code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # How to Create a US Census Bar Chart Race
# ## Overview
# The beginning of every data science project starts with data and this will be the focus of this project. Usually, this part involves three steps:
# * data acquisition,
# * data cleaning and
# * data visualization.
#
# Data visualization can mean anything from simple print statements to fancy plots and animations.
#
# ## Data Acquisition
# As a data source, we use Wikipedia's census data websites. Since this data is only available on the web, we need to parse the website and extract the useful data. We use the popular [pandas](https://pandas.pydata.org/) package for this task.
import pandas as pd
# We also be using two more packages, `pickle` which allows us to easily save data to disk and `re` which allows us to use [regular expression](https://regexr.com/) syntax. Finally, for nicer table formatting, we import `IPython.display`.
# +
import pickle
import re
from IPython.display import display, HTML
# Define empty Census data dictionary - the keys will be the census years.
Census_raw = {}
# -
# ### Parsing Wikipedia
# Generally, parsing is not recommended to get data from the web as we often have to deal with messy formatting and a lot of cleaning up is needed. However, in this case, Wikipedia's layout of the US census years seems relatively consistent and thus we give parsing a shot. The following code goes through all Wikipedia census websites from [1790](https://en.wikipedia.org/wiki/1790_United_States_Census) to [2010](https://en.wikipedia.org/wiki/2010_United_States_Census) and collects all tables from the pages. We also add the latest census estimate from 2019.
# +
# Uncomment if you like to parse the data again (not recommended!)
# for year in range(1790,2020,10):
# Census_raw[year]=pd.read_html('https://en.wikipedia.org/wiki/{}_United_States_Census'.format(year))
# Census_raw[2019]=pd.read_html('https://en.wikipedia.org/wiki/List_of_states_and_territories_of_the_United_States_by_population')
# -
# Once, we have parsed the data for the first time, we save it to disk and when we run our notebook the next time, we can skip the parsing step.
if len(Census_raw.keys()):
with open('rawdata.pickle', 'wb') as handle:
pickle.dump(Census_raw, handle, protocol=pickle.HIGHEST_PROTOCOL)
else:
with open('rawdata.pickle', 'rb') as handle:
Census_raw = pickle.load(handle)
# ## Data Cleaning
#
# All Census pages contain several tables but we are only interested in those tables that contain `State` (or `District` as in [1800](https://en.wikipedia.org/wiki/1800_United_States_Census)) as one of their columns. Most pages also have a seperate statistic on largest cities and we thus ignore all tables that have `City` as one of their columns.
Census = {}
for key in Census_raw.keys():
Census[key] = [table for table in Census_raw[key] if ('State' in table.columns) or (('District' in table.columns))]
if len(Census[key]) > 1:
Census[key] = [table for table in Census[key] if ('City' not in table.columns)][0]
else:
Census[key] = Census[key][0]
# After these relatively easy manipulations, we need to get our hands dirty and make sure that all tables are of the correct format. Often, it is best to visualize the tables (or at least the first entry) to get a better idea. So let's do that.
for key in Census.keys():
print("Displaying values for {}".format(key))
display(Census[key].head(1))
# Ok, so it looks like 1790 and 1800 use `Total` instead of `Population` and 1800 also uses `District` instead of `State`. Let's change that!
Census[1790].rename(columns = {'Total':'Population'}, inplace=True)
Census[1800].rename(columns = {'Total':'Population', 'District':'State'}, inplace=True)
# Also, from 1990 to 2010, the population of the previous decade is listed and we need to make sure we get the right one. For this task, we use regular expressions to select the column that contains the current year. Once we have found the right column, we rename it to `Population`. For 2019, we need to change it from `Population estimate, July 1, 2019[5]` to `Population`.
for year in range(1990,2020,10):
d = {'Population as of{}.*'.format(year): 'Population'}
Census[year].columns = Census[year].columns.to_series().replace(d, regex=True)
Census[2019].columns = Census[2019].columns.to_series().replace({'Population estimate, July 1, 2019.*': 'Population'}, regex=True)
# Next, we get rid of all columns other than `State` and `Population`. We also remove all non-numerical characters from the `Population` column with some regular expression magic. Once we have done these two steps, we display the first 8 entries of all tables.
for key in Census.keys():
print("Displaying values for {}".format(key))
Census[key] = Census[key][['State','Population']]
Census[key].loc[:,'Population']=Census[key].loc[:,'Population'].apply(
lambda s: int(re.sub(r'\[.*','', str(s)).replace(',','')))
display(Census[key].head(8))
# ## The year 1800 😒
display(Census[1800].head(10))
# Unfortunately, the year 1800 doesn't play well with the other years and requires some additional cleaning up (that's why parsing is not recommended). We can see that instead of `States`, the 1800 census data has smaller subdivisions. To remedy this, we match our list of the 50+1 states against the district names and rename them to a single state. For example, our row with `New York (Duchess, Ulster, Orange counties)` will just become `New York`. Once we have renamed them, we group them by state and sum up the populations on all rows that share the same state. Just to be safe, we apply the same action to every census year. Grouping also has the nice side effect of setting the index to our column that we group by (`State`). Finally, we rename our `Population` column to the census year.
# +
States = pd.read_csv('States.csv')['States']
StatesList = States.tolist()
if isinstance(Census[1790].index[0], int):
for key in Census.keys():
for i, row in Census[key].iterrows():
for State in StatesList:
if State in row['State']:
Census[key].loc[i,'State'] = State
Census[key] = Census[key][Census[key]['State'].isin(StatesList)]
Census[key] = Census[key].groupby(['State'],squeeze=True).sum()
Census[key].rename(columns={'Population':str(key)},inplace=True)
display(Census[1800].head(14))
# -
# It worked! Great, now we are ready to put everything in one big table and make some final checks.
#
result_df = pd.DataFrame(States).set_index('States')
for key in Census.keys():
result_df = result_df.add(Census[key],fill_value=0)
result_df.to_csv('State_Census_Historical.csv')
result_df.tail(5)
# It looks like some years have missing information because records were lost. For instance, there is no value for `West Virginia` in 1860. To fix this, we interpolate all missing values.
result_df.interpolate(axis=1, inplace = True)
result_df.tail()
# We can see how `West Virginia`'s 1860 result was successfully interpolated between its neighbors. Now we are ready to go and visualize the census data over the past 230 years!
# ## Data Visualization
# Getting and cleaning the data was by far the hardest part. Now that we got this out of the way, we can use a visualization tool such as Flourish to get our final **US Census Bar Chart Race** which is available at https://public.flourish.studio/visualisation/1322083.
#
# Congrats on completing this tutorial and I hope it can help you in your next data science project!
HTML("<iframe src='https://public.flourish.studio/visualisation/1322083/embed' frameborder='0' scrolling='no' style='width:100%;height:600px;'></iframe><div style='width:100%!;margin-top:4px!important;text-align:right!important;'><a class='flourish-credit' href='https://public.flourish.studio/visualisation/1322083/?utm_source=embed&utm_campaign=visualisation/1322083' target='_top' style='text-decoration:none!important'><img alt='Made with Flourish' src='https://public.flourish.studio/resources/made_with_flourish.svg' style='width:105px!important;height:16px!important;border:none!important;margin:0!important;'> </a></div>")
# ## Licensing
# This work is licensed under a [Creative Commons Attribution 4.0 International
# License][cc-by]. This means that you are free to:
#
# * **Share** — copy and redistribute the material in any medium or format
# * **Adapt** — remix, transform, and build upon the material for any purpose, even commercially.
#
# Under the following terms:
#
# * **Attribution** — a link to this [github repo](https://github.com/AntonMu/Census2020)!
#
# [![CC BY 4.0][cc-by-image]][cc-by]
#
# [cc-by]: http://creativecommons.org/licenses/by/4.0/
# [cc-by-image]: https://i.creativecommons.org/l/by/4.0/88x31.png
# [cc-by-shield]: https://img.shields.io/badge/License-CC%20BY%204.0-lightgrey.svg
| USCensus2020.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: fe_test
# language: python
# name: fe_test
# ---
# ## Outlier Engineering
#
#
# An outlier is a data point which is significantly different from the remaining data. “An outlier is an observation which deviates so much from the other observations as to arouse suspicions that it was generated by a different mechanism.” [<NAME>. Identification of Outliers, Chapman and Hall , 1980].
#
# Statistics such as the mean and variance are very susceptible to outliers. In addition, **some Machine Learning models are sensitive to outliers** which may decrease their performance. Thus, depending on which algorithm we wish to train, we often remove outliers from our variables.
#
# We discussed in section 3 of this course how to identify outliers. In this section, we we discuss how we can process them to train our machine learning models.
#
#
# ## How can we pre-process outliers?
#
# - Trimming: remove the outliers from our dataset
# - Treat outliers as missing data, and proceed with any missing data imputation technique
# - Discrestisation: outliers are placed in border bins together with higher or lower values of the distribution
# - Censoring: capping the variable distribution at a max and / or minimum value
#
# **Censoring** is also known as:
#
# - top and bottom coding
# - windsorisation
# - capping
#
#
# ## Censoring or Capping.
#
# **Censoring**, or **capping**, means capping the maximum and /or minimum of a distribution at an arbitrary value. On other words, values bigger or smaller than the arbitrarily determined ones are **censored**.
#
# Capping can be done at both tails, or just one of the tails, depending on the variable and the user.
#
# Check my talk in [pydata](https://www.youtube.com/watch?v=KHGGlozsRtA) for an example of capping used in a finance company.
#
# The numbers at which to cap the distribution can be determined:
#
# - arbitrarily
# - using the inter-quantal range proximity rule
# - using the gaussian approximation
# - using quantiles
#
#
# ### Advantages
#
# - does not remove data
#
# ### Limitations
#
# - distorts the distributions of the variables
# - distorts the relationships among variables
#
#
# ## In this Demo
#
# We will see how to perform capping with the quantiles using the Boston House Dataset
#
# ## Important
#
# When doing capping, we tend to cap values both in train and test set. It is important to remember that the capping values MUST be derived from the train set. And then use those same values to cap the variables in the test set
#
# I will not do that in this demo, but please keep that in mind when setting up your pipelines
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# for Q-Q plots
import scipy.stats as stats
# boston house dataset for the demo
from sklearn.datasets import load_boston
from feature_engine.outlier_removers import Winsorizer
# +
# load the the Boston House price data
# load the boston dataset from sklearn
boston_dataset = load_boston()
# create a dataframe with the independent variables
# I will use only 3 of the total variables for this demo
boston = pd.DataFrame(boston_dataset.data,
columns=boston_dataset.feature_names)[[
'RM', 'LSTAT', 'CRIM'
]]
# add the target
boston['MEDV'] = boston_dataset.target
boston.head()
# +
# function to create histogram, Q-Q plot and
# boxplot. We learned this in section 3 of the course
def diagnostic_plots(df, variable):
# function takes a dataframe (df) and
# the variable of interest as arguments
# define figure size
plt.figure(figsize=(16, 4))
# histogram
plt.subplot(1, 3, 1)
sns.distplot(df[variable], bins=30)
plt.title('Histogram')
# Q-Q plot
plt.subplot(1, 3, 2)
stats.probplot(df[variable], dist="norm", plot=plt)
plt.ylabel('RM quantiles')
# boxplot
plt.subplot(1, 3, 3)
sns.boxplot(y=df[variable])
plt.title('Boxplot')
plt.show()
# +
# let's find outliers in RM
diagnostic_plots(boston, 'RM')
# +
# visualise outliers in LSTAT
diagnostic_plots(boston, 'LSTAT')
# +
# outliers in CRIM
diagnostic_plots(boston, 'CRIM')
# -
# There are outliers in all of the above variables. RM shows outliers in both tails, whereas LSTAT and CRIM only on the right tail.
#
# To find the outliers, let's re-utilise the function we learned in section 3:
def find_boundaries(df, variable):
# the boundaries are the quantiles
lower_boundary = df[variable].quantile(0.05)
upper_boundary = df[variable].quantile(0.95)
return upper_boundary, lower_boundary
# +
# find limits for RM
RM_upper_limit, RM_lower_limit = find_boundaries(boston, 'RM')
RM_upper_limit, RM_lower_limit
# +
# limits for LSTAT
LSTAT_upper_limit, LSTAT_lower_limit = find_boundaries(boston, 'LSTAT')
LSTAT_upper_limit, LSTAT_lower_limit
# +
# limits for CRIM
CRIM_upper_limit, CRIM_lower_limit = find_boundaries(boston, 'CRIM')
CRIM_upper_limit, CRIM_lower_limit
# +
# Now let's replace the outliers by the maximum and minimum limit
boston['RM']= np.where(boston['RM'] > RM_upper_limit, RM_upper_limit,
np.where(boston['RM'] < RM_lower_limit, RM_lower_limit, boston['RM']))
# +
# Now let's replace the outliers by the maximum and minimum limit
boston['LSTAT']= np.where(boston['LSTAT'] > LSTAT_upper_limit, LSTAT_upper_limit,
np.where(boston['LSTAT'] < LSTAT_lower_limit, LSTAT_lower_limit, boston['LSTAT']))
# +
# Now let's replace the outliers by the maximum and minimum limit
boston['CRIM']= np.where(boston['CRIM'] > CRIM_upper_limit, CRIM_upper_limit,
np.where(boston['CRIM'] < CRIM_lower_limit, CRIM_lower_limit, boston['CRIM']))
# +
# let's explore outliers in the trimmed dataset
# for RM we see much less outliers as in the original dataset
diagnostic_plots(boston, 'RM')
# -
diagnostic_plots(boston, 'LSTAT')
diagnostic_plots(boston, 'CRIM')
# We can see that the outliers are gone, but the variable distribution was distorted quite a bit.
# ## Censoring with feature-engine
# +
# load the the Boston House price data
# load the boston dataset from sklearn
boston_dataset = load_boston()
# create a dataframe with the independent variables
# I will use only 3 of the total variables for this demo
boston = pd.DataFrame(boston_dataset.data,
columns=boston_dataset.feature_names)[[
'RM', 'LSTAT', 'CRIM'
]]
# add the target
boston['MEDV'] = boston_dataset.target
boston.head()
# +
# create the capper
windsoriser = Winsorizer(distribution='quantiles', # choose from skewed, gaussian or quantiles
tail='both', # cap left, right or both tails
fold=0.05,
variables=['RM', 'LSTAT', 'CRIM'])
windsoriser.fit(boston)
# -
boston_t = windsoriser.transform(boston)
diagnostic_plots(boston, 'RM')
diagnostic_plots(boston_t, 'RM')
# we can inspect the minimum caps for each variable
windsoriser.left_tail_caps_
# we can inspect the maximum caps for each variable
windsoriser.right_tail_caps_
| Section-09-Outlier-Engineering/09.04-Capping-Quantiles.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Boston(42.3601° N, 71.0589° W) is located on the east coast of the US, whereas Seattle(47.6062° N, 122.3321° W) is located on the west coast.
#
# If we compare the weathers, this is what we see:<br>
# Boston, Massachusetts <br>
# Summer High: the July high is around 82.4 degrees<br>
# Winter Low: the January low is 19.2<br>
# Rain: averages 47.4 inches of rain a year<br>
# Snow: averages 48.1 inches of snow a year<br>
#
# Seattle, Washington <br>
# Summer High: the July high is around 75.8 degrees<br>
# Winter Low: the January low is 37<br>
# Rain: averages 38 inches of rain a year<br>
# Snow: averages 4.6 inches of snow a year<br>
# Source: https://www.bestplaces.net/climate/?c1=55363000&c2=52507000<br>
#
#
# # 1. Business Understanding
# Since, they are located almost in the same latitude, and bost are coastal cities, it's worth comparing the cities and find out which is a better place to travel?<br>
# As we can see, **Seattle** is a slightly more comfortable place to visit. I used Airbnb data to find out travelling pattern for the cities. Let’s see what we can infer using the Airbnb data.
#
# This projects aims to solve 4 business questions related to Boston and Seattle data.
#
# **Question 1:** Which place is cheaper to stay? Boston or Seattle. <br>
# If we compare the average housing price between Boston and Seattle, we can determine which place is cheaper.
#
# **Question 2:** What is the best time to visit these places?<br>
# We can find out the pricing change throughout the year.
#
# **Question 3:** Which city is more popular with visitors?<br>
# If we ca find out the monthly availibility of the properties throughout the year, we can answer this question.
#
# **Question 4:** What are the popular areas to stay and what is the recommended type of housing?<br>
# We need to find out the pricing change with the property type, bedrooms, bathrooms, neighbourhood, zipcode etc to determine the popular areas. In this project I tried to find out the relation between the neighbourhood and price and review vs price.
# For the second part we can investigate the relation between property type and price.
# # 2. Data Understanding
# +
# Import useful libraries
# for computation
import numpy as np
import pandas as pd
# for visualization
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# -
# #### Gather the Data
# There are four types of datasets used for this project. "Boston_listing.csv", "Seattle_listings.csv", "Boston_calendar.csv" and "Seattle_calendar.csv".
# +
# reading the dataset
# get the data from here : http://insideairbnb.com/get-the-data.html
# Boston
boston_calendar_sep2016 = pd.read_csv('./data/Boston_calendar_sep2016.csv')
boston_calendar_oct2017 = pd.read_csv('./data/Boston_calendar_oct2017.csv')
boston_calendar_apr2018 = pd.read_csv('./data/Boston_calendar_apr2018.csv')
boston_calendar_jan2019 = pd.read_csv('./data/Boston_calendar_jan2019.csv')
boston_calendar_jan2020 = pd.read_csv('./data/Boston_calendar_jan2020.csv')
boston_listing = pd.read_csv('./data/Boston_listings_jan2020.csv')
# reading the dataset
# get the data from here : http://insideairbnb.com/get-the-data.html
# Seattle
seattle_calendar_jan2016 = pd.read_csv('./data/Seattle_calendar_jan2016.csv')
seattle_calendar_apr2018 = pd.read_csv('./data/Seattle_calendar_apr2018.csv')
seattle_calendar_jan2019 = pd.read_csv('./data/Seattle_calendar_jan2019.csv')
seattle_calendar_jan2020 = pd.read_csv('./data/Seattle_calendar_jan2020.csv')
seattle_listing = pd.read_csv('./data/Seattle_listings_jan2020.csv')
# -
# #### Assess the data
#
# Let's look at one of the boston_calendar and one of the seattle_calnedar data first. There 4 boston_calendar data and 4 seattle_calnedar data taken at different years. Let's check the first 5 rows of one of the datasets (one for Boston one for Seattle)
boston_calendar_oct2017.head()
# rows and columns
print (f"boston_calendar_oct2017 has {boston_calendar_oct2017.shape[0]} rows and {boston_calendar_oct2017.shape[1]} columns")
print ( "Columns are = ", list(boston_calendar_oct2017.columns))
seattle_calendar_jan2016.head()
print (f"seattle_calendar_oct2017 has {seattle_calendar_jan2016.shape[0]} rows and {seattle_calendar_jan2016.shape[1]} columns")
print ( "Columns are = ", list(seattle_calendar_jan2016.columns))
# Which columns had no missing values? Provide a set of column names that have no missing values.
# +
boston_calendar_no_nulls = set(boston_calendar_oct2017.columns[boston_calendar_oct2017.isnull().mean()==0])
boston_calendar_most_nulls = set(boston_calendar_oct2017.columns[boston_calendar_oct2017.isnull().mean() > 0.75])
print("boston_calendar data columns with no missing values \n",boston_calendar_no_nulls) #columns with no missing value
print("\nboston_calendar data columns with atleast values \n",boston_calendar_most_nulls) #columns with atleast 75% missing value
# +
seattle_calendar_no_nulls = set(seattle_calendar_jan2016.columns[seattle_calendar_jan2016.isnull().mean()==0])
seattle_calendar_most_nulls = set(seattle_calendar_jan2016.columns[seattle_calendar_jan2016.isnull().mean() > 0.75])
print("seattle_calendar data columns with no missing values \n",seattle_calendar_no_nulls) #columns with no missing value
print("\n seattle_calendar data columns with atleast values \n",seattle_calendar_most_nulls) #columns with atleast 75% missing value
# -
boston_calendar_oct2017.available.value_counts() # Categorical values
seattle_calendar_jan2016.available.value_counts() # Categorical values
# As we can see, boston_calendar_oct2017 and seattle_calendar_jan2016 has no missing values in **'date', 'available', 'listing_id'** columns.<br>
# **available** column has 2 categorical features **t** means available and **f** means not available.<br>
# Also, the **price** column is NAN is the housing is not available.<br>
# **listing_id** corresponds to host category.
# Let's look at one of the boston_listing and one of the seattle_listing data
boston_listing.head()
print (f"boston_listing has {boston_listing.shape[0]} rows and {boston_listing.shape[1]} columns. \n")
print ( "Columns are = ", list(boston_listing.columns))
seattle_listing.head()
print (f"seattle_listing has {seattle_listing.shape[0]} rows and {seattle_listing.shape[1]} columns. \n")
print ( "Columns are = ", list(seattle_listing.columns))
# As we can see in terms of number of columns, **boston_calendar and seattle_calendar** are identical. Similarly **boston_listing and seattle_listing** are identical.
# Which columns had no missing values? Provide a set of column names that have no missing values.
# +
boston_listing_no_nulls = set(boston_listing.columns[boston_listing.isnull().mean()==0])
boston_listing_most_nulls = set(boston_listing.columns[boston_listing.isnull().mean() > 0.75])
print("boston_listing data columns with no missing values \n",boston_listing_no_nulls) #columns with no missing value
print("\nboston_listing data columns with atleast values \n",boston_listing_most_nulls) #columns with atleast 75% missing value
# +
seattle_listing_no_nulls = set(seattle_listing.columns[seattle_listing.isnull().mean()==0])
seattle_listing_most_nulls = set(seattle_listing.columns[seattle_listing.isnull().mean() > 0.75])
print("seattle_listing data columns with no missing values \n",seattle_listing_no_nulls) #columns with no missing value
print("\n seattle_listing data columns with atleast values \n",seattle_listing_most_nulls) #columns with atleast 75% missing value
# -
# It's better to ignore **'monthly_price', 'neighbourhood_group_cleansed','square_feet', 'weekly_price', 'host_acceptance_rate', 'medium_url', 'thumbnail_url', 'xl_picture_url'** columns from boston_listing and seattle_listing as more than 75% values are missing.
# #### Data insight
#
# For the first 3 business understanding questions we only need boston_calendar/seattle_calendar dataset with **available, date and price** columns.<br>
# there are missing values in the price column. I will discuss how to handle with that.
#
#
# For the last question, we can use **"neighbourhood_cleansed" and "price"** and **"review_scores_rating" and "price"** columns to find out the popular areas and **"property_type" and "price"** columns to find out the recommended housing.<br>
#
# **The good news is there are no missing values! except "review_scores_rating" :)**
# # 3. Data Preparation
# First, we have to remove the '$' sign of the **price** column and convert srting into float
# +
# function to remove $ sign and return float
def str_to_float(string):
"""
INPUT
string - string of the price (ex : $250.00)
OUTPUT
float - returns float value of the price (ex: 250.0)
"""
if string[:1] == '$':
return float(string[1:].replace(',', ''))
else:
return np.nan
# -
# We can extract the month and year from the **date** column
# ### Data Cleaning
#
# ### Boston/Seattle calendar data
# +
# Boston
# Sep 2016
boston_calendar_sep2016["price"] = boston_calendar_sep2016["price"].dropna().apply(str_to_float)
boston_calendar_sep2016["year"] = pd.DatetimeIndex(boston_calendar_sep2016['date']).year # exracting year
boston_calendar_sep2016["month"] = pd.DatetimeIndex(boston_calendar_sep2016['date']).month # extracting month
# Oct 2017
boston_calendar_oct2017["price"] = boston_calendar_oct2017["price"].dropna().apply(str_to_float)
boston_calendar_oct2017["year"] = pd.DatetimeIndex(boston_calendar_oct2017['date']).year
boston_calendar_oct2017["month"] = pd.DatetimeIndex(boston_calendar_oct2017['date']).month
# Apr2018
boston_calendar_apr2018["price"] = boston_calendar_apr2018["price"].dropna().apply(str_to_float)
boston_calendar_apr2018["year"] = pd.DatetimeIndex(boston_calendar_apr2018['date']).year
boston_calendar_apr2018["month"] = pd.DatetimeIndex(boston_calendar_apr2018['date']).month
# Jan 2019
boston_calendar_jan2019["price"] = boston_calendar_jan2019["price"].dropna().apply(str_to_float)
boston_calendar_jan2019["year"] = pd.DatetimeIndex(boston_calendar_jan2019['date']).year
boston_calendar_jan2019["month"] = pd.DatetimeIndex(boston_calendar_jan2019['date']).month
# Jan 2020
boston_calendar_jan2020["price"] = boston_calendar_jan2020["price"].dropna().apply(str_to_float)
boston_calendar_jan2020["year"] = pd.DatetimeIndex(boston_calendar_jan2020['date']).year
boston_calendar_jan2020["month"] = pd.DatetimeIndex(boston_calendar_jan2020['date']).month
# boston listing
boston_listing["price"] = boston_listing["price"].apply(str_to_float)
# Seattle
# Jan 2016
seattle_calendar_jan2016["price"] = seattle_calendar_jan2016["price"].dropna().apply(str_to_float)
seattle_calendar_jan2016["year"] = pd.DatetimeIndex(seattle_calendar_jan2016['date']).year
seattle_calendar_jan2016["month"] = pd.DatetimeIndex(seattle_calendar_jan2016['date']).month
# Apr2018
seattle_calendar_apr2018["price"] = seattle_calendar_apr2018["price"].dropna().apply(str_to_float)
seattle_calendar_apr2018["year"] = pd.DatetimeIndex(seattle_calendar_apr2018['date']).year
seattle_calendar_apr2018["month"] = pd.DatetimeIndex(seattle_calendar_apr2018['date']).month
# Jan 2019
seattle_calendar_jan2019["price"] = seattle_calendar_jan2019["price"].dropna().apply(str_to_float)
seattle_calendar_jan2019["year"] = pd.DatetimeIndex(seattle_calendar_jan2019['date']).year
seattle_calendar_jan2019["month"] = pd.DatetimeIndex(seattle_calendar_jan2019['date']).month
# Jan 2020
seattle_calendar_jan2020["price"] = seattle_calendar_jan2020["price"].dropna().apply(str_to_float)
seattle_calendar_jan2020["year"] = pd.DatetimeIndex(seattle_calendar_jan2020['date']).year
seattle_calendar_jan2020["month"] = pd.DatetimeIndex(seattle_calendar_jan2020['date']).month
# Seattle listing
seattle_listing["price"] = seattle_listing["price"].apply(str_to_float)
# -
# Our plan is to see how much the yearly average pricing has changed from the year 2016-2020 and investigate any general trend. <br>
#
# Let's extract the yearly data from the above dataset. We only need **year ,month, available, and price** columns
#
#
# We are going to filter the Boston/Seattle data w.r.t year(2017-2020 for Boston and 2016,2018-2020 for seattle as 2017 was not available in Airbnb database)
#
# +
# Boston
# Year 2017
boston_2017_p1 = boston_calendar_sep2016[boston_calendar_sep2016["year"] == 2017]
boston_2017_p2 = boston_calendar_oct2017[boston_calendar_oct2017["year"] == 2017]
boston_2017 = pd.concat([boston_2017_p1,boston_2017_p2], axis = 0)
boston_2017 = boston_2017[["year" ,"month", "available" ,"price"]]
# Year 2018
boston_2018_p1 = boston_calendar_oct2017[boston_calendar_oct2017["year"] == 2018]
boston_2018_p2 = boston_calendar_apr2018[boston_calendar_apr2018["year"] == 2018]
boston_2018 = pd.concat([boston_2018_p1,boston_2018_p2], axis = 0)
boston_2018 = boston_2018[["year" ,"month", "available" ,"price"]]
# Year 2019
boston_2019_p1 = boston_calendar_apr2018[boston_calendar_apr2018["year"] == 2019]
boston_2019_p2 = boston_calendar_jan2019[boston_calendar_jan2019["year"] == 2019]
boston_2019 = pd.concat([boston_2019_p1,boston_2019_p2], axis = 0)
boston_2019 = boston_2019[["year" ,"month", "available" ,"price"]]
# Year 2020
boston_2020_p1 = boston_calendar_jan2019[boston_calendar_jan2019["year"] == 2020]
boston_2020_p2 = boston_calendar_jan2020[boston_calendar_jan2020["year"] == 2020]
boston_2020 = pd.concat([boston_2020_p1,boston_2020_p2], axis = 0)
boston_2020 = boston_2020[["year" ,"month", "available" ,"price"]]
# Seattle
# Year 2016
seattle_2016 = seattle_calendar_jan2016[seattle_calendar_jan2016["year"] == 2016]
seattle_2016 = seattle_2016[["year" ,"month", "available" ,"price"]]
# Year 2018
seattle_2018 = seattle_calendar_apr2018[seattle_calendar_apr2018["year"] == 2018]
seattle_2018 = seattle_2018[["year" ,"month", "available" ,"price"]]
# Year 2019
seattle_2019_p1 = seattle_calendar_apr2018[seattle_calendar_apr2018["year"] == 2019]
seattle_2019_p2 = seattle_calendar_jan2019[seattle_calendar_jan2019["year"] == 2019]
seattle_2019 = pd.concat([seattle_2019_p1,seattle_2019_p2], axis = 0)
seattle_2019 = seattle_2019[["year" ,"month", "available" ,"price"]]
# Year 2020
seattle_2020_p1 = seattle_calendar_jan2019[seattle_calendar_jan2019["year"] == 2020]
seattle_2020_p2 = seattle_calendar_jan2020[seattle_calendar_jan2020["year"] == 2020]
seattle_2020 = pd.concat([seattle_2020_p1,seattle_2020_p2], axis = 0)
seattle_2020 = seattle_2020[["year" ,"month", "available" ,"price"]]
# -
# ### Handling categorical values in Boston/Seattle calendar data ("available" column)
# Since **available** column has 2 values "t"(available) and "f"(not-available) Let's expand the **available** column using one-hot encoding
# +
# Boston
# 2017
boston_one_hot_2017 = pd.get_dummies(boston_2017['available'])
boston_2017 = boston_2017.join(boston_one_hot_2017)
# 2018
boston_one_hot_2018 = pd.get_dummies(boston_2018['available'])
boston_2018 = boston_2018.join(boston_one_hot_2018)
#2019
boston_one_hot_2019 = pd.get_dummies(boston_2019['available'])
boston_2019 = boston_2019.join(boston_one_hot_2019)
#2020
boston_one_hot_2020 = pd.get_dummies(boston_2020['available'])
boston_2020 = boston_2020.join(boston_one_hot_2020)
# Seattle
# 2016
seattle_one_hot_2016 = pd.get_dummies(seattle_2016['available'])
seattle_2016 = seattle_2016.join(seattle_one_hot_2016)
# 2018
seattle_one_hot_2018 = pd.get_dummies(seattle_2018['available'])
seattle_2018 = seattle_2018.join(seattle_one_hot_2018)
#2019
seattle_one_hot_2019 = pd.get_dummies(seattle_2019['available'])
seattle_2019 = seattle_2019.join(seattle_one_hot_2019)
#2020
seattle_one_hot_2020 = pd.get_dummies(seattle_2020['available'])
seattle_2020 = seattle_2020.join(seattle_one_hot_2020)
# -
# This will create two new columns **t** and **f** where, **t** = 1 means the housing is available and **f** = 1 means it's not available
# ### Merge the calendar data¶
#
# Finally we will merge all all boston and seattle calendar data as **boston_total** and **seattle_total** respectively
# +
boston_total = pd.concat([boston_2017,boston_2018,boston_2019,boston_2020], axis = 0)
seattle_total = pd.concat([seattle_2016,seattle_2018,seattle_2019,seattle_2020], axis = 0)
# +
# Free up some memory
boston_calendar_sep2016 = []
boston_calendar_oct2017 = []
boston_calendar_apr2018 = []
boston_calendar_jan2019 = []
boston_calendar_jan2020 = []
boston_one_hot_2017 = []
boston_one_hot_2018 = []
boston_one_hot_2019 = []
boston_one_hot_2020 = []
boston_2017 = []
boston_2018 = []
boston_2019 = []
boston_2020 = []
# Free up some memory
seattle_calendar_sep2016 = []
seattle_calendar_apr2018 = []
seattle_calendar_jan2019 = []
seattle_calendar_jan2020 = []
seattle_one_hot_2016 = []
seattle_one_hot_2019 = []
seattle_one_hot_2020 = []
seattle_2018 = []
seattle_2019 = []
# -
# ### Handling missing values
#
# Let's see how much missing values does boston_total and seattle_total's **price** has, since this is the only column with missing values
# Also, boston_listing and seattle_listing has **review_scores_rating** which can be used to find recommended housing. We have to look for missing values in this column as well.
# +
boston_missing_value = boston_total.price.isnull().mean()
seattle_missing_value = seattle_total.price.isnull().mean()
boston_review_missing_value = boston_listing.review_scores_rating.isnull().mean()
seattle_review_missing_value = seattle_listing.review_scores_rating.isnull().mean()
print(f"boston_total has {boston_missing_value*100 : .2f}% missing value in the price column")
print(f"seattle_total has {seattle_missing_value*100 : .2f}% missing value in the price column")
print(f"boston_listing has {boston_review_missing_value*100 : .2f}% missing value in the review_scores_rating column")
print(f"seattle_listing has {seattle_review_missing_value*100 : .2f}% missing value in the review_scores_rating column")
# -
# Since both **boston_total and seattle_total "price"** columns have less than 40% missing values and we will mostly work with average price, it's safe to just **drop** the corresponding rows whenever necessary.<br>
# Similarly both **boston_listing and seattle_listing "review_scores_rating"** columns have less than 20% missing values and we will mostly work with average price, it's safe to just **drop** the corresponding rows whenever necessary.<br>
# The only disadvantage will be, we'll have lesser data points. One possible solution could be to download more data from Airbnb wesite. But we have enough dataset to usderstand the general trend
# #### Boston/Seattle listing data
#
# we only need the **"neighbourhood_cleansed","property_type" and "price"** columns for this project. So let's just pick those and drop the rest. These columns have no missing values!
boston_listing = boston_listing[["neighbourhood_cleansed","property_type", "price",'review_scores_rating' ]]
seattle_listing = seattle_listing[["neighbourhood_cleansed","property_type", "price",'review_scores_rating']]
# ## 4. Analysis, Modeling, Visualization
# We can ask 4 business questions from the data. Answer to each question can be analyzed through visualization
# ## Question 1: Which place is cheaper to stay? Boston or Seattle.
# To answer this question I downloaded the Boston and Seattle data for the past few years and calculated the average pricing for each year. The following figures explain it.
figure, axes = plt.subplots(1, 2,figsize=(10,5))
seattle_total.groupby(["month","year"]).price.mean().unstack().plot(kind='box',title = "Seattle", ax = axes[1]);
boston_total.groupby(["month","year"]).price.mean().unstack().plot(kind='box',title = "Boston",ax = axes[0]);
axes[0].set_ylabel("Price in USD")
axes[0].set_xlabel("Year");
axes[1].set_xlabel("Year");
#plt.savefig('avg_price.png')
# The graph shows the mean price and spread of price (standard diviation) for each year
# Average housing price in Boston by year
boston_total.groupby(["year"]).price.mean()
# Average housing price in Seattle by year
seattle_total.groupby(["year"]).price.mean()
# So, from the above graph we can see that Seattle is cheaper to visit. For the past two years the average price of seattle is **159 dollars** whereas, the average price of Boston is **206 dollars**
# ## Question 2: What is the best time to visit these places?
# To address this question, we have to look into the availability of Airbnb housing and price change during the season.
# +
# average price of boston from 2017-2020
print (f"average price of boston from 2017-2020 is {boston_total.price.mean() :.0f} USD")
# average price of seattle from 2016-2020
print (f"average price of seattle from 2016-2020 is {seattle_total.price.mean() :.0f} USD")
# +
figure, axes = plt.subplots(1, 2,figsize=(10,5))
(boston_total.groupby(["month"]).price.mean() - boston_total.price.mean()).plot(kind='bar', title = "Boston price from 2017-2020",ax = axes[0])
(seattle_total.groupby(["month"]).price.mean() - seattle_total.price.mean()).plot(kind='bar', title = "Seattle price from 2016-2020",ax = axes[1])
axes[0].set_ylabel("change in price by month (in USD)");
axes[0].set_xlabel("Month");
axes[1].set_xlabel("Month");
#plt.savefig('price_change.png')
# -
# Here 0 means the **216 USD** for **Boston**. Anything more above 0 means more expensive during that month <br>
# Here 0 means the **159 USD** for **Seattle**. Anything more above 0 means more expensive during that month
# From this graph, we can see that since Boston has more extreme weather compared to Seattle, Boston has more price variation. Also, from the end of spring to the end of fall (March — November), Boston Airbnb's price is higher than average. So for Boston, **May and July** should be avoided if you want to save money. **March and November** are the best months to visit. During the winter season, **February** has the least price.<br>
# Similarly for Seattle, **July and August** should be avoided and **February** is the cheapest month of the year. From spring to the end of fall **March, April, May, October, and November** are the best months to visit.
# ## Question 3: Which city is more popular and what is the ideal time to visit?
# ### Average occupancy
# +
# Average yearly occupancy of Boston (in percentage)
boston_total.groupby(["year"]).f.mean() * 100
# +
# Average yearly occupancy of Seattle (in percentage)
seattle_total.groupby(["year"]).f.mean() * 100
# +
# average price of boston from 2017-2020
print (f"average occupancy of boston from 2017-2020 is {boston_total.f.mean()*100 :.0f} %") # f = 1 means occupied
# average price of seattle from 2016-2020
print (f"average 0ccupancy of seattle from 2016-2020 is {seattle_total.f.mean()*100 :.0f} %") # f = 1 means occupied
# -
# Over the last few years, the average occupancy of Boston and Seattle is 58% and 59% respectively. which suggests that Seattle is more popular among visitors. So what time people usually try to visit these cities?
# +
figure, axes = plt.subplots(1, 2,figsize=(10,5))
(boston_total.groupby(["month"]).f.mean() - boston_total.f.mean()).plot(kind='bar',title = "Boston occupancy from 2017-2020 (avg 58%)",ax = axes[0]);
(seattle_total.groupby(["month"]).f.mean() - seattle_total.f.mean()).plot(kind='bar',title = "Seattle occupancy from 2016-2020 (avg 58%)",ax = axes[1]);
axes[0].set_ylabel("Change in Availibility from yearly average (in fraction)");
axes[0].set_xlabel("Month");
axes[1].set_xlabel("Month");
#plt.savefig('accupancy.png')
# -
# As we can see, during early winter Boston is more occupied than the yearly average (more than 4% of the yearly average).<br>
# Seattle has more occupancy in **January** and from **July — December**, which also suggests Seattle being more popular among the two.
#
# ## Question 4: What are the popular areas to stay and what is the recommended type of housing?
# ### where are most of the hosts located in Boston?
# +
plt.figure(figsize = (10,10))
host_loc = boston_listing.neighbourhood_cleansed.value_counts()
(host_loc*100/boston_listing.neighbourhood_cleansed.value_counts().dropna().sum()).plot(kind="bar");
plt.ylabel('percentage of the neighbourhood')
plt.xlabel( "Neighborhood")
plt.title("Where are the hosts located in Boston?");
plt.xticks(rotation=45, ha='right');
#plt.savefig('most_host.png')
# +
boston_host_loc = seattle_listing.neighbourhood_cleansed.value_counts() # number of properties by neighbourhood
boston_host_loc[:6].sum()/boston_listing.neighbourhood_cleansed.value_counts().dropna().sum() # fraction of properties in the first 5 neighbourhood
# -
# As we can see from the above graph the more that roughly 50% of the hosing is located in **Dorchester, Downtown, Jamaica Plain, South End, and Back Bay** <br>
# ### So what are the recommended neighborhoods in Boston?
#
# If 75% of the hosts in each neighbourhood are rated more than 8, that neighbourhood can be recommended.
# We can check the rating of these neighbourhoods and find out which neighbourhood has ratings more than 80 out of 100 and we can filter out places with less than 70/100 rating.
above_80 = boston_listing[boston_listing['review_scores_rating'] > 80.0] #data with review above 80/100 rating
below_70 = boston_listing[boston_listing['review_scores_rating'] < 70.0] #data with review less than 70/100 rating
host_loc80 = above_80.neighbourhood_cleansed.value_counts()
(host_loc80*100/host_loc - 75).dropna().plot(kind="bar");
plt.ylabel('Percentage diviation from 75%')
plt.xlabel('Neighbourhood')
plt.title("Neighbourhood with more than 75% hosts rated 8/10");
#plt.savefig('recomended_palce.png')
# Here 0 marks the neighborhood with 75% of the hosts. Above 0 are the recommended places.
# From the graph above the recommended properties are **Allston, Beacon Hill, Charlestown, Dorchester, East Boston, Hyde Park, Jamaica Plain, Leather District, Mattapan, North End, Roslindale, Roxbury, South Boston, South End, West Roxbury.** <br>
# If you prefer a popular neighbourhood, **Jamaica Plain, Dorchester, and South End** is recommended. <br>
# In terms of rating, most places in **Hyde Park, Leather District, Mattapan, Roslindale, West Roxbury** are highly rated, but further research is needed since these places have lesser number of hosts.
#
# ### Which places should be avoided in Boston?
host_loc70 = below_70.neighbourhood_cleansed.value_counts()
(host_loc70/host_loc).dropna().plot(kind="bar");
plt.ylabel('fraction of the hosts\' rated less than 7')
plt.xlabel('Neighbourhood')
plt.title("Neighbourhood with hosts' rated less than 7/10");
#plt.savefig('avoid.png')
# From the graph above We can see that **Downtown and Misson Hill** have the most number of hosts' with less than 7 rating. So these places should be avoided.
# ### What is the recommended type of housing in Boston?
# Property type vs Price
(boston_listing.groupby(["property_type"]).price.mean()).dropna().plot(kind='bar');
plt.ylabel('Price in USD');
#plt.savefig('recomended_housing.png')
# If our goal is to save money, then we see that **Bungalow, Guesthouse, In-law, and Villa** are among the cheaper options to stay in Boston (Costs $150 or less).
# #### Similar analysis on Seattle
# #### where are most of the hosts located in seattle?
# +
plt.figure(figsize=(20,5))
seattle_host_loc = seattle_listing.neighbourhood_cleansed.value_counts().dropna()
(seattle_host_loc*100/seattle_listing.neighbourhood_cleansed.value_counts().dropna().sum()).plot(kind="bar");
plt.ylabel('percentage of the neighbourhood')
plt.title("Where are the hosts located?");
# -
seattle_host_loc[:16].sum()/seattle_listing.neighbourhood_cleansed.value_counts().dropna().sum() # fraction of properties in the first 15 neighbourhood
# First 15 neighbourhoods has 50% of the hosts
#
# As we can see from above graph top 50% neighbourhoods are located in **Broadway, Belltown, Central Business District, Wallingford, and Minor,First Hill, Univeristy District, Pike-Market, ..**
#
# #### So what is the recomended neighbourhood in Seattle?
seattle_above_80 = seattle_listing[seattle_listing['review_scores_rating'] > 80.0]
seattle_below_70 = seattle_listing[seattle_listing['review_scores_rating'] < 70.0]
# We can check the rating of these neighbourhoods and find out which neighbourhood has ratings more than 80 out of 100 and we can filter out places with less than 70/100 rating.
plt.figure(figsize=(20,5))
seattle_host_loc80 = seattle_above_80.neighbourhood_cleansed.value_counts().dropna()
(seattle_host_loc80*100/seattle_host_loc - 90).dropna().plot(kind="bar");
plt.ylabel('Percentage diviation from 90%')
plt.xlabel('Neighbourhood')
plt.title("Neighbourhood with more than 90% hosts rated 8/10");
# Here 0 marks the neighborhood with 90% of the hosts. Above 0 are the recommended places.
#More than 95% housing reted 8/10
[(seattle_host_loc80*100/seattle_host_loc - 95).index[i] for i, value in enumerate(seattle_host_loc80*100/seattle_host_loc - 95) if value > 0]
# Seattle is more spread out than Boston, and most of the neighbourhood is highly rated. <br>
# If 90% of the hosts in each neighbourhood are rated more than 8, that neighbourhood can be rocomended. So the recomended properties are **Arbor Heights, Atlantic, Broadview, Columbia City, Crown Hill, Fremont,Genesee,Georgetown, Harbor Island,Leschi,Loyal Heights,Madison Park,Madrona,Mann,Maple Leaf,Mount Baker,North Admiral,North Beacon Hill,North Delridge,Olympic Hills, Phinney Ridge,Rainier Beach,Ravenna, Seaview, Seward Park,South Beacon Hill,Victory Heights, West Queen Anne, and West Woodland** <br>
# If you prefer poplular neighbourhood, **Fremont, North Beacon Hill, and Mann** is recomended. <br>
# In terms of rating, most places in **Arbor Heights,Crown Hill,Georgetown,Harbor Island,Madrona,Rainier Beach,Seaview** are highly rated, but further research is needed since these palecs have leeser number of hosts.
# From the graph below We can see that **International District,Montlake, South Park, and View Ridge** have the most number of hosts with less than 7 rating. So these places should be avoided.
# #### Which places should be avoided in Seattle?
seattle_host_loc70 = seattle_below_70.neighbourhood_cleansed.value_counts()
(seattle_host_loc70/seattle_host_loc).dropna().plot(kind="bar");
plt.ylabel('fraction of the hosts\' rated less than 7')
plt.xlabel('Neighbourhood')
plt.title("Neighbourhood with hosts' rated less than 7/10");
# Property_type vs Price
#
(seattle_listing.groupby(["property_type"]).price.mean()).dropna().plot(kind='bar');
plt.ylabel('Price in USD');
# From the graph above we see that **Camper/RV, Dome house, Hostel, and Treehouse** are among the cheaper options to stay in Seattle (Costs less than $100).
# ## 5. Evaluation
#
# Let's put all the questions and answers togather and dicuss the the final remarks.
#
# ---
#
# ### Question 1: Which place is cheaper to stay? Boston or Seattle.
#
# **Seattle** is cheaper to visit. For the past two years the average price of seattle is **159 dollars** whereas, the average price of Boston is **206 dollars**.
#
# ---
#
# ### Question 2: What is the best time to visit these places?
#
# From the end of spring to the end of fall (March — November), Boston Airbnb's price is higher than average. So for Boston, **May and July** should be avoided if you want to save money. **March and November** are the best months to visit. During the winter season, **February** has the least price.<br>
# Similarly for Seattle, **July and August** should be avoided and **February** is the cheapest month of the year. From spring to the end of fall **March, April, May, October, and November** are the best months to visit.
#
# ---
#
# ### Question 3: Which city is more popular and what is the ideal time to visit?
#
# Over the last few years, the average occupancy of Boston and Seattle is 58% and 59% respectively. which suggests that Seattle is slightly more popular among visitors. <br>
# During early winter Boston is more occupied than the yearly average (more than 4% of the yearly average).<br>
# Seattle has more occupancy in **January** and from **July — December**.
#
# ---
#
# ### Question 4: What are the popular areas to stay and what is the recommended type of housing?
#
# **Bosoton:**
#
# The recommended properties are **Allston, Beacon Hill, Charlestown, Dorchester, East Boston, Hyde Park, Jamaica Plain, Leather District, Mattapan, North End, Roslindale, Roxbury, South Boston, South End, West Roxbury.** <br>
# If you prefer a popular neighbourhood, **Jamaica Plain, Dorchester, and South End** is recommended. <br>
# In terms of rating, most places in **Hyde Park, Leather District, Mattapan, Roslindale, West Roxbury** are highly rated, but further research is needed since these places have lesser number of hosts.<br>
# **Downtown and Misson Hill** have the most number of hosts' with less than 7 rating. So these places should be **avoided**.
# If our goal is to save money, then we see that **Bungalow, Guesthouse, In-law, and Villa** are among the cheaper options to stay in Boston (Costs $150 or less).
#
# **Seattle:**
#
# the recommended properties are **Arbor Heights, Atlantic, Broadview, Columbia City, Crown Hill, Fremont, Genesee, Georgetown, Harbor Island, Leschi, Loyal Heights, Madison Park, Madrona, Mann, Maple Leaf, Mount Baker, North Admiral, North Beacon Hill, North Delridge, Olympic Hills, Phinney Ridge, Rainier Beach, Ravenna, Seaview, Seward Park, South Beacon Hill, Victory Heights, West Queen Anne, and West Woodland** <br>
# If you prefer a popular neighborhood, **Fremont, North Beacon Hill, and Mann** are recommended.
# Most places in **Arbor Heights, Crown Hill, Georgetown, Harbor Island, Madrona, Rainier Beach, Seaview** are highly rated, but further research is needed since these places have a lesser number of hosts.
# We can see that **Montlake, South Park, and View Ridge** have the most number of hosts with less than 7 ratings. So these places should be **avoided**.<br>
# We see that **Camper/RV, Dome house, Hostel, and Treehouse** are among the cheaper options to stay in Seattle (Costs less than $100).
#
# ---
#
# In conclusion, I found out that in terms of weather conditions and housing prices, Seattle is a better place to travel compared to Boston.
# In future work, I am planning to predict the Airbnb price using machine learning.
#
| What is your next destination? Boston or Seattle!.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.0.1
# language: julia
# name: julia-1.0
# ---
# ### GASS utils
#
# Misc. methods for the GASS
# +
using Distances , Random
using PyCall
using DataFrames
import CSV
rootdir = "/home/stephane/Science/ALMA/ArrayConfig/GASS"
push!(LOAD_PATH,"$rootdir/master/src")
using GASS
import PyPlot
@pyimport astropy.units as u
## directory
datadir = "$rootdir/master/data"
wdir = "$rootdir/products"
plotdir = "$rootdir/products/test"
cd(wdir)
# +
## metatype for population
struct _cfg
arr::AbstractDataFrame
obs::observation
sub::subarrayParameters
wei::weight
ga::GA
end
# -
function _read_input_cfg(inpfile)
res= input_parameters(inpfile)
inpcfg= parse_input(res)
println(inpcfg)
## parameters inputs
res= input_parameters(inpcfg.file_parameters)
paramcfg= parse_parameters(res)
println(paramcfg)
return(inpcfg , paramcfg)
end
# +
## init population ...
function _init_cfg(obs , sub , wei , ga )
arrcfg = CSV.read(obs.Array_Configuration_File, datarow=4 , header=["X" , "Y", "Z" , "diam" , "name"] , delim= " ")
pop = _cfg(arrcfg , obs , sub , wei , ga)
return(pop)
end
# +
macro main(inpfile)
inpcfg , paramcfg = read_input_cfg(inpfile)
cfg = _init_cfg(paramcfg[1] , paramcfg[2], paramcfg[3], paramcfg[4])
end
@main("../master/data/GA_Inputs_O-10.txt.julia")
| notebooks/.ipynb_checkpoints/GA-util-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/richardtml/riiaa-20-aa/blob/master/notebooks/2a_cnn.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# -
# # Red convolucional
#
# #### Bere & <NAME>
#
# En esta libreta veremos un ejemplo de clasificación multiclase de imágenes de dígitos implementando una red convolucional en PyTorch.
#
# Emplearemos un conjunto referencia llamado [MNIST](http://yann.lecun.com/exdb/mnist/) recolectado por [Yann LeCun](http://yann.lecun.com). Está compuesto de imágenes en escala de grises de 28 × 28 píxeles que contienen dígitos entre 0 y 9 escritos a mano. El conjunto cuenta con 60,000 imágenes de entrenamiento y 10,000 de prueba.
#
# 
# ## 1 Preparación
# ### 1.1 Bibliotecas
# + colab={} colab_type="code" id="Ny0L2LzogTN-"
# funciones aleatorias
import random
# tomar n elementos de una secuencia
from itertools import islice as take
# gráficas
import matplotlib.pyplot as plt
# arreglos multidimensionales
import numpy as np
# redes neuronales
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
# imágenes
from skimage import io
# redes neuronales
from torch.utils.data import DataLoader
from torchvision import transforms
from torchvision.datasets import MNIST
# barras de progreso
from tqdm import tqdm
# directorio de datos
DATA_DIR = '../data'
# MNIST
MEAN = (0.1307)
STD = (0.3081)
# tamaño del lote
BATCH_SIZE = 128
# reproducibilidad
SEED = 0
random.seed(SEED)
np.random.seed(SEED)
torch.manual_seed(SEED)
# -
# ### 1.2 Auxiliares
def display_grid(xs, titles, rows, cols):
fig, ax = plt.subplots(rows, cols)
for r in range(rows):
for c in range(cols):
i = r * rows + c
ax[r, c].imshow(xs[i], cmap='gray')
ax[r, c].set_title(titles[i])
ax[r, c].set_xticklabels([])
ax[r, c].set_yticklabels([])
fig.tight_layout()
plt.show()
# + [markdown] colab_type="text" id="MjKxreAkoZeT"
# ## 2 Datos
# -
# ### 2.1 Tuberias de datos con PyTorch
#
# 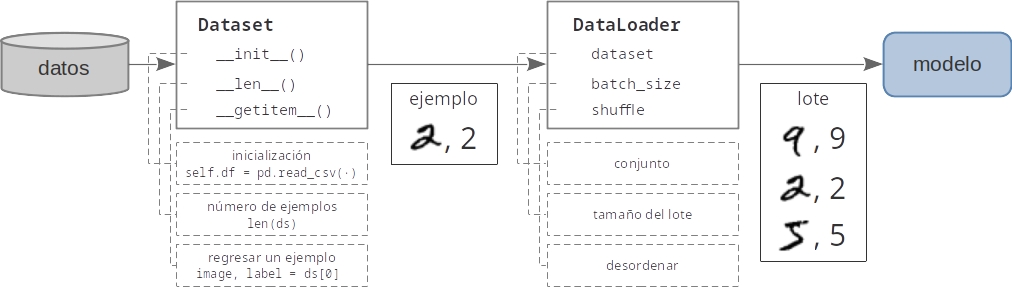
# ### 2.2 Exploración
# creamos un Dataset
ds = MNIST(
# directorio de datos
root=DATA_DIR,
# subconjunto de entrenamiento
train=True,
# convertir la imagen a ndarray
transform=np.array,
# descargar el conjunto
download=True
)
# +
# cargamos algunas imágenes
images, labels = [], []
for i in range(12):
x, y = ds[i]
images.append(x)
labels.append(y)
# desplegamos
print(f'images[0] shape={images[0].shape} dtype={images[0].dtype}')
titles = [str(y) for y in labels]
display_grid(images, titles, 3, 4)
# + [markdown] colab_type="text" id="9p_BsiITogUA"
# ### 2.3 Cargadores de datos
# -
# #### Entrenamiento
# + colab={"base_uri": "https://localhost:8080/", "height": 54} colab_type="code" id="E1aEVpYtuadH" outputId="8df25761-3201-461a-e82b-26b5befd0302"
# transformaciones para la imagen
trn_tsfm = transforms.Compose([
# convertimos a torch.Tensor y escalamos a [0,1]
transforms.ToTensor(),
# estandarizamos: restamos la media y dividimos sobre la varianza
transforms.Normalize(MEAN, STD),
])
# creamos un Dataset
trn_ds = MNIST(
# directorio de datos
root=DATA_DIR,
# subconjunto de entrenamiento
train=True,
# transformación
transform=trn_tsfm
)
# creamos un DataLoader
trn_dl = DataLoader(
# conjunto
trn_ds,
# tamaño del lote
batch_size=BATCH_SIZE,
# desordenar
shuffle=True
)
# desplegamos un lote de imágenes
for x, y in take(trn_dl, 1):
print(f'x shape={x.shape} dtype={x.dtype}')
print(f'y shape={y.shape} dtype={y.dtype}')
# -
# #### Prueba
# + colab={"base_uri": "https://localhost:8080/", "height": 54} colab_type="code" id="QXMXXc9DPgqY" outputId="100e6a58-5552-4c5f-b08e-dc83c23c3b21"
# transformaciones para la imagen
tst_tsfm = transforms.Compose([
# convertimos a torch.Tensor y escalamos a [0,1]
transforms.ToTensor(),
# estandarizamos: restamos la media y dividimos sobre la varianza
transforms.Normalize(MEAN, STD),
])
# creamos un Dataset
tst_ds = MNIST(
# directorio de datos
root=DATA_DIR,
# subconjunto de entrenamiento
train=False,
# transformación
transform=tst_tsfm
)
# creamos un DataLoader
tst_dl = DataLoader(
# subconjunto
tst_ds,
# tamaño del lote
batch_size=BATCH_SIZE,
# desordenar
shuffle=True
)
# imprimimos forma y tipo del lote
for x, y in take(tst_dl, 1):
print(f'x shape={x.shape} dtype={x.dtype}')
print(f'y shape={y.shape} dtype={y.dtype}')
# -
# ## 3 Modelo
#
# 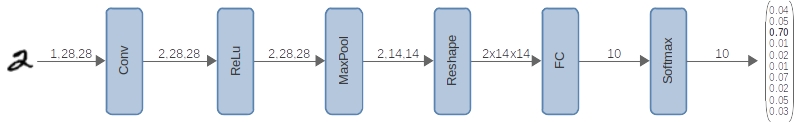
# ### 3.1 Definición de la arquitectura
# definición del modelo
class CNN(nn.Module):
# inicializador
def __init__(self):
# inicilización del objeto padre, obligatorio
super(CNN, self).__init__()
# tamaño de las capas
C1 = 2
FC1 = 10
self.num_feats = 2 * 14 * 14
# capas convolucionales
self.cnn = nn.Sequential(
# bloque conv1
# [N, 1, 28, 28] => [N, 2, 28, 28]
nn.Conv2d(in_channels=1, out_channels=C1, kernel_size=3, padding=1),
# [N, 2, 28, 28]
nn.ReLU(),
# [N, 2, 28, 28] => [N, 2, 14, 14]
nn.MaxPool2d(kernel_size=2, stride=2)
)
# capas completamente conectadas
self.cls = nn.Sequential(
# bloque fc1
# [N, 2x14x14] => [N, 10]
nn.Linear(self.num_feats, FC1),
)
# metodo para inferencia
def forward(self, x):
# [N, 1, 28, 28] => [N, 2, 14, 14]
x = self.cnn(x)
# aplanamos los pixeles de la imagen
# [N, 2, 14, 14] => [N, 2x14x14]
x = x.view(-1, self.num_feats)
# [N, 2x14x14] => [N, 10]
x = self.cls(x)
return x
# ### 3.2 Impresión de la arquitectura
model = CNN()
print(model)
# ### 3.3 Prueba de la arquitectura
# inferencia con datos sinteticos
x = torch.zeros(1, 1, 28, 28)
y = model(x)
print(y.shape)
# ## 4 Entrenamiento
#
# 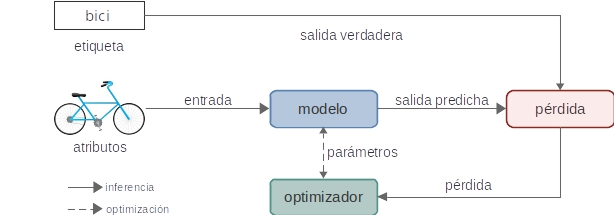
# ### 4.1 Ciclo de entrenamiento
# + colab={"base_uri": "https://localhost:8080/", "height": 568} colab_type="code" id="xCqwGRD1nz1a" outputId="7dc4823c-865a-41ee-b54b-67117e5d4e95"
# creamos un modelo
model = CNN()
# optimizador
opt = optim.SGD(model.parameters(), lr=1e-3)
# historial de pérdida
loss_hist = []
# ciclo de entrenamiento
EPOCHS = 20
for epoch in range(EPOCHS):
# entrenamiento de una época
for x, y_true in trn_dl:
# vaciamos los gradientes
opt.zero_grad()
# hacemos inferencia para obtener los logits
y_lgts = model(x)
# calculamos la pérdida
loss = F.cross_entropy(y_lgts, y_true)
# retropropagamos
loss.backward()
# actulizamos parámetros
opt.step()
# evitamos que se registren las operaciones
# en la gráfica de cómputo
with torch.no_grad():
losses, accs = [], []
# validación de la época
for x, y_true in take(tst_dl, 10):
# hacemos inferencia para obtener los logits
y_lgts = model(x)
# calculamos las probabilidades
y_prob = F.softmax(y_lgts, 1)
# obtenemos la clase predicha
y_pred = torch.argmax(y_prob, 1)
# calculamos la pérdida
loss = F.cross_entropy(y_lgts, y_true)
# calculamos la exactitud
acc = (y_true == y_pred).type(torch.float32).mean()
# guardamos históricos
losses.append(loss.item() * 100)
accs.append(acc.item() * 100)
# imprimimos métricas
loss = np.mean(losses)
acc = np.mean(accs)
print(f'E{epoch:2} loss={loss:6.2f} acc={acc:.2f}')
# agregagmos al historial de pérdidas
loss_hist.append(loss)
# -
# ### 4.2 Gráfica de la pérdida
plt.plot(loss_hist, color='red')
plt.xlabel('época')
plt.ylabel('pérdida')
plt.show()
# ## 5 Evaluación
#
# 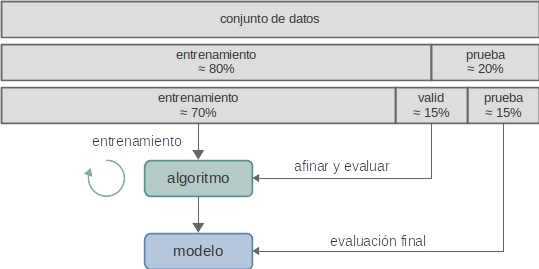
# ### 5.1 Conjunto de validación
# +
# modelo en modo de evaluación
model.eval()
# evitamos que se registren las operaciones
# en la gráfica de cómputo
with torch.no_grad():
accs = []
# validación de la época
for x, y_true in tst_dl:
# hacemos inferencia para obtener los logits
y_lgts = model(x)
# calculamos las probabilidades
y_prob = F.softmax(y_lgts, 1)
# obtenemos la clase predicha
y_pred = torch.argmax(y_prob, 1)
# calculamos la exactitud
acc = (y_true == y_pred).type(torch.float32).mean()
accs.append(acc.item() * 100)
acc = np.mean(accs)
print(f'Exactitud = {acc:.2f}')
# -
# ### 5.2 Inferencia
# +
with torch.no_grad():
for x, y_true in take(tst_dl, 1):
y_lgts = model(x)
y_prob = F.softmax(y_lgts, 1)
y_pred = torch.argmax(y_prob, 1)
x = x[:12].squeeze().numpy()
y_true = y_true[:12].numpy()
y_pred = y_pred[:12].numpy()
titles = [f'V={t} P={p}' for t, p in zip(y_true, y_pred)]
display_grid(x, titles, 3, 4)
| notebooks/2a_cnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/seryeongi/exec_machinlearning/blob/master/wholesalecustomers_kmeans.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="51xtoNUHeb7T" outputId="3cdce381-10d3-4df1-f528-5b4bc61e8b4e"
# !ls #list
# + colab={"base_uri": "https://localhost:8080/"} id="mu3R39ibfY7c" outputId="55b83626-e357-4ebb-f1ab-0a9da0c1fc6d"
# !ls -l
# + [markdown] id="16TsWVn4gaiT"
# 자세하게(맨 앞과 맨 뒤 보기)
# **d**rwxr-xr-x 1 root root 4096 Jun 15 13:37 **sample_data**(directory)
# **-**rwxr-xr-x 1 root root 1697 Jan 1 2000 **anscombe.json**(file)
# + colab={"base_uri": "https://localhost:8080/"} id="gpQ6IkBdfgl-" outputId="b934b590-aca7-4d80-a849-a963c2a5541e"
# !pwd # 현재 자기 위치
# + colab={"base_uri": "https://localhost:8080/"} id="XFluru-rfxYp" outputId="4983700d-f1e5-41c3-cb56-4638ffda8e0d"
# !ls -l ./sample_data # 내가 원하는 디렉토리명 or 파일명
# + colab={"base_uri": "https://localhost:8080/"} id="aEdUFX4iguS0" outputId="f67cf3c7-41c3-4816-9042-e199231e706e"
# !ls -l ./
# + colab={"base_uri": "https://localhost:8080/"} id="YLzjUUfVjHTz" outputId="fec0c244-35d7-4c00-e924-15ecda45d178"
# !ls -l ./Wholesale_customers_data.csv
# + id="BJ60umwVin5K"
import pandas as pd
df = pd.read_csv('./Wholesale_customers_data.csv')
# + colab={"base_uri": "https://localhost:8080/"} id="1C66FKgsmMn4" outputId="e309fbff-e015-44fe-ec0f-74a4d1212cea"
df.info()
# + id="kF2ljLJqr5Sr"
X = df.iloc[:,:]
# + colab={"base_uri": "https://localhost:8080/"} id="_2bZ5GeCsSeh" outputId="6533be6e-ebec-4888-e2c1-6495c468a768"
X.shape
# + id="JbPlcc8FsVNz"
# split 할 필요없음 -> 측정할 방법 없음(y가 없어서)
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(X)
X = scaler.transform(X)
# + id="n2ABo3iCtUGJ"
from sklearn import cluster
kmeans = cluster.KMeans(n_clusters = 5)
# + colab={"base_uri": "https://localhost:8080/"} id="a2l2u4vBun4Q" outputId="01cf10e5-f7c1-4230-a457-c0ff178a657d"
kmeans.fit(X)
# + colab={"base_uri": "https://localhost:8080/"} id="1mIxUCKvuuGL" outputId="858147a8-31df-4b12-de43-13c2b8f6eb75"
kmeans.labels_
# + id="cOIltUgFvGFQ"
df['label'] = kmeans.labels_
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="m3xPXuifve3O" outputId="f52d384b-f762-4bba-dd49-19d2155c8941"
df.head()
# + [markdown] id="JCtEWxfzwphT"
# 시각화의 중심은 x, y 무조건 x, y 두개의 컬럼으로 만들기
# 이차원으로 무조건 만들어야 한다.
# + colab={"base_uri": "https://localhost:8080/", "height": 602} id="nAzmPf_ZvffF" outputId="2adde85c-d3cd-4c5e-9684-1caab4a62485"
df.plot(kind='scatter', x='Grocery', y='Frozen', c='label', cmap='Set1', figsize=(10,10)) # 종류, x, y
# + colab={"base_uri": "https://localhost:8080/"} id="pqw4Q6BJxNgT" outputId="41ee32dd-74ee-46a2-ea63-69746a38d62c"
# for ...:
# if ~((df['label'] == 0) | (df['label'] == 1)):
dfx = df[~((df['label'] == 0) | (df['label'] == 1))]
df.shape, dfx.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 602} id="mLNeDKC82LwS" outputId="c7c220cd-ae8e-46ac-e18f-327c45c8af6d"
dfx.plot(kind='scatter', x='Grocery', y='Frozen', c='label', cmap='Set1', figsize=(10,10)) # 종류, x, y
# + id="VmASp-Bi26H0"
df.to_excel('./wholesale.xls')
# + id="SvIPbbWP3JBp"
| wholesalecustomers_kmeans.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# prerequisite package imports
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sb
# %matplotlib inline
from solutions_biv import scatterplot_solution_1, scatterplot_solution_2
# -
# In this workspace, you'll make use of this data set describing various car attributes, such as fuel efficiency. The cars in this dataset represent about 3900 sedans tested by the EPA from 2013 to 2018. This dataset is a trimmed-down version of the data found [here](https://catalog.data.gov/dataset/fuel-economy-data).
fuel_econ = pd.read_csv('./data/fuel_econ.csv')
fuel_econ.head()
# **Task 1**: Let's look at the relationship between fuel mileage ratings for city vs. highway driving, as stored in the 'city' and 'highway' variables (in miles per gallon, or mpg). Use a _scatter plot_ to depict the data. What is the general relationship between these variables? Are there any points that appear unusual against these trends?
plt.scatter(data=fuel_econ, x="city", y="highway", alpha=0.05)
plt.xlabel("City fuel efficency")
plt.ylabel("Highway fuel efficency")
plt.xlim((10, 60))
plt.ylim((10, 60))
# run this cell to check your work against ours
scatterplot_solution_1()
# **Task 2**: Let's look at the relationship between two other numeric variables. How does the engine size relate to a car's CO2 footprint? The 'displ' variable has the former (in liters), while the 'co2' variable has the latter (in grams per mile). Use a heat map to depict the data. How strong is this trend?
x_vals = fuel_econ["displ"]
x_bins = np.arange(0, np.ceil(x_vals.max()), np.ceil(x_vals.std())/5)
y_vals = fuel_econ["co2"]
y_bins = np.arange(np.floor(y_vals.min()), np.ceil(y_vals.max()), np.ceil(y_vals.std())/2)
plt.hist2d(x=x_vals, y=y_vals, cmin=1, bins=[x_bins, y_bins], cmap="viridis");
plt.colorbar()
y_vals.describe()
# run this cell to check your work against ours
scatterplot_solution_2()
| exercises/Data & Plotting/Scatterplot_Practice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from collections import defaultdict
import json
import os
import matplotlib.pyplot as plt
from matplotlib.ticker import FuncFormatter
import numpy as np
import pandas as pd
from tti_explorer import config, sensitivity, utils
from tti_explorer.strategies import RETURN_KEYS
# +
perc_formatter = FuncFormatter(lambda x, _: f"{100*x:.0f}%")
formatters = {
'app_cov': perc_formatter,
'trace_adherence': perc_formatter
}
rc_dct = {
'figure.figsize': (7, 6),
'figure.max_open_warning': 1000,
"errorbar.capsize": 2.5,
'font.size': 16
}
plt.rcParams.update(rc_dct)
plt.style.use("seaborn-ticks")
# +
chart_folder = os.path.join(os.environ['REPOS'], 'tti-explorer', 'outputs', 'charts', 'new-style')
input_folder = os.path.join(os.environ['DATA'], "tti-explorer")
measures_order = [
'no_TTI',
'symptom_based_TTI',
'test_based_TTI',
'test_based_TTI_test_contacts',
]
# +
def config_files(folder):
return filter(lambda x: x.startswith("config") and x.endswith('.json'), os.listdir(folder))
def run_fname(cfg_fname):
return cfg_fname.replace("config", "run").replace("json", 'csv')
def resgen(data_dir):
lockdowns = next(os.walk(data_dir))[1]
for lockdown in lockdowns:
folder = os.path.join(data_dir, lockdown)
for cfg_file in config_files(folder):
cfg = utils.read_json(os.path.join(folder, cfg_file))
target = cfg[sensitivity.TARGET_KEY]
results = pd.read_csv(os.path.join(folder, run_fname(cfg_file)), index_col=0)
yield lockdown, target, cfg[sensitivity.CONFIG_KEY], results
def errorbar(ax, xaxis, means, stds, label, **kwds):
conf_intervals = 1.96 * stds
ax.errorbar(xaxis, means, yerr=conf_intervals, label=label, **kwds)
ax.set_xticks(xaxis)
ax.set_xticklabels(xaxis)
def legend(ax, **kwds):
defaults = dict(
loc="best",
frameon=True,
framealpha=0.5,
fancybox=False,
fontsize=12
)
defaults.update(kwds)
return ax.legend(**defaults)
def plot_lockdown(plotter, lockdown_dct, deck, key_to_plot, order, formatters={}, title=False):
for param_name, sim_results in lockdown_dct.items():
tick_formatter = formatters.get(param_name)
fig, ax = plt.subplots(1)
for measure in order:
xaxis, means, std_errs = arrange_sim_results(sim_results[measure], key_to_plot)
plotter(ax, xaxis, means, std_errs, label=nice_lockdown_name(measure))
legend(ax)
if title:
ax.set_title(param_name)
ax.set_ylabel(key_to_plot, fontsize=14)
ax.set_xlabel(nice_param_name(param_name))
if tick_formatter is not None:
ax.xaxis.set_major_formatter(tick_formatter)
deck.add_figure(fig, name=param_name+"_"+key_to_plot.lower().replace(" ", "_").replace("%", ""))
return fig
def make_tables(entry, key):
coords, reslist = zip(*entry)
means, stds = zip(*[(k[key].loc['mean'], k[key].loc['std']) for k in reslist])
means_mat = pd.DataFrame(np.array(utils.sort_by(means, coords, return_idx=Faelse)).reshape(3, 3))
stds_mat = pd.DataFrame(np.array(utils.sort_by(stds, coords, return_idx=False)).reshape(3, 3))
return means_mat, stds_mat
def format_table(means, stds):
t1, t2 = ['TTI Delay (days)', 'NPI severity']
table = means.applymap(
lambda x: f"{x:.2f}"
).add(
" \pm "
).add(
stds.mul(
1.96
).applymap(
lambda x: f"{x:.2f}"
)
).applymap(lambda x: f"${x}$")
new_order = [[2,2], [2,1], [1,2], [2,0], [1,1], [0,2], [1,0], [0,1], [0,0]]
new_order = list(tuple(x) for x in new_order)
table.index = ['No TTI'] + new_order
table.columns = sorted(test_trace_results.keys(), reverse = True)
table.index.name = t1
table.columns.name = t2
return table
def format_mean_only(means):
t1, t2 = ['TTI Delay (days)', 'NPI severity']
table = means.applymap(
lambda x: f"{x:.0f}\%"
)
new_order = [[2,2], [2,1], [1,2], [2,0], [1,1], [0,2], [1,0], [0,1], [0,0]]
new_order = list(tuple(x) for x in new_order)
table.index = ['No TTI'] + new_order
table.columns = sorted(test_trace_results.keys(), reverse = True)
table.index.name = t1
table.columns.name = t2
return table
def make_new_tables(big_dict, name='test_based_TTI'):
new_order = [[2,2], [2,1], [1,2], [2,0], [1,1], [0,2], [1,0], [0,1], [0,0]]
mean_mat = np.zeros((10, 5))
stds_mat = np.zeros((10, 5))
no_tti, test_based_tti = ('no_TTI', name)
s_levels = sorted(test_trace_results.keys(), reverse=True)
for s_idx, s_level in enumerate(s_levels):
mean_mat[0, s_idx] = big_dict[s_level][no_tti]['means'].iloc[0].loc[0]
stds_mat[0, s_idx] = big_dict[s_level][no_tti]['stds'].iloc[0].loc[0]
for new_row_idx, (row_idx, col_idx) in enumerate(new_order):
mean_mat[new_row_idx+1, s_idx] = big_dict[s_level][test_based_tti]['means'].iloc[row_idx].loc[col_idx]
stds_mat[new_row_idx+1, s_idx] = big_dict[s_level][test_based_tti]['stds'].iloc[row_idx].loc[col_idx]
mean_mat = pd.DataFrame(mean_mat, columns = s_levels)
stds_mat = pd.DataFrame(stds_mat, columns = s_levels)
return mean_mat, stds_mat
def nice_lockdown_name(name):
# this is so nasty!
mapp = {
'test_based_TTI_test_contacts': "Test-based TTI, test contacts",
'no_TTI': 'No TTI',
'symptom_based_TTI': 'Symptom-based TTI',
'test_based_TTI': 'Test-based TTI',
'test_based_TTI_full_compliance': 'Test-based TTI'
}
return mapp[name]
# -
# # Pinch Points
pinch_points_dir = os.path.join(input_folder, "pinch-points")
pinch_points_results = defaultdict(lambda: defaultdict(lambda: defaultdict(list)))
for lockdown, target, cfg, results in resgen(pinch_points_dir):
if int(lockdown[1]) > 0:
level, measures = lockdown.split('_', maxsplit=1)
results = results.set_index(config.STATISTIC_COLNAME, drop=True)
pinch_points_results[level][target][measures].append((cfg[target], results))
tt_dir = os.path.join(input_folder, "pinch-points-test-trace-delay")
test_trace_results = defaultdict(lambda: defaultdict(list))
for lockdown, target, cfg, results in resgen(tt_dir):
if int(lockdown[1]) > 0:
vals = [cfg[k] for k in target]
level, measures = lockdown.split('_', maxsplit=1)
results = results.set_index(config.STATISTIC_COLNAME, drop=True)
test_trace_results[level][measures].append((vals, results))
# # Compliance Tables
perc_keys = [
RETURN_KEYS.stopped_by_social_distancing_percentage,
RETURN_KEYS.stopped_by_symptom_isolating_percentage,
RETURN_KEYS.stopped_by_tracing_percentage,
RETURN_KEYS.not_stopped_by_tti,
]
# +
def take_key(lst_of_df, key, stat_name):
x, dfs = zip(*lst_of_df)
means = pd.Series([df.loc[stat_name][key] for df in dfs], index=x)
return means.sort_index()
def arrange_sim_results(sim_results, key):
means = take_key(sim_results, key, 'mean')
stds = take_key(sim_results, key, 'std')
return means.index.values, means, stds
def to_perc(f):
return f"{100 * f:.0f}\%"
def format_table(tab, fmt_fnc):
tab.index.name = "Compliance"
tab.columns.name = "NPI Severity"
return tab.rename(index=fmt_fnc).applymap(fmt_fnc).sort_index(axis=1, ascending=False)
def param_tables(pp_res_dict, param_name, measure, keys, stat_name):
param_res = {k: v[param_name] for k, v in pp_res_dict.items()}
table_data = utils.swaplevel({
key:
utils.swaplevel(utils.map_lowest(lambda x: take_key(x, key, stat_name), param_res)) for key in perc_keys
})[measure]
return {k: format_table(pd.concat(v, axis=1) / 100, to_perc) for k, v in table_data.items()}
# +
measure = 'test_based_TTI'
for param_name in ['trace_adherence', 'app_cov']:
table_deck = utils.LatexTableDeck()
tables = param_tables(pinch_points_results, param_name, measure, keys=perc_keys, stat_name='mean')
for key, tab in tables.items():
table_deck.add_table(
tex_table=tab.to_latex(escape=False),
caption=key.replace("%", "\%")
)
table_deck.make(os.path.join(chart_folder, f"{param_name}_{measure.lower()}_perc_tables.tex"))
# -
# # Test Trace Tables
# +
def get_big_dict(tt_results, measure, keys):
big_dict = defaultdict(dict)
for k in sorted(list(tt_results.keys())):
for policy, entry in tt_results[k].items():
if policy in [measure, "no_TTI"]:
means, stds = make_tables(entry, key=keys)
big_dict[k][policy] = {"means": means, "stds": stds}
return big_dict
def increment_tup(tup):
return tuple(k+1 for k in tup) if isinstance(tup, tuple) else tup
# -
name = "test_based_TTI"
table_deck = utils.LatexTableDeck()
for key in perc_keys:
big_dict = get_big_dict(test_trace_results, name, key)
mean_mat, stds_mat = make_new_tables(big_dict, name=name)
table_str = format_mean_only(mean_mat).rename(index=increment_tup).to_latex(escape=False)
table_deck.add_table(
tex_table=table_str,
caption=nice_lockdown_name(name) + " " + key.replace("%", "\%")
)
table_deck.make(os.path.join(chart_folder, "test-trace-perc-prevented.tex"))
# ### New style chart
# +
levels = [f"S{i}" for i in range(1, 6)]
kwds_list = [
{'marker': "o", "markersize": 4, "color": "C0"},# "markerfacecolor": "none"},
{'marker': "+", "markersize": 4, "color": "C1"}, #"markerfacecolor": "none"},
{'marker': "*", "markersize": 4, "color": "C2"}, #"markerfacecolor": "none"},
{'marker': "x", "markersize": 4, "color": "C3"}, #"markerfacecolor": "none"},
{'marker': "s", "markersize": 4, "color": "C4"}, #"markerfacecolor": "none"},
]
# -
key = RETURN_KEYS.reduced_r
to_plot = np.array([8, 7, 4, 3, 0])
original_order = np.array([(1, 1), (1, 2), (1, 3), (2, 1), (2, 2), (2, 3), (3, 1), (3, 2), (3, 3)])
all_yticklabs = np.array(["/".join(l.astype(str)) for l in original_order])
xaxis = [1, 2, 3, 4, 5]
test_based_tti = utils.swaplevel(test_trace_results)['test_based_TTI']
no_tti = utils.swaplevel(test_trace_results)['no_TTI']
# +
def get_stat(dct, stat, key):
return {k: take_key(v, key, stat) for k,v in dct.items()}
tt_means = get_stat(test_based_tti, "mean", key=RETURN_KEYS.reduced_r)
tt_stds = get_stat(test_based_tti, "std", key=RETURN_KEYS.reduced_r)
no_tt_means = utils.map_lowest(lambda x: x[0], get_stat(no_tti, "mean", key=RETURN_KEYS.reduced_r))
no_tt_stds = utils.map_lowest(lambda x: x[0], get_stat(no_tti, "std", key=RETURN_KEYS.reduced_r))
# -
pd.concat(tt_means, axis=1)
# +
fig, (ax1, ax2) = plt.subplots(1, 2, sharey=True, figsize=(13, 6))
for l, kwds in zip(levels, kwds_list):
ax1.errorbar(xaxis, tt_means[l][to_plot], 1.96 * tt_stds[l][to_plot], label=l, **kwds)
ax1.errorbar([0], [no_tt_means[l]], [1.96 * no_tt_stds[l]], **kwds )
xaxis_2, means, stds = arrange_sim_results(
pinch_points_results[l]['trace_adherence']['test_based_TTI'],
key
)
ax2.errorbar(xaxis_2, means, 1.96 * stds, label=l, **kwds)
ax1.grid(alpha=1, axis="y")
ax2.grid(alpha=1, axis="y")
ax1.set_xticks(range(-1, len(to_plot) + 1 + 1))
ax1.set_xticklabels(["", "No TTI", *all_yticklabs[to_plot], ""])
ax1.set_ylabel(key)
ax1.set_xlabel("Test/Trace Delay, 80% Compliance")
ax2.xaxis.set_major_formatter(perc_formatter)
ax2.set_xlabel("Compliance, 2/1 Test/Trace Delay")
ax2.yaxis.set_tick_params(which='both',length=0)
plt.subplots_adjust(wspace=0.1)
fig.legend(
*ax1.get_legend_handles_labels(),
framealpha=0,
ncol=5,
bbox_to_anchor=(0.45, -0.03),
loc="lower center"
)
for fmt in ['pdf', 'png']:
fig.savefig(os.path.join(chart_folder, f"tt_compliance.{fmt}"), format=fmt, bbox_inches="tight", dpi=1500)
# -
assert False
# # Sensitivity
inf_props_mapping = {
'oxteam_infection_proportions0_seed0': "vary_flu0",
'oxteam_infection_proportions1_seed0': "vary_flu1",
'oxteam_infection_proportions2_seed0': "vary_flu2",
'oxteam_infection_proportions3_seed0': "vary_flu3",
'oxteam_infection_proportions4_seed0': "vary_covid0",
'oxteam_infection_proportions5_seed0': "vary_covid1",
'oxteam_infection_proportions6_seed0': "vary_covid2"
}
# +
def make_tables(entry, key=RETURN_KEYS.reduced_r):
coords, reslist = zip(*entry)
means, stds = zip(*[(k[key]['mean'], k[key]['std']) for k in reslist])
means_mat = pd.DataFrame(np.array(utils.sort_by(means, coords, return_idx=False)).reshape(3, 3))
stds_mat = pd.DataFrame(np.array(utils.sort_by(stds, coords, return_idx=False)).reshape(3, 3))
return means_mat, stds_mat
def format_table(means, stds):
t1, t2 = ['Test Delay (days)', 'Manual Trace Delay (days)']
table = means.applymap(
lambda x: f"{x:.2f}"
).add(
" \pm "
).add(
stds.mul(
1.96
).applymap(
lambda x: f"{x:.2f}"
)
).applymap(lambda x: f"${x}$")
table.index = range(1, 4)
table.columns = range(1, 4)
table.index.name = t1
table.columns.name = t2
return table
# -
case_sensitivity_results = defaultdict(dict)
case_sensitivity_dir = os.path.join(input_folder, 'case-sensitivity-quick')
for lockdown, target, cfg, results in resgen(case_sensitivity_dir):
if lockdown[1] == '0':
continue
level, measures = lockdown.split("_", maxsplit=1)
# just load all the run_dfs and concat them...
folder = os.path.join(case_sensitivity_dir, lockdown)
runs = [
pd.read_csv(
os.path.join(folder, fname),
index_col=0
)
for fname in os.listdir(folder) if fname.startswith("run") and fname.endswith(".csv")
]
case_sensitivity_results[level][measures] = pd.concat(
runs,
axis=0
).drop('oxteam_infection_proportions7_seed0').rename(index=inf_props_mapping)
# +
import config
k_formatter = FuncFormatter(lambda x, _: f"{x:.0f}k")
class LockdownSensitivityPlotter:
# BE: need to sort out this naming bullshit
def __init__(self):
self.ticklabel_maps = {
'inf_profile': np.array([f"{int(x)}" for x in (np.array([2.11, 2.8, 3.49]) - 1)/0.69]),
'vary_flu': np.array([f"{k}k" for k in [50, 100, 200, 300]]),
'vary_covid': np.array([f"{k}k" for k in [10, 20, 30]]),
'p_day_noticed_symptoms': (np.array([2, 3, 4]) - 1).astype(str),
}
self.xlabels = {
'inf_profile': "Day on which primary case is most infectious",
'vary_flu': "# COVID negative cases with COVID like symptoms (daily)",
'vary_covid': "# COVID positive cases (daily)",
'p_day_noticed_symptoms': (
"Expected number of days"
)
}
self.param_map = {
'oxteam_inf_profile11_seed0': 0,
'oxteam_inf_profile12_seed0': 1,
'oxteam_inf_profile13_seed0': 2,
'oxteam_p_day_noticed_symptoms10_seed0': 0,
'oxteam_p_day_noticed_symptoms8_seed0': 2,
'oxteam_p_day_noticed_symptoms9_seed0': 1,
'vary_flu0': 0,
'vary_flu1': 1,
'vary_flu2': 2,
'vary_flu3': 3,
'vary_covid0': 0,
'vary_covid1': 1,
'vary_covid2': 2
}
self.y_formatters = {'vary_covid': k_formatter, 'vary_flu': k_formatter}
def _grouper(self, s): ## not neat!
for i in range(10):
s = s.replace(f"{i}", "")
return s.replace("oxteam_", "").replace("_seed", "")
def _plot_parameter(self, ax, results_df, key):
for measure, dfstats in results_df[[key, config.STATISTIC_COLNAME]].groupby(level=0):
dfstats = dfstats.droplevel(0).rename(self.param_map).sort_index()
mean = dfstats.query(
f"{config.STATISTIC_COLNAME}=='mean'"
).drop(config.STATISTIC_COLNAME, axis=1).squeeze()
std = dfstats.query(
f"{config.STATISTIC_COLNAME}=='std'"
).drop(config.STATISTIC_COLNAME, axis=1).squeeze()
xaxis = mean.index.values
errorbar(ax, xaxis, mean.values, std.values, label=nice_lockdown_name(measure))
return xaxis
def __call__(self, results, deck, keys_to_plot):
by_param = pd.concat(results, axis=0).groupby(level=1, by=self._grouper)
for param_name, v in by_param:
fig, ax = plt.subplots(1)
xaxis = self._plot_parameter(ax, v, keys_to_plot[param_name])
ax.set_xticks(xaxis)
ax.set_xticklabels(self.ticklabel_maps[param_name][xaxis])
ax.set_xlabel(self.xlabels[param_name])
ax.set_ylabel(keys_to_plot[param_name])
formatter = self.y_formatters.get(param_name, None)
if formatter is not None:
ax.yaxis.set_major_formatter(formatter)
legend(ax)
deck.add_figure(fig, name=param_name)
# +
# self = plotter
# key = RETURN_KEYS.reduced_r
# by_param = pd.concat(results, axis=0).groupby(level=1, by=self._grouper)
# for param_name, results_df in by_param:
# for measure, dfstats in results_df[[key, config.STATISTIC_COLNAME]].groupby(level=0):
# break
# dfstats = dfstats.droplevel(0).rename(self.param_map)
# mean = dfstats.query(
# f"{config.STATISTIC_COLNAME}=='mean'"
# ).drop(config.STATISTIC_COLNAME, axis=1).squeeze()
# std = dfstats.query(
# f"{config.STATISTIC_COLNAME}=='std'"
# ).drop(config.STATISTIC_COLNAME, axis=1).squeeze()
# xaxis = mean.index.values
# errorbar(ax, xaxis, mean.values, std.values, label=nice_lockdown_name(measure))
# +
keys_to_plot = {
'inf_profile': RETURN_KEYS.reduced_r,
'p_day_noticed_symptoms': RETURN_KEYS.reduced_r,
'vary_covid': RETURN_KEYS.tests,
'vary_flu': RETURN_KEYS.tests
}
with plt.rc_context(rc_dct):
plotter = LockdownSensitivityPlotter()
for level, results in case_sensitivity_results.items():
deck = utils.PdfDeck()
plotter(results, deck, keys_to_plot)
deck.make(os.path.join(chart_folder, f"{level}_sensitivity.pdf"))
for fig in deck.figs:
fig.tight_layout()
individual_dir = os.path.join(os.path.join(chart_folder, f"{level}_individual"))
os.makedirs(individual_dir, exist_ok=True)
deck.make_individual(folder=individual_dir)
| tti/notebooks/plots/plots-bryn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.ticker as mtick
# %matplotlib inline
np.random.seed(1)
KNN = np.loadtxt('result_KNN_median.dat')
NB = np.loadtxt('result_NB_median.dat')
DT = np.loadtxt('result_DT_median.dat')
SVM = np.loadtxt('result_SVM_median.dat')
LR = np.loadtxt('result_LR_median.dat')
RF = np.loadtxt('result_RF_median.dat')
XGB = np.loadtxt('result_XGB.dat')
MLP = np.loadtxt('result_MLP_median_morehyperparameters.dat')
ER = np.loadtxt('result_ER_LAD_median.dat')
# +
#number of methods
m = 9
# number of data sets
n = LR.shape[1]
print(m,n)
## accuracy
acc = np.zeros((m,n))
# -
def number_winnings(acc,alpha=0.):
## find number of winning times for each method
m,n = acc.shape
n_wins = np.zeros(m)
## for each data
#j = 0
for j in range(n):
#print('j:',acc[:,j])
acc_max = max(acc[:,j])
for i in range(m):
if acc[i,j] >= (1-alpha)*acc_max:
n_wins[i] += 1
return n_wins
def select_metric(i):
acc[0,:],acc[1,:],acc[2,:],acc[3,:],acc[4,:],acc[5,:],acc[6,:],acc[7,:],acc[8,:] = \
KNN[i,:],NB[i,:],DT[i,:],SVM[i,:],MLP[i,:],LR[i,:,],XGB[i,:],RF[i,:],ER[i,:]
return acc
# ### Accuracy
# +
# accuracy:
acc = select_metric(i=0)
acc_av = acc.mean(axis=1)
acc_std = acc.std(axis=1)
print('mean:',acc_av)
print('std:',acc_std)
n_wins = number_winnings(acc,alpha=0.005)
#n_wins = n_wins/n
print(n_wins)
# -
acc.shape
# +
nx,ny = 2,1
nfig = nx*ny
fig, ax = plt.subplots(ny,nx,figsize=(nx*4.2,ny*2.8))
xvalue = np.arange(m+1)
labels = ['KNN','NB','DT','SVM','MLP','LR','XGB','RF','ER']
colors = ['pink','orange','red','green','olive','brown','gray','purple','blue']
#patterns = ["|", "/", "\\", "-", ".", "*" ]
for i in range(m):
#ax[0].barh(i,acc_av[i],color='white',edgecolor='black',\
# width=0.8,hatch=patterns[i],label=labels[i])
ax[0].bar(i,acc_av[i],color=colors[i],edgecolor='black',width=0.7,label=labels[i],alpha=1.,zorder=0)
ax[1].bar(i,n_wins[i],color=colors[i],edgecolor='black',width=0.7,label=labels[i],alpha=1.,zorder=0)
for j in range(2):
ax[j].set_xticks(xvalue)
ax[j].set_xticklabels(labels)
ax[j].yaxis.grid(linestyle='--',linewidth='0.5',zorder=-1)
ax[j].yaxis.set_zorder(level=-1)
ax[0].set_ylabel('accuracy mean')
ax[0].set_ylim([0.82,0.92])
ax[1].set_ylabel('number of winnings')
ax[1].set_ylim([0,20])
plt.tight_layout(h_pad=1, w_pad=2.)
#plt.show()
#plt.savefig('fig1.pdf', format='pdf', dpi=100)
# -
| 2020.07.2400_classification/.ipynb_checkpoints/performance_comparison_v3-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + id="TEYu6zYJx-k2"
import gensim
import scipy
import operator
import pandas as pd
import numpy as np
from scipy.cluster import hierarchy
from sklearn.cluster import KMeans
from gensim.models import Word2Vec
import pickle
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/"} id="kgmAVRFqsRqD" outputId="360528f9-7da7-4519-b310-8a1cc61f33c7"
from google.colab import drive
drive.mount ('/content/drive', force_remount=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 383} id="FWAb15zHgOme" outputId="d018c912-e8d2-46cb-b458-4315f875788a"
df = pd.read_csv('/content/drive/MyDrive/Colab Notebooks/Deteksi Depresi Bismillah/Preprocessing Data/Preprocessing_v2.csv')
df = df.apply(lambda x: x.str.strip() if isinstance(x, str) else x).replace('', np.nan)
df = df.dropna()
df.iloc[:3]
# + colab={"base_uri": "https://localhost:8080/"} id="D7Vxs8BNZ2Sf" outputId="c5125b88-4b0f-44a0-85cb-c71a8834678e"
print(len(df))
# + id="GCUz3BLevQ7g"
import gensim
# Training the wor2vec model using train dataset
w2v_model=gensim.models.Word2Vec(df['string'],size=100, workers=4)
w2v_model.save('/content/drive/MyDrive/Colab Notebooks/Deteksi Depresi Bismillah/Model Word2vec.model')
# + id="0RLeCLnLKUTF"
w2v_model = gensim.models.Word2Vec.load("/content/drive/MyDrive/Colab Notebooks/Deteksi Depresi Bismillah/Model Word2vec.model")
# + colab={"base_uri": "https://localhost:8080/"} id="-MaF-zhis70E" outputId="ea8159a7-6267-4885-a5c1-0ff3adad3aa8"
sent_vectors =[]
for sent in df['string']:
sent_vec = np.zeros(100) #karena panjang vektor kata adalah 0
cnt_words = 0
for word in sent:
try :
vec = w2v_model.wv[word]
sent_vec += vec
cnt_words += 1
except:
pass
sent_vec /= cnt_words
sent_vectors.append(sent_vec)
sent_vectors = np.array(sent_vectors)
sent_vectors = np.nan_to_num(sent_vectors)
sent_vectors.shape
# + colab={"base_uri": "https://localhost:8080/"} id="9kGQ_ovtHSMi" outputId="36d3a2be-1521-479c-866c-427d30382b21"
print(sent_vectors)
# + id="OHgE61NfCaVg"
from sklearn.cluster import DBSCAN
# + colab={"base_uri": "https://localhost:8080/"} id="cMonK2O99cMw" outputId="349002b8-3073-479c-8a29-ed412b50c1c4"
coba = {"Tweet":df['string'],"Username":df['usernameTweet']}
coba = pd.DataFrame(coba)
for eps in np.arange(0.1, 0.4, 0.1):
epsilon = []
epsilon.append(eps)
for min_sam in np.arange(1,80,1):
dbscan_model = DBSCAN(eps=eps, min_samples=min_sam, metric_params=None, algorithm="auto", leaf_size=30, p=None, n_jobs=1)
labels = dbscan_model.fit_predict(sent_vectors)
min_samp = []
noise = []
cluster = []
min_samp.append(min_sam)
clusters = {}
for i, w in enumerate(df['string']):
clusters[w] = labels[i]
dbscan_clusters = sorted(clusters.items(), key=operator.itemgetter(1))
n_clusters = len(set(labels)) - (1 if -1 in labels else 0)
n_noise = len([lab for lab in labels if lab == -1])
cluster.append(n_clusters)
noise.append(n_noise)
print("EPS: ", eps, "\tmin_sample: ", min_sam, "\tClusters: ", n_clusters, "\tNoise: ", n_noise)
DBSCANsave = (f'/content/drive/MyDrive/Colab Notebooks/Deteksi Depresi Bismillah/DBSCAN Model/DBSCAN Model eps {eps} min_sample {min_sam}.sav')
pickle.dump(dbscan_model, open(DBSCANsave, 'wb'))
coba[f'Clustering DBSCAN w2v {eps}{min_sam}']= dbscan_model.labels_
# + id="y1rbN95eKXhY"
dbscan_model = DBSCAN(eps=0.1, min_samples=31, metric_params=None, algorithm="auto", leaf_size=30, p=None, n_jobs=1)
labels = dbscan_model.fit_predict(sent_vectors)
# + colab={"base_uri": "https://localhost:8080/"} id="OVaY7wHHKmKg" outputId="7d3274af-abdc-4165-bc84-ea5f11ad164e"
print(len(dbscan_model.labels_))
# + colab={"base_uri": "https://localhost:8080/", "height": 429} id="Wl6Ic2IlBL6o" outputId="7b8b399b-6f7e-4cb3-a2b7-c3e654134814"
coba.iloc[:3]
# + colab={"base_uri": "https://localhost:8080/"} id="iN5CqWtt85M5" outputId="07fa86ca-f9e4-4139-8d13-e39000a507d7"
coba.groupby(['Clustering DBSCAN w2v 0.137'])['Tweet'].count()
# + id="uhf8ATJeXMe-"
coba.to_csv('/content/drive/MyDrive/Colab Notebooks/Deteksi Depresi Bismillah/DBSCAN Model/DBSCAN Clustering W2v Result.csv ')
# + colab={"base_uri": "https://localhost:8080/", "height": 429} id="0fpM1eaqIKDV" outputId="f8ed80b7-2ef6-47eb-e091-750a1effc429"
coba = pd.read_csv('/content/drive/MyDrive/Colab Notebooks/Deteksi Depresi Bismillah/DBSCAN Model/DBSCAN Clustering W2v Result.csv ')
coba.loc[:2]
# + id="-Tho_8S7Y7Sp"
data = pd.read_csv('/content/drive/MyDrive/Colab Notebooks/Deteksi Depresi Bismillah/Data Clustering.csv')
# + id="dolaL1P_C5sM"
n_clusters=4
kmeans = KMeans(n_clusters).fit(sent_vectors)
clusters = kmeans.labels_.tolist()
# + colab={"base_uri": "https://localhost:8080/"} id="BuBbLtYXsfxr" outputId="a6b24cd4-eaaa-411e-db4d-b0f8601d7d95"
data['Clustering Kmeans W2V 4'] = clusters
data.groupby(['Clustering Kmeans W2V 3'])['Tweet'].count()
# + id="LsIMFTFNC77d"
data.to_csv('/content/drive/MyDrive/Colab Notebooks/Deteksi Depresi Bismillah/Data Clustering.csv', index=False)
# + colab={"base_uri": "https://localhost:8080/", "height": 247} id="mLg4aST1upeT" outputId="46f138d5-0962-4481-a16c-feb9fff89ed9"
dt = pd.read_csv('/content/drive/MyDrive/Colab Notebooks/Deteksi Depresi Bismillah/Data Clustering.csv')
dt.iloc[:2]
# + colab={"base_uri": "https://localhost:8080/", "height": 285} id="lA5zv0X2VrfC" outputId="eed1915a-a48f-4dd7-d7d8-ebfc44e41c83"
import scipy
from scipy.cluster import hierarchy
dendro=hierarchy.dendrogram(hierarchy.linkage(sent_vectors,method='ward'))
plt.axhline(y=13)# cut at 30 to get 5 clusters
# + id="QQcNkKrzWFAd"
from sklearn.cluster import AgglomerativeClustering
cluster = AgglomerativeClustering(n_clusters=3, affinity='euclidean', linkage='ward') #took n=5 from dendrogram curve
Agg=cluster.fit_predict(sent_vectors)
# + colab={"base_uri": "https://localhost:8080/"} id="PqVRwQZQWtna" outputId="8fcba1bd-a947-49de-ce9e-e3ffe2d56385"
dt['Clustering Agglomerative W2V 3'] = cluster.labels_
dt.groupby(['Clustering Agglomerative W2V 3'])['Tweet'].count()
# + colab={"base_uri": "https://localhost:8080/", "height": 335} id="ELcX5cWRXBFl" outputId="81919795-0244-43f5-8e01-9f9325917931"
dt.iloc[:2]
# + id="0rJp1xQ4oIY4"
dt.to_csv('/content/drive/MyDrive/Colab Notebooks/Deteksi Depresi Bismillah/Data Clustering.csv', index=False)
# + id="DelxRo17uI33"
| analytics/Clustering_W2v.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Name : <NAME>
# ## Mat No: 21120612455
# ## E-mail: <EMAIL>
# ## PROJECT 1
# 
# <br>BEGIN
# <br>INPUT num
# <br>LET diff = num - 17
# <br>IF num > 17 THEN
# * PRINT 2 * diff
#
# <br>ELSE
# * LET abs_val = -1 * diff
# * PRINT abs_val
# <br>END IF
# <br>END
# program to get difference of a number and 17 and perform operations on the difference
num = float(input("What number do you want please? "))
diff = num - 17
if num > 17:
print (2* diff)
else:
abs_val = abs(diff)
print(abs_val)
# ## PROJECT 2
# 
# <br>BEGIN
# <br>INPUT num1
# <br>INPUT num2
# <br>INPUT num3
# <br>IF num1 EQUALS num2 and num1 EQUALS num3 THEN
# * LET add = 3 * num1
# * PRINT add
#
# <br>ELSE
# * LET add = num1 + num2 + num3
# * PRINT add
#
# <br>END IF
# <br>END
# +
# program to find the sum of 3 numbers
# get the 3 numbers
num1 = float(input("What is the 1st number? "))
num2 = float(input("What is the 2nd number? "))
num3 = float(input("What is the 3rd number? "))
if num1 == num2 and num1 == num3:
add = 3 * num1
print(add)
else:
add = num1 + num2 + num3
print(add)
# -
# ## PROJECT 3
# 
# <br>BEGIN
# <br>INPUT num1
# <br>INPUT num2
# <br>LET add = num1 + num2
# <br>IF num1 > num2 THEN
# * LET diff = num1 - num2
#
# <br>ELSE
# * LET diff = num2 - num1
#
# <br>END IF
# <br>IF num1 EQUALS num2 OR add EQUALS 5 OR diff EQUALS 5 THEN
# * PRINT TRUE
#
# <br>ELSE
# * PRINT FALSE
#
# <br>END IF
# <br>END
# +
# program to confirm values of numbers, their sum and their difference
num1 = float(input("What is the 1st number? "))
num2 = float(input("What is the 2nd number? "))
# get the sum and difference of the numbers
add = num1 + num2
if num1 > num2:
diff = num1 - num2
else:
diff = num2 - num1
if num1 == num2 or add == 5 or diff == 5:
print("True")
else:
print("False")
# -
# ## PROJECT 4
# 
# <br>BEGIN
# <br>INPUT num1
# <br>INPUT num2
# <br>INPUT num3
# <br>FIND the biggest number
# <br>FIND the middle number
# <br>FIND the smallest number
# <br>LET order = ARRANGE smallest number, middle number, biggest number
# <br>PRINT order
# <br>END
# +
# program to arrange numbers in ascending order
num1 = float(input("What is the 1st number? "))
num2 = float(input("What is the 2nd number? "))
num3 = float(input("What is the 3rd number? "))
maximum = max(num1, num2,num3)
minimum = min(num1,num2,num3)
middle = (num1 + num2 + num3) - maximum - minimum
order = (minimum, middle, maximum)
print(order)
# -
# ## PROJECT 5
# 
# <br>BEGIN
# <br>INPUT num
# <br>SET N = 0
# <br>LET num = num-1
# <br>SET numbers = list starting with num
# <br>WHILE num > 0
# * num = num-1
# * IF last element in numbers IS NOT EQUAL to 1:
# * ADD num^3 to the list
# * ELSE
# * PRINT numbers
# * END IF
# * END WHILE
# <br>FOR i in numbers
# * N = N + i
# <br>END FOR
# <br>PRINT N
# <br> END
#
#program that adds the cubes of numbers less than the input
num = int(input("Enter a number: "))
def cubed(num):
N =0
num = num - 1
#create a list starting with the cube of the 1st number less than the input
numbers = [num**3]
while num > 0:
num -=1
#when the last element is not 1, keep adding cubes of numbers less than the input to list
if numbers[-1] != 1:
numbers.append(num**3)
else:
print("The numbers being added:",numbers)
#add each list element to N
for i in numbers:
N += i
print ("The summation is", N)
cubed(num)
| week_3/class-projects.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sklearn.datasets import load_iris
type(load_iris)
iris = load_iris()
print(iris)
print(iris.data.shape)
print(type(iris.data))
print(iris.target)
print(iris.target_names)
x=iris.data
y=iris.target
# # 2. Write a python program to build a K-Nearest Neighbour classifier using Scikit learn and predict the class of a unknown sample
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier(n_neighbors=1)
knn.fit(x,y)
knn.predict([[5, 3, 1, 0],[6,3,5,2]])
knn5 = KNeighborsClassifier(n_neighbors=5)
x_new = [[5, 3, 1, 0],[6,3,5,2]]
knn5.fit(x,y)
knn5.predict(x_new)
# ### 3. Write a Python program to create separate ndarrays for features and targets from data using Scikit learn library. Use train_test_split function to split the dataset into training and testing
#
#from sklearn.cross_validation import train_test_split
from sklearn.model_selection import train_test_split
help(train_test_split)
x_train,x_test,y_train,y_test = train_test_split(x,y,test_size=0.4,random_state = 4)
knn = KNeighborsClassifier(n_neighbors=5)
knn.fit(x_train,y_train)
# ### 4. Write a python program to build a K-Nearest Neighbour classifier using Scikit learn and test it using the test dataset. Find the accuracy using accuracy_score() function
from sklearn import metrics
type(metrics)
y_pred = knn.predict(x_test)
metrics.accuracy_score(y_test,y_pred)
metrics.confusion_matrix(y_test,y_pred)
# ### 5. Write a python program to build a K-Nearest Neighbour classifier using Scikit learn and find the optimal value of k by plotting the accuracies for different values of k using matplotlib library
k_range = range(1,30)
score = []
for k in k_range:
knn = KNeighborsClassifier(n_neighbors=k)
knn.fit(x_train,y_train)
y_pred = knn.predict(x_test)
score.append(metrics.accuracy_score(y_test,y_pred))
import matplotlib.pyplot as plt
# %matplotlib inline
plt.plot(k_range,score)
plt.xlabel('Values of k')
plt.ylabel('Accuracy')
| 1 Data-Sciecne-Basics/iris_dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <a href="https://www.python.org/"><img src="https://upload.wikimedia.org/wikipedia/commons/thumb/f/f8/Python_logo_and_wordmark.svg/390px-Python_logo_and_wordmark.svg.png" style="max-width: 200px; display: inline" alt="Python"/></a> [pour Statistique et Science des Données](https://github.com/wikistat/Intro-Python)
# # Eléments de programmation en <a href="https://www.python.org/"><img src="https://upload.wikimedia.org/wikipedia/commons/thumb/f/f8/Python_logo_and_wordmark.svg/390px-Python_logo_and_wordmark.svg.png" style="max-width: 150px; display: inline" alt="Python"/></a> pour Calcul Scientifique - Statistique
# **Résumé**: Compléments de programmation en python: structures de contrôle, programmation fonctionnelle (map, reduce, lambda), introduction aux classes et objets.
#
# ## Introduction
#
# L'objectif de ce tutoriel est d'introduire quelques outils et concepts plus avancés de la programmation en Python pour dans le but d'améliorer la performance et la lisibilité des codes. Les notions de classe, de programmation objet et celle de *programmation fonctionnelle* qui en découle sont fondamentales. Elles sont fondamentales pour le bon usage de certaines librairie dont *Scikit-learn*.
# ## 1 Structures de contrôle
#
# ### 1.1 Structure itérative `for`
#
# Une boucle `for` permet, comme dans la pluspart des langages de programmation, de parcourir les éléments d'un objet *itérable*. En python cela peut-être une liste, un tuple, une chaîne de caractères (*string*), mais également des objets spécialement conçus pour cela, appelés *iterator*.
#
# Syntaxe de la structure `for`:
#
# `for` *variable* `in range` *iterator*:
# <br/> instruction
#
# La ligne `for` se termine par deux points ':'. Le bloc de codes à l'intérieur de la boucle est *indenté*.
#
# Voici comment parcourir différentes structures itératives en se souvenant que: **de façon générale, il faut éviter les boucles `for` dans un langage interprété** en utilisant les autres fonctionnalités (section 2) prévues dans les librairies pour parcourir des tableaux ou matrices.
#
#
#
# ##### `range`
# La fonction `range` produit une liste d'entiers mais aussi un objet itérable d'entiers utilisés au fur et à mesure des itérations de la boucle.
#
# **Attention** `xrange` de python2 est remplacé par `range` en python3.
# + code_folding=[]
for i in range(5):
print (i)
# -
# L'appel à la fonction range(n) permet d'itérer sur les entiers de 0 à n-1 mais il est possible de spécifier des intervalles des valeurs de chaque pas d'itération: `range(2,11,2) range(10,0,-1)`
# ##### Strings
# Les chaînes de caractères sont également des objets parcourables avec la boucle `for`: caractère après caractère.
for character in "Hi There!":
print (character)
# ##### Dictionnaires
# Un dictionnaire peut être parcouru terme à terme, cependant, comme la pluspart des objets python, les dictionnaires possèdent des fonctions permettant de les transformer en une liste:
#
# .items()
# .keys()
# .values()
#
# ou un *iterator*
#
# .iteritems()
# .iterkeys()
# .itervalues()
#
#
# Les dictionnaires n'ont pas de structure ordonnée, les valeurs ne s'affichent pas forcément dans l'ordre dans lequel on les a entrées.
dico={"a":1,"b":2,"c":3}
for k in dico.keys():
print (k)
# Les fonctions `.items()` et `.iteritems()` permettent de parcourir les couples (clés, objets) des dictionnaires.
# Si une seule variable itérative est spécifiée, celle-ci est un-tuple.
for kv in dico.items():
print (kv)
# Si deux variables itératives sont spécifiées, la première est la clé, la seconde la valeur correspondante
for k,v in dico.items():
print (k)
print (v)
# Si la clé ou la valeur du dictionnaire n'est pas nécessaire, elle n'est pas stockée dans une variable en utilisant "_"
for k,_ in dico.items():
print (k)
# #### Librarie *Itertools*
# La librairie native de python *itertools* possède de nombreuses fonctions générant des *iterators*.
# Plus d'exemples dans la [documentation](https://docs.python.org/3/library/itertools.html).
import itertools
#zip : Concatenne les éléments de plusieurs objets itérables
zip_list = []
for k in zip('ABCD',[1,2,3,4],"wxyz"):
zip_list.append(k)
print("zip")
print(zip_list)
#permutation : retourne tous les arrangements possibles de liste de longueur n.
permutation_list = []
for k in itertools.permutations("ABCD",2):
permutation_list.append(k)
print("permutations")
print(permutation_list)
# #### One-Line Statement
# Après avoir parouru différents éléments d'un objet itérable et pour enregistrer un résultat pour chaque étape dans une liste, il est possible d'écrire la boucle `for` sur une seule ligne de la manière suivante:
#
# [result *for* variable in *iterator*]
# +
# Version1
A1=[]
for k in range(10):
A1.append(k*k)
# Version 2
A2 = [k*k for k in range(10)]
print(A1,A2)
# -
# #### Affectation des indices
# Des expressions telles que x = x + 1 ou x = x - 2 apparaissent très souvent dans le corps des boucles. Le language Python permet de simplifier ces notations.
a = 17
s = "hi"
a += 3 # Equivalent to a = a + 3
a -= 3 # Equivalent to a = a - 3
a *= 3 # Equivalent to a = a * 3
a /= 3 # Equivalent to a = a / 3
a %= 3 # Equivalent to a = a % 3
s += " there" # Equivalent to s = s + “ there"
# ### 1.2 Structure itérative `while`
# La boucle *for* permet de parcourir l'ensemble des éléments d'un objet itérable, ou un nombre déterminé d'éléments de ce dernier. Cependant, ce nombre n'est pas toujours prévisible et il est possible de parcourir ces éléments et arrêter le parcours lorsqu'une condition est respéctée ou non. C'est l'objet de l'instruction `while` avec la syntaxe:
#
# `while` *condition*:
# <br/> instructions
#
# Cette instruction permet de répéter en boucle les instructions jusqu'à ce que la *condition* soit vérifiée.
# Incrémentation de `count` jusqu'à ce qu'elle dépasse la valeur 100.000
count = 1
while count <= 100000:
count += 1
print (count)
# L'instruction `break` permet de sortir de la boucle `while` même si sa condition est respectée.
while True:
number = int(input("Enter the numeric grade: "))
if number >= 0 and number <= 100:
break
else:
print ("Error: grade must be between 100 and 0" )
print (number)
# ### 1.3 Structures conditionnelles `if - else`
# L'instruction conditionelle `if-else` est une des plus communes en programmation informatique. En python, elle se présente sous la forme suivante:
#
# `if` *condition*:
# <br/> instructions 1
# <br/> `else`:
# <br/> Instructions 2
#
number=1
if number==1:
print (True)
else:
print (False)
# Lorsque plus de deux alternative sont possibles, utiliser l'instruction `elif` pour énumérer les différentes possibilités.
number=13
if number<5:
print("A")
elif number <10:
print("B")
elif number <20:
print("C")
else:
print("D")
# #### One-Line Statement
# Comme pour la boucle for, il est possible d'écrire l'instruction `if_else` en une seule ligne lorsque le code est simple.
number=10
"A" if number >10 else "B"
# #### One line with for Loop
# Différentes combinaisons sont possibles pour associer à la fois l'instruction `if_else` avec une boucle `for` en une ligne.
#Sélectionne uniquement les valeur paire
l1 = [k for k in range(10) if k%2==0]
l1
#Retourne "even" si l'élement k est pair, "odd" sinon.
l2 = ["even" if k%2==0 else "odd" for k in range(10)]
l2
# ## 2 Programmation fonctionnelle
# L'utilisation de *higher-order functions* permet d'éxecuter rapidement des schémas classiques:
#
# * appliquer la même fonction aux éléments d'une liste,
# * séléctionner, ou non, les différents éléments d'une liste selon une certaine condition,
# * ...
#
# **Important**: il s'agit ici d'introduitre les éléments de *programmation fonctionnelle*, présents dans Python, et utilisés systématiquement, car *parallélisable*, dans des architectures distribuées (*e. g. Hadoop, Spark*).
#
# ### 2.1 `map`
#
# La première de cette fonction est la fonction `map`. Elle permet d'appliquer une fonction sur toutes les valeurs d'une liste et retourne une nouvelle liste avec les résultats correspondants.
#
import random
numbers = [random.randrange(-10,10) for k in range(10)]
abs_numbers = map(abs,numbers) # Applique la fonction "valeur absolue" à tout les élements de la liste
print(numbers,list(abs_numbers))
# +
def first_capital_letters(txt):
if txt[0].islower():
txt = txt[0].upper()+txt[1:]
return txt
name=["Jason","bryan","hercule","Karim"]
list(map(first_capital_letters,name))
# -
# ### 2.1 `filter`
#
# La fonction `filter` permet d'appliquer une fonction test à chaque valeur d'une liste. Si la fonction test est vérifiée, la valeur est ajoutée dans une nouvelle liste. Sinon, la valeur n'est pas prise en compte. La nouvelle liste, constitué de toutes les valeures "positive" selon la fonction test, est retournée.
def is_odd(n):
return n % 2 == 1
list(filter(is_odd,range(20)))
# ### 2.2 `reduce`
#
# La dernière fonction est la fonction `reduce`. Cette approche est loin d'être intuitive. Le meilleur moyen de comprendre le mode d'emploi de cette fonction est d'utiliser un exemple.
#
# L'objectif est de calculer la somme de tous les entier de 0 à 9.
#
# - Générer dans un premier temps la liste contenant tout ces éléments `r10 = [0,1,2,3,4,5,6,7,8,9]`
# - La fonction `reduce`, applique une première fois la fonction `sum_and_print` sur les deux premiers éléments de la liste.
# - Exécution récursive: la fonction `sum_and_print` est appliquée sur le résultat de la première opération et sur le 3ème éléments de la liste
# - Itération récursive jusqu'à ce que tous les éléments de la liste soient parcourus
#
# La fonction `reduce` a donc deux arguments: une fonction et une liste.
#
# +
import functools
def sum_and_print(x,y):
print("Input: ", x,y)
print("Output: ", y)
return x+y
r10 = range(10)
res =functools.reduce(sum_and_print, r10)
print(res)
# -
# Par défaut, la fonction passée en paramètre de la fonction `reduce` effectue sa première opérations sur les deux premiers éléments de la listes passés en paramètre. Mais il est possible de spécifier une valeur initiale en troisième paramètre. La première opération sera alors effectuée sur cette valeur initiale et le premier élément de la liste.
# +
def somme(x,y):
return x+y
r10 = range(10)
res =functools.reduce(somme, r10,1000)
print(res)
# -
# ### 2.4 `lambda`
# L'utilisation de fonctions génériques permet de simplifier le code mais il est coûteux de définir une nouvelle fonction qui peut ne pas être réutilisée, comme par exemple celle de l'exemple précédent.
# L'appel `lambda` permet de créer une fonction de façon temporaire. La définition de ces fonctions est assez restrictive, puisqu'elle implique une définition sur *une seule ligne*, et ne permet pas d'assignation.
#
# Autre point important, l'exécution de cette fonction sur des données distribuées est implicitement parallélisée.
#
# Ainsi les précédents exemples peuvent-être ré-écrits de la manière suivante:
name=["Jason","bryan","hercule","Karim"]
list(map(lambda x : x[0].upper()+x[1:] if x[0].islower() else x,name))
list(filter(lambda x : x % 2 == 1 ,range(10)))
r10 = range(10)
res =functools.reduce(lambda x,y:x+y, r10,1000)
res
# ## 3 Classes et objets
# ### 3.1 Définitions et exemples
# Les classes sont des objets communs à tous les langages orientés objets. Ce sont des objets constitués de
#
# - *attributs*: des paramètres fixes, de différentes natures, attribués à l'objet,
# - *méthodes*: des fonctions qui permettent d'appliquer des transformations sur ces attributs.
#
# Ci dessous on définit une classe "Elève" dans laquelle un élève est décrit par son nom, son prénom et ses notes. On notera la *convention de nommage* des méthodes qui commence par une minuscule, mais qui possède une majuscule à chaque début de nouveau mot.
class Eleve:
"""Classe définissant un élève caractérisé par:
- son nom
- son prénom
- ses notes
"""
def __init__(self, nom, prenom): #constructeur de la classe
""" Construit un élève avec les nom et prenom passé en paramètre et une liste de notes vide."""
self._nom = nom
self._prenom=prenom
self._notes = []
def getNom(self):
""" retourne le nom de l'élève """
return self._nom
def getNotes(self):
""" retourne les notes de l'élève"""
return self._notes
def getNoteMax(self):
""" retourne la note max de l'élève"""
return max(self._notes)
def getMean(self):
""" retourne la moyenne de l'élève"""
return np.mean(self._notes)
def getNbNote(self):
""" retourne le nombre de note de l'élève"""
return len(self._notes)
def addNote(self, note):
""" ajoute la note 'note' à la liste de note de l'élève"""
self._notes.append(note)
# Toutes les classes sont composées d'un *constructeur* qui a pour nom \_\_init\_\_. Il s'agit d'une méthode spéciale d'instance que Python reconnaît et sait utiliser dans certains contextes. La fonction \_\_init\_\_ est automatiquement appelée à la création d'une nouvelle classe et prend en paramètre `self`, qui représente l'objet instantié, et les différents attributs nécessaires à sa création.
eleve1 = Eleve("Jean","Bon")
eleve1._nom
# Les attributs de la classe sont directement accessibles de la manière suivante:
#
# `objet.nomDeLAtribbut`
#
# Cependant, par convention, il est conseillé de définir une méthode pour avoir accès à cet objet.
# Les méthodes qui permettent d'accéder à des attributs de l'objets sont appelés des *accessors*. Dans la classe élève, les méthodes commençant par `get` sont des *accessors*.
eleve1.getNom()
# Les méthodes permettant de modifier les attributs d'un objet sont appelées des *mutators*. La fonction `addNote`, qui permet d'ajouter une note à la liste de notes de l'élève est un *mutator*.
print(eleve1.getNotes())
eleve1.addNote(15)
print(eleve1.getNotes())
for k in range(10):
eleve1.addNote(np.random.randint(20))
print (eleve1.getNbNote())
print (eleve1.getNoteMax())
print (eleve1.getMean())
# ### 3.2 Héritage
#
# L'*héritage* est une fonctionnalité qui permet de définir une classe "fille" à partir d'une autre classe "mère". La classe fille hérite alors automatiquement de tous les attributs et de toutes les méthodes de la classe mère.
class EleveSpecial(Eleve):
def __init__(self, nom, prenom, optionName):
Eleve.__init__(self, nom, prenom)
self._optionName = optionName
self._optionNotes = []
def getNotesOption(self):
""" retourne les notes de l'élève"""
return self._optionNotes
def addNoteOption(self, note):
""" ajoute la note 'note' à la liste de note de l'élève"""
self._optionNotes.append(note)
eleve2 = EleveSpecial("Sam","Stress","latin")
eleve2.addNote(14)
print (eleve2.getNotes())
eleve2.addNoteOption(12)
print (eleve2.getNotesOption())
# ### 3.3 Classes de *Scikit-learn*
#
# Les méthodes d'apprentissage statistique de la librairie *Scikit-learn* sont un parfait exemple d'utilisation de classes. Sans entrer en détail dans l'implémentation de cette méthode, voici l'exemple de la Régression linéaire par moindres carrés.
#
# **Toutes** les fonctions de *Scikit-learn* sont définies sur de modèle. Voir dans le [tutoriel d'apprentissage statistique](https://github.com/wikistat/Intro-Python) avec *Scikit-learn* comment ces propriétés sont utilisées pour enchaîner (*pipeline*) des exécutions.
#
# L'objet [`LinearRegression`](http://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression.html) est une classe qui est initée avec des attributs suivants:
#
# - `fit_intercept` : si le terme constant doit être estimé ou considéré à 0
# - `normalize` : si le jeux d'apprentissage doit être normalisé avant la regression
# - `copy_X` : si le jeux d'apprentissage doit être copié pour éviter les effets de bords
# - `n_jobs` : le nombre de processeur à utiliser
from sklearn.linear_model import LinearRegression
lr = LinearRegression(fit_intercept=True, normalize=False, copy_X=True, n_jobs=1)
print (lr.fit_intercept, lr.normalize, lr.copy_X, lr.n_jobs)
# La classe `LinearRegression` possède également des attributs qui sont mis à jour à l'aide des *méthodes*:
#
# - `coef_` : coefficients estimés
# - `residues_` : somme des résidus
# - `intercept_` : terme constant
#
# La méthode `fit` de cette classe est un *mutator*. Elle prend en paramètre le jeux d'apprentissage, `X_train` et la variable réponse correspondante `Y_train` pour estimer les paramètres de la regression linéaire et ainsi mettre à jour les attributs correspondants.
X_train=[[0, 0], [1, 1], [2, 2]]
Y_train = [0, 1, 2]
lr.fit (X_train, Y_train)
lr.coef_
# La classe `LinearRegression` possède aussi d'autres *méthodes* qui utilisent les attributs de la classe. Par exemple:
#
# - `predict`: estime la prévision de la variable réponse d'un jeu test `X_test`
# - `score`: retourne le coefficient *R2* de qualité de la prévision.
#
X_test = [[1.5,1.5],[2,4],[7.3,7.1]]
Y_test = [1.5,2.4,7]
pred = lr.predict(X_test)
s = lr.score(X_test,Y_test)
print(pred,s)
# ## 4 *Packing* et *Unpacking*
# Section plus technique qui peut être sautée en première lecture.
#
# L'opérateur `*` permet, selon la situation, de "paquéter" ou "dépaquéter" les éléments d'une liste.
#
# L'opérateur `**` permet, selon la situation, de "paquéter" ou "dépaquéter" les éléments d'un dictionnaire.
# ### 4.1 *Unpacking*
#
# Dans l'exemple ci-dessous. Les opérateurs \* et \*\* dépaquettent les listes et dictionnaires pour les passer en arguments de fonctions.
# +
def unpacking_list_and_print(a, b):
print (a)
print (b)
listarg = [3,4]
unpacking_list_and_print(*listarg)
def unpacking_dict_and_print(k1=0, k2=0):
print (k1)
print (k2)
dictarg = {'k1':4, 'k2':8}
unpacking_dict_and_print(**dictarg)
# -
# ### 4.2 *Packing*
# Ces opérateurs sont surtout utiles dans le sens du "packing". Les fonctions sont alors définies de sorte à recevoir un nombre inconnu d'argument qui seront ensuite "paquétés" et traités dans la fonction.
#
# L'argument `*args` permet à la fonction de recevoir un nombre supplémentaire inconnu d'arguments sans mot-clef associé.
# +
def packing_and_print_args(required_arg, *args):
print ("arg Nécessaire:", required_arg)
for i, arg in enumerate(args):
print ("args %d:" %i, arg)
packing_and_print_args(1, "two", 3)
packing_and_print_args(1, "two", [1,2,3],{"a":1,"b":2,"c":3})
# -
# L'argument `**kwargs` permet à la fonction de recevoir un nombre supplémentaire inconnu d'arguments avec mot-clef.
# +
def packing_and_print_kwargs(def_kwarg=2, **kwargs):
print ("kwarg défini:", def_kwarg)
for i,(k,v) in enumerate(kwargs.items()):
print ("kwarg %d:" %i ,k , v)
packing_and_print_kwargs(def_kwarg=1, sup_arg1="two", sup_arg2=3)
packing_and_print_kwargs(sup_arg1="two", sup_arg2=3, sup_arg3=[1,2,3])
# -
# Les arguments `*args` et `**kwargs` peuvent être combinés dans une autre fonctions.
# +
def packing_and_print_args_and_kwargs(required_arg ,def_kwarg=2, *args, **kwargs):
print ("arg Nécessaire:", required_arg)
for i, arg in enumerate(args):
print ("args %d:" %i, arg)
print ("kwarg défini:", def_kwarg)
for i,(k,v) in enumerate(kwargs.items()):
print ("kwarg %d:" %i ,k , v )
packing_and_print_args_and_kwargs(1, "two", [1,2,3] ,sup_arg1="two", sup_arg2=3 )
# -
# Ces deux opérateurs sont très utiles pour gérer des classes liées par des héritages. Les arguments `*args **kwargs` permettent alors de gérer la tranmission de cet héritage sans avoir à redéfinir les arguments à chaque étape.
# +
class Objet(object):
def __init__(self, attribut=None, *args, **kwargs):
print (attribut)
class Objet2Point0(Objet):
def __init__(self, *args, **kwargs):
super(Objet, self).__init__(*args, **kwargs)
class Objet3Point0(Objet2Point0):
def __init__(self,attribut2=None, *args, **kwargs):
super(Objet2Point0, self).__init__(*args, **kwargs)
print (attribut2)
my_data = {'attribut': 'Argument1', 'attribut2': 'Argument2'}
Objet3Point0(**my_data)
# -
# ## Référence
#
# **<NAME>. et <NAME>.** (2010). *Fundamentals of Python: From First Programs Through Data Structures*, Course Technology.
#
# **Wikistat** (2019) *Programmation, classes, objets, programmation fonctionnelle.* [notebook](https://github.com/wikistat/Intro-Python/blob/master/Cal4-PythonProg.ipynb)
| TPs/TP2-PythonProg.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.1 64-bit
# language: python
# name: python38164bitb3ebfd1fa0594a1c9d5c617333c2c1a4
# ---
# # Promotion Feature
# This notebook follows the orders
# ## Loading our data...
# +
import numpy as np
import pandas as pd
from utils import read_data, process_time, merge_data, promo_detector, promo_detector_fixed
import seaborn as sns
import matplotlib.pyplot as plt
import sys
sys.path.append("../../main/datasets/")
# -
# !ls ../../main/datasets/
infos, items, orders = read_data("../../main/datasets/")
print("Sanity checks...", infos.shape, items.shape, orders.shape)
orders.head()
# ## Preprocessing our orders
# These cells were taken from Bruno's "1.1-First Model" Notebook, that can be found in this repository.
process_time(orders)
def promo_detector_fixed(orders, aggregation=True, mode=True):
"""
This function adds a "promotion" column at "orders.csv".
It verifies if an item of an order is being sold cheaper than it's prices "mode"/"mean".
Case affirmative, a '1' will be added in 'promotion' column in the line of the order.
Parameters: orders -> Orders DataFrame
aggregation -> Flag that mantains or not the "salesPriceMode" in our returned DataFrame
True => Return will have the column
mode -> Decision method flag (Default 'True'). If "True", the function will
use the 'mode' of the prices to decide if an item is being sold below it's normal price.
If 'False', we'll use the "mean" of the prices.
Returns: our orders Dataframe with 2 new columns ("salesPriceMode" and "promotion")
"""
new_df = pd.DataFrame()
def agregationMode(x): return x.value_counts().index[0] if mode else 'mean'
for i in range(13, -1, -1):
# Getting an itemID / salesPriceMode Dataframe
# salesPriceMode column will store the
# 'mean'/'mode' of our items
current_agg = orders.loc[orders.group_backwards > i].groupby(['itemID']).agg(salesPriceMode=('salesPrice', agregationMode))
current_agg['promotion'] = 0
orders_copy = orders.loc[orders.group_backwards == i + 1].copy()
current_orders_with_promotion = pd.merge(orders_copy, current_agg, how='inner', left_on='itemID', right_on='itemID')
# For every item whose salesPrice is lower than the 'mean'/'mode',
# we'll attribute 1 to it's position in 'promotion' column
current_orders_with_promotion.loc[current_orders_with_promotion['salesPrice'] <
current_orders_with_promotion['salesPriceMode'], 'promotion'] = 1
new_df = pd.concat([new_df, current_orders_with_promotion])
week_13 = orders.loc[orders.group_backwards == 13].copy()
week_13['salesPriceMode'] = 0
week_13['promotion'] = 0
new_df = pd.concat([new_df, week_13])
if (not(aggregation)):
new_df.drop(
'salesPriceMode', axis=1, inplace=True)
new_df.sort_values(by=['group_backwards', 'itemID'], inplace=True)
return new_df
new_df = promo_detector_fixed(orders)
new_df.sort_values(by=['group_backwards', 'itemID'], inplace=True)
orders = promo_detector(orders)
# Sanity checking...
orders.loc[orders['promotion'] == 1]
def promotionAggregation(orders, items, promotionMode='mean', timeScale='group_backwards', salesPriceMode='mean'):
"""The 'promotion' feature is, originally, given by sale. This function aggregates it into the selected
time scale.
Parameters
-------------
orders : A pandas DataFrame with all the sales.
items: A pandas DataFrame with the infos about all items
promotionMode : A pandas aggregation compatible data type;
The aggregation mode of the 'promotion' feature
timeScale : A String with the name of the column containing the time signature.
E.g.: 'group_backwards'
salesPriceMode : A pandas aggregation compatible data type;
The aggregation mode of the 'salesPrice' feature
"""
df = orders.groupby([timeScale, 'itemID'], as_index=False).agg(
{'order': 'sum', 'promotion': promotionMode, 'salesPrice': salesPriceMode})
print(df)
items_copy = items.copy()
df.rename(columns={'order': 'orderSum', 'promotion': f'promotion_{promotionMode}',
'salesPrice': f'salesPrice_{salesPriceMode}'}, inplace=True)
return pd.merge(df, items_copy, how='left', left_on=['itemID'], right_on=['itemID'])
df = promotionAggregation(orders, items)
df.loc[df.itemID == 1]
| dora/pre-processing-features/promotionFeatureLeakFix.ipynb |
# +
from __future__ import print_function
import numpy as np
import tensorflow as tf
from six.moves import cPickle as pickle
from six.moves import range
pickle_file = 'notMNIST.pickle'
with open(pickle_file, 'rb') as f:
save = pickle.load(f)
train_dataset = save['train_dataset']
train_labels = save['train_labels']
valid_dataset = save['valid_dataset']
valid_labels = save['valid_labels']
test_dataset = save['test_dataset']
test_labels = save['test_labels']
del save # hint to help gc free up memory
print('Training set', train_dataset.shape, train_labels.shape)
print('Validation set', valid_dataset.shape, valid_labels.shape)
print('Test set', test_dataset.shape, test_labels.shape)
image_size = 28
num_labels = 10
def reformat(dataset, labels):
dataset = dataset.reshape((-1, image_size * image_size)).astype(np.float32)
# Map 0 to [1.0, 0.0, 0.0 ...], 1 to [0.0, 1.0, 0.0 ...]
labels = (np.arange(num_labels) == labels[:,None]).astype(np.float32)
return dataset, labels
# +
train_dataset, train_labels = reformat(train_dataset, train_labels)
valid_dataset, valid_labels = reformat(valid_dataset, valid_labels)
test_dataset, test_labels = reformat(test_dataset, test_labels)
print('Training set', train_dataset.shape, train_labels.shape)
print('Validation set', valid_dataset.shape, valid_labels.shape)
print('Test set', test_dataset.shape, test_labels.shape)
train_subset = 10000
batch_size = 128
graph = tf.Graph()
with graph.as_default():
# Input data. For the training data, we use a placeholder that will be fed
# at run time with a training minibatch.
tf_train_dataset = tf.placeholder(tf.float32,
shape=(batch_size, image_size * image_size))
tf_train_labels = tf.placeholder(tf.float32, shape=(batch_size, num_labels))
tf_valid_dataset = tf.constant(valid_dataset)
tf_test_dataset = tf.constant(test_dataset)
# Variables.
weights = tf.Variable(
tf.truncated_normal([image_size * image_size, num_labels]))
biases = tf.Variable(tf.zeros([num_labels]))
# Training computation.
logits = tf.matmul(tf_train_dataset, weights) + biases
loss = tf.reduce_mean(
tf.nn.softmax_cross_entropy_with_logits(labels=tf_train_labels, logits=logits))
# Optimizer.
optimizer = tf.train.GradientDescentOptimizer(0.5).minimize(loss)
# Predictions for the training, validation, and test data.
train_prediction = tf.nn.softmax(logits)
valid_prediction = tf.nn.softmax(
tf.matmul(tf_valid_dataset, weights) + biases)
test_prediction = tf.nn.softmax(tf.matmul(tf_test_dataset, weights) + biases)
num_steps = 801
def accuracy(predictions, labels):
return (100.0 * np.sum(np.argmax(predictions, 1) == np.argmax(labels, 1))
/ predictions.shape[0])
with tf.Session(graph=graph) as session:
tf.global_variables_initializer().run()
print("Initialized")
for step in range(num_steps):
# Pick an offset within the training data, which has been randomized.
# Note: we could use better randomization across epochs.
offset = (step * batch_size) % (train_labels.shape[0] - batch_size)
# Generate a minibatch.
batch_data = train_dataset[offset:(offset + batch_size), :]
batch_labels = train_labels[offset:(offset + batch_size), :]
# Prepare a dictionary telling the session where to feed the minibatch.
# The key of the dictionary is the placeholder node of the graph to be fed,
# and the value is the numpy array to feed to it.
feed_dict = {tf_train_dataset : batch_data, tf_train_labels : batch_labels}
_, l, predictions = session.run(
[optimizer, loss, train_prediction], feed_dict=feed_dict)
if (step % 500 == 0):
print("Minibatch loss at step %d: %f" % (step, l))
print("Minibatch accuracy: %.1f%%" % accuracy(predictions, batch_labels))
print("Validation accuracy: %.1f%%" % accuracy(
valid_prediction.eval(), valid_labels))
print("Test accuracy: %.1f%%" % accuracy(test_prediction.eval(), test_labels))
| tutorial/01_DNN/dnn_SGD.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# ## 2d prediction maps
#
# Inspirations:
#
# * [Plot classifier comparison - scikit-learn](https://scikit-learn.org/stable/auto_examples/classification/plot_classifier_comparison.html)
# * [TensorFlow playground](https://playground.tensorflow.org/)
# * [Which Machine Learning algorithm are you?](https://github.com/stared/which-ml-are-you)
#
# Take a note that is feature is experimental as of 0.4.1.
#
# In this case it uses PyTorch, but can be attached to other things. You need to use a different `predict` method.
# +
from sklearn import datasets
from sklearn.model_selection import train_test_split
import torch
from torch import nn, optim
from torch.utils.data import TensorDataset, DataLoader
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
from livelossplot import PlotLosses
from livelossplot import matplotlib_subplots
# -
# try with make_moons
X, y = datasets.make_circles(noise=0.2, factor=0.5, random_state=1)
X_train, X_test, y_train, y_test = \
train_test_split(X, y, test_size=.4, random_state=42)
# plot them
cm_bright = ListedColormap(['#FF0000', '#0000FF'])
plt.scatter(X_train[:, 0], X_train[:, 1], c=y_train, cmap=cm_bright)
plt.scatter(X_test[:, 0], X_test[:, 1], c=y_test, cmap=cm_bright, alpha=0.3)
# +
# PyTorch technicalities - loading and trainin
dataloaders = {
'train':
DataLoader(TensorDataset(torch.from_numpy(X_train).float(), torch.from_numpy(y_train).long()),
batch_size=32,
shuffle=True, num_workers=4),
'validation':
DataLoader(TensorDataset(torch.from_numpy(X_test).float(), torch.from_numpy(y_test).long()),
batch_size=32,
shuffle=False, num_workers=4)
}
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
def train_model(model, criterion, optimizer, num_epochs=10,
liveloss=PlotLosses()):
model = model.to(device)
for epoch in range(num_epochs):
logs = {}
for phase in ['train', 'validation']:
if phase == 'train':
model.train()
else:
model.eval()
running_loss = 0.0
running_corrects = 0
for inputs, labels in dataloaders[phase]:
inputs = inputs.to(device)
labels = labels.to(device)
outputs = model(inputs)
loss = criterion(outputs, labels)
if phase == 'train':
optimizer.zero_grad()
loss.backward()
optimizer.step()
_, preds = torch.max(outputs, 1)
running_loss += loss.item() * inputs.size(0)
running_corrects += (preds == labels.data).sum().item()
epoch_loss = running_loss / len(dataloaders[phase].dataset)
epoch_acc = running_corrects / len(dataloaders[phase].dataset)
prefix = ''
if phase == 'validation':
prefix = 'val_'
logs[prefix + 'log loss'] = epoch_loss
logs[prefix + 'accuracy'] = epoch_acc
liveloss.update(logs)
liveloss.draw()
# +
# an old-school neural network: a multi-layer perceptron
class MLP(nn.Module):
def __init__(self, hidden_size=3, activation=nn.ReLU()):
super().__init__()
self.fc = nn.Sequential(
nn.Linear(2, hidden_size),
activation,
nn.Linear(hidden_size, 2)
)
def forward(self, x):
x = self.fc(x)
return x
# +
model = MLP(6)
plot2d = matplotlib_subplots.Plot2d(model, X_train, y_train,
valiation_data=(X_test, y_test),
margin=0.2, h=0.02)
plot2d.predict = plot2d._predict_pytorch
liveloss = PlotLosses(cell_size=(5, 6), extra_plots=[plot2d])
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters(), lr=1e-1)
train_model(model, criterion, optimizer, num_epochs=30,
liveloss=liveloss)
# +
model = MLP(3, activation=nn.Sigmoid())
plot2d = matplotlib_subplots.Plot2d(model, X_train, y_train,
valiation_data=(X_test, y_test),
margin=0.2, h=0.02)
plot2d.predict = plot2d._predict_pytorch
liveloss = PlotLosses(cell_size=(5, 6), extra_plots=[plot2d])
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters(), lr=1e-1)
train_model(model, criterion, optimizer, num_epochs=100,
liveloss=liveloss)
# -
# if needed, we can draw it as a separate plot
plot2d.draw()
| examples/2d_prediction_maps.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Problem 4
# ### In this notebook I would like to highlight my visualization, machine learning, and data analysis skills as well as my work using neural networks to perform image classification.
# ### The goal of this problem was to create a model that could accurately classify the images that were presented to it (see below).
import numpy as np
np.random.seed(1337) # for reproducibility
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense, Dropout, Activation, Flatten
from keras.layers import Convolution2D, MaxPooling2D
from keras.utils import np_utils
import matplotlib.pyplot as plt
(X_train, y_train), (X_test, y_test) = mnist.load_data()
# +
batch_size = 256
nb_classes = 10
nb_epoch = 16
# input image dimensions
img_rows, img_cols = 28, 28
# number of convolutional filters to use
nb_filters = 36
# size of pooling area for max pooling
pool_size = (3, 3)
# convolution kernel size
kernel_size = 3 #(3, 3)
# -
X_train = X_train.reshape(X_train.shape[0], img_rows, img_cols,1)
X_test = X_test.reshape(X_test.shape[0], img_rows, img_cols,1)
input_shape = (img_rows, img_cols,1)
X_train = X_train.astype('float32')
X_test = X_test.astype('float32')
X_train /= 255
X_test /= 255
print('X_train shape:', X_train.shape)
print(X_train.shape[0], 'train samples')
print(X_test.shape[0], 'test samples')
y_train
# convert class vectors to binary class matrices
Y_train = np_utils.to_categorical(y_train, nb_classes)
Y_test = np_utils.to_categorical(y_test, nb_classes)
Y_test
plt.imshow(X_train[5999][:,:,:]);
plt.imshow(X_test[500][:,:,:]);
# +
# Creating Model that is different than class example
# -
def reset_weights(model):
"""This function re-initializes model weights at each compile"""
for layer in model.layers:
if isinstance(layer, tf.keras.Model):
reset_weights(layer)
continue
for k, initializer in layer.__dict__.items():
if "initializer" not in k:
continue
# find the corresponding variable
var = getattr(layer, k.replace("_initializer", ""))
var.assign(initializer(var.shape, var.dtype))
# +
model = Sequential()
model.add(Convolution2D(nb_filters, kernel_size,
padding='same',
input_shape=input_shape))
model.add(Activation('relu'))
model.add(Convolution2D(nb_filters, kernel_size))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=pool_size))
model.add(Flatten())
model.add(Dense(256))
model.add(Activation('relu'))
model.add(Dense(nb_classes))
model.add(Activation('softmax'))
model.compile(loss='categorical_crossentropy',
optimizer='adadelta',
metrics=['accuracy'])
# -
# train model
history = model.fit(X_train, Y_train, batch_size=batch_size, epochs=nb_epoch,
verbose=1, validation_data=(X_test, Y_test))
score = model.evaluate(X_test, Y_test, verbose=0)
print('Test score:', score[0])
print('Test accuracy:', score[1])
model.summary()
# ### (a)
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# ### (b)
from sklearn.metrics import classification_report, confusion_matrix
from sklearn.metrics import multilabel_confusion_matrix
y_pred = model.predict(X_test, batch_size=64, verbose=1)
y_pred_bool = np.argmax(y_pred, axis=1)
print(classification_report(y_test, y_pred_bool))
# ### (c)
print(multilabel_confusion_matrix(y_test, y_pred_bool))
# ### (d) Including more dropout layers
# +
model1 = Sequential()
model1.add(Convolution2D(nb_filters, kernel_size,
padding='same',
input_shape=input_shape))
model1.add(Activation('relu'))
model1.add(Convolution2D(nb_filters, kernel_size))
model1.add(Dropout(0.45))
model1.add(Activation('relu'))
model1.add(MaxPooling2D(pool_size=pool_size))
model1.add(Dropout(0.4))
model1.add(Flatten())
model1.add(Dense(128))
model1.add(Activation('relu'))
model1.add(Dropout(0.5))
model1.add(Dense(nb_classes))
model1.add(Activation('softmax'))
model1.compile(loss='categorical_crossentropy',
optimizer='adadelta',
metrics=['accuracy'])
# -
history = model.fit(X_train, Y_train, batch_size=batch_size, epochs=nb_epoch,
verbose=1, validation_data=(X_test, Y_test))
# ### After some experimentation with this dropout function, the model is definitely performing better, I used values between 0.1-0.5. It is interesting to see an improvement in the accuracy of the model, as the dropout is randomly selected neurons to be ignored during training. This ensures that the model doesn't become too specialized to the training data. The final accuracy was 0.797 which is better than the previous model which had a test accuracy of 0.7019
| Machine Learning/Machine Learning (Image Classification).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="FfybwkHr2CrY" colab={"base_uri": "https://localhost:8080/", "height": 404} outputId="de174192-1bc4-4d54-f22b-897b0117fe02"
from google.colab import drive
drive.mount('/content/drive')
# + id="uEuHnBKJ3wqH"
import os
os.chdir('/content/drive/MyDrive/Handwritten_Text_Recognition/src')
# + id="oHJBMNNM4X1e"
# !python main.py
# + id="PwLkdEPk4cZe"
# !python main.py --validate
# + id="LjYXyF-65Oro"
| Handwritten-Text-Recognition-main/Handwritten_Text_Recognition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Find worst cases
# \begin{equation}
# \begin{array}{rl}
# \mathcal{F}_L =& \dfrac{4 K I H r}{Q_{in}(1+f)}\\
# u_{c} =& \dfrac{-KI\mathcal{F}_L}{\theta \left(\mathcal{F}_L+1\right)}\\
# \tau =& -\dfrac{r}{|u_{c}|}\\
# C_{\tau,{\rm decay}}=& C_0 \exp{\left(-\lambda \tau \right)}\\
# C_{\tau,{\rm filtr}}=& C_0 \exp{\left(-k_{\rm att} \tau \right)}\\
# C_{\tau,{\rm dilut}} =& C_{in} \left( \dfrac{Q_{in}}{u_c \Delta y \Delta z} \right)\\
# C_{\tau,{\rm both}} =& \dfrac{C_{\rm in}Q_{\rm in}}{u_c \Delta y H \theta} \exp{\left(-\lambda\dfrac{r}{|u_c|}\right)}
# \end{array}
# \end{equation}
# +
# %reset -f
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
from os import system
import os
from matplotlib.gridspec import GridSpec
from drawStuff import *
import jupypft.attachmentRateCFT as CFT
import jupypft.plotBTC as BTC
''' GLOBAL CONSTANTS '''
PI = 3.141592
THETA = 0.35
# +
def flowNumber():
return (4.0*K*I*H*r) / (Qin*(1+f))
def uChar():
'''Interstitial water velocity'''
return -(K*I*flowNumber())/(THETA*(flowNumber() + 1))
def tChar():
return -r/uChar()
def cDecay():
return C0 * np.exp(-decayRate * tChar())
def cAttach():
return C0 * np.exp(-attchRate * tChar())
def cDilut():
return (C0 * Qin) / (-uChar() * delY * delZ * THETA)
def cBoth():
return (C0 * Qin) / (-uChar() * delY * delZ * THETA) * np.exp(-decayRate * tChar())
def cTrice():
return (C0 * Qin) / (-uChar() * delY * delZ * THETA) * np.exp(-(decayRate+attchRate) * tChar())
def findSweet():
deltaConc = np.abs(cBoth() - np.max(cBoth()))
return np.argmin(deltaConc)
# +
K = 10**-2
Qin = 0.24/86400
f = 10
H = 20
r = 40
I = 0.001
C0 = 1.0
qabs = np.abs(uChar()*THETA)
kattDict = dict(
dp = 1.0E-7,
dc = 2.0E-3,
q = qabs,
theta = THETA,
visco = 0.0008891,
rho_f = 999.79,
rho_p = 1050.0,
A = 5.0E-21,
T = 10. + 273.15,
alpha = 0.01)
decayRate = 3.5353E-06
attchRate,_ = CFT.attachmentRate(**kattDict)
delY,delZ = 1.35,H
# -
print("Nondim Flow = {:.2E}".format(flowNumber()))
print("Charac. Vel = {:.2E} m/s".format(uChar()))
print("Charac. time = {:.2E} s".format(tChar()))
print("Rel concenc. due decay = {:.2E}".format(cDecay()))
print("Rel conc. due dilution = {:.2E}".format(cDilut()))
print("Rel conc. due attachmt = {:.2E}".format(cAttach()))
print("Rel conc. due both eff = {:.2E}".format(cBoth()))
print("Rel conc. due three ef = {:.2E}".format(cTrice()))
# # Plot v. 1
# + active=""
# I = 10**np.linspace(-5,0,num=100)
#
# cDec = cDecay()
# cDil = cDilut()
# cAtt = cAttach()
# cBot = cBoth()
# cAll = cTrice()
#
# i = findSweet()
#
# worstC = cBot[i]
# worstI = I[i]
#
# fig, axs = plt.subplots(2,2,sharex=True, sharey=False,\
# figsize=(12,8),gridspec_kw={"height_ratios":[1,4],"hspace":0.04,"wspace":0.35})
#
# bbox = dict(boxstyle='round', facecolor='mintcream', alpha=0.90)
#
# arrowprops = dict(
# arrowstyle="->",
# connectionstyle="angle,angleA=90,angleB=40,rad=5")
#
# fontdict = dict(size=12)
#
# annotation = \
# r"$\bf{-\log(C/C_0)} = $" + "{:.1f}".format(-np.log10(worstC)) + \
# "\n@" + r" $\bf{I} = $" + "{:.1E}".format(worstI)
#
# information = \
# r"$\bf{K}$" + " = {:.1E} m/s".format(K) + "\n"\
# r"$\bf{H}$" + " = {:.1f} m".format(H) + "\n"\
# r"$\bf{r}$" + " = {:.1f} m".format(r) + "\n"\
# r"$\bf{Q_{in}}$" + " = {:.2f} m³/d".format(Qin*86400) + "\n"\
# r"$\bf{f}$" + " = {:.1f}".format(f) + "\n"\
# r"$\bf{\lambda}$" + " = {:.2E} 1/s".format(decayRate)
#
# #########################################
# # Ax1 - Relative concentration
# ax = axs[1,0]
# ax.plot(I,cDec,label="Due decay",lw=3,ls="dashed",alpha=0.8)
# ax.plot(I,cDil,label="Due dilution",lw=3,ls="dashed",alpha=0.8)
# ax.plot(I,cAtt,label="Due attachment",lw=2,ls="dashed",alpha=0.6)
# ax.plot(I,cBot,label="Decay + dilution",lw=3,c='k',alpha=0.9)
# ax.plot(I,cAll,label="Overall effect",lw=3,c='gray',alpha=0.9)
#
# ax.set(xscale="log",yscale="log")
# ax.set(xlim=(1.0E-4,1.0E-1),ylim=(1.0E-10,1))
# ax.legend(loc="lower left",shadow=True)
#
# ax.annotate(annotation,(worstI,worstC),
# xytext=(0.05,0.85), textcoords='axes fraction',
# bbox=bbox, arrowprops=arrowprops)
#
# ax.text(0.65,0.05,information,bbox=bbox,transform=ax.transAxes)
#
# ax.set_xlabel("Water table gradient\n$I$ [m/m]",fontdict=fontdict)
# ax.set_ylabel("Relative Concentration\n$C/C_0$ [-]",fontdict=fontdict)
# ####################################
# # Ax2 - log-removals
# ax = axs[1,1]
# ax.plot(I,-np.log10(cDec),label="Due decay",lw=3,ls="dashed",alpha=0.8)
# ax.plot(I,-np.log10(cDil),label="Due dilution",lw=3,ls="dashed",alpha=0.8)
# ax.plot(I,-np.log10(cAtt),label="Due attachment",lw=2,ls="dashed",alpha=0.6)
# ax.plot(I,-np.log10(cBot),label="Decay + dilution",lw=3,c='k',alpha=0.9)
# ax.plot(I,-np.log10(cAll),label="Overall effect",lw=3,c='gray',alpha=0.9)
#
# ax.set(xscale="log")
# ax.set(xlim=(1.0E-4,1.0E-1),ylim=(0,10))
# ax.legend(loc="upper left",shadow=True)
#
# ax.annotate(annotation,(worstI,-np.log10(worstC)),
# xytext=(0.65,0.55), textcoords='axes fraction',
# bbox=bbox, arrowprops=arrowprops)
#
# ax.text(0.65,0.70,information,bbox=bbox,transform=ax.transAxes)
#
# ax.set_xlabel("Water table gradient\n$I$ [m/m]",fontdict=fontdict)
# ax.set_ylabel("log-reductions\n$-\log(C/C_0)$ [-]",fontdict=fontdict)
#
# ####################################
# #Flow number
# for ax in axs[0,:]:
# ax.plot(I,flowNumber(),label="flowNumber",lw=3,c="gray")
# ax.axhline(y=1.0)
# ax.set_xscale("log")
# ax.set_yscale("log")
# ax.xaxis.set_tick_params(which="both",labeltop='on',top=True,bottom=False)
# ax.set_ylabel("Nondim. flow number\n$\mathcal{F}_L$ [-]")
#
# ####################################<
# #Line worst case scenario
# for axc in axs:
# for ax in axc:
# ax.axvline(x=I[i], lw=1, ls="dashed", c="red",alpha=0.5)
#
# plt.show()
# -
# # v.2 With PFLOTRAN result
# +
listOfFiles = os.listdir("LittleValidation_MASSBALANCES")
listOfFiles.sort()
IPFLO = [float(s[9:16]) for s in listOfFiles]
CPFLO = BTC.get_endConcentrations(
"LittleValidation_MASSBALANCES",
indices={'t':"Time [d]",\
'q':"ExtractWell Water Mass [kg/d]",\
'm':"ExtractWell Vaq [mol/d]"
},
normalizeWith=dict(t=1.0,q=kattDict['rho_f']/1000.,m=1.0))
# +
listOfFiles = os.listdir("LittleValidation_MASSBALANCES_Att")
listOfFiles.sort()
IPFLO2 = [float(s[8:15]) for s in listOfFiles]
CPFLO2 = BTC.get_endConcentrations(
"LittleValidation_MASSBALANCES_Att",
indices={'t':"Time [d]",\
'q':"ExtractWell Water Mass [kg/d]",\
'm':"ExtractWell Vaq [mol/d]"
},
normalizeWith=dict(t=1.0,q=kattDict['rho_f']/1000.,m=1.0))
# -
# Theoretical stuff
I = 10**np.linspace(-5,0,num=100)
cDec = cDecay()
cDil = cDilut()
cAtt = cAttach()
cBot = cBoth()
cAll = cTrice()
i = findSweet()
worstC = cBot[i]
worstI = I[i]
# + active=""
# fig, axs = plt.subplots(2,2,sharex=True, sharey=False,\
# figsize=(10,8),gridspec_kw={"height_ratios":[1,10],"hspace":0.04,"wspace":0.02})
#
# bbox = dict(boxstyle='round', facecolor='mintcream', alpha=0.90)
#
# arrowprops = dict(
# arrowstyle="->",
# connectionstyle="angle,angleA=90,angleB=40,rad=5")
#
# fontdict = dict(size=12)
#
# annotation = \
# r"$\bf{-\log(C/C_0)} = $" + "{:.1f}".format(-np.log10(worstC)) + \
# "\n@" + r" $\bf{I} = $" + BTC.sci_notation(worstI)
#
# information = \
# r"$\bf{K} = $" + BTC.sci_notation(K) + " m/s\n"\
# r"$\bf{H}$" + " = {:.1f} m".format(H) + "\n"\
# r"$\bf{r}$" + " = {:.1f} m".format(r) + "\n"\
# r"$\bf{Q_{in}}$" + " = {:.2f} m³/d".format(Qin*86400) + "\n"\
# r"$\bf{f}$" + " = {:.1f}".format(f) + "\n"\
# r"$\bf{\lambda_{\rm aq}} = $" + BTC.sci_notation(decayRate) + " s⁻¹\n"\
# r"$\bf{k_{\rm att}} = $" + BTC.sci_notation(attchRate) + " s⁻¹"
#
# ####################################
# # Ax2 - log-removals
# ax = axs[1,0]
#
# symbols=dict(dil="\u25A2",dec="\u25B3",att="\u25CE")
#
# ax.plot(I,-np.log10(cDil),\
# label="Due dilution " + symbols['dil'],\
# lw=3,ls="dashed",alpha=0.6,c='crimson')
# ax.plot(I,-np.log10(cDec),\
# label="Due decay " + symbols['dec'],\
# lw=3,ls="dashed",alpha=0.6,c='indigo')
# ax.plot(I,-np.log10(cAtt),\
# label="Due attachment " + symbols['att'],\
# lw=2,ls="dashed",alpha=0.5,c='olive')
# ax.plot(I,-np.log10(cBot),\
# label=symbols['dil'] + " + " + symbols['dec'],\
# lw=3,c='k',alpha=0.9,zorder=2)
# ax.plot(I,-np.log10(cAll),\
# label=symbols['dil'] + " + " + symbols['dec'] + " + " + symbols['att'],\
# lw=2,c='gray',alpha=0.9,zorder=2)
#
# ax.set(xscale="log")
# ax.set(xlim=(1.0E-4,1.0E-1),ylim=(0,9.9))
# ax.legend(loc="upper right",shadow=True,ncol=1,\
# title="Potential flow prediction",title_fontsize=11)
#
# ax.annotate(annotation,(worstI,-np.log10(worstC)),
# xytext=(0.65,0.55), textcoords='axes fraction',
# bbox=bbox, arrowprops=arrowprops)
#
# ax.set_xlabel("Water table gradient\n$I$ [m/m]",fontdict=fontdict)
# ax.set_ylabel("log-reductions\n$-\log(C/C_0)$ [-]",fontdict=fontdict)
#
# ####################################
# # Ax2 - log-removals
# ax = axs[1,1]
# ax.plot(I,-np.log10(cBot),
# lw=3,c='k',alpha=0.9)
# ax.plot(IPFLO,-np.log10(CPFLO),zorder=2,\
# label= symbols['dil'] + " + " + symbols['dec'],\
# lw=0,ls='dotted',c='k',alpha=0.9,\
# marker="$\u25CA$",mec='k',mfc='k',ms=10)
#
# ax.plot(I,-np.log10(cAll),\
# lw=2,c='gray',alpha=0.9)
# ax.plot(IPFLO2,-np.log10(CPFLO2),zorder=2,\
# label= symbols['dil'] + " + " + symbols['dec'] + " + " + symbols['att'],\
# lw=0,ls='dotted',c='gray',alpha=0.9,\
# marker="$\u2217$",mec='gray',mfc='gray',ms=10)
# #####
#
# ax.set(xscale="log")
# ax.set(xlim=(1.0E-4,9.0E-2),ylim=(0,9.9))
# ax.legend(loc="lower left",shadow=True,ncol=1,\
# labelspacing=0.4,mode=None,\
# title="PFLOTRAN run",title_fontsize=11)
# ax.set_xlabel("Water table gradient\n$I$ [m/m]",fontdict=fontdict)
# ax.yaxis.tick_right()
#
# ax.text(0.60,0.70,information,bbox=bbox,transform=ax.transAxes)
# ax.text(I[i],0.5,"Worst\ncase",\
# ha='center',va='center',weight='semibold',\
# bbox=dict(boxstyle='square', fc='white', ec='red'))
#
# ####################################
# #Flow number
# ax = axs[0,0]
# ax.plot(I,flowNumber(),label="flowNumber",lw=3,c="gray")
# #ax.axhline(y=1.0)
# ax.set_xscale("log")
# ax.set_yscale("log")
# ax.xaxis.set_tick_params(which="both",labeltop='on',top=False,bottom=False)
# #ax.set_ylabel("Nondim. flow number\n$\mathcal{F}_L$ [-]")
#
# ####################################<
# #Line worst case scenario
# axs[0,0].axvline(x=I[i], lw=2, ls="dotted", c="red",alpha=0.5)
# axs[1,0].axvline(x=I[i], lw=2, ls="dotted", c="red",alpha=0.5)
# axs[1,1].axvline(x=I[i], lw=2, ls="dotted", c="red",alpha=0.5)
#
# ###############################
# # Information box
# ax = axs[0,1]
# ax.axis('off')
#
# plt.show()
# +
fig, axs = plt.subplots(1,1,figsize=(5,5))
fontdict = dict(size=12)
lines = {}
information = \
r"$\bf{K} = $" + BTC.sci_notation(K) + " m/s\n"\
r"$\bf{H}$" + " = {:.1f} m".format(H) + "\n"\
r"$\bf{r}$" + " = {:.1f} m".format(r) + "\n"\
r"$\bf{Q_{in}}$" + " = {:.2f} m³/d".format(Qin*86400) + "\n"\
r"$\bf{f}$" + " = {:.1f}".format(f) + "\n"\
r"$\bf{\lambda_{\rm aq}} = $" + BTC.sci_notation(decayRate) + " s⁻¹"\
####################################
# Ax2 - log-removals
ax = axs
lines['Dilution'] = ax.plot(I,-np.log10(cDil),\
label="Due dilution",\
lw=3,ls="dashed",alpha=0.99,c='#f1a340')
lines['Decay'] = ax.plot(I,-np.log10(cDec),\
label="Due decay",\
lw=3,ls="dashdot",alpha=0.99,c='#998ec3')
#ax.plot(I,-np.log10(cAtt),\
# label="Due attachment " + symbols['att'],\
# lw=2,ls="dashed",alpha=0.5,c='olive')
lines['Both'] = ax.plot(I,-np.log10(cBot),\
label="Combined effect",\
lw=3,c='k',alpha=0.9,zorder=2)
#ax.plot(I,-np.log10(cAll),\
# label=symbols['dil'] + " + " + symbols['dec'] + " + " + symbols['att'],\
# lw=2,c='gray',alpha=0.9,zorder=2)
lines['PFLOT'] = ax.plot(IPFLO,-np.log10(CPFLO),zorder=2,\
label= "BIOPARTICLE model",\
lw=0,ls='dotted',c='k',alpha=0.9,\
marker="$\u25CA$",mec='k',mfc='k',ms=10)
ax.set(xscale="log")
ax.set(xlim=(5.0E-5,5.0E-1),ylim=(-0.5,9.9))
ax.set_xlabel("Water table gradient\n$I$ [m/m]",fontdict=fontdict)
ax.set_ylabel("log-reductions\n$-\log(C/C_0)$ [-]",fontdict=fontdict)
## Legend
whitebox = ax.scatter([1],[1],c="white",marker="o",s=1,alpha=0)
handles = [lines['PFLOT'][0],
lines['Both'][0]]
labels = [lines['PFLOT'][0].get_label(),
"PF model"]
plt.legend(handles, labels,
loc="upper right",ncol=1,\
edgecolor='gray',facecolor='mintcream',labelspacing=1)
##aNNOTATIONS
rotang = -1.0
bbox = dict(boxstyle='square', fc='w', ec='#998ec3',lw=1.5)
ax.text(0.85,0.10,'Inactivation',ha='center',va='center',
fontsize=11,rotation=rotang,bbox=bbox,transform=ax.transAxes)
rotang = 21.0
bbox = dict(boxstyle='square', fc='w', ec='#f1a340',lw=1.5)
ax.text(0.12,0.21,'Dilution',ha='center',va='center',
fontsize=11,rotation=rotang,bbox=bbox,transform=ax.transAxes)
rotang = -0.0
bbox = dict(boxstyle='square', fc='k', ec=None,lw=0.0)
ax.text(0.55,0.45,'Combined',c='w',ha='center',va='center',weight='bold',
fontsize=11,rotation=rotang,bbox=bbox,transform=ax.transAxes)
bbox = dict(boxstyle='round,pad=0.5,rounding_size=0.3', fc='whitesmoke', alpha=0.90,ec='whitesmoke')
#ax.text(0.60,0.64,information,bbox=bbox,transform=ax.transAxes,fontsize=10)
fig.savefig(fname="SweetpointOnlyDecay.png",transparent=False,dpi=300,bbox_inches="tight")
#plt.show()
# +
fig, axs = plt.subplots(1,1,figsize=(5,5))
fontdict = dict(size=12)
lines = {}
information = \
r"$\bf{K} = $" + BTC.sci_notation(K) + " m/s\n"\
r"$\bf{H}$" + " = {:.1f} m".format(H) + "\n"\
r"$\bf{r}$" + " = {:.1f} m".format(r) + "\n"\
r"$\bf{Q_{in}}$" + " = {:.2f} m³/d".format(Qin*86400) + "\n"\
r"$\bf{f}$" + " = {:.1f}".format(f) + "\n"\
r"$\bf{\lambda_{\rm aq}} = $" + BTC.sci_notation(decayRate) + " s⁻¹\n"\
r"$\bf{k_{\rm att}} = $" + BTC.sci_notation(attchRate) + " s⁻¹"
####################################
# Ax2 - log-removals
ax = axs
lines['Dilution'] = ax.plot(I,-np.log10(cDil),\
label="Due dilution",\
lw=3,ls="dashed",alpha=0.99,c='#f1a340')
lines['Decay'] = ax.plot(I,-np.log10(cDec),\
label="Due decay",\
lw=3,ls="dashdot",alpha=0.99,c='#998ec3')
lines['Attach'] = ax.plot(I,-np.log10(cAtt),\
label="Due attachment",\
lw=3,ls="dotted",alpha=0.95,c='#4dac26')
#lines['Both'] = ax.plot(I,-np.log10(cBot),\
# label="Combined effect",\
# lw=3,c='k',alpha=0.9,zorder=2)
lines['Both'] = ax.plot(I,-np.log10(cAll),\
label="Combined\neffect",\
lw=3,c='#101613',alpha=0.99,zorder=2)
lines['PFLOT'] = ax.plot(IPFLO2,-np.log10(CPFLO2),zorder=2,\
label= "BIOPARTICLE model",\
lw=0,ls='dotted',c='k',alpha=0.9,\
marker="$\u2217$",mec='gray',mfc='k',ms=10)
ax.set(xscale="log")
ax.set(xlim=(5.0E-5,5.0E-1),ylim=(-0.5,9.9))
ax.set_xlabel("Water table gradient\n$I$ [m/m]",fontdict=fontdict)
ax.set_ylabel("log-reductions\n$-\log(C/C_0)$ [-]",fontdict=fontdict)
## Legend
whitebox = ax.scatter([1],[1],c="white",marker="o",s=1,alpha=0)
handles = [lines['PFLOT'][0],
lines['Both'][0]]
labels = [lines['PFLOT'][0].get_label(),
"PF model"]
plt.legend(handles, labels,
loc="upper right",ncol=1,\
edgecolor='gray',facecolor='mintcream',labelspacing=1)
##aNNOTATIONS
rotang = -82.0
bbox = dict(boxstyle='square', fc='w', ec='#998ec3',lw=1.5)
ax.text(0.12,0.85,'Inactivation',c='k',fontweight='normal',ha='center',va='center',
fontsize=11,rotation=rotang,bbox=bbox,transform=ax.transAxes)
rotang = 21.0
bbox = dict(boxstyle='square', fc='w', ec='#f1a340',lw=1.5)
ax.text(0.12,0.21,'Dilution',ha='center',va='center',
fontsize=11,rotation=rotang,bbox=bbox,transform=ax.transAxes)
rotang = -1.5
bbox = dict(boxstyle='square', fc='w', ec='#4dac26',ls='-',lw=1.5)
ax.text(0.85,0.10,'Filtration',ha='center',va='center',
fontsize=11,rotation=rotang,bbox=bbox,transform=ax.transAxes)
rotang = -0.0
bbox = dict(boxstyle='square', fc='#101613', ec='#101613')
ax.text(0.65,0.52,'Combined',c='w',ha='center',va='center',weight='bold',
fontsize=11,rotation=rotang,bbox=bbox,transform=ax.transAxes)
bbox = dict(boxstyle='round,pad=0.5,rounding_size=0.3', fc='whitesmoke', alpha=0.90,ec='whitesmoke')
#ax.text(0.60,0.58,information,bbox=bbox,transform=ax.transAxes,fontsize=10)
fig.savefig(fname="SweetpointAllConsidered.png",transparent=False,dpi=300,bbox_inches="tight")
plt.show()
# -
# # v.3 Together as filtration is assumed
# + active=""
# fig, axs = plt.subplots(1,2,figsize=(10,5))
# bbox = dict(boxstyle='round,pad=0.5,rounding_size=0.3', facecolor='mintcream', alpha=0.90)
# fontdict = dict(size=12)
# lines = {}
# information = \
# "Parameters:\n" + \
# r"$\bf{K} = $" + BTC.sci_notation(K) + " m/s\n"\
# r"$\bf{H}$" + " = {:.1f} m".format(H) + "\n"\
# r"$\bf{r}$" + " = {:.1f} m".format(r) + "\n"\
# r"$\bf{Q_{in}}$" + " = {:.2f} m³/d".format(Qin*86400) + "\n"\
# r"$\bf{f}$" + " = {:.1f}".format(f) + "\n"\
# r"$\bf{\lambda_{\rm aq}} = $" + BTC.sci_notation(decayRate) + " s⁻¹"\
#
# ####################################
# # Ax2 - log-removals
# ax = axs[0]
#
# lines['Dilution'] = ax.plot(I,-np.log10(cDil),\
# label="Due dilution",\
# lw=3,ls="dashed",alpha=0.99,c='#f1a340')
# lines['Decay'] = ax.plot(I,-np.log10(cDec),\
# label="Due decay",\
# lw=3,ls="dashdot",alpha=0.99,c='#998ec3')
# #ax.plot(I,-np.log10(cAtt),\
# # label="Due attachment " + symbols['att'],\
# # lw=2,ls="dashed",alpha=0.5,c='olive')
# lines['Both'] = ax.plot(I,-np.log10(cBot),\
# label="Combined effect",\
# lw=3,c='k',alpha=0.9,zorder=2)
# #ax.plot(I,-np.log10(cAll),\
# # label=symbols['dil'] + " + " + symbols['dec'] + " + " + symbols['att'],\
# # lw=2,c='gray',alpha=0.9,zorder=2)
# lines['PFLOT'] = ax.plot(IPFLO,-np.log10(CPFLO),zorder=2,\
# label= "PFLOTRAN results",\
# lw=0,ls='dotted',c='k',alpha=0.9,\
# marker="$\u25CA$",mec='k',mfc='k',ms=10)
#
# ax.set(xscale="log")
# ax.set(xlim=(5.0E-5,5.0E-1),ylim=(0,9.9))
# ax.text(1.04,0.55,information,bbox=bbox,transform=ax.transAxes)
#
# ax.set_xlabel("Water table gradient\n$I$ [m/m]",fontdict=fontdict)
# ax.set_ylabel("log-reductions\n$-\log(C/C_0)$ [-]",fontdict=fontdict)
#
# ## Legend
# whitebox = ax.scatter([1],[1],c="white",marker="o",s=1,alpha=0)
#
# handles = [whitebox,
# lines['Dilution'][0],
# lines['Decay'][0],
# lines['Both'][0],
# whitebox,
# lines['PFLOT'][0]]
#
# labels = ['PF model',
# lines['Dilution'][0].get_label(),
# lines['Decay'][0].get_label(),
# lines['Both'][0].get_label(),
# '',
# lines['PFLOT'][0].get_label(),]
#
# plt.legend(handles, labels,
# loc="center left",ncol=1,bbox_to_anchor=(1.01,0.25),\
# edgecolor='k',facecolor='mintcream',labelspacing=1)
#
# bbox = dict(boxstyle='round,pad=0.5,rounding_size=0.3', facecolor='mintcream', alpha=0.90)
# fontdict = dict(size=12)
# lines = {}
# information = \
# "Parameters:\n" + \
# r"$\bf{K} = $" + BTC.sci_notation(K) + " m/s\n"\
# r"$\bf{H}$" + " = {:.1f} m".format(H) + "\n"\
# r"$\bf{r}$" + " = {:.1f} m".format(r) + "\n"\
# r"$\bf{Q_{in}}$" + " = {:.2f} m³/d".format(Qin*86400) + "\n"\
# r"$\bf{f}$" + " = {:.1f}".format(f) + "\n"\
# r"$\bf{\lambda_{\rm aq}} = $" + BTC.sci_notation(decayRate) + " s⁻¹\n"\
# r"$\bf{k_{\rm att}} = $" + BTC.sci_notation(attchRate) + " s⁻¹"
#
# ####################################
# # Ax2 - log-removals
# ax = axs[1]
#
# lines['Dilution'] = ax.plot(I,-np.log10(cDil),\
# label="Due dilution",\
# lw=3,ls="dashed",alpha=0.99,c='#f1a340')
# lines['Decay'] = ax.plot(I,-np.log10(cDec),\
# label="Due decay",\
# lw=3,ls="dashdot",alpha=0.99,c='#998ec3')
# lines['Attach'] = ax.plot(I,-np.log10(cAtt),\
# label="Due attachment",\
# lw=3,ls="dotted",alpha=0.95,c='#4dac26')
# #lines['Both'] = ax.plot(I,-np.log10(cBot),\
# # label="Combined effect",\
# # lw=3,c='k',alpha=0.9,zorder=2)
# lines['Both'] = ax.plot(I,-np.log10(cAll),\
# label="Combined effect",\
# lw=3,c='k',alpha=0.99,zorder=2)
#
# lines['PFLOT'] = ax.plot(IPFLO2,-np.log10(CPFLO2),zorder=2,\
# label= "PFLOTRAN results",\
# lw=0,ls='dotted',c='k',alpha=0.9,\
# marker="$\u2217$",mec='gray',mfc='k',ms=10)
#
# ax.set(xscale="log")
# ax.set(xlim=(5.0E-5,5.0E-1),ylim=(0,9.9))
# ax.text(1.04,0.55,information,bbox=bbox,transform=ax.transAxes)
#
# ax.set_xlabel("Water table gradient\n$I$ [m/m]",fontdict=fontdict)
# ax.set_ylabel("log-reductions\n$-\log(C/C_0)$ [-]",fontdict=fontdict)
#
# ## Legend
# whitebox = ax.scatter([1],[1],c="white",marker="o",s=1,alpha=0)
#
# handles = [whitebox,
# lines['Dilution'][0],
# lines['Decay'][0],
# lines['Attach'][0],
# lines['Both'][0],
# whitebox,
# lines['PFLOT'][0]]
#
# labels = ['PF model',
# lines['Dilution'][0].get_label(),
# lines['Decay'][0].get_label(),
# lines['Attach'][0].get_label(),
# lines['Both'][0].get_label(),
# '',
# lines['PFLOT'][0].get_label(),]
#
# plt.legend(handles, labels,
# loc="center left",ncol=1,bbox_to_anchor=(1.01,0.25),\
# edgecolor='k',facecolor='mintcream',labelspacing=1)
#
# plt.show()
# -
# ____
# # Find the worst case
# ## >> Geometric parameters $H$ and $r$
# + active=""
# K = 10**-2
# Qin = 0.24/86400
# f = 10
# C0 = 1.0
# decayRate = 3.5353E-06
#
# Harray = np.array([2.,5.,10.,20.,50.])
# rarray = np.array([5.,10.,40.,100.])
# Iarray = 10**np.linspace(-5,0,num=100)
#
# Ci = np.zeros([len(rarray),len(Harray)])
# Ii = np.zeros([len(rarray),len(Harray)])
# FLi = np.zeros([len(rarray),len(Harray)])
#
# for hi,H in enumerate(Harray):
# for ri,r in enumerate(rarray):
# i = findSweet()
#
# worstC = -np.log10(cBoth()[i])
# worstGradient = Iarray[i]
# worstFlowNumber = flowNumber()[i]
#
# Ci[ri,hi] = worstC
# Ii[ri,hi] = worstGradient
# FLi[ri,hi] = worstFlowNumber
# + active=""
# myLabels={"Title": { 0: r"$\bf{-\log (C_{\tau}/C_0)}$",
# 1: r"$\bf{I}$ (%)",
# 2: r"$\log(\mathcal{F}_L)$"},
# "Y": "Aquifer thickness\n$\\bf{H}$ (m)",
# "X": "Setback distance\n$\\bf{r}$ (m)"}
#
# threeHeatplots(data={"I":Ii.T,"C":Ci.T,"FL":FLi.T},\
# xlabel=Harray,ylabel=rarray,myLabels=myLabels);
# -
# ## >>Well parameters
# + active=""
# K = 10**-2
# H = 20
# r = 40
# C0 = 1.0
# decayRate = 3.5353E-06
#
# Qin_array = np.array([0.24,1.0,10.0,100.])/86400.
# f_array = np.array([1,10.,100.,1000.,10000.])
# Iarray = 10**np.linspace(-5,0,num=100)
#
# Ci = np.zeros([len(Qin_array),len(f_array)])
# Ii = np.zeros([len(Qin_array),len(f_array)])
# FLi = np.zeros([len(Qin_array),len(f_array)])
#
# for fi,f in enumerate(f_array):
# for qi,Qin in enumerate(Qin_array):
# i = findSweet()
# worstC = -np.log10(cBoth()[i])
# worstGradient = Iarray[i]
# worstFlowNumber = flowNumber()[i]
#
# Ci[qi,fi] = worstC
# Ii[qi,fi] = worstGradient
# FLi[qi,fi] = worstFlowNumber
#
# ### Plot heatmap
# myLabels={"Title": { 0: r"$\bf{-\log (C_{\tau}/C_0)}$",
# 1: r"$\bf{I}$ (%)",
# 2: r"$\log(\mathcal{F}_L)$"},
# "Y": "Extraction to injection ratio\n$\\bf{f}$ (-)",
# "X": "Injection flow rate\n$\\bf{Q_{in}}$ (m³/d)"}
#
# threeHeatplots(data={"I":Ii.T,"C":Ci.T,"FL":FLi.T},\
# xlabel=f_array,ylabel=np.round(Qin_array*86400,decimals=2),myLabels=myLabels);
# -
# ## Hydraulic conductivity
# + active=""
# Qin = 0.24/86400
# f = 10
# C0 = 1.0
# decayRate = 3.5353E-06
#
# Karray = 10.**np.array([-1.,-2.,-3.,-4.,-5.])
# rarray = np.array([5.,10.,40.,100.])
# Iarray = 10**np.linspace(-5,0,num=100)
#
# Ci = np.zeros([len(rarray),len(Karray)])
# Ii = np.zeros([len(rarray),len(Karray)])
# FLi = np.zeros([len(rarray),len(Karray)])
#
# for ki,K in enumerate(Karray):
# for ri,r in enumerate(rarray):
# i = findSweet()
#
# worstC = -np.log10(cBoth()[i])
# worstGradient = Iarray[i]
# worstFlowNumber = flowNumber()[i]
#
# Ci[ri,ki] = worstC
# Ii[ri,ki] = worstGradient
# FLi[ri,ki] = worstFlowNumber
#
# ### Plot heatmap
# myLabels={"Title": { 0: r"$\bf{-\log (C_{\tau}/C_0)}$",
# 1: r"$\bf{I}$ (%)",
# 2: r"$\log(\mathcal{F}_L)$"},
# "Y": "Hydraulic conductivity\n$\\bf{K}$ (m/s)",
# "X": "Setback distance\n$\\bf{r}$ (m)"}
#
# threeHeatplots(data={"I":Ii.T,"C":Ci.T,"FL":FLi.T},\
# xlabel=Karray,ylabel=rarray,myLabels=myLabels);
# + active=""
# K = 10**-2
# H = 20
# f = 10
# C0 = 1.0
# decayRate = 3.5353E-06
#
# Qin_array = np.array([0.24,1.0,10.0,100.])/86400.
# rarray = np.array([5.,10.,40.,100.])
# Iarray = 10**np.linspace(-5,0,num=100)
#
# Ci = np.zeros([len(rarray),len(Qin_array)])
# Ii = np.zeros([len(rarray),len(Qin_array)])
# FLi = np.zeros([len(rarray),len(Qin_array)])
#
# for qi,Qin in enumerate(Qin_array):
# for ri,r in enumerate(rarray):
# i = findSweet()
#
# worstC = -np.log10(cBoth()[i])
# worstGradient = Iarray[i]
# worstFlowNumber = flowNumber()[i]
#
# Ci[ri,qi] = worstC
# Ii[ri,qi] = worstGradient
# FLi[ri,qi] = worstFlowNumber
#
# myLabels={"Title": { 0: r"$\bf{-\log (C_{\tau}/C_0)}$",
# 1: r"$\bf{I}$ (%)",
# 2: r"$\log(\mathcal{F}_L)$"},
# "Y": "Injection flow rate\n$\\bf{Q_{in}}$ (m³/d)",
# "X": "Setback distance\n$\\bf{r}$ (m)"}
#
# threeHeatplots(data={"I":Ii.T,"C":Ci.T,"FL":FLi.T},\
# ylabel=rarray,xlabel=np.round(Qin_array*86400,decimals=2),myLabels=myLabels);
# + active=""
# K = 10**-2
# H = 20
# f = 10
# C0 = 1.0
# decayRate = 1.119E-5
#
# Qin_array = np.array([0.24,1.0,10.0,100.])/86400.
# rarray = np.array([5.,10.,40.,100.])
# Iarray = 10**np.linspace(-5,0,num=100)
#
# Ci = np.zeros([len(rarray),len(Qin_array)])
# Ii = np.zeros([len(rarray),len(Qin_array)])
# FLi = np.zeros([len(rarray),len(Qin_array)])
#
# for qi,Qin in enumerate(Qin_array):
# for ri,r in enumerate(rarray):
# i = findSweet()
#
# worstC = -np.log10(cBoth()[i])
# worstGradient = Iarray[i]
# worstFlowNumber = flowNumber()[i]
#
# Ci[ri,qi] = worstC
# Ii[ri,qi] = worstGradient
# FLi[ri,qi] = worstFlowNumber
#
# myLabels={"Title": { 0: r"$\bf{-\log (C_{\tau}/C_0)}$",
# 1: r"$\bf{I}$ (%)",
# 2: r"$\log(\mathcal{F}_L)$"},
# "Y": "Injection flow rate\n$\\bf{Q_{in}}$ (m³/d)",
# "X": "Setback distance\n$\\bf{r}$ (m)"}
#
# threeHeatplots(data={"I":Ii.T,"C":Ci.T,"FL":FLi.T},\
# ylabel=rarray,xlabel=np.round(Qin_array*86400,decimals=2),myLabels=myLabels);
# -
# ## PLOTRAN SIMULATION RESULTS
# + active=""
# minI = np.array([0.00046667, 0.0013 , 0.0048 , 0.012 , 0.00046667,
# 0.0013 , 0.0048 , 0.012 , 0.00053333, 0.0013 ,
# 0.0048 , 0.012 , 0.00056667, 0.0021 , 0.0053 ,
# 0.012 ])
#
# minC = np.array([2.2514572 , 2.62298917, 3.14213329, 3.51421485, 1.64182175,
# 2.00913676, 2.52461269, 2.89537637, 0.74130696, 1.0754177 ,
# 1.55071976, 1.90646243, 0.18705258, 0.39222131, 0.73428991,
# 1.00387133])
# + active=""
# Qin_array = np.array([0.24,1.0,10.0,100.])/86400.
# rarray = np.array([5.,10.,40.,100.])
# Iarray = 10**np.linspace(-5,0,num=100)
#
# Ci = np.zeros([len(rarray),len(Qin_array)])
# Ii = np.zeros([len(rarray),len(Qin_array)])
# FLi = np.zeros([len(rarray),len(Qin_array)])
#
# i = 0
# for qi,Qin in enumerate(Qin_array):
# for ri,r in enumerate(rarray):
# worstC = minC[i]
# worstGradient = minI[i]
#
# Ci[ri,qi] = worstC
# Ii[ri,qi] = worstGradient
#
# i += 1
#
# myLabels={"Title": { 0: r"$\bf{-\log (C_{\tau}/C_0)}$",
# 1: r"$\bf{I}$ (%)",
# 2: r"$\log(\mathcal{F}_L)$"},
# "Y": "Injection flow rate\n$\\bf{Q_{in}}$ (m³/d)",
# "X": "Setback distance\n$\\bf{r}$ (m)"}
#
# threeHeatplots(data={"I":Ii.T,"C":Ci.T,"FL":Ii.T},\
# ylabel=rarray,xlabel=np.round(Qin_array*86400,decimals=2),myLabels=myLabels);
# -
| notebooks/Concepts/Find worst case (1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
# https://news.daum.net/economic#1
economic = requests.get('https://news.daum.net/economic')
economic.status_code
from bs4 import BeautifulSoup
soup = BeautifulSoup(economic.content,'html.parser')
soup,type(soup)
soup.select('li > strong.tit_timenews > a[class="link_txt"]')
| scraping_bs4_daum_economic_exer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="1czVdIlqnImH"
# # Data Augmentation
# + [markdown] colab_type="text" id="1KD3ZgLs80vY"
# ### Goals
# In this notebook you're going to build a generator that can be used to help create data to train a classifier. There are many cases where this might be useful. If you are interested in any of these topics, you are welcome to explore the linked papers and articles!
#
# - With smaller datasets, GANs can provide useful data augmentation that substantially [improve classifier performance](https://arxiv.org/abs/1711.04340).
# - You have one type of data already labeled and would like to make predictions on [another related dataset for which you have no labels](https://www.nature.com/articles/s41598-019-52737-x). (You'll learn about the techniques for this use case in future notebooks!)
# - You want to protect the privacy of the people who provided their information so you can provide access to a [generator instead of real data](https://www.ahajournals.org/doi/full/10.1161/CIRCOUTCOMES.118.005122).
# - You have [input data with many missing values](https://arxiv.org/abs/1806.02920), where the input dimensions are correlated and you would like to train a model on complete inputs.
# - You would like to be able to identify a real-world abnormal feature in an image [for the purpose of diagnosis](https://link.springer.com/chapter/10.1007/978-3-030-00946-5_11), but have limited access to real examples of the condition.
#
# In this assignment, you're going to be acting as a bug enthusiast — more on that later.
#
# ### Learning Objectives
# 1. Understand some use cases for data augmentation and why GANs suit this task.
# 2. Implement a classifier that takes a mixed dataset of reals/fakes and analyze its accuracy.
# + [markdown] colab_type="text" id="wU8DDM6l9rZb"
# ## Getting Started
#
# ### Data Augmentation
# Before you implement GAN-based data augmentation, you should know a bit about data augmentation in general, specifically for image datasets. It is [very common practice](https://arxiv.org/abs/1712.04621) to augment image-based datasets in ways that are appropriate for a given dataset. This may include having your dataloader randomly flipping images across their vertical axis, randomly cropping your image to a particular size, randomly adding a bit of noise or color to an image in ways that are true-to-life.
#
# In general, data augmentation helps to stop your model from overfitting to the data, and allows you to make small datasets many times larger. However, a sufficiently powerful classifier often still overfits to the original examples which is why GANs are particularly useful here. They can generate new images instead of simply modifying existing ones.
#
# ### CIFAR
# The [CIFAR-10 and CIFAR-100](https://www.cs.toronto.edu/~kriz/learning-features-2009-TR.pdf) datasets are extremely widely used within machine learning -- they contain many thousands of “tiny” 32x32 color images of different classes representing relatively common real-world objects like airplanes and dogs, with 10 classes in CIFAR-10 and 100 classes in CIFAR-100. In CIFAR-100, there are 20 “superclasses” which each contain five classes. For example, the “fish” superclass contains “aquarium fish, flatfish, ray, shark, trout”. For the purposes of this assignment, you’ll be looking at a small subset of these images to simulate a small data regime, with only 40 images of each class for training.
#
# 
#
# ### Initializations
# You will begin by importing some useful libraries and packages and defining a visualization function that has been provided. You will also be re-using your conditional generator and functions code from earlier assignments. This will let you control what class of images to augment for your classifier.
# + colab={} colab_type="code" id="JfkorNJrnmNO"
import torch
import torch.nn.functional as F
import matplotlib.pyplot as plt
from torch import nn
from tqdm.auto import tqdm
from torchvision import transforms
from torchvision.utils import make_grid
from torch.utils.data import DataLoader
torch.manual_seed(0) # Set for our testing purposes, please do not change!
def show_tensor_images(image_tensor, num_images=25, size=(3, 32, 32), nrow=5, show=True):
'''
Function for visualizing images: Given a tensor of images, number of images, and
size per image, plots and prints the images in an uniform grid.
'''
image_tensor = (image_tensor + 1) / 2
image_unflat = image_tensor.detach().cpu()
image_grid = make_grid(image_unflat[:num_images], nrow=nrow)
plt.imshow(image_grid.permute(1, 2, 0).squeeze())
if show:
plt.show()
# + [markdown] colab_type="text" id="P1A1M6kpnfxw"
# #### Generator
# + colab={} colab_type="code" id="EvO7h0LYnEJZ"
class Generator(nn.Module):
'''
Generator Class
Values:
input_dim: the dimension of the input vector, a scalar
im_chan: the number of channels of the output image, a scalar
(CIFAR100 is in color (red, green, blue), so 3 is your default)
hidden_dim: the inner dimension, a scalar
'''
def __init__(self, input_dim=10, im_chan=3, hidden_dim=64):
super(Generator, self).__init__()
self.input_dim = input_dim
# Build the neural network
self.gen = nn.Sequential(
self.make_gen_block(input_dim, hidden_dim * 4, kernel_size=4),
self.make_gen_block(hidden_dim * 4, hidden_dim * 2, kernel_size=4, stride=1),
self.make_gen_block(hidden_dim * 2, hidden_dim, kernel_size=4),
self.make_gen_block(hidden_dim, im_chan, kernel_size=2, final_layer=True),
)
def make_gen_block(self, input_channels, output_channels, kernel_size=3, stride=2, final_layer=False):
'''
Function to return a sequence of operations corresponding to a generator block of DCGAN;
a transposed convolution, a batchnorm (except in the final layer), and an activation.
Parameters:
input_channels: how many channels the input feature representation has
output_channels: how many channels the output feature representation should have
kernel_size: the size of each convolutional filter, equivalent to (kernel_size, kernel_size)
stride: the stride of the convolution
final_layer: a boolean, true if it is the final layer and false otherwise
(affects activation and batchnorm)
'''
if not final_layer:
return nn.Sequential(
nn.ConvTranspose2d(input_channels, output_channels, kernel_size, stride),
nn.BatchNorm2d(output_channels),
nn.ReLU(inplace=True),
)
else:
return nn.Sequential(
nn.ConvTranspose2d(input_channels, output_channels, kernel_size, stride),
nn.Tanh(),
)
def forward(self, noise):
'''
Function for completing a forward pass of the generator: Given a noise tensor,
returns generated images.
Parameters:
noise: a noise tensor with dimensions (n_samples, input_dim)
'''
x = noise.view(len(noise), self.input_dim, 1, 1)
return self.gen(x)
def get_noise(n_samples, input_dim, device='cpu'):
'''
Function for creating noise vectors: Given the dimensions (n_samples, input_dim)
creates a tensor of that shape filled with random numbers from the normal distribution.
Parameters:
n_samples: the number of samples to generate, a scalar
input_dim: the dimension of the input vector, a scalar
device: the device type
'''
return torch.randn(n_samples, input_dim, device=device)
def combine_vectors(x, y):
'''
Function for combining two vectors with shapes (n_samples, ?) and (n_samples, ?)
Parameters:
x: (n_samples, ?) the first vector.
In this assignment, this will be the noise vector of shape (n_samples, z_dim),
but you shouldn't need to know the second dimension's size.
y: (n_samples, ?) the second vector.
Once again, in this assignment this will be the one-hot class vector
with the shape (n_samples, n_classes), but you shouldn't assume this in your code.
'''
return torch.cat([x, y], 1)
def get_one_hot_labels(labels, n_classes):
'''
Function for combining two vectors with shapes (n_samples, ?) and (n_samples, ?)
Parameters:
labels: (n_samples, 1)
n_classes: a single integer corresponding to the total number of classes in the dataset
'''
return F.one_hot(labels, n_classes)
# + [markdown] colab_type="text" id="qRk_8azSq3tF"
# ## Training
# Now you can begin training your models.
# First, you will define some new parameters:
#
# * cifar100_shape: the number of pixels in each CIFAR image, which has dimensions 32 x 32 and three channel (for red, green, and blue) so 3 x 32 x 32
# * n_classes: the number of classes in CIFAR100 (e.g. airplane, automobile, bird, cat, deer, dog, frog, horse, ship, truck)
# + colab={} colab_type="code" id="UpfJifVcmMhJ"
cifar100_shape = (3, 32, 32)
n_classes = 100
# + [markdown] colab_type="text" id="gJM9afuu0IuD"
# And you also include the same parameters from previous assignments:
#
# * criterion: the loss function
# * n_epochs: the number of times you iterate through the entire dataset when training
# * z_dim: the dimension of the noise vector
# * display_step: how often to display/visualize the images
# * batch_size: the number of images per forward/backward pass
# * lr: the learning rate
# * device: the device type
# + colab={} colab_type="code" id="sJlx2W71lUCv"
n_epochs = 10000
z_dim = 64
display_step = 500
batch_size = 64
lr = 0.0002
device = 'cuda'
# + [markdown] colab_type="text" id="jltxAMd00TRE"
# Then, you want to set your generator's input dimension. Recall that for conditional GANs, the generator's input is the noise vector concatenated with the class vector.
# + colab={} colab_type="code" id="tuSOzzpwlXl7"
generator_input_dim = z_dim + n_classes
# + [markdown] colab_type="text" id="ccQZRSYFXsHh"
# #### Classifier
#
# For the classifier, you will use the same code that you wrote in an earlier assignment (the same as previous code for the discriminator as well since the discriminator is a real/fake classifier).
# + colab={} colab_type="code" id="cVPxAjGSfYlX"
class Classifier(nn.Module):
'''
Classifier Class
Values:
im_chan: the number of channels of the output image, a scalar
n_classes: the total number of classes in the dataset, an integer scalar
hidden_dim: the inner dimension, a scalar
'''
def __init__(self, im_chan, n_classes, hidden_dim=32):
super(Classifier, self).__init__()
self.disc = nn.Sequential(
self.make_classifier_block(im_chan, hidden_dim),
self.make_classifier_block(hidden_dim, hidden_dim * 2),
self.make_classifier_block(hidden_dim * 2, hidden_dim * 4),
self.make_classifier_block(hidden_dim * 4, n_classes, final_layer=True),
)
def make_classifier_block(self, input_channels, output_channels, kernel_size=3, stride=2, final_layer=False):
'''
Function to return a sequence of operations corresponding to a classifier block;
a convolution, a batchnorm (except in the final layer), and an activation (except in the final
Parameters:
input_channels: how many channels the input feature representation has
output_channels: how many channels the output feature representation should have
kernel_size: the size of each convolutional filter, equivalent to (kernel_size, kernel_size)
stride: the stride of the convolution
final_layer: a boolean, true if it is the final layer and false otherwise
(affects activation and batchnorm)
'''
if not final_layer:
return nn.Sequential(
nn.Conv2d(input_channels, output_channels, kernel_size, stride),
nn.BatchNorm2d(output_channels),
nn.LeakyReLU(0.2, inplace=True),
)
else:
return nn.Sequential(
nn.Conv2d(input_channels, output_channels, kernel_size, stride),
)
def forward(self, image):
'''
Function for completing a forward pass of the classifier: Given an image tensor,
returns an n_classes-dimension tensor representing fake/real.
Parameters:
image: a flattened image tensor with im_chan channels
'''
class_pred = self.disc(image)
return class_pred.view(len(class_pred), -1)
# + [markdown] colab_type="text" id="tYXJTxM9pzZK"
# #### Pre-training (Optional)
#
# You are provided the code to pre-train the models (GAN and classifier) given to you in this assignment. However, this is intended only for your personal curiosity -- for the assignment to run as intended, you should not use any checkpoints besides the ones given to you.
# + colab={} colab_type="code" id="UXptQZcwrBrq"
# This code is here for you to train your own generator or classifier
# outside the assignment on the full dataset if you'd like -- for the purposes
# of this assignment, please use the provided checkpoints
class Discriminator(nn.Module):
'''
Discriminator Class
Values:
im_chan: the number of channels of the output image, a scalar
(MNIST is black-and-white, so 1 channel is your default)
hidden_dim: the inner dimension, a scalar
'''
def __init__(self, im_chan=3, hidden_dim=64):
super(Discriminator, self).__init__()
self.disc = nn.Sequential(
self.make_disc_block(im_chan, hidden_dim, stride=1),
self.make_disc_block(hidden_dim, hidden_dim * 2),
self.make_disc_block(hidden_dim * 2, hidden_dim * 4),
self.make_disc_block(hidden_dim * 4, 1, final_layer=True),
)
def make_disc_block(self, input_channels, output_channels, kernel_size=4, stride=2, final_layer=False):
'''
Function to return a sequence of operations corresponding to a discriminator block of the DCGAN;
a convolution, a batchnorm (except in the final layer), and an activation (except in the final layer).
Parameters:
input_channels: how many channels the input feature representation has
output_channels: how many channels the output feature representation should have
kernel_size: the size of each convolutional filter, equivalent to (kernel_size, kernel_size)
stride: the stride of the convolution
final_layer: a boolean, true if it is the final layer and false otherwise
(affects activation and batchnorm)
'''
if not final_layer:
return nn.Sequential(
nn.Conv2d(input_channels, output_channels, kernel_size, stride),
nn.BatchNorm2d(output_channels),
nn.LeakyReLU(0.2, inplace=True),
)
else:
return nn.Sequential(
nn.Conv2d(input_channels, output_channels, kernel_size, stride),
)
def forward(self, image):
'''
Function for completing a forward pass of the discriminator: Given an image tensor,
returns a 1-dimension tensor representing fake/real.
Parameters:
image: a flattened image tensor with dimension (im_chan)
'''
disc_pred = self.disc(image)
return disc_pred.view(len(disc_pred), -1)
def train_generator():
gen = Generator(generator_input_dim).to(device)
gen_opt = torch.optim.Adam(gen.parameters(), lr=lr)
discriminator_input_dim = cifar100_shape[0] + n_classes
disc = Discriminator(discriminator_input_dim).to(device)
disc_opt = torch.optim.Adam(disc.parameters(), lr=lr)
def weights_init(m):
if isinstance(m, nn.Conv2d) or isinstance(m, nn.ConvTranspose2d):
torch.nn.init.normal_(m.weight, 0.0, 0.02)
if isinstance(m, nn.BatchNorm2d):
torch.nn.init.normal_(m.weight, 0.0, 0.02)
torch.nn.init.constant_(m.bias, 0)
gen = gen.apply(weights_init)
disc = disc.apply(weights_init)
criterion = nn.BCEWithLogitsLoss()
cur_step = 0
mean_generator_loss = 0
mean_discriminator_loss = 0
for epoch in range(n_epochs):
# Dataloader returns the batches and the labels
for real, labels in dataloader:
cur_batch_size = len(real)
# Flatten the batch of real images from the dataset
real = real.to(device)
# Convert the labels from the dataloader into one-hot versions of those labels
one_hot_labels = get_one_hot_labels(labels.to(device), n_classes).float()
image_one_hot_labels = one_hot_labels[:, :, None, None]
image_one_hot_labels = image_one_hot_labels.repeat(1, 1, cifar100_shape[1], cifar100_shape[2])
### Update discriminator ###
# Zero out the discriminator gradients
disc_opt.zero_grad()
# Get noise corresponding to the current batch_size
fake_noise = get_noise(cur_batch_size, z_dim, device=device)
# Combine the vectors of the noise and the one-hot labels for the generator
noise_and_labels = combine_vectors(fake_noise, one_hot_labels)
fake = gen(noise_and_labels)
# Combine the vectors of the images and the one-hot labels for the discriminator
fake_image_and_labels = combine_vectors(fake.detach(), image_one_hot_labels)
real_image_and_labels = combine_vectors(real, image_one_hot_labels)
disc_fake_pred = disc(fake_image_and_labels)
disc_real_pred = disc(real_image_and_labels)
disc_fake_loss = criterion(disc_fake_pred, torch.zeros_like(disc_fake_pred))
disc_real_loss = criterion(disc_real_pred, torch.ones_like(disc_real_pred))
disc_loss = (disc_fake_loss + disc_real_loss) / 2
disc_loss.backward(retain_graph=True)
disc_opt.step()
# Keep track of the average discriminator loss
mean_discriminator_loss += disc_loss.item() / display_step
### Update generator ###
# Zero out the generator gradients
gen_opt.zero_grad()
# Pass the discriminator the combination of the fake images and the one-hot labels
fake_image_and_labels = combine_vectors(fake, image_one_hot_labels)
disc_fake_pred = disc(fake_image_and_labels)
gen_loss = criterion(disc_fake_pred, torch.ones_like(disc_fake_pred))
gen_loss.backward()
gen_opt.step()
# Keep track of the average generator loss
mean_generator_loss += gen_loss.item() / display_step
if cur_step % display_step == 0 and cur_step > 0:
print(f"Step {cur_step}: Generator loss: {mean_generator_loss}, discriminator loss: {mean_discriminator_loss}")
show_tensor_images(fake)
show_tensor_images(real)
mean_generator_loss = 0
mean_discriminator_loss = 0
cur_step += 1
def train_classifier():
criterion = nn.CrossEntropyLoss()
n_epochs = 10
validation_dataloader = DataLoader(
CIFAR100(".", train=False, download=True, transform=transform),
batch_size=batch_size)
display_step = 10
batch_size = 512
lr = 0.0002
device = 'cuda'
classifier = Classifier(cifar100_shape[0], n_classes).to(device)
classifier_opt = torch.optim.Adam(classifier.parameters(), lr=lr)
cur_step = 0
for epoch in range(n_epochs):
for real, labels in tqdm(dataloader):
cur_batch_size = len(real)
real = real.to(device)
labels = labels.to(device)
### Update classifier ###
# Get noise corresponding to the current batch_size
classifier_opt.zero_grad()
labels_hat = classifier(real.detach())
classifier_loss = criterion(labels_hat, labels)
classifier_loss.backward()
classifier_opt.step()
if cur_step % display_step == 0:
classifier_val_loss = 0
classifier_correct = 0
num_validation = 0
for val_example, val_label in validation_dataloader:
cur_batch_size = len(val_example)
num_validation += cur_batch_size
val_example = val_example.to(device)
val_label = val_label.to(device)
labels_hat = classifier(val_example)
classifier_val_loss += criterion(labels_hat, val_label) * cur_batch_size
classifier_correct += (labels_hat.argmax(1) == val_label).float().sum()
print(f"Step {cur_step}: "
f"Classifier loss: {classifier_val_loss.item() / num_validation}, "
f"classifier accuracy: {classifier_correct.item() / num_validation}")
cur_step += 1
# + [markdown] colab_type="text" id="ZYGOiy-xWHOH"
# ## Tuning the Classifier
# After two courses, you've probably had some fun debugging your GANs and have started to consider yourself a bug master. For this assignment, your mastery will be put to the test on some interesting bugs... well, bugs as in insects.
#
# As a bug master, you want a classifier capable of classifying different species of bugs: bees, beetles, butterflies, caterpillar, and more. Luckily, you found a great dataset with a lot of animal species and objects, and you trained your classifier on that.
#
# But the bug classes don't do as well as you would like. Now your plan is to train a GAN on the same data so it can generate new bugs to make your classifier better at distinguishing between all of your favorite bugs!
#
# You will fine-tune your model by augmenting the original real data with fake data and during that process, observe how to increase the accuracy of your classifier with these fake, GAN-generated bugs. After this, you will prove your worth as a bug master.
# + [markdown] colab_type="text" id="oSuAJTuYYr2o"
# #### Sampling Ratio
#
# Suppose that you've decided that although you have this pre-trained general generator and this general classifier, capable of identifying 100 classes with some accuracy (~17%), what you'd really like is a model that can classify the five different kinds of bugs in the dataset. You'll fine-tune your model by augmenting your data with the generated images. Keep in mind that both the generator and the classifier were trained on the same images: the 40 images per class you painstakingly found so your generator may not be great. This is the caveat with data augmentation, ultimately you are still bound by the real data that you have but you want to try and create more. To make your models even better, you would need to take some more bug photos, label them, and add them to your training set and/or use higher quality photos.
#
# To start, you'll first need to write some code to sample a combination of real and generated images. Given a probability, `p_real`, you'll need to generate a combined tensor where roughly `p_real` of the returned images are sampled from the real images. Note that you should not interpolate the images here: you should choose each image from the real or fake set with a given probability. For example, if your real images are a tensor of `[[1, 2, 3, 4, 5]]` and your fake images are a tensor of `[[-1, -2, -3, -4, -5]]`, and `p_real = 0.2`, two potential random return values are `[[1, -2, 3, -4, -5]]` or `[[-1, 2, -3, -4, -5]]`.
#
#
# Notice that `p_real = 0.2` does not guarantee that exactly 20% of the samples are real, just that when choosing an image for the combined set, there is a 20% probability that that image will be chosen from the real images, and an 80% probability that it will be selected from the fake images.
#
# In addition, we will expect the images to remain in the same order to maintain their alignment with their labels (this applies to the fake images too!).
#
# <details>
# <summary>
# <font size="3" color="green">
# <b>Optional hints for <code><font size="4">combine_sample</font></code></b>
# </font>
# </summary>
#
# 1. This code probably shouldn't be much longer than 3 lines
# 2. You can index using a set of booleans which have the same length as your tensor
# 3. You want to generate an unbiased sample, which you can do (for example) with `torch.rand(length_reals) > p`.
# 4. There are many approaches here that will give a correct answer here. You may find [`torch.rand`](https://pytorch.org/docs/stable/generated/torch.rand.html) or [`torch.bernoulli`](https://pytorch.org/docs/master/generated/torch.bernoulli.html) useful.
# 5. You don't want to edit an argument in place, so you may find [`cur_tensor.clone()`](https://pytorch.org/docs/stable/tensors.html) useful too, which makes a copy of `cur_tensor`.
#
# </details>
# + colab={} colab_type="code" id="16JJ7RlKxrsY"
# UNQ_C1 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# GRADED FUNCTION: combine_sample
def combine_sample(real, fake, p_real):
'''
Function to take a set of real and fake images of the same length (x)
and produce a combined tensor with length (x) and sampled at the target probability
Parameters:
real: a tensor of real images, length (x)
fake: a tensor of fake images, length (x)
p_real: the probability the images are sampled from the real set
'''
#### START CODE HERE ####
make_fake = torch.rand(len(real)) > p_real
target_images = real.clone()
target_images[make_fake] = fake[make_fake]
#### END CODE HERE ####
return target_images
# + colab={} colab_type="code" id="1kDmOc81zJGN"
n_test_samples = 9999
test_combination = combine_sample(
torch.ones(n_test_samples, 1),
torch.zeros(n_test_samples, 1),
0.3
)
# Check that the shape is right
assert tuple(test_combination.shape) == (n_test_samples, 1)
# Check that the ratio is right
assert torch.abs(test_combination.mean() - 0.3) < 0.05
# Make sure that no mixing happened
assert test_combination.median() < 1e-5
test_combination = combine_sample(
torch.ones(n_test_samples, 10, 10),
torch.zeros(n_test_samples, 10, 10),
0.8
)
# Check that the shape is right
assert tuple(test_combination.shape) == (n_test_samples, 10, 10)
# Make sure that no mixing happened
assert torch.abs((test_combination.sum([1, 2]).median()) - 100) < 1e-5
test_reals = torch.arange(n_test_samples)[:, None].float()
test_fakes = torch.zeros(n_test_samples, 1)
test_saved = (test_reals.clone(), test_fakes.clone())
test_combination = combine_sample(test_reals, test_fakes, 0.3)
# Make sure that the sample isn't biased
assert torch.abs((test_combination.mean() - 1500)) < 100
# Make sure no inputs were changed
assert torch.abs(test_saved[0] - test_reals).sum() < 1e-3
assert torch.abs(test_saved[1] - test_fakes).sum() < 1e-3
test_fakes = torch.arange(n_test_samples)[:, None].float()
test_combination = combine_sample(test_reals, test_fakes, 0.3)
# Make sure that the order is maintained
assert torch.abs(test_combination - test_reals).sum() < 1e-4
if torch.cuda.is_available():
# Check that the solution matches the input device
assert str(combine_sample(
torch.ones(n_test_samples, 10, 10).cuda(),
torch.zeros(n_test_samples, 10, 10).cuda(),
0.8
).device).startswith("cuda")
print("Success!")
# + [markdown] colab_type="text" id="LpMGXMYU1a4O"
# Now you have a challenge: find a `p_real` and a generator image such that your classifier gets an average of a 51% accuracy or higher on the insects, when evaluated with the `eval_augmentation` function. **You'll need to fill in `find_optimal` to find these parameters to solve this part!** Note that if your answer takes a very long time to run, you may need to hard-code the solution it finds.
#
# When you're training a generator, you will often have to look at different checkpoints and choose one that does the best (either empirically or using some evaluation method). Here, you are given four generator checkpoints: `gen_1.pt`, `gen_2.pt`, `gen_3.pt`, `gen_4.pt`. You'll also have some scratch area to write whatever code you'd like to solve this problem, but you must return a `p_real` and an image name of your selected generator checkpoint. You can hard-code/brute-force these numbers if you would like, but you are encouraged to try to solve this problem in a more general way. In practice, you would also want a test set (since it is possible to overfit on a validation set), but for simplicity you can just focus on the validation set.
# + colab={} colab_type="code" id="Fc7mFIVRVT_2"
# UNQ_C2 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# GRADED FUNCTION: find_optimal
def find_optimal():
# In the following section, you can write the code to choose your optimal answer
# You can even use the eval_augmentation function in your code if you'd like!
gen_names = [
"gen_1.pt",
"gen_2.pt",
"gen_3.pt",
"gen_4.pt"
]
#### START CODE HERE ####
best_p_real, best_gen_name = 0, "gen_1.pt"
max_eval = -1
for gen_name in gen_names:
p_real_all = torch.linspace(0, 1, 21)
for p_real in tqdm(p_real_all):
curr_eval = eval_augmentation(p_real, gen_name, n_test=20)
if curr_eval > max_eval:
max_eval = curr_eval
best_p_real = p_real
best_gen_name = gen_name
return best_p_real, best_gen_name
#best_p_real, best_gen_name = None, None
#### END CODE HERE ####
#return best_p_real, best_gen_name
def augmented_train(p_real, gen_name):
gen = Generator(generator_input_dim).to(device)
gen.load_state_dict(torch.load(gen_name))
classifier = Classifier(cifar100_shape[0], n_classes).to(device)
classifier.load_state_dict(torch.load("class.pt"))
criterion = nn.CrossEntropyLoss()
batch_size = 256
train_set = torch.load("insect_train.pt")
val_set = torch.load("insect_val.pt")
dataloader = DataLoader(
torch.utils.data.TensorDataset(train_set["images"], train_set["labels"]),
batch_size=batch_size,
shuffle=True
)
validation_dataloader = DataLoader(
torch.utils.data.TensorDataset(val_set["images"], val_set["labels"]),
batch_size=batch_size
)
display_step = 1
lr = 0.0002
n_epochs = 20
classifier_opt = torch.optim.Adam(classifier.parameters(), lr=lr)
cur_step = 0
best_score = 0
for epoch in range(n_epochs):
for real, labels in dataloader:
real = real.to(device)
# Flatten the image
labels = labels.to(device)
one_hot_labels = get_one_hot_labels(labels.to(device), n_classes).float()
### Update classifier ###
# Get noise corresponding to the current batch_size
classifier_opt.zero_grad()
cur_batch_size = len(labels)
fake_noise = get_noise(cur_batch_size, z_dim, device=device)
noise_and_labels = combine_vectors(fake_noise, one_hot_labels)
fake = gen(noise_and_labels)
target_images = combine_sample(real.clone(), fake.clone(), p_real)
labels_hat = classifier(target_images.detach())
classifier_loss = criterion(labels_hat, labels)
classifier_loss.backward()
classifier_opt.step()
# Calculate the accuracy on the validation set
if cur_step % display_step == 0 and cur_step > 0:
classifier_val_loss = 0
classifier_correct = 0
num_validation = 0
with torch.no_grad():
for val_example, val_label in validation_dataloader:
cur_batch_size = len(val_example)
num_validation += cur_batch_size
val_example = val_example.to(device)
val_label = val_label.to(device)
labels_hat = classifier(val_example)
classifier_val_loss += criterion(labels_hat, val_label) * cur_batch_size
classifier_correct += (labels_hat.argmax(1) == val_label).float().sum()
accuracy = classifier_correct.item() / num_validation
if accuracy > best_score:
best_score = accuracy
cur_step += 1
return best_score
def eval_augmentation(p_real, gen_name, n_test=20):
total = 0
for i in range(n_test):
total += augmented_train(p_real, gen_name)
return total / n_test
best_p_real, best_gen_name = find_optimal()
performance = eval_augmentation(best_p_real, best_gen_name)
print(f"Your model had an accuracy of {performance:0.1%}")
assert performance > 0.512
print("Success!")
# + [markdown] colab_type="text" id="mmqeeBjE32ls"
# You'll likely find that the worst performance is when the generator is performing alone: this corresponds to the case where you might be trying to hide the underlying examples from the classifier. Perhaps you don't want other people to know about your specific bugs!
# + colab={} colab_type="code" id="aLRFjtb_HEuP"
accuracies = []
p_real_all = torch.linspace(0, 1, 21)
for p_real_vis in tqdm(p_real_all):
accuracies += [eval_augmentation(p_real_vis, best_gen_name, n_test=4)]
plt.plot(p_real_all.tolist(), accuracies)
plt.ylabel("Accuracy")
_ = plt.xlabel("Percent Real Images")
# + [markdown] colab_type="text" id="e2j-xodd1ykT"
# Here's a visualization of what the generator is actually generating, with real examples of each class above the corresponding generated image.
# + colab={} colab_type="code" id="HpcnjIK_0WdF"
examples = [4, 41, 80, 122, 160]
train_images = torch.load("insect_train.pt")["images"][examples]
train_labels = torch.load("insect_train.pt")["labels"][examples]
one_hot_labels = get_one_hot_labels(train_labels.to(device), n_classes).float()
fake_noise = get_noise(len(train_images), z_dim, device=device)
noise_and_labels = combine_vectors(fake_noise, one_hot_labels)
gen = Generator(generator_input_dim).to(device)
gen.load_state_dict(torch.load(best_gen_name))
fake = gen(noise_and_labels)
show_tensor_images(torch.cat([train_images.cpu(), fake.cpu()]))
# -
| Apply Generative Adversarial Networks (GANs)/C3W1_Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.01442, "end_time": "2021-05-09T14:47:17.999735", "exception": false, "start_time": "2021-05-09T14:47:17.985315", "status": "completed"} tags=[]
# 
#
# The Herbarium 2021: Half-Earth Challenge is to identify vascular plant specimens provided by the New York Botanical Garden (NY), Bishop Museum (BPBM), Naturalis Biodiversity Center (NL), Queensland Herbarium (BRI), and Auckland War Memorial Museum (AK).
#
# The Herbarium 2021: Half-Earth Challenge dataset includes more than 2.5M images representing nearly 65,000 species from the Americas and Oceania that have been aligned to a standardized plant list (LCVP v1.0.2).
#
# This kernel covers how to train a **EfficientNet** using a TFRecords dataset. The notebook is intended to be used on TPU.
# + [markdown] papermill={"duration": 0.012975, "end_time": "2021-05-09T14:47:18.026392", "exception": false, "start_time": "2021-05-09T14:47:18.013417", "status": "completed"} tags=[]
# <a id = 'basic'></a>
# # Packages 📦 and Basic Setup
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _kg_hide-input=true _kg_hide-output=true _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 20.63138, "end_time": "2021-05-09T14:47:38.670542", "exception": false, "start_time": "2021-05-09T14:47:18.039162", "status": "completed"} tags=[]
# %%capture
# Install Weights and Biases
# !pip3 install wandb --upgrade >> /dev/null
# Packages
import os
import time
import logging
import re, math
import numpy as np
import pandas as pd
import tensorflow as tf
import matplotlib.pyplot as plt
import tensorflow.keras.backend as K
from sklearn.model_selection import KFold
from kaggle_datasets import KaggleDatasets
# Configure Logging Level
logger = tf.get_logger()
logger.setLevel(logging.ERROR)
# Weights and Biases Setup
import wandb
from wandb.keras import WandbCallback
from kaggle_secrets import UserSecretsClient
user_secrets = UserSecretsClient()
api_key = user_secrets.get_secret("WANDB_API_KEY")
wandb.login(key=api_key);
# + [markdown] papermill={"duration": 0.012442, "end_time": "2021-05-09T14:47:38.696231", "exception": false, "start_time": "2021-05-09T14:47:38.683789", "status": "completed"} tags=[]
# ## Basic Hyperparameters 🪡
# + papermill={"duration": 0.37306, "end_time": "2021-05-09T14:47:39.082041", "exception": false, "start_time": "2021-05-09T14:47:38.708981", "status": "completed"} tags=[]
DEVICE = "TPU"
GCS_PATH = KaggleDatasets().get_gcs_path('herb2021-256')
IMG_SIZES = 256
IMAGE_SIZE = [IMG_SIZES, IMG_SIZES]
BATCH_SIZE_SINGLE = 64
EPOCHS = 40
FOLDS = 10
N_CLASSES = 64500
# + [markdown] papermill={"duration": 0.012704, "end_time": "2021-05-09T14:47:39.107859", "exception": false, "start_time": "2021-05-09T14:47:39.095155", "status": "completed"} tags=[]
# ## Device Configuration 🔌
# + _kg_hide-input=true _kg_hide-output=false papermill={"duration": 5.699874, "end_time": "2021-05-09T14:47:44.820841", "exception": false, "start_time": "2021-05-09T14:47:39.120967", "status": "completed"} tags=[]
if DEVICE == "TPU":
print("connecting to TPU...")
try:
tpu = tf.distribute.cluster_resolver.TPUClusterResolver()
print('Running on TPU ', tpu.master())
except ValueError:
print("Could not connect to TPU")
tpu = None
if tpu:
try:
print("initializing TPU ...")
tf.config.experimental_connect_to_cluster(tpu)
tf.tpu.experimental.initialize_tpu_system(tpu)
strategy = tf.distribute.experimental.TPUStrategy(tpu)
print("TPU initialized")
except _:
print("failed to initialize TPU")
else:
DEVICE = "GPU"
if DEVICE == "GPU":
n_gpu = len(tf.config.experimental.list_physical_devices('GPU'))
print("Num GPUs Available: ", n_gpu)
if n_gpu > 1:
print("Using strategy for multiple GPU")
strategy = tf.distribute.MirroredStrategy()
else:
print('Standard strategy for GPU...')
strategy = tf.distribute.get_strategy()
AUTO = tf.data.experimental.AUTOTUNE
REPLICAS = strategy.num_replicas_in_sync
print(f'REPLICAS: {REPLICAS}')
BATCH_SIZE = BATCH_SIZE_SINGLE * REPLICAS
print(f'BATCH_SIZE: {BATCH_SIZE}')
# + [markdown] papermill={"duration": 0.013749, "end_time": "2021-05-09T14:47:44.849823", "exception": false, "start_time": "2021-05-09T14:47:44.836074", "status": "completed"} tags=[]
# <a id = 'data'></a>
# # 💿 Tensorflow Dataset from TFRecords
# + _kg_hide-input=true _kg_hide-output=true papermill={"duration": 0.050858, "end_time": "2021-05-09T14:47:44.914425", "exception": false, "start_time": "2021-05-09T14:47:44.863567", "status": "completed"} tags=[]
data_augmentation = tf.keras.Sequential([
tf.keras.layers.experimental.preprocessing.RandomRotation(0.2, seed=12345),
])
def read_labeled_tfrecord(example):
LABELED_TFREC_FORMAT = {
"image": tf.io.FixedLenFeature([], tf.string),
"image_idx": tf.io.FixedLenFeature([], tf.string),
'label' : tf.io.FixedLenFeature([], tf.int64)
}
example = tf.io.parse_single_example(example, LABELED_TFREC_FORMAT)
image = decode_image(example['image'])
label = example['label']
return image, label
def read_labeled_tfrecord_for_test(example):
LABELED_TFREC_FORMAT = {
"image": tf.io.FixedLenFeature([], tf.string),
"image_idx": tf.io.FixedLenFeature([], tf.string),
'label' : tf.io.FixedLenFeature([], tf.int64)
}
example = tf.io.parse_single_example(example, LABELED_TFREC_FORMAT)
image = decode_image(example['image'])
label = example['label']
return image, label
def decode_image(image_data):
image = tf.image.decode_jpeg(image_data, channels=3)
image = tf.cast(image, tf.float32)
image = tf.reshape(image, [*IMAGE_SIZE, 3])
return image
def count_data_items(filenames):
n = [int(re.compile(r"-([0-9]*)\.").search(filename).group(1))
for filename in filenames]
return np.sum(n)
def load_dataset(filenames, labeled=True, ordered=False, isTest=False):
ignore_order = tf.data.Options()
if not ordered:
ignore_order.experimental_deterministic = False
dataset = tf.data.TFRecordDataset(filenames, num_parallel_reads=AUTO)
dataset = dataset.with_options(ignore_order)
if isTest == False:
dataset = dataset.map(read_labeled_tfrecord)
else:
dataset = dataset.map(read_labeled_tfrecord_for_test)
return dataset
def get_training_dataset(filenames):
dataset = load_dataset(filenames, labeled=True, isTest = False)
dataset = dataset.repeat()
dataset = dataset.shuffle(2048)
dataset = dataset.batch(BATCH_SIZE)
dataset = dataset.prefetch(AUTO)
return dataset
def get_valid_dataset(filenames):
dataset = load_dataset(filenames, labeled=True, isTest = True)
dataset = dataset.batch(BATCH_SIZE)
dataset = dataset.prefetch(AUTO)
return dataset
def get_test_dataset(filenames):
dataset = load_dataset(filenames, labeled=True, isTest = True, ordered=True)
dataset = dataset.batch(BATCH_SIZE)
dataset = dataset.prefetch(AUTO)
return dataset
# + [markdown] papermill={"duration": 0.013402, "end_time": "2021-05-09T14:47:44.941850", "exception": false, "start_time": "2021-05-09T14:47:44.928448", "status": "completed"} tags=[]
# # The Model 👷♀️
# + [markdown] papermill={"duration": 0.01333, "end_time": "2021-05-09T14:47:44.968966", "exception": false, "start_time": "2021-05-09T14:47:44.955636", "status": "completed"} tags=[]
# ## Transfer Learning
#
# The main aim of transfer learning (TL) is to implement a model quickly i.e. instead of creating a DNN (dense neural network) from scratch, the model will transfer the features it has learned from the different dataset that has performed the same task. This transaction is also known as **knowledge transfer**.
#
# ---
#
# ## EfficientNetB4
#
# 
#
# > Excerpt from Google AI Blog
#
# **Convolutional neural networks (CNNs)** are commonly developed at a fixed resource cost, and then scaled up in order to achieve better accuracy when more resources are made available. For example, ResNet can be scaled up from ResNet-18 to ResNet-200 by increasing the number of layers. The conventional practice for model scaling is to arbitrarily increase the CNN depth or width, or to use larger input image resolution for training and evaluation. While these methods do improve accuracy, they usually require tedious manual tuning, and still often yield suboptimal performance. Instead, the authors of [**"EfficientNet: Rethinking Model Scaling for Convolutional Neural Networks (ICML 2019)"**](https://arxiv.org/abs/1905.11946) found a more principled method to scale up a CNN to obtain better accuracy and efficiency.
#
# They proposed a novel model scaling method that uses a simple yet highly effective **compound coefficient** to scale up CNNs in a more structured manner. Unlike conventional approaches that arbitrarily scale network dimensions, such as width, depth and resolution, their method uniformly scales each dimension with a fixed set of scaling coefficients. The resulting models named **EfficientNets**, superpassed state-of-the-art accuracy with up to **10x** better efficiency (**smaller and faster**).
#
# In this project we'll use **`EfficientNetB4`** for training our Classifier. The Model can easily be instantiated using the **`tf.keras.applications`** Module, which provides canned architectures with pre-trained weights. For more details kindly visit [this](https://www.tensorflow.org/api_docs/python/tf/keras/applications) link. Unhide the below cell to see the `build_model()` function
# + _kg_hide-input=true _kg_hide-output=true papermill={"duration": 0.037069, "end_time": "2021-05-09T14:47:45.020126", "exception": false, "start_time": "2021-05-09T14:47:44.983057", "status": "completed"} tags=[]
tpu_data_augmentation = tf.keras.Sequential([
tf.keras.layers.experimental.preprocessing.RandomFlip("horizontal_and_vertical", seed=12345),
])
def build_model(dim = IMG_SIZES, ef = 0):
inp = tf.keras.layers.Input(shape=(*IMAGE_SIZE, 3))
base = tf.keras.applications.EfficientNetB3(include_top=False, weights='imagenet',
input_shape=(*IMAGE_SIZE, 3), pooling='avg')
x = tpu_data_augmentation(inp)
x = base(x)
x = tf.keras.layers.BatchNormalization()(x)
x = tf.keras.layers.Dense(512)(x)
x = tf.keras.layers.BatchNormalization()(x)
x = tf.keras.layers.Activation('relu')(x)
x = tf.keras.layers.Dense(N_CLASSES, activation='softmax')(x)
model = tf.keras.Model(inputs = inp,outputs = x)
opt = tf.keras.optimizers.Adam(learning_rate = 0.001)
fn_loss = tf.keras.losses.SparseCategoricalCrossentropy()
model.compile(optimizer = opt, loss = [fn_loss], metrics=['accuracy'])
return model
# + _kg_hide-input=false papermill={"duration": 5.477577, "end_time": "2021-05-09T14:47:50.512107", "exception": false, "start_time": "2021-05-09T14:47:45.034530", "status": "completed"} tags=[]
display_model = build_model(dim=IMG_SIZES)
display_model.summary()
# + [markdown] papermill={"duration": 0.016355, "end_time": "2021-05-09T14:47:50.545396", "exception": false, "start_time": "2021-05-09T14:47:50.529041", "status": "completed"} tags=[]
# <a id = 'train'></a>
# # Training 💪🏻
# + [markdown] papermill={"duration": 0.016512, "end_time": "2021-05-09T14:47:50.578656", "exception": false, "start_time": "2021-05-09T14:47:50.562144", "status": "completed"} tags=[]
# ## LearningRate Scheduler
#
# > From a [TowardsDataScience article](https://towardsdatascience.com/learning-rate-scheduler-d8a55747dd90)
#
# In training deep networks, it is helpful to reduce the learning rate as the number of training epochs increases. This is **based on the intuition** that with a high learning rate, the deep learning model would possess high kinetic energy. As a result, it’s parameter vector bounces around chaotically. Thus, it’s unable to settle down into deeper and narrower parts of the loss function (local minima). If the learning rate, on the other hand, was very small, the system then would have low kinetic energy. Thus, it would settle down into shallow and narrower parts of the loss function (false minima).
#
# <center> <img src = "https://miro.medium.com/max/668/1*iYWyu8hemMyaBlK6V-2vqg.png"> </center>
#
# The above figure depicts that a high learning rate will lead to random to and fro moment of the vector around local minima while a slow learning rate results in getting stuck into false minima. Thus, knowing when to decay the learning rate can be hard to find out.
#
# Decreasing the learning rate during training can lead to improved accuracy and (most perplexingly) reduced overfitting of the model. A piecewise decrease of the learning rate whenever progress has plateaued is effective in practice. Essentially this ensures that we converge efficiently to a suitable solution and only then reduce the inherent variance of the parameters by reducing the learning rate.
#
# Here, we'll demonstrate how to use LearningRate schedules to automatically **adapt learning rates** that achieve the **optimal rate of convergence** for stochastic gradient descent. Unhide the cell to see the custom callback.
# + _kg_hide-input=true _kg_hide-output=true papermill={"duration": 0.027147, "end_time": "2021-05-09T14:47:50.622312", "exception": false, "start_time": "2021-05-09T14:47:50.595165", "status": "completed"} tags=[]
def get_lr_callback(batch_size=8):
lr_start = 0.0002
lr_max = 0.0002 * 10
lr_min = lr_start/2
lr_ramp_ep = 6
lr_sus_ep = 10
lr_decay = 0.8
def lrfn(epoch):
if epoch < lr_ramp_ep:
lr = (lr_max - lr_start) / lr_ramp_ep * epoch + lr_start
elif epoch < lr_ramp_ep + lr_sus_ep:
lr = lr_max
else:
lr = (lr_max - lr_min) * lr_decay**(epoch - lr_ramp_ep - lr_sus_ep) + lr_min
return lr
lr_callback = tf.keras.callbacks.LearningRateScheduler(lrfn, verbose=True)
return lr_callback
# + papermill={"duration": 0.112837, "end_time": "2021-05-09T14:47:50.752094", "exception": false, "start_time": "2021-05-09T14:47:50.639257", "status": "completed"} tags=[]
all_files = tf.io.gfile.glob(GCS_PATH + '/train*.tfrec')
num_total_files = len(all_files)
n_images = count_data_items(all_files)
print('Total number of files for train-validation:', num_total_files)
print('Total number of image for train-validation:', n_images)
# + _kg_hide-input=true _kg_hide-output=true papermill={"duration": 0.032552, "end_time": "2021-05-09T14:47:50.801959", "exception": false, "start_time": "2021-05-09T14:47:50.769407", "status": "completed"} tags=[]
def train_one_fold(fold, files_train, files_valid):
VERBOSE = 1
tStart = time.time()
# Better Performance
if DEVICE=='TPU':
tf.tpu.experimental.initialize_tpu_system(tpu)
# Build the Model
K.clear_session()
with strategy.scope():
print('Building model...')
model = build_model(dim=IMG_SIZES)
# Callback to Save Model
sv = tf.keras.callbacks.ModelCheckpoint('fold-%i.h5'%fold, monitor='val_loss', verbose=1, save_best_only=True,
save_weights_only=True, mode='min', save_freq='epoch')
# Train for One Fold
history = model.fit(get_training_dataset(files_train),
epochs=EPOCHS,
callbacks = [sv, get_lr_callback(BATCH_SIZE), WandbCallback()],
steps_per_epoch = count_data_items(files_train)/BATCH_SIZE//REPLICAS,
validation_data = get_valid_dataset(files_valid),
validation_steps = count_data_items(files_valid)/BATCH_SIZE//REPLICAS,
verbose=VERBOSE)
model.save('b3-aug.h5')
#save it as model artifact on W&B
artifact = wandb.Artifact(name="b3-aug", type="weights")
artifact.add_file('b3-aug.h5')
wandb.log_artifact(artifact)
# Record the Time Spent
tElapsed = round(time.time() - tStart, 1)
print(' ')
print('Time (sec) elapsed: ', tElapsed)
print('...')
print('...')
return history
# + _kg_hide-input=false _kg_hide-output=true papermill={"duration": 9986.558595, "end_time": "2021-05-09T17:34:17.378511", "exception": false, "start_time": "2021-05-09T14:47:50.819916", "status": "completed"} tags=[]
SHOW_FILES = True
STOP_FOLDS = 0
skf = KFold(n_splits = FOLDS, shuffle = True, random_state=54321)
histories = []
for fold,(idxT,idxV) in enumerate(skf.split(np.arange(num_total_files))):
print('')
print('#'*60)
print('#### FOLD', fold+1)
print('#### Epochs: %i' %(EPOCHS))
print('#'*60)
train_files = tf.io.gfile.glob([GCS_PATH + '/train%.3i*.tfrec'%x for x in idxT])
valid_files = tf.io.gfile.glob([GCS_PATH + '/train%.3i*.tfrec'%x for x in idxV])
if SHOW_FILES:
print('Number of training images', count_data_items(train_files))
print('Number of validation images', count_data_items(valid_files))
run = wandb.init(project='Herbarium 2021', entity='sauravmaheshkar', reinit=True)
history = train_one_fold(fold+1, train_files, valid_files)
run.finish()
histories.append(history)
if fold >= STOP_FOLDS:
break
| notebooks/EfficientNet/herbarium2021-tensorflow-weights-biases.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import mxnet as mx
import numpy as np
import random
import bisect
# set up logging
import logging
reload(logging)
logging.basicConfig(format='%(asctime)s %(levelname)s:%(message)s', level=logging.DEBUG, datefmt='%I:%M:%S')
# # A Glance of LSTM structure and embedding layer
#
# We will build a LSTM network to learn from char only. At each time, input is a char. We will see this LSTM is able to learn words and grammers from sequence of chars.
#
# The following figure is showing an unrolled LSTM network, and how we generate embedding of a char. The one-hot to embedding operation is a special case of fully connected network.
#
# <img src="http://webdocs.cs.ualberta.ca/~bx3/char-rnn_1.png">
#
# <img src="http://webdocs.cs.ualberta.ca/~bx3/char-rnn_2.png">
from lstm import lstm_unroll, lstm_inference_symbol
from bucket_io import BucketSentenceIter
from rnn_model import LSTMInferenceModel
# +
# Read from doc
def read_content(path):
with open(path) as ins:
content = ins.read()
return content
# Build a vocabulary of what char we have in the content
def build_vocab(path):
content = read_content(path)
content = list(content)
idx = 1 # 0 is left for zero-padding
the_vocab = {}
for word in content:
if len(word) == 0:
continue
if not word in the_vocab:
the_vocab[word] = idx
idx += 1
return the_vocab
# We will assign each char with a special numerical id
def text2id(sentence, the_vocab):
words = list(sentence)
words = [the_vocab[w] for w in words if len(w) > 0]
return words
# -
# Evaluation
def Perplexity(label, pred):
loss = 0.
for i in range(pred.shape[0]):
loss += -np.log(max(1e-10, pred[i][int(label[i])]))
return np.exp(loss / label.size)
# # Get Data
import os
data_url = "http://webdocs.cs.ualberta.ca/~bx3/lab_data.zip"
os.system("wget %s" % data_url)
os.system("unzip -o lab_data.zip")
# Sample training data:
# ```
# all to Renewal Keynote Address Call to Renewal Pt 1Call to Renewal Part 2 TOPIC: Our Past, Our Future & Vision for America June
# 28, 2006 Call to Renewal' Keynote Address Complete Text Good morning. I appreciate the opportunity to speak here at the Call to R
# enewal's Building a Covenant for a New America conference. I've had the opportunity to take a look at your Covenant for a New Ame
# rica. It is filled with outstanding policies and prescriptions for much of what ails this country. So I'd like to congratulate yo
# u all on the thoughtful presentations you've given so far about poverty and justice in America, and for putting fire under the fe
# et of the political leadership here in Washington.But today I'd like to talk about the connection between religion and politics a
# nd perhaps offer some thoughts about how we can sort through some of the often bitter arguments that we've been seeing over the l
# ast several years.I do so because, as you all know, we can affirm the importance of poverty in the Bible; and we can raise up and
# pass out this Covenant for a New America. We can talk to the press, and we can discuss the religious call to address poverty and
# environmental stewardship all we want, but it won't have an impact unless we tackle head-on the mutual suspicion that sometimes
# ```
# # LSTM Hyperparameters
# The batch size for training
batch_size = 32
# We can support various length input
# For this problem, we cut each input sentence to length of 129
# So we only need fix length bucket
buckets = [129]
# hidden unit in LSTM cell
num_hidden = 512
# embedding dimension, which is, map a char to a 256 dim vector
num_embed = 256
# number of lstm layer
num_lstm_layer = 3
# we will show a quick demo in 2 epoch
# and we will see result by training 75 epoch
num_epoch = 2
# learning rate
learning_rate = 0.01
# we will use pure sgd without momentum
momentum = 0.0
# we can select multi-gpu for training
# for this demo we only use one
devs = [mx.context.gpu(i) for i in range(1)]
# build char vocabluary from input
vocab = build_vocab("./obama.txt")
# generate symbol for a length
def sym_gen(seq_len):
return lstm_unroll(num_lstm_layer, seq_len, len(vocab) + 1,
num_hidden=num_hidden, num_embed=num_embed,
num_label=len(vocab) + 1, dropout=0.2)
# initalize states for LSTM
init_c = [('l%d_init_c'%l, (batch_size, num_hidden)) for l in range(num_lstm_layer)]
init_h = [('l%d_init_h'%l, (batch_size, num_hidden)) for l in range(num_lstm_layer)]
init_states = init_c + init_h
# we can build an iterator for text
data_train = BucketSentenceIter("./obama.txt", vocab, buckets, batch_size,
init_states, seperate_char='\n',
text2id=text2id, read_content=read_content)
# the network symbol
symbol = sym_gen(buckets[0])
# # Train model
# Train a LSTM network as simple as feedforward network
model = mx.model.FeedForward(ctx=devs,
symbol=symbol,
num_epoch=num_epoch,
learning_rate=learning_rate,
momentum=momentum,
wd=0.0001,
initializer=mx.init.Xavier(factor_type="in", magnitude=2.34))
# Fit it
model.fit(X=data_train,
eval_metric = mx.metric.np(Perplexity),
batch_end_callback=mx.callback.Speedometer(batch_size, 50),
epoch_end_callback=mx.callback.do_checkpoint("obama"))
# # Inference from model
# helper strcuture for prediction
def MakeRevertVocab(vocab):
dic = {}
for k, v in vocab.items():
dic[v] = k
return dic
# make input from char
def MakeInput(char, vocab, arr):
idx = vocab[char]
tmp = np.zeros((1,))
tmp[0] = idx
arr[:] = tmp
# +
# helper function for random sample
def _cdf(weights):
total = sum(weights)
result = []
cumsum = 0
for w in weights:
cumsum += w
result.append(cumsum / total)
return result
def _choice(population, weights):
assert len(population) == len(weights)
cdf_vals = _cdf(weights)
x = random.random()
idx = bisect.bisect(cdf_vals, x)
return population[idx]
# we can use random output or fixed output by choosing largest probability
def MakeOutput(prob, vocab, sample=False, temperature=1.):
if sample == False:
idx = np.argmax(prob, axis=1)[0]
else:
fix_dict = [""] + [vocab[i] for i in range(1, len(vocab) + 1)]
scale_prob = np.clip(prob, 1e-6, 1 - 1e-6)
rescale = np.exp(np.log(scale_prob) / temperature)
rescale[:] /= rescale.sum()
return _choice(fix_dict, rescale[0, :])
try:
char = vocab[idx]
except:
char = ''
return char
# -
# load from check-point
_, arg_params, __ = mx.model.load_checkpoint("obama", 75)
# build an inference model
model = LSTMInferenceModel(num_lstm_layer, len(vocab) + 1,
num_hidden=num_hidden, num_embed=num_embed,
num_label=len(vocab) + 1, arg_params=arg_params, ctx=mx.gpu(), dropout=0.2)
# +
# generate a sequence of 1200 chars
seq_length = 1200
input_ndarray = mx.nd.zeros((1,))
revert_vocab = MakeRevertVocab(vocab)
# Feel free to change the starter sentence
output ='The joke'
random_sample = True
new_sentence = True
ignore_length = len(output)
for i in range(seq_length):
if i <= ignore_length - 1:
MakeInput(output[i], vocab, input_ndarray)
else:
MakeInput(output[-1], vocab, input_ndarray)
prob = model.forward(input_ndarray, new_sentence)
new_sentence = False
next_char = MakeOutput(prob, revert_vocab, random_sample)
if next_char == '':
new_sentence = True
if i >= ignore_length - 1:
output += next_char
# -
# Let's see what we can learned from char in Obama's speech.
print(output)
| example/rnn/char-rnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Introduction to NumPy
# by <NAME>
#
# Part of the Quantopian Lecture Series:
#
# * [www.quantopian.com/lectures](https://www.quantopian.com/lectures)
# * [github.com/quantopian/research_public](https://github.com/quantopian/research_public)
#
# Notebook released under the Creative Commons Attribution 4.0 License.
# NumPy is an incredibly powerful package in Python that is ubiquitous throughout the Quantopian platform. It has strong integration with Pandas, another tool we will be covering in the lecture series. NumPy adds support for multi-dimensional arrays and mathematical functions that allow you to easily perform linear algebra calculations. This lecture will be a collection of linear algebra examples computed using NumPy.
import numpy as np
import matplotlib.pyplot as plt
# ### Basic NumPy arrays
# The most basic way that we could make use of NumPy in finance is calculation the mean return of a portfolio. Say that we have a list containing the historical return of several stocks.
stock_list = [3.5, 5, 2, 8, 4.2]
# We can make an array by calling a function on the list:
returns = np.array(stock_list)
print returns, type(returns)
# You'll notice that the type of our array is 'ndarray', not just 'array'. This is because NumPy arrays can be created with multiple dimensions. If we pass np.array() a list of lists, it will create a 2-dimensional array. If pass a list of lists of lists, it will create a 3-dimensional array, and so on and so forth.
A = np.array([[1, 2], [3, 4]])
print A, type(A)
# We can access the dimensions of an array by looking at its `shape` member variable.
print A.shape
# Arrays are indexed in much the same way as lists in Python. Elements of a list begin indexing from $0$ and end at $n - 1$, where $n$ is the length of the array.
print returns[0], returns[len(returns) - 1]
# We can take a slice of an array using a colon, just like in a list.
print returns[1:3]
# A slice of an array, like in a list, will select a group of elements in the array starting from the first element indicated and going up to (but not including) the last element indicated.
#
# In the case of multidimensional arrays, many of the same conventions with slicing and indexing hold. We can access the first column of a 2-dimensional array like so:
print A[:, 0]
# And the first row of a 2-dimensional array like so:
print A[0, :]
# Notice that each slice of the array returns yet another array!
print type(A[0,:])
# Passing only one index to a 2-dimensional array will result in returning the row with the given index as well, providing us with another way to access individual rows.
print A[0]
# Accessing the index of an individual element will return only the element.
print A[1, 1]
# #### Array functions
#
# Functions built into NumPy can be easily called on arrays. Most functions are applied to an array element-wise (as scalar multiplication is). For example, if we call `log()` on an array, the logarithm will be taken of each element.
print np.log(returns)
# Some functions return a single value. This is because they treat the array as a collection (similar to a list), performing the designated function. For example, the `mean()` function will do exactly what you expect, calculating the mean of an array.
print np.mean(returns)
# Or the `max()` function will return the maximum element of an array.
print np.max(returns)
# For further reading on the universal functions in NumPy, check out the [documentation](https://docs.scipy.org/doc/numpy/user/quickstart.html#universal-functions).
# ### Return to the returns
#
# Now let's modify our returns array with scalar values. If we add a scalar value to an array it will be added to every element of the array. If we multiply an array by a scalar value it will be multiplied against every element of the array. If we do both, both will happen!
returns*2 + 5
# NumPy also has functions specifically built to operate on arrays. Let's take the mean and standard deviation of this group of returns.
print "Mean: ", np.mean(returns), "Std Dev: ", np.std(returns)
# Let's simulate a universe of stocks using NumPy's functions. First we need to create the arrays to hold the assets and returns that we will use to build a portfolio. This is because arrays are created with a fixed size. Their dimensions can't be changed without creating a new array.
N = 10
assets = np.zeros((N, 100))
returns = np.zeros((N, 100))
# This function, `zeroes()`, creates a NumPy array with the given dimensions that is entirely filled in with $0$. We can pass a single value or a tuple of as many dimensions as we like. Passing in the tuple `(N, 100)`, will return a two-dimensional array with $N$ rows and $100$ columns. Our result is a $N \times 100$ array.
#
# Now we will simulate a base asset. We want the universe of stocks to be correlated with each other so we will use this initial value to generate the others.
R_1 = np.random.normal(1.01, 0.03, 100)
returns[0] = R_1
assets[0] = np.cumprod(R_1)
# The `random` module in NumPy is exceedingly useful. It contains methods for sampling from many different probability distributions, some of which are covered in the [random variables lecture](https://www.quantopian.com/lectures/random-variables) in the Quantopian lecture series. In this case we draw $N = 100$ random samples from a normal distribution with mean $1.01$ and standard deviation $0.03$. We treat these as the daily percentage returns of our asset and take the cumulative product of these samples to get the current price.
#
# The way we have generated our universe, the the individual $R_i$ vectors are each 1-dimensional arrays and the `returns` and `assets` variables contain 2-dimensional arrays. Above, we set the initial row of both `returns` and `assets` to be the first $R_i$ vector and the cumulative asset price based on those returns, respectively.
#
# We will now use this base asset to create a few other random assets that are correlated with it.
# +
# Generate assets that are correlated with R_1
for i in range(1, N):
R_i = R_1 + np.random.normal(0.001, 0.02, 100)
returns[i] = R_i # Set each row of returns equal to the new R_i array
assets[i] = np.cumprod(R_i)
mean_returns = [(np.mean(R) - 1)*100 for R in returns]
return_volatilities = [np.std(R) for R in returns]
# -
# Here we generate the remaining $N - 1$ securities that we want in our universe by adding random noise to $R_1$. This ensures that our $N - 1$ other assets will be correlated with the base asset because they have some underlying information that is shared.
#
# Let's plot what the mean return of each asset looks like:
plt.bar(np.arange(len(mean_returns)), mean_returns)
plt.xlabel('Stock')
plt.ylabel('Returns')
plt.title('Returns for {0} Random Assets'.format(N));
# ### Calculating Expected Return
#
# So we have a universe of stocks. Great! Now let's put them together in a portfolio and calculate its expected return and risk.
#
# We will start off by generating $N$ random weights for each asset in our portfolio.
weights = np.random.uniform(0, 1, N)
weights = weights/np.sum(weights)
# We have to rescale the weights so that they all add up to $1$. We do this by scaling the weights vector by the sum total of all the weights. This step ensures that we will be using $100\%$ of the portfolio's cash.
#
# To calculate the mean return of the portfolio, we have to scale each asset's return by its designated weight. We can pull each element of each array and multiply them individually, but it's quicker to use NumPy's linear algebra methods. The function that we want is `dot()`. This will calculate the dot product between two arrays for us. So if $v = \left[ 1, 2, 3 \right]$ and $w = \left[4, 5, 6 \right]$, then:
#
# $$ v \cdot w = 1 \times 4 + 2 \times 5 + 3 \times 6 $$
#
# For a one-dimensional vector, the dot product will multiply each element pointwise and add all the products together! In our case, we have a vector of weights, $\omega = \left[ \omega_1, \omega_2, \dots \omega_N\right]$ and a vector of returns, $\mu = \left[ \mu_1, \mu_2, \dots, \mu_N\right]$. If we take the dot product of these two we will get:
#
# $$ \omega \cdot \mu = \omega_1\mu_1 + \omega_2\mu_2 + \dots + \omega_N\mu_N = \mu_P $$
#
# This yields the sum of all the asset returns scaled by their respective weights. This the the portfolio's overall expected return!
p_returns = np.dot(weights, mean_returns)
print "Expected return of the portfolio: ", p_returns
# Calculating the mean return is fairly intuitive and does not require too much explanation of linear algebra. However, calculating the variance of our portfolio requires a bit more background.
# #### Beware of NaN values
#
# Most of the time, all of these calculations will work without an issue. However, when working with real data we run the risk of having `nan` values in our arrays. This is NumPy's way of saying that the data there is missing or doesn't exist. These `nan` values can lead to errors in mathematical calculations so it is important to be aware of whether your array contains `nan` values and to know how to drop them.
v = np.array([1, 2, np.nan, 4, 5])
print v
# Let's see what happens when we try to take the mean of this array.
print np.mean(v)
# Clearly, `nan` values can have a large impact on our calculations. Fortunately, we can check for `nan` values with the `isnan()` function.
np.isnan(v)
# Calling `isnan()` on an array will call the function on each value of the array, returning a value of `True` if the element is `nan` and `False` if the element is valid. Now, knowing whether your array contains `nan` values is all well and good, but how do we remove `nan`s? Handily enough, NumPy arrays can be indexed by boolean values (`True` or `False`). If we use a boolean array to index an array, we will remove all values of the array that register as `False` under the condition. We use the `isnan()` function in create a boolean array, assigning a `True` value to everything that is *not* `nan` and a `False` to the `nan`s and we use that to index the same array.
ix = ~np.isnan(v) # the ~ indicates a logical not, inverting the bools
print v[ix] # We can also just write v = v[~np.isnan(v)]
print np.mean(v[ix])
# There are a few shortcuts to this process in the form of NumPy functions specifically built to handle them, such as `nanmean()`.
print np.nanmean(v)
# The `nanmean()` function simply calculates the mean of the array as if there were no `nan` values at all! There are a few more of these functions, so feel free to read more about them in the [documentation](https://docs.scipy.org/doc/numpy/user/index.html). These indeterminate values are more an issue with data than linear algebra itself so it is helpful that there are ways to handle them.
# ### Conclusion
#
# Linear algebra is pervasive in finance and in general. For example, the calculation of *optimal* weights according to modern portfolio theory is done using linear algebra techniques. The arrays and functions in NumPy allow us to handle these calculations in an intuitive way. For a quick intro to linear algebra and how to use NumPy to do more significant matrix calculations, proceed to the next section.
# ## A brief foray into linear algebra
#
# Let's start with a basic overview of some linear algebra. Linear algebra comes down to the mutiplication and composition of scalar and matrix values. A scalar value is just a real number that we multiply against an array. When we scale a matrix or array using a scalar, we multiply each individual element of that matrix or array by the scalar.
#
# A matrix is a collection of values, typically represented by an $m \times n$ grid, where $m$ is the number of rows and $n$ is the number of columns. The edge lengths $m$ and $n$ do not necessarily have to be different. If we have $m = n$, we call this a square matrix. A particularly interesting case of a matrix is when $m = 1$ or $n = 1$. In this case we have a special case of a matrix that we call a vector. While there is a matrix object in NumPy we will be doing everything using NumPy arrays because they can have dimensions greater than $2$. For the purpose of this section, we will be using matrix and array interchangeably.
#
# We can express the matrix equation as:
#
# $$ y = A\cdot x $$
#
# Where $A$ is an $m \times n$ matrix, $y$ is a $m \times 1$ vector, and $x$ is a $n \times 1$ vector. On the right-hand side of the equation we are multiplying a matrix by a vector. This requires a little bit more clarification, lest we think that we can go about multiplying any matrices by any other matrices.
#
# #### Matrix multiplication
#
# With matrix multiplication, the order in which the matrices are multiplied matters. Multiplying a matrix on the left side by another matrix may be just fine, but multiplying on the right may be undefined.
A = np.array([
[1, 2, 3, 12, 6],
[4, 5, 6, 15, 20],
[7, 8, 9, 10, 10]
])
B = np.array([
[4, 4, 2],
[2, 3, 1],
[6, 5, 8],
[9, 9, 9]
])
# Notice that the above-defined matrices, $A$ and $B$, have different dimensions. $A$ is $3 \times 5$ and $B$ is $4 \times 3$. The general rule of what can and cannot be multiplied in which order is based on the dimensions of the matrices. Specifically, the number of columns in the matrix on the left must be equal to the number of rows in the matrix on the right. In super informal terms, let's say that we have an $m \times n$ matrix and a $p \times q$ matrix. If we multiply the first by the second on the right, we get the following:
#
# $$ (m \times n) \cdot (p \times q) = (m \times q) $$
#
# So the resultant product has the same number of rows as the left matrix and the same number of columns as the right matrix. This limitation of matrix multiplication with regards to dimensions is important to keep track of when writing code. To demonstrate this, we use the `dot()` function to multiply our matrices below:
print np.dot(A, B)
# These results make sense in accordance with our rule. Multiplying a $3 \times 5$ matrix on the right by a $4 \times 3$ matrix results in an error while multiplying a $4 \times 3$ matrix on the right by a $3 \times 5$ matrix results in a $4 \times 5$ matrix.
print np.dot(B, A)
# ### Portfolio Variance
#
# Let's return to our portfolio example from before. We calculated the expected return of the portfolio, but how do we calculate the variance We start by trying to evaluate the portfolio as a sum of each individual asset, scaled by it's weight.
#
# $$ VAR[P] = VAR[\omega_1 S_1 + \omega_2 S_2 + \cdots + \omega_N S_N] $$
#
# Where $S_0, \cdots, S_N$ are the assets contained within our universe. If all of our assets were independent of each other, we could simply evaluate this as
#
# $$ VAR[P] = VAR[\omega_1 S_1] + VAR[\omega_2 S_2] + \cdots + VAR[\omega_N S_N] = \omega_1^2\sigma_1^2 + \omega_2^2\sigma_2^2 + \cdots + \omega_N^2\sigma_N^2 $$
#
# However, all of our assets depend on each other by their construction. They are all in some way related to our base asset and therefore each other. We thus have to calculate the variance of the portfolio by including the individual pairwise covariances of each asset. Our formula for the variance of the portfolio:
#
# $$ VAR[P] = \sigma_P^2 = \sum_i \omega_i^2\sigma_i^2 + \sum_i\sum_{i\neq j} \omega_i\omega_j\sigma_i\sigma_j\rho_{i, j}, \ i, j \in \lbrace 1, 2, \cdots, N \rbrace $$
#
# Where $\rho_{i,j}$ is the correlation between $S_i$ and $S_j$, $\rho_{i, j} = \frac{COV[S_i, S_j]}{\sigma_i\sigma_j}$. This seems exceedingly complicated, but we can easily handle all of this using NumPy arrays. First, we calculate the covariance matrix that relates all the individual stocks in our universe.
cov_mat = np.cov(returns)
print cov_mat
# This array is not formatted particularly nicely, but a covariance matrix is a very important concept. The covariance matrix is of the form:
#
# $$ \left[\begin{matrix}
# VAR[S_1] & COV[S_1, S_2] & \cdots & COV[S_1, S_N] \\
# COV[S_2, S_1] & VAR[S_2] & \cdots & COV[S_2, S_N] \\
# \vdots & \vdots & \ddots & \vdots \\
# COV[S_N, S_1] & COV[S_N, S_2] & \cdots & VAR[S_N]
# \end{matrix}\right] $$
#
# So each diagonal entry is the variance of that asset at that index and each off-diagonal holds the covariance of two assets indexed by the column and row number. What is important is that once we have the covariance matrix we are able to do some very quick linear algebra to calculate the variance of the overall portfolio. We can represent the variance of the portfolio in array form as:
#
# $$ \sigma_p^2 = \omega \ C \ \omega^\intercal$$
#
# Where $C$ is the covariance matrix of all the assets and $\omega$ is the array containing the weights of each individual asset. The superscript $\intercal$ on the second $\omega$ listed above denotes the **transpose** of $\omega$. For a reference on the evaluation of the variance of a portfolio as a matrix equation, please see the Wikipedia article on [modern portfolio theory](https://en.wikipedia.org/wiki/Modern_portfolio_theory).
#
# The transpose of an array is what you get when you switch the rows and columns of an array. This has the effect of reflecting an array across what you might imagine as a diagonal. For example, take our array $A$ from before:
print A
# The transpose looks like a mirror image of the same array.
print np.transpose(A)
# But $\omega$ here is a 1-dimensional array, a vector! It makes perfect to take the transpose of $A$, a $3 \times 5$ array, as the output will be a $5 \times 3$ array, but a 1-dimensional array is not quite as intuitive. A typical 1-dimensional array can be thought of as a $1 \times n$ horizontal vector. Thus, taking the tranpose of this array essentially means changing it into a $n \times 1$ vertical vector. This makes sense because 1-dimensional arrays are still arrays and any multiplication done between 1-dimensional and higher dimensional arrays must keep in line with our dimensionality issue of matrix multiplication.
#
# To make a long story short, we think of $\omega$ as $1 \times N$ since we have $N$ securities. This makes it so that $\omega^\intercal$ is $N \times 1$. Again, our covariance matrix is $N \times N$. So the overall multiplication works out like so, in informal terms:
#
# $$ \text{Dimensions}(\sigma_p^2) = \text{Dimensions}(\omega C \omega^\intercal) = (1 \times N)\cdot (N \times N)\cdot (N \times 1) = (1 \times 1)$$
#
# Multiplying the covariance matrix on the left by the plain horizontal vector and on the right by that vector's transpose results in the calculation of a single scalar ($1 \times 1$) value, our portfolio's variance.
#
# So knowing this, let's proceed and calculate the portfolio variance! We can easily calculate the product of these arrays by using `dot()` for matrix multiplication, though this time we have to do it twice.
# Calculating the portfolio volatility
var_p = np.dot(np.dot(weights, cov_mat), weights.T)
vol_p = np.sqrt(var_p)
print "Portfolio volatility: ", vol_p
# To confirm this calculation, let's simply evaluate the volatility of the portfolio using only NumPy functions.
# Confirming calculation
vol_p_alt = np.sqrt(np.var(np.dot(weights, returns), ddof=1))
print "Portfolio volatility: ", vol_p_alt
# The `ddof` parameter is a simple integer input that tells the function the number of degrees of freedom to take into account. This is a more statistical concept, but what this tells us that our matrix calculation is correct!
#
# A lot of this might not make sense at first glance. It helps to go back and forth between the theory and the code representations until you have a better grasp of the mathematics involved. It is definitely not necessary to be an expert on linear algebra and on matrix operations, but linear algebra can help to streamline the process of working with large amounts of data. For further reading on NumPy, check out the [documentation](https://docs.scipy.org/doc/numpy/user/index.html).
# *This presentation is for informational purposes only and does not constitute an offer to sell, a solicitation to buy, or a recommendation for any security; nor does it constitute an offer to provide investment advisory or other services by Quantopian, Inc. ("Quantopian"). Nothing contained herein constitutes investment advice or offers any opinion with respect to the suitability of any security, and any views expressed herein should not be taken as advice to buy, sell, or hold any security or as an endorsement of any security or company. In preparing the information contained herein, Quantopian, Inc. has not taken into account the investment needs, objectives, and financial circumstances of any particular investor. Any views expressed and data illustrated herein were prepared based upon information, believed to be reliable, available to Quantopian, Inc. at the time of publication. Quantopian makes no guarantees as to their accuracy or completeness. All information is subject to change and may quickly become unreliable for various reasons, including changes in market conditions or economic circumstances.*
| docs/memo/notebooks/lectures/Introduction_to_NumPy/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/NoCodeProgram/CodingTest/blob/main/mathBit/gcd.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="lM7G97XYQKzg"
# Title : Greatest Common Divisor
#
# Chapter : Math, Bit
#
# Link :
#
# ChapterLink :
#
# 문제: 주어진 두 숫자의 최대공약수를 구하여라
# + colab={"base_uri": "https://localhost:8080/"} id="Sa04gI_aQDqS" outputId="b9e7194b-f5ad-422f-a26b-f28e282ba946"
#Brute Force
def gcdBruteForce(m: int, n: int) -> int:
def getDivisors(n: int):
divisors = []
for num in range(1,n+1):
if n%num == 0:
divisors.append(num)
return divisors
m_d = getDivisors(m)
n_d = getDivisors(n)
idx_m = 0
idx_n = 0
gcd = 1
while True:
if len(m_d) <=idx_m or len(n_d) <= idx_n:
break
if m_d[idx_m] == n_d[idx_n]:
gcd = m_d[idx_m]
idx_m += 1
idx_n += 1
continue
elif m_d[idx_m] < n_d[idx_n]:
idx_m += 1
continue
else:
idx_n += 1
continue
return gcd
gcdBruteForce(24,54)
# + colab={"base_uri": "https://localhost:8080/"} id="_B5yCsntRCM7" outputId="1fd5760e-8a65-4251-b560-c241b265087b"
#Euclidean Algorithm
def gcdEuclidean(a,b):
if (b == 0):
return a
return gcdEuclidean(b, a%b)
print(gcdEuclidean(24,54))
| mathBit/gcd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# argv:
# - /opt/anaconda3/bin/python3
# - -m
# - ipykernel_launcher
# - -f
# - '{connection_file}'
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nteract={"transient": {"deleting": false}}
# # Ciclos
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
import math
def signo(x):
return x and (1-2*(x<0))
# + [markdown] nteract={"transient": {"deleting": false}}
# ## Problemas Centrales
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
A = [[4. , 3.8, 4. , 4.2, 4. , 3.8, 3.9, 4.1, 4.1],
[4.4, 4.1, 3.8, 3.9, 4.1, 4.1, 4. , 4.1, 4.2],
[4.4, 4.6, 4.2, 3.8, 3.9, 4.1, 4. , 4.2, 4.5],
[4.1, 4.3, 4.4, 4.1, 3.8, 3.9, 4.1, 4.1, 4.5],
[4.1, 4.2, 4.2, 4.2, 4.1, 4.1, 4.3, 4.4, 4.6],
[4.3, 4.4, 4.3, 4.3, 4.2, 4.1, 4.1, 4.5, 4.9],
[4.4, 4.2, 4.1, 4.1, 3.8, 3.7, 4. , 4.3, 5.2],
[4.5, 4.7, 4.5, 4.2, 4.2, 3.8, 4. , 4.5, 5.1],
[4.3, 4.4, 4.5, 4.5, 4.4, 4.2, 4. , 4.5, 5.3]]
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
import matplotlib.pyplot as plt
plt.imshow(A, cmap=plt.cm.gray)
plt.axis('off')
# + [markdown] nteract={"transient": {"deleting": false}}
# ## Ciclos *for*
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
secuenciaADN = "ACCTTTGTTAACCACACTAG"
frecuencia = [0,0,0,0] # inicialización de la lista de
# frecuencias
for i in secuenciaADN: # se itera sobre la secuencia de ADN
if i == "A": # determinar el tipo de nucleótido
frecuencia[0] += 1
elif i == "C":
frecuencia[1] += 1
elif i == "G":
frecuencia[2] += 1
else:
frecuencia[3] += 1
print(frecuencia)
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
def ConteoNucleotidos(S, nucleotidos = 'ACGT'):
Frecuencias = []
S = S.upper()
for n in nucleotidos.upper():
Frecuencias.append(S.count(n))
return Frecuencias
ConteoNucleotidos(secuenciaADN)
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
def ConteoNucleotidosL(S, nucleotidos = 'ACGT'):
S = S.upper()
Frecuencias = {}
for n in nucleotidos.upper():
Frecuencias[n] = S.count(n)
return Frecuencias
ConteoNucleotidosL(secuenciaADN)
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
def multiplicacionListas(lista1,lista2):
l = []
suma = 0
if len(lista1)==len(lista2):
for i in range(len(lista1)):
l.append(lista1[i]*lista2[i])
suma += l[i]
print("La suma del producto de los elementos es", suma)
else:
print("Listas con distintas longitudes")
return l
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
def promedioMatriz(matrizOriginal):
n = len(matrizOriginal) # número de filas
m = len(matrizOriginal[0]) # número de columnas
suma = 0 # inicialización de la suma
for i in range(n):
for j in range(m):
suma += matrizOriginal[i][j] # suma sobre todo los
# elementos de la lista
return suma/(n*m)
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
listaInicial = [2,3.5,-2,-4.5, 4, 5, -3.5,7]
listaRaizCuadrada = [ math.sqrt(abs(i)) for i in listaInicial]
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
lRaizCuadradaPos = [ math.sqrt(i) for i in listaInicial if i > 0]
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
lista1 = [3,2,4,7,5]
lista2 = [9,3,4,6,1]
multiplicacion = [i*j for i,j in zip(lista1,lista2)]
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
def multiplicacionListas(lista1,lista2):
if len(lista1)==len(lista2):
l = [i*j for i,j in zip(lista1,lista2)]
print("La suma del producto de los elementos es", sum(l))
else:
print("Listas con distintas longitudes")
l = None
return l
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
[ [ j*i for i in lista1 ] for j in range(1,8)]
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
promedioA = promedioMatriz(A)
B = [[promedioA for i in range(len(A)+2)] \
for j in range(len(A[0])+2)]
for i in range(len(A)):
for j in range(len(A[0])):
B[i+1][j+1] = A[i][j]
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
def imagenBorrosa(matOrig):
n = len(matOrig)
m = len(matOrig[0])
promedio = promedioMatriz(matOrig)
# Inicialización de las matrices B y C
B = [[promedio for j in range(m+2)] for i in range(n+2)]
C = [[matOrig[i][j] for j in range(m)] for i in range(n)]
for i in range(n): # Creación matriz aumentada
for j in range(m):
B[i+1][j+1] = matOrig[i][j]
for i in range(n): # Cálculo del filtro
for j in range(m):
Bij = [[B[k][l] for k in range(i,i+3)] \
for l in range(j,j+3)]
C[i][j] = promedioMatriz(Bij)
return(C)
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
plt.imshow(imagenBorrosa(A), cmap=plt.cm.gray)
plt.axis('off')
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
def contenidoGC(S, W = 64, salto =32):
rhos = []
for k in range(0, len(S), salto):
conteos = ConteoNucleotidos(S[k:(k+W)])
cg = conteos[1]+conteos[2]
rhos.append(cg/(conteos[0]+conteos[3]+cg))
return rhos
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
conSeq1 = contenidoGC('TTGACCGATGACCCCGGTTCAGGCTTCACCACAGT',8,4)
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
import collections
def ConteoNucleotidosC(S, nucleotidos = 'ACGT'):
Frecuencias = collections.Counter(S)
Fsel= {n:Frecuencias[n] for n in Frecuencias.keys()\
& nucleotidos}
return Fsel
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
def contenidoGC2(S, W= 64, salto=32):
return [sum(ConteoNucleotidosC(S[k:(k+W)],'CG').values())\
/len(S[k:(k+W)]) for k in range(0,len(S),salto)]
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
conSeq2=contenidoGC2('TTGACCGATGACCCCGGTTCAGGCTTCACCACAGT',8,4)
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
import os
import urllib
def lecturaGen(archivo):
if not os.path.isfile(archivo):
paginaD = 'https://alexrojas.netlify.app/Data/Prog/'
direccion = paginaD+archivo
with urllib.request.urlopen(direccion) as respuestaURL:
lineas = respuestaURL.readlines()
else:
with open(archivo) as archivoLocal:
lineas = archivoLocal.readlines()
gen = ''.join([l.strip().decode("utf-8") for l in lineas])
return gen
LCT = lecturaGen('LCT.txt')
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
LCT_GC = contenidoGC2(LCT, 1000,500)
# + [markdown] nteract={"transient": {"deleting": false}}
# ## Ciclos *while*
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
for i in range(210):
if (107*i)%210 == 1:
k = i
print(k)
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
k = 0
while (107*k)%210 != 1:
k += 1
print(k)
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
def mcd0(a,b):
minAB = min(a,b)
maxComDiv = 1
for x in range(2,minAB+1):
if b%x == 0 and a%x == 0:
maxComDiv = x
return(maxComDiv)
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
def mcd1(a,b):
res = a%b
while res>0:
a = b
b = res
res = a%b
return(b)
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
def mcd2(a,b):
while b:
a, b = b, a%b
return(a)
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
def BusquedaRaiz(f,l,u, h=0.001):
r = l
fr = f(r)
s = r+h
fs = f(s)
while signo(fr) == signo(fs):
if r > u:
return None
r = s
fr = fs
s = r + h
fs = f(s)
else:
return r,s
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
interesMensual = lambda r: (1+r)**(-60)+40*r-1
lTasa, uTasa = BusquedaRaiz(interesMensual,0.001,0.02)
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
def metodoNewton(f, df, r0, epsilon=1e-4):
delta = abs(0-f(r0))
while delta > epsilon:
dfr0 = df(r0)
if dfr0 == 0:
print("Derivada igual a cero. Método falla")
else:
r0 = r0 - f(r0)/df(r0)
delta = abs(0-f(r0))
return r0
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
d_interesMensual = lambda r: 40 - 60/(1+r)**61
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
rSolucion = metodoNewton(interesMensual, d_interesMensual,
0.014, 1e-5)
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
print(interesMensual(rSolucion))
| static/code/Prog/PPCap3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from pyspark.ml.feature import HashingTF, IDF, Tokenizer
from pyspark.context import SparkContext
from pyspark.sql.session import SparkSession
from pyspark.ml.classification import LogisticRegression
import pyspark.sql.functions as F
import pyspark.sql.types as T
sc = SparkContext('local')
spark = SparkSession(sc)
# -
df = spark.read.format("csv").option("inferschema","true").option("header", "true").option("delimiter", "\t").load("trainReviews.tsv")
tokenizer = Tokenizer(inputCol="text", outputCol="words")
wordsData = tokenizer.transform(df)
wordsData.show(5)
hashingTF = HashingTF(inputCol="words", outputCol="rawFeatures")
tf = hashingTF.transform(wordsData)
tf.show(10)
tf.head().rawFeatures
idf = IDF(inputCol="rawFeatures", outputCol="features", minDocFreq=2).fit(tf)
tfidf = idf.transform(tf)
tfidf.show(5)
ml = LogisticRegression(featuresCol="features", labelCol='category', regParam=0.01)
mlModel = ml.fit(tfidf.limit(5000))
res_train = mlModel.transform(tfidf)
extract_prob = F.udf(lambda x: float(x[1]), T.FloatType())
res_train.withColumn("proba", extract_prob("probability")).select("id", "proba", "prediction").show()
test_df = spark.read.format("csv").option("inferschema","true").option("header", "true").option("delimiter", "\t").load("testReviews.tsv")
tokenizer = Tokenizer(inputCol="text", outputCol="words")
wordsData = tokenizer.transform(test_df)
wordsData.show(5)
test_tf = hashingTF.transform(wordsData)
test_tf.show(10)
test_idf = IDF(inputCol="rawFeatures", outputCol="features", minDocFreq=2).fit(test_tf)
test_tfidf = idf.transform(test_tf)
test_tfidf.show(5)
res_test = mlModel.transform(test_tfidf)
res_test.show(2)
res_test.withColumn("proba", extract_prob("probability")).select("id", "proba", "prediction").show(10)
| ClassifyText.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# # All decoders (except KF, NB, and ensemble) run with subsampled neurons
# ## User Options
# Define what folder you're saving to
# save_folder=''
save_folder='/home/jglaser/Files/Neural_Decoding/Results/'
# Define what folder you're loading the files from
# load_folder=''
load_folder='/home/jglaser/Data/DecData/'
# Define what dataset you are using
dataset='s1'
# dataset='m1'
# dataset='hc'
# Define which decoder to run
run_wf=1 #Wiener Filter
run_wc=0 #Wiener Cascade
run_svr=0 #Support vector regression
run_xgb=0 #XGBoost
run_dnn=0 #Feedforward (dense) neural network
run_rnn=0 #Recurrent neural network
run_gru=0 #Gated recurrent units
run_lstm=0 #Long short term memory network
# Determine how many neurons you're subsampling, and how many times to do this subsampling
# +
num_nrns_used=10 #Number of neurons you're subsampling
num_folds=10 #Number of times to subsample (Note that we've kept it called "num_folds" from our previous codes, even though this isnt cross validation folds)
# -
# ## 1. Import Packages
#
# We import standard packages and functions from the accompanying .py files
# +
#Import standard packages
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from scipy import io
from scipy import stats
import pickle
import time
import sys
#Add the main folder to the path, so we have access to the files there.
#Note that if your working directory is not the Paper_code folder, you may need to manually specify the path to the main folder. For example: sys.path.append('/home/jglaser/GitProj/Neural_Decoding')
sys.path.append('..')
#Import function to get the covariate matrix that includes spike history from previous bins
from preprocessing_funcs import get_spikes_with_history
#Import metrics
from metrics import get_R2
from metrics import get_rho
#Import decoder functions
from decoders import WienerCascadeDecoder
from decoders import WienerFilterDecoder
from decoders import DenseNNDecoder
from decoders import SimpleRNNDecoder
from decoders import GRUDecoder
from decoders import LSTMDecoder
from decoders import XGBoostDecoder
from decoders import SVRDecoder
#Import Bayesian Optimization package
from bayes_opt import BayesianOptimization
# +
#Turn off deprecation warnings
import warnings
warnings.filterwarnings("ignore", category=DeprecationWarning)
# -
# ## 2. Load Data
#
# The data that we load is in the format described below. We have another example script, "Example_format_data" that may be helpful towards putting the data in this format.
#
# Neural data should be a matrix of size "number of time bins" x "number of neurons", where each entry is the firing rate of a given neuron in a given time bin
#
# The output you are decoding should be a matrix of size "number of time bins" x "number of features you are decoding"
# +
if dataset=='s1':
with open(load_folder+'example_data_s1.pickle','rb') as f:
# neural_data,vels_binned=pickle.load(f,encoding='latin1')
neural_data,vels_binned=pickle.load(f)
if dataset=='m1':
with open(load_folder+'example_data_m1.pickle','rb') as f:
# neural_data,vels_binned=pickle.load(f,encoding='latin1')
neural_data,vels_binned=pickle.load(f)
if dataset=='hc':
with open(load_folder+'example_data_hc.pickle','rb') as f:
# neural_data,pos_binned=pickle.load(f,encoding='latin1')
neural_data,pos_binned=pickle.load(f)
# -
# ## 3. Preprocess Data
# ### 3A. User Inputs
# The user can define what time period to use spikes from (with respect to the output).
# +
if dataset=='s1':
bins_before=6 #How many bins of neural data prior to the output are used for decoding
bins_current=1 #Whether to use concurrent time bin of neural data
bins_after=6 #How many bins of neural data after (and including) the output are used for decoding
if dataset=='m1':
bins_before=13 #How many bins of neural data prior to the output are used for decoding
bins_current=1 #Whether to use concurrent time bin of neural data
bins_after=0 #How many bins of neural data after (and including) the output are used for decoding
if dataset=='hc':
bins_before=4 #How many bins of neural data prior to the output are used for decoding
bins_current=1 #Whether to use concurrent time bin of neural data
bins_after=5 #How many bins of neural data after (and including) the output are used for decoding
# -
# ### 3B. Format Covariates
# #### Format Input Covariates
#Remove neurons with too few spikes in HC dataset
if dataset=='hc':
nd_sum=np.nansum(neural_data,axis=0)
rmv_nrn=np.where(nd_sum<100)
neural_data=np.delete(neural_data,rmv_nrn,1)
# Format for recurrent neural networks (SimpleRNN, GRU, LSTM)
# Function to get the covariate matrix that includes spike history from previous bins
X=get_spikes_with_history(neural_data,bins_before,bins_after,bins_current)
# #### Format Output Covariates
#Set decoding output
if dataset=='s1' or dataset=='m1':
y=vels_binned
if dataset=='hc':
y=pos_binned
# #### In HC dataset, remove time bins with no output (y value)
if dataset=='hc':
#Remove time bins with no output (y value)
rmv_time=np.where(np.isnan(y[:,0]) | np.isnan(y[:,1]))
X=np.delete(X,rmv_time,0)
y=np.delete(y,rmv_time,0)
# ### 3C. Define training/testing/validation sets
# We use the same training/testing/validation sets used for the largest training set in Fig. 6
# +
if dataset=='s1' or dataset=='m1':
dt=.05
if dataset=='hc':
dt=.2
if dataset=='hc':
#Size of sets
test_size=int(450/dt) #7.5 min
valid_size=test_size #validation size is the same as the test size
train_size=int(2250/dt) #37.5 min
#End indices
end_idx=np.int(X.shape[0]*.8) #End of test set
tr_end_idx=end_idx-test_size-valid_size #End of training set
if dataset=='s1':
#Size of sets
test_size=int(300/dt) #5 min
valid_size=test_size #validation size is the same as the test size
train_size=int(1200/dt) # 20 min
#End indices
end_idx=np.int(X.shape[0]*.9)#End of test set
tr_end_idx=end_idx-test_size-valid_size #End of training set
if dataset=='m1':
#Size of sets
test_size=int(300/dt) #5 min
valid_size=test_size #validation size is the same as the test size
train_size=int(600/dt) # 10 min
#End indices
end_idx=np.int(X.shape[0]*1)#End of test set
tr_end_idx=end_idx-test_size-valid_size #End of training set
#Range of sets
testing_range=[end_idx-test_size,end_idx] #Testing set (length of test_size, goes up until end_idx)
valid_range=[end_idx-test_size-valid_size,end_idx-test_size] #Validation set (length of valid_size, goes up until beginning of test set)
training_range=[tr_end_idx-train_size,tr_end_idx] #Training set (length of train_size, goes up until beginning of validation set)
# -
# ## 4. Run Decoders
# **Initialize lists of results**
# +
#R2 values
mean_r2_wf=np.empty(num_folds)
mean_r2_wc=np.empty(num_folds)
mean_r2_xgb=np.empty(num_folds)
mean_r2_svr=np.empty(num_folds)
mean_r2_dnn=np.empty(num_folds)
mean_r2_rnn=np.empty(num_folds)
mean_r2_gru=np.empty(num_folds)
mean_r2_lstm=np.empty(num_folds)
#Actual data
y_test_all=[]
y_train_all=[]
y_valid_all=[]
#Test predictions
y_pred_wf_all=[]
y_pred_wc_all=[]
y_pred_xgb_all=[]
y_pred_dnn_all=[]
y_pred_rnn_all=[]
y_pred_gru_all=[]
y_pred_lstm_all=[]
y_pred_svr_all=[]
#Training predictions
y_train_pred_wf_all=[]
y_train_pred_wc_all=[]
y_train_pred_xgb_all=[]
y_train_pred_dnn_all=[]
y_train_pred_rnn_all=[]
y_train_pred_gru_all=[]
y_train_pred_lstm_all=[]
y_train_pred_svr_all=[]
#Validation predictions
y_valid_pred_wf_all=[]
y_valid_pred_wc_all=[]
y_valid_pred_xgb_all=[]
y_valid_pred_dnn_all=[]
y_valid_pred_rnn_all=[]
y_valid_pred_gru_all=[]
y_valid_pred_lstm_all=[]
y_valid_pred_svr_all=[]
# -
# **In the following section, we**
# 1. Loop across iterations (each iteration we subsample different neurons)
# 2. Extract the training/validation/testing data
# 3. Preprocess the data
# 4. Run the individual decoders (whichever have been specified in user options). This includes the hyperparameter optimization
# 5. Save the results
#
# Note that the Wiener Filter, Wiener Cascade, and XGBoost decoders are commented most fully. So look at those for the best understanding.
# +
t1=time.time() #If I want to keep track of how much time has elapsed
num_examples=X.shape[0] #number of examples (rows in the X matrix)
for i in range(num_folds): #Loop through the iterations
#### SUBSAMPLE NEURONS ####
#Randomly subsample "num_nrns_used" neurons
nrn_idxs=np.random.permutation(X.shape[2])[0:num_nrns_used]
X_sub=np.copy(X[:,:,nrn_idxs])
# Format for Wiener Filter, Wiener Cascade, SVR, XGBoost, and Dense Neural Network
#Put in "flat" format, so each "neuron / time" is a single feature
X_flat_sub=X_sub.reshape(X_sub.shape[0],(X_sub.shape[1]*X_sub.shape[2]))
######### SPLIT DATA INTO TRAINING/TESTING/VALIDATION #########
#Note that all sets have a buffer of"bins_before" bins at the beginning, and "bins_after" bins at the end
#This makes it so that the different sets don't include overlapping neural data
#Testing set
testing_set=np.arange(testing_range[0]+bins_before,testing_range[1]-bins_after)
#Validation set
valid_set=np.arange(valid_range[0]+bins_before,valid_range[1]-bins_after)
#Training_set
training_set=np.arange(training_range[0]+bins_before,training_range[1]-bins_after)
#Get training data
X_train=X_sub[training_set,:,:]
X_flat_train=X_flat_sub[training_set,:]
y_train=y[training_set,:]
#Get testing data
X_test=X_sub[testing_set,:,:]
X_flat_test=X_flat_sub[testing_set,:]
y_test=y[testing_set,:]
#Get validation data
X_valid=X_sub[valid_set,:,:]
X_flat_valid=X_flat_sub[valid_set,:]
y_valid=y[valid_set,:]
##### PREPROCESS DATA #####
#Z-score "X" inputs.
X_train_mean=np.nanmean(X_train,axis=0) #Mean of training data
X_train_std=np.nanstd(X_train,axis=0) #Stdev of training data
X_train=(X_train-X_train_mean)/X_train_std #Z-score training data
X_test=(X_test-X_train_mean)/X_train_std #Preprocess testing data in same manner as training data
X_valid=(X_valid-X_train_mean)/X_train_std #Preprocess validation data in same manner as training data
#Z-score "X_flat" inputs.
X_flat_train_mean=np.nanmean(X_flat_train,axis=0)
X_flat_train_std=np.nanstd(X_flat_train,axis=0)
X_flat_train=(X_flat_train-X_flat_train_mean)/X_flat_train_std
X_flat_test=(X_flat_test-X_flat_train_mean)/X_flat_train_std
X_flat_valid=(X_flat_valid-X_flat_train_mean)/X_flat_train_std
#Zero-center outputs
y_train_mean=np.nanmean(y_train,axis=0) #Mean of training data outputs
y_train=y_train-y_train_mean #Zero-center training output
y_test=y_test-y_train_mean #Preprocess testing data in same manner as training data
y_valid=y_valid-y_train_mean #Preprocess validation data in same manner as training data
#Z-score outputs (for SVR)
y_train_std=np.nanstd(y_train,axis=0)
y_zscore_train=y_train/y_train_std
y_zscore_test=y_test/y_train_std
y_zscore_valid=y_valid/y_train_std
################# DECODING #################
#Add actual train/valid/test data to lists (for saving)
y_test_all.append(y_test)
y_train_all.append(y_train)
y_valid_all.append(y_valid)
###### WIENER FILTER ######
if run_wf:
#Note - the Wiener Filter has no hyperparameters to fit, unlike all other methods
#Declare model
model_wf=WienerFilterDecoder()
#Fit model on training data
model_wf.fit(X_flat_train,y_train)
#Get test set predictions
y_test_predicted_wf=model_wf.predict(X_flat_test)
#Get R2 of test set (mean of x and y values of position/velocity)
mean_r2_wf[i]=np.mean(get_R2(y_test,y_test_predicted_wf))
#Print R2 values on test set
R2s_wf=get_R2(y_test,y_test_predicted_wf)
print('R2s_wf:', R2s_wf)
#Add predictions of training/validation/testing to lists (for saving)
y_pred_wf_all.append(y_test_predicted_wf)
y_train_pred_wf_all.append(model_wf.predict(X_flat_train))
y_valid_pred_wf_all.append(model_wf.predict(X_flat_valid))
###### WIENER CASCADE ######
if run_wc:
### Get hyperparameters using Bayesian optimization based on validation set R2 values###
#Define a function that returns the metric we are trying to optimize (R2 value of the validation set)
#as a function of the hyperparameter we are fitting (here, degree)
def wc_evaluate(degree):
model_wc=WienerCascadeDecoder(degree) #Define model
model_wc.fit(X_flat_train,y_train) #Fit model
y_valid_predicted_wc=model_wc.predict(X_flat_valid) #Validation set predictions
return np.mean(get_R2(y_valid,y_valid_predicted_wc)) #R2 value of validation set (mean over x and y position/velocity)
#Do bayesian optimization
wcBO = BayesianOptimization(wc_evaluate, {'degree': (1, 5.01)}, verbose=0) #Define Bayesian optimization, and set limits of hyperparameters
wcBO.maximize(init_points=3, n_iter=3) #Set number of initial runs and subsequent tests, and do the optimization
best_params=wcBO.res['max']['max_params'] #Get the hyperparameters that give rise to the best fit
degree=best_params['degree']
# print("degree=", degree)
### Run model w/ above hyperparameters
model_wc=WienerCascadeDecoder(degree) #Declare model
model_wc.fit(X_flat_train,y_train) #Fit model on training data
y_test_predicted_wc=model_wc.predict(X_flat_test) #Get test set predictions
mean_r2_wc[i]=np.mean(get_R2(y_test,y_test_predicted_wc)) #Get test set R2 (mean across x and y position/velocity)
#Print R2 values on test set
R2s_wc=get_R2(y_test,y_test_predicted_wc)
print('R2s_wc:', R2s_wc)
#Add predictions of training/validation/testing to lists (for saving)
y_pred_wc_all.append(y_test_predicted_wc)
y_train_pred_wc_all.append(model_wc.predict(X_flat_train))
y_valid_pred_wc_all.append(model_wc.predict(X_flat_valid))
###### SVR ######
if run_svr:
### Get hyperparameters using Bayesian optimization based on validation set R2 values###
#Set the maximum number of iterations (to save time) - 2000 for M1 and S1, 4000 for HC which is faster
if dataset=='hc':
max_iter=4000
else:
max_iter=2000
#Define a function that returns the metric we are trying to optimize (R2 value of the validation set)
#as a function of the hyperparameter we are fitting (here, C)
def svr_evaluate(C):
model_svr=SVRDecoder(C=C, max_iter=max_iter)
model_svr.fit(X_flat_train,y_zscore_train) #Note for SVR that we use z-scored y values
y_valid_predicted_svr=model_svr.predict(X_flat_valid)
return np.mean(get_R2(y_zscore_valid,y_valid_predicted_svr))
#Do bayesian optimization
svrBO = BayesianOptimization(svr_evaluate, {'C': (.5, 10)}, verbose=0)
svrBO.maximize(init_points=5, n_iter=5)
best_params=svrBO.res['max']['max_params']
C=best_params['C']
# print("C=", C)
# Run model w/ above hyperparameters
model_svr=SVRDecoder(C=C, max_iter=max_iter)
model_svr.fit(X_flat_train,y_zscore_train) #Note for SVR that we use z-scored y values
y_test_predicted_svr=model_svr.predict(X_flat_test)
mean_r2_svr[i]=np.mean(get_R2(y_zscore_test,y_test_predicted_svr))
#Print R2 values on test set
R2s_svr=get_R2(y_zscore_test,y_test_predicted_svr)
print('R2s_svr:', R2s_svr)
#Add predictions of training/validation/testing to lists (for saving)
y_pred_svr_all.append(y_test_predicted_svr)
y_train_pred_svr_all.append(model_svr.predict(X_flat_train))
y_valid_pred_svr_all.append(model_svr.predict(X_flat_valid))
##### XGBOOST ######
if run_xgb:
### Get hyperparameters using Bayesian optimization based on validation set R2 values###
#Define a function that returns the metric we are trying to optimize (R2 value of the validation set)
#as a function of the hyperparameter we are fitting (max_depth, num_round, eta)
def xgb_evaluate(max_depth,num_round,eta):
max_depth=int(max_depth) #Put in proper format (Bayesian optimization uses floats, and we just want to test the integer)
num_round=int(num_round) #Put in proper format
eta=float(eta) #Put in proper format
model_xgb=XGBoostDecoder(max_depth=max_depth, num_round=num_round, eta=eta) #Define model
model_xgb.fit(X_flat_train,y_train) #Fit model
y_valid_predicted_xgb=model_xgb.predict(X_flat_valid) #Get validation set predictions
return np.mean(get_R2(y_valid,y_valid_predicted_xgb)) #Return mean validation set R2
#Do bayesian optimization
xgbBO = BayesianOptimization(xgb_evaluate, {'max_depth': (2, 10.01), 'num_round': (100,700), 'eta': (0, 1)}) #Define Bayesian optimization, and set limits of hyperparameters
#Set number of initial runs and subsequent tests, and do the optimization. Also, we set kappa=10 (greater than the default) so there is more exploration when there are more hyperparameters
xgbBO.maximize(init_points=20, n_iter=20, kappa=10)
best_params=xgbBO.res['max']['max_params'] #Get the hyperparameters that give rise to the best fit
num_round=np.int(best_params['num_round']) #We want the integer value associated with the best "num_round" parameter (which is what the xgb_evaluate function does above)
max_depth=np.int(best_params['max_depth']) #We want the integer value associated with the best "max_depth" parameter (which is what the xgb_evaluate function does above)
eta=best_params['eta']
# Run model w/ above hyperparameters
model_xgb=XGBoostDecoder(max_depth=max_depth, num_round=num_round, eta=eta) #Declare model w/ fit hyperparameters
model_xgb.fit(X_flat_train,y_train) #Fit model
y_test_predicted_xgb=model_xgb.predict(X_flat_test) #Get test set predictions
mean_r2_xgb[i]=np.mean(get_R2(y_test,y_test_predicted_xgb)) #Get test set R2 (mean across x and y position/velocity)
#Print R2 values on test set
R2s_xgb=get_R2(y_test,y_test_predicted_xgb)
print('R2s:', R2s_xgb)
#Add predictions of training/validation/testing to lists (for saving)
y_pred_xgb_all.append(y_test_predicted_xgb)
y_train_pred_xgb_all.append(model_xgb.predict(X_flat_train))
y_valid_pred_xgb_all.append(model_xgb.predict(X_flat_valid))
##### Dense (Feedforward) NN ######
if run_dnn:
### Get hyperparameters using Bayesian optimization based on validation set R2 values###
#Define a function that returns the metric we are trying to optimize (R2 value of the validation set)
#as a function of the hyperparameter we are fitting
def dnn_evaluate(num_units,frac_dropout,n_epochs):
num_units=int(num_units)
frac_dropout=float(frac_dropout)
n_epochs=int(n_epochs)
model_dnn=DenseNNDecoder(units=[num_units,num_units],dropout=frac_dropout,num_epochs=n_epochs)
model_dnn.fit(X_flat_train,y_train)
y_valid_predicted_dnn=model_dnn.predict(X_flat_valid)
return np.mean(get_R2(y_valid,y_valid_predicted_dnn))
#Do bayesian optimization
dnnBO = BayesianOptimization(dnn_evaluate, {'num_units': (50, 600), 'frac_dropout': (0,.5), 'n_epochs': (2,21)})
dnnBO.maximize(init_points=20, n_iter=20, kappa=10)
best_params=dnnBO.res['max']['max_params']
frac_dropout=float(best_params['frac_dropout'])
n_epochs=np.int(best_params['n_epochs'])
num_units=np.int(best_params['num_units'])
# Run model w/ above hyperparameters
model_dnn=DenseNNDecoder(units=[num_units,num_units],dropout=frac_dropout,num_epochs=n_epochs)
model_dnn.fit(X_flat_train,y_train)
y_test_predicted_dnn=model_dnn.predict(X_flat_test)
mean_r2_dnn[i]=np.mean(get_R2(y_test,y_test_predicted_dnn))
#Print R2 values on test set
R2s_dnn=get_R2(y_test,y_test_predicted_dnn)
print('R2s:', R2s_dnn)
#Add predictions of training/validation/testing to lists (for saving)
y_pred_dnn_all.append(y_test_predicted_dnn)
y_train_pred_dnn_all.append(model_dnn.predict(X_flat_train))
y_valid_pred_dnn_all.append(model_dnn.predict(X_flat_valid))
##### SIMPLE RNN ######
if run_rnn:
### Get hyperparameters using Bayesian optimization based on validation set R2 values###
#Define a function that returns the metric we are trying to optimize (R2 value of the validation set)
#as a function of the hyperparameter we are fitting
def rnn_evaluate(num_units,frac_dropout,n_epochs):
num_units=int(num_units)
frac_dropout=float(frac_dropout)
n_epochs=int(n_epochs)
model_rnn=SimpleRNNDecoder(units=num_units,dropout=frac_dropout,num_epochs=n_epochs)
model_rnn.fit(X_train,y_train)
y_valid_predicted_rnn=model_rnn.predict(X_valid)
return np.mean(get_R2(y_valid,y_valid_predicted_rnn))
#Do bayesian optimization
rnnBO = BayesianOptimization(rnn_evaluate, {'num_units': (50, 600), 'frac_dropout': (0,.5), 'n_epochs': (2,21)})
rnnBO.maximize(init_points=20, n_iter=20, kappa=10)
best_params=rnnBO.res['max']['max_params']
frac_dropout=float(best_params['frac_dropout'])
n_epochs=np.int(best_params['n_epochs'])
num_units=np.int(best_params['num_units'])
# Run model w/ above hyperparameters
model_rnn=SimpleRNNDecoder(units=num_units,dropout=frac_dropout,num_epochs=n_epochs)
model_rnn.fit(X_train,y_train)
y_test_predicted_rnn=model_rnn.predict(X_test)
mean_r2_rnn[i]=np.mean(get_R2(y_test,y_test_predicted_rnn))
#Print R2 values on test set
R2s_rnn=get_R2(y_test,y_test_predicted_rnn)
print('R2s:', R2s_rnn)
#Add predictions of training/validation/testing to lists (for saving)
y_pred_rnn_all.append(y_test_predicted_rnn)
y_train_pred_rnn_all.append(model_rnn.predict(X_train))
y_valid_pred_rnn_all.append(model_rnn.predict(X_valid))
##### GRU ######
if run_gru:
### Get hyperparameters using Bayesian optimization based on validation set R2 values###
#Define a function that returns the metric we are trying to optimize (R2 value of the validation set)
#as a function of the hyperparameter we are fitting
def gru_evaluate(num_units,frac_dropout,n_epochs):
num_units=int(num_units)
frac_dropout=float(frac_dropout)
n_epochs=int(n_epochs)
model_gru=GRUDecoder(units=num_units,dropout=frac_dropout,num_epochs=n_epochs)
model_gru.fit(X_train,y_train)
y_valid_predicted_gru=model_gru.predict(X_valid)
return np.mean(get_R2(y_valid,y_valid_predicted_gru))
#Do bayesian optimization
gruBO = BayesianOptimization(gru_evaluate, {'num_units': (50, 600), 'frac_dropout': (0,.5), 'n_epochs': (2,21)})
gruBO.maximize(init_points=20, n_iter=20,kappa=10)
best_params=gruBO.res['max']['max_params']
frac_dropout=float(best_params['frac_dropout'])
n_epochs=np.int(best_params['n_epochs'])
num_units=np.int(best_params['num_units'])
# Run model w/ above hyperparameters
model_gru=GRUDecoder(units=num_units,dropout=frac_dropout,num_epochs=n_epochs)
model_gru.fit(X_train,y_train)
y_test_predicted_gru=model_gru.predict(X_test)
mean_r2_gru[i]=np.mean(get_R2(y_test,y_test_predicted_gru))
#Print test set R2 values
R2s_gru=get_R2(y_test,y_test_predicted_gru)
print('R2s:', R2s_gru)
#Add predictions of training/validation/testing to lists (for saving)
y_pred_gru_all.append(y_test_predicted_gru)
y_train_pred_gru_all.append(model_gru.predict(X_train))
y_valid_pred_gru_all.append(model_gru.predict(X_valid))
##### LSTM ######
if run_lstm:
### Get hyperparameters using Bayesian optimization based on validation set R2 values###
#Define a function that returns the metric we are trying to optimize (R2 value of the validation set)
#as a function of the hyperparameter we are fitting
def lstm_evaluate(num_units,frac_dropout,n_epochs):
num_units=int(num_units)
frac_dropout=float(frac_dropout)
n_epochs=int(n_epochs)
model_lstm=LSTMDecoder(units=num_units,dropout=frac_dropout,num_epochs=n_epochs)
model_lstm.fit(X_train,y_train)
y_valid_predicted_lstm=model_lstm.predict(X_valid)
return np.mean(get_R2(y_valid,y_valid_predicted_lstm))
#Do bayesian optimization
lstmBO = BayesianOptimization(lstm_evaluate, {'num_units': (50, 600), 'frac_dropout': (0,.5), 'n_epochs': (2,21)})
lstmBO.maximize(init_points=20, n_iter=20, kappa=10)
best_params=lstmBO.res['max']['max_params']
frac_dropout=float(best_params['frac_dropout'])
n_epochs=np.int(best_params['n_epochs'])
num_units=np.int(best_params['num_units'])
# Run model w/ above hyperparameters
model_lstm=LSTMDecoder(units=num_units,dropout=frac_dropout,num_epochs=n_epochs)
model_lstm.fit(X_train,y_train)
y_test_predicted_lstm=model_lstm.predict(X_test)
mean_r2_lstm[i]=np.mean(get_R2(y_test,y_test_predicted_lstm))
#Print test set R2
R2s_lstm=get_R2(y_test,y_test_predicted_lstm)
print('R2s:', R2s_lstm)
#Add predictions of training/validation/testing to lists (for saving)
y_pred_lstm_all.append(y_test_predicted_lstm)
y_train_pred_lstm_all.append(model_lstm.predict(X_train))
y_valid_pred_lstm_all.append(model_lstm.predict(X_valid))
print ("\n") #Line break after each fold
time_elapsed=time.time()-t1 #How much time has passed
###### SAVE RESULTS #####
#Note that I save them after every cross-validation fold rather than at the end in case the code/computer crashes for some reason while running
#Only save results for the decoder we chose to run
if run_wf:
with open(save_folder+dataset+'_results_nrn'+str(num_nrns_used)+'_wf.pickle','wb') as f:
pickle.dump([mean_r2_wf,y_pred_wf_all,y_train_pred_wf_all,y_valid_pred_wf_all],f)
if run_wc:
with open(save_folder+dataset+'_results_nrn'+str(num_nrns_used)+'_wc.pickle','wb') as f:
pickle.dump([mean_r2_wc,y_pred_wc_all,y_train_pred_wc_all,y_valid_pred_wc_all],f)
if run_xgb:
with open(save_folder+dataset+'_results_nrn'+str(num_nrns_used)+'_xgb.pickle','wb') as f:
pickle.dump([mean_r2_xgb,y_pred_xgb_all,y_train_pred_xgb_all,y_valid_pred_xgb_all,time_elapsed],f)
if run_dnn:
with open(save_folder+dataset+'_results_nrn'+str(num_nrns_used)+'_dnn.pickle','wb') as f:
pickle.dump([mean_r2_dnn,y_pred_dnn_all,y_train_pred_dnn_all,y_valid_pred_dnn_all,time_elapsed],f)
if run_rnn:
with open(save_folder+dataset+'_results_nrn'+str(num_nrns_used)+'_rnn.pickle','wb') as f:
pickle.dump([mean_r2_rnn,y_pred_rnn_all,y_train_pred_rnn_all,y_valid_pred_rnn_all,time_elapsed],f)
if run_gru:
with open(save_folder+dataset+'_results_nrn'+str(num_nrns_used)+'_gru.pickle','wb') as f:
pickle.dump([mean_r2_gru,y_pred_gru_all,y_train_pred_gru_all,y_valid_pred_gru_all,time_elapsed],f)
if run_lstm:
with open(save_folder+dataset+'_results_nrn'+str(num_nrns_used)+'_lstm.pickle','wb') as f:
pickle.dump([mean_r2_lstm,y_pred_lstm_all,y_train_pred_lstm_all,y_valid_pred_lstm_all,time_elapsed],f)
if run_svr:
with open(save_folder+dataset+'_results_nrn'+str(num_nrns_used)+'_svr.pickle','wb') as f:
pickle.dump([mean_r2_svr,y_pred_svr_all,y_train_pred_svr_all,y_valid_pred_svr_all,time_elapsed],f)
#Save ground truth results
with open(save_folder+dataset+'_ground_truth_nrns.pickle','wb') as f:
pickle.dump([y_test_all,y_train_all,y_valid_all],f)
print("time_elapsed:",time_elapsed)
# -
# ### Quick check of results
mean_r2_wf
np.mean(mean_r2_wf)
plt.plot(y_test_all[1][0:1000,0])
plt.plot(y_pred_wf_all[1][0:1000,0])
| Paper_code/ManyDecoders_FewNeurons.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# # Tensorflow js model
# > Creating and serving a Tensorflow javascript model in the browser.
#
# - toc: false
# - badges: true
# - comments: true
# - categories: [tfjs, keras, serving]
# + [markdown] colab_type="text" id="taEElfKSBHge"
# In this tutorial we learn how to
#
#
# 1. Train a model with Keras with GPU
# 2. Convert a model to web format
# 3. Upload the model to GitHub Pages
# 4. Prediction using TensorFlow.js
#
#
# + [markdown] colab_type="text" id="b1JCrGrePvKp"
# We will create a simple model that models XOR operation. Given two inputs $(x_0, x_1)$ it outputs $y$
#
# $$\left[\begin{array}{cc|c}
# x_0 & x_1 & y\\
# 0 & 0 & 0\\
# 0 & 1 & 1\\
# 1 & 0 & 1\\
# 1 & 1 & 0
# \end{array}\right]$$
# -
# ## Build the model
# + [markdown] colab_type="text" id="WKYiL-oYR0yk"
# Imports
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="a-UbSG-DR3ID" outputId="c8c804fc-d97d-45df-dff4-6cc729f21951"
from keras.models import Sequential
from keras.layers.core import Dense, Dropout, Activation
from keras.optimizers import SGD
import numpy as np
# + [markdown] colab_type="text" id="UvBySAixR4Ca"
# Initialize the inputs
# + colab={} colab_type="code" id="Hj65iQS6R6pO"
X = np.array([[0,0],[0,1],[1,0],[1,1]])
y = np.array([[0],[1],[1],[0]])
# + [markdown] colab_type="text" id="TTyQKnEgSBQb"
# Create the model
# + colab={} colab_type="code" id="ivnpyw3ZSAF9"
model = Sequential()
model.add(Dense(8, input_dim=2))
model.add(Activation('tanh'))
model.add(Dense(1))
model.add(Activation('sigmoid'))
sgd = SGD(lr=0.1)
model.compile(loss='binary_crossentropy', optimizer=sgd)
# + [markdown] colab_type="text" id="zzrpHO1XSIeJ"
# Train the model
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="jRwYsPJxRrYT" outputId="733d8a80-8589-4d5e-cbc9-9e8f4c8f2c1e"
model.fit(X, y, batch_size=1, epochs=1000, verbose= 0)
# + [markdown] colab_type="text" id="VHlJ2cmpSbZ7"
# Predict the output
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" id="ky1bM2EiSHYt" outputId="cd528c04-469d-479f-eda5-ed610716a9f0"
print(model.predict_proba(X))
# + [markdown] colab_type="text" id="vvdWZCRslZUz"
# Save the model
# + colab={} colab_type="code" id="mxRke-l9lXfY"
model.save('saved_model/keras.h5')
# + [markdown] colab_type="text" id="glkP5CvySfgK"
# ## Convert the model
# + [markdown] colab_type="text" id="q30sPc63lbvw"
# Download the library
# + colab={} colab_type="code" id="-FSJVtS9SiVi"
# !pip install tensorflowjs
# + [markdown] colab_type="text" id="HWCP02udldLr"
# Convert the model
# + colab={} colab_type="code" id="DuQP_mkeSkKL"
# !tensorflowjs_converter --input_format keras saved_model/keras.h5 web_model
# + [markdown] colab_type="text" id="dr8MnQUbUBY7"
# ## Create a web page to serve the model
# + [markdown] colab_type="text" id="SC9QaDQreTDr"
# Import TensorFlow.js
# + colab={} colab_type="code" id="iwJQerK2eA_u"
header = '<head><script src="https://cdn.jsdelivr.net/npm/@tensorflow/tfjs@1.5.2/dist/tf.min.js"> </script>\n'
# + [markdown] colab_type="text" id="GE_Y5U3UeW6U"
# Code for loading the web model. We predict a tensor of zeros and show the result in the page.
# + colab={} colab_type="code" id="kpGEMkjJecBM"
script = '\
<script>\n\
async function loadModel(){ \n\
model = await tf.loadLayersModel(\'web_model/model.json\') \n\
y = model.predict(tf.zeros([1,2])) \n\
document.getElementById(\'out\').innerHTML = y.dataSync()[0] \n\
} \n\
loadModel() \n\
</script>\n\
</head> \n'
# + [markdown] colab_type="text" id="0TDOfXR6f9tp"
# Body of the page
# + colab={} colab_type="code" id="cf5VErepf9H0"
body = '\
<body>\n\
<p id =\'out\'></p> \n\
</body>'
# + [markdown] colab_type="text" id="2DaBOiA-jTER"
# Save the code as html file
# + colab={} colab_type="code" id="pM6JIkRCglMu"
with open('index.html','w') as f:
f.write(header+script+body)
f.close()
| _notebooks/2020-07-02-tfjs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Customer Churn em Operadoras de Telecom
# ## 1. Definição do Problema
# 
# **Customer Churn** (ou Rotatividade de Clientes, em uma tradução livre)
# refere-se a uma decisão tomada pelo cliente sobre o término do relacionamento
# comercial. Refere-se também à perda de clientes. A fidelidade do cliente e a
# rotatividade de clientes sempre somam 100%. Se uma empresa tem uma taxa de
# fidelidade de 60%, então a taxa de perda de clientes é de 40%. De acordo com a
# regra de lucratividade do cliente 80/20, 20% dos clientes estão gerando 80% da
# receita. Portanto, é muito importante prever os usuários que provavelmente
# abandonarão o relacionamento comercial e os fatores que afetam as decisões do
# cliente.
# **Objetivo:** Realizar a previsão do customer churn em uma operadora de telecom. Criar um modelo de aprendizagem de máquina (regressão logística) que possa prever se um cliente vai cancelar seu plano (1 ou 0/Sim ou Não) e a probabilidade de uma opção ou outra.
# ## 2. Dados
# ### 2.1. Importando as bibliotecas
# +
# Manipulação de dados
import pandas as pd
# Computação científica
import numpy as np
# Plotagem de gráficos
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# Filtro de Warnings
import warnings
warnings.filterwarnings('ignore')
# Normalização e padronização
from sklearn.preprocessing import MinMaxScaler, StandardScaler
# Balanceamento
from imblearn.over_sampling import SMOTE
# Multicolinearidade
from statsmodels.stats.outliers_influence import variance_inflation_factor
from statsmodels.tools.tools import add_constant
# Machine Learning
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
from sklearn.metrics import classification_report
from sklearn.metrics import roc_auc_score
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_val_score
from sklearn.linear_model import LogisticRegression
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.neighbors import KNeighborsClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.tree import DecisionTreeClassifier
from sklearn.svm import SVC
from sklearn.ensemble import RandomForestClassifier
import pickle
# -
# ### 2.2. Carregando os Dados
# Os dados são fornecidos pela plataforma de ensino Data Science Academy, sendo que, estão divididos em datasets de treino e teste. Dessa forma, todas as manipulações, transformações, entre outros aspectos relacionados ao tratamento dos dados, precisam ser realizados nos dados de treino, como também nos dados de teste.
# Dados de treino
dataTrain = pd.read_csv('C:/Users/otavio/Mundo/Big-Data-Analytics-Python-Spark/Projeto4-Prevendo-Customer-Churn/projeto4_telecom_treino.csv', index_col = 0)
# Primeiras linhas dados de treino
dataTrain.head()
# Dados de teste
dataTest = pd.read_csv('C:/Users/otavio/Mundo/Big-Data-Analytics-Python-Spark/Projeto4-Prevendo-Customer-Churn/projeto4_telecom_teste.csv', index_col = 0)
# Primeiras linhas dados de teste
dataTest.head()
# ### 2.3. Dicionário dos Dados
# Nome das colunas/variávies
dataTrain.columns
# Descrição de cada coluna
descricao = ['Estado', 'Número de dias na qual a conta está ativa', 'Código da área ', 'Plano internacional', \
'Plano de voz', 'Número de menssagens de voz realizadas', \
'Total de minutos falados durante o dia', 'Total de chamadas realizadas durante o dia', \
'Total pago por chamadas realizadas durante o dia', 'Total de minutos falados no início da noite', \
'Total de chamadas realizadas no início da noite', 'Total pago por ligações realizadas no início da noite', \
'Total de minutos falados no fim da noite', 'Total de chamadas realizadas no fim da noite', \
'Total pago por ligações realizadas no fim da noite', 'Total de minutos falados em chamadas internacionais', \
'Total de chamadas internacionais realizadas', 'Total cobrado por chamadas internacionais', \
'Número de chamadas feitas para o atendimento ao cliente', 'Se o cliente cancelou ou não o serviço (variável target)']
# +
# Dicionário dos dados
dict_dados = {'Variaveis' : [dataTrain.columns[i] for i in range(20)],
'Descricao' : descricao,
'Tipo de dados' : [dataTrain.dtypes[i] for i in range(20)]}
# Dataframe do dicionário de dados
pd.set_option('max_colwidth', 100) # ajustando o tamanho das colunas
df_dict_dados = pd.DataFrame(dict_dados)
df_dict_dados
# -
# Salvando o dicionário de dados
df_dict_dados.to_csv('dicionario_dados.csv', sep = ',')
# Retornando o tamanho normal das colunas
pd.reset_option('max_colwidth')
# ## 3. Data Munging
# Shape dos dados de treino
dataTrain.shape
# Visão geral dos dados de treino
dataTrain.info()
# Os dados de treino possuem 20 colunas e 3333 linhas, ou seja, cada uma das colunas possui 3333 registros. Outro ponto importante é o fato do dataset não conter valores nulos. Além disso, será realizada algumas transformações nas variáveis, como também a formação de variáveis novas.
dataTrain.head()
# Função na qual realiza a transformação e criação de features
def transformFeatures(dataset):
# Transforma as features que retornam yes/no em 1/0 e atribui a uma nova feature
dictYesNo = {'yes' : 1, 'no' : 0}
dataset['international_plan_num'] = dataset.international_plan.map(dictYesNo)
dataset['voice_mail_plan_num'] = dataset.voice_mail_plan.map(dictYesNo)
dataset['churn_num'] = dataset.churn.map(dictYesNo)
# Remove as strings dos códigos de Area, preservando somente o código em um novo campo
dictAreaCode = {"area_code_415": 0, "area_code_408": 1, "area_code_510": 2}
dataset['area_code_num'] = dataset['area_code'].map(dictAreaCode)
# Transforma o valor dos status em Numeros em um novo campo
dictState = {'KS': 1000, 'OH': 1001, 'NJ': 1002, 'OK': 1003, 'AL': 1004, 'MA': 1005,
'MO': 1006, 'LA': 1007, 'WV': 1008, 'IN': 1009, 'RI': 1010,
'IA': 1011, 'MT': 1012, 'NY': 1013, 'ID': 1014, 'VT': 1015,
'VA': 1016, 'TX': 1017, 'FL': 1018, 'CO': 1019, 'AZ': 1020,
'SC': 1021, 'NE': 1022, 'WY': 1023, 'HI': 1024, 'IL': 1025,
'NH': 1026, 'GA': 1027, 'AK': 1028, 'MD': 1029, 'AR': 1030,
'WI': 1031, 'OR': 1032, 'MI': 1033, 'DE': 1034, 'UT': 1035,
'CA': 1036, 'MN': 1037, 'SD': 1038, 'NC': 1039, 'WA': 1040,
'NM': 1041, 'NV': 1042, 'DC': 1043, 'KY': 1044, 'ME': 1045,
'MS': 1046, 'TN': 1047, 'PA': 1048, 'CT': 1049, 'ND': 1050}
dataset['state_num'] = dataset['state'].map(dictState)
# Transforma algumas features em variáveis categóricas
dataset['international_plan'] = dataset['international_plan'].astype('category')
dataset['voice_mail_plan'] = dataset['voice_mail_plan'].astype('category')
dataset['area_code'] = dataset['area_code'].astype('category')
dataset['state'] = dataset['state'].astype('category')
# Criando novas features: total de minutos, total de ligações e total pago pelo cliente
dataset['total_minutes'] = dataset.total_day_minutes + dataset.total_eve_minutes + dataset.total_night_minutes + dataset.total_intl_minutes
dataset['total_calls'] = dataset.total_day_calls + dataset.total_eve_calls + dataset.total_night_calls + dataset.total_intl_calls
dataset['total_charge'] = dataset.total_day_charge + dataset.total_eve_charge + dataset.total_night_charge + dataset.total_intl_charge
# Período na qual o cliente mais utliza o serviço
shift = []
for i in dataset.values:
if i[6] > i[9] and i[6] > i[12]:
shift.append('day')
elif i[9] > i[6] and i[9] > i[12]:
shift.append('evening')
elif i[12] > i[6] and i[12] > i[9]:
shift.append('night')
else:
shift.append('no preference')
dataset['shift'] = shift
dataset['shift'] = dataset['shift'].astype('category')
return dataset
# Aplicação da função transformFeatures
dataTrain = transformFeatures(dataTrain)
dataTrain.info()
# Novo dataset construído após as transformações aplicadas
dataTrain.head()
# ## 4. Análise Exploratória dos Dados
# Algumas estatísticas descritivas dos dados
dataTrain.describe()
# Através da breve análise realizada acima, pode-se perceber que será necessário aplicar uma normalização aos dados, uma vez que encontram-se em diferentes escalas. Além disso, será utilizado o algoritmo de regressão logística, na qual obtém melhores resultados quando os dados estão normalizados.
# Distribuição da variável churn, a variável target, aquela a ser prevista
churn_dist = dataTrain.churn.value_counts()
churn_dist
# +
# Percentual de clientes que cancelaram e que não cancelaram o serviço
churn_no = str(round((churn_dist[0] * 100 / (churn_dist[0] + churn_dist[1])), 2)) + '%'
churn_yes = str(round((churn_dist[1] * 100 / (churn_dist[0] + churn_dist[1])), 2)) + '%'
# Construção do gráfico
fig, ax = plt.subplots(figsize = (8, 6))
sns.countplot(x = 'churn', data = dataTrain)
ax.set_title('Porcentagem de Churn', fontsize = 15)
plt.xlabel('Churn', fontsize = 13)
plt.ylabel('Contagem', fontsize = 13)
ax.text(0, 1400, churn_no,fontsize = 15,color = 'white', ha = 'center', va = 'center')
ax.text(1, 200, churn_yes, fontsize = 15,color = 'white', ha = 'center', va = 'center')
# Mostrar gráfico
plt.show()
# -
# O dataset encontra-se desbalanceado, ou seja, possui muito mais registros de não cancelamento do que o oposto. Este fato pode causar um viés no modelo de machine learning. Portanto, será preciso balancear a variável churn para que, dessa forma, seja possível obter um resultado mais confiável.
# +
# Histogramas: minutos, ligações e pagamentos
# Área de plotagem
fig, ax = plt.subplots(1, 3, figsize = (18, 6))
# Gráfico minutos
ax[0].hist(dataTrain.total_minutes, bins = 10)
ax[0].set_title('Minutos')
# Gráfico ligações
ax[1].hist(dataTrain.total_calls, bins = 10)
ax[1].set_title('Ligações')
# Gráfico pagamentos
ax[2].hist(dataTrain.total_charge, bins = 10)
ax[2].set_title('Pagamentos')
plt.show()
# -
# A grande maioria dos clientes desta empresa realizam entre 250 e 350 ligações, que possuem duração entre 450 e 700 minutos, totalizando pagamentos entre 50 e 70 dólares. Além disso, é importante salientar que estes dados encontram-se normalizados, isto é, respeitam uma distribuição normal.
# +
# Relacionamento entre a variável churn e algumas outras
fig, ax = plt.subplots(1, 3, figsize = (15, 3))
# Gráfico 1: state x churn
sns.countplot(x = 'state', hue = 'churn', data = dataTrain, ax = ax[0])
ax[0].set_title('Estado de residência dos clientes')
ax[0].set_xlabel('')
# Gráfico 2: area_code x churn
sns.countplot(x = 'area_code', hue = 'churn', data = dataTrain, ax = ax[1])
ax[1].set_title('Código de área dos clientes')
ax[1].set_xlabel('')
ax[1].set_ylabel('')
# Gráfico 3: international_plan x churn
sns.countplot(x = 'international_plan', hue = 'churn', data = dataTrain, ax = ax[2])
ax[2].set_title('Clientes com planos internacionais')
ax[2].set_xlabel('')
ax[2].set_ylabel('')
plt.show()
# -
# Obervamos que o estado na qual o cliente se encontra pouco interfere na sua decisão de cancelar ou não o serviço. O mesmo é observado ao código de área dos clientes, embora, proporcionalmente os clientes dos códigos de área 408 e 510 cancelem mais o serviço quando comparamos com os clientes do cogido de área 415.
#
# Em relação aos clientes com planos internacionais, nota-se um alto cancelamento do serviço daqueles clientes que adquirem o plano, enquanto os que não adquirem possuem uma taxa de cancelamento baixa. Planos internacionais são mais caros e necessitam de um bom funcionamento. Outro ponto importante refere-se ao fato desses clientes possivelmente estarem dentro daqueles 20% de clientes que representam 80% dos lucros da empresa.
# +
# Relacionamento entre a variável churn e algumas outras
fig, ax = plt.subplots(1, 3, figsize = (15, 3))
# Gráfico 1: voice_mail_plan x churn
sns.countplot(x = 'voice_mail_plan', hue = 'churn', data = dataTrain, ax = ax[0])
ax[0].set_title('Clientes com plano de mensagem de voz')
ax[0].set_xlabel('')
# Gráfico 2: shift x churn
sns.countplot(x = 'shift', hue = 'churn', data = dataTrain, ax = ax[1])
ax[1].set_title('Período do serviço mais utilizado pelos clientes')
ax[1].set_xlabel('Período')
ax[1].set_ylabel('')
# Gráfico 3: number_customer_service_calls x churn
sns.countplot(x = 'number_customer_service_calls', hue = 'churn', data = dataTrain, ax = ax[2])
ax[2].set_title('Clientes que buscaram atendimento')
ax[2].set_xlabel('Número de atendimentos')
ax[2].set_ylabel('')
plt.show()
# -
# O fato de clientes possuírem ou não plano com mensagem de voz parece não influenciar no cancelamento do serviço. Assim como também, o período na qual o serviço é utilizado não parece ter influência nessa decisão.
#
# Por outro lado, em relação aos clientes que buscam atendimento, os clientes na qual pouco procuram o atendimento não costumam realizar o cancelamento. Para os usuários do serviço que procuram atendimento 4 vezes ou mais, esse parece ser um fator determinante na hora de cancelar o serviço, devido ao alto churn.
# +
# Histogramas: As features apresentam distribuição normal?
features = dataTrain.select_dtypes(exclude = ['category', 'object']).columns
for item in features:
sns.distplot(dataTrain[item])
plt.title('Histograma')
plt.show()
# -
# Através dos histogramas pode-se perceber que a grande maioria das features estão de acordo com uma distribuição gaussiana, ou seja, estão normalizadas. A excessão são as features categóricas, aonde já era esperado que não estivessem de acordo com uma distribuição normal. Porém, as features number_vmail_messages, total_int_calls e number_customer_service_calls não encontram-se normalizadas, dessa forma será preciso realizar a padronização dos dados, de forma que todas as features representem uma distribuição gaussiana.
# ## 5. Transformações dos dados
# Remoção das variávies categóricas, uma vez que já foram criadas outras variávies numéricas em seus lugares.
# Serão removidas também variávies criadas apenas para a análise exploratória.
data = dataTrain.drop(['state', 'area_code', 'international_plan', 'voice_mail_plan', 'churn', 'total_minutes',
'total_calls', 'total_charge', 'shift'], axis = 1)
# Variável target
target = data['churn_num']
# Removendo a variável target do conjunto de dados
data = data.drop(['churn_num'], axis = 1)
# ### 5.1. Normalização
# Processo que confere aos dados estarem em uma mesma escala, ou seja, em um range que varia de 0 a 1.
# +
# Transformando o dataframe em um array numpy
data_np = data.values
# Normalização
scaler = MinMaxScaler(feature_range = (0, 1))
data_normalized = scaler.fit_transform(data_np)
# -
# ### 5.2. Padronização
# Processo que transforma os dados de acordo com uma distribuição normal. Possui melhor desempenho com os dados normalizados.
scaler = StandardScaler().fit(data_normalized)
data_stardard = scaler.transform(data_normalized)
# ### 5.3. Balanceamento
# O método a ser utilizado para resolver o problema do balanceamento dos dados é o SMOTE. Esse método consiste em gerar dados sintéticos (não duplicados) da classe minoritária a partir de vizinhos. Ele calcula quais são os vizinhos mais próximos e as características desses vizinhos para criar novos dados.
balancer = SMOTE()
data_stardard, target_balancer = balancer.fit_resample(data_stardard, target)
# ### 5.4. Correlação e Colinearidade
# Caso as variáveis forem muito correlacionadas, as inferências baseadas no modelo de regressão podem ser errôneas ou pouco confiáveis. Por isso, é importante verificar a colinearidade entre as variávies.
# Matriz de correlação
corr = pd.DataFrame(data_stardard, columns = data.columns).corr(method = 'pearson')
# +
# Gráfico matriz de correlação
fig, ax = plt.subplots(figsize = (20, 20))
sns.heatmap(corr, vmin = 0, vmax = 1, linewidths = 1, annot = True, cmap = 'Spectral')
ax.set_title('Matriz de Correlação', fontsize = 20)
plt.show()
# -
# Podemos diagnosticar Multicolinearidade por meio do VIF (Variance Inflation Factor). Os VIFs medem o quanto a variância de um coeficiente de regressão estimado aumenta se seus preditores estão correlacionados. Se todos os VIFs forem 1, não há multicolinearidade, mas se alguns VIFs forem maiores do que 1, os preditores estão correlacionados. Quando um VIF é 5 > 10, o coeficiente de regressão para esse termo não é estimado de maneira apropriada.
# +
# Datafram com os dados prontos
vif = pd.DataFrame(data_stardard, columns = data.columns)
# Coeficiente
x = add_constant(vif)
# VIF
vif_series = pd.Series([variance_inflation_factor(x.values, i) for i in range(x.shape[1])], index = x.columns)
vif = pd.DataFrame(vif_series, columns = ['VIF Factor'])
vif
# -
# De acordo com o VIF, as variáveis total_night_minutes, total_night_charge, total_intl_minutes e total_intl_charge ultrapassam o valor máximo de 5, dessa forma, serão excluídas do dataset.
# Remoção das variáveis colineares
data_stardard = pd.DataFrame(data_stardard).drop(columns = [8, 10, 11, 13]).values
# ## 6. Machine Learning
# Com as features devidamente tratadas e escolhidas é possível dar início ao processo de Machine Learning. A proposta inicial era utilizar o modelo de Regressão Logística para fazer as previsões, entretanto, vou utilizar também outros modelos para fazer comparações e, dessa forma, escolher aquele mais adequado.
# Divisão do dataset em dados de treino e dados de teste
test_size = 0.3
x_treino, x_teste, y_treino, y_teste = train_test_split(data_stardard, target_balancer, test_size = test_size)
# ### 6.1. Regressão Logística
# +
# Criação do modelo
modelo = LogisticRegression()
# Treinamento do modelo
modelo.fit(x_treino, y_treino)
# Previsoes
previsoes1 = modelo.predict(x_teste)
# -
# Relatório de classificação
print('Relatório de Classificação:\n', classification_report(y_teste, previsoes1))
print("Acurácia: %.3f" % (accuracy_score(y_teste, previsoes1) * 100))
print('AUC: %.3f' % (roc_auc_score(y_teste, previsoes1) * 100))
# Salvando o modelo
arquivo1 = 'modelo/model_reg_log.sav'
pickle.dump(modelo, open(arquivo1, 'wb'))
# O modelo apresentou um bom desempenho, uma vez que sua acurácia encontra-se acima de 70%, faixa considerada boa para o problema a ser resolvido. A métrica AUC, área do grpafico que fica sob a curva, ou seja, a precisão do modelo, também obteve porcentagem satisfatória.
# ### 6.1.1. Regressão Logística com Cross Validation
# +
# Definindo os valores para o número de folds
num_folds = 10
# Separando os dados em folds
kfold = KFold(num_folds, True)
# Criando o modelo
modelo = LogisticRegression()
# Cross Validation
resultado = cross_val_score(modelo, data_stardard, target_balancer, cv = kfold)
# Print do resultado
print("Acurácia: %.3f" % (resultado.mean() * 100))
# -
# Ao utilizar cross validation para realizar a partição dos dados de treino e de teste, não foi notada nenhuma melhora significativa do modelo, pelo contrário, verificou-se uma leve queda na sua performance.
# ### 6.2. Linear Discriminant Analysis
# +
# Criação do modelo
modelo = LinearDiscriminantAnalysis()
# Treinamento do modelo
modelo.fit(x_treino, y_treino)
# Previsoes
previsoes2 = modelo.predict(x_teste)
# -
# Relatório de classificação
print('Relatório de Classificação:\n', classification_report(y_teste, previsoes2))
print("Acurácia: %.3f" % (accuracy_score(y_teste, previsoes2) * 100))
print('AUC: %.3f' % (roc_auc_score(y_teste, previsoes2) * 100))
# Foi observada uma leve melhora nesse modelo, porém nada significativo.
# Salvando o modelo
arquivo2 = 'modelo/model_LDA.sav'
pickle.dump(modelo, open(arquivo2, 'wb'))
# ### 6.3. KNN - K-Nearest Neighbors
# +
# Criação do modelo
modelo = KNeighborsClassifier()
# Treinamento do modelo
modelo.fit(x_treino, y_treino)
# Previsoes
previsoes3 = modelo.predict(x_teste)
# -
# Relatório de classificação
print('Relatório de Classificação:\n', classification_report(y_teste, previsoes3))
print("Acurácia: %.3f" % (accuracy_score(y_teste, previsoes3) * 100))
print('AUC: %.3f' % (roc_auc_score(y_teste, previsoes3) * 100))
# Com o algoritmo KNN foi observada uma grande melhora na performance do modelo, porém o modelo apredeu de maneira enviesada (recall).
# Salvando o modelo
arquivo3 = 'modelo/model_KNN.sav'
pickle.dump(modelo, open(arquivo3, 'wb'))
# ### 6.4. Naive Bayes
# +
# Criação do modelo
modelo = GaussianNB()
# Treinamento do modelo
modelo.fit(x_treino, y_treino)
# Previsoes
previsoes4 = modelo.predict(x_teste)
# -
# Relatório de classificação
print('Relatório de Classificação:\n', classification_report(y_teste, previsoes4))
print("Acurácia: %.3f" % (accuracy_score(y_teste, previsoes4) * 100))
print('AUC: %.3f' % (roc_auc_score(y_teste, previsoes4) * 100))
# Esse modelo apresentou melhora em relação ao de Regressão Logística e o Linear Discrimiant Analysis, porém não foi superior ao KNN, apesar de ter aprendido de maneira mais equilibrada (recall).
# Salvando o modelo
arquivo4 = 'modelo/model_naive_bayes.sav'
pickle.dump(modelo, open(arquivo4, 'wb'))
# ### 6.5. CART (Classification and Regression Trees)
# +
# Criação do modelo
modelo = DecisionTreeClassifier()
# Treinamento do modelo
modelo.fit(x_treino, y_treino)
# Previsoes
previsoes5 = modelo.predict(x_teste)
# -
# Relatório de classificação
print('Relatório de Classificação:\n', classification_report(y_teste, previsoes5))
print("Acurácia: %.3f" % (accuracy_score(y_teste, previsoes5) * 100))
print('AUC: %.3f' % (roc_auc_score(y_teste, previsoes5) * 100))
# Até o momento é o modelo com melhor desempenho, uma vez que apresenta a maior acurácia, assimo como também a maior AUC. É notável também que o modelo aprendeu de maneira equilibrada entre as classes.
# Salvando o modelo
arquivo5 = 'modelo/model_CART.sav'
pickle.dump(modelo, open(arquivo5, 'wb'))
# ### 6.6. SVM - Support Vector Machines
# +
# Criação do modelo
modelo = SVC()
# Treinamento do modelo
modelo.fit(x_treino, y_treino)
# Previsoes
previsoes6 = modelo.predict(x_teste)
# -
# Relatório de classificação
print('Relatório de Classificação:\n', classification_report(y_teste, previsoes6))
print("Acurácia: %.3f" % (accuracy_score(y_teste, previsoes6) * 100))
print('AUC: %.3f' % (roc_auc_score(y_teste, previsoes6) * 100))
# Modelo com resultado muito semelhante ao CART, porém com leve queda na acurácia e AUC, além de um leve desbalanceamente entre as classes.
# Salvando o modelo
arquivo6 = 'modelo/model_SVM.sav'
pickle.dump(modelo, open(arquivo6, 'wb'))
# ### 6.7. Random Forest
# +
# Criação do modelo
modelo = RandomForestClassifier()
# Treinamento do modelo
modelo.fit(x_treino, y_treino)
# Previsoes
previsoes7 = modelo.predict(x_teste)
# -
# Relatório de classificação
print('Relatório de Classificação:\n', classification_report(y_teste, previsoes7))
print("Acurácia: %.3f" % (accuracy_score(y_teste, previsoes7) * 100))
print('AUC: %.3f' % (roc_auc_score(y_teste, previsoes7) * 100))
# Apresentou um ótimo resultado, na acurácia e AUC, como também no balanceamento entre classes. Portanto, este modelo será utilizado na sequência do trabalho.
# Salvando o modelo
arquivo7 = 'modelo/model_random_forest.sav'
pickle.dump(modelo, open(arquivo7, 'wb'))
# ## 7. Previsão
# Para realizar as previsões, primeiramente é necessário realizar as mesmas transformações que os dados de treino tiveram nos dados de teste, para que dessa forma eles estejam nivelados.
# Aplicando a função transformFeatures
dataTest2 = transformFeatures(dataTest)
# Remocção de variáveis
dataTeste3 = dataTest2.drop(['state', 'area_code', 'international_plan', 'voice_mail_plan', 'churn', 'total_minutes',
'total_calls', 'total_charge', 'shift', 'churn_num'], axis = 1)
# +
# Normalização
# Transformando o dataframe em um array numpy
data_np = dataTeste3.values
# Normalização
scaler = MinMaxScaler(feature_range = (0, 1))
data_normalized_test = scaler.fit_transform(data_np)
# -
# Padronização
scaler = StandardScaler().fit(data_normalized_test)
data_stardard_test = scaler.transform(data_normalized_test)
# Remoção das variáveis colineares
data_stardard_test = pd.DataFrame(data_stardard_test).drop(columns = [8, 10, 11, 13]).values
# ### 7.1. Aplicação do modelo Random Forest
# Baixando modelo
model = pickle.load(open(arquivo7, 'rb'))
# Previsões de churn e probabilidade
predicted_churn = model.predict(data_stardard_test)
predicted_proba = model.predict_proba(data_stardard_test)
# Dataframe com as previsões
predicted_churn_df = pd.DataFrame(predicted_churn, columns = ['Churn'])
predict_proba_df = pd.DataFrame(predicted_proba, columns = ['Prob. Não Cancelamento', 'Prob. Cancelamento'])
# Resultado
result = pd.concat([predicted_churn_df, predict_proba_df], axis = 1)
result.head(20)
# Churn: 0/não - 1/sim
result.groupby('Churn').size()
# ## 8. Conclusão
# De acordo com este trabalho, 1394 clientes vão optar por continuarem com o serviço, enquanto que 273 clientes irão optar pelo seu cancelamento. Dessa forma, podemos perceber o quanto é importante para uma empresa saber extrair resultados a partir dos seus dados, para que então possam tomar aquelas decisões que considerem como sendo as mais corretas para o negócio em questão.
# Para que um modelo obtenha um bom resultado é de suma importância realizar as etapas de análise exploratória e pré-processamento, para que então os modelos consigam atingir seu potencial máximo.
# Com este trabalho, foi possível determinar a rotatitividade dos clientes da empresa de Telecom, estipulando se o cliente vai cancelar seu serviço ou não e qual a probabilidade de que uma coisa ou outra ocorra. Em um primeiro momento, o algoritmo de Regressão Logística iria ser utilizado, porém ao analisarmos os demais algoritmos foi possível verificar que o modelo Random Forest era mais adequado para essa situação, uma vez que obteve melhor acurácia, AUC, além de ter feito um aprendizado bastante equilibrado entre as classes.
| .ipynb_checkpoints/Projeto4-Customer-Churn-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %pylab inline
import scipy.integrate as integrate
from IPython.html.widgets import interact
# # El Oscilador armonico.
#
# Dibujamos el espacio de fases para la ecuacion $$\ddot{x} = -\omega^2x$$
#
# Para eso lo pasamos a un sistema:
#
# $$
# \begin{cases}
# \dot{V_{x}} = -\omega^2 x\\
# \dot{x} = V_{x}
# \end{cases}
# $$
#
# +
@interact(xin=(-5,5,0.1),yin=(-5,5,0.1))
def plotInt(xin,yin):
xmax = 2
vmax = 5
x = linspace(-xmax, xmax, 15) # Definimos el rango en el que se mueven las variables y el paso
v = linspace(-vmax, vmax, 15)
X, V = meshgrid(x,v) # Creamos una grilla con eso
# Definimos las constantes
w = 3
# Definimos las ecuaciones
Vp = -w**2*X
Xp = V
def resorte(y, t):
yp = y[1]
vp = -w**2*y[0]
return [yp, vp]
x0 = [xin, yin]
t = linspace(0,10,2000)
sh = integrate.odeint(resorte, x0, t)
fig = figure(figsize(10,5))
ax1 = subplot(121) # Hacer el grafico
quiver(X, V, Xp, Vp, angles='xy')
plot(x, [0]*len(x) ,[0]*len(v), v)
lfase = plot(sh[:,0],sh[:,1],'.')
ylim((-vmax,vmax))
xlim((-xmax,xmax))
# Retocarlo: tamanios, colores, leyendas, etc...
xlabel('$x$', fontsize=16)
ylabel('$\\dot{x}$',fontsize=16)
ax1.set_title('Espacio de fases')
ax2 = subplot(122) # Hacer otro grafico
lines = plot(t,sh )
xlabel('Tiempo [s]')
ax2.set_title('Espacio de tiempo')
legend(['Posicion','Velocidad'])
tight_layout()
ylim((-xmax, xmax))
# -
# # El Pendulo
#
# Dibujamos el espacio de fases para la ecuacion $$\ddot{\theta} = -\frac{g}{l}sin(\theta)$$
#
# Para eso lo pasamos a un sistema:
#
# $$
# \begin{cases}
# \dot{V_{\theta}} = -\frac{g}{l}sin(\theta)\\
# \dot{\theta} = V_{\theta}
# \end{cases}
# $$
#
@interact(thI=(0,np.pi,0.1),vI=(0,5,0.1))
def plotInt(thI, vI):
h = linspace(-pi,pi,15) # Definimos el rango en el que se mueven las variables y el paso
v = linspace(-10,10,15)
H, V = meshgrid(h,v) # Creamos una grilla con eso
# Definimos las constantes
g = 10
l = 1
# Definimos las ecuaciones
Vp = -g/l*sin(H)
Hp = V
def pendulo(y, t):
hp = y[1]
vp = -g/l*sin(y[0])
return [hp, vp]
y0 = [thI, vI]
t = linspace(0,10,2000)
sh = integrate.odeint(pendulo, y0, t)
fig = figure(figsize(10,5))
ax1 = subplot(121) # Hacer el grafico
quiver(H, V, Hp, Vp, angles='xy')
plot(h, [0]*len(h) ,[0]*len(v), v)
sh[:,0] = np.mod(sh[:,0] + np.pi, 2*np.pi) - np.pi
lfase = plot(sh[:,0], sh[:,1],'.')
# Retocarlo: tamanios, colores, leyendas, etc...
xlabel('$\\theta$', fontsize=16)
ylabel('$\\dot{\\theta}$', fontsize=16)
xlim((-pi,pi))
ylim((-10,10))
xtick = arange(-1,1.5,0.5)
x_label = [ r"$-\pi$",
r"$-\frac{\pi}{2}$", r"$0$",
r"$+\frac{\pi}{2}$", r"$+\pi$",
]
ax1.set_xticks(xtick*pi)
ax1.set_xticklabels(x_label, fontsize=20)
ax1.set_title('Espacio de fases')
ax2 = subplot(122) # Hacer otro grafico
lines = plot(t,sh )
ylim((-pi, pi))
ytick = [-pi, 0, pi]
y_label = [ r"$-\pi$", r"$0$", r"$+\pi$"]
ax2.set_yticks(ytick)
ax2.set_yticklabels(y_label, fontsize=20)
xlabel('Tiempo [s]')
ax2.set_title('Espacio de tiempo')
legend(['Posicion','Velocidad'])
tight_layout()
# # El Pendulo con perdidas
#
# Dibujamos el espacio de fases para la ecuacion $$\ddot{\theta} = -\frac{g}{l}sin(\theta) - \gamma \dot \theta$$
#
# Para eso lo pasamos a un sistema:
#
# $$
# \begin{cases}
# \dot{V_{\theta}} = -\frac{g}{l}sin(\theta) - \gamma \dot \theta\\
# \dot{\theta} = V_{\theta}
# \end{cases}
# $$
#
@interact(th0=(-2*np.pi,2*np.pi,0.1),v0=(-2,2,0.1))
def f(th0 = np.pi/3, v0 = 0):
h = linspace(-pi,pi,15) # Definimos el rango en el que se mueven las variables y el paso
v = linspace(-10,10,15)
H, V = meshgrid(h,v) # Creamos una grilla con eso
# Definimos las constantes
g = 10
l = 1
ga = 0.5
# Definimos las ecuaciones
Vp = -g/l*sin(H) - ga*V #SOLO CAMBIA ACA
Hp = V
def pendulo(y, t):
hp = y[1]
vp = -g/l*sin(y[0]) - ga* y[1] # Y ACAA
return [hp, vp]
y0 = [th0, v0]
t = linspace(0,10,2000)
sh = integrate.odeint(pendulo, y0, t)
fig = figure(figsize(10,5))
ax1 = subplot(121) # Hacer el grafico
quiver(H, V, Hp, Vp, angles='xy')
plot(h, [0]*len(h) , h , -g/l/ga*sin(h)) # Dibujar nulclinas
lfase = plot(sh[:,0],sh[:,1],'.')
# Retocarlo: tamanios, colores, leyendas, etc...
xlabel('$\\theta$', fontsize=16)
ylabel('$\\dot{\\theta}$',fontsize=16)
xlim((-pi,pi))
ylim((-10,10))
xtick = arange(-1,1.5,0.5)
x_label = [ r"$-\pi$",
r"$-\frac{\pi}{2}$", r"$0$",
r"$+\frac{\pi}{2}$", r"$+\pi$",
]
ax1.set_xticks(xtick*pi)
ax1.set_xticklabels(x_label, fontsize=20)
ax1.set_title('Espacio de fases')
ax2 = subplot(122) # Hacer otro grafico
lines = plot(t,sh )
ylim((-pi, pi))
ytick = [-pi, 0, pi]
y_label = [ r"$-\pi$", r"$0$", r"$+\pi$"]
ax2.set_yticks(ytick)
ax2.set_yticklabels(y_label, fontsize=20)
xlabel('Tiempo [s]')
ax2.set_title('Espacio de tiempo')
legend(['Posicion','Velocidad'])
tight_layout()
# # El resorte Oscilaciones longitudinales.
#
# Dibujamos el espacio de fases para la ecuacion $$\ddot{y} = -2\frac{k}{m}\left(1-\frac{l_0}{\sqrt{y^2+l^2}}\right)y$$
#
# Para eso lo pasamos a un sistema:
#
# $$
# \begin{cases}
# \dot{V_{y}} = -2\frac{k}{m}\left(1-\frac{l_0}{\sqrt{y^2+l^2}}\right)y\\
# \dot{y} = V_{y}
# \end{cases}
# $$
#
@interact(x0=(-1,1,0.1),v0=(0,1,0.1))
def f(x0=0,v0=1):
ymax = 2
vmax = 5
y = linspace(-ymax, ymax, 15) # Definimos el rango en el que se mueven las variables y el paso
v = linspace(-vmax, vmax, 15)
Y, V = meshgrid(y,v) # Creamos una grilla con eso
# Definimos las constantes
k = 10
l = 1
l0 = 1.2
m = 1
# Definimos las ecuaciones
Vp = -2*k/m*(1-l0/(sqrt(Y**2+l**2)))*Y
Yp = V
def resorte(y, t):
yp = y[1]
vp = -2*k/m*(1-l0/(sqrt(y[0]**2+l**2)))*y[0]
return [yp, vp]
y0 = [x0, v0]
t = linspace(0,10,2000)
sh = integrate.odeint(resorte, y0, t)
fig = figure(figsize(10,5))
ax1 = subplot(121) # Hacer el grafico
quiver(Y, V, Yp, Vp, angles='xy')
plot(y, [0]*len(y) ,[0]*len(v), v)
lfase = plot(sh[:,0],sh[:,1],'.')
ylim((-vmax,vmax))
xlim((-ymax,ymax))
# Retocarlo: tamanios, colores, leyendas, etc...
xlabel('$y$', fontsize=16)
ylabel('$\\dot{y}$', fontsize=16)
ax1.set_title('Espacio de fases')
ax2 = subplot(122) # Hacer otro grafico
lines = plot(t,sh )
xlabel('Tiempo [s]')
ax2.set_title('Espacio de tiempo')
legend(['Posicion','Velocidad'])
tight_layout()
ylim((-ymax, ymax))
| python/Extras/Fisica2/Espacio de Fases.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## April 25, 2021 ##
#
# Dear all,
#
# The following notebook covers necessary syntaxes in Python, mainly about repetition (for-loop and while-loop) and conditionals (if-else and if-else if)
#
# I wrote this code using Jupyter Notebook, feel free to test each block and see if you get the same results as mine.
# ## Part 1: For Loop ##
#
# a. For-Loop that increments by one
#
# b. For-Loop that increment with more than one
#
# c. For-Loop that decrements by one (counting down)
#
# d. For-Loop that decrements with more than one
#
# e. For-Loop through a container
## a. For-Loop that increments by one
for i in range(5):
print("The counter is at {}".format(i))
## b. For-Loop that increments by more than one
step = 2
## In this case, three arguments are required: starting point, ending point, and increment value
for i in range(0, 10, step):
print("The counter is at {}".format(i))
## c. For-Loop that decrements by one
step = -1
for i in range(10, 0, step):
print("The counter is at {}".format(i))
## d. For-Loop that decrements by more than one
step = -2
for i in range(10, 0, step):
print("The counter is at {}".format(i))
# +
## e. For-Loop through a container
test_array = ["Apple", "Banana", "Carrots", "Pineapple", "Grapes", "Lychee"]
print("The length of my container: {}".format(len(test_array)))
for i in range( len(test_array) ) :
print("The name of the fruit is {}".format(test_array[i]))
# -
# ## Part 2: While Loop ##
## a. While-Loop that increments by one
step = 1; final = 10; i = 0
while i < final:
print("The counter is at {}".format(i))
i = i+step
## b. While-Loop that increments by more than one
step = 2; final = 10; i = 0
while i < final:
print("The counter is at {}".format(i))
i = i+step
## c. While-Loop that decrements by one
step = -1; final = 0; i = 10
while i > final:
print("The counter is at {}".format(i))
i = i+step
## d. While-Loop that decrements by more than one
step = -2; final = 0; i = 10
while i > final:
print("The counter is at {}".format(i))
i = i+step
# +
## e. While-Loop through a container (list or array)
test_array = ["Apple", "Banana", "Carrots", "Pineapple", "Grapes", "Lychee"]
print("The length of my container: {}".format(len(test_array)))
i = 0 # Declare a counter and do not forget to set its initial value
while i < len(test_array):
print("The name of the fruit is {}".format(test_array[i]))
i = i+1 # Make sure to increment the counter or otherwise the while loop runs in infinite loop
# -
# ## Part 3: If-Else Conditional ##
# ## Part 4: If-Else If Conditional ##
| 2021_04_23_Functions/.ipynb_checkpoints/python_syntaxes-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# =======================
# Whats New 0.98.4 Legend
# =======================
#
# Create a legend and tweak it with a shadow and a box.
#
# +
import matplotlib.pyplot as plt
import numpy as np
ax = plt.subplot(111)
t1 = np.arange(0.0, 1.0, 0.01)
for n in [1, 2, 3, 4]:
plt.plot(t1, t1**n, label="n=%d"%(n,))
leg = plt.legend(loc='best', ncol=2, mode="expand", shadow=True, fancybox=True)
leg.get_frame().set_alpha(0.5)
plt.show()
# -
# ------------
#
# References
# """"""""""
#
# The use of the following functions, methods, classes and modules is shown
# in this example:
#
#
import matplotlib
matplotlib.axes.Axes.legend
matplotlib.pyplot.legend
matplotlib.legend.Legend
matplotlib.legend.Legend.get_frame
| matplotlib/gallery_jupyter/pyplots/whats_new_98_4_legend.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# $\newcommand{\xv}{\mathbf{x}}
# \newcommand{\wv}{\mathbf{w}}
# \newcommand{\yv}{\mathbf{y}}
# \newcommand{\zv}{\mathbf{z}}
# \newcommand{\uv}{\mathbf{u}}
# \newcommand{\vv}{\mathbf{v}}
# \newcommand{\Chi}{\mathcal{X}}
# \newcommand{\R}{\rm I\!R}
# \newcommand{\sign}{\text{sign}}
# \newcommand{\Tm}{\mathbf{T}}
# \newcommand{\Xm}{\mathbf{X}}
# \newcommand{\Zm}{\mathbf{Z}}
# \newcommand{\I}{\mathbf{I}}
# \newcommand{\Um}{\mathbf{U}}
# \newcommand{\Vm}{\mathbf{V}}
# \newcommand{\muv}{\boldsymbol\mu}
# \newcommand{\Sigmav}{\boldsymbol\Sigma}
# \newcommand{\Lambdav}{\boldsymbol\Lambda}
# \newcommand\numberthis{\addtocounter{equation}{1}\tag{\theequation}}
# $
#
# ### ITCS 8010: Machine Learning with Graphs and Large Networks (Fall 2020)
#
# # Problem Set 2
# <font color="blue"> <NAME> </font>
#
# The purpose of these exercises is to explore the `Signed` and `Disease Spreading` network. To get more understanding about the topics, we further implemented a couple of graph analysis algotithms. For example, to analyze the deviation of users behavior of a `Signed` network, we computed the `generative` and `receptive` surprise. For `Disease Spreading` network, we implemented the `SIR Model` to understand the likelihood of an epidemic in a given network. We further test our implementations on the provided datasets.
#
# For this homework, I installed the `NetworkX` [[2]]() network analysis package. The details about the installation of `NetworkX` can be found in [[3]](https://networkx.github.io/documentation/stable/install.html). Here is the software packages I used in this assignment:
# * Python 3.7.6
# * NetworkX 2.5
# * Numpy
# * Matplotlib
# ## 1. Analyzing the Signed networks [50 points]
#
# Online networks are very useful in analyzing the social theory of structural balance and status inequality.
# * Reading: [Signed Networks in Social Media](https://cs.stanford.edu/~jure/pubs/triads-chi10.pdf)
# * Reading: [Predicting Positive and Negative Links in Online Social Networks](https://www.cs.cornell.edu/home/kleinber/www10-signed.pdf)
#
# I have downloaded the [Slashdot](http://snap.stanford.edu/data/soc-sign-Slashdot081106.html) dataset and conduct analysis to answer the following questions:
# 1. <font color="blue">Compute the number of triangles in the network.</font>
# * <b>Answer:</b>
# * Number of cycles: 391205
# * Number of triangles: 22936
# 2. <font color="blue">Report the fraction of balanced triangles and unbalanced triangles. (assume network is undirected; if there is a sign for each direction, randomly pick one.)</font>
# * <b>Answer:</b>
# * Number of balanced triangles: 19238
# * Number of unbalanced triangles: 3698
# 3. <font color="blue">Compare the frequency of signed triads in real and "shuffled" networks (refer slides) (assume network is undirected; if there is a sign for each direction, randomly pick one.)</font>
# * <b>Answer:</b>
# * Edge-list size with negative sign: 114959
# * Number of cycles: 391205
# * Number of triad: 22936
# * ($+++$) Original Vs. Shuffeled: 14605 - 9605
# * ($--+$) Original Vs. Shuffeled: 4633 - 3347
# * ($+--$) Original Vs. Shuffeled: 2918 - 9576
# * ($---$) Original Vs. Shuffeled: 780 - 408
# <table>
# <tr>
# <th>Is Balanced?</th>
# <th>Triad</th>
# <th colspan="2" align="middle">Slashdot</th>
# </tr>
# <tr>
# <th></th>
# <th></th>
# <th>Real Network</th>
# <th>Shuffled Network</th>
# </tr>
# <tr>
# <th rowspan="2">Balanced</th>
# <th>+++</th>
# <td>0.64</td>
# <td>0.42</td>
# </tr>
# <tr>
# <th>--+</th>
# <td>0.20</td>
# <td>0.15</td>
# </tr>
# <tr>
# <th rowspan="2">Unbalanced</th>
# <th>+--</th>
# <td>0.13</td>
# <td>0.42</td>
# </tr>
# <tr>
# <th>---</th>
# <td>0.03</td>
# <td>0.02</td>
# </tr>
# <tr>
# <td colspan="4" align="middle"><b>Table 1: Compare frequencies of signed triads in real and "shuffled" signs.</b></td>
# </tr>
# </table>
# 4. <font color="blue">Compute "Gen. Surprise" (assume directed signed networks) for each of the 16 types.</font>
# * <b>Answer:</b>
# <table>
# <tr>
# <th>Type</th>
# <th>Generative Surprise</th>
# </tr>
# <tr>
# <td>Type-1</td>
# <td>388.25</td>
# </tr>
# <tr>
# <td>Type-2</td>
# <td>-3.74</td>
# </tr>
# <tr>
# <td>Type-3</td>
# <td>388.54</td>
# </tr>
# <tr>
# <td>Type-4</td>
# <td>-4.08</td>
# </tr>
# <tr>
# <td>Type-5</td>
# <td>6.46</td>
# </tr>
# <tr>
# <td>Type-6</td>
# <td>4.66</td>
# </tr>
# <tr>
# <td>Type-7</td>
# <td>6.17</td>
# </tr>
# <tr>
# <td>Type-8</td>
# <td>2.28</td>
# </tr>
# <tr>
# <td>Type-9</td>
# <td>405.85</td>
# </tr>
# <tr>
# <td>Type-10</td>
# <td>4.59</td>
# </tr>
# <tr>
# <td>Type-11</td>
# <td>104.56</td>
# </tr>
# <tr>
# <td>Type-12</td>
# <td>-2.70</td>
# </tr>
# <tr>
# <td>Type-13</td>
# <td>3.60</td>
# </tr>
# <tr>
# <td>Type-14</td>
# <td>26.36</td>
# </tr>
# <tr>
# <td>Type-15</td>
# <td>-2.60</td>
# </tr>
# <tr>
# <td>Type-16</td>
# <td>-1.84</td>
# </tr>
# <tr>
# <td colspan="2" align="middle"><b>Table 2: "Generative Surprise" on Slashdot dataset</b></td>
# </tr>
# </table>
# 5. <font color="blue">Rewrite the formula for "Rec. Surprise" using the idea introduced in "Gen. Surprise".</font>
# * <b>Answer:</b>
#
# `Generative baseline` ($p_g$) represents the fraction of positive feedback ($+$) given by a user. On the other hand, `receptive baseline` ($p_r$) the fraction of positive feedback ($+$) received by a user. In this context, `surprise` represents the behavioural deviation of two users ($A/B$) from baseline w.r.t. another user ($X$).
#
# The formula for `Generative Surprise` of context X is:
# $$
# \begin{align*}
# s_g(X) &= \frac{k - \sum_{i=1}^{n} p_g(A_i)}{\sqrt{\sum_{i=1}^{n} p_g(A_i) * (1 - p_g(A_i))}}\\
# \end{align*}
# $$
#
# Here,
# * $p_g(A_i)$ represents generative baseline of $A_i$
# * Context $X$ represents all the ($A, B, X$) triads, i.e., ($A_1, B_1| X_1$),..., ($A_n, B_n| X_n$)
# * $k$ is the number of triad instances where $X$ closed with a plus edges
#
# Receptive surprise is similar, just use $p_r(A_i)$. So, the formula for `Receptive Surprise` of context X is:
# $$
# \begin{align*}
# s_r(X) &= \frac{k - \sum_{i=1}^{n} p_r(A_i)}{\sqrt{\sum_{i=1}^{n} p_r(A_i) * (1 - p_r(A_i))}}\\
# \end{align*}
# $$
#
# Here, $p_r(A_i)$ represents receptive baseline of $A_i$.
#
# 6. <font color="blue">Compute “Rec. Surprise” for all each of the 16 types.</font>
# * <b>Answer:</b>
# <table>
# <tr>
# <th>Type</th>
# <th>Receptive Surprise</th>
# </tr>
# <tr>
# <td>Type-1</td>
# <td>401.58</td>
# </tr>
# <tr>
# <td>Type-2</td>
# <td>-5.02</td>
# </tr>
# <tr>
# <td>Type-3</td>
# <td>400.85</td>
# </tr>
# <tr>
# <td>Type-4</td>
# <td>-5.10</td>
# </tr>
# <tr>
# <td>Type-5</td>
# <td>4.77</td>
# </tr>
# <tr>
# <td>Type-6</td>
# <td>3.14</td>
# </tr>
# <tr>
# <td>Type-7</td>
# <td>5.65</td>
# </tr>
# <tr>
# <td>Type-8</td>
# <td>0.55</td>
# </tr>
# <tr>
# <td>Type-9</td>
# <td>402.80</td>
# </tr>
# <tr>
# <td>Type-10</td>
# <td>4.31</td>
# </tr>
# <tr>
# <td>Type-11</td>
# <td>102.21</td>
# </tr>
# <tr>
# <td>Type-12</td>
# <td>-2.17</td>
# </tr>
# <tr>
# <td>Type-13</td>
# <td>5.95</td>
# </tr>
# <tr>
# <td>Type-14</td>
# <td>26.76</td>
# </tr>
# <tr>
# <td>Type-15</td>
# <td>-0.87</td>
# </tr>
# <tr>
# <td>Type-16</td>
# <td>-0.58</td>
# </tr>
# <tr>
# <td colspan="2" align="middle"><b>Table 3: "Receptive Surprise" on Slashdot dataset</b></td>
# </tr>
# </table>
import networkx as nx
import numpy as np
import matplotlib.pyplot as plt
import collections as collec
import math
import random
from random import randrange
# %matplotlib inline
# Reading `Slashdot` graph as an weighted undirected graph:
G_slash_undir = nx.read_edgelist("data/soc-sign-Slashdot.txt", nodetype=int, data=(("sign", int),), edgetype=int, comments='#', create_using=nx.Graph)
# Check basic graph properties:
print("Number of nodes: {}".format(nx.number_of_nodes(G_slash_undir)))
print("Number of edges: {}".format(nx.number_of_edges(G_slash_undir)))
# <font color="blue">1. Compute the number of triangles in the network.</font>
# +
cycle_list = nx.cycle_basis(G_slash_undir)
print("Number of cycles: {}".format(len(cycle_list)))
triangle_count = 0
for cycle in cycle_list:
if len(cycle) == 3:
triangle_count += 1
print("Number of triangles: {}".format(triangle_count))
# -
# <font color="blue">2. Report the fraction of balanced triangles and unbalanced triangles. (assume network is undirected; if there is a sign for each direction, randomly pick one.)</font>
# +
cycle_list = nx.cycle_basis(G_slash_undir)
triad_count = 0
print("Number of cycles: {}".format(len(cycle_list)))
fff_tri_count = 0
eef_tri_count = 0
ffe_tri_count = 0
eee_tri_count = 0
for cycle in cycle_list:
if len(cycle) == 3:
triad_count += 1
# print(cycle)
sum_sign = G_slash_undir[cycle[0]][cycle[1]]['sign'] + G_slash_undir[cycle[1]][cycle[2]]['sign'] + G_slash_undir[cycle[2]][cycle[0]]['sign']
# print(G_slash_undir[cycle[0]][cycle[1]]['sign'])
# print(G_slash_undir[cycle[1]][cycle[2]]['sign'])
# print(G_slash_undir[cycle[2]][cycle[0]]['sign'])
if sum_sign == 3:
fff_tri_count += 1
if sum_sign == -1:
eef_tri_count += 1
if sum_sign == 1:
ffe_tri_count += 1
if sum_sign == -3:
eee_tri_count += 1
print("Number of triads: {}".format(triad_count))
print("Number of balanced triangles: {}".format(fff_tri_count + eef_tri_count))
print("Number of unbalanced triangles: {}".format(ffe_tri_count + eee_tri_count))
# -
# <font color="blue">3. Compare the frequency of signed triads in real and “shuffled” networks (refer slides) (assume network is undirected; if there is a sign for each direction, randomly pick one.)</font>
# +
G_slash_undir_shuffle = nx.read_edgelist("data/soc-sign-Slashdot.txt", nodetype=int, data=(("sign", int),), edgetype=int, comments='#', create_using=nx.Graph)
num_nodes = nx.number_of_nodes(G_slash_undir_shuffle)
#print("Edges: {}".format(G_slash_undir_shuffle.edges(data=True)))
# selecting negative edges
edge_list = [(u,v) for u,v,e in G_slash_undir_shuffle.edges(data=True) if e['sign'] == -1]
random.seed()
print("Edge-list size with negative sign: {}".format(len(edge_list)))
#print(edge_list)
for edge in edge_list:
shuffeled = False
while shuffeled == False:
u = random.randint(0, num_nodes)
v = random.randint(0, num_nodes)
#print(u)
#print(v)
if G_slash_undir_shuffle.has_edge(u, v) and G_slash_undir_shuffle[u][v]['sign'] == 1:
shuffeled = True
G_slash_undir_shuffle[u][v]['sign'] = -1
G_slash_undir_shuffle[edge[0]][edge[1]]['sign'] = 1
print("Shuffeled the edge signs!")
cycle_list = nx.cycle_basis(G_slash_undir_shuffle)
triad_count = 0
print("Number of cycles: {}".format(len(cycle_list)))
s_fff_tri_count = 0
s_eef_tri_count = 0
s_ffe_tri_count = 0
s_eee_tri_count = 0
for cycle in cycle_list:
if len(cycle) == 3:
triad_count += 1
# print(cycle)
sum_sign = G_slash_undir_shuffle[cycle[0]][cycle[1]]['sign'] + G_slash_undir_shuffle[cycle[1]][cycle[2]]['sign'] + G_slash_undir_shuffle[cycle[2]][cycle[0]]['sign']
# print(G_slash_undir[cycle[0]][cycle[1]]['sign'])
# print(G_slash_undir[cycle[1]][cycle[2]]['sign'])
# print(G_slash_undir[cycle[2]][cycle[0]]['sign'])
if sum_sign == 3:
s_fff_tri_count += 1
if sum_sign == -1:
s_eef_tri_count += 1
if sum_sign == 1:
s_ffe_tri_count += 1
if sum_sign == -3:
s_eee_tri_count += 1
print("Number of triads: {}".format(triad_count))
print("fff Original Vs. Shuffeled: {} - {}".format(fff_tri_count, s_fff_tri_count))
print("eef Original Vs. Shuffeled: {} - {}".format(eef_tri_count, s_eef_tri_count))
print("ffe Original Vs. Shuffeled: {} - {}".format(ffe_tri_count, s_ffe_tri_count))
print("eee Original Vs. Shuffeled: {} - {}".format(eee_tri_count, s_eee_tri_count))
# -
# Reading `Slashdot` graph as an weighted directed graph:
G_slash_dir = nx.read_edgelist("data/soc-sign-Slashdot.txt", nodetype=int, data=(("sign", int),), edgetype=int, comments='#', create_using=nx.DiGraph)
# Check basic graph properties:
print("Number of nodes: {}".format(nx.number_of_nodes(G_slash_dir)))
print("Number of edges: {}".format(nx.number_of_edges(G_slash_dir)))
# <font color="blue">4. Compute “Gen. Surprise” (assume directed signed networks) for each of the 16 types</font>
# +
# G_slash_dir = nx.read_edgelist("data/tmp.txt", nodetype=int, data=(("sign", int),), edgetype=int, comments='#', create_using=nx.DiGraph)
# store K value for each type
K = {1: 0, 2: 0, 3: 0, 4: 0, 5: 0, 6: 0, 7: 0, 8: 0, 9: 0, 10: 0, 11: 0, 12: 0, 13: 0, 14: 0, 15: 0, 16: 0}
# store P_g value for each type
P_g = {1: 0.0, 2: 0.0, 3: 0.0, 4: 0.0, 5: 0.0, 6: 0.0, 7: 0.0, 8: 0.0, 9: 0.0, 10: 0.0, 11: 0.0, 12: 0.0, 13: 0.0, 14: 0.0, 15: 0.0, 16: 0.0}
# store Denominator value for each type's S_g formula, needs to make square-root over this value
Denominator = {1: 0.0, 2: 0.0, 3: 0.0, 4: 0.0, 5: 0.0, 6: 0.0, 7: 0.0, 8: 0.0, 9: 0.0, 10: 0.0, 11: 0.0, 12: 0.0, 13: 0.0, 14: 0.0, 15: 0.0, 16: 0.0}
"""
Triad type matcher on graph "G_slash_dir".
This function assumes there is a directed edge from node "a" to node "b".
attributes
==========
a node-id
b node-id
x node-id
"""
def triad_type_matcher(a, b, x):
ret = []
# print("a: {}, b: {}, x: {}".format(a, b, x))
if G_slash_dir.has_edge(a, x) and G_slash_dir.has_edge(x, b):
# print("a->x: {}, x->b: {}".format(G_slash_dir[a][x]['sign'], G_slash_dir[x][b]['sign']))
if G_slash_dir[a][x]['sign'] == 1 and G_slash_dir[x][b]['sign'] == 1:
ret.append(1)
elif G_slash_dir[a][x]['sign'] == 1 and G_slash_dir[x][b]['sign'] == -1:
ret.append(2)
elif G_slash_dir[a][x]['sign'] == -1 and G_slash_dir[x][b]['sign'] == 1:
ret.append(5)
elif G_slash_dir[a][x]['sign'] == -1 and G_slash_dir[x][b]['sign'] == -1:
ret.append(6)
if G_slash_dir.has_edge(a, x) and G_slash_dir.has_edge(b, x):
# print("a->x: {}, b->x: {}".format(G_slash_dir[a][x]['sign'], G_slash_dir[b][x]['sign']))
if G_slash_dir[a][x]['sign'] == 1 and G_slash_dir[b][x]['sign'] == 1:
ret.append(3)
elif G_slash_dir[a][x]['sign'] == 1 and G_slash_dir[b][x]['sign'] == -1:
ret.append(4)
elif G_slash_dir[a][x]['sign'] == -1 and G_slash_dir[b][x]['sign'] == 1:
ret.append(7)
elif G_slash_dir[a][x]['sign'] == -1 and G_slash_dir[b][x]['sign'] == -1:
ret.append(8)
if G_slash_dir.has_edge(x, a) and G_slash_dir.has_edge(x, b):
# print("x->a: {}, x->b: {}".format(G_slash_dir[x][a]['sign'], G_slash_dir[x][b]['sign']))
if G_slash_dir[x][a]['sign'] == 1 and G_slash_dir[x][b]['sign'] == 1:
ret.append(9)
elif G_slash_dir[x][a]['sign'] == 1 and G_slash_dir[x][b]['sign'] == -1:
ret.append(10)
elif G_slash_dir[x][a]['sign'] == -1 and G_slash_dir[x][b]['sign'] == 1:
ret.append(13)
elif G_slash_dir[x][a]['sign'] == -1 and G_slash_dir[x][b]['sign'] == -1:
ret.append(14)
if G_slash_dir.has_edge(x, a) and G_slash_dir.has_edge(b, x):
# print("x->a: {}, b->x: {}".format(G_slash_dir[x][a]['sign'], G_slash_dir[b][x]['sign']))
if G_slash_dir[x][a]['sign'] == 1 and G_slash_dir[b][x]['sign'] == 1:
ret.append(11)
elif G_slash_dir[x][a]['sign'] == 1 and G_slash_dir[b][x]['sign'] == -1:
ret.append(12)
elif G_slash_dir[x][a]['sign'] == -1 and G_slash_dir[b][x]['sign'] == 1:
ret.append(15)
elif G_slash_dir[x][a]['sign'] == -1 and G_slash_dir[b][x]['sign'] == -1:
ret.append(16)
return ret
"""
Triad type matcher on graph "G_slash_dir".
This function assumes there is a directed edge from node "a" to node "b".
attributes
==========
a node-id
b node-id
x node-id
"""
def update_dictionary(a, b, x, type_):
if G_slash_dir[a][b]['sign'] == 1:
K[type_] += 1
out_deg = G_slash_dir.out_degree(a)
pos_out_deg = len([(u,v) for u,v,e in G_slash_dir.out_edges(a, data=True) if e['sign'] == 1])
# print("Node-id: {}, Pos-OutEdges: {}".format(a, [(u,v) for u,v,e in G_slash_dir.out_edges(data=True) if e['sign'] == 1]))
# print("Pos-OutDeg: {}, OutDeg: {}".format(pos_out_deg, out_deg))
P_g[type_] += (pos_out_deg/float(out_deg))
Denominator[type_] += ((pos_out_deg/float(out_deg)) * (1 - (pos_out_deg/float(out_deg))))
def print_gen_surprise():
for type_ in range(1, 17):
gen_surp = 0.0
if Denominator[type_] != 0.0:
gen_surp = K[type_] - P_g[type_] / math.sqrt(Denominator[type_])
# else:
# print("Type-{} is not covered. K: {}, P_g: {}, Denominator: {}".format(type_, K[type_], P_g[type_], Denominator[type_]))
print("Generative Surprise for type-{}: {}".format(type_, gen_surp))
def calculate_gen_surprise():
triad_gen = nx.all_triads(G_slash_dir)
triad_nums = 0
for triad in triad_gen:
triad_nodes = list(triad.nodes())
# print(type(triad_nodes))
# print(triad_nodes)
triad_nums += 1
if triad_nums % 100000 == 0:
print("Finished {} triads.".format(triad_nums))
if triad_nums >= 10000000:
break
if G_slash_dir.has_edge(triad_nodes[0], triad_nodes[1]):
types = triad_type_matcher(triad_nodes[0], triad_nodes[1], triad_nodes[2])
# print("a: {} to b: {} got: {}".format(triad_nodes[0], triad_nodes[1], types))
for type_ in types:
update_dictionary(triad_nodes[0], triad_nodes[1], triad_nodes[2], type_)
if G_slash_dir.has_edge(triad_nodes[1], triad_nodes[0]):
types = triad_type_matcher(triad_nodes[1], triad_nodes[0], triad_nodes[2])
# print("a: {} to b: {} got: {}".format(triad_nodes[1], triad_nodes[0], types))
for type_ in types:
update_dictionary(triad_nodes[1], triad_nodes[0], triad_nodes[2], type_)
if G_slash_dir.has_edge(triad_nodes[1], triad_nodes[2]):
types = triad_type_matcher(triad_nodes[1], triad_nodes[2], triad_nodes[0])
# print("a: {} to b: {} got: {}".format(triad_nodes[1], triad_nodes[2], types))
for type_ in types:
update_dictionary(triad_nodes[1], triad_nodes[2], triad_nodes[0], type_)
if G_slash_dir.has_edge(triad_nodes[2], triad_nodes[1]):
types = triad_type_matcher(triad_nodes[2], triad_nodes[1], triad_nodes[0])
# print("a: {} to b: {} got: {}".format(triad_nodes[2], triad_nodes[1], types))
for type_ in types:
update_dictionary(triad_nodes[2], triad_nodes[1], triad_nodes[0], type_)
if G_slash_dir.has_edge(triad_nodes[0], triad_nodes[2]):
types = triad_type_matcher(triad_nodes[0], triad_nodes[2], triad_nodes[1])
# print("a: {} to b: {} got: {}".format(triad_nodes[0], triad_nodes[2], types))
for type_ in types:
update_dictionary(triad_nodes[0], triad_nodes[2], triad_nodes[1], type_)
if G_slash_dir.has_edge(triad_nodes[2], triad_nodes[0]):
types = triad_type_matcher(triad_nodes[2], triad_nodes[0], triad_nodes[1])
# print("a: {} to b: {} got: {}".format(triad_nodes[2], triad_nodes[0], types))
for type_ in types:
update_dictionary(triad_nodes[2], triad_nodes[0], triad_nodes[1], type_)
calculate_gen_surprise()
print_gen_surprise()
# -
# <font color="blue">5. Rewrite the formula for “Rec. Surprise” using the idea introduced in “Gen. Surprise”</font>
# `Generative baseline` ($p_g$) represents the fraction of positive feedback ($+$) given by a user. On the other hand, `receptive baseline` ($p_r$) the fraction of positive feedback ($+$) received by a user. In this context, `surprise` represents the behavioural deviation of two users ($A/B$) from baseline w.r.t. another user ($X$).
#
# The formula for `Generative Surprise` of context X is:
# $$
# \begin{align*}
# s_g(X) &= \frac{k - \sum_{i=1}^{n} p_g(A_i)}{\sqrt{\sum_{i=1}^{n} p_g(A_i) * (1 - p_g(A_i))}}\\
# \end{align*}
# $$
#
# Here,
# * $p_g(A_i)$ represents generative baseline of $A_i$
# * Context $X$ represents all the ($A, B, X$) triads, i.e., ($A_1, B_1| X_1$),..., ($A_n, B_n| X_n$)
# * $k$ is the number of triad instances where $X$ closed with a plus edges
#
# Receptive surprise is similar, just use $p_r(A_i)$. So, the formula for `Receptive Surprise` of context X is:
# $$
# \begin{align*}
# s_r(X) &= \frac{k - \sum_{i=1}^{n} p_r(A_i)}{\sqrt{\sum_{i=1}^{n} p_r(A_i) * (1 - p_r(A_i))}}\\
# \end{align*}
# $$
#
# Here, $p_r(A_i)$ represents receptive baseline of $A_i$.
# <font color="blue">6. Compute “Rec. Surprise” for all each of the 16 types.</font>
# +
# G_slash_dir = nx.read_edgelist("data/tmp.txt", nodetype=int, data=(("sign", int),), edgetype=int, comments='#', create_using=nx.DiGraph)
# store K value for each type
K_r = {1: 0, 2: 0, 3: 0, 4: 0, 5: 0, 6: 0, 7: 0, 8: 0, 9: 0, 10: 0, 11: 0, 12: 0, 13: 0, 14: 0, 15: 0, 16: 0}
# store P_r value for each type
P_r = {1: 0.0, 2: 0.0, 3: 0.0, 4: 0.0, 5: 0.0, 6: 0.0, 7: 0.0, 8: 0.0, 9: 0.0, 10: 0.0, 11: 0.0, 12: 0.0, 13: 0.0, 14: 0.0, 15: 0.0, 16: 0.0}
# store Denominator value for each type's S_g formula, needs to make square-root over this value
Denominator_r = {1: 0.0, 2: 0.0, 3: 0.0, 4: 0.0, 5: 0.0, 6: 0.0, 7: 0.0, 8: 0.0, 9: 0.0, 10: 0.0, 11: 0.0, 12: 0.0, 13: 0.0, 14: 0.0, 15: 0.0, 16: 0.0}
"""
Triad type matcher on graph "G_slash_dir".
This function assumes there is a directed edge from node "a" to node "b".
attributes
==========
a node-id
b node-id
x node-id
"""
def update_rec_dictionary(a, b, x, type_):
if G_slash_dir[a][b]['sign'] == 1:
K_r[type_] += 1
in_deg = G_slash_dir.in_degree(a)
pos_in_deg = len([(u,v) for u,v,e in G_slash_dir.in_edges(a, data=True) if e['sign'] == 1])
# print("Node-id: {}, Pos-InEdges: {}".format(a, [(u,v) for u,v,e in G_slash_dir.in_edges(data=True) if e['sign'] == 1]))
# print("Pos-InDeg: {}, InDeg: {}".format(pos_in_deg, in_deg))
if in_deg != 0:
P_r[type_] += (pos_in_deg/float(in_deg))
Denominator_r[type_] += ((pos_in_deg/float(in_deg)) * (1 - (pos_in_deg/float(in_deg))))
def print_rec_surprise():
for type_ in range(1, 17):
rec_surp = 0.0
if Denominator_r[type_] != 0.0:
rec_surp = K_r[type_] - P_r[type_] / math.sqrt(Denominator_r[type_])
# else:
# print("Type-{} is not covered. K: {}, P_r: {}, Denominator: {}".format(type_, K_r[type_], P_r[type_], Denominator_r[type_]))
print("Receptive Surprise for type-{}: {}".format(type_, rec_surp))
def calculate_rec_surprise():
triad_gen = nx.all_triads(G_slash_dir)
triad_nums = 0
for triad in triad_gen:
triad_nodes = list(triad.nodes())
# print(type(triad_nodes))
# print(triad_nodes)
triad_nums += 1
if triad_nums % 100000 == 0:
print("Finished {} triads.".format(triad_nums))
if triad_nums >= 10000000:
break
if G_slash_dir.has_edge(triad_nodes[0], triad_nodes[1]):
types = triad_type_matcher(triad_nodes[0], triad_nodes[1], triad_nodes[2])
# print("a: {} to b: {} got: {}".format(triad_nodes[0], triad_nodes[1], types))
for type_ in types:
update_rec_dictionary(triad_nodes[0], triad_nodes[1], triad_nodes[2], type_)
if G_slash_dir.has_edge(triad_nodes[1], triad_nodes[0]):
types = triad_type_matcher(triad_nodes[1], triad_nodes[0], triad_nodes[2])
# print("a: {} to b: {} got: {}".format(triad_nodes[1], triad_nodes[0], types))
for type_ in types:
update_rec_dictionary(triad_nodes[1], triad_nodes[0], triad_nodes[2], type_)
if G_slash_dir.has_edge(triad_nodes[1], triad_nodes[2]):
types = triad_type_matcher(triad_nodes[1], triad_nodes[2], triad_nodes[0])
# print("a: {} to b: {} got: {}".format(triad_nodes[1], triad_nodes[2], types))
for type_ in types:
update_rec_dictionary(triad_nodes[1], triad_nodes[2], triad_nodes[0], type_)
if G_slash_dir.has_edge(triad_nodes[2], triad_nodes[1]):
types = triad_type_matcher(triad_nodes[2], triad_nodes[1], triad_nodes[0])
# print("a: {} to b: {} got: {}".format(triad_nodes[2], triad_nodes[1], types))
for type_ in types:
update_rec_dictionary(triad_nodes[2], triad_nodes[1], triad_nodes[0], type_)
if G_slash_dir.has_edge(triad_nodes[0], triad_nodes[2]):
types = triad_type_matcher(triad_nodes[0], triad_nodes[2], triad_nodes[1])
# print("a: {} to b: {} got: {}".format(triad_nodes[0], triad_nodes[2], types))
for type_ in types:
update_rec_dictionary(triad_nodes[0], triad_nodes[2], triad_nodes[1], type_)
if G_slash_dir.has_edge(triad_nodes[2], triad_nodes[0]):
types = triad_type_matcher(triad_nodes[2], triad_nodes[0], triad_nodes[1])
# print("a: {} to b: {} got: {}".format(triad_nodes[2], triad_nodes[0], types))
for type_ in types:
update_rec_dictionary(triad_nodes[2], triad_nodes[0], triad_nodes[1], type_)
calculate_rec_surprise()
print_rec_surprise()
# -
# ## 2. The SIR Model of Disease Spreading [50 points]
#
# In this question, we will explore how varying the set of initially infected nodes in the SIR model can affect how a contagion spreads through a network.
#
# For the `2005 Graph Drawing` conference, a data set was provided of the IMDB movie database. We will use a reduced version of this dataset, which derived all actor-actor collaboration edges where the actors co-starred in at least $2$ movies together between $1995$ and $2004$. The following files are in the `data` directory:
# * [imdb_actor_edges.tsv]()
# * [imdb_actors_keys.tsv]()
#
# We will be comparing our results to two other null models, the `Erdos-Renyi` graph and the `Preferential Attachment` graph, with the same number of nodes and expected degree. The following files are in the `data` directory:
# * [SIR_erdos_renyi.txt]()
# * [SIR_preferential_attachment.txt]()
#
# Under the SIR model, every node can be either `susceptible`, `infected`, or `recovered` and every node starts as either `susceptible` or `infected`. Every infected neighbor of a susceptible node infects the `susceptible node` with probability $β$, and `infected nodes` can recover with probability $δ$. `Recovered nodes` are no longer susceptible and cannot be infected again. In the problem statement, $Algorithm 1$ describes for pseudo-code of this process.
#
# 1. For a node with $d$ neighbors, we need to find the probability of getting infected in a given round.
# 2. Need to implement the SIR model (described above) and run $100$ simulations with $β = 0.05$ and $δ = 0.5$ for each of the three graphs (e.g. imdb, erdos-renyi, and preferential attachment). Initialize the infected set with a single node chosen uniformly at random. Record the total percentage of nodes that became `infected` in each simulation. Note that a simulation ends when there are no more infected nodes; the total percentage of nodes that became infected at some point is thus the number of `recovered nodes` at the end of your simulation divided by the total number of nodes in the network.
#
# Some simulations may die out very quickly as not able to create an epidemic in the netwotk. While others may become epidemics and infect a large proportion of the networks, and thus may need a longer simulation time. For all three graphs if the proportion of simulations that infected at least $50%$ of the network; we will consider these events as `epidemics`. To compare the likelihood of an epidemic starting across graphs, and more importantly, test whether or not the observed differences are actually significant, we will use pairwise `Chi-Square tests`. For each pair of networks, compute:
#
# $scipy.stats.chi2 contingency([[e_1, 100-e_1],[e_2, 100-e_2]])$
#
# where $e_1$ is the number of trials where more than $50%$ were infected in $network 1$ and $e_2$ is the number of trials where more than $50%$ were infected in $network 2$. We need to report both the $χ2-statistic$ and $p-values$. See the problem statement for details on interpreting the output of the function call.
#
# Finally, we like to answer the following questions about the two synthetic networks:
# * <font color="blue">Does the `Erdos-Renyi` graph appear to be more/less susceptible to epidemics than the `Preferential Attachment` graph?</font>
# * <font color="blue">In cases where an epidemic does take off, does `Erdos-Renyi` graph appear to have higher/lower final percentage infected?</font>
# * <font color="blue">Overall, which of these two networks seems to be more susceptible to the spread of disease?</font>
# * <font color="blue">Give one good reason why we might expect to see these significant differences (or lack thereof) between `Erdos-Renyi` and `Preferential Attachment`? (2–3 sentences).</font>
#
# For further analysis on different network, I highly encourage to first try with a smaller number of simulations and only run with $100$ simulations once you are confident that this code works fine for your graph. Running $100$ simulations is necessary to ensure statistical significance in some of the comparisons.
def SIR_simulation(G, beta, delta):
# declaration of required data-structures
# susceptible, infected, and recovered nodes
S = set()
I = set()
R = set()
# list of all the nodes
nodes = []
for n in list(G.nodes):
nodes.append(int(n))
S.add(n)
num_nodes = len(nodes)
# print(num_nodes)
# print(nodes)
# choosing the initial single infected node randomly
initial_infected_idx = random.randint(0, num_nodes)
initial_infected_node = nodes[initial_infected_idx]
# initializing infected set
I.add(initial_infected_node)
# removing the initial infected node from the susceptible list
S.remove(initial_infected_node)
while len(I) > 0:
# declaration of required data-structures
S_ = set()
I_ = set()
J_ = set()
R_ = set()
for u in nodes:
if u in S:
for v in G.neighbors(u):
if v in I:
toss = random.random()
if toss <= beta:
S_.add(u)
I_.add(u)
break
elif u in I:
toss = random.random()
if toss <= delta:
J_.add(u)
R_.add(u)
S = S - S_
I = (I | I_) - J_
R = (R | R_)
return len(R)/float(num_nodes)
# +
import random
from random import randrange
random.seed()
beta = 0.05
delta = 0.5
# +
G_imdb = nx.read_edgelist("data/imdb_actor_edges.tsv", nodetype=int, data=(('weight', int),), comments='#', create_using=nx.Graph)
G_imdb_sir = []
for itr in range(100):
infected_ratio = SIR_simulation(G_imdb, beta, delta)
G_imdb_sir.append(infected_ratio)
print("Iteration: {} infected ratio: {}".format(itr, infected_ratio))
# +
G_erdos = nx.read_edgelist("data/SIR_erdos_renyi.txt", nodetype=int, comments='#', create_using=nx.Graph)
G_erdos_sir = []
for itr in range(100):
infected_ratio = SIR_simulation(G_erdos, beta, delta)
G_erdos_sir.append(infected_ratio)
print("Iteration: {} infected ratio: {}".format(itr, infected_ratio))
# +
G_pref = nx.read_edgelist("data/SIR_preferential_attachment.txt", nodetype=int, comments='#', create_using=nx.Graph)
G_pref_sir = []
for itr in range(100):
infected_ratio = SIR_simulation(G_pref, beta, delta)
G_pref_sir.append(infected_ratio)
print("Iteration: {} infected ratio: {}".format(itr, infected_ratio))
# +
# print(G_erdos_sir)
# print(G_pref_sir)
# print(G_imdb_sir)
import scipy
from scipy.stats import chi2_contingency
# scipy.__version__
e_imdb = len([i for i in G_imdb_sir if i >= 0.5])
e_erdos = len([i for i in G_erdos_sir if i >= 0.5])
e_pref = len([i for i in G_pref_sir if i >= 0.5])
print("e_imdb: {}".format(e_imdb))
print("e_erdos: {}".format(e_erdos))
print("e_pref: {}".format(e_pref))
imdb_vs_erdos_obs = np.array([[e_imdb, (100-e_imdb)], [e_erdos, (100-e_erdos)]])
imdb_vs_erdos_contingent = chi2_contingency(imdb_vs_erdos_obs)
print("imdb Vs. erdos contingent: {}".format(imdb_vs_erdos_contingent))
imdb_vs_pref_obs = np.array([[e_imdb, (100-e_imdb)], [e_pref, (100-e_pref)]])
imdb_vs_pref_contingent = chi2_contingency(imdb_vs_pref_obs)
print("imdb Vs. pref contingent: {}".format(imdb_vs_pref_contingent))
erdos_vs_pref_obs = np.array([[e_erdos, (100-e_erdos)], [e_pref, (100-e_pref)]])
erdos_vs_pref_contingent = chi2_contingency(erdos_vs_pref_obs)
print("erdos Vs. pref contingent: {}".format(erdos_vs_pref_contingent))
# +
def plot_degree_dist(G, plt_title):
m=3
degree_freq = nx.degree_histogram(G)
degrees = range(len(degree_freq))
plt.figure(figsize=(12, 8))
plt.loglog(degrees[m:], degree_freq[m:],'go-')
plt.title(plt_title)
plt.xlabel('Degree')
plt.ylabel('Frequency')
plot_degree_dist(G_imdb, "Degree distribution (Log-Log) of imdb graph")
plot_degree_dist(G_erdos, "Degree distribution (Log-Log) of Erdos-Renyi graph")
plot_degree_dist(G_pref, "Degree distribution (Log-Log) of Preferential Attachment graph")
# +
import seaborn as sns
def plot_infection_spread(sir_data, plt_title):
plt.figure(figsize=(12, 8))
# seaborn histogram
sns.distplot(sir_data, hist=True, kde=False,
bins=int(1000/5), color = 'blue',
hist_kws={'edgecolor':'black'})
# # matplotlib histogram
# plt.hist(sir_data, color = 'blue', edgecolor = 'black',
# bins = int(100/5))
# Add labels
plt.title(plt_title)
plt.xlabel('Ratio of infected nodes')
plt.ylabel('# of trials')
plot_infection_spread(G_imdb_sir, 'Histogram of infection spread ratio in imdb graph')
plot_infection_spread(G_erdos_sir, 'Histogram of infection spread ratio in Erdos-Renyi graph')
plot_infection_spread(G_pref_sir, 'Histogram of infection spread ratio in Preferential Attachment graph')
# +
cc_imdb = nx.number_connected_components(G_imdb)
cc_erdos = nx.number_connected_components(G_erdos)
cc_pref = nx.number_connected_components(G_pref)
print("Connected components in imdb graph: {}".format(cc_imdb))
print("Connected components in Erdos-Renyi graph: {}".format(cc_erdos))
print("Connected components in Preferential Attachment graph: {}".format(cc_pref))
# -
# To further analyzing the SIR model results and the susceptibility to epidemics, we further plotted the degree distribution and infection spread ratio of the three network. Let's first plot the occurance of epidemics (more than 50% node infected) in the networks within the 100 simulations:
#
# <table>
# <tr>
# <th>Graph</th>
# <th># of epidemics (in 100 simulations)</th>
# </tr>
# <tr>
# <td>imdb</td>
# <td>56</td>
# </tr>
# <tr>
# <td>Erdos-Renyi</td>
# <td>68</td>
# </tr>
# <tr>
# <td>Preferential Attachment</td>
# <td>73</td>
# </tr>
# <tr>
# <td colspan="2" align="middle"><b>Table 4: Number of epidemics (in 100 simulations)</b></td>
# </tr>
# </table>
#
# To compare the likelihood of an epidemic starting across graphs, and more importantly, to understand whether or not the observed differences are actually significant, we used pairwise Chi-Square tests. Here is the $χ²$ (i.e., test statistic) and p-value result we observed from this test:
#
# <table>
# <tr>
# <th>$e_1$</th>
# <th>$e_2$</th>
# <th>$χ²$ (test statistic)</th>
# <th>p-value</th>
# </tr>
# <tr>
# <td>imdb</td>
# <td>Erdos-Renyi</td>
# <td>2.57</td>
# <td>0.10</td>
# </tr>
# <tr>
# <td>imdb</td>
# <td>Preferential Attachment</td>
# <td>5.59</td>
# <td>0.02</td>
# </tr>
# <tr>
# <td>Erdos-Renyi</td>
# <td>Preferential Attachment</td>
# <td>0.38</td>
# <td>0.54</td>
# </tr>
# <tr>
# <td colspan="4" align="middle"><b>Table 5: chi2 contingency results</b></td>
# </tr>
# </table>
#
# Here the $χ²$ (i.e., `test statistic`) resembles a normalized sum of squared deviations of between two comparing network property (in our case, we considered the susceptibility to epidemics). The `p-value` is the probability of obtaining test results at least as extreme as the results actually observed. During this, it makes the assumption that the null hypothesis is correct.
# * <font color="blue">Does the `Erdos-Renyi` graph appear to be more/less susceptible to epidemics than the `Preferential Attachment` graph?</font>
# * <b>Answer:</b> From `Table-4` we can see, `Erdos-Renyi` graph appear to be less susceptible to epidemics than the `preferential attachment` graph. This is because, the `preferential attachment` graph build in a way so that the nodes with higher degree have higher probability to be attached to more nodes, thus, causing higher spreading of the contagion. We can observe this from the degree distribution we plotted earlier. However, there is no significant difference in epidemic likelihood.
# * <font color="blue">In cases where an epidemic does take off, does `Erdos-Renyi` graph appear to have higher/lower final percentage infected?</font>
# * <b>Answer:</b> From the histogram of infection spread ratio in Erdos-Renyi graph, we can see it appears to have higher final percentage infected node in cases where an epidemic does take off.
# * <font color="blue">Overall, which of these two networks seems to be more susceptible to the spread of disease?</font>
# * <b>Answer:</b> From the degree distribution plot, we can observe that the `preferential attachment` graph have a heavy tail, meaning it have more higher degree nodes comparing to `Erdos-Renyi` graph. Besides this, both of the graph have a single connected component. This implies, if a node is infected in the `preferential attachment` graph, it have a higher probability that it will infect the neighboring nodes quickly.
# * <font color="blue">Give one good reason why we might expect to see these significant differences (or lack thereof) between `Erdos-Renyi` and `Preferential Attachment`? (2–3 sentences).</font>
# * <b>Answer:</b> Both of the `Erdos-Renyi` and `Preferential Attachment` have lots of similarities that we can see from the degree distribution and the histogram plotting of infection spread ratio. It is worth observing that, while comparing the with the `imdb` graph through Chi-Square tests, it seems `Erdos-Renyi` graph is more susceptible to the spread of disease (as it gives lower $χ²$ with higher `p-value`). To compare the likelihood of an epidemic between `Erdos-Renyi` and `Preferential Attachment` graph, we further used pairwise Chi-Square tests in between them. From that test we got higher likelihood of an epidemic in between them.
# # Conclusion
#
# In the first part of this assignment, we have explored the `Signed` network. For `signed network`, we did experiment with [Slashdot](http://snap.stanford.edu/data/soc-sign-Slashdot081106.html) dataset and calculate the frequency of signed triads in both "real" and "shuffled" network to understand whether it hold structural balance. Later we explored `generative` and `receptive` surprise and computed those metric on the same dataset. It helped us to understand the deviation of users behavior from the baseline in context $X$.
#
# In the second part of this assignment, we have explored the `Disease Spreading` network and implemented the `SIR Model` to understand the likelihood of an epidemic in a given network. For this problem, we did experiment with three datasets, e.g. [imdb](), [erdos-renyi](), and [preferential-attachment](). We have done the pairwise `Chi-Square test` to compare the likelihood of an epidemic starting across graphs.
# # References
#
#
# [1] <NAME>, and <NAME>. 1960. “On the Evolution of Random Graphs.” Bull. Inst. Internat. Statis. 38 (4): 343–47.
#
# [2] NetworkX, “Software for Complex Networks,” https://networkx.github.io/documentation/stable/, 2020, accessed: 2020-10.
#
# [3] NetworkX, “Install NetworkX Documentation,” https://networkx.github.io/documentation/stable/install.html, 2020, accessed: 2020-10.
| assignments/assignment_2/assignment_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Spyder)
# language: python3
# name: python3
# ---
# # Module 3.5 Loops, Summarized
#
# Created By: <NAME> from http://learningdata.io
#
# Each code block is designed to be an independent program for ease of use!
#
# ---
#
# ***Disclaimer***
#
# > Copyright (c) 2020 <NAME>
#
# > Permission is hereby granted, free of charge, to any person obtaining a copy of this software and associated documentation files (the "Software"), to deal in the Software without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Software, and to permit persons to whom the Software is furnished to do so, subject to the following conditions:
#
# > The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Software.
#
# > THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
# ## Part 1: For Loops
#
# For loops in Python allow us to run an iterative chunk of code across an array of items. This could be in a list, set, dictionary, or even just characters in a string.
#
# ```python
# my_list = [1,2,3]
#
# for item in my_list:
# # ###
# # This code is executed on each item in my list
# # ###
# ```
#
# The placeholder `item` can be replaed with anything. For numbers, we often use `i` or `n` to denote significance.
#
# The for loop creates a local variable `item` that is able to be used during the execution of the single loop.
# ### Print Out All Values in a List
# +
items = ["Apples", "Grapes", "Avocados", "Cucumbers"]
for item in items:
print(item)
# -
# ### For Loops with Conditional Logic
# +
numbers = [0,1,2,3,4,5,6,7,8,9]
for n in numbers:
if (n%2==0):
print(n)
# -
# ## Part 2: While Loops
#
# While loops execute a block of code repeatedly until a conditional statement evaluates `False`
#
# Be careful to note code an infinite loop! There needs to be a stopping condition.
#
# ```python
# while (True):
# # ###
# # This Block of Code Iterates
# # ###
# # And part of the code needs to progress towards or trigger a stopping condition
# # ###
# else:
# # ###
# # Optional code block that evaluates upon stopping the loop
# # ###
# ```
# ### Simple Countdown
# +
i = 10
while (i > 0):
print(i)
i -= 1
else:
print("Blast Off!")
# -
# ### Process & Remove Elements from List
#
# Tip: If a list contains elements, it evaluates `True`
# +
shopping_list = ["Green Onion", "Tortillas", "Avocados", "Black Beans", "Paper Towels"]
while (shopping_list):
item = shopping_list.pop()
print("Adding {} to your Amazon Cart.".format(item))
else:
print("Ready to Checkout!")
# -
# ## There is a lot more to loops!
#
# We can also integrate these within our functions to process an input, or call a function within our loop for each element!
| modules/module-03/module3-loops.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Sensors in SMRT
# ===========
#
# __Goal__:
# - plot the diagram of thermal emission + backscattering coefficient from a simple snowpack at 13 GHz
# - plot the diagram of thermal emission from a simple snowpack at multiple frequencies (e.g. 19 and 37 GHz)
#
# __Learning__: Sensor and Result object
#
# For the snowpack, you can take the following properties:
# - thickness = 1000 (means semi-infinite)
# - sticky_hard_spheres microstructure model
# - radius = 100e-6
# - density = 300
# - temperature = 260
# - stickiness = 0.15
#
# The following imports are valid for both excercices:
#
#
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib notebook
from smrt import make_model, make_snowpack, sensor_list
from smrt.utils import dB
# -
# Emission / backscatter diagram
# -----------------------------------
# prepare the snowpack
radius = 100e-6
density=300
temperature = 260
sp = make_snowpack(thickness=[1000], microstructure_model='sticky_hard_spheres',
density=density, radius=radius, temperature=temperature,
stickiness=0.15)
# prepare a list for theta from 5 to 65 by step of 5
theta = [5, 10, 15, 20, 25, 30, 35, 40, 45, 50, 55, 60, 65]
# prepare two sensors (one active, on passive) at 13 GHz
radiometer = sensor_list.passive(13e9, theta)
radar = sensor_list.active(13e9, theta)
# prepare the model and run it successively for each sensor
m = make_model("iba", "dort")
res_a = m.run(radar, sp)
res_p = m.run(radiometer, sp)
# +
# for plotting two side by side graphs, the best solution is:
f, axs = plt.subplots(1, 2, figsize=(8, 3.5))
# plot on left graph
axs[0].plot(theta, res_a.sigmaVV()) # adapt x and y to your need
# plot on right graph
#axs[1].plot(x, y) # adapt x and y to your need
# to set axis labels:
axs[0].set_xlabel("Viewing angle")
# ...
# -
# multi-frequency emission diagram
# -----------------------------------
# +
# prepare 1 sensor object with two frequencies
# +
# prepare the model and run it
# +
# plot the results on a single graph
# see results documentation for selecting by frequency
# http://smrt.readthedocs.io/en/latest/smrt.core.result.html
# -
| 02_using_smrt/01_sensor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Answer Key to the Data Wrangling with Spark SQL Quiz
#
# This quiz uses the same dataset and most of the same questions from the earlier "Quiz - Data Wrangling with Data Frames Jupyter Notebook." For this quiz, however, use Spark SQL instead of Spark Data Frames.
#
# Helpful resources:
# http://spark.apache.org/docs/latest/api/python/pyspark.sql.html
# +
import findspark
findspark.init()
from pyspark.sql import SparkSession
# from pyspark.sql.functions import isnan, count, when, col, desc, udf, col, sort_array, asc, avg
# from pyspark.sql.functions import sum as Fsum
# from pyspark.sql.window import Window
# from pyspark.sql.types import IntegerType
# +
# 1) import any other libraries you might need
# 2) instantiate a Spark session
# 3) read in the data set located at the path "data/sparkify_log_small.json"
# 4) create a view to use with your SQL queries
# 5) write code to answer the quiz questions
spark = SparkSession \
.builder \
.appName("Spark SQL Quiz") \
.getOrCreate()
user_log = spark.read.json("../data/sparkify_log_small.json")
user_log.createOrReplaceTempView("log_table")
# -
# # Question 1
#
# Which page did user id "" (empty string) NOT visit?
user_log.printSchema()
# SELECT distinct pages for the blank user and distinc pages for all users
# Right join the results to find pages that blank visitor did not visit
spark.sql("""
SELECT *
FROM (
SELECT DISTINCT page
FROM log_table
WHERE userID='') AS user_pages
RIGHT JOIN (
SELECT DISTINCT page
FROM log_table) AS all_pages
ON user_pages.page = all_pages.page
WHERE user_pages.page IS NULL
""").show()
# # Question 2 - Reflect
#
# Why might you prefer to use SQL over data frames? Why might you prefer data frames over SQL?
#
# Both Spark SQL and Spark Data Frames are part of the Spark SQL library. Hence, they both use the Spark SQL Catalyst Optimizer to optimize queries.
#
# You might prefer SQL over data frames because the syntax is clearer especially for teams already experienced in SQL.
#
# Spark data frames give you more control. You can break down your queries into smaller steps, which can make debugging easier. You can also [cache](https://unraveldata.com/to-cache-or-not-to-cache/) intermediate results or [repartition](https://hackernoon.com/managing-spark-partitions-with-coalesce-and-repartition-4050c57ad5c4) intermediate results.
# # Question 3
#
# How many female users do we have in the data set?
spark.sql("""
SELECT COUNT(DISTINCT userID)
FROM log_table
WHERE gender = 'F'
""").show()
# # Question 4
#
# How many songs were played from the most played artist?
# +
# Here is one solution
spark.sql("""
SELECT Artist, COUNT(Artist) AS plays
FROM log_table
GROUP BY Artist
ORDER BY plays DESC
LIMIT 1
""").show()
# Here is an alternative solution
# Get the artist play counts
play_counts = spark.sql("""
SELECT Artist, COUNT(Artist) AS plays
FROM log_table
GROUP BY Artist
""")
# save the results in a new view
play_counts.createOrReplaceTempView("artist_counts")
# use a self join to find where the max play equals the count value
spark.sql("""
SELECT a2.Artist, a2.plays
FROM (
SELECT max(plays) AS max_plays
FROM artist_counts
) AS a1
JOIN artist_counts AS a2
ON a1.max_plays = a2.plays
""").show()
# -
# # Question 5 (challenge)
#
# How many songs do users listen to on average between visiting our home page? Please round your answer to the closest integer.
#
#
# +
# SELECT CASE WHEN 1 > 0 THEN 1 WHEN 2 > 0 THEN 2.0 ELSE 1.2 END;
is_home = spark.sql("""
SELECT userID, page, ts, CASE WHEN page = 'Home' THEN 1 ELSE 0 END AS is_home
FROM log_table
WHERE (page = 'NextSong') or (page = 'Home')
""")
# keep the results in a new view
is_home.createOrReplaceTempView("is_home_table")
# find the cumulative sum over the is_home column
cumulative_sum = spark.sql("""
SELECT *, SUM(is_home) OVER
(PARTITION BY userID ORDER BY ts DESC ROWS BETWEEN UNBOUNDED PRECEDING AND CURRENT ROW) AS period
FROM is_home_table""")
# keep the results in a view
cumulative_sum.createOrReplaceTempView("period_table")
# find the average count for NextSong
spark.sql("""
SELECT AVG(count_results)
FROM (
SELECT COUNT(*) AS count_results
FROM period_table
GROUP BY userID, period, page
HAVING page = 'NextSong') AS counts
""").show()
# -
| Data-Lake/notebooks/9_spark_sql_quiz.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mrksntndr/Linear-Algebra-58019/blob/main/Vectors.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="oBFbGmYV9eHS" outputId="975af7bb-4e08-4f82-e12d-de66d77db628"
#numpy
import numpy as np
a = np.array([1,2,3])
print(a)
# + colab={"base_uri": "https://localhost:8080/"} id="os2rf0qT-kxI" outputId="1d4f9e23-fecc-4be0-bffe-537ea3e329ef"
import numpy as np
b = np.array([[1,2,3],[4,5,6]])
print(b)
# + colab={"base_uri": "https://localhost:8080/"} id="ixl1HEFg-m6a" outputId="2659ffb3-bede-43f8-8b62-68fb13652749"
import numpy as np
c = np.array([[1,2,3],[4,5,6],[7,8,9]])
print(c)
# + colab={"base_uri": "https://localhost:8080/"} id="GrNXjiwL-pgv" outputId="398a2b44-9dfd-463a-b6ae-58fbb413829e"
import numpy as np
d = np.full ((3,3),7)
print(d)
# + colab={"base_uri": "https://localhost:8080/"} id="EeHaSsMD-qw1" outputId="0b216c22-49df-40ed-ba4a-452a1d0b5123"
import numpy as np
e = np.array([[1,2,3],[4,5,6],[7,8,9]])
print(e)
e = np.diagonal([[1,2,3],[4,5,6],[7,8,9]])
print(e)
# + colab={"base_uri": "https://localhost:8080/"} id="0OeACQKo-sjr" outputId="45d66439-43ef-4e7b-922a-a3f59cd80f9e"
import numpy as np
f = np.eye(3)
print(f)
# + colab={"base_uri": "https://localhost:8080/"} id="gMVdYW-u-uwD" outputId="a4ca8a51-af20-444c-8e98-a82368849c67"
import numpy as np
g = np.zeros((3,3))
print(g)
# + colab={"base_uri": "https://localhost:8080/"} id="wrJ26gWm-vkm" outputId="882da873-930d-4455-899a-5fae24be8ff0"
import numpy as np
h = np.empty((0,12))
print(h)
| Vectors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
##imports
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from interpret.glassbox import ExplainableBoostingClassifier
# +
valid_Xs = pd.read_csv("../preprocessed_data/valid_Xs.csv")
valid_ys = pd.read_csv("../preprocessed_data/valid_ys.csv")
train_Xs = pd.read_csv("../preprocessed_data/train_Xs.csv")
train_ys = pd.read_csv("../preprocessed_data/train_ys.csv")
def relevel_gd(df):
df.loc[df['goal_diff']>3, 'goal_diff'] = 3
df.loc[df['goal_diff']<-3, 'goal_diff'] = -3
return df
valid_Xs = relevel_gd(valid_Xs)
train_Xs = relevel_gd(train_Xs)
# -
train_Xs.head()
# +
# %%time
models = {}
for col in ["scoring", "conceding"]:
ebm = ExplainableBoostingClassifier(interactions=3, random_state=42, validation_size=0.2)
ebm.fit(train_Xs, train_ys[col])
models[col] = ebm
# +
scoring_probs = models["scoring"].predict_proba(valid_Xs)[:, 1]
conceding_probs = models["conceding"].predict_proba(valid_Xs)[:, 1]
vaep_values = scoring_probs - conceding_probs
# -
with plt.style.context("ggplot"):
fig, ax = plt.subplots(figsize=(12, 8))
ax.hist(vaep_values, bins=26, ec="k", fc="xkcd:salmon")
ax.set(xlabel="VAEP Values", ylabel="Frequency", title="Histogram of Derived VAEP Values")
fig.savefig("vaep_values", dpi=150)
# +
from interpret import show
ebm_global = ebm.explain_global()
show(ebm_global)
# -
ebm.explain_global()
# + [markdown] tags=[]
# #### Model Evaluation
# +
from sklearn.metrics import brier_score_loss
model_name = "scoring"
predicted_probs = models[model_name].predict_proba(valid_Xs)[:, 1]
baseline_preds = np.ones(len(valid_ys))*valid_ys[model_name].mean() ##according to the paper predicting all values as the class distribution
baseline_brier_score = brier_score_loss(valid_ys[model_name].values, baseline_preds)
print(f"Baseline Prediction Brier Score: {baseline_brier_score:.5f}")
model_brier_score = brier_score_loss(valid_ys[model_name].values, predicted_probs)
print(f"Model Brier Score: {model_brier_score:.5f}")
normalized_brier_score_loss = model_brier_score/baseline_brier_score
print(f"Normalized Brier Score: {normalized_brier_score_loss:.5f}") ##lower=better; 0 = perfect, >1 equals bogus, <1 is okay
# +
from interpret.blackbox import PartialDependence
pdp = PartialDependence(predict_fn=ebm.predict_proba, data=train_Xs)
pdp_global = pdp.explain_global(name='Partial Dependence')
show(pdp_global)
# -
from xgboost import XGBClassifier
xgb_model = XGBClassifier(random_state=42).fit(train_Xs.values, train_ys["scoring"].values)
# +
from sklearn.metrics import brier_score_loss
model_name = "scoring"
predicted_probs = xgb_model.predict_proba(valid_Xs.values)[:, 1]
baseline_preds = np.ones(len(valid_ys))*valid_ys[model_name].mean() ##according to the paper predicting all values as the class distribution
baseline_brier_score = brier_score_loss(valid_ys[model_name].values, baseline_preds)
print(f"Baseline Prediction Brier Score: {baseline_brier_score:.5f}")
model_brier_score = brier_score_loss(valid_ys[model_name].values, predicted_probs)
print(f"Model Brier Score: {model_brier_score:.5f}")
normalized_brier_score_loss = model_brier_score/baseline_brier_score
print(f"Normalized Brier Score: {normalized_brier_score_loss:.5f}") ##lower=better; 0 = perfect, >1 equals bogus, <1 is okay
# -
| vaep_gam_implementation/notebooks/modelling_interactML.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Graph4NLP Demo: Math Word Problem
#
# ---
#
# In this demo, we will have a closer look at how to apply **Graph2Tree model to the task of math word problem automatically solving**.
# Math word problem solving aims to infer reasonable equations from given natural language problem descriptions. It is important for exploring automatic solutions to mathematical problems and improving the reasoning ability of neural networks.
# In this demo, we use the Graph4NLP library to build a GNN-based math word problem (MWP) solving model.
#
# The **Graph2Tree** model consists of:
#
# - graph construction module (e.g., node embedding based dynamic graph)
# - graph embedding module (e.g., undirected GraphSage)
# - predictoin module (e.g., tree decoder with attention and copy mechanisms)
#
# As shown in the picture below, we firstly construct graph input from problem description by syntactic parsing (CoreNLP) and then represent the output equation with a hierarchical structure (Node ``N`` stands for non-terminal node).
#
# <p align="center">
# <img src="./imgs/g2t.png" width="600" class="center" alt="graph2tree_mwp"/>
# <br/>
# </p>
#
# We will use the built-in Graph2Tree model APIs to build the model, and evaluate it on the Mawps dataset.
# ## Environment setup
# ---
#
# Please follow the instructions [here](https://github.com/graph4ai/graph4nlp_demo#environment-setup) to set up the environment. Please also run the following commands to install extra packages used in this demo.
# ```
# pip install sympy
# pip install ipywidgets
# ```
#
#
# This notebook was tested on :
#
# ```
# torch == 1.9.0
# torchtext == 0.10.0
# ```
# ## Load the config file
# +
from graph4nlp.pytorch.modules.config import get_basic_args
from graph4nlp.pytorch.modules.utils.config_utils import update_values, get_yaml_config
def get_args():
config = {'dataset_yaml': "./config.yaml",
'learning_rate': 1e-3,
'gpuid': -1,
'seed': 123,
'init_weight': 0.08,
'graph_type': 'static',
'weight_decay': 0,
'max_epochs': 20,
'min_freq': 1,
'grad_clip': 5,
'batch_size': 20,
'share_vocab': True,
'pretrained_word_emb_name': None,
'pretrained_word_emb_url': None,
'pretrained_word_emb_cache_dir': ".vector_cache",
'checkpoint_save_path': "./checkpoint_save",
'beam_size': 4
}
our_args = get_yaml_config(config['dataset_yaml'])
template = get_basic_args(graph_construction_name=our_args["graph_construction_name"],
graph_embedding_name=our_args["graph_embedding_name"],
decoder_name=our_args["decoder_name"])
update_values(to_args=template, from_args_list=[our_args, config])
return template
# show our config
cfg_g2t = get_args()
from pprint import pprint
pprint(cfg_g2t)
# +
import copy
import torch
import random
import argparse
import numpy as np
import torch.optim as optim
from torch.utils.data import DataLoader
from tqdm.notebook import tqdm
from graph4nlp.pytorch.data.data import to_batch
from graph4nlp.pytorch.datasets.mawps import MawpsDatasetForTree
from graph4nlp.pytorch.modules.graph_construction import DependencyBasedGraphConstruction
from graph4nlp.pytorch.modules.graph_embedding import *
from graph4nlp.pytorch.models.graph2tree import Graph2Tree
from graph4nlp.pytorch.modules.utils.tree_utils import Tree
from utils import convert_to_string, compute_tree_accuracy, prepare_oov
# -
class Mawps:
def __init__(self, opt=None):
super(Mawps, self).__init__()
self.opt = opt
seed = self.opt["seed"]
random.seed(seed)
np.random.seed(seed)
torch.manual_seed(seed)
if self.opt["gpuid"] == -1:
self.device = torch.device("cpu")
else:
self.device = torch.device("cuda:{}".format(self.opt["gpuid"]))
self.use_copy = self.opt["decoder_args"]["rnn_decoder_share"]["use_copy"]
self.use_share_vocab = self.opt["graph_construction_args"]["graph_construction_share"]["share_vocab"]
self.data_dir = self.opt["graph_construction_args"]["graph_construction_share"]["root_dir"]
self._build_dataloader()
self._build_model()
self._build_optimizer()
def _build_dataloader(self):
para_dic = {'root_dir': self.data_dir,
'word_emb_size': self.opt["graph_construction_args"]["node_embedding"]["input_size"],
'topology_builder': DependencyBasedGraphConstruction,
'topology_subdir': self.opt["graph_construction_args"]["graph_construction_share"]["topology_subdir"],
'edge_strategy': self.opt["graph_construction_args"]["graph_construction_private"]["edge_strategy"],
'graph_type': 'static',
'dynamic_graph_type': self.opt["graph_construction_args"]["graph_construction_share"]["graph_type"],
'share_vocab': self.use_share_vocab,
'enc_emb_size': self.opt["graph_construction_args"]["node_embedding"]["input_size"],
'dec_emb_size': self.opt["decoder_args"]["rnn_decoder_share"]["input_size"],
'dynamic_init_topology_builder': None,
'min_word_vocab_freq': self.opt["min_freq"],
'pretrained_word_emb_name': self.opt["pretrained_word_emb_name"],
'pretrained_word_emb_url': self.opt["pretrained_word_emb_url"],
'pretrained_word_emb_cache_dir': self.opt["pretrained_word_emb_cache_dir"]
}
dataset = MawpsDatasetForTree(**para_dic)
self.train_data_loader = DataLoader(dataset.train, batch_size=self.opt["batch_size"], shuffle=True,
num_workers=0,
collate_fn=dataset.collate_fn)
self.test_data_loader = DataLoader(dataset.test, batch_size=1, shuffle=False, num_workers=0,
collate_fn=dataset.collate_fn)
self.valid_data_loader = DataLoader(dataset.val, batch_size=1, shuffle=False, num_workers=0,
collate_fn=dataset.collate_fn)
self.vocab_model = dataset.vocab_model
self.src_vocab = self.vocab_model.in_word_vocab
self.tgt_vocab = self.vocab_model.out_word_vocab
self.share_vocab = self.vocab_model.share_vocab if self.use_share_vocab else None
def _build_model(self):
'''For encoder-decoder'''
self.model = Graph2Tree.from_args(self.opt,
vocab_model=self.vocab_model)
self.model.init(self.opt["init_weight"])
self.model.to(self.device)
def _build_optimizer(self):
optim_state = {"learningRate": self.opt["learning_rate"], "weight_decay": self.opt["weight_decay"]}
parameters = [p for p in self.model.parameters() if p.requires_grad]
self.optimizer = optim.Adam(parameters, lr=optim_state['learningRate'], weight_decay=optim_state['weight_decay'])
def train_epoch(self, epoch):
loss_to_print = 0
num_batch = len(self.train_data_loader)
for step, data in tqdm(enumerate(self.train_data_loader), desc=f'Epoch {epoch:02d}', total=len(self.train_data_loader)):
batch_graph, batch_tree_list, batch_original_tree_list = data['graph_data'], data['dec_tree_batch'], data['original_dec_tree_batch']
batch_graph = batch_graph.to(self.device)
self.optimizer.zero_grad()
oov_dict = prepare_oov(
batch_graph, self.src_vocab, self.device) if self.use_copy else None
if self.use_copy:
batch_tree_list_refined = []
for item in batch_original_tree_list:
tgt_list = oov_dict.get_symbol_idx_for_list(item.strip().split())
tgt_tree = Tree.convert_to_tree(tgt_list, 0, len(tgt_list), oov_dict)
batch_tree_list_refined.append(tgt_tree)
loss = self.model(batch_graph, batch_tree_list_refined if self.use_copy else batch_tree_list, oov_dict=oov_dict)
loss.backward()
torch.nn.utils.clip_grad_value_(
self.model.parameters(), self.opt["grad_clip"])
self.optimizer.step()
loss_to_print += loss
return loss_to_print/num_batch
def train(self):
best_acc = -1
best_model = None
print("-------------\nStarting training.")
for epoch in range(1, self.opt["max_epochs"]+1):
self.model.train()
loss_to_print = self.train_epoch(epoch)
print("epochs = {}, train_loss = {:.3f}".format(epoch, loss_to_print))
if epoch > 15:
val_acc = self.eval(self.model, mode="val")
if val_acc > best_acc:
best_acc = val_acc
best_model = self.model
self.eval(best_model, mode="test")
best_model.save_checkpoint(self.opt["checkpoint_save_path"], "best.pt")
def eval(self, model, mode="val"):
model.eval()
reference_list = []
candidate_list = []
data_loader = self.test_data_loader if mode == "test" else self.valid_data_loader
for data in tqdm(data_loader, desc="Eval: "):
eval_input_graph, batch_tree_list, batch_original_tree_list = data['graph_data'], data['dec_tree_batch'], data['original_dec_tree_batch']
eval_input_graph = eval_input_graph.to(self.device)
oov_dict = prepare_oov(eval_input_graph, self.src_vocab, self.device)
if self.use_copy:
assert len(batch_original_tree_list) == 1
reference = oov_dict.get_symbol_idx_for_list(batch_original_tree_list[0].split())
eval_vocab = oov_dict
else:
assert len(batch_original_tree_list) == 1
reference = model.tgt_vocab.get_symbol_idx_for_list(batch_original_tree_list[0].split())
eval_vocab = self.tgt_vocab
candidate = model.translate(eval_input_graph,
oov_dict=oov_dict,
use_beam_search=True,
beam_size=self.opt["beam_size"])
candidate = [int(c) for c in candidate]
num_left_paren = sum(
1 for c in candidate if eval_vocab.idx2symbol[int(c)] == "(")
num_right_paren = sum(
1 for c in candidate if eval_vocab.idx2symbol[int(c)] == ")")
diff = num_left_paren - num_right_paren
if diff > 0:
for i in range(diff):
candidate.append(
self.test_data_loader.tgt_vocab.symbol2idx[")"])
elif diff < 0:
candidate = candidate[:diff]
ref_str = convert_to_string(
reference, eval_vocab)
cand_str = convert_to_string(
candidate, eval_vocab)
reference_list.append(reference)
candidate_list.append(candidate)
eval_acc = compute_tree_accuracy(
candidate_list, reference_list, eval_vocab)
print("{} accuracy = {:.3f}\n".format(mode, eval_acc))
return eval_acc
a = Mawps(cfg_g2t)
best_acc = a.train()
| IJCAI2021_demo/math_word_problem_solving/math_word_problem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="S-O5GRG6c0H5" colab_type="text"
# # Customer Life Time Value
# + id="Ing-T-CPiwm_" colab_type="code" colab={}
import pandas as pd
import matplotlib.pyplot as plt
# + id="QFwbRb-2iwnG" colab_type="code" outputId="de34d317-c8d5-42a4-d209-de4370a696cf" executionInfo={"status": "ok", "timestamp": 1575757677411, "user_tz": 240, "elapsed": 45009, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCGo6aIm0tOcd5EhqWlYb0rime9sBvHS9YMpx0D2w=s64", "userId": "08597265227091462140"}} colab={"base_uri": "https://localhost:8080/", "height": 202}
# Uncomment this line if using this notebook locally
#df = pd.read_excel('./data/retail/Online Retail.xlsx', sheet_name='Online Retail')
file_name = "https://raw.githubusercontent.com/rajeevratan84/datascienceforbusiness/master/OnlineRetail.xlsx"
df = pd.read_excel(file_name, sheet_name='Online Retail')
df.head()
# + id="pRzLcCMIiwnN" colab_type="code" outputId="6f26d1f3-d0e1-4e5a-a6d4-43e782a040f2" executionInfo={"status": "ok", "timestamp": 1575757677414, "user_tz": 240, "elapsed": 44997, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCGo6aIm0tOcd5EhqWlYb0rime9sBvHS9YMpx0D2w=s64", "userId": "08597265227091462140"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
df.shape
# + id="BrjkOFBdiwnT" colab_type="code" colab={}
# Remove quantities that are less than 0 (possibly returned items)
df = df.loc[df['Quantity'] > 0]
# + id="AmbYpkKQiwnW" colab_type="code" colab={}
# Remove blank customer IDs
df = df[pd.notnull(df['CustomerID'])]
# + id="LfQz7k1KiwnZ" colab_type="code" outputId="4d5a98b9-4f8b-4ce9-e3db-4714339102d6" executionInfo={"status": "ok", "timestamp": 1575757677421, "user_tz": 240, "elapsed": 44983, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCGo6aIm0tOcd5EhqWlYb0rime9sBvHS9YMpx0D2w=s64", "userId": "08597265227091462140"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
# View our data range
print('Date Range: %s to %s' % (df['InvoiceDate'].min(), df['InvoiceDate'].max()))
# + id="VfycwOcaiwqI" colab_type="code" colab={}
# taking all of the transactions that occurred before December 01, 2011
df = df.loc[df['InvoiceDate'] < '2011-12-01']
# + id="90GQlX-NiwqL" colab_type="code" colab={}
# Create a Sales Revenue Column
df['Sales'] = df['Quantity'] * df['UnitPrice']
# + id="btbKr9WpiwqQ" colab_type="code" colab={}
# Get a orders summary dataset that shows the total in sales made per customer invoice
orders_df = df.groupby(['CustomerID', 'InvoiceNo']).agg({'Sales': sum,'InvoiceDate': max})
# + id="fKLA_05TiwqS" colab_type="code" outputId="0140348e-4463-44a9-d349-3c1d87cd566a" executionInfo={"status": "ok", "timestamp": 1575757677959, "user_tz": 240, "elapsed": 45495, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCGo6aIm0tOcd5EhqWlYb0rime9sBvHS9YMpx0D2w=s64", "userId": "08597265227091462140"}} colab={"base_uri": "https://localhost:8080/", "height": 386}
orders_df.head(10)
# + id="0e73UEPRiwqV" colab_type="code" colab={}
# Create some simple functions we'll be using to create our summary dataframe
def groupby_mean(x):
return x.mean()
def groupby_count(x):
return x.count()
def purchase_duration(x):
return (x.max() - x.min()).days
def avg_frequency(x):
'''returns the average days between sales'''
return (x.max() - x.min()).days/x.count()
groupby_mean.__name__ = 'avg'
groupby_count.__name__ = 'count'
purchase_duration.__name__ = 'purchase_duration'
avg_frequency.__name__ = 'purchase_frequency'
summary_df = orders_df.reset_index().groupby('CustomerID').agg({
'Sales': [min, max, sum, groupby_mean, groupby_count],
'InvoiceDate': [min, max, purchase_duration, avg_frequency]
})
# + id="NuccuQqAiwqY" colab_type="code" outputId="16408745-f447-4876-a406-f2968aa43713" executionInfo={"status": "ok", "timestamp": 1575757681325, "user_tz": 240, "elapsed": 47826, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCGo6aIm0tOcd5EhqWlYb0rime9sBvHS9YMpx0D2w=s64", "userId": "08597265227091462140"}} colab={"base_uri": "https://localhost:8080/", "height": 417}
summary_df.head(10)
# + id="2XdZG2W-iwqc" colab_type="code" colab={}
# Rename our columns by combinng it with the top row
summary_df.columns = ['_'.join(col).lower() for col in summary_df.columns]
# + id="yThWPIDLiwqf" colab_type="code" outputId="4aef9da6-0386-4055-f217-2fb0b21a6ab0" executionInfo={"status": "ok", "timestamp": 1575757681327, "user_tz": 240, "elapsed": 16080, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCGo6aIm0tOcd5EhqWlYb0rime9sBvHS9YMpx0D2w=s64", "userId": "08597265227091462140"}} colab={"base_uri": "https://localhost:8080/", "height": 233}
summary_df.head()
# + [markdown] id="bvlk1j_Yd7bQ" colab_type="text"
# ## Visualization
# + id="5kZfIAxhjXXr" colab_type="code" outputId="06282ca5-a0a9-490a-d883-51894e9c5281" executionInfo={"status": "ok", "timestamp": 1575757749299, "user_tz": 240, "elapsed": 802, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCGo6aIm0tOcd5EhqWlYb0rime9sBvHS9YMpx0D2w=s64", "userId": "08597265227091462140"}} colab={"base_uri": "https://localhost:8080/", "height": 399}
# Let's look at the first 20 sales counts and the average spend for that sale
# We firstly group by Sales_Counts and then look at the sales_avg
summary_df.groupby('sales_count').count()['sales_avg'][:20]
# + id="akRUdf9iey4j" colab_type="code" outputId="a4a2abed-68e6-4d8a-ccb7-8b4d865ba3a7" executionInfo={"status": "ok", "timestamp": 1575757864043, "user_tz": 240, "elapsed": 673, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCGo6aIm0tOcd5EhqWlYb0rime9sBvHS9YMpx0D2w=s64", "userId": "08597265227091462140"}} colab={"base_uri": "https://localhost:8080/", "height": 446}
summary_df = summary_df.loc[summary_df['invoicedate_purchase_duration'] > 0]
summary_df
# + [markdown] id="AeqAnW1Tpemz" colab_type="text"
# Because we're trying to get the time between sales (plotted in the next cell). We need to eliminate single purchases. That's what this line does.
#
# `summary_df = summary_df.loc[summary_df['invoicedate_purchase_duration'] > 0]`
#
# Purchase durations that are equal to 0 indicate that only purchase was ever made.
# + id="pPtfdRgXiwqj" colab_type="code" outputId="87e74ca1-991d-4efa-979d-56594fe6ccdf" executionInfo={"status": "ok", "timestamp": 1575757750955, "user_tz": 240, "elapsed": 939, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCGo6aIm0tOcd5EhqWlYb0rime9sBvHS9YMpx0D2w=s64", "userId": "08597265227091462140"}} colab={"base_uri": "https://localhost:8080/", "height": 456}
# We filter on invoicedate_purchase_duration greater than 0
summary_df = summary_df.loc[summary_df['invoicedate_purchase_duration'] > 0]
ax = summary_df.groupby('sales_count').count()['sales_avg'][:20].plot(kind='bar', color='skyblue',figsize=(12,7), grid=True)
ax.set_ylabel('count')
plt.show()
# + id="LPv2_UvUiwqm" colab_type="code" outputId="2957fb65-99b1-4e90-8664-367634788f1f" executionInfo={"status": "ok", "timestamp": 1574033174406, "user_tz": 0, "elapsed": 769, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCGo6aIm0tOcd5EhqWlYb0rime9sBvHS9YMpx0D2w=s64", "userId": "08597265227091462140"}} colab={"base_uri": "https://localhost:8080/", "height": 442}
# Now we create a distribution for the average times between sales
ax = summary_df['invoicedate_purchase_frequency'].hist(bins=20, color='skyblue', rwidth=0.7, figsize=(12,7))
ax.set_xlabel('avg. number of days between purchases')
ax.set_ylabel('count')
plt.show()
# + [markdown] id="nAj-YCbAiwqo" colab_type="text"
# # Predicting the 3 month Customer Lifetime Value
# + id="Rq8fkYcqk8sU" colab_type="code" outputId="0c9ee755-e871-4f8d-ad23-3d5ea84fb47e" executionInfo={"status": "ok", "timestamp": 1574033287564, "user_tz": 0, "elapsed": 471, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCGo6aIm0tOcd5EhqWlYb0rime9sBvHS9YMpx0D2w=s64", "userId": "08597265227091462140"}} colab={"base_uri": "https://localhost:8080/", "height": 233}
orders_df.head()
# + id="uL7RK54Biwqp" colab_type="code" outputId="f6b65f4e-eb6f-4466-ab94-cd54b7334229" executionInfo={"status": "ok", "timestamp": 1574033380558, "user_tz": 0, "elapsed": 1398, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCGo6aIm0tOcd5EhqWlYb0rime9sBvHS9YMpx0D2w=s64", "userId": "08597265227091462140"}} colab={"base_uri": "https://localhost:8080/", "height": 202}
# Create summary statistics in 3 month intervals for customer Sales
# Use 3M for 3 month
clv_freq = '3M'
# Groupby customer ID now, and group all invoices for 3 month cycles.
# Aggregate on sum, mean and counts
data_df = orders_df.reset_index().groupby(['CustomerID', pd.Grouper(key='InvoiceDate', freq=clv_freq)]).agg({
'Sales': [sum, groupby_mean, groupby_count],})
data_df.columns = ['_'.join(col).lower() for col in data_df.columns]
data_df = data_df.reset_index()
data_df.head()
# + id="O-3VU10Oiwqv" colab_type="code" outputId="b3f67d40-5d1e-430c-bbf2-4a7ada37ea39" executionInfo={"status": "ok", "timestamp": 1574033674407, "user_tz": 0, "elapsed": 478, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCGo6aIm0tOcd5EhqWlYb0rime9sBvHS9YMpx0D2w=s64", "userId": "08597265227091462140"}} colab={"base_uri": "https://localhost:8080/", "height": 202}
# Adding Recency 'M'
# Get the M
date_month_map = {str(x)[:10]: 'M_%s' % (i+1) for i, x in enumerate(sorted(data_df.reset_index()['InvoiceDate'].unique(), reverse=True))}
# Create our M Column
data_df['M'] = data_df['InvoiceDate'].apply(lambda x: date_month_map[str(x)[:10]])
data_df.head()
# + id="LH3zIEqeiwqz" colab_type="code" outputId="77732623-b57e-4963-b678-329607ee09af" executionInfo={"status": "ok", "timestamp": 1574033678180, "user_tz": 0, "elapsed": 417, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCGo6aIm0tOcd5EhqWlYb0rime9sBvHS9YMpx0D2w=s64", "userId": "08597265227091462140"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
data_df['M'].unique()
# + [markdown] id="YrHDX2xmtoH0" colab_type="text"
# **RFM**
#
# By definition, RFM represents:
#
# - R(ecency): how recently did customer purchase?
# - F(rquency): how often do customer purchase?
# - M(onetary Value): how much do they spend (each time on average)?
# + id="ukJKMBXViwq6" colab_type="code" outputId="b79f3b06-933a-41c8-c612-ad15afa94c53" executionInfo={"status": "ok", "timestamp": 1574033680711, "user_tz": 0, "elapsed": 470, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCGo6aIm0tOcd5EhqWlYb0rime9sBvHS9YMpx0D2w=s64", "userId": "08597265227091462140"}} colab={"base_uri": "https://localhost:8080/", "height": 253}
# Create our Features DF
features_df = pd.pivot_table(data_df.loc[data_df['M'] != 'M_1'], values=['sales_sum', 'sales_avg', 'sales_count'], columns='M', index='CustomerID')
features_df.columns = ['_'.join(col) for col in features_df.columns]
features_df.head()
# + id="mGASBDdEiwq-" colab_type="code" outputId="abe342e3-e904-45f8-f39a-3b204171ade6" executionInfo={"status": "ok", "timestamp": 1574033708202, "user_tz": 0, "elapsed": 460, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCGo6aIm0tOcd5EhqWlYb0rime9sBvHS9YMpx0D2w=s64", "userId": "08597265227091462140"}} colab={"base_uri": "https://localhost:8080/", "height": 202}
# Create our Response DF
response_df = data_df.loc[data_df['M'] == 'M_1', ['CustomerID', 'sales_sum']]
response_df.columns = ['CustomerID', 'CLV_'+clv_freq]
response_df.head()
# + id="FbstDoBOiwrA" colab_type="code" outputId="5b2fc512-b15d-4d02-b904-2900b68cc211" executionInfo={"status": "ok", "timestamp": 1574033711445, "user_tz": 0, "elapsed": 497, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCGo6aIm0tOcd5EhqWlYb0rime9sBvHS9YMpx0D2w=s64", "userId": "08597265227091462140"}} colab={"base_uri": "https://localhost:8080/", "height": 222}
# Create our Sample Set by merging features with response_df
sample_set_df = features_df.merge(
response_df,
left_index=True,
right_on='CustomerID',
how='left'
)
sample_set_df = sample_set_df.fillna(0)
sample_set_df.head()
# + [markdown] id="d3Uv53jioMaI" colab_type="text"
# # Using Linear Regression
# + id="vsKfw9WriwrE" colab_type="code" outputId="6c107304-a220-4801-f668-41ccb0655f6c" executionInfo={"status": "ok", "timestamp": 1574033742477, "user_tz": 0, "elapsed": 502, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCGo6aIm0tOcd5EhqWlYb0rime9sBvHS9YMpx0D2w=s64", "userId": "08597265227091462140"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
# Linear regression
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
# Our Target Variable
target_var = 'CLV_'+clv_freq
# Our Features
all_features = [x for x in sample_set_df.columns if x not in ['CustomerID', target_var]]
# Split data in Train and Test
x_train, x_test, y_train, y_test = train_test_split(sample_set_df[all_features], sample_set_df[target_var], test_size=0.3)
# Fit Model
reg_fit = LinearRegression()
reg_fit.fit(x_train, y_train)
# + id="WmafmzosiwrI" colab_type="code" outputId="6449d65c-ac8c-4037-e2dc-e613164a0ceb" executionInfo={"status": "ok", "timestamp": 1574033744432, "user_tz": 0, "elapsed": 482, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCGo6aIm0tOcd5EhqWlYb0rime9sBvHS9YMpx0D2w=s64", "userId": "08597265227091462140"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
# Get our y intercept
reg_fit.intercept_
# + id="LBIAPWL6iwrK" colab_type="code" outputId="05725d3d-51bf-4e99-bbe8-30cfabcee05e" executionInfo={"status": "ok", "timestamp": 1574033745241, "user_tz": 0, "elapsed": 312, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCGo6aIm0tOcd5EhqWlYb0rime9sBvHS9YMpx0D2w=s64", "userId": "08597265227091462140"}} colab={"base_uri": "https://localhost:8080/", "height": 69}
# Get our coefficents
reg_fit.coef_
# + id="rc0Qt0M-iwrM" colab_type="code" outputId="0eb18af0-fd9a-49e7-842f-029baa7792dc" executionInfo={"status": "ok", "timestamp": 1574033747197, "user_tz": 0, "elapsed": 575, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCGo6aIm0tOcd5EhqWlYb0rime9sBvHS9YMpx0D2w=s64", "userId": "08597265227091462140"}} colab={"base_uri": "https://localhost:8080/", "height": 417}
# Show them here per feature
coef = pd.DataFrame(list(zip(all_features, reg_fit.coef_)))
coef.columns = ['feature', 'coef']
coef
# + id="sQj7tVW5iwrN" colab_type="code" colab={}
# Run our input data into our model predictors
train_preds = reg_fit.predict(x_train)
test_preds = reg_fit.predict(x_test)
# + id="oPDYpDNHiwrQ" colab_type="code" outputId="32c4d9df-9d8c-43c1-d629-e60a5a2602ec" executionInfo={"status": "ok", "timestamp": 1574033779159, "user_tz": 0, "elapsed": 1073, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCGo6aIm0tOcd5EhqWlYb0rime9sBvHS9YMpx0D2w=s64", "userId": "08597265227091462140"}} colab={"base_uri": "https://localhost:8080/", "height": 52}
# Assess performance Training Data
from sklearn.metrics import r2_score, median_absolute_error
print(r2_score(y_true=y_train, y_pred=train_preds))
print(median_absolute_error(y_true=y_train, y_pred=train_preds))
# + id="vS-yETOuiwrS" colab_type="code" outputId="09b4203f-93e7-4923-fe54-50299abcfb43" executionInfo={"status": "ok", "timestamp": 1574033786655, "user_tz": 0, "elapsed": 709, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCGo6aIm0tOcd5EhqWlYb0rime9sBvHS9YMpx0D2w=s64", "userId": "08597265227091462140"}} colab={"base_uri": "https://localhost:8080/", "height": 52}
# Compare R-Squared Performance on Training and Test Datasets
print('In-Sample R-Squared: %0.4f' % r2_score(y_true = y_train, y_pred = train_preds))
print('Out-of-Sample R-Squared: %0.4f' % r2_score(y_true = y_test, y_pred = test_preds))
# + id="wDWgZJXKiwrU" colab_type="code" outputId="a3ae55b7-e9b3-40d2-e68a-2b61662c079d" executionInfo={"status": "ok", "timestamp": 1574033803913, "user_tz": 0, "elapsed": 461, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCGo6aIm0tOcd5EhqWlYb0rime9sBvHS9YMpx0D2w=s64", "userId": "08597265227091462140"}} colab={"base_uri": "https://localhost:8080/", "height": 52}
# Compare MSE on Training and Test Datasets
print('In-Sample MSE: %0.4f' % median_absolute_error(y_true = y_train, y_pred = train_preds))
print('Out-of-Sample MSE: %0.4f' % median_absolute_error(y_true = y_test, y_pred = test_preds))
# + id="6lSxjyr5iwrW" colab_type="code" outputId="24a841d4-b1fc-4f86-94b9-ea221a36f137" executionInfo={"status": "ok", "timestamp": 1574033815556, "user_tz": 0, "elapsed": 662, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCGo6aIm0tOcd5EhqWlYb0rime9sBvHS9YMpx0D2w=s64", "userId": "08597265227091462140"}} colab={"base_uri": "https://localhost:8080/", "height": 295}
# Plot the predicted vs actual values using our Test Dataset
plt.scatter(y_test, test_preds)
plt.plot([0, max(y_test)], [0, max(test_preds)], color='gray', lw=1, linestyle='--')
plt.xlabel('actual')
plt.ylabel('predicted')
plt.title('Out-of-Sample Actual vs. Predicted')
plt.grid()
plt.show()
# + id="0h2e3UAqqEjU" colab_type="code" colab={}
| notebooks/21.0 Case Study 11 - Customer Lifetime Value.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from sklearn.preprocessing import LabelEncoder, StandardScaler, MinMaxScaler
from sklearn.pipeline import make_pipeline
from sklearn.compose import make_column_transformer
from sklearn.model_selection import train_test_split, cross_val_score
from sklearn import svm
from sklearn.tree import DecisionTreeClassifier
import matplotlib.pyplot as plt
from imblearn.over_sampling import SMOTE
from helper import get_performance
EPOCHS = 700
BATCH_SIZE = 2048
ACTIVATION = 'swish'
LEARNING_RATE = 0.0007
FOLDS = 5
# +
# Reading the dataset
data = pd.read_csv("dataset/Job_Change/aug_train.csv")
aug_train = data.sample(frac=1, replace=True, random_state=1).reset_index(drop=True)
# Seperate aug_train into target and features
y = aug_train['target']
X_aug_train = aug_train.drop('target',axis = 'columns')
# save the index for X_aug_train
X_aug_train_index = X_aug_train.index.to_list()
class MultiColumnLabelEncoder:
def __init__(self,columns = None):
self.columns = columns # array of column names to encode
def fit(self,X,y=None):
return self # not relevant here
def transform(self,X):
'''
Transforms columns of X specified in self.columns using
LabelEncoder(). If no columns specified, transforms all
columns in X.
'''
output = X.copy()
if self.columns is not None:
for col in self.columns:
# convert float NaN --> string NaN
output[col] = output[col].fillna('NaN')
output[col] = LabelEncoder().fit_transform(output[col])
else:
for colname,col in output.iteritems():
output[colname] = LabelEncoder().fit_transform(col)
return output
def fit_transform(self,X,y=None):
return self.fit(X,y).transform(X)
# store the catagorical features names as a list
cat_features = X_aug_train.select_dtypes(['object']).columns.to_list()
# use MultiColumnLabelEncoder to apply LabelEncoding on cat_features
# uses NaN as a value , no imputation will be used for missing data
X = MultiColumnLabelEncoder(columns = cat_features).fit_transform(X_aug_train)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.30, random_state = 42)
print("Train data: ", X_train.shape)
print("Test data: ", X_test.shape)
# -
y_train.value_counts()
y_test.value_counts()
# # Default Model
model_default = svm.SVC(kernel='sigmoid')
scores_default = cross_val_score(model_default, X=X_train, y=y_train, cv = FOLDS)
model_default.fit(X_train, y_train)
y_pred_default = model_default.predict(X_test)
get_performance(X_test, y_test, y_pred_default)
pd.DataFrame(y_pred_default).value_counts()
import time
import sys
sys.path.insert(1, './mmd')
from mmd import diagnoser
from scipy import stats as st
import numpy
#notebook's library
# %matplotlib inline
from helper import get_top_f1_rules, get_relevent_attributs_target, get_MMD_results, get_biased_features, get_BGMD_results
from helper import generateTrain_data_Weights
default_result = pd.concat([X_test, y_test], axis=1, join='inner')
default_result.loc[:,"pred"] = y_pred_default
def mispredict_label(row):
if row['target'] == row['pred']:
return False
return True
default_result_copy = default_result.copy()
X_test_copy = X_test.copy()
X_test_copy['mispredict'] = default_result_copy.apply(lambda row: mispredict_label(row), axis=1)
# +
settings = diagnoser.Settings
settings.all_rules = True
# Get relevent attributes and target
relevant_attributes, Target = get_relevent_attributs_target(X_test_copy)
# Generate MMD rules and correspodning information
MMD_rules, MMD_time, MMD_Features = get_MMD_results(X_test_copy, relevant_attributes, Target)
#Get biased attributes this time
biased_attributes = get_biased_features(X_test_copy, relevant_attributes)
BGMD_rules, BGMD_time, BGMD_Features = get_BGMD_results(X_test_copy, biased_attributes, Target)
print('MMD Spent:', MMD_time, 'BGMD Spent:', BGMD_time)
MMD_rules, BGMD_rules
# -
# # Decision Tree
model_default = DecisionTreeClassifier()
scores_default = cross_val_score(model_default, X=X_train, y=y_train, cv = FOLDS)
model_default.fit(X_train, y_train)
y_pred_default = model_default.predict(X_test)
get_performance(X_test, y_test, y_pred_default)
# +
default_result = pd.concat([X_test, y_test], axis=1, join='inner')
default_result.loc[:,"pred"] = y_pred_default
default_result_copy = default_result.copy()
X_test_copy = X_test.copy()
X_test_copy['mispredict'] = default_result_copy.apply(lambda row: mispredict_label(row), axis=1)
settings = diagnoser.Settings
settings.all_rules = True
# Get relevent attributes and target
relevant_attributes, Target = get_relevent_attributs_target(X_test_copy)
# Generate MMD rules and correspodning information
MMD_rules, MMD_time, MMD_Features = get_MMD_results(X_test_copy, relevant_attributes, Target)
#Get biased attributes this time
biased_attributes = get_biased_features(X_test_copy, relevant_attributes)
BGMD_rules, BGMD_time, BGMD_Features = get_BGMD_results(X_test_copy, biased_attributes, Target)
print('MMD Spent:', MMD_time, 'BGMD Spent:', BGMD_time)
MMD_rules, BGMD_rules
| BGMD/RQ1_Job_Change.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab_type="code"
# !pip install -r https://raw.githubusercontent.com/datamllab/automl-in-action-notebooks/master/requirements.txt
# + colab_type="code"
from tensorflow.keras.datasets import fashion_mnist
(x_train, _), (x_test, _) = fashion_mnist.load_data()
x_train = x_train.astype("float32") / 255.0
x_test = x_test.astype("float32") / 255.0
# + colab_type="code"
import tensorflow as tf
from tensorflow.keras.models import Model
from tensorflow.keras import layers, losses
class AutoencoderModel(Model):
def __init__(self, latent_dim):
super().__init__()
self.latent_dim = latent_dim
self.encoder_layer = layers.Dense(latent_dim, activation="relu")
self.decoder_layer = layers.Dense(784, activation="sigmoid")
def encode(self, encoder_input):
encoder_output = layers.Flatten()(encoder_input)
encoder_output = self.encoder_layer(encoder_output)
return encoder_output
def decode(self, decoder_input):
decoder_output = decoder_input
decoder_output = self.decoder_layer(decoder_output)
decoder_output = layers.Reshape((28, 28))(decoder_output)
return decoder_output
def call(self, x):
return self.decode(self.encode(x))
# + colab_type="code"
import numpy as np
tf.random.set_seed(5)
np.random.seed(5)
autoencoder = AutoencoderModel(64)
autoencoder.compile(optimizer="adam", loss="mse")
autoencoder.fit(
x_train, x_train, epochs=10, shuffle=True, validation_data=(x_test, x_test)
)
autoencoder.evaluate(x_test, x_test)
# + colab_type="code"
autoencoder.encode(x_test[:1])
# + colab_type="code"
import matplotlib.pyplot as plt
def show_images(model, images):
encoded_imgs = model.encode(images).numpy()
decoded_imgs = model.decode(encoded_imgs).numpy()
n = 10
plt.figure(figsize=(20, 4))
for i in range(n):
ax = plt.subplot(2, n, i + 1)
plt.imshow(images[i])
plt.title("original")
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
ax = plt.subplot(2, n, i + 1 + n)
plt.imshow(decoded_imgs[i])
plt.title("reconstructed")
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()
show_images(autoencoder, x_test)
# + colab_type="code"
import keras_tuner
from tensorflow import keras
from keras_tuner import RandomSearch
class AutoencoderBlock(keras.Model):
def __init__(self, latent_dim, hp):
super().__init__()
self.latent_dim = latent_dim
self.encoder_layers = []
for i in range(
hp.Int("encoder_layers", min_value=0, max_value=2, step=1, default=0)
):
self.encoder_layers.append(
layers.Dense(
units=hp.Choice("encoder_layers_{i}".format(i=i), [64, 128, 256]),
activation="relu",
)
)
self.encoder_layers.append(layers.Dense(latent_dim, activation="relu"))
self.decoder_layers = []
for i in range(
hp.Int("decoder_layers", min_value=0, max_value=2, step=1, default=0)
):
self.decoder_layers.append(
layers.Dense(
units=hp.Choice("decoder_layers_{i}".format(i=i), [64, 128, 256]),
activation="relu",
)
)
self.decoder_layers.append(layers.Dense(784, activation="sigmoid"))
def encode(self, encoder_input):
encoder_output = layers.Flatten()(encoder_input)
for layer in self.encoder_layers:
encoder_output = layer(encoder_output)
return encoder_output
def decode(self, decoder_input):
decoder_output = decoder_input
for layer in self.decoder_layers:
decoder_output = layer(decoder_output)
decoder_output = layers.Reshape((28, 28))(decoder_output)
return decoder_output
def call(self, x):
return self.decode(self.encode(x))
# + colab_type="code"
def build_model(hp):
latent_dim = 20
autoencoder = AutoencoderBlock(latent_dim, hp)
autoencoder.compile(optimizer="adam", loss="mse")
return autoencoder
tuner = RandomSearch(
build_model,
objective="val_loss",
max_trials=10,
overwrite=True,
directory="my_dir",
project_name="helloworld",
)
tuner.search(x_train, x_train, epochs=10, validation_data=(x_test, x_test))
# + colab_type="code"
autoencoder = tuner.get_best_models(num_models=1)[0]
tuner.results_summary(1)
autoencoder.evaluate(x_test, x_test)
show_images(autoencoder, x_test)
| 6.1.2-Tuning-an-autoencoder-model-for-unsupervised-learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Reconocimiento de dígitos escritos a mano
import keras
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense, Dropout
from keras.optimizers import RMSprop
# ## Paso 1. Cargar la información
# the data, shuffled and split between train and test sets
(x_train, y_train), (x_test, y_test) = mnist.load_data()
# ## Paso 2 y 3. Comprender la información y modificarla
#
# Recibimos una lista con 60mil ejemplos. En esa lista, tenemos una matriz (arreglo de arreglos) de 28x28. Para que una red neuronal normal pueda recibir como entrada esto, necesita recibir un vector, por lo que vamos a convertir la matrix en un vector de 784 (28x28). Luego, vamos a escalar los valores para que estén entre 0 y 1.
print(x_train.shape)
print(x_test.shape)
x_train = x_train.reshape(60000, 784)
x_test = x_test.reshape(10000, 784)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
print(x_train.shape)
print(x_test.shape)
# La siguiente función lo que hace es mostrar un dígito. Veamos como se ve el primer dígito de nuestro dataset como imagen, luego su valor en x y finalmente su valor en y.
# +
import matplotlib.pyplot as plt
# %matplotlib inline
def show_digit(index):
label = y_train[index].argmax(axis=0)
image = x_train[index].reshape([28,28])
plt.title('Training data. Label: {}'.format(label))
plt.imshow(image, cmap='gray_r')
plt.show()
show_digit(0)
# -
x_train[0]
y_train[0]
# ### One-hot encoding
#
# Ya has visto cómo se hace one-hot encoding. Utiliza el método ```keras.utils.to_categorical```para convertir y_train y y_test en vectores de 10 valores en vez de un valor.
#
# https://keras.io/utils/#to_categorical
# +
## TODO: Has el one hot encoding para que 5 se convierta en [0, 0, 0, 0, 1, 0, 0, 0, 0]
# -
# ## Paso 4. Construir Arquitectura del Modelo
# +
## TODO: Construye un modelo secuencial
## TODO: Compila el modelo con un optimizador y una función de pérdida
# -
# ## Paso 5. Entrenamos el modelo
#
# Intenta lograr más de 97% de precisión.
# +
## TODO: Corre el modelo. Experimenta con diferentes tamaños de batch y número de epochs.
# Usa verbose=2 para ver cómo va progresando el modelo.
# Empieza con 2 ó 3 epochs. Este es un algoritmo pesado. Al final intenta con 10 epochs.
# -
# ## Paso 6. Evaluamos el modelo
# +
## TODO: Evalua tu modelo
# -
# ## Soluciones: No las revises hasta haber terminado
# +
## TODO: Has el one hot encoding para que 5 se convierta en [0, 0, 0, 0, 1, 0, 0, 0, 0]
y_train = keras.utils.to_categorical(y_train, 10)
y_test = keras.utils.to_categorical(y_test, 10)
y_train[0]
## TODO: Construye un modelo secuencial
model = Sequential()
model.add(Dense(512, activation='relu', input_shape=(784,)))
model.add(Dropout(0.2))
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(10, activation='softmax'))
model.summary()
## TODO: Compila el modelo con un optimizador y una función de pérdida
model.compile(loss='categorical_crossentropy',
optimizer=RMSprop(),
metrics=['accuracy'])
## TODO: Corre el modelo. Experimenta con diferentes tamaños de batch y número de epochs.
# Usa verbose=2 para ver cómo va progresando el modelo.
# Empieza con 2 ó 3 epochs. Este es un algoritmo pesado. Al final intenta con 10 epochs.
history = model.fit(x_train, y_train,
batch_size=128,
epochs=3,
verbose=1,
validation_data=(x_test, y_test))
## TODO: Evalua tu modelo
score = model.evaluate(x_test, y_test, verbose=0)
print('Test accuracy:', score[1])
| Kata Intro to AI/6. DigitRecognition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Quickdraw Data
#
# If machine learning is rocket science then data is your fuel! So before
# doing anything we will have a close look at the data available and spend
# some time bringing it into the "right" form (i.e.
# [tf.train.Example](https://www.tensorflow.org/versions/r1.0/api_docs/python/tf/train/Example)).
#
# That's why we start by spending quite a lot on this notebook, downloading
# the data, understanding it, and transforming it into the right format for
# Tensorflow.
#
# The data used in this workshop is taken from Google's quickdraw (click on
# the images to see loads of examples):
#
# https://quickdraw.withgoogle.com/data
#
# Table of contents:
#
# - [ 1 Get the data](#1-Get-the-data)
# - [ 2 Inspect the data](#2-Inspect-the-data)
# - [ 3 Rasterize](#3-Rasterize)
# - [ 4 tf.train.Example data format](#4-tf.train.Example-data-format)
# - [ 5 Create dataset](#5-Create-dataset)
# - [ 6 Prepare dataset for RNN – bonus!](#6-Prepare-dataset-for-RNN-%E2%80%93-bonus!)
# +
import base64, io, itertools, json, os, random, re, time
import numpy as np
import tensorflow as tf
from matplotlib import pyplot
from PIL import Image, ImageDraw
from IPython import display
from six.moves.urllib import request
from xml.dom import minidom
# %matplotlib inline
# Always make sure you are using running the expected version.
# There are considerable differences between versions...
tf.__version__
# -
# # 1 Get the data
#
# In this section we download a set of raw data files from the web.
# +
# Retrieve list of classes.
def list_bucket(bucket, regexp='.*'):
"""Returns a (filtered) list of Keys in specified GCE bucket."""
keys = []
fh = request.urlopen('https://storage.googleapis.com/%s' % bucket)
content = minidom.parseString(fh.read())
for e in content.getElementsByTagName('Contents'):
key = e.getElementsByTagName('Key')[0].firstChild.data
if re.match(regexp, key):
keys.append(key)
return keys
all_ndjsons = list_bucket('quickdraw_dataset', '.*ndjson$')
print 'available: (%d)' % len(all_ndjsons)
print ' '.join([key.split('/')[-1].split('.')[0] for key in all_ndjsons])
# +
# Store all data locally in this directory.
data_path = '../data'
# Mini group of two animals.
pets = ['cat', 'dog']
# Somewhat larger group of zoo animals.
zoo = ['elephant', 'giraffe', 'kangaroo', 'lion', 'monkey', 'panda',
'penguin', 'rhinoceros', 'tiger', 'zebra']
# Even larger group of all animals.
animals = ['bat', 'bird', 'butterfly', 'camel', 'cat', 'cow', 'crab',
'crocodile', 'dog', 'dolphin', 'duck', 'elephant', 'fish',
'frog', 'giraffe', 'hedgehog', 'horse', 'kangaroo', 'lion',
'lobster', 'monkey', 'mosquito', 'mouse', 'octopus', 'owl',
'panda', 'parrot', 'penguin', 'pig', 'rabbit', 'raccoon',
'rhinoceros', 'scorpion', 'sea turtle', 'shark', 'sheep',
'snail', 'spider', 'squirrel', 'teddy-bear', 'tiger',
'whale', 'zebra']
# Create your own group -- the more classes you include the more challenging
# the classification task will be...
# Choose one of above groups for remainder of workshop.
# Note: This will result in ~100MB of download per class.
classes, classes_name = zoo, 'zoo'
# +
# Download above chosen group.
def valid_ndjson(filename):
"""Checks presence + completeness of .ndjson file."""
try:
json.loads(open(filename).readlines()[-1])
return True
except ValueError, IOError:
return False
def retrieve(bucket, key, filename):
"""Returns a file specified by its Key from a GCE bucket."""
url = 'https://storage.googleapis.com/%s/%s' % (bucket, key)
if not os.path.isfile(filename):
request.urlretrieve(url=url, filename=filename)
while not valid_ndjson(filename):
print '*** Corrupted download (%.2f MB), retrying...' % (os.path.getsize(filename) / 2.**20)
request.urlretrieve(url=url, filename=filename)
if not os.path.exists(data_path):
os.mkdir(data_path)
print '\n%d classes:' % len(classes)
for name in classes:
print name,
dst = '%s/%s.ndjson' % (data_path, name)
retrieve('quickdraw_dataset', 'full/simplified/%s.ndjson' % name, dst)
print '%.2f MB' % (os.path.getsize(dst) / 2.**20)
print '\nDONE :)'
# -
# # 2 Inspect the data
#
# What is the format of the downloaded files?
# So let's check out the downloaded files...
# !ls $data_path
# What is the NDJSON file format?
# Seems to be one JSON dictionary per line...
path = tf.gfile.Glob(os.path.join(data_path, '*.ndjson'))[1]
print file(path).read()[:1000] + '...'
# Parse single line.
data_json = json.loads(file(path).readline())
data_json.keys()
# So we have some meta information...
for k, v in data_json.iteritems():
if k != 'drawing':
print '%20s -> %s' % (k, v)
# ...and the actual drawing.
drawing = data_json['drawing']
# The drawing consists of a series of strokes:
[np.array(stroke).shape for stroke in drawing]
# Draw the image -- the strokes all have have shape (2, n)
# so the first index seems to be x/y coordinate:
for stroke in drawing:
pyplot.plot(np.array(stroke[0]), -np.array(stroke[1]))
# Would YOU recognize this drawing successfully?
# +
# %%time
# Some more code to load many sketches at once.
# Let's ignore the difficult "unrecognized" sketches for now...
# (i.e. unrecognized by the official quickdraw classifier)
def convert(line):
"""Converts single line to JSON + converts 'drawing' to list of np.array."""
d = json.loads(line)
d['drawing'] = [np.array(stroke) for stroke in d['drawing']]
return d
def loaditer(name, unrecognized=False):
"""Returns iterable of drawings in specified file.
Args:
name: Name of the downloaded object (e.g. "elephant").
unrecognized: Whether to include drawings that were not recognized
by Google AI (i.e. the hard ones).
"""
for line in open('%s/%s.ndjson' % (data_path, name)):
d = convert(line)
if d['recognized'] or unrecognized:
yield d
def loadn(name, n, unrecognized=False):
"""Returns list of drawings.
Args:
name: Name of the downloaded object (e.g. "elephant").
n: Number of drawings to load.
unrecognized: Whether to include drawings that were not recognized
by Google AI (i.e. the hard ones).
"""
it = loaditer(name, unrecognized=unrecognized)
return list(itertools.islice(it, 0, n))
print 'loading some "%s"...' % classes[0]
sample = loadn(classes[0], 100)
# -
# Some more drawings...
pyplot.figure(figsize=(10, 10))
n = 3
for x in range(n):
for y in range(n):
i = x * n + y
pyplot.subplot(n, n, i + 1)
for stroke in sample[i]['drawing']:
pyplot.plot(np.array(stroke[0]), -np.array(stroke[1]))
# # 3 Rasterize
#
# Idea: After converting the raw drawing data into rasterized images, we can
# use [MNIST](https://www.tensorflow.org/get_started/mnist/beginners)-like
# image processing to classify the drawings.
# +
# %%writefile _derived/1_json_to_img.py
# (Written into separate file for sharing between notebooks.)
# Function that converts drawing (specified by individual strokes)
# to a rendered black/white image.
def json_to_img(drawing, img_sz=64, lw=3, maximize=True):
img = Image.new('L', (img_sz, img_sz))
draw = ImageDraw.Draw(img)
lines = np.array([
stroke[0:2, i:i+2]
for stroke in drawing['drawing']
for i in range(stroke.shape[1] - 1)
], dtype=np.float32)
if maximize:
for i in range(2):
min_, max_ = lines[:,i,:].min() * 0.95, lines[:,i,:].max() * 1.05
lines[:,i,:] = (lines[:,i,:] - min_) / max(max_ - min_, 1)
else:
lines /= 1024
for line in lines:
draw.line(tuple(line.T.reshape((-1,)) * img_sz), fill='white', width=lw)
return img
# +
# (Load code from previous cell -- make sure to have executed above cell first.)
# %run -i _derived/1_json_to_img.py
# Show some examples.
def showimg(img):
if isinstance(img, np.ndarray):
img = Image.fromarray(img, 'L')
b = io.BytesIO()
img.convert('RGB').save(b, format='png')
enc = base64.b64encode(b.getvalue()).decode('utf-8')
display.display(display.HTML(
'<img src="data:image/png;base64,%s">' % enc))
# Fetch some images + shuffle order.
rows, cols = 10, 10
n_per_class = rows * cols // len(classes) + 1
drawings_matrix = [loadn(name, rows*cols) for name in classes]
drawings_list = reduce(lambda x, y: x + y, drawings_matrix, [])
drawings_list = np.random.permutation(drawings_list)
# Create mosaic of rendered images.
lw = 4
img_sz = 64
tableau = np.zeros((img_sz * rows, img_sz * cols), dtype=np.uint8)
for y in range(rows):
for x in range(cols):
i = y * rows + x
img = json_to_img(drawings_list[i], img_sz=img_sz, lw=lw, maximize=True)
tableau[y*img_sz:(y+1)*img_sz, x*img_sz:(x+1)*img_sz] = np.asarray(img)
showimg(tableau)
# -
# # 4 tf.train.Example data format
#
# Tensorflow's "native" format for data storage is the `tf.train.Example`
# [protocol buffer](https://en.wikipedia.org/wiki/Protocol_Buffers).
#
# In this section we briefly explore the API needed to access the data
# inside the `tf.train.Example` protocol buffer. It's **not necessary** to read
# through the
# [Python API documentation](https://developers.google.com/protocol-buffers/docs/pythontutorial).
# Create a new (empty) instance.
example = tf.train.Example()
# (empty example will print nothing)
print example
# +
# An example contains a map from feature name to "Feature".
# Every "Feature" contains a list of elements of the same
# type, which is one of:
# - bytes_list (similar to Python's "str")
# - float_list (float number)
# - int64_list (integer number)
# These values can be accessed as follows (no need to understand
# details):
# Add float value "3.1416" to feature "magic_numbers"
example.features.feature['magic_numbers'].float_list.value.append(3.1416)
# Add some more values to the float list "magic_numbers".
example.features.feature['magic_numbers'].float_list.value.extend([2.7183, 1.4142, 1.6180])
### YOUR ACTION REQUIRED:
# Create a second feature named "adversaries" and add the elements
# "Alice" and "Bob".
example.features.feature['adversaries'].bytes_list.value.extend(['Alice', 'Bob']) #example.features.feature['adversaries'].
# This will now print a serialized representation of our protocol buffer
# with features "magic_numbers" and "adversaries" set...
print example
# .. et voila : that's all you need to know about protocol buffers
# for this workshop.
# -
# # 5 Create dataset
#
# Now let's create a "dataset" of `tf.train.Example`
# [protocol buffers](https://developers.google.com/protocol-buffers/) ("protos").
#
# A single example contains all the information for a drawing (i.e. rasterized
# image, label, and meta information).
#
# A dataset consists of non-overlapping sets of examples that will be used for
# training and evaluation of the classifier (the "test" set will be used for the
# final evaluation). Because these files can quickly become very large, we
# "shard" them into multiple smaller files of equal size.
# Let's first check how many [recognized=True] examples we have in each class.
# Depending on your choice of classes you could generate up to 200k examples...
for name in classes:
print name, len(list(open('%s/%s.ndjson' % (data_path, name)))), 'recognized', len(list(loaditer(name)))
# +
# Helper code to create sharded recordio files.
# (No need to read through this.)
# Well... Since you continue to read through this cell, I could as
# well explain in more detail what it is about :-)
# Because we work with large amounts of data, we will create "sharded"
# files, that is, we split a single dataset into a number of files, like
# train-00000-of-00005, ..., train-00004-of-00005 (if we're using 5 shards).
# This way we have smaller individual files, and we can also easily access
# e.g. 20% of all data, or have 5 threads reading through the data
# simultaneously.
# The code in this cell simply takes a list of iterators and then
# randomly distributes the values returned by these iterators into sharded
# datasets (e.g. a train/eval/test split).
def rand_key(counts):
"""Returns a random key from "counts", using values as distribution."""
r = random.randint(0, sum(counts.values()))
for key, count in counts.iteritems():
if r > count or count == 0:
r -= count
else:
counts[key] -= 1
return key
def make_sharded_files(make_example, path, classes, iters, splits,
shards=10, overwrite=False, report_dt=10):
"""Create sharded files from "iters".
Args:
make_example: Converts object returned by elements of "iters"
to tf.train.Example() proto.
path: Directory that will contain recordio files.
classes: Names of classes, will be written to "labels.txt".
splits: Dictionary mapping filename to number of examples (of
every class).
shards: Number of files to be created per split.
overwrite: Whether a pre-existing directory should be overwritten.
report_dt: Number of seconds between status updates (0=no updates).
Returns:
Total number of examples written to disk (this should be equal to
the number of classes times the sum of the number of examples of
all the splits).
"""
assert len(iters) == len(classes)
if not os.path.exists(path):
os.makedirs(path)
paths = {
split: ['%s/%s-%05d-of-%05d' % (path, split, i, shards)
for i in range(shards)]
for split in splits
}
assert overwrite or not os.path.exists(paths.values()[0][0])
writers = {
split: [tf.python_io.TFRecordWriter(ps[i]) for i in range(shards)]
for split, ps in paths.iteritems()
}
t0 = time.time()
n = sum(splits.values())
examples = 0
for i in range(n):
split = rand_key(splits)
writer = writers[split][splits[split] % shards]
for j in range(len(classes)):
example = make_example(j, iters[j].next())
writer.write(example.SerializeToString())
examples += 1
remaining = sum(splits.values())
if report_dt > 0 and time.time() - t0 > report_dt:
print 'processed %d/%d (%.2f%%)' % (i, n, 100. * i / n)
t0 = time.time()
for split in splits:
for writer in writers[split]:
writer.close()
with open('%s/labels.txt' % path, 'w') as f:
f.write('\n'.join(classes))
return examples
# +
# %%writefile _derived/1_make_example_img.py
# (Written into separate file for sharing between notebooks.)
# Convert drawing tf.train.Example proto.
# Uses json_to_img() from previous cell to create raster image.
def make_example_img(label, drawing):
example = tf.train.Example()
example.features.feature['label'].int64_list.value.append(label)
img_64 = np.asarray(json_to_img(drawing, img_sz=64, lw=4, maximize=True)).reshape(-1)
example.features.feature['img_64'].int64_list.value.extend(img_64)
example.features.feature['countrycode'].bytes_list.value.append(drawing['countrycode'].encode())
example.features.feature['recognized'].int64_list.value.append(drawing['recognized'])
example.features.feature['word'].bytes_list.value.append(drawing['word'].encode())
ts = drawing['timestamp']
ts = time.mktime(time.strptime(ts[:ts.index('.')], '%Y-%m-%d %H:%M:%S'))
example.features.feature['timestamp'].int64_list.value.append(long(ts))
example.features.feature['key_id'].int64_list.value.append(long(drawing['key_id']))
return example
# +
# (Load code from previous cell -- make sure to have executed above cell first.)
# %run -i _derived/1_make_example_img.py
# Create the (rasterized) dataset.
path = '%s/dataset_img' % data_path
t0 = time.time()
n = make_sharded_files(
make_example=make_example_img,
path=path,
classes=classes,
iters=[loaditer(name) for name in classes],
# Note: We only generate few examples here so you won't be
# blocked for too long while waiting for this cell to finish.
# You can re-run the cell with larger values (don't forget to
# update "path" above) in the background to get a larger
# dataset...
splits=dict(train=5000, eval=1000, test=1000),
overwrite=True,
)
print 'stored data to "%s"' % path
print 'generated %d examples in %d seconds' % (n, time.time() - t0)
# -
# # 6 Prepare dataset for RNN – bonus!
#
# This section creates another dataset of example protos that contain the raw
# stroke data, suitable for usage with a recurrent neural network.
#
# Note that later notebooks will have a "bonus" section that uses this dataset,
# but the "non-bonus" parts can be worked through without executing below
# cells...
# +
# %%writefile _derived/1_json_to_stroke.py
# (Written into separate file for sharing between notebooks.)
# Convert stroke coordinates into normalized relative coordinates,
# one single list, and add a "third dimension" that indicates when
# a new stroke starts.
def json_to_stroke(d):
norm = lambda x: (x - x.min()) / max(1, (x.max() - x.min()))
xy = np.concatenate([np.array(s, dtype=np.float32) for s in d['drawing']], axis=1)
z = np.zeros(xy.shape[1])
if len(d['drawing']) > 1:
z[np.cumsum(np.array(map(lambda x: x.shape[1], d['drawing'][:-1])))] = 1
dxy = np.diff(norm(xy))
return np.concatenate([dxy, z.reshape((1, -1))[:, 1:]])
# +
# (Load code from previous cell -- make sure to have executed above cell first.)
# %run -i _derived/1_json_to_stroke.py
# Visualize / control output of json_to_stroke().
stroke = json_to_stroke(sample[3])
# First 2 dimensions are normalized dx/dy coordinates
# third dimension indicates "new stroke".
xy = stroke[:2, :].cumsum(axis=1)
pyplot.plot(*xy)
pxy = xy[:, stroke[2] != 0]
pyplot.plot(pxy[0], pxy[1], 'ro')
# +
# %%writefile _derived/1_make_example_stroke.py
# (Written into separate file for sharing between notebooks.)
# Convert drawing tf.train.Example proto.
# Uses json_to_stroke() from previous cell to create raster image.
def make_example_stroke(label, drawing):
example = tf.train.Example()
example.features.feature['label'].int64_list.value.append(label)
stroke = json_to_stroke(drawing)
example.features.feature['stroke_x'].float_list.value.extend(stroke[0, :])
example.features.feature['stroke_y'].float_list.value.extend(stroke[1, :])
example.features.feature['stroke_z'].float_list.value.extend(stroke[2, :])
example.features.feature['stroke_len'].int64_list.value.append(stroke.shape[1])
example.features.feature['countrycode'].bytes_list.value.append(drawing['countrycode'].encode())
example.features.feature['recognized'].int64_list.value.append(drawing['recognized'])
example.features.feature['word'].bytes_list.value.append(drawing['word'].encode())
ts = drawing['timestamp']
ts = time.mktime(time.strptime(ts[:ts.index('.')], '%Y-%m-%d %H:%M:%S'))
example.features.feature['timestamp'].int64_list.value.append(long(ts))
example.features.feature['key_id'].int64_list.value.append(long(drawing['key_id']))
return example
# +
# (Load code from previous cell -- make sure to have executed above cell first.)
# %run -i _derived/1_make_example_stroke.py
# Create the (stroke) dataset.
path = '%s/dataset_stroke' % data_path
t0 = time.time()
n = make_sharded_files(
make_example=make_example_stroke,
path=path,
classes=classes,
iters=[loaditer(name) for name in classes],
splits=dict(train=50000, eval=10000, test=10000),
overwrite=True,
)
print 'stored examples to "%s"' % path
print 'generated %d examples in %d seconds' % (n, time.time() - t0)
| extras/amld/notebooks/solutions/1_qd_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sos
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SoS
# language: sos
# name: sos
# ---
# + [markdown] kernel="SoS"
# # SoS Notebook
# + [markdown] kernel="SoS" workflow_cell=true
# SoS Notebook is a **multi-language notebook** that allows the use of multiple kernels in a single notebook with data exchange among live kernels, a **Jupyter frontend to the SoS workflow engine** to allows local and remote analysis of large datasets, with an **enhanced Jupyter frontend** that provides features such as line by line execution of cell contents. The unique combination of a multi-language kernel and a workflow engine allows easy transition from interactive data analysis to batch data processing workflows.
# + kernel="SoS"
# %revisions -n 5
# + [markdown] kernel="SoS" workflow_cell=true
# ## Installing the SoS kernel <a id="Installing_the_SoS_kernel"></a>
# + [markdown] kernel="SoS"
# Please follow the instructions in [Running SoS](https://vatlab.github.io/sos-docs/#runningsos) to install `sos`, `sos-notebook`, and relevant language modules. After the installation of `sos-notebook`, you should install the sos kernel to Jupyter with command
#
# ```bash
# % python -m sos_notebook.install
# ```
#
# `sos` should then appear in the output of:
# + kernel="SoS"
!jupyter kernelspec list
# + [markdown] kernel="SoS"
# To create a SoS notebook, start a Jupyter server using command
# ```
# $ jupyter notebook
# ```
# and select a `SoS` kernel for your new notebook.
# + [markdown] kernel="SoS"
# SoS by default records runtime signatures and skip commands that have been executed successfully. Here we change the default signature mode to disallow this feature.
# + kernel="R"
# %set -s ignore
# + [markdown] kernel="SoS"
# ## User Interface <a id="User_Interface"></a>
# + [markdown] kernel="sos"
# 
# + [markdown] kernel="SoS"
# The SoS frontend is based on the Jupyter notebook frontend but adds a dropdown list and a button to the menu bar, a side panel, and dropdown lists for all code cell.
# + [markdown] kernel="SoS"
# ### cell content <a id="cell_content"></a>
# + [markdown] kernel="SoS"
# A SoS Jupyter notebook accepts the following types of cells:
#
# |Cell type | Content |Interpreted by| Behavior |
# |----------|-----------------|----------|
# |**Markdown**| Markdown text |Jupyter| can be evaluated to have title, table etc|
# |**Subkernel**| Statements in other languages with optional SoS magics| Subkernels| SoS prepares statements and evaluate them in subkernels such as R. |
# |**SoS**| SoS statements without section header| SoS | Evaluate the cell as a SoS step in a persistent SoS dictionary.|
# |**Workflow**| SoS statements with section header | SoS | Can only be executed by magic `%run` (workflow in the current cell) or `%sosrun` (workflow in the entire notebook).|
# + [markdown] kernel="SoS"
# The **markdown cells** contains Markdown text and are rendered by Jupyter. These cells are used for displaying rich-format texts such as title and table. Most of the content of this documentation is written in such cells.
# + [markdown] kernel="SoS"
# The kernel of each **code cell** is marked by the language selector at the top-right corner of each code cell, and by the color of the prompt area of the cell. For example, the following cell is a code cell with kernel `R`.
# + kernel="R"
paste('This', 'is', 'cool', '!')
# + [markdown] kernel="SoS"
# Similar to other Jupyter kernels, SoS defines a number of *magics*, which are special commands that start with character `%`. For example, expression `{2**4}` in the following code cell is expanded by magic `%expand` before it is passed to the underlying R kernel.
# + kernel="R"
# %expand
cat("The value of expression '2**4' in Python is {2**4}")
# + [markdown] kernel="SoS"
# A note of caution is that because the underlying kernels also accepts their own magics, SoS magics in subkernel cells must start from the first lines. The following code will send `%expand` to the underlying Python3 kernel and cause an error. SoS cells are not affected by this restriction so you can put magics, new lines, even comments before actual cell content.
# + kernel="Python3"
# %expand
# + [markdown] kernel="SoS"
# A cell with a SoS kernel can be either a **SoS cell** or a **Workflow cell**, with the latter containing **section headers**. A scratch cell is executed immediately with a persistent SoS dictionary so that you can execute arbitrary SoS (Python) statements,
# + kernel="SoS" workflow_cell=false
print('This is a scratch cell')
# + [markdown] kernel="SoS"
# or a single SoS step without header
# + kernel="SoS" workflow_cell=false
# %preview -n rand.txt
output: 'rand.txt'
R: expand=True
cat(rnorm(5), file='{_output}')
# + [markdown] kernel="SoS"
# The last type of SoS cell contains formal definition of SoS steps. These cells define complete SoS workflows and can only be executed by SoS magics (with magics `%run` or `%sosrun`) in Jupyter or by SoS commands from command line. For example, executing the following cell would not execute any statement.
# + kernel="SoS" workflow_cell=true
[test_20]
print(f"This is step {step_name} of a workflow")
# + [markdown] kernel="SoS"
# and you can execute the cell with a SoS magic `%run`.
# + kernel="SoS" workflow_cell=false
# %run wf
[wf_20]
print(f"This is step {step_name} of a workflow")
# + [markdown] kernel="SoS"
# What is even more magical about these cells is that they form **notebook workflows** that consist of all sections defined in the Jupyter notebook. A `%sosrun` magic would collect all workflow stepss in a notebook and execute them.
# + kernel="SoS" workflow_cell=false
# %sosrun wf
[wf_30]
print(f"This is step {step_name} of a workflow")
# + [markdown] kernel="SoS"
# SoS provides a rich environment for you to analyze data in different languages. Generally speaking, you can
#
# * Use subkernels to analyze data interactively using different languages.
# * Use SoS cells to execute complete (and separate) scripts to analyze data or test steps of workflows, and
# * If needed, convert SoS cells to workflow cells to create complete workflows to analyze data in batch mode.
# + [markdown] kernel="SoS"
# ### Switch between kernels <a id="Switch_between_kernels"></a>
# + [markdown] kernel="SoS"
# One of the most important features of the SoS kernel is its support for multiple Jupyter subkernel. A `subkernel` can be any [Jupyter supported kernels](https://github.com/ipython/ipython/wiki/IPython-kernels-for-other-languages) that have been installed locally (or a [remote ikernel](https://bitbucket.org/tdaff/remote_ikernel) with a local definition).
#
# A subkernel has the following properties:
#
# | Property | Example | Options of magics `%use` and `%with` | Comments |
# |---| ---| | ---|
# |**name** | `R`, `R2` | (positional) |Name to identify the subkernel, usually the same as language name |
# |**kernel** | `ir`, `python`, `R_remote` | `-k`, `--kernel`| Name of Jupyter kernel, must be show in the output of command `jupyter kernelspec list`|
# |**language** | `R`, `Python2` |`-l`, `--language`| SoS definition of the language, which enables magics `%get` and `%get` for the kernel.|
# |**color** | `red`, `#FAEBD7`|`-c`, `--color`|Background color of the cell, with default defined by language definition.|
# + [markdown] kernel="SoS"
# You can switch the kernel of a code cell from a dropdown list at the top right corner of the cell or using the `%use` or `%with` magic. Despite of the flexibility on the use of local and remote kernels, multiple instances of the same kernel, use of self-defined languages, the majority of the times you will be using magics like
#
# ```
# # # %use R
# ```
#
# to switch to a language with default kernel (`ir`), color (`FDEDEC`), and name (`R`).
# + kernel="R"
# %use R
# + [markdown] kernel="SoS"
# starts and switches to a `ir` kernel so that you can enter any R commands as if you are working on a notebook with a `ir` kernel. Note that `R` stands for a SoS language extension that uses kernel `ir` and you have to use the kernel name (e.g. `iperl`) if there is no language extension for the kernel.
# + kernel="R" show_output=true
require(graphics)
pairs(mtcars, main = "mtcars data")
# + [markdown] kernel="SoS"
# As you can see, a different style is used for cells executed by the `ir` kernel. After you are done with the `ir` kernel, you can switch back to the SoS kernel using magic
# + kernel="SoS"
# %use sos
# + [markdown] kernel="SoS"
# The `%with` magic is similar to magic `%use` but it does not start a new kernel, and it accepts options `--in` (`-i`) and `--out` (`-o`) to pass specified input variables to the kernel, and return specified output variables from the kernel after the completion of the evaluation.
#
# For example, if you have
# + kernel="SoS"
n = 3
# + [markdown] kernel="SoS"
# you can pass the input and output variables to magic `%with`
# + kernel="R"
# %with R -i n -o arr
arr = rnorm(n)
# + [markdown] kernel="R"
# and obtain the result in the SoS kernel
# + kernel="SoS"
arr
# + [markdown] kernel="SoS"
# Note that any new cell will inherit the kernel of its previous code cell.
# + [markdown] kernel="SoS"
# ### Side panel <a id="Side_panel"></a>
# + [markdown] kernel="SoS"
# SoS provides a side panel that can be toggled by a cube icon next to the language selection dropdown. The side panel contains a special cell that is used for two purposes:
#
# 1. A scratch cell using which you can evaluate any expression and check its results.
# 2. A preview cell in which the `%preview` magic of any cell would be executed if the side panel is open.
#
# The input area of the panel cell has a dropdown button that allows you to execute a few frequently executed magics and previous executed statements.
#
# Because this cell is not part of the main notebook, its output will not be saved with the notebook. This allows you to test commands, check environment, values of variables and content of files without affecting the content of the notebook.
#
# + [markdown] kernel="SoS"
# ### Keyboard Shortcuts <a id="Keyboard_Shortcuts"></a>
# + [markdown] kernel="SoS"
# In addition to shortcuts defined by Jupyter (e.g. `Ctrl-Enter` to evaluate a cell and `Shift-Enter` to evaluate a cell and move next), the SoS kernel defines the following shortcuts
#
# 1. `Ctrl-Shift-Enter` This shortcut **send current line or selected text to the panel cell for evaluation**. This effectively allows you to evaluate content of a cell line by line. This shortcut works for both code and markdown cells. The panel cell will switch to the kernel of the sending cell if the sending cell is a code cell.
# 2. `Ctrl-Shift-t` executes magic `%toc` that **displays the table of content of the current notebook** in the side panel, allowing you to easily navigate within a (long) notebook.
# 3. `Ctrl-Shift-O` (output) **toggles a code cell tag `show_output`** and mark the output with a gray bar to the right of the output area, or toggle a markdown cell tag `hide_output`. The tags will be rendered accordingly in HTML reports generated using sos templates.
# 4. `Ctrl-Shift-v` (paste-table) If you have a table copied from external sources such as HTML page or excel file and if you are inside of a markdown cell, this shortcut **pastes table as markdown code to the current cell**. This allows easy copy of tables to SoS notebook.
# 5. `Ctrl-Shift-m` (markdown) **toggle a cell between markdown and code type**, which can be easier to use than select code or markdown cell type from tool bar.
# 6. `Ctrl-B` toggles side-panel. This is for compatibility with the toc2 extension of Jupyter notebook.
# + [markdown] kernel="SoS"
# ### Other usage hints <a id="Other_usage_hints"></a>
# + [markdown] kernel="SoS"
# 1. **Paste pictures from clipboard**: You can paste image from clipboard directly to a markdown cell of the Jupyter notebook using `Ctrl-V` (or `Cmd-V` under Mac OS X). The key here is that you should select a markdown cell before pasting.
# 2. **Drag and drop figures**: You can drag a picture and drop it to a markdown cell of the Jupyter notebook.
# 3. **Tab completion**: You can use tab to complete keyword, magics (enter `%` and press `TAB` to get a list of magics), variable name, function name, file name etc.
# 4. **Inspect name**: You can place your cursor inside a keyword, function name, magic etc, and press `CTRL-TAB` to inspect it. SoS will show variable name,, help message etc depending on the keyword that is being inspected.
# + [markdown] kernel="SoS"
# ### How to get help <a id="How_to_get_help"></a>
# + [markdown] kernel="SoS"
# It is recommended that you go through this document and understand how SoS Notebook works, but it is of course not possible for you to memorize all the magics, SoS actions and their options. To get help, you can
#
# 1. For a complete reference, visit the [documentation page of SoS](https://vatlab.github.io/SoS/#documentation).
# 2. For a list of available magics, type `%` and `TAB` (completion).
# 3. For help on a particular magic, execute the magic with option `-h` (e.g. `%run -h`) in the side panel. Alternatively, you can place your cursor on `%run` and press `CTRL-TAB` to get a short description of magic.
# 4. For help on a SoS action (e.g. `python`), place your cursor on the action name (e.g. before `:` on `python:`), press `CTRL-TAB` to get the help message of the action.
# + [markdown] kernel="SoS"
# ## Data exchange among kernels
# + [markdown] kernel="SoS"
# A SoS notebook can have multiple live kernels. A SoS notebook can be used as a collection of cells from multiple independent notebooks. However, the real power of SoS Notebook lies in its ability to exchange data among kernels.
# + [markdown] kernel="SoS"
# ### String interpolation (magic `%expand`) <a id="String_Interpolation"></a>
# + [markdown] kernel="SoS"
# Cell content is by default sent to SoS or subkernels untouched. A `expand` magic interpolate cell content with variables in the SoS kernel as if the content is a Python f-string. For example,
# + kernel="SoS"
filename = 'somefile'
# + kernel="Bash"
echo {filename}
# + kernel="Bash"
# %expand
echo {filename}
# + kernel="Python3"
filename = 'myfile'
print(f'A filename "{filename}" is passed ')
# + kernel="Python3"
# %expand
filename = 'myfile'
print(f'A filename "{filename}" is passed ')
# + [markdown] kernel="SoS"
# In the last example, `{filename}` is expanded by the `%expand` magic so the following statements are sent to Python 3:
# ```
# filename = 'myfile'
# print(f'A filename "somefile" is passed ')
# ```
# + [markdown] kernel="SoS"
# As you can imagine, you can keep constants such as filenames and parameters in SoS and use these information to compose scripts in subkernels for execution.
# + [markdown] kernel="SoS"
# By default, the `expand` magic expands expressions in braces `{ }`. However, as you can see from the last example, `%expand` can be awkward if the script already contains `{ }`. Although the official method is to double the braces, as in
# + kernel="Python3"
# %expand
filename = 'myfile'
print(f'A filename "{{filename}}" is passed ')
# + [markdown] kernel="SoS"
# SoS recommend the use of an alternative sigil so that you do not have to change the syntax of the original script
# + kernel="Python3"
# %expand ${ }
filename = 'myfile'
print(f'SoS filename ${filename}, Python 3 filename: "{filename}"')
# + [markdown] kernel="SoS"
# ### Markdown cell and markdown kernel
# + [markdown] kernel="SoS"
# You can include headers, lists, figures, tables in your Jupyter notebook using markdown cells. These markdown cells are rendered by Jupyter itself and do not interact with the kernels. Consequently, it is not possible to pass information (e.g. results from analysis) to markdown cells to generate dynamic output. In contrast, RStudio/RMarkdown has long allowed the inclusion of expressions in markdown texts.
#
# To overcome this problem, you can install a markdown kernel with commands
#
# ```
# pip install markdown-kernel
# python -m markdown.kernel install
# ```
# and write markdown code in code cells with a markdown kernel.
# + kernel="markdown"
Hello, this is a **code cell in markdown kernel**, not a markdown cell.
# + [markdown] kernel="SoS"
# The significance of the markdown kernel is that you can pass information from SoS to it through the `%expand` magic. For example, suppose you have defined a function to calculate Fibonacci sequence,
# + kernel="SoS"
def fibo(n):
return n if n <= 1 else (fibo(n-1) + fibo(n-2))
# + [markdown] kernel="SoS"
# You can write use it in Python expressions as follows:
# + kernel="markdown"
# %expand
The Fibonacci sequence has value {fibo(1)} when `n=1` and {fibo(10)}
when `n=10`, which can be calculated recursively by
`fibo(10)=fibo(9) + fib(8)={fibo(9)}+{fibo(8)}`, and so on.
# + [markdown] kernel="SoS"
# ### Explicit data exchange (magic `%get`) <a id="Explicit_data_exchange"></a>
# + [markdown] kernel="SoS"
# The SoS kernel provides a mechanism to pass variables between SoS and some subkernels using SoS magics.
#
# For example, magic `%get` can get specified SoS variables from the SoS kernel to the subkernel `ir`.
# + kernel="R"
# %get data
data
# + [markdown] kernel="SoS"
# SoS tries its best to find the best-matching data types between SoS and the subkernel and convert the data in the subkernel's native datatypes (e.g. Python's `DataFrame` to R's `data.frame`), so the variables you get will always be in the subkernel's native data types, not a wrapper of a foreign object (for example objects provided by `rpy2`).
# + kernel="R"
class(data)
# + [markdown] kernel="SoS"
# Similarly, using magic `%put`, you can put a variable in the subkernel to the sos kernel.
# + kernel="R"
data[ data < 1] = 0
data[ data > 1] = 1
# + kernel="R"
# %put data
# + kernel="SoS" workflow_cell=false
# %use sos
data
# + [markdown] kernel="SoS"
# Variables can also be transferred with options `--in` (`-i`) and `--out` (`-o`) of magics `%use` and `%with`. For example, if you would like to add `2` to all elements in `data` but not sure if `pandas` can do that, you can send the dataframe to `R`, add `2` and send it back.
# + kernel="R"
# %with R -i data -o data
data <- data + 2
# + kernel="SoS" workflow_cell=false
data
# + [markdown] kernel="SoS"
# ### Implicit data exchange (`sos*` variables)<a id="Implicit_data_exchange"></a>
# + [markdown] kernel="SoS"
# In addition to the use of magics `%put` and `%get` and parameters `--in` and `--out` of magics `%use` and `%with` to explicitly exchange variables between SoS and subkernels, SoS automatically shares variables with names starting with `sos` among all subkernels.
#
# For example,
# + kernel="R"
# %use R
sos_var = 100
# + kernel="Python3"
# %with Python3
sos_var += 100
print("sos_var is changed to {}".format(sos_var))
# + kernel="R"
# I am still in R
sos_var
# + kernel="SoS" workflow_cell=false
# %use sos
sos_var
# + [markdown] kernel="SoS"
# SoS supports an increasing number of languages and provides [an interface to add support for other languages](Language_Module.html). Please refer to chapter [Supported Language](Supported_Languages.html) for details on each supported language. If your language of choice is not yet supported, please considering adding SoS support for your favoriate kernel with a pull request.
# + kernel="SoS"
# clean up
# %use sos
!rm mydata.csv
# + [markdown] kernel="SoS"
# ### Capture output (Magic `%capture`)
# + [markdown] kernel="SoS"
# Magic `capture` captures output from a cell, optionally parse it in specified format (option `--as`, and save or append the result to a variable in SoS (`--to` or `--append`). The output of the cell (namely the input to magic `%capture`) can be
#
# * standard output, `%capture stdout` or just `%capture`
# * standard error, `%capture stderr`
# * plain text, `%capture text` from `text/plain` of message `display_data`
# * markdown, `%capture markdown` from `text/markdown` of message `display_data`
# * html, `%capture html` from `text/html` of message `display_data`, or
# * raw, which simply returns a list of messages.
#
# For example
# + kernel="R"
# %capture --to r_var
cat(paste(c(1,2,3)))
# + [markdown] kernel="R"
# captures standard output of the cell to variable `r_var`
# + kernel="SoS"
r_var
# + kernel="R"
# %capture text --to r_var
paste(c(1,2,3))
# + kernel="SoS"
r_var
# + [markdown] kernel="SoS"
# The `capture` magic allows passing information from the output of cells back to SoS. If the output text is structured, it can even parse the output in `json`, `csv` (comma separated values), or `tsv` (tab separated values) formats and save them in Python `dict` or `Pandas.DataFrame`.
# + [markdown] kernel="SoS"
# For example,
# + kernel="SoS"
# %capture --as csv --to table
print('a,b\n1,2\n3,4')
# + kernel="SoS"
table
# + [markdown] kernel="SoS"
# This method is especially suitable for kernels without a language module so there is no way to use a `%get` magic to retrieve information from it. For example, a [`sparql` kernel](https://www.w3.org/TR/rdf-sparql-query/) simply executes sparql queries and return results. It does not have a concept of variable so you will have to capture its output to handle it in SoS:
# + kernel="sparql"
# #%capture json --from output.txt --to sparql
# %endpoint http://dbpedia.org/sparql
# %format JSON
SELECT DISTINCT ?property
WHERE {
?s ?property ?person .
?person rdf:type foaf:Person .
}
LIMIT 3
# + kernel="SoS"
import json
json.loads(sparql)
# + [markdown] kernel="SoS"
# In addition to option `--to`, the `%capture` magic can also append cell output to an existing variable using paramter `--append`. How newly captured data is appended to the existing variable depends on type of existing and new data. Basically,
#
# 1. If the variable does not exist, `--append` is equivalent to `--to`.
# 2. If the existing variable is a list, the new data is appended to the list.
# 3. If the new data has the same type with the existing one, the new data will be appended to existing variable. That it to say, strings are appended to existing string, dictionaries are merged to existing dictionaries, and rows of data frames are appended to existing data frame.
#
# For example, we already have a variable `table`
# + kernel="SoS"
table
# + kernel="Python3"
# %capture csv --append table
print('a,b\n5,6\n7,8')
# + kernel="SoS"
table
# + [markdown] kernel="SoS"
# If you would like to collect results from multiple cells, you can create a list and use it to capture them
# + kernel="SoS"
results = []
# + kernel="SoS"
# %capture --append results
sh:
echo result from sh
# + kernel="SoS"
# %capture --append results
python:
print('result from python')
# + kernel="SoS"
results
# + [markdown] kernel="SoS"
# ## Preview of results <a id="Preview_of_results"></a>
# + [markdown] kernel="SoS"
# Instant preview of intermediate results is extrmely useful for interactive data analysis. SoS provides rich and extensible preview features to
#
# * preview files in many different formats,
# * preview variables and expressions in SoS and subkernels,
# * show preview results temporarily in the side panel or permanently in the main notebook, and
# * generate interactive tables and plots for better presentation of data both in Jupyter notebook and in converted HTML reports.
# + [markdown] kernel="SoS"
# ### Magic `%preview` <a id="Magic_preview"></a>
# + [markdown] kernel="SoS"
# SoS provides a `%preview` magic to preview files and variables (and their expressions) after the completion of a cell. By default, `%preview` displays results in the side panel if the side panel is open, and otherwise in the main notebook. You can override this behavior with options `-p` (`--panel`) or `-n` (`--notebook`) to always display results in the side panel or notebook.
#
# For example, in a subkernel R, you can do
# + kernel="R"
# %preview a.png
# %use R
png("a.png")
plot(seq(1,10, 0.1), sin(seq(1,10, 0.1)))
dev.off()
# + [markdown] kernel="SoS"
# to preview `a.png` generated by this cell. The figure will be displayed in the side panel if the side panel is open, or otherwise in the main notebook.
# + [markdown] kernel="SoS"
# The `%preview` magic also accept sos variable and expressions. For example, the following example previes variable `mtcars` in a R kernel.
# + kernel="R" workflow_cell=false
# %preview -n mtcars
# + [markdown] kernel="SoS"
# You can also specify expressions to be reviewed, but similar to command line arguments, you will need to quote the expression if it contains spaces. For example,
# + kernel="R"
# %preview -n rownames(mtcars) mtcars['gear']
# + [markdown] kernel="SoS"
# If option `-w` (`--workflow`) is specified, sos collects workflow steps from the current notebook and preview it. This would give you a better sense of what would be saved with magic `%sossave` or executed with magic `%sosrun`.
# + kernel="SoS"
# %preview --workflow
# + [markdown] kernel="SoS"
# Magic `%preview` also accepts an option `--off`, in which case it turns off the preview of output files from the SoS workflow.
# + [markdown] kernel="SoS"
# ### Preview variables from another kernel <a id="Preview_variables_from_another_kernel"></a>
# + [markdown] kernel="SoS"
# The `%preview` magic previews the variables with the kernel using which the cell is evaluated. For example, even if you have `var` defined in both SoS and `R` kernel,
# + kernel="SoS"
val = [2,3]*2
# + kernel="R"
val = rep(c(1, 2), 3)
# + kernel="R"
# %preview -n val
# + kernel="SoS"
# %preview -n val
# + [markdown] kernel="SoS"
# Note that SoS also displays type information of variables in the SoS kernel, including row and columns for data frames, but not in subkernels.
# + [markdown] kernel="SoS"
# If you would like to preview variable in another kernel, you can specify it using option `--kernel`, for example, the following cell previews a variable `r.val` in a SoS kernel.
# + kernel="SoS"
# %preview -n val --kernel R
# + [markdown] kernel="SoS"
# ### Preview of files <a id="Preview_of_files"></a>
# + [markdown] kernel="SoS"
# The `%preview` magic can be used to preview files in a variety of formats. For example, the following cell download a bam file and use `%preview` magic to preview the content of the file to make sure the file has been downloaded correctly.
# + kernel="SoS"
# %preview -n issue225.bam
download:
https://github.com/lomereiter/sambamba/raw/master/test/issue225.bam
# + [markdown] kernel="SoS"
# Similarly, you can preview files produced by another kernel
# + kernel="R"
# %preview test.png -n -s png
# %use R
png('test.png', width=600, height=400)
attach(mtcars)
plot(wt, mpg, main="Scatterplot Example",
xlab="Car Weight ", ylab="Miles Per Gallon ", pch=19)
dev.off()
# + [markdown] kernel="SoS"
# Note that SoS by default preview PDF files as an embeded object (iframe), but you can use option `-s png` to convert PDF to png to preview it as a image, if you have imagematick and Python package `wand` installed.
# + [markdown] kernel="SoS"
# ### Preview options <a id="Preview_options"></a>
# + [markdown] kernel="SoS"
# The `%magic` accepts options that are format dependent. This table lists options that are available to specific file types. Options for previewing dataframes will be described later.
#
# |file type| Option | description|
# |--|--|--|
# |pdf | `--style png` (`-s png`)| Convert PDF to png before preview. All pages are combined to produce a sngle PNG figure if the pdf file contains multiple pages. This option requires Python module `wand`.|
# | | `--pages 2 3 `| With `--style png`, preview specified pages (page numbers starting from 1) from a multi-page PDF file. |
# |txt | `--limit lines` (`-l`) | Preview number of lines to preview for text files (not limited to files with extension `.txt`), default to `5`|
# + [markdown] kernel="SoS"
# ### Automatic preview <a id="Automatic_preview"></a>
# + [markdown] kernel="SoS"
# SoS will automatically preview results of Python assignments when you send statements to the sidepanel for execution. For example, if you do execute the following cell with `Ctrl-Shift-Enter`,
# + kernel="SoS"
s='12345'
# + [markdown] kernel="SoS"
# The cell would be executed in the side panel as
# ```
# # # %preview s
# s='12345'
# ```
# which allows instant feedback when you step through your code.
# + [markdown] kernel="SoS"
# SoS would also automatic preview results from SoS workflows that are specified with statement `output:`. For example, the following SoS cell executes a scratch step (step without section head) and generates output `a.png`. The figure would be automatically displayed in the side panel after the step is executed.
# + kernel="SoS"
output: 'a.png'
R: expand=True
png("{_output}")
plot(seq(1,10, 0.1), sin(seq(1,10, 0.1)))
dev.off()
# + [markdown] kernel="SoS"
# ### Preview of remote files <a id="Preview_of_remote_files"></a>
# + [markdown] kernel="SoS"
# When a workflow or a task is executed remotely (see [Remote Execution](Remote_Execution.html) for details, result files might be on a remote host that is unavailable for local preview. In this case, you can specify the host with option `-r HOST` and preview files remotely. This essentially executes a command `sos preview -r HOST FILE` on the remote host and allows you to preview content of (large) files without copying them locally.
#
# For example, you can execute a workflow on a remote host `dev` to generate a file (`mygraphic.png`) and use `%preview -r dev` to preview it.
# + kernel="SoS"
# %preview -n mygraphic.png -r dev
# %run -r dev
R:
png(file="mygraphic.png",width=400,height=350)
plot(x=rnorm(10),y=rnorm(10),main="example")
dev.off()
# + [markdown] kernel="SoS"
# ### Preview of tables <a id="Preview_of_tables"></a>
# + [markdown] kernel="SoS"
# If a data to be previewed is a Pandas DataFrame (or a csv or excel file which would be previewed as a DataFrame), SoS will preview it as a sortable and searchable table. For example, the following cell get a data.frame `mtcar` from the R kernel (as a pandas DataFrame) and preview it in the main notebook:
# + kernel="SoS"
# %get mtcars --from R
# %preview -n mtcars
# + [markdown] kernel="R"
# Compared to previewing the same variable in R, you have the addition features of
#
# 1. sorting table by clicking the sort icon at the header of each column
# 2. displaying a subset of rows that matches specified text in the input text
#
# Note that SoS by default outputs the first 200 rows of the table. You can use option `-l` (`--limit`) to change this threshold.
#
# What makes this feature particularly interesting is that the table will be sortable and searchable when you save the jupyter notebook in HTML format (through command `sos convert analysis.ipynb analysis.html --template sos-report` or magic `%sossave --to html`).
# + [markdown] kernel="SoS"
# ### Scatter Plot <a id="Scatter_Plot"></a>
# + [markdown] kernel="SoS"
# The tablular preview of Pandas DataFrame is actually using the default `table` style of `%preview`. If you have one or more numeric columns, you can use the `scatterplot` style of `%preview` to view the data. For example, the following command plots `mpg` vs `disp` of the `mtcars` dataset, stratified by `cyl`.
# + kernel="SoS"
# %preview mtcars -n -s scatterplot mpg disp --by cyl
# + [markdown] kernel="SoS"
# The advantage of this scatterplot is that you can see a description of data when you hover over the data points, which can be more informative than static figures produced by, for example, R.
#
# The `scatterplot` style provides a number of options and you can use option `-h` with `-s` to display them:
# + kernel="SoS"
# %preview mtcars -s scatterplot -h
# + [markdown] kernel="SoS"
# For example, you can show more tooltips and multiple columns as follows:
# + kernel="SoS"
# %preview mtcars -n -s scatterplot _index disp hp mpg --tooltip wt qsec
# + [markdown] kernel="SoS"
# ### Command `sos preview` <a id="Command_sos_preview"></a>
# + [markdown] kernel="SoS"
# The `%preview` magic has a command line counterpart `sos preview`. This command cannot display any figure but can be convenient to preview content of compressed files and files that are not previewed by the operating system (e.g. `bam` files). The `-r` option is especially useful in previewing files on a remote host without logging to the remote host or copying the files to local host.
# + [markdown] kernel="SoS"
# ## Execution of Workflows <a id="Execution_of_Workflows"></a>
# + [markdown] kernel="SoS"
# We have discussed markdown cells, subkernel cells, and SoS cells, which can be considered as a subkernel cell with SoS (Python) kernel. SoS notebook supports another type of cell, namely **workflow cell**.
#
# A workflow cell is a SoS cell with one or more formal **SoS steps**, which are marked by a **section header**. In summary,
#
# * A workflow cell can contain a complete workflow and be executed by magic `%run`.
# * Sections defined in all workflow cells in a notebook form **notebook workflows**, which can be executed by magic `%sosrun` or by command `sos run` (or `sos-runner`) from command line.
# + [markdown] kernel="SoS"
# ### Magic `%run` <a id="Magic_run"></a>
# + [markdown] kernel="SoS"
# Magic `%sos` executes the content of a cell as a complete SoS workflow in a separate namespace.
#
# For example, the following workflow cell defines a SoS step, but SoS Notebook will ignore it when you execute it with `Ctrl-Enter`.
# + kernel="SoS"
[example_step]
print(f"This is {step_name}")
# + [markdown] kernel="SoS"
# You can only execute such a cell with magic `%run`:
# + kernel="SoS"
# %run
[example_step_1]
print(f"This is {step_name}")
# + [markdown] kernel="SoS"
# The `%run` magic treats the content of the cell as a SoS workflow so you can put multiple sections in the cell
# + kernel="SoS"
# %run
[example_step_10]
print(f"This is {step_name}")
[example_step_20]
print(f"This is {step_name}")
# + [markdown] kernel="SoS"
# You can have global section, define parameter, and execute the workflow multiple times with multiple `%run` magics:
# + kernel="SoS"
# %run
# %run --var 200
parameter: var = 100
[example_step_15]
print(f"This is {step_name} with option {var}")
[example_step_25]
print(f"This is {step_name}")
# + [markdown] kernel="SoS"
# It is important to remember that the workflow is executed in its own namespace so it needs to be self-contained. That is to say, if you define a variable in the SoS namespace,
# + kernel="SoS"
var = 500
# + [markdown] kernel="SoS"
# The variable is not available to the workflow
# + kernel="SoS"
# %sandbox --expect-error
# %run
print(var)
# + [markdown] kernel="SoS"
# so you will have to pass it to the workflow as command line options
# + kernel="SoS"
# %run --var {var}
parameter: var = int
print(var)
# + [markdown] kernel="SoS"
# Similar to command line tool `sos run`, magic `%run` accepts a large number of parameters. Please refer to the output of `%run -h` for details.
# + [markdown] kernel="SoS"
# ### Magic `%sosrun` <a id="Magic_sosrun"></a>
# + [markdown] kernel="SoS"
# Magic `%sosrun` is similar to `%sos` in its ability to execute workflows with many options, with the major difference in that it execute **notebook workflows**, which are defined by all sections defined in the notebook. The notebook workflow can be displayed with magic `%preview --workflow` and is the workflow that will be executed if the notebook is executed from the command line with command `sos run` or `sos-runner`.
#
# It is worth noting that magics `%sosrun` and `%run` can be used together, so that you can run single workflows with `%run` and multiple workflows with `%sosrun`. For example, you can debug and execute single workflows with magic %run`
# + kernel="SoS"
# %run stepA --var 10
[stepA]
parameter: var = 20
print(f"{step_name} with parameter {var}")
# + [markdown] kernel="SoS"
# and execute many such workflows in batch mode with magic `%sosrun`:
# + kernel="SoS"
# %sosrun master
[master]
sos_run('stepA', var=30)
# + [markdown] kernel="SoS"
# ## Conversion between `.ipynb` and `.sos` files
# <a id="Conversion between `.ipynb` and `.sos` files
# "></a>
# You can save a Jupyter notebook with SoS kernel to a SoS script using `File -> Download As -> SoS` from the browser, or using command
#
# ```
# $ sos convert myscript.ipynb myscript.sos
# ```
#
# By default, **only workflow cells will be saved to a `.sos` file to create a SoS script in correct syntax**. You can also save all cells, including cells in other kernels in a '.sos' file using option `--all`, although the resulting `.sos` file might not be executable in bath mode.
#
# You can convert an `.sos` script to `.ipynb` format using command
#
# ```
# $ sos convert myscript.sos myscript.ipynb
# ```
#
# or even to an executed notebook with option `--execute`
#
# ```
# $ sos convert myscript.sos myscript.ipynb --execute
# ```
#
# and SoS will either assign each SoS step to a cell, or split the workflow according to some cell magic if the `.sos` was exported with the `--all` option.
#
# Please refer to the tutorial on [File Conversion](../tutorials/File_Conversion.html) for details of these commands.
# + [markdown] kernel="SoS"
# ## Convert `.ipynb` to HTML format <a id="Convert_ipynb_to_HTML_format"></a>
# + [markdown] kernel="SoS"
# Because it is not particularly easy to open an `.ipynb` file (a live Jupyter server is required) and because of risk of changing the content of an `.ipynb` file, it is often desired to save an `.ipynb` file in HTML format.
#
# Jupyter makes use of a template system to control the content and style of exported HTML file. For example, you can use the default Jupyter template (`File` -> `Save As` -> `HTML`) to save all input and output cells, or you can use the [hide code Jupyter extension](https://github.com/kirbs-/hide_code) to manually hide input or output of each cell and produce a customized .HTML file.
#
# SoS provides its own `template` called `sos-report`, which can be used from command line using command
#
# ```
# sos convert analysis.ipynb analysis.html --template sos-report
# ```
# or from within Jupyter with magic `%sossave` with specified filename
#
# ```
# # # %sossave analysis.html -f
# ```
# or using the same name as the notebook with option `--to html`
# ```
# # # %sossave --to html -f
# ```
#
# Option `-f` overrides existing file if an output file already exists. You could also use other `sos` based templates such as `sos-full`, which displays all cells.
#
# The generated HTML file has the following properties
#
# 1. It by default only displays markdown cells, input and output cells with tag `report_cell`, and output cells with tag `report_output`. This is the **report view** that only displays the results of interest.
#
# 2. If you point your mose to the left top corner of the window, a display control panel will be displayed for you to select additional items to display, including all input and output cells, input and output prompts, and various messages. This is the **notebook view** that displays all the details of the analysis.
#
# The `report_output` tag can be toggled using keyboard shortcut `Ctrl-Shift-O` and cells with this tag will be marked by a gray bar to the right of the output area. The `report_cell` and `scratch` tag has to be manually added through the `tag` toolbar (`View` -> `Cell Toolbar` -> `Tags`). Whereas `report_output` only works for output of code cells, the `scratch` tag applies to both markdown and code cells. Finally, if you have created your own customized template, you can define `sos-default-template` in a configuration file (eg. `sos config --set sos-default-template mytemplate.tpl`) instead of specifying it with option `--template` each time.
#
| src/documentation/Notebook_Interface.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tensores
# Tensor é a principal estrutura de dados utilizada em um framework de deep learning, simplificadamente podemos dizer que o que o pytorch faz é realizar operações sobre os tensores.
# Tensores são um containner para dados, no nosso caso dados numéricos, você pode pensar em tensores como uma generalização de matrizes e vetores, assim, um escalar seria um tensor com 0 eixos, um vetor um tensor de 1 eixo e uma matriz um tensor de 2 eixos.
#
# Os tensores tem 3 atributos principais:
# * **Rank**: o número de eixos que o tensor tem, por exemplo, um tensor de 3 dimensões tem rank 3, pois tem três eixos.
# * **Shape**: é representado por uma tupla de inteiros que descreve quantas dimensões o tensor possui ao longo de cada eixo, por exemplo, uma matriz de 2 dimensões tem shape (3, 5), isso quer dizer que ela possui 3 linhas e 5 colunas.
# * **Type**: é o tipo do tensor, por exemplo, um tensor pode ser composto de inteiro de 32 bits com sinal, sem sinal, pontos flutuante, e assim por diante, para ver a lista dos tipos suportados pelo pytorch acesse o link: https://pytorch.org/docs/stable/tensors.html
# ### Importando PyTorch
import torch
# ## Criando tensores
# O PyTorch utiliza sua propria biblioteca de tensores, porque ele acelera as operações de tensores utilizando GPU, no entanto se você está acostumado com o numpy, converter os tensores do PyTorch para numpy ou vice e versa é muito fácil.
# #### A partir de lista do python
torch.tensor([[1., -1.], [1., -1.]])
# especifica tipo de dados do tensor
torch.tensor([[1., -1.], [1., -1.]], dtype=torch.int32)
# #### A partir de tensores do numpy
# +
import numpy as np
# tensor do pytorch a partir de um tensor do numpy
torch.tensor(np.array([[1, 2, 3], [4, 5, 6]]))
# -
# #### Tensores de números aleatórios
# Tensores com números aleatorios
torch.rand(2,4)
# Tensores aleatórios com semente, para reproduzir mesma sequência pseudoaleatória
torch.manual_seed(123456)
torch.rand(2,4)
# #### Tensores de zeros
torch.zeros([2, 4], dtype=torch.int32)
# #### Tensores de uns
torch.ones(2,3)
# #### Matriz identidade
torch.eye(4)
# #### Tensores na gpu
if torch.cuda.is_available():
cuda0 = torch.device('cuda:0')
torch.ones([2, 4], dtype=torch.float64, device=cuda0)
# ## Operações com tensores
# #### visualizando o shape do tensor
t = torch.ones(2,3)
t.size()
# #### Modificando o shape do tensor
t1 = torch.ones(4,4)
print(t1.size())
t2 = t1.view(8,2)
print(t2.size())
t3 = t1.view(1, 4,4)
print(t3.size())
# Sempre que for modificar o shape de um tensor, observe que a multiplicação do valor da dimensão de cada eixo deve ter sempre o mesmo valor, ou seja
# \begin{equation*}4*4 = 8*2 = 1*4*4\end{equation*}
# ### Conversões entre NumPy e Tensores PyTorch
# #### Numpy para PyTorch
# <font color='red'>ATENÇÃO</font>: na conversão de tensores do numpy para pytorch existe um detalhe a ser que considerado que são que as funções de rede neurais do PyTorch utilizam o tipo FloatTensor e o numpy utiliza como default o tipo float64, o que faz uma conversão automática para DoubleTensor do PyTorch e consequentemente gerando um erro. Portanto devemos utilizar a função **FloatTensor** para realizar essa conversão.
np_t = np.ones((4,5))
pt_t = torch.FloatTensor(np_t)
pt_t
# #### PyTorch para numpy
pt_t = torch.ones(2,3)
np_t = pt_t.numpy()
np_t
# #### Visualização do shape (size)
pt_t = torch.ones(5,3,2)
pt_t.size()
# #### Reshape com a função (view)
pt_t = torch.ones(5,3,2)
b = pt_t.view(2,5,3)
b.size()
# #### Adição e subtração elemento por elemento
a = torch.arange(0,24).view(4,6)
b = torch.arange(0,24).view(4,6)
c = a + b
c
d = a - b
d
# forma funcional
e = a.add(b)
e
# forma funcional
f = a.sub(b)
f
# operação inplace
a.add_(b)
a
# operação inplace
a.sub_(b)
a
# #### Muitiplicação elemento por elemento
# usando sobrecarga de operadores
c = a * b
c
# usando chamada de função
d = a.mul(b)
d
# operação inplace
a.mul_(b)
a
# #### Divisão por um escalar
c = a/2
c
# #### Média
# O metodo type é utilizado para converter o tipo do tensor
a = torch.arange(0,20).type(torch.FloatTensor).view(4,5)
a
# o metodo mean calcula a média do tensor inteiro resultando um escalar
u = a.mean()
u
# #### Média em um eixo
# calcula a média em cada linha, como existem 4 linhas, retorna um vetor de 4 elementos
u_row = a.mean(dim=1)
u_row
# calcula a média em cada coluna, como existem 5 colunas, retorna um vetor de 5 elementos
u_col = a.mean(dim=0)
u_col
# #### Desvio padrão
std = a.std()
std
std_row = a.std(dim=1)
std_row
std_col = a.std(dim=0)
std_col
| Tensores.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.2 64-bit (''matching'': conda)'
# name: python382jvsc74a57bd0454522b50f89d4a3a328678ff7012a805cf35c0635c83b4d2eabf1d7699bd5aa
# ---
# # actual_connectedness.ipynb
# adapted code to allow for computing connectedness given an actual sequence of states
# +
# Jupyter's default tab completion is broken right now;
# this seems to be a workaround until it's fixed, per
# https://github.com/jupyter/notebook/issues/3763#issuecomment-745701270
# %config Completer.use_jedi = False
# %load_ext autoreload
# %autoreload 2
# %load_ext blackcellmagic
# +
import pandas as pd
from tqdm.auto import tqdm
import pyphi
import numpy as np
import matplotlib.pyplot as plt
import connectedness as con
import compute
from utils import get_substate
from networks import get_dynamics_from_sbs as sim
import adjacency
import grid_search
from pyphi.convert import sbn2sbs, sbs2sbn, to_2d, state2le_index
from pyphi.tpm import condition_tpm, marginalize_out
# -
tqdm.pandas()
# ## Set up network
# +
def build_weights(params):
"""Return the weight matrix for layers of grids (no vertex units)."""
return adjacency.weighted_adjacency_matrix_input_and_system(
n=params["layer_size"],
n_layers=params["grid_layers"],
# Grid
w_lateral=params["grid_lateral"],
w_self=params["grid_self"],
w_in=params["grid_input"],
k=params["k"],
# Input array
w_input_lateral=params["world_lateral"],
w_input_self=params["world_self"],
k_input=params["k"],
# All
toroidal=params["toroidal"],
)
def build_network(params):
W = build_weights(params)
node_labels = [
"iA",
"iB",
"iC",
"iD",
"A1",
"B1",
"C1",
"D1",
"sA",
"sB",
"sC",
"sD",
]
temp = np.ones(params["layer_size"] * 3) * params["temp_grid"]
field = np.ones(params["layer_size"] * 3) * params["field_grid"]
network = grid_search.build_network(
params, W=W, field=field, temperature=temp, node_labels=node_labels
)
return network
# +
def get_conditioned_tpms(network, input_states, input_indices):
non_inputs = tuple(set(network.node_indices) - set(input_indices))
return [
np.squeeze(
sbn2sbs(condition_tpm(network.tpm, input_indices, state)[..., non_inputs])
)
for state in input_states
]
def get_marginal_tpms(network, input_indices, tau):
non_inputs = tuple(set(network.node_indices) - set(input_indices))
marginal = np.squeeze(
sbn2sbs(marginalize_out(input_indices, network.tpm)[..., non_inputs])
)
return [marginal] * tau
def get_input_constrained_tpms(conditioned_tpms):
input_constrained_tpms = [conditioned_tpms[-1]]
for tau, tpm in enumerate(reversed(conditioned_tpms[:-1])):
input_constrained_tpms.append(np.matmul(tpm, input_constrained_tpms[tau]))
return list(input_constrained_tpms)
def get_unconstrained_tpms(marginal_tpms):
unconstrained_tpms = [marginal_tpms[0]]
for tau, tpm in enumerate(marginal_tpms[1:]):
unconstrained_tpms.append(np.matmul(tpm, unconstrained_tpms[tau]))
return unconstrained_tpms
def get_subset_connectedness(
subset_indices, subset_state, input_constrained_tpms, unconstrained_tpms, n_inputs
):
translated_subset_indices = tuple([i - n_inputs for i in subset_indices])
connectedness = []
for conditioned_tpm, marginal_tpm in zip(
input_constrained_tpms, unconstrained_tpms
):
Pr_subsetstate_given_input = sbn2sbs(
to_2d(sbs2sbn(conditioned_tpm))[..., translated_subset_indices]
).mean(0)[state2le_index(subset_state)]
Pr_subsetstate = sbn2sbs(
to_2d(sbs2sbn(marginal_tpm))[..., translated_subset_indices]
).mean(0)[state2le_index(subset_state)]
# get intrinsic difference
connectedness.append(np.log2(Pr_subsetstate_given_input / Pr_subsetstate))
return connectedness
def get_change_pr_step(data):
return np.sum(np.abs(data[:-1]-data[1:]),1)
def get_time_to_change(data):
changes = get_change_pr_step(data)
time_to_change = []
t = 1
for c in changes:
if c == 0:
t+=1
else:
time_to_change.append(t)
t=1
return time_to_change
def get_data_sample(data,t,tau,subset_indices,input_indices):
subset_state = data[t,subset_indices]
input_sequence = data[t-tau-1:t,input_indices]
return subset_state,input_sequence
# -
# # Look at a simple example for the use of this notebook
# +
# Define a network
n = 4
layers = 2
params = {
"layer_size": n,
"grid_layers": layers,
"grid_input": 0.025,
"grid_lateral": 0.2,
"grid_self": 0.8,
"world_lateral": 0.01,
"world_self": 0.8,
"temp_grid": 0.4,
"field_grid": 0,
"k": 1,
"toroidal": True,
}
network = build_network(params)
# -
# set parameters and generate an exampl input sequence
subset_indices = (9, 10)
subset_state = (1,1)
input_indices = (0, 1, 2, 3)
input_sequence = [
(0, 0, 0, 0),
(0, 0, 0, 0),
(0, 0, 0, 0),
(0, 0, 0, 0),
(0, 0, 0, 0),
(0, 0, 0, 0),
(0, 0, 0, 0),
(0, 0, 0, 0),
(0, 0, 0, 0),
(0, 0, 0, 0),
(0, 0, 0, 0),
(0, 0, 0, 0),
(0, 0, 0, 0),
(0, 0, 0, 0),
(0, 0, 0, 0),
(0, 0, 0, 0),
]
tau = len(input_sequence)
n_inputs = len(input_indices)
# +
# compute connectedness for the example data sequence
conditioned_tpms = get_conditioned_tpms(network, input_sequence, input_indices)
marginal_tpms = get_marginal_tpms(network, input_indices, tau)
input_constrained_tpms = get_input_constrained_tpms(conditioned_tpms)
unconstrained_tpms = get_unconstrained_tpms(marginal_tpms)
connectedness = get_subset_connectedness(subset_indices, subset_state, input_constrained_tpms, unconstrained_tpms, n_inputs)
connectedness
# -
# ## Simulate some real data
# +
# Prepare RNG
ENTROPY = 19283591005102385916723052837195786192730157108218751289951982
SS = np.random.SeedSequence(ENTROPY)
RNG = np.random.default_rng(SS)
data = sim(pyphi.convert.sbn2sbs(network.tpm), 0, 1000,RNG)
# + [markdown] tags=[]
# ## Separate out data in the input array, and inspect its variability
# +
input_data_sequences = data[:,:4]
plt.subplot(211)
plt.hist(get_time_to_change(input_data_sequences),range(1,20))
plt.title('Time with constant input')
plt.subplot(212)
plt.hist(get_change_pr_step(input_data_sequences),range(5))
plt.title('flips per step')
plt.tight_layout()
# -
# ## Get a sample of data, and compute connectedness
# +
# define parameters to select the input data
t = 200
tau = 20
subset_indices = (9,10)
input_indices = (0,1,2,3)
subset_state, input_sequence = get_data_sample(data,t,tau,subset_indices,input_indices)
# +
# compute connectedness for this data sample
conditioned_tpms = get_conditioned_tpms(network, input_sequence, input_indices)
marginal_tpms = get_marginal_tpms(network, input_indices, tau)
input_constrained_tpms = get_input_constrained_tpms(conditioned_tpms)
unconstrained_tpms = get_unconstrained_tpms(marginal_tpms)
connectedness = get_subset_connectedness(subset_indices, subset_state, input_constrained_tpms, unconstrained_tpms, n_inputs)
# -
plt.plot(range(tau),connectedness)
plt.ylabel('connectedness')
plt.xticks(range(tau+1),input_sequence,rotation = 90)
plt.xlabel('input states in the past\n (present ---> past)')
plt.title(subset_state);
# # Get connectedness for multiple samples to get an idea of domain connectedness
#
# +
tau = 20
times = range(tau + 1, 1000, tau)
subset_indices = (9, 10)
input_indices = (0, 1, 2, 3)
structure_connectedness = []
for t in tqdm(list(times)):
subset_state, input_sequence = get_data_sample(
data, t, tau, subset_indices, input_indices
)
# compute connectedness for this data sample
conditioned_tpms = get_conditioned_tpms(network, input_sequence, input_indices)
marginal_tpms = get_marginal_tpms(network, input_indices, tau)
input_constrained_tpms = get_input_constrained_tpms(conditioned_tpms)
unconstrained_tpms = get_unconstrained_tpms(marginal_tpms)
structure_connectedness.append(
get_subset_connectedness(
subset_indices,
subset_state,
input_constrained_tpms,
unconstrained_tpms,
n_inputs,
)
)
# -
# plot the results
for c in structure_connectedness:
plt.plot(range(tau),c,'b',alpha=0.1)
plt.plot(range(tau),np.mean(structure_connectedness,0),'b',linewidth=3)
plt.ylabel('connectedness')
plt.xlabel('tau (timesteps into the past)')
# # Repeating the experiment, but with no structure in the world
# +
# define a new network with no connectivity between world elments
n = 4
layers = 2
params = {
"layer_size": n,
"grid_layers": layers,
"grid_input": 0.022,
"grid_lateral": 0.01,
"grid_self": 0.8,
"world_lateral": 0,
"world_self": 0,
"temp_grid": 0.4,
"field_grid": 0,
"k": 1,
"toroidal": True,
}
network_no_structure = build_network(params)
# simulate data
data_no_structure = sim(pyphi.convert.sbn2sbs(network_no_structure.tpm), 0, 1000,RNG)
# +
# compute connectedness
tau = 20
times = range(tau + 1, 1000, tau)
subset_indices = (9, 10)
input_indices = (0, 1, 2, 3)
no_structure_connectedness = []
for t in tqdm(list(times)):
subset_state, input_sequence = get_data_sample(
data_no_structure, t, tau, subset_indices, input_indices
)
# compute connectedness for this data sample
conditioned_tpms_no_structure = get_conditioned_tpms(network_no_structure, input_sequence, input_indices)
marginal_tpms_no_structure = get_marginal_tpms(network_no_structure, input_indices, tau)
input_constrained_tpms_no_structure = get_input_constrained_tpms(conditioned_tpms_no_structure)
unconstrained_tpms_no_structure = get_unconstrained_tpms(marginal_tpms_no_structure)
no_structure_connectedness.append(
get_subset_connectedness(
subset_indices,
subset_state,
input_constrained_tpms_no_structure,
unconstrained_tpms_no_structure,
len(input_indices),
)
)
# +
# plot the results (comparing connectedness with and without structure in the world)
for c in structure_connectedness:
plt.plot(range(tau),c,'b',alpha=0.1)
plt.plot(range(tau),np.mean(structure_connectedness,0),'b',linewidth=3)
import matplotlib.pyplot as plt
for c in no_structure_connectedness:
plt.plot(range(tau),c,'r',alpha=0.1)
plt.plot(range(tau),np.mean(no_structure_connectedness,0),'r',linewidth=3)
plt.ylabel('connectedness');
plt.xlabel('tau [number of past states included]')
plt.xticks(range(tau),range(tau))
plt.title('BC connectedness to input states in the past');
# -
# Notice that for these parameters, the mean connectedness (thick lines) is basically the same for a structured (blue) and an unstructured (red) world. The main difference between the two is that the variance in connectedness values is different (smaller for no structure)
# # Trying the comparison for different values of input strength
# ## input strength: 0.2
# +
n = 4
layers = 2
tau = 20
times = range(tau + 1, 1000, tau)
subset_indices = (9, 10)
input_indices = (0, 1, 2, 3)
# Prepare RNG
ENTROPY = 19283591005102385916723052837195786192730157108218751289951982
SS = np.random.SeedSequence(ENTROPY)
RNG = np.random.default_rng(SS)
params = {
"layer_size": n,
"grid_layers": layers,
"grid_input": 0.2,
"grid_lateral": 0.01,
"grid_self": 0.8,
"world_lateral": 0.01,
"world_self": 0.8,
"temp_grid": 0.4,
"field_grid": 0,
"k": 1,
"toroidal": True,
}
network = build_network(params)
params["world_self"] = 0
params["world_lateral"] = 0
network_no_structure = build_network(params)
# simulate data
data = sim(pyphi.convert.sbn2sbs(network.tpm), 0, 1000,RNG)
data_no_structure = sim(pyphi.convert.sbn2sbs(network_no_structure.tpm), 0, 1000,RNG)
# computing connectedness
# for structured world
structure_connectedness = []
for t in tqdm(list(times)):
subset_state, input_sequence = get_data_sample(
data, t, tau, subset_indices, input_indices
)
# compute connectedness for this data sample
conditioned_tpms = get_conditioned_tpms(network, input_sequence, input_indices)
marginal_tpms = get_marginal_tpms(network, input_indices, tau)
input_constrained_tpms = get_input_constrained_tpms(conditioned_tpms)
unconstrained_tpms = get_unconstrained_tpms(marginal_tpms)
structure_connectedness.append(
get_subset_connectedness(
subset_indices,
subset_state,
input_constrained_tpms,
unconstrained_tpms,
n_inputs,
)
)
# for unstructured world
no_structure_connectedness = []
for t in tqdm(list(times)):
subset_state, input_sequence = get_data_sample(
data_no_structure, t, tau, subset_indices, input_indices
)
# compute connectedness for this data sample
conditioned_tpms_no_structure = get_conditioned_tpms(network_no_structure, input_sequence, input_indices)
marginal_tpms_no_structure = get_marginal_tpms(network_no_structure, input_indices, tau)
input_constrained_tpms_no_structure = get_input_constrained_tpms(conditioned_tpms_no_structure)
unconstrained_tpms_no_structure = get_unconstrained_tpms(marginal_tpms_no_structure)
no_structure_connectedness.append(
get_subset_connectedness(
subset_indices,
subset_state,
input_constrained_tpms_no_structure,
unconstrained_tpms_no_structure,
len(input_indices),
)
)
# -
for c in structure_connectedness:
plt.plot(range(tau),c,'b',alpha=0.1)
plt.plot(range(tau),np.mean(structure_connectedness,0),'b',linewidth=3)
import matplotlib.pyplot as plt
for c in no_structure_connectedness:
plt.plot(range(tau),c,'r',alpha=0.1)
plt.plot(range(tau),np.mean(no_structure_connectedness,0),'r',linewidth=3)
plt.ylabel('connectedness');
plt.xlabel('tau [number of past states included]')
plt.xticks(range(tau),range(tau))
plt.title('BC connectedness to input states in the past');
# # input strength: 0.5
# +
n = 4
layers = 2
tau = 20
times = range(tau + 1, 1000, tau)
subset_indices = (9, 10)
input_indices = (0, 1, 2, 3)
# Prepare RNG
ENTROPY = 19283591005102385916723052837195786192730157108218751289951982
SS = np.random.SeedSequence(ENTROPY)
RNG = np.random.default_rng(SS)
params = {
"layer_size": n,
"grid_layers": layers,
"grid_input": 0.5,
"grid_lateral": 0.01,
"grid_self": 0.8,
"world_lateral": 0.01,
"world_self": 0.8,
"temp_grid": 0.4,
"field_grid": 0,
"k": 1,
"toroidal": True,
}
network = build_network(params)
params["world_self"] = 0
params["world_lateral"] = 0
network_no_structure = build_network(params)
# simulate data
data = sim(pyphi.convert.sbn2sbs(network.tpm), 0, 1000,RNG)
data_no_structure = sim(pyphi.convert.sbn2sbs(network_no_structure.tpm), 0, 1000,RNG)
# computing connectedness
# for structured world
structure_connectedness = []
for t in tqdm(list(times)):
subset_state, input_sequence = get_data_sample(
data, t, tau, subset_indices, input_indices
)
# compute connectedness for this data sample
conditioned_tpms = get_conditioned_tpms(network, input_sequence, input_indices)
marginal_tpms = get_marginal_tpms(network, input_indices, tau)
input_constrained_tpms = get_input_constrained_tpms(conditioned_tpms)
unconstrained_tpms = get_unconstrained_tpms(marginal_tpms)
structure_connectedness.append(
get_subset_connectedness(
subset_indices,
subset_state,
input_constrained_tpms,
unconstrained_tpms,
n_inputs,
)
)
# for unstructured world
no_structure_connectedness = []
for t in tqdm(list(times)):
subset_state, input_sequence = get_data_sample(
data_no_structure, t, tau, subset_indices, input_indices
)
# compute connectedness for this data sample
conditioned_tpms_no_structure = get_conditioned_tpms(network_no_structure, input_sequence, input_indices)
marginal_tpms_no_structure = get_marginal_tpms(network_no_structure, input_indices, tau)
input_constrained_tpms_no_structure = get_input_constrained_tpms(conditioned_tpms_no_structure)
unconstrained_tpms_no_structure = get_unconstrained_tpms(marginal_tpms_no_structure)
no_structure_connectedness.append(
get_subset_connectedness(
subset_indices,
subset_state,
input_constrained_tpms_no_structure,
unconstrained_tpms_no_structure,
len(input_indices),
)
)
# -
for c in structure_connectedness:
plt.plot(range(tau),c,'b',alpha=0.1)
plt.plot(range(tau),np.mean(structure_connectedness,0),'b',linewidth=3)
import matplotlib.pyplot as plt
for c in no_structure_connectedness:
plt.plot(range(tau),c,'r',alpha=0.1)
plt.plot(range(tau),np.mean(no_structure_connectedness,0),'r',linewidth=3)
plt.ylabel('connectedness');
plt.xlabel('tau [number of past states included]')
plt.xticks(range(tau),range(tau))
plt.title('BC connectedness to input states in the past');
# # Experimenting with dataframe representation
def get_timevarying_connectedness(
network, data, times, tau, input_indices, mechanisms, purviews=None
):
content = []
for t in tqdm(times):
input_sequence = data[t - tau - 1 : t - 1, input_indices]
state = data[t, :]
system_indices = tuple(set(network.node_indices) - set(input_indices))
system = pyphi.subsystem.Subsystem(network, tuple(state), system_indices)
ces = pyphi.compute.ces(system, mechanisms, purviews)
# compute connectedness for this data sample
conditioned_tpms = get_conditioned_tpms(network, input_sequence, input_indices)
marginal_tpms = get_marginal_tpms(network, input_indices, tau)
input_constrained_tpms = get_input_constrained_tpms(conditioned_tpms)
unconstrained_tpms = get_unconstrained_tpms(marginal_tpms)
content.extend(
[
pd.DataFrame.from_dict(
{
(
t,
-(tt + 1),
concept.mechanism,
tuple(state[m] for m in concept.mechanism),
tuple(input_sequence[-(tt + 1), :]),
): pd.Series(
{
"connectedness": connectedness,
"cause phi": concept.cause.phi,
"cause purview": concept.cause.purview,
"cause maximal state": concept.cause.maximal_state,
"cause actual state": tuple(
data[t - 1, concept.cause.purview]
),
"congruent": any(
[
tuple(s)
== tuple(data[t - 1, concept.cause.purview])
for s in concept.cause.maximal_state
]
),
}
)
for tt, connectedness in enumerate(
get_subset_connectedness(
concept.cause.purview,
tuple(data[t - 1, concept.cause.purview]),
input_constrained_tpms,
unconstrained_tpms,
len(input_indices),
)
)
},
orient="index",
)
for concept in ces
]
)
df = pd.concat(content)
df.index.names = ["t", "tau", "mechanism", "mech state", "input state"]
return df
# + tags=[]
# creating a new network
n = 4
layers = 2
params = {
"layer_size": n,
"grid_layers": layers,
"grid_input": 0.8,
"grid_lateral": 0.2,
"grid_self": 0.8,
"world_lateral": 0.2,
"world_self": 0.8,
"temp_grid": 0.4,
"field_grid": 0,
"k": 1,
"toroidal": True,
}
network = build_network(params)
params["world_lateral"] = 0
params["world_self"] = 0
network_ns = build_network(params)
# +
# simulate data
# Prepare RNG
ENTROPY = 19283591005102385916723052837195786192730157108218751289951982
SS = np.random.SeedSequence(ENTROPY)
RNG = np.random.default_rng(SS)
time = 1000
data = sim(pyphi.convert.sbn2sbs(network.tpm), 0, time,RNG)
data_ns = sim(pyphi.convert.sbn2sbs(network_ns.tpm), 0, time,RNG)
# +
max_tau = 25
times = list(range(max_tau+1,time,5))
inputs = (0, 1, 2, 3)
mechanisms = ((5,), (6,), (5, 6), (9,), (10,), (11,), (9, 10), (10, 11), (9, 10, 11))
purviews = tuple(pyphi.utils.powerset((5, 6, 7, 9, 10, 11), nonempty=True, max_size=3))
df = get_timevarying_connectedness(
network, data, times, max_tau, inputs, mechanisms, purviews,
)
df_ns = get_timevarying_connectedness(
network_ns, data_ns, times, max_tau, inputs, mechanisms, purviews,
)
# -
df.to_csv('data/connectedness/actual-connectedness_structure_strong-input.csv')
df_ns.to_csv('data/connectedness/actual-connectedness_no-structure_strong-input.csv')
# + jupyter={"outputs_hidden": true} tags=[]
df.head(45)
# -
df_reset = df.reset_index()
df_reset.loc[(df_reset.mechanism ==(9,10)) & (df_reset['mech state']==(1,0)) & ~(df_reset['congruent'])]
# + tags=[]
import seaborn as sb
# -
sb.relplot(
data=df_reset, kind="line",
x="tau", y="connectedness",
hue="congruent",col="mechanism",
col_wrap = 3, facet_kws=dict(sharey=True)
)
df_reset_ns = df_ns.reset_index()
sb.relplot(
data=df_reset_ns, kind="line",
x="tau", y="connectedness",
hue="congruent",col="mechanism",
col_wrap = 3, facet_kws=dict(sharey=True)
)
df1 = pd.read_csv('data/connectedness/actual-connectedness_structure_strong-input.csv')
df2 = pd.read_csv('data/connectedness/actual-connectedness_no-structure_strong-input.csv')
df1.loc[(df1.mechanism =='(9, 10)')]
# +
input_data_sequences = data[:,:4]
input_data_sequences_ns = data_ns[:,:4]
plt.subplot(221)
plt.hist(get_time_to_change(input_data_sequences),range(1,20))
plt.title('Time with constant input')
plt.subplot(222)
plt.hist(get_change_pr_step(input_data_sequences),range(5))
plt.title('flips per step')
plt.subplot(223)
plt.hist(get_time_to_change(input_data_sequences_ns),range(1,20))
plt.title('Time with constant input')
plt.subplot(224)
plt.hist(get_change_pr_step(input_data_sequences_ns),range(5))
plt.title('flips per step')
plt.tight_layout()
# -
data[100:120,:]
# +
# creating a new network
n = 4
layers = 2
params = {
"layer_size": n,
"grid_layers": layers,
"grid_input": 0.7,
"grid_lateral": 0.2,
"grid_self": 0.8,
"world_lateral": 0.2,
"world_self": 0.8,
"temp_grid": 0.4,
"field_grid": 0,
"k": 1,
"toroidal": True,
}
network = build_network(params)
params["world_lateral"] = 0
params["world_self"] = 0
network_ns = build_network(params)
# -
tpm = network.tpm
import extrinsic_information as ei
import visualization as viz
viz.plot_distribution(ei.get_stationary_distribution(np.squeeze(condition_tpm(tpm,(0,1,2,3),(0,1,0,1))[...,(4,5,6,7,8,9,10,11)])))
# +
t = 25
s = 50
clamped_tpm_hom = np.zeros((t+1,8,s))
clamped_tpm_het = np.zeros((t+1,8,s))
for i in range(s):
clamped_tpm_hom[:,:,i] = sim(sbn2sbs(np.squeeze(condition_tpm(tpm,(0,1,2,3),(0,0,0,0))[...,(4,5,6,7,8,9,10,11)])),25,t)
clamped_tpm_het[:,:,i] = sim(sbn2sbs(np.squeeze(condition_tpm(tpm,(0,1,2,3),(1,0,1,0))[...,(4,5,6,7,8,9,10,11)])),25,t)
# -
plt.imshow(np.mean(clamped_tpm_hom,2),vmin=0,vmax=1)
plt.imshow(np.mean(clamped_tpm_het,2),vmin=0,vmax=1)
clamped_tpm_het[0]
| actual-connectedness.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
# # %matplotlib inline
# -
#export
from exp.nb_06 import *
# # ConvNet
x_train, y_train, x_valid, y_valid = get_data()
x_train.mean(), x_train.std()
train_ds, valid_ds = Dataset(x_train, y_train), Dataset(x_valid, y_valid)
nh, bs = 50, 512
c = y_train.max().item() + 1
loss_func = F.cross_entropy
data = DataBunch(*get_dls(train_ds, valid_ds, bs), c)
mnist_view = view_tfm(1,28,28)
cbfs = [Recorder,
partial(AvgStatsCallback, accuracy),
CudaCallback,
partial(BatchTransformXCallback, mnist_view)]
nfs = [8, 16, 32, 64, 64]
sched = combine_scheds([0.5, 0.5], [sched_cos(0.2, 1), sched_cos(1., 0.1)])
# +
# learn, run = get_learn_run(nfs, data, 1., conv_layer, cbs=cbfs+[partial(ParamScheduler, 'lr', sched)])
# -
cbfs = [partial(BatchTransformXCallback, mnist_view),
Recorder,
partial(AvgStatsCallback, accuracy),
CudaCallback]
learn, run = get_learn_run(nfs, data, 0.4, conv_layer, cbs=cbfs + [partial(ParamScheduler, 'lr', sched)])
run.fit(2, learn)
# # Batchnorm
# ## Custom
class BatchNorm(nn.Module):
def __init__(self, nf, mom=0.1, eps=1e-5):
super().__init__()
self.mom, self.eps = mom, eps
self.mults = nn.Parameter(torch.ones(nf,1,1))
self.adds = nn.Parameter(torch.zeros(nf,1,1))
self.register_buffer('means', torch.zeros(1, nf, 1, 1))
self.register_buffer('vars', torch.ones(1, nf, 1, 1))
def forward(self, x):
if self.training:
with torch.no_grad(): m,v = self.update_stats(x)
else: m,v = self.means, self.vars
x = (x - m) / (v + self.eps).sqrt()
return x * self.mults + self.adds
def update_stats(self, x):
m = x.mean((0,2,3), keepdim=True)
v = x.var((0,2,3), keepdim=True)
self.means.lerp_(m, self.mom)
self.vars.lerp_(v, self.mom)
return m,v
def conv_layer(ni, nf, ks=3, stride=2, bn=True, **kwargs):
# no bias needed if using bn
layers = [nn.Conv2d(ni, nf, ks, stride, padding=ks//2, bias=not bn),
GeneralRelu(**kwargs)]
if bn: layers.append(BatchNorm(nf))
return nn.Sequential(*layers)
#export
def get_learn_run(nfs, data, lr, layer, cbs=None, opt_func=None, uniform=False, **kwargs):
model = get_cnn_model(data, nfs, layer, **kwargs)
init_cnn(model, uniform=uniform)
return get_runner(model, data, lr=lr, cbs=cbs, opt_func=opt_func)
#export
def init_cnn(m, uniform=False):
f = init.kaiming_uniform_ if uniform else init.kaiming_uniform_
init_cnn_(m, f)
#export
def init_cnn_(m, f):
if isinstance(m, nn.Conv2d):
f(m.weight, a=0.1)
if getattr(m, 'bias', None) is not None: m.bias.data.zero_()
for l in m.children(): init_cnn_(l, f)
learn, run = get_learn_run(nfs, data, 0.9, conv_layer, cbs=cbfs)
with Hooks(learn.model, append_stats) as hooks:
run.fit(1, learn)
fig, (ax0,ax1) = plt.subplots(1,2, figsize=(10,4))
for h in hooks[:-1]:
ms, ss = h.stats
ax0.plot(ms[:10])
ax1.plot(ss[:10])
h.remove()
plt.legend(range(6));
fig, (ax0,ax1) = plt.subplots(1,2, figsize=(10,4))
for h in hooks[:-1]:
ms, ss = h.stats
ax0.plot(ms)
ax1.plot(ss)
plt.legend(range(6));
learn, run = get_learn_run(nfs, data, 1.0, conv_layer, cbs=cbfs)
# %time run.fit(3, learn)
#export
def conv_layer(ni, nf, ks=3, stride=2, bn=True, **kwargs):
layers = [nn.Conv2d(ni, nf, ks, stride, padding=ks//2, bias=not bn),
GeneralRelu(**kwargs)]
if bn: layers.append(nn.BatchNorm2d(nf, eps=1e-5, momentum=0.1))
return nn.Sequential(*layers)
learn, run = get_learn_run(nfs, data, 1.0, conv_layer, cbs=cbfs)
# %time run.fit(3, learn)
sched = combine_scheds([0.3, 0.7], [sched_lin(0.6, 2), sched_lin(2., 0.1)])
learn, run = get_learn_run(nfs, data, 0.9, conv_layer, cbs=cbfs + [partial(ParamScheduler, 'lr', sched)])
run.fit(8, learn)
# ## More norms
# ### Layer norm
class LayerNorm(nn.Module):
__constants__ = ['eps']
def __init__(self, eps=1e-5):
super().__init__()
self.eps = eps
self.mult = nn.Parameter(tensor(1.))
self.add = nn.Parameter(tensor(0.))
def forward(self, x):
m = x.mean((1,2,3), keepdim=True)
v = x.var((1,2,3), keepdim=True)
x = (x - m) / (v + self.eps).sqrt()
return x * self.mult + self.add
def conv_ln(ni, nf, ks=3, stride=2, bn=True, **kwargs):
layers = [nn.Conv2d(ni, nf, ks, stride, padding=ks//2, bias=True),
GeneralRelu(**kwargs)]
if bn: layers.append(LayerNorm())
return nn.Sequential(*layers)
learn,run = get_learn_run(nfs, data, 0.8, conv_ln, cbs=cbfs)
# %time run.fit(3, learn)
# ### Instance Norm
class InstanceNorm(nn.Module):
__constans__ = ['eps']
def __init__(self, nf, eps=1e-0):
super().__init__()
self.eps = eps
self.mults = nn.Parameter(torch.ones(nf,1,1))
self.adds = nn.Parameter(torch.zeros(nf,1,1))
def forward(self, x):
m = x.mean((2,3), keepdim=True)
v = x.var((2,3), keepdim=True)
res = (x - m) / (v + self.eps).sqrt()
return res * self.mults + self.adds
def conv_in(ni, nf, ks=3, stride=2, bn=True, **kwargs):
layers = [nn.Conv2d(ni, nf, ks, stride, padding=ks//2, bias=True),
GeneralRelu(**kwargs)]
if bn: layers.append(InstanceNorm(nf))
return nn.Sequential(*layers)
learn,run = get_learn_run(nfs, data, 0.1, conv_in, cbs=cbfs)
# %time run.fit(3, learn)
# # Fix small batch sizes
data = DataBunch(*get_dls(train_ds, valid_ds, 2), c)
def conv_layer(ni, nf, ks=3, stride=2, bn=True, **kwargs):
layers = [nn.Conv2d(ni, nf, ks, padding=ks//2, stride=stride, bias=not bn),
GeneralRelu(**kwargs)]
if bn: layers.append(nn.BatchNorm2d(nf, eps=1e-5, momentum=0.1))
return nn.Sequential(*layers)
learn,run = get_learn_run(nfs, data, 0.4, conv_layer, cbs=cbfs)
# %time run.fit(1, learn)
# ## Running Batch Norm
class RunningBatchNorm(nn.Module):
def __init__(self, nf, mom=0.1, eps=1e-5):
super().__init__()
self.mom, self.eps = mom, eps
self.mults = nn.Parameter(torch.ones(nf,1,1))
self.adds = nn.Parameter(torch.zeros(nf,1,1))
self.register_buffer('sums', torch.zeros(1,nf,1,1))
self.register_buffer('sqrs', torch.zeros(1,nf,1,1))
self.register_buffer('batch', tensor(0.))
self.register_buffer('count', tensor(0.))
self.register_buffer('step', tensor(0.))
self.register_buffer('dbias', tensor(0.))
def forward(self, x):
if self.training: self.update_stats(x)
sums = self.sums
sqrs = self.sqrs
c = self.count
if self.step < 100:
sums /= self.dbias
sqrs /= self.dbias
c /= self.dbias
means = sums / c
vars = (sqrs / c).sub_(means*means)
if bool(self.batch < 20): vars.clamp_min_(0.01)
x = (x - means).div_((vars.add_(self.eps)).sqrt())
return x.mul_(self.mults).add_(self.adds)
def update_stats(self, x):
bs, nc, *_ = x.shape
self.sums.detach_()
self.sqrs.detach_()
dims = (0, 2, 3)
s = x.sum(dims, keepdim=True)
ss = (x*x).sum(dims, keepdim=True)
c = self.count.new_tensor(x.numel() / nc)
mom1 = 1 - (1 - self.mom) / math.sqrt(bs-1) # < 0.9 for self.mom=0.1
self.mom1 = self.dbias.new_tensor(mom1)
self.sums.lerp_(s, self.mom1)
self.sqrs.lerp_(ss, self.mom1)
self.count.lerp_(c, self.mom1)
self.dbias = self.dbias * (1 - self.mom1) + self.mom1
self.batch += bs
self.step += 1
def conv_rbn(ni, nf, ks=3, stride=2, bn=True, **kwargs):
layers = [nn.Conv2d(ni, nf, ks, padding=ks//2, stride=stride, bias=not bn),
GeneralRelu(**kwargs)]
if bn: layers.append(RunningBatchNorm(nf))
return nn.Sequential(*layers)
data = DataBunch(*get_dls(train_ds, valid_ds, 32), c)
learn,run = get_learn_run(nfs, data, 0.4, conv_rbn, cbs=cbfs)
# %time run.fit(1, learn)
# # What can we do in a single epoch?
learn, run = get_learn_run(nfs, data, 0.9, conv_rbn, cbs=cbfs + [partial(ParamScheduler, 'lr', sched_lin(1., 0.2))])
# %time run.fit(1, learn)
# !python notebook2script.py 07_batchnorm.ipynb
| nbs/dl2/selfmade/07_batchnorm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ### Save to Ocean Data View file
# *Load a biofloat DataFrame, apply WOA calibrated gain factor, and save it as an ODV spreadsheet*
# Use the local cache file for float 5903891 that drifted around ocean station Papa. It's the file that was produced for [compare_oxygen_calibrations.ipynb](compare_oxygen_calibrations.ipynb).
from biofloat import ArgoData, converters
from os.path import join, expanduser
ad = ArgoData(cache_file=join(expanduser('~'),'6881StnP_5903891.hdf'), verbosity=2)
wmo_list = ad.get_cache_file_all_wmo_list()
df = ad.get_float_dataframe(wmo_list)
# Show top 5 records.
df.head()
# Remove NaNs and apply the gain factor from [compare_oxygen_calibrations.ipynb](compare_oxygen_calibrations.ipynb).
corr_df = df.dropna().copy()
corr_df['DOXY_ADJUSTED'] *= 1.12
corr_df.head()
# Convert to ODV format and save in a .txt file.
converters.to_odv(corr_df, '6881StnP_5903891.txt')
# Import as an ODV Spreadsheet and use the tool.
from IPython.display import Image
Image('../doc/screenshots/Screen_Shot_2015-11-25_at_1.42.00_PM.png')
| notebooks/save_to_odv.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.1.0
# language: julia
# name: julia-1.1
# ---
using Distributed
addprocs(8)
# +
# %%
@everywhere begin
using Pkg; Pkg.activate("..")
# Pkg.instantiate()
using GraphFusedLasso
# using Base.Threads
# using Revise
using Distributions
import StatsBase.weights
using Random
using RCall
using ProgressBars
using FileIO
using DataFrames, CSV
using Printf
# using Base.Threads
end
# +
println("Running with $(Threads.nthreads()) thread(s)")
# println("Running with $(nprocs()) process(ses)")
R"library('tidyverse')"
# Random.seed!(418916);
@everywhere begin
# %%
function generate_trace(task, N)
cuts = [N ÷ 3, 2(N ÷ 3)]
# cuts = sort(sample(2:N-1, 2, replace=false))
x1 = 1:cuts[1]
x2 = (cuts[1] + 1):cuts[2]
x3 = (cuts[2] + 1):N
x = [x1; x2; x3]
values1 = 2.0(rand(Uniform(), 4) .- 0.5)
if task == "smooth"
y1 = values1[1] .+ (values1[2] - values1[1]) .* (x1 ./ cuts[1])
y2 = values1[2] .+ (values1[3] - values1[2]) .* (x2 .- cuts[1]) ./ (cuts[2] - cuts[1])
y3 = values1[3] .+ (values1[4] - values1[3]) .* (x3 .- cuts[2]) ./ (N - cuts[2])
elseif task == "constant"
y1 = fill(values1[1], cuts[1])
y2 = fill(values1[2], cuts[2] - cuts[1])
y3 = fill(values1[3], N - cuts[2])
elseif task == "mixed"
y1 = values1[1] .+ Int(rand() < 0.5) .* (values1[2] - values1[1]) .* (x1 ./ cuts[1])
y2 = values1[2] .+ Int(rand() < 0.5) .* (values1[3] - values1[2]) .* (x2 .- cuts[1]) ./ (cuts[2] - cuts[1])
y3 = values1[3] .+ Int(rand() < 0.5) .* (values1[4] - values1[3]) .* (x3 .- cuts[2]) ./ (N - cuts[2])
else
throw(ArgumentError)
end
μ = [y1; y2; y3]
return μ
end
function generate_spt_task(task_space, task_time, N, pmiss; σ=0.3, outliers=false)
μ1s = generate_trace(task_space, N)
μ2s = generate_trace(task_space, N)
ts = generate_trace(task_time, N)
evalpts = collect(range(-2.5, 2.5, length=100))
μs = [(t + μ1, t + μ2) for (μ1, μ2) in zip(μ1s, μ2s), t in ts]
dmodels = [MixtureModel([Normal(μ1, σ), Normal(μ2, σ)]) for (μ1, μ2) in μs]
devals = [pdf.(d, evalpts) for d in dmodels];
ndata = [sample([0, 10], weights([pmiss, 1.0 - pmiss])) for d in dmodels]
y = [rand(d, n) for (d, n) in zip(dmodels, ndata)]
if outliers
Nobs = sum([1 for n in ndata if n > 0])
K = Int(floor(Nobs * 0.5))
idx = sample([i for (i, n) in enumerate(ndata) if n > 0], K, replace=false)
for i in idx
j = rand(1:length(y[i]))
y[i][j] += rand([-1, 1]) * 5.0
end
end
# make matrix pts
xrange = collect(1:N)
# temporal
ptr = Int[]
brks = Int[1]
for i in 1:N
append!(ptr, xrange .+ (i - 1) * N)
push!(brks, brks[end] + N)
end
istemporal = fill(true, N^2)
# spatial
xrange = [(i - 1) * N + 1 for i in 1:N]
for i in 1:N
append!(ptr, xrange .+ (i - 1))
push!(brks, brks[end] + N)
end
append!(istemporal, fill(false, N^2))
return Dict("evalpts" => evalpts,
"dmodels" => dmodels,
"devals" => devals,
"y" => y,
"ndata" => ndata,
"mean1" => μ1s,
"mean2" => μ2s,
"t" => ts,
"means" => μs,
"ptr" => ptr,
"brks" => brks,
"istemporal" => istemporal)
end
# function for cross-validation fit
function generate_cvsets(y, nsplits)
# make the cv splits
N = length(y)
cvsets = [Set{Int}() for i in 1:nsplits]
iobs = shuffle([i for (i, yi) in enumerate(y) if !isempty(yi)])
Nobs = length(iobs)
splitsize = Nobs ÷ nsplits
for k in 1:nsplits
for i in ((k - 1) * splitsize + 1):(k * splitsize)
push!(cvsets[k], iobs[i])
end
end
return cvsets
end
function fit2(ytrain, ptr, brks, λ1, λ2, η1, η2, istemporal)
N = length(ytrain)
lambdasl1 = Float64[temp ? η1 : λ1 for temp in istemporal]
lambdasl2 = Float64[temp ? η2 : λ2 for temp in istemporal]
# create the tree
M = 33
splits = collect(range(-2.5, 2.5, length=M))
tree = DensityTree(splits)
bins2counts = Dict()
for (j, (li, ui)) in enumerate([tree.bins; [(i, i+1) for i in 1:M-1]])
lower = splits[li]
upper = splits[ui]
k = [sum(lower .< yi .< upper) for yi in ytrain]
bins2counts[(li, ui)] = k
end
# fit binomial model in each tree
beta = zeros(N, M - 2)
for j in 1:M - 2
li, ui = tree.bins[j]
mi = (ui + li) ÷ 2
parent_counts = bins2counts[(li, ui)] .+ 0.1
left_counts = bins2counts[(li, mi)] .+ 0.05
level = trunc(Int, log2(j))
model = BinomialEnet(
ptr, brks,
lambdasl1 .* 2.0 ^ (0.25 * level),
lambdasl2 .* 2.0 ^ (0.25 * level);
abstol=0.0,
reltol=1e-3)
fit!(model, left_counts, parent_counts; steps=100, parallel=false)
beta[:, j] = model.beta
end
tree.beta = beta
return tree
end
function cv_fit2(y, evalpts, ptr, brks, istemporal, lambdas, cvsets, models)
# for each cv split get the mse error
N = length(y)
nsplits = length(cvsets)
nlambdas = length(lambdas)
# prepare the tree structure and the bint
test_loglikelihood = pmap(1:nlambdas) do k
λ1, λ2, η1, η2 = lambdas[k]
Ntest = 0.0
loglikelihood = 0.0
for i = 1:nsplits
# get the cv vector with missing data
ytrain = [j in cvsets[i] ? Float64[] : yi
for (j, yi) in enumerate(y)]
tree = fit2(ytrain, ptr, brks, λ1, λ2, η1, η2, istemporal)
# compute the out-of-sample likelihood
for j in collect(cvsets[i])
test_eval = y[j]
Ntest += length(test_eval)
ll = - log.(predict(tree, sort(test_eval), j) .+ 1e-4)
loglikelihood += sum(ll)
end
end
loglikelihood / Ntest
end
# println(".")
# now choose the best lambdas
# display(collect(zip(test_loglikelihood, lambdas)))
best_lambdas = lambdas[argmin(test_loglikelihood)]
best_loglikelihood = minimum(test_loglikelihood)
# compute validation likelihood
nsims = 100
samples = [rand(model, nsims) for model in models]
λ1, λ2, η1, η2 = best_lambdas
tree = fit2(y, ptr, brks, λ1, λ2, η1, η2, istemporal)
lls = [- mean(log.(predict(tree, sort(x), j) .+ 1e-4)) for (j, x) in enumerate(samples)]
validation_loglikelihood = mean(lls)
return Dict("best_lambdas" => best_lambdas,
"cv_loglikelihood" => best_loglikelihood,
"val_loglikelihood" => validation_loglikelihood)
end
function get_hypers()
# fl_choices = [1e-12, 0.33, 0.66, 1.0]
# kal_choices = [1e-12, 0.75, 1.5, 3.0]
fl_choices = [1e-6, 0.33, 0.66, 1.0] ./ 2.0
kal_choices = [1e-6, 0.75, 1.5, 3.0] ./ 2.0
lambdas_dict = Dict(
"fl" => [(λ1, 1e-12, η1, 1e-12) for λ1 in fl_choices for η1 in fl_choices],
"kal" => [(1e-12, λ2, 1e-12, η2) for λ2 in kal_choices for η2 in kal_choices],
"enet" => [(λ1, λ2, η1, η2) for λ1 in fl_choices for λ2 in kal_choices
for η1 in fl_choices for η2 in kal_choices])
return lambdas_dict
end
function run_benchmarks(N, pmiss;
nsims=100,
nsplits=5,
tasks=("constant", "smooth", "mixed"),
outliers=false)
experiment_results = []
for task_space in tasks
for task_time in tasks
data = [
generate_spt_task(task_space, task_time, N, pmiss, outliers=outliers)
for _ in 1:nsims
]
lambdas_dict = get_hypers()
for method in ("fl", "kal", "enet")
println("Running task_space $task_space task_time $task_time for method $method, outliers $outliers")
new_records = []
for (l, D) in ProgressBar(collect(enumerate(data)))
# for (l, D) in enumerate(data)
y = vec(D["y"])
models = vec(D["dmodels"])
ndata = vec(D["ndata"])
devals = vec(D["devals"])
ptr = D["ptr"]
brks = D["brks"]
evalpts = D["evalpts"]
istemporal = D["istemporal"]
cvsets = generate_cvsets(y, nsplits)
lambdas = lambdas_dict[method]
results = cv_fit2(y, evalpts, ptr, brks, istemporal, lambdas, cvsets, models)
λ1, λ2, η1, η2 = results["best_lambdas"]
record = Dict(
:experiment => l,
:task_space => task_space,
:task_time => task_time,
:method => method,
:cv_ll => results["cv_loglikelihood"],
:val_ll => results["val_loglikelihood"],
:lamb1 => λ1,
:lamb2 => λ2,
:eta1 => η1,
:eta2 => η2,
:outliers => Int(outliers))
push!(new_records, record)
display(record)
end
append!(experiment_results, new_records)
end
end
end
return experiment_results
end
end
# +
# %%
# @everywhere begin
N = 30
nsims = 10
tasks = ("smooth", "constant", "mixed")
# end
pmiss = 0.8
experiment_results = run_benchmarks(N, pmiss, nsims=nsims, tasks=tasks)
experiment_results_out = run_benchmarks(N, pmiss, nsims=nsims, tasks=("mixed", ), outliers=true)
# +
# %%
df = DataFrame(experiment = Int[],
task_space=String[],
task_time=String[],
method=String[],
cv_ll=Float64[],
val_ll=Float64[],
lamb1=Float64[],
lamb2=Float64[],
eta1=Float64[],
eta2=Float64[],
outliers=Int[])
for record in [experiment_results; experiment_results_out]
push!(df, record)
end
CSV.write("benchmarks-results-spt_p08_scaled.csv", df)
# %%
head(df, 10)
# %%
res = R"""
df = $df %>%
group_by(task_space, task_time, method, outliers) %>%
summarize(val_loglikelihood = mean(val_ll),
cv_loglikelihood = mean(cv_ll))
df
"""
res = rcopy(res)
# %%
# -
print(df)
# +
# %%
# @everywhere begin
N = 30
nsims = 10
tasks = ("smooth", "constant", "mixed")
# end
pmiss = 0.1
experiment_results = run_benchmarks(N, pmiss, nsims=nsims, tasks=tasks)
experiment_results_out = run_benchmarks(N, pmiss, nsims=nsims, tasks=("mixed", ), outliers=true)
# +
# %%
df = DataFrame(experiment = Int[],
task_space=String[],
task_time=String[],
method=String[],
cv_ll=Float64[],
val_ll=Float64[],
lamb1=Float64[],
lamb2=Float64[],
eta1=Float64[],
eta2=Float64[],
outliers=Int[])
for record in [experiment_results; experiment_results_out]
push!(df, record)
end
CSV.write("benchmarks-results-spt_p01_scaled.csv", df)
# %%
head(df, 10)
# %%
res = R"""
df = $df %>%
group_by(task_space, task_time, method, outliers) %>%
summarize(val_loglikelihood = mean(val_ll),
cv_loglikelihood = mean(cv_ll))
df
"""
res = rcopy(res)
# %%
# -
print(df)
| simulations/benchmarks/simulation-automated.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#default_exp ranker
# -
#hide
from nbdev.showdoc import *
# # Ranker
#
# Takes a query and an index and finds the nearest neighbors or most similar scores. Ideally this is just a simple Annoy `get_nns_by_vector`, or in the simple case a similarity score across all the vectors.
# +
import torch
from pathlib import Path
from memery.loader import treemap_loader, db_loader
from memery.encoder import text_encoder
# -
treemap = treemap_loader(Path('./images/memery.ann'))
if treemap:
treemap.get_n_items()
#export
def ranker(query_vec, treemap):
nn_indexes = treemap.get_nns_by_vector(query_vec[0], treemap.get_n_items())
return(nn_indexes)
#export
def nns_to_files(db, indexes):
# return([[v['fpath'] for k,v in db.items() if v['index'] == ind][0] for ind in indexes])
return([db[ind]['fpath'] for ind in indexes])
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
db = db_loader(Path('images/memery.pt'), device)
query = 'dog'
query_vec = text_encoder(query, device)
indexes = ranker(query_vec, treemap)
ranked_files = nns_to_files(db, indexes)
ranked_files[:5]
| notebooks/05_ranker.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={}
# # CLT vs PJ
# + pycharm={"is_executing": false}
import pandas as pd
import DescontosBeneficiosCLT as CLT
import DescontosPJ as PJ
# -
# ## Introdução
# Este documento visa apresentar um comparativo entre os valores líquidos recebidos entre as modalidades de contratação CLT e PJ. Auxiliando profissionais (principalmente na área de TI) a avaliar propostas de trabalho em diferente modalidades. Auxiliando também na compreensão dos direitos e deveres associados a cada uma das modalidades de contratação.
#
# Para os cálculos apresentados serão apresentados resultados para os diferentes métodos de pagamento do Imposto de Renda sobre Pessoa Física (pagamento simplificado e completo).
#
# Os resultados apresentados são estimativas e podem variar por diversos fatores: numero de hora trabalhadas (PJ), abatimentos nos cálculos do IR, reajustes, ....
# \* *É permitido a livre divulgação, modificação e utilização deste documento e deste repositório de acordo com os termos de [licença](files/LICENSE) MIT. Caso queira contribuir com melhorias e correções, faça-o através do repositório no Github.*
# + [markdown] pycharm={}
# ## Parâmetros
# -
# Apenas as variáveis abaixo (dentro da seção "Parâmetros") precisam ser alteradas. Não há necessidade de alterar nenhum outro campo dentro de outras seções deste documento.
# Informações referentes à modalidade **CLT**:
# +
salario_mensal_bruto = 10000 # Salário bruto recebido mensalmente na modalidade CLT
PLR = 2*salario_mensal_bruto # Valor da PLR ou Bonus anual recebidos na modalidade CLT
VA_VR = 1500.00 # Valor mensal de Vale Alimentação e Vale Refeição somados
# -
# Informações referentes à modalidade **PJ**. Normalmente, para a modalidade PJ o contrato estipula o pagamento por horas trabalhadas, dessa forma é solicitado que seja informado o salário/hora.
#
# Na modalidade PJ, férias normalmente não são remuneradas, dessa forma deve-se informar o número de dias de férias no ano para que não seja considerado nenhum faturamento no período de férias.
#
# Deve-se considerar também custos relacionados como contratação de Contados, custo de aluguel de escritório (físico ou virtual), contratação de plano de saúde (para que a comparação fique justa com a proposta CLT que oferece plano de saúde), ...
# +
salario_PJ_hora = 100.00 # Salário/hora (R$/h) na modalidade PJ
horas_mes = 170 # Número de horas trabalhadas no mês
dias_ferias = 30 # Número de dias corridos de férias por ano (não remunerados na modalidade PJ)
contador = 100.00 # Custos mensais com contador
outros_custos = 0.00 # Outros custos mensais para operação da empresa e.g.: aluguel de escritório, custos com equipamentos, ...
seguro_saude = 0.00 # Custo mensal do Plano de saúde de saúde que precisa serm contratado a parte.
# -
# Informações complementares
numero_dependentes = 1 # Número de dependentes para abatimento no IR - será utilizado tanto para cálculo CLT quanto PJ
# + [markdown] pycharm={}
# ## Cálculos CLT
# -
# Cálculo dos descontos e benefícios na modalidade CLT
# #### INSS (mensal)
INSS_mensal = CLT.INSS(salario_mensal_bruto)
print(f"Valor mensal do INSS: R$ {INSS_mensal:,.2f}")
# #### Imposto de Renda mensal sobre o salário (mensal)
# +
salario_base = salario_mensal_bruto - INSS_mensal # Salário Base para cálculo do IR, com os devidos descontos
IR_mensal_simplificado = CLT.IR_Mensal_Simplificado(salario_mensal_bruto) # Opção de IR sobre salário com cálculo simplificado
IR_mensal_completo = CLT.IR_Mensal(salario_base, numero_dependentes) # Opção de IR sobre salário com cálculo completo
print(f"IR Simplificado sobre o salário: R$ {IR_mensal_simplificado:,.2f}")
print(f"IR Completo sobre o salário: R$ {IR_mensal_completo:,.2f}")
# -
# #### FGTS
# Para uma comparação justa com a modalidade PJ, o rendimento do FGTS não será considerado. Além disso, é importante salientar que o valor recebido via FGTS não é totalmente líquido, tendo sua utilização e resgate condicionados à ocasiões estabelecidas pela legislação.
#
# Os cálculos abaixo também não consideram os optantes pela modalidade de Saque-Aniversário, dado que nesta modalidade sim há o resgate parcial de forma líquida deste benefício.
# +
numero_meses = 1 # número de meses a considerar
incluir_multa = False # Incluir o valor de 40% de Multa de demissão sem justa causa?
incluir_rendimento = False # Incluir o cálculo do rendimento de juros do FGTS?
FGTS_mensal = CLT.FGTS(salario_mensal_bruto, numero_meses, incluir_multa, incluir_rendimento)
print(f"Valor mensal do FGTS: R$ {FGTS_mensal:,.2f}")
# -
# #### Salário de Férias
# Neste cálculo é considerado o salário de férias no valor de 1+1/3 salário. E em seguida já é realizado o cálculo dos devidos descontos.
# +
salario_ferias = CLT.Salario_Ferias_Bruto(salario_mensal_bruto)
INSS_ferias = CLT.INSS(salario_ferias) # Desconto de INSS no salário de férias
salario_base_ferias = salario_ferias - INSS_ferias # Salário de Férias Base para cálculo do IR, com os devidos descontos
IR_ferias_completo = CLT.IR_Mensal(salario_base_ferias, numero_dependentes) # Opção de IR sobre ferias com cálculo completo
IR_ferias_simplificado = CLT.IR_Mensal_Simplificado(salario_ferias) # Opção de IR sobre ferias com cálculo simplificado
print(f"Valor do salário de férias bruto: R$ {salario_ferias:,.2f}")
print(f"INSS sobre o salário de férias: R$ {INSS_ferias:,.2f}")
print(f"IR Simplificado sobre o salário de férias: R$ {IR_ferias_simplificado:,.2f}")
print(f"IR Completo sobre o salário de férias: R$ {IR_ferias_completo:,.2f}")
# -
# #### 13º Salário
# +
salario_13_bruto = salario_mensal_bruto
INSS_13o = CLT.INSS(salario_13_bruto) # Desconto de INSS no 13º salário
salario_base_13o = salario_13_bruto - INSS_13o # 13o Salário Base para cálculo do IR, com os devidos descontos
IR_13o_completo = CLT.IR_Mensal(salario_base_13o, numero_dependentes) # Opção de IR sobre 13º com cálculo completo
IR_13o_simplificado = CLT.IR_Mensal_Simplificado(salario_13_bruto) # Opção de IR sobre 13º com cálculo simplificado
print(f"Valor bruto do 13º salário: R$ {salario_13_bruto:,.2f}")
print(f"INSS sobre o 13º salário: R$ {INSS_13o:,.2f}")
print(f"IR Simplificado sobre o 13º salário: R$ {IR_13o_simplificado:,.2f}")
print(f"IR Completo sobre o 13º salário: R$ {IR_13o_completo:,.2f}")
# -
# #### PLR / Bônus
# Cálculo dos descontos aplicados sobre a PLR
# +
IR_PLR = CLT.IR_PLR(PLR)
print(f"IR Completo sobre o PLR anual: R$ {IR_PLR:,.2f}")
# + [markdown] pycharm={}
# ### Resumo dos Cálculos Anuais
# -
# Para o cálculo anual serão considerados os sálarios de 11 meses de trabalho mais 1 mês de férias. Os valores podem variar dentro do primeiro ano de contratação, dado que neste período o funcinário não possui o benefício das férias.
# + pycharm={"is_executing": false}
# 11 meses de trabalho (1 mês de férias)
salario_anual_liq_simp = (salario_mensal_bruto - INSS_mensal - IR_mensal_simplificado) * 11 # Salário líquido anual com IR Simplificado
salario_anual_liq_comp = (salario_mensal_bruto - INSS_mensal - IR_mensal_completo) * 11 # Salário líquido anual com IR Completo
salario_13_liq_simp = salario_13_bruto - INSS_13o - IR_13o_simplificado # 13º Salário Líquido com desconto de IR Simplificado
salario_13_liq_comp = salario_13_bruto - INSS_13o - IR_13o_completo # 13º Salário Líquido com desconto de IR Completo
salario_ferias_liq_simp = salario_ferias - INSS_ferias - IR_ferias_simplificado # Salário de Férias Líquido com desconto de IR Simplificado
salario_ferias_liq_comp = salario_ferias - INSS_ferias - IR_ferias_completo # Salário de Férias Líquido com desconto de IR Completo
PLR_liquido = PLR - IR_PLR # Valor da PLR líquido no ano
# +
# Salario + benefícios Brutos
CLT_bruto_anual = salario_mensal_bruto*12 + salario_ferias + PLR + FGTS_mensal*(13+1/3) + VA_VR*12
# Salario + benefícios Líquidos
CLT_liquido_anual_simp = salario_anual_liq_simp + salario_13_liq_simp + salario_ferias_liq_simp + PLR_liquido + FGTS_mensal*(13+1/3) + VA_VR*12
CLT_liquido_anual_comp = salario_anual_liq_comp + salario_13_liq_comp + salario_ferias_liq_comp + PLR_liquido + FGTS_mensal*(13+1/3) + VA_VR*12
print(f"Salário mensal bruto: R$ {salario_mensal_bruto:,.2f}") # Sálario mensal Bruto
print(f"Salário anual bruto: R$ {CLT_bruto_anual:,.2f}") # Salario anual Bruto
print(f"Salário anual líquido + benefícios (IR Simplificado): R$ {CLT_liquido_anual_simp:,.2f}") # Salario Líquido + benefícios
print(f"Salário anual líquido + benefícios (IR Completo): R$ {CLT_liquido_anual_comp:,.2f}") # Salario Líquido + benefícios
# + pycharm={"is_executing": false}
# Agregando os resultados dos cálculos para CLT Simplificado
resultado_CLT_Simp = [] # Montar variável com resultados CLT Simplificado
resultado_CLT_Simp.append(f'R$ {salario_mensal_bruto:,.2f} /mês') # Ref
resultado_CLT_Simp.append(salario_mensal_bruto) # Salário Mensal
resultado_CLT_Simp.append(salario_mensal_bruto-INSS_mensal-IR_mensal_simplificado) # Salário Mensal Líquido
resultado_CLT_Simp.append((salario_mensal_bruto-INSS_mensal-IR_mensal_simplificado)*12) # Salário Anual Líquido
resultado_CLT_Simp.append(CLT_liquido_anual_simp) # Liquido + Benefícios (Anual)
# +
# Agregando os resultados dos cálculos para CLT Completo
resultado_CLT_Comp = [] # Montar variável com resultados CLT Completo
resultado_CLT_Comp.append(f'R$ {salario_mensal_bruto:,.2f} /mês') # Ref
resultado_CLT_Comp.append(salario_mensal_bruto) # Salário Mensal
resultado_CLT_Comp.append(salario_mensal_bruto-INSS_mensal-IR_mensal_completo) # Salário Mensal Líquido
resultado_CLT_Comp.append((salario_mensal_bruto-INSS_mensal-IR_mensal_completo)*12) # Salário Anual Líquido
resultado_CLT_Comp.append(CLT_liquido_anual_comp) # Liquido + Benefícios (Anual)
# + [markdown] pycharm={}
# ## PJ SIMPLES NACIONAL (ANEXO III)
# -
# Nesta seção serão considerados apenas os cálculos para empresas que se enquadram no Anexo III do Simples Nacional.
#
# Para que prestadores de serviço possam se enquadrar nessa modalidade é necessário que o fator R (cálculado abaixo) esteja dentro do limite estabelecido.
# #### Faturamento mensal e anual PJ
# +
receita_mensal_PJ = salario_PJ_hora * horas_mes # Faturamento mensal PJ
meses_ferias = dias_ferias / 30 # Converte o numero de dias de férias para mêses para auxiliar no cálculo anual. Está aproximado o valor de 30 dias em um mês.
receita_anual_PJ = receita_mensal_PJ * (12 - meses_ferias) # Cáculo da receita anual descontado os dias de férias não remuneradas
print(f"Faturamento mensal PJ: R$ {receita_mensal_PJ:,.2f}") # Faturamento mensal PJ
print(f"Faturamento mensal PJ: R$ {receita_anual_PJ:,.2f}") # Faturamento anual PJ
# -
# #### Cálculo do Pro-Labore
# Para enquadramento de prestadores de serviço no Anexo III do Simples Nacional, a empresa deve possuir um Fator R acima de **0.28** (ou **28%**).
# O fator R é calculado dividindo o Pro-Labore pela receita bruta.
#
# O Pró-Labore é a remunueração dos sócios da empresa\*, neste documento estamos considerando um sócio único (uma única pessoa trabalhando dentro da modalidade PJ).
#
# Há incidência de Imposto de Renda sobre a remuneração do Pró-Labore, dessa forma, é desejável mantê-lo próximo ao limite permitido. Para os cálculos a seguir será considerado um Fator R de 30% (como margem de segurança).
#
# \* *Além do Pró-Labore, os demais rendimentos líquidos da empresa serão repassados ao sócios como **dividendos**, dado que por enquanto ainda não há incidência de Imposto sobre dividendos.*
# +
pro_labore = PJ.ProLabore_FatorR(receita_mensal_PJ, 0.30) # Cálculo do valor mensal do Pro-labore, utilizando fator R de 30%
print(f"Valor do Pro-Labore mensal: R$ {pro_labore:,.2f}") # Pro-Labore mensal
# -
# #### Recolhimento de imposto DAS.
# A DAS é uma via única de recolhimento de imposto para PJ.
#
# O valor anual da DAS durante os primeiros 12 meses de atividade da empresa pode vir acima do valor calculado a seguir, devido ao cálculo proporcionalizado durante os 12 primeiros meses. Neste caso é possivel solicitar a restituição de pagamento a maior junto à Receita Federal.
# +
imposto_DAS_anual = PJ.DAS_SimplesNacionalIII(receita_anual_PJ) # Cálculo do imposto a ser recolhido via DAS
print(f"Imposto DAS anual: R$ {imposto_DAS_anual:,.2f}") # Pro-Labore mensal
# -
# #### Impostos Pessoa Física -- Pro-Labore
# Além da incidência de impostos sobre a Pessoa Jurídica, há tambéma incidência sobre a Pessoa Física, sobre o valor do Pró-Labore.
# +
### INSS sobre pro-labore 11%
INSS_prolabore_mensal = pro_labore * 0.11 # INSS sobre Pró-labore 11%
print(f"Valor mensal do INSS: R$ {INSS_prolabore_mensal:,.2f}")
## IRRF sobre pro-labore
prolabore_base = pro_labore - INSS_prolabore_mensal
IR_prolabore_mensal_simplificado = CLT.IR_Mensal_Simplificado(prolabore_base) # Opção de IR sobre pró-labore com cálculo simplificado
IR_prolabore_mensal_completo = CLT.IR_Mensal(prolabore_base, numero_dependentes) # Opção de IR sobre pró-labore com cálculo completo
print(f"IR Simplificado sobre o pró-labore: R$ {IR_prolabore_mensal_simplificado:,.2f}")
print(f"IR Completo sobre o pró-labore: R$ {IR_prolabore_mensal_completo:,.2f}")
# +
rendimento_liquido_anual_PJ_simpl = receita_anual_PJ - imposto_DAS_anual - contador - outros_custos - seguro_saude - (INSS_prolabore_mensal + IR_prolabore_mensal_simplificado) * (12 - meses_ferias)
rendimento_liquido_anual_PJ_compl = receita_anual_PJ - imposto_DAS_anual - contador - outros_custos - seguro_saude - (INSS_prolabore_mensal + IR_prolabore_mensal_completo) * (12 - meses_ferias)
print(f"Rendimento líquido anual para PJ (com IR simplificado sobre Pró-Labore): R$ {rendimento_liquido_anual_PJ_simpl:,.2f}")
print(f"Rendimento líquido anual para PJ (com IR completo sobre Pró-Labore): R$ {rendimento_liquido_anual_PJ_compl:,.2f}")
# +
# Agregando os resultados dos cálculos para PJ Simplificado
resultado_PJ_Simp = [] # Montar variável com resultados PJ Simplificado
resultado_PJ_Simp.append(f'R$ {salario_PJ_hora:,.2f} /hora') # Ref
resultado_PJ_Simp.append(receita_mensal_PJ) # Salário Mensal
resultado_PJ_Simp.append(receita_mensal_PJ-INSS_prolabore_mensal-IR_prolabore_mensal_simplificado-imposto_DAS_anual/12) # <NAME>
resultado_PJ_Simp.append(rendimento_liquido_anual_PJ_simpl) # <NAME>
resultado_PJ_Simp.append(rendimento_liquido_anual_PJ_simpl) # Liquido + Benefícios (Anual)
# +
# Agregando os resultados dos cálculos para PJ Completo
resultado_PJ_Comp = [] # Montar variável com resultados PJ Completo
resultado_PJ_Comp.append(f'R$ {salario_PJ_hora:,.2f} /hora') # Ref
resultado_PJ_Comp.append(receita_mensal_PJ) # <NAME>
resultado_PJ_Comp.append(receita_mensal_PJ-INSS_prolabore_mensal-IR_prolabore_mensal_completo-imposto_DAS_anual/12) # <NAME>
resultado_PJ_Comp.append(rendimento_liquido_anual_PJ_compl) # Sal<NAME>
resultado_PJ_Comp.append(rendimento_liquido_anual_PJ_compl) # Liquido + Benefícios (Anual)
# + [markdown] pycharm={}
# # Resultado
# -
# A tabela apresenta o resultado comparativo entre as opções de contratação CLT e PJ. Para cada uma das modalidades são consideradas as diferentes formas de tributação (Simplificada e Completa).
#
# O resultado aparecerá ordenado pelo valor Líquido + Benefícios (Anual) de forma decescente.
#
# Reforçando que os resultados apresentados são uma estimativa e podem variar na remuneração real.
# +
# Parâmetros, opções e formatação da tabela
table_columns = ["Ref", "Salário Mensal", "Salário Líquido Mensal","Salário Líquido Anual","Líquido + Benefícios (Anual)"]
table_index = ['CLT (Simplificado)', 'CLT (Completo)', 'PJ (Simplificado)', 'PJ (Completo)']
pd.options.display.max_columns = None
pd.options.display.float_format = 'R$ {:,.2f}'.format # Format float numbers in dataTable
# +
df = pd.DataFrame([resultado_CLT_Simp, resultado_CLT_Comp, resultado_PJ_Simp, resultado_PJ_Comp],
columns=table_columns,
index=table_index)
# Novas colunas calculadas
df['Diferença Anual'] = df['Líquido + Benefícios (Anual)'] - df['Líquido + Benefícios (Anual)'].max() # Diferença para o maior valor
df['Equivalente Mensal Líquido'] = df['Líquido + Benefícios (Anual)']/12 # Define column Equivalente Mensal
df = df.sort_values(by='Líquido + Benefícios (Anual)', ascending=False)
display(df)
# -
# _**Legenda:**_
#
# | Coluna | Descrição |
# | :---------- | :---------- |
# | Ref | *Referência da linha* |
# | Salário Mensal | *Salário (ou receita) bruto mensal* |
# | Salário Líquido Mensal | *Salário (ou receita) mensal já descontados os impostos* |
# | Salário Líquido Anual | *Salário líquido mensal multiplicado pelo número de meses trabalhados (não incluso benefícios)* |
# | Liquido + Benefícios (Anual) | *Salário Líquido + Benefícios anuais já aplicados os descontos*|
# ## Referências
#
# https://fdr.com.br/calculadora/fgts/
#
# https://www.idinheiro.com.br/calculadoras/calculadora-fgts/
#
# https://www.contabilizei.com.br/contabilidade-online/irpf-2021-o-que-e-descontado-na-fonte-e-como-funciona-o-ajuste-anual/
#
# https://www.idinheiro.com.br/calculadoras/calculadora-de-ferias/
#
# https://www.contabilizei.com.br/contabilidade-online/desconto-inss/
#
# https://wkrh.com.br/impostos-profissional-pj/
#
# https://www.contratopj.com.br/quais-impostos-a-pagar-trabalhando-como-pj/
#
# https://www.contratopj.com.br/fator-r-simples-nacional/
#
| CLT_vs_PJ.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Setup rendering dependencies for Google Colaboratory.
# !pip install gym pyvirtualdisplay > /dev/null 2>&1
# !apt-get install -y xvfb python-opengl ffmpeg > /dev/null 2>&1
# Install d3rlpy!
# !git clone https://github.com/takuseno/d3rlpy
# !cd d3rlpy && pip install -e .
import site
site.main()
# Setup cartpole environment.
# +
import gym
env = gym.make('CartPole-v0')
eval_env = gym.make('CartPole-v0')
# -
# Setup data-driven deep reinforcement learning algorithm.
# +
from d3rlpy.algos import DQN
from d3rlpy.online.explorers import ConstantEpsilonGreedy
from d3rlpy.online.buffers import ReplayBuffer
from d3rlpy.metrics.scorer import evaluate_on_environment
# setup DQN algorithm
dqn = DQN(learning_rate=1e-3, target_update_interval=100, use_gpu=False)
# setup explorer
explorer = ConstantEpsilonGreedy(epsilon=0.3)
# setup replay buffer
buffer = ReplayBuffer(maxlen=50000, env=env)
# start training
dqn.fit_online(env,
buffer,
explorer,
eval_env=eval_env,
n_steps=50000,
n_steps_per_epoch=10000)
# -
# Setup rendering utilities for Google Colaboratory.
# +
import glob
import io
import base64
from gym.wrappers import Monitor
from IPython.display import HTML
from IPython import display as ipythondisplay
from pyvirtualdisplay import Display
# start virtual display
display = Display(visible=0, size=(1400, 900))
display.start()
# play recorded video
def show_video():
mp4list = glob.glob('video/*.mp4')
if len(mp4list) > 0:
mp4 = mp4list[0]
video = io.open(mp4, 'r+b').read()
encoded = base64.b64encode(video)
ipythondisplay.display(HTML(data='''
<video alt="test" autoplay loop controls style="height: 400px;">
<source src="data:video/mp4;base64,{0}" type="video/mp4" />
</video>'''.format(encoded.decode('ascii'))))
else:
print("Could not find video")
# -
# Record video!
# +
# wrap Monitor wrapper
env = Monitor(env, './video', force=True)
# evaluate
evaluate_on_environment(env)(dqn)
# -
# Let's see how it works!
show_video()
| tutorials/online.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Exercise 1 (Description of an outlier)
import matplotlib.pyplot as plt
from math import cos, pi
ys = [cos(i*(pi/4)) for i in range(50)]
plt.plot(ys)
ys[4] = ys[4] + 5.0
ys[20] = ys[20] + 8.0
plt.plot(ys)
# ### Exercise 2 (Box plot)
plt.boxplot(ys)
# ### Exercise 3 (Z score)
# !pip install scipy
from scipy import stats
cos_arr_z_score = stats.zscore(ys)
cos_arr_z_score
# ### Exercise 4: Removing outliers based on Z score.
import pandas as pd
df_original = pd.DataFrame(ys)
df_original
cos_arr_without_outliers = df_original[(cos_arr_z_score < 3)]
cos_arr_without_outliers.shape
df_original.shape
plt.plot(cos_arr_without_outliers)
# ### Exercise 5 (levenshtein distance)
# !pip install python-Levenshtein
ship_data = {"Sea Princess": {"date": "12/08/18", "load": 40000},
"Sea Pincess": {"date": "10/06/18", "load": 30000},
"Sea Princes": {"date": "12/04/18", "load": 30000},
}
from Levenshtein import distance
name_of_ship = "Sea Princess"
for k, v in ship_data.items():
print("{} {} {}".format(k, name_of_ship, distance(name_of_ship, k)))
| Lesson06/Exercise79-81/Identifying and cleaning outliers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="17jGSw2UD-Di" outputId="546ae7c4-e2f5-488d-869f-8a39618147ea"
# !pip install pywavelets
# + id="tMc1SjLgEk1K"
import numpy as np
import matplotlib.pyplot as plt
# + id="egQo6WoZFn-T"
from scipy.fftpack import fft
def get_fft_values(y_values, T, N, f_s):
f_values = np.linspace(0.0, 1.0/(2.0*T), N//2)
fft_values_ = fft(y_values)
fft_values = 2.0/N * np.abs(fft_values_[0:N//2])
return f_values, fft_values
# + colab={"base_uri": "https://localhost:8080/"} id="OcZthC7pGMu_" outputId="debc8650-8d2e-4e80-f1d9-addd0a1b6b9f"
import pywt
print(pywt.families(short=False))
# + colab={"base_uri": "https://localhost:8080/", "height": 619} id="JiOdmHkSGfmH" outputId="bf817a91-0e91-4b02-f3a0-60b7de17523e"
discrete_wavelets = ['db5', 'sym5', 'coif5', 'bior2.4']
continuous_wavelets = ['mexh', 'morl', 'cgau5', 'gaus5']
list_list_wavelets = [discrete_wavelets, continuous_wavelets]
list_funcs = [pywt.Wavelet, pywt.ContinuousWavelet]
fig, axarr = plt.subplots(nrows=2, ncols=4, figsize=(16,8))
for ii, list_wavelets in enumerate(list_list_wavelets):
func = list_funcs[ii]
row_no = ii
for col_no, waveletname in enumerate(list_wavelets):
wavelet = func(waveletname)
family_name = wavelet.family_name
biorthogonal = wavelet.biorthogonal
orthogonal = wavelet.orthogonal
symmetry = wavelet.symmetry
if ii == 0:
_ = wavelet.wavefun()
wavelet_function = _[0]
x_values = _[-1]
else:
wavelet_function, x_values = wavelet.wavefun()
if col_no == 0 and ii == 0:
axarr[row_no, col_no].set_ylabel("Discrete Wavelets", fontsize=16)
if col_no == 0 and ii == 1:
axarr[row_no, col_no].set_ylabel("Continuous Wavelets", fontsize=16)
axarr[row_no, col_no].set_title("{}".format(family_name), fontsize=16)
axarr[row_no, col_no].plot(x_values, wavelet_function)
axarr[row_no, col_no].set_yticks([])
axarr[row_no, col_no].set_yticklabels([])
plt.tight_layout()
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="NswNeARTGm76" outputId="886ec9f1-74a7-402a-ed94-1411ad3781a8"
import pywt
import matplotlib.pyplot as plt
db_wavelets = pywt.wavelist('db')[:5]
print(db_wavelets)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="SMQgjwKmG-Pb" outputId="db679630-d784-4d66-c708-ef5f1c06b7f2"
fig, axarr = plt.subplots(ncols=5, nrows=5, figsize=(20,16))
fig.suptitle('Daubechies family of wavelets', fontsize=16)
for col_no, waveletname in enumerate(db_wavelets):
wavelet = pywt.Wavelet(waveletname)
no_moments = wavelet.vanishing_moments_psi
family_name = wavelet.family_name
for row_no, level in enumerate(range(1,6)):
wavelet_function, scaling_function, x_values = wavelet.wavefun(level = level)
axarr[row_no, col_no].set_title("{} - level {}\n{} vanishing moments\n{} samples".format(
waveletname, level, no_moments, len(x_values)), loc='left')
axarr[row_no, col_no].plot(x_values, wavelet_function, 'bD--')
axarr[row_no, col_no].set_yticks([])
axarr[row_no, col_no].set_yticklabels([])
plt.tight_layout()
plt.subplots_adjust(top=0.9)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="6RnqWdPmHDhB" outputId="a09dce50-f063-422a-c7bf-d21f67a46932"
import pywt
x = np.linspace(0, 1, num=2048)
chirp_signal = np.sin(250 * np.pi * x**2)
fig, ax = plt.subplots(figsize=(6,1))
ax.set_title("Original Chirp Signal: ")
ax.plot(chirp_signal)
plt.show()
data = chirp_signal
waveletname = 'sym5'
fig, axarr = plt.subplots(nrows=5, ncols=2, figsize=(6,6))
for ii in range(5):
(data, coeff_d) = pywt.dwt(data, waveletname)
axarr[ii, 0].plot(data, 'r')
axarr[ii, 1].plot(coeff_d, 'g')
axarr[ii, 0].set_ylabel("Level {}".format(ii + 1), fontsize=14, rotation=90)
axarr[ii, 0].set_yticklabels([])
if ii == 0:
axarr[ii, 0].set_title("Approximation coefficients", fontsize=14)
axarr[ii, 1].set_title("Detail coefficients", fontsize=14)
axarr[ii, 1].set_yticklabels([])
plt.tight_layout()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="3AdaT5w7HPAo" outputId="8fe45c41-71ec-4dc7-bdd9-eaf7e98cbc96"
import pandas as pd
def plot_wavelet(time, signal, scales,
waveletname = 'cmor',
cmap = plt.cm.seismic,
title = 'Wavelet Transform (Power Spectrum) of signal',
ylabel = 'Period (years)',
xlabel = 'Time'):
dt = time[1] - time[0]
[coefficients, frequencies] = pywt.cwt(signal, scales, waveletname, dt)
power = (abs(coefficients)) ** 2
period = 1. / frequencies
levels = [0.0625, 0.125, 0.25, 0.5, 1, 2, 4, 8]
contourlevels = np.log2(levels)
fig, ax = plt.subplots(figsize=(15, 10))
im = ax.contourf(time, np.log2(period), np.log2(power), contourlevels, extend='both',cmap=cmap)
ax.set_title(title, fontsize=20)
ax.set_ylabel(ylabel, fontsize=18)
ax.set_xlabel(xlabel, fontsize=18)
yticks = 2**np.arange(np.ceil(np.log2(period.min())), np.ceil(np.log2(period.max())))
ax.set_yticks(np.log2(yticks))
ax.set_yticklabels(yticks)
ax.invert_yaxis()
ylim = ax.get_ylim()
ax.set_ylim(ylim[0], -1)
cbar_ax = fig.add_axes([0.95, 0.5, 0.03, 0.25])
fig.colorbar(im, cax=cbar_ax, orientation="vertical")
plt.show()
def get_ave_values(xvalues, yvalues, n = 5):
signal_length = len(xvalues)
if signal_length % n == 0:
padding_length = 0
else:
padding_length = n - signal_length//n % n
xarr = np.array(xvalues)
yarr = np.array(yvalues)
xarr.resize(signal_length//n, n)
yarr.resize(signal_length//n, n)
xarr_reshaped = xarr.reshape((-1,n))
yarr_reshaped = yarr.reshape((-1,n))
x_ave = xarr_reshaped[:,0]
y_ave = np.nanmean(yarr_reshaped, axis=1)
return x_ave, y_ave
def plot_signal_plus_average(time, signal, average_over = 5):
fig, ax = plt.subplots(figsize=(15, 3))
time_ave, signal_ave = get_ave_values(time, signal, average_over)
ax.plot(time, signal, label='signal')
ax.plot(time_ave, signal_ave, label = 'time average (n={})'.format(5))
ax.set_xlim([time[0], time[-1]])
ax.set_ylabel('Signal Amplitude', fontsize=18)
ax.set_title('Signal + Time Average', fontsize=18)
ax.set_xlabel('Time', fontsize=18)
ax.legend()
plt.show()
def get_fft_values(y_values, T, N, f_s):
f_values = np.linspace(0.0, 1.0/(2.0*T), N//2)
fft_values_ = fft(y_values)
fft_values = 2.0/N * np.abs(fft_values_[0:N//2])
return f_values, fft_values
def plot_fft_plus_power(time, signal):
dt = time[1] - time[0]
N = len(signal)
fs = 1/dt
fig, ax = plt.subplots(figsize=(15, 3))
variance = np.std(signal)**2
f_values, fft_values = get_fft_values(signal, dt, N, fs)
fft_power = variance * abs(fft_values) ** 2 # FFT power spectrum
ax.plot(f_values, fft_values, 'r-', label='Fourier Transform')
ax.plot(f_values, fft_power, 'k--', linewidth=1, label='FFT Power Spectrum')
ax.set_xlabel('Frequency [Hz / year]', fontsize=18)
ax.set_ylabel('Amplitude', fontsize=18)
ax.legend()
plt.show()
dataset = "http://paos.colorado.edu/research/wavelets/wave_idl/sst_nino3.dat"
df_nino = pd.read_table(dataset)
N = df_nino.shape[0]
t0=1871
dt=0.25
time = np.arange(0, N) * dt + t0
signal = df_nino.values.squeeze()
scales = np.arange(1, 128)
plot_signal_plus_average(time, signal)
plot_fft_plus_power(time, signal)
plot_wavelet(time, signal, scales)
| WaveletTransform.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pickle
# load the model from disk
loaded_model = pickle.load(open('rev2_dt_final.sav', 'rb'))
#result = loaded_model.score(X_test, Y_test)
#print(result)
# +
# Natural Language Processing
# Importing the libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# Importing the dataset
dataset = pd.read_csv('train.tsv', delimiter = '\t', quoting = 3)
# Cleaning the texts
import re
import nltk
nltk.download('stopwords')
from nltk.corpus import stopwords
from nltk.stem.porter import PorterStemmer
corpus = []
for i in range(0, 156060):
review = re.sub('[^a-zA-Z]', ' ', dataset['Phrase'][i])
review = review.lower()
review = review.split()
ps = PorterStemmer()
review = [ps.stem(word) for word in review ]
review = ' '.join(review)
corpus.append(review)
# Creating the Bag of Words model
from sklearn.feature_extraction.text import CountVectorizer
cv = CountVectorizer(max_features = 1500)
X = cv.fit_transform(corpus).toarray()
y = dataset.iloc[:, 3].values
# Splitting the dataset into the Training set and Test set
from sklearn.cross_validation import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.20, random_state = 0)
# +
# Natural Language Processing
# Importing the libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# Importing the dataset
dataset = pd.read_csv('train.tsv', delimiter = '\t', quoting = 3)
# Cleaning the texts
import re
import nltk
nltk.download('stopwords')
from nltk.corpus import stopwords
from nltk.stem.porter import PorterStemmer
corpus = []
for i in range(0, 156060):
review = re.sub('[^a-zA-Z]', ' ', dataset['Phrase'][i])
review = review.lower()
review = review.split()
ps = PorterStemmer()
review = [ps.stem(word) for word in review ]
review = ' '.join(review)
corpus.append(review)
# Creating the Bag of Words model
from sklearn.feature_extraction.text import CountVectorizer
cv = CountVectorizer(max_features = 1500)
X = cv.fit_transform(corpus).toarray()
y = dataset.iloc[:, 3].values
# Splitting the dataset into the Training set and Test set
from sklearn.cross_validation import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.20, random_state = 0)
| dt_load.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Information: Under development
# # Setup:
#
# We recommend you follow this course in a jupyter notebook which includes the course material and it's own terminal. If for whatever reason you don't see below for setup on your own machine.
# ## Jupyter Notebook
#
# Jupyter notebooks contain bash 'magic' which executes terminal commands within the jupyter interface. We have modified one of these for the purpose of this training, so where you see %%bash2 this is what you should run in a terminal.
# %%bash2
# This is a comment
# echo "This will be output"
# Slide Type
#
# You can also use the %%bash2 magic to keep your solutions to exercises in the notebook as a record. They will still be executed at the command line, so be careful! We can extract the files for this training by unzipping the data-shell.zip file in the repository. This can be done by executing the cell below:
#
# %%bash2 --dir ~/library/data
unzip data-shell.zip
A
# Slide Type
#
# We can check this has suceeded by changing into the newly created data-shell directory, using the bash2 magic and listing the contents (see the first lesson for more information).
# %%bash2 --dir ~/library/data/data-shell
# ls
# You are now ready to begin lesson 1.
| nbplain/99_bash_magic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Модель движения квадрокоптера
# Движение квадрокоптера определяется силами гравитации, аэродинамического сопротивления и тягой пропеллеров
# \begin{equation} \label{eq:common_traslational_motion}
# m \ddot{{r}} = {F}_g + {F}_{aero} + {F}_{thr}. \quad (IF)
# \end{equation}
#
# Сила тяжести определяется общей массой аппарата $m$ и вектором ускорения свободного падения $g$
# \begin{equation} \label{eq:gravity_force}
# {F}_g = mg.
# \end{equation}
#
# Сила аэродинамического сопротивления зависит от площади сечения корпуса $S_{\perp}$, плотности воздуха $\rho_{air}$ и аэродинамической константы $C$
#
# \begin{equation} \label{eq:aerodynamic_force}
# {F}_{aero} = - \frac{1}{2} \rho_{air} C S_{\perp} |\dot{{r}}| \dot{{r}}.
# \end{equation}
#
# Сила тяги пропеллеров зависит от их оборотов $\tilde\omega$ и аэродинамического коэффициента $k$,
# а ее направление зависит от ориентации корпуса $q$
# \begin{equation} \label{eq:thrust_force}
# {F}_{thr} = q \circ \Big(\sum_{i=1}^{4}{ f_i } \Big) \circ \tilde q =
# q \circ \Big( \sum_{i=1}^{4}{ { k \tilde\omega^2_i e_z} \Big) \circ \tilde q
# .}
# \end{equation}
#
#
# Вращательное движение зависит от моментов сил со стороны роторов с пропеллерами
#
# \begin{equation} \label{eq:common_rotational_motion}
# \sum_{i=1}^{4}{{r_i} \times {f}_{i}} - \sum_{i=1}^{4}{{\tau}_{i}}
# = {J}\dot{{\Omega}} + {\Omega} \times {J}{{\Omega}}, \quad (BF)
# \end{equation}
#
# \begin{equation} \label{eq:poison_eq}
# \dot{q} = \frac{1}{2} q \circ \Omega, \\
# \end{equation}
#
# \begin{equation} \label{eq:m_rotors_dyn}
# {\tau}_i + {\varsigma}_{i} =
# {j_i} \dot{\omega}_i + \omega_i \times j_i \omega_i, \quad (RF)
# \end{equation}
#
# \begin{equation} \label{eq:rotor_ext_torque}
# {\varsigma}_{i} = -b \tilde{\omega}^2_i {z_i}.
# \end{equation}
#
# Или, в упрощенном виде
# \begin{equation} \label{eq:simple_rotational_motion}
# T \tilde \omega
# = {J}\dot{{\Omega}} + {\Omega} \times {J}{{\Omega}}, \quad (BF) \\
# \quad \\
# \tilde \omega =
# \begin{bmatrix}
# \tilde \omega^{2}_{1}\\
# \tilde \omega^{2}_{2}\\
# \tilde \omega^{2}_{3}\\
# \tilde \omega^{2}_{4}
# \end{bmatrix}, \\
# \quad \\
# T =
# \begin{bmatrix}
# 0 & kL & 0 & -kL\\
# -kL & 0 & kL & 0\\
# b & -b & b & -b
# \end{bmatrix}.
# \end{equation}
#
# +
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from scipy.integrate import solve_ivp
# Вспомогательные функции
def normalize(obj):
return obj / np.linalg.norm(obj)
def cross_product(a, b):
def check_dimensions(vec, string):
if vec.ndim != 1:
raise Exception("The {} input is not a vector".format(string))
if len(vec) != 3:
raise Exception("Wrong number of coordinates in the {0} vector: {1}, should be 3".format(string, len(vec)))
check_dimensions(a, 'first')
check_dimensions(b, 'second')
return np.array([a[1]*b[2]-a[2]*b[1], a[2]*b[0]-a[0]*b[2], a[0]*b[1] - a[1]*b[0]])
def quat_product(q1, q2):
def check_dimensions(q, string):
if q.ndim != 1:
raise Exception("The {} input is not a quaternion".format(string))
if len(q) != 4:
raise Exception("Wrong number of coordinates in the {0} quaternion: {1}, should be 4".format(string, len(q)))
check_dimensions(q1, 'first')
check_dimensions(q2, 'second')
q = np.zeros(4)
q[0] = q1[0] * q2[0] - q1[1:].dot(q2[1:])
q[1:] = q1[0] * q2[1:] + q2[0] * q1[1:] + cross_product(q1[1:], q2[1:])
return q
def rotate_vec_with_quat(q, vec):
def check_dimensions(obj, is_quat):
if obj.ndim != 1:
raise Exception("Not a {}".format('quaternion' * is_quat + 'vector' * (1 - is_quat)))
if len(obj) != (3 + 1 * is_quat):
raise Exception("Wrong number of coordinates in the {0}: {1}, should be {2}"
.format('quaternion' * is_quat + 'vector' * (1 - is_quat), len(obj), 3 + 1 * is_quat))
check_dimensions(q, True)
check_dimensions(vec, False)
q = quat_conjugate(q)
qxvec = cross_product(q[1:], vec)
return q[1:].dot(vec) * q[1:] + q[0]**2. * vec + 2. * q[0] * qxvec + cross_product(q[1:], qxvec)
def quat2rpy(q0, q1, q2, q3):
roll = np.arctan2(2. * (q0 * q1 + q2 * q3), 1. - 2. * (q1**2 + q2**2))
pitch = np.arcsin(2. * (q0 * q2 - q1 * q3))
yaw = np.arctan2(2. * (q0 * q3 + q1 * q2), 1. - 2. * (q2**2 + q3**2))
return [roll, pitch, yaw]
def quat2rpy_deg(q0, q1, q2, q3):
roll = np.arctan2(2. * (q0 * q1 + q2 * q3), 1. - 2. * (q1**2 + q2**2))*180/np.pi
pitch = np.arcsin(2. * (q0 * q2 - q1 * q3))*180/np.pi
yaw = np.arctan2(2. * (q0 * q3 + q1 * q2), 1. - 2. * (q2**2 + q3**2))*180/np.pi
return [roll, pitch, yaw]
def quat_conjugate(q):
q_new = np.copy(q)
q_new[1:] *= -1.
return q_new
# -
# # Управление квадрокоптером
# [<NAME>., <NAME>. Minimum snap trajectory generation, control for
# quadrotors (https://ieeexplore.ieee.org/document/5980409)]
#
# Построим вектор управления таким образом,
# что первая его компонента будет отвечать за абсолютную величину тяги пропеллеров,
# а остальные три -- за три компоненты момента силы, действующие на корпус квадрокоптера
# \begin{equation} \label{eq:mellinger_control_vector}
# \begin{aligned}
# u = [|f| \quad \tau]^T =
# \begin{bmatrix}
# k & k & k & k\\
# 0 & kL & 0 & -kL\\
# -kL & 0 & kL & 0\\
# b & -b & b & -b
# \end{bmatrix}
# \begin{bmatrix}
# \tilde \omega^{2}_{1}\\
# \tilde \omega^{2}_{2}\\
# \tilde \omega^{2}_{3}\\
# \tilde \omega^{2}_{4}
# \end{bmatrix}.
# \end{aligned} \quad (BF)
# \end{equation}
#
# Для того, чтобы управлять положением квадрокоптера, построим ПД регулятор
# \begin{equation} \label{eq:pos_reg_1}
# \ddot{r}_{des} = k_r e_r + k_v e_v,
# \end{equation}
# где
# \begin{equation} \label{eq:mellinger_pos_err}
# {e}_r = {r}_{des} - {r},
# \end{equation}
# \begin{equation} \label{eq:mellinger_vel_err}
# {e}_v = {v}_{des} - {v}.
# \end{equation}
# +
# целевые траектории
def des_traj_fcn_0(t):
return np.array([0, 0])
def des_traj_fcn_1(t):
k = 0.05
return np.array([k * t**2, 2 * k * t])
def des_traj_fcn_2(t):
k = 3
p = 15
return np.array([k * np.sin(t * 2 * np.pi / p), k * 2 * np.pi / p * np.cos(t * 2 * np.pi/p)])
# выбор целевой траектории
def des_traj_fcn(t):
return des_traj_fcn_2(t)
# регулятор
def pd(t, x, kr, kv):
x_des = des_traj_fcn(t)
er = x_des[0] - x[0]
ev = x_des[1] - x[1]
x_dot = np.array([0.,0.])
x_dot[0] = x[1]
x_dot[1] = kr * er + kv * ev
return x_dot
# t
t0 = 0.
tf = 15.
t_eval = np.arange(t0, tf, 0.01)
# des
x_des = np.array([0, 0])
i = 0;
for t in t_eval:
if i == 0:
x_des = des_traj_fcn(t)
else:
x_des = np.vstack((x_des, des_traj_fcn(t)))
i += 1
# Выбор коэффициентов PD регулятора
kr = 1.5;
kv = 1.5;
# initial
x_0 = np.array([10., 6.])
# solve
sol = solve_ivp(lambda t, x: pd(t, x, kr, kv), (t0,tf), x_0, t_eval = t_eval)
x = sol.y.T
t = sol.t[1:]
# plot
fig1 = plt.figure(figsize=(8,8))
ax1 = fig1.add_subplot(1,1,1)
ax1.set_title("PD")
ax1.plot(t, x[1:,0], label = 'r', color = 'red')
ax1.plot(t, x[1:,1], label = 'v', color = 'green')
ax1.plot(t, x_des[1:,0], 'r--', label = 'r_des')
ax1.plot(t, x_des[1:,1], 'g--', label = 'v_des')
ax1.set_ylabel(r'val')
ax1.set_xlabel(r't, [s]')
ax1.grid(True)
ax1.legend()
# -
# Тогда,
# \begin{equation} \label{eq:mellinger_pos_reg}
# {F}_{des} = m k_r {e}_r + m k_v {e}_v - F_g - F_{aero}.
# \end{equation}
#
# Чтобы обеспечить соответствие управляющего вектора выходу регулятора,
# нужно изменить ориентацию корпуса квадрокоптера так,
# чтобы направление его тяги совпадало с требуемой
#
# \begin{equation} \label{eq:mellinger_Rdes}
# q_{des} \circ e_z \circ \tilde q_{des} = \frac{{F}_{des}}{||{F}_{des}||}.
# \end{equation}
#
# Решений для $q_{des}$ будет множество,
# т.к. наклонить таким образом корпус можно, изменяя тангаж и крен,
# при этом угол рысканья может быть произвольным.
# Вычислив $q_{des}$, например, для фиксированного угла рысканья,
# можно воспользоваться рассмотренным ранее регулятором
# для отслеживания необходимой ориентации
#
# \begin{equation} \label{eq:qf_reg}
# \tau_{des} = -{\Omega} \times {J}{{\Omega}} + k_{\Omega} e_{\Omega} + k_{q} e_{q}.
# \end{equation}
#
# Тогда, $u = [{F}_{des} \cdot {z}_B \quad \tau_{des}]^T$.
# +
q_des_list = np.array([0, 0, 0, 0])
r_des_list = np.array([0, 0, 0])
v_des_list = np.array([0, 0, 0])
tilde_w_sat_list = np.array([0, 0, 0, 0])
# Управление квадрокоптером
def des_traj_fcn_0(t):
return np.zeros((6,))
def des_traj_fcn_1(t):
k = 10
p = 60
rx = k * np.sin(t * 2 * np.pi / p)
ry = k * np.sin(2 * t * 2 * np.pi / p)
rz = 0.3*rx
vx = 2 * np.pi / p * k * np.cos(t * 2 * np.pi / p)
vy = 2 * 2 * np.pi / p * k * np.cos(2 * t * 2 * np.pi / p)
vz = 0.3 * vx
return np.array([rx, ry, rz, vx, vy, vz])
def des_traj_fcn(t):
return des_traj_fcn_1(t)
def uav_euler_int(x0, T, dt, UAV):
xs = np.array([x0])
ts = np.array([0.])
while ts[-1] < T:
x_dot = model(ts[-1], xs[-1,:], UAV)
x_next = xs[-1,:] + x_dot * dt
t_next = ts[-1] + dt
xs = np.vstack((xs, x_next))
ts = np.append(ts, t_next)
return ts, xs
def model(t, x, UAV):
# x [r v q omega]
r = x[0:3]
v = x[3:6]
q = x[6:10]
w = x[10:13]
# reg
x_des = des_traj_fcn(t)
r_des = x_des[0:3]
v_des = x_des[3:6]
f_des = pose_reg(t, x, UAV, r_des, v_des)
q_des = quat_des(f_des, yaw=1.)
tau_des = qf_reg(t, x, q_des, UAV)
# ctrl vec
u = np.array([np.linalg.norm(f_des), tau_des[0], tau_des[1], tau_des[2]])
tilde_w2 = UAV.A_inv @ u
tilde_w = np.sqrt(tilde_w2)
tilde_w_sat = np.clip(tilde_w, 0., UAV.w_lim)
tilde_w2_sat = np.power(tilde_w_sat, 2)
u_sat = UAV.A @ tilde_w2_sat
# f tau
ft_abs = u_sat[0]
ft = rotate_vec_with_quat(quat_conjugate(q), np.array([0.,0.,ft_abs]))
Fg = - np.array([0., 0., 10]) * UAV.m
Fa = - 0.5 * UAV.C * UAV.S * UAV.rho * v * np.linalg.norm(v)
f = ft + Fg + Fa
taut = u_sat[1:4]
tau = taut - cross_product(w, np.matmul(UAV.J, w))
# x_dot
a = f/UAV.m
nu = np.matmul(UAV.J_inv, tau)
x_dot = np.zeros((13,))
x_dot[0:3] = v
x_dot[3:6] = a
x_dot[6] = -0.5 * q[1:4].dot(w)
x_dot[7:10] = 0.5 * (q[0] * w + cross_product(q[1:4], w))
x_dot[10:13] = nu
# hist
global q_des_list
global r_des_list
global v_des_list
global tilde_w_sat_list
q_des_list = np.vstack((q_des_list, q_des))
r_des_list = np.vstack((r_des_list, r_des))
v_des_list = np.vstack((v_des_list, v_des))
tilde_w_sat_list = np.vstack((tilde_w_sat_list, tilde_w_sat))
return x_dot
def quat_des(v, yaw):
q_yaw = np.array([np.cos(yaw / 2.0), 0.0, 0.0, np.sin(yaw / 2.0)])
if (np.linalg.norm(v) < 1e-3):
return q_yaw
ez = np.array([0., 0., 1.])
pin = np.cross(ez, v)
if (np.linalg.norm(pin) < 1e-3):
return q_yaw
pin = pin / np.linalg.norm(pin)
vn = v / np.linalg.norm(v)
cosA = np.dot(vn, ez)
A = np.arccos(cosA)
sinHalfA = np.sin(A / 2.0)
q_bow = np.array([np.cos(A / 2.0), sinHalfA * pin[0], sinHalfA * pin[1], sinHalfA * pin[2]])
return quat_product(q_bow, q_yaw)
def pose_reg(t, x, UAV, r_des, v_des):
# x [r v q omega]
r = x[0:3]
v = x[3:6]
q = x[6:10]
w = x[10:13]
er = r_des - r
ev = v_des - v
Fg = - np.array([0., 0., 10]) * UAV.m
Fa = - 0.5 * UAV.C * UAV.S * UAV.rho * v * np.linalg.norm(v)
ez_I = rotate_vec_with_quat(quat_conjugate(q), np.array([0., 0., 1]))
f = UAV.m * UAV.kr * er + UAV.m * UAV.kv * ev - Fg - Fa
return f
def qf_reg(t, x, q_des, UAV):
# x [r v q omega]
r = x[0:3]
v = x[3:6]
q = x[6:10]
w = x[10:13]
x_des = np.zeros((7))
x_des[0:4] = q_des
dw = -w
dq = -quat_product(quat_conjugate(x_des[0:4]), q)
tau = cross_product(w, np.matmul(UAV.J, w)) + UAV.ko * np.matmul(UAV.J, dw) + UAV.kq * np.matmul(UAV.J, dq[1:])
return tau
# +
q_des_list = np.array([0, 0, 0, 0])
r_des_list = np.array([0, 0, 0])
v_des_list = np.array([0, 0, 0])
tilde_w_sat_list = np.array([0, 0, 0, 0])
# Моделирование
# params
class UAV:
m = 1.0
J = np.diag(np.array([1, 1, 1])) * 1e-2
J_inv = np.linalg.inv(J)
L = 0.25
k = 1e-5
b = 1e-6
w_lim = 1e3
S = 0.12
C = 1.0
rho = 1.0
kr = 0.1
kv = 1.
kq = 25.
ko = 30.
A = np.array([[k, k, k, k],
[0, k*L, 0, -k*L],
[-k*L, 0, k*L, 0],
[b, -b, b, -b]])
A_inv = np.linalg.inv(A)
UAVobject = UAV();
# t
t0 = 0.
tf = 90.
dt = 0.01
t_eval = np.arange(t0, tf, dt)
# initial
x_0 = np.array([-45., 0., -15., 0., 0., 0., 1., 0., 0., 0., 0., 0., 0.])
ts, xs = uav_euler_int(x_0, tf, dt, UAVobject)
# plot
fig1 = plt.figure(figsize=(16,8))
ax1 = fig1.add_subplot(1,2,1)
ax2 = fig1.add_subplot(1,2,2)
# r
ax1.set_title("r")
n = 0
ax1.plot(ts, xs[:,n], 'r', label = 'x')
ax1.plot(ts, xs[:,n+1], 'g', label = 'y')
ax1.plot(ts, xs[:,n+2], 'b', label = 'z')
ax1.plot(ts[1:], r_des_list[1:,0], 'r--', label = 'x_des')
ax1.plot(ts[1:], r_des_list[1:,1], 'g--', label = 'y_des')
ax1.plot(ts[1:], r_des_list[1:,2], 'b--', label = 'z_des')
ax1.set_ylabel(r'r')
ax1.set_xlabel(r't, [s]')
ax1.grid(True)
ax1.legend()
# q
ax2.set_title("qv")
n = 7
ax2.plot(ts, xs[:,n], 'r', label = 'qx')
ax2.plot(ts, xs[:,n+1], 'g', label = 'qy')
ax2.plot(ts, xs[:,n+2], 'b', label = 'qz')
ax2.plot(ts[1:], q_des_list[1:,1], 'r--', label = 'qx_des')
ax2.plot(ts[1:], q_des_list[1:,2], 'g--', label = 'qy_des')
ax2.plot(ts[1:], q_des_list[1:,3], 'b--', label = 'qz_des')
ax2.set_ylabel(r'qv')
ax2.set_xlabel(r't, [s]')
ax2.grid(True)
ax2.legend()
fig3 = plt.figure(figsize=(16,8))
ax3 = fig3.add_subplot(1,2,1)
ax4 = fig3.add_subplot(1,2,2)
# v
ax3.set_title("v")
n = 3
ax3.plot(ts, xs[:,n], 'r', label = 'vx')
ax3.plot(ts, xs[:,n+1], 'g', label = 'vy')
ax3.plot(ts, xs[:,n+2], 'b', label = 'vz')
ax3.plot(ts[1:], v_des_list[1:,0], 'r--', label = 'x_des')
ax3.plot(ts[1:], v_des_list[1:,1], 'g--', label = 'y_des')
ax3.plot(ts[1:], v_des_list[1:,2], 'b--', label = 'z_des')
ax3.set_ylabel(r'v')
ax3.set_xlabel(r't, [s]')
ax3.grid(True)
ax3.legend()
# w
ax4.set_title(r'$\Omega$')
n = 10
ax4.plot(ts, xs[:,n], 'r', label = r'$\Omega_x$')
ax4.plot(ts, xs[:,n+1], 'g', label = r'$\Omega_y$')
ax4.plot(ts, xs[:,n+2], 'b', label = r'$\Omega_z$')
ax4.set_ylabel(r'w')
ax4.set_xlabel(r't, [s]')
ax4.grid(True)
ax4.legend()
# rotors
fig4 = plt.figure(figsize=(16,8))
ax5 = fig4.add_subplot(1,2,1)
ax6 = fig4.add_subplot(1,2,2)
ax5.set_title(r'$\tilde{\omega}$')
ax6.set_title(r'$\tilde{\omega} - \tilde{\omega}_0$')
ax5.plot(ts[1:], tilde_w_sat_list[1:,0], 'k', label = '')
ax5.plot(ts[1:], tilde_w_sat_list[1:,1], 'k', label = '')
ax5.plot(ts[1:], tilde_w_sat_list[1:,2], 'k', label = '')
ax5.plot(ts[1:], tilde_w_sat_list[1:,3], 'k', label = '')
ax6.plot(ts[1:], tilde_w_sat_list[1:,0]-tilde_w_sat_list[1:,0], 'k', label = '')
ax6.plot(ts[1:], tilde_w_sat_list[1:,1]-tilde_w_sat_list[1:,0], 'r', label = '')
ax6.plot(ts[1:], tilde_w_sat_list[1:,2]-tilde_w_sat_list[1:,0], 'g', label = '')
ax6.plot(ts[1:], tilde_w_sat_list[1:,3]-tilde_w_sat_list[1:,0], 'b', label = '')
ax5.set_ylabel(r'$\tilde{\omega}$')
ax5.set_xlabel(r't, [s]')
ax6.set_ylabel(r'$\tilde{\omega} - \tilde{\omega}_0$')
ax6.set_xlabel(r't, [s]')
ax5.grid(True)
ax6.grid(True)
#ax5.legend()
#ax6.legend()
# -
# 3d
fig3d = plt.figure(figsize=(8,8))
ax3d = fig3d.add_subplot(111, projection='3d')
ax3d.plot(xs[:,0], xs[:,1], xs[:,2], 'b', label = 'traj')
ax3d.plot(r_des_list[1:,0], r_des_list[1:,1], r_des_list[1:,2], 'k', label = 'des_traj')
ax3d.set_xlabel(r'x')
ax3d.set_ylabel(r'y')
ax3d.set_zlabel(r'z')
ax3d.grid(True)
lim = 10.
ax3d.set_xlim(-lim, lim)
ax3d.set_ylim(-lim, lim)
ax3d.set_zlim(-lim, lim)
#ax5.legend()
# В качестве упражнения предлагается усовершенствовать контур управления квадрокоптером.
#
# С помощью выражения
# \begin{equation} \label{eq:mellinger_Rdes}
# q_{des} \circ e_z \circ \tilde q_{des} = \frac{{F}_{des}}{||{F}_{des}||}.
# \end{equation}
# производится расчет целевой ориентации БЛА, при этом на вход quarternion feedback регулятора можно подавать также целевую угловую скорость.
#
# Как найти целевую угловую скорость и как ее использование в регуляторе повлияет на производительность алгоритма управления?
| Lecture Jupyter Notebooks/Lecture 7 Simplified Quadcopter Control Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lecture 15 - Advanced Curve Fitting: Gaussian Process Regression
import numpy as np
import scipy.stats as st
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
sns.set_context('talk')
sns.set_style('white')
# ## Objectives
#
# + To do regression using a GP.
# + To introduce some diagnostics for how good a probabilistic regression is.
# + To find the hyperparameters of the GP by maximizing the (marginal) likelihood.
# ## Readings
#
# + Please read [this](http://www.kyb.mpg.de/fileadmin/user_upload/files/publications/pdfs/pdf2903.pdf) OR watch [this video lecture](http://videolectures.net/mlss03_rasmussen_gp/?q=MLSS).
#
# + [Chapter 3 from C.E. Rasmussen's textbook on Gaussian processes](http://www.gaussianprocess.org/gpml/chapters/RW2.pdf)
#
# + [Section 5.4 in GP for ML textbook](http://www.gaussianprocess.org/gpml/chapters/RW5.pdf).
# ## Motivation: A fully Bayesian paradigm for curve fitting
#
# Remember why we are doing this:
#
# + Let's say that you have to learn some function $f(\cdot)$ from some space $\mathcal{X}$ to $\mathbb{R}$ (this could either be a supervised learning problem (regression or classification) or even an unsupervised learning problem.
#
# + You sit down and you think about $f(\cdot)$. What do you know about it? How large do you expect it be? How small do you expect it be? Is it continuous? Is it differentiable? Is it periodic? How fast does it change as you change its inputs?
#
# + You create a probability measure on the space of functions in which $f(\cdot)$ lives which is compatible with everything you know about it. Abusing mathematical notation a lot, let's write this probability measure as $p(f(\cdot))$. Now you can sample from it. Any sample you take is compatible with your prior beliefs. You cannot tell which one is better than any other. Any of them could be the true $f(\cdot)$.
#
# + Then, you get a little bit of data, say $\mathcal{D}$. You model the likelihood of the data, $p(\mathcal{D}|f(\cdot))$, i.e., you model how the data may have been generated if you knew $f(\cdot)$.
#
# + Finally, you use Bayes' rule to come up with your posterior probability measure over the space of functions:
# $$
# p(f(\cdot)|\mathcal{D}) \propto p(\mathcal{D}|f(\cdot)) p(f(\cdot)),
# $$
# which is simultaneously compatible with your prior beliefs and the data.
#
# In the last lecture, we formalized mathematically the prior $p(f(\cdot))$.
# Today, we will mathematically formalize the posterior $p(f(\cdot)|\mathcal{D})$.
# ## Reminder: The prior $p(f(\cdot))$
#
# In the previous lecture, we defined $p(f(\cdot))$ through the concept of a Gaussian process (GP), a generalization of a multivariate Gaussian distribution to
# *infinite* dimensions.
# We argued that it defines a probability measure on a function space.
# We wrote:
# $$
# f(\cdot) \sim \text{GP}\left(m(\cdot), k(\cdot, \cdot) \right),
# $$
# where
# $m:\mathbb{R}^d \rightarrow \mathbb{R}$ is the *mean function* and
# $k:\mathbb{R}^d \times \mathbb{R}^d \rightarrow \mathbb{R}$ is the *covariance function*.
#
# We also discuss in detail the meaning of these equations.
# Namely that for any points $\mathbf{x}_{1:n}=(\mathbf{x}_1,\dots,\mathbf{x}_n)$, the joint probability density of the function values:
# $$
# \mathbf{f}_{1:n} =
# \left(
# f(\mathbf{x}_1),
# \dots,
# f(\mathbf{x}_n)
# \right).
# $$
# is the multivariate Gaussian:
# $$
# \mathbf{f}_{1:n} | \mathbf{x}_{1:n} \sim \mathcal{N}\left(\mathbf{m}(\mathbf{x}_{1:n}), \mathbf{K}(\mathbf{x}_{1:n}, \mathbf{x}_{1:n}) \right),
# $$
# with mean vector:
# $$
# \mathbf{m}(\mathbf{x}_{1:n}) =
# \left(
# m(\mathbf{x}_1),
# \dots,
# m(\mathbf{x}_n)
# \right),
# $$
# and covariance matrix:
# $$
# \mathbf{K}(\mathbf{x}_{1:n}, \mathbf{x}_{1:n}) = \left(
# \begin{array}{ccc}
# k(\mathbf{x}_1,\mathbf{x}_1) & \dots & k(\mathbf{x}_1, \mathbf{x}_n)\\
# \vdots & \ddots & \vdots\\
# k(\mathbf{x}_n, \mathbf{x}_1) & \dots & k(\mathbf{x}_n, \mathbf{x}_n)
# \end{array}
# \right).
# $$
# Please note that all the above expressions are conditional on the hyperparameters of the covariance function, e.g., lengthscales and signal variance for the squared exponential.
# However, for now, we do not explicitly show this dependence.
# ## Gaussian process regression
#
# Assume that the data we observe is:
# $$
# \mathcal{D} = (\mathbf{x}_{1:n}, y_{1:n}),
# $$
# where $y_i$ is not exactly $f(\mathbf{x}_i)$, but it may have some noise.
# For the sake of simplicity, let's assume that the noise is Gaussian with variance $\sigma$ (you can relax this, but things will no longer be analytically available).
# Assume that $\sigma$ is known (for now).
# So, we have:
# $$
# y_i|f(\mathbf{x}_i) \sim \mathcal{N}(f(\mathbf{x}_i), \sigma^2),
# $$
# for a single observation.
# For all the observations together, we can write:
# $$
# y_{1:n}| \mathbf{f}_{1:n} \sim \mathcal{N}(\mathbf{f}_{1:n}, \sigma^2\mathbf{I}_n),
# $$
# where $\mathbf{I}_n$ is the $n\times n$ unit matrix.
# Let's draw a graphical model representation what we have here.
# Remember that $\sigma$ and the hyperparameters of the covariance function, let's call them $\theta$, are (for the time being) fixed.
from graphviz import Digraph
g = Digraph('gp')
g.node('sigma', label='<σ>', style='filled')
g.node('theta', label='<θ>', style='filled')
g.node('x1n', label='<<b>x</b><sub>1:n</sub>>', style='filled')
g.node('f', label='<<b>f</b><sub>1:n</sub>>')
g.node('y1n', label='<<b>y</b><sub>1:n</sub>>', style='filled')
g.edge('theta', 'f')
g.edge('x1n', 'f')
g.edge('f', 'y1n')
g.edge('sigma', 'y1n')
g.render('gp_reg1', format='png')
g
# So, far so good, but how do we make predictions?
# How do we get $p(f(\cdot)|\mathcal{D})$.
# There are some nuances here.
# We are looking for a posterior measure over a function space.
# This is a strange beast.
# The only way we can describe it is through the joint probability density of the function values at any arbitrary collection of **test** points.
# So, let's take $n^*$ such test points:
# $$
# \mathbf{x}^*_{1:n^*} = \left(\mathbf{x}^*_1,\dots,\mathbf{x}^*_{n^*}\right).
# $$
# Imagine that these cover the input space as densely as we wish.
# Consider the vector of function values at these test points:
# $$
# \mathbf{f}^*_{1:n^*} = \left(f(\mathbf{x}_1^*),\dots,f(\mathbf{x}^*_{n^*})\right).
# $$
# We will derive the posterior over these points, i.e., we will derive $p(\mathbf{f}^*|\mathcal{D})$.
# And we will be happy with that.
#
# From the definition of the GP, we can now write the joint probability density of $\mathbf{f}$ and $\mathbf{f}^*$.
# It is just a multivariate Gaussian.
# We have:
# $$
# p(\mathbf{f}_{1:n}, \mathbf{f}^*_{1:n^*} | \mathbf{x}_{1:n}, \mathbf{x}^*_{1:n^*}) = \mathcal{N}\left(
# \left(
# \begin{array}{c}
# \mathbf{f}_{1:n}\\
# \mathbf{f}^*_{1:n^*}
# \end{array}
# \right)\middle |
# \left(
# \begin{array}{c}
# \mathbf{m}(\mathbf{x}_{1:n})\\
# \mathbf{m}(\mathbf{x}^*_{1:n^*})
# \end{array}
# \right),
# \left(
# \begin{array}{cc}
# \mathbf{K}(\mathbf{x}_{1:n}, \mathbf{x}_{1:n}) & \mathbf{K}(\mathbf{x}_{1:n}, \mathbf{x}^*_{1:n^*})\\
# \mathbf{K}(\mathbf{x}^*_{1:n^*}, \mathbf{x}_{1:n}) & \mathbf{K}(\mathbf{x}^*_{1:n^*}, \mathbf{x}_{1:n^*})
# \end{array}
# \right)
# \right),
# $$
# where the block matrix is just the covariance matrix of all inputs $\mathbf{x}_{1:n}$ (observed) and $\mathbf{x}^*_{1:n^*}$ (test).
# Let's visualize the situation again.
g2 = Digraph('gp2')
g2.node('sigma', label='<σ>', style='filled')
g2.node('theta', label='<θ>', style='filled')
g2.node('x1n', label='<<b>x</b><sub>1:n</sub>>', style='filled')
g2.node('f', label='<<b>f</b><sub>1:n</sub>, <b>f</b>*<sub>1:n*</sub>>')
g2.node('y1n', label='<<b>y</b><sub>1:n</sub>>', style='filled')
g2.node('xs1ns', label='<<b>x</b>*<sub>1:n*</sub>>', style='filled')
g2.edge('theta', 'f')
g2.edge('x1n', 'f')
g2.edge('f', 'y1n')
g2.edge('xs1ns', 'f')
g2.edge('sigma', 'y1n')
g2.render('gp_reg2', format='png')
g2
# Ok, now we have only finite dimensional probability densities.
# This is great. We know what to do next.
# We will use the basic probability rules:
# $$
# \begin{array}{ccc}
# p(\mathbf{f}^*_{1:n^*} | \mathbf{x}^*_{1:n^*}, \mathcal{D}) &=& p(\mathbf{f}^*_{1:n^*} | \mathbf{x}^*_{1:^*}, \mathbf{x}_{1:n}, \mathbf{y}_{1:n})\\
# &=& \int p(\mathbf{f}_{1:n}, \mathbf{f}^*_{1:n^*} | \mathbf{x}^*_{1:^*}, \mathbf{x}_{1:n}, \mathbf{y}_{1:n})d\mathbf{f}_{1:n}\;\text{(sum rule)}\\
# &\propto& \int p(\mathbf{y}_{1:n}| \mathbf{f}_{1:n}) p(\mathbf{f}_{1:n}, \mathbf{f}^*_{1:n^*}|\mathbf{x}^*_{1:^*}, \mathbf{x}_{1:n}) d\mathbf{f}_{1:n}\;\text{(Bayes' rule)}.
# \end{array}
# $$
# This is the integral of a Gaussian times a Gaussian.
# We are not going to go into the details, but you can actually do it analytically.
# The result is... a Gaussian:
# $$
# p(\mathbf{f}^*_{1:n^*}| \mathbf{x}^*_{1:n^*}, \mathcal{D}) = \mathcal{N}\left(\mathbf{f}^*_{1:n^*}\middle| \mathbf{m}_n(\mathbf{x}^*_{1:n^*}), \mathbf{K}_n(\mathbf{x}^*_{1:n^*},\mathbf{x}^*_{1:n^*})\right),
# $$
# where *posterior mean function* is:
# $$
# m_n(x) = m(x) + \mathbf{k}(x,\mathbf{x}_{1:n})\left(\mathbf{K}(\mathbf{x}_{1:n},\mathbf{x}_{1:n})+\sigma^2I_n\right)^{-1}\left(\mathbf{y}_{1:n} - \mathbf{m}(\mathbf{x}_{1:n})\right),
# $$
# and the *posterior covariance function* is:
# $$
# k_n(x, x') = k(x,x') - \mathbf{k}(x,\mathbf{x}_{1:n})\left(\mathbf{K}(\mathbf{x}_{1:n},\mathbf{x}_{1:n})+\sigma^2I_n\right)^{-1}\mathbf{k}^T(x,\mathbf{x}_{1:n}),
# $$
# with
# $$
# \mathbf{k}(x,\mathbf{x}_{1:n}) = \left(k(x,\mathbf{x}_1),\dots,k(x,\mathbf{x}_n)\right)
# $$
# being the cross-covariance vector.
#
# Now notice that the test points $\mathbf{x}^*_{1:n^*}$ are arbitrary and that the joint distribution of the function values at these points, $\mathbf{f}^*$, conditioned on the observations $\mathcal{D}$ is a multivariate Gaussian with a mean and covariance matrix specified by the posterior mean and covariance functions, respectively.
# This is the defintion of a Gaussian process!
# Therefore, we conclude that the posterior probability measure over the space of functions is also a Gaussian process:
# $$
# f(\cdot)|\mathcal{D} \sim \operatorname{GP}(m_n(\cdot), k_n(\cdot, \cdot)).
# $$
# ### The point predictive distribution
#
# What if you just want to make a prediction at a single point?
# How do you do that?
# Well, this is quite simple.
# Your "test points" $\mathbf{x}^*_{1:n^*}$ are now just a single point, say $\mathbf{x}^*$.
# Your prediction about the function value at this point is captured by:
# $$
# p\left(f(\mathbf{x}^*) | \mathcal{D}\right) = \mathcal{N}\left(f(\mathbf{x}^*)\middle|m_n(\mathbf{x}^*), \sigma_n^2(\mathbf{x}^*)\right),
# $$
# where the *predictive variance* is just:
# $$
# \sigma_n^2(\mathbf{x}^*) = k_n(\mathbf{x}^*,\mathbf{x}^*).
# $$
# This is what we will be using the most.
#
# Now, if you wanted to predict $y^*$ at $\mathbf{x}^*$, i.e., the measurement at $\mathbf{x}^*$, you have do this:
# $$
# p(y^*|\mathbf{x}^*, \mathcal{D}) = \int p(y^*|f(\mathbf{x}^*)) p(f(\mathbf{x}^*)|\mathcal{D}) df(\mathbf{x}^*) = \mathcal{N}\left(f(\mathbf{x}^*)\middle|m_n(\mathbf{x}^*), \sigma_n^2(\mathbf{x}^*)+\sigma^2\right).
# $$
# Notice that you need to add the noise variance when you are talking about the measurement.
# ## Example: Gaussian process regression in 1D with fixed hyper-parameters
#
# Let's generate some synthetic 1D data to work with:
# Fixing the seed so that we all see the same data
np.random.seed(1234)
n = 10
# The inputs are in [0, 1]
X = np.random.rand(n, 1) # Needs to be an n x 1 vector
# The outputs are given from a function plus some noise
# The standard deviation of the noise is:
sigma = 0.4
# The true function that we will try to identify
f_true = lambda x: -np.cos(np.pi * x) + np.sin(4. * np.pi * x)
# Some data to train on
Y = f_true(X) + sigma * np.random.randn(X.shape[0], 1)
# Let's visualize the data
fig, ax = plt.subplots(dpi=100)
ax.plot(X, Y, 'kx', markersize=10, markeredgewidth=2)
ax.set_xlabel('$x$')
ax.set_ylabel('$y$');
# Now, we will get started with the regression
# First, import GPy
import GPy
# Second, pick a kernel. Let's pick a squared exponential (RBF = Radial Basis Function)
k = GPy.kern.RBF(1) # The parameter here is the dimension of the input (here 1)
# Let's print the kernel object to see what it includes:
print(k)
# The ``variance`` of the kernel is one. This seems reasonable.
# Let's leave it like that.
# The ``lengthscale`` seems to big.
# Let's change it to something reasonable (based on our expectations):
k.lengthscale = 0.1
print(k)
# There is a possibility to choose a mean function, but for simplicity we are going to pick a zero mean function:
# $$
# m(x) = 0.
# $$
# Now we put together the GP regression model as follows:
gpm = GPy.models.GPRegression(X, Y, k) # It is input, output, kernel
# This model is automatically assuming that the likelihood is Gaussian (you can modify it if you wish).
# Where do can you find the $\sigma^2$ parameter specifying the likelihood noise? Here it is:
print(gpm)
# We will talk about the meaning of all that later. For now, let's just fix the noise variance to something reasonable (actually the correct value):
gpm.likelihood.variance = sigma ** 2
print(gpm)
# That's it. We have now specified the model completely.
# The posterior GP is completely defined.
# Where is the posterior mean $m_n(x)$ and variance $\sigma_n^2(x)$? You can get them like this:
# First the mean on some test points
x_star = np.linspace(0, 1, 100)[:, None] # This is needed to turn the array into a column vector
m_star, v_star = gpm.predict(x_star)
# Let's plot the mean first
fig, ax = plt.subplots(dpi=100)
ax.plot(X, Y, 'kx', markersize=10, markeredgewidth=2, label='data')
ax.set_xlabel('$x$')
ax.set_ylabel('$y$')
ax.plot(x_star, m_star, lw=2, label='$m_n(x)$')
plt.legend(loc='best');
# Extracting the variance is a bit more involved.
# Just a tiny bit though.
# This is because ``v_star`` returned by ``gpm.predict`` is not exactly $\sigma_n^2(x)$.
# It is actually $\sigma_n^2(x) + \sigma^2$ and not just $\sigma_n^2(x)$.
# Here, see it:
# Now the variance on the same test points
fig, ax = plt.subplots(dpi=100)
ax.plot(x_star, v_star, lw=2, label='$\sigma_n^2(x) + \sigma^2$')
ax.plot(x_star, gpm.likelihood.variance * np.ones(x_star.shape[0]), 'r--', lw=2, label='$\sigma^2$')
ax.set_xlabel('$x$')
ax.set_ylabel('$\sigma_n^2(x)$')
plt.legend(loc='best');
# Notice that the variance is small wherever we have an observation.
# It is not, however, exactly, $\sigma^2$.
# It will become exactly $\sigma^2$ in the limit of many observations.
# Having the posterior mean and variance, we can derive 95\% predictive intervals for $f(x^*)$ and $y^*$.
# For $f(x^*)$ these are:
# $$
# m_n(\mathbf{x}^*)) - 2\sigma_n(\mathbf{x}^*) \le f(\mathbf{x}^*) \le m_n(\mathbf{x}^*)) + 2\sigma_n(\mathbf{x}^*).
# $$
# Let's plot this:
fig, ax = plt.subplots(dpi=100)
ax.set_xlabel('$x$')
ax.set_ylabel('$y$')
f_lower = m_star - 2.0 * np.sqrt(v_star - gpm.likelihood.variance)
f_upper = m_star + 2.0 * np.sqrt(v_star - gpm.likelihood.variance)
ax.fill_between(x_star.flatten(), f_lower.flatten(), f_upper.flatten(), alpha=0.5)
ax.plot(x_star, m_star, lw=2, label='$m_n(x)$')
ax.plot(X, Y, 'kx', markersize=10, markeredgewidth=2, label='data')
plt.legend(loc='best');
# Now, on the same plot, let's superimpose our predictive error bar about $y^*$.
# This is:
# $$
# m_n(\mathbf{x}^*)) - 2\sqrt{\sigma_n^2(\mathbf{x}^*)+\sigma^2}\le f(\mathbf{x}^*) \le m_n(\mathbf{x}^*)) + 2\sqrt{\sigma_n(\mathbf{x}^*) + \sigma^2}.
# $$
# Let's use red color for this:
fig, ax = plt.subplots(dpi=100)
ax.set_xlabel('$x$')
ax.set_ylabel('$y$')
f_lower = m_star - 2.0 * np.sqrt(v_star - gpm.likelihood.variance)
f_upper = m_star + 2.0 * np.sqrt(v_star - gpm.likelihood.variance)
y_lower = m_star - 2.0 * np.sqrt(v_star)
y_upper = m_star + 2.0 * np.sqrt(v_star)
ax.fill_between(x_star.flatten(), y_lower.flatten(), y_upper.flatten(), color='red', alpha=0.25, label='$y^*$ 95% pred.')
ax.fill_between(x_star.flatten(), f_lower.flatten(), f_upper.flatten(), alpha=0.5, label='$f(\mathbf{x}^*)$ 95% pred.')
ax.plot(x_star, m_star, lw=2, label='$m_n(x)$')
ax.plot(X, Y, 'kx', markersize=10, markeredgewidth=2, label='data')
#plt.legend(loc='best');
# Let's also put the correct function there for comparison:
fig, ax = plt.subplots(dpi=100)
ax.set_xlabel('$x$')
ax.set_ylabel('$y$')
f_lower = m_star - 2.0 * np.sqrt(v_star - gpm.likelihood.variance)
f_upper = m_star + 2.0 * np.sqrt(v_star - gpm.likelihood.variance)
y_lower = m_star - 2.0 * np.sqrt(v_star)
y_upper = m_star + 2.0 * np.sqrt(v_star)
ax.fill_between(x_star.flatten(), y_lower.flatten(), y_upper.flatten(), color='red', alpha=0.25, label='$y^*$ 95% pred.')
ax.fill_between(x_star.flatten(), f_lower.flatten(), f_upper.flatten(), alpha=0.5, label='$f(\mathbf{x}^*)$ 95% pred.')
ax.plot(x_star, m_star, lw=2, label='$m_n(x)$')
ax.plot(x_star, f_true(x_star), 'm-.', label='True function')
ax.plot(X, Y, 'kx', markersize=10, markeredgewidth=2, label='data');
#plt.legend(loc='best');
# You see that the true function is almost entirely within the blue bounds.
# It is ok that it is a little bit off, becuase these are 95% prediction intervals.
# About 5% of the function can be off.
# That's good.
# However, we have much more information encoded in the posterior GP.
# It is actually a probability measure over the space of functions.
# How do we sample functions?
# Well, you can't sample functions...
# They are infinite objects.
# But you can sample the *function values* on a bunch of test points.
# As a matter of fact, the joint probability density of the function values at any collection of set points is a multivariate Gaussian.
# We did it manually in the last lecture.
# In this lecture, we are going to use the capabilities of ``GPy``.
# Here it is:
# Here is how you take the samples
f_post_samples = gpm.posterior_samples_f(x_star, 10) # Test points, how many samples you want
# Here is the size of f_post_samples
print(f_post_samples.shape)
# This is ``test points x number of outputs (1 here) x number of samples``.
# Let's plot them along with the data and the truth:
fig, ax = plt.subplots(dpi=100)
ax.plot(x_star, f_post_samples[:, 0, :], 'r', lw=0.5)
ax.plot(X, Y, 'kx', markersize=10, markeredgewidth=2, label='data');
ax.plot(x_star, f_true(x_star), 'm-.', label='True function')
ax.set_xlabel('$x$')
ax.set_ylabel('$y$');
# Ok, we see that the lengthscale we have assumed does not match the lengthscale of the true function perfectly.
# But that's how it is.
# In real problems, you won't know the true function anyway.
# The following interactive function regenerates the figures above allowing you to experiment with various choices of the hyperparameters.
# +
from ipywidgets import interact_manual
def analyze_and_plot_gp_ex1(kern_variance=1.0, kern_lengthscale=0.1, like_variance=0.4):
"""
Performs GP regression with given kernel variance, lengthcale and likelihood variance.
"""
#k = GPy.kern.RBF(1)
k = GPy.kern.Matern32(1)
gp_model = GPy.models.GPRegression(X, Y, k)
# Set the parameters
gp_model.kern.variance = kern_variance
gp_model.kern.lengthscale = kern_lengthscale
gp_model.likelihood.variance = like_variance
# Print model for sanity check
print(gp_model)
# Pick test points
x_star = np.linspace(0, 1, 100)[:, None]
# Get posterior mean and variance
m_star, v_star = gp_model.predict(x_star)
# Plot 1: Mean and 95% predictive interval
fig1, ax1 = plt.subplots()
ax1.set_xlabel('$x$')
ax1.set_ylabel('$y$')
f_lower = m_star - 2.0 * np.sqrt(v_star - gp_model.likelihood.variance)
f_upper = m_star + 2.0 * np.sqrt(v_star - gp_model.likelihood.variance)
y_lower = m_star - 2.0 * np.sqrt(v_star)
y_upper = m_star + 2.0 * np.sqrt(v_star)
ax1.fill_between(x_star.flatten(), y_lower.flatten(), y_upper.flatten(), color='red', alpha=0.25, label='$y^*$ 95% pred.')
ax1.fill_between(x_star.flatten(), f_lower.flatten(), f_upper.flatten(), alpha=0.5, label='$f(\mathbf{x}^*)$ 95% pred.')
ax1.plot(x_star, m_star, lw=2, label='$m_n(x)$')
ax1.plot(x_star, f_true(x_star), 'm-.', label='True function')
ax1.plot(X, Y, 'kx', markersize=10, markeredgewidth=2, label='data')
#plt.legend(loc='best');
# Plot 2: Data plus posterior samples
fig2, ax2 = plt.subplots()
f_post_samples = gp_model.posterior_samples_f(x_star, 10)
ax2.plot(x_star, f_post_samples[:, 0, :], 'r', lw=0.5)
ax2.plot(X, Y, 'kx', markersize=10, markeredgewidth=2, label='data');
ax2.plot(x_star, f_true(x_star), 'm-.', label='True function')
ax2.set_xlabel('$x$')
ax2.set_ylabel('$y$')
#plt.legend(loc='best');
interact_manual(analyze_and_plot_gp_ex1,
kern_variance=(0.01, 1.0, 0.01),
kern_lengthscale=(0.01, 1.0, 0.01),
like_variance=(0.01, 1.0, 0.01));
# -
# ### Questions
#
# In the interactive tool above:
#
# + Experiment with differnet lengthscales for the kernel. You need to click on ``Run Interact`` for the code to run.
# What happens to the posterior mean and the 95% predictive error bar as the lengthscale increases (decreases)?
#
# + Experiment with difference likelihood variances. What happens for very big variances? What happens for very small variances?
#
# + Experiment with different kernel variances. This the $s^2$ parameter of the squared exponential covariance function. It specifies our prior variance about the function values. What is its effect?
#
# + Imagine that, as it would be the case in reality, you do not know the true function. How would you pick the correct values for the hyperparameters specifying the kernel?
#
# + Try some other kernels. Edit the function ``analyze_and_plot_gp_ex1`` and change the line ``k = GPy.kern.RBF(1)`` to ``k = GPy.kern.Matern52(1)``. This is a kernel that is less regular than the RBF. What do you observe?
# Then try ``k = GPy.kern.Matern32(1)``. Then ``k = GPy.kern.Exponential(1)``. The last one is continuous but nowhere differentiable.
# How can you pick the right kernel?
# ## Diagnostics: How do you know if the fit is good?
#
# To objective test the resulting model we need a *validation dataset* consisting of inputs:
# $$
# \mathbf{x}^v_{1:n^v} = \left(\mathbf{x}^v_1,\dots,\mathbf{x}^v_{n^v}\right),
# $$
# and corresponding, observed outputs:
# $$
# \mathbf{y}^v_{1:n^v} = \left(y^v_1,\dots,y^v_{n^v}\right).
# $$
# We will use this validation dataset to define some diagnostics.
# Let's do it directly through the 1D example above.
# First, we generate some validation data:
n_v = 100
X_v = np.random.rand(n_v)[:, None]
Y_v = f_true(X_v) + sigma * np.random.randn(n_v, 1)
# #### Point-predictions
#
# Point-predictions only use $m_n\left(\mathbf{x}^v_i\right)$.
# Of course, when there is a lot of noise, they are not very useful.
# But let's look at what we get anyway.
# (In the questions section I will ask you to reduce the noise and repeat).
#
# The simplest thing we can do is to compare $y^v_i$ to $m_n\left(\mathbf{x}^v_i\right)$.
# We start with the *mean square error*:
# $$
# \operatorname{MSE} := \frac{1}{n^v}\sum_{i=1}^{n^v}\left[y^v_i-m_n\left(\mathbf{x}^v_i\right)\right]^2.
# $$
m_v, v_v = gpm.predict(X_v)
mse = np.mean((Y_v - m_v) ** 2)
print('MSE = {0:1.2f}'.format(mse))
# This is not very intuitive though.
# An somewhat intuitive measure is [coefficient of determination](https://en.wikipedia.org/wiki/Coefficient_of_determination) also known as $R^2$, *R squared*.
# It is defined as:
# $$
# R^2 = 1 - \frac{\sum_{i=1}^{n^v}\left[y_i^v - m_n(\mathbf{x}_i^v)\right]^2}{\sum_{i=1}^{n^v}\left[y_i^v-\bar{y}^v\right]^2},
# $$
# where $\bar{y}^v$ is the mean of the observed data:
# $$
# \bar{y}^v = \frac{1}{n^v}\sum_{i=1}^{n^v}y_i^v.
# $$
# The interpretation of $R^2$, and take this with a grain of salt, is that it gives the percentage of variance of the data explained by the model.
# A score of $R^2=1$, is a perfect fit.
# In our data we get:
R2 = 1.0 - np.sum((Y_v - m_v) ** 2) / np.sum((Y_v - np.mean(Y_v)) ** 2)
print('R2 = {0:1.2f}'.format(R2))
# Finally, on point-predictions, we can simply plot the predictions vs the observations:
fig, ax = plt.subplots(dpi=100)
y_range = np.linspace(Y_v.min(), Y_v.max(), 50)
ax.plot(y_range, y_range, 'r', lw=2)
ax.plot(Y_v, m_v, 'bo')
ax.set_xlabel('Prediction')
ax.set_ylabel('Observation');
# ### Statistical diagnostics
#
# Statistical diagnostics compare the predictive distribution to the distribution of the validation dataset.
# The way to start, are the standarized errors defined by:
# $$
# e_i = \frac{y_i^v - m_n\left(\mathbf{x}^v_i\right)}{\sigma_n\left(\mathbf{x}^v_i\right)}.
# $$
# Now, if our model is correct, the standarized errors must be distributed as a standard normal $N(0,1)$ (why?).
# There are various plots that you can do to test that.
# First, the histogram of the standarized errors:
s_v = np.sqrt(v_v)
e = (Y_v - m_v) / s_v
fig, ax = plt.subplots(dpi=100)
zs = np.linspace(-3.0, 3.0, 100)
ax.plot(zs, st.norm.pdf(zs))
ax.hist(e, density=True, alpha=0.25)
ax.set_xlabel('Std. error')
# Close, but not perfect.
# Another common plot is this:
fig, ax = plt.subplots(dpi=100)
ax.plot(e, 'o')
ax.plot(np.arange(e.shape[0]), 2.0 * np.ones(e.shape[0]), 'r--')
ax.plot(np.arange(e.shape[0]), -2.0 * np.ones(e.shape[0]), 'r--')
ax.set_xlabel('$i$')
ax.set_ylabel('$e_i$');
# Where the red lines indicate the 95% quantiles of the standard normal.
# This means that if 5\% of the errors are inside, then we are good to go.
#
# Yet another plot yielding the same information is the q-q plot comparing the empirical quantiles of the standarized errors to what they are supposed to be, i.e., to the quantiles of $N(0,1)$:
fig, ax = plt.subplots(dpi=100)
st.probplot(e.flatten(), dist=st.norm, plot=ax);
# ### Note on Gaussian process diagnostics
#
# For a more detailed description of GP regression diagnostics, please see this [paper](https://www.jstor.org/stable/40586652).
# ### Questions
#
# + Experiment with larger number of training points $n$. Are the models becoming better according to the metrics we defined above?
# + Experiment with smaller measurement noises $\sigma$. What do you observe? Which diagnostics make sense for very small $\sigma$'s?
# ## Calibrating the Hyperparameters of a Gaussian Process
#
# So, we saw how GP regression works but everything we did was conditional on knowing the hyperparameters of the covariance function, we called them $\theta$, and the likelihood variance $\sigma^2$.
# But if what do we do if we are not sure about them?
# We will do what we always do:
#
# + We summarize our state of knowledge about them by assigning prior probability density $p(\theta)$ and $p(\sigma)$.
#
# + We use the Bayes rule to derive our posterior state of knowledge about them:
# $$
# \begin{array}{ccc}
# p(\theta,\sigma | \mathcal{D}) &\propto& p(\mathcal{D}|\theta,\sigma)p(\theta)p(\sigma) \\
# &=& \int p(\mathbf{y}_{1:n}|\mathbf{f}_{1:n},\sigma) p(\mathbf{f}_{1:n} | \mathbf{x}_{1:n},\theta)d\mathbf{f}_{1:n} p(\theta)p(\sigma).
# \end{array}
# $$
#
# + We somehow approximate this posterior. So far, we only know of one way of approximating this posterior, and that is by a maximum a posteriori estimate.
# ### Making a little bit of analytical progress in the posterior
# We stated without proof that the posterior of the hyperparameters is:
# $$
# p(\theta,\sigma|\mathcal{D}) \propto \int p(\mathbf{y}_{1:n}|\mathbf{f}_{1:n},\sigma)p(\mathbf{f}_{1:n}|\mathbf{x}_{1:n},\theta)d\mathbf{f}_{1:n}p(\theta)p(\sigma).
# $$
# You should ry to familiriaze yourself with these expressions.
# How can you just see the validity of these expressions?
# It's quite simple if you look at the graph.
# So, let's draw the graph.
g3 = Digraph('gp')
g3.node('sigma', label='<σ>')
g3.node('theta', label='<θ>')
g3.node('x1n', label='<<b>x</b><sub>1:n</sub>>', style='filled')
g3.node('f', label='<<b>f</b><sub>1:n</sub>>')
g3.node('y1n', label='<<b>y</b><sub>1:n</sub>>', style='filled')
g3.edge('theta', 'f')
g3.edge('x1n', 'f')
g3.edge('f', 'y1n')
g3.edge('sigma', 'y1n')
g3.render('gp_reg3', format='png')
g3
# The graph, tells you how the joint distribution of all the variables decomposes in conditional probabilities.
# You know that the parent nodes condition the children nodes.
# Here is the decomposition here:
# $$
# p(\mathbf{x}_{1:n}, \mathbf{y}_{1:n}, \mathbf{f}_{1:n}, \theta, \sigma) =
# p(\mathbf{y}_{1:n} | \mathbf{f}_{1:n}, \sigma) p(\mathbf{f}_{1:n} | \mathbf{x}_{1:n}, \theta)p(\theta)p(\sigma)p(\mathbf{x}_{1:n}).
# $$
# Now, by Bayes' rule, we know that the conditional joint probability density of all unobserved variables is proportional to the joint:
# $$
# p(\theta, \sigma, \mathbf{f}_{1:n} | \mathbf{x}_{1:n}, \mathbf{y}_{1:n}) \propto p(\mathbf{x}_{1:n}, \mathbf{y}_{1:n}, \mathbf{f}_{1:n}, \theta, \sigma).
# $$
# The normalization constant does not matter, i.e., we can drop $p(\mathbf{x}_{1:n})$, so we get:
# $$
# p(\theta, \sigma, \mathbf{f}_{1:n} | \mathbf{x}_{1:n}, \mathbf{y}_{1:n}) \propto p(\mathbf{y}_{1:n} | \mathbf{f}_{1:n}, \sigma) p(\mathbf{f}_{1:n} | \mathbf{x}_{1:n}, \theta)p(\theta)p(\sigma).
# $$
# Finally, to get the posterior over $\theta$ and $\sigma$ only, we *marginalize* (i.e., integrate out) the unobserved variable $\mathbf{f}_{1:n}$.
# Here, the integral is actually analytically available (integral of a Gaussian times a Gaussian which is a Gaussian).
# If you do the math, you will get:
# $$
# p(\theta,\sigma | \mathcal{D}) \propto \mathcal{N}\left(\mathbf{y}_{1:n}\middle|
# \mathbf{m}(\mathbf{x}_{1:n}), \mathbf{K}(\mathbf{x}_{1:n},\mathbf{x}_{1:n}) + \sigma^2\mathbf{I}_n\right) p(\theta)p(\sigma).
# $$
# ### Maximum a Posteriori Estimate of the Hyperparameters
#
# In the maximum a posteriori estimate (MAP) we are basically approximating the posterior with the $\delta$-function centered at its pick.
# That is, we are approximating:
# $$
# p(\theta,\sigma|\mathcal{D}) \approx \delta(\theta-\theta^*)\delta(\sigma-\sigma^*),
# $$
# where $\theta^*$ and $\sigma^*$ are maximizing $\log p(\theta,\sigma|\mathcal{D})$.
# It is instructive to see what $\log p(\theta,\sigma|\mathcal{D})$ looks like and see if we can assign any intuitive meaning to its terms.
# It is:
# $$
# \begin{array}{ccc}
# \log p(\theta,\sigma|\mathcal{D}) &=& \log \mathcal{N}\left(\mathbf{y}_{1:n}\middle|
# \mathbf{m}(\mathbf{x}_{1:n}), \mathbf{K}(\mathbf{x}_{1:n},\mathbf{x}_{1:n}) + \sigma^2\mathbf{I}_n\right) + \log p(\theta) + \log p(\sigma) \\
# &=&
# -\frac{1}{2}\left(\mathbf{y}_{1:n}-\mathbf{m}(\mathbf{x}_{1:n})\right)^T\left(\mathbf{K}(\mathbf{x}_{1:n},\mathbf{x}_{1:n}) + \sigma^2\mathbf{I}_n\right)^{-1}\left(\mathbf{y}_{1:n}-\mathbf{m}(\mathbf{x}_{1:n})\right)\\
# &&-\frac{1}{2}\log |\mathbf{K}(\mathbf{x}_{1:n},\mathbf{x}_{1:n}) + \sigma^2\mathbf{I}_n|\\
# &&+\log p(\theta) + \log p(\sigma)\\
# && + \text{constants}.
# \end{array}
# $$
# The constants are terms that do not depend on $\theta$ or $\sigma$.
# The first term is a familiar one.
# It kind of looks like least squares (it is actually a form of least squares).
# The third and forth term are familiar regularizers stemming from our prior knowledge.
# The second term is a naturally occuring regularizer.
#
# Now, back to solving the optimization problem that yields the MAP.
# Of course, you need to get the deratives of $\log p(\theta,\sigma|\mathcal{D})$ and use an optimization algorithm.
# Back in the stone age, we were doing this by hand.
# Now you don't have to worry about it.
# Automatic differentiation can work through the entire expression (including the matrix determinant).
# Once you have the derivative you can use a gradient-based optimization algorithm from [scipy.optimize](https://docs.scipy.org/doc/scipy/reference/optimize.html).
# ``GPy`` is using by default the [L-BFGS algorithm](https://docs.scipy.org/doc/scipy/reference/optimize.minimize-lbfgsb.html) but you could change it if you want.
# These are minimization algorithms.
# So, ``GPy`` is actually minimizing $-\log p(\theta,\sigma)$.
# Let's see how it works through our previous example:
# ## Example: Gaussian process regression with fitted hyperparameters
#
# Make sure that you still have the data from the previous example:
# Fixing the seed so that we all see the same data
np.random.seed(1234)
n = 10
# The inputs are in [0, 1]
X = np.random.rand(n, 1) # Needs to be an n x 1 vector
# The outputs are given from a function plus some noise
# The standard deviation of the noise is:
sigma = 0.4
# The true function that we will try to identify
# Some data to train on
Y = f_true(X) + sigma * np.random.randn(X.shape[0], 1)
fig, ax = plt.subplots(dpi=100)
ax.plot(X, Y, 'kx', markersize=10, markeredgewidth=2)
ax.set_xlabel('$x$')
ax.set_ylabel('$y$');
# Let's pick a squared exponential kernel and make a model with Gaussian likelihood (the default choice):
k = GPy.kern.RBF(1) # GPy.kern.RBF(60, ARD=True)
gpm = GPy.models.GPRegression(X, Y, k)
print(gpm)
# Let's explain what all this means.
# Notice that there are some default values for the hyperparameters (they are all one).
# Also, notice that ``GPy`` is keeping track of how many parameters it needs to fit.
# Here we have three parameters ($s,\ell$ and $\sigma$).
# The second column are constraints for the parameters.
# The ``+ve`` term means that the corresponding hyperparamer has to be positive.
# Notice that there is nothing in the ``priors`` column.
# This is because we have set no priors right now.
# When this happens, ``GPy`` assumes that we assign a flat prior, i.e., here it assumes that we have assigned $p(\theta)\propto 1$ and $p(\sigma)\propto 1$.
# That's not the best choice, but it should be ok for now.
#
# Now, pay attention to the ``Objective``. This is the $-\log p(\theta,\sigma|\mathcal{D})$ for the current choice of parameters.
# Let's now find the MAP:
gpm.optimize_restarts(messages=True) # we use multiple restarts to avoid being trapped to a local minimum
print(gpm)
# Ok. We did find something with higher posterior value.
# Let's plot the prediction:
fig, ax = plt.subplots(dpi=100)
ax.set_xlabel('$x$')
ax.set_ylabel('$y$')
m_star, v_star = gpm.predict(x_star)
f_lower = m_star - 2.0 * np.sqrt(v_star - gpm.likelihood.variance)
f_upper = m_star + 2.0 * np.sqrt(v_star - gpm.likelihood.variance)
y_lower = m_star - 2.0 * np.sqrt(v_star)
y_upper = m_star + 2.0 * np.sqrt(v_star)
ax.fill_between(x_star.flatten(), y_lower.flatten(), y_upper.flatten(), color='red', alpha=0.25, label='$y^*$ 95% pred.')
ax.fill_between(x_star.flatten(), f_lower.flatten(), f_upper.flatten(), alpha=0.5, label='$f(\mathbf{x}^*)$ 95% pred.')
ax.plot(x_star, m_star, lw=2, label='$m_n(x)$')
ax.plot(x_star, f_true(x_star), 'm-.', label='True function')
ax.plot(X, Y, 'kx', markersize=10, markeredgewidth=2, label='data');
# Admitedly, this doesn't look very good.
# (Of course, we can tell this only because we know the truth).
# It seems that the assign lengthscale is too small.
# Also, the likelihood variance seems smaller than it really is.
# What do we do now?
# You have two choices:
# + You encode some prior knowledge and repeat.
# + You add some more data and repeat.
#
# Let's start with some prior knowledge and let the other item for the questions section.
# Let's say that we know that the noise variance.
# How do we encode this?
# Here you go:
gpm.likelihood.variance.constrain_fixed(sigma ** 2)
print(gpm)
# Notice that it now ``GPy`` reports that the likelihood variance is fixed.
# Let's repeat the optimization:
gpm.optimize_restarts(messages=True) # we use multiple restarts to avoid being trapped to a local minimum
print(gpm)
fig, ax = plt.subplots(dpi=100)
ax.set_xlabel('$x$')
ax.set_ylabel('$y$')
m_star, v_star = gpm.predict(x_star)
f_lower = m_star - 2.0 * np.sqrt(v_star - gpm.likelihood.variance)
f_upper = m_star + 2.0 * np.sqrt(v_star - gpm.likelihood.variance)
y_lower = m_star - 2.0 * np.sqrt(v_star)
y_upper = m_star + 2.0 * np.sqrt(v_star)
ax.fill_between(x_star.flatten(), y_lower.flatten(), y_upper.flatten(), color='red', alpha=0.25, label='$y^*$ 95% pred.')
ax.fill_between(x_star.flatten(), f_lower.flatten(), f_upper.flatten(), alpha=0.5, label='$f(\mathbf{x}^*)$ 95% pred.')
ax.plot(x_star, m_star, lw=2, label='$m_n(x)$')
ax.plot(x_star, f_true(x_star), 'm-.', label='True function')
ax.plot(X, Y, 'kx', markersize=10, markeredgewidth=2, label='data');
# This looks better.
# But it seems that the automatically selected lengthscale is smaller than the true one.
# (Of course don't really know the true lengthscale is).
# Let's assign a prior probability density on the lengthscale which pushes it to be greater.
# Since we are dealing with a positie parameter and we don't know much about it, let's assign an exponential prior with a rate of 2 (which will yield an expectation of 0.5):
# $$
# \ell \sim \operatorname{Log-N}(0.2, 1).
# $$
ell_prior = GPy.priors.LogGaussian(.2, 1.0)
# Let's visualize it to make sure it's ok
fig, ax = plt.subplots(dpi=100)
ells = np.linspace(0.01, 2.0, 100)
ax.plot(ells, ell_prior.pdf(ells))
# Now here is how you can set it:
gpm.kern.lengthscale.set_prior(ell_prior)
print(gpm)
gpm.optimize_restarts(messages=True)
print(gpm)
fig, ax = plt.subplots(dpi=100)
ax.set_xlabel('$x$')
ax.set_ylabel('$y$')
m_star, v_star = gpm.predict(x_star)
f_lower = m_star - 2.0 * np.sqrt(v_star - gpm.likelihood.variance)
f_upper = m_star + 2.0 * np.sqrt(v_star - gpm.likelihood.variance)
y_lower = m_star - 2.0 * np.sqrt(v_star)
y_upper = m_star + 2.0 * np.sqrt(v_star)
ax.fill_between(x_star.flatten(), y_lower.flatten(), y_upper.flatten(), color='red', alpha=0.25, label='$y^*$ 95% pred.')
ax.fill_between(x_star.flatten(), f_lower.flatten(), f_upper.flatten(), alpha=0.5, label='$f(\mathbf{x}^*)$ 95% pred.')
ax.plot(x_star, m_star, lw=2, label='$m_n(x)$')
ax.plot(x_star, f_true(x_star), 'm-.', label='True function')
ax.plot(X, Y, 'kx', markersize=10, markeredgewidth=2, label='data');
# That's better, but not perfect.
# But remember: You don't know what the truth is...
# ## Questions
#
# Let's investigate what happens to the previous examples as you increase the number of observations.
#
# + Rerun everything gradually increasing the number of samples from $n=10$ to $n=100$.
# Notice that as the number of samples increases it doesn't really matter what your prior knowledge is.
# As a matter of fact, for the largest number of samples you try, pick a very wrong prior probability for $\ell$.
# See what happens.
#
# + Rerun everything with $\sigma=0.01$ (small noise) and gradually increasing the number of samples from $n=10$ to $n=100$.
# For small noise, the model is trying to interpolate.
# Is it capable of figuring out that the noise is small when the number of observations is limited? When does the method realize the noise is indeed small?
| lectures/lecture_15.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/risker93/Hello_World/blob/main/algorithm/ai_week_10.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="lj1KGzpq7gOa"
# #퍼셉트론
#
# - 입력 값을 놓고 활성화 함수에 의해 일정한 수준을 넘으면 참을1, 그렇지 않으면 거짓0을 내보내는 이 간단한 회로가 하는 일이 뉴런과 같음.
#
# - 여러 층의 퍼셉트론을 서로 연결시키고 복잡하게 조합하여 주어진 입력 값에 대한 판단을 하게 하는것, 그것이 바로 신경망의 기본 구조임.
# - 신경망을 이루는 가장 중요한 기본단위는 퍼셉트론(perceptron)
# - 퍼셉트론은 입력 값과 활성화 함수를 사용해 출력 값을 다음으로 넘기는 가장 작은 신경망 단위
#
# - 기울기a 나 y절편 b 같은 용어를 퍼셉트론의 개념에 맞춰 좀 더 '딥러닝 답게' 표현해 보면 다음과 같음
#
# y = ax+b (a는 기울기, b는 y절편)
#
# -> y= wx + b (w는 가중치, b는 바이어스)
#
# - 가중합: 입력값(x)과 가중치(w)의 곱을 모두 더한 다음 거기에 바이어스(b)를 더한 값.
#
# - 활성화 함수(activation function):
# 가중합의 결과를 놓고 1또는 0을 출력해서 다음으로 보냄,
# 여기서 0과 1을 판단하는 함수
#
# - 단 하나의 퍼셉트론으로는 많은 것을 기대할 수가 없음
# - 퍼셉트론의 한계와 이를 해결하는 과정을 보며 신경망의 기본 개념을 확립해 보자
#
#
# #XOR 문제
# 논리 회로에 등장하는 개념
#
# - 게이트(gate): 컴퓨터는 두 가지의 디지털 값, 즉 0과 1을 입력해 하나의 값을 출력하는 회로가 모여 만들어지는 회로.
#
# AND진리표 OR진리표 XOR진리표
# 0 0 0 0 0 0 0 0 0
# 0 1 0 0 1 1 0 1 1
# 1 0 0 1 0 1 1 0 1
# 1 1 1 1 1 1 1 1 0
#
# AND (conjunction) 논리곱 , 둘다 참일때만 참 , 나머진 거짓
#
# OR (disjunction) 논리합, 둘다 거짓일때만 거짓, 나머진 참
#
# NAND (negation of conjunction) 부정 논리곱, 둘다 참일때만 거짓, 나머진 참
#
# NOR(negation of disjunction) 부정 논리합, 둘다 거짓일때만 참, 나머진 거짓
#
# XOR(exclusive disjunction) 베타적 논리합 , 둘의 참, 거짓이 다르면 참, 같으면 거짓
#
# XNOR(exclusive negation of disjunction)베타적부정논리합 둘의 참 거짓이 다르면 거짓, 같으면 참.
#
# - AND와 OR게이트는 직선을 그어 결과값 0, 1을 구별 할 수 있음.
# - XOR의 경우 선을 그어 구분할 수 없음
# 뉴런 → 신경망 → 지능 이라는 도식을 따라 퍼셉트론 → 인공신경망 → 인공지능 이 가능하리라 꿈꾸던 당시 사람들은 이것이 생각처럼 쉽지 않다는 사실을 깨닫게 됨
#
# - 알고 보니 간단한 XOR 문제조차 해결할 수 없었음
# - 이후 10여년이 지난 후에야 이 문제가 해결되는데, 이를 해결한 개념이 바로 다층 퍼셉트론(multilayer perceptron)
#
#
# + [markdown] id="4zW4TyNjCINN"
# #다층 퍼셉트론
# - 종이 위에 각각 엇갈려 놓인 검은점 두개와 흰점 두개를 하나의 선으로는 구별 할 수 없다는 것을 살펴봄
#
# - 언뜻 보기에 해답이 없어 보이는 이 문제를 해결하려면 새로운 접근이 필요함.
# - 골똘히 연구해도 답을 찾지 못했던 이 문제는 2차원 평면에서만 해결하려는 고정 관념을 깨고 피라미드 모양으로 성냥개비를 쌓아 올리니 해결됨.
#
# - 인공 신경망을 개발하기 위해서는 반드시 XOR문제를 극복 해야만 했음
# - 이 문제 역시 고정관념을 깬 기발한 아이디어에서 해결점이 보였음.
#
# - 좌표 평면 자체에 변화를 주는것
# - XOR 문제를 해결하기 위해서 우리는 두 개의 퍼셉트론을 한 번에 계산할 수 있어야 함
# - 이를 가능하게 하려면 숨어있는 층, 즉 은닉층(hidden layer)을 만들면 됨
#
# - 딥러닝의 딥이 의미 하는것은 은닉층이 2개 이상이면 딥.
#
# - 딥러닝은 입력과 출력은 아는데 안에를 모름 그래서 블랙박스 모델이라 부름
#
# - 그거를 찾는 방법중 하나가 오차 역전파.
#
# - 은닉층이 좌표 평면을 왜곡시키는 결과를 가져옴.
# + colab={"base_uri": "https://localhost:8080/"} id="DdvdkkKEEp_J" outputId="dbfaee12-ef3d-4b3f-e9fb-bf14571167aa"
#가짜 동전 찾기 알고리즘 - 반씩 그룹으로 나누어 비교하기
#주어진 동전 n개 중에 가짜 동전(fake)을 찾아내는 알고리즘
#입력: 전체 동전 위치의 시작과 끝(0, n -1)
#출력: 가짜 동전의 위치 번호
#무게 재기 함수
#a에서 b까지 놓인 동전과
#c에서 d까지 넣인 동전의 무게를 비교
#a에서 b까지 가짜 동전이 있으면(가벼우면): -1
#c에서 d까지 가짜 동전이 있으면(가벼우면): 1
#가짜 동전이 없으면(양쪽 무게가 같으면):0
def weigh(a,b,c,d):
fake = 29 #가짜 동전의 위치(알고리즘은 weigh() 함수를 이용하여 이 값을 맞혀야 함)
if a <= fake and fake <= b:
return -1
if c <= fake and fake <= d:
return 1
return 0
#weigh()함수(저울질) 를 이용하여 left와 right 까지에 놓인 가짜 동전의 위치를 찾아냄
def find_fakecoin(left, right):
#종료 조건: 가짜 동전이 있을 범위 안에 동전이 한 개뿐이면 그 동전이 가짜 동전임
#left, right 는 어디서부터 어디까지 숫자임 즉 left와 right가 같다는 말은둘이 같은 숫자 라는말.
if left == right:
return left
#left 와 right 까지 놓인 동전을 두 그룹(g1_left~ g1_right, g2_left~g2_right) 으로 나눔
#동전 수가 홀수면 두 그룹으로 나누고 한개가 남음
half = (right - left + 1)//2
g1_left = left
g1_right = left + half - 1
g2_left = left + half
g2_right = g2_left + half - 1
#나눠진 두 그룹을 weigh() 함수를 이용하여 저울질함
result = weigh(g1_left, g1_right, g2_left, g2_right)
if result == -1: #그룹 1이 가벼움( 가짜 동전이 이 그룹에 있음)
return find_fakecoin(g1_left, g1_right) # 그룹 1 범위를 재귀호출로 다시 조사
elif result == 1: #그룹 2가 가벼움(가짜 동전이 이 그룹에 있음)
return find_fakecoin(g2_left, g2_right) #그룹2 범위를 재귀 호출로 다시 조사
else: #두 그룹의 무게가 같으면
return right #두 그룹으로 나뉘지 않고 남은 나머지 한개의 동전이 가짜
n = 100 #전체 동전 개수
print(find_fakecoin(0, n - 1))
# + [markdown] id="vL6F_W6OPzkg"
# - 이 문제의 알고리즘 효율성을 '저울질 횟수' 를 기준으로 생각해 보자
# - 0번 동전과 나머지 동전을 일일이 비교하는 방법인 프로그램은 17-1은 저울질이 최대 n-1번 필요 → 계간 복잡도가 O(n)
# - 동전 n개를 절반씩 나누어 후보를 좁히며 비교하는 방법은 → 계산 복잡도가 O(log n)
#
# - 가짜 동전 문제와 순차/ 이분 탐색 알고리즘 비교
# - 순차 탐색에서는 하나씩 비교하여 값을 찾아내므로 계산 복잡도가 O(n)
# - 이분 탐색에서는 리스트가 이미 정렬되어 있다는 것을 전제로, 중간 값을 비교한 후 값이 있을 가능성이 없는 절반을 제외해 나가면서 값을 찾아내므로 계산 복잡도가 O(log n)
# - 리스트 탐색 문제와 가짜 동전 문제는 잘 생각해보면 구조가 비슷한 문제
#
#
# + id="VHgyOuMmYs9t"
#이중 포문 이해를 위해 아래 코드에서 잘라옴
n = 10
for i in range(0, n - 1):
for j in range(i+1, n):
print(i,'',j)
# + colab={"base_uri": "https://localhost:8080/"} id="rJkwfddKW8Ja" outputId="26bfe04f-d5c1-40e4-8e97-cb6b8389fe76"
#최대 수익을 구하는 알고리즘
#주어진 주식 가격을 보고 얻을 수 있는 최대 수익을 구하는 알고리즘
#입력: 주식 가격의 변화 값(리스트 prices)
#출력: 한 주를 한번 사고 팔아 얻을 수 있는 최대 수익 값
def max_profit(prices):
n = len(prices)
max_profit = 0 #최대 수익은 항상 0 이상의 값
for i in range(0, n - 1):
for j in range(i+1, n): #첫번째 포문에서 가져온 값을 빼고 나머지 값을 모두 비교
profit = prices[j] - prices[i] #i날에 사서 j날에 팔았을 때 얻을 수 있는 수익
if profit > max_profit: #이 수익이 지금까지 최대 수익보다 크면 값을 고침
max_profit = profit
return max_profit
stock = [10300, 9600, 9800, 8200, 7800, 8300, 9500, 9800, 10200, 9500]
print(max_profit(stock))
# + id="-EL_WKb0YzjZ" colab={"base_uri": "https://localhost:8080/"} outputId="4aeafa67-cd78-4fd6-bac8-a9fda325c8f3"
#최대 수익을 구하는 알고리즘2
#한번 반복으로 최대 수익 찾기
#주어진 주식 가격을 보고 얻을 수 있는 최대 수익을 구하는 알고리즘
#입력: 주식 가격의 변화값(prices)
#출력: 한 주를 한번 사고 팔아 얻을 수 있는 최대 수익 값
def max_profit(prices):
n = len(prices)
max_profit = 0 #최대 수익은 항상 0 이상의 값
min_price = prices[0] #첫째 날의 주가를 주가의 최솟값으로 기억
for i in range(1, n): #1부터 n-1까지 반복
profit = prices[i] - min_price #지금까지의 최솟값에 주식을 사서 i날에 팔 때의 수익
if profit > max_profit: #이 수익이 지금까지 최대 수익보다 크면 값을 고침
max_profit = profit
if prices[i] < min_price: #i날 주가가 치솟값보다 작으면 값을 고침
min_price = prices[i]
return max_profit
stock = [10300, 9600, 9800, 8200, 7800, 8300, 9500, 9800, 10200, 9500]
print(max_profit(stock))
# + [markdown] id="uWRJJ4jjqPi4"
# 첫 번째 알고리즘보다 두 번째 알고리즘이 더 효율적임
#
# 모든 경우를 비교한 첫 번째 알고리즘은 동명이인 찾기와 비슷한 구조 → 계산 복잡도는O(n2)
#
# 리스트를 한 번 탐색하면서 최대 수익을 계산한 두 번째 알고리즘은 문제 2 최댓값 찾기와 비슷한 구조 → 계산 복잡도는 O(n)
#
#
# + id="qAAbUthYqwYg"
#최대 수익을 구하는 두 알고리즘의 계산 속도를 비교하기
#최대 수익 문제를 푸는 두 알고리즘의 계산 속도 비교하기
#최대 수익 문제를 O(n*n)과 O(n)으로 푸는 알고리즘을 각각 수행하여
#걸린 시간을 출력/비교함
import time #시간 측정을 위한 time 모듈
import random #테스트 주가 생성을 위한 random 모듈
#최대 수익: 느린O(n*n)알고리즘
def max_porfit_slow(prices):
n = len(prices)
max_profit = 0
for i in range(0, n-1):
for j in range(i +1, n ):
profit = prices[j] - prices[i]
if profit > max_profit:
max_profit = profit
return max_profit
#최대 수익: 빠른O(n) 알고리즘
def max_profit_fast(prices):
n = len(prices)
max_profit = 0
min_price = prices[0]
for i in range(1, n):
profit = prices[i] - min_price
if profit > max_profit:
max_profit = profit
if prices[i] < min_price:
min_price = prices[i]
return max_profit
def test(n):
#테스트 자료 만들기(5000부터 20000까지의 난수를 주가로 사용)
a = []
for i in range(0,n):
a.append(random.randint(5000,20000))
#느린 O(n*n) 알고리즘 테스트
start = time.time() #계산 시작 직전 시각을 기억
mps = max_porfit_slow(a) #계산 수행
end = time.time() #계산 시작 직후 시각을 기억
time_slow = end-start #두 시각을 빼면 계산에 걸린 시간
#빠른 O(n)알고리즘 테스트
start = time.time() #계산시간 직전 시간을 기억
mpf = max_profit_fast(a) #계산수행
end = time.time() #계산 시작 직후 시각을 기억
time_fast = end - start #두 시각을 빼면 계산에 걸린 시간
#결과 출력: 계산 결과
print(n, mps, mpf) #입력 크기, 각각 알고리즘이 계산한 최대 수익 값(같아야함)
m = 0 #느린 알고리즘과 빠른 알고리즘의 수행 시간 비율을 저장할 변수
if time_fast > 0: #컴퓨터 환경에 따라 빠른 알고리즘 시간이 0으로 측정 될 수 있음( 이럴때는 0을출력)
m = time_slow/time_fast #느린 알고리즘시간/ 빠른 알고리즘 시간
#입력크기, 느린알고리즘수행시간, 빠른 알고리즘 수행시간, 느린알고/빠른알고 시간
print("%d %5.f %5f %.2f" %(n, time_slow, time_fast, m))
#test(100)
test(1000)
# + [markdown] id="0QWsc2BmxAlm"
# #다익스트라 알고리즘
# 1. 최단 경로 문제
# - 그래프(노드와 선으로 연결된 그래프임) 내의 한 정점에서 다른 정점으로 이동할때 찾는 가중치 합이 최소값이 되도록 만드는 경로를 찾는 알고리즘
# - 가중간선(Edge Weight) : 간선들이 할당된 값을 가짐
# - 가중 경로(Path Weight): 경로에 속하는 모든 간선의 값을 더한 값
#
# - 모든 간선 가중치가 음이 아닌 일반적인 경우
# - 다익스트라 알고리즘
# - 음의 가중치가 존재하는 경우
# - 벨만 -포드 알고리즘
# - 음의 가중치는 허용하지만 가중치 합이 음인 사이클은 절대 허용하지 않음
# - 음의 사이클이 존재하면 해당 사이클을 계속 반복한다면 가중치 합이 무한이 작아질 수 있으므로 최단 경로 문제 자체가 성립하지 않음
#
# 2. 다익스트라 알고리즘
# - 어떠한 간선도 음수 값을 갖지 않는 방향 그래프에서 주어진 출발점과 도착점 사이의 최단 경로를 찾아주는 알고리즘
# - 방향그래프, 무방향 그래프 모두가능
# - 프림 알고리즘이 단순히 간선의 길이를 이용해 어떤 간선을 먼저 연결할지를 결정하는데 반해 다익스트라 알고리즘은 경로의 길이를 감안해서 간선을 연결함
# - 하나의 출발점에서 다른 모든 정점까지의 최단 경로를 구하는 방법
# - 모든 노드를 순회해야 하므로 시간복잡도에 결정적인 영향을 미치게 되는데 제대로 구현된 우선순위 큐를 활용하면 비용을 줄일 수 있음
# - 처음 고안한 알고리즘은 O(V^2)의 시간 복잡도를 가졌으나 이후 우선순위 큐(=힙 트리) 등을 이용하면 더욱 개선된 알고리즘이 나와 O(E logV) 의 시간복잡도를 가짐
# - 다익스트라 알고리즘은 벨만-포드 알고리즘과 동일단 작업을 수행하고 실행 속도도 더 빠름
# - 다익스트라 알고리즘은 가중치가 음수인 경우는 처리할 수 없으므로 이런 경우에는 벨만-포드 알고리즘을 사용함
#
# - 가능한 적은 비용으로 가장 빠르게 해답에 도달하는 경로를 찾아내는 대부분의 문제에 응용됨
#
# - 내비게이션에서 각 도시들을 정점으로 도로들을 간선으로 갖는 그래프로 간주한다면 두 도시를 잇는 가장 빠른 길을 찾는 문제
#
# - 미로 탐색 알고리즘으로 사용할 수 있음
# - 라우팅에서도 IP 라우팅 프로토콜의 한 종류인 OSPF 방식의 프로토콜에서 사용
#
#
# + [markdown] id="z_bU7MxE3r9D"
# 1. 다익스트라 알고리즘
# 1. 각 정점 위에 시작점으로 부터 자신에게 이르는 경로의 길이를 저장할 곳을 준비하고 모든 정점 위에 있는 경로의 길이를 무한대로 초기화함
# 2. 시작 정점의 경로 길이를 0으로 초기화하고(시작 정점에서 시작 정점까지의 거리는 0 이기 때문) 최단 경로에 추가함
# 3. 최단 경로에 새로 추가된 정점의 인접 정점들에 대해 경로 길이를 갱신하고 이들을 최단 경로에 추가함, 만약 추가하려는 인접 정점이 이미 최단 경로안에 존재한다면 갱신되기 이전의 경로 길이가 새로운 경로의 길이보다 더 큰 경우에 한해 다른 선행 정점을 지나던 기존의 경로를 현재 정점을 경유하도록 수정함
# 4. 그래프 내의 모든 정점이 최단 경로에 소속될 때까지 3 과정을 반복함.
| algorithm/ai_week_10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
data_path = "pokemon_data.csv"
pokemon_data = pd.read_csv(data_path, index_col=0)
pokemon_data.head()
pokemon_data_numeric = pokemon_data[["Attack", "Defense"]]
pokemon_data_numeric.head()
pokemon_data.plot.scatter(x="Attack", y="Defense", figsize=(16,6))
pokemon_data.plot.scatter(x="Total", y="HP", figsize=(16,6))
pokemon_data["Type 1"].value_counts().plot.bar(figsize=(16,6))
pokemon_data.plot.hexbin(x="Attack", y="Defense", figsize=(14,8), gridsize=20)
pokemon_stats = pokemon_data.groupby(["Generation", "Legendary"]).mean()[["Attack", "Defense"]]
pokemon_stats.plot.bar(stacked=True, figsize=(16,6))
pokemon_stats = pokemon_data.groupby(["Generation", "Legendary"]).mean()[["Attack", "Defense"]]
pokemon_stats.plot.line(stacked=True, figsize=(16,6))
pokemon_stats_gen = pokemon_data.groupby("Generation").mean()[["HP", "Attack", "Defense", "Sp. Atk", "Sp. Def", "Speed"]]
pokemon_stats_gen.plot.line(figsize=(16,6))
plt.figure(figsize=(16,6))
sns.countplot(pokemon_data["Generation"])
grid = sns.jointplot(x="Attack", y="Defense", data=pokemon_data)
grid.fig.set_figwidth(14)
grid.fig.set_figheight(8)
plt.figure(figsize=(16,6))
sns.distplot(pokemon_data["HP"])
grid = sns.jointplot(x="Attack", y="Defense", data=pokemon_data, kind="hex")
grid.fig.set_figwidth(10)
grid.fig.set_figheight(10)
plt.figure(figsize=(12,8))
sns.kdeplot(pokemon_data["Attack"], pokemon_data["Defense"])
plt.figure(figsize=(16,6))
sns.boxplot(pokemon_data["Type 1"], pokemon_data["Attack"])
plt.figure(figsize=(16,6))
sns.violinplot(pokemon_data["Type 2"], pokemon_data["Attack"])
plt.figure(figsize=(16,6))
sns.countplot(pokemon_data["Type 1"])
| 5. Pokemon Data Analysis/pokemon_data_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# *Accompanying code examples of the book "Introduction to Artificial Neural Networks and Deep Learning: A Practical Guide with Applications in Python" by [<NAME>](https://sebastianraschka.com). All code examples are released under the [MIT license](https://github.com/rasbt/deep-learning-book/blob/master/LICENSE). If you find this content useful, please consider supporting the work by buying a [copy of the book](https://leanpub.com/ann-and-deeplearning).*
#
# Other code examples and content are available on [GitHub](https://github.com/rasbt/deep-learning-book). The PDF and ebook versions of the book are available through [Leanpub](https://leanpub.com/ann-and-deeplearning).
# %load_ext watermark
# %watermark -a '<NAME>' -v -p torch
# - Runs on CPU or GPU (if available)
# # Model Zoo -- Ordinal Regression CNN -- Beckham and Pal 2016
# Implementation of a method for ordinal regression by Beckham and Pal [1] applied to predicting age from face images in the AFAD [2] (Asian Face) dataset using a simple ResNet-34 [3] convolutional network architecture.
#
# Note that in order to reduce training time, only a subset of AFAD (AFAD-Lite) is being used.
#
# - [1] Beckham, Christopher, and <NAME>. "[A simple squared-error reformulation for ordinal classification](https://arxiv.org/abs/1612.00775)." arXiv preprint arXiv:1612.00775 (2016).
# - [2] Niu, Zhenxing, <NAME>, <NAME>, <NAME>, and <NAME>. "[Ordinal regression with multiple output cnn for age estimation](https://ieeexplore.ieee.org/document/7780901/)." In Proceedings of the IEEE conference on computer vision and pattern recognition, pp. 4920-4928. 2016.
# - [3] He, Kaiming, <NAME>, <NAME>, and <NAME>. "[Deep residual learning for image recognition](http://openaccess.thecvf.com/content_cvpr_2016/html/He_Deep_Residual_Learning_CVPR_2016_paper.html)." In Proceedings of the IEEE conference on computer vision and pattern recognition, pp. 770-778. 2016.
#
#
#
# ## Imports
# +
import time
import numpy as np
import pandas as pd
import os
import torch.nn as nn
import torch.nn.functional as F
import torch
from torch.utils.data import Dataset
from torch.utils.data import DataLoader
from torchvision import transforms
from PIL import Image
if torch.cuda.is_available():
torch.backends.cudnn.deterministic = True
# -
# ## Downloading the Dataset
# !git clone https://github.com/afad-dataset/tarball-lite.git
# !cat tarball-lite/AFAD-Lite.tar.xz* > tarball-lite/AFAD-Lite.tar.xz
# !tar xf tarball-lite/AFAD-Lite.tar.xz
# +
rootDir = 'AFAD-Lite'
files = [os.path.relpath(os.path.join(dirpath, file), rootDir)
for (dirpath, dirnames, filenames) in os.walk(rootDir)
for file in filenames if file.endswith('.jpg')]
# -
len(files)
# +
d = {}
d['age'] = []
d['gender'] = []
d['file'] = []
d['path'] = []
for f in files:
age, gender, fname = f.split('/')
if gender == '111':
gender = 'male'
else:
gender = 'female'
d['age'].append(age)
d['gender'].append(gender)
d['file'].append(fname)
d['path'].append(f)
# -
df = pd.DataFrame.from_dict(d)
df.head()
df['age'].min()
df['age'] = df['age'].values.astype(int) - 18
np.random.seed(123)
msk = np.random.rand(len(df)) < 0.8
df_train = df[msk]
df_test = df[~msk]
df_train.set_index('file', inplace=True)
df_train.to_csv('training_set_lite.csv')
df_test.set_index('file', inplace=True)
df_test.to_csv('test_set_lite.csv')
num_ages = np.unique(df['age'].values).shape[0]
print(num_ages)
# ## Settings
# +
##########################
### SETTINGS
##########################
# Device
DEVICE = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
NUM_WORKERS = 4
NUM_CLASSES = 22
BATCH_SIZE = 512
NUM_EPOCHS = 150
LEARNING_RATE = 0.0005
RANDOM_SEED = 123
GRAYSCALE = False
TRAIN_CSV_PATH = 'training_set_lite.csv'
TEST_CSV_PATH = 'test_set_lite.csv'
IMAGE_PATH = 'AFAD-Lite'
# -
# ## Dataset Loaders
# +
class AFADDatasetAge(Dataset):
"""Custom Dataset for loading AFAD face images"""
def __init__(self, csv_path, img_dir, transform=None):
df = pd.read_csv(csv_path, index_col=0)
self.img_dir = img_dir
self.csv_path = csv_path
self.img_paths = df['path']
self.y = df['age'].values
self.transform = transform
def __getitem__(self, index):
img = Image.open(os.path.join(self.img_dir,
self.img_paths[index]))
if self.transform is not None:
img = self.transform(img)
label = self.y[index]
return img, label
def __len__(self):
return self.y.shape[0]
custom_transform = transforms.Compose([transforms.Resize((128, 128)),
transforms.RandomCrop((120, 120)),
transforms.ToTensor()])
train_dataset = AFADDatasetAge(csv_path=TRAIN_CSV_PATH,
img_dir=IMAGE_PATH,
transform=custom_transform)
custom_transform2 = transforms.Compose([transforms.Resize((128, 128)),
transforms.CenterCrop((120, 120)),
transforms.ToTensor()])
test_dataset = AFADDatasetAge(csv_path=TEST_CSV_PATH,
img_dir=IMAGE_PATH,
transform=custom_transform2)
train_loader = DataLoader(dataset=train_dataset,
batch_size=BATCH_SIZE,
shuffle=True,
num_workers=NUM_WORKERS)
test_loader = DataLoader(dataset=test_dataset,
batch_size=BATCH_SIZE,
shuffle=False,
num_workers=NUM_WORKERS)
# -
# ## Model
# +
##########################
# MODEL
##########################
def conv3x3(in_planes, out_planes, stride=1):
"""3x3 convolution with padding"""
return nn.Conv2d(in_planes, out_planes, kernel_size=3, stride=stride,
padding=1, bias=False)
class BasicBlock(nn.Module):
expansion = 1
def __init__(self, inplanes, planes, stride=1, downsample=None):
super(BasicBlock, self).__init__()
self.conv1 = conv3x3(inplanes, planes, stride)
self.bn1 = nn.BatchNorm2d(planes)
self.relu = nn.ReLU(inplace=True)
self.conv2 = conv3x3(planes, planes)
self.bn2 = nn.BatchNorm2d(planes)
self.downsample = downsample
self.stride = stride
def forward(self, x):
residual = x
out = self.conv1(x)
out = self.bn1(out)
out = self.relu(out)
out = self.conv2(out)
out = self.bn2(out)
if self.downsample is not None:
residual = self.downsample(x)
out += residual
out = self.relu(out)
return out
class ResNet(nn.Module):
def __init__(self, block, layers, num_classes, grayscale):
self.num_classes = num_classes
self.inplanes = 64
if grayscale:
in_dim = 1
else:
in_dim = 3
super(ResNet, self).__init__()
self.conv1 = nn.Conv2d(in_dim, 64, kernel_size=7, stride=2, padding=3,
bias=False)
self.bn1 = nn.BatchNorm2d(64)
self.relu = nn.ReLU(inplace=True)
self.maxpool = nn.MaxPool2d(kernel_size=3, stride=2, padding=1)
self.layer1 = self._make_layer(block, 64, layers[0])
self.layer2 = self._make_layer(block, 128, layers[1], stride=2)
self.layer3 = self._make_layer(block, 256, layers[2], stride=2)
self.layer4 = self._make_layer(block, 512, layers[3], stride=2)
self.avgpool = nn.AvgPool2d(7, stride=1, padding=2)
self.fc = nn.Linear(2048 * block.expansion, num_classes)
self.a = torch.nn.Parameter(torch.zeros(
self.num_classes).float().normal_(0.0, 0.1).view(-1, 1))
for m in self.modules():
if isinstance(m, nn.Conv2d):
n = m.kernel_size[0] * m.kernel_size[1] * m.out_channels
m.weight.data.normal_(0, (2. / n)**.5)
elif isinstance(m, nn.BatchNorm2d):
m.weight.data.fill_(1)
m.bias.data.zero_()
def _make_layer(self, block, planes, blocks, stride=1):
downsample = None
if stride != 1 or self.inplanes != planes * block.expansion:
downsample = nn.Sequential(
nn.Conv2d(self.inplanes, planes * block.expansion,
kernel_size=1, stride=stride, bias=False),
nn.BatchNorm2d(planes * block.expansion),
)
layers = []
layers.append(block(self.inplanes, planes, stride, downsample))
self.inplanes = planes * block.expansion
for i in range(1, blocks):
layers.append(block(self.inplanes, planes))
return nn.Sequential(*layers)
def forward(self, x):
x = self.conv1(x)
x = self.bn1(x)
x = self.relu(x)
x = self.maxpool(x)
x = self.layer1(x)
x = self.layer2(x)
x = self.layer3(x)
x = self.layer4(x)
x = self.avgpool(x)
x = x.view(x.size(0), -1)
logits = self.fc(x)
probas = torch.softmax(logits, dim=1)
predictions = ((self.num_classes-1)
* torch.sigmoid(probas.mm(self.a).view(-1)))
return logits, probas, predictions
def resnet34(num_classes, grayscale):
"""Constructs a ResNet-34 model."""
model = ResNet(block=BasicBlock,
layers=[3, 4, 6, 3],
num_classes=num_classes,
grayscale=grayscale)
return model
# +
###########################################
# Initialize Cost, Model, and Optimizer
###########################################
def cost_fn(targets, predictions):
return torch.mean((targets.float() - predictions)**2)
torch.manual_seed(RANDOM_SEED)
torch.cuda.manual_seed(RANDOM_SEED)
model = resnet34(NUM_CLASSES, GRAYSCALE)
model.to(DEVICE)
optimizer = torch.optim.Adam(model.parameters(), lr=LEARNING_RATE)
# -
# ## Training
# +
def compute_mae_and_mse(model, data_loader):
mae, mse, num_examples = torch.tensor([0.]), torch.tensor([0.]), 0
for features, targets in data_loader:
features = features.to(DEVICE)
targets = targets.float().to(DEVICE)
logits, probas, predictions = model(features)
assert len(targets.size()) == 1
assert len(predictions.size()) == 1
predicted_labels = torch.round(predictions).float()
num_examples += targets.size(0)
mae += torch.abs(predicted_labels - targets).sum()
mse += torch.sum((predicted_labels - targets)**2)
mae = mae / num_examples
mse = mse / num_examples
return mae, mse
start_time = time.time()
for epoch in range(NUM_EPOCHS):
model.train()
for batch_idx, (features, targets) in enumerate(train_loader):
features = features.to(DEVICE)
targets = targets.to(DEVICE)
# FORWARD AND BACK PROP
logits, probas, predictions = model(features)
assert len(targets.size()) == 1
assert len(predictions.size()) == 1
cost = cost_fn(targets, predictions)
optimizer.zero_grad()
cost.backward()
# UPDATE MODEL PARAMETERS
optimizer.step()
# LOGGING
if not batch_idx % 150:
s = ('Epoch: %03d/%03d | Batch %04d/%04d | Cost: %.4f'
% (epoch+1, NUM_EPOCHS, batch_idx,
len(train_dataset)//BATCH_SIZE, cost))
print(s)
s = 'Time elapsed: %.2f min' % ((time.time() - start_time)/60)
print(s)
# -
# ## Evaluation
# +
model.eval()
with torch.set_grad_enabled(False): # save memory during inference
train_mae, train_mse = compute_mae_and_mse(model, train_loader)
test_mae, test_mse = compute_mae_and_mse(model, test_loader)
s = 'MAE/RMSE: | Train: %.2f/%.2f | Test: %.2f/%.2f' % (
train_mae, torch.sqrt(train_mse), test_mae, torch.sqrt(test_mse))
print(s)
s = 'Total Training Time: %.2f min' % ((time.time() - start_time)/60)
print(s)
# -
# %watermark -iv
| pytorch_ipynb/ordinal/ordinal-cnn-beckham2016-afadlite.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Slicer 4.11
# language: python
# name: slicer-4.11
# ---
# +
# Trained tensorflow model
model_file_name = r"SagittalSpine_2019-08-14_00-18-11.h5"
models_folder_name = r"SavedModels"
# Input ultrasound sequence names
input_browser_name = r"SagittalScan"
input_image_name = r"Image_Image"
# Output will be saved using these names
output_browser_name = r"BoneSequenceBrowser"
output_sequence_name = r"SegmentationSequence"
output_image_name = r"Segmented_Image"
# Optionally save output to numpy arrays
array_output = True
array_folder_name = r"Temp"
array_segmentation_name = r"segmentation"
array_ultrasound_name = r"ultrasound"
# Image processing parameters
# Erases the side of prediction images. 1.0 means the whole prediction is erased.
# Background should be the first component (i.e. y[:,:,:,0]) in the prediction output array.
clip_side_ratio = 0.3
apply_logarithmic_transformation = True
logarithmic_transformation_decimals = 5
# +
import datetime
import numpy as np
import os
import scipy.ndimage
from keras.models import load_model
from local_vars import root_folder
# +
# Check if keras model file exists. Abort if not found. Load model otherwise.
models_path = os.path.join(root_folder, models_folder_name)
model_fullpath = os.path.join(models_path, model_file_name)
if not os.path.exists(model_fullpath):
raise Exception("Could not find model: " + model_fullpath)
print("Loading model from: " + model_fullpath)
if array_output:
array_output_fullpath = os.path.join(root_folder, array_folder_name)
array_segmentation_fullname = os.path.join(array_output_fullpath, array_segmentation_name)
array_ultrasound_fullname = os.path.join(array_output_fullpath, array_ultrasound_name)
if not os.path.exists(array_output_fullpath):
os.mkdir(array_output_fullpath)
print("Folder created: {}".format(array_output_fullpath))
print("Will save segmentation output to {}".format(array_segmentation_fullname))
print("Will save ultrasound output to {}".format(array_ultrasound_fullname))
model = load_model(model_fullpath)
# model.summary()
# +
# Check input. Abort if browser or image doesn't exist.
input_browser_node = slicer.util.getFirstNodeByName(input_browser_name, className='vtkMRMLSequenceBrowserNode')
input_image_node = slicer.util.getFirstNodeByName(input_image_name, className="vtkMRMLScalarVolumeNode")
if input_browser_node is None:
logging.error("Could not find input browser node: {}".format(input_browser_node))
raise
if input_image_node is None:
logging.error("Could not find input image node: {}".format(input_image_name))
raise
# +
# Create output image and browser for segmentation output.
output_browser_node = slicer.util.getFirstNodeByName(output_browser_name, className='vtkMRMLSequenceBrowserNode')
if output_browser_node is None:
output_browser_node = slicer.mrmlScene.AddNewNodeByClass('vtkMRMLSequenceBrowserNode', output_browser_name)
output_sequence_node = slicer.util.getFirstNodeByName(output_sequence_name, className="vtkMRMLSequenceNode")
if output_sequence_node is None:
output_sequence_node = slicer.mrmlScene.AddNewNodeByClass('vtkMRMLSequenceNode', output_sequence_name)
output_browser_node.AddSynchronizedSequenceNode(output_sequence_node)
output_image_node = slicer.util.getFirstNodeByName(output_image_name, className="vtkMRMLScalarVolumeNode")
if output_image_node is None:
volumes_logic = slicer.modules.volumes.logic()
output_image_node = volumes_logic.CloneVolume(slicer.mrmlScene, input_image_node, output_image_name)
browser_logic = slicer.modules.sequencebrowser.logic()
browser_logic.AddSynchronizedNode(output_sequence_node, output_image_node, output_browser_node)
output_browser_node.SetRecording(output_sequence_node, True)
# +
# Add all input sequences to the output browser for being able to conveniently replay everything
proxy_collection = vtk.vtkCollection()
input_browser_node.GetAllProxyNodes(proxy_collection)
for i in range(proxy_collection.GetNumberOfItems()):
proxy_node = proxy_collection.GetItemAsObject(i)
output_sequence = slicer.mrmlScene.AddNewNodeByClass('vtkMRMLSequenceNode')
browser_logic.AddSynchronizedNode(output_sequence, proxy_node, output_browser_node)
output_browser_node.SetRecording(output_sequence, True)
# +
# Iterate input sequence, compute segmentation for each frame, record output sequence.
num_items = input_browser_node.GetNumberOfItems()
n = num_items
input_browser_node.SelectFirstItem()
input_array = slicer.util.array(input_image_node.GetID())
slicer_to_model_scaling = model.layers[0].input_shape[1] / input_array.shape[1]
model_to_slicer_scaling = input_array.shape[1] / model.layers[0].input_shape[1]
print("Will segment {} images".format(n))
if array_output:
array_output_ultrasound = np.zeros((n, input_array.shape[1], input_array.shape[1]))
array_output_segmentation = np.zeros((n, input_array.shape[1], input_array.shape[1]), dtype=np.uint8)
# +
model_output_size = model.layers[-1].output_shape[1]
num_output_components = model.layers[-1].output_shape[3]
mask_model = np.ones([model_output_size, model_output_size])
mask_model_background = np.zeros([model_output_size, model_output_size])
columns_to_mask = int(model_output_size / 2 * clip_side_ratio)
print("Will mask {} columns on both sides".format(columns_to_mask))
mask_model[:,:columns_to_mask] = 0
mask_model[:,-columns_to_mask:] = 0
mask_model_background[:,:columns_to_mask] = 1
mask_model_background[:,-columns_to_mask:] = 1
# Display mask
# import matplotlib
# matplotlib.use('WXAgg')
# from matplotlib import pyplot as plt
# plt.imshow(mask_model[:,:])
# plt.show()
# +
print("Processing started at: {}".format(datetime.datetime.now().strftime('%H-%M-%S')))
for i in range(n):
# if i > 10: # todo Just for debugging
# break
input_array = slicer.util.array(input_image_node.GetID())
if array_output:
array_output_ultrasound[i, :, :] = input_array[0, :, :]
resized_input_array = scipy.ndimage.zoom(input_array[0,:,:], slicer_to_model_scaling)
resized_input_array = np.flip(resized_input_array, axis=0)
resized_input_array = resized_input_array / resized_input_array.max() # Scaling intensity to 0-1
resized_input_array = np.expand_dims(resized_input_array, axis=0)
resized_input_array = np.expand_dims(resized_input_array, axis=3)
y = model.predict(resized_input_array)
if apply_logarithmic_transformation:
e = logarithmic_transformation_decimals
y = np.log10(np.clip(y, 10**(-e), 1.0)*(10**e))/e
y[0,:,:,:] = np.flip(y[0,:,:,:], axis=0)
for component in range(1, num_output_components):
y[0,:,:,component] = y[0,:,:,component] * mask_model[:,:]
y[0,:,:,0] = np.maximum(y[0,:,:,0], mask_model_background)
upscaled_output_array = scipy.ndimage.zoom(y[0,:,:,1], model_to_slicer_scaling)
upscaled_output_array = upscaled_output_array * 255
upscaled_output_array = np.clip(upscaled_output_array, 0, 255)
if array_output:
array_output_segmentation[i, :, :] = upscaled_output_array[:, :].astype(np.uint8)
# output_array = slicer.util.array(output_image_node.GetID())
# output_array[0, :, :] = upscaled_output_array[:, :].astype(np.uint8)
slicer.util.updateVolumeFromArray(output_image_node, upscaled_output_array.astype(np.uint8)[np.newaxis, ...])
output_browser_node.SaveProxyNodesState()
input_browser_node.SelectNextItem()
slicer.app.processEvents()
# print("Processed frame {:02d} at {}".format(i, datetime.datetime.now().strftime('%H-%M-%S')))
print("Processing finished at: {}".format(datetime.datetime.now().strftime('%H-%M-%S')))
if array_output:
np.save(array_ultrasound_fullname, array_output_ultrasound)
np.save(array_segmentation_fullname, array_output_segmentation)
print("Saved {}".format(array_ultrasound_fullname))
print("Saved {}".format(array_segmentation_fullname))
# -
| Notebooks/Slicer/SequenceSegmentation2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.3 64-bit (conda)
# metadata:
# interpreter:
# hash: 1d8a5bbb0756dea561c5110081a5788c97ee82690e858540714922886ca24d79
# name: python3
# ---
# # 1A Subject Introduction
#
# ## 1A.1 Problem Solving
#
# With both typical and computational problem solving, the following steps are performed:
#
# 1. Identify the problem
# 2. Structure the problem - i.e., analyse and understand the problem
# 3. Look for possible solutions
# 4. Make a decision
# 5. Implement the solution
# 6. Monitor and/or seek feedback
#
# Do note that each step should be revisited as greater insights into the problem are gleaned.
#
# Computational thinking primarily encompasses several primary concepts, which suggests how we should address problem solving from a computational perspective:
#
# <center><img src="https://www.wcpss.net/cms/lib/NC01911451/Centricity/Domain/17003/Computational%20Thinking.PNG" alt='https://www.wcpss.net/cms/lib/NC01911451/Centricity/Domain/17003/Computational%20Thinking.PNG' style='width:40%; height:auto'></center>
#
#
# ## 1B.1 Programs and Programming Languages
#
# >A **program** is the actual expression of an algorithm in a specific programming language. It allows the computer to execute the problem solution through a sequence of instructions.
#
# Essentially, an algorithm is generally produced in the design phase of a problem solution, while a program that expresses the algorithm is produced in the programming stage. A program is also made up of lines of code that can be categorized into either an **expression** or a **statement**.
#
# >An **expression** is a syntactic entity in a programming language that may be evaluated to determine its value.
#
# >A **statement**, on the other hand, is a syntactic entity, which has no value (merely an instruction.)
#
# Example:
# >```python
# >answers = 1+1: # '1+1' is an expression
# >if answers == 2:
# > print('Good Outlook') # print('Good Outlook') is a statement.
# >```
#
# There are typically 3 basic statements:
#
# 1. Input statements
# 2. Output statements
# 3. Assignment statements
#
# In general, input and output (I/O) allows the communication between an information processing system (such as a computer) and an external entity (such as a human).
#
# Input is information supplied to a computer or program.
#
# Output is information provided by a computer or program.
#
# Assignment statements will be elaborated further in the later section.
#
# >```python
# >x = input("Enter a positive integer value for x: ") #example of input statement. Note that the user input is assigned to the variable x
# >print(x) #print statement to display the output
# >```
#
# The most basic statement, and typically the first code you write when learning a programming language, is the output statement, which outputs certain data - e.g., printing `“hello world”`. There is typically also an input statement that will request input from the user.
#
# ### 1B.1.1 `print()` function
#
# The `print(YOUR_SPECIFIED_MESSAGE)` function prints `YOUR_SPECIFIED_MESSAGE` to the screen, or other standard output device.
#
# > It's demo time.
#
# ### 1B.1.2 `input()` function
#
# The `input(YOUR_MESSAGE_HERE)` function allows user input, where `YOUR_MESSAGE_HERE` is a `str` object. Variable assigned is also of `str` type.
#
# > It's demo time. Talk about type casting.
#
# ### 1B.1.3 Formatting output with `.format()` method or `f-string`
#
# > It's demo time.
a = int(input("Enter a value for a:"))
print( f"My marks are {a+b} and {b**2}")
# ## Exercise 1
# What is the output of the following:
#
# 1. `print(5 == 5.0)`
# 2. `print(float(1/2))`
# 3. `print(float(1)/2)`
# 4. `print(5 == "5")`
# 5. `print("sdf" != "sdf")`
# 6. `print(True and (False or not True))`
# 7. `print(str(53) + str(True))`
# 8. code
# >```python
# >a = 20
# >print(str(15 - (a - 15)) + ", ", end = " ")
# >a = 10
# >print(15 - (a - 15))
# >```
# ## Exercise 2
# The goal of this programming exercise is simply to get you more comfortable with using IDLE, and to begin using simple elements of Python. Standard elements of a program include the ability to print out results (using the `print()` function), the ability to read input from a user at the console (e.g., using the `input()` function), and the ability to store values in a variable, so that the program can access that value as needed.
#
# The problem:
#
# - Ask the user to enter his/her last name.
# - Ask the user to enter his/her first name.
# - Print out the users last and first names in that order.
#
# Example interaction between the program and the user. (note: words on the 2nd and 4th line are from the computer, based on the program, while the words on 3rd and 5th line ('Daren', 'Ler' respectively) are a user's input):
# >```python
# >>>>
# >Enter your last name:
# >Daren
# >Enter your first name:
# >Ler
# >Ler #output
# >Daren #output
# >>>>
# # 1C Data Types
#
# >In computer science, a data is defined to be a sequence of one or more symbols.
#
# Note that under this definition, data doesn't need to carry information or even be meaningful. However, to make use of data, we need to endow it with more structure and make it meaningful.
#
# ## 1C.1 Basic Data Types
#
# Data can be categorised into different types, i.e. a data type is a category/class of data. The following are 5 basic data types which are found in most programming languages:
#
# | No | Data Type | Definition | Examples | Benefits | Limitations |
# |---|-----------------------|-------------------------------------------------------------------|----------------------------|-----------------------------------------------------------------|----------------------------------------------------|
# | 1 | Integer `int` | A whole number or number without a fractional part | `-1`,`0`,`1`,`1000` | Full precision with finite digits | Typically limited to a specific range |
# | 2 | Floating Point Number `float`| A number with ( or calculated with)a fractional part | `-1.5`,`0.1`,$\tt{\frac{1}{3}}$,`2.5`,$\pi$ | Stores rational/irrational numbers with reasonable accuracy | Not exact; not 100% precise |
# | 3 | Boolean `Boolean` | Two values representing either true or false in a logic statement | `True`, `False` | Space efficient when needed to represent values that are binary | Only 2 possible values |
# | 4 | String `str` | A collection of symbols | `a`,`abc`,`123`,`a string` | Allows for more human-readable information | Mapped values cannot be manipulated arithmetically |
# | 5 | `None` | A null variable or object | | | |
#
# We may also manipulate these values by performing various operations on them. Essentially, when we write code, we may form expressions via the use of operators to manipulate data.
#
# ## 1C.2 Operators
#
# >An operator is a symbol that *operates* on one or more values, i.e. it is a symbol that represents an action or process.
#
# ### 1C.2.1 Arithmetic Operations
#
# An arithmetic operator is an operator that work on numeric data types. The typical operations that may be performed on numbers include:
#
# | Operation | Symbol |
# |:----------------:|:------:|
# | Addition | `+` |
# | Subtraction | `-` |
# | Multiplication | `*` |
# | Division | `/` |
# | Integer Division | `//` |
# | Modulo | `%` |
# | Power | `**` |
#
# >It's demo time. Try the operations with the following pairs `(15,10)`, `(3,4)`, `(124,20)`
# ## Exercise 3
#
# It is important to know the type of the value that a variable refers to – e.g., this would allow us to use the correct operators. Python automatically infers the type from the value you assign to a variable. Write down the type of the values stored in each of the variables below. Pay special attention to punctuation: values are not always the type they seem!
#
# 1. `a = False`
# 2. `b = 3.7`
# 3. `c = 'Alex'`
# 4. `d = 7`
# 5. `e = 'True'`
# 6. `f = 12 ** 3`
# 7. `g = '17'`
# 8. `h = True`
# 9. `i = '3.14159'`
# 10. `j = 12 / 27`
# 11. `k = 2.0 / 1`
# 12. `l = (5 == "5")`
# 13. `m = str((-4 + abs(-5) / 2 ** 3) + 321 - ((64 / 16) % 4) ** 2)`
#
# To verify your answers, you can use the `type` function in interactive Python shell (as shown below). However, first try to do the exercise without the help of the shell.
#
# >```python
# >>>> x = 100
# >>>> type(x)
# ><type 'int'>
# >>>>
# >```
# ### 1C.2.2 Logical and Comparison Operations
#
# There are several Boolean or logical operations. Among the most common are the following:
#
# | Operation | Symbol |
# |:-------------:|:--------:|
# | Logical AND | `and` |
# | Logical OR | `or` |
# | Logical NOT | `not` |
#
# Each logical operation is associated with a truth table, which defines all possible pairs of operand values, and the corresponding resultant value that is attained when the operator in question is applied to the given operands. The truth table for the above logical operations are as follows.
#
# | `x` | `y` | `x and y` | `x or y` | `not x` |
# |-------|-------|---------|--------|-------|
# | `True` | `True` | `True` | `True` | `False` |
# | `True` | `False` | `False` | `True` | `False` |
# | `False` | `True` | `False` | `True` | `True` |
# | `False` | `False` | `False` | `False` | `True` |
#
# Note that:
# > 1. The result is `True` if both `x` and `y` are `True`, or else, the result is `False`.
# > 2. The result is `True` if `x` is `True` or `y` is `True`, or else, the result is `False`.
# > 3. The result is the opposite of the Boolean value of `x`.
#
# Additionally, there are several comparison operations that do not require Boolean operands, but have a Boolean value result:
#
# | Operation | Symbol |
# |:----------------:|:------:|
# | Less Than | `<` |
# | Less Than or Equals | `<=` |
# | Equality | `==` |
# | Greater Than | `>` |
# | Greater Than or Equals | `>=` |
# | Not Equals | `!=` |
#
# As with the arithmetic and logical operations, the above comparison operations take 2 operands, evaluate the associated test, and then have Boolean result. For example, `10 < 5` will result in `False`, since 10 is actually greater than 5, not less; `“abc” == “cba”` will result in False, since the 2 strings are not equivalent.
#
# ## Exercise 4
#
# Boolean operators can seem tricky at first, and it takes practice to evaluate them correctly. Write the value (`True` or `False`) produced by each expression below, using the assigned values of the variables `a`, `b`, and `c`. Try to do this without using your interpreter, but you should check yourself when you think you've got it.
#
# >Hint: Work from the inside out, starting with the innermost expressions, similar to arithmetic.
#
# Let:
#
# - `a = False`
# - `b = True`
# - `c = False`
#
# Would the expressions below evaluate to `True` or `False`?
#
# 1. `b and c`
# 2. `b or c`
# 3. `not a and b`
# 4. `(a and b) or not c`
# 5. `not b and not (a or c)`
# ### 1C.2.2 String Operations
#
# The most basic string operation is concatenation `+`, which simply combines the contents of 2 strings. For example, given 2 strings, `“abc”` and `“xyz”`, the concatenation of `“abc”` and `“xyz”` would thus be the string `“abcxyz”`.
#
# Note that strings are typically denoted via the open and closed inverted commas - i.e., the value of the string corresponds to the symbols in between the open and closed inverted commas.
#
# You can also repeat strings by using `*` operator.
#
# ### 1C.2.3 Expressions and Operation Precedence
#
# As mentioned in Section 2.1., we utilise operations within expressions.
#
# Expressions have the general form:
#
# `<OPERAND> <OPERATOR> <OPERAND>`
#
# For example, in the expression `10 + 20`, `10` corresponds to the first operand, `20` to the second operand, and `+` is the operator (i.e., the operation being applied to the 2 operands).
#
# All operators require certain operand types to work properly:
#
# | Operator Type | Acceptable Operand Types |
# |:----------------:|:------:|
# | Arithmetic | `int`,`float` |
# | Logical | `Boolean` |
# | Comparison | Any (Result is `Boolean`) |
# | String | `str` |
#
#
# Expressions may also be nested (just as mathematical expressions are). For example, `10 + ( 20 * 30 )`. When expressions are nested, they are executed using a certain order of precedence. Going back to the example above, we know that `20 * 30` must be evaluated first (to `600`), before we evaluate the resultant `10 + 600` expression.
#
# This precedence is typically consistent across all programming languages, the higher the operator is in the list below, the higher its precedence, i.e. it will get evaluated first.
#
# | Operator Type | Operation |
# |:----------------:|:------:|
# | Exponentiation | `**` |
# | Multiplication and Division | `*`,`/`,`//`,`%` |
# | Addition and Subtraction | `+`,`-` |
# | All Comparison | `<`,`<=`,`>`,`>=`,`==`,`!=` |
# | Logical NOT | `not` |
# | Logical AND | `and` |
# | Logical OR | `or` |
#
# However, to enforce precedence, we may typically utilise brackets, just as we do in mathematics. Thus, for example, despite the precedence table shown above, when `(10 + 20) * 30` is evaluated, we now instead evaluate `10 + 20` first (since it is bracketed). As with mathematics, the innermost brackets must be evaluated first.
# # 1D Programming Constructs
#
# >A programming construct is an abstract way of describing a concept in terms of the syntax of programming language. These constructs are building blocks that forms the basis of all programs.
#
# These are basic programming constructs that we need to know:
#
# - Sequence,
# - Assignment,
# - Selection,
# - Iteration.
#
# We will explore the Assignment and Selection Construct in the sections below.
#
# ## 1D.0 Programming Construct : Sequence
#
# The most general construct of programming corresponds to the sequence of instructions, or more specifically, the sequence of statements that are specified within the program code.
#
# ## 1D.1 Programming Construct : Assignment
#
# In order to process data, we typically need to store it in computer memory. This is done via the use of variables. You might be familiar with variables given their use in mathematics. However, variables in mathematics and computing are different.
#
# In computing, a variable corresponds to an identifier that references a unit of data in computer memory.
#
# In order to define a variable, we must utilise the first programming construct: an **assignment statement**.
#
# To understand how assignment and variables work, let us refer to the following example:
#
# >```python
# >#x is the variable, 100 is the data, ← is the assignment operator
# >#and the whole line below is the assignment statement
# >x ← 100
# >```
#
# The above is a typical assignment statement. We are specifying a variable, i.e., an identifier `x`, which will reference some part of the computer’s memory, and in that segment of computer memory, store the integer value `100`.
#
# Essentially, the assignment statement does 3 things:
#
# 1. Stores the variable identifier in an identifier table
# 2. Allocates computer memory for the storage of the data type value specified; the memory allocated will correspond to some specific location in memory - i.e., a memory address
# 3. Links the variable identifier to the computer memory location
#
# In this manner, whenever the variable is used, the computer knows that we are referring to the value that is stored in the associated location in computer memory.
#
# Assignment also allows programmers to:
#
# 1. Assign new values to a variable
# 2. Copy variable values
# 3. Swap values between variables
#
# It should also be noted that when assigning a value, that value may be computed using expressions consisting of arithmetic, logical, comparison and/or string operations. For example:
#
# >```python
# >x ← 10+20
# >```
#
# Such expressions can, and or course, often do include the specification of other variables. For example:
#
# >```python
# >result ← a*x**2 + b*x + c
# >```
#
# In Python, the assignment operator is `=` , which should not be confused with the equal sign $=$ from mathematics.
#
# In Python, we can do multivariable assignment in a single line too. Example:
# >```python
# > a,b = 10, 20
# > a,b = b, a
# >```
#
# ### 1D.1.1 Python Naming Conventions
#
# To prevent conflict and to keep consistency between programs, programming languages normally has some naming conventions for the variables, please check [https://www.python.org/dev/peps/pep-0008/#naming-conventions](https://www.python.org/dev/peps/pep-0008/#naming-conventions) for updated style guide for python. For example, never use the characters `l` (lowercase letter el), `O` (uppercase letter oh), or `I` (uppercase letter eye) as single character variable names. In some fonts, these characters are indistinguishable from the numerals one and zero. When tempted to use `l`, use `L` instead.
#
# Legal variable names of a variable in Python :
# - Cannot begin with a digit
# - Cannot include operator symbols
# - Cannot be reserved words (e.g., `or`, `and`, `not`, `in`, `is`, `def`, `return`, `pass`, `break`, `continue`)
# - Should not be built-in function names (e.g., `print`, `input`, `range`, `len`, `min`, `max`, `int`, `str`)
#
# ## Exercise 5
#
# What is the output of the following code?
#
# >```python
# >a = 5
# >b = a + 7
# >a = 10
# >print(str(b))
# >```
# ## Exercise 6
#
# The Python interpreter has strict rules for variable names. Which of the following are legal Python variable names? If the name is not legal, state the reason.
#
# 1. `and`
# 2. `_and`
# 3. `Var`
# 4. `var1`
# 5. `1var`
# 6. `my-name`
# 7. `your_name`
# 8. `COLOUR`
# ### Programming Construct :Selection (Conditional Branching)
# <ul>
# <li> Comparision operators: ==, !=, >,>=,<,<= </li>
# <li> is, in </li>
# <li> evaluates to a Boolean value</li>
# <li> if .. elif .. else ..</li>
# </ul>
# ## 1D.2 Programming Construct : Selection
# Often, a program consisting of a sequence of statements is insufficient. This is because it is common to face a problem that requires the execution of different sets of statements dependent upon the current state of the data.
#
# To remedy this, we utilise the selection programming construct.
#
# With the selection programming construct, a query/condition is evaluated. Depending on its result, the program will take one of two possible courses of action. As such, the selection construct is typically referred to as an `if-else` statement (or `if` statement for short), which selects different sets of statements to execute.
#
# For example:
#
# >```coffeescript
# >INPUT x
# >IF x ≥ 50
# > THEN
# > OUTPUT “PASS”
# > OUTPUT “Well done!”
# > ELSE
# > OUTPUT “FAIL”
# > OUTPUT “Please try harder.”
# >ENDIF
# >```
#
# It should be noted that it is common practice to indent the statements within the `if` or `else` parts of an `if` statement so that it is easier to read the code.
#
# Finally, it should also be noted that `if` statements may be nested, i.e., the statements in the `if` or `else` parts may themselves include another `if` statement.
#
#
if <condition>:
pass
pass
elif <condition>:
pass
pass
else:
pass
# ## Example 7
#
# Convert marks to grade: > 70 -> A, 60-69 -> B, 55-59 -> C , < 55 -> D
mark = int(input("Enter mark:"))
if mark >= 70:
print("A")
elif mark >=60:
print("B")
elif mark >= 55:
print("C")
else:
print("Fail")
# -----
# ## Exercise 8
# Show that for any integer $n$, $n^5$ and $n$ always have the same last digit.
# -----
# ## Exercise 9
#
# Consider the following code.
#
# >```python
# >x = input("Enter a positive integer value for x: ")
# >
# >if x > 10 and x % 6 == 3:
# > print("a", end = "")
# >elif x > 10 and x < 20:
# > print("b", end = "")
# >else:
# > print("c", end = "")
# >print("")
# >```
#
# Given the following outputs, determine the corresponding value(s) for `x` that would have generated that output. If no values of a can produce the listed output, write none.
#
# 1. `ab`
# 2. `a`
# 3. `b`
# 4. `c`
# -----
# # 2A Programming Constructs : Iteration
#
# The last programming construct is the iteration construct, which allows us to specify loops.
#
# Often, we will notice that our code requires the execution of similar repeated steps.
#
# The iteration, or loop construct allows us to automate repeated steps. More specifically, iteration is the act of repeating a process to achieve a specific end goal. When iteration is used, the sequence/set of instructions within the loop will keep repeating until a certain condition/requirement has been reached, or is no longer satisfied. Each repetition is also known as an iteration. The results of one iteration are used as the starting point for the next iteration.
#
# Iteration usually involves the use of loops. Most commonly used loops include:
#
# 1. **For** loops : Loops with an explicit counter for every iteration. This allows the body of the `for` loop code to know about the sequencing (i.e. index number) of the iteration.
# 2. **While-Do** loops : Loops which continually execute a statement while a particular expression (of a condition/set of conditions) is true. It is executed zero or more times.
# 3. **Do-While** (or Repeat-Until) loops : Similar to while-do loops, except the expression is evaluated at the end. Hence, the block of code is executed at once, then only repeats when the specified condition or set of conditions is met.
# ### Programming Construct: Iterations
# <ul>
# <li> while </li>
# <li> for.. in range(start,stop,step) -> start,start+step,..stop-1</li>
# </ul>
# ## Example 10
# Convert marks to grade repeatedly
in_str = input("Enter mark:")
while in_str.isdigit():
if mark >= 70:
print("A")
elif mark >=60:
print("B")
elif mark >= 55:
print("C")
else:
print("Fail")
in_str = input("Enter mark:")
for i in "abcdefg":
print(i)
# +
## Problem 2: Extract digits from a number, 123 -> 3, 2, 1
# -
num = int(input("Enter a number:"))
while num != 0:
print ( num % 10)
num = num //10
# +
## Problem 3: find all the factors for a +ve integer n
for i in range(1, n+1):
if n % i == 0:
print(i)
# -
# ## Exercise 11
#
# What is the output for each of the following pieces of code?
#
# Note: if the code does not terminate, write infinite loop.
#
# 1. Code 1
#
# >```python
# >a = 5
# >while a < 8:
# > print("X", end = "")
# >print("")
# >```
#
# 2. Code 2
#
# >```python
# >a = -1
# >while a < 3:
# > print("X" , end = "")
# >a = a + 1
# >print("")
# >```
#
# 3. Code 3
#
# >```python
# >a = 1
# >while a % 7 != 0:
# > if a % 2 == 0:
# > print("O" , end = "")
# > if a == 2:
# > print("X" , end = "")
# > a = a + 1
# >print("")
# >```
# ## Exercise 12
#
# What is the output for each of the following pieces of code?
#
# 1. Code 1
#
# >```python
# >num = 10
# >while num > 3:
# > print(num)
# > num = num – 1
# >```
#
# 2. Code 2
#
# >```python
# >divisor = 2
# >for i in range(0, 10, 2):
# > print(i // divisor)
# >```
#
# 3. Code 3
#
# >```python
# >num = 10
# >while True:
# > if num < 7:
# > break
# > print(num)
# > num -= 1
# >```
#
# 4. Code 4
#
# >```python
# >count = 0
# >for letter in "Snow!":
# > print("Letter #" + str(count), " is ", letter)
# > count += 1
# >```
# ## Exercise 13
#
# What is the output for each of the following pieces of code?
#
# 1. Code 1
#
# >```python
# >keep_going = True
# >a = 0
# >b = 0
# >while keep_going:
# > print("O", end = "")
# > a = a + 5
# > b = b + 7
# > if a + b >= 24:
# > keep_going = False
# >print("")
# >```
#
# 2. Code 2
#
# >```python
# >keep_going = True
# >a = 0
# >b = 0
# >while keep_going:
# > print("O" , end = "")
# > if a + b >= 24:
# > keep_going = False
# > a = a + 5
# > b = b + 7
# >print("")
# >```
#
# 3. Code 3
#
# >```python
# >keep_going = True
# >a = 0
# >b = 0
# >while keep_going:
# > print("O" , end = "")
# > a = a + 5
# > b = b + 7
# > if a + b > 24: # note that ">" is used here … vs ">=" in (i)
# > keep_going = False
# >print("")
# >```
#
# 4. Code 4
#
# >```python
# >keep_going = True
# >a = 0
# >b = 0
# >while keep_going:
# > print("O" , end = "")
# > if a + b > 24:
# > keep_going = False # note that ">" is used here … vs ">=" in (ii)
# > a = a + 5
# > b = b + 7
# >print("")
# >```
# ## Exercise 14
#
# What is the output for each of the following pieces of code? Note: if the code does not terminate, write infinite loop.
#
# 1. Code 1
#
# >```python
# >a = 0
# >while a < 3:
# > while True:
# > print("X", end = "")
# > break
# > print("O", end = "")
# > a = a + 1
# >print("")
# >```
#
# 2. Code 2
#
# >```python
# >a = 1
# >while a < 3:
# > while a < 3:
# > print("O", end = "")
# > a = a + 1
# >print("")
# >```
#
# 3. Code 3
#
# >```python
# >a = 1
# >while a < 3:
# > if a % 2 == 0:
# > b = 1
# > while b < 3:
# > print("X", end = "")
# > b = b + 1
# > print("O", end = "")
# > a = a + 1
# >print("")
# >```
# ## Exercise 15
#
# The following code loops infinitely when run, fix the code so that its output is `OOOXOXOO`.
#
# >```python
# >a = 1
# >while a < 3:
# > b = 1
# > while b < 3:
# > if a == 2:
# > print("X", end = "")
# > print "O",
# > b = b + 1
# > print("O", end = "")
# >print("")
# >```
# ## Exercise 16
#
# <NAME> (1777-1855) was one of those remarkable infant prodigies whose natural aptitude for mathematics soon becomes apparent. As a child of three, according to a well-authenticated story, he corrected an error in his father's payroll calculations. His arithmetical powers so overwhelmed his schoolmasters that, by the time Gauss was 10 years old, they admitted that there was nothing more they could teach the boy. It is said that in his first arithmetic class Gauss astonished his teacher by instantly solving what was intended to be a "busy work" problem: Find the sum of all the numbers from 1 to 100.
#
# How would you find this sum yourself? No Googling for answer kthx.
# # 3B Basic Data Structures
#
# In computer science, a data structure is a data organization, management, and storage format that enables efficient access and modification. More precisely, a data structure is a collection of data values, the relationships among them, and the functions or operations that can be applied to the data.
#
# ### 3B.1.1.1 Lists and Its Manipulation
#
# `list` is a Python data type that
#
# - stores data in order it was added to it,
# - is **mutable**, meaning the values can change.
#
# Lists are created by using square brackets `[]`. For example
#
# >```python
# >mylist=['Mathematics','Computing','General Paper']
# >```
#
# - Use `mylist[n-1]` to access the $n$th element in the list `mylist`. Note that index is 0-based
# - Sometimes, you want a running sublist of the list instead. `mylist[i:k]` will the list that contains the $i$th element of `mylist` to the $k$th element. This is called **slicing** and more generally, `mylist[i:k:step]`
# - `mylist.append(myelement)` will add the element `myelement` into the list `mylist`.
# - to concatenate lists `mylist` and `myanotherlist` to a list, we use the `+` operator. So the combined list is `mylist + myanotherlist`. Alternatively, you can also use the `.extend()` method, i.e., `mylist.extend(myanotherlist)` to get the same result. Also, `*`
# - to locate the position of the element `myelement` in the list `mylist`, we can use `.index()` method for lists. i.e., `mylist.index(myelement)`
# - to remove the element `myelement`, which has the index `myindex`, from the list `mylist`, we use `.pop()` method. The syntax is `mylist.pop(myindex)`
# - we can also use `for` loops to iterate over a list.
# - elements in the list can be arranged into numerical or alphabetical order via the `sorted()` function. We can also pass in the argument `reverse=True` inside the function to get a descending order instead.
# - `.count()`, `.remove()`, `.insert()`
L=[1,"two",3.0, 4, 5]
# ### Collection Types: String, List
# <ul>
# <li> index, zero-based </li>
# <li> slicing, [start:stop:step]</li>
# <li> concatenate , + , *n</li>
# <li> for .. in iterable </li>
# <li> for .. in range(len(L)) </li>
# </ul>
# ## Exercise
#
# - Create a list called `my_subjects` with the subjects '`Mathematics'`, `'General Paper'`, `'Further Mathematics'` and `'Computing'`.
# - Use the `.extend()` method on `my_subjects` to add your 2 other subjects that you are offering in A-Level and print the list `my_subjects`.
# - Use the `.index()` method to find the position of `'Further Mathematics'` in the list. Save the result as `my_index`.
# - Use the `.pop()` method with `my_index` to remove `'Further Mathematics'` from the list.
# - Print the `my_subjects` list again.
#
# >```python
# ># Create a list containing the subjects: my_subjects
# >my_subjects = ---
# >
# ># Extend my_subjects with your other subjects
# >---
# >
# ># Print subject names
# >print(my_subjects)
# >
# ># Find the index of 'Further Mathematics7' in the list
# >my_index = ---
# >
# ># Remove 'Further Mathematics' from my_subjects
# >---
# >
# ># Print subject names
# >print(my_subjects)
# >```
# ### List methods
# <ul>
# <li> creating list</li>
# <li> .append(object) </li>
# <li> .index(object) </li>
# <li> .count(object)</li>
# <li> .pop(index)</li>
# <li> .remove(object)</li>
# <li> .insert(index,object)</li>
#
# </ul>
L=[]
L=list()
# ### Dictionary
# <ul>
# <li> collection of key,value pairs</li>
# </ul>
# ## Additional Practice Questions
# **[Introduction to Electrical Engineering and Computer Science I: Types, Values, Expressions; Variables and Binding](https://ocw.mit.edu/courses/electrical-engineering-and-computer-science/6-01sc-introduction-to-electrical-engineering-and-computer-science-i-spring-2011/python-tutorial/part-i/)**<br>
# **[Introduction to Electrical Engineering and Computer Science I: Using if, else, and while](https://ocw.mit.edu/courses/electrical-engineering-and-computer-science/6-01sc-introduction-to-electrical-engineering-and-computer-science-i-spring-2011/python-tutorial/part-3/)**
# +
import matplotlib.pyplot as plt
import csv
X = []
Y = []
with open('60 mm.txt', 'r') as datafile:
ploting = csv.reader(datafile, delimiter=',')
for ROWS in ploting:
print(ROWS)
X.append(float(ROWS[0]))
Y.append(float(ROWS[1]))
plt.plot(X, Y)
plt.title('Transmission spectrum')
plt.xlabel('Wavelength(nm)')
plt.ylabel('Transmission')
plt.show()
# -
| Working Folder/Fundamentals1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:tflearn]
# language: python
# name: conda-env-tflearn-py
# ---
# <h1>Lists</h1>
# <li>Sequential, Ordered Collection
#
# <h2>Creating lists</h2>
x = [4,2,6,3] #Create a list with values
y = list() # Create an empty list
y = [] #Create an empty list
print(x)
print(y)
# <h3>Adding items to a list</h3>
#
x=list()
print(x)
x.append('One') #Adds 'One' to the back of the empty list
print(x)
x.append('Two') #Adds 'Two' to the back of the list ['One']
print(x)
x.insert(0,'Half') #Inserts 'Half' at location 0. Items will shift to make roomw
print(x)
x=list()
x.extend([1,2,3]) #Unpacks the list and adds each item to the back of the list
print(x)
x.extend([4, 5])
print(x)
x.append([7,8])
print(x)
# <h3>Indexing and slicing</h3>
x=[1,7,2,5,3,5,67,32]
print(len(x))
print(x[3])
print(x[2:5])
print(x[-1])
print(x[::-1])
# <h3>Removing items from a list</h3>
x=[1,7,2,5,3,5,67,32]
x.pop() #Removes the last element from a list
print(x)
x.pop(3) #Removes element at item 3 from a list
print(x)
x.remove(7) #Removes the first 7 from the list
print(x)
# <h3>Anything you want to remove must be in the list or the location must be inside the list</h3>
x.remove(20)
# <h2>Mutablility of lists</h2>
y=['a','b']
x = [1,y,3]
print(x)
print(y)
y[1] = 4
print(y)
print(x)
x="Hello"
print(x,id(x))
x+=" You!"
print(x,id(x)) #x is not the same object it was
y=["Hello"]
print(y,id(y))
y+=["You!"]
print(y,id(y)) #y is still the same object. Lists are mutable. Strings are immutable
# +
def eggs(item,total=0):
total+=item
return total
def spam(elem,some_list=[]):
some_list.append(elem)
print(id(some_list))
return some_list
# +
print(eggs(1))
print(eggs(2))
print(spam(1))
print(spam(2))
# -
# <h1>Iteration</h1>
# <h2>Range iteration</h2>
#The for loop creates a new variable (e.g., index below)
#range(len(x)) generates values from 0 to len(x)
x=[1,7,2,5,3,5,67,32]
for index in range(len(x)):
print(x[index])
list(range(len(x)))
# generator 好处: 节省内存. 如果需要创建一个1billion 的数组, 可以使用这个来创建计数器.
# <h3>List element iteration</h3>
x=[1,7,2,5,3,5,67,32]
for element in x: #The for draws elements - sequentially - from the list x and uses the variable "element" to store values
print(element)
# <h3>Practice problem</h3>
# Write a function search_list that searches a list of tuple pairs and returns the value associated with the first element of the pair
def search_list(list_of_tuples,value):
#Write the function here
result = 0
for item in list_of_tuples:
if item[0] == value:
return item[1]
return result
prices = [('AAPL',96.43),('IONS',39.28),('GS',159.53)]
ticker = 'IONS'
print(search_list(prices,ticker))
ticker = 'ION'
print(search_list(prices,ticker))
# <h1>Dictionaries</h1>
mktcaps = {'AAPL':538.7,'GOOG':68.7,'IONS':4.6}
mktcaps['AAPL'] #Returns the value associated with the key "AAPL"
mktcaps['GS'] #Error because GS is not in mktcaps
mktcaps.get('GS') #Returns None because GS is not in mktcaps
mktcaps['GS'] = 88.65 #Adds GS to the dictionary
print(mktcaps)
del(mktcaps['GOOG']) #Removes GOOG from mktcaps
print(mktcaps)
mktcaps.keys() #Returns all the keys
mktcaps.values() #Returns all the values
sorted(mktcaps.keys())
sorted(mktcaps.values())
print(mktcaps)
test = {'x':1, 'x':2, 'y':3}
test['x']
for item in test:
print(item[0])
len(test)
# <h1>Sets</h1>
tickers = {"AAPL", "GE", "NFLX", "IONS"}
regions = {"North East", "South", "West coast", "Mid-West"}
"AAPL" in tickers
"IBK" in tickers
pharma_tickers = {"IONS", "IMCL"}
tickers.isdisjoint(pharma_tickers) # empty intersection 是否0交集
tickers & pharma_tickers # intersection 交集
tickers | pharma_tickers # union 并集
tickers - pharma_tickers # set diference 不同
tickers > pharma_tickers # superset 是否为父集合
pharma_tickers <= tickers # subset 是否为子集合
pharma_tickers < tickers # proper-subset 是否为子集合
for item in tickers:
print(item)
s = {1, 2, 4, 3}
s[3] - 4
dict1 = {"john":40, "peter":45}
dict2 = {"john":466, "peter":45}
dict1 > dict2
del dict1["john"]
dict1
| BAMM.101x/Collections.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Querying Preprint Metrics!
#
# This notebook will contain examples of how to use the preprint date analytics endpoints developed for the Sloan grant.
# # Main API Queries
#
# First, we'll focus on making queries to the main views and downloads endpoints. The request is controlled by adding various query parameters at the end of the URL. To add the first query param, add a "?" to the end of the url, followed by the param name, an "=", and the value. So to look for downloads metrics for the preprint with the guid `abcde`, your URL would look like:
#
# `/_/metrics/preprints/downloads/?guids=abcde`
#
# To add a second query param to the URL, add a "&" and follow the same pattern. To then look for metrics for this guid on the date `2019-01-01`, your URL would look like:
#
# `/_/metrics/preprints/downloads/?guids=abcde&on_date=2019-01-01`
#
# To search for results for a list of guids, seperate them with a ",":
#
# `/_/metrics/preprints/downloads/?guids=abcde,efghi,jklmn&on_date=2019-01-01`
#
# Query params:
#
# - `guids`: The guids, seperated by commas, to request metrics for
#
# - `on_date`: metrics for this specific day. If you include an on_date, you cannot include other date parameters. Must be in the format of `YYYY-MM-DD`
#
# - `start_datetime`: restrict the results to starting on this datetime. Can either be in the format `YYYY-MM-DD` and results will start at midnight of that day, or `YYYY-MM-DDThh:mm` where h and m are the hour and minute. If you provide a start datetime with no end datetime, the end datetime will default to a full day ago at 11:59pm UTC. If you provide a start or end datetime including minutes, the other value must also include minutes.
#
# - `end_datetime`: restrict the results to ending on this date. You cannot provide an `end_datetime` with no start datetime. See formatting rules for `start_datetime` for more specifics.
#
# - `interval`: how fine grained you'd like the results you'd like to be returned to you. To see what you can enter in this section, check out [the elasticsearch docs on intervals](https://www.elastic.co/guide/en/elasticsearch/reference/current/_intervals.html)
#
#
# If no time period is specified, metrics will be returned for the previous 5 full days - 6 days ago at midnight UTC to one day ago at 11:59pm UTC.
#
# ## Notes
#
# * There is no verification server-side that any of the guids are *actually* preprints or, indeed, exist. Any guids that are requested that have no data will not return data.
# * The api calls start from `/_/` rather than `/v2/` because this is a private endpoint
# * The example on withdrawn preprints below doesn't have any data because it would take me a while to figure out how to withdraw a preprint, but the elastic index doesn't know anything about the current status of a preprint; if there was an access, then there will be an entry.
# # Query Examples
#
# In the following cells, replace the variables with ones you'd like to use.
# +
# python setup
import os
import json
import requests
# +
# Change me! Uncomment and/or change the variables you'd like to use
METRICS_BASE = 'http://localhost:8000/_/metrics/preprints/'
# METRICS_BASE = 'https://api.osf.io/v2/metrics/preprints/'
TOKEN = os.environ['LOCAL_OSF_TOKEN']
# TOKEN = os.environ['LOCAL_HENRIQUE_OSF_TOKEN']
# LOCAL_TOKEN = '<PASSWORD>'
headers = {
'Content-Type': 'application/vnd.api+json',
'Authorization': 'Bearer {}'.format(TOKEN)
}
SINGLE_PREPRINT_GUID = 'jpftg'
WITHDRAWN_PREPRINT_GUID = 'hg89q'
LIST_OF_GUIDS = ['jpftg', 'f8ph9', 'xr4jv', 'mdpcb']
# +
# get download results for one preprint from the past 5 full days
url = '{}downloads/?guids={}'.format(METRICS_BASE, SINGLE_PREPRINT_GUID)
res = requests.get(url, headers=headers)
print('Request URL was: {}'.format(url))
print(json.dumps(res.json(), indent=4))
# +
# get view results for one preprint from January 3, 2019
url = '{}views/?guids={}&on_date=2019-01-03'.format(METRICS_BASE, SINGLE_PREPRINT_GUID)
res = requests.get(url, headers=headers)
print('Request URL was: {}'.format(url))
print(json.dumps(res.json(), indent=4))
# -
','.join(LIST_OF_GUIDS)
# +
# get view results for a list of preprints from January 2, 2018
url = '{}views/?guids={}&on_date=2018-01-02'.format(
METRICS_BASE,
','.join(LIST_OF_GUIDS)
)
res = requests.get(url, headers=headers)
print('Request URL was: {}'.format(url))
print(json.dumps(res.json(), indent=4))
# +
# get view results from January 3, 2019 to today for a list of guids
url = '{}views/?guids={}&start_datetime=2019-01-03'.format(
METRICS_BASE,
','.join(LIST_OF_GUIDS)
)
res = requests.get(url, headers=headers)
print('Request URL was: {}'.format(url))
print(json.dumps(res.json(), indent=4))
# +
# get view results from Jan 1, 2019 to Jan 3, 2019
url = '{}views/?guids={}&start_datetime=2019-01-01&end_datetime=2019-01-03'.format(METRICS_BASE, SINGLE_PREPRINT_GUID)
res = requests.get(url, headers=headers)
print('Request URL was: {}'.format(url))
print(json.dumps(res.json(), indent=4))
# +
# get view results from March 1, 2019 at 1:00am UTC to March 1, 2019 at 1:30am UTC by 5 min intervals
url = '{}views/?guids={}&start_datetime=2019-03-01T01:00&end_datetime=2019-03-01T01:30&interval=5m'.format(METRICS_BASE, SINGLE_PREPRINT_GUID)
res = requests.get(url, headers=headers)
print('Request URL was: {}'.format(url))
print(json.dumps(res.json(), indent=4))
# +
# Access metrics for a withdrawn preprint from March 1, 2019 to yesterday at 11:59pm UTC
url = '{}views/?guids={}&start_datetime=2019-01-03'.format(METRICS_BASE, WITHDRAWN_PREPRINT_GUID)
res = requests.get(url, headers=headers)
print('Request URL was: {}'.format(url))
print(json.dumps(res.json(), indent=4))
# -
# # Advanced Query Examples
#
# The preprint metrics API also allows `POST` requests containing more complicated raw queries for preprint metrics. These requests are made to just the bare `/v2/metrics/preprints/views/` and `/v2/metrics/preprints/downloads/` endpoints, without any query parameters. All of the data for the query is contained in a JSON object included in the request's `POST` data.
#
# These queries can be anything at all conforming to [the elasticsearch query DSL](https://www.elastic.co/guide/en/elasticsearch/reference/current/query-dsl.html), so there are many, many options not just limited to what you see here.
#
# Results will be formatted in the raw elasticsearch format, and so won't conform to the specific format outlined above.
#
# From the [official preprint metrics docs](https://www.notion.so/cos/Impact-API-documentation-6d7c638c0cb642f8989287a1794580b2), each data point is stored with the following fields:
#
# - timestamp
# - provider_id, e.g. "socarxiv"
# - preprint_id, e.g. "qmdc4"
# - user_id, e.g. "q7fts"
# - version (file version)
# - path
#
# You can use any of those fields when building a custom query. Note, you'll have to add "keyword" to queries for `path`, `provider_id`, or `preprint_id` - see examples below.
#
#
# +
post_url = '{}downloads/'.format(METRICS_BASE) # TODO - here's where to leave off
# total preprint downloads per year
query = {
"aggs" : {
"preprints_over_time" : {
"date_histogram" : {
"field" : "timestamp",
"interval" : "year"
}
}
}
}
payload = {
'data': {
'type': 'preprint_metrics',
'attributes': {
'query': query
}
}
}
res = requests.post(post_url, headers=headers, json=payload)
res.json()['aggregations']['preprints_over_time']['buckets']
# +
# see views/downloads broken down by month for one provider
# restricted to one year
query = {
"query": {
"term": {"provider_id.keyword": "psyarxiv"}
},
"aggs" : {
"preprints_from_2017": {
"filter": {
"range" : {
"timestamp" : {
"gte" : "2017-01-01",
"lt" : "2017-12-31"
}
}
},
"aggs": {
"preprints_per_month" : {
"date_histogram" : {
"field" : "timestamp",
"interval" : "month"
}
}
}
}
}
}
payload = {
'data': {
'type': 'preprint_metrics',
'attributes': {
'query': query
}
}
}
res = requests.post(post_url, headers=headers, json=payload)
res.json()['aggregations']['preprints_from_2017']['preprints_per_month']['buckets']
# +
# downloads that come from logged in users
logged_in_query = {
"query": {
"exists" : { "field" : "user_id" }
},
"size": 0,
"aggs" : {
"preprints_per_year" : {
"date_histogram" : {
"field" : "timestamp",
"interval" : "year"
}
}
}
}
payload = {
'data': {
'type': 'preprint_metrics',
'attributes': {
'query': logged_in_query
}
}
}
res = requests.post(post_url, headers=headers, json=payload)
res.json()['aggregations']['preprints_per_year']['buckets']
# +
# downloads that come from NON-logged in users
logged_out_query = {
"query": {
"bool": {
"must_not": {
"exists": {
"field": "user_id"
}
}
}
},
"size": 0,
"aggs" : {
"preprints_per_year" : {
"date_histogram" : {
"field" : "timestamp",
"interval" : "year"
}
}
}
}
payload = {
'data': {
'type': 'preprint_metrics',
'attributes': {
'query': logged_out_query
}
}
}
res = requests.post(post_url, headers=headers, json=payload)
res.json()['aggregations']['preprints_per_year']['buckets']
# -
# # Notes and Extras
#
# Code below here is for reference, or to run little one off adjustments in the terminal for adding more metrics.
#
# It's commented out because it won't run in this notebook "as is" and would need adjustment by whomever was running it for local testing purposes.
# +
# A little bit of code to add views and downloads to certain preprints for a developer
# uncomment, adjust, and run me in an interactive shell to add views/downloads
# from datetime import datetime
# from osf.metrics import PreprintView, PreprintDownload
# from osf.models import Preprint
# me = OSFUser.objects.get(username='<EMAIL>')
# user_to_use = OSFUser.objects.get(username='<EMAIL>')
# metric_dates = ['2017-01-01', '2018-01-02', '2019-01-03']
# times = ['T00:00', 'T01:00', 'T02:00']
# preps = [Preprint.load('ythm7'), Preprint.load('h5rgp'), Preprint.load('e3fq4')]
# metrics = [PreprintView, PreprintDownload]
# for preprint_to_add in preps:
# for metric in metrics:
# for date in metric_dates:
# for time in times:
# metric.record_for_preprint(
# preprint=preprint_to_add,
# user=user_to_use,
# path=preprint_to_add.primary_file.path,
# timestamp=datetime.strptime(date + time, '%Y-%m-%dT%H:%M')
# )
| Querying_Preprint_Metrics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import seaborn as sns
from sklearn.preprocessing import LabelEncoder, MinMaxScaler, OneHotEncoder
import os
import matplotlib.pyplot as plt
import collections
from sklearn.model_selection import train_test_split, StratifiedKFold, KFold
import gc
from sklearn.metrics import accuracy_score, roc_auc_score
from tqdm.notebook import tqdm
import lightgbm as lgb
import shap
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
pd.options.mode.chained_assignment = None
# -
path = '../input/weather-dataset-rattle-package'
def chunk_shapper(X,model, n = 20, verbose=True, dim = 20000):
pos = np.sort(np.random.randint(X.shape[0], size=dim))
X = X.iloc[pos, :].reset_index(drop = True)
predict_fold = KFold(n)
result = np.zeros(X.shape)
if verbose:
for trn_idx, val_idx in tqdm(predict_fold.split(X)):
result[val_idx,:] = model.shap_values(X.iloc[val_idx,:])[1]
else:
for trn_idx, val_idx in predict_fold.split(X):
result[val_idx,:] = model.shap_values(X.iloc[val_idx,:])[1]
return(result,pos)
# +
def sd(col, max_loss_limit=0.001, avg_loss_limit=0.001, na_loss_limit=0, n_uniq_loss_limit=0, fillna=0):
"""
max_loss_limit - don't allow any float to lose precision more than this value. Any values are ok for GBT algorithms as long as you don't unique values.
See https://en.wikipedia.org/wiki/Half-precision_floating-point_format#Precision_limitations_on_decimal_values_in_[0,_1]
avg_loss_limit - same but calculates avg throughout the series.
na_loss_limit - not really useful.
n_uniq_loss_limit - very important parameter. If you have a float field with very high cardinality you can set this value to something like n_records * 0.01 in order to allow some field relaxing.
"""
is_float = str(col.dtypes)[:5] == 'float'
na_count = col.isna().sum()
n_uniq = col.nunique(dropna=False)
try_types = ['float16', 'float32']
if na_count <= na_loss_limit:
try_types = ['int8', 'int16', 'float16', 'int32', 'float32']
for type in try_types:
col_tmp = col
# float to int conversion => try to round to minimize casting error
if is_float and (str(type)[:3] == 'int'):
col_tmp = col_tmp.copy().fillna(fillna).round()
col_tmp = col_tmp.astype(type)
max_loss = (col_tmp - col).abs().max()
avg_loss = (col_tmp - col).abs().mean()
na_loss = np.abs(na_count - col_tmp.isna().sum())
n_uniq_loss = np.abs(n_uniq - col_tmp.nunique(dropna=False))
if max_loss <= max_loss_limit and avg_loss <= avg_loss_limit and na_loss <= na_loss_limit and n_uniq_loss <= n_uniq_loss_limit:
return col_tmp
# field can't be converted
return col
def reduce_mem_usage_sd(df, deep=True, verbose=False, obj_to_cat=False):
numerics = ['int16', 'uint16', 'int32', 'uint32', 'int64', 'uint64', 'float16', 'float32', 'float64']
start_mem = df.memory_usage(deep=deep).sum() / 1024 ** 2
for col in tqdm(df.columns):
col_type = df[col].dtypes
# collect stats
na_count = df[col].isna().sum()
n_uniq = df[col].nunique(dropna=False)
# numerics
if col_type in numerics:
df[col] = sd(df[col])
# strings
if (col_type == 'object') and obj_to_cat:
df[col] = df[col].astype('category')
if verbose:
print(f'Column {col}: {col_type} -> {df[col].dtypes}, na_count={na_count}, n_uniq={n_uniq}')
new_na_count = df[col].isna().sum()
if (na_count != new_na_count):
print(f'Warning: column {col}, {col_type} -> {df[col].dtypes} lost na values. Before: {na_count}, after: {new_na_count}')
new_n_uniq = df[col].nunique(dropna=False)
if (n_uniq != new_n_uniq):
print(f'Warning: column {col}, {col_type} -> {df[col].dtypes} lost unique values. Before: {n_uniq}, after: {new_n_uniq}')
end_mem = df.memory_usage(deep=deep).sum() / 1024 ** 2
percent = 100 * (start_mem - end_mem) / start_mem
print('Mem. usage decreased from {:5.2f} Mb to {:5.2f} Mb ({:.1f}% reduction)'.format(start_mem, end_mem, percent))
return df
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
def feature_engineer_pipeline(data):
data['Date'] = pd.to_datetime(data['Date'])
data["month"] = data['Date'].apply(lambda x : x.month)
data["day"] = data['Date'].apply(lambda x : x.day)
data["year"] = data['Date'].apply(lambda x : x.day)
data['delta_temp'] = data['MinTemp'] - data['MaxTemp']
data['delta_wind_speed'] = data['WindSpeed3pm'] - data['WindSpeed9am']
data['delta_humidity'] = data['Humidity3pm'] - data['Humidity9am']
data['delta_pressure'] = data['Pressure3pm'] - data['Pressure9am']
data['delta_cloud'] = data['Cloud3pm'] - data['Cloud9am']
data['delta_temp'] = data['Temp3pm'] - data['Temp9am']
#add country
to_drop = ['RISK_MM', 'Date']
data = data.drop(to_drop, axis = 1)
return(data)
def aggregation_pipeline(data, numeric_col):
for col in tqdm(numeric_col):
gc.collect()
data[f'{col}_agg_month_loc'] = data[['month', 'Location', col]].groupby(['month', 'Location']).transform(np.nanmean)
data[f'{col}_diff_month_loc'] = data[col] - data[f'{col}_agg_month_loc']
gc.collect()
data[f'{col}_agg_year_loc'] = data[['year', 'Location', col]].groupby(['year', 'Location']).transform(np.nanmean)
data[f'{col}_diff_year_loc'] = data[col] - data[f'{col}_agg_year_loc']
gc.collect()
data[f'{col}_agg_day_loc'] = data[['day', 'Location', col]].groupby(['day', 'Location']).transform(np.nanmean)
data[f'{col}_diff_day_loc'] = data[col] - data[f'{col}_agg_day_loc']
return data
def remover_pipeline(data, categoric_column, numeric_column):
to_erase = ['RainToday']
data = data.drop('RainToday', axis = 1)
categoric_column = [x for x in categoric_column if x not in to_erase]
numeric_column = [x for x in numeric_column if x not in to_erase]
return data, categoric_column, numeric_column
def loader(path = path):
data = pd.read_csv(os.path.join(path, 'weatherAUS.csv'))
data = reduce_mem_usage_sd(data)
data = feature_engineer_pipeline(data)
data = reduce_mem_usage_sd(data)
categoric_col = ['Location', 'WindGustDir', 'WindDir9am', 'WindDir3pm', 'RainToday', 'month', 'day', 'year', 'RainTomorrow']
numeric_col = [x for x in data.columns if x not in categoric_col]
data[categoric_col] = data[categoric_col].astype(str, copy = True)
data, categoric_col, numeric_col = remover_pipeline(data, categoric_col, numeric_col)
data = aggregation_pipeline(data, numeric_col)
col_strat = ['RainTomorrow', 'Location']
strat = data[col_strat].astype(str).apply(lambda row: '_'.join(row.values.astype(str)), axis=1)
le = LabelEncoder()
strat = le.fit_transform(strat).astype(np.int16)
assert collections.Counter(strat).most_common()[-1][1] > 15
for col in categoric_col:
le = LabelEncoder()
data[col] = le.fit_transform(data[col]).astype(np.int16)
categoric_col = [x for x in categoric_col if x != 'RainTomorrow']
data = reduce_mem_usage_sd(data)
return data, numeric_col, categoric_col, strat
def split_pipeline(data, strat):
train, test, train_strat, _ = train_test_split(data, strat, test_size = 0.33, random_state = 12345, shuffle = True, stratify = strat)
train.reset_index(drop = True, inplace = True)
test.reset_index(drop = True, inplace = True)
y_train, y_test = train.pop('RainTomorrow').values, test.pop('RainTomorrow').values
return(train, test, y_train, y_test, train_strat)
# -
data, numeric_col, categoric_col, strat = loader()
df_train, df_test, y_train, y_test, train_strat = split_pipeline(data, strat)
gc.collect()
seed = 1
params = {
'objective': 'binary',
'boosting_type':'gbdt',
'metric':'auc',
'learning_rate':0.05,
'colsample_bytree': 0.8,
# 'lambda_l1': 2,
# 'lambda_l2': 2,
'max_depth': -1,
'num_leaves': 2**8,
'subsample': 0.75,
'tree_learner':'serial',
'max_bin':255,
'seed': seed,
'n_jobs':-1,
'importance_type':'gain',
}
# +
n_fold = 5
folds = StratifiedKFold(n_splits=n_fold, shuffle=True, random_state=0)
pred = np.zeros(df_train.shape[0])
pred_test = np.zeros(df_test.shape[0])
shap_matrix = np.zeros(df_train.shape)
score = 0
model5 =[]
gc.collect()
for fold_ , (trn_idx, val_idx) in tqdm(enumerate(folds.split(df_train, train_strat))):
gc.collect()
train_x, train_y = df_train.loc[trn_idx,:], y_train[trn_idx]
valid_x, valid_y = df_train.loc[val_idx,:], y_train[val_idx]
model = lgb.train(
params,
lgb.Dataset(train_x, label = train_y, categorical_feature = categoric_col),
3500, valid_sets = lgb.Dataset(valid_x, label = valid_y, categorical_feature = categoric_col),
categorical_feature = categoric_col,
valid_names ='validation',
verbose_eval = 50, early_stopping_rounds = 50
)
temp_y = model.predict(valid_x)
pred[val_idx] = temp_y
pred_test += model.predict(df_test)/n_fold
score += roc_auc_score(valid_y, temp_y)/n_fold
print('\nFold: {}; Auc: {:.3f}\n'.format(fold_, roc_auc_score(valid_y, temp_y)))
model5.append(model)
print('CV - Auc : {}\n'.format(score))
test_score = roc_auc_score(y_test, pred_test)
print('Test - Auc : {}\n'.format(test_score))
# +
feature_importances = pd.DataFrame()
feature_importances['feature'] = df_train.columns
for fold_, mod in tqdm(enumerate(model5)):
feature_importances['fold_{}'.format(fold_ + 1)] = mod.feature_importance(importance_type='gain')
scaler = MinMaxScaler(feature_range=(0, 100))
feature_importances['average'] = scaler.fit_transform(X=pd.DataFrame(feature_importances[['fold_{}'.format(fold + 1) for fold in range(folds.n_splits)]].mean(axis=1)))
fig = plt.figure(figsize=(12,8))
sns.barplot(data=feature_importances.sort_values(by='average', ascending=False).head(50), x='average', y='feature');
plt.title('50 TOP feature importance over {} average'.format(fold_+1))
| Other/weather-lgbm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
# To make data visualisations display in Jupyter Notebooks
import numpy as np # linear algebra
import pandas as pd # Data processing, Input & Output load
import matplotlib.pyplot as plt # Visualization & plotting
import datetime
from sklearn.svm import SVC
import joblib #Joblib is a set of tools to provide lightweight pipelining in Python (Avoid computing twice the same thing)
from sklearn.model_selection import train_test_split, cross_val_score, GridSearchCV
# GridSearchCV - Implements a “fit” and a “score” method
# train_test_split - Split arrays or matrices into random train and test subsets
# cross_val_score - Evaluate a score by cross-validation
from sklearn.metrics import f1_score, roc_auc_score, recall_score, precision_score, make_scorer, accuracy_score, roc_curve, confusion_matrix, classification_report
# Differnt metrics to evaluate the model
import warnings # To avoid warning messages in the code run
warnings.filterwarnings("ignore")
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', None)
# -
def plot_roc_curve(y_train_actual, train_pred_prob, y_test_actual, test_pred_prob, *args):
'''
Generate the train & test roc curve
'''
AUC_Train = roc_auc_score(y_train_actual, train_pred_prob)
AUC_Test = roc_auc_score(y_test_actual, test_pred_prob)
if len(args) == 0:
print("Train AUC = ", AUC_Train)
print("Test AUC = ", AUC_Test)
fpr, tpr, thresholds = roc_curve(y_train_actual, train_pred_prob)
fpr_tst, tpr_tst, thresholds = roc_curve(y_test_actual, test_pred_prob)
roc_plot(fpr, tpr, fpr_tst, tpr_tst)
else:
AUC_Valid = roc_auc_score(args[0], args[1])
print("Train AUC = ", AUC_Train)
print("Test AUC = ", AUC_Test)
print("Validation AUC = ", AUC_Valid)
fpr, tpr, thresholds = roc_curve(y_train_actual, train_pred_prob)
fpr_tst, tpr_tst, thresholds = roc_curve(y_test_actual, test_pred_prob)
fpr_val, tpr_val, thresholds = roc_curve(args[0], args[1])
roc_plot(fpr, tpr, fpr_tst, tpr_tst, fpr_val, tpr_val)
def roc_plot(fpr, tpr, fpr_tst, tpr_tst, *args):
'''
Generates roc plot
'''
fig = plt.plot(fpr, tpr, label='Train')
fig = plt.plot(fpr_tst, tpr_tst, label='Test')
if len(args) == 0:
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.0])
plt.title("ROC curve using ")
plt.xlabel('False Positive Rate (1 - Specificity)')
plt.ylabel('True Positive Rate (Sensitivity)')
plt.legend(loc='lower right')
plt.grid(True)
plt.show()
else:
fig = plt.plot(args[0], args[1], label='Validation')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.0])
plt.title("ROC curve using ")
plt.xlabel('False Positive Rate (1 - Specificity)')
plt.ylabel('True Positive Rate (Sensitivity)')
plt.legend(loc='lower right')
plt.grid(True)
plt.show()
# Read-in the dataset
Insurance_Data = pd.read_csv('carInsurance_train.csv')
print('Train Data Shape - ', Insurance_Data.shape)
Insurance_Data.head()
# What type of values are stored in the columns?
Insurance_Data.info()
# Let's look at some statistical information about our dataframe.
Insurance_Data.describe(include='all')
# This is how we can get summary for the categorical data
Insurance_Data.describe(include=np.object)
Target = 'CarInsurance'
pd.crosstab(Insurance_Data[Target], columns='N', normalize=True)
# pd.crosstab(Insurance_Data[Target], columns='N')
# count every row of class 1 as 2 rows of Class 1
0.599/0.401
num_cols = Insurance_Data.select_dtypes(include=[np.number]).columns.tolist()
non_num_cols = Insurance_Data.select_dtypes(exclude=[np.number]).columns.tolist()
num_cols
non_num_cols
# Lets drop columns which we will not use
num_cols = Insurance_Data.drop(['Id', 'CarInsurance'],axis=1).select_dtypes(include=[np.number]).columns.tolist()
non_num_cols = Insurance_Data.drop(['CallStart', 'CallEnd'],axis=1).select_dtypes(exclude=[np.number]).columns.tolist()
print('Numeric Columns \n', num_cols)
print('Non-Numeric Columns \n', non_num_cols)
# +
# Lets drop CarLoan, HHInsurance, Default from the numeric columns as these are dummies
num_cols_viz = ['DaysPassed', 'Age', 'NoOfContacts', 'PrevAttempts', 'LastContactDay', 'Balance']
fig, axes = plt.subplots(3,2,sharex=False,sharey=False, figsize=(15,15))
Insurance_Data.loc[:,[Target]+num_cols_viz].boxplot(by=Target, ax=axes, return_type='axes')
# -
non_num_cols_viz = non_num_cols+['CarLoan', 'HHInsurance', 'Default']
fig, axes = plt.subplots(len(non_num_cols_viz),sharex=False,sharey=False, figsize=(15,50))
for i in range(len(non_num_cols_viz)):
pd.crosstab(Insurance_Data[non_num_cols_viz[i]], Insurance_Data[Target]).plot(kind='bar',
stacked=True,
grid=False,
ax=axes[i],
rot=0)
Insurance_Data.isnull().sum()
Insurance_Data_Org = Insurance_Data.copy()
Insurance_Data['Job'].value_counts(dropna=False)
Insurance_Data['Job'] = Insurance_Data['Job'].fillna('None')
Insurance_Data['Job'].isnull().sum()
Insurance_Data['Job'].value_counts()
# +
# Fill missing education with the most common education level by job type
# Create job-education level mode mapping
edu_mode=[]
# What are different Job Types
job_types = Insurance_Data.Job.value_counts().index
job_types
# +
# Now according to the job type we will crate a mapping where the job and mode of education is there.
# It means when there are many people in the managment job then most of them are in which education.
# We can find that in below mapping
for job in job_types:
mode = Insurance_Data[Insurance_Data.Job==job]['Education'].value_counts().nlargest(1).index
edu_mode = np.append(edu_mode,mode)
edu_map=pd.Series(edu_mode,index=Insurance_Data.Job.value_counts().index)
edu_map
# +
# Apply the mapping to missing education obs. We will replace education now by jobs value
for j in job_types:
Insurance_Data.loc[(Insurance_Data['Education'].isnull()) & (Insurance_Data['Job']==j),'Education'] = edu_map.loc[edu_map.index==j][0]
# For those who are not getting mapped we will create a new category as None
Insurance_Data['Education'].fillna('None',inplace=True)
# -
Insurance_Data.isnull().sum()
# Fill missing communication with none
Insurance_Data['Communication'].value_counts(dropna=False)
Insurance_Data['Communication'] = Insurance_Data['Communication'].fillna('None')
# Check for missing value in Outcome
Insurance_Data['Outcome'].value_counts(dropna=False)
# +
# Fill missing outcome as not in previous campaign, we are adding one category to Outcome
# We will add category if the value of DaysPassed is -1
Insurance_Data.loc[Insurance_Data['DaysPassed']==-1,'Outcome']= 'NoPrev'
Insurance_Data['Outcome'].value_counts(dropna=False)
# -
# Check if we have any missing values left
Insurance_Data.isnull().sum()
Insurance_Data_num = Insurance_Data[num_cols+['Id', 'CarInsurance']]
# Categorical columns data
Insurance_Data_cat = Insurance_Data[non_num_cols]
non_num_cols
# Create dummies
Insurance_Data_cat_dummies = pd.get_dummies(Insurance_Data_cat) #One-Hot Embedding
print(Insurance_Data_cat_dummies.shape)
Insurance_Data_cat_dummies.head()
Insurance_Data_final = pd.concat([Insurance_Data_num, Insurance_Data_cat_dummies], axis=1)
print(Insurance_Data_final.shape)
Insurance_Data_final.head()
# Checking if there are missing values before we run model
Insurance_Data_final.isnull().sum(axis = 0)
train_df = Insurance_Data_final.drop(['Id', 'CarInsurance'], axis=1) #X
train_label = Insurance_Data_final['CarInsurance'] #y
#random_state is the seed used by the random number generator. It can be any integer.
# Train test split
X_train, X_test, y_train, y_test = train_test_split(train_df, train_label, train_size=0.7 , stratify=train_label, random_state=100)
# +
# Stratify option will make sure that train has both the classes, and also test has both the classes in 70-30
# Guarantee that:
# train 7 rows(5 rows class 0, 2 rows class 1)
# test 3 rows (2 rows class 0, 1 row of class1)
# -
print(y_train.shape[0]) # 2800, 1123 are 1s and the rest (2800-1123=1677) 0s
print(np.sum(y_train))
print(y_test.shape[0]) # 1200, 481 are 1s and the rest (1200-481=719) 0s
print(np.sum(y_test))
print('Train shape - ', X_train.shape)
print('Test shape - ', X_test.shape)
# Define Model parameters to tune
model_parameters = {
'kernel':['rbf'], #['rbf', 'poly', 'linear', 'sigmoid']
'C': [1], # [0, 1, 10, 100, 1000, 1e15] C is reciprocal of Lambda, strength of Lambda;
# Lambda = 0(Plain Logistic Regression , No Regularization), C = 1/0, feed very very big number to C (1e15)
# Lambda = Very big number(Heavy Regularization, weights will almost die,0), C = 1/inf, C = 0
'class_weight': ['balanced'], #['balanced', None]
'gamma': [0.0001] #[0.0001, 'scale', 'auto']
}
# +
# Gridsearch the parameters to find the best parameters.
model = SVC(probability=True, random_state=34) # Support vector classifier
gscv = GridSearchCV(estimator=model,
param_grid=model_parameters,
cv=3, # 3-Fold Cross Validation
#
verbose=3, #To print what it is doing
n_jobs=-1, #fastest possible depending in the laptop
scoring='roc_auc')
# scoring='roc_auc' (Almost always try that.)
gscv.fit(X_train, y_train)
# -
print('The best parameter are -', gscv.best_params_)
print(gscv.best_score_)
print(gscv.best_estimator_)
print(gscv.scorer_)
print('AUC on test by gscv =', roc_auc_score(y_true=y_test,
y_score=gscv.predict_proba(X_test)[:, 1]))
# +
# Generate ROC
plt.subplots(figsize=(10, 5))
train_prob = gscv.predict_proba(X_train)[:, 1]
test_prob = gscv.predict_proba(X_test)[:, 1]
plot_roc_curve(y_train, train_prob,
y_test, test_prob)
# -
| SVM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %load_ext autoreload
autoreload 2
# %matplotlib inline
# +
import matplotlib.pyplot as plt
import numpy as np
import sympy as sym
import seaborn as sn
import pycollocation
# -
# ## The basic model: amplification and persistance
# +
# need to specify some production function for gatherers
def output(k, alpha):
return k**alpha
def marginal_product_capital(k, alpha, **params):
return alpha * k**(alpha - 1)
def K_dot(t, K, B, q, u, a, R):
return (1 / u) * ((a + q) * K - R * B) - K
def B_dot(t, K, B, q, u, R):
return (1 / R) * q * K - B
def q_dot(t, K, B, q, u, R):
return (R - 1) * q - R * u
def u_dot(t, K, B, q, u, m, K_bar, R, **params):
return (1 / R) * marginal_product_capital((1 / m) * (K_bar - K), **params) - u
def rhs(t, K, B, q, u, a, m, K_bar, R, **params):
out = [K_dot(t, K, B, q, u, a, R),
B_dot(t, K, B, q, u, R),
q_dot(t, K, B, q, u, R),
u_dot(t, K, B, q, u, m, K_bar, R, **params)]
return out
# +
def steady_state_capital(a, m, K_bar, R, alpha, **params):
return K_bar - m * (alpha / (a * R))**(1 / (1 - alpha))
def steady_state_debt(a, m, K_bar, R, **params):
Kstar = steady_state_capital(a, m, K_bar, R, **params)
return (a / (R - 1)) * Kstar
def steady_state_land_price(a, R, **params):
return (R / (R - 1)) * a
def steady_state_user_cost(a, **params):
return a
def bcs_lower(t, K, B, q, u, K0, **params):
return [K - K0]
def bcs_upper(t, K, B, q, u, a, m, K_bar, R, **params):
Bstar = steady_state_debt(a, m, K_bar, R, **params)
qstar = steady_state_land_price(a, R)
ustar = steady_state_user_cost(a)
return [B - Bstar, q - qstar, u - ustar]
# -
params = {'a': 1.01, 'm': 10.0, 'alpha': 0.33, 'R': 1.5, 'K_bar': 100, 'K0': 95}
Kstar
B0
Bstar
# +
# specify an initial guess
domain = [0, 10]
ts = np.linspace(domain[0], domain[1], 1000)
Kstar = steady_state_capital(**params)
Ks = Kstar - (Kstar - params['K0']) * np.exp(-ts)
initial_capital_poly = np.polynomial.Chebyshev.fit(ts, Ks, 5, domain)
# initial value of debt is some multiple of capital stock
B0 = 1.5 * params['K0']
Bstar = steady_state_debt(**params)
Bs = Bstar - (Bstar - B0) * np.exp(-ts)
initial_debt_poly = np.polynomial.Chebyshev.fit(ts, Bs, 5, domain)
# starting with K0 < Kstar, must be that u0 > ustar
ustar = steady_state_user_cost(**params)
u0 = 1.5 * ustar
us = ustar - (ustar - u0) * np.exp(-ts)
initial_user_cost_poly = np.polynomial.Chebyshev.fit(ts, us, 5, domain)
# starting with K0 < Kstar, must be that q0 > qstar
qstar = steady_state_land_price(**params)
q0 = 1.05 * qstar
qs = qstar + (qstar - q0) * np.exp(-ts)
initial_land_price_poly = np.polynomial.Chebyshev.fit(ts, qs, 5, domain)
initial_coefs = np.hstack([initial_capital_poly.coef, initial_debt_poly.coef,
initial_user_cost_poly.coef, initial_land_price_poly.coef])
# -
nodes = pycollocation.PolynomialSolver.collocation_nodes(5, domain, "Chebyshev")
problem = pycollocation.TwoPointBVP(bcs_lower, bcs_upper, 1, 4, rhs, params)
solution = pycollocation.PolynomialSolver.solve({'kind': "Chebyshev"},
initial_coefs,
domain,
nodes,
problem)
pycollocation.PolynomialSolver._array_to_list(initial_coefs, 4)
initial_capital_poly.coef
solution.result
K_hat, B_hat, q_hat, u_hat = solution.functions
# +
pts = np.linspace(domain[0], domain[1], 1000)
fig, axes = plt.subplots(4, 1)
axes[0].plot(pts, K_hat(pts))
axes[1].plot(pts, B_hat(pts))
axes[2].plot(pts, q_hat(pts))
axes[3].plot(pts, q_hat(pts))
fig.tight_layout()
plt.show()
# -
K_resids, B_resids, q_resids, u_resids = solution.residuals(pts)
# +
pts = np.linspace(domain[0], domain[1], 1000)
fig, axes = plt.subplots(4, 1)
axes[0].plot(pts, K_resids)
axes[1].plot(pts, B_resids)
axes[2].plot(pts, q_resids)
axes[3].plot(pts, u_resids)
fig.tight_layout()
plt.show()
# -
basic_model_solver.solve(kind="Chebyshev",
coefs_dict=initial_coefs,
domain=domain)
basic_model_solver.result["success"]
basic_model_viz = pycollocation.Visualizer(basic_model_solver)
basic_model_viz.interpolation_knots = np.linspace(domain[0], domain[1], 1000)
basic_model_viz.solution.plot(subplots=True, style=['r', 'b'])
plt.show()
# Solution is not as accurate as I would like...
basic_model_viz.residuals.plot(subplots=True, style=['r', 'b'])
plt.show()
# ...actually, when using noramlized residuals eveything looks great!
basic_model_viz.normalized_residuals.plot(logy=True, sharey=True)
plt.show()
assets = basic_model_viz.solution[['q', 'K']].prod(axis=1)
liabilities = basic_model_viz.solution.B
equity = assets - liabilities
leverage = assets / equity
leverage.plot()
# +
def credit_cycles(t, X, a, m, alpha, R, K_bar):
out = np.array([(1 / X[3]) * ((a + X[2]) * X[0] - R * X[1]) - X[0],
(1 / R) * X[2] * X[0] - X[1],
(R - 1) * X[2] - R * X[3],
(alpha / R) * ((1 / m) * (K_bar - X[0]))**(alpha - 1) - X[3]])
return out
def jacobian(t, X, a, m, alpha, R, K_bar):
out = np.array([[((a + X[2]) / X[3]) - 1.0, -R / X[3], X[0] / X[3], -X[3]**(-2)],
[(1 / R) * X[2], -1.0, (1 / R) * X[0], 0.0],
[0.0, 0.0, R - 1, -R],
[-(1 / m) * (alpha - 1) * (alpha / R) * ((1 / m) * (K_bar - X[0]))**(alpha - 2), 0.0, 0.0, -1.0]])
return out
def Kstar(a, m, alpha, R, K_bar):
return K_bar - m * (alpha / (a * R))**(1 / (1 - alpha))
def Bstar(a, m, alpha, R, K_bar):
return (a / (R - 1)) * Kstar(a, m, alpha, R, K_bar)
# -
initial_condition = np.array([Kstar(a, m, alpha, R, K_bar), Bstar(a, m, alpha, R, K_bar), (R / (R - 1)) * a, a])
initial_condition
credit_cycles(0, initial_condition)
jacobian(0, initial_condition)
from scipy import linalg
from IPython.html.widgets import fixed, interact, FloatSliderWidget
def eigenvalues(a=1.0, m=1.0, alpha=0.33, R=1.05, K_bar=10.0):
steady_state = np.array([Kstar(a, m, alpha, R, K_bar),
Bstar(a, m, alpha, R, K_bar),
(R / (R - 1)) * a,
a])
vals, vecs = linalg.eig(jacobian(0, steady_state, a, m, alpha, R, K_bar))
print vals
interact(eigenvalues, a=(0.0, 1e3, 1e0), m=(0.0, 1e2, 1e-1), R=(0.0, 1e2, 1e-2), K_bar=(0.0, 1e4, 1e1))
params = 2.0, 0.5, 0.33, 1.01, 500.0
problem = ivp.IVP(credit_cycles, jacobian)
problem.f_params = params
problem.jac_params = params
# <h2> Full model </h2>
# +
lamda, pi, phi, = sym.symbols('lamda, pi, phi')
# full model from Kiyotaki and Moore "credit-cycles" paper
K_dot = (pi / (phi + u)) * ((a + q + lamda * phi) * K - R * B) - pi * lamda * K
B_dot = (R - 1) * B + (phi * (1 - lamda) - a) * K
q_dot = (R - 1) * q - R * u
u_dot = (1 / R) * mpk.subs({k: (1 / m) * (K_bar - K)}) - u
rhs = {'K': K_dot, 'B': B_dot, 'q': q_dot, 'u': u_dot}
# +
bcs = {}
ustar = ((pi * a - (1 - lamda) * (1 - R + pi * R) * phi) /
(lamda * pi + (1 - lamda) * (1 - R + pi * R)))
qstar = (R / (R - 1)) * ustar
Kstar = K_bar - m * (alpha / (ustar * R))**(1 / (1 - alpha))
Bstar = ((a - (1 - lamda) * phi) / (R - 1)) * Kstar
# initial conditions for K and B are given
K0 = 75
bcs['lower'] = [K - K0]
# boundary conditions on B, q and u can be written in terms of steady state values
bcs['upper'] = [B - Bstar, q - qstar, u - ustar]
# -
params = {'a': 1.05, 'pi': 0.05, 'phi': 20.0, 'lamda': 0.975,'m': 1.0, 'alpha': 0.16,
'R': 1.01, 'K_bar': 100}
# +
# set up the model and solver
full_model = pycollocation.SymbolicBoundaryValueProblem(dependent_vars=['K', 'B', 'q', 'u'],
independent_var='t',
rhs=rhs,
boundary_conditions=bcs,
params=params)
full_model_solver = pycollocation.OrthogonalPolynomialSolver(full_model)
# +
def Kstar(a, phi, R, alpha, pi, m, lamda, K_bar):
return K_bar - m * (alpha / (ustar(a, phi, R, alpha, pi, m, lamda, K_bar) * R))**(1 / (1 - alpha))
def Bstar(a, phi, R, alpha, pi, m, lamda, K_bar):
return ((a - (1 - lamda) * phi) / (R - 1)) * Kstar(a, phi, R, alpha, pi, m, lamda, K_bar)
def qstar(a, phi, R, alpha, pi, m, lamda, K_bar):
return (R / (R - 1)) * ustar(a, phi, R, alpha, pi, m, lamda, K_bar)
def ustar(a, phi, R, alpha, pi, m, lamda, K_bar):
u = ((pi * a - (1 - lamda) * (1 - R + pi * R) * phi) /
(lamda * pi + (1 - lamda) * (1 - R + pi * R)))
return u
# specify an initial guess
domain = [0, 25]
ts = np.linspace(domain[0], domain[1], 1000)
Ks = Kstar(**params) - (Kstar(**params) - K0) * np.exp(-ts) * np.cos(2.0 * np.pi * ts)
initial_capital_poly = np.polynomial.Chebyshev.fit(ts, Ks, 25, domain)
# initial value of debt is some multiple of capital stock
B0 = 1.5 * K0
Bs = Bstar(**params) - (Bstar(**params) - B0) * np.exp(-ts) #* np.cos(2.0 * np.pi * ts)
initial_debt_poly = np.polynomial.Chebyshev.fit(ts, Bs, 25, domain)
# starting with K0 > Kstar, must be that u0 > ustar
us = ustar(**params) - (ustar(**params) - 1.5 * ustar(**params)) * np.exp(-ts) #* np.cos(2.0 * np.pi * ts)
initial_user_cost_poly = np.polynomial.Chebyshev.fit(ts, us, 25, domain)
# starting with K0 > Kstar, must be that q0 > qstar
qs = qstar(**params) - (qstar(**params) - 1.5 * qstar(**params)) * np.exp(-ts) #* np.cos(2.0 * np.pi * ts)
initial_land_price_poly = np.polynomial.Chebyshev.fit(ts, qs, 25, domain)
initial_coefs = {'K': initial_capital_poly.coef, 'B': initial_debt_poly.coef,
'u': initial_user_cost_poly.coef, 'q': initial_land_price_poly.coef}
# +
def jacobian(t, X, a, phi, R, alpha, pi, m, lamda, K_bar):
out = np.array([[(pi / (phi + X[3])) * (a + X[2] + lamda * phi) - pi * lamda, -(pi / (phi + X[3])) * R, (pi / (phi + X[3])) * X[0], -(pi / (phi + X[3])**2) * ((a + X[2] + lamda * phi) * X[0] - R * X[1])],
[(R - 1) * X[1] + (phi * (1 - lamda) - a), (R - 1), 0.0, 0.0],
[0.0, 0.0, R - 1, -R],
[-(1 / m) * (alpha - 1) * (alpha / R) * ((1 / m) * (K_bar - X[0]))**(alpha - 2), 0.0, 0.0, -1.0]])
return out
def eigenvalues(a=1.0, phi=20.0, pi=0.05, lamda=0.975, m=1.0, alpha=0.33, R=1.05, K_bar=10.0):
steady_state = np.array([Kstar(a, phi, R, alpha, pi, m, lamda, K_bar),
Bstar(a, phi, R, alpha, pi, m, lamda, K_bar),
qstar(a, phi, R, alpha, pi, m, lamda, K_bar),
ustar(a, phi, R, alpha, pi, m, lamda, K_bar)])
vals, vecs = linalg.eig(jacobian(0, steady_state, a, phi, R, alpha, pi, m, lamda, K_bar))
print vals
print np.absolute(vals)
# -
interact(eigenvalues, a=(1.0, 2.0, 1e-2), alpha=(1e-2, 1-1e-2, 1e-2), m=(0.0, 1e2, 1e-1), R=(0.0, 1e2, 1e-2), K_bar=(0.0, 1e4, 1e1))
full_model_solver.solve(kind="Chebyshev",
coefs_dict=initial_coefs,
domain=domain)
full_model_solver.result["success"]
full_model_viz = pycollocation.Visualizer(full_model_solver)
full_model_viz.interpolation_knots = np.linspace(domain[0], domain[1], 1000)
full_model_viz.solution.plot(subplots=True)
full_model_viz.normalized_residuals.plot(subplots=True)
plt.show()
| examples/credit-cycles.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import pickle
from tqdm.notebook import tqdm
# model
from sklearn.cross_decomposition import PLSRegression
# math
from sklearn.model_selection import cross_val_score
from sklearn.metrics import mean_squared_error
from math import sqrt, isnan
from sklearn.metrics import r2_score
fp = "G:\\My Drive\\Darby Work\\Ytsma and Dyar 2021 (LOD paper)\\"
# -
# #### Compositions
# generate comps
comps_path = fp + "tables\\TableS1_sample_compositions.xlsx"
lanl_comps = pd.read_excel(comps_path, sheet_name = "LANL")
mhc_comps = pd.read_excel(comps_path, sheet_name = "MHC")
comps = pd.merge(mhc_comps, lanl_comps, how = "outer") # merge comps
comps.columns = comps.columns.map(lambda x: x.split()[0])
comps = comps.drop_duplicates(subset = 'Sample') # remove duplicates
comps['Sample'] = comps['Sample'].astype(str)
comps = comps.sort_values(by='Sample')
comps = comps.replace(np.nan, "", regex=True)
cols = comps.columns.drop('Sample')
comps[cols] = comps[cols].apply(pd.to_numeric) # make columns numeric
# add random number assignment
rd = pd.read_excel('Z:\\Millennium Set\\Millennium_COMPS_viewonly.xlsx', usecols=[0,2])
rd = rd.drop([0,1]).rename(columns={'DO NOT TOUCH THIS':'Sample',
'X.1':'rand_num'}).reset_index(drop=True)
comps = pd.merge(rd, comps, how='right', on='Sample')
# #### Datasets (baseline removal and normalization already applied)
cl_earth = pd.read_csv(fp+'CL_all_Earth_spectra.csv')
cl_mars = pd.read_csv(fp+'CL_all_Mars_spectra.csv')
cl_vac = pd.read_csv(fp+'CL_all_Vacuum_spectra.csv')
cc_mars = pd.read_csv(fp+'CC_all_Mars_spectra.csv')
# #### Sensitivities
sensitivities = pd.read_csv(fp+'instrument_sensitivities.csv')
# #### Split test and train
train_250_1000 = comps[comps.rand_num >= 250].reset_index(drop=True)
train_0_750 = comps[comps.rand_num <= 750].reset_index(drop=True)
test_250_1000 = comps[comps.rand_num < 250].reset_index(drop=True)
test_0_750 = comps[comps.rand_num > 750].reset_index(drop=True)
# #### Outlier limits
# Calculated by 1.5*IQR + Q3 on entire MHC dataset or highest natural sample for doped elements
outlier_limits = pd.read_csv('Z:\\Millennium Set\\NEW_OUTLIER_LIMITS.csv')
iqr_outliers = dict(zip(outlier_limits.element, outlier_limits.iqr_q3_outlier_limit))
dope_outliers = dict(zip(outlier_limits.element, outlier_limits.highest_natural_for_doped))
# #### Make models per element
elements = ['MnO', 'Na2O', 'SiO2', 'Li', 'Ni', 'Pb', 'Rb', 'Sr', 'Zn']
n_ranges = ['0-750', '250-1000']
factors = {
'LOB' : 1.645,
'LOD' : 3.3,
'LOQ' : 10
}
methods = ['braga', 'metals']
dfs = [cl_earth,cl_mars,cl_vac,cc_mars]
df_names = ['CL_Earth', 'CL_Mars', 'CL_Vac', 'CC_Mars']
mhc_list = [cl_earth,cl_mars,cl_vac]
outliers = [iqr_outliers, dope_outliers]
# +
# PLS parameters
n_folds = 5
max_components = 30
# prep for results
n_range_list = []
element_list = []
atm_list = []
inst_list = []
n_train_list = []
rmsecv_list = []
component_list = []
rmsec_list = []
train_r2_list = []
train_adj_r2_list = []
lob_list = []
lod_list = []
loq_list = []
outlier_list = []
n_test_list = []
rmsep_list = []
test_r2_list = []
test_adj_r2_list = []
method_list = []
for n_range in tqdm(n_ranges, desc='Number ranges'):
if n_range == '0-750':
all_train = train_0_750
all_test = test_0_750
else:
all_train = train_250_1000
all_test = test_250_1000
for element in tqdm(elements, leave=False, desc='Elements'):
count = 0
for df in tqdm(dfs, leave=False, desc='Dataset'):
if df_names[count].split('_')[0]=='CC':
inst='LANL'
else:
inst='ChemLIBS'
if df_names[count].split('_')[1]=='Vac':
atm = 'Vacuum'
else:
atm = df_names[count].split('_')[1]
outpath = "{}\\python_models\\{}_{}\\".format(fp, df_names[count], n_range)
count +=1
count1 = 0
for outlier in outliers:
n_folds = 5
max_components = 30
if count1 == 0:
o = 'iqr_q3'
else:
o = 'highest_natural'
count1 += 1
out_lim = outlier[element]
if isnan(out_lim):
temp_train = all_train.dropna(subset=[element]).reset_index(drop=True)[['Sample', element]]
temp_test = all_test.dropna(subset=[element]).reset_index(drop=True)[['Sample', element]]
else:
temp_train = all_train[all_train[element] <= out_lim].reset_index(drop=True)[['Sample', element]]
temp_test = all_test[all_test[element] <= out_lim].reset_index(drop=True)[['Sample', element]]
# train metadata
train_names = sorted(set(temp_train.Sample).intersection(df.columns)) # sorted
y_train = temp_train[temp_train.Sample.isin(train_names)][element].values # already alphabetized
n_train = len(y_train)
if n_train < n_folds:
n_folds = n_train
# train spectra
X_train = df[train_names]
spec_list = []
for column in X_train.columns:
spectrum = list(X_train[column])
spec_list.append(spectrum)
X_train = np.array(spec_list)
#---------------------CROSS--VALIDATION and TRAINING--------------------------#
cv_dict = {}
for n_components in np.arange(start=2, stop=max_components+1, step=1):
# define model
temp_pls = PLSRegression(n_components = n_components, scale=False)
# run CV and get RMSE
temp_rmsecv = (-cross_val_score(
temp_pls, X_train, y_train, cv=n_folds, scoring='neg_root_mean_squared_error'
)).mean()
# add results to dictionary
cv_dict.update({temp_rmsecv : n_components})
# select parameters of model with lowest rmsecv
rmsecv = min(list(cv_dict.keys()))
component = cv_dict[rmsecv]
model = PLSRegression(n_components = component, scale=False)
model.fit(X_train, y_train)
pickle.dump(model, open(outpath+element+'_'+o+'_model.asc', 'wb'), protocol=0)
coeff = pd.DataFrame(model.coef_)
coeff.to_csv(outpath+element+'_'+o+'_coeffs.csv', index=False)
for method in methods:
#---------------------CALCULATE---LBDQ--------------------------#
sensitivity = sensitivities[
(sensitivities.instrument == inst) &
(sensitivities.atmosphere == atm) &
(sensitivities.method == method)
]['sensitivity'].iloc[0]
# calculate regression vector
vector = pow(coeff, 2).sum().pow(.5) #square root of sum of squares
# calculate values
lob = factors['LOB'] * sensitivity * vector[0]
lod = factors['LOD'] * sensitivity * vector[0]
loq = factors['LOQ'] * sensitivity * vector[0]
#---------------------CALIBRATION--ERROR--------------------------#
train_pred = model.predict(X_train)
train_pred_true = pd.DataFrame({
'sample' : train_names,
'actual' : y_train.flatten().tolist(),
'pred' : train_pred.flatten().tolist()
})
temp = train_pred_true[(train_pred_true.pred < 100) &
(train_pred_true.pred > loq)].copy(deep=True)
if len(temp) == 0:
rmsec='NA'
train_r2 = 'NA'
train_adj_r2 = 'NA'
test_r2 = 'NA'
test_adj_r2 = 'NA'
rmsep = 'NA'
n_test = 'NA'
n_range_list.append(n_range)
outlier_list.append(o)
method_list.append(method)
element_list.append(element)
atm_list.append(atm)
inst_list.append(inst)
n_train_list.append(n_train)
rmsecv_list.append(rmsecv)
component_list.append(component)
lob_list.append(lob)
lod_list.append(lod)
loq_list.append(loq)
rmsec_list.append(rmsec)
train_r2_list.append(train_r2)
train_adj_r2_list.append(train_adj_r2)
n_test_list.append(n_test)
rmsep_list