code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Scraping Tweets using Python
# +
import pandas as pd
import tweepy
tweepy.__version__
# -
# ## Authentication
consumer_key = ''
consumer_secret = ''
access_token = ''
access_token_secret = ''
# +
auth = tweepy.OAuthHandler(consumer_key, consumer_secret)
auth.set_access_token(access_token, access_token_secret)
# Access Twitter API
api = tweepy.API(auth, wait_on_rate_limit=True)
api
# -
# ## Get tweets from user timeline
tweets_user = api.user_timeline('narendramodi', count=100)
len(tweets_user)
type(tweets_user)
# +
tweets = pd.DataFrame(columns=['id', 'created_at', 'likes',
'retweets', 'text', 'source'])
for tweet in tweets_user:
#print(tweet.id, tweet.created_at, tweet.favorite_count, tweet.retweet_count,
# tweet.text, tweet.source)
curr_tweet = {'id': tweet.id,
'created_at': tweet.created_at,
'likes': tweet.favorite_count,
'retweets': tweet.retweet_count,
'text': tweet.text,
'source': tweet.source}
tweets = tweets.append(curr_tweet, ignore_index=True)
tweets['created_at'] = tweets['created_at'].astype(str)
tweets['created_at'] = pd.to_datetime(tweets['created_at'],
format='%Y-%m-%d %H:%M:%S')
tweets.to_csv('modi_tweets.csv', index=False)
# -
tweets.head()
tweets['created_at'] = tweets['created_at'].astype(str)
tweets['created_at'].head()
tweets['created_at'] = pd.to_datetime(tweets['created_at'],
format='%Y-%m-%d %H:%M:%S')
tweets['created_at'].head()
tweets['created_at'].min(), tweets['created_at'].max()
tweets.index = tweets['created_at']
#tweets.head()
# %matplotlib inline
tweets.resample('1D')['id'].count().plot.line()
tweets.to_csv('modi_tweets.csv', index=False)
os.getcwd()
# ### Text Analysis
# 
# ## Search tweets from a hashtag
tweets_datascience = api.search('#olacabs', count=500)
len(tweets_datascience)
# ## Convert tweets to data frame
tweets_df = pd.DataFrame(columns=['text', 'retweets', 'likes', 'user', 'screen_name'])
for tweet in list(tweets_datascience):
tweet_series = pd.Series({'text': tweet.text, 'date': tweet.created_at,
'retweets': tweet.retweet_count,
'likes': tweet.favorite_count,
'user': tweet.user.name,
'screen_name': tweet.user.screen_name})
tweets_df = tweets_df.append(tweet_series, ignore_index=True)
tweets_df.head()
| Notebooks/Day 1/Scraping_Tweets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Line2D
# API
#
# matplotlib.lines.Line2D
#
# 在matplotlib中line这个名,实指Line2D类的实例。包括顶点vertex与连接顶点的线段。顶点有标记样式,线段有风格样式,以及绘画风格。我们知道matplotlib的制图方式是由Renderer将Artist描绘在Canvas上。
#
# class matplotlib.lines.Line2D(
# xdata, ydata,
# linewidth=None, linestyle=None, color=None,
# marker=None, markersize=None, markeredgewidth=None, markeredgecolor=None, markerfacecolor=None, markerfacecoloralt='none',
# fillstyle=None, antialiased=None, dash_capstyle=None, solid_capstyle=None, dash_joinstyle=None, solid_joinstyle=None, pickradius=5,
# drawstyle=None, markevery=None, **kwargs)
#
# 顶点 vertex
# vertex = (x, y) [vertex for vertex in zip(xdata, ydata)] == [*zip(xdata, ydata)]
# 7个相关方法。
#
# get_data(self, orig=True) : Return the xdata, ydata. If orig is True, return the original data.
# get_xdata(self, orig=True) : Return the xdata. If orig is True, return the original data, else the processed data.
# get_xydata(self) : Return the xy data as a Nx2 numpy array.
# get_ydata(self, orig=True) : Return the ydata. If orig is True, return the original data, else the processed data.
# set_data(self, *args) : Set the x and y data. *args(2, N) array or two 1D arrays
# set_xdata(self, x) : Set the data array for x. x <- 1D array
# set_ydata(self, y) : Set the data array for y. y <- 1D array
#
# xdata = [.1,.9,.9,.1]
# ydata = [.9,.9,.1,.1]
# fig = plt.figure()
# l1 = lines.Line2D(xdata,ydata)
# l2 = fig.add_artist(l1)
#
# 配置好Line2D,将它添加进Figura容器,渲染器就会将它描绘在画布上。在jupyter lab中的%matplotlib widget模式,自动完成了大部分事情。
# l1 == l2
# 线 Line
#
# lineStyles = {
# '': '_draw_nothing',
# ' ': '_draw_nothing',
# '-': '_draw_solid',
# '--': '_draw_dashed',
# '-.': '_draw_dash_dot',
# ':': '_draw_dotted',
# 'None': '_draw_nothing'}
#
# set_linestyle(self, ls) : Set the linestyle of the line.
# set_ls(self, ls) : Alias for set_linestyle.
# get_linestyle(self) : Return the linestyle.
# get_ls(self) : Alias for get_linestyle.
# set_dashes(self, seq) : Set the dash sequence.
# is_dashed(self) : Return whether line has a dashed linestyle.
#
# dash序列是浮点数序列,样式同(线,间,线,间,...),不知道追多支持几对。我们试一下(1,1,2,2,3,3,4,4,5,5,6,6,7,7,8,8,9,9) ok 我们可以假装它没限制了。数值的单位是点point,点是像素的集合,一个点有几个像素?应该由dpi定义。
#
# l1.set_dashes((1,1,2,2,3,3,4,4,5,5,6,6,7,7,8,8,9,9))
# l2 = fig.add_artist(l1)
#
# set_linestyle方法是set_dashes方法的超集,除了为几个虚线样式预定义了名称,额外增加了左移偏置功能。
#
# l1.set_linestyle((10,(20,10)))
# l2 = fig.add_artist(l1)
# A line - the line can have both a solid linestyle connecting all the vertices, and a marker at each vertex. Additionally, the drawing of the solid line is influenced by the drawstyle, e.g., one can create "stepped" lines in various styles.
# 在matplotlib中line这个名,实指Line2D类的实例。包括顶点vertex与连接顶点的线段。顶点有标记样式,线段有风格样式,以及绘画风格。我们知道matplotlib的制图方式是由Renderer将Artist描绘在Canvas上。
# %matplotlib widget
import matplotlib.lines as lines
import matplotlib.pyplot as plt
import numpy as np
# ```python
# class matplotlib.lines.Line2D(
# xdata, ydata,
# linewidth=None, linestyle=None, color=None,
# marker=None, markersize=None, markeredgewidth=None, markeredgecolor=None, markerfacecolor=None, markerfacecoloralt='none',
# fillstyle=None, antialiased=None, dash_capstyle=None, solid_capstyle=None, dash_joinstyle=None, solid_joinstyle=None, pickradius=5,
# drawstyle=None, markevery=None, **kwargs)
# ```
# ----
# ## 顶点 vertex
# vertex = (x, y)
# ```[vertex for vertex in zip(xdata, ydata)] == [*zip(xdata, ydata)]```
#
# 7个相关方法。
# - get_data(self, orig=True) : Return the xdata, ydata. If orig is True, return the original data.
# - get_xdata(self, orig=True) : Return the xdata. If orig is True, return the original data, else the processed data.
# - get_xydata(self) : Return the xy data as a Nx2 numpy array.
# - get_ydata(self, orig=True) : Return the ydata. If orig is True, return the original data, else the processed data.
# - set_data(self, \*args) : Set the x and y data. *args(2, N) array or two 1D arrays
# - set_xdata(self, x) : Set the data array for x. x <- 1D array
# - set_ydata(self, y) : Set the data array for y. y <- 1D array
xdata = [.1,.9,.9,.1]
ydata = [.9,.9,.1,.1]
fig = plt.figure()
l1 = lines.Line2D(xdata,ydata)
l2 = fig.add_artist(l1)
l1
l2
# 配置好Line2D,将它添加进Figura容器,渲染器就会将它描绘在画布上。在jupyter lab中的%matplotlib widget模式,自动完成了大部分事情。
# ----
# ## 线 Line
# > linewidth=None, linestyle=None, color=None
# ### linestyle or ls : {'-', '--', '-.', ':', '', (offset, on-off-seq), ...}
#
# ```python
# lineStyles = {
# '': '_draw_nothing',
# ' ': '_draw_nothing',
# '-': '_draw_solid',
# '--': '_draw_dashed',
# '-.': '_draw_dash_dot',
# ':': '_draw_dotted',
# 'None': '_draw_nothing'}
# ```
# - set_linestyle(self, ls) : Set the linestyle of the line.
# - set_ls(self, ls) : Alias for set_linestyle.
# - get_linestyle(self) : Return the linestyle.
# - get_ls(self) : Alias for get_linestyle.
# - set_dashes(self, seq) : Set the dash sequence.
# - is_dashed(self) : Return whether line has a dashed linestyle.
# The dash sequence is a sequence of floats of even length describing the length of dashes and spaces in points.
# dash序列是浮点数序列,样式同(线,间,线,间,...),不知道追多支持几对。我们试一下(1,1,2,2,3,3,4,4,5,5,6,6,7,7,8,8,9,9) ok 我们可以假装它没限制了。数值的单位是点point,点是像素的集合,一个点有几个像素?应该由dpi定义。
xdata = [.1,.9,.9,.1]
ydata = [.9,.9,.1,.1]
fig = plt.figure()
l1 = lines.Line2D(xdata,ydata)
l1.set_dashes((1,1,2,2,3,3,4,4,5,5,6,6,7,7,8,8,9,9))
l2 = fig.add_artist(l1)
l1.set_dashes((1,1,2,2,3,3,4,4,5,5,6,6,7,7,8,8,9,9))
l2 = fig.add_artist(l1)
# > ls{'-', '--', '-.', ':', '', (offset, on-off-seq), ...}
#
# set_linestyle方法是set_dashes方法的超级,除了为几个虚线样式预定义了名称,额外增加了偏置功能。
l1.set_linestyle((10,(20,10)))
l2 = fig.add_artist(l1)
# ----
# set_linewidth(self, w)
#
# 比想象中神奇
# - get_dash_capstyle(self) : Return the cap style for dashed lines.
# - get_dash_joinstyle(self) : Return the join style for dashed lines.
# - set_dash_joinstyle(self, s) : Set the join style for dashed lines. s <- {'miter', 'round', 'bevel'}
# - set_dash_capstyle(self, s) : Set the cap style for dashed lines. s <- {'butt', 'round', 'projecting'}
l2.get_dash_joinstyle()
l2.get_dash_capstyle()
from matplotlib.markers import MarkerStyle
# + jupyter={"outputs_hidden": true}
# MarkerStyle??
# -
MarkerStyle('.')
def set_marker(self, marker):
if (isinstance(marker, np.ndarray) and marker.ndim == 2 and
marker.shape[1] == 2):
self._marker_function = self._set_vertices
elif isinstance(marker, str) and cbook.is_math_text(marker):
self._marker_function = self._set_mathtext_path
elif isinstance(marker, Path):
self._marker_function = self._set_path_marker
elif (isinstance(marker, Sized) and len(marker) in (2, 3) and
marker[1] in (0, 1, 2, 3)):
self._marker_function = self._set_tuple_marker
elif (not isinstance(marker, (np.ndarray, list)) and
marker in self.markers):
self._marker_function = getattr(
self, '_set_' + self.markers[marker])
else:
try:
Path(marker)
self._marker_function = self._set_vertices
except ValueError:
raise ValueError('Unrecognized marker style {!r}'
.format(marker))
self._marker = marker
self._recache()
import matplotlib.cbook as cbook
from matplotlib.path import Path
def set_marker(marker):
if (isinstance(marker, np.ndarray) and marker.ndim == 2 and
marker.shape[1] == 2):
print(1)
elif isinstance(marker, str) and cbook.is_math_text(marker):
print(2)
elif isinstance(marker, Path):
print(3)
elif (not isinstance(marker, (np.ndarray, list))):
print(4)
else:
try:
print(5)
Path(marker)
except ValueError:
raise ValueError('Unrecognized marker style {!r}'
.format(marker))
marker = '.'
set_marker(marker)
cbook.is_math_text(marker)
isinstance(marker, Path)
markers = {
'.': 'point',
',': 'pixel',
'o': 'circle',
'v': 'triangle_down',
'^': 'triangle_up',
'<': 'triangle_left',
'>': 'triangle_right',
'1': 'tri_down',
'2': 'tri_up',
'3': 'tri_left',
'4': 'tri_right',
'8': 'octagon',
's': 'square',
'p': 'pentagon',
'*': 'star',
'h': 'hexagon1',
'H': 'hexagon2',
'+': 'plus',
'x': 'x',
'D': 'diamond',
'd': 'thin_diamond',
'|': 'vline',
'_': 'hline',
'P': 'plus_filled',
'X': 'x_filled',
TICKLEFT: 'tickleft',
TICKRIGHT: 'tickright',
TICKUP: 'tickup',
TICKDOWN: 'tickdown',
CARETLEFT: 'caretleft',
CARETRIGHT: 'caretright',
CARETUP: 'caretup',
CARETDOWN: 'caretdown',
CARETLEFTBASE: 'caretleftbase',
CARETRIGHTBASE: 'caretrightbase',
CARETUPBASE: 'caretupbase',
CARETDOWNBASE: 'caretdownbase',
"None": 'nothing',
None: 'nothing',
' ': 'nothing',
'': 'nothing'
}
def _set_circle(self, reduction=1.0):
self._transform = Affine2D().scale(0.5 * reduction)
self._snap_threshold = np.inf
fs = self.get_fillstyle()
if not self._half_fill():
self._path = Path.unit_circle()
else:
# build a right-half circle
if fs == 'bottom':
rotate = 270.
elif fs == 'top':
rotate = 90.
elif fs == 'left':
rotate = 180.
else:
rotate = 0.
self._path = self._alt_path = Path.unit_circle_righthalf()
self._transform.rotate_deg(rotate)
self._alt_transform = self._transform.frozen().rotate_deg(180.)
Path.unit_circle()
| api_Line2d.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PyCharm (python)
# language: python
# name: pycharm-56d7c5fb
# ---
#loading the dataset
from sklearn.datasets import load_iris
import numpy as np
iris_data=load_iris()
# print(iris_data)
print(iris_data.keys())
#keys of data set
print(iris_data['target_names'])
#the value of the key target_names is an array of strings,
#containing the species of flowers which we want to predict
print(iris_data['feature_names'])
#The value of feature_names is a list of strings, giving the description of each feature:
X=iris_data['data']
print(X)
y=iris_data['target']
print(y)
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X,y,random_state=0)
#we have assigned random_state as 0 so that each time when we run this ,we get the same output
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier(n_neighbors=4, p=1)
# p=1 manhatan distance
knn.fit(X_train,y_train)
sepel_len=eval(input("Enter the sepel length- "))
sepel_wid=eval(input("Enter the sepel width- "))
petal_len=eval(input("Enter the petal length- "))
petal_wid=eval(input("Enter the petal width- "))
X_new = np.array([[sepel_len,sepel_wid,petal_len,petal_wid]])
#calling predict method from knn for making prediction
prediction = knn.predict(X_new)
print("Predicted target name",iris_data['target_names'][prediction])
y_pred = knn.predict(X_test)
print("Test set prediction:\n {}".format(y_pred))
#accuracy
print('Test score is: {:.2f}'.format(knn.score(X_test,y_test)))
| 3_Naive Bayes dan K-Nearest Neighbor/iris_KNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Radial Profile
# This example shows how to create a radial profile from a SOXS event file, including using an exposure map to get flux-based quantities. We'll simulate a simple isothermal cluster.
import matplotlib
matplotlib.rc("font", size=18)
import matplotlib.pyplot as plt
import soxs
import astropy.io.fits as pyfits
# First, create the spectrum for the cluster using an absorbed thermal APEC model:
emin = 0.05 # keV
emax = 20.0 # keV
nbins = 20000
agen = soxs.ApecGenerator(emin, emax, nbins)
kT = 6.0
abund = 0.3
redshift = 0.05
norm = 1.0
spec = agen.get_spectrum(kT, abund, redshift, norm)
spec.rescale_flux(1.0e-13, emin=0.5, emax=2.0, flux_type="energy")
spec.apply_foreground_absorption(0.02)
# And a spatial distribution based on a $\beta$-model:
pos = soxs.BetaModel(30.0, 45.0, 50.0, 0.67)
# Generate a SIMPUT catalog from these two models, and write it to a file:
width = 10.0 # arcmin by default
nx = 1024 # resolution of image
cluster = soxs.SimputSpectrum.from_models("beta_model", spec, pos, width, nx)
cluster_cat = soxs.SimputCatalog.from_source("beta_model.simput", cluster, overwrite=True)
# and run the instrument simulation (for simplicity we'll turn off the point-source background):
soxs.instrument_simulator("beta_model.simput", "evt.fits", (100.0, "ks"), "lynx_hdxi", [30., 45.],
overwrite=True, ptsrc_bkgnd=False)
# Make an exposure map so that we can obtain flux-based quantities:
soxs.make_exposure_map("evt.fits", "expmap.fits", 2.3, overwrite=True)
# Make the radial profile, using energies between 0.5 and 5.0 keV, between radii of 0 and 200 arcseconds, with 50 bins:
soxs.write_radial_profile("evt.fits", "profile.fits", [30.0, 45.0],
0, 200, 50, emin=0.5, emax=5.0, expmap_file="expmap.fits", overwrite=True)
# Now we can use AstroPy's FITS reader to open the profile and have a look at the columns that are inside:
f = pyfits.open("profile.fits")
f["PROFILE"].columns
# and use Matplotlib to plot some quantities. We can plot the surface brightness:
plt.figure(figsize=(8,8))
plt.errorbar(f["profile"].data["rmid"], f["profile"].data["sur_bri"], lw=2, yerr=f["profile"].data["sur_bri_err"])
plt.xscale('log')
plt.yscale('log')
plt.xlabel("r (arcsec)")
plt.ylabel("S (cts/s/arcsec**2)")
# and, since we used an exposure map, the surface flux:
plt.figure(figsize=(8,8))
plt.errorbar(f["profile"].data["rmid"], f["profile"].data["sur_flux"], lw=2, yerr=f["profile"].data["sur_flux_err"])
plt.xscale('log')
plt.yscale('log')
plt.xlabel("r (arcsec)")
plt.ylabel("S (cts/s/cm**2/arcsec**2)")
| doc/source/cookbook/Radial_Profile.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Valuación de opciones asiáticas
# - Las opciones que tratamos la clase pasada dependen sólo del valor del precio del subyacente $S_t$, en el instante que se ejerce.
#
# - Cambios bruscos en el precio, cambian que la opción esté *in the money* a estar *out the money*.
#
# - **Posibilidad de evitar esto** $\longrightarrow$ suscribir un contrato sobre el valor promedio del precio del subyacente.
#
# - <font color ='red'> Puede proveer protección contra fluctuaciones extremas del precio en mercados volátiles. </font>
#
# - **Nombre**: Banco Trust de Tokio ofreció este tipo de opciones
#
# ### ¿Dónde se negocian?
#
# - Mercados OTC (Over the Counter / Independientes). Una explicación de esto podría ser el último punto de la lámina anterior.
#
# - Las condiciones para el cálculo matemático del promedio y otras condiciones son especificadas en el contrato. Lo que las hace un poco más “personalizables”.
#
# Existen diversos tipos de opciones asiáticas y se clasiflcan de acuerdo con lo siguiente.
#
# 1. La media que se utiliza puede ser **aritmética** o geométrica.
#
# 2. Media se calcula para $S_t \longrightarrow$ "Precio de ejercicio fijo". Media se calcula para precio de ejercicio $\longrightarrow$ "Precio de ejercicio flotante".
#
# 3. Si la opción sólo se puede ejercer al final del tiempo del contrato se dice que es asiática de tipo europeo o **euroasiática**, y si puede ejercer en cualquier instante, durante la vigencia del contrato se denomina **asiática de tipo americano.**
#
# Los tipos de opciones euroasiáticas son:
#
# - Call con precio de ejercicio fijo, función de pago: $\max\{A-K,0\}$.
# - Put con precio de ejercicio fijo, función de pago: $\max\{K-A,0\}$.
# - Call con precio de ejercicio flotante, función de pago: $\max\{S-K,0\}$.
# - Put con precio de ejercicio flotante, función de pago: $\max\{K-S,0\}$.
#
# Donde $A$ es el promedio del precio del subyacente.
#
# $$\text{Promedio aritmético} \quad A={1\over T} \int_0^TS_tdt$$
# $$\text{Promedio geométrico} \quad A=\exp\Big({1\over T} \int_0^T Ln(S_t) dt\Big)$$
#
# De aquí en adelante denominaremos **Asiática ** $\longrightarrow$ Euroasiática y se analizará el call asiático con **K Fijo**.
#
# Se supondrá un solo activo con riesgo, cuyos proceso de precios $\{S_t | t\in [0,T]\}$ satisface un movimiento browniano geométrico, en un mercado que satisface las suposiciones del modelo de Black y Scholes. Recordemos que bajo esta medida de probabilidad, $P^*$, denominada de riesgo neutro, bajo la cual el precio del activo, $S_t$, satisface:
#
# $$dS_t = rS_tdt+\sigma S_tdW_t,\quad 0\leq t \leq T, S_0 >0$$
#
# Para un call asiático de promedio aritmético y con precio de ejercicios fijo, está dado por
# $$\max \{A(T)-K,0\} = (A(T)-K)_+$$
#
# con $A(x)={1\over x} \int_0^x S_u du$
# Se puede ver que el valor en el tiempo t de la opción call asiática está dado por:
#
# $$ V_t(K) = e^{-r(T-t)}E^*[(A(T)-K)_+]$$
#
# Para el caso de interés, *Valución de la opción*, donde $t_0=0$ y $t=0$, se tiene:
#
# $$\textbf{Valor call asiático}\longrightarrow V_0(K)=e^{-rT}E\Bigg[ \Big({1\over T} \int_0^T S_u du -K\Big)_+\Bigg]$$
# ## Usando Monte Carlo
#
# Para usar este método es necesario que se calcule el promedio $S_u$ en el intervalo $[0,T]$. Para esto se debe aproximar el valor de la integral por los siguiente dos métodos.
#
# Para los dos esquemas se dividirá el intervalo $[0,T]$ en N subintervalos de igual longitud, $h={T\over N}$, esto determina los tiempos $t_0,t_1,\cdots,t_{N-1},t_N $, en donde $t_i=ih$ para $i=0,1,\cdots,N$
#
# ### Sumas de Riemann
#
# $$\int_0^T S_u du \approx h \sum_{i=0}^{n-1} S_{t_i}$$
#
# Reemplanzando esta aproximación en el valor del call asiático obtenemos la siguiente aproximación:
#
# $$\hat V_0^{(1)}= {e^{-rT} \over M} \sum_{j=1}^{M} \Bigg({1\over N} \sum_{i=0}^{N-1} S_{t_i}-K \Bigg)_+$$
#
# ## Mejorando la aproximación de las sumas de Riemann (esquema del trapecio)
#
# 
# Desarrollando la exponencial en serie de taylor y suponiendo que $h$ es pequeña, sólo se conservan los términos de orden uno, se tiene la siguiente aproximación:
# $$\int_0^T S_u du \approx {h \over 2}\sum_{i=0}^{N-1}S_{t_i}(2+rh+(W_{t_{i+1}}-W_{t_i})\sigma)$$
#
# Reemplazando esta aproximación en el precio del call, se tiene la siguiente estimación:
# $$\hat V_0^{(2)}= {e^{-rT} \over M} \sum_{j=1}^{M} \Bigg({h\over 2T} \sum_{i=0}^{N-1} S_{t_i}(2+rh+(W_{t_{i+1}}-W_{t_i})\sigma)-K \Bigg)_+$$
# ## Ejemplo
#
# Como caso de prueba se seleccionó el de un call asiático con precio inicial, $S_0 = 100$, precio de ejercicio $K = 100$, tasa libre de riesgo $r = 0.10$, volatilidad $\sigma = 0.20$ y $T = 1$ año. Cuyo precio es $\approx 7.04$.
#importar los paquetes que se van a usar
import pandas as pd
import pandas_datareader.data as web
import numpy as np
import datetime
import matplotlib.pyplot as plt
import scipy.stats as st
import seaborn as sns
# %matplotlib inline
#algunas opciones para Pandas
pd.set_option('display.notebook_repr_html', True)
pd.set_option('display.max_columns', 6)
pd.set_option('display.max_rows', 10)
pd.set_option('display.width', 78)
pd.set_option('precision', 3)
# +
def BSprices(mu,sigma,S0,NbTraj,NbStep):
T = 1
nu = mu-(sigma**2)/2
DeltaT = T/NbStep
SqDeltaT = np.sqrt(DeltaT)
#for i in range(NbStep):
DeltaW = SqDeltaT*np.random.randn(NbTraj,NbStep-1)
increments = nu*DeltaT + sigma*DeltaW
concat = np.concatenate((np.log(S0)*np.ones([NbTraj,1]),increments),axis=1)
LogSt = np.cumsum(concat,axis=1)
St = np.exp(LogSt)
t = np.arange(0,1,DeltaT)
return St.T,t
def calc_daily_ret(closes):
return np.log(closes/closes.shift(1)).iloc[1:]
# +
NbTraj = 2
NbStep = 100
S0 = 100
r = 0.10
sigma = 0.2
K = 100
T = 1
# Resolvemos la ecuación de black scholes para obtener los precios
St,t = BSprices(r,sigma,S0,NbTraj,NbStep)
t = t*NbStep
plt.plot(t,St,label='precios')
prices = pd.DataFrame(St,index=t)
# Obtenemos los precios promedios en todo el tiempo
Average_t = prices.expanding().mean()
plt.plot(t,Average_t,label='Promedio de precios')
plt.legend()
plt.show()#
# -
# ### Método sumas de Riemann
# +
#### Sumas de Riemann
strike = pd.DataFrame(K*np.ones([NbStep,NbTraj]), index=t)
call = pd.DataFrame({'Prima':np.exp(-r*T) \
*np.fmax(Average_t-strike,np.zeros([NbStep,NbTraj])).mean(axis=1)}, index=t)
# .mean(axis=1) realiza el promedio entre las filas de np.fmax()
call.plot()
print(call.iloc[-1])
# intervalos de confianza
confianza = 0.95
sigma_est = prices.iloc[-1].sem()
mean_est = call.iloc[-1].Prima
i1 = st.t.interval(confianza,NbTraj-1, loc=mean_est, scale=sigma_est)
i2 = st.norm.interval(confianza, loc=mean_est, scale=sigma_est)
print(i1)
print(i2)
# -
# Ahora hagamos pruebas variando la cantidad de trayectorias `NbTraj` y la cantidad de números de puntos `NbStep` para obtener el
# +
NbTraj = [1000,5000,10000]
NbStep = [10,50,100]
S0 = 100 # Precio inicial
r = 0.10 # Tasa libre de riesgo
sigma = 0.2 # volatilidad
K = 100 # Strike price
T = 1 # Tiempo de cierre - años
Call = np.zeros([len(NbTraj),len(NbStep)])
intervalos = []#np.zeros([len(NbTraj),len(NbStep)])
i = 0 # controla las filas
j = 0 # controla las columnas
for tray in NbTraj:
j = 0
for nescen in NbStep:
St,t = BSprices(r,sigma,S0,tray,nescen)
prices = pd.DataFrame(St,index=t)
Average_t = prices.expanding().mean()
strike = pd.DataFrame(K*np.ones([nescen,tray]), index=t)
call = pd.DataFrame({'Prima':np.exp(-r*T) \
*np.fmax(Average_t-strike,np.zeros([nescen,tray])).mean(axis=1)}, index=t)
Call[i,j]= call.iloc[-1]
# intervalos de confianza
confianza = 0.95
sigma_est = prices.iloc[-1].sem()
mean_est = call.iloc[-1].Prima
# i1 = st.t.interval(confianza,nescen-1, loc=mean_est, scale=sigma_est)
i2 = st.norm.interval(confianza, loc=mean_est, scale=sigma_est)
intervalos.append(np.asarray(i2))
j+=1
i+=1
intervalos = np.reshape(np.asarray(intervalos),[3,6])
filas = ['Nbtray = %i' %i for i in NbTraj]
col = ['NbStep = %i' %i for i in NbStep]+['int %i' %j for j in range(6)]
df = pd.DataFrame(index=filas,columns=col)
df.loc[0:4,0:3] = Call
df.loc[0:,3:] = intervalos
df
# -
# <script>
# $(document).ready(function(){
# $('div.prompt').hide();
# $('div.back-to-top').hide();
# $('nav#menubar').hide();
# $('.breadcrumb').hide();
# $('.hidden-print').hide();
# });
# </script>
#
# <footer id="attribution" style="float:right; color:#808080; background:#fff;">
# Created with Jupyter by <NAME>.
# </footer>
| TEMA-3/Clase19_ValuacionOpcionesAsiaticas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # test note
#
#
# * jupyterはコンテナ起動すること
# * テストベッド一式起動済みであること
#
# !pip install --upgrade pip
# !pip install --force-reinstall ../lib/ait_sdk-0.1.7-py3-none-any.whl
from pathlib import Path
import pprint
from ait_sdk.test.hepler import Helper
import json
# +
# settings cell
# mounted dir
root_dir = Path('/workdir/root/ait')
ait_name='eval_regression_analyze_rmse_and_mae_tf2.3'
ait_version='0.1'
ait_full_name=f'{ait_name}_{ait_version}'
ait_dir = root_dir / ait_full_name
td_name=f'{ait_name}_test'
# (dockerホスト側の)インベントリ登録用アセット格納ルートフォルダ
# current_dir = %pwd
with open(f'{current_dir}/config.json', encoding='utf-8') as f:
json_ = json.load(f)
root_dir = json_['host_ait_root_dir']
is_container = json_['is_container']
invenotory_root_dir = f'{root_dir}\\ait\\{ait_full_name}\\local_qai\\inventory'
# entry point address
# コンテナ起動かどうかでポート番号が変わるため、切り替える
if is_container:
backend_entry_point = 'http://host.docker.internal:8888/qai-testbed/api/0.0.1'
ip_entry_point = 'http://host.docker.internal:8888/qai-ip/api/0.0.1'
else:
backend_entry_point = 'http://host.docker.internal:5000/qai-testbed/api/0.0.1'
ip_entry_point = 'http://host.docker.internal:6000/qai-ip/api/0.0.1'
# aitのデプロイフラグ
# 一度実施すれば、それ以降は実施しなくてOK
is_init_ait = True
# インベントリの登録フラグ
# 一度実施すれば、それ以降は実施しなくてOK
is_init_inventory = True
# -
helper = Helper(backend_entry_point=backend_entry_point,
ip_entry_point=ip_entry_point,
ait_dir=ait_dir,
ait_full_name=ait_full_name)
# +
# health check
helper.get_bk('/health-check')
helper.get_ip('/health-check')
# -
# create ml-component
res = helper.post_ml_component(name=f'MLComponent_{ait_full_name}', description=f'Description of {ait_full_name}', problem_domain=f'ProbremDomain of {ait_full_name}')
helper.set_ml_component_id(res['MLComponentId'])
# deploy AIT
if is_init_ait:
helper.deploy_ait_non_build()
else:
print('skip deploy AIT')
res = helper.get_data_types()
model_data_type_id = [d for d in res['DataTypes'] if d['Name'] == 'model'][0]['Id']
dataset_data_type_id = [d for d in res['DataTypes'] if d['Name'] == 'dataset'][0]['Id']
res = helper.get_file_systems()
unix_file_system_id = [f for f in res['FileSystems'] if f['Name'] == 'UNIX_FILE_SYSTEM'][0]['Id']
windows_file_system_id = [f for f in res['FileSystems'] if f['Name'] == 'WINDOWS_FILE'][0]['Id']
# +
# add inventories
if is_init_inventory:
inv1_name = helper.post_inventory('dataset_for_verification', dataset_data_type_id, windows_file_system_id,
f'{invenotory_root_dir}\\dataset_for_verification\\test_dataset.csv',
'幸福度調査データで使用した学習用データ', ['csv'])
inv2_name = helper.post_inventory('trained_model', model_data_type_id, windows_file_system_id,
f'{invenotory_root_dir}\\trained_model\\model_2.h5',
'幸福度調査データで構築した幸福度予測モデル', ['h5'])
else:
print('skip add inventories')
# +
# get ait_json and inventory_jsons
res_json = helper.get_bk('/QualityMeasurements/RelationalOperators', is_print_json=False).json()
eq_id = int([r['Id'] for r in res_json['RelationalOperator'] if r['Expression'] == '=='][0])
nq_id = int([r['Id'] for r in res_json['RelationalOperator'] if r['Expression'] == '!='][0])
gt_id = int([r['Id'] for r in res_json['RelationalOperator'] if r['Expression'] == '>'][0])
ge_id = int([r['Id'] for r in res_json['RelationalOperator'] if r['Expression'] == '>='][0])
lt_id = int([r['Id'] for r in res_json['RelationalOperator'] if r['Expression'] == '<'][0])
le_id = int([r['Id'] for r in res_json['RelationalOperator'] if r['Expression'] == '<='][0])
res_json = helper.get_bk('/testRunners', is_print_json=False).json()
ait_json = [j for j in res_json['TestRunners'] if j['Name'] == ait_name][-1]
inv_1_json = helper.get_inventory(inv1_name)
inv_2_json = helper.get_inventory(inv2_name)
# +
# add teast_descriptions
helper.post_td(td_name, ait_json['QualityDimensionId'],
quality_measurements=[
{"Id":ait_json['Report']['Measures'][0]['Id'], "Value":"0.5", "RelationalOperatorId":lt_id, "Enable":True},
{"Id":ait_json['Report']['Measures'][1]['Id'], "Value":"0.5", "RelationalOperatorId":lt_id, "Enable":True}
],
target_inventories=[
{"Id":1, "InventoryId": inv_2_json['Id'], "TemplateInventoryId": ait_json['TargetInventories'][0]['Id']},
{"Id":2, "InventoryId": inv_1_json['Id'], "TemplateInventoryId": ait_json['TargetInventories'][1]['Id']}
],
test_runner={
"Id":ait_json['Id'],
"Params":[
{"TestRunnerParamTemplateId":ait_json['ParamTemplates'][0]['Id'], "Value":"Happiness Score"}
]
})
# -
# get test_description_jsons
td_1_json = helper.get_td(td_name)
# run test_descriptions
helper.post_run_and_wait(td_1_json['Id'])
res_json = helper.get_td_detail(td_1_json['Id'])
pprint.pprint(res_json)
# generate report
res = helper.post_report(td_1_json['Id'])
| ait_repository/test/tests/eval_regression_analyze_rmse_and_mae_tf2.3.ipynb |
# # An embedding-based test
# +
from pkg.utils import set_warnings
set_warnings()
import datetime
import time
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
from graspologic.embed import select_svd
from graspologic.plot import pairplot
from graspologic.utils import augment_diagonal
from myst_nb import glue as default_glue
from pkg.data import load_network_palette, load_node_palette, load_unmatched
from pkg.io import savefig
from pkg.plot import set_theme
from pkg.stats import rdpg_test
from pkg.utils import get_seeds
DISPLAY_FIGS = False
FILENAME = "rdpg_unmatched_test"
rng = np.random.default_rng(8888)
def gluefig(name, fig, **kwargs):
savefig(name, foldername=FILENAME, **kwargs)
glue(name, fig, prefix="fig")
if not DISPLAY_FIGS:
plt.close()
def glue(name, var, prefix=None):
savename = f"{FILENAME}-{name}"
if prefix is not None:
savename = prefix + ":" + savename
default_glue(savename, var, display=False)
t0 = time.time()
set_theme(font_scale=1.25)
network_palette, NETWORK_KEY = load_network_palette()
node_palette, NODE_KEY = load_node_palette()
left_adj, left_nodes = load_unmatched("left")
right_adj, right_nodes = load_unmatched("right")
left_nodes["inds"] = range(len(left_nodes))
right_nodes["inds"] = range(len(right_nodes))
seeds = get_seeds(left_nodes, right_nodes)
# -
# ## A test based on latent positions
# ### Look at the singular values
# +
def screeplot(sing_vals, elbow_inds=None, color=None, ax=None, label=None):
if ax is None:
_, ax = plt.subplots(1, 1, figsize=(8, 4))
plt.plot(range(1, len(sing_vals) + 1), sing_vals, color=color, label=label)
if elbow_inds is not None:
plt.scatter(
elbow_inds,
sing_vals[elbow_inds - 1],
marker="x",
s=50,
zorder=10,
color=color,
)
ax.set(ylabel="Singular value", xlabel="Index")
return ax
max_n_components = 64
_, left_singular_values, _ = select_svd(
augment_diagonal(left_adj), n_elbows=6, n_components=max_n_components
)
_, right_singular_values, _ = select_svd(
augment_diagonal(right_adj), n_elbows=6, n_components=max_n_components
)
fig, ax = plt.subplots(1, 1, figsize=(8, 6))
screeplot(
left_singular_values,
color=network_palette["Left"],
ax=ax,
label="Left",
)
screeplot(
right_singular_values,
color=network_palette["Right"],
ax=ax,
label="Right",
)
ax.legend()
gluefig("screeplot", fig)
# -
#
# ```{glue:figure} fig:rdpg_unmatched_test-screeplot
# :name: "fig:rdpg_unmatched_test-screeplot"
#
# Comparison of the singular values from the spectral decompositions of the left and
# right hemisphere adjacency matrices. Note that the right hemisphere singular values
# tend to be slightly higher than the corresponding singular value on the left
# hemisphere, which is consistent with an increased density on the right hemisphere as
# seen in [](er_unmatched_test.ipynb).
# ```
# ### Run the test
n_components = 8 # TODO trouble is that this choice is somewhat arbitrary...
stat, pvalue, misc = rdpg_test(
left_adj, right_adj, seeds=seeds, n_components=n_components
)
glue("pvalue", pvalue)
# ### Look at the embeddings
# +
Z1 = misc["Z1"]
Z2 = misc["Z2"]
def plot_latents(
left,
right,
title="",
n_show=4,
alpha=0.6,
linewidth=0.4,
s=10,
connections=False,
palette=None,
):
if n_show > left.shape[1]:
n_show = left.shape[1]
plot_data = np.concatenate([left, right], axis=0)
labels = np.array(["Left"] * len(left) + ["Right"] * len(right))
pg = pairplot(
plot_data[:, :n_show],
labels=labels,
title=title,
size=s,
palette=palette,
)
# pg._legend.remove()
return pg
n_show = 4
pg = plot_latents(Z1, Z2, palette=network_palette, n_show=n_show)
fig = pg.figure
eff_n_components = Z1.shape[1]
glue("n_show", n_show)
glue("eff_n_components", eff_n_components)
gluefig("latents", fig)
# -
#
# ```{glue:figure} fig:rdpg_unmatched_test-latents
# :name: "fig:rdpg_unmatched_test-latents"
#
# Comparison of the latent positions used for the test based on the random dot product
# graph. This plot shows only the first {glue:text}`rdpg_unmatched_test-n_show`
# dimensions, though the test was run in {glue:text}`rdpg_unmatched_test-eff_n_components`.
# The p-value for the test comparing the multivariate distribution of latent positions
# for the left vs. the right hemispheres (distance correlation 2-sample test) is
# {glue:text}`rdpg_unmatched_test-pvalue:.2f`, indicating that we fail to reject our
# null hypothesis of bilateral symmetry under this null model.
# ```
| docs/rdpg_unmatched_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: rscube
# language: python
# name: rscube
# ---
# This notebook simply downloads the data used for the datacube from ASF and Hansen. This is so that the tutorial cna be done independent of available data. This is not the thrust of what we are doing, as we expect users will incorporate their own data for segmentation.
# # Downloading the Data over the Waxlake
from pathlib import Path
import rasterio
from skimage.restoration import denoise_tv_bregman
import numpy as np
import lxml.etree as etree
import datetime
import matplotlib.pyplot as plt
from rscube import get_cropped_profile, interpolate_nn
from tqdm import tqdm
from itertools import starmap
# ## Making the Data Directory
#
# We are downloading into a subdirectory within data called `full_coverage`.
DATA_DIR = Path('data/full_coverage/')
DATA_DIR.mkdir(exist_ok=True, parents=True)
# ## ASF ALOS-1 Radiometrically and Terrain Corrected data
# We used the ASF search tool to download data over mangroves in Malaysia. Here is the specific areas within the [search tool](https://search.asf.alaska.edu/#/?zoom=11.066666666666666¢er=100.744323,4.539299&polygon=POLYGON((100.4527%204.584,100.746%204.584,100.746%204.9416,100.4527%204.9416,100.4527%204.584))&dataset=ALOS&productTypes=RTC_LOW_RES&beamModes=FBD&resultsLoaded=true&granule=ALPSRP252880080-RTC_LOW_RES&path=490-490&frame=80-80). We then add these images to the queue and download the python script. We have the data in our `data/asf_data/download-alos1-data-malaysaia.py` which we now use. You will need an [Nasa Earthdata login](https://urs.earthdata.nasa.gov/) to proceed.
#
# The first command **should be run in your terminal without the `!`**. We run in it in the cell is so that you can *input your login credentials*. Below is what the output will be.
# +
# #!python data/asf_data/download-alos1-plr-wax-lake.py
# -
# You can run this from the terminal or the cell.
# !unzip '*.zip' -d 'data/asf_data/'
# rm -rf AP_*.zip
# ## Hansen Data
#
# This is aquite a large file so be patient.
# +
# # !curl https://storage.googleapis.com/earthenginepartners-hansen/GFC-2019-v1.7/Hansen_GFC-2019-v1.7_last_30N_100W.tif --output data/full_coverage/Hansen_GFC-2019-v1.7_last_30N_100W.tif
# -
# # Denoising and Renaming ALOS Data
asf_dir = Path('data/asf_data/')
alos_paths = sorted(list(asf_dir.glob('AP_*/*RT1_H*.tif'))) + sorted(list(asf_dir.glob('AP_*/*RT1_VV.tif')))
alos_paths
def get_alos_date(metadata_xml_path):
tree = etree.parse(open(metadata_xml_path))
root = tree.getroot()
dataAcquisition_elements = root.xpath('//gml:beginPosition', namespaces=root.nsmap)
assert(len(dataAcquisition_elements) == 1)
element = dataAcquisition_elements[0].text
date = datetime.date(int(element[:4]), int(element[5:7]), int(element[8:10]))
return date
metadata_path = list(asf_dir.glob('AP_*/*.iso.xml'))[0]
retreival_date = get_alos_date(metadata_path)
retreival_date
def read_one(path, k=1):
with rasterio.open(path) as ds:
arr = ds.read(k)
arr = np.clip(arr, 0, 1)
arr[arr == 0] = np.nan
return arr
# We get the metadata from the ASF data.
with rasterio.open(alos_paths[0]) as ds:
ASF_ALOS_PROFILE = ds.profile
quad_pol_data = list(map(read_one, alos_paths))
plt.imshow(quad_pol_data[0], vmax=.5)
sy = np.s_[:2000]
sx = np.s_[:2700]
plt.imshow(quad_pol_data[0][sy, sx], vmax=.8)
cropped_profile = get_cropped_profile(ASF_ALOS_PROFILE, sx, sy)
cropped_profile
# Now, we are going to crop each image according to these slices.
quad_pol_data_c = list(map(lambda img: img[sy, sx], quad_pol_data))
# Our focus is for segmentation, so we need to despeckle our image. We are going to use [`Mulog`](https://www.charles-deledalle.fr/pages/files/mulog_high_quality_version.pdf). We ignore the so-called debiasing because we are going to scale the dynamic range before segmentation to incorporate this data with the Landsat mosaic from Hansen.
def denoise(img):
# Fill in nodata areas with nearest neighbor
img_nn = interpolate_nn(img)
# Convert to db and make noise additive
img_nn = 10 * np.log10(img_nn)
# Use TV denoising
# This parameter lambda = .2 worked well
# Higher values mean less denoising and lower mean
# image will appear smoother.
img_nn_tv = denoise_tv_bregman(img_nn, .2)
img_denoised = 10**(img_nn_tv / 10.)
img_denoised[np.isnan(img)] = np.nan
return img_denoised
# We inspect this denoising on a small subset.
plt.figure(figsize=(10, 10))
X = quad_pol_data_c[0][750:1250, 1000: 2000]
plt.imshow(X)
plt.figure(figsize=(10, 10))
X_d = denoise(X)
plt.imshow(X_d)
quad_pol_data_c_tv = list(map(denoise, tqdm(quad_pol_data_c)))
def write_one(arr, dest_path):
with rasterio.open(dest_path, 'w', **cropped_profile) as ds:
ds.write(arr.astype(np.float32), 1)
return dest_path
y, m, d = retreival_date.year, retreival_date.month, retreival_date.day
dest_paths = [DATA_DIR/f'ALOS1_RTC_{pol}_{y}{m}{d}_tv.tif' for pol in ['HH', 'HV', 'VV']]
dest_paths
list(starmap(write_one, zip(tqdm(quad_pol_data_c_tv), dest_paths)))
| notebooks/superpixel_segmentation/0 - Downloading and Formatting the Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="z2Id0h3kTG5F" outputId="747ff533-9c91-4a20-9de4-56fb5a3eaec2"
from google.colab import drive
drive.mount('/content/gdrive', force_remount=True)
# + colab={"base_uri": "https://localhost:8080/"} id="mLzXIwx9WI9f" outputId="2279dff3-5613-4397-dd43-03437559198f"
# !pip install pypinyin pywubi zhconv overrides boto3
# + colab={"base_uri": "https://localhost:8080/"} id="7Oob8oexy_N4" outputId="f6de1fb1-4671-4526-86b5-72554e1d80f5"
import time
import torch
print(torch.cuda.get_device_name(0))
start = time.perf_counter()
# + colab={"base_uri": "https://localhost:8080/"} id="AxJvcOtUTLkH" outputId="00f6a9c1-37e6-4125-f211-36b6e1d45ebb"
# !python3 "/content/gdrive/MyDrive/Colab Data/Chinese Characters/glyce/glyce/bin/run_bert_glyce_tagger.py" \
# --data_sign resume_ner \
# --task_name ner \
# --config_path "/content/gdrive/MyDrive/Colab Data/Chinese Characters/glyce/glyce/configs/ctb6cws_glyce_bert.json" \
# --data_dir "/content/gdrive/MyDrive/Colab Data/Chinese Characters/ner/resume" \
# --output_name "/content/gdrive/MyDrive/Colab Data/Chinese Characters/glyce/glyce/my_export_models/resume_ner glyce.bin" \
# --bert_model "/content/gdrive/MyDrive/Colab Data/Chinese Characters/chinese_L-12_H-768_A-12" \
# --checkpoint 239 \
# --seed 8008 \
# --max_seq_length 150 \
# --do_train \
# --do_eval \
# --train_batch_size 16 \
# --dev_batch_size 16 \
# --test_batch_size 16 \
# --learning_rate 3e-5 \
# --num_train_epochs 15 \
# --warmup_proportion 0 \
# --training_strat "bert-glyce-joint"
# + colab={"base_uri": "https://localhost:8080/"} id="n6tw6wYio3rR" outputId="e9f0f50a-7066-4b71-a72b-17474c861669"
end = time.perf_counter()
elapsed = end-start
hours = elapsed//(60*60)
mins = (elapsed - hours*60*60)//60
secs = (elapsed - hours*60*60 - mins*60)
print("Time elapsed: %02d:%02d:%02d" % (hours,mins,secs))
| glyce/glyce/experiments/Resume_NER_Glyce.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # First challenge: https://app.codility.com/c/run/trainingDUVMCF-8E6/
N = int(input())
# +
def solution(N):
print(len(max(bin(N)[2:].strip('0').strip('1').split('1'))))
solution(N)
# -
# **Deconstructing this solution:**
# When we convert a number to binary in python using 'bin(N)', it always gives us a '0b' in the beginning:
bin(304)
# So we slice after the first two characters with '[2:]':
bin(304)[2:]
# As we are just trying to count the maximum gap in between the 1s, we use '.strip()'. So .strip('0') strips any outer 0s:
bin(304)[2:].strip('0')
# Getting there! Now we strip the outer 1s with .strip('1'):
bin(304)[2:].strip('0').strip('1')
# at this point we could just count the length of the zeros with 'len()', but this disregards some edge cases. What if there was two gaps of zeros? Let's input a different value this time:
bin(1041)[2:].strip('0').strip('1')
# aha. So we split the remaining values at the 1 with .split('1'):
bin(1041)[2:].strip('0').strip('1').split('1')
# Great. Now we count the greater gap with 'max()':
max(bin(1041)[2:].strip('0').strip('1').split('1'))
# Now we count the length of the maximum gap! (len())
len(max(bin(1041)[2:].strip('0').strip('1').split('1')))
# # Question 2: https://app.codility.com/c/run/training8VJ2XN-9P5/
# Ok, gonna try to learn something additional right now. I want to figure out how I can input a list along with the K number.
#
# *note: I will delete any cells that i use for experimenting and just save the working answers, as the experiment cells won't do me any good in the future.*
A = list(map(int,input("\nEnter the numbers : ").strip().split()))
K = int(input())
A
# So I know that 'map(int, input())' will tell("map") each value in the input to be an integer. If we get more than one value, '.strip()' will strip all spaces from the ends of the input.
A = list(map(int,input("\nEnter the numbers : ").strip()))
# 'split()' alone will split the remaining values up by the default separator (if nothing is specified), which is a space, into a list. We still have to coerce that with list() though, otherwise A will just be a map object.
a = list(map(int,input("\nEnter the numbers : ").strip().split()))
a
# Back to the question.
# +
#3 8 9 7 6
#3
# -
A = list(map(int,input("\nEnter the numbers : ").strip().split()))
K = int(input())
last = (A.pop(-1))
A
# +
def solution(A, K):
for i in range(K):
last = (A.pop(-1))
A.insert(0, last)
return A
solution(A, K)
# -
# holy shit. I did it!!
# Oh wait. So I entered my answer and got an 87% on codility because I didn't account for an extreme case. Which is if A is just an empty list. Gonna put my for loop into an if/else statement.
# +
def solution(A, K):
if len(A) == 0:
pass
else:
for i in range(K):
last = (A.pop(-1))
A.insert(0, last)
return A
solution(A, K)
# -
# And 100%!!!
# Breaking that down: If the length of A is 0, meaning an empty list, then just skip everything and output that empty list. If there is anything in the list of A, then we iterate through each value in the range of K. For each value in the range of K, I pop the last '(-1)' number off the list and assign that to a variable 'last'. Then I insert that value 'last' into the first, 0, space of the list A. Return list A!
# Wow codility really hid that assumption there:
#
# "Assume that:
#
# N and K are integers within the range [0..100];
# **each element of array A is an integer within the range [−1,000..1,000].**"
# # Question 3: https://app.codility.com/programmers/lessons/2-arrays/odd_occurrences_in_array/
A = list(map(int, input("\nEnter the numbers: ").strip().split()))
A
no_dups = [9, 3, 7]
# I think 'set()' removes duplicates. let's see
# +
def solution(A):
no_dups = set(A)
for i in A:
if i in no_dups:
A.pop()
return list(A)
solution(A)
# -
# We can do a true/false check for the duplicates in the array:
a_set = set(A)
len(A) != len(a_set)
# OK set is not gonna for for us
# +
def solution(A):
for element in A:
if A.count(element) == 1:
return element
solution(A)
# -
# 50% Success!! because of the time complexity. Some of the test inputs are extremely large to test for this:
#
# Detected time complexity:
# O(N**2)
#
# Let's break down the answer: For each element in A, if the count of that element is greater is just 1, return that element.
#
# Now let's try to fix the runtime complexity.
#
# This is an answer I found which uses a "bitwise" operator. New to me!
# +
def solution(A):
result = 0
for number in A:
result ^= number
return result
solution(A)
# +
three = 9
three ^= 1
three
# -
# good youtube vid for explaining bitwise operators https://www.youtube.com/watch?v=PyfKCvHALj8
print(0 ^ 9)
print(0 ^ 7)
# Ok, think I got it. So bitwise operators work in binary. When we use any bitwise operators (&, |, ^). The computer is converting to or thinking in binary. If there is an odd number of values in an array, the "^" will return that number.
# # https://app.codility.com/programmers/lessons/3-time_complexity/frog_jmp/
X, Y, D = map(int, input().split())
# So what we are trying to do here is have the frog start at X, then jump a distance, D, just enough times to equal or pass Y.
# +
import math
def solution(X, Y, D):
return math.ceil(((Y - X)/D))
solution(X, Y, D)
# -
# great success! I hope we are able to import extra libraries like math, although finding another way to round up an integer wouldn't be so hard.
# # : https://app.codility.com/programmers/lessons/3-time_complexity/perm_missing_elem/
# So we have an array, A, with N different integers in it. The integers in the arrary have a range of (1, (N+1)). Since there are only N integers, that extra one is missing. return that missing element.
A = list(map(int, input().split()))
# +
def solution(A):
n = len(A)
total = (n + 1)*(n + 2)/2
return int(total - sum(A))
solution(A)
# -
# breaking this down: so we need to know the len of A, N.
n = len(A)
n + 1
n+2
5*6/2
sum(A)
# If we multiply 1+len(N) by 2+len(N) and divide by 2, we get the sum of what would be the list without the missing value. If we subtract this by the sum of the actual array A (with missing value), the differene is that missing value.
# # https://app.codility.com/programmers/lessons/3-time_complexity/tape_equilibrium/
A = list(map(int, input().split()))
# Ok, this one was a little confusing. We have an array A, consisting of N values. P is any value from 1 to N-1. In other words 0 > P > N. P can split the array at A[P] into two separate arrays. We are looking at the minimum difference between the sum of the two split arrays.
# +
def solution(A):
first_half = 0
second_half = 0
diffs = []
for i in range(0, len(A)-1):
first_half += A[i]
second_half = sum(A) - A[i]
diffs.append(abs(first_half-second_half))
return min(diffs)
solution(A)
# -
# nevermind, this did horribly
# I iterate through each value that P could be.
#
# For each, I can add each corresponding value in A to the first half, and the second_half with be the sum of A minus that corresponding P value.
#
# Then I append the difference of the halfs for each potential P to diffs and selection the minimum,
# +
def solution(A):
m = float('inf') #It acts as an unbounded upper value for comparison. sets the value to infit.
left_sum = 0
for i in range(len(A)-1):
left_sum += A[i]
m = min(abs(sum(A) - 2*left_sum), m)
return m
solution(A)
# -
# This was a working answer I found.
# Broken down: so for each value in the potential range of P, which is anything from 1 to N[-1], i add that to the first half. Then, m becomes either the minimum of the absolute difference of 2 x left_sum, or the two halves, or infinity- still in each iteration of the for loop. Still not totally sure why we need the infinity in there.
print(min(3, float('inf')))
print(min(5, float('inf')))
# So for each iteration, it is re-storing the minimum difference and then selecting the minimum of either that or infinity?? tf.
# +
def solution(A):
mins = []
left_sum = 0
for i in range(len(A)-1):
left_sum += i
mins.append(abs(sum(A) - 2*left_sum))
return min(mins)
solution(A)
# -
# Just 100% correct with 0% performance. WHY
# Ok, I think I understand now. In the for loop, we get our minimum and **reassign itself**, always keeping the minimum number it comes across. Fuck.
def solution(A):
m = float('inf')
left_sum = 0
for i in A[:-1]:
left_sum += i
m = min(abs(sum(A) - 2*left_sum), m)
return m
# 53% performance. ?!?!
# +
def solution(A):
s = sum(A)
m = float('inf')
left_sum = 0
for i in A[:-1]:
left_sum += i
m = min(abs(s - 2*left_sum), m)
return m
solution(yerp)
# -
# Wow!! I just learned some shit right there. even if I still only have the one for loop, because I was adding the sum(A), that was too much calculation to do every iteration in the extreme cases. So the sum(A) added a magnitude of complexity when inside the for loop, as opposed to just calculating sum(A) once in the outside.
def solution(A):
mins = []
s = sum(A)
left_sum = 0
for i in range(len(A)-1):
left_sum += A[i]
mins.append(abs(s - 2*left_sum))
return min(mins)
# final answer.
# # https://app.codility.com/programmers/lessons/4-counting_elements/frog_river_one/
A = int(input())
X = list(map(int, input().split()))
# so we want to go through each. add to a list called 'steps'. and stop when all values of range(A) are in steps.
# +
def solution(A, X):
steps = []
for i in range(0, len(A)):
if X not in steps:
steps.append(A[i])
return i
solution(A, X)
# -
A
A[1]-1
# found genius, simple answer on youtube! thank the lord. they use set() and enumerate() which i've thought about but wasn't sure how.
# +
def solution(A, X):
steps = set() #set compiles only unique values
for i, j in enumerate(X): #for index, value in enumerate (gives index to array X)
steps.add(j) #add the value to steps set,
if len(steps) == A: #once the len(steps), or number of leafs equals A, return that position!
return i
return -1 #if the steps never equals the length of A, meaning frog can't cross, return -1
solution(A, X)
# -
# # https://app.codility.com/programmers/lessons/4-counting_elements/max_counters/
N = int(input())
A = list(map(int, input().split()))
counters = [0]*N
counters
counters[3] += 1
counters[3]
# +
def solution(N, A):
counters = [0] * N
for value in A:
if 1 <= value <= N: #if the value in A is greater than 1 and less than N, that position of that value in counters is increased by 1
counters[value-1] += 1
else:
counters = [max(counters)]*N
return counters
solution(N, A)
# -
# 66%! 100% correct, just lacking on runtime complexity because of max(counters) in the for loop.
# +
def solution(N, A):
counters = [0] * N
for value in A:
if 1 <= value <= N: #if the value in A is greater than 1 and less than N, that position of that value in counters is increased by 1
counters[value-1] += 1
else:
counters = [max(counters)]*N
return counters
solution(N, A)
# -
# # https://app.codility.com/programmers/lessons/4-counting_elements/missing_integer/
A = list(map(int, input().split()))
# +
def solution(A):
smallest = 1
unique = set(A)
for int in unique:
if int == smallest:
smallest += 1
return smallest
solution(A)
# +
def solution(A):
smallest = 1
unique = set(A)
for i in unique:
if i == smallest:
smallest += 1
return smallest
solution(A)
# -
# 88%
a = -1
int(a)
# # https://app.codility.com/programmers/lessons/4-counting_elements/perm_check/
A = list(map(int, input().split()))
# +
def solution(A):
if len(A) == max(A):
return 1
else:
return 0
solution(A)
# -
# 83%
# +
def solution(A):
A = sorted(A)
if A == list(range(1, max(A)+1)):
return 1
else:
return 0
solution(A)
# -
list(range(1, max(A)+1))
A
# 83%
# # balanced password
# A string is balanced if it consists of exactly two different characters and both of those characters appear exactly the same number of times. For example: "aabbab" is balanced (both 'a' and 'b' occur three times) but "aabba" is not balanced ('a' occurs three times, 'b' occurs two times). String "aabbcc" is also not balanced (it contains three different letters).
S = str(input())
# +
def solution(S):
dict = {char: S.count(char) for char in set(S)}
if len(list(set(list(dict.values())))) == 1 and len(dict) == 2:
return 'Balanced'
else:
return 'Not balanced'
solution(S)
# -
dict = {char: S.count(char) for char in set(S)}
dict
set(S)
len(list(set(list(dict.values())))) == 1
#coerce the unique dict values into a set. if there is more than 1 unique value, then the value counts are not equal, and therefore not balanced.
# +
print(list(dict.values()))
# -
set(list(dict.values()))
len(set(S))
set(S)
for i in enumerate
# _____
def solution(A, B, C, D):
clock = []
if A in range(0, 3)
list(range(0, 3))
| Deconstructing Challenge Solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [toc]
#
# > 原文地址 https://www.linuxidc.com/Linux/2016-12/138979.htm
#
# ## [CentOS](http://www.linuxidc.com/topicnews.aspx?tid=14 "CentOS")6.5
#
# ### CentOS6.5查看防火墙的状态
#
# ```sh
# service iptable status
# ```
#
# 显示结果:
#
# ```sh
# [linuxidc@localhost ~]$service iptable status
# Redirecting to /bin/systemctl status iptable.service
# ● iptable.service
# Loaded: not-found (Reason: No such file or directory)
# Active: inactive (dead) --表示防火墙已经关闭
# ```
#
# ### CentOS 6.5 关闭防火墙
#
# - 临时关闭防火墙
# ```sh
# servcie iptables stop
# ```
# - 永久关闭防火墙
# ```sh
# chkconfig iptables off
# ```
#
# ## CentOS 7.2
#
# ### 查看防火墙状态
#
# 关闭后显示 not running,开启后显示 running
# ```sh
# firewall-cmd --state
# ```
# 或者
# ```sh
# systemctl list-unit-files|grep firewalld.service
# ```
#
# 或者
#
# ```sh
# systemctl status firewalld.service
# ```
#
# ### 其他常用命令
#
# - 启动服务:systemctl start firewalld.service
# - 关闭服务:systemctl stop firewalld.service
# - 重启服务:systemctl restart firewalld.service
# - 显示服务的状态:systemctl status firewalld.service
# - 在开机时启用服务:systemctl enable firewalld.service
# - 在开机时禁用服务:systemctl disable firewalld.service
# - 查看服务是否开机启动:systemctl is-enabled firewalld.service;echo $?
# - 查看已启动的服务列表:systemctl list-unit-files|grep enabled
#
# ## Centos 7 firewall 命令
#
# ### 查看已经开放的端口
#
# ```sh
# firewall-cmd --list-ports
# ```
#
# ### 开启端口
#
# ```sh
# firewall-cmd --zone=public --add-port=80/tcp --permanent
#
# # 删除一个端口
# firewall-cmd --zone=public --remove-port=80/tcp --permanent
#
# # 开放多个端口
# firewall-cmd --zone=public --add-port=10000-10020/tcp --permanent
# ```
#
# 命令含义
#
# - --zone #作用域
# - --add-port=80/tcp #添加端口,格式为:端口 / 通讯协议
# - permanent #永久生效,没有此参数重启后失效
#
# ### 重启防火墙
#
# ```sh
# firewall-cmd --reload #重启firewall
# systemctl stop firewalld.service #停止firewall
# systemctl disable firewalld.service #禁止firewall开机启动
# firewall-cmd --state #查看默认防火墙状态(关闭后显示notrunning,开启后显示running)
# ```
#
# ## CentOS 7 以下版本 iptables 命令
#
# 如要开放 80,22,8080 端口,输入以下命令即可
#
# ```sh
# /sbin/iptables -I INPUT -p tcp --dport 80 -j ACCEPT
# /sbin/iptables -I INPUT -p tcp --dport 22 -j ACCEPT
# /sbin/iptables -I INPUT -p tcp --dport 8080 -j ACCEPT
# ```
#
# 然后保存:
#
# ```sh
# /etc/rc.d/init.d/iptables save
# ```
#
# 查看打开的端口:
#
# ```sh
# /etc/init.d/iptables status
# ```
#
# ### 关闭防火墙
#
# 1. 永久性生效,重启后不会复原
#
# 开启: chkconfig iptables on
#
# 关闭: chkconfig iptables off
#
# 2. 即时生效,重启后复原
#
# 开启: service iptables start
#
# 关闭: service iptables stop
#
# 3. 查看防火墙状态: service iptables status
#
# ## CentOS7 和 6 的默认防火墙的区别
#
# CentOS 7 默认使用的是 firewall 作为防火墙,使用 iptables 必须重新设置一下
#
# 1、直接关闭防火墙
#
# ```sh
# systemctl stop firewalld.service # 停止 firewall
# systemctl disable firewalld.service # 禁止 firewall 开机启动
# ```
#
# 2、设置 iptables service
#
# ```sh
# yum -y install iptables-services
# ```
#
# 如果要修改防火墙配置,如增加防火墙端口 3306
#
# ```sh
# vi /etc/sysconfig/iptables
# ```
#
# 增加规则
# ```sh
# -A INPUT -m state --state NEW -m tcp -p tcp --dport 3306 -j ACCEPT
# ```
# 保存退出后
#
# ```sh
# systemctl restart iptables.service # 重启防火墙使配置生效
# systemctl enable iptables.service # 设置防火墙开机启动
# ```
#
# 最后重启系统使设置生效即可。
# ```sh
# systemctl start iptables.service # 打开防火墙
# systemctl stop iptables.service # 关闭防火墙
# ```
#
# ### 解决主机不能访问虚拟机 CentOS 中的站点
#
# 前阵子在虚拟机上装好了 CentOS6.2,并配好了 apache+php+mysql,但是本机就是无法访问。一直就没去折腾了。
#
# 具体情况如下
#
# > 1. 本机能 ping 通虚拟机
# >
# > 2. 虚拟机也能 ping 通本机
# >
# > 3. 虚拟机能访问自己的 web
# >
# > 4. 本机无法访问虚拟机的 web
#
# 后来发现是**防火墙将 80 端口屏蔽了的缘故。**
#
# 检查是不是服务器的 80 端口被防火墙堵了,可以通过命令:telnet server_ip 80 来测试。
#
# 解决方法如下:
#
# > /sbin/iptables -I INPUT -p tcp --dport 80 -j ACCEPT
#
# 然后保存:
#
# > /etc/rc.d/init.d/iptables save
#
# 重启防火墙
#
# > /etc/init.d/iptables restart
#
# CentOS 防火墙的关闭,关闭其服务即可:
#
# > 查看 CentOS 防火墙信息:/etc/init.d/iptables status
# >
# > 关闭 CentOS 防火墙服务:/etc/init.d/iptables stop
#
# 更多 CentOS 相关信息见[CentOS](https://www.linuxidc.com/topicnews.aspx?tid=14)专题页面[http://www.linuxidc.com/topicnews.aspx?tid=14](https://www.linuxidc.com/topicnews.aspx?tid=14 "CentOS")
#
# **本文永久更新链接地址**:[http://www.linuxidc.com/Linux/2016-12/138979.htm](https://www.linuxidc.com/Linux/2016-12/138979.htm)
| 技术/CentOS - 防火墙.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
import time
import numpy as np
import pandas as pd
import json
import matplotlib.pyplot as plt
import datetime
import boto3
import sagemaker
from sagemaker import get_execution_role
with_categories = False
# Set a good base job name when building different models
# It will help in identifying trained models and endpoints
base_job_name = 'deepar-biketrain-with-dynamic-feat'
# +
bucket = 'chandra-ml-sagemaker'
prefix = 'deepar/bikerental'
# This structure allows multiple training and test files for model development and testing
s3_data_path = "{}/{}/data_dynamic".format(bucket, prefix)
s3_output_path = "{}/{}/output".format(bucket, prefix)
# -
s3_data_path,s3_output_path
# File name is referred as key name in S3
# Files stored in S3 are automatically replicated across
# three different availability zones in the region where the bucket was created.
# http://boto3.readthedocs.io/en/latest/guide/s3.html
def write_to_s3(filename, bucket, key):
with open(filename,'rb') as f: # Read in binary mode
return boto3.Session().resource('s3').Bucket(bucket).Object(key).upload_fileobj(f)
# Upload one or more training files and test files to S3
write_to_s3('train_dynamic_feat.json',bucket,'deepar/bikerental/data_dynamic/train/train_dynamic_feat.json')
write_to_s3('test_dynamic_feat.json',bucket,'deepar/bikerental/data_dynamic/test/test_dynamic_feat.json')
sagemaker_session = sagemaker.Session()
role = get_execution_role()
# We no longer have to maintain a mapping of container images by region
# Simply use the convenience method provided by sagemaker
# https://docs.aws.amazon.com/sagemaker/latest/dg/sagemaker-algo-docker-registry-paths.html
from sagemaker.amazon.amazon_estimator import get_image_uri
image_name = get_image_uri(boto3.Session().region_name, 'forecasting-deepar')
image_name
# +
freq='H' # Timeseries consists Hourly Data and we need to predict hourly rental count
# how far in the future predictions can be made
# 12 days worth of hourly forecast
prediction_length = 288
# aws recommends setting context same as prediction length as a starting point.
# This controls how far in the past the network can see
context_length = 288
# +
# Check Free Tier (if you are still under free-tier)
# At this time, m4.xlarge is offered as part of 2 months free tier
# https://aws.amazon.com/sagemaker/pricing/
# If you are outside of free-tier, you can also use ml.m5.xlarge (newer generation instance)
# In this example, I am using ml.m5.xlarge for training
# Dynamic Feat - Using a large instance ml.c5.4xlarge = 16 CPU, 32 GB
# 'ml.c4.xlarge' -> 'ml.c5.4xlarge'. out of memory error with c4.xlarge
estimator = sagemaker.estimator.Estimator(
sagemaker_session=sagemaker_session,
image_name=image_name,
role=role,
train_instance_count=1,
train_instance_type='ml.c5.4xlarge',
base_job_name=base_job_name,
output_path="s3://" + s3_output_path
)
# -
freq, context_length, prediction_length
# https://docs.aws.amazon.com/sagemaker/latest/dg/deepar_hyperparameters.html
hyperparameters = {
"time_freq": freq,
"epochs": "400",
"early_stopping_patience": "40",
"mini_batch_size": "64",
"learning_rate": "5E-4",
"context_length": str(context_length),
"prediction_length": str(prediction_length),
"cardinality" : "auto" if with_categories else ''
}
hyperparameters
estimator.set_hyperparameters(**hyperparameters)
# Here, we are simply referring to train path and test path
# You can have multiple files in each path
# SageMaker will use all the files
data_channels = {
"train": "s3://{}/train/".format(s3_data_path),
"test": "s3://{}/test/".format(s3_data_path)
}
data_channels
# This step takes around 35 minutes to train the model with m4.xlarge instance
estimator.fit(inputs=data_channels)
job_name = estimator.latest_training_job.name
print ('job name: {0}'.format(job_name))
# Create an endpoint for real-time predictions
endpoint_name = sagemaker_session.endpoint_from_job(
job_name=job_name,
initial_instance_count=1,
instance_type='ml.m4.xlarge',
deployment_image=image_name,
role=role
)
print ('endpoint name: {0}'.format(endpoint_name))
# +
# Don't forget to terminate the end point after completing the demo
# Otherwise, you account will accumulate hourly charges
# you can delete from sagemaker management console or through command line or throught code
#sagemaker_session.delete_endpoint(endpoint_name)
# -
| 8 DeepAR/BikeRental/sdk1.7/deepar_biketrain_dynamic_feat_cloud_training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="zfDcmTIqtf8j"
# # Pre-processing examples on modcloth dataset
#
# ### Attribution: <NAME>, Kaggle
#
# The notebook was released under the [Apache 2.0](http://www.apache.org/licenses/LICENSE-2.0) open source license.
# + [markdown] id="_b_falCzsxI3" papermill={"duration": 0.03365, "end_time": "2020-10-07T13:01:49.380394", "exception": false, "start_time": "2020-10-07T13:01:49.346744", "status": "completed"} tags=[]
# # **About the dataset**
#
# In this notebook, we will use modcloth_final_data.json as input dataset
# + id="psZFn_WLsxI3" papermill={"duration": 1.23869, "end_time": "2020-10-07T13:01:50.652746", "exception": false, "start_time": "2020-10-07T13:01:49.414056", "status": "completed"} tags=[]
# import necessary libraries
# File read and EDA(Data Cleansing & Transformations)
import numpy as np
import pandas as pd
# EDA Visualization
import matplotlib.pyplot as plt
import seaborn as sns
# + [markdown] id="XD_u28pnuGP_"
# ## Mounting gDrive
# + id="tjrT33QduFiX"
#Mounting gDrive in Colaboratory
try:
from google.colab import drive
drive.mount("/content/drive/", force_remount=True)
google_drive_prefix = "/content/drive/My Drive"
data_prefix = "{}/mnist/".format(google_drive_prefix)
except ModuleNotFoundError:
data_prefix = "data/"
# + id="hK0Nd6qWuY7M"
#Change directory to my folder for analytics labs where I have cloned my gitHub repositories with magic command.
# %cd drive/My Drive/Data_analytics_lab
# + [markdown] id="ekMls2ZmsxI3" papermill={"duration": 0.034773, "end_time": "2020-10-07T13:01:50.722822", "exception": false, "start_time": "2020-10-07T13:01:50.688049", "status": "completed"} tags=[]
# # Read input json data
# + id="cVHvyXpFsxI3" papermill={"duration": 1.522854, "end_time": "2020-10-07T13:01:52.279963", "exception": false, "start_time": "2020-10-07T13:01:50.757109", "status": "completed"} tags=[]
#Read file and view first ten rows
mc_data= pd.read_json("data_code_along/modcloth_final_data.json", lines=True)
mc_data.head() # displays first 5 records in the dataframe
# + [markdown] id="rT0bM85GsxI4" papermill={"duration": 0.050436, "end_time": "2020-10-07T13:01:52.381097", "exception": false, "start_time": "2020-10-07T13:01:52.330661", "status": "completed"} tags=[]
# ## EDA - Exploratory Data Analysis
# + [markdown] id="nmtChSFRsxI4" papermill={"duration": 0.049735, "end_time": "2020-10-07T13:01:52.481036", "exception": false, "start_time": "2020-10-07T13:01:52.431301", "status": "completed"} tags=[]
# # Column names are inconsistent
# Some of the column names are having space and rest of them are having underscore in between them. Hence try to be consistent by adding underscore instead of space
#
# > for your information, size is the keyword in pandas , make sure to change the feature name "size" to some user defined name like "mc_size"
# https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.size.html
# + id="QmpOm7FdsxI4" papermill={"duration": 0.053752, "end_time": "2020-10-07T13:01:52.586622", "exception": false, "start_time": "2020-10-07T13:01:52.532870", "status": "completed"} tags=[]
mc_data.columns = ['item_id', 'waist', 'mc_size', 'quality', 'cup_size', 'hips', 'bra_size', 'category', 'bust', 'height', 'user_name', 'length', 'fit', 'user_id', 'shoe_size', 'shoe_width', 'review_summary', 'review_test']
# + [markdown] id="2tufuE6WsxI5" papermill={"duration": 0.034055, "end_time": "2020-10-07T13:01:52.655611", "exception": false, "start_time": "2020-10-07T13:01:52.621556", "status": "completed"} tags=[]
# > see the total number of observations, column names and datatypes info
# + id="zMBJT6NRsxI5" papermill={"duration": 0.136337, "end_time": "2020-10-07T13:01:52.826289", "exception": false, "start_time": "2020-10-07T13:01:52.689952", "status": "completed"} tags=[]
mc_data.info()
# + [markdown] id="26PMR8C4sxI5" papermill={"duration": 0.034869, "end_time": "2020-10-07T13:01:52.896385", "exception": false, "start_time": "2020-10-07T13:01:52.861516", "status": "completed"} tags=[]
# # Sparse Data
#
# Given data is having lot of missing values , for example look at the columns such as shoe_size and show_width.
#
# > lets check the missing values percentage for each feature
# + id="4Ui9o69DsxI5" papermill={"duration": 0.183048, "end_time": "2020-10-07T13:01:53.115403", "exception": false, "start_time": "2020-10-07T13:01:52.932355", "status": "completed"} tags=[]
missing_data_sum = mc_data.isnull().sum()
missing_data = pd.DataFrame({'total_missing_values': missing_data_sum,'percentage_of_missing_values': (missing_data_sum/mc_data.shape[0])*100})
missing_data
# + [markdown] id="GmJ3IVcIsxI6" papermill={"duration": 0.034981, "end_time": "2020-10-07T13:01:53.185402", "exception": false, "start_time": "2020-10-07T13:01:53.150421", "status": "completed"} tags=[]
# *Out of 18 columns, only 6 columns have complete data. And columns such as waist , bust, shoe_size, show_width and hips are highly sparse*
# + [markdown] id="1jdjY6RXsxI6" papermill={"duration": 0.034739, "end_time": "2020-10-07T13:01:53.255064", "exception": false, "start_time": "2020-10-07T13:01:53.220325", "status": "completed"} tags=[]
# > Check Data types which are having numerical/categorical data
# + id="Eu4_BkshsxI6" papermill={"duration": 0.044227, "end_time": "2020-10-07T13:01:53.334514", "exception": false, "start_time": "2020-10-07T13:01:53.290287", "status": "completed"} tags=[]
mc_data.dtypes
# + [markdown] id="hWLmj3tLsxI6" papermill={"duration": 0.036486, "end_time": "2020-10-07T13:01:53.406680", "exception": false, "start_time": "2020-10-07T13:01:53.370194", "status": "completed"} tags=[]
# *pandas library identifies item_id, waist, mc_size, quality, hips, bra_size, user_id, shoe_size are numeric And cup_size, category, bust, height, user_name, length, fit, shoe_width, review_summary , review_test are object type . Take away from this is "There are some numeric data columns are fall under Object types" . Hence we need to handle the misclassification of these data types. For example, bust data contains numeric values but its dtype is Object.*
# + [markdown] id="IT69P8BOsxI7" papermill={"duration": 0.035696, "end_time": "2020-10-07T13:01:53.478268", "exception": false, "start_time": "2020-10-07T13:01:53.442572", "status": "completed"} tags=[]
# # Unique number of observations for each feature
# + [markdown] id="12cwMjaasxI7" papermill={"duration": 0.036718, "end_time": "2020-10-07T13:01:53.550334", "exception": false, "start_time": "2020-10-07T13:01:53.513616", "status": "completed"} tags=[]
# > If the dataset is having less number of observations then we can see the unique data that resides in each feature(There are 82790 observations)
# + id="YCk1qC9osxI7" papermill={"duration": 0.220401, "end_time": "2020-10-07T13:01:53.806347", "exception": false, "start_time": "2020-10-07T13:01:53.585946", "status": "completed"} tags=[]
mc_data.nunique()
# + [markdown] id="MTpmVe7ksxI7" papermill={"duration": 0.036537, "end_time": "2020-10-07T13:01:53.880003", "exception": false, "start_time": "2020-10-07T13:01:53.843466", "status": "completed"} tags=[]
# *With this, we can clearly understand there is no unique data entirely for one feature to make the index if any.
# And just to make a note the columns item_id and user_id are repeating.*
# + [markdown] id="rb9o0xLjsxI7" papermill={"duration": 0.036336, "end_time": "2020-10-07T13:01:53.952864", "exception": false, "start_time": "2020-10-07T13:01:53.916528", "status": "completed"} tags=[]
# > Lets look into unique observations which are having less uniqueness
# + id="yU9ngyJDsxI8" papermill={"duration": 0.045479, "end_time": "2020-10-07T13:01:54.035455", "exception": false, "start_time": "2020-10-07T13:01:53.989976", "status": "completed"} tags=[]
def countplot(indipendent_features):
plt.figure(figsize=(25, 25))
for loc, feature in enumerate(indipendent_features):
ax = plt.subplot(3, 4, loc+1)
ax.set_xlabel('{}'.format(feature), fontsize=10)
chart = sns.countplot(mc_data[feature])
chart.set_xticklabels(chart.get_xticklabels(), rotation=90)
return None
# + id="ZyjITizHsxI8" papermill={"duration": 2.080919, "end_time": "2020-10-07T13:01:56.170181", "exception": false, "start_time": "2020-10-07T13:01:54.089262", "status": "completed"} tags=[]
uniques_data = ['quality', 'cup_size', 'bra_size', 'category', 'length', 'fit', 'shoe_size', 'shoe_width', 'height', 'bust', 'mc_size']
countplot(uniques_data)
# + [markdown] id="nCM02xHZsxI8" papermill={"duration": 0.055171, "end_time": "2020-10-07T13:01:56.281008", "exception": false, "start_time": "2020-10-07T13:01:56.225837", "status": "completed"} tags=[]
# *Few observations*
# * cup_size contains some format which might represents the measurement
# * shoe_size 38 is an outlier , there we can see lot of variance
# * height column also having few outliers (May be we can see those things after converting categorical data into numeric values)
# * there are categorical data exists such as shoe_width, category, length, fit and height.
# * For the feature bust - clearly there is one observation with different data, hence we need to format it i.e "37 - 39". Will try to replace this value with mean
#
# + id="5nXl92ETsxI8" papermill={"duration": 0.070394, "end_time": "2020-10-07T13:01:56.405966", "exception": false, "start_time": "2020-10-07T13:01:56.335572", "status": "completed"} tags=[]
# replacing bust unformatted value with mean 38 which is taken from the values 37 & 39
mc_data.at[mc_data[mc_data.bust == '37-39'].index[0],'bust'] = '38'
# + [markdown] id="WzlIbH56sxI8" papermill={"duration": 0.037623, "end_time": "2020-10-07T13:01:56.482203", "exception": false, "start_time": "2020-10-07T13:01:56.444580", "status": "completed"} tags=[]
# # Height feature - Convert US units to Metric units (ft & in to cm).
# + id="2S8GUZopsxI9" papermill={"duration": 0.361257, "end_time": "2020-10-07T13:01:56.881541", "exception": false, "start_time": "2020-10-07T13:01:56.520284", "status": "completed"} tags=[]
def height_in_cms(ht):
if ht.lower() != 'nan':
ht = ht.replace('ft','').replace('in', '')
h_ft = int(ht.split()[0])
if len(ht.split()) > 1:
h_inch = int(ht.split()[1])
else:
h_inch = 0
h_inch += h_ft * 12
h_cm = round(h_inch * 2.54, 1)
return h_cm
mc_data.height = mc_data.height.astype(str).apply(height_in_cms)
mc_data.head()
# + [markdown] id="YK4csn9asxI9" papermill={"duration": 0.038316, "end_time": "2020-10-07T13:01:56.958670", "exception": false, "start_time": "2020-10-07T13:01:56.920354", "status": "completed"} tags=[]
# > we successfully converted metrics to centimetres. Now lets handle the missing values with mean imputation and then look into the outliers for this height feature. Use box/scatter plot for outliers visualization
# + id="6SbMrdWfsxI9" papermill={"duration": 0.050139, "end_time": "2020-10-07T13:01:57.049667", "exception": false, "start_time": "2020-10-07T13:01:56.999528", "status": "completed"} tags=[]
mc_data.height.fillna(value=mc_data.height.mean(), inplace=True)
mc_data.height.isnull().sum()
# + id="Fv5KxtrKsxI-" papermill={"duration": 0.154647, "end_time": "2020-10-07T13:01:57.244188", "exception": false, "start_time": "2020-10-07T13:01:57.089541", "status": "completed"} tags=[]
def plot_outlier(feature):
plt.figure(figsize=(25, 6))
ax = sns.boxplot(x=feature, linewidth=2.5)
plot_outlier(mc_data.height)
# + [markdown] id="m6wi1uMUsxI-" papermill={"duration": 0.038802, "end_time": "2020-10-07T13:01:57.322282", "exception": false, "start_time": "2020-10-07T13:01:57.283480", "status": "completed"} tags=[]
# > Check the lower and upper cutoff range values for the outliers
# + id="em-I_ABCsxI-" papermill={"duration": 0.046529, "end_time": "2020-10-07T13:01:57.408002", "exception": false, "start_time": "2020-10-07T13:01:57.361473", "status": "completed"} tags=[]
def get_outliers_range(datacolumn):
sorted(datacolumn)
Q1,Q3 = np.percentile(datacolumn , [25,75])
IQR = Q3 - Q1
lower_range = Q1 - (1.5 * IQR)
upper_range = Q3 + (1.5 * IQR)
return lower_range,upper_range
# + id="K5eTw2N6sxI-" papermill={"duration": 0.066125, "end_time": "2020-10-07T13:01:57.514753", "exception": false, "start_time": "2020-10-07T13:01:57.448628", "status": "completed"} tags=[]
ht_lower_range,ht_upper_range = get_outliers_range(mc_data.height)
ht_lower_range,ht_upper_range
# + [markdown] id="GU7ZLNY1sxI-" papermill={"duration": 0.040367, "end_time": "2020-10-07T13:01:57.596233", "exception": false, "start_time": "2020-10-07T13:01:57.555866", "status": "completed"} tags=[]
# > Take away "there are so many outliers" exists. Here I have used Inter Quartile Range calculation to find the lower range and upper range cutoff.
# So the outlier would be anything less than the lower range cutoff(144.7) or anything more than the upper range cutoff(185.5) is an outlier. \
#
# Note: there are different techniques to identify the outliers
# Please check out this link for more details on this
# https://statisticsbyjim.com/basics/outliers/
#
# + [markdown] id="zUYtGKVzsxI_" papermill={"duration": 0.040198, "end_time": "2020-10-07T13:01:57.677007", "exception": false, "start_time": "2020-10-07T13:01:57.636809", "status": "completed"} tags=[]
# > Lets count how many outliers exists for this height feature
#
# + id="w7LiRuyCsxI_" papermill={"duration": 0.104176, "end_time": "2020-10-07T13:01:57.821988", "exception": false, "start_time": "2020-10-07T13:01:57.717812", "status": "completed"} tags=[]
mc_data[(mc_data.height < ht_lower_range) | (mc_data.height > ht_upper_range)]
# + [markdown] id="CfBn4vObsxI_" papermill={"duration": 0.042027, "end_time": "2020-10-07T13:01:57.905016", "exception": false, "start_time": "2020-10-07T13:01:57.862989", "status": "completed"} tags=[]
# > Take Away "There are 199 outliers". which is 0.2% of total number observations. Hence we can remove/drop/delete these outliers
# + id="NI7gQoVfsxI_" papermill={"duration": 0.072994, "end_time": "2020-10-07T13:01:58.019553", "exception": false, "start_time": "2020-10-07T13:01:57.946559", "status": "completed"} tags=[]
mc_df = mc_data.drop(mc_data[(mc_data.height < ht_lower_range) | (mc_data.height > ht_upper_range)].index)
mc_df.reset_index(drop=True, inplace=True)
mc_df.shape
# + [markdown] id="iooYNNvYsxJA" papermill={"duration": 0.042094, "end_time": "2020-10-07T13:01:58.103287", "exception": false, "start_time": "2020-10-07T13:01:58.061193", "status": "completed"} tags=[]
# > Lets look again the height feature using box plot to see the handling of outlier
# + id="duEORYTesxJA" papermill={"duration": 0.169618, "end_time": "2020-10-07T13:01:58.315263", "exception": false, "start_time": "2020-10-07T13:01:58.145645", "status": "completed"} tags=[]
plot_outlier(mc_df.height)
# + [markdown] id="4-vWNHSqsxJA" papermill={"duration": 0.043231, "end_time": "2020-10-07T13:01:58.401165", "exception": false, "start_time": "2020-10-07T13:01:58.357934", "status": "completed"} tags=[]
# # Numeric features distributions Visualization
# + id="LDZ_OSgIsxJA" papermill={"duration": 0.051569, "end_time": "2020-10-07T13:01:58.497065", "exception": false, "start_time": "2020-10-07T13:01:58.445496", "status": "completed"} tags=[]
def plot_dist(df, indipendent_features):
plt.figure(figsize=(25, 20))
for loc, feature in enumerate(indipendent_features):
ax = plt.subplot(3, 3, loc+1)
sns.distplot(df[feature]) # you can try histplot as well
return None
# + id="VIoaJgG8sxJA" papermill={"duration": 1.338943, "end_time": "2020-10-07T13:01:59.879258", "exception": false, "start_time": "2020-10-07T13:01:58.540315", "status": "completed"} tags=[]
plot_dist(mc_data, ['height', 'waist', 'mc_size', 'quality', 'hips', 'bra_size', 'shoe_size'])
# + [markdown] id="MpX6x5IIsxJA" papermill={"duration": 0.045302, "end_time": "2020-10-07T13:01:59.970311", "exception": false, "start_time": "2020-10-07T13:01:59.925009", "status": "completed"} tags=[]
# # Missing Values Handling for numeric features
#
# > as we see there are lot of missing values in this dataset. Since the data is highly sparse, i am trying to use KNN algorith to impute the relavant features.
# + id="GSzwlhuosxJB" papermill={"duration": 264.070364, "end_time": "2020-10-07T13:06:24.086033", "exception": false, "start_time": "2020-10-07T13:02:00.015669", "status": "completed"} tags=[]
from sklearn.impute import KNNImputer
imputer = KNNImputer(n_neighbors=10)
# finding imputation using other features (it will take couple of minutes to complete the execution)
mc_data_knn_ind_features = mc_df[['waist', 'hips', 'bra_size', 'bust', 'height', 'shoe_size']]
df_filled = imputer.fit_transform(mc_data_knn_ind_features)
knn_numeric_imputations = pd.DataFrame(data=df_filled, columns=['waist', 'hips', 'bra_size', 'bust', 'height', 'shoe_size'])
# remove the existing numeric columns (waist, height, hips, bra_size, bust, shoe_size ) from the main dataframe and concatenate with knn imputed data
#mc_df = mc_data
mc_new_df = mc_df.drop(['waist', 'hips', 'bra_size', 'bust', 'height', 'shoe_size'], axis=1)
# + id="GoLkjtRNsxJB" papermill={"duration": 0.139941, "end_time": "2020-10-07T13:06:24.272893", "exception": false, "start_time": "2020-10-07T13:06:24.132952", "status": "completed"} tags=[]
# concat the imputations data with mc data frame
mc = pd.concat([mc_new_df, knn_numeric_imputations], axis=1)
mc.isnull().sum()
# + [markdown] id="1q_GYj0vsxJB" papermill={"duration": 0.045941, "end_time": "2020-10-07T13:06:24.364511", "exception": false, "start_time": "2020-10-07T13:06:24.318570", "status": "completed"} tags=[]
# > we successfully done the imputations for some of the numeric features
# + [markdown] id="mm5x7U-ksxJB" papermill={"duration": 0.045661, "end_time": "2020-10-07T13:06:24.456728", "exception": false, "start_time": "2020-10-07T13:06:24.411067", "status": "completed"} tags=[]
# # Handling shoe-size outliers
# + id="_OVkyofZsxJB" papermill={"duration": 0.154279, "end_time": "2020-10-07T13:06:24.656849", "exception": false, "start_time": "2020-10-07T13:06:24.502570", "status": "completed"} tags=[]
plot_outlier(mc.shoe_size)
# + [markdown] id="6zK1lAAEsxJC" papermill={"duration": 0.045063, "end_time": "2020-10-07T13:06:24.747638", "exception": false, "start_time": "2020-10-07T13:06:24.702575", "status": "completed"} tags=[]
# clearly, there are few outliers, using IQR cutoff range values remove there observations
# + id="2O6w0_QUsxJC" papermill={"duration": 0.224317, "end_time": "2020-10-07T13:06:25.017569", "exception": false, "start_time": "2020-10-07T13:06:24.793252", "status": "completed"} tags=[]
ss_lower_range,ss_upper_range = get_outliers_range(mc.shoe_size)
#print(ss_lower_range,ss_upper_range)
mc.drop(mc[(mc.shoe_size < ss_lower_range) | (mc.shoe_size > ss_upper_range)].index, axis=0, inplace=True) # found 390 observations
plot_outlier(mc.shoe_size)
# + id="lloay3tqsxJC" papermill={"duration": 0.046937, "end_time": "2020-10-07T13:06:25.111506", "exception": false, "start_time": "2020-10-07T13:06:25.064569", "status": "completed"} tags=[]
# + [markdown] id="Uh4iGqygsxJC" papermill={"duration": 0.04623, "end_time": "2020-10-07T13:06:25.204093", "exception": false, "start_time": "2020-10-07T13:06:25.157863", "status": "completed"} tags=[]
# # Different solutions to transform categorical variables to numeric ones.
#
# In real world datasets, variables (features) are often categorial, most often such variables are represented by strings. Most machine learning models, however, cannot process strings, they can only handle numerical values i.e. numbers. The categorial features therefore needs to be transformed to numerical values, but at the same time it is important not to change the meaning and interpretations of the values.
#
# To read more about transforming categorial features in different ways (there are several all with different weaknesses and strengths depending on the data), see for instance [here](https://pbpython.com/categorical-encoding.html) and [here](https://towardsdatascience.com/beyond-one-hot-17-ways-of-transforming-categorical-features-into-numeric-features-57f54f199ea4), also pandas have a dtype called category which can also be helpful, see documentation [here](https://pandas.pydata.org/pandas-docs/stable/user_guide/categorical.html).
#
# + [markdown] id="tbqzWpRfsxJC" papermill={"duration": 0.046763, "end_time": "2020-10-07T13:06:25.297343", "exception": false, "start_time": "2020-10-07T13:06:25.250580", "status": "completed"} tags=[]
# ### Example 1:
#
# Applied to feature *Cup size* - used open source data to convert measurements into numerical data
# + [markdown] id="sEnkNeMPsxJC" papermill={"duration": 0.046726, "end_time": "2020-10-07T13:06:25.391285", "exception": false, "start_time": "2020-10-07T13:06:25.344559", "status": "completed"} tags=[]
# source taken from https://www.blitzresults.com/en/bra-size/
#
# Adding two new columns for the feture "cup_size" in order to convert the measurements into centimeters and then imputing missing values with mean values.
# + id="EiPGJPy4sxJD" papermill={"duration": 0.060177, "end_time": "2020-10-07T13:06:25.498089", "exception": false, "start_time": "2020-10-07T13:06:25.437912", "status": "completed"} tags=[]
def convert_cup_size_to_cms(cup_size_code):
if cup_size_code == 'aa':
return 10, 11
if cup_size_code == 'a':
return 12, 13
if cup_size_code == 'b':
return 14, 15
if cup_size_code == 'c':
return 16, 17
if cup_size_code == 'd':
return 18, 19
if cup_size_code == 'dd/e':
return 20, 21
if cup_size_code == 'ddd/f':
return 22, 23
if cup_size_code == 'dddd/g':
return 24, 25
if cup_size_code == 'h':
return 26, 27
if cup_size_code == 'i':
return 28, 29
if cup_size_code == 'j':
return 30, 31
if cup_size_code == 'k':
return 32, 33
else:
return str('unknown')
# + id="fj5RWuQTsxJD" papermill={"duration": 0.09651, "end_time": "2020-10-07T13:06:25.641808", "exception": false, "start_time": "2020-10-07T13:06:25.545298", "status": "completed"} tags=[]
mc['cup_size_in_cms'] = mc.cup_size.apply(convert_cup_size_to_cms)
mc.head()
# + id="Pschzw4wsxJD" papermill={"duration": 0.280551, "end_time": "2020-10-07T13:06:25.969661", "exception": false, "start_time": "2020-10-07T13:06:25.689110", "status": "completed"} tags=[]
def split_cup_size_data(data, index):
if data.lower() == 'unknown':
return 0
value = data.replace('(','').replace(')','').replace(',','')
return value.split()[index]
mc['cup_size_start_in_cms'] = mc.cup_size_in_cms.astype(str).apply(lambda x : split_cup_size_data(x, 0))
mc['cup_size_end_in_cms'] = mc.cup_size_in_cms.astype(str).apply(lambda x : split_cup_size_data(x, 1))
mc.head()
# + id="vwjtJ6DisxJD" papermill={"duration": 0.078579, "end_time": "2020-10-07T13:06:26.096410", "exception": false, "start_time": "2020-10-07T13:06:26.017831", "status": "completed"} tags=[]
mc['cup_size_start_in_cms'] = mc.cup_size_start_in_cms.astype('int')
mc['cup_size_end_in_cms'] = mc.cup_size_end_in_cms.astype('int')
# missing values imputation with mean
mc['cup_size_start_in_cms'] = mc.cup_size_start_in_cms.mask(mc.cup_size_start_in_cms==0).fillna(value=mc.cup_size_start_in_cms.mean())
mc['cup_size_end_in_cms'] = mc.cup_size_end_in_cms.mask(mc.cup_size_end_in_cms==0).fillna(value=mc.cup_size_end_in_cms.mean())
# + [markdown] id="6tDwjOZOsxJD" papermill={"duration": 0.067577, "end_time": "2020-10-07T13:06:26.232612", "exception": false, "start_time": "2020-10-07T13:06:26.165035", "status": "completed"} tags=[]
# > lets double check the NaN values imputations for the newly added features
# + id="GZDJRoy6sxJD" papermill={"duration": 0.138601, "end_time": "2020-10-07T13:06:26.438838", "exception": false, "start_time": "2020-10-07T13:06:26.300237", "status": "completed"} tags=[]
mc[mc.cup_size.isnull()]
# + id="mrSeGsq0sxJE" papermill={"duration": 0.07683, "end_time": "2020-10-07T13:06:26.564872", "exception": false, "start_time": "2020-10-07T13:06:26.488042", "status": "completed"} tags=[]
# drop the columns which are used for reference
mc = mc.drop(['cup_size', 'cup_size_in_cms'], axis = 1)
mc.reset_index(drop=True, inplace=True)
# + [markdown] id="lkgbhUxNsxJE" papermill={"duration": 0.047982, "end_time": "2020-10-07T13:06:26.661074", "exception": false, "start_time": "2020-10-07T13:06:26.613092", "status": "completed"} tags=[]
# #Example 2 Shoe_with
# + [markdown] id="n-DQrkdAsxJE" papermill={"duration": 0.04782, "end_time": "2020-10-07T13:06:26.757042", "exception": false, "start_time": "2020-10-07T13:06:26.709222", "status": "completed"} tags=[]
# > lets try to see the visualization for categorical data against the dependent feature fit
#
# + id="AZC6dh3fsxJE" papermill={"duration": 0.058542, "end_time": "2020-10-07T13:06:26.863748", "exception": false, "start_time": "2020-10-07T13:06:26.805206", "status": "completed"} tags=[]
def countplot_wrt_target(indipendent_features, df):
plt.figure(figsize=(28, 10))
for loc, feature in enumerate(indipendent_features):
ax = plt.subplot(1, 3, loc+1)
ax.set_xlabel('{}'.format(feature), fontsize=10)
chart = sns.countplot(x=df[feature], hue=df.fit)
chart.set_xticklabels(chart.get_xticklabels(), rotation=90)
return None
# + id="WlytbsXLsxJE" papermill={"duration": 1.124155, "end_time": "2020-10-07T13:06:28.037536", "exception": false, "start_time": "2020-10-07T13:06:26.913381", "status": "completed"} tags=[]
countplot_wrt_target(['category', 'length', 'quality'], mc)
# + [markdown] id="X2jQ81OCsxJE" papermill={"duration": 0.07125, "end_time": "2020-10-07T13:06:28.180349", "exception": false, "start_time": "2020-10-07T13:06:28.109099", "status": "completed"} tags=[]
# ## Example 2:
# Applied to feature *shoe_width* : used open source data to identify shoe width based on shoe size
#
# Reference link : https://images-na.ssl-images-amazon.com/images/I/71u90X9oX3S.pdf
# + id="nwYSHPe8sxJE" papermill={"duration": 0.086999, "end_time": "2020-10-07T13:06:28.338222", "exception": false, "start_time": "2020-10-07T13:06:28.251223", "status": "completed"} tags=[]
# fill NaN with average shoe width category (this is just an assumption)
mc.shoe_width = mc.shoe_width.fillna('average')
# + id="Ce1UU83-sxJF" papermill={"duration": 0.420654, "end_time": "2020-10-07T13:06:28.807750", "exception": false, "start_time": "2020-10-07T13:06:28.387096", "status": "completed"} tags=[]
# Use above chart to convert shoe width data such as 'wide','average','narrow' to inches
mc['shoe_width_in_inches'] = np.where(((mc.shoe_size >= 5) & (mc.shoe_size < 5.5)) & (mc.shoe_width == 'narrow') , 2.81,
np.where(((mc.shoe_size >= 5) & (mc.shoe_size < 5.5)) & (mc.shoe_width == 'average') , 3.19,
np.where(((mc.shoe_size >= 5) & (mc.shoe_size < 5.5)) & (mc.shoe_width == 'wide') , 3.56,
np.where(((mc.shoe_size >= 5.5) & (mc.shoe_size < 6)) & (mc.shoe_width == 'narrow') , 2.87,
np.where(((mc.shoe_size >= 5.5) & (mc.shoe_size < 6)) & (mc.shoe_width == 'average') , 3.25,
np.where(((mc.shoe_size >= 5.5) & (mc.shoe_size < 6)) & (mc.shoe_width == 'wide') , 3.62,
np.where(((mc.shoe_size >= 6) & (mc.shoe_size < 6.5)) & (mc.shoe_width == 'narrow') , 2.94,
np.where(((mc.shoe_size >= 6) & (mc.shoe_size < 6.5)) & (mc.shoe_width == 'average') , 3.31,
np.where(((mc.shoe_size >= 6) & (mc.shoe_size < 6.5)) & (mc.shoe_width == 'wide') , 3.69,
np.where(((mc.shoe_size >= 6.5) & (mc.shoe_size < 7)) & (mc.shoe_width == 'narrow') , 3,
np.where(((mc.shoe_size >= 6.5) & (mc.shoe_size < 7)) & (mc.shoe_width == 'average') , 3.37,
np.where(((mc.shoe_size >= 6.5) & (mc.shoe_size < 7)) & (mc.shoe_width == 'wide') , 3.75,
np.where(((mc.shoe_size >= 7) & (mc.shoe_size < 7.5)) & (mc.shoe_width == 'narrow') , 3.06,
np.where(((mc.shoe_size >= 7) & (mc.shoe_size < 7.5)) & (mc.shoe_width == 'average') , 3.44,
np.where(((mc.shoe_size >= 7) & (mc.shoe_size < 7.5)) & (mc.shoe_width == 'wide') , 3.81,
np.where(((mc.shoe_size >= 7.5) & (mc.shoe_size < 8)) & (mc.shoe_width == 'narrow') , 3.12,
np.where(((mc.shoe_size >= 7.5) & (mc.shoe_size < 8)) & (mc.shoe_width == 'average') , 3.5,
np.where(((mc.shoe_size >= 7.5) & (mc.shoe_size < 8)) & (mc.shoe_width == 'wide') , 3.87,
np.where(((mc.shoe_size >= 8) & (mc.shoe_size < 8.5)) & (mc.shoe_width == 'narrow') , 3.19,
np.where(((mc.shoe_size >= 8) & (mc.shoe_size < 8.5)) & (mc.shoe_width == 'average') , 3.56,
np.where(((mc.shoe_size >= 8) & (mc.shoe_size < 8.5)) & (mc.shoe_width == 'wide') , 3.94,
np.where(((mc.shoe_size >= 8.5) & (mc.shoe_size < 9)) & (mc.shoe_width == 'narrow') , 3.25,
np.where(((mc.shoe_size >= 8.5) & (mc.shoe_size < 9)) & (mc.shoe_width == 'average') , 3.62,
np.where(((mc.shoe_size >= 8.5) & (mc.shoe_size < 9)) & (mc.shoe_width == 'wide') , 4,
np.where(((mc.shoe_size >= 9) & (mc.shoe_size < 9.5)) & (mc.shoe_width == 'narrow') , 3.37,
np.where(((mc.shoe_size >= 9) & (mc.shoe_size < 9.5)) & (mc.shoe_width == 'average') , 3.69,
np.where(((mc.shoe_size >= 9) & (mc.shoe_size < 9.5)) & (mc.shoe_width == 'wide') , 4.06,
np.where(((mc.shoe_size >= 9.5) & (mc.shoe_size < 10)) & (mc.shoe_width == 'narrow') , 3.37,
np.where(((mc.shoe_size >= 9.5) & (mc.shoe_size < 10)) & (mc.shoe_width == 'average') , 3.75,
np.where(((mc.shoe_size >= 9.5) & (mc.shoe_size < 10)) & (mc.shoe_width == 'wide') , 4.12,
np.where(((mc.shoe_size >= 10) & (mc.shoe_size < 10.5)) & (mc.shoe_width == 'narrow') , 3.44,
np.where(((mc.shoe_size >= 10) & (mc.shoe_size < 10.5)) & (mc.shoe_width == 'average') , 3.75,
np.where(((mc.shoe_size >= 10) & (mc.shoe_size < 10.5)) & (mc.shoe_width == 'wide') , 4.19,
np.where(((mc.shoe_size >= 10.5) & (mc.shoe_size < 11)) & (mc.shoe_width == 'narrow') , 3.5,
np.where(((mc.shoe_size >= 10.5) & (mc.shoe_size < 11)) & (mc.shoe_width == 'average') , 3.87,
np.where(((mc.shoe_size >= 10.5) & (mc.shoe_size < 11)) & (mc.shoe_width == 'wide') , 4.19,
np.where(((mc.shoe_size >= 11) & (mc.shoe_size < 12)) & (mc.shoe_width == 'narrow') , 3.56,
np.where(((mc.shoe_size >= 11) & (mc.shoe_size < 12)) & (mc.shoe_width == 'average') , 3.94,
np.where(((mc.shoe_size >= 11) & (mc.shoe_size < 12)) & (mc.shoe_width == 'wide') , 4.19,
np.nan)))))))))))))))))))))))))))))))))))))))
# + id="n67G9iCqsxJF" papermill={"duration": 0.076844, "end_time": "2020-10-07T13:06:28.938463", "exception": false, "start_time": "2020-10-07T13:06:28.861619", "status": "completed"} tags=[]
# drop the refrence colum shoe_width
mc.drop(['shoe_width'], axis=1, inplace=True)
# + [markdown] id="R4qWL5vssxJF" papermill={"duration": 0.072428, "end_time": "2020-10-07T13:06:29.081892", "exception": false, "start_time": "2020-10-07T13:06:29.009464", "status": "completed"} tags=[]
# # Example 3:
#
# Applied to features *lenght & category* Using one-hot encoding to change categorial data to numeric
# + [markdown] id="xpj23rN9sxJF" papermill={"duration": 0.071255, "end_time": "2020-10-07T13:06:29.230869", "exception": false, "start_time": "2020-10-07T13:06:29.159614", "status": "completed"} tags=[]
# > One hot encoding is a common way to change categorial (often string) data to numeric data without changing the scale of the feature. This type of transformation is suitable when the data is in nominal scale i.e. don't have any order.
#
#
# + id="NQuouTp-sxJF" papermill={"duration": 0.094769, "end_time": "2020-10-07T13:06:29.396560", "exception": false, "start_time": "2020-10-07T13:06:29.301791", "status": "completed"} tags=[]
# lets replace NaN values with unknown for the feature length
mc.length = mc.length.fillna('unknown')
# + id="bfJzy8rFsxJF" papermill={"duration": 0.25441, "end_time": "2020-10-07T13:06:29.700910", "exception": false, "start_time": "2020-10-07T13:06:29.446500", "status": "completed"} tags=[]
# apply one hot encoding using dummies
length_dummies = pd.get_dummies(mc['length'])
length_dummies.columns = ['just_right','slightly_long','very_short','slightly_short','very_long', 'length_unkown']
category_dummies = pd.get_dummies(mc['category'])
category_dummies.columns = ['new','dresses','wedding','sale','tops', 'bottoms','outerwear']
model_input_df = pd.concat([mc, length_dummies,category_dummies], axis = 1)
model_input_df.drop(['length'], axis=1, inplace=True)
model_input_df.drop(['category'], axis=1, inplace=True)
# target variable
fit = {'small':0, 'fit':1, 'large':2}
model_input_df['fit'] = model_input_df['fit'].map(fit)
# + id="og21gKO5sxJF" papermill={"duration": 0.097724, "end_time": "2020-10-07T13:06:29.848380", "exception": false, "start_time": "2020-10-07T13:06:29.750656", "status": "completed"} tags=[]
# since there is no value add to the features like item_id , user_id and user_name
model_input_df.drop(['item_id'], axis=1, inplace=True)
model_input_df.drop(['user_id'], axis=1, inplace=True)
model_input_df.drop(['user_name'], axis=1, inplace=True)
model_input_df.head()
# -
# # Example 4:
#
# Applied to feature *fit* Change into ordinal (dtype categorial).
#
mc.fit = mc.fit.astype('category').cat.as_ordered()
| Data_precossesing_modcloth.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Boston house data
# Example taken from [here](https://machinelearningmastery.com/regression-tutorial-keras-deep-learning-library-python/)
# +
import pandas
from keras.models import Sequential
from keras.layers import Dense
from keras.wrappers.scikit_learn import KerasRegressor
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import KFold
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import Pipeline
from sklearn.model_selection import train_test_split
import matplotlib.pylab as plt
from pylab import rcParams
rcParams['figure.figsize'] = 15, 10
# -
# ### Load data
df = pandas.read_csv("housing.data.txt", delim_whitespace=True, header=None)
dataset = df.values
# ### split into input (X) and output (Y) variables
X = dataset[:,0:13]
Y = dataset[:,13]
# ### define base model
def baseline_model():
# create model
model = Sequential()
model.add(Dense(13, input_dim=13, kernel_initializer='normal', activation='relu'))
model.add(Dense(1, kernel_initializer='normal'))
# Compile model
model.compile(loss='mean_squared_error', optimizer='adam')
return model
def wider_model():
# create model
model = Sequential()
model.add(Dense(20, input_dim=13, kernel_initializer='normal', activation='relu'))
model.add(Dense(1, kernel_initializer='normal'))
# Compile model
model.compile(loss='mean_squared_error', optimizer='adam')
return model
# +
estimators = []
estimators.append(('standardize', StandardScaler()))
estimators.append(('mlp', KerasRegressor(build_fn=wider_model, epochs=100, batch_size=5, verbose=0)))
pipeline = Pipeline(estimators)
# -
# ## Cross validation
# +
kfold = KFold(n_splits=10)
results = cross_val_score(pipeline, X, Y, cv=kfold)
print("Results: %.2f (%.2f) MSE" % (results.mean(), results.std()))
# -
df.hist(bins=30);
X.shape
X_train, X_test, y_train, y_test = train_test_split(X,Y,test_size=0.33)
pipeline.fit(X=X_train, y=y_train)
y_pred = pipeline.predict(X_test)
from sklearn.metrics import r2_score
r2_score(y_test, y_pred)
linear_estimator = LinearRegression()
linear_estimator.fit(X_train, y_train)
y_pred_linear = linear_estimator.predict(X_test)
linear_estimator.score(X_test, y_test)
r2_score(y_test, y_pred_linear)
pipeline.score(X_test, y_test)
fig,ax = plt.subplots()
ax.plot(y_test, y_pred, '.', label='keras')
ax.plot(y_test, y_pred_linear, '.', label='linear')
ax.set_xlabel('Actual price')
ax.set_ylabel('Predicted price');
| 07_boston_house_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # How to build up a labelled dataset for unet
#
#
# Here we introduce how to build up dataset for unet.We use one event waveform for example.
#
# # load modules
# #!/usr/bin/env python
import os,glob,re
import numpy as np
import matplotlib
#matplotlib.use('Agg')
import matplotlib.pyplot as plt
import scipy.stats as stats
from quakenet.data_pipeline_unet import DataWriter
import tensorflow as tf
from obspy.core import read,Stream
from quakenet.data_io import load_catalog
from obspy.core.utcdatetime import UTCDateTime
import fnmatch,math
import json
# # useful subroutines
# +
def preprocess_stream(stream):
stream = stream.detrend('constant')
##add by mingzhao,2017/12/2
stream =stream.filter('bandpass', freqmin=0.5, freqmax=20)
##########
return stream
def write_json(metadata,output_metadata):
with open(output_metadata, 'w') as outfile:
json.dump(metadata, outfile)
def normfun(x, mu, sigma):
pdf = np.exp(-((x - mu) ** 2) / (2 * sigma ** 2)) / (sigma * np.sqrt(2 * np.pi))
return pdf
# -
# # load event data
# +
#this data has a time range [tp-5,ts+15],tp,ts is manual picks by expert,which is cutted from continous waveform previously
stream_file = "2012-08-30T00:14:25.105000Z.BJ.BBS.00.BHZ.D.SAC"
output_name = "test.tfrecords"
output_path = os.path.join('.', output_name)
writer = DataWriter(output_path)
stream_file1 = re.sub('HZ.D.SAC', 'HE.D.SAC', str(stream_file))
stream_file2 = re.sub('HZ.D.SAC', 'HN.D.SAC', str(stream_file))
stream = read(stream_file)
stream += read(stream_file1)
stream += read(stream_file2)
print ('+ Preprocessing stream',stream)
stream = preprocess_stream(stream)
start_date = stream[0].stats.starttime
end_date = stream[-1].stats.endtime
print("-- Start Date={}, End Date={}".format(start_date, end_date))
# -
# ### resample to 100 sample points,trim it to the window_size,use 0 to fill if the data is not as long as window_size
x = np.random.randint(0, 4)
##the cut start randomly before 1~5s of the p arrival,and end after 30s
window_size=30
st_event = stream.resample(100).trim(start_date+x, start_date+x+window_size,pad=True, fill_value=0.0).copy()
print (st_event)
st_event.plot()
# ### necessary parameters for labelling
n_samples = len(st_event[0].data)
sample_rate = st_event[0].stats.sampling_rate
n_pts = sample_rate * window_size + 1
cluster_id_p = 5-x
cluster_id_s = end_date - start_date-x-15
# +
# p,s and noise labelling
# +
# for p picks
label_obj = st_event.copy()
label_obj[0].data[...] = 1
label_obj[1].data[...] = 0
label_obj[2].data[...] = 0
u1 = cluster_id_p * sample_rate # mean value miu
lower = int(u1 - 0.5*sample_rate)
upper = int(u1 + 0.5*sample_rate)
label_obj[1].data[lower:upper] = 1
# for s pick
u2 = cluster_id_s * sample_rate # mean value miu
lower2, upper2 = int(u2 - sample_rate), int(u2 + sample_rate)
try:
label_obj[2].data[lower2:upper2] = 2
except:
nnn = int(n_samples) - int(u2 + sample_rate)
#print (nnn, n_samples)
label_obj[2].data[lower2:n_samples] = 2
label_obj.normalize()
label_obj[0].data = label_obj[0].data - label_obj[1].data - label_obj[2].data
writer.write(st_event.copy().normalize(), label_obj)
# -
# ### Plot one trace and its label
# +
traces = Stream()
traces += st_event[0]
label_obj[0].stats.channel="N"
label_obj[1].stats.channel="P"
label_obj[2].stats.channel="S"
traces += label_obj
traces.normalize().plot()
# -
# # (optional) Gaussian label
# +
label_obj = st_event.copy()
label_obj[0].data[...] = 1
label_obj[1].data[...] = 0
label_obj[2].data[...] = 0
u1 = cluster_id_p * sample_rate # mean value miu
lower = int(u1 - 0.2 * sample_rate)
upper = int(u1 + 0.2 * sample_rate)
mu, sigma = u1, 0.1
X = stats.truncnorm((lower - mu) / sigma, (upper - mu) / sigma, loc=mu, scale=sigma)
n, bins, patches = plt.hist(X.rvs(10000), int(0.4*sample_rate), normed=1, alpha=0.5)
label_obj[1].data[lower:upper] = n[0:int(0.4*sample_rate)]
# for s pick
u2 = cluster_id_s * sample_rate # mean value miu
lower2, upper2 = int(u2 - 0.2*sample_rate), int(u2 + 0.2*sample_rate)
mu, sigma = u2, 0.1
X = stats.truncnorm((lower2 - mu) / sigma, (upper2 - mu) / sigma, loc=mu, scale=sigma)
n1, bins1, patches1 = plt.hist(X.rvs(10000), int(0.4 * sample_rate), normed=1, alpha=0.5)
try:
label_obj[2].data[int(u2 - 0.2*sample_rate):int(u2 + 0.2*sample_rate)] = n1[0:int(0.4 * sample_rate)]
# label_obj.data[int(u2 - sample_rate):int(u2 + sample_rate)] =2
except:
nnn = int(n_samples) - int(u2 + 0.4*sample_rate)
print nnn, n_samples
label_obj[2].data[int(u2 - 0.2*sample_rate):n_samples] = n1[0:nnn]
label_obj.normalize()
label_obj[0].data = label_obj[0].data - label_obj[1].data - label_obj[2].data
writer.write(st_event.copy().normalize(), label_obj)
# -
traces = Stream()
traces += st_event[0]
label_obj[0].stats.channel="N"
label_obj[1].stats.channel="P"
label_obj[2].stats.channel="S"
traces += label_obj
traces.normalize().plot()
| build_dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise Solution
import pandas as pd
from sklearn.datasets import load_wine
wine = load_wine()
dir(wine)
df = pd.DataFrame(wine.data, columns=wine.feature_names)
df.head()
df['target']=wine.target
df.head()
wine.target_names
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(df.drop('target',axis='columns'),df.target,test_size=0.2)
X_train.shape
X_test[:3]
y_test[:5]
# ### Using Gaussian Naive Bayes
from sklearn.naive_bayes import GaussianNB
Gmodel = GaussianNB()
Gmodel.fit(X_train,y_train)
Gmodel.score(X_test,y_test) # Accuracy
Gmodel.predict(X_test[0:10])
# ### Using Multinomial Naive Bayes
from sklearn.naive_bayes import MultinomialNB
Mmodel = MultinomialNB()
Mmodel.fit(X_train,y_train)
Mmodel.score(X_test,y_test) # Accuracy
Gmodel.predict(X_test[0:10])
# **Gaussian Naive Bayes gave optimal result so far for this dataset**
| Machine Learning/14. Naive Bayes 3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# ### Density
# -
import os
import sys
module_path = os.path.abspath(os.path.join('..\\..'))
if module_path not in sys.path:
sys.path.append(module_path)
import civilpy
rho = civilpy.general.density.water / civilpy.general.physics.gravity
round(rho.to('slugs/ft**3'), 3)
round(rho.to('kg/m**3'), 3)
# ### Specific Gravity
# +
co2_gas_constant = 35.1 * (civilpy.unit.ft*civilpy.unit.lb) / (civilpy.unit.lbf * civilpy.unit.rankine)
co2_pressure = 20 * civilpy.unit.lb / civilpy.unit.inch ** 2
Q = civilpy.unit.Quantity
temp_i = Q(150, civilpy.unit.delta_degF)
temp_f = Q(460, civilpy.unit.degR)
density_co2 = (co2_pressure.to('lb/ft**2') / (co2_gas_constant * (temp_i + temp_f)))
sg = civilpy.water_resources.hydraulics.specific_gravity(density_co2, phase='gas')
round(sg, 2)
# -
# ### Specific Weight
density = 1.95 * civilpy.unit.slug / civilpy.unit.ft ** 3
SW = civilpy.general.specific_weight(density)
round(SW.to('lbf / ft ** 3'), 2)
# ### Manometer
# +
rhom = civilpy.general.density.mercury
rho2 = civilpy.general.density.water
hw = 120 * civilpy.unit.inch
hm = 17 * civilpy.unit.inch
delta_pressure = civilpy.water_resources.manometer_pressure(rhom, hm, rho2=rho2, h2=hw)
round(delta_pressure.to('psi'), 2)
# +
h = 100 * civilpy.unit.ft
p = civilpy.water_resources.pressure_at_depth(h)
p.to('psi')
# -
rho = 997.0474 * civilpy.unit.kg / civilpy.unit.m ** 3
rho.to('lb/ft**3')
civilpy.general.density.water
civilpy.general.physics.gravity / civilpy.general.physics.gravity_c
| tests/notebooks/water_resources_test_notebook.ipynb |
% ---
% jupyter:
% jupytext:
% text_representation:
% extension: .m
% format_name: light
% format_version: '1.5'
% jupytext_version: 1.14.4
% kernelspec:
% display_name: Octave
% language: octave
% name: octave
% ---
% some housekeeping stuff
register_graphics_toolkit("gnuplot");
available_graphics_toolkits();
graphics_toolkit("gnuplot")
clear
% end of housekeeping
% # REE (rare earth elements) for nutrient removal
%
% Rare earth elements form sparingly soluble salts with phosphate. Their use in wastewater treatment for nutrient removal is being explored.
%
% Phosphate is a regulated chemical at the "end of pipe" of a wastewater treatment plant because excess phosphate can cause eutrophication in the receiving waters.
%
% Europium is a rare earth element that forms low solubility precipitates with phosphate.
%
% $EuPO4(s) = Eu^{3+}+ PO_4^{3-}$ Ksp=10$^{-24.0}$
%
% So let's model for a fixed pH the removal of phosphate from solution after addition of increasing amounts of europium nitrate (europium nitrate is soluble).
%
% We derived the equations in class. Assume total phosphate of 10 $\mu$M.
%
%
%
% +
%plot -s 600,500 -f 'svg'
PT=1e-5; EuT=[1e-7 1e-6:2e-6:10e-5]; pH=5.5;
H=10^-pH;
Ka1=7.5*10^-3; Ka2=6.2*10^-8; Ka3=2.14*10^-13; %mol/l
Ksp=10^-24;
for i=1:size(EuT,2)
% first test for supersaturation
PO4tst=(PT*Ka1*Ka2*Ka3)./(Ka1*Ka2*Ka3+Ka1*Ka2*H+Ka1*H^2+H^3);
Eutst=EuT(i);
Qsp=Eutst*PO4tst;
SItst=Qsp/Ksp;
if SItst<1; %disp('undersaturated');
EuPO4s(i)=0; end
if SItst>=1
%disp('supersaturated');
a=1+H/Ka3+(H^2)/(Ka3*Ka2)+(H^3)/(Ka3*Ka2*Ka1);
b=EuT(i)-PT;
c=-Ksp;
t=roots([a b c]);
t(imag(t)==0); %sets any imaginary roots to zero
t=t(t>0); %t=t(t<EuT(i));
PO4(i)=t; Eu(i)=Ksp/PO4(i); EuPO4s(i)=EuT(i)-Eu(i);
end
end
plot(EuT*1e6,EuPO4s*1e6,'ko','markerfacecolor','b','markersize',4)
xlabel('EuT (\muM)'); ylabel('EuPO4s (\muM)')
set(gca,'linewidth',1.5)
| .ipynb_checkpoints/EuPO4s_wastewater_treatment-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# name: python3
# ---
# +
#all credit goes to <NAME> (PhantomJL, stemaway repository) for working program
import pandas as pd
data = pd.read_csv('combined_csv.csv')
data.head()
# -
data["Leading Comment"][3]
data.drop(['Unnamed: 0','Commenters'],axis = 1,inplace=True)
data.columns
# ## Strategy 1
# Lower case all the words
#
# Replaced by space these [/(){}[]\'\“\”\’|@,;]
#
# Removed these [^0-9a-z #+_]
import logging
import pandas as pd
import numpy as np
from numpy import random
import gensim
import nltk
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer
from sklearn.metrics import accuracy_score, confusion_matrix
import matplotlib.pyplot as plt
from nltk.corpus import stopwords
import re
from bs4 import BeautifulSoup
# %matplotlib inline
df = data.copy()
df.info()
df['post'] = df['Topic Title'] + ' ' + df['Leading Comment'] + ' ' + df['Other Comments'] + ' ' + df['Tags']
df['post'].head()
print(df['post'].apply(lambda x: len(str(x).split(' '))).sum())
df['Category'].unique()
df = df[df.Category != ' 1990. My husband and I rented a spiffy new Cadillac in Indiana with plans to pick up my parents in Erie']
df['Category'].unique()
df.Category.value_counts()
df.loc[df.Category == 'Buying/Selling', 'Category'] = "Buying/Selling & Ask Someone Who Owns One"
df.loc[df.Category == 'Ask Someone Who Owns One', 'Category'] = "Buying/Selling & Ask Someone Who Owns One"
df.Category.value_counts()
df['Category'].unique()
df["post"].isnull().sum()
df = df[df['post'].notna()]
df["post"].isnull().sum()
df["post"].head()
my_categories = ['Buying/Selling & Ask Someone Who Owns One', 'Safety',
'Driving/Drivers', 'Site Feedback', 'Auto Type/Class',
'Maintenance/Repairs', 'The Show', 'Power/Fuel',
'General Discussion']
plt.figure(figsize=(10,4))
df.Category.value_counts().plot(kind='bar');
def print_plot(index):
example = df[df.index == index][['post', 'Category']].values[0]
if len(example) > 0:
print(example[0])
print('Category:', example[1])
print_plot(10)
# Text Pre-processing
test_df = df.copy()
test_df['post'] = test_df['post'].astype(str)
# +
REPLACE_BY_SPACE_RE = re.compile('[/(){}\[\]\'\“\”\’\|@,;]')
BAD_SYMBOLS_RE = re.compile('[^0-9a-z #+_]')
#STOPWORDS = set(stopwords.words('english'))
def clean_text(text):
"""
text: a string
return: modified initial string
"""
text = re.sub(r'^https?:\/\/.*[\r\n]*', '', text, flags=re.MULTILINE)
text = BeautifulSoup(text, "lxml").text # HTML decoding
text = text.lower() # lowercase text
text = REPLACE_BY_SPACE_RE.sub(' ', text)# replace REPLACE_BY_SPACE_RE symbols by space in text
#text = text.replace(r'\n', '')
text = BAD_SYMBOLS_RE.sub('', text) # delete symbols which are in BAD_SYMBOLS_RE from text
#text = ''.join(word for word in text if word not in punct) # remove punctuation
#text = ' '.join(word for word in text.split() if word not in STOPWORDS) # delete stopwors from text
return text
test_df['post'] = test_df['post'].apply(clean_text)
# -
def test_print_plot(index):
example = test_df[test_df.index == index][['post', 'Category']].values[0]
if len(example) > 0:
print(example[0])
print('Category:', example[1])
test_print_plot(10)
test_df['post'].apply(lambda x: len(x.split(' '))).sum()
print("lost info",4765518-4286502)
# Modeling the data
X = test_df.post
y = test_df.Category
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state = 42)
X_train.head()
y_train.head()
# ## Naive Bayes Classifier for Multinomial Models
# CountVectorizer + TF-IDFTransformer + MultinomialNB
# +
from sklearn.naive_bayes import MultinomialNB
from sklearn.pipeline import Pipeline
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.metrics import classification_report
def Naive_Bayes_Classifier(X_train, X_test, y_train, y_test):
nb = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', MultinomialNB()),
])
nb.fit(X_train, y_train)
y_pred = nb.predict(X_test)
print('accuracy %s' % accuracy_score(y_pred, y_test))
print(classification_report(y_test, y_pred,target_names=my_categories))
return accuracy_score(y_pred, y_test)
res1311 = Naive_Bayes_Classifier(X_train, X_test, y_train, y_test)
# -
# ## Linear Support Vector Machine
# CountVectorizer + TF-IDFTransformer + SGDClassifier
# +
from sklearn.linear_model import SGDClassifier
def Linear_Support_Vector(X_train, X_test, y_train, y_test):
sgd = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', SGDClassifier(loss='hinge', penalty='l2',alpha=1e-3, random_state=42, max_iter=5, tol=None)),
])
sgd.fit(X_train, y_train)
y_pred = sgd.predict(X_test)
print('accuracy %s' % accuracy_score(y_pred, y_test))
print(classification_report(y_test, y_pred,target_names=my_categories))
return accuracy_score(y_pred, y_test)
res1321 = Linear_Support_Vector(X_train, X_test, y_train, y_test)
# -
# ## Logistic Regression
# CountVectorizer + TF-IDFTransformer + Logistic Regression
# +
from sklearn.linear_model import LogisticRegression
def Logistic_Regression(X_train, X_test, y_train, y_test):
logreg = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', LogisticRegression(n_jobs=1, C=1e5)),
])
logreg.fit(X_train, y_train)
y_pred = logreg.predict (X_test)
print('accuracy %s' % accuracy_score(y_pred, y_test))
print(classification_report(y_test, y_pred,target_names=my_categories))
return accuracy_score(y_pred, y_test)
res1331 = Logistic_Regression(X_train, X_test, y_train, y_test)
# -
# Decision Tree
# CountVectorizer + TF-IDFTransformer + DecisionTreeClassifier
# ## Decision Tree
# CountVectorizer + TF-IDFTransformer + DecisionTreeClassifier
# +
from sklearn.tree import DecisionTreeClassifier
def Decision_Tree(X_train, X_test, y_train, y_test):
dtree = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', DecisionTreeClassifier(random_state=0)),
])
dtree.fit(X_train, y_train)
y_pred = dtree.predict (X_test)
print('accuracy %s' % accuracy_score(y_pred, y_test))
print(classification_report(y_test, y_pred,target_names=my_categories))
return accuracy_score(y_pred, y_test)
res1341 = Decision_Tree(X_train, X_test, y_train, y_test)
# -
# ## Results
# Results of the previously trained models
# +
import pandas as pd
results = pd.DataFrame({'Model': ['Naive Bayes MultinomialNB', 'Linear SVM', 'Logistic Regression', 'Decision Tree'],
'Accuracy': [res1311, res1321, res1331, res1341]})
results.set_index('Model')
results.sort_values(by='Accuracy')
# -
# Cross Validation with linear SVM
# +
from sklearn.linear_model import SGDClassifier
from sklearn.model_selection import cross_val_score
import numpy as np
def Mean_cv_res_sgd(X_train,y_train):
sgd = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', SGDClassifier(loss='hinge', penalty='l2',alpha=1e-3, random_state=42, max_iter=5, tol=None)),
])
cv_res_sgd = cross_val_score(sgd, X_train, y_train, cv=10)
mean_cv_res_sgd = np.mean(cv_res_sgd)
return (mean_cv_res_sgd)
mean_cv_res_sgd = Mean_cv_res_sgd(X_train,y_train)
# -
print(mean_cv_res_sgd)
# ## Strategy 2
# Remove stop words (+ what was done in Strategy 1)
# +
# test_df2 = test_df.copy()
# +
# REPLACE_BY_SPACE_RE = re.compile('[/(){}\[\]\'\“\”\’\|@,;]')
# BAD_SYMBOLS_RE = re.compile('[^0-9a-z #+_]')
# STOPWORDS = set(stopwords.words('english'))
# def clean_text(text):
# """
# text: a string
# return: modified initial string
# """
# text = BeautifulSoup(text, "lxml").text # HTML decoding
# text = text.lower() # lowercase text
# text = REPLACE_BY_SPACE_RE.sub(' ', text) # replace REPLACE_BY_SPACE_RE symbols by space in text
# #text = BAD_SYMBOLS_RE.sub('', text) # delete symbols which are in BAD_SYMBOLS_RE from text
# #text = ''.join(word for word in text if word not in punct) # remove punctuation
# text = ' '.join(word for word in text.split() if word not in STOPWORDS) # delete stopwors from text
# return text
# test_df2['post'] = test_df2['post'].apply(clean_text)
# +
# def test_print_plot(index):
# example = test_df2[test_df2.index == index][['post', 'Category']].values[0]
# if len(example) > 0:
# print(example[0])
# print('Category:', example[1])
# test_print_plot(10)
# -
# ## Modeling the data
# +
# X = test_df2.post
# y = test_df2.Category
# X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state = 42)
# -
# ## Naive Bayes Classifier for Multinomial Models
# CountVectorizer + TF-IDFTransformer + MultinomialNB
# +
# res2311 = Naive_Bayes_Classifier(X_train, X_test, y_train, y_test)
# -
# ## Linear Support Vector Machine
# CountVectorizer + TF-IDFTransformer + SGDClassifier
# +
# res2321 = Linear_Support_Vector(X_train, X_test, y_train, y_test)
# -
# ## Logistic Regression
# CountVectorizer + TF-IDFTransformer + Logistic Regression
# +
# res2331 = Logistic_Regression(X_train, X_test, y_train, y_test)
# -
# ## Decision Tree
# CountVectorizer + TF-IDFTransformer + DecisionTreeClassifier
# +
# res2341 = Decision_Tree(X_train, X_test, y_train, y_test)
# +
# from sklearn.metrics import confusion_matrix
# import seaborn as sns
# conf_mat = confusion_matrix(y_test, y_pred)
# fig, ax = plt.subplots(figsize=(10,10))
# sns.heatmap(conf_mat, annot=True, fmt='d',
# xticklabels=my_categories, yticklabels=my_categories)
# plt.ylabel('Actual')
# plt.xlabel('Predicted')
# plt.show()
# -
# ## Results
# Results of the previously trained models
# +
# import pandas as pd
# results = pd.DataFrame({'Model': ['Naive Bayes MultinomialNB', 'Linear SVM', 'Logistic Regression', 'Decision Tree'],
# 'Accuracy': [res2311, res2321, res2331, res2341]})
# results.set_index('Model')
# results.sort_values(by='Accuracy')
# -
# ### Cross Validation with linear SVM
# +
# mean_cv_res_sgd2 = Mean_cv_res_sgd(X_train, y_train)
# print(mean_cv_res_sgd2)
# -
# ## Strategy 3
# text.replace(r'\n', ' ') + Strategy 2
# +
# test_df3 = test_df2.copy()
# +
# REPLACE_BY_SPACE_RE = re.compile('[/(){}\[\]\'\“\”\’\|@,;]')
# BAD_SYMBOLS_RE = re.compile('[^0-9a-z #+_]')
# STOPWORDS = set(stopwords.words('english'))
# def clean_text(text):
# """
# text: a string
# return: modified initial string
# """
# text = BeautifulSoup(text, "lxml").text # HTML decoding
# text = text.lower() # lowercase text
# text = REPLACE_BY_SPACE_RE.sub(' ', text) # replace REPLACE_BY_SPACE_RE symbols by space in text
# text = text.replace(r'\n', ' ')
# #text = BAD_SYMBOLS_RE.sub('', text) # delete symbols which are in BAD_SYMBOLS_RE from text
# #text = ''.join(word for word in text if word not in punct) # remove punctuation
# text = ' '.join(word for word in text.split() if word not in STOPWORDS) # delete stopwors from text
# return text
# test_df3['post'] = test_df3['post'].apply(clean_text)
# +
# def test_print_plot(index):
# example = test_df3[test_df3.index == index][['post', 'Category']].values[0]
# if len(example) > 0:
# print(example[0])
# print('Category:', example[1])
# test_print_plot(10)
# +
# res3311 = Naive_Bayes_Classifier(X_train, X_test, y_train, y_test)
# +
# res3321 = Linear_Support_Vector(X_train, X_test, y_train, y_test)
# +
# res3331 = Logistic_Regression(X_train, X_test, y_train, y_test)
# +
# res3341 = Decision_Tree(X_train, X_test, y_train, y_test)
# +
# import pandas as pd
# results = pd.DataFrame({'Model': ['Naive Bayes MultinomialNB', 'Linear SVM', 'Logistic Regression', 'Decision Tree'],
# 'Accuracy': [res3311, res3321, res3331, res3341]})
# results.set_index('Model')
# results.sort_values(by='Accuracy')
# -
# ## Cross Validation with linear SVM
# +
# X = test_df3.post
# y = test_df3.Category
# X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state = 42)
# +
# mean_cv_res_sgd3 = Mean_cv_res_sgd(X_train, y_train)
# print(mean_cv_res_sgd3)
# -
# ## Random Forest
# +
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import cross_val_score
import numpy as np
def Mean_cv_res_rf(X_train, y_train,m,n):
rf = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', RandomForestClassifier(n_estimators=n, max_depth=m, random_state=0)),
])
cv_res_rf = cross_val_score(rf, X_train, y_train, cv=10)
mean_cv_res_rf = np.mean(cv_res_rf)
return mean_cv_res_rf
# +
# print(Mean_cv_res_rf(X_train, y_train,10,300))
# +
# print(Mean_cv_res_rf(X_train, y_train,100,300))
# +
# print(Mean_cv_res_rf(X_train, y_train,500,300))
# +
# print(Mean_cv_res_rf(X_train, y_train,500,400))
# +
# print(Mean_cv_res_rf(X_train, y_train,1000,400))
# +
# print(Mean_cv_res_rf(X_train, y_train,500,500))
# -
def Mean_cv_res_tfidf(X_train, y_train,m,n):
rf = Pipeline([('vect', TfidfVectorizer()),
('tfidf', TfidfTransformer()),
('clf', RandomForestClassifier(n_estimators=n, max_depth=m, random_state=0)),
])
cv_res_rf_tfidf = cross_val_score(rf, X_train, y_train, cv=10)
mean_cv_res_rf_tfidf = np.mean(cv_res_rf_tfidf)
return mean_cv_res_rf_tfidf
print(Mean_cv_res_tfidf(X_train, y_train,500,500))
# +
# print(Mean_cv_res_tfidf(X_train, y_train,200,100))
# -
# Using TF-IDF as a vectorizer and transformer is slighly better than using CountVectorizer as a Vectorizer.
def Mean_cv_res_sgd_tfidf(X_train, y_train, max_iter):
sgd = Pipeline([('vect', TfidfVectorizer()),
('tfidf', TfidfTransformer()),
('clf', SGDClassifier(loss='hinge', penalty='l2',alpha=1e-3, random_state=42, max_iter=max_iter, tol=None)),
])
cv_res_sgd_tfidf = cross_val_score(sgd, X_train, y_train, cv=10)
mean_cv_res_sgd_tfidf = np.mean(cv_res_sgd_tfidf)
return mean_cv_res_sgd_tfidf
mean_cv_res_sgd_tfidf = Mean_cv_res_sgd_tfidf(X_train, y_train, 10)
mean_cv_res_sgd_tfidf = Mean_cv_res_sgd_tfidf(X_train, y_train, 100)
print(mean_cv_res_sgd_tfidf)
# ## XGBoost
import sys
# !{sys.executable} -m pip install xgboost
# +
import xgboost as xgb
xgb = Pipeline(steps=[('vect', TfidfVectorizer()),
('tfidf', TfidfTransformer()),
('xgboost', xgb.XGBClassifier(objective='multi:softmax'))])
cv_res_xgb_tfidf = cross_val_score(xgb, X_train, y_train, cv=10)
mean_cv_res_xgb_tfidf = np.mean(cv_res_xgb_tfidf)
# -
mean_cv_res_xgb_tfidf
xgb.fit(X_train, y_train)
y_pred = xgb.predict(X_test)
print('accuracy %s' % accuracy_score(y_pred, y_test))
res_xgb = accuracy_score(y_pred, y_test)
print(classification_report(y_test, y_pred,target_names=my_categories))
import seaborn as sns
conf_mat = confusion_matrix(y_test, y_pred)
fig, ax = plt.subplots(figsize=(10,10))
sns.heatmap(conf_mat, annot=True, fmt='d',
xticklabels=my_categories, yticklabels=my_categories)
plt.ylabel('Actual')
plt.xlabel('Predicted')
plt.show()
# ## Light GBM
import sys
# !{sys.executable} -m pip install lightgbm
# +
import lightgbm as lgbm
tfidf_vec = TfidfVectorizer(dtype=np.float32, sublinear_tf=True, use_idf=True, smooth_idf=True)
X_data_tfidf = tfidf_vec.fit_transform(df['post'])
X_train_tfidf = tfidf_vec.transform(X_train)
X_test_tfidf = tfidf_vec.transform(X_test)
clf_LGBM = lgbm.LGBMClassifier(objective='multiclass', verbose=-1, learning_rate=0.5, max_depth=20, num_leaves=50, n_estimators=120, max_bin=2000,)
clf_LGBM.fit(X_train_tfidf, y_train, verbose=-1)
predicted_LGBM = clf_LGBM.predict(X_test_tfidf)
# -
cv_res_lgbm_tfidf = cross_val_score(clf_LGBM, X_train_tfidf, y_train, cv=10)
mean_cv_res_lgbm_tfidf = np.mean(cv_res_lgbm_tfidf)
mean_cv_res_lgbm_tfidf
print('accuracy %s' % accuracy_score(predicted_LGBM, y_test))
res_lgbm = accuracy_score(predicted_LGBM, y_test)
print(classification_report(y_test, predicted_LGBM,target_names=my_categories))
# ## Other: Investigating Abbreviations
data = pd.read_csv('combined_csv.csv')
STOPWORDS = set(stopwords.words('english'))
STOPWORDSv2 = [word.upper() for word in STOPWORDS]
STOPWORDSv2
# +
def clean_data(text):
"""
text: a string
return: modified initial string
"""
text = BeautifulSoup(str(text), "lxml").text # HTML decoding
text = ' '.join(word for word in str(text).split() if word not in STOPWORDSv2) # delete stopwors from text
return text
data['Leading Comment'] = data['Leading Comment'].apply(clean_data)
# -
data['Leading Comment'].head()
# +
upcase_words = {}
regex = r"([A-Z]{3})"
for i, row in zip(range(500), data['Leading Comment'][:500]):
row = data['Leading Comment'].str.findall(regex)[i]
if len(row) != 0:
upcase_words[i] = row
# -
df_upword = pd.DataFrame(upcase_words.items(), columns=['Original Index', 'Upper Case Words'])
df_upword.head()
df_upword['Number of UpCase Words'] = df_upword['Upper Case Words'].apply(lambda x: len(x))
df_upword.head()
df_upword['Number of UpCase Words'].values.max()
df_upword['Number of UpCase Words'][df_upword['Number of UpCase Words'].values == 16]
df_upword['Number of UpCase Words'][0]
df_upword['Number of UpCase Words'][29]
df_upword['Upper Case Words'][0]
df_upword['Upper Case Words'][29]
def ABB(n):
abb = {}
for word in df_upword['Upper Case Words'][0]:
if word in abb.keys():
abb[word] += 1
else:
abb[word] = 1
return abb
ABB(0)
ABB(29)
| data-cleaning-simple-modeling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 9.2
# language: sage
# name: sagemath
# ---
# 2021-05-26
#
# # Julia Key Agreement in SageMath
#
# This worksheet accompanies the paper *Julia: Fast and Secure Key Agreement for IoT Devices*
# by <NAME> and <NAME>, 2021. SageMath 9.2 was used to create this worksheet. The latest version is hosted at [github.com/assaabloy-ppi/julia-in-sage/](https://github.com/assaabloy-ppi/julia-in-sage/).
#
# The example code uses the well-known Curve25519. Any other crypto group could have been chosen. This code should be seen as example code to illustrate the Julia Key Agreement (JKA) as it is described in the paper. Details, such as how to serialize a group element into bytes, is not of importance here. A full secure channel specification and implementation would need to consider many details that are out of scope here.
# # Init
#
# Run the code blocks below to initialize the field (field), the elliptic curve (ec), the generator (G), and some functions.
field = GF(2^255-19)
ec = EllipticCurve(field, [0,486662,0,1,0])
G = ec([9, 14781619447589544791020593568409986887264606134616475288964881837755586237401])
show(ec)
# Generates and returns a secret key.
def secret(): return 2^254 + 8*randint(1, 2^251-1)
# Returns a random byte array of length 32 (256 bits).
def rand():
array = bytearray(32)
for i in range(len(array)):
array[i] = randint(0, 255)
return array
# +
import hashlib
# Ordinary hash function, returns byte array
def hash1(a):
hasher = hashlib.sha256()
hasher.update(a)
return hasher.digest();
# Hashes and converts to 256-bit integer
def hash2(a):
digest = hash1(a)
return int.from_bytes(digest, byteorder='big', signed=False)
# Converts a string, Integer, or group element to a byte array.
def to_bytes(A):
if isinstance(A, Integer):
return str(A).encode("US-ASCII")
elif isinstance(A, str):
return A.encode("US-ASCII")
else:
return str(A[0]).encode("US-ASCII") + str(A[1]).encode("US_ASCII") + str(A[2]).encode("US_ASCII")
# -
# # Notes
#
# The following sections illustrate a number of key agreement protocols. Many details are not included, some of which are essential for a real implementation of a full secure channel handshake.
#
# Note, the following:
#
# * Two parties, P1 and P2, communicate over a realible, but insecure communication channel.
# * The attacker, Mallory has full power to modify and resend messages. She has access
# to all sessions ever executed and all sessions currently executing.
# * The communication can be assumed to be half-duplex. Only one of the parties send data
# at a time.
# * P1 initiates the communication and P2 responds to the initial message from P1.
# * The computation of one or multiple keys for the symmetric
# cryptography that follows in full secure channel protocol is not included.
# * To be concrete, we can assume that a shared symmetric key is computed
# from a hash of all session data and the result of one or multiple scalar multiplications
# that can be computed by both parties.
# * The goal of the handshake is to achieve one or more secrets that are shared between
# P1 and P2, but are not available to Mallory. For more information on the security properties
# of JKA, see the paper.
# # Key agreement with one scalar multiplication
#
# Each party computes only one scalar multiplication. A shared secret (D) is attained, but forward secrecy and compromised-key impersonation is not support.
# +
# ==== Static keys ====
# P1:
s1 = secret()
S1 = s1*G
# P2:
s2 = secret()
S2 = s2*G
# +
# ==== Pre-handshake ====
# P1:
c1 = rand()
# P2:
c2 = rand()
# +
# ==== Handshake ====
# P1:
# -> Send c1
# P2:
# <- Send c2
# P1:
D1 = s1*S2
# P2:
D2 = s2*S1
# -
D1 == D2
# # Key agreement with three scalar multiplications
#
# This key agreement is conceptually equivalent to protocol "XX" of the Noise Protocol Framework
# [noiseprotocol.org](https://noiseprotocol.org/).
# +
# ==== Static keys ====
# P1:
s1 = secret()
S1 = s1*G
# P2:
s2 = secret()
S2 = s2*G
# +
# ==== Pre-handshake ====
# P1:
e1 = secret()
E1 = e1*G
# P2:
e2 = secret()
E2 = e2*G
# +
# P1:
# -> Send E1
# P2:
D21 = E1*e2
D22 = E1*s2
# <- Send E2, S2
# P1:
D11 = e1*E2
D12 = e1*S2
D13 = s1*E2
# -> Send S1, app1
# P2:
D23 = S1*e2
# -
D11 == D21, D12 == D22, D13 == D23
# A common secret for encrypting application data can computed based on the session hash (hash of all data transferred) and the Dxx values. The application data is encrypted using all three scalar products (D11, D12, D13) while D11, D12 are used to encrypt S1 and S2.
# # Julia Key Agreement
#
# Between P1 and P2. The baseline version. t is computed jointly by P1 and P2.
# +
# ==== Static keys ====
# P1:
s1 = secret()
S1 = s1*G
# P2:
s2 = secret()
S2 = s2*G
# +
# ==== Pre-handshake ====
# P1:
e1 = secret()
E1 = e1*G
t1 = hash2(to_bytes("t1") + to_bytes(E1))
h1 = hash1(to_bytes("h1") + to_bytes(E1))
# P2:
e2 = secret()
E2 = e2*G
t2 = hash2(to_bytes("t2") + to_bytes(E2))
# +
# ==== Handshake ====
# P1:
# -> Send h1
# P2:
# <- Send E2
# P1:
t2 = hash2(to_bytes("t1") + to_bytes(E2))
t = t1+t2
D1 = (t*s1+e1)*(t*S2+E2)
# -> Send E1, app1
# P2:
# Verify h1 = hash1(to_bytes("h1") + to_bytes(E1))
t = t1+t2
D2 = (t*S1+E1)*(t*s2+e2)
# -
D1 == D2
# # Julia Key Agreement, one scalar multiplication
#
# t1 and t2 are computed independently by the parties. P1 can pre-compute t1, P2 can pre-compute t2.
# +
# ==== Pre-handshake ====
# P1:
e1 = secret()
E1 = e1*G
t1 = hash2(to_bytes("t1") + to_bytes(E1))
h1 = hash1(to_bytes("h1") + to_bytes(E1))
# P2:
e2 = secret()
E2 = e2*G
t2 = hash2(to_bytes("t2") + to_bytes(E2))
# +
# ==== Handshake ====
# P1:
# -> Send h1
# P2:
# <- Send E2
# P1:
t2 = hash2(to_bytes("t1") + to_bytes(E2))
D1 = (t2*s1+e1)*(t1*S2+E2)
# -> Send E1, app1
# P2:
# Verify h1 = hash(to_bytes("h1") + to_bytes(E1))
t = t1+t2
D2 = (t2*S1+E1)*(t1*s2+e2)
# -
D1 == D2
| julia.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Question1. & Solution.
#
# * Suppose ABCD is __not a frequent itemset__, while ABC and ACD are __frequent itemsets__. Which of the following is definitely true?
# * BC is a frequent itemset.
# * __definitely true__
# * ABCE is not a frequent itemset.
# * non-informable
# * ABCD is in the negative border.
# * Which means that ABC, ACD, BCD, ABD are all frequent itemsets => non-informable
# * ABD is a frequent itemset.
# * non-informable
# ## Question 2.
# * Suppose we are __representing sets__ by strings and __indexing the strings according to both the symbol and its position within the prefix__. We want to find strings within Jaccard distance at most 0.2 (i.e., similarity at least 0.8), and we are given __a probe string of length 24__. Into how many buckets must we look?
#
# ## Solution.
# * we must look through floor( J * L + 1) = floor( 0.2 * 24 + 1) = floor( 4.8 + 1 ) = 5
#
# ##Question 3.
# * In the following question we consider an example of __the implementation of the PCY algorithm__. All numbers should be treated as decimal; e.g., "one million" is 1,000,000, NOT $2^{20}$ = 1,048,576. All integers (item counts and bucket counts) require __4 bytes(32 bits)__.
# * We have __one billion bytes of main memory available for the first pass__. There are 100,000,000 items, and also 100,000,000 baskets, each of which contains exactly 10 items. Say that PCY is effective __if the average count of a bucket is at most half the support value__. For the given data, what is the smallest support value for which PCY will be effective?
# * 6
# * 60
# * 600
# * 6000
# ## Solution.
# * I believe the order of magnitude will be 10 or 100, so I choose 600
# ## Question 4.
# * Suppose we want to represent the multiplication of two 10-by-10 matrices as a "problem" in the sense used for our discussion of __the theory of MapReduce algorithms__. How many pairs are in the input-output mapping?
#
# ## Solution.
# * In a naive manner, a reducer get a triple ($A_i$, $B_j$, $C_{i,j}$). So in this case, the total number of pairs are 10 * 10 = 100 pairs
# ##Question 5.
# * The "all-triples" problem is described by __n inputs, (n choose 3) outputs__, and an input-output mapping where each output is connected to __a different set of three inputs__.
# * Suppose q is the __reducer size__. Which of the following functions of n and q approximates, to within a constant factor, __the lowest possible replication rate__ for a mapping schema that solves this problem?
# ##Question 6.
# * Suppose we are running the __DGIM algorithm__ (approximate counting of 1's in a window. At time t, the list of bucket sizes being maintained is 8,4,4,2,1,1. At times t+1, t+2, and t+3, 1's arrive on the input. Assuming no buckets are deleted because they fall outside the window, what are the numbers of buckets after each of the times t+1, t+2, and t+3?
#
# ## Solution.
# * T + 1: 1, 2, 2, 4, 4, 8 => 6 BUCKETS
# * T + 2: 1, 1, 2, 2, 4, 4, 8 => 7 BUCKETS
# * T + 3: 1, 1, 1, 2, 2, 4, 4, 8 => 1, 2, 4, 8, 8 => 5 BUCKETS
#
# ## Question 7.
# * Apply the HITS algorithm to a network with four pages (nodes) A, B, C, and D, arranged in a chain:
# A-->B-->C-->D
#
# * Compute the hubbiness and authority of each of these pages (scale doesn't matter, because you only have to identify pages with the same hubbiness or the same authority). Which of the following is FALSE.
#
# ## Solution.
# ##Question 8.
# * Let G be the complete graph on five nodes (i.e., there is an edge in G between every pair of distinct nodes). What is the sum of the squares of the elements of the Laplacian matrix for G?
# ##Question 9.
# * Note: This problem is similar to one on the Basic Final, but involves a combiner.
# * Consider the following MapReduce algorithm. The input is a collection of positive integers. Given integer X, the Map function produces a tuple with key Y and value X for each prime divisor Y of X. For example, if X = 20, there are two key-value pairs: (2,20) and (5,20). The Reduce function, given a key K and list L of values, produces a tuple with key K and value sum(L) i.e., the sum of the values in the list.
#
# * Suppose we process the input 9, 15, 16, 23, 25, 27, 28, 56, using a Combiner. There are 4 Map tasks and 1 Reduce task. The first Map task processes the first two inputs, the second the next two, and so on. How many input tuples does the Reduce task receive?
# ## Question 10
# * Consider an AdWords scenario with 4 advertisers competing for the same query Q, all with the same budget of 100 dollar and the same clickthrough rate. The table below shows the bid and the dollars spent by each advertiser until this point. Suppose we use Generalized BALANCE, and show one ad for each query. Which advertiser do we pick the next time query Q comes up?
#
# <pre>
#
# Advertiser Bid Spend
# A $1 $20
# B $2 $40
# C $3 $60
# D $4 $80
# </pre>
# ##Question 11.
# * Suppose we wish to estimate the rating of movie M by user U using item-Item Collaborative Filtering, but there are no movies really similar to movie M. The average of all ratings is 3.5, user U's average rating is 3.1, and movie M's average rating is 4.3. What is our best guess for the rating of movie M by user U using a global baseline estimate?
#
# ##Question 12
# * The table below shows data from ten people showing whether they like four different ice cream flavors.
# 
# * Fit a decision tree that predicts whether somebody would like Peanut ice cream based on whether she liked the other three flavors. Use Information gain as the measure to make the splits. What is the order of splits?
#
# ##Question 14
# * A is a users times movie-ratings matrix like the one seen in class. Each column in A represents a movie, and there are 5 movies in total. Recall that a Singular Value Decomposition of a matrix is a multiplication of three matrices: U, Σ and V. is the following is such a decomposition for matrix A:
#
# 
#
# * If we get three new users with the following rating vectors: User 1: [5,0,0,0,0] User 2: [0,5,0,0,0] User 3: [0,0,0,0,4] If for advertising purposes we want to cluster these three customers into two clusters using the movie concepts as features. How would you cluster them? (use cosine distance).
# ##Question 15
# * The soft margin SVM optimization problem is:
#
# * If for some i we have ξ=0, this indicates that the point xi is (check the true option):
# * A support vector √
# * Exactly in the decision boundary
# * Incorrectly classified
# * Correctly classified
| final/.ipynb_checkpoints/Final-advance-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:playground]
# language: python
# name: conda-env-playground-py
# ---
# %matplotlib inline
# +
import numpy as np
def _idcg(l):
return sum((1.0 / np.log(i + 2) for i in range(l)))
_idcgs = [_idcg(i) for i in range(101)]
def ndcg(gt, rec):
dcg = 0.0
for i, r in enumerate(rec):
if r in gt:
dcg += 1.0 / np.log(i + 2)
return dcg / _idcgs[len(gt)]
# +
import enum
class QuestionType(enum.Enum):
ALL = enum.auto()
SONG_TAG = enum.auto()
SONG_TITLE = enum.auto()
TAG_TITLE = enum.auto()
SONG_ONLY = enum.auto()
TAG_ONLY = enum.auto()
TITLE_ONLY = enum.auto()
NOTHING = enum.auto()
QUESTION_TYPE_MAP = {
# (songs, tags, title): question_type
(True, True, True): QuestionType.ALL,
(True, True, False): QuestionType.SONG_TAG,
(True, False, True): QuestionType.SONG_TITLE,
(False, True, True): QuestionType.TAG_TITLE,
(True, False, False): QuestionType.SONG_ONLY,
(False, True, False): QuestionType.TAG_ONLY,
(False, False, True): QuestionType.TITLE_ONLY,
(False, False, False): QuestionType.NOTHING,
}
def get_question_type(question):
songs = question['songs']
tags = question['tags']
title = question['plylst_title']
has_songs = len(songs) > 0
has_tags = len(tags) > 0
has_title = title != ""
return QUESTION_TYPE_MAP[has_songs, has_tags, has_title]
# +
import matplotlib.pyplot as plt
import numpy as np
from collections import defaultdict
from statistics import mean
def print_score(music_ndcgs, tag_ndcgs, scores):
music_ndcg = mean(music_ndcgs)
tag_ndcg = mean(tag_ndcgs)
score = mean(scores)
print(f"Music nDCG: {music_ndcg:.6}")
print(f"Tag nDCG: {tag_ndcg:.6}")
print(f"Score: {score:.6}")
def evaluate(results, questions, answers):
if len(results) < len(answers):
print("[Warning] 제출한 정답이 부족합니다.")
questions_dict = {p['id']: p for p in questions}
answers_dict = {p['id']: p for p in answers}
total_music_ndcgs = list()
total_tag_ndcgs = list()
total_scores = list()
case_music_ndcgs = defaultdict(list)
case_tag_ndcgs = defaultdict(list)
case_scores = defaultdict(list)
for p in results:
pid = p['id']
songs = p['songs']
tags = p['tags']
if pid not in questions_dict:
raise Exception(f"questions에 없습니다: {pid}")
if pid not in answers_dict:
raise Exception(f"answers 없습니다: {pid}")
question = questions_dict[pid]
answer = answers_dict[pid]
question_type = get_question_type(question)
# Validate playlist
if len(songs) != 100:
raise Exception(f"추천 곡 결과의 개수가 맞지 않습니다: {pid}")
if len(tags) != 10:
raise Exception(f"추천 태그 결과의 개수가 맞지 않습니다: {pid}")
if len(set(songs)) != 100:
raise Exception(f"한 플레이리스트에 중복된 곡 추천은 허용되지 않습니다: {pid}")
if len(set(tags)) != 10:
raise Exception(f"한 플레이리스트에 중복된 태그 추천은 허용되지 않습니다: {pid}")
cur_music_ndcg = ndcg(answer['songs'], songs)
cur_tag_ndcg = ndcg(answer['tags'], tags)
cur_score = cur_music_ndcg * 0.85 + cur_tag_ndcg * 0.15
# Update total score
total_music_ndcgs.append(cur_music_ndcg)
total_tag_ndcgs.append(cur_tag_ndcg)
total_scores.append(cur_score)
# Update case score
case_music_ndcgs[question_type].append(cur_music_ndcg)
case_tag_ndcgs[question_type].append(cur_tag_ndcg)
case_scores[question_type].append(cur_score)
return (
total_music_ndcgs, total_tag_ndcgs, total_scores,
case_music_ndcgs, case_tag_ndcgs, case_scores,
)
def print_scores(
total_music_ndcgs, total_tag_ndcgs, total_scores,
case_music_ndcgs, case_tag_ndcgs, case_scores,
):
print("=== Total score ===")
print_score(total_music_ndcgs, total_tag_ndcgs, total_scores)
for question_type in QuestionType:
if question_type not in case_music_ndcgs:
continue
print(f"=== {question_type.name} score ===")
print_score(case_music_ndcgs[question_type], case_tag_ndcgs[question_type], case_scores[question_type])
def create_histogram(music_ndcgs, tag_ndcgs, scores, ax):
bins=np.linspace(0, 1, 10)
# ax.hist([music_ndcgs, tag_ndcgs, scores], bins, label=["music_ndcgs", "tag_ndcgs", "score"])
ax.hist([music_ndcgs, tag_ndcgs], bins, alpha=1, label=["music_ndcgs", "tag_ndcgs"])
ax.hist(scores, bins, alpha=0.33, label="score")
# ax.set_xlim(0, 1)
# ax.set_ylim(0, 400)
ax.legend(loc='upper right')
# -
from utils import read_json
questions = read_json('./arena_data/questions/val.json')
answers = read_json('./arena_data/answers/val.json')
results = read_json('./arena_data/results/results.json')
(total_music_ndcgs, total_tag_ndcgs, total_scores,
case_music_ndcgs, case_tag_ndcgs, case_scores) = evaluate(results, questions, answers)
print_scores(
total_music_ndcgs, total_tag_ndcgs, total_scores,
case_music_ndcgs, case_tag_ndcgs, case_scores,
)
ax = plt.subplot()
plt.title("Total")
create_histogram(total_music_ndcgs, total_tag_ndcgs, total_scores, ax)
plt.show()
# +
plt.figure(figsize=(20, 10))
i = 1
ax = None
for question_type in QuestionType:
if question_type not in case_music_ndcgs:
continue
ax = plt.subplot(2, 4, i, sharex=ax, sharey=ax)
ax.set_title(question_type.name)
create_histogram(case_music_ndcgs[question_type], case_tag_ndcgs[question_type], case_scores[question_type], ax)
i += 1
plt.show()
ax = None
# -
| graph/detailed_evaluate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Giới thiệu phân phối SinhArcsinh
# BS <NAME>
#
# Phân phối SinhArcsinh được giới thiệu bởi <NAME> và <NAME> vào năm 2009 như một dạng tổng quát hoá của phân phối normal. Trong khi phân phối normal là đối xứng, có hai đuôi từ nhỏ đến vừa và có thể được định nghĩa bằng chỉ hai tham số ($\mu$-`loc` và $\sigma$-`scale`), phân phối SinhArcsinh cần thêm hai tham số để kiểm soát độ lệch ($\nu$-`skewness`) và trọng lượng của đuôi ($\tau$-`tailweight`).
#
# Các bạn có thể nhìn thấy ảnh hưởng của các tham số `nu` ($\nu$) và `tau`($\tau$) ở các hình sau:
# 
# 
# 
#
# `tensorflow` đã chuyển nó thành một dạng transformation để sử dụng được với tất cả các phân phối khác. Gần đây, `numpyro` đã cho phép sử dụng các phân phối có trong `tensorflow` để suy luận Bayes.
#
# Hôm nay tôi sẽ dùng phân phối này để suy luận tham số của một biến X. Biến X có phân phối lệch (T) và có một đuôi rất nhỏ.
import numpy as np
import pandas as pd
import seaborn as sns
import arviz as az
import numpyro
import numpyro.distributions as dist
import numpyro.contrib.tfp.distributions as tfd
from numpyro.infer import MCMC, NUTS, Predictive
from jax import random
import jax.numpy as jnp
import matplotlib.pyplot as plt
az.style.use('fivethirtyeight')
# ## Mô phỏng biến X
x = np.concatenate([
np.random.uniform(0,1, 100),
np.random.normal(1.1,0.1,50),
np.random.normal(0.9,0.1,500)
])
sns.kdeplot(x, bw_adjust=0.5)
plt.title('Phân phối của biến X');
# ## Suy luận
# Ta đặt prior cho 4 tham số, vì giá trị của mình khoảng nhỏ nên prior của `loc` cũng đặt gần giá trị 1.
def model(x=None):
loc = numpyro.sample('loc', dist.Normal(1,1))
scale = numpyro.sample('scale', dist.Exponential(1))
skewness = numpyro.sample('skewness', dist.Normal(0,1))
tailweight = numpyro.sample('tailweight', dist.Exponential(1))
numpyro.sample('x',
tfd.SinhArcsinh(loc, scale, skewness, tailweight,
distribution=tfd.Normal(0,1)),
obs=x
)
mcmc = MCMC(NUTS(model), 500,1000, 4,chain_method="sequential")
mcmc.run(random.PRNGKey(1), x=x)
mcmc.print_summary()
# ## Kiểm tra converge của các chuỗi MCMC.
az.plot_trace(mcmc);
# ## Kiểm tra fit của model
post = mcmc.get_samples()
x_seq = np.linspace(0,1.5,100)
plt.plot(x_seq,
np.exp(tfd.SinhArcsinh(
post['loc'].mean(),
post['scale'].mean(),
post['skewness'].mean(),
post['tailweight'].mean()
).log_prob(x_seq)))
plt.hist(x, density=True, bins=50, alpha=0.5);
# # Tổng kết:
# Phân phối SinhArcsinh là một phân phối rất flexible. R package `gamlss` đã ứng dụng nó làm phân phối chính, kết hợp thêm các kỹ thuật trong GAM để suy luận.
#
# Trong bối cảnh Generalized Linear Model, phân phối này là một phân phối hữu ích, bên cạnh Gamma Regression, Beta Regression. Để thành "SinhArcsinh regression", các bạn chỉ cần thêm linear model vào tham số `loc` như các GLM khác.
#
# Chúc các bạn thành công.
dist.transforms.AbsTransform
a = dist.TransformedDistribution(
tfd.SinhArcsinh(0,1,0,1),
dist.transforms.AbsTransform())
a.log_prob(1)
| notebooks/sinharchsinh.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 5. Single-layer Neural Network with Pattern images based on Lengyel-Epstein model
# - X : imges, Z = W * gradient(X) + b
# - optimizer : Adam
# ## 1) Import Packages
from PIL import Image
import numpy as np
import matplotlib.pyplot as plt
import random
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import OneHotEncoder
import math
import sklearn.metrics as metrics
# ## 2) Make Dataset
# +
# Make dataset(144)
x_orig = []
y_orig = np.zeros((1,48))
for i in range(1,145):
if i <= 48 :
folder = 0
elif i <=96 :
folder = 1
else:
folder = 2
img = Image.open('144/{0}/pattern_{1}.jpg'.format(folder,i))
data = np.array(img)
x_orig.append(data)
for i in range(1,3):
y_orig = np.append(y_orig, np.full((1, 48),i), axis = 1)
# +
# Make dataset(360)
x_orig = []
y_orig = np.zeros((1,120))
for i in range(1,361):
if i <= 120 :
folder = 0
elif i <=240 :
folder = 1
else:
folder = 2
img = Image.open('360/{0}/pattern_{1}.jpg'.format(folder,i))
data = np.array(img)
x_orig.append(data)
for i in range(1,3):
y_orig = np.append(y_orig, np.full((1, 120),i), axis = 1)
# +
# Make dataset(720)
x_orig = []
y_orig = np.zeros((1,240))
for i in range(1,721):
if i <= 240 :
folder = 0
elif i <=480 :
folder = 1
else:
folder = 2
img = Image.open('720/{0}/pattern_{1}.jpg'.format(folder,i))
data = np.array(img)
x_orig.append(data)
for i in range(1,3):
y_orig = np.append(y_orig, np.full((1, 240),i), axis = 1)
# -
x_orig = np.array(x_orig)
print(x_orig.shape)
print(y_orig.shape)
# +
# Random shuffle
s = np.arange(x_orig.shape[0])
np.random.shuffle(s)
x_shuffle = x_orig[s,:]
y_shuffle = y_orig[:,s]
print(x_shuffle.shape)
print(y_shuffle.shape)
# y_shuffle
# -
# Split train and test datasets
x_train_orig, x_test_orig, y_train_orig, y_test_orig = train_test_split(x_shuffle,y_shuffle.T,
test_size=0.3, shuffle=True, random_state=1004)
print(x_train_orig.shape)
print (y_train_orig.shape)
# +
# Flatten the training and test images
x_train_flatten = x_train_orig.reshape(x_train_orig.shape[0], -1).T
x_test_flatten = x_test_orig.reshape(x_test_orig.shape[0], -1).T
# Normalize image vectors
x_train = x_train_flatten/255.
x_test = x_test_flatten/255.
# Convert training and test labels to one hot matrices
enc = OneHotEncoder()
y1 = y_train_orig.reshape(-1,1)
enc.fit(y1)
y_train = enc.transform(y1).toarray()
y_train = y_train.T
y2 = y_test_orig.reshape(-1,1)
enc.fit(y2)
y_test = enc.transform(y2).toarray()
y_test = y_test.T
# Explore dataset
print ("number of training examples = " + str(x_train.shape[1]))
print ("number of test examples = " + str(x_test.shape[1]))
print ("x_train shape: " + str(x_train.shape))
print ("y_train shape: " + str(y_train.shape))
print ("x_test shape: " + str(x_test.shape))
print ("y_test shape: " + str(y_test.shape))
# -
# ## 3) Define required functions
def initialize_parameters(nx, ny):
"""
Argument:
nx -- size of the input layer (4096)
ny -- size of the output layer (3)
Returns:
W -- weight matrix of shape (ny, nx)
b -- bias vector of shape (ny, 1)
"""
np.random.seed(1)
W = np.random.randn(ny,nx)*0.01
b = np.zeros((ny,1))
assert(W.shape == (ny, nx))
assert(b.shape == (ny, 1))
return W, b
def softmax(Z):
# compute the softmax activation
S = np.exp(Z + np.max(Z)) / np.sum(np.exp(Z + np.max(Z)), axis = 0)
return S
def classlabel(Z):
# probabilities back into class labels
y_hat = Z.argmax(axis=0)
return y_hat
def gradient_vec(X):
g_X_r = np.gradient(X, axis = 1)
g_X_c = np.gradient(X, axis = 0)
g_X = g_X_r**2 + g_X_c**2
return g_X
# +
def propagate(W, b, X, Y):
m = X.shape[1]
# n = Y.shape[0]
# Forward Propagation
Z = np.dot(W, gradient_vec(X))+ b
A = softmax(Z) # compute activation
cost = (-1/m) * np.sum(Y * np.log(A)) # compute cost (Cross_entropy)
# Backward propagation
dW = (1/m) * (np.dot(gradient_vec(X),(A-Y).T)).T
db = (1/m) * (np.sum(A-Y))
# assert(dW.shape == W.shape)
# assert(db.dtype == float)
# cost = np.squeeze(cost)
# assert(cost.shape == (Y.shape[0],1))
grads = {"dW": dW,
"db": db}
return grads, cost
# -
# ## 4) Single-Gradient Layer Neural Network with Adam
def optimize(X, Y, num_iterations, learning_rate, t, beta1 = 0.9, beta2 = 0.999, epsilon = 1e-8, print_cost = False):
costs = []
W, b = initialize_parameters(4096,3)
v_dW = np.zeros((W.shape[0],W.shape[1]))
v_db = np.zeros((b.shape[0],b.shape[1]))
s_dW = np.zeros((W.shape[0],W.shape[1]))
s_db = np.zeros((b.shape[0],b.shape[1]))
for i in range(num_iterations):
grads, cost = propagate(W, b, X, Y)
dW = grads["dW"]
db = grads["db"]
# update parameters with adam
v_dW = beta1 * v_dW + (1-beta1) * dW
v_db = beta1 * v_db + (1-beta1) * db
# Compute bias-corrected first moment estimate
v_corrected_dW = v_dW / (1-beta1**t)
v_corrected_db = v_db / (1-beta1**t)
# Moving average of the squared gradients
s_dW = beta2 * s_dW + (1-beta2) * dW ** 2
s_db = beta2 * s_db + (1-beta2) * db ** 2
# Compute bias-corrected second raw moment estimate.
s_corrected_dW = s_dW / (1-beta2**t)
s_corrected_db = s_db / (1-beta2**t)
# Update parameters
W = W - (learning_rate) * ((v_corrected_dW) / (s_corrected_dW **(1/2) + epsilon))
b = b - (learning_rate) * ((v_corrected_db) / (s_corrected_db ** (1/2) + epsilon))
# Record the costs for plotting
if i % 200 == 0:
costs.append(cost)
# Print the cost every 100 training iterations
if print_cost and i % 200 == 0:
print ("Cost after iteration %i: %f" %(i, cost))
# plot the cost
plt.plot(costs)
plt.ylabel('cost')
plt.xlabel('iterations (per 200)')
plt.title("Learning rate =" + str(learning_rate))
plt.show()
# Lets save the trainded parameters in a variable
params = {"W": W,
"b": b}
grads = {"dW": dW,
"db": db}
return params, grads, costs
# +
params, grads, costs = optimize(x_train, y_train, num_iterations=2000, learning_rate=0.9,
t=2, beta1 = 0.9, beta2 = 0.999, epsilon = 1e-8, print_cost = True)
print ("W = " + str(params["W"]))
print ("b = " + str(params["b"]))
print ("dw = " + str(grads["dW"]))
print ("db = " + str(grads["db"]))
# -
# ## 5) Accuracy Analysis
# +
def predict(W, b, X) :
'''
Predict the label(0,1,2) using argmax
Arguments:
X : data of size (num_px * num_px, number of examples)
Returns:
y_prediction : predictions (0/1/2) for the examples(my_image)
'''
m = X.shape[1]
# Compute "A" predicting the probabilities
Z = np.dot(W, gradient_vec(X))+ b
A = softmax(Z)
# Convert probabilities A to actual predictions
y_prediction = A.argmax(axis=0)
# assert(y_prediction.shape == (1, m))
return y_prediction
# -
# Predict test/train set
W1 = params['W']
b1 = params['b']
y_prediction_train = predict(W1, b1, x_train)
y_prediction_test = predict(W1, b1, x_test)
print(y_prediction_train)
print(y_prediction_test)
# Print train/test Errors
print("train accuracy : ", metrics.accuracy_score(y_prediction_train, y_train_orig))
print("test accuracy : ", metrics.accuracy_score(y_prediction_test, y_test_orig))
| Classification_of_Pattern_Images/5_1gradient_layer_NN_with_Adam.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
x = np.arange(0, 10, 1)
y = np.sin(x)
z = np.cos(x)
plt.plot(x, y)
plt.plot(x, z)
# +
plt.plot(x, y, marker="o", color="r", linestyle="--", label="sin curve")
plt.plot(x, z, marker="x", color="k", linestyle="--", label="cos curve")
plt.xlabel("X axis");
plt.ylabel("Y axis")
plt.legend()
plt.xlim(0,9)
plt.ylim(-1.25, 1.25)
plt.title("Sine and Cosine curves")
plt.grid()
# -
plt.scatter(x, y, label="sine")
plt.scatter(x, z, label="cosine")
plt.legend()
# ### Matplotlib functions may also be accessed through a lower level interface: axes
#
# The "plt" interface does not allow us as much control over the plot elements. For greater control in more complicated plots, we need the axes interface
# +
fig = plt.figure(figsize=(190/25.4, 100/25.4) )
ax = plt.axes()
line1, = ax.plot(x, y, marker="o", color="r", linestyle="--", label="sin curve")
line2, = ax.plot(x, z, marker="x", color="k", linestyle="--", label="cos curve")
#ax.plot(x, np.exp(x), marker="+", color="b", linestyle="--", label="exp curve")
axr = plt.twinx()
line3, = axr.plot(x, np.exp(x), marker="v", color="b", linestyle="--", label="exp curve")
ax.set_xlabel("X axis");
ax.set_ylabel("Sine and Cosine curves")
ax.set_xlim(0,9)
ax.set_ylim(-1.25, 1.25)
ax.set_xticks(np.arange(10))
ax.set_xticks(np.arange(0.5, 9, 1), minor=True)
ax.set_yticks([-0.25, 0, 0.25], minor=True)
axr.set_ylabel("Exponential curve (Log scale)", color="b")
axr.set_yscale("log")
ax.set_title("Matplotlib plot functions using axes")
ax.grid()
ax.grid(which="minor", linestyle="--", linewidth=0.25)
ax.legend([line1, line2, line3], ["sin curve", "cos curve", "exp curve"], loc="lower right")
#plt.legend()
plt.savefig("./sin_cos_exp.jpg", dpi=150, bbox_inches="tight")
# -
# An exponential function on a log scale is linear, note the right hand side Y axis having a log scale!
# alternatively, use get_legend_handles_labels()
handles, labels = ax.get_legend_handles_labels()
labels
np.std(y)
# ### Errorbars
yerr_for_the_redLine = np.array([np.arange(0, 1, 0.1), np.arange(1, 2, 0.1)])
# +
fig = plt.figure(figsize=(190/25.4, 100/25.4) )
ax = plt.axes()
#line1, = ax.plot(x, y, color="r", linestyle="--", label="sin curve")
line1 = ax.errorbar(x, y, yerr=yerr_for_the_redLine, marker="o", color="r", linestyle="--", label="sin curve", capsize=5)
line2 = ax.errorbar(x, z, yerr=np.std(z), marker="x", color="k", linestyle="--", label="cos curve", capsize=5)
#ax.plot(x, np.exp(x), marker="+", color="b", linestyle="--", label="exp curve")
axr = plt.twinx()
line3, = axr.plot(x, np.exp(x), marker="+", color="b", linestyle="--", label="exp curve")
ax.set_xlabel("X axis");
ax.set_ylabel("Sine and Cosine curves")
ax.set_xlim(0,9)
ax.set_ylim(-2, 2)
ax.set_xticks(np.arange(0.5, 9), minor=True)
axr.set_ylabel("Exponential curve (Log scale)", color="b")
axr.set_yscale("log")
ax.set_title("Matplotlib plot functions using axes")
ax.grid()
ax.grid(which="minor", linestyle="--", linewidth=0.25)
ax.legend([line1, line2, line3], ["sin curve", "cos curve", "exp curve"])
# -
fig = plt.figure(figsize=(190/25.4, 100/25.4) )
ax = plt.axes()
ax.fill_between(x, y-np.std(y), y+np.std(y), color="0.75")
ax.plot(x, y, color="k")
ax.grid()
ax.set_xlim(0,9)
ax.set_ylim(-2, 2)
# ## Bar plots
plt.hist(np.random.randn(1000)*100, bins=100);
plt.bar(np.arange(1,11), np.random.randn(10)*5+20 )
# ## Box plots
x = np.array([np.random.randn(1000), np.random.randn(1000)*1.5+1]).T
x.shape
x = np.array([np.random.randn(1000), np.random.randn(1000)*1.5+1]).T
plt.boxplot(x);
# The box encloses the Q1 (1st quantile, or the 25th percentile value) to Q3 (3rd quantile, or the 75th percentile value) range.
np.quantile(x, 0.25), np.quantile(x, 0.75)
# The whiskers are drawn at Q1 - 1.5\*IQR and Q3 + 1.5\*IQR, where IQR is the interquantile range. The circles are the outlier values beyond the whiskers. You can customize all these elements of the plot.
Q1 = np.quantile(x, 0.25)
Q3 = np.quantile(x, 0.75)
IQR = Q3 - Q1
print(Q1 - 1.5*IQR, Q3 + 1.5*IQR)
# ## Representing 2 dimensional data using pcolormesh, contour, contourf
x
x[1] - x[0]
# +
x = np.linspace(-2*np.pi, 2*np.pi, 100)
xc = x[:-1] + np.diff(x)[0] * 0.5 # center points of X axis
y = np.linspace(-2*np.pi, 2*np.pi, 100)
yc = y[:-1] + np.diff(y)[0] * 0.5 # center points of Y axis
XX, YY = np.meshgrid(xc,yc)
# -
x, xc
XX
Z = np.sin(XX) + np.sin(YY)
Z.shape
# + jupyter={"outputs_hidden": true}
# plt.pcolormesh?
# -
PC = plt.pcolormesh(x, y, Z, cmap="RdBu_r")
plt.colorbar(PC)
# + jupyter={"outputs_hidden": true}
# plt.contour?
# -
CR = plt.contour(xc, yc, Z, colors="k")
plt.clabel(CR)
#plt.colorbar(CR)
CF = plt.contourf(xc, yc, Z)
plt.colorbar(CF)
plt.style.available[:5]
with plt.style.context("bmh"):
plt.plot(x, np.sin(x))
# Try changing the style context and have a look at available styles at the matplotlib gallery: https://matplotlib.org/stable/gallery/style_sheets/style_sheets_reference.html
# ## Exercise for Matplotlib
#
# 1. Pick two 1D functions of your choice y = f(x), z = g(x)
# 1. Plot the functions using the plt.plot command, remember to label the functions.
# 1. Label the X and Y axes of the plot.
# 1. Set an appropriate title for your plot.
# 1. Set the X and Y axis limits appropriately.
# 1. Add errorbars to the plot corresponding to $\pm 1\sigma$, where $\sigma$ is the standard deviation of the function.
# 1. Set appropriate X and Y ticks, including minor ticks.
# 1. Call the legend() function to plot the legend.
# +
fig = plt.figure(figsize = (190*0.5/25.4, 230*0.25/25.4))
ax = plt.subplot()
x = np.arange(0, 1, 0.1)
y = 2 * x
z = x**2
y_std = np.std(y)
z_std = np.std(z)
ax.plot(x, y, label="linear func.", color="b")
ax.plot(x, z, label = "exp. func.", color="k")
ax.errorbar(x, y, yerr=y_std, color="b")
ax.errorbar(x, z, yerr=z_std, color="k")
plt.title("Title")
plt.xlim(0, 0.6)
#ax.set_xticks(np.arange(0.05, 0.6, 0.1), minor=True)
ax.set_xticks([])
plt.xlabel("Time (s)")
plt.ylabel("f(x), g(x)")
plt.legend()
plt.grid()
# -
plt.plot(x, z)
# <br></br>
# ### Example using GridSpec for plot layouts
# +
x = np.arange(0,10,0.1)
temp = 2 * x + np.random.randn(x.shape[0])*5
productivity = 0.5 * temp
salinity = np.random.randn(x.shape[0]) * 2.0 + 33
# -
import matplotlib.gridspec as gridspec
# + jupyter={"outputs_hidden": true}
# gridspec.GridSpec?
# +
fig = plt.figure(figsize=(190/25.4, 230/25.4))
gs = gridspec.GridSpec(2, 2, height_ratios=[1, 1], width_ratios=[1, 0.5])
ax1 = plt.subplot(gs[0,0])
ax1.plot(x, temp, label="Temp.")
ax1.set_title("Temperature")
ax1.set_ylabel("$^\circ$C")
ax2 = plt.subplot(gs[:,1])
ax2.plot(x, productivity, label = "productivity")
ax2.set_title("Productivity")
ax3 = plt.subplot(gs[1, 0])
ax3.plot(x, salinity, label="salinity")
ax3.set_title("Salinity")
ax3.set_ylabel("PSU")
# +
# ?
# -
gs = gridspec.GridSpec
| Matplotlib_intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="zHHjQL_MhT_s"
# 
# + [markdown] id="VVL1jdg3hXfb"
# [](https://colab.research.google.com/github/JohnSnowLabs/spark-nlp-workshop/blob/master/tutorials/Certification_Trainings/Healthcare/10.1.Clinical_Relation_Extraction_BodyParts_Models.ipynb)
# + [markdown] id="PSk7NM6_hXOs"
# # 10.1 Clinical Relation Extraction BodyPart Models
#
# (requires Spark NLP 2.7.1 and Spark NLP Healthcare 2.7.2 and above)
# + id="6Yr81i2yc4td"
import os
jsl_secret = os.getenv('SECRET')
import sparknlp
sparknlp_version = sparknlp.version()
import sparknlp_jsl
jsl_version = sparknlp_jsl.version()
print (jsl_secret)
# + colab={"base_uri": "https://localhost:8080/"} id="v2QJSWNVFDrH" outputId="bbb81391-e246-4f32-b14e-22577f762a17"
import json
import os
from pyspark.ml import Pipeline
from pyspark.sql import SparkSession
from sparknlp.annotator import *
from sparknlp_jsl.annotator import *
from sparknlp.base import *
import sparknlp_jsl
import sparknlp
params = {"spark.driver.memory":"16G",
"spark.kryoserializer.buffer.max":"2000M",
"spark.driver.maxResultSize":"2000M"}
spark = sparknlp_jsl.start(jsl_secret, params=params)
print (sparknlp.version())
print (sparknlp_jsl.version())
# + colab={"base_uri": "https://localhost:8080/", "height": 208} id="NDLcNaDyhN6J" outputId="49bdc79f-07ff-4946-c91b-a9bcf462460a"
spark
# + [markdown] id="52YRfABhhN6K"
# ## 1. Prediction Pipeline for Clinical Binary Relation Models
# + [markdown] id="JOob76OLZNHt"
# Basic Pipeline without Re Models. Run it once and we can add custom Re models to the same pipeline
# + colab={"base_uri": "https://localhost:8080/"} id="mnyPwdRIhN6M" outputId="b0211c7f-c602-4389-db41-8029a4feaa43"
documenter = DocumentAssembler()\
.setInputCol("text")\
.setOutputCol("document")
sentencer = SentenceDetector()\
.setInputCols(["document"])\
.setOutputCol("sentences")
tokenizer = Tokenizer()\
.setInputCols(["sentences"])\
.setOutputCol("tokens")\
words_embedder = WordEmbeddingsModel()\
.pretrained("embeddings_clinical", "en", "clinical/models")\
.setInputCols(["sentences", "tokens"])\
.setOutputCol("embeddings")
pos_tagger = PerceptronModel()\
.pretrained("pos_clinical", "en", "clinical/models") \
.setInputCols(["sentences", "tokens"])\
.setOutputCol("pos_tags")
dependency_parser = DependencyParserModel()\
.pretrained("dependency_conllu", "en")\
.setInputCols(["sentences", "pos_tags", "tokens"])\
.setOutputCol("dependencies")
# get pretrained ner model
clinical_ner_tagger = MedicalNerModel()\
.pretrained('jsl_ner_wip_greedy_clinical','en','clinical/models')\
.setInputCols("sentences", "tokens", "embeddings")\
.setOutputCol("ner_tags")
ner_chunker = NerConverter()\
.setInputCols(["sentences", "tokens", "ner_tags"])\
.setOutputCol("ner_chunks")
# + id="pAmatZ3dAF4n"
import pandas as pd
# This function will be utilized to show prediction results in a dataframe
def get_relations_df (results, col='relations'):
rel_pairs=[]
for rel in results[0][col]:
rel_pairs.append((
rel.result,
rel.metadata['entity1'],
rel.metadata['entity1_begin'],
rel.metadata['entity1_end'],
rel.metadata['chunk1'],
rel.metadata['entity2'],
rel.metadata['entity2_begin'],
rel.metadata['entity2_end'],
rel.metadata['chunk2'],
rel.metadata['confidence']
))
rel_df = pd.DataFrame(rel_pairs, columns=['relations',
'entity1','entity1_begin','entity1_end','chunk1',
'entity2','entity2_end','entity2_end','chunk2',
'confidence'])
# limit df columns to get entity and chunks with results only
rel_df = rel_df.iloc[:,[0,1,4,5,8,9]]
return rel_df
# + [markdown] id="Wa00p42s-1vj"
# ### Example pairs for relation entities
# + id="SSkP-5aKhN6N"
# bodypart entities >> ['external_body_part_or_region', 'internal_organ_or_component']
# 1. bodypart vs problem
pair1 = ['symptom-external_body_part_or_region', 'external_body_part_or_region-symptom']
# 2. bodypart vs procedure and test
pair2 = ['internal_organ_or_component-imagingtest',
'imagingtest-internal_organ_or_component',
'internal_organ_or_component-procedure',
'procedure-internal_organ_or_component',
'internal_organ_or_component-test',
'test-internal_organ_or_component',
'external_body_part_or_region-imagingtest',
'imagingtest-external_body_part_or_region',
'external_body_part_or_region-procedure',
'procedure-external_body_part_or_region',
'external_body_part_or_region-test',
'test-external_body_part_or_region']
# 3. bodypart vs direction
pair3 = ['direction-external_body_part_or_region', 'external_body_part_or_region-direction',
'internal_organ_or_component-direction','direction-internal_organ_or_component']
# 4. date vs other clinical entities
# date entities >> ['Date', 'RelativeDate', 'Duration', 'RelativeTime', 'Time']
pair4 = ['symptom-date', 'date-procedure', 'relativedate-test', 'test-date']
# + [markdown] id="6g_Rw2_0hN6N"
# **Pretrained relation model names**; use this names in `RelationExtractionModel()` ;
#
# + `re_bodypart_problem`
#
# + `re_bodypart_directions`
#
# + `re_bodypart_proceduretest`
#
# + `re_date_clinical`
# + [markdown] id="szA2kg2B-9TG"
# ## 2. Example of how custom RE models can be added to the same pipeline
# + [markdown] id="FyAR5U18ZNHz"
# ### 2.1 Relation Extraction Model
# + colab={"base_uri": "https://localhost:8080/"} id="O-0YDDdwhN6O" outputId="4ba29339-76ba-469a-b61f-ea77cfe1ba41"
re_model = RelationExtractionModel()\
.pretrained("re_bodypart_directions", "en", 'clinical/models')\
.setInputCols(["embeddings", "pos_tags", "ner_chunks", "dependencies"])\
.setOutputCol("relations")\
.setRelationPairs(['direction-external_body_part_or_region',
'external_body_part_or_region-direction',
'direction-internal_organ_or_component',
'internal_organ_or_component-direction'
])\
.setMaxSyntacticDistance(4)\
.setPredictionThreshold(0.9)
trained_pipeline = Pipeline(stages=[
documenter,
sentencer,
tokenizer,
words_embedder,
pos_tagger,
clinical_ner_tagger,
ner_chunker,
dependency_parser,
re_model
])
empty_data = spark.createDataFrame([[""]]).toDF("text")
loaded_re_model = trained_pipeline.fit(empty_data)
# + [markdown] id="NDLzlhI9ZNH1"
# ### 2.2 ReDL Model - based on end-to-end trained Bert Model
# + colab={"base_uri": "https://localhost:8080/"} id="28ICav0cZNH2" outputId="086768a8-5176-41ca-bde7-a19386c7c0ee"
re_ner_chunk_filter = RENerChunksFilter() \
.setInputCols(["ner_chunks", "dependencies"])\
.setOutputCol("re_ner_chunks")\
.setMaxSyntacticDistance(4)\
.setRelationPairs(['direction-external_body_part_or_region',
'external_body_part_or_region-direction',
'direction-internal_organ_or_component',
'internal_organ_or_component-direction'
])
re_model = RelationExtractionDLModel() \
.pretrained('redl_bodypart_direction_biobert', "en", "clinical/models")\
.setPredictionThreshold(0.5)\
.setInputCols(["re_ner_chunks", "sentences"]) \
.setOutputCol("relations")
trained_pipeline = Pipeline(stages=[
documenter,
sentencer,
tokenizer,
words_embedder,
pos_tagger,
clinical_ner_tagger,
ner_chunker,
dependency_parser,
re_ner_chunk_filter,
re_model
])
empty_data = spark.createDataFrame([[""]]).toDF("text")
loaded_redl_model = trained_pipeline.fit(empty_data)
# + [markdown] id="A0HO7NYlhN6O"
# ## 3. Sample clinical tetxs
# + id="dHAWmLXkhN6P"
# bodypart vs problem
text1 = '''No neurologic deficits other than some numbness in his left hand.'''
# bodypart vs procedure and test
#text2 = 'Common bile duct was noted to be 10 mm in size on that ultrasound.'
#text2 = 'Biopsies of the distal duodenum, gastric antrum, distalesophagus were taken and sent for pathological evaluation.'
text2 = 'TECHNIQUE IN DETAIL: After informed consent was obtained from the patient and his mother, the chest was scanned with portable ultrasound.'
# bodypart direction
text3 = '''MRI demonstrated infarction in the upper brain stem , left cerebellum and right basil ganglia'''
# date vs other clinical entities
text4 = ''' This 73 y/o patient had Brain CT on 1/12/95, with progressive memory and cognitive decline since 8/11/94.'''
# + [markdown] id="RmTkYtm9hN6P"
# **Get Single Prediction** with `LightPipeline()`
# + [markdown] id="e5P-pPuBZNH5"
# ### 3. 1 Using Relation Extraction Model
# + colab={"base_uri": "https://localhost:8080/", "height": 171} id="9tuTJp81hN6Q" outputId="54c3c17b-a0fd-4597-a8f3-483430b1fef5"
# choose one of the sample texts depending on the pretrained relation model you want to use
text = text3
loaded_re_model_light = LightPipeline(loaded_re_model)
annotations = loaded_re_model_light.fullAnnotate(text)
rel_df = get_relations_df(annotations) # << get_relations_df() is the function defined in the 3rd cell
print('\n',text)
rel_df[rel_df.relations!="0"]
#rel_df
# + [markdown] id="mkYhDW9lZNH7"
# ### 3.2 Using Relation Extraction DL Model
# + colab={"base_uri": "https://localhost:8080/", "height": 171} id="VUeWINUCZNH7" outputId="acde26fd-1d7f-41e7-d30a-a35b4d3fc5ea"
# choose one of the sample texts depending on the pretrained relation model you want to use
text = text3
loaded_re_model_light = LightPipeline(loaded_redl_model)
annotations = loaded_re_model_light.fullAnnotate(text)
rel_df = get_relations_df(annotations) # << get_relations_df() is the function defined in the 3rd cell
print('\n',text)
rel_df[rel_df.relations!="0"]
#rel_df
# + [markdown] id="jLUZZSIL_Kzt"
# ## Custom Function
# + id="bDVfQ7pmhN6R"
# Previous cell content is merged in this custom function to get quick predictions, for custom cases please check parameters in RelationExtractionModel()
def relation_exraction(model_name, pairs, text):
re_model = RelationExtractionModel()\
.pretrained(model_name, "en", 'clinical/models')\
.setInputCols(["embeddings", "pos_tags", "ner_chunks", "dependencies"])\
.setOutputCol("relations")\
.setRelationPairs(pairs)\
.setMaxSyntacticDistance(3)\
.setPredictionThreshold(0.9)
trained_pipeline = Pipeline(stages=[
documenter,
sentencer,
tokenizer,
words_embedder,
pos_tagger,
clinical_ner_tagger,
ner_chunker,
dependency_parser,
re_model
])
empty_data = spark.createDataFrame([[""]]).toDF("text")
loaded_re_model = trained_pipeline.fit(empty_data)
loaded_re_model_light = LightPipeline(loaded_re_model)
annotations = loaded_re_model_light.fullAnnotate(text)
rel_df = get_relations_df(annotations) # << get_relations_df() is the function defined in the 3rd cell
print('\n','Target Text : ',text, '\n')
#rel_df
return rel_df[rel_df.relations!="0"]
def relation_exraction_dl(model_name, pairs, text):
re_ner_chunk_filter = RENerChunksFilter() \
.setInputCols(["ner_chunks", "dependencies"])\
.setOutputCol("re_ner_chunks")\
.setRelationPairs(pairs)
re_model = RelationExtractionDLModel() \
.pretrained(model_name, "en", "clinical/models")\
.setPredictionThreshold(0.0)\
.setInputCols(["re_ner_chunks", "sentences"]) \
.setOutputCol("relations")
trained_pipeline = Pipeline(stages=[
documenter,
sentencer,
tokenizer,
words_embedder,
pos_tagger,
clinical_ner_tagger,
ner_chunker,
dependency_parser,
re_ner_chunk_filter,
re_model
])
empty_data = spark.createDataFrame([[""]]).toDF("text")
loaded_re_model = trained_pipeline.fit(empty_data)
loaded_re_model_light = LightPipeline(loaded_re_model)
annotations = loaded_re_model_light.fullAnnotate(text)
rel_df = get_relations_df(annotations) # << get_relations_df() is the function defined in the 3rd cell
print('\n','Target Text : ',text, '\n')
#rel_df
return rel_df[rel_df.relations!="0"]
# + [markdown] id="T6jxxTJS_Q6i"
# ## Predictions with Custom Function
# + [markdown] id="1iA1uHynZNH9"
# ### 4.1 Bodypart vs Problem - RelationExtractionModel
# + colab={"base_uri": "https://localhost:8080/", "height": 176} id="TBfM2LXVhN6R" outputId="05925250-507c-4014-8429-7971142b20d3"
# bodypart vs problem
model_name = 're_bodypart_problem'
pairs = ['symptom-external_body_part_or_region', 'external_body_part_or_region-symptom']
text = "Some numbness in his left hand noted, no other neurologic deficts."
relation_exraction(model_name, pairs, text)
# + [markdown] id="fOFg3sabZNH-"
# ### 4.2 Bodypart vs Problem - RelationExtractionDLModel
# + colab={"base_uri": "https://localhost:8080/", "height": 176} id="Aac4wUI1ZNH-" outputId="123a253e-4425-4b9f-d69e-0ddfe179845d"
# bodypart vs problem
model_name = 'redl_bodypart_problem_biobert'
pairs = ['symptom-external_body_part_or_region', 'external_body_part_or_region-symptom']
text = "Some numbness in his left hand noted, no other neurologic deficts."
relation_exraction_dl(model_name, pairs, text)
# + [markdown] id="QHeHj2xmZNH_"
# ### 5.1 Bodypart vs Procedure & Test - RelationExtractionModel
# + colab={"base_uri": "https://localhost:8080/", "height": 176} id="PqywwOKZhN6S" outputId="a59f96f6-0671-4083-a1f7-f8efeda5c264"
# bodypart vs procedure and test
model_name = 're_bodypart_proceduretest'
pairs = pair2
text = text2
relation_exraction(model_name, pairs, text)
# + [markdown] id="9j3lVFtAZNIA"
# ### 5.2 Bodypart vs Procedure & Test - RelationExtractionDLModel
# + colab={"base_uri": "https://localhost:8080/", "height": 176} id="nRa7_pYxZNIA" outputId="e0942b9d-3759-454e-8c48-163e5183e44b"
# bodypart vs procedure and test
model_name = 'redl_bodypart_procedure_test_biobert'
pairs = pair2
text = text2
relation_exraction_dl(model_name, pairs, text)
# + [markdown] id="RBLGvr1aZNIA"
# ### 6.1 Bodypart vs Directions - RelationExtractionModel
# + colab={"base_uri": "https://localhost:8080/", "height": 236} id="QSLzQFY2hN6T" outputId="07a9dfee-e2ce-4447-d85e-e7babed1b493"
# bodypart vs directions
model_name = 're_bodypart_directions'
pairs = pair3
text = text3
relation_exraction(model_name, pairs, text)
# + [markdown] id="7RRWfsVzZNIB"
# ### 6.2 Bodypart vs Directions - RelationExtractionDLModel
# + colab={"base_uri": "https://localhost:8080/", "height": 236} id="_p3dO7vRZNIC" outputId="a5a199dc-47fa-4776-b73c-3b9d4e8ebd13"
# bodypart vs directions
model_name = 'redl_bodypart_direction_biobert'
pairs = pair3
text = text3
relation_exraction_dl(model_name, pairs, text)
# + [markdown] id="Iv4OGxWZZNIC"
# ### 7.1 Date vs Clinical Entities - RelationExtractionModel
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="SBEyTj4MhN6T" outputId="d13178e5-e4f6-400e-cc45-96cf0f0a1944"
# date vs clinical date entities
model_name = 're_date_clinical'
pairs = pair4
text = text4
relation_exraction(model_name, pairs, text)
# + [markdown] id="7-m6k94_ZNID"
# ### 7.2 Date vs Clinical Entities - RelationExtractionDLModel
# + colab={"base_uri": "https://localhost:8080/", "height": 267} id="3nHeUDt1HFwh" outputId="6866d991-d2bf-4251-9c0c-aef3b8e9ac17"
# date vs clinical date entities
model_name = 'redl_date_clinical_biobert'
pairs = pair4
text = text4
relation_exraction_dl(model_name, pairs, text)
| jupyter/docker_enterprise/docker_notebooks/Spark_NLP/Healthcare/10.1.Clinical_Relation_Extraction_BodyParts_Models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.9 64-bit (''Python3.7'': conda)'
# name: python3
# ---
# # We plot some utility function
import numpy as np
import matplotlib.pyplot as plt
x = np.linspace(0.0, 1.0, 101)
f = lambda a : (1 - np.exp(-a*x))/a
plt.figure(figsize=(6,4))
plt.plot(x,f(1.0), label=r'$a=1$ (concave)')
plt.plot(x,f(1.0e-6), label=r'$a=0$ (affine)')
plt.plot(x,f(-1.0), label=r'$a=-1$ (convex)')
plt.legend()
plt.xlabel(r'$x$')
plt.ylabel(r'$u(x)$')
plt.tight_layout()
plt.savefig('ExponentialUtility.png')
| ExponentialUtility/Utility.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + jupyter={"outputs_hidden": true}
# !pip install xlrd
# !pip install nltk
# !pip install gensim
# !pip install openpyxl
# !pip install spacy
# !python -m spacy download en_core_web_sm
# -
# ### Import libraries
# +
import numpy as np
import tqdm
import os
import pandas as pd
import nltk
import re
import gensim
from gensim.utils import simple_preprocess
from nltk.corpus import stopwords
import spacy
import gensim.corpora as corpora
from pprint import pprint
from gensim.models import CoherenceModel
import pickle
from sklearn.decomposition import LatentDirichletAllocation
from sklearn.feature_extraction.text import CountVectorizer
# -
# ### Download stopwords
nltk.download('stopwords')
nltk.download('punkt')
# ### Import data
# read xls file
df = pd.read_excel('Final.xlsx', sheet_name='Data',engine='openpyxl')
# if above import doesn't work, use the one below
#df = pd.read_excel('Final.xlsx', sheet_name='Data')
df = df.astype(str)
print(df.shape)
df.head()
# ### Preprocessing data
# remove duplicates and n/a
df2 = df.dropna(how='all',subset=['body'])
df2 = df2.drop_duplicates(subset=['link', 'body'], keep='first')
df2.reset_index(drop=True, inplace=True)
df2
# +
# Remove punctuation
#df2['body_processed'] = df2['body'].map(lambda x: re.sub('[,\.!?]', '', x))
df2['body_processed'] = df2['body'].map(lambda x: re.sub('^[a-z]+$', '', x))
# Convert the titles to lowercase
df2['body_processed'] = df2['body_processed'].map(lambda x: x.lower())
# Print out the first rows of papers
df2['body_processed'].head()
# +
def sent_to_words(sentences):
for sentence in sentences:
yield(gensim.utils.simple_preprocess(str(sentence), deacc=True)) # deacc=True removes punctuations
data = df2.body_processed.values.tolist()
data_words = list(sent_to_words(data))
# -
# ### Bigrams/Trigrams
# +
# Build the bigram and trigram models
bigram = gensim.models.Phrases(data_words, min_count=5, threshold=100) # higher threshold fewer phrases.
trigram = gensim.models.Phrases(bigram[data_words], threshold=100)
# Faster way to get a sentence clubbed as a trigram/bigram
bigram_mod = gensim.models.phrases.Phraser(bigram)
trigram_mod = gensim.models.phrases.Phraser(trigram)
# -
# stopwords
stop_words = stopwords.words('english')
stop_words.extend(['company', 'market', 'week', 'month', 'year', 'country', 'include', 'business', 'll', 've', '0', '1', '10', '2', '2012', '2013', '2014', '3', '4', '5', '6', '7', '8', '9', 'a', 'as', 'able', 'about', 'above', 'abst', 'accordance', 'according', 'accordingly', 'across', 'act', 'actually', 'added', 'adj', 'affected', 'affecting', 'affects', 'after', 'afterwards', 'again', 'against', 'ah', 'aint', 'ako', 'all', 'allow', 'allows', 'almost', 'alone', 'along', 'already', 'also', 'although', 'always', 'am', 'among', 'amongst', 'amoungst', 'amount', 'an', 'and', 'ang', 'announce', 'another', 'any', 'anybody', 'anyhow', 'anymore', 'anyone', 'anything', 'anyway', 'anyways', 'anywhere', 'apart', 'apparently', 'appear', 'appreciate', 'appropriate', 'approximately', 'are', 'aren', 'arent', 'arent', 'arise', 'around', 'as', 'aside', 'ask', 'asking', 'associated', 'at', 'august', 'auth', 'available', 'away', 'awfully', 'b', 'back', 'be', 'became', 'because', 'become', 'becomes', 'becoming', 'been', 'before', 'beforehand', 'begin', 'beginning', 'beginnings', 'begins', 'behind', 'being', 'believe', 'below', 'beside', 'besides', 'best', 'better', 'between', 'beyond', 'biol', 'both', 'bottom', 'brief', 'briefly', 'but', 'by', 'c', 'cmon', 'cs', 'ca', 'call', 'came', 'can', 'cant', 'cannot', 'cant', 'cause', 'causes', 'certain', 'certainly', 'changes', 'clearly', 'co', 'com', 'come', 'comes', 'con', 'concerning', 'consequently', 'consider', 'considering', 'contain', 'containing', 'contains', 'corresponding', 'could', 'couldnt', 'couldnt', 'course', 'currently', 'd', 'date', 'day', 'de', 'december', 'definitely', 'describe', 'described', 'despite', 'detail', 'did', 'didnt', 'different', 'do', 'does', 'doesnt', 'doing', 'don', 'dont', 'done', 'down', 'downwards', 'due', 'during', 'e', 'each', 'ed', 'edu', 'effect', 'eg', 'eight', 'eighty', 'either', 'eleven', 'else', 'elsewhere', 'empty', 'end', 'ending', 'enough', 'entirely', 'especially', 'et', 'et-al', 'etc', 'even', 'ever', 'every', 'everybody', 'everyone', 'everything', 'everywhere', 'ex', 'exactly', 'example', 'except', 'f', 'far', 'february', 'few', 'ff', 'fifteen', 'fifth', 'fifty', 'fill', 'find', 'fire', 'first', 'five', 'fix', 'followed', 'following', 'follows', 'for', 'former', 'formerly', 'forth', 'forty', 'found', 'four', 'fri', 'friday', 'from', 'front', 'full', 'further', 'furthermore', 'g', 'gave', 'get', 'gets', 'getting', 'give', 'given', 'gives', 'giving', 'go', 'goes', 'going', 'gone', 'got', 'gotten', 'government', 'greetings', 'h', 'had', 'hadnt', 'happens', 'hardly', 'has', 'hasnt', 'hasnt', 'have', 'havent', 'having', 'he', 'hed', 'hell', 'hes', 'hed', 'hello', 'help', 'hence', 'her', 'here', 'heres', 'hereafter', 'hereby', 'herein', 'heres', 'hereupon', 'hers', 'herself', 'hes', 'hi', 'hid', 'him', 'himself', 'his', 'hither', 'home', 'homepage', 'hopefully', 'how', 'hows', 'howbeit', 'however', 'hundred', 'i', 'id', 'ill', 'im', 'ive', 'id', 'ie', 'if', 'ignored', 'im', 'immediate', 'immediately', 'importance', 'important', 'in', 'inasmuch', 'inc', 'indeed', 'index', 'indicate', 'indicated', 'indicates', 'information', 'inner', 'inquirer', 'insofar', 'instead', 'into', 'invention', 'inward', 'is', 'isnt', 'it', 'itd', 'itll', 'its', 'itd', 'its', 'itself', 'j', 'january', 'july', 'june', 'just', 'k', 'keep', 'keeps', 'kept', 'kg', 'km', 'know', 'known', 'knows', 'ko', 'l', 'largely', 'last', 'lately', 'later', 'latter', 'latterly', 'least', 'less', 'lest', 'let', 'lets', 'lets', 'like', 'liked', 'likely', 'line', 'little', 'look', 'looking', 'looks', 'ltd', 'm', 'made', 'mainly', 'make', 'makes', 'manila', 'many', 'march', 'may', 'maybe', 'me', 'mean', 'means', 'meantime', 'meanwhile', 'merely', 'mg', 'might', 'mill', 'million', 'mine', 'miss', 'ml', 'mon', 'monday', 'more', 'moreover', 'most', 'mostly', 'move', 'mr', 'mrs', 'much', 'mug', 'must', 'mustnt', 'my', 'myself', 'n', 'na', 'name', 'namely', 'nay', 'nd', 'near', 'nearly', 'necessarily', 'necessary', 'need', 'needs', 'neither', 'never', 'nevertheless', 'new', 'newsinfo', 'next', 'ng', 'nine', 'ninety', 'no', 'nobody', 'non', 'none', 'nonetheless', 'noone', 'nor', 'normally', 'nos', 'not', 'noted', 'nothing', 'novel', 'november', 'now', 'nowhere', 'o', 'obtain', 'obtained', 'obviously', 'october', 'of', 'off', 'often', 'oh', 'ok', 'okay', 'old', 'omitted', 'on', 'once', 'one', 'ones', 'only', 'onto', 'or', 'ord', 'other', 'others', 'otherwise', 'ought', 'our', 'ours', 'ourselves', 'out', 'outside', 'over', 'overall', 'owing', 'own', 'p', 'page', 'pages', 'part', 'particular', 'particularly', 'past', 'people', 'per', 'percent', 'perhaps', 'philippine', 'photo', 'placed', 'please', 'plus', 'pm', 'police', 'poorly', 'possible', 'possibly', 'potentially', 'pp', 'predominantly', 'present', 'presumably', 'previously', 'primarily', 'probably', 'promptly', 'proud', 'provides', 'put', 'q', 'que', 'quickly', 'quite', 'qv', 'r', 'ran', 'rappler', 'rapplercom', 'rather', 'rd', 're', 'readily', 'really', 'reasonably', 'recent', 'recently', 'ref', 'refs', 'regarding', 'regardless', 'regards', 'related', 'relatively', 'research', 'respectively', 'resulted', 'resulting', 'results', 'reuters', 'right', 'run', 's', 'sa', 'said', 'same', 'sat', 'saturday', 'saw', 'say', 'saying', 'says', 'sec', 'second', 'secondly', 'section', 'see', 'seeing', 'seem', 'seemed', 'seeming', 'seems', 'seen', 'self', 'selves', 'sensible', 'sent', 'september', 'serious', 'seriously', 'seven', 'several', 'shall', 'shant', 'she', 'shed', 'shell', 'shes', 'shed', 'shes', 'should', 'shouldnt', 'show', 'showed', 'shown', 'showns', 'shows', 'side', 'significant', 'significantly', 'similar', 'similarly', 'since', 'sincere', 'singapore', 'six', 'sixty', 'slightly', 'so', 'some', 'somebody', 'somehow', 'someone', 'somethan', 'something', 'sometime', 'sometimes', 'somewhat', 'somewhere', 'soon', 'sorry', 'specifically', 'specified', 'specify', 'specifying', 'sports', 'still', 'stop', 'stories', 'story', 'strongly', 'sub', 'substantially', 'successfully', 'such', 'sufficiently', 'suggest', 'sunday', 'sup', 'sure', 't', 'ts', 'take', 'taken', 'taking', 'tell', 'ten', 'tends', 'th', 'than', 'thank', 'thanks', 'thanx', 'that', 'thatll', 'thats', 'thatve', 'thats', 'the', 'their', 'theirs', 'them', 'themselves', 'then', 'thence', 'there', 'therell', 'theres', 'thereve', 'thereafter', 'thereby', 'thered', 'therefore', 'therein', 'thereof', 'therere', 'theres', 'thereto', 'thereupon', 'these', 'they', 'theyd', 'theyll', 'theyre', 'theyve', 'theyd', 'theyre', 'thickv', 'thin', 'think', 'third', 'this', 'thorough', 'thoroughly', 'those', 'thou', 'though', 'thoughh', 'thousand', 'three', 'throug', 'through', 'throughout', 'thru', 'thu', 'thursday', 'thus', 'til', 'time', 'tip', 'to', 'together', 'told', 'too', 'took', 'top', 'toward', 'towards', 'tried', 'tries', 'truly', 'try', 'trying', 'ts', 'tue', 'tuesday', 'tweet', 'twelve', 'twenty', 'twice', 'two', 'u', 'un', 'under', 'unfortunately', 'unless', 'unlike', 'unlikely', 'until', 'unto', 'up', 'upon', 'ups', 'us', 'use', 'used', 'useful', 'usefully', 'usefulness', 'uses', 'using', 'usually', 'v', 'value', 'various', 'very', 'via', 'viz', 'vol', 'vols', 'vs', 'w', 'want', 'wants', 'was', 'wasnt', 'wasnt', 'way', 'we', 'wed', 'well', 'were', 'weve', 'wed', 'wednesday', 'welcome', 'well', 'went', 'were', 'werent', 'werent', 'what', 'whatll', 'whats', 'whatever', 'whats', 'when', 'whens', 'whence', 'whenever', 'where', 'wheres', 'whereafter', 'whereas', 'whereby', 'wherein', 'wheres', 'whereupon', 'wherever', 'whether', 'which', 'while', 'whim', 'whither', 'who', 'wholl', 'whos', 'whod', 'whoever', 'whole', 'whom', 'whomever', 'whos', 'whose', 'why', 'whys', 'widely', 'will', 'willing', 'wish', 'with', 'within', 'without', 'wont', 'wonder', 'wont', 'words', 'world', 'would', 'wouldnt', 'wouldnt', 'www', 'x', 'y', 'yahoo', 'year', 'years', 'yes', 'yet', 'you', 'youd', 'youll', 'youre', 'youve', 'youd', 'your', 'youre', 'yours', 'yourself', 'yourselves', 'yung', 'z', 'zero'])
# ### Further preprocessing
# +
# Define functions for stopwords, bigrams, trigrams and lemmatization
def remove_stopwords(texts):
return [[word for word in simple_preprocess(str(doc)) if word not in stop_words] for doc in texts]
def make_bigrams(texts):
return [bigram_mod[doc] for doc in texts]
def make_trigrams(texts):
return [trigram_mod[bigram_mod[doc]] for doc in texts]
def lemmatization(nlp, texts, allowed_postags=['NOUN', 'ADJ', 'VERB', 'ADV']):
texts_out = []
for sent in texts:
doc = nlp(" ".join(sent))
texts_out.append([token.lemma_ for token in doc if token.pos_ in allowed_postags])
return texts_out
# +
# Remove Stop Words
data_words_nostops = remove_stopwords(data_words)
## Form Bigrams
#data_words_bigrams = make_bigrams(data_words_nostops)
# Form Bigrams
data_words_trigrams = make_trigrams(data_words_nostops)
# Initialize spacy 'en' model, keeping only tagger component (for efficiency)
nlp = spacy.load("en_core_web_sm", disable=['parser', 'ner'])
# Do lemmatization keeping only noun, adj, vb, adv
data_lemmatized = lemmatization(nlp, data_words_trigrams, allowed_postags=['NOUN', 'ADJ', 'VERB', 'ADV'])
# -
# ### Create Corpus
# +
# Create Dictionary
id2word = corpora.Dictionary(data_lemmatized)
# Create Corpus
texts = data_lemmatized
# Term Document Frequency
corpus = [id2word.doc2bow(text) for text in texts]
# View
print(corpus[0][0][:30])
# +
def save_obj(obj, name ):
with open('model/' + name +'_tri.pkl', 'wb') as f:
pickle.dump(obj, f, pickle.HIGHEST_PROTOCOL)
def load_obj(name):
with open('model/' + name + '_tri.pkl', 'rb') as f:
return pickle.load(f)
# -
save_obj(corpus, 'corpus')
save_obj(id2word, 'id2word')
# ### Test LDA model
# Build LDA model
lda_model = gensim.models.LdaMulticore(corpus=corpus,
id2word=id2word,
num_topics=20,
random_state=100,
chunksize=100,
passes=10,
per_word_topics=True)
# Print the Keyword in the 10 topics
pprint(lda_model.print_topics())
doc_lda = lda_model[corpus]
# Compute Coherence Score
coherence_model_lda = CoherenceModel(model=lda_model, texts=texts,
dictionary=id2word, coherence='c_v')
print('Coherence Score: ', coherence_model_lda.get_coherence())
# ### Hyper-parameter tuning
# supporting function
def compute_coherence_values(corpus, dictionary, k, a, e):
lda_model = gensim.models.LdaMulticore(corpus=corpus,
id2word=dictionary,
num_topics=k,
random_state=100,
chunksize=100,
passes=10,
alpha=a,
eta=e)
coherence_model_lda = CoherenceModel(model=lda_model, texts=texts, dictionary=id2word, coherence='c_v')
return coherence_model_lda.get_coherence()
# +
# Topics range
min_topics = 10
max_topics = 25
step_size = 2
topics_range = range(min_topics, max_topics, step_size)
# alpha parameter
alpha = list(np.arange(0.01, 1, 0.25))
alpha.append('symmetric')
alpha.append('asymmetric')
# eta parameter
eta = list(np.arange(0.01, 1, 0.25))
eta.append('symmetric')
# Validation sets
num_of_docs = len(corpus)
corpus_sets = [gensim.utils.ClippedCorpus(corpus, int(num_of_docs*0.75)),
corpus]
corpus_title = ['75% Corpus', '100% Corpus']
model_results = {'Validation_Set': [],
'Topics': [],
'Alpha': [],
'Eta': [],
'Coherence': []
}
# +
# ## running code on docker
# import os
# os.environ['CUDA_DEVICE_ORDER'] = 'PCI_BUS_ID'
# os.environ['CUDA_VISIBLE_DEVICES'] = "6"
# -
if 1 == 1:
pbar = tqdm.tqdm(total=(len(eta)*len(alpha)*len(topics_range)*len(corpus_title)))
# iterate through validation corpuses
for i in range(len(corpus_sets)):
# iterate through number of topics
for k in topics_range:
# iterate through alpha values
for a in alpha:
# iterare through eta values
for e in eta:
# get the coherence score for the given parameters
cv = compute_coherence_values(corpus=corpus_sets[i], dictionary=id2word,
k=k, a=a, e=e)
# Save the model results
model_results['Validation_Set'].append(corpus_title[i])
model_results['Topics'].append(k)
model_results['Alpha'].append(a)
model_results['Eta'].append(e)
model_results['Coherence'].append(cv)
pbar.update(1)
pd.DataFrame(model_results).to_csv('lda_tuning_results_2.csv', index=False)
pbar.close()
tuned_results_df = pd.DataFrame(data=model_results)
print(tuned_results_df.shape)
tuned_results_df = tuned_results_df.sort_values(['Coherence'], ascending=[False])
tuned_results_df.head()
# ### Select final LDA model
# +
# from the training above below are the final selected paramters
k=16
a=0.01
e=0.01
lda_model_final = gensim.models.LdaMulticore(corpus=corpus,
id2word=id2word,
num_topics=k,
random_state=100,
chunksize=100,
passes=10,
alpha=a,
eta=e)
# -
# Print the Keyword in the 10 topics
pprint(lda_model_final.print_topics())
doc_lda_final = lda_model_final[corpus]
# ### Save model to file
lda_model_final.save('model/lda_final_16.model')
# IMPORTANT
# MODEL CAN BE LOADED AS BELOW
#lda_model_load = gensim.models.LdaModel.load('model/lda_final_16.model')
# ### Visualize Results
# + jupyter={"outputs_hidden": true}
# !pip install pyLDAvis
import pyLDAvis
pyLDAvis.enable_notebook()
# -
# load the saved model
lda_model_load = gensim.models.LdaModel.load('model/lda_final_16.model')
num_topics= 16
# load corpus and id2word from pickle
# since training was done on docker and this visualozation ia performed in local system
corpus = load_obj('corpus')
id2word = load_obj('id2word')
# +
# Visualize the topics
LDAvis_data_filepath = 'model/ldavis_tuned_16.html'
# # this is a bit time consuming - make the if statement True
# # if you want to execute visualization prep yourself
if 1 == 1:
LDAvis_prepared = pyLDAvis.gensim.prepare(lda_model_load, corpus, id2word)
with open(LDAvis_data_filepath, 'wb') as f:
pickle.dump(LDAvis_prepared, f)
# load the pre-prepared pyLDAvis data from disk
with open(LDAvis_data_filepath, 'rb') as f:
LDAvis_prepared = pickle.load(f)
pyLDAvis.save_html(LDAvis_prepared, LDAvis_data_filepath)
LDAvis_prepared
# -
# ### Get dominant topic
def format_topics_sentences(ldamodel, corpus, texts):
# Init output
sent_topics_df = pd.DataFrame()
# Get main topic in each document
for i, row in enumerate(ldamodel[corpus]):
row = sorted(row, key=lambda x: (x[1]), reverse=True)
# Get the Dominant topic, Perc Contribution and Keywords for each document
for j, (topic_num, prop_topic) in enumerate(row):
if j == 0: # => dominant topic
wp = ldamodel.show_topic(topic_num)
topic_keywords = ", ".join([word for word, prop in wp])
sent_topics_df = sent_topics_df.append(pd.Series([int(topic_num), round(prop_topic,4), topic_keywords]), ignore_index=True)
else:
break
sent_topics_df.columns = ['Dominant_Topic', 'Perc_Contribution', 'Topic_Keywords']
# Add original text to the end of the output
contents = pd.Series(texts)
sent_topics_df = pd.concat([sent_topics_df, contents], axis=1)
return(sent_topics_df)
# +
df_topic_sents_keywords = format_topics_sentences(ldamodel=lda_model_final, corpus=corpus, texts=texts)
# Format
df_dominant_topic = df_topic_sents_keywords.reset_index()
df_dominant_topic.columns = ['Document_No', 'Dominant_Topic', 'Topic_Perc_Contrib', 'Keywords', 'Text']
# Show
print(df_dominant_topic.shape)
df_dominant_topic.head(10)
# -
merged = df2.merge(df_dominant_topic, left_index=True, right_index=True, how='inner')
# reformat data
merged['Dominant_Topic'] = merged['Dominant_Topic'].astype(int)
merged = merged.drop(columns=['Document_No'], axis=1)
merged = merged.drop(columns=['body_processed'], axis=1)
merged['company'] = df2['company']
# show
print(merged.shape)
merged.head()
# ### Save LDA output to excel
# +
# writing to Excel
datatoexcel = pd.ExcelWriter('model/LDA_Results.xlsx')
# write DataFrame to excel
merged.to_excel(datatoexcel, index=False, sheet_name='Data')
# save the excel
datatoexcel.save()
# -
# ### Company word cloud for keywords from dominant topics
# + jupyter={"outputs_hidden": true}
# !pip install wordcloud
import matplotlib.pyplot as plt
from wordcloud import WordCloud
# -
# create copy of dataframe and drop unnecessary columns
company_keywords_df = merged.copy()
company_keywords_df = company_keywords_df.drop(columns=['body','Dominant_Topic','Topic_Perc_Contrib','Text'], axis=1)
# reformat and group by company keywords
company_keywords_df = company_keywords_df.astype(str)
company_keywords_df = company_keywords_df.groupby('company')['Keywords'].apply(', '.join).reset_index()
company_keywords_df.head()
def show_wordcloud(company_name):
word_cloud = WordCloud(collocations = False,
background_color = 'white').generate(company_keywords_df.loc[company_keywords_df['company']==company_name, 'Keywords'].iloc[0])
plt.imshow(word_cloud, interpolation='bilinear')
plt.axis("off")
plt.show()
# ### Display wordcloud randomly
# +
import random
comps = random.sample(list(merged['company']),5)
for c in comps:
print(c)
show_wordcloud(c)
# -
| 02_Topic_modelling/01_Topic_modelling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise 2b: Modeling steady-state heat flow in the lithosphere
#
# <NAME>
#
# November 2019
#
# <<EMAIL>>
#
# ## Objectives
#
# * Learn to model heat flow in the crust and lithosphere using Python
# * Learn what typical geothermal gradients in the lithosphere look like and how they are affected by thermal parameters and the thickness of the lithosphere
#
#
# **Deadline**: **17 Jan 2020**. Hand in a version of your jupyter notebook and a short word document with answers to the assignments and the result figures of your numerical model.
#
# **Grading**: Each assignment is 1 point, for a total of 5 points.
#
# Do not hesitate to ask questions if you get stuck anywhere. You can reach me by email <<EMAIL>> or pass by at my office, room 122 in the Structural Geology dept.
#
# *Good luck !*
# ## Introduction
# In this exercise we will update the notebook that you completed in exercise 2a to model steady-state heat flow instead of groundwater flow. Heat conduction and groundwater flow are governed by very similar diffusion laws. We will adjust the parameters and boundary conditions of your groundwater model to instead calculate steady-state geothermal gradients in the lithosphere.
# ## Rewrite your notebook to model heat flow
#
# The heat flow equation that we will use in this exercise is a combination of Fouriers's law:
#
# \begin{equation}
# q = - K \dfrac{\partial T}{\partial x}
# \end{equation}
#
# and the heat balance equation:
#
# \begin{equation}
# \dfrac{\partial q}{\partial x} = W
# \end{equation}
#
# Note the similarity of Fourier's law and Darcy's law. Instead of hydraulic head (*h*) we use temperature (*T*), and instead of hydraulic conductivity we use thermal conductivity (both denoted by *K*). The source term in this case is not recharge or groundwater pumping, but the generation of heat by the decay of radioactive elements.
#
# Copy the fast version of the diffusion function (``solve_steady_state_diffusion_eq_faster``) from your notebook of exercise 2a into this notebook below.
#
# Change the function to model heat flow model by changing all the variables called ``h``, ``h_new``, ``h_old`` to ``T``, ``T_new``, ``T_old``.
# importing external python modules
# numpy for working with arrays:
import numpy as np
# and matplotlib to make nice looking figures
import matplotlib.pyplot as pl
# +
# # copy your finite difference diffusion function here...
# -
# **Assignment 1** Write down the finite difference approximation of the steady-state heat flow equation. This is the heat flow version of equation 10 in your handout of exercise 1. What units do heat flow (*q*) and heat production (*W*) have?
# ## Assigning new parameters
#
# Next we have to assign new parameter values to our notebook. We will model the geothermal gradient in the lithosphere. We will increase the size of our model (variable ``L``) to a 100 km, which is a value representative of the thickness of the continental lithosphere. To change this, copy the code block where you assign parameters in exercise 2a below and change the line assigning ``L`` in your notebook like this:
#
# ~~~~python
# L = 100000
# ~~~~
#
# For thermal conductivity we will use an initial value of 2.5 W m^-1 K^-1. Our model will be one dimensional, so the thickness parameter ``b`` should be equal to 1. And for the grid cell size of our numerical model we will use a value of 200 m (``dx = 200``). For now we will leave the source term zero, so ``W_array[:] = 0``.
# +
# # copy the code blocks where you assign parameters here....
# -
# ## Adjust the boundary conditions
#
# In contrast to the groundwater model, the right hand side / bottom of our model domain is not insulated. Instead there is a heat flux to the lithosphere from the astenosphere. The base of the lithosphere is usually defined as a thermal boundary, and this boundary is commonly assumed to be a temperature of 1300 °C. This means that we can define the right hand boundary condition as a specified temperature of 1300 °C. Change the following line in the ``solve_steady_state_diffusion_eq_faster`` function:
#
# ~~~~python
# T_new[-1] = ...
# ~~~~
#
# to
#
# ~~~~python
# T_new[-1] = 1300.0
# ~~~~
#
# This makes sure that the last node in our model always has a temperature of 1300 °C. For the top boundary we can use an average global surface temperature of 10 °C.
# ## Calculate a steady-state geothermal gradient
#
# With the newly assigned variables and boundary condition we are ready to run our numerical model and calculate an average steady-state geothermal gradient in the lithosphere.
#
# **Assignment 2** Copy the line where you run the diffusion function (``h = solve_steady....``) from exercise 2a. Run your new heat flow model. Try to experiment with the number of iterations. What number of iterations do you need (approximately) to reach a steady-state geothermal gradient?
# +
# # copy the line that calls the diffusion function here:
# -
# ## Adding realistic thermal parameters
#
# In exercise 1 we have already have set up an array for the source term (``W``), which we can now use to vary heat production in the lithosphere. As you have learned in the Python tutorial you can perform operations like assigning numbers to sections of arrays like this:
#
# ~~~~python
# W_array[10:30] = 1e-6
# ~~~~
#
# This assign a heat production of 1 x 10-6^ W m^-3 from node 10 to 30. Note: this is an example, do not insert this line into your script yet. There is an even more convenient way to assign values based on their depth. We can also assign values to parts of arrays like this:
#
# ~~~~python
# W_array[x < 10000] = 1e-6
# ~~~~
#
# This will change the heat flow values in the upper 10000 m of you model domain. For example, to assign numbers to a section running from 10000 to 20000 m, try the following:
#
# ~~~~python
# W_array[(x >= 10000) & (x < 20000)] = 1e-6
# ~~~~
#
# Note that the sign ``>=`` means larger than or equal to a number.
#
# Look up heat production for the upper crust, lower crust and the mantle from Cloetingh et al. (2005), Table 1. Use the reference at the bottom of this notebook and look up the paper on google scholar. Follow the examples above to assign heat production values for these three sections of the lithosphere. For the depth of the upper crust we can use a value of 12 km and for the depth of the lower crust 35 km. Try to place the new lines of code where you define heat production *after* the line where you calculate the first steady-state geothermal gradient (ie., after first call of the ``solve_steady_state_diffusion_eq function``). Next, add a line to calculate the new geothermal gradient including heat production:
#
# ~~~~python
# T2 = solve_steady_state_diffusion_eq_faster(dx, K, W_array, T0)
# ~~~~
#
# This makes sure that you store the new more exact geothermal gradient in a new variable ``T2``, which you can then compare to the old geothermal gradient ``T``.
#
# Note that thermal conductivity also varies in the crust and in the mantle. However, the solution we derived for steady-state groundwater flow and heat flow equations assumes that *K* is constant, since we moved *K* out of the derivative. See the handout of exercise 1, equations 5 and 6. Therefore with our current simplified model we cannot model a variable thermal conductivity.
# **Assignment 3** Run your new model with improved thermal parameters. Plot both the old and new temperature curve in the same panel. Try to explain the shape of the new geothermal gradient in a few words or sentences, why is there a curvature?
# ## Calculate heat flow
#
# We can use Fourier's law to calculate heat flow, using the values of the geothermal gradient that you just calculated. Use the following lines to calculate the gradient and the heat flow in the lithosphere:
T_gradient = (T2[1:] - T2[:-1]) / dx
q = T_gradient * K
# ## Add temperature and heat flow to a figure
#
# We will try to make a figure that shows both the change in temperature and heat flow in the lithosphere. First copy the code block that generates a figure from exercise 2a below:
# +
# # copy code block that makes a figure here:
# -
# First try to change the command where you plot temperature so that temperature is shown on the x-axis and depth (x) on the y-axis, instead of the other way around.
#
# Next, we will try to set up a figure with two panels, one for temperature and one for heat flow. Add one more panel to the figure to show temperature and heat flow side by side. The following line creates a new figure containing two panels side by side.
#
# ~~~~python
# fig, panels = pl.subplots(1, 2, figsize=(10, 6))
# ~~~~
#
# *Replace* the current line that creates the new figure with this new line. The variable ``panels`` is now a list containing two panels. Each panel can be called using either ``panels[0]`` or ``panels[1]``.
# Replace all existing lines of code where you use ``panel.`` with ``panels[0].``. Then add some lines to plot heat flow in the second panel (``panels[1]``):
#
# ~~~~python
# x_mid = (x[1:] + x[:-1]) / 2.0
# panels[1].plot(q, x_mid, color='black')
# ~~~~
#
# we can also flip the y-axis to make sure the surface (x=0) is at the top of the figure and not the bottom:
#
# ~~~~python
# panels[0].set_ylim(100000, 0)
# panels[1].set_ylim(100000, 0)
# ~~~~
# **Assignment 4** Calculate heat flow in the lithosphere and make a figure of temperature and heat flow. What percentage of the heat flow at the surface is supplied by the mantle? (hint: compare *q* at x=0 with *q* at x=mantle depth).
#
# Note: make sure that your model has reached steady state, you may need to run significantly more iterations once you add heat production.
# ## Heat flow in different geological settings
#
# The thickness of the lithosphere exerts a strong influence on geothermal gradients, because it is a thermal boundary. We will calculate representative geothermal gradients for three geological settings with a different lithosperic thickness: 1) a craton, 2) average continental crust and 3) oceanic crust and pick a fourth location that your are interested in (the Goettingen campus, mount everest, antarctica, etc...).
# Pick each of these settings and look up values for the lithosphere thickness for a location of choice. For oceanic lithosphere look up thickness in Conrad et al. (2006). Refer to Artemieva et al. (2006) for estimates of the thickness of the continental lithosphere. Estimate the thickness of the crust using Reguzzoni et al. (2013), Fig. 12. You can find these references below.
#
# The thermal parameters of oceanic and continental lithosphere also vary. For the continental lithosphere you can use the parameters that you have already assigned. The heat generation in the oceanic lithosphere is relatively low, on average 0.5 x 10^-6 W m^-3 (Allen & Allen 2005, note: this reference is not available online). The average thermal conductivity (*K*) of the oceanic lithosphere is 3.14 W m^-1 K^-1 (Stein 1992).
#
# **Assignment 5** Model heat flow for the four different geological settings, make a figure of temperature and heat flow and compare the modeled heat flow with a database of global heat flow (Davies & Davies 2010): http://www.heatflow.und.edu/. Describe in words why the heat flow and geothermal gradients vary between the different settings, and why the models match the global heat flow data or why there is a difference.
# ## References
#
# <NAME>., <NAME>., 2005. Basin analysis: principles and applications. Blackwell publishing, Oxford.
#
# <NAME>., 2006. Global thermal model TC1 for the continental lithosphere: Implications for lithosphere secular evolution. Tectonophysics 416, 245–277.
#
# <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., 2005. Intraplate deformation and 3D rheological structure of the Rhine Rift System and adjacent areas of the northern Alpine foreland. Int. J. Earth Sci. 94, 758–778.
#
# <NAME>., <NAME>., 2006. Influence of continental roots and asthenosphere on plate-mantle coupling. Geophys. Res. Lett. 33, 2–5. doi:10.1029/2005GL025621
#
# <NAME>., <NAME>., 2010. Earth’s surface heat flux. Solid Earth 1, 5–24. doi:10.5194/se-1-5-2010
#
# <NAME>., <NAME>., <NAME>., 2013. Global moho from the combination of the CRUST2.0 model and GOCE data. Geophys. J. Int. 195, 222–237. doi:10.1093/gji/ggt247
#
# <NAME>., <NAME>., 1992. A model for the global variation in oceanic depth and heat flow with lithospheric age. Nature 356, 133–135. doi:10.1038/359123a0
#
# (note use google scholar to find any of these papers, https://scholar.google.com/)
| exercises/exercise_2_steady_state_flow_models/.ipynb_checkpoints/exercise_2b-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# split_at_heading: true
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/butchland/build-your-own-image-classifier/blob/master/colab-test-image-classifier.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="jaglhNLw3id1"
# # Test your Image Classifier
#
# ## Instructions
#
# 1. In the **Specify Project Name** section below, fill out the project name first _(this is the name of the project you used in the previous notebook. If you didn't change the name of the default project in the previous notebook, you shouldn't have to change the default project name here either so just leave the project name as is)_.
#
# This notebook assumes that you have already built and exported your image classifier (i.e. the _`export.pkl`_ file already been saved to your Google Drive under the _`/My Drive/build-your-own-image-classifier/models/pets`_ directory or its equivalent.
#
# If the exported image classifier does not exist, this will trigger an error. Please make sure to run the previous notebook ([Build your Image Classifier](https://colab.research.google.com/github/butchland/build-your-own-image-classifier/blob/master/colab-build-image-classifier.ipynb)) before running this one.
#
# 1. Click on the `Connect` button on the top right area of the page. This will change into a checkmark with the RAM and Disk health bars once the connection is complete.
# 1. Press `Cmd/Ctrl+F9` or Click on the menu `Runtime/Run all`
# 1. Click on the link to `accounts.google.com` that appears and login in to your Google Account if neccessary or select the Google Account to use for your Google Drive. (This will open a new tab)
# 1. Authorize `Google Drive File Stream` to access your Google Drive.
#
# 1. Copy the generated authentication token and paste it on the input box that appears.
#
# 1. Once the text 'RUNNING APPLICATION...' is displayed at the **Run App** section near the bottom of the notebook, click on the **`Upload`** button to upload an image and click on the **`Classify`** button to see if your images have been classified correctly.
#
# 1. Once you are satisfied that the application is running correctly, you can click on the menu `Runtime/Factory reset runtime` and click `Yes` on the dialog box to end your session.
#
# + [markdown] id="bcSBCY-uBh-0"
# ## What is going on?
#
# This section explains the code behind this notebook
#
# _(Click on SHOW CODE to display the code)_
# + [markdown] id="Bj32tdG9CAFo"
# ### Connect to your Google Drive
#
# We'll need to connect to your Google Drive in order to retrieve your exported image classifier.
# + id="8s2G8vXx258T"
#@title {display-mode: "form"}
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="K9QxPd3QCrUz"
# ### Specify Project Name and Dataset Type
#
# Fill out the `project name` -- the project name should be the same one used as the project name used in the previous notebook.
#
# + id="etO-wuOqw1hN"
#@title Enter your project name {display-mode: "form"}
project = "pets" #@param {type: "string"}
# + [markdown] id="Sks-KHVGDzeh"
# ### Install Python Packages
#
# Install all the python packages to run your app
# + id="nnNp9sOhluAP"
#@title {display-mode: "form"}
# !pip install -Uqq fastai --upgrade
# + [markdown] id="xH1ApfyFtTyO"
# ### Copy your Image Classifier from Google Drive
# + id="JMLn_leFtgM_"
#@title {display-mode: "form"}
file_name = f'export.pkl'
folder_path = f'build-your-own-image-classifier/models/{project}'
# !cp /content/drive/My\ Drive/{folder_path}/{file_name} /content/.
# + id="o0xKg1RZiAwK"
#@title {display-mode: "form"}
from fastai.vision.all import *
from fastai.vision.widgets import *
from ipywidgets import widgets
path = Path(f'/content')
Path.BASE_PATH = path
if not (path/file_name).is_file():
raise RuntimeError("Could not find export.pkl -- Please run notebook to build your classifier first!")
learn = load_learner(path/file_name)
# + id="jw1mprNPmYrq"
#@title {display-mode: "form"}
# !mkdir -p /content/images
# !curl -o /content/images/purple_dog.jpg -s https://raw.githubusercontent.com/butchland/build-your-own-image-classifier/master/images/purple_dog.jpg
# + [markdown] id="mH3pwpMHiAy7"
# ### Run App
#
# We will now run the app which should show below.
# + id="PMmZaHnwtKyc"
#@title {display-mode: "form"}
print('RUNNING APPLICATION...')
# + id="JAJdyynpp_PS"
#@title {display-mode: "form"}
btn_upload = SimpleNamespace(data = ['/content/images/purple_dog.jpg'])
img = PILImage.create(btn_upload.data[-1])
out_pl = widgets.Output()
out_pl.clear_output()
with out_pl: display(img.to_thumb(128,128))
pred,pred_idx,probs = learn.predict(img)
lbl_pred = widgets.Label()
lbl_pred.value = f'Prediction: {pred}; Probability: {probs[pred_idx]:.04f}'
btn_run = widgets.Button(description='Classify')
def on_click_classify(change):
img = PILImage.create(btn_upload.data[-1])
out_pl.clear_output()
with out_pl: display(img.to_thumb(128,128))
pred,pred_idx,probs = learn.predict(img)
lbl_pred.value = f'Prediction: {pred}; Probability: {probs[pred_idx]:.04f}'
btn_run.on_click(on_click_classify)
#Putting back btn_upload to a widget for next cell
btn_upload = widgets.FileUpload()
#hide_output
VBox([widgets.Label('UPLOAD AN IMAGE AND CLASSIFY!'),
btn_upload, btn_run, out_pl, lbl_pred])
# + id="s0mFpvLvwZJY"
#@title {display-mode: "form"}
print("DONE! DONE! DONE!")
print("Make sure to end your session (Click on menu Runtime/Factory reset runtime and click 'Yes' on the dialog box to end your session)")
print("before closing this notebook.")
| colab-test-image-classifier.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.2
# language: julia
# name: julia-1.5.2
# ---
using Revise
using MDToolbox, PyPlot, Statistics, LinearAlgebra
PyPlot.plt.style.use("seaborn-colorblind")
ENV["COLUMNS"] = 110; #display width for MDToolbox
# -----
# +
pdb_xtal = readpdb("/data/higashi/grest_4krn_swissmodel_DIHEDRAL/ground_truth.pdb");
pdb_init = readpdb("/data/higashi/grest_4krn_swissmodel_DIHEDRAL/prot.pdb");
ta1 = readdcd("/data/higashi/grest_4krn_swissmodel_DIHEDRAL/07_remd_convert/run_param1.dcd", isbox=false);
#ta2 = readdcd("/data/yasu/vhh_grest/grest_4krn_swissmodel/08_remd_convert/run_param1.dcd", isbox=false);
#ta3 = readdcd("/data/yasu/vhh_grest/grest_4krn_swissmodel/09_remd_convert/run_param1.dcd", isbox=false);
#ta4 = readdcd("/data/yasu/vhh_grest/grest_4krn_swissmodel/10_remd_convert/run_param1.dcd", isbox=false);
#ta5 = readdcd("/data/yasu/vhh_grest/grest_4krn_swissmodel/11_remd_convert/run_param1.dcd", isbox=false);
#ta = [pdb_init; ta1; ta2; ta3; ta4; ta5];
#ta = [pdb_init; ta1; ta2; ta3; ta4];
ta = [pdb_init; ta1]
# -
pdb_xtal = pdb_xtal["(atomname CA) and (resid 100:120)"]
pdb_init = pdb_init["(atomname CA) and (resid 100:120)"]
ta = ta["(atomname CA) and (resid 100:120)"];
#ta = ta["(atomname CA) and (resid 20:125)"];
x_xtal = MDToolbox.compute_distancemap(pdb_xtal);
x_init = MDToolbox.compute_distancemap(pdb_init);
X = MDToolbox.compute_distancemap(ta);
X_mean = mean(X, dims=1)
X .= X .- X_mean
e = eigen(X' * X ./ size(X, 1))
lambda = e.values[end:-1:1]
W = e.vectors[:, end:-1:1]
P = X * W
p_xtal = (x_xtal .- X_mean) * W
p_init = (x_init .- X_mean) * W
#fig, ax = subplots(figsize=(13, 5));
figure(figsize=(13,5),)
plot((1:size(P, 1)).*0.25*0.001, P[:, 1])
plot((1:size(P, 1)).*0.25*0.001, ones(size(P,1))*p_xtal[1], color=:red)
plot((1:size(P, 1)).*0.25*0.001, ones(size(P,1))*p_init[1], color=:green)
#ax[:set_xlim]([0,100])
xlim([0, 25])
ylabel("1st principal component [Å]",fontsize=20);
xlabel("time [ns]",fontsize=20);
xticks(fontsize=20);
yticks(fontsize=20);
savefig("timeseries_grest.png", dpi=350, bbox_inches="tight");
P
# +
fig, ax = subplots(figsize=(7, 6));
ax.scatter(P[:, 1], P[:, 2], s=2.0);
ax.scatter(p_xtal[1], p_xtal[2], s=100.0, color=:red);
ax.scatter(p_init[1, 1], p_init[1, 2], s=100.0, color=:green);
xlabel("1st principal component [Å]",fontsize=20);
ylabel("2nd principal component [Å]",fontsize=20);
# detailed options (not necessary, just for reference)
#ax.set(xlim=[-20, 10], ylim=[-10, 15]);
#ax.xaxis.set_tick_params(which="major",labelsize=15);
#ax.yaxis.set_tick_params(which="major",labelsize=15);
#ax.grid(linestyle="--", linewidth=0.5);
#tight_layout();
xticks(fontsize=20);
yticks(fontsize=20);
savefig("scatter_grest.png", dpi=350);
# -
#figure(figsize=(13,5),)
figure(figsize=(8,5),)
ta_fitted = superimpose(pdb_xtal, ta);
rmsd = compute_rmsd(pdb_xtal, ta_fitted)
plot((1:size(P, 1)).*0.25*0.001, rmsd)
xticks(fontsize=20);
yticks(fontsize=20);
ylabel("RMSD [Å]",fontsize=20);
xlabel("time [ns]",fontsize=20);
ylim([1.2, 6])
#ylim([1.5, 6.6])
xlim([0, 25])
savefig("rmsd.png", dpi=350, bbox_inches="tight");
| 4krn_dihedral/analysis/.ipynb_checkpoints/analysis_DIHEDRAL-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Voronoi Tesselation with color
# +
import json
import random
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from scipy.spatial import Voronoi
from shapely.geometry import Polygon
np.random.seed(783)
# -
def voronoi_finite_polygons_2d(vor, radius=None):
"""
Reconstruct infinite voronoi regions in a 2D diagram to finite
regions.
Parameters
----------
vor : Voronoi
Input diagram
radius : float, optional
Distance to 'points at infinity'.
Returns
-------
regions : list of tuples
Indices of vertices in each revised Voronoi regions.
vertices : list of tuples
Coordinates for revised Voronoi vertices. Same as coordinates
of input vertices, with 'points at infinity' appended to the
end.
"""
if vor.points.shape[1] != 2:
raise ValueError("Requires 2D input")
new_regions = []
new_vertices = vor.vertices.tolist()
center = vor.points.mean(axis=0)
if radius is None:
radius = vor.points.ptp().max()*2
# Construct a map containing all ridges for a given point
all_ridges = {}
for (p1, p2), (v1, v2) in zip(vor.ridge_points, vor.ridge_vertices):
all_ridges.setdefault(p1, []).append((p2, v1, v2))
all_ridges.setdefault(p2, []).append((p1, v1, v2))
# Reconstruct infinite regions
for p1, region in enumerate(vor.point_region):
vertices = vor.regions[region]
if all(v >= 0 for v in vertices):
# finite region
new_regions.append(vertices)
continue
# reconstruct a non-finite region
ridges = all_ridges[p1]
new_region = [v for v in vertices if v >= 0]
for p2, v1, v2 in ridges:
if v2 < 0:
v1, v2 = v2, v1
if v1 >= 0:
# finite ridge: already in the region
continue
# Compute the missing endpoint of an infinite ridge
t = vor.points[p2] - vor.points[p1] # tangent
t /= np.linalg.norm(t)
n = np.array([-t[1], t[0]]) # normal
midpoint = vor.points[[p1, p2]].mean(axis=0)
direction = np.sign(np.dot(midpoint - center, n)) * n
far_point = vor.vertices[v2] + direction * radius
new_region.append(len(new_vertices))
new_vertices.append(far_point.tolist())
# sort region counterclockwise
vs = np.asarray([new_vertices[v] for v in new_region])
c = vs.mean(axis=0)
angles = np.arctan2(vs[:,1] - c[1], vs[:,0] - c[0])
new_region = np.array(new_region)[np.argsort(angles)]
# finish
new_regions.append(new_region.tolist())
return new_regions, np.asarray(new_vertices)
# +
with open('../data/general/matches_2017_2018_v1.json') as matches_json:
matches = pd.read_json(matches_json)
match_id = random.choice(matches['id'])
x_position = np.load('../data/match_' + str(match_id) + '/x_position.npy')
y_hasball = np.load('../data/match_' + str(match_id) + '/y_player_data.npy')
y_pass_data = np.load('../data/match_' + str(match_id) + '/y_pass_data.npy')
y_pass_pred = np.load('../data/match_' + str(match_id) + '/y_pass_pred.npy')
y_pass_time = np.load('../data/match_' + str(match_id) + '/y_pass_time.npy')
y_pass_controller = np.arange(y_pass_data.shape[0])
pass_id = random.choice(y_pass_controller)
min_x = 0
max_x = 105
min_y = 0
max_y = 68
box = Polygon([[min_x, min_y], [min_x, max_y], [max_x, max_y], [max_x, min_y]])
for index in range(y_pass_time[pass_id]-5, y_pass_time[pass_id]+1):
if index == y_pass_time[pass_id]:
hasball = y_pass_pred[pass_id]
hasball = np.insert(hasball, 0, 0, axis=0)
else:
hasball = y_hasball[index]
controller = np.arange(len(x_position[index]))
deleted = []
for i in range(1, len(x_position[index])):
if x_position[index][i][0] == 0 and x_position[index][i][1] == 0:
deleted.append(i)
deleted.append(0)
points = np.delete(x_position[index], deleted, axis=0)
hasball = np.delete(hasball, deleted, axis=0)
control = np.delete(controller, deleted, axis=0)
# compute Voronoi tesselation
vor = Voronoi(points)
regions, vertices = voronoi_finite_polygons_2d(vor)
# colorize
for region in regions:
polygon = vertices[region]
# Clipping polygon
poly = Polygon(polygon)
poly = poly.intersection(box)
polygon = [p for p in poly.exterior.coords]
plt.fill(*zip(*polygon), alpha=0.3, c=np.random.rand(3,))
for i in range(len(points)):
if control[i] < 15:
if hasball[i] == 1:
plt.scatter(points[i:i+1, 0], points[i:i+1, 1], marker='^', c='red')
else:
plt.scatter(points[i:i+1, 0], points[i:i+1, 1], marker='o', c='red')
else:
if hasball[i] == 1:
plt.scatter(points[i:i+1, 0], points[i:i+1, 1], marker='^', c='blue')
else:
plt.scatter(points[i:i+1, 0], points[i:i+1, 1], marker='o', c='blue')
plt.scatter(x_position[index][0][0], x_position[index][0][1], marker='s', c='black')
plt.axis('equal')
plt.xlim([-5, 110])
plt.ylim([-5, 73])
plt.savefig('../graphs/voronoi_match_' + str(match_id) + '_second_' + str(index) + '.png')
plt.show()
# -
# Voronoi Tesselation without color
# +
import matplotlib.pyplot as pl
import numpy as np
import scipy as sp
import scipy.spatial
import sys
id = 60561
x_position = np.load('../data/match_' + str(id) + '/x_position.npy')
deleted = []
for i in range(29):
if x_position[65][i][0] == 0:
deleted.append(i)
towers = np.delete(x_position[65], deleted, axis=0)
print(towers)
eps = sys.float_info.epsilon
bounding_box = np.array([0, 105, 0, 68]) # [x_min, x_max, y_min, y_max]
def in_box(towers, bounding_box):
return np.logical_and(np.logical_and(bounding_box[0] <= towers[:, 0],
towers[:, 0] <= bounding_box[1]),
np.logical_and(bounding_box[2] <= towers[:, 1],
towers[:, 1] <= bounding_box[3]))
def voronoi(towers, bounding_box):
# Select towers inside the bounding box
i = in_box(towers, bounding_box)
# Mirror points
points_center = towers[i, :]
points_left = np.copy(points_center)
points_left[:, 0] = bounding_box[0] - (points_left[:, 0] - bounding_box[0])
points_right = np.copy(points_center)
points_right[:, 0] = bounding_box[1] + (bounding_box[1] - points_right[:, 0])
points_down = np.copy(points_center)
points_down[:, 1] = bounding_box[2] - (points_down[:, 1] - bounding_box[2])
points_up = np.copy(points_center)
points_up[:, 1] = bounding_box[3] + (bounding_box[3] - points_up[:, 1])
points = np.append(points_center,
np.append(np.append(points_left,
points_right,
axis=0),
np.append(points_down,
points_up,
axis=0),
axis=0),
axis=0)
# Compute Voronoi
vor = sp.spatial.Voronoi(points)
# Filter regions
regions = []
for region in vor.regions:
flag = True
for index in region:
if index == -1:
flag = False
break
else:
x = vor.vertices[index, 0]
y = vor.vertices[index, 1]
if not(bounding_box[0] - eps <= x and x <= bounding_box[1] + eps and
bounding_box[2] - eps <= y and y <= bounding_box[3] + eps):
flag = False
break
if region != [] and flag:
regions.append(region)
vor.filtered_points = points_center
vor.filtered_regions = regions
return vor
def centroid_region(vertices):
# Polygon's signed area
A = 0
# Centroid's x
C_x = 0
# Centroid's y
C_y = 0
for i in range(0, len(vertices) - 1):
s = (vertices[i, 0] * vertices[i + 1, 1] - vertices[i + 1, 0] * vertices[i, 1])
A = A + s
C_x = C_x + (vertices[i, 0] + vertices[i + 1, 0]) * s
C_y = C_y + (vertices[i, 1] + vertices[i + 1, 1]) * s
A = 0.5 * A
C_x = (1.0 / (6.0 * A)) * C_x
C_y = (1.0 / (6.0 * A)) * C_y
return np.array([[C_x, C_y]])
vor = voronoi(towers, bounding_box)
fig = pl.figure()
ax = fig.gca()
# Plot initial points
ax.plot(vor.filtered_points[:, 0], vor.filtered_points[:, 1], 'b.')
# Plot ridges points
for region in vor.filtered_regions:
vertices = vor.vertices[region, :]
ax.plot(vertices[:, 0], vertices[:, 1], 'go')
# Plot ridges
for region in vor.filtered_regions:
vertices = vor.vertices[region + [region[0]], :]
ax.plot(vertices[:, 0], vertices[:, 1], 'k-')
# Compute and plot centroids
centroids = []
for region in vor.filtered_regions:
vertices = vor.vertices[region + [region[0]], :]
centroid = centroid_region(vertices)
centroids.append(list(centroid[0, :]))
#ax.plot(centroid[:, 0], centroid[:, 1], 'r.')
print(len(centroids))
ax.set_xlim([-5, 110])
ax.set_ylim([-5, 73])
pl.savefig("bounded_voronoi.png")
sp.spatial.voronoi_plot_2d(vor)
pl.savefig("voronoi.png")
| src/visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
"""
The MIT License (MIT)
Copyright (c) 2021 NVIDIA
Permission is hereby granted, free of charge, to any person obtaining a copy of
this software and associated documentation files (the "Software"), to deal in
the Software without restriction, including without limitation the rights to
use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of
the Software, and to permit persons to whom the Software is furnished to do so,
subject to the following conditions:
The above copyright notice and this permission notice shall be included in all
copies or substantial portions of the Software.
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS
FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR
COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER
IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN
CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
"""
# This code example demonstrates how to use a convolutional neural network to solve an image classification problem, using the CIFAR-10 dataset. More context for this code example can be found in the section "Programming Example: Image Classification with a Convolutional Network" in Chapter 7 in the book Learning Deep Learning by <NAME> (ISBN: 9780137470358).
#
# The first code snippet shows the initialization code for our CNN program. Among the import statements, we now import a new layer called Conv2D, which is a 2D convolutional layer. We load and standardize the CIFAR-10 dataset.
# +
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import Flatten
from tensorflow.keras.layers import Conv2D
import numpy as np
import logging
tf.get_logger().setLevel(logging.ERROR)
EPOCHS = 128
BATCH_SIZE = 32
# Load dataset.
cifar_dataset = keras.datasets.cifar10
(train_images, train_labels), (test_images,
test_labels) = cifar_dataset.load_data()
# Standardize dataset.
mean = np.mean(train_images)
stddev = np.std(train_images)
train_images = (train_images - mean) / stddev
test_images = (test_images - mean) / stddev
print('mean: ', mean)
print('stddev: ', stddev)
# Change labels to one-hot.
train_labels = to_categorical(train_labels,
num_classes=10)
test_labels = to_categorical(test_labels,
num_classes=10)
# -
# The actual model is created by the next code snippet, which first declares a Sequential model and then adds layers. We are now working with a 2D convolutional layer, so there is no need to start with a Flatten layer because the dimensions of the input image already match the required dimension of the first layer. We tell the layer that the input shape of the image is 32×32×3. We also state that we want 64 channels, a kernel size of 5×5, a stride of (2, 2), and padding=’same’. We specify the neuron type as ReLU because that has been shown to be a good activation function. The combination of padding=’same’ and strides=(2,2) results in half as many neurons in each dimension as in the previous layer (i.e., 16×16 neurons per channel because the input image has 32×32 pixels).
#
# The next convolutional layer is similar but with a smaller kernel size. There is no need to specify the input shape—it is implicitly defined by the outputs of the previous layer. The number of neurons per channel is implicitly defined as 8×8 because the previous layer was 16×16 outputs per channel, and we choose a stride of 2, 2 for this layer as well.
#
# Before we can add the fully connected (Dense) layer, we need to flatten (convert from three dimensions to a single dimension) the outputs from the second convolutional layer. We use softmax activation for the fully connected layer so we can interpret the one-hot encoded outputs as probabilities.
#
# We finally select the categorical_crossentropy loss function and use the Adam optimizer in our call to compile. We then print out a description of the network with a call to model.summary().
#
# +
# Model with two convolutional and one fully connected layer.
model = Sequential()
model.add(Conv2D(64, (5, 5), strides=(2,2),
activation='relu', padding='same',
input_shape=(32, 32, 3),
kernel_initializer='he_normal',
bias_initializer='zeros'))
model.add(Conv2D(64, (3, 3), strides=(2,2),
activation='relu', padding='same',
kernel_initializer='he_normal',
bias_initializer='zeros'))
model.add(Flatten())
model.add(Dense(10, activation='softmax',
kernel_initializer='glorot_uniform',
bias_initializer='zeros'))
model.compile(loss='categorical_crossentropy',
optimizer='adam', metrics =['accuracy'])
model.summary()
# -
# We are now ready to train the network by calling fit() on the model.
#
history = model.fit(
train_images, train_labels, validation_data =
(test_images, test_labels), epochs=EPOCHS,
batch_size=BATCH_SIZE, verbose=2, shuffle=True)
| tf_framework/c7e2_convnet_cifar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_tensorflow_p36
# language: python
# name: conda_tensorflow_p36
# ---
# # Next-item recommendation on top of Amason’s explicit rating dataset
#
# Frame the recommendation system as a rating prediction machine learning problem and create a hybrid architecture that mixes the collaborative and content based filtering approaches:
# - Collaborative part: Predict items ratings in order to recommend to the user items that he is likely to rate high.
# - Content based: use metadata inputs (such as price and title) about items to find similar items to recommend.
#
# Create 2 explicit recommendation engine models based on 2 machine learning architecture using Keras: a matrix factorization model and a deep neural network model.
#
# For training validation and prediction, I used the electronics reviews dataset from amazon which contains explicit item ranking
#
# Compare the results of the different models and configurations to find the "best" predicting model
#
# I used the best model for recommending items to users
# +
# Check Jave version
# # !sudo yum -y update
# +
# # !sudo yum remove jre-1.7.0-openjdk -y
# -
# !java -version
# +
# # !sudo update-alternatives --config java
# -
# !pip install ipython-autotime
# +
#### To measure all running time
# https://github.com/cpcloud/ipython-autotime
# %load_ext autotime
# -
# !pip install tqdm pydot pydotplus pydot_ng
# +
# %pylab inline
import warnings
warnings.filterwarnings("ignore")
# %matplotlib inline
import re
import seaborn as sbn
import nltk
import tqdm as tqdm
import sqlite3
import pandas as pd
import numpy as np
from pandas import DataFrame
import string
import matplotlib.pyplot as plt
from math import floor,ceil
#from nltk.corpus import stopwords
#stop = stopwords.words("english")
from nltk.stem.porter import PorterStemmer
english_stemmer=nltk.stem.SnowballStemmer('english')
from nltk.tokenize import word_tokenize
from sklearn.metrics import accuracy_score, confusion_matrix,roc_curve, auc,classification_report, mean_squared_error, mean_absolute_error
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
from sklearn.model_selection import train_test_split
from sklearn import metrics
from sklearn.svm import LinearSVC
from sklearn.neighbors import NearestNeighbors
from sklearn.linear_model import LogisticRegression
from sklearn import neighbors
from scipy.spatial.distance import cosine
from sklearn.feature_selection import SelectKBest
from IPython.display import SVG
import pydot
import pydotplus
import pydot_ng
import pickle
import time
import gzip
import os
os.getcwd()
# +
# Tensorflow
import tensorflow as tf
#Keras
from keras.models import Sequential, Model, load_model, save_model
from keras.callbacks import ModelCheckpoint
from keras.layers import Dense, Activation, Dropout, Input, Masking, TimeDistributed, LSTM, Conv1D, Embedding
from keras.layers import GRU, Bidirectional, BatchNormalization, Reshape
from keras.optimizers import Adam
from keras.layers.core import Reshape, Dropout, Dense
from keras.layers.merge import Multiply, Dot, Concatenate
from keras.layers.embeddings import Embedding
from keras import optimizers
from keras.callbacks import ModelCheckpoint
from keras.utils.vis_utils import model_to_dot
# -
# ### Set GPUs
# +
#Session
from keras import backend as K
cfg = K.tf.ConfigProto()
cfg.gpu_options.per_process_gpu_memory_fraction =1 # allow all of the GPU memory to be allocated
# for 8 GPUs
# cfg.gpu_options.visible_device_list = "0,1,2,3,4,5,6,7" # "0,1"
# for 1 GPU
cfg.gpu_options.visible_device_list = "0"
#cfg.gpu_options.allow_growth = True # # Don't pre-allocate memory; dynamically allocate the memory used on the GPU as-needed
#cfg.log_device_placement = True # to log device placement (on which device the operation ran)
sess = K.tf.Session(config=cfg)
K.set_session(sess) # set this TensorFlow session as the default session for Keras
# -
print("* TF version: ", [tf.__version__, tf.test.is_gpu_available()])
print("* List of GPU(s): ", tf.config.experimental.list_physical_devices() )
print("* Num GPUs Available: ", len(tf.config.experimental.list_physical_devices('GPU')))
# +
def set_check_gpu():
os.environ["CUDA_DEVICE_ORDER"] = "PCI_BUS_ID";
# set for 8 GPUs
# os.environ["CUDA_VISIBLE_DEVICES"] = "0,1,2,3,4,5,6,7";
# set for 1 GPU
os.environ["CUDA_VISIBLE_DEVICES"] = "0";
# Tf debugging option
tf.debugging.set_log_device_placement(True)
gpus = tf.config.experimental.list_physical_devices('GPU')
if gpus:
try:
# Currently, memory growth needs to be the same across GPUs
for gpu in gpus:
tf.config.experimental.set_memory_growth(gpu, True)
logical_gpus = tf.config.experimental.list_logical_devices('GPU')
print(len(gpus), "Physical GPUs,", len(logical_gpus), "Logical GPUs")
except RuntimeError as e:
# Memory growth must be set before GPUs have been initialized
print(e)
# print(tf.config.list_logical_devices('GPU'))
print(tf.config.experimental.list_physical_devices('GPU'))
print("Num GPUs Available: ", len(tf.config.experimental.list_physical_devices('GPU')))
# -
set_check_gpu()
# reset GPU memory& Keras Session
def reset_keras():
try:
del classifier
del model
except:
pass
K.clear_session()
K.get_session().close()
# sess = K.get_session()
cfg = K.tf.ConfigProto()
cfg.gpu_options.per_process_gpu_memory_fraction
cfg.gpu_options.visible_device_list = "0,1,2,3,4,5,6,7" # "0,1"
cfg.gpu_options.allow_growth = True # dynamically grow the memory used on the GPU
sess = K.tf.Session(config=cfg)
K.set_session(sess) # set this TensorFlow session as the default session for Keras
# ## Load dataset and analysis
# #### Download and prepare Data:
# ###### 1. Read the data:
# Read the data from the electronics reviews dataset of amazon. Use the dastaset in which all users and items have at least 5 reviews ("small" dataset), or a sample of all the reviews regardless of reviews number ("large" dataset).
#
#
# - https://nijianmo.github.io/amazon/index.html
# +
import sys
# !{sys.executable} -m pip install --upgrade pip
# !{sys.executable} -m pip install sagemaker-experiments
# !{sys.executable} -m pip install pandas
# !{sys.executable} -m pip install numpy
# !{sys.executable} -m pip install matplotlib
# !{sys.executable} -m pip install boto3
# !{sys.executable} -m pip install sagemaker
# !{sys.executable} -m pip install pyspark
# !{sys.executable} -m pip install ipython-autotime
# !{sys.executable} -m pip install surprise
# !{sys.executable} -m pip install smart_open
# !{sys.executable} -m pip install pyarrow
# !{sys.executable} -m pip install fastparquet
# +
import pandas as pd
import boto3
import sagemaker
from sagemaker import get_execution_role
from sagemaker.session import Session
from sagemaker.analytics import ExperimentAnalytics
import gzip
import json
from pyspark.ml import Pipeline
from pyspark.sql.types import StructField, StructType, StringType, DoubleType
from pyspark.ml.feature import StringIndexer, VectorIndexer, OneHotEncoder, VectorAssembler
from pyspark.sql.functions import *
# spark imports
from pyspark.sql import SparkSession
from pyspark.sql.functions import UserDefinedFunction, explode, desc
from pyspark.sql.types import StringType, ArrayType
from pyspark.ml.evaluation import RegressionEvaluator
import os
import pandas as pd
from smart_open import smart_open
# from pandas_profiling import ProfileReport
# +
# # !wget http://deepyeti.ucsd.edu/jianmo/amazon/categoryFilesSmall/Clothing_Shoes_and_Jewelry_5.json.gz
# +
number_cores = 16
memory_gb = 64
spark = SparkSession \
.builder \
.appName("amazon recommendation") \
.config("spark.driver.memory", '{}g'.format(memory_gb)) \
.config("spark.master", 'local[{}]'.format(number_cores)) \
.getOrCreate()
# get spark context
sc = spark.sparkContext
# -
# #### Loading Clothing_Shoes_and_Jewelry_5.json.gz
# +
# DATA_PATH = './'
# REVIEW_DATA = 'Clothing_Shoes_and_Jewelry_5.json.gz'
# ratings = spark.read.load(DATA_PATH+REVIEW_DATA, format='json', header=True, inferSchema=True)
# print(ratings.show(3))
# print(type(ratings))
# print(ratings.head(n=2))
# +
# clean_ratings = ratings.na.drop(how='any', subset='vote')
# print(clean_ratings.columns)
# product_ratings = clean_ratings.drop(
# 'helpful',
# 'unixReviewTime',
# 'style',
# 'verified',
# 'vote'
# )
# +
# print(product_ratings.columns)
# print(product_ratings.show(3))
# print(type(product_ratings))
# +
# # !rm -rf output
# +
## DONOT USE CSV..... SUCKS....
## create csv file
# product_ratings.write.csv("./Clean_Clothing_Shoes_and_Jewelry_5_clean")
# +
# product_ratings.write.parquet("output/Clothing_Shoes_and_Jewelry_5.parquet")
# +
# # !ls -ahl output/Clothing_Shoes_and_Jewelry_5.parquet
# +
# # !aws s3 cp ./output/Clothing_Shoes_and_Jewelry_5.parquet/*.parquet s3://dse-cohort5-group1/2-Keras-DeepRecommender/dataset/Clean_Clothing_Shoes_and_Jewelry_5_clean.parquet
# -
# !aws s3 cp s3://dse-cohort5-group1/2-Keras-DeepRecommender/dataset/Clean_Clothing_Shoes_and_Jewelry_5_clean.parquet .
# !ls -alh
review_data = pd.read_parquet("./Clean_Clothing_Shoes_and_Jewelry_5_clean.parquet")
review_data.head(n=10)
review_data.shape
# Thus I have 1069568 rows and 8 columns dataset.
#
# ## 2. Arrange and clean the data
# Rearrange the columns by relevance and rename column names
review_data.columns
# +
review_data = review_data[['asin', 'image', 'summary', 'reviewText', 'overall', 'reviewerID', 'reviewerName', 'reviewTime']]
review_data.rename(columns={ 'overall': 'score','reviewerID': 'user_id', 'reviewerName': 'user_name'}, inplace=True)
#the variables names after rename in the modified data frame
list(review_data)
# -
# ## The Data
# ###### The datasets are composed of:
#
# “item_id”(“asin”): a unique id for an item - Independent explanatory variable
#
# “user_id”: a unique id for a user -Independent explanatory variable
#
# “score”(“overall”): the review score, between 1 and 5 - The dependent variable I want to predict
#
# “user_name”: the name of the user.
#
# “reviewTime”: the date when the user posted the review. Will not be used in this work.
#
# “reviewText”: For some entries, there is also a typed review. Will not be used in this work.
#
# “summary”: summary of the review. Will not be used in this work.
#
#
#
# $\underline{Investigate}$:
# +
# review_data["score"]
# -
# Ratings distribution using pandas:
review_data["score"] = review_data["score"].fillna(review_data["score"].median())
review_data["score"].describe()
# Plot the distribution
review_data["score"].hist(bins=10)
# The median in both datasets is 5. This means that the data is skewed towards high ratings.
# This is a common bias in internet ratings, where people tend to rate items that they liked, and rarely spend time to comment something they dislike or are indiferent to. This will have a huge impact on the way I model the recommendation problem.
#
# ###### Key conclusions from above:
#
# Reviews are skewed towards positive
#
# Many people agree with score 5 reviews
#
# ## Add metadata
# +
# # !wget http://deepyeti.ucsd.edu/jianmo/amazon/metaFiles/meta_Clothing_Shoes_and_Jewelry.json.gz
# +
## Read the input and convert ro DF
def parse_gz(path):
g = gzip.open(path, 'rb')
for l in g:
yield eval(l)
def convert_to_DF(path):
i = 0
df = {}
for d in parse_gz(path):
df[i] = d
i += 1
return pd.DataFrame.from_dict(df, orient='index')
# -
# ### Load meta_Clothing_Shoes_and_Jewelry.json.gz
# +
# all_info = spark.read.load('./meta_Clothing_Shoes_and_Jewelry.json.gz', format='json', header=True, inferSchema=True)
# +
# all_info.columns
# +
# print(all_info.columns)
# all_info = all_info.drop(
# 'date',
# 'details',
# 'feature',
# 'fit',
# 'main_cat',
# 'rank',
# 'similar_item',
# 'tech1',
# 'tech2'
# )
# print(all_info.columns)
# +
# # Save it as parquet format
# all_info.write.parquet("output/Cleaned_meta_Clothing_Shoes_and_Jewelry.parquet")
# +
# # !aws s3 cp ./output/Cleaned_meta_Clothing_Shoes_and_Jewelry.parquet/*.parquet s3://dse-cohort5-group1/2-Keras-DeepRecommender/dataset/Cleaned_meta_Clothing_Shoes_and_Jewelry.parquet
# +
# # !aws s3 cp s3://dse-cohort5-group1/2-Keras-DeepRecommender/dataset/Cleaned_meta_Clothing_Shoes_and_Jewelry.parquet .
# -
all_info = pd.read_parquet("./Cleaned_meta_Clothing_Shoes_and_Jewelry.parquet")
all_info.head(n=5)
# ###### Arrange and clean the data
# Cleaning, handling missing data, normalization, etc:
#
# For the algorithm in keras to work, remap all item_ids and user_ids to an interger between 0 and the total number of users or the total number of items
#
#
all_info.columns
items = all_info.asin.unique()
item_map = {i:val for i,val in enumerate(items)}
inverse_item_map = {val:i for i,val in enumerate(items)}
all_info["old_item_id"] = all_info["asin"] # copying for join with metadata
all_info["item_id"] = all_info["asin"].map(inverse_item_map)
items = all_info.item_id.unique()
print ("We have %d unique items in metadata "%items.shape[0])
all_info['description'] = all_info['description'].fillna(all_info['title'].fillna('no_data'))
all_info['title'] = all_info['title'].fillna(all_info['description'].fillna('no_data').apply(str).str[:20])
all_info['image'] = all_info['image'].fillna('no_data')
all_info['price'] = pd.to_numeric(all_info['price'],errors="coerce")
all_info['price'] = all_info['price'].fillna(all_info['price'].median())
# +
users = review_data.user_id.unique()
user_map = {i:val for i,val in enumerate(users)}
inverse_user_map = {val:i for i,val in enumerate(users)}
review_data["old_user_id"] = review_data["user_id"]
review_data["user_id"] = review_data["user_id"].map(inverse_user_map)
items_reviewed = review_data.asin.unique()
review_data["old_item_id"] = review_data["asin"] # copying for join with metadata
review_data["item_id"] = review_data["asin"].map(inverse_item_map)
items_reviewed = review_data.item_id.unique()
users = review_data.user_id.unique()
# -
print ("We have %d unique users"%users.shape[0])
print ("We have %d unique items reviewed"%items_reviewed.shape[0])
# We have 192403 unique users in the "small" dataset
# We have 63001 unique items reviewed in the "small" dataset
review_data.head(3)
#
#
# $\underline{Investigate}$:
#
# ###### Check the Distribution of number of ratings per user:
users_ratings = review_data['old_user_id'].value_counts().reset_index()
users_ratings.columns= ['old_user_id','num_ratings']
users_ratings['num_ratings'].describe()
# The distribution of number of ratings per user is very skewed in both datasets, with 50% of people having done a small number of reviews, and few made many ratings. I will check if it gives us enough information for generating good recommendations.
#
# ###### Check the Distribution of the number of ratings per item:
#items_nb = review_data['old_item_id'].value_counts().reset_index()
items_avg = review_data.drop(['summary','reviewText','user_id','asin','user_name','reviewTime','old_user_id','item_id'],axis=1).groupby('old_item_id').agg(['count','mean']).reset_index()
items_avg.columns= ['old_item_id','num_ratings','avg_rating']
#items_avg.head(5)
items_avg['num_ratings'].describe()
# Again, the distribution of number of ratings per item is very skewed in both datasets, with 50% of the items having a small number of ratings whereas few got many ratings.
# I will check if the distribution of items rating is good enough to generate good recommendations.
#
# ###### Adding the review count and avarage to the metadata
all_info = pd.merge(all_info,items_avg,how='left',left_on='asin',right_on='old_item_id')
pd.set_option('display.max_colwidth', 100)
all_info.head(2)
# Top 10 Reviewed Products
print("Top 10 Reviewed Products:\n")
all_info[['asin','item_id','title','avg_rating','num_ratings']].sort_values('num_ratings',ascending=False).head(10)
# Check how many rows contain empty columns
a = review_data.dropna()[['item_id','user_id','score']]
#a.shape[0] / float(review_data.shape[0]) ## 1664458 /1689188.0
b = all_info.dropna()[['item_id','price']]
#b.shape[0] / float(all_info.shape[0]) ## 1664458 /1689188.0
# Get the bottom rated items that are rated more than 500 times.
all_info[all_info['num_ratings']>=500][['asin','title','avg_rating','num_ratings']].sort_values('avg_rating',ascending=True).head(5)
# Look at the top items rated more than 3000 times
all_info[all_info['num_ratings']>=3000][['asin','title','avg_rating','num_ratings']].sort_values('avg_rating',ascending=False).head(5)
# After investigating and having a good overview of the data. Let's start recommending the items
# # Explicit feedback Recommender System
# Explicit feedback is when users gives voluntarily the rating information on what they like and dislike.
#
# In this case, I have explicit item ratings ranging from one to five.
#
# `framed the recommendation system as a rating prediction machine learning problem:
# Predict an item's ratings in order to be able to recommend to a user an item that he is likely to rate high if he buys it. `
#
# ###### To evaluate the model, I randomly separate the data into a training and test set.
ratings_train, ratings_test = train_test_split( review_data, test_size=0.1, random_state=0)
ratings_train.shape
ratings_test.shape
# ### Adding Metadata to the train set
# Create an architecture that mixes the collaborative and content based filtering approaches:
# ```
# - Collaborative Part: Predict items ratings to recommend to the user items which he is likely to rate high according to learnt item & user embeddings (learn similarity from interactions).
# - Content based part: Use metadata inputs (such as price and title) about items to recommend to the user contents similar to those he rated high (learn similarity of item attributes).
# ```
#
# ### Adding the title and price
# Add the metadata of the items in the training and test datasets.
# +
# creating metadata mappings
titles = all_info['title'].unique()
titles_map = {i:val for i,val in enumerate(titles)}
inverse_titles_map = {val:i for i,val in enumerate(titles)}
price = all_info['price'].unique()
price_map = {i:val for i,val in enumerate(price)}
inverse_price_map = {val:i for i,val in enumerate(price)}
print ("We have %d prices" %price.shape)
print ("We have %d titles" %titles.shape)
all_info['price_id'] = all_info['price'].map(inverse_price_map)
all_info['title_id'] = all_info['title'].map(inverse_titles_map)
# creating dict from
item2prices = {}
for val in all_info[['item_id','price_id']].dropna().drop_duplicates().iterrows():
item2prices[val[1]["item_id"]] = val[1]["price_id"]
item2titles = {}
for val in all_info[['item_id','title_id']].dropna().drop_duplicates().iterrows():
item2titles[val[1]["item_id"]] = val[1]["title_id"]
# populating the rating dataset with item metadata info
ratings_train["price_id"] = ratings_train["item_id"].map(lambda x : item2prices[x])
ratings_train["title_id"] = ratings_train["item_id"].map(lambda x : item2titles[x])
# populating the test dataset with item metadata info
ratings_test["price_id"] = ratings_test["item_id"].map(lambda x : item2prices[x])
ratings_test["title_id"] = ratings_test["item_id"].map(lambda x : item2titles[x])
# +
# ratings_train.to_parquet("output/ratings_train.parquet")
# ratings_test.to_parquet("output/ratings_test.parquet")
# +
# # !ls -alh output
# +
# # !aws s3 cp ./output/ratings_test.parquet s3://dse-cohort5-group1/2-Keras-DeepRecommender/dataset/ratings_test.parquet
# # !aws s3 cp ./output/ratings_train.parquet s3://dse-cohort5-group1/2-Keras-DeepRecommender/dataset/ratings_train.parquet
# -
# !aws s3 cp s3://dse-cohort5-group1/2-Keras-DeepRecommender/dataset/ratings_test.parquet .
# !aws s3 cp s3://dse-cohort5-group1/2-Keras-DeepRecommender/dataset/ratings_train.parquet .
ratings_test = pd.read_parquet('ratings_test.parquet')
ratings_train = pd.read_parquet('ratings_train.parquet')
ratings_train[:3]
ratings_train.shape
# Thus we have 1,520,269 rows and 12 columns in train set
#
#
#
# # **Define embeddings
# The $\underline{embeddings}$ are low-dimensional hidden representations of users and items, i.e. for each item I can find its properties and for each user I can encode how much they like those properties so I can determine attitudes or preferences of users by a small number of hidden factors
#
# Throughout the training, I learn two new low-dimensional dense representations: one embedding for the users and another one for the items.
#
# +
# declare input embeddings to the model
#User input
user_id_input = Input(shape=[1], name='user')
#Item Input
item_id_input = Input(shape=[1], name='item')
price_id_input = Input(shape=[1], name='price')
title_id_input = Input(shape=[1], name='title')
# define the size of embeddings as a parameter
user_embedding_size = 15 # Check 5, 10 , 15, 20, 50
item_embedding_size = 15 # Check 5, 10 , 15, 20, 50
price_embedding_size = 15 # Check 5, 10 , 15, 20, 50
title_embedding_size = 15 # Check 5, 10 , 15, 20, 50
# apply an embedding layer to all inputs
user_embedding = Embedding(output_dim=user_embedding_size, input_dim=users.shape[0],
input_length=1, name='user_embedding')(user_id_input)
item_embedding = Embedding(output_dim=item_embedding_size, input_dim=items_reviewed.shape[0],
input_length=1, name='item_embedding')(item_id_input)
price_embedding = Embedding(output_dim=price_embedding_size, input_dim=price.shape[0],
input_length=1, name='price_embedding')(price_id_input)
title_embedding = Embedding(output_dim=title_embedding_size, input_dim=titles.shape[0],
input_length=1, name='title_embedding')(title_id_input)
# reshape from shape (batch_size, input_length,embedding_size) to (batch_size, embedding_size).
user_vecs = Reshape([user_embedding_size])(user_embedding)
item_vecs = Reshape([item_embedding_size])(item_embedding)
price_vecs = Reshape([price_embedding_size])(price_embedding)
title_vecs = Reshape([title_embedding_size])(title_embedding)
# -
# ### Applying matrix factorization approach
# 
# Matrix Factorisation works on the principle that we can learn the user and the item embeddings, and then predict the rating for each user-item by performing a dot (or scalar) product between the respective user and item embedding.
#
#
# Applying matrix factorization: declare the output as being the dot product between the two embeddings: items and users
y = Dot(1, normalize=False)([user_vecs, item_vecs])
#
#
#
# ## Going deeper
#
# Instead of taking a dot product of the user and the item embedding, concatenate or multiply them and use them as features for a neural network.
#
# Thus, we are not constrained to the dot product way of combining the embeddings, and can learn complex non-linear relationships.
#
# 
#
#
#
#
#
# ##### Check Performance
#
# I searched around deep network architecture and checked:
# - What happens if I add other layers on top of the first one?
# - What happens if I increase or decrease the embedding size?
# - What happens if I add dense layers on top of the embeddings before merging?
# - What happens if I change the number of hidden units in each dense layer?
# - What happens if I change the number of epochs?
# - What happens if I use Dropout or not?
#
# Modifications are commented in the code below
#
# ###### Here is the model representation for deep neural network that can be compared to the Matrix Factorisation implementation:
# Try add dense layers on top of the embeddings before merging (Comment to drop this idea.)
user_vecs = Dense(64, activation='relu')(user_vecs)
item_vecs = Dense(64, activation='relu')(item_vecs)
price_vecs = Dense(64, activation='relu')(price_vecs)
title_vecs = Dense(64, activation='relu')(title_vecs)
# +
# Concatenate the item embeddings :
item_vecs_complete = Concatenate()([item_vecs, price_vecs,title_vecs])
# Concatenate user and item embeddings and use them as features for the neural network:
input_vecs = Concatenate()([user_vecs, item_vecs_complete]) # can be changed by Multiply
#input_vecs = Concatenate()([user_vecs, item_vecs]) # can be changed by Multiply
# Multiply user and item embeddings and use them as features for the neural network:
#input_vecs = Multiply()([user_vecs, item_vecs]) # can be changed by concat
# Dropout is a technique where randomly selected neurons are ignored during training to prevent overfitting
input_vecs = Dropout(0.1)(input_vecs)
# Check one dense 128 or two dense layers (128,128) or (128,64) or three denses layers (128,64,32))
# First layer
# Dense(128) is a fully-connected layer with 128 hidden units.
# Use rectified linear units (ReLU) f(x)=max(0,x) as an activation function.
x = Dense(128, activation='relu')(input_vecs)
x = Dropout(0.1)(x) # Add droupout or not # To improve the performance
# Next Layers
#x = Dense(128, activation='relu')(x) # Add dense again or not
x = Dense(64, activation='relu')(x) # Add dense again or not
x = Dropout(0.1)(x) # Add droupout or not # To improve the performance
x = Dense(32, activation='relu')(x) # Add dense again or not #
x = Dropout(0.1)(x) # Add droupout or not # To improve the performance
# The output
y = Dense(1)(x)
# -
# ###### declare a model that takes items and users as input and output y, our prediction.
model = Model(inputs=[user_id_input
, item_id_input
, price_id_input
, title_id_input
],
outputs=y)
# ###### compile the model with 'mse' loss, and "adam" loss optimization
# I would optimise the model such that I minimise the mean squared error ('mse') on the ratings from the train set.
# The Adam optimization algorithm used, is an extension to stochastic gradient
model.compile(loss='mse',
optimizer="adam" )
# ###### Save different histories and best models using keras ModelCheckpoint callback
#
save_path = "./models"
mytime = time.strftime("%Y_%m_%d_%H_%M")
# modname = 'dense_2_15_embeddings_2_epochs' + mytime
modname = 'dense_2_15_embeddings_2_epochs'
thename = save_path + '/' + modname + '.h5'
mcheck = ModelCheckpoint(thename , monitor='val_loss', save_best_only=True)
# ###### Train the model by calling the model’s fit method
#
# When training the model, the embeddings parameters are learnt too.
#
# Use the internal keras random cross validation scheme (the validation_split=0.1 parameter below) instead of the test set to evaluate the models
#
# The test set will be kept to verify the quality of recommendations at the end.
# ## ***Set model fit
history = model.fit([ratings_train["user_id"]
, ratings_train["item_id"]
, ratings_train["price_id"]
, ratings_train["title_id"]
]
, ratings_train["score"]
, batch_size=64
, epochs=2
, validation_split=0.1
, callbacks=[mcheck]
, shuffle=True)
# ###### Save the fitted model history to a file
def plot_history(history):
# acc = history.history['accuracy']
# val_acc = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
# x = range(1, len(acc) + 1)
x = range(1, len(loss) + 1)
plt.figure(figsize=(12, 5))
# plt.subplot(1, 2, 1)
# plt.plot(x, acc, 'b', label='Training acc')
# plt.plot(x, val_acc, 'r', label='Validation acc')
# plt.title('Training and validation accuracy')
# plt.legend()
# plt.subplot(1, 2, 2)
plt.plot(x, loss, 'b', label='Training loss')
plt.plot(x, val_loss, 'r', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.savefig('loss_validation_acc.jpg')
history.history
plot_history(history)
# +
modname='dense_2_15_embeddings_2_epochs'
with open('./histories/' + modname + '.pkl' , 'wb') as file_pi:
pickle.dump(history.history, file_pi)
# -
#
# ## Compare the results of matrix factorization and different configurations of neural networks:
#
# +
def disp_model(path,file,suffix):
model = load_model(path+file+suffix)
## Summarise the model
model.summary()
# Extract the learnt user and item embeddings, i.e., a table with number of items and users rows and columns, with number of columns is the dimension of the trained embedding.
# In our case, the embeddings correspond exactly to the weights of the model:
weights = model.get_weights()
print ("embeddings \ weights shapes",[w.shape for w in weights])
return model
# model_path = "./models/save/"
model_path = "./models/"
# -
# Running the model and looking at the corresponding history I get the following
# plots of the training MSE loss for the different the embedding and epoches sizes :
# +
def plt_pickle(path,file,suffix):
with open(path+file+suffix , 'rb') as file_pi:
thepickle= pickle.load(file_pi)
plot(thepickle["loss"],label ='Train Error ' + file,linestyle="--")
plot(thepickle["val_loss"],label='Validation Error ' + file)
plt.legend()
plt.xlabel("Epoch")
plt.ylabel("Error")
##plt.ylim(0, 0.1)
return pd.DataFrame(thepickle,columns =['loss','val_loss'])
hist_path = "./histories/"
# -
model=disp_model(model_path , modname , '.h5')
# Display the model using keras
SVG(model_to_dot(model).create(prog='dot', format='svg'))
x=plt_pickle(hist_path , modname , '.pkl')
x.head(20).transpose()
# ###### Using matrix factorization with 10 embeddings and 3 epochs, the training and validation loss are very high and just go higher with each epoch, with no convergence.
model=disp_model(model_path , 'matrix_facto_10_embeddings_20_epochs' , '.h5')
# Display the model using keras
SVG(model_to_dot(model).create(prog='dot', format='svg'))
x=plt_pickle(hist_path , 'matrix_facto_10_embeddings_20_epochs' , '.pkl')
x.head(20).transpose()
# +
#disp_model(model_path , 'matrix_facto_10_embeddings_100_epochs' , '.h5')
# -
x=plt_pickle(hist_path , 'matrix_facto_10_embeddings_100_epochs' , '.pkl')
x.head(20).transpose()
# ###### Increasing number of epochs of matrix factorization to 20 or 100 does not help. Training and validation errors are still very high and just go higher with each epoch, the model does not converge
#
# ###### Check the deep network results:
#
# +
#model=disp_model(model_path , 'dense_2_Concatenate_10_embeddings_10_epochs' , '.h5')
# -
# Display the model using keras
SVG(model_to_dot(model).create(prog='dot', format='svg'))
x=plt_pickle(hist_path , 'dense_2_Concatenate_10_embeddings_10_epochs' , '.pkl')
x.head(20).transpose()
# ###### Using 2 layers (each with 128 hidden units) neural network with 10 embeddings , Concatenate user and item embeddings , with Dropout and 10 epochs- the training error stabilises around 1.6 after 9 epochs and MSE validation error stabilises around 1.4 after 7 epochs. Lowest validation error is reached also after 7 epochs
#
#
#
#
model=disp_model(model_path , 'dense_3_Multiply_10_embeddings_100_epochs' , '.h5')
# Display the model using keras
SVG(model_to_dot(model).create(prog='dot', format='svg'))
x=plt_pickle(hist_path , 'dense_3_Multiply_10_embeddings_100_epochs' , '.pkl')
x.head(30).transpose()
# ###### Using 3 layers (128,64,32) neural network with 10 embeddings, multiply user and item embeddings , with Dropout and 100 epochs, the training error stabilises around 1.2 after 7 epochs and MSE validation error stabilises around 1.4 after 7 epochs. Lowest validation error received only after 1 epoch. This is better than the above but not so good either.
# ###### The fact that training error is lower and and MSE validation error is about the same suggests overfitting
#
#
# +
#model=disp_model(model_path , 'dense_2_Concatenate_20_embeddings_25_epochs' , '.h5')
# -
# Display the model using keras
SVG(model_to_dot(model).create(prog='dot', format='svg'))
x=plt_pickle(hist_path , 'dense_2_Concatenate_20_embeddings_25_epochs' , '.pkl')
x.head(30).transpose()
# ###### Using 2 layers (128,64) neural network with 20 embeddings, Concatenate user and item embeddings , with Dropout and 25 epochs, MSE validation error stabilises again around 1.4 after 7 epochs.
# ###### No improuvment
#
#
# +
#model=disp_model(model_path , 'dense_4_Multiply_5_embeddings_7_epochs' , '.h5')
# -
# Display the model using keras
SVG(model_to_dot(model).create(prog='dot', format='svg'))
x=plt_pickle(hist_path , 'dense_4_Multiply_5_embeddings_7_epochs' , '.pkl')
x.head(30).transpose()
# ###### Using 4 layers (128,128,64,32) neural network with 5 embeddings, Multiply user and item embeddings , with no Dropout and 7 epochs, the training error is getting down and MSE validation error getting up after 2 epochs. Lowest validation error 1.4 is reached only after 2 epochs
# ###### The system is overfitting quickly
# Check only one epoch with high number of embeddings
# +
#model=disp_model(model_path , 'dense_1_Multiply_50_embeddings_7_epochs' , '.h5')
# -
# Display the model using keras
SVG(model_to_dot(model).create(prog='dot', format='svg'))
x=plt_pickle(hist_path , 'dense_1_Multiply_50_embeddings_7_epochs' , '.pkl')
x.head(30).transpose()
# ###### Using 1 layer (128) network with 50 embeddings, Multiply user and item embeddings , with no Dropout and 7 epochs, the training error is getting down and MSE validation error getting up after 2 epochs. Lowest validation error is still 1.4 and reached only after 1 epochs
# ###### The system is overfitting quickly with no improuvment
# ##### Concatenating user and item embeddings, with embedding size of 15, 2 layers and 5 epochs
#
model=disp_model(model_path , 'dense_2_Concatenate_15_embeddings_5_epochs' , '.h5')
#Display the model using keras
SVG(model_to_dot(model).create(prog='dot', format='svg'))
x=plt_pickle(hist_path , 'dense_2_Concatenate_15_embeddings_5_epochs' , '.pkl')
x.head(30).transpose()
# ###### Concatenating user and item embeddings, with embedding size of 15, 2 layers , with Dropouts and 5 epochs, the training error is getting down and MSE validation error getting up after 2 epochs. Lowest validation error of 1.29 is reached only after 2 epochs. This is a little better
#
#
#
# ###### Adding metadata with 15 embeddings, dense layers on top of the embeddings before merging , with Dropouts and 5 epochs:
#Display the model using keras
model=disp_model(model_path , 'dense_2_metadata_10-15_embeddings_5_epochs' , '.h5')
SVG(model_to_dot(model).create(prog='dot', format='svg'))
x=plt_pickle(hist_path , 'dense_2_metadata_10-15_embeddings_5_epochs' , '.pkl')
x.head(30).transpose()
# ###### Using 2 layers, adding metadata with 15 embeddings, dense layers on top of the embeddings before concatenating, with Dropout and 5 epochs, the MSE validation error stabilises around 1.28 after 2 epochs
# ###### Adding metadata with 20 embeddings, dense layers on top of the embeddings before merging , with Dropouts and 4 epochs:
#Display the model using keras
#model=disp_model(model_path , 'dense_2_metadata_20_embeddings_4_epochs' , '.h5')
SVG(model_to_dot(model).create(prog='dot', format='svg'))
x=plt_pickle(hist_path , 'dense_2_metadata_20_embeddings_4_epochs' , '.pkl')
x.head(30).transpose()
# ##### Adding metadata with 20 embeddings, dense layers on top of the embeddings before concatenating , with Dropouts and 4 epochs, the training error is getting down and MSE validation error getting up after less than 2 epochs. Lowest validation error is reached only after 2 epochs, and it is the lowest so far
#
# ###### Adding metadata with 50 embeddings, dense layers on top of the embeddings before merging , with Dropouts and 3 epochs:
#Display the model using keras
SVG(model_to_dot(model).create(prog='dot', format='svg'))
x=plt_pickle(hist_path , 'dense_3_metadata_50_embeddings_3_epochs' , '.pkl')
x.head(30).transpose()
# ###### Using 3 layers (128,64,32), adding metadata with 50 embeddings, dense layers on top of the embeddings before concatenating, with Dropout and 3 epochs, the MSE validation error stabilises around 1.195 after 2 epochs.
# ###### This is the best result I could reach
# ##### We can notice the following points from the above:
#
# - Performance got way better when using neural network comparing to using matrix factorization.
#
# - When using neural network, I converge to the best model very quickly, sometimes after 2 epochs and after that the model starts overfitting or at least the validation error does not seem to go down anymore. Matrix factorization does not converge at all.
#
# - Adding epochs lead to overfitting
#
# - Adding layers (over 3) does not help much and actually leads to overfitting
#
# - Changing the number of hidden units does not help.
#
# - Simplifying the model by reducing embedding size does not help either.
#
# - Choosing large values of embedding has made a small improvement in the results.
#
# - Multiply or concatenate user and item embeddings does not seem to matter, but concatenate seems to give little better results
#
# - Training with Dropout seem to prevent some overfitting
#
# - Adding dense layers on top of the embeddings before the merge helps a bit.
#
# - Adding some metadata lead to some improvement in the results.
#
# - Running on a larger dataset does not help either, because the data in both datasets is very skewed.
#
# ## Evaluate and compare the different models
# Using the 10% of cross validated training set records and the history I saved:
# +
hist_path = "./histories/"
validation_error = {}
train_error = {}
# models =[
# 'matrix_facto_10_embeddings_20_epochs'
# ,'matrix_facto_10_embeddings_3_epochs'
# ,'matrix_facto_10_embeddings_100_epochs'
# ,'dense_1_Concatenate_1_embeddings_1_epochs'
# ,'dense_4_Multiply_5_embeddings_7_epochs'
# ,'dense_3_Multiply_10_embeddings_100_epochs'
# ,'dense_2_Concatenate_20_embeddings_25_epochs'
# ,'dense_2_Concatenate_10_embeddings_10_epochs'
# ,'dense_1_Multiply_50_embeddings_7_epochs'
# ,'dense_2_Concatenate_10_embeddings_1_epochs'
# ,'dense_2_Concatenate_15_embeddings_5_epochs'
# ,'dense_2_metadata_10-15_embeddings_5_epochs'
# ,'dense_2_metadata_20_embeddings_4_epochs'
# ,'dense_3_metadata_50_embeddings_3_epochs'
# ]
models =[modname]
for val in models:
with open(hist_path + val +'.pkl', 'rb') as file_pi:
thepickle = pickle.load(file_pi)
validation_error[val]=np.min(thepickle["val_loss"])
train_error[val]=np.min(thepickle["loss"])
validation_error = pd.Series(validation_error)
train_error = pd.Series(train_error)
print ("MSE validation error \n",validation_error.sort_values(ascending=True).head(20))
print ("\nTrain error \n",train_error.sort_values(ascending=True).head(20))
# -
# ###### Real test results
# ## Predict - Verifying the performance on the test set.
# Check whether our results are reproducible on unseen data.
#
# Test on new data using previously saved models.
#
# I got the following results on the test set:
#
ratings_test.head(n=3)
# +
load_path = "./models/"
perfs = {}
# models =[
# 'matrix_facto_10_embeddings_20_epochs'
# ,'matrix_facto_10_embeddings_3_epochs'
# ,'dense_1_Concatenate_1_embeddings_1_epochs'
# ,'dense_4_Multiply_5_embeddings_7_epochs'
# ,'dense_3_Multiply_10_embeddings_100_epochs'
# ,'dense_2_Concatenate_20_embeddings_25_epochs'
# ,'dense_2_Concatenate_10_embeddings_10_epochs'
# ,'dense_1_Multiply_50_embeddings_7_epochs'
# ,'dense_2_Concatenate_10_embeddings_1_epochs'
# ,'dense_2_Concatenate_15_embeddings_5_epochs'
# ,'dense_2_metadata_10-15_embeddings_5_epochs'
# ]
models =[modname]
for mod in models:
model = load_model(load_path+mod+'.h5')
ratings_test['preds_' + mod] = model.predict([ratings_test['user_id'],
ratings_test['item_id'],
ratings_test["price_id"],
ratings_test["title_id"]])
perfs[mod] = mean_squared_error(ratings_test['score'], ratings_test['preds_'+mod])
perfs= pd.Series(perfs)
perfs.sort_values(ascending=True).head(20)
# +
# load_path = "./models/save/"
# #reset_keras()
# perfs = {}
# models =[
# 'dense_2_metadata_20_embeddings_4_epochs'
# , 'dense_3_metadata_50_embeddings_3_epochs'
# ]
# for mod in models:
# model = load_model(load_path+mod+'.h5')
# ratings_test['preds_' + mod] = model.predict([ratings_test["user_id"]
# , ratings_test["item_id"]
# , ratings_test["price_id"]
# , ratings_test["title_id"]
# ])
# perfs[mod] = mean_squared_error(ratings_test['score'], ratings_test['preds_'+mod]) ## MSE between real score and prdicted score
# perfs= pd.Series(perfs)
# #perfs.sort()
# perfs
# -
# ###### MSE on test data is very similar to what I got on the evaluation data
# ###### The best result on both the internal keras random cross validation scheme and test-set acheived when using 2 layers, 15 layered concatenated embeddings, Dropout and 5 epochs
# ###### I will use this model further for executing recommendations (dense_2_Concatenate_15_embeddings_5_epochs )
# # Recommend:
# The last thing to do is to use our saved models to recommend items to users:
#
#
# For the requested user:
# - Calculate the score for every item.
# - Sort the items based on the score and output the top results.
#
# ###### Check which users exist on the test set
ratings_test.user_id.value_counts().sort_values(ascending=False).head(10)
# ###### Create a recommendation example dataset of 100 users from the test set and 100 random items for each and predict recommendations for them
items = all_info.item_id.unique()
df_items = pd.DataFrame(data=items.flatten(),columns=['item_id'])
df_items = pd.merge(df_items,all_info,how='left',left_on=('item_id'),right_on=('item_id'))
df_items= df_items.sample(100)
df_items['key'] = 1
print ("We have %d unique items "%df_items['item_id'].shape[0])
# df_items= df_items[['item_id', 'description', 'category', 'title', 'title_id', 'price', 'price_id', 'brand', 'key']]
df_items= df_items[['item_id', 'description', 'category', 'title', 'title_id', 'price', 'price_id', 'key']]
df_items.head(2)
users = ratings_test.user_id.unique()
df_users = pd.DataFrame(data=users.flatten(),columns=['user_id'])
df_users = pd.merge(df_users,ratings_test,how='left',left_on=('user_id'),right_on=('user_id'))
df_users= df_users.sample(100)
df_users['key'] = 1
print ("We have %d unique users "%df_users['user_id'].shape[0])
df_users= df_users[['user_id', 'user_name', 'key']]
df_users.head(2)
# ###### Merge users and item and items metadata
df_unseenData= pd.merge(df_users, df_items, on='key')
del df_unseenData['key']
print ("We have %d unique records in the recommendation example dataset "%df_unseenData.shape[0])
df_unseenData.head(2)
df_unseenData.columns
# ###### Predict the ratings for the items and users in the a recommendation example dataset:
# +
load_path = "./models/"
# models =[
# 'matrix_facto_10_embeddings_20_epochs'
# ,'matrix_facto_10_embeddings_3_epochs'
# ,'dense_1_Concatenate_1_embeddings_1_epochs'
# ,'dense_4_Multiply_5_embeddings_7_epochs'
# ,'dense_3_Multiply_10_embeddings_100_epochs'
# ,'dense_2_Concatenate_20_embeddings_25_epochs'
# ,'dense_2_Concatenate_10_embeddings_10_epochs'
# ,'dense_1_Multiply_50_embeddings_7_epochs'
# ,'dense_2_Concatenate_10_embeddings_1_epochs'
# ,'dense_2_Concatenate_15_embeddings_5_epochs'
# ,'dense_2_metadata_10-15_embeddings_5_epochs'
# ]
models =[modname]
for mod in models:
model = load_model(load_path+mod+'.h5')
df_unseenData['preds_' + mod] = model.predict([df_unseenData['user_id'],
df_unseenData['item_id'],
df_unseenData['price_id'],
df_unseenData['title_id']])
df_unseenData.head(2)
# -
df_unseenData['user_id'].head(n=2)
df_unseenData.columns
# Check which users exist on the example set
df_unseenData.user_id.value_counts().sort_values(ascending=False).head(5)
df_unseenData[['user_id','preds_dense_2_15_embeddings_2_epochs']].sort_values('preds_dense_2_15_embeddings_2_epochs',ascending=True).head(5)
# ###### A function that will return recommendation list for a given user
df_unseenData.head(n=3)
# +
load_path = "./models/"
def get_recommendations(userID , model_scr, df_Data):
if userID not in df_Data['user_id'].values:
print("\nUser ID not found %d" %userID)
return userID
print("\nRecommendations for user id %d Name: %s is:" % (userID, df_Data.loc[df_Data['user_id'] == userID, 'user_name'].values[0]))
df_output=df_Data.loc[df_Data['user_id'] == userID][['item_id','title','description','category','price', model_scr,
]].sort_values(model_scr,ascending=False).set_index('item_id')
# print(df_output)
df_output.rename(columns={model_scr: 'score'}, inplace=True)
return df_output
# -
# ### Recommend items to a given user
####### User ID: 502656
df_output = get_recommendations(userID=502656 ,model_scr='preds_dense_2_15_embeddings_2_epochs', df_Data=df_unseenData)
df_output.head(10)
# ##### Make predictions for another user using another model:
# +
# ####### 20818 14398 79321
# df_output = get_recommendations(userID=20818 ,model_scr='dense_3_Multiply_10_embeddings_100_epochs',df_Data=df_unseenData)
# df_output.head(10)
# -
# ## Conclusion
# - In this work I created and compared 2 models for predicting user's ratings on top of Amazon's review data: a matrix factorization model and deep network model, and used the models for recommending items to users.
#
# - I showed that using deep neural networks can achieve better performance than using matrix factorization.
#
# - Going deeper (more than 3 layers) seems to lead to overfitting and not to further improvement.
#
# - Adding epochs, reducing embedding size or change hidden units numbers does not help either.
#
# - Running on a larger dataset does not help either, because the data in both datasets is very skewed.
#
# - Choosing large values of embedding (50) and adding dense layers on top of the embeddings before concatenating helps a bit.
#
# - Adding metadata and training with Dropout lead to some improvement in the results.
#
# - The fact that the data is so sparsed and skewed has a huge impact on the ability to model the recommendation problem and to achieve smaller test MSE.
# - <EMAIL>
#
| Achiveves/2-Keras-DeepRecommender/DeepRecommendation_Keras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Support Vector Machine
# ## 先玩一个 Demo
# 在使用 Python 实现之前推荐一个 demo ,这个 demo 可以让你通过控制几个参数调整分类结果。玩过之后会对 SVM 有一个直观的理解。
#
# https://cs.stanford.edu/people/karpathy/svmjs/demo/
# ## 正式开始实现 SVM
# 这里我们使用 sklearn 库中的 SVM,它可以用来做分类、回归和异常值检测,不过用得最多的还是做分类任务。这里我们也只举分类的例子。
#
# 这部分包含了三种实现 SVC,NuSVC,LinearSVC。SVC 即 Support Vector Classifier。其中 SVC 和 NuSVC 比较相似,在参数设置上略有不同。LinearSVC 顾名思义只支持线性核函数。于是我们在这里只介绍 SVC 的用法。
#
# 数据集依然是之前使用过的 iris。
# 同样地导入数据集和所需的函数。
from sklearn import svm, datasets
import numpy as np
import matplotlib.pyplot as plt
from sklearn.metrics import confusion_matrix
# 这里我们要可视化地展示分类结果,所以构造两个函数 make_meshgrid 和 plot_contours
# +
def make_meshgrid(x, y, h=.02):
"""Create a mesh of points to plot in
Parameters
----------
x: data to base x-axis meshgrid on
y: data to base y-axis meshgrid on
h: stepsize for meshgrid, optional
Returns
-------
xx, yy : ndarray
"""
x_min, x_max = x.min() - 1, x.max() + 1
y_min, y_max = y.min() - 1, y.max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
return xx, yy
def plot_contours(ax, clf, xx, yy, **params):
"""Plot the decision boundaries for a classifier.
Parameters
----------
ax: matplotlib axes object
clf: a classifier
xx: meshgrid ndarray
yy: meshgrid ndarray
params: dictionary of params to pass to contourf, optional
"""
Z = clf.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
out = ax.contourf(xx, yy, Z, **params)
return out
# -
# 导入数据。因为我们要做可视化,所以只取两个特征作为分类依据。
iris = datasets.load_iris()
X = iris.data[:,:2]
y = iris.target
## 不做额外设置的话惩罚系数 C 的默认值为 1.0
## 这里可以通过设置参数 decision_function_shape 更改多分类类型
clf_linear = svm.SVC(kernel='linear')
clf_gaussian = svm.SVC(kernel='rbf') ## gamma 默认等于0.7,可调
clf_poly = svm.SVC(kernel='poly') ## poly 默认等于3,可调
model_linear = clf_linear.fit(X, y)
model_gaussion = clf_gaussian.fit(X, y)
model_poly = clf_poly.fit(X, y)
# +
## 可视化专用部分,不必改动
## 目的是生成一堆点(xx,yy)形成网格将现有的数据都覆盖进去,用于后续的预测
X0, X1 = X[:, 0], X[:, 1]
xx, yy = make_meshgrid(X0, X1)
print xx.shape, yy.shape
# -
## 线性核
Z = model_linear.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
ax = plt.gca()
ax.contourf(xx, yy, Z)
ax.scatter(X0, X1, c=y, s=20, edgecolors='k')
ax.set_xlim(xx.min(), xx.max())
ax.set_ylim(yy.min(), yy.max())
ax.set_xlabel('Sepal length')
ax.set_ylabel('Sepal width')
ax.set_xticks(())
ax.set_yticks(())
ax.set_title('SVC with linear kernel')
## 高斯核和多项式核
fig, sub = plt.subplots(1, 2)
titles = ('SVC with RBF kernel', 'SVC with polynomial kernel')
models = (model_gaussion, model_poly)
for clf, title, ax in zip(models, titles, sub.flatten()):
plot_contours(ax, clf, xx, yy)
ax.scatter(X0, X1, c=y, s=50, edgecolors='k')
ax.set_xlim(xx.min(), xx.max())
ax.set_ylim(yy.min(), yy.max())
ax.set_xlabel('Sepal length')
ax.set_ylabel('Sepal width')
ax.set_xticks(())
ax.set_yticks(())
ax.set_title(title)
plt.tight_layout()
plt.show()
# 可以自己调整各种参数感受 decision surface 的区别。
# ## 用于预测
# 以高斯核的 SVM 为例说明其他的一些功能。
#
# 训练好的 SVM 依然可以通过 predict 函数来预测结果。
model_gaussion.predict(X[:10,:])
# 这里我们用 confusion matrix 混淆矩阵开查看分类准确程度
y_pred = model_gaussion.predict(X)
cnf_matrix = confusion_matrix(y_pred, y)
print cnf_matrix
# 在这个矩阵中行表示真实类别1,2,3;列表示预测的类别1,2,3。对角线上的数字表示预测正确的样本个数,14表示有14个本来是第二类的样本被认为是第三类了。以此类推。
## 通过调用 score 查看预测的 mean accuracy
model_gaussion.score(X, y)
## 通过调用 decision_function 查看样本到各个超平面的距离
## 默认采用ovr的方式进行多分类
model_gaussion.decision_function(X[:10,:])
# ## 支撑向量
# 也可以查看支撑向量 support vectors 的相关信息
## 获得支撑向量
model_gaussion.support_vectors_
## 获得支撑向量的 indices
model_gaussion.support_
## 获得每个 class 的支撑向量个数
model_gaussion.n_support_
# 请比较 SVM 与逻辑回归、决策树在同一个数据集 iris 上做分类时 decision boundary 的差异
# 请用上这个数据集的全部特征,并且调整各个参数,看分类效果是否有改善
| Support Vector Machine - Examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + active=""
# Text provided under a Creative Commons Attribution license, CC-BY. All code is made available under the FSF-approved BSD-3 license. (c) <NAME>, <NAME> 2017. Thanks to NSF for support via CAREER award #1149784.
# -
# [@LorenaABarba](https://twitter.com/LorenaABarba)
# This notebook complements the [interactive CFD online](https://bitbucket.org/cfdpython/cfd-python-class/overview) module **12 steps to Navier-Stokes**, addressing the issue of high performance with Python.
# Optimizing Loops with Numba
# ----
# ***
# You will recall from our exploration of [array operations with NumPy](./06_Array_Operations_with_NumPy.ipynb) that there are large speed gains to be had from implementing our discretizations using NumPy-optimized array operations instead of many nested loops.
#
# [Numba](http://numba.pydata.org/) is a tool that offers another approach to optimizing our Python code. Numba is a library for Python which turns Python functions into C-style compiled functions using LLVM. Depending on the original code and the size of the problem, Numba can provide a significant speedup over NumPy optimized code.
#
# Let's revisit the 2D Laplace Equation:
# +
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
from matplotlib import pyplot
import numpy
##variable declarations
nx = 81
ny = 81
c = 1
dx = 2.0/(nx-1)
dy = 2.0/(ny-1)
##initial conditions
p = numpy.zeros((ny,nx)) ##create a XxY vector of 0's
##plotting aids
x = numpy.linspace(0,2,nx)
y = numpy.linspace(0,1,ny)
##boundary conditions
p[:,0] = 0 ##p = 0 @ x = 0
p[:,-1] = y ##p = y @ x = 2
p[0,:] = p[1,:] ##dp/dy = 0 @ y = 0
p[-1,:] = p[-2,:] ##dp/dy = 0 @ y = 1
# -
# Here is the function for iterating over the Laplace Equation that we wrote in Step 9:
def laplace2d(p, y, dx, dy, l1norm_target):
l1norm = 1
pn = numpy.empty_like(p)
while l1norm > l1norm_target:
pn = p.copy()
p[1:-1,1:-1] = (dy**2*(pn[2:,1:-1]+pn[0:-2,1:-1])+dx**2*(pn[1:-1,2:]+pn[1:-1,0:-2]))/(2*(dx**2+dy**2))
p[0,0] = (dy**2*(pn[1,0]+pn[-1,0])+dx**2*(pn[0,1]+pn[0,-1]))/(2*(dx**2+dy**2))
p[-1,-1] = (dy**2*(pn[0,-1]+pn[-2,-1])+dx**2*(pn[-1,0]+pn[-1,-2]))/(2*(dx**2+dy**2))
p[:,0] = 0 ##p = 0 @ x = 0
p[:,-1] = y ##p = y @ x = 2
p[0,:] = p[1,:] ##dp/dy = 0 @ y = 0
p[-1,:] = p[-2,:] ##dp/dy = 0 @ y = 1
l1norm = (numpy.sum(np.abs(p[:])-np.abs(pn[:])))/np.sum(np.abs(pn[:]))
return p
# Let's use the `%%timeit` cell-magic to see how fast it runs:
# %%timeit
laplace2d(p, y, dx, dy, .00001)
# Ok! Our function `laplace2d` takes around 206 *micro*-seconds to complete. That's pretty fast and we have our array operations to thank for that. Let's take a look at how long it takes using a more 'vanilla' Python version.
def laplace2d_vanilla(p, y, dx, dy, l1norm_target):
l1norm = 1
pn = numpy.empty_like(p)
nx, ny = len(y), len(y)
while l1norm > l1norm_target:
pn = p.copy()
for i in range(1, nx-1):
for j in range(1, ny-1):
p[i,j] = (dy**2*(pn[i+1,j]+pn[i-1,j])+dx**2*(pn[i,j+1]-pn[i,j-1]))/(2*(dx**2+dy**2))
p[0,0] = (dy**2*(pn[1,0]+pn[-1,0])+dx**2*(pn[0,1]+pn[0,-1]))/(2*(dx**2+dy**2))
p[-1,-1] = (dy**2*(pn[0,-1]+pn[-2,-1])+dx**2*(pn[-1,0]+pn[-1,-2]))/(2*(dx**2+dy**2))
p[:,0] = 0 ##p = 0 @ x = 0
p[:,-1] = y ##p = y @ x = 2
p[0,:] = p[1,:] ##dp/dy = 0 @ y = 0
p[-1,:] = p[-2,:] ##dp/dy = 0 @ y = 1
l1norm = (numpy.sum(np.abs(p[:])-np.abs(pn[:])))/np.sum(np.abs(pn[:]))
return p
# %%timeit
laplace2d_vanilla(p, y, dx, dy, .00001)
# The simple Python version takes 32 *milli*-seconds to complete. Let's calculate the speedup we gained in using array operations:
32*1e-3/(206*1e-6)
# So NumPy gives us a 155x speed increase over regular Python code. That said, sometimes implementing our discretizations in array operations can be a little bit tricky.
#
# Let's see what Numba can do. We'll start by importing the special function decorator `autojit` from the `numba` library:
from numba import autojit
# To integrate Numba with our existing function, all we have to do it is prepend the `@autojit` function decorator before our `def` statement:
@autojit
def laplace2d_numba(p, y, dx, dy, l1norm_target):
l1norm = 1
pn = numpy.empty_like(p)
while l1norm > l1norm_target:
pn = p.copy()
p[1:-1,1:-1] = (dy**2*(pn[2:,1:-1]+pn[0:-2,1:-1])+dx**2*(pn[1:-1,2:]+pn[1:-1,0:-2]))/(2*(dx**2+dy**2))
p[0,0] = (dy**2*(pn[1,0]+pn[-1,0])+dx**2*(pn[0,1]+pn[0,-1]))/(2*(dx**2+dy**2))
p[-1,-1] = (dy**2*(pn[0,-1]+pn[-2,-1])+dx**2*(pn[-1,0]+pn[-1,-2]))/(2*(dx**2+dy**2))
p[:,0] = 0 ##p = 0 @ x = 0
p[:,-1] = y ##p = y @ x = 2
p[0,:] = p[1,:] ##dp/dy = 0 @ y = 0
p[-1,:] = p[-2,:] ##dp/dy = 0 @ y = 1
l1norm = (numpy.sum(np.abs(p[:])-np.abs(pn[:])))/np.sum(np.abs(pn[:]))
return p
# The only lines that have changed are the `@autojit` line and also the function name, which has been changed so we can compare performance. Now let's see what happens:
# %%timeit
laplace2d_numba(p, y, dx, dy, .00001)
# Ok! So it's not a 155x speed increase like we saw between vanilla Python and NumPy, but it is a non-trivial gain in performance time, especially given how easy it was to implement. Another cool feature of Numba is that you can use the `@autojit` decorator on non-array operation functions, too. Let's try adding it onto our vanilla version:
@autojit
def laplace2d_vanilla_numba(p, y, dx, dy, l1norm_target):
l1norm = 1
pn = numpy.empty_like(p)
nx, ny = len(y), len(y)
while l1norm > l1norm_target:
pn = p.copy()
for i in range(1, nx-1):
for j in range(1, ny-1):
p[i,j] = (dy**2*(pn[i+1,j]+pn[i-1,j])+dx**2*(pn[i,j+1]-pn[i,j-1]))/(2*(dx**2+dy**2))
p[0,0] = (dy**2*(pn[1,0]+pn[-1,0])+dx**2*(pn[0,1]+pn[0,-1]))/(2*(dx**2+dy**2))
p[-1,-1] = (dy**2*(pn[0,-1]+pn[-2,-1])+dx**2*(pn[-1,0]+pn[-1,-2]))/(2*(dx**2+dy**2))
p[:,0] = 0 ##p = 0 @ x = 0
p[:,-1] = y ##p = y @ x = 2
p[0,:] = p[1,:] ##dp/dy = 0 @ y = 0
p[-1,:] = p[-2,:] ##dp/dy = 0 @ y = 1
l1norm = (numpy.sum(np.abs(p[:])-np.abs(pn[:])))/np.sum(np.abs(pn[:]))
return p
# %%timeit
laplace2d_vanilla_numba(p, y, dx, dy, .00001)
# 561 micro-seconds. That's not quite the 155x increase we saw with NumPy, but it's close. And all we did was add one line of code.
#
# So we have:
#
# Vanilla Python: 32 milliseconds
#
# NumPy Python: 206 microseconds
#
# Vanilla + Numba: 561 microseconds
#
# NumPy + Numba: 137 microseconds
#
# Clearly the NumPy + Numba combination is the fastest, but the ability to quickly optimize code with nested loops can also come in very handy in certain applications.
#
#
from IPython.core.display import HTML
def css_styling():
styles = open("../styles/custom.css", "r").read()
return HTML(styles)
css_styling()
# > (The cell above executes the style for this notebook. We modified a style we found on the GitHub of [CamDavidsonPilon](https://github.com/CamDavidsonPilon), [@Cmrn_DP](https://twitter.com/cmrn_dp).)
| lessons/14_Optimizing_Loops_with_Numba.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PyCharm (Face2Text)
# language: python
# name: pycharm-548e7b6f
# ---
# # Notebook used to create a validation set out of the training set
import os, random, math
# +
cwd = os.getcwd()
datasetName = 'Segmentation_Dataset'
validationDir = os.path.join(cwd, datasetName, 'validation')
trainingDir = os.path.join(cwd, datasetName, 'training')
nTrainImg = len(os.listdir(os.path.join(trainingDir, 'images', 'img')))
valRatio = 11
if not os.path.exists(validationDir):
os.makedirs(validationDir)
if not os.path.exists(os.path.join(validationDir, 'images', 'img')):
os.makedirs(os.path.join(validationDir, 'images', 'img'))
if not os.path.exists(os.path.join(validationDir, 'masks', 'img')):
os.makedirs(os.path.join(validationDir, 'masks', 'img'))
random.seed(1234)
if valRatio > 1:
raise Exception('The validation ratio must be <= 1.')
for i in range(math.floor(valRatio * nTrainImg)):
chosenFile = random.choice(os.listdir(os.path.join(trainingDir, 'images', 'img')))
os.rename(os.path.join(trainingDir, 'images', 'img', chosenFile), os.path.join(validationDir, 'images', 'img', chosenFile))
os.rename(os.path.join(trainingDir, 'masks', 'img', chosenFile), os.path.join(validationDir, 'masks', 'img', chosenFile))
| Competition 2/Notebooks/ValSetCreation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # How to implement Graphs:
# ## 1) Edge Lists:
# In this we maintain two lists one for storing vertices and the other for storing edges.
# There's some problem in this, if I ask you whether there's an edge between any two vertices V1 and V2,
# you need to traverse the complete Edge array taking O(E) time. {O(E) is O(N**2}}. It is taking lots of
# time for simple computation
# 
#
# 
# ## 2) Adjacency List:
# Each vertex will maintain all the vertices which is adjacent to. Searching a vertex in a list take O(N) time but this can be reduced if we use dictionary and search if a vertex (say 3) is there are not.
#
#
# 
# ## 3) Adjacency Matrix: We maintain a matrix and store 1 if there's an edge else 0
#
# If adj[v1][v2] is 0 then there's no edge in between v1 and v2
#
# Space: O(N**2)
#
#
#
# 
# iF EDGES ARE LESS THEN ADJACENCY LIST IS BETTER
# IF EDGES ARE MORE THEN ADJACENCY MATRIX IS BETTER. **SO USE ACCORDINGLY
| 19 Graphs - 1/19.04 How to implement Graphs.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.7.0
# language: julia
# name: julia-1.7
# ---
# # Week 1
#
# This problem sheet tests the representation of numbers on the computer, using
# modular arithmetic. We also use floating point rounding modes to implement
# interval arithmetic, and thereby
# produce rigorous bounds on the exponential.
using ColorBitstring, SetRounding
# Questions marked with a ⋆ are meant to be completed without using a computer.
#
# ## 1. Binary representation
#
#
# **Problem 1.1** What is the binary representations of $1/5$? (Hint: use `printbits` to derive a proposed form.)
#
#
#
# **Problem 1.2⋆** What is $\pi$ to 5 binary places? Hint: recall that $\pi \approx 3.14$.
#
#
# ## 2. Integers
#
# **Problem 2.1⋆** With 8-bit signed integers, find the bits for the following: $10, 120, -10$.
#
#
# **Problem 2.2⋆** What will `Int8(120) + Int8(10)` return?
#
#
# ## 3. Floating point numbers
#
# **Problem 3.1⋆** What are the single precision $F_{32}$ (`Float32`) floating point representations for the following:
# $$
# 2, 31, 32, 23/4, (23/4)\times 2^{100}
# $$
# Check your answers using `printbits`.
#
#
# **Problem 3.2** Let $m(y) = \min\{x \in F_{32} : x > y \}$ be the smallest single precision number
# greater than $y$. What is $m(2) - 2$ and $m(1024) - 1024$? Check your answer using the `nextfloat`
# command.
#
#
#
# ## 4. Arithmetic
#
#
# **Problem 4.1⋆** Suppose $x = 1.25$ and consider 16-bit floating point arithmetic (`Float16`).
# What is the error in approximating $x$ by the nearest float point number ${\rm fl}(x)$?
# What is the error in approximating $2x$, $x/2$, $x + 2$ and $x - 2$ by $2 \otimes x$, $x \oslash 2$, $x ⊕ 2$ and $x \ominus 2$?
#
# **Problem 4.2⋆** For what floating point numbers is $x \oslash 2 \neq x/2$ and $x ⊕ 2 \neq x + 2$?
#
#
#
#
# **Problem 4.3⋆** Explain why the following return `true`. What is the largest floating point number `y` such that `y + 1 ≠ y`?
x = 10.0^100
x + 1 == x
# **Problem 4.4⋆** What are the exact bits for $1/5$, $1/5 + 1$ computed
# using half-precision arithmetic (`Float16`) (using default rounding)?
#
#
#
# **Problem 4.5⋆** Explain why the following does not return `1`. Can you compute the bits explicitly?
Float16(0.1) / (Float16(1.1) - 1)
# **Problem 4.4⋆** Find a bound on the _absolute error_ in terms of a constant times
# $ϵ_{\rm m}$ for the following computations
# $$
# \begin{align*}
# (1.1 * 1.2) &+ 1.3 \\
# (1.1 - 1) &/ 0.1
# \end{align*}
# $$
# implemented using floating point arithmetic (with any precision).
#
#
#
#
# ## 5. Interval arithmetic
#
#
# The following problems consider implementation of interval arithmetic for
# proving precise bounds on arithmetic operations. That is recall the set operations
# $$
# A + B = \{x + y : x \in A, y \in B\}, AB = \{xy : x \in A, y \in B\}.
# $$
#
# **Problem 5.1⋆** For intervals $A = [a,b]$ and $B = [c,d]$ such that $0 \notin A,B$
# and integer $n \neq 0$,
# deduce formulas for the minimum and maximum of $A/n$, $A+B$ and $AB$.
#
#
#
#
#
#
# **Problem 5.2**
# We want to implement floating point variants such that, for $S = [a,b] + [c,d]$
# $P = [a,b] * [c,d]$, and $D = [a,b]/n$ for an integer $n$,
# $$
# \begin{align*}
# [a,b] ⊕ [c,d] &:= [{\rm fl}^{\rm down}(\min S), {\rm fl}^{\rm up}(\max S)] \\
# [a,b] ⊗ [c,d] &:= [{\rm fl}^{\rm down}(\min P), {\rm fl}^{\rm up}(\max P)] \\
# [a,b] ⊘ n &:= [{\rm fl}^{\rm down}(\min D), {\rm fl}^{\rm up}(\max D)]
# \end{align*}
# $$
# This guarantees $S ⊆ [a,b] ⊕ [c,d]$, $P ⊆ [a,b] ⊗ [c,d]$, and
# $D ⊆ [a,b] ⊘ n$.
# In other words, if $x \in [a,b]$ and
# $y \in [c,d]$ then $x +y \in [a,b] ⊕ [c,d]$, and we thereby have bounds on $x + y$.
#
# Use the formulae from Problem 5.1 to complete (by replacing the `# TODO: …` comments with code)
# the following implementation of an
# `Interval`
# so that `+`, `-`, and `/` implement $⊕$, $⊖$, and $⊘$ as defined above.
# +
# Interval(a,b) represents the closed interval [a,b]
struct Interval{T}
a::T
b::T
end
import Base: *, +, -, /, one, in
# create an interval corresponding to [1,1]
one(x::Interval) = Interval(one(x.a), one(x.b))
# Support x in Interval(a,b)
in(x, y::Interval) = y.a ≤ x ≤ y.b
# Following should implement ⊕
function +(x::Interval, y::Interval)
T = promote_type(typeof(x.a), typeof(x.b))
a = setrounding(T, RoundDown) do
# TODO: upper bound
end
b = setrounding(T, RoundUp) do
# TODO: upper bound
end
Interval(a, b)
end
# Following should implement ⊘
function /(x::Interval, n::Integer)
T = typeof(x.a)
if iszero(n)
error("Dividing by zero not support")
end
a = setrounding(T, RoundDown) do
# TODO: lower bound
end
b = setrounding(T, RoundUp) do
# TODO: upper bound
end
Interval(a, b)
end
# Following should implement ⊗
function *(x::Interval, y::Interval)
T = promote_type(typeof(x.a), typeof(x.b))
if 0 in x || 0 in y
error("Multiplying with intervals containing 0 not supported.")
end
a = setrounding(T, RoundDown) do
# TODO: lower bound
end
b = setrounding(T, RoundUp) do
# TODO: upper bound
end
Interval(a, b)
end
# -
# **Problem 5.3** The following function computes the first `n+1` terms of the Taylor series of $\exp(x)$:
# $$
# \sum_{k=0}^n {x^k \over k!}
# $$
function exp_t(x, n)
ret = one(x) # 1 of same type as x
s = one(x)
for k = 1:n
s = s/k * x
ret = ret + s
end
ret
end
# Bound the tail of the Taylor series for ${\rm e}^x$ assuming $|x| \leq 1$.
# (Hint: ${\rm e}^x \leq 3$ for $x \leq 1$.)
# Use the bound
# to complete the function `exp_bound` which computes ${\rm e}^x$ with rigorous error bounds, that is
# so that when applied to an interval $[a,b]$ it returns an interval that is
# guaranteed to contain the interval $[{\rm e}^a, {\rm e}^b]$.
function exp_bound(x::Interval, n)
if abs(x.a) > 1 || abs(x.b) > 1
error("Interval must be a subset of [-1, 1]")
end
ret = exp_t(x, n) # the code for Taylor series should work on Interval unmodified
f = factorial(min(20, n + 1)) # avoid overflow in computing factorial
T = typeof(ret.a)
# TODO: modify ret so that exp(x) is guaranteed to lie in it
end
# Check your result by assuring that
# the following returns `true`:
exp(big(1)) in exp_bound(Interval(1.0,1.0), 20) && exp(big(-1)) in exp_bound(Interval(-1.0,-1.0), 20)
| sheets/week1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:pablocarreira-py39] *
# language: python
# name: conda-env-pablocarreira-py39-py
# ---
# <br>
#
# # Introdução
import os
import gspread
import pandas as pd
from datetime import datetime
from oauth2client.service_account import ServiceAccountCredentials
gspread.__version__
# <br>
#
# # Autenticação
#
# Códigos necessários para "entrar" em modo de edição de uma planilha, a partir do _python_.
# +
# Escopo Utilizado
scope = [
'https://spreadsheets.google.com/feeds',
'https://www.googleapis.com/auth/drive',
]
# API
api_file = 'python-gspread-personal.json'
# Lê a Credencial para Autenticação
credentials = ServiceAccountCredentials.from_json_keyfile_name(
os.path.join('..', '..', '..', 'my_vault', api_file),
scope)
# Autenticação, de fato
gc = gspread.authorize(credentials)
# -
# <br>
#
# Escolhe a planilha e aba a ser editada<br>
# https://docs.google.com/spreadsheets/d/1bRwjoieInaRkmoyvlisMtDi-2kZ-5CvyKEQ2V_Xk1U8/edit#gid=0
# Ou Autenticar assim
gc = gspread.service_account(filename=os.path.join('..', '..', '..', 'my_vault', api_file))
# Autorizar
gc = gspread.authorize(credentials)
gc.list_permissions('1bRwjoieInaRkmoyvlisMtDi-2kZ-5CvyKEQ2V_Xk1U8')
# Abre a Planilha
url = 'https://docs.google.com/spreadsheets/d/1bRwjoieInaRkmoyvlisMtDi-2kZ-5CvyKEQ2V_Xk1U8/edit#gid=0'
sh = gc.open_by_key('<KEY>')
sh = gc.open_by_url('https://docs.google.com/spreadsheets/d/1bRwjoieInaRkmoyvlisMtDi-2kZ-5CvyKEQ2V_Xk1U8/edit#gid=0')
sh = gc.open('python gspread').sheet1
sh = gc.open('python gspread')
# Lista Abas
sh.worksheets()
# Adiciona Aba
worksheet = sh.add_worksheet(title='worksheet', rows='100', cols='20')
# Deleta Aba
sh.del_worksheet(worksheet)
# Seleciona Aba
worksheet = sh.get_worksheet(0) # Pela ordem, começando com 0
worksheet = sh.worksheet('Página1') # Pelo nome da aba
# Get URL
worksheet.url
# Nome da Aba
worksheet.title
# Limpa
worksheet.clear()
# +
#sh = gc.create('spreadsheet')
#sh.share('<EMAIL>', perm_type='user', role='writer')
# -
# <br>
#
# # Escrevendo Dados
# ## Célula Individual
# +
# Limpa
worksheet.clear()
# Pela notação de coordenadas
worksheet.update_cell(1, 2, 'Bingo!')
# +
# Limpa
worksheet.clear()
# Pela notação padrão do Google Spreadshhet
worksheet.update_acell('A1', 'Dia e Hora')
# +
# Limpa
worksheet.clear()
# Valores de Função
#worksheet.update_acell('A2', datetime.today().strftime('%Y-%m-%d'))
worksheet.update_acell('A2', datetime.today().strftime('%d/%m/%Y'))
# +
# Autorizar
gc = gspread.authorize(credentials)
# Limpa
worksheet.clear()
# Update
sh.values_update(
'Página1!A1',
params={
'valueInputOption': 'USER_ENTERED'
},
body={
'values': [[1, 2.3, 3]]
}
)
# -
# <br>
#
# ## Range
# +
# Limpa
worksheet.clear()
# Atualiza um Conjunto de Células iguais
cell_list = worksheet.range('A1:D7')
# Loop
for cell in cell_list:
cell.value = '00'
# Update in batch
worksheet.update_cells(cell_list)
# -
# <br>
#
# ## Append
# +
# Limpa
#worksheet.clear()
worksheet.append_row(
['Test1', 2.4, 'Test2'],
table_range='A1'
)
# -
# <br>
#
# ## *Dictionary*
# <br>
#
# ### *Dictionary* to *GoogleSheets* 1
# +
# Limpa
worksheet.clear()
# Cabeçalho
worksheet.update_acell('A1', 'Estado')
worksheet.update_acell('B1', 'Capital')
# Dicionário com os estados e as capitais
capitais = {
'Paraíba': 'João Pessoa',
'Santa Catarina': 'Florianópolis',
'São Paulo': 'São Paulo'
}
# Contador de colunas e celulas
row = 2
column = 1
# Loop
for key, value in capitais.items():
# Atualiza a celula 2 da coluna 1 com o nome do estado
worksheet.update_cell(row, column, key)
# A coluna agora é a B
column = 2
# Atualiza a celula 2 da coluna 2 com o nome da capital
worksheet.update_cell(row, column, value)
# A coluna agora é a A
column = 1
# Acrescenta mais um valor no numero da celula
row += 1
# -
# <br>
#
# ### *Dictionary* to *GoogleSheets* 2
# +
# Limpa
worksheet.clear()
# Dataframe
df = pd.DataFrame.from_dict(
{
'col_1': [3, 2, 1.5, 0.214],
'col_2': ['a', 'banana', 'c', 'd']
}
)
# Update
worksheet.update([df.columns.values.tolist()] + df.values.tolist())
# -
# <br>
#
# ## *GoogleSheets* to *Dictionary*
# Get Values
worksheet.get_all_records()
pd.DataFrame(worksheet.get_all_records())
# <br>
#
# # Lendo Dados
# ## Célula Individual
# +
# Autorizar
gc = gspread.authorize(credentials)
# Pega o valor específico de uma célula
worksheet.cell(1,1).value
# -
worksheet.acell('A3').value
# Seleciona todos os dados de uma coluna
worksheet.col_values(1)
# Seleciona todos os dados de uma linha
worksheet.row_values(1)
# <br>
#
# # Procura
# Procurando uma localização específica
cell = worksheet.find('banana')
'Encontrado na celula {} coluna {}'.format(cell.row, cell.col)
# <br>
#
# ## Conjunto de Células
# Gera uma lista de listas
list_of_lists = worksheet.get_all_values()
list_of_lists[0:3]
# +
# Autorizar
gc = gspread.authorize(credentials)
# Get All Values
data = worksheet.get_all_values()
headers = data.pop(0)
# Data
df = pd.DataFrame(
data,
columns=headers
)
# Results
df.head()
# +
# Autorizar
gc = gspread.authorize(credentials)
# Get Data
data = worksheet.get_all_records(
#empty2zero=True,
#head=1,
#default_blank='',
#allow_underscores_in_numeric_literals=False,
)
# Results
data
# -
# <br>
#
# # Package *df2gspread*
from df2gspread import gspread2df as g2d
from df2gspread import df2gspread as d2g
# <br>
#
# ## *Google Sheet* para *Dataframe*
# +
df = g2d.download(
gfile='1bRwjoieInaRkmoyvlisMtDi-2kZ-5CvyKEQ2V_Xk1U8',
wks_name='Página1',
start_cell='A1',
credentials=credentials,
col_names=True,
row_names=False
)
# Results
df.head(2)
# -
# <br>
#
# ## *Dataframe* para *Google Sheet*
# +
# Dataframe
df = pd.DataFrame.from_dict(
{
'col_1': [3, 2, 1.5, 0.214],
'col_2': ['a', 'b', 'c', 'd']
}
)
# Send to Google Spreadsheet
d2g.upload(
df,
gfile='1bRwjoieInaRkmoyvlisMtDi-2kZ-5CvyKEQ2V_Xk1U8',
wks_name='Página1',
start_cell='A1',
credentials=credentials,
col_names=True,
row_names=False,
clean=True,
)
# Results
df.head(2)
# +
# Lendo e filtrando dados
df = pd.read_csv(os.path.join('data', 'empresas.xz'))
df = df[df['state'] == 'SP']
df = df[df['city'] == 'SANTOS']
# Results
df.head(2)
# -
# Convert df to list
for i in df.values.tolist():
print(i)
| test/gspread.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.9 (''.venv'': venv)'
# language: python
# name: python3
# ---
# +
import tensorflow as tf
import tensorflow_hub as hub
print("TF Version: ", tf.__version__)
print("Hub Version: ", hub.__version__)
print("GPU is", "available" if tf.config.list_physical_devices('GPU') else "not available")
# -
do_finetuning = False
| model exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Coding with Clarkey: Lesson 1!
#
# This aims to build a little dictionaries which we covered in person.
#
# * There is a library that holds a collection of books (mostly just the ones that
# I could remember an author for :p)
#
# * There is a simple loop set up that will go through the dictionary, getting both
# the title (the 'key'), and the info about the book (the 'value')
# * Currently, this loop just prints out very unsexy information
# +
library = {
'A Tale of Two Cities': {
'author': '<NAME>',
'year': 1859
},
'The Thorn Birds': {
'author': '<NAME>',
'year': 1977
},
'The Handmaid\'s Tale': {
'author': '<NAME>',
'year': 1985
},
'The Book Thief': {
'author': '<NAME>',
'year': 2005
}
}
def display():
for title, info in library.items():
print(title, info)
# now, let's call our display() method to see what it looks like
display()
# -
| beginner/lesson001/lesson001_collections.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Введение в оспользование [antspy](https://github.com/ANTsX/ANTsPy) и MRTDataset
import ants
# Скармливаете `ants.image_read()` путь до `.nii.gz` или `.mni`
img = ants.image_read(ants.get_ants_data('mni'))
img.plot()
# Это все легко перегоняется в numpy массивы
img_array = img.numpy()
img_array.shape, img_array.max()
# Это все дело можно нормировать
ants.iMath_normalize(img).max()
# Также удобно ресайзить
print(img.shape)
img.resample_image((10, 20, 30), 1, 0).shape
# Также можно поворачивать
ants.reorient_image(img, (1,0))
# Ну и гонять массивы назад в мозги
# +
new_img1 = img.new_image_like(img_array)
# doesnt copy any information
new_img2 = ants.from_numpy(img_array)
# verbose way to copy information
new_img3 = ants.from_numpy(img_array, spacing=img.spacing,
origin=img.origin, direction=img.direction)
# -
# ## Теперь перейдем к датасету
# +
# %matplotlib notebook
import os, sys
from matplotlib import pyplot
sys.path.append('/root/cAAE')
from model.tools.config import Config, read_conf
from model.generator import generator, net
config = read_conf(f'/root/cAAE/config/{os.environ["CONFIG_NAME"]}.json')
# -
def plot_all_axis(brain):
n=2/3
fig, axs = pyplot.subplots(1, 3)
axs[0].imshow(brain[int(brain.shape[0]*n), :, :])
axs[0].set_title('Brain axis = 0')
axs[1].imshow(brain[:, int(brain.shape[1]*n), :])
axs[1].set_title('Brain axis = 1')
axs[2].imshow(brain[:, :, int(brain.shape[1]*n)])
axs[2].set_title('Brain axis = 2')
# Давайте посмотрим на датасеты. Сначала на тренировочный (`train_flg = True`)
dataset = generator(config, train_flg=True)
# Основа датасета это объект класса `Person`. Датасет веде себя и как словарь и как список
person = dataset.get_person(11) #Или случайный dataset.get_random()
person
# Если вызывать `dataset[i]`, то будет вызвано `person.__call__` с `transform`, которая есть в датасете
print(dataset.transform)
brain = dataset[11]
brain.shape
# Давайте посмотрим на преобразованный тензор. Желательно следить за направлением мозга по каждой из осей, чтобы на трейне и на тесте данные были соноправленны.
plot_all_axis(brain)
# А теперь посмотрим на то, как оно выглядело в оригинале
person.plot(axis=2)
# Для тестовых данных работает все тоже самое, но появляется еще и возможность посмотреть на опухоли
dataset_test = generator(config, train_flg=False)
person_test = dataset_test.get_person(11)
print(dataset_test.transform) #должен быть такой же, как и трейн
brain_test = dataset_test[11]
brain_test.shape
# Должны смотреть в том же направлении, что и трейн
plot_all_axis(brain_test)
# Так получается маска, она вам понадобится
person_test.get_mask().plot()
# Для того, чтобы получить опухоль, надо сначала ее подвязать
person_test.set_tumor()
tumor = person_test.get_tumor(dataset_test.transform)
plot_all_axis(tumor)
| jupyter/MRTDataset_and_antspy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Dataset and Connection to SAP HANA
from hana_ml import dataframe
from hana_ml.algorithms.pal.neural_network import MLPClassifier, MLPRegressor
from data_load_utils import Settings
from hana_ml.dataframe import create_dataframe_from_pandas
import numpy as np
import pandas as pd
from sklearn.datasets import load_iris
# save load_iris() sklearn dataset to iris
# if you'd like to check dataset type use: type(load_iris())
# if you'd like to view list of attributes use: dir(load_iris())
iris = load_iris()
# np.c_ is the numpy concatenate function
# which is used to concat iris['data'] and iris['target'] arrays
# for pandas column argument: concat iris['feature_names'] list
# and string list (in this case one string); you can make this anything you'd like..
# the original dataset would probably call this ['Species']
data = pd.DataFrame(data= np.c_[iris['data'], iris['target']],
columns= iris['feature_names'] + ['target'])
url, port, user, pwd = Settings.load_config("../../config/e2edata.ini")
connection_context = dataframe.ConnectionContext(url, port, user, pwd)
hana_df = create_dataframe_from_pandas(connection_context, data, 'IRIS', force=True)
hana_df.head(10).collect()
# +
from hana_ml.model_storage import ModelStorage
#from hana_ml.model_storage_services import ModelSavingServices
model = MLPRegressor(hidden_layer_size=[10,],
activation='TANH',
output_activation='TANH',
learning_rate=0.01,
momentum=0.001)
model.fit(hana_df, label='target')
# Creates an object called model_storage which must use the same connection with the model
model_storage = ModelStorage(connection_context=connection_context)
model_storage.clean_up()
# Saves the model for the first time
model.name = 'Model A' # The model name is mandatory
model.version = 1
model_storage.save_model(model=model)
# Lists models
model_storage.list_models()
# -
# save the model for the second time with 'replace' option and expect to have a model called 'Model A' and version is 1
model_storage.save_model(model=model, if_exists='replace')
model_storage.list_models()
# save the model for the second time with 'upgrade' option and expect to have two model called 'Model A'
# and version is 1 and 2
model_storage.save_model(model=model, if_exists='upgrade')
print(model.version)
model_storage.list_models()
# See the detail of model
model_storage.list_models('Model A', 2)['JSON'].iloc[0]
# +
# Load the model
new_model = model_storage.load_model(name='Model A', version=1)
# Predict with the loaded model
test = pd.DataFrame()
test['id'] = [1]
test['sepal length (cm)'] = [4.2]
test['sepal width (cm)'] = [3.2]
test['petal length (cm)'] = [1.2]
test['petal width (cm)'] = [0.2]
test_df = create_dataframe_from_pandas(connection_context, test, 'IRIS_TEST', force=True)
new_model.predict(test_df, key='id').collect()
# -
new_model.model_.select_statement
# enable persistent memory of a model 'Model A' with version 1
model_storage.enable_persistent_memory('Model A', 1)
# see the Runtime data of column tables M_CS_TABLES
# LOADED: If the table is loaded in memory
# PERSISTENT_MEMORY: Flag to indicate that new main part will be written to disk during table delta merge unless requested differently
import json
connection_context.sql("SELECT TABLE_NAME, LOADED, PERSISTENT_MEMORY FROM M_CS_TABLES WHERE TABLE_NAME='{}'".format(json.loads(model_storage.list_models('Model A',1)["JSON"][0])["artifacts"]["model_tables"])).collect()
# Unload from the memory
model_storage.unload_from_memory("Model A", 1, persistent_memory='retain')
# see the Runtime data of column tables M_CS_TABLES
connection_context.sql("SELECT TABLE_NAME, LOADED, PERSISTENT_MEMORY FROM M_CS_TABLES WHERE TABLE_NAME='{}'".format(json.loads(model_storage.list_models('Model A',1)["JSON"][0])["artifacts"]["model_tables"])).collect()
model_storage.disable_persistent_memory('Model A', 1)
connection_context.sql("SELECT TABLE_NAME, LOADED, PERSISTENT_MEMORY FROM M_CS_TABLES WHERE TABLE_NAME='{}'".format(json.loads(model_storage.list_models('Model A',1)["JSON"][0])["artifacts"]["model_tables"])).collect()
model_storage.load_into_memory("Model A", 1)
connection_context.sql("SELECT TABLE_NAME, LOADED, PERSISTENT_MEMORY FROM M_CS_TABLES WHERE TABLE_NAME='{}'".format(json.loads(model_storage.list_models('Model A',1)["JSON"][0])["artifacts"]["model_tables"])).collect()
# Delete a model
model_storage.delete_model('Model A', 1)
model_storage.list_models()
model_storage.list_models(name='Model A')
# Detele models with specified name and a specified range of time
start_time = '2019-07-27 07:49:48'
end_time = '2021-07-29 10:26:47'
model_storage.delete_models('Model A', start_time=start_time, end_time=end_time)
model_storage.list_models()
# Clean up all models at once
model_storage.clean_up()
# Close the connection to SAP HANA
connection_context.close()
| Python-API/pal/notebooks/irisModelStorageMLP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Imports
# +
import numpy as np
from sklearn import manifold
from matplotlib import pyplot as plt
from matplotlib.lines import Line2D
from adjustText import adjust_text
import unittest
import types
import os
from enum import Enum, auto
from ibm_watson import ApiException
from ibm_watson import NaturalLanguageUnderstandingV1
from ibm_watson.natural_language_understanding_v1 import Features, SentimentOptions
from ibm_cloud_sdk_core.authenticators import IAMAuthenticator
# Install:
# pip install numpy
# pip install -U scikit-learn
# pip install -U matplotlib
# pip install adjustText
# pip install --upgrade ibm-watson
# -
# # Classes
# +
class SentimentAnalyzer:
"""Class for analyzing sentiments in a text."""
def __init__(
self,
nlu_api_key = '<KEY>',
nlu_url = 'https://api.eu-gb.natural-language-understanding.watson.cloud.ibm.com/instances/c644c8ed-897a-4890-b692-e913c32b36bc',
version = '2021-03-25'
):
"""Constructor for Sentiment Analyzer object.
:param nlu_api_key (str, optional): API-key for IBM Watson Natural Language Understanding. Defaults to '<KEY>'.
:param nlu_url (str, optional): URL for IBM Watson Natural Language Understanding. Defaults to 'https://api.eu-gb.natural-language-understanding.watson.cloud.ibm.com/instances/c644c8ed-897a-4890-b692-e913c32b36bc'.
:param version (str, optional): Version of IBM Watson Natural Language Understanding. Defaults to '2021-03-25'.
"""
authenticator = IAMAuthenticator(nlu_api_key)
self._service = NaturalLanguageUnderstandingV1(version=version, authenticator=authenticator)
self._service.set_service_url(nlu_url)
def analyze(self, analyzable, keywords = ['deep learning', 'singularity', 'logic', 'planning', 'autonomous']):
"""Analyze a text with IBM Watson Natural Language Understanding.
:param analyzable (Analyzable): Analyzable object to analyze.
:param keywords (list(str), optional): Keywords to target for sentiment analysis. Defaults to ['deep learning', 'singularity', 'logic', 'planning', 'autonomous'].
:returns: List of Sentiment objects.
:raises TypeError: If param types are incorrect.
:raises ApiException: If API-key, URL or version params are invalid.
"""
if not isinstance(analyzable, Analyzable):
raise TypeError('Param analyzable should be Analyzable object')
elif not isinstance(keywords, list):
raise TypeError('Param keywords should be list of strings')
else:
for keyword in keywords:
if not isinstance(keyword, str):
raise TypeError('Param keywords should be list of strings')
try:
# API call
response = self._service.analyze(
text = analyzable.get_text(),
features = Features(sentiment = SentimentOptions(targets = keywords)),
)
results = response.result['sentiment']['targets']
# Create Sentiment objects per keyword
sentiments = []
for keyword in keywords:
is_present = False
for result in results:
if result['text'] == keyword:
is_present = True
score = result['score']
label = SentimentTypes[result['label'].upper()]
sentiments.append(Sentiment(keyword, score, label))
# If keyword is missing, create neutral Sentiment object
if is_present == False:
sentiments.append(Sentiment(keyword, 0, SentimentTypes.NEUTRAL))
return sentiments
except ApiException: raise
class Sentiment:
"""Class to record sentiment toward a keyword."""
def __init__(self, keyword, score, label):
"""Constructor for Sentiment object.
:param keyword (str): Keyword to which the sentiment relates.
:param score (float): Sentiment score toward the keyword.
:param label (SentimentTypes): Label of the sentiment score toward the keyword.
"""
self._keyword = keyword
self._score = score
self._label = label
def get_keyword(self):
return self._keyword
def get_score(self):
return self._score
def get_label(self):
return self._label
class SentimentTypes(Enum):
"""Enumeration of sentiment types: POSITIVE, NEUTRAL or NEGATIVE."""
POSITIVE = auto()
NEUTRAL = auto()
NEGATIVE = auto()
def get_color(self):
"""Return color related to sentiment type.
:returns: String with color name.
"""
if self == SentimentTypes.POSITIVE:
return 'green'
elif self == SentimentTypes.NEUTRAL:
return 'orange'
else:
return 'red'
class Analyzable:
"""Class for analyzable object."""
def __init__(self, text, id_ = None):
"""Constructor for Analyzable object.
:param text (str): Text to be analyzed.
:param id_ (any type): Id of the object. Defaults to None, which will induce id(self).
:raises TypeError: If type of text param is incorrect.
"""
if not isinstance(text, str):
raise TypeError('Param should be string')
self._text = text
if id_ == None:
self._id = id(self)
else:
self._id = id_
self._sentiments = []
def get_text(self):
return self._text
def get_id(self):
return self._id
def get_sentiments(self):
return self._sentiments
def get_sentiment_toward_keyword(self, keyword):
"""Return sentiment toward a given keyword.
:param keyword (str): Keyword to which the sentiment relates.
:returns: Sentiment object.
:raises NameError: If there is no sentiment toward the given keyword.
"""
if not isinstance(keyword, str):
raise TypeError('Param should be string')
is_present = False
for sentiment in self._sentiments:
if sentiment.get_keyword() == keyword:
is_present = True
return sentiment
if is_present == False:
raise NameError('No sentiment found toward given keyword')
def set_sentiments(self, sentiments):
self._sentiments = sentiments
class Person(Analyzable):
"""Class for analyzable Person."""
def __init__(self, name, text):
"""Constructor for Person object.
:param text (str): Text to be analyzed.
:param name (str): Name of the person, will be used as id of the object.
:raises TypeError: If type of name param is incorrect.
"""
if not isinstance(name, str):
raise TypeError('Param name should be string')
super().__init__(text, name)
class AnalyzableObjects:
"""Class to list and group analyzable objects."""
def __init__(self):
self._analyzables = []
def get_analyzables(self):
return self._analyzables
def append(self, obj):
"""Add analyzable object to analyzables list.
:param obj (Analyzable): Object to be added to the analyzables list.
:raises TypeError: If param type is incorrect.
"""
if not isinstance(obj, Analyzable):
raise TypeError('Param analyzable should be Analyzable object')
self._analyzables.append(obj)
def group_by_sentiment_type_toward_keyword(self, keyword):
"""Split analyzables in groups per sentiment type, based on sentiment toward a given keyword.
:param keyword (str): Keyword to which the sentiment relates.
:returns: Tuple of keyword and list of respectively positive, neutral and negative group lists.
"""
positive_group = []
neutral_group = []
negative_group = []
for obj in self._analyzables:
sentiment_type = obj.get_sentiment_toward_keyword(keyword).get_label()
if sentiment_type == SentimentTypes.POSITIVE:
positive_group.append(obj)
elif sentiment_type == SentimentTypes.NEUTRAL:
neutral_group.append(obj)
else:
negative_group.append(obj)
group = [positive_group, neutral_group, negative_group]
return (keyword, group)
def group_by_sentiment_score_toward_keyword(self, keyword, num_groups = 3):
"""Split analyzables in groups of equal size, based on sentiment score toward a given keyword.
:param keyword (str): Keyword to which the sentiment relates.
:param num_groups (int): Number of groups that should be created. Defaults to 3.
:returns: Tuple of keyword and list of given number of group lists, divided by similar sentiment scores.
:raises TypeError: If type of param num_groups is incorrect.
"""
if not isinstance(num_groups, int):
raise TypeError('Param num_groups should be integer')
copied_group = self._analyzables.copy()
copied_group.sort(key = lambda obj: obj.get_sentiment_toward_keyword(keyword).get_score())
group = [arr.tolist() for arr in np.array_split(copied_group, num_groups)]
return (keyword, group)
def print_group(self, keyword_and_group):
"""Print group members per group with id, sentiment score and sentiment type toward a given keyword.
:param keyword_and_group (tuple(str, list(list(Analyzable))): Tuple of keyword and list of group lists.
:raises TypeError: If param type is incorrect.
"""
if not isinstance(keyword_and_group, tuple) \
or not isinstance(keyword_and_group[0], str) \
or not isinstance(keyword_and_group[1], list):
raise TypeError('Param keyword_and_group should be tuple of str and list')
keyword = keyword_and_group[0]
group = keyword_and_group[1]
i = 1
print(f'Groups are divided by their sentiment toward the keyword "{keyword}".')
for lst in [lst for lst in group if len(lst) != 0]:
print(f'Group {i}:')
i += 1
for member in lst:
id_ = member.get_id()
sentiment = member.get_sentiment_toward_keyword(keyword)
sentiment_score = sentiment.get_score()
sentiment_label = sentiment.get_label().name.lower()
print(f'- {id_}: {sentiment_score} ({sentiment_label})')
def create_dissimilarities_matrix(self, keyword):
"""Create dissimilarities matrix of group members' sentiment scores toward a given keyword.
:param keyword (str): Keyword to which the sentiment relates.
:returns: Tuple of keyword and 2D ndarray with sentiment score differences among group members.
:raises ValueError: If self._analyzables is empty.
"""
if len(self._analyzables) == 0:
raise ValueError('The list of analyzable objects (self._analyzables) is empty. Add Analyzable objects with the append method.')
length = len(self._analyzables)
dissimilarities_matrix = np.empty([length, length])
for x in range(length):
for y in range(length):
score_x = self._analyzables[x].get_sentiment_toward_keyword(keyword).get_score()
score_y = self._analyzables[y].get_sentiment_toward_keyword(keyword).get_score()
dissimilarities_matrix[x,y] = abs(score_x - score_y)
return (keyword, dissimilarities_matrix)
def print_scatterplot(self, keyword_and_dissimilarities_matrix):
"""Print scatterplot of dissimilarities matrix of group members' sentiment scores toward a given keyword.
:param keyword_and_dissimilarities_matrix (tuple(str, ndarray)): Tuple of keyword and 2D ndarray of group members' sentiment scores toward a given keyword.
:raises TypeError: If param type is incorrect.
"""
if not isinstance(keyword_and_dissimilarities_matrix, tuple) \
or not isinstance(keyword_and_dissimilarities_matrix[0], str) \
or not isinstance(keyword_and_dissimilarities_matrix[1], np.ndarray):
raise TypeError('Param keyword_and_dissimilarities_matrix should be tuple of str and ndarray')
keyword = keyword_and_dissimilarities_matrix[0]
dissimilarities_matrix = keyword_and_dissimilarities_matrix[1]
# Create array with ids and colors of group members
group_members = []
for member in self._analyzables:
id_ = member.get_id()
sentiment_type = member.get_sentiment_toward_keyword(keyword).get_label()
group_members.append((id_, sentiment_type.get_color()))
# Create scatterplot
mds = model = manifold.MDS(dissimilarity='precomputed')
out = model.fit_transform(dissimilarities_matrix)
plt.figure(figsize=(12,8), dpi= 100)
plt.scatter(out[:, 0], out[:, 1], c = [member[1] for member in group_members])
plt.axis('equal')
plt.title(f'Sentiment toward "{keyword}"')
plt.legend([Line2D([0], [0], linewidth = 0, marker = 'o', color = 'green'),
Line2D([0], [0], linewidth = 0, marker = 'o', color = 'orange'),
Line2D([0], [0], linewidth = 0, marker = 'o', color = 'red')],
['Positive', 'Neutral', 'Negative'])
# Place marker labels
# Option 1: dynamically, improved label placement (slower)
labels = [member[0] for member in group_members]
texts = []
for x, y, text in zip(out[:, 0], out[:, 1], labels):
texts.append(plt.text(x, y, text))
adjust_text(texts, force_text=0.05, arrowprops=dict(arrowstyle="-|>", color='grey', alpha=0.5))
## Option 2: statically, overlapping labels (faster)
# for i, id_ in enumerate([member[0] for member in group_members]):
# plt.annotate(id_, (out[i, 0], out[i, 1]))
# -
# # Program
# +
input = [
['Auri', "I think that AI can be very interesting, because computers think with more logic than humans. The decisions are based on the results that are processed with deep learning. There are also AI systems that develop themselves with their own planning. That way systems will evolve that can reason autonomous. In the end a singularity can be generated and that will be terrible for humanity. Humans will become slaves to machines as is demonstrated in many movies and books."],
['Bast', "I think that AI can be very interesting, because computers think with more logic than humans. The decisions are based on the results that are processed with deep learning. There are also AI systems that develop themselves with their own planning. That way systems will evolve that can reason autonomous. In the end a singularity can be generated and that will be terrible for humanity. Humans will become slaves to machines as is demonstrated in many movies and books."],
['Cinder', "Deep learning is already a blessing. We are quite far from Singularity. AI logic would probably work differently than human logic. Automatic planning and scheduling can solve more complex problems. Autonomous operations by vehicles and (virtual) robots driven by AI will be a significant part of our future."],
['Deoch', "Singularity, when computer intelligence will be beyond human brain capacity and still growing might be a bit scary but fortunately we can still pull out the plug ! I guess some day autonomous computers might create conceptual new computers themselves by means of deep learning and recursive self improvement. Logic thinking is a powerfull tool in biological behaviour and survival but essential in Python programming! If you are failing to plan, you're planning to fail."],
['Elodin', "Deep learning has moved from theory to practice in the field of artificial intelligence. A singularity as a doomsday scenario is not realistic at the current and near future state of technology. But military use of autonomous devices will become a problem if you look at history and use of technology. Both strategy and tactics are a matter of planning and logic, what is what Artificial Intelligence is good at."],
['Felurian', "Deep learning is an interesting way of doing machine learning. Singularity is both fascinating and frightening. If more systems were to become autonomous, our lives would become easier. Logic is the basis for all we have discovered. Planning is a difficult thing but important."],
['Geoffrey', "Does it rain? No it is deep learning. When AI reach singularity the weather will not be predicted but AI will autonomous be planning the weather so we people can talk about the weather when we have nothing to do. That’s logic."],
['Hemme', "Deep learning could be dangerous. If a machine learns autonomously, the logic can't be traced. There is a chance of singularity happening, although then I would expect it to have happened already. There are good uses for deep learning, e.g. planning."],
['Iax', "A singularity within deep learning supposes exponential growth of computer capacity. In nature exponential growth always stops itself so this goes against any logic. Autonomous learning does make sence and will grow according to planning."],
['Jaxim', "Deep learning is a form of machine learning through multi-layered neural networks. It will lead to singularity whereby superhuman intelligences will set different priorities than humanity. AI will no longer follow human logic. I don’t know exactly when AI is planning to become autonomous, but probably within some decades."],
['Kvothe', "Today the machines are learning deeply from the logic we provide them. The singularity will happend in a very near future and if machines become autonomous, we humans will have to change the planning of our future life."],
['Lanre', "The movie 'Terminator' predicted singularity of autonomous machines. A terrible perspective on our future. The machines used logic and deep learning (I guess) for the purpose of their own existence. They acted as humans, planning their own survival, but without the ethical boundaries. If we proceed with AI development, we need to do so with caution."],
['Mola', "The singularity has been depicted as the start of a dystopian future in many science fiction movies. Fully autonomous machines with flawed logic and poor planning seem to be a catalyst to arrive at such a future. Deep learning however seems to be a mostly harmless technique."],
['Nina', "Deep learning is a machine learning technique that teaches computers to do what humans do. I hope they don't achieve singularity. That would seriously challenge all logic. Even with proper planning this will lead to poor human performance. I only have one question. Will these autonomous machines make a decent cup of coffee?"],
['<NAME>', "Deep learning is an artificial intelligence (AI) function that imitates the workings of the human brain. Deep learning will never bring us to a hypothetical point in time at which technological growth becomes uncontrollable and irreversible, resulting in unforeseeable changes to human civilization. We call the aforementioned technological singularity. However, at the hypothetical point of technological singularity indeed no autonomous machines, nor any logic reasoning, and or planning, is required anymore."],
['Penthe', "In my humble opinion the singularity will not become a reality. When I watch my daughter, and I did this for many hours, how she discovers the world I realize how limited intelligent agents are. Even with techniques like deep learning we are not even close to how a toddler is able to learn. Where the toddler will only need a bit of observation, a handful of examples and a few corrections, you will need to give the machine thousands of image to start being good at this game. Although for example AI planning and logic are tools which have a lot of practical use I do not think we will develop a fully autonomous machine that will change the course of history quite soon."],
['Rethe', "Deep learning can benefit human life significantly. As of now it is hard to tell if singularity is good or bad. The logic and planning used by autonomous machines makes them excel in tasks humans can do just mediocre."],
['Simmon', "Of course Deep Learning is interesting from a technology perspective. The technology can tell us more about how our brain works, and give us detailed insights in ourselves. From a socilogical perspective deep learning might be challenging. Whwre automation had a major effect on the blue collar worker, deep learning might have a huge effect on the white collar worker. Are we ready for those effects? On top of that if we support a medical doctor in their diagnosis, based on deep learning, who will be leading, and who will be responsible in case of a misdiagnosis. Will that be the devloper who wrote the algoritm or will it be the doctor who missed. Challenges whihc are not so much in technology but in society, and we need to solve them, before we start to use the technology at large scale. Logic can be defined as the proof or validation behind any reason provided. In the run for transparency in the use of artifcial intelligence, but also in the run for automation, show the logic is important. You need to show how you deduct towards a certain conclucsion or action. I sure hope we learn this in this course. Before Corona I had a large discussion with an architect from RDW about Autonomous cars. He liked the concept and really wanted to start pilots with it. And the technology is cool he said. But society is not ready, because a complete autonomous car has huge implications on the insurance, the road infrastructure etc. What are you going to do when an autonomous vehicle makes an accident. Who is responsible? The passenger, because the people in the car are not driving, it is the software. Is it the developer of the software, or is the supplier of the sensors? Is this car manufacturer? How do you solve this? And then how do you handle exceptions that you have not planned for. because in life there will be exceptions which is the key capability of a human. So I do think it is difficult to easily state what is going to happen. The technological singularity—also, simply, the singularity —is a hypothetical point in time at which technological growth becomes uncontrollable and irreversible, resulting in unforeseeable changes to human civilization. With the read of 'The Shallows' from N. Carr I would state that we already entered this inflection point. With the use of navigation systems I do experience myself that I am less able to navigate without the solution. What will it be for my kids. Also if we look at education, I learned fact and figures, important to get a proper understanding of history timelines. I do see with my kids, that they will look for details at the internet, and I do see it with myself during this course also. That changes learning, and approaches in life. And I do see in the practice of midwifery a change, people come with their Googled diagnosis, and have less respect for the professional opinion of the midwife. let alone the discussion on how we get news, and how this influences us. So I do see a huge change in society, and I think this is irreversible. Planning is the process of thinking about the activities required to achieve a desired goal. For tasks that are done repeatedly I think AI is wonderful solution. And you could embrace several data sources to extend this."],
['Tempi', "Deep learning is an exciting and promising application of computing technology that can facilitate further automation. Enabling computers to become autonomous in tasks that previously required human intelligence will further increase productivity and increase prosperity. A key aspect is that computers are now able to move away from simple logic-based tasks and perform tasks that involve learning, planning, etcetera. Though this technology is promising, dystopian (or utopian) prospects of a techonolocigal singularity seem unfounded in the foreseeable future. Progress is still very much incremental, with machines being adapted to new tasks gradually and mostly under human supervision. Even a state-of-the-art deep learning algorithm such as Deep Mind requires extensive work to adapt to new tasks, and it is still headline news when it outperforms human experts."],
]
analyzer = SentimentAnalyzer()
analyzables = AnalyzableObjects()
# Create Person objects with sentiments and add them to the analyzables list
for arr in input:
person = Person(arr[0], arr[1])
sentiments = analyzer.analyze(person)
person.set_sentiments(sentiments)
analyzables.append(person)
keyword = 'deep learning'
# Divide analyzables in groups of equal size by sentiment score toward keyword
group_by_sentiment_score = analyzables.group_by_sentiment_score_toward_keyword(keyword)
analyzables.print_group(group_by_sentiment_score)
print('')
# Divide analyzables in groups per sentiment type toward keyword
group_by_sentiment_type = analyzables.group_by_sentiment_type_toward_keyword(keyword)
analyzables.print_group(group_by_sentiment_type)
print('')
# Print scatterplot of analyzables' sentiment toward keyword
dissimilarities_matrix = analyzables.create_dissimilarities_matrix(keyword)
analyzables.print_scatterplot(dissimilarities_matrix)
# -
# # Unittests
# +
class TestSentimentAnalyzer(unittest.TestCase):
"""Unittest for class SentimentAnalyzer."""
def setUp(self):
self.analyzer = SentimentAnalyzer()
self.analyzable = Analyzable('This is a test.')
def test_sentiment_analyzer_analyze_invalid_analyzable_arg(self):
with self.assertRaises(TypeError):
self.analyzer.analyze('test')
obj = types.SimpleNamespace()
obj.text = 'This is a test.'
with self.assertRaises(TypeError):
self.analyzer.analyze(obj)
def test_sentiment_analyzer_analyze_invalid_keywords_arg(self):
with self.assertRaises(TypeError):
self.analyzer.analyze(self.analyzable, 'test')
with self.assertRaises(TypeError):
self.analyzer.analyze(self.analyzable, [1, 2])
class TestSentimentTypes(unittest.TestCase):
"""Unittest for class SentimentTypes"""
def test_sentiment_types_get_color_correct_result_type(self):
color = SentimentTypes.POSITIVE.get_color()
self.assertIsInstance(color, str)
def test_sentiment_types_get_color_correct_result_values(self):
self.assertEqual(SentimentTypes.POSITIVE.get_color(),'green')
self.assertEqual(SentimentTypes.NEUTRAL.get_color(), 'orange')
self.assertEqual(SentimentTypes.NEGATIVE.get_color(), 'red')
class TestAnalyzable(unittest.TestCase):
"""Unittest for class Analyzable."""
def setUp(self):
self.analyzable = Analyzable('This is a test.')
self.sentiments = [Sentiment('test', 0.6, SentimentTypes.POSITIVE),
Sentiment('deep learning', -0.2, SentimentTypes.NEGATIVE),
Sentiment('university', 0, SentimentTypes.NEUTRAL)]
self.analyzable.set_sentiments(self.sentiments)
def test_analyzable_invalid_text_arg(self):
with self.assertRaises(TypeError):
Analyzable(1)
with self.assertRaises(TypeError):
Analyzable(['test 1', 'test 2'])
def test_analyzable_get_sentiment_toward_keyword_invalid_keyword_arg(self):
with self.assertRaises(TypeError):
self.analyzable.get_sentiment_toward_keyword(1)
with self.assertRaises(TypeError):
self.analyzable.get_sentiment_toward_keyword(['test 1', 'test 2'])
def test_analyzable_get_sentiment_toward_keyword_no_sentiment_found(self):
with self.assertRaises(NameError):
self.analyzable.get_sentiment_toward_keyword('test 1')
def test_analyzable_get_sentiment_toward_keyword_correct_result_type(self):
sentiment = self.analyzable.get_sentiment_toward_keyword('test')
self.assertIsInstance(sentiment, Sentiment)
def test_analyzable_get_sentiment_toward_keyword_correct_result_values(self):
sentiment = self.analyzable.get_sentiment_toward_keyword('test')
self.assertEqual(sentiment, self.sentiments[0])
sentiment = self.analyzable.get_sentiment_toward_keyword('deep learning')
self.assertEqual(sentiment, self.sentiments[1])
sentiment = self.analyzable.get_sentiment_toward_keyword('university')
self.assertEqual(sentiment, self.sentiments[2])
class TestPerson(unittest.TestCase):
"""Unittest for class Person."""
def setUp(self):
self.person = Person('Name', 'This is a test.')
def test_person_invalid_name_arg(self):
text = 'This is a test.'
with self.assertRaises(TypeError):
Person(True, text)
with self.assertRaises(TypeError):
Person(1, text)
with self.assertRaises(TypeError):
Person(['Name 1', 'Name 2'], text)
def test_person_correct_inheritance_of_analyzable(self):
self.assertIsInstance(self.person, Analyzable)
def test_person_correct_transfer_of_text_arg_to_analyzable_object(self):
self.assertEqual(self.person.get_text(), 'This is a test.')
def test_person_correct_transfer_of_name_arg_to_analyzable_object(self):
self.assertEqual(self.person.get_id(), 'Name')
class TestAnalyzableObjects(unittest.TestCase):
"""Unittest for class AnalyzableObjects."""
def setUp(self):
self.person1 = Person('Name 1', 'This is a test.')
sentiments1 = [Sentiment('test', 0.6, SentimentTypes.POSITIVE)]
self.person1.set_sentiments(sentiments1)
self.person2 = Person('Name 2', 'This is another unit test.')
sentiments2 = [Sentiment('test', -0.4, SentimentTypes.NEGATIVE)]
self.person2.set_sentiments(sentiments2)
self.analyzables = AnalyzableObjects()
self.keyword = 'test'
def test_analyzable_objects_append_invalid_obj_arg(self):
with self.assertRaises(TypeError):
self.analyzables.append('Test')
obj = types.SimpleNamespace()
obj.text = 'This is a test.'
with self.assertRaises(TypeError):
self.analyzables.append(obj)
def test_analyzable_objects_append_correct_result_values(self):
self.analyzables.append(self.person1)
self.assertEqual(len(self.analyzables.get_analyzables()), 1)
self.analyzables.append(self.person2)
self.assertEqual(len(self.analyzables.get_analyzables()), 2)
self.assertEqual(self.analyzables.get_analyzables()[0], self.person1)
self.assertEqual(self.analyzables.get_analyzables()[1], self.person2)
def test_analyzable_objects_group_by_sentiment_type_toward_keyword_correct_result_type(self):
self.analyzables.append(self.person1)
result = self.analyzables.group_by_sentiment_type_toward_keyword(self.keyword)
self.assertIsInstance(result, tuple)
self.assertIsInstance(result[0], str)
self.assertIsInstance(result[1], list)
for lst in result[1]:
for obj in lst:
self.assertIsInstance(obj, Analyzable)
def test_analyzable_objects_group_by_sentiment_type_toward_keyword_correct_result_values(self):
self.analyzables.append(self.person1)
result = self.analyzables.group_by_sentiment_type_toward_keyword(self.keyword)
for lst in result[1]:
if len(lst) > 0:
lst_set = set([obj.get_sentiment_toward_keyword(self.keyword).get_label() for obj in lst])
self.assertEqual(len(lst_set), 1)
self.assertTrue(self.person1 in result[1][0])
def test_analyzable_objects_group_by_sentiment_score_toward_keyword_invalid_num_groups_arg(self):
self.analyzables.append(self.person1)
with self.assertRaises(TypeError):
self.analyzables.group_by_sentiment_score_toward_keyword(self.keyword, 3.5)
with self.assertRaises(TypeError):
self.analyzables.group_by_sentiment_score_toward_keyword(self.keyword, 'test')
with self.assertRaises(TypeError):
self.analyzables.group_by_sentiment_score_toward_keyword(self.keyword, [1, 2])
def test_analyzable_objects_group_by_sentiment_score_toward_keyword_correct_result_type(self):
self.analyzables.append(self.person1)
result = self.analyzables.group_by_sentiment_score_toward_keyword(self.keyword)
self.assertIsInstance(result, tuple)
self.assertIsInstance(result[0], str)
self.assertIsInstance(result[1], list)
for lst in result[1]:
for obj in lst:
self.assertIsInstance(obj, Analyzable)
def test_analyzable_objects_group_by_sentiment_score_toward_keyword_correct_result_values(self):
num_groups = 3
self.analyzables.append(self.person1)
result = self.analyzables.group_by_sentiment_score_toward_keyword(self.keyword, num_groups)
self.assertEqual(len(result[1]), num_groups)
self.assertTrue(self.person1 in result[1][0])
def test_analyzable_objects_print_group_invalid_keyword_and_group_arg_no_tuple(self):
self.analyzables.append(self.person1)
group = self.analyzables.group_by_sentiment_type_toward_keyword(self.keyword)
with self.assertRaises(TypeError):
self.analyzables.print_group(self.keyword)
with self.assertRaises(TypeError):
self.analyzables.print_group([self.keyword, group])
def test_analyzable_objects_print_group_invalid_keyword_and_group_arg_tuple_without_str(self):
self.analyzables.append(self.person1)
group = self.analyzables.group_by_sentiment_type_toward_keyword(self.keyword)
with self.assertRaises(TypeError):
self.analyzables.print_group((1, group))
with self.assertRaises(TypeError):
self.analyzables.print_group(([self.keyword], group))
def test_analyzable_objects_print_group_invalid_keyword_and_group_arg_tuple_without_list(self):
self.analyzables.append(self.person1)
with self.assertRaises(TypeError):
self.analyzables.print_group((self.keyword, 'Object'))
with self.assertRaises(TypeError):
self.analyzables.print_group((self.keyword, self.person1))
def test_analyzable_objects_create_dissimilarities_matrix_empty_analyzables_list(self):
with self.assertRaises(ValueError):
self.analyzables.create_dissimilarities_matrix(self.keyword)[1]
def test_analyzable_objects_create_dissimilarities_matrix_correct_result_type(self):
self.analyzables.append(self.person1)
self.analyzables.append(self.person2)
dissimilarities_matrix = self.analyzables.create_dissimilarities_matrix(self.keyword)[1]
self.assertIsInstance(dissimilarities_matrix, np.ndarray)
for x in range(len(dissimilarities_matrix)):
for y in range(len(dissimilarities_matrix)):
self.assertIsInstance(dissimilarities_matrix[x,y], float)
def test_analyzable_objects_create_dissimilarities_matrix_correct_result_values(self):
self.analyzables.append(self.person1)
self.analyzables.append(self.person2)
dissimilarities_matrix = self.analyzables.create_dissimilarities_matrix(self.keyword)[1]
self.assertEqual(dissimilarities_matrix.shape, (2,2))
self.assertEqual(dissimilarities_matrix[0,0], 0.0)
self.assertEqual(dissimilarities_matrix[1,1], 0.0)
self.assertEqual(dissimilarities_matrix[0,1], 1.0)
self.assertEqual(dissimilarities_matrix[1,0], 1.0)
def test_analyzable_objects_print_scatterplot_invalid_keyword_and_dissimilarities_matrix_arg_no_tuple(self):
self.analyzables.append(self.person1)
dissimilarities_matrix = self.analyzables.create_dissimilarities_matrix(self.keyword)
with self.assertRaises(TypeError):
self.analyzables.print_scatterplot(self.keyword)
with self.assertRaises(TypeError):
self.analyzables.print_scatterplot([self.keyword, dissimilarities_matrix])
def test_analyzable_objects_print_scatterplot_invalid_keyword_and_dissimilarities_matrix_tuple_without_str(self):
self.analyzables.append(self.person1)
dissimilarities_matrix = self.analyzables.create_dissimilarities_matrix(self.keyword)
with self.assertRaises(TypeError):
self.analyzables.print_scatterplot((1, dissimilarities_matrix))
with self.assertRaises(TypeError):
self.analyzables.print_scatterplot(([self.keyword], dissimilarities_matrix))
def test_analyzable_objects_print_scatterplot_invalid_keyword_and_dissimilarities_matrix_tuple_without_ndarray(self):
self.analyzables.append(self.person1)
with self.assertRaises(TypeError):
self.analyzables.print_scatterplot((self.keyword, 'Matrix'))
with self.assertRaises(TypeError):
self.analyzables.print_scatterplot((self.keyword, 1))
with self.assertRaises(TypeError):
self.analyzables.print_scatterplot((self.keyword, [1, 2]))
unittest.main(argv=[''], verbosity=2, exit=False)
# -
| ai_student_grouping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import csv
# +
temple_ids = {}
with open('chinese_temple_list.csv', encoding="utf8") as csv_file:
csv_reader = csv.reader(csv_file, delimiter=',')
chinese_temple_list = list(csv_reader)
for temple in chinese_temple_list:
# # check duplicated temple names
# if temple[4] in temple_ids:
# print(temple[4])
temple_ids[temple[4]] = temple[0]
# -
temple_ids
import pandas as pd
file = '112aec6807a515c3.xlsx'
df = pd.read_excel(file)
print(df)
traditional_names = df['tarditional_chinese'].tolist()
traditional_names = [str(t).strip() for t in traditional_names]
traditional_names
fixed_folders = df['Folder Name'].tolist()
fixed_folders = [str(t).strip() for t in fixed_folders]
fixed_folders
folder_path = r'D:\照片分类\zhaopian\Gallery\Chinese Temple'
folderlist = os.listdir(folder_path)
folderlist
converted_pairs_df = pd.DataFrame(columns=['original_name', 'converted_name'])
__original_names = []
__converted_names = []
# +
folders_not_found = []
converted_folders = []
for folder in folderlist:
chunks = folder.split('_')
found = False
for chunk in chunks:
if chunk.strip() in traditional_names:
found = True
converted = df['objectid'].iloc[traditional_names.index(chunk.strip())]
if converted in converted_folders:
_converted = str(converted) + '_' + str(converted_folders.count(converted))
# _converted will be the converted folder name
__original_names.append(folder)
__converted_names.append(_converted)
print(folder, '=>', _converted)
else:
# converted will be the converted folder name
__original_names.append(folder)
__converted_names.append(converted)
print(folder, '=>', converted)
converted_folders.append(converted)
break
if not found:
if folder.strip() in fixed_folders:
fixed_folder = df['objectid'].iloc[fixed_folders.index(folder.strip())]
if fixed_folder in converted_folders:
_fixed_folder = str(fixed_folder) + '_' + str(converted_folders.count(fixed_folder))
# _fixed_folder will be the converted folder name
__original_names.append(folder)
__converted_names.append(_fixed_folder)
print('###', folder, '=>', _fixed_folder)
else:
# fixed_folder will be the converted folder name
__original_names.append(folder)
__converted_names.append(fixed_folder)
print('###', folder, '=>', fixed_folder)
converted_folders.append(fixed_folder)
else:
# these are not found
folders_not_found.append(folder)
for f in folders_not_found:
print('---', f)
# -
len(folders_not_found)
data = {'original_name':__original_names, 'converted_name':__converted_names}
converted_pairs_df = pd.DataFrame.from_dict(data)
converted_pairs_df
converted_pairs_df.to_csv ('converted_pairs.csv', index = None, header=True)
# test renaming folders
os.rename(r'C:\Users\eva\Desktop\to_rename_this_文件夹', r'C:\Users\eva\Desktop\to_rename_this_文件夹_')
for index, row in converted_pairs_df.iterrows():
# print(row['original_name'], row['converted_name'])
o_name = 'D:\\照片分类\\zhaopian\\Gallery\\Chinese Temple\\'
o_name += str(row['original_name'])
c_name = 'D:\\照片分类\\zhaopian\\Gallery\\Chinese Temple\\'
c_name += str(row['converted_name'])
if os.path.exists(o_name):
os.rename(o_name, c_name)
print(o_name)
print(c_name)
| folder name converter/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.2 64-bit (''DS_38'': conda)'
# name: python38264bitds38condadd9b54eec7b04875a6bc324f2f32b4a4
# ---
# # Is the goal time random ?
# Goal is a rare event in soccer games. The goal time is random in the sense that for a specific game, it is not possible to give any sound prediction on it. However, from a statistical point of view, there should be some pattern for the time distribution of goal time. Is it evenly distributed for every minute, or there is more chance to have a goal in the second half as the defending side is not as energetic as in the first half ? Or maybe the opposite is true, because it is more likely to be an open game in the first half ?
#
# Let's try to answer these questions using the [Wyscount dataset](https://www.nature.com/articles/s41597-019-0247-7). This is a very valuable dataset made open by Wyscout including spatial and temporal events in matches for the entire 2017-2018 season of the first league in England, Frence, Germany, Italy and Spain, the European Champions League, and the 2018 FIFA World Cup as well.
# +
# load some packages
# %load_ext autoreload
# %autoreload 2
import matplotlib.pyplot as plt
import numpy as np
import json
import pandas as pd
import os
from utils import wyscout
# -
# load the dataset from https://figshare.com/collections/Soccer_match_event_dataset/4415000/5
wyscout.download_data()
# use the Premier League as an example
dirname = './data/Wyscout/events'
fn = os.path.join(dirname, 'events_England.json')
season = '2017-2018'
league = 'Premier League'
# read in event data using pandas --> DataFrame
df_events = pd.read_json(fn)
df_events.head()
# find goal tag
# check the meaning of the tags @https://apidocs.wyscout.com/matches-wyid-events
tag2name = pd.read_csv('./data/Wyscout/tags2name.csv', sep=';')
goal_tag = tag2name[tag2name['Description']=='Goal'].Tag.values[0]
print('Goal event is labelled as tag {}'.format(goal_tag))
# +
# add a tag_list column
df_events['tag_list'] = df_events.tags.apply(lambda x: [i['id'] for i in x])
# extract goal events
goal_events = df_events[df_events.tag_list.apply(lambda x: goal_tag in x)]
goal_events.tail()
# -
# note that among these events, there are also Save attempts that need to be excluded
print(goal_events.eventName.unique())
goal_events = goal_events[goal_events.eventName!='Save attempt']
# Among these goals, some from shots, some from free kicks including penalties
shot_goals = goal_events.groupby('eventName').get_group('Shot')
freekick_goals = goal_events.groupby('eventName').get_group('Free Kick')
print('In total {} goals, {} from shots, {} from free kicks'.format(len(goal_events), len(shot_goals), len(freekick_goals)))
# ## First or Second Half ?
# +
# separate goals in first and second half time
goal_1H_sec = goal_events[goal_events.matchPeriod=='1H'].eventSec.values
goal_2H_sec = goal_events[goal_events.matchPeriod=='2H'].eventSec.values
# transform from seconds to minutes
goal_1H_min = goal_1H_sec//60
goal_2H_min = goal_2H_sec//60
# +
plt.style.use('./utils/default_plot_style.mplstyle')
max_H = int(max([np.max(goal_1H_min), np.max(goal_2H_min)]))
bins_1H = range(0, 51, 5)
bins_2H = range(55, 106, 5)
xticks = range(0, 101, 5)
xticklabels = [str(i)+'-'+str(i+5) for i in range(0, 90, 5)]
xticklabels.insert(9, '>45')
xticklabels.insert(10, '')
xticklabels.append('>90')
fig, ax = plt.subplots(1,1,figsize=(12,6))
ax.hist(goal_1H_min, bins=bins_1H, rwidth=0.75, align='left')
ax.hist(goal_2H_min + 55, bins=bins_2H, rwidth=0.75, align='left')
ax.axvline(x=50, color='r', linestyle='--') # add half time line
ax.text(x=0.5, y=0.75, s='half time', transform=ax.transAxes, ha='center', va='center', rotation=90, fontsize=20, bbox=dict(facecolor='white'))
ax.set_title(league + ' ' + season)
ax.set_xlabel('Match Time (min)')
ax.set_ylabel('Goals')
ax.set_xticks(xticks)
ax.set_xticklabels(xticklabels, rotation=30)
plt.show()
fig.savefig('./figs/PL_score_time.png', dpi=300)
# -
n_matches = df_events.groupby('matchId').ngroups
goal_diff_per_match = (len(goal_2H_min) - len(goal_1H_min))/n_matches
goal_ratio = len(goal_2H_min)/len(goal_1H_min) - 1
print('Second half has a {:.2} more goals in average'.format(goal_diff_per_match))
print('Second half has a {:.1%} higher goal probability'.format(goal_ratio))
# ### Other National Leagues
# Goals in the second half clearly outnumber that in the first half. Let us take a look at the other national leagues.
# +
# First, wrap up the steps above into functions
def extract_goal_events(df):
# make a tag_list
tag_list = df.tags.apply(lambda x: [i['id'] for i in x])
# extract goal events
goal_events = df[tag_list.apply(lambda x: goal_tag in x)] # goal_tag is global
# exclude save attempt
goal_events = goal_events[goal_events.eventName!='Save attempt']
return goal_events
def plot_scoring_time(goal, time='sec', ax=None, title=None):
goal_1H = goal[0]
goal_2H = goal[1]
if time=='sec':
# transform from seconds to minutes
goal_1H = goal_1H//60
goal_2H = goal_2H//60
max_H = int(max([np.max(goal_1H), np.max(goal_2H)]))
bins_1H = range(0, 51, 5)
bins_2H = range(55, 106, 5)
xticks = range(0, 101, 5)
xticklabels = [str(i)+'-'+str(i+5) for i in range(0, 90, 5)]
xticklabels.insert(9, '>45')
xticklabels.insert(10, '')
xticklabels.append('>90')
if ax is None:
fig, ax = plt.subplots(1,1,figsize=(12,6))
ax.hist(goal_1H, bins=bins_1H, rwidth=0.75, align='left')
ax.hist(goal_2H + 55, bins=bins_2H, rwidth=0.75, align='left')
ax.axvline(x=50, color='r', linestyle='--') # add half time line
ax.text(x=0.5, y=0.75, s='half time', transform=ax.transAxes, ha='center', va='center', rotation=90, fontsize=20, bbox=dict(facecolor='white'))
ax.set_title(title)
ax.set_ylabel('Goals')
ax.set_xticks(xticks)
ax.set_xticklabels(xticklabels, rotation=30)
return
# +
# plot the time distribution of goals for all the five leagues
leagues = ['England', 'France', 'Germany', 'Italy', 'Spain']
goal_time_league = []
fig, axes = plt.subplots(nrows=len(leagues), ncols=1, figsize=(12, len(leagues)*5))
for league, ax in zip(leagues, axes):
fn = os.path.join(dirname, 'events_'+league+'.json')
df = pd.read_json(fn)
goal_events = extract_goal_events(df)
goal_1H = goal_events[goal_events.matchPeriod=='1H'].eventSec.values
goal_2H = goal_events[goal_events.matchPeriod=='2H'].eventSec.values
goal_time_league.append((goal_1H, goal_2H))
plt.style.use('./utils/default_plot_style.mplstyle')
plot_scoring_time((goal_1H, goal_2H), ax=ax, title=league)
plt.xlabel('Match Time (min)')
plt.show()
fig.savefig('./figs/leagues_scoring_time.png', dpi=300)
# -
for league, (goal_1H, goal_2H) in zip(leagues, goal_time_league):
n_goal_1H = len(goal_1H)
n_goal_2H = len(goal_2H)
a = n_goal_2H/n_goal_1H - 1
print('{:8} : 1H - {} goals, 2H - {} goals ({:.1%} higher)'.format(league, n_goal_1H, n_goal_2H, a))
# An interesting observation here is that the English and Spanish leagues have much more second-half goals.
# ### Home or Away Matters ?
#
# The home tends to attack because they don't want to accept a draw or lose in front of the home fans.
# match information is stored in the matches_[League].json file
match_file = './data/Wyscout/matches/matches_England.json'
# add homeaway feature in the dataframe
events = wyscout.add_event_cols(df_events, ['homeaway'], match_file)
goal = extract_goal_events(events)
goal_home = goal[goal['homeaway']=='home']
goal_away= goal[goal['homeaway']=='away']
# +
homeaway = ['Home', 'Away']
goal_events = [goal_home, goal_away]
goal_time_ha = []
fig, axes = plt.subplots(nrows=len(homeaway), ncols=1, figsize=(12, 14))
for ha, events, ax in zip(homeaway, goal_events, axes):
fn = os.path.join(dirname, 'events_'+league+'.json')
df = pd.read_json(fn)
goal_1H = events[events.matchPeriod=='1H'].eventSec.values
goal_2H = events[events.matchPeriod=='2H'].eventSec.values
goal_time_ha.append((goal_1H, goal_2H))
plt.style.use('./utils/default_plot_style.mplstyle')
plot_scoring_time((goal_1H, goal_2H), ax=ax, title=ha+'\n Premier League 2017-2018')
plt.xlabel('Match Time (min)')
plt.show()
fig.savefig('./figs/homeaway_scoring_time.png', dpi=300)
# -
for ha, (goal_1H, goal_2H) in zip(homeaway, goal_time_ha):
n_goal_1H = len(goal_1H)
n_goal_2H = len(goal_2H)
a = n_goal_2H/n_goal_1H - 1
print('{:5} : 1H - {} goals, 2H - {} goals ({:.1%} higher)'.format(ha, n_goal_1H, n_goal_2H, a))
# Clearly, the home team has a much higher probability to score in the second half by a increase of over 50%, while the away team only has 20% increase for a second-half goal.
| score_time.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Modelling of Atmospheric Clodus
# #### copyright: Jagiellonian University
# #### licence: CC-BY
# #### author: <NAME>
# ## Class 4 (March 16, virtual)
#
# Let's try to reproduce Figure 7.4 from Rogers & Yau
# (first and last panel this week, middle panels next week)
# 
# ### previously on...
import numpy as np
import pint
from matplotlib import pyplot
si = pint.UnitRegistry()
si.setup_matplotlib()
class Constants:
from scipy import constants
import mendeleev as pt
# polynomial fot to equilibrium vapour pressure wrt water (coefficients from Flatau et al. 1992)
# doi:10.1175/1520-0450(1992)031<1507%3APFTSVP>2.0.CO%3B2
c_w = (6.115836990e000, 0.444606896e000, 0.143177157e-01, 0.264224321e-03, 0.299291081e-05,
0.203154182e-07, 0.702620698e-10, 0.379534310e-13, -.321582393e-15)
T0 = T0 = constants.zero_Celsius * si.kelvin
def __molar_mass(x):
return x.atomic_weight * si.gram / si.mole
M_a = (
0.78 * __molar_mass(pt.N) * 2 +
0.21 * __molar_mass(pt.O) * 2 +
0.01 * __molar_mass(pt.Ar)
)
M_v = __molar_mass(pt.O) + __molar_mass(pt.H) * 2
R_str = constants.R * si.joule / si.kelvin / si.mole
R_a = R_str / M_a
R_v = R_str / M_v
g = constants.g * si.metre / si.second**2
# the only new things!
l_v = 2.5e6 * si.joule / si.kilogram
c_p = 1000 * si.joule / si.kilogram / si.kelvin
class Formulae:
@staticmethod
def rho(p, R, T):
return p / (R * T)
@staticmethod
def __p_sat(temperature, coefficients, valid_range):
from numpy.polynomial.polynomial import polyval
value = polyval(temperature.to(si.celsius).magnitude, coefficients)
if isinstance(temperature.magnitude, np.ndarray):
value[np.logical_or(temperature < valid_range[0], temperature > valid_range[1])] = np.nan
else:
value = np.nan if not valid_range[0] < temperature <= valid_range[1] else value
return value * si.hectopascals
@staticmethod
def p_eq(T):
return Formulae.__p_sat(T, Constants.c_w, (Constants.T0-85 * si.kelvin, np.inf * si.kelvin))
# ### let's play with numpy, pint and scipy
class Storage:
class __Impl(np.ndarray):
p_idx, p_unit = 0, si.hectopascals
T_idx, T_unit = 1, si.kelvins
m_idx, m_unit = 2, si.grams
z_unit = si.metres
@property
def p(self):
return self[self.p_idx] * self.p_unit
@p.setter
def p(self, value):
self[self.p_idx] = value.to(self.p_unit) / self.p_unit
@property
def T(self):
return self[self.T_idx] * self.T_unit
@T.setter
def T(self, value):
self[self.T_idx] = value.to(self.T_unit) / self.T_unit
@property
def m(self):
return self[self.m_idx] * self.m_unit
@m.setter
def m(self, value):
self[self.m_idx] = value.to(self.m_unit) / self.m_unit
@staticmethod
def __make_storage(shape):
storage = Storage.__Impl(shape)
return storage
@staticmethod
def make_state():
return Storage.__make_storage((3,))
@staticmethod
def make_deriv(state):
storage = Storage.__make_storage(state.shape)
storage.p_unit /= storage.z_unit
storage.T_unit /= storage.z_unit
storage.m_unit /= storage.z_unit
return storage
@staticmethod
def view_state(array):
storage = Storage.__make_storage(array.shape)
storage[:] = array[:]
return storage
# ### the system we need to solve (super simplified "parcel model")
# $$
# \begin{eqnarray}
# \frac{dp}{dz} &=& - \rho g \\
# \frac{dm}{dz} &=& \frac{\max(0, m_v - m_{eq})}{dz} \\
# \frac{dT}{dz} &=& \frac{1}{c_p} \left(\frac{1}{\rho}\frac{dp}{dz} + \frac{l_v}{m_a} \frac{dm}{dz} \right)
# \end{eqnarray}
# $$
# $p$: pressure
# $z$: vertical displacement
# $\rho$: density
# $g$: gravitational acceleration
# $m$: mass of liquid water
# $m_v$: mass of water vapour
# $m_{eq}$: mass of water vapour at saturation (in phase equilibrium wrt water surface)
# $T$: temperature
# $c_p$: specific heat of air
# $l_v$: latent heat of vapourisation
# $m_a$: mass of air
# ### let's implement the system according to SciPy API
class System:
def __init__(self, pv0, volume, state):
assert state.m == 0
self.m_a = Formulae.rho(state.p, Constants.R_a, state.T) * volume
self.m_w = Formulae.rho(pv0, Constants.R_v, state.T) * volume
def __call__(self, _, state, dz):
state = Storage.view_state(state)
deriv = Storage.make_deriv(state)
rho = Formulae.rho(state.p, Constants.R_a, state.T)
volume = self.m_a / rho
p_eq = Formulae.p_eq(state.T)
m_eq = Formulae.rho(p_eq, Constants.R_v, state.T) * volume
m_v = self.m_w - state.m
# hydrostatic pressure
deriv.p = -Formulae.rho(state.p, Constants.R_a, state.T) * Constants.g
# saturation adjustment
deriv.m = max(0 * state.m_unit, m_v - m_eq) / dz
# heat budget
deriv.T = (deriv.p/rho + deriv.m/self.m_a * Constants.l_v) / Constants.c_p
return deriv
# ### before using sophisticated solvers from SciPy (next week), let's use a simplest one
def solve(system, state, displacement, dz):
nz = int(displacement / dz)
z = [0.] * nz * state.z_unit
states = np.repeat(state.reshape(-1, 1), nz, axis=1)
dz_magn = dz.to(state.z_unit).magnitude
for i, _ in enumerate(z[1:], start=1):
z[i] = z[i-1] + dz
states[:, i] = states[:, i-1] + dz_magn * system(_, states[:, i-1], dz)
return states, z
# ### and let's finally do the calculations ...
# +
state = Storage.make_state()
state.p = 1000 * si.hectopascals
state.T = 300 * si.kelvins
state.m = 0 * si.grams
volume = 1 * si.metre**3
pv0 = .995 * Formulae.p_eq(state.T)
displacement = 100 * si.metres
system = System(pv0, volume, state)
integ, z = solve(system, state, displacement, dz = .1 * si.metre)
# -
# ### ... and plotting
# +
m_v = system.m_w - integ.m
volume = system.m_a / Formulae.rho(integ.p, Constants.R_a, integ.T)
p_v = m_v / volume * Constants.R_v * integ.T
p_eq = Formulae.p_eq(integ.T)
fig, axs = pyplot.subplots(1, 2, sharey=True)
axs[0].step(100 * (p_v / p_eq - 1), z, label='Supersaturation [%]')
axs[1].step(integ.m / volume, z, color='red', label='Liquid water content')
fig.legend()
# -
# ## TBC!
| notebooks/04_parcel_bulk.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: voila_opencv
# language: python
# name: voila_opencv
# ---
# # Purpose
#
# Develop an example of using [Loky](https://github.com/joblib/loky) to do multiprocessing on my laptop.
# # Approach
#
# Install loky in the virtual environment I am using for my Jupyter kernel for this notebook.
#
# $ source /Users/nordin/python_envs/voila_opencv/.venv/bin/activate
# $ pip install loky
# Installing collected packages: cloudpickle, loky
# Successfully installed cloudpickle-1.6.0 loky-2.9.0
import numpy as np
from loky import get_reusable_executor
# +
# https://cosmiccoding.com.au/tutorials/multiprocessing
def slow_fn(args):
""" Simulated an optimisation problem with args coming in
and function value being output """
n = 1000
y = 0
for j in range(n):
j = j / n
for i, p in enumerate(args):
y += j * (p ** (i + 1))
return y / n
def get_jobs(num_jobs=512, num_args=5):
""" Simulated sampling our parameter space multiple times """
return [j for j in np.random.random((num_jobs, num_args))]
jobs = get_jobs()
# -
# %%time
# Check out this single core performance
for job in jobs:
slow_fn(job)
# %%time
executor = get_reusable_executor(max_workers=6)
results = list(executor.map(slow_fn, jobs, chunksize=16));
len(results)
| python/multiprocessing/example_use_loky.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/microprediction/humpday/blob/main/black_box_optimization_package_recommender.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="dOItaUcAIxs8"
# !pip install --upgrade git+https://github.com/microprediction/humpday.git
# !pip install scikit-optimize
# !pip install nevergrad
# !pip install optuna
# !pip install platypus-opt
# !pip install poap
# !pip install pysot
# !pip install bayesian-optimization
# !pip install cmake
# !pip install ultraopt
# !pip install dlib
# !pip install ax-platform
# !pip install py-bobyqa
# !pip install hebo
# !pip install nlopt
# + id="GGfPDj6ERPU5"
from humpday import recommend
import time
import math
from pprint import pprint
# + [markdown] id="V1yKlwYgJLuQ"
# # A script to recommend a black box optimizer
# There are lots of package out there. This will generate a shortlist of Python global derivative free optimizers.
# + colab={"base_uri": "https://localhost:8080/"} id="e2Ff1ZyVJGBR" outputId="03aafbfc-9e3e-430c-fcd9-e79bbaba597e"
def my_objective(u):
# Swap this out for your own.
# Domain must be [0,1]^n
time.sleep(0.01)
return u[0]*math.sin(u[1])
my_objective(u=[0.2,0.5])
# + [markdown] id="x3myV_06JO09"
# ## Run the recommender
# This will puke a list of optimizers, informed by ongoing battles reported in the [Optimizer Elo Ratings](https://microprediction.github.io/optimizer-elo-ratings/html_leaderboards/all_d08_n80.html)
# + colab={"base_uri": "https://localhost:8080/"} id="dWwUwleVJIQT" outputId="ed3866f3-9f3b-4797-d303-41042e6624b9"
recommendations = recommend(my_objective, n_dim=4, n_trials=130)
pprint(recommendations[:7])
# + [markdown] id="7IP3OxYNO4mo"
# Any of these can be called from the humpday package.
# + [markdown] id="dGIFSpq0M9pm"
# # Feeling lucky?
# Go ahead and use the meta-minimizer. This uses the top recommendation.
# + colab={"base_uri": "https://localhost:8080/", "height": 334} id="aHG1YJGiM_-T" outputId="7b82a666-1713-4aaa-ea99-2a1c9b6bba60"
from humpday import minimize
best_val, best_x = minimize(my_objective, n_dim=3, n_trials=80 )
print('Best x ='+str(best_x)+' corresponding to a value of '+str(best_val))
# + [markdown] id="aikEN8PsQcMC"
# No luck? Occasionally these optimizers fail on corner cases. Occasionally.
# + [markdown] id="oIQvl8G6Qm6r"
# # Rather choose your own?
# If you see something you like, you can instantiate it easily:
# + colab={"base_uri": "https://localhost:8080/", "height": 198} id="Mjgt3DuUQPr4" outputId="23f9fa89-1215-4cc8-f4d4-0c794b7e3707"
from humpday.optimizers.alloptimizers import optimizer_from_name
optim = optimizer_from_name('pymoo_pattern_cube')
optim(my_objective,n_dim=4, n_trials=130)
# + [markdown] id="ncULVsRVPLUs"
# # Points race
# Here's something else you can do that is a bit more time intensive, but probably very helpful. The 'points race' utility will see how all the recommended optimizers perform on your objective function (or set of objective functions). It reports a running total of points, where 3 points are assigned for first place, 2 for second and 1 for third.
# + id="Hd0gHMSJQ7d9"
def my_other_objective(u):
# Swap this out for your own.
# Domain must be [0,1]^n
time.sleep(0.01)
return u[0]*u[0]*math.cos(u[1])
# + colab={"base_uri": "https://localhost:8080/"} id="-HK2wVpvQv12" outputId="849b2777-c98a-460e-d300-7df635dee4a7"
from humpday import points_race
points_race(objectives=[my_objective, my_other_objective],n_dim=5, n_trials=100)
| black_box_optimization_package_recommender.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Sankey_story
#
# - toc: true
# - badges: true
# - comments: true
# - sticky_rank: 1
# - author: <NAME>
# - image: images/diagram.png
# - categories: [fastpages, jupytern sankey]
#
# +
#hide
from influxdb_client import InfluxDBClient
import panel as pn
pn.extension('plotly')
import pandas as pd
import time
import requests
from decouple import config
from urllib.parse import urlparse
import influxdb_client
from influxdb_client.client.write_api import SYNCHRONOUS
from snippets import *
import plotly.graph_objs as go
TOKEN = config("TOKEN")
ORG = config("ORG")
INFLUXDB_URL = config("INFLUXDB_URL")
# -
#hide
scenarios = {}
# # 100% Elec
# Sur les différentes années de ce projet, la maison et son usage ont évolués, d'un coté l'isolation a été améliorée ponctuellement par certains travaux, d'un autre coté la famille s'est agrandie. Je fais le choix de figer les besoins de la maison en chauffage, ECS et électricité domestique afin de pouvoir comparer différentes configuration sur une base commune.
#
# Par rapport au DPE, j'ai considéré les besoins suivants:
#
# - Besoin en chauffage : 15 200 kWh annuel, soit +30% par rapport à l'estimation du DPE initial. Cette valeur est estimée à partir du suivi de consommation des factures sur plusieurs années. Il est cependant difficile de suivre correctement la consommation de bois d'une année sur l'autre. L'estimation du besoin du DPE est fait par rapport à une valeur de consigne de chauffage à 19°C {% fn 1 %}, en pratique nous avons une température de consigne à 21°C.
# - Production d'ECS 1800kWh annuel. Il est estimée à 600 kWh/personne/an {% fn 2 %}, en considérant 2 adultes et 2 enfants,
# - Besoin en électricité domestique : 3998kWh annuel. Il est estimé à 1100 kWh/an/personne {% fn 3 %} soit 3300 kWh annuel pour 2 adultes et 2 enfants. Il faut ajouter la consommation spécifique de la piscine, en considérant 6 h de fonctionnement par jour pendant 5 mois et une puissance de pompe de 750 W, la consommation annuelle est de 697 kWh. Ces données sont cohérentes avec les consommations constatées sur plusieurs années.
#
# Je vais utiliser des diagrammes Sankey, en partant de la configuration la plus simple ou seule l'électricité est utilisée pour répondre aux besoins de la maison:
# +
Ratio_pertes = 2.58
Frac_nucleaire=0.65
Frac_ENR_reseau=0.25
Frac_gaz=1-Frac_ENR_reseau-Frac_nucleaire
rdt_chaud=0.95
rdt_insert=0.75 #(0.90 sinon)
PV_frac_autoconso = 1
nb_pers = 3
Total_chauff_SdB = 15200
Total_ECS = 600*nb_pers
Total_pompe_piscine = 6*5*31*0.750
Total_conso_elec = 1100*nb_pers + Total_chauff_SdB + Total_ECS + Total_pompe_piscine
Total_conso_elec_reseau = Total_conso_elec #*PV_frac_autoconso
Part_PAC=0
Total_conso_gaz = 0*(1-Part_PAC)
Total_conso_bois = 3500*0
Frac_recup_bois = 0
Total_prod_PV = 0
Total_revente_PV = (Total_prod_PV-Total_conso_elec*(1-PV_frac_autoconso))
Total_charge_batterie = Total_prod_PV *0.3*0
Total_decharge_batterie = Total_charge_batterie*0.85*0
Total_prod_thermique = 0
Total_prod_thermique_piscine = 0
Total_th_PAC = 10000*Part_PAC
sCOP = 1.8*2.5
Total_thermique = Total_conso_gaz*rdt_chaud + Total_prod_thermique + Frac_recup_bois*Total_conso_bois + Total_th_PAC
# +
#hide
fig = go.Figure(go.Sankey(
arrangement = "snap",
hoverinfo="all",
textfont=dict(color="black", size=25),
node = {
"label": ["Non renouvelable", # 0
"ENR", # 1
"Nucléaire", # 2
"Gaz", # 3
"ENR Réseau", # 4
"Photovoltaïque", # 5
"Solaire thermique", # 6
"Air (PAC)", # 7
"Bois", # 8
"Electricité Réseau", # 9
"Pertes", # 10
"Electricité domestique", # 11
"Usage direct", # 12
"Batterie", # 13
"Revente", # 14
"Production eau chaude", # 15
"Chauffage", # 16
"ECS", # 17
"Chauffage piscine", # 18
"PAC", # 19
"Autre", # 20
"Piscine", # 21
"Piscine" # 22
],
# "x": [0.2, 0.1, 0.5, 0.7, 0.3, 0.5],
# "y": [0.7, 0.5, 0.2, 0.4, 0.2, 0.3],
'pad':25}, # 10 Pixels
link = {
"source": [0, 0,
1, 1, 1, 1, 1,
2,
3,
4,
9,9,
5,
11,11,11,
14,
13,
3,
6,
7,
8,
15,12,
6,
12,
8,
12,
19,
12,
13,
3,
8,
12,
18,
21
],
"target": [2, 3,
9, 5, 6, 7, 8,
9,
9,
9,
12,10,
11,
12,13,14,
4,
12,
15,
15,
19,
15,
16,17,
18,
16,
16,
19,
15,
20,
10,
10,
10,
21,
22,
22
],
"value": [Total_conso_elec_reseau*Ratio_pertes*Frac_nucleaire, Total_conso_elec_reseau*Ratio_pertes*Frac_gaz+ Total_conso_gaz,
Total_conso_elec_reseau*Ratio_pertes*Frac_ENR_reseau, Total_prod_PV, Total_prod_thermique+Total_prod_thermique_piscine, Total_th_PAC*(1-1/sCOP), Total_conso_bois,
Total_conso_elec_reseau*Ratio_pertes*Frac_nucleaire,
Total_conso_elec_reseau*Ratio_pertes*Frac_gaz,
Total_conso_elec_reseau*Ratio_pertes*Frac_ENR_reseau*0, # + INJ
Total_conso_elec_reseau, Total_conso_elec_reseau*(Ratio_pertes-1),
Total_prod_PV,
Total_conso_elec*0-Total_decharge_batterie*0, Total_charge_batterie, Total_revente_PV, ###
Total_revente_PV,
Total_decharge_batterie,
Total_conso_gaz*rdt_chaud,
Total_prod_thermique,
Total_th_PAC*(1-1/sCOP),
Total_conso_bois*rdt_insert*Frac_recup_bois,
Total_thermique-Total_ECS,Total_ECS,
Total_prod_thermique_piscine,
Total_chauff_SdB,
Total_conso_bois*rdt_insert*(1-Frac_recup_bois),
Total_th_PAC/sCOP,
Total_th_PAC,
Total_conso_elec-Total_th_PAC/sCOP-Total_chauff_SdB-Total_pompe_piscine -Total_ECS,
Total_charge_batterie-Total_decharge_batterie,
Total_conso_gaz*(1-rdt_chaud),
Total_conso_bois*(1-rdt_insert),
Total_pompe_piscine,
Total_prod_thermique_piscine,
Total_pompe_piscine*0
]
}))
sankey_plotly_elec = pn.pane.Plotly(fig)
scenarios['100% électricité'] = sankey_plotly_elec
# -
#hide
filename = '../_includes/sankey_plotly_elec.html'
export_plot_fastpages(pn.panel(sankey_plotly_elec), filename)
# {% include sankey_plotly_elec.html %}
# # Elec + Bois
# +
#hide
import plotly.graph_objects as go
Ratio_pertes = 2.58
Frac_nucleaire=0.65
Frac_ENR_reseau=0.25
Frac_gaz=1-Frac_ENR_reseau-Frac_nucleaire
rdt_chaud=0.95
rdt_insert=0.75 #(0.90 sinon)
PV_frac_autoconso = 1
nb_pers = 3
Total_conso_bois = 3500
Total_chauff_SdB = 15200 - Total_conso_bois*rdt_insert
Total_ECS = 600*nb_pers
Total_pompe_piscine = 6*5*31*0.750
Total_conso_elec = 1100*nb_pers + Total_chauff_SdB + Total_ECS + Total_pompe_piscine
Total_conso_elec_reseau = Total_conso_elec #*PV_frac_autoconso
Part_PAC=0
Total_conso_gaz = 0*(1-Part_PAC)
Frac_recup_bois = 0
Total_prod_PV = 0
Total_revente_PV = (Total_prod_PV-Total_conso_elec*(1-PV_frac_autoconso))
Total_charge_batterie = Total_prod_PV *0.3*0
Total_decharge_batterie = Total_charge_batterie*0.85*0
Total_prod_thermique = 0
Total_prod_thermique_piscine = 0
Total_th_PAC = 10000*Part_PAC
sCOP = 1.8*2.5
Total_thermique = Total_conso_gaz*rdt_chaud + Total_prod_thermique + Frac_recup_bois*Total_conso_bois + Total_th_PAC
fig = go.Figure(go.Sankey(
arrangement = "snap",
textfont=dict(color="black", size=25),
node = {
"label": ["Non renouvelable", # 0
"ENR", # 1
"Nucléaire", # 2
"Gaz", # 3
"ENR Réseau", # 4
"Photovoltaïque", # 5
"Solaire thermique", # 6
"Air (PAC)", # 7
"Bois", # 8
"Electricité Réseau", # 9
"Pertes", # 10
"Electricité domestique", # 11
"Usage direct", # 12
"Batterie", # 13
"Revente", # 14
"Prod eau chaude", # 15
"Chauffage", # 16
"ECS", # 17
"Chauffage piscine", # 18
"PAC", # 19
"Autre", # 20
"Piscine", # 21
"Piscine" # 22
],
# "x": [0.2, 0.1, 0.5, 0.7, 0.3, 0.5],
# "y": [0.7, 0.5, 0.2, 0.4, 0.2, 0.3],
'pad':20}, # 10 Pixels
link = {
"source": [0, 0,
1, 1, 1, 1, 1,
2,
3,
4,
9,9,
5,
11,11,11,
14,
13,
3,
6,
7,
8,
15,12,
6,
12,
8,
12,
19,
12,
13,
3,
8,
12,
18,
21
],
"target": [2, 3,
4, 5, 6, 7, 8,
9,
9,
9,
12,10,
11,
12,13,14,
4,
12,
15,
15,
19,
15,
16,17,
18,
16,
16,
19,
15,
20,
10,
10,
10,
21,
22,
22
],
"value": [Total_conso_elec_reseau*Ratio_pertes*Frac_nucleaire, Total_conso_elec_reseau*Ratio_pertes*Frac_gaz+ Total_conso_gaz,
Total_conso_elec_reseau*Ratio_pertes*Frac_ENR_reseau, Total_prod_PV, Total_prod_thermique+Total_prod_thermique_piscine, Total_th_PAC*(1-1/sCOP), Total_conso_bois,
Total_conso_elec_reseau*Ratio_pertes*Frac_nucleaire,
Total_conso_elec_reseau*Ratio_pertes*Frac_gaz,
Total_conso_elec_reseau*Ratio_pertes*Frac_ENR_reseau, # + INJ
Total_conso_elec_reseau, Total_conso_elec_reseau*(Ratio_pertes-1),
Total_prod_PV,
Total_conso_elec*0-Total_decharge_batterie*0, Total_charge_batterie, Total_revente_PV, ###
Total_revente_PV,
Total_decharge_batterie,
Total_conso_gaz*rdt_chaud,
Total_prod_thermique,
Total_th_PAC*(1-1/sCOP),
Total_conso_bois*rdt_insert*Frac_recup_bois,
Total_thermique-Total_ECS,Total_ECS,
Total_prod_thermique_piscine,
Total_chauff_SdB,
Total_conso_bois*rdt_insert*(1-Frac_recup_bois),
Total_th_PAC/sCOP,
Total_th_PAC,
Total_conso_elec-Total_th_PAC/sCOP-Total_chauff_SdB-Total_pompe_piscine -Total_ECS,
Total_charge_batterie-Total_decharge_batterie,
Total_conso_gaz*(1-rdt_chaud),
Total_conso_bois*(1-rdt_insert),
Total_pompe_piscine,
Total_prod_thermique_piscine,
Total_pompe_piscine*0
]
}))
sankey_plotly_elec_bois = pn.pane.Plotly(fig)
scenarios['Electricité & Bois'] = sankey_plotly_elec_bois
filename = '../_includes/sankey_plotly_elec_bois.html'
export_plot_fastpages(pn.panel(sankey_plotly_elec_bois), filename)
# -
# {% include sankey_plotly_elec_bois.html %}
# # Rotex
# +
#hide
import plotly.graph_objects as go
Ratio_pertes = 2.58
Frac_nucleaire=0.65
Frac_ENR_reseau=0.25
Frac_gaz=1-Frac_ENR_reseau-Frac_nucleaire
rdt_chaud=0.95
rdt_insert=0.75 #(0.90 sinon)
PV_frac_autoconso = 1
nb_pers = 3
Total_conso_bois = 3500
Total_chauff_SdB = 200
Total_ECS = 600*nb_pers
Total_pompe_piscine = 6*5*31*0.750
Total_conso_elec = 1100*nb_pers + Total_chauff_SdB + Total_pompe_piscine
Total_conso_elec_reseau = Total_conso_elec*PV_frac_autoconso
Part_PAC=0
Total_prod_thermique = 3000
Total_conso_gaz = (15200 - Total_conso_bois*rdt_insert - Total_chauff_SdB - Total_prod_thermique )*(1-Part_PAC)+ Total_ECS
Frac_recup_bois = 0.3*0
Total_prod_PV = 5900*0
Total_revente_PV = (Total_prod_PV-Total_conso_elec*(1-PV_frac_autoconso))*0
Total_charge_batterie = Total_prod_PV *0.3*0
Total_decharge_batterie = Total_charge_batterie*0.85*0
Total_prod_thermique_piscine = 3800*0
Total_th_PAC = (15200 - Total_conso_bois*rdt_insert - Total_chauff_SdB )*Part_PAC
sCOP = 1.8*2.5
Total_thermique = Total_conso_gaz*rdt_chaud + Total_prod_thermique + Frac_recup_bois*Total_conso_bois + Total_th_PAC
fig = go.Figure(go.Sankey(
arrangement = "snap",
textfont=dict(color="black", size=25),
node = {
"label": ["Non renouvelable", # 0
"ENR", # 1
"Nucléaire", # 2
"Gaz", # 3
"ENR Réseau", # 4
"Photovoltaïque", # 5
"Solaire thermique", # 6
"Air (PAC)", # 7
"Bois", # 8
"Electricité Réseau", # 9
"Pertes", # 10
"Electricité domestique", # 11
"Usage direct", # 12
"Batterie", # 13
"Revente", # 14
"Prod eau chaude", # 15
"Chauffage", # 16
"ECS", # 17
"Chauffage piscine", # 18
"PAC", # 19
"Autre", # 20
"Piscine", # 21
"Piscine" # 22
],
# "x": [0.2, 0.1, 0.5, 0.7, 0.3, 0.5],
# "y": [0.7, 0.5, 0.2, 0.4, 0.2, 0.3],
'pad':20}, # 10 Pixels
link = {
"source": [0, 0,
1, 1, 1, 1, 1,
2,
3,
4,
9,9,
5,
11,11,11,
14,
13,
3,
6,
7,
8,
15,15,
6,
12,
8,
12,
19,
12,
13,
3,
8,
12,
18,
21
],
"target": [2, 3,
4, 5, 6, 7, 8,
9,
9,
9,
12,10,
11,
12,13,14,
4,
12,
15,
15,
19,
15,
16,17,
18,
16,
16,
19,
15,
20,
10,
10,
10,
21,
22,
22
],
"value": [Total_conso_elec_reseau*Ratio_pertes*Frac_nucleaire, Total_conso_elec_reseau*Ratio_pertes*Frac_gaz+ Total_conso_gaz,
Total_conso_elec_reseau*Ratio_pertes*Frac_ENR_reseau, Total_prod_PV, Total_prod_thermique+Total_prod_thermique_piscine, Total_th_PAC*(1-1/sCOP), Total_conso_bois,
Total_conso_elec_reseau*Ratio_pertes*Frac_nucleaire,
Total_conso_elec_reseau*Ratio_pertes*Frac_gaz,
Total_conso_elec_reseau*Ratio_pertes*Frac_ENR_reseau, # + INJ
Total_conso_elec_reseau, Total_conso_elec_reseau*(Ratio_pertes-1),
Total_prod_PV,
Total_conso_elec*0-Total_decharge_batterie, Total_charge_batterie, Total_revente_PV, ###
Total_revente_PV,
Total_decharge_batterie,
Total_conso_gaz*rdt_chaud,
Total_prod_thermique,
Total_th_PAC*(1-1/sCOP),
Total_conso_bois*rdt_insert*Frac_recup_bois,
Total_thermique-Total_ECS,Total_ECS,
Total_prod_thermique_piscine,
Total_chauff_SdB,
Total_conso_bois*rdt_insert*(1-Frac_recup_bois),
Total_th_PAC/sCOP,
Total_th_PAC,
Total_conso_elec-Total_th_PAC/sCOP-Total_chauff_SdB-Total_pompe_piscine,
Total_charge_batterie-Total_decharge_batterie,
Total_conso_gaz*(1-rdt_chaud),
Total_conso_bois*(1-rdt_insert),
Total_pompe_piscine,
Total_prod_thermique_piscine,
Total_pompe_piscine*0
]
}))
sankey_plotly_rotex = pn.pane.Plotly(fig)
scenarios['Combiné gaz condensation/solaire'] = sankey_plotly_rotex
filename = '../_includes/sankey_plotly_rotex.html'
export_plot_fastpages(pn.panel(sankey_plotly_rotex), filename)
# -
# {% include sankey_plotly_rotex.html %}
# ## Rotex gaz uniquement
# +
#hide
import plotly.graph_objects as go
Ratio_pertes = 2.58
Frac_nucleaire=0.65
Frac_ENR_reseau=0.25
Frac_gaz=1-Frac_ENR_reseau-Frac_nucleaire
rdt_chaud=0.95
rdt_insert=0.75 #(0.90 sinon)
PV_frac_autoconso = 1
nb_pers = 3
Total_conso_bois = 3500
Total_chauff_SdB = 200
Total_ECS = 600*nb_pers
Total_pompe_piscine = 6*5*31*0.750
Total_conso_elec = 1100*nb_pers + Total_chauff_SdB + Total_pompe_piscine
Total_conso_elec_reseau = Total_conso_elec*PV_frac_autoconso
Part_PAC=0
Total_prod_thermique = 3000*0
Total_conso_gaz = (15200 - Total_conso_bois*rdt_insert - Total_chauff_SdB - Total_prod_thermique )*(1-Part_PAC)+ Total_ECS
Frac_recup_bois = 0.3*0
Total_prod_PV = 5900*0
Total_revente_PV = (Total_prod_PV-Total_conso_elec*(1-PV_frac_autoconso))*0
Total_charge_batterie = Total_prod_PV *0.3*0
Total_decharge_batterie = Total_charge_batterie*0.85*0
Total_prod_thermique_piscine = 3800*0
Total_th_PAC = (15200 - Total_conso_bois*rdt_insert - Total_chauff_SdB )*Part_PAC
sCOP = 1.8*2.5
Total_thermique = Total_conso_gaz*rdt_chaud + Total_prod_thermique + Frac_recup_bois*Total_conso_bois + Total_th_PAC
fig = go.Figure(go.Sankey(
arrangement = "snap",
textfont=dict(color="black", size=25),
node = {
"label": ["Non renouvelable", # 0
"ENR", # 1
"Nucléaire", # 2
"Gaz", # 3
"ENR Réseau", # 4
"Photovoltaïque", # 5
"Solaire thermique", # 6
"Air (PAC)", # 7
"Bois", # 8
"Electricité Réseau", # 9
"Pertes", # 10
"Electricité domestique", # 11
"Usage direct", # 12
"Batterie", # 13
"Revente", # 14
"Prod eau chaude", # 15
"Chauffage", # 16
"ECS", # 17
"Chauffage piscine", # 18
"PAC", # 19
"Autre", # 20
"Piscine", # 21
"Piscine" # 22
],
# "x": [0.2, 0.1, 0.5, 0.7, 0.3, 0.5],
# "y": [0.7, 0.5, 0.2, 0.4, 0.2, 0.3],
'pad':20}, # 10 Pixels
link = {
"source": [0, 0,
1, 1, 1, 1, 1,
2,
3,
4,
9,9,
5,
11,11,11,
14,
13,
3,
6,
7,
8,
15,15,
6,
12,
8,
12,
19,
12,
13,
3,
8,
12,
18,
21
],
"target": [2, 3,
4, 5, 6, 7, 8,
9,
9,
9,
12,10,
11,
12,13,14,
4,
12,
15,
15,
19,
15,
16,17,
18,
16,
16,
19,
15,
20,
10,
10,
10,
21,
22,
22
],
"value": [Total_conso_elec_reseau*Ratio_pertes*Frac_nucleaire, Total_conso_elec_reseau*Ratio_pertes*Frac_gaz+ Total_conso_gaz,
Total_conso_elec_reseau*Ratio_pertes*Frac_ENR_reseau, Total_prod_PV, Total_prod_thermique+Total_prod_thermique_piscine, Total_th_PAC*(1-1/sCOP), Total_conso_bois,
Total_conso_elec_reseau*Ratio_pertes*Frac_nucleaire,
Total_conso_elec_reseau*Ratio_pertes*Frac_gaz,
Total_conso_elec_reseau*Ratio_pertes*Frac_ENR_reseau, # + INJ
Total_conso_elec_reseau, Total_conso_elec_reseau*(Ratio_pertes-1),
Total_prod_PV,
Total_conso_elec*0-Total_decharge_batterie, Total_charge_batterie, Total_revente_PV, ###
Total_revente_PV,
Total_decharge_batterie,
Total_conso_gaz*rdt_chaud,
Total_prod_thermique,
Total_th_PAC*(1-1/sCOP),
Total_conso_bois*rdt_insert*Frac_recup_bois,
Total_thermique-Total_ECS,Total_ECS,
Total_prod_thermique_piscine,
Total_chauff_SdB,
Total_conso_bois*rdt_insert*(1-Frac_recup_bois),
Total_th_PAC/sCOP,
Total_th_PAC,
Total_conso_elec-Total_th_PAC/sCOP-Total_chauff_SdB-Total_pompe_piscine,
Total_charge_batterie-Total_decharge_batterie,
Total_conso_gaz*(1-rdt_chaud),
Total_conso_bois*(1-rdt_insert),
Total_pompe_piscine,
Total_prod_thermique_piscine,
Total_pompe_piscine*0
]
}))
sankey_plotly_rotex_gaz = pn.pane.Plotly(fig)
scenarios['Gaz seul'] = sankey_plotly_rotex_gaz
filename = '../_includes/sankey_plotly_rotex_gaz.html'
export_plot_fastpages(pn.panel(sankey_plotly_rotex_gaz), filename)
# -
# {% include sankey_plotly_rotex_gaz.html %}
# # Rotex + piscine
# +
#hide
import plotly.graph_objects as go
Ratio_pertes = 2.58
Frac_nucleaire=0.65
Frac_ENR_reseau=0.25
Frac_gaz=1-Frac_ENR_reseau-Frac_nucleaire
rdt_chaud=0.95
rdt_insert=0.75 #(0.90 sinon)
PV_frac_autoconso = 1
nb_pers = 3
Total_conso_bois = 3500
Total_chauff_SdB = 200
Total_ECS = 600*nb_pers
Total_pompe_piscine = 6*5*31*0.750
Total_conso_elec = 1100*nb_pers + Total_chauff_SdB + Total_pompe_piscine
Total_conso_elec_reseau = Total_conso_elec*PV_frac_autoconso
Part_PAC=0
Total_prod_thermique = 3000
Total_conso_gaz = (15200 - Total_conso_bois*rdt_insert - Total_chauff_SdB - Total_prod_thermique)*(1-Part_PAC) + Total_ECS
Frac_recup_bois = 0.3*0
Total_prod_PV = 5900*0
Total_revente_PV = (Total_prod_PV-Total_conso_elec*(1-PV_frac_autoconso))*0
Total_charge_batterie = Total_prod_PV *0.3*0
Total_decharge_batterie = Total_charge_batterie*0.85*0
Total_prod_thermique_piscine = 3800
Total_th_PAC = (15200 - Total_conso_bois*rdt_insert - Total_chauff_SdB )*Part_PAC
sCOP = 1.8*2.5
Total_thermique = Total_conso_gaz*rdt_chaud + Total_prod_thermique + Frac_recup_bois*Total_conso_bois + Total_th_PAC
fig = go.Figure(go.Sankey(
arrangement = "snap",
textfont=dict(color="black", size=25),
node = {
"label": ["Non renouvelable", # 0
"ENR", # 1
"Nucléaire", # 2
"Gaz", # 3
"ENR Réseau", # 4
"Photovoltaïque", # 5
"Solaire thermique", # 6
"Air (PAC)", # 7
"Bois", # 8
"Electricité Réseau", # 9
"Pertes", # 10
"Electricité domestique", # 11
"Usage direct", # 12
"Batterie", # 13
"Revente", # 14
"Prod eau chaude", # 15
"Chauffage", # 16
"ECS", # 17
"Chauffage piscine", # 18
"PAC", # 19
"Autre", # 20
"P<NAME>", # 21
"Piscine" # 22
],
# "x": [0.2, 0.1, 0.5, 0.7, 0.3, 0.5],
# "y": [0.7, 0.5, 0.2, 0.4, 0.2, 0.3],
'pad':40}, # 10 Pixels
link = {
"source": [0, 0,
1, 1, 1, 1, 1,
2,
3,
4,
9,9,
5,
11,11,11,
14,
13,
3,
6,
7,
8,
15,15,
6,
12,
8,
12,
19,
12,
13,
3,
8,
12,
18,
21
],
"target": [2, 3,
4, 5, 6, 7, 8,
9,
9,
9,
12,10,
11,
12,13,14,
4,
12,
15,
15,
19,
15,
16,17,
18,
16,
16,
19,
15,
20,
10,
10,
10,
21,
22,
22
],
"value": [Total_conso_elec_reseau*Ratio_pertes*Frac_nucleaire, Total_conso_elec_reseau*Ratio_pertes*Frac_gaz+ Total_conso_gaz,
Total_conso_elec_reseau*Ratio_pertes*Frac_ENR_reseau, Total_prod_PV, Total_prod_thermique+Total_prod_thermique_piscine, Total_th_PAC*(1-1/sCOP), Total_conso_bois,
Total_conso_elec_reseau*Ratio_pertes*Frac_nucleaire,
Total_conso_elec_reseau*Ratio_pertes*Frac_gaz,
Total_conso_elec_reseau*Ratio_pertes*Frac_ENR_reseau, # + INJ
Total_conso_elec_reseau, Total_conso_elec_reseau*(Ratio_pertes-1),
Total_prod_PV,
Total_conso_elec*0-Total_decharge_batterie, Total_charge_batterie, Total_revente_PV, ###
Total_revente_PV,
Total_decharge_batterie,
Total_conso_gaz*rdt_chaud,
Total_prod_thermique,
Total_th_PAC*(1-1/sCOP),
Total_conso_bois*rdt_insert*Frac_recup_bois,
Total_thermique-Total_ECS,Total_ECS,
Total_prod_thermique_piscine,
Total_chauff_SdB,
Total_conso_bois*rdt_insert*(1-Frac_recup_bois),
Total_th_PAC/sCOP,
Total_th_PAC,
Total_conso_elec-Total_th_PAC/sCOP-Total_chauff_SdB-Total_pompe_piscine,
Total_charge_batterie-Total_decharge_batterie,
Total_conso_gaz*(1-rdt_chaud),
Total_conso_bois*(1-rdt_insert),
Total_pompe_piscine,
Total_prod_thermique_piscine,
Total_pompe_piscine
]
}))
sankey_plotly_rotex_piscine = pn.pane.Plotly(fig)
scenarios['Combiné gaz condensation/solaire & Piscine'] = sankey_plotly_rotex_piscine
filename = '../_includes/sankey_plotly_rotex_piscine.html'
export_plot_fastpages(pn.panel(sankey_plotly_rotex_piscine), filename)
# -
# {% include sankey_plotly_rotex_piscine.html %}
# # Rotex + piscine + bouilleur
# +
#hide
import plotly.graph_objects as go
Ratio_pertes = 2.58
Frac_nucleaire=0.65
Frac_ENR_reseau=0.25
Frac_gaz=1-Frac_ENR_reseau-Frac_nucleaire
rdt_chaud=0.95
rdt_insert=0.90
PV_frac_autoconso = 1
nb_pers = 3
Total_conso_bois = 3500
Total_chauff_SdB = 200
Total_ECS = 600*nb_pers
Total_pompe_piscine = 6*5*31*0.750
Total_conso_elec = 1100*nb_pers + Total_chauff_SdB + Total_pompe_piscine
Total_conso_elec_reseau = Total_conso_elec*PV_frac_autoconso
Part_PAC=0
Total_prod_thermique = 3000
Total_conso_gaz = (15200 - Total_conso_bois*rdt_insert - Total_chauff_SdB - Total_prod_thermique)*(1-Part_PAC)+ Total_ECS
Frac_recup_bois = 0.3
Total_prod_PV = 5900*0
Total_revente_PV = (Total_prod_PV-Total_conso_elec*(1-PV_frac_autoconso))*0
Total_charge_batterie = Total_prod_PV *0.3*0
Total_decharge_batterie = Total_charge_batterie*0.85*0
Total_prod_thermique_piscine = 3800
Total_th_PAC = (15200 - Total_conso_bois*rdt_insert - Total_chauff_SdB)*Part_PAC
sCOP = 1.8*2.5
Total_thermique = Total_conso_gaz*rdt_chaud + Total_prod_thermique + Frac_recup_bois*Total_conso_bois + Total_th_PAC
fig = go.Figure(go.Sankey(
arrangement = "snap",
textfont=dict(color="black", size=25),
node = {
"label": ["Non renouvelable", # 0
"ENR", # 1
"Nucléaire", # 2
"Gaz", # 3
"ENR Réseau", # 4
"Photovoltaïque", # 5
"Solaire thermique", # 6
"Air (PAC)", # 7
"Bois", # 8
"Electricité Réseau", # 9
"Pertes", # 10
"Electricité domestique", # 11
"Usage direct", # 12
"Batterie", # 13
"Revente", # 14
"Prod eau chaude", # 15
"Chauffage", # 16
"ECS", # 17
"Chauffage piscine", # 18
"PAC", # 19
"Autre", # 20
"Pompe Piscine", # 21
"Piscine" # 22
],
# "x": [0.2, 0.1, 0.5, 0.7, 0.3, 0.5],
# "y": [0.7, 0.5, 0.2, 0.4, 0.2, 0.3],
'pad':40}, # 10 Pixels
link = {
"source": [0, 0,
1, 1, 1, 1, 1,
2,
3,
4,
9,9,
5,
11,11,11,
14,
13,
3,
6,
7,
8,
15,15,
6,
12,
8,
12,
19,
12,
13,
3,
8,
12,
18,
21
],
"target": [2, 3,
4, 5, 6, 7, 8,
9,
9,
9,
12,10,
11,
12,13,14,
4,
12,
15,
15,
19,
15,
16,17,
18,
16,
16,
19,
15,
20,
10,
10,
10,
21,
22,
22
],
"value": [Total_conso_elec_reseau*Ratio_pertes*Frac_nucleaire, Total_conso_elec_reseau*Ratio_pertes*Frac_gaz+ Total_conso_gaz,
Total_conso_elec_reseau*Ratio_pertes*Frac_ENR_reseau, Total_prod_PV, Total_prod_thermique+Total_prod_thermique_piscine, Total_th_PAC*(1-1/sCOP), Total_conso_bois,
Total_conso_elec_reseau*Ratio_pertes*Frac_nucleaire,
Total_conso_elec_reseau*Ratio_pertes*Frac_gaz,
Total_conso_elec_reseau*Ratio_pertes*Frac_ENR_reseau, # + INJ
Total_conso_elec_reseau, Total_conso_elec_reseau*(Ratio_pertes-1),
Total_prod_PV,
Total_conso_elec*0-Total_decharge_batterie, Total_charge_batterie, Total_revente_PV, ###
Total_revente_PV,
Total_decharge_batterie,
Total_conso_gaz*rdt_chaud,
Total_prod_thermique,
Total_th_PAC*(1-1/sCOP),
Total_conso_bois*rdt_insert*Frac_recup_bois,
Total_thermique-Total_ECS,Total_ECS,
Total_prod_thermique_piscine,
Total_chauff_SdB,
Total_conso_bois*rdt_insert*(1-Frac_recup_bois),
Total_th_PAC/sCOP,
Total_th_PAC,
Total_conso_elec-Total_th_PAC/sCOP-Total_chauff_SdB-Total_pompe_piscine,
Total_charge_batterie-Total_decharge_batterie,
Total_conso_gaz*(1-rdt_chaud),
Total_conso_bois*(1-rdt_insert),
Total_pompe_piscine,
Total_prod_thermique_piscine,
Total_pompe_piscine
]
}))
sankey_plotly_rotex_piscine_builleur = pn.pane.Plotly(fig)
scenarios['Combiné gaz condensation/solaire & Piscine & Bouilleur'] = sankey_plotly_rotex_piscine_builleur
filename = '../_includes/sankey_plotly_rotex_piscine_builleur.html'
export_plot_fastpages(pn.panel(sankey_plotly_rotex_piscine_builleur), filename)
# -
# {% include sankey_plotly_rotex_piscine_builleur.html %}
# # Rotex + piscine + bouilleur + PAC
# +
#hide
import plotly.graph_objects as go
Ratio_pertes = 2.58
Frac_nucleaire=0.65
Frac_ENR_reseau=0.25
Frac_gaz=1-Frac_ENR_reseau-Frac_nucleaire
rdt_chaud=0.95
rdt_insert=0.90
PV_frac_autoconso = 1
nb_pers = 3
Total_conso_bois = 3500
Total_chauff_SdB = 200
Total_ECS = 600*nb_pers
Total_pompe_piscine = 6*5*31*0.750
alpha=0.77
Part_PAC=0.87
Total_prod_thermique = 3000
Total_conso_gaz = (15200 - Total_conso_bois*rdt_insert - Total_chauff_SdB - Total_prod_thermique+ Total_ECS)*(1-Part_PAC)*alpha
Frac_recup_bois = 0.3
Total_th_PAC = (15200 - Total_conso_bois*rdt_insert - Total_chauff_SdB+ Total_ECS)*Part_PAC*alpha
sCOP = 1.8*2.5
P_elec_PAC = Total_th_PAC / sCOP
Total_conso_elec = 1100*nb_pers + Total_chauff_SdB + Total_pompe_piscine + P_elec_PAC
Total_conso_elec_reseau = Total_conso_elec*PV_frac_autoconso
Total_prod_PV = 5900*0
Total_revente_PV = (Total_prod_PV-Total_conso_elec*(1-PV_frac_autoconso))*0
Total_charge_batterie = Total_prod_PV *0.3*0
Total_decharge_batterie = Total_charge_batterie*0.85*0
Total_prod_thermique_piscine = 3800
Total_thermique = Total_conso_gaz*rdt_chaud + Total_prod_thermique + Frac_recup_bois*Total_conso_bois + Total_th_PAC
fig = go.Figure(go.Sankey(
arrangement = "snap",
textfont=dict(color="black", size=25),
node = {
"label": ["Non renouvelable", # 0
"ENR", # 1
"Nucléaire", # 2
"Gaz", # 3
"ENR Réseau", # 4
"Photovoltaïque", # 5
"Solaire thermique", # 6
"Air (PAC)", # 7
"Bois", # 8
"Electricité Réseau", # 9
"Pertes", # 10
"Electricité domestique", # 11
"Usage direct", # 12
"Batterie", # 13
"Revente", # 14
"Prod eau chaude", # 15
"Chauffage", # 16
"ECS", # 17
"Chauffage piscine", # 18
"PAC", # 19
"Autre", # 20
"Pompe Piscine", # 21
"Piscine" # 22
],
# "x": [0.2, 0.1, 0.5, 0.7, 0.3, 0.5],
# "y": [0.7, 0.5, 0.2, 0.4, 0.2, 0.3],
'pad':40}, # 10 Pixels
link = {
"source": [0, 0,
1, 1, 1, 1, 1,
2,
3,
4,
9,9,
5,
11,11,11,
14,
13,
3,
6,
7,
8,
15,15,
6,
12,
8,
12,
19,
12,
13,
3,
8,
12,
18,
21
],
"target": [2, 3,
4, 5, 6, 7, 8,
9,
9,
9,
12,10,
11,
12,13,14,
4,
12,
15,
15,
19,
15,
16,17,
18,
16,
16,
19,
15,
20,
10,
10,
10,
21,
22,
22
],
"value": [Total_conso_elec_reseau*Ratio_pertes*Frac_nucleaire, Total_conso_elec_reseau*Ratio_pertes*Frac_gaz+ Total_conso_gaz,
Total_conso_elec_reseau*Ratio_pertes*Frac_ENR_reseau, Total_prod_PV, Total_prod_thermique+Total_prod_thermique_piscine, Total_th_PAC*(1-1/sCOP), Total_conso_bois,
Total_conso_elec_reseau*Ratio_pertes*Frac_nucleaire,
Total_conso_elec_reseau*Ratio_pertes*Frac_gaz,
Total_conso_elec_reseau*Ratio_pertes*Frac_ENR_reseau, # + INJ
Total_conso_elec_reseau, Total_conso_elec_reseau*(Ratio_pertes-1),
Total_prod_PV,
Total_conso_elec*0-Total_decharge_batterie, Total_charge_batterie, Total_revente_PV, ###
Total_revente_PV,
Total_decharge_batterie,
Total_conso_gaz*rdt_chaud,
Total_prod_thermique,
Total_th_PAC*(1-1/sCOP),
Total_conso_bois*rdt_insert*Frac_recup_bois,
Total_thermique-Total_ECS,Total_ECS,
Total_prod_thermique_piscine,
Total_chauff_SdB,
Total_conso_bois*rdt_insert*(1-Frac_recup_bois),
Total_th_PAC/sCOP,
Total_th_PAC,
Total_conso_elec-Total_th_PAC/sCOP-Total_chauff_SdB-Total_pompe_piscine,
Total_charge_batterie-Total_decharge_batterie,
Total_conso_gaz*(1-rdt_chaud),
Total_conso_bois*(1-rdt_insert),
Total_pompe_piscine,
Total_prod_thermique_piscine,
Total_pompe_piscine
]
}))
sankey_plotly_rotex_piscine_builleur_pac = pn.pane.Plotly(fig)
scenarios['Combiné gaz condensation/solaire & Piscine & Bouilleur & PAC'] = sankey_plotly_rotex_piscine_builleur_pac
filename = '../_includes/sankey_plotly_rotex_piscine_builleur_pac.html'
export_plot_fastpages(pn.panel(sankey_plotly_rotex_piscine_builleur_pac), filename)
# -
# {% include sankey_plotly_rotex_piscine_builleur_pac.html %}
# # Rotex + piscine + bouilleur + PAC + PV
# +
#hide
import plotly.graph_objects as go
Ratio_pertes = 2.58
Frac_nucleaire=0.65
Frac_ENR_reseau=0.25
Frac_gaz=1-Frac_ENR_reseau-Frac_nucleaire
rdt_chaud=0.95
rdt_insert=0.90
PV_frac_autoconso = 0.3
nb_pers = 3
Total_conso_bois = 3500
Total_chauff_SdB = 200
Total_ECS = 600*nb_pers
Total_pompe_piscine = 6*5*31*0.750
alpha=0.77
Part_PAC=0.87
Total_prod_thermique = 3000
Total_conso_gaz = (15200 - Total_conso_bois*rdt_insert - Total_chauff_SdB - Total_prod_thermique+ Total_ECS)*(1-Part_PAC)*alpha
Frac_recup_bois = 0.3
Total_th_PAC = (15200 - Total_conso_bois*rdt_insert - Total_chauff_SdB+ Total_ECS)*Part_PAC*alpha
sCOP = 1.8*2.5
P_elec_PAC = Total_th_PAC / sCOP
Total_conso_elec = 1100*nb_pers + Total_chauff_SdB + Total_pompe_piscine + P_elec_PAC
Total_prod_PV = 5900
Total_revente_PV = (Total_prod_PV-Total_conso_elec*(1-PV_frac_autoconso))
Total_charge_batterie = Total_prod_PV *0.3
Total_decharge_batterie = Total_charge_batterie*0.85
Total_conso_elec_reseau = 2140 #Total_conso_elec*PV_frac_autoconso
Total_prod_thermique_piscine = 3800
Total_thermique = Total_conso_gaz*rdt_chaud + Total_prod_thermique + Frac_recup_bois*Total_conso_bois + Total_th_PAC
fig = go.Figure(go.Sankey(
arrangement = "snap",
textfont=dict(color="black", size=25),
node = {
"label": ["Non renouvelable", # 0
"ENR", # 1
"Nucléaire", # 2
"Gaz", # 3
"ENR Réseau", # 4
"Photovoltaïque", # 5
"Solaire thermique", # 6
"Air (PAC)", # 7
"Bois", # 8
"Electricité Réseau", # 9
"Pertes", # 10
"Electricité domestique", # 11
"Usage direct", # 12
"Batterie", # 13
"Revente", # 14
"Prod eau chaude", # 15
"Chauffage", # 16
"ECS", # 17
"Chauffage piscine", # 18
"PAC", # 19
"Autre", # 20
"Pompe piscine", # 21
"Piscine" # 22
],
# "x": [0.2, 0.1, 0.5, 0.7, 0.3, 0.5],
# "y": [0.7, 0.5, 0.2, 0.4, 0.2, 0.3],
'pad':20}, # 10 Pixels
link = {
"source": [0, 0,
1, 1, 1, 1, 1,
2,
3,
4,
9,9,
5,
11,11,11,
14,
13,
3,
6,
7,
8,
15,15,
6,
12,
8,
12,
19,
12,
13,
3,
8,
12,
18,
21
],
"target": [2, 3,
4, 5, 6, 7, 8,
9,
9,
9,
11,10,
11,
12,13,14,
4,
12,
15,
15,
19,
15,
16,17,
18,
16,
16,
19,
15,
20,
10,
10,
10,
21,
22,
22
],
"value": [Total_conso_elec_reseau*Ratio_pertes*Frac_nucleaire, Total_conso_elec_reseau*Ratio_pertes*Frac_gaz+ Total_conso_gaz,
Total_conso_elec_reseau*Ratio_pertes*Frac_ENR_reseau, Total_prod_PV, Total_prod_thermique+Total_prod_thermique_piscine, Total_th_PAC*(1-1/sCOP), Total_conso_bois,
Total_conso_elec_reseau*Ratio_pertes*Frac_nucleaire,
Total_conso_elec_reseau*Ratio_pertes*Frac_gaz,
Total_conso_elec_reseau*Ratio_pertes*Frac_ENR_reseau, # + INJ
Total_conso_elec_reseau, Total_conso_elec_reseau*(Ratio_pertes-1),
Total_prod_PV,
Total_conso_elec-Total_decharge_batterie, Total_charge_batterie, Total_revente_PV, ###
Total_revente_PV,
Total_decharge_batterie,
Total_conso_gaz*rdt_chaud,
Total_prod_thermique,
Total_th_PAC*(1-1/sCOP),
Total_conso_bois*rdt_insert*Frac_recup_bois,
Total_thermique-Total_ECS,Total_ECS,
Total_prod_thermique_piscine,
Total_chauff_SdB,
Total_conso_bois*rdt_insert*(1-Frac_recup_bois),
Total_th_PAC/sCOP,
Total_th_PAC,
Total_conso_elec-Total_th_PAC/sCOP-Total_chauff_SdB-Total_pompe_piscine,
Total_charge_batterie-Total_decharge_batterie,
Total_conso_gaz*(1-rdt_chaud),
Total_conso_bois*(1-rdt_insert),
Total_pompe_piscine,
Total_prod_thermique_piscine,
Total_pompe_piscine
]
}))
sankey_plotly_rotex_piscine_builleur_pac_pv = pn.pane.Plotly(fig)
scenarios['Combiné gaz condensation/solaire & Piscine & Bouilleur & PAC & PV' ] = sankey_plotly_rotex_piscine_builleur_pac_pv
filename = '../_includes/sankey_plotly_rotex_piscine_builleur_pac_pv.html'
export_plot_fastpages(pn.panel(sankey_plotly_rotex_piscine_builleur_pac_pv), filename)
# sankey_plotly_rotex_piscine_builleur_pac_pv
# -
# {% include sankey_plotly_rotex_piscine_builleur_pac_pv.html %}
# +
#hide
import plotly.graph_objects as go
Ratio_pertes = 1.52
Frac_nucleaire=0.0
Frac_ENR_reseau=1
Frac_gaz=1-Frac_ENR_reseau-Frac_nucleaire
rdt_chaud=0.95
rdt_insert=0.90
PV_frac_autoconso = 0.3
nb_pers = 3
Total_conso_bois = 3500
Total_chauff_SdB = 200
Total_ECS = 600*nb_pers
Total_pompe_piscine = 6*5*31*0.750
alpha=0.77
Part_PAC=0.87
Total_prod_thermique = 3000
Total_conso_gaz = (15200 - Total_conso_bois*rdt_insert - Total_chauff_SdB - Total_prod_thermique+ Total_ECS)*(1-Part_PAC)*alpha
Frac_recup_bois = 0.3
Total_th_PAC = (15200 - Total_conso_bois*rdt_insert - Total_chauff_SdB+ Total_ECS)*Part_PAC*alpha
sCOP = 1.8*2.5
P_elec_PAC = Total_th_PAC / sCOP
Total_conso_elec = 1100*nb_pers + Total_chauff_SdB + Total_pompe_piscine + P_elec_PAC
Total_prod_PV = 5900
Total_revente_PV = (Total_prod_PV-Total_conso_elec*(1-PV_frac_autoconso))
Total_charge_batterie = Total_prod_PV *0.3
Total_decharge_batterie = Total_charge_batterie*0.85
Total_conso_elec_reseau = 2140 #Total_conso_elec*PV_frac_autoconso
Total_prod_thermique_piscine = 3800
Total_thermique = Total_conso_gaz*rdt_chaud + Total_prod_thermique + Frac_recup_bois*Total_conso_bois + Total_th_PAC
fig = go.Figure(go.Sankey(
arrangement = "snap",
textfont=dict(color="black", size=25),
node = {
"label": ["Non renouvelable", # 0
"ENR", # 1
"Nucléaire", # 2
"BioGaz", # 3
"ENR Réseau", # 4
"Photovoltaïque", # 5
"Solaire thermique", # 6
"Air (PAC)", # 7
"Bois", # 8
"Electricité Réseau", # 9
"Pertes", # 10
"Electricité domestique", # 11
"Usage direct", # 12
"Batterie", # 13
"Revente", # 14
"Prod eau chaude", # 15
"Chauffage", # 16
"ECS", # 17
"Chauffage piscine", # 18
"PAC", # 19
"Autre", # 20
"Pompe piscine", # 21
"Piscine" # 22
],
# "x": [0.2, 0.1, 0.5, 0.7, 0.3, 0.5],
# "y": [0.7, 0.5, 0.2, 0.4, 0.2, 0.3],
'pad':20}, # 10 Pixels
link = {
"source": [0, 1,
1, 1, 1, 1, 1,
2,
3,
4,
9,9,
5,
11,11,11,
14,
13,
3,
6,
7,
8,
15,15,
6,
12,
8,
12,
19,
12,
13,
3,
8,
12,
18,
21
],
"target": [2, 3,
4, 5, 6, 7, 8,
9,
9,
9,
11,10,
11,
12,13,14,
4,
12,
15,
15,
19,
15,
16,17,
18,
16,
16,
19,
15,
20,
10,
10,
10,
21,
22,
22
],
"value": [Total_conso_elec_reseau*Ratio_pertes*Frac_nucleaire, Total_conso_elec_reseau*Ratio_pertes*Frac_gaz+ Total_conso_gaz,
Total_conso_elec_reseau*Ratio_pertes*Frac_ENR_reseau, Total_prod_PV, Total_prod_thermique+Total_prod_thermique_piscine, Total_th_PAC*(1-1/sCOP), Total_conso_bois,
Total_conso_elec_reseau*Ratio_pertes*Frac_nucleaire,
Total_conso_elec_reseau*Ratio_pertes*Frac_gaz,
Total_conso_elec_reseau*Ratio_pertes*Frac_ENR_reseau, # + INJ
Total_conso_elec_reseau, Total_conso_elec_reseau*(Ratio_pertes-1),
Total_prod_PV,
Total_conso_elec-Total_decharge_batterie, Total_charge_batterie, Total_revente_PV, ###
Total_revente_PV,
Total_decharge_batterie,
Total_conso_gaz*rdt_chaud,
Total_prod_thermique,
Total_th_PAC*(1-1/sCOP),
Total_conso_bois*rdt_insert*Frac_recup_bois,
Total_thermique-Total_ECS,Total_ECS,
Total_prod_thermique_piscine,
Total_chauff_SdB,
Total_conso_bois*rdt_insert*(1-Frac_recup_bois),
Total_th_PAC/sCOP,
Total_th_PAC,
Total_conso_elec-Total_th_PAC/sCOP-Total_chauff_SdB-Total_pompe_piscine,
Total_charge_batterie-Total_decharge_batterie,
Total_conso_gaz*(1-rdt_chaud),
Total_conso_bois*(1-rdt_insert),
Total_pompe_piscine,
Total_prod_thermique_piscine,
Total_pompe_piscine
]
}))
#fig.show()
#import plotly
#plotly.offline.plot(fig, validate=False)
# -
# # Comparaison
# +
#hide
# ['100% électricité']
select = pn.widgets.Select(name='Select', options=list(scenarios.keys()))
@pn.depends(select.param.value)
def callback(value):
return scenarios[value]
row = pn.Column(select, callback)
# options = {"embed_states":{select: list(range(0, 20, 1))}, "embed":True}
options = {"embed":True}
filename = '../_includes/sank_hvplot_test.html'
export_plot_fastpages_panel(row, filename, options=options)
# -
# {% include sank_hvplot_test.html %}
# # Références
# {{ "This is the actual footnote" | fndetail: 1 }}
# {{ "This is the actual footnote" | fndetail: 2 }}
# {{ "This is the actual footnote" | fndetail: 3 }}
| _notebooks/2021-04-19 - Sankey_story - DP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install pendulum
import pendulum
help(pendulum)
# +
from datetime import datetime
import pendulum
utc = pendulum.timezone('UTC')
pst = pendulum.timezone('America/Los_Angeles')
ist = pendulum.timezone('Asia/Calcutta')
print(type(utc))
print('Current Date Time in UTC =', datetime.now(utc))
print('Current Date Time in PST =', datetime.now(pst))
print('Current Date Time in IST =', datetime.now(ist))
print(type(datetime.now(ist)))
#Let’s see how to use pendulum module as a replacement of datetime module. However, if you are already using datetime module then it’s better to not mix them up.
utc_time = pendulum.now('UTC')
print(type(utc_time))
print('Current Date Time in UTC =', utc_time)
# -
# ### Converting Timezones
#
# +
utc_time = pendulum.now('UTC')
ist_time = utc_time.in_timezone('Asia/Calcutta')
print(type(ist_time))
print('Current Date Time in IST =', ist_time)
tz = pendulum.timezone('Europe/Paris')
paris_time = tz.convert(ist_time)
print('Current Date Time in Paris =', paris_time)
# -
# ### Date Time Manipulations
# We can use add() and subtract() functions for date time manipulations.
utc_time.add(years=1)
utc_time.subtract(months=2)
print('Updated UTC Time', utc_time)
# ### Date Time Formatting
# There are some useful methods to convert date time to standard formatted string. Pendulum module also has strftime() function where we can specify our own format.
# +
print(utc_time.to_iso8601_string())
print(utc_time.to_formatted_date_string())
print(utc_time.to_w3c_string())
print(utc_time.to_date_string())
# supports strftime() too
print(utc_time.strftime('%Y-%m-%d %H:%M:%S %Z%z'))
# -
# ### Parse String to Date Time
# We can use parse() function to parse a string having commonly used formats to datetime object. If you want to specify format string, then use from_format() function.
# +
dt = pendulum.parse('2018-05-21T22:00:00')
print(dt)
dt = pendulum.parse('2018-05-21T22:00:00', tz='Europe/Paris')
print(dt)
# parsing using specified format string
dt = pendulum.from_format('2018/05/21', 'YYYY/MM/DD')
print(dt)
# -
# ### Duration – timedelta replacement
# +
time_delta = pendulum.duration(days=1, hours=10, years=2)
print(time_delta)
print('time_delta years =', time_delta.years)
print('time_delta in seconds =', time_delta.in_seconds())
print('time_delta in words =', time_delta.in_words())
print('future date =', pendulum.now() + time_delta)
# -
# ### Period of Time
# +
current_date = pendulum.now()
future_date = current_date.add(days=4)
period_time = future_date - current_date
print('period in words =', period_time.in_words())
# period is iterable with days
for dt in period_time:
print(dt)
# +
import pendulum
dt = pendulum.datetime(2015, 2, 5)
isinstance(dt, datetime)
dt.timezone.name
# -
| Pendulum Module.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from performance_visualizer import load_search_performances, plot_search_performances
dataset = '185_baseball_MIN_METADATA'
file_path = 'resource/search_from_scratch.json'
search_performances = load_search_performances(file_path, 'Scratch')
search_results_scratch = pd.DataFrame.from_dict(search_performances)
search_results_scratch['time'] = pd.to_datetime(search_results_scratch['time'])
plot_search_performances(search_results_scratch, dataset)
file_path = 'resource/search_from_pretrained.json'
search_performances = load_search_performances(file_path, 'Pre-trained')
search_results_pretrained = pd.DataFrame.from_dict(search_performances)
search_results_pretrained['time'] = pd.to_datetime(search_results_pretrained['time'])
plot_search_performances(search_results_pretrained, dataset)
search_results_all = pd.concat([search_results_scratch, search_results_pretrained], ignore_index=True)
plot_search_performances(search_results_all, dataset)
| scripts/comparison_scratch_vs_pretrained.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Welcome to Transformer Reinforcement Learning (trl)
#
# > Train transformer language models with reinforcement learning.
# ## What is it?
# With `trl` you can train transformer language models with Proximal Policy Optimization (PPO). The library is built with the `transformer` library by 🤗 Hugging Face ([link](https://github.com/huggingface/transformers)). Therefore, pre-trained language models can be directly loaded via the transformer interface. At this point only GTP2 is implemented.
#
# **Highlights:**
# - GPT2 model with a value head: A transformer model with an additional scalar output for each token which can be used as a value function in reinforcement learning.
# - PPOTrainer: A PPO trainer for language models that just needs (query, response, reward) triplets to optimise the language model.
# - Example: Train GPT2 to generate positive movie reviews with a BERT sentiment classifier.
# ## How it works
# Fine-tuning a language model via PPO consists of roughly three steps:
#
# 1. **Rollout**: The language model generates a response or continuation based on query which could be the start of a sentence.
# 2. **Evaluation**: The query and response are evaluated with a function, model, human feedback or some combination of them. The important thing is that this process should yield a scalar value for each query/response pair.
# 3. **Optimization**: This is the most complex part. In the optimisation step the query/response pairs are used to calculate the log-probabilities of the tokens in the sequences. This is done with the model that is trained and and a reference model, which is usually the pre-trained model before fine-tuning. The KL-divergence between the two outputs is used as an additional reward signal to make sure the generated responses don't deviate to far from the reference language model. The active language model is then trained with PPO.
#
# This process is illustrated in the sketch below:
#
#
# <div style="text-align: center">
# <img src='images/trl_overview.png' width='800'>
# <p style="text-align: center;"> <b>Figure:</b> Sketch of the workflow. </p>
# </div>
# ## Installation
# ### Python package
# Install the library with pip:
#
# `pip install trl`
#
# ### Repository
# If you want to run the examples in the repository a few additional libraries are required. Clone the repository and install it with pip:
#
# `git clone https://github.com/lvwerra/trl.git`
#
# `cd tlr/`
#
# `pip install -r requirements.txt`
#
# ### Jupyter notebooks
#
# If you run Jupyter notebooks you might need to run the following:
#
# `jupyter nbextension enable --py --sys-prefix widgetsnbextension`
#
# For Jupyterlab additionally this command:
#
# `jupyter labextension install @jupyter-widgets/jupyterlab-manager`
# ## How to use
# ### Example
# This is a basic example on how to use the library. Based on a query the language model creates a response which is then evaluated. The evaluation could be a human in the loop or another model's output.
# +
# imports
import torch
from transformers import GPT2Tokenizer
from trl.gpt2 import GPT2HeadWithValueModel, respond_to_batch
from trl.ppo import PPOTrainer
# get models
gpt2_model = GPT2HeadWithValueModel.from_pretrained('gpt2')
gpt2_model_ref = GPT2HeadWithValueModel.from_pretrained('gpt2')
gpt2_tokenizer = GPT2Tokenizer.from_pretrained('gpt2')
# initialize trainer
ppo_config = {'batch_size': 1, 'forward_batch_size': 1}
ppo_trainer = PPOTrainer(gpt2_model, gpt2_model_ref, **ppo_config)
# encode a query
query_txt = "This morning I went to the "
query_tensor = gpt2_tokenizer.encode(query_txt, return_tensors="pt")
# get model response
response_tensor = respond_to_batch(gpt2_model, query_tensor)
response_txt = gpt2_tokenizer.decode(response_tensor[0,:])
# define a reward for response
# (this could be any reward such as human feedback or output from another model)
reward = torch.tensor([1.0])
# train model with ppo
train_stats = ppo_trainer.step(query_tensor, response_tensor, reward)
# -
# ### Advanced example: IMDB sentiment
# For a detailed example check out the notebook *Tune GPT2 to generate positive reviews*, where GPT2 is fine-tuned to generate positive movie reviews. An few examples from the language models before and after optimisation are given below:
#
# <div style="text-align: center">
# <img src='images/table_imdb_preview.png' width='800'>
# <p style="text-align: center;"> <b>Figure:</b> A few review continuations before and after optimisation. </p>
# </div>
#
# ## Notebooks
# This library is built with `nbdev` and as such all the library code as well as examples are in Jupyter notebooks. The following list gives an overview:
#
# - `index.ipynb`: Generates the README and the overview page.
# - `00-core.ipynb`: Contains the utility functions used throughout the library and examples.
# - `01-gpt2-with-value-head.ipynb`: Implementation of a `transformer` compatible GPT2 model with an additional value head as well as a function to generate sequences.
# - `02-ppo.ipynb`: Implementation of the PPOTrainer used to train language models.
# - `03-bert-imdb-training.ipynb`: Training of BERT with `simpletransformers` to classify sentiment on the IMDB dataset.
# - `04-gpt2-sentiment-ppo-training.ipynb`: Fine-tune GPT2 with the BERT sentiment classifier to produce positive movie reviews.
#
# ## References
#
# ### Proximal Policy Optimisation
# The PPO implementation largely follows the structure introduced in the paper **"Fine-Tuning Language Models from Human Preferences"** by <NAME> et al. \[[paper](https://arxiv.org/pdf/1909.08593.pdf), [code](https://github.com/openai/lm-human-preferences)].
#
# ### Language models
# The language models utilize the `transformer` library by 🤗Hugging Face.
| nbs/index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# a collection of instructions
# a collection of code
def function1():
print("ahhhh")
print("ahhhhh 2")
print("this is outside the function")
function1()
function1()
function1()
# a mapping
# input or an argument
def function2(x):
return 2*x
a = function2(3)
# return value or output
print(a)
b = function2(4)
print(b)
c = function2(5)
print(c)
d = function2()
def function3(x, y):
return x + y
e = function3(1, 2)
print(e)
def function4(x):
print(x)
print("still in this function")
return 3*x
f = function4(4)
print(f)
def function5(some_argument):
print(some_argument)
print("weeee")
function5(4)
# +
# BMI calculator
name1 = "YK"
height_m1 = 2
weight_kg1 = 90
name2 = "<NAME>"
height_m2 = 1.8
weight_kg2 = 70
name3 = "<NAME>"
height_m3 = 2.5
weight_kg3 = 160
# -
def bmi_calculator(name, height_m, weight_kg):
bmi = weight_kg / (height_m ** 2)
print("bmi: ")
print(bmi)
if bmi < 25:
return name + " is not overweight"
else:
return name + " is overweight"
result1 = bmi_calculator(name1, height_m1, weight_kg1)
result2 = bmi_calculator(name2, height_m2, weight_kg2)
result3 = bmi_calculator(name3, height_m3, weight_kg3)
print(result1)
print(result2)
print(result3)
# Solution to the task:
# The following function converts miles to kilometers.
# km = 1.6 * miles
def convert(miles):
return 1.6 * miles
print(convert(1))
print(convert(2))
# +
# csdojo.io/python3
# csdojo.io/news - 2-4
# YK from CS Dojo
| Python X/.ipynb_checkpoints/What+are+functions_-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # [SC57 - Working with big, multi-dimensional geoscientific datasets in Python: a tutorial introduction to xarray](http://meetingorganizer.copernicus.org/EGU2017/session/25651)
#
#
# Original notebook by [<NAME>](http://stephanhoyer.com), Rossbypalooza, 2016.
# Modified by <NAME>, <NAME> and [<NAME>](http://fabienmaussion.info/) for EGU General Assembly 2017, Vienna, Austria
# Modified by <NAME> for GHRSST Science Team Tutorial 2019, Rome, Italy
#
# Sunday, 31 May 2019, 9:00 - 2:00 Hotel Flora
#
#
# **Convenors**
# * [Dr <NAME>](mailto:<EMAIL>) - Earth and Space Research, USA
# * [Dr <NAME>](mailto:<EMAIL>) - Farallon Institute, USA
# -------------
# 
#
# # Structure of this tutorial
#
# 1. Opening data
# 1. Collocating satellite data with a cruise dataset
#
#
# # 1. Key features of `xarray`
# -------------------
#
# ## Import python packages
#
# You are going to want numpy, pandas, matplotlib.pyplot and xarray
# +
import warnings
warnings.simplefilter('ignore') # filter some warning messages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import xarray as xr
#for search capabilites import podaacpy
import podaac.podaac as podaac
# then create an instance of the Podaac class
p = podaac.Podaac()
# -
# ## A nice cartopy tutorial is [here](http://earthpy.org/tag/visualization.html)
# # Collocate a Saildrone cruise with AVHRR SST data
# ### Let's open 2 months of 0.2 km AVHRR OI SST data
# `xarray`can open multiple files at once using string pattern matching.
#
# In this case we open all the files that match our `filestr`, i.e. all the files for the 2080s.
#
# Each of these files (compressed) is approximately 800 MB.
from glob import glob
filepath = './data/avhrr_oi/*.nc'
files = glob(filepath,recursive=True)
files[:10]
# # Let's see what one day looks like
ds_sst = xr.open_dataset(files[0])
f = plt.figure(figsize=(8, 4))
ax = plt.axes(projection=ccrs.Orthographic(-80, 35))
ds_sst.sst[0,:,:].plot(ax=ax, transform=ccrs.PlateCarree())
ax.coastlines(); ax.gridlines();
# # Change the colormap, colorscale, and add land
f = plt.figure(figsize=(8, 4))
ax = plt.axes(projection=ccrs.Orthographic(-80, 35))
ds_sst.sst[0,:,:].plot(ax=ax, transform=ccrs.PlateCarree(),cmap='jet',vmin=-1,vmax=34)
ax.coastlines(); ax.gridlines();
ax.stock_img();
#fig_fname='./images/sst_avhrroi.png'
#f.savefig(fig_fname, transparent=False, format='png')
# ## Now open multiple files (lazy) using [.open_mfdataset](http://xarray.pydata.org/en/stable/generated/xarray.open_mfdataset.html#xarray.open_mfdataset)
#
# * use the option `coords = 'minimal'`
ds_sst = xr.open_mfdataset(files,coords='minimal')
ds_sst
# ## Again with the 0-360 vs -180-180. Change it up below!
# * Also, look at the coordinates, there is an extra one `zlev`. Drop it using .isel
#
#
ds_sst.coords['lon'] = np.mod(ds_sst.coords['lon'] + 180,360) - 180
ds_sst = ds_sst.sortby(ds_sst.lon)
ds_sst = ds_sst.isel(zlev=0)
ds_sst
# `xarray` even puts them in the right order for you.
ds_sst.time
# How big is all this data uncompressed? Will it fit into memory?
# Use `.nbytes` / 1e9 to convert it into gigabytes
ds_sst.nbytes / 1e9
# # Collocating Saildrone cruise data with MUR SSTs
#
# * read in the Saildrone data
url = 'https://podaac-opendap.jpl.nasa.gov/opendap/hyrax/allData/insitu/L2/saildrone/Baja/saildrone-gen_4-baja_2018-sd1002-20180411T180000-20180611T055959-1_minutes-v1.nc'
ds_usv = xr.open_dataset(url)
ds_usv
# ## The NCEI trajectory format uses 'obs' as the coordinate. This is an example of an 'older' style of data formatting that doesn't really mesh well with modern software capabilities.
#
# * So, let's change that by using [.swap_dims](http://xarray.pydata.org/en/stable/generated/xarray.DataArray.swap_dims.html) to change the coordinate from `obs` to `time`
# * Another thing, `latitude` and `longitude` are just long and annoying, lets [.rename](http://xarray.pydata.org/en/stable/generated/xarray.Dataset.rename.html) them to `lat` and `lon`
#
# * Finally, the first and last part of the cruise the USV is being towed, so let's only include data from `2018-04-12T02` to `2018-06-10T18`
#
ds_usv = ds_usv.isel(trajectory=0).swap_dims({'obs':'time'}).rename({'longitude':'lon','latitude':'lat'})
ds_usv_subset = ds_usv.sel(time=slice('2018-04-12T02','2018-06-10T18'))
# # Xarray interpolation won't run on chunked dimensions.
# 1. First let's subset the data to make it smaller to deal with by using the cruise lat/lons
# * Find the max/min of the lat/lon using `.lon.min().data`
#
# 1. Now load the data into memory (de-Dask-ify) it using `.load()`
#
#Step 1 from above
print('min max lat lon:', ds_usv_subset.lon.min().data,ds_usv_subset.lon.max().data,ds_usv_subset.lat.min().data,ds_usv_subset.lat.max().data)
subset = ds_sst.sel(lon=slice(ds_usv_subset.lon.min().data,ds_usv_subset.lon.max().data),
lat=slice(ds_usv_subset.lat.min().data,ds_usv_subset.lat.max().data))
#Step 2 from above
subset.load()
# # Collocate USV data with MUR data
# There are different options when you interpolate. First, let's just do a linear interpolation using [.interp()](http://xarray.pydata.org/en/stable/generated/xarray.Dataset.interp.html#xarray.Dataset.interp)
#
# `Dataset.interp(coords=None, method='linear', assume_sorted=False, kwargs={}, **coords_kwargs))`
#
#
ds_collocated = subset.interp(lat=ds_usv_subset.lat,lon=ds_usv_subset.lon,time=ds_usv_subset.time,method='linear')
# # Collocate USV data with MUR data
# There are different options when you interpolate. First, let's just do a nearest point rather than interpolate the data
# `method = 'nearest'`
ds_collocated_nearest = subset.interp(lat=ds_usv_subset.lat,lon=ds_usv_subset.lon,time=ds_usv_subset.time,method='nearest')
# ## Now, calculate the different in SSTs and print the [.mean()](http://xarray.pydata.org/en/stable/generated/xarray.DataArray.mean.html#xarray.DataArray.mean) and [.std()](http://xarray.pydata.org/en/stable/generated/xarray.DataArray.std.html#xarray.DataArray.std)
# For the satellite data we need to use `sst` and for the USV data we need to use `TEMP_CTD_MEAN`
dif = ds_collocated_nearest.sst-ds_usv.TEMP_CTD_MEAN
print('mean difference = ',dif.mean().data)
print('STD = ',dif.std().data)
# # xarray can do more!
#
# * concatentaion
# * open network located files with openDAP
# * import and export Pandas DataFrames
# * .nc dump to
# * groupby_bins
# * resampling and reduction
#
# For more details, read this blog post: http://continuum.io/blog/xray-dask
#
# +
#ds_collocated_nearest.to_netcdf('./data/new file.nc')
# -
# ## Where can I find more info?
#
# ### For more information about xarray
#
# - Read the [online documentation](http://xarray.pydata.org/)
# - Ask questions on [StackOverflow](http://stackoverflow.com/questions/tagged/python-xarray)
# - View the source code and file bug reports on [GitHub](http://github.com/pydata/xarray/)
#
# ### For more doing data analysis with Python:
#
# - <NAME>, [A modern guide to getting started with Data Science and Python](http://twiecki.github.io/blog/2014/11/18/python-for-data-science/)
# - <NAME>, [Python for Data Analysis](http://shop.oreilly.com/product/0636920023784.do) (book)
#
# ### Packages building on xarray for the geophysical sciences
#
# For analyzing GCM output:
#
# - [xgcm](https://github.com/xgcm/xgcm) by <NAME>
# - [oogcm](https://github.com/lesommer/oocgcm) by <NAME>
# - [MPAS xarray](https://github.com/pwolfram/mpas_xarray) by <NAME>
# - [marc_analysis](https://github.com/darothen/marc_analysis) by <NAME>
#
# Other tools:
#
# - [windspharm](https://github.com/ajdawson/windspharm): wind spherical harmonics by <NAME>
# - [eofs](https://github.com/ajdawson/eofs): empirical orthogonal functions by <NAME>
# - [infinite-diff](https://github.com/spencerahill/infinite-diff) by <NAME>
# - [aospy](https://github.com/spencerahill/aospy) by <NAME> and <NAME>
# - [regionmask](https://github.com/mathause/regionmask) by <NAME>
# - [salem](https://github.com/fmaussion/salem) by <NAME>
#
# Resources for teaching and learning xarray in geosciences:
# - [Fabien's teaching repo](https://github.com/fmaussion/teaching): courses that combine teaching climatology and xarray
#
| ocean_python_tutorial/Intro_08_Xarray-Advanced_cruise_data_case_study.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia (4 threads) 1.7.2
# language: julia
# name: julia-(4-threads)-1.7
# ---
# # Multithreading
# By default Julia only using a single thread. To start it with multiple threads we must tell it explicitly:
#
# ##### Command line argument
#
# ```bash
# julia -t 4
# ```
#
# or
#
# ```bash
# julia --threads 4
# ```
#
# ##### Environmental variable
#
# On Linux/MacOS:
#
# ```bash
# export JULIA_NUM_THREADS=4
# ```
#
# On Windows:
#
# ```bash
# set JULIA_NUM_THREADS=4
# ```
#
# Afterwards start julia *in the same terminal*.
#
# ##### Jupyter kernel
#
# You can also create a *Jupyter kernel* for multithreaded Julia:
#
# ```julia
# using IJulia
# installkernel("Julia (4 threads)", "--project=@.", env=Dict("JULIA_NUM_THREADS"=>"4"))
# ```
#
# *Note:* This has to be manually redone for every new Julia version and you have to restart your Jupyter process to see an effect.
# To check this has worked we we use:
Threads.nthreads()
# ## `Threads.@spawn`
# The `Threads.@spawn` macro dynamically spawns a new thread to execute a command in the background. Programmatically, it creates a `Task` and puts it on the todo-list. Whenever a thread is free, the task is dynamically assigned to a thread and executing the work starts.
Threads.@spawn println("test")
# **Important:** `Threads.@spawn` returns the created task *immediately*, but we might have to wait until the task is done and fetch the result later:
t = Threads.@spawn begin
sleep(3)
4
end
# We immediately get here
@time fetch(t) # This waits until the task is done
# To prevent the immediate return, we need to explicitly synchronise the execution using an `@sync` macro barrier.
# For example:
# +
@sync begin
t = Threads.@spawn begin
sleep(3)
4
end
end
@time fetch(t) # No need to wait, the task is already done
# -
# ## Filling an array in parallel
#
# Now, let's use this to actually parallelise something: We will fill an array in parallel:
# +
function fill_array_parallel(a)
@sync for i in 1:length(a)
Threads.@spawn a[i] = Threads.threadid()
end
a
end
a = zeros(Threads.nthreads()*10);
fill_array_parallel(a)
# -
@show count(a .== 1.0)
@show count(a .== 2.0)
@show count(a .== 3.0)
@show count(a .== 4.0)
# Note: Due to the **dynamic scheduling** some threads actually do more work (more values of i) than others!
# ## Nesting threading
#
# A key motion in the Julia ecosystem is to support **nested threading**:
function threaded_fun()
x = Threads.threadid()
Threads.@spawn println("job1", " (spawned from $x, processed by $(Threads.threadid()))")
Threads.@spawn println("job2", " (spawned from $x, processed by $(Threads.threadid()))")
Threads.@spawn println("job3", " (spawned from $x, processed by $(Threads.threadid()))")
end
@sync for i in 1:Threads.nthreads()
Threads.@spawn threaded_fun()
end
# The key point about this is that in this way the threading of different layers of functions does not interfer by causing more threads to be spawned than there are workers (CPU cores).
#
# The issue happens rather easily whenever a parallelised routine like `threaded_fun` (e.g. a numerical integration routine) is again called from a parallelised outer loop (e.g. a solver). To avoid the problem one needs to introduce some kind of coupling between the routines to communicate to the inner routine (`threaded_fun`) how many threads it may use. To avoid the need to do this explicitly, Julia implemented has decided to base its threading mostly on dynamic scheduling and the `@spawn` formalism.
# ## Threading takes extra care: Parallel summation
#
# We consider the case of a parallel summation
function mysum(xs)
s = zero(eltype(xs))
for x in xs
s += x
end
s
end
function mysum_parallel_naive(xs)
s = zero(eltype(xs))
@sync for x in xs
Threads.@spawn (s += x)
end
s
end
xs = rand(100_000);
@show sum(xs);
@show mysum(xs);
@show mysum_parallel_naive(xs);
# Hmmm ... the problem is a so-called **race condition**, a clash due to the parallel writing access from multiple threads.
#
# One way to solve this is by using [Atomic Operations](https://docs.julialang.org/en/v1/manual/multi-threading/#Atomic-Operations):
# +
import Base.Threads: Atomic, atomic_add!
function mysum_parallel_atomics(xs)
T = eltype(xs)
s = Atomic{T}(zero(T))
@sync for x in xs
Threads.@spawn atomic_add!(s, x)
end
s[]
end
# -
@show mysum(xs);
@show mysum_parallel_atomics(xs);
@btime mysum($xs);
@btime mysum_parallel_atomics($xs);
@btime mysum_parallel_naive($xs);
# **Note:** Atomics are generally bad. Don't use this paradigm in production unless you know what you are doing. Use FLoops.jl (see below).
# ## Is there no static scheduling option in Julia?
#
# Yes there is and it can sometimes be faster than dynamic threading:
function mysum_parallel_threads(xs)
T = eltype(xs)
s = Atomic{T}(zero(T))
Threads.@threads :static for x in xs
atomic_add!(s, x)
end
s[]
end
@btime mysum_parallel_atomics($xs);
@btime mysum_parallel_threads($xs);
# While on a first look this has advantages in form of a 10-fold reduced speed, the disadvantages are that there is no nested threading and there can be severe load imbalancing since work is split statically at startup of the loop.
# ## FLoops.jl: Easy and fast dynamic threads
#
# As a way out the Julia ecosystem has brought forward a number of carefully optimised packages for threaded execution based on *dynamic* scheduling. One example is [FLoops.jl](https://github.com/JuliaFolds/FLoops.jl). Our `mysum` function is parallelised using FLoops by just adding two macros:
# +
using FLoops
function mysum_parallel_floops(xs)
s = zero(eltype(xs))
@floop for x in xs
@reduce s += x
end
s
end
# -
# Still it gives the right result and is faster than our statically scheduled `@threads` version:
@show mysum(xs);
@show mysum_parallel_floops(xs);
@btime mysum_parallel_threads(xs);
@btime mysum_parallel_floops($xs);
# **Note:** The fact that `FLoops` is faster is a little misleading at first sight, but illustrates an important point nevertheless:
#
# - If *perfectly* written *statically scheduled* threads are faster than dynamically scheduled threads
# - But this requires deep insight to obtain optimal load balancing, careful use of atomics etc.
# - If you are not a parallelisation expert carefully optimised packages based on *dynamical scheduling* will likely be faster for your use case. The plain reason is that the *learning time* to understand all the neccessary tricks and the time needed to *fix all the subtle bugs* is not to be underestimated.
# ### Takeaways
# - Julia's thread infrastructure is mostly based on *dynamic* threading
# - The advantages are thread nesting and better load balancing in cases where load per iteration is not uniform.
# - The disadvantage is a larger startup time per thread
# - Packages like FLoops.jl make it easy to write fast parallel code.
# ##### More details
# - https://juliafolds.github.io/data-parallelism/
| 08_Multithreading_Basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="W1AytsknY8Nb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c137940e-64e8-4f8c-cbc8-7bca26a4f9d5"
print("LetsUpgrade - Day 2 Assignment of python ");
# + [markdown] id="LncIJMD_5SSb" colab_type="text"
# # ***LIST***
# + id="OUeSuBvpbFvd" colab_type="code" colab={}
# To Create a List
lst = ["Sema", "Angle", 75439, 74569.53, [7, 5, 9, 2]]
# + id="p2drbS0wcnN3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7df39dd5-97c1-4a22-d446-4e6c516ab4aa"
# To Display List
lst
# + id="3BQ0V8BDc4ps" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="f819c7e7-d02a-49c8-d6b9-ebde8a29dcb0"
lst [0]
# + id="_k7elqaydi2N" colab_type="code" colab={}
# Append is used to add a subscript in a List a Last
lst.append("Komal")
# + id="-8JvgmbgeKIr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f42e8a4a-b38b-4d52-d38c-bd2ead456b5b"
# Subscript is added in the List at Last
lst
# + id="Z5VA4ZkAeNFp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9d6af47e-f21d-420a-d405-d59f6df3e67a"
# Used to show the Subscript of the index
lst [4]
# + id="H9GV90WgeVxj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0d70605c-2612-4ba3-fbec-d68a9439f80c"
lst [4][2]
# + id="BXKADsYlefrg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="189f3624-0b96-44aa-c464-e8de911b81f2"
# Pop is used to delete the Subscript from the List
lst.pop(4)
# + id="TdMeKgnnerFH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="fb789410-6628-4555-c826-19e2ae67d448"
# List is displayed
lst
# + id="4_Co-JsSf3rk" colab_type="code" colab={}
# reverse is used to reverse the List
lst.reverse()
# + id="pGKgeWBKf6-o" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="bbe48441-b40a-48a3-94d8-61f252ae3884"
# reversed List is displayed
lst
# + id="dXWqW-vDgAmK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1dcb30d8-dea6-4504-c5d6-4b817fcc2232"
# count is used to count the number of time Subscript have same element
lst.count(75439)
# + id="3-_kY09QgIwk" colab_type="code" colab={}
# Second List is created
lst2 = ["Riya", 345]
# + id="8GaTj-ZKyiaj" colab_type="code" colab={}
# extend is use to modify the original list
lst2.extend(lst)
# + id="VvygJtN1zijp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4828bbb7-6dff-41f0-9dae-120f4f2c29e2"
# due to extend function lst Subscript are added to lst2 Subscripts
lst2
# + [markdown] id="H4G_khf95Ctk" colab_type="text"
#
# # ***DICTIONARY***
# ---
#
#
# + id="5u8HMTH91jl0" colab_type="code" colab={}
# dit is used to create a dictionary
dit = {"Name":"<NAME>", "Age":"22", "Mobile No.":7291054963, "Email ID":"<EMAIL>"}
# + id="cXfkK8qZ25zQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="da4580ad-1f95-44bc-c547-52330cb1a3a2"
# dit is created
dit
# + id="w75ecOBM27Fo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="1a0eed13-37c8-4c00-97a6-5ce28956cb56"
# setdefault add new subscript in dit
dit.setdefault("Course","B.Tech")
# + id="AYy_N00i6bGM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="da707261-1912-422d-c243-e978f7fe7a43"
# dit is displayed
dit
# + id="0RAXPH8V9tnh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="f9512697-ffe9-4bb3-bbcc-03d6b40f1a01"
# get will help to show the Name
dit.get("Name")
# + id="Df_0Ax41-Kta" colab_type="code" colab={}
# dit2 is created and copy function will copy dit in dit2
dit2=dit.copy()
# + id="e2dNNX2g_T1m" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="16f445e6-648a-4022-efc3-6bcc204ed2d1"
# dit2 is displayed
dit2
# + id="r5cU1ppP_nw5" colab_type="code" colab={}
# clear function will make dit2 empty
dit2.clear()
# + id="Nl2tLWgX_70S" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1b73cc57-4b16-4a18-8d5c-c2a687642ed9"
# dit2 is empty
dit2
# + id="sv2OTtNz_-RC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9ee6a8b8-2b0d-496c-80a7-36a35cbb2c39"
# fromkeys() is used to specify keys and its value
x=('maths','english','hindi','computer')
y=('100')
dit.fromkeys(x,y)
# + [markdown] id="PSWturY7Cu0t" colab_type="text"
# ## ***SETS***
# + id="uYLRXQ_uC6LW" colab_type="code" colab={}
# Two Sets are created st and st1
st = {"SAM", "c+=", 10, 10, 20, 10, 50, 30, 50, 70}
st1 = {"REHA", "C+", 10, 15, 20, 10, 50, 25, 40, 70}
# + id="Ea4ykTXRDx0G" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7298e4e9-7592-48f0-dd48-578943b68dc9"
# st set is displyed
st
# + id="4ZFQpgf3D03T" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0b411246-179e-4035-bc20-b3e55f795031"
# st1 set is displyed
st1
# + id="Tc9x57NhD6Fk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="82dedb85-b1db-4424-ed3d-b52c3dabe035"
# difference is used to find all the difference of st1 from st
st1.difference(st)
# + id="7tThwXDRFQ-6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="52a1dc62-97e3-47bf-ecc2-066fac312922"
# issuperset is used to see that is st1 is the super set of st
st.issuperset(st1)
# + id="zRaHwBvfFZty" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="205bdc05-0afb-4f7c-d796-2f671a2cce53"
# issubset is used to see that is st1 is the sub set of st1(itself)
st1.issubset(st1)
# + id="uTxJKD_iFdoq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="96128db2-a7e8-435d-e6ee-de18b3ded1f6"
# intersection is used to see that is st1 is the intersection of st
st1.intersection(st)
# + id="0KPkb4EZFklD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d40e7824-ef2e-4868-e06e-589a530d5345"
# isdisjoint is used to see that is st1 is the isdisjoint of st
st1.isdisjoint(st)
# + [markdown] id="O5duv2M_Il3B" colab_type="text"
# ## ***TUPLE***
# + id="jMqRjgW0IsY9" colab_type="code" colab={}
# tup is used to creat Tpule
tup = ["Raja", "Ram", "Mohan", "Roy"]
# + id="MWrewhlxJT0K" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ce882c4f-c2f4-4320-9fd8-7e325cd19510"
# Tuple is created
tup
# + id="CqQgiei7JVZO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="505d580a-0f7f-4859-968e-0ec26a9f5232"
# index of roy is shown
tup.index("Roy")
# + id="UJr84NGzJeLL" colab_type="code" colab={}
# new subscript Raja is added in the Tuple
tup.insert(4,'Raja')
# + id="u4ivMEaJKPWu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0435d655-b06c-4e98-ac96-20337797f3e6"
# updated tuple is displayed
tup
# + id="ZWYqu08DK78d" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="88a94cd4-beaf-4372-e738-94307f0a1ad5"
# count is used to count how many time raja is there in the Tuple
tup.count("Raja")
# + id="g5ehqjfkLCec" colab_type="code" colab={}
# Tuple is sorted in alphabetical order
tup.sort()
# + id="0h6PivSVLNWe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c821dd6d-1b26-421d-b289-dddcdfb3faca"
# sorted Tuple is displayed
tup
# + id="xVK9TJL_LQEj" colab_type="code" colab={}
# remove is used to remove Raja from Tuple
tup.remove("Raja")
# + id="ugGqU4y-LZ39" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="33b6391e-9c3d-4f28-d64a-e4e396b31589"
# updated Tuple is displayed
tup
# + [markdown] id="jVnZszbGNHRn" colab_type="text"
# ## ***STRING***
# + id="sZg4Qn4sNMH3" colab_type="code" colab={}
# two string is created name and name1
name = "<NAME>"
name1 = "<NAME>"
# + id="8f-4y0LsOKOE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="24ec54ec-fa72-417c-8dd2-81c23a0e0c45"
# name string is displayed
name
# + id="zquMqqzoOMQl" colab_type="code" colab={}
# new string is created with the name newname which is the addition of name and name1
newname = name + ' ' + name1
# + id="9Q3n5kR_QCmE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="781b023f-5962-4bdc-b1e5-02d9c227cc73"
# newname string is displayed
newname
# + id="Yr5KPiUmOWep" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="2d7f98c8-5dcf-44ab-b724-6d5248bc9215"
# type() is used for knowing the data type
type(newname)
# + id="C9jcVuSGPANC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="bd9a71ae-ca74-43ac-a23f-44d2106da878"
# endswith() is used for knowing that from what string is ending with
newname.endswith(".")
# + id="YZvblfXOPRyI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="433f0546-2768-4bd6-d231-a6ec045a2e33"
# expandtabs() is used to specified number of whitespaces.
newname.expandtabs(2)
# + id="aynH_9k0Qc2U" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="56a73364-f409-4e80-8487-2dec689c8934"
# isspace() is used for string handling
newname.isspace()
# + id="JR04THAAQ14i" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="11e9c1cd-053f-4114-ab87-f2f6020e2251"
# formate() is used for formatted string
newname.format()
# + id="VuCW1FwaR4X0" colab_type="code" colab={}
| Python_Day_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="Y52zwvlpHvCC"
# ## deblurring
# repairing images motion blur
# - channel attention mechanism [[paper]](https://arxiv.org/abs/1807.02758)
# - residual in residual architecture [[paper]](https://arxiv.org/abs/1505.04597)
# - subpixel convolution / pixelshuffle [[paper]](https://arxiv.org/abs/1609.05158)
# - running on [tensorflow/google colab](https://colab.research.google.com/) AND on [plaidml](https://www.intel.ai/plaidml/)
# - using the famous [Set14](https://www.google.com/search?q=set14) dataset ONLY (with heavy augmentation) - no validation needed
#
# jupyter notebook by [<NAME>](https://scholar.google.de/citations?user=yEn9St8AAAAJ) from [github](https://www.github.com/BenjaminWegener/keras-examples)
# + [markdown] colab_type="text" id="s-U33BDYHvCF"
# ### options
#
# + colab_type="code" id="bDQFzxuJHvCH" colab={}
run_on_google_colab = True #use PlaidML as Backend, change this to 'True' to run on colab/tf
epochs = 250 #Number of epochs to train
channels = 3 #channels of low resolution image
batch_size = 14 #what batch-size should we use (decrease if you encounter video memory errors)
steps_per_epoch = 1000 #How much iterations per epoch to train
height_lr = 256 #height of low resolution image (must be dividable by 4)
width_lr = height_lr #width of low resolution image (must be dividable by 4)
gen_lr = 0.001 #learning rate of generator
logging_steps = 50 #how often to update the training log
rotation_max = 0.33 # max max degree of rotation
max_move = height_lr / 100 #how much pixels to shift the blurred image
blur_copies = 5 # how many copies of image to generate and merge back together
# + [markdown] colab_type="text" id="GzyzlkF3HvCL"
# ### imports
# + colab_type="code" id="v78MsA4CHvCM" outputId="55545f87-9a09-450a-a202-1bb3c5e70fbb" colab={"base_uri": "https://localhost:8080/", "height": 67}
import os
if run_on_google_colab:
# %cd /content
# !git clone https://github.com/BenjaminWegener/keras-examples #download Dataset
# %cd keras-examples
else:
os.environ['KERAS_BACKEND'] = 'plaidml.keras.backend'
import numpy as np
from keras.models import Model, Input, load_model
from keras.layers import *
from keras.optimizers import Adam
from keras import backend as K
from keras.callbacks import LambdaCallback
from IPython.display import clear_output
import matplotlib.pyplot as plt
from keras.preprocessing.image import ImageDataGenerator
import random
from scipy.ndimage import rotate, shift
# %matplotlib inline
# + [markdown] colab_type="text" id="QsOYTVq4KpRZ"
# ### function for image visualization
# + colab_type="code" id="Ygs-7DxwHvCS" colab={}
def show(tensors):
plt.rcParams['figure.figsize'] = [20, 10]
fig = plt.figure()
for i in range(len(tensors)):
try:
tensors[i] = np.squeeze(tensors[i], axis = 0)
except:
pass
tensors[i] = (tensors[i] + 1.) * 127.5
fig.add_subplot(1,len(tensors), i + 1)
plt.imshow(tensors[i].astype(np.uint8), interpolation = 'nearest')
plt.setp(plt.gcf().get_axes(), xticks=[], yticks=[]);
plt.show()
# + [markdown] colab_type="text" id="SJi-rBgoHvCU"
# ### dataset function
# + colab_type="code" id="m5lxEgg6HvCV" colab={}
# return batch of augmented train and target images with quantity n_samples
def get_batch(n_samples, height, width, channels):
# define a ImageGenerator instance from keras with augmentations
image_gen = ImageDataGenerator(rotation_range=360,
width_shift_range=0.5,
height_shift_range=0.5,
zoom_range=[0.2, 0.7],
horizontal_flip=True,
vertical_flip=True,
fill_mode='reflect',
data_format='channels_last',
brightness_range=[0.5, 1.5])
#seed for random augmentations
random_seed = int(random.random() * 100000)
#generate augmented images
y_train = image_gen.flow_from_directory('.', target_size = (height, width), batch_size = n_samples, class_mode = None, seed = random_seed)
y_train = y_train.__getitem__(0).copy() #fix for 'array doesn't own its data'
x_train = y_train.copy()
for i in range(n_samples):
# source images are blurred (slightly rotated, moved and merged together)
copy = x_train[i].copy()
counter = 1
for j in range(blur_copies):
if (random.random() * 2 > 1) or (j == 1): #50% chance to make more than one copy
rotation_angle = random.random() * rotation_max / 2 + rotation_max #minimum ratation is the half of maximum rotation
copy = rotate(copy, rotation_angle, reshape=False, order=5, mode='reflect')
move_amount_x = int(random.random() * max_move + 1) * 2 #allow for negative values
move_amount_x = move_amount_x - move_amount_x / 2
move_amount_y = int(random.random() * max_move + 1) * 2
move_amount_y = move_amount_y - move_amount_y / 2
copy = shift(copy, [move_amount_x, move_amount_y, 0], order=5, mode='reflect')
counter = counter + 1
x_train[i] = (1 - 1 / counter) * x_train[i] + 1 / counter * copy #blend back together
x_train[i] = np.clip(x_train[i], 0, 255)
#normalize images to [-1, 1]
x_train = x_train/127.5 - 1.
y_train = y_train/127.5 - 1.
return x_train, y_train
# + [markdown] colab_type="text" id="A9P-WqYbHvCX"
# ### base functions
# + colab_type="code" id="tdeUTBRaHvCY" colab={}
def fast_normalization(x): # use clipping instead of batchnormalization for network stabilization
return Lambda(lambda x: K.clip(x, -1, 1), output_shape=lambda s: (s[0], s[1], s[2], s[3]))(x)
def residual_block(inputs): #combined pixel shuffle and squeeze
x = inputs
x = Conv2D(32, kernel_size = 9, activation = 'tanh', padding = 'same', strides = 2)(x)
x = SeparableConv2D(128, kernel_size = 9, activation = 'tanh', padding = 'same')(x) # rapidly increase speed at slightly worse results
x = fast_normalization(x)
x = Lambda(lambda x: K.reshape(x, (K.shape(x)[0], K.shape(x)[1], K.shape(x)[2], 32, 2, 2)), output_shape = lambda s: (s[0], s[1], s[2], s[3] // 4, 2, 2))(x)
x = Permute((3, 2, 4, 1, 5))(x)
x = Lambda(lambda x: K.reshape(x, (K.shape(x)[0], K.shape(x)[1], K.shape(x)[2] * K.shape(x)[3], K.shape(x)[4] * K.shape(x)[5])), output_shape = lambda s: (s[0], s[1], s[2] * s[3], s[4] * s[5]))(x)
x = Permute((3, 2, 1))(x)
#---
x1 = x
x = GlobalAveragePooling2D()(x)
x = Dense(8, activation = 'relu')(x) #reduction like in RCAN
x = Dense(32, activation = 'hard_sigmoid')(x)
x = Reshape((1, 1, 32))(x)
x = Multiply()([x1, x])
x = Add()([inputs, x])
return x
# + [markdown] colab_type="text" id="Gvy5AbKwHvCi"
# ### build generator model
# + colab_type="code" id="SKFcfdh_HvCf" outputId="2bb99668-6eb8-49c5-aae2-2ead416be9e6" colab={"base_uri": "https://localhost:8080/", "height": 1000}
x = inputs = Input(shape = (height_lr, width_lr, channels))
x = Conv2D(32, kernel_size = 3, padding = 'same', activation = 'tanh')(x)
x = residual_block(x)
x = residual_block(x)
x = residual_block(x)
x = residual_block(x)
x = Conv2D(3, kernel_size = 3, padding = 'same', activation = 'tanh')(x)
x = fast_normalization(x)
generator = Model(inputs = inputs, outputs = x)
generator.summary()
# + [markdown] colab_type="text" id="EmBC-aMQHvCk"
# ### train
# + colab_type="code" id="hkYfqMpN2MIQ" colab={}
#load checkpoint & compile the generator network
print('trying to load last saved weights...', end = ' ')
try:
generator.load_weights('deblurring_weights')
print('success.')
except:
print('failed')
pass
generator.compile(optimizer = Adam(gen_lr), loss = 'mae')
# Train generator
def logging(epoch, logs):
if epoch % logging_steps == 0:
testX, testY = get_batch(1, height_lr, width_lr, channels)
clear_output()
print('epoch', real_epoch + 1, '/', epochs, '--> step', epoch, '/', steps_per_epoch, ': loss', logs['loss'])
testZ = generator.predict(testX)
show([testX, testZ, testY])
print('test_loss:', generator.evaluate(testX, testY, verbose = 0))
logging_callback = LambdaCallback(
on_epoch_end=lambda epoch, logs: logging(epoch, logs)
)
for real_epoch in range(epochs):
X, Y = get_batch(batch_size, height_lr, width_lr, channels)
generator.fit(X, Y, batch_size, epochs = steps_per_epoch, verbose = 0, callbacks = [logging_callback], shuffle = True)
try:
print('trying to save weights...', end = ' ')
generator.save_weights('deblurring_weights')
print('success.')
except:
print('failed.')
# + [markdown] colab_type="text" id="xo-_GBp6NE8s"
# ### validate on complete picture
# + [markdown] id="nGLX-J7rxLD5" colab_type="text"
#
# + colab_type="code" id="XzRnnhw6fRkH" colab={}
from PIL import Image
testY = np.array(Image.open('./Set14/lenna.png'))
testX = testY.copy()
height = testX.shape[0]
width = testX.shape[1]
max_move = height / 100
# source image distortion wirth motionblur
copy = testX.copy()
counter = 1
for j in range(blur_copies):
if (random.random() * 2 > 1) or (j == 1): #50% chance to make more than one copy
rotation_angle = random.random() * rotation_max / 2 + rotation_max #minimum ratation is the half of maximum rotation
copy = rotate(copy, rotation_angle, reshape=False, order=5, mode='reflect')
move_amount_x = int(random.random() * max_move + 1) * 2 #allow for negative values
move_amount_x = move_amount_x - move_amount_x / 2
move_amount_y = int(random.random() * max_move + 1) * 2
move_amount_y = move_amount_y - move_amount_y / 2
copy = shift(copy, [move_amount_x, move_amount_y, 0], order=5, mode='reflect')
counter = counter + 1
testX = (1 - 1 / counter) * testX + 1 / counter * copy #blend back together
testX = np.clip(testX, 0, 255)
testX = testX /127.5 - 1
testY = testY /127.5 - 1
x = inputs = Input(shape = testX.shape)
x = Conv2D(32, kernel_size = 3, padding = 'same', activation = 'tanh')(x)
x = residual_block(x)
x = residual_block(x)
x = residual_block(x)
x = residual_block(x)
x = Conv2D(3, kernel_size = 3, padding = 'same', activation = 'tanh')(x)
x = fast_normalization(x)
generator = Model(inputs = inputs, outputs = x)
print('trying to load last saved weights...', end = ' ')
try:
generator.load_weights('deblurring_weights')
print('success.')
except:
print('failed')
pass
predicted = generator.predict(np.expand_dims((testX), 0))
show([testX, predicted, testY])
predicted = np.squeeze(predicted)
predicted = Image.fromarray(((predicted + 1) * 127.5).astype(np.uint8))
print('trying to save image as \'debluring_result.png\'...', end = ' ')
try:
predicted.save('debluring_result.png', "PNG")
print('success.')
except:
print('failed.')
pass
| deblurring.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Objectives" data-toc-modified-id="Objectives-1"><span class="toc-item-num">1 </span>Objectives</a></span></li><li><span><a href="#What-Are-Time-Series-Data?" data-toc-modified-id="What-Are-Time-Series-Data?-2"><span class="toc-item-num">2 </span>What Are Time Series Data?</a></span><ul class="toc-item"><li><span><a href="#Some-Examples" data-toc-modified-id="Some-Examples-2.1"><span class="toc-item-num">2.1 </span>Some Examples</a></span></li><li><span><a href="#Uses-for-Time-Series" data-toc-modified-id="Uses-for-Time-Series-2.2"><span class="toc-item-num">2.2 </span>Uses for Time Series</a></span></li><li><span><a href="#Example-Data" data-toc-modified-id="Example-Data-2.3"><span class="toc-item-num">2.3 </span>Example Data</a></span></li></ul></li><li><span><a href="#Datetime-Objects" data-toc-modified-id="Datetime-Objects-3"><span class="toc-item-num">3 </span>Datetime Objects</a></span><ul class="toc-item"><li><span><a href="#Setting-Datetime-Objects-as-the-Index" data-toc-modified-id="Setting-Datetime-Objects-as-the-Index-3.1"><span class="toc-item-num">3.1 </span>Setting Datetime Objects as the Index</a></span></li><li><span><a href="#Investigating-Time-Series-with-Datetime-Objects" data-toc-modified-id="Investigating-Time-Series-with-Datetime-Objects-3.2"><span class="toc-item-num">3.2 </span>Investigating Time Series with Datetime Objects</a></span></li></ul></li><li><span><a href="#Resampling-Techniques" data-toc-modified-id="Resampling-Techniques-4"><span class="toc-item-num">4 </span>Resampling Techniques</a></span><ul class="toc-item"><li><span><a href="#Aside:-Deeper-Exploration" data-toc-modified-id="Aside:-Deeper-Exploration-4.1"><span class="toc-item-num">4.1 </span>Aside: Deeper Exploration</a></span></li></ul></li><li><span><a href="#Visualizing-Time-Series" data-toc-modified-id="Visualizing-Time-Series-5"><span class="toc-item-num">5 </span>Visualizing Time Series</a></span><ul class="toc-item"><li><span><a href="#Showing-Changes-Over-Time" data-toc-modified-id="Showing-Changes-Over-Time-5.1"><span class="toc-item-num">5.1 </span>Showing Changes Over Time</a></span><ul class="toc-item"><li><span><a href="#Line-Plot" data-toc-modified-id="Line-Plot-5.1.1"><span class="toc-item-num">5.1.1 </span>Line Plot</a></span></li><li><span><a href="#Dot-Plot" data-toc-modified-id="Dot-Plot-5.1.2"><span class="toc-item-num">5.1.2 </span>Dot Plot</a></span></li><li><span><a href="#Grouping-Plots" data-toc-modified-id="Grouping-Plots-5.1.3"><span class="toc-item-num">5.1.3 </span>Grouping Plots</a></span><ul class="toc-item"><li><span><a href="#All-Annual-Separated" data-toc-modified-id="All-Annual-Separated-5.1.3.1"><span class="toc-item-num">5.1.3.1 </span>All Annual Separated</a></span></li><li><span><a href="#All-Annual-Together" data-toc-modified-id="All-Annual-Together-5.1.3.2"><span class="toc-item-num">5.1.3.2 </span>All Annual Together</a></span></li></ul></li></ul></li><li><span><a href="#Showing-Distributions" data-toc-modified-id="Showing-Distributions-5.2"><span class="toc-item-num">5.2 </span>Showing Distributions</a></span><ul class="toc-item"><li><span><a href="#Histogram" data-toc-modified-id="Histogram-5.2.1"><span class="toc-item-num">5.2.1 </span>Histogram</a></span></li><li><span><a href="#Density" data-toc-modified-id="Density-5.2.2"><span class="toc-item-num">5.2.2 </span>Density</a></span></li><li><span><a href="#Box-Plot" data-toc-modified-id="Box-Plot-5.2.3"><span class="toc-item-num">5.2.3 </span>Box Plot</a></span></li><li><span><a href="#Heat-Maps" data-toc-modified-id="Heat-Maps-5.2.4"><span class="toc-item-num">5.2.4 </span>Heat Maps</a></span><ul class="toc-item"><li><span><a href="#Example-of-how-heat-maps-are-useful" data-toc-modified-id="Example-of-how-heat-maps-are-useful-5.2.4.1"><span class="toc-item-num">5.2.4.1 </span>Example of how heat maps are useful</a></span></li></ul></li></ul></li></ul></li><li><span><a href="#Level-Up" data-toc-modified-id="Level-Up-6"><span class="toc-item-num">6 </span>Level Up</a></span><ul class="toc-item"><li><span><a href="#EDA" data-toc-modified-id="EDA-6.1"><span class="toc-item-num">6.1 </span>EDA</a></span></li></ul></li></ul></div>
# +
import pandas as pd
import numpy as np
pd.set_option('display.max_rows', 1000)
import matplotlib.pyplot as plt
import seaborn as sns
plt.style.use('ggplot')
# + [markdown] heading_collapsed=true
# # Objectives
# + [markdown] hidden=true
# - Understand the use case for time series data
# - Manipulate datetime objects
# - Understand different resampling techniques
# - Implement different visualization techniques for time series data
# + [markdown] heading_collapsed=true
# # What Are Time Series Data?
# + [markdown] hidden=true
# > We can saw data is a **time series** when the temporal information is a key focus of the data.
# + [markdown] hidden=true
# Data in a time series can stem from historical data or data that is dependent on past values.
# + [markdown] heading_collapsed=true hidden=true
# ## Some Examples
# + [markdown] hidden=true
# - Stock prices
# - Atmospheric changes over the course of decades
# - Audio samples
# - Heart rate data
# + [markdown] heading_collapsed=true hidden=true
# ## Uses for Time Series
# + [markdown] hidden=true
# - Understand some underlying process
# - Forecasting (what we'll mostly focus on)
# - Imputation (filling missing "past" data)
# - Anomaly detection
# + [markdown] heading_collapsed=true hidden=true
# ## Example Data
# + hidden=true
# Define a function that will help us load and
# clean up a dataset.
def load_trend(trend_name='football', country_code='us'):
df = pd.read_csv('data/google-trends_'
+ trend_name + '_'
+ country_code
+ '.csv').iloc[1:, :]
df.columns = ['counts']
df['counts'] = df['counts'].str.replace('<1', '0').astype(int)
return df
# + hidden=true
df = load_trend(**{'trend_name': 'data-science', 'country_code': 'us'})
df.head()
# + [markdown] hidden=true
# Now we can do this with multiple time series data!
# + hidden=true
trends = [
{'trend_name': 'data-science', 'country_code': 'us'},
{'trend_name': 'football', 'country_code': 'us'},
{'trend_name': 'football', 'country_code': 'uk'},
{'trend_name': 'coronavirus', 'country_code': 'us'},
{'trend_name': 'trump', 'country_code': 'us'},
{'trend_name': 'taxes', 'country_code': 'us'},
{'trend_name': 'avengers', 'country_code': 'us'}
]
# + hidden=true
trend_dfs = [load_trend(**trend) for trend in trends]
# + [markdown] heading_collapsed=true
# # Datetime Objects
# + [markdown] hidden=true
# Datetime objects make our time series modeling lives easier. They will allow us to perform essential data prep tasks with a few lines of code.
#
# We need our time series **index** to be datetime objects, since our models will rely on being able to identify the previous chronological value.
# + [markdown] hidden=true
# There is a `datetime` [library](https://docs.python.org/2/library/datetime.html), and inside `pandas` there is a datetime module as well as a to_datetime() function.
#
# For time series modeling, the first step often is to make sure that the index is a datetime object.
# + [markdown] heading_collapsed=true hidden=true
# ## Setting Datetime Objects as the Index
# + [markdown] hidden=true
# There are a few ways to **reindex** our series to datetime.
#
# We can use `pandas.to_datetime()` method:
# + hidden=true
ts_no_datetime = pd.read_csv('data/Gun_Crimes_Heat_Map.csv')
# + hidden=true
ts_no_datetime.head()
# + hidden=true
ts_no_datetime.index
# + hidden=true
ts = ts_no_datetime.set_index(pd.to_datetime(ts_no_datetime['Date']), drop=True)
# + [markdown] hidden=true
# > Alternatively, we can parse the dates directly on import
# + hidden=true
ts = pd.read_csv('data/Gun_Crimes_Heat_Map.csv', index_col='Date', parse_dates=True)
# + hidden=true
print(f"Now our index is a {type(ts.index)}")
# + hidden=true
ts.head()
# + [markdown] heading_collapsed=true hidden=true
# ## Investigating Time Series with Datetime Objects
# + [markdown] hidden=true
# Datetime objects include aspects of the date as attributes, like month and year:
# + hidden=true
ts.index[0]
# + hidden=true
ts.index[0].month
# + hidden=true
ts.index[0].year
# + [markdown] hidden=true
# We can also use the date to directly slice the DataFrame
# + hidden=true
# Only data after 2021
ts['2021':]
# + hidden=true
# Only data from this time period
ts['2020-02-01 00:00':'2020-02-01 01:00']
# + [markdown] hidden=true
# We can easily see now whether offenses happen, for example, during business hours.
# + hidden=true
fig, ax = plt.subplots()
ts['hour'] = ts.index
ts['hour'] = ts.hour.apply(lambda x: x.hour)
ts['business_hours'] = ts.hour.apply(lambda x: 9 <= x <= 17)
bh_ratio = ts.business_hours.value_counts()[1]/len(ts)
x = ts.business_hours.value_counts().index
y = ts.business_hours.value_counts()
sns.barplot(x=x, y=y)
ax.set_title(f'{bh_ratio: 0.2%} of Offenses\n Happen Btwn 9 and 5');
# + [markdown] heading_collapsed=true
# # Resampling Techniques
# + [markdown] hidden=true
# > **Resampling** allows us to convert the time series into a particular frequency
#
# https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.resample.html
# https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#resampling
# + [markdown] hidden=true
# With a Datetime index, we also have new abilities, such as **resampling**.
#
# To create our timeseries, we will count the number of gun offenses reported per day.
# + hidden=true
ts.resample('D')
# + [markdown] hidden=true
# There are many possible units for resampling, each with its own alias:
# + [markdown] hidden=true
# <table style="display: inline-block">
# <caption style="text-align: center"><strong>TIME SERIES OFFSET ALIASES</strong></caption>
# <tr><th>ALIAS</th><th>DESCRIPTION</th></tr>
# <tr><td>B</td><td>business day frequency</td></tr>
# <tr><td>C</td><td>custom business day frequency (experimental)</td></tr>
# <tr><td>D</td><td>calendar day frequency</td></tr>
# <tr><td>W</td><td>weekly frequency</td></tr>
# <tr><td>M</td><td>month end frequency</td></tr>
# <tr><td>SM</td><td>semi-month end frequency (15th and end of month)</td></tr>
# <tr><td>BM</td><td>business month end frequency</td></tr>
# <tr><td>CBM</td><td>custom business month end frequency</td></tr>
# <tr><td>MS</td><td>month start frequency</td></tr>
# <tr><td>SMS</td><td>semi-month start frequency (1st and 15th)</td></tr>
# <tr><td>BMS</td><td>business month start frequency</td></tr>
# <tr><td>CBMS</td><td>custom business month start frequency</td></tr>
# <tr><td>Q</td><td>quarter end frequency</td></tr>
# <tr><td></td><td><font color=white>intentionally left blank</font></td></tr></table>
#
# <table style="display: inline-block; margin-left: 40px">
# <caption style="text-align: center"></caption>
# <tr><th>ALIAS</th><th>DESCRIPTION</th></tr>
# <tr><td>BQ</td><td>business quarter endfrequency</td></tr>
# <tr><td>QS</td><td>quarter start frequency</td></tr>
# <tr><td>BQS</td><td>business quarter start frequency</td></tr>
# <tr><td>A</td><td>year end frequency</td></tr>
# <tr><td>BA</td><td>business year end frequency</td></tr>
# <tr><td>AS</td><td>year start frequency</td></tr>
# <tr><td>BAS</td><td>business year start frequency</td></tr>
# <tr><td>BH</td><td>business hour frequency</td></tr>
# <tr><td>H</td><td>hourly frequency</td></tr>
# <tr><td>T, min</td><td>minutely frequency</td></tr>
# <tr><td>S</td><td>secondly frequency</td></tr>
# <tr><td>L, ms</td><td>milliseconds</td></tr>
# <tr><td>U, us</td><td>microseconds</td></tr>
# <tr><td>N</td><td>nanoseconds</td></tr></table>
# + [markdown] hidden=true
# When resampling, we have to provide a rule to resample by, and an **aggregate function**.
# + [markdown] hidden=true
# **To upsample** is to increase the frequency of the data of interest.
# **To downsample** is to decrease the frequency of the data of interest.
#
# For our purposes, we will downsample, and count the number of occurences per day.
# + hidden=true
ts.resample('D').count()
# + [markdown] hidden=true
# Our time series will consist of a series of counts of gun reports per day.
# + hidden=true
# ID is unimportant. We could choose any column, since the counts are the same.
ts = ts.resample('D').count()['ID']
# + hidden=true
ts
# + [markdown] hidden=true
# Let's visualize our timeseries with a plot.
# + hidden=true
fig, ax = plt.subplots(figsize=(10, 5))
ax.plot(ts.index, ts.values)
ax.set_title('Gun Crimes per day in Chicago')
ax.set_ylabel('Reported Gun Crimes');
# + [markdown] hidden=true
# There seems to be some abnormal activity happening towards the end of our series.
#
# **[sun-times](https://chicago.suntimes.com/crime/2020/6/8/21281998/chicago-deadliest-day-violence-murder-history-police-crime)**
# + [markdown] heading_collapsed=true hidden=true
# ## Aside: Deeper Exploration
# + hidden=true
ts.sort_values(ascending=False)[:10]
# + [markdown] hidden=true
# Let's treat the span of days from 5-31 to 6-03 as outliers.
#
# There are several ways to do this, but let's first remove the outliers, and populate an an empty array with the original date range. That will introduce us to the `pandas.date_range()` method.
# + hidden=true
daily_count = ts[ts < 90]
ts_dr = pd.date_range(daily_count.index[0], daily_count.index[-1])
ts_daily = np.empty(shape=len(ts_dr))
ts_daily = pd.Series(ts_daily)
ts_daily = ts_daily.reindex(ts_dr)
ts = ts_daily.fillna(daily_count)
# + hidden=true
ts
# + hidden=true
fig, ax = plt.subplots(figsize=(10, 5))
ts.plot(ax=ax)
ax.set_title('Gun Crimes in Chicago with Deadliest Days Removed');
# + [markdown] hidden=true
# Let's zoom in on that week again:
# + hidden=true
fig, ax = plt.subplots()
ax.plot(ts[(ts.index > '2020-05-20')
& (ts.index < '2020-06-07')]
)
ax.tick_params(rotation=45)
ax.set_title('We have some gaps now');
# + [markdown] hidden=true
# The datetime object allows us several options of how to fill those gaps:
# + hidden=true
# .ffill()
fig, (ax1, ax2) = plt.subplots(1, 2, figsize = (10, 5))
ax1.plot(ts.ffill()[(ts.index > '2020-05-20')
& (ts.index < '2020-06-07')]
)
ax1.tick_params(rotation=45)
ax1.set_title('Forward Fill')
ax2.plot(ts[(ts.index > '2020-05-20')
& (ts.index < '2020-06-07')]
)
ax2.tick_params(rotation=45)
ax2.set_title('Original');
# + hidden=true
# .bfill()
fig, (ax1, ax2) = plt.subplots(1, 2, figsize = (10, 5))
ax1.plot(ts.bfill()[(ts.index > '2020-05-20')
& (ts.index < '2020-06-07')]
)
ax1.tick_params(rotation=45)
ax1.set_title('Back Fill')
ax2.plot(ts[(ts.index > '2020-05-20')
& (ts.index < '2020-06-07')]
)
ax2.tick_params(rotation=45)
ax2.set_title('Original');
# + hidden=true
# .interpolate()
fig, (ax1, ax2) = plt.subplots(1, 2, figsize = (10, 5))
ax1.plot(ts.interpolate()[(ts.index > '2020-05-20')
& (ts.index < '2020-06-07')]
)
ax1.tick_params(rotation=45)
ax1.set_title('Interpolation')
ax2.plot(ts[(ts.index > '2020-05-20')
& (ts.index < '2020-06-07')]
)
ax2.tick_params(rotation=45)
ax2.set_title('Original');
# + [markdown] hidden=true
# Let's proceed with the interpolated data.
# + hidden=true
ts = ts.interpolate()
ts.isna().sum()
# + [markdown] hidden=true
# Now that we've cleaned up a few data points, let's downsample to the week level.
# + hidden=true
ts_weekly = ts.resample('W').mean()
# + hidden=true
ts_weekly.plot();
# + [markdown] heading_collapsed=true
# # Visualizing Time Series
# + [markdown] hidden=true
# There can be a lot information to be found in time series! Visualizations can help us tease out this information to something we can more easily observe.
# + [markdown] heading_collapsed=true hidden=true
# ## Showing Changes Over Time
# + [markdown] hidden=true
# Can identify patterns and trends with visualizations
# + hidden=true
# New York Stock Exchange average monthly returns [1961-1966] from curriculum
nyse = pd.read_csv("data/NYSE_monthly.csv")
col_name= 'Month'
nyse[col_name] = pd.to_datetime(nyse[col_name])
nyse.set_index(col_name, inplace=True)
# + hidden=true
display(nyse.head(10))
display(nyse.info())
# + [markdown] heading_collapsed=true hidden=true
# ### Line Plot
# + hidden=true
nyse.plot(figsize = (16,6))
plt.show()
# + [markdown] heading_collapsed=true hidden=true
# ### Dot Plot
# + hidden=true
nyse.plot(figsize = (16,6), style="*")
plt.show()
# + [markdown] hidden=true
# > Note the difference between this and the line plot.
# >
# > When might you want a dot vs a line plot?
# + [markdown] heading_collapsed=true hidden=true
# ### Grouping Plots
# + [markdown] hidden=true
# What if we wanted to look at year-to-year (e.g., temperature throughout many years)
#
# There are a couple options to choose from.
# + [markdown] heading_collapsed=true hidden=true
# #### All Annual Separated
# -
year_groups == nyse.groupby()
# + hidden=true
# Annual Frequency
year_groups = nyse.groupby(pd.Grouper(freq ='A'))
#Create a new DataFrame and store yearly values in columns
nyse_annual = pd.DataFrame()
for yr, group in year_groups:
nyse_annual[yr.year] = group.values.ravel()
# Plot the yearly groups as subplots
nyse_annual.plot(figsize = (13,8), subplots=True, legend=True)
plt.show()
# + [markdown] heading_collapsed=true hidden=true
# #### All Annual Together
# + hidden=true
# Plot overlapping yearly groups
nyse_annual.plot(figsize = (15,5), subplots=False, legend=True)
plt.show()
# + [markdown] heading_collapsed=true hidden=true
# ## Showing Distributions
# + [markdown] hidden=true
# Sometimes the distribution of the values are important.
#
# What are some reasons?
#
# - Checking for normality (for stat testing)
# - First check on raw & transformed data
# + [markdown] heading_collapsed=true hidden=true
# ### Histogram
# + hidden=true
nyse.hist(figsize = (10,6))
plt.show()
# + hidden=true
# Bin it to make it more obvious if normal
nyse.hist(figsize = (10,6), bins = 7)
plt.show()
# + [markdown] heading_collapsed=true hidden=true
# ### Density
# + hidden=true
nyse.plot(kind='kde', figsize = (15,10))
plt.show()
# + [markdown] heading_collapsed=true hidden=true
# ### Box Plot
# + [markdown] hidden=true
# - Shows distribution over time
# - Can help show outliers
# - Seasonal trends
# + hidden=true
# Generate a box and whiskers plot for temp_annual dataframe
nyse_annual.boxplot(figsize = (12,7))
plt.show()
# + [markdown] heading_collapsed=true hidden=true
# ### Heat Maps
# + [markdown] hidden=true
# Use color to show patterns throughout a time period for data
# + [markdown] heading_collapsed=true hidden=true
# #### Example of how heat maps are useful
# + hidden=true
df_temp = pd.read_csv(
'data/min_temp.csv', # Data to read
index_col=0, # Use the first column as index ('Date')
parse_dates=True, # Have Pandas parse the dates
infer_datetime_format=True, # Make Pandas try to parse dates automatically
dayfirst=True # Impoprtant to know format is DD/MM
)
# + hidden=true
display(df_temp.head())
display(df_temp.info())
# + hidden=true
# Create a new DataFrame and store yearly values in columns for temperature
temp_annual = pd.DataFrame()
for yr, group in df_temp.groupby(pd.Grouper(freq ='A')):
temp_annual[yr.year] = group.values.ravel()
# + [markdown] heading_collapsed=true hidden=true
# ##### Plotting each line plot in a subplot
# + [markdown] hidden=true
# Let's use our strategy in plotting multiple line plots to see if we can see a pattern:
# + hidden=true
# Plot the yearly groups as subplots
temp_annual.plot(figsize = (16,8), subplots=True, legend=True)
plt.show()
# + [markdown] hidden=true
# You likely will have a hard time seeing exactly the temperature shift is throughout the year (if it even exists!)
#
# We can try plotting all the lines together to see if a pattern is more obvious in our visual.
# + [markdown] heading_collapsed=true hidden=true
# ##### Plotting all line plots in one plot
# + hidden=true
# Plot overlapping yearly groups
temp_annual.plot(figsize = (15,5), subplots=False, legend=True)
plt.show()
# + [markdown] hidden=true
# That's great we can see that the temperature decreases in the middle of the data! But now we sacrificed being able to observe any pattern for an individual year.
#
# This is where using a heat map can help visualize patterns throughout the year for temperature! And of course, the heat map can be used for more than just temperature related data.
# + [markdown] heading_collapsed=true hidden=true
# ##### And finally, using a heat map to visualize a pattern
# + hidden=true
# Year and month
year_matrix = temp_annual.T
plt.matshow(year_matrix, interpolation=None, aspect='auto', cmap=plt.cm.Spectral_r)
plt.show()
# + [markdown] hidden=true
# ☝🏼 Look at that beautiful visual pattern! Makes me want to weep with joy for all the information density available to us!
# + [markdown] heading_collapsed=true
# # Level Up
# + [markdown] heading_collapsed=true hidden=true
# ## EDA
#
# Let's import some data on **gun violence in Chicago**.
#
# [source](https://data.cityofchicago.org/Public-Safety/Gun-Crimes-Heat-Map/iinq-m3rg)
# + hidden=true
ts = pd.read_csv('data/Gun_Crimes_Heat_Map.csv')
# + hidden=true
ts.head()
# + [markdown] hidden=true
# Let's look at some summary stats:
# + hidden=true
print(f"There are {ts.shape[0]} records in our timeseries")
# + hidden=true
# Definitely some messy input of our Desciption data
ts['Description'].value_counts()
# + hidden=true
height = ts['Description'].value_counts()[:10]
offense_names = ts['Description'].value_counts()[:10].index
fig, ax = plt.subplots()
sns.barplot(height, offense_names, color='r', ax=ax)
ax.set_title('Mostly Handgun offenses');
# + hidden=true
# Mostly non-domestic offenses
fig, ax = plt.subplots()
sns.barplot( ts['Domestic'].value_counts().index,
ts['Domestic'].value_counts(),
palette=[ 'r', 'b'], ax=ax
)
ax.set_title("Overwhelmingly Non-Domestic Offenses");
# + hidden=true
# Mostly non-domestic offenses
arrest_rate = ts['Arrest'].value_counts()[1]/len(ts)
fig, ax = plt.subplots()
sns.barplot( ts['Arrest'].value_counts().index,
ts['Arrest'].value_counts(),
palette=['r', 'g'], ax=ax
)
ax.set_title(f'{arrest_rate: 0.2%} of Total Cases\n Result in Arrest');
# + hidden=true
fig, ax = plt.subplots()
sns.barplot( ts['Year'].value_counts().index,
ts['Year'].value_counts(),
color= 'r', ax=ax
)
ax.set_title("Offenses By Year");
# + [markdown] hidden=true
# While this does show some interesting information that will be relevant to our time series analysis, we are going to get more granular.
| Phase_4/Time Series/ds-time_series_data_manipulation-kvo32-main/time_series.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import torch
from ask_attack import ASKAttack
from dknn import DKNN
from models.vgg import VGG16
from data_utils import get_dataloaders
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# -
# # ASK attack on CIFAR-10
# ## Adversarial trained model
# +
model = VGG16()
model.load_state_dict(torch.load("./checkpoints/cifar10_vgg16_at.pt"))
model.to(device)
model.eval()
trainloader, testloader = get_dataloaders(
"cifar10",
root="./datasets",
batch_size=1000,
download=False,
augmentation=False,
train_shuffle=False,
num_workers=1
)
train_data, train_targets = [], []
for x, y in trainloader:
train_data.append(x)
train_targets.append(y)
train_data = torch.cat(train_data, dim=0)
train_targets = torch.cat(train_targets)
ask_attack = ASKAttack(
model,
train_data,
train_targets,
max_iter=20,
temperature=0.01,
hidden_layers=[3, ],
class_samp_size=2000,
metric="cosine",
random_seed=3,
device=device
)
dknn = DKNN(
model,
torch.cat(ask_attack.train_data, dim=0),
torch.arange(ask_attack.n_class).repeat_interleave(ask_attack.class_samp_size),
hidden_layers=ask_attack.hidden_layers,
metric=ask_attack.metric,
device=device
)
# +
x_batch, y_batch = [], []
batch_count = 5
for i,(x,y) in enumerate(testloader):
if i == batch_count:
break
x_batch.append(x)
y_batch.append(y)
x_batch = torch.cat(x_batch, dim=0)
y_batch = torch.cat(y_batch)
pred_dknn_clean = dknn.predict(x_batch)
print("Clean accuracy of DkNN is {}".format(
(pred_dknn_clean.argmax(axis=1) == y_batch.numpy()).astype("float").mean()
))
x_adv = ask_attack.generate(x_batch, y_batch)
pred_dknn_adv = dknn.predict(x_adv)
print("Adversarial accuracy of DkNN is {}".format(
(pred_dknn_adv.argmax(axis=1) == y_batch.numpy()).astype("float").mean()
))
# -
# # Ask defense
# +
model = VGG16()
model.load_state_dict(torch.load("./checkpoints/cifar10_vgg16_askdef.pt"))
model.to(device)
model.eval()
train_data, train_targets = [], []
for x, y in trainloader:
train_data.append(x)
train_targets.append(y)
train_data = torch.cat(train_data, dim=0)
train_targets = torch.cat(train_targets)
ask_attack = ASKAttack(
model,
train_data,
train_targets,
max_iter=20,
temperature=0.01,
hidden_layers=[3, ],
class_samp_size=2000,
metric="cosine",
random_seed=3,
device=device
)
dknn = DKNN(
model,
torch.cat(ask_attack.train_data, dim=0),
torch.arange(ask_attack.n_class).repeat_interleave(ask_attack.class_samp_size),
hidden_layers=ask_attack.hidden_layers,
metric=ask_attack.metric,
device=device
)
pred_dknn_clean = dknn.predict(x_batch)
print("Clean accuracy of DkNN is {}".format(
(pred_dknn_clean.argmax(axis=1) == y_batch.numpy()).astype("float").mean()
))
x_adv = ask_attack.generate(x_batch, y_batch)
pred_dknn_adv = dknn.predict(x_adv)
print("Adversarial accuracy of DkNN is {}".format(
(pred_dknn_adv.argmax(axis=1) == y_batch.numpy()).astype("float").mean()
))
| run_attack.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
# +
# Source: Texas Health and Human Services Commission
# Data provided Sept. 6, 2018
dat_orig = pd.read_excel(
'../src/Andrea Ball #16064 Child Injury Data 9.6.18.xlsx',
skiprows=4,
skipfooter=3
)
# Data provided Nov. 14, 2018
dat = pd.read_excel(
'../src/D91909_Inv_InjuryData_Center_Home.xlsx',
skiprows=4,
skipfooter=3
)
# +
print(
'Sept. 6 data:\n{:,} total injuries\n{:,} serious injuries'.format(
dat_orig.shape[0],
dat_orig.rename(
columns = lambda x: x.replace(' ', '')
).query('InjuryDetermination == "Serious"').shape[0]
)
)
print(
'\n\nNov. 14 data:\n{:,} total injuries\n{:,} serious injuries'.format(
dat.shape[0],
dat.rename(
columns = lambda x: x.replace(' ', '')
).query('InjuryDetermination == "Serious"').shape[0]
)
)
# -
pd.crosstab(
dat['Injury Related to Abuse/Neglect'],
dat['Injury Determination']
)
dat['Injury Determination'].value_counts()
dat['Injury Related to Abuse/Neglect'].value_counts()
dat['Injury Determination'].value_counts(normalize=True).apply('{:.2%}'.format)
dat['Operation Type3'].value_counts(normalize=True).apply('{:.2%}'.format)
dat['Injury Caused By'].value_counts().apply('{:,}'.format)
| notebooks/injury_data_request_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DoE - Modern DoE example: Metamodels for black-box systems
#
# The purpose of this notebook is to show you an example of how to apply some of the modern DoE techniques available through simulation, see [the associated presentation](doe_erigrid_dtu_summerschool_2018-08-28.pdf).
# %matplotlib notebook
import pandas as pd
import HEMS_sim
import sobol_seq # you likely need to install this first
import numpy as np
import matplotlib.pyplot as plt
from util import block_print, enable_print
# +
# Dictionary with basic configuration of the simulation
basic_conf = {
'ID':'00',
'batt_storage_capacity':20,
'batt_charge_capacity':5,
'pv1_scaling':1,
'controller_change_rate':0.5,
'climate_conditions':'sunny',
'season':'autumn',
'random_weather':False,
'stochastic':False, # this time we exclude the fluctuation (deterministic simulation)
'noise_scale':1}
# +
# Scenario name which determines the name of the files we will be saving with the results
scenario_name = 'without_noise'
# Do deterministic space-filling sampling with Sobol sequence
samples = 15 # more samples would be better if you have the time ;-)
z = sobol_seq.i4_sobol_generate(2, samples)
z = np.array(z) * np.array([10, 5]) + np.array([15, 2]) # Sobol samples are between 0 and 1, here we scale
# Range for factor 1 (battery storage capacity): 15 to 25
# Range for factor 2 (battery charge capacity): 2 to 7
variations = {}
for i in range(samples):
run_id = 'run_' + str(i)
variations[run_id] = {'ID': str(i), 'batt_storage_capacity': z[i][0], 'batt_charge_capacity': z[i][1]}
# Merging of the basic configuration and the variations
recipes = {key: basic_conf.copy() for key in variations}
recipes = {key: {**recipes[key],**data} for key,data in variations.items()}
# -
# These are the parameter combinations which were selected by the Sobol sequence
plt.figure()
plt.plot(*(z.T), '*')
plt.xlabel('Battery Storage Capacity [kWh]')
plt.ylabel('Battery Charge Capacity [kW]')
# Run the simulations (delete temp files first)
for recipe in recipes:
print("Starting run {0}".format(recipes[recipe]['ID']))
block_print() # Print statements are blocked due to notebook message length limitations
HEMS_sim.run_simulation(scenario_name,recipes[recipe])
enable_print()
# Store results and get response summaries
run_store.close()
run_store = pd.HDFStore('temp_files/runs_summary_{}.h5'.format(scenario_name))
summaries = [run_store[k] for k in run_store.keys()]
run_store.close()
summaries = pd.concat(summaries, axis=0).set_index('ID')
# Get the used treatments (battery size and power)
exog = summaries[['battery storage capacity [kWh]', 'battery charge capacity[kW]']]
# Get the self consumption index as response for each treatment
endog = summaries[['Self consumption index']]
# We want to fit a metamodel to our system
# We chose Kriging (Gaussian Process) this time around. You can also choose other metamodels if you want
# (simplest would be linear interpolation)
from sklearn.gaussian_process import GaussianProcessRegressor
from sklearn.gaussian_process.kernels import RBF, ConstantKernel as C
# Kriging needs a kernel. This kernel parameterization should work, but you can also play around with it.
kernel = C(1.0, (1e-3, 1e3)) * RBF(10, (1e-2, 1e2))
gp = GaussianProcessRegressor(kernel=kernel, n_restarts_optimizer=9)
gp.fit(exog.values, endog.values) # Fit the Kriging model to our samples and associated responses
# +
# Next we need to evaluate our metamodel. Thus, we set up a 50x50 point evaluation grid:
x_vec, y_vec = np.meshgrid(np.linspace(15, 25, 50), np.linspace(2, 7, 50))
# x_vec and y_vec is for plotting. We reshape them to get vectors for predicting:
evalgrid = np.array([x_vec.flatten(), y_vec.flatten()]).T
# Using our metamodel for predictions (Kriging also gives us a sense of uncertainty via the sigma):
scipred, sigma = gp.predict(evalgrid, return_std=True)
# For plotting we need to reshape our prediction array:
pltsci = np.reshape(scipred, (50, 50))
# +
from matplotlib import pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
# %matplotlib
# This gives us the figure in an extra window
fig = plt.figure()
ax = fig.gca(projection='3d')
# Get 3D surface plot:
surf = ax.plot_surface(x_vec, y_vec, pltsci)
# Include our original sample points:
ax.plot(exog.values[:,0], exog.values[:,1], endog.values.ravel(), 'r*')
plt.show()
# Some questions to ask yourself in the end:
# - Does the metamodel make sense to you? If not, why?
# - Did you use enough samples?
| DoE-exercises/4_Modern_DoE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
warnings.filterwarnings("ignore")
# -
# ### Data Import
# +
usecols = [
'room_type', 'neighbourhood_group',
'latitude','longitude','price','minimum_nights','number_of_reviews','reviews_per_month','calculated_host_listings_count','availability_365']
data = pd.read_csv('/Users/muzalevskiy/Downloads/data-science-ml-master/DS_KnowHow/EDA_Preprocessing/data/AB_NYC_2019.csv', usecols=usecols)
# -
print(data.shape)
data.head(10)
data['reviews_per_month']=data.reviews_per_month.fillna(0)
# ### Frequent Values
data.describe(include=["O"])
data['neighbourhood_group'].mode()
# #### Answer for Question №1: 'Manhattan'
data_numeric = data.copy()
data_numeric = data.drop(["neighbourhood_group","room_type", "price"], axis=1)
data_numeric.describe()
# ### Correlation Calculation
data_numeric.corr()
plt.figure(figsize=(15,10))
sns.heatmap(data_numeric.corr(),annot=True,linewidths=.5, cmap="Blues")
plt.title('Heatmap showing correlations between numerical data')
plt.show()
data_numeric.corr().unstack().sort_values(ascending = False)
# #### Answer for Question №2: 'number_of_reviews' and 'reviews_per_month'
# +
data_class = data.copy()
mean = data_class['price'].mean()
data_class['above_average'] = np.where(data_class['price']>=mean,1,0)
# -
data_class = data_class.drop('price', axis=1)
from sklearn.model_selection import train_test_split
df_train_full, df_test = train_test_split(data_class, test_size=0.2, random_state=42)
df_train, df_val = train_test_split(df_train_full, test_size=0.25, random_state=42)
df_train = df_train.reset_index(drop=True)
df_val = df_val.reset_index(drop=True)
df_test = df_test.reset_index(drop=True)
y_train = df_train.above_average.values
y_val = df_val.above_average.values
y_test = df_test.above_average.values
# ### Mutual information
from sklearn.metrics import mutual_info_score
cat = ['neighbourhood_group', 'room_type']
# +
def calculate_mi(series):
return mutual_info_score(series, df_train.above_average)
df_mi = df_train[cat].apply(calculate_mi)
df_mi = df_mi.sort_values(ascending=False).to_frame(name='MI')
# -
df_mi
# #### Answer for Question №3: 'room_type'
df_train = df_train.drop('above_average', axis=1)
df_val = df_val.drop('above_average', axis=1)
df_test = df_test.drop('above_average', axis=1)
# ### Classification and accuracy of a model
from sklearn.feature_extraction import DictVectorizer
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score
num = ['latitude','longitude','minimum_nights','number_of_reviews','reviews_per_month','calculated_host_listings_count','availability_365']
# +
train_dict = df_train[cat + num].to_dict(orient='records')
dv = DictVectorizer(sparse=False)
dv.fit(train_dict)
X_train = dv.transform(train_dict)
# +
model = LogisticRegression(solver='lbfgs', C=1.0, random_state=42)
model.fit(X_train, y_train)
val_dict = df_val[cat + num].to_dict(orient='records')
X_val = dv.transform(val_dict)
y_pred = model.predict(X_val)
accuracy = np.round(accuracy_score(y_val, y_pred),2)
print(accuracy)
# -
# #### Answer for Question №4: 0.79
features = cat + num
features
# ### Feature elimination
# +
orig_score = accuracy
for c in features:
subset = features.copy()
subset.remove(c)
train_dict = df_train[subset].to_dict(orient='records')
dv = DictVectorizer(sparse=False)
dv.fit(train_dict)
X_train = dv.transform(train_dict)
model = LogisticRegression(solver='lbfgs', C=1.0, random_state=42)
model.fit(X_train, y_train)
val_dict = df_val[subset].to_dict(orient='records')
X_val = dv.transform(val_dict)
y_pred = model.predict(X_val)
score = accuracy_score(y_val, y_pred)
print(c, orig_score - score, score)
# -
# #### Answer for Question №5: smallest difference - 'number_of_reviews'
data['price']=np.log1p(data['price'])
df_train_full, df_test = train_test_split(data, test_size=0.2, random_state=42)
df_train, df_val = train_test_split(df_train_full, test_size=0.25, random_state=42)
df_train = df_train.reset_index(drop=True)
df_val = df_val.reset_index(drop=True)
df_test = df_test.reset_index(drop=True)
y_train = df_train.price.values
y_val = df_val.price.values
y_test = df_test.price.values
del df_train['price']
del df_val['price']
del df_test['price']
# ### Ridge Regression
train_dict = df_train[cat + num].to_dict(orient='records')
# +
dv = DictVectorizer(sparse=False)
dv.fit(train_dict)
X_train = dv.transform(train_dict)
val_dict = df_val[cat + num].to_dict(orient='records')
X_val = dv.transform(val_dict)
# -
from sklearn.linear_model import Ridge
from sklearn.metrics import mean_squared_error
for a in [0, 0.01, 0.1, 1, 10]:
model = Ridge(alpha=a,random_state=42)
model.fit(X_train, y_train)
y_pred = model.predict(X_val)
score = np.sqrt(mean_squared_error(y_val, y_pred))
print(a, round(score, 3))
# #### Answer for Question №6: 0
| 03_ML_classification/homework_solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# Run the command below if necessary, for example with Google Colab
# #!python3 -m pip install mxnet-cu110
# +
# Global Libs
import matplotlib.pyplot as plt
import mxnet as mx
import numpy as np
import pickle
import random
# Local libs
import model
# -
with open("losses.pkl", "rb") as f:
training_loss, validation_loss = pickle.load(f)
# +
# plot the losses
epochs = 200
plt.style.use("ggplot")
plt.figure()
plt.plot(np.arange(0, epochs), validation_loss[:epochs], label="Validation Loss")
plt.plot(np.arange(0, epochs), training_loss[:epochs], label="Training Loss")
plt.xlabel("Epoch")
plt.ylabel("Loss")
plt.legend(loc="upper right")
plt.title("Losses")
plt.show()
# +
# Qualitative Evaluation
# Load Best Model
model_file_name = "net.params"
ctx = mx.gpu()
net = model.create_regression_network()
net.load_parameters(model_file_name, ctx=ctx)
# +
# Loading data for evaluation
# Saving data to analyse further
with open("data.pkl", "rb") as f:
X_train, y_train, X_val, y_val, X_test, y_test = pickle.load(f)
with open("scaled_data.pkl", "rb") as f:
(scaled_X_train_onehot_df, scaled_X_val_onehot_df, scaled_X_test_onehot_df, sc_X,
scaled_y_train, scaled_y_val, scaled_y_test, sc_y) = pickle.load(f)
# +
random_index = random.randrange(0, len(X_test))
# scaled_input = mx.nd.array([scaled_X_test[random_index]])
scaled_input = mx.nd.array([scaled_X_train_onehot_df.values[random_index]])
# Unscaled Expected Output
expected_output = y_test[random_index]
print("Unscaled Expected Output:", expected_output)
# Scaled Expected Output
scaled_expected_output = scaled_y_test[random_index]
print("Scaled Expected Output:", scaled_expected_output)
# Model Output (scaled)
output = net(scaled_input.as_in_context(ctx)).asnumpy()[0]
print("Model Output (scaled):", output)
# Unscaled Output
unscaled_output = sc_y.inverse_transform(output)
print("Unscaled Output:", unscaled_output)
# Absolute Error
abs_error = abs(expected_output - unscaled_output)
print("Absolute error: ", abs_error)
# Percentage Error
perc_error = abs_error / expected_output * 100.0
print("Percentage Error: ", perc_error)
# +
# Quantitative Evaluation
# Evaluating on Test Set (unseend data)
expected_outputs = y_test
# Model Outputs (common for all quantitative evaluations)
inputs = mx.nd.array(scaled_X_test_onehot_df.values, ctx=ctx)
outputs = net(inputs)
# Transform to numpy (CPU)
outputs = outputs.asnumpy()
unscaled_outputs = sc_y.inverse_transform(outputs)
# +
# Mean Absolute Error
abs_error = abs(expected_outputs - unscaled_outputs)
mae = sum(abs_error) / len(abs_error)
print("Mean Absolute Error (MAE):", mae)
# +
# Mean Absolute Percentage Error
perc_error = abs_error / expected_outputs * 100.0
mape = sum(perc_error) / len(perc_error)
print("Mean Absolute Percentage Error (MAPE):", mape)
# +
# Thresholds and Percentage
# How many houses in our dataset have been correctly estimated?
# We deem an estimation to be correct, when the error in the value is less than 20%
perc_threshold = 25.0
houses_in_threshold = (perc_error <= perc_threshold)
number_houses_in_threshold = sum(houses_in_threshold)
number_houses_in_threshold_perc = 100.0 * number_houses_in_threshold / len(perc_error)
print("% Houses with a predicted price error below", str(perc_threshold), "%:", number_houses_in_threshold_perc)
# +
# Calculate the % of houses depending on the threshold
perc_thresholds = np.arange(0, 100, 0.1)
houses_percentages = []
for perc_threshold in perc_thresholds:
houses_in_threshold = (perc_error <= perc_threshold)
number_houses_in_threshold = sum(houses_in_threshold)
number_houses_in_threshold_perc = 100.0 * number_houses_in_threshold / len(perc_error)
houses_percentages.append(number_houses_in_threshold_perc)
# -
# plot the graph
plt.style.use("ggplot")
plt.figure()
plt.plot(perc_thresholds, houses_percentages)
plt.xlabel("Error in the estimation")
plt.ylabel("Percentage of houses within error")
plt.title("Percentage of Correct Estimations")
plt.show()
| ch03/3_4_Evaluating_Regression_Models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# **[MLE-01]** モジュールをインポートします。
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
from numpy.random import multivariate_normal, permutation
import pandas as pd
from pandas import DataFrame, Series
# **[MLE-02]** トレーニングセットのデータを用意します。
# +
np.random.seed(20160512)
n0, mu0, variance0 = 20, [10, 11], 20
data0 = multivariate_normal(mu0, np.eye(2)*variance0 ,n0)
df0 = DataFrame(data0, columns=['x1','x2'])
df0['t'] = 0
n1, mu1, variance1 = 15, [18, 20], 22
data1 = multivariate_normal(mu1, np.eye(2)*variance1 ,n1)
df1 = DataFrame(data1, columns=['x1','x2'])
df1['t'] = 1
df = pd.concat([df0, df1], ignore_index=True)
train_set = df.reindex(permutation(df.index)).reset_index(drop=True)
# -
# **[MLE-03]** トレーニングセットのデータの内容を確認します。
train_set
# **[MLE-04]** (x1, x2) と t を別々に集めたものをNumPyのarrayオブジェクトとして取り出しておきます。
train_x = train_set[['x1','x2']].as_matrix()
train_t = train_set['t'].as_matrix().reshape([len(train_set), 1])
# **[MLE-05]** トレーニングセットのデータについて、t=1 である確率を求める計算式 p を用意します。
x = tf.placeholder(tf.float32, [None, 2])
w = tf.Variable(tf.zeros([2, 1]))
w0 = tf.Variable(tf.zeros([1]))
f = tf.matmul(x, w) + w0
p = tf.sigmoid(f)
# **[MLE-06]** 誤差関数 loss とトレーニングアルゴリズム train_step を定義します。
t = tf.placeholder(tf.float32, [None, 1])
loss = -tf.reduce_sum(t*tf.log(p) + (1-t)*tf.log(1-p))
train_step = tf.train.AdamOptimizer().minimize(loss)
# **[MLE-07]** 正解率 accuracy を定義します。
correct_prediction = tf.equal(tf.sign(p-0.5), tf.sign(t-0.5))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
# **[MLE-08]** セッションを用意して、Variableを初期化します。
sess = tf.Session()
sess.run(tf.initialize_all_variables())
# **[MLE-09]** 勾配降下法によるパラメーターの最適化を20000回繰り返します。
i = 0
for _ in range(20000):
i += 1
sess.run(train_step, feed_dict={x:train_x, t:train_t})
if i % 2000 == 0:
loss_val, acc_val = sess.run(
[loss, accuracy], feed_dict={x:train_x, t:train_t})
print ('Step: %d, Loss: %f, Accuracy: %f'
% (i, loss_val, acc_val))
# **[MLE-10]** この時点のパラメーターの値を取り出します。
w0_val, w_val = sess.run([w0, w])
w0_val, w1_val, w2_val = w0_val[0], w_val[0][0], w_val[1][0]
print w0_val, w1_val, w2_val
# **[MLE-11]** 取り出したパラメーターの値を用いて、結果をグラフに表示します。
# +
train_set0 = train_set[train_set['t']==0]
train_set1 = train_set[train_set['t']==1]
fig = plt.figure(figsize=(6,6))
subplot = fig.add_subplot(1,1,1)
subplot.set_ylim([0,30])
subplot.set_xlim([0,30])
subplot.scatter(train_set1.x1, train_set1.x2, marker='x')
subplot.scatter(train_set0.x1, train_set0.x2, marker='o')
linex = np.linspace(0,30,10)
liney = - (w1_val*linex/w2_val + w0_val/w2_val)
subplot.plot(linex, liney)
field = [[(1 / (1 + np.exp(-(w0_val + w1_val*x1 + w2_val*x2))))
for x1 in np.linspace(0,30,100)]
for x2 in np.linspace(0,30,100)]
subplot.imshow(field, origin='lower', extent=(0,30,0,30),
cmap=plt.cm.gray_r, alpha=0.5)
# -
| Chapter02/Maximum likelihood estimation example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: DESI master
# language: python
# name: desi-master
# ---
# # Spectrum and Cutout Plotting
#
# This is a basic notebook that will plot spectra, redrock fits, and Legacy Survey cutouts in a nice way given the following:
# - TARGETID
# - DATE
# - TILE
# +
import os
from glob import glob
from astropy.io import ascii, fits
from astropy.table import join, hstack, vstack, unique, Table
from desispec.spectra import stack as specstack
from desispec.io import read_spectra, write_spectra
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
# -
mpl.rc('font', size=18)
mpl.rc('axes', titlesize='small')
# ## Extract Object Redshifts
#
# Specify the TARGETID, date, tile, and version of the spectroscopic reduction desired.
#
# Note that `targetid` can be a list.
# +
redux = 'daily/tiles/cumulative'
# tile = 20750
# date = 20210521
# targetid = 39632941629837556
tile = 21202
date = 20210514
targetid = 39632956406369645
# +
folder = '{}/{}/{}/{}'.format(os.environ['DESI_SPECTRO_REDUX'], redux, tile, date)
coadds = sorted(glob('{}/coadd-*.fits'.format(folder)))
myspec = None
for i, coadd in enumerate(coadds):
zbfile = coadd.replace('coadd', 'zbest')
if os.path.exists(zbfile) and os.path.exists(coadd):
spectra = read_spectra(coadd)
sselect = np.in1d(spectra.fibermap['TARGETID'], targetid)
if np.any(sselect):
spectra = spectra[sselect]
if not hasattr(spectra, 'scores_comments'):
spectra.scores_comments = None
# Select matching target IDs from the list and access the redshift fit.
zbest = fits.open(zbfile)['ZBEST'].data
zselect = np.in1d(zbest['TARGETID'], targetid)
zbest = zbest[zselect]
# Append spectra to a larger list of spectra, stored in memory.
# Note that an EXPID is required in the update step below, but the coadded spectra
# have a FIRST_EXPID and a LAST_EXPID. So copy one of these as a hack.
spectra.fibermap['EXPID'] = spectra.fibermap['LAST_EXPID']
spectra.extra_catalog = zbest
myspec = spectra
break
# -
myspec.num_spectra()
# ### Output Redrock Spectral Types
types_, counts_ = np.unique(myspec.extra_catalog['SPECTYPE'], return_counts=True)
for t, c in zip(types_, counts_):
print('{:10s} : {:10d} ({:.1f}%)'.format(t, c, 100*c/len(myspec.extra_catalog)))
# ## Plot Spectra
# ### Grab Legacy Survey Cutouts
#
# Some code to access legacy survey cutouts.
# +
import requests
def get_cutout(targetid, ra, dec, verbose=False):
"""Grab and cache legacy survey cutouts.
Parameters
----------
targetid : int
DESI target ID.
ra : float
Right ascension (degrees).
dec : float
Declination (degrees).
verbose : bool
Add some status messages if true.
Returns
-------
img_name : str
Name of JPG cutout file written after query.
"""
img_name = '{}.jpg'.format(targetid)
if os.path.exists(img_name):
if verbose:
print('{} exists.'.format(img_name))
else:
if verbose:
print('Accessing {}'.format(img_name))
img_url = 'https://www.legacysurvey.org/viewer/cutout.jpg?ra={}&dec={}&%22/pix=0.25&layer=dr8&size=180'.format(ra, dec)
with open(img_name, 'wb') as handle:
response = requests.get(img_url, stream=True)
if not response.ok:
print(response)
for block in response.iter_content(1024):
if not block:
break
handle.write(block)
return img_name
# -
# #### Spectrum Plot Options
#
# Pull in a basic Gaussian smoothing filter and some code for plotting the same emission and absorption features used in Prospect.
# +
from scipy.ndimage import gaussian_filter1d
mpl.rc('figure', max_open_warning = 0)
# +
# Emission and absorption lines from Prospect tables.
emi_lines = ascii.read('emission_lines.txt', comment='#', names=['name','longname','lambda','vacuum','major'])
abs_lines = ascii.read('absorption_lines.txt', comment='#', names=['name','longname','lambda','vacuum','major'])
emi_lines_major = emi_lines[emi_lines['major']=='True']
abs_lines_major = abs_lines[abs_lines['major']=='True']
# +
for i in range(myspec.num_spectra()):
fig, axes = plt.subplots(1,2, figsize=(16,5), gridspec_kw={'width_ratios':[3,1.1]},
tight_layout=True)
# Plot the spectra.
ax = axes[0]
fmin, fmax = 1e99, -1e99
for band in 'brz':
smoothed = gaussian_filter1d(myspec.flux[band][i], 5)
fmin = np.minimum(fmin, np.min(smoothed))
fmax = np.maximum(fmax, np.max(smoothed))
ax.plot(myspec.wave[band], smoothed)
zbest = myspec.extra_catalog[i]
z = zbest['Z']
dchi2 = zbest['DELTACHI2']
zwarn = zbest['ZWARN']
sptype = zbest['SPECTYPE']
print(dchi2, zwarn)
for eline in emi_lines:
wl = eline['lambda']*(1 + z)
if wl > 3600 and wl < 9800:
ax.axvline(wl, ls='--', color='k', alpha=0.3)
ax.text(wl+20, fmin, eline['name'], fontsize=8, rotation=90, alpha=0.3)
for aline in abs_lines:
wl = aline['lambda']*(1 + z)
if wl > 3600 and wl < 9800:
ax.axvline(wl, ls='--', color='r', alpha=0.3)
ax.text(wl+20, 0.95*fmax, aline['name'], color='r', fontsize=8, rotation=90, alpha=0.3)
ax.set(xlabel=r'$\lambda_{\mathrm{obs}}$ [$\AA$]',
xlim=(3500,9900),
ylabel=r'flux [erg s$^{-1}$ cm$^{-2}$ $\AA^{-1}$]',
title=r'{}; $z={:.3f}$ ($\Delta\chi^2={:.5g}$; ZWARN=0x{:x}; SPECTYPE={:s})'.format(myspec.fibermap[i]['TARGETID'], z, dchi2, zwarn, sptype),
)
# Plot the image cutout.
ax = axes[1]
obj = myspec.fibermap[i]
img_file = get_cutout(obj['TARGETID'], obj['TARGET_RA'], obj['TARGET_DEC'])
img = mpl.image.imread(img_file)
ax.imshow(img)
x1, x2, x3, x4 = [90, 90], [70, 80], [90, 90], [100,110]
y1, y2, y3, y4 = [70, 80], [90, 90], [100,110], [90,90]
ax.plot(x1, y1, x2, y2, x3, y3, x4, y4, color='r', linewidth=2, alpha=0.7)
ax.text(5,15, '{:3s} = {}\n{:3s} = {}'.format('RA', obj['TARGET_RA'], 'Dec', obj['TARGET_DEC']), color='yellow', fontsize=9)
ax.set(aspect='equal',
title='{}, Tile {}, Exp {}'.format(obj['LAST_NIGHT'], obj['TILEID'], obj['EXPID']))
ax.axis('off')
fig.savefig('spec_cutout_{}_{}_{:06d}_{:06d}.png'.format(obj['TARGETID'], obj['LAST_NIGHT'], obj['TILEID'], obj['EXPID']), dpi=100)
# # Clean up
# fig.clear()
# plt.close(fig)
# -
# ### Rest Frame Plot
# +
for i in range(myspec.num_spectra()):
fig, axes = plt.subplots(1,2, figsize=(14.5,5), gridspec_kw={'width_ratios':[2.75,1.2]},
tight_layout=True)
# Plot the spectra.
ax = axes[0]
zbest = myspec.extra_catalog[i]
z = zbest['Z']
dchi2 = zbest['DELTACHI2']
zwarn = zbest['ZWARN']
sptype = zbest['SPECTYPE']
print(dchi2, zwarn)
fmin, fmax = 1e99, -1e99
for band in 'brz':
smoothed = gaussian_filter1d(myspec.flux[band][i], 5)
fmin = np.minimum(fmin, np.min(smoothed))
fmax = np.maximum(fmax, np.max(smoothed))
ax.plot(myspec.wave[band]/(1 + z), smoothed)
fmax = 7.
for eline in emi_lines:
wl = eline['lambda']
if wl > 3600/(1+z) and wl < 9800/(1+z):
ax.axvline(wl, ls='--', color='k', alpha=0.3)
ax.text(wl+20, -0.5, eline['name'], fontsize=8, rotation=90, alpha=0.3)
for aline in abs_lines:
wl = aline['lambda']
if wl > 3600/(1+z) and wl < 9800/(1+z):
ax.axvline(wl, ls='--', color='r', alpha=0.3)
ax.text(wl+20, 0.95*fmax, aline['name'], color='r', fontsize=8, rotation=90, alpha=0.3)
ax.set(xlabel=r'$\lambda_{\mathrm{rest}}$ [$\AA$]',
xlim=(3600/(1 + z),9800/(1 + z)),
ylabel=r'flux [erg s$^{-1}$ cm$^{-2}$ $\AA^{-1}$]',
ylim=(-1, fmax),
title=r'{}; $z={:.3f}$ ($\Delta\chi^2={:.5g}$; ZWARN=0x{:x}; SPECTYPE={:s})'.format(myspec.fibermap[i]['TARGETID'], z, dchi2, zwarn, sptype),
)
# Plot the image cutout.
ax = axes[1]
obj = myspec.fibermap[i]
img_file = get_cutout(obj['TARGETID'], obj['TARGET_RA'], obj['TARGET_DEC'])
img = mpl.image.imread(img_file)
ax.imshow(img)
x1, x2, x3, x4 = [90, 90], [70, 80], [90, 90], [100,110]
y1, y2, y3, y4 = [70, 80], [90, 90], [100,110], [90,90]
ax.plot(x1, y1, x2, y2, x3, y3, x4, y4, color='r', linewidth=2, alpha=0.7)
ax.text(5,15, '{:3s} = {}\n{:3s} = {}'.format('RA', obj['TARGET_RA'], 'Dec', obj['TARGET_DEC']), color='yellow', fontsize=9)
ax.set(aspect='equal',
title='{}, Tile {}, Exp {}'.format(obj['LAST_NIGHT'], obj['TILEID'], obj['EXPID']))
ax.axis('off')
fig.savefig('spec_cutout_{}_{}_{:06d}_{:06d}.png'.format(obj['TARGETID'], obj['LAST_NIGHT'], obj['TILEID'], obj['EXPID']), dpi=120)
fig.savefig('spec_cutout_{}_{}_{:06d}_{:06d}.pdf'.format(obj['TARGETID'], obj['LAST_NIGHT'], obj['TILEID'], obj['EXPID']))
# # Clean up
# fig.clear()
# plt.close(fig)
# -
# ## Compute and Subtract Redrock Model
#
# Extract the redrock templates and use the best fit coefficients to plot and remove the best-fit model from the data.
# +
import redrock.templates
templates = dict()
for f in redrock.templates.find_templates():
t = redrock.templates.Template(f)
templates[(t.template_type, t.sub_type)] = t
# -
zbest = myspec.extra_catalog[0]
z = zbest['Z']
sptype = zbest['SPECTYPE']
sbtype = zbest['SUBTYPE']
fulltype = (sptype, sbtype)
ncoeff = templates[fulltype].flux.shape[0]
coeff = zbest['COEFF'][0:ncoeff]
# +
from desispec.interpolation import resample_flux
from desispec.resolution import Resolution
tflux = templates[fulltype].flux.T.dot(coeff)
twave = templates[fulltype].wave
plt.plot(twave, tflux)
# +
for i in range(myspec.num_spectra()):
fig, axes = plt.subplots(2,1, figsize=(10,10), sharex=True, tight_layout=True)
# Plot the spectra.
ax = axes[0]
zbest = myspec.extra_catalog[i]
z = zbest['Z']
dchi2 = zbest['DELTACHI2']
zwarn = zbest['ZWARN']
sptype = zbest['SPECTYPE']
print(dchi2, zwarn)
sptype = zbest['SPECTYPE']
sbtype = zbest['SUBTYPE']
fulltype = (sptype, sbtype)
ncoeff = templates[fulltype].flux.shape[0]
coeff = zbest['COEFF'][0:ncoeff]
tflux = templates[fulltype].flux.T.dot(coeff)
twave = templates[fulltype].wave*(1+z)
fmin, fmax = 1e99, -1e99
for band in 'brz':
smoothed = gaussian_filter1d(myspec.flux[band][i], 5)
fmin = np.minimum(fmin, np.min(smoothed))
fmax = np.maximum(fmax, np.max(smoothed))
ax.plot(myspec.wave[band]/(1 + z), smoothed)
R = Resolution(myspec.resolution_data[band][i])
txflux = R.dot(resample_flux(myspec.wave[band], twave, tflux))
smoothed = gaussian_filter1d(txflux, 5)
ax.plot(myspec.wave[band]/(1+z), smoothed, color='k', lw=1, ls='--')
residual = myspec.flux[band][i] - txflux
smoothed = gaussian_filter1d(residual, 5)
axes[1].plot(myspec.wave[band]/(1+z), smoothed)
fmax = 7.
for eline in emi_lines:
wl = eline['lambda']
if wl > 3600/(1+z) and wl < 9800/(1+z):
ax.axvline(wl, ls='--', color='k', alpha=0.3)
ax.text(wl+20, -0.5, eline['name'], fontsize=8, rotation=90, alpha=0.3)
for aline in abs_lines:
wl = aline['lambda']
if wl > 3600/(1+z) and wl < 9800/(1+z):
ax.axvline(wl, ls='--', color='r', alpha=0.3)
ax.text(wl+20, 0.95*fmax, aline['name'], color='r', fontsize=8, rotation=90, alpha=0.3)
ax.set(#xlabel=r'$\lambda_{\mathrm{rest}}$ [$\AA$]',
xlim=(3600/(1 + z),9800/(1 + z)),
ylabel=r'flux [erg s$^{-1}$ cm$^{-2}$ $\AA^{-1}$]',
ylim=(-1, fmax),
title=r'{}; $z={:.3f}$ ($\Delta\chi^2={:.5g}$; ZWARN=0x{:x}; SPECTYPE={:s})'.format(myspec.fibermap[i]['TARGETID'], z, dchi2, zwarn, sptype),
)
ax = axes[1]
ax.set(xlabel=r'$\lambda_{\mathrm{rest}}$ [$\AA$]',
ylabel=r'residual [erg s$^{-1}$ cm$^{-2}$ $\AA^{-1}$]',
ylim=(-4,4))
ax.grid(ls=':')
fig.savefig('spec_residual_{}_{}_{:06d}_{:06d}.png'.format(obj['TARGETID'], obj['LAST_NIGHT'], obj['TILEID'], obj['EXPID']), dpi=120)
# fig.savefig('spec_nocutout_{}_{}_{:06d}_{:06d}.pdf'.format(obj['TARGETID'], obj['LAST_NIGHT'], obj['TILEID'], obj['EXPID']))
# # Clean up
# fig.clear()
# plt.close(fig)
# -
| db/plot_spectra_cutouts.ipynb |
#!/usr/bin/env python
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The Transformation and Accept/Reject (aka. Von Neumann) methods
# Python notebook with exercise on how to generate random numbers following a specific PDF using uniformly distributed random numbers, in this case a third degree polynomial: $f(x) = x^3$, in the interval $[0, C]$ where $C$ is a number that ensures that the function if properly normalized.
#
# Both the Accept-Reject (Von Neumann) and transformation method should be considered for the problem:
# - Transformation method (if function can be integrated and then inverted).
# - Accept-Reject (or Hit & Miss) method (by <NAME> and <NAME>).
#
# ### References:
# - <NAME>: Chapter 3
# - <NAME>: page 81-84
# - __[40. Monte Carlo Techniques PDG](http://pdg.lbl.gov/2020/reviews/rpp2020-rev-monte-carlo-techniques.pdf)__
#
# ### Authors:
# - <NAME> (Niels Bohr Institute)
#
# ### Date:
# - 29-11-2021 (latest update)
#
# ***
import numpy as np # Matlab like syntax for linear algebra and functions
import matplotlib.pyplot as plt # Plots and figures like you know them from Matlab
import seaborn as sns # Make the plots nicer to look at
from iminuit import Minuit # The actual fitting tool, better than scipy's
import sys # Modules to see files and folders in directories
from scipy import stats
# Load external functions:
# +
sys.path.append('../../../External_Functions')
from ExternalFunctions import Chi2Regression, BinnedLH, UnbinnedLH
from ExternalFunctions import nice_string_output, add_text_to_ax # Useful functions to print fit results on figure
plt.rcParams['font.size'] = 18 # Set some basic plotting parameters
# -
# Set the parameters of the program:
# +
r = np.random
r.seed(42)
save_plots = False
N_points = 10000 # Number of random points to be generated
# -
# ## Problem 1:
#
# #### Produce random points following $f(x) \sim \exp(-x/3), x \in [0, \infty]$.
# +
# Define a reasonable range to plot in:
xmin = 0
xmax = 20
N_bins = 100
x_exp = 20.0*r.uniform(size=N_points) # This is NOT the solution - just something to plot!
# -
# And to make sure that we have computed the points correctly, we plot a histogram of the data:
fig, ax = plt.subplots(figsize=(10, 6))
ax.hist(x_exp, bins=100, range=(xmin, xmax), histtype='step', label='histogram' )
ax.set(xlabel="x_exp", ylabel="Frequency", xlim=(xmin-1.0, xmax+1.0));
# ## Problem 2:
#
# #### Produce random points following $g(x) \sim x \cos(x), x \in [0, \frac{\pi}{2}]$.
xmin2, xmax2 = 0, np.pi/2 # Note the simple/short notation.
# ## Problem 3:
#
# #### Produce random points following $h(x) \sim \exp(-x/3)\cos(x)^2$ in the interval $[0, \infty]$ and estimate the integral of $h(x)$ in the defined range.
| AppStat2022/Week3/original/TransformationAcceptReject/TransformationAcceptReject_general.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] origin_pos=0
# # Layers and Blocks
# :label:`sec_model_construction`
#
# When we first introduced neural networks,
# we focused on linear models with a single output.
# Here, the entire model consists of just a single neuron.
# Note that a single neuron
# (i) takes some set of inputs;
# (ii) generates a corresponding scalar output;
# and (iii) has a set of associated parameters that can be updated
# to optimize some objective function of interest.
# Then, once we started thinking about networks with multiple outputs,
# we leveraged vectorized arithmetic
# to characterize an entire layer of neurons.
# Just like individual neurons,
# layers (i) take a set of inputs,
# (ii) generate corresponding outputs,
# and (iii) are described by a set of tunable parameters.
# When we worked through softmax regression,
# a single layer was itself the model.
# However, even when we subsequently
# introduced MLPs,
# we could still think of the model as
# retaining this same basic structure.
#
# Interestingly, for MLPs,
# both the entire model and its constituent layers
# share this structure.
# The entire model takes in raw inputs (the features),
# generates outputs (the predictions),
# and possesses parameters
# (the combined parameters from all constituent layers).
# Likewise, each individual layer ingests inputs
# (supplied by the previous layer)
# generates outputs (the inputs to the subsequent layer),
# and possesses a set of tunable parameters that are updated
# according to the signal that flows backwards
# from the subsequent layer.
#
#
# While you might think that neurons, layers, and models
# give us enough abstractions to go about our business,
# it turns out that we often find it convenient
# to speak about components that are
# larger than an individual layer
# but smaller than the entire model.
# For example, the ResNet-152 architecture,
# which is wildly popular in computer vision,
# possesses hundreds of layers.
# These layers consist of repeating patterns of *groups of layers*. Implementing such a network one layer at a time can grow tedious.
# This concern is not just hypothetical---such
# design patterns are common in practice.
# The ResNet architecture mentioned above
# won the 2015 ImageNet and COCO computer vision competitions
# for both recognition and detection :cite:`He.Zhang.Ren.ea.2016`
# and remains a go-to architecture for many vision tasks.
# Similar architectures in which layers are arranged
# in various repeating patterns
# are now ubiquitous in other domains,
# including natural language processing and speech.
#
# To implement these complex networks,
# we introduce the concept of a neural network *block*.
# A block could describe a single layer,
# a component consisting of multiple layers,
# or the entire model itself!
# One benefit of working with the block abstraction
# is that they can be combined into larger artifacts,
# often recursively. This is illustrated in :numref:`fig_blocks`. By defining code to generate blocks
# of arbitrary complexity on demand,
# we can write surprisingly compact code
# and still implement complex neural networks.
#
# 
# :label:`fig_blocks`
#
#
# From a programing standpoint, a block is represented by a *class*.
# Any subclass of it must define a forward propagation function
# that transforms its input into output
# and must store any necessary parameters.
# Note that some blocks do not require any parameters at all.
# Finally a block must possess a backpropagation function,
# for purposes of calculating gradients.
# Fortunately, due to some behind-the-scenes magic
# supplied by the auto differentiation
# (introduced in :numref:`sec_autograd`)
# when defining our own block,
# we only need to worry about parameters
# and the forward propagation function.
#
# [**To begin, we revisit the code
# that we used to implement MLPs**]
# (:numref:`sec_mlp_concise`).
# The following code generates a network
# with one fully-connected hidden layer
# with 256 units and ReLU activation,
# followed by a fully-connected output layer
# with 10 units (no activation function).
#
# + origin_pos=3 tab=["tensorflow"]
import tensorflow as tf
net = tf.keras.models.Sequential([
tf.keras.layers.Dense(256, activation=tf.nn.relu),
tf.keras.layers.Dense(10),
])
X = tf.random.uniform((2, 20))
net(X)
# + [markdown] origin_pos=6 tab=["tensorflow"]
# In this example, we constructed
# our model by instantiating an `keras.models.Sequential`, with layers in the order
# that they should be executed passed as arguments.
# In short, `Sequential` defines a special kind of `keras.Model`,
# the class that presents a block in Keras.
# It maintains an ordered list of constituent `Model`s.
# Note that each of the two fully-connected layers is an instance of the `Dense` class
# which is itself a subclass of `Model`.
# The forward propagation (`call`) function is also remarkably simple:
# it chains each block in the list together,
# passing the output of each as the input to the next.
# Note that until now, we have been invoking our models
# via the construction `net(X)` to obtain their outputs.
# This is actually just shorthand for `net.call(X)`,
# a slick Python trick achieved via
# the Block class's `__call__` function.
#
# + [markdown] origin_pos=7
# ## [**A Custom Block**]
#
# Perhaps the easiest way to develop intuition
# about how a block works
# is to implement one ourselves.
# Before we implement our own custom block,
# we briefly summarize the basic functionality
# that each block must provide:
#
# + [markdown] origin_pos=8 tab=["tensorflow"]
# 1. Ingest input data as arguments to its forward propagation function.
# 1. Generate an output by having the forward propagation function return a value. Note that the output may have a different shape from the input. For example, the first fully-connected layer in our model above ingests an input of arbitrary dimension but returns an output of dimension 256.
# 1. Calculate the gradient of its output with respect to its input, which can be accessed via its backpropagation function. Typically this happens automatically.
# 1. Store and provide access to those parameters necessary
# to execute the forward propagation computation.
# 1. Initialize model parameters as needed.
#
# + [markdown] origin_pos=10
# In the following snippet,
# we code up a block from scratch
# corresponding to an MLP
# with one hidden layer with 256 hidden units,
# and a 10-dimensional output layer.
# Note that the `MLP` class below inherits the class that represents a block.
# We will heavily rely on the parent class's functions,
# supplying only our own constructor (the `__init__` function in Python) and the forward propagation function.
#
# + origin_pos=13 tab=["tensorflow"]
class MLP(tf.keras.Model):
# Declare a layer with model parameters. Here, we declare two fully
# connected layers
def __init__(self):
# Call the constructor of the `MLP` parent class `Model` to perform
# the necessary initialization. In this way, other function arguments
# can also be specified during class instantiation, such as the model
# parameters, `params` (to be described later)
super().__init__()
# Hidden layer
self.hidden = tf.keras.layers.Dense(units=256, activation=tf.nn.relu)
self.out = tf.keras.layers.Dense(units=10) # Output layer
# Define the forward propagation of the model, that is, how to return the
# required model output based on the input `X`
def call(self, X):
return self.out(self.hidden((X)))
# + [markdown] origin_pos=14
# Let us first focus on the forward propagation function.
# Note that it takes `X` as the input,
# calculates the hidden representation
# with the activation function applied,
# and outputs its logits.
# In this `MLP` implementation,
# both layers are instance variables.
# To see why this is reasonable, imagine
# instantiating two MLPs, `net1` and `net2`,
# and training them on different data.
# Naturally, we would expect them
# to represent two different learned models.
#
# We [**instantiate the MLP's layers**]
# in the constructor
# (**and subsequently invoke these layers**)
# on each call to the forward propagation function.
# Note a few key details.
# First, our customized `__init__` function
# invokes the parent class's `__init__` function
# via `super().__init__()`
# sparing us the pain of restating
# boilerplate code applicable to most blocks.
# We then instantiate our two fully-connected layers,
# assigning them to `self.hidden` and `self.out`.
# Note that unless we implement a new operator,
# we need not worry about the backpropagation function
# or parameter initialization.
# The system will generate these functions automatically.
# Let us try this out.
#
# + origin_pos=17 tab=["tensorflow"]
net = MLP()
net(X)
# + [markdown] origin_pos=18
# A key virtue of the block abstraction is its versatility.
# We can subclass a block to create layers
# (such as the fully-connected layer class),
# entire models (such as the `MLP` class above),
# or various components of intermediate complexity.
# We exploit this versatility
# throughout the following chapters,
# such as when addressing
# convolutional neural networks.
#
#
# ## [**The Sequential Block**]
#
# We can now take a closer look
# at how the `Sequential` class works.
# Recall that `Sequential` was designed
# to daisy-chain other blocks together.
# To build our own simplified `MySequential`,
# we just need to define two key function:
# 1. A function to append blocks one by one to a list.
# 2. A forward propagation function to pass an input through the chain of blocks, in the same order as they were appended.
#
# The following `MySequential` class delivers the same
# functionality of the default `Sequential` class.
#
# + origin_pos=21 tab=["tensorflow"]
class MySequential(tf.keras.Model):
def __init__(self, *args):
super().__init__()
self.modules = []
for block in args:
# Here, `block` is an instance of a `tf.keras.layers.Layer`
# subclass
self.modules.append(block)
def call(self, X):
for module in self.modules:
X = module(X)
return X
# + [markdown] origin_pos=24
# When our `MySequential`'s forward propagation function is invoked,
# each added block is executed
# in the order in which they were added.
# We can now reimplement an MLP
# using our `MySequential` class.
#
# + origin_pos=27 tab=["tensorflow"]
net = MySequential(
tf.keras.layers.Dense(units=256, activation=tf.nn.relu),
tf.keras.layers.Dense(10))
net(X)
# + [markdown] origin_pos=28
# Note that this use of `MySequential`
# is identical to the code we previously wrote
# for the `Sequential` class
# (as described in :numref:`sec_mlp_concise`).
#
#
# ## [**Executing Code in the Forward Propagation Function**]
#
# The `Sequential` class makes model construction easy,
# allowing us to assemble new architectures
# without having to define our own class.
# However, not all architectures are simple daisy chains.
# When greater flexibility is required,
# we will want to define our own blocks.
# For example, we might want to execute
# Python's control flow within the forward propagation function.
# Moreover, we might want to perform
# arbitrary mathematical operations,
# not simply relying on predefined neural network layers.
#
# You might have noticed that until now,
# all of the operations in our networks
# have acted upon our network's activations
# and its parameters.
# Sometimes, however, we might want to
# incorporate terms
# that are neither the result of previous layers
# nor updatable parameters.
# We call these *constant parameters*.
# Say for example that we want a layer
# that calculates the function
# $f(\mathbf{x},\mathbf{w}) = c \cdot \mathbf{w}^\top \mathbf{x}$,
# where $\mathbf{x}$ is the input, $\mathbf{w}$ is our parameter,
# and $c$ is some specified constant
# that is not updated during optimization.
# So we implement a `FixedHiddenMLP` class as follows.
#
# + origin_pos=31 tab=["tensorflow"]
class FixedHiddenMLP(tf.keras.Model):
def __init__(self):
super().__init__()
self.flatten = tf.keras.layers.Flatten()
# Random weight parameters created with `tf.constant` are not updated
# during training (i.e., constant parameters)
self.rand_weight = tf.constant(tf.random.uniform((20, 20)))
self.dense = tf.keras.layers.Dense(20, activation=tf.nn.relu)
def call(self, inputs):
X = self.flatten(inputs)
# Use the created constant parameters, as well as the `relu` and
# `matmul` functions
X = tf.nn.relu(tf.matmul(X, self.rand_weight) + 1)
# Reuse the fully-connected layer. This is equivalent to sharing
# parameters with two fully-connected layers
X = self.dense(X)
# Control flow
while tf.reduce_sum(tf.math.abs(X)) > 1:
X /= 2
return tf.reduce_sum(X)
# + [markdown] origin_pos=32
# In this `FixedHiddenMLP` model,
# we implement a hidden layer whose weights
# (`self.rand_weight`) are initialized randomly
# at instantiation and are thereafter constant.
# This weight is not a model parameter
# and thus it is never updated by backpropagation.
# The network then passes the output of this "fixed" layer
# through a fully-connected layer.
#
# Note that before returning the output,
# our model did something unusual.
# We ran a while-loop, testing
# on the condition its $L_1$ norm is larger than $1$,
# and dividing our output vector by $2$
# until it satisfied the condition.
# Finally, we returned the sum of the entries in `X`.
# To our knowledge, no standard neural network
# performs this operation.
# Note that this particular operation may not be useful
# in any real-world task.
# Our point is only to show you how to integrate
# arbitrary code into the flow of your
# neural network computations.
#
# + origin_pos=34 tab=["tensorflow"]
net = FixedHiddenMLP()
net(X)
# + [markdown] origin_pos=35
# We can [**mix and match various
# ways of assembling blocks together.**]
# In the following example, we nest blocks
# in some creative ways.
#
# + origin_pos=38 tab=["tensorflow"]
class NestMLP(tf.keras.Model):
def __init__(self):
super().__init__()
self.net = tf.keras.Sequential()
self.net.add(tf.keras.layers.Dense(64, activation=tf.nn.relu))
self.net.add(tf.keras.layers.Dense(32, activation=tf.nn.relu))
self.dense = tf.keras.layers.Dense(16, activation=tf.nn.relu)
def call(self, inputs):
return self.dense(self.net(inputs))
chimera = tf.keras.Sequential()
chimera.add(NestMLP())
chimera.add(tf.keras.layers.Dense(20))
chimera.add(FixedHiddenMLP())
chimera(X)
# + [markdown] origin_pos=39
# ## Efficiency
#
# + [markdown] origin_pos=42 tab=["tensorflow"]
# The avid reader might start to worry
# about the efficiency of some of these operations.
# After all, we have lots of dictionary lookups,
# code execution, and lots of other Pythonic things
# taking place in what is supposed to be
# a high-performance deep learning library.
# The problems of Python's [global interpreter lock](https://wiki.python.org/moin/GlobalInterpreterLock) are well known.
# In the context of deep learning,
# we may worry that our extremely fast GPU(s)
# might have to wait until a puny CPU
# runs Python code before it gets another job to run.
# The best way to speed up Python is by avoiding it altogether.
#
# + [markdown] origin_pos=43
# ## Summary
#
# * Layers are blocks.
# * Many layers can comprise a block.
# * Many blocks can comprise a block.
# * A block can contain code.
# * Blocks take care of lots of housekeeping, including parameter initialization and backpropagation.
# * Sequential concatenations of layers and blocks are handled by the `Sequential` block.
#
#
# ## Exercises
#
# 1. What kinds of problems will occur if you change `MySequential` to store blocks in a Python list?
# 1. Implement a block that takes two blocks as an argument, say `net1` and `net2` and returns the concatenated output of both networks in the forward propagation. This is also called a parallel block.
# 1. Assume that you want to concatenate multiple instances of the same network. Implement a factory function that generates multiple instances of the same block and build a larger network from it.
#
# + [markdown] origin_pos=46 tab=["tensorflow"]
# [Discussions](https://discuss.d2l.ai/t/264)
#
| d2l/tensorflow/chapter_deep-learning-computation/model-construction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# +
#@title Imports, initial setup (Ctrl+F9 to run all)
try:
import gamry_parser
except:
subprocess.run(
["pip", "install", "gamry-parser"],
encoding="utf-8",
shell=False)
finally:
import gamry_parser
import os
import re
import pandas as pd
import matplotlib.pyplot as plt
z = gamry_parser.Impedance()
print('Done.')
# + id="7f1iTOecIISA" cellView="form"
"""
### SCRIPT CONFIGURATION SETTINGS ###
"""
#@markdown **Load data** Parse Gamry *.DTA files from folder
#@markdown Where are the Gamry DTA files located?
file_path = "/path/to/gamry/files/" #@param {type:"string"}
#@markdown Which of the DTA files do we want to compare? (Regular expression matching)
file_pattern = "EIS" #@param {type:"string"}
#@markdown Which impedance frequencies should be shown? (separated by comma, e.g. `4, 1000, 10000`)
frequencies_to_show = "1, 5, 10, 10000" #@param {type:"string"}
frequencies_to_show = [int(val.strip()) for val in frequencies_to_show.split(",")]
files = [f for f in os.listdir(file_path) if
os.path.splitext(f)[1].lower() == ".dta" and
len(re.findall(file_pattern, f)) > 0
]
# For repeating EIS, we need to properly sort files -- by chronological run-order instead of alphanumeric filename.
run_pattern = re.compile("[0-9]+_Run[0-9]+\.DTA", re.IGNORECASE)
files.sort(key=lambda fname: "_".join(["".join(filter(str.isdigit, x)).zfill(4) for x in run_pattern.search(fname).group().split("_")]))
if len(files) == 0:
assert False, "No files matching the file filter [{}] were found.".format(file_pattern)
else:
print('Found [{}] data files matching [{}]'.format(len(files), file_pattern))
# store aggregated start time, magnitude, phase, real, and imaginary impedance into separate variables
start_times = []
df_mag = pd.DataFrame()
df_phz = pd.DataFrame()
df_real = pd.DataFrame()
df_imag = pd.DataFrame()
# iterate through gamry files
index = 0
for dataf in files:
name = os.path.splitext(dataf)[0].split('-')
name = ", ".join(name[1:])
# load file
f = os.path.join(file_path, dataf)
z.load(f)
# process data header metadata
header = z.get_header()
start_time = pd.Timestamp("{} {}".format(header.get("DATE"), header.get("TIME")))
print('{} [{}] ocp={}'.format(start_time, name, z.get_ocv_value()))
# extract EIS curve
res = z.get_curve_data()
start_times.append(start_time)
df_mag[name] = res['Zmod']
df_phz[name] = res['Zphz']
df_real[name] = res['Zreal']
df_imag[name] = res['Zimag']
# post-processing for all collected curves
# validate the collected data, set frequency as dataframe index
df_mag["Freq"] = res["Freq"]
df_mag.set_index("Freq", inplace=True)
df_mag.mask(df_mag < 0, inplace=True)
df_phz["Freq"] = res["Freq"]
df_phz.set_index("Freq", inplace=True)
df_phz.mask(df_phz > 0, inplace=True)
df_phz.mask(df_phz < -90, inplace=True)
df_real["Freq"] = res["Freq"]
df_real.set_index("Freq", inplace=True)
df_real.mask(df_real < 0, inplace=True)
df_imag["Freq"] = res["Freq"]
df_imag.set_index("Freq", inplace=True)
df_imag.mask(df_imag > 0, inplace=True)
df_imag = df_imag.applymap(abs)
# print to screen impedance magnitude for the desired frequency
def freq_lookup(df, freq):
return df.index.get_loc(freq, method='nearest')
for freq in frequencies_to_show:
row_index = freq_lookup(df_mag, freq)
print("\n Showing Z_mag @ {} Hz [actual={:0.2f} Hz]".format(freq, df_mag.index[row_index]))
print(df_mag.iloc[row_index])
# + id="t850lx00MlBL" cellView="form"
#@markdown **Bode Plot**: Display Zmag, Zphase vs. Freq
from plotly.subplots import make_subplots
import plotly.graph_objects as go
from plotly.colors import DEFAULT_PLOTLY_COLORS
show_legend = False #@param{type:"boolean"}
fig = make_subplots(rows=2, cols=1, shared_xaxes=True, vertical_spacing=0.02)
data = []
# Yields a tuple of column name and series for each column in the dataframe
for (index, (columnName, columnData)) in enumerate(df_mag.iteritems()):
newTrace = go.Scatter(
x=df_mag.index,
y=columnData,
mode='lines',
name=columnName,
legendgroup=columnName,
line=dict(color=DEFAULT_PLOTLY_COLORS[index % len(DEFAULT_PLOTLY_COLORS)]),
)
fig.add_trace(newTrace, row=1, col=1)
newTrace = go.Scatter(
x=df_mag.index,
y=-1*df_phz[columnName],
mode='lines',
name=columnName,
legendgroup=columnName,
line=dict(color=DEFAULT_PLOTLY_COLORS[index % len(DEFAULT_PLOTLY_COLORS)]),
showlegend=False
)
fig.add_trace(newTrace, row=2, col=1)
# variation = df_mag.std(axis=1) / newTrace['y']
# fig.add_trace({'x': df_mag.index, 'y': variation, 'name': 'Signal Variation'}, row=2, col=1)
layout = {
'title': {'text': 'Bode Plot [{}]'.format(experiment_name),
'yanchor': 'top',
'y': 0.95,
'x': 0.5 },
'xaxis': {
'anchor': 'x',
'type': 'log'
},
'xaxis2': {
'title': 'Frequency, Hz',
'type': 'log',
'matches': 'x'
},
'yaxis': {
'title': 'Magnitude, Ohm',
'type': 'log'
''
},
'yaxis2': {
'title': 'Phase, deg',
},
'legend': {'x': 0.85, 'y': 0.97},
'margin': dict(l=30, r=20, t=60, b=20),
'width': 1200,
'height': 500,
}
fig.update_layout(layout)
if not show_legend:
fig.update_layout({"showlegend": False})
config={
'displaylogo': False,
'modeBarButtonsToRemove': ['select2d', 'lasso2d', 'hoverClosestCartesian', 'toggleSpikelines','hoverCompareCartesian']
}
fig.show(config=config)
# + id="V6LKqpiECgb5" cellView="form"
#@markdown **Polar Coordinate Plot**: Display -Zimag vs. Zreal
from plotly.subplots import make_subplots
import plotly.graph_objects as go
from plotly.colors import DEFAULT_PLOTLY_COLORS
logx = True #@param {type:"boolean"}
logy = True #@param {type:"boolean"}
fig = make_subplots(rows=1, cols=1,vertical_spacing=0.02, horizontal_spacing=0.02)
data = []
# Yields a tuple of column name and series for each column in the dataframe
for (index, columnName) in enumerate(df_real.columns):
newTrace = go.Scatter(
x=df_real[columnName],
y=-df_imag[columnName],
mode='markers+lines',
name=columnName,
legendgroup=columnName,
text='Freq: ' + df_real.index.astype(str),
line=dict(color=DEFAULT_PLOTLY_COLORS[index % len(DEFAULT_PLOTLY_COLORS)]),
)
fig.add_trace(newTrace, row=1, col=1)
# variation = df_mag.std(axis=1) / newTrace['y']
# fig.add_trace({'x': df_mag.index, 'y': variation, 'name': 'Signal Variation'}, row=2, col=1)
layout = {
'title': {'text': 'Impendance Plot, Polar Coord. [tests matching: {}]'.format(file_pattern),
'yanchor': 'top',
'y': 0.95,
'x': 0.5 },
'xaxis': {
'anchor': 'x',
'title': 'Zreal, Ohm',
'type': 'log' if logx else 'linear',
},
'yaxis': {
'title': '-Zimag, Ohm',
'type': 'log' if logy else 'linear',
},
'legend': {'x': 0.03, 'y': 0.97},
'margin': dict(l=30, r=20, t=60, b=20),
'width': 600,
'height': 600,
}
fig.update_layout(layout)
config={
'displaylogo': False,
'modeBarButtonsToRemove': ['select2d', 'lasso2d', 'hoverClosestCartesian', 'toggleSpikelines','hoverCompareCartesian']
}
fig.show(config=config)
# + id="JVPwUpRcSdXM" cellView="form"
#@markdown **Time-series Plot (Z_mag)**
from plotly.subplots import make_subplots
import plotly.graph_objects as go
from plotly.colors import DEFAULT_PLOTLY_COLORS
impedance_type = "magnitude" #@param ["magnitude", "phase", "real", "imaginary"]
#@markdown Which impedance frequencies should be shown? (separated by comma, e.g. `4, 1000, 10000`)
frequencies_to_show = "3,5,5000" #@param {type:"string"}
impedance_map = dict(magnitude=df_mag, phase=df_phz, real=df_real, imaginary=df_imag)
impedance_yaxis_map = {
"magnitude": {'title': 'Impedance Magnitude, Ohm', 'type': 'log'},
"phase": {'title': 'Phase Angle, deg', 'type': 'linear'},
"real": {'title': 'Real Impedance, Ohm', 'type': 'log'},
"imaginary": {'title': '- Imaginary Impedance, Ohm', 'type': 'log'},
}
frequencies_to_show = [int(val.strip()) for val in frequencies_to_show.split(",")]
def freq_lookup(df, freq):
return df.index.get_loc(freq, method='nearest')
ilocs = [freq_lookup(df_mag, freq) for freq in frequencies_to_show]
source = impedance_map.get(impedance_type)
source_yaxis_config = impedance_yaxis_map.get(impedance_type)
# df_mag.T.to_csv("magnitude_transpose.csv")
df_time = source.copy()
df_time.columns = start_times
df_time.T.to_csv("longitudinal.csv")
df_time = source.copy().iloc[ilocs]
df_time.columns = start_times
df_time = df_time.T
df_time.to_csv("longitudinal-filtered.csv")
fig = make_subplots(rows=1, cols=1, shared_xaxes=True, vertical_spacing=0.02)
# Yields a tuple of column name and series for each column in the dataframe
for (index, (columnName, columnData)) in enumerate(df_time.iteritems()):
newTrace = go.Scatter(
x=df_time.index,
y=columnData,
mode='lines',
name="{} Hz".format(round(columnName,1)),
legendgroup=columnName,
line=dict(color=DEFAULT_PLOTLY_COLORS[index % len(DEFAULT_PLOTLY_COLORS)]),
)
fig.add_trace(newTrace, row=1, col=1)
layout = {
'title': {'text': 'Time-series Plot [{}]'.format(experiment_name),
'yanchor': 'top',
'y': 0.95,
'x': 0.5 },
'xaxis': {
'anchor': 'x',
# 'type': 'log'
},
# 'xaxis2': {
# 'title': 'Frequency, Hz',
# # 'type': 'log',
# 'matches': 'x'
# },
'yaxis': source_yaxis_config,
# 'yaxis2': {
# 'title': 'Phase, deg',
# },
'legend': {'x': 0.85, 'y': 0.97},
'margin': dict(l=30, r=20, t=60, b=20),
'width': 1200,
'height': 500,
}
fig.update_layout(layout)
config={
'displaylogo': False,
'modeBarButtonsToRemove': ['select2d', 'lasso2d', 'hoverClosestCartesian', 'toggleSpikelines','hoverCompareCartesian']
}
fig.show(config=config)
# + id="V17rqu3-A74s" cellView="form"
#@markdown WORK IN PROGRESS **Simulation**: Fit collected data to an electrochemical model
equivalent_model = "Body Impedance Model" #@param ["Randles", "Randles + Diffusion", "Randles + Corrosion", "Body Impedance Model"]
# import additional math libs
import numpy as np
from numpy import pi, sqrt
try:
from lmfit import Model
except:
subprocess.run(
["pip", "install", "--upgrade", "lmfit"],
encoding="utf-8",
shell=False)
finally:
from lmfit import Model
def reference_model(freq, Rct, Cdl_C, Cdl_a, Rsol):
# modifeid randle circuit -- The dual layer capacitance is non-ideal due to
# diffusion-related limitations. It has been replaced with a constant phase
# element. Circuit layout: SERIES(Rsol, PARALLEL(Rct, CPE))
CPE1 = 1/(Cdl_C*(1j*2*pi*freq)**(Cdl_a))
return 1/((1/Rct) + (1/CPE1)) + Rsol
def body_impedance_model(freq, R1, C1, R2, C2, R3, C3, Rs, Cs): # P1, R2, C2, P2, R3, C3, P3, Rs, Cs):
# Layer1 = 1/((1/R1) + (C1*(1j*2*pi*(freq + P1))))
# Layer2 = 1/((1/R2) + (C2*(1j*2*pi*(freq + P2))))
# Layer3 = 1/((1/R3) + (C3*(1j*2*pi*(freq + P3))))
Layer1 = 1/((1/R1) + (C1*(1j*2*pi*(freq))))
Layer2 = 1/((1/R2) + (C2*(1j*2*pi*(freq))))
Layer3 = 1/((1/R3) + (C3*(1j*2*pi*(freq))))
Zc_s = 1/(Cs * 1j * 2 * pi * (freq))
Zr_s = Rs
return Zc_s + Zr_s + Layer1 + Layer2 + Layer3
def diffusion_model(freq, Rct, Cdl_C, Cdl_a, Rsol, Zdf):
# A modified Randle circuit with a warburg component included.
# Circuit layout: SERIES(Rsol, PARALLEL(SERIES(Warburg, Rct), CPE))
CPE1 = 1/(Cdl_C*(1j*2*pi*freq)**(Cdl_a))
## use finite length diffusion constant
# Warburg = Zdf * (np.tanh(sqrt(2j*pi*freq*TCdf))/sqrt(2j*pi*freq*TCdf))
## use infinite warburg coeff (simplified)
Warburg = 1/(Zdf*sqrt(1j*2*pi*freq))
return 1/((1/(Rct + Warburg)) + (1/CPE1)) + Rsol
def corrosion_model(freq, Rc, Cdl_C, Cdl_a, Rsol, Ra, La):
# split cathodic and anodic resistances with inductive component
CPE1 = 1/(Cdl_C*(1j*2*pi*freq)**(Cdl_a))
Za = Ra + 2j*pi*freq*La
Rct = 1 / ((1/Rc) + (1/Za))
return 1/((1/Rct) + (1/CPE1)) + Rsol
def corrosion2_model(freq, Rc, Cdl_C, Cdl_a, Rsol, Ra, La):
# split cathodic and anodic resistances with inductive component
CPE1 = 1/(Cdl_C*(1j*2*pi*freq)**(Cdl_a))
Za = Ra + 2j*pi*freq*La
Rct = 1 / ((1/Rc) + (1/Za))
return 1/((1/Rct) + (1/CPE1)) + Rsol
# create the model
if equivalent_model == "Body Impedance Model":
# model a membrane as resistor and capacitor in parallel.
gmodel = Model(body_impedance_model)
gmodel.set_param_hint('C1', value = 85e-9, min = 1e-9, max=1e-6)
gmodel.set_param_hint('C2', value = 85e-9, min = 1e-9, max=1e-6)
gmodel.set_param_hint('C3', value = 85e-9, min = 1e-9, max=1e-6)
gmodel.set_param_hint('R1', value = 45e3, min=1e3, max=1e6)
gmodel.set_param_hint('R2', value = 875e3, min=1e3, max=1e6)
gmodel.set_param_hint('R3', value = 750, min=0, max=1e3)
gmodel.set_param_hint('Rs', value = 400, min=200, max=600)
gmodel.set_param_hint('Cs', value = 150e-9, min=50e-9, max=5e-6)
elif equivalent_model == 'Randles':
# default, use a randle's circuit with non-ideal capacitor assumption
gmodel = Model(reference_model)
gmodel.set_param_hint('Rct', value = 1e7, min = 1e3, max = 1e9)
elif equivalent_model == 'Randles + Diffusion':
# use previous model and add a warburg
gmodel = Model(diffusion_model)
gmodel.set_param_hint('Rct', value = 1e6, min = 1e3, max = 1e10)
gmodel.set_param_hint('Zdf', value = 1e4, min = 1e3, max = 1e6)
else:
# Randle + Corrosion
gmodel = Model(corrosion_model)
gmodel.set_param_hint('Rc', value = 5e5, min = 1e3, max = 1e10)
gmodel.set_param_hint('Ra', value = 5e5, min = 1e3, max = 1e10)
gmodel.set_param_hint('La', value = 1e6, min = 0, max = 1e9)
# initial guess shared across all models, with defined acceptable limits
gmodel.set_param_hint('Cdl_C', value = 5e-5, min = 1e-12)
gmodel.set_param_hint('Cdl_a', value = 0.9, min = 0, max = 1)
gmodel.set_param_hint('Rsol', value = 1000, min = 100, max = 5e5)
# now solve for each loaded sensor
a = []
freq = np.asarray(df_mag.index)
for (index, columnName) in enumerate(df_mag.columns):
print('Model Simulation [{}] on [{}]'.format(equivalent_model, columnName))
impedance = np.asarray(df_real[columnName]) - 1j * np.asarray(df_imag[columnName])
# fit_weights = (np.arange(len(freq))**.6)/len(freq) #weight ~ freq
fit_weights = np.ones(len(freq))/len(freq) #equal weight
result = gmodel.fit(impedance, freq=freq, weights = fit_weights)
print(result.fit_report(show_correl=False))
fig = make_subplots(rows=2, cols=1, shared_xaxes=True, vertical_spacing=0.02)
data = []
# Yields a tuple of column name and series for each column in the dataframe
rawTrace = go.Scatter(
x=freq,
y=df_mag[columnName],
mode='markers',
name=columnName,
legendgroup='raw',
marker=dict(color=DEFAULT_PLOTLY_COLORS[0]),
)
fig.add_trace(rawTrace, row=1, col=1)
fitTrace = go.Scatter(
x=freq,
y=np.abs(result.best_fit),
mode='lines',
name=columnName,
legendgroup='model',
line=dict(color=DEFAULT_PLOTLY_COLORS[1]),
)
fig.add_trace(fitTrace, row=1, col=1)
rawTrace = go.Scatter(
x=freq,
y=-df_phz[columnName],
mode='markers',
name=columnName,
legendgroup='raw',
showlegend=False,
marker=dict(color=DEFAULT_PLOTLY_COLORS[0]),
)
fig.add_trace(rawTrace, row=2, col=1)
fitTrace = go.Scatter(
x=freq,
y=-180/pi*np.angle(result.best_fit),
mode='lines',
name=columnName,
legendgroup='model',
showlegend=False,
line=dict(color=DEFAULT_PLOTLY_COLORS[1]),
)
fig.add_trace(fitTrace, row=2, col=1)
layout = {
'title': 'Model Fit [{}] for [{}]'.format(equivalent_model, columnName),
'xaxis': {
'anchor': 'x',
'type': 'log'
},
'xaxis2': {
'anchor': 'x',
'type': 'log'
},
'yaxis': {
'type': 'log'
},
'legend': {'x': 0.85, 'y': 0.97},
'margin': dict(l=30, r=20, t=60, b=20),
'width': 1200,
'height': 500,
}
fig.update_layout(layout)
fig.show()
| demo/notebook_potentiostatic_eis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Importing **Numpy**
import numpy as np
# ### Checking Numpy version
np.version.version
np.__version__
# ### Creation
# Creating two arrays:
# * Python array
# * Numpy array.
# +
# python ordinary array
py_array = [0,1,2,3,4,5,6,7,8,9]
# numpy array
np_array = np.array([0,1,2,3,4,5,6,7,8,9])
# -
# Printing python array
print(py_array)
# Printing Numpy array
print(np_array)
# ### Type
type(py_array)
type(np_array)
# ### Shape
# printing the length of python list/array
len(py_array)
# printing the length of numpy ndarray
len(np_array)
# printing the length of the outter array of np_matrix
len(np_matrix)
# +
# create numpy array contains five elements
np_array = np.array([1, 2, 3, 4, 5])
# printing np_array shape(length per axis)
np_array.shape
# -
# Shape of Matrix
# +
# create numpy multi-dimensional array contains six elements(2 rows, 3 columns)
np_matrix = np.array([[1,2,3],[4,5,6]])
# printing np_matrix shape(length per axis/ number of rows and columns)
np_matrix.shape
# -
# ### `np.reshape()`
# ### `np.arrange()`
np.arange(0,10)
np.arange(0,10,2)
np.arange(0,10) == np.arange(0,10,1)
# ### np.zeros()
# ### np.ones()
# ### np.eye()
# ### np.dot()
# ### np.sum()
# ### Testing the speed
# %%timeit
count = 0
for i in range(0, 1000):
count = count + 1
# %%timeit
np_count = np.sum(np.ones((1000,), dtype=int), axis=0)
count
np_count
| numpy/numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
df = pd.read_csv('wat_good_routers.csv')
df.shape
df.max()
df.dtypes
interval = 99
r0count = 0
r1count = 0
r2count = 0
r3count = 0
r4count = 0
r5count = 0
r6count = 0
r7count = 0
r8count = 0
r9count = 0
r10count = 0
r11count = 0
r12count = 0
r13count = 0
r14count = 0
r15count = 0
r0arr = []
r1arr = []
r2arr = []
r3arr = []
r4arr = []
r5arr = []
r6arr = []
r7arr = []
r8arr = []
r9arr = []
r10arr = []
r11arr = []
r12arr = []
r13arr = []
r14arr = []
r15arr = []
for index, row in df.iterrows():
if row['time']<=interval:
if row['router0']==1:
r0count+=1
elif row['router1']==1:
r1count+=1
elif row['router2']==1:
r2count+=1
elif row['router3']==1:
r3count+=1
elif row['router4']==1:
r4count+=1
elif row['router5']==1:
r5count+=1
elif row['router6']==1:
r6count+=1
elif row['router7']==1:
r7count+=1
elif row['router8']==1:
r8count+=1
elif row['router9']==1:
r9count+=1
elif row['router10']==1:
r10count+=1
elif row['router11']==1:
r11count+=1
elif row['router12']==1:
r12count+=1
elif row['router13']==1:
r13count+=1
elif row['router14']==1:
r14count+=1
elif row['router15']==1:
r15count+=1
else:
r0arr.append(r0count)
r1arr.append(r1count)
r2arr.append(r2count)
r3arr.append(r3count)
r4arr.append(r4count)
r5arr.append(r5count)
r6arr.append(r6count)
r7arr.append(r7count)
r8arr.append(r8count)
r9arr.append(r9count)
r10arr.append(r10count)
r11arr.append(r11count)
r12arr.append(r12count)
r13arr.append(r13count)
r14arr.append(r14count)
r15arr.append(r15count)
if row['router0']==1:
r0count = 1
else:
r0count = 0
if row['router1']==1:
r1count = 1
else:
r1count = 0
if row['router2']==1:
r2count = 1
else:
r2count = 0
if row['router3']==1:
r3count = 1
else:
r3count = 0
if row['router4']==1:
r4count = 1
else:
r4count = 0
if row['router5']==1:
r5count = 1
else:
r5count = 0
if row['router6']==1:
r6count = 1
else:
r6count = 0
if row['router7']==1:
r7count = 1
else:
r7count = 0
if row['router8']==1:
r8count = 1
else:
r8count = 0
if row['router9']==1:
r9count = 1
else:
r9count = 0
if row['router10']==1:
r10count = 1
else:
r10count = 0
if row['router11']==1:
r11count = 1
else:
r11count = 0
if row['router12']==1:
r12count = 1
else:
r12count = 0
if row['router13']==1:
r13count = 1
else:
r13count = 0
if row['router14']==1:
r14count = 1
else:
r14count = 0
if row['router15']==1:
r15count = 1
else:
r15count = 0
interval+=100
df = pd.DataFrame(range(100,3152969,100), columns=["time_intervals"])
df
len(r0arr)
df = df.assign(r0=r0arr)
df = df.assign(r1=r1arr)
df = df.assign(r2=r2arr)
df = df.assign(r3=r3arr)
df = df.assign(r4=r4arr)
df = df.assign(r5=r5arr)
df = df.assign(r6=r6arr)
df = df.assign(r7=r7arr)
df = df.assign(r8=r8arr)
df = df.assign(r9=r9arr)
df = df.assign(r10=r10arr)
df = df.assign(r11=r11arr)
df = df.assign(r12=r12arr)
df = df.assign(r13=r13arr)
df = df.assign(r14=r14arr)
df = df.assign(r15=r15arr)
df['target']=1
df
df.to_csv('wat-good-time.csv',index=False)
| [02 - Modeling]/dos ver 4/wat-good-editable.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/harveenchadha/bol/blob/main/demos/hf/hindi/hf_hindi_him_4200_demo.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="BUR_s3tt_IJF"
# # Vakyansh + Hugging Face : Hindi Speech To text Demo
# + [markdown] id="gQCiOn_9_ZeI"
# ## Install requirements
# + id="X_UDaMuH_GnL"
# %%capture
# !apt-get -y install sox ffmpeg
# !pip install transformers ffmpeg-python sox
# !wget https://raw.githubusercontent.com/harveenchadha/bol/main/demos/colab/record.py
# + [markdown] id="Ikp7WVluBacM"
# ## Load Hindi Model
# + id="pZ5rnYAt_rB1"
import soundfile as sf
import torch
from transformers import Wav2Vec2ForCTC, Wav2Vec2Processor
def load_model():
# load pretrained model
processor = Wav2Vec2Processor.from_pretrained("Harveenchadha/vakyansh-wav2vec2-hindi-him-4200")
model = Wav2Vec2ForCTC.from_pretrained("Harveenchadha/vakyansh-wav2vec2-hindi-him-4200")
return processor, model
processor, model = load_model()
# + id="eT5eskX5KqEZ"
def parse_transcription(wav_file):
# load audio
audio_input, sample_rate = sf.read(wav_file)
# pad input values and return pt tensor
input_values = processor(audio_input, sampling_rate=16_000, return_tensors="pt").input_values
# INFERENCE
# retrieve logits & take argmax
logits = model(input_values).logits
predicted_ids = torch.argmax(logits, dim=-1)
# transcribe
transcription = processor.decode(predicted_ids[0], skip_special_tokens=True)
return transcription
# + [markdown] id="v6a1YbhPAXY0"
# ## Record file using colab
# + colab={"base_uri": "https://localhost:8080/", "height": 167} id="irfbRvhbAVs8" outputId="1a66139c-127a-424d-871b-bebe547e4d48"
from record import record_audio
record_audio('test')
# + [markdown] id="sHiDi3a0JKNk"
# ## Run Model on recorded file
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="uIccgORyCm0S" outputId="30e73a16-fd08-4bef-eaa9-ab765d0182cd"
parse_transcription('test.wav')
| demos/hf/hindi/hf_hindi_him_4200_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
from __future__ import division, print_function, absolute_import
import numpy as np
import matplotlib.pyplot as plt
import tflearn
# %matplotlib inline
# Data loading and preprocessing
import tflearn.datasets.mnist as mnist
X, Y, testX, testY = mnist.load_data(one_hot=True)
# Building the encoder
encoder = tflearn.input_data(shape=[None, 784])
encoder = tflearn.fully_connected(encoder, 256)
encoder = tflearn.fully_connected(encoder, 64)
# Building the decoder
decoder = tflearn.fully_connected(encoder, 256)
decoder = tflearn.fully_connected(decoder, 784)
# Regression, with mean square error
net = tflearn.regression(decoder, optimizer='adam', learning_rate=0.001,
loss='mean_square', metric=None)
# Training the auto encoder
model = tflearn.DNN(net, tensorboard_verbose=0)
model.fit(X, X, n_epoch=10, validation_set=(testX, testX),
run_id="auto_encoder", batch_size=256)
# Encoding X[0] for test
print("\nTest encoding of X[0]:")
# New model, re-using the same session, for weights sharing
encoding_model = tflearn.DNN(encoder, session=model.session)
print(encoding_model.predict([X[0]]))
# Testing the image reconstruction on new data (test set)
print("\nVisualizing results after being encoded and decoded:")
testX = tflearn.data_utils.shuffle(testX)[0]
# Applying encode and decode over test set
encode_decode = model.predict(testX)
# Compare original images with their reconstructions
f, a = plt.subplots(2, 10, figsize=(10, 2))
for i in range(10):
a[0][i].imshow(np.reshape(testX[i], (28, 28)))
a[1][i].imshow(np.reshape(encode_decode[i], (28, 28)))
f.show()
plt.draw()
plt.waitforbuttonpress()
# -
X[0]
| autoencoders/mnist_encoder_decoder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ##### Preamble
# +
import numpy as np
from scipy.spatial.distance import cdist
from scipy.special import expit
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.pipeline import make_pipeline, make_union
from sklearn.preprocessing import PolynomialFeatures
from sklearn.linear_model import LinearRegression
from sklearn.datasets import load_diabetes
# -
# [Feature Union with Heterogeneous Data Sources](http://scikit-learn.org/stable/auto_examples/hetero_feature_union.html)
# ### Polynomial basis function
# The polynomial basis function is provided by `scikit-learn` in the [sklearn.preprocessing](http://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.PolynomialFeatures.html) module.
X = np.arange(1, 9).reshape(4, 2)
X
PolynomialFeatures(degree=2).fit_transform(X)
# ### Custom basis functions
# Unfortunately, this is pretty much the extent of what `scikit-learn` provides in the way of basis functions. Here we define some standard basis functions, while adhering to the `scikit-learn` interface. This will be important when we try to incorporate our basis functions in pipelines and feature unions later on. While this is not strictly required, it will certainly make life easier for us down the road.
# #### Radial Basis Function
class RadialFeatures(BaseEstimator, TransformerMixin):
def __init__(self, mu=0, s=1):
self.mu = mu
self.s = s
def fit(self, X, y=None):
# this basis function stateless
# need only return self
return self
def transform(self, X, y=None):
return np.exp(-cdist(X, self.mu, 'sqeuclidean')/(2*self.s**2))
# #### Sigmoidal Basis Function
class SigmoidalFeatures(BaseEstimator, TransformerMixin):
def __init__(self, mu=0, s=1):
self.mu = mu
self.s = s
def fit(self, X, y=None):
# this basis function stateless
# need only return self
return self
def transform(self, X, y=None):
return expit(cdist(X, self.mu)/self.s)
mu = np.linspace(0.1, 1, 10).reshape(5, 2)
mu
RadialFeatures(mu=mu).fit_transform(X).round(2)
SigmoidalFeatures(mu=mu).fit_transform(X).round(2)
# ### Real-world Dataset
# Now that we have a few basis functions at our disposal, let's try to apply different basis functions to different features of a dataset. We use the diabetes dataset, a real-world dataset with 442 instances and 10 features. We first work through each step manually, and show how the steps can be combined using `scikit-learn`'s feature unions and pipelines to form a single model that will perform all the necessary steps in one fell swoop.
diabetes = load_diabetes()
X, y = diabetes.data, diabetes.target
X.shape
y.shape
# We print every other feature for just the first few instances, just to get an idea of what the data looks like
# sanity check
X[:5, ::2]
# sanity check
y[:5]
# Assume for some reason we are interested in training a model using, say, features 2 and 5 with a *polynomial basis*, and features 6, 8 and 9 with a *radial basis*. We first slice up our original dataset.
X1 = X[:, np.array([2, 5])]
X1.shape
# sanity check
X1[:5]
X2 = X[:, np.array([6, 8, 9])]
X2.shape
# sanity check
X2[:5]
# Now we apply the respective basis functions.
# #### Polynomial
X1_poly = PolynomialFeatures().fit_transform(X1)
X1_poly.shape
# sanity check
X1_poly[:5].round(2)
# #### Radial
mu = np.linspace(0, 1, 6).reshape(2, 3)
mu
X2_radial = RadialFeatures(mu).fit_transform(X2)
X2_radial.shape
# sanity check
X2_radial[:5].round(2)
# Now we're ready to concatenate these augmented datasets.
X_concat = np.hstack((X1_poly, X2_radial))
X_concat.shape
# sanity check
X_concat[:5, ::2].round(2)
# Now we are ready to train a regressor with this augmented dataset. For this example, we'll simply use a linear regression model.
model = LinearRegression()
model.fit(X_concat, y)
model.score(X_concat, y)
# *(To no one's surprise, our model performs quite poorly, since zero effort was made to identify and incorporate the most informative features or appropriate basis functions. Rather, they were chosen solely to maximize clarity of exposition.)*
# #### Recap
#
# So let's recap what we've done.
#
# 1. We started out with a dataset with 442 samples and 10 features, represented by **442x10 matrix `X`**
# 2. For one reason or another, we wanted to use different basis functions for different subsets of features. Apparently, we wanted features 2 and 5 for one basis function and features 6, 8 and 9 for another. Therefore, we
# 1. sliced the matrix `X` to obtain **442 by 2 matrix `X1`** and
# 2. sliced the matrix `X` to obtain **442 by 3 matrix `X2`**.
# 3. We
# 1. applied a polynomial basis function of degree 2 to `X1` with 2 features and 442 samples. This returns a dataset `X1_poly` with $\begin{pmatrix} 4 \\ 2 \end{pmatrix} = 6$ features and 442 samples. (**NB:** In general, the number of output features for a polynomial basis function of degree $d$ on $n$ features is the number of multisets of cardinality $d$, with elements taken from a finite set of cardinality $n+1$, which is given by the multiset coefficient $\begin{pmatrix} \begin{pmatrix} n + 1 \\ d \end{pmatrix} \end{pmatrix} = \begin{pmatrix} n + d \\ d \end{pmatrix}$.) So from 442 by 2 matrix `X1` we obtain **442 by 6 matrix `X1_poly`**
# 2. applied a radial basis function with 2 mean vectors $\mu_1 = \begin{pmatrix} 0 & 0.2 & 0.4 \end{pmatrix}^T$ and $\mu_2 = \begin{pmatrix} 0.6 & 0.8 & 1.0 \end{pmatrix}^T$, which is represented by the 2 by 3 matrix `mu`. From the 442 by 3 matrix `X2`, we obtain **442 by 2 matrix `X2_radial`**
# 4. Next, we horizontally concatenated 442 by 6 matrix `X1_poly` with 442 by 2 matrix `X2_radial` to obtain the final **442 by 8 matrix `X_concat`**
# 5. Finally, we fitted a linear model on `X_concat`.
#
# So this is how we went from a 442x**10** matrix `X` to the 442x**8** matrix `X_concat`.
# ### With Pipeline and Feature Union
# First we define a transformer that slices up the input data. Note instead of working with (tuples of) slice objects, it is usually more convenient to use the Numpy function `np.index_exp`. We explain later why this is necessary.
class ArraySlicer(BaseEstimator, TransformerMixin):
def __init__(self, index_exp):
self.index_exp = index_exp
def fit(self, X, y=None):
return self
def transform(self, X, y=None):
return X[self.index_exp]
model = \
make_pipeline(
make_union(
make_pipeline(
ArraySlicer(np.index_exp[:, np.array([2, 5])]),
PolynomialFeatures()
),
make_pipeline(
ArraySlicer(np.index_exp[:, np.array([6, 8, 9])]),
RadialFeatures(mu)
)
)
)
model.fit(X)
model.transform(X).shape
# sanity check
model.transform(X)[:5, ::2].round(2)
# This effectively composes each of the steps we had to manually perform and amalgamated it into a single transformer. We can even append a regressor at the end to make it a complete estimator/predictor.
model = \
make_pipeline(
make_union(
make_pipeline(
ArraySlicer(np.index_exp[:, np.array([2, 5])]),
PolynomialFeatures()
),
make_pipeline(
ArraySlicer(np.index_exp[:, np.array([6, 8, 9])]),
RadialFeatures(mu)
)
),
LinearRegression()
)
model.fit(X, y)
model.score(X, y)
# ### Breaking it Down
# The most important thing to note is that everything in `scikit-learn` is either a transformer or a predictor, and are almost always an estimator. An estimator is simply a class that implements the `fit` method, while a transfromer and predictor implements a, well, `transform` and `predict` method respectively. From this simple interface, we get a surprising hight amount of functionality and flexibility.
# #### Pipeline
# A pipeline behaves as a transformer or a predictor depending on what the last step of the pipleline is. If the last step is a transformer, the entire pipeline is a transformer and one can call `fit`, `transform` or `fit_transform` like an ordinary transformer. The same is true if the last step is a predictor. Essentially, all it does is chain the `fit_transform` calls of every transformer in the pipeline. If we think of ordinary transformers like functions, pipelines can be thought of as a higher-order function that simply composes an arbitary number of functions.
model = \
make_pipeline(
PolynomialFeatures(), # transformer
LinearRegression() # predictor
)
model.fit(X, y)
model.score(X, y)
# #### Union
# A union is a transformer that is initialized with an arbitrary number of transformers. When `fit_transform` is called on a dataset, it simply calls `fit_transform` of the transformers it was given and horizontally concatenates its results.
mu_ = np.linspace(0, 10, 30).reshape(3, 10)
model = \
make_union(
PolynomialFeatures(),
RadialFeatures(mu_)
)
# If we run this on the original 442x10 dataset, we expect to get a dataset with the same number of samples and $\begin{pmatrix} 12 \\ 2 \end{pmatrix} + 3 = 66 + 3 = 69$ features.
model.fit_transform(X).shape
# ### Putting it all together
# The above union applies the basis functions on the entire dataset, but we're interested in applying different basis functions to different features. To do this, we can simply define a rather frivolous transformer that simply slices the input data, and that's exactly what `ArraySlicer` was for.
model = \
make_pipeline(
ArraySlicer(np.index_exp[:, np.array([2, 5])]),
PolynomialFeatures()
)
model.fit(X)
model.transform(X).shape
# sanity check
model.transform(X)[:5].round(2)
# Then we can combine this all together to form our mega-transformer which we showed earlier.
model = \
make_pipeline(
make_union(
make_pipeline(
ArraySlicer(np.index_exp[:, np.array([2, 5])]),
PolynomialFeatures()
),
make_pipeline(
ArraySlicer(np.index_exp[:, np.array([6, 8, 9])]),
RadialFeatures(mu)
)
),
LinearRegression()
)
# This gives us a predictor which takes some input, slices up the respective features, churns it through a basis function and finally trains a linear regressor on it, all in one go!
model.fit(X, y)
model.score(X, y)
# #### Inter
# ## Propagating Variable and Keyword arguments in a pipeline
| Concatenating Basis Functions with Heterogeneous Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Creation of tables
import sqlite3
import pandas as pd
import numpy as np
# ### Initial read
# +
# connect to database and get data
conn = sqlite3.connect('data/youtube-may-2016-2018.db')
conn.text_factory = str
# read whole data from raw dataset for aggregation
date_cols = ['dtime']
traceroute = pd.read_sql_query('select * from traceroute', conn, parse_dates = date_cols)
msmpoint = pd.read_sql_query('select * from msmpoint', conn)
conn.close()
# +
# open connection to new database (which will be used for all analyses)
conn = sqlite3.connect('data/youtube-traceroute.db')
conn.text_factory = str
# write table msmpoint to new db
msmpoint.to_sql(name = 'msmpoint', con = conn, index = False, if_exists = 'replace')
# -
# ### Filtering unusable data
# +
# drop all duplicate rows that are identical in all columns (i.e. redundant information)
traceroute = traceroute.drop_duplicates()
# drop measurements from August 2017 to January 2018 due to faulty measurement runs at that time
traceroute = traceroute[(traceroute['dtime'] < '2017-08-01') | (traceroute['dtime'] >= '2018-02-01')]
# -- Manual removal of probes that are problematic
# drop probes that have shown non-dualstacked behavior
traceroute = traceroute[~traceroute['unit_id'].isin([525884, 658929])]
# drop probes that use Hurricane Electric as a tunneling service for IPv6 connectivity
traceroute = traceroute[~traceroute['unit_id'].isin([19602, 632406, 660076])]
# Storing to database
traceroute.to_sql(name = 'traceroute', con = conn, index = False, if_exists = 'replace')
# -
traceroute
# ### General aggregation
#
# We only analyze COMPLETED traces from now on, as failure analysis showed that only a few percent of traces did not succeed (see failures.ipynb notebook).
# +
# get grouped data for each ~hourly measurement from each device
df = pd.read_sql_query('select unit_id, dtime, source, destination, max(ttl), rtt \
from traceroute \
where status="COMPLETED" and \
destination = endpoint \
group by unit_id, dtime, source, destination', conn)
# parse date from string to datetime and set type of RTT to float
df['dtime'] = pd.to_datetime(df['dtime'])
df.rtt = df.rtt.astype(float)
# query two separate versions of the dataframe, one for v6, one for v4
df_v6 = df[df['source'].str.contains(':')]
df_v4 = df[~df['source'].str.contains(':')]
# Write dataframes into table in database, replace if already exists
df_v6.to_sql(name = 'traceroute_v6', con = conn, index = False, if_exists = 'replace')
df_v4.to_sql(name = 'traceroute_v4', con = conn, index = False, if_exists = 'replace')
# -
df_v6
df_v4
# ## Path Medians (grouped by unit_id, src, dst)
# +
# group by paths and compute median TTL and RTT for each group, this removes time as a dimension
df_v6_median = df_v6.groupby(['unit_id', 'source', 'destination'], as_index = False).median()
df_v4_median = df_v4.groupby(['unit_id', 'source', 'destination'], as_index = False).median()
# rename columns to represent what was done
df_v6_median.columns = ['unit_id', 'source', 'destination', 'median(ttl)', 'median(rtt)']
df_v4_median.columns = ['unit_id', 'source', 'destination', 'median(ttl)', 'median(rtt)']
# round medians of RTT to 3 decimals
df_v6_median['median(rtt)'] = df_v6_median['median(rtt)'].round(3)
df_v4_median['median(rtt)'] = df_v4_median['median(rtt)'].round(3)
# write dataframe into table in database
df_v6_median.to_sql(name = 'path_medians_v6', con = conn, index = False, if_exists = 'replace')
df_v4_median.to_sql(name = 'path_medians_v4', con = conn, index = False, if_exists = 'replace')
# -
df_v6_median
df_v4_median
# ## AS Mappings
#
# Load and write collected metadata into database tables
#
# - dst IP address to ASN
# - endpoint IP address to ASN
# +
# destination <-> ASN
dst_asn_df = pd.read_csv('metadata/dst_ip_to_asn.csv', sep = ';')
dst_asn_df.to_sql(name = 'dst_asn_mapping', con = conn, index = False, if_exists = 'replace')
# intermediate endpoints <-> ASN
endpoint_asn_df = pd.read_csv('metadata/endpoint_asn_lookup.csv', sep = ';')
endpoint_asn_df.to_sql(name = 'endpoint_asn_mapping', con = conn, index = False, if_exists = 'replace')
# -
dst_asn_df
endpoint_asn_df
# ## Destination address to hostnames (through reverse DNS lookup data)
#
# Load and write collected metadata into database table
# load hostnames from reverse DNS lookup into database
hostnames_df = pd.read_csv('metadata/dst_ip_to_hostname.csv', sep = ';')
hostnames_df.to_sql(name = 'hostnames', con = conn, index = False, if_exists = 'replace')
hostnames_df
# ## Deltas of both versions for hourly measurements
# +
# round down/cut off minutes and seconds for grouping
df_v6['dtime'] = df_v6['dtime'].dt.floor('h')
df_v4['dtime'] = df_v4['dtime'].dt.floor('h')
# ignore SettingWithCopyWarnings; working as intended, see below
# +
# merge v4 and v6 dataframes on unit_id and dtime (inner join)
merged = pd.merge(df_v4, df_v6, on = ['unit_id', 'dtime'])
# add column for TTL delta, i.e. TTL_IPv4 - TTL_IPv6
merged['ttl_delta'] = merged['max(ttl)_x'] - merged['max(ttl)_y']
# add column for RTT delta, i.e. RTT_IPv4 - RTT_IPv6
merged['rtt_delta'] = merged['rtt_x'] - merged['rtt_y']
# round RTT deltas to 3 decimals
merged['rtt_delta'] = merged['rtt_delta'].round(3)
# define clear column names to replace _x and _y columns
cols = ['unit_id', 'dtime', 'source_v4', 'destination_v4', 'max(ttl)_v4', 'rtt_v4', 'source_v6', 'destination_v6', 'max(ttl)_v6', 'rtt_v6', 'ttl_delta', 'rtt_delta']
merged.columns = cols
# Write dataframe into table in database
merged.to_sql(name = 'deltas', con = conn, index = False, if_exists = 'replace')
# -
merged
# ## Medians of destination pairs
# make use of delta table from above and use grouping for destination pair medians
pair_medians = merged.groupby(['unit_id', 'destination_v4', 'destination_v6'], as_index = False).median()
pair_medians.to_sql(name = 'pair_medians', con = conn, index = False, if_exists = 'replace')
pair_medians
# ## Add meta data to table of destination pair medians
# +
# read measurement point information which contain msmIDs and ASNs
msmpoint = pd.read_sql_query('select * from msmpoint', conn)
# load metadata of probes; required to get skunitid for merging
probes_metadata = pd.read_csv('metadata/probes_metadata.txt', engine = 'python', sep = '|', skiprows = 3, skipfooter = 1, names = ['probe', 'since', 'hardware', 'asnv4', 'asnv6', 'location', 'type', 'test'])
probes_metadata = probes_metadata[['probe', 'since', 'hardware', 'asnv4', 'asnv6', 'location', 'type']].reset_index().drop('index', 1)
# replace probe number with name to merge with table 'msmpoint' to get skunitid
# strip whitespace from beginning and end of string
probes_metadata['probe'] = probes_metadata['probe'].str.replace('#', 'samsbox').str.strip()
probes = pd.merge(probes_metadata, msmpoint, left_on = 'probe', right_on = 'name')
probes = probes.rename(columns = {'skunitid' : 'unit_id', 'asnv4' : 'src_asn_v4', 'asnv6' : 'src_asn_v6'})
probes = probes[['unit_id', 'src_asn_v4', 'src_asn_v6']]
# -
# some probes are not assigned an ASN for IPv6
probes[probes['src_asn_v6'].str.contains('-')]
# in those cases, assume that source ASNs are the same over both address families
probes['src_asn_v6'] = np.where(probes['src_asn_v6'].str.contains('-'), probes['src_asn_v4'], probes['src_asn_v6'])
# +
# AS numbers from string to int
probes['src_asn_v4'] = probes['src_asn_v4'].str.replace('AS', '').astype(int)
probes['src_asn_v6'] = probes['src_asn_v6'].str.replace('AS', '').astype(int)
# cast unit_id to int
probes['unit_id'] = probes['unit_id'].astype(int)
probes
# -
# determine holders by AS numbers
src = pd.read_csv('metadata/src_asn_holders.csv', sep = ';')
probes = probes.merge(src, left_on = 'src_asn_v4', right_on = 'asn').rename(columns = {'holder' : 'src_holder_v4'})
probes = probes.merge(src, left_on = 'src_asn_v6', right_on = 'asn').rename(columns = {'holder' : 'src_holder_v6'})
probes = probes.drop(['asn_x', 'asn_y'], axis = 1)
probes
probes.to_sql(name = 'probes_asns' , con = conn, index = False, if_exists = 'replace')
# +
# merge probe src ASN to pair_medians table
pair_medians = pd.read_sql_query('select * from pair_medians', conn)
pair_medians = pair_medians.merge(probes, on = 'unit_id')
# +
# read remaining mappings from database (rather than CSVs)
dst = pd.read_sql_query('select * from dst_asn_mapping', conn)
hostnames = pd.read_sql_query('select * from hostnames', conn)
# distinguish between v4 and v6 data
dst_v6 = dst[dst.ip.str.contains(':')]
dst_v4 = dst[~dst.ip.str.contains(':')]
hostnames_v6 = hostnames[hostnames.ip.str.contains(':')]
hostnames_v4 = hostnames[~hostnames.ip.str.contains(':')]
# add destination ASNs
pair_medians = pair_medians.merge(dst_v4, left_on = 'destination_v4', right_on = 'ip').rename(columns = {'ip' : 'dst_v4', 'asn' : 'dst_asn_v4', 'holder' : 'dst_holder_v4'})
pair_medians = pair_medians.merge(dst_v6, left_on = 'destination_v6', right_on = 'ip').rename(columns = {'ip' : 'dst_v6', 'asn' : 'dst_asn_v6', 'holder' : 'dst_holder_v6'})
# add hostnames
pair_medians = pair_medians.merge(hostnames_v4, how = 'left', left_on = 'destination_v4', right_on = 'ip').rename(columns = {'ip' : 'ip_hostname_v4', 'hostname' : 'hostname_v4'})
pair_medians = pair_medians.merge(hostnames_v6, how = 'left', left_on = 'destination_v6', right_on = 'ip').rename(columns = {'ip' : 'ip_hostname_v6', 'hostname' : 'hostname_v6'})
# pick correct columns and rearrange order
pair_medians = pair_medians[['unit_id',
'src_asn_v4', 'src_holder_v4', 'dst_v4', 'hostname_v4', 'dst_asn_v4', 'dst_holder_v4', 'max(ttl)_v4', 'rtt_v4',
'src_asn_v6', 'src_holder_v6', 'dst_v6', 'hostname_v6', 'dst_asn_v6', 'dst_holder_v6', 'max(ttl)_v6', 'rtt_v6',
'ttl_delta', 'rtt_delta']]
# rename to remind these are medians
pair_medians.rename(columns = {'max(ttl)_v4' : 'm_ttl_v4', 'max(ttl)_v6' : 'm_ttl_v6',
'rtt_v4' : 'm_rtt_v4', 'rtt_v6' : 'm_rtt_v6',
'ttl_delta' : 'm_ttl_delta', 'rtt_delta' : 'm_rtt_delta'},
inplace = True)
# remove rows where src_asn_v6 = 6939, which is Hurricane Electrics -> affected measurements use tunneling service for v6, not native
pair_medians = pair_medians[pair_medians['src_asn_v6'] != 6939].reset_index() # Note: this should have already been done using filter_data.py, just to double-check here!
# sort dataframe to avoid some weird interactions that occured when merging
pair_medians.sort_values(by = 'unit_id', inplace = True)
pair_medians = pair_medians.drop('index', axis = 1) # drop index column that got in through merging
# writeback
pair_medians.to_sql(name = 'pair_medians_meta' , con = conn, index = False, if_exists = 'replace')
# -
conn.close()
pair_medians
| nb-create_tables.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# >原文地址 https://blog.csdn.net/qq_33733970/article/details/77719427?utm_medium=distribute.pc_relevant.none-task-blog-baidujs-1
#
# GIL 与互斥锁再理解
# ===========
#
# 
#
# 线程一要把 python 代码交给解释器去执行,而此时垃圾回收线程和线程二也需要将自己的任务交给 python 解释器去执行,为了防止各个线程之间的数据产生冲突,谁拿到 GIL 锁的权限谁才能执行自己的任务,这就避免了不同任务之间的数据不会产生冲突,这是在同一个进程中加 GIL 锁会保证数据的安全,不同的数据要加不同的锁。
# 死锁与递归锁
# ======
#
# 死锁
# --
#
# ### 代码演示
#
# ```py
# from threading import Thread, Lock
# import time
# mutexA = Lock()
# mutexB = Lock()
#
# class MyThread(Thread):
# def run(self):
# self.f1()
# self.f2()
#
# def f1(self):
# mutexA.acquire()
# print('%s 拿到A锁' % self.name)
# mutexB.acquire()
# print('%s 拿到B锁' % self.name)
# mutexB.release()
# mutexA.release()
#
# def f2(self):
# mutexB.acquire()
# time.sleep(1)
# print('%s 拿到B锁' % self.name)
# mutexA.acquire()
# print('%s 拿到A锁' % self.name)
# mutexA.release()
# mutexB.release()
#
#
# if __name__ == '__main__':
# for i in range(10):
# t = MyThread()
# t.start()
# ```
#
# 该种情况出现死锁:
# 
#
# ### 代码讲解
#
# 由于 Thread-1 创建的比较快,所以 Thread-1 先抢到 A 锁,继而顺利成章的拿到 B 锁,当 Thread-1 释放掉 A 锁时,另外 9 个线程抢 A 锁,于此同时,Thread-1 抢到 B 锁,而此时 Thread-2 抢到 A 锁,这样 Thread-1、Thread-2 就等待彼此把锁释放掉,这样程序就卡住了,解决这个问题就用到了递归锁。
# 递归锁
# ---
#
# ### 代码演示
#
# ```py
# from threading import Thread, Lock, RLock
# import time
#
# # mutexA = Lock()
# # mutexB = Lock()
# mutexA = mutexB = RLock()
#
#
# class MyThread(Thread):
# def run(self):
# self.f1()
# self.f2()
#
# def f1(self):
# mutexA.acquire()
# print('%s 拿到A锁' % self.name)
# mutexB.acquire()
# print('%s 拿到B锁' % self.name)
# mutexB.release()
# mutexA.release()
#
# def f2(self):
# mutexB.acquire()
# time.sleep(1)
# print('%s 拿到B锁' % self.name)
# mutexA.acquire()
# print('%s 拿到A锁' % self.name)
# mutexA.release()
# mutexB.release()
#
#
# if __name__ == '__main__':
# for i in range(10):
# t = MyThread()
# t.start()
# ```
#
# ### 代码讲解
#
# 递归锁时通过计数完成对锁的控制的,当 acquire 一次,count+=1,release 一次,count-=1,当 count=0,所有的线程都可以对锁进行抢夺。从而避免了死锁的产生。
# 信号量 Semaphore
# =============
#
# 代码演示
# ----
# ```py
# from threading import Thread, Semaphore, currentThread
# import time
# smph = Semaphore(5)
#
#
# def do_task():
# smph.acquire()
# print('\033[45m%s\033[0m 获得了权限' % currentThread().name)
# time.sleep(2)
# print('\033[46m%s\033[0m 放弃了权限' % currentThread().name)
# smph.release()
#
#
# if __name__ == '__main__':
# for i in range(10)
# t = Thread(target=do_task, )
# t.start()
# ```
# 代码效果
# ----
#
# 
#
# 代码讲解
# ----
#
# 信号量 Semaphore 本质也是一把锁,但是这把锁可以限定允许多个任务同时执行任务,但是不能超出规定的限制,下面的代码参数 5 就代表可以执行 5 个任务,如果第 6 个任务要执行,必须等 5 个任务中的一个结束,然后第六个才能进入执行。
#
# ```
# smph = Semaphore(5)
# ```
#
# 这有点像进程池,只不过进程池规定了进程数量,多个任务进入进程池只能有数量一定的进程进行处理。,但是 Semaphore 可以产生多个线程。
# 线程 Queue
# ========
#
# 队列 Queue
# --------
#
# ### 代码演示
# +
import queue
q = queue.Queue()
q.put('1')
q.put(1)
q.put({'a': 1})
print(q.get())
print(q.get())
print(q.get())
# -
# ### 代码讲解
#
# 1. 先进先出
# 2. 可以存放任意类型数据
#
# 堆栈 Queue
# --------
#
# ### 代码演示
# +
import queue
q = queue.LifoQueue()
q.put(1)
q.put('1')
q.put({'a': 1})
print(q.get())
print(q.get())
print(q.get())
# -
# ### 代码讲解
#
# 1. 可以存放任意数据类型
# 2. Lifo 代表后进先出
#
# 优先级 Queue
# ---------
#
# ### 代码演示
# +
import queue
q = queue.PriorityQueue()
q.put((10, 'Q'))
q.put((30, 'Z'))
q.put((20, 'A'))
print(q.get())
print(q.get())
print(q.get())
# -
# ### 代码讲解
#
# 1. 存放的数据是元组类型,带有优先级数字越小优先级越高。
# 2. 数据优先级高的优先被取出。
# 3. 用于 VIP 用户数据优先被取出场景,因为上面两种都要挨个取出。
# Event
# =====
#
# 代码演示
# ----
#
# ```py
# from threading import Thread, Event, currentThread
# import time
#
# e = Event()
#
#
# def traffic_lights():
# time.sleep(5)
# e.set()
#
#
# def cars():
# print('\033[45m%s\033[0m is waiting' % currentThread().name)
# e.wait()
# print('\033[45m%s\033[0m is running' % currentThread().name)
#
#
# if __name__ == '__main__':
# for i in range(10):
# t = Thread(target=cars, )
# t.start()
# traffic_lights = Thread(target=traffic_lights, )
# traffic_lights.start()
# ```
#
# ### 代码讲解
#
# 首先创建 10 个线程代表 10 辆车正在等信号灯,创建 1 个线程代表信号灯,当 10 辆汽车被创建后就等着信号灯发信号起跑,当遇到 e.wait() 时程序被挂起,等待信号灯变绿,而 e.set() 就是来改变这个状态让信号灯变绿,当 e.set 被设置后 cars 等到了信号,就可以继续往后跑了,代码可以继续执行了。e.set() 默认 False,e.set() 调用后值变为 True,e.wait() 接收到后程序由挂起变为可执行。
#
# ### 应用场景
#
# #### 代码演示
#
# ```py
# from threading import Thread, Event, currentThread
# import time
#
# e = Event()
#
#
# def check_sql():
# print('%s is checking mySQL' % currentThread().name)
# time.sleep(5)
# e.set()
#
#
# def link_sql():
# count = 1
# while not e.is_set():#e.isSet是一个绑定方法,自带布尔值为True,e.is_set()默认值为False
# e.wait(timeout=1)
# print('%s is trying %s' % (currentThread().name, count))
# if count > 3:
# raise ConnectionError('连接超时')
# count += 1
# print('%s is connecting' % currentThread().name)
#
#
# if __name__ == '__main__':
# t_check = Thread(target=check_sql, )
# t_check.start()
# for i in range(3):
# t_link = Thread(target=link_sql, )
# t_link.start()
# ```
#
# #### 代码讲解
#
# 1. 数据库远程连接
# 2. e.isSet 是一个绑定方法,自带布尔值为 True,e.is_set() 默认值为 False
# 定时器
# ===
#
# 代码演示
# ----
# +
from threading import Timer
import time
def deal_task(n):
print('%s 我被执行了~' % n)
print(int(time.time()))
t = Timer(3, deal_task, args=(10,))
t.start()
t.join()
print(int(time.time()))
# -
# 代码讲解
# ----
#
# 注意传参时必须是元组形式
| 并发&异步/多线程常用场景.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Apply convolutional neural network to MNIST hand-written digits dataset
# +
# As usual, a bit of setup
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
plt.rcParams['figure.figsize'] = (10.0, 8.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# for auto-reloading external modules
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
# %load_ext autoreload
# %autoreload 2
def rel_error(x, y):
""" returns relative error """
return np.max(np.abs(x - y) / (np.maximum(1e-8, np.abs(x) + np.abs(y))))
# +
# Copyright 2015 Google Inc. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
"""Simple, end-to-end, LeNet-5-like convolutional MNIST model example.
This should achieve a test error of 0.7%. Please keep this model as simple and
linear as possible, it is meant as a tutorial for simple convolutional models.
Run with --self_test on the command line to execute a short self-test.
"""
import gzip
import os
import sys
import time
import numpy
from six.moves import urllib
from six.moves import xrange # pylint: disable=redefined-builtin
import tensorflow as tf
SOURCE_URL = 'http://yann.lecun.com/exdb/mnist/'
WORK_DIRECTORY = 'data'
IMAGE_SIZE = 28
NUM_CHANNELS = 1
PIXEL_DEPTH = 255
NUM_LABELS = 10
VALIDATION_SIZE = 5000 # Size of the validation set.
SEED = 66478 # Set to None for random seed.
BATCH_SIZE = 64
NUM_EPOCHS = 10
EVAL_BATCH_SIZE = 64
EVAL_FREQUENCY = 100 # Number of steps between evaluations.
tf.app.flags.DEFINE_boolean("self_test", False, "True if running a self test.")
FLAGS = tf.app.flags.FLAGS
# +
def maybe_download(filename):
"""Download the data from Yann's website, unless it's already here."""
if not tf.gfile.Exists(WORK_DIRECTORY):
tf.gfile.MakeDirs(WORK_DIRECTORY)
filepath = os.path.join(WORK_DIRECTORY, filename)
if not tf.gfile.Exists(filepath):
filepath, _ = urllib.request.urlretrieve(SOURCE_URL + filename, filepath)
with tf.gfile.GFile(filepath) as f:
size = f.Size()
print(('Successfully downloaded', filename, size, 'bytes.'))
return filepath
def extract_data(filename, num_images):
"""Extract the images into a 4D tensor [image index, y, x, channels].
Values are rescaled from [0, 255] down to [-0.5, 0.5].
"""
print(('Extracting', filename))
with gzip.open(filename) as bytestream:
bytestream.read(16)
buf = bytestream.read(IMAGE_SIZE * IMAGE_SIZE * num_images)
data = numpy.frombuffer(buf, dtype=numpy.uint8).astype(numpy.float32)
data = (data - (PIXEL_DEPTH / 2.0)) / PIXEL_DEPTH
data = data.reshape(num_images, IMAGE_SIZE, IMAGE_SIZE, 1)
return data
def extract_labels(filename, num_images):
"""Extract the labels into a vector of int64 label IDs."""
print(('Extracting', filename))
with gzip.open(filename) as bytestream:
bytestream.read(8)
buf = bytestream.read(1 * num_images)
labels = numpy.frombuffer(buf, dtype=numpy.uint8).astype(numpy.int64)
return labels
def fake_data(num_images):
"""Generate a fake dataset that matches the dimensions of MNIST."""
data = numpy.ndarray(
shape=(num_images, IMAGE_SIZE, IMAGE_SIZE, NUM_CHANNELS),
dtype=numpy.float32)
labels = numpy.zeros(shape=(num_images,), dtype=numpy.int64)
for image in range(num_images):
label = image % 2
data[image, :, :, 0] = label - 0.5
labels[image] = label
return data, labels
def error_rate(predictions, labels):
"""Return the error rate based on dense predictions and sparse labels."""
return 100.0 - (
100.0 *
numpy.sum(numpy.argmax(predictions, 1) == labels) /
predictions.shape[0])
def load_test_data():
print('Running self-test.')
train_data, train_labels = fake_data(256)
validation_data, validation_labels = fake_data(EVAL_BATCH_SIZE)
test_data, test_labels = fake_data(EVAL_BATCH_SIZE)
return {
'X_train': train_data,
'y_train': train_labels,
'X_test': test_data,
'y_test': test_labels,
'X_val': validation_data,
'y_val': validation_labels,
}
def load_data():
# Get the data.
train_data_filename = maybe_download('train-images-idx3-ubyte.gz')
train_labels_filename = maybe_download('train-labels-idx1-ubyte.gz')
test_data_filename = maybe_download('t10k-images-idx3-ubyte.gz')
test_labels_filename = maybe_download('t10k-labels-idx1-ubyte.gz')
# Extract it into numpy arrays.
train_data = extract_data(train_data_filename, 60000)
train_labels = extract_labels(train_labels_filename, 60000)
test_data = extract_data(test_data_filename, 10000)
test_labels = extract_labels(test_labels_filename, 10000)
# Generate a validation set.
validation_data = train_data[:VALIDATION_SIZE, ...]
validation_labels = train_labels[:VALIDATION_SIZE]
train_data = train_data[VALIDATION_SIZE:, ...]
train_labels = train_labels[VALIDATION_SIZE:]
data = {
'X_train': train_data,
'y_train': train_labels,
'X_test': test_data,
'y_test': test_labels,
'X_val': validation_data,
'y_val': validation_labels,
}
for k, v in sorted(data.items()):
print(('%s: ' % k, v.shape))
return data
# -
data = load_data()
# # load data and visualization
# +
num_epochs = 1
num_epochs = NUM_EPOCHS
train_size = data['y_train'].shape[0]
classes = set(data['y_train'])
num_classes = len(classes)
samples_per_class = 7
for y, cls in enumerate(classes):
idxs = np.flatnonzero(data['y_train'] == y)
idxs = np.random.choice(idxs, samples_per_class, replace=False)
for i, idx in enumerate(idxs):
plt_idx = i * num_classes + y + 1
plt.subplot(samples_per_class, num_classes, plt_idx)
plt.imshow(data['X_train'][idx].reshape((28, 28)))
plt.axis('off')
if i == 0:
plt.title(cls)
plt.show()
# -
# # data node, parameter initialization and the model
# +
# This is where training samples and labels are fed to the graph.
# These placeholder nodes will be fed a batch of training data at each
# training step using the {feed_dict} argument to the Run() call below.
train_data_node = tf.placeholder(
tf.float32,
shape=(BATCH_SIZE, IMAGE_SIZE, IMAGE_SIZE, NUM_CHANNELS))
train_labels_node = tf.placeholder(tf.int64, shape=(BATCH_SIZE,))
eval_data_node = tf.placeholder(
tf.float32,
shape=(EVAL_BATCH_SIZE, IMAGE_SIZE, IMAGE_SIZE, NUM_CHANNELS))
# The variables below hold all the trainable weights. They are passed an
# initial value which will be assigned when we call:
# {tf.initialize_all_variables().run()}
conv1_weights = tf.Variable(
tf.truncated_normal([5, 5, NUM_CHANNELS, 32], # 5x5 filter, depth 32.
stddev=0.1,
seed=SEED))
conv1_biases = tf.Variable(tf.zeros([32]))
conv2_weights = tf.Variable(
tf.truncated_normal([5, 5, 32, 64],
stddev=0.1,
seed=SEED))
conv2_biases = tf.Variable(tf.constant(0.1, shape=[64]))
fc1_weights = tf.Variable( # fully connected, depth 512.
tf.truncated_normal(
[IMAGE_SIZE // 4 * IMAGE_SIZE // 4 * 64, 512],
stddev=0.1,
seed=SEED))
fc1_biases = tf.Variable(tf.constant(0.1, shape=[512]))
fc2_weights = tf.Variable(
tf.truncated_normal([512, NUM_LABELS],
stddev=0.1,
seed=SEED))
fc2_biases = tf.Variable(tf.constant(0.1, shape=[NUM_LABELS]))
# We will replicate the model structure for the training subgraph, as well
# as the evaluation subgraphs, while sharing the trainable parameters.
def model(data, train=False):
"""The Model definition."""
# 2D convolution, with 'SAME' padding (i.e. the output feature map has
# the same size as the input). Note that {strides} is a 4D array whose
# shape matches the data layout: [image index, y, x, depth].
conv = tf.nn.conv2d(data,
conv1_weights,
strides=[1, 1, 1, 1],
padding='SAME')
# Bias and rectified linear non-linearity.
relu = tf.nn.relu(tf.nn.bias_add(conv, conv1_biases))
# Max pooling. The kernel size spec {ksize} also follows the layout of
# the data. Here we have a pooling window of 2, and a stride of 2.
pool = tf.nn.max_pool(relu,
ksize=[1, 2, 2, 1],
strides=[1, 2, 2, 1],
padding='SAME')
conv = tf.nn.conv2d(pool,
conv2_weights,
strides=[1, 1, 1, 1],
padding='SAME')
relu = tf.nn.relu(tf.nn.bias_add(conv, conv2_biases))
pool = tf.nn.max_pool(relu,
ksize=[1, 2, 2, 1],
strides=[1, 2, 2, 1],
padding='SAME')
# Reshape the feature map cuboid into a 2D matrix to feed it to the
# fully connected layers.
pool_shape = pool.get_shape().as_list()
reshape = tf.reshape(
pool,
[pool_shape[0], pool_shape[1] * pool_shape[2] * pool_shape[3]])
# Fully connected layer. Note that the '+' operation automatically
# broadcasts the biases.
hidden = tf.nn.relu(tf.matmul(reshape, fc1_weights) + fc1_biases)
# Add a 50% dropout during training only. Dropout also scales
# activations such that no rescaling is needed at evaluation time.
if train:
hidden = tf.nn.dropout(hidden, 0.5, seed=SEED)
return tf.matmul(hidden, fc2_weights) + fc2_biases
# -
# # Build the computation graph
# +
# Training computation: logits + cross-entropy loss.
logits = model(train_data_node, True)
loss = tf.reduce_mean(tf.nn.sparse_softmax_cross_entropy_with_logits(
logits, train_labels_node))
# L2 regularization for the fully connected parameters.
regularizers = (tf.nn.l2_loss(fc1_weights) + tf.nn.l2_loss(fc1_biases) +
tf.nn.l2_loss(fc2_weights) + tf.nn.l2_loss(fc2_biases))
# Add the regularization term to the loss.
loss += 5e-4 * regularizers
# Optimizer: set up a variable that's incremented once per batch and
# controls the learning rate decay.
batch = tf.Variable(0)
# Decay once per epoch, using an exponential schedule starting at 0.01.
learning_rate = tf.train.exponential_decay(
0.01, # Base learning rate.
batch * BATCH_SIZE, # Current index into the dataset.
train_size, # Decay step.
0.95, # Decay rate.
staircase=True)
# Use simple momentum for the optimization.
optimizer = tf.train.MomentumOptimizer(learning_rate,
0.9).minimize(loss,
global_step=batch)
# Predictions for the current training minibatch.
train_prediction = tf.nn.softmax(logits)
# Predictions for the test and validation, which we'll compute less often.
eval_prediction = tf.nn.softmax(model(eval_data_node))
# Small utility function to evaluate a dataset by feeding batches of data to
# {eval_data} and pulling the results from {eval_predictions}.
# Saves memory and enables this to run on smaller GPUs.
def eval_in_batches(data, sess):
"""Get all predictions for a dataset by running it in small batches."""
size = data.shape[0]
if size < EVAL_BATCH_SIZE:
raise ValueError(
"batch size for evals larger than dataset: %d" %
size)
predictions = numpy.ndarray(
shape=(
size,
NUM_LABELS),
dtype=numpy.float32)
for begin in range(0, size, EVAL_BATCH_SIZE):
end = begin + EVAL_BATCH_SIZE
if end <= size:
predictions[begin:end, :] = sess.run(
eval_prediction,
feed_dict={eval_data_node: data[begin:end, ...]})
else:
batch_predictions = sess.run(
eval_prediction,
feed_dict={eval_data_node: data[-EVAL_BATCH_SIZE:, ...]})
predictions[begin:, :] = batch_predictions[begin - size:, :]
return predictions
# -
# # run the computation graph
def train(data, sess):
train_data = data['X_train']
train_labels = data['y_train']
validation_data = data['X_val']
validation_labels = data['y_val']
test_data = data['X_test']
test_labels = data['y_test']
# Create a local session to run the training.
start_time = time.time()
print('Initialized!')
# Loop through training steps.
for step in range(int(num_epochs * train_size) // BATCH_SIZE):
# Compute the offset of the current minibatch in the data.
# Note that we could use better randomization across epochs.
offset = (step * BATCH_SIZE) % (train_size - BATCH_SIZE)
batch_data = train_data[offset:(offset + BATCH_SIZE), ...]
batch_labels = train_labels[offset:(offset + BATCH_SIZE)]
# This dictionary maps the batch data (as a numpy array) to the
# node in the graph it should be fed to.
feed_dict = {train_data_node: batch_data,
train_labels_node: batch_labels}
# Run the graph and fetch some of the nodes.
_, l, lr, predictions = sess.run(
[optimizer, loss, learning_rate, train_prediction],
feed_dict=feed_dict)
if step % EVAL_FREQUENCY == 0:
elapsed_time = time.time() - start_time
start_time = time.time()
print(('Step %d (epoch %.2f), %.1f ms' %
(step, float(step) * BATCH_SIZE / train_size,
1000 * elapsed_time / EVAL_FREQUENCY)))
print(('Minibatch loss: %.3f, learning rate: %.6f' % (l, lr)))
print((
'Minibatch error: %.1f%%' %
error_rate(
predictions,
batch_labels)))
print(('Validation error: %.1f%%' % error_rate(
eval_in_batches(validation_data, sess), validation_labels)))
sys.stdout.flush()
# Finally print the result!
test_labels_predict = eval_in_batches(test_data, sess)
test_error = error_rate(test_labels_predict, test_labels)
print(('Test error: %.1f%%' % test_error))
if FLAGS.self_test:
print(('test_error', test_error))
assert test_error == 0.0, 'expected 0.0 test_error, got %.2f' % (
test_error,)
sess = tf.Session()
# Run all the initializers to prepare the trainable parameters.
tf.initialize_all_variables().run(session=sess)
train(data, sess)
saver = tf.train.Saver()
saver.save(sess, 'mnist-sess')
test_sample_idxs = np.random.choice(data['X_test'].shape[0], samples_per_class * num_classes)
test_sample_images = data['X_test'][test_sample_idxs, ...]
test_sample_predict = eval_in_batches(test_sample_images, sess)
for i, sample_predict in enumerate(test_sample_idxs):
plt_idx = i + 1
plt.subplot(samples_per_class, num_classes, plt_idx)
plt.imshow(test_sample_images[i].reshape((28, 28)))
plt.axis('off')
plt.title(np.argmax(test_sample_predict[i]))
plt.show()
# +
# load dataset from kaggle for education purpose, not competition
import pandas as pd
IMAGE_TO_DISPLAY = 8
# read training data from CSV file
train_data_kaggle = pd.read_csv('~/Documents/mine/data/mnist/train.csv')
test_data_kaggle = pd.read_csv('~/Documents/mine/data/mnist/test.csv')
print(('data({0[0]},{0[1]})'.format(train_data_kaggle.shape)))
print((train_data_kaggle.head()))
# -
# # preprocess data
# +
images = train_data_kaggle.iloc[:,1:].values
images = images.astype(np.float).reshape(
images.shape[0], IMAGE_SIZE, IMAGE_SIZE, NUM_CHANNELS)
# convert from [0:255] => [0.0:1.0]
images = (images - (PIXEL_DEPTH / 2.0)) / PIXEL_DEPTH
print(('images({0})'.format(images.shape)))
test_images = test_data_kaggle.values.reshape(
-1, IMAGE_SIZE, IMAGE_SIZE, NUM_CHANNELS)
test_images = test_images.astype(np.float)
# convert from [0:255] => [0.0:1.0]
test_images = (test_images - (PIXEL_DEPTH / 2.0)) / PIXEL_DEPTH
print(('images({0[0]},{0[1]},{0[2]},{0[3]})'.format(test_images.shape)))
# +
image_size = images.shape[1]
print(('image_size => {0}'.format(image_size)))
# in this case all images are square
image_width = image_height = np.ceil(np.sqrt(image_size)).astype(np.uint8)
print(('image_width => {0}\nimage_height => {1}'.format(image_width,image_height)))
# +
labels_flat = train_data_kaggle[[0]].values.ravel()
print(('labels_flat({0})'.format(len(labels_flat))))
print(('labels_flat[{0}] => {1}'.format(IMAGE_TO_DISPLAY,labels_flat[IMAGE_TO_DISPLAY])))
# -
# For most classification problems "one-hot vectors" are used. A one-hot vector is a vector that contains a single element equal to $1$ and the rest of the elements equal to $0$. In this case, the $n$th digit is represented as a zero vector with $1$ in the $n$th position.
# However, tensorflow supports dense labels, so we don't need to convert it
#
# +
# convert class labels from scalars to one-hot vectors
# 0 => [1 0 0 0 0 0 0 0 0 0]
# 1 => [0 1 0 0 0 0 0 0 0 0]
# ...
# 9 => [0 0 0 0 0 0 0 0 0 1]
def dense_to_one_hot(labels_dense, num_classes):
num_labels = labels_dense.shape[0]
index_offset = np.arange(num_labels) * num_classes
labels_one_hot = np.zeros((num_labels, num_classes))
labels_one_hot.flat[index_offset + labels_dense.ravel()] = 1
return labels_one_hot
labels_count = np.unique(labels_flat).shape[0]
# labels = dense_to_one_hot(labels_flat, labels_count)
labels = labels_flat
# labels = labels.astype(np.uint8)
print(('labels({0})'.format(labels.shape)))
print(('labels[{0}] => {1}'.format(IMAGE_TO_DISPLAY,labels[IMAGE_TO_DISPLAY])))
# +
# split data into training & validation
validation_images = images[:VALIDATION_SIZE]
validation_labels = labels[:VALIDATION_SIZE]
train_images = images[VALIDATION_SIZE:]
train_labels = labels[VALIDATION_SIZE:]
print(('train_images({0[0]},{0[1]})'.format(train_images.shape)))
print(('validation_images({0[0]},{0[1]})'.format(validation_images.shape)))
# -
labels_predict_kaggle = eval_in_batches(train_images, sess)
print(('trains error: %.1f%%' % error_rate(labels_predict_kaggle, train_labels)))
# # predict and write predictions to file
labels_predict_kaggle_test = eval_in_batches(test_images, sess)
with open('submit.csv', 'w') as f:
f.write('ImageId,Label\n')
for i in range(labels_predict_kaggle_test.shape[0]):
f.write('%d,%d\n' % (i+1, np.argmax(labels_predict_kaggle_test[i])))
| notebooks/ConvNet-MNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + tags=["remove-input"]
from datascience import *
path_data = '../data/'
import matplotlib
matplotlib.use('Agg')
# %matplotlib inline
import matplotlib.pyplot as plots
plots.style.use('fivethirtyeight')
import numpy as np
# -
# # The Monty Hall Problem
# This [problem](https://en.wikipedia.org/wiki/Monty_Hall_problem) has flummoxed many people over the years, [mathematicians included](https://web.archive.org/web/20140413131827/http://www.decisionsciences.org/DecisionLine/Vol30/30_1/vazs30_1.pdf). Let's see if we can work it out by simulation.
#
# The setting is derived from a television game show called "Let's Make a Deal". Monty Hall hosted this show in the 1960's, and it has since led to a number of spin-offs. An exciting part of the show was that while the contestants had the chance to win great prizes, they might instead end up with "zonks" that were less desirable. This is the basis for what is now known as *the Monty Hall problem*.
#
# The setting is a game show in which the contestant is faced with three closed doors. Behind one of the doors is a fancy car, and behind each of the other two there is a goat. The contestant doesn't know where the car is, and has to attempt to find it under the following rules.
#
# - The contestant makes an initial choice, but that door isn't opened.
# - At least one of the other two doors must have a goat behind it. Monty opens one of these doors to reveal a goat, displayed in all its glory in [Wikipedia](https://en.wikipedia.org/wiki/Monty_Hall_problem):
#
# 
#
# - There are two doors left, one of which was the contestant's original choice. One of the doors has the car behind it, and the other one has a goat. The contestant now gets to choose which of the two doors to open.
#
# The contestant has a decision to make. Which door should she choose to open, if she wants the car? Should she stick with her initial choice, or switch to the other door? That is the Monty Hall problem.
# ## The Solution
#
# In any problem involving chances, the assumptions about randomness are important. It's reasonable to assume that there is a 1/3 chance that the contestant's initial choice is the door that has the car behind it.
#
# The solution to the problem is quite straightforward under this assumption, though the straightforward solution doesn't convince everyone. Here it is anyway.
#
# - The chance that the car is behind the originally chosen door is 1/3.
# - The car is behind either the originally chosen door or the door that remains. It can't be anywhere else.
# - Therefore, the chance that the car is behind the door that remains is 2/3.
# - Therefore, the contestant should switch.
#
# That's it. End of story.
#
# Not convinced? Then let's simulate the game and see how the results turn out.
# ## Simulation ##
# The simulation will be more complex that those we have done so far. Let's break it down.
#
# **Step 1: What to Simulate**
#
# For each play we will simulate what's behind all three doors:
# - the one the contestant first picks
# - the one that Monty opens
# - the remaining door
#
# So we will be keeping track of three quantitites, not just one.
# **Step 2: Simulating One Play**
#
# As is often the case in simulating a game, the bulk of the work consists of simulating one play of the game. This involves several pieces.
#
# **The goats:** We start by setting up an array `goats` that contains unimaginative names for the two goats.
goats = make_array('first goat', 'second goat')
# To help Monty conduct the game, we are going to have to identify which goat is selected and which one is revealed behind the open door. The function `other_goat` takes one goat and returns the other.
def other_goat(x):
if x == 'first goat':
return 'second goat'
elif x == 'second goat':
return 'first goat'
other_goat('first goat'), other_goat('second goat'), other_goat('watermelon')
# The string `watermelon` is not the name of one of the goats, so when `watermelon` is the input then `other_goat` does nothing.
# **The options:** The array `hidden_behind_doors` contains the three things that are
# behind the doors.
hidden_behind_doors = np.append(goats, 'car')
hidden_behind_doors
# We are now ready to simulate one play. To do this, we will define a function `monty_hall_game` that takes no arguments. When the function is called, it plays Monty's game once and returns a list consisting of:
#
# - the contestant's guess
# - what Monty reveals when he opens a door
# - what remains behind the other door
#
# The game starts with the contestant choosing one door at random. In doing so, the contestant makes a random choice from among the first goat, the second goat, and the car.
#
# If the contestant happens to pick one of the goats, then the other goat is revealed and the car is behind the remaining door.
#
# If the contestant happens to pick the car, then Monty reveals one of the goats and the other goat is behind the remaining door.
def monty_hall_game():
"""Return
[contestant's guess, what Monty reveals, what remains behind the other door]"""
contestant_guess = np.random.choice(hidden_behind_doors)
if contestant_guess == 'first goat':
return [contestant_guess, 'second goat', 'car']
if contestant_guess == 'second goat':
return [contestant_guess, 'first goat', 'car']
if contestant_guess == 'car':
revealed = np.random.choice(goats)
return [contestant_guess, revealed, other_goat(revealed)]
# Let's play! Run the cell several times and see how the results change.
monty_hall_game()
# **Step 3: Number of Repetitions**
#
# To gauge the frequency with which the different results occur, we have to play the game many times and collect the results. Let's run 10,000 repetitions.
#
# **Step 4: Simulating Multiple Repetitions**
#
# It's time to run the whole simulation. But unlike our previous simulations in which each simulation produced a single value, in this example we simulate a list of three values each time.
#
# We will treat this simulated list as a row of a table. So instead of starting with an empty array and augmenting it with each new simulated value, we will start with an empty table and
# augment the table with each new simulated row. Each row will contain the complete result of one play.
#
# One way to grow a table by adding a new row is to use the `append` method. If `my_table` is a table and `new_row` is a list containing the entries in a new row, then `my_table.append(new_row)` adds the new row to the bottom of `my_table`.
#
# Note that `append` does not create a new table. It changes `my_table` to have one more row than it did before.
# We will start with a collection table `games` that has three empty columns. We can do this by just specifying a list of the column labels in the same order in which `monty_hall_game` returns the result of one game.
#
# Now we can add 10,000 rows to `games`. Each row will represent the result of one play of Monty's game.
# +
# empty collection table
games = Table(['Guess', 'Revealed', 'Remaining'])
# Play the game 10000 times and
# record the results in the table games
for i in np.arange(10000):
games.append(monty_hall_game())
# -
# The simulation is done. Notice how short the code is. The majority of the work was done in simulating the outcome of one game.
games.show(3)
# ## Visualization
#
# To see whether the contestant should stick with her original choice or switch, let's see how frequently the car is behind each of her two options.
#
# It is no surprise that the three doors appear about equally often as the contestant's original guess.
original_choice = games.group('Guess')
original_choice
# Once Monty has eliminated a goat, how often is the car behind the remaining door?
remaining_door = games.group('Remaining')
remaining_door
# As our earlier solution said, the car is behind the remaining door two-thirds of the time, to a pretty good approximation. The contestant is twice as likely to get the car if she switches than if she sticks with her original choice.
#
# To see this graphically, we can join the two tables above and draw overlaid bar charts.
joined = original_choice.join('Guess', remaining_door, 'Remaining')
combined = joined.relabeled(0, 'Item').relabeled(1, 'Original Door').relabeled(2, 'Remaining Door')
combined
combined.barh(0)
# Notice how the three blue bars are almost equal – the original choice is equally likely to be any of the three available items. But the gold bar corresponding to `Car` is twice as long as the blue.
#
# The simulation confirms that the contestant is twice as likely to win if she switches.
| Mathematics/Statistics/Statistics and Probability Python Notebooks/Computational and Inferential Thinking - The Foundations of Data Science (book)/Notebooks - by chapter/9. Randomness and Probabiltities/4. Monty_Hall_Problem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Setting up Python for machine learning: scikit-learn and IPython Notebook
# *From the video series: [Introduction to machine learning with scikit-learn](https://github.com/justmarkham/scikit-learn-videos)*
# ## Agenda
#
# - What are the benefits and drawbacks of scikit-learn?
# - How do I install scikit-learn?
# - How do I use the IPython Notebook?
# - What are some good resources for learning Python?
# 
# ## Benefits and drawbacks of scikit-learn
#
# ### Benefits:
#
# - **Consistent interface** to machine learning models
# - Provides many **tuning parameters** but with **sensible defaults**
# - Exceptional **documentation**
# - Rich set of functionality for **companion tasks**
# - **Active community** for development and support
#
# ### Potential drawbacks:
#
# - Harder (than R) to **get started with machine learning**
# - Less emphasis (than R) on **model interpretability**
#
# ### Further reading:
#
# - <NAME>: [Six reasons why I recommend scikit-learn](http://radar.oreilly.com/2013/12/six-reasons-why-i-recommend-scikit-learn.html)
# - scikit-learn authors: [API design for machine learning software](http://arxiv.org/pdf/1309.0238v1.pdf)
# - Data School: [Should you teach Python or R for data science?](http://www.dataschool.io/python-or-r-for-data-science/)
# 
# ## Installing scikit-learn
#
# **Option 1:** [Install scikit-learn library](http://scikit-learn.org/stable/install.html) and dependencies (NumPy and SciPy)
#
# **Option 2:** [Install Anaconda distribution](https://www.continuum.io/downloads) of Python, which includes:
#
# - Hundreds of useful packages (including scikit-learn)
# - IPython and IPython Notebook
# - conda package manager
# - Spyder IDE
# 
# ## Using the IPython Notebook
#
# ### Components:
#
# - **IPython interpreter:** enhanced version of the standard Python interpreter
# - **Browser-based notebook interface:** weave together code, formatted text, and plots
#
# ### Installation:
#
# - **Option 1:** Install [IPython](http://ipython.org/install.html) and the [notebook](https://jupyter.readthedocs.io/en/latest/install.html)
# - **Option 2:** Included with the Anaconda distribution
#
# ### Launching the Notebook:
#
# - Type **ipython notebook** at the command line to open the dashboard
# - Don't close the command line window while the Notebook is running
#
# ### Keyboard shortcuts:
#
# **Command mode** (gray border)
#
# - Create new cells above (**a**) or below (**b**) the current cell
# - Navigate using the **up arrow** and **down arrow**
# - Convert the cell type to Markdown (**m**) or code (**y**)
# - See keyboard shortcuts using **h**
# - Switch to Edit mode using **Enter**
#
# **Edit mode** (green border)
#
# - **Ctrl+Enter** to run a cell
# - Switch to Command mode using **Esc**
#
# ### IPython and Markdown resources:
#
# - [nbviewer](http://nbviewer.jupyter.org/): view notebooks online as static documents
# - [IPython documentation](http://ipython.readthedocs.io/en/stable/): focuses on the interpreter
# - [IPython Notebook tutorials](http://jupyter.readthedocs.io/en/latest/content-quickstart.html): in-depth introduction
# - [GitHub's Mastering Markdown](https://guides.github.com/features/mastering-markdown/): short guide with lots of examples
# ## Resources for learning Python
#
# - [Codecademy's Python course](https://www.codecademy.com/learn/python): browser-based, tons of exercises
# - [DataQuest](https://www.dataquest.io/): browser-based, teaches Python in the context of data science
# - [Google's Python class](https://developers.google.com/edu/python/): slightly more advanced, includes videos and downloadable exercises (with solutions)
# - [Python for Informatics](http://www.pythonlearn.com/): beginner-oriented book, includes slides and videos
# ## Comments or Questions?
#
# - Email: <<EMAIL>>
# - Website: http://dataschool.io
# - Twitter: [@justmarkham](https://twitter.com/justmarkham)
from IPython.core.display import HTML
def css_styling():
styles = open("styles/custom.css", "r").read()
return HTML(styles)
css_styling()
| scikit-learn-videos/02_machine_learning_setup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="fcFjxdPMktzm"
# #Section 1
# + id="A-ZrAb4fZGiL"
import numpy as np
import matplotlib.pyplot as plt
# + id="WKNVuHRNDdGW"
class NeuralNetwork(object):
def __init__(self, layers = [1 , 10, 1], activations=['sigmoid', 'sigmoid']):
self.layers = layers
self.activations = activations
self.weights = []
self.biases = []
for i in range(len(layers)-1):
self.weights.append(np.random.randn(layers[i+1], layers[i]))
self.biases.append(np.random.randn(layers[i+1], 1))
def fit(self, x):
a = np.copy(x)
z_s = []
a_s = [a]
for i in range(len(self.weights)):
activation_function = self.getActivationFunction(self.activations[i])
z_s.append(self.weights[i].dot(a) + self.biases[i])
a = activation_function(z_s[-1])
a_s.append(a)
return (z_s, a_s)
def backpropagation(self,y, z_s, a_s):
dw = []
db = []
errors = [None] * len(self.weights)
errors[-1] = ((y-a_s[-1])*(self.getDerivitiveActivationFunction(self.activations[-1]))(z_s[-1]))
for i in reversed(range(len(errors)-1)):
errors[i] = self.weights[i+1].T.dot(errors[i+1])*(self.getDerivitiveActivationFunction(self.activations[i])(z_s[i]))
batch_size = y.shape[1]
db = [d.dot(np.ones((batch_size,1)))/float(batch_size) for d in errors]
dw = [d.dot(a_s[i].T)/float(batch_size) for i,d in enumerate(errors)]
return dw, db
def train(self, x, y, batch_size=512, epochs=1000, lr = 0.1):
for e in range(epochs):
i=0
while(i<len(y)):
x_batch = x[i:i+batch_size]
y_batch = y[i:i+batch_size]
i = i+batch_size
z_s, a_s = self.fit(x_batch)
dw, db = self.backpropagation(y_batch, z_s, a_s)
self.weights = [w+lr*dweight for w,dweight in zip(self.weights, dw)]
self.biases = [w+lr*dbias for w,dbias in zip(self.biases, db)]
if e == epochs - 1 : print("loss = {}".format(np.linalg.norm(a_s[-1]-y_batch) ))
@staticmethod
def getActivationFunction(name):
return lambda x : np.exp(x)/(1+np.exp(x))
@staticmethod
def getDerivitiveActivationFunction(name):
sig = lambda x : np.exp(x)/(1+np.exp(x))
return lambda x :sig(x)*(1-sig(x))
# + id="CVKsfvQkY6zI"
def main(layer , activations , X, Y, X_test, Y_test, epochs = 50, lr = 0.5, batch_size = 512):
nn = NeuralNetwork(layer,activations)
nn.train(X, Y, epochs, batch_size, lr )
_, train = nn.fit(X)
plt.scatter(X.flatten(), Y.flatten())
plt.scatter(X.flatten(), train[-1].flatten())
plt.show()
_, prediction = nn.fit(X_test)
plt.plot(X_test.flatten(), Y_test.flatten(), label='Actual')
plt.plot(X_test.flatten() ,prediction[-1].flatten(), label='Predicted')
plt.legend()
plt.show()
# + [markdown] id="sS2Hf5hSk9VJ"
# #Section 2
# + [markdown] id="YMm-BQQgdUv6"
# #Sin(x)
# + id="sF3ivsnSf3_W" colab={"base_uri": "https://localhost:8080/", "height": 529} outputId="743ebd77-23f2-4571-b3a2-e0cfecc219a5"
x = 2*np.pi*np.random.rand(1000).reshape(1, -1)
y = np.sin(x)
x_test = np.arange(-2*np.pi, 2*np.pi, 0.1).reshape(1, -1)
y_test = np.sin(x_test)
layer = [1, 10, 1]
activations =['sigmoid', 'sigmoid']
main(layer , activations, x, y, x_test, y_test, 1500, 0.5)
# + id="0lkqKpLxgv1C" colab={"base_uri": "https://localhost:8080/", "height": 512} outputId="c1294940-acd6-4822-e583-b46b41224d2d"
x = 2*np.pi*np.random.rand(1000).reshape(1, -1)
y = np.sin(x)
x_test = np.arange(0, 10*np.pi, 0.1).reshape(1, -1)
y_test = np.sin(x_test)
layer = [1, 100, 1]
activations =['sigmoid', 'sigmoid']
main(layer , activations, x, y, x_test, y_test, 1500, 0.5)
# + id="hUSnZTd3g2iG" colab={"base_uri": "https://localhost:8080/", "height": 529} outputId="d446c737-53ae-46b9-85a3-07faa3c8b76f"
x = 2*np.pi*np.random.rand(5000).reshape(1, -1)
y = np.sin(x)
x_test = np.arange(-2*np.pi, 2*np.pi, 0.1).reshape(1, -1)
y_test = np.sin(x_test)
layer = [1, 128, 64, 1]
activations =['sigmoid', 'sigmoid', 'sigmoid']
main(layer , activations, x, y, x_test, y_test, 1000, 0.6)
# + id="pjsT9TkYhFRK" colab={"base_uri": "https://localhost:8080/", "height": 512} outputId="99857a7f-b537-4737-da1b-ad5be5b6c018"
x = 4*np.pi*np.random.rand(4000).reshape(1, -1)
y = np.sin(x)
x_test = np.arange(-10*np.pi, 10*np.pi, 0.1).reshape(1, -1)
y_test = np.sin(x_test)
layer = [1, 128, 64, 32, 1]
activations =['sigmoid', 'sigmoid', 'sigmoid', 'sigmoid']
main(layer , activations, x, y, x_test, y_test, 5000, 0.6)
# + [markdown] id="Hzq6boYzlLbX"
# ## Same as Keras
# + id="jU_tLmnxkOzU" colab={"base_uri": "https://localhost:8080/", "height": 512} outputId="228ce5f6-427c-4313-ffce-be27bfc1de6a"
x = 4*np.pi*np.random.rand(10000).reshape(1, -1)
y = np.sin(x)
x_test = np.arange(-10*np.pi, 10*np.pi, 0.1).reshape(1, -1)
y_test = np.sin(x_test)
layer = [1, 128, 64, 16, 1]
activations =['sigmoid', 'sigmoid', 'sigmoid', 'sigmoid']
main(layer , activations, x, y, x_test, y_test, 1000, 0.6)
# + [markdown] id="ETDRoGT5dX1g"
# #X^2
# + [markdown] id="bihgCj6EC8-N"
# ##same as keras
# + id="iSBFhMJv5CS5" colab={"base_uri": "https://localhost:8080/", "height": 529} outputId="029617b1-6091-48e0-bb15-d92f064629df"
x = np.random.uniform(-1, 1, 2000).reshape(1, -1)
y = x**2
x_test = np.arange(-2, 2, 0.1).reshape(1, -1)
y_test = x_test**2
layer = [1, 128, 64, 16, 1]
activations =['sigmoid', 'sigmoid', 'sigmoid', 'sigmoid']
main(layer , activations, x, y, x_test, y_test, 1000, 0.6)
# + id="OS4RN1-KdCwU" colab={"base_uri": "https://localhost:8080/", "height": 529} outputId="bce13ccc-14d3-4a6f-c3bc-e4c69755c9e3"
x = np.random.uniform(-1, 1, 2000).reshape(1, -1)
y = x**2
x_test = np.arange(-2, 2, 0.1).reshape(1, -1)
y_test = x_test**2
layer = [1, 512, 512, 512, 1]
activations =['sigmoid', 'sigmoid', 'sigmoid', 'sigmoid']
main(layer , activations, x, y, x_test, y_test, 10000, 0.5)
# + [markdown] id="WDkLL2m1LT9d"
# #sin(x) + x
# + id="TJuUEY5iLYnR" colab={"base_uri": "https://localhost:8080/", "height": 532} outputId="eaa1fdc8-184c-460a-a182-33731a1b8b8f"
x = np.random.uniform(-1, 1, 5000).reshape(1, -1)
y = x + np.sin(x)
x_test = np.arange(-2, 2, 0.1).reshape(1, -1)
y_test = x_test + np.sin(x_test)
layer = [1, 128, 64, 32, 1]
activations =['sigmoid', 'sigmoid', 'sigmoid', 'sigmoid']
main(layer , activations, x, y, x_test, y_test, 2000, 0.5)
# + id="Icg7M-HsrkWl"
x = np.random.uniform(-5, 5, 5000).reshape(1, -1)
y = x + np.sin(x)
x_test = np.arange(-2, 2, 0.1).reshape(1, -1)
y_test = x_test + np.sin(x_test)
layer = [1, 128, 64, 32, 1]
activations =['sigmoid', 'sigmoid', 'sigmoid', 'sigmoid']
main(layer , activations, x, y, x_test, y_test, 2000, 0.5)
# + id="-AG97O9bcIk7" colab={"base_uri": "https://localhost:8080/", "height": 532} outputId="83e9526d-ff24-45e3-bae1-c2b28ee2e8fd"
x = np.random.uniform(-1, 1, 8000).reshape(1, -1)
y = x + np.sin(x)
x_test = np.arange(-2, 2, 0.1).reshape(1, -1)
y_test = x_test + np.sin(x_test)
layer = [1, 128, 64, 32, 1]
activations =['sigmoid', 'sigmoid', 'sigmoid', 'sigmoid']
main(layer , activations, x, y, x_test, y_test, 10000, 0.7)
# + [markdown] id="cQM4W8eVMMG8"
# #Section 3 noisy data
# + [markdown] id="3m4KpPskMVET"
# #sin(x) --noisy
# + id="WW5yTTtVMPBA" colab={"base_uri": "https://localhost:8080/", "height": 529} outputId="04fcd069-3c1c-4608-f826-e386ecbf3781"
x = 2*np.pi*np.random.rand(1000).reshape(1, -1)
y = np.sin(x) + np.random.uniform(low=-0.1, high=0.1, size=(len(x)))
x_test = np.arange(0, 2*np.pi, 0.1).reshape(1, -1)
y_test = np.sin(x_test)
layer = [1, 128, 64, 1]
activations =['sigmoid', 'sigmoid', 'sigmoid']
main(layer , activations, x, y, x_test, y_test, 1000, 0.6)
# + id="cyG-lOVUMYtp" colab={"base_uri": "https://localhost:8080/", "height": 529} outputId="08b65fe7-ef48-4c9e-c83a-e6608b930716"
x = 2*np.pi*np.random.rand(1000).reshape(1, -1)
y = np.sin(x) + np.random.uniform(low=-0.7, high=0.7, size=(len(x)))
x_test = np.arange(0, 2*np.pi, 0.1).reshape(1, -1)
y_test = np.sin(x_test)
layer = [1, 128, 64, 1]
activations =['sigmoid', 'sigmoid', 'sigmoid']
main(layer , activations, x, y, x_test, y_test, 1000, 0.6)
# + id="rUvSx48eMbRW" colab={"base_uri": "https://localhost:8080/", "height": 529} outputId="52629748-66c4-4218-99a7-667ca662a495"
x = 2*np.pi*np.random.rand(1000).reshape(1, -1)
y = np.sin(x) + np.random.uniform(low=-9, high=9, size=(len(x)))
x_test = np.arange(0, 2*np.pi, 0.1).reshape(1, -1)
y_test = np.sin(x_test)
layer = [1, 128, 64, 1]
activations =['sigmoid', 'sigmoid', 'sigmoid']
main(layer , activations, x, y, x_test, y_test, 1000, 0.6)
# + [markdown] id="TBXEGO_8MxAi"
# #X^2 -- noisy
# + id="Da6M5WsqMnda" colab={"base_uri": "https://localhost:8080/", "height": 529} outputId="36bb3285-ef0b-4f01-f539-eea584a624f1"
x = np.random.uniform(-1, 1, 2000).reshape(1, -1)
y = x**2 + np.random.uniform(low=-0.1, high=0.1, size=(len(x)))
x_test = np.arange(-2, 2, 0.1).reshape(1, -1)
y_test = x_test**2
layer = [1, 128, 64, 16, 1]
activations =['sigmoid', 'sigmoid', 'sigmoid', 'sigmoid']
main(layer , activations, x, y, x_test, y_test, 1000, 0.6)
# + id="Mdktx8DoTxPy" colab={"base_uri": "https://localhost:8080/", "height": 529} outputId="f3914365-9d8c-48e5-c427-8194af8f01b3"
x = np.random.uniform(-1, 1, 2000).reshape(1, -1)
y = x**2 + np.random.uniform(low=0.5, high=0.5, size=(len(x)))
x_test = np.arange(-2, 2, 0.1).reshape(1, -1)
y_test = x_test**2
layer = [1, 128, 64, 16, 1]
activations =['sigmoid', 'sigmoid', 'sigmoid', 'sigmoid']
main(layer , activations, x, y, x_test, y_test, 1000, 0.6)
# + id="KuEUW42RT05k" colab={"base_uri": "https://localhost:8080/", "height": 529} outputId="990612e0-650e-42de-c1cb-685e541400ab"
x = np.random.uniform(-1, 1, 2000).reshape(1, -1)
y = x**2 + np.random.uniform(low=0.9, high=0.9, size=(len(x)))
x_test = np.arange(-2, 2, 0.1).reshape(1, -1)
y_test = x_test**2
layer = [1, 128, 64, 16, 1]
activations =['sigmoid', 'sigmoid', 'sigmoid', 'sigmoid']
main(layer , activations, x, y, x_test, y_test, 1000, 0.6)
# + [markdown] id="9uHzry_WYwiI"
# #sin(x) + x --noisy
# + id="iZQc_q8OT3r8" colab={"base_uri": "https://localhost:8080/", "height": 529} outputId="28da07a6-ebba-4861-f63a-8e371b90abd1"
x = np.random.uniform(-1, 1, 2000).reshape(1, -1)
y = x + np.sin(x) + np.random.uniform(low=0.9, high=0.9, size=(len(x)))
x_test = np.arange(-2, 2, 0.1).reshape(1, -1)
y_test = x_test + np.sin(x_test)
layer = [1, 512, 255, 128, 1]
activations =['sigmoid', 'sigmoid', 'sigmoid', 'sigmoid']
main(layer , activations, x, y, x_test, y_test, 10000, 0.5)
| MLP_with_Numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Lesson 6 Practice: Data Wrangling
# Use this notebook to follow along with the lesson in the corresponding lesson notebook: [L06-Data_Wrangling-Lesson.ipynb](./L06-Data_Wrangling-Lesson.ipynb).
#
# ## Instructions
# Follow along with the teaching material in the lesson. Throughout the tutorial sections labeled as "Tasks" are interspersed and indicated with the icon: . You should follow the instructions provided in these sections by performing them in the practice notebook. When the tutorial is completed you can turn in the final practice notebook. For each task, use the cell below it to write and test your code. You may add additional cells for any task as needed or desired.
# ## Task 1a: Setup
#
#
# Import the following packages:
# + `pandas` as `pd`
# + `numpy` as `np`
| L06-Data_Wrangling-Practice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Configure plotting in Jupyter
from matplotlib import pyplot as plt
# %matplotlib inline
plt.rcParams.update({
'figure.figsize': (7.5, 7.5),
'axes.spines.right': False,
'axes.spines.left': False,
'axes.spines.top': False,
'axes.spines.bottom': False})
# Seed random number generator
import random
from numpy import random as nprand
seed = hash("Network Science in Python") % 2**32
nprand.seed(seed)
random.seed(seed)
# Import NetworkX
import networkx as nx
# ## Strong and Weak Ties
G = nx.karate_club_graph()
# Annotate with splinter club label
member_club = [
0, 0, 0, 0, 0, 0, 0, 0, 1, 1,
0, 0, 0, 0, 1, 1, 0, 0, 1, 0,
1, 0, 1, 1, 1, 1, 1, 1, 1, 1,
1, 1, 1, 1]
nx.set_node_attributes(G, dict(enumerate(member_club)), 'club')
# Find internal and external edges
internal = [(v, w) for v, w in G.edges if G.nodes[v]['club'] == G.nodes[w]['club']]
external = [(v, w) for v, w in G.edges if G.nodes[v]['club'] != G.nodes[w]['club']]
# Update edge attributes
nx.set_edge_attributes(G, dict((e, 'internal') for e in internal), 'internal')
nx.set_edge_attributes(G, dict((e, 'external') for e in external), 'internal')
# +
def tie_strength(G, v, w):
# Get neighbors of nodes v and w in G
v_neighbors = set(G.neighbors(v))
w_neighbors = set(G.neighbors(w))
# Return size of the set intersection
return 1 + len(v_neighbors & w_neighbors)
strength = dict(
((v,w), tie_strength(G, v, w))
for v, w in G.edges())
# +
def bridge_span(G):
# Get list of edges
edges = G.edges()
# Copy G
G = nx.Graph(G)
# Create result dict
result = dict()
for v, w in edges:
# Temporarily remove edge
G.remove_edge(v, w)
# Find distance with edge removed
try:
d = nx.shortest_path_length(G, v, w)
result[(v, w)] = d
except nx.NetworkXNoPath:
result[(v, w)] = float('inf')
# Restore edge
G.add_edge(v, w)
return result
span = bridge_span(G)
# -
# Order edges by tie strength
ordered_edges = sorted(strength.items(), key=lambda x: x[1])
print('Edge\t Strength\tSpan\tInternal')
# Print 10 strongest
for e, edge_strength in ordered_edges[:10]:
print('{:10}{}\t\t{}\t{}'.format(
str(e), edge_strength, span[e], G.edges[e]['internal']
))
print('...')
# Print 10 weakest
for e, edge_strength in ordered_edges[-10:]:
print('{:10}{}\t\t{}\t{}'.format(
str(e), edge_strength, span[e], G.edges[e]['internal']
))
# ## The Small World Problem
G_small_ring = nx.watts_strogatz_graph(16, 4, 0)
pos = nx.circular_layout(G_small_ring)
nx.draw_networkx(G_small_ring, pos=pos, with_labels=False)
G_ring = nx.watts_strogatz_graph(4000, 10, 0)
nx.average_shortest_path_length(G_ring)
nx.average_clustering(G_ring)
# ### A Real Social Network
# Load data file into network
from pathlib import Path
data_dir = Path('.') / 'data'
G_social = nx.read_edgelist(data_dir / 'mcauley2012' / 'facebook_combined.txt')
nx.average_shortest_path_length(G_social)
nx.average_clustering(G_social)
# # Random Network
G_small_random = nx.watts_strogatz_graph(16, 4, 1)
pos = nx.circular_layout(G_small_random)
nx.draw_networkx(G_small_random, pos=pos, with_labels=False)
G_random = nx.watts_strogatz_graph(4000, 10, 1)
nx.average_shortest_path_length(G_random)
nx.average_clustering(G_random)
# ### Watts-Strogatz Model
path = []
clustering = []
# Try a range of rewiring probabilities
p = [10**(x) for x in range(-6, 1)]
for p_i in p:
path_i = []
clustering_i =[]
# Create 10 models for each probability
for n in range(10):
G = nx.watts_strogatz_graph(1000, 10, p_i)
path_i.append(nx.average_shortest_path_length(G))
clustering_i.append(nx.average_clustering(G))
# Average the properties for each p_i
path.append(sum(path_i) / len(path_i))
clustering.append(sum(clustering_i) / len(clustering_i))
# Plot the results
fig, ax = plt.subplots()
for spine in ax.spines.values():
spine.set_visible(True)
plt.semilogx(p, [x / path[0] for x in path], label='Mean Path / Initial')
plt.semilogx(p, [x / clustering[0] for x in clustering], label='Clustering / Initial')
plt.tick_params(axis='both', which='major', labelsize=16)
plt.xlabel('Rewiring Probability p', fontsize=16)
plt.legend(fontsize=16)
# ## Contagion
# ### Simple Contagion
def propagate_simple(G):
to_infect = set([])
# Find infected nodes
for v in G.nodes():
if G.nodes[v]['infected'] == False:
# Mark all neighbors for infection
for w in nx.neighbors(G, v):
if G.nodes[w]['infected']:
to_infect.add(v)
break
# Infect marked nodes
for v in to_infect:
G.nodes[v]['infected'] = True
# +
# Infect two nodes
nx.set_node_attributes(
G_small_ring,
dict((i, False) for i in range(16)),
'infected')
for i in range(2):
G_small_ring.nodes[i]['infected'] = True
# Visualize first three time steps
plt.figure(figsize=(7.5, 2.5))
for i in range(3):
# Visualize
plt.subplot(1, 3, i + 1)
node_color = [
'#bfbf7f' if G_small_ring.nodes[v]['infected'] else '#9f9fff'
for v in G_small_ring.nodes]
nx.draw_networkx(
G_small_ring,
pos=nx.circular_layout(G_small_ring),
node_color=node_color)
# Propagate the contagion
propagate_simple(G_small_ring)
plt.title("Step {}".format(i))
plt.tight_layout()
# +
# Infect two nodes
nx.set_node_attributes(
G_small_random,
dict((i, False) for i in range(16)),
'infected')
for i in range(2):
G_small_random.nodes[i]['infected'] = True
# Visualize first three time steps
plt.figure(figsize=(7.5, 2.5))
for i in range(3):
# Visualize
plt.subplot(1, 3, i + 1)
node_color = [
'#bfbf7f' if G_small_random.nodes[v]['infected'] else '#9f9fff'
for v in G_small_random.nodes]
nx.draw_networkx(
G_small_random,
pos=nx.circular_layout(G_small_random),
node_color=node_color)
# Propagate the contagion
propagate_simple(G_small_random)
plt.title("Step {}".format(i))
plt.tight_layout()
# -
# ### Complex Contagion
def propagate_complex(G):
to_infect = set([])
# Find uninfected nodes
for v in G.nodes():
if G.nodes[v]['infected'] == False:
infected_neighbors = 0
# Count infected neighbors
for w in nx.neighbors(G, v):
if G.nodes[w]['infected']:
infected_neighbors += 1
# Remember nodes with 2+ infected neighbors
if infected_neighbors >= 2:
to_infect.add(v)
# Infect new nodes
for v in to_infect:
G.nodes[v]['infected'] = True
# +
# Infect two nodes
nx.set_node_attributes(
G_small_ring,
dict((i, False) for i in range(16)),
'infected')
for i in range(2):
G_small_ring.nodes[i]['infected'] = True
# Visualize first three time steps
plt.figure(figsize=(7.5, 2.5))
for i in range(3):
# Visualize
plt.subplot(1, 3, i + 1)
node_color = [
'#bfbf7f' if G_small_ring.nodes[v]['infected'] else '#9f9fff'
for v in G_small_ring.nodes]
nx.draw_networkx(
G_small_ring,
pos=nx.circular_layout(G_small_ring),
node_color=node_color)
# Propagate the contagion
propagate_complex(G_small_ring)
plt.title("Step {}".format(i))
plt.tight_layout()
# +
# Infect two nodes
nx.set_node_attributes(
G_small_random,
dict((i, False) for i in range(16)),
'infected')
for i in range(2):
G_small_random.nodes[i]['infected'] = True
# Visualize first three time steps
plt.figure(figsize=(7.5, 2.5))
for i in range(3):
# Visualize
plt.subplot(1, 3, i + 1)
node_color = [
'#bfbf7f' if G_small_random.nodes[v]['infected'] else '#9f9fff'
for v in G_small_random.nodes]
nx.draw_networkx(
G_small_random,
pos=nx.circular_layout(G_small_random),
node_color=node_color)
# Propagate the contagion
propagate_complex(G_small_random)
plt.title("Step {}".format(i))
plt.tight_layout()
# -
| Chapter08/Chapter_08.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:ptlesson] *
# language: python
# name: conda-env-ptlesson-py
# ---
# +
# Example 5-5: 와인 분류하기
## - Optimizer: SGD
## - Loss function: Cross entropy
##(5.2)
# PyTorch Library
import torch
from torch.autograd import Variable
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.utils.data import DataLoader, TensorDataset
# scikit-learn Library
from sklearn.datasets import load_wine
from sklearn.model_selection import train_test_split
# Pandas Library
import pandas as pd
# -
##(5.3) load wine data
wine = load_wine()
##(5.4) print data using DataFrame
pd.DataFrame(wine.data, columns=wine.feature_names)
##(5.5) print target
wine.target
##(5.6) assign 130 of data and target into variable
wine_data = wine.data[0:130]
wine_target = wine.target[0:130]
##(5.7) divide dataset: 80% of training set and 20% of test set
train_X, test_X, train_Y, test_Y = train_test_split(wine_data, wine_target, test_size=0.2)
print(len(train_X))
print(len(test_X))
##(5.8) convert numpy to torch tensor
train_X = torch.from_numpy(train_X).float()
train_Y = torch.from_numpy(train_Y).long()
test_X = torch.from_numpy(test_X).float()
test_Y = torch.from_numpy(test_Y).long()
print(train_X.shape)
print(train_Y.shape)
##(5.9) merge data and target into one tensor
### TensorDataset: Dataset wrapping tensors that have the same size of first dimension.
train = TensorDataset(train_X, train_Y)
print(train[0])
### DataLoader: set minibatches which are size of 16 and randomly shuffled
train_loader = DataLoader(train, batch_size=16, shuffle=True)
# +
##(5.10) Establish Neural Network
### class torch.nn.Module: Base class for all NN modules
### Net class inherits nn.Module(parent class)
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__() ## utilize __init__ method of parent class
self.fc1 = nn.Linear(13,96) ## torch.nn.Linear(in,out,bias=True)
self.fc2 = nn.Linear(96, 2) ## : applies linear transformation
def forward(self, x):
x = F.relu(self.fc1(x)) ## Hidden layer: ReLU
x = self.fc2(x)
return F.log_softmax(x,dim=0) ## Output layer: softmax
### create instance
model = Net()
# +
##(5.11)
### loss function: cross entropy
criterion = nn.CrossEntropyLoss()
### optimizer: stochastic gradient descent, learning rate 1%
optimizer = optim.SGD(model.parameters(), lr=0.01)
### start learning
for epoch in range(300):
total_loss = 0
### pop minibatche set out from train_loader
for train_x, train_y in train_loader:
### graph
train_x, train_y = Variable(train_x), Variable(train_y)
### initialize gradient as PyTorch accumulates gradients
optimizer.zero_grad()
### calc. forward propagation
output = model(train_x)
### calc. loss function
loss = criterion(output, train_y)
### calc. backward propagation
loss.backward()
### update weights
optimizer.step()
### calc. cumulative loss func
total_loss += loss.data.item()
if (epoch+1)%50 == 0:
print(epoch+1, total_loss)
# +
##(5.12)
### graph
### torch.autograd.Variable(data): wrapping tensor and logging
test_x, test_y = Variable(test_X), Variable(test_Y)
### make output 0 or 1
result = torch.max(model(test_x).data, 1)[1]
### accuracy of model
accuracy = sum(test_y.data.numpy() == result.numpy()) / len(test_y.data.numpy())
accuracy
# -
| ch5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py3.plutoalert
# language: python
# name: py3.plutoalert
# ---
import datetime
import pytz # pip install tzlocal pytz
from tzlocal import get_localzone # pip install tzlocal
# +
start_in="2022-03-02T02:33:00.000Z"
start = datetime.datetime.strptime(start_in[:19]+" +0000", "%Y-%m-%dT%H:%M:%S %z")
print(start.strftime("%H:%M %b %d %Z"))
start_localtime = start.astimezone(get_localzone())
start_print = start_localtime.strftime("%H:%M %b %d %Z")
print(start_print)
| interactive/timezones.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Python : How to Save and Load ML Models
# **object serialization**
# This process / procedure of saving a ML Model is also known as object serialization - representing an object with a stream of bytes, in order to store it on disk, send it over a network or save to a database.
#
# **deserialization**
# While the restoring/reloading of ML Model procedure is known as deserialization.
#
# In this notebook, we explore 2 ways to Save and Reload ML Models in Python and scikit-learn, we will also discuss about the pros and cons of each method.
# We will be covering 2 approaches of Saving and Reloading a ML Model -
#
# 1) Pickle Approach
# 2) Joblib Approach
# **ML Model Creation**
#
# For the purpose of Demo , we will create a basic Logistic Regression Model on IRIS Dataset.
# Dataset used : IRIS
# Model : Logistic Regression using Scikit Learn
# Import Required packages
from sklearn.linear_model import LogisticRegression
from sklearn.datasets import load_iris
from sklearn.model_selection import train_test_split
# Load the data
Iris_data = load_iris()
# Split data
x_train, x_test, y_train, y_test = train_test_split(Iris_data.data,
Iris_data.target,
test_size = 0.2,
random_state = 18)
# +
# Define the Model
log_reg = LogisticRegression(C = 0.1,
max_iter = 20,
fit_intercept = True,
solver = 'liblinear')
# Train the Model
log_reg.fit(x_train, y_train)
# -
# **Approach 1 : Pickle approach**
#
# Following lines of code, the log_reg which we created in the previous step is saved to file, and then loaded as a new object called pickle_log_reg.
# The loaded model is then used to calculate the accuracy score and predict outcomes on new unseen (test) data.
# Import pickle Package
import pickle
# +
# Save the Modle to file in the current working directory
pkl_filename = "save/pickle_log_reg.pkl"
with open(pkl_filename, 'wb') as file:
pickle.dump(log_reg, file)
# +
# Load the Model back from file
with open(pkl_filename, 'rb') as file:
pickle_log_reg = pickle.load(file)
pickle_log_reg
# +
# Use the Reloaded Model to
# Calculate the accuracy score and predict target values
# Calculate the Score
score = pickle_log_reg.score(x_test, y_test)
# Print the Score
print("Test score: {0:.2f} %".format(100 * score))
# Predict the Labels using the reloaded Model
y_pred = pickle_log_reg.predict(x_test)
y_pred
# -
# **Let's Reflect back on Pickle approach :**
#
# PROs of Pickle :
#
# 1) save and restore our learning models is quick - we can do it in two lines of code.
# 2) It is useful if you have optimized the model's parameters on the training data, so you don't need to repeat this step again.
#
#
# CONs of Pickle :
#
# 1) it doesn't save the test results or any data.
# **Approach 2 - Joblib** :
#
# The Joblib Module is available from Scikit Learn package and is intended to be a replacement for Pickle, for objects containing large data.
#
# This approach will save our ML Model in the pickle format only but we dont need to load additional libraries as the 'Pickling' facility is available within Scikit Learn package itself which we will use invariably for developing our ML models.
# Import Joblib Module from Scikit Learn
import joblib
# Save RL_Model to file in the current working directory
joblib_file = "save/joblib_log_reg.joblib"
joblib.dump(log_reg, joblib_file)
# +
# Load from file
joblib_log_reg = joblib.load(joblib_file)
joblib_log_reg
# +
# Use the Reloaded Joblib Model to
# Calculate the accuracy score and predict target values
# Calculate the Score
score2 = joblib_log_reg.score(x_test, y_test)
# Print the Score
print("Test score: {0:.2f} %".format(100 * score2))
# Predict the Labels using the reloaded Model
y_pred2 = joblib_log_reg.predict(x_test)
y_pred2
# -
# **Let's Reflect back on Joblib approach :**
#
# PROs of Joblib :
#
# 1) the Joblib library offers a bit simpler workflow compared to Pickle.
# 2) While Pickle requires a file object to be passed as an argument, Joblib works with both file objects and string filenames.
# 3) In case our model contains large arrays of data, each array will be stored in a separate file, but the save and restore procedure will remain the same.
# 4) Joblib also allows different compression methods, such as 'zlib', 'gzip', 'bz2', and different levels of compression.
| ai_ml/local/save-and-load-ml-models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2
import matplotlib.pyplot as plt
import numpy as np
def GetCircles(img,dp,minDist,para1,para2,minRadius,maxradius):
circles = cv2.HoughCircles(img,cv2.HOUGH_GRADIENT,dp,minDist,
param1=para1,param2=para2,minRadius=minradius,maxRadius=maxradius)
circles = np.uint16(np.around(circles))
particle_radius = circle[:,3]
for i in circles[0,:]:
# draw the outer circle
cv2.circle(img,(i[0],i[1]),i[2],(0,255,0),2)
# draw the center of the circle
cv2.circle(img,(i[0],i[1]),2,(0,0,255),3)
return plt.imshow(img)
import circle
img = cv2.imread("Opal_Tecopa_near_gem.jpg",0)
img = cv2.medianBlur(img,5)
circle.GetCircles(img,3,20,150,50,0,30)
plt.imshow(img)
| notebooks/Get Circle Function.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 + Jaspy
# language: python
# name: jaspy
# ---
# ## Demostrator 0
# #### Purpose:
# - In this example we download ERA5 data on-demand for training the IceNet2 model within the Azure/Pangeo environment
# - We also compare two methods (with/without Dask) to load the downloaded ERA5 data
#
# #### Comments:
# - We recommend to install the libraries into a virtual or conda environment
# - For instance, the notebook only works if you have in before
# - Installed the Copernicus Climate Data Store cdsapi package
# - Registered and setup your CDS API key as described on [their website here](https://cds.climate.copernicus.eu/api-how-to)
#
# #### Credits:
# - Some code snippets were adapted from the IceNet2 repository authored by <NAME> (BAS)
#
# #### Author:
# <NAME> (The Alan Turing Institute)
# ## Load libraries
import cdsapi
import xarray as xr
import numpy as np
import time
import pandas as pd
from pathlib import Path
import matplotlib.pyplot as plt
# ## Set distributed client
from dask.distributed import Client
client = Client()
client
# ## Request ERA5 data from CDI API
cds = cdsapi.Client()
# +
# Settings - as defined by TAndersson
## Variables
variables = {
'tas': {
'cdi_name': '2m_temperature',
},
'tos': {
'cdi_name': 'sea_surface_temperature',
},
'ta500': {
'plevel': '500',
'cdi_name': 'temperature',
},
}
## Target months, days and times
months = [
'01', '02', '03', '04', '05', '06',
'07', '08', '09', '10', '11', '12'
]
days = [
'01', '02', '03',
'04', '05', '06',
'07', '08', '09',
'10', '11', '12',
'13', '14', '15',
'16', '17', '18',
'19', '20', '21',
'22', '23', '24',
'25', '26', '27',
'28', '29', '30',
'31',
]
times = [
'00:00', '01:00', '02:00',
'03:00', '04:00', '05:00',
'06:00', '07:00', '08:00',
'09:00', '10:00', '11:00',
'12:00', '13:00', '14:00',
'15:00', '16:00', '17:00',
'18:00', '19:00', '20:00',
'21:00', '22:00', '23:00',
]
# +
# Variables - as defined by TAndersson
var = 'tas'
hemisphere = 'nh'
year = 2020
# Output folder and files
notebook_folder = '/mnt/batch/tasks/shared/LS_root/mounts/clusters/ci-alejandroc-943cade0/code'
var_folder = os.path.join(notebook_folder,'data', hemisphere, var)
if not os.path.exists(var_folder):
os.makedirs(var_folder)
download_path = os.path.join(var_folder, '{}_latlon_hourly_{}_{}.nc'.format(var, year, year))
daily_fpath = os.path.join(var_folder, '{}_latlon_{}_{}.nc'.format(var, year, year))
# +
# Settings - as defined by TAndersson
## Hemisphere
if hemisphere == 'nh':
area = [90, -180, 0, 180]
elif hemisphere == 'sh':
area = [-90, -180, 0, 180]
elif hemisphere == 'nh_sh':
area = [-90, -180, 90, 180]
## Target var dictionary
var_dict = variables[var]
## Retrieve dictionary
retrieve_dict = {
'product_type': 'reanalysis',
'variable': var_dict['cdi_name'],
'year': year,
'month': months,
'day': days,
'time': times,
'format': 'netcdf',
'area': area
}
if 'plevel' not in var_dict.keys():
dataset_str = 'reanalysis-era5-single-levels'
elif 'plevel' in var_dict.keys():
dataset_str = 'reanalysis-era5-pressure-levels'
retrieve_dict['pressure_level'] = var_dict['plevel']
# -
# %%time
# check if file exist, otherwise retrieve the target variable from CDI
if not Path(download_path).is_file():
cds.retrieve(dataset_str, retrieve_dict, download_path)
# ## Load and resample from hourly to daily
# #### Method 1 - Using open_mfdataset function and DASK to chunk the data
# %%time
# Import data as xarray dataset from the directory
dask = True
if dask:
# Import with dask
da = xr.open_mfdataset(download_path, parallel=True,
combine='by_coords', chunks={'time': 50}
, engine='netcdf4')
print(f'The chunk size for time dimension is {da.chunks["time"][0]}\n')
print(f'Dataset, thus, have {len(da.time)/da.chunks["time"][0]} chunks\n')
else:
# Import without dask for debugging
da = xr.open_mfdataset(download_path, parallel=False, combine='by_coords', engine='netcdf4')
# +
# %%time
print('\n\nComputing daily averages... ', end='', flush=True)
da_daily_mfdataset = da.resample(time='1D').reduce(np.mean)
if var == 'zg500' or var == 'zg250':
da_daily_mfdataset = da_daily_mfdataset / 9.80665
if var == 'tos':
# Replace every value outside of SST < 1000 with zeros (the ERA5 masked values)
da_daily_mfdataset = da_daily_mfdataset.where(da_daily_mfdataset < 1000., 0)
# -
# #### Method 2 - Using open_dataarray
# %%time
# Import using open_dataarray
da = xr.open_dataarray(download_path)
# +
# %%time
print('\n\nComputing daily averages... ', end='', flush=True)
da_daily = da.resample(time='1D').reduce(np.mean)
if var == 'zg500' or var == 'zg250':
da_daily = da_daily / 9.80665
if var == 'tos':
# Replace every value outside of SST < 1000 with zeros (the ERA5 masked values)
da_daily = da_daily.where(da_daily < 1000., 0)
# -
try:
xr.testing.assert_allclose(da_daily,da_daily_mfdataset,atol=1e-6)
except AssertionError:
print("There are differences between the resample datasets according to the assert test")
if not Path(daily_fpath).is_file():
print(np.unique(da_daily.data)[[0, -1]]) # TEMP
print('saving new daily year file... ', end='', flush=True)
da_daily.to_netcdf(daily_fpath)
# ## Visualization and comparison amongts the Xarray methods
# %%time
# plot the last timestep using the daily dataset generated using open_dataarray
da_daily.isel(time=-1).plot()
# %%time
# plot the last timestep using the daily dataset generated using open_mfdataset
da_daily_mfdataset.isel(time=-1)['t2m'].plot()
# %%time
# plot a timeseries at one location using the daily dataset generated using open_dataarray
da_daily.sel(longitude=114.3055, latitude=30.5928, method='nearest').plot()
# %%time
# plot a timeseries at one location using the daily dataset generated using open_dataarray
# notes1: slower than the daily dataset generated using open_mfdataset
# notes2: it doesn't work when using open_mfdataset and dask FALSE (memory issues)
da_daily_mfdataset.sel(longitude=114.3055, latitude=30.5928, method='nearest')['t2m'].plot()
# ## Close the Client
#
# Before moving on to the next exercise, make sure to close your client or stop this kernel.
client.close()
| copernicus/demo0_Pangeo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lab 4: Functional Programming
# ## Overview
# Explore functional programming's place in the Python landscape, and gain practice with powerful tools like `map`, `filter`, iterators, generators, and decorators.
#
# *Surprisingly, a few people have asked for longer labs - we think we've delivered! We've added lots of challenge problems to this lab, many of which are domain-specific, but we don't expect you to complete them all. If you're short on time, or don't know exactly what a challenge problem is asking, skip it! Challenge problems are intended to be challenging, and are reserved for when you've finished the rest of the lab.*
# ## Functional Tools
# ### Lambdas
#
# Recall that lambda functions are anonymous, unnamed function objects created on the fly, usually to accomplish a small transformation. For example,
#
# ```Python
# (lambda val: val ** 2)(5) # => 25
# (lambda x, y: x * y)(3, 8) # => 24
# (lambda s: s.strip().lower()[:2])(' PyTHon') # => 'py'
# ```
#
# On their own, `lambda`s aren't particularly useful, as demonstrated above. Usually, `lambda`s are used to avoid creating a formal function definiton for small throwaway functions, not only because they involves less typing (no `def` or `return` statement needed) but also, and perhaps more importantly, because these small functions won't pollute the enclosing namespace.
#
# Lambdas are also frequently used as arguments to or return values from higher-order functions, such as `map` and `filter`.
# ### Map
#
# Recall from class that `map(func, iterable)` applies a function over elements of an iterable.
#
# For each of the following rows, write a single statement using `map` that converts the left column into the right column:
#
# | From | To|
# | --- | --- |
# | `['12', '-2', '0']` | `[12, -2, 0]` |
# | `['hello', 'world']` | `[5, 5]` |
# | `['hello', 'world']`|`['olleh', 'dlrow']` |
# | `range(2, 6)`|`[(2, 4, 8), (3, 9, 27), (4, 16, 64), (5, 25, 125)]` |
# | `zip(range(2, 5), range(3, 9, 2))`|`[6, 15, 28]` |
#
# *Hint: you may need to wrap the output in a `list()` constructor to see it printed to console - that is, `list(map(..., ...))`*
# #### Using Multiple Iterables
# The `map` function can accept a variable number of iterables as arguments. Thus, `map(func, iterA, iterB, iterC)` is equivalent to `map(func, zip(iterA, iterB, iterC))`. This can be used as follows:
#
# ```Python
# map(int, ('10110', '0xCAFE', '42'), (2, 16, 10)) # generates 22, 51966, 42
# ```
# *This works because* `int` *takes an optional second argument specifying the conversion base*
#
# ### Filter
#
# Recall from class that `filter(pred, iterable)` keeps only those elements from an iterable that satisfy a predicate function.
#
# Write statements using `filter` that convert the following sequences from the left column to the right column:
#
# From | To
# --- | ---
# `['12', '-2', '0']` | `['12', '0']`
# `['hello', 'world']` | `['world']`
# `['Stanford', 'Cal', 'UCLA']`|`['Stanford']`
# `range(20)`|`[0, 3, 5, 6, 9, 10, 12, 15, 18]`
# ### More Useful Tools (optional)
# #### Module: `functools`
# The functools module is "for higher order functions; functions that act on or return other functions."
#
# There is a utility in the `functools` module called `reduce`, which in Python 2.x was a builtin language feature but has since been relegated to this module. The `reduce` function is explained best by the [official documentation](https://docs.python.org/3.4/library/functools.html#functools.reduce):
#
# > `functools.reduce(function, iterable[, initializer])`
# >> Apply `function` of two arguments cumulatively to the items of `iterable`, from left to right, so as to reduce the iterable to a single value. For example, `functools.reduce(lambda x, y: x + y, [1, 2, 3, 4, 5])` calculates `((((1 + 2) + 3) + 4) + 5)`. The left argument, `x`, is the accumulated value and the right argument, `y`, is the update value from the sequence. If the optional `initializer` is present, it is placed before the items of the sequence in the calculation, and serves as a default when the iterable is empty. If `initializer` is not given and `iterable` contains only one item, the first item is returned.
#
# Use the `reduce` function to find the least common multiple (LCM) of an arbitrary amount of positive integer arguments. This can be accomplished in one line of Python.
#
# ```Python
# import operator
# from functools import reduce
# from fractions import gcd
#
# def lcm(*nums):
# pass
# # Your implementation here: Use reduce. This function can be implemented in only one line!
#
# lcm()
# ```
#
# Hint: Recall that, mathematically, the LCM of two numbers `x` and `y` can be expressed as `(x*y) // gcd(x, y)`, and that the LCM of a list of numbers `[x, y, z, ...]` is the same as the `LCM(...(LCM(LCM(x, y), z), ...)`.
# #### Module: `operator`
#
# Frequently, you might find yourself writing anonymous functions similar to `lambda x, y: x + y`. This feels a little redundant, since Python already knows how to add two values together. Unfortunately, we can't just refer to `+` as a function - it's a builtin syntax element. To solve this problem, The `operator` module exports callable functions for each builtin operation. These operators can simplify some common uses of lambdas, and should be used wherever possible, since in almost all cases they are faster than constructing and repeatedly invoking a lambda function.
#
# ```Python
# import operator
# operator.add(1, 2) # => 3
# operator.mul(3, 10) # => 30
# operator.pow(2, 3) # => 8
# operator.itemgetter(1)([1, 2, 3]) # => 2
# ```
#
# Take a moment to skim over the [official documentation for the `operator` module](https://docs.python.org/3.4/library/operator.html).
#
# Use `reduce` in conjunction with a function from the `operator` module to compute factorials in one line of Python:
#
# ```Python
# import operator
# from functools import reduce
#
# def fact(n):
# # Your implementation here: Use reduce, an operator, and only one line!
#
# fact(3) # => 6
# fact(7) # => 5040
# ```
# #### Custom comparison for `sort`, `max`, and `min`
#
# When ordering sequences (or finding the largest or smallest element), Python defaults a default ordering for sequence elements. For instance, a collection of strings will be sorted alphabetically (by ASCII value), and a collection of tuples will sort lexicographically. Sometimes, however, we need to sort based on a custom key value. In Python, we can supply an optional `key` argument to `sorted(seq)`, `max(seq)`, `min(seq)`, or `seq.sort()` to determine the values used for ordering elements in a sequence. In Python, both `sorted(seq)` and `seq.sort()` are stable.
#
# For example:
#
# ```Python
# words = ['pear', 'cabbage', 'apple', 'bananas']
# min(words) # => 'apple'
# words.sort(key=lambda s: s[-1]) # Alternatively, key=operator.itemgetter(-1)
# words # => ['cabbage', 'apple', 'pear', 'bananas'] ... Why 'cabbage' > 'apple'?
# max(words, key=len) # 'cabbage' ... Why not 'bananas'?
# min(words, key=lambda s: s[1::2]) # What will this value be?
# ```
#
# Write a function to return the two words with the highest alphanumeric score of uppercase letters:
#
# ```Python
# def alpha_score(upper_letters):
# """Computers the alphanumeric sum of letters in a string.
# Prerequisite: upper_letters is composed entirely of capital letters.
# """
# return sum(map(lambda l: 1 + ord(l) - ord('A'), upper_letters))
#
# alpha_score('ABC') # => 6 = 1 ('A') + 2 ('B') + 3 ('C')
#
# def two_best(words):
# pass # Your implementation here
#
# two_best(['hEllO', 'wOrLD', 'i', 'aM', 'PyThOn'])
# ```
#
# You may want to use `filter` too.
#
# ## Purely Functional Programming (optional)
#
# As an academic thought exercise, let's investigate how we would use Python in a purely functional programming paradigm. Ultimately, we will try to remove statements and replace them with expressions.
# ### Replacing Control Flow
#
# The first thing that needs to go are control flow statements - `if/elif/else`. Luckily, Python, like many other languages, short circuits boolean expressions. This means that we can rewrite
#
# ```Python
# if <cond1>: func1()
# elif <cond2>: func2()
# else: func3()
# ```
#
# as the equivalent expression
#
# ```Python
# (<cond1> and func1()) or (<cond2> and func2()) or (func3())
# ```
#
# Note: The above will work if and only if all of the functions return truthy values.
#
# Rewrite the following code block without using `if/elif/else`:
#
# ```Python
# if score == 1:
# return "Winner"
# elif score == -1:
# return "Loser"
# else:
# return "Tied"
# ```
# ### Replacing Returns
#
# However, we would still need return values to do anything useful. Since lambdas implicitly return their expression, we will use lambdas to eliminate return statements. We can bind these temporary conditional conjunctive normal form expressions to a lambda function.
#
# ```Python
# echo = lambda arg: arg # In practice, you should never bind lambdas to local names
# cond_fn = lambda x: (x==1 and echo("one")) \
# or (x==2 and echo("two")) \
# or (echo("other"))
# ```
# ### Replacing Loops
#
# Getting rid of loops is easy! We can `map` over a sequence instead of looping over the sequence. For example:
#
# ```Python
# for e in lst:
# func(e)
# ```
#
# becomes
#
# ```Python
# map(func, lst)
# ```
# ### Replacing Action Sequence
# Most programs take the form a sequence of steps, written out line by line. By using a `just_do_it` function and `map`, we can replicate a sequence of function calls.
#
# ```Python
# just_do_it = lambda f: f()
#
# # Suppose f1, f2, f3 are actions
# map(just_do_it, [f1, f2, f3])
# ```
#
# Our main program execution can then be a single call to such a map expression.
#
# #### Note
# In fact, Python has `eval` and `exec` functions builtin. Don't use them! They are dangerous.
# ### Closing
# Python supports functional programming paradigms, but as you can see, in some cases FP introduces unnecessary complexity.
#
# If you really enjoyed this section, read [Part 1](http://www.ibm.com/developerworks/linux/library/l-prog/index.html), [Part 2](http://www.ibm.com/developerworks/linux/library/l-prog2/index.html), and [Part 3](http://www.ibm.com/developerworks/linux/library/l-prog3/index.html) of IBM's articles on FP in Python.
#
# ## Iterators
#
# Recall from class than an iterator is an object that represents a stream of data returned one value at a time.
# ### Iterator Consumption
# Suppose the following two lines of code have been run:
#
# ```Python
# it = iter(range(100))
# 67 in it # => True
# ```
#
# What is the output of each of the following lines of code?
#
# ```Python
# next(it) # => ??
# 37 in it # => ??
# next(it) # => ??
# ```
#
# With a partner, discuss why we see these results.
# ### Module: `itertools`
#
# Python ships with a spectacular module for manipulating iterators called `itertools`. Take a moment to read through the [documentation page for itertools](https://docs.python.org/3.4/library/itertools.html).
#
# Predict the output of the following pieces of code:
#
# ```Python
# import itertools
# import operator
#
# for el in itertools.permutations('XKCD', 2):
# print(el, end=', ')
#
# for el in itertools.cycle('LO'):
# print(el, end='') # Don't run this one. Why not?
#
# itertools.starmap(operator.mul, itertools.zip_longest([3,5,7],[2,3], fillvalue=1))
# ```
# ### (Challenge) Linear Algebra
#
# These challenge problems test your ability to write compact Python functions using the tools of functional programming and some good old-fashioned cleverness. As always, challenge problems are optional. These challenge problems focus heavily on linear algebra.
# #### Dot Product
# Write a one-liner in Python that takes the dot product of two lists `u` and `v`. You can assume that the lists are the same size, and are standard Python lists (not anything special, like numpy arrays). You should use
#
# ```Python
# def dot_product(u, v):
# pass
# ```
#
# For example, `dot_product([1, 3, 5], [2, 4, 6])` should return `44` (since `1 * 2 + 3 * 4 + 5 * 6 = 44`).
# #### Matrix Transposition
# Write a one-liner in Python to transpose a matrix. Assume that the input matrix is a tuple-of-tuples that represents a valid matrix, not necessarily square. Again, do not use numpy or any other libraries - just raw data structure manipulation.
#
# ```Python
# def transpose(m):
# pass
# ```
#
# Not only can you do this in one line - you can do it in 14 characters!
#
# For example,
#
# ```Python
# matrix = (
# (1, 2, 3, 4),
# (5, 6, 7, 8),
# (9,10,11,12)
# )
# transpose(matrix)
# # returns
# # (
# # (1, 5, 9),
# # (2, 6, 10),
# # (3, 7, 11),
# # (4, 8, 12)
# # )
# ```
#
# #### Matrix Multiplication
# Write another one-liner in Python to take the product of two matrices `m1` and `m2`. You can use the `dot_product` and `transpose` functions you already wrote.
#
# ```Python
# def matmul(m1, m2):
# pass
# ```
# #### Lazy Generation
# Rewrite your `transpose` and `matmul` functions above so that they are lazily evaluated.
# ## Generator Expressions
#
# Recall that generator expressions are a way to lazily compute values on the fly, without buffering the entire contents of the list in place.
#
# For each of the following scenarios, discuss whether it would be more appropriate to use a generator expression or a list comprehension:
#
# 1. Searching for a given entity in the entries of a 1TB database.
# 2. Calculate cheap airfare using lots of journey-to-destination flight information.
# 3. Finding the first palindromic Fibonacci number greater than 1,000,000.
# 4. Determine all multi-word anagrams of user-supplied 1000-character-or-more strings (very expensive to do).
# 5. Generate a list of names of Stanford students whose SUNet ID numbers are less than 5000000.
# 6. Return a list of all startups within 50 miles of Stanford.
#
# ## Generators
# ### Triangle Generator
#
# Write a infinite generator that successively yields the triangle numbers `0, 1, 3, 6, 10, ...`
#
# ```Python
# def generate_triangles():
# pass # Your implementation here
# ```
#
# Use your generator to write a function `triangles_under(n)` that prints out all triangle numbers strictly less than the parameter `n`.
#
# ```Python
# def triangles_under(n):
# pass
# ```
#
# ## Functions in Data Structures
#
# In class, we quickly showed a highly unusual way to generate primes. Take some time to read through it again, and talk with a partner about how and why this successfully generates prime numbers.
#
# ```Python
# def primes_under(n):
# tests = []
# for i in range(2, n):
# if not any(map(lambda test: test(i), tests)):
# tests.append(make_divisibility_test(i))
# yield i
# ```
#
# How would you modify the code above to yield all composite numbers, rather than all prime numbers? Test your solution. What is the 1000th composite number?
# ## Nested Functions and Closures
#
# In class, we saw that functions can be defined within the scope of another function (recall from Week 3 that functions introduce new scopes via a new local symbol table). An inner function is only in scope inside of the outer function, so this type of function definition is usually only used when the inner function is being returned to the outside world.
#
# ```Python
# def outer():
# def inner(a):
# return a
# return inner
#
# f = outer()
# print(f) # <function outer.<locals>.inner at 0x1044b61e0>
# f(10) # => 10
#
# f2 = outer()
# print(f2) # <function outer.<locals>.inner at 0x1044b6268> (Different from above!)
# f2(11) # => 11
# ```
#
# Why are the memory addresses different for `f` and `f2`? Discuss with a partner.
# ### Closure
# As we saw above, the definition of the inner function occurs during the execution of the outer function. This implies that a nested function has access to the environment in which it was defined. Therefore, it is possible to return an inner function that remembers the state of the outer function, even after the outer function has completed execution. This model is referred to as a closure.
#
# ```Python
# def make_adder(n):
# def add_n(m): # Captures the outer variable `n` in a closure
# return m + n
# return add_n
#
# add1 = make_adder(1)
# print(add1) # <function make_adder.<locals>.add_n at 0x103edf8c8>
# add1(4) # => 4
# add1(5) # => 6
# add2 = make_adder(2)
# print(add2) # <function make_adder.<locals>.add_n at 0x103ecbf28>
# add2(4) # => 6
# add2(5) # => 7
# ```
#
# The information in a closure is available in the function's `__closure__` attribute. For example:
#
# ```Python
# closure = add1.__closure__
# cell0 = closure[0]
# cell0.cell_contents # => 1 (this is the n = 1 passed into make_adder)
# ```
#
# As another example, consider the function:
#
# ```Python
# def foo(a, b, c=-1, *d, e=-2, f=-3, **g):
# def wraps():
# print(a, c, e, g)
# ```
#
# The `print` call induces a closure of `wraps` over `a`, `c`, `e`, `g` from the enclosing scope of `foo`. Or, you can imagine that wraps "knows" that it will need `a`, `c`, `e`, and `g` from the enclosing scope, so at the time `wraps` is defined, Python takes a "screenshot" of these variables from the enclosing scope and stores references to the underlying objects in the `__closure__` attribute of the `wraps` function.
#
# ```Python
# w = foo(1, 2, 3, 4, 5, e=6, f=7, y=2, z=3)
# list(map(lambda cell: cell.cell_contents, w.__closure__))
# # = > [1, 3, 6, {'y': 2, 'z': 3}]
# ```
#
# What happens in the following situation? Why?
# ```Python
# def outer(l):
# def inner(n):
# return l * n
# return inner
#
# l = [1, 2, 3]
# f = outer(l)
# print(f(3)) # => ??
#
# l.append(4)
# print(f(3)) # => ??
# ```
#
# ## Building Decorators
#
# Recall that a decorator is a special type of function that accepts a function as an argument and (usually) returns a modified version of that function. In class, we saw the `debug` decorator - review the slides if you still feel uncomfortable with the idea of a decorator.
#
# Furthermore, recall that the `@decorator` syntax is syntactic sugar.
#
# ```Python
# @decorator
# def fn():
# pass
# ```
#
# is equivalent to
#
# ```Python
# def fn():
# pass
# fn = decorator(fn)
# ```
#
# ### Review
#
# In lecture, we implemented the `debug` decorator.
#
# ```Python
# def debug(function):
# def wrapper(*args, **kwargs):
# print("Arguments:", args, kwargs)
# return function(*args, **kwargs)
# return wrapper
# ```
#
# Take a moment, with a partner, and make sure you understand what is happening in the above lines. Why are the arguments to wrapper on the second line `*args` and `**kwargs` instead of something else? What would happen if we didn't `return wrapper` at the end of the function body?
# ### Automatic Caching
# Write a decorator `cache` that will automatically cache any calls to the decorated function. You can assume that all arguments passed to the decorated function will always be hashable types.
#
# ```Python
# def cache(function):
# pass # Your implementation here
# ```
#
# For example:
#
# ```Python
# @cache
# def fib(n):
# return fib(n-1) + fib(n-2) if n > 2 else 1
#
# fib(10) # 55 (takes a moment to execute)
# fib(10) # 55 (returns immediately)
# fib(100) # doesn't take forever
# fib(400) # doesn't raise RuntimeError
# ```
#
# Hint: You can set arbitrary attributes on a function (e.g. `fn._cache`). When you do so, the attribute-value pair also gets inserted into `fn.__dict__`. Take a look for yourself. Are the extra attributes and `.__dict__` always in sync?
# #### Challenge: Cache Options
#
# Add `max_size` and `eviction_policy` keyword arguments, with reasonable defaults (perhaps `max_size=None` as a sentinel), to your `cache` decorator. `eviction_policy` should be `'LRU'`, `'MRU'`, or `'random'`.
#
# #### Note
# This is actually implemented as part of the language in `functools.lru_cache`
# ### `print_args`
# The `debug` decorator we wrote in class isn't very good. It doesn't tell us which function is being called, and it just gives us a tuple of positional arguments and a dictionary of keyword arguments - it doesn't even know what the names of the positional arguments are! If the default arguments aren't overridden, it won't show us their value either.
#
# Use function attributes to improve our `debug` decorator into a `print_args` decorator that is as good as you can make it.
#
# ```Python
# def print_args(function):
# def wrapper(*args, **kwargs):
# # (1) You could do something here
# retval = function(*args, **kwargs)
# # (2) You could also do something here
# return retval
# return wrapper
# ```
#
# *Hint: Consider using the attributes `fn.__name__` and `fn.__code__`. You'll have to investigate these attributes, but I will say that the `fn.__code__` code object contains a number of useful attributes - for instance, `fn.__code__.co_varnames`. Check it out! More information is available in the latter half of Lab 3.*
#
# #### Note
# There are a lot of subtleties to this function, since functions can be called in a number of different ways. How does your `print_args` handle keyword arguments or even keyword-only arguments? Variadic positional arguments? Variadic keyword arguments? For more customization, look at `fn.__defaults__`, `fn.__kwdefaults__`, as well as other attributes of `fn.__code__`.
# ### Dynamic Type Checker
#
# Functions in Python can be optionally annotated by semantically-useless but structurally-valuable type annotations. For example:
#
# ```Python
# def foo(a: int, b: str) -> bool:
# return b[a] == 'X'
#
# foo.__annotations__ # => {'a': int, 'b': str, 'return': bool}
# ```
#
# Write a dynamic type checker, implemented as a decorator, that enforces the parameter types and return type of Python objects, if supplied.
#
# ```Python
# def enforce_types(function):
# pass # Your implementation here
# ```
#
# For example:
#
# ```Python
# @enforce_types
# def foo(a: int, b: str) -> bool:
# if a == -1:
# return 'Gotcha!'
# return b[a] == 'X'
#
# foo(3, 'abcXde') # => True
# foo(2, 'python') # => False
# foo(1, 4) # prints "Invalid argument type for b: expected str, received int
# foo(-1, '') # prints "Invalid return type: expected bool, received str
# ```
#
# There are lots of nuances to this function. What happens if some annotations are missing? How are keyword arguments and variadic arguments handled? What happens if the expected type of a parameter is not a primitive type? Can you annotate a function to describe that a parameter should be a list of strings? A tuple of (str, bool) pairs? A dictionary mapping strings to lists of ints?
#
# If you think you're done, show your decorator to a member of the course staff.
# #### Bonus: Optional Severity Argument
# *Warning! This extension is very hard*
#
# Extend the `enforce_types` decorator to accept a keyword argument `severity` which modifies the extent of the enforcement. If `severity == 0`, disable type checking. If `severity == 1` (which is the default), just print a message if there are any type violations. If `severity == 2`, raise a TypeError if there are any type violations.
#
# For example:
#
# ```Python
# @enforce_types_challenge(severity=2)
# def bar(a: list, b: str) -> int:
# return 0
#
# @enforce_types_challenge() # Why are there parentheses here?
# def baz(a: bool, b: str) -> str:
# return ''
# ```
#
# ## Credit
# Credit goes to a lot of websites, whose names I've unfortunately forgotten along the way. Credit to everyone!
# > With <3 by @sredmond
| notebooks/lab4-fp-notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R 4.0
# language: R
# name: ir40
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Dependencies" data-toc-modified-id="Dependencies-1">Dependencies</a></span></li><li><span><a href="#Paths" data-toc-modified-id="Paths-2">Paths</a></span></li><li><span><a href="#Main" data-toc-modified-id="Main-3">Main</a></span><ul class="toc-item"><li><span><a href="#Remove-cell-lines" data-toc-modified-id="Remove-cell-lines-3.1">Remove cell lines</a></span></li><li><span><a href="#Duplicated-samples-(includes-cell-lines)" data-toc-modified-id="Duplicated-samples-(includes-cell-lines)-3.2">Duplicated samples (includes cell lines)</a></span></li><li><span><a href="#Sample-types" data-toc-modified-id="Sample-types-3.3">Sample types</a></span></li><li><span><a href="#Sufficient-sample-size" data-toc-modified-id="Sufficient-sample-size-3.4">Sufficient sample size</a></span></li><li><span><a href="#Clustering-cleanup" data-toc-modified-id="Clustering-cleanup-3.5">Clustering cleanup</a></span></li></ul></li><li><span><a href="#Sankey-plot" data-toc-modified-id="Sankey-plot-4">Sankey plot</a></span></li></ul></div>
# -
# # Dependencies
library(ggforce)
source("~/OneDrive - UHN/R_src/ggplot2_theme.R")
# # Paths
manifestpath <- "/Users/anabbi/OneDrive - UHN/Documents/IPD2/Manifests/"
datapath <- "/Users/anabbi/OneDrive - UHN/Documents/IPD2/Data/"
plotpath <- "/Users/anabbi/OneDrive - UHN/Documents/IPD2/Plots/"
# # Main
# Start with RSEM manifest file
CBTN_RNA_manifest_genes <- read.csv(paste0(manifestpath, "CBTTC_RSEM-manifest.csv"),header = TRUE, stringsAsFactors = FALSE)
dim(CBTN_RNA_manifest_genes)
CBTN_samples <- cbind.data.frame(CBTN_RNA_manifest_genes$sample_id,
CBTN_RNA_manifest_genes$aliquot_id, stringsAsFactors = F)
colnames(CBTN_samples) <- c("CBTN_sample_id", "CBTN_aliquot_id")
CBTN_samples$CBTN_sample_aliquot <- paste0(CBTN_samples$CBTN_sample_id,"_", CBTN_samples$CBTN_aliquot_id)
CBTN_samples$CBTN <- "CBTN\nsamples\n(n = 996)"
head(CBTN_samples)
# ## Remove cell lines
CBTN_celllines <- read.csv(paste0(manifestpath,"cell_line_RNAalqids_CBTTC.csv"), header = TRUE, stringsAsFactors = F)
head(CBTN_celllines)
CBTN_celllines$sample_aliquot_id <- paste0(CBTN_celllines$clinical_event, "_", CBTN_celllines$cell_lines_rnaalq_id)
cellines <- CBTN_samples$CBTN_sample_aliquot[ CBTN_samples$CBTN_sample_aliquot %in% CBTN_celllines$sample_aliquot_id]
not_celllines <- CBTN_samples$CBTN_sample_aliquot[!CBTN_samples$CBTN_sample_aliquot %in% CBTN_celllines$sample_aliquot_id]
# +
CBTN_samples$Cell_lines <- NA
CBTN_samples$Cell_lines[ CBTN_samples$CBTN_sample_aliquot %in% cellines] <-
paste0("Cell lines\n(n = ", length(cellines), ")")
CBTN_samples$Cell_lines[CBTN_samples$CBTN_sample_aliquot %in% not_celllines] <-
paste0("Tumours\n(n = ", length(not_celllines), ")")
# -
table(CBTN_samples$Cell_lines)
head(CBTN_samples)
# ## Duplicated samples (includes cell lines)
CBTN_RNA_manifest_genes$sample_aliquot_id <- paste0(CBTN_RNA_manifest_genes$sample_id, "_", CBTN_RNA_manifest_genes$aliquot_id)
CBTN_RNA_manifest_genes_dedup <- CBTN_RNA_manifest_genes[!duplicated(CBTN_RNA_manifest_genes$sample_id),]
dim(CBTN_RNA_manifest_genes_dedup)
dedup_ids <- CBTN_RNA_manifest_genes_dedup$sample_aliquot_id
dup_ids <- CBTN_RNA_manifest_genes$sample_aliquot_id[duplicated(CBTN_RNA_manifest_genes$sample_id)]
dup_ids
# +
CBTN_samples$duplicated_samples <- NA
CBTN_samples$duplicated_samples[CBTN_samples$CBTN_sample_aliquot %in% dup_ids] <-
paste0("Duplicated\nsamples\n(n = ",length(dup_ids), ")")
CBTN_samples$duplicated_samples[CBTN_samples$CBTN_sample_aliquot %in% dedup_ids] <-
paste0("Unique\nsamples\n(n = ",length(dedup_ids), ")")
# -
table(CBTN_samples$duplicated_samples, useNA = "always")
# ## Sample types
load(file = paste0(manifestpath,"Master_manifest.RData"))
cbtn_metadata <- IPD_metadata[ IPD_metadata$group == "CBTTC",]
cbtn_metadata$sample_aliquot_id <- paste0(cbtn_metadata$sample_id, "_", cbtn_metadata$aliquot_id)
table(cbtn_metadata$sample_type, useNA = "always")
length(cbtn_metadata$sample_type)
primaries <- cbtn_metadata$sample_id[cbtn_metadata$sample_type == "Initial CNS Tumor"]
progressives <- cbtn_metadata$sample_id[cbtn_metadata$sample_type== "Progressive"]
recurrences <- cbtn_metadata$sample_id[cbtn_metadata$sample_type == "Recurrence"]
secondmalig <- cbtn_metadata$sample_id[cbtn_metadata$sample_type == "Second Malignancy"]
# +
CBTN_samples$tumour_type <- NA
CBTN_samples$tumour_type[CBTN_samples$CBTN_sample_id %in% primaries] <-
paste0("Primary\n(n = ",
length(CBTN_samples$CBTN_sample_id[CBTN_samples$CBTN_sample_id %in% primaries]), ")")
CBTN_samples$tumour_type[ CBTN_samples$CBTN_sample_id %in% progressives] <-
paste0("Progressive\n(n = ",
length(CBTN_samples$CBTN_sample_id[CBTN_samples$CBTN_sample_id %in% progressives]), ")")
CBTN_samples$tumour_type[ CBTN_samples$CBTN_sample_id %in% recurrences ] <-
paste0("Recurrence\n(n = ",
length(CBTN_samples$CBTN_sample_id[CBTN_samples$CBTN_sample_id %in% recurrences]), ")")
CBTN_samples$tumour_type[ CBTN_samples$CBTN_sample_id %in% secondmalig ] <-
paste0("Second\nmalignancy\n(n = ",
length(CBTN_samples$CBTN_sample_id[CBTN_samples$CBTN_sample_id %in% secondmalig]), ")")
# -
table(CBTN_samples$tumour_type, useNA = "always")
CBTN_samples$CBTN_sample_id[is.na(CBTN_samples$tumour_type)]
#all are cellline entries from initial CNS tumours
# add those 4 unannotated samples using original clinical csv file
CBTTC_clinical <- read.csv(paste0(manifestpath,"CBTTC_clinical.csv"),header = TRUE, stringsAsFactors = FALSE, na.strings = c("", NA))
CBTTC_clinical[ CBTTC_clinical$CBTTC.Event.ID %in% CBTN_samples$CBTN_sample_id[is.na(CBTN_samples$tumour_type)],]
primaries <- c(primaries, CBTN_samples$CBTN_sample_id[is.na(CBTN_samples$tumour_type)])
CBTN_samples$tumour_type[CBTN_samples$CBTN_sample_id %in% primaries] <-
paste0("Primary\n(n = ",
length(CBTN_samples$CBTN_sample_id[CBTN_samples$CBTN_sample_id %in% primaries]), ")")
table(CBTN_samples$tumour_type, useNA = "always")
# ## Sufficient sample size
load(file = paste0(datapath,"ESTIMATE/estimate_manifest_primary.RData"))
load(file = paste0(datapath,"ESTIMATE/estimate_manifest_primary_clean.RData"))
cbtn_primary <- estimate_manifest_primary[ estimate_manifest_primary$group == "CBTTC",]
cbtn_primary_clean <- estimate_manifest_primary_clean[ estimate_manifest_primary_clean$group == "CBTN",]
dim(cbtn_primary)
dim(cbtn_primary_clean)
cbtn_primary$sample_aliquot_id <- paste0(cbtn_primary$sample_id, "_", cbtn_primary$aliquot_id)
cbtn_primary_clean$sample_aliquot_id <- paste0(cbtn_primary_clean$sample_id, "_", cbtn_primary_clean$aliquot_id)
goodsamplesize <- cbtn_primary_clean$sample_aliquot_id
badsamplesize <- cbtn_primary$sample_aliquot_id[ ! cbtn_primary$sample_aliquot_id %in% cbtn_primary_clean$sample_aliquot_id]
length(badsamplesize)
# +
CBTN_samples$sufficient_samplesize <- NA
CBTN_samples$sufficient_samplesize[ CBTN_samples$CBTN_sample_aliquot %in% goodsamplesize ] <-
paste0("Sufficient\nsample size\n(n = ", length(goodsamplesize), ")")
CBTN_samples$sufficient_samplesize[ CBTN_samples$CBTN_sample_aliquot %in% badsamplesize ] <-
paste0("Rare/unannotated\n(n = ", length(badsamplesize), ")")
# -
table(CBTN_samples$sufficient_samplesize, useNA = "always")
# ## Clustering cleanup
load(file = paste0(datapath, "/ESTIMATE/estimate_manifest_primary_clean.RData"))
cbtn_preclustering <- estimate_manifest_primary_clean[ estimate_manifest_primary_clean$group == "CBTN",]
dim(cbtn_preclustering)
load(file = paste0(datapath, "ESTIMATE/estimate_manifest_primary_clean_postclustering.RData"))
cbtn_postclustering <- estimate_manifest_primary_clean[ estimate_manifest_primary_clean$group == "CBTN",]
dim(cbtn_postclustering)
# +
cbtn_preclustering$sample_aliquot_id <- paste0(cbtn_preclustering$sample_id, "_",cbtn_preclustering$aliquot_id)
cbtn_postclustering$sample_aliquot_id <- paste0(cbtn_postclustering$sample_id, "_",cbtn_postclustering$aliquot_id)
# -
goodexpression <- cbtn_postclustering$sample_aliquot_id
badexpression <- cbtn_preclustering$sample_aliquot_id[!cbtn_preclustering$sample_aliquot_id %in% cbtn_postclustering$sample_aliquot_id]
length(goodexpression)
length(badexpression)
# +
CBTN_samples$tumorexpression <- NA
CBTN_samples$tumorexpression[ CBTN_samples$CBTN_sample_aliquot %in% goodexpression ] <- paste0("Matched with\npathology\n(n = ",length(goodexpression), ")")
CBTN_samples$tumorexpression[ CBTN_samples$CBTN_sample_aliquot %in% badexpression ] <-
paste0("Not matched with\npathology\n(n = ",length(badexpression), ")")
# -
table(CBTN_samples$tumorexpression, useNA = "always")
# # Sankey plot
freqtab <- as.data.frame(table(CBTN_samples$CBTN, CBTN_samples$Cell_lines, CBTN_samples$duplicated_samples,
CBTN_samples$tumour_type, CBTN_samples$sufficient_samplesize,
CBTN_samples$tumorexpression), stringsAsFactors = F)
freqtab[order(freqtab$Freq, decreasing = T),]
# +
freqtab$Var7 <- NA
freqtab$Var7[ freqtab$Freq == 581] <- "Included\nin the study\n(n = 581)"
freqtab$Var7[ freqtab$Freq != 581] <- "Not included\nin the study\n(n = 415)"
# -
freqtab_ggforce <- gather_set_data(freqtab, c(1:6,8))
freqtab_ggforce$includedsamples[ freqtab_ggforce$Freq == 581] <- "Yes"
freqtab_ggforce$includedsamples[ freqtab_ggforce$Freq != 581] <- "No"
p_plot <- ggplot(freqtab_ggforce, aes(x, id = id, split = y, value = Freq)) +
geom_parallel_sets(aes(fill = includedsamples), alpha = 0.5, axis.width = 0.1) +
geom_parallel_sets_axes(axis.width = 0.3, fill = "dark gray") +
geom_parallel_sets_labels(colour = 'black', size = 8, angle = 0) +
myaxis + myplot +
theme(plot.margin = unit(c(1,10,1,1),"cm"),
legend.position = "none",
axis.title.x = element_blank(),
axis.text.x = element_blank(),
axis.line.x = element_blank(),
axis.ticks.x = element_blank()) +
scale_y_continuous(expand = c(0.05, 0.5)) +
scale_x_discrete(expand = c(0.1, 0.1)) +
scale_fill_manual(values = c("Yes" = "#d7191c", "No" = "#2c7bb6"))
p_plot
# +
pdf(paste0(plotpath, "CBTN_alluvial.pdf"),
width = 30, height = 10,
useDingbats = FALSE)
p_plot
dev.off()
# -
| notebooks/CBTN_Sankey.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.1 64-bit
# name: python391jvsc74a57bd04cd7ab41f5fca4b9b44701077e38c5ffd31fe66a6cab21e0214b68d958d0e462
# ---
# +
#@title Default title text
import numpy as np
import pandas as pd
import yfinance as yf
import riskfolio.Portfolio as pf
import riskfolio.Reports as rp
from stocks import stocks
yf.pdr_override()
# Date range
start = '2019-06-06'
end = '2021-04-02'
# Tickers of assets
tickers = stocks
tickers.sort()
# Downloading the data
data = yf.download(tickers, start = start, end = end)
data = data.loc[:,('Adj Close', slice(None))]
data.columns = tickers
assets = data.pct_change().dropna()
Y = assets
# Creating the Portfolio Object
port = pf.Portfolio(returns=Y)
# To display dataframes values in percentage format
pd.options.display.float_format = '{:.4%}'.format
# Choose the risk measure
rm = 'MV' # Standard Deviation
# Estimate inputs of the model (historical estimates)
method_mu='hist' # Method to estimate expected returns based on historical data.
method_cov='hist' # Method to estimate covariance matrix based on historical data.
port.assets_stats(method_mu=method_mu, method_cov=method_cov, d=0.94)
returns = port.returns
# Estimate the portfolio that maximizes the risk adjusted return ratio
w = port.optimization(model='Classic', rm=rm, obj='Sharpe', rf=0.0, l=0, hist=True)
ax = rp.jupyter_report(returns, w, rm='MV', rf=0, alpha=0.05, height=6, width=14,
others=0.05, nrow=25)
| notebooks/basic_porfolio.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
airbnb_df = pd.read_csv("../data/processed/sf_airbnb_listings_cleaned.csv").drop(columns=["Unnamed: 0", "Unnamed: 0.1"])
damage_df = pd.read_csv("../data/processed/damagedProperty.csv").drop(columns=["Unnamed: 0"])
homeless_df = pd.read_csv("../data/processed/homelessConcerns.csv").drop(columns=["Unnamed: 0"])
noise_df = pd.read_csv("../data/processed/noiseReports.csv").drop(columns=["Unnamed: 0"])
airbnb_df.iloc[0]
damage_df.iloc[0]
homeless_df.iloc[0]
noise_df.iloc[0]
# +
airbnb_in_d = airbnb_df["supervisor_district"].value_counts().rename("num_airbnb_in_sd")
damages_in_d = damage_df["Supervisor District"].value_counts().rename("num_property_damages_in_sd")
homeless_in_d = homeless_df["Supervisor District"].value_counts().rename("num_homeless_concerns_in_sd")
noises_in_d = noise_df["Supervisor District"].value_counts().rename("num_noise_complaints_in_sd")
combined_df = airbnb_df.join(damages_in_d, on=["supervisor_district"])
combined_df = combined_df.join(homeless_in_d, on=["supervisor_district"])
combined_df = combined_df.join(noises_in_d, on=["supervisor_district"])
combined_df.head()
# +
sup_dis = pd.DataFrame(pd.Series(list(range(1, 12)), index=list(range(1,12)), name="supervisor_district"))
# pd.concat([sup_dis, airbnb_in_d, damages_in_d], axis=1)
airbnb_in_d_df = pd.DataFrame(airbnb_in_d.rename("value"))
airbnb_in_d_df["feature"] = "Airbnb Listings"
damages_in_d_df = pd.DataFrame(damages_in_d.rename("value"))
damages_in_d_df["feature"] = "Property Damages"
homeless_in_d_df = pd.DataFrame(homeless_in_d.rename("value"))
homeless_in_d_df["feature"] = "Homeless Concerns"
noises_in_d_df = pd.DataFrame(noises_in_d.rename("value"))
noises_in_d_df["feature"] = "Noise Complaints"
airbnb_in_d_df = pd.merge(sup_dis, airbnb_in_d_df, left_index=True, right_index=True)
damages_in_d_df = pd.merge(sup_dis, damages_in_d_df, left_index=True, right_index=True)
homeless_in_d_df = pd.merge(sup_dis, homeless_in_d_df, left_index=True, right_index=True)
noises_in_d_df = pd.merge(sup_dis, noises_in_d_df, left_index=True, right_index=True)
combined_nums_df = pd.concat([airbnb_in_d_df, damages_in_d_df, homeless_in_d_df, noises_in_d_df], axis=0).reset_index()
# -
plt.figure(figsize=(12, 5))
plot = sns.barplot(x="supervisor_district", y="value", hue="feature", data=combined_nums_df)
plt.legend(bbox_to_anchor=(1.01, 1), loc=2, borderaxespad=0.)
plt.title("Airbnb Listings per District, Dependent on Factors")
fig = plot.get_figure()
fig.savefig( 'airbnb_combined_sd.png', dpi=300, transparent=False, format="png", bbox_inches="tight")
# +
def normalize_value(x):
min_max = (x["value"].min(), x["value"].max())
x["value_norm"] = x["value"].apply(lambda x: x / min_max[1])
return x
combined_nums_df = combined_nums_df.groupby("feature").apply(normalize_value).reset_index().drop(columns=["index"])
plt.figure(figsize=(12, 5))
plot = sns.barplot(x="supervisor_district", y="value_norm", hue="feature", data=combined_nums_df)
plt.legend(bbox_to_anchor=(1.01, 1), loc=2, borderaxespad=0.)
plt.title("Airbnb Listings per District, Dependent on Factors, Normalized per Feature")
fig = plot.get_figure()
fig.savefig( 'airbnb_combined_norm_sd.png', dpi=300, transparent=False, format="png", bbox_inches="tight")
| notebooks/Airbnb Combo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: py35-paddle1.2.0
# ---
#
# # Text Recognition Algorithm Theory
#
# This chapter mainly introduces the theoretical knowledge of text recognition algorithms, including background introduction, algorithm classification and some classic paper ideas.
#
# Through the study of this chapter, you can master:
#
# 1. The goal of text recognition
#
# 2. Classification of text recognition algorithms
#
# 3. Typical ideas of various algorithms
#
#
# ## 1 Background Introduction
#
# Text recognition is a subtask of OCR (Optical Character Recognition), and its task is to recognize the text content of a fixed area. In the two-stage method of OCR, it is followed by text detection and converts image information into text information.
#
# Specifically, the model inputs a positioned text line, and the model predicts the text content and confidence level in the picture. The visualization results are shown in the following figure:
#
# <center><img src=https://ai-studio-static-online.cdn.bcebos.com/a7c3404f778b489db9c1f686c7d2ff4d63b67c429b454f98b91ade7b89f8e903 width="600"></center>
#
# <center><img src=https://ai-studio-static-online.cdn.bcebos.com/e72b1d6f80c342ac951d092bc8c325149cebb3763ec849ec8a2f54e7c8ad60ca width="600"></center>
# <br><center>Figure 1: Visualization results of model predicttion</center>
#
# There are many application scenarios for text recognition, including document recognition, road sign recognition, license plate recognition, industrial number recognition, etc. According to actual scenarios, text recognition tasks can be divided into two categories: **Regular text recognition** and **Irregular Text recognition**.
#
# * Regular text recognition: mainly refers to printed fonts, scanned text, etc., and the text is considered to be roughly in the horizontal position
#
# * Irregular text recognition: It often appears in natural scenes, and due to the huge differences in text curvature, direction, deformation, etc., the text is often not in the horizontal position, and there are problems such as bending, occlusion, and blurring.
#
#
# The figure below shows the data patterns of IC15 and IC13, which represent irregular text and regular text respectively. It can be seen that irregular text often has problems such as distortion, blurring, and large font differences. It is closer to the real scene and is also more challenging.
#
# Therefore, the current major algorithms are trying to obtain higher indicators on irregular data sets.
#
# <center><img src=https://ai-studio-static-online.cdn.bcebos.com/bae4fce1370b4751a3779542323d0765a02a44eace7b44d2a87a241c13c6f8cf width="400">
# <br><center>Figure 2: IC15 picture sample (irregular text)</center>
# <img src=https://ai-studio-static-online.cdn.bcebos.com/b55800d3276f4f5fad170ea1b567eb770177fce226f945fba5d3247a48c15c34 width="400"></center>
# <br><center>Figure 3: IC13 picture sample (rule text)</center>
#
#
# When comparing the capabilities of different recognition algorithms, they are often compared on these two types of public data sets. Comparing the effects on multiple dimensions, currently the more common English benchmark data sets are classified as follows:
#
# <center><img src=https://ai-studio-static-online.cdn.bcebos.com/4d0aada261064031a16816b39a37f2ff6af70dbb57004cb7a106ae6485f14684 width="600"></center>
# <br><center>Figure 4: Common English benchmark data sets</center>
#
# ## 2 Text Recognition Algorithm Classification
#
# In the traditional text recognition method, the task is divided into 3 steps, namely image preprocessing, character segmentation and character recognition. It is necessary to model a specific scene, and it will become invalid once the scene changes. In the face of complex text backgrounds and scene changes, methods based on deep learning have better performance.
#
# Most existing recognition algorithms can be represented by the following unified framework, and the algorithm flow is divided into 4 stages:
#
# 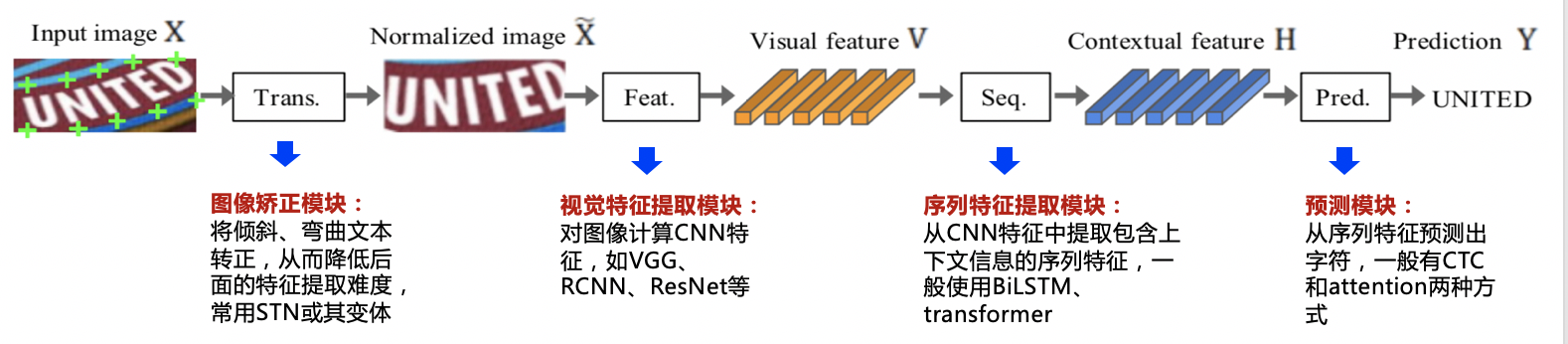
#
#
# We have sorted out the mainstream algorithm categories and main papers, refer to the following table:
# <center>
#
# | Algorithm category | Main ideas | Main papers |
# | -------- | --------------- | -------- |
# | Traditional algorithm | Sliding window, character extraction, dynamic programming |-|
# | ctc | Based on ctc method, sequence is not aligned, faster recognition | CRNN, Rosetta |
# | Attention | Attention-based method, applied to unconventional text | RARE, DAN, PREN |
# | Transformer | Transformer-based method | SRN, NRTR, Master, ABINet |
# | Correction | The correction module learns the text boundary and corrects it to the horizontal direction | RARE, ASTER, SAR |
# | Segmentation | Based on the method of segmentation, extract the character position and then do classification | Text Scanner, Mask TextSpotter |
#
# </center>
#
#
# ### 2.1 Regular Text Recognition
#
#
# There are two mainstream algorithms for text recognition, namely the CTC (Conectionist Temporal Classification)-based algorithm and the Sequence2Sequence algorithm. The difference is mainly in the decoding stage.
#
# The CTC-based algorithm connects the encoded sequence to the CTC for decoding; the Sequence2Sequence-based method connects the sequence to the Recurrent Neural Network (RNN) module for cyclic decoding. Both methods have been verified to be effective and mainstream. Two major practices.
#
# <center><img src=https://ai-studio-static-online.cdn.bcebos.com/f64eee66e4a6426f934c1befc3b138629324cf7360c74f72bd6cf3c0de9d49bd width="600"></center>
# <br><center>Figure 5: Left: CTC-based method, right: Sequece2Sequence-based method </center>
#
#
# #### 2.1.1 Algorithm Based on CTC
#
# The most typical algorithm based on CTC is CRNN (Convolutional Recurrent Neural Network) [1], and its feature extraction part uses mainstream convolutional structures, commonly used ResNet, MobileNet, VGG, etc. Due to the particularity of text recognition tasks, there is a large amount of contextual information in the input data. The convolution kernel characteristics of convolutional neural networks make it more focused on local information and lack long-dependent modeling capabilities, so it is difficult to use only convolutional networks. Dig into the contextual connections between texts. In order to solve this problem, the CRNN text recognition algorithm introduces the bidirectional LSTM (Long Short-Term Memory) to enhance the context modeling. Experiments prove that the bidirectional LSTM module can effectively extract the context information in the picture. Finally, the output feature sequence is input to the CTC module, and the sequence result is directly decoded. This structure has been verified to be effective and widely used in text recognition tasks. Rosetta [2] is a recognition network proposed by FaceBook, which consists of a fully convolutional model and CTC. Gao Y [3] et al. used CNN convolution instead of LSTM, with fewer parameters, and the performance improvement accuracy was the same.
#
# <center><img src=https://ai-studio-static-online.cdn.bcebos.com/d3c96dd9e9794fddb12fa16f926abdd3485194f0a2b749e792e436037490899b width="600"></center>
# <center>Figure 6: CRNN structure diagram </center>
#
#
# #### 2.1.2 Sequence2Sequence algorithm
#
# In the Sequence2Sequence algorithm, the Encoder encodes all input sequences into a unified semantic vector, which is then decoded by the Decoder. In the decoding process of the decoder, the output of the previous moment is continuously used as the input of the next moment, and the decoding is performed in a loop until the stop character is output. The general encoder is an RNN. For each input word, the encoder outputs a vector and hidden state, and uses the hidden state for the next input word to get the semantic vector in a loop; the decoder is another RNN, which receives the encoder Output a vector and output a series of words to create a transformation. Inspired by Sequence2Sequence in the field of translation, Shi [4] proposed an attention-based codec framework to recognize text. In this way, rnn can learn character-level language models hidden in strings from training data.
#
# <center><img src=https://ai-studio-static-online.cdn.bcebos.com/f575333696b7438d919975dc218e61ccda1305b638c5497f92b46a7ec3b85243 width="400" hight="500"></center>
# <center>Figure 7: Sequence2Sequence structure diagram </center>
#
# The above two algorithms have very good effects on regular text, but due to the limitations of network design, this type of method is difficult to solve the task of irregular text recognition of bending and rotation. In order to solve such problems, some algorithm researchers have proposed a series of improved algorithms on the basis of the above two types of algorithms.
#
# ### 2.2 Irregular Text Recognition
#
# * Irregular text recognition algorithms can be divided into 4 categories: correction-based methods; Attention-based methods; segmentation-based methods; and Transformer-based methods.
#
# #### 2.2.1 Correction-based Method
#
# The correction-based method uses some visual transformation modules to convert irregular text into regular text as much as possible, and then uses conventional methods for recognition.
#
# The RARE [4] model first proposed a correction scheme for irregular text. The entire network is divided into two main parts: a spatial transformation network STN (Spatial Transformer Network) and a recognition network based on Sequence2Squence. Among them, STN is the correction module. Irregular text images enter STN and are transformed into a horizontal image through TPS (Thin-Plate-Spline). This transformation can correct curved and transmissive text to a certain extent, and send it to sequence recognition after correction. Network for decoding.
#
# <center><img src=https://ai-studio-static-online.cdn.bcebos.com/66406f89507245e8a57969b9bed26bfe0227a8cf17a84873902dd4a464b97bb5 width="600"></center>
# <center>Figure 8: RARE structure diagram </center>
#
# The RARE paper pointed out that this method has greater advantages in irregular text data sets, especially comparing the two data sets CUTE80 and SVTP, which are more than 5 percentage points higher than CRNN, which proves the effectiveness of the correction module. Based on this [6] also combines a text recognition system with a spatial transformation network (STN) and an attention-based sequence recognition network.
#
# Correction-based methods have better migration. In addition to Attention-based methods such as RARE, STAR-Net [5] applies correction modules to CTC-based algorithms, which is also a good improvement compared to traditional CRNN.
#
# #### 2.2.2 Attention-based Method
#
# The Attention-based method mainly focuses on the correlation between the parts of the sequence. This method was first proposed in the field of machine translation. It is believed that the result of the current word in the process of text translation is mainly affected by certain words, so it needs to be The decisive word has greater weight. The same is true in the field of text recognition. When decoding the encoded sequence, each step selects the appropriate context to generate the next state, which is conducive to obtaining more accurate results.
#
# R^2AM [7] first introduced Attention into the field of text recognition. The model first extracts the encoded image features from the input image through a recursive convolutional layer, and then uses the implicitly learned character-level language statistics to decode the output through a recurrent neural network character. In the decoding process, the Attention mechanism is introduced to realize soft feature selection to make better use of image features. This selective processing method is more in line with human intuition.
#
# <center><img src=https://ai-studio-static-online.cdn.bcebos.com/a64ef10d4082422c8ac81dcda4ab75bf1db285d6b5fd462a8f309240445654d5 width="600"></center>
# <center>Figure 9: R^2AM structure drawing </center>
#
# A large number of algorithms will be explored and updated in the field of Attention in the future. For example, SAR[8] extends 1D attention to 2D attention. The RARE mentioned in the correction module is also a method based on Attention. Experiments prove that the Attention-based method has a good accuracy improvement compared with the CTC method.
#
# <center><img src=https://ai-studio-static-online.cdn.bcebos.com/4e2507fb58d94ec7a9b4d17151a986c84c5053114e05440cb1e7df423d32cb02 width="600"></center>
# <center>Figure 10: Attention diagram</center>
#
#
# #### 2.2.3 Method Based on Segmentation
#
# The method based on segmentation is to treat each character of the text line as an independent individual, and it is easier to recognize the segmented individual characters than to recognize the entire text line after correction. It attempts to locate the position of each character in the input text image, and applies a character classifier to obtain these recognition results, simplifying the complex global problem into a local problem solving, and it has a relatively good effect in the irregular text scene. However, this method requires character-level labeling, and there is a certain degree of difficulty in data acquisition. Lyu [9] et al. proposed an instance word segmentation model for word recognition, which uses a method based on FCN (Fully Convolutional Network) in its recognition part. [10] Considering the problem of text recognition from a two-dimensional perspective, a character attention FCN is designed to solve the problem of text recognition. When the text is bent or severely distorted, this method has better positioning results for both regular and irregular text.
#
# <center><img src=https://ai-studio-static-online.cdn.bcebos.com/fd3e8ef0d6ce4249b01c072de31297ca5d02fc84649846388f890163b624ff10 width="800"></center>
# <center>Figure 11: Mask TextSpotter structure diagram </center>
#
#
#
# #### 2.2.4 Transformer-based Method
#
# With the rapid development of Transformer, both classification and detection fields have verified the effectiveness of Transformer in visual tasks. As mentioned in the regular text recognition part, CNN has limitations in long-dependency modeling. The Transformer structure just solves this problem. It can focus on global information in the feature extractor and can replace additional context modeling modules (LSTM ).
#
# Part of the text recognition algorithm uses Transformer's Encoder structure and convolution to extract sequence features. The Encoder is composed of multiple blocks stacked by MultiHeadAttentionLayer and Positionwise Feedforward Layer. The self-attention in MulitHeadAttention uses matrix multiplication to simulate the timing calculation of RNN, breaking the barrier of long-term dependence on timing in RNN. There are also some algorithms that use Transformer's Decoder module to decode, which can obtain stronger semantic information than traditional RNNs, and parallel computing has higher efficiency.
#
# The SRN[11] algorithm connects the Encoder module of Transformer to ResNet50 to enhance the 2D visual features. A parallel attention module is proposed, which uses the reading order as a query, making the calculation independent of time, and finally outputs the aligned visual features of all time steps in parallel. In addition, SRN also uses Transformer's Eecoder as a semantic module to integrate the visual information and semantic information of the picture, which has greater benefits in irregular text such as occlusion and blur.
#
# NRTR [12] uses a complete Transformer structure to encode and decode the input picture, and only uses a few simple convolutional layers for high-level feature extraction, and verifies the effectiveness of the Transformer structure in text recognition.
#
# <center><img src=https://ai-studio-static-online.cdn.bcebos.com/e7859f4469a842f0bd450e7e793a679d6e828007544241d09785c9b4ea2424a2 width="800"></center>
# <center>Figure 12: NRTR structure drawing </center>
#
# SRACN [13] uses Transformer's decoder to replace LSTM, once again verifying the efficiency and accuracy advantages of parallel training.
#
# ## 3 Summary
#
# This section mainly introduces the theoretical knowledge and mainstream algorithms related to text recognition, including CTC-based methods, Sequence2Sequence-based methods, and segmentation-based methods. The ideas and contributions of classic papers are listed respectively. The next section will explain the practical course based on the CRNN algorithm, from networking to optimization to complete the entire training process,
#
# ## 4 Reference
#
#
# [1]<NAME>., <NAME>., & <NAME>. (2016). An end-to-end trainable neural network for image-based sequence recognition and its application to scene text recognition. IEEE transactions on pattern analysis and machine intelligence, 39(11), 2298-2304.
#
# [2]<NAME>, <NAME>, and <NAME>. Rosetta: Large scale system for text detection and recognition in images. In Proceedings of the 24th ACM SIGKDD International Conference on Knowledge Discovery & Data Mining, pages 71–79. ACM, 2018.
#
# [3]<NAME>., <NAME>., <NAME>., & <NAME>. (2017). Reading scene text with attention convolutional sequence modeling. arXiv preprint arXiv:1709.04303.
#
# [4]<NAME>., <NAME>., <NAME>., <NAME>., & <NAME>. (2016). Robust scene text recognition with automatic rectification. In Proceedings of the IEEE conference on computer vision and pattern recognition (pp. 4168-4176).
#
# [5] Star-Net <NAME>, <NAME>, <NAME>, et al. Spa- tial transformer networks. In Advances in neural information processing systems, pages 2017–2025, 2015.
#
# [6]<NAME>, <NAME>, <NAME>, <NAME>, <NAME>, and <NAME>. Aster: An attentional scene text recognizer with flexible rectification. IEEE transactions on pattern analysis and machine intelligence, 31(11):855–868, 2018.
#
# [7] <NAME> Y , Osindero S . Recursive Recurrent Nets with Attention Modeling for OCR in the Wild[C]// IEEE Conference on Computer Vision & Pattern Recognition. IEEE, 2016.
#
# [8]<NAME>., <NAME>., <NAME>., & <NAME>. (2019, July). Show, attend and read: A simple and strong baseline for irregular text recognition. In Proceedings of the AAAI Conference on Artificial Intelligence (Vol. 33, No. 01, pp. 8610-8617).
#
# [9]<NAME>, <NAME>, <NAME>, <NAME>, and <NAME>. Multi-oriented scene text detection via corner localization and region segmentation. In Proc. CVPR, pages 7553–7563, 2018.
#
# [10] <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., ... & <NAME>. (2019, July). Scene text recognition from two-dimensional perspective. In Proceedings of the AAAI Conference on Artificial Intelligence (Vol. 33, No. 01, pp. 8714-8721).
#
# [11] <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., & <NAME>. (2020). Towards accurate scene text recognition with semantic reasoning networks. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (pp. 12113-12122).
#
# [12] <NAME>., <NAME>., & <NAME>. (2019, September). NRTR: A no-recurrence sequence-to-sequence model for scene text recognition. In 2019 International Conference on Document Analysis and Recognition (ICDAR) (pp. 781-786). IEEE.
#
# [13]<NAME>., <NAME>., <NAME>., <NAME>., & <NAME>. (2020). A holistic representation guided attention network for scene text recognition. Neurocomputing, 414, 67-75.
#
# [14]<NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., ... & <NAME>. (2020, April). Decoupled attention network for text recognition. In Proceedings of the AAAI Conference on Artificial Intelligence (Vol. 34, No. 07, pp. 12216-12224).
#
# [15] <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., & <NAME>. (2021). From two to one: A new scene text recognizer with visual language modeling network. In Proceedings of the IEEE/CVF International Conference on Computer Vision (pp. 14194-14203).
#
# [16] <NAME>., <NAME>., <NAME>., <NAME>., & <NAME>. (2021). Read Like Humans: Autonomous, Bidirectional and Iterative Language Modeling for Scene Text Recognition. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (pp. 7098-7107).
#
# [17] <NAME>., <NAME>., <NAME>., & <NAME>. (2021). Primitive Representation Learning for Scene Text Recognition. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (pp. 284-293).
| notebook/notebook_en/3.text_recognition/text_recognition_theory.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="Q-T844HDTKad"
# <a href="https://colab.research.google.com/github/adasegroup/ML2022_seminars/blob/master/seminar5/Trees_Bagging_Random_Forest.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="Ha8adx-dTKah"
# # Seminar: Trees, Bootstrap Aggregation (Bagging) and Random Forest
# Machine Learning by professor <NAME>
# <br\>
# Author: <NAME>
# -
# ! pip install --upgrade scikit-learn
# + colab={} colab_type="code" id="4aofO-_oTKai"
import numpy as np
import pandas as pd
import math
import matplotlib.pyplot as plt
# %matplotlib inline
pd.set_option('display.max_rows', 5)
# + [markdown] colab_type="text" id="5fM68yzJTKam"
# # Example 1: Regression tree
# + colab={"base_uri": "https://localhost:8080/", "height": 446} colab_type="code" id="Pn8YBI0STKan" outputId="1514c45e-b1c3-466b-8477-cca1ef5c95b0"
# prepare and show a dataset
n = 1 # number of features
N = 100**n # number of samples
np.random.seed(0)
X = np.random.random((N, n))*3
coeffs = 1 + 2 * np.random.random((n, 1))
y = np.sin(np.matmul(X*X, coeffs)) + np.random.random((N, 1))/3
y = y.ravel()
print((pd.DataFrame({'x1': X[:, 0], 'y': y})))
plt.plot(X, y, '*')
plt.title('1 predictor: x1; target: y')
plt.xlabel('x1')
plt.ylabel('y')
# train and predict a regression tree
from sklearn.tree import DecisionTreeRegressor
from sklearn.metrics import mean_squared_error
clf = DecisionTreeRegressor(max_depth=1)
clf.fit(X, y)
y_pred = clf.predict(X)
plt.plot(X, y_pred, '.r')
plt.show()
print('Mean Squared Error: ', mean_squared_error(y, y_pred))
# + [markdown] colab_type="text" id="TARmhz1jTKaq"
# ### Question 1.1: Change the number of levels in a regression tree above until the best approximation of the training set. What is the best MSE?
# + [markdown] colab_type="text" id="edrWTpBATKas"
# ### Question 1.2: Calculate MSE above without `mean_squared_error()` calling.
# + [markdown] colab_type="text" id="ovQfL2AvTKat"
# ## Example 2: Regression tree with $n=2$ features
# + [markdown] colab_type="text" id="sb3_gvdWTKau"
# ### Question 2.1: Lets try an example with $n=2$ features. Train a regression stump (a tree of a depth 1) and see the optimal threshold (border between colors) for the best feature chosen for split among $x_1$ and $x_2$. What feature was chosen and why? Change something in the string
# ```
# coeffs = np.array([[0.2], [1.5]])
# ```
# ### to make another feature is chosen as the best for split.
#
#
# + [markdown] colab_type="text" id="x9LEj01XTKav"
# ### Question 2.2: Find the optimal `max_depth` hyperparameter when MSE on the training set is the best.
# + colab={"base_uri": "https://localhost:8080/", "height": 942} colab_type="code" id="c73kkkA9TKav" outputId="2d5e57b0-0f87-46ee-e3ae-d7ad93777bb4"
# prepare and show a dataset in 2D
n = 2 # number of features
N = 100**n # number of samples
np.random.seed(0)
X = np.random.random((N, n))*1.8
# Change something to make another feature is chosen for split by some optimal threshold
coeffs = np.array([[0.2], [1.5]])
y = np.sin(np.matmul(X*X, coeffs)) + np.random.random((N, 1))/3
# print dataset
print((pd.DataFrame({'x1': X[:, 0], 'x2': X[:, 1], 'y': y.ravel()})))
# show target y in (x1, x2) space
plt.figure(figsize=[9, 6])
sc = plt.scatter(X[:, 0], X[:, 1], c=y.ravel(), s=5)
plt.colorbar(sc)
plt.title('training data: predictors x1, x2 and target y')
plt.xlabel('x1')
plt.ylabel('x2')
plt.show()
# train and predict by a regression tree
from sklearn.tree import DecisionTreeRegressor
from sklearn.metrics import mean_squared_error
clf = DecisionTreeRegressor(max_depth=5)
clf.fit(X, y)
y_pred = clf.predict(X)
# show prediction in (x1, x2) space
plt.figure(figsize=[9, 6])
plt.scatter(X[:, 0], X[:, 1], c=y_pred, s=5)
plt.colorbar(sc)
plt.title('prediction on the training data')
plt.xlabel('x1')
plt.ylabel('x2')
plt.show()
print('Mean Squared Error: ', mean_squared_error(y, y_pred))
# + [markdown] colab_type="text" id="uh-op1b-TKaz"
# ## Example 3: Regression tree: training and testing sets
# + [markdown] colab_type="text" id="Nmo0dNQQTKa0"
# ### Question 3.1: Now we consider training and testing sets. Try different depths of a decision tree to see when the model is underfitted and when the one is overfitted to the training set. Plot the MSE on the testing set depending on `max_depth` hyperparameter. What is the optimal value?
# + colab={"base_uri": "https://localhost:8080/", "height": 825} colab_type="code" id="geXnQ2_GTKa0" outputId="d14083f2-5c64-4b53-ba3c-716c2e9f9e2d"
from sklearn.tree import DecisionTreeRegressor
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import train_test_split
# prepare dataset
n = 1 # number of features
N = 200**n # number of samples
np.random.seed(0)
X = np.random.random((N, n))*3
coeffs = 1 + 2 * np.random.random((n, 1))
y = np.sin(np.matmul(X*X, coeffs)) + np.random.random((N, 1))
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=1/2, random_state=0)
# --- change this block to select the best max_depth
clf = DecisionTreeRegressor(max_depth=1)
clf.fit(X_train, y_train)
print('train MSE: ', mean_squared_error(y_train, clf.predict(X_train)))
print('test MSE: ', mean_squared_error(y_test, clf.predict(X_test)))
# ---
plt.figure(figsize=[9, 6])
plt.plot(X_train, y_train, '*')
plt.plot(X_train, clf.predict(X_train), '.r')
plt.title('training dataset')
plt.xlabel('x1')
plt.ylabel('y')
plt.show()
plt.figure(figsize=[9, 6])
plt.plot(X_test, y_test, '*')
plt.plot(X_test, clf.predict(X_test), '.r')
plt.title('testing dataset')
plt.xlabel('x1')
plt.ylabel('y')
plt.show()
# + [markdown] colab_type="text" id="G8DthEy1Y-TY"
# ### Question 3.2. How many constant-valued regions of red points are on the picture when `max_depth=5`?
#
#
# + [markdown] colab_type="text" id="ODCAU-cxTKa3"
# # Example 4: Bagging = Decision Tree + Bootstrap
# ### In the question above we've found the optimal `max_depth` for the case of single Decision Tree. By limiting the tree depth we distort the fitting to the training dataset and prevent the model from overfitting.
# ### The second way to prevent overfitting is to distort the ... training dataset itself. What is Bagging?
# ### We train many trees each on a Bootstraped training dataset (it contains the same number of samples but some of them are included with some number of their copies, and some of them are not included). Then we average over all such trees. It is called Bootstrap aggregation - Bagging.
# + [markdown] colab_type="text" id="bye47AOmTKa4"
# ### Question 4.1: Compare the prediction above (single tree) with Bagging all with `max_depth=5`. Why Bagging approximation red dots does not look like constant-valued regions as in a single decision tree? Tune the best number of trees. Has Bagging improved the single tree model regarding `test MSE`?
# + [markdown] colab_type="text" id="7d8jyfDATKa5"
# ### Question 4.2: Sometimes it is good to be a perfectionist and unittest any code :). Unittest scikit-learn! Should `DecisionTreeRegressor` and `BaggingRegressor` give the same results in some special case? Simplify Bagging to the single Decision Tree and show the same pictures as above when `max_depth=5`. What parameter `n_estimators` have to be set to? Do we need to change any other hyperparameters?
# + colab={"base_uri": "https://localhost:8080/", "height": 825} colab_type="code" id="AsxzenebTKa5" outputId="c21e9e12-dab6-4614-95c4-e42ebc63ac69"
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import BaggingRegressor
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import train_test_split
# prepare dataset
n = 1 # number of features
N = 200**n # number of samples
np.random.seed(0)
X = np.random.random((N, n))*3
coeffs = 1 + 2 * np.random.random((n, 1))
y = np.sin(np.matmul(X*X, coeffs)) + np.random.random((N, 1))
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=1/2, random_state=0)
# --- 1. change this block to select the best n_estimators
# --- 2. change this block to simplify Bagging to ordinary single decision tree
clf = BaggingRegressor(DecisionTreeRegressor(max_depth=12), n_estimators=10, bootstrap=True, random_state=0)
clf.fit(X_train, y_train.ravel())
print('train MSE: ', mean_squared_error(y_train, clf.predict(X_train)))
print('test MSE: ', mean_squared_error(y_test, clf.predict(X_test)))
# ---
# plt.figure(figsize=[9, 6])
plt.plot(X_train, y_train, '*')
plt.plot(X_train, clf.predict(X_train), '.r')
plt.title('training dataset')
plt.xlabel('x1')
plt.ylabel('y')
plt.show()
# plt.figure(figsize=[9, 6])
plt.plot(X_test, y_test, '*')
plt.plot(X_test, clf.predict(X_test), '.r')
plt.title('testing dataset')
plt.xlabel('x1')
plt.ylabel('y')
plt.show()
# + [markdown] colab_type="text" id="5_nx2p4NfCBo"
# ### Question 4.3: Estimate the part of points that are not selected after Bootstrap procedure analytically. Consider the sample of a size $N\to\infty$.
# + [markdown] colab_type="text" id="qujVV30ATKa9"
# # Example 5: Random Forest = Bagging + (`max_features' < $n$)
# + [markdown] colab_type="text" id="5nXxzoGdTKa-"
# ## Can we reduce the overfitting more? - YES!!! <br> Along with Bootstrap reduce the number of features among which the best feature for each tree in ensemble is chosen!
# + [markdown] colab_type="text" id="j-xYYau3TKa_"
# ### Question 5.1: Tune `max_features` to improve the tesing MSE. Check that testing MSE becomes better. And what happens with training MSE and why?
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="vF0oi4utTKa_" outputId="19f922e5-e6ed-4c72-d1d0-f40a32621215"
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import train_test_split
# prepare dataset
n = 100 # number of features
N = 10**4 # number of samples
np.random.seed(0)
X = np.random.random((N, n))*3
coeffs = 1 + 2 * np.random.random((n, 1))
y = np.sin(np.matmul(X*X, coeffs)) + np.random.random((N, 1))
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=1/2, random_state=0)
# --- change this block to select the best max_features
clf = RandomForestRegressor(max_depth=5, n_estimators=10, max_features=n, random_state=0)
clf.fit(X_train, y_train.ravel())
print('train MSE: ', mean_squared_error(y_train, clf.predict(X_train)))
print('test MSE: ', mean_squared_error(y_test, clf.predict(X_test)))
# ---
# -
# ## Random Forest on Kaggle data
# ### Below shorter trees and 'max_features' < all features performs better.
# +
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import roc_curve, roc_auc_score
X_train = pd.read_csv('https://raw.githubusercontent.com/adasegroup/ML2022_seminars/master/seminar5/give_me_some_credit.csv', index_col=0)
X_train = X_train.dropna()
y_train = X_train['SeriousDlqin2yrs']
X_train = X_train.drop(['SeriousDlqin2yrs'], axis=1)
X_train = X_train.iloc[::10, :] # use only each 10th sample to save a time
y_train = y_train.iloc[::10]
n_features = X_train.shape[1]
clf = GridSearchCV(RandomForestClassifier(random_state=0),
# you can play with tuning, up to your CPU performance:
{'max_depth': [10, 15], 'max_features': ['auto', n_features]},
scoring = 'roc_auc',
cv = 3,
n_jobs=-1)
# You can see below that detailed trees i.e. with high depth and all features on a split are worse,
# GridSearchCV() choses shorter depth and sqrt() of all features.
# This is because we use cross-validation (cv=3), and it prevents from overfitting
clf.fit(X_train, y_train)
print('best parameters:', clf.best_params_)
print('ROC_AUC score:', roc_auc_score(y_train, clf.predict(X_train)))
# now let's draw ROC curve
plt.figure(figsize=[9, 6])
fpr, tpr, _ = roc_curve(y_train, clf.predict_proba(X_train)[:, 1])
plt.plot(fpr, tpr, 'r', label='train')
plt.title('ROC curve')
plt.xlabel('False positive rate')
plt.ylabel('True positive rate')
plt.show()
# + [markdown] colab_type="text" id="MVEms_2pTKbC"
# # Example 6: Feature Importance
# +
X_train = pd.read_csv('https://raw.githubusercontent.com/adasegroup/ML2022_seminars/master/seminar5/give_me_some_credit.csv', index_col=0)
y_train = X_train['SeriousDlqin2yrs']
X_train = X_train.drop(['SeriousDlqin2yrs'], axis=1)
X_train = X_train.iloc[::10, :] # use only each 10th sample to save a time
y_train = y_train.iloc[::10]
clf = RandomForestClassifier(max_depth = 10, max_features = 'auto', random_state=0)
clf.fit(X_train, y_train)
print('ROC_AUC score:', roc_auc_score(y_train, clf.predict(X_train)))
# feature importances
fi = pd.Series(clf.feature_importances_, index=X_train.columns)
fi.sort_values(ascending=False).plot(kind='bar')
plt.title('feature importances')
plt.ylim([0, 0.2])
plt.show()
# + [markdown] colab_type="text" id="n_3ZE7fUTKbI"
# ### Be careful with `feature_importances`: after adding extremly correlated features (here they are even copies) the values of importance decrease (see the scale along vertical axis). This is expected, because similar features can share their common similar importance.
# + colab={"base_uri": "https://localhost:8080/", "height": 491} colab_type="code" id="W0PtboZXTKbJ" outputId="74415fbe-d62f-409e-92f4-4f82e888e797"
X_train = pd.read_csv('https://raw.githubusercontent.com/adasegroup/ML2022_seminars/master/seminar5/give_me_some_credit.csv', index_col=0)
X_train = X_train.dropna()
y_train = X_train['SeriousDlqin2yrs']
X_train = X_train.drop(['SeriousDlqin2yrs'], axis=1)
X_train = X_train.iloc[::10, :] # use only each 10th sample to save a time
y_train = y_train.iloc[::10]
X_train['NumberOfTimes90DaysLate_2'] = X_train['NumberOfTimes90DaysLate']
X_train['NumberOfTimes90DaysLate_3'] = X_train['NumberOfTimes90DaysLate']
X_train['NumberOfTimes90DaysLate_4'] = X_train['NumberOfTimes90DaysLate']
X_train['NumberOfTimes90DaysLate_5'] = X_train['NumberOfTimes90DaysLate']
clf = RandomForestClassifier(max_depth = 10, max_features = 'auto', random_state=0)
clf.fit(X_train, y_train)
print('ROC_AUC score:', roc_auc_score(y_train, clf.predict(X_train)))
# now let's draw ROC curve
plt.figure(figsize=[9, 6])
fpr, tpr, _ = roc_curve(y_train, clf.predict_proba(X_train)[:, 1])
plt.plot(fpr, tpr, 'r', label='train')
plt.title('ROC curve')
plt.xlabel('False positive rate')
plt.ylabel('True positive rate')
plt.show()
# feature importances
fi = pd.Series(clf.feature_importances_, index=X_train.columns)
fi.sort_values(ascending=False).plot(kind='bar')
plt.title('feature importances')
plt.ylim([0, 0.2])
plt.show()
# -
# ## Feature Selection and Feature Importance
#
# ### Above, the feature 'NumberOfTimes90DaysLate' moved from the top to the tail together with its highly correlated (even cloned) 'friends'. Now they share their importance together. Do not think that top features are really strong and tail ones are weak! The can correlate significantly.
#
# ### So, do not rely only on Feature Importance! Try to reduce the dimension, for. ex. by Feature Selection
#
# ### In the following example if sequentially add the strongest feature one can recover the original feature set
# +
# from sklearn.feature_selection import RFE
from sklearn.feature_selection import SequentialFeatureSelector
from sklearn.metrics import roc_curve, roc_auc_score
X_train = pd.read_csv('https://raw.githubusercontent.com/adasegroup/ML2022_seminars/master/seminar5/give_me_some_credit.csv', index_col=0)
X_train = X_train.dropna()
y_train = X_train['SeriousDlqin2yrs']
X_train = X_train.drop(['SeriousDlqin2yrs'], axis=1)
X_train = X_train.iloc[::10, :] # use only each 10th sample to save a time
y_train = y_train.iloc[::10]
X_train['NumberOfTimes90DaysLate_2'] = X_train['NumberOfTimes90DaysLate']
X_train['NumberOfTimes90DaysLate_3'] = X_train['NumberOfTimes90DaysLate']
X_train['NumberOfTimes90DaysLate_4'] = X_train['NumberOfTimes90DaysLate']
X_train['NumberOfTimes90DaysLate_5'] = X_train['NumberOfTimes90DaysLate']
clf = RandomForestClassifier(max_depth = 10, max_features ='auto', random_state = 0)
clf_SFS = SequentialFeatureSelector(clf, n_features_to_select=10, direction='forward', scoring='roc_auc', cv=3, n_jobs=-1)
# clf = RFE(clf, n_features_to_select=3, step=1)
clf_SFS.fit(X_train, y_train)
X_selected = X_train.iloc[:, clf_SFS.get_support()]
clf.fit(X_selected, y_train)
print('ROC_AUC score:', roc_auc_score(y_train, clf.predict(X_selected)))
# now let's draw ROC curve
plt.figure(figsize=[9, 6])
fpr, tpr, _ = roc_curve(y_train, clf.predict_proba(X_selected)[:, 1])
plt.plot(fpr, tpr, 'r', label='train')
plt.title('ROC curve')
plt.xlabel('False positive rate')
plt.ylabel('True positive rate')
plt.show()
# feature importances
fi = pd.Series(clf.feature_importances_, index=X_selected.columns)
fi.sort_values(ascending=False).plot(kind='bar')
plt.title('feature importances')
# plt.ylim([0, 0.2])
plt.show()
| seminar5/Trees_Bagging_Random_Forest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: TensorFlow 2.3 on Python 3.6 (CUDA 10.1)
# language: python
# name: python3
# ---
# **도구 - 판다스(pandas)**
#
# *`pandas` 라이브러리는 사용하기 쉬운 고성능 데이터 구조와 데이터 분석 도구를 제공합니다. 주 데이터 구조는 `DataFrame`입니다. 이를 인-메모리(in-memory) 2D 테이블로 생각할 수 있습니다(열 이름과 행 레이블이 있는 스프레드시트와 비슷합니다). 엑셀에 있는 많은 기능을 프로그램에서 사용할 수 있습니다. 여기에는 피봇 테이블이나 다른 열을 기반으로 열을 계산하고 그래프 출력하는 기능 등이 포함됩니다. 열 값으로 행을 그룹핑할 수도 있습니다. 또한 SQL과 비슷하게 테이블을 조인할 수 있습니다. 판다스는 시계열 데이터를 다루는데도 뛰어납니다.*
#
# 필요 라이브러리:
#
# * 넘파이(NumPy) – 넘파이에 익숙하지 않다면 지금 [넘파이 튜토리얼](tools_numpy.ipynb)을 둘러 보세요.
# <table align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/rickiepark/handson-ml2/blob/master/tools_pandas.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />구글 코랩에서 실행하기</a>
# </td>
# </table>
# # 설정
# 먼저 `pandas`를 임포트합니다. 보통 `pd`로 임포트합니다:
import pandas as pd
# # `Series` 객체
#
# `pandas` 라이브러리는 다음과 같은 유용한 데이터 구조를 포함하고 있습니다:
#
# * `Series` 객체를 곧 이어서 설명하겠습니다. `Series` 객체는 1D 배열입니다. (열 이름과 행 레이블을 가진) 스프레드시트의 열과 비슷합니다.
# * `DataFrame` 객체는 2D 테이블입니다. (열 이름과 행 레이블을 가진) 스프레드시트와 비슷합니다.
# ## `Series` 만들기
#
# 첫 번째 `Series` 객체를 만들어 보죠!
s = pd.Series([2,-1,3,5])
s
# ## 1D `ndarray`와 비슷합니다
#
# `Series` 객체는 넘파이 `ndarray`와 비슷하게 동작합니다. 넘파이 함수에 매개변수로 종종 전달할 수 있습니다:
import numpy as np
np.exp(s)
# `Series` 객체에 대한 산술 연산도 가능합니다. `ndarray`와 비슷하게 *원소별*로 적용됩니다:
s + [1000,2000,3000,4000]
# 넘파이와 비슷하게 `Series`에 하나의 숫자를 더하면 `Series`에 있는 모든 원소에 더해집니다. 이를 *브로드캐스팅*(broadcasting)이라고 합니다:
s + 1000
# `*`나 `/` 같은 모든 이항 연산과 심지어 조건 연산에서도 마찬가지입니다:
s < 0
# ## 인덱스 레이블
#
# `Series` 객체에 있는 각 원소는 *인덱스 레이블*(index label)이라 불리는 고유한 식별자를 가지고 있습니다. 기본적으로 `Series`에 있는 원소의 순서입니다(`0`에서 시작합니다). 하지만 수동으로 인덱스 레이블을 지정할 수도 있습니다:
s2 = pd.Series([68, 83, 112, 68], index=["alice", "bob", "charles", "darwin"])
s2
# 그다음 `dict`처럼 `Series`를 사용할 수 있습니다:
s2["bob"]
# 일반 배열처럼 정수 인덱스를 사용하여 계속 원소에 접근할 수 있습니다:
s2[1]
# 레이블이나 정수를 사용해 접근할 때 명확하게 하기 위해 레이블은 `loc` 속성을 사용하고 정수는 `iloc` 속성을 사용하는 것이 좋습니다:
s2.loc["bob"]
s2.iloc[1]
# `Series`는 인덱스 레이블을 슬라이싱할 수도 있습니다:
s2.iloc[1:3]
# 기본 정수 레이블을 사용할 때 예상 외의 결과를 만들 수 있기 때문에 주의해야 합니다:
surprise = pd.Series([1000, 1001, 1002, 1003])
surprise
surprise_slice = surprise[2:]
surprise_slice
# 보세요. 첫 번째 원소의 인덱스 레이블이 `2`입니다. 따라서 슬라이싱 결과에서 인덱스 레이블 `0`인 원소는 없습니다:
try:
surprise_slice[0]
except KeyError as e:
print("키 에러:", e)
# 하지만 `iloc` 속성을 사용해 정수 인덱스로 원소에 접근할 수 있습니다. `Series` 객체를 사용할 때 `loc`와 `iloc`를 사용하는 것이 좋은 이유입니다:
surprise_slice.iloc[0]
# ## `dict`에서 초기화
#
# `dict`에서 `Series` 객체를 만들 수 있습니다. 키는 인덱스 레이블로 사용됩니다:
weights = {"alice": 68, "bob": 83, "colin": 86, "darwin": 68}
s3 = pd.Series(weights)
s3
# `Series`에 포함할 원소를 제어하고 `index`를 지정하여 명시적으로 순서를 결정할 수 있습니다:
s4 = pd.Series(weights, index = ["colin", "alice"])
s4
# ## 자동 정렬
#
# 여러 개의 `Series` 객체를 다룰 때 `pandas`는 자동으로 인덱스 레이블에 따라 원소를 정렬합니다.
# +
print(s2.keys())
print(s3.keys())
s2 + s3
# -
# 만들어진 `Series`는 `s2`와 `s3`의 인덱스 레이블의 합집합을 담고 있습니다. `s2`에 `"colin"`이 없고 `s3`에 `"charles"`가 없기 때문에 이 원소는 `NaN` 값을 가집니다(Not-a-Number는 *누락*이란 의미입니다).
#
# 자동 정렬은 구조가 다고 누락된 값이 있는 여러 데이터를 다룰 때 매우 편리합니다. 하지만 올바른 인덱스 레이블을 지정하는 것을 잊는다면 원치않는 결과를 얻을 수 있습니다:
# +
s5 = pd.Series([1000,1000,1000,1000])
print("s2 =", s2.values)
print("s5 =", s5.values)
s2 + s5
# -
# 레이블이 하나도 맞지 않기 때문에 판다스가 이 `Series`를 정렬할 수 없습니다. 따라서 모두 `NaN`이 되었습니다.
# ## 스칼라로 초기화
#
# 스칼라와 인덱스 레이블의 리스트로 `Series` 객체를 초기화할 수도 있습니다: 모든 원소가 이 스칼라 값으로 설정됩니다.
meaning = pd.Series(42, ["life", "universe", "everything"])
meaning
# ## `Series` 이름
#
# `Series`는 `name`을 가질 수 있습니다:
s6 = pd.Series([83, 68], index=["bob", "alice"], name="weights")
s6
# ## `Series` 그래프 출력
#
# 맷플롯립을 사용해 `Series` 데이터를 쉽게 그래프로 출력할 수 있습니다(맷플롯립에 대한 자세한 설명은 [맷플롯립 튜토리얼](tools_matplotlib.ipynb)을 참고하세요). 맷플롯립을 임포트하고 `plot()` 메서드를 호출하면 끝입니다:
# %matplotlib inline
import matplotlib.pyplot as plt
temperatures = [4.4,5.1,6.1,6.2,6.1,6.1,5.7,5.2,4.7,4.1,3.9,3.5]
s7 = pd.Series(temperatures, name="Temperature")
s7.plot()
plt.show()
# 데이터를 그래프로 출력하는데 많은 *옵션*이 있습니다. 여기에서 모두 나열할 필요는 없습니다. 특정 종류의 그래프(히스토그램, 파이 차트 등)가 필요하면 판다스 문서의 [시각화](http://pandas.pydata.org/pandas-docs/stable/visualization.html) 섹션에서 예제 코드를 참고하세요.
# # 시간 다루기
#
# 많은 데이터셋에 타임스탬프가 포함되어 있습니다. 판다스는 이런 데이터를 다루는데 뛰어납니다:
# * (2016Q3 같은) 기간과 ("monthly" 같은) 빈도를 표현할 수 있습니다.
# * 기간을 실제 타임스탬프로 변환하거나 그 반대로 변환할 수 있습니다.
# * 데이터를 리샘플링하고 원하는 방식으로 값을 모을 수 있습니다.
# * 시간대를 다룰 수 있습니다.
#
# ## 시간 범위
#
# 먼저 `pd.date_range()`를 사용해 시계열을 만들어 보죠. 이 함수는 2016년 10월 29일 5:30pm에서 시작하여 12시간마다 하나의 datetime을 담고 있는 `DatetimeIndex`를 반환합니다.
dates = pd.date_range('2016/10/29 5:30pm', periods=12, freq='H')
dates
# 이 `DatetimeIndex`를 `Series`의 인덱스로 사용할수 있습니다:
temp_series = pd.Series(temperatures, dates)
temp_series
# 이 시리즈를 그래프로 출력해 보죠:
# +
temp_series.plot(kind="bar")
plt.grid(True)
plt.show()
# -
# ## 리샘플링
#
# 판다스는 매우 간단하게 시계열을 리샘플링할 수 있습니다. `resample()` 메서드를 호출하고 새로운 주기를 지정하면 됩니다:
temp_series_freq_2H = temp_series.resample("2H")
temp_series_freq_2H
# 리샘플링 연산은 사실 지연된 연산입니다. 그래서 `Series` 객체 대신 `DatetimeIndexResampler` 객체가 반환됩니다. 실제 리샘플링 연산을 수행하려면 `mean()` 같은 메서드를 호출할 수 있습니다. 이 메서드는 연속적인 시간 쌍에 대해 평균을 계산합니다:
temp_series_freq_2H = temp_series_freq_2H.mean()
# 결과를 그래프로 출력해 보죠:
temp_series_freq_2H.plot(kind="bar")
plt.show()
# 2시간 간격으로 어떻게 값이 수집되었는지 확인해 보세요. 예를 들어 6-8pm 간격을 보면 6:30pm에서 `5.1`이고 7:30pm에서 `6.1`입니다. 리샘플링 후에 `5.1`과 `6.1`의 평균인 `5.6` 하나를 얻었습니다. 평균말고 어떤 집계 함수(aggregation function)도 사용할 수 있습니다. 예를 들어 각 기간에서 최솟값을 찾을 수 있습니다:
temp_series_freq_2H = temp_series.resample("2H").min()
temp_series_freq_2H
# 또는 동일한 효과를 내는 `apply()` 메서드를 사용할 수 있습니다:
temp_series_freq_2H = temp_series.resample("2H").apply(np.min)
temp_series_freq_2H
# ## 업샘플링과 보간
#
# 다운샘플링의 예를 보았습니다. 하지만 업샘플링(즉, 빈도를 높입니다)도 할 수 있습니다. 하지만 데이터에 구멍을 만듭니다:
temp_series_freq_15min = temp_series.resample("15Min").mean()
temp_series_freq_15min.head(n=10) # `head`는 상위 n 개의 값만 출력합니다
# 한가지 방법은 보간으로 사이를 채우는 것입니다. 이렇게 하려면 `interpolate()` 메서드를 호출합니다. 기본값은 선형 보간이지만 3차 보간(cubic interpolation) 같은 다른 방법을 선택할 수 있습니다:
temp_series_freq_15min = temp_series.resample("15Min").interpolate(method="cubic")
temp_series_freq_15min.head(n=10)
temp_series.plot(label="Period: 1 hour")
temp_series_freq_15min.plot(label="Period: 15 minutes")
plt.legend()
plt.show()
# ## 시간대
#
# 기본적으로 datetime은 *단순*합니다. 시간대(timezone)을 고려하지 않죠. 따라서 2016-10-30 02:30는 파리나 뉴욕이나 2016년 10월 30일 2:30pm입니다. `tz_localize()` 메서드로 시간대를 고려한 datetime을 만들 수 있습니다:
temp_series_ny = temp_series.tz_localize("America/New_York")
temp_series_ny
# 모든 datetime에 `-04:00`이 추가됩니다. 즉 모든 시간은 [UTC](https://en.wikipedia.org/wiki/Coordinated_Universal_Time) - 4시간을 의미합니다.
#
# 다음처럼 파리 시간대로 바꿀 수 있습니다:
temp_series_paris = temp_series_ny.tz_convert("Europe/Paris")
temp_series_paris
# UTC와의 차이가 `+02:00`에서 `+01:00`으로 바뀐 것을 알 수 있습니다. 이는 프랑스가 10월 30일 3am에 겨울 시간으로 바꾸기 때문입니다(2am으로 바뀝니다). 따라서 2:30am이 두 번 등장합니다! 시간대가 없는 표현으로 돌아가 보죠(시간대가 없이 지역 시간으로 매시간 로그를 기록하는 경우 이와 비슷할 것입니다):
temp_series_paris_naive = temp_series_paris.tz_localize(None)
temp_series_paris_naive
# 이렇게 되면 `02:30`이 정말 애매합니다. 시간대가 없는 datetime을 파리 시간대로 바꿀 때 에러가 발생합니다:
try:
temp_series_paris_naive.tz_localize("Europe/Paris")
except Exception as e:
print(type(e))
print(e)
# 다행히 `ambiguous` 매개변수를 사용하면 판다스가 타임스탬프의 순서를 기반으로 적절한 DST(일광 절약 시간제)를 추측합니다:
temp_series_paris_naive.tz_localize("Europe/Paris", ambiguous="infer")
# ## 기간
#
# `pd.period_range()` 함수는 `DatetimeIndex`가 아니라 `PeriodIndex`를 반환합니다. 예를 들어 2016과 2017년의 전체 분기를 가져와 보죠:
quarters = pd.period_range('2016Q1', periods=8, freq='Q')
quarters
# `PeriodIndex`에 숫자 `N`을 추가하면 `PeriodIndex` 빈도의 `N` 배만큼 이동시킵니다:
quarters + 3
# `asfreq()` 메서드를 사용하면 `PeriodIndex`의 빈도를 바꿀 수 있습니다. 모든 기간이 늘어나거나 줄어듭니다. 예를 들어 분기 기간을 모두 월별 기간으로 바꾸어 보죠:
quarters.asfreq("M")
# 기본적으로 `asfreq`는 각 기간의 끝에 맞춥니다. 기간의 시작에 맞추도록 변경할 수 있습니다:
quarters.asfreq("M", how="start")
# 간격을 늘릴 수도 있습니다:
quarters.asfreq("A")
# 물론 `PeriodIndex`로 `Series`를 만들 수 있습니다:
quarterly_revenue = pd.Series([300, 320, 290, 390, 320, 360, 310, 410], index = quarters)
quarterly_revenue
quarterly_revenue.plot(kind="line")
plt.show()
# `to_timestamp`를 호출해서 기간을 타임스탬프로 변경할 수 있습니다. 기본적으로 기간의 첫 번째 날을 반환합니다. 하지만 `how`와 `freq`를 지정해서 기간의 마지막 시간을 얻을 수 있습니다:
last_hours = quarterly_revenue.to_timestamp(how="end", freq="H")
last_hours
# `to_peroid`를 호출하면 다시 기간으로 돌아갑니다:
last_hours.to_period()
# 판다스는 여러 가지 시간 관련 함수를 많이 제공합니다. [온라인 문서](http://pandas.pydata.org/pandas-docs/stable/timeseries.html)를 확인해 보세요. 예를 하나 들면 2016년 매월 마지막 업무일의 9시를 얻는 방법은 다음과 같습니다:
months_2016 = pd.period_range("2016", periods=12, freq="M")
one_day_after_last_days = months_2016.asfreq("D") + 1
last_bdays = one_day_after_last_days.to_timestamp() - pd.tseries.offsets.BDay()
last_bdays.to_period("H") + 9
# # `DataFrame` 객체
#
# 데이터프레임 객체는 스프레드시트를 표현합니다. 셀 값, 열 이름, 행 인덱스 레이블을 가집니다. 다른 열을 바탕으로 열을 계산하는 식을 쓸 수 있고 피봇 테이블을 만들고, 행을 그룹핑하고, 그래프를 그릴 수 있습니다. `DataFrame`을 `Series`의 딕셔너리로 볼 수 있습니다.
#
# ## `DataFrame` 만들기
#
# `Series` 객체의 딕셔너리를 전달하여 데이터프레임을 만들 수 있습니다:
people_dict = {
"weight": pd.Series([68, 83, 112], index=["alice", "bob", "charles"]),
"birthyear": pd.Series([1984, 1985, 1992], index=["bob", "alice", "charles"], name="year"),
"children": pd.Series([0, 3], index=["charles", "bob"]),
"hobby": pd.Series(["Biking", "Dancing"], index=["alice", "bob"]),
}
people = pd.DataFrame(people_dict)
people
# 몇가지 알아 두어야 할 것은 다음과 같습니다:
#
# * `Series`는 인덱스를 기반으로 자동으로 정렬됩니다.
# * 누란된 값은 `NaN`으로 표현됩니다.
# * `Series` 이름은 무시됩니다(`"year"`란 이름은 삭제됩니다).
# * `DataFrame`은 주피터 노트북에서 멋지게 출력됩니다!
# 예상하는 방식으로 열을 참조할 수 있고 `Serires` 객체가 반환됩니다:
people["birthyear"]
# 동시에 여러 개의 열을 선택할 수 있습니다:
people[["birthyear", "hobby"]]
# 열 리스트나 행 인덱스 레이블을 `DataFrame` 생성자에 전달하면 해당 열과 행으로 채워진 데이터프레임이 반환됩니다. 예를 들면:
d2 = pd.DataFrame(
people_dict,
columns=["birthyear", "weight", "height"],
index=["bob", "alice", "eugene"]
)
d2
# `DataFrame`을 만드는 또 다른 편리한 방법은 `ndarray`나 리스트의 리스트로 모든 값을 생성자에게 전달하고 열 이름과 행 인덱스 레이블을 각기 지정하는 것입니다:
values = [
[1985, np.nan, "Biking", 68],
[1984, 3, "Dancing", 83],
[1992, 0, np.nan, 112]
]
d3 = pd.DataFrame(
values,
columns=["birthyear", "children", "hobby", "weight"],
index=["alice", "bob", "charles"]
)
d3
# 누락된 값을 지정하려면 `np.nan`이나 넘파이 마스크 배열을 사용합니다:
masked_array = np.ma.asarray(values, dtype=np.object)
masked_array[(0, 2), (1, 2)] = np.ma.masked
d3 = pd.DataFrame(
masked_array,
columns=["birthyear", "children", "hobby", "weight"],
index=["alice", "bob", "charles"]
)
d3
# `ndarray` 대신에 `DataFrame` 객체를 전달할 수도 있습니다:
d4 = pd.DataFrame(
d3,
columns=["hobby", "children"],
index=["alice", "bob"]
)
d4
# 딕셔너리의 딕셔너리(또는 리스트의 리스트)로 `DataFrame`을 만들 수 있습니다:
people = pd.DataFrame({
"birthyear": {"alice":1985, "bob": 1984, "charles": 1992},
"hobby": {"alice":"Biking", "bob": "Dancing"},
"weight": {"alice":68, "bob": 83, "charles": 112},
"children": {"bob": 3, "charles": 0}
})
people
# ## 멀티 인덱싱
#
# 모든 열이 같은 크기의 튜플이면 멀티 인덱스로 인식합니다. 열 인덱스 레이블에도 같은 방식이 적용됩니다. 예를 들면:
d5 = pd.DataFrame(
{
("public", "birthyear"):
{("Paris","alice"):1985, ("Paris","bob"): 1984, ("London","charles"): 1992},
("public", "hobby"):
{("Paris","alice"):"Biking", ("Paris","bob"): "Dancing"},
("private", "weight"):
{("Paris","alice"):68, ("Paris","bob"): 83, ("London","charles"): 112},
("private", "children"):
{("Paris", "alice"):np.nan, ("Paris","bob"): 3, ("London","charles"): 0}
}
)
d5
# 이제 `"public"` 열을 모두 담은 `DataFrame`을 손쉽게 만들 수 있습니다:
d5["public"]
d5["public", "hobby"] # d5["public"]["hobby"]와 같습니다.
# ## 레벨 낮추기
#
# `d5`를 다시 확인해 보죠:
d5
# 열의 레벨(level)이 2개이고 인덱스 레벨이 2개입니다. `droplevel()`을 사용해 열 레벨을 낮출 수 있습니다(인덱스도 마찬가지입니다):
d5.columns = d5.columns.droplevel(level = 0)
d5
# ## 전치
#
# `T` 속성을 사용해 열과 인덱스를 바꿀 수 있습니다:
d6 = d5.T
d6
# ## 레벨 스택과 언스택
#
# `stack()` 메서드는 가장 낮은 열 레벨을 가장 낮은 인덱스 뒤에 추가합니다:
d7 = d6.stack()
d7
# `NaN` 값이 생겼습니다. 이전에 없던 조합이 생겼기 때문입니다(예를 들어 `London`에 `bob`이 없었습니다).
#
# `unstack()`을 호출하면 반대가 됩니다. 여기에서도 많은 `NaN` 값이 생성됩니다.
d8 = d7.unstack()
d8
# `unstack`을 다시 호출하면 `Series` 객체가 만들어 집니다:
d9 = d8.unstack()
d9
# `stack()`과 `unstack()` 메서드를 사용할 때 스택/언스택할 `level`을 선택할 수 있습니다. 심지어 한 번에 여러 개의 레벨을 스택/언스택할 수도 있습니다:
d10 = d9.unstack(level = (0,1))
d10
# ## 대부분의 메서드는 수정된 복사본을 반환합니다
#
# 눈치챘겠지만 `stack()`과 `unstack()` 메서드는 객체를 수정하지 않습니다. 대신 복사본을 만들어 반환합니다. 판다스에 있는 대부분의 메서드들이 이렇게 동작합니다.
# ## 행 참조하기
#
# `people` `DataFrame`으로 돌아가 보죠:
people
# `loc` 속성으로 열 대신 행을 참조할 수 있습니다. `DataFrame`의 열 이름이 행 인덱스 레이블로 매핑된 `Series` 객체가 반환됩니다:
people.loc["charles"]
# `iloc` 속성을 사용해 정수 인덱스로 행을 참조할 수 있습니다:
people.iloc[2]
# 행을 슬라이싱할 수 있으며 `DataFrame` 객체가 반환됩니다:
people.iloc[1:3]
# 마자믹으로 불리언 배열을 전달하여 해당하는 행을 가져올 수 있습니다:
people[np.array([True, False, True])]
# 불리언 표현식을 사용할 때 아주 유용합니다:
people[people["birthyear"] < 1990]
# ## 열 추가, 삭제
#
# `DataFrame`을 `Series`의 딕셔너리처럼 다룰 수 있습니다. 따라서 다음 같이 쓸 수 있습니다:
people
# +
people["age"] = 2018 - people["birthyear"] # "age" 열을 추가합니다
people["over 30"] = people["age"] > 30 # "over 30" 열을 추가합니다
birthyears = people.pop("birthyear")
del people["children"]
people
# -
birthyears
# 새로운 열을 추가할 때 행의 개수는 같아야 합니다. 누락된 행은 `NaN`으로 채워지고 추가적인 행은 무시됩니다:
people["pets"] = pd.Series({"bob": 0, "charles": 5, "eugene":1}) # alice 누락됨, eugene은 무시됨
people
# 새로운 열을 추가할 때 기본적으로 (오른쪽) 끝에 추가됩니다. `insert()` 메서드를 사용해 다른 곳에 열을 추가할 수 있습니다:
people.insert(1, "height", [172, 181, 185])
people
# ## 새로운 열 할당하기
#
# `assign()` 메서드를 호출하여 새로운 열을 만들 수도 있습니다. 이는 새로운 `DataFrame` 객체를 반환하며 원본 객체는 변경되지 않습니다:
people.assign(
body_mass_index = people["weight"] / (people["height"] / 100) ** 2,
has_pets = people["pets"] > 0
)
# 할당문 안에서 만든 열은 접근할 수 없습니다:
try:
people.assign(
body_mass_index = people["weight"] / (people["height"] / 100) ** 2,
overweight = people["body_mass_index"] > 25
)
except KeyError as e:
print("키 에러:", e)
# 해결책은 두 개의 연속된 할당문으로 나누는 것입니다:
d6 = people.assign(body_mass_index = people["weight"] / (people["height"] / 100) ** 2)
d6.assign(overweight = d6["body_mass_index"] > 25)
# 임시 변수 `d6`를 만들면 불편합니다. `assign()` 메서드를 연결하고 싶겠지만 `people` 객체가 첫 번째 할당문에서 실제로 수정되지 않기 때문에 작동하지 않습니다:
try:
(people
.assign(body_mass_index = people["weight"] / (people["height"] / 100) ** 2)
.assign(overweight = people["body_mass_index"] > 25)
)
except KeyError as e:
print("키 에러:", e)
# 하지만 걱정하지 마세요. 간단한 방법이 있습니다. `assign()` 메서드에 함수(전형적으로 `lambda` 함수)를 전달하면 `DataFrame`을 매개변수로 이 함수를 호출할 것입니다:
(people
.assign(body_mass_index = lambda df: df["weight"] / (df["height"] / 100) ** 2)
.assign(overweight = lambda df: df["body_mass_index"] > 25)
)
# 문제가 해결되었군요!
# ## 표현식 평가
#
# 판다스가 제공하는 뛰어난 기능 하나는 표현식 평가입니다. 이는 `numexpr` 라이브러리에 의존하기 때문에 설치가 되어 있어야 합니다.
people.eval("weight / (height/100) ** 2 > 25")
# 할당 표현식도 지원됩니다. `inplace=True`로 지정하면 수정된 복사본을 만들지 않고 바로 `DataFrame`을 변경합니다:
people.eval("body_mass_index = weight / (height/100) ** 2", inplace=True)
people
# `'@'`를 접두어로 사용하여 지역 변수나 전역 변수를 참조할 수 있습니다:
overweight_threshold = 30
people.eval("overweight = body_mass_index > @overweight_threshold", inplace=True)
people
# ## `DataFrame` 쿼리하기
#
# `query()` 메서드를 사용하면 쿼리 표현식에 기반하여 `DataFrame`을 필터링할 수 있습니다:
people.query("age > 30 and pets == 0")
# ## `DataFrame` 정렬
#
# `sort_index` 메서드를 호출하여 `DataFrame`을 정렬할 수 있습니다. 기본적으로 인덱스 레이블을 기준으로 오름차순으로 행을 정렬합니다. 여기에서는 내림차순으로 정렬해 보죠:
people.sort_index(ascending=False)
# `sort_index`는 `DataFrame`의 정렬된 *복사본*을 반환합니다. `people`을 직접 수정하려면 `inplace` 매개변수를 `True`로 지정합니다. 또한 `axis=1`로 지정하여 열 대신 행을 정렬할 수 있습니다:
people.sort_index(axis=1, inplace=True)
people
# 레이블이 아니라 값을 기준으로 `DataFrame`을 정렬하려면 `sort_values`에 정렬하려는 열을 지정합니다:
people.sort_values(by="age", inplace=True)
people
# ## `DataFrame` 그래프 그리기
#
# `Series`와 마찬가지로 판다스는 `DataFrame` 기반으로 멋진 그래프를 손쉽게 그릴 수 있습니다.
#
# 예를 들어 `plot` 메서드를 호출하여 `DataFrame`의 데이터에서 선 그래프를 쉽게 그릴 수 있습니다:
people.plot(kind = "line", x = "body_mass_index", y = ["height", "weight"])
plt.show()
# 맷플롯립의 함수가 지원하는 다른 매개변수를 사용할 수 있습니다. 예를 들어, 산점도를 그릴 때 맷플롯립의 `scatter()` 함수의 `s` 매개변수를 사용해 크기를 지정할 수 있습니다:
people.plot(kind = "scatter", x = "height", y = "weight", s=[40, 120, 200])
plt.show()
# 선택할 수 있는 옵션이 많습니다. 판다스 문서의 [시각화](http://pandas.pydata.org/pandas-docs/stable/visualization.html) 페이지에서 마음에 드는 그래프를 찾아 예제 코드를 살펴 보세요.
# ## `DataFrame` 연산
#
# `DataFrame`이 넘파이 배열을 흉내내려는 것은 아니지만 몇 가지 비슷한 점이 있습니다. 예제 `DataFrame`을 만들어 보죠:
grades_array = np.array([[8,8,9],[10,9,9],[4, 8, 2], [9, 10, 10]])
grades = pd.DataFrame(grades_array, columns=["sep", "oct", "nov"], index=["alice","bob","charles","darwin"])
grades
# `DataFrame`에 넘파이 수학 함수를 적용하면 모든 값에 이 함수가 적용됩니다:
np.sqrt(grades)
# 비슷하게 `DataFrame`에 하나의 값을 더하면 `DataFrame`의 모든 원소에 이 값이 더해집니다. 이를 *브로드캐스팅*이라고 합니다:
grades + 1
# 물론 산술 연산(`*`,`/`,`**`...)과 조건 연산(`>`, `==`...)을 포함해 모든 이항 연산에도 마찬가지 입니다:
grades >= 5
# `DataFrame`의 `max`, `sum`, `mean` 같은 집계 연산은 각 열에 적용되어 `Series` 객체가 반환됩니다:
grades.mean()
# `all` 메서드도 집계 연산입니다: 모든 값이 `True`인지 아닌지 확인합니다. 모든 학생의 점수가 `5` 이상인 월을 찾아 보죠:
(grades > 5).all()
# Most of these functions take an optional `axis` parameter which lets you specify along which axis of the `DataFrame` you want the operation executed. The default is `axis=0`, meaning that the operation is executed vertically (on each column). You can set `axis=1` to execute the operation horizontally (on each row). For example, let's find out which students had all grades greater than `5`:
(grades > 5).all(axis = 1)
# `any` 메서드는 하나라도 참이면 `True`를 반환합니다. 한 번이라도 10점을 받은 사람을 찾아 보죠:
(grades == 10).any(axis = 1)
# `DataFrame`에 `Series` 객체를 더하면 (또는 다른 이항 연산을 수행하면) 판다스는 `DataFrame`에 있는 모든 *행*에 이 연산을 브로드캐스팅합니다. 이는 `Series` 객체가 `DataFrame`의 행의 개수와 크기가 같을 때만 동작합니다. 예를 들어 `DataFrame`의 `mean`(`Series` 객체)을 빼보죠:
grades - grades.mean() # grades - [7.75, 8.75, 7.50] 와 동일
# 모든 9월 성적에서 `7.75`를 빼고, 10월 성적에서 `8.75`를 빼고, 11월 성적에서 `7.50`을 뺍니다. 이는 다음 `DataFrame`을 빼는 것과 같습니다:
pd.DataFrame([[7.75, 8.75, 7.50]]*4, index=grades.index, columns=grades.columns)
# 모든 성적의 전체 평균을 빼고 싶다면 다음과 같은 방법을 사용합니다:
grades - grades.values.mean() # 모든 점수에서 전체 평균(8.00)을 뺍니다
# ## 자동 정렬
#
# `Series`와 비슷하게 여러 개의 `DataFrame`에 대한 연산을 수행하면 판다스는 자동으로 행 인덱스 레이블로 정렬하지만 열 이름으로도 정렬할 수 있습니다. 10월부터 12월까지 보너스 포인트를 담은 `DataFrame`을 만들어 보겠습니다:
bonus_array = np.array([[0,np.nan,2],[np.nan,1,0],[0, 1, 0], [3, 3, 0]])
bonus_points = pd.DataFrame(bonus_array, columns=["oct", "nov", "dec"], index=["bob","colin", "darwin", "charles"])
bonus_points
grades + bonus_points
# 덧셈 연산이 수행되었지만 너무 많은 원소가 `NaN`이 되었습니다. `DataFrame`을 정렬할 때 일부 열과 행이 한 쪽에만 있기 때문입니다. 다른 쪽에는 누란되었다고 간주합니다(`NaN`). `NaN`에 어떤 수를 더하면 `NaN`이 됩니다.
#
# ## 누락된 데이터 다루기
#
# 실제 데이터에서 누락된 데이터를 다루는 경우는 자주 발생합니다. 판다스는 누락된 데이터를 다룰 수 있는 몇 가지 방법을 제공합니다.
#
# 위 데이터에 있는 문제를 해결해 보죠. 예를 들어, 누락된 데이터는 `NaN`이 아니라 0이 되어야 한다고 결정할 수 있습니다. `fillna()` 메서드를 사용해 모든 `NaN` 값을 어떤 값으로 바꿀 수 있습니다:
(grades + bonus_points).fillna(0)
# 9월의 점수를 0으로 만드는 것은 공정하지 않습니다. 누락된 점수는 그대로 두고, 누락된 보너스 포인트는 0으로 바꿀 수 있습니다:
fixed_bonus_points = bonus_points.fillna(0)
fixed_bonus_points.insert(0, "sep", 0)
fixed_bonus_points.loc["alice"] = 0
grades + fixed_bonus_points
# 훨씬 낫네요: 일부 데이터를 꾸며냈지만 덜 불공정합니다.
#
# 누락된 값을 다루는 또 다른 방법은 보간입니다. `bonus_points` `DataFrame`을 다시 보죠:
bonus_points
# `interpolate` 메서드를 사용해 보죠. 기본적으로 수직 방향(`axis=0`)으로 보간합니다. 따라서 수평으로(`axis=1`)으로 보간하도록 지정합니다.
bonus_points.interpolate(axis=1)
# bob의 보너스 포인트는 10월에 0이고 12월에 2입니다. 11월을 보간하면 평균 보너스 포인트 1을 얻습니다. colin의 보너스 포인트는 11월에 1이지만 9월에 포인트는 얼마인지 모릅니다. 따라서 보간할 수 없고 10월의 포인트는 그대로 누락된 값으로 남아 있습니다. 이를 해결하려면 보간하기 전에 9월의 보너스 포인트를 0으로 설정해야 합니다.
better_bonus_points = bonus_points.copy()
better_bonus_points.insert(0, "sep", 0)
better_bonus_points.loc["alice"] = 0
better_bonus_points = better_bonus_points.interpolate(axis=1)
better_bonus_points
# 좋습니다. 이제 모든 보너스 포인트가 합리적으로 보간되었습니다. 최종 점수를 확인해 보죠:
grades + better_bonus_points
# 9월 열이 오른쪽에 추가되었는데 좀 이상합니다. 이는 더하려는 `DataFrame`이 정확히 같은 열을 가지고 있지 않기 때문입니다(`grade` `DataFrame`에는 `"dec"` 열이 없습니다). 따라서 판다스는 알파벳 순서로 최종 열을 정렬합니다. 이를 해결하려면 덧셈을 하기 전에 누락된 열을 추가하면 됩니다:
grades["dec"] = np.nan
final_grades = grades + better_bonus_points
final_grades
# 12월과 colin에 대해 할 수 있는 것이 많지 않습니다. 보너스 포인트를 만드는 것이 나쁘지만 점수를 합리적으로 올릴 수는 없습니다(어떤 선생님들은 그럴 수 있지만). `dropna()` 메서드를 사용해 모두 `NaN`인 행을 삭제합니다:
final_grades_clean = final_grades.dropna(how="all")
final_grades_clean
# 그다음 `axis` 매개변수를 `1`로 지정하여 모두 `NaN`인 열을 삭제합니다:
final_grades_clean = final_grades_clean.dropna(axis=1, how="all")
final_grades_clean
# ## `groupby`로 집계하기
#
# SQL과 비슷하게 판다스는 데이터를 그룹핑하고 각 그룹에 대해 연산을 수행할 수 있습니다.
#
# 먼저 그루핑을 위해 각 사람의 데이터를 추가로 만들겠습니다. `NaN` 값을 어떻게 다루는지 보기 위해 `final_grades` `DataFrame`을 다시 사용하겠습니다:
final_grades["hobby"] = ["Biking", "Dancing", np.nan, "Dancing", "Biking"]
final_grades
# hobby로 이 `DataFrame`을 그룹핑해 보죠:
grouped_grades = final_grades.groupby("hobby")
grouped_grades
# 이제 hobby마다 평균 점수를 계산할 수 있습니다:
grouped_grades.mean()
# 아주 쉽네요! 평균을 계산할 때 `NaN` 값은 그냥 무시됩니다.
# ## 피봇 테이블
#
# 판다스는 스프레드시트와 비슷하 [피봇 테이블](https://en.wikipedia.org/wiki/Pivot_table)을 지원하여 데이터를 빠르게 요약할 수 있습니다. 어떻게 동작하는 알아 보기 위해 간단한 `DataFrame`을 만들어 보죠:
bonus_points
more_grades = final_grades_clean.stack().reset_index()
more_grades.columns = ["name", "month", "grade"]
more_grades["bonus"] = [np.nan, np.nan, np.nan, 0, np.nan, 2, 3, 3, 0, 0, 1, 0]
more_grades
# 이제 이 `DataFrame`에 대해 `pd.pivot_table()` 함수를 호출하고 `name` 열로 그룹핑합니다. 기본적으로 `pivot_table()`은 수치 열의 평균을 계산합니다:
pd.pivot_table(more_grades, index="name")
# 집계 함수를 `aggfunc` 매개변수로 바꿀 수 있습니다. 또한 집계 대상의 열을 리스트로 지정할 수 있습니다:
pd.pivot_table(more_grades, index="name", values=["grade","bonus"], aggfunc=np.max)
# `columns` 매개변수를 지정하여 수평으로 집계할 수 있고 `margins=True`로 설정해 각 행과 열에 대해 전체 합을 계산할 수 있습니다:
pd.pivot_table(more_grades, index="name", values="grade", columns="month", margins=True)
# 마지막으로 여러 개의 인덱스나 열 이름을 지정하면 판다스가 다중 레벨 인덱스를 만듭니다:
pd.pivot_table(more_grades, index=("name", "month"), margins=True)
# ## 함수
#
# 큰 `DataFrame`을 다룰 때 내용을 간단히 요약하는 것이 도움이 됩니다. 판다스는 이를 위한 몇 가지 함수를 제공합니다. 먼저 수치 값, 누락된 값, 텍스트 값이 섞인 큰 `DataFrame`을 만들어 보죠. 주피터 노트북은 이 `DataFrame`의 일부만 보여줍니다:
much_data = np.fromfunction(lambda x,y: (x+y*y)%17*11, (10000, 26))
large_df = pd.DataFrame(much_data, columns=list("ABCDEFGHIJKLMNOPQRSTUVWXYZ"))
large_df[large_df % 16 == 0] = np.nan
large_df.insert(3,"some_text", "Blabla")
large_df
# `head()` 메서드는 처음 5개 행을 반환합니다:
large_df.head()
# 마지막 5개 행을 반환하는 `tail()` 함수도 있습니다. 원하는 행 개수를 전달할 수도 있습니다:
large_df.tail(n=2)
# `info()` 메서드는 각 열의 내용을 요약하여 출력합니다:
large_df.info()
# 마지막으로 `describe()` 메서드는 각 열에 대한 주요 집계 연산을 수행한 결과를 보여줍니다:
#
# Finally, the `describe()` method gives a nice overview of the main aggregated values over each column:
# * `count`: null(NaN)이 아닌 값의 개수
# * `mean`: null이 아닌 값의 평균
# * `std`: null이 아닌 값의 [표준 편차](https://ko.wikipedia.org/wiki/%ED%91%9C%EC%A4%80_%ED%8E%B8%EC%B0%A8)
# * `min`: null이 아닌 값의 최솟값
# * `25%`, `50%`, `75%`: null이 아닌 값의 25번째, 50번째, 75번째 [백분위수](https://ko.wikipedia.org/wiki/%EB%B0%B1%EB%B6%84%EC%9C%84%EC%88%98)
# * `max`: null이 아닌 값의 최댓값
large_df.describe()
# # 저장 & 로딩
#
# 판다스는 `DataFrame`를 여러 가지 포맷으로 저장할 수 있습니다. CSV, Excel, JSON, HTML, HDF5, SQL 데이터베이스 같은 포맷이 가능합니다. 예제를 위해 `DataFrame`을 하나 만들어 보겠습니다:
my_df = pd.DataFrame(
[["Biking", 68.5, 1985, np.nan], ["Dancing", 83.1, 1984, 3]],
columns=["hobby","weight","birthyear","children"],
index=["alice", "bob"]
)
my_df
# ## 저장
#
# CSV, HTML, JSON로 저장해 보죠:
my_df.to_csv("my_df.csv")
my_df.to_html("my_df.html")
my_df.to_json("my_df.json")
# 저장된 내용을 확인해 보죠:
for filename in ("my_df.csv", "my_df.html", "my_df.json"):
print("#", filename)
with open(filename, "rt") as f:
print(f.read())
print()
# 인덱스는 (이름 없이) CSV 파일의 첫 번째 열에 저장되었습니다. HTML에서는 `<th>` 태그와 JSON에서는 키로 저장되었습니다.
#
# 다른 포맷으로 저장하는 것도 비슷합니다. 하지만 일부 포맷은 추가적인 라이브러리 설치가 필요합니다. 예를 들어, 엑셀로 저장하려면 openpyxl 라이브러리가 필요합니다:
try:
my_df.to_excel("my_df.xlsx", sheet_name='People')
except ImportError as e:
print(e)
# ## 로딩
#
# CSV 파일을 `DataFrame`으로 로드해 보죠:
my_df_loaded = pd.read_csv("my_df.csv", index_col=0)
my_df_loaded
# 예상할 수 있듯이 `read_json`, `read_html`, `read_excel` 함수도 있습니다. 인터넷에서 데이터를 바로 읽을 수도 있습니다. 예를 들어 [simplemaps.com](http://simplemaps.com/)에서 U.S. 도시를 로드해 보죠:
us_cities = None
try:
csv_url = "http://simplemaps.com/files/cities.csv"
us_cities = pd.read_csv(csv_url, index_col=0)
us_cities = us_cities.head()
except IOError as e:
print(e)
us_cities
# 이외에도 많은 옵션이 있습니다. 특히 datetime 포맷에 관련된 옵션이 많습니다. 더 자세한 내용은 온라인 [문서](http://pandas.pydata.org/pandas-docs/stable/io.html)를 참고하세요.
# # `DataFrame` 합치기
#
# ## SQL 조인
#
# 판다스의 강력한 기능 중 하나는 `DataFrame`에 대해 SQL 같은 조인(join)을 수행할 수 있는 것입니다. 여러 종류의 조인이 지원됩니다. 이너 조인(inner join), 레프트/라이트 아우터 조인(left/right outer join), 풀 조인(full join)입니다. 이에 대해 알아 보기 위해 간단한 `DataFrame`을 만들어 보죠:
city_loc = pd.DataFrame(
[
["CA", "San Francisco", 37.781334, -122.416728],
["NY", "New York", 40.705649, -74.008344],
["FL", "Miami", 25.791100, -80.320733],
["OH", "Cleveland", 41.473508, -81.739791],
["UT", "Salt Lake City", 40.755851, -111.896657]
], columns=["state", "city", "lat", "lng"])
city_loc
city_pop = pd.DataFrame(
[
[808976, "San Francisco", "California"],
[8363710, "New York", "New-York"],
[413201, "Miami", "Florida"],
[2242193, "Houston", "Texas"]
], index=[3,4,5,6], columns=["population", "city", "state"])
city_pop
# 이제 `merge()` 함수를 사용해 이 `DataFrame`을 조인해 보죠:
pd.merge(left=city_loc, right=city_pop, on="city")
# 두 `DataFrame`은 `state`란 이름의 열을 가지고 있으므로 `state_x`와 `state_y`로 이름이 바뀌었습니다.
#
# 또한 Cleveland, Salt Lake City, Houston은 두 `DataFrame`에 모두 존재하지 않기 때문에 삭제되었습니다. SQL의 `INNER JOIN`과 동일합니다. 도시를 삭제하지 않고 `NaN`으로 채우는 `FULL OUTER JOIN`을 원하면 `how="outer"`로 지정합니다:
all_cities = pd.merge(left=city_loc, right=city_pop, on="city", how="outer")
all_cities
# 물론 `LEFT OUTER JOIN`은 `how="left"`로 지정할 수 있습니다. 왼쪽의 `DataFrame`에 있는 도시만 남습니다. 비슷하게 `how="right"`는 오른쪽 `DataFrame`에 있는 도시만 결과에 남습니다. 예를 들면:
pd.merge(left=city_loc, right=city_pop, on="city", how="right")
# 조인할 키가 `DataFrame` 인덱스라면 `left_index=True`나 `right_index=True`로 지정해야 합니다. 키 열의 이름이 다르면 `left_on`과 `right_on`을 사용합니다. 예를 들어:
city_pop2 = city_pop.copy()
city_pop2.columns = ["population", "name", "state"]
pd.merge(left=city_loc, right=city_pop2, left_on="city", right_on="name")
# ## 연결
#
# `DataFrame`을 조인하는 대신 그냥 연결할 수도 있습니다. `concat()` 함수가 하는 일입니다:
result_concat = pd.concat([city_loc, city_pop])
result_concat
# 이 연산은 (행을 따라) 수직적으로 데이터를 연결하고 (열을 따라) 수평으로 연결하지 않습니다. 이 예에서 동일한 인덱스를 가진 행이 있습니다(예를 들면 3). 판다스는 이를 우아하게 처리합니다:
result_concat.loc[3]
# 또는 인덱스를 무시하도록 설정할 수 있습니다:
pd.concat([city_loc, city_pop], ignore_index=True)
# 한 `DataFrame`에 열이 없을 때 `NaN`이 채워져 있는 것처럼 동작합니다. `join="inner"`로 설정하면 *양쪽*의 `DataFrame`에 존재하는 열만 반환됩니다:
pd.concat([city_loc, city_pop], join="inner")
# `axis=1`로 설정하면 `DataFrame`을 수직이 아니라 수평으로 연결할 수 있습니다:
pd.concat([city_loc, city_pop], axis=1)
# 이 경우 인덱스가 잘 정렬되지 않기 때문에 의미가 없습니다(예를 들어 Cleveland와 San Francisco의 인덱스 레이블이 `3`이기 때문에 동일한 행에 놓여 있습니다). 이 `DataFrame`을 연결하기 전에 도시로 인덱스를 재설정해 보죠:
pd.concat([city_loc.set_index("city"), city_pop.set_index("city")], axis=1)
# `FULL OUTER JOIN`을 수행한 것과 비슷합니다. 하지만 `state` 열이 `state_x`와 `state_y`로 바뀌지 않았고 `city` 열이 인덱스가 되었습니다.
# `append()` 메서드는 `DataFrame`을 수직으로 연결하는 단축 메서드입니다:
city_loc.append(city_pop)
# 판다스의 다른 메서드와 마찬가지로 `append()` 메서드는 실제 `city_loc`을 수정하지 않습니다. 복사본을 만들어 수정한 다음 반환합니다.
# # 범주
#
# 범주로 표현된 값을 가진 경우가 흔합니다. 예를 들어 `1`은 여성, `2`는 남성이거나 `"A"`는 좋은 것, `"B"`는 평균, `"C"`는 나쁜 것 등입니다. 범주형 값을 읽기 힘들고 다루기 번거롭습니다. 하지만 판다스에서는 간단합니다. 예를 들기 위해 앞서 만든 `city_pop` `DataFrame`에 범주를 표현하는 열을 추가해 보겠습니다:
city_eco = city_pop.copy()
city_eco["eco_code"] = [17, 17, 34, 20]
city_eco
# 이제 `eco_code`열은 의미없는 코드입니다. 이를 바꿔 보죠. 먼저 `eco_code`를 기반으로 새로운 범주형 열을 만듭니다:
city_eco["economy"] = city_eco["eco_code"].astype('category')
city_eco["economy"].cat.categories
# 의미있는 이름을 가진 범주를 지정할 수 있습니다:
city_eco["economy"].cat.categories = ["Finance", "Energy", "Tourism"]
city_eco
# 범주형 값은 알파벳 순서가 아니라 범주형 순서로 정렬합니다:
city_eco.sort_values(by="economy", ascending=False)
# # 그 다음엔?
#
# 이제 알았겠지만 판다스는 매우 커다란 라이브러리이고 기능이 많습니다. 가장 중요한 기능들을 둘러 보았지만 빙산의 일각일 뿐입니다. 더 많은 것을 익히려면 실전 데이터로 직접 실습해 보는 것이 제일 좋습니다. 판다스의 훌륭한 [문서](http://pandas.pydata.org/pandas-docs/stable/index.html)와 [쿡북](http://pandas.pydata.org/pandas-docs/stable/cookbook.html)을 보는 것도 좋습니다.
| tools_pandas.ipynb |
# +
# Copyright 2010-2018 Google LLC
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""MaxFlow and MinCostFlow examples."""
from __future__ import print_function
from ortools.graph import pywrapgraph
def MaxFlow():
"""MaxFlow simple interface example."""
print('MaxFlow on a simple network.')
tails = [0, 0, 0, 0, 1, 2, 3, 3, 4]
heads = [1, 2, 3, 4, 3, 4, 4, 5, 5]
capacities = [5, 8, 5, 3, 4, 5, 6, 6, 4]
expected_total_flow = 10
max_flow = pywrapgraph.SimpleMaxFlow()
for i in range(0, len(tails)):
max_flow.AddArcWithCapacity(tails[i], heads[i], capacities[i])
if max_flow.Solve(0, 5) == max_flow.OPTIMAL:
print('Total flow', max_flow.OptimalFlow(), '/', expected_total_flow)
for i in range(max_flow.NumArcs()):
print(('From source %d to target %d: %d / %d' %
(max_flow.Tail(i), max_flow.Head(i), max_flow.Flow(i),
max_flow.Capacity(i))))
print('Source side min-cut:', max_flow.GetSourceSideMinCut())
print('Sink side min-cut:', max_flow.GetSinkSideMinCut())
else:
print('There was an issue with the max flow input.')
def MinCostFlow():
"""MinCostFlow simple interface example.
Note that this example is actually a linear sum assignment example and will
be more efficiently solved with the pywrapgraph.LinearSumAssignement class.
"""
print('MinCostFlow on 4x4 matrix.')
num_sources = 4
num_targets = 4
costs = [[90, 75, 75, 80], [35, 85, 55, 65], [125, 95, 90, 105],
[45, 110, 95, 115]]
expected_cost = 275
min_cost_flow = pywrapgraph.SimpleMinCostFlow()
for source in range(0, num_sources):
for target in range(0, num_targets):
min_cost_flow.AddArcWithCapacityAndUnitCost(
source, num_sources + target, 1, costs[source][target])
for node in range(0, num_sources):
min_cost_flow.SetNodeSupply(node, 1)
min_cost_flow.SetNodeSupply(num_sources + node, -1)
status = min_cost_flow.Solve()
if status == min_cost_flow.OPTIMAL:
print('Total flow', min_cost_flow.OptimalCost(), '/', expected_cost)
for i in range(0, min_cost_flow.NumArcs()):
if min_cost_flow.Flow(i) > 0:
print('From source %d to target %d: cost %d' %
(min_cost_flow.Tail(i),
min_cost_flow.Head(i) - num_sources,
min_cost_flow.UnitCost(i)))
else:
print('There was an issue with the min cost flow input.')
MaxFlow()
MinCostFlow()
| examples/notebook/examples/pyflow_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.2 ('scikit-learn-course')
# language: python
# name: python3
# ---
# +
import pandas as pd
ames_housing = pd.read_csv("../datasets/ames_housing_no_missing.csv")
target_name = "SalePrice"
data, target = ames_housing.drop(columns=target_name), ames_housing[target_name]
target = (target > 200_000).astype(int)
| notebooks/warpup_quiz_01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# %load_ext autoreload
# %autoreload 2
# +
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# +
# one document in the corpus is a book
from adhtools.utils import corpus_wordlist
from nlppln.utils import get_files
#xml_file1 = '/home/jvdzwaan/data/tmp/adh/chapters/1266MuhammadHasanNajafiJawhari.JawahirKalam.xml'
#xml_file2 = '/home/jvdzwaan/data/tmp/adh/chapters/0381IbnBabawayh.Hidaya.xml'
in_dir = '/home/jvdzwaan/Downloads/2019-02-08-fiqh-newfiles-alkhalil/'
in_files = get_files(in_dir)
c = corpus_wordlist(in_files, analyzer=True, field='word')
# -
# %%time
from tqdm import tqdm_notebook as tqdm
data = [list(terms) for terms in tqdm(c, total=len(in_files))]
corpus_file = 'fiqh-works-alkhalil.pkl'
# +
# %%time
import pickle
with open(corpus_file, 'wb') as f:
pickle.dump(data, f)
# +
# %%time
import pickle
with open(corpus_file, 'rb') as f:
data = pickle.load(f)
# -
#calculate pwc per book
df = calculate(data, w=0.001, top_k=200)
df
# +
# selected works:
#0483 Sarakhsī, Mabsūṭ
#0450 Māwardī, Ḥāwī
#0684 Qarāfi, Dhakhīra
#0884 Ibn Mufliḥ, Mubdiʿ
#0620 Ibn Qudāma, Mughnī
#0460 Ṭūsī, Mabsūṭ
#0676 Ḥillī, Sharāʾiʿ
import os
indices = {}
for i, in_file in enumerate(in_files):
bn = os.path.basename(in_file)
for num in ('0483', '0450', '0684', '0884', '0620', '0460', '0676'):
if bn.startswith(num):
print(i, os.path.splitext(bn)[0])
indices[i] = os.path.splitext(bn)[0]
# -
indices
del(indices[21])
indices
# +
# select columns
cols1 = ['{}_p'.format(k) for k in indices.keys()]
cols2 = ['{}_term'.format(k) for k in indices.keys()]
print(cols1)
cols = []
for c1, c2 in zip(cols1, cols2):
cols.append(c1)
cols.append(c2)
print(cols)
df[cols]
# +
# rename columns
cols1 = ['{}_p'.format(v) for k, v in indices.items()]
cols2 = ['{}_term'.format(v) for k, v in indices.items()]
name_cols = []
for c1, c2 in zip(cols1, cols2):
name_cols.append(c1)
name_cols.append(c2)
print(name_cols)
df = df[cols]
df.columns = name_cols
df
# -
df.to_csv('pwc-0.001-fiqh-works.csv')
# +
# %%time
# one document in the corpus is a school
import os
import codecs
import glob
from itertools import chain
import pandas as pd
from nlppln.utils import get_files
from adhtools.utils import read_file_analyzer, read_file_stemmer
#xml_file1 = '/home/jvdzwaan/data/tmp/adh/chapters/1266MuhammadHasanNajafiJawhari.JawahirKalam.xml'
#xml_file2 = '/home/jvdzwaan/data/tmp/adh/chapters/0381IbnBabawayh.Hidaya.xml'
md_file = '/home/jvdzwaan/data/adh-corpora/fiqh_corpus/Meta/Metadata_Fiqh.csv'
in_dir = '/home/jvdzwaan/data/tmp/adh/2019-02-08-fiqh-newfiles-alkhalil-chapters/'
metadata = pd.read_csv(md_file, encoding='utf-8', sep=';|,')
#print(metadata.head())
schools = metadata.groupby('BookSUBJ')
def corpus(schools, in_dir, analyzer=True):
for i, (name, data) in enumerate(schools):
print(i, name)
#print(data['BookURI'])
words = []
#with codecs.open('{}.txt'.format(i), 'w', encoding='utf-8') as f:
for book in data['BookURI']:
#print(book)
#in_file = os.path.join(in_dir, '{}.xml'.format(book))
in_files = glob.glob('{}/{}*.xml'.format(in_dir, book))
#print(in_files)
for in_file in in_files:
if analyzer:
ws = read_file_analyzer(in_file)
else:
ws = read_file_stemmer(in_file)
#print(ws[0])
words.append(ws)
yield(chain(*words))
#print(len(ws))
#print(ws[0])
#f.write(' '.join(ws))
#f.write('\n')
c = corpus(schools, in_dir, analyzer=True)
# +
# %%time
# make a sample for tuning the parameter. We use one book per school.
import os
import codecs
import glob
from itertools import chain
from collections import defaultdict
import pandas as pd
from nlppln.utils import get_files
from adhtools.utils import read_file_analyzer, read_file_stemmer
#xml_file1 = '/home/jvdzwaan/data/tmp/adh/chapters/1266MuhammadHasanNajafiJawhari.JawahirKalam.xml'
#xml_file2 = '/home/jvdzwaan/data/tmp/adh/chapters/0381IbnBabawayh.Hidaya.xml'
md_file = '/home/jvdzwaan/data/adh-corpora/fiqh_corpus/Meta/Metadata_Fiqh.csv'
in_dir = '/home/jvdzwaan/data/tmp/adh/2019-02-08-fiqh-newfiles-alkhalil-chapters/'
metadata = pd.read_csv(md_file, encoding='utf-8')
#print(metadata.head())
schools = metadata.groupby('BookSUBJ')
for i, (name, data) in enumerate(schools):
# we use the oldest book for every school
print(name)
#print(data)
for book in data.iterrows():
# and sample 30.000 words
b = book[1]
#print(book[1])
print(b['BookURI'], b['Number_of_tokens'])
in_files = glob.glob('{}/{}*.xml'.format(in_dir, book))
#print(in_files)
counts = defaultdict(int)
for in_file in in_files:
if analyzer:
ws = read_file_analyzer(in_file)
else:
ws = read_file_stemmer(in_file)
for word in ws:
counts[word] += 1
break
# +
# %%time
# make a sample for tuning the parameter. We use one book per school.
import os
import codecs
import glob
from itertools import chain
import pandas as pd
from nlppln.utils import get_files
from adhtools.utils import read_file_analyzer, read_file_stemmer
#xml_file1 = '/home/jvdzwaan/data/tmp/adh/chapters/1266MuhammadHasanNajafiJawhari.JawahirKalam.xml'
#xml_file2 = '/home/jvdzwaan/data/tmp/adh/chapters/0381IbnBabawayh.Hidaya.xml'
md_file = '/home/jvdzwaan/data/adh-corpora/fiqh_corpus/Meta/Metadata_Fiqh.csv'
in_dir = '/home/jvdzwaan/data/tmp/adh/2019-02-08-fiqh-newfiles-alkhalil-chapters/'
metadata = pd.read_csv(md_file, encoding='utf-8')
#print(metadata.head())
schools = metadata.groupby('BookSUBJ')
def corpus(schools, in_dir, analyzer=True):
for i, (name, data) in enumerate(schools):
print(i, name)
#print(data['BookURI'])
words = []
#with codecs.open('{}.txt'.format(i), 'w', encoding='utf-8') as f:
for book in data['BookURI']:
#print(book)
#in_file = os.path.join(in_dir, '{}.xml'.format(book))
in_files = glob.glob('{}/{}*.xml'.format(in_dir, book))
#print(in_files)
for in_file in in_files:
if analyzer:
ws = read_file_analyzer(in_file)
else:
ws = read_file_stemmer(in_file)
#print(ws[0])
words.append(ws)
yield(chain(*words))
#print(len(ws))
#print(ws[0])
#f.write(' '.join(ws))
#f.write('\n')
c = corpus(schools, in_dir, analyzer=True)
# -
words = read_file('/home/jvdzwaan/data/tmp/adh/analysis/alkhalil/0311AbuBakrKhallal.WuqufWaTarajjul.xml')
print(len(words))
def read_text_file(in_file):
with codecs.open(in_file, encoding='utf-8') as f:
for ln in f:
for word in ln.split():
#print(word)
yield word
# %%time
data = [list(terms) for terms in c]
corpus_file = 'fiqh-schools-alkhalil.pkl'
# +
# %%time
import pickle
with open(corpus_file, 'wb') as f:
pickle.dump(data, f)
# +
# %%time
import pickle
with open(corpus_file, 'rb') as f:
data = pickle.load(f)
# -
for terms in data:
print(len(terms))
print(data[0][0])
# +
# %%time
from weighwords import ParsimoniousLM
model = ParsimoniousLM([terms for terms in data], w=.01)
# +
# %%time
dfs = []
top_k = 25
for i, terms in enumerate(data):
result = []
for term, p in model.top(top_k, terms, max_iter=100):
result.append({'{}_term'.format(i): term, '{}_p'.format(i): np.exp(p)})
#print(" %s %.4f" % (term, np.exp(p)))
#print(term)
dfs.append(pd.DataFrame(result))
#print('---')
# -
pd.concat(dfs, axis=1, sort=False)
# +
from weighwords import ParsimoniousLM
def calculate(data, w, top_k=25):
model = ParsimoniousLM([terms for terms in data], w=w)
# calculate terms and weights
dfs = []
for i, terms in enumerate(data):
result = []
for term, p in model.top(top_k, terms, max_iter=100):
result.append({'{}_term'.format(i): term, '{}_p'.format(i): np.exp(p)})
dfs.append(pd.DataFrame(result))
return pd.concat(dfs, axis=1, sort=False)
# +
import logging
logger = logging.getLogger(__name__)
logging.basicConfig(level=logging.INFO)
# +
# %%time
wordcloud_data = []
weights = (1.0, 0.5, 0.1, 0.05, 0.01, 0.005, 0.001, 0.0005, 0.0001)
for w in weights:
wordcloud_data.append(calculate(data, w=w))
# +
# %%time
# pwc6 (selected by Christian) for schools
df = calculate(data, w=0.001, top_k=200)
# +
label_0 = 'حنبلي'
label_1 = 'حنفي'
label_2 = 'شافعي'
label_3 = 'شيعي'
label_4 = 'مالكي'
labels = [label_0, label_1, label_2, label_3, label_4]
# -
df.columns = ['{}_p'.format(label_0), '{}_term'.format(label_0),
'{}_p'.format(label_1), '{}_term'.format(label_1),
'{}_p'.format(label_2), '{}_term'.format(label_2),
'{}_p'.format(label_3), '{}_term'.format(label_3),
'{}_p'.format(label_4), '{}_term'.format(label_4)]
df
df.to_csv('pwc-0.001-fiqh-schools.csv')
def get_terms(txt_file):
# get the terms list
terms = pd.read_csv(txt_file, encoding='utf-8', index_col=None, header=None)
t = terms[0].tolist()
print('total number of terms:', len(t))
terms = set(t)
print('number of unique terms:', len(terms))
return terms
stopwords = get_terms('/home/jvdzwaan/data/adh/stopwords/custom.txt')
def sw(term):
return 'background-color: yellow' if term in stopwords else ''
print(len(wordcloud_data))
wordcloud_data[0].style.applymap(sw)
wordcloud_data[1].style.applymap(sw)
wordcloud_data[2].style.applymap(sw)
wordcloud_data[3].style.applymap(sw)
wordcloud_data[4].style.applymap(sw)
wordcloud_data[5].style.applymap(sw)
wordcloud_data[6].style.applymap(sw)
wordcloud_data[7].style.applymap(sw)
wordcloud_data[8].style.applymap(sw)
for i, w in enumerate(weights):
wordcloud_data[i].style.applymap(sw).to_excel('pwc{}.xls'.format(i), engine='openpyxl')
print('\n'.join(list(wordcloud_data[5]['0_term'])))
# +
# %%time
import codecs
c_from_text = [read_text_file(t) for t in ('0.txt', '1.txt', '2.txt', '3.txt', '4.txt')]
[len(list(terms)) for terms in c_from_text]
# +
import os
top_k = 20
words = {}
for fname, doc in zip(in_files, terms):
print("Top %d words in %s:" % (top_k, os.path.basename(fname)))
words[os.path.basename(fname)] = {}
for term, p in model.top(top_k, doc):
print(" %s %.4f" % (term, np.exp(p)))
words[os.path.basename(fname)][term] = np.exp(p)
print("")
# +
# Boek dat <NAME>
# 0620IbnQudamaMaqdisi.MughniFiFiqh.xml
# -
print(len(terms))
# +
from wordcloud import WordCloud
wc = WordCloud(background_color="white", font_path='/usr/share/fonts/opentype/fonts-hosny-amiri/amiri-quran.ttf')
# generate word cloud
wc.generate_from_frequencies(words['0179MalikIbnAnas.Muwatta.xml'])
# show
plt.imshow(wc, interpolation="bilinear")
plt.axis("off")
plt.show()
# +
wc.generate_from_frequencies(words['0483IbnAhmadSarakhsi.Mabsut.xml'])
# show
plt.imshow(wc, interpolation="bilinear")
plt.axis("off")
plt.show()
# -
for word, w in words['0483IbnAhmadSarakhsi.Mabsut.xml'].items():
print(word, w)
| notebooks/parsimonious-wordclouds.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Continuous training pipeline with Kubeflow Pipeline and AI Platform
# **Learning Objectives:**
# 1. Learn how to use Kubeflow Pipeline (KFP) pre-build components (BiqQuery, AI Platform training and predictions)
# 1. Learn how to use KFP lightweight python components
# 1. Learn how to build a KFP with these components
# 1. Learn how to compile, upload, and run a KFP with the command line
#
#
# In this lab, you will build, deploy, and run a KFP pipeline that orchestrates **BigQuery** and **AI Platform** services to train, tune, and deploy a **scikit-learn** model.
#
# ## Understanding the pipeline design
#
# The workflow implemented by the pipeline is defined using a Python based Domain Specific Language (DSL). The pipeline's DSL is in the `covertype_training_pipeline.py` file that we will generate below.
#
# The pipeline's DSL has been designed to avoid hardcoding any environment specific settings like file paths or connection strings. These settings are provided to the pipeline code through a set of environment variables.
#
#
#
# !grep 'BASE_IMAGE =' -A 5 pipeline/covertype_training_pipeline.py
# The pipeline uses a mix of custom and pre-build components.
#
# - Pre-build components. The pipeline uses the following pre-build components that are included with the KFP distribution:
# - [BigQuery query component](https://github.com/kubeflow/pipelines/tree/0.2.5/components/gcp/bigquery/query)
# - [AI Platform Training component](https://github.com/kubeflow/pipelines/tree/0.2.5/components/gcp/ml_engine/train)
# - [AI Platform Deploy component](https://github.com/kubeflow/pipelines/tree/0.2.5/components/gcp/ml_engine/deploy)
# - Custom components. The pipeline uses two custom helper components that encapsulate functionality not available in any of the pre-build components. The components are implemented using the KFP SDK's [Lightweight Python Components](https://www.kubeflow.org/docs/pipelines/sdk/lightweight-python-components/) mechanism. The code for the components is in the `helper_components.py` file:
# - **Retrieve Best Run**. This component retrieves a tuning metric and hyperparameter values for the best run of a AI Platform Training hyperparameter tuning job.
# - **Evaluate Model**. This component evaluates a *sklearn* trained model using a provided metric and a testing dataset.
#
# ### Exercise
#
# Complete TO DOs the pipeline file below.
#
# <ql-infobox><b>NOTE:</b> If you need help, you may take a look at the complete solution by navigating to **mlops-on-gcp > workshops > kfp-caip-sklearn > lab-02-kfp-pipeline** and opening **lab-02.ipynb**.
# </ql-infobox>
# +
# %%writefile ./pipeline/covertype_training_pipeline.py
# Copyright 2019 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""KFP orchestrating BigQuery and Cloud AI Platform services."""
import os
from helper_components import evaluate_model
from helper_components import retrieve_best_run
from jinja2 import Template
import kfp
from kfp.components import func_to_container_op
from kfp.dsl.types import Dict
from kfp.dsl.types import GCPProjectID
from kfp.dsl.types import GCPRegion
from kfp.dsl.types import GCSPath
from kfp.dsl.types import String
from kfp.gcp import use_gcp_secret
# Defaults and environment settings
BASE_IMAGE = os.getenv('BASE_IMAGE')
TRAINER_IMAGE = os.getenv('TRAINER_IMAGE')
RUNTIME_VERSION = os.getenv('RUNTIME_VERSION')
PYTHON_VERSION = os.getenv('PYTHON_VERSION')
COMPONENT_URL_SEARCH_PREFIX = os.getenv('COMPONENT_URL_SEARCH_PREFIX')
USE_KFP_SA = os.getenv('USE_KFP_SA')
TRAINING_FILE_PATH = 'datasets/training/data.csv'
VALIDATION_FILE_PATH = 'datasets/validation/data.csv'
TESTING_FILE_PATH = 'datasets/testing/data.csv'
# Parameter defaults
SPLITS_DATASET_ID = 'splits'
HYPERTUNE_SETTINGS = """
{
"hyperparameters": {
"goal": "MAXIMIZE",
"maxTrials": 6,
"maxParallelTrials": 3,
"hyperparameterMetricTag": "accuracy",
"enableTrialEarlyStopping": True,
"params": [
{
"parameterName": "max_iter",
"type": "DISCRETE",
"discreteValues": [500, 1000]
},
{
"parameterName": "alpha",
"type": "DOUBLE",
"minValue": 0.0001,
"maxValue": 0.001,
"scaleType": "UNIT_LINEAR_SCALE"
}
]
}
}
"""
# Helper functions
def generate_sampling_query(source_table_name, num_lots, lots):
"""Prepares the data sampling query."""
sampling_query_template = """
SELECT *
FROM
`{{ source_table }}` AS cover
WHERE
MOD(ABS(FARM_FINGERPRINT(TO_JSON_STRING(cover))), {{ num_lots }}) IN ({{ lots }})
"""
query = Template(sampling_query_template).render(
source_table=source_table_name, num_lots=num_lots, lots=str(lots)[1:-1])
return query
# Create component factories
component_store = kfp.components.ComponentStore(
local_search_paths=None, url_search_prefixes=[COMPONENT_URL_SEARCH_PREFIX])
bigquery_query_op = component_store.load_component('bigquery/query')
mlengine_train_op = component_store.load_component('ml_engine/train')
mlengine_deploy_op = component_store.load_component('ml_engine/deploy')
retrieve_best_run_op = func_to_container_op(
retrieve_best_run, base_image=BASE_IMAGE)
evaluate_model_op = func_to_container_op(evaluate_model, base_image=BASE_IMAGE)
@kfp.dsl.pipeline(
name='Covertype Classifier Training',
description='The pipeline training and deploying the Covertype classifierpipeline_yaml'
)
def covertype_train(project_id,
region,
source_table_name,
gcs_root,
dataset_id,
evaluation_metric_name,
evaluation_metric_threshold,
model_id,
version_id,
replace_existing_version,
hypertune_settings=HYPERTUNE_SETTINGS,
dataset_location='US'):
"""Orchestrates training and deployment of an sklearn model."""
# Create the training split
query = generate_sampling_query(
source_table_name=source_table_name, num_lots=10, lots=[1, 2, 3, 4])
training_file_path = '{}/{}'.format(gcs_root, TRAINING_FILE_PATH)
create_training_split = bigquery_query_op(
query=query,
project_id=project_id,
dataset_id=dataset_id,
table_id='',
output_gcs_path=training_file_path,
dataset_location=dataset_location)
# Create the validation split
query = generate_sampling_query(
source_table_name=source_table_name, num_lots=10, lots=[8])
validation_file_path = '{}/{}'.format(gcs_root, VALIDATION_FILE_PATH)
create_validation_split = bigquery_query_op(
query=query,
project_id=project_id,
dataset_id=dataset_id,
table_id='',
output_gcs_path=validation_file_path,
dataset_location=dataset_location)
# Create the testing split
query = generate_sampling_query(
source_table_name=source_table_name, num_lots=10, lots=[9])
testing_file_path = '{}/{}'.format(gcs_root, TESTING_FILE_PATH)
create_testing_split = bigquery_query_op(
query=query,
project_id=project_id,
dataset_id=dataset_id,
table_id='',
output_gcs_path=testing_file_path,
dataset_location=dataset_location)
# Tune hyperparameters
tune_args = [
'--training_dataset_path',
create_training_split.outputs['output_gcs_path'],
'--validation_dataset_path',
create_validation_split.outputs['output_gcs_path'], '--hptune', 'True'
]
job_dir = '{}/{}/{}'.format(gcs_root, 'jobdir/hypertune',
kfp.dsl.RUN_ID_PLACEHOLDER)
hypertune = mlengine_train_op(
project_id=project_id,
region=region,
master_image_uri=TRAINER_IMAGE,
job_dir=job_dir,
args=tune_args,
training_input=hypertune_settings)
# Retrieve the best trial
get_best_trial = retrieve_best_run_op(
project_id, hypertune.outputs['job_id'])
# Train the model on a combined training and validation datasets
job_dir = '{}/{}/{}'.format(gcs_root, 'jobdir', kfp.dsl.RUN_ID_PLACEHOLDER)
train_args = [
'--training_dataset_path',
create_training_split.outputs['output_gcs_path'],
'--validation_dataset_path',
create_validation_split.outputs['output_gcs_path'], '--alpha',
get_best_trial.outputs['alpha'], '--max_iter',
get_best_trial.outputs['max_iter'], '--hptune', 'False'
]
train_model = mlengine_train_op(
project_id=project_id,
region=region,
master_image_uri=TRAINER_IMAGE,
job_dir=job_dir,
args=train_args)
# Evaluate the model on the testing split
eval_model = evaluate_model_op(
dataset_path=str(create_testing_split.outputs['output_gcs_path']),
model_path=str(train_model.outputs['job_dir']),
metric_name=evaluation_metric_name)
# Deploy the model if the primary metric is better than threshold
with kfp.dsl.Condition(eval_model.outputs['metric_value'] > evaluation_metric_threshold):
deploy_model = mlengine_deploy_op(
model_uri=train_model.outputs['job_dir'],
project_id=project_id,
model_id=model_id,
version_id=version_id,
runtime_version=RUNTIME_VERSION,
python_version=PYTHON_VERSION,
replace_existing_version=replace_existing_version)
# Configure the pipeline to run using the service account defined
# in the user-gcp-sa k8s secret
if USE_KFP_SA == 'True':
kfp.dsl.get_pipeline_conf().add_op_transformer(
use_gcp_secret('user-gcp-sa'))
# -
# The custom components execute in a container image defined in `base_image/Dockerfile`.
# !cat base_image/Dockerfile
# The training step in the pipeline employes the AI Platform Training component to schedule a AI Platform Training job in a custom training container. The custom training image is defined in `trainer_image/Dockerfile`.
# !cat trainer_image/Dockerfile
# ## Building and deploying the pipeline
#
# Before deploying to AI Platform Pipelines, the pipeline DSL has to be compiled into a pipeline runtime format, also refered to as a pipeline package. The runtime format is based on [Argo Workflow](https://github.com/argoproj/argo), which is expressed in YAML.
#
# ### Configure environment settings
#
# Update the below constants with the settings reflecting your lab environment.
#
# - `REGION` - the compute region for AI Platform Training and Prediction
# - `ARTIFACT_STORE` - the GCS bucket created during installation of AI Platform Pipelines. The bucket name will be similar to `qwiklabs-gcp-xx-xxxxxxx-kubeflowpipelines-default`.
# - `ENDPOINT` - set the `ENDPOINT` constant to the endpoint to your AI Platform Pipelines instance. Then endpoint to the AI Platform Pipelines instance can be found on the [AI Platform Pipelines](https://console.cloud.google.com/ai-platform/pipelines/clusters) page in the Google Cloud Console.
#
# 1. Open the **SETTINGS** for your instance
# 2. Use the value of the `host` variable in the **Connect to this Kubeflow Pipelines instance from a Python client via Kubeflow Pipelines SKD** section of the **SETTINGS** window.
#
# Run gsutil ls without URLs to list all of the Cloud Storage buckets under your default project ID.
# !gsutil ls
# **HINT:**
#
# For **ENDPOINT**, use the value of the `host` variable in the **Connect to this Kubeflow Pipelines instance from a Python client via Kubeflow Pipelines SDK** section of the **SETTINGS** window.
#
# For **ARTIFACT_STORE_URI**, copy the bucket name which starts with the qwiklabs-gcp-xx-xxxxxxx-kubeflowpipelines-default prefix from the previous cell output. Your copied value should look like **'gs://qwiklabs-gcp-xx-xxxxxxx-kubeflowpipelines-default'**
#
REGION = 'us-central1'
ENDPOINT = '337dd39580cbcbd2-dot-us-central2.pipelines.googleusercontent.com' # TO DO: REPLACE WITH YOUR ENDPOINT
ARTIFACT_STORE_URI = 'gs://qwiklabs-gcp-04-568443837277-kubeflowpipelines-default' # TO DO: REPLACE WITH YOUR ARTIFACT_STORE NAME
PROJECT_ID = !(gcloud config get-value core/project)
PROJECT_ID = PROJECT_ID[0]
# ### Build the trainer image
IMAGE_NAME='trainer_image'
TAG='latest'
TRAINER_IMAGE='gcr.io/{}/{}:{}'.format(PROJECT_ID, IMAGE_NAME, TAG)
# #### **Note**: Please ignore any **incompatibility ERROR** that may appear for the packages visions as it will not affect the lab's functionality.
# !gcloud builds submit --timeout 15m --tag $TRAINER_IMAGE trainer_image
# ### Build the base image for custom components
IMAGE_NAME='base_image'
TAG='latest'
BASE_IMAGE='gcr.io/{}/{}:{}'.format(PROJECT_ID, IMAGE_NAME, TAG)
# !gcloud builds submit --timeout 15m --tag $BASE_IMAGE base_image
# ### Compile the pipeline
#
# You can compile the DSL using an API from the **KFP SDK** or using the **KFP** compiler.
#
# To compile the pipeline DSL using the **KFP** compiler.
# #### Set the pipeline's compile time settings
#
# The pipeline can run using a security context of the GKE default node pool's service account or the service account defined in the `user-gcp-sa` secret of the Kubernetes namespace hosting KFP. If you want to use the `user-gcp-sa` service account you change the value of `USE_KFP_SA` to `True`.
#
# Note that the default AI Platform Pipelines configuration does not define the `user-gcp-sa` secret.
# +
USE_KFP_SA = False
COMPONENT_URL_SEARCH_PREFIX = 'https://raw.githubusercontent.com/kubeflow/pipelines/0.2.5/components/gcp/'
RUNTIME_VERSION = '1.15'
PYTHON_VERSION = '3.7'
# %env USE_KFP_SA={USE_KFP_SA}
# %env BASE_IMAGE={BASE_IMAGE}
# %env TRAINER_IMAGE={TRAINER_IMAGE}
# %env COMPONENT_URL_SEARCH_PREFIX={COMPONENT_URL_SEARCH_PREFIX}
# %env RUNTIME_VERSION={RUNTIME_VERSION}
# %env PYTHON_VERSION={PYTHON_VERSION}
# -
# #### Use the CLI compiler to compile the pipeline
# ### Exercise
#
# Compile the `covertype_training_pipeline.py` with the `dsl-compile` command line:
#
# <ql-infobox><b>NOTE:</b> If you need help, you may take a look at the complete solution by navigating to **mlops-on-gcp > workshops > kfp-caip-sklearn > lab-02-kfp-pipeline** and opening **lab-02.ipynb**.
# </ql-infobox>
# !dsl-compile --py pipeline/covertype_training_pipeline.py --output covertype_training_pipeline.yaml
# The result is the `covertype_training_pipeline.yaml` file.
# !head covertype_training_pipeline.yaml
# ### Deploy the pipeline package
# ### Exercise
#
# Upload the pipeline to the Kubeflow cluster using the `kfp` command line:
#
# <ql-infobox><b>NOTE:</b> If you need help, you may take a look at the complete solution by navigating to **mlops-on-gcp > workshops > kfp-caip-sklearn > lab-02-kfp-pipeline** and opening **lab-02.ipynb**.
# </ql-infobox>
# +
PIPELINE_NAME='covertype_continuous_training'
# !kfp --endpoint $ENDPOINT pipeline upload \
# -p $PIPELINE_NAME \
# covertype_training_pipeline.yaml
# -
# ## Submitting pipeline runs
#
# You can trigger pipeline runs using an API from the KFP SDK or using KFP CLI. To submit the run using KFP CLI, execute the following commands. Notice how the pipeline's parameters are passed to the pipeline run.
#
# ### List the pipelines in AI Platform Pipelines
# !kfp --endpoint $ENDPOINT pipeline list
# ### Submit a run
#
# Find the ID of the `covertype_continuous_training` pipeline you uploaded in the previous step and update the value of `PIPELINE_ID` .
#
#
#
PIPELINE_ID='0918568d-758c-46cf-9752-e04a4403cd84' # TO DO: REPLACE WITH YOUR PIPELINE ID
# +
EXPERIMENT_NAME = 'Covertype_Classifier_Training'
RUN_ID = 'Run_001'
SOURCE_TABLE = 'covertype_dataset.covertype'
DATASET_ID = 'splits'
EVALUATION_METRIC = 'accuracy'
EVALUATION_METRIC_THRESHOLD = '0.69'
MODEL_ID = 'covertype_classifier'
VERSION_ID = 'v01'
REPLACE_EXISTING_VERSION = 'True'
GCS_STAGING_PATH = '{}/staging'.format(ARTIFACT_STORE_URI)
# -
# ### Exercise
#
# Run the pipeline using the `kfp` command line. Here are some of the variable
# you will have to use to pass to the pipeline:
#
# - EXPERIMENT_NAME is set to the experiment used to run the pipeline. You can choose any name you want. If the experiment does not exist it will be created by the command
# - RUN_ID is the name of the run. You can use an arbitrary name
# - PIPELINE_ID is the id of your pipeline. Use the value retrieved by the `kfp pipeline list` command
# - GCS_STAGING_PATH is the URI to the Cloud Storage location used by the pipeline to store intermediate files. By default, it is set to the `staging` folder in your artifact store.
# - REGION is a compute region for AI Platform Training and Prediction.
#
#
# <ql-infobox><b>NOTE:</b> If you need help, you may take a look at the complete solution by navigating to **mlops-on-gcp > workshops > kfp-caip-sklearn > lab-02-kfp-pipeline** and opening **lab-02.ipynb**.
# </ql-infobox>
# !kfp --endpoint $ENDPOINT run submit \
# -e $EXPERIMENT_NAME \
# -r $RUN_ID \
# -p $PIPELINE_ID \
# project_id=$PROJECT_ID \
# gcs_root=$GCS_STAGING_PATH \
# region=$REGION \
# source_table_name=$SOURCE_TABLE \
# dataset_id=$DATASET_ID \
# evaluation_metric_name=$EVALUATION_METRIC \
# evaluation_metric_threshold=$EVALUATION_METRIC_THRESHOLD \
# model_id=$MODEL_ID \
# version_id=$VERSION_ID \
# replace_existing_version=$REPLACE_EXISTING_VERSION
# ### Monitoring the run
#
# You can monitor the run using KFP UI. Follow the instructor who will walk you through the KFP UI and monitoring techniques.
#
# To access the KFP UI in your environment use the following URI:
#
# https://[ENDPOINT]
#
#
# **NOTE that your pipeline run may fail due to the bug in a BigQuery component that does not handle certain race conditions. If you observe the pipeline failure, re-run the last cell of the notebook to submit another pipeline run or retry the run from the KFP UI**
#
# <font size=-1>Licensed under the Apache License, Version 2.0 (the \"License\");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at [https://www.apache.org/licenses/LICENSE-2.0](https://www.apache.org/licenses/LICENSE-2.0)
#
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an \"AS IS\" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.</font>
| workshops/kfp-caip-sklearn/lab-02-kfp-pipeline/exercises/lab-02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# <div style="text-align:center"><font size=6><b>EJERCICIOS SEABORN</b></font></div>
#
# <div style="text-align:center"><img src="https://media4.giphy.com/media/vf3LO38xXNRrq/giphy.gif?cid=ecf05e47cl9pyl4yub1755skxt1tbhfl1359573ak9cd4d1n&rid=giphy.gif" /></div>
#
# ### 1. Importa pandas, pyplot y seaborn
import seaborn as sns
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# ### 2. Activa matplotlib de forma estática (eso del ``inline``)
# %matplotlib inline
# ### 3. Lee 'Pokemon.csv', dentro de la carpeta ``data`` algunos niveles más arriba, y lee la primera columna del csv como índice del DataFrame
import pandas as pd
df = pd.read_csv("../../../data/Pokemon.csv", encoding='latin1')
df
# ### 4. Muestra las primeras filas del dataset para hacerte una idea de los datos cargados
df.head()
# ### 5. Haz un gráfico de dispersión con ``lmplot()``, que es un método que además pinta una regresión lineal, para representar cómo varía el Ataque (eje X) en función de la Defensa (eje Y)
sns.lmplot(data=df, x='Attack', y='Defense')
# ¿Se ajusta bien? ¿No? Al fin y al cabo, lo que estamos haciendo es ver la relación lineal entre Ataque y Defensa, ¿podrías cuantificar esta relación de alguna manera y justificar así lo que acabas de ver?
df[['Attack', 'Defense']].corr().loc['Attack', 'Defense']
# ### 6. No queremos ver la línea recta del modelo de regresión lineal que relaciona Ataque y Defensa
#
# Prueba a consultar la [documentación](https://seaborn.pydata.org/generated/seaborn.lmplot.html) para ver qué parámetro debes modificar:
sns.lmplot('Attack', 'Defense', data=df, fit_reg=False)
# ### 7. Ahora queremos añadir otra dimensión más a nuestro gráfico, queremos ver el Stage junto con el Ataque y la Defensa
#
# Consulta la documentación (o básate en lo que hemos visto en clase en otros métodos) para representar Stage con el color:
sns.lmplot('Attack', 'Defense', data=df, fit_reg=False, hue='Stage')
# Limita ahora esta representación entre los valores 20 y 80 de Ataque. Para hacerlo, puede que no exista un parámetro en la documentación, y que lo tengas que hacer por otras vías:
fig = plt.figure(figsize=(15,15))
ax = sns.lmplot(data=df, x= 'Attack', y='Defense', fit_reg=False, hue='Stage')
plt.xlim(20,80);
# ### 8. Con las 3 gráficas juntas no seaprecian muy bien las relaciones, separa la gráfica en otras 3: una por cada valor de Stage
#
# Para ello, puedes comprobar cómo lo hemos hecho en el notebook anterior (aunque fuera para otro método de representación), cuando utilizábamos un parámetro para que nos dividiera las gráficas en función de su valor. Si no, también puedes consultar la documentación.
#
# (En este caso, no mantengas los límites del último apartado)
fig = plt.figure(figsize=(15,15))
ax = sns.lmplot(data=df, x= 'Attack', y='Defense', col='Stage', hue='Stage')
# Comprueba ahora, para cada uno de ellos, si alguno tiene una relación lineal lo suficientemente buena como para considerar que existe esta relación (establece el umbral en +/-0.7).
#
# Para ello, utiliza lo que hemos visto con Pandas para separar el Dataframe.
print(df[df['Stage']==1].corr().loc['Attack', 'Defense'])
print(df[df['Stage']==2].corr().loc['Attack', 'Defense'])
print(df[df['Stage']==3].corr().loc['Attack', 'Defense'])
# ### 9. Dibuja un diagrama de cajas con el dataframe
#
# ¿De qué? Pues de todo lo que puedas.
df.dtypes
sns.boxplot(data=df)
# ### 10. Como puedes observar, representar todo lo que podemos quizás no sea la mejor idea.
#
# Parece que las columnas Total, Stage y Legendary nos desvirtúan lo que estamos viendo, y la columna # tampoco parece que nos vaya a aportar mucho. Vuelve a dibujar los diagramas de caja sin incluirlas:
sns.boxplot(data=df.loc[:, 'HP':'Speed'])
sns.boxplot(data=df)
# ### 11. Dejemos los gráficos al lado por un momento. ¿Cuántos 'Type 1' diferentes hay en el DataFrame? ¿Cuáles? ¿Cuántos registros existen de cada tipo?
#
# Si puedes, hazlo con una sola instrucción que nos de toda esa información.
df['Type 1'].value_counts()
# ### 12. Quédate del Dataframe solo con los 'Type 1' que tengan más de 7 registros. ¿En cuáles de ellos puedes apreciar relación lineal entre el Ataque y la Defensa? (Umbral 0.7)
tipos = df['Type 1'].value_counts().apply(lambda x: x > 7)
df_join = tipos[tipos == True]
df_cruce = df.set_index("Type 1").join(df_join)
df_cruce[df_cruce['Type 1'].notnull()]
df_join
# +
tipos = df['Type 1'].value_counts().apply(lambda x: x > 7)
tipos = tipos[tipos == True].index
df_type = df[df['Type 1'].apply(lambda x: x in tipos)]
tipos_corr = []
for tipo in tipos:
if df_type[df_type['Type 1']==tipo][['Attack', 'Defense']].corr().loc['Attack', 'Defense'] >= 0.7:
tipos_corr.append(tipo)
tipos_corr
df_type
for i in df_type['Type 1'].unique():
df_type_2 = df_type[df_type['Type 1'] == i]
if round(df_type_2[['Attack', 'Defense']].corr().loc['Attack', 'Defense'], 4) > 0.7:
sns.set_style("whitegrid")
a = sns.pairplot(df_type_2[['Attack', 'Defense', 'HP']],
kind='reg')
a.fig.suptitle(i, y = 1.05);
# +
# tipos_corr = []
# for tipo in tipos:
# corr_value = df_type[df_type['Type 1']==tipo][['Attack', 'Defense']].corr().loc['Attack', 'Defense']
# if corr_value >= 0.7:
# tipos_corr.append((tipo, corr_value))
# tipos_corr
# -
# ### 13. Usa un estilo 'whitegrid' y representa un correlograma de las estadísticas Ataque, Defensa y HP, para cada uno de los tipos que has obtenido en el último ejercicio
#
# Para cambiar el estulo, puedes consultar este apartado de la [documentación](http://seaborn.pydata.org/generated/seaborn.set_theme.html#seaborn.set_theme). Y los correlogramas deberán representar también la regresión lineal.
# +
df_type
for i in df_type['Type 1'].unique():
df_type_2 = df_type[df_type['Type 1'] == i]
if round(df_type_2[['Attack', 'Defense']].corr().loc['Attack', 'Defense'], 4) > 0.7:
sns.set_style("whitegrid")
a = sns.pairplot(df_type_2[['Attack', 'Defense', 'HP']],
kind='reg')
print("AAAAAA")
plt.show()
a.fig.suptitle(i, y = 1.05);
# -
# ### 14. Representa las mismas relaciones que en el apartado anterior pero cuantificándolas en un mapa de calor:
#
# Si hacemos un bucle for para representar varios mapas de calor, se sobreescribirán uno sobre otro, por lo que no se verán bien. Para por verlo en 3 gráficas diferentes, deberíamos crear una nueva figura en cada iteración del bucle for con la siguiente línea ``fig = plt.figure()``.
#
# Por otra parte, si queremos fijar la barra de color, podemos utilizar un par de parámetros del método, descritos en la [documentación](https://seaborn.pydata.org/generated/seaborn.heatmap.html?highlight=heatmap#seaborn.heatmap) (si vas por orden no tardarás mucho).
for i in df_type['Type 1'].unique():
df_type_2 = df_type[df_type['Type 1'] == i]
if round(df_type_2[['Attack', 'Defense']].corr().loc['Attack', 'Defense'], 4) > 0.7:
fig = plt.figure()
sns.heatmap(df_type_2[['Attack', 'Defense', 'HP']].corr(), annot=True, linewidth=.5, vmin=-1, vmax=1)
for i in df['Type 1'].unique():
df_type_2 = df[df['Type 1'] == i]
if round(df_type_2[['Attack', 'Defense']].corr().loc['Attack', 'Defense'], 4) > 0.7:
fig = plt.figure()
# ### 15. Selecciona los tipos 'Grass', 'Fire' y 'Water', y refleja, en un diagrama de violín, el 'Type 1' (eje X) frente al Ataque (eje Y):
#
# Para ello, utiliza la función ``catplot``.
# ### 16. Está bien, pero los colores no se corresponden con lo que uno espera para agua, fuego y planta
#
# Para seleccionar los colores que quieras, puedes usar la paleta de colores mediante el siguiente parámetro:
#
# ``palette=sns.color_palette([color1, color2, color3, color4...])``
#
# donde colorX es el color especificado como lo harías para cualquier otro gráfico. El tamaño de la lista de colores se debe corresponder conlos diferentes valores de lo que estés representando en el eje X.
#
# Ahora que ya sabes, utiliza los colores aporpiados para cada uno
# Repite este gráfico de forma que en cada violín se represente tanto el Ataque como la Defensa, para lo que deberás modificar el dataframe:
# ### 17. Con el diagrama de violín vemos la forma de la distribución, pero no sabemos si Grass tiene muchos puntos y Fire tiene muy pocos.
#
# Para ver esto, podemos usar swarmplot. Pinta el primero de los diagramas de violines del ejercicio anterior pero con un swarmplot en lugar de un violín, para lo que tendrás que cambiar un parámetro del ``catplot``
# ### 18. Queremos ver los violines a la vez que los swarm, los podemos pintar juntos.
#
# Para ello, deberemos utilizar por separado ambas representaciones, no pudiéndolas combinar con catplot. Revisa cómo lo hemos hecho en el notebook anterior. Además, elimina los diagramas de caja dentro de los violines con el parámetro inner = None y pon una transparencia de 0.9
#
# ### 19. Veamos la función melt() de pandas. ``pd.melt()`` toma los valores de varias columnas y los junta todos en una misma columna nueva.
#
# Dado un DataFrame, ``melt`` recibe varios parámetros:
# * qué columnas del DataFrame se quedan igual
# * qué nombres de columnas del dataframe pasan a ser una columna nueva
# * cómo queremos llamar esas nuevas columnas
#
# Primero, crea un dataframe ``df_grass_fire_water`` quedándote con los valores de "Type 1": "Water", "Fire" y "Grass", y muestra las primeras filas para recordar su forma:
df_grass_fire_water = df[df['Type 1'].isin(["Grass", "Fire", "Water"])]
df_grass_fire_water.head()
# ### 20. Queremos dejar igual las variables 'Name' y 'Type 1', así como añadir una columa nueva cuyos valores sean las columnas que queremos apilar y que se llame 'New Column'
#
# Para llevar esto a cabo escribimos df_grass_fire_water.melt(id_vars=['Name', 'Type 1'], value_vars=[columna1, columna2...], var_name='New Column')
#
# donde \[columna1, columna2...\] es la lista con las columnas a apilar, que en este caso serán 'HP', 'Attack', 'Defense' y 'Speed'. Lo guardaremos en una variable llamada ``df_melted``
len(df_grass_fire_water[['Name', 'Type 1', 'HP', 'Attack', 'Defense', 'Speed']])
# +
df_melted = df_grass_fire_water.melt(id_vars=['Name', 'Type 1'], value_vars=['HP', 'Attack', 'Defense', 'Speed'], var_name='New Column')
df_melted
# -
# ### 21. Pinta el swarmplot con el melted y entiende qué está pasando
# ¿Cómo lo has hecho? ¿Has utilizado ``catplot`` o directamente ``swarmplot``? Utiliza la contraria:
# ### 22. Basándote en el gráfico anterior, añade color en función del 'Type 1'
#
# Para ello, utiliza el parámetro ``hue`` y usa la anterior paleta de colores Pokemon
# Ahora, prueba qué ocurre al utilizar el parámetro ``dodge``:
# ### 23. Pinta el histograma y la densidad de probabilidad de la Defensa del dataframe original
# ### 24. Compara, en 3 gráficas diferentes, las densidades de probabilidad del Defensa (primera) y los histogramas del Ataque (segunda) para los que tienen 'Fire' como 'Type 1' frente a los que tienen "Water" y "Grass"
#
# Respeta los colores que hemos usado con anterioridad y añade una leyenda
# ¿Cuál te parece mejor? Utilízala para comparar la Velocidad de estos 3 tipos:
# ### 25. Pinta un diagrama de barras de la columna 'Type 1' usando el DataFrame original
#
# Para ello, utiliza ``sns.countplot``. Usa ``plt.xticks`` para rotar las etiquetas de los tipos. Si lo necesitas, fíjate en los ejemplos de [aquí](https://matplotlib.org/3.1.1/api/_as_gen/matplotlib.pyplot.xticks.html).
#
# Además, muéstralos ordenados de mayor a menor, para ello, podrás utilizar el parámetro ``order`` del ``countplot``, al que deberás pasarle una lista de strings con los valores de la variable cualitativa.
# ### 26. Usa catplot para realizar 2 subplots: 1 (izquierda) para el Type 1 y otro (derecha) para el Type 2, donde se diferencie si es Legendary o no con el color
# ### 27. Muestra una función distribución de probabilidad conjunta de la Defensa respecto al Ataque.
#
# Una vez lo tengas, identifica, apoximadamente, en qué bin 2D donde caen la mayoría de puntos (ataque, defensa).
#
# Identifica las etiquetas de los ejes y ponle un título a la gráfica.
# Realiza ahora la misma comparación pero de tal manera que se vean los puntos agrupados en bins de forma hexagonal, y que también se muestren los histogramas de cada una, todo en el mismo gráfico:
# ### 28. Selecciona los Pokémons 3, 149 y 150, y representa con gráficos de barras, en 3 figuras diferentes, los valores de los stats 'HP', 'Attack', 'Defense', 'Sp. Atk', 'Sp. Def', 'Speed' del cada uno de estos Pokémon frente a los valores máximos para su Type 1:
#
# Para construir el dataframe, recuerda que puedes utilizar la función ``pd.melt()``
# ### 29. Quédate con los Pokémon de Stage 3 y calcula la media de sus stats principales ('HP', 'Attack', 'Defense', 'Sp. Atk', 'Sp. Def' y 'Speed')
#
# Representa, en un diagrama de violín, la suma de los stats de cada uno de ellos, marcando cada uno de los valores dentro del propio violín.
# Ahora agrupa todos los pokémons de en función de su Stage (diferenciando, en el caso de que sea Stage 1, si es legendario o no; que sería otro grupo más), y representa la suma de sus stats en un diagrama de tarta.
# Quédate ahora con los que empiecen por "D" y representa, en un gráfico de donut, la media de sus totales en función del Tipo 1:
# Finalmente, repítelo con todos los registros:
| Bloque 2 - Data_Analyst/04_Visualización/03_Seaborn/03_Ejercicios_Seaborn.ipynb |