playing with synthetic data. We will start by introducing the tensor,
# PyTorch's primary tool for storing and transforming data. If you have worked with NumPy before, you will notice that tensors are, by design, similar to NumPy's multi-dimensional array. Tensors support asynchronous computation on CPU, GPU and provide support for automatic differentiation.
书上2.2
# ## Getting Started
import torch
# Tensors represent (possibly multi-dimensional) arrays of numerical values.
# The simplest object we can create is a vector. To start, we can use `arange` to create a row vector with 12 consecutive integers.
x = torch.arange(12, dtype=torch.float64)
x
# We can get the tensor shape through the shape attribute.
x.shape
# .shape is an alias for .size(), and was added to more closely match numpy
# 在numpy里面size返回的是实例中元素(element)的总数
x.size()
# We use the `reshape` function to change the shape of one (possibly multi-dimensional) array, to another that contains the same number of elements.
# For example, we can transform the shape of our line vector `x` to (3, 4), which contains the same values but interprets them as a matrix containing 3 rows and 4 columns. Note that although the shape has changed, the elements in `x` have not.
x = x.reshape((3, 4))
x
# Reshaping by manually specifying each of the dimensions can get annoying. Once we know one of the dimensions, why should we have to perform the division our selves to determine the other? For example, above, to get a matrix with 3 rows, we had to specify that it should have 4 columns (to account for the 12 elements). Fortunately, PyTorch can automatically work out one dimension given the other.
# We can invoke this capability by placing `-1` for the dimension that we would like PyTorch to automatically infer. In our case, instead of
# `x.reshape((3, 4))`, we could have equivalently used `x.reshape((-1, 4))` or `x.reshape((3, -1))`.
# 当我们已知x的元素个数,那么-1是可以通过元素个数和其他维度的大小推断出来的
torch.FloatTensor(2, 3)
torch.Tensor(2, 3)
torch.empty(2, 3)
# torch.Tensor() is just an alias to torch.FloatTensor() which is the default type of tensor, when no dtype is specified during tensor construction.
#
# From the torch for numpy users notes, it seems that torch.Tensor() is a drop-in replacement of numpy.empty()
#
# So, in essence torch.FloatTensor() and torch.empty() does the same job.
#
# torch.FloatTensor() 和torch.empty()做了同样的工作
#
# The `empty` method just grabs some memory and hands us back a matrix without setting the values of any of its entries. This is very efficient but it means that the entries might take any arbitrary values, including very big ones! 注意,使用上述三种声明方式创建出来的数组是非常随机的,相当于只声明了空间。 Typically, we'll want our matrices initialized either with ones, zeros, some known constant or numbers randomly sampled from a known distribution.
#
# Perhaps most often, we want an array of all zeros. To create tensor with all elements set to 0 and a shape of (2, 3, 4) we can invoke:
torch.zeros((2, 3, 4))
# We can create tensors with each element set to 1 works via
torch.ones((2, 3, 4))
# We can also specify the value of each element in the desired NDArray by supplying a Python list containing the numerical values.
y = torch.tensor([[2, 1, 4, 3], [1, 2, 3, 4], [4, 3, 2, 1]])
y
# In some cases, we will want to randomly sample the values of each element in the tensor according to some known probability distribution. This is especially common when we intend to use the tensor as a parameter in a neural network. The following snippet creates an tensor with a shape of (3,4). Each of its elements is randomly sampled in a normal distribution with zero mean and unit variance.
torch.randn(3, 4)
# ## Operations
#
# Oftentimes, we want to apply functions to arrays. Some of the simplest and most useful functions are the element-wise functions. These operate by performing a single scalar operation on the corresponding elements of two arrays. We can create an element-wise function from any function that maps from the scalars to the scalars. In math notations we would denote such a function as $f: \mathbb{R} \rightarrow \mathbb{R}$. Given any two vectors $\mathbf{u}$ and $\mathbf{v}$ *of the same shape*, and the function f,
# we can produce a vector $\mathbf{c} = F(\mathbf{u},\mathbf{v})$ by setting $c_i \gets f(u_i, v_i)$ for all $i$. Here, we produced the vector-valued $F: \mathbb{R}^d \rightarrow \mathbb{R}^d$ by *lifting* the scalar function to an element-wise vector operation. In PyTorch, the common standard arithmetic operators (+,-,/,\*,\*\*) have all been *lifted* to element-wise operations for identically-shaped tensors of arbitrary shape. We can call element-wise operations on any two tensors of the same shape, including matrices.
x = torch.tensor([1, 2, 4, 8], dtype=torch.float32)
y = torch.ones_like(x) * 2
print('x =', x)
print('x + y', x + y)
print('x - y', x - y)
# 按位乘法和按位除法
print('x * y', x * y)
print('x / y', x / y)
# Many more operations can be applied element-wise, such as exponentiation:
torch.exp(x)
# Note: torch.exp is not implemented for 'torch.LongTensor'.
# In addition to computations by element, we can also perform matrix operations, like matrix multiplication using the `mm` or `matmul` function. Next, we will perform matrix multiplication of `x` and the transpose of `y`. We define `x` as a matrix of 3 rows and 4 columns, and `y` is transposed into a matrix of 4 rows and 3 columns. The two matrices are multiplied to obtain a matrix of 3 rows and 3 columns.
x = torch.arange(12, dtype=torch.float32).reshape((3,4))
y = torch.tensor([[2, 1, 4, 3], [1, 2, 3, 4], [4, 3, 2, 1]], dtype=torch.float32)
print(x.dtype)
print(y)
# 矩阵乘法,需要维数match
torch.mm(x, y.t())
# Note that torch.dot() behaves differently to np.dot(). There's been some discussion about what would be desirable here. Specifically, torch.dot() treats both a and b as 1D vectors (irrespective of their original shape) and computes their inner product.
#
# 注意torch.dot()只能用于两个张量的点积(内积),返回的是一个标量。
#
# torch.dot(torch.tensor([2, 3]), torch.tensor([2, 1])) #即对应位置相乘再相加
#
# 返回7 tensor(7)
#
#
# torch.dot(torch.rand(2, 3), torch.rand(2, 2)) #报错,只允许一维的tensor
# **Concatenate连结操作**
#
# We can also merge multiple tensors. For that, we need to tell the system along which dimension to merge. The example below merges two matrices along dimension 0 (along rows) and dimension 1 (along columns) respectively.
torch.cat((x, y), dim=0)
torch.cat((x, y), dim=1)
# Sometimes, we may want to construct binary tensors via logical statements. Take `x == y` as an example. If `x` and `y` are equal for some entry, the new tensor has a value of 1 at the same position; otherwise it is 0.
#
# 使用条件判别式可以得到元素为0或1的新array,在下面例子中X,Y在相同位置的条件判断为真
x == y
# Summing all the elements in the tensor yields an tensor with only one element.
x.sum()
# We can transform the result into a scalar in Python using the `asscalar` function of `numpy`(from numpy v1.16 onwards it issues a deprication warning. We can ignore that at this moment). In the following example, the $\ell_2$ norm of `x` yields a single element tensor. The final result is transformed into a scalar.
import numpy as np
np.asscalar(x.norm())
# ## Broadcast Mechanism 广播机制
#
# In the above section, we saw how to perform operations on two tensors of the same shape. When their shapes differ, a broadcasting mechanism may be triggered analogous to NumPy: first, copy the elements appropriately so that the two tensors have the same shape, and then carry out operations by element.
a = torch.arange(3, dtype=torch.float).reshape((3, 1))
b = torch.arange(2, dtype=torch.float).reshape((1, 2))
a, b
# Since `a` and `b` are (3x1) and (1x2) matrices respectively, their shapes do not match up if we want to add them. PyTorch addresses this by 'broadcasting' the entries of both matrices into a larger (3x2) matrix as follows: for matrix `a` it replicates the columns, for matrix `b` it replicates the rows before adding up both element-wise.
