code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Dependencies and Setup
import pandas as pd
import numpy as np
import time
from pprint import pprint
import requests
from datetime import date, timedelta, datetime
import json
from pprint import pprint
from tqdm import tqdm
from tqdm import tqdm_notebook
# sqlite Dependencies
# ----------------------------------
# Imports the method used for connecting to DBs
from sqlalchemy import create_engine
# Allow us to declare column types
from sqlalchemy import Column, Integer, String, Text, DateTime, Float, Boolean, ForeignKey
from sqlalchemy.orm import relationship
from sqlalchemy.ext.declarative import declarative_base
from sqlalchemy.orm import Session
from nba_api.stats.endpoints import playercareerstats, drafthistory, commonplayerinfo, playerawards
# +
# cumestatsplayer,draftcombinedrillresults,playerdashboardbyteamperformance, leagueleaders, leaguedashplayerstats, draftcombinenonstationaryshooting, draftcombinestats, commonallplayers,
# +
# pd.show_versions()
# -
# ## Static Examples
import cudf, io, requests
from io import StringIO
# +
import cudf, io, requests
from io import StringIO
url = "https://github.com/plotly/datasets/raw/master/tips.csv"
content = requests.get(url).content.decode('utf-8')
tips_df = cudf.read_csv(StringIO(content))
tips_df['tip_percentage'] = tips_df['tip'] / tips_df['total_bill'] * 100
# display average tip by dining party size
print(tips_df.groupby('size').tip_percentage.mean())
# -
# +
from nba_api.stats.static import players
# Find players by full name.
players.find_players_by_full_name('james')
# Find players by first name.
players.find_players_by_first_name('lebron')
# Find players by last name.
players.find_players_by_last_name('^(james|love)$')
# Get all players.
# players.get_players()
# -
# + [markdown] heading_collapsed=true
# ## Begin Database Connection (sqlite)
# + hidden=true
# + hidden=true
# Base = declarative_base()
# engine = create_engine('sqlite:///db.sqlite', echo=False)
# conn = engine.connect()
# Create (if not already in existence) the tables associated with our classes.
# Base.metadata.create_all(engine)
# # Create a Session Object to Connect to DB
# # ----------------------------------
# session = Session(bind=engine)
# + hidden=true
# Use this to clear out the db
# ----------------------------------
# Session.rollback(self)
# Base.metadata.drop_all(engine)
# session.commit()
# -
# ## List of all players
# find specific player
from nba_api.stats.static import players
# get_players returns a list of dictionaries, each representing a player.
nba_players = players.get_players()
print('Number of players fetched: {}'.format(len(nba_players)))
nba_players[:]
all_players = pd.DataFrame(nba_players)
# ### Find specific player
## loop to find player
player_name = [player for player in nba_players
if player['full_name'] == '<NAME>'][0]
player_name
# ## Get Career Stats by player id number
# Get player_id number from get_players above
# #### Key:
#
# * GP: Games Played
# * MIN: Minutes Played
# * FGM: Field Goals Made
# * FGA: Field Goals Attempted
# * FG%: Field Goal Percentage
# * 3PM: 3 Point Field Goals Made
# * 3PA: 3 Point Field Goals Attempted
# * 3P%: 3 Point Field Goals Percentage
# * FTM: Free Throws Made
# * FTA: Free Throws Attempted
# * FT%: Free Throw Percentage
# * OREB: Offensive Rebounds
# * DREB: Defensive Rebounds
# * REB: Rebounds
# * AST: Assists
# * TOV: Turnovers
# * STL: Steals
# * BLK: Blocks
# * PF: Personal Fouls
# * DD2: Double Doubles
# * TD3: Trible Doubles
# * PTS: Points
#
#
#
# example
# +
# Display all columns
pd.set_option('display.max_columns', 500)
# <NAME>
career = playercareerstats.PlayerCareerStats(player_id='203076')
career_df = career.get_data_frames()[0]
career_dict = career_df.to_dict('records')
first_5_years = career_df[0:5]
# -
career_df.columns
# -----------
# + [markdown] heading_collapsed=true
# #### PySpark databases
# + hidden=true
# import os
# import findspark
# findspark.init()
# + hidden=true
# # Start Spark session
# from pyspark.sql import SparkSession
# from pyspark import SparkFiles
# spark = SparkSession.builder.appName("nbaStats").getOrCreate()
# + hidden=true
# with open("sample.json", "w") as outfile:
# json.dump(career_dict, outfile)
# + hidden=true
# sc = spark.sparkContext
# path = "./sample.json"
# players_json_spark = spark.read.json(path)
# + hidden=true
# players_json_spark.printSchema()
# + hidden=true
# Creates a temporary view using the DataFrame
# players_json_spark.createOrReplaceTempView("people")
# + hidden=true
# players_json_spark.show()
# + hidden=true
# # playersDF_spark = spark.range(3).toDF("myCol")
# newRow = players_json_spark
# appended = players_json_spark.union(newRow)
# display(appended)
# + hidden=true
# appended.count()
# + hidden=true
# def customFunction(row):
# return (row)
# sample2 = appended.rdd.map(customFunction)
# sample2
# + hidden=true
# for ids in all_players_ids:
# player_to_pass = ids
# + hidden=true
# for f in appended.collect():
# print (f)
# + hidden=true
# playersDF_spark.withColumn('newprice', dataframe]).show()
# + hidden=true
# otherPeople = spark.read.json(path)
# otherPeople.show()
# otherPeople.select(otherPeople["AST"]).show()
# + hidden=true
# Create DataFrame manually
# dataframe = spark.createDataFrame(path, schema=main_df_columns)
# dataframe.show()
# + hidden=true
# + [markdown] hidden=true
# ------------
# -
# ## Get first 5 years of each player id.
all_players_ids = list(all_players['id'].values.astype(str))
len(all_players_ids)
all_players_col = ['PLAYER_ID', 'SEASON_ID', 'LEAGUE_ID', 'TEAM_ID', 'TEAM_ABBREVIATION',
'PLAYER_AGE', 'GP', 'GS', 'MIN', 'FGM', 'FGA', 'FG_PCT', 'FG3M', 'FG3A',
'FG3_PCT', 'FTM', 'FTA', 'FT_PCT', 'OREB', 'DREB', 'REB', 'AST', 'STL',
'BLK', 'TOV', 'PF', 'PTS']
len(all_players_col)
# +
main_df = pd.DataFrame(columns=all_players_col)
pbar = tqdm(total=len(all_players_ids))
try:
for ids in all_players_ids:
player_to_pass = ids
career = playercareerstats.PlayerCareerStats(player_id=player_to_pass)
career_df = career.get_data_frames()[0]
first_5_years = career_df[0:5]
main_df = main_df.append(first_5_years, ignore_index = True)
pbar.update(1)
pbar.close()
except:
print("something happened")
# -
all_players_ids
five_year_all_players = main_df.sort_values(by='SEASON_ID', ascending=False).copy()
five_year_all_players = five_year_all_players.reset_index(drop=True)
five_year_all_players
five_year_all_players_to_json = five_year_all_players.to_json(orient='records')
# SAVE: Player_position
with open(f'./_players_all_data.json', 'w') as fp:
json.dump(five_year_all_players_to_json, fp)
# -------------
five_year_all_players.set_index('SEASON_ID')
# +
## Save all players data to sqlite db 'all_players'
# all_players.to_sql('all_players', con=engine)
## Retreive all players from sqlite db
# engine.execute("SELECT * FROM all_players").fetchall()
# -
# ## Draft History
draft_history = drafthistory.DraftHistory()
draft_history.get_data_frames()[0]
# ## Player Awards
player_awards = playerawards.PlayerAwards(player_id='203076')
player_awards.get_data_frames()[0]
# ## common_player_info
common_player_info = commonplayerinfo.CommonPlayerInfo(player_id='203076')
common_player_info.get_data_frames()[0]
# ## common_all_players
common_all_players = commonallplayers.CommonAllPlayers()
common_all_players.get_data_frames()[0]
# ------------
# Not so useful
# ## league_dash_player_stats
league_dash_player_bio_stats = leaguedashplayerbiostats.LeagueDashPlayerBioStats()
league_dash_player_bio_stats_df = league_dash_player_bio_stats.get_data_frames()[0]
league_dash_player_bio_stats_df
league_dash_player_bio_stats_df.columns
# ## League Leaders
league_leaders_df.columns
league_leaders = leagueleaders.LeagueLeaders()
league_leaders_df = league_leaders.get_data_frames()[0]
league_leaders_df
# ## player_dashboard_by_team_performance
# requires player_id number
player_dashboard_by_team_performance = playerdashboardbyteamperformance.PlayerDashboardByTeamPerformance(player_id='203076')
player_dashboard_by_team_performance.get_data_frames()[0]
boxscoreadvancedv22 = drafthistory.DraftHistory()
boxscoreadvancedv22.get_data_frames()[0]
# ## Draft Combine Drill Results
## Draft Combine Drill Results
draft_combine_drill = draftcombinedrillresults.DraftCombineDrillResults()
draft_combine_drill.get_data_frames()[0]
# ## Draft Combine Non Stationary Shooting
draft_combine_non_stationary_shooting = draftcombinenonstationaryshooting.DraftCombineNonStationaryShooting()
draft_combine_non_stationary_shooting.get_data_frames()[0]
# ## Draft Combine Stats
draft_combine_stats = draftcombinestats.DraftCombineStats()
draft_combine_stats.get_data_frames()[0]
| group_files/cpompa/notebooks/testing_purposes/nba-filter-5-years-player-data_cuDF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py36]
# language: python
# name: conda-env-py36-py
# ---
# # Threshold & GMM Analysis
# +
# %reload_ext autoreload
# %autoreload 2
from IPython.core.pylabtools import figsize
figsize(9, 9)
from tifffile import imread, imsave
from BlobDetector import BlobDetector
from BlobMetrics import BlobMetrics
from sklearn.mixture import GaussianMixture
import util
import matplotlib.pyplot as plt
import numpy as np
from scipy.stats import norm
from datetime import datetime
from skimage import morphology, transform
from skimage.filters import gaussian
from skimage.feature import blob_log
plt.set_cmap('gray')
source_dict = {
'cell_detection_0': 'COLM',
'cell_detection_1': 'COLM',
'cell_detection_2': 'COLM',
'cell_detection_3': 'COLM',
'cell_detection_4': 'laVision',
'cell_detection_5': 'laVision',
'cell_detection_6': 'laVision',
'cell_detection_7': 'laVision',
'cell_detection_8': 'laVision',
'cell_detection_9': 'laVision'
}
exp_name = 'cell_detection_8'
input_tif_path = '../data/s3617/tifs/{}.tiff'.format(exp_name)
output_csv_path = '../data/s3617/prediction_csv/{}.csv'.format(exp_name)
img = imread(input_tif_path)
ground_truth_path = '../data/s3617/annotations/{}.csv'.format(exp_name)
ground_truth = util.get_list_from_csv(ground_truth_path)
detector = BlobDetector(input_tif_path, data_source=source_dict[exp_name])
centroids = np.asarray(detector.get_blob_centroids())
util.write_list_to_csv(centroids, output_csv_path)
util.plot_csv_on_rgb_tif(centroids, input_tif_path, 'marked_final.tiff', color=[img.max(), 0, 0])
#util.plot_csv_on_rgb_tif(centroids, 'marked_final.tiff', 'marked_final.tiff', color=[img.max(), 0, 0])
print('detector threshold', detector.threshold)
metrics = BlobMetrics(ground_truth, centroids)
print('Precision: {}\nRecall: {}\nF-Measure: {}\n\n'.format(metrics.precision(), metrics.recall(), metrics.f_measure()))
# -
| notebooks/Threshold Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Comparison between `modin` + `ray` cvs file reading performance and `pandas`, `pyarrow` and `dask`
# ## Install modin
# + hide_input=false
# # !pip install 'modin[ray]'
# +
# # !wget https://s3.amazonaws.com/nyc-tlc/trip+data/yellow_tripdata_2020-01.csv
# -
# Number of CPUs
import os
print(f"CPUs: {os.cpu_count()}")
# ## Init ray
# +
import ray
ray.init(_plasma_directory="/tmp")
# -
# ## Import all libs
import modin.pandas as mpd
import pandas as pd
import pyarrow.csv as csv
import dask.dataframe as dd
file_name = 'yellow_tripdata_2020-01.csv'
# ## modin
# +
# %%timeit
df = mpd.read_csv(file_name)
# -
# ## pandas
# +
# %%timeit
df = pd.read_csv(file_name)
# -
# ## pyarrow
# +
# %%timeit
table = csv.read_csv(file_name)
df = table.to_pandas()
# -
# ## dask
# +
# %%timeit
df = dd.read_csv(file_name, assume_missing=True)
# convert to pandas
df = df.compute()
| snippets/modin-csv-test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import re
text = "I love data science and data munging, how about you ? ? ? "
text = re.sub('[\W]', ' ', text)
text
text = text.lower()
text = text.split()
text
from nltk.stem import WordNetLemmatizer
# +
lemmatizer = WordNetLemmatizer()
print(lemmatizer.lemmatize("cats"))
print(lemmatizer.lemmatize("cacti"))
print(lemmatizer.lemmatize("geese"))
print(lemmatizer.lemmatize("rocks"))
print(lemmatizer.lemmatize("python"))
print(lemmatizer.lemmatize("better", pos="a"))
print(lemmatizer.lemmatize("best", pos="a"))
print(lemmatizer.lemmatize("run"))
print(lemmatizer.lemmatize("run",'v'))
# -
| python regular expression for text processing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''fastai'': conda)'
# language: python
# name: python38564bitfastaicondad52d12c5a30a4725bf9d3e235cf1271c
# ---
# %load_ext autoreload
# %autoreload 2
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
from fastai.vision.all import *
import sklearn.metrics as skm
from tqdm.notebook import tqdm
import sklearn.feature_extraction.text
from transformers import (BertTokenizer, BertModel,
DistilBertTokenizer, DistilBertModel)
# -
from shopee_utils import *
from train_utils import *
PATH = Path('../input/shopee-product-matching')
model_file = '../input/resnet-model/bert814.pth'
if not PATH.is_dir():
PATH = Path('/home/slex/data/shopee')
model_file ='models/bert814.pth'
BERT_PATH = './bert_indonesian'
BERT_PATH='cahya/distilbert-base-indonesian'
train_df = pd.read_csv(PATH/'train_split.csv')
train_df['is_valid'] = train_df.split==0
def get_img_file(row):
img =row.image
fn = PATH/'train_images'/img
if not fn.is_file():
fn = PATH/'test_images'/img
return fn
class TitleTransform(Transform):
def __init__(self):
super().__init__()
self.tokenizer = DistilBertTokenizer.from_pretrained(BERT_PATH)
def encodes(self, row):
text = row.title
encodings = self.tokenizer(text, padding = 'max_length', max_length=50, truncation=True,return_tensors='pt')
keys =['input_ids', 'attention_mask']#, 'token_type_ids']
return tuple(encodings[key].squeeze() for key in keys)
# +
text_tfm = TitleTransform()
data_block = DataBlock(blocks = (ImageBlock(), TransformBlock(type_tfms=text_tfm),
CategoryBlock(vocab=train_df.label_group.to_list())),
splitter=ColSplitter(),
#splitter=RandomSplitter(),
get_y=ColReader('label_group'),
get_x=[get_img_file,lambda x:x],
item_tfms=Resize(460),
batch_tfms=aug_transforms(size=224, min_scale=0.75),
)
dls = data_block.dataloaders(train_df, bs=64,num_workers=16)
# -
b_im,b_txt,by=dls.one_batch()
b_im.shape
class ArcFaceClassifier(nn.Module):
def __init__(self, in_features, output_classes):
super().__init__()
self.W = nn.Parameter(torch.Tensor(in_features, output_classes))
nn.init.kaiming_uniform_(self.W)
def forward(self, x):
x_norm = F.normalize(x)
W_norm = F.normalize(self.W, dim=0)
return x_norm @ W_norm
class MultiModalModel(nn.Module):
def __init__(self, img_stem, text_stem):
super().__init__()
emb_dim = 1024+768
self.img_stem = img_stem
self.text_stem = text_stem
self.regularizers = nn.Sequential(
nn.BatchNorm1d(emb_dim),
nn.Dropout()
)
self.classifier=ArcFaceClassifier(emb_dim, dls.c)
#self.classifier=nn.Linear(emb_dim, dls.c)
self.outputEmbs = False
def forward(self, img_x, text_x):
img_out = self.img_stem(img_x)
text_out = self.text_stem(*text_x)
text_out = text_out.last_hidden_state[:,0,:]
embs = torch.cat([img_out, text_out],dim=1)
embs = self.regularizers(embs)
if self.outputEmbs:
return embs
return self.classifier(embs)
def new_model():
img_stem = nn.Sequential(create_body(resnet34,cut=-2), AdaptiveConcatPool2d(), Flatten())
bert_model = DistilBertModel.from_pretrained(BERT_PATH)
return MultiModalModel(img_stem, bert_model).cuda()
def split_2way(model):
return L(params(model.img_stem)+params(model.text_stem),
params(model.classifier)+params(model.regularizers))
learn = Learner(dls,new_model(), loss_func=arcface_loss, splitter=split_2way,cbs = F1FromEmbs, metrics=FakeMetric())
learn.fine_tune(20,1e-3,freeze_epochs=2)
| MultiTrain.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from os import listdir
from os import path
from keras.preprocessing.image import load_img
from keras.preprocessing.image import img_to_array
from keras.applications.vgg16 import preprocess_input
def load_photos(directory):
images = dict()
for name in listdir(directory):
# load an image from file
filename = path.join(directory, name)
image = load_img(filename, target_size=(224, 224))
# convert the image pixels to a numpy array
image = img_to_array(image)
# reshape data for the model
image = image.reshape((1, image.shape[0], image.shape[1], image.shape[2]))
# prepare the image for the VGG model
image = preprocess_input(image)
# get image id
image_id = name.split('.')[0]
images[image_id] = image
return images
# load images
directory = 'Flicker8k_Dataset'
images = load_photos(directory)
print('Loaded Images: %d' % len(images))
# +
from os import listdir
from os import path
from pickle import dump
from keras.applications.vgg16 import VGG16
from keras.preprocessing.image import load_img
from keras.preprocessing.image import img_to_array
from keras.applications.vgg16 import preprocess_input
from keras.models import Model
# extract features from each photo in the directory
def extract_features(directory):
# load the model
model = VGG16()
# re-structure the model
model.layers.pop()
model = Model(inputs=model.inputs, outputs=model.layers[-1].output)
# summarize
model.summary()
# extract features from each photo
features = dict()
for name in listdir(directory):
# load an image from file
filename = path.join(directory, name)
image = load_img(filename, target_size=(224, 224))
# convert the image pixels to a numpy array
image = img_to_array(image)
# reshape data for the model
image = image.reshape((1, image.shape[0], image.shape[1], image.shape[2]))
# prepare the image for the VGG model
image = preprocess_input(image)
# get features
feature = model.predict(image, verbose=0)
# get image id
image_id = name.split('.')[0]
# store feature
features[image_id] = feature
print('>%s' % name)
return features
# extract features from all images
directory = 'Flicker8k_Dataset'
features = extract_features(directory)
print('Extracted Features: %d' % len(features))
# save to file
dump(features, open('features.pkl', 'wb'))
# +
import string
import re
# load doc into memory
def load_doc(filename):
# open the file as read only
file = open(filename, 'r')
# read all text
text = file.read()
# close the file
file.close()
return text
# extract descriptions for images
def load_descriptions(doc):
mapping = dict()
# process lines
for line in doc.split('\n'):
# split line by white space
tokens = line.split()
if len(line) < 2:
continue
# take the first token as the image id, the rest as the description
image_id, image_desc = tokens[0], tokens[1:]
# remove filename from image id
image_id = image_id.split('.')[0]
# convert description tokens back to string
image_desc = ' '.join(image_desc)
# store the first description for each image
if image_id not in mapping:
mapping[image_id] = image_desc
return mapping
# clean description text
def clean_descriptions(descriptions):
# prepare regex for char filtering
re_punc = re.compile('[%s]' % re.escape(string.punctuation))
for key, desc in descriptions.items():
# tokenize
desc = desc.split()
# convert to lower case
desc = [word.lower() for word in desc]
# remove punctuation from each word
desc = [re_punc.sub('', w) for w in desc]
# remove hanging 's' and 'a'
desc = [word for word in desc if len(word)>1]
# store as string
descriptions[key] = ' '.join(desc)
# save descriptions to file, one per line
def save_doc(descriptions, filename):
lines = list()
for key, desc in descriptions.items():
lines.append(key + ' ' + desc)
data = '\n'.join(lines)
file = open(filename, 'w')
file.write(data)
file.close()
filename = 'Flickr8k.token.txt'
# load descriptions
doc = load_doc(filename)
# parse descriptions
descriptions = load_descriptions(doc)
print('Loaded: %d ' % len(descriptions))
# clean descriptions
clean_descriptions(descriptions)
# summarize vocabulary
all_tokens = ' '.join(descriptions.values()).split()
vocabulary = set(all_tokens)
print('Vocabulary Size: %d' % len(vocabulary))
# save descriptions
save_doc(descriptions, 'descriptions.txt')
# +
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.utils import to_categorical
# load doc into memory
def load_doc(filename):
# open the file as read only
file = open(filename, 'r')
# read all text
text = file.read()
# close the file
file.close()
return text
# load clean descriptions into memory
def load_clean_descriptions(filename):
doc = load_doc(filename)
descriptions = dict()
for line in doc.split('\n'):
# split line by white space
tokens = line.split()
# split id from description
image_id, image_desc = tokens[0], tokens[1:]
# store
descriptions[image_id] = ' '.join(image_desc)
return descriptions
descriptions = load_clean_descriptions('descriptions.txt')
print('Loaded %d' % (len(descriptions)))
# extract all text
desc_text = list(descriptions.values())
# prepare tokenizer
tokenizer = Tokenizer()
tokenizer.fit_on_texts(desc_text)
vocab_size = len(tokenizer.word_index) + 1
print('Vocabulary Size: %d' % vocab_size)
# integer encode descriptions
sequences = tokenizer.texts_to_sequences(desc_text)
# pad all sequences to a fixed length
max_length = max(len(s) for s in sequences)
print('Description Length: %d' % max_length)
padded = pad_sequences(sequences, maxlen=max_length, padding='post')
# one hot encode
y = to_categorical(padded, num_classes=vocab_size)
y = y.reshape((len(descriptions), max_length, vocab_size))
print(y.shape)
# +
from numpy import array
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.utils import to_categorical
# load doc into memory
def load_doc(filename):
# open the file as read only
file = open(filename, 'r')
# read all text
text = file.read()
# close the file
file.close()
return text
# load clean descriptions into memory
def load_clean_descriptions(filename):
doc = load_doc(filename)
descriptions = dict()
for line in doc.split('\n'):
# split line by white space
tokens = line.split()
# split id from description
image_id, image_desc = tokens[0], tokens[1:]
# store
descriptions[image_id] = ' '.join(image_desc)
return descriptions
descriptions = load_clean_descriptions('descriptions.txt')
print('Loaded %d' % (len(descriptions)))
# extract all text
desc_text = list(descriptions.values())
# prepare tokenizer
tokenizer = Tokenizer()
tokenizer.fit_on_texts(desc_text)
vocab_size = len(tokenizer.word_index) + 1
print('Vocabulary Size: %d' % vocab_size)
# integer encode descriptions
sequences = tokenizer.texts_to_sequences(desc_text)
# determine the maximum sequence length
max_length = max(len(s) for s in sequences)
print('Description Length: %d' % max_length)
X, y = list(), list()
for img_no, seq in enumerate(sequences):
# split one sequence into multiple X,y pairs
for i in range(1, len(seq)):
# split into input and output pair
in_seq, out_seq = seq[:i], seq[i]
# pad input sequence
in_seq = pad_sequences([in_seq], maxlen=max_length)[0]
# encode output sequence
out_seq = to_categorical([out_seq], num_classes=vocab_size)[0]
# store
X.append(in_seq)
y.append(out_seq)
# convert to numpy arrays
X, y = array(X), array(y)
print(X.shape)
print(y.shape)
# +
from os import listdir
from os import path
from numpy import array
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.utils import to_categorical
from keras.preprocessing.image import load_img
from keras.preprocessing.image import img_to_array
from keras.applications.vgg16 import preprocess_input
# load doc into memory
def load_doc(filename):
# open the file as read only
file = open(filename, 'r')
# read all text
text = file.read()
# close the file
file.close()
return text
# load clean descriptions into memory
def load_clean_descriptions(filename):
doc = load_doc(filename)
descriptions = dict()
for line in doc.split('\n'):
# split line by white space
tokens = line.split()
# split id from description
image_id, image_desc = tokens[0], tokens[1:]
# store
descriptions[image_id] = ' '.join(image_desc)
return descriptions
# fit a tokenizer given caption descriptions
def create_tokenizer(descriptions):
lines = list(descriptions.values())
tokenizer = Tokenizer()
tokenizer.fit_on_texts(lines)
return tokenizer
# load a single photo intended as input for the VGG feature extractor model
def load_photo(filename):
image = load_img(filename, target_size=(224, 224))
# convert the image pixels to a numpy array
image = img_to_array(image)
# reshape data for the model
image = image.reshape((1, image.shape[0], image.shape[1], image.shape[2]))
# prepare the image for the VGG model
image = preprocess_input(image)[0]
# get image id
image_id = path.basename(filename).split('.')[0]
return image, image_id
# create sequences of images, input sequences and output words for an image
def create_sequences(tokenizer, max_length, desc, image):
Ximages, XSeq, y = list(), list(),list()
vocab_size = len(tokenizer.word_index) + 1
# integer encode the description
seq = tokenizer.texts_to_sequences([desc])[0]
# split one sequence into multiple X,y pairs
for i in range(1, len(seq)):
# select
in_seq, out_seq = seq[:i], seq[i]
# pad input sequence
in_seq = pad_sequences([in_seq], maxlen=max_length)[0]
# encode output sequence
out_seq = to_categorical([out_seq], num_classes=vocab_size)[0]
# store
Ximages.append(image)
XSeq.append(in_seq)
y.append(out_seq)
Ximages, XSeq, y = array(Ximages), array(XSeq), array(y)
return [Ximages, XSeq, y]
# data generator, intended to be used in a call to model.fit_generator()
def data_generator(descriptions, tokenizer, max_length):
# loop for ever over images
directory = 'Flicker8k_Dataset'
while 1:
for name in listdir(directory):
# load an image from file
filename = path.join(directory, name)
image, image_id = load_photo(filename)
# create word sequences
desc = descriptions[image_id]
in_img, in_seq, out_word = create_sequences(tokenizer, max_length, desc, image)
yield [[in_img, in_seq], out_word]
# load mapping of ids to descriptions
descriptions = load_clean_descriptions('descriptions.txt')
# integer encode sequences of words
tokenizer = create_tokenizer(descriptions)
# pad to fixed length
max_length = max(len(s.split()) for s in list(descriptions.values()))
print('Description Length: %d' % max_length)
# test the data generator
generator = data_generator(descriptions, tokenizer, max_length)
inputs, outputs = next(generator)
print(inputs[0].shape)
print(inputs[1].shape)
print(outputs.shape)
# +
import string
import re
# load doc into memory
def load_doc(filename):
# open the file as read only
file = open(filename, 'r')
# read all text
text = file.read()
# close the file
file.close()
return text
# extract descriptions for images
def load_descriptions(doc):
mapping = dict()
# process lines
for line in doc.split('\n'):
# split line by white space
tokens = line.split()
if len(line) < 2:
continue
# take the first token as the image id, the rest as the description
image_id, image_desc = tokens[0], tokens[1:]
# remove filename from image id
image_id = image_id.split('.')[0]
# convert description tokens back to string
image_desc = ' '.join(image_desc)
# create the list if needed
if image_id not in mapping:
mapping[image_id] = list()
# store description
mapping[image_id].append(image_desc)
return mapping
def clean_descriptions(descriptions):
# prepare regex for char filtering
re_punc = re.compile('[%s]' % re.escape(string.punctuation))
for _, desc_list in descriptions.items():
for i in range(len(desc_list)):
desc = desc_list[i]
# tokenize
desc = desc.split()
# convert to lower case
desc = [word.lower() for word in desc]
# remove punctuation from each token
desc = [re_punc.sub('', w) for w in desc]
# remove hanging 's' and 'a'
desc = [word for word in desc if len(word)>1]
# remove tokens with numbers in them
desc = [word for word in desc if word.isalpha()]
# store as string
desc_list[i] = ' '.join(desc)
# convert the loaded descriptions into a vocabulary of words
def to_vocabulary(descriptions):
# build a list of all description strings
all_desc = set()
for key in descriptions.keys():
[all_desc.update(d.split()) for d in descriptions[key]]
return all_desc
# save descriptions to file, one per line
def save_descriptions(descriptions, filename):
lines = list()
for key, desc_list in descriptions.items():
for desc in desc_list:
lines.append(key + ' ' + desc)
data = '\n'.join(lines)
file = open(filename, 'w')
file.write(data)
file.close()
filename = 'Flickr8k.token.txt'
# load descriptions
doc = load_doc(filename)
# parse descriptions
descriptions = load_descriptions(doc)
print('Loaded: %d ' % len(descriptions))
# clean descriptions
clean_descriptions(descriptions)
# summarize vocabulary
vocabulary = to_vocabulary(descriptions)
print('Vocabulary Size: %d' % len(vocabulary))
# save to file
save_descriptions(descriptions, 'descriptions.txt')
# +
from numpy import array
from pickle import load
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.utils import to_categorical
from keras.utils import plot_model
from keras.models import Model
from keras.layers import Input
from keras.layers import Dense
from keras.layers import LSTM
from keras.layers import Embedding
from keras.layers import Dropout
from keras.layers.merge import add
from keras.callbacks import ModelCheckpoint
# load doc into memory
def load_doc(filename):
# open the file as read only
file = open(filename, 'r')
# read all text
text = file.read()
# close the file
file.close()
return text
# load a pre-defined list of photo identifiers
def load_set(filename):
doc = load_doc(filename)
dataset = list()
# process line by line
for line in doc.split('\n'):
# skip empty lines
if len(line) < 1:
continue
# get the image identifier
identifier = line.split('.')[0]
dataset.append(identifier)
return set(dataset)
# load clean descriptions into memory
def load_clean_descriptions(filename, dataset):
# load document
doc = load_doc(filename)
descriptions = dict()
for line in doc.split('\n'):
# split line by white space
tokens = line.split()
# split id from description
image_id, image_desc = tokens[0], tokens[1:]
# skip images not in the set
if image_id in dataset:
# create list
if image_id not in descriptions:
descriptions[image_id] = list()
# wrap description in tokens
desc = 'startseq ' + ' '.join(image_desc) + ' endseq'
# store
descriptions[image_id].append(desc)
return descriptions
# load photo features
def load_photo_features(filename, dataset):
# load all features
all_features = load(open(filename, 'rb'))
# filter features
features = {k: all_features[k] for k in dataset}
return features
# covert a dictionary of clean descriptions to a list of descriptions
def to_lines(descriptions):
all_desc = list()
for key in descriptions.keys():
[all_desc.append(d) for d in descriptions[key]]
return all_desc
# fit a tokenizer given caption descriptions
def create_tokenizer(descriptions):
lines = to_lines(descriptions)
tokenizer = Tokenizer()
tokenizer.fit_on_texts(lines)
return tokenizer
# calculate the length of the description with the most words
def max_length(descriptions):
lines = to_lines(descriptions)
return max(len(d.split()) for d in lines)
# create sequences of images, input sequences and output words for an image
def create_sequences(tokenizer, max_length, descriptions, photos):
X1, X2, y = list(), list(), list()
# walk through each image identifier
for key, desc_list in descriptions.items():
# walk through each description for the image
for desc in desc_list:
# encode the sequence
seq = tokenizer.texts_to_sequences([desc])[0]
# split one sequence into multiple X,y pairs
for i in range(1, len(seq)):
# split into input and output pair
in_seq, out_seq = seq[:i], seq[i]
# pad input sequence
in_seq = pad_sequences([in_seq], maxlen=max_length)[0]
# encode output sequence
out_seq = to_categorical([out_seq], num_classes=vocab_size)[0]
# store
X1.append(photos[key][0])
X2.append(in_seq)
y.append(out_seq)
return array(X1), array(X2), array(y)
# define the captioning model
def define_model(vocab_size, max_length):
# feature extractor model
inputs1 = Input(shape=(4096,))
fe1 = Dropout(0.5)(inputs1)
fe2 = Dense(256, activation='relu')(fe1)
# sequence model
inputs2 = Input(shape=(max_length,))
se1 = Embedding(vocab_size, 256, mask_zero=True)(inputs2)
se2 = Dropout(0.5)(se1)
se3 = LSTM(256)(se2)
# decoder model
decoder1 = add([fe2, se3])
decoder2 = Dense(256, activation='relu')(decoder1)
outputs = Dense(vocab_size, activation='softmax')(decoder2)
# tie it together [image, seq] [word]
model = Model(inputs=[inputs1, inputs2], outputs=outputs)
# compile model
model.compile(loss='categorical_crossentropy', optimizer='adam')
# summarize model
model.summary()
return model
# load training dataset (6K)
filename = 'Flickr_8k.trainImages.txt'
train = load_set(filename)
print('Dataset: %d' % len(train))
# descriptions
train_descriptions = load_clean_descriptions('descriptions.txt', train)
print('Descriptions: train=%d' % len(train_descriptions))
# photo features
train_features = load_photo_features('features.pkl', train)
print('Photos: train=%d' % len(train_features))
# prepare tokenizer
tokenizer = create_tokenizer(train_descriptions)
vocab_size = len(tokenizer.word_index) + 1
print('Vocabulary Size: %d' % vocab_size)
# determine the maximum sequence length
max_length = max_length(train_descriptions)
print('Description Length: %d' % max_length)
# prepare sequences
X1train, X2train, ytrain = create_sequences(tokenizer, max_length, train_descriptions,train_features)
# load test set
filename = 'Flickr_8k.devImages.txt'
test = load_set(filename)
print('Dataset: %d' % len(test))
# descriptions
test_descriptions = load_clean_descriptions('descriptions.txt', test)
print('Descriptions: test=%d' % len(test_descriptions))
# photo features
test_features = load_photo_features('features.pkl', test)
print('Photos: test=%d' % len(test_features))
# prepare sequences
X1test, X2test, ytest = create_sequences(tokenizer, max_length, test_descriptions,test_features)
# define the model
model = define_model(vocab_size, max_length)
# define checkpoint callback
checkpoint = ModelCheckpoint('model.h5', monitor='val_loss', verbose=1,save_best_only=True, mode='min')
# -
X1train.shape,X2train.shape,max_length
# fit model
model.fit([X1train, X2train], ytrain, epochs=30, verbose=1, callbacks=[checkpoint],validation_data=([X1test, X2test], ytest))
# +
from numpy import argmax
from pickle import load
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.models import load_model
from nltk.translate.bleu_score import corpus_bleu
# load doc into memory
def load_doc(filename):
# open the file as read only
file = open(filename, 'r')
# read all text
text = file.read()
# close the file
file.close()
return text
# load a pre-defined list of photo identifiers
def load_set(filename):
doc = load_doc(filename)
dataset = list()
# process line by line
for line in doc.split('\n'):
# skip empty lines
if len(line) < 1:
continue
# get the image identifier
identifier = line.split('.')[0]
dataset.append(identifier)
return set(dataset)
# load clean descriptions into memory
def load_clean_descriptions(filename, dataset):
# load document
doc = load_doc(filename)
descriptions = dict()
for line in doc.split('\n'):
# split line by white space
tokens = line.split()
# split id from description
image_id, image_desc = tokens[0], tokens[1:]
# skip images not in the set
if image_id in dataset:
# create list
if image_id not in descriptions:
descriptions[image_id] = list()
# wrap description in tokens
desc = 'startseq ' + ' '.join(image_desc) + ' endseq'
# store
descriptions[image_id].append(desc)
return descriptions
# load photo features
def load_photo_features(filename, dataset):
# load all features
all_features = load(open(filename, 'rb'))
# filter features
features = {k: all_features[k] for k in dataset}
return features
# covert a dictionary of clean descriptions to a list of descriptions
def to_lines(descriptions):
all_desc = list()
for key in descriptions.keys():
[all_desc.append(d) for d in descriptions[key]]
return all_desc
# fit a tokenizer given caption descriptions
def create_tokenizer(descriptions):
lines = to_lines(descriptions)
tokenizer = Tokenizer()
tokenizer.fit_on_texts(lines)
return tokenizer
# calculate the length of the description with the most words
def max_length(descriptions):
lines = to_lines(descriptions)
return max(len(d.split()) for d in lines)
# map an integer to a word
def word_for_id(integer, tokenizer):
for word, index in tokenizer.word_index.items():
if index == integer:
return word
return None
# generate a description for an image
def generate_desc(model, tokenizer, photo, max_length):
# seed the generation process
in_text = 'startseq'
# iterate over the whole length of the sequence
for _ in range(max_length):
# integer encode input sequence
sequence = tokenizer.texts_to_sequences([in_text])[0]
# pad input
sequence = pad_sequences([sequence], maxlen=max_length)
# predict next word
yhat = model.predict([photo,sequence], verbose=0)
# convert probability to integer
yhat = argmax(yhat)
# map integer to word
word = word_for_id(yhat, tokenizer)
# stop if we cannot map the word
if word is None:
break
# append as input for generating the next word
in_text += ' ' + word
# stop if we predict the end of the sequence
if word == 'endseq':
break
return in_text
# remove start/end sequence tokens from a summary
def cleanup_summary(summary):
# remove start of sequence token
index = summary.find('startseq ')
if index > -1:
summary = summary[len('startseq '):]
# remove end of sequence token
index = summary.find(' endseq')
if index > -1:
summary = summary[:index]
return summary
def evaluate_model(model, descriptions, photos, tokenizer, max_length):
actual, predicted = list(), list()
# step over the whole set
for key, desc_list in descriptions.items():
# generate description
yhat = generate_desc(model, tokenizer, photos[key], max_length)
# clean up prediction
yhat = cleanup_summary(yhat)
# store actual and predicted
references = [cleanup_summary(d).split() for d in desc_list]
actual.append(references)
predicted.append(yhat.split())
# calculate BLEU score
print('BLEU-1: %f' % corpus_bleu(actual, predicted, weights=(1.0, 0, 0, 0)))
print('BLEU-2: %f' % corpus_bleu(actual, predicted, weights=(0.5, 0.5, 0, 0)))
print('BLEU-3: %f' % corpus_bleu(actual, predicted, weights=(0.3, 0.3, 0.3, 0)))
print('BLEU-4: %f' % corpus_bleu(actual, predicted, weights=(0.25, 0.25, 0.25, 0.25)))
# load training dataset (6K)
filename = 'Flickr_8k.trainImages.txt'
train = load_set(filename)
print('Dataset: %d' % len(train))
# descriptions
train_descriptions = load_clean_descriptions('descriptions.txt', train)
print('Descriptions: train=%d' % len(train_descriptions))
# prepare tokenizer
tokenizer = create_tokenizer(train_descriptions)
vocab_size = len(tokenizer.word_index) + 1
print('Vocabulary Size: %d' % vocab_size)
# determine the maximum sequence length
max_length = max_length(train_descriptions)-4
print('Description Length: %d' % max_length)
# load test set
filename = 'Flickr_8k.testImages.txt'
test = load_set(filename)
print('Dataset: %d' % len(test))
# descriptions
test_descriptions = load_clean_descriptions('descriptions.txt', test)
print('Descriptions: test=%d' % len(test_descriptions))
# photo features
test_features = load_photo_features('features.pkl', test)
print('Photos: test=%d' % len(test_features))
# load the model
filename = 'model.h5'
model = load_model(filename)
# evaluate model
evaluate_model(model, test_descriptions, test_features, tokenizer, max_length)
# +
from keras.preprocessing.text import Tokenizer
from pickle import dump
# load doc into memory
def load_doc(filename):
# open the file as read only
file = open(filename, 'r')
# read all text
text = file.read()
# close the file
file.close()
return text
# load a pre-defined list of photo identifiers
def load_set(filename):
doc = load_doc(filename)
dataset = list()
# process line by line
for line in doc.split('\n'):
# skip empty lines
if len(line) < 1:
continue
# get the image identifier
identifier = line.split('.')[0]
dataset.append(identifier)
return set(dataset)
# load clean descriptions into memory
def load_clean_descriptions(filename, dataset):
# load document
doc = load_doc(filename)
descriptions = dict()
for line in doc.split('\n'):
# split line by white space
tokens = line.split()
# split id from description
image_id, image_desc = tokens[0], tokens[1:]
# skip images not in the set
if image_id in dataset:
# create list
if image_id not in descriptions:
descriptions[image_id] = list()
# wrap description in tokens
desc = 'startseq ' + ' '.join(image_desc) + ' endseq'
# store
descriptions[image_id].append(desc)
return descriptions
# covert a dictionary of clean descriptions to a list of descriptions
def to_lines(descriptions):
all_desc = list()
for key in descriptions.keys():
[all_desc.append(d) for d in descriptions[key]]
return all_desc
# fit a tokenizer given caption descriptions
def create_tokenizer(descriptions):
lines = to_lines(descriptions)
tokenizer = Tokenizer()
tokenizer.fit_on_texts(lines)
return tokenizer
# load training dataset
filename = 'Flickr_8k.trainImages.txt'
train = load_set(filename)
print('Dataset: %d' % len(train))
# descriptions
train_descriptions = load_clean_descriptions('descriptions.txt', train)
print('Descriptions: train=%d' % len(train_descriptions))
# prepare tokenizer
tokenizer = create_tokenizer(train_descriptions)
# save the tokenizer
dump(tokenizer, open('tokenizer.pkl', 'wb'))
# +
from pickle import load
from numpy import argmax
from keras.preprocessing.sequence import pad_sequences
from keras.applications.vgg16 import VGG16
from keras.preprocessing.image import load_img
from keras.preprocessing.image import img_to_array
from keras.applications.vgg16 import preprocess_input
from keras.models import Model
from keras.models import load_model
# extract features from each photo in the directory
def extract_features(filename):
# load the model
model = VGG16()
# re-structure the model
model.layers.pop()
model = Model(inputs=model.inputs, outputs=model.layers[-1].output)
# load the photo
image = load_img(filename, target_size=(224, 224))
# convert the image pixels to a numpy array
image = img_to_array(image)
# reshape data for the model
image = image.reshape((1, image.shape[0], image.shape[1], image.shape[2]))
# prepare the image for the VGG model
image = preprocess_input(image)
# get features
feature = model.predict(image, verbose=0)
return feature
# map an integer to a word
def word_for_id(integer, tokenizer):
for word, index in tokenizer.word_index.items():
if index == integer:
return word
return None
# remove start/end sequence tokens from a summary
def cleanup_summary(summary):
# remove start of sequence token
index = summary.find('startseq ')
if index > -1:
summary = summary[len('startseq '):]
# remove end of sequence token
index = summary.find(' endseq')
if index > -1:
summary = summary[:index]
return summary
# generate a description for an image
def generate_desc(model, tokenizer, photo, max_length):
# seed the generation process
in_text = 'startseq'
# iterate over the whole length of the sequence
for _ in range(max_length):
# integer encode input sequence
sequence = tokenizer.texts_to_sequences([in_text])[0]
# pad input
sequence = pad_sequences([sequence], maxlen=max_length)
# predict next word
yhat = model.predict([photo,sequence], verbose=0)
# convert probability to integer
yhat = argmax(yhat)
# map integer to word
word = word_for_id(yhat, tokenizer)
# stop if we cannot map the word
if word is None:
break
# append as input for generating the next word
in_text += ' ' + word
# stop if we predict the end of the sequence
if word == 'endseq':
break
return in_text
# load the tokenizer
tokenizer = load(open('tokenizer.pkl', 'rb'))
# pre-define the max sequence length (from training)
max_length = 30
# load the model
model = load_model('model.h5')
# load and prepare the photograph
photo = extract_features('example.jpg')
# generate description
description = generate_desc(model, tokenizer, photo, max_length)
description = cleanup_summary(description)
print(description)
# -
| Neural Image Captioning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <center><font size="+4">Programming and Data Analytics 1 2021/2022</font></center>
# <center><font size="+2">Sant'Anna School of Advanced Studies, Pisa, Italy</font></center>
# <center><img src="https://github.com/EMbeDS-education/StatsAndComputing20212022/raw/main/PDA/jupyter/jupyterNotebooks/images/SSSA.png" width="700" alt="The extensible parallel architecture of MultiVeStA"></center>
#
# <center><font size="+2">Course responsible</font></center>
# <center><font size="+2"><NAME> <EMAIL></font></center>
#
# <center><font size="+2">Co-lecturer </font></center>
# <center><font size="+2"><NAME> <EMAIL></font></center>
#
# ---
# + [markdown] id="PzTk_3lR7k5D"
# <center><font size="+4">Assignments for</font></center>
# <center><font size="+4">Lecture 4: Control and Repetition Statements</font><br/></center>
# <center><font size="+2"> and CSV manipulation/visualization applied on COVID-19 data</font></center>
#
# ---
#
# + cellView="form" colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1975, "status": "ok", "timestamp": 1622909713233, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgLeN60kpJtdBKQxyRrU7GwxagDOGnFA3G3Z8BwMA=s64", "userId": "01113523768495748338"}, "user_tz": -120} id="p666zbf89eCD" outputId="cafe9bca-03cf-4d92-f4cb-0e9aab51ab82"
#@title RUN, BUT DO NOT MODIFY
# !curl -O https://raw.githubusercontent.com/EMbeDS-education/StatsAndComputing20212022/main/PDA/jupyter/jupyterNotebooks/assignments/auto_testing.py
# !curl -O https://raw.githubusercontent.com/EMbeDS-education/StatsAndComputing20212022/main/PDA/jupyter/jupyterNotebooks/assignments/dpc-covid19-ita-andamento-nazionale.csv
# %reload_ext autoreload
# %autoreload 2
from auto_testing import *
# + [markdown] id="Om177ojl7k5J"
# # Assignment 04.01: If-Linear equation
# ## Statement
#
# Write a program that solves a linear equation _ax = b_ in **integers**.
#
# Given two integers _a_ and _b_ (_a_ may be zero),
# - print a single integer root if it exists or
# - print `no solution` if no integer solution exists
# - print `many solutions` if many integer solutions exist.
#
# ## Example input #1
#
# ```
# 1
# ```
#
# ```
# -2
# ```
#
# ## Example output #1
#
# ```
# -2
# ```
#
# ## Example input #2
#
# ```
# 2
# ```
#
# ```
# -1
# ```
#
# ## Example output #2
#
# ```
# no solution
# ```
#
#
# + [markdown] id="FMZE1vmMbRLz"
# ## Write your solution here
#
#
# * Do not change the first line (`def ...():`)
# * Maintain the given indentation
# * You can run some tests by yourself by decommenting the last line
#
#
# + executionInfo={"elapsed": 13, "status": "ok", "timestamp": 1622909899205, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgLeN60kpJtdBKQxyRrU7GwxagDOGnFA3G3Z8BwMA=s64", "userId": "01113523768495748338"}, "user_tz": -120} id="Yg4jeBnA7k5K"
def asgn04_01If_Linear_equation():
# This program prints 'Hello, world!':
print('Hello, world!')
# Change it to solve the assignment
#You can test independently your solution by executing the following line
#asgn04_01If_Linear_equation()
# + [markdown] id="_lqTangP8R46"
# ## Run the following cells to perform the provided tests
# + cellView="form" id="uv3gdWMQ9FmP"
#@title RUN and TEST ALL
from IPython.display import display, Markdown
inputs=[[1,-2],[2,-1],[0,0],[5,0],[0,7],[10,11],[1,30000]]
expected_outputs=[["-2"],["no solution"],["many solutions"],['0'],\
["no solution"],["no solution"],["30000"]]
for k in range(len(inputs)):
display(Markdown(f'{k+1}. TEST {inputs[k]} = {",".join(expected_outputs[k])}'))
print('-'*60)
run_and_test(inputs[k],expected_outputs[k],asgn04_01If_Linear_equation)
# + [markdown] id="1JMDln-vIKRB"
# # Assignment 04.02: If-Queen move
# ## Statement
#
# Chess queen moves horizontally, vertically or diagonally in any number of squares. <br/>
# Given two different squares of the chessboard, determine whether a queen can go from the first square to the second one in a single move. <br/>
# The chessboard is assumed to not contain further pieces.
#
# The program receives four numbers from 1 to 8 each specifying the column and the row number (the first two refer to the starting square, the last two refer to the target one). <br/>
# The program should output
# - `YES` if a queen can go from the first square to the second one in a single move or
# - `NO` otherwise.
#
#
# ## Example input
#
# ```
# 1
# 1
# 2
# 2
# ```
#
# ## Example output
#
# ```
# YES
# ```
#
# We suggest you to use intelligible (self-commenting) names for the variables...
#
# + [markdown] id="IROCYUDQb0t7"
# ## Write your solution here
#
#
# * Do not change the first line (`def ...():`)
# * Maintain the given indentation
# * You can run some tests by yourself by decommenting the last line
#
#
# + executionInfo={"elapsed": 284, "status": "ok", "timestamp": 1622909965077, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgLeN60kpJtdBKQxyRrU7GwxagDOGnFA3G3Z8BwMA=s64", "userId": "01113523768495748338"}, "user_tz": -120} id="QtPMkR0aJRYv"
def asgn04_02If_Queen_move():
# This program reads a string and prints it
str1=input()
print(str1)
# Can you change it to solve the assignment?
#You can test independently your solution by executing the following line
#asgn04_02If_Queen_move()
# + [markdown] id="HcoyoBpScQ7f"
# ## Run the following cells to perform the provided tests
# + cellView="form" id="X6tvTspi-MiJ"
#@title RUN and TEST ALL
from IPython.display import display, Markdown
inputs=[[1,1,2,2],[1,1,2,3],[5,6,3,3],[3,3,1,1],[6,5,2,5],\
[7,6,5,2],[2,7,6,7],[2,7,4,6],[7,4,2,5],[7,5,1,1],\
[2,4,5,7],[3,5,7,1],[5,2,5,8],[1,2,3,1],[2,1,1,3]]
expected_outputs=[["YES"],["NO"],["NO"],["YES"],["YES"],\
["NO"],["YES"],["NO"],["NO"],["NO"],\
["YES"],["YES"],["YES"],["NO"],["NO"]]
for k in range(len(inputs)):
display(Markdown(f'{k+1}. TEST {inputs[k]} = {",".join(expected_outputs[k])}'))
print('-'*60)
run_and_test(inputs[k],expected_outputs[k],asgn04_02If_Queen_move)
# + [markdown] id="CvRjQW_ITQ8T"
# # Assignment 04.03: Read integers
# ## Statement
#
# Write a program that reads positive integer numbers from the keyboard as long as the user provides them. <br/>
# The program should terminate as soon as the user provides an input that is not numeric (e.g. `'end'`, `'ciao'`, ...), which is considered as a terminating token.
#
# Upon reading the termination token, the program prints in separate lines:
#
# - how many numbers were given
# - how many odd numbers were given
# - the mean of the given numbers
# - the maximum given number
#
# **Untested extra challenge:** some of you might be tempted to solve this assignment using a list to store all numbers, and then iterate it. This is problematic in case of big data, or live data, as there might too many numbers to store. Also, it would be inefficient, as you would have to iterate the list of numbers twice.
#
# - Try to solve this assignment without using a list. You should define a variable per quantity we want to observe (or more if necessary), and update them every time a new number arrives.
#
# ## Example input 1
#
# ```
# 1
# ```
#
# ```
# 2
# ```
#
# ```
# 3
# ```
#
# ```
# end
# ```
#
# ## Example output 1
#
# ```
# 3
# ```
#
# ```
# 2
# ```
#
# ```
# 2.0
# ```
#
# ```
# 3
# ```
#
# ## Hints
#
# Here a suggestion on how to compute the maximum:
#
# - Define a variable `m` with value smaller than any value that might be given.
# - Every time a new number `n` is given, compare `m` with `n`, and update `m` if `n` is bigger than `m`.
#
# Whenever you read with `input()`, you read a string. Strings have a method to check if they represent a number...
# + [markdown] id="Lr-UYAK6b18z"
# ## Write your solution here
#
#
# * Do not change the first line (`def ...():`)
# * Maintain the given indentation
# * You can run some tests by yourself by decommenting the last line
#
#
# + executionInfo={"elapsed": 261, "status": "ok", "timestamp": 1622910617438, "user": {"displayName": "<NAME>ari", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgLeN60kpJtdBKQxyRrU7GwxagDOGnFA3G3Z8BwMA=s64", "userId": "01113523768495748338"}, "user_tz": -120} id="ghJWGiEPUD9v"
def asgn04_03Read_integers():
# This program reads a numbwe and prints it
a = int(input())
print(a)
# Change it according to the assignment description
#You can test independently your solution by executing the following line
#asgn04_03Read_integers()
# + [markdown] id="XJA5H0ULcVAM"
# ## Run the following cells to perform the provided tests
# + id="42vQBC_gAm4e"
#@title RUN and TEST ALL
from IPython.display import display, Markdown
inputs=[['1', '2', '3', 'end'],['1', '2', '3', 'fine'],['1', '3', '5', '7', '9', '11', 'ciao'],\
['11', '9', '7', '5', '3', '1', 'ciao']]
expected_outputs=[["3","2","2.0","3"],["3","2","2.0","3"],['6', '6', '6.0', '11'],\
['6', '6', '6.0', '11']]
for k in range(len(inputs)):
display(Markdown(f'{k+1}. TEST {inputs[k]} = {",".join(expected_outputs[k])}'))
print('-'*60)
run_and_test(inputs[k],expected_outputs[k],asgn04_03Read_integers)
# + [markdown] id="XYSmynIm83GP"
# # Assignment 04.04: While-Average of sequence
# ## Statement
#
# Read a sequence of non-negative integers, where each number is written in a separate line. The sequence ends with 0. Upon termination, print the average of the sequence.
#
# This assignment should be solved using while loops
#
# ## Example input
#
# ```
# 10
# ```
#
# ```
# 30
# ```
#
# ```
# 0
# ```
#
# ## Example output
#
# ```
# 20.0
# ```
#
#
# + [markdown] id="_XdWvj1R83GQ"
# ## Write your solution here
#
#
# * Do not change the first line (`def ...():`)
# * Maintain the given indentation
# * You can run some tests by yourself by decommenting the last line
#
#
# + executionInfo={"elapsed": 292, "status": "ok", "timestamp": 1622910780571, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgLeN60kpJtdBKQxyRrU7GwxagDOGnFA3G3Z8BwMA=s64", "userId": "01113523768495748338"}, "user_tz": -120} id="ewoiaJGW83GQ"
def asgn04_04While_Average_of_sequence():
# This program reads a numbwe and prints it
a = int(input())
print(a)
# Change it according to the assignment description
#You can test independently your solution by executing the following line
#asgn04_04While_Average_of_sequence()
# + [markdown] id="FCNXKoAJ83GV"
# ## Run the following cells to perform the provided tests
# + cellView="form" id="niuOKfaUBJUv"
#@title RUN and TEST ALL
from IPython.display import display, Markdown
inputs=[['10', '30', '0'],['1', '1', '1', '1', '0'],['1', '2', '0'],\
['1', '2', '3', '4', '0'],['1234', '0'],['1', '2', '3', '4', '5', '6', '7', '0']]
expected_outputs=[["20.0"],["1.0"],["1.5"],["2.5"],["1234.0"],["4.0"]]
for k in range(len(inputs)):
display(Markdown(f'{k+1}. TEST {inputs[k]} = {",".join(expected_outputs[k])}'))
print('-'*60)
run_and_test(inputs[k],expected_outputs[k],asgn04_04While_Average_of_sequence)
# + [markdown] id="OrzHQ6gm83GZ"
# # Assignment 04.05: While-Fibonacci
# ## Statement
#
# Fibonacci numbers are the numbers in the [integer sequence ](https://en.wikipedia.org/wiki/Integer_sequence)starting with 1, 1 where every number after the first two is the sum of the two preceding ones:
#
# 1, 1, 2, 3, 5, 8, 13, 21, 34, ...
#
# Given a positive integer **n**, print the **n**th Fibonacci number.
#
# This assignment should be solved using while loops
#
# ## Example input
#
# ```
# 6
# ```
#
# ## Example output
#
# ```
# 8
# ```
# + [markdown] id="4URO_dHJ83Ga"
# ## Write your solution here
#
#
# * Do not change the first line (`def ...():`)
# * Maintain the given indentation
# * You can run some tests by yourself by decommenting the last line
#
#
# + executionInfo={"elapsed": 269, "status": "ok", "timestamp": 1622910949398, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgLeN60kpJtdBKQxyRrU7GwxagDOGnFA3G3Z8BwMA=s64", "userId": "01113523768495748338"}, "user_tz": -120} id="x4ETfW8Z83Ga"
def asgn04_05While_Fibonacci():
# This program reads a numbwe and prints it
a = int(input())
print(a)
# Change it according to the assignment description
#You can test independently your solution by executing the following line
#asgn04_05While_Fibonacci()
# + [markdown] id="0NWXYSq483Gb"
# ## Run the following cells to perform the provided tests
# + cellView="form" id="YpB2xJG0BtVh"
#@title RUN and TEST ALL
from IPython.display import display, Markdown
inputs=[[6],[1],[2],[3],[4],[7],[14],[18]]
expected_outputs=[["8"],["1"],["1"],["2"],["3"],["13"],["377"],["2584"]]
for k in range(len(inputs)):
display(Markdown(f'{k+1}. TEST {inputs[k]} = {",".join(expected_outputs[k])}'))
print('-'*60)
run_and_test(inputs[k],expected_outputs[k],asgn04_05While_Fibonacci)
# + [markdown] id="ciulVFG383Gg"
# # Assignment 04.06: For-Sum of N numbers
# ## Statement
#
# `N` numbers are given in the input. Read them and print their sum.
#
# The first line of input contains the integer `N`, which is the number of integers to follow. Each of the next `N` lines contains one integer. Print the sum of these `N` integers.
#
# This assignment should be solved using for loops
#
# ## Example input
#
# ```
# 10
# 1
# 2
# 1
# 1
# 1
# 1
# 3
# 1
# 1
# 1
# ```
#
# ## Example output
#
# ```
# 13
# ```
#
#
# + [markdown] id="DGWTK0kR83Gi"
# ## Write your solution here
#
#
# * Do not change the first line (`def ...():`)
# * Maintain the given indentation
# * You can run some tests by yourself by decommenting the last line
#
#
# + executionInfo={"elapsed": 255, "status": "ok", "timestamp": 1622911176380, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgLeN60kpJtdBKQxyRrU7GwxagDOGnFA3G3Z8BwMA=s64", "userId": "01113523768495748338"}, "user_tz": -120} id="Enz3bEGR83Gi"
def asgn04_06For_Sum_of_N_numbers():
# This program reads a numbwe and prints it
a = int(input())
print(a)
# Change it according to the assignment description
#You can test independently your solution by executing the following line
#asgn04_06For_Sum_of_N_numbers()
# + [markdown] id="JeXi4A__83Gi"
# ## Run the following cells to perform the provided tests
# + id="JNnqG1QYCbv8"
#@title RUN and TEST ALL
from IPython.display import display, Markdown
inputs=[['1', '891'],['0'],['10', '1', '2', '1', '1', '1', '1', '3', '1', '1', '1'],\
['10', '1', '2', '3', '4', '5', '6', '7', '8', '9', '10'],['2', '235', '56'],\
['4', '4', '4', '4', '4']]
expected_outputs=[["891"],["0"],["13"],["55"],["291"],["16"]]
for k in range(len(inputs)):
display(Markdown(f'{k+1}. TEST {inputs[k]} = {",".join(expected_outputs[k])}'))
print('-'*60)
run_and_test(inputs[k],expected_outputs[k],asgn04_06For_Sum_of_N_numbers)
# + [markdown] id="3kuaF8ZI83Gm"
# # Assignment 04.07: For-Series
# ## Statement
#
# Given two integers `A` and `B`. Print all numbers from `A` to `B` inclusively, all in the same line separated by a space.
# In particular, the numbers should be printed
# - in increasing order, if `A < B`, or
# - in decreasing order, if `A >= B`.
#
# This assignment should be solved using for loops
#
# ## Example input 1
#
# ```
# 8
# ```
#
# ```
# 5
# ```
#
# ## Example output 1
#
# ```
# 8 7 6 5
# ```
#
# ## Example input 2
#
# ```
# 5
# ```
#
# ```
# 8
# ```
#
# ## Example output 2
#
# ```
# 5 6 7 8
# ```
#
# + [markdown] id="ubsNGAQY83Gm"
# ## Write your solution here
#
#
# * Do not change the first line (`def ...():`)
# * Maintain the given indentation
# * You can run some tests by yourself by decommenting the last line
#
#
# + executionInfo={"elapsed": 272, "status": "ok", "timestamp": 1622911281891, "user": {"displayName": "<NAME>ari", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgLeN60kpJtdBKQxyRrU7GwxagDOGnFA3G3Z8BwMA=s64", "userId": "01113523768495748338"}, "user_tz": -120} id="w-aeIJEw83Gn"
def asgn04_07For_Series():
# This program reads a numbwe and prints it
a = int(input())
print(a)
# Change it according to the assignment description
#You can test independently your solution by executing the following line
#asgn04_07For_Series()
# + [markdown] id="RF_4ydwf83Gr"
# ## Run the following cells to perform the provided tests
# + cellView="form" id="HukDAeBaC47-"
#@title RUN and TEST ALL
from IPython.display import display, Markdown
inputs=[['8', '5'],['5', '8'],['1', '10'],['179', '179'],['-14', '7'],['12', '-5']]
expected_outputs=[["8 7 6 5"],["5 6 7 8"],["1 2 3 4 5 6 7 8 9 10"],["179"],["-14 -13 -12 -11 -10 -9 -8 -7 -6 -5 -4 -3 -2 -1 0 1 2 3 4 5 6 7"],["12 11 10 9 8 7 6 5 4 3 2 1 0 -1 -2 -3 -4 -5"]]
for k in range(len(inputs)):
display(Markdown(f'{k+1}. TEST {inputs[k]} = {",".join(expected_outputs[k])}'))
print('-'*60)
run_and_test(inputs[k],expected_outputs[k],asgn04_07For_Series)
# + [markdown] id="Upj8pqov83HI"
# # Assignment 04.08: For-Sum of cubes
# ## Statement
#
# For the given integer `N`, calculate the following sum:
#
# $1^3$ + $2^3$ + ... + $N^3$
#
# This assignment should be solved using for loops
#
# ## Example input
#
# ```
# 3
# ```
#
# ## Example output
#
# ```
# 36
# ```
#
#
# + [markdown] id="eZlHiITb83HI"
# ## Write your solution here
#
#
# * Do not change the first line (`def ...():`)
# * Maintain the given indentation
# * You can run some tests by yourself by decommenting the last line
#
#
# + executionInfo={"elapsed": 6, "status": "ok", "timestamp": 1622911321436, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgLeN60kpJtdBKQxyRrU7GwxagDOGnFA3G3Z8BwMA=s64", "userId": "01113523768495748338"}, "user_tz": -120} id="AHo4vbwm83HJ"
def asgn04_08For_Sum_of_cubes():
# This program reads a numbwe and prints it
a = int(input())
print(a)
# Change it according to the assignment description
#You can test independently your solution by executing the following line
#asgn04_08For_Sum_of_cubes()
# + [markdown] id="bGdYWPpB83HJ"
# ## Run the following cells to perform the provided tests
# + id="32tr5T_yDXOQ"
#@title RUN and TEST ALL
from IPython.display import display, Markdown
inputs=[['1'],['2'],['3'],['4'],['9'],['20']]
expected_outputs=[["1"],["9"],["36"],["100"],["2025"],["44100"]]
for k in range(len(inputs)):
display(Markdown(f'{k+1}. TEST {inputs[k]} = {",".join(expected_outputs[k])}'))
print('-'*60)
run_and_test(inputs[k],expected_outputs[k],asgn04_08For_Sum_of_cubes)
# + [markdown] id="2lv3rrdh83HN"
# # Assignment 04.09: For-Sum of cubes
# ## Statement
#
# Write a program that
# - reads the CSV file **dpc-covid19-ita-andamento-nazionale.csv** provided here
# - It contains Italian official COVID'19 data (downloaded from [here](https://github.com/pcm-dpc/COVID-19) on 2020-05-08)
# - computes some quantities on the data in the CSV file.
#
# For this assignment, we are interested only in
# - the number of currently infected individuals (label totale_positivi), and
# - how they divide among being 'hospitalized' or in 'home isolation'.
#
# In particular, the infected individuals can be either
#
# - 'hospitalized' (label totale_ospedalizzati), or
# - in 'home isolation' (label isolamento_domiciliare)
#
# Your program should compute two lists with one entry per row in the CSV file:
#
# - `currently_hosp_perc` containing the percentage of infected individuals hospitalized
# - `currently_home_perc` containing the percentage of infected individuals in home isolation
#
# After computing these two lists, you should:
#
# - print the minimum value in `currently_hosp_perc`
# - print the maximum value in `currently_home_perc`
# - for each day for each we have measurements, print the corresponding entries in `currently_hosp_perc` and `currently_home_perc` in the same line but separated by a space. Each pair of numbers should be printed in a different row
#
# In all the cases, **you should print rounding up to the second decimal digit, using 4 digits in total.**
#
# We have shown in class that this can be obtained by doing:
#
# - `print( "%4.2f" % n ). # old way of doing this`
# - `print("{:4.2f}".format(n) ) # new way of doing this`
#
# where `n` is the float we want to print
#
# ## Hint
#
# What you are required to di is not so different from what we saw in class.
#
# We suggest you to first create a list per label of interest mentioned above:
#
# - currently_infected
# - currently_hosp
# - currently_home
#
# After the data has been loaded in the three lists, you should just properly iterate such lists to compute the required quantities, and print the required results.
#
# The function zip might help you in iterating the elements of `currently_hosp_perc` and `currently_home_perc` at the same time
#
# ## Expected output
#
# ```
# 17.97
# ```
#
# ```
# 82.03
# ```
#
# ```
# 57.47 42.53
# ```
#
# ```
# 48.08 51.92
# ```
#
# ```
# 42.60 57.40
# ```
#
# ```
# 51.70 48.30
# ```
#
# ```
# 49.82 50.18
# ```
#
# ```
# 48.24 51.76
# ```
#
# ```
# 49.40 50.60
# ```
#
# ```
# 49.48 50.52
# ```
#
# ```
# 55.81 44.19
# ```
#
# ```
# 60.64 39.36
# ```
#
# ```
# 64.96 35.04
# ```
#
# ```
# 72.93 27.07
# ```
#
# ```
# 63.58 36.42
# ```
#
# ```
# 65.87 34.13
# ```
#
# ```
# 63.23 36.77
# ```
#
# ```
# 69.47 30.53
# ```
#
# ```
# 64.83 35.17
# ```
#
# ```
# 60.78 39.22
# ```
#
# ```
# 58.54 41.46
# ```
#
# ```
# 55.72 44.28
# ```
#
# ```
# 55.02 44.98
# ```
#
# ```
# 55.81 44.19
# ```
#
# ```
# 57.38 42.62
# ```
#
# ```
# 57.89 42.11
# ```
#
# ```
# 55.00 45.00
# ```
#
# ```
# 49.33 50.67
# ```
#
# ```
# 48.18 51.82
# ```
#
# ```
# 49.01 50.99
# ```
#
# ```
# 47.40 52.60
# ```
#
# ```
# 46.89 53.11
# ```
#
# ```
# 46.25 53.75
# ```
#
# ```
# 45.74 54.26
# ```
#
# ```
# 44.81 55.19
# ```
#
# ```
# 43.58 56.42
# ```
#
# ```
# 42.36 57.64
# ```
#
# ```
# 42.07 57.93
# ```
#
# ```
# 41.50 58.50
# ```
#
# ```
# 40.26 59.74
# ```
#
# ```
# 39.25 60.75
# ```
#
# ```
# 38.42 61.58
# ```
#
# ```
# 37.39 62.61
# ```
#
# ```
# 36.08 63.92
# ```
#
# ```
# 35.28 64.72
# ```
#
# ```
# 34.56 65.44
# ```
#
# ```
# 33.78 66.22
# ```
#
# ```
# 33.04 66.96
# ```
#
# ```
# 32.30 67.70
# ```
#
# ```
# 31.44 68.56
# ```
#
# ```
# 30.50 69.50
# ```
#
# ```
# 30.19 69.81
# ```
#
# ```
# 29.91 70.09
# ```
#
# ```
# 29.14 70.86
# ```
#
# ```
# 27.98 72.02
# ```
#
# ```
# 26.74 73.26
# ```
#
# ```
# 25.74 74.26
# ```
#
# ```
# 25.56 74.44
# ```
#
# ```
# 25.39 74.61
# ```
#
# ```
# 24.70 75.30
# ```
#
# ```
# 24.32 75.68
# ```
#
# ```
# 23.53 76.47
# ```
#
# ```
# 22.76 77.24
# ```
#
# ```
# 22.33 77.67
# ```
#
# ```
# 22.04 77.96
# ```
#
# ```
# 21.08 78.92
# ```
#
# ```
# 20.52 79.48
# ```
#
# ```
# 20.07 79.93
# ```
#
# ```
# 19.54 80.46
# ```
#
# ```
# 18.97 81.03
# ```
#
# ```
# 18.76 81.24
# ```
#
# ```
# 18.71 81.29
# ```
#
# ```
# 18.31 81.69
# ```
#
# ```
# 17.97 82.03
# ```
#
# ```
# 18.68 81.32
# ```
#
# ```
# 18.39 81.61
# ```
#
# ```
# 17.97 82.03
# ```
#
#
# + [markdown] id="xl4cj0gU83HP"
# ## Write your solution here
#
#
# * Do not change the first line (`def ...():`)
# * Maintain the given indentation
# * You can run some tests by yourself by decommenting the last line
#
#
# + executionInfo={"elapsed": 251, "status": "ok", "timestamp": 1622911454100, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgLeN60kpJtdBKQxyRrU7GwxagDOGnFA3G3Z8BwMA=s64", "userId": "01113523768495748338"}, "user_tz": -120} id="shQim6bm83HQ"
import csv
def asgn04_09COVID_19():
#This program reads the csv file used and
# stores its header in the list header,
# then it prints all its rows
file_name='dpc-covid19-ita-andamento-nazionale.csv'
with open(file_name, 'r') as csvfile:
rows = csv.reader(csvfile, delimiter=',')
header = next(rows,None)
r=0
for row in rows :
print(r)
print(row)
r=r+1
#You can test independently your solution by executing the following line
#asgn04_09COVID_19()
# + [markdown] id="pDaBAJtu83HQ"
# ## Run the following cells to perform the provided tests
# + id="-O0Sz80383HQ"
#@title RUN AND TEST
inputs=[]
expected_outputs=['17.97', '82.03', '57.47 42.53', '48.08 51.92', '42.60 57.40', '51.70 48.30', '49.82 50.18', '48.24 51.76', '49.40 50.60', '49.48 50.52', '55.81 44.19', '60.64 39.36', '64.96 35.04', '72.93 27.07', '63.58 36.42', '65.87 34.13', '63.23 36.77', '69.47 30.53', '64.83 35.17', '60.78 39.22', '58.54 41.46', '55.72 44.28', '55.02 44.98', '55.81 44.19', '57.38 42.62', '57.89 42.11', '55.00 45.00', '49.33 50.67', '48.18 51.82', '49.01 50.99', '47.40 52.60', '46.89 53.11', '46.25 53.75', '45.74 54.26', '44.81 55.19', '43.58 56.42', '42.36 57.64', '42.07 57.93', '41.50 58.50', '40.26 59.74', '39.25 60.75', '38.42 61.58', '37.39 62.61', '36.08 63.92', '35.28 64.72', '34.56 65.44', '33.78 66.22', '33.04 66.96', '32.30 67.70', '31.44 68.56', '30.50 69.50', '30.19 69.81', '29.91 70.09', '29.14 70.86', '27.98 72.02', '26.74 73.26', '25.74 74.26', '25.56 74.44', '25.39 74.61', '24.70 75.30', '24.32 75.68', '23.53 76.47', '22.76 77.24', '22.33 77.67', '22.04 77.96', '21.08 78.92', '20.52 79.48', '20.07 79.93', '19.54 80.46', '18.97 81.03', '18.76 81.24', '18.71 81.29', '18.31 81.69', '17.97 82.03', '18.68 81.32', '18.39 81.61', '17.97 82.03']
run_and_test(inputs,expected_outputs,asgn04_09COVID_19)
| PDA/jupyter/jupyterNotebooks/assignments/.ipynb_checkpoints/04ControlAndRepetitionStatements_Assignments-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: env
# language: python
# name: env
# ---
# From the useful sheets, we have to formalize a GRL problem.
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import os
import requests
import io
import seaborn as sns
PATH = '../data/'
files = os.listdir(PATH)
dfs = {f[:-4] : pd.read_csv(PATH + f)
for f in files if f[-3:] == 'csv'
}
# All Users
# member_id is unique and reflected across all csvs (total 1207)
# all other things like DoB, IP, etc are most likely spoof
# Difference between name and member_title
# What is member_group_id?
# ID 4, 6, 10 are core moderators (from core_moderators)
dfs['core_members'].head(4);
core_members = dfs['core_members'][['member_id', 'name', 'member_title', 'member_posts']]
core_members.to_csv(PATH + 'modified/core_members.csv')
#3. 170 comments. comment_eid (event id?), comment_mid (member id?),
#comment_text,
# comment_author (string name) (doesn't match with any of member_title or name)
# might be useful to get some active users
# only comment_mid would be useful ig
dfs['calendar_event_comments'].head(4)
# 8066 follow connections
# follow_rel_id (?) and follow_member_id may be useful.
dfs['core_follow'].head(5)
dfs['core_follow'].follow_app.value_counts()
dfs['core_follow'].follow_area.value_counts()
dfs['core_follow'].follow_rel_id.nunique(), dfs['core_follow'].follow_member_id.nunique()
min(dfs['core_members'].member_id), max(dfs['core_members'].member_id)
min(dfs['core_follow'].follow_rel_id), max(dfs['core_follow'].follow_rel_id)
min(dfs['core_follow'].follow_member_id), max(dfs['core_follow'].follow_member_id)
# Important. All Private Messages. Total 21715.
# msg_topic_id is the thread ID; each convo bw two users has its own thread/topic
# msg_author_id is the same as member_id in core_members
# msg_post is the first message in the thread, with HTML tags
dfs['core_message_posts'].head(3)
#30. All Private Message Topics (total 4475)
# mt_id is same as msg_topic_id from core_message_posts
# mt_starter_id is the member_id of the person who put the first message on that topic
# mt_to_member_id is the member_id of the recipient of this private message
dfs['core_message_topics'].head(5)
#35. Pfields for each member. Total 1208. Could be used for node classification.
# dfs['core_pfields_content'].field_5[dfs['core_pfields_content'].field_5.isnull()]
# dfs['core_pfields_content'].field_5[190:196]
# print(sum(dfs['core_pfields_content'].field_5.isnull()))
# print(sum(dfs['core_pfields_content'].field_6.isnull()))
# print(sum(dfs['core_pfields_content'].field_7.isnull()))
# print(sum(dfs['core_pfields_content'].field_12.isnull()))
# print(sum(dfs['core_pfields_content'].field_13.isnull()))
# print(sum(dfs['core_pfields_content'].field_3.isnull()))
dfs['core_pfields_content'].head(10)
#43. Important. All Posts. Total 196042.
# index_content has the post content.
# index_author is the member_id of the authon of that post.
dfs['core_search_index'].head(4)
min(dfs['core_search_index'].index_author), max(dfs['core_search_index'].index_author)
#75. Useful. Notification Graph could give us very informative edges.
# from notify_from_id to notify_to_id based on notify_type_key
# and notify_type_key could be a nice edge feature
dfs['orig_inline_notifications'].notify_type_key.value_counts();
dfs['orig_inline_notifications'].head(5)
# +
#76. 763 Original members.
# check if all orig members have emails filled in
sum(dfs['orig_members'].email.isnull()) # (== 0) = True
orig_members = [name for name in dfs['orig_members'].name]
core_members = [name for name in dfs['core_members'].name]
print('based on username')
print(len(core_members), len(orig_members))
print(round(np.mean([
user in core_members
for user in orig_members
]), 2),
'= fraction of orig members in core members.')
print(round(np.mean([
user in orig_members
for user in core_members
]), 2),
'= fraction of core members in orig members.')
print('based on member_id')
orig_members = [user for user in dfs['orig_members'].member_id]
core_members = [user for user in dfs['core_members'].member_id]
print(len(core_members), len(orig_members))
print(round(np.mean([
user in core_members
for user in orig_members
]), 2),
'= fraction of orig members in core members.')
print(round(np.mean([
user in orig_members
for user in core_members
]), 2),
'= fraction of core members in orig members.')
# -
#78. Useful. But don't fully understand.
# Mapping from user_id to topic_id, might help connect users.
dfs['orig_message_topic_user_map'].head(4)
# TODO:
# Check if users following the same topic are same in
# this map and core_message_topics and orig_message_topics
# reference with topic_title and compare the topic_id v topic_title mapping of both
# orig and core
ids = 0
for i in range(len(dfs['orig_message_topics'].mt_title)):
if dfs['orig_message_topics'].mt_title[i] == dfs['core_message_topics'].mt_title[i]:
ids += 1
print(ids)
len(dfs['orig_message_topics'].mt_title), len(dfs['core_message_topics'].mt_title)
#79 All Orig Message Topics (total 3101)
# mt_id is same as map_topic_id from orig_message_topic_user_map
# mt_starter_id is the member_id of the person who put the first message on that topic
# mt_to_member_id is the member_id of the recipient of this message
dfs['orig_message_topics'].head(5)
#82. pfields of 764 members. Might help in node features.
dfs['orig_pfields_content'].head(3)
# What is reputation? Total 141550
# 635 users have a reputation index, could be used for node classification or features?
members = set(dfs['orig_reputation_index'].member_id)
# print(members)
freq = [[m, sum(dfs['orig_reputation_index'].member_id == m)]
for m in members]
# dfs['orig_reputation_index'].head(3)
freq_sort = sorted(freq, key = lambda z: z[1], reverse=True)
freq_sorted = pd.DataFrame(freq_sort)
i = 0
while freq_sorted[1][i] > 30:
i += 1
print(i)
print(len(freq_sorted[1]) - i)
plt.plot(freq_sorted[1])
plt.grid()
| src/.ipynb_checkpoints/37 > steel IS NOT graphite-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TP1 NLTK
# Natural Language Toolkit
# The Natural Language Toolkit, or more commonly NLTK, is a suite of libraries and programs for symbolic and statistical natural language processing (NLP) for English written in the Python programming language.
# +
## install nltk
# -
# !pip install nltk
# +
import nltk
from nltk.tokenize import sent_tokenize, word_tokenize,PunktSentenceTokenizer
# +
arabic_text="""ربما كانت أحد أهم التطورات التي قامت بها الرياضيات العربية التي بدأت في هذا الوقت بعمل الخوارزمي و هي بدايات الجبر، و من المهم فهم كيف كانت هذه الفكرة الجديدة مهمة، فقد كانت خطوة ثورية بعيدا عن المفهوم اليوناني للرياضيات التي هي في جوهرها هندسة، الجبر كان نظرية موحدة تتيح الأعداد الكسرية و الأعداد اللا كسرية، و قدم وسيلة للتنمية في هذا الموضوع مستقبلا. و جانب آخر مهم لإدخال أفكار الجبر و هو أنه سمح بتطبيق الرياضيات على نفسها بطريقة لم تحدث من قبل"""
english_text= """Perhaps one of the most significant advances made by Arabic mathematics began at this time with the work of al-Khwarizmi, namely
the beginnings of algebra. It is important to understand just how significant this new idea was. It was a revolutionary move away from
the Greek concept of mathematics which was essentially geometry. Algebra was a unifying theory which allowed rational
numbers, irrational numbers, geometrical magnitudes, etc., to all be treated as "algebraic objects". It gave mathematics a whole new
development path so much broader in concept to that which had existed before, and provided a vehicle for future development of the
subject. Another important aspect of the introduction of algebraic ideas was that it allowed mathematics to be applied to itself in a
way which had not happened before.
"""
# -
# # ***Tokenize text by Syllable***
tk = nltk.SyllableTokenizer()
print(tk.tokenize(english_text))
tk = nltk.SyllableTokenizer()
print(tk.tokenize(arabic_text))
# # tokenize text by word
# + active=""
# tokenize english text by word
# -
import nltk
from nltk.tokenize import sent_tokenize, word_tokenize,PunktSentenceTokenizer
english_text= """Perhaps one of the most significant advances made by Arabic mathematics began at this time with the work of al-Khwarizmi, namely
the beginnings of algebra. It is important to understand just how significant this new idea was. It was a revolutionary move away from
the Greek concept of mathematics which was essentially geometry. Algebra was a unifying theory which allowed rational
numbers, irrational numbers, geometrical magnitudes, etc., to all be treated as "algebraic objects". It gave mathematics a whole new
development path so much broader in concept to that which had existed before, and provided a vehicle for future development of the
subject. Another important aspect of the introduction of algebraic ideas was that it allowed mathematics to be applied to itself in a
way which had not happened before.
"""
for i in range(len(word_tokenize((english_text)))) :
print(word_tokenize((english_text))[i])
# # tokenize arabic text by word
# +
import nltk
from nltk.tokenize import sent_tokenize, word_tokenize,PunktSentenceTokenizer
arabic_text="""ربما كانت أحد أهم التطورات التي قامت بها الرياضيات العربية التي بدأت في هذا الوقت بعمل الخوارزمي و هي بدايات الجبر، و من المهم فهم كيف كانت هذه الفكرة الجديدة مهمة، فقد كانت خطوة ثورية بعيدا عن المفهوم اليوناني للرياضيات التي هي في جوهرها هندسة، الجبر كان نظرية موحدة تتيح الأعداد الكسرية و الأعداد اللا كسرية، و قدم وسيلة للتنمية في هذا الموضوع مستقبلا. و جانب آخر مهم لإدخال أفكار الجبر و هو أنه سمح بتطبيق الرياضيات على نفسها بطريقة لم تحدث من قبل"""
for i in range(len(word_tokenize((arabic_text)))) :
print(word_tokenize((arabic_text))[i])
# -
# # Sentence Tokenizing
# + active=""
# ----------------------tokenize english text by sentence
# -
for i in range(len(sent_tokenize(english_text))) :
print(sent_tokenize(english_text)[i])
print()
# + active=""
# ----------------------tokenize arabic text by sentence
# -
for i in range(len(sent_tokenize(arabic_text))) :
print(sent_tokenize(arabic_text)[i])
print()
# # Filtering Stop Words
nltk.download("stopwords")
# NLTK has by default a bunch of words that it considers to be stop words. It can be accessed via the NLTK corpus with:
from nltk.corpus import stopwords
#
# Example to incorporate the stop_words set to remove the stop words from a given text:
# +
english_text= """Perhaps one of the most significant advances made by Arabic mathematics began at this time with the work of al-Khwarizmi, namely
the beginnings of algebra. It is important to understand just how significant this new idea was. It was a revolutionary move away from
the Greek concept of mathematics which was essentially geometry. Algebra was a unifying theory which allowed rational
numbers, irrational numbers, geometrical magnitudes, etc., to all be treated as "algebraic objects". It gave mathematics a whole new
development path so much broader in concept to that which had existed before, and provided a vehicle for future development of the
subject. Another important aspect of the introduction of algebraic ideas was that it allowed mathematics to be applied to itself in a
way which had not happened before.
"""
words = nltk.word_tokenize(english_text)
print("Unfiltered: ", words)
stopwords = nltk.corpus.stopwords.words("english")
cleaned = [word for word in words if word not in stopwords]
print("Filtered: ", cleaned)
# -
# # You can observe the differences between the two lists Unfiltered & Filtered
# # Stemming English Text
# +
# importing modules
from nltk.stem import PorterStemmer
from nltk.tokenize import word_tokenize
ps = PorterStemmer()
words = word_tokenize(english_text)
for w in words:
print(w, " : ", ps.stem(w))
# -
# # Stemming Arabic Text
# +
# importing modules
from nltk.stem import PorterStemmer
from nltk.tokenize import word_tokenize
ps = PorterStemmer()
words = word_tokenize(arabic_text)
for w in words:
print(w, " : ", ps.stem(w))
# -
# # Tagging Parts of Speech
eng_pos_tag = nltk.pos_tag(word_tokenize(english_text))
nltk.help.upenn_tagset()
en_pos_tag = nltk.pos_tag(word_tokenize(english_text))
# # Lemmatizing
from nltk.stem import WordNetLemmatizer
lemmatizer = WordNetLemmatizer()
lemmatized_words = [lemmatizer.lemmatize(word) for word in word_tokenize(english_text)]
print(lemmatized_words)
# # mplementation: Chunking in NLP using nltk
# let us try to extract all the noun phrases from a sentence using the steps defined above. First, we’ll import the required libraries and then tokenize the sentence before applying POS_tagging to it.
# +
# Importing the required libraries
from nltk import pos_tag
from nltk import word_tokenize
from nltk import RegexpParser
# english_text
english_text= """Perhaps one of the most significant advances made by Arabic mathematics began at this time with the work of al-Khwarizmi, namely
the beginnings of algebra. It is important to understand just how significant this new idea was. It was a revolutionary move away from
the Greek concept of mathematics which was essentially geometry. Algebra was a unifying theory which allowed rational
numbers, irrational numbers, geometrical magnitudes, etc., to all be treated as "algebraic objects". It gave mathematics a whole new
development path so much broader in concept to that which had existed before, and provided a vehicle for future development of the
subject. Another important aspect of the introduction of algebraic ideas was that it allowed mathematics to be applied to itself in a
way which had not happened before.
"""
# Splitiing the sentence into words
list_of_words = word_tokenize(english_text)
# Applying POS_tagging
tagged_words = pos_tag(list_of_words)
# -
# We then define our chunk keeping in mind that our aim is to extract all the noun phrases present in our sentence.
# Extracting the Noun Phrases
chunk_to_be_extracted = r''' Chunk: {<DT>*<NNP>*<NN>*} '''
# Applying chunking to the text
chunkParser = nltk.chunk.RegexpParser(chunk_to_be_extracted)
chunked_sentence = chunkParser.parse(tagged_words)
# The ‘chunked_sentence’ variable is an NLTK tree which can be viewed using the draw() method.
# To view the NLTK tree
chunked_sentence.draw()
#
| nltk .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Given a Binary Search Tree (BST), convert it to a Greater Tree such that every key of the original BST is changed to the original key plus sum of all keys greater than the original key in BST.
#
# Example:
#
# Input: The root of a Binary Search Tree like this:
# 5
# / \
# 2 13
#
# Output: The root of a Greater Tree like this:
# 18
# / \
# 20 13
# ref:
# - [Approach #1 Recursion [Accepted]](https://leetcode.com/problems/convert-bst-to-greater-tree/solution/)
#
# > Intuition
# >
# > One way to perform a reverse in-order traversal is via recursion. By using the call stack to return to previous nodes, we can easily visit the nodes in reverse order.
# >
# >
# > Algorithm
# >
# > For the recursive approach, we maintain some minor "global" state so each recursive call can access and modify the current total sum. Essentially, we ensure that the current node exists, recurse on the right subtree, visit the current node by updating its value and the total sum, and finally recurse on the left subtree. If we know that recursing on root.right properly updates the right subtree and that recursing on root.left properly updates the left subtree, then we are guaranteed to update all nodes with larger values before the current node and all nodes with smaller values after.
#
#
# +
# Definition for a binary tree node.
# class TreeNode:
# def __init__(self, x):
# self.val = x
# self.left = None
# self.right = None
class Solution:
def __init__(self):
self.valsum = 0
def convertBST(self, root: TreeNode) -> TreeNode:
if root:
self.convertBST(root.right)
self.valsum += root.val
root.val = self.valsum
self.convertBST(root.left)
return root
| DSA/tree/convertBST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="IZ-z33sXQ_Pt"
# Implemente as funções abaixo. Você não deverá usar variáveis globais.
# + id="KFwX1rjCQ_P2"
from typing import List, Set
import random
# + id="oP-SSFJMKxbw"
def escolhe_palavra(arr_palavras:List[str]) -> str:
"""
Atividade 1: Escolhe aleatoriamente uma posição do vetor de arr_palavras
use e entenda a biblioteca random do python para isso.
Não esqueça de dar import na biblioteca. Coloque isso n célula acima
"""
indice_palavra = random.randint(0, len(arr_palavras)-1)
return arr_palavras[indice_palavra]
# + colab={"base_uri": "https://localhost:8080/", "height": 36} id="LzFJmwhULVJP" outputId="fc60499c-9fa4-4024-c699-25249ce43051"
#Faça aqui um teste desta função
vetor1 = ['abacaxi', 'maca', 'banana', 'pera', 'limao']
escolhe_palavra(vetor1)
# + id="-m_T1uUiK-jp"
def obtem_posicoes_letra(palavra:str, letra:str):
"""
Atividade 2: Em uma determinada palavra, obtem as posições em que
ocorre a letra, ambas passadas como paramtro
"""
set_posicoes = set()
#navega na palavra, substitua o "None" corretamente
#é possivel navegar em uma palavra como se fosse uma lista de letras
#ou seja, os mesmos comandos que funcionam em uma lista funcionam em uma string
for i,letra_palavra in zip(range(len(palavra)),palavra):
#substitua o pass para adicionar em set_posicoes as posições desta letra (passad como parametro)
#na palavra
if letra_palavra == letra:
set_posicoes.add(i)
return set_posicoes
# + colab={"base_uri": "https://localhost:8080/"} id="k3LulOUBLbiC" outputId="dadc171b-5220-4204-a81c-fdec74dc7e64"
#Faça aqui um teste desta função
obtem_posicoes_letra('abacaxi', 'a')
# + id="gdIDb7KtIVPY"
from IPython.display import clear_output
def imprime_forca(palavra_forca:str, posicoes_descobertas:Set[int]):
"""
Atividade 3: Imprime a palavra a ser exibida na tela
exibindo apenas os caracteres das posições descobertas.
Por exemplo, se a palavra for "casaco" e as posições descobertas
forem {1,3,5}, será impresso "_ a _ a _ o"
Nesta atividade, você deverá alterar apenas o if apropriadamente.
"""
clear_output(wait=False)
#palavra_exibida = obtem_palavra_exibida(palavra_forca, posicoes_descobertas)
for posicao,letra in enumerate(palavra_forca):
if posicao in posicoes_descobertas:
print(letra,end=" ")
else:
print("_",end=" ")
# + colab={"base_uri": "https://localhost:8080/"} id="WhGOf0uSJ9wj" outputId="b009f698-1971-4d79-a479-1b4f47a0bf19"
#Faça aqui um teste desta função
imprime_forca('casaco',{1,3,5})
# + id="dk8OJgxGLOog"
from IPython.display import clear_output
def desenha_bonequinho(erros):
switcher = {
0: "\n",
1: "\nO",
2: "\nO\n/",
3: "\nO\n/\\",
4: "\nO\n/\\\n||",
5: "\nO\n/\\\n||\n/",
6: "\nO\n/\\\n||\n/\\",
}
return switcher.get(erros)
# + id="uGLnnSeCLTLf"
def jogo_forca(palavra):
"""
Atividade 4: implemente o jogo da forca
usando as funções criadas
"""
erros = 0
conj = set()
while True:
while True:
letra = input("Informe uma letra: ")
if letra.isalpha():
break
else:
print("Apenas letras são permitidas!")
print(erros)
conj.update(obtem_posicoes_letra(palavra, letra))
imprime_forca(palavra, conj)
if len(obtem_posicoes_letra(palavra, letra)) == 0:
erros = erros + 1
print(desenha_bonequinho(erros))
if len(conj) == len(palavra):
print("\nVoce venceu!")
break
if erros == 6:
print("\nVoce perdeu!")
break
# + id="bKT3yVHJLjNZ" colab={"base_uri": "https://localhost:8080/", "height": 659} outputId="bd828a11-0d28-4421-fe7b-02acaf90e539"
#Faça aqui um teste da ultima função e se divirta :)
vetor1 = ['abacaxi', 'maca', 'banana', 'pera', 'limao']
palavra = escolhe_palavra(vetor1)
jogo_forca(palavra)
| Laboratório/Tafera 1/Tarefa___Forca_resolucao.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from coppertop.core import *
from coppertop.pipe import *
from bones.core.types import num
from bones.metatypes import BType
from coppertop.std import typeOf, struct
from coppertop.dm.canvas import *
# +
import io, urllib, base64
from IPython.display import HTML
@coppertop
def toJup(fig):
# http://nbviewer.ipython.org/github/ipython-books/cookbook-code/blob/master/notebooks/chapter01_basic/06_kernel.ipynb
imgdata = io.BytesIO()
fig.savefig(imgdata, format='png')
imgdata.seek(0)
b64 = urllib.parse.quote(base64.b64encode(imgdata.getvalue()))
return HTML('<div ><img src="data:image/png;base64,{0}"></img></div>'.format(b64))
# +
c = rough(height=300, width=300)
c \
>> style(_,fill='green', stroke='red') \
>> fill(_,1,1,300,1) >> fill(_,1,1,1,300) >> fill(_,1,300,300,300) >> fill(_,300,300,300,1) \
>> fill(_,3,3,298,298) >> clear(_,4,4,297,297) \
>> outline(_, 6,6,295,295) \
>> style(_,stroke='blue',width=1) \
>> line(_,10,10,291,291) \
>> style(_,font='32px "Comic Sans MS"') \
>> text(_, 50, 50, "Fred rocks!")
# >> style(_,font='font-family: "Comic Sans MS";') \
# +
(rough(height=300, width=300)
>> fill(_, 50, 250, 55, 255)
>> fill(_, 100, 250, 105, 255)
>> fill(_, 75-2.5, 200, 75+2.5, 205)
>> style(_, width=10)
# smile
>> line(_, 40,170,50,150)
>> line(_, 50,150,100,150)
>> line(_, 100,150,110,170)
# HI
>> style(_,font='32px "Comic Sans MS"')
>> text(_, 55, 100, "Hi<NAME>!")