# a,b理论上维数不同无法进行相加,但是由于广播机制,a,b都相加时进行维数match(复制),从而得以相加
a + b
# ## Indexing and Slicing
#
# Just like in any other Python array, elements in a tensor can be accessed by its index. In good Python tradition the first element has index 0 and ranges are specified to include the first but not the last element. By this logic `1:3` selects the second and third element. Let's try this out by selecting the respective rows in a matrix.
x[1:3]
# Beyond reading, we can also write elements of a matrix.
x[1, 2] = 9
x
# If we want to assign multiple elements the same value, we simply index all of them and then assign them the value. For instance, `[0:2, :]` accesses the first and second rows. While we discussed indexing for matrices, this obviously also works for vectors and for tensors of more than 2 dimensions.
x[0:2, :] = 12
x
# ## Saving Memory
#
# In the previous example, every time we ran an operation, we allocated new memory to host its results. For example, if we write `y = x + y`, we will dereference the matrix that `y` used to point to and instead point it at the newly allocated memory. In the following example we demonstrate this with Python's `id()` function, which gives us the exact address of the referenced object in memory. After running `y = y + x`, we will find that `id(y)` points to a different location. That is because Python first evaluates `y + x`, allocating new memory for the result and then subsequently redirects `y` to point at this new location in memory.
before = id(y)
y = y + x
id(y) == before
# This might be undesirable for two reasons. First, we do not want to run around allocating memory unnecessarily all the time. In machine learning, we might have hundreds of megabytes of parameters and update all of them multiple times per second. Typically, we will want to perform these updates *in place*. Second, we might point at the same parameters from multiple variables. If we do not update in place, this could cause a memory leak, making it possible for us to inadvertently reference stale parameters.
#
# Fortunately, performing in-place operations in PyTorch is easy. We can assign the result of an operation to a previously allocated array with slice notation, e.g., `y[:] = <expression>`. To illustrate the behavior, we first clone the shape of a matrix using `zeros_like` to allocate a block of 0 entries.
z = torch.zeros_like(y)
print('id(z):', id(z))
z[:] = x + y
print('id(z):', id(z))
# While this looks pretty, `x+y` here will still allocate a temporary buffer to store the result of `x+y` before copying it to `z[:]`. To make even better use of memory, we can directly invoke the underlying `tensor` operation, in this case `add`, avoiding temporary buffers. We do this by specifying the `out` keyword argument, which every `tensor` operator supports:
before = id(z)
torch.add(x, y, out=z)
id(z) == before
# If the value of `x ` is not reused in subsequent computations, we can also use `x[:] = x + y` or `x += y` to reduce the memory overhead of the operation.
before = id(x)
x += y
id(x) == before
# ## Mutual Transformation of PyTorch and NumPy torch矩阵和numpy矩阵的转换
#
# Converting PyTorch Tensors to and from NumPy Arrays is easy. The converted arrays do *not* share memory. This minor inconvenience is actually quite important: when you perform operations on the CPU or one of the GPUs, you do not want PyTorch having to wait whether NumPy might want to be doing something else with the same chunk of memory. `.tensor` and `.numpy` do the trick.
a = x.numpy()
print(type(a))
b = torch.tensor(a)
print(type(b))
# ## Exercises
#
# 1. Run the code in this section. Change the conditional statement `x == y` in this section to `x < y` or `x > y`, and then see what kind of tensor you can get.
# 1. Replace the two tensors that operate by element in the broadcast mechanism with other shapes, e.g. three dimensional tensors. Is the result the same as expected?
# 1. Assume that we have three matrices `a`, `b` and `c`. Rewrite `c = torch.mm(a, b.t()) + c` in the most memory efficient manner.
x > y
x < y
| Ch04_The_Preliminaries_A_Crashcourse/Data_Manipulation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib ipympl
import ipywidgets as widgets
import matplotlib.pyplot as plt
import numpy as np
# %load_ext autoreload
# %autoreload 2
from mpl_interactions import *
# +
fig, ax = plt.subplots()
def y(heck):
return heck
def xmin(heck):
return heck
# controls = interactive_axhline(y,0,1, heck=(0,1))
controls = interactive_axhline(y, xmin, 1, heck=(0, 1))
# ret.set_ydata([.5,1])
# -
plt.axhline()
# +
def interactive_axhline(
y=0,
xmin=0,
xmax=1,
ax=None,
slider_formats=None,
title=None,
force_ipywidgets=False,
play_buttons=False,
play_button_pos="right",
controls=None,
display_controls=True,
**kwargs,
):
"""
Control an horizontal line using widgets.
parameters
----------
y : float or function
y position in data coordinates of the horizontal line.
xmin : float or function
Should be between 0 and 1, 0 being the far left of the plot, 1 the
far right of the plot.
xmax : float or function
Should be between 0 and 1, 0 being the far left of the plot, 1 the
far right of the plot.
ax : matplotlib axis, optional
If None a new figure and axis will be created
slider_formats : None, string, or dict
If None a default value of decimal points will be used. Uses the new {} style formatting
force_ipywidgets : boolean
If True ipywidgets will always be used, even if not using the ipympl backend.
If False the function will try to detect if it is ok to use ipywidgets
If ipywidgets are not used the function will fall back on matplotlib widgets
play_buttons : bool or dict, optional
Whether to attach an ipywidgets.Play widget to any sliders that get created.
If a boolean it will apply to all kwargs, if a dictionary you choose which sliders you
want to attach play buttons too.
play_button_pos : str, or dict, or list(str)
'left' or 'right'. Whether to position the play widget(s) to the left or right of the slider(s)
controls : mpl_interactions.controller.Controls
An existing controls object if you want to tie multiple plot elements to the same set of
controls
display_controls : boolean
Whether the controls should display on creation. Ignored if controls is specified.
returns
-------
controls
"""
ipympl = notebook_backend()
fig, ax = gogogo_figure(ipympl, ax)
use_ipywidgets = ipympl or force_ipywidgets
slider_formats = create_slider_format_dict(slider_formats)
controls, params = gogogo_controls(
kwargs, controls, display_controls, slider_formats, play_buttons, play_button_pos
)
def update(params, indices, cache):
if title is not None:
ax.set_title(title.format(**params))
y_ = callable_else_value(y, params, cache).item()
line.set_ydata([y_, y_])
xmin_ = callable_else_value(xmin, params, cache).item()
xmax_ = callable_else_value(xmax, params, cache).item()
line.set_xdata([xmin_, xmax_])
# TODO consider updating just the ydatalim here
controls.register_function(update, fig, params)
line = ax.axhline(
callable_else_value(y, params).item(),
callable_else_value(xmin, params).item(),
callable_else_value(xmax, params).item(),
)
return controls
def interactive_axvline(
x=0,
ymin=0,
ymax=1,
ax=None,
slider_formats=None,
title=None,
force_ipywidgets=False,
play_buttons=False,
play_button_pos="right",
controls=None,
display_controls=True,
**kwargs,
):
ipympl = notebook_backend()
fig, ax = gogogo_figure(ipympl, ax)
use_ipywidgets = ipympl or force_ipywidgets
slider_formats = create_slider_format_dict(slider_formats)
controls, params = gogogo_controls(
kwargs, controls, display_controls, slider_formats, play_buttons, play_button_pos
)
def update(params, indices, cache):
if title is not None:
ax.set_title(title.format(**params))
x_ = callable_else_value(x, params, cache).item()
line.set_ydata([x_, x_])
ymin_ = callable_else_value(ymin, params, cache).item()
ymax_ = callable_else_value(ymax, params, cache).item()
line.set_xdata([ymin_, ymax_])
# TODO consider updating just the ydatalim here
controls.register_function(update, fig, params)
line = ax.axvline(
callable_else_value(x, params).item(),
callable_else_value(ymin, params).item(),
callable_else_value(ymax, params).item(),
)
return controls
# -
ret.set_ydata([0.5, 1])
ret = ax.axhline(0, np.array(0), 1)
arr = np.array(0)
arr.item()
np.asscalar(arr)
ret.set_xdata([0, 1.5])
def axhline(y=0, xmin=0, xmax=1):
ret.get_ydata()
| docs/examples/devlop/devlop-axline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def sol(lst):
new_lst =[]
for i in lst:
# compare against earlier tuples in new_lst
for j in new_lst:
if len(i) == len(j) and sum(i) == sum(j):
break
else:
new_lst.append(i)
return new_lst
#lst = [(1, 2, 0), (3, 0), (2, 1)]
lst = [(1, 2, 0), (3, 0, 0), (0, 2, 1)]
print(sol(lst))
| Intermediate programs/ 32. Remove Equilength and Equisum Tuple Duplicates from List..ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import statsmodels.api as sm
from sklearn.linear_model import LinearRegression, Lasso, Ridge, LassoCV, RidgeCV
from sklearn.cross_decomposition import PLSRegression
from numpy.linalg import svd
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
data = pd.read_excel("single_name_return_data.xlsx", sheet_name="total returns").set_index("Date")
data.head(5)
# # 1 Penalized Regression
# ## 1
# ### a)
y = data['PFE']
X = data.drop(columns=['PFE'])
model_ols = LinearRegression().fit(X,y)
params_ols = pd.DataFrame(index=X.columns, columns=['Intercept', 'Beta'])
params_ols['Intercept'] = model_ols.intercept_
params_ols['Beta'] = model_ols.coef_
params_ols
# ### b)
model_ols.score(X, y)
# ### c)
params_ols['Beta'].abs().idxmax()
# ### d)
params_ols['Beta*vol'] = params_ols['Beta'] * X.std()
params_ols
params_ols['Beta*vol'].abs().idxmax()
# ## 2
# ### a)
model_ridge = Ridge(alpha=0.5).fit(X,y)
params_ridge = pd.DataFrame(index=X.columns, columns=['Intercept', 'Beta'])
params_ridge['Intercept'] = model_ridge.intercept_
params_ridge['Beta'] = model_ridge.coef_
params_ridge
# ### b)
model_ridge.score(X, y)
# ### c)
params_ridge['Beta*vol'] = params_ridge['Beta'] * X.std()
params_ridge
params_ridge['Beta*vol'].abs().idxmax()
# ## 3
# ### a)
model_lasso = Lasso(alpha=3e-4).fit(X,y)
params_lasso = pd.DataFrame(index=X.columns, columns=['Intercept', 'Beta'])
params_lasso['Intercept'] = model_lasso.intercept_
params_lasso['Beta'] = model_lasso.coef_
params_lasso
# ### b)
model_lasso.score(X, y)
# ### c)
params_lasso['Beta*vol'] = params_lasso['Beta'] * X.std()
params_lasso
params_lasso['Beta*vol'].abs().idxmax()
# ### d)
(params_lasso['Beta']!=0).sum()
# ### e)
penalty = 1000
while (Lasso(alpha=penalty*1e-6).fit(X,y).coef_ != 0).sum() != 3:
penalty += 1
print(f'The penalty is: {penalty*1e-6}')
# ## 4
# ### a)
iterables = [["OLS", "Ridge", "LASSO"], ["Intercept", "Beta"]]
params = pd.DataFrame(columns=pd.MultiIndex.from_product(iterables), index=X.columns)
params['OLS'] = params_ols[['Intercept', 'Beta']]
params['Ridge'] = params_ridge[['Intercept', 'Beta']]
params['LASSO'] = params_lasso[['Intercept', 'Beta']]
params
fig, ax = plt.subplots(figsize=(20,8))
ax.plot(params[('OLS', 'Beta')])
ax.plot(params[('Ridge', 'Beta')], color='darkorange')
ax.plot(params[('LASSO', 'Beta')], color='darkgreen', alpha=0.7)
ax.legend(['OLS', 'Ridge', 'LASSO'])
ax.set_title("Beta Comparison")
plt.show()
# - Only LASSO gives us zero estimated betas
# - They all gives both positive and negative values
# - Ridge and LASSO yield smaller variation magnitude, comparing to OLS
# ### b)
# OLS gives the largest R-squared. This is not surprising since both Ridge and LASSO are penalizing the betas from being too influencial to the fitting. In other words, OLS is more prone to overfit the dataset, and therefore is reasonable to have a higher in-sample R-squared.
# ## 5
X_train = X[:"2018"].copy()
y_train = y[:"2018"].copy()
X_test = X["2019":].copy()
y_test = y["2019":].copy()
LinearRegression().fit(X_train,y_train).score(X_test,y_test)
Ridge(alpha=0.5).fit(X_train,y_train).score(X_test,y_test)
Lasso(alpha=3e-4).fit(X_train,y_train).score(X_test,y_test)
# OLS appears to have the best OOS R-squared, but of course this greatly depends on what penalization parameter we choose. If we choose difference alphas, Ridge or LASSO may perform better in OOS estimations.
# ## 6
model_ridgeCV = RidgeCV().fit(X,y)
model_ridgeCV.alpha_
model_ridgeCV.score(X_test, y_test)
model_lassoCV = LassoCV().fit(X,y)
model_lassoCV.alpha_
model_lassoCV.score(X_test, y_test)
# # 2 Principal Components
equities = data.drop(columns=['SPY', 'SHV'])
equities.head(5)
# ## 1
clean = equities - equities.mean()
u, s, vh = svd(clean)
factors = clean @ vh.T
factors.columns = np.arange(1,23)
factors
# ## 2
report = pd.DataFrame(index=factors.columns, columns=['Eigen Value', 'Percentage Explained'])
report['Eigen Value'] = s**2
report['Percentage Explained'] = s**2 / (s**2).sum()
report
# ## 3
report['Percentage Explained'].cumsum()
# We need the first 5 PCs to explain at least 75% of the variation
# ## 4
correlations = factors[[1]].merge(equities, how='left', left_index=True, right_index=True).corr().iloc[0,1:]
correlations
correlations.abs().idxmax()
# ## 5
factors[[1,2,3]].merge(data['SPY'], how='left', left_index=True, right_index=True).corr()
# # 3 Optional
# ## 1
X = factors[[1,2,3]]
y = data['SPY']
model = LinearRegression().fit(X,y)
model.score(X, y)
# ## 2
X = equities
y = data['SPY']
penalty = 1000
while (Lasso(alpha=penalty*1e-6).fit(X,y).coef_ != 0).sum() != 3:
penalty += 1
print(f'The penalty is: {penalty*1e-6}')
Lasso(alpha=penalty*1e-6).fit(X,y).score(X, y)
# ## 3
PLSRegression(n_components=3).fit(X, y).score(X, y)
# ## 4
# Yes, this result makes sense. OLS and PLS are both seeking to maximize the ability to explain the variation in y variable, and therefore we see they have high R-squared. When using LASSO to select only 3 regressors out of 22, we are making an extremely conservative selections. This makes in-sample R-squared lower, but may make more robust OOS predictions.
| notebooks/HW9.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (banet_dev)
# language: python
# name: banet_dev
# ---
# +
# default_exp models
# -
# # Models
#
# > Deep Learning models used to map burned areas
#hide
from nbdev.export import notebook2script
#export
import os, sys
import requests
from fastai.vision.all import *
import FireHR
# +
#export
def expand_filter(x, ks=3):
with torch.no_grad():
k5 = nn.Conv2d(1, 1, kernel_size=ks, padding=ks//2, padding_mode='reflect', bias=False)
k5.weight.data = torch.ones(1, 1, ks, ks)/(ks*ks)
xbuffer = k5(x[:,-1].unsqueeze(1))
x = torch.cat([x[:,:-1], xbuffer], dim=1)
return x
class ChLin(Module):
def __init__(self, ni, nf):
self.chlin = nn.Sequential(
nn.Linear(ni, nf, bias=False), nn.BatchNorm1d(nf), nn.ReLU(inplace=True))
def forward(self, x):
sh = x.shape
x = x.permute(0,2,3,1).contiguous().view(sh[0]*sh[2]*sh[3], sh[1])
x = self.chlin(x).view(sh[0],sh[2],sh[3], -1).permute(0,3,1,2).contiguous()
return x
class FireHR(Module):
def __init__(self, ni, nc):
self.conv = ConvLayer(1, 8)
self.chlin = nn.Sequential(ChLin(ni+8, 128), ChLin(128, 64))
self.middleconv = nn.Sequential(ConvLayer(64, 128), ConvLayer(128, 64))
self.finalconv = nn.Conv2d(64, nc, kernel_size=1, bias=True)
def forward(self, x):
x = torch.cat([x[:,:-1], self.conv(x[:,-1].unsqueeze(1))], dim=1)
x = self.chlin(x)
x = self.middleconv(x)
return self.finalconv(x)
# -
FireHR(6,1)
#export
def download_model_weights(weight_file='model512.pth'):
"""Download model weights if they don't exist yet on ~/.firehr."""
path_save = Path(os.path.expandvars('$HOME'))/'.firehr'
path_save.mkdir(exist_ok=True)
file_save = path_save/weight_file
if not file_save.is_file():
print(f'Downloading model weights {weight_file}.')
url = 'https://github.com/mnpinto/FireHR_weights/raw/main/model512.pth'
file = requests.get(url)
open(str(file_save), 'wb').write(file.content)
else:
print(f'Using local model weights {file_save}')
download_model_weights()
#export
_WEIGHTS = Path(os.path.expandvars('$HOME'))/'.firehr/model512.pth'
def load_pretrained_model(weights=_WEIGHTS, ni=6, nc=1, half_precision=True, gpu=True):
download_model_weights()
model = FireHR(ni,nc)
st = torch.load(weights, map_location=torch.device('cpu'))
model.load_state_dict(st['model'])
if gpu:
if half_precision: model = model.half()
if torch.cuda.is_available():
model = model.cuda()
else:
warnings.warn('GPU is not available. torch.cuda.is_available() returned False.')
return model
#local
model = load_pretrained_model()
model.conv[0].weight.device, model.conv[0].weight.type()
#hide
notebook2script()
| nbs/02_models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import requests
from src.data.download_DHS import GBD_API_KEY
url = 'fhttps://api.healthdata.org/healthdata/v1/data/gbd/cause/?cause_id=294&location_id=102&measure_id=1&metric_id=1&year_id=2016&authorization={GBD_API_KEY}'
# +
# I DON'T THINK COVARIATES WORK
# url = f'https://api.healthdata.org/healthdata/v1/data/gbd/covariate/?covariate_id=8&location_id=102&year_id=2016&authorization={GBD_API_KEY}'
# -
# Risk
# Attributable Risk
url = 'https://api.healthdata.org/healthdata/v1/data/gbd/risk/?risk_id=82&cause_id=294&location_id=102&measure_id=1&metric_id=1&year_id=2016&age_group_id=1&authorization={GBD_API_KEY}'
# SEV - Remember these must be most detailed risks!
url = 'https://api.healthdata.org/healthdata/v1/data/gbd/sev/?risk_id=82&measure_id=29&metric_id=3&year_id=2016&age_group_id=1&authorization={GBD_API_KEY}'
url = 'https://api.healthdata.org/healthdata/v1/data/gbd/sev/?risk_id=83&measure_id=29&location_id=102&age_group_id=5&sex_id=3&authorization={GBD_API_KEY}'
r = requests.get(url)
status = r.json()['meta']['status']
if status['code'] != '200':
raise AssertionError(f'Failed HealthData API query - {status["message"]}')
cols = r.json()['meta']['fields']
df = pd.DataFrame(r.json()['data'], columns=cols)
df.head()
| notebooks/_archive/2019_01_24_GBD_API_testing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="N2_J4Rw2r0SQ" outputId="add44b90-08ec-4f95-e873-a9b43366f34d"
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
from tqdm import tqdm
# %matplotlib inline
from torch.utils.data import Dataset, DataLoader
import torch
import torchvision
import torch.nn as nn
import torch.optim as optim
from torch.nn import functional as F
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print(device)
# + id="ZTJPyL45DiJ3"
m = 10 # 5, 50, 100, 500, 2000
# + id="g-do5dZWK5L6"
train_size = 10000 # 100, 500, 2000, 10000
# + [markdown] id="F6fjud_Fr0Sa"
# # Generate dataset
# + colab={"base_uri": "https://localhost:8080/"} id="CqdXHO0Cr0Sd" outputId="24cee382-ea88-41f4-c0f2-92a4acb23374"
np.random.seed(12)
y = np.random.randint(0,10,5000)
idx= []
for i in range(10):
print(i,sum(y==i))
idx.append(y==i)
# + id="ddhXyODwr0Sk"
x = np.zeros((5000,2))
# + id="DyV3N2DIr0Sp"
np.random.seed(12)
x[idx[0],:] = np.random.multivariate_normal(mean = [7,4],cov=[[0.1,0],[0,0.1]],size=sum(idx[0]))
x[idx[1],:] = np.random.multivariate_normal(mean = [8,6.5],cov=[[0.1,0],[0,0.1]],size=sum(idx[1]))
x[idx[2],:] = np.random.multivariate_normal(mean = [5.5,6.5],cov=[[0.1,0],[0,0.1]],size=sum(idx[2]))
x[idx[3],:] = np.random.multivariate_normal(mean = [-1,0],cov=[[0.1,0],[0,0.1]],size=sum(idx[3]))
x[idx[4],:] = np.random.multivariate_normal(mean = [0,2],cov=[[0.1,0],[0,0.1]],size=sum(idx[4]))
x[idx[5],:] = np.random.multivariate_normal(mean = [1,0],cov=[[0.1,0],[0,0.1]],size=sum(idx[5]))
x[idx[6],:] = np.random.multivariate_normal(mean = [0,-1],cov=[[0.1,0],[0,0.1]],size=sum(idx[6]))
x[idx[7],:] = np.random.multivariate_normal(mean = [0,0],cov=[[0.1,0],[0,0.1]],size=sum(idx[7]))
x[idx[8],:] = np.random.multivariate_normal(mean = [-0.5,-0.5],cov=[[0.1,0],[0,0.1]],size=sum(idx[8]))
x[idx[9],:] = np.random.multivariate_normal(mean = [0.4,0.2],cov=[[0.1,0],[0,0.1]],size=sum(idx[9]))
# + colab={"base_uri": "https://localhost:8080/"} id="qh1mDScsU07I" outputId="4605919d-f584-45f4-e894-7608076caa12"
x[idx[0]][0], x[idx[5]][5]
# + colab={"base_uri": "https://localhost:8080/", "height": 287} id="hJ8Jm7YUr0St" outputId="7da2978b-568d-478a-83aa-7e0747ef017b"
for i in range(10):
plt.scatter(x[idx[i],0],x[idx[i],1],label="class_"+str(i))
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
# + colab={"base_uri": "https://localhost:8080/"} id="3lMBZEHNBlF2" outputId="faae2e4a-78e2-4972-ed15-307ccaefc590"
bg_idx = [ np.where(idx[3] == True)[0],
np.where(idx[4] == True)[0],
np.where(idx[5] == True)[0],
np.where(idx[6] == True)[0],
np.where(idx[7] == True)[0],
np.where(idx[8] == True)[0],
np.where(idx[9] == True)[0]]
bg_idx = np.concatenate(bg_idx, axis = 0)
bg_idx.shape
# + colab={"base_uri": "https://localhost:8080/"} id="blRbGZHeCwXU" outputId="ba506046-5c00-47f9-f631-1af74570e44f"
np.unique(bg_idx).shape
# + id="Y43sWeX7C15F"
x = x - np.mean(x[bg_idx], axis = 0, keepdims = True)
# + colab={"base_uri": "https://localhost:8080/"} id="ooII7N6UDWe0" outputId="63e2fa23-fbb3-4232-f518-8782eeab12c9"
np.mean(x[bg_idx], axis = 0, keepdims = True), np.mean(x, axis = 0, keepdims = True)
# + id="g21bvPRYDL9k"
x = x/np.std(x[bg_idx], axis = 0, keepdims = True)
# + colab={"base_uri": "https://localhost:8080/"} id="GtFvIeHsDZJk" outputId="89497515-bbbe-4dfd-cec7-43f1efb01041"
np.std(x[bg_idx], axis = 0, keepdims = True), np.std(x, axis = 0, keepdims = True)
# + colab={"base_uri": "https://localhost:8080/", "height": 283} id="8-VLhUfDDeHt" outputId="873a8dc4-f6b9-47a7-bf14-5359ef3707ad"
for i in range(10):
plt.scatter(x[idx[i],0],x[idx[i],1],label="class_"+str(i))
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
# + id="UfFHcZJOr0Sz"
foreground_classes = {'class_0','class_1', 'class_2'}
background_classes = {'class_3','class_4', 'class_5', 'class_6','class_7', 'class_8', 'class_9'}
# + colab={"base_uri": "https://localhost:8080/"} id="OplNpNQVr0S2" outputId="9049b153-e669-4377-8093-f90051ed4a6e"
fg_class = np.random.randint(0,3)
fg_idx = np.random.randint(0,m)
train_data=[]
a = []
fg_instance = np.array([[0.0,0.0]])
bg_instance = np.array([[0.0,0.0]])
for i in range(m):
if i == fg_idx:
b = np.random.choice(np.where(idx[fg_class]==True)[0],size=1)
fg_instance += x[b]
a.append(x[b])
print("foreground "+str(fg_class)+" present at " + str(fg_idx))
else:
bg_class = np.random.randint(3,10)
b = np.random.choice(np.where(idx[bg_class]==True)[0],size=1)
bg_instance += x[b]
a.append(x[b])
print("background "+str(bg_class)+" present at " + str(i))
a = np.concatenate(a,axis=0)
print(a.shape)
print(fg_class , fg_idx)
# + colab={"base_uri": "https://localhost:8080/"} id="Rx6cMmR3baVf" outputId="20e1543a-4593-46f7-ec1a-cbd8ffe12a4a"
a
# + colab={"base_uri": "https://localhost:8080/"} id="dEdaFZPQhc7D" outputId="6055ab89-588d-40ef-9637-5e94dbbef430"
fg_instance
# + colab={"base_uri": "https://localhost:8080/"} id="IqdAVbG1hftM" outputId="b51e00d9-cd57-45fd-ff48-1589b1f06551"
bg_instance
# + colab={"base_uri": "https://localhost:8080/"} id="Ho98SSXEhx6L" outputId="31397ba8-2252-46ea-c466-cc4b7f4bf74a"
(fg_instance+bg_instance)/m , m
# + id="jqbvfbwVr0TN"
# mosaic_list_of_images =[]
# mosaic_label = []
train_label=[]
fore_idx=[]
train_data = []
for j in range(train_size):
np.random.seed(j)
fg_instance = torch.zeros([2], dtype=torch.float64) #np.array([[0.0,0.0]])
bg_instance = torch.zeros([2], dtype=torch.float64) #np.array([[0.0,0.0]])
# a=[]
for i in range(m):
if i == fg_idx:
fg_class = np.random.randint(0,3)
b = np.random.choice(np.where(idx[fg_class]==True)[0],size=1)
fg_instance += x[b]
# a.append(x[b])
# print("foreground "+str(fg_class)+" present at " + str(fg_idx))
else:
bg_class = np.random.randint(3,10)
b = np.random.choice(np.where(idx[bg_class]==True)[0],size=1)
bg_instance += x[b]
# a.append(x[b])
# print("background "+str(bg_class)+" present at " + str(i))
train_data.append((fg_instance+bg_instance)/m)
# a = np.concatenate(a,axis=0)
# mosaic_list_of_images.append(np.reshape(a,(2*m,1)))
train_label.append(fg_class)
fore_idx.append(fg_idx)
# + colab={"base_uri": "https://localhost:8080/"} id="X_g6lxkVjN0N" outputId="e5adf0d7-e063-4495-8586-602fdcbd03f1"
train_data[0], train_label[0]
# + colab={"base_uri": "https://localhost:8080/"} id="Tz5wQCevobYr" outputId="b3d198a9-4838-486f-8a0d-aba4fd68dc63"
train_data = torch.stack(train_data, axis=0)
train_data.shape, len(train_label)
# + id="CVAlImGYA49A"
test_label=[]
# fore_idx=[]
test_data = []
for j in range(1000):
np.random.seed(j)
fg_instance = torch.zeros([2], dtype=torch.float64) #np.array([[0.0,0.0]])
fg_class = np.random.randint(0,3)
b = np.random.choice(np.where(idx[fg_class]==True)[0],size=1)
fg_instance += x[b]
# a.append(x[b])
# print("foreground "+str(fg_class)+" present at " + str(fg_idx))
test_data.append((fg_instance)/m)
# a = np.concatenate(a,axis=0)
# mosaic_list_of_images.append(np.reshape(a,(2*m,1)))
test_label.append(fg_class)
# fore_idx.append(fg_idx)
# + colab={"base_uri": "https://localhost:8080/"} id="XltBz3__BT_C" outputId="ab293c6a-8ead-4381-f267-0783b6519427"
test_data[0], test_label[0]
# + colab={"base_uri": "https://localhost:8080/"} id="JqQfjkCnBX8E" outputId="1fd7a08c-dd64-4213-feeb-41bae11f623e"
test_data = torch.stack(test_data, axis=0)
test_data.shape, len(test_label)
# + id="nQhRf1ICBvW6"
x1 = (train_data).numpy()
y1 = np.array(train_label)
# + colab={"base_uri": "https://localhost:8080/"} id="vDAz96DSCXn5" outputId="d5c0ce62-0e72-4974-cb3f-268eb6dcf089"
x1[y1==0,0]
# + colab={"base_uri": "https://localhost:8080/"} id="kMwwZhv7Bv1U" outputId="cf6ee831-31fa-4035-dfca-bcd21b334103"
x1[y1==0,0][:,0]
# + colab={"base_uri": "https://localhost:8080/"} id="ydbu3OV3CYxB" outputId="acface5d-ea77-479f-e47e-1ec6653304b5"
x1[y1==0,0][:,1]
# + colab={"base_uri": "https://localhost:8080/", "height": 299} id="bT9-kEI7NAnR" outputId="93150a30-9d71-44e0-f117-fcb4093f688d"
x1 = (train_data).numpy()
y1 = np.array(train_label)
plt.scatter(x1[y1==0,0][:,0], x1[y1==0,0][:,1], label='class 0')
plt.scatter(x1[y1==1,0][:,0], x1[y1==1,0][:,1], label='class 1')
plt.scatter(x1[y1==2,0][:,0], x1[y1==2,0][:,1], label='class 2')
plt.legend()
plt.title("dataset4 CIN with alpha = 1/"+str(m))
# + colab={"base_uri": "https://localhost:8080/", "height": 299} id="Sc8W2N7LQQ_l" outputId="9c95738e-77d9-4ed0-d574-972608079e1a"
x1 = (test_data).numpy()
y1 = np.array(test_label)
plt.scatter(x1[y1==0,0][:,0], x1[y1==0,0][:,1], label='class 0')
plt.scatter(x1[y1==1,0][:,0], x1[y1==1,0][:,1], label='class 1')
plt.scatter(x1[y1==2,0][:,0], x1[y1==2,0][:,1], label='class 2')
plt.legend()
plt.title("test dataset4")
# + id="yL0BRf8er0TX"
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list_of_images, mosaic_label):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list_of_images
self.label = mosaic_label
#self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx] #, self.fore_idx[idx]
# + colab={"base_uri": "https://localhost:8080/"} id="4KsrW9qL9xgS" outputId="746924be-ede3-42d5-83ae-8247440c60f4"
train_data[0].shape, train_data[0]
# + id="EY2l62APygaV"
batch = 200
traindata_1 = MosaicDataset(train_data, train_label )
trainloader_1 = DataLoader( traindata_1 , batch_size= batch ,shuffle=True)
# + id="9suUslCj8YZK"
testdata_1 = MosaicDataset(test_data, test_label )
testloader_1 = DataLoader( testdata_1 , batch_size= batch ,shuffle=False)
# + id="Nh3mBQHZ8bEj"
# testdata_11 = MosaicDataset(test_dataset, labels )
# testloader_11 = DataLoader( testdata_11 , batch_size= batch ,shuffle=False)
# + id="5_XeIUk0r0Tl"
class Whatnet(nn.Module):
def __init__(self):
super(Whatnet,self).__init__()
self.linear1 = nn.Linear(2,50)
self.linear2 = nn.Linear(50,3)
torch.nn.init.xavier_normal_(self.linear1.weight)
torch.nn.init.zeros_(self.linear1.bias)
torch.nn.init.xavier_normal_(self.linear2.weight)
torch.nn.init.zeros_(self.linear2.bias)
def forward(self,x):
x = F.relu(self.linear1(x))
x = (self.linear2(x))
return x[:,0]
# + id="pjD2VZuV9Ed4"
def calculate_loss(dataloader,model,criter):
model.eval()
r_loss = 0
with torch.no_grad():
for i, data in enumerate(dataloader, 0):
inputs, labels = data
inputs, labels = inputs.to("cuda"),labels.to("cuda")
outputs = model(inputs)
# print(outputs.shape)
loss = criter(outputs, labels)
r_loss += loss.item()
return r_loss/(i+1)
# + id="uALi25pmzQHV"
def test_all(number, testloader,net):
correct = 0
total = 0
out = []
pred = []
with torch.no_grad():
for data in testloader:
images, labels = data
images, labels = images.to("cuda"),labels.to("cuda")
out.append(labels.cpu().numpy())
outputs= net(images)
_, predicted = torch.max(outputs.data, 1)
pred.append(predicted.cpu().numpy())
total += labels.size(0)
correct += (predicted == labels).sum().item()
pred = np.concatenate(pred, axis = 0)
out = np.concatenate(out, axis = 0)
print("unique out: ", np.unique(out), "unique pred: ", np.unique(pred) )
print("correct: ", correct, "total ", total)
print('Accuracy of the network on the %d test dataset %d: %.2f %%' % (total, number , 100 * correct / total))
# + id="4vmNprlPzTjP"
def train_all(trainloader, ds_number, testloader_list, lr_list):
final_loss = []
for LR in lr_list:
print("--"*20, "Learning Rate used is", LR)
torch.manual_seed(12)
net = Whatnet().double()
net = net.to("cuda")
criterion_net = nn.CrossEntropyLoss()
optimizer_net = optim.Adam(net.parameters(), lr=0.001 ) #, momentum=0.9)
acti = []
loss_curi = []
epochs = 1000
running_loss = calculate_loss(trainloader,net,criterion_net)
loss_curi.append(running_loss)
print('epoch: [%d ] loss: %.3f' %(0,running_loss))
for epoch in range(epochs): # loop over the dataset multiple times
ep_lossi = []
running_loss = 0.0
net.train()
for i, data in enumerate(trainloader, 0):
# get the inputs
inputs, labels = data
inputs, labels = inputs.to("cuda"),labels.to("cuda")
# zero the parameter gradients
optimizer_net.zero_grad()
# forward + backward + optimize
outputs = net(inputs)
# print(outputs.shape)
loss = criterion_net(outputs, labels)
# print statistics
running_loss += loss.item()
loss.backward()
optimizer_net.step()
running_loss = calculate_loss(trainloader,net,criterion_net)
if(epoch%200 == 0):
print('epoch: [%d] loss: %.3f' %(epoch + 1,running_loss))
loss_curi.append(running_loss) #loss per epoch
if running_loss<=0.05:
print('epoch: [%d] loss: %.3f' %(epoch + 1,running_loss))
break
print('Finished Training')
correct = 0
total = 0
with torch.no_grad():
for data in trainloader:
images, labels = data
images, labels = images.to("cuda"), labels.to("cuda")
outputs = net(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the %d train images: %.2f %%' % (total, 100 * correct / total))
for i, j in enumerate(testloader_list):
test_all(i+1, j,net)
print("--"*40)
final_loss.append(loss_curi)
return final_loss
# + id="Yl41sE8vFERk"
train_loss_all=[]
testloader_list= [ testloader_1]
lr_list = [0.0001, 0.0005, 0.001, 0.005, 0.01, 0.05, 0.1, 0.5 ]
# + colab={"base_uri": "https://localhost:8080/"} id="5gQoPST5zW2t" outputId="e61908e0-4cb1-41cc-f2e2-b5ba9c203de9"
fin_loss = train_all(trainloader_1, 1, testloader_list, lr_list)
train_loss_all.append(fin_loss)
# + id="In76SYH_zZHV"
# %matplotlib inline
# + id="_GsE3qHR_A5c"
len(fin_loss)
# + id="BS4HtOHEzZ0E"
for i,j in enumerate(fin_loss):
plt.plot(j,label ="LR = "+str(lr_list[i]))
plt.xlabel("Epochs")
plt.ylabel("Training_loss")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
# + id="1UbTkfLUINTI"
| AAAI/Learnability/CIN/MLP/ds3/synthetic_type3_MLP_size_10000_m_10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TIE Reconstruction Template
# In this notebook we demonstrate solving the transport of intensity equation (TIE) and finding magnetic induction from an experimental through focal series (tfs) of TEM images. For more information please check out our [wiki/documentation pages](https://pylorentztem.readthedocs.io/en/latest/).
#
# If you have never used a Jupyter Notebook before we recommend familiarizing yourself with the [notebook basics](https://jupyter-notebook.readthedocs.io/en/stable/examples/Notebook/Notebook%20Basics.html). While Python knowledge is not strictly required for running PyLorentz code, a certain familiarity is helpful for understanding how to modify the examples to suit your needs.
#
# This notebook is ready to be run as-is. The first time, however, you will need to download the example data. This can be done through the notebook and is demonstrated below. Cells can be executed by selecting and pressing ``shift + enter``, and you can proceed with running all of the cells in order. The python docstrings for these functions can be found on the wiki, but are also viewable in Jupyter. E.g. for the TIE() function, executing a cell containing: `?TIE` will pull up the documentation for that function.
#
# All of the features available in this notebook are also available in the TIE GUI, which additionally contains processes for aligning experimental datasets. For details on running that please refer to the README in the ``Pylorentz/GUI/`` directory.
#
# Authors: <NAME>, <NAME>
# V1.0, ANL, July 2020
# ### Importing the necessary modules
# +
import sys
import numpy as np
import matplotlib.pyplot as plt
sys.path.append("../PyTIE")
from microscopes import Microscope
from TIE_helper import *
from TIE_reconstruct import TIE, SITIE
# %matplotlib notebook
# -
# ## Downloading the data
# This first section shows how to download example data from the [Materials Data Facility](https://doi.org/10.18126/z9tc-i8bf). This experimental dataset contains a full through focus series (tfs) of an artificial spin ice structure with the sample in both flipped and unflipped orientations.
#
# The files will be placed in `Examples/cphatak_pytie_example_v1.1/`, and you only need to run this following cell the first time through the notebook. Incidentally there seems to be a bug in the mdf-forge package and the progress bar doesn't move, but it generally takes ~5-10 minutes for the download depending on your internet speed.
#
# If you are having trouble downloading the data please make sure
# * You have a Globus endpoint on your computer
# * You have Globus Connect running
#
# If it still isn't working you can download the data with globus from the [data page](https://doi.org/10.18126/z9tc-i8bf). Click "Globus" under "Get the Data" and download using their online app.
# +
from mdf_forge import Forge
mdf = Forge()
# Match the dataset by title and only collect "dataset" resource types
res = mdf.match_resource_types('dataset').match_titles('pytie').search()
# Download the data using Globus
mdf.globus_download(res, download_datasets=True)
# -
# ## Loading the data into PyTIE
# Now that we have our data on the computer, we need to load it into PyTIE with the `load_data()` function. We set a few parameters here:
#
# - path (str): We specify the filepath as the one created with the example data.
# - fls_file (str): The name of the .fls file, which itself contains the image filenames and defocus values.
# - flip_fls_file (str): We explicitly show that there isn't a flip fls file, though this could be omitted as it's the default value.
# - al_file (str): The name of the aligned image file.
# - flip (bool): Whether or not there is a flip tfs along with the unflip.
# +
data_loc = './cphatak_pytie_example_v1.1/'
fls_name = "dataset1.fls"
flip_fls_name = None # just one fls file
stack_name = 'd1_full_align.tif'
flip = True # there is/isn't a flipped tfs and flipstack
imstack, flipstack, ptie = load_data(path=data_loc,
fls_file=fls_name,
flip_fls_file=flip_fls_name,
al_file=stack_name,
flip=flip)
print("Defocus values: ", ptie.defvals, "nm")
# create microscope object:
#Only relevant parameter for TIE reconstruction is accelerating voltage.
pscope = Microscope(E=200e3)
# -
# Now view the masked infocus image.
#
# + code_folding=[]
# Scroll through and view the aligned images
show_stack(imstack + flipstack)
# -
show_im(ptie.infocus, title='Masked infocus image', scale=ptie.scale)
# ## Select a smaller image size (optional)
# There may be times when you would want to only reconstruct part of an image, e.g. because a window edge introduces a high-contrast edge that interferes with the reconstruction. Although this is available here in the Jupyter notebook, it is somewhat finnicky. We recommend using the GUI if you want to be precise with region selection, as it also allows for rotating the images.
#
# The select_region() method of the TIE_params class opens an interface to select a region, and running the crop_ims() method saves it. So you can change the rectangular region after running crop_ims() but the changes will not take effect until it is run again.
#
# * If you choose a non-square subregion, the results are no longer quantitative. Qualitatively they will appear correct except beyond aspect ratios of ~3:1, where Fourier edge effects become more prominent. These can be reduced by symmetrizing the image.
# * Default is 1/4 image size centered in the middle
# * Drag mask and corners to get the section you want to reconstruct
# - It's finnicky, if the corner isn't dragging, try moving mouse cursor more slowly and make sure you're on the corner.
# - if it still isn't working you can change ptie.crop by hand which sets the boundary (in pixels) for which to crop.
#
#
ptie.select_region()
### Run this when the region is what you want.
ptie.crop_ims()
print(f"Ptie.crop: {ptie.crop}")
show_stack(ptie.imstack+ptie.flipstack, ptie)
# ## The actual phase reconstruction
# Here we perform the TIE reconstruction. The following parameters are set:
#
# * Set which focus values you want to use with index i
# * images will be saved as /dataloc/images/dataname_focusval_imagename.tif
# * if you want to symmetrize the images set sym = True
# * if you want to run with a Tikhonov frequency
# - set qc = value that you want, (currently needs large values ~>10) or
# - set qc = 'percent' and it will use 15% of q (not that it makes much difference) or
# - leave qc = None and it will run normally
# * Save options:
# - True -- saves all images results directory
# - 'b' -- saves 'color_b', 'bxt', 'byt'
# - 'color' -- saves just 'color_b'
# - False -- Does not save images, but they can still be viewed in the notebook
# - If any images are saved, a dataname_params.txt file will be saved as well giving reconstruction parameters.
print('actual defocus values: ', ptie.defvals)
print('reconstruct with i = ', ' 0 ', ' 1 ', ' 2 ...')
i = 2 # Select which defocus value you'll be working with if 3-point deriv
dataname = 'example_TIE'# str
sym = False # bool
qc = False # 0.001 is a reasonable value for the test dataset, though it has an overall small effect here.
save = 'b' # str or bool
print(f'Set to reconstruct for defocus value: {ptie.defvals[i]} nm ')
results = TIE(i, ptie, pscope,
dataname = dataname,
sym=sym,
qc = qc,
save=save)
# ## Viewing the images
# You can of course just look at the data where it's saved, but if you don't want to save it and move it to a local computer you can view the images here, as they're all in the results dictionary.
# * Results:
# - 'byt' : y-component of integrated magnetic induction,
# - 'bxt' : x-component of integrated magnetic induction,
# - 'bbt' : magnitude of integrated magnetic induction,
# - 'phase_m' : magnetic phase shift (radians),
# - 'phase_e' : electrostatic phase shift (if using flip stack) (radians),
# - 'dIdZ_m' : intensity derivative for calculating phase_m, (useful for troubleshooting)
# - 'dIdZ_e' : intensity derivative for calculating phase_e (if using flip stack),
# - 'color_b' : RGB image of magnetization,
# - 'inf_im' : the in-focus image
show_im(results['phase_m'], title='magnetic phase shift', scale=ptie.scale,
cbar_title="Radians")
show_im(results['phase_e'], 'electrostatic phase shift',scale=ptie.scale,
cbar_title="Radians")
# Show the image with a four-fold colorwheel instead of a 3-fold hsv colormap
from colorwheel import color_im
show_im(color_im(results['bxt'], results['byt'], hsvwheel=False), "Magnetization with four-fold colormap", simple=True)
# Show a vector plot of the magnetization over the colormap image. The arrows size is fixed so you will have to zoom in to see them. There are few parameters required to have the vectormap look good.
# - a (int): Number of arrows to show in the x and y directions
# - l (float): Scale factor of arrows. Larger l -> shorter arrows.
# - w (float): Width scaling of arrows.
# - title (str): (*optional*) Title for plot. Default None.
# - color (bool): (*optional*) Whether or not to show a colormap underneath the arrow plot.
# - hsv (bool): (*optional*) Only relevant if color == True. Whether to use an hsv or 4-fold colorwheel in the color image.
# - origin (str): (*optional*) Control image orientation.
# - save (str): (*optional*) Path to save the figure.
show_2D(results['bxt'], results['byt'], a=100, l=1, w=1, title='Arrow overlay image', color=True, hsv=True,
origin='upper', save = ptie.data_loc+'images/arrow_colormap.png')
# ### Single Image Reconstruction (SITIE)
# Uniformly thin magnetic samples can be reconstructed from a single defocused image if their only source of contrast is magnetic Fresnel contrast. As this sample is an island structure that is not the case, and we require two TFS to reconstruct the phase. For an example of SITIE on simulated data please refer to SIM_template.ipynb
# --- end notebook ---
| Examples/TIE_template.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Description
#
# This notebook is used to request computation of Water Productivity for customized season period, crop multiplier and area from WaPOR's Gross Biomass Water Productivity (GBWP) or Net Biomass Water Productivity (NBWP) layers using WaPOR API.
#
# You will need WaPOR API Token to use this notebook (See README.md)
# + [markdown] colab_type="text" id="SeLbBDiMTh2X"
# # Step 1: Read APIToken
#
# Get your APItoken from https://wapor.apps.fao.org/profile. Enter your API Token when running the cell below.
# + colab={} colab_type="code" id="-TPJcW3sNzCH"
import requests
import json
path_query=r'https://io.apps.fao.org/gismgr/api/v1/query/'
path_sign_in=r'https://io.apps.fao.org/gismgr/api/v1/iam/sign-in/'
APIToken=input('Your API token: ')
# + [markdown] colab_type="text" id="LgJQvG49TmV6"
# # Step 2: Get Authorization AccessToken
#
# Using the input API token to get AccessToken for authorization
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="QWZsCiGGQZOM" outputId="fecb35f9-1dc9-433c-cb77-ae2d71a6b839"
resp_signin=requests.post(path_sign_in,headers={'X-GISMGR-API-KEY':APIToken})
resp_signin = resp_signin.json()
AccessToken=resp_signin['response']['accessToken']
AccessToken
# + [markdown] colab_type="text" id="Ux4ehfFEX2_o"
# # Step 3: Write Query Payload
#
# **Arguments** 'waterProductivity' type, 'startSeason', 'endSeason', 'resolution','shape'
#
# Read [Explanatory notes on Custom water productvity](./data/explanatory-notes.pdf) to know how to calculate cropMultiplier
# -
waterProductivity="GBWP" #"GBWP" or "NBWP"
cropMultiplier=0.5
startSeason="2015-01" # "YYYY-DK" (Dekad)
endSeason= "2015-18" # "YYYY-DK" (Dekad)
resolution="100m" #"250m" or "100m" , maybe "30m" works for some area
# Define Polygon shape of the area of interest
bbox= [37.95883206252312, 7.89534, 43.32093, 12.3873979377346] #latlon
xmin,ymin,xmax,ymax=bbox[0],bbox[1],bbox[2],bbox[3]
Polygon=[
[xmin,ymin],
[xmin,ymax],
[xmax,ymax],
[xmax,ymin],
[xmin,ymin]
]
# + colab={"base_uri": "https://localhost:8080/", "height": 504} colab_type="code" id="nElLc1RKPYPy" outputId="61b4b7c2-a12c-4094-ae4b-7bc4a9331582"
query ={
"type": "CustomWaterProductivity",
"params": {
"waterProductivity": waterProductivity, #"GBWP" or "NBWP"
"resolution": resolution, #"250m" or "100m" , maybe "30m" works for some area
"startSeason": startSeason, # "YYYY-DK" (Dekad)
"endSeason": endSeason, # "YYYY-DK" (Dekad),
"cropMultiplier":cropMultiplier,
"shape": {
"type": "Polygon", #define coordinates of the area in geojson format
"coordinates": [Polygon]
}
}
}
# -
# # Step 4: Post the QueryPayload with AccessToken in Header
#
# In responses, get an url to query job.
# +
resp_query=requests.post(path_query,headers={'Authorization':'Bearer {0}'.format(AccessToken)},
json=query)
resp_query = resp_query.json()
job_url=resp_query['response']['links'][0]['href']
job_url
# + [markdown] colab_type="text" id="w9HOBa5lTyWc"
# # Step 5: Get Job Results.
#
# It will take some time for the job to be finished. When the job is finished, its status will be changed from 'RUNNING' to 'COMPLETED' or 'COMPLETED WITH ERRORS'. If it is COMPLETED, the url to download custom water productivity map can be achieved from Response 'output'.
# + colab={"base_uri": "https://localhost:8080/", "height": 504} colab_type="code" id="bBNIuHKmRdcm" outputId="2f41f6d3-00bf-457e-c656-449fa0b73eac"
i=0
print('RUNNING',end=" ")
while i==0:
resp = requests.get(job_url)
resp=resp.json()
if resp['response']['status']=='RUNNING':
print('.',end =" ")
if resp['response']['status']=='COMPLETED':
results=resp['response']['output']
print('Link to download GBWP',results['bwpDownloadUrl'])
print('Link to download TBP',results['tbpDownloadUrl'])
print('Link to download AETI',results['wtrDownloadUrl'])
i=1
if resp['response']['status']=='COMPLETED WITH ERRORS':
print(resp['response']['log'])
i=1
| notebooks/Module1_unit5/6_CustomWaterProductivity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import math as math
import os
exec(open("../header.py").read())
# -
threshold = 40
raw_train_data = pd.read_csv(processed_root("03-bag-of-words/threshold-"+str(threshold)+"/bow_train_data.csv"))
raw_val_data = pd.read_csv(processed_root("03-bag-of-words/threshold-"+str(threshold)+"/bow_val_data.csv"))
raw_test_data = pd.read_csv(processed_root("03-bag-of-words/threshold-"+str(threshold)+"/bow_test_data.csv"))
raw_train_data.columns
raw_val_data.shape
raw_val_data.poetry_author.value_counts()
# ### Import
X_train = raw_train_data\
.drop(['poetry_text', 'poetry_author'], axis = 1)
X_val = raw_val_data\
.drop(['poetry_text', 'poetry_author'], axis = 1)
y_train = raw_train_data['poetry_author']
y_val = raw_val_data['poetry_author']
# ### Model
from sklearn.neural_network import MLPClassifier
mlp = MLPClassifier(hidden_layer_sizes = (50,10,), activation='relu')
# ### Train
y_pred = mlp.fit(X_train, y_train)
# ### Evaluate
# +
# Training
y_pred_train = mlp.predict(X_train)
train_accuracy = np.mean(y_pred_train == y_train) * 100
print("Training Accuracy: %.2f%%"%(train_accuracy))
# Validation
y_pred_val = mlp.predict(X_val)
val_accuracy = np.mean(y_pred_val == y_val) * 100
print("Validation Accuracy: %.2f%%"%(val_accuracy))
# -
# # Accuracy by author
# +
acc = pd.DataFrame({'pred':y_pred_val,
'true':y_val})
acc_by_author = acc\
.assign(correct = lambda x:x.pred == x.true)\
.assign(count_one = 1)\
.groupby('true')\
.agg({'correct':'mean',
'count_one':'sum'})\
.rename({'count_one':'total'}, axis = 1)
acc_by_author
# -
# ## Tuning process
def tuning_loop(hl_choices):
train_accuracies = []
val_accuracies = []
for hl in hl_choices:
model = MLPClassifier(hidden_layer_sizes = hl, activation='relu')
model.fit(X_train, y_train)
# Training
y_pred_train = model.predict(X_train)
train_accuracy = np.mean(y_pred_train == y_train) * 100
train_accuracies.append(train_accuracy)
# Validation
y_pred_val = model.predict(X_val)
val_accuracy = np.mean(y_pred_val == y_val) * 100
val_accuracies.append(val_accuracy)
return train_accuracies, val_accuracies
def plot_results(tune_var, train, val):
fig, ax = plt.subplots()
ax.plot(tune_var, train, label = "Train")
ax.plot(tune_var, val, label = "Validation")
ax.set_title("Multi-layer Perceptron Accuracy versus Max Depth")
ax.set_xlabel("Hidden Layer Size")
ax.set_ylabel("Accuracy (%)")
ax.legend()
print("Maximum validation accuracy, %.2f%% found a hidden layer size of %.2f."\
%(max(val), tune_var[np.argmax(val)]))
# ## Tune 1
hl_choices1 = [5,20,50,100]
t1, v1 = tuning_loop(hl_choices1)
plot_results(hl_choices1, t1, v1)
| code/03-notebooks/04-mlp.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.4.0
# language: julia
# name: julia-1.4
# ---
# *This notebook provides a comparison of several models for water waves on the results of an experiment of
# [Hammack\&Segur](https://dx.doi.org/10.1017/S002211207800020) experiment, in the spirit of [Carter](https://dx.doi.org/10.1016/j.wavemoti.2018.07.004)).*
# ## Initialization
# Import package
Pkg.activate("..")
using WaterWaves1D
#include("../src/dependencies.jl");
include("../src/models/WaterWaves.jl"); #include the model which will be used
include("../src/models/WWn.jl");
include("../src/models/SerreGreenNaghdi.jl");
include("../src/models/WhithamGreenNaghdi.jl");
# Define parameters of the problem
d = 0.1; # depth of the basin (in m)
g = 9.81; # gravitational acceleration
λ = 0.1; # typical horizontal length (=d to simplify)
T = 40; # final time (in s)
L = 156.16;# half-length of the numerical tank (in m)
param = (
# Physical parameters. Variables are non-dimensionalized as in Lannes, The water waves problem, isbn:978-0-8218-9470-5
μ = 1, # shallow-water dimensionless parameter
ϵ = 1, # nonlinearity dimensionless parameter
# Numerical parameters
N = 2^12, # number of collocation points
L = L/d, # half-length of the numerical tank ( = 156.16 m)
T = T*sqrt(g*d)/λ, # final time of computation (= 50 s)
dt = T*sqrt(g*d)/λ/10^3, # timestep
)
# Define initial data (as in [Carter](https://dx.doi.org/10.1016/j.wavemoti.2018.07.004)).
#
# +
using Elliptic,Plots
a = 0.005 # half-amplitude of the initial data (in m)
sn(x) = getindex.(ellipj.(9.25434*x*d,0.9999^2),1); # regularized step function
η0(x) = (-1/2*a.+1/2*a*sn(x)).*(x*d.<.61).*(x*d.>-1.83)/d;
init = Init(x->2*η0(x),x->zero(x)); # generate the initial data with correct type
plot(x,[init.η(Mesh(param).x)*d init.v(x)],
title="initial data",
label=["surface deformation" "velocity"],
xlims=(-20,10)
)
# -
# Set up initial-value problems for different models to compare
model_WW2=WWn(param;n=2,dealias=1,δ=1/10,verbose=false) # The quadratic water waves model (WW2)
model_SGN=SerreGreenNaghdi(param;verbose=false) # The Serre-Green-Naghdi model (SGN)
model_WGN=WhithamGreenNaghdi(param;verbose=false) # The fully dispersive Whitham-Green-Naghdi model (WGN)
# type `?WaterWaves` or `?WWn`, etc. to see details and signification of arguments
WW2=Problem(model_WW2, init, param) ;
SGN=Problem(model_SGN, init, param) ;
WGN=Problem(model_WGN, init, param) ;
# ## Computation
# Solve integration in time
solve!(WW);solve!(WW2);solve!(SGN);solve!(WGN);
# ## Visualization
# Import the data (from the [Hammack\&Segur](https://dx.doi.org/10.1017/S0022112078000208) experiment, kindly provided by [Carter](https://dx.doi.org/10.1016/j.wavemoti.2018.07.004))
using DelimitedFiles
data2a = readdlm("./Hammack-Segur/Fig2aFixed.out",'\t');
data2b = readdlm("./Hammack-Segur/Fig2bFixed.out",'\t');
data2c = readdlm("./Hammack-Segur/Fig2cFixed.out",'\t');
data2d = readdlm("./Hammack-Segur/Fig2dFixed.out",'\t');
data2e = readdlm("./Hammack-Segur/Fig2eFixed.out",'\t');
"""
gauge(p::Problem;x,t)
Give the surface elevation provided by a problem
at a specific location `x` (by default, `x=0`),
and at times defined by `t` if provided (otherwise at all computed times).
"""
function gauge(p::Problem;x=0,t=nothing)
if t==nothing
times=p.times.ts
elseif t[1]==t
times=[t]
else
times=t
end
times/sqrt(g*d)*λ,[solution(p,t=ti,x=[x])[1][1] for ti in times]*d
end
# Plot data and numerics at first gauge
plt=plot(data2a[:,1]/sqrt(g*d)*λ,data2a[:,2]*d*2/3,
title="first gauge",
label="experiment",
xlabel="time (in s)",
ylabel="surface deformation (in m)")
plot!([gauge(WW2,x=1),gauge(SGN,x=1),gauge(WGN,x=1)],
label=["WW2" "SGN" "WGN"])
xlims!(0,5)
# Plot data and numerics at second gauge
plt=plot((data2b[:,1].+50)/sqrt(g*d)*λ,data2b[:,2]*d*2/3,
title="second gauge",
label="experiment",
xlabel="time (in s)",
ylabel="surface deformation (in m)")
plot!([gauge(WW2,x=51.),gauge(SGN,x=51.),gauge(WGN,x=51.)],
label=["WW2" "SGN" "WGN"])
xlims!(5,17)
# Plot data and numerics at third gauge
plt=plot((data2c[:,1].+100)/sqrt(g*d)*λ,data2c[:,2]*d*2/3,
title="third gauge",
label="experiment",
xlabel="time (in s)",
ylabel="surface deformation (in m)")
plot!([gauge(WW2,x=101.),gauge(SGN,x=101.),gauge(WGN,x=101.)],
label=["WW2" "SGN" "WGN"])
xlims!(10,30)
# Plot data and numerics at fourth gauge
plt=plot((data2d[:,1].+150)/sqrt(g*d)*λ,data2d[:,2]*d*2/3,
title="fourth gauge",
label="experiment",
xlabel="time",
ylabel="surface deformation")
plot!([gauge(WW2,x=151.),gauge(SGN,x=151.),gauge(WGN,x=151.)],
label=["WW2" "SGN" "WGN"])
xlims!(15,35)
# Plot data and numerics at fifth gauge
plt=plot((data2e[:,1].+200)/sqrt(g*d)*λ,data2e[:,2]*d*2/3,
title="fifth gauge",
label="experiment",
xlabel="time",
ylabel="surface deformation")
plot!([gauge(WW2,x=201.),gauge(SGN,x=201.),gauge(WGN,x=201.)],
label=["WW2" "SGN" "WGN"])
xlims!(20,40)
# 50[1]==50
plot(x,solution(WW2))
plot!(x, solution(WW2).-solution(WGN))
| notebooks/Hammack-Segur/.ipynb_checkpoints/Hammack-Segur-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.3
# language: julia
# name: julia-1.6
# ---
# +
using Pkg
Pkg.activate(".")
using PlotlyJS
layout = Layout(;
title = "",
width = 600,
height = 400,
xaxis = attr(title = "Date"),
yaxis = attr(title = "Requests Count"),
)
# -
PlotElement = GenericTrace{Dict{Symbol, Any}}
PlotArray(;n=9) = Vector{PlotElement}(undef,0)
data = PlotArray()
append!(data, [scatter(x=[0,1],y=[0,1]), scatter(x=[1,0],y=[0,1])])
plot(data,layout)
data = [pie(labels=["a","b"],values=[20,30])]
plot(data)
d = Dict("a"=>1,"b"=>2,"c"=>3)
[d[x] for x in ["a","b"]]
typeof("") === String
plot(pie(values=[1,2,3]))
d = [Dict("a"=>1)]
println(d)
| Testing.ipynb |