)
# +
label = BType.ensure('label').setConstructor(struct)
spacer = BType.ensure('spacer').setConstructor(struct)
xy = BType.ensure('xy').setConstructor(struct)
scatter = BType.ensure('scatter').setConstructor(struct)
axis = BType.ensure('axis').setConstructor(struct)
defaults = struct(align='centre')
space3 = spacer(size=3)
space5 = spacer(size=5)
dataCanvas = xy(id=1, minDataX=0.0, maxDataX=1.0, minDataY=0.0, maxDataX=1.0)
title = label(text='My cool Tufte-compliant scatter graph')
layout = [
space5,
title,
space5,
[space5, yTitle, space3, yAxis, space5, dataCanvas, space5],
space5,
xAxis,
space3,
xTitle,
space5
]
myScatter = scatter(id='myscatter', xs=xs, ys=ys)
# -
c = rough(x1=-1, x2=302, y1=-1, y2=302) # 304 by 304
c \
>> style(_,fill='green', stroke='red') \
>> fill(_,1,1,300,1) >> fill(_,1,1,1,300) >> fill(_,1,300,300,300) >> fill(_,300,300,300,1) \
>> fill(_,3,3,298,298) >> clear(_,4,4,297,297) \
>> outline(_, 6,6,295,295) \
>> style(_,stroke='blue',width=1) \
>> line(_,10,10,291,291)
c.rotate >> DD
# +
from matplotlib.backends.backend_agg import FigureCanvasAgg
from matplotlib.figure import Figure
import numpy as np
fig = Figure(figsize=(5, 4), dpi=100)
# A canvas must be manually attached to the figure (pyplot would automatically
# do it). This is done by instantiating the canvas with the figure as
# argument.
canvas = FigureCanvasAgg(fig)
# -
canvas >> DD
# # %matplotlib inline
import matplotlib.pyplot as plt, numpy as np
fig >> DD
fig2 = plt.figure(figsize=(5, 5), dpi= 80, facecolor='w', edgecolor='k')
fig.canvas
from matplotlib.artist import Artist
class Fred(Artist):
pass
fred = Fred()
fig.add_artist(fred)
fred.show()
# + active=""
# fred
# -
fred >> DD
fig >> DD
fig.artists
import matplotlib.pyplot as plt
import numpy as np
N = 5
menMeans = (20, 35, 30, 35, -27)
womenMeans = (25, 32, 34, 20, -25)
menStd = (2, 3, 4, 1, 2)
womenStd = (3, 5, 2, 3, 3)
ind = np.arange(N) # the x locations for the groups
width = 0.35 # the width of the bars: can also be len(x) sequence
# fig, ax = plt.subplots()
fig = plt.figure()
fig >> DD
# +
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
p1 = ax.bar(ind, menMeans, width, yerr=menStd, label='Men')
p2 = ax.bar(ind, womenMeans, width,
bottom=menMeans, yerr=womenStd, label='Women')
ax.axhline(0, color='grey', linewidth=0.8)
ax.set_ylabel('Scores')
ax.set_title('Scores by group and gender')
ax.set_xticks(ind)
ax.set_xticklabels(('G1', 'G2', 'G3', 'G4', 'G5'))
ax.legend()
# Label with label_type 'center' instead of the default 'edge'
ax.bar_label(p1, label_type='center')
ax.bar_label(p2, label_type='center')
ax.bar_label(p2)
None
# -
x = np.linspace(0, 3*np.pi, 500)
plt.plot(x, np.sin(x**2))
plt.title('A simple chirp')
;
fig = plt.figure()
# +
fig = plt.figure()
fred2 = Fred()
fig.add_artist(fred2)
# -
plt.show()
fig.canvas.get_supported_filetypes()
from matplotlib.text import Text
fig = plt.figure()
fig.text(100,100,"hello")
plt.show()
hAx = plt.figure(figsize = (10, 10)).gca()
hAx >> DD
# +
import matplotlib.pyplot as plt
from matplotlib.figure import Figure
import numpy as np
class WatermarkFigure(Figure):
"""A figure with a text watermark."""
def __init__(self, *args, watermark=None, **kwargs):
super().__init__(*args, **kwargs)
if watermark is not None:
bbox = dict(boxstyle='square', lw=3, ec='gray', fc=(0.9, 0.9, .9, .5), alpha=0.5)
self.text(0.5, 0.5, watermark, ha='center', va='center', rotation=30, fontsize=40, color='gray', alpha=0.5, bbox=bbox)
x = np.linspace(-3, 3, 201)
y = np.tanh(x) + 0.1 * np.cos(5 * x)
fig2 = plt.figure(figsize = (5, 5), FigureClass=WatermarkFigure, watermark='draft')
# ax = fig.add_subplot(1,1,1)
# ax.set_visible(False)
# ax.plot(x, y)
# plt.plot(x, y)
# ax = fig.add_subplot(1,1,1)
# p1 = ax.bar(ind, menMeans, width, yerr=menStd, label='Men')
# plt.show(fig2)
fig2 >> toJup
# +
import matplotlib.pyplot as plt
fig = plt.figure()
plt.axis([0, 10, 0, 10])
t = ("This is a really long string that I'd rather have wrapped so that it "
"doesn't go outside of the figure, but if it's long enough it will go "
"off the top or bottom!")
plt.text(4, 1, t, ha='left', rotation=15, wrap=True)
plt.text(6, 5, t, ha='left', rotation=15, wrap=True)
# plt.text(5, 5, t, ha='right', rotation=-15, wrap=True)
# plt.text(5, 10, t, fontsize=18, style='oblique', ha='center',
# va='top', wrap=True)
# plt.text(3, 4, t, family='serif', style='italic', ha='right', wrap=True)
# plt.text(-1, 0, t, ha='left', rotation=-15, wrap=True)
fig >> toJup
# -
fig2 >> toJup
widgets.Image(value=_to_png(fig2))
from ipywidgets import widgets
# +
# # %matplotlib
# -
| canvas/basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python Pandas I/O
# ### Creating DataFrames, Reading and Writing to CSV & JSON files
# [Documentation](https://pandas.pydata.org/docs/index.html)
import numpy as np
import pandas as pd
import random
# ### Creating DataFrames from Lists and Dicts
# ▶ New DataFrame from a **List**
# Pandas automatically assigns numerical row indexes.
data1 = [random.random() for i in range(10000)]
df = pd.DataFrame(data1)
df.head()
# ▶ New DataFrame from a **2D List**
# Column names default to integers. Each subList is a row.
data2 = [[i, random.randint(10,99)] for i in 'ABCDEFGHIJKLMNOPQRSTUVWXYZ']
df = pd.DataFrame(data2)
df.head()
# ▶ New DataFrame from a **Dictionary**
# Dict Keys become column names
data3 = {
'Model':['T57','T61','T64','T65'],
'Price':[1.42,1.48,1.73,1.95],
'Size':['57 inches','61 inches','64 inches','65 inches']}
df = pd.DataFrame(data3)
df
# Change previous example to use Model number as index.
df = pd.DataFrame(
{'Price':data3['Price'],'Size':data3['Size']},
index=data3['Model'])
df
# ▶ New DataFrame from a List of Dictionaries
# Note, missing Length is populated with NaN (not a number).
data4 = [
{'Ht':63, 'Len':45, 'Wt':2.6},
{'Ht':29, 'Wt':1.7},
{'Ht':37, 'Len':71, 'Wt':4.2}]
df = pd.DataFrame(data4)
df
# ### Reading & Writing DataFrames to CSV Files
# [Documentation](https://pandas.pydata.org/docs/user_guide/io.html#csv-text-files) of numerous optional parameters.
#
# ▶ Write DataFrame to CSV file
df = pd.DataFrame(data4)
df.to_csv('outfile.csv', index=False) #, sep=';')
# ▶ Read CSV file into DataFrame
# Missing numerical data are given value NaN by default.
df = pd.read_csv('outfile.csv')
df
# ▶ Convert DataFrame to_string
df = pd.DataFrame(data4)
d4str = df.to_string()
d4str
# ### Reading & Writing DataFrames to JSON files
# [Documentation](https://pandas.pydata.org/docs/user_guide/io.html#csv-text-files) of numerous optional parameters.
#
# ▶ Convert DataFrame to **JSON** string
# No argument - json by columns is default, {column -> {index -> value}}
data4_json = df.to_json()
data4_json
# Use orient='index' to structure the json by rows, {index -> {column -> value}}.
# You can also strip out the row indices by using orient='records'.
data4_json = df.to_json(orient='index')
data4_json
# ▶ Write to a text file in JSON format.
data4_json = df.to_json('outjson.txt')
data4_json
# ▶ Read same JSON data back in to a DataFrame.
data4 = pd.read_json('outjson.txt')
data4
| Pandas/Python Pandas Input-Output.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: reddit
# language: python
# name: reddit
# ---
# # Stock Entity Recognition Unmasked
# +
import pandas
import re
import json
import math
import numpy
import os
import tensorflow as tf
from itertools import chain
from multiprocessing import Pool
from functools import partial
from transformers import TFBertForTokenClassification, BertTokenizerFast
from sklearn.model_selection import train_test_split
# -
# ## Config
# +
THREADS = 48
DEVICE = '/cpu:0'
TOKENS_AND_LABELS_TRAIN_LOC = 'data/ner/tokens_and_labels_train.parquet'
TOKENS_AND_LABELS_TEST_LOC = 'data/ner/tokens_and_labels_test.parquet'
MODEL_LOC = 'data/ner/unmasked/best.ckpt'
# -
# ## Training
tokens_and_labels_train = pandas.read_parquet(TOKENS_AND_LABELS_TRAIN_LOC)
tokens_and_labels_train
# +
tokenizer = BertTokenizerFast.from_pretrained('bert-base-uncased')
def encode_labels(max_length, chunk):
# loss function doesn't compute loss for labels if -100 so default to that
labels_encoded = numpy.ones((len(chunk), max_length), dtype=numpy.int8) * -100
# start filling in labels using the offset_mapping
for observation_i, (offset_mapping, labels_raw) in enumerate(chunk):
labels = numpy.array(labels_raw.split(' ')).astype(int)
label_i = 0
for offset_i, offset in enumerate(offset_mapping):
if offset[0] == 0 and offset[1] != 0:
labels_encoded[observation_i][offset_i] = labels[label_i]
label_i += 1
return labels_encoded
def encode_df(df, max_length=256):
# encode everything
inputs_encoded = tokenizer(
# split ourselves so we can align with labels
list(map(lambda o: o.split(' '), df['tokens'])),
return_tensors="tf",
is_split_into_words=True,
# offset mappings to align labels to the first word piece
return_offsets_mapping=True,
# make sure the same length across all encodings
max_length=max_length,
padding='max_length',
truncation=True
)
offsets_with_labels = list(zip(inputs_encoded.offset_mapping.numpy(), df['labels']))
chunk_size = len(offsets_with_labels) / THREADS
offsets_with_labels_chunks = [offsets_with_labels[round(chunk_size *i):round(chunk_size * (i + 1))] for i in range(0, THREADS)]
with Pool(THREADS) as pool:
encoded_labels = pool.map(partial(encode_labels, max_length), offsets_with_labels_chunks)
return inputs_encoded, numpy.stack(list(chain(*encoded_labels)))
# +
encoded = encode_df(tokens_and_labels_train)
tf_train_dataset = tf.data.Dataset.from_tensor_slices(({
'input_ids': encoded[0]['input_ids'],
'token_type_ids': encoded[0]['token_type_ids'],
'attention_mask': encoded[0]['attention_mask']
}, encoded[1])).batch(32)
# +
# %%time
model = TFBertForTokenClassification.from_pretrained('bert-base-uncased', num_labels = 2)
optimizer = tf.optimizers.Adam(learning_rate=5e-5)
model.compile(optimizer=optimizer, loss=model.compute_loss, metrics=['accuracy'])
model.summary()
cp_callback = tf.keras.callbacks.ModelCheckpoint(
filepath=MODEL_LOC,
save_weights_only=True,
verbose=1
)
model.fit(tf_train_dataset, epochs=5, callbacks=[cp_callback])
# -
# ## Inference
# load up the tokenizer and model
tokenizer = BertTokenizerFast.from_pretrained('bert-base-uncased')
model = TFBertForTokenClassification.from_pretrained('bert-base-uncased', num_labels = 2)
model.load_weights(MODEL_LOC)
def predict(sentence, max_length=256):
tokens = sentence.split(' ')
test_encoding = tokenizer(
tokens,
return_tensors="tf",
is_split_into_words=True,
max_length=max_length,
padding='max_length',
truncation=True,
return_offsets_mapping=True
)
# grab the offset mappings
offset_mapping = test_encoding.offset_mapping
del test_encoding['offset_mapping']
test_encoding
prediction = tf.argsort(model(test_encoding).logits[0])
token_predictions = []
num_tokens = len(test_encoding.attention_mask[0][test_encoding.attention_mask[0] == 1])
token_i = 0
for i in range(num_tokens):
offset = offset_mapping[0][i]
token_prediction = prediction[i]
if offset[0] == 0 and offset[1] != 0:
token_predictions.append([tokens[token_i], bool(token_prediction[1] == 1)])
token_i += 1
return token_predictions
with tf.device(DEVICE):
display(predict('my msft is crm'))
| StockEntityRecognitionUnmasked.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: MyPython
# language: Python
# name: mypython
# ---
# ##%overwritefile
# ## %file:src/templatefile.py
# ##%fnlist:CMake Mermaid Gradle
# ##%fnlist:Python C Dart Kotlin Groovy Java Batch PS R Swift Lua Perl PHP Tcl Vimscript M4 Dot
# ##%fnlist:Gjs Vala Nodejs Bash VBScript WLS Go Html TypeScript Make Ruby Rust
# ##%fileforlist:../../jupyter-My$fnlist-kernel/jupyter_My$fnlist_kernel/plugins/templatefile.py
# ##%noruncode
import typing as t
from typing import Dict, Tuple, Sequence,List
from plugins.ISpecialID import IStag,IDtag,IBtag,ITag
import os
class MyTemplatefile(IStag):
kobj=None
def getName(self) -> str:
# self.kobj._write_to_stdout("setKernelobj setKernelobj setKernelobj\n")
return 'MyTemplatefile'
def getAuthor(self) -> str:
return 'Author'
def getIntroduction(self) -> str:
return 'MyTemplatefile'
def getPriority(self)->int:
return 0
def getExcludeID(self)->List[str]:
return []
def getIDSptag(self) -> List[str]:
return ['templatefile']
def setKernelobj(self,obj):
self.kobj=obj
# self.kobj._write_to_stdout("setKernelobj setKernelobj setKernelobj\n")
return
def on_shutdown(self, restart):
return
def on_ISpCodescanning(self,key, value,magics,line) -> str:
# self.kobj._write_to_stdout(line+" on_ISpCodescanning\n")
self.kobj.addkey2dict(magics,'templatefile')
return self.templatehander(self,key, value,magics,line)
##在代码预处理前扫描代码时调用
def on_Codescanning(self,magics,code)->Tuple[bool,str]:
pass
return False,code
##生成文件时调用
def on_before_buildfile(self,code,magics)->Tuple[bool,str]:
return False,''
def on_after_buildfile(self,returncode,srcfile,magics)->bool:
return False
def on_before_compile(self,code,magics)->Tuple[bool,str]:
return False,''
def on_after_compile(self,returncode,binfile,magics)->bool:
return False
def on_before_exec(self,code,magics)->Tuple[bool,str]:
return False,''
def on_after_exec(self,returncode,srcfile,magics)->bool:
return False
def on_after_completion(self,returncode,execfile,magics)->bool:
return False
def templatehander(self,key, value,magics,line):
# self.kobj._write_to_stdout(value+"\n")
newline=line
index1=line.find('//%')
if len(value)>0:
magics[key] =value.split(" ",1)
else:
magics[key] =None
return ''
templatefile=magics['templatefile'][0]
if len(magics['templatefile'])>1:
argsstr=magics['templatefile'][1]
templateargsdict=self.kobj.resolving_eqval2dict(argsstr)
else:
templateargsdict=None
if len(magics['templatefile'])>0:
newline=self.readtemplatefile(self,templatefile,index1,templateargsdict)
return newline + '\n'
def readtemplatefile(self,filename,spacecount=0,*args: t.Any, **kwargs: t.Any):
filecode=''
newfilecode=''
codelist1=None
filenm=os.path.join(os.path.abspath(''),filename);
if not os.path.exists(filenm):
return filecode;
template = self.jinja2_env.get_template(filenm)
filecode=template.render(*args,**kwargs)
for line in filecode.splitlines():
if len(line)>0:
for t in line:
newfilecode+=' '*spacecount + t+'\n'
return newfilecode
| plugins/_Stemplatefile.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
# %matplotlib notebook
# To use this notebook, make sure to copy the two .parquest.gzip files from Google Drive
# (Team Transpo / Datasets & Notebooks / Toad (Congestion) / toad_stop_to_stop_durations )
# and put them in the <data science repo> / data / interim / TOAD folder.
# (you will need to make the TOAD folder if it doesn't exist.)
# Or change the path to point to them however you want :)
# -
# stop_to_stop is all the bus stops that either stop at a stop or pass by a stop (basically not disturbance stops)
stops_df = pd.read_parquet('../data/interim/TOAD/stop_to_stop.parquet.gzip')
# routes is the progression of stops along a route
routes_df = pd.read_parquet('../data/interim/TOAD/route_stops.parquet.gzip')
# There's ~ 54 million here!
stops_df.info()
routes_df.info()
# +
# I chose route 35 for this
route = 35
# stop_to_stop has the current stop and previous stop on the same line to get the duration between them.
# This creates the same structure.
line = routes_df[routes_df['rte'] == route].copy()
line['prev_stop_id'] = line['stop_id'].shift(1)
line.fillna(0, inplace=True)
line['prev_stop_id'] = line['prev_stop_id'].astype('uint16')
line.head()
# +
# Edit these to pick different stops, and adjust histogram.
row_num = 13
max_duration = 200
num_bins = 25
# Essentially this pulls all the occurances of stop to stop which gives us a distribution of those durations.
S = line.iloc[row_num][['stop_id', 'prev_stop_id']]
line_stops = stops_df[(stops_df['stop_id'] == S['stop_id']) & (stops_df['prev_stop_id'] == S['prev_stop_id'])]
ax = line_stops[line_stops['elapsed_time_seconds'] < max_duration].hist('elapsed_time_seconds', bins=num_bins)
xlabel = "Arrival to Arrival Duration [s]"
ylabel = "Count"
title = f"Line 35 Stops {S['prev_stop_id']} to {S['stop_id']}"
_ = ax[0][0].set(title=title, xlabel=xlabel, ylabel=ylabel)
ax[0][0].get_figure().set_size_inches((9.8,6))
# -
| notebooks/5.0-jab-stop-to-stop-analysis.ipynb |
/ ---
/ jupyter:
/ jupytext:
/ text_representation:
/ extension: .q
/ format_name: light
/ format_version: '1.5'
/ jupytext_version: 1.14.4
/ kernelspec:
/ display_name: SQL
/ language: sql
/ name: SQL
/ ---
/ + [markdown] azdata_cell_guid="8461e664-47ea-4734-a571-c227a8a8e504"
/ # Step 1: Create a view on the data
/
/ Analyze the data from the ORACLE table and create a view on top of it. The users don't have to know the data comes from Oracle. It just looks like it comes from SQL Server.
/ + azdata_cell_guid="32b3ab87-f3ce-49ab-9aec-f82b9de6b88e"
USE [TutorialDB]
GO
DROP VIEW IF EXISTS rental_data;
GO
CREATE VIEW rental_data AS
SELECT * FROM oracle.rental_data;
GO
/ + [markdown] azdata_cell_guid="a53ea1a5-41f3-492c-bdea-c4f6b055982c"
/ # Step 2: Train and generate a model
/
/ Create a table to store the model, create a procedure to generate the model based on prediction data, and then execute the procedure and store the results of the model (binary) into the table. The source of the data is the view that uses Oracle as a base.
/ + azdata_cell_guid="b44ddbb8-0b01-4d14-97c5-98341ec7268e"
USE TutorialDB
GO
DROP TABLE IF EXISTS rental_py_models;
GO
CREATE TABLE rental_py_models (
model_name VARCHAR(30) NOT NULL DEFAULT('default model') PRIMARY KEY,
model VARBINARY(MAX) NOT NULL
);
GO
-- Stored procedure that trains and generates a Python model using the rental_data and a decision tree algorithm
DROP PROCEDURE IF EXISTS generate_rental_py_model;
go
CREATE PROCEDURE generate_rental_py_model (@trained_model varbinary(max) OUTPUT)
AS
BEGIN
EXECUTE sp_execute_external_script
@language = N'Python'
, @script = N'
from sklearn.linear_model import LinearRegression
import pickle
df = rental_train_data
# Get all the columns from the dataframe.
columns = df.columns.tolist()
# Store the variable well be predicting on.
target = "RentalCount"
# Initialize the model class.
lin_model = LinearRegression()
# Fit the model to the training data.
lin_model.fit(df[columns], df[target])
#Before saving the model to the DB table, we need to convert it to a binary object
trained_model = pickle.dumps(lin_model)'
, @input_data_1 = N'select "RentalCount", "Year", "Month", "Day", "WeekDay", "Snow", "Holiday" from dbo.rental_data where Year < 2019'
, @input_data_1_name = N'rental_train_data'
, @params = N'@trained_model varbinary(max) OUTPUT'
, @trained_model = @trained_model OUTPUT;
END;
GO
TRUNCATE TABLE rental_py_models;
GO
DECLARE @model VARBINARY(MAX);
EXEC generate_rental_py_model @model OUTPUT;
INSERT INTO rental_py_models (model_name, model) VALUES('linear_model', @model);
GO
/ + [markdown] azdata_cell_guid="032f309b-d8a8-4f07-b6a8-9abe99a3169f"
/ # Step 3: Create objects to run the prediction model and store the results
/
/ Create a procedure and table to store the results of the prediction
/ + azdata_cell_guid="3c84a3e5-ce41-4d25-91ef-9e922691d56b"
DROP PROCEDURE IF EXISTS py_predict_rentalcount;
GO
CREATE PROCEDURE py_predict_rentalcount (@model varchar(100))
AS
BEGIN
DECLARE @py_model varbinary(max) = (select model from rental_py_models where model_name = @model);
EXEC sp_execute_external_script
@language = N'Python',
@script = N'
# Import the scikit-learn function to compute error.
from sklearn.metrics import mean_squared_error
import pickle
import pandas as pd
rental_model = pickle.loads(py_model)
df = rental_score_data
# Get all the columns from the dataframe.
columns = df.columns.tolist()
# variable we will be predicting on.
target = "RentalCount"
# Generate our predictions for the test set.
lin_predictions = rental_model.predict(df[columns])
print(lin_predictions)
# Compute error between our test predictions and the actual values.
lin_mse = mean_squared_error(lin_predictions, df[target])
#print(lin_mse)
predictions_df = pd.DataFrame(lin_predictions)
OutputDataSet = pd.concat([predictions_df, df["RentalCount"], df["Month"], df["Day"], df["WeekDay"], df["Snow"], df["Holiday"], df["Year"]], axis=1)
'
, @input_data_1 = N'Select "RentalCount", "Year" ,"Month", "Day", "WeekDay", "Snow", "Holiday" from rental_data where Year = 2019'
, @input_data_1_name = N'rental_score_data'
, @params = N'@py_model varbinary(max)'
, @py_model = @py_model
with result sets (("RentalCount_Predicted" float, "RentalCount" float, "Month" float,"Day" float,"WeekDay" float,"Snow" float,"Holiday" float, "Year" float));
END;
GO
DROP TABLE IF EXISTS [dbo].[py_rental_predictions];
GO
--Create a table to store the predictions in
CREATE TABLE [dbo].[py_rental_predictions](
[RentalCount_Predicted] [int] NULL,
[RentalCount_Actual] [int] NULL,
[Month] [int] NULL,
[Day] [int] NULL,
[WeekDay] [int] NULL,
[Snow] [int] NULL,
[Holiday] [int] NULL,
[Year] [int] NULL
) ON [PRIMARY];
GO
/ + [markdown] azdata_cell_guid="70ab9d0f-aa40-4a2a-a92e-807bedf039fa"
/ # Step 4: Run the prediction model, store and display the results
/
/ Run the procedure to run the prediction model and store the results in a table. Query out the results.
/ + azdata_cell_guid="593d03fa-5a77-4841-9e1e-8f79b54c021b"
USE TutorialDB;
GO
TRUNCATE TABLE py_rental_predictions;
--Insert the results of the predictions for test set into a table
INSERT INTO py_rental_predictions
EXEC py_predict_rentalcount 'linear_model';
GO
-- Select contents of the table
SELECT * FROM py_rental_predictions;
GO
| demos/build2020/sqllinux/rental_predictions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 0.688297, "end_time": "2021-10-01T06:39:41.529346", "exception": false, "start_time": "2021-10-01T06:39:40.841049", "status": "completed"} tags=[]
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import spacy
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# You can write up to 20GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
# + [markdown] papermill={"duration": 0.017895, "end_time": "2021-10-01T06:39:41.567499", "exception": false, "start_time": "2021-10-01T06:39:41.549604", "status": "completed"} tags=[]
# **Read the Tweets**
# + papermill={"duration": 2.693293, "end_time": "2021-10-01T06:39:44.274036", "exception": false, "start_time": "2021-10-01T06:39:41.580743", "status": "completed"} tags=[]
tweets = pd.read_csv('../input/covid19-tweets/covid19_tweets.csv')
# + papermill={"duration": 0.041963, "end_time": "2021-10-01T06:39:44.328325", "exception": false, "start_time": "2021-10-01T06:39:44.286362", "status": "completed"} tags=[]
tweets.head()
# + [markdown] papermill={"duration": 0.012928, "end_time": "2021-10-01T06:39:44.354134", "exception": false, "start_time": "2021-10-01T06:39:44.341206", "status": "completed"} tags=[]
# **Load the pre-trained model**
# + papermill={"duration": 1.355268, "end_time": "2021-10-01T06:39:45.722562", "exception": false, "start_time": "2021-10-01T06:39:44.367294", "status": "completed"} tags=[]
nlp = spacy.load("en_core_web_sm")
# + [markdown] papermill={"duration": 0.012312, "end_time": "2021-10-01T06:39:45.747908", "exception": false, "start_time": "2021-10-01T06:39:45.735596", "status": "completed"} tags=[]
# **Extract text from Pandas Colum**
# + papermill={"duration": 0.024141, "end_time": "2021-10-01T06:39:45.784644", "exception": false, "start_time": "2021-10-01T06:39:45.760503", "status": "completed"} tags=[]
text=str(tweets['text'])
# + papermill={"duration": 0.073369, "end_time": "2021-10-01T06:39:45.871095", "exception": false, "start_time": "2021-10-01T06:39:45.797726", "status": "completed"} tags=[]
doc = nlp(text)
# + [markdown] papermill={"duration": 0.012603, "end_time": "2021-10-01T06:39:45.897367", "exception": false, "start_time": "2021-10-01T06:39:45.884764", "status": "completed"} tags=[]
# **Parts of Speech**
# + papermill={"duration": 0.054786, "end_time": "2021-10-01T06:39:45.965136", "exception": false, "start_time": "2021-10-01T06:39:45.910350", "status": "completed"} tags=[]
#Parts of Speech - pos
for token in doc:
print(token.text, token.pos_)
# + [markdown] papermill={"duration": 0.013446, "end_time": "2021-10-01T06:39:45.993162", "exception": false, "start_time": "2021-10-01T06:39:45.979716", "status": "completed"} tags=[]
# **Named Entity Recognition**
# + papermill={"duration": 0.023662, "end_time": "2021-10-01T06:39:46.030399", "exception": false, "start_time": "2021-10-01T06:39:46.006737", "status": "completed"} tags=[]
spacy.displacy.render(doc,style='ent',jupyter=True)
# + [markdown] papermill={"duration": 0.01394, "end_time": "2021-10-01T06:39:46.058729", "exception": false, "start_time": "2021-10-01T06:39:46.044789", "status": "completed"} tags=[]
# **Extracting a particular entity type**
# + papermill={"duration": 0.022796, "end_time": "2021-10-01T06:39:46.096267", "exception": false, "start_time": "2021-10-01T06:39:46.073471", "status": "completed"} tags=[]
for ent in doc.ents:
if ent.label_ == 'PERSON':
print(ent.text,ent.label_)
# + papermill={"duration": 0.014088, "end_time": "2021-10-01T06:39:46.125096", "exception": false, "start_time": "2021-10-01T06:39:46.111008", "status": "completed"} tags=[]
| basic-nlp-with-covid-tweets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
with open('input.txt') as f:
data = f.read().splitlines()
ingredients = [set(line.split(' (contains ')[0].split(' ')) for line in data]
allergens = [line.split(' (contains ')[1].replace(')', '').split(', ') for line in data]
all_ingredients = [_ for line in ingredients for _ in line]
# +
d = {}
for i, a in zip(ingredients, allergens):
for k in a:
if k in d:
d[k] = d[k].intersection(i)
else:
d[k] = i
probable_allergen_ingredient = set.union(*list(d.values()))
no_allergen_ingredients = [i for i in all_ingredients if i not in probable_allergen_ingredient]
print(f"Answer part one: {len(no_allergen_ingredients)}")
# +
dd = {}
while True:
for k, v in d.items():
tmp = [_ for _ in list(v) if _ not in dd]
if len(tmp) == 1:
dd[tmp[0]] = k
if all(k in dd for k in probable_allergen_ingredient):
break
dd_sorted = {k: v for k, v in sorted(dd.items(), key=lambda item: item[1])}
can_in = list(dd_sorted.keys())
print(f"Answer part two: {','.join(can_in)}")
| day_21/Day 21 - Allergen Assessment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext watermark
# %watermark -v -p numpy,sklearn,scipy,matplotlib,tensorflow
# **15장 – 오토인코더**
# _이 노트북은 15장에 있는 모든 샘플 코드와 연습문제 해답을 가지고 있습니다._
# # 설정
# 파이썬 2와 3을 모두 지원합니다. 공통 모듈을 임포트하고 맷플롯립 그림이 노트북 안에 포함되도록 설정하고 생성한 그림을 저장하기 위한 함수를 준비합니다:
# +
# 파이썬 2와 파이썬 3 지원
from __future__ import division, print_function, unicode_literals
# 공통
import numpy as np
import os
import sys
# 일관된 출력을 위해 유사난수 초기화
def reset_graph(seed=42):
tf.reset_default_graph()
tf.set_random_seed(seed)
np.random.seed(seed)
# 맷플롯립 설정
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
plt.rcParams['axes.labelsize'] = 14
plt.rcParams['xtick.labelsize'] = 12
plt.rcParams['ytick.labelsize'] = 12
# 한글출력
plt.rcParams['font.family'] = 'NanumBarunGothic'
plt.rcParams['axes.unicode_minus'] = False
# 그림을 저장할 폴더
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "autoencoders"
def save_fig(fig_id, tight_layout=True):
path = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID, fig_id + ".png")
if tight_layout:
plt.tight_layout()
plt.savefig(path, format='png', dpi=300)
# -
# 28x28 흑백 이미지를 그리기 위한 유틸리티 함수:
def plot_image(image, shape=[28, 28]):
plt.imshow(image.reshape(shape), cmap="Greys", interpolation="nearest")
plt.axis("off")
def plot_multiple_images(images, n_rows, n_cols, pad=2):
images = images - images.min() # 최소값을 0으로 만들어 패딩이 하얗게 보이도록 합니다.
w,h = images.shape[1:]
image = np.zeros(((w+pad)*n_rows+pad, (h+pad)*n_cols+pad))
for y in range(n_rows):
for x in range(n_cols):
image[(y*(h+pad)+pad):(y*(h+pad)+pad+h),(x*(w+pad)+pad):(x*(w+pad)+pad+w)] = images[y*n_cols+x]
plt.imshow(image, cmap="Greys", interpolation="nearest")
plt.axis("off")
# # 선형 오토인코더를 사용한 PCA
# 3D 데이터셋을 만듭니다:
# +
import numpy.random as rnd
rnd.seed(4)
m = 200
w1, w2 = 0.1, 0.3
noise = 0.1
angles = rnd.rand(m) * 3 * np.pi / 2 - 0.5
data = np.empty((m, 3))
data[:, 0] = np.cos(angles) + np.sin(angles)/2 + noise * rnd.randn(m) / 2
data[:, 1] = np.sin(angles) * 0.7 + noise * rnd.randn(m) / 2
data[:, 2] = data[:, 0] * w1 + data[:, 1] * w2 + noise * rnd.randn(m)
# -
# 데이터를 정규화합니다:
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
X_train = scaler.fit_transform(data[:100])
X_test = scaler.transform(data[100:])
# 오토인코더를 만듭니다:
# +
import tensorflow as tf
reset_graph()
n_inputs = 3
n_hidden = 2 # 코딩 유닛
n_outputs = n_inputs
learning_rate = 0.01
X = tf.placeholder(tf.float32, shape=[None, n_inputs])
hidden = tf.layers.dense(X, n_hidden)
outputs = tf.layers.dense(hidden, n_outputs)
reconstruction_loss = tf.reduce_mean(tf.square(outputs - X))
optimizer = tf.train.AdamOptimizer(learning_rate)
training_op = optimizer.minimize(reconstruction_loss)
init = tf.global_variables_initializer()
# +
n_iterations = 1000
codings = hidden
with tf.Session() as sess:
init.run()
for iteration in range(n_iterations):
training_op.run(feed_dict={X: X_train})
codings_val = codings.eval(feed_dict={X: X_test})
# -
fig = plt.figure(figsize=(4,3))
plt.plot(codings_val[:,0], codings_val[:, 1], "b.")
plt.xlabel("$z_1$", fontsize=18)
plt.ylabel("$z_2$", fontsize=18, rotation=0)
save_fig("linear_autoencoder_pca_plot")
plt.show()
# # 적층 오토인코더
# MNIST 데이터셋을 사용합니다:
# 주의: `tf.examples.tutorials.mnist`은 삭제될 예정이므로 대신 `tf.keras.datasets.mnist`를 사용하겠습니다.
(X_train, y_train), (X_test, y_test) = tf.keras.datasets.mnist.load_data()
X_train = X_train.astype(np.float32).reshape(-1, 28*28) / 255.0
X_test = X_test.astype(np.float32).reshape(-1, 28*28) / 255.0
y_train = y_train.astype(np.int32)
y_test = y_test.astype(np.int32)
X_valid, X_train = X_train[:5000], X_train[5000:]
y_valid, y_train = y_train[:5000], y_train[5000:]
def shuffle_batch(X, y, batch_size):
rnd_idx = np.random.permutation(len(X))
n_batches = len(X) // batch_size
for batch_idx in np.array_split(rnd_idx, n_batches):
X_batch, y_batch = X[batch_idx], y[batch_idx]
yield X_batch, y_batch
# +
# from tensorflow.examples.tutorials.mnist import input_data
# mnist = input_data.read_data_sets("/tmp/data/")
# -
# ## 한 번에 모든 층을 훈련하기
# 3개의 은닉층과 1개의 출력층(즉, 두 개를 적층)을 가진 적층 오토인코더를 만들어 보겠습니다. ELU 활성화 함수와 He 초기화, L2 정규화를 사용하겠습니다.
# +
reset_graph()
from functools import partial
n_inputs = 28 * 28
n_hidden1 = 300
n_hidden2 = 150 # 코딩 유닛
n_hidden3 = n_hidden1
n_outputs = n_inputs
learning_rate = 0.01
l2_reg = 0.0001
X = tf.placeholder(tf.float32, shape=[None, n_inputs])
he_init = tf.variance_scaling_initializer() # He 초기화
#아래와 동일합니다:
#he_init = lambda shape, dtype=tf.float32: tf.truncated_normal(shape, 0., stddev=np.sqrt(2/shape[0]))
l2_regularizer = tf.contrib.layers.l2_regularizer(l2_reg)
my_dense_layer = partial(tf.layers.dense,
activation=tf.nn.elu,
kernel_initializer=he_init,
kernel_regularizer=l2_regularizer)
hidden1 = my_dense_layer(X, n_hidden1)
hidden2 = my_dense_layer(hidden1, n_hidden2)
hidden3 = my_dense_layer(hidden2, n_hidden3)
outputs = my_dense_layer(hidden3, n_outputs, activation=None)
reconstruction_loss = tf.reduce_mean(tf.square(outputs - X))
reg_losses = tf.get_collection(tf.GraphKeys.REGULARIZATION_LOSSES)
loss = tf.add_n([reconstruction_loss] + reg_losses)
optimizer = tf.train.AdamOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
init = tf.global_variables_initializer()
saver = tf.train.Saver() # 책에는 없음
# -
# 이제 훈련시켜 보죠! 여기에서는 타깃 값을 주입하지 않습니다(`y_batch`가 사용되지 않습니다). 이는 비지도 학습입니다.
# +
n_epochs = 5
batch_size = 150
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
n_batches = len(X_train) // batch_size
for iteration in range(n_batches):
print("\r{}%".format(100 * iteration // n_batches), end="") # 책에는 없음
sys.stdout.flush() # 책에는 없음
X_batch, y_batch = next(shuffle_batch(X_train, y_train, batch_size))
sess.run(training_op, feed_dict={X: X_batch})
loss_train = reconstruction_loss.eval(feed_dict={X: X_batch}) # 책에는 없음
print("\r{}".format(epoch), "훈련 MSE:", loss_train) # 책에는 없음
saver.save(sess, "./my_model_all_layers.ckpt") # 책에는 없음
# -
# 이 함수는 모델을 로드하고 테스트 세트에서 이를 평가합니다(재구성 오차를 측정합니다). 그런 다음 원본 이미지와 재구성 이미지를 그립니다:
def show_reconstructed_digits(X, outputs, model_path = None, n_test_digits = 2):
with tf.Session() as sess:
if model_path:
saver.restore(sess, model_path)
# X_test = mnist.test.images[:n_test_digits]
outputs_val = outputs.eval(feed_dict={X: X_test[:n_test_digits]})
fig = plt.figure(figsize=(8, 3 * n_test_digits))
for digit_index in range(n_test_digits):
plt.subplot(n_test_digits, 2, digit_index * 2 + 1)
plot_image(X_test[digit_index])
plt.subplot(n_test_digits, 2, digit_index * 2 + 2)
plot_image(outputs_val[digit_index])
show_reconstructed_digits(X, outputs, "./my_model_all_layers.ckpt")
save_fig("reconstruction_plot")
# ## 가중치 묶기
# 인코더와 디코더의 가중치를 묶는 일은 자주 있습니다(`weights_decoder = tf.transpose(weights_encoder)`). 안타깝지만 `tf.layers.dense()` 함수를 사용해서 이렇게 하기는 불가능합니다(또는 매우 어렵습니다). 수동으로 직접 오토인코더를 만들어야 합니다:
# +
reset_graph()
n_inputs = 28 * 28
n_hidden1 = 300
n_hidden2 = 150 # 코딩 유닛
n_hidden3 = n_hidden1
n_outputs = n_inputs
learning_rate = 0.01
l2_reg = 0.0005
# +
activation = tf.nn.elu
regularizer = tf.contrib.layers.l2_regularizer(l2_reg)
initializer = tf.variance_scaling_initializer()
X = tf.placeholder(tf.float32, shape=[None, n_inputs])
weights1_init = initializer([n_inputs, n_hidden1])
weights2_init = initializer([n_hidden1, n_hidden2])
weights1 = tf.Variable(weights1_init, dtype=tf.float32, name="weights1")
weights2 = tf.Variable(weights2_init, dtype=tf.float32, name="weights2")
weights3 = tf.transpose(weights2, name="weights3") # 가중치 묶기
weights4 = tf.transpose(weights1, name="weights4") # 가중치 묶기
biases1 = tf.Variable(tf.zeros(n_hidden1), name="biases1")
biases2 = tf.Variable(tf.zeros(n_hidden2), name="biases2")
biases3 = tf.Variable(tf.zeros(n_hidden3), name="biases3")
biases4 = tf.Variable(tf.zeros(n_outputs), name="biases4")
hidden1 = activation(tf.matmul(X, weights1) + biases1)
hidden2 = activation(tf.matmul(hidden1, weights2) + biases2)
hidden3 = activation(tf.matmul(hidden2, weights3) + biases3)
outputs = tf.matmul(hidden3, weights4) + biases4
reconstruction_loss = tf.reduce_mean(tf.square(outputs - X))
reg_loss = regularizer(weights1) + regularizer(weights2)
loss = reconstruction_loss + reg_loss
optimizer = tf.train.AdamOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
init = tf.global_variables_initializer()
# -
saver = tf.train.Saver()
# +
n_epochs = 5
batch_size = 150
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
n_batches = len(X_train) // batch_size
for iteration in range(n_batches):
print("\r{}%".format(100 * iteration // n_batches), end="")
sys.stdout.flush()
X_batch, y_batch = next(shuffle_batch(X_train, y_train, batch_size))
sess.run(training_op, feed_dict={X: X_batch})
loss_train = reconstruction_loss.eval(feed_dict={X: X_batch})
print("\r{}".format(epoch), "훈련 MSE:", loss_train)
saver.save(sess, "./my_model_tying_weights.ckpt")
# -
show_reconstructed_digits(X, outputs, "./my_model_tying_weights.ckpt")
# ## 여러 개의 그래프에서 오토인토더를 따로따로 훈련하기
# 하나의 오토인코더를 따로따로 훈련하는 방법이 많이 있습니다. 첫 번째 방법은 각 오토인코더를 다른 그래프를 사용하여 훈련하는 것입니다. 그런 다음 이런 오토인코더의 가중치와 편향을 복사해 초깃값으로 지정해서 적층 오토인코더를 만듭니다.
# 하나의 오토인코더를 훈련하고 변환된 훈련 세트(즉, 은닉층의 출력)와 모델 파라미터를 반환하는 함수를 만들겠습니다.
# +
reset_graph()
from functools import partial
def train_autoencoder(X_train, n_neurons, n_epochs, batch_size,
learning_rate = 0.01, l2_reg = 0.0005, seed=42,
hidden_activation=tf.nn.elu,
output_activation=tf.nn.elu):
graph = tf.Graph()
with graph.as_default():
tf.set_random_seed(seed)
n_inputs = X_train.shape[1]
X = tf.placeholder(tf.float32, shape=[None, n_inputs])
my_dense_layer = partial(
tf.layers.dense,
kernel_initializer=tf.variance_scaling_initializer(),
kernel_regularizer=tf.contrib.layers.l2_regularizer(l2_reg))
hidden = my_dense_layer(X, n_neurons, activation=hidden_activation, name="hidden")
outputs = my_dense_layer(hidden, n_inputs, activation=output_activation, name="outputs")
reconstruction_loss = tf.reduce_mean(tf.square(outputs - X))
reg_losses = tf.get_collection(tf.GraphKeys.REGULARIZATION_LOSSES)
loss = tf.add_n([reconstruction_loss] + reg_losses)
optimizer = tf.train.AdamOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
init = tf.global_variables_initializer()
with tf.Session(graph=graph) as sess:
init.run()
for epoch in range(n_epochs):
n_batches = len(X_train) // batch_size
for iteration in range(n_batches):
print("\r{}%".format(100 * iteration // n_batches), end="")
sys.stdout.flush()
indices = rnd.permutation(len(X_train))[:batch_size]
X_batch = X_train[indices]
sess.run(training_op, feed_dict={X: X_batch})
loss_train = reconstruction_loss.eval(feed_dict={X: X_batch})
print("\r{}".format(epoch), "훈련 MSE:", loss_train)
params = dict([(var.name, var.eval()) for var in tf.get_collection(tf.GraphKeys.TRAINABLE_VARIABLES)])
hidden_val = hidden.eval(feed_dict={X: X_train})
return hidden_val, params["hidden/kernel:0"], params["hidden/bias:0"], params["outputs/kernel:0"], params["outputs/bias:0"]
# -
# 이제 두 개의 오토인코더를 훈련시켜 보죠. 첫 번째는 훈련 데이터를 사용하고 두 번째는 첫 번째 오토인코더의 은닉층 출력을 사용해 훈련시킵니다:
hidden_output, W1, b1, W4, b4 = train_autoencoder(X_train, n_neurons=300, n_epochs=4, batch_size=150,
output_activation=None)
_, W2, b2, W3, b3 = train_autoencoder(hidden_output, n_neurons=150, n_epochs=4, batch_size=150)
# 마지막으로 방금전 훈련한 오토인코더의 가중치와 편향을 재사용하여 적층 오토인코더를 만듭니다:
# +
reset_graph()
n_inputs = 28*28
X = tf.placeholder(tf.float32, shape=[None, n_inputs])
hidden1 = tf.nn.elu(tf.matmul(X, W1) + b1)
hidden2 = tf.nn.elu(tf.matmul(hidden1, W2) + b2)
hidden3 = tf.nn.elu(tf.matmul(hidden2, W3) + b3)
outputs = tf.matmul(hidden3, W4) + b4
# -
show_reconstructed_digits(X, outputs)
# ## 하나의 그래프에서 오토인코더를 따로따로 훈련하기
# 하나의 그래프를 사용하는 방법도 있습니다. 이 방법은 전체 적층 오토인코더를 위한 그래프를 만들지만 각 오토인코더를 독립적으로 훈련하기 위한 연산도 추가합니다. 단계 1은 맨 아래층과 맨 윗층을 훈련하고(즉, 첫 번째 오토인코더), 단계 2는 두 개의 가운데 층을 훈련합니다(즉, 두 번째 오토인코더).
# +
reset_graph()
n_inputs = 28 * 28
n_hidden1 = 300
n_hidden2 = 150 # 코딩 유닛
n_hidden3 = n_hidden1
n_outputs = n_inputs
learning_rate = 0.01
l2_reg = 0.0001
activation = tf.nn.elu
regularizer = tf.contrib.layers.l2_regularizer(l2_reg)
initializer = tf.variance_scaling_initializer()
X = tf.placeholder(tf.float32, shape=[None, n_inputs])
weights1_init = initializer([n_inputs, n_hidden1])
weights2_init = initializer([n_hidden1, n_hidden2])
weights3_init = initializer([n_hidden2, n_hidden3])
weights4_init = initializer([n_hidden3, n_outputs])
weights1 = tf.Variable(weights1_init, dtype=tf.float32, name="weights1")
weights2 = tf.Variable(weights2_init, dtype=tf.float32, name="weights2")
weights3 = tf.Variable(weights3_init, dtype=tf.float32, name="weights3")
weights4 = tf.Variable(weights4_init, dtype=tf.float32, name="weights4")
biases1 = tf.Variable(tf.zeros(n_hidden1), name="biases1")
biases2 = tf.Variable(tf.zeros(n_hidden2), name="biases2")
biases3 = tf.Variable(tf.zeros(n_hidden3), name="biases3")
biases4 = tf.Variable(tf.zeros(n_outputs), name="biases4")
hidden1 = activation(tf.matmul(X, weights1) + biases1)
hidden2 = activation(tf.matmul(hidden1, weights2) + biases2)
hidden3 = activation(tf.matmul(hidden2, weights3) + biases3)
outputs = tf.matmul(hidden3, weights4) + biases4
reconstruction_loss = tf.reduce_mean(tf.square(outputs - X))
# +
optimizer = tf.train.AdamOptimizer(learning_rate)
with tf.name_scope("phase1"):
phase1_outputs = tf.matmul(hidden1, weights4) + biases4 # hidden2와 hidden3 통과합니다
phase1_reconstruction_loss = tf.reduce_mean(tf.square(phase1_outputs - X))
phase1_reg_loss = regularizer(weights1) + regularizer(weights4)
phase1_loss = phase1_reconstruction_loss + phase1_reg_loss
phase1_training_op = optimizer.minimize(phase1_loss)
with tf.name_scope("phase2"):
phase2_reconstruction_loss = tf.reduce_mean(tf.square(hidden3 - hidden1))
phase2_reg_loss = regularizer(weights2) + regularizer(weights3)
phase2_loss = phase2_reconstruction_loss + phase2_reg_loss
train_vars = [weights2, biases2, weights3, biases3]
phase2_training_op = optimizer.minimize(phase2_loss, var_list=train_vars) # hidden1 동결
# -
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# +
training_ops = [phase1_training_op, phase2_training_op]
reconstruction_losses = [phase1_reconstruction_loss, phase2_reconstruction_loss]
n_epochs = [4, 4]
batch_sizes = [150, 150]
with tf.Session() as sess:
init.run()
for phase in range(2):
print("훈련 단계 #{}".format(phase + 1))
for epoch in range(n_epochs[phase]):
n_batches = len(X_train) // batch_sizes[phase]
for iteration in range(n_batches):
print("\r{}%".format(100 * iteration // n_batches), end="")
sys.stdout.flush()
X_batch, y_batch = next(shuffle_batch(X_train, y_train, batch_sizes[phase]))
sess.run(training_ops[phase], feed_dict={X: X_batch})
loss_train = reconstruction_losses[phase].eval(feed_dict={X: X_batch})
print("\r{}".format(epoch), "훈련 MSE:", loss_train)
saver.save(sess, "./my_model_one_at_a_time.ckpt")
loss_test = reconstruction_loss.eval(feed_dict={X: X_test})
print("테스트 MSE:", loss_test)
# -
# ## 동결 층의 출력을 캐싱하기
# +
training_ops = [phase1_training_op, phase2_training_op]
reconstruction_losses = [phase1_reconstruction_loss, phase2_reconstruction_loss]
n_epochs = [4, 4]
batch_sizes = [150, 150]
with tf.Session() as sess:
init.run()
for phase in range(2):
print("훈련 단계 #{}".format(phase + 1))
if phase == 1:
hidden1_cache = hidden1.eval(feed_dict={X: X_train})
for epoch in range(n_epochs[phase]):
n_batches = len(X_train) // batch_sizes[phase]
for iteration in range(n_batches):
print("\r{}%".format(100 * iteration // n_batches), end="")
sys.stdout.flush()
if phase == 1:
indices = rnd.permutation(len(X_train))
hidden1_batch = hidden1_cache[indices[:batch_sizes[phase]]]
feed_dict = {hidden1: hidden1_batch}
sess.run(training_ops[phase], feed_dict=feed_dict)
else:
X_batch, y_batch = next(shuffle_batch(X_train, y_train, batch_sizes[phase]))
feed_dict = {X: X_batch}
sess.run(training_ops[phase], feed_dict=feed_dict)
loss_train = reconstruction_losses[phase].eval(feed_dict=feed_dict)
print("\r{}".format(epoch), "훈련 MSE:", loss_train)
saver.save(sess, "./my_model_cache_frozen.ckpt")
loss_test = reconstruction_loss.eval(feed_dict={X: X_test})
print("테스트 MSE:", loss_test)
# -
# ## 재구성 시각화
# +
n_test_digits = 2
# X_test = mnist.test.images[:n_test_digits]
with tf.Session() as sess:
saver.restore(sess, "./my_model_one_at_a_time.ckpt") # not shown in the book
outputs_val = outputs.eval(feed_dict={X: X_test[:n_test_digits]})
def plot_image(image, shape=[28, 28]):
plt.imshow(image.reshape(shape), cmap="Greys", interpolation="nearest")
plt.axis("off")
for digit_index in range(n_test_digits):
plt.subplot(n_test_digits, 2, digit_index * 2 + 1)
plot_image(X_test[digit_index])
plt.subplot(n_test_digits, 2, digit_index * 2 + 2)
plot_image(outputs_val[digit_index])
# -
# ## 특성 시각화
# +
with tf.Session() as sess:
saver.restore(sess, "./my_model_one_at_a_time.ckpt") # 책에는 없음
weights1_val = weights1.eval()
for i in range(5):
plt.subplot(1, 5, i + 1)
plot_image(weights1_val.T[i])
save_fig("extracted_features_plot") # 책에는 없음
plt.show() # 책에는 없음
# -
# # 비지도 사전훈련
# MNIST 분류 문제를 위한 작은 신경망을 만들겠습니다:
# +
reset_graph()
n_inputs = 28 * 28
n_hidden1 = 300
n_hidden2 = 150
n_outputs = 10
learning_rate = 0.01
l2_reg = 0.0005
activation = tf.nn.elu
regularizer = tf.contrib.layers.l2_regularizer(l2_reg)
initializer = tf.variance_scaling_initializer()
X = tf.placeholder(tf.float32, shape=[None, n_inputs])
y = tf.placeholder(tf.int32, shape=[None])
weights1_init = initializer([n_inputs, n_hidden1])
weights2_init = initializer([n_hidden1, n_hidden2])
weights3_init = initializer([n_hidden2, n_outputs])
weights1 = tf.Variable(weights1_init, dtype=tf.float32, name="weights1")
weights2 = tf.Variable(weights2_init, dtype=tf.float32, name="weights2")
weights3 = tf.Variable(weights3_init, dtype=tf.float32, name="weights3")
biases1 = tf.Variable(tf.zeros(n_hidden1), name="biases1")
biases2 = tf.Variable(tf.zeros(n_hidden2), name="biases2")
biases3 = tf.Variable(tf.zeros(n_outputs), name="biases3")
hidden1 = activation(tf.matmul(X, weights1) + biases1)
hidden2 = activation(tf.matmul(hidden1, weights2) + biases2)
logits = tf.matmul(hidden2, weights3) + biases3
cross_entropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
reg_loss = regularizer(weights1) + regularizer(weights2) + regularizer(weights3)
loss = cross_entropy + reg_loss
optimizer = tf.train.AdamOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32))
init = tf.global_variables_initializer()
pretrain_saver = tf.train.Saver([weights1, weights2, biases1, biases2])
saver = tf.train.Saver()
# -
# (사전훈련 없이)평범하게 훈련시킵니다:
# +
n_epochs = 4
batch_size = 150
n_labeled_instances = 20000
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
n_batches = n_labeled_instances // batch_size
for iteration in range(n_batches):
print("\r{}%".format(100 * iteration // n_batches), end="")
sys.stdout.flush()
indices = rnd.permutation(n_labeled_instances)[:batch_size]
X_batch, y_batch = X_train[indices], y_train[indices]
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: X_batch, y: y_batch})
print("\r{}".format(epoch), "검증 세트 정확도:", accuracy_val, end=" ")
saver.save(sess, "./my_model_supervised.ckpt")
test_val = accuracy.eval(feed_dict={X: X_test, y: y_test})
print("테스트 정확도:", test_val)
# -
# 사전 훈련된 오토인코더의 첫 두개의 층을 재사용해 보겠습니다:
# +
n_epochs = 4
batch_size = 150
n_labeled_instances = 20000
#training_op = optimizer.minimize(loss, var_list=[weights3, biases3]) # layers 1와 2를 동결 (선택사항)
with tf.Session() as sess:
init.run()
pretrain_saver.restore(sess, "./my_model_cache_frozen.ckpt")
for epoch in range(n_epochs):
n_batches = n_labeled_instances // batch_size
for iteration in range(n_batches):
print("\r{}%".format(100 * iteration // n_batches), end="")
sys.stdout.flush()
indices = rnd.permutation(n_labeled_instances)[:batch_size]
X_batch, y_batch = X_train[indices], y_train[indices]
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: X_batch, y: y_batch})
print("\r{}".format(epoch), "훈련 정확도:", accuracy_val, end="\t")
saver.save(sess, "./my_model_supervised_pretrained.ckpt")
test_val = accuracy.eval(feed_dict={X: X_test, y: y_test})
print("테스트 정확도:", test_val)
# -
# # 적층 잡음제거 오토인코더
# 가우시안 잡음을 사용합니다:
# +
reset_graph()
n_inputs = 28 * 28
n_hidden1 = 300
n_hidden2 = 150 # 코딩 유닛
n_hidden3 = n_hidden1
n_outputs = n_inputs
learning_rate = 0.01
# +
noise_level = 1.0
X = tf.placeholder(tf.float32, shape=[None, n_inputs])
X_noisy = X + noise_level * tf.random_normal(tf.shape(X))
hidden1 = tf.layers.dense(X_noisy, n_hidden1, activation=tf.nn.relu,
name="hidden1")
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=tf.nn.relu, # 책에는 없음
name="hidden2") # 책에는 없음
hidden3 = tf.layers.dense(hidden2, n_hidden3, activation=tf.nn.relu, # 책에는 없음
name="hidden3") # 책에는 없음
outputs = tf.layers.dense(hidden3, n_outputs, name="outputs") # 책에는 없음
reconstruction_loss = tf.reduce_mean(tf.square(outputs - X)) # MSE
# +
optimizer = tf.train.AdamOptimizer(learning_rate)
training_op = optimizer.minimize(reconstruction_loss)
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# +
n_epochs = 10
batch_size = 150
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
n_batches = len(X_train) // batch_size
for iteration in range(n_batches):
print("\r{}%".format(100 * iteration // n_batches), end="")
sys.stdout.flush()
X_batch, y_batch = next(shuffle_batch(X_train, y_train, batch_size))
sess.run(training_op, feed_dict={X: X_batch})
loss_train = reconstruction_loss.eval(feed_dict={X: X_batch})
print("\r{}".format(epoch), "훈련 MSE:", loss_train)
saver.save(sess, "./my_model_stacked_denoising_gaussian.ckpt")
# -
# 드롭아웃을 사용합니다:
# +
reset_graph()
n_inputs = 28 * 28
n_hidden1 = 300
n_hidden2 = 150 # 코딩 유닛
n_hidden3 = n_hidden1
n_outputs = n_inputs
learning_rate = 0.01
# +
dropout_rate = 0.3
training = tf.placeholder_with_default(False, shape=(), name='training')
X = tf.placeholder(tf.float32, shape=[None, n_inputs])
X_drop = tf.layers.dropout(X, dropout_rate, training=training)
hidden1 = tf.layers.dense(X_drop, n_hidden1, activation=tf.nn.relu,
name="hidden1")
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=tf.nn.relu, # 책에는 없음
name="hidden2") # 책에는 없음
hidden3 = tf.layers.dense(hidden2, n_hidden3, activation=tf.nn.relu, # 책에는 없음
name="hidden3") # 책에는 없음
outputs = tf.layers.dense(hidden3, n_outputs, name="outputs") # 책에는 없음
reconstruction_loss = tf.reduce_mean(tf.square(outputs - X)) # MSE
# +
optimizer = tf.train.AdamOptimizer(learning_rate)
training_op = optimizer.minimize(reconstruction_loss)
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# +
n_epochs = 10
batch_size = 150
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
n_batches = len(X_train) // batch_size
for iteration in range(n_batches):
print("\r{}%".format(100 * iteration // n_batches), end="")
sys.stdout.flush()
X_batch, y_batch = next(shuffle_batch(X_train, y_train, batch_size))
sess.run(training_op, feed_dict={X: X_batch, training: True})
loss_train = reconstruction_loss.eval(feed_dict={X: X_batch})
print("\r{}".format(epoch), "훈련 MSE:", loss_train)
saver.save(sess, "./my_model_stacked_denoising_dropout.ckpt")
# -
show_reconstructed_digits(X, outputs, "./my_model_stacked_denoising_dropout.ckpt")
# # 희소 오토인코더
p = 0.1
q = np.linspace(0.001, 0.999, 500)
kl_div = p * np.log(p / q) + (1 - p) * np.log((1 - p) / (1 - q))
mse = (p - q)**2
plt.plot([p, p], [0, 0.3], "k:")
plt.text(0.05, 0.32, "목표 희소", fontsize=14)
plt.plot(q, kl_div, "b-", label="쿨백 라이블러 발산")
plt.plot(q, mse, "r--", label="MSE")
plt.legend(loc="upper left")
plt.xlabel("실제 희소")
plt.ylabel("비용", rotation=0)
plt.axis([0, 1, 0, 0.95])
save_fig("sparsity_loss_plot")
# +
reset_graph()
n_inputs = 28 * 28
n_hidden1 = 1000 # 희소 코딩 유닛
n_outputs = n_inputs
# +
def kl_divergence(p, q):
# 쿨백 라이블러 발산
return p * tf.log(p / q) + (1 - p) * tf.log((1 - p) / (1 - q))
learning_rate = 0.01
sparsity_target = 0.1
sparsity_weight = 0.2
X = tf.placeholder(tf.float32, shape=[None, n_inputs]) # 책에는 없음
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.sigmoid) # 책에는 없음
outputs = tf.layers.dense(hidden1, n_outputs) # 책에는 없음
hidden1_mean = tf.reduce_mean(hidden1, axis=0) # 배치 평균
sparsity_loss = tf.reduce_sum(kl_divergence(sparsity_target, hidden1_mean))
reconstruction_loss = tf.reduce_mean(tf.square(outputs - X)) # MSE
loss = reconstruction_loss + sparsity_weight * sparsity_loss
optimizer = tf.train.AdamOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
# -
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# +
n_epochs = 100
batch_size = 1000
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
n_batches = len(X_train) // batch_size
for iteration in range(n_batches):
print("\r{}%".format(100 * iteration // n_batches), end="")
sys.stdout.flush()
X_batch, y_batch = next(shuffle_batch(X_train, y_train, batch_size))
sess.run(training_op, feed_dict={X: X_batch})
reconstruction_loss_val, sparsity_loss_val, loss_val = sess.run([reconstruction_loss, sparsity_loss, loss], feed_dict={X: X_batch})
print("\r{}".format(epoch), "훈련 MSE:", reconstruction_loss_val, "\t희소 손실:", sparsity_loss_val, "\t전체 손실:", loss_val)
saver.save(sess, "./my_model_sparse.ckpt")
# -
show_reconstructed_digits(X, outputs, "./my_model_sparse.ckpt")
# 코딩층은 0에서 1사이의 값을 출력해야 하므로 시그모이드 활성화 함수를 사용합니다:
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.sigmoid)
# 훈련 속도를 높이기 위해 입력을 0과 1사이로 정규화하고 비용 함수로 MSE 대신 크로스엔트로피를 사용합니다:
# +
logits = tf.layers.dense(hidden1, n_outputs)
outputs = tf.nn.sigmoid(logits)
xentropy = tf.nn.sigmoid_cross_entropy_with_logits(labels=X, logits=logits)
reconstruction_loss = tf.reduce_mean(xentropy)
# -
# # 변이형 오토인코더
# +
reset_graph()
from functools import partial
n_inputs = 28 * 28
n_hidden1 = 500
n_hidden2 = 500
n_hidden3 = 20 # 코딩 유닛
n_hidden4 = n_hidden2
n_hidden5 = n_hidden1
n_outputs = n_inputs
learning_rate = 0.001
initializer = tf.variance_scaling_initializer()
my_dense_layer = partial(
tf.layers.dense,
activation=tf.nn.elu,
kernel_initializer=initializer)
X = tf.placeholder(tf.float32, [None, n_inputs])
hidden1 = my_dense_layer(X, n_hidden1)
hidden2 = my_dense_layer(hidden1, n_hidden2)
hidden3_mean = my_dense_layer(hidden2, n_hidden3, activation=None)
hidden3_sigma = my_dense_layer(hidden2, n_hidden3, activation=None)
noise = tf.random_normal(tf.shape(hidden3_sigma), dtype=tf.float32)
hidden3 = hidden3_mean + hidden3_sigma * noise
hidden4 = my_dense_layer(hidden3, n_hidden4)
hidden5 = my_dense_layer(hidden4, n_hidden5)
logits = my_dense_layer(hidden5, n_outputs, activation=None)
outputs = tf.sigmoid(logits)
xentropy = tf.nn.sigmoid_cross_entropy_with_logits(labels=X, logits=logits)
reconstruction_loss = tf.reduce_sum(xentropy)
# -
eps = 1e-10 # NaN을 반환하는 log(0)을 피하기 위한 안전항
latent_loss = 0.5 * tf.reduce_sum(
tf.square(hidden3_sigma) + tf.square(hidden3_mean)
- 1 - tf.log(eps + tf.square(hidden3_sigma)))
# +
loss = reconstruction_loss + latent_loss
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate)
training_op = optimizer.minimize(loss)
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# +
n_epochs = 50
batch_size = 150
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
n_batches = len(X_train) // batch_size
for iteration in range(n_batches):
print("\r{}%".format(100 * iteration // n_batches), end="")
sys.stdout.flush()
X_batch, y_batch = next(shuffle_batch(X_train, y_train, batch_size))
sess.run(training_op, feed_dict={X: X_batch})
loss_val, reconstruction_loss_val, latent_loss_val = sess.run([loss, reconstruction_loss, latent_loss], feed_dict={X: X_batch})
print("\r{}".format(epoch), "훈련 전체 손실:", loss_val, "\t재구성 손실:", reconstruction_loss_val, "\t잠재 손실:", latent_loss_val)
saver.save(sess, "./my_model_variational.ckpt")
# +
reset_graph()
from functools import partial
n_inputs = 28 * 28
n_hidden1 = 500
n_hidden2 = 500
n_hidden3 = 20 # 코딩 유닛
n_hidden4 = n_hidden2
n_hidden5 = n_hidden1
n_outputs = n_inputs
learning_rate = 0.001
initializer = tf.variance_scaling_initializer()
my_dense_layer = partial(
tf.layers.dense,
activation=tf.nn.elu,
kernel_initializer=initializer)
X = tf.placeholder(tf.float32, [None, n_inputs])
hidden1 = my_dense_layer(X, n_hidden1)
hidden2 = my_dense_layer(hidden1, n_hidden2)
hidden3_mean = my_dense_layer(hidden2, n_hidden3, activation=None)
hidden3_gamma = my_dense_layer(hidden2, n_hidden3, activation=None)
noise = tf.random_normal(tf.shape(hidden3_gamma), dtype=tf.float32)
hidden3 = hidden3_mean + tf.exp(0.5 * hidden3_gamma) * noise
hidden4 = my_dense_layer(hidden3, n_hidden4)
hidden5 = my_dense_layer(hidden4, n_hidden5)
logits = my_dense_layer(hidden5, n_outputs, activation=None)
outputs = tf.sigmoid(logits)
xentropy = tf.nn.sigmoid_cross_entropy_with_logits(labels=X, logits=logits)
reconstruction_loss = tf.reduce_sum(xentropy)
latent_loss = 0.5 * tf.reduce_sum(
tf.exp(hidden3_gamma) + tf.square(hidden3_mean) - 1 - hidden3_gamma)
loss = reconstruction_loss + latent_loss
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate)
training_op = optimizer.minimize(loss)
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# -
# ## 숫자 이미지 생성
# 모델을 훈련시켜 랜덤한 이미지를 생성해 보겠습니다:
# +
import numpy as np
n_digits = 60
n_epochs = 50
batch_size = 150
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
n_batches = len(X_train) // batch_size
for iteration in range(n_batches):
print("\r{}%".format(100 * iteration // n_batches), end="") # not shown in the book
sys.stdout.flush() # not shown
X_batch, y_batch = next(shuffle_batch(X_train, y_train, batch_size))
sess.run(training_op, feed_dict={X: X_batch})
loss_val, reconstruction_loss_val, latent_loss_val = sess.run([loss, reconstruction_loss, latent_loss], feed_dict={X: X_batch}) # not shown
print("\r{}".format(epoch), "훈련 전체 손실:", loss_val, "\t재구성 손실:", reconstruction_loss_val, "\t잠재 손실:", latent_loss_val) # not shown
saver.save(sess, "./my_model_variational.ckpt") # not shown
codings_rnd = np.random.normal(size=[n_digits, n_hidden3])
outputs_val = outputs.eval(feed_dict={hidden3: codings_rnd})
# -
plt.figure(figsize=(8,50)) # 책에는 없음
for iteration in range(n_digits):
plt.subplot(n_digits, 10, iteration + 1)
plot_image(outputs_val[iteration])
n_rows = 6
n_cols = 10
plot_multiple_images(outputs_val.reshape(-1, 28, 28), n_rows, n_cols)
save_fig("generated_digits_plot")
plt.show()
# 여기에서 잠재 변수 손실은 조금 다르게 계산된 점을 주목하세요:
latent_loss = 0.5 * tf.reduce_sum(
tf.exp(hidden3_gamma) + tf.square(hidden3_mean) - 1 - hidden3_gamma)
# ## 인코드 & 디코드
# 인코드:
# +
n_digits = 3
# X_test, y_test = mnist.test.next_batch(batch_size)
X_test_batch, y_test_batch = next(shuffle_batch(X_test, y_test, batch_size))
codings = hidden3
with tf.Session() as sess:
saver.restore(sess, "./my_model_variational.ckpt")
codings_val = codings.eval(feed_dict={X: X_test_batch})
# -
# 디코드:
with tf.Session() as sess:
saver.restore(sess, "./my_model_variational.ckpt")
outputs_val = outputs.eval(feed_dict={codings: codings_val})
# 재구성 이미지를 그려봅니다:
fig = plt.figure(figsize=(8, 2.5 * n_digits))
for iteration in range(n_digits):
plt.subplot(n_digits, 2, 1 + 2 * iteration)
plot_image(X_test_batch[iteration])
plt.subplot(n_digits, 2, 2 + 2 * iteration)
plot_image(outputs_val[iteration])
# ## 숫자 이미지 보간
# +
n_iterations = 3
n_digits = 6
codings_rnd = np.random.normal(size=[n_digits, n_hidden3])
with tf.Session() as sess:
saver.restore(sess, "./my_model_variational.ckpt")
target_codings = np.roll(codings_rnd, -1, axis=0)
for iteration in range(n_iterations + 1):
codings_interpolate = codings_rnd + (target_codings - codings_rnd) * iteration / n_iterations
outputs_val = outputs.eval(feed_dict={codings: codings_interpolate})
plt.figure(figsize=(11, 1.5*n_iterations))
for digit_index in range(n_digits):
plt.subplot(1, n_digits, digit_index + 1)
plot_image(outputs_val[digit_index])
plt.show()
# -
# # 연습문제 해답
# Coming soon...
| 15_autoencoders.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # cuSignal Cheat Sheets sample code
#
# (c) 2020 NVIDIA, Blazing SQL
#
# Distributed under Apache License 2.0
# # Imports
# +
import cusignal
import cupy as cp
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# # cuSignal I/O
#
# Data downloaded from: https://sccn.ucsd.edu/~arno/fam2data/publicly_available_EEG_data.html
# #### cusignal.io.reader.read_bin()
# +
data_eeg = cusignal.read_bin(
'../data/sub-01_ses-1_task-BreathCounting_eeg_SMALL.bdf'
, dtype=cp.uint8
, num_samples=50000
, offset=1000
)
plt.figure(figsize=(12,9))
plt.plot(cp.asnumpy(data_eeg[0:500]))
# -
# #### cusignal.io.reader.read_sigmf()
# +
data_wifi = cusignal.read_sigmf(
data_file='../data/Demod_WiFi_cable_X310_3123D76_IQ#1_run1.sigmf-data'
, meta_file='../data/Demod_WiFi_cable_X310_3123D76_IQ#1_run1.sigmf-meta'
, num_samples=50000
, offset=1000
)
plt.figure(figsize=(12,9))
plt.plot(cp.asnumpy(data_wifi[0:500]))
# -
# #### cusignal.io.reader.unpack_bin()
cusignal.unpack_bin(
cusignal.pack_bin(data_eeg)
, dtype=cp.uint8
)
# #### cusignal.io.writer.pack_bin()
cusignal.pack_bin(data_eeg)
# #### cusignal.io.writer.write_bin()
cusignal.write_bin(
'../data/sub-01_ses-1_task-BreathCounting_eeg_SMALL.bdf'
, data_eeg
, append=False
)
# #### cusignal.io.writer.write_sigmf()
cusignal.write_sigmf(
'../data/Demod_WiFi_cable_X310_3123D76_IQ#1_run1_SMALL.sigmf-data'
, data_wifi
, append=False
)
| cheatsheets/cuSignal/cusignal_IO.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # CCDC - Submission Model Training
# ### LIBRARIES
# + tags=[]
# %%capture
# !pip install pandas_path pytorch_lightning cloudpathlib loguru typer
# !pip install --upgrade pandas==1.2.4
# !pip install albumentations
# -
# %load_ext autoreload
# %autoreload 2
# +
import shutil
import rasterio
import pyproj
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from pandas_path import path # noqa
from pathlib import Path
from PIL import Image
import pytorch_lightning as pl
import torch
import albumentations
# + [markdown] tags=[]
# ### DATA
# +
DATA_DIR = Path("/driven-data/cloud-cover")
TRAIN_FEATURES = DATA_DIR / "train_features"
TRAIN_LABELS = DATA_DIR / "train_labels"
assert TRAIN_FEATURES.exists()
# -
BANDS = ["B02", "B03", "B04", "B08"]
# #### Metadata
train_meta = pd.read_csv(DATA_DIR / "train_metadata.csv")
train_meta.head()
# how many different chip ids, locations, and datetimes are there?
train_meta[["chip_id", "location", "datetime"]].nunique()
train_location_counts = (
train_meta.groupby("location")["chip_id"].nunique().sort_values(ascending=False)
)
train_meta["datetime"] = pd.to_datetime(train_meta["datetime"])
train_meta["year"] = train_meta.datetime.dt.year
train_meta.groupby("year")[["chip_id"]].nunique().sort_index().rename(
columns={"chip_id": "chip_count"}
)
train_meta["datetime"].min(), train_meta["datetime"].max()
chips_per_locationtime = (
train_meta.groupby(["location", "datetime"])[["chip_id"]]
.nunique()
.sort_values(by="chip_id", ascending=False)
.rename(columns={"chip_id": "chip_count"})
)
chips_per_locationtime.head(10)
# +
def add_paths(df, feature_dir, label_dir=None, bands=BANDS):
"""
Given dataframe with a column for chip_id, returns a dataframe with a column
added indicating the path to each band's TIF image as "{band}_path", eg "B02_path".
A column is also added to the dataframe with paths to the label TIF, if the
path to the labels directory is provided.
"""
for band in bands:
df[f"{band}_path"] = feature_dir / df["chip_id"] / f"{band}.tif"
# make sure a random sample of paths exist
assert df.sample(n=40, random_state=5)[f"{band}_path"].path.exists().all()
if label_dir is not None:
df["label_path"] = label_dir / (df["chip_id"] + ".tif")
# make sure a random sample of paths exist
assert df.sample(n=40, random_state=5)["label_path"].path.exists().all()
return df
train_meta = add_paths(train_meta, TRAIN_FEATURES, TRAIN_LABELS)
train_meta.head(3)
# -
def lat_long_bounds(filepath):
"""Given the path to a GeoTIFF, returns the image bounds in latitude and
longitude coordinates.
Returns points as a tuple of (left, bottom, right, top)
"""
with rasterio.open(filepath) as im:
bounds = im.bounds
meta = im.meta
# create a converter starting with the current projection
current_crs = pyproj.CRS(meta["crs"])
crs_transform = pyproj.Transformer.from_crs(current_crs, current_crs.geodetic_crs)
# returns left, bottom, right, top
return crs_transform.transform_bounds(*bounds)
def true_color_img(chip_id, data_dir=TRAIN_FEATURES):
"""Given the path to the directory of Sentinel-2 chip feature images,
plots the true color image"""
chip_dir = data_dir / chip_id
red = rioxarray.open_rasterio(chip_dir / "B04.tif").squeeze()
green = rioxarray.open_rasterio(chip_dir / "B03.tif").squeeze()
blue = rioxarray.open_rasterio(chip_dir / "B02.tif").squeeze()
return ms.true_color(r=red, g=green, b=blue)
# ### Train / Test Split
# We have chosen the simplest route, and split our training chips randomly into 1/3 validation and 2/3 training. You may want to think about splitting by location instead of by chip, to better check how your model will do in new settings.
import random
# +
random.seed(9) # set a seed for reproducibility
# put 1/3 of chips into the validation set
chip_ids = train_meta.chip_id.unique().tolist()
val_chip_ids = random.sample(chip_ids, round(len(chip_ids) * 0.33))
val_mask = train_meta.chip_id.isin(val_chip_ids)
val = train_meta[val_mask].copy().reset_index(drop=True)
train = train_meta[~val_mask].copy().reset_index(drop=True)
val.shape, train.shape
# +
# separate features from labels
feature_cols = ["chip_id"] + [f"{band}_path" for band in BANDS]
val_x = val[feature_cols].copy()
val_y = val[["chip_id", "label_path"]].copy()
train_x = train[feature_cols].copy()
train_y = train[["chip_id", "label_path"]].copy()
# -
val_x.head()
val_y.head()
# ### SUBMISSION BY DESIGN
# +
# create benchmark_src folder
submission_dir = Path("train_src")
if submission_dir.exists():
shutil.rmtree(submission_dir)
submission_dir.mkdir(parents=True)
# -
# ### The Model
# +
# %%file {submission_dir}/cloud_dataset.py
import numpy as np
import pandas as pd
import rasterio
import torch
from typing import Optional, List
class CloudDataset(torch.utils.data.Dataset):
"""Reads in images, transforms pixel values, and serves a
dictionary containing chip ids, image tensors, and
label masks (where available).
"""
def __init__(
self,
x_paths: pd.DataFrame,
bands: List[str],
y_paths: Optional[pd.DataFrame] = None,
transforms: Optional[list] = None,
):
"""
Instantiate the CloudDataset class.
Args:
x_paths (pd.DataFrame): a dataframe with a row for each chip. There must be a column for chip_id,
and a column with the path to the TIF for each of bands
bands (list[str]): list of the bands included in the data
y_paths (pd.DataFrame, optional): a dataframe with a for each chip and columns for chip_id
and the path to the label TIF with ground truth cloud cover
transforms (list, optional): list of transforms to apply to the feature data (eg augmentations)
"""
self.data = x_paths
self.label = y_paths
self.transforms = transforms
self.bands = bands
def __len__(self):
return len(self.data)
def __getitem__(self, idx: int):
# Loads an n-channel image from a chip-level dataframe
img = self.data.loc[idx]
band_arrs = []
for band in self.bands:
with rasterio.open(img[f"{band}_path"]) as b:
band_arr = b.read(1).astype("float32")
band_arrs.append(band_arr)
x_arr = np.stack(band_arrs, axis=-1)
# Apply data augmentations, if provided
if self.transforms:
x_arr = self.transforms(image=x_arr)["image"]
x_arr = np.transpose(x_arr, [2, 0, 1])
# Prepare dictionary for item
item = {"chip_id": img.chip_id, "chip": x_arr}
# Load label if available
if self.label is not None:
label_path = self.label.loc[idx].label_path
with rasterio.open(label_path) as lp:
y_arr = lp.read(1).astype("float32")
# Apply same data augmentations to the label
if self.transforms:
y_arr = self.transforms(image=y_arr)["image"]
item["label"] = y_arr
return item
# + [markdown] tags=[]
# #### Loss class
# +
# %%file {submission_dir}/losses.py
import numpy as np
def intersection_over_union(pred, true):
"""
Calculates intersection and union for a batch of images.
Args:
pred (torch.Tensor): a tensor of predictions
true (torc.Tensor): a tensor of labels
Returns:
intersection (int): total intersection of pixels
union (int): total union of pixels
"""
valid_pixel_mask = true.ne(255) # valid pixel mask
true = true.masked_select(valid_pixel_mask).to("cpu")
pred = pred.masked_select(valid_pixel_mask).to("cpu")
# Intersection and union totals
intersection = np.logical_and(true, pred)
union = np.logical_or(true, pred)
return intersection.sum() / union.sum()
# -
# #### `CloudModel`
#
# Now is the moment we've all been waiting for - coding our actual model!
#
# Again, we'll make our lives simpler by starting with the [`pl.LightningModule`](https://pytorch-lightning.readthedocs.io/en/latest/common/lightning_module.html) class from Pytorch Lightning. This comes with most of the logic we need, so we only have to specify components that are specific to our modeling setup. Our custom `CloudModel` class will define:
#
# - `__init__`: how to instantiate a `CloudModel` class
#
# - `forward`: forward pass for an image in the neural network propogation
#
# - `training_step`: switch the model to train mode, implement the forward pass, and calculate training loss (cross-entropy) for a batch
#
# - `validation_step`: switch the model to eval mode and calculate validation loss (IOU) for the batch
#
# - `train_dataloader`: call an iterable over the training dataset for automatic batching
#
# - `val_dataloader`: call an iterable over the validation dataset for automatic batching
#
# - `configure_optimizers`: configure an [optimizer](https://pytorch.org/docs/stable/optim.html) and a [scheduler](https://pytorch.org/docs/stable/generated/torch.optim.lr_scheduler.ReduceLROnPlateau.html) to dynamically adjust the learning rate based on the number of epochs
#
# - `_prepare_model`: load the U-Net model with a ResNet34 backbone from the `segmentation_models_pytorch` package
# ### MODEL VERSIONING
# | No | Model | Backbone | Weights | Folder | Accuracy |
# | --- | --- | --- | -- | -- | -- |
# | 1 | unet | resnet35 | imagenet | version_3/checkpoints/epoch=25-step=25583.ckpt | .877 |
# | 2 | unet | resnet50 | imagenet | lightning_logs/version_9/checkpoints/epoch=33-step=33455.ckpt| .882 |
# | 3 | unet | resnet101 | imagenet | version_12/checkpoints/epoch=8-step=8855.ckpt | .878 |
# | 4 | unet | resnet50 | imagenet - DiceLoss + 0.5TverskyFocal | version_13/checkpoints/epoch=12-step=12791.ckpt | .880 |
# +
# training_transformations = albumentations.Compose(
# [
# albumentations.RandomCrop(256, 256),
# albumentations.RandomRotate90(),
# albumentations.HorizontalFlip(),
# albumentations.VerticalFlip(),
# ]
# )
# +
# %%file {submission_dir}/cloud_model.py
from typing import Optional, List
import pandas as pd
import pytorch_lightning as pl
import segmentation_models_pytorch as smp
import segmentation_models_pytorch.losses as Loss
import torch
try:
from cloud_dataset import CloudDataset
from losses import intersection_over_union
except ImportError:
from train_src.cloud_dataset import CloudDataset
from train_src.losses import intersection_over_union
class CloudModel(pl.LightningModule):
def __init__(
self,
bands: List[str],
x_train: Optional[pd.DataFrame] = None,
y_train: Optional[pd.DataFrame] = None,
x_val: Optional[pd.DataFrame] = None,
y_val: Optional[pd.DataFrame] = None,
hparams: dict = {},
):
"""
Instantiate the CloudModel class based on the pl.LightningModule
(https://pytorch-lightning.readthedocs.io/en/latest/common/lightning_module.html).
Args:
bands (list[str]): Names of the bands provided for each chip
x_train (pd.DataFrame, optional): a dataframe of the training features with a row for each chip.
There must be a column for chip_id, and a column with the path to the TIF for each of bands.
Required for model training
y_train (pd.DataFrame, optional): a dataframe of the training labels with a for each chip
and columns for chip_id and the path to the label TIF with ground truth cloud cover.
Required for model training
x_val (pd.DataFrame, optional): a dataframe of the validation features with a row for each chip.
There must be a column for chip_id, and a column with the path to the TIF for each of bands.
Required for model training
y_val (pd.DataFrame, optional): a dataframe of the validation labels with a for each chip
and columns for chip_id and the path to the label TIF with ground truth cloud cover.
Required for model training
hparams (dict, optional): Dictionary of additional modeling parameters.
"""
super().__init__()
self.hparams.update(hparams)
self.save_hyperparameters()
# required
self.bands = bands
# optional modeling params
self.backbone = self.hparams.get("backbone", "resnet50")
self.weights = self.hparams.get("weights", "imagenet")
self.learning_rate = self.hparams.get("lr", 1e-3)
self.transforms = self.hparams.get("transforms", None)
self.patience = self.hparams.get("patience", 4)
self.num_workers = self.hparams.get("num_workers", 2)
self.batch_size = self.hparams.get("batch_size", 32)
self.gpu = self.hparams.get("gpu", False)
# Instantiate datasets, model, and trainer params if provided
self.train_dataset = CloudDataset(
x_paths=x_train,
bands=self.bands,
y_paths=y_train,
transforms=self.transforms,
)
self.val_dataset = CloudDataset(
x_paths=x_val,
bands=self.bands,
y_paths=y_val,
transforms=None,
)
self.model = self._prepare_model()
## Required LightningModule methods ##
def forward(self, image: torch.Tensor):
# Forward pass
return self.model(image)
def training_step(self, batch: dict, batch_idx: int):
"""
Training step.
Args:
batch (dict): dictionary of items from CloudDataset of the form
{'chip_id': list[str], 'chip': list[torch.Tensor], 'label': list[torch.Tensor]}
batch_idx (int): batch number
"""
if self.train_dataset.data is None:
raise ValueError(
"x_train and y_train must be specified when CloudModel is instantiated to run training"
)
# Switch on training mode
self.model.train()
torch.set_grad_enabled(True)
# Load images and labels
x = batch["chip"]
# Change to int.
y = batch["label"].long()
if self.gpu:
x, y = x.cuda(non_blocking=True), y.cuda(non_blocking=True)
# Forward pass
preds = self.forward(x)
# Log batch loss
# Combined loss.
loss = Loss.DiceLoss(mode='multiclass', smooth=1.0)(preds, y).mean() \
+ 0.5 * Loss.TverskyLoss(mode='multiclass')(preds, y).mean()
self.log(
"loss",
loss,
on_step=True,
on_epoch=True,
prog_bar=True,
logger=True,
)
return loss
def validation_step(self, batch: dict, batch_idx: int):
"""
Validation step.
Args:
batch (dict): dictionary of items from CloudDataset of the form
{'chip_id': list[str], 'chip': list[torch.Tensor], 'label': list[torch.Tensor]}
batch_idx (int): batch number
"""
if self.val_dataset.data is None:
raise ValueError(
"x_val and y_val must be specified when CloudModel is instantiated to run validation"
)
# Switch on validation mode
self.model.eval()
torch.set_grad_enabled(False)
# Load images and labels
x = batch["chip"]
y = batch["label"].long()
if self.gpu:
x, y = x.cuda(non_blocking=True), y.cuda(non_blocking=True)
# Forward pass & softmax
preds = self.forward(x)
preds = torch.softmax(preds, dim=1)[:, 1]
preds = (preds > 0.5) * 1 # convert to int
# Log batch IOU
batch_iou = intersection_over_union(preds, y)
self.log(
"iou", batch_iou, on_step=True, on_epoch=True, prog_bar=True, logger=True
)
return batch_iou
def train_dataloader(self):
# DataLoader class for training
return torch.utils.data.DataLoader(
self.train_dataset,
batch_size=self.batch_size,
num_workers=self.num_workers,
shuffle=True,
pin_memory=True,
)
def val_dataloader(self):
# DataLoader class for validation
return torch.utils.data.DataLoader(
self.val_dataset,
batch_size=self.batch_size,
num_workers=self.num_workers,
shuffle=False,
pin_memory=True,
)
def configure_optimizers(self):
opt = torch.optim.Adam(self.model.parameters(), lr=self.learning_rate)
sch = torch.optim.lr_scheduler.CosineAnnealingLR(opt, T_max=10)
return [opt], [sch]
## Convenience Methods ##
def _prepare_model(self):
# Instantiate U-Net model
unet_model = smp.Unet(
encoder_name=self.backbone,
encoder_weights=self.weights,
in_channels=4,
classes=2,
)
if self.gpu:
unet_model.cuda()
return unet_model
# + [markdown] tags=[]
# ### Fit the model
# +
import warnings
# warnings.filterwarnings("ignore")
# +
from train_src.cloud_model import CloudModel
# Set up pytorch_lightning.Trainer object
cloud_model = CloudModel(
bands=BANDS,
x_train=train_x,
y_train=train_y,
x_val=val_x,
y_val=val_y,
hparams={
"num_workers": 4,
"batch_size": 8,
},
)
# +
checkpoint_callback = pl.callbacks.ModelCheckpoint(
monitor="iou_epoch", mode="max", verbose=True,
)
early_stopping_callback = pl.callbacks.early_stopping.EarlyStopping(
monitor="iou_epoch",
patience=(cloud_model.patience * 3),
mode="max",
verbose=True,
)
trainer = pl.Trainer(
gpus=1,
fast_dev_run=False,
callbacks=[checkpoint_callback, early_stopping_callback],
)
# -
# + jupyter={"outputs_hidden": false}
# Fit the model
trainer.fit(model=cloud_model, )
# -
raise
# Last checkpoint, noted - 0.87612
# Our best IOU on the validation split is 0.887.
#
# If you'd like to track changes in performance more closely, you could log information about metrics across batches, epochs, and models through the [TensorBoard](https://pytorch.org/tutorials/recipes/recipes/tensorboard_with_pytorch.html) UI.
# + [markdown] tags=[]
# <a id='generate-submission'></a>
#
# ### Generate a submission
#
# Now that we have our trained model, we can generate a full submission. **Remember that this is a [code execution](https://www.drivendata.org/competitions/83/cloud-cover/page/412/) competition,** so you will be submitting our inference code rather than our predictions. We've already written out our key class definition to scripts in the folder `benchmark_src`, which now contains:
#
# ```
# benchmark_src
# ├── cloud_dataset.py
# ├── cloud_model.py
# └── losses.py
# ```
#
# To submit to the competition, we still need to:
#
# 1. Store our trained model weights in `benchmark_src` so that they can be loaded during inference
#
# 2. Write a `main.py` file that loads our model weights, generates predictions for each chip, and saves the predictions to a folder called `predictions` in the same directory as itself
#
# 3. Zip the contents of `benchmark_src/` - not the directory itself - into a file called `submission.zip`.
#
# 4. Upload `submission.zip` to the competition submissions page. The file will be unzipped and `main.py` will be run in a [containerized execution environment](https://github.com/drivendataorg/cloud-cover-runtime) to calculate our model's IOU.
# -
# #### 1. Save our model
#
# First, let's make a folder for our model assets, and save the weights for our trained model using PyTorch's handy `model.save()` method. The below saves the weights to `benchmark_src/assets/cloud_model.pt`.
# +
# save the model
submission_assets_dir = submission_dir / "assets"
submission_assets_dir.mkdir(parents=True, exist_ok=True)
model_weight_path = submission_assets_dir / "cloud_model.pt"
torch.save(cloud_model.state_dict(), model_weight_path)
# -
# #### 2. Write `main.py`
#
# Now we'll write out a script called `main.py` to `benchmark_src`, which runs the whole inference process using the saved model weights.
# +
# %%file benchmark_src/main.py
import os
from pathlib import Path
from typing import List
from loguru import logger
import pandas as pd
from PIL import Image
import torch
import typer
try:
from cloud_dataset import CloudDataset
from cloud_model import CloudModel
except ImportError:
from benchmark_src.cloud_dataset import CloudDataset
from benchmark_src.cloud_model import CloudModel
ROOT_DIRECTORY = Path("/codeexecution")
PREDICTIONS_DIRECTORY = ROOT_DIRECTORY / "predictions"
ASSETS_DIRECTORY = ROOT_DIRECTORY / "assets"
DATA_DIRECTORY = ROOT_DIRECTORY / "data"
INPUT_IMAGES_DIRECTORY = DATA_DIRECTORY / "test_features"
# Set the pytorch cache directory and include cached models in your submission.zip
os.environ["TORCH_HOME"] = str(ASSETS_DIRECTORY / "assets/torch")
def get_metadata(features_dir: os.PathLike, bands: List[str]):
"""
Given a folder of feature data, return a dataframe where the index is the chip id
and there is a column for the path to each band's TIF image.
Args:
features_dir (os.PathLike): path to the directory of feature data, which should have
a folder for each chip
bands (list[str]): list of bands provided for each chip
"""
chip_metadata = pd.DataFrame(index=[f"{band}_path" for band in bands])
chip_ids = (
pth.name for pth in features_dir.iterdir() if not pth.name.startswith(".")
)
for chip_id in chip_ids:
chip_bands = [features_dir / chip_id / f"{band}.tif" for band in bands]
chip_metadata[chip_id] = chip_bands
return chip_metadata.transpose().reset_index().rename(columns={"index": "chip_id"})
def make_predictions(
model: CloudModel,
x_paths: pd.DataFrame,
bands: List[str],
predictions_dir: os.PathLike,
):
"""Predicts cloud cover and saves results to the predictions directory.
Args:
model (CloudModel): an instantiated CloudModel based on pl.LightningModule
x_paths (pd.DataFrame): a dataframe with a row for each chip. There must be a column for chip_id,
and a column with the path to the TIF for each of bands provided
bands (list[str]): list of bands provided for each chip
predictions_dir (os.PathLike): Destination directory to save the predicted TIF masks
"""
test_dataset = CloudDataset(x_paths=x_paths, bands=bands)
test_dataloader = torch.utils.data.DataLoader(
test_dataset,
batch_size=model.batch_size,
num_workers=model.num_workers,
shuffle=False,
pin_memory=True,
)
for batch_index, batch in enumerate(test_dataloader):
logger.debug(f"Predicting batch {batch_index} of {len(test_dataloader)}")
x = batch["chip"]
preds = model.forward(x)
preds = torch.softmax(preds, dim=1)[:, 1]
preds = (preds > 0.5).detach().numpy().astype("uint8")
for chip_id, pred in zip(batch["chip_id"], preds):
chip_pred_path = predictions_dir / f"{chip_id}.tif"
chip_pred_im = Image.fromarray(pred)
chip_pred_im.save(chip_pred_path)
def main(
model_weights_path: Path = ASSETS_DIRECTORY / "cloud_model.pt",
test_features_dir: Path = DATA_DIRECTORY / "test_features",
predictions_dir: Path = PREDICTIONS_DIRECTORY,
bands: List[str] = ["B02", "B03", "B04", "B08"],
fast_dev_run: bool = False,
):
"""
Generate predictions for the chips in test_features_dir using the model saved at
model_weights_path.
Predictions are saved in predictions_dir. The default paths to all three files are based on
the structure of the code execution runtime.
Args:
model_weights_path (os.PathLike): Path to the weights of a trained CloudModel.
test_features_dir (os.PathLike, optional): Path to the features for the test data. Defaults
to 'data/test_features' in the same directory as main.py
predictions_dir (os.PathLike, optional): Destination directory to save the predicted TIF masks
Defaults to 'predictions' in the same directory as main.py
bands (List[str], optional): List of bands provided for each chip
"""
if not test_features_dir.exists():
raise ValueError(
f"The directory for test feature images must exist and {test_features_dir} does not exist"
)
predictions_dir.mkdir(exist_ok=True, parents=True)
logger.info("Loading model")
model = CloudModel(bands=bands, hparams={"weights": None})
model.load_state_dict(torch.load(model_weights_path))
logger.info("Loading test metadata")
test_metadata = get_metadata(test_features_dir, bands=bands)
if fast_dev_run:
test_metadata = test_metadata.head()
logger.info(f"Found {len(test_metadata)} chips")
logger.info("Generating predictions in batches")
make_predictions(model, test_metadata, bands, predictions_dir)
logger.info(f"""Saved {len(list(predictions_dir.glob("*.tif")))} predictions""")
if __name__ == "__main__":
typer.run(main)
# -
# If we wanted to test out running `main` from this notebook, we could execute:
#
# ```python
# from benchmark_src.main import main
#
# main(
# model_weights_path=submission_dir / "assets/cloud_model.pt",
# test_features_dir=TRAIN_FEATURES,
# predictions_dir=submission_dir / "predictions",
# fast_dev_run=True,
# )
# ```
# #### 3. Zip submission contents
#
# Compress all of the submission files in `benchmark_src` into a .zip called `submission.zip`. Our final submission directory has:
# clear out our pycache before zipping up submission
# !rm -rf benchmark_src/__pycache__
# !tree benchmark_src
# Remember to make sure that your submission does *not* include any prediction files.
# Zip submission
# !cd benchmark_src && zip -r ../submission.zip *
# !du -h submission.zip
# ### Upload submission
# We can now head over to the competition [submissions page](https://www.drivendata.org/competitions/83/cloud-cover/submissions/) to upload our code and get our model's IOU!
#
# 
#
# Our submission took about 20 minutes to execute. You can monitor progress during scoring with the Code Execution Status [tab](https://www.drivendata.org/competitions/83/submissions/code/). Finally, we see that we got an IOU of **0.817** - that's pretty good! It means that 81.7% of the area covered by either the ground truth labels or our predictions was shared between the two.
#
# There is still plenty of room for improvement! Head over to the On Cloud N challenge [homepage](https://www.drivendata.org/competitions/83/cloud-cover/page/396/) to get started on your own model. We're excited to see what you create!
#
| misc/training_script.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Diagnostics" data-toc-modified-id="Diagnostics-1"><span class="toc-item-num">1 </span>Diagnostics</a></span><ul class="toc-item"><li><span><a href="#Summary-Statistics-Selection" data-toc-modified-id="Summary-Statistics-Selection-1.1"><span class="toc-item-num">1.1 </span>Summary Statistics Selection</a></span><ul class="toc-item"><li><span><a href="#Example:-Summary-statistics-selection-for-the-MA2-model" data-toc-modified-id="Example:-Summary-statistics-selection-for-the-MA2-model-1.1.1"><span class="toc-item-num">1.1.1 </span>Example: Summary statistics selection for the MA2 model</a></span></li></ul></li></ul></li></ul></div>
# -
# # Diagnostics
# + init_cell=true
from functools import partial
import numpy as np
import scipy.stats as ss
import elfi
import logging
logging.basicConfig(level=logging.INFO)
# + init_cell=true
from elfi.methods import diagnostics
# -
# ## Summary Statistics Selection
# One of the most difficult aspects in likelihood-free inference is finding a good way to compare simulated and observed data. This is typically accomplished via a set of summary statistics, but they tend to be very problem-specific. ELFI includes tools to aid this selection in the form of the Two Stage Procedure proposed by [Nunes & Balding (2010)](https://www.degruyter.com/view/j/sagmb.2010.9.1/sagmb.2010.9.1.1576/sagmb.2010.9.1.1576.xml), which determines a well-performing summary statistics combination. The procedure can be summarised as follows:
#
# - First, all possible combinations of the candidate summary statistics are generated (can also be user-given)
# - Stage 1:
# - Each summary statistics combination is evaluated using the Minimum Entropy algorithm
# - The minimum entropy combination is selected, and the 'closest' datasets are identified
# - Stage 2:
# - For each summary statistics combination, the mean root sum of squared errors (MRSSE) is calculated over all 'closest' datasets
# - The minimum MRSSE combination is chosen as the one with the optimal performance.
# ### Example: Summary statistics selection for the MA2 model
# We will again use the MA2 example introduced in the [tutorial](tutorial.ipynb), where we used the first two autocovariances as the summary statistics: one with `lag=1`, another with `lag=2`.
from elfi.examples import ma2
m = ma2.get_model()
elfi.draw(m)
# Let's see if it would be beneficial try other summary statistics as well, for example the mean and the variance. To use the Two-Stage Selection process, we have to define the ElfiModel up until **the node for which the summary statistics will be applied**, which is typically the simulator node (here, named *MA2*). Because the MA2 example defines a complete ElfiModel, we have to remove the summary statistics (and anything after them, in this case the distance) from it:
m.remove_node('S1')
m.remove_node('S2')
m.remove_node('d')
elfi.draw(m)
# Next we need to define a list of candidate summary statistics:
# +
autocov1 = ma2.autocov
autocov2 = partial(ma2.autocov, lag=2)
autocov2.__name__ = 'autocov2' # Note: the name must be given separately if using partial
def mean(y):
return np.mean(y, axis=1)
def var(y):
return np.var(y, axis=1)
# Initialising the list of assessed summary statistics.
list_ss = [autocov1, autocov2, mean, var]
# -
# ELFI will generate all possible combinations of these candidates, and build an ElfiModel for each combination by generating child nodes to the user-given node (here, the simulator node MA2). A distance node between the summary statistics can be given as a function or string as with `elfi.Distance` or `elfi.Discrepancy` (here, 'euclidean'):
selection = diagnostics.TwoStageSelection(m['MA2'], 'euclidean', list_ss=list_ss)
# Sometimes the generated list of combinations may be very long. If you are able to make an educated guess about which combinations are the most promising, you can save computational time by providing these combinations to ELFI. This can be done by replacing the `list_ss` keyword argument with for example:
#
# `prepared_ss=[[autocov1], [autocov1, autocov2], [mean, var]]`
#
# and then ELFI will only consider these combinations.
# After these preparations, we can execute the selection process as follows:
ss = selection.run(n_sim=100000, batch_size=10000)
ss
# So the Two-Stage Procedure supports our earlier decision to use the autocovariances with lags 1 and 2. :)
#
# The method includes further options for tuning the selection process, please check the [documentation](http://elfi.readthedocs.io) for more details.
| diagnostics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ml2
# language: python
# name: ml2
# ---
# # Fast Explorer
#
# > Work in progress
# A utility to visualize models.
#
# Main idea:
# * Use this library to attach a learner and initialize a proxy server.
# * Connect a javascript client via [fastexplorer-js](https://github.com/renato145/fastexplorer-js).
# * Send back and forth relevant information to be visualized.
#
# This will allow to use tools like d3, react and threejs to visualize NN information.
# ## Install
# `pip install git+https://github.com/renato145/fastexplorer.git`
# ## How to use
# all_slow
# %reload_ext autoreload
# %autoreload 2
# Load you Learner as usual and import fastexplorer:
# +
from fastai.vision.all import *
from fastexplorer.all import *
path = untar_data(URLs.PETS)
files = get_image_files(path/"images")
def label_func(f): return f[0].isupper()
dls = ImageDataLoaders.from_name_func(path, files, label_func, item_tfms=Resize(224))
learn = cnn_learner(dls, resnet34, metrics=accuracy)
# +
#hide
# learn.fine_tune(1)
# -
# When ready, start serving the server:
#srv
learn.fastexplorer(True)
# Finally, go to [https://renato145.github.io/fastexplorer-js/](https://renato145.github.io/fastexplorer-js/) to visualize the model:
# 
#
# Also, you can visualize the [Loss Landscape](https://arxiv.org/abs/1712.09913):
# > Note that the original work calculates the landscape using the whole dataset which can take a lot of time. In this library, I am using just one batch of data to calculate the landscape (this will probably change once I find a faster way to calculate it).
#
# 
| nbs/index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Labelling
#
# ## Objetivos
#
# - Revisar algunos algortimos de Advances in Financial Machine Learning. Capítulos 2, 3 (y algo del 20).
# - Tópicos de Labelling.
#
#
# ## Bibliografia
#
# - <NAME>. Advances in Financial Machine Learning. Wiley
#
# ## Recursos
#
# - Python 3.7+
# - Jupyter notebook.
# - Pandas
# - numpy
# - matplotlib
#
# ## Descripcion
#
# En la siguiente notebook vamos a seguir la linea de trabajo del capítulo 3 del libro.
# A partir de un dataset descargado de Yahoo Finance del ETF [MTUM](https://www.ishares.com/us/products/251614/ishares-msci-usa-momentum-factor-etf) vamos a
# generar labels de entrada y salida con el método de la triple frontera:
#
# - Frontera de profit taking (frontera horizontal superior)
# - Frontera de stop loss (frontera horizontal inferior)
# - Frontera temporal (frontera vertical)
#
# En el libro se discute brevemente que hacer cuando la señal no toca ninguna de las fronteras
# horizontales y alcanza el span temporal. <NAME> recomienda tomar el signo del retorno
# en ese instante temporal, aquí se opta por la innacción, es decir, mantenemos la pocisión.
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# +
# # %load mpfin.py
import multiprocessing as mp
import datetime as dt
import time
import sys
def mpPandasObj(func,pdObj,numThreads=24,mpBatches=1,linMols=True,**kargs):
'''
Parallelize jobs, return a dataframe or series
multiprocessing snippet [20.7]
+ func: function to be parallelized. Returns a DataFrame
+ pdObj[0]: Name of argument used to pass the molecule
+ pdObj[1]: List of atoms that will be grouped into molecules
+ kwds: any other argument needed by func
Example: df1=mpPandasObj(func,('molecule',df0.index),24,**kwds)
'''
import pandas as pd
#if linMols:parts=linParts(len(argList[1]),numThreads*mpBatches)
#else:parts=nestedParts(len(argList[1]),numThreads*mpBatches)
if linMols:parts=linParts(len(pdObj[1]),numThreads*mpBatches)
else:parts=nestedParts(len(pdObj[1]),numThreads*mpBatches)
jobs=[]
for i in range(1,len(parts)):
job={pdObj[0]:pdObj[1][parts[i-1]:parts[i]],'func':func}
job.update(kargs)
jobs.append(job)
if numThreads==1:out=processJobs_(jobs)
else: out=processJobs(jobs,numThreads=numThreads)
if isinstance(out[0],pd.DataFrame):df0=pd.DataFrame()
elif isinstance(out[0],pd.Series):df0=pd.Series()
else:return out
for i in out:df0=df0.append(i)
df0=df0.sort_index()
return df0
def processJobs_(jobs):
# Run jobs sequentially, for debugging
out=[]
for job in jobs:
out_=expandCall(job)
out.append(out_)
return out
def linParts(numAtoms,numThreads):
# partition of atoms with a single loop
parts=np.linspace(0,numAtoms,min(numThreads,numAtoms)+1)
parts=np.ceil(parts).astype(int)
return parts
def nestedParts(numAtoms,numThreads,upperTriang=False):
# partition of atoms with an inner loop
parts,numThreads_=[0],min(numThreads,numAtoms)
for num in range(numThreads_):
part=1+4*(parts[-1]**2+parts[-1]+numAtoms*(numAtoms+1.)/numThreads_)
part=(-1+part**.5)/2.
parts.append(part)
parts=np.round(parts).astype(int)
if upperTriang: # the first rows are heaviest
parts=np.cumsum(np.diff(parts)[::-1])
parts=np.append(np.array([0]),parts)
return parts
import datetime as dt
def reportProgress(jobNum,numJobs,time0,task):
# Report progress as asynch jobs are completed
msg=[float(jobNum)/numJobs, (time.time()-time0)/60.]
msg.append(msg[1]*(1/msg[0]-1))
timeStamp=str(dt.datetime.fromtimestamp(time.time()))
msg=timeStamp+' '+str(round(msg[0]*100,2))+'% '+task+' done after '+ \
str(round(msg[1],2))+' minutes. Remaining '+str(round(msg[2],2))+' minutes.'
if jobNum<numJobs:sys.stderr.write(msg+'\r')
else:sys.stderr.write(msg+'\n')
return
def processJobs(jobs,task=None,numThreads=24):
# Run in parallel.
# jobs must contain a 'func' callback, for expandCall
if task is None:task=jobs[0]['func'].__name__
pool=mp.Pool(processes=numThreads)
outputs,out,time0=pool.imap_unordered(expandCall,jobs),[],time.time()
# Process asyn output, report progress
for i,out_ in enumerate(outputs,1):
out.append(out_)
reportProgress(i,len(jobs),time0,task)
pool.close();pool.join() # this is needed to prevent memory leaks
return out
def expandCall(kargs):
# Expand the arguments of a callback function, kargs['func']
func=kargs['func']
del kargs['func']
out=func(**kargs)
return out
# +
# # %load labelling.py
import numpy as np
import pandas as pd
def getDailyVol(close, span0=100):
'''
Computes the daily volatility of price returns.
It takes a closing price series, applies a diff sample to sample
(assumes each sample is the closing price), computes an EWM with
`span0` samples and then the standard deviation of it.
See Advances in Financial Analytics, snippet 3.1
@param[in] close A series of prices where each value is the closing price of an asset.
The index of the series must be a valid datetime type.
@param[in] span0 The sample size of the EWM.
@return A pandas series of daily return volatility.
'''
df0 = close.index.searchsorted(close.index-pd.Timedelta(days=1))
df0 = df0[df0 > 0]
df0 = pd.Series(close.index[df0-1], index=close.index[close.shape[0]-df0.shape[0]:])
df0 = close.loc[df0.index] / close.loc[df0.values].values-1 # Daily returns
df0 = df0.ewm(span=span0).std()
return df0
def getVerticalBarrier(tEvents, close, numDays=0):
"""
Adding a Vertical Barrier
For each index in t_events, it finds the timestamp of the next price bar at or immediately after
a number of days num_days. This vertical barrier can be passed as an optional argument t1 in get_events.
This function creates a series that has all the timestamps of when the vertical barrier would be reached.
Advances in Financial Machine Learning, Snippet 3.4 page 49.
@param tEvents A pd.DateTimeIndex of events.
@param close A pd.Series of close prices.
@param numDays The number of days to add for vertical barrier.
@return A pd.Series of Timestamps of vertical barriers
"""
verticalBarrier = close.index.searchsorted(tEvents + pd.Timedelta(days=numDays))
verticalBarrier = verticalBarrier[verticalBarrier < close.shape[0]]
return pd.Series(close.index[verticalBarrier], index = tEvents[:verticalBarrier.shape[0]]) # NaNs at the end
def applyPtSlOnT1(close, events, ptSl, molecule):
'''
Apply stop loss/profit taking, if it takes place before t1 (vertical barrier)
(end of event).
Advances in Financial Machine Learning, snippet 3.2 page 45.
@param close
@param events
@param ptSl
@param molecule
@return
'''
events_ = events.loc[molecule]
out = events_[['t1']].copy(deep=True)
if ptSl[0] > 0:
pt = ptSl[0] * events_['trgt']
else:
pt = pd.Series(index=events.index) # NaNs
if ptSl[1] > 0:
sl = -ptSl[1] * events_['trgt']
else:
sl=pd.Series(index=events.index) # NaNs
for loc, t1 in events_['t1'].fillna(close.index[-1]).iteritems():
df0 = close[loc:t1] # path prices
df0 = (df0 / close[loc] - 1) * events_.at[loc,'side'] # path returns
out.loc[loc,'sl'] = df0[df0<sl[loc]].index.min() # earliest stop loss
out.loc[loc,'pt'] = df0[df0>pt[loc]].index.min() # earliest profit taking
return out
def getEvents(close, tEvents, ptSl, trgt, minRet, numThreads, t1=False, side=None):
#1) get target
trgt=trgt.loc[tEvents]
trgt=trgt[trgt>minRet] # minRet
#2) get t1 (max holding period)
if t1 is False:
t1 = pd.Series(pd.NaT, index=tEvents)
#3) form events object, apply stop loss on t1
if side is None:
side_, ptSl_ = pd.Series(1.,index=trgt.index), [ptSl[0],ptSl[0]]
else:
side_, ptSl_= side.loc[trgt.index],ptSl[:2]
events = (pd.concat({'t1':t1,'trgt':trgt,'side':side_}, axis=1) .dropna(subset=['trgt']))
df0=mpPandasObj(func=applyPtSlOnT1,pdObj=('molecule',events.index),
numThreads=numThreads,close=close,events=events,
ptSl=ptSl_)
events['t1'] = df0.dropna(how='all').min(axis=1) # pd.min ignores nan
if side is None:events=events.drop('side',axis=1)
return events
def getBinsOld(events,close):
# Snippet 3.5
#1) prices aligned with events
events_=events.dropna(subset=['t1'])
px=events_.index.union(events_['t1'].values).drop_duplicates()
px=close.reindex(px,method='bfill')
#2) create out object
out=pd.DataFrame(index=events_.index)
out['ret']=px.loc[events_['t1'].values].values/px.loc[events_.index]-1
out['bin']=np.sign(out['ret'])
# Where out index and t1 (vertical barrier) intersect label 0
# See page 49, it is a suggested exercise.
try:
locs = out.query('index in @t1').index
out.loc[locs, 'bin'] = 0
except:
pass
return out
def getBins(events, close):
'''
Compute event's outcome (including side information, if provided).
Snippet 3.7
Case 1: ('side' not in events): bin in (-1,1) <-label by price action
Case 2: ('side' in events): bin in (0,1) <-label by pnl (meta-labeling)
@param events It's a dataframe whose
- index is event's starttime
- Column t1 is event's endtime
- Column trgt is event's target
- Column side (optional) implies the algo's position side.
@param close It's a close price series.
'''
#1) prices aligned with events
events_=events.dropna(subset=['t1'])
px=events_.index.union(events_['t1'].values).drop_duplicates()
px=close.reindex(px,method='bfill')
#2) create out object
out=pd.DataFrame(index=events_.index)
out['ret']=px.loc[events_['t1'].values].values/px.loc[events_.index]-1
if 'side' in events_:
out['ret']*=events_['side'] # meta-labeling
out['bin']=np.sign(out['ret'])
if 'side' in events_:
out.loc[out['ret']<=0,'bin']=0 # meta-labeling
return out
def dropLabels(events, minPct=.05):
'''
Takes a dataframe of events and removes those labels that fall
below minPct (minimum percentil).
Snippet 3.8
@param events An events dataframe, such as the output of getBins()
@param minPct The minimum percentil of rare labels to have.
@return The input @p events dataframe but filtered.
'''
# apply weights, drop labels with insufficient examples
while True:
df0=events['bin'].value_counts(normalize=True)
if df0.min()>minPct or df0.shape[0]<3:break
print('dropped label: ', df0.argmin(),df0.min())
events=events[events['bin']!=df0.argmin()]
return events
# +
# # %load events.py
def cusumFilterEvents(close, threshold):
'''
Symmetric CUSUM Filter.
It is a quality-control method, designed to detect a shift in
the mean value of the measured quantity away from a target value.
The value of each filter is:
S^{+}_t = max{0, S^{+}_{t-1} + y_t - E_{t-1}[y_t]}, S^{+}_0 = 0
S^{-}_t = mix{0, S^{-}_{t-1} + y_t - E_{t-1}[y_t]}, S^{-}_0 = 0
S_t = max{S^{+}_t, -S^{-}_t}
See Advances in Financial Analytics, snippet 2.4, page 39.
@param close A price series.
@param threshold A positive threshold to flag a positive or negative
event when either S^{+}_t or S^{-}_t is greater than it.
@return A subset of the @p close index series where the events of
filter are triggered.
'''
tEvents = []
sPos = 0
sNeg = 0
diff = close.diff()
for i in diff.index[1:]:
sPos, sNeg = max(0, sPos+diff.loc[i]), min(0, sNeg+diff.loc[i])
if sNeg < -threshold:
sNeg = 0
tEvents.append(i)
elif sPos > threshold:
sPos = 0
tEvents.append(i)
return pd.to_datetime(tEvents)
def getEwmDf(close, fast_window_num_days=3, slow_window_num_days=7):
'''
Generates a close prices dataframe with three columns, where:
- Column "close" is the `close` price.
- Column "fast" is the `fast` EWM with @p fast_window_num_days days window size.
- Column "slow" is the `slow` EWM with @p slow_window_num_days days window size.
@param close A pandas series of close daily prices.
@param fast_window_num_days A positive integer indicating the fast window size in days.
@param slow_window_num_days A positive integer indicating the slow window size in days,
which is greater than @p fast_window_num_days.
@return A dataframe as described above.
'''
close_df = (pd.DataFrame()
.assign(close=close)
.assign(fast=close.ewm(fast_window_num_days).mean())
.assign(slow=close.ewm(slow_window_num_days).mean()))
return close_df
def get_up_cross(df):
'''
@return A pandas series of events from @p df (as provided in getEwmDf()) when the
fast signal crosses over the slow signal positively.
'''
crit1 = df.fast.shift(1) < df.slow.shift(1)
crit2 = df.fast > df.slow
return df.fast[(crit1) & (crit2)]
def get_down_cross(df):
'''
@return A pandas series of events from @p df (as provided in getEwmDf()) when the
fast signal crosses over the slow signal positively.
'''
crit1 = df.fast.shift(1) > df.slow.shift(1)
crit2 = df.fast < df.slow
return df.fast[(crit1) & (crit2)]
def getEwmEvents(close, fast_window_num_days=3, slow_window_num_days=7):
'''
Generates a pandas series from @p close price series whose events are generated
from the EWM fast and slow signal crosses. When the fast signal crosses with a
positive slope the slow signal a "1" is flagged and when the opposite happens
a "-1" is flagged.
@param close A pandas series of close daily prices.
@param fast_window_num_days A positive integer indicating the fast window size in days.
@param slow_window_num_days A positive integer indicating the slow window size in days,
which is greater than @p fast_window_num_days.
@return A pandas series of events of buy and sell signals.
'''
close_df = getEwmDf(close, fast_window_num_days, slow_window_num_days)
up_events = get_up_cross(close_df)
down_events = get_down_cross(close_df)
side_up = pd.Series(1, index=up_events.index)
side_down = pd.Series(-1, index=down_events.index)
return pd.concat([side_up,side_down]).sort_index()
# -
# ### 1.- Exploración del dataset
#
# - Levantamos el datset desde un CSV
# - Nota: indexamos el dataset con la columna de fechas para poder aprovechar las funciones del libro
# como estan dadas.
# - Describimos el dataset.
# - Mostramos la evolución de las series.
#
#
# A saber, las columnas son:
#
# - `Date`: es la fecha que aplica una estampa temporal a la fila.
# - `Open`: es el precio en USD al que abre la cotización del fondo.
# - `High`: es el mayor valor en USD que alcanza la cotización del fondo.
# - `Low`: es el menor valor en USD que alcanza la cotización del fondo.
# - `Close`: es el valor de cierre en USD que alcanza la cotización del fondo.
# - `Adj Close`: es el precio ajustado que alcanzan algunos activos producto de multiplicadores por regalías y dividendos. Ver https://help.yahoo.com/kb/SLN28256.html para una descripción más detallada al respecto.
# - `Volume`: cantidad total de USD operados.
MTUM_PATH = '../datasets/mtum.csv'
# Función que permite convertir el formato de las fechas como string
# en un objeto de fecha.
string_to_date = lambda x: pd.datetime.strptime(x, "%Y-%m-%d")
mtum = pd.read_csv(MTUM_PATH, parse_dates=['Date'], date_parser=string_to_date, index_col='Date')
mtum.head() # Filas iniciales del dataset.
mtum.tail() # Filas finales del dataset.
# [#Filas, #Columnas]
mtum.shape[0], mtum.shape[1]
# Tipo de dato de cada columna.
mtum.dtypes
# Ploteamos la series.
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(20,10))
mtum.plot(kind='line',y='Close', color='blue', ax=ax)
mtum.plot(kind='line',y='High', color='green', ax=ax)
mtum.plot(kind='line',y='Low', color='red', ax=ax)
mtum.plot(kind='line',y='Open', color='orange', ax=ax)
plt.title('MTUM prices')
plt.grid()
plt.show()
# En particular los va a interesar el precio de cierre.
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(20,10))
mtum.plot(kind='line',y='Close', color='blue', ax=ax)
plt.title('MTUM Close Price')
plt.grid()
plt.show()
# Solo para comparar, miramos el volumen operado.
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(20,10))
mtum.plot(kind='line',y='Volume', color='blue', ax=ax)
plt.title('MTUM volume')
plt.grid()
plt.show()
# ### 2.- Desarrollo con labels
#
# El procedimiento propuesto en el capítulo 3 del Lopez de Prado sugiere que a
# partir de una estrategia donde conozcamos en qué momento comprar y vender nuestro
# activo, generemos un modelo de ML que nos permita aprender el tamaño de la apuesta.
# En lo que sigue vamos a usar el algoritmo CUSUM simétrico del snippet 2.4 para
# generar las señales de compra y venta.
#
# Cada una de las señales de compra y venta actúa como un disparador que posiciona
# el inicio de la ventana de triple frontera. Debemos entonces generar una marca temporal
# que indique cuando termina dicha ventana.
#
# Una vez que tenemos los inicios de ventana (provistos por nuestra estrategia de trading)
# y las terminaciones como frontera vertical (con un valor temporal a aprender, es un hiperparametro
# de nuestra estrategia como lo son los retornos de entrada y salida!) debemos analizar
# que sucede con la señal de precios para generar el label correcto. Esto implica entender
# si la señal toca la barrera de profit taking, stop loss o la vertical primero.
#
# Finalmente, generamos un data frame que nos indica en qué momento, con qué retorno
# y que señal (entrada o salida). Esto será la información que le daremos a un algoritmo
# de ML para que aprenda el tamaño de la apuesta.
# Computamos la volatilidad diaria, suavizada con una media
# movil pesada de 50 dias (arbitraria, también se podria aprender)
daily_vol = getDailyVol(close=mtum['Close'], span0=50)
# Volatilidad media.
daily_vol.mean()
# Graficamos la volatilidad a lo largo de la historia del ETF.
# La línea roja muestra el valor medio.
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(15,7))
daily_vol.plot(kind='line',y='dailyVol', color='blue', ax=ax)
ax.axhline(daily_vol.mean(),ls='--',color='r')
plt.title('MTUM daily volatility')
plt.grid()
plt.show()
# Usamos CUSUM simetrico como generador de eventos.
# Usamos la media de la volatilidad como umbral que nos indique los eventos
# en que el que operaremos.
tEvents = cusumFilterEvents(mtum['Close'], threshold=daily_vol.mean())
tEvents
# Definimos el tiempo de la barrera vertical. Esto es algo a optimizar
# y debe ajustarse en conjunto con los valores de retorno PT y SL.
num_days = 6
# Computamos las marcas temporales de la ventana. Para cada evento
# en tEvents (inicio de la ventana), obtenemos el final de la ventana.
t1 = getVerticalBarrier(tEvents, mtum['Close'], numDays=num_days)
t1
# Definimos las bandas de PT y SL.
ptsl = [0.015, 0.015]
target = daily_vol
# Seleccionamos el mínimo retorno considerado.
minRet = 0.01
# Numero de CPUs para el procesamiento en paralelo.
cpus = 4
# Generamos los eventos de la triple frontera. En esta funcion obtenemos
# un dataframe cuyo indice es cuando ocurre el evento y tiene 2 columnas
# - t1: momento en el que sucede el evento.
# - trgt: retorno obtenido en ese momento.
triple_barrier_events = getEvents(mtum['Close'],tEvents,ptsl,target,minRet,cpus,t1=t1)
triple_barrier_events
# Obtenemos los labels! Los labels nos dan la siguiente informacion:
# - Indice: momento en el que ocurre el evento segun nuestra estrategia.
# - Columna ret: el retorno que vamos a obtener.
# - Columna bin: lo que sucede con la señal de precio:
# - 1: la señal impacta la barrera de profit taking (horizontal superior).
# - 0: la señal impacta la barrera vertical (no se hace nada).
# - -1: la señal impacta la barrea de stop loss (horizontal inferior).
labels = getBinsOld(triple_barrier_events, mtum['Close'])
labels
# Una pequeña tabla de contención que nos indica como se distribuyen
# los labels.
labels['bin'].value_counts()
| notebooks/labelling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.10 64-bit
# language: python
# name: python3810jvsc74a57bd031f2aee4e71d21fbe5cf8b01ff0e069b9275f58929596ceb00d14d90e3e16cd6
# ---
# # Building a classifier to predict the churn in bank customers
# ### 0.1. Clear the cache
try:
from IPython import get_ipython
get_ipython().magic('clear')
get_ipython().magic('reset -f')
except:
pass
# ### 0.2. Import libraries
# +
import pandas as pd
import numpy as np
import joblib
# standard Data Visualization libraries:
import seaborn as sns
import matplotlib.pyplot as plt
import matplotlib.pylab as pylab
import matplotlib
import matplotlib.ticker as mtick
from IPython.display import display
from pandas.plotting import scatter_matrix
from sklearn.metrics import roc_curve
# sklearn modules for Data Preprocessing:
from sklearn.preprocessing import LabelEncoder, OneHotEncoder
from sklearn.preprocessing import StandardScaler, RobustScaler
# -
# advanced Data Visualization libraries:
import sweetviz as sv
import pandas_profiling as pp
from pandas_profiling import ProfileReport
# modules to use before modeling
from statsmodels.stats.outliers_influence import variance_inflation_factor
from sklearn.model_selection import train_test_split
# +
# sklearn modules for Model Selection:
from sklearn.neighbors import KNeighborsClassifier
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
from sklearn.neighbors import KNeighborsClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
# +
#sklearn modules for Model Evaluation & Improvement:
from sklearn import model_selection
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_val_score
from sklearn.metrics import classification_report, precision_recall_curve
from sklearn.metrics import auc, roc_auc_score, roc_curve, accuracy_score
from sklearn.metrics import confusion_matrix, plot_confusion_matrix
from sklearn.metrics import f1_score, precision_score, recall_score, fbeta_score
# -
# This is to supress the warning messages (if any) generated in our code:
import warnings
warnings.filterwarnings('ignore')
# Setting the seaborn style
sns.set(style="white")
# Setting the matplotlib style
parameters = {'axes.labelsize': 'medium',
'axes.titlesize': 25,
'xtick.labelsize': 15,
'ytick.labelsize' : 15,
'legend.fontsize' : 15,
'legend.title_fontsize' : 20,
'font.family' : 'sans-serif',
'font.style' : "normal"}
plt.rcParams.update(parameters)
# ### 0.3 Functions
def initial_cleaning(df: pd.DataFrame) -> pd.DataFrame:
"""
function for initial cleaning tasks like
removing columns, duplicates etc.
"""
# remove last 2 columns not needed for analysis
df = df.iloc[: , :-2]
return df
def handle_categorical_features(df : pd.DataFrame) -> pd.DataFrame:
"""
function to encode categorical variables
source : https://pbpython.com/categorical-encoding.html
"""
# creating a column list for subset dataframe for categorical variables
col_list = list(df.select_dtypes(include=['object']))
# using Label encoding for columns with 2 or less unique categorical values
le = LabelEncoder()
le_list = []
for col in col_list:
if len(list(df[col].unique())) <= 2:
le.fit(df[col])
df[col] = le.transform(df[col])
le_list.append(col)
col_list = [i for i in col_list if i not in le_list]
# using One Hot encoding for columns with 3 or more unique categorical values
encoded_df = pd.get_dummies(df, columns= col_list, drop_first = True)
return encoded_df
def calc_vif(X : pd.DataFrame) -> pd.DataFrame:
"""
function to calculate VIF which is a measure of multicollinearity
"""
vif = pd.DataFrame()
vif["variables"] = X.columns
vif["VIF"] = [variance_inflation_factor(X.values, i) for i in range(X.shape[1])]
return vif
def feature_scaling(df : pd.DataFrame, method : str ) -> pd.DataFrame:
"""
function to scale the given data as per method input
"""
# choosing the scaler acc to input
which_scaler = {
"Standard": StandardScaler(),
"Robust": RobustScaler()
}
scaler = which_scaler.get(method)
# saving the scaled results in scaled_df
df2 = pd.DataFrame(scaler.fit_transform(df))
df2.columns = df.columns.values
df2.index = df.index.values
scaled_df = df2
return scaled_df
def models_table(df : pd.DataFrame) -> pd.DataFrame:
"""
function to study different classifers using the test dataset.
a results table is returned for comparison.
"""
# Defining different Models to use for classification
models = []
models.append(('Logistic Regression', LogisticRegression(solver='liblinear', random_state = 0,
class_weight='balanced')))
models.append(('SVC', SVC(kernel = 'linear', random_state = 0)))
models.append(('Kernel SVM', SVC(kernel = 'rbf', random_state = 0)))
models.append(('KNN', KNeighborsClassifier(n_neighbors = 5, metric = 'minkowski', p = 2)))
models.append(('Gaussian NB', GaussianNB()))
models.append(('Decision Tree Classifier', DecisionTreeClassifier(criterion = 'entropy', random_state = 0)))
models.append(('Random Forest', RandomForestClassifier(
n_estimators=100, criterion = 'entropy', random_state = 0)))
#Evaluating Model Results:
# set table to populate with performance results
col = ['Algorithm', 'ROC AUC Mean', 'ROC AUC STD', 'Accuracy Mean', 'Accuracy STD']
model_results = pd.DataFrame(columns=col)
i = 0
# Evaluate each model using k-fold cross-validation:
for name, model in models:
kfold = model_selection.KFold( n_splits=10)
# accuracy scoring:
cv_acc_results = model_selection.cross_val_score(model,df, y_train, cv=kfold, scoring='accuracy')
# roc_auc scoring:
cv_auc_results = model_selection.cross_val_score( model, df, y_train, cv=kfold, scoring='roc_auc')
model_results.loc[i] = [name,
round(cv_auc_results.mean()*100, 2),
round(cv_auc_results.std()*100, 2),
round(cv_acc_results.mean()*100, 2),
round(cv_acc_results.std()*100, 2)
]
i += 1
# Showcasing the results table
model_results.set_index('Algorithm',inplace=True )
model_results = model_results.sort_values(by=['ROC AUC Mean'], ascending=False)
return model_results
def parameter_finding(n : int) -> float:
"""
function to create a graph for different number of estimators
returns the maximum score obtained
"""
# finding optimal Random Forest parameter
score_array = []
for each in range(1,n):
cl = RandomForestClassifier(n_estimators = each, random_state = 1)
cl.fit(rs_X_train,y_train)
score_array.append(cl.score(rs_X_test,y_test))
# plotting the graph
fig = plt.figure(figsize=(15, 7))
plt.plot(range(1,n),score_array, color = '#ec838a')
plt.ylabel('Score\n',horizontalalignment="center")
plt.xlabel('Range\n',horizontalalignment="center")
plt.title('Optimal Number of Random Forest Estimators \n')
plt.legend(loc='upper right')
plt.savefig('Estimator finding.png', bbox_inches='tight')
plt.show()
return round(max(score_array),5)
def models_table_tuned(df, df2, m, n):
"""
function to see different impact of different hyperparameters
"""
models = []
models.append(('Logistic Regression', LogisticRegression(solver='liblinear', random_state = 0,
class_weight='balanced')))
models.append(('SVC', SVC(kernel = 'linear', random_state = 0)))
models.append(('Kernel SVM', SVC(kernel = 'rbf', random_state = 0)))
models.append(('KNN', KNeighborsClassifier(n_neighbors = m, metric = 'minkowski', p = 2)))
models.append(('Gaussian NB', GaussianNB()))
models.append(('Decision Tree Classifier',
DecisionTreeClassifier(criterion = 'entropy', random_state = 0)))
models.append(('Random Forest', RandomForestClassifier(
n_estimators=n, criterion = 'entropy', random_state = 0)))
# Evaluating Model Results:
acc_results, prec_results, rec_results, f1_results, f2_results = ([] for i in range(5))
auc_results = []
names = []# set table to table to populate with performance results
col = ['Algorithm', 'ROC AUC','Accuracy', 'Precision', 'Recall', 'F1 Score', 'F2 Score']
model_results = pd.DataFrame(columns=col)
i = 0
for name, model in models:
# execute search
model.fit(df, y_train)
# get the best performing model fit on the whole training set
#best_model = result.best_estimator_
# evaluate model on the hold out dataset
y_pred = model.predict(df2)
# Evaluate results
roc_auc = roc_auc_score(y_test, y_pred )
acc = accuracy_score(y_test, y_pred )
prec = precision_score(y_test, y_pred )
rec = recall_score(y_test, y_pred )
f1 = f1_score(y_test, y_pred )
f2 = fbeta_score(y_test, y_pred, beta=2.0)
model_results.loc[i] = [name,
round(roc_auc.mean()*100, 2),
round(acc.mean()*100, 2),
round(prec.mean()*100, 2),
round(rec.mean()*100, 2),
round(f1.mean()*100, 2),
round(f2.mean()*100, 2)
]
i += 1
model_results.set_index('Algorithm', inplace=True)
model_results = model_results.sort_values(by=["Precision", "Recall", "F2 Score"], ascending=False)
return model_results
def color_high_green(threshold : float):
"""
function to highlight certain values
"""
color = 'lightgreen' if threshold > round(95.50,2) else 'white'
return 'background: %s' % color
def plotting_confusion_matrix(df : pd.DataFrame):
"""
function to plot confusion matrix for given dataframe
"""
plt.figure(figsize = (28,20))
fig, ax = plt.subplots()
sns.set(font_scale=1.0)
sns.heatmap(df_cm, annot=True, fmt='g')
class_names=['Attrited','Existing']
tick_marks = np.arange(len(class_names))
plt.tight_layout();
plt.title('Error matrix\n', y=1.1)
plt.xticks(tick_marks, class_names)
plt.yticks(tick_marks, class_names)
ax.xaxis.set_label_position("top")
plt.ylabel('Actual label\n')
plt.xlabel('Predicted label\n');
plt.savefig('test confusion matrix.png');
return ;
# ## 1. Getting started with bankchurners data
# Data and information about data publicly available [here](https://www.kaggle.com/sakshigoyal7/credit-card-customers?select=BankChurners.csv)
# ### 1.1 Importing the data
# +
# import data as pandas dataframe with CLIENTNUM as the index
df = pd.read_csv('BankChurners.csv',index_col = 0)
# looking at first few rows
df.head(2)
# -
# ### 1.2 Data Exploration
# +
# visual exploration using pandas profiling library
#profile = pp.ProfileReport(df, title = 'Profile Report : Bank Churners - Credit Card Data')
#profile.to_file('CC_pp_Analysis.html')
# +
# visual exploration using sweetviz library
#my_report = sv.analyze(df)
#my_report.show_html('CC_sz_Analysis.html')
# -
# #### From the above two reports we gather that there are **No missing values, No duplicates, No Nan or null values**
# 1. There are 16 categorical and 6 categorical variables.
#
#
# 2. Attrition Rate is the target or response variable and rest being the features.
#
#
# 3. Existing Customer 8500
# Attrited Customer 1627
#
# Female 5358
# Male 4769
#
#
# 4. Most common values
# | Feature Name | Most Common Values |
# | --- | --- |
# | Customer Age | 40-50 |
# | Dependent Count | 2 and 3 |
# | Education Level | Graduate |
# | Marital Status | Marital |
# | Income Category | Less than 40K dollars |
# | Card Category | Blue |
# | Months on Book | 36 |
# | Total_Relationship_Count | 3 |
# | Months_Inactive_12_mon | 3 |
# | Contacts_Count_12_mon | 2 and 3 |
# | Credit_Limit | 1438.3 |
# | Total_Trans_Amt | 3800 - 3900 |
# | Avg_Utilization_Ratio | 2470 zeros |
#
# 5. Correlation exists between certain features that will be studied later in detail
# ### 1.3 Detailed EDA
# +
sns.catplot(x="Marital_Status", hue="Income_Category", col="Gender",
data= df, kind="count",
height=4, aspect=2);
plt.savefig('gender_income_marital', bbox_inches='tight')
# -
not_attrited = df[df.Attrition_Flag != 'Attrited Customer']
not_attrited = not_attrited.drop(columns = 'Attrition_Flag')
not_attrited.head(2)
df_120 = not_attrited[not_attrited.Income_Category == '$120K +' ]
df_120_silver = df_120[df_120.Card_Category == 'Silver' ]
sns.catplot(x="Contacts_Count_12_mon", hue="Months_Inactive_12_mon", col="Card_Category",
data= df_120_silver, kind="count",
height=4, aspect=1.5);
# ### 1.4 Data Preprocessing
# +
# perform basic cleaning on dataset to remove last 2 irrelevant columns
df = initial_cleaning(df)
# looking at first few rows
df.head(2)
# +
# encoding the categorical features
encoded_df = handle_categorical_features(df)
# looking at first few rows
encoded_df.head(2)
# -
# ### 1.5 Preparing data for machine learning Removing multicollinearity
# +
# dataframe containing features
X = encoded_df.drop('Attrition_Flag',axis=1)
# the target
y = encoded_df['Attrition_Flag']
# -
#
# Now we ready to experiment with **Data Modeling**
# ## 2.Data Modeling
# ### 2.1 Splitting the dataset
# 80% of data is used for training
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)
# ### 2.2 Feature scaling
# As all features are scaled differently, we need to scale them.
# This would help in finding the columns with high multicollinearity and then if needed remove those columns.
# We here use the Robust method, as it is more suitable for our data
# +
# first scaling training set
rs_X_train = feature_scaling(X_train, "Robust")
# now calculating vif for to look for features having strong multicollinearity
calc_vif(rs_X_train)
# +
# dropping columns that are highly collinear
ls = ["Total_Revolving_Bal","Avg_Open_To_Buy"]
rs_X_train = rs_X_train.drop(columns = ls)
# revalidate Colinearity
calc_vif(rs_X_train)
# +
# replicating the exact process on test set
rs_X_test = feature_scaling(X_test, "Robust")
rs_X_test = rs_X_test.drop(columns = ls)
# revalidated colinearity and all is aligned.
# -
# ### 2.3 Model Comparison
# We will see below the comparison of results from different classifiers
# calling function to build models
models_table(rs_X_train)
# ### 2.4 Optimizing parameters for baseline models
# We see that Random Forest Classifier provides the best result and we will try to finetune the hyperparameters.
# graphically checking the impact of varying number of estimators(trees) in the random forest
parameter_finding(40)
# ### 2.5 More evaluation of different models
model_results = models_table_tuned(rs_X_train,rs_X_test, 5, 15)
model_results.style.applymap(color_high_green)
# ### By observations above, Random Forest is the best performing overall
# ## 3.Model Evaluation of Random Forest
# ### 3.1 Kfold cross validation for this chosen model
# +
# Fitting Random Forest to the Training set:
classifier = RandomForestClassifier(n_estimators = 15,bootstrap = True,
max_features = 'sqrt', criterion = 'entropy', random_state = 0)
classifier.fit(rs_X_train, y_train)
# Predicting the Test set results
y_pred = classifier.predict(rs_X_test)
#Evaluate Model Results on Test Set:
acc = accuracy_score(y_test, y_pred )
prec = precision_score(y_test, y_pred )
rec = recall_score(y_test, y_pred )
f1 = f1_score(y_test, y_pred )
f2 = fbeta_score(y_test, y_pred, beta=2.0)
results = pd.DataFrame([['Random Forest Classifier',acc, prec, rec, f1, f2]],
columns = ['Model', 'Accuracy', 'Precision', 'Recall', 'F1 Score', 'F2 Score'])
results.set_index('Model', inplace = True)
results
# -
accuracies = cross_val_score(estimator = classifier, X = rs_X_train, y = y_train, cv = 10)
print("Random Forest Classifier Accuracy: %0.2f (+/- %0.2f)" % (accuracies.mean(), accuracies.std() * 2))
# ### 3.2 Confusion matrix
# +
#create a confusion matrix for test data
cm = confusion_matrix(y_test, y_pred)
df_cm = pd.DataFrame(cm, index = ('Attrited','Existing'), columns = ('Attrited','Existing'))
# plotting
plotting_confusion_matrix(df_cm)
# +
# plotting the confusion matrix for train and test data
disp = plot_confusion_matrix(classifier, rs_X_train, y_train, cmap = "GnBu" )
disp.figure_.suptitle("Training Data Confusion Matrix")
disp = plot_confusion_matrix(classifier, rs_X_test, y_test, cmap = "GnBu" )
disp.figure_.suptitle("Testing Data Confusion Matrix")
# -
# detailed clasification report
print(classification_report(y_test,y_pred))
# ### 3.3 ROC curve
# +
classifier.fit(rs_X_train, y_train)
probs = classifier.predict_proba(rs_X_test)
probs = probs[:, 1]
classifier_roc_auc = accuracy_score(y_test, y_pred )
rf_fpr, rf_tpr, rf_thresholds = roc_curve(y_test, classifier.predict_proba(rs_X_test)[:,1])
plt.figure(figsize=(8,4))
# Plot Logistic Regression ROC
plt.plot(rf_fpr, rf_tpr, label='RF Classifier (area = %0.2f)' % classifier_roc_auc)
# Plot Base Rate ROC
plt.plot([0,1], [0,1],label='Base Rate' 'k--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.ylabel('True Positive Rate \n')
plt.xlabel('\nFalse Positive Rate \n')
plt.title('ROC Graph \n')
plt.legend(loc="lower right")
plt.savefig('roc graph.png', dpi=100)
plt.show()
# -
# ### 3.4 Feature Importance
# +
# view the feature scores
feature_scores = pd.Series(classifier.feature_importances_, index=rs_X_train.columns).sort_values(ascending=False)
# most important features
feature_scores.nlargest()
# -
# least important features
feature_scores.nsmallest()
# +
plt.figure(figsize=(48,30))
g = sns.barplot(x=feature_scores.nlargest(15), y=feature_scores.nlargest(15).index)
g.set_yticklabels(['Total Transaction Count','Total Transaction Amount','Avg Utilization Ratio',
'Change in Tr. Count','Change in Tr. Amount','Total Relationship count',
'Credit limit', 'Customer Age','Months of Inacticity', 'Months on Book',
'Contacts made', 'No. of Dependents','Gender', 'Married', 'Single'])
# Add labels to the graph
plt.xlabel('Feature Importance Score')
plt.ylabel('Features')
# Add title to the graph
plt.title("Most Important Features")
# Visualize the graph
plt.savefig('imp features.png');
plt.show()
# -
# ## 4.Final Results
# ### 4.1 Here we will do the **Risk Profiling** of the client
#getting the CLIENTNUM
train_identity = rs_X_train.index.astype(int)
test_identity = pd.Series(rs_X_test.index.values)
# +
# geenrating final results data frame for the test data
final_results = y_test.to_frame()
final_results['predictions'] = y_pred
final_results["propensity_to_retain(%)"] = probs*100
final_results["propensity_to_retain(%)"] = final_results["propensity_to_retain(%)"].round(2)
#final_results['Ranking'] = pd.qcut(final_results['propensity_to_churn(%)'].rank(method = 'first'),10,labels=range(10,0,-1))
# create a list of our conditions
conditions = [
(final_results["propensity_to_retain(%)"] <= 25),
(final_results["propensity_to_retain(%)"] > 25) & (final_results["propensity_to_retain(%)"] <= 50),
(final_results["propensity_to_retain(%)"] > 50) & (final_results["propensity_to_retain(%)"] <= 75),
(final_results["propensity_to_retain(%)"] > 75)
]
# create a list of the values we want to assign for each condition
values = ['Very High', 'High', 'Medium', 'Low']
# create a new column and use np.select to assign values to it using our lists as arguments
final_results['Risk of Attrition'] = np.select(conditions, values)
final_results.head()
# -
# exporting final classification result for test set to csv file
final_results.to_csv(r'final_classification_result.csv', header = True)
# ### 4.2 Digging deeper based on Propensity to retain
# looking at number of clients in each % group
final_results_retain = final_results.groupby(by=["propensity_to_retain(%)"]).count()
final_results_retain['Number of Customers'] = final_results_retain.predictions
final_results_retain.drop(columns = ['Attrition_Flag','predictions','Risk of Attrition'], inplace = True)
final_results_retain.to_csv(r'final_classification_clients_that_retain.csv', header = True)
final_results_retain
# +
# looking at number of clients in each risk group
final_results_riskwise = final_results.groupby(by=["Risk of Attrition"]).count()
reorderlist = ['Very High', 'High', 'Medium', 'Low']
final_results_riskwise = final_results_riskwise.reindex(reorderlist)
final_results_riskwise
# +
sns.barplot(x=final_results_riskwise.predictions, y=final_results_riskwise.index, data=final_results_riskwise)
# Add labels to the graph
plt.xlabel('Number of Clients')
plt.ylabel('Risk of Attrition')
# Add title to the graph
plt.title("Test Data Risk Profile")
# Visualize the graph
plt.savefig('Test_Data_Risk_Profile.png', bbox_inches='tight')
plt.show()
# -
# ### 4.3 Looking at predictions that were not right
#
# +
# mismatch is data frame for type I and type II errors
mismatch = final_results.loc[final_results.Attrition_Flag != final_results.predictions ]
mismatch = mismatch.sort_values( by = ['propensity_to_retain(%)'])
mismatch.head(47)
# -
mismatch = mismatch[mismatch.predictions == 1]
mismatch
# +
# detailed profiling
mismatch = mismatch.groupby(by=[ "Risk of Attrition"]).count()
mismatch['Number of Customers']=mismatch.predictions
mismatch.drop(columns = ['Attrition_Flag','predictions', 'propensity_to_retain(%)'], inplace = True)
# +
mismatch.to_csv(r'mismatch.csv', header = True)
mismatch
# -
# ## 5.Preparing for Deployment
# saving the classifier for use in deployment
filename = 'final_model.model'
i = [classifier]
joblib.dump(i,filename)
# ## Future Analysis
#
# More Specific Analyis based on Risk groups, Age etc needs to be done.
# Profile of Clients who were most likely to be mismatched need to be studied and special attention to be paid to them.
| CleanClassication.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3-azureml
# kernelspec:
# display_name: Python 3.6 - AzureML
# language: python
# name: python3-azureml
# ---
# + gather={"logged": 1618668640956}
import numpy as np
from keras.models import load_model
from glob import glob
from keras.preprocessing.image import load_img, img_to_array, ImageDataGenerator
#from keras.applications.vgg16 import preprocess_input
from keras.applications.resnet50 import preprocess_input, decode_predictions
from PIL import Image
from keras.preprocessing import image
import matplotlib.image as npimg
import matplotlib.pyplot as plt
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1618667294394}
model = load_model('cov_v_nor.h5py')
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1618669541151}
my_image = load_img('api_call_image/x-ray-1261.png', target_size=(224, 224))
my_image = my_image.convert('L')
my_image = image.img_to_array(my_image)
my_image/=255
my_image = np.expand_dims(my_image, axis=0) # image shape is (1, 12, 12, 3)
print(my_image.shape)
prediction = model.predict(my_image)[0][0]#
confidence = abs(.5-prediction)*2
if(prediction>.5):
answer = "positive"
else:
answer = "negative"
print("prediction is ", prediction)
print("confidence is ", confidence)
print("diagnosis is ", answer)
| runTest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # pypdb demos
#
# This is a set of basic examples of the usage and outputs of the various individual functions included in. There are generally three types of functions.
# ### Preamble
# +
# %pylab inline
from IPython.display import HTML
# Import from local directory
# import sys
# sys.path.insert(0, '../pypdb')
# from pypdb import *
# Import from installed package
from pypdb import *
# %load_ext autoreload
# %autoreload 2
# -
# # Search functions that return lists of PDB IDs
# #### Get a list of PDBs for a specific search term
found_pdbs = Query("actin network").search()
print(found_pdbs[:10])
# #### Search by PubMed ID Number
found_pdbs = Query(27499440, "PubmedIdQuery").search()
print(found_pdbs[:10])
# #### Search by source organism using NCBI TaxId
found_pdbs = Query('6239', 'TreeEntityQuery').search() #TaxID for C elegans
print(found_pdbs[:5])
# #### Search by a specific experimental method
found_pdbs = Query('SOLID-STATE NMR', query_type='ExpTypeQuery').search()
print(found_pdbs[:10])
# #### Search by protein structure similarity
found_pdbs = Query('2E8D', query_type="structure").search()
print(found_pdbs[:10])
# #### Search by Author
found_pdbs = Query('<NAME>.', query_type='AdvancedAuthorQuery').search()
print(found_pdbs)
# #### Search by organism
q = Query("Dictyostelium", query_type="OrganismQuery")
print(q.search()[:10])
# # Information Search functions
# While the basic functions described in the previous section are useful for looking up and manipulating individual unique entries, these functions are intended to be more user-facing: they take search keywords and return lists of authors or dates
# #### Find papers for a given keyword
matching_papers = find_papers('crispr', max_results=10)
print(list(matching_papers)[:10])
# # Functions that return information about single PDB IDs
# #### Get the full PDB file
pdb_file = get_pdb_file('4lza', filetype='cif', compression=False)
print(pdb_file[:400])
# #### Get a general description of the entry's metadata
all_info = get_info('4LZA')
print(list(all_info.keys()))
# #### Run a Sequence search
#
# Formerly using BLAST, this method now uses MMseqs2
# +
q = Query("VLSPADKTNVKAAWGKVGAHAGEYGAEALERMFLSFPTTKTYFPHFDLSHGSAQVKGHGKKVADALTAVAHVDDMPNAL",
query_type="sequence",
return_type="polymer_entity")
print(q.search())
# -
# #### Search by PFAM number
pfam_info = Query("PF00008", query_type="pfam").search()
print(pfam_info[:5])
# ## Search for all entries that mention the word 'ribosome'
# +
from pypdb.clients.search.search_client import perform_search
from pypdb.clients.search.search_client import ReturnType
from pypdb.clients.search.operators import text_operators
search_operator = text_operators.DefaultOperator(value="ribosome")
return_type = ReturnType.ENTRY
results = perform_search(search_operator, return_type)
print(results[:10])
# -
# ## Search for polymers from 'Mus musculus'
# +
from pypdb.clients.search.search_client import perform_search
from pypdb.clients.search.search_client import ReturnType
from pypdb.clients.search.operators import text_operators
search_operator = text_operators.ExactMatchOperator(value="Mus musculus",
attribute="rcsb_entity_source_organism.taxonomy_lineage.name")
return_type = ReturnType.POLYMER_ENTITY
results = perform_search(search_operator, return_type)
print(results[:5])
# -
# ## Search for non-polymers from 'Mus musculus' or 'Homo sapiens'
# +
from pypdb.clients.search.search_client import perform_search
from pypdb.clients.search.search_client import ReturnType
from pypdb.clients.search.operators import text_operators
search_operator = text_operators.InOperator(values=["Mus musculus", "Homo sapiens"],
attribute="rcsb_entity_source_organism.taxonomy_lineage.name")
return_type = ReturnType.NON_POLYMER_ENTITY
results = perform_search(search_operator, return_type)
print(results[:5])
# -
# ## Search for polymer instances whose titles contain "actin" or "binding" or "protein"
# +
from pypdb.clients.search.search_client import perform_search
from pypdb.clients.search.search_client import ReturnType
from pypdb.clients.search.operators import text_operators
search_operator = text_operators.ContainsWordsOperator(value="actin-binding protein",
attribute="struct.title")
return_type = ReturnType.POLYMER_INSTANCE
results = perform_search(search_operator, return_type)
print(results[:5])
# -
# ## Search for assemblies that contain the words "actin binding protein"
# (must be in that order).
#
# For example, "actin-binding protein" and "actin binding protein" will match,
# but "protein binding actin" will not.
# +
from pypdb.clients.search.search_client import perform_search
from pypdb.clients.search.search_client import ReturnType
from pypdb.clients.search.operators import text_operators
search_operator = text_operators.ContainsPhraseOperator(value="actin-binding protein",
attribute="struct.title")
return_type = ReturnType.ASSEMBLY
results = perform_search(search_operator, return_type)
print(results[:5])
# -
# ## Search for entries released in 2019 or later
# +
from pypdb.clients.search.search_client import perform_search
from pypdb.clients.search.search_client import ReturnType
from pypdb.clients.search.operators import text_operators
search_operator = text_operators.ComparisonOperator(
value="2019-01-01T00:00:00Z",
attribute="rcsb_accession_info.initial_release_date",
comparison_type=text_operators.ComparisonType.GREATER)
return_type = ReturnType.ENTRY
results = perform_search(search_operator, return_type)
print(results[:5])
# -
# ## Search for entries released only in 2019
# +
from pypdb.clients.search.search_client import perform_search
from pypdb.clients.search.search_client import ReturnType
from pypdb.clients.search.operators import text_operators
search_operator = text_operators.RangeOperator(
from_value="2019-01-01T00:00:00Z",
to_value="2020-01-01T00:00:00Z",
include_lower=True,
include_upper=False,
attribute="rcsb_accession_info.initial_release_date")
return_type = ReturnType.ENTRY
results = perform_search(search_operator, return_type)
print(results[:5])
# -
# ## Search for structures under 4 angstroms of resolution
# +
from pypdb.clients.search.search_client import perform_search
from pypdb.clients.search.search_client import ReturnType
from pypdb.clients.search.operators import text_operators
search_operator = text_operators.ComparisonOperator(
value=4,
attribute="rcsb_entry_info.resolution_combined",
comparison_type=text_operators.ComparisonType.LESS)
return_type = ReturnType.ENTRY
results = perform_search(search_operator, return_type)
print(results[:5])
# -
# ## Search for structures with a given attribute.
#
# (Admittedly every structure has a release date, but the same logic would
# apply for a more sparse RCSB attribute).
#
# +
from pypdb.clients.search.search_client import perform_search
from pypdb.clients.search.search_client import ReturnType
from pypdb.clients.search.operators import text_operators
search_operator = text_operators.ExistsOperator(
attribute="rcsb_accession_info.initial_release_date")
return_type = ReturnType.ENTRY
results = perform_search(search_operator, return_type)
print(results[:5])
# -
# ## Search for 'Mus musculus' or 'Homo sapiens' structures after 2019 using graph search
#
# +
from pypdb.clients.search.search_client import perform_search_with_graph
from pypdb.clients.search.search_client import ReturnType
from pypdb.clients.search.search_client import QueryGroup, LogicalOperator
from pypdb.clients.search.operators import text_operators
# SearchOperator associated with structures with under 4 Angstroms of resolution
under_4A_resolution_operator = text_operators.ComparisonOperator(
value=4,
attribute="rcsb_entry_info.resolution_combined",
comparison_type=text_operators.ComparisonType.GREATER)
# SearchOperator associated with entities containing 'Mus musculus' lineage
is_mus_operator = text_operators.ExactMatchOperator(
value="Mus musculus",
attribute="rcsb_entity_source_organism.taxonomy_lineage.name")
# SearchOperator associated with entities containing 'Homo sapiens' lineage
is_human_operator = text_operators.ExactMatchOperator(
value="Homo sapiens",
attribute="rcsb_entity_source_organism.taxonomy_lineage.name")
# QueryGroup associated with being either human or `Mus musculus`
is_human_or_mus_group = QueryGroup(
queries = [is_mus_operator, is_human_operator],
logical_operator = LogicalOperator.OR
)
# QueryGroup associated with being ((Human OR Mus) AND (Under 4 Angstroms))
is_under_4A_and_human_or_mus_group = QueryGroup(
queries = [is_human_or_mus_group, under_4A_resolution_operator],
logical_operator = LogicalOperator.AND
)
return_type = ReturnType.ENTRY
results = perform_search_with_graph(
query_object=is_under_4A_and_human_or_mus_group,
return_type=return_type)
print("\n", results[:10]) # Huzzah
# -
| demos/demos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="http://cocl.us/pytorch_link_top">
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0110EN/notebook_images%20/Pytochtop.png" width="750" alt="IBM Product " />
# </a>
#
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0110EN/notebook_images%20/cc-logo-square.png" width="200" alt="cognitiveclass.ai logo" />
#
# <h1>Simple One Hidden Layer Neural Network</h1>
#
# <h2>Table of Contents</h2>
# <p>In this lab, you will use a single-layer neural network to classify non linearly seprable data in 1-Ddatabase.</p>
#
# <ul>
# <li><a href="#Model">Neural Network Module and Training Function</a></li>
# <li><a href="#Makeup_Data">Make Some Data</a></li>
# <li><a href="#Train">Define the Neural Network, Criterion Function, Optimizer, and Train the Model</a></li>
# </ul>
# <p>Estimated Time Needed: <strong>25 min</strong></p>
#
# <hr>
#
# <h2>Preparation</h2>
#
# We'll need the following libraries
#
# +
# Import the libraries we need for this lab
import torch
import torch.nn as nn
from torch import sigmoid
import matplotlib.pylab as plt
import numpy as np
torch.manual_seed(0)
# -
# Used for plotting the model
#
# +
# The function for plotting the model
def PlotStuff(X, Y, model, epoch, leg=True):
plt.plot(X.numpy(), model(X).detach().numpy(), label=('epoch ' + str(epoch)))
plt.plot(X.numpy(), Y.numpy(), 'r')
plt.xlabel('x')
if leg == True:
plt.legend()
else:
pass
# -
# <!--Empty Space for separating topics-->
#
# <h2 id="Model">Neural Network Module and Training Function</h2>
#
# Define the activations and the output of the first linear layer as an attribute. Note that this is not good practice.
#
# +
# Define the class Net
class Net(nn.Module):
# Constructor
def __init__(self, D_in, H, D_out):
super(Net, self).__init__()
# hidden layer
self.linear1 = nn.Linear(D_in, H)
self.linear2 = nn.Linear(H, D_out)
# Define the first linear layer as an attribute, this is not good practice
self.a1 = None
self.l1 = None
self.l2=None
# Prediction
def forward(self, x):
self.l1 = self.linear1(x)
self.a1 = sigmoid(self.l1)
self.l2=self.linear2(self.a1)
yhat = sigmoid(self.linear2(self.a1))
return yhat
# -
# Define the training function:
#
# +
# Define the training function
def train(Y, X, model, optimizer, criterion, epochs=1000):
cost = []
total=0
for epoch in range(epochs):
total=0
for y, x in zip(Y, X):
yhat = model(x)
loss = criterion(yhat, y)
loss.backward()
optimizer.step()
optimizer.zero_grad()
#cumulative loss
total+=loss.item()
cost.append(total)
if epoch % 300 == 0:
PlotStuff(X, Y, model, epoch, leg=True)
plt.show()
model(X)
plt.scatter(model.a1.detach().numpy()[:, 0], model.a1.detach().numpy()[:, 1], c=Y.numpy().reshape(-1))
plt.title('activations')
plt.show()
return cost
# -
# <!--Empty Space for separating topics-->
#
# <h2 id="Makeup_Data">Make Some Data</h2>
#
# +
# Make some data
X = torch.arange(-20, 20, 1).view(-1, 1).type(torch.FloatTensor)
Y = torch.zeros(X.shape[0])
Y[(X[:, 0] > -4) & (X[:, 0] < 4)] = 1.0
# -
# <!--Empty Space for separating topics-->
#
# <h2 id="Train">Define the Neural Network, Criterion Function, Optimizer and Train the Model</h2>
#
# Create the Cross-Entropy loss function:
#
# +
# The loss function
def criterion_cross(outputs, labels):
out = -1 * torch.mean(labels * torch.log(outputs) + (1 - labels) * torch.log(1 - outputs))
return out
# -
# Define the Neural Network, Optimizer, and Train the Model:
#
# Train the model
# size of input
D_in = 1
# size of hidden layer
H = 2
# number of outputs
D_out = 1
# learning rate
learning_rate = 0.1
# create the model
model = Net(D_in, H, D_out)
#optimizer
optimizer = torch.optim.SGD(model.parameters(), lr=learning_rate)
#train the model usein
cost_cross = train(Y, X, model, optimizer, criterion_cross, epochs=1000)
#plot the loss
plt.plot(cost_cross)
plt.xlabel('epoch')
plt.title('cross entropy loss')
# By examining the output of the activation, you see by the 600th epoch that the data has been mapped to a linearly separable space.
#
# we can make a prediction for a arbitrary one tensors
#
x=torch.tensor([0.0])
yhat=model(x)
yhat
# we can make a prediction for some arbitrary one tensors
#
X_=torch.tensor([[0.0],[2.0],[3.0]])
Yhat=model(X_)
Yhat
# we can threshold the predication
#
Yhat=Yhat>0.5
Yhat
# <h3>Practice</h3>
#
# Repeat the previous steps above by using the MSE cost or total loss:
#
# +
# Practice: Train the model with MSE Loss Function
learning_rate = 0.1
MSE_criterion = nn.MSELoss()
model=Net(D_in,H,D_out)
optimizer=torch.optim.SGD(model.parameters(), lr=learning_rate)
MSE_cost = train(Y, X, model, optimizer, MSE_criterion, epochs=1000)
#plot the loss
plt.plot(MSE_cost)
plt.xlabel('epoch')
plt.title('MSE loss')
# Type your code here
# -
x=torch.tensor([0.0])
yhat=model(x)
print(yhat)
X_=torch.tensor([[0.0],[2.0],[3.0]])
Yhat=model(X_)
print(Yhat)
# Double-click <b>here</b> for the solution.
#
# <!--
# learning_rate = 0.1
# criterion_mse=nn.MSELoss()
# model=Net(D_in,H,D_out)
# optimizer=torch.optim.SGD(model.parameters(), lr=learning_rate)
# cost_mse=train(Y,X,model,optimizer,criterion_mse,epochs=1000)
# plt.plot(cost_mse)
# plt.xlabel('epoch')
# plt.title('MSE loss ')
# -->
#
# <!--Empty Space for separating topics-->
#
# <a href="http://cocl.us/pytorch_link_bottom">
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0110EN/notebook_images%20/notebook_bottom%20.png" width="750" alt="PyTorch Bottom" />
# </a>
#
# <h2>About the Authors:</h2>
#
# <a href="https://www.linkedin.com/in/joseph-s-50398b136/"><NAME></a> has a PhD in Electrical Engineering, his research focused on using machine learning, signal processing, and computer vision to determine how videos impact human cognition. Joseph has been working for IBM since he completed his PhD.
#
# Other contributors: <a href="https://www.linkedin.com/in/michelleccarey/"><NAME></a>, <a href="www.linkedin.com/in/jiahui-mavis-zhou-a4537814a"><NAME></a>
#
# <hr>
#
# Copyright © 2018 <a href="cognitiveclass.ai?utm_source=bducopyrightlink&utm_medium=dswb&utm_campaign=bdu">cognitiveclass.ai</a>. This notebook and its source code are released under the terms of the <a href="https://bigdatauniversity.com/mit-license/">MIT License</a>.
#
| 7.1_simple1hiddenlayer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [py36]
# language: python
# name: Python [py36]
# ---
# + [markdown] deletable=true editable=true
# # Data Dive 3: Cleaning Data
# ### Making Sense of NYC Restaurant Inspection Data
#
# [This dataset](https://data.cityofnewyork.us/Health/DOHMH-New-York-City-Restaurant-Inspection-Results/43nn-pn8j) provides restaurant inspections, violations, grades and adjudication information. It is freely available through the NYC Open Data portal, though for the purposes of this exercise we'll be using a static version I pulled in early September 2018 that is available at the url below. A Data Dictionary is available [here](https://data.cityofnewyork.us/api/views/43nn-pn8j/files/e3160d37-1b15-41ef-af6b-a830ed39cbc0?download=true&filename=Restaurant_Inspection_Open_Data_Dictionary_082214.xlsx).
#
# Throughout this exercise, we'll be drawing on skills highlighted in the [DataCamp course](https://www.datacamp.com/courses/cleaning-data-in-python), as well as miscellaneous other commands I've introduced in the first two classes. [Ten Minutes to Pandas](https://pandas.pydata.org/pandas-docs/stable/10min.html) is a great reference for these.
#
#
# `https://grantmlong.com/data/DOHMH_New_York_City_Restaurant_Inspection_Results_20180911.csv`.
#
#
# + deletable=true editable=true
import pandas as pd
import requests
# %matplotlib inline
# + [markdown] deletable=true editable=true
# ## Part 1: Load and Inspect
# 1. Load data from DataFrame
# 2. Find the number of rows and columns in the data
# 3. List the columns in the DataFrame
# 4. Display the first twenty rows
#
# + [markdown] deletable=true editable=true
# Load data from DataFrame
# + deletable=true editable=true
df = pd.read_csv('https://grantmlong.com/data/DOHMH_New_York_City_Restaurant_Inspection_Results_20180911.csv')
# + [markdown] deletable=true editable=true
# Find the number of rows and columns in the data
# + deletable=true editable=true
# + [markdown] deletable=true editable=true
# List the columns in the DataFrame
# + deletable=true editable=true
# + [markdown] deletable=true editable=true
# Display the first twenty rows
# + deletable=true editable=true
# + deletable=true editable=true
# + deletable=true editable=true
# + [markdown] deletable=true editable=true
# ## Part 2: Explore and Summarize
# 1. Count the number of unique restaurants in the DataFrame.
# 2. Calculate the share of critical inpections.
# 3. Show a histogram of `SCORE`.
# 4. Create a boxplot of `GRADE` against `SCORE`.
# 5. Describe the `INSPECTION DATE` field.
# 6. Count the number of null values for `VIOLATION DESCRIPTION`.
# 7. Print twenty unique non-null values for `VIOLATION DESCRIPTION`.
#
# + [markdown] deletable=true editable=true
# Count the number of unique restaurants in the DataFrame.
# + deletable=true editable=true
# + [markdown] deletable=true editable=true
# Calculate the share of critical inpections.
# + deletable=true editable=true
# + [markdown] deletable=true editable=true
# Show a histogram of `SCORE`.
# + deletable=true editable=true
# + [markdown] deletable=true editable=true
# Create a boxplot of `GRADE` against `SCORE`.
# + deletable=true editable=true
# + [markdown] deletable=true editable=true
# Describe the `INSPECTION DATE` field.
# + deletable=true editable=true
# + [markdown] deletable=true editable=true
# Count the number of null values for VIOLATION DESCRIPTION.
# + deletable=true editable=true
# + [markdown] deletable=true editable=true
# Print twenty unique violation descriptions.
# + deletable=true editable=true
# + deletable=true editable=true
# + [markdown] deletable=true editable=true
# ## Part 3: Create Clean Variables
# 1. Transform `INSPECTION DATE` to datetime in new variable `inspection_datetime`.
# 2. Create a `inspection_year` variable with the year of the `INSPECTION DATE`.
# 4. Drop observations with `inspection_year` before 2014.
# 5. Drop observations with null values for `VIOLATION DESCRIPTION`.
# 6. Create a `found_vermin` variable for any `VIOLATION DESCRIPTION` containing *vermin*, *mouse*, *mice*, or *rat*.
# 7. Create a `found_bugs` variable for any `VIOLATION DESCRIPTION` containing *insect*, *roach*, or *flies*.
# 8. Create a `bad_temp` variable for any `VIOLATION DESCRIPTION` containing *temperature* or *º F*.
#
#
# Transform `INSPECTION DATE` to datetime in new variable `inspection_datetime`.
# + deletable=true editable=true
# + [markdown] deletable=true editable=true
# Create an `inspection_year` variable with the year of the `INSPECTION DATE`.
#
# + deletable=true editable=true
# + [markdown] deletable=true editable=true
# Drop observations with `inspection_year` before 2014.
# + deletable=true editable=true
# + [markdown] deletable=true editable=true
# Drop observations with null values for `VIOLATION DESCRIPTION`.
# + deletable=true editable=true
# + [markdown] deletable=true editable=true
# Create a `found_vermin` variable for any `VIOLATION DESCRIPTION` containing *vermin*, *mouse*, *mice*, or *rat*.
#
# + deletable=true editable=true
# + [markdown] deletable=true editable=true
# Create a `found_bugs` variable for any `VIOLATION DESCRIPTION` containing *insect*, *roach*, or *flies*.
# + deletable=true editable=true
# + deletable=true editable=true
# + deletable=true editable=true
# + [markdown] deletable=true editable=true
# ## Part 4: Create a Working Subset
#
# 1. Create a working subset DataFrame called `rest_df` with data grouped by restaurant - take the max value for the following fields: `'CAMIS', 'DBA', 'BORO', 'BUILDING', 'STREET', 'ZIPCODE', 'PHONE', 'CUISINE DESCRIPTION', 'inspection_datetime',` and `'inspection_year'`.
# 2. Create another working subset DataFrame called `violation_df` with data grouped by restaurant - take the sum value for `'found_vermin'` and `'found_bugs'`.
# 3. Merge `rest_df` with `violation_df` to create `new_df`.
# 4. Show the top 20 value_counts for `CUISINE DESCRIPTION`.
# 5. Use the `cuisine_dict` to create a `cuisine_new` column with the `CUISINE DESCRIPTION`
# 6. Replace the `CUISINE DESCRIPTION` for `Café/Coffee/Tea` with `Coffee`.
# + [markdown] deletable=true editable=true
# Create a working subset DataFrame called `rest_df` with data grouped by restaurant - take the max value for the following fields: `'CAMIS', 'DBA', 'BORO', 'BUILDING', 'STREET', 'ZIPCODE', 'PHONE', 'CUISINE DESCRIPTION', 'inspection_datetime',` and `'inspection_year'`.
#
# + deletable=true editable=true
# + [markdown] deletable=true editable=true
# Create another working subset DataFrame called `violation_df` with data grouped by restaurant - take the sum value for `'found_vermin'` and `'found_bugs'`.
#
# + deletable=true editable=true
# + [markdown] deletable=true editable=true
# Join `rest_df` with `violation_df` to create `new_df`.
#
# + deletable=true editable=true
# + [markdown] deletable=true editable=true
# Show the top 20 value_counts for `CUISINE DESCRIPTION`.
# + deletable=true editable=true
# + [markdown] deletable=true editable=true
# Replace the `CUISINE DESCRIPTION` for `Café/Coffee/Tea` with `Coffee`.
# + deletable=true editable=true
# + deletable=true editable=true
# + deletable=true editable=true
# + [markdown] deletable=true editable=true
# ## Bonus Round: Using Outside Resources to Clean Data
#
# Oftentimes, external services - or even services from other teams within your own company - will exist to help process data. One handy example case we can use here is the [NYC Geoclient](https://api.cityofnewyork.us/geoclient/v1/doc), a REST api that returns location information for an arbitrary address in New York City. It's an awesome resource!
#
# For the purposes of this exercise, I've included an API id below and gave you the key in class, but you can sign up for your own key at the [NYC Developer Portal](https://developer.cityofnewyork.us/).
#
# We can use this to find the exact location for each coffee shop in our data set.
#
# 1. First, create a function to return the latitude and longitude for a given building number, street address, borough, and zip code.
# 2. Next, create a new subset of data for a single cuisine.
# 3. Apply the function from Step 1 to the df from Step 2.
#
# + deletable=true editable=true
def get_coordinates(row):
url = 'https://api.cityofnewyork.us/geoclient/v1/address.json'
params = {
'houseNumber' : row['BUILDING'],
'street' : row['STREET'],
'borough' : row['BORO'],
'zip' : row['ZIPCODE'],
'app_id' : '7cc1b653',
'app_key' : '<KEY>',
}
raw_response = requests.get(url, params)
try:
lat = raw_response.json()['address']['latitude']
long = raw_response.json()['address']['longitude']
value = str(lat) + ',' + str(long)
except KeyError:
value = None
return value
# + deletable=true editable=true
cuisine_df = new_df.loc[new_df['CUISINE DESCRIPTION']=='Ice Cream, Gelato, Yogurt, Ices', ]
# + deletable=true editable=true
cuisine_df['coordinates'] = cuisine_df.apply(get_coordinates, axis=1)
# + deletable=true editable=true
cuisine_df['latitude'] = cuisine_df.coordinates.str.split(',').str.get(0).astype(float)
cuisine_df['longitude'] = cuisine_df.coordinates.str.split(',').str.get(1).astype(float)
# + deletable=true editable=true
cuisine_df.plot.scatter('latitude', 'longitude')
# + deletable=true editable=true
# + deletable=true editable=true
# + deletable=true editable=true
# + deletable=true editable=true
# + deletable=true editable=true
# + deletable=true editable=true
# + deletable=true editable=true
# + deletable=true editable=true
# + deletable=true editable=true
| lecture-3/DataDive-Lecture3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# This notebook shows the scripts to load data and plotting the Polarimetric phase curve.
#
# %config InlineBackend.figure_format = 'retina'
InteractiveShell.ast_node_interactivity = 'last_expr'
# +
from io import StringIO
from pathlib import Path
import numpy as np
import pandas as pd
from scipy.optimize import curve_fit
from matplotlib import pyplot as plt
from matplotlib import rcParams
import ysvisutilpy2005ud as yvu
import polutil2005ud as util
# -
# We need to do it in a separate cell. See:
# https://github.com/jupyter/notebook/issues/3385
plt.style.use('default')
rcParams.update({'font.size':15})
# +
dats_ast = util.dats_ast
dats = util.dats
fitfunc = util.trigp
DATAPATH = util.DATAPATH
SAVEPATH = util.SAVEPATH
alpha_msi = util.alpha_msi
polr_msi = util.polr_msi
dpolr_msi = util.dpolr_msi
alpha_oth = util.alpha_oth
polr_oth = util.polr_oth
dpolr_oth = util.dpolr_oth
popt_ud, _, pol_ud, _ = util.cfit_pol(fitfunc, "trigp_f", util.dats, use_error=True)
xx = np.arange(0.1, 160, 0.1)
xx_plot = np.arange(0.1, 120, 0.1)
pol_g = dats_ast.groupby("reference")
phae_2016 = dats_ast.loc[(dats_ast['label'] == "Phaethon")
&(dats_ast['reference'].isin(["2018NatCo...9.2486I"]))]
phae_2017 = dats_ast.loc[(dats_ast['label'] == "Phaethon")
&(dats_ast['reference'].isin(["2018ApJ...864L..33S", "2018MNRAS.479.3498D"]))]
popt_2016, _, pol_phae_2016, _ = util.cfit_pol(fitfunc, "trigp_f", phae_2016, use_error=True)
popt_2017, _, pol_phae_2017, _ = util.cfit_pol(fitfunc, "trigp_f", phae_2017, use_error=True)
kws = {
# B-type
"2018ApJ...864L..33S": dict(label="Phaethon 2017 (Shinnaka+18)", marker='D', mfc='none', ms=4, color='dodgerblue', alpha=0.8),
"2018MNRAS.479.3498D": dict(label="Phaethon 2017 (Devogèle+18)", marker='s', mfc='none', ms=5, color='dodgerblue', alpha=0.8),
"2018NatCo...9.2486I": dict(label="Phaethon 2016 (Ito+18)" , marker='P', mfc='none', ms=7, color='b', alpha=1),
# S-types
"1995Icar..113..200L": dict(label="Toutatis (Lupishko+95)", marker='1', mfc='none', ms=6, color='gray'),
"1997Icar..127..452M": dict(label="Toutatis (Mukai+97)" , marker='2', mfc='none', ms=6, color='gray'),
"1997PASJ...49L..31I": dict(label="Toutatis (Ishiguro+97)", marker='3', mfc='none', ms=6, color='gray'),
"2017AJ....154..180I": dict(label="Icarus (Ishiguro+17)", marker='D', mfc='none', ms=5, color='lime'),
# C-types
"2018MNRAS.481L..49C": dict(label="Bennu (Cellino+18)" , marker='s', mfc='none', ms=5, color='k'),
"2018A&A...611A..31K": dict(label="1999 KU2 (Kuroda+18)" , marker='d', mfc='none', ms=5, color='k'),
"1999Icar..140..464K": dict(label="Ra-Shalom (Kiselev+99)" , marker='D', mfc='none', ms=5, color='k'),
"KurodaD2021arXiv": dict(label="Ryugu (Kuroda+21)" , marker='s', mfc='k', ms=3, color='k'),
}
# +
fig, axs = plt.subplots(1, 1, figsize=(9, 9),
sharex=False, sharey=False, gridspec_kw=None)
ax0 = axs
ax1 = ax0.twinx() # ghost axes
ax1.axis('off')
axi = ax0.inset_axes([0.65, 0.28, 0.32, 0.30])
util.plot_data(
np.array([ax0, axi]),
ylims=[(-5, 65), (-2, 4)],
mkw_msi=dict(color='r', marker='o', ms=8, mfc='none', label="2005 UD (MSI)"),
mkw_oth=dict(color='r', marker='s', ms=8, mfc='none', label="2005 UD (Others)")
)
[ax.plot(xx_plot, fitfunc(xx_plot, *popt_ud), color='r') for ax in [ax0, axi]]
for ref, kw in kws.items():
g = pol_g.get_group(ref)
ax0.plot(g.alpha, g.Pr, ls='', **kws[ref])
# axi.plot(g.alpha, g.Pr, ls='', **kws[ref])
if kw['label'].startswith("Phaethon"):
axi.errorbar(g.alpha, g.Pr, g.dPr, **kws[ref])
for popt, ls, c, y in zip([popt_2016, popt_2017], ['-', '--'], ['b', 'royalblue'], ['2016', '2017']):
[ax.plot(xx_plot[150:], fitfunc(xx_plot[150:], *popt), ls=ls, color=c, lw=0.6) for ax in [ax0, axi]]
ax1.plot(np.nan, np.nan, ls=ls, color=c, lw=0.6, label=f"Phaethon {y}")
# ax0.text(xx_plot[-1] + 5, fitfunc(xx_plot, *popt)[-1], f"Phaethon\n{y}", color=c)
# ax0.text(110, 63, "2005 UD", color='r')
ax1.plot(np.nan, np.nan, 'r-', label=f"2005 UD\n(all data)")
ax0.legend(loc=2, title="Observations", prop={'family': 'monospace', 'size': 12}, framealpha=0.6)
ax1.legend(loc=1, title="Best fits", prop={'family': 'monospace', 'size': 12})
plt.tight_layout()
plt.savefig(SAVEPATH/"phasecurve.pdf")
# -
| polarimetry/Phase_Curve.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Vanishing Gradients
# In this notebook, we will demonstrate the difference between using sigmoid and ReLU nonlinearities in a simple neural network with two hidden layers. This notebook is built off of a minimal net demo done by <NAME> for CS 231n, which you can check out here: http://cs231n.github.io/neural-networks-case-study/
# +
# Setup
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
plt.rcParams['figure.figsize'] = (10.0, 8.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# for auto-reloading external modules
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
# %load_ext autoreload
# %autoreload 2
# -
# generate random data -- not linearly separable
np.random.seed(0)
N = 100 # number of points per class
D = 2 # dimensionality
K = 3 # number of classes
X = np.zeros((N*K, D))
num_train_examples = X.shape[0]
y = np.zeros(N*K, dtype='uint8')
for j in range(K):
ix = range(N*j, N*(j+1))
r = np.linspace(0.0,1,N) # radius
t = np.linspace(j*4,(j+1)*4,N) + np.random.randn(N)*0.2 # theta
X[ix] = np.c_[r*np.sin(t), r*np.cos(t)]
y[ix] = j
fig = plt.figure()
plt.scatter(X[:, 0], X[:, 1], c=y, s=40, cmap=plt.cm.Spectral)
plt.xlim([-1,1])
plt.ylim([-1,1])
# The sigmoid function "squashes" inputs to lie between 0 and 1. Unfortunately, this means that for inputs with sigmoid output close to 0 or 1, the gradient with respect to those inputs are close to zero. This leads to the phenomenon of vanishing gradients, where gradients drop close to zero, and the net does not learn well.
#
# On the other hand, the relu function (max(0, x)) does not saturate with input size. Plot these functions to gain intution.
# +
def sigmoid(x):
x = 1 / (1 + np.exp(-x))
return x
def sigmoid_grad(x):
return (x) * (1 - x)
def relu(x):
return np.maximum(0,x)
# -
# Let's try and see now how the two kinds of nonlinearities change deep neural net training in practice. Below, we build a very simple neural net with three layers (two hidden layers), for which you can swap out ReLU/ sigmoid nonlinearities.
# +
#function to train a three layer neural net with either RELU or sigmoid nonlinearity via vanilla grad descent
def three_layer_net(NONLINEARITY, X, y, model, step_size, reg):
#parameter initialization
h = model['h']
h2= model['h2']
W1= model['W1']
W2= model['W2']
W3= model['W3']
b1= model['b1']
b2= model['b2']
b3= model['b3']
# some hyperparameters
# gradient descent loop
num_examples = X.shape[0]
plot_array_1=[]
plot_array_2=[]
for i in range(50000):
#FOWARD PROP
if NONLINEARITY== 'RELU':
hidden_layer = relu(np.dot(X, W1) + b1)
hidden_layer2 = relu(np.dot(hidden_layer, W2) + b2)
scores = np.dot(hidden_layer2, W3) + b3
elif NONLINEARITY == 'SIGM':
hidden_layer = sigmoid(np.dot(X, W1) + b1)
hidden_layer2 = sigmoid(np.dot(hidden_layer, W2) + b2)
scores = np.dot(hidden_layer2, W3) + b3
exp_scores = np.exp(scores)
probs = exp_scores / np.sum(exp_scores, axis=1, keepdims=True) # [N x K]
#print(X.shape)
#print(scores.shape)
#print(np.sum(exp_scores, axis=1, keepdims=True).shape)
#print(probs.shape)
#assert False
# compute the loss: average cross-entropy loss and regularization
# v = probs[range(num_examples), y] -> 1d vector v[i] = probs[i, y[i]]]
corect_logprobs = -np.log(probs[range(num_examples), y])
data_loss = np.sum(corect_logprobs) / num_examples
reg_loss = 0.5*reg*np.sum(W1*W1) + 0.5*reg*np.sum(W2*W2)+ 0.5*reg*np.sum(W3*W3)
loss = data_loss + reg_loss
if i % 1000 == 0:
print("iteration %d: loss %f" % (i, loss))
# compute the gradient on scores
dscores = probs
dscores[range(num_examples), y] -= 1
dscores /= num_examples
# BACKPROP HERE
dW3 = (hidden_layer2.T).dot(dscores)
db3 = np.sum(dscores, axis=0, keepdims=True)
if NONLINEARITY == 'RELU':
#backprop ReLU nonlinearity here
dhidden2 = np.dot(dscores, W3.T)
dhidden2[hidden_layer2 <= 0] = 0
dW2 = np.dot( hidden_layer.T, dhidden2)
plot_array_2.append(np.sum(np.abs(dW2)) / np.sum(np.abs(dW2.shape)))
db2 = np.sum(dhidden2, axis=0)
dhidden = np.dot(dhidden2, W2.T)
dhidden[hidden_layer <= 0] = 0
elif NONLINEARITY == 'SIGM':
#backprop sigmoid nonlinearity here
dhidden2 = dscores.dot(W3.T)*sigmoid_grad(hidden_layer2)
dW2 = (hidden_layer.T).dot(dhidden2)
plot_array_2.append(np.sum(np.abs(dW2))/np.sum(np.abs(dW2.shape)))
db2 = np.sum(dhidden2, axis=0)
dhidden = dhidden2.dot(W2.T)*sigmoid_grad(hidden_layer)
dW1 = np.dot(X.T, dhidden)
plot_array_1.append(np.sum(np.abs(dW1))/np.sum(np.abs(dW1.shape)))
db1 = np.sum(dhidden, axis=0)
# add regularization
dW3 += reg * W3
dW2 += reg * W2
dW1 += reg * W1
#option to return loss, grads -- uncomment next comment
grads={}
grads['W1']=dW1
grads['W2']=dW2
grads['W3']=dW3
grads['b1']=db1
grads['b2']=db2
grads['b3']=db3
#return loss, grads
# update
W1 += -step_size * dW1
b1 += -step_size * db1
W2 += -step_size * dW2
b2 += -step_size * db2
W3 += -step_size * dW3
b3 += -step_size * db3
# evaluate training set accuracy
if NONLINEARITY == 'RELU':
hidden_layer = relu(np.dot(X, W1) + b1)
hidden_layer2 = relu(np.dot(hidden_layer, W2) + b2)
elif NONLINEARITY == 'SIGM':
hidden_layer = sigmoid(np.dot(X, W1) + b1)
hidden_layer2 = sigmoid(np.dot(hidden_layer, W2) + b2)
scores = np.dot(hidden_layer2, W3) + b3
predicted_class = np.argmax(scores, axis=1)
print('training accuracy: %.2f' % (np.mean(predicted_class == y)))
#return cost, grads
return plot_array_1, plot_array_2, W1, W2, W3, b1, b2, b3
# -
# #### Train net with sigmoid nonlinearity first
# +
#Initialize toy model, train sigmoid net
N = 100 # number of points per class
D = 2 # dimensionality
K = 3 # number of classes
h=50
h2=50
num_train_examples = X.shape[0]
model={}
model['h'] = h # size of hidden layer 1
model['h2']= h2# size of hidden layer 2
model['W1']= 0.1 * np.random.randn(D,h)
model['b1'] = np.zeros((1,h))
model['W2'] = 0.1 * np.random.randn(h,h2)
model['b2']= np.zeros((1,h2))
model['W3'] = 0.1 * np.random.randn(h2,K)
model['b3'] = np.zeros((1,K))
(sigm_array_1, sigm_array_2, s_W1, s_W2,s_W3, s_b1, s_b2,s_b3) = three_layer_net('SIGM', X,y,model, step_size=1e-1, reg=1e-3)
# -
# #### Now train net with ReLU nonlinearity
# +
#Re-initialize model, train relu net
model={}
model['h'] = h # size of hidden layer 1
model['h2']= h2# size of hidden layer 2
model['W1']= 0.1 * np.random.randn(D,h)
model['b1'] = np.zeros((1,h))
model['W2'] = 0.1 * np.random.randn(h,h2)
model['b2']= np.zeros((1,h2))
model['W3'] = 0.1 * np.random.randn(h2,K)
model['b3'] = np.zeros((1,K))
(relu_array_1, relu_array_2, r_W1, r_W2,r_W3, r_b1, r_b2,r_b3) = three_layer_net('RELU', X,y,model, step_size=1e-1, reg=1e-3)
# -
# # The Vanishing Gradient Issue
# We can use the sum of the magnitude of gradients for the weights between hidden layers as a cheap heuristic to measure speed of learning (you can also use the magnitude of gradients for each neuron in the hidden layer here). Intuitevely, when the magnitude of the gradients of the weight vectors or of each neuron are large, the net is learning faster. (NOTE: For our net, each hidden layer has the same number of neurons. If you want to play around with this, make sure to adjust the heuristic to account for the number of neurons in the layer).
plt.plot(np.array(sigm_array_1))
plt.plot(np.array(sigm_array_2))
plt.title('Sum of magnitudes of gradients -- SIGM weights')
plt.legend(("sigm first layer", "sigm second layer"))
plt.plot(np.array(relu_array_1))
plt.plot(np.array(relu_array_2))
plt.title('Sum of magnitudes of gradients -- ReLU weights')
plt.legend(("relu first layer", "relu second layer"))
# Overlaying the two plots to compare
plt.plot(np.array(relu_array_1))
plt.plot(np.array(relu_array_2))
plt.plot(np.array(sigm_array_1))
plt.plot(np.array(sigm_array_2))
plt.title('Sum of magnitudes of gradients -- hidden layer neurons')
plt.legend(("relu first layer", "relu second layer","sigm first layer", "sigm second layer"))
# #### Feel free to play around with this notebook to gain intuition. Things you might want to try:
#
# - Adding additional layers to the nets and seeing how early layers continue to train slowly for the sigmoid net
# - Experiment with hyperparameter tuning for the nets -- changing regularization and gradient descent step size
# - Experiment with different nonlinearities -- Leaky ReLU, Maxout. How quickly do different layers learn now?
#
#
# We can see how well each classifier does in terms of distinguishing the toy data classes. As expected, since the ReLU net trains faster, for a set number of epochs it performs better compared to the sigmoid net.
# plot the classifiers- SIGMOID
h = 0.02
x_min, x_max = X[:, 0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
Z = np.dot(sigmoid(np.dot(sigmoid(np.dot(np.c_[xx.ravel(), yy.ravel()], s_W1)
+ s_b1), s_W2) + s_b2), s_W3) + s_b3
Z = np.argmax(Z, axis=1)
Z = Z.reshape(xx.shape)
fig = plt.figure()
plt.contourf(xx, yy, Z, cmap=plt.cm.Spectral, alpha=0.8)
plt.scatter(X[:, 0], X[:, 1], c=y, s=40, cmap=plt.cm.Spectral)
plt.xlim(xx.min(), xx.max())
plt.ylim(yy.min(), yy.max())
# plot the classifiers-- RELU
h = 0.02
x_min, x_max = X[:, 0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
Z = np.dot(relu(np.dot(relu(np.dot(np.c_[xx.ravel(), yy.ravel()], r_W1)
+ r_b1), r_W2) + r_b2), r_W3) + r_b3
Z = np.argmax(Z, axis=1)
Z = Z.reshape(xx.shape)
fig = plt.figure()
plt.contourf(xx, yy, Z, cmap=plt.cm.Spectral, alpha=0.8)
plt.scatter(X[:, 0], X[:, 1], c=y, s=40, cmap=plt.cm.Spectral)
plt.xlim(xx.min(), xx.max())
plt.ylim(yy.min(), yy.max())
| vanishing_grad_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + run_control={"frozen": false, "read_only": false}
# %matplotlib inline
# %load_ext ipycache
import pandas as pd
import numpy as np
import scipy
import sklearn as sk
import xgboost as xgb
from eli5 import show_weights
import seaborn as sns
sns.set()
import matplotlib.pyplot as plt
# + run_control={"frozen": false, "read_only": false}
import math
#A function to calculate Root Mean Squared Logarithmic Error (RMSLE)
def rmsle(y, y_pred):
assert len(y) == len(y_pred)
terms_to_sum = [
(math.log(y_pred[i] + 1) - math.log(y[i] + 1)) ** 2.0
for i,pred in enumerate(y_pred)
]
return (sum(terms_to_sum) * (1.0/len(y))) ** 0.5
def rmse(y, y_pred):
return np.sqrt(((y_pred - y) ** 2).mean())
# + [markdown] run_control={"frozen": false, "read_only": false}
# # Препроцессинг фич
# + run_control={"frozen": false, "read_only": false}
# train_raw = pd.read_csv("data/train.csv")
train_raw = pd.read_csv("data/train_without_noise.csv")
test = pd.read_csv("data/test.csv")
macro = pd.read_csv("data/macro.csv")
train_raw.head()
# + run_control={"frozen": false, "read_only": false}
def preprocess_anomaly(df):
df["full_sq"] = map(lambda x: x if x > 10 else float("NaN"), df["full_sq"])
df["life_sq"] = map(lambda x: x if x > 5 else float("NaN"), df["life_sq"])
df["kitch_sq"] = map(lambda x: x if x > 2 else float("NaN"), df["kitch_sq"])
# superclean
# https://www.kaggle.com/keremt/very-extensive-cleaning-by-sberbank-discussions
df.ix[df[df.life_sq > df.full_sq].index, "life_sq"] = np.NaN
df.ix[df[df.kitch_sq >= df.life_sq].index, "kitch_sq"] = np.NaN
df.ix[df[df.kitch_sq == 0].index, "kitch_sq"] = np.NaN
df.ix[df[df.kitch_sq == 1].index, "kitch_sq"] = np.NaN
df.ix[df[df.build_year < 1500].index, "build_year"] = np.NaN
df.ix[df[df.build_year > 1500].index, "build_year"] = np.NaN
df.ix[df[df.num_room == 0].index, "num_room"] = np.NaN
df.ix[df[df.floor == 0].index, "floor"] = np.NaN
df.ix[df[df.max_floor == 0].index, "max_floor"] = np.NaN
df.ix[df[df.floor > df.max_floor].index, "max_floor"] = np.NaN
df.ix[df[df.state == 33].index, "state"] = np.NaN
return df
# + code_folding=[] run_control={"frozen": false, "read_only": false}
def preprocess_categorial(df):
# df = mess_y_categorial(df, 5)
for c in df.columns:
if df[c].dtype == 'object':
lbl = sk.preprocessing.LabelEncoder()
lbl.fit(list(train_raw[c].values) + list(test[c].values))
df[c] = lbl.transform(list(df[c].values))
df = df.select_dtypes(exclude=['object'])
return df
def apply_categorial(test, train):
# test = mess_y_categorial_fold(test, train)
# test = test.select_dtypes(exclude=['object'])
return preprocess_categorial(test)
def smoothed_likelihood(targ_mean, nrows, globalmean, alpha=10):
try:
return (targ_mean * nrows + globalmean * alpha) / (nrows + alpha)
except Exception:
return float("NaN")
def mess_y_categorial(df, nfolds=3, alpha=10):
from sklearn.utils import shuffle
from copy import copy
folds = np.array_split(shuffle(df), nfolds)
newfolds = []
for i in range(nfolds):
fold = folds[i]
other_folds = copy(folds)
other_folds.pop(i)
other_fold = pd.concat(other_folds)
newfolds.append(mess_y_categorial_fold(fold, other_fold, alpha=10))
return pd.concat(newfolds)
def mess_y_categorial_fold(fold_raw, other_fold, cols=None, y_col="price_doc", alpha=10):
fold = fold_raw.copy()
if not cols:
cols = list(fold.select_dtypes(include=["object"]).columns)
globalmean = other_fold[y_col].mean()
for c in cols:
target_mean = other_fold[[c, y_col]].groupby(c).mean().to_dict()[y_col]
nrows = other_fold[c].value_counts().to_dict()
fold[c + "_sll"] = fold[c].apply(
lambda x: smoothed_likelihood(target_mean.get(x), nrows.get(x), globalmean, alpha) if x else float("NaN")
)
return fold
# + run_control={"frozen": false, "read_only": false}
def apply_macro(df):
macro_cols = [
'timestamp', "balance_trade", "balance_trade_growth", "eurrub", "average_provision_of_build_contract",
"micex_rgbi_tr", "micex_cbi_tr", "deposits_rate", "mortgage_value", "mortgage_rate",
"income_per_cap", "rent_price_4+room_bus", "museum_visitis_per_100_cap", "apartment_build"
]
return pd.merge(df, macro, on='timestamp', how='left')
# + run_control={"frozen": false, "read_only": false}
def preprocess(df):
from sklearn.preprocessing import OneHotEncoder, FunctionTransformer
# df = apply_macro(df)
# df["timestamp_year"] = df["timestamp"].apply(lambda x: x.split("-")[0])
# df["timestamp_month"] = df["timestamp"].apply(lambda x: x.split("-")[1])
# df["timestamp_year_month"] = df["timestamp"].apply(lambda x: x.split("-")[0] + "-" + x.split("-")[1])
ecology = ["no data", "poor", "satisfactory", "good", "excellent"]
df["ecology_index"] = map(ecology.index, df["ecology"].values)
bool_feats = [
"thermal_power_plant_raion",
"incineration_raion",
"oil_chemistry_raion",
"radiation_raion",
"railroad_terminal_raion",
"big_market_raion",
"nuclear_reactor_raion",
"detention_facility_raion",
"water_1line",
"big_road1_1line",
"railroad_1line",
"culture_objects_top_25"
]
for bf in bool_feats:
df[bf + "_bool"] = map(lambda x: x == "yes", df[bf].values)
df = preprocess_anomaly(df)
df['rel_floor'] = df['floor'] / df['max_floor'].astype(float)
df['rel_kitch_sq'] = df['kitch_sq'] / df['full_sq'].astype(float)
df['rel_life_sq'] = df['life_sq'] / df['full_sq'].astype(float)
df["material_cat"] = df.material.fillna(0).astype(int).astype(str).replace("0", "")
df["state_cat"] = df.state.fillna(0).astype(int).astype(str).replace("0", "")
df["num_room_cat"] = df.num_room.fillna(0).astype(int).astype(str).replace("0", "")
df = df.drop(["id", "timestamp"], axis=1)
return df
# + run_control={"frozen": false, "read_only": false}
train_pr = preprocess(train_raw)
train = preprocess_categorial(train_pr)
# train = train.fillna(-1)
X = train.drop(["price_doc"], axis=1)
y = train["price_doc"].values
# + [markdown] run_control={"frozen": false, "read_only": false}
# # Обучение моделей
# + run_control={"frozen": false, "read_only": false}
from sklearn.model_selection import train_test_split
X_train, X_val, y_train, y_val = train_test_split(X.values, y, test_size=0.20, random_state=43)
dtrain_all = xgb.DMatrix(X.values, y, feature_names=X.columns)
dtrain = xgb.DMatrix(X_train, y_train, feature_names=X.columns)
dval = xgb.DMatrix(X_val, y_val, feature_names=X.columns)
# + run_control={"frozen": false, "read_only": false}
xgb_params = {
'max_depth': 5,
'n_estimators': 200,
'learning_rate': 0.01,
'objective': 'reg:linear',
'eval_metric': 'rmse',
'silent': 1
}
# Uncomment to tune XGB `num_boost_rounds`
model = xgb.train(xgb_params, dtrain, num_boost_round=4000, evals=[(dval, 'val')],
early_stopping_rounds=40, verbose_eval=40)
num_boost_round = model.best_iteration
# + run_control={"frozen": false, "read_only": false}
cv_output = xgb.cv(dict(xgb_params, silent=0), dtrain_all, num_boost_round=num_boost_round, verbose_eval=40)
cv_output[['train-rmse-mean', 'test-rmse-mean']].plot()
# + run_control={"frozen": false, "read_only": false}
model = xgb.train(dict(xgb_params, silent=0), dtrain_all, num_boost_round=num_boost_round, verbose_eval=40)
print "predict-train:", rmse(model.predict(dtrain_all), y)
# + run_control={"frozen": true, "read_only": true}
# model = xgb.XGBRegressor(max_depth=5, n_estimators=100, learning_rate=0.01, nthread=-1, silent=False)
# model.fit(X.values, y, verbose=20)
#
# with open("scores.tsv", "a") as sf:
# sf.write("%s\n" % rmsle(model.predict(X.values), y))
#
# !tail scores.tsv
# + run_control={"frozen": true, "read_only": true}
# show_weights(model, feature_names=list(X.columns), importance_type="weight")
# + run_control={"frozen": true, "read_only": true}
# from sklearn.model_selection import cross_val_score
# from sklearn.metrics import make_scorer
#
# def validate(clf):c
# cval = np.abs(cross_val_score(clf, X.values, y, cv=3,
# scoring=make_scorer(rmsle, False), verbose=2))
# return np.mean(cval), cval
#
# print validate(model)
# + [markdown] run_control={"frozen": false, "read_only": false}
# # Submission
# + run_control={"frozen": false, "read_only": false}
test_pr = preprocess(test)
test_pr = apply_categorial(test_pr, train_pr)
# test_pr = test_pr.fillna(-1)
dtest = xgb.DMatrix(test_pr.values, feature_names=test_pr.columns)
y_pred = model.predict(dtest)
# y_pred = model.predict(test_pr.values)
# y_pred = np.exp(y_pred) - 1
submdf = pd.DataFrame({"id": test["id"], "price_doc": y_pred})
submdf.to_csv("data/submission.csv", header=True, index=False)
# !head data/submission.csv
# + [markdown] run_control={"frozen": false, "read_only": false}
# without noise xgb logarithmic y
#
# val-rmse:0.478924
#
# macro 10*400
#
# val-rmse:0.480618
#
# macro 5*200
#
# val-rmse:0.476849
#
# macro 5*200 no month and year
#
# val-rmse:0.477861
#
# macro 5*200 no month and year
#
# val-rmse:0.473012
#
# macro 5*200 no month and year
#
# val-rmse:0.471758
# predict-train: 0.427215115875
#
# macro 5*200 no month and year, train_without_noise
#
# val-rmse:0.461684
# train-rmse:0.411116+0.00299259 test-rmse:0.472202+0.00166791
# predict-train: 0.423849149218
# kaggle: 0.36027
#
# 5*200, no macro no add features, train_without_noise:
#
# val-rmse:0.471989
# train-rmse:0.425924+0.00643495 test-rmse:0.473873+0.0131213
# predict-train: 0.43508730101
#
# 5*200, no macro add rel features, train_without_noise:
#
# val-rmse:0.471808
# train-rmse:0.425264+0.00595741 test-rmse:0.47383+0.0130655
# predict-train: 0.435635092773
# kaggle: 0.32837
#
# 5*200, no macro, add rel features, no log price, train_without_noise:
#
# val-rmse:2.63772e+06
# train-rmse:1.9989e+06+10986.4 test-rmse:2.69158e+06+53020
# predict-train: 2076010.27131
# kaggle: 0.31720
#
# 5*200, no macro, add rel features, no log price, train_with_noise:
#
# val-rmse:2.53378e+06
# train-rmse:1.95069e+06+16166.4 test-rmse:2.69703e+06+61455.1
# predict-train: 2054421.59869
# kaggle: 0.32056
#
# 5*200, macro, add rel features, no log price, train_without_noise:
#
# val-rmse:2.79632e+06
# train-rmse:1.81015e+06+19781.2 test-rmse:2.6641e+06+123875
# predict-train: 1904063.27368
# kaggle: 0.32976
#
# 5*200, no macro, add rel features, no log price, train_without_noise:
#
# val-rmse:2.61682e+06
# train-rmse:1.81123e+06+27681.2 test-rmse:2.66923e+06+53925.7
# predict-train: 1899129.43771
# kaggle: 0.31592
#
# 5*200, no macro, add rel features, no log price, train_without_noise, 4000 iter:
#
# val-rmse:2.61055e+06
# train-rmse:1.71826e+06+30076.1 test-rmse:2.66515e+06+54583.5
# predict-train: 1814572.97424
# kaggle: 0.31602
#
# 7*300, no macro, add rel features, no log price, train_without_noise, 4000 iter:
#
# val-rmse:2.59955e+06
# train-rmse:1.41393e+06+21208.1 test-rmse:2.6763e+06+35553.3
# predict-train: 1548257.49121
# kaggle: 0.31768
#
# 4*300, no macro, add rel features, no log price, train_without_noise, 4000 iter:
#
# val-rmse:2.63407e+06
# train-rmse:1.96513e+06+21470.8 test-rmse:2.69417e+06+74288.3
# predict-train: 2062299.41091
# kaggle: 0.31952
#
# 7*200, no macro, add rel features, no log price, train_without_noise, 4000 iter:
#
# val-rmse:2.59955e+06
# train-rmse:1.41393e+06+21208.1 test-rmse:2.6763e+06+35553.3
# predict-train: 1548257.49121
#
# 5*300, no macro, add rel features, no log price, train_without_noise, 4000 iter:
#
# val-rmse:2.61055e+06
# train-rmse:1.71826e+06+30076.1 test-rmse:2.66515e+06+54583.5
# predict-train: 1814572.97424
#
# 5*200, no macro, add rel features, no log price, train_without_noise, 4000 iter, not fillna:
#
# val-rmse:2.61664e+06
# train-rmse:1.77892e+06+23111 test-rmse:2.65829e+06+56398.6
# predict-train: 1875799.54634
# kaggle: 0.31521
#
# 5*200, no macro, add rel features, no log price, train_without_noise, 4000 iter, not fillna, superclean:
#
# val-rmse:2.6265e+06
# train-rmse:1.78478e+06+22545.4 test-rmse:2.66179e+06+60626.3
# predict-train: 1881672.27588
#
# + run_control={"frozen": false, "read_only": false}
| sber/Model-0.31434.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import gym
import numpy as np
import torch
from matplotlib import pyplot as plt
env = gym.make('CartPole-v0')
l1 = 4
l2 = 150
l3 = 2
model = torch.nn.Sequential(
torch.nn.Linear(l1, l2),
torch.nn.LeakyReLU(),
torch.nn.Linear(l2, l3),
torch.nn.Softmax()
)
learning_rate = 0.0009
optimizer = torch.optim.Adam(model.parameters(), lr = learning_rate)
# -
torch.arange(5).float()
torch.pow(0.9, torch.arange(5).float())
state1 = env.reset()
pred = model(torch.from_numpy(state1).float())
action = np.random.choice(np.array([0,1]), p = pred.data.numpy())
state2, reward, done, info = env.step(action)
def discount_rewards(rewards, gamma = 0.99):
lenr = len(rewards)
disc_return = torch.pow(gamma, torch.arange(lenr).float())*rewards
disc_return/=disc_return.max()
return disc_return
def loss_fn(preds, r):
return -1*torch.sum(r*torch.log(preds))
MAX_DUR = 200
MAX_EPISODES = 500
gamma = 0.99
score = []
expectation = 0.0
for episode in range(MAX_EPISODES):
curr_state = env.reset()
done = False
transitions = []
for t in range(MAX_DUR):
act_prob = model(torch.from_numpy(curr_state).float())
action = np.random.choice(np.array([0,1]), p=act_prob.data.numpy())
prev_state = curr_state
curr_state, _, done, info = env.step(action)
transitions.append((prev_state, action, t+1))
if done:
break
ep_len = len(transitions)
score.append(ep_len)
reward_batch = torch.Tensor([r for (s,a,r) in transitions]).flip(dims=(0,))
disc_returns = discount_rewards(reward_batch)
state_batch = torch.Tensor([s for (s,a,r) in transitions])
action_batch = torch.Tensor([a for (s,a,r) in transitions])
pred_batch = model(state_batch)
prob_batch = pred_batch.gather(dim=1,index=action_batch.long().view(-1,1)).squeeze()
loss = loss_fn(prob_batch, disc_returns)
optimizer.zero_grad()
loss.backward()
optimizer.step()
def running_mean(x, N=50):
kernel = np.ones(N)
conv_len = x.shape[0]-N
y = np.zeros(conv_len)
for i in range(conv_len):
y[i] = kernel @ x[i:i+N]
y[i] /= N
return y
score = np.array(score)
avg_score = running_mean(score, 50)
plt.figure(figsize=(10,7))
plt.ylabel("Episode Duration",fontsize=22)
plt.xlabel("Training Epochs",fontsize=22)
plt.plot(avg_score, color='green')
| Chapter 4/Ch4_prac.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Not Filter Demo
#
# Example how to wrap a filter in a not filter to negate a filter
#
# ## Imports
from pyspark import SparkConf, SparkContext
from mmtfPyspark.io import mmtfReader
from mmtfPyspark.filters import ContainsDnaChain, ContainsLProteinChain, NotFilter
from mmtfPyspark.structureViewer import view_structure
# ## Configure Spark
conf = SparkConf().setMaster("local[*]") \
.setAppName("notFilterExample")
sc = SparkContext(conf = conf)
# ## Read in MMTF Files
# +
path = "../../resources/mmtf_reduced_sample/"
pdb = mmtfReader.read_sequence_file(path, sc)
# -
# ## Filter by contains L Protein Chain
structures = pdb.filter(ContainsLProteinChain())
# ## Using Not filter to reverse a filter
#
# Get entires that does not contain DNA Chains
structures = structures.filter(NotFilter(ContainsDnaChain()))
# ## Count number of entires
# +
count = structures.count()
print(f"PDB entires without DNA chains : {count}")
# -
# ## Visualize Structures
view_structure(structures.keys().collect())
# ## Terminate Spark
sc.stop()
| docs/_static/demos/filters/NotFilterDemo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Допустим, у нас есть $n$ товаров с заданными стоимостями $v_i$ и массой $w_i$. В сумку убирается $С$ кг. Сколько какого товара взять, чтобы сумма всех стоимостей товаров была наибольшей?
values = [4, 2, 1, 7, 3, 6]
weights = [5, 9, 8, 2, 6, 5]
C = 15
n = 6
# Сформулируем задачу:
# $$\max\sum v_i x_i$$
# $$\sum w_i x_i \leq C $$
# Как должна выглядеть задача:
# $$\min c^T x$$
# $$A x \leq b $$
# Получается, что $c=-v$, $A=w^T$, $b=(C)$
# +
import numpy as np
c = - np.array(values)
A = np.array(weights, ndmin=2)
b = np.array([C])
# -
linprog(c=c, A_ub=A, b_ub=b)
import numpy as np
c = - np.array(values)
A = np.array(weights) #shape = (6,)
A = np.expand_dims(A, 0) #shape = (1,6)
b = np.array([C])
from scipy.optimize import linprog
linprog(c=c, A_ub=A, b_ub=b)
# !pip install cvxopt
import cvxpy
x = cvxpy.Variable(shape=n, integer = True)
constraint = (A @ x <= b)
total_value = c * x
problem = cvxpy.Problem(cvxpy.Minimize(total_value), constraints=[constraint])
problem.solve(solver='ECOS_BB')
x.value
# Теперь положительные $x$
x = cvxpy.Variable(shape=n, integer=True)
constraint = (A @ x <= b)
x_positive = (x >= 0)
total_value = c * x
problem = cvxpy.Problem(cvxpy.Minimize(total_value), constraints=[constraint, x_positive])
problem.solve(solver='ECOS_BB')
x.value
# Теперь $x = 0$ или $1$
x = cvxpy.Variable(shape=n, boolean=True)
constraint = A @ x <= b
x_positive = x >= 0
total_value = c * x
problem = cvxpy.Problem(cvxpy.Minimize(total_value), constraints=[constraint, x_positive])
problem.solve(solver='ECOS_BB')
x.value
# 
c = np.array([[2, 5, 3], [7, 7, 6]])
x = cvxpy.Variable(shape=c.shape, integer=True)
constraint = [
cvxpy.sum(x[0]) <= 180,
cvxpy.sum(x[1]) <= 220,
cvxpy.sum(x[:, 0]) == 110,
cvxpy.sum(x[:, 1]) == 150,
cvxpy.sum(x[:, 2]) == 140,
x >= 0
]
total_value = cvxpy.sum(cvxpy.multiply(c, x))
problem = cvxpy.Problem(cvxpy.Minimize(total_value), constraints=constraint)
problem.solve(solver='ECOS_BB')
# 
c = np.array([[1000, 12, 10, 19, 8],
[12, 1000, 3, 7, 2],
[10, 3, 1000, 6, 20],
[19, 7, 6, 1000, 4],
[8, 2, 20, 4, 1000]])
x = cvxpy.Variable(shape=c.shape, boolean=True)
# +
constraint = [
cvxpy.sum(x, axis=0) == np.ones(5),
cvxpy.sum(x, axis=1) == np.ones(5)
]
total_value = cvxpy.sum(cvxpy.multiply(c, x))
# -
problem = cvxpy.Problem(cvxpy.Minimize(total_value), constraints=constraint)
problem.solve(solver='ECOS_BB')
| module_5/Module_suppl_notebooks/5.6.Math&Stat6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Image Browser
# This example shows how to browse through a set of images with a slider.
# %matplotlib inline
import matplotlib.pyplot as plt
from IPython.html.widgets import interact
from sklearn import datasets
# We will use the digits dataset from [scikit-learn](http://scikit-learn.org/stable/).
digits = datasets.load_digits()
def browse_images(digits):
n = len(digits.images)
def view_image(i):
plt.imshow(digits.images[i], cmap=plt.cm.gray_r, interpolation='nearest')
plt.title('Training: %s' % digits.target[i])
plt.show()
interact(view_image, i=(0,n-1))
browse_images(digits)
| notebooks/1 - IPython Notebook Examples/IPython Project Examples/Interactive Widgets/Image Browser.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
x = [1,2,3]
y = [2,4,1]
plt.plot(x,y)
plt.xlabel('x-axis')
plt.ylabel('y-axis')
plt.title('First graph')
plt.show()
# +
import numpy as np
import matplotlib as mpl
from matplotlib import pyplot as plt
#parameters
r0 = 1533 #in pc
sigb = 5.12e6 #solar masses per pc^2
x_0=1533.0 #using this instead of r. in parsec
sig_b = lambda x: np.exp(-np.power(x/x_0,(1/4)))
vec_sig_b = np.vectorize(sig_b)
fig,axes = plt.subplots(1,1,figsize=(9.0,8.0),sharex=True)
ax1 = axes
X = np.arange(0.0,10,0.01)
ax1.plot(X,vec_sig_b(X), color='k', linestyle='-', \
label='mass density of bulge', linewidth=2)
ax1.set_yscale('log')
ax1.set_xlim(0, 10) #in pairs
ax1.set_ylim(6e-1,1)
ax1.set_xlabel('disc coordinate [pc]')
ax1.set_ylabel('\Sigma')
ax1.grid(True)
ax1.yaxis.grid(True,which='minor',linestyle='--')
ax1.legend(loc=1,prop={'size':22})
for axis in ['top','bottom','left','right']: #sets the axis linewidth to 2
ax1.spines[axis].set_linewidth(2)
plt.tight_layout()
plt.show()
# +
#notes from practicing plotting
import numpy as np
import matplotlib as mpl
from matplotlib import pyplot as plt
plt.style.use('seaborn-whitegrid')
#parameters for central bulge graph
r0 = 1533 #in pc
sigb = 5.12e6 #solar masses per pc^2
#adjusting the plot
fig = plt.figure()
ax = plt.axes()
x = np.linspace(0, 10, 1000) #x from/to and line smoothness
plt.plot(x, np.sin(x), linestyle='solid', label='sin(x)') #functions
plt.plot(x, np.cos(x), linestyle='dashdot', label='cos(x)')
plt.plot(x, x - 1, ':k', label='x - 1'); #dotted black
plt.axis([-1, 11, -1.5, 1.5]); #xmin, xmax, ymin, ymax
#or use: plt.xlim(10, 0) plt.ylim(1.2, -1.2);
#or to tighten the bounds: plt.axis('tight')
#labeling plots
ax.set(title='A Sine and Cosine Curve', #labeling axes
xlabel='x', ylabel='sin(x)');
plt.legend(); #adding legend
# +
import numpy as np
import matplotlib as mpl
from matplotlib import pyplot as plt
plt.style.use('seaborn-whitegrid')
#parameters for graphs
r_0 = 500 #bulge/characteristic radius of bulge (in pc)
sig_b = 5.12e6 #bulge/central mass density (solar masses per pc^2)
sig_g1 = 11 #gas/peak surface density (solar masses per pc^2)
sig_g2 = 11 #gas/peak surface density (solar masses per pc^2)
r_g1 = 5.0e3 #gas/location 1 (in pc)
r_g2 = 12.0e3 #gas/location 2 (in pc)
w_g1 = 4.0e3 #gas/gaussian width (in pc)
w_g2 = 4.0e3 #gas/gaussian width (in pc)
r_b = 2.0e3 #disk/bulge radius (in pc)
sig_s = 611 #disk/max surface density (solar masses per pc^2)
l_c = 2.5e3 #disk/characteristic lenght (in pc)
#adjusting the plot
fig = plt.figure(figsize=(9.0,8.0)) #size of the plot
ax = plt.axes()
ax.grid(True)
ax.yaxis.grid(True,which='minor',linestyle='--')
ax.set_yscale('log') #setting the axis to log
x = np.linspace(0, 16000, 1000) #x from/to and line smoothness
#functions
plt.plot(x, sig_b*np.exp(-np.power(x/r_0, (1/4))), linestyle='solid', label='Bulge')
plt.plot(x, (sig_g1*np.exp(-np.power(((x-r_g1)/w_g1), 2))) + (sig_g2*np.exp(-np.power(((x-r_g2)/w_g2), 2))), linestyle='dashdot', label='Gas')
plt.plot(x, (sig_s*l_c)/(np.sqrt(np.power((x-r_b), 2) + np.power(l_c, 2))), ':k', label='Stellar Disk'); #dotted black
plt.axis('tight'); #xmin, xmax, ymin, ymax
#or use: plt.xlim(10, 0) plt.ylim(1.2, -1.2);
#or to tighten the bounds: plt.axis('tight')
#or use: plt.axis([0, 15000, 0.5e6, 2.5e6])
#labeling plots
fig.suptitle('Mass Density of Milky Way', fontsize=16)
ax.set(title='"Modelling Mass Distribution of the Milky Way Galaxy Using GAIAs Billion-Star Map" paper') #labeling axes
ax.set(xlabel='Disc Coordinate [pc]', ylabel='Mass Density [Solar Mass/pc^2]');
plt.legend(fontsize=14); #adding legend
# +
import numpy as np
import matplotlib as mpl
from matplotlib import pyplot as plt
plt.style.use('seaborn-whitegrid')
#parameters for graphs
r_0 = 500 #bulge/half-mass scale radius (in pc)
sig_b = 3.2e3 #bulge/surface mass density at half-mass scale radius (solar masses per pc^2)
sig_g1 = 0.17*8.44e2 #rings/peak density (solar masses per pc^2)
sig_g2 = 0.34*8.44e2 #rings/peak density (solar masses per pc^2)
r_g1 = 3e3 #rings/radii of wave nodes (in pc)
r_g2 = 9.5e3 #rings/radii of wave nodes (in pc)
w_g1 = 1.0e3 #rings/width (in pc)
w_g2 = 2.0e3 #rings/width (in pc)
r_b = 3.5e3 #disk/scale radius (in pc)
sig_s = 8.44e2 #disk/center surface mass density (solar masses per pc^2)
l_c = 1.7e3 #disk/characteristic lenght (in pc)
#adjusting the plot
fig = plt.figure(figsize=(10.0,10.0)) #size of the plot
ax = plt.axes()
ax.grid(True)
ax.yaxis.grid(True,which='minor',linestyle='--')
ax.set_yscale('log')
x = np.linspace(0, 16000, 1000) #x from/to and line smoothness
#functions
plt.plot(x, sig_b*np.exp(-np.power(x/r_0, (1/4))), linestyle='solid', label='Bulge')
plt.plot(x, (sig_g1*np.exp(-np.power(((x-r_g1)/w_g1), 2))) + (sig_g2*np.exp(-np.power(((x-r_g2)/w_g2), 2))), linestyle='dashdot', label='Gas')
plt.plot(x, (sig_s*l_c)/(np.sqrt(np.power((x-r_b), 2) + np.power(l_c, 2))), ':k', label='Stellar Disk');
plt.plot(x, sig_b*np.exp(-np.power(x/r_0, (1/4)))+(sig_g1*np.exp(-np.power(((x-r_g1)/w_g1), 2))) + (sig_g2*np.exp(-np.power(((x-r_g2)/w_g2), 2)))+(sig_s*l_c)/(np.sqrt(np.power((x-r_b), 2) + np.power(l_c, 2))), '--', label='Bulge+Gas+Disk');
plt.axis('tight'); #xmin, xmax, ymin, ymax
#or use: plt.xlim(10, 0) plt.ylim(1.2, -1.2);
#or to tighten the bounds: plt.axis('tight')
#or use: plt.axis([0, 15000, 0.5e6, 2.5e6])
#labeling plots
fig.suptitle('Mass Density of Milky Way', fontsize=16)
ax.set(title='"Unified Rotation Curve of the Milky Way Galaxy" numbers') #labeling axes
ax.set(xlabel='Disc Coordinate [pc]', ylabel='Mass Density [Solar Mass/pc^2]');
plt.legend(fontsize=14); #adding legend
# +
import numpy as np
import matplotlib as mpl
from matplotlib import pyplot as plt
plt.style.use('seaborn-whitegrid')
#parameters for graphs
r_0 = 500 #bulge/half-mass scale radius (in pc)
sig_b = 3.2e3 #bulge/surface mass density at half-mass scale radius (solar masses per pc^2)
sig_g1 = 0.17*8.44e2 #rings/peak density (solar masses per pc^2)
sig_g2 = 0.34*8.44e2 #rings/peak density (solar masses per pc^2)
r_g1 = 3e3 #rings/radii of wave nodes (in pc)
r_g2 = 9.5e3 #rings/radii of wave nodes (in pc)
w_g1 = 1.0e3 #rings/width (in pc)
w_g2 = 2.0e3 #rings/width (in pc)
r_b = 3.5e3 #disk/scale radius (in pc)
sig_s = 8.44e2 #disk/center surface mass density (solar masses per pc^2)
l_c = 1.7e3 #disk/characteristic lenght (in pc)
#adjusting the plot
fig = plt.figure(figsize=(10.0,10.0)) #size of the plot
ax = plt.axes()
ax.grid(True)
ax.yaxis.grid(True,which='minor',linestyle='--')
ax.set_yscale('log')
x = np.linspace(0, 16000, 1000) #x from/to and line smoothness
#functions
plt.plot(x, sig_b*np.exp(-np.power(x/r_0, (1/4)))+(sig_g1*np.exp(-np.power(((x-r_g1)/w_g1), 2))) + (sig_g2*np.exp(-np.power(((x-r_g2)/w_g2), 2)))+(sig_s*l_c)/(np.sqrt(np.power((x-r_b), 2) + np.power(l_c, 2))), '--', label='Bulge+Gas+Disk');
plt.axis('tight'); #xmin, xmax, ymin, ymax
#or use: plt.xlim(10, 0) plt.ylim(1.2, -1.2);
#or to tighten the bounds: plt.axis('tight')
#or use: plt.axis([0, 15000, 0.5e6, 2.5e6])
#labeling plots
fig.suptitle('Mass Density of Milky Way', fontsize=16)
ax.set(title='"Unified Rotation Curve of the Milky Way Galaxy" numbers') #labeling axes
ax.set(xlabel='Disc Coordinate [pc]', ylabel='Mass Density [Solar Mass/pc^2]');
plt.legend(fontsize=14); #adding legend
# +
import numpy as np
import matplotlib as mpl
from matplotlib import pyplot as plt
plt.style.use('seaborn-whitegrid')
#parameters for graphs
#Bulge
sig_be = 3.2e3 #surface mass density at half-mass scale radius (in solar masses per pc^2)
K = 7.6675
r_b = 0.5e3 #half-mass scale radius (in pc)
#Disk
sig_dc = 8.44e2 #central value (in solar masses per pc^2)
r_d = 3.5e3 #scale radius (in pc)
#Dark Halo
rho_0 = 18.3e-3 #scale density (in solar masses per pc^2)
h = 10.7e3 #scale radius (in pc)
#adjusting the plot
fig = plt.figure(figsize=(9.0,8.0)) #size of the plot
ax = plt.axes()
ax.grid(True)
ax.yaxis.grid(True,which='minor',linestyle='--')
ax.set_yscale('log')
x = np.linspace(0, 25000, 1000) #x from/to and line smoothness
#functions
plt.plot(x, sig_be*np.exp(-K*((np.power(x/r_b, (1/4))-1))), linestyle='solid', label='Bulge')
plt.plot(x, sig_dc*np.exp(-x/r_d), linestyle='dashdot', label='Disk')
plt.plot(x, (sig_be*np.exp(-K*((np.power(x/r_b, (1/4))-1)))) + (sig_dc*np.exp(-x/r_d)), '--', label='Bulge+Disk');
#plt.plot(x, rho_0/((x/h)*np.exp(1+x/h, 2)), ':k', label='Dark Halo') #dotted black
plt.axis('tight'); #xmin, xmax, ymin, ymax
#or use: plt.xlim(10, 0) plt.ylim(1.2, -1.2);
#or to tighten the bounds: plt.axis('tight')
#or use: plt.axis([0, 15000, 0.5e6, 2.5e6])
#labeling plots
fig.suptitle('Mass Density of Milky Way', fontsize=16)
ax.set(title='"Mass Distribution and Rotation Curve in the Galaxy" equation+numbers') #labeling axes
ax.set(xlabel='Disc Coordinate [pc]', ylabel='Mass Density [Solar Mass/pc^2]');
plt.legend(fontsize=14); #adding legend
# +
import numpy as np
import matplotlib as mpl
from matplotlib import pyplot as plt
plt.style.use('seaborn-whitegrid')
#parameters for graphs
r_0 = 500 #bulge/characteristic radius of bulge (in pc)
sig_b = 3.2e3 #bulge/central mass density (solar masses per pc^2)
sig_g1 = 4.4e2 #gas/peak surface density (solar masses per pc^2)
sig_g2 = 4.4e2 #gas/peak surface density (solar masses per pc^2)
r_g1 = 5.0e3 #gas/location 1 (in pc)
r_g2 = 12.0e3 #gas/location 2 (in pc)
w_g1 = 4.0e3 #gas/gaussian width (in pc)
w_g2 = 4.0e3 #gas/gaussian width (in pc)
r_b = 2.0e3 #disk/bulge radius (in pc)
sig_s = 8.44e2 #disk/max surface density (solar masses per pc^2)
l_c = 2.5e3 #disk/characteristic lenght (in pc)
#adjusting the plot
fig = plt.figure(figsize=(9.0,8.0)) #size of the plot
ax = plt.axes()
ax.grid(True)
ax.yaxis.grid(True,which='minor',linestyle='--')
ax.set_yscale('log')
x = np.linspace(0, 16000, 1000) #x from/to and line smoothness
#functions
plt.plot(x, sig_b*np.exp(-np.power(x/r_0, (1/4))), linestyle='solid', label='Bulge')
plt.plot(x, (sig_g1*np.exp(-np.power(((x-r_g1)/w_g1), 2))) + (sig_g2*np.exp(-np.power(((x-r_g2)/w_g2), 2))), linestyle='dashdot', label='Gas')
plt.plot(x, (sig_s*l_c)/(np.sqrt(np.power((x-r_b), 2) + np.power(l_c, 2))), ':k', label='Stellar Disk'); #dotted black
plt.plot(x, sig_b*np.exp(-np.power(x/r_0, (1/4)))+(sig_g1*np.exp(-np.power(((x-r_g1)/w_g1), 2))) + (sig_g2*np.exp(-np.power(((x-r_g2)/w_g2), 2)))+(sig_s*l_c)/(np.sqrt(np.power((x-r_b), 2) + np.power(l_c, 2))), '--', label='Bulge+Gas+Disk');
plt.axis('tight'); #xmin, xmax, ymin, ymax
#or use: plt.xlim(10, 0) plt.ylim(1.2, -1.2);
#or to tighten the bounds: plt.axis('tight')
#or use: plt.axis([0, 15000, 0.5e6, 2.5e6])
#labeling plots
fig.suptitle('Mass Density of Milky Way', fontsize=16)
ax.set(title='"Modelling Mass Distribution of the Milky Way using GAIAs Billion-Star Map" equation + "Mass Distribution and Rotation Curve in the Galaxy" numbers') #labeling axes
ax.set(xlabel='Disc Coordinate [pc]', ylabel='Mass Density [Solar Mass/pc^2]');
plt.legend(fontsize=14); #adding legend
# -
y = lambda x: x**2
print(y(5))
a = 3
z = lambda x,y: a*x*y
print(z(4,5))
# +
from mpl_toolkits import mplot3d
def f(x, y):
return np.sin(np.sqrt(x ** 2 + y ** 2))
x = np.linspace(-6, 6, 30)
y = np.linspace(-6, 6, 30)
X, Y = np.meshgrid(x, y)
Z = f(X, Y)
fig = plt.figure()
ax = plt.axes(projection='3d')
ax.contour3D(X, Y, Z, 50, cmap='binary')
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.set_zlabel('z');
print(f(1,1))
# -
| binder/testing/Timing_and_Old_Tests/JB_Jupyter_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.9 64-bit (''stoch_env'': venv)'
# name: python3
# ---
#from pychastic.sde_solver import SDESolver
#from pychastic.sde_solver import VectorSDESolver
import pychastic
import matplotlib.pyplot as plt
import numpy as np
import jax.numpy as jnp
from tqdm import tqdm
# # Testing Vector Euler
# +
problem = pychastic.sde_problem.VectorSDEProblem(
lambda x: jnp.array([1/(2*x[0]),0]), # [1/2r,0]
lambda x: jnp.array([
[jnp.cos(x[1]),jnp.sin(x[1])], # cos \phi, sin \phi
[-jnp.sin(x[1])/x[0],jnp.cos(x[1])/x[0]] # -sin \phi / r, cos \phi / r
]),
dimension = 2,
noiseterms= 2,
x0 = jnp.array([1.0,0.0]), # r=1.0, \phi=0.0
tmax=1.0
)
solvers = [
pychastic.sde_solver.VectorSDESolver(scheme='euler'),
#pychastic.sde_solver.VectorSDESolver(scheme='milstein'),
]
dts = [2**-5, 2**-6, 2**-7, 2**-8]
n_rows = 10
n_wieners_per_cell = 10
results = np.zeros((len(solvers), len(dts), n_rows, n_wieners_per_cell),dtype=dict) # solver x dt x wiener
for n_solver, solver in enumerate(solvers):
for n_dt, dt in tqdm(enumerate(dts)):
solver.dt = dt
#for n_row in tqdm(range(n_rows)):
for n_row in range(n_rows):
#solutions = solver.solve_many(problem,n_trajectories=n_wieners_per_cell)
solutions = np.array([solver.solve(problem,seed = np.random.randint(2**10)) for it in range(n_wieners_per_cell)])
results[n_solver, n_dt, n_row] = solutions
# -
s = results.shape
results_flat = results.flatten()
def to_cartesian(x):
return x[0]*jnp.array([jnp.cos(x[1]),jnp.sin(x[1])])
def f(sol):
x = to_cartesian(sol['last_value'])
x_exact = sol['last_wiener'] + jnp.array([1,0])
e = jnp.sqrt(jnp.sum((x - x_exact)**2)) # L2 norm error
return e
errors = np.array([f(x) for x in results_flat]).reshape(s)
errors.shape
# +
table_euler = errors[0].mean(axis=-1).T
mu_euler = table_euler.mean(axis=0)
sd_euler = table_euler.std(axis=0)/np.sqrt(n_wieners_per_cell)
#table_milstein = errors[1].mean(axis=-1).T
#mu_milstein = table_milstein.mean(axis=0)
#sd_milstein = table_milstein.std(axis=0)/np.sqrt(n_wieners_per_cell)
lsp = jnp.linspace(0.0,0.035,100)
mu_pred_euler = [1.6*x**(1/2) for x in lsp]
#mu_pred_milstein = [10*x for x in lsp]
#mu_pred_milstein_wolfram = [2*x**(2/3) for x in lsp]
plt.errorbar(dts, mu_euler, 2*sd_euler, capsize=5, marker='o', label='Euler-Maruyama')
#plt.errorbar(dts, mu_milstein, 2*sd_milstein, capsize=5, marker='o', label='Milstein')
plt.xlabel('Step size')
plt.ylabel('$L_2$ error')
plt.ylim((0,.5))
plt.plot(lsp, mu_pred_euler)
#plt.plot(lsp, mu_pred_milstein)
#plt.plot(lsp, mu_pred_milstein_wolfram)
plt.legend()
| benchmarks/benchmark_vectors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example use case of the Generative Conditional Independence Test
#
# This notebook provides a simple use case of the Generative Conditional Independence Test for testing independence of two variables given we know about other variables that may be related to our quantities of interest. Please find more details the paper *"Conditional Independence Testing using Generative Adversarial Networks"* by *<NAME> and <NAME>*.
# ## What is the Generative Conditional Independence Test?
# Conditional independence tests are concerned with the question of whether two variables $X$ and $Y$ behave independently of each other, after accounting for the effect of confounders $Z$. Such questions can be written as a hypothesis testing problem: $\mathcal H_0: X\indep Y|Z $ versus the general alternative of no independence. A conditional independence test, given a significance level, determines whether to reject or not the null hypothesis $\mathcal H_0$.
# A number of studies have shown such tests to fail when a large number of variables $Z$ confound the relationship between $X$ and $Y$. This work describes a test that is empirically more robust and whose performance guarantees do not depend on the number of variables involved.
# Our test is based on a modification of Generative Adversarial Networks (GANs) that simulates from a distribution under the assumption of conditional independence, while maintaining good power in high dimensional data. In our procedure, after training, the first step involves simulating from our network to generate data sets consistent with $\mathcal H_0$. We then define a test statistic to capture the $X-Y$ dependency in each sample and compute an empirical distribution which approximates the behaviour of the statistic under $\mathcal H_0$ and can be directly compared to the statistic observed on the real data to make a decision.
# Let us first generate some data from the GCIT.data_utils module in GCIT as follows:
# +
from utils import *
x, y, z = generate_samples_random(size=500, sType='CI', dx=1, dy=1, dz=100,fixed_function='nonlinear', dist_z='gaussian')
# -
# Here we sample gaussian random variables transformed through non-linear functions, such that $Y$ depends on both $X$ and $Z$.
# ### Now, can we say whether there is an independent relationship between $X$ and $Y$ that is not due to $Z$?
# To answer this question, the GCIT takes into account the associations of $Z$ and $Y$ separately to mimic a setting where the null hypothesis holds, i.e. $X$ is irrelevant for inferring $Y$, and then compares this synthetic setting to the actual observations. To arrive at a *p-value* we simply call GCIT with arguments the arrays of data variables as follows:
# +
from GCIT import *
alpha = 0.05
pval = GCIT(x, y, z, verbose=False)
# This is a two-sided test. For a level 0.05 test, set alpha = 0.05/2
if pval < alpha/2:
print('p-value is',pval,'- There is enough evidence in the data to reject the null hypoyhesis.')
else:
print('p-value is',pval,'- There is not enough evidence in the data to reject the null hypothesis.')
| alg/gcit/tutorial_gcit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Some ideas for future development
#
# 1. Metrics ideas:
# - Correlation with returns to find similar stock profiles (and finish MA based correlation).
# - Correlation of day to day change.
# - Event analysis for major moves ( >5% per day ).
# - Event analysis for results publication.
# - Event analysis of news (good thing is no negative response to negative news) - split the news into categories based on PAP?
# - Volatility of some sort.
# - Wave/movement size analysis between local max and min (minor / major).
# 2. Fundamental analysis:
# - Add basic financial data for stocks that are interesting and store them in some kind of database.
# - Calculate automatically interesting metrics from cashflow, balance sheet and profit report.
# 3. Machine learning:
# - Use clustering methods with metrics from 1. and 2. if available to find similar stocks / create GPW map.
# - Use ensamble methods based on metrics to try to create some prediction tool?
# - Use Reinforced Learning to create trading algos (especially interesting for Forex, Oil, US500). (Keras + CNTK?)
# 4. Hot get-rich idea 1:
# - Do daily / weekly percentage change charts + daily / weekly range charts for stocks that went nuts and try to find patters
# - Count days up vs days down ratio
# - Count average change ratio (above or below 0%)
# - Average daily range based on my calc (high-low)/close
# - Threshold for calm vs volatile periods
# - Average time distance between high daily changes for given threshold (positive, negative and both)
# - Identify 10 possible stocks that are starting to rise -> invest 1k in each if odds are good based on your analysis
# - Do basic metrics, like what is the average increase rate, when it should go up, when it will not go up etc.
| examples/notebooks/200_AnalysisIdeas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:stable]
# language: python
# name: conda-env-stable-py
# ---
# + [markdown] toc="true"
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"></ul></div>
# -
# The following command line launches the parallel ipython cores:
# !ipcluster start -n 4
from nbtools import execute_in_parallel
from planet4 import io
db = io.DBManager()
data = db.get_image_id_markings('bo3')[:10]
def calc_stuff(fpath):
d = {'path': fpath}
d['length'] = len(pd.read_csv(fpath))
return d
results = execute_in_parallel(calc_stuff, data)
df = pd.DataFrame(results.result)
from planet4 import markings, io, plotting
import seaborn as sns
sns.set_context('paper')
from planet4 import io
db= io.DBManager()
print(db.dbname)
data = db.get_image_id_markings('bo3')
print('Shape:',data.shape)
from configparser import ConfigParser
config = ConfigParser()
config.read("/Users/klay6683/.pyciss.ini")
config['paths']['my_mac']
df = db.get_image_id_markings('6n3')
df.classification_id.nunique()
p4id = markings.ImageID('6n3')
# %matplotlib inline
fig, ax = plt.subplots(ncols=2, figsize=(16,5))
p4id.show_subframe(ax[0])
p4id.plot_fans(lw=1, ax=ax[1])
fig.tight_layout()
ax[0].set_title(f"{p4id.imgid}, original input")
ax[1].set_title(f"{p4id.imgid}, Planet Four fan markings")
plt.savefig("/Users/klay6683/Dropbox/src/p4_paper1/figures/fan_markings.png",
bbox_inches='tight',
dpi=200)
print(id_.data.image_url.iloc[0])
p4id = markings.ImageID('1aa')
plotting.plot_image_id_pipeline('1aa', datapath='catalog_1.0b2', save=True, savetitle='pipeline',
saveroot='/Users/klay6683/Dropbox/src/p4_paper1/figures/')
| notebooks/io development.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # 7장. 텍스트 데이터 다루기
# *아래 링크를 통해 이 노트북을 주피터 노트북 뷰어(nbviewer.org)로 보거나 구글 코랩(colab.research.google.com)에서 실행할 수 있습니다.*
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://nbviewer.org/github/rickiepark/intro_ml_with_python_2nd_revised/blob/main/07-working-with-text-data.ipynb"><img src="https://jupyter.org/assets/share.png" width="60" />주피터 노트북 뷰어로 보기</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/rickiepark/intro_ml_with_python_2nd_revised/blob/main/07-working-with-text-data.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />구글 코랩(Colab)에서 실행하기</a>
# </td>
# </table>
# <b><font size=2>이 노트북은 맷플롯립 그래프에 한글을 쓰기 위해 나눔 폰트를 사용합니다. 컴퓨터에 나눔 폰트가 없다면 설치해 주세요.<br><br><font color='red'>주의: 코랩에서 실행하는 경우 아래 셀을 실행하고 ⌘+M . 또는 Ctrl+M . 을 눌러 런타임을 재시작한 다음 처음부터 다시 실행해 주세요.</font></b>
# 노트북이 코랩에서 실행 중인지 체크합니다.
import os
import sys
if 'google.colab' in sys.modules and not os.path.isdir('mglearn'):
# 사이킷런 최신 버전을 설치합니다.
# !pip install -q --upgrade scikit-learn
# mglearn을 다운받고 압축을 풉니다.
# !wget -q -O mglearn.tar.gz https://bit.ly/mglearn-tar-gz
# !tar -xzf mglearn.tar.gz
# 나눔 폰트를 설치합니다.
# !sudo apt-get -qq -y install fonts-nanum
import matplotlib.font_manager as fm
fm._rebuild()
# +
import sklearn
from preamble import *
import matplotlib
# 나눔 폰트를 사용합니다.
matplotlib.rc('font', family='NanumBarunGothic')
matplotlib.rcParams['axes.unicode_minus'] = False
# -
# ## 7.1 문자열 데이터 타입
# ## 7.2 예제 어플리케이션: 영화 리뷰 감성 분석
# +
import os.path
if not os.path.isfile('data/aclImdb_v1.tar.gz'):
# !wget -q http://ai.stanford.edu/~amaas/data/sentiment/aclImdb_v1.tar.gz -P data
# !tar -xzf data/aclImdb_v1.tar.gz -C data
# -
# !은 셸(shell) 명령을 실행해주는 IPython의 매직 명령어입니다.
# tree 명령이 없다면 find ./data -type d 명령을 사용해 하위 폴더의 목록을
# 볼 수 있습니다. 윈도에서는 !tree data/aclImdb 와 같이 사용하세요.
# # !tree -dL 2 data/aclImdb
# !find ./data -type d
# !rm -r data/aclImdb/train/unsup
# +
from sklearn.datasets import load_files
reviews_train = load_files("data/aclImdb/train/")
# 텍스트와 레이블을 포함하고 있는 Bunch 오브젝트를 반환합니다.
text_train, y_train = reviews_train.data, reviews_train.target
print("text_train의 타입:", type(text_train))
print("text_train의 길이:", len(text_train))
print("text_train[6]:\n", text_train[6])
# -
text_train = [doc.replace(b"<br />", b" ") for doc in text_train]
print("클래스별 샘플 수 (훈련 데이터):", np.bincount(y_train))
reviews_test = load_files("data/aclImdb/test/")
text_test, y_test = reviews_test.data, reviews_test.target
print("테스트 데이터의 문서 수:", len(text_test))
print("클래스별 샘플 수 (테스트 데이터):", np.bincount(y_test))
text_test = [doc.replace(b"<br />", b" ") for doc in text_test]
# ## 7.3 텍스트 데이터를 BOW로 표현하기
# ### 7.3.1 샘플 데이터에 BOW 적용하기
bards_words =["The fool doth think he is wise,",
"but the wise man knows himself to be a fool"]
from sklearn.feature_extraction.text import CountVectorizer
vect = CountVectorizer()
vect.fit(bards_words)
print("어휘 사전의 크기:", len(vect.vocabulary_))
print("어휘 사전의 내용:\n", vect.vocabulary_)
bag_of_words = vect.transform(bards_words)
print("BOW:", repr(bag_of_words))
print("BOW의 밀집 표현:\n", bag_of_words.toarray())
# ### 7.3.2 영화 리뷰에 대한 BOW
vect = CountVectorizer().fit(text_train)
X_train = vect.transform(text_train)
print("X_train:\n", repr(X_train))
# get_feature_names() 메서드가 1.0에서 deprecated 되었고 1.2 버전에서 삭제될 예정입니다.
# 대신 get_feature_names_out()을 사용합니다.
feature_names = vect.get_feature_names_out()
print("특성 개수:", len(feature_names))
print("처음 20개 특성:\n", feature_names[:20])
print("20010에서 20030까지 특성:\n", feature_names[20010:20030])
print("매 2000번째 특성:\n", feature_names[::2000])
# +
from sklearn.model_selection import cross_val_score
from sklearn.linear_model import LogisticRegression
scores = cross_val_score(LogisticRegression(max_iter=1000), X_train, y_train, n_jobs=-1)
print("교차 검증 평균 점수: {:.2f}".format(np.mean(scores)))
# -
from sklearn.model_selection import GridSearchCV
param_grid = {'C': [0.001, 0.01, 0.1, 1, 10]}
grid = GridSearchCV(LogisticRegression(max_iter=5000), param_grid, n_jobs=-1)
grid.fit(X_train, y_train)
print("최상의 교차 검증 점수: {:.2f}".format(grid.best_score_))
print("최적의 매개변수: ", grid.best_params_)
X_test = vect.transform(text_test)
print("테스트 점수: {:.2f}".format(grid.score(X_test, y_test)))
vect = CountVectorizer(min_df=5).fit(text_train)
X_train = vect.transform(text_train)
print("min_df로 제한한 X_train:", repr(X_train))
# +
# get_feature_names() 메서드가 1.0에서 deprecated 되었고 1.2 버전에서 삭제될 예정입니다.
# 대신 get_feature_names_out()을 사용합니다.
feature_names = vect.get_feature_names_out()
print("처음 50개 특성:\n", feature_names[:50])
print("20,010부터 20,030까지 특성:\n", feature_names[20010:20030])
print("매 700번째 특성:\n", feature_names[::700])
# -
grid = GridSearchCV(LogisticRegression(max_iter=5000), param_grid, n_jobs=-1)
grid.fit(X_train, y_train)
print("최상의 교차 검증 점수: {:.2f}".format(grid.best_score_))
# ## 7.4 불용어
from sklearn.feature_extraction.text import ENGLISH_STOP_WORDS
print("불용어 개수:", len(ENGLISH_STOP_WORDS))
print("매 10번째 불용어:\n", list(ENGLISH_STOP_WORDS)[::10])
# stop_words="english"라고 지정하면 내장된 불용어를 사용합니다.
# 내장된 불용어에 추가할 수도 있고 자신만의 목록을 사용할 수도 있습니다.
vect = CountVectorizer(min_df=5, stop_words="english").fit(text_train)
X_train = vect.transform(text_train)
print("불용어가 제거된 X_train:\n", repr(X_train))
grid = GridSearchCV(LogisticRegression(max_iter=5000), param_grid, n_jobs=-1)
grid.fit(X_train, y_train)
print("최상의 교차 검증 점수: {:.2f}".format(grid.best_score_))
from sklearn.pipeline import make_pipeline
pipe = make_pipeline(CountVectorizer(), LogisticRegression(max_iter=5000))
param_grid = {'countvectorizer__max_df': [100, 1000, 10000, 20000], 'logisticregression__C': [0.001, 0.01, 0.1, 1, 10]}
grid = GridSearchCV(pipe, param_grid, n_jobs=-1)
grid.fit(text_train, y_train)
print("최상의 교차 검증 점수: {:.2f}".format(grid.best_score_))
print(grid.best_params_)
len(grid.best_estimator_.named_steps['countvectorizer'].vocabulary_)
scores = grid.cv_results_['mean_test_score'].reshape(-1, 5)
# 히트맵을 그립니다
heatmap = mglearn.tools.heatmap(
scores, xlabel="C", ylabel="max_df", cmap="viridis", fmt="%.3f",
xticklabels=param_grid['logisticregression__C'],
yticklabels=param_grid['countvectorizer__max_df'])
plt.colorbar(heatmap)
plt.show() # 책에는 없음
# ## 7.5 tf–idf로 데이터 스케일 변경
# \begin{equation*}
# \text{tfidf}(w, d) = \text{tf} \times (\log\big(\frac{N + 1}{N_w + 1}\big) + 1)
# \end{equation*}
# +
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.pipeline import make_pipeline
pipe = make_pipeline(TfidfVectorizer(min_df=5), LogisticRegression(max_iter=5000))
param_grid = {'logisticregression__C': [0.001, 0.01, 0.1, 1, 10]}
grid = GridSearchCV(pipe, param_grid, n_jobs=-1)
grid.fit(text_train, y_train)
print("최상의 교차 검증 점수: {:.2f}".format(grid.best_score_))
# +
vectorizer = grid.best_estimator_.named_steps["tfidfvectorizer"]
# 훈련 데이터셋을 변환합니다
X_train = vectorizer.transform(text_train)
# 특성별로 가장 큰 값을 찾습니다
max_value = X_train.max(axis=0).toarray().ravel()
sorted_by_tfidf = max_value.argsort()
# get_feature_names() 메서드가 1.0에서 deprecated 되었고 1.2 버전에서 삭제될 예정입니다.
# 대신 get_feature_names_out()을 사용합니다.
# 특성 이름을 구합니다
feature_names = np.array(vectorizer.get_feature_names_out())
print("가장 낮은 tfidf를 가진 특성:\n",
feature_names[sorted_by_tfidf[:20]])
print("가장 높은 tfidf를 가진 특성: \n",
feature_names[sorted_by_tfidf[-20:]])
# -
sorted_by_idf = np.argsort(vectorizer.idf_)
print("가장 낮은 idf를 가진 특성:\n",
feature_names[sorted_by_idf[:100]])
# ## 7.6 모델 계수 조사
grid.best_estimator_.named_steps["logisticregression"].coef_
mglearn.tools.visualize_coefficients(
grid.best_estimator_.named_steps["logisticregression"].coef_[0],
feature_names, n_top_features=40)
# ## 7.7 여러 단어로 만든 BOW (n-그램)
print("bards_words:\n", bards_words)
cv = CountVectorizer(ngram_range=(1, 1)).fit(bards_words)
print("어휘 사전 크기:", len(cv.vocabulary_))
# get_feature_names() 메서드가 1.0에서 deprecated 되었고 1.2 버전에서 삭제될 예정입니다.
# 대신 get_feature_names_out()을 사용합니다.
print("어휘 사전:\n", cv.get_feature_names_out())
cv = CountVectorizer(ngram_range=(2, 2)).fit(bards_words)
print("어휘 사전 크기:", len(cv.vocabulary_))
# get_feature_names() 메서드가 1.0에서 deprecated 되었고 1.2 버전에서 삭제될 예정입니다.
# 대신 get_feature_names_out()을 사용합니다.
print("어휘 사전:\n", cv.get_feature_names_out())
print("변환된 데이터 (밀집 배열):\n", cv.transform(bards_words).toarray())
cv = CountVectorizer(ngram_range=(1, 3)).fit(bards_words)
print("어휘 사전 크기:", len(cv.vocabulary_))
# get_feature_names() 메서드가 1.0에서 deprecated 되었고 1.2 버전에서 삭제될 예정입니다.
# 대신 get_feature_names_out()을 사용합니다.
print("어휘 사전:\n", cv.get_feature_names_out())
# +
pipe = make_pipeline(TfidfVectorizer(min_df=5), LogisticRegression(max_iter=5000))
# 매개변수 조합이 많고 트라이그램이 포함되어 있기 때문에
# 그리드 서치 실행에 시간이 오래 걸립니다
param_grid = {'logisticregression__C': [0.001, 0.01, 0.1, 1, 10, 100],
"tfidfvectorizer__ngram_range": [(1, 1), (1, 2), (1, 3)]}
grid = GridSearchCV(pipe, param_grid, n_jobs=-1)
grid.fit(text_train, y_train)
print("최상의 교차 검증 점수: {:.2f}".format(grid.best_score_))
print("최적의 매개변수:\n", grid.best_params_)
# -
# 그리드 서치에서 테스트 점수를 추출합니다
scores = grid.cv_results_['mean_test_score'].reshape(-1, 3).T
# 히트맵을 그립니다
heatmap = mglearn.tools.heatmap(
scores, xlabel="C", ylabel="ngram_range", cmap="viridis", fmt="%.3f",
xticklabels=param_grid['logisticregression__C'],
yticklabels=param_grid['tfidfvectorizer__ngram_range'])
plt.colorbar(heatmap)
plt.show() # 책에는 없음
# 특성 이름과 계수를 추출합니다
vect = grid.best_estimator_.named_steps['tfidfvectorizer']
# get_feature_names() 메서드가 1.0에서 deprecated 되었고 1.2 버전에서 삭제될 예정입니다.
# 대신 get_feature_names_out()을 사용합니다.
feature_names = np.array(vect.get_feature_names_out())
coef = grid.best_estimator_.named_steps['logisticregression'].coef_
mglearn.tools.visualize_coefficients(coef[0], feature_names, n_top_features=40)
plt.ylim(-22, 22)
plt.show() # 책에는 없음
# 트라이그램 특성을 찾습니다
mask = np.array([len(feature.split(" ")) for feature in feature_names]) == 3
# 트라이그램 특성만 그래프로 나타냅니다
mglearn.tools.visualize_coefficients(coef.ravel()[mask],
feature_names[mask], n_top_features=40)
plt.ylim(-22, 22)
plt.show() # 책에는 없음
# ## 7.8 고급 토큰화, 어간 추출, 표제어 추출
# nltk와 spacy를 설치합니다.
# +
# !pip install -q nltk spacy
import spacy
try:
en_nlp = spacy.load('en_core_web_sm')
except:
# !pip install spacy
# !python -m spacy download en
# +
import spacy
import nltk
# spacy의 영어 모델을 로드합니다
en_nlp = spacy.load('en_core_web_sm')
# nltk의 PorterStemmer 객체를 만듭니다
stemmer = nltk.stem.PorterStemmer()
# spacy의 표제어 추출과 nltk의 어간 추출을 비교하는 함수입니다
def compare_normalization(doc):
# spacy로 문서를 토큰화합니다
doc_spacy = en_nlp(doc)
# spacy로 찾은 표제어를 출력합니다
print("표제어:")
print([token.lemma_ for token in doc_spacy])
# PorterStemmer로 찾은 토큰을 출력합니다
print("어간:")
print([stemmer.stem(token.norm_.lower()) for token in doc_spacy])
# -
compare_normalization(u"Our meeting today was worse than yesterday, "
"I'm scared of meeting the clients tomorrow.")
# +
# 요구사항: spacy에서 표제어 추출 기능과 CountVectorizer의 토큰 분할기를 사용합니다.
# spacy의 언어 모델을 로드합니다
en_nlp = spacy.load('en_core_web_sm', disable=['parser', 'ner'])
# spacy 문서 처리 파이프라인을 사용해 자작 토큰 분할기를 만듭니다
# (우리만의 토큰 분할기를 사용합니다)
def custom_tokenizer(document):
doc_spacy = en_nlp(document)
return [token.lemma_ for token in doc_spacy]
# 자작 토큰 분할기를 사용해 CountVectorizer 객체를 만듭니다
lemma_vect = CountVectorizer(tokenizer=custom_tokenizer, min_df=5)
# +
# 표제어 추출이 가능한 CountVectorizer 객체로 text_train을 변환합니다
X_train_lemma = lemma_vect.fit_transform(text_train)
print("X_train_lemma.shape:", X_train_lemma.shape)
# 비교를 위해 표준 CountVectorizer를 사용합니다
vect = CountVectorizer(min_df=5).fit(text_train)
X_train = vect.transform(text_train)
print("X_train.shape:", X_train.shape)
# +
# 훈련 세트의 1%만 사용해서 그리드 서치를 만듭니다
from sklearn.model_selection import StratifiedShuffleSplit
param_grid = {'C': [0.001, 0.01, 0.1, 1, 10]}
cv = StratifiedShuffleSplit(n_splits=5, test_size=0.99,
train_size=0.01, random_state=0)
grid = GridSearchCV(LogisticRegression(max_iter=5000), param_grid, cv=cv, n_jobs=-1)
# 기본 CountVectorizer로 그리드 서치를 수행합니다
grid.fit(X_train, y_train)
print("최상의 교차 검증 점수 "
"(기본 CountVectorizer): {:.3f}".format(grid.best_score_))
# 표제어를 사용해서 그리드 서치를 수행합니다
grid.fit(X_train_lemma, y_train)
print("최상의 교차 검증 점수 "
"(표제어): {:.3f}".format(grid.best_score_))
# -
# ## 7.9 토픽 모델링과 문서 군집화
# ### 7.9.1 LDA
vect = CountVectorizer(max_features=10000, max_df=.15)
X = vect.fit_transform(text_train)
from sklearn.decomposition import LatentDirichletAllocation
lda = LatentDirichletAllocation(n_components=10, learning_method="batch",
max_iter=25, random_state=0, n_jobs=-1)
# 모델 생성과 변환을 한 번에 합니다
# 변환 시간이 좀 걸리므로 시간을 절약하기 위해 동시에 처리합니다
document_topics = lda.fit_transform(X)
print("lda.components_.shape:", lda.components_.shape)
# 토픽마다(components_의 행) 특성을 오름차순으로 정렬합니다
# 내림차순이 되도록 [:, ::-1] 사용해 행의 정렬을 반대로 바꿉니다
sorting = np.argsort(lda.components_, axis=1)[:, ::-1]
# get_feature_names() 메서드가 1.0에서 deprecated 되었고 1.2 버전에서 삭제될 예정입니다.
# 대신 get_feature_names_out()을 사용합니다.
# CountVectorizer 객체에서 특성 이름을 구합니다.
feature_names = np.array(vect.get_feature_names_out())
# 10개의 토픽을 출력합니다
mglearn.tools.print_topics(topics=range(10), feature_names=feature_names,
sorting=sorting, topics_per_chunk=5, n_words=10)
lda100 = LatentDirichletAllocation(n_components=100, learning_method="batch",
max_iter=25, random_state=0, n_jobs=-1)
document_topics100 = lda100.fit_transform(X)
topics = np.array([7, 16, 24, 25, 28, 36, 37, 41, 45, 51, 53, 54, 63, 89, 97])
sorting = np.argsort(lda100.components_, axis=1)[:, ::-1]
# get_feature_names() 메서드가 1.0에서 deprecated 되었고 1.2 버전에서 삭제될 예정입니다.
# 대신 get_feature_names_out()을 사용합니다.
feature_names = np.array(vect.get_feature_names_out())
mglearn.tools.print_topics(topics=topics, feature_names=feature_names,
sorting=sorting, topics_per_chunk=5, n_words=20)
# 음악적인 토픽 45를 가중치로 정렬합니다
music = np.argsort(document_topics100[:, 45])[::-1]
# 이 토픽이 가장 비중이 큰 문서 다섯개를 출력합니다
for i in music[:10]:
# 첫 두 문장을 출력합니다
print(b".".join(text_train[i].split(b".")[:2]) + b".\n")
fig, ax = plt.subplots(1, 2, figsize=(10, 10))
topic_names = ["{:>2} ".format(i) + " ".join(words)
for i, words in enumerate(feature_names[sorting[:, :2]])]
# 두 개의 열이 있는 막대 그래프
for col in [0, 1]:
start = col * 50
end = (col + 1) * 50
ax[col].barh(np.arange(50), np.sum(document_topics100, axis=0)[start:end])
ax[col].set_yticks(np.arange(50))
ax[col].set_yticklabels(topic_names[start:end], ha="left", va="top")
ax[col].invert_yaxis()
ax[col].set_xlim(0, 2000)
yax = ax[col].get_yaxis()
yax.set_tick_params(pad=130)
plt.tight_layout()
# ## 7.10 요약 및 정리
| 07-working-with-text-data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="../assets/images/science_logo_400x400.jpg" style="width: 150px; float: right;">
#
# # 02 - Programming for `UniProt` (browser/notebook)
# ## Table of Contents
#
# 1. [Introduction](#introduction)
# 2. [Python imports](#imports)
# 3. [Running a remote `UniProt` query](#uniprot)
# 1. [Connecting to `UniProt`](#connect)
# 2. [Constructing a query](#query)
# 3. [Perform the query](#search)
# 4. [EXAMPLE: Putting it together](#example01)
# 4. [Advanced queries](#advanced)
# 1. [`key:value` queries](#keyvalue)
# 2. [Exercise 01](#ex01)
# 3. [Combining queries](#combine)
# 4. [Exercise 02](#ex02)
# 4. [Processing query results](#processing)
# 1. [Tabular](#tabular)
# 2. [Excel](#excel)
# 3. [FASTA sequence](#fasta)
# 4. [`pandas` dataframe](#pandas)
# <a id="introduction"></a>
# ## Introduction
#
# The `UniProt` browser interface is very powerful, but you will have noticed from the previous exercises that even the most complex queries can be converted into a single string that describes the search being made of the `UniProt` databases. Using the browser interface, this string is generated for you, and placed into the search field at the top of the `UniProt` webpage every time you run a search.
#
# <p></p>
# <div class="alert-danger">
# <b>It can be tedious and time-consuming to point-and-click your way through a large number of browser-based searches, but by using the <code>UniProt</code> <i>webservice</i>, the search strings you've already seen, and a Python module called <code>bioservices</code>, we can compose and run as many searches as we like using a small amount of code, and pull the results of those searches down to our local machines.</b>
# </div>
#
# This notebook presents examples of methods for using `UniProt` programmatically, via a *webservice*, and you will be controlling the searches using Python code in this notebook.
#
# There are a number of advantages to this approach:
#
# <p></p>
# <div class="alert-success"><b>
# <ul>
# <li> It is easy to set up repeatable searches for many sequences, or collections of sequences
# <li> It is easy to read in the search results and conduct downstream analyses that add value to your search
# </ul>
# </b></div>
#
# Where it is not practical to submit a large number of simultaneous queries via a web form (because it is tiresome to point-and-click over and over again), this can be handled programmatically instead. You have the opportunity to change custom options to help refine your query, compared to the website interface. If you need to repeat a query, it can be trivial to apply the same settings every time, if you use a programmatic approach.
# <a id="imports"></a>
# ## Python imports
#
# To use the Python programming language to query `UniProt`, we have to *import* helpful *packages* (collections of Python code that perform specialised tasks.
# +
# io is a standard library package that lets us manipulate data
import io
# Import Seaborn for graphics and plotting
import seaborn as sns
# Import bioservices module, to run remote UniProt queries
from bioservices import UniProt
# Import Pandas, so we can use dataframes
import pandas as pd
# -
# <a id="uniprot"></a>
# ## Running a remote `UniProt` query
#
# <p></p>
# <div class="alert-success">
# <b>There are three key steps to running a remote <code>UniProt</code> query with <code>bioservices</code>:</b>
# </div>
#
# 1. Make a link to the `UniProt` webservice
# 2. Construct a query string
# 3. Send the query to `UniProt`, and catch the result in a *variable*
#
# Once the search result is caught and contained in a *variable*, that *variable* can be processed in any way you like, written to a file, or ignored.
# <a id="connect"></a>
# ### Connecting to `UniProt`
#
# To open a connection to `UniProt`, you make an *instance* of the `UniProt()` *class* from `bioservices`. This can be made to be persistent so that, once a single connection to the database is created, you can interact with it over and over again to make multiple queries.
#
# <p></p>
# <div class="alert-success">
# <b>To make a persistent instance, you can assign <code>UniProt()</code> to a variable:</b>
# </div>
#
# ```python
# service = UniProt() # it is good practice to have a meaningful variable name
# ```
# <a id="query"></a>
# ### Constructing a query
#
# <p></p>
# <div class="alert-success">
# <b><code>UniProt</code> allows for the construction of complex searches by combining <i>fields</i>. A full discussion is beyond the scope of this lesson, but you will have seen in <a href="01-uniprot_browser.ipynb">the preceding notebook</a> that the searches you constructed by pointing and clicking on the <code>UniProt</code> website were converted into text in the search field at the top. </b>
# </div>
#
# To describe the format briefly: there are a set of defined *keywords* (or *keys*) that indicate the specific type of data you want to search in (such as `host`, `annotation`, or sequence `length`), and these are combined with a particular *value* you want to search for (such as `mouse`, or `40674`) in a `key:value` pair, separated by a colon, such as `host:mouse` or `ec:172.16.58.3`.
#
# * `UniProt` query fields: [http://www.uniprot.org/help/query-fields](http://www.uniprot.org/help/query-fields)
#
# If you provide a string, instead of a `key:value` pair, `UniProt` will search in all *fields* for your search term.
#
# Programmatically, we construct the query as a *string*, e.g.
#
# ```python
# query = "Q9AJE3" # this query means we want to look in all fields for Q9AJE3
# ```
# <a id="search"></a>
# ### Perform the query
#
# To send the query to `UniProt`, you will use the `.search()` *method* of your active *instance* of the `UniProt()` *class*.
#
# <p></p>
# <div class="alert-success">
# <b>If you have assigned your instance to the variable <code>service</code> (as above), then you can run the <code>query</code> string as a remote search with the line:</b>
# </div>
#
# ```python
# result = service.search(query) # Run a query and catch the output in result
# ```
#
# In the line above, the output of the search (i.e. your result) is stored in a new variable (created when the search is complete) called `result`. It is good practice to make variable names short and descriptive - this makes your code easier to read.
# <a id="example01"></a>
# ### EXAMPLE: Putting it together
#
# The code in the cell below uses the example code above to create an instance of the `UniProt()` class, and uses it to submit a pre-stored query to the `UniProt` service, then catch the result in a variable called `result`. The `print()` statement then shows us what the result returned by the service looks like.
# +
# Make a link to the UniProt webservice (UniProt())
service = UniProt()
# Build a query string ("Q9AJE3")
query = "Q9AJE3"
# Send the query to UniProt, and catch the search result in a variable (service.search())
result = service.search(query)
# Inspect the result
print(result)
# -
# The `UniProt()` instance defined in the cell above is *persistent*, so you can reuse it to make another query, as in the cell below:
# +
# Make a new query string "Q01844", and run a remote search at UniProt
new_query = "Q01844"
new_result = service.search(new_query)
# Inspect the result
print(new_result)
# -
# <a id="advanced"></a>
# ## Advanced queries
#
# <p></p>
# <div class="alert-success">
# <b>The examples above built queries that were simple strings. They did not exploit the <code>key:value</code> search structure, or combine search terms. In this section, you will explore some queries that use the <code>UniProt</code> query fields, and combine them into powerful, filtering searches.</b>
# </div>
# <a id="keyvalue"></a>
# ### `key:value` queries
#
# As noted above (and at [http://www.uniprot.org/help/query-fields](http://www.uniprot.org/help/query-fields)) particular values of specific data can be requested by using `key:value` pairs to restrict searches to named *fields* in the `UniProt` database.
#
# As a first example, you will note that the result returned for the query `"Q01844"` has multiple entries. Only one of these is the sequence with `accession` value equal to `"Q01844"`, but the other entries make reference to this sequence somewhere in their database record. If we want to restrict our result only to the particular entry `"Q01844"`, we can specify the field we want to search as `accession`, and build the following query:
#
# ```python
# query = "accession:Q01844" # specify a search on the accession field
# ```
#
# Note that we can use the same variable name `query` as earlier (this overwrites the previous value in `query`). The code below runs the search and shows the output:
# +
# Make a new query string ("accession:Q01844"), and run a remote search at UniProt
query = "accession:Q01844"
result = service.search(query)
# Inspect the result
print(result)
# -
# <div class="alert-success">
# <b>By using this and other <code>key:value</code> constructions, we can refine our searches to give us only the entries we're interested in</b>
# </div>
# <img src="../assets/images/exercise.png" style="width: 100px; float: left;">
#
# <a id="ex01"></a>
# ### Exercise 01 (10min)
#
# Using `key:value` searches, can you find and download sets of entries for proteins that satisfy the following requirements? (**HINT:** this <a href="http://www.uniprot.org/help/query-fields">link</a> to the `UniProt` query fields may be helpful, here):
#
# <p></p>
# <div class="alert-danger">
# <ul>
# <li> Have publications authored by someone with the surname Broadhurst
# <li> Have protein length between 9000aa and 9010aa
# <li> Derive from the taipan snake
# <li> Have been found in the wing
# </ul>
# </div>
# SOLUTION - EXERCISE 01
query = "citation:(author:broadhurst)"
result = service.search(query)
print(result)
# SOLUTION - EXERCISE 01
query = "length:[9000 TO 9010]"
result = service.search(query)
print(result)
# SOLUTION - EXERCISE 01
query = "organism:taipan"
result = service.search(query)
print(result)
# SOLUTION - EXERCISE 01
query = 'annotation:(type:"tissue specificity" wing)'
result = service.search(query)
print(result)
# <a id="combine"></a>
# ### Combining queries
#
# <p></p>
# <div class="alert-success">
# <b>Combining terms in a <code>UniProt</code> query can be as straightforward as putting them in the same string, separated by a space.</b>
# </div>
#
# For example:
#
# ```python
# query = "organism:rabbit tissue:eye"
# ```
#
# will search for all entries deriving from rabbits that are found in the eye
# Combine queries for rabbit (organism) and eye (tissue), and search
query = "organism:rabbit tissue:eye"
result = service.search(query)
print(result)
# <img src="../assets/images/exercise.png" style="width: 100px; float: left;">
#
# <a id="ex02"></a>
# ### Exercise 02 (10min)
#
# Using `key:value` searches, can you find and download sets of entries for proteins that satisfy the following requirements? (**HINT:** this <a href="http://www.uniprot.org/help/query-fields">link</a> to the `UniProt` query fields may be helpful, here):
#
# <p></p>
# <div class="alert-danger">
# <ul>
# <li> Found in sheep spleen
# <li> Have "rxlr" in their name, have a publication with author name Pritchard, and are between 70aa and 80aa in length
# <li> Derive from a quokka and have had their annotations manually reviewed
# <li> Are found in cell membranes of horse heart tissue, and have had their annotations manually reviewed
# </ul>
# </div>
# SOLUTION - EXERCISE 02
query = "name:rxlr author:pritchard length:[70 TO 80]"
result = service.search(query)
print(result)
# SOLUTION - EXERCISE 02
query = "organism:quokka AND reviewed:yes"
result = service.search(query)
print(result)
# SOLUTION - EXERCISE 02
query = 'organism:horse annotation:(type:"tissue specificity" heart) locations:(location:membrane) reviewed:yes'
result = service.search(query)
print(result)
# <a id="boolean"></a>
# ### Combining terms with Boolean logic
#
# Boolean logic allows you to combine search terms with each other in arbitrary ways using three *operators*, specifying whether:
#
# * both terms are required (`AND`) **NOTE:** this is implicitly what you have been doing in the examples above
# * either term is allowed (`OR`)
# * a term is disallowed (`NOT`)
#
# Searches are read from left-to right, but the logic of a search can be controlled by placing the combinations you want to resolve first in parentheses (`()`). Combining these operators can build some extremely powerful searches. For example, to get all proteins from horses and sheep, identified in the ovary, and having length greater than 200aa, you could use the query:
#
# ```
# query = "tissue:ovary AND (organism:sheep OR organism:horse) NOT length:[1 TO 200]"
# ```
# <a id="processing"></a>
# ## Processing query results
#
# So far you have worked with the default output from `bioservices`, although you know from [the previous notebook](01-uniprot_browser.ipynb) that `UniProt` can provide output in a number of useful formats for searches in the browser.
#
# The default output is `tabular`, and gives a good idea of the nature and content of the entries you recover. In this section, you will see some ways to download search results in alternative formats, which can be useful for analysis.
#
# All the output format options are controlled in a similar way, using the `frmt=<format>` argument when you conduct your search - with `<format>` being one of the allowed terms (see the [`bioservices` documentation](https://pythonhosted.org/bioservices/references.html#module-bioservices.uniprot) for a full list).
# <a id="tabular"></a>
# ### Tabular
#
# <p></p>
# <div class="alert-success">
# <b>The default datatype is the most flexible datatype for download: <i>tabular</i>.</b>
# </div>
#
# This can be specified explicitly with the `tab` format:
#
# ```python
# result = service.search(query, frmt="tab")
# ```
# +
# Make a query string ("Q01844"), and run a remote search at UniProt,
# getting the result as tabular format (frmt="tab")
query = "Q01844"
result = service.search(query, frmt="tab")
# Inspect the result
print(result)
# -
# By default, the columns that are returned are: `Entry`, `Entry name`, `Status`, `Protein names`, `Gene names`, `Organism`, and `Length`. But these can be modified by passing the `columns=<list>` argument, where the `<list>` is a comma-separated list of column names. For example:
#
# ```python
# columnlist = "id,entry name,length,organism,mass,domains,domain,pathway"
# result = service.search(query, frmt="tab", columns=columnlist)
# ```
#
# The list of allowed column names can be found by inspecting the content of the variable `service._valid_columns`.
# +
# Make a query string ("Q01844")
query = "Q01844"
# Define a list of columns we want to retrive
columnlist = "id,entry name,length,mass,go(cellular component)"
# Run the remote search (columns=columnlist)
result = service.search(query, columns=columnlist)
# View the result
print(result)
# -
# #### Converting to a dataframe
#
# <p></p>
# <div class="alert-success">
# <b>The <code>pandas</code> module allows us to process tabular data into dataframes, just like in <code>R</code>.</b>
# </div>
#
# To do this, we have to use the `io.StringIO()` class to make it think that our downloaded results are a file
#
# ```python
# df = pd.read_table(io.StringIO(result))
# ```
# +
# Convert the last search result into a dataframe in Pandas
df = pd.read_table(io.StringIO(result))
# View the dataframe
df
# -
# Doing this will produce a `pandas` dataframe that can be manipulated and analysed just like any other dataframe. We can, for instance, view a histogram of sequence lengths from the table above:
# Plot histogram of protein sequence lengths from dataframe
df.hist();
# <a id="excel"></a>
# ### Excel
#
# <p></p>
# <div class="alert-success">
# <b>You can download Excel spreadsheets directly from <code>UniProt</code>, just as with the browser interface.</b>
# </div>
#
# ```python
# result = service.search(query, frmt="xls")
# ```
#
# You can't use the Excel output directly in your code without some file manipulation, and you'll have to save it to a file, as in the example below. Also, the downloaded format is not guaranteed to be current for your version of Excel, and the application may ask to repair it. But, if you want Excel output to share with/display to others, you can get it programmatically.
#
# <p></p>
# <div class="alert-danger">
# <b>NOTE: the downloaded format is actually `.xlsx`, rather than `.xls` which is implied by the format</b>
# </div>
# +
# Make a query string, and run a remote search at UniProt,
# getting the result as an Excel spreadsheer
query = "Q01844"
result = service.search(query, frmt="xls")
# Write the Excel spreadsheet to file
outfile = '../assets/downloads/Q01844.xlsx'
with open(outfile, 'wb') as ofh:
ofh.write(result)
# -
# <a id="fasta"></a>
# ### FASTA sequence
#
# <p></p>
# <div class="alert-success">
# <b>If you're interested only in the FASTA format sequence for an entry, you can use the <code>fasta</code> option with <code>frmt</code> to recover the sequences directly, as in the example below:</b>
# </div>
# +
# Make a query string, and run a remote search at UniProt,
# getting the result as FASTA sequence
query = "go:membrane organism:horse tissue:heart reviewed:yes"
result = service.search(query, frmt="fasta")
# Inspect the result
print(result)
# -
# <a id="pandas"></a>
# ### `pandas` dataframe
#
# <p></p>
# <div class="alert-success">
# <b>In addition to the conversion of tabular output to a <code>pandas</code> dataframe above, you can ask the <code>UniProt()</code> instance to return a <code>pandas</code> dataframe directly, with the <code>.get_df()</code> method.</b>
# </div>
#
# ```python
# result = service.get_df("tissue:venom (organism:viper OR organism:mamba)", limit=None)
# ```
#
# However, this is slow compared to the other methods above and can take a long time for queries with thousands of results
# +
# Get a dataframe for all venom proteins from snakes (or mambas, if "snake" is not in the annotation)
df = service.get_df('tissue:venom (organism:viper OR organism:mamba)', limit=None)
# View the dataframe
df.head()
# -
# This dataframe works like any other dataframe. You can get a complete list of returned columns:
print(list(df.columns))
# Or, for instance, the number of rows and columns in the results:
print(df.shape)
# and use the convenient features of a dataframe, such as built-in plotting:
# Construct a histogram of returned sequence lengths
df.hist('Length', bins=100);
# and grouping/subsetting:
# +
# Subset out pit vipers
pits = df.loc[df["Organism"].str.contains("pit viper")]
pits.head()
# Plot a strip plot of sequence size by organism in the dataframe
output = sns.stripplot(y="Organism", x="Length",
data=pits) # Render strip plot
# -
# <img src="../assets/images/exercise.png" style="width: 100px; float: left;">
#
# <a id="ex03"></a>
# ### Exercise 03 (10min)
#
# Can you use `bioservices`, `UniProt` and `pandas` to:
#
# <br />
# <p></p>
# <div class="alert-danger"><b>
# <ul>
# <li> download a dataframe for all proteins that have "rxlr" in their name
# <li> render a violin plot (<code>sns.violinplot()</code>) that shows the distribution of protein lengths grouped according to the evidence for the protein
# </ul>
# </b></div>
# +
# SOLUTION - EXERCISE 03
# Get dataframe
df = service.get_df("name:rxlr", limit=None)
# Draw violin plot
output = sns.violinplot(y="Protein existence", x="Length", data=df)
# Profit
# -
| notebooks/02-uniprot_programming_with_code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### 1. Which type of network cable is commonly used in backbone networks and telephone companies?
# ##### Ans: fiber-optic cable
# #### 2. Which type of Ethernet cable should be used to directly connect two devices that both use the same pins for transmitting and receiving data?
# ##### Ans: crossover twisted-pair cable
# #### 3. What are three characteristics of UTP cabling? (Choose three.)
# ##### Ans:
# - easiest type of networking cable to install
# - susceptible to EMI and RFI
# - most commonly used networking cable
# #### 4. Which two criteria are used to help select a network medium for a network? (Choose two.)
# ##### Ans:
# - the environment where the selected medium is to be installed
# - the distance the selected medium can successfully carry a signal
# #### 5. Which type of network cable contains multiple copper wires and uses extra shielding to prevent interference?
# ##### Ans: STP
# #### 6. Which type of network cable is used to connect the components that make up satellite communication systems?
# ##### Ans: coaxial
# #### 7. What are two wiring schemes defined by the TIA/EIA organization for Ethernet installation in homes and businesses? (Choose two.)
# ##### Ans:
# - T568A
# - T568B
# #### 8. Which term describes the interference when electrical impulses from one cable cross over to an adjacent cable?
# ##### Ans: crosstalk
# #### 9. What are two sources of electromagnetic interference that can affect data transmissions? (Choose two.)
# ##### Ans:
# - fluorescent light fixture
# - microwave oven
# #### 10. What is the purpose of an IP address?
# ##### Ans: It identifies the source and destination of data packets on a network.
# #### 11. What type of network is defined by two computers that can both send and receive requests for resources?
# ##### Ans: peer-to-peer
# #### 12. Which statement describes the ping and tracert commands?
# ##### Ans: Tracert shows each hop, while ping shows a destination reply only.
# #### 13. What is an advantage of the peer-to-peer network model?
# ##### Ans: ease of setup
| Coursera/Cisco Networking Basics Specializations/Course_1-Internet_Connection_How_to_Get_Online/Week-4/Quiz/Building-a-Simple-Peer-to-Peer-Network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: sina
# language: python
# name: sina
# ---
# Fukushima Heatmap: Subsecting Data
# ===============================
# In some cases, the amount of data available is too much to graph all at once. While the Fukushima set is small enough to fit comfortably in memory, we can still use it to showcase some techniques for handling much larger sets. In this case, we will still process the entire set of data, but do so in coordinate cells; each cell is averaged to create a heatmap. You can configure the number of cells created; the more there are, the less data is held in memory at a time, but the more queries are done overall.
#
#
# Setting number of cells and opening a connection
# -----------------------------------------------------------
#
# As mentioned above, we choose a number of cells that's a reasonable compromise between amount of data in memory and amount of querying needing done (also, a higher number of cells will naturally give a graph with greater fidelity). We then open a connection to our database of interest and find what dates are available to us. We'll track data from each date separately.
# +
from collections import defaultdict
from IPython.display import clear_output, display
import matplotlib.pyplot as plt
from matplotlib import patheffects
import numpy as np
import sina
from sina.utils import get_example_path, DataRange
# Number of cells along a side; this number squared is the total number of cells
CELLS_PER_SIDE = 12
database = get_example_path('fukushima/data.sqlite')
print("Using database {}".format(database))
# Identify the coordinates, label, and label orientation for the power
# plant and selected cities as points of reference.
CITIES = [ # (lon, lat), desc, horizontal alignment
[(141.0281, 37.4213), ' Daiichi Nuclear Power Plant', 'left'],
[(141.0125, 37.4492), 'Futaba ', 'right'],
[(141.0000, 37.4833), ' Namie', 'left'],
[(140.9836, 37.4044), ' Okuma ', 'right'],
[(141.0088, 37.3454), ' Tomioka', 'left']]
# The coordinates our analysis will cover
X_COORDS = (140.9, 141.3)
Y_COORDS = (37.0, 37.83)
# The city coordinates need to be normalized to our grid (whose size depends on CELLS_PER_SIDE)
norm_x = [CELLS_PER_SIDE * ((c[0][0] - X_COORDS[0]) / (X_COORDS[1] - X_COORDS[0])) for c in CITIES]
norm_y = [CELLS_PER_SIDE * ((c[0][1] - Y_COORDS[0]) / (Y_COORDS[1] - Y_COORDS[0])) for c in CITIES]
# Access the data
ds = sina.connect(database)
records = ds.records
relationships = ds.relationships
# Get the ids of the experiments (which are their dates)
dates = list(records.find_with_type("exp", ids_only=True))
print('Database has the following dates available: {}'.format(', '.join(dates)))
# -
# Filter the Data: Filtering Logic
# ========================
# We subdivide our coordinate range (37.3-37.8, 140.9-141.4) into $cells\_per\_side^2$ regions and find the records whose coordinates are within each range. We separate these out based on which day each Record is associated with. We then find that Record's gcnorm (counts per sec) and average to get that cell's average for the day, and also track the total number of records per cell per day (so we know around how confident we are in that average).
#
# This cell adds the functions to memory, plus does a bit of preprocessing. The functions themselves will be called once it's time to create the graph.
# +
# First, we figure out which record ids are associated with which dates
records_at_dates = {}
for date in dates:
records_at_dates[date] = set([str(x.object_id) for x in relationships.find(subject_id=date,
predicate="contains")])
# Jupyter sometimes has an issue with the first call to plt.show(), so we make a dummy call
plt.show()
def calculate_cell(lat_min, lat_max, long_min, long_max):
"""
Calculate the avg_gcnorm and count for the cell across each day in records_at_dates.
:param lat_min: The minimum latitude of this cell (inclusive)
:param lat_max: The maximum latitude of this cell (exclusive)
:param long_min: The minimum longitude of this cell (inclusive)
:param long_max: The maximum longitude of this cell (exclusive)
:returns: a dictionary mapping total and average gcnorm & num samples in a cell to a day
"""
out = defaultdict(lambda: {"total": 0.0, "count": 0, "average": 0.0})
record_ids = set(records.find_with_data(latitude=DataRange(lat_min, lat_max),
longitude=DataRange(long_min, long_max)))
for date in dates:
ids_at_date = record_ids.intersection(records_at_dates[date])
# Quit fast if there's nothing on this day at this coord
if not ids_at_date:
continue
# While we could get_date_for_records() across all record_ids, we expect to run on a system
# that caps the number of SQL variables per query (common practice). So we split by date:
data = records.get_data(id_list=ids_at_date, data_list=["gcnorm"])
for id in ids_at_date:
out[date]["total"] += data[id]["gcnorm"]["value"]
out[date]["count"] += 1
if out[date]["count"] > 0:
out[date]["average"] = out[date]["total"] / (out[date]["count"])
return out
print("Functions loaded and date mappings built!")
# -
# Create the Graphs
# ===============
# Now we divide up based on the number of cells and collect the information for each cell independently. Since we're only *reading* the underlying database, this could, in theory, be parallelized. Generating this graph may take some time; see the progress indicator beneath the code for an idea of how much is left.
# +
x_increment = (X_COORDS[1] - X_COORDS[0]) / CELLS_PER_SIDE
x_range = [x_increment * offset + X_COORDS[0] for offset in range(CELLS_PER_SIDE)]
y_increment = (Y_COORDS[1] - Y_COORDS[0]) / CELLS_PER_SIDE
y_range = [y_increment * offset + Y_COORDS[0] for offset in range(CELLS_PER_SIDE)]
avgs_at_date = defaultdict(lambda: np.zeros((CELLS_PER_SIDE, CELLS_PER_SIDE)))
counts_at_date = defaultdict(lambda: np.zeros((CELLS_PER_SIDE, CELLS_PER_SIDE)))
# This may take awhile! (around a minute for a 12*12 map)
def gen_data():
"""Generate the plot, including calculating the data it contains."""
cells_completed = 0
for x_offset, x_coord in enumerate(x_range):
for y_offset, y_coord in enumerate(y_range):
norms_at_dates = calculate_cell(lat_min=y_coord,
lat_max=y_coord + y_increment,
long_min=x_coord,
long_max=x_coord + x_increment)
cells_completed += 1
if cells_completed % CELLS_PER_SIDE == 0:
progress = ("Progress: {}/{}, finished ([{},{}), [{}, {}))"
.format(cells_completed, CELLS_PER_SIDE ** 2,
'{:.3f}'.format(x_coord),
'{:.3f}'.format(x_coord + x_increment),
'{:.3f}'.format(y_coord),
'{:.3f}'.format(y_coord + y_increment)))
clear_output(wait=True)
display(progress)
for date in norms_at_dates:
avgs_at_date[date][y_offset, x_offset] = (norms_at_dates[date]["average"])
counts_at_date[date][y_offset, x_offset] = (norms_at_dates[date]["count"])
gen_data()
print("All cells calculated! You can now generate the graph (next cell).")
# -
# Configuring and Displaying the Graph
# --------------------------------------------
#
# There's a fair bit of configuration that goes into how the heatmap is displayed. Feel free to tweak these settings to maximize how readable the data is for you personally. Once you're ready (and the cell above has completed), run this cell to display the graph! You can re-run this cell after tweaking the config options to re-create your graph relatively quickly.
# +
# How the cities are marked. Font/marker color and size, label outline and size
COLOR_CITIES = 'white'
COLOR_OUTLINE = 'black'
SIZE_CITY_FONT = 14
SIZE_OUTLINE = 5
AREA_CITIES = 50
# Heatmap colormap, see https://matplotlib.org/users/colormaps.html#grayscale-conversion
COLORMAP = "plasma"
def create_graph():
"""Configure and display the graph itself. Dependent on the data from gen_plot()."""
for date in dates:
fig, ax = plt.subplots(figsize=(9, 9))
plt.xlabel('Longitude')
plt.ylabel('Latitude')
heatmap_avg = ax.imshow(avgs_at_date[date], origin='lower', cmap=COLORMAP)
plt.colorbar(heatmap_avg, label="Counts Per Second")
plt.title("Fukushima Radiation: Flight {}".format(date))
_ = ax.scatter(x=norm_x,
y=norm_y,
s=AREA_CITIES,
c=COLOR_CITIES,
linewidths=SIZE_OUTLINE / 2, # Correction to be around same size as font outline
edgecolor=COLOR_OUTLINE)
for x_coord, y_coord, city_info in zip(norm_x, norm_y, CITIES):
_, desc, alignment = city_info
text = ax.text(x_coord, y_coord, desc,
va="center", ha=alignment,
size=SIZE_CITY_FONT, color=COLOR_CITIES)
text.set_path_effects([patheffects.withStroke(linewidth=SIZE_OUTLINE,
foreground=COLOR_OUTLINE)])
# Matplotlib labels the boxes themselves, rather than their
# borders/the origins, so we need to calculate the centers
ax.set_xticks(range(len(x_range)))
ax.set_xticklabels(('{:.3f}'.format(x + x_increment / 2) for x in x_range))
ax.set_yticks(range(len(y_range)))
ax.set_yticklabels(('{:.3f}'.format(y + y_increment / 2) for y in y_range))
plt.setp(ax.get_xticklabels(), rotation=45, ha="right",
rotation_mode="anchor")
plt.show()
create_graph()
# -
# Releasing Resources
# -------------------------
# Don't forget to release resources when you're all done!
ds.close()
| examples/fukushima/fukushima_subsecting_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="mjx_q5sbW79R" outputId="a186a673-4b32-4e89-a13c-89a2b87b5f94"
from torchvision.datasets.utils import download_url
from torch.utils.data import Dataset, DataLoader, random_split
from torch.nn.utils.rnn import pad_sequence
from torch import nn
from torch.nn import functional as F
from torch import optim
from tqdm.autonotebook import tqdm
from textwrap import wrap
import random
import sys
import io
import random
import numpy as np
import torch
import matplotlib.pyplot as plt
import seaborn as sns
torch.manual_seed(0)
# + colab={"base_uri": "https://localhost:8080/", "height": 84, "referenced_widgets": ["8de9035cf6af40a88278986b578e1c4a", "046139b2ceaa4607b921bb239695cbf8", "6ac68c0cbdb64a9daa1fb4322754a0c6", "6753015b7953433ca379857196331e4f", "<KEY>", "<KEY>", "c5a18aa0eb424b1c85ac636b0be54471", "a9f9e03ff0c5463faf156575d5f49905"]} id="n-BukCCaXf-1" outputId="132442d7-f3cc-4973-970a-893bfe36a425"
class CodeDataset(Dataset):
def __init__(self):
download_url('https://github.com/ecs-vlc/COMP6248/raw/master/exercises/lab7/dataset.txt', '.', 'dataset.txt', None)
with io.open('dataset.txt', 'r') as f:
self.data = f.readlines()
self.PAD='_'
self.SOS='^'
self.EOS='$'
self.PAD_IDX=0
# construct the vocabularies to numericalise the data
self.alphabet = "*".join(self.PAD+self.SOS+self.EOS+"abcdefghijklmnopqrstuvwxyz ").split('*')
self.alphabet_indices = dict((c, i) for i, c in enumerate(self.alphabet))
self.indices_alphabet = dict((i, c) for i, c in enumerate(self.alphabet))
self.morsebet = self.PAD+self.SOS+self.EOS+'.- /'
self.morse_indices = dict((c, i) for i, c in enumerate(self.morsebet))
self.indices_morse = dict((i, c) for i, c in enumerate(self.morsebet))
def encode_alpha(self, inp):
x = torch.zeros(len(inp), dtype=torch.long)
for t, char in enumerate(inp):
x[t] = self.alphabet_indices[char]
return x
def decode_alpha(self, ten, skip_tok=False):
s = ''
ten = ten.view(-1)
for v in ten.view(-1):
if not skip_tok:
s += self.indices_alphabet[v.item()]
elif v>2:
s += self.indices_alphabet[v.item()]
return s
def encode_morse(self, inp):
x = torch.zeros(len(inp), dtype=torch.long)
for t, char in enumerate(inp):
x[t] = self.morse_indices[char]
return x
def decode_morse(self, ten):
s = ''
for v in ten:
s += self.indices_morse[v]
return s
def __len__(self):
return len(self.data)
def __getitem__(self, i):
inp, out = self.data[i].strip().split('|')
x = self.encode_morse(inp)
y = self.encode_alpha(out[::-1])
return x, y
# This will be used to automatically pad all batch items to the same length
def pad_collate(batch):
data = [item[0] for item in batch]
data = pad_sequence(data)
targets = [item[1] for item in batch]
targets = pad_sequence(targets)
return [data, targets]
# Load the data and split randomly into training and val subsets
ds = CodeDataset()
tr, va = random_split(ds, [len(ds) - len(ds)//3, len(ds)//3])
trainloader = DataLoader(tr, batch_size=1024, shuffle=True, collate_fn=pad_collate)
valloader = DataLoader(va, batch_size=1024, shuffle=False, collate_fn=pad_collate)
# + id="nDTCvWVJXj40"
class Encoder(nn.Module):
def __init__(self, input_dim, emb_dim, hid_dim):
super().__init__()
self.hid_dim = hid_dim
self.embedding = nn.Embedding(input_dim, emb_dim)
self.rnn = nn.LSTM(emb_dim, hid_dim)
def forward(self, src):
embedded = self.embedding(src)
lstm_out, (hidden, cell) = self.rnn(embedded)
return hidden, cell
class Decoder(nn.Module):
def __init__(self, output_dim, emb_dim, hid_dim):
super().__init__()
self.output_dim = output_dim
self.hid_dim = hid_dim
self.embedding = nn.Embedding(output_dim, emb_dim)
self.rnn = nn.LSTM(emb_dim, hid_dim)
self.fc_out = nn.Linear(hid_dim, output_dim)
def forward(self, input, hidden, cell):
input = input.unsqueeze(0)
embedded = self.embedding(input)
output, (hidden, cell) = self.rnn(embedded, (hidden, cell))
prediction = self.fc_out(output.squeeze(0))
return prediction, hidden, cell
class Seq2Seq(nn.Module):
def __init__(self, encoder, decoder):
super().__init__()
self.encoder = encoder
self.decoder = decoder
def forward(self, src, trg=None, teacher_forcing_ratio = 0.5, maxlen=5, padding_idx=0):
batch_size = src.shape[1]
trg_len = trg.shape[0] if trg is not None else maxlen
trg_vocab_size = self.decoder.output_dim
outputs = torch.zeros(trg_len, batch_size, trg_vocab_size).to(src.device)
hidden, cell = self.encoder(src)
input = torch.ones(batch_size, dtype=torch.long, device=src.device) * padding_idx
for t in range(1, trg_len):
output, hidden, cell = self.decoder(input, hidden, cell)
outputs[t] = output
teacher_force = random.random() < teacher_forcing_ratio
top1 = output.argmax(1)
input = trg[t] if teacher_force and trg is not None else top1
return outputs
INPUT_DIM = len(ds.morsebet)
OUTPUT_DIM = len(ds.alphabet)
ENC_EMB_DIM = 128
DEC_EMB_DIM = 128
HID_DIM = 256
enc = Encoder(INPUT_DIM, ENC_EMB_DIM, HID_DIM)
dec = Decoder(OUTPUT_DIM, DEC_EMB_DIM, HID_DIM)
model = Seq2Seq(enc, dec).cuda()
# + id="ylZk5LXGXpmf"
def train_val_plot(loss_plot, val_loss_plot, name):
print("Final training loss: ", loss_plot[len(loss_plot)-1])
print("Final validation loss: ", val_loss_plot[len(val_loss_plot)-1])
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(8, 5))
#plt.tight_layout()
ax.plot(loss_plot, label="Training Loss", color="g", linewidth=7)
ax.plot(val_loss_plot, label="Validation Loss", color="r", linewidth=7)
ax.legend(fontsize=18)
ax.grid(True)
ax.set_ylim(bottom=0)
ax.set_title("Training and Validation Loss", fontsize=20);
ax.set_xlabel("Epoch", fontsize=18);
ax.set_ylabel("Loss", fontsize=18);
plt.savefig(name + "_Training.png", format='png', dpi=1200)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000, "referenced_widgets": ["44f88cd8e4e24fed80a9406999136ffb", "e963614b75784745b37a6ee4e79dbd6b", "f50db6c3562c4eef97604fa5418de0d5", "fc8593eb83b54da6ab7ac8c1b24ad8f4", "bbf322c318f941a8a6f7d959c27ddbf4", "e77e4a137e6b45db80ea674f14d070f2", "3c261ac980584c23b07f5ec7cfa21fd8", "a28ac2f174d34e30b605b85419e1f639", "<KEY>", "<KEY>", "d5493125b6b34b5da6c0ac4aac145eb0", "<KEY>", "<KEY>", "f1fe535b233b47f38db4321e13f0be81", "<KEY>", "d8dba396aba14320b5f6f24f29a0d2f3", "651144f367364498aff9574ff8b7a865", "f21633fdecc14786969ac90ae2e7b27e", "18f06ec778e04c998d99e49322391abd", "<KEY>", "f20986de6dc44e58bdc5aab2e196a7bf", "c24a35d1f61844e8ac00c805c9b96d46", "e5411d95148441a488fe3ba010ba4aaf", "ad67378a3ba94c4abd0ee55820e2a57a", "830438c003af4336b97e405d400f11f7", "90eb82df59e14e41bce6dbc28383ed64", "<KEY>", "d8b296e7235a4c29a591c8e9cb254267", "fa85489d41574717b5a223ab14f8e1dd", "8ccd21e2fe764faca8fb4007c819d9b4", "<KEY>", "97776f32b0d7450eaff865c1866d718e", "<KEY>", "8ee7f0644dd0430ab0eb04be7aebb6a5", "414af243bd8e43568b945186ac349ea3", "<KEY>", "<KEY>", "3ce838e12232406a9ffd880e42c195ae", "4c27e0ffacbb4830bcaa18d3f473e57a", "<KEY>", "80a27b8688534da98c29f39329e5a41f", "4264be91b7f443af826f5fd41f1a603b", "dcca6e4e7f204938bdb1ed74c13d354b", "b5a4f45e863d4f3f9c6494d4a213e1bc", "aac9a4bd8e9d4b1e858ee6fa90a28c21", "67acebe12fae471d9aa2bc72ac1bf385", "9571e853f2214238b5018db8e267615d", "2d1d9e3e10e74110b963ed2419c015f7", "1d123cd9ceed49e4892b6384baeddbd3", "<KEY>", "b46590aba2c643cf83c7e15ca4985e60", "<KEY>", "22563ba9bed045b9be4472bd728489b0", "<KEY>", "486b63ffe206418e92592ca8f13baace", "<KEY>", "3347a0624c43470d87809b78475a82d1", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "5550bb38d27a442883241b8e5564f213", "<KEY>", "7acfa15c38b9428eaa9fffd24452d58f", "<KEY>", "<KEY>", "<KEY>", "ac5ad3eb50b9444b9a42b409a241c3f8", "4f7d36a506c04a978a992bb7df1077a4", "<KEY>", "db002199376044e483db7d1d7588569e", "4642e9cd2f8440a7beac9eb90b957769", "<KEY>", "a7d82860909c4eae928268a80c460322", "e2ff733efc1d496d917c9e51b96618bb", "b43927158a254c0598cdea0bf2a9402c", "79af43ed61ad468d92511abee8e5fab7", "<KEY>", "efc912cc720e484bbefcf13174253acd", "<KEY>", "<KEY>", "cb139fc4990049298537e5e11d311e0d", "<KEY>", "e97772d9ffda47a2b9281c3a361f4597", "<KEY>", "2774992a911145eb9e373581ff844229", "<KEY>", "<KEY>", "dc329d8d109349cea94048e5e17f8a2b", "cd99b82085ff4265a7a58f7a28d9ea31", "04264834cbdf404e95a0014554c4a177", "5e121d8170f0483795801b15cbedbec2", "<KEY>", "c9a6e72a38274bc7892935b54a01be15", "<KEY>", "cbd2f1311c084136a6fc29e8233e336d", "<KEY>", "<KEY>", "<KEY>", "acc4738f5e964fce821625f315666931", "d04d80ec1a2d45cdb030f828ae5f245f", "a05ed74b018544da9c5ce05db6ffd903", "<KEY>", "af0e9637a3244c2bace8b80c8a7ee1bb", "<KEY>", "<KEY>", "25afe01576c44f70adc6bed1866f10c2", "<KEY>", "<KEY>", "720d432b6e4c4ef68a2d8fc108ec9035", "<KEY>", "4e4e2a29b2494e258c2d7de0cab95c7b", "<KEY>", "e01290d5e0a14d42b306e18f23c52f5f", "520de697f9b742b88d9b90acb9a1d59c", "<KEY>", "408425c6dd7248898e93f89a8ea1c464", "fe832f5c02ca410e988d2182ddd59b3e", "c7482a1d1a554a9996f9894d55ef16cf", "5482e1a147904b5992d2ace3ba8e1523", "<KEY>", "6e48f071ed8441098ce9039ea26d7c2e", "<KEY>", "3798fa7ee94549168a21113328692def", "3a8e790e49c8459985d10a1fc4de4384", "d6ec3a13971943f7a0e5f9c3efa69b96", "<KEY>", "f26666e43ac04e6499a437b69306abd7", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "96aa6de65e43421a869d8fa5012ab32b", "25b332c655ec4f64a6749f385ba78105", "<KEY>", "<KEY>", "<KEY>", "3b910d692ec34a6698ad130d3594528c", "25d6b44319ef4624bedfeb898595b8ac", "<KEY>", "<KEY>", "dd0ff64d5e624422be45d8067763fb37", "09fafe4d010343da88beb7e98c6f8276", "<KEY>", "<KEY>", "d6809ffeb66f41309426b815c8eeedb1", "<KEY>", "3c2a26e49aca4c819c9485a5d0eb9801", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "576e717737a44f75a4d50970f5c34734", "28ea3b6efc284a7c8c4104a15a68b0a5", "<KEY>", "670d15948b4f48e49d41d8da6cf94860", "f78135797e8f43fc8a27a870de807beb", "2588ad10ccc341cba2adb530651a87f3", "60c547a57b0f4aedb237402fa9230f92", "e00790163d11439ca17fa4ad6625b43a"]} id="ucdEyJKkXrgl" outputId="763f9c6d-ed9d-43f7-a9f0-022ec3941067"
crit = nn.CrossEntropyLoss(ignore_index=ds.PAD_IDX)
opt = optim.Adam(model.parameters())
train_loss, val_loss = [], []
for e in range(10):
model.train()
with tqdm(total=len(trainloader), desc='train') as t:
epoch_loss = 0
for i, (x, y) in enumerate(trainloader):
x = x.cuda()
y = y.cuda()
opt.zero_grad()
pred = model(x, y, padding_idx=ds.PAD_IDX)
pred_dim = pred.shape[-1]
pred = pred[1:].view(-1, pred_dim)
y = y[1:].view(-1)
loss = crit(pred, y)
loss.backward()
opt.step()
epoch_loss = (epoch_loss*i + loss.item()) / (i+1)
t.set_postfix(loss='{:05.3f}'.format(epoch_loss))
t.update()
train_loss.append(epoch_loss)
model.eval()
with tqdm(total=len(valloader), desc='val') as t:
with torch.no_grad():
epoch_loss = 0
for i, (x, y) in enumerate(valloader):
x = x.cuda()
y = y.cuda()
pred = model(x, y, teacher_forcing_ratio=0, padding_idx=ds.PAD_IDX)
pred_dim = pred.shape[-1]
pred = pred[1:].view(-1, pred_dim)
y = y[1:].view(-1)
loss = crit(pred, y)
epoch_loss = (epoch_loss*i + loss.item()) / (i+1)
t.set_postfix(loss='{:05.3f}'.format(epoch_loss))
t.update()
val_loss.append(epoch_loss)
# + colab={"base_uri": "https://localhost:8080/", "height": 398} id="dO1GydVd93zJ" outputId="10eac1cf-a604-4c34-e16b-f23884b946ce"
name = 'lab7'
train_val_plot(train_loss, val_loss, name="")
# + id="yF82Q8VP96sY"
torch.save(model.state_dict(), '/res')
#model = TheModelClass(*args, **kwargs)
#model.load_state_dict(torch.load(PATH))
#model.eval()
# + colab={"base_uri": "https://localhost:8080/", "height": 36} id="mM6T9zA498nd" outputId="7314983f-a3ab-4aab-a880-a3c18525378f"
def decode(code):
out = ''
for i, chunk in enumerate(code.split(' ')):
num = ds.encode_morse('^ ' + chunk + ' $').unsqueeze(1).cuda()
pred = model(num, teacher_forcing_ratio = 0.5, maxlen=2, padding_idx=0)
_, pred = torch.topk(pred[1:].view(-1, pred_dim), 1)
out += ds.decode_alpha(pred.cpu())[::-1]
return out
decode('.- -. ... .-- . .-. / - .... . / ..-. --- .-.. .-.. --- .-- .. -. --.')
# + colab={"base_uri": "https://localhost:8080/", "height": 36} id="rUBggQxqzbtc" outputId="91330e1e-3001-4ce4-e363-d501d6589a7c"
def decode(code):
out = ''
for i, chunk in enumerate(code.split(' ')):
num = ds.encode_morse('^ ' + chunk + ' $').unsqueeze(1).cuda()
pred = model(num.cuda(), maxlen=2)
pred = pred[1:].view(-1, pred_dim).argmax (-1)
out += ds.decode_alpha(pred.cpu())[::-1]
return out
decode('.- -. ... .-- . .-. / - .... . / ..-. --- .-.. .-.. --- .-- .. -. --.')
# + colab={"base_uri": "https://localhost:8080/", "height": 36} id="7od-hLpK9-d2" outputId="3ff84b7c-3c32-4818-caeb-318232c1ff11"
decode('.-- .... -.-- / .. ... / - .... . / --- .-. -.. . .-. / --- ..-. / - .... . / --- ..- - .--. ..- - / .-. . ...- . .-. ... . -..')
# + colab={"base_uri": "https://localhost:8080/", "height": 36} id="w5hQBaDM9_ul" outputId="f053adad-501d-4a06-aaa2-7222f8843be6"
decode('.-- .... .- - / .. ... / - .... . / .--. --- .. -. - / --- ..-. / - . .- -.-. .... . .-. / ..-. --- .-. -.-. .. -. --.')
# + colab={"base_uri": "https://localhost:8080/", "height": 36} id="in8j8KW8-CS2" outputId="75af4e5e-cac6-4f84-e45e-d13faf603c50"
decode('- .-. .- -. ... ..-. --- .-. -- .. -. --. / ... . --.- ..- . -. -.-. . ...')
# + colab={"base_uri": "https://localhost:8080/", "height": 36} id="dhqAyjJS-Ike" outputId="6c280d7b-0a37-41b1-8371-b1075decd10d"
def decode2(code):
out = ''
for i, chunk in enumerate(code.split(' / ')):
num = ds.encode_morse('^ ' + chunk + ' $').unsqueeze(1).cuda()
pred = model(num, teacher_forcing_ratio = 0.5, maxlen=len(chunk.split(' '))+1, padding_idx=0)
# _, pred = torch.topk(pred[1:].view(-1, pred_dim), 1)
pred = pred[1:].view(-1, pred_dim).argmax(1)
out += ds.decode_alpha(pred.cpu())[::-1] +' '
return out
decode2('.- -. ... .-- . .-. / - .... . / ..-. --- .-.. .-.. --- .-- .. -. --.')
# + colab={"base_uri": "https://localhost:8080/", "height": 36} id="cyGDTMBG-Juw" outputId="1c22e63b-eee0-4ac9-a694-c77654a0485e"
decode2('.-- .... -.-- / .. ... / - .... . / --- .-. -.. . .-. / --- ..-. / - .... . / --- ..- - .--. ..- - / .-. . ...- . .-. ... . -..')
# + colab={"base_uri": "https://localhost:8080/", "height": 36} id="2O8qVysO-Ty6" outputId="d0f8bf36-2137-4f70-87fd-9b57efc574f3"
decode2('.-- .... .- - / .. ... / - .... . / .--. --- .. -. - / --- ..-. / - . .- -.-. .... . .-. / ..-. --- .-. -.-. .. -. --.')
# + colab={"base_uri": "https://localhost:8080/", "height": 36} id="4YFH84fc-Vkz" outputId="c627b3bc-4e06-4cd8-9ffa-06b2913a7c26"
decode2('- .-. .- -. ... ..-. --- .-. -- .. -. --. / ... . --.- ..- . -. -.-. . ...')
| Pytorch Projects/DecypherMorseCode.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2.7.16 64-bit
# name: python2716jvsc74a57bd0e7370f93d1d0cde622a1f8e1c04877d8463912d04d973331ad4851f04de6915a
# ---
from sklearn.datasets import load_breast_cancer
data = load_breast_cancer()
data.target[[10, 50, 85]]
list(data.target_names)
| src/tvm/singlestore_sagemaker_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="FgK_h8x4hmKa"
import math
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
from IPython.display import display, HTML
# + [markdown] colab_type="text" id="wQu-eR3feTDs"
# # IE550-A
#
# ### EFC 1 - Data de Entrega: 20/03
# + [markdown] colab_type="text" id="1O9sj6_Webpy"
# ### Questão 1. Convolução
#
# -------------------------
# **(a)** Determine o comprimento P da sequência $y[n]$ gerada na saída do sistema em função de K e D.
# Como tanto a entrada $x[n]$ quanto a resposta ao impulso $h[n]$ são sequencias de comprimento finito, ́e possıvel determinar a sa ́ıda $y[n]$ explorando uma representação vetorial. Seja $y = [y[0], y[1] ... y[P−1]]$ T o vetor que descreve a saída $y[n]$. Então, podemos escrever que: $y = Hx$, (2)
#
# onde $H ∈ R ^ {PxK}$ é a matriz de convolução do sistema e x denota o vetor associado ao sinal de entrada.
#
#
# **Resposta:**
#
# Tendo que a convolução descreve um operador linear entre duas funções que resulta em uma terceira com tamanho total a somatória de ambas, o tamanho do vetor de saída $y[n]$ é igual a $K+D-1$.
#
# + [markdown] colab_type="text" id="820m7n1yQPVS"
# -------------------------
# **(b)** Mostre que este procedimento para o calculo da convolução esta correto, identificando quem e a matriz *H* e o vetor *x*.
#
# **Resposta:**
#
# Se *K*=5 e *D*=3, portanto:
#
# $P = K + D - 1$
#
# $P = 5 + 3 - 1 \Rightarrow P = 7$
#
#
# Exemplo de convolução *K*=2 e *D*=3:
#
# $y[0] = x[0] . h[0]$
#
# $y[1] = x[0] . h[1] + x[1] . h[0]$
#
# $y[2] = x[0] . h[2] + x[1] . h[1] + x[2] . h[0]$
#
# $y[3] = x[1] . h[2] + x[2] . h[1] + x[3] . h[0]$
#
# ...
#
#
# Para criar um programa que calcule a convolução, precisamos gerar uma matriz com a multiplicação entre cada elemento do vetor $x[n]$ com o de $h[n]$. Em seguinda, devemos realizar a somatória da diagonal dessa matriz, a qual tera como saída o vetor $y[n]$.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 221, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 469, "status": "ok", "timestamp": 1521479920497, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-w1LieAxhQKw/AAAAAAAAAAI/AAAAAAAAFJs/zKX2da1ciEE/s50-c-k-no/photo.jpg", "userId": "110919010500485674069"}, "user_tz": 180} id="zkKnkkkeeSCw" outputId="ec0e0239-69d3-47fd-8089-2842be87e31f"
def convolve1D(x, h):
"""Função para calcular a convolução entre dois vetores."""
K, D = len(x), len(h)
S = K + D - 1
# Montamos uma matriz com o calculo entre cada uma das colunas de K e D
matrix = np.zeros((K, D))
for i, k in enumerate(x):
for j, d in enumerate(h):
matrix[i, j] = k * d
# Precisamos inverter a matriz no eixo x, para usar a função np.diagonal abaixo
matrix = np.flip(np.matrix(matrix), axis=1)
# Neste momento, vamos calcular todos os valores das diagonais...
# Vamos somar todos os valores para retornar o vetor da convolução
y = np.array([np.sum(np.diagonal(matrix, axis1=1, axis2=0, offset=m-(D-1))) for m in range(S)])
return y.astype(int)
# Definimos um vetor randômico como x
x = np.random.randint(0, high=10, size=7)
# Definimos um vetor randômico como h
h = np.flip(np.random.randint(0, high=10, size=5), axis=0)
# numpy convolve
np_conv = np.convolve(x, h)
# my convolve
y = convolve1D(x, h)
print('Resultados:')
print('-' * 20)
print('Tam. x: ', len(x))
print('Tam. h: ', len(h))
print('Tam. (x * h): ', len(np_conv))
print('convolve == np.convolve: ', 'Verdadeiro' if np.array_equal(y, np_conv) else 'Falso')
print('-' * 20)
print('Vetores:')
print('x: ', x)
print('h: ', h)
print('y: ', y)
print('np:', np_conv)
# + [markdown] colab_type="text" id="qftJyk76ilX-"
# ### Questão 2. Resposta em Frequência
#
# -------------------------
# Sabendo que é possível controlar o sinal $x[n]$ colocado na entrada do sistema LIT, assim como observar a saida $y[n]$ gerada, proponha uma estratégia que realize a determinação da resposta em frequência $H(e^{jw})$ do sistema, na faixa de frequências $w \in [0, \pi]$ rad. Explique claramente quais conceitos estão sendo explorados na solução proposta, assim como todos os passos envolvidos no processo.
#
# **Resposta:**
#
# Sabendo que:
#
# $Y(e^{jw}) = H(e^{jw}) \times X(e^{jw})$
#
# Dessa maneira, podemos calcular os valores de $H(e^{jw})$ utilizando a operação contrária a multiplicação das transformadas de Fourier de cada sinal:
#
# $H(e^{jw}) = \frac {Y(e^{jw})}{X(e^{jw})}$
#
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="ywhWF4F8i7l2"
# SISTEMA DESCONHECIDO... IGNORAR O CONTEÚDO
def sistema_desconhecido(x):
N = 400
n = np.arange(N)
hn = (1/2) * np.exp(-0.3 * abs(n - 100)) * np.cos(math.pi/10 * n)
y = np.convolve(hn, x)
return y
# + [markdown] colab_type="text" id="-ItViaAQRQdi"
# -------------------------
# A função sistema_desconhecido fornecido contém a implementação de um sistema LIT cujo comportamento não é conhecido. Ele recebe uma sequencia $x[n]$ de entrada e devolve a saída $y[n]$ correspondente. As únicas informações que temos sobre o sistema é que ele possui uma resposta ao impulso $h[n]$ com duração finita, e que $H(e^{jw}), w$ é real. Neste cenário, sua missão é levantar a resposta em frequência do sistema utilizando a estratégia proposta.
#
# **(c)** Uma vez que vamos trabalhar em um computador digital, algumas limitações surgem com relação ao tipo de sequência que pode ser gerada (e.g., somente de comprimento finito). Tendo em mente questões como esta, discuta como a estratégia planejada para obter a resposta em frequência pode ser adaptada, ou, em outras palavras, de que modo os valores de $||H(e^{jw})||$ podem ser obtidos com segurançaa considerando a faixa de frequências sugerida. Mostre os passos do procedimento (i.e., quem é a entrada? Qual foi a saída observada? Como o valor de $||H(e^{jw})||$ foi obtido?) considerando duas frequências específicas.
#
# **(d)** Prepare um programa que faça a captura dos valores de $||H(e^{jw})||$, na faixa de 0 rad a $\pi$ rad, de forma automática.
#
# **(e)** Plote o módulo da resposta em frequência obtida em função de $w$. Por fim, comente de maneira sucinta como o sistema em questão se comporta no domínio da frequência.
#
# **Resposta:**
#
# Vamos colocar como sinal de entrada acordes musicais misturados (através da convolução dos mesmos), para testar a resolução do filtro.
#
# Usando a resposta da questão acima, iremos converter os sinais para sua representação em frequência, usando a transformada de Fourier e depois retornaremos para a representação em tempo usando a transformada inversa.
#
# Farei a divisão do sinal de entrada pelo de saída para encontrarmos o valor do sistema desconhecido e verificaremos sua função matemática.
#
# Primeiro, vamos definir algumas notas musicais e as frequências que elas estão localizadas.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 204, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 459, "status": "ok", "timestamp": 1521479921745, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-w1LieAxhQKw/AAAAAAAAAAI/AAAAAAAAFJs/zKX2da1ciEE/s50-c-k-no/photo.jpg", "userId": "110919010500485674069"}, "user_tz": 180} id="8jMQ0l0rRZzu" outputId="35fcdc30-6aa3-4a55-a8aa-c02391b46251"
df = pd.DataFrame(
[['C',130.82], ['C#',138.59], ['D',146.83], ['D#',155.56], ['E',164.81], ['F',174.61], ['F#',185], ['G',196], ['G#',207.65], ['A',220], ['A#',233.08], ['B',246.94], ['C',261.63], ['C#',277.18], ['D',293.66], ['D#',311.13], ['E',329.63], ['F',349.23], ['F#',369.99], ['G',392], ['G#',415.3], ['A',440], ['A#',466.16], ['B',493.88], ['C',523.25], ['C#',554.37], ['D',587.33], ['D#',622.25], ['E',659.26], ['F',698.46], ['F#',739.99], ['G',783.99], ['G#',830.61], ['A',880], ['A#',932.33], ['B',987.77], ['C',1046.5], ['C#',1108.73], ['D',1174.66], ['D#',1244.51], ['E',1318.51], ['F',1396.91], ['F#',1479.98], ['G',1567.98], ['G#',1661.22], ['A',1760], ['A#',1864.66], ['B',1975.53], ['C',2093]],
columns=['Note', 'Frequency']
)
display(df.head(5))
# + [markdown] colab_type="text" id="cqkieYtXRItS"
# Criamos duas funções abaixo, para gerar os acordes e os respectivos sinais gerados a partir da convolução deles. Criamos dois sinais diferentes para constatar que o retorno do sistema desconhecido será o mesmo.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 373, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 1260, "status": "ok", "timestamp": 1521479923153, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-w1LieAxhQKw/AAAAAAAAAAI/AAAAAAAAFJs/zKX2da1ciEE/s50-c-k-no/photo.jpg", "userId": "110919010500485674069"}, "user_tz": 180} id="sOzAv33HhUp8" outputId="c57c0779-9d30-4825-c98b-aedc1952ae20"
def gen_coords(f=440, peak=4096, Fs=44100, duration=1, partial=False):
"""Generate a sine wave of a coord... default is A."""
# Fs = 48000 # sampling rate, Hz, must be integer
# f = 440 # sine frequency, Hz, may be float
# duration = 5 # in seconds, may be float
n = np.arange(Fs * duration)
signal = (peak * np.sin(2 * np.pi * f * n / Fs)).astype('int16')
if partial:
return signal[:1000]
return signal
def create_signal(freqs, partial=False):
"""Mix various coords to create a new signal to process."""
coords = [gen_coords(f=f, partial=partial) for f in freqs]
signal = np.ones([len(coords[0])])
for c in coords:
signal = np.convolve(signal, c)
return signal
signal1 = create_signal([440, 130.82, 349.23], True)
signal2 = create_signal([df.iat[5, 1], df.iat[15, 1], df.iat[12, 1]], True)
plt.figure(1, figsize=(16, 12))
plt.subplot(221)
plt.plot(signal1, label='Signal 1')
plt.title('Sinal de entrada 1')
plt.xlabel('sample(n)')
plt.ylabel('amplitude')
plt.legend()
plt.subplot(222)
plt.plot(signal2, label='Signal 2', color='C1')
plt.title('Sinal de entrada 2')
plt.xlabel('sample(n)')
plt.ylabel('amplitude')
plt.legend()
plt.show()
# + [markdown] colab_type="text" id="7zczoiVic2Yc"
# Utilizando a função que exerce sobre o sinal de entrada uma operação, temos como resultado ambos os sinais de saída abaixo.
# + cellView="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 373, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 1041, "status": "ok", "timestamp": 1521479924216, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-w1LieAxhQKw/AAAAAAAAAAI/AAAAAAAAFJs/zKX2da1ciEE/s50-c-k-no/photo.jpg", "userId": "110919010500485674069"}, "user_tz": 180} id="qZq4yhiecodE" outputId="531fdf76-913e-4ea9-bc4f-2270e9d751c5"
y1 = sistema_desconhecido(signal1)
y2 = sistema_desconhecido(signal2)
plt.figure(1, figsize=(16, 12))
plt.subplot(221)
plt.plot(y1, label='y1')
plt.title('Sinal de saída 1 após passar pelo filtro desconhecido')
plt.xlabel('sample(n)')
plt.ylabel('amplitude')
plt.legend()
plt.subplot(222)
plt.plot(y2, label='y2', color='C1')
plt.title('Sinal de saída 1 após passar pelo filtro desconhecido')
plt.xlabel('sample(n)')
plt.ylabel('amplitude')
plt.legend()
plt.show()
# + [markdown] colab_type="text" id="4578rrEJc_eo"
# Podemo verificar em ambos os gráficos abaixo que ambos os sinais de saída sofreram uma mudança na amplitude e fase com relação aos sinais de entradas originais.
#
# Em azul os sinais de entrada e em verde os sinais de saída.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 373, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 1135, "status": "ok", "timestamp": 1521479925678, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-w1LieAxhQKw/AAAAAAAAAAI/AAAAAAAAFJs/zKX2da1ciEE/s50-c-k-no/photo.jpg", "userId": "110919010500485674069"}, "user_tz": 180} id="SGgEem-zggO4" outputId="1e9f3806-83ef-4532-c973-0dcdcbf26d56"
plt.figure(1, figsize=(16, 12))
plt.subplot(221)
plt.plot(signal1, label='entrada')
plt.plot(y1, label='saída')
plt.title('Sinal de entrada/saída 1.')
plt.xlabel('sample(n)')
plt.ylabel('amplitude')
plt.grid(True)
plt.legend()
plt.subplot(222)
plt.plot(signal2, label='entrada')
plt.plot(y2, label='saída')
plt.title('Sinais de entrada/saída 2.')
plt.xlabel('sample(n)')
plt.ylabel('amplitude')
plt.legend()
plt.grid(True)
plt.show()
# + [markdown] colab_type="text" id="Mkxuj4cPjtp2"
# Vamos criar uma função que represente a equeção definida na resposta inicial desta questão:
#
# $H(e^{jw}) = \frac {Y(e^{jw})}{X(e^{jw})}$
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 475, "output_extras": [{"item_id": 1}, {"item_id": 2}]} colab_type="code" executionInfo={"elapsed": 821, "status": "ok", "timestamp": 1521479926646, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-w1LieAxhQKw/AAAAAAAAAAI/AAAAAAAAFJs/zKX2da1ciEE/s50-c-k-no/photo.jpg", "userId": "110919010500485674069"}, "user_tz": 180} id="rvsaw7oLi2iE" outputId="1c7af4b5-b3fb-45ae-9afb-e71d0de7eade"
def descobre_sistema(y, x):
xn = np.zeros(y.shape)
xn[:len(x)] = x
return (np.fft.ifft(np.fft.fft(y) / np.fft.fft(xn))).real
# Filtro do sinal1 == h1
h1 = descobre_sistema(y1, signal1)
# Filtro do sinal2 == h2
h2 = descobre_sistema(y2, signal2)
print('-' * 20)
print('Média de h1 :', np.mean(h1))
print('Média de h2 :', np.mean(h2))
print('Diferença h1 - h2 :', np.sum(h1 - h2))
print('-' * 20)
print()
plt.figure(1, figsize=(16, 12))
plt.subplot(221)
plt.plot(h1, label='h1')
plt.title('Filtro h1 calculado usando F(y1) / F(x1)')
plt.xlabel('sample(n)')
plt.ylabel('amplitude')
plt.legend()
plt.subplot(222)
plt.plot(h2, label='h2', color='C1')
plt.title('Filtro h2 calculado usando F(y2) / F(x2)')
plt.xlabel('sample(n)')
plt.ylabel('amplitude')
plt.legend()
plt.show()
# + [markdown] colab_type="text" id="_XBszDUPkOMy"
# Para confirmar se o sinal do sistema encontrado esta correto iremos restaurar o valor do sinal original realizando o cáluclo inverso.
#
# $X(e^{jw}) = \frac {Y(e^{jw})}{H(e^{jw})}$
#
# É possível notar que a alteração da fase do sinal original gera uma quantidade de amostras extras com valores zeros nos sinais de saída.
#
# Verifica-se que a diferença entre os sinais de entrada original e saídas restaurados é pequeno.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 492, "output_extras": [{"item_id": 1}, {"item_id": 2}]} colab_type="code" executionInfo={"elapsed": 1107, "status": "ok", "timestamp": 1521479927914, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-w1LieAxhQKw/AAAAAAAAAAI/AAAAAAAAFJs/zKX2da1ciEE/s50-c-k-no/photo.jpg", "userId": "110919010500485674069"}, "user_tz": 180} id="JzZvghQ7jFV4" outputId="814d9a66-196a-4137-c435-2320dd8bcef3"
def restaurando_sinal_original(y, h):
hn = np.zeros(y.shape)
hn[:len(h)] = h
return (np.fft.ifft(np.fft.fft(y) / np.fft.fft(hn))).real[:len(h)]
xn1 = restaurando_sinal_original(y1, h1)
print('-' * 20)
print('Média de sinal restaurado y / h :', np.mean(xn1))
print('Média de signal1 :', np.mean(signal1))
print('Igualdade signal 1 == xn1 :', np.array_equal(xn1, signal1))
print('Diferença signal 1 - xn1 :', np.sum(signal1 - xn1[:3997]))
print('-' * 20)
print()
plt.figure(1, figsize=(16, 12))
plt.subplot(221)
plt.plot(xn1, label='xn1')
plt.plot(signal1, label='signal 1')
plt.title('Sobreposição do signal 1 e xn1 (restaurado)')
plt.xlabel('sample(n)')
plt.ylabel('amplitude')
plt.legend()
plt.subplot(222)
plt.plot(xn1[700:1000], label='xn1')
plt.plot(signal1[700:1000], label='signal 1')
plt.title('Zoom da sobreposição do signal 1 e xn1 (restaurado), 300 samples')
plt.xlabel('sample(n)')
plt.ylabel('amplitude')
plt.legend()
plt.show()
| notebooks/efc/efc1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
import numpy as np
import os
import pickle
import random
import sys
from tqdm import tqdm
import tensorflow as tf
# <project_root>/ddnet/ddnet.py
sys.path.insert(0, os.path.join(os.path.abspath(''), '..', 'ddnet'))
import ddnet
# -
# # Initialize the setting
# +
# os.environ["CUDA_DEVICE_ORDER"]="PCI_BUS_ID"
# os.environ["CUDA_VISIBLE_DEVICES"]="0"
# +
random.seed(123)
# directory that contains pickle files
data_dir = os.path.join(os.path.abspath(''), '..', 'data', 'openpose_zeros_all')
# -
# # Helper functions
def data_generator(T, C, le):
"""
Generate X (list of arrays) and Y (array) from a dict
"""
X = T['pose'] # list of arrays
Y = np.zeros(shape=(len(T['label']), C.clc_num)) # 2D array one-hot encoding of labels
Y[range(Y.shape[0]), le.transform(T['label'])] = 1
return X, Y
# +
# helper functions for plotting
# history is a history object from keras
def plot_accuracy(history):
# Plot training & validation accuracy values
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('Model accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
def plot_loss(history):
# Plot training & validation loss values
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model Loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# -
# ## Load and Preprocess Data
# +
Train = pickle.load(open(os.path.join(data_dir, "GT_train_1.pkl"), "rb"))
Test = pickle.load(open(os.path.join(data_dir, "GT_test_1.pkl"), "rb"))
from sklearn import preprocessing
le = preprocessing.LabelEncoder()
le.fit(Train['label'])
# +
C = ddnet.DDNetConfig(frame_length=32, num_joints=25, joint_dim=2, num_classes=21, num_filters=32)
X, Y = data_generator(Train,C,le)
X_test,Y_test = data_generator(Test,C,le)
print(len(X), X[0].shape, Y.shape)
print(len(X_test), X_test[0].shape, Y_test.shape)
# -
# ### Convert Invisible Joints with `nan`
# +
def make_nan(p, copy=True):
"""
Convert 0 values to np.nan
"""
assert isinstance(p, np.ndarray)
q = p.copy() if copy else p
q[q == 0] = np.nan
return q
def has_nan(p):
assert isinstance(p, np.ndarray)
return np.isnan(p).any()
def count_nan(p):
assert isinstance(p, np.ndarray)
return np.isnan(p).sum()
X_nan = list(map(make_nan, X))
X_test_nan = list(map(make_nan, X_test))
print("Video without any nan: {} out of {}".format(len([p for p in X_nan if not has_nan(p)]), len(X_nan)))
print("nan entries in X_nan: {} out of {}".format(sum(map(count_nan, X_nan)), sum([p.size for p in X_nan])))
# -
# ### Preprocessing
# * Select a subset of frequently-detected joints
# * Temporal interpolate
# * Fill the others with mean
# +
def find_top_joints(X_nan, top=15):
"""
Find the indices of the `top` most frequently-detected joints """
count_nan_per_joint = np.array([sum([count_nan(p[:, j, :]) for p in X_nan]) for j in range(X_nan[0].shape[1])])
print(count_nan_per_joint)
# print(np.sort(count_nan_per_joint))
good_joint_idx = np.argsort(count_nan_per_joint)[:top]
return good_joint_idx
good_joint_idx = find_top_joints(X_nan)
print("Good joint indices:", sorted(good_joint_idx.tolist()))
# note: the most frequently visible joints are not the same for train and test
test_good_joint_idx = find_top_joints(X_test_nan)
print("Good joint indices of test set: ", sorted(test_good_joint_idx.tolist()))
HAND_PICKED_GOOD_JOINTS = [0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 12, 13, 15, 16]
good_joint_idx = HAND_PICKED_GOOD_JOINTS
def filter_joints(p, good_joint_idx):
"""
Filter a point by only keeping joints in good_joint_idx
"""
return p[:, good_joint_idx, :]
X_topj = [filter_joints(p, good_joint_idx) for p in X_nan]
X_test_topj = [filter_joints(p, good_joint_idx) for p in X_test_nan]
print("Video with nan before/after selecting top joints: {} / {}".format(
sum(map(has_nan, X_nan)),
sum(map(has_nan, X_topj))
))
print("nan entries in before/after selecting top joints: {} / {}. Total {}".format(
sum(map(count_nan, X_nan)),
sum(map(count_nan, X_topj)),
sum([p.size for p in X_topj])
))
# +
def nan_helper(y):
"""Helper function to handle real indices and logical indices of NaNs.
Input:
- y, 1d numpy array with possible NaNs
Output:
- nans, logical indices of NaNs
- index, a function, with signature indices= index(logical_indices),
to convert logical indices of NaNs to 'equivalent' indices
Example:
>>> # linear interpolation of NaNs
>>> nans, x= nan_helper(y)
>>> y[nans]= np.interp(x(nans), x(~nans), y[~nans])
"""
return np.isnan(y), lambda z: z.nonzero()[0]
def temporal_interp(p):
"""
If a joint is detected in at least one frame in a video,
we interpolate the nan coordinates from other frames.
This is done independently for each joint.
Note: it can still leave some all-nan columns if a joint is never detected in any frame.
"""
p = p.copy()
for j in range(p.shape[1]): # joint
for coord in range(p.shape[2]): # x, y (,z)
view = p[:, j, coord]
if np.isnan(view).all() or not np.isnan(view).any():
continue
nans, idx = nan_helper(view)
view[nans]= np.interp(idx(nans), idx(~nans), view[~nans])
return p
X_interp = list(map(temporal_interp, X_topj))
X_test_interp = list(map(temporal_interp, X_test_topj))
print("Video with nan before/after temporal interp: {} / {}".format(
sum(map(has_nan, X_topj)),
sum(map(has_nan, X_interp))
))
print("nan entries in before/after temporal interp: {} / {}".format(
sum(map(count_nan, X_topj)),
sum(map(count_nan, X_interp))
))
# +
def per_video_normalize(p, copy=True):
"""
For x,y[, z] independently:
Normalize into between -0.5~0.5
"""
q = p.copy() if copy else p
for coord in range(p.shape[2]):
view = q[:, :, coord]
a, b = np.nanmin(view), np.nanmax(view)
# q[:,:, coord] = (view - np.mean(view)) / np.std(view)
view[:] = ((view - a) / (b-a)) - 0.5
return q
X_norm = [per_video_normalize(p) for p in X_interp]
X_test_norm = [per_video_normalize(p) for p in X_test_interp]
# print(X_norm[0][:,:,1])
# print(X_test_norm[0][:,1,1])
# +
# fill in the remaining nans
# def per_frame_fill_mean(p, copy=True):
# """
# For each frame independently:
# for x, y[, z] independently:
# Fill nan entries with the mean of all other joints' coordinates
# This is defnitely not perfect, but may help.
# """
# q = p.copy() if copy else p
# for f in range(q.shape[0]):
# for coord in range(q.shape[2]): # x,y
# view = q[f, :, coord]
# view[np.isnan(view)] = np.nanmean(view)
# return q
def fill_nan_random(p, copy=True, sigma=.5):
"""
Fill nan values with normal distribution
"""
q = p.copy() if copy else p
q[np.isnan(q)] = np.random.randn(np.count_nonzero(np.isnan(q))) * sigma
return q
def fill_nan_uniform(p, copy=True, a=-0.5, b=0.5):
"""
Fill nan values with normal distribution
"""
q = p.copy() if copy else p
q[np.isnan(q)] = np.random.random((np.count_nonzero(np.isnan(q)),)) * (b-a) + a
return q
def fill_nan_bottom(p, copy=True):
q = p.copy() if copy else p
xview = q[:,:,0]
xview[np.isnan(xview)] = 0.
yview = q[:,:,1]
yview[np.isnan(yview)] = 0.5
return q
# def fill_nan_col_random(p, copy=True, sigma=.1)
# """
# Fill each nan column with the same value drawn from normal distribution
# """
# q = p.copy() if copy else p
# for j in range(q.shape[1]):
# for coord in range(q.shape[2]):
# view = q[:, j, coord]
# if np.all(np.isnan(view)):
# view[:] = np.random.randn(1) * sigma
# return q
def augment_nan(X, Y, num=5):
"""
Data augmentation.
X is a list of arrays.
"""
Xa = []
Ya = []
for p1, y1 in zip(X, Y):
Xa.extend([fill_nan_uniform(p1) for _ in range(num)])
Ya.extend([y1] * num)
Ya = np.stack(Ya)
assert len(Xa) == Ya.shape[0]
return Xa, Ya
X_aug, Y_aug = augment_nan(X_norm, Y)
print(len(X_aug), Y_aug.shape)
# X_fillnan = [fill_nan_random(p, sigma=0.5) for p in X_norm]
X_test_fillnan = [fill_nan_random(p, sigma=0.0) for p in X_test_norm]
print(X_norm[0][0], "\n\n", X_aug[0][0], "\n\n", X_aug[1][1])
# assert not any (map(has_nan, X_fillnan))
# X_fillnan[0][:,:,1]
# -
X_input, Y_input = X_aug, Y_aug
X_test_input, Y_test_input = X_test_fillnan, Y_test
print(X_input[0].shape)
# ### DDNet's preprocess and config
# +
# redefine config with new # of joints
C = ddnet.DDNetConfig(frame_length=32, num_joints=len(good_joint_idx), joint_dim=2, num_classes=21, num_filters=32)
X_0, X_1 = ddnet.preprocess_batch(X_input, C)
X_test_0, X_test_1 = ddnet.preprocess_batch(X_test_input, C)
# -
# # Building the model
DD_Net = ddnet.create_DDNet(C)
DD_Net.summary()
# # Train, Test and Save/Load the Model
# ### Train and plot loss/accuracy
# +
import keras
from keras import backend as K
from keras.optimizers import *
# K.set_session(tf.Session(config=tf.ConfigProto(intra_op_parallelism_threads=32, inter_op_parallelism_threads=16)))
lr = 1e-3
DD_Net.compile(loss="categorical_crossentropy",optimizer=adam(lr),metrics=['accuracy'])
lrScheduler = keras.callbacks.ReduceLROnPlateau(monitor='loss', factor=0.5, patience=5, cooldown=5, min_lr=1e-5)
history1 = DD_Net.fit([X_0,X_1],Y_input,
batch_size=len(Y_input),
epochs=800,
verbose=True,
shuffle=True,
callbacks=[lrScheduler],
validation_data=([X_test_0,X_test_1],Y_test_input)
)
lr = 1e-4
DD_Net.compile(loss="categorical_crossentropy",optimizer=adam(lr),metrics=['accuracy'])
lrScheduler = keras.callbacks.ReduceLROnPlateau(monitor='loss', factor=0.5, patience=5, cooldown=5, min_lr=5e-6)
history2 = DD_Net.fit([X_0,X_1],Y_input,
batch_size=len(Y_input),
epochs=600,
verbose=True,
shuffle=True,
callbacks=[lrScheduler],
validation_data=([X_test_0,X_test_1],Y_test_input)
)
# -
# %matplotlib inline
# the first 600 epochs
plot_accuracy(history1)
plot_loss(history1)
# the next 500 epochs
plot_accuracy(history2)
plot_loss(history2)
# ### Plot confusion matrix
Y_test_pred = DD_Net.predict([X_test_0, X_test_1])
# +
Y_test_pred_cls = np.argmax(Y_test_pred, axis=1)
Y_test_cls = np.argmax(Y_test, axis=1)
Y_test_cls[:10], Y_test_pred_cls[:10]
from sklearn.metrics import confusion_matrix, ConfusionMatrixDisplay
normalize= None # 'true'
cm = confusion_matrix(Y_test_cls, Y_test_pred_cls, normalize=normalize)
# print(cm)
# print(np.sum(np.diagonal(cm)) / np.sum(cm)) # accuracy
disp = ConfusionMatrixDisplay(
confusion_matrix=cm,
display_labels=le.classes_)
fig, ax = plt.subplots(figsize=(10,10))
disp.plot(xticks_rotation=90, ax=ax)
# -
# ### Save/Load Model
model_path = 'jhmdb_lite_model.h5'
ddnet.save_DDNet(DD_Net, model_path)
# Load the model back from disk
new_net = ddnet.load_DDNet(model_path)
# Evaluate against test set, you should get the same accuracy
new_net.evaluate([X_test_0,X_test_1],Y_test)
from keras import backend as K
K.tensorflow_backend._get_available_gpus()
sess = tf.Session(config=tf.ConfigProto(log_device_placement=True))
from tensorflow.python.client import device_lib
print(device_lib.list_local_devices())
# +
##########################
## Imputation
##########################
from sklearn.preprocessing import MinMaxScaler
def per_frame_normalized(p):
# mean and std is calculated within a frame, across all joints
# separately for x,y
mean = np.nanmean(p, axis=(1,))
std = np.nanstd(p, axis=(1,))
return (p - np.expand_dims(mean, 1)) / np.expand_dims(std, 1)
def per_video_normalize(p):
# noramlize x and y separate
mean = np.nanmean(p, axis=(0,1))
std = np.nanstd(p, axis=(0,1))
return (p - mean) / std
# all_frames_normalized = np.concatenate(list(map(per_frame_normalized, X_interp)))
# print(all_frames_normalized.shape)
# print(all_frames_normalized[0])
# print(count_nan(all_frames_normalized))
from sklearn.experimental import enable_iterative_imputer
from sklearn.impute import IterativeImputer, SimpleImputer
imp = IterativeImputer(max_iter=10, random_state=0, initial_strategy='mean', verbose=1)
# imp = SimpleImputer(missing_values=np.nan, strategy='mean')
all_frames_normalized_flat_imputed = imp.fit_transform(all_frames_normalized.reshape((all_frames_normalized.shape[0], -1)))
print(all_frames_normalized_flat_imputed[0])
def impute(p, imp):
# per-frame normalize
mean = np.nanmean(p, axis=(1,))
std = np.nanstd(p, axis=(1,))
p_normalized = (p - np.expand_dims(mean, 1)) / np.expand_dims(std, 1)
# impute
q = p_normalized.reshape((p_normalized.shape[0], -1))
q = imp.transform(q)
q = q.reshape(p.shape)
print(q.shape)
# per-frame de-normalize
return (q * np.expand_dims(std, 1) ) + np.expand_dims(mean, 1)
# def per_frame_impute(p, imp):
# q = np.empty_like(p)
# for i, frame in enumerate(p):
# scaler = MinMaxScaler()
# frame_scaled = scaler.fit_transform(frame)
# f_flat = frame_scaled.reshape((1, -1))
# f_flat_imputed = imp.transform(f_flat)
# f_imputed = f_flat_imputed.reshape(frame.shape)
# frame_imputed = scaler.inverse_transform(f_imputed)
# q[i] = frame_imputed
# return q
print(impute(X_interp[0], imp)[0])
X_imputed = [per_video_normalize(impute(p, imp)) for p in X_interp]
X_test_imputed = [per_video_normalize(impute(p, imp)) for p in X_test_interp]
| JHMDB/jhmdb_openpose.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="ikbLmgDxX8Om" colab_type="code" colab={}
# + id="QzaYGO_OZonA" colab_type="code" colab={}
# + id="zH1zHVLtZyNz" colab_type="code" colab={}
# + id="EmT2NuHFZyRY" colab_type="code" colab={}
# ! pip install fastai2 -q
# + id="Nhc7uqL0ZyU0" colab_type="code" colab={}
# + id="gSnn-sBMZyYX" colab_type="code" colab={}
# + id="-zGtJj_5Zoqy" colab_type="code" colab={}
from fastai2.basics import *
from fastai2.callback.all import *
from fastai2.text.all import *
# + id="sJD8piSuZouo" colab_type="code" colab={}
# + id="Rm018hPiZoxn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="583a873e-a4ba-4821-d1a5-f6feebd92fca"
path = untar_data(URLs.WIKITEXT_TINY)
path.ls()
# + id="t23PviY-Zo1c" colab_type="code" colab={}
# + id="C_uSSBMXZ-NQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 459} outputId="0d92232f-1cee-41e1-ecbf-c00aa17a717f"
df_train = pd.read_csv(path/"train.csv", header=None, names=['text'])
df_valid = pd.read_csv(path/"test.csv", header=None, names=['text'])
df_all = pd.concat([df_train, df_valid])
df_all.head()
# + id="u2ESO-qwZo40" colab_type="code" colab={}
# + id="cVkw6lhRaTnI" colab_type="code" colab={}
# df_train.iloc[-1].values
# + id="ztBAe9WLaWRY" colab_type="code" colab={}
# + id="ztgIUQsvanRj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 17} outputId="3fc65d0b-c395-4a5c-d1fb-baf3ea317654"
splits = [list(range_of(df_train)), list(range(len(df_train), len(df_all)))]
tfms = [attrgetter("text"), Tokenizer.from_df(0), Numericalize(), ToTensor()]
dsets = Datasets(df_all, [tfms], splits=splits, dl_type=LMDataLoader)
# dsets
# + id="CmjkGi_Yar1z" colab_type="code" colab={}
bs, sl = 64, 72
dls = dsets.dataloaders(bs=bs, seq_len=sl)
# + id="oYfZjRwrar5t" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 476} outputId="74fb9087-74f0-423a-9797-f5b562ac6e5e"
dls.show_batch(max_n=5)
# + id="dKeBYW5bar85" colab_type="code" colab={}
# + [markdown] id="5mjRjznHhRkc" colab_type="text"
# # Model
# + id="auYAwUXphRov" colab_type="code" colab={}
config = awd_lstm_lm_config.copy()
config.update({'input_p': 0.6, 'output_p': 0.4, 'weight_p': 0.5, 'embed_p': 0.1, 'hidden_p': 0.2})
model = get_language_model(AWD_LSTM, len(dls.vocab), config=config)
# + id="3sVVYwjuhRsz" colab_type="code" colab={}
opt_func = partial(Adam, wd=0.1, eps=1e-7)
cbs = [MixedPrecision(clip=0.1), ModelReseter, RNNRegularizer(alpha=2, beta=1)]
# + id="rhLvUFV_hR0t" colab_type="code" colab={}
learn = Learner(dls, model, loss_func=CrossEntropyLossFlat(), opt_func=opt_func, cbs=cbs, metrics=[accuracy, Perplexity()])
# + id="VP6mzuQBjnlD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="06733718-ccf1-4525-e54f-bc4121bc9778"
learn.validate()
# + id="QkRsDkhKnVTT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 300} outputId="d01e0959-3968-4c29-fced-7cb5fffd2548"
learn.lr_find()
# + id="P5veLh9vhRyg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="ba814b64-bdd4-4cf7-b38f-1da0cc83b4a7"
learn.fit_one_cycle(6, 3e-2, moms=(0.8, 0.7, 0.8), div=10)
# + id="5J5HOcORlUjA" colab_type="code" colab={}
learn.save("stage1")
# + id="i5ADZpT7lX_6" colab_type="code" colab={}
learn.freeze_to(-2)
# + id="79TZD_9DlZdn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 300} outputId="c4004123-ec32-4900-e2bf-a25f3b2ee4b0"
learn.lr_find()
# + id="jk18KJApiV4t" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="7357803b-3b31-4689-9d8b-01de177bd317"
learn.fit_one_cycle(8, slice(3e-5, 3e-3), moms=(0.8, 0.7, 0.8), div=10)
# + id="2XKFMplatBeI" colab_type="code" colab={}
learn.save("stage2")
# + colab_type="code" id="LGB9XwFDs7Sg" colab={}
learn.unfreeze()
# + colab_type="code" id="tkTE8ekis7Sr" colab={"base_uri": "https://localhost:8080/", "height": 300} outputId="62dd1c6e-4f5e-4e8f-ef8d-e61cb84557f5"
learn.lr_find()
# + colab_type="code" id="2kGf4CmVs7Sv" colab={"base_uri": "https://localhost:8080/", "height": 359} outputId="11cd6125-e8fc-4088-e249-cb4515d78458"
learn.fit_one_cycle(10, slice(1e-6, 1e-4), moms=(0.8, 0.7, 0.8), div=10)
# + id="qf2Dl0kqtDbg" colab_type="code" colab={}
learn.save("stage3")
# + id="pLzs_Mb2uc5Q" colab_type="code" colab={}
# + id="uoWGfdptucty" colab_type="code" colab={}
# + [markdown] id="0EcWlVODanWP" colab_type="text"
# # Credit
# + [markdown] id="XDO11OpVapGE" colab_type="text"
# * https://dev.fast.ai/tutorial.wikitext
# + id="_ksZJwrBarBR" colab_type="code" colab={}
| nbs/26m_wikitext_fastai2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + papermill={"duration": 0.030999, "end_time": "2021-05-21T15:53:18.487220", "exception": false, "start_time": "2021-05-21T15:53:18.456221", "status": "completed"} tags=["injected-parameters"]
# Parameters
name = "2020-01-01-till-2021-02-28-tycho-brahe"
n_rows = None
# + [markdown] papermill={"duration": 0.015624, "end_time": "2021-05-21T15:53:18.518461", "exception": false, "start_time": "2021-05-21T15:53:18.502837", "status": "completed"} tags=[]
# # Clean : trip statistics
# + [markdown] papermill={"duration": 0.015653, "end_time": "2021-05-21T15:53:18.565325", "exception": false, "start_time": "2021-05-21T15:53:18.549672", "status": "completed"} tags=[]
# # Purpose
# There are some outliers in the trip statistics that will need to be removed.
# + [markdown] papermill={"duration": 0.015622, "end_time": "2021-05-21T15:53:18.643400", "exception": false, "start_time": "2021-05-21T15:53:18.627778", "status": "completed"} tags=[]
# # Setup
# + papermill={"duration": 3.700467, "end_time": "2021-05-21T15:53:22.375142", "exception": false, "start_time": "2021-05-21T15:53:18.674675", "status": "completed"} tags=[]
# # %load ../imports.py
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.pyplot as plt
import seaborn as sns
width=20
height=3
plt.rcParams["figure.figsize"] = (width,height)
sns.set(rc={'figure.figsize':(width,height)})
#import seaborn as sns
import os
from collections import OrderedDict
from IPython.display import display
pd.options.display.max_rows = 999
pd.options.display.max_columns = 999
pd.set_option("display.max_columns", None)
import folium
import plotly.express as px
import plotly.graph_objects as go
import sys
import os
import scipy.integrate
import seaborn as sns
import pyarrow as pa
import pyarrow.parquet as pq
import dask.dataframe
sys.path.append('../')
from src.visualization import visualize
import scipy.integrate
try:
import trip_statistics
except:
sys.path.append('../../../../src/models/pipelines/longterm/scripts/prepdata/trip_statistics')
import trip_statistics
try:
import trip_id,prepare_dataset,trips
except:
sys.path.append('../../../../src/models/pipelines/longterm/scripts/prepdata/trip')
import trip_id,prepare_dataset,trips
try:
import clean_statistics
except:
sys.path.append('../../../../src/models/pipelines/longterm/scripts/prepdata/clean_statistics')
import clean_statistics
# + papermill={"duration": 0.234474, "end_time": "2021-05-21T15:53:22.625141", "exception": false, "start_time": "2021-05-21T15:53:22.390667", "status": "completed"} tags=[]
df_stat = trip_statistics.load_output_as_pandas_dataframe('id_statistics.parquet')
df_stat.head()
# + papermill={"duration": 0.171935, "end_time": "2021-05-21T15:53:22.812733", "exception": false, "start_time": "2021-05-21T15:53:22.640798", "status": "completed"} tags=[]
df_stat.shape
# + papermill={"duration": 0.17193, "end_time": "2021-05-21T15:53:23.000185", "exception": false, "start_time": "2021-05-21T15:53:22.828255", "status": "completed"} tags=[]
df_stat.info()
# + papermill={"duration": 0.218799, "end_time": "2021-05-21T15:53:23.234510", "exception": false, "start_time": "2021-05-21T15:53:23.015711", "status": "completed"} tags=[]
df_stat.describe()
# -
# # Clean
df_clean = clean_statistics.process(df=df_stat, path='id_statistics_clean.parquet')
# +
df_compare = df_stat.copy()
df_compare['item'] = 'raw'
df_clean['item'] = 'clean'
df_compare = df_compare.append(df_clean)
# -
sns.displot(df_compare, x='P', hue='item', binwidth=25, aspect=3)
sns.displot(df_compare, x='trip_time', hue='item', binwidth=25, aspect=3)
| notebooks/pipelines/longterm/steps/02.2_clean_trip_statistics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] deletable=true editable=true
# #### New to Plotly?
# Plotly's Python library is free and open source! [Get started](https://plot.ly/python/getting-started/) by downloading the client and [reading the primer](https://plot.ly/python/getting-started/).
# <br>You can set up Plotly to work in [online](https://plot.ly/python/getting-started/#initialization-for-online-plotting) or [offline](https://plot.ly/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plot.ly/python/getting-started/#start-plotting-online).
# <br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
# + deletable=true editable=true
import plotly.plotly as py
import plotly.graph_objs as go
import pandas as pd
# + [markdown] deletable=true editable=true
# #### Import Data
# + deletable=true editable=true
import plotly.plotly as py
import plotly.graph_objs as go
import pandas as pd
# Read data from a csv
z_data = pd.read_csv('https://raw.githubusercontent.com/plotly/datasets/master/api_docs/mt_bruno_elevation.csv')
data = [
go.Surface(
z=z_data.as_matrix()
)
]
layout = go.Layout(
title='Mt Bruno Elevation',
autosize=False,
width=600,
height=600,
margin=dict(
l=65,
r=50,
b=65,
t=90
)
)
fig = go.Figure(data=data, layout=layout)
# + [markdown] deletable=true editable=true
# #### Default Params
# The camera position is determined by three vectors: *up*, *center*, *eye*.
#
# The up vector determines the up direction on the page. The default is $(x=0, y=0, z=1)$, that is, the z-axis points up.
#
# The center vector determines the translation about the center of the scene. By default, there is no translation: the center vector is $(x=0, y=0, z=0)$.
#
# The eye vector determines the camera view point about the origin. The default is $(x=1.25, y=1.25, z=1.25)$.
# + deletable=true editable=true
name = 'default'
camera = dict(
up=dict(x=0, y=0, z=1),
center=dict(x=0, y=0, z=0),
eye=dict(x=1.25, y=1.25, z=1.25)
)
py.iplot(fig, validate=False, filename=name)
# + [markdown] deletable=true editable=true
# #### Lower the View Point
# + deletable=true editable=true
name = 'eye = (x:2, y:2, z:0.1)'
camera = dict(
up=dict(x=0, y=0, z=1),
center=dict(x=0, y=0, z=0),
eye=dict(x=2, y=2, z=0.1)
)
fig['layout'].update(
scene=dict(camera=camera),
title=name
)
py.iplot(fig, validate=False, filename=name)
# + [markdown] deletable=true editable=true
# #### X-Z plane
# + deletable=true editable=true
name = 'eye = (x:0.1, y:2.5, z:0.1)'
camera = dict(
up=dict(x=0, y=0, z=1),
center=dict(x=0, y=0, z=0),
eye=dict(x=0.1, y=2.5, z=0.1)
)
fig['layout'].update(
scene=dict(camera=camera),
title=name
)
py.iplot(fig, validate=False, filename=name)
# + [markdown] deletable=true editable=true
# #### Y-Z plane
# + deletable=true editable=true
name = 'eye = (x:2.5, y:0.1, z:0.1)'
camera = dict(
up=dict(x=0, y=0, z=1),
center=dict(x=0, y=0, z=0),
eye=dict(x=2.5, y=0.1, z=0.1)
)
fig['layout'].update(
scene=dict(camera=camera),
title=name
)
py.iplot(fig, validate=False, filename=name)
# + [markdown] deletable=true editable=true
# #### View from Above
# + deletable=true editable=true
name = 'eye = (x:0.1, y:0.1, z:2.5)'
camera = dict(
up=dict(x=0, y=0, z=1),
center=dict(x=0, y=0, z=0),
eye=dict(x=0.1, y=0.1, z=2.5)
)
fig['layout'].update(
scene=dict(camera=camera),
title=name
)
py.iplot(fig, validate=False, filename=name)
# + [markdown] deletable=true editable=true
# #### Zooming In
# ... by reducing the norm the eye vector.
# + deletable=true editable=true
name = 'eye = (x:0.1, y:0.1, z:1)'
camera = dict(
up=dict(x=0, y=0, z=1),
center=dict(x=0, y=0, z=0),
eye=dict(x=0.1, y=0.1, z=1)
)
fig['layout'].update(
scene=dict(camera=camera),
title=name
)
py.iplot(fig, validate=False, filename=name)
# + [markdown] deletable=true editable=true
# #### Reference
# + [markdown] deletable=true editable=true
# See https://plot.ly/python/reference/#layout-scene-camera for more information and chart attribute options!
# + deletable=true editable=true
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
# ! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'3d-camera-controls.ipynb', 'python/3d-camera-controls/', 'Python 3D Camera Controls | plotly',
'How to Control the Camera in your 3D Charts in Python with Plotly.',
title= 'Python 3D Camera Controls | plotly',
name = '3D Camera Controls',
has_thumbnail='true', thumbnail='thumbnail/3d-camera-controls.jpg',
language='python',
display_as='3d_charts', order=0.108,
ipynb= '~notebook_demo/78')
# + deletable=true editable=true
| _posts/python-v3/3d/3d-camera-controls/3d-camera-controls.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # PCA CHALLENGE
#
# **File:** PCAChallenge.ipynb
#
# **Course:** Data Science Foundations: Data Mining in Python
# # CHALLENGE
#
# In this challenge, I invite you to do the following:
#
# 1. Set up the PCA object.
# 1. Project the data onto the principal directions found by PCA.
# 1. Plot the ratio of variances explained by each direction.
# 1. Create a scatter plot of projected data along the first two principal directions.
# # IMPORT LIBRARIES
import pandas as pd # For dataframes
import matplotlib.pyplot as plt # For plotting data
import seaborn as sns # For plotting data
from sklearn.decomposition import PCA # For PCA
# # LOAD DATA
#
# For this challenge, we'll use the `swiss` dataset, which is saved in the data folder as "swiss.csv." This dataset contains a standardized fertility measure and socio-economic indicators for each of 47 French-speaking provinces of Switzerland at about 1888. (For more information, see https://opr.princeton.edu/archive/pefp/switz.aspx.)
#
# We'll use the complete dataset for this challenge, as opposed to separating it into training and testing sets.
# Imports the data
df = pd.read_csv('data/swiss.csv')
# Shows the first few rows of the training data
df.head()
# +
# Set up the PCA object
pca = PCA()
# Transform the data ('tf' = 'transformed')
df_tf = pca.fit_transform(df)
# Plot the variance explained by each component
plt.plot(pca.explained_variance_ratio_)
# -
# Plot the projected data set on the first two principal components and colors by class
sns.scatterplot(
x=df_tf[:, 0],
y=df_tf[:, 1])
# # CLEAN UP
#
# - If desired, clear the results with Cell > All Output > Clear.
# - Save your work by selecting File > Save and Checkpoint.
# - Shut down the Python kernel and close the file by selecting File > Close and Halt.
| AE_PCAChallenge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="9PmgWaXKRbP8"
# ## Find key data points from multiple documents
#
# Download <a href="https://drive.google.com/file/d/1V6hmJhCqMyR65e4tal1Q70Lc_jvtZm0F/view?usp=sharing">these documents</a>.
#
# They all have an identical structure to them.
#
# Using regex, capture and export as a CSV the following data points in all the documents:
#
# - The case number.
# - Whether the decision was to accept or reject the appeal.
# - The request date.
# - The decision date.
# - Source file name
#
#
#
# + id="qoXViR6c1k-s"
## import libraries
import re
import pandas as pd
import glob
# from google.colab import files ## for google colab only
# +
### COLAB ONLY
## import colab's file uploader
# files.upload()
# -
## path to documents
## my documents are stored in a folder called docs
path = "docs/*.txt"
myfiles = sorted(glob.glob(path))
myfiles
## quick read reminder
for file in myfiles:
with open(file, "r") as my_text_doc:
print(my_text_doc.read())
## find date pattern
date_pat = re.compile(r"request:\s(\w+\s\d{1,2},\s\d{4})")
## find date pattern and store findings in a list
for file in myfiles:
with open(file, "r") as my_doc:
all_text = my_doc.read()
all_text = all_text.lower()
date = date_pat.findall(all_text)
print(date[0])
# +
## call request dates list hold
request_dates_list = []
for file in myfiles:
with open(file, "r") as my_doc:
all_text = my_doc.read()
all_text = all_text.lower()
date = date_pat.findall(all_text)
request_dates_list.append(date[0])
request_dates_list
# +
## add case number pattern and store findings in a list
date_pat = re.compile(r"request:\s(\w+\s\d{1,2},\s\d{4})") ## date regex pattern
case_pat = re.compile(r"case #:\s(\d+\w)") ## case number regex pattern
decision_pat = re.compile(r"decision:\n{1,2}.+is\s(\w+)") ## ## decision regex pattern
decision_date_pat = re.compile(r"decision:\n.*dated\s(\w+\s\d{1,2},\s\d{4})")
## initializing lists
request_dates_list = []
case_list = []
decision_list = []
dec_list = []
dec_date_list =[]
## iterate through docs to find, capture and store relevant data
for file in myfiles:
with open(file, "r") as my_doc:
all_text = my_doc.read()
all_text = all_text.lower()
date = date_pat.findall(all_text)
request_dates_list.append(date[0])
case = case_pat.findall(all_text)
case_list.append(case[0])
decision = decision_pat.findall(all_text)
decision_list.append(decision[0])
dec_list.append(decision_pat.findall(all_text)[0])
dec_date_list.append(decision_date_pat.findall(all_text)[0])
# -
## call different lists to confirm capture
dec_date_list
## call the case number
case_list
# +
## zip all lists together
final_decision = []
for (request_date, case_number, decision, decision_date, source)\
in zip(request_dates_list, case_list, decision_list, dec_date_list, myfiles):
decision_dict = {"request_date": request_date,
"case_number": case_number,
"decision": decision,
"decision_date": decision_date,
"source_file": source}
final_decision.append(decision_dict)
# -
## call final decisions list
final_decision
## export to csv
df = pd.DataFrame(final_decision)
df.to_csv("decisions.csv", encoding = "UTF-8", index = False)
df
| homework/homework_for_week_14_regex_SOLUTION.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
qpath = r'../../' # path to quetzal here
data = r'inputs/'
import sys
sys.path.append(qpath)
# import class
from quetzal.io.gtfs_reader import importer
# -
# ### Read GTFS
feed = importer.GtfsImporter(path=data + r'bilbao.zip', dist_units='m')
feed = feed.clean()
feed.describe()
# Frequency conversion currently work only for one specific service and date, and on one given time period.
# It computes the average headway over this time period.
# ### Restrict to one date and merge services
feed = feed.restrict(dates=['20191210'])
feed.group_services()
feed.describe()
feed = feed.create_shapes()
feed.map_trips(
feed.trips.groupby('route_id').first().trip_id.head(50) # Map the first trip of each route
)
# ### Build patterns
# In a non-frequency based GTFS, a route has many trips, each corresponding to a start at a specific time. It is necessary to group these trips in order to compute their headway. A group of trips is refered as a pattern.
# The default method to build the patterns is by grouping trips based on their ordered list of stop_ids, without knowledge of time or duration.
# Other methods to build the patterns are available: by parent stations, or clusters. See example 4-advanced-patterns.ipynb
feed.build_patterns()
feed.describe()
# ### Convert to frequencies
time_range = ['06:00:00', '09:00:00'] # time format must be HH:MM:SS
feed_f = feed.convert_to_frequencies(time_range=time_range)
feed_f.describe()
# The average headway computed is the interval length divided by the number of trip starts within the interval.
# - a trip starting exactly at the start of the time range (6:00:00) is taken into account
# - a trip starting exactly at the end of the time range (9:00:00) is not considered.
| examples-ipynb/gtfs_reader/.ipynb_checkpoints/3-convert-to-frequency-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="4f3CKqFUqL2-"
# # Synthetic Features and Outliers
# + [markdown] colab_type="text" id="jnKgkN5fHbGy"
# **Learning Objectives:**
# * Create a synthetic feature that is the ratio of two other features
# * Use this new feature as an input to a linear regression model
# * Improve the effectiveness of the model by identifying and clipping (removing) outliers out of the input data
# + [markdown] colab_type="text" id="S8gm6BpqRRuh"
# ## Setup
# -
# Install latest `2.x.x` release for tensorflow
# !pip install tensorflow==2.0.0-beta1
from matplotlib import cm
from matplotlib import gridspec
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
import logging
from packaging import version
from IPython.display import display
pd.options.display.max_rows = 10
pd.options.display.float_format = '{:.1f}'.format
logging.getLogger('tensorflow').disabled = True
# +
import tensorflow as tf
# %load_ext tensorboard
# -
# First, we'll import the California housing data into a *pandas* `DataFrame`:
# + colab={} colab_type="code" id="9D8GgUovHbG0"
from datetime import datetime
import io
logging.getLogger('tensorboard').disabled = True
california_housing_dataframe = pd.read_csv("https://download.mlcc.google.com/mledu-datasets/california_housing_train.csv", sep=",")
california_housing_dataframe = california_housing_dataframe.reindex(
np.random.permutation(california_housing_dataframe.index))
california_housing_dataframe["median_house_value"] /= 1000.0
california_housing_dataframe
# + [markdown] colab_type="text" id="I6kNgrwCO_ms"
# We'll set up our `plot_to_image` function to convert the matplotlib plot specified by figure to a PNG image
# + colab={} colab_type="code" id="sjBzqxL_okLq"
def plot_to_image(figure):
"""Converts the matplotlib plot specified by 'figure' to a PNG image and
returns it. The supplied figure is closed and inaccessible after this call."""
# Save the plot to a PNG in memory.
buf = io.BytesIO()
plt.savefig(buf, format='png')
# Closing the figure prevents it from being displayed directly inside
# the notebook.
plt.close(figure)
buf.seek(0)
# Convert PNG buffer to TF image
image = tf.image.decode_png(buf.getvalue(), channels=4)
# Add the batch dimension
image = tf.expand_dims(image, 0)
return image
# -
# Next, we'll define the function for model training
# + colab={} colab_type="code" id="VgQPftrpHbG3"
def fit_model(learning_rate,
steps_per_epoch,
batch_size,
input_feature):
"""Trains a linear regression model of one feature.
Args:
learning_rate: A `float`, the learning rate.
steps_per_epoch: A non-zero `int`, the total number of training steps. A training step
consists of a forward and backward pass using a single batch.
batch_size: A non-zero `int`, the batch size.
input_feature: A `string` specifying a column from `california_housing_dataframe`
to use as input feature.
Returns:
A Pandas `DataFrame` containing targets and the corresponding predictions done
after training the model.
"""
epochs = 10
features = california_housing_dataframe[[input_feature]].values
label = "median_house_value"
labels = california_housing_dataframe[label].values
model = tf.keras.models.Sequential([
tf.keras.layers.Dense(1, activation='linear', kernel_initializer='zeros')
])
model.compile(optimizer=tf.keras.optimizers.SGD(learning_rate=learning_rate, clipnorm=5.0),
loss='mse',
metrics=[tf.keras.metrics.RootMeanSquaredError()])
sample = california_housing_dataframe.sample(n=300)
logdir = "logs/synthetic_features_and_outliers/plots" + datetime.now().strftime("%Y%m%d-%H%M%S")
scalars_logdir = "logs/synthetic_features_and_outliers/scalars" + datetime.now().strftime("%Y%m%d-%H%M%S")
file_writer = tf.summary.create_file_writer(logdir)
# Set up to plot the state of our model's line each epoch.
def create_plt_params(feature, label, epochs=10):
colors = [cm.coolwarm(x) for x in np.linspace(-1, 1, epochs)]
return (colors,
(sample[feature].min(), sample[feature].max()),
(0, sample[label].max()))
def create_figure(feature, label, epochs=10):
figure = plt.figure(figsize=(15, 6))
plt.title("Learned Line by Epoch")
plt.ylabel(label)
plt.xlabel(feature)
plt.scatter(sample[feature], sample[label])
return figure
colors, x_min_max, y_min_max = create_plt_params(input_feature, label, epochs)
def log(epoch, logs):
root_mean_squared_error = logs["root_mean_squared_error"]
print(" epoch %02d : %0.2f" % (epoch, root_mean_squared_error))
weight, bias = [x.flatten()[0] for x in model.layers[0].get_weights()]
# Apply some math to ensure that the data and line are plotted neatly.
y_extents = np.array(y_min_max)
x_extents = (y_extents - bias) / weight
x_extents = np.maximum(np.minimum(x_extents,
x_min_max[1]),
x_min_max[0])
y_extents = weight * x_extents + bias
figure = create_figure(input_feature, label, epochs)
plt.plot(x_extents, y_extents, color=colors[epoch])
with file_writer.as_default():
tf.summary.image("Learned Line by Epoch",
plot_to_image(figure),
step=epoch)
model_callback = tf.keras.callbacks.LambdaCallback(
on_epoch_end=lambda epoch, logs: log(epoch, logs))
tensorboard_callback = tf.keras.callbacks.TensorBoard(log_dir=scalars_logdir,
update_freq='epoch')
print("Train model...")
print("RMSE (on training data):")
history = model.fit(features,
labels,
epochs=epochs,
steps_per_epoch=steps_per_epoch,
batch_size=batch_size,
callbacks=[model_callback, tensorboard_callback],
verbose=0).history
print("Model training finished.")
calibration_data = pd.DataFrame()
calibration_data["predictions"] = model.predict_on_batch(features).flatten()
calibration_data["targets"] = pd.Series(labels)
display(calibration_data.describe())
root_mean_squared_error = history["root_mean_squared_error"][9]
print("Final RMSE (on training data): %0.2f" % root_mean_squared_error)
return calibration_data
# + [markdown] colab_type="text" id="FJ6xUNVRm-do"
# ## Task 1: Try a Synthetic Feature
#
# Both the `total_rooms` and `population` features count totals for a given city block.
#
# But what if one city block were more densely populated than another? We can explore how block density relates to median house value by creating a synthetic feature that's a ratio of `total_rooms` and `population`.
# + colab={} colab_type="code" id="suyBB7DZnkGj"
# !rm -rf logs/synthetic_features_and_outliers
# -
# In the cell below, create a feature called `rooms_per_person`, and use that as the `input_feature` to `fit_model()`.
# + cellView="both" colab={} colab_type="code" id="5ihcVutnnu1D"
#
# YOUR CODE HERE
#
california_housing_dataframe["rooms_per_person"] =
calibration_data = fit_model(
learning_rate=0.00005,
steps_per_epoch=500,
batch_size=5,
input_feature="rooms_per_person"
)
# -
# What's the best performance you can get with this single feature by tweaking the learning rate? (The better the performance, the better your regression line should fit the data, and the lower
# the final RMSE should be.)
# + colab={} colab_type="code" id="QVAke92Yn0VT"
from google.datalab.ml import TensorBoard
TensorBoard().start('logs/synthetic_features_and_outliers')
# + [markdown] colab_type="text" id="ZjQrZ8mcHFiU"
# ## Task 2: Identify Outliers
#
# We can visualize the performance of our model by creating a scatter plot of predictions vs. target values. Ideally, these would lie on a perfectly correlated diagonal line.
#
# Use Pyplot's [`scatter()`](https://matplotlib.org/gallery/shapes_and_collections/scatter.html) to create a scatter plot of predictions vs. targets, using the rooms-per-person model you trained in Task 1.
#
# Do you see any oddities? Trace these back to the source data by looking at the distribution of values in `rooms_per_person`.
# -
logdir = "logs/synthetic_features_and_outliers/plots"
file_writer = tf.summary.create_file_writer(logdir + datetime.now().strftime("%Y%m%d-%H%M%S"))
# __#TODO:__ Plot a scatter graph to show the scatter points.
# + colab={} colab_type="code" id="P0BDOec4HbG_"
# YOUR CODE HERE
# -
# The calibration data shows most scatter points aligned to a line. The line is almost vertical, but we'll come back to that later. Right now let's focus on the ones that deviate from the line. We notice that they are relatively few in number.
# If we plot a histogram of `rooms_per_person`, we find that we have a few outliers in our input data:
figure = plt.figure()
plt.subplot(1, 2, 2)
_ = california_housing_dataframe["rooms_per_person"].hist()
with file_writer.as_default():
tf.summary.image("Rooms per person",
plot_to_image(figure),
step=0)
# + colab={} colab_type="code" id="ZquKIjwkq9QA"
TensorBoard().start('logs/synthetic_features_and_outliers')
# + [markdown] colab_type="text" id="9l0KYpBQu8ed"
# ## Task 3: Clip Outliers
#
# See if you can further improve the model fit by setting the outlier values of `rooms_per_person` to some reasonable minimum or maximum.
#
# For reference, here's a quick example of how to apply a function to a Pandas `Series`:
#
# clipped_feature = my_dataframe["my_feature_name"].apply(lambda x: max(x, 0))
#
# The above `clipped_feature` will have no values less than `0`.
# -
# The histogram we created in Task 2 shows that the majority of values are less than `5`.
# __#TODO:__ Let's clip rooms_per_person to 5, and plot a histogram to double-check the results.
# + colab={} colab_type="code" id="rGxjRoYlHbHC"
# YOUR CODE HERE
# + colab={} colab_type="code" id="9lfR8NyQsxi1"
TensorBoard().start('logs/synthetic_features_and_outliers')
# + [markdown] colab_type="text" id="vO0e1p_aSgKA"
# To verify that clipping worked, let's train again and print the calibration data once more:
# + colab={} colab_type="code" id="ZgSP2HKfSoOH"
calibration_data = fit_model(
learning_rate=0.05,
steps_per_epoch=1000,
batch_size=5,
input_feature="rooms_per_person")
# + colab={} colab_type="code" id="gySE-UgfSony"
file_writer = tf.summary.create_file_writer(logdir + datetime.now().strftime("%Y%m%d-%H%M%S"))
figure = plt.figure()
_ = plt.scatter(calibration_data["predictions"], calibration_data["targets"])
with file_writer.as_default():
tf.summary.image("Predictions vs Targets",
plot_to_image(figure),
step=0)
# + colab={} colab_type="code" id="T0WkfcNQtYkc"
TensorBoard().start('logs/synthetic_features_and_outliers')
| self-paced-labs/tensorflow-2.x/synthetic_features_and_outliers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
# ## algorithm
def encode(parity_bits, data):
n = len(data) + parity_bits
assert 2 ** parity_bits == n + 1
# copy data to code
code = np.zeros(n, dtype=int)
code[np.arange(n) & np.arange(n) + 1 > 0] = data
# parity mask
mask = np.zeros(n, dtype=int)
mask[::2] = 1
# compute parity
i = 0
while i < n:
code[i] = code[i:][mask == 1].sum() & 1
i += i + 1
mask = np.repeat(mask, 2)[:n - i]
# result
return code
def decode(code):
n = len(code)
# parity mask
mask = np.zeros(n, dtype=int)
mask[::2] = 1
# compute parity
error, i = -1, 0
while i < n:
error += (i + 1) * (code[i:][mask == 1].sum() & 1)
i += i + 1
mask = np.repeat(mask, 2)[:n - i]
# fix error
if error >= 0:
code[error] ^= 1
# get data from code
data = code[np.arange(n) & np.arange(n) + 1 > 0]
# result
return error, data
# ## encoding
# +
parity_bits = 3
data = np.random.randint(0, 2, 4)
# generate code
code = encode(parity_bits, data)
print('hamming code', data, '->', code)
# make error
code[3] ^= 1
print('with error', code)
# reconstruct
error, recon = decode(code)
print('error @', error, '->', recon)
# +
parity_bits = 4
data = np.random.randint(0, 2, 11)
# generate code
code = encode(parity_bits, data)
print('hamming code', data, '->', code)
# make error
code[14] ^= 1
print('with error', code)
# reconstruct
error, recon = decode(code)
print('error @', error, '->', recon)
# -
| 100days/day 42 - hamming codes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # **Multiple Sequence Alignment Piepline**
# This notebook documents the configuration of select MSA tools to be used in our work:
# 1. MUSCLE
# 2. MAFFT v7
# 3. T_COFFEE
# 4. PASTA
# 5. SATé
# Thorough literature reviews were conducted and the following functionalities of the above tools were settled on and tested:
# ### **1. MUSCLE:**
# 1. **Large data alignment:** Not the best output: see bash function below
# + language="bash"
# muscle_large() { #muscle aligment of large datasets, long execution times is an issue
# #-maxiters 2: Iterations beyond 2 attempt refinement, which often results in a small improvement, at most
# usage $@
# echo "muscle starting alinment..."
#
# for i in $@
# do
# if [ ! -f $i ]
# then
# echo "input error: file $i is non-existent!"
# elif [[ ( -f $i ) && ( `basename $i` =~ .*\.(afa|fasta|fa) ) ]]
# then
# rename
# echo -e "\nproceeding with file `basename $i`..."
# muscle -in $i -fastaout ${muscle_dest}./aligned/${output_filename}.afa -clwout ${muscle_dest}./aligned/${output_filename}.aln -maxiters 2
# else
# echo "input file error in `basename $i`: input file should be a .fasta file format"
# continue
# fi
# done
# }
# -
# Test run:
# + language="bash"
# pwd
# + language="bash"
# source ./aling.sh
# muscle_large ../data/input/test_data/COI_testa00_data.fasta
# -
# 2. **MSA alignment refinement:** Proved useful in refining the a large alignment, especially from PASTA
# + language="bash"
# muscle_refine() {
# #attempt to improve an existing alignment. Input is an existing MSA
# usage $@
# echo "starting refinment of existing MSAs..."
#
# for i in $@
# do
# if [ ! -f $i ]
# then
# echo "input error: file $i is non-existent!"
# elif [[ ( -f $i ) && ( `basename $i` =~ .*\.(afa|fasta|fa|aln) ) ]]
# then
# rename
# echo -e "\nproceeding with file `basename $i`..."
# muscle -in $i -fastaout ${muscle_dest}./refined/\r${output_filename}.afa -clwout ${muscle_dest}./refined/\r${output_filename}.aln -refine
# else
# echo "input file error in `basename $i`: input file should be a .afa file format"
# continue
# fi
# done
# }
# -
# Test run:(using a PASTA alignment output)
# + language="bash"
# source ./aling.sh
# muscle_refine ../data/output/alignment/pasta_output/aligned/COI_testa00_data.aln
# -
# 3. **MSA profile to profile alignment:** Useful in merging two alignment files of homologous sequences
# + language="bash"
# muscle_p2p() {
# #takes two existing MSAs ("profiles") and aligns them to each other, keeping the columns in each MSA intact.The final alignment is made by inserting columns of gaps into the MSAs as needed. The alignments of sequences in each input MSAs are thus fully preserved in the output alignment.
# usage $@
# unset $in1_outname && echo "good: output filename var in1_outname is empty"
# unset $in2_outname && echo "good: output filename var in2_outname is empty"
#
#
# for i in $@
# do
# if [ $# -ne 2 ]
# then
# echo "input file error: only two input files are allowed!"
# break
# elif [[ ( -f $i ) && ( `basename $i` =~ .*\.(afa|fasta|fa) ) ]]
# then
# if [ -z $in1_outname ]
# then
# rename $i
# in1_outname=$output_filename
# else
# rename $i
# in2_outname=$output_filename
# fi
# continue
# elif [ ! -f $i ]
# then
# echo "input error: file $i is non-existent!"
# break
# else
# echo "input file error in `basename $i`: input file should be a .afa file format"
# break
# fi
# done
#
# if [[ ( -n $in1_outname ) && ( -n $in2_outname ) ]]
# then
# echo -e "\nproceeding with file `basename $1` and `basename $2`..."
# muscle -profile -in1 $1 -in2 $2 -fastaout ${muscle_dest}./merged/${in1_outname}_${in2_outname}.afa -clwout ${muscle_dest}./merged/${in1_outname}_${in2_outname}.aln
# else
# echo " A error with output_filenames: in1_outname and in2_outname "
# fi
# }
# -
# Test run: Using two PASTA alignment output files
# + language="bash"
# source ./aling.sh
# muscle_p2p ../data/output/alignment/pasta_output/aligned/COI_testa00_data.aln ../data/output/alignment/pasta_output/aligned/COI_testa01_data.aln
# -
# ### **2. MAFFT:**
# MAFFT version 7 was used and the following functionalities tested:
# 1. **G-INS-1 option for large data alignment**: Just as accurate as MAFFT global alignment option G-INS-1
# + language="bash"
# ###mafft :highly similar ∼50,000 – ∼100,000 sequences × ∼5,000 sites incl. gaps (2016/Jul)
#
# ##G-INS-1 option is applicable to large data, when huge RAM and a large number of CPU cores are available (at most 26.0 GB)
# #By a new flag, --large, the G-INS-1 option has become applicable to large data without using huge RAM.This option uses files, instead of RAM, to store temporary data. The default location of temporary files is $HOME/maffttmp/ (linux, mac and cygwin) or %TMP% (windows) and can be changed by setting the MAFFT_TMPDIR environmental variable.
#
# #syntax: mafft --large --globalpair --thread n in > out
#
# mafft_GlINS1() {
# usage $@
#
# for i in $@
# do
# if [ ! -f $i ]
# then
# echo "input error: file $i is non-existent!"
# elif [[ ( -f $i ) && ( `basename $i` =~ .*\.(afa|fasta|fa) ) ]]
# then
# rename
# echo -e "\nmafft G-large-INS-1 MSA: proceeding with file `basename $i`..."
# printf "Choose from the following output formats: \n"
# select output_formats in fasta_output_format clustal_output_format none_exit
# do
# case $output_formats in
# fasta_output_format)
# echo -e "\nGenerating .fasta output\n"
# mafft --large --globalpair --thread -1 --reorder $i > ${mafft_dest}aligned/${output_filename}.fasta
# break
# ;;
# clustal_output_format)
# echo -e "\nGenerating a clustal format output\n"
# mafft --large --globalpair --thread -1 --reorder --clustalout $1 > ${mafft_dest}aligned/${output_filename}.aln
# break
# ;;
# none_exit)
# break
# ;;
# *) echo "error: Invalid selection!"
# esac
# done
# else
# echo "input file error in `basename $i`: input file should be a .fasta file format"
# continue
# fi
# done
# }
# -
# Test run:
# + language="bash"
# source ./aling.sh
# mafft_GlINS1 ../data/input/test_data/COI_testa00_data.fasta << EOF
# 1
# EOF
# -
# 2. **L-INS-I option for large data aligment:** More accurate than G-INS-I option in some cases but less suitable for large data; in this case we use the MPI(Massage Parsing Interface) version
# + language="bash"
# #MPI version of high-accuracy progressive options, [GLE]-large-INS-1; Two environmental variables, MAFFT_N_THREADS_PER_PROCESS and MAFFT_MPIRUN, have to be set:
# #The number of threads to run in a process: Set "1" unless using a MPI/Pthreads hybrid mode.
# # export MAFFT_N_THREADS_PER_PROCESS="1"
# #Location of mpirun/mpiexec and options: mpirun or mpiexec must be from the same library as mpicc that was used in compiling
# # export MAFFT_MPIRUN="/somewhere/bin/mpirun -n 160 -npernode 16 -bind-to none ..." (for OpenMPI)
# #OR export MAFFT_MPIRUN="/somewhere/bin/mpirun -n 160 -perhost 16 -binding none ..." (for MPICH)
#
# #mpi command: Add "--mpi --large" to the normal command of G-INS-1, L-INS-1 or E-INS-1
# # mafft --mpi --large --localpair --thread 16 input
#
# #mafft L-INS-I command:
# #mafft --localpair --maxiterate 1000 input_file > output_file
#
# mafft_local() {
# usage $@
#
# for i in $@
# do
# if [ ! -f $i ]
# then
# echo "input error: file $i is non-existent!"
# elif [[ ( -f $i ) && ( `basename $i` =~ .*\.(afa|fasta|fa) ) ]]
# then
# rename
# echo -e "\nmafft G-large-INS-1 MSA: proceeding with file `basename $i`..."
# printf "Choose from the following output formats: \n"
# select output_formats in fasta_output_format clustal_output_format none_exit
# do
# case $output_formats in
# fasta_output_format)
# echo -e "\nGenerating .fasta output\n"
# bash $mpionly
# mafft --mpi --large --globalpair --thread -1 --reorder $i > ${mafft_dest}aligned/${output_filename}_l.fasta
# break
# ;;
# clustal_output_format)
# echo -e "\nGenerating a clustal format output\n"
# bash $mpionly
# mafft --mpi --large --globalpair --thread -1 --reorder --clustalout $1 > ${mafft_dest}aligned/${output_filename}_l.aln
# break
# ;;
# none_exit)
# break
# ;;
# *) echo "error: Invalid selection!"
# esac
# done
# else
# echo "input file error in `basename $i`: input file should be a .fasta file format"
# continue
# fi
# done
# }
# -
# Test run:
# + language="bash"
# source ./aling.sh
# mafft_local ../data/input/test_data/COI_testb01_data.aln << EOF
# 1
# EOF
# -
# 3. **MSA merging option**: Just like MUSCLE merging option, it merges two MSA profiles, but with a different strategy.
# + language="bash"
# ##Merge multiple sub-MSAs into a single MSA
# #Each sub-MSA is forecd to form a monophyletic cluster in version 7.043 and higher (2013/May/26).
#
# #syntax: cat subMSA1 subMSA2 > input
# # ruby makemergetable.rb subMSA1 subMSA2 > subMSAtable
# # mafft --merge subMSAtable input > output
#
# mafft_merge() {
# echo -e "\nwarning: Each sub-MSA is forced to form a monophyletic cluster in version 7.043 and higher (2013/May/26)."
# printf "Enter [Yes] to continue or [No] to exit: "
# read choice
# case $choice in
# [yY][eE][sS] | [yY] )
# usage $@ #testing the arguments
# RUBY_EXEC=$( which ruby )
#
# inputfiletest $@ #assesing the validity of the input files
#
# outputfilename $@ # generating output file name
#
# cat $@ > ${mafft_dest}merged/input.fasta
# ${RUBY_EXEC} ${makemergetable} $@ > ${mafft_dest}merged/subMSAtable
# mafft --merge ${mafft_dest}merged/subMSAtable ${mafft_dest}merged/input.fasta > ${mafft_dest}merged/${outname}.fasta
# ;;
# [nN][oO] | [nN] )
# echo "exiting the operation"
# ;;
# *)
# echo "Invalid input: please enter [Yes] or [No]"
# ;;
# esac
#
# }
# -
# Test run: uses two PASTA alignment output files
# + language="bash"
# source ./aling.sh
# mafft_merge ../data/output/alignment/pasta_output/aligned/COI_testa00_data.aln ../data/output/alignment/pasta_output/aligned/COI_testa01_data.aln
# -
# 4. **MAFFT -add option:** For adding unaligned full-length sequence(s) into an existing alignment
# + language="bash"
# #new_sequences and existing_alignment files are single multi-FASTA format file. Gaps in existing_alignment are preserved, but the alignment length may be changed in the default setting. If the --keeplength option is given, then the alignment length is unchanged. Insertions at the new sequences are deleted. --reorder to rearrange sequence order.
#
# #syntax: % mafft --add new_sequences --reorder existing_alignment > output
#
# mafft_add() {
# echo -e "\n \$1 should be the new_sequences: unaligned full-length sequence(s) to be added into the existing alignment (\$2) "
# printf "\nEnter [Yes] to continue or [No] to exit: "
# read choice
# case $choice in
# [yY][eE][sS] | [yY] )
# usage $@
#
# inputfiletest $@
#
# outputfilename $@
#
# mafft --add $1 --reorder $2 > ${mafft_dest}addseq/${outname}.fasta
# ;;
# [nN][oO] | [nN] )
# echo "exiting the --add sequences operation"
# ;;
# *)
# echo "Invalid input: please enter [Yes] or [No]"
# ;;
# esac
# }
# -
# Test run: Uses one PASTA alignment output and an unaligned sample sequence file
# + language="bash"
# source ./aling.sh
# mafft_add ../data/output/alignment/pasta_output/aligned/COI_testa00_data.aln ../data/input/test_data/COI_testb01_data.aln
# -
# 5. **MAFFT --addfragments option:** For adding unaligned fragmentary sequence(s) into an existing alignment
# + language="bash"
# ## --addfragments: Adding unaligned fragmentary sequence(s) into an existing alignment
#
# #syntax: Accurate option
# #syntax: % mafft --addfragments fragments --reorder --thread -1 existing_alignment > output
# # Fast option (accurate enough for highly similar sequences):
# # % mafft --addfragments fragments --reorder --6merpair --thread -1 existing_alignment > output
#
# mafft_addfragmets() {
# echo -e "\n \$1 is fragments is a single multi-FASTA format file and \$2 existing_alignment is a single multi-FASTA format file "
# printf "\nEnter [Yes] to continue or [No] to exit: "
# read choice
# case $choice in
# [yY][eE][sS] | [yY] )
# usage $@
#
# inputfiletest $@
#
# outputfilename $@
#
# mafft --addfragments $1 --reorder --thread -1 $2 > ${mafft_dest}addfragments/${outname}.fasta
# ;;
# [nN][oO] | [nN] )
# echo "exiting the --addfragments operation"
# ;;
# *)
# echo "Invalid input: please enter [Yes] or [No]"
# ;;
# esac
# }
# -
# Test run: using sapmles from those with less than 500 nucleotides to those with between 650 to 660 nucleotides
# + language="bash"
# source ./aling.sh
# mafft_addfragments ../data/input/test_data/COI_testd06_data.fasta ../data/output/alignment/pasta_output/aligned/COI_testc04_data.aln
# -
# ### **3. T_Coffee:**
# 1. **T_Coffee regressive mode for very large sequence alignment:** Uses a regressive algorithm opposed to the common progressive algorithm.
# + language="bash"
# # t_coffee: the regressive mode of T-Coffee is meant to align very large datasets with a high accuracy.
# #It starts by aligning the most distantly related sequences first.
# #It uses this guide tree to extract the N most diverse sequences.
# #In this first intermediate MSA, each sequence is either a leaf or the representative of a subtree.
# #The algorithm is re-aplied recursively onto every representative sequence until all sequences have been incorporated in an internediate MSA of max size N.
# #The final MSA is then obtained by merging all the intermediate MSAs into the final MSA.
#
# #Fast and accurate: the regressive alignment is used to align the sequences in FASTA format. The tree is estimated using the mbed method of Clustal Omega (-reg_tree=mbed), the size of the groups is 100 (-reg_nseq=100) and the method used to align the groups is Clustal Omega:
#
# #syntax: $ t_coffee -reg -seq proteases_large.fasta -reg_nseq 100 -reg_tree mbed -reg_method clustalo_msa -outfile proteases_large.aln -outtree proteases_large.mbed
#
# #-seq :provide sequences. must be in FASTA format
# #-reg_tree :defines the method to be used to estimste the tree
# #-outtree :defines the name of newly computed out tree. mbed method of Clustal Omega is used.
# #-outfile** :defines the name of output file of the MSA
# #-reg_nseq :sets the max size of the subsequence alignments; the groups is 100
# #-reg_thread :sets max threads to be used
# #-reg_method** :defines the method to be used to estimate MSA: Clustal Omega
# #-multi_core :Specifies that T-Coffee should be multithreaded or not; by default all relevant steps are parallelized; DEFAULT: templates_jobs_relax_msa_evaluate OR templates_jobs_relax_msa_evaluate (when flag set)
# #-n_core :Number of cores to be used by machine [default=0 => all those defined in the environement]
#
# tcoffee_large() {
# usage $@
# echo "t-coffee starting alinment..."
#
# for i in $@
# do
# if [ ! -f $i ]
# then
# echo "input error: file $i is non-existent!"
# elif [[ ( -f $i ) && ( `basename $i` =~ .*\.(afa|fasta|fa) ) ]]
# then
# rename
# echo -e "\nproceeding with file `basename $i`..."
# t_coffee -reg -multi_core -n_core=32 -seq $i -reg_nseq 100 -reg_tree mbed -reg_method clustalo_msa -outfile ${tcoffee_dest}aligned/${output_filename}.fasta -newtree ${tcoffee_dest}trees/${output_filename}.mbed
# else
# echo "input file error in `basename $i`: input file should be a .fasta file format"
# continue
#
# fi
# done
#
# }
# -
# Test Run:
# + language="bash"
# source ./align.sh
# tcoffee_large ../data/input/test_data/COI_testa00_data.fasta
# -
# 2. **T_Coffee MSA evaluation option CORE index**:
# Useful in scoring the various results from different alignment algorithms and choosing the most accurate/suitable algorithm
# + language="bash"
# ##Evaluating Your Alignment: Most of T_coffee evalution methods are designed for protein sequences (notably structure based methods), however, T-Coffee via sequence-based-methods (TCS and CORE index) offers some possibilities to evaluate also DNA alignments
#
# #The CORE index is the basis of T-Coffee is an estimation of the consistency between your alignment and the computed library( by default a list of pairs of residues that align in possible global and 10 best local pairwise alignments). The higher the consistency, the better the alignment.
# #Computing the CORE index of any alignment: To evaluate any existing alignment with the CORE index, provide that alignment with the -infile flag and specify that you want to evaluate it
#
# #syntax: $ t_coffee -infile=proteases_small_g10.aln -output=html -score
#
# COREindex() { #Evaluating an existing alignment with the CORE index
# usage $@
# echo "t_coffee starting MSA alignment evaluation using CORE index... "
#
# for i in $@
# do
# if [ ! -f $i ]
# then
# echo "input error: file $i is non-existent!"
# elif [[ ( -f $i ) && ( `basename $i` =~ .*\.(afa|fasta|fa|aln) ) ]]
# then
# rename
# outfile_dest
# echo -e "\nproceeding with `basename $i` alignment file evaluatio..."
# t_coffee -infile=$i -multi_core -n_core=32 -output=html -score -outfile ${output_dest}scores/coreindex/${output_filename}.html
# else
# echo "input file error in `basename $i`: input file should be a *.aln file format"
# continue
# fi
# done
# }
# -
# **Test run**
# + language="bash"
# source ./align.sh
# COREindex ../data/output/alignment/pasta_output/aligned/COI_testa00_data.aln ../data/output/alignment/pasta_output/aligned/COI_testa01_data.aln
# -
# 2. **T_Coffee MSA evaluation Transitive Consistency Score (TCS) option**:
# Like CORE index above it is used in scoring the different alignments and choosing the most accurate/suitable algorithm. It's output can also be used in phylogenetic inference to give different weights to the alignment collumns
# + language="bash"
# # However, to evaluate an alignment, the use of Transitive Consistency Score (TCS) procedure is recommended. TCS is an alignment evaluation score that makes it possible to identify the most correct positions in an MSA. It has been shown that these positions are the most likely to be structuraly correct and also the most informative when estimating phylogenetic trees.
# #Evaluating an existing MSA with Transitive Consistency Score (TCS): most informative when used to identify low-scoring portions within an MSA. *.score_ascii file displays the score of the MSA, the sequences and the residues. *.score_html file displays a colored version score of the MSA, the sequences and the residues
#
# #syntax: $ t_coffee -infile sample_seq1.aln -evaluate -output=score_ascii,aln,score_html
#
# TCSeval() { #Evaluating an existing alignment with the TCS
# usage $@
# echo "t_coffee starting MSA alignment evaluation using TCS... "
#
# for i in $@
# do
# if [ ! -f $i ]
# then
# echo "input error: file $i is non-existent!"
# elif [[ ( -f $i ) && ( `basename $i` =~ .*\.(afa|fasta|fa|aln) ) ]]
# then
# rename
# outfile_dest
# if [ -z "$output_dest" ]
# then
# echo -e "\noutput destination folder not set: input can only be sourced from: \n$muscle_dest; \n$mafft_dest; \n$tcoffee_dest; \n$pasta_dest; and \n$sate_dest"
# continue
# else
# echo -e "\nproceeding with `basename $i` alignment file evaluatio..."
# select TCS_library_estimation_format in proba_pair fast_mafft_kalign_muscle_combo none_exit
# do
# case $TCS_library_estimation_format in
# proba_pair)
# echo -e "\nTCS evaluation using default aligner proba_pair"
# t_coffee -multi_core -n_core=32 -infile $i -evaluate -method proba_pair -output=score_ascii,html -outfile ${output_dest}scores/tcs/${output_filename}_score
# break
# ;;
# fast_mafft_kalign_muscle_combo)
# echo -e "\nTCS evaluation using a series of fast multiple aligners; mafft_msa,kalign_msa,muscle_msa. \nThis option is not accurate and can not be relied on in filtering sequences"
# t_coffee -multi_core -n_core=32 -infile $i -evaluate -method mafft_msa,kalign_msa,muscle_msa -output=score_ascii,html -outfile ${output_dest}scores/tcs/${output_filename}_fastscore
# break
# ;;
# none_exit)
# break
# ;;
# *)
# echo "error: Invalid selection!"
# esac
# done
# fi
# else
# echo "input file error in `basename $i`: input file should be a .fasta file format"
# continue
# fi
# done
# }
# -
# **Test run**
# + language="bash"
# source ./align.sh
# TCSeval ../data/output/alignment/pasta_output/aligned/COI_testa00_data.aln ../data/output/alignment/pasta_output/aligned/COI_testa01_data.aln
# -
# Other T_Coffee functionalities to look at
# + language="bash"
# #=====================================================================================
# #Filtering unreliable MSA positions: columns
# #TCS allows you to filter out from your alignment regions that appears unreliable according to the consistency score; the filtering can be made at the residue level or the column level:
# # t_coffee -infile sample_seq1.aln -evaluate -output tcs_residue_filter3,tcs_column_filter3,tcs_residue_lower4
#
# #sample_seq1.tcs_residue_filter3 :All residues with a TCS score lower than 3 are filtered out
# #sample_seq1.tcs_column_filter3 :All columns with a TCS score lower than 3 are filtered out
# #sample_seq1.tcs_residue_lower4 :All residues with a TCS score lower than 3 are in lower case
#
# #t_coffee -infile sample_seq1.aln -evaluate -output tcs_residue_filter1,tcs_column_filter1
#
# #=====================================================================================
#
# ##Estimating the diversity in your alignment:
#
# # The "-other_pg" flag: call a collection of tools that perform other operations: reformatting, evaluating results, comparing methods. After the flag -other_pg, the common T-Coffee flags are not recognized. "-seq_reformat" flag: calls one of several tools to reformat/trim/clean/select your input data but also your output results, from a very powerful reformatting utility named seq_reformat
#
# # "-output" option of "seq_reformat", will output all the pairwise identities, as well as the average level of identity between each sequence and the others:
# # "-output sim_idscore" realign your sequences pairwise so it can accept unaligned or aligned sequences alike. "-output sim" computes the identity using the sequences as they are in your input file so it is only suited for MSAs
#
# #Syntax: $ t_coffee -other_pg seq_reformat -in sample_seq1.aln -output sim
#
# # "-in" and "in2" flags: define the input file(s)
# # "-output" flag: defines output format*
#
# -
# ### **4. PASTA:**
# This was the most important and was used a lot in the actual analysis. It si a progressive algorithm that subsets the data set into a maximum of 200 sequences per subset; aligns the subsets using one of many third party tools (MAFFT L-INS-i) and; merges the subsets using transivity using third party tools (OPAL).
# 1. **PASTA MSA alignment using MAFFT, OPAL and FastTree**:
# Estimates a start tree using HMM, subsets the data, align using Mafft L-INS-I, marge using OPAL and estimate a tree using FastTree, then use this tree in the next iteration to subset the data. set to 3 iterations(default).
# + language="bash"
# #Usage: $run_pasta.py [options] <settings_file1> <settings_file2> ...
# #syntax: $run_pasta.py -i <input_fasta_file> -j <job_name> --temporaries <TEMP_DIR> -o <output_dir>
#
# pasta_aln() { #MSA alignment using pasta
# usage $@
# echo "PASTA starting alinment..."
#
# PYTHON3_EXEC=$( which python3 )
# runpasta=${co1_path}code/tools/pasta_code/pasta/run_pasta.py
#
# for i in $@
# do
# if [ ! -f $i ]
# then
# echo "input error: file $i is non-existent!"
# elif [[ ( -f $i ) && ( `basename $i` =~ .*\.(afa|fasta|fa|aln) ) ]]
# then
# echo -e "\tProceeding with `basename $i`"
# echo -e "\tPlease select the mafft alignment method;\n\tlocal[mafft_linsi] or global[mafft_ginsi]:"
# select type_of_alignment in mafft_linsi mafft_ginsi mafft_linsi_with_starting_tree mafft_ginsi_with_starting_tree none_exit
# do
# case $type_of_alignment in
# mafft_linsi)
# rename
# echo -e "\nDoing local alignment of `basename $i`..."
# ${PYTHON3_EXEC} ${runpasta} --num-cpus=32 --aligner=mafft -i $i -j ${output_filename} --temporaries=${pasta_dest}temporaries/ -o ${pasta_dest}\jobs/
# cp ${pasta_dest}\jobs/*.${output_filename}.aln ${pasta_dest}aligned/ && mv ${pasta_dest}aligned/{*.${output_filename}.aln,${output_filename}.aln}
# cp ${pasta_dest}\jobs/${output_filename}.tre ${pasta_dest}aligned/${output_filename}.tre
# break
# ;;
# mafft_ginsi)
# rename
# echo -e "\nDoing global alignment of `basename $i`..."
# ${PYTHON3_EXEC} ${runpasta} --num-cpus=32 --aligner=ginsi -i $i -j ${output_filename} --temporaries=${pasta_dest}temporaries/ -o ${pasta_dest}\jobs/
# cp ${pasta_dest}\jobs/*.${output_filename}.aln ${pasta_dest}aligned/ && mv ${pasta_dest}aligned/{*.${output_filename}.aln,${output_filename}.aln}
# cp ${pasta_dest}\jobs/${output_filename}*.tre ${pasta_dest}aligned/${output_filename}.tre
# break
# ;;
# mafft_linsi_with_starting_tree)
# rename
# unset start_tree
# echo -e "\nDoing local alignment of `basename $i` using a starting tree..."
# until [[ ( -f "$start_tree" ) && ( `basename -- "$start_tree"` =~ .*\.(tre) ) ]]
# do
# echo -e "\n\tFor the starting tree provide the full path to the file, the filename included."
# read -p "Please enter the file to be used as the starting tree: " start_tree
# done
# ${PYTHON3_EXEC} ${runpasta} --num-cpus=32 --aligner=mafft -i $i -t $start_tree -j ${output_filename} --temporaries=${pasta_dest}temporaries/ -o ${pasta_dest}\jobs/
# cp ${pasta_dest}\jobs/*.${output_filename}.aln ${pasta_dest}aligned/ && mv ${pasta_dest}aligned/{*.${output_filename}.aln,${output_filename}.aln}
# cp ${pasta_dest}\jobs/${output_filename}.tre ${pasta_dest}aligned/${output_filename}.tre
# break
# ;;
# mafft_ginsi_with_starting_tree)
# rename
# unset start_tree
# echo -e "\nDoing global alignment of `basename $i` using a starting tree..."
# until [[ ( -f "$start_tree" ) && ( `basename -- "$start_tree"` =~ .*\.(tre) ) ]]
# do
# echo -e "\n\tFor the starting tree provide the full path to the file, the filename included."
# read -p "Please enter the file to be used as the starting tree: " start_tree
# done
# ${PYTHON3_EXEC} ${runpasta} --num-cpus=32 --aligner=ginsi -i $i -j ${output_filename} --temporaries=${pasta_dest}temporaries/ -o ${pasta_dest}\jobs/
# cp ${pasta_dest}\jobs/*.${output_filename}.aln ${pasta_dest}aligned/ && mv ${pasta_dest}aligned/{*.${output_filename}.aln,${output_filename}.aln}
# cp ${pasta_dest}\jobs/${output_filename}*.tre ${pasta_dest}aligned/${output_filename}.tre
# break
# ;;
# none_exit)
# break
# ;;
# *)
# echo "error: Invalid selection!"
# esac
# done
# else
# echo "input file error in `basename $i`: input file should be a .fasta file format"
# continue
# fi
# done
# }
# -
# **Test run**
# + language="bash"
# source ./align.sh
# pasta_aln ../data/output/alignment/pasta_output/aligned/COI_testa00_data.aln ../data/output/alignment/pasta_output/aligned/COI_testb01_data.fasta ../data/output/alignment/pasta_output/aligned/COI_testc04_data.fasta ../data/output/alignment/pasta_output/aligned/COI_teste07_data.fasta << EOF
# 1
# 2
# 1
# 1
# EOF
# -
# ### **5. UPP:Ultra-large alignments using Phylogeny-aware Profiles**
# "addresses the problem of alignment of very large datasets, potentially containing fragmentary data. UPP can align datasets with up to 1,000,000 sequences"
# Dependent on Python, PASTA and SEPP(SATe-enabled Phylogenetic Placement)
# + language="bash"
# upp_align() { #UPP stands for Ultra-large alignments using Phylogeny-aware Profiles. A modification of SEPP, SATé-Enabled Phylogenetic Placement, for performing alignments of ultra-large and fragmentary datasets.
# #Usage: $ python <bin>/run_upp.py -s <unaligned_sequences>
# #To run UPP with a pre-computed backbone alignment and tree, run
# # $ python <bin>/run_upp.py -s input.fas -a <alignment_file> -t <tree_file>
# #To run the parallelized version of UPP, run
# # $ python <bin>/run_upp.py -s input.fas -x <cpus>
# usage $@
# echo "UPP starting alinment..."
#
# PYTHON3_EXEC=$( which python3 )
# run_upp=${co1_path}code/tools/sepp/run_upp.py
#
# for i in $@
# do
# if [ ! -f $i ]
# then
# echo "input error: file $i is non-existent!"
# elif [[ ( -f $i ) && ( `basename $i` =~ .*\.(afa|fasta|fa|aln) ) ]]
# then
# echo -e "\tProceeding with `basename $i`"
# echo -e "\tPlease select the type of alignment method;\n\tUsing unaligned sequences only[using_sequences_only] or using a backbone[using_precomputed_backbone]:"
# select type_of_alignment in using_sequences_only using_precomputed_backbone none_exit
# do
# case $type_of_alignment in
# using_sequences_only)
# rename
# echo -e "\nDoing Multiple Sequence Alignment of `basename $i` based on the fragmentary sequences alone"
# ${PYTHON3_EXEC} ${run_upp} -s $i -o ${output_filename} --tempdir ${pasta_dest}temporaries/sepp/ -d ${pasta_dest}jobs_upp/ #-x 32
# cp ${pasta_dest}\jobs/*.${output_filename}_alignment.fasta ${pasta_dest}aligned/
# break
# ;;
# using_precomputed_backbone)
# rename
# unset backbone
# unset start_tree
# echo -e "\nDoing Multiple Sequence Alignment of `basename $i` using a backbone alignment and a starting tree..."
#
# until [[ ( -f "$start_tree" ) && ( `basename -- "$start_tree"` =~ .*\.(tre) ) ]]
# do
# echo -e "\n\tFor the starting tree provide the full path to the file, the filename included."
# read -p "Please enter the file to be used as the starting tree: " start_tree
# done
#
# until [[ ( -f "$backbone" ) && ( `basename -- "$backbone"` =~ .*\.(aln|fasta|fa|afa) ) ]]
# do
# echo -e "\n\tFor the backbone alignment provide the full path to the file, the filename included."
# read -p "Please enter the file to be used as the backbone alignment: " backbone
# done
#
# ${PYTHON3_EXEC} ${run_upp} -s $i -a ${backbone} -t ${start_tree} -o ${output_filename} --tempdir ${pasta_dest}temporaries/sepp/ -d ${pasta_dest}jobs_upp/ #-x 32
# cp ${pasta_dest}\jobs/*.${output_filename}_alignment.fasta ${pasta_dest}aligned/
# break
# ;;
# none_exit)
# break
# ;;
# *)
# echo "error: Invalid selection!"
# esac
# done
# else
# echo "input file error in `basename $i`: input file should be a .fasta file format"
# continue
# fi
# done
#
# }
# -
# **Test run:**
# + language="bash"
# source ./align.sh
# upp_align ../data/output/alignment/pasta_output/aligned/COI_testa00_data.aln
| code/03.01.MSA_pipeline_development.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: squidward_env
# language: python
# name: squidward_env
# ---
# # Singular Matrices
#
# Recently I've been working a lot on coding models and functions 'from scratch'. The point is to force myself to understand not only how each model/method/algorithm works, but also really understand the implementations and parts of the models.
#
# One of the problems I've hit with a few different statistical models is singular matrices. In this post I'm going to talk about:
#
# 1. What is a singular matrix?
# 2. Why are singular matrices a problem?
# 3. How can I identify a singular matrix?
# 4. How can I work with/around a singular matrix?
# ## What is a singular matrix?
#
# A singular (or degenerate) matrix is a matrix that can not be inverted. A lot of this post will discuss the bridge between theory and practice, so I'll further specify that a singular amtrix is a matrix that can't theoretically be inverted.
#
# Of course, practically we can do all sorts of silly things. There are many functions that one can run on a computer that will try and 'succeed' at inverting a sigular matrix. I put succeed in quotes as any function that returns a result when trying to invert a singular matrix is returning nonsense. This is why it is important not to blindly trust functions for random packages you find on the internet! On the other hand, there may be matrices that are theoretically invertable but impossible to practically invert for a variety of reasons that I'll discuss in a bit.
#
# First, let's review the definition of 'invertable'. A matrix is invertable if there exists a square ($nxn$) matrix B such that $AB = BA = I$ where $I$ is the identity matrix. Matrix inversion is the process of finding matrix $B$ for matrix $A$ that satisfies the equation above.
#
# **Technical Asside:** I'd like to burrow a little deeper into what a singular matrix is, but it's a bit mathy. Feel free to skip this if you aren't a hard core math nerd. One can show that non-singular coefficient matrices lead to unique solutions for every vector of constants one could choose. Singular matrices, on the other hand, have non-trivial nullspaces (see proof NMTNS at bottom). For vector contraints b, the system $\mathcal{LS}(A,b)$ could be inconsistant (i.e. no solution). However, if $\mathcal{LS}(A,b)$ has at elast one solution $(w)$, then it will have infinitely many solutions (see proof PSPHS)! A system of equations with a singular coefficient matrix will never have a unique solution.
#
# We'll also note that for singular matrices that there will often be a way to write one row of the matrix as a linear combination of the other rows (may also be true for the columns).
# ## Why are singular matrices a problem?
#
# Why are singular matrices a problem? Well, as it turns out, we often need to invert matrices. For example, what if we want to evaluate the probability density function of a multivariate gaussian distribution?
#
# $$
# p(x;\mu,\Sigma)= \frac{1}{(2\pi)^{\frac{n}{2}} \left\lvert \Sigma \right\rvert ^\frac{1}{2}} \exp\biggl( \frac{-1}{2}(x-\mu)^T \Sigma^{-1} (x-\mu) \biggr)
# $$
#
# We would need to find $\Sigma^{-1}$, the inverse of the caovariance matrix. Or what if we wanted to evaulate the posterior of a Gaussian Process Model?
#
# $$
# \overline{\mathcal{f}}_* = k^T_*(K+\sigma^2_nI)^-1y
# $$
#
# $$
# \mathcal{V}[\mathcal{f}_*] = k(x+*,x_*)-k^T_*(K+\sigma^2_nI)^-1k_*
# $$
#
# I borrowed the notation above from Gaussian Processes for MAchine Learning Eq. 2.25-26. I could go on listing examples of important equations that require matrix inversions but I think you get the point.
#
# The problem is, if we ever need to invert a singular matrix we are in big trouble!
# ## How can I identify a singular matrix?
#
# In many classrooms we teach that the simplest way to find out if a matrix is singular is to find the determinant. If the determinant is zero, then the matrix is singular. This would work fine if theory and practice always went hand in hand, but in the real world things can go terribly wrong with using the determinant to find out if a matrix is singular.
#
# Here is a good example (courtesy of anonymous user 85109 on stack exchange). Let's take the determinant of the matrix below.
# +
import numpy as np
arr = np.array([
[16, 2, 3, 13],
[5, 11, 10, 8],
[9, 7, 6, 12],
[4, 14, 15, 1]
])
np.linalg.det(arr)
# -
# Well that's not zero! Awesome, so the matrix must be non-singular, right? Nope. We can see that there is a way to write a row on this matrix as a linear combination of the other raws (and the same for the columns). This implies that the matrix is singular!
#
# Let's check the symbolic determinant to get a second opinion.
# +
import sympy as sym
M = sym.Matrix(arr)
M.det()
# -
# Wow! The symbolic determinant is exactly what we expect for a singular matrix (zero). So why did numpy give us a different answer?
#
# Well, calcualting the determinant of large matrices is very inefficient. A nice approximation that is commonly leveraged by apckages like numpy is to use the product of the diagonal elements of a specific matrix factorization of the array (LU factorization as of version 1.15). Let's look at this factorization below.
# +
import scipy.linalg as la
P, L, U = la.lu(arr)
print(L)
print(U)
print(P)
# -
# The diagonal of the lower triable (L) are all ones and the diagnonal of the upper triangle (U) are all on-zero! This makes for nice easy math then writing statistical/scientific computing packages. We can take the product of the diagonal of the upper triagle to approximate the determinant of the original matrix.
np.prod(np.diag(U))
# We got the same answer as when we called the determinant function from numpy! Neat. Now this LU decomposition technique is super fast, but it relies on floating point arithmatic. The product of the daigonal of the upper triangle is not quite zero as we would expect. This is why using standard functions that calcualte determinants to identify singualr matrices is a bad idea.
#
# Here are a few other weird examples where using the determinant misleads us! Now, the identity matrix is NOT singular,
np.linalg.det(np.eye(100))
# But by multiplying our matrix by a very samll number, we suddenly see a determinant value that is WAY closer to zero than the determinant value for the singular matrix above!
np.linalg.det(0.1*np.eye(100))
# Now this matrix is NOT singular (for any constant $c$ with identiy matrix $I$, $c*I=D$, matrix D is non-singular jsut like $I$), but with a determinant of $1e^{-100}$ we might easily be fooled into thinking that it is....just wait, it gets worse. Look at the example below.
np.linalg.det(.0001*np.eye(100))
# The determinant should jsut be the determinant of $I$ scaled by $.0001^{-100}$...but numpy can't represent that number! Instead the number underflows and becomes zero, thus tricking us into thinking that this matrix is singular. We could easily invert this matrix and get the correct inversion. We can try the same trick with a large constant to get overflow issues (at least this time numpy warns us!).
np.linalg.det(10000*np.eye(100))
# What other tests might we try for identifying if a matrix is singular? One common tool is using the matrix rank. If the rank of an NxM matrix is less than the minimum of N and M, then we call the matrix singular.
#
# The [rank](https://stattrek.com/matrix-algebra/matrix-rank.aspx) of a matrix is defined as either 1) the maximum number of linearly independent column vectors in the matrix or 2) the maximum number of linearly independent row vectors in the matrix. Both definitions are equivalent.
# +
A = .0001*np.eye(100)
rank = np.linalg.matrix_rank(A)
size_M = A.shape[0]
det = np.linalg.det(A)
print("rank {} = dimension {}".format(rank, size_M))
print("determinant {}".format(det))
# -
# The scaled identity matrix from above still fails to pass the determinant test (due to underflow issues), passes the rank test. We can try this for our original array as well!
# +
rank = np.linalg.matrix_rank(arr)
size_M = arr.shape[0]
det = np.linalg.det(arr)
print("rank {} != dimension {}".format(rank, size_M))
print("determinant {}".format(det))
# -
# This array passes the determinant test (even though it is singular), but fails to pass the rank test.
#
# Another test that we can try is the [condition](https://en.wikipedia.org/wiki/Condition_number) test. The condition of a matrix can be thought of as a measure of how easy the matrix is to invert. The best condition is one. The higher the condition number the harder a matrix is to invert and the more errors may propagate through to the inverted matrix. This is nice, because it not only gives us a clue as to whether a matrix is singular, but also whether the matrix is close enough to singular that we can expect errors when computing the inversion on a computer (due to floating point errors and what not).
#
# The condition is technically the norm of a matrix times the norm of it's 'inverse' (or the matrix the computer gets when it tries to invert the matrix). If these two norms are very dissimilar (meaning the norm changed a lot when the matrix was inverted) then we say that the matrix is poorly (or ill) conditioned. The condition number will be high in this case.
#
# Now the computer may still invert ill conditioned matrices. In fact, it takes the same amount of steps to invert a matrix using Gaussian elimination no matter the condition. However, ill conditioned matrices will have many errors in their inverted counter parts (even to the point of being completely useless). The condition becomes a kind of error multiplier.
#
# When solving the linear system $Ax = b$, you might expect that a small error in $b$ would result in a small error in $x$. That’s true if $A$ is well-conditioned. But small changes in $b$ could result in large changes in x if $A$ is ill-conditioned. Any error (like measurement error from real world observations) will be multiplied by poor conditioning (not just floating point errors).
#
# As a rule of thumb in double precision, a condition greater than 1e15 is really bad.
np.linalg.cond(arr)
# Our original matrix (the tricky singular one) has a HUGE condition and is probably even singular based only on looking at the condition. Obviously it will be bad to try to invert this matrix without taking the proper precautions.
#
# One good check is to see if the reciprocal of the condition is larger than than float epsilon. If it is clsot to epsilon then you are bound to run into some issues.
# +
import sys
1.0 / np.linalg.cond(arr) >= sys.float_info.epsilon
# -
# Finally there is [svd](https://en.wikipedia.org/wiki/Singular_value_decomposition) (singular value decomposition). This is what rank and condition are based on! When an of the singular values of a matrix are small compared to the largest singular value...beware!
# +
np.set_printoptions(suppress=True)
singular_values = np.linalg.svd(arr)[1]
max_sv = np.max(singular_values)
min_sv = np.min(singular_values)
min_sv / max_sv
# -
# We notice that the ratio of the largest and smallest value is REALLY small...that's a bad sign. The svd can tell us if a amtrix is close to singularity. If multiple singualr values are really small it can tell us about the matrix rank.
#
# All of the tools above are easy to use and pretty efficient. A careful scientific coder should always check if his/her matrices are invertable.
#
# So what do we do if we find a singular matrix?!
# ## How can I work with/around a singular matrix?
#
# Singular marticies are, as it turns out, a very small subset of the space of all possible square matricies. In fact, if you were to fill matricies with random uniform samples, you would almost NEVER get a singular matrix.
#
# So the easiest trick to work with a singular amtrix is to add a very small value to the diagonal of the matrix to 'nudge' it out of the singular subset.
arr_nudged = arr + np.eye(arr.shape[0])*1e-10
print("Original Matrix Condition: {}".format(np.linalg.cond(arr)))
print("Nudged Matrix Condition: {}".format(np.linalg.cond(arr_nudged)))
# The condition of our nudged matrix is still really big...but not NEARLY as bad as it's original condition! Adding a tiny value like 1e-10 to the diagonal of a covariance matrix (for example) might not change the matrix in any meaningful way from a scientific standpoint, but it can be many fewer errors when calculating the matrix's inverse.
#
# Another good piece of advice to to looka t different methods of inverting matrices. Instead of using [Cramer's formula](https://en.wikipedia.org/wiki/Invertible_matrix#Analytic_solution) or standard funtions like `np.linal.inv`, try using SVD decomposition or LU decomposition. You can even find some very nice numerically stable methods leveraging cholesky decomposition (a favorite for Gaussian Process models).
#
# **Author's Note:** The observant reader may note that earlier I said that singular matrices are very rare...so why worry about them? Well, they are rare in the sense that you are unlikely to stumble arocss one when randomly sampling many common distributions from the exponential family. However, there are good reasons why we may run acorss them commonly in the real world. Covariances matrices, for example, are often build around multiple samples from a test set. Many data points/samples may be identical or very close resulting in rows/columns in the matrix that are identical/close to identical. This is why we regularize the matrix we want to invert by adding a very small number to the 'ridge' or 'principal diagonal' of the matrix (just like in [ridge regression](https://link.springer.com/content/pdf/10.3758/BF03208332.pdf)) in the same way that we might add a noise term in the noisy case of Gaussian Process regression! In layman's terms: this is why we add a small number to the matrix diagonal. If you'd like to read more about this in the case of Gaussian Processes, you can check out equation 3.26 on page 45 of Gaussian Processes for Machine Learning.
# ## Closing Remarks
#
# Well now you know how to find hot singular matrices in your area and even how to work around them! My advice is to always check your matrices before you try to invert them and have a plan for how to treat the matrix if it is poorly conditioned.
# ## Proofs
# I credit these to 'A First Course in Linear Algrebra' by <NAME> from which I took these proofs. I thank Robert for releasing this great reference for free under the GNU open source liscence!
#
# **Theorem NMTNS:** Nonsingular Matrices have Trivial Null Spaces. <br>
# Suppose that $A$ is a square matrix. Then $A$ is nonsingular if and only if the null space of $A$ is the set containing only the zero vector, i.e. $\mathcal{N}(A)=\{0\}$.
#
# Proof: The null space of a square matrix, $A$, is equal to the set of solutions to the homogeneous system, $\mathcal{LS}(A,0)$. A matrix is nonsingular if and only if the set of solutions to the homogeneous system, $\mathcal{LS}(A,0)$, has only a trivial solution. These two observations may be chained together to construct the two proofs necessary for each half of this theorem.
#
# **Theorem PSPHS:** Particular Solution Plus Homogeneous Solutions. <br> Suppose that $w$ s one solution to the linear system of equations $\mathcal{LS}(A,b)$. Then $y$ is a solution to $\mathcal{LS}(A,b)$ if and only if $y=w+z$ for some vector $z \in \mathcal{N}(A)$.
#
# Proof: [PSPHS Proof](http://linear.ups.edu/html/section-LC.html)
| singular_matrices/Singular_Matrices.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_amazonei_mxnet_p36)
# language: python
# name: conda_amazonei_mxnet_p36
# ---
# +
# !pip install TensorFlow
# !pip install gym
# %matplotlib inline
import numpy as np
from rnn.arch import RNN
from vae.arch import VAE
import matplotlib.pyplot as plt
from gym.utils import seeding
from IPython import display
import time
from tensorflow.keras.utils import plot_model
np.set_printoptions(precision=4, suppress = True)
# -
rnn = RNN()
rnn.set_weights('./rnn/weights.h5')
plot_model(rnn.model, to_file='./rnn/model.png', show_shapes=True)
vae = VAE()
vae.set_weights('./vae/weights2.h5')
# +
# obs_data = np.load('./data/obs_data_car_racing_' + str(batch_num) + '.npy')
# action_data = np.load('./data/action_data_car_racing_' + str(batch_num) + '.npy')
# reward_data = np.load('./data/reward_data_car_racing_' + str(batch_num) + '.npy')
# done_data = np.load('./data/done_data_car_racing_' + str(batch_num) + '.npy')
rnn_files = np.load('./data/rnn_files.npz')
rnn_input = rnn_files['rnn_input']
rnn_output = rnn_files['rnn_output']
initial_mu = np.load('./data/initial_z.npz')['initial_mu']
initial_log_var = np.load('./data/initial_z.npz')['initial_log_var']
# +
rollout_files = np.load('./data/rollout/350636408.npz')
obs_file = rollout_files['obs']
action_file = rollout_files['action']
reward_file = rollout_files['reward']
done_file = rollout_files['done']
series_files = np.load('./data/series/350636408.npz')
mu_file = series_files['mu']
log_var_file = series_files['log_var']
action_2_file = series_files['action']
reward_2_file = series_files['reward']
done_2_file = series_files['done']
# +
hot_zs = np.where(np.exp(initial_log_var[0]/2) < 0.5)[0]
hot_zs
# +
GAUSSIAN_MIXTURES = 5
Z_DIM = 32
z_dim = 32
def get_mixture_coef(z_pred):
log_pi, mu, log_sigma = np.split(z_pred, 3, 1)
log_pi = log_pi - np.log(np.sum(np.exp(log_pi), axis = 1, keepdims = True))
return log_pi, mu, log_sigma
# -
def get_pi_idx(x, pdf):
# samples from a categorial distribution
N = pdf.size
accumulate = 0
for i in range(0, N):
accumulate += pdf[i]
if (accumulate >= x):
return i
random_value = np.random.randint(N)
#print('error with sampling ensemble, returning random', random_value)
return random_value
def sample_z(mu, log_sigma):
z = mu + (np.exp(log_sigma)) * np_random.randn(*log_sigma.shape) * 0.5
return z
np_random, seed = seeding.np_random()
# +
def sample_next_mdn_output(obs, h, c):
d = GAUSSIAN_MIXTURES * Z_DIM
# print(np.array([[obs]]))
# print(np.array([h]))
# print(np.array([c]))
# print('------')
# print(np.array([[obs]]).shape)
# print(np.array([h]).shape)
# print(np.array([c]).shape)
# print('------')
out = rnn.forward.predict([np.array([[obs]]),np.array([h]),np.array([c])])
y_pred = out[0][0][0]
new_h = out[1][0]
new_c = out[2][0]
z_pred = y_pred[:3*d]
rew_pred = y_pred[-1]
z_pred = np.reshape(z_pred, [-1, GAUSSIAN_MIXTURES * 3])
log_pi, mu, log_sigma = get_mixture_coef(z_pred)
chosen_log_pi = np.zeros(z_dim)
chosen_mu = np.zeros(z_dim)
chosen_log_sigma = np.zeros(z_dim)
# adjust temperatures
pi = np.copy(log_pi)
# pi -= pi.max()
pi = np.exp(pi)
pi /= pi.sum(axis=1).reshape(z_dim, 1)
# print(pi)
for j in range(z_dim):
idx = get_pi_idx(np_random.rand(), pi[j])
# print(idx)
chosen_log_pi[j] = idx
chosen_mu[j] = mu[j,idx]
chosen_log_sigma[j] = log_sigma[j,idx]
next_z = sample_z(chosen_mu, chosen_log_sigma)
if rew_pred > 0:
next_reward = 1
else:
next_reward = 0
return next_z, chosen_mu, chosen_log_sigma, chosen_log_pi, rew_pred, next_reward, new_h, new_c
# +
run_idx = 10
idx = 0
# real = obs_data[run_idx]
obs = rnn_input[run_idx]
actual = rnn_output[run_idx]
print(obs.shape)
print(actual.shape)
# +
d = GAUSSIAN_MIXTURES * Z_DIM
print(sum(np.abs(obs[idx,hot_zs])))
plt.plot(actual[idx,hot_zs])
y_pred = rnn.model.predict(np.array([[obs[idx,:]]]))
# print(y_pred)
z_pred = y_pred[:,:,:(3*d)]
rew_pred = y_pred[:,:,-1]
print(rew_pred)
z_pred = np.reshape(z_pred, [-1, GAUSSIAN_MIXTURES * 3])
log_pi, mu, log_sigma = np.split(z_pred, 3, 1)
plt.plot(mu[hot_zs])
log_pi = log_pi - np.log(np.sum(np.exp(log_pi), axis = 1, keepdims = True))
print(sum(np.abs(mu)))
# -
current_z = obs[idx]
i = idx
current_h = np.zeros(256)
current_c = np.zeros(256)
next_z = obs[idx,:]
next_h = np.zeros(256)
next_c = np.zeros(256)
# next_z_decoded = vae.decoder.predict(np.array([next_z]))[0]
# plt.imshow( next_z_decoded)
# plt.show()
#SIMULATE TURNING IN THE DREAM
for i in range(300):
next_z, chosen_mu, chosen_log_sigma, chosen_pi, rew_pred, next_reward, next_h, next_c \
= sample_next_mdn_output(next_z, next_h, next_c)
next_z_decoded = vae.decoder.predict(np.array([next_z]))[0]
next_z = np.concatenate([next_z, [1,1,0], [next_reward]])
plt.gca().cla()
plt.imshow(next_z_decoded)
print(next_reward)
display.clear_output(wait=True)
display.display(plt.gcf())
print(i)
# +
b = np.swapaxes(actual,0,1)
print(b.shape)
actual_rew = b[-1,:]
next_zs = np.zeros(shape = (32,299))
next_mus = np.zeros(shape = (32,299))
next_rews = np.zeros(shape = (1,299))
# next_z = np.copy(obs)
# print(next_z.shape)
next_z = obs[idx,:]
next_mus[:,0] = next_z[:32]
next_zs[:,0] = next_z[:32]
next_rews[:,0] = next_z[-1]
for i in range(1,299):
next_z, chosen_mu, chosen_log_sigma, chosen_pi, rew_pred, next_reward, next_h, next_c = sample_next_mdn_output(next_z, next_h, next_c)
next_mus[:,i] = chosen_mu
next_zs[:,i] = next_z
next_rews[:,i] = rew_pred
next_z_decoded = vae.decoder.predict(np.array([next_z]))[0]
# plt.gca().cla()
# plt.imshow( next_z_decoded)
# # plt.show()
# display.clear_output(wait=True)
# display.display(plt.gcf())
next_z = np.concatenate([next_z, [-1,1,0], [next_reward]])
plt.figure(figsize=(20,30))
for i in hot_zs:
plt.subplot(511)
plt.plot( b[i,:])
plt.subplot(512)
plt.plot( next_zs[i,:])
plt.subplot(513)
plt.plot( next_mus[i,:])
plt.subplot(514)
plt.plot(next_rews[0,:])
plt.subplot(515)
plt.plot(actual_rew[:])
# print(next_zs)
# -
| check_04_rnn.ipynb |
# # Prior Distributions with `Bilby`
#
# Prior distributions are a core component of any Bayesian problem and specifying them in codes can be one of the most confusing elements of a code.
# The `prior` modules in `Bilby` provide functionality for specifying prior distributions in a natural way.
#
# We have a range of predefined types of prior distribution and each kind has methods to:
#
# 1. draw samples, `prior.sample`.
# 2. calculate the prior probability, `prior.prob`.
# 3. rescale samples from a unit cube to the prior distribution, `prior.rescale`.
# This is especially useful when using nested samplers as it avoids the need for rejection sampling.
# 4. Calculate the log prior probability, `prior.log_prob`.
#
# In addition to the predefined prior distributions there is functionality to specify your own prior, either from a pair of arrays, or from a file.
#
# Each prior distribution can also be given a `name` and may have a different `latex_label` for plotting.
# If no name is provided, the default is `None` (this should probably by `''`).
# +
import bilby
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
# -
# ## Prior Instantiation
#
# Below we demonstrate instantiating a range of prior distributions.
#
# Note that when a `latex_label` is not specified, the `name` is used.
# +
fig = plt.figure(figsize=(12, 5))
priors = [
bilby.core.prior.Uniform(minimum=5, maximum=50),
bilby.core.prior.LogUniform(minimum=5, maximum=50),
bilby.core.prior.PowerLaw(name="name", alpha=2, minimum=100, maximum=1000),
bilby.gw.prior.UniformComovingVolume(
name="luminosity_distance", minimum=100, maximum=1000, latex_label="label"
),
bilby.gw.prior.AlignedSpin(),
bilby.core.prior.Gaussian(name="name", mu=0, sigma=1, latex_label="label"),
bilby.core.prior.TruncatedGaussian(
name="name", mu=1, sigma=0.4, minimum=-1, maximum=1, latex_label="label"
),
bilby.core.prior.Cosine(name="name", latex_label="label"),
bilby.core.prior.Sine(name="name", latex_label="label"),
bilby.core.prior.Interped(
name="name",
xx=np.linspace(0, 10, 1000),
yy=np.linspace(0, 10, 1000) ** 4,
minimum=3,
maximum=5,
latex_label="label",
),
]
for ii, prior in enumerate(priors):
fig.add_subplot(2, 5, 1 + ii)
plt.hist(prior.sample(100000), bins=100, histtype="step", density=True)
if not isinstance(prior, bilby.core.prior.Gaussian):
plt.plot(
np.linspace(prior.minimum, prior.maximum, 1000),
prior.prob(np.linspace(prior.minimum, prior.maximum, 1000)),
)
else:
plt.plot(np.linspace(-5, 5, 1000), prior.prob(np.linspace(-5, 5, 1000)))
plt.xlabel("{}".format(prior.latex_label))
plt.tight_layout()
plt.show()
plt.close()
# -
# ## Define an Analytic Prior
#
# Adding a new analytic is achieved as follows.
class Exponential(bilby.core.prior.Prior):
"""Define a new prior class where p(x) ~ exp(alpha * x)"""
def __init__(self, alpha, minimum, maximum, name=None, latex_label=None):
super(Exponential, self).__init__(
name=name, latex_label=latex_label, minimum=minimum, maximum=maximum
)
self.alpha = alpha
def rescale(self, val):
return (
np.log(
(np.exp(self.alpha * self.maximum) - np.exp(self.alpha * self.minimum))
* val
+ np.exp(self.alpha * self.minimum)
)
/ self.alpha
)
def prob(self, val):
in_prior = (val >= self.minimum) & (val <= self.maximum)
return (
self.alpha
* np.exp(self.alpha * val)
/ (np.exp(self.alpha * self.maximum) - np.exp(self.alpha * self.minimum))
* in_prior
)
# +
prior = Exponential(name="name", alpha=-1, minimum=0, maximum=10)
plt.figure(figsize=(12, 5))
plt.hist(prior.sample(100000), bins=100, histtype="step", density=True)
plt.plot(
np.linspace(prior.minimum, prior.maximum, 1000),
prior.prob(np.linspace(prior.minimum, prior.maximum, 1000)),
)
plt.xlabel("{}".format(prior.latex_label))
plt.show()
plt.close()
| examples/tutorials/making_priors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Machine Learning for Engineers: [XGBoostRegressor](https://www.apmonitor.com/pds/index.php/Main/XGBoostRegressor)
# - [XGBoost Regressor](https://www.apmonitor.com/pds/index.php/Main/XGBoostRegressor)
# - Source Blocks: 2
# - Description: Introduction to XGBoost for Regression
# - [Course Overview](https://apmonitor.com/pds)
# - [Course Schedule](https://apmonitor.com/pds/index.php/Main/CourseSchedule)
#
import xgboost as xgb
xgbc = xgb.XGBRegressor()
xgbc.fit(XA,yA)
yP = xgbc.predict(XB)
# +
import xgboost as xgb
from sklearn.datasets import make_regression
from sklearn.metrics import r2_score
from sklearn.model_selection import train_test_split
import pandas as pd
X, y = make_regression(n_samples=1000, n_features=10, n_informative=8)
Xa,Xb,ya,yb = train_test_split(X, y, test_size=0.2, shuffle=True)
xgbr = xgb.XGBRegressor()
xgbr.fit(Xa,ya)
yp = xgbr.predict(Xb)
acc = r2_score(yb,yp)
print('R2='+str(acc))
# importance_types = ['weight', 'gain', 'cover', 'total_gain', 'total_cover']
f = xgbr.get_booster().get_score(importance_type='gain')
t = pd.DataFrame(f.items(),columns=['Feature','Gain'])
print(t.sort_values('Gain',ascending=False))
# create table from feature_importances_
fi = xgbr.feature_importances_
n = ['Feature '+str(i) for i in range(10)]
d = pd.DataFrame({'Feature':n,'Importance':fi})
print(d.sort_values('Importance',ascending=False))
# plot the importance by 'weight'
xgb.plot_importance(xgbr)
| All_Source_Code/XGBoostRegressor/XGBoostRegressor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="tYKr0WV0mGT0" outputId="62298ba5-bc61-4ec1-fcda-0509462efe49"
# !ls -l ./autompg_lr.pkl
# + id="xpCVUFrpm4eP"
import pickle
# + [markdown] id="l-zGeDQknWRe"
# # load pickle with linear regression and scaler
# + colab={"base_uri": "https://localhost:8080/"} id="x3-X83cHnMJV" outputId="e933b4f0-76bb-42c6-c5ad-2907c65c0e49"
import pickle
lr = pickle.load(open('./autompg_lr.pkl', 'rb'))
type(lr)
# + colab={"base_uri": "https://localhost:8080/"} id="umnUZj6juNFn" outputId="95b89974-fa41-46fe-c340-a8161680d850"
scaler = pickle.load(open('./autompg_standardscaler.pkl','rb'))
type(scaler)
# + [markdown] id="IHX80qZh_0h3"
# # Predict with Linear Regression
# + id="J6DCdoQHun0D"
# [[307.0, 130.0, 3504.0, 12.0]]
displacement = 307.0
horsepower = 130.0
weight = 3504.0
acceleration = 12.0
x_custom = [[displacement, horsepower, weight, acceleration]]
# + colab={"base_uri": "https://localhost:8080/"} id="VI6dgxYFCAon" outputId="25916b4f-6c8a-4a5d-98bb-d9c1d3d45a22"
x_custom = scaler.transform(x_custom)
x_custom.shape
# + colab={"base_uri": "https://localhost:8080/"} id="dVksUZxeBVf2" outputId="9b52ac41-44ac-430f-e5dd-fde494d069a6"
result = lr.predict(x_custom)
result[0]
# + id="54onq6RjCeZM"
| autompg_linearregression_service.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### 1. What is DHCP used for?
# ##### Ans: It is a network protocol used to assign IP addresses.
# #### 2. What is an SSH server?
# ##### Ans: A program which responds to incoming ssh requests
# #### 3. When an ssh client is invoked at the command-line with the “ssh” command, what other argument must be provided on the command line?
# ##### Ans: A domain name or IP address to connect to
# #### 4. What command will reveal the IP address of your Raspberry Pi?
# ##### Ans: ifconfig
# #### 5. What is a fair definition of the term "protocol"?
# ##### Ans: A set of rules for communication
# #### 6. What does an IP address uniquely identify?
# ##### Ans: A host communicating on the Internet
# #### 7. How big is a port TCP/UDP number?
# ##### Ans: 16 bits
# #### 8. The nslookup command will show the IP address corresponding to a given domain name.
# ##### Ans: True
| Coursera/Interfacing with the Raspberry Pi/Week-1/Quiz/Module-1-Quiz.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Performance measurement and analysis on HPCL machines
#
# These instructions assume that you have an account on the HPCL servers, where most of the tool you require have already been installed. If you need to install TAU and other tools somewhere else, please follow the instructions in the [Getting Started](GettingStarted.ipynb) notebook.
#
# To see a list of available modules on each system, use the `module avail` command. To load all modules required for using TAU Commander (with TAU and PAPI) and Hatchet, do
#
# ```
# module load perf
# ```
# Next, you need to set up your python environment using the Conda installation in the perf module. To do that, run:
#
# ```
# conda init
# ```
#
# The changes will take effect in a new shell, so you can log out and log in again, or simply run
#
# ```
# exec bash
# ```
#
# You should see a slightly different prompt, starting with `(base)`, e.g.:
# ```
# $ exec bash
# (base) username@arya:~$
# ```
# You are now ready to use TAU Commander and Hatchet as described in the [Getting Started](GettingStarted.ipynb) notebook without having to install any tools.
| tau_analysis/HPCL_Machines.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: visualization-curriculum-gF8wUgMm
# language: python
# name: python3
# ---
# # Simple Neural Network in NumPy Experimentation Notebook
#
# - toc: true
# - badges: true
# - comments: false
# - categories: []
# - hide: true
# - search_exclude: true
# # Note
#
# This is an additional notebook for the post Simple Neural Network in NumPy. You should click on one of the badges (colab, binder, etc.) to get a kernel to run experiments.
# # Setup
# +
from enum import IntEnum
import numpy as np
import matplotlib.pyplot as plt
def linear(inp, w, b):
return inp @ w + b
def linear_backward(inp, w, b, dout):
db = dout.mean(axis=0)
dw = inp.T @ dout
dinp = dout @ w.T
return dinp, dw, db
def relu(inp):
return np.maximum(0, inp)
def relu_backward(inp, dout):
return (inp > 0) * dout
def mse(inp, true):
return np.square(inp - true).mean()
def mse_backward(inp, true):
return (inp - true) * (2 / np.prod(inp.shape))
class WeightInit(IntEnum):
simple=0
kaiming=1
class SimpleNN:
def __init__(self, input_dim, hidden_dim, out_dim, weight_init=WeightInit.simple):
self.w1, self.b1 = self._get_weights(input_dim, hidden_dim, weight_init)
self.w2, self.b2 = self._get_weights(hidden_dim, out_dim, weight_init)
def _get_weights(self, input_dim, output_dim, weight_init):
scale = 1.0
if weight_init == WeightInit.kaiming:
scale = np.sqrt(2 / input_dim)
w = np.random.normal(size=(input_dim, output_dim), scale=scale)
b = np.zeros(output_dim)
return w, b
def forward(self, inp):
self.inp = inp
self.linear1 = linear(self.inp, self.w1, self.b1)
self.relu1 = relu(self.linear1)
self.linear2 = linear(self.relu1, self.w2, self.b2)
return self.linear2
def backward(self, dlinear2):
drelu1, self.dw2, self.db2 = linear_backward(self.relu1, self.w2, self.b2, dlinear2)
dlinear1 = relu_backward(self.linear1, drelu1)
dinp, self.dw1, self.db1 = linear_backward(self.inp, self.w1, self.b1, dlinear1)
def sgd_update(self, lr):
self.w1 -= self.dw1 * lr
self.b1 -= self.db1 * lr
self.w2 -= self.dw2 * lr
self.b2 -= self.db2 * lr
def _print_mean_and_var(self, X):
y = self.forward(X)
print_stats("input", X)
print_stats("layer1", self.relu1)
print_stats("layer2", self.linear2)
def noop(x):
return x
def square(X, w, b):
return linear(np.square(X), w, b)
def wave(X, w, b):
return linear(np.sin(X), w, b)
def get_data(n, input_features, output_features, transform, x_spread=3.0, noise_spread=4.0):
w = np.random.normal(loc=0.0, scale=3.0, size=(input_features, output_features))
b = np.random.normal(loc=0.0, scale=5.0)
noise = np.random.normal(size=(n, output_features), scale=noise_spread)
X = np.random.normal(scale=x_spread, size=(n, input_features))
y = transform(X, w, b) + noise
return X, y, w, b
def get_linear_data(n, input_features, output_features):
return get_data(n, input_features, output_features, linear)
def get_square_data(n, input_features, output_features):
return get_data(n, input_features, output_features, square)
def get_wave_data(n, input_features, output_features):
return get_data(n, input_features, output_features, wave, 5.0, 0.3)
def print_stats(name, X):
print(f"After {name}: mean={X.mean():.2f}, variance={X.var():.3f}")
def print_final_loss(y_train, y_val, train_loss, val_loss):
avg_train = y_train.mean()
predict_avg_loss_train = mse(avg_train, y_train)
predict_avg_loss_val = mse(avg_train, y_val)
print(f"Train loss={train_loss:.4f}, val. loss={val_loss:.4f}")
print(f"Using avg. response: train loss={predict_avg_loss_train:.4f}, val. loss={predict_avg_loss_val:.4f}")
def plot_losses(train_losses, val_losses):
plt.figure(figsize=(16, 9))
plt.plot(range(len(train_losses)), train_losses, label="Train loss")
plt.plot(range(len(val_losses)), val_losses, label="Val. loss")
plt.yscale("log")
plt.legend()
if save_plots: plt.savefig(f"loss_for_{name}")
if show_plots: plt.show()
plt.close()
def plot_data(X, X_val, y_val, X_train, y_train, true_data_func, model, normalize_param):
X = np.sort(X, axis=0)
plt.figure(figsize=(16, 9))
plt.scatter(X_val, y_val, color="blue", label="Val. data")
plt.scatter(X_train, y_train, color="black", label="Train data")
plt.plot(X, true_data_func(X * normalize_param[1] + normalize_param[0]), color="black", label="True function")
plt.scatter(X_val, model.forward(X_val), color="orange", label="Predicted for val. data")
plt.plot(X, model.forward(X), color="orange", label="Model function")
plt.legend()
if save_plots: plt.savefig(f"data_for_{name}")
if show_plots: plt.show()
plt.close()
def run_experiment():
train_size = int(n * train_split)
val_size = n - train_size
print(f"Train size = {train_size}, validation size = {val_size}")
for (name, get_data_func, data_func) in data_types:
print(f"\n_______________\nFitting {name} data\n_______________")
X, y, true_w, true_b = get_data_func(n, inp_dim, out_dim)
true_data_func = lambda X: data_func(X, true_w, true_b)
normalize_param = (0.0, 1.0)
if normalize_input: normalize_param = (X.mean(), X.std()); X = (X - X.mean()) / X.std()
X_train, y_train = X[:train_size], y[:train_size]
X_val, y_val = X[train_size:], y[train_size:]
model = SimpleNN(inp_dim, hidden_dim, out_dim, weight_init)
if print_mean_and_var: model._print_mean_and_var(X_train)
train_losses, val_losses = [], []
for i in range(updates):
train_predicted = model.forward(X_train)
train_loss = mse(train_predicted, y_train)
model.backward(mse_backward(train_predicted, y_train))
model.sgd_update(lr * (1 - i / updates))
val_predicted = model.forward(X_val)
val_loss = mse(val_predicted, y_val)
train_losses.append(train_loss)
val_losses.append(val_loss)
if i % 200 == 0:
print(f"Update {i}/{updates}, train loss={train_loss:.4f}, val. loss={val_loss:.4f}", end=" \r")
print()
print_final_loss(y_train, y_val, train_loss, val_loss)
plot_losses(train_losses, val_losses)
if inp_dim == 1 and out_dim == 1:
plot_data(X, X_val, y_val, X_train, y_train, true_data_func, model, normalize_param)
# -
# # Config and Run Experiment
# +
data_types = [("linear", get_linear_data, linear), ("square", get_square_data, square), ("wave", get_wave_data, wave)]
np.random.seed(0)
show_plots = True
save_plots = False
print_mean_and_var = True # Print mean and variance of activations before training (variance is important for stable training)
normalize_input = True
weight_init = WeightInit.simple
n = 200 # Number of data points to generate
inp_dim = 1 # Number of features
out_dim = 1 # Number of outputs per example
train_split = 0.8 # Percentage of examples to use for training
hidden_dim = 20 # Number of neurons in the hidden layer
lr = 0.05
updates = 10000 # Number of updates to do
run_experiment()
| _notebooks/1980-01-01-snnn_exp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Author : <NAME>
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.model_selection import cross_val_score
from sklearn.tree import DecisionTreeClassifier
from sklearn import metrics
# ## 1.Let’s attempt to predict the survival of a horse based on various observed medical conditions. Load the data from ‘horses.csv’ and observe whether it contains missing values.[Hint: Pandas dataframe has a method isnull]
df = pd.read_csv(r"C:\Users\saqla\Desktop\Python Certification for Data Science\Case Studies\Module 7\Dataset\horse.csv")
df
df.isna().sum()
# ## 2.This dataset contains many categorical features, replace them with label encoding.[Hint: Refer to get_dummies methods in pandas dataframe or Label encoder in scikit-learn]
x = df.drop(columns=['hospital_number','outcome'])
y = df['outcome']
x = pd.get_dummies(x)
x
x.columns
# ## 3.Replace the missing values by the most frequent value in each column.[Hint: Refer to Imputer class in Scikit learn preprocessing module]
from sklearn.impute import SimpleImputer
imp = SimpleImputer(strategy='most_frequent')
imp.fit_transform(x)
x = pd.DataFrame(imp.fit_transform(x), columns=x.columns, index=x.index)
pd.set_option('display.max_columns',75)
x
# ## 4.Fit a decision tree classifier and observe the accuracy.
# +
x = x.drop(columns=['surgery_no','age_young','capillary_refill_time_3','capillary_refill_time_less_3_sec','surgical_lesion_no','cp_data_no'])
y = df['outcome']
# -
x_train, x_test, y_train, y_test = train_test_split(x,y, test_size = 0.3, random_state = 1)
model_tree = DecisionTreeClassifier()
model_tree.fit(x_train,y_train)
y_pred = model_tree.predict(x_test)
metrics.accuracy_score(y_test,y_pred)
# ## 5.Fit a random forest classifier and observe the accuracy.
from sklearn.ensemble import RandomForestClassifier
model_rf = RandomForestClassifier()
model_rf.fit(x_train,y_train)
rf_y_pred = model_rf.predict(x_test)
metrics.accuracy_score(y_test,rf_y_pred)
| Machine Learning/Supervised Learning/Module 7 - Case Studies (Logistic Regression, Decision Trees, Random Forest)/Module 7 - Case Study 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # 梯度下降和随机梯度下降
#
# 本节中,我们将介绍梯度下降(gradient descent)的工作原理。虽然梯度下降在深度学习中很少被直接使用,但理解梯度的意义以及沿着梯度反方向更新自变量可能降低目标函数值的原因是学习后续优化算法的基础。随后,我们将引出随机梯度下降(stochastic gradient descent)。
#
# ## 一维梯度下降
#
# 我们先以简单的一维梯度下降为例,解释梯度下降算法可能降低目标函数值的原因。假设连续可导的函数$f: \mathbb{R} \rightarrow \mathbb{R}$的输入和输出都是标量。给定绝对值足够小的数$\epsilon$,根据泰勒展开公式(参见[“数学基础”](../chapter_appendix/math.md)一节),我们得到以下的近似
#
# $$f(x + \epsilon) \approx f(x) + \epsilon f'(x) .$$
#
# 这里$f'(x)$是函数$f$在$x$处的梯度。一维函数的梯度是一个标量,也称导数。
#
# 接下来,找到一个常数$\eta > 0$,使得$\left|\eta f'(x)\right|$足够小,那么可以将$\epsilon$替换为$-\eta f'(x)$并得到
#
# $$f(x - \eta f'(x)) \approx f(x) - \eta f'(x)^2.$$
#
# 如果导数$f'(x) \neq 0$,那么$\eta f'(x)^2>0$,所以
#
# $$f(x - \eta f'(x)) \lesssim f(x).$$
#
# 这意味着,如果我们通过
#
# $$x \leftarrow x - \eta f'(x)$$
#
# 来迭代$x$,函数$f(x)$的值可能会降低。因此在梯度下降中,我们先选取一个初始值$x$和常数$\eta > 0$,然后不断通过上式来迭代$x$,直到达到停止条件,例如$f'(x)^2$的值已足够小或迭代次数已达到某个值。
#
# 下面我们以目标函数$f(x)=x^2$为例来看一看梯度下降是如何执行的。虽然我们知道最小化$f(x)$的解为$x=0$,这里我们依然使用这个简单函数来观察$x$是如何被迭代的。首先,导入本节实验所需的包或模块。
# + attributes={"classes": [], "id": "", "n": "3"}
# %matplotlib inline
import d2lzh as d2l
import math
from mxnet import nd
import numpy as np
# -
# 接下来我们使用$x=10$作为初始值,并设$\eta=0.2$。使用梯度下降对$x$迭代10次,可见最终$x$的值较接近最优解。
# + attributes={"classes": [], "id": "", "n": "4"}
def gd(eta):
x = 10
results = [x]
for i in range(10):
x -= eta * 2 * x # f(x) = x * x的导数为f'(x) = 2 * x
results.append(x)
print('epoch 10, x:', x)
return results
res = gd(0.2)
# -
# 下面将绘制出自变量$x$的迭代轨迹。
# + attributes={"classes": [], "id": "", "n": "5"}
def show_trace(res):
n = max(abs(min(res)), abs(max(res)), 10)
f_line = np.arange(-n, n, 0.1)
d2l.set_figsize()
d2l.plt.plot(f_line, [x * x for x in f_line])
d2l.plt.plot(res, [x * x for x in res], '-o')
d2l.plt.xlabel('x')
d2l.plt.ylabel('f(x)')
show_trace(res)
# -
# ## 学习率
#
# 上述梯度下降算法中的正数$\eta$通常叫做学习率。这是一个超参数,需要人工设定。如果使用过小的学习率,会导致$x$更新缓慢从而需要更多的迭代才能得到较好的解。下面展示了使用学习率$\eta=0.05$时自变量$x$的迭代轨迹。可见,同样迭代10次后,当学习率过小时,最终$x$的值依然与最优解存在较大偏差。
# + attributes={"classes": [], "id": "", "n": "6"}
show_trace(gd(0.05))
# -
# 如果使用过大的学习率,$\left|\eta f'(x)\right|$可能会过大从而使前面提到的一阶泰勒展开公式不再成立:这时我们无法保证迭代$x$会降低$f(x)$的值。举个例子,当我们设学习率$\eta=1.1$时,可以看到$x$不断越过(overshoot)最优解$x=0$并逐渐发散。
# + attributes={"classes": [], "id": "", "n": "7"}
show_trace(gd(1.1))
# -
# ## 多维梯度下降
#
# 在了解了一维梯度下降之后,我们再考虑一种更广义的情况:目标函数的输入为向量,输出为标量。假设目标函数$f: \mathbb{R}^d \rightarrow \mathbb{R}$的输入是一个$d$维向量$\boldsymbol{x} = [x_1, x_2, \ldots, x_d]^\top$。目标函数$f(\boldsymbol{x})$有关$\boldsymbol{x}$的梯度是一个由$d$个偏导数组成的向量:
#
# $$\nabla_{\boldsymbol{x}} f(\boldsymbol{x}) = \bigg[\frac{\partial f(\boldsymbol{x})}{\partial x_1}, \frac{\partial f(\boldsymbol{x})}{\partial x_2}, \ldots, \frac{\partial f(\boldsymbol{x})}{\partial x_d}\bigg]^\top.$$
#
#
# 为表示简洁,我们用$\nabla f(\boldsymbol{x})$代替$\nabla_{\boldsymbol{x}} f(\boldsymbol{x})$。梯度中每个偏导数元素$\partial f(\boldsymbol{x})/\partial x_i$代表着$f$在$\boldsymbol{x}$有关输入$x_i$的变化率。为了测量$f$沿着单位向量$\boldsymbol{u}$(即$\|\boldsymbol{u}\|=1$)方向上的变化率,在多元微积分中,我们定义$f$在$\boldsymbol{x}$上沿着$\boldsymbol{u}$方向的方向导数为
#
# $$\text{D}_{\boldsymbol{u}} f(\boldsymbol{x}) = \lim_{h \rightarrow 0} \frac{f(\boldsymbol{x} + h \boldsymbol{u}) - f(\boldsymbol{x})}{h}.$$
#
# 依据方向导数性质 \[1,14.6节定理三\],以上的方向导数可以改写为
#
# $$\text{D}_{\boldsymbol{u}} f(\boldsymbol{x}) = \nabla f(\boldsymbol{x}) \cdot \boldsymbol{u}.$$
#
# 方向导数$\text{D}_{\boldsymbol{u}} f(\boldsymbol{x})$给出了$f$在$\boldsymbol{x}$上沿着所有可能方向的变化率。为了最小化$f$,我们希望找到$f$能被降低最快的方向。因此,我们可以通过单位向量$\boldsymbol{u}$来最小化方向导数$\text{D}_{\boldsymbol{u}} f(\boldsymbol{x})$。
#
# 由于$\text{D}_{\boldsymbol{u}} f(\boldsymbol{x}) = \|\nabla f(\boldsymbol{x})\| \cdot \|\boldsymbol{u}\| \cdot \text{cos} (\theta) = \|\nabla f(\boldsymbol{x})\| \cdot \text{cos} (\theta)$,
# 其中$\theta$为梯度$\nabla f(\boldsymbol{x})$和单位向量$\boldsymbol{u}$之间的夹角,当$\theta = \pi$时,$\text{cos}(\theta)$取得最小值$-1$。因此,当$\boldsymbol{u}$在梯度方向$\nabla f(\boldsymbol{x})$的相反方向时,方向导数$\text{D}_{\boldsymbol{u}} f(\boldsymbol{x})$被最小化。所以,我们可能通过梯度下降算法来不断降低目标函数$f$的值:
#
# $$\boldsymbol{x} \leftarrow \boldsymbol{x} - \eta \nabla f(\boldsymbol{x}).$$
#
# 相同地,其中$\eta$(取正数)称作学习率。
#
# 下面我们构造一个输入为二维向量$\boldsymbol{x} = [x_1, x_2]^\top$和输出为标量的目标函数$f(\boldsymbol{x})=x_1^2+2x_2^2$。那么,梯度$\nabla f(\boldsymbol{x}) = [2x_1, 4x_2]^\top$。我们将观察梯度下降从初始位置$[-5,-2]$开始对自变量$\boldsymbol{x}$的迭代轨迹。我们先定义两个辅助函数。第一个函数使用给定的自变量更新函数,从初始位置$[-5,-2]$开始迭代自变量$\boldsymbol{x}$共20次。第二个函数将可视化自变量$\boldsymbol{x}$的迭代轨迹。
# + attributes={"classes": [], "id": "", "n": "10"}
def train_2d(trainer): # 本函数将保存在d2lzh包中方便以后使用
x1, x2, s1, s2 = -5, -2, 0, 0 # s1和s2是自变量状态,之后章节会使用
results = [(x1, x2)]
for i in range(20):
x1, x2, s1, s2 = trainer(x1, x2, s1, s2)
results.append((x1, x2))
print('epoch %d, x1 %f, x2 %f' % (i + 1, x1, x2))
return results
def show_trace_2d(f, results): # 本函数将保存在d2lzh包中方便以后使用
d2l.plt.plot(*zip(*results), '-o', color='#ff7f0e')
x1, x2 = np.meshgrid(np.arange(-5.5, 1.0, 0.1), np.arange(-3.0, 1.0, 0.1))
d2l.plt.contour(x1, x2, f(x1, x2), colors='#1f77b4')
d2l.plt.xlabel('x1')
d2l.plt.ylabel('x2')
# -
# 然后,观察学习率为$0.1$时自变量的迭代轨迹。使用梯度下降对自变量$\boldsymbol{x}$迭代20次后,可见最终$\boldsymbol{x}$的值较接近最优解$[0,0]$。
# + attributes={"classes": [], "id": "", "n": "15"}
eta = 0.1
def f_2d(x1, x2): # 目标函数
return x1 ** 2 + 2 * x2 ** 2
def gd_2d(x1, x2, s1, s2):
return (x1 - eta * 2 * x1, x2 - eta * 4 * x2, 0, 0)
show_trace_2d(f_2d, train_2d(gd_2d))
# -
# ## 随机梯度下降
#
# 在深度学习里,目标函数通常是训练数据集中有关各个样本的损失函数的平均。设$f_i(\boldsymbol{x})$是有关索引为$i$的训练数据样本的损失函数,$n$是训练数据样本数,$\boldsymbol{x}$是模型的参数向量,那么目标函数定义为
#
# $$f(\boldsymbol{x}) = \frac{1}{n} \sum_{i = 1}^n f_i(\boldsymbol{x}).$$
#
# 目标函数在$\boldsymbol{x}$处的梯度计算为
#
# $$\nabla f(\boldsymbol{x}) = \frac{1}{n} \sum_{i = 1}^n \nabla f_i(\boldsymbol{x}).$$
#
# 如果使用梯度下降,每次自变量迭代的计算开销为$\mathcal{O}(n)$,它随着$n$线性增长。因此,当训练数据样本数很大时,梯度下降每次迭代的计算开销很高。
#
# 随机梯度下降(stochastic gradient descent,简称SGD)减少了每次迭代的计算开销。在随机梯度下降的每次迭代中,我们随机均匀采样的一个样本索引$i\in\{1,\ldots,n\}$,并计算梯度$\nabla f_i(\boldsymbol{x})$来迭代$\boldsymbol{x}$:
#
# $$\boldsymbol{x} \leftarrow \boldsymbol{x} - \eta \nabla f_i(\boldsymbol{x}).$$
#
# 这里$\eta$同样是学习率。可以看到每次迭代的计算开销从梯度下降的$\mathcal{O}(n)$降到了常数$\mathcal{O}(1)$。值得强调的是,随机梯度$\nabla f_i(\boldsymbol{x})$是对梯度$\nabla f(\boldsymbol{x})$的无偏估计:
#
# $$\mathbb{E}_i \nabla f_i(\boldsymbol{x}) = \frac{1}{n} \sum_{i = 1}^n \nabla f_i(\boldsymbol{x}) = \nabla f(\boldsymbol{x}).$$
#
# 这意味着,平均来说,随机梯度是对梯度的一个良好的估计。
#
# 下面我们通过在梯度中添加均值为0的随机噪音来模拟随机梯度下降,以此来比较它与梯度下降的区别。
# + attributes={"classes": [], "id": "", "n": "17"}
def sgd_2d(x1, x2, s1, s2):
return (x1 - eta * (2 * x1 + np.random.normal(0.1)),
x2 - eta * (4 * x2 + np.random.normal(0.1)), 0, 0)
show_trace_2d(f_2d, train_2d(sgd_2d))
# -
# 可以看到,随机梯度下降中自变量的迭代轨迹相对于梯度下降中的来说更为曲折。这是由于实验所添加的噪音使得模拟的随机梯度的准确度下降。在实际中,这些噪音通常来自于训练数据集中的各个样本。
#
#
# ## 小结
#
# * 使用适当的学习率,沿着梯度反方向更新自变量可能降低目标函数值。梯度下降重复这一更新过程直到得到满足要求的解。
# * 学习率过大过小都有问题。一个合适的学习率通常是需要通过多次实验找到的。
# * 当训练数据集的样本较多时,梯度下降每次迭代计算开销较大,因而随机梯度下降通常更受青睐。
#
#
# ## 练习
#
# * 使用一个不同的目标函数,观察梯度下降和随机梯度下降中自变量的迭代轨迹。
# * 在二维梯度下降的实验中尝试使用不同的学习率,观察并分析实验现象。
#
#
# ## 扫码直达[讨论区](https://discuss.gluon.ai/t/topic/1877)
#
# 
| chapter_optimization/gd-sgd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.6 (''torch110'': venv)'
# language: python
# name: python3
# ---
# # Network Pruning
#
# Network pruning is a commonly-used technique to speed up your model during inference. We will talk about this topic in this tutorial.
#
# ## Basic concept
# As we all know, the majority of the runtime is attributed to the generic matrix multiply (a.k.a. GEMM) operations. So naturally, the problem comes out that whether we can speed up the operation by reducing the number of the elements in the matrices. By setting the weights, biases and the corresponding input and output items to 0, we can then just skip those calculations.
#
# There are generally two kinds of pruning, structured pruning and unstructured pruning. For structured pruning, the weight connections are removed in groups. e.g. The entire channel is deleted. It has the effect of changing the input and output shapes of layers and the weight matrices. Because of this, nearly every system can benefit from it. Unstructured pruning, on the other hand, removes individual weight connections from a network by setting them to 0. So, it is highly dependent on the inference backends.
#
# Currently, only structured pruning is supported in TinyNeuralNetwork.
#
# ### How structured pruning is implemented in DNN frameworks?
# ```py
# model = Net(pretrained=True)
# sparsity = 0.5
#
# masks = {None: None}
#
# def register_masks(layer):
# parent_layer = get_parent(layer)
# input_mask = masks[parent_layer]
# if is_passthrough_layer(layer):
# output_mask = input_mask
# else:
# output_mask = get_mask(layer, sparsity)
# register_mask(layer, input_mask, output_mask)
# masks[layer] = output_mask
#
# model.apply(register_masks)
# model.fit(train_data)
#
# def apply_masks(layer):
# parent_layer = get_parent(layer)
# input_mask = masks[parent_layer]
# output_mask = masks[layer]
# apply_mask(layer, input_mask, output_mask)
#
# model.apply(apply_masks)
# ```
#
# ### Network Pruning in TinyNerualNetwork
# The problem in the previous code example is that only one parent layer is expected. But in some recent DNN models, there are a few complicated operations like `cat`, `add` and `split`. We need to resolve the dependencies of those operations as well.
#
# To solve the aforementioned problem, first we go through some basic definitions. When the input shape and output shape of a node are not related during pruning, it is called a node with isolation. For example, the `conv`, `linear` and `lstm` nodes are nodes with isolation. We want to find out a group of nodes, which is called a subgraph, that starts with and ends with nodes with isolation and doesn't contain a subgraph in it. We use the nodes with isolation for finding out the candidate subgraphs in the model.
#
# ```py
# def find_subgraph(layer, input_modify, output_modify, nodes):
# if layer in nodes:
# return None
#
# nodes.append(layer)
#
# if is_layer_with_isolation(layer):
# if input_modify:
# for prev_layer in get_parent(layer):
# return find_subgraph(prev_layer, False, True, nodes)
# if output_modify:
# for next_layer in get_child(layer):
# return find_subgraph(next_layer, True, False, nodes)
# else:
# for prev_layer in get_parent(layer):
# return find_subgraph(prev_layer, input_modify, output_modify, nodes)
# for next_layer in get_child(layer):
# return find_subgraph(next_layer, input_modify, output_modify, nodes)
#
# candidate_subgraphs = []
#
# def construct_candidate_subgraphs(layer):
# if is_layer_with_isolation(layer):
# nodes = []
# find_subgraph(layer, True, False, nodes)
# candidate_subgraphs.append(nodes)
#
# nodes = []
# find_subgraph(layer, False, True, nodes)
# candidate_subgraphs.append(nodes)
#
# model.apply(construct_subgraphs)
# ```
#
# With all candidate subgraphs, the next step we do is to remove the duplicated and invalid ones in them. Due to space limitations, we will not cover this section in detail. When we get the final subgraphs, the first node in it is called the center node. During configuration, we use the name of the center node to represent the subgraph it constructs. Some properties can be set at the subgraph level by the user, like sparsity.
#
# Although we have the subgraphs, the mapping of channels between nodes is still unknown. So we need to resolve channel dependency. Similarly, we pass the channel information recursively so as to get the correct mapping at each node. It may be a bit more complicated since each node has its own logic for sharing channel mapping. Operations like `add` require shared mapping in all the input and output tensors, while `cat` allows the inputs to have independent mappings, however the output mapping and the combined input mapping is shared. As this is too detailed, we will not expand on it.
#
# After resolving the channel dependency, we follow the ordinary pruning process, that is to register the masks of the weight and bias tensors. And then you may just finetune the model. When the training process is finished, then it is time to apply the masks, so that the model actually gets smaller. Alternatively, you may apply the masks just after registering them if the masks won't change during training. As a result, the training process will be significantly faster. That's all the story for pruning.
#
# ### Using the pruner in TinyNeuralNetwork
# It is really simple to use the pruner in our framework. You can use the code below.
#
# +
import sys
sys.path.append('../..')
import torch
import torchvision
from tinynn.prune.oneshot_pruner import OneShotChannelPruner
model = torchvision.models.mobilenet_v2(pretrained=True)
model.train()
dummy_input = torch.randn(1, 3, 224, 224)
pruner = OneShotChannelPruner(model, dummy_input, config={'sparsity': 0.25, 'metrics': 'l2_norm'})
st_flops = pruner.calc_flops()
pruner.prune()
ed_flops = pruner.calc_flops()
print(f"Pruning over, reduced FLOPS {100 * (st_flops - ed_flops) / st_flops:.2f}% ({st_flops} -> {ed_flops})")
# You should start finetuning the model here
| tutorials/pruning/basic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Predict Happiness Source
# - Importing the Packages
# +
# importing packages
import pandas as pd
import numpy as np # For mathematical calculations
import seaborn as sns # For data visualization
import matplotlib.pyplot as plt # For plotting graphs
# %matplotlib inline
import warnings # To ignore any warnings warnings.filterwarnings("ignore")
import seaborn as sns
from sklearn.preprocessing import MinMaxScaler
#from sklearn.cross_validation import train_test_split
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import f1_score
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
from nltk.stem.porter import PorterStemmer
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import cross_val_score
import nltk
import re
import codecs
import seaborn as sns
from sklearn.preprocessing import LabelEncoder, OneHotEncoder
#from sklearn.cross_validation import train_test_split
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import f1_score
# -
import pandas as pd
import numpy as np
#import xgboost as xgb
from tqdm import tqdm
from sklearn.svm import SVC
from keras.models import Sequential
from keras.layers.recurrent import LSTM, GRU
from keras.layers.core import Dense, Activation, Dropout
from keras.layers.embeddings import Embedding
from keras.layers.normalization import BatchNormalization
from keras.utils import np_utils
from sklearn import preprocessing, decomposition, model_selection, metrics, pipeline
from sklearn.model_selection import GridSearchCV
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
from sklearn.decomposition import TruncatedSVD
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn.naive_bayes import MultinomialNB
from keras.layers import GlobalMaxPooling1D, Conv1D, MaxPooling1D, Flatten, Bidirectional, SpatialDropout1D
from keras.preprocessing import sequence, text
from keras.callbacks import EarlyStopping
from nltk import word_tokenize
from nltk.corpus import stopwords
stop_words = stopwords.words('english')
# - Reading the data
train=pd.read_csv('hm_train.csv')
test=pd.read_csv('hm_test.csv')
submission=pd.read_csv('sample_submission.csv')
print(train.columns,test.columns,submission.columns)
print(train.head(),test.head(),submission.head())
print(train.shape,test.shape,submission.shape)
# - Let’s make a copy of train and test data so that even if we have to make any changes in these datasets we would not lose the original datasets.
#
#
train_copy=pd.read_csv('hm_train.csv').copy()
test_copy=pd.read_csv('hm_test.csv').copy()
submission_copy=pd.read_csv('sample_submission.csv').copy()
# # Univariate Analysis
print(train.dtypes,test.dtypes,submission.dtypes)
train['predicted_category'].value_counts(normalize=True)
# Read as percentage after multiplying by 100
train['predicted_category'].value_counts(normalize=True).plot.bar()
# - exercise featurescontributes approx 2%, nature contributes 3.5 %, leisure contributes 7.5%, enjoy_the_moment does 10%, bonding does 10%, achievement does 34%, and affection does34.5 % to the population sample.
# On Looking at the datsets, we identified that there are 3 Data Types:
#
# - Continuous : reflection_period, cleaned_hm ,num_sentence
#
# - Categorical :Category
#
# - Text : cleaned_hm
# Let's go for Continuous Data Type Exploration. We know that we use BarGraphs for Categorical variable, Histogram or ScatterPlot for continuous variables.
# reflection_period
plt.figure(figsize=(6, 6))
sns.countplot(train["reflection_period"])
plt.title('reflection_period')
plt.show()
# num_sentence
plt.figure(figsize=(10, 6))
sns.countplot(train["num_sentence"])
plt.title('num_sentence')
plt.show()
# predicted_category
plt.figure(figsize=(13, 6))
sns.countplot(train["predicted_category"])
plt.title('predicted_category')
plt.show()
# Pairplot for cross visualisation of continuous variables
plt.figure(figsize=(30,30))
sns.pairplot(train, diag_kind='kde');
# Data Preprocessing
# - Checking Missing Values
def missing_value(df):
total = df.isnull().sum().sort_values(ascending = False)
percent = (df.isnull().sum()/df.isnull().count()*100).sort_values(ascending=False)
missing_df = pd.concat([total, percent], axis=1, keys = ['Total', 'Percent'])
return missing_df
mis_train = missing_value(train)
mis_train
mis_test = missing_value(test)
mis_test
# Text Preprocessing
# - Let's try to understand the writing style if possible :)
grouped_df = train.groupby('predicted_category')
for name, group in grouped_df:
print("Text : ", name)
cnt = 0
for ind, row in group.iterrows():
print(row["cleaned_hm"])
cnt += 1
if cnt == 5:
break
print("\n")
# Though, there are not very special Characters but some are present like expression of emoji's i.e. :), vs, etc
# ### Feature Engineering:
# Now let us come try to do some feature engineering. This consists of two main parts.
#
# - Meta features - features that are extracted from the text like number of words, number of stop words, number of punctuations etc
# - Text based features - features directly based on the text / words like frequency, svd, word2vec etc.
#
# #### Meta Features:
# We will start with creating meta featues and see how good are they at predicting the happiness source. The feature list is as follows:
#
# - Number of words in the text
# - Number of unique words in the text
# - Number of characters in the text
# - Number of stopwords
# - Number of punctuations
# - Number of upper case words
# - Number of title case words
# - Average length of the words
#
# Feature Engineering function
import string
def feature_engineering(df):
## Number of words in the text ##
df["num_words"] = df['cleaned_hm'].apply(lambda x: len(str(x).split()))
## Number of unique words in the text ##
df["num_unique_words"] = df['cleaned_hm'].apply(lambda x: len(set(str(x).split())))
## Number of characters in the text ##
df["num_chars"] = df['cleaned_hm'].apply(lambda x: len(str(x)))
## Number of stopwords in the text ##
from nltk.corpus import stopwords
eng_stopwords = stopwords.words('english')
df["num_stopwords"] = df['cleaned_hm'].apply(lambda x: len([w for w in str(x).lower().split() if w in eng_stopwords]))
## Number of punctuations in the text ##
df["num_punctuations"] =df['cleaned_hm'].apply(lambda x: len([c for c in str(x) if c in string.punctuation]))
## Number of title case words in the text ##
df["num_words_upper"] = df['cleaned_hm'].apply(lambda x: len([w for w in str(x).split() if w.isupper()]))
## Number of title case words in the text ##
df["num_words_title"] = df['cleaned_hm'].apply(lambda x: len([w for w in str(x).split() if w.istitle()]))
## Average length of the words in the text ##
df["mean_word_len"] = df['cleaned_hm'].apply(lambda x: np.mean([len(w) for w in str(x).split()]))
df = pd.concat([df, df["num_words"],df["num_unique_words"],df["num_chars"],df["num_stopwords"],df["num_punctuations"],
df["num_words_upper"],df["num_words_title"],df["mean_word_len"]],axis=1)
#X = dataset.loc[:,['Transaction-Type','Complaint-reason','Company-response','Consumer-disputes','delayed','converted_text','convertion_language']]
return df
train = feature_engineering(train)
test = feature_engineering(test)
train.head()
# Let us now plot some of our new variables to see of they will be helpful in predictions.
# CLEANING THE TEXT !!!
def preprocess_text(df):
""" Here, The function preprocesses the text. It performs stemming, lemmatization
, stopwords removal, common words and rare words removal and removal of unwanted chractaers. """
#removing non-letter symbols and converting text in 'cleaned_hm' to lowercase
df = df.apply(lambda x: "".join(re.sub(r"[^A-Za-z\s]", '',str(x))))
# lower casing the Text
df = df.apply(lambda x: " ".join(x.lower() for x in x.split()))
#Removing punctuations
#adding characters list which needs to remove that is PUNCTUATION
punc = ['.', ',', '"', "'", '?','#', '!', ':','vs',':)', ';', '(', ')', '[', ']', '{', '}',"%",'/','<','>','br','�','^','XX','XXXX','xxxx','xx']
#removing extra characters
df = df.apply(lambda x: " ".join(x for x in x.split() if x not in punc))
import nltk
nltk.download('stopwords')
#removal of stopwords
from nltk.corpus import stopwords
stop = stopwords.words('english')
df = df.apply(lambda x: " ".join(x for x in x.split() if x not in stop))
#common words removal
freq_df = pd.Series(' '.join(df).split()).value_counts()[:10]
freq_df = list(freq_df.index)
df = df.apply(lambda x: " ".join(x for x in x.split() if x not in freq_df))
#rare words removal
freq_df_rare = pd.Series(' '.join(df).split()).value_counts()[-10:]
freq_df_rare = list(freq_df_rare.index)
df = df.apply(lambda x: " ".join(x for x in x.split() if x not in freq_df_rare))
#STEMMING
st = PorterStemmer()
df=df.apply(lambda x: " ".join([st.stem(w) for w in x.split()]))
# WordNet lexical database for lemmatization
#from nltk.stem import WordNetLemmatizer
#lem = WordNetLemmatizer()
#df=df.apply(lambda x: " ".join([lem.lemmatize(w) for w in x.split()]))
return df
train['cleaned_hm'] = preprocess_text(train['cleaned_hm'])
test['cleaned_hm'] = preprocess_text(test['cleaned_hm'])
# Removing 'h or m extension
train['reflection_period'] = train['reflection_period'].str.rstrip('h | m')
test['reflection_period'] = test['reflection_period'].str.rstrip('h | m')
train['reflection_period'].fillna(train['reflection_period'].mode()[0], inplace=True)
train['cleaned_hm'].fillna(train['cleaned_hm'].mode()[0], inplace=True)
train['num_sentence'].fillna(train['num_sentence'].mode()[0], inplace=True)
train['predicted_category'].fillna(train['predicted_category'].mode()[0], inplace=True)
test['reflection_period'].fillna(test['reflection_period'].mode()[0], inplace=True)
test['cleaned_hm'].fillna(test['cleaned_hm'].mode()[0], inplace=True)
test['num_sentence'].fillna(test['num_sentence'].mode()[0], inplace=True)
#NOw We will convert the target variable into LabelEncoder
y_train = train.loc[:,['predicted_category']]
labelencoder1 = LabelEncoder()
labelencoder1.fit(y_train.values)
y_train=labelencoder1.transform(y_train)
train.columns
# +
x = train.loc[:,["num_sentence","reflection_period",'num_sentence',
'num_words', 'num_unique_words', 'num_chars',
'num_stopwords', 'num_punctuations', 'num_words_upper',
'num_words_title', 'mean_word_len', 'num_words', 'num_unique_words',
'num_chars', 'num_stopwords', 'num_punctuations', 'num_words_upper',
'num_words_title', 'mean_word_len']]
x1 = test.loc[:,["num_sentence","reflection_period",'num_sentence',
'num_words', 'num_unique_words', 'num_chars',
'num_stopwords', 'num_punctuations', 'num_words_upper',
'num_words_title', 'mean_word_len', 'num_words', 'num_unique_words',
'num_chars', 'num_stopwords', 'num_punctuations', 'num_words_upper',
'num_words_title', 'mean_word_len']]
# -
from sklearn.pipeline import Pipeline
from sklearn.feature_extraction.text import TfidfTransformer
import gensim
# +
from gensim.models import Word2Vec
wv = gensim.models.KeyedVectors.load_word2vec_format("GoogleNews-vectors-negative300.bin.gz",limit=50000, binary=True)
wv.init_sims(replace=True)
# -
from itertools import islice
list(islice(wv.vocab, 13030, 13050))
# +
def word_averaging(wv, words):
all_words, mean = set(), []
for word in words:
if isinstance(word, np.ndarray):
mean.append(word)
elif word in wv.vocab:
mean.append(wv.syn0norm[wv.vocab[word].index])
all_words.add(wv.vocab[word].index)
if not mean:
logging.warning("cannot compute similarity with no input %s", words)
# FIXME: remove these examples in pre-processing
return np.zeros(wv.vector_size,)
mean = gensim.matutils.unitvec(np.array(mean).mean(axis=0)).astype(np.float32)
return mean
def word_averaging_list(wv, text_list):
return np.vstack([word_averaging(wv, post) for post in text_list ])
# -
def w2v_tokenize_text(text):
tokens = []
for sent in nltk.sent_tokenize(text, language='english'):
for word in nltk.word_tokenize(sent, language='english'):
if len(word) < 2:
continue
tokens.append(word)
return tokens
nltk.download('punkt')
alldata = pd.concat([train, test], axis=1)
# initialise the functions - we'll create separate models for each type.
countvec = CountVectorizer(analyzer='word', ngram_range = (1,2), max_features=500)
tfidfvec = TfidfVectorizer(analyzer='word', ngram_range = (1,2), max_features=500)
# create features
bagofwords = countvec.fit_transform(alldata['cleaned_hm'])
tfidfdata = tfidfvec.fit_transform(alldata['cleaned_hm'])
# create dataframe for features
bow_df = pd.DataFrame(bagofwords.todense())
tfidf_df = pd.DataFrame(tfidfdata.todense())
# set column names
bow_df.columns = ['col'+ str(x) for x in bow_df.columns]
tfidf_df.columns = ['col' + str(x) for x in tfidf_df.columns]
# +
# create separate data frame for bag of words and tf-idf
bow_df_train = bow_df[:len(train)]
bow_df_test = bow_df[len(train):]
tfid_df_train = tfidf_df[:len(train)]
tfid_df_test = tfidf_df[len(train):]
# -
# split the merged data file into train and test respectively
train_feats = alldata[~pd.isnull(alldata.predicted_category)]
test_feats = alldata[pd.isnull(alldata.predicted_category)]
# +
# merge count (bag of word) features into train
x_train = pd.concat([x, bow_df_train], axis = 1)
x_test = pd.concat([x1, bow_df_test], axis=1)
x_test.reset_index(drop=True, inplace=True)
# -
from sklearn.linear_model import LogisticRegression
logreg = LogisticRegression(n_jobs=-1, penalty = 'l1',C=1.0,random_state = 0)
logreg = logreg.fit(x_train, train['cleaned_hm'])
y_pred = logreg.predict(x_test)
ans = labelencoder1.inverse_transform(y_pred)
type(ans)
ans = pd.DataFrame(ans)
id1=test.loc[:,['hmid']]
final_ans = [id1, ans]
final_ans = pd.concat(final_ans, axis=1)
final_ans.columns = ['hmid', 'predicted_category']
final_ans.to_csv('vdemo_HK.csv',index=False)
| Final_Submission_Hackerearth - Copy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Power of Two
# + active=""
# Given an integer, write a function to determine if it is a power of two.
# Example 1:
# Input: 1
# Output: true
# Explanation: 2^0 = 1
#
# Example 2:
# Input: 16
# Output: true
# Explanation: 2^4 = 16
#
# Example 3:
# Input: 218
# Output: false
# -
class Solution:
def isPowerOfTwo(self, n): # 99.83
"""
:type n: int
:rtype: bool
"""
# n(>0)負值為False, &:位元運算子, 比較二進位!! 2^n -> 10..0 2^n-1 -> 01..1, (2^n & 2^n-1) -> 00..0
return n > 0 and not (n & n-1) # not 轉 bool
n = 2
ans = Solution()
ans.isPowerOfTwo(n)
| 231. Power of Two.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Solves the 1D Wave equation with MDM and position Verlet algorithm
import numpy as np
from core import *
import matplotlib.pyplot as plt
from matplotlib import animation, rcParams
from IPython.display import HTML
rcParams['animation.html'] = 'html5'
def animate_wave(solution):
fig = plt.figure(figsize=(8,6))
ax = plt.gca()
ax.set_xlim((0, 1))
ax.set_ylim((-1.5, 1.5))
ax.set_xlabel('x')
ax.set_ylabel('u(x)')
plt.grid(True)
line, = ax.plot([], [], lw=2)
plt.close()
def animate(i):
line.set_data(xgrid, solution[:, i])
ax.set_title('1D Wave Equation\nt = {:01.2f}'.format(i * dt))
return line,
return animation.FuncAnimation(fig, animate, frames=solution.shape[1], interval=50, blit=True)
# +
# Spatial discretization
# Order of accuracy (spatial)
k = 2
# Number of cells
m = 50
# Left boundary
a = 0
# Right boundary
b = 1
# Step length
dx = (b - a) / m
verlet = False # If verlet = 0 then it uses 4th order accurate for time (FR)
# 1D Staggered grid
xgrid = np.append(np.insert(np.arange(a+dx/2, b, dx), 0, a), b)
# Mimetic operator (Laplacian)
L = lap1D(k, m, dx)
# Wave propagation speed
c = 2. # (T/p) Tension over density
# "Force" function
F = lambda x : (c**2) * L * x # c^2 DivGrad x
# Simulation time
TIME = 1.
# Temporal discretization based on CFL condition
dt = dx / (2 * c) # dt = h on Young's paper
# Initial condition
ICU = lambda x : np.sin(np.pi * x) # Initial position of particles
ICV = lambda x : np.zeros(m+1) # Initial velocity of particles
uold = ICU(xgrid)
vold = ICV(xgrid)
vold = np.append(vold, vold[-1])
theta = 1. / (2. - 2.**(1./3.)) # From <NAME>'s paper
Nt = int(TIME/dt)
solution = np.zeros((uold.shape[0], Nt))
# Time integration loop
for t in range(0, Nt):
# Apply "position Verlet" algorithm (2nd-order in time)
if verlet:
uold = uold + 0.5 * dt * vold
vnew = vold + dt * F(uold)
unew = uold + 0.5 * dt * vnew
# Apply "Forest-Ruth" algorithm (4th-order in time)
else:
unew = uold + theta * 0.5 * dt * vold
vnew = vold + theta * dt * F(unew)
unew = unew + (1 - theta) * 0.5 * dt * vnew
vnew = vnew + (1 - 2*theta) * dt * F(unew)
unew = unew + (1 - theta) * 0.5 * dt * vnew
vnew = vnew + theta * dt * F(unew)
unew = unew + theta * 0.5 * dt * vnew
uold = unew
vold = vnew
solution[:, t] = unew
# -
anim = animate_wave(solution)
anim
| wave1D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: transforming-autoencoders
# language: python
# name: transforming-autoencoders
# ---
# Change to project root
import os
os.chdir('..')
# Autoreload
# %load_ext autoreload
# %autoreload 2
import tensorflow as tf
import numpy as np
min_trans = -4
max_trans = 4
parallel_calls = 8
# # Load the data
# +
from transforming_autoencoders import mnist, data
mnist_data = mnist.MNIST('data')
def train_input_fn(batch_size):
return mnist_data.train_input_fn()(batch_size,
lambda:data.translation_fn(batch_size, min_trans,max_trans),
parallel_calls)
def eval_input_fn(batch_size):
return mnist_data.eval_input_fn()(batch_size,
lambda:data.translation_fn(batch_size, min_trans,max_trans),
parallel_calls)
# -
# # Create the estimator
from transforming_autoencoders import trans_autoencoder_estimator
model = trans_autoencoder_estimator.create_transforming_autoencoder(model_dir='model/translation-2',
num_capsules=30,
num_rec=50,
num_gen=50,
trans_size=2,
trans_fn='translation')
# # Train the model
batch_size = 64
epochs = 6
steps = int(mnist_data.train_size() * epochs / batch_size)
model.train(input_fn=lambda:train_input_fn(batch_size), steps=steps)
# # Evaluate the model
test_data = eval_input_fn(4)
with tf.Session() as sess:
img, trans, expec = sess.run([test_data[0]['image'],
test_data[0]['transformation'],
test_data[1]])
predict_input_fn = tf.estimator.inputs.numpy_input_fn(
x={"image": img, "transformation": trans},
num_epochs=1,
shuffle=False)
yhat = list(model.predict(input_fn=predict_input_fn))
pred = np.array(list(map(lambda x: x['out_image'], yhat)))
from transforming_autoencoders import plot
plot.plot_some(img)
plot.plot_some(expec)
plot.plot_some(pred)
print(trans)
| nbs/MNIST-translation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Convolutions
#
# In this notebook, we explore the concept of convolutional neural networks.
#
#
# You may want to read this [wikipedia page](https://en.wikipedia.org/wiki/Convolution) if you're not familiar with the concept of a convolution.
#
# In a convolutional neural network
# ### 1. Definition of the (discrete) convolution
#
# You may read Wikipedia's [web page](https://en.wikipedia.org/wiki/Convolution#Discrete_convolution)
#
# I we consider two functions $f$ and $g$ taking values from $\mathbb{Z} \to \mathbb{R}$ then:
# $ (f * g)[n] = \sum_{m = -\infty}^{+\infty} f[m] \cdot g[n - m] $
#
# In our case, we consider the two vectors $x$ and $w$ :
# $ x = (x_1, x_2, ..., x_{n-1}, x_n) $
# $ w = (w_1, w_2) $
#
# And get :
# $ x * w = (w_1 x_1 + w_2 x_2, w_1 x_2 + w_2 x_3, ..., w_1 x_{n-1} + w_2 x_n)$
#
#
# #### Deep learning subtility :
#
# In most of deep learning framewoks, you'll get to chose in between three paddings:
# - **Same**: $(f*g)$ has the same shape as x (we pad the entry with zeros)
# - **valid**: $(f*g)$ has the shape of x minus the shape of w plus 1 (no padding on x)
# - **Causal**: $(f*g)(n_t)$ does not depend on any $(n_{t+1})$
# +
# Which is easily implemented on python :
def _convolve(x, w, type='valid'):
# x and w are np vectors
conv = []
for i in range(len(x)):
if type == 'valid':
conv.append((x[i: i+len(w)] * w).sum())
return np.array(conv)
def convolve(X, w):
# Convolves a batch X to w
w = np.array(w)
X = np.array(X)
conv = []
for i in range(len(X)):
conv.append(_convolve(X[i], w))
return np.array(conv)
# -
# ### 2. Derive the Convolution !!
#
# As we use it, the convolution is parametrised by two vectors $x$ and $w$ and outputs a vector $z$. We have:
# $ x * w = z$
# $ z_i = (w_1 x_i + w_2 x_{i+1})$
#
# We want to derive $z$ with respect to some weights $w_j$:
# $\frac{\delta z_i}{\delta w_j} = x_{i+j}$
# $\frac{\delta z_i}{\delta w} = (x_{i}, x_{i+1}, ..., x_{i+n})$
#
# ### Example of convolutions :
#
# We consider a classification problem where we want to distinguish 2 signals. One is going upward and the other is going downwards
# +
from utils import *
import utils
reload(utils)
from utils import *
(x_train, y_train), (x_test, y_test) = load_up_down(50)
plt.plot(x_train.T)
plt.show()
# -
# ### Train a convolutional neural net
# +
# Rename y_silver to X and y_gold to Y
X, Y = [x_train, ], y_train
# Initilize the parameters
Ws = [0.5, 0.5]
alphas = (0.01, 0.01)
# Load Trainer
t = Trainer(X, Y, Ws, alphas)
# Define Prediction and Loss
t.pred = lambda X : convolve(X[0], (t.Ws[0], t.Ws[1])).mean(axis=1)
t.loss = lambda : (np.power((t.Y - t.pred(t.X)), 2) * 1 / 2.).mean()
print t.pred(X)
t.acc = lambda X, Y : t.pred(X)
# Define the gradient functions
dl_dp = lambda : -(t.Y - t.pred(X))
dl_dw0 = lambda : (t.X[0][:-1]).mean()
dl_dw1 = lambda : (t.X[0][1:]).mean()
t.dWs = (dl_dw0, dl_dw1)
# Start training
anim = t.animated_train(is_notebook=True)
from IPython.display import HTML
HTML(anim.to_html5_video())
# -
t.loss()
# ### Applied to image
#
# We demonstrate how 2D convolutions applies to images (In this case, we designe the kernels of the conlutions).
# +
from scipy import signal
# Load MNIST
(x_train, y_train), (x_test, y_test) = load_MNIST()
img = x_train[2]
# Design the kernels
kernels = [[[-1, 2, -1],[-1, 2, -1],[-1, 2, -1]],
[[-1, -1, -1],[2, 2, 2],[-1, -1, -1]],
[[2, -1, -1],[-1, 2, -1],[-1, -1, 2]],
[[-1, -1, 2],[-1, 2, -1],[2, -1, -1]], ]
# Plot and convolve them to the image
for i, k in enumerate(kernels):
i = i*2+1
plt.subplot(3,4,i)
plt.imshow(k, cmap='gray')
plt.subplot(3,4,i+1)
conv = signal.convolve2d(img, k)
plt.imshow(conv > 1.5, cmap='gray')
plt.subplot(349)
plt.imshow(img, cmap='gray')
plt.show()
| notebooks/Convolution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sklearn import datasets
import matplotlib.pyplot as plt
from sklearn.decomposition import PCA
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report,confusion_matrix
from sklearn import ensemble
import time
lfw=datasets.fetch_lfw_people(min_faces_per_person=100,resize=0.4)
lfw.keys()
lfw.data.shape
fig=plt.figure(figsize=(8,8))
for i in range(64):
ax=fig.add_subplot(8,8,i+1)
ax.imshow(lfw.images[i],cmap=plt.cm.bone)
plt.show()
x,y=lfw.data,lfw.target
x_train,x_test,y_train,y_test=train_test_split(x,y,random_state=0)
pca_oliv=PCA()
pca_oliv.fit(x_train)
k=0
total=0
while total<0.99:
total=total+pca_oliv.explained_variance_ratio_[k]
k=k+1
k
pca_lfw=PCA(n_components=k,whiten=True)
x_transformed=pca_lfw.fit_transform(x_train)
x_transformed.shape
x_approx=pca_lfw.inverse_transform(x_transformed)
x_approx=x_approx.reshape(855,50,37)
fig=plt.figure(figsize=(8,8))
for i in range(64):
ax=fig.add_subplot(8,8,i+1)
ax.imshow(x_approx[i],cmap=plt.cm.bone)
plt.show()
pca_lfw.components_.shape
eigenv=pca_lfw.components_.reshape(k,50,37)
fig=plt.figure(figsize=(8,8))
for i in range(64):
ax=fig.add_subplot(8,8,i+1,xticks=[],yticks=[])
ax.imshow(eigenv[i],cmap=plt.cm.bone)
plt.show()
x_train_pca=x_transformed
x_test_pca=pca_lfw.transform(x_test)
rf=ensemble.RandomForestClassifier()
start=time.time()
rf.fit(x_train,y_train)
print(time.time()-start)
y_pred=rf.predict(x_test)
print(classification_report(y_test,y_pred))
rf=ensemble.RandomForestClassifier()
start=time.time()
rf.fit(x_train_pca,y_train)
print(time.time()-start)
y_pred=rf.predict(x_test_pca)
print(classification_report(y_test,y_pred))
print(confusion_matrix(y_test,y_pred))
| Lecture 20 PCA-2/Classification of LFW Images/Classification of LFW Images-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### *Assessment of pathways using the IAMC 1.5°C Scenario Data*
#
# <img style="float: right; height: 60px; padding-left: 20px;" src="../_static/IIASA_logo.png">
# <img style="float: right; height: 60px;" src="../_static/iamc-logo.png">
#
# # CO2 emissions in the Sustainable Development Scenario (IEA-ETP 2020)
#
# This notebook illustrates the CO2 emissions in the **Sustainable Development Scenario** from the 2020 IEA Energy Technology Perspectives report ([press release](https://www.iea.org/news/reaching-energy-and-climate-goals-demands-a-dramatic-scaling-up-of-clean-energy-technologies-starting-now)) compared to the IAMC 1.5°C Scenario Ensemble compiled for the IPCC's _"Special Report on Global Warming of 1.5°C"_.
#
# The scenario data used in this analysis can be accessed and downloaded at [https://data.ene.iiasa.ac.at/iamc-1.5c-explorer](https://data.ene.iiasa.ac.at/iamc-1.5c-explorer).
# ## Load ``pyam`` package and other dependences
import pandas as pd
import numpy as np
import io
import itertools
import yaml
import math
import matplotlib.pyplot as plt
plt.style.use('../assessment/style_sr15.mplstyle')
# %matplotlib inline
import pyam
# ## Import scenario data, categorization and specifications files
#
# The metadata file must be generated from the notebook `sr15_2.0_categories_indicators`
# in the repository https://github.com/iiasa/ipcc_sr15_scenario_analysis.
# If the snapshot file has been updated, make sure that you rerun the categorization notebook.
#
# The last cell of this section loads and assigns a number of auxiliary lists as defined in the categorization notebook.
sr1p5 = pyam.IamDataFrame(data='../data/iamc15_scenario_data_world_r2.0.xlsx')
sr1p5.load_meta('../assessment/sr15_metadata_indicators.xlsx')
# +
with open("../assessment/sr15_specs.yaml", 'r') as stream:
specs = yaml.load(stream, Loader=yaml.FullLoader)
rc = pyam.run_control()
for item in specs.pop('run_control').items():
rc.update({item[0]: item[1]})
cats = specs.pop('cats')
cats_15 = specs.pop('cats_15')
cats_15_no_lo = specs.pop('cats_15_no_lo')
# -
# ## Downselect scenario ensemble
years = range(2010, 2101, 10)
df = (
sr1p5
.filter(category='Above 2C', keep=False)
.filter(variable='Emissions|CO2', year=years)
)
df.convert_unit('Mt CO2/yr', 'Gt CO2/yr', inplace=True)
# ## Append IEA ETP 2020 SDS scenario data
#
# Data collected from this chart: https://www.iea.org/data-and-statistics/charts/global-energy-sector-co2-emissions-by-sector-in-the-sustainable-development-scenario-2019-2070
_sds = pd.DataFrame([33.4, 27.4, 16.7, 9.4, 3.8, 0], columns=['year'], index=range(2020, 2071, 10)).T
df.append(_sds, model='IEA ETP 2020', scenario='SDS',
variable='Emissions|CO2', unit='Gt CO2/yr', region='year', inplace=True)
# ## Plot the IEA ETP 2020 CO2 emissions
# +
fig, ax = plt.subplots()
_cats = df.filter(category=['1.5C low overshoot', 'Lower 2C'])
_cats.line_plot(ax=ax, color='category', linewidth=0, fill_between=True, final_ranges=True, legend=False)
ax.axhline(y=0, xmin=0.09, xmax=0.83, color='black', linewidth=0.8)
etp_2020 = df.filter(model='IEA ETP 2020')
etp_2020.line_plot(ax=ax, color='black', linewidth=1.5, linestyle='--', legend=False)
etp_2017 = df.filter(model='IEA Energy Technology Perspective Model 2017')
etp_2017.line_plot(ax=ax, color='black', linewidth=1.5, linestyle=':', legend=False)
ax.set_title('CO2 emissions in IEA ETP (2020/2017) scenarios relative to IAMC 1.5°C scenarios')
# -
| further_analysis/iamc15_iea_etp_sds_2020.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # The NetworkX Module
# NetworkX is a python module. To start exploring NetworkX we simply need to start a python session (Like the IPython session you are in now!), and type
import networkx
# All of NetworkX's data structures and functions can then be accessed using the syntax `networkx.[Object]`, where `[Object]` is the function or data structure you need. Of course you would replace `[Object]` with the function you wanted. For example to make a graph, we'd write:
G = networkx.Graph()
# Usually to save ourselves some keystrokes, we'll import NetworkX using a shorter variable name
import networkx as nx
# # Basic Graph Data Structures
# One of the main strengths of NetworkX is its flexible graph data structures. There are four data structures
# - `Graph`: Undirected Graphs
# - `DiGraph`: Directed Graphs
# - `MultiGraph`: Undirected multigraphs, ie graphs which allow for multiple edges between nodes
# - `MultiDiGraph`: Directed Multigraphs
#
# Each of these has the same basic structure, attributes and features, with a few minor differences.
# # Creating Graphs
# Creating Graphs is as simple as calling the appropriate constructor.
G = nx.Graph()
D = nx.DiGraph()
M = nx.MultiGraph()
MD = nx.MultiDiGraph()
# You can also add attributes to a graph during creation, either by providing a dictionary, or simply using keyword arguments
G = nx.Graph(DateCreated='2015-01-10',name="Terry")
G.graph
# The graph attribute is just a dictionary and can be treated as one, so you can add and delete more information from it.
G.graph['Current']=False
del G.graph['name']
G.graph
# ## Nodes
# Next we'll cover how to add and remove nodes, as well as check for their existance in a graph and add attributes to both!
# ### Adding Nodes
# There are two main functions for adding nodes. `add_node`, and `add_nodes_from`. The former takes single values, and the latter takes any iterable (list, set, iterator, generator). Nodes can be of any _immutable_ type. This means numbers (ints and floats complex), strings, bytes, tuples or frozen sets. They cannot be _mutable_, such as lists, dictionaries or sets. Nodes in the same graph do not have to be of the same type
# Adding single nodes of various types
G.add_node(0)
G.add_node('A')
G.add_node(('x',1.2))
# Adding collections of nodes
G.add_nodes_from([2,4,6,8,10])
G.add_nodes_from(set([10+(3*i)%5 for i in range(10,50)]))
# ### Listing Nodes
# Accessing nodes is done using the `nodes` function which is a member of the `Graph` object.
G.nodes()
# Sometimes to save memory we might only want to access a list of nodes one at a time, so we can use an _iterator_. These are especially useful in long running loops to save memory.
for n in G.nodes_iter():
if type(n)== str:
print(n + ' is a string!')
else:
print(str(n) + " is not a string!")
# In the future more functions of NetworkX will exclusively use iterators to save memory and be more Python 3 like...
# ### Checking whether nodes are in a Graph
# We can also check to see if a graph has a node several different ways. The easiest is just using the `in` keyword in python, but there is also the `has_node` function.
13 in G
9 in G
G.has_node(13)
G.has_node(9)
# ### Node attributes
# You can also add attributes to nodes. This can be handy for storing information about nodes within the graph object. This can be done when you create new nodes using keyword arguments to the `add_node` and `add_nodes_from` function
G.add_node('Spam',company='Hormel',food='meat')
# When using `add_nodes_from` you provide a tuple with the first element being the node, and the second being a dictionary of attributes for that node. You can also add attributes which will be applied to all added nodes using keyword arguments
G.add_nodes_from([('Bologna',{'company':'<NAME>'}),
('Bacon',{'company':'Wright'}),
('Sausage',{'company':'<NAME>'})],food='meat')
# To list node attributes you need to provide the `data=True` keyword to the `nodes` and `nodes_iter` functions
G.nodes(data=True)
# Attributes are stored in a special dictionary within the graph called `node` you can access, edit and remove attributes there
G.node['Spam']
G.node['Spam']['Delicious'] = True
G.node[6]['integer'] = True
G.nodes(data=True)
del G.node[6]['integer']
G.nodes(data=True)
# Similiarly, you can remove nodes with the `remove_node` and `remove_nodes_from` functions
G.remove_node(14)
G.remove_nodes_from([10,11,12,13])
G.nodes()
# ### Exercises
# #### Repeated Nodes
# 1. What happens when you add nodes to a graph that already exist?
# 2. What happens when you add nodes to the graph that already exist but have new attributes?
# 3. What happens when you add nodes to a graph with attributes different from existing nodes?
# 4. Try removing a node that doesn't exist, what happens?
# #### The FizzBuzz Graph
# Using the spaces provided below make a new graph, `FizzBuzz`. Add nodes labeled 0 to 100 to the graph. Each node should have an attribute 'fizz' and 'buzz'. If the nodes label is divisble by 3 `fizz=True` if it is divisble by 5 `buzz=True`, otherwise both are false.
# ## Edges
# Adding edges is similar to adding nodes. They can be added, using either `add_edge` or `add_edges_from`. They can also have attributes in the same way nodes can. If you add an edge that includes a node that doesn't exist it will create it for you
G.add_edge('Bacon','Sausage',breakfast=True)
G.add_edge('Ham','Bacon',breakfast=True)
G.add_edge('Spam','Eggs',breakfast=True)
# Here we are using a list comprehension. This is an easy way to construct lists using a single line. Learn more about list comprehensions [here](https://docs.python.org/2/tutorial/datastructures.html#list-comprehensions).
G.add_edges_from([(i,i+2) for i in range(2,8,2)])
G.edges()
G.edges(data=True)
# Removing edges is accomplished by using the `remove_edge` or `remove_edges_from` function. Remove edge attributes can be done by indexing into the graph
G['Spam']['Eggs']
del G['Spam']['Eggs']['breakfast']
G.remove_edge(2,4)
G.edges(data=True)
# You can check for the existance of edges with `has_edge`
G.has_edge(2,4)
G.has_edge('Ham','Bacon')
# For directed graphs, ordering matters. `add_edge(u,v)` will add an edge from `u` to `v`
D.add_nodes_from(range(10))
D.add_edges_from([(i,i+1 % 10) for i in range(0,10)])
D.edges()
D.has_edge(0,1)
D.has_edge(1,0)
# You can also access edges for only a subset of nodes by passing edges a collection of nodes
D.edges([3,4,5])
# ### Exercises
# For the `FizzBuzz` graph above, add edges betweeen two nodes `u` and `v` if they are both divisible by 2 or by 7. Each edge should include attributes `div2` and `div7` which are true if `u` and `v` are divisible by 2 and 7 respecitively. Exclude self loops.
# ## Multigraphs
# Multigraphs can have multiple edges between any two nodes. They are referenced by a key.
M.add_edge(0,1)
M.add_edge(0,1)
M.edges()
# The keys of the edges can be accessed by using the keyword `keys=True`. This will give a tuple of `(u,v,k)`, with the edge being `u` and `v` and the key being `k`.
M.edges(keys=True)
# `MultiDraphs` and `MultiDiGraphs` are similar to `Graphs` and `DiGraphs` in most respects
# ## Adding Graph Motifs
# In addition to adding nodes and edges one at a time networkx has some convenient functions for adding complete subgraphs. But beware, these may be removed, or the API changed in the future.
G.add_cycle(range(100,110))
G.edges()
# # Basic Graph Properties
# Basic graph properties are functions which are member of the `Graph` class itself. We'll explore different metrics in part III.
# ## Node and Edge Counts
# The _order_ of a graph is the number of nodes, it can be accessed by calling `G.order()` or using the builtin length function: `len(G)`.
G.order()
len(G)
# The number of edges is usually referred to as the _size_ of the graph, and can be accessed by `G.size()`. You could also find out by calling `len(G.edges())`, but this is much slower.
G.size()
# For multigraphs it counts the number of edges includeing multiplicity
M.size()
# ## Node Neighbors
# Node neighbors can be accessed via the `neighbors`
G.neighbors('Bacon')
# In the case of directed graphs, neighbors are only those originating at the node.
D.add_edges_from([(0,i) for i in range(5,10)])
D.neighbors(0)
# For multigraphs, neighbors are only reported once.
M.neighbors(0)
# ## Degree
# The degree of a graph can be found using the `degree` function for undirected graphs, and `in_degree` and `out_degree` for directed graphs. They both return a dictionary with the node as the keys of the dictionary and the degree as the value
G.degree()
D.in_degree()
D.out_degree()
# Both of these can be called on a single node or a subset of nodes if not all degrees are needed
D.in_degree(5)
D.out_degree([0,1,2])
# You can also calculate weighted degree. To do this each edge has to have specific attribute to be used as a weight.
WG = nx.Graph()
WG.add_star(range(5))
WG.add_star(range(5,10))
WG.add_edges_from([(i,2*i %10) for i in range(10)])
for (u,v) in WG.edges_iter():
WG[u][v]['product'] = (u+1)*(v+1)
WG.degree(weight='product')
# # Exercises
# ## Create A Classroom Graph
# Let's make a network of the people in this room. First, create a graph called `C`. Everyone state their name (one at a time) and where they are from. Add nodes to the graph representing each individual, with an attribute denoting where they are from. Add edges to the graph between an individual and their closest three classmates. Have each edge have an attribute that indicates whether there was a previous relationship between the two. If none existed have `relationship=None`, if it does exist have the relationship stated, e.g. `relationship='Cousin-in-law'`
# How many nodes are in the Graph? How many Edges? What is the degree of the graph?
# # Quickly Saving a Graph
# In the next section we'll learn more about saving and loading graphs, as well as operations on graphs, but for now just run the code below.
nx.write_gpickle(C,'./data/Classroom.pickle')
| I. The Graph Data Structures.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Test MetaCNN
# In this notebook we show how to use the flat CNN meta-model on the task active enhancers vs active promoters.
from meta_models.meta_models import CNN1DMetaModel
from keras_bed_sequence import BedSequence
from keras_mixed_sequence import MixedSequence
from ucsc_genomes_downloader import Genome
from crr_labels import fantom
hg38 = Genome("hg38")
model = CNN1DMetaModel()
| Test MetaCNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 - AzureML
# language: python
# name: python3-azureml
# ---
# +
#Connect to AML workspace
from azureml.core import Workspace
ws = Workspace.from_config()
# +
#Select AML Compute Cluster
cpu_cluster_name = 'amlcluster'
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
# Verify that cluster does not exist already
try:
cpu_cluster = ComputeTarget(workspace=ws, name=cpu_cluster_name)
print('Found an existing cluster, using it instead.')
except ComputeTargetException:
#When creating a new cluster, use the identity_type argument to attach a system- or user-assigned managed identity
compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_D13_V2',
min_nodes=0,
max_nodes=10,
identity_type='SystemAssigned')
cpu_cluster = ComputeTarget.create(ws, cpu_cluster_name, compute_config)
cpu_cluster.wait_for_completion(show_output=True)
# -
#Create Experiment
from azureml.core import Experiment
experiment = Experiment(ws, 'managed-identity-test')
# +
from azureml.core.runconfig import RunConfiguration
from azureml.core.conda_dependencies import CondaDependencies
from azureml.core.runconfig import DEFAULT_CPU_IMAGE
from azureml.pipeline.core import Pipeline
from azureml.pipeline.steps import PythonScriptStep
from azureml.pipeline.core import PipelineParameter, PipelineData
from azureml.data import OutputFileDatasetConfig
# create a new runconfig object
test_kv_run_config = RunConfiguration()
# enable Docker
test_kv_run_config.environment.docker.enabled = True
# set Docker base image to the default CPU-based image
test_kv_run_config.environment.docker.base_image = DEFAULT_CPU_IMAGE
# use conda_dependencies.yml to create a conda environment in the Docker image for execution
test_kv_run_config.environment.python.user_managed_dependencies = False
test_kv_run_config.environment.python.conda_dependencies.add_pip_package('azure-keyvault')
# -
#Create PythonScriptStep to gather data from remote source and register as AML dataset
test_kv_access_step = PythonScriptStep(
script_name="test_kv_access.py",
compute_target=cpu_cluster,
source_directory='.',
allow_reuse=False,
runconfig=test_kv_run_config
)
pipeline = Pipeline(workspace=ws, steps=[test_kv_access_step])
run = experiment.submit(pipeline)
run.wait_for_completion(show_output=True)
| KV_Access_Sample_Pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Matching Market - testing Parameter NL production decline
# + [markdown] slideshow={"slide_type": "-"}
# This simple model consists of a buyer, a supplier, and a market.
#
# The buyer represents a group of customers whose willingness to pay for a single unit of the good is captured by a vector of prices _wta_. You can initiate the buyer with a set_quantity function which randomly assigns the willingness to pay according to your specifications. You may ask for these willingness to pay quantities with a _getbid_ function.
#
# The supplier is similar, but instead the supplier is willing to be paid to sell a unit of technology. The supplier for instance may have non-zero variable costs that make them unwilling to produce the good unless they receive a specified price. Similarly the supplier has a get_ask function which returns a list of desired prices.
#
# The willingness to pay or sell are set randomly using uniform random distributions. The resultant lists of bids are effectively a demand curve. Likewise the list of asks is effectively a supply curve. A more complex determination of bids and asks is possible, for instance using time of year to vary the quantities being demanded.
#
# ## New in version 20
# - fixed bug in clearing mechanism, included a logic check to avoid wierd behavior around zero
#
# ## Microeconomic Foundations
#
# The market assumes the presence of an auctioneer which will create a _book_, which seeks to match the bids and the asks as much as possible. If the auctioneer is neutral, then it is incentive compatible for the buyer and the supplier to truthfully announce their bids and asks. The auctioneer will find a single price which clears as much of the market as possible. Clearing the market means that as many willing swaps happens as possible. You may ask the market object at what price the market clears with the get_clearing_price function. You may also ask the market how many units were exchanged with the get_units_cleared function.
#
# ## Paramsweep
#
# Within this version we test a scenario where the Netherlands choose to decrease their production, irrigardless of market conditions. For example due to safety concerns in the production area.
# -
# ## Agent-Based Objects
#
# The following section presents three objects which can be used to make an agent-based model of an efficient, two-sided market.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import random as rnd
import pandas as pd
import numpy as np
import time
import datetime
import calendar
import json
import statistics
# fix what is missing with the datetime/time/calendar package
def add_months(sourcedate,months):
month = sourcedate.month - 1 + months
year = int(sourcedate.year + month / 12 )
month = month % 12 + 1
day = min(sourcedate.day,calendar.monthrange(year, month)[1])
return datetime.date(year,month,day)
# measure how long it takes to run the script
startit = time.time()
dtstartit = datetime.datetime.now()
# -
# ## classes buyers and sellers
# Below we are constructing the buyers and sellers in classes.
# +
class Seller():
def __init__(self, name):
self.name = name
self.wta = []
self.step = 0
self.prod = 2000
self.lb_price = 10
self.lb_multiplier = 0
self.ub_price = 20
self.ub_multiplier = 0
self.init_reserve = 500000
self.reserve = 500000
self.init_unproven_reserve = 0
self.unproven_reserve = 0
#multiple market idea, also 'go away from market'
self.subscr_market = {}
self.last_price = 15
self.state_hist = {}
self.cur_scenario = ''
self.count = 0
self.storage = 0
self.q_to_market = 0
self.ratio_sold = 0
self.ratio_sold_hist = []
# the supplier has n quantities that they can sell
# they may be willing to sell this quantity anywhere from a lower price of l
# to a higher price of u
def set_quantity(self):
self.count = 0
self.update_price()
n = self.prod
l = self.lb_price + self.lb_multiplier
u = self.ub_price + self.ub_multiplier
wta = []
for i in range(n):
p = rnd.uniform(l, u)
wta.append(p)
if len(wta) < self.reserve:
self.wta = wta
else:
self.wta = wta[0:(self.reserve-1)]
self.prod = self.reserve
if len(self.wta) > 0:
self.wta = self.wta #sorted(self.wta, reverse=False)
self.q_to_market = len(self.wta)
def get_name(self):
return self.name
def get_asks(self):
return self.wta
def extract(self, cur_extraction):
if self.reserve > 0:
self.reserve = self.reserve - cur_extraction
else:
self.prod = 0
# production costs rise a 100%
def update_price(self):
depletion = (self.init_reserve - self.reserve) / self.init_reserve
self.ub_multiplier = int(self.ub_price * depletion)
self.lb_multiplier = int(self.lb_price * depletion)
def return_not_cleared(self, not_cleared):
self.count = self.count + (len(self.wta) - len(not_cleared))
self.wta = not_cleared
def get_price(self, price):
self.last_price = price
def update_production(self):
if (self.step/12).is_integer():
if self.prod > 0 and self.q_to_market > 0:
rp_ratio = self.reserve / self.prod
self.ratio_sold = self.count / self.q_to_market
self.ratio_sold_hist.append(self.ratio_sold)
yearly_average = statistics.mean(self.ratio_sold_hist[-12:])
if self.name != 'Netherlands':
if (rp_ratio > 15) and (yearly_average > .9):
self.prod = int(self.prod * 1.1)
if print_details:
print("%s evaluate production" % self.name)
else:
self.prod = int(self.prod * .9)
if (self.unproven_reserve > 0) and (self.cur_scenario == 'PACES'):
self.reserve = self.reserve + int(0.1 * self.init_unproven_reserve)
self.unproven_reserve = self.unproven_reserve - int(0.1 * self.init_unproven_reserve)
def evaluate_timestep(self):
self.update_production()
# record every step into an dictionary, nog pythonic look into (vars)
def book_keeping(self):
self.state_hist[self.step] = self.__dict__
class Buyer():
def __init__(self, name):
self.name = name
self.type = 0
self.rof = 0
self.wtp = []
self.step = 0
self.offset= 0
self.base_demand = 0
self.max_demand = 0
self.lb_price = 10
self.ub_price = 20
self.last_price = 15
self.subscr_market = {}
self.state_hist = {}
self.cur_scenario = ''
self.count = 0
self.real_demand = 0
self.storage_cap = 1
self.storage = 0
self.storage_q = 0
# the supplier has n quantities that they can buy
# they may be willing to sell this quantity anywhere from a lower price of l
# to a higher price of u
def set_quantity(self):
self.count = 0
self.update_price()
n = int(self.consumption(self.step))
l = self.lb_price
u = self.ub_price
wtp = []
for i in range(n):
p = rnd.uniform(l, u)
wtp.append(p)
self.wtp = wtp #sorted(wtp, reverse=True)
# gets a little to obvious
def get_name(self):
return self.name
# return list of willingness to pay
def get_bids(self):
return self.wtp
def consumption(self, x):
# make it initialise to seller
b = self.base_demand
m = self.max_demand
y = b + m * (.5 * (1 + np.cos(((x+self.offset)/6)*np.pi)))
self.real_demand = y
s = self.storage_manager()
return(y+s)
def update_price(self):
# adjust Q
if self.type == 1: #home
if (self.step/12).is_integer():
self.base_demand = home_savings[self.cur_scenario] * self.base_demand
self.max_demand = home_savings[self.cur_scenario] * self.max_demand
if self.type == 2: # elec for eu + us
if (self.step/12).is_integer():
cur_elec_df = elec_space['RELATIVE'][self.cur_scenario]
period_now = add_months(period_null, self.step)
index_year = int(period_now.strftime('%Y'))
#change_in_demand = cur_elec_df[index_year]
self.base_demand = self.base_demand * cur_elec_df[index_year]
self.max_demand = self.max_demand * cur_elec_df[index_year]
if self.type == 3: #indu
if (self.step/12).is_integer():
if (self.rof == 0) and (self.cur_scenario == 'PACES'):
#cur_df = economic_growth['ECONOMIC GROWTH'][self.cur_scenario]
period_now = add_months(period_null, self.step)
index_year = int(period_now.strftime('%Y'))
#growth = cur_df[index_year]
growth = np.arctan((index_year-2013)/10)/(.5*np.pi)*.05+0.03
self.base_demand = (1 + growth) * self.base_demand
self.max_demand = (1 + growth) * self.max_demand
else:
cur_df = economic_growth['ECONOMIC GROWTH'][self.cur_scenario]
period_now = add_months(period_null, self.step)
index_year = int(period_now.strftime('%Y'))
growth = cur_df[index_year]
self.base_demand = (1 + growth) * self.base_demand
self.max_demand = (1 + growth) * self.max_demand
## adjust P now to get_price, but adress later
## moved to get_price, rename update_price function (?)
#self.lb_price = self.last_price * .75
#self.ub_price= self.last_price * 1.25
def return_not_cleared(self, not_cleared):
self.count = self.count + (len(self.wtp)-len(not_cleared))
self.wtp = not_cleared
def get_price(self, price):
self.last_price = price
if self.last_price > 100:
self.last_price = 100
self.lb_price = self.last_price * .75
self.ub_price= self.last_price * 1.25
# writes complete state to a dictionary, see if usefull
def book_keeping(self):
self.state_hist[self.step] = self.__dict__
# there has to be some accountability for uncleared bids of the buyers
# integrate get_price in here somehow
def evaluate_timestep(self):
if self.type==1:
not_cleared = len(self.wtp)
#total_demand = self.real_demand + self.storage_q
storage_delta = self.storage_q - not_cleared
self.storage = self.storage + storage_delta
if print_details:
print(self.name, storage_delta)
def storage_manager(self):
# check if buyer is household buyer
if self.type==1:
if self.storage < 0:
self.storage_q = -self.storage
else:
self.storage_q = 0
return(self.storage_q)
else:
return(0)
# -
# ## Construct the market
# For the market two classes are made. The market itself, which controls the buyers and the sellers, and the book. The market has a book where the results of the clearing procedure are stored.
# +
# the book is an object of the market used for the clearing procedure
class Book():
def __init__(self):
self.ledger = pd.DataFrame(columns = ("role","name","price","cleared"))
def set_asks(self,seller_list):
# ask each seller their name
# ask each seller their willingness
# for each willingness append the data frame
for seller in seller_list:
seller_name = seller.get_name()
seller_price = seller.get_asks()
ar_role = np.full((1,len(seller_price)),'seller', dtype=object)
ar_name = np.full((1,len(seller_price)),seller_name, dtype=object)
ar_cleared = np.full((1,len(seller_price)),'in process', dtype=object)
temp_ledger = pd.DataFrame([*ar_role,*ar_name,seller_price,*ar_cleared]).T
temp_ledger.columns= ["role","name","price","cleared"]
self.ledger = self.ledger.append(temp_ledger, ignore_index=True)
def set_bids(self,buyer_list):
# ask each seller their name
# ask each seller their willingness
# for each willingness append the data frame
for buyer in buyer_list:
buyer_name = buyer.get_name()
buyer_price = buyer.get_bids()
ar_role = np.full((1,len(buyer_price)),'buyer', dtype=object)
ar_name = np.full((1,len(buyer_price)),buyer_name, dtype=object)
ar_cleared = np.full((1,len(buyer_price)),'in process', dtype=object)
temp_ledger = pd.DataFrame([*ar_role,*ar_name,buyer_price,*ar_cleared]).T
temp_ledger.columns= ["role","name","price","cleared"]
self.ledger = self.ledger.append(temp_ledger, ignore_index=True)
def update_ledger(self,ledger):
self.ledger = ledger
def get_ledger(self):
return self.ledger
def clean_ledger(self):
self.ledger = pd.DataFrame(columns = ("role","name","price","cleared"))
class Market():
def __init__(self, name):
self.name= name
self.count = 0
self.last_price = ''
self.book = Book()
self.b = []
self.s = []
self.buyer_list = []
self.seller_list = []
self.buyer_dict = {}
self.seller_dict = {}
self.ledger = ''
self.seller_analytics = {}
self.buyer_analytics = {}
def book_keeping_all(self):
for i in self.buyer_dict:
self.buyer_dict[i].book_keeping()
for i in self.seller_dict:
self.seller_dict[i].book_keeping()
def add_buyer(self,buyer):
if buyer.subscr_market[self.name] == 1:
self.buyer_list.append(buyer)
def add_seller(self,seller):
if seller.subscr_market[self.name] == 1:
self.seller_list.append(seller)
def set_book(self):
self.book.set_bids(self.buyer_list)
self.book.set_asks(self.seller_list)
def get_bids(self):
# this is a data frame
ledger = self.book.get_ledger()
rows= ledger.loc[ledger['role'] == 'buyer']
# this is a series
prices=rows['price']
# this is a list
bids = prices.tolist()
return bids
def get_asks(self):
# this is a data frame
ledger = self.book.get_ledger()
rows = ledger.loc[ledger['role'] == 'seller']
# this is a series
prices=rows['price']
# this is a list
asks = prices.tolist()
return asks
# return the price at which the market clears
# this fails because there are more buyers then sellers
def get_clearing_price(self):
# buyer makes a bid starting with the buyer which wants it most
b = self.get_bids()
s = self.get_asks()
# highest to lowest
self.b=sorted(b, reverse=True)
# lowest to highest
self.s=sorted(s, reverse=False)
# find out whether there are more buyers or sellers
# then drop the excess buyers or sellers; they won't compete
n = len(b)
m = len(s)
# there are more sellers than buyers
# drop off the highest priced sellers
if (m > n):
s = s[0:n]
matcher = n
# There are more buyers than sellers
# drop off the lowest bidding buyers
else:
b = b[0:m]
matcher = m
# -It's possible that not all items sold actually clear the market here
# -Produces an error when one of the two lists are empty
# something like 'can't compare string and float'
count = 0
for i in range(matcher):
if (self.b[i] > self.s[i]):
count +=1
self.last_price = self.b[i]
# copy count to market object
self.count = count
return self.last_price
# TODO: Annotate the ledger
# this procedure takes up 80% of processing time
def annotate_ledger(self,clearing_price):
ledger = self.book.get_ledger()
# logic test
# b or s can not be zero, probably error or unreliable results
# so annote everything as false in that case and move on
b = self.get_bids()
s = self.get_asks()
if (len(s)==0 or len(b)==0):
new_col = [ 'False' for i in range(len(ledger['cleared']))]
ledger['cleared'] = new_col
self.book.update_ledger(ledger)
return
# end logic test
for index, row in ledger.iterrows():
if (row['role'] == 'seller'):
if (row['price'] < clearing_price):
ledger.loc[index,'cleared'] = 'True'
else:
ledger.loc[index,'cleared'] = 'False'
else:
if (row['price'] > clearing_price):
ledger.loc[index,'cleared'] = 'True'
else:
ledger.loc[index,'cleared'] = 'False'
self.book.update_ledger(ledger)
def get_units_cleared(self):
return self.count
def clean_ledger(self):
self.ledger = ''
self.book.clean_ledger()
def run_it(self):
self.pre_clearing_operation()
self.clearing_operation()
self.after_clearing_operation()
# pre clearing empty out the last run and start
# clean ledger is kind of sloppy, rewrite functions to overide the ledger
def pre_clearing_operation(self):
self.clean_ledger()
def clearing_operation(self):
self.set_book()
clearing_price = self.get_clearing_price()
if print_details:
print(self.name, clearing_price)
self.annotate_ledger(clearing_price)
def after_clearing_operation(self):
for agent in self.seller_list:
name = agent.name
cur_extract = len(self.book.ledger[(self.book.ledger['cleared'] == 'True') &
(self.book.ledger['name'] == name)])
agent.extract(cur_extract)
agent.get_price(self.last_price)
self.seller_analytics[name] = cur_extract
if cur_extract >0:
agent_asks = agent.get_asks()
agent_asks = sorted(agent_asks, reverse=False)
not_cleared = agent_asks[cur_extract:len(agent_asks)]
agent.return_not_cleared(not_cleared)
for agent in self.buyer_list:
name = agent.name
cur_extract = len(self.book.ledger[(self.book.ledger['cleared'] == 'True') &
(self.book.ledger['name'] == name)])
agent.get_price(self.last_price)
self.buyer_analytics[name] = cur_extract
if cur_extract >0:
agent_bids = agent.get_bids()
agent_bids = sorted(agent_bids, reverse=True)
not_cleared = agent_bids[cur_extract:len(agent_bids)]
agent.return_not_cleared(not_cleared)
# cleaning up the books
self.book_keeping_all()
# -
# ## Observer
# The observer holds the clock and collects data. In this setup it tells the market another tick has past and it is time to act. The market will instruct the other agents. The observer initializes the model, thereby making real objects out of the classes defined above.
class Observer():
def __init__(self, init_buyer, init_seller, timesteps, scenario):
self.init_buyer = init_buyer
self.init_seller = init_seller
self.init_market = init_market
self.maxrun = timesteps
self.cur_scenario = scenario
self.buyer_dict = {}
self.seller_dict = {}
self.market_dict = {}
self.timetick = 0
self.gas_market = ''
self.market_hist = []
self.seller_hist = []
self.buyer_hist = []
self.market_origin = []
self.market_origin_df = pd.DataFrame(columns=['seller_analytics','buyer_analytics'])
self.all_data = {}
def set_buyer(self, buyer_info):
for name in buyer_info:
self.buyer_dict[name] = Buyer('%s' % name)
self.buyer_dict[name].base_demand = buyer_info[name]['offset']
self.buyer_dict[name].base_demand = buyer_info[name]['b']
self.buyer_dict[name].max_demand = buyer_info[name]['m']
self.buyer_dict[name].lb_price = buyer_info[name]['lb_price']
self.buyer_dict[name].ub_price = buyer_info[name]['ub_price']
self.buyer_dict[name].type = buyer_info[name]['type']
self.buyer_dict[name].rof = buyer_info[name]['rof']
self.buyer_dict[name].cur_scenario = self.cur_scenario
self.buyer_dict[name].subscr_market = dict.fromkeys(init_market,0)
for market in buyer_info[name]['market']:
self.buyer_dict[name].subscr_market[market] = 1
def set_seller(self, seller_info):
for name in seller_info:
self.seller_dict[name] = Seller('%s' % name)
self.seller_dict[name].prod = seller_info[name]['prod']
self.seller_dict[name].lb_price = seller_info[name]['lb_price']
self.seller_dict[name].ub_price = seller_info[name]['ub_price']
self.seller_dict[name].reserve = seller_info[name]['reserve']
self.seller_dict[name].init_reserve = seller_info[name]['reserve']
self.seller_dict[name].unproven_reserve = seller_info[name]['UP_reserve']
self.seller_dict[name].init_unproven_reserve = seller_info[name]['UP_reserve']
#self.seller_dict[name].rof = seller_info[name]['rof']
self.seller_dict[name].cur_scenario = self.cur_scenario
self.seller_dict[name].subscr_market = dict.fromkeys(init_market,0)
for market in seller_info[name]['market']:
self.seller_dict[name].subscr_market[market] = 1
def set_market(self, market_info):
for name in market_info:
self.market_dict[name] = Market('%s' % name)
#add suplliers and buyers to this market
for supplier in self.seller_dict.values():
self.market_dict[name].add_seller(supplier)
for buyer in self.buyer_dict.values():
self.market_dict[name].add_buyer(buyer)
self.market_dict[name].seller_dict = self.seller_dict
self.market_dict[name].buyer_dict = self.buyer_dict
def update_buyer(self):
for i in self.buyer_dict:
self.buyer_dict[i].step += 1
self.buyer_dict[i].set_quantity()
def update_seller(self):
for i in self.seller_dict:
self.seller_dict[i].step += 1
self.seller_dict[i].set_quantity()
def evaluate_timestep(self):
for i in self.buyer_dict:
self.buyer_dict[i].evaluate_timestep()
for i in self.seller_dict:
self.seller_dict[i].evaluate_timestep()
def get_reserve(self):
reserve = []
for name in self.seller_dict:
reserve.append(self.seller_dict[name].reserve)
return reserve
def get_data(self):
for name in self.seller_dict:
self.all_data[name] = self.seller_dict[name].state_hist
for name in self.buyer_dict:
self.all_data[name] = self.buyer_dict[name].state_hist
def run_it(self):
# Timing
# time initialising
startit_init = time.time()
# initialise, setting up all the agents (firstrun not really needed anymore, since outside the loop)
# might become useful again if run_it is used for parametersweep
first_run = True
if first_run:
self.set_buyer(self.init_buyer)
self.set_seller(self.init_seller)
self.set_market(self.init_market)
first_run=False
# time init stop
stopit_init = time.time() - startit_init
if print_details:
print('%s : initialisation time' % stopit_init)
# building the multiindex for origin dataframe
listing = []
for m in self.market_dict:
listing_buyer = [(runname, m,'buyer_analytics',v.name) for v in self.market_dict[m].buyer_list]
listing = listing + listing_buyer
listing_seller = [(runname, m,'seller_analytics',v.name) for v in self.market_dict[m].seller_list]
listing = listing + listing_seller
multi_listing = pd.MultiIndex.from_tuples(listing)
# recording everything in dataframes, more dependable than lists?
#reserve_df = pd.DataFrame(data=None, columns=[i for i in self.seller_dict])
#iterables = [[i for i in self.market_dict], ['buyer_analytics', 'seller_analytics']]
#index = pd.MultiIndex.from_product(iterables)
market_origin_df = pd.DataFrame(data=None, columns=multi_listing)
for period in range(self.maxrun):
# time the period
startit_period = time.time()
self.timetick += 1
period_now = add_months(period_null, self.timetick-1)
if print_details:
print('#######################################')
print(period_now.strftime('%Y-%b'), self.cur_scenario)
# update the buyers and sellers (timetick+ set Q)
self.update_buyer()
self.update_seller()
# real action on the market
for market in self.market_dict:
if market != 'lng':
self.market_dict[market].run_it()
self.market_dict['lng'].run_it()
#tell buyers timetick has past
self.evaluate_timestep()
# data collection
for name in self.market_dict:
p_clearing = self.market_dict[name].last_price
q_sold = self.market_dict[name].count
self.market_hist.append([period_now.strftime('%Y-%b'), p_clearing, q_sold, name])
for name in self.seller_dict:
reserve = self.seller_dict[name].reserve
produced = self.seller_dict[name].count
self.seller_hist.append([period_now.strftime('%Y-%b'), reserve, produced, name])
for name in self.buyer_dict:
storage = self.buyer_dict[name].storage
consumed = self.buyer_dict[name].count
self.buyer_hist.append([period_now.strftime('%Y-%b'), storage, consumed, name])
# means to caption the origin of stuff sold on the market,
# but since dictionaries are declared global of some sort
# Dataframe has to be used to capture the real values
for name in self.market_dict:
seller_analytics = self.market_dict[name].seller_analytics
buyer_analytics = self.market_dict[name].buyer_analytics
for seller in seller_analytics:
market_origin_df.loc[period_now.strftime('%Y-%b'),
(runname, name,'seller_analytics',seller)] = seller_analytics[seller]
for buyer in buyer_analytics:
market_origin_df.loc[period_now.strftime('%Y-%b'),
(runname, name,'buyer_analytics',buyer)] = buyer_analytics[buyer]
# recording the step_info
# since this operation can take quite a while, print after every operation
period_time = time.time() - startit_period
if print_details:
print('%.2f : seconds to clear period' % period_time)
#safe df as attribute
self.market_origin_df = market_origin_df
# ## Example Market
#
# In the following code example we use the buyer and supplier objects to create a market. At the market a single price is announced which causes as many units of goods to be swapped as possible. The buyers and sellers stop trading when it is no longer in their own interest to continue.
# +
# import scenarios
inputfile = 'economic growth scenarios.xlsx'
# economic growth percentages
economic_growth = pd.read_excel(inputfile, sheetname='ec_growth', index_col=0, header=[0,1])
## demand for electricity import scenarios spaced by excel
#elec_space = pd.read_excel(inputfile, sheetname='elec_space', skiprows=1, index_col=0, header=0)
# demand for electricity import scenarios spaced by excel
elec_space = pd.read_excel(inputfile, sheetname='elec_space', index_col=0, header=[0,1])
# gasdemand home (percentage increases)
home_savings = {'PACES': 1.01, 'TIDES': .99, 'CIRCLES': .97}
# multilevel ecgrowth
economic_growth2 = pd.read_excel(inputfile, sheetname='ec_growth', index_col=0, header=[0,1])
#economic_growth2['ECONOMIC GROWTH']
# +
# reading excel initialization data back
read_file = 'init_buyers_sellers_lng.xlsx'
df_buyer = pd.read_excel(read_file,orient='index',sheetname='buyers')
df_seller = pd.read_excel(read_file,orient='index',sheetname='sellers')
df_buyer['market'] = [eval(i) for i in df_buyer['market'].values]
df_seller['market'] = [eval(i) for i in df_seller['market'].values]
init_buyer = df_buyer.to_dict('index')
init_seller = df_seller.to_dict('index')
#init_market = {'eu', 'us','as'}, construct markets by unique values
market = []
for i in init_seller:
for x in init_seller[i]['market']: market.append(x)
for i in init_buyer:
for x in init_buyer[i]['market']: market.append(x)
market = list(set(market))
init_market = market
# set the starting time
period_null= datetime.date(2013,1,1)
# -
# ## run the model
# To run the model we create the observer. The observer creates all the other objects and runs the model.
# create observer and run the model
# first data about buyers then sellers and then model ticks
years = 35
# timestep = 12
print_details = False
run_market = {}
run_seller = {}
run_buyer = {}
run_market_origin = {}
run_market_origin_df = {}
for i in ['PACES', 'CIRCLES', 'TIDES']:
runname = i
dtrunstart = datetime.datetime.now()
print('\n%s scenario %d year run started' %(i,years))
obser1 = Observer(init_buyer, init_seller, years*12, i)
obser1.run_it()
#get the info from the observer
run_market[i] = obser1.market_hist
run_seller[i] = obser1.seller_hist
run_buyer[i] = obser1.buyer_hist
run_market_origin_df[i] = obser1.market_origin_df
#run_data[i] = obser1.all_data
dtrunstop = datetime.datetime.now()
print('%s scenario %d year run finished' %(i,years))
print('this run took %s (h:m:s) to complete'% (dtrunstop - dtrunstart))
# +
# timeit
stopit = time.time()
dtstopit = datetime.datetime.now()
print('it took us %s seconds to get to this conclusion' % (stopit-startit))
print('in another notation (h:m:s) %s'% (dtstopit - dtstartit))
# -
# ## Operations Research Formulation
#
# The market can also be formulated as a very simple linear program or linear complementarity problem. It is clearer and easier to implement this market clearing mechanism with agents. One merit of the agent-based approach is that we don't need linear or linearizable supply and demand function.
#
# The auctioneer is effectively following a very simple linear program subject to constraints on units sold. The auctioneer is, in the primal model, maximizing the consumer utility received by customers, with respect to the price being paid, subject to a fixed supply curve. On the dual side the auctioneer is minimizing the cost of production for the supplier, with respect to quantity sold, subject to a fixed demand curve. It is the presumed neutrality of the auctioneer which justifies the honest statement of supply and demand.
#
# An alternative formulation is a linear complementarity problem. Here the presence of an optimal space of trades ensures that there is a Pareto optimal front of possible trades. The perfect opposition of interests in dividing the consumer and producer surplus means that this is a zero sum game. Furthermore the solution to this zero-sum game maximizes societal welfare and is therefore the Hicks optimal solution.
#
# ## Next Steps
#
# A possible addition of this model would be to have a weekly varying demand of customers, for instance caused by the use of natural gas as a heating agent. This would require the bids and asks to be time varying, and for the market to be run over successive time periods. A second addition would be to create transport costs, or enable intermediate goods to be produced. This would need a more elaborate market operator. Another possible addition would be to add a profit maximizing broker. This may require adding belief, fictitious play, or message passing.
#
# The object-orientation of the models will probably need to be further rationalized. Right now the market requires very particular ordering of calls to function correctly.
# ## Time of last run
# Time and date of the last run of this notebook file
# print the time of last run
print('last run of this notebook:')
time.strftime("%a, %d %b %Y %H:%M:%S", time.localtime())
# ## Plotting scenario runs
# For the scenario runs we vary the external factors according to the scenarios. Real plotting is done in a seperate visualization file
plt.subplots()
for market in init_market:
for i in run_market:
run_df = pd.DataFrame(run_market[i])
run_df = run_df[run_df[3]==market]
run_df.set_index(0, inplace=True)
run_df.index = pd.to_datetime(run_df.index)
run_df.index.name = 'month'
run_df.rename(columns={1: 'price', 2: 'quantity'}, inplace=True)
run_df = run_df['price'].resample('A').mean().plot(label=i, title=market)
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
plt.ylabel('€/MWh')
plt.xlabel('Year')
plt.show();
# ### saving data for later
# To keep this file as clear as possible and for efficiency we visualize the results in a separate file. To transfer the model run data we use the Json library (and possibly excel).
# +
today = datetime.date.today().strftime('%Y%m%d')
outputexcel = '.\exceloutput\%srun.xlsx' %today
writer = pd.ExcelWriter(outputexcel)
def write_to_excel():
for i in run_market:
run_df = pd.DataFrame(run_market[i])
run_df.set_index(0, inplace=True)
run_df.index = pd.to_datetime(run_df.index)
run_df.index.name = 'month'
run_df.rename(columns={1: 'price', 2: 'quantity'}, inplace=True)
run_df.to_excel(writer, sheet_name=i)
# uncomment if wanted to write to excel file
#write_to_excel()
# -
# Writing JSON data
# market data
data = run_market
with open('marketdata.json', 'w') as f:
json.dump(data, f)
# seller/reserve data
data = run_seller
with open('sellerdata.json', 'w') as f:
json.dump(data, f)
# buyer data
data = run_buyer
with open('buyerdata.json', 'w') as f:
json.dump(data, f)
# complex dataframes do not work well with Json, so use Pickle
# Merge Dataframes
result = pd.concat([run_market_origin_df[i] for i in run_market_origin_df], axis=1)
#pickle does the job
result.to_pickle('marketdataorigin.pickle', compression='infer', protocol=4)
# testing if complex frames did what it is expected to do
df_pickle = result
for i in df_pickle.columns.levels[0]:
scen=i
market='eu'
df = df_pickle[scen][market]['seller_analytics']
df.index = pd.to_datetime(df.index)
df.resample('A').sum().plot.area(title='%s %s'%(scen,market), colormap='tab20')
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
plt.show()
| longrun/parametersweep/NL dry/Matching Market v20- dry NL.ipynb |
-- ---
-- jupyter:
-- jupytext:
-- text_representation:
-- extension: .hs
-- format_name: light
-- format_version: '1.5'
-- jupytext_version: 1.14.4
-- kernelspec:
-- display_name: Haskell
-- language: haskell
-- name: haskell
-- ---
-- # Day 1: Report Repair
-- https://adventofcode.com/2020/day/1
inputLines = lines <$> readFile "input/day01.txt"
-- Convert input to list of integer numbers:
inputNumbers :: IO [Int]
inputNumbers = map read <$> inputLines
-- This is the test input list:
testInput = [1721, 979, 366, 299, 675, 1456]
-- We need sets for efficient lookup and prevent $\mathcal{O}(N^3)$ complexity in part 2:
import qualified Data.Set as Set
-- # Part 1
-- In part 1, we are looking for the product of two input numbers whose sum is 2020:
solution1 :: [Int] -> Int
solution1 inputList =
let
target = 2020
inputSet = Set.fromList inputList
(x, y) = head
. filter ((`Set.member` inputSet) . snd)
. map (\x -> (x, target - x))
$ inputList
in
x * y
solution1 testInput
-- ## Solution, part 1:
solution1 <$> inputNumbers
-- # Part 2
-- In part 2, we need the product of three input numbers whose sum is 2020:
solution2 :: [Int] -> Int
solution2 inputList =
let
target = 2020
minInput = minimum inputList
inputPairs = [(x, y) | x <- inputList, y <- inputList, x <= y && x + y <= target - minInput]
inputSet = Set.fromList inputList
(x, y, z) = head
. filter (\ (_, _, z) -> Set.member z inputSet)
. map (\ (x, y) -> (x, y, target - x - y))
$ inputPairs
in
x * y * z
solution2 testInput
-- ## Solution, part 2
solution2 <$> inputNumbers
| 2020/day01-haskell.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/davidcavazos/predictive-maintenance/blob/master/data-generator.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="DGU8LFQcjc0M" colab_type="text"
# # Predictive Maintenance Data
# + [markdown] id="2_D0oJxNaYzk" colab_type="text"
# This notebook will guide you through the process of generating statistically significant random data. This data will then be used to train a BigQuery Machine Learning model to predict a target value.
# + [markdown] id="Q4PFXufajhNA" colab_type="text"
# # Setup
# + [markdown] id="z9fidNixagEw" colab_type="text"
# Before you begin, please fill up your Google Cloud project ID and your Cloud Storage bucket name without the `gs://`.
# + id="028LmRwPtbqe" colab_type="code" cellView="form" colab={}
project = "" #@param {type:"string"}
bucket = "" #@param {type:"string"}
dataset = "water_utilities" #@param {type:"string"}
table_training = "pipes_training" #@param {type:"string"}
table = "pipes" #@param {type:"string"}
# + id="GhTugQR4jXoO" colab_type="code" colab={}
import numpy as np
import logging
from pprint import pprint
from google.colab import auth
auth.authenticate_user()
def run(command):
print(f">> {command}")
# !{command}
# + id="1BjoSDLOWKTJ" colab_type="code" colab={}
# Configure bigquery to your project.
run(f"gcloud config set project {project}")
bq = f"bq --project {project}"
# + id="l-FplU9a7ayC" colab_type="code" colab={}
# Create a dataset.
run(f"{bq} mk --dataset {dataset}")
# + [markdown] id="2uvgYcCbsXd1" colab_type="text"
# # Data Generator
# + [markdown] id="kkmD0pPPWSA9" colab_type="text"
# There are different types of fields we can have. For this simple example, we'll only deal with categorical, numerical and dates.
#
# We'll first create a base class `Field` that other fields will inherit from.
# + id="h_PtvRWmsYmi" colab_type="code" colab={}
class Field:
def __init__(self, name):
self.name = name
def generate_random(self, seed=None):
raise NotImplementedError('Field.generate_random(seed)')
def estimate_label(self, value):
raise NotImplementedError('Field.estimate_label(value)')
# + [markdown] id="n21KCuGLWtEg" colab_type="text"
# The `generate_random()` method will allow us to create a random instance of this field. That is, a random element from the categories, or a random number within a range, or a random date within a range.
#
# The `estimate_label()` method will allow us to *evaluate* a value and get a *score* from `0` to `1`. This value will help us calculate the value we want the model to learn to predict, called *label*. They don't have to be strictly between `0` and `1`, but as long as they are mostly smallish numbers.
#
# We will use the scores from all the fields in a pipe to calculate the *label*, which is the value we want to predict. In this case, our label is the number of years between repairs.
# + [markdown] id="xJh5WlZTSV_o" colab_type="text"
# ## CategoricalField
# + [markdown] id="DPj9KfU8Whx_" colab_type="text"
# For categorical fields, we have a finite number of options.
#
# To estimate the label, we will assume the values have a linear relationship. That is, the first value will be `0`, the next ones will increment by fixed steps until the last one which will be `1`.
#
# You could also make them follow a normal distrbution to get values from `0` to `1`, making different values have different weights.
# + id="95An5_sFxHcP" colab_type="code" outputId="93837b30-4637-4f39-dbd9-f7c0eca02c8d" colab={"base_uri": "https://localhost:8080/", "height": 153}
class CategoricalField(Field):
def __init__(self, name, categories):
super().__init__(name)
self.categories = categories
def generate_random(self):
# We only need to make a random choice from all possible categories.
return np.random.choice(self.categories)
def estimate_label(self, value):
# We will assume the last values have more weight than the last,
# so the first value will give 0 and the last value will give 1.
try:
return self.categories.index(value) / (len(self.categories) - 1)
except Exception as e:
logging.error(f"{e}: value={value}, valid={self.categories}")
return 0.5
np.random.seed(42)
field = CategoricalField('Test field name', ['A', 'B', 'C', 'D', 'E'])
print(f"random: {field.generate_random()}")
for value in field.categories:
print(f" {value} label: {field.estimate_label(value)}")
print(f" X label: {field.estimate_label('X')}")
# + id="dMQ0Nksyl4nM" colab_type="code" outputId="73400540-73f6-46f5-bbef-b284db8398af" colab={"base_uri": "https://localhost:8080/", "height": 173}
materials = [
'pvc',
'ceramic',
'copper',
'galvanized iron',
'cast iron',
]
class Material(CategoricalField):
def __init__(self):
super().__init__('material', materials)
def estimate_label(self, value):
return 1 - super().estimate_label(value)
np.random.seed(42)
field = Material()
print(f"random: {field.generate_random()}")
for value in field.categories:
print(f" {value} label: {field.estimate_label(value)}")
print(f" X label: {field.estimate_label('X')}")
# + id="l--Z3gL3mCy-" colab_type="code" outputId="ec34f07d-86f3-4054-be67-888f6caf3662" colab={"base_uri": "https://localhost:8080/", "height": 394}
# Diameter in inches
diameters = [
1/8, 1/4, 3/8, 1/2, 3/4,
1, 1 + 1/4, 1 + 1/2,
2, 2 + 1/2,
3, 4, 5, 6, 8, 10, 12,
]
class Diameter(CategoricalField):
def __init__(self):
super().__init__('diameter_in_inches', diameters)
def estimate_label(self, value):
# If the diameter is not one of the predefined categories, we would still
# like to get a weight that is proportional to where it would have fit
# within the rest of the numbers.
if value in self.categories:
return super().estimate_label(value)
try:
return np.searchsorted(self.categories, value) / len(self.categories)
except Exception as e:
logging.error(f"{e}: value={value}, valid={self.categories}")
return 0.5
np.random.seed(42)
field = Diameter()
print(f"random: {field.generate_random()}")
for value in field.categories:
print(f" {value} label: {field.estimate_label(value)}")
print(f" 9 label: {field.estimate_label(9)}")
print(f" X label: {field.estimate_label('X')}")
# + [markdown] id="0MePjHa3SYl1" colab_type="text"
# ## NumericalField
# + [markdown] id="x--i-3Mpct09" colab_type="text"
# For numbers, we'll also just use a linear uniform distribution, but you could also use any kind of distribution or function that best fits your data.
# + id="q7Qf6vEqzHeK" colab_type="code" outputId="752aacfe-ee39-4e69-eaaa-36ddaecb296e" colab={"base_uri": "https://localhost:8080/", "height": 292}
class NumericalField(Field):
def __init__(self, name, min_value, max_value):
super().__init__(name)
self.min_value = min_value
self.max_value = max_value
def generate_random(self):
# We'll just get a uniform distribution between two values.
return np.random.uniform(self.min_value, self.max_value)
def estimate_label(self, value):
# The first value will be 0 and the last will be 1.
try:
return (value - self.min_value) / (self.max_value - self.min_value)
except Exception as e:
logging.error(f"{e}: value={value}, range=({self.min_value}, {self.max_value})")
return 0.5
np.random.seed(42)
field = NumericalField('Test field name', 5, 10)
print(f"random: {field.generate_random()}")
for value in np.linspace(field.min_value, field.max_value, 10):
print(f" {value} label: {field.estimate_label(value)}")
print(f" 0.0 label: {field.estimate_label(0.0)}")
print(f" 15.0 label: {field.estimate_label(15.0)}")
print(f" X label: {field.estimate_label('X')}")
# + id="eV6XdBSKpjtX" colab_type="code" outputId="1ea12430-0113-43eb-a520-e0b4935311a7" colab={"base_uri": "https://localhost:8080/", "height": 275}
# Length in miles
length_min = 0.1
length_max = 10
class Length(NumericalField):
def __init__(self):
super().__init__('length_in_miles', length_min, length_max)
np.random.seed(42)
field = Length()
print(f"random: {field.generate_random()}")
for value in np.linspace(field.min_value, field.max_value, 10):
print(f" {value} label: {field.estimate_label(value)}")
print(f" 15.0 label: {field.estimate_label(15.0)}")
print(f" X label: {field.estimate_label('X')}")
# + [markdown] id="BeAJWRxtSaxv" colab_type="text"
# ## DateField
# + [markdown] id="XbsKfBehYpBu" colab_type="text"
# Dates are a bit more complicated since they have years, months, days, hours, minutes, seconds and milliseconds. And not even taking into account timezones and leap seconds.
#
# Fortunately there is a simple solution. We can transform them to *Unix time*, which is basically a number representing the number of seconds that have passed since January 1, 1970. The standard library already has functions that take care of this conversion for us, taking leap seconds and everything into account.
#
# Transformed as a number, we can now use the same method as with numbers going between 0 and the current time.
# + id="xPb0Emf-RLOi" colab_type="code" outputId="8e426c7a-9bd9-46ae-af0f-318861efc56d" colab={"base_uri": "https://localhost:8080/", "height": 258}
from datetime import datetime
class DateField(NumericalField):
def __init__(self, name):
super().__init__(name, 0, datetime.now().timestamp())
np.random.seed(42)
field = DateField('Test field name')
print(f"random: {field.generate_random()}")
for value in np.linspace(field.min_value, field.max_value, 10):
print(f" {value} label: {field.estimate_label(value)}")
print(f" X label: {field.estimate_label('X')}")
# + id="c0IOKbePjvVs" colab_type="code" outputId="d13e5cc1-8cea-4dc8-c957-cc8161f81133" colab={"base_uri": "https://localhost:8080/", "height": 258}
class InstallationDate(DateField):
def __init__(self):
super().__init__('installation_date')
np.random.seed(42)
field = InstallationDate()
print(f"random: {field.generate_random()}")
for value in np.linspace(field.min_value, field.max_value, 10):
print(f" {value} label: {field.estimate_label(value)}")
print(f" X label: {field.estimate_label('X')}")
# + [markdown] id="zv9mnF6s9t1x" colab_type="text"
# ## Schema
# + [markdown] id="BzdIIYrrZd_Y" colab_type="text"
# We'll now put together all the fields into a schema. This will allow us to access the columns and to generate new pipes.
#
# Note that we also have a `columns_training()` method, which basically adds a label column. This `label` column is the value the Machine Learning model will try to learn to predict.
#
# In this case we'll name the number of years between repairs as the **`lifespan`**.
# + id="0nHy9OSoUsOq" colab_type="code" outputId="47b66cc7-e2b3-4e66-c416-2c9d13ecf653" colab={"base_uri": "https://localhost:8080/", "height": 156}
import re
class Schema:
def __init__(self, fields, id_name='id', label_name='label'):
self.fields = fields
self.id_name = id_name
self.label_name = label_name
def columns(self):
return [self.id_name] + [field.name for field in self.fields]
def columns_training(self):
return self.columns() + [self.label_name]
def generate_random(self, element_id, seed=None):
np.random.seed(seed)
pipe = {field.name: field.generate_random() for field in self.fields}
pipe[self.id_name] = element_id
return pipe
fields = [
Material(),
InstallationDate(),
Diameter(),
Length(),
]
schema = Schema(fields, id_name='pipe_id', label_name='lifespan')
print(schema.columns())
print(schema.columns_training())
schema.generate_random(0, seed=42)
# + [markdown] id="cu1MZx5s7rUP" colab_type="text"
# # Training the model
# + [markdown] id="QZw0yldBaQ7P" colab_type="text"
# We're almost ready to generate the data, but we're still missing the data labels. We need to calculate the value we want the model to learn to predict. And we'll use the `estimate_label()` methods we created earlier.
# + [markdown] id="TWTyB9grfQBN" colab_type="text"
# ## Data labeling
# + [markdown] id="rgnmc2HrdYrQ" colab_type="text"
# First, we'll calculate a *weight* from `0` to `1` using the `estimate_label()` methods. We can then scale this into any other range. Eventually, we'll scale it to values from `0` to `100`, since it's roughly a good estimate of the lifespan of pipes.
#
# Remember, the weights don't have to be strictly between `0` and `1`, because in the end the model might predict `-0.1` or `1.2`, or even an occasional `3.14`, but most values will be between `0` and `1`. So predictions can fall outside our predefined range of `0` to `100` years.
# + id="Gbh0f75FbnPZ" colab_type="code" outputId="23a06b6c-df9a-4127-ac8b-64834590151d" colab={"base_uri": "https://localhost:8080/", "height": 119}
pipe = schema.generate_random(0, seed=42)
lifespan_weight = 0.0
print("field: value; weight * lifespan = weighted_lifespan")
for weight, field in zip(weights, schema.fields):
label = field.estimate_label(pipe[field.name])
weighted_label = weight * label
print(f" {field.name}: {weight} * {label} = {weighted_label}")
lifespan_weight += weighted_label
print(f"lifespan_weight: {lifespan_weight}")
# + id="tsr8w1OykPG8" colab_type="code" outputId="ac68cd83-ac42-4417-f2bc-47323c747657" colab={"base_uri": "https://localhost:8080/", "height": 119}
def calculate_lifespan_years(schema, pipe):
# We'll assume the fields are sorted by importance.
# That is, the first field will have the largest weight to lifespan, and the
# last field will have the least weight.
indices = np.array(range(len(schema.fields), 0, -1))
weights = indices / indices.sum()
# Calculate a weighted sum for the lifespan estimate of each field.
lifespan_weight = 0.0
for weight, field in zip(weights, schema.fields):
lifespan_weight += weight * field.estimate_label(pipe[field.name])
# Return in a scale from 0 to 100 years.
return 100.0 * lifespan_weight
pipe = schema.generate_random(0, seed=42)
pprint(pipe)
lifespan_years = calculate_lifespan_years(schema, pipe)
print(f"lifespan_years: {lifespan_years}")
# + [markdown] id="77Uqekx3pioV" colab_type="text"
# ## Generating the training data
# + [markdown] id="C42RbNQafVlx" colab_type="text"
# We're finally ready to generate our training data!
#
# We'll just write it to a CSV file since we can easily load that into BigQuery.
# + id="JYfraO7vj10V" colab_type="code" cellView="form" colab={}
training_points = 1000000 #@param {type:"integer"}
# + id="sKfhVnNoe1bt" colab_type="code" outputId="81258c7a-7f02-47b3-c128-64f9089c8a18" colab={"base_uri": "https://localhost:8080/", "height": 204}
import csv
np.random.seed(42)
training_data_file = 'data.training.csv'
with open(training_data_file, 'w') as f:
writer = csv.DictWriter(f, schema.columns_training())
writer.writerow({col: col for col in schema.columns_training()})
for i in range(training_points):
# Get all the fields in the schema plus the lifespan (the model label).
pipe = schema.generate_random(f"pipe {i}")
pipe[schema.label_name] = calculate_lifespan_years(schema, pipe)
writer.writerow(pipe)
# !ls -lh {training_data_file}
# !head {training_data_file}
# + [markdown] id="Gj3vyEU9e2F0" colab_type="text"
# ## Uploading data to BigQuery
# + [markdown] id="Kg9Az0mafjTk" colab_type="text"
# First, we'll copy it to Cloud Storage so BigQuery can load it.
# + id="vIWJvCGYpSyW" colab_type="code" outputId="0c5a9109-b085-4951-c5f9-95c826bb4e12" colab={"base_uri": "https://localhost:8080/", "height": 85}
gcs_training_data_file = f"gs://{bucket}/{training_data_file}"
run(f"gsutil cp {training_data_file} {gcs_training_data_file}")
# + id="l9nC_kDvpm4v" colab_type="code" outputId="8a63af0e-659d-4602-ac53-eeb21ea4915b" colab={"base_uri": "https://localhost:8080/", "height": 71}
# Load the data into the table.
run(f"{bq} load --source_format=CSV --autodetect --replace "
f"{dataset}.{table_training} {gcs_training_data_file}")
# + [markdown] id="vLs8_fLixCKw" colab_type="text"
# ## Training a BigQuery ML Model
# + [markdown] id="AGsAeOP4fxBu" colab_type="text"
# First, let's get 3 rows to see how our data looks like.
# + id="5NayVOUJxFH2" colab_type="code" outputId="674ca008-f591-4aa1-88d9-ffdeb22d41df" colab={"base_uri": "https://localhost:8080/", "height": 142}
# %%bigquery --project {project}
SELECT * FROM `water_utilities.pipes_training` LIMIT 3
# + [markdown] id="RtznLhbLf4A_" colab_type="text"
# Next, we can create and train a BigQuery ML model with a very simple SQL [CREATE MODEL statement](https://cloud.google.com/bigquery/docs/reference/standard-sql/bigqueryml-syntax-create). It is very similar to creating a table, and we'll be able to access it as if it were a table. The only difference is that rather than having the values *stored* in the database, they will be computed as needed.
#
# We'll use linear regression to train our model since we're predicting a number. If you're predicting a classification, you would want to use logistic regression (`logistic_reg`) instead.
# + id="dQw61HFTyia_" colab_type="code" outputId="ba5073cf-71a1-4323-86a9-f354b8edef49" colab={"base_uri": "https://localhost:8080/", "height": 32}
# %%bigquery --project {project}
CREATE OR REPLACE MODEL
`water_utilities.pipe_lifespan`
OPTIONS (
model_type='linear_reg',
input_label_cols=['lifespan']
)
AS SELECT * FROM `water_utilities.pipes_training`
# + [markdown] id="YwPoF88j1wlb" colab_type="text"
# ## Evaluating the model
# + [markdown] id="PmnDxNSEgqR8" colab_type="text"
# So, something happened, but we don't really know any details of it. How can we measure if the model is doing good or bad?
#
# Fortunately, BigQuery ML provides us with an [ML.EVALUATE function](https://cloud.google.com/bigquery/docs/reference/standard-sql/bigqueryml-syntax-evaluate) which will give us more details on the error and variance of our model.
# + id="wlswwo7a0zfF" colab_type="code" outputId="98248fce-3780-45f3-a368-9413b93a2d99" colab={"base_uri": "https://localhost:8080/", "height": 80}
# %%bigquery --project {project}
SELECT *
FROM
ML.EVALUATE(
MODEL `water_utilities.pipe_lifespan`,
(SELECT * FROM `water_utilities.pipes_training`)
)
# + [markdown] id="RxkeudV20xQo" colab_type="text"
# ## Getting predictions
# + [markdown] id="7ZLLkD5OjCZS" colab_type="text"
# We can get prediction results with a simple SELECT statement using the [ML.PREDICT function](https://cloud.google.com/bigquery/docs/reference/standard-sql/bigqueryml-syntax-predict), and access it as if it were another table.
# + id="GhG6yOWx35Pa" colab_type="code" outputId="4b680aa3-ee22-41c8-a30f-93b558b05e96" colab={"base_uri": "https://localhost:8080/", "height": 142}
# %%bigquery --project {project}
SELECT
predicted_lifespan
FROM
ML.PREDICT(
MODEL `water_utilities.pipe_lifespan`,
(SELECT * FROM `water_utilities.pipes_training` LIMIT 3)
)
# + [markdown] id="oZ8I0ghajRhQ" colab_type="text"
# Let's compare some of our generated labels with the predicted values. They are surprisingly similar, so it's looking good!
# + id="taY7dOjO16o3" colab_type="code" outputId="1e57dd59-8821-4334-91ce-482d6d44e547" colab={"base_uri": "https://localhost:8080/", "height": 359}
# %%bigquery --project {project}
SELECT
lifespan,
predicted_lifespan
FROM
ML.PREDICT(
MODEL `water_utilities.pipe_lifespan`,
(SELECT * FROM `water_utilities.pipes_training` LIMIT 10)
)
# + [markdown] id="jrrLjzLpBTlC" colab_type="text"
# # Generating the app data
# + [markdown] id="FRMQPWpdjzN7" colab_type="text"
# We'll now generate a second dataset. This will be to populate the app since we don't have real data, but you could skip this step if you already have data.
#
# For this dataset, however, we no longer need the label. And we'll also add `last_repair_date` and `repair_cost` columns. We'll add the predicted lifespan to the last repair date to get the predicted break date.
#
# We'll also create a simple repair cost estimate to test the budget analysis on the app.
# + colab_type="code" cellView="form" id="2w-4NfhwtNmD" colab={}
total_pipes = 1000 #@param {type:"integer"}
# + id="MW1ORyle3Vqh" colab_type="code" outputId="2860222f-3ca4-4729-9b77-237da2a82e3c" colab={"base_uri": "https://localhost:8080/", "height": 224}
import csv
np.random.seed(42)
data_file = 'data.csv'
with open(data_file, 'w') as f:
columns = schema.columns() + ['last_repair_date', 'repair_cost']
writer = csv.DictWriter(f, columns)
writer.writerow({col: col for col in columns})
for i in range(total_pipes):
pipe = schema.generate_random(f"pipe {i}", seed=i)
# The last repair date is any date between the installation date and now.
pipe['last_repair_date'] = np.random.uniform(pipe['installation_date'], datetime.now().timestamp())
# The repair cost will be around $10,000 and affected by the pipe length.
pipe['repair_cost'] = np.log(pipe['length_in_miles']) * 10000
writer.writerow(pipe)
# !ls -lh {data_file}
# !head {data_file}
# + [markdown] id="ksjFYJMKkod4" colab_type="text"
# Next, we'll also copy it to Cloud Storage and upload it into BigQuery to another table.
# + id="h81jAkVk4krI" colab_type="code" outputId="098712a3-ef9a-4d18-baa7-0f3b0a7e798b" colab={"base_uri": "https://localhost:8080/", "height": 105}
gcs_data_file = f"gs://{bucket}/{data_file}"
run(f"gsutil cp {data_file} {gcs_data_file}")
# + id="Pd9-SZTd4YUL" colab_type="code" outputId="4ddfe870-1390-4a50-e062-7f44b65cfa88" colab={"base_uri": "https://localhost:8080/", "height": 71}
# Load the data into the table.
run(f"{bq} load --source_format=CSV --autodetect --replace "
f"{dataset}.{table} {gcs_data_file}")
# + id="rU7UGe5M5XQi" colab_type="code" outputId="1793890f-2028-4b5d-e346-21374c31583f" colab={"base_uri": "https://localhost:8080/", "height": 142}
# %%bigquery --project {project}
SELECT * FROM `water_utilities.pipes` LIMIT 3
# + [markdown] id="FAjQkEa4kvKH" colab_type="text"
# Finally, we can get predictions on our newly created dataset.
# + id="uFgDXg1u5can" colab_type="code" outputId="3c0fa8cd-ac78-4009-d2e8-05250056f64c" colab={"base_uri": "https://localhost:8080/", "height": 162}
# %%bigquery --project {project}
SELECT
pipe_id,
material,
diameter_in_inches,
length_in_miles,
installation_date,
last_repair_date,
last_repair_date + predicted_lifespan*60*60*24*365 AS predicted_break_date
FROM
ML.PREDICT(
MODEL `water_utilities.pipe_lifespan`,
(SELECT * FROM `water_utilities.pipes` LIMIT 3)
)
| data-generator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
### 1. Write a Python Program to Display Fibonacci Sequence Using Recursion?
def fibonacci(a, b, n):
"""
Prints n fibonacci numbers starting from a, second element b
:param a: first element of your sequence
:param b: second element of your sequence
:return: None
"""
print(a, end = " ")
a, b = b, a+b
if n > 0:
n -=1
fibonacci(a, b, n)
fibonacci(0, 1, 10)
# +
### 2. Write a Python Program to find Factorial of Number Using Recursion?
def factorial(n):
"""
Returns the factorial of given number
:param n: number to find the factorial of
:return: None
"""
if n > 0:
return n * factorial(n-1)
return 1
factorial(6)
# +
### 3. Write a Python Program to calculate your Body Mass Index?
def bmi_calculate(height, weight):
"""
Calculates and shows your BMI score depending on your weight and height
:param height: height in meters
:param weight: weight in Kgs
"""
bmi = float(weight)/float(height*height)
if(bmi>0):
print(f"Your Body Mass Index (BMI) score is: {round(bmi, 2)}")
if(bmi<=16):
print("You are very underweight")
elif(bmi<=18.5):
print("You are underweight")
elif(bmi<=25):
print("Congrats! You are Healthy")
elif(bmi<=30):
print("You are overweight")
else:
print("You are very overweight")
else:
print("Could not calculate your BMI Score, enter valid details!")
h=float(input("Enter your height in meters: "))
w=float(input("Enter your Weight in Kg: "))
bmi_calculate(h, w)
# +
### 4. Write a Python Program to calculate the natural logarithm of any number?
import math
num = float(input("Enter a number to calculat its natural logarithm (base e): "))
print(f"natural logarithm of {num} is: {round(math.log(num), 4)}")
# +
### 5. Write a Python Program for cube sum of first n natural numbers?
# formula: (n(n+1)*0.5)^2
inp = int(input("Enter a number to find sum of cubes of natural numbers till: "))
print(f"cube sum of first {inp} natural numbers is : {(inp*(inp+1)*0.5)**2}")
| assignments/PythonBasicsProgramming/Programming_Assignment_6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
# Licensed under the MIT License.
# 
# # Facial Expression Recognition (FER+) using ONNX Runtime on Azure ML
#
# This example shows how to deploy an image classification neural network using the Facial Expression Recognition ([FER](https://www.kaggle.com/c/challenges-in-representation-learning-facial-expression-recognition-challenge/data)) dataset and Open Neural Network eXchange format ([ONNX](http://aka.ms/onnxdocarticle)) on the Azure Machine Learning platform. This tutorial will show you how to deploy a FER+ model from the [ONNX model zoo](https://github.com/onnx/models), use it to make predictions using ONNX Runtime Inference, and deploy it as a web service in Azure.
#
# Throughout this tutorial, we will be referring to ONNX, a neural network exchange format used to represent deep learning models. With ONNX, AI developers can more easily move models between state-of-the-art tools (CNTK, PyTorch, Caffe, MXNet, TensorFlow) and choose the combination that is best for them. ONNX is developed and supported by a community of partners including Microsoft AI, Facebook, and Amazon. For more information, explore the [ONNX website](http://onnx.ai) and [open source files](https://github.com/onnx).
#
# [ONNX Runtime](https://aka.ms/onnxruntime-python) is the runtime engine that enables evaluation of trained machine learning (Traditional ML and Deep Learning) models with high performance and low resource utilization. We use the CPU version of ONNX Runtime in this tutorial, but will soon be releasing an additional tutorial for deploying this model using ONNX Runtime GPU.
#
# #### Tutorial Objectives:
#
# 1. Describe the FER+ dataset and pretrained Convolutional Neural Net ONNX model for Emotion Recognition, stored in the ONNX model zoo.
# 2. Deploy and run the pretrained FER+ ONNX model on an Azure Machine Learning instance
# 3. Predict labels for test set data points in the cloud using ONNX Runtime and Azure ML
# ## Prerequisites
#
# ### 1. Install Azure ML SDK and create a new workspace
# If you are using an Azure Machine Learning Notebook VM, you are all set. Otherwise, please follow [Azure ML configuration notebook](../../../configuration.ipynb) to set up your environment.
#
# ### 2. Install additional packages needed for this Notebook
# You need to install the popular plotting library `matplotlib`, the image manipulation library `opencv`, and the `onnx` library in the conda environment where Azure Maching Learning SDK is installed.
#
# ```sh
# (myenv) $ pip install matplotlib onnx opencv-python
# ```
#
# **Debugging tip**: Make sure that to activate your virtual environment (myenv) before you re-launch this notebook using the `jupyter notebook` comand. Choose the respective Python kernel for your new virtual environment using the `Kernel > Change Kernel` menu above. If you have completed the steps correctly, the upper right corner of your screen should state `Python [conda env:myenv]` instead of `Python [default]`.
#
# ### 3. Download sample data and pre-trained ONNX model from ONNX Model Zoo.
#
# In the following lines of code, we download [the trained ONNX Emotion FER+ model and corresponding test data](https://github.com/onnx/models/tree/master/vision/body_analysis/emotion_ferplus) and place them in the same folder as this tutorial notebook. For more information about the FER+ dataset, please visit Microsoft Researcher <NAME>'s [FER+ source data repository](https://github.com/ebarsoum/FERPlus).
# +
# urllib is a built-in Python library to download files from URLs
# Objective: retrieve the latest version of the ONNX Emotion FER+ model files from the
# ONNX Model Zoo and save it in the same folder as this tutorial
import urllib.request
onnx_model_url = "https://www.cntk.ai/OnnxModels/emotion_ferplus/opset_7/emotion_ferplus.tar.gz"
urllib.request.urlretrieve(onnx_model_url, filename="emotion_ferplus.tar.gz")
# the ! magic command tells our jupyter notebook kernel to run the following line of
# code from the command line instead of the notebook kernel
# We use tar and xvcf to unzip the files we just retrieved from the ONNX model zoo
# !tar xvzf emotion_ferplus.tar.gz
# -
# ## Deploy a VM with your ONNX model in the Cloud
#
# ### Load Azure ML workspace
#
# We begin by instantiating a workspace object from the existing workspace created earlier in the configuration notebook.
# +
# Check core SDK version number
import azureml.core
print("SDK version:", azureml.core.VERSION)
# +
from azureml.core import Workspace
ws = Workspace.from_config()
print(ws.name, ws.location, ws.resource_group, sep = '\n')
# -
# ### Registering your model with Azure ML
# +
model_dir = "emotion_ferplus" # replace this with the location of your model files
# leave as is if it's in the same folder as this notebook
# +
from azureml.core.model import Model
model = Model.register(model_path = model_dir + "/" + "model.onnx",
model_name = "onnx_emotion",
tags = {"onnx": "demo"},
description = "FER+ emotion recognition CNN from ONNX Model Zoo",
workspace = ws)
# -
# ### Optional: Displaying your registered models
#
# This step is not required, so feel free to skip it.
models = ws.models
for name, m in models.items():
print("Name:", name,"\tVersion:", m.version, "\tDescription:", m.description, m.tags)
# ### ONNX FER+ Model Methodology
#
# The image classification model we are using is pre-trained using Microsoft's deep learning cognitive toolkit, [CNTK](https://github.com/Microsoft/CNTK), from the [ONNX model zoo](http://github.com/onnx/models). The model zoo has many other models that can be deployed on cloud providers like AzureML without any additional training. To ensure that our cloud deployed model works, we use testing data from the well-known FER+ data set, provided as part of the [trained Emotion Recognition model](https://github.com/onnx/models/tree/master/vision/body_analysis/emotion_ferplus) in the ONNX model zoo.
#
# The original Facial Emotion Recognition (FER) Dataset was released in 2013 by <NAME> and <NAME> as part of a [Kaggle Competition](https://www.kaggle.com/c/challenges-in-representation-learning-facial-expression-recognition-challenge/data), but some of the labels are not entirely appropriate for the expression. In the FER+ Dataset, each photo was evaluated by at least 10 croud sourced reviewers, creating a more accurate basis for ground truth.
#
# You can see the difference of label quality in the sample model input below. The FER labels are the first word below each image, and the FER+ labels are the second word below each image.
#
# 
#
# ***Input: Photos of cropped faces from FER+ Dataset***
#
# ***Task: Classify each facial image into its appropriate emotions in the emotion table***
#
# ``` emotion_table = {'neutral':0, 'happiness':1, 'surprise':2, 'sadness':3, 'anger':4, 'disgust':5, 'fear':6, 'contempt':7} ```
#
# ***Output: Emotion prediction for input image***
#
#
# Remember, once the application is deployed in Azure ML, you can use your own images as input for the model to classify.
# +
# for images and plots in this notebook
import matplotlib.pyplot as plt
# display images inline
# %matplotlib inline
# -
# ### Model Description
#
# The FER+ model from the ONNX Model Zoo is summarized by the graphic below. You can see the entire workflow of our pre-trained model in the following image from Barsoum et. al's paper ["Training Deep Networks for Facial Expression Recognition
# with Crowd-Sourced Label Distribution"](https://arxiv.org/pdf/1608.01041.pdf), with our (64 x 64) input images and our output probabilities for each of the labels.
# 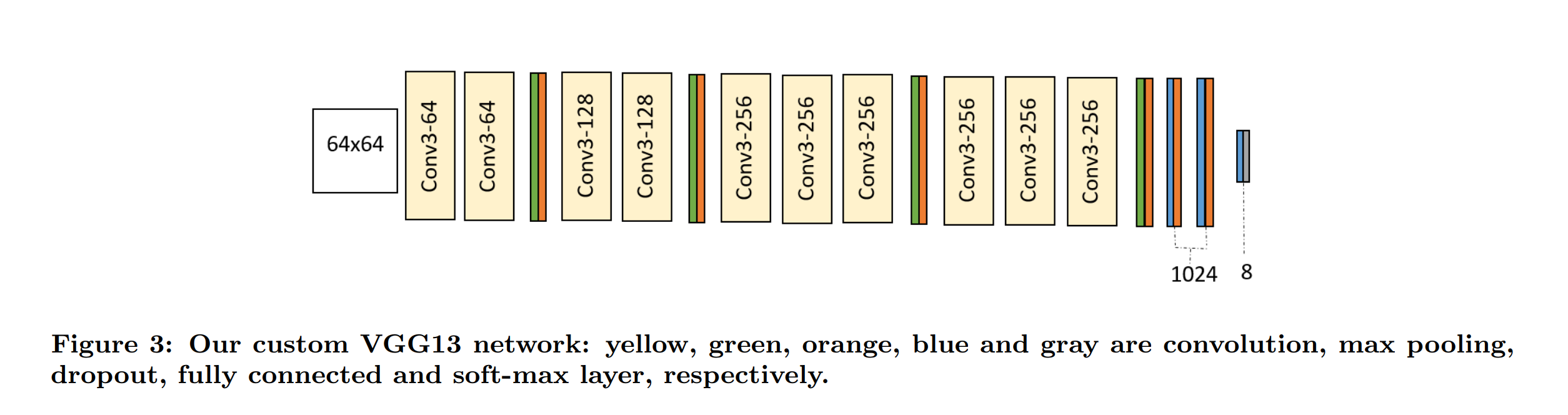
# ### Specify our Score and Environment Files
# We are now going to deploy our ONNX Model on AML with inference in ONNX Runtime. We begin by writing a score.py file, which will help us run the model in our Azure ML virtual machine (VM), and then specify our environment by writing a yml file. You will also notice that we import the onnxruntime library to do runtime inference on our ONNX models (passing in input and evaluating out model's predicted output). More information on the API and commands can be found in the [ONNX Runtime documentation](https://aka.ms/onnxruntime).
#
# ### Write Score File
#
# A score file is what tells our Azure cloud service what to do. After initializing our model using azureml.core.model, we start an ONNX Runtime inference session to evaluate the data passed in on our function calls.
# +
# %%writefile score.py
import json
import numpy as np
import onnxruntime
import sys
import os
import time
def init():
global session, input_name, output_name
model = os.path.join(os.getenv('AZUREML_MODEL_DIR'), 'model.onnx')
session = onnxruntime.InferenceSession(model, None)
input_name = session.get_inputs()[0].name
output_name = session.get_outputs()[0].name
def run(input_data):
'''Purpose: evaluate test input in Azure Cloud using onnxruntime.
We will call the run function later from our Jupyter Notebook
so our azure service can evaluate our model input in the cloud. '''
try:
# load in our data, convert to readable format
data = np.array(json.loads(input_data)['data']).astype('float32')
start = time.time()
r = session.run([output_name], {input_name : data})
end = time.time()
result = emotion_map(postprocess(r[0]))
result_dict = {"result": result,
"time_in_sec": [end - start]}
except Exception as e:
result_dict = {"error": str(e)}
return json.dumps(result_dict)
def emotion_map(classes, N=1):
"""Take the most probable labels (output of postprocess) and returns the
top N emotional labels that fit the picture."""
emotion_table = {'neutral':0, 'happiness':1, 'surprise':2, 'sadness':3,
'anger':4, 'disgust':5, 'fear':6, 'contempt':7}
emotion_keys = list(emotion_table.keys())
emotions = []
for i in range(N):
emotions.append(emotion_keys[classes[i]])
return emotions
def softmax(x):
"""Compute softmax values (probabilities from 0 to 1) for each possible label."""
x = x.reshape(-1)
e_x = np.exp(x - np.max(x))
return e_x / e_x.sum(axis=0)
def postprocess(scores):
"""This function takes the scores generated by the network and
returns the class IDs in decreasing order of probability."""
prob = softmax(scores)
prob = np.squeeze(prob)
classes = np.argsort(prob)[::-1]
return classes
# -
# ### Write Environment File
# +
from azureml.core.conda_dependencies import CondaDependencies
myenv = CondaDependencies.create(pip_packages=["numpy", "onnxruntime", "azureml-core"])
with open("myenv.yml","w") as f:
f.write(myenv.serialize_to_string())
# -
# ### Setup inference configuration
# +
from azureml.core.model import InferenceConfig
inference_config = InferenceConfig(runtime= "python",
entry_script="score.py",
conda_file="myenv.yml",
extra_docker_file_steps = "Dockerfile")
# -
# ### Deploy the model
# +
from azureml.core.webservice import AciWebservice
aciconfig = AciWebservice.deploy_configuration(cpu_cores = 1,
memory_gb = 1,
tags = {'demo': 'onnx'},
description = 'ONNX for emotion recognition model')
# -
# The following cell will likely take a few minutes to run as well.
aci_service_name = 'onnx-demo-emotion'
print("Service", aci_service_name)
aci_service = Model.deploy(ws, aci_service_name, [model], inference_config, aciconfig)
aci_service.wait_for_deployment(True)
print(aci_service.state)
if aci_service.state != 'Healthy':
# run this command for debugging.
print(aci_service.get_logs())
# If your deployment fails, make sure to delete your aci_service before trying again!
# aci_service.delete()
# ### Success!
#
# If you've made it this far, you've deployed a working VM with a facial emotion recognition model running in the cloud using Azure ML. Congratulations!
#
# Let's see how well our model deals with our test images.
# ## Testing and Evaluation
#
# ### Useful Helper Functions
#
# We preprocess and postprocess our data (see score.py file) using the helper functions specified in the [ONNX FER+ Model page in the Model Zoo repository](https://github.com/onnx/models/tree/master/vision/body_analysis/emotion_ferplus).
# +
def emotion_map(classes, N=1):
"""Take the most probable labels (output of postprocess) and returns the
top N emotional labels that fit the picture."""
emotion_table = {'neutral':0, 'happiness':1, 'surprise':2, 'sadness':3,
'anger':4, 'disgust':5, 'fear':6, 'contempt':7}
emotion_keys = list(emotion_table.keys())
emotions = []
for c in range(N):
emotions.append(emotion_keys[classes[c]])
return emotions
def softmax(x):
"""Compute softmax values (probabilities from 0 to 1) for each possible label."""
x = x.reshape(-1)
e_x = np.exp(x - np.max(x))
return e_x / e_x.sum(axis=0)
def postprocess(scores):
"""This function takes the scores generated by the network and
returns the class IDs in decreasing order of probability."""
prob = softmax(scores)
prob = np.squeeze(prob)
classes = np.argsort(prob)[::-1]
return classes
# -
# ### Load Test Data
#
# These are already in your directory from your ONNX model download (from the model zoo).
#
# Notice that our Model Zoo files have a .pb extension. This is because they are [protobuf files (Protocol Buffers)](https://developers.google.com/protocol-buffers/docs/pythontutorial), so we need to read in our data through our ONNX TensorProto reader into a format we can work with, like numerical arrays.
# +
# to manipulate our arrays
import numpy as np
# read in test data protobuf files included with the model
import onnx
from onnx import numpy_helper
# to use parsers to read in our model/data
import json
import os
test_inputs = []
test_outputs = []
# read in 3 testing images from .pb files
test_data_size = 3
for num in np.arange(test_data_size):
input_test_data = os.path.join(model_dir, 'test_data_set_{0}'.format(num), 'input_0.pb')
output_test_data = os.path.join(model_dir, 'test_data_set_{0}'.format(num), 'output_0.pb')
# convert protobuf tensors to np arrays using the TensorProto reader from ONNX
tensor = onnx.TensorProto()
with open(input_test_data, 'rb') as f:
tensor.ParseFromString(f.read())
input_data = numpy_helper.to_array(tensor)
test_inputs.append(input_data)
with open(output_test_data, 'rb') as f:
tensor.ParseFromString(f.read())
output_data = numpy_helper.to_array(tensor)
output_processed = emotion_map(postprocess(output_data[0]))[0]
test_outputs.append(output_processed)
# + [markdown] nbpresent={"id": "c3f2f57c-7454-4d3e-b38d-b0946cf066ea"}
# ### Show some sample images
# We use `matplotlib` to plot 3 test images from the dataset.
# + nbpresent={"id": "396d478b-34aa-4afa-9898-cdce8222a516"}
plt.figure(figsize = (20, 20))
for test_image in np.arange(3):
test_inputs[test_image].reshape(1, 64, 64)
plt.subplot(1, 8, test_image+1)
plt.axhline('')
plt.axvline('')
plt.text(x = 10, y = -10, s = test_outputs[test_image], fontsize = 18)
plt.imshow(test_inputs[test_image].reshape(64, 64), cmap = plt.cm.gray)
plt.show()
# -
# ### Run evaluation / prediction
# +
plt.figure(figsize = (16, 6), frameon=False)
plt.subplot(1, 8, 1)
plt.text(x = 0, y = -30, s = "True Label: ", fontsize = 13, color = 'black')
plt.text(x = 0, y = -20, s = "Result: ", fontsize = 13, color = 'black')
plt.text(x = 0, y = -10, s = "Inference Time: ", fontsize = 13, color = 'black')
plt.text(x = 3, y = 14, s = "Model Input", fontsize = 12, color = 'black')
plt.text(x = 6, y = 18, s = "(64 x 64)", fontsize = 12, color = 'black')
plt.imshow(np.ones((28,28)), cmap=plt.cm.Greys)
for i in np.arange(test_data_size):
input_data = json.dumps({'data': test_inputs[i].tolist()})
# predict using the deployed model
r = json.loads(aci_service.run(input_data))
if "error" in r:
print(r['error'])
break
result = r['result'][0]
time_ms = np.round(r['time_in_sec'][0] * 1000, 2)
ground_truth = test_outputs[i]
# compare actual value vs. the predicted values:
plt.subplot(1, 8, i+2)
plt.axhline('')
plt.axvline('')
# use different color for misclassified sample
font_color = 'red' if ground_truth != result else 'black'
clr_map = plt.cm.Greys if ground_truth != result else plt.cm.gray
# ground truth labels are in blue
plt.text(x = 10, y = -70, s = ground_truth, fontsize = 18, color = 'blue')
# predictions are in black if correct, red if incorrect
plt.text(x = 10, y = -45, s = result, fontsize = 18, color = font_color)
plt.text(x = 5, y = -22, s = str(time_ms) + ' ms', fontsize = 14, color = font_color)
plt.imshow(test_inputs[i].reshape(64, 64), cmap = clr_map)
plt.show()
# -
# ### Try classifying your own images!
# +
# Preprocessing functions take your image and format it so it can be passed
# as input into our ONNX model
import cv2
def rgb2gray(rgb):
"""Convert the input image into grayscale"""
return np.dot(rgb[...,:3], [0.299, 0.587, 0.114])
def resize_img(img_to_resize):
"""Resize image to FER+ model input dimensions"""
r_img = cv2.resize(img_to_resize, dsize=(64, 64), interpolation=cv2.INTER_AREA)
r_img.resize((1, 1, 64, 64))
return r_img
def preprocess(img_to_preprocess):
"""Resize input images and convert them to grayscale."""
if img_to_preprocess.shape == (64, 64):
img_to_preprocess.resize((1, 1, 64, 64))
return img_to_preprocess
grayscale = rgb2gray(img_to_preprocess)
processed_img = resize_img(grayscale)
return processed_img
# +
# Replace the following string with your own path/test image
# Make sure your image is square and the dimensions are equal (i.e. 100 * 100 pixels or 28 * 28 pixels)
# Any PNG or JPG image file should work
# Make sure to include the entire path with // instead of /
# e.g. your_test_image = "C:/Users/vinitra.swamy/Pictures/face.png"
your_test_image = "<path to file>"
import matplotlib.image as mpimg
if your_test_image != "<path to file>":
img = mpimg.imread(your_test_image)
plt.subplot(1,3,1)
plt.imshow(img, cmap = plt.cm.Greys)
print("Old Dimensions: ", img.shape)
img = preprocess(img)
print("New Dimensions: ", img.shape)
else:
img = None
# -
if img is None:
print("Add the path for your image data.")
else:
input_data = json.dumps({'data': img.tolist()})
try:
r = json.loads(aci_service.run(input_data))
result = r['result'][0]
time_ms = np.round(r['time_in_sec'][0] * 1000, 2)
except KeyError as e:
print(str(e))
plt.figure(figsize = (16, 6))
plt.subplot(1,8,1)
plt.axhline('')
plt.axvline('')
plt.text(x = -10, y = -40, s = "Model prediction: ", fontsize = 14)
plt.text(x = -10, y = -25, s = "Inference time: ", fontsize = 14)
plt.text(x = 100, y = -40, s = str(result), fontsize = 14)
plt.text(x = 100, y = -25, s = str(time_ms) + " ms", fontsize = 14)
plt.text(x = -10, y = -10, s = "Model Input image: ", fontsize = 14)
plt.imshow(img.reshape((64, 64)), cmap = plt.cm.gray)
# +
# remember to delete your service after you are done using it!
# aci_service.delete()
# -
# ## Conclusion
#
# Congratulations!
#
# In this tutorial, you have:
# - familiarized yourself with ONNX Runtime inference and the pretrained models in the ONNX model zoo
# - understood a state-of-the-art convolutional neural net image classification model (FER+ in ONNX) and deployed it in the Azure ML cloud
# - ensured that your deep learning model is working perfectly (in the cloud) on test data, and checked it against some of your own!
#
# Next steps:
# - If you have not already, check out another interesting ONNX/AML application that lets you set up a state-of-the-art [handwritten image classification model (MNIST)](https://github.com/Azure/MachineLearningNotebooks/blob/master/how-to-use-azureml/deployment/onnx/onnx-inference-mnist-deploy.ipynb) in the cloud! This tutorial deploys a pre-trained ONNX Computer Vision model for handwritten digit classification in an Azure ML virtual machine.
# - Keep an eye out for an updated version of this tutorial that uses ONNX Runtime GPU.
# - Contribute to our [open source ONNX repository on github](http://github.com/onnx/onnx) and/or add to our [ONNX model zoo](http://github.com/onnx/models)
| notebooks/onnx/onnx-inference-facial-expression-recognition-deploy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:m5_env] *
# language: python
# name: conda-env-m5_env-py
# ---
# +
import pandas as pd
import numpy as np
import pickle
from joblib import Parallel, delayed
from fbprophet import Prophet
# -
# ## Params
# +
IS_EVAL = True
DATA_PATH = '../data/'
if IS_EVAL:
PERIOD_LABEL = 'evaluation'
else:
PERIOD_LABEL = 'validation'
# -
# ## Load needed data
prophet_df = pd.read_pickle(DATA_PATH + 'refined/prophet_df_' + PERIOD_LABEL + '.pkl')
prophet_params = pd.read_csv(DATA_PATH + 'external/params_prophet_store_dpt_' + PERIOD_LABEL + '.csv')
sample_submission = pd.read_csv(DATA_PATH + 'raw/sample_submission.csv')
# ## Forecast
def forecast_prophet(store_id, dept_id):
# Reduce df & params on current
df = prophet_df[(prophet_df['store_id'] == store_id) & (prophet_df['dept_id'] == dept_id)].copy()
params = prophet_params.loc[(prophet_params['store_id'] == store_id) &
(prophet_params['dept_id'] == dept_id), 'params'].values[0]
params = eval(params) # String to dict
# Define model
m = Prophet(
yearly_seasonality=False,
weekly_seasonality=False,
daily_seasonality=False,
uncertainty_samples=False,
changepoint_range=params['changepoint_range'],
changepoint_prior_scale=params['changepoint_prior_scale'],
holidays_prior_scale=params['holidays_prior_scale'],
seasonality_mode=params['seasonality_mode']
)
m.add_seasonality(
name='yearly',
period=365.25,
fourier_order=params['yearly_order'],
prior_scale=params['yearly_prior_scale']
)
m.add_seasonality(
name='monthly',
period=365.25/12,
fourier_order=params['monthly_order'],
prior_scale=params['monthly_prior_scale']
)
m.add_seasonality(
name='weekly',
period=7,
fourier_order=params['weekly_order'],
prior_scale=params['weekly_prior_scale']
)
# Add holidays/regressor
m.add_country_holidays(country_name='US')
m.add_regressor('snap', mode=params['snap_mode'])
m.add_regressor('price', mode=params['price_mode'])
m.add_regressor('dom', mode=params['dom_mode'])
# Fit
m.fit(df.dropna(subset=['y'])) # drop pred period
# Predict
future = m.make_future_dataframe(periods=28, freq='D', include_history=False)
future['snap'] = df['snap'].values[-28:]
future['price'] = df['price'].values[-28:]
future['dom'] = df['dom'].values[-28:]
fcst = m.predict(future)
# Add ids & d to the output
fcst['store_id'] = store_id
fcst['dept_id'] = dept_id
fcst['d'] = df['d'].values[-28:]
# Round forecast
fcst['yhat'] = fcst['yhat'].round().astype(int)
fcst.loc[fcst['yhat'] < 0, 'yhat'] = 0
return fcst[['store_id', 'dept_id', 'd', 'yhat']]
# +
ts_ids = prophet_df[['store_id', 'dept_id']].drop_duplicates().reset_index(drop=True)
res = Parallel(n_jobs=-1, verbose=1)\
(delayed(forecast_prophet)\
(row['store_id'], row['dept_id']) for _, row in ts_ids.iterrows())
# -
# retrieve forecasts
forecast_df = pd.concat(res)
forecast_df.head()
# ## Format and save as external data
forecast_df = pd.pivot_table(forecast_df, index=['store_id', 'dept_id'], columns='d', values='yhat')
forecast_df.columns = ['F' + str(int(c) + 1) for c in range(28)]
forecast_df = forecast_df.reset_index()
forecast_df.head()
forecast_df.to_csv(DATA_PATH + 'external/forecast_prophet_store_dpt_' + PERIOD_LABEL + '.csv', index=False)
| prophet_store_dpt_pipeline/prophet_store_dpt_fit_predict.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import tensorflow as tf
import keras
import os
import scipy.io as sio
import pickle
import matplotlib.pyplot as plt
from scipy import stats
from os import listdir
from tensorflow.python.client import device_lib
from keras.models import Sequential, load_model
from keras.layers import CuDNNGRU, Bidirectional, LeakyReLU, Dense, Dropout, Input, Convolution1D, Layer,Flatten, Reshape
from keras.callbacks import ModelCheckpoint, LearningRateScheduler, EarlyStopping
from keras.models import Model
from keras.layers.normalization import BatchNormalization
from keras import regularizers, initializers, constraints
from keras import backend as K
from sklearn.model_selection import StratifiedKFold
from sklearn.metrics import f1_score
from keras.utils import plot_model
random_seed = 34
batch_size = 16
epochs = 100
# -
from tensorflow.python.client import device_lib
print(device_lib.list_local_devices())
# +
class PharmacophoreException(Exception):
pass
class PharmacophoreFileEndException(PharmacophoreException):
pass
class PharmacophorePoint(object):
def __init__(self, code, cx, cy, cz, alpha, norm, nx, ny, nz):
self.code = code
self.cx = float(cx)
self.cy = float(cy)
self.cz = float(cz)
self.alpha = float(alpha)
self.norm = int(norm)
self.nx = float(nx)
self.ny = float(ny)
self.nz = float(nz)
@classmethod
def from_line(cls, line):
return cls(*line.split())
def to_line(self):
return "{} {} {} {} {} {} {} {} {}".format(self.code, self.cx, self.cy, self.cz, self.alpha, self.norm,\
self.nx, self.ny, self.nz)
def __str__(self):
return self.to_line()
class Pharmacophore(object):
def __init__(self, name, points):
self.name = name
self.points = points
@classmethod
def from_stream(cls, stream):
name = stream.readline().strip()
points = []
line = stream.readline().strip()
if not line:
raise PharmacophoreFileEndException("End of file")
while line != "$$$$" or not line:
points.append(PharmacophorePoint.from_line(line))
line = stream.readline().strip()
if not line:
raise PharmacophoreException("Wrong format, no end line")
return cls(name, points)
@classmethod
def from_file(cls, file_path):
with open(file_path) as fd:
return cls.from_stream(fd)
def write_to_stream(self, stream):
stream.write("{}\n".format(self.name))
for point in self.points:
stream.write("{}\n".format(point.to_line()))
stream.write("$$$$\n".format(self.name))
def write_to_file(self, file_path):
with open(file_path, "w") as fd:
self.write_to_stream(fd)
def __str__(self):
return "{}\n{}\n$$$$".format(self.name,
"\n".join(str(x) for x in self.points))
def __len__(self):
return len(self.points)
def sample(self, name, n):
points = sample(self.points, min(n, len(self)))
return Pharmacophore(name, points)
class PharmDatabaseException(Exception):
pass
def calc_pharmacophore(lig_path, ph_path):
proc = Popen(
"align-it --dbase {} --pharmacophore {}".format(lig_path, ph_path),
shell=True,
stdout=PIPE, stderr=PIPE)
_ = proc.communicate()
class PharmDatabase(object):
def __init__(self, path_to_ligands, path_to_ph_db, is_calculated=False):
self.path_to_ligands = path_to_ligands
self.path_to_ph_db = path_to_ph_db
self.is_calculated = is_calculated
def repair_database(self):
pass
def calc_database(self):
if not self.path_to_ph_db:
self.calc_pharmacophore(self.path_to_ligands, self.path_to_ph_db)
def sample_database(self):
pass
def iter_database(self):
if not self.is_calculated:
raise PharmDatabaseException("Not calculated")
with open(self.path_to_ph_db, 'r') as fd:
while True:
try:
pharmacophore = Pharmacophore.from_stream(fd)
yield pharmacophore
except PharmacophoreFileEndException:
break
# -
from rdkit import Chem
from rdkit.Chem import AllChem
def smi_to_morganfingerprint(smi, radius, MORGAN_SIZE):
mol = Chem.MolFromSmiles(smi)
if mol is not None:
tempReturn = np.zeros(MORGAN_SIZE, dtype=np.int8)
vec = AllChem.GetMorganFingerprintAsBitVect(mol,2,nBits=MORGAN_SIZE)
for i in range(tempReturn.shape[0]):
tempReturn[i] = vec[i]
return tempReturn
else:
return np.zeros(MORGAN_SIZE)
def get_fasta(fasta_name, training_data):
training_data['sequence'] = None
file = open(fasta_name)
index = 0
seq = ''
for line in file:
if line.startswith(">"):
if index >= 1:
training_data['sequence'][training_data['target_id'] == name] = seq
print(index,name,seq[:10])
seq = ''
name = line[4:10]
index = index + 1
else:
seq = seq + line[:-1]
return training_data
# # Data Prepreprocessing
training_data = pd.read_csv('PDR_SMILES.csv')
training_data.head()
# ,'Scientific name'
data3 = pd.read_csv("PDR_SMILES.csv")
# data3[data3 == '*'] = np.nan
data3 = pd.concat([data3[data3['IN or ON matrix'] == 'O'], data3[data3['IN or ON matrix'] == 'I']])
data3 = pd.concat([data3[data3['Field or Undercover'] == 'F'], data3[data3['Field or Undercover'] == 'U']])
data4 = data3[['Plant','Scientific name','Study location','Matrix','IN or ON matrix','Field or Undercover','SMILES','Min (days)','Max (days)','Mean (days)']].copy()
remained_index = data4[['Plant','Scientific name','Study location','Matrix','IN or ON matrix','Field or Undercover','SMILES']].drop_duplicates(keep = False ).index
Data_properties = data4[['Plant','Scientific name','Study location','Matrix','IN or ON matrix','Field or Undercover']].loc[remained_index]
smiles = data4['SMILES'].loc[remained_index]
Target_properties = data4[['Min (days)','Max (days)','Mean (days)']].loc[remained_index]
data5 = Data_properties.copy()
# data5["Type"] = data5["IN or ON matrix"].map(str) + data5["Field or Undercover"].map(str)
# types = np.argmax(pd.get_dummies(data5['Type']).values,axis = 1)
types = data4['Mean (days)'].loc[remained_index].copy()
types = types.astype('int32').copy()
types[types <16] = 0
types2 = types.copy()
types2[types2 >= 60] = 0
types[types2 !=0] = 1
types[types >= 60] = 2
types.shape
radius = 6
MORGAN_SIZE = 2048
SMILES_MORGAN = np.zeros((smiles.shape[0], MORGAN_SIZE), dtype=np.int8)
for ind, smi in enumerate(smiles):
SMILES_MORGAN[ind] = smi_to_morganfingerprint(smi, radius, MORGAN_SIZE)
# SMILES_MORGAN[SMILES_MORGAN == 0] = -1
SMILES_MORGAN.shape
Data_properties['Study location'].unique()
np.save("PDR_SMILES_MORGAN_2048_6.npy", SMILES_MORGAN)
np.save("PDR_Target_properties.npy", Target_properties.values)
np.save("PDR_types.npy", types)
Data_properties.head()
pd.get_dummies(Data_properties).head()
np.save("PDR_properties.npy", pd.get_dummies(Data_properties).values)
# # Training
# +
X_list = np.concatenate((np.load('PDR_SMILES_MORGAN_2048_6.npy'),np.load('PDR_properties.npy')),axis = -1)
# Y_list = np.log10(np.load('PDR_Target_properties.npy'))
Y_list = np.load('PDR_Target_properties.npy')
types = np.load('PDR_types.npy')
X_list.shape,Y_list.shape,types.shape
# -
for i in range(len(np.unique(types))):
print(len(types[types == i]))
def pearson_r(y_true, y_pred):
x = y_true
y = y_pred
mx = K.mean(x, axis=0)
my = K.mean(y, axis=0)
xm, ym = x - mx, y - my
r_num = K.sum(xm * ym)
x_square_sum = K.sum(xm * xm)
y_square_sum = K.sum(ym * ym)
r_den = K.sqrt(x_square_sum * y_square_sum)
r = r_num / r_den
return K.mean(r)
def root_mean_squared_error(y_true, y_pred):
return K.sqrt(K.mean(K.square(y_pred - y_true)))
def root_mean_squared_error_loss(y_true, y_pred):
X = 10**(-y_pred)
Y = 10**(-y_true)
return K.sqrt(K.mean(K.square(X - Y)))
# +
def dot_product(x, kernel):
if K.backend() == 'tensorflow':
return K.squeeze(K.dot(x, K.expand_dims(kernel)), axis=-1)
else:
return K.dot(x, kernel)
class AttentionWithContext(Layer):
def __init__(self,
W_regularizer=None, u_regularizer=None, b_regularizer=None,
W_constraint=None, u_constraint=None, b_constraint=None,
bias=True, **kwargs):
self.supports_masking = True
self.init = initializers.get('glorot_uniform')
self.W_regularizer = regularizers.get(W_regularizer)
self.u_regularizer = regularizers.get(u_regularizer)
self.b_regularizer = regularizers.get(b_regularizer)
self.W_constraint = constraints.get(W_constraint)
self.u_constraint = constraints.get(u_constraint)
self.b_constraint = constraints.get(b_constraint)
self.bias = bias
super(AttentionWithContext, self).__init__(**kwargs)
def build(self, input_shape):
assert len(input_shape) == 3
self.W = self.add_weight((input_shape[-1], input_shape[-1],),
initializer=self.init,
name='{}_W'.format(self.name),
regularizer=self.W_regularizer,
constraint=self.W_constraint)
if self.bias:
self.b = self.add_weight((input_shape[-1],),
initializer='zero',
name='{}_b'.format(self.name),
regularizer=self.b_regularizer,
constraint=self.b_constraint)
self.u = self.add_weight((input_shape[-1],),
initializer=self.init,
name='{}_u'.format(self.name),
regularizer=self.u_regularizer,
constraint=self.u_constraint)
super(AttentionWithContext, self).build(input_shape)
def compute_mask(self, input, input_mask=None):
return None
def call(self, x, mask=None):
uit = dot_product(x, self.W)
if self.bias:
uit += self.b
uit = K.tanh(uit)
ait = dot_product(uit, self.u)
a = K.exp(ait)
if mask is not None:
a *= K.cast(mask, K.floatx())
a /= K.cast(K.sum(a, axis=1, keepdims=True) + K.epsilon(), K.floatx())
a = K.expand_dims(a)
weighted_input = x * a
return K.sum(weighted_input, axis=1)
def compute_output_shape(self, input_shape):
return input_shape[0], input_shape[-1]
# -
from sklearn.model_selection import train_test_split
x_train_from_train, x_val_from_train, y_train_from_train, y_val_from_train = train_test_split(X_list, Y_list, test_size=0.2, random_state=random_seed, stratify = types)
x_train_from_train.shape, x_val_from_train.shape, y_train_from_train.shape, y_val_from_train.shape
# +
#model structure
model_name = 'ACTHON_model_2048_6'
auxiliary_input1 = Input(shape=(3094,), dtype='float32', name='main_input')
x = Dense(1524)(auxiliary_input1)
x = BatchNormalization()(x)
x = LeakyReLU(alpha=0.3)(x)
r = Dropout(0.2)(x)
x = Dense(768)(x)
x = BatchNormalization()(x)
x = LeakyReLU(alpha=0.3)(x)
r = Dropout(0.2)(x)
x = Dense(384)(x)
x = BatchNormalization()(x)
x = LeakyReLU(alpha=0.3)(x)
r = Dropout(0.2)(x)
x = Dense(192)(x)
x = BatchNormalization()(x)
x = LeakyReLU(alpha=0.3)(x)
r = Dropout(0.2)(x)
x = Dense(96)(x)
x = BatchNormalization()(x)
x = LeakyReLU(alpha=0.3)(x)
r = Dropout(0.2)(x)
x = Dense(48)(x)
x = BatchNormalization()(x)
x = LeakyReLU(alpha=0.3)(x)
r = Dropout(0.2)(x)
x = Dense(24)(x)
x = BatchNormalization()(x)
x = LeakyReLU(alpha=0.3)(x)
r = Dropout(0.2)(x)
x = Dense(12)(x)
x = BatchNormalization()(x)
x = LeakyReLU(alpha=0.3)(x)
r = Dropout(0.2)(x)
x = Dense(6)(x)
x = BatchNormalization()(x)
x = LeakyReLU(alpha=0.3)(x)
x = Dropout(0.2)(x)
main_output = Dense(3,activation='relu')(x)
model = Model(inputs=auxiliary_input1, outputs=main_output)
print(model.summary())
opt = keras.optimizers.Adam()
model.compile(loss=root_mean_squared_error,
optimizer=opt,
metrics=[pearson_r])
checkpointer = ModelCheckpoint(model_name, verbose=1, save_best_only=True)
# -
model_history = model.fit(x_train_from_train, np.log10(y_train_from_train),
batch_size=batch_size, epochs=100, verbose=1, callbacks=[checkpointer],
validation_data=(x_val_from_train, np.log10(y_val_from_train)), class_weight = 'auto')
file = open(model_name+'_history.pickle', 'wb')
pickle.dump(model.history.history, file)
file.close()
# +
"""
@author: <NAME>
"""
import numpy as np
import copy
from math import sqrt
from scipy import stats
from sklearn import preprocessing,metrics
def rmse(y,f):
"""
Task: To compute root mean squared error (RMSE)
Input: y Vector with original labels (pKd [M])
f Vector with predicted labels (pKd [M])
Output: rmse RSME
"""
rmse = sqrt(((y - f)**2).mean(axis=0))
return rmse
def pearson(y,f):
"""
Task: To compute Pearson correlation coefficient
Input: y Vector with original labels (pKd [M])
f Vector with predicted labels (pKd [M])
Output: rp Pearson correlation coefficient
"""
rp = np.corrcoef(y, f)[0,1]
return rp
def spearman(y,f):
"""
Task: To compute Spearman's rank correlation coefficient
Input: y Vector with original labels (pKd [M])
f Vector with predicted labels (pKd [M])
Output: rs Spearman's rank correlation coefficient
"""
rs = stats.spearmanr(y, f)[0]
return rs
def ci(y,f):
"""
Task: To compute concordance index (CI)
Input: y Vector with original labels (pKd [M])
f Vector with predicted labels (pKd [M])
Output: ci CI
References:
[1] <NAME>, <NAME>, <NAME>, <NAME>,
<NAME>, JingTang and <NAME>.
Toward more realistic drug-target interaction predictions.
Briefings in Bioinformatics, 16, pages 325-337, 2014.
"""
ind = np.argsort(y)
y = y[ind]
f = f[ind]
i = len(y)-1
j = i-1
z = 0.0
S = 0.0
while i > 0:
while j >= 0:
if y[i] > y[j]:
z = z+1
u = f[i] - f[j]
if u > 0:
S = S + 1
elif u == 0:
S = S + 0.5
j = j - 1
i = i - 1
j = i-1
ci = S/z
return ci
def weighted_f1(y,f):
"""
Task: To compute F1 score using the threshold of 7 M
to binarize pKd's into true class labels.
Input: y Vector with original labels (pKd [M])
f Vector with predicted labels (pKd [M])
Output: f1 F1 score
"""
y_binary = y.astype('int32').copy()
y_binary[y_binary <16] = 0
y_binary2 = y_binary.copy()
y_binary2[y_binary2 >= 60] = 0
y_binary[y_binary2 !=0] = 1
y_binary[y_binary >= 60] = 2
f_binary = f.astype('int32').copy()
f_binary[f_binary <16] = 0
f_binary2 = f_binary.copy()
f_binary2[f_binary2 >= 60] = 0
f_binary[f_binary2 !=0] = 1
f_binary[f_binary >= 60] = 2
f1 = metrics.f1_score(y_binary, f_binary, average='weighted')
return f1
def average_AUC(y,f):
thr = np.linspace(16,60,10)
auc = np.empty(np.shape(thr)); auc[:] = np.nan
for i in range(len(thr)):
y_binary = copy.deepcopy(y)
y_binary = preprocessing.binarize(y_binary.reshape(1,-1), threshold=thr[i], copy=False)[0]
fpr, tpr, thresholds = metrics.roc_curve(y_binary, f, pos_label=1)
auc[i] = metrics.auc(fpr, tpr)
avAUC = np.mean(auc)
# y_binary = y.astype('int32').copy()
# y_binary[y_binary <16] = 0
# y_binary2 = y_binary.copy()
# y_binary2[y_binary2 >= 60] = 0
# y_binary[y_binary2 !=0] = 1
# y_binary[y_binary >= 60] = 2
# f_binary = f.astype('int32').copy()
# f_binary[f_binary <16] = 0
# f_binary2 = f_binary.copy()
# f_binary2[f_binary2 >= 60] = 0
# f_binary[f_binary2 !=0] = 1
# f_binary[f_binary >= 60] = 2
# avAUC = 0
# for i in range(3):
# fpr, tpr, thresholds = metrics.roc_curve(y_binary, f_binary, pos_label=i)
# avAUC = avAUC + metrics.auc(fpr, tpr)
# avAUC = avAUC/3
return avAUC
def accuracy(y,f):
"""
Task: To compute F1 score using the threshold of 7 M
to binarize pKd's into true class labels.
Input: y Vector with original labels (pKd [M])
f Vector with predicted labels (pKd [M])
Output: f1 F1 score
"""
y_binary = y.astype('int32').copy()
y_binary[y_binary <16] = 0
y_binary2 = y_binary.copy()
y_binary2[y_binary2 >= 60] = 0
y_binary[y_binary2 !=0] = 1
y_binary[y_binary >= 60] = 2
f_binary = f.astype('int32').copy()
f_binary[f_binary <16] = 0
f_binary2 = f_binary.copy()
f_binary2[f_binary2 >= 60] = 0
f_binary[f_binary2 !=0] = 1
f_binary[f_binary >= 60] = 2
accuracy = metrics.accuracy_score(y_binary, f_binary)
return accuracy
# -
# reload a file to a variable
for i in range(10):
with open(model_name+'_history.pickle', 'rb') as file:
historyList = pickle.load(file)
# %matplotlib inline
BIGGER_SIZE = 20
plt.figure(figsize=(25, 25))
plt.plot(historyList['loss'], label='Train loss')
plt.plot(historyList['val_loss'], label='Val loss')
plt.xlabel('Epochs', size=BIGGER_SIZE)
plt.ylabel('Loss', size=BIGGER_SIZE)
plt.xticks(fontsize=BIGGER_SIZE)
plt.yticks(fontsize=BIGGER_SIZE)
# plt.ylim((-0.1, 4))
plt.title(model_name, size=BIGGER_SIZE)
plt.legend()
BIGGER_SIZE = 20
plt.figure(figsize=(25, 25))
plt.plot(historyList['pearson_r'], label='Train pearson correlation')
plt.plot(historyList['val_pearson_r'], label='Val pearson correlation')
plt.xlabel('Epochs', size=BIGGER_SIZE)
plt.ylabel('Loss', size=BIGGER_SIZE)
plt.xticks(fontsize=BIGGER_SIZE)
plt.yticks(fontsize=BIGGER_SIZE)
# plt.ylim((0.25, 0.75))
plt.title(model_name, size=BIGGER_SIZE)
plt.legend()
model.load_weights(model_name)
predict = 10**(model.predict(x_val_from_train))
true = y_val_from_train
print('rmse: ',rmse(true[:,2],predict[:,2]))
print('pearson: ',pearson(true[:,2],predict[:,2]))
print('spearman: ',spearman(true[:,2],predict[:,2]))
print('ci: ',ci(true[:,2],predict[:,2]))
print('f1: ',f1(true[:,2],predict[:,2]))
print('average_AUC: ',average_AUC(true[:,2],predict[:,2]))
print('accuracy: ',accuracy(true[:,2],predict[:,2]))
# %matplotlib inline
import matplotlib.pyplot as plt
BIGGER_SIZE = 20
plt.figure(figsize=(25, 25))
plt.scatter(true[:,2],predict[:,2])
plt.xlabel('true', size=BIGGER_SIZE)
plt.ylabel('predict', size=BIGGER_SIZE)
plt.xticks(fontsize=BIGGER_SIZE)
plt.xlim((-0.1, 100))
plt.yticks(fontsize=BIGGER_SIZE)
plt.ylim((-0.1, 100))
plt.legend()
plt.show()
model.load_weights(model_name)
predict_train = 10**(model.predict(x_train_from_train))
true_train = y_train_from_train
print('rmse: ',rmse(true_train[:,2],predict_train[:,2]))
print('pearson: ',pearson(true_train[:,2],predict_train[:,2]))
print('spearman: ',spearman(true_train[:,2],predict_train[:,2]))
print('ci: ',ci(true_train[:,2],predict_train[:,2]))
print('f1: ',f1(true_train[:,2],predict_train[:,2]))
print('average_AUC: ',average_AUC(true_train[:,2],predict_train[:,2]))
print('accuracy: ',accuracy(true_train[:,2],predict_train[:,2]))
# %matplotlib inline
import matplotlib.pyplot as plt
BIGGER_SIZE = 20
plt.figure(figsize=(25, 25))
plt.scatter(true_train[:,2],predict_train[:,2])
plt.xlabel('true', size=BIGGER_SIZE)
plt.ylabel('predict', size=BIGGER_SIZE)
plt.xticks(fontsize=BIGGER_SIZE)
plt.xlim((-0.1, 100))
plt.yticks(fontsize=BIGGER_SIZE)
plt.ylim((-0.1, 100))
plt.legend()
plt.show()
| PrecisionFuturePesticide.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pytube
from pytube import YouTube
link=input('Enter Your Youtube Link')
yt=pytube.YouTube(link)
video=yt.streams.first()
video.download('D:/')
| Download Youtube Videos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="vnNVFgX6Ehei"
import numpy as np
np.random.seed(10)
import tensorflow as tf
tf.random.set_seed(10)
import matplotlib.pyplot as plt
tf.keras.backend.set_floatx('float64')
# Load MNIST data set
(train_images, _), (test_images, _) = tf.keras.datasets.mnist.load_data()
train_images = train_images.reshape(-1,28*28)/255
test_images = test_images.reshape(-1,28*28)/255
np.random.shuffle(train_images)
# + id="D5_4j0W7Et97"
class vi_nf_model(tf.keras.Model):
def __init__(self,num_latent,num_flow_iters):
super(vi_nf_model,self).__init__()
# Constants
self.num_latent = num_latent
self.num_flow_iters = num_flow_iters
# Initializer
xavier = tf.keras.initializers.GlorotUniform()
# Define inference architecture
self.l1 = tf.keras.layers.Dense(150,activation='swish')
self.l2 = tf.keras.layers.Dense(100,activation='swish')
self.l3 = tf.keras.layers.Dense(50,activation='swish')
# Variational stuff
self.zz_mu = tf.keras.layers.Dense(num_latent,activation='swish')
self.zz_logvar = tf.keras.layers.Dense(num_latent,activation='swish')
# Normalizing flow stuff
self.ww_list = []
self.uu_list = []
self.bb_list = []
for _ in range(num_flow_iters):
self.ww_list.append(xavier(shape=(num_latent,1),dtype='float64'))
self.uu_list.append(xavier(shape=(num_latent,1),dtype='float64'))
self.bb_list.append(xavier(shape=(1,),dtype='float64'))
# Define reconstruction architecture
self.l5 = tf.keras.layers.Dense(50,activation='swish')
self.l6 = tf.keras.layers.Dense(100,activation='swish')
self.l7 = tf.keras.layers.Dense(150,activation='swish')
self.l8 = tf.keras.layers.Dense(28*28,activation='sigmoid')
# Optimizer
self.train_op = tf.keras.optimizers.Adam(learning_rate=0.001)
self.bce = tf.keras.losses.BinaryCrossentropy()
def call(self,X):
# Call inference
h1 = self.l1(X)
h2 = self.l2(h1)
h3 = self.l3(h2)
# Latent space quantities
self.mu = self.zz_mu(h3)
self.logvar = self.zz_logvar(h3)
# Shapes
batch_size = tf.shape(self.mu)[0]
nl = self.num_latent
# Batch wise latent space operations
eps = tf.random.normal(shape=(batch_size,nl),mean=0.0,stddev=1.0,dtype='float64')
self.Z = self.mu + tf.math.exp(0.5*self.logvar)*eps
# Find density value of initial sample
self.logZ0 = -0.5*(tf.cast(tf.math.log(2.0*np.pi),dtype='float64') - self.logvar) - (0.5*(self.Z-self.mu)**2)/(2.0*tf.exp(self.logvar))
# Use normalizing flow and get energy (needed for loss function)
self.Zk, flow_energy = self.normalizing_flow(self.Z,self.logZ0)
# Decode
h5 = self.l5(self.Zk)
h6 = self.l6(h5)
h7 = self.l7(h6)
out = self.l8(h7)
return out, flow_energy
def normalizing_flow(self,Z,logstart):
# Number of transformations
Kval = self.num_flow_iters
zprev = Z
for i in range(Kval):
w_i = self.ww_list[i]
u_i = self.uu_list[i]
b_i = self.bb_list[i]
wz = tf.matmul(zprev,w_i)
tt = tf.math.tanh(wz+b_i)
dtt = 1.0-(tf.math.tanh(wz+b_i))**2
zprev = zprev + tf.matmul(tt,tf.transpose(u_i))
phi = tf.matmul(tt,tf.transpose(w_i))
utphi = tf.matmul(phi,u_i)
logstart = logstart - tf.math.log(tf.math.abs(1.0+utphi))
return zprev, logstart
def sample_latent(self,X,num_samples):
sample_list = []
for sample in range(num_samples):
sample_list.append(self.call(X)[0].numpy())
return sample_list
def get_loss(self,X):
Ypred, flow_energy = self.call(X)
return self.bce(X,Ypred) + tf.reduce_mean(flow_energy)
def get_grad(self,X):
with tf.GradientTape() as tape:
tape.watch(self.trainable_variables)
L = self.get_loss(X)
g = tape.gradient(L, self.trainable_variables)
return g
# perform gradient descent - regular
def network_learn(self,X):
g = self.get_grad(X)
self.train_op.apply_gradients(zip(g, self.trainable_variables))
# Train the model
def train_model(self,Xtrain):
plot_iter = 0
stop_iter = 0
patience = 10
best_loss = 999999.0 # Some large number
self.num_batches = 10
self.train_batch_size = int(Xtrain.shape[0]/self.num_batches)
for i in range(200):
# Training loss
print('Training iteration:',i)
loss = 0.0
for batch in range(self.num_batches):
input_batch = Xtrain[batch*self.train_batch_size:(batch+1)*self.train_batch_size]
self.network_learn(input_batch)
batch_loss = self.get_loss(input_batch).numpy()
loss+=batch_loss
print('Batch loss:',batch_loss)
loss = loss/self.num_batches
print('Epoch loss:',loss)
# Check early stopping criteria
if loss < best_loss:
print('Improved loss from:',best_loss,' to:', loss)
best_loss = loss
self.save_weights('./checkpoints/my_checkpoint')
stop_iter = 0
else:
print('Loss (no improvement):',loss)
stop_iter = stop_iter + 1
if stop_iter == patience:
break
def test_model(self,Xtest):
# Check accuracy on test
print('Test loss:',self.get_loss(Xtest).numpy())
# Load weights
def restore_model(self):
self.load_weights('./checkpoints/my_checkpoint') # Load pretrained model
# + id="3MeupfDlEv5V"
model = vi_nf_model(40,10)
# + colab={"base_uri": "https://localhost:8080/", "height": 513} id="_KaM8m-K09aQ" outputId="86be3469-c095-415b-bc76-7e4794386fef"
# Sample before training
sample = 1000
num_samples = 100
sample_list = np.asarray(model.sample_latent(train_images[sample:sample+1],num_samples))[:,0,:]
plt.figure()
plt.imshow(np.mean(sample_list,axis=0).reshape(28,28))
plt.colorbar()
plt.show()
plt.figure()
plt.imshow(np.std(sample_list,axis=0).reshape(28,28))
plt.colorbar()
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="CJZwGFfzEx5Z" outputId="60a49725-04b9-4bc8-91b8-6707128bdd40"
model.train_model(train_images)
# + colab={"base_uri": "https://localhost:8080/", "height": 513} id="cpljGJAiE0Bz" outputId="848ff4cc-0ce0-42f2-ca78-b1d906f05da6"
# Sample after training
sample = 1000
num_samples = 100
sample_list = np.asarray(model.sample_latent(train_images[sample:sample+1],num_samples))[:,0,:]
plt.figure()
plt.imshow(np.mean(sample_list,axis=0).reshape(28,28))
plt.colorbar()
plt.show()
plt.figure()
plt.imshow(np.std(sample_list,axis=0).reshape(28,28))
plt.colorbar()
plt.show()
# + id="lKyAIpkZIc4Z" colab={"base_uri": "https://localhost:8080/", "height": 513} outputId="c03fadb7-04ae-418c-f668-e08086ea327c"
# Sample after training
sample = 0
num_samples = 100
sample_list = np.asarray(model.sample_latent(train_images[sample:sample+1],num_samples))[:,0,:]
plt.figure()
plt.imshow(np.mean(sample_list,axis=0).reshape(28,28))
plt.colorbar()
plt.show()
plt.figure()
plt.imshow(np.std(sample_list,axis=0).reshape(28,28))
plt.colorbar()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 513} id="_VthYWr73r4V" outputId="57350096-cf81-477a-e749-0ed75b33299b"
# Sample after training
sample = 50
num_samples = 100
sample_list = np.asarray(model.sample_latent(train_images[sample:sample+1],num_samples))[:,0,:]
plt.figure()
plt.imshow(np.mean(sample_list,axis=0).reshape(28,28))
plt.colorbar()
plt.show()
plt.figure()
plt.imshow(np.std(sample_list,axis=0).reshape(28,28))
plt.colorbar()
plt.show()
# + id="kTVm_lPD5O3A"
| Generative_Models/VI_NF/VI_NF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python3
# ---
# # Classification data using scikit-learn
#
#
# Classification problems are those in which the feature to be predicted contains categories of values. Each of these categories are considered as a class into which the predicted value will fall into and hence has its name, classification.
#
# In this notebook, we'll use scikit-learn to predict classes. scikit-learn provides implementations of many classification algorithms. In here, we have done a comparative study of 5 different classification algorithms.
#
# To help visualize what we are doing, we'll use 2D and 3D charts to show how the classes look (with 3 selected dimensions) with matplotlib and scikitplot python libraries.
#
#
# <a id="top"></a>
# ## Table of Contents
#
# 1. [Load libraries](#load_libraries)
# 2. [Data exploration](#explore_data)
# 3. [Prepare data for building classification model](#prepare_data)
# 4. [Split data into train and test sets](#split_data)
# 5. [Helper methods for graph generation](#helper_methods)
# 6. [Build Naive Bayes classification model](#model_nb)
# 7. [Build Logistic Regression classification model](#model_lrc)
# 8. [Build K-Nearest classification model](#model_knn)
# 9. [Build Kernel SVM classification model](#model_svm)
# 10. [Build Random Forest classification model](#model_rfc)
# 11. [Comparative study of different classification algorithms](#compare_classification)
# ### Quick set of instructions to work through the notebook
#
# If you are new to Notebooks, here's a quick overview of how to work in this environment.
#
# 1. The notebook has 2 types of cells - markdown (text) such as this and code such as the one below.
# 2. Each cell with code can be executed independently or together (see options under the Cell menu). When working in this notebook, we will be running one cell at a time because we need to make code changes to some of the cells.
# 3. To run the cell, position cursor in the code cell and click the Run (arrow) icon. The cell is running when you see the * next to it. Some cells have printable output.
# 4. Work through this notebook by reading the instructions and executing code cell by cell. Some cells will require modifications before you run them.
# <a id="load_libraries"></a>
# ## 1. Load libraries
# [Top](#top)
#
# Install python modules
# NOTE! Some pip installs require a kernel restart.
# The shell command pip install is used to install Python modules. Some installs require a kernel restart to complete. To avoid confusing errors, run the following cell once and then use the Kernel menu to restart the kernel before proceeding.
# !pip install pandas==0.24.2
# !pip install --user pandas_ml==0.6.1
#downgrade matplotlib to bypass issue with confusion matrix being chopped out
# !pip install matplotlib==3.1.0
# !pip install --user scikit-learn==0.21.3
# !pip install -q scikit-plot
# +
import sys
import io
from sklearn.preprocessing import LabelEncoder, OneHotEncoder
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import StandardScaler
from sklearn.compose import ColumnTransformer, make_column_transformer
from sklearn.pipeline import Pipeline
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score, classification_report
import pandas as pd, numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.colors as mcolors
import matplotlib.patches as mpatches
import scikitplot as skplt
# -
# <a id="explore_data"></a>
# ## 2. Data exploration
# [Top](#top)
#
# Data can be easily loaded within IBM Watson Studio. Instructions to load data within IBM Watson Studio can be found [here](https://ibmdev1.rtp.raleigh.ibm.com/tutorials/watson-studio-using-jupyter-notebook/). The data set can be located by its name and inserted into the notebook as a pandas DataFrame as shown below.
#
# 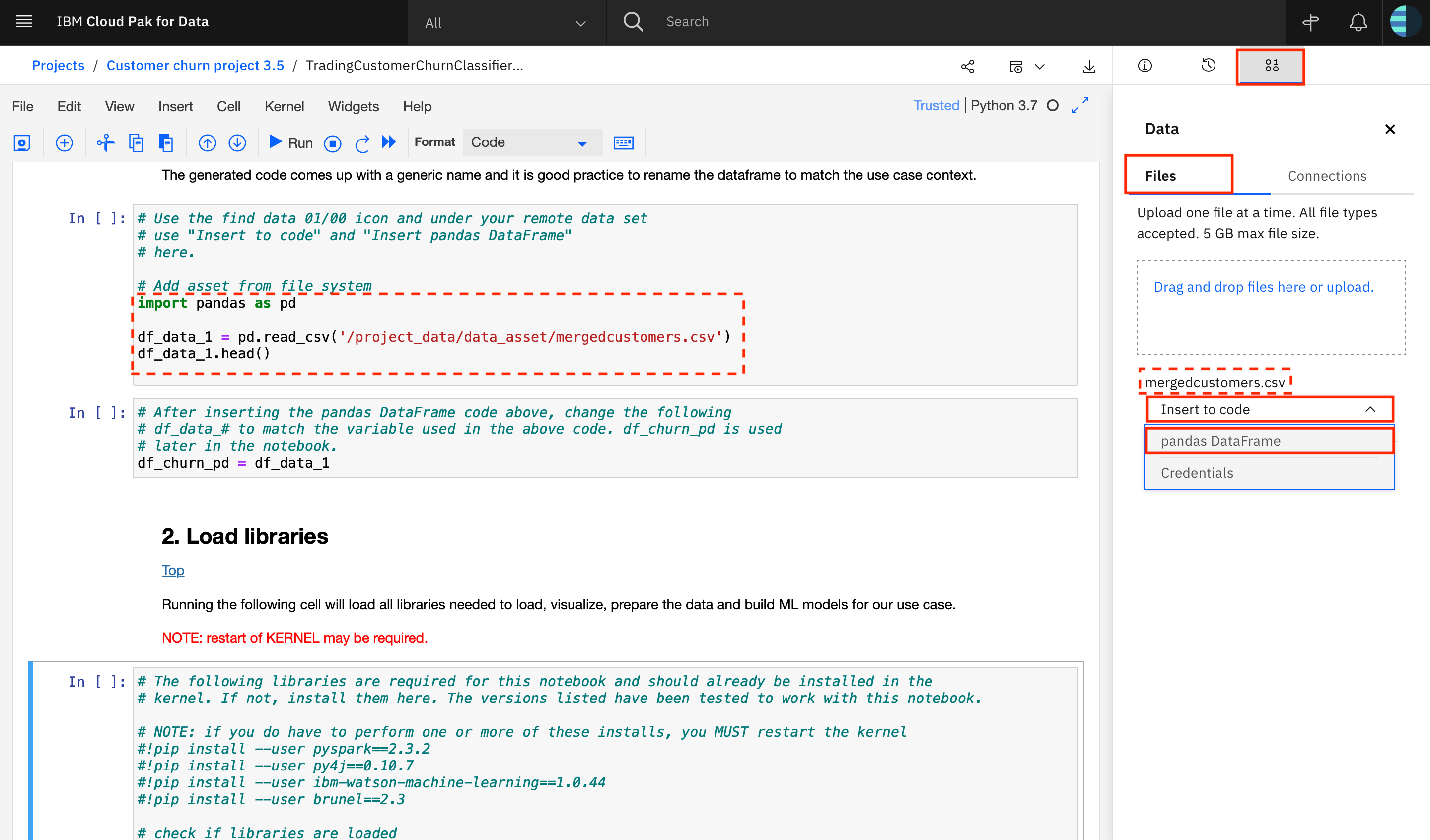
#
# The generated code comes up with a generic name and it is good practice to rename the dataframe to match the use case context.
df_churn_pd = pd.read_csv("https://raw.githubusercontent.com/IBM/ml-learning-path-assets/master/data/mergedcustomers_missing_values_GENDER.csv")
df_churn_pd.head()
# +
print("The dataset contains columns of the following data types : \n" +str(df_churn_pd.dtypes))
# -
#notice that Gender has three missing values. This will be handled in one of the preprocessing steps that is to follow.
print("The dataset contains following number of records for each of the columns : \n" +str(df_churn_pd.count()))
print( "Each category within the churnrisk column has the following count : ")
print(df_churn_pd.groupby(['CHURNRISK']).size())
#bar chart to show split of data
index = ['High','Medium','Low']
churn_plot = df_churn_pd['CHURNRISK'].value_counts(sort=True, ascending=False).plot(kind='bar',figsize=(4,4),title="Total number for occurences of churn risk " + str(df_churn_pd['CHURNRISK'].count()), color=['#BB6B5A','#8CCB9B','#E5E88B'])
churn_plot.set_xlabel("Churn Risk")
churn_plot.set_ylabel("Frequency")
# <a id="prepare_data"></a>
# ## 3. Data preparation
# [Top](#top)
#
# Data preparation is a very important step in machine learning model building. This is because the model can perform well only when the data it is trained on is good and well prepared. Hence, this step consumes the bulk of a data scientist's time spent building models.
#
# During this process, we identify categorical columns in the dataset. Categories needed to be indexed, which means the string labels are converted to label indices. These label indices are encoded using One-hot encoding to a binary vector with at most a single-value indicating the presence of a specific feature value from among the set of all feature values. This encoding allows algorithms which expect continuous features to use categorical features.
#
# Final step in the data preparation process is to assemble all the categorical and non-categorical columns into a feature vector. We use VectorAssembler for this. VectorAssembler is a transformer that combines a given list of columns into a single vector column. It is useful for combining raw features and features generated by different feature transformers into a single feature vector, in order to train ML models.
# +
#remove columns that are not required
df_churn_pd = df_churn_pd.drop(['ID'], axis=1)
df_churn_pd.head()
# +
# Defining the categorical columns
categoricalColumns = ['GENDER', 'STATUS', 'HOMEOWNER']
print("Categorical columns : " )
print(categoricalColumns)
impute_categorical = SimpleImputer(strategy="most_frequent")
onehot_categorical = OneHotEncoder(handle_unknown='ignore')
categorical_transformer = Pipeline(steps=[('impute',impute_categorical),('onehot',onehot_categorical)])
# +
# Defining the numerical columns
numericalColumns = df_churn_pd.select_dtypes(include=[np.float,np.int]).columns
print("Numerical columns : " )
print(numericalColumns)
scaler_numerical = StandardScaler()
numerical_transformer = Pipeline(steps=[('scale',scaler_numerical)])
# +
preprocessorForCategoricalColumns = ColumnTransformer(transformers=[('cat', categorical_transformer, categoricalColumns)],
remainder="passthrough")
preprocessorForAllColumns = ColumnTransformer(transformers=[('cat', categorical_transformer, categoricalColumns),('num',numerical_transformer,numericalColumns)],
remainder="passthrough")
#. The transformation happens in the pipeline. Temporarily done here to show what intermediate value looks like
df_churn_pd_temp = preprocessorForCategoricalColumns.fit_transform(df_churn_pd)
print("Data after transforming :")
print(df_churn_pd_temp)
df_churn_pd_temp_2 = preprocessorForAllColumns.fit_transform(df_churn_pd)
print("Data after transforming :")
print(df_churn_pd_temp_2)
# +
# prepare data frame for splitting data into train and test datasets
features = []
features = df_churn_pd.drop(['CHURNRISK'], axis=1)
label_churn = pd.DataFrame(df_churn_pd, columns = ['CHURNRISK'])
label_encoder = LabelEncoder()
label = df_churn_pd['CHURNRISK']
label = label_encoder.fit_transform(label)
print("Encoded value of Churnrisk after applying label encoder : " + str(label))
# +
area = 75
x = df_churn_pd['ESTINCOME']
y = df_churn_pd['DAYSSINCELASTTRADE']
z = df_churn_pd['TOTALDOLLARVALUETRADED']
pop_a = mpatches.Patch(color='#BB6B5A', label='High')
pop_b = mpatches.Patch(color='#E5E88B', label='Medium')
pop_c = mpatches.Patch(color='#8CCB9B', label='Low')
def colormap(risk_list):
cols=[]
for l in risk_list:
if l==0:
cols.append('#BB6B5A')
elif l==2:
cols.append('#E5E88B')
elif l==1:
cols.append('#8CCB9B')
return cols
fig = plt.figure(figsize=(12,6))
fig.suptitle('2D and 3D view of churnrisk data')
# First subplot
ax = fig.add_subplot(1, 2,1)
ax.scatter(x, y, alpha=0.8, c=colormap(label), s= area)
ax.set_ylabel('DAYS SINCE LAST TRADE')
ax.set_xlabel('ESTIMATED INCOME')
plt.legend(handles=[pop_a,pop_b,pop_c])
# Second subplot
ax = fig.add_subplot(1,2,2, projection='3d')
ax.scatter(z, x, y, c=colormap(label), marker='o')
ax.set_xlabel('TOTAL DOLLAR VALUE TRADED')
ax.set_ylabel('ESTIMATED INCOME')
ax.set_zlabel('DAYS SINCE LAST TRADE')
plt.legend(handles=[pop_a,pop_b,pop_c])
plt.show()
# -
#
# <a id="split_data"></a>
# ## 4. Split data into test and train
# [Top](#top)
#
#
# +
X_train, X_test, y_train, y_test = train_test_split(features,label , random_state=0)
print("Dimensions of datasets that will be used for training : Input features"+str(X_train.shape)+
" Output label" + str(y_train.shape))
print("Dimensions of datasets that will be used for testing : Input features"+str(X_test.shape)+
" Output label" + str(y_test.shape))
# -
#
# <a id="helper_methods"></a>
# ## 5. Helper methods for graph generation
# [Top](#top)
#
#
# +
def colormap(risk_list):
cols=[]
for l in risk_list:
if l==0:
cols.append('#BB6B5A')
elif l==2:
cols.append('#E5E88B')
elif l==1:
cols.append('#8CCB9B')
return cols
def two_d_compare(y_test,y_pred,model_name):
#y_pred = label_encoder.fit_transform(y_pred)
#y_test = label_encoder.fit_transform(y_test)
area = (12 * np.random.rand(40))**2
plt.subplots(ncols=2, figsize=(10,4))
plt.suptitle('Actual vs Predicted data : ' +model_name + '. Accuracy : %.2f' % accuracy_score(y_test, y_pred))
plt.subplot(121)
plt.scatter(X_test['ESTINCOME'], X_test['DAYSSINCELASTTRADE'], alpha=0.8, c=colormap(y_test))
plt.title('Actual')
plt.legend(handles=[pop_a,pop_b,pop_c])
plt.subplot(122)
plt.scatter(X_test['ESTINCOME'], X_test['DAYSSINCELASTTRADE'],alpha=0.8, c=colormap(y_pred))
plt.title('Predicted')
plt.legend(handles=[pop_a,pop_b,pop_c])
plt.show()
x = X_test['TOTALDOLLARVALUETRADED']
y = X_test['ESTINCOME']
z = X_test['DAYSSINCELASTTRADE']
pop_a = mpatches.Patch(color='#BB6B5A', label='High')
pop_b = mpatches.Patch(color='#E5E88B', label='Medium')
pop_c = mpatches.Patch(color='#8CCB9B', label='Low')
def three_d_compare(y_test,y_pred,model_name):
fig = plt.figure(figsize=(12,10))
fig.suptitle('Actual vs Predicted (3D) data : ' +model_name + '. Accuracy : %.2f' % accuracy_score(y_test, y_pred))
ax = fig.add_subplot(121, projection='3d')
ax.scatter(x, y, z, c=colormap(y_test), marker='o')
ax.set_xlabel('TOTAL DOLLAR VALUE TRADED')
ax.set_ylabel('ESTIMATED INCOME')
ax.set_zlabel('DAYS SINCE LAST TRADE')
plt.legend(handles=[pop_a,pop_b,pop_c])
plt.title('Actual')
ax = fig.add_subplot(122, projection='3d')
ax.scatter(x, y, z, c=colormap(y_pred), marker='o')
ax.set_xlabel('TOTAL DOLLAR VALUE TRADED')
ax.set_ylabel('ESTIMATED INCOME')
ax.set_zlabel('DAYS SINCE LAST TRADE')
plt.legend(handles=[pop_a,pop_b,pop_c])
plt.title('Predicted')
plt.show()
def model_metrics(y_test,y_pred):
print("Decoded values of Churnrisk after applying inverse of label encoder : " + str(np.unique(y_pred)))
skplt.metrics.plot_confusion_matrix(y_test,y_pred,text_fontsize="small",cmap='Greens',figsize=(6,4))
plt.show()
#print("The classification report for the model : \n\n"+ classification_report(y_test, y_pred))
# -
# <a id="model_nb"></a>
# ## 6. Build Naive Bayes classification model
# [Top](#top)
#
# Applies the bayesian theorem to calculate the probabilty of a data point belonging to a particular class. Given the probability of certain related values,the formula to calculate the probabilty of an event B given event A to occur is calculated as follows.
# +
from sklearn.naive_bayes import MultinomialNB
model_name = 'Naive Bayes Classifier'
nbClassifier = MultinomialNB(alpha=1.0, class_prior=None, fit_prior=True)
nb_model = Pipeline(steps=[('preprocessor', preprocessorForCategoricalColumns),('classifier', nbClassifier)])
nb_model.fit(X_train,y_train)
y_pred_nb= nb_model.predict(X_test)
# -
two_d_compare(y_test,y_pred_nb,model_name)
y_test = label_encoder.inverse_transform(y_test)
y_pred_nb = label_encoder.inverse_transform(y_pred_nb)
model_metrics(y_test,y_pred_nb)
# <a id="model_lrc"></a>
# ## 7. Build Logistic Regression classification model
# [Top](#top)
#
# A logistic function is applied to the outcome of linear regression. The logistic function is also referred to as sigmoid function. This outputs a value between 0 and 1. We then select a line that depends on the use case, and any data point with probability value above the line is classified into the class represented by 1 and the data point below the line is classified into the class represented by 0.
# +
from sklearn.linear_model import LogisticRegression
model_name = "Logistic Regression Classifier"
logisticRegressionClassifier = LogisticRegression(random_state=0,multi_class='auto',solver='lbfgs',max_iter=1000)
lrc_model = Pipeline(steps=[('preprocessor', preprocessorForCategoricalColumns),
('classifier', logisticRegressionClassifier)])
lrc_model.fit(X_train,y_train)
y_pred_lrc = lrc_model.predict(X_test)
# -
y_test = label_encoder.transform(y_test)
two_d_compare(y_test,y_pred_lrc,model_name)
y_test = label_encoder.inverse_transform(y_test)
y_pred_lrc = label_encoder.inverse_transform(y_pred_lrc)
model_metrics(y_test,y_pred_lrc)
# <a id="model_knn"></a>
# ## 8. Build K-Nearest classification model
# [Top](#top)
#
# K number of nearest points around the data point to be predeicted are taken into consideration. These K points at this time, already belong to a class. The data point under consideration, is said to belong to the class with which most number of points from these k points belong to.
# +
from sklearn.neighbors import KNeighborsClassifier
model_name = "K-Nearest Neighbor Classifier"
knnClassifier = KNeighborsClassifier(n_neighbors = 5, metric='minkowski', p=2)
knn_model = Pipeline(steps=[('preprocessorAll',preprocessorForAllColumns),('classifier', knnClassifier)])
knn_model.fit(X_train,y_train)
y_pred_knn = knn_model.predict(X_test)
# -
y_test = label_encoder.transform(y_test)
two_d_compare(y_test,y_pred_knn,model_name)
y_test = label_encoder.inverse_transform(y_test)
y_pred_knn = label_encoder.inverse_transform(y_pred_knn)
model_metrics(y_test,y_pred_knn)
# <a id="model_svm"></a>
# ## 9. Build Kernel SVM classification model
# [Top](#top)
#
# Support Vector Machines outputs an optimal line of separation between the classes based on the training data served as input. This line of separation is called a hyperplane in a multi dimensional environment. SVM takes outliers that lie pretty close to another class into consideration to derive this separating hyperplane. Once the model is constructed with this hyperplane, any new point to be predicted will now check to see which side of the hyperplane this values lies in.
# +
from sklearn.svm import SVC
model_name = 'Kernel SVM Classifier'
svmClassifier = SVC(kernel='rbf', gamma= 'auto')
svm_model = Pipeline(steps=[('preprocessorAll',preprocessorForAllColumns),('classifier', svmClassifier)])
svm_model.fit(X_train,y_train)
y_pred_svm = svm_model.predict(X_test)
# -
y_test = label_encoder.transform(y_test)
#y_pred_svm = label_encoder.transform(y_pred_svm)
two_d_compare(y_test,y_pred_svm,model_name)
y_test = label_encoder.inverse_transform(y_test)
y_pred_svm = label_encoder.inverse_transform(y_pred_svm)
model_metrics(y_test,y_pred_svm)
# <a id="model_rfc"></a>
# ## 10. Build Random Forest classification model
# [Top](#top)
# Decision tree algorithms are efficient in eliminating columns that don't add value in predicting the output and in some cases, we are even able to see how a prediction was derived by backtracking the tree. However, this algorithm doesn't perform individually when the trees are huge and are hard to interpret. Such models are often referred to as weak models. The model performance is however improvised by taking an average of several such decision trees derived from the subsets of the training data. This approach is called the Random Forest classification.
# +
from sklearn.ensemble import RandomForestClassifier
model_name = "Random Forest Classifier"
randomForestClassifier = RandomForestClassifier(n_estimators=100, max_depth=2,random_state=0)
rfc_model = Pipeline(steps=[('preprocessorAll',preprocessorForAllColumns),('classifier', randomForestClassifier)])
rfc_model.fit(X_train,y_train)
y_pred_rfc = rfc_model.predict(X_test)
# +
y_test = label_encoder.transform(y_test)
two_d_compare(y_test,y_pred_rfc,model_name)
#three_d_compare(y_test,y_pred_rfc,model_name)
# -
y_test = label_encoder.inverse_transform(y_test)
y_pred_rfc = label_encoder.inverse_transform(y_pred_rfc)
model_metrics(y_test,y_pred_rfc)
#
# <a id="compare_classification"></a>
# ## 11. Comparative study of different classification algorithms.
# [Top](#top)
#
# In the bar chart below, we have compared the different classification algorithm against the actual values.
#
# +
uniqueValues, occurCount = np.unique(y_test, return_counts=True)
frequency_actual = (occurCount[0],occurCount[2],occurCount[1])
uniqueValues, occurCount = np.unique(y_pred_knn, return_counts=True)
frequency_predicted_knn = (occurCount[0],occurCount[2],occurCount[1])
uniqueValues, occurCount = np.unique(y_pred_rfc, return_counts=True)
frequency_predicted_rfc = (occurCount[0],occurCount[2],occurCount[1])
uniqueValues, occurCount = np.unique(y_pred_lrc, return_counts=True)
frequency_predicted_lrc = (occurCount[0],occurCount[2],occurCount[1])
uniqueValues, occurCount = np.unique(y_pred_svm, return_counts=True)
frequency_predicted_svm = (occurCount[0],occurCount[2],occurCount[1])
uniqueValues, occurCount = np.unique(y_pred_nb, return_counts=True)
frequency_predicted_nb = (occurCount[0],occurCount[2],occurCount[1])
n_groups = 3
fig, ax = plt.subplots(figsize=(10,5))
index = np.arange(n_groups)
bar_width = 0.1
opacity = 0.8
rects1 = plt.bar(index, frequency_actual, bar_width,
alpha=opacity,
color='g',
label='Actual')
rects2 = plt.bar(index + bar_width, frequency_predicted_nb, bar_width,
alpha=opacity,
color='pink',
label='Naive Bayesian - Predicted')
rects3 = plt.bar(index + bar_width*2, frequency_predicted_lrc, bar_width,
alpha=opacity,
color='y',
label='Logistic Regression - Predicted')
rects4 = plt.bar(index + bar_width*3, frequency_predicted_knn, bar_width,
alpha=opacity,
color='b',
label='K-Nearest Neighbor - Predicted')
rects5 = plt.bar(index + bar_width*4, frequency_predicted_svm, bar_width,
alpha=opacity,
color='red',
label='Kernel SVM - Predicted')
rects6 = plt.bar(index + bar_width*5, frequency_predicted_rfc, bar_width,
alpha=opacity,
color='purple',
label='Random Forest - Predicted')
plt.xlabel('Churn Risk')
plt.ylabel('Frequency')
plt.title('Actual vs Predicted frequency.')
plt.xticks(index + bar_width, ('High', 'Medium', 'Low'))
plt.legend()
plt.tight_layout()
plt.show()
# -
# In general, we see that the predictions around the Medium values is low on accuracy. One reason for that could be that the number of entries for the Medium values were much less represented than High and Low values. Further testing can be done by either increasing the number of Medium entries or involving several data fabrication techniques.
# <p><font size=-1 color=gray>
# © Copyright 2019 IBM Corp. All Rights Reserved.
# <p>
# Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file
# except in compliance with the License. You may obtain a copy of the License at
# https://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software distributed under the
# License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either
# express or implied. See the License for the specific language governing permissions and
# limitations under the License.
# </font></p>
| notebooks/classification_with_scikit-learn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] graffitiCellId="id_ow7e8a4"
# ### Problem Statement
#
# Given a linked list with integer data, arrange the elements in such a manner that all nodes with even numbers are placed after odd numbers. **Do not create any new nodes and avoid using any other data structure. The relative order of even and odd elements must not change.**
#
# **Example:**
# * `linked list = 1 2 3 4 5 6`
# * `output = 1 3 5 2 4 6`
# + graffitiCellId="id_7vmjo6u"
class Node:
def __init__(self, data):
self.data = data
self.next = None
# + [markdown] graffitiCellId="id_kefqagf"
# ### Exercise - Write the function definition here
#
# + graffitiCellId="id_hhuh98d"
def even_after_odd(head):
"""
:param - head - head of linked list
return - updated list with all even elements are odd elements
"""
pass
# + [markdown] graffitiCellId="id_xpuflcm"
# <span class="graffiti-highlight graffiti-id_xpuflcm-id_9q4n7o8"><i></i><button>Show Solution</button></span>
# + [markdown] graffitiCellId="id_m63s5ow"
# ### Test - Let's test your function
# + graffitiCellId="id_u1fjo92"
# helper functions for testing purpose
def create_linked_list(arr):
if len(arr)==0:
return None
head = Node(arr[0])
tail = head
for data in arr[1:]:
tail.next = Node(data)
tail = tail.next
return head
def print_linked_list(head):
while head:
print(head.data, end=' ')
head = head.next
print()
# + graffitiCellId="id_c1r40y6"
def test_function(test_case):
head = test_case[0]
solution = test_case[1]
node_tracker = dict({})
node_tracker['nodes'] = list()
temp = head
while temp:
node_tracker['nodes'].append(temp)
temp = temp.next
head = even_after_odd(head)
temp = head
index = 0
try:
while temp:
if temp.data != solution[index] or temp not in node_tracker['nodes']:
print("Fail")
return
temp = temp.next
index += 1
print("Pass")
except Exception as e:
print("Fail")
# + graffitiCellId="id_g689uo0"
arr = [1, 2, 3, 4, 5, 6]
solution = [1, 3, 5, 2, 4, 6]
head = create_linked_list(arr)
test_case = [head, solution]
test_function(test_case)
# + graffitiCellId="id_f2x6gq4"
arr = [1, 3, 5, 7]
solution = [1, 3, 5, 7]
head = create_linked_list(arr)
test_case = [head, solution]
test_function(test_case)
# + graffitiCellId="id_ejterzp"
arr = [2, 4, 6, 8]
solution = [2, 4, 6, 8]
head = create_linked_list(arr)
test_case = [head, solution]
test_function(test_case)
| Data Structures/Arrays and Linked List/linked_lists/Even-After-Odd-Nodes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# # San Diego Burrito Analytics: California burritos
#
# <NAME>
#
# 27 August 2016
#
# * This notebook formats the burrito data set in order to analyze California burritos
# * The data set is format so that it can be read into Tableau to make a visualization
#
# * Analyze the different kinds of California burritos served at the different locations (with guac?)
# ### Default imports
# +
# %config InlineBackend.figure_format = 'retina'
# %matplotlib inline
import numpy as np
import scipy as sp
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
sns.set_style("white")
# -
# ### Load data
import util2
df, dfRestaurants, dfIngredients = util2.load_burritos()
N = df.shape[0]
# ### Limit data to California burritos
dfCali = df.loc[df.Burrito.str.contains('.*[Cc]ali.*')]
dfCaliIngredients = dfIngredients.loc[df.Burrito.str.contains('.*[Cc]ali.*')][['Beef','Pico','Guac','Cheese','Fries','Sour cream','Chicken']]
dfRestaurants=dfRestaurants.reset_index().drop('index',axis=1)
dfCaliRestaurants = dfRestaurants.loc[[i for i,x in enumerate(dfRestaurants.Location) if x in dfCali.Location.unique()]]
# ### Process Cali burrito data: Averages for each restaurant
dfCaliAvg = dfCali.groupby('Location').agg({'Cost': np.mean,'Volume': np.mean,'Hunger': np.mean,
'Tortilla': np.mean,'Temp': np.mean,'Meat': np.mean,
'Fillings': np.mean,'Meat:filling': np.mean,'Uniformity': np.mean,
'Salsa': np.mean,'Synergy': np.mean,'Wrap': np.mean,
'overall': np.mean, 'Location':np.size})
dfCaliAvg.rename(columns={'Location': 'N'}, inplace=True)
dfCaliAvg['Location'] = list(dfCaliAvg.index)
# +
# Calculate latitutude and longitude for each city
import geocoder
addresses = dfCaliRestaurants['Address'] + ', San Diego, CA'
lats = np.zeros(len(addresses))
longs = np.zeros(len(addresses))
for i, address in enumerate(addresses):
g = geocoder.google(address)
Ntries = 1
while g.latlng ==[]:
g = geocoder.google(address)
Ntries +=1
print 'try again: ' + address
if Ntries >= 5:
raise ValueError('Address not found: ' + address)
lats[i], longs[i] = g.latlng
# # Check for nonsense lats and longs
if sum(np.logical_or(lats>34,lats<32)):
raise ValueError('Address not in san diego')
if sum(np.logical_or(longs<-118,longs>-117)):
raise ValueError('Address not in san diego')
# -
# Incorporate lats and longs into restaurants data
dfCaliRestaurants['Latitude'] = lats
dfCaliRestaurants['Longitude'] = longs
# Merge restaurant data with burrito data
dfCaliTableau = pd.merge(dfCaliRestaurants,dfCaliAvg,on='Location')
dfCaliTableau.to_csv('cali_now.csv')
| burrito/Burrito_California.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=false editable=false
# # Assignment 3
# Welcome to the third programming assignment for the course. This assignments will help to familiarise you with Boolean function oracles while revisiting the topics discussed in this week's lectures.
#
# ### Submission Guidelines
# For final submission, and to ensure that you have no errors in your solution, please use the 'Restart and Run All' option availble in the Kernel menu at the top of the page.
# To submit your solution, run the completed notebook and attach the solved notebook (with results visible) as a .ipynb file using the 'Add or Create' option under the 'Your Work' heading on the assignment page in Google Classroom.
# + deletable=false editable=false
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
# Importing standard Qiskit libraries
from qiskit import QuantumCircuit, execute
from qiskit.providers.aer import QasmSimulator
from qiskit.visualization import *
from qiskit.quantum_info import *
basis_gates = ['id', 'x', 'y', 'z', 's', 't', 'sdg', 'tdg', 'h', 'p', 'sx' ,'r', 'rx', 'ry', 'rz', 'u', 'u1', 'u2', 'u3', 'cx', 'ccx', 'barrier', 'measure', 'snapshot']
# + [markdown] deletable=false editable=false
# ## A quantum oracle implementation of the classical OR operation
# We've already seen that the Toffoli gate implements the quantum version of the classical AND operation. The first part of this exercise will require you to construct such a quantum implementation for the OR operation.
# The logical OR operation takes two Boolean inputs and returns 1 as the result if either or both of the inputs are 1. It is often denoted using the $\vee$ symbol (it is also called the disjunction operation). The truth table for the classical OR operation is given below:
#
# | $x$ | $y$ | $x\vee y$ |
# |----- |----- |----------- |
# | 0 | 0 | 0 |
# | 0 | 1 | 1 |
# | 1 | 0 | 1 |
# | 1 | 1 | 1 |
#
# ### De Morgan's laws
# Finding a gate that is the direct quantum analogue of the OR operation might prove to be difficult. Luckily, there are a set of two relation in Boolean algebra that can provide a helpful workaround.
# $$\overline{x\vee y} = \overline{x} \wedge \overline{y}$$
# This is read as _not ($x$ or $y$) = not $x$ and not $y$_
# $$\overline{x\wedge y} = \overline{x} \vee \overline{y}$$
# This is read as _not ($x$ and $y$) = not $x$ or not $y$_
# ## **Problem 1**
# 1. Using the expressions for De Morgan's laws above, construct a Boolean formula for $x \vee y$ consisting only of the logical AND and NOT operations.
# 2. We have provided the `QuantumCircuit()` for a quantum bit oracle to implement the OR operation. Apply the appropriate gates to this circuit based on the expression calculated in Step 1. Do NOT add a measurement
#
# <div class="alert alert-block alert-warning"><b>Warning: </b>Please be careful to ensure that the circuit below matches the oracle structure i.e. the input qubit states are not altered after the operation of the oracle.</div>
# +
or_oracle = QuantumCircuit(3)
# Do not change below this line
or_oracle.draw(output='mpl')
# + deletable=false editable=false
or_tt = ['000', '011', '101', '111']
def check_or_oracle(tt_row):
check_qc = QuantumCircuit(3)
for i in range(2):
if (tt_row[i] == '1'):
check_qc.x(i)
check_qc.extend(or_oracle)
check_qc.measure_all()
return (execute(check_qc.reverse_bits(),backend=QasmSimulator(), shots=1).result().get_counts().most_frequent() == tt_row)
try:
assert list(or_oracle.count_ops()) != [], f"Circuit cannot be empty"
assert 'measure' not in or_oracle.count_ops(), f"Please remove measurements"
assert set(or_oracle.count_ops().keys()).difference(basis_gates) == set(), f"Only the following basic gates are allowed: {basis_gates}"
for tt_row in or_tt:
assert check_or_oracle(tt_row), f" Input {tt_row[0:2]}: Your encoding is not correct"
print("Your oracle construction passed all checks")
except AssertionError as e:
print(f'Your code has an error: {e.args[0]}')
except Exception as e:
print(f'This error occured: {e.args[0]}')
# + [markdown] deletable=false editable=false
# ## Linear functions and the Bernstein-Vazirani Algorithm
# The Deutch-Jozsa algorithm allows us to distinguish between constant and balanced Boolean functions. There is an extension to the Deutsch-Jozsa algorithm that allows us to extract some information about a certain other class of functions. This is what we will be exploring now.
#
# An $n$-bit Boolean function $f(x)$ is called linear if it can be written as the bitwise product of a particular $n$-bit binary string $a$ and the function variable $x$ (which is also a binary string of length $n$), i.e., linear functions can be written as
# $$f(x) = a\cdot x \;(\text{ mod } 2)$$
#
# You might recall from the discussion on the Hadamard transform, that for any general $n$-qubit computational basis state, the Hadamard transform has the following effect
# $$H^{\otimes n}|a\rangle = \frac{1}{2^{n/2}}\sum\limits_{x=0}^{2^n-1}(-1)^{a\cdot x}|x\rangle$$
# Due to the self-inverting nature of the Hadamard transformation, we can apply $H^{\otimes n}$ to both sides of the above equation and get (after flipping sides)
# $$H^{\otimes n} \left( \frac{1}{2^{n/2}}\sum\limits_{x=0}^{2^n-1}(-1)^{a\cdot x}|x\rangle \right) = |a\rangle$$
# The term inside the brackets on the left hand side of the equation looks like what we would get if we passed an equal superposition state through a phase oracle for the Boolean function $f(x) = a\cdot x \;(\text{ mod } 2)$. This is depicted in the equation below:
# $$\frac{1}{2^{n/2}}\sum\limits_{x=0}^{2^n-1}|x\rangle \xrightarrow{U_f} \frac{1}{2^{n/2}}\sum\limits_{x=0}^{2^n-1}(-1)^{a\cdot x}|x\rangle$$
#
# The Bernstein-Vazirani algorithm uses all the things discussed above. Given an oracle for a function that we know is linear, we can find the binary string $a$ corresponding to the linear function. The steps of the algorithm are shown in the equation below and then described in words.
# $$|0^{\otimes n}\rangle \xrightarrow{H^{\otimes n}} \frac{1}{2^{n/2}}\sum\limits_{x=0}^{2^n-1}|x\rangle \xrightarrow{U_f} \frac{1}{2^{n/2}}\sum\limits_{x=0}^{2^n-1}(-1)^{a\cdot x}|x\rangle \xrightarrow{H^{\otimes n}} |a\rangle$$
# In the expression above, we've omitted (for readability) the mention of the extra qubit in the $|-\rangle$ state that is required for the oracle output, but it is necessary.
#
# ## **Problem 2**
# Consider the Boolean function $f(x) = (\overline{x_1} \wedge x_0) \vee (x_1 \wedge \overline{x_0})$. Take it as given that this function is a linear function. We want to find the 2-bit binary string $a$. Your objective is to use this expression above to implement the quantum bit oracle for this Boolean function.
# This is more complex than any expression we have seen so far, so the implementation will be carried out in a few steps.
#
# A `QuantumCircuit()` with 3 qubits is provided below.
# - $q_0$ and $q_1$ are the input qubits for the variables $x_0$ and $x_1$ respectively.
# - $q_2$ is the output qubit and stores the value of the final Boolean function expression
#
#
# + deletable=false editable=false
bv_oracle = QuantumCircuit(3)
bv_oracle.cx(0,2)
bv_oracle.cx(1,2)
bv_oracle.draw('mpl')
# + [markdown] deletable=false editable=false
# Using the bit oracle provided above, construct a circuit for the Bernstein-Vazirani algorithm.The steps for the algorithm are as follows:
# 1. Start will $(n+1)$ qubits in the $|0\rangle$ state. Here $n=2$. The first two qubits $q_0$ and $q_1$ will serve as input to the oracle. The extra qubit is used for the oracle output. Since we need a phase oracle, add gates to prepare the state $|-\rangle$ in this qubit ($q_2$).
# 2. Apply an $H$ gate to all the input qubits.
# 3. Apply the oracle $U_f$
# 4. Apply an $H$ gate to all the input qubits.
# 5. Measure the $n$ input qubits.
# If the function corresponding to $U_f$ is linear, the final state measured will be the binary string $a$.
#
# Astute readers will notice that the steps followed in the Bernstein-Vazirani and the Deutsch-jozsa algorithms are the same. `bv_circ` is a `QuantumCircuit(3,2)` given below. Add necessary operations to the circuit below to realise the steps for the Bernstein-Vazirani algorithm.
# +
bv_circ = QuantumCircuit(3,2)
# Do not remove this line
bv_circ.draw(output='mpl')
# + deletable=false editable=false
try:
assert list(bv_circ.count_ops()) != [], f"Circuit cannot be empty"
assert set(bv_circ.count_ops().keys()).difference(basis_gates) == set(), f"Only the following basic gates are allowed: {basis_gates}"
counts = execute(bv_circ.reverse_bits(), backend=QasmSimulator(), shots=8192).result().get_counts()
assert list(counts.keys()) == ['11'], "Your circuit did not produce the right answer"
print(" Your circuit produced the correct output. Please submit for evaluation.")
except AssertionError as e:
print(f'Your code has an error: {e.args[0]}')
except Exception as e:
print(f'This error occured: {e.args[0]}')
plot_histogram(counts)
| assignments/assignment3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import hvplot.pandas
import numpy as np
import ast
# +
df = pd.read_csv('agent_performance.csv')
df['Gradient'] = df['Gradient'].apply(ast.literal_eval)
df[['g1','g2','g3','g4', 'g5']] = pd.DataFrame(df.Gradient.tolist(), index= df.index)
df['State'] = df['State'].apply(ast.literal_eval)
df[['s1','s2','s3','s4', 's5']] = pd.DataFrame(df.State.tolist(), index= df.index)
df['a1'] = df['Action'].apply(lambda x: int(x[1]))
df['a2'] = df['Action'].apply(lambda x: int(x[3]))
df['a3'] = df['Action'].apply(lambda x: int(x[5]))
# -
df.hvplot(y='Reward')
df.hvplot(y=['g1','g2','g3','g4', 'g5'])
df.hvplot(y=['s1','s2','s3','s4', 's5'])
df.hvplot(y='Balance')
df.hvplot(y=['a1','a2','a3'])
| test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import norm
# Increase the font size (since this will be a small plot figure)
plt.rcParams.update({'font.size': 20})
# Create the x-values and pdf-values
x = np.linspace(0,1,1000)
pdf = norm.pdf(x,loc=0.5,scale=0.01)
# Create the plot, then set the labels and title
plt.plot(x, pdf,'b-', label='PDF ');
plt.xlabel('p')
plt.ylabel('PDF');
plt.title('Prior Probabilty for $p=P($Heads$)$');
| figure_fair_coin_prior.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
#
# <div>
# <img src="https://blogs.nasa.gov/swesarr/wp-content/uploads/sites/305/2021/06/swesarr.png" width="1589"/>
# </div>
#
#
# <center>
# <h1><font size="+3">SWESARR Tutorial</font></h1>
# </center>
#
# ---
#
# <CENTER>
# <H1 style="color:red">
# Introduction
# </H1>
# </CENTER>
# <div class="alert alert-block alert-info">
# <b>Objectives:</b>
# This is a 30-minute tutorial where we will ...
# <ol>
# <li> Introduce SWESARR </li>
# <li> Briefly introduce active and passive microwave remote sensing </li>
# <li> Learn how to access, filter, and visualize SWESARR data </li>
# </ol>
# </div>
# # SWESARR Tutorial
# ## Quick References
#
# <OL>
# <LI> <A HREF="https://glihtdata.gsfc.nasa.gov/files/radar/SWESARR/prerelease/">SWESARR SAR Data Pre-release FTP Server</A>
# <LI> <A HREF="https://nsidc.org/data/SNEX20_SWESARR_TB/versions/1"> SWESARR Radiometer Data, SnowEx20, v1</A>
# <LI> <A HREF="https://blogs.nasa.gov/swesarr/">SWESARR Blogspot</A>
# </OL>
# ## What is SWESARR?
from IPython.display import Audio,Image, YouTubeVideo; id='5hVQusosGSg'; YouTubeVideo(id=id,width=600,height=300,start=210,end=238)
# courtesy of this github post
# https://gist.github.com/christopherlovell/e3e70880c0b0ad666e7b5fe311320a62
# <UL>
# <LI> Airborne sensor system measuring active and passive microwave measurements
# <LI> Colocated measurements are taken simultaneously using an ultra-wideband antenna
# </UL>
# <P>
#
# SWESARR gives us insights on the different ways active and passive signals are influenced by snow over large areas.
# ## Active and Passive? Microwave Remote Sensing?
#
# ### Passive Systems
#
# * All materials can naturally emit electromagnetic waves
# * What is the cause?
#
# <div>
# <img src="https://blogs.nasa.gov/swesarr/wp-content/uploads/sites/305/2021/07/particles.png" width="360"/>
# </div>
# <br><br>
#
# * Material above zero Kelvin will display some vibration or movement of particles
# * These moving, charged particles will induce electromagnetic waves
# * If we're careful, we can measure these waves with a radio wave measuring tool, or "radiometer"
# <br>
#
# <div>
# <img src="https://blogs.nasa.gov/swesarr/wp-content/uploads/sites/305/2021/07/radiometer.png" width="500"/>
# </div>
#
#
# * Radiometers see emissions from many sources, but they're usually very weak
# * It's important to design a radiometer that (1) minimizes side lobes and (2) allows for averaging over the main beam
# * For this reason, radiometers often have low spatial resolution
#
# | ✏️ | Radiometers allow us to study earth materials through incoherent averaging of naturally emitted signals |
# |---------------|:----------------------------------------------------------------------------------------------------------|
#
# <br><br><br>
#
# ### Active Systems
# * While radiometers generally measure natural electromagnetic waves, radars measure man-made electromagnetic waves
# * Transmit your own wave, and listen for the returns
# * The return of this signal is dependent on the surface and volume characteristics of the material it contacts
#
# <div>
# <img src="https://blogs.nasa.gov/swesarr/wp-content/uploads/sites/305/2021/07/radar.png" width="500"/>
# </div>
#
# | ✏️ | Synthetic aperture radar allows for high spatial resolution through processing of coherent signals |
# |---------------|:----------------------------------------------------------------------------------------------------------|
#
# %%HTML
<style>
td { font-size: 15px }
th { font-size: 15px }
</style>
# ## SWESARR Sensors
# <center>
# <h1><font size="+1">SWESARR Frequencies, Polarization, and Bandwidth Specification </font></h1>
# </center>
#
# | Center-Frequency (GHz) | Band | Sensor | Bandwidth (MHz) | Polarization |
# | ---------------------- | ---------- | ------------ | --------------- | ------------ |
# | 9.65 | X | SAR | 200 | VH and VV |
# | 13.6 | Ku | SAR | 200 | VH and VV |
# | 17.25 | Ku | SAR | 200 | VH and VV |
# | 10.65 | X | Radiometer | 200 | H |
# | 18.7 | K | Radiometer | 200 | H |
# | 36.5 | Ka | Radiometer | 1,000 | H |
#
# ## SWESARR Instrument
#
# <br>
# <center>
# <img src="https://blogs.nasa.gov/swesarr/wp-content/uploads/sites/305/wppa/1.jpg", width="400", title="Plane" /> <br>
# <img src="https://blogs.nasa.gov/swesarr/wp-content/uploads/sites/305/wppa/4.jpg", width="400", title="Instrument" />
# </center>
#
# ## SWESARR Spatiotemporal Coverage
#
#
# * Currently, there are two primary dataset coverages
# * **2019**: 04 November through 06 November
# * **2020**: 10 February through 12 February
# * Below: radiometer coverage for all passes made between February 10 to February 12, 2020
# * SWESARR flights cover many snowpit locations over the Grand Mesa area as shown by the dots in blue
# <div>
# <img src="https://blogs.nasa.gov/swesarr/wp-content/uploads/sites/305/2021/06/passes.png", width="500", title="SWESARR passes over Grand Mesa in 2020" />
# </div>
# ## Reading SWESARR Data
#
# - SWESARR's SAR data is organized with a common file naming convention for finding the time, location, and type of data
# - [Lets look at the prerelease data on its homepage](https://glihtdata.gsfc.nasa.gov/files/radar/SWESARR/prerelease/)
# <div>
# <img src="https://blogs.nasa.gov/swesarr/wp-content/uploads/sites/305/2020/09/SWESARR-Naming-Convention_recolor_v2.png" width="1000"/>
# </div>
#
# ***
# <br><br>
# <CENTER>
# <H1 style="color:red">
# Accessing Data: SAR
# </H1>
# </CENTER>
#
# ### SAR Data Example
# +
# Import several libraries.
# comments to the right could be useful for local installation on Windows.
from shapely import speedups # https://www.lfd.uci.edu/~gohlke/pythonlibs/
speedups.disable() # <-- handle a potential error in cartopy
# downloader library
import requests # !conda install -c anaconda requests
# raster manipulation libraries
import rasterio # https://www.lfd.uci.edu/~gohlke/pythonlibs/
from osgeo import gdal # https://www.lfd.uci.edu/~gohlke/pythonlibs/
import cartopy.crs as ccrs # https://www.lfd.uci.edu/~gohlke/pythonlibs/
import rioxarray as rxr # !conda install -c conda-forge rioxarray
import xarray as xr # !conda install -c conda-forge xarray dask netCDF4 bottleneck
# plotting tools
from matplotlib import pyplot # !conda install matplotlib
import datashader as ds # https://www.lfd.uci.edu/~gohlke/pythonlibs/
import hvplot.xarray # !conda install hvplot
# append the subfolders of the current working directory to pythons path
import os
import sys
swesarr_subdirs = ["data", "util"]
tmp = [sys.path.append(os.getcwd() + "/swesarr/" + sd) for sd in swesarr_subdirs]
del tmp # suppress Jupyter notebook output, delete variable
from helper import gdal_corners, join_files, join_sar_radiom
# -
# #### Select your data
# +
# select files to download
# SWESARR data website
source_repo = 'https://glihtdata.gsfc.nasa.gov/files/radar/SWESARR/prerelease/'
# Example flight line
flight_line = 'GRMCT2_31801_20007_016_200211_225_XX_01/'
# SAR files within this folder
data_files = [
'GRMCT2_31801_20007_016_200211_09225VV_XX_01.tif',
'GRMCT2_31801_20007_016_200211_09225VH_XX_01.tif',
'GRMCT2_31801_20007_016_200211_13225VV_XX_01.tif',
'GRMCT2_31801_20007_016_200211_13225VH_XX_01.tif',
'GRMCT2_31801_20007_016_200211_17225VV_XX_01.tif',
'GRMCT2_31801_20007_016_200211_17225VH_XX_01.tif'
]
# store the location of the SAR tiles as they're located on the SWESARR data server
remote_tiles = [source_repo + flight_line + d for d in data_files]
# create local output data directory
output_dir = '/tmp/swesarr/data/'
try:
os.makedirs(output_dir)
except FileExistsError:
print('output directory prepared!')
# store individual TIF files locally on our computer / server
output_paths = [output_dir + d for d in data_files]
# -
# #### Download SAR data and place into data folder
# +
## for each file selected, store the data locally
##
## only run this block if you want to store data on the current
## server/hard drive this notebook is located.
##
################################################################
for remote_tile, output_path in zip(remote_tiles, output_paths):
# download data
r = requests.get(remote_tile)
# Store data (~= 65 MB/file)
if r.status_code == 200:
with open(output_path, 'wb') as f:
f.write(r.content)
# -
# #### Merge SAR datasets into single xarray file
da = join_files(output_paths)
da
# #### Plot data with hvplot
# +
# Set clim directly:
clim=(-20,20)
cmap='gray'
crs = ccrs.UTM(zone='12n')
tiles='EsriImagery'
da.hvplot.image(x='x',y='y',groupby='band',cmap=cmap,clim=clim,rasterize=True,
xlabel='Longitude',ylabel='Latitude',
frame_height=500, frame_width=500,
xformatter='%.1f',yformatter='%.1f', crs=crs, tiles=tiles, alpha=0.8)
# -
# | 🎉 | Congratulations! You now know how to download and display a SWESARR SAR dataset ! | 🎉 |
# | :--- | :--- | :--- |
# <br>
# ### Radiometer Data Example
# * SWESARR's radiometer data is publicly available at NSIDC
# * [Radiometer Data v1 Available Here](https://nsidc.org/data/SNEX20_SWESARR_TB/versions/1)
#
#
# +
import pandas as pd # !conda install pandas
import numpy as np # !conda install numpy
import xarray as xr # !conda install -c anaconda xarray
import hvplot # !conda install hvplot
import hvplot.pandas
import holoviews as hv # !conda install -c conda-forge holoviews
from holoviews.operation.datashader import datashade
from geopy.distance import distance #!conda install -c conda-forge geopy
# -
# #### Downloading SWESARR Radiometer Data with `wget`
#
# * If you are running this on the SnowEx Hackweek server, `wget` should be configured.
# * If you are using this tutorial on your local machine, you'll need `wget`.
# * Linux Users
# - You should be fine. This is likely baked into your operating systems. Congratulations! You chose correctly.
# * Apple Users
# - The author of this textbox has never used a Mac. There are many command-line suggestions online. `sudo brew install wget`, `sudo port install wget`, etc. Try searching online!
# * Windows Users
# - [Check out this tutorial, page 2](https://blogs.nasa.gov/swesarr/wp-content/uploads/sites/305/2020/10/how_to_download_SWESARR_radar_data.pdf) You'll need to download binaries for `wget`, and you should really make it an environment variable!
#
# Be sure to be diligent before installing anything to your computer.
#
# Regardless, fill in your NASA Earthdata Login credentials and follow along!
# !wget --quiet https://n5eil01u.ecs.nsidc.org/SNOWEX/SNEX20_SWESARR_TB.001/2020.02.11/SNEX20_SWESARR_TB_GRMCT2_13801_20007_000_200211_XKKa225H_v01.csv -O {output_dir}/SNEX20_SWESARR_TB_GRMCT2_13801_20007_000_200211_XKuKa225H_v03.csv
# #### Select an example radiometer data file
# +
# use the file we downloaded with wget above
excel_path = f'{output_dir}/SNEX20_SWESARR_TB_GRMCT2_13801_20007_000_200211_XKuKa225H_v03.csv'
# read data
radiom = pd.read_csv(excel_path)
# -
# #### Lets examine the radiometer data files content
radiom.hvplot.table(width=1100)
# #### Plot radiometer data with hvplot
# +
# create several series from pandas dataframe
lon_ser = pd.Series( radiom['Longitude (deg)'].to_list() * (3) )
lat_ser = pd.Series( radiom['Latitude (deg)'].to_list() * (3) )
tb_ser = pd.Series(
radiom['TB X (K)'].to_list() + radiom['TB K (K)'].to_list() +
radiom['TB Ka (K)'].to_list(), name="Tb"
)
# get series length, create IDs for plotting
sl = len(radiom['TB X (K)'])
id_ser = pd.Series(
['X-band']*sl + ['K-band']*sl + ['Ka-band']*sl, name="ID"
)
frame = {'Longitude (deg)' : lon_ser, 'Latitude (deg)' : lat_ser,
'TB' : tb_ser, 'ID' : id_ser}
radiom_p = pd.DataFrame(frame)
del sl, lon_ser, lat_ser, tb_ser, id_ser, frame
radiom_p.hvplot.points('Longitude (deg)', 'Latitude (deg)', groupby='ID', geo=True, color='TB', alpha=1,
tiles='ESRI', height=400, width=500)
# -
# | 🎉 | Congratulations! You now know how to download and display a SWESARR radiometer dataset ! | 🎉 |
# | :--- | :--- | :--- |
# ## SAR and Radiometer Together
# * The novelty of SWESARR lies in its colocated SAR and radiometer systems
# * Lets try filtering the SAR dataset and plotting both datasets together
# * For this session, I've made the code a function in {download}`swesarr/util/helper.py <./swesarr/util/helper.py>`
# +
data_p, data_ser = join_sar_radiom(da, radiom)
data_p.hvplot.points('Longitude (deg)', 'Latitude (deg)', groupby='ID', geo=True, color='Measurements', alpha=1,
tiles='ESRI', height=400, width=500)
# -
# ## Exercise
# <div class="alert alert-block alert-info">
# <b>Exercise:</b>
# <ol>
# <li>Plot a time-series visualization of the filtered SAR channels from the output of the
# <font face="Courier New" > join_sar_radiom()</font> function
# </li>
# <li>Plot a time-series visualization of the radiometer channels from the output of the
# <font face="Courier New" > join_sar_radiom()</font> function
# </li>
# <li>Hint: the data series variable (<font face="Courier New" > data_ser </font>) is a pandas data series.
# Use some of the methods shown above to read and plot the data!
# </li>
# </ol>
# </div>
# +
### Your Code Here #############################################################################################################
#
# Two of Many Options:
# 1.) Go the matplotlib route
# a.) Further reading below:
# https://matplotlib.org/stable/tutorials/introductory/pyplot.html
#
# 2.) Try using hvplot tools if you like
# a.) Further reading below:
# https://hvplot.holoviz.org/user_guide/Plotting.html
#
# Remember, if you don't use a library all of the time, you'll end up <search engine of your choice>-ing it. Go crazy!
#
################################################################################################################################
# configure some inline parameters to make things pretty / readable if you'd like to go with matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = (16, 9) # (w, h)
# -
# <br><br><br>
# ## [Warnings](#Table-of-Contents)
# <div class="alert alert-block alert-danger">
# <b>Interpreting Data:</b> After the 2019 and 2020 measurement periods for SWESARR, an internal timing error was found in the flight data which affects the spatial precision of the measurements. While we are working to correct this geospatial error, please consider this offset before drawing conclusions from SWESARR data if you are using a dataset prior to this correction. The SWESARR website will announce the update of the geospatially corrected dataset.
# </div>
#
| book/tutorials/sar/swesarr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:.conda-alphapept]
# language: python
# name: conda-env-.conda-alphapept-py
# ---
# +
# default_exp recalibration
# -
# # Recalibration
#
# > Functions related to recalibrating
# This notebook contains evertyhing related to recalibration of data.
#hide
from nbdev.showdoc import *
# ## Recalibration after search
#
# ### Precursor mass calibration
#
# Recalibration refers to the computational step where masses are recalibrated after a first search. The identified peptides are used to calculate the deviations of experimental masses to their theoretical masses. After recalibration, a second search with decreased precursor tolerance is performed.
#
# The recalibration is largely motivated by the software lock mass paper:
#
# [<NAME>, <NAME>, <NAME>. Software lock mass by two-dimensional minimization of peptide mass errors. J Am Soc Mass Spectrom. 2011;22(8):1373-1380. doi:10.1007/s13361-011-0142-8](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3231580/)
#
# Here, mass offsets are piecewise linearly approximated. The positions for approximation need to fulfill a number of criteria (e.g., a minimum number of samples and a minimum distance). The AlphaPept implementation is slightly modified by employing a more general `KNeighborsRegressor`-approach. In brief, the calibration is calculated for each point individually by estimating the deviation from its identified neighbors in n-dimensional space (e.g., retention time, mass, mobility).
#
# More specifically, the algorithm consists of the following steps:
#
# 1. Outlier removal: We remove outliers from the identified peptides by only accepting identifications with a mass offset that is within n (default 3) standard deviations to the mean.
# 2. For each point, we perform a neighbors lookup of the next n (default 100) neighbors. For the neighbor's lookup we need to scale the axis, which is done with a transform function either absolute or relative.
# 3. Next, we perform a regression based on the neighbors to determine the mass offset. The contribution of each neighbor is weighted by their distance.
#
# ### Fragment mass calibration
#
# The fragment mass calibration is based on the identified ions (i.e., b-hits and y-hits). For each hit, we calculate the offset to its theoretical mass. The correction is then applied by taking the median offset in ppm and applying it globally.
# +
#export
import numpy as np
import pandas as pd
def remove_outliers(
df: pd.DataFrame,
outlier_std: float) -> pd.DataFrame:
"""Helper function to remove outliers from a dataframe.
Outliers are removed based on the precursor offset mass (prec_offset).
All values within x standard deviations to the median are kept.
Args:
df (pd.DataFrame): Input dataframe that contains a prec_offset_ppm-column.
outlier_std (float): Range of standard deviations to filter outliers
Raises:
ValueError: An error if the column is not present in the dataframe.
Returns:
pd.DataFrame: A dataframe w/o outliers.
"""
if 'prec_offset_ppm' not in df.columns:
raise ValueError(f"Column prec_offset_ppm not in df")
else:
# Remove outliers for calibration
o_mass_std = np.abs(df['prec_offset_ppm'].std())
o_mass_median = df['prec_offset_ppm'].median()
df_sub = df.query('prec_offset_ppm < @o_mass_median+@outlier_std*@o_mass_std and prec_offset_ppm > @o_mass_median-@outlier_std*@o_mass_std').copy()
return df_sub
# +
#hide
def test_remove_outliers():
df = pd.DataFrame({'prec_offset_ppm':[1,1,1,1,1,3,5]})
assert remove_outliers(df, 2).values.max() == 3
assert remove_outliers(df, 1).values.max() == 1
test_remove_outliers()
# +
#export
def transform(
x: np.ndarray,
column: str,
scaling_dict: dict) -> np.ndarray:
"""Helper function to transform an input array for neighbors lookup used for calibration
Note: The scaling_dict stores information about how scaling is applied and is defined in get_calibration
Relative transformation: Compare distances relatively, for mz that is ppm, for mobility %.
Absolute transformation: Compare distance absolute, for RT it is the timedelta.
An example definition is below:
scaling_dict = {}
scaling_dict['mz'] = ('relative', calib_mz_range/1e6)
scaling_dict['rt'] = ('absolute', calib_rt_range)
scaling_dict['mobility'] = ('relative', calib_mob_range)
Args:
x (np.ndarray): Input array.
column (str): String to lookup what scaling should be applied.
scaling_dict (dict): Lookup dict to retrieve the scaling operation and factor for the column.
Raises:
KeyError: An error if the column is not present in the dict.
NotImplementedError: An error if the column is not present in the dict.
Returns:
np.ndarray: A scaled array.
"""
if column not in scaling_dict:
raise KeyError(f"Column {_} not in scaling_dict")
else:
type_, scale_ = scaling_dict[column]
if type_ == 'relative':
return np.log(x, out=np.zeros_like(x), where=(x>0))/scale_
elif type_ == 'absolute':
return x/scale_
else:
raise NotImplementedError(f"Type {type_} not known.")
# +
#hide
def test_transform():
scaling_dict = {'A':('relative', 10), 'B':('absolute', 20)}
x = np.array([1,2,3,4,5], dtype=float)
assert np.allclose(transform(x, 'A', scaling_dict), np.log(x)/10)
assert np.allclose(transform(x, 'B', scaling_dict), x/20)
test_transform()
# +
#export
from sklearn.neighbors import KNeighborsRegressor
def kneighbors_calibration(df: pd.DataFrame, features: pd.DataFrame, cols: list, target: str, scaling_dict: dict, calib_n_neighbors: int) -> np.ndarray:
"""Calibration using a KNeighborsRegressor.
Input arrays from are transformed to be used with a nearest-neighbor approach.
Based on neighboring points a calibration is calculated for each input point.
Args:
df (pd.DataFrame): Input dataframe that contains identified peptides (w/o outliers).
features (pd.DataFrame): Features dataframe for which the masses are calibrated.
cols (list): List of input columns for the calibration.
target (str): Target column on which offset is calculated.
scaling_dict (dict): A dictionary that contains how scaling operations are applied.
calib_n_neighbors (int): Number of neighbors for calibration.
Returns:
np.ndarray: A numpy array with calibrated masses.
"""
data = df[cols]
tree_points = data.values
for idx, _ in enumerate(data.columns):
tree_points[:, idx] = transform(tree_points[:, idx], _, scaling_dict)
target_points = features[[_+'_matched' for _ in cols]].values
for idx, _ in enumerate(data.columns):
target_points[:, idx] = transform(target_points[:, idx], _, scaling_dict)
neigh = KNeighborsRegressor(n_neighbors=calib_n_neighbors, weights = 'distance')
neigh.fit(tree_points, df[target].values)
y_hat = neigh.predict(target_points)
return y_hat
# +
#hide
def test_kneighbors_calibration():
scaling_dict = {'A':('relative', 10), 'B':('absolute', 20)}
df = pd.DataFrame({'prec_offset_ppm':[1,1,1,2], 'A':np.array([1,2,3,4],dtype=float)})
features = pd.DataFrame({'mass':[1,1,1,1], 'A_matched':np.array([1,2,3,4],dtype=float)})
cols = ['A']
target = 'prec_offset_ppm'
calib_n_neighbors = 3
assert np.allclose(kneighbors_calibration(df, features, cols, target, scaling_dict, calib_n_neighbors), np.array([1,1,1,2]))
test_kneighbors_calibration()
# +
#export
import logging
def get_calibration(
df: pd.DataFrame,
features:pd.DataFrame,
outlier_std: float = 3,
calib_n_neighbors: int = 100,
calib_mz_range: int = 20,
calib_rt_range: float = 0.5,
calib_mob_range: float = 0.3,
**kwargs) -> (np.ndarray, float):
"""Wrapper function to get calibrated values for the precursor mass.
Args:
df (pd.DataFrame): Input dataframe that contains identified peptides.
features (pd.DataFrame): Features dataframe for which the masses are calibrated.
outlier_std (float, optional): Range in standard deviations for outlier removal. Defaults to 3.
calib_n_neighbors (int, optional): Number of neighbors used for regression. Defaults to 100.
calib_mz_range (int, optional): Scaling factor for mz range. Defaults to 20.
calib_rt_range (float, optional): Scaling factor for rt_range. Defaults to 0.5.
calib_mob_range (float, optional): Scaling factor for mobility range. Defaults to 0.3.
**kwargs: Arbitrary keyword arguments so that settings can be passes as whole.
Returns:
corrected_mass (np.ndarray): The calibrated mass
y_hat_std (float): The standard deviation of the precursor offset after calibration
"""
if len(df) > calib_n_neighbors:
target = 'prec_offset_ppm'
cols = ['mz','rt']
if 'mobility' in df.columns:
cols += ['mobility']
scaling_dict = {}
scaling_dict['mz'] = ('relative', calib_mz_range/1e6)
scaling_dict['rt'] = ('absolute', calib_rt_range)
scaling_dict['mobility'] = ('relative', calib_mob_range)
df_sub = remove_outliers(df, outlier_std)
y_hat = kneighbors_calibration(df, features, cols, target, scaling_dict, calib_n_neighbors)
corrected_mass = (1-y_hat/1e6) * features['mass_matched']
y_hat_std = y_hat.std()
mad_offset = np.median(np.absolute(y_hat - np.median(y_hat)))
logging.info(f'Precursor calibration std {y_hat_std:.2f}, {mad_offset:.2f}')
return corrected_mass, y_hat_std, mad_offset
else:
logging.info('Not enough data points present. Skipping recalibration.')
mad_offset = np.median(np.absolute(df['prec_offset_ppm'].values - np.median(df['prec_offset_ppm'].values)))
return features['mass_matched'], np.abs(df['prec_offset_ppm'].std()), mad_offset
# +
#hide
def test_get_calibration():
df = pd.DataFrame({'prec_offset_ppm':[0,0,0,0],
'mz':np.array([10,10,10,10], dtype=float),
'rt':np.array([1,2,3,4], dtype=float)})
features = pd.DataFrame({'mass_matched':[100,100,100,100],
'mz_matched':np.array([10,10,10,10], dtype=float),
'rt_matched':np.array([1,2,3,4], dtype=float)})
corrected_mass, y_hat_std, mad_offset = get_calibration(df, features, calib_n_neighbors=3)
assert np.allclose(corrected_mass.values, np.array([100,100,100,100]))
assert y_hat_std == 0
# Test calibration on files
import alphapept.io
ms_data = alphapept.io.MS_Data_File('../testfiles/test.ms_data.hdf')
features = ms_data.read(dataset_name="features")
df = ms_data.read(dataset_name="first_search")
corrected_mass, y_hat_std, mad_offset = get_calibration(df, features, calib_n_neighbors = 10)
assert y_hat_std < df['prec_offset_ppm'].std()
test_get_calibration()
# +
#export
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import cm
from matplotlib.colors import Normalize
from scipy.interpolate import interpn
def density_scatter( x , y, ax = None, sort = True, bins = 20, **kwargs ) :
"""
Scatter plot colored by 2d histogram
Adapted from https://stackoverflow.com/questions/20105364/how-can-i-make-a-scatter-plot-colored-by-density-in-matplotlib
"""
data , x_e, y_e = np.histogram2d( x, y, bins = bins, density = True )
z = interpn( ( 0.5*(x_e[1:] + x_e[:-1]) , 0.5*(y_e[1:]+y_e[:-1]) ) , data , np.vstack([x,y]).T , method = "splinef2d", bounds_error = False)
#To be sure to plot all data
z[np.where(np.isnan(z))] = 0.0
# Sort the points by density, so that the densest points are plotted last
if sort :
idx = z.argsort()
x, y, z = x[idx], y[idx], z[idx]
ax.scatter( x, y, c=z, cmap='turbo', **kwargs )
return ax
def save_fragment_calibration(ions, corrected, std_offset, file_name, settings):
f, axes = plt.subplots(2, 2, figsize=(20,10))
ax1 = axes[0,0]
ax2 = axes[1,0]
ax3 = axes[0,1]
ax4 = axes[1,1]
ax1 = density_scatter(ions['rt'].values, ions['delta_ppm'].values, ax = ax1)
ax1.set_title('Fragment error before correction')
ax1.axhline(0, color='w', linestyle='-', alpha=0.5)
ax1.set_ylabel('Error (ppm)')
ax1.set_xlabel('RT (min)')
ax2 = density_scatter(ions['rt'].values, corrected.values, ax = ax2)
ax1.axhline(0, color='w', linestyle='-', alpha=0.5)
ax2.axhline(0, color='w', linestyle='-', alpha=0.5)
ax2.axhline(0+std_offset*settings['search']['calibration_std_frag'], color='w', linestyle=':', alpha=0.5)
ax2.axhline(0-std_offset*settings['search']['calibration_std_frag'], color='w', linestyle=':', alpha=0.5)
ax2.set_title('Fragment error after correction')
ax2.set_ylabel('Error (ppm)')
ax2.set_xlabel('RT (min)')
ax3 = density_scatter(ions['ion_mass'].values, ions['delta_ppm'].values, bins=50, ax = ax3)
ax3.axhline(0, color='w', linestyle='-', alpha=0.5)
ax3.set_ylabel('Error (ppm)')
ax3.set_xlabel('m/z')
ax3.set_xlim([100,1500])
ax3.set_title('Fragment error before correction')
ax4 = density_scatter(ions['ion_mass'].values, corrected.values, bins=50, ax = ax4)
ax4.set_ylabel('Error (ppm)')
ax4.set_xlabel('m/z')
ax4.set_xlim([100, 1500])
ax4.set_title('Fragment error after correction')
ax4.axhline(0, color='w', linestyle='-', alpha=0.5)
ax4.axhline(0+std_offset*settings['search']['calibration_std_frag'], color='w', linestyle=':', alpha=0.5)
ax4.axhline(0-std_offset*settings['search']['calibration_std_frag'], color='w', linestyle=':', alpha=0.5)
base, ext = os.path.splitext(file_name)
plt.suptitle(os.path.split(file_name)[1])
for ax in [ax1, ax2, ax3, ax4]:
ax.set_ylim([-settings['search']['frag_tol'], settings['search']['frag_tol']])
plt.savefig(base+'_calibration.png')
def calibrate_fragments_nn(ms_file_, file_name, settings):
logging.info('Starting fragment calibration.')
skip = False
try:
logging.info(f'Calibrating fragments with neighbors')
ions = ms_file_.read(dataset_name='ions')
except KeyError:
logging.info('No ions to calibrate fragment masses found')
skip = True
if not skip:
calib_n_neighbors = 400
psms = ms_file_.read(dataset_name='first_search')
#Calculate offset
ions['rt'] = psms['rt'][ions['psms_idx'].values.astype('int')].values
ions['delta_ppm'] = ((ions['db_mass'] - ions['ion_mass'])/((ions['db_mass'] + ions['ion_mass'])/2)*1e6).values
ions['hits'] = psms['hits'][ions['psms_idx'].values.astype('int')].values
#Min score to only use "true hits"
min_score = 12
ions = ions[ions['hits']> min_score]
#Train regressor
neigh = KNeighborsRegressor(n_neighbors=calib_n_neighbors, weights = 'distance')
neigh.fit(ions['rt'].values.reshape(-1, 1), ions['delta_ppm'].values)
#Read required datasets
rt_list_ms2 = ms_file_.read_DDA_query_data()['rt_list_ms2']
mass_list_ms2 = ms_file_.read_DDA_query_data()['mass_list_ms2']
incides_ms2 = ms_file_.read_DDA_query_data()['indices_ms2']
scan_idx = np.searchsorted(incides_ms2, np.arange(len(mass_list_ms2)), side='right') - 1
#Estimate offset
y_hat = neigh.predict(rt_list_ms2.reshape(-1, 1))
y_hat_ = neigh.predict(ions['rt'].values.reshape(-1, 1))
delta_ppm_corrected = ions['delta_ppm'] - y_hat_
median_off_corrected = np.median(delta_ppm_corrected.values)
delta_ppm_median_corrected = delta_ppm_corrected - median_off_corrected
mad_offset = np.median(np.abs(delta_ppm_median_corrected))
try:
offset = ms_file_.read(dataset_name = 'corrected_fragment_mzs')
except KeyError:
offset = np.zeros(len(mass_list_ms2))
offset += -y_hat[scan_idx] - median_off_corrected
delta_ppm_median = ions['delta_ppm'].median()
delta_ppm_std = ions['delta_ppm'].std()
delta_ppm_median_corrected_median = delta_ppm_median_corrected.median()
delta_ppm_median_corrected_std = delta_ppm_median_corrected.std()
logging.info(f'Median offset (std) {delta_ppm_median:.2f} ({delta_ppm_std:.2f}) - after calibration {delta_ppm_median_corrected_median:.2f} ({delta_ppm_median_corrected_std:.2f}) Mad offset {mad_offset:.2f}')
logging.info('Saving calibration')
save_fragment_calibration(ions, delta_ppm_median_corrected, delta_ppm_median_corrected_std, file_name, settings)
ms_file_.write(
offset,
dataset_name="corrected_fragment_mzs",
)
ms_file_.write(np.array([delta_ppm_median_corrected_std]), dataset_name="estimated_max_fragment_ppm")
# +
#export
from typing import Union
import alphapept.io
from alphapept.score import score_x_tandem
import os
def calibrate_hdf(
to_process: tuple, callback=None, parallel=True) -> Union[str,bool]:
"""Wrapper function to get calibrate a hdf file when using the parallel executor.
The function loads the respective dataframes from the hdf, calls the calibration function and applies the offset.
Args:
to_process (tuple): Tuple that contains the file index and the settings dictionary.
callback ([type], optional): Placeholder for callback (unused).
parallel (bool, optional): Placeholder for parallel usage (unused).
Returns:
Union[str,bool]: Either True as boolean when calibration is successfull or the Error message as string.
"""
try:
index, settings = to_process
file_name = settings['experiment']['file_paths'][index]
base_file_name, ext = os.path.splitext(file_name)
ms_file = base_file_name+".ms_data.hdf"
ms_file_ = alphapept.io.MS_Data_File(ms_file, is_overwritable=True)
features = ms_file_.read(dataset_name='features')
try:
psms = ms_file_.read(dataset_name='first_search')
except KeyError: #no elements in search
psms = pd.DataFrame()
if len(psms) > 0 :
df = score_x_tandem(
psms,
fdr_level=settings["search"]["peptide_fdr"],
plot=False,
verbose=False,
**settings["search"]
)
corrected_mass, prec_offset_ppm_std, prec_offset_ppm_mad = get_calibration(
df,
features,
**settings["calibration"]
)
ms_file_.write(
corrected_mass,
dataset_name="corrected_mass",
group_name="features"
)
else:
ms_file_.write(
features['mass_matched'],
dataset_name="corrected_mass",
group_name="features"
)
prec_offset_ppm_std = 0
ms_file_.write(
prec_offset_ppm_std,
dataset_name="corrected_mass",
group_name="features",
attr_name="estimated_max_precursor_ppm"
)
logging.info(f'Precursor calibration of file {ms_file} complete.')
# Calibration of fragments
calibrate_fragments_nn(ms_file_, file_name, settings)
logging.info(f'Fragment calibration of file {ms_file} complete.')
return True
except Exception as e:
logging.error(f'Calibration of file {ms_file} failed. Exception {e}.')
return f"{e}" #Can't return exception object, cast as string
# -
# #### Database calibration
#
# Another way to calibrate the fragment and precursor masses is by directly comparing them to a previously generated theoretical mass database. Here, peaks in the distribution of databases are used to align the experimental masses.
# +
#export
import scipy.stats
import scipy.signal
import scipy.interpolate
import alphapept.fasta
#The following function does not have an own unit test but is run by test_calibrate_fragments.
def get_db_targets(
db_file_name: str,
max_ppm: int=100,
min_distance: float=0.5,
ms_level: int=2,
) ->np.ndarray:
"""Function to extract database targets for database-calibration.
Based on the FASTA database it finds masses that occur often. These will be used for calibration.
Args:
db_file_name (str): Path to the database.
max_ppm (int, optional): Maximum distance in ppm between two peaks. Defaults to 100.
min_distance (float, optional): Minimum distance between two calibration peaks. Defaults to 0.5.
ms_level (int, optional): MS-Level used for calibration, either precursors (1) or fragmasses (2). Defaults to 2.
Raises:
ValueError: When ms_level is not valid.
Returns:
np.ndarray: Numpy array with calibration masses.
"""
if ms_level == 1:
db_mzs_ = alphapept.fasta.read_database(db_file_name, 'precursors')
elif ms_level == 2:
db_mzs_ = alphapept.fasta.read_database(db_file_name, 'fragmasses')
else:
raise ValueError(f"{ms_level} is not a valid ms level")
tmp_result = np.bincount(
(
np.log10(
db_mzs_[
np.isfinite(db_mzs_) & (db_mzs_ > 0)
].flatten()
) * 10**6
).astype(np.int64)
)
db_mz_distribution = np.zeros_like(tmp_result)
for i in range(1, max_ppm):
db_mz_distribution[i:] += tmp_result[:-i]
db_mz_distribution[:-i] += tmp_result[i:]
peaks = scipy.signal.find_peaks(db_mz_distribution, distance=max_ppm)[0]
db_targets = 10 ** (peaks / 10**6)
db_array = np.zeros(int(db_targets[-1]) + 1, dtype=np.float64)
last_int_mz = -1
last_mz = -1
for mz in db_targets:
mz_int = int(mz)
if (mz_int != last_int_mz) & (mz > (last_mz + min_distance)):
db_array[mz_int] = mz
else:
db_array[mz_int] = 0
last_int_mz = mz_int
last_mz = mz
return db_array
# +
#export
#The following function does not have an own unit test but is run by test_calibrate_fragments.
def align_run_to_db(
ms_data_file_name: str,
db_array: np.ndarray,
max_ppm_distance: int=1000000,
rt_step_size:float =0.1,
plot_ppms: bool=False,
ms_level: int=2,
) ->np.ndarray:
"""Function align a run to it's theoretical FASTA database.
Args:
ms_data_file_name (str): Path to the run.
db_array (np.ndarray): Numpy array containing the database targets.
max_ppm_distance (int, optional): Maximum distance in ppm. Defaults to 1000000.
rt_step_size (float, optional): Stepsize for rt calibration. Defaults to 0.1.
plot_ppms (bool, optional): Flag to indicate plotting. Defaults to False.
ms_level (int, optional): ms_level for calibration. Defaults to 2.
Raises:
ValueError: When ms_level is not valid.
Returns:
np.ndarray: Estimated errors
"""
ms_data = alphapept.io.MS_Data_File(ms_data_file_name)
if ms_level == 1:
mzs = ms_data.read(dataset_name="mass_matched", group_name="features")
rts = ms_data.read(dataset_name="rt_matched", group_name="features")
elif ms_level == 2:
mzs = ms_data.read(dataset_name="Raw/MS2_scans/mass_list_ms2")
inds = ms_data.read(dataset_name="Raw/MS2_scans/indices_ms2")
precursor_rts = ms_data.read(dataset_name="Raw/MS2_scans/rt_list_ms2")
rts = np.repeat(precursor_rts, np.diff(inds))
else:
raise ValueError(f"{ms_level} is not a valid ms level")
selected = mzs.astype(np.int64)
ds = np.zeros((3, len(selected)))
if len(db_array) < len(selected) + 1:
tmp = np.zeros(len(selected) + 1)
tmp[:len(db_array)] = db_array
db_array = tmp
ds[0] = mzs - db_array[selected - 1]
ds[1] = mzs - db_array[selected]
ds[2] = mzs - db_array[selected + 1]
min_ds = np.take_along_axis(
ds,
np.expand_dims(np.argmin(np.abs(ds), axis=0), axis=0),
axis=0
).squeeze(axis=0)
ppm_ds = min_ds / mzs * 10**6
selected = np.abs(ppm_ds) < max_ppm_distance
selected &= np.isfinite(rts)
rt_order = np.argsort(rts)
rt_order = rt_order[selected[rt_order]]
ordered_rt = rts[rt_order]
ordered_ppm = ppm_ds[rt_order]
rt_idx_break = np.searchsorted(
ordered_rt,
np.arange(ordered_rt[0], ordered_rt[-1], rt_step_size),
"left"
)
median_ppms = np.empty(len(rt_idx_break) - 1)
for i in range(len(median_ppms)):
median_ppms[i] = np.median(
ordered_ppm[rt_idx_break[i]: rt_idx_break[i + 1]]
)
if plot_ppms:
import matplotlib.pyplot as plt
plt.plot(
rt_step_size + np.arange(
ordered_rt[0],
ordered_rt[-1],
rt_step_size
)[:-1],
median_ppms
)
plt.show()
estimated_errors = scipy.interpolate.griddata(
rt_step_size / 2 + np.arange(
ordered_rt[0],
ordered_rt[-1] - 2 * rt_step_size,
rt_step_size
),
median_ppms,
rts,
fill_value=0,
method="linear",
rescale=True
)
estimated_errors[~np.isfinite(estimated_errors)] = 0
return estimated_errors
# +
#export
def calibrate_fragments(
db_file_name: str,
ms_data_file_name: str,
ms_level: int=2,
write = True,
plot_ppms = False,
):
"""Wrapper function to calibrate fragments.
Calibrated values are saved to corrected_fragment_mzs
Args:
db_file_name (str): Path to database
ms_data_file_name (str): Path to ms_data file
ms_level (int, optional): MS-level for calibration. Defaults to 2.
write (bool, optional): Boolean flag for test purposes to avoid writing to testfile. Defaults to True.
plot_ppms (bool, optional): Boolean flag to plot the calibration. Defaults to False.
"""
db_array = get_db_targets(
db_file_name,
max_ppm=100,
min_distance=0.5,
ms_level=ms_level,
)
estimated_errors = align_run_to_db(
ms_data_file_name,
db_array=db_array,
ms_level=ms_level,
plot_ppms=plot_ppms,
)
if write:
ms_file = alphapept.io.MS_Data_File(ms_data_file_name, is_overwritable=True)
ms_file.write(
estimated_errors,
dataset_name="corrected_fragment_mzs",
)
# +
#hide
def test_calibrate_fragments():
"""
This tests if the function can be called with the sample files.
TODO: add functional tests
"""
calibrate_fragments('../testfiles/database.hdf', '../testfiles/test.ms_data.hdf', write=False)
test_calibrate_fragments()
# -
#hide
from nbdev.export import *
notebook2script()
| nbs/07_recalibration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="LlkGjhyZgXqB"
# # Desafio:
#
# Você faz parte da equipe de Analytics de uma grande marca de vestuário com mais de 25 lojas espalhadas em Shoppings de todo o Brasil.
#
# Toda semana você precisa enviar para a diretoria um ranking atualizado com as 25 lojas contendo 3 informações:
# - Faturamento de cada Loja
# - Quantidade de Produtos Vendidos de cada Loja
# - Ticket Médio dos Produto de cada Loja
#
# Além disso, cada loja tem 1 gerente que precisa receber o resumo das informações da loja dele. Por isso, cada gerente deve receber no e-mail:
# - Faturamento da sua loja
# - Quantidade de Produtos Vendidos da sua loja
# - Ticket Médio dos Produto da sua Loja
#
# Esse relatório é sempre enviado como um resumo de todos os dados disponíveis no ano.
# + [markdown] id="DrMBBB_XpNzA"
# # Solução do Desafio:
#
# Para resolver o desafio vamos seguir a seguinte lógica:
#
# - Passo 1 - Importar a base de Dados
# - Passo 2 - Visualizar a Base de Dados para ver se precisamos fazer algum tratamento
# - Passo 3 - Calcular os indicadores de todas as lojas:
# - Faturamento por Loja
# - Quantidade de Produtos Vendidos por Loja
# - Ticket Médio dos Produto por Loja
# - Passo 4 - Calcular os indicadores de cada loja
# - Passo 5 - Enviar e-mail para cada loja
# - Passo 6 - Enviar e-mail para a diretoria
# + [markdown] id="LfrZ3LGvp9S7"
# ### Passo 1 - Importando a Base de Dados + Passo 2 - Visualizando os Dados
# + colab={"base_uri": "https://localhost:8080/", "height": 417} id="xMpovGrClBSh" executionInfo={"status": "ok", "timestamp": 1612710107820, "user_tz": 180, "elapsed": 19118, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhErqX2JU50uJaYinI4De9eiLkDS3VoBYK4CnzlHA=s64", "userId": "10609963884889021877"}} outputId="2340a59a-b817-44d3-efe9-5ca445a95f67"
import pandas as pd
df = pd.read_excel(r'/content/drive/MyDrive/Colab Notebooks/Projetos Intensivão de Python/Aula 1/Vendas.xlsx')
display(df)
# + [markdown] id="VajUvKlIqGfy"
# ### Passo 3.1 - Calculando o Faturamento por Loja
# + colab={"base_uri": "https://localhost:8080/", "height": 849} id="yvCRCMfV-88_" executionInfo={"status": "ok", "timestamp": 1612280130861, "user_tz": 180, "elapsed": 915, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhErqX2JU50uJaYinI4De9eiLkDS3VoBYK4CnzlHA=s64", "userId": "10609963884889021877"}} outputId="092b7a78-955b-46f3-b853-02193f2ac6a0"
faturamento = df[['ID Loja', 'Valor Final']].groupby('ID Loja').sum()
faturamento = faturamento.sort_values(by='Valor Final', ascending=False)
display(faturamento)
# + [markdown] id="pUf-OWDiqSNg"
# ### Passo 3.2 - Calculando a Quantidade Vendida por Loja
# + colab={"base_uri": "https://localhost:8080/", "height": 849} id="a_MAXNz8_Wkf" executionInfo={"status": "ok", "timestamp": 1612280135301, "user_tz": 180, "elapsed": 1037, "user": {"displayName": "Jo\u00e3<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhErqX2JU50uJaYinI4De9eiLkDS3VoBYK4CnzlHA=s64", "userId": "10609963884889021877"}} outputId="c64825f9-518d-4213-bd49-54fd7091d078"
quantidade = df[['ID Loja', 'Quantidade']].groupby('ID Loja').sum()
quantidade = quantidade.sort_values(by='ID Loja', ascending=False)
display(quantidade)
# + [markdown] id="55E2osWYqWr0"
# ### Passo 3.3 - Calculando o Ticket Médio dos Produtos por Loja
# + colab={"base_uri": "https://localhost:8080/", "height": 849} id="QDdIQVg4APsM" executionInfo={"status": "ok", "timestamp": 1612280139877, "user_tz": 180, "elapsed": 932, "user": {"displayName": "Jo\u00e3<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhErqX2JU50uJaYinI4De9eiLkDS3VoBYK4CnzlHA=s64", "userId": "10609963884889021877"}} outputId="495d6ab2-192e-44c0-b4be-10d37e12d170"
ticket_medio = (faturamento['Valor Final'] / quantidade['Quantidade']).to_frame()
ticket_medio = ticket_medio.rename(columns={0: 'Ticket Medio'})
ticket_medio = ticket_medio.sort_values(by='Ticket Medio', ascending=False)
display(ticket_medio)
# + [markdown] id="OIyFO49Aqiv5"
# ### Criando a função de enviar e-mail
# + id="6w561mVoCcMQ"
# função enviar_email
import smtplib
import email.message
def enviar_email(resumo_loja, loja):
server = smtplib.SMTP('smtp.gmail.com:587')
email_content = f'''
<p>Coe Lira,</p>
{resumo_loja.to_html()}
<p>Tmj</p>'''
msg = email.message.Message()
msg['Subject'] = f'Lira Rules - Loja: {loja}'
msg['From'] = '<EMAIL>'
msg['To'] = email
password = <PASSWORD>
msg.add_header('Content-Type', 'text/html')
msg.set_payload(email_content)
s = smtplib.SMTP('smtp.gmail.com: 587')
s.starttls()
# Login Credentials for sending the mail
s.login(msg['From'], password)
s.sendmail(msg['From'], [msg['To']], msg.as_string().encode('utf-8'))
# + [markdown] id="Xby-JPcdqmdK"
# ### Calculando Indicadores por Loja + Enviar E-mail para todas as lojas
# + id="6NLWIl6dRVIl"
# email para diretoria
tabela_diretoria = faturamento.join(quantidade).join(ticket_medio)
enviar_email(tabela_diretoria, 'Todas as Lojas')
# + id="JHdM0HSHBAEP"
lojas = df['ID Loja'].unique()
for loja in lojas:
tabela_loja = df.loc[df['ID Loja'] == loja, ['ID Loja', 'Quantidade', 'Valor Final']]
resumo_loja = tabela_loja.groupby('ID Loja').sum()
resumo_loja['Ticket Médio'] = resumo_loja['Valor Final'] / resumo_loja['Quantidade']
enviar_email(resumo_loja, loja)
# + [markdown] id="cZwGA9GEqrj8"
# ### Enviar e-mail para a diretoria
| docs/Gabarito - Projeto1 - Intensivao de Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json
import os
json_files = [pos_json for pos_json in os.listdir("../allignment/") if pos_json.endswith('.json')]
len(json_files)
json_files[0]
for json_file_path in json_files:
with open('../allignment/'+json_file_path) as json_file:
dev = json.load(json_file)
json_object = json.dumps(dev, indent = 4,ensure_ascii=False)
with open('../allignment/'+"pretty_" + json_file_path, "w", encoding='utf-8') as outfile:
outfile.write(json_object)
| enelpi/json process/pretty_json_formatter_in_specific_directory.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# # Armillaria Penetration map
# ## Left Hemisphere
#
# this uses the new function which imports xyz coordinates from celldb
#
# I started recording coordinates after some sites, so the early ones were later estimations
# not perfectly in line with reality.
#
# A composite image with the penetrations overlayed on a picture of the brain is found at
# /auto/data/lbhb/photos/Craniotomies/Armillaria
# + pycharm={"name": "#%%\n"}
import pathlib as pl
from nems_lbhb.penetration_map import penetration_map
import matplotlib.pyplot as plt
sites = ['ARM012d', 'ARM013b', 'ARM014b', 'ARM015b', 'ARM016c', 'ARM017a', 'ARM018a', 'ARM019a', 'ARM020a',
'ARM021b', 'ARM022b', 'ARM023a', 'ARM024a', 'ARM025a', 'ARM026b', 'ARM027a', 'ARM028b', 'ARM029a', 'ARM030a',
'ARM031a', 'ARM032a', 'ARM033a', 'ARM034a']
# fig, coords = penetration_map(sites, cubic=False, flip_X=True, flatten=False)
fig, coords = penetration_map(sites, equal_aspect=True, flip_X=True, flatten=True)
print('ARM034a is a post mortem dye staining penetration')
# saves the scatter
mappath = pl.Path('/auto/users/mateo/Pictures/Armillaria/ARM_LH_map.png')
fig.savefig(mappath, transparent=True)
| reports/210209_ARM_penetration_map.ipynb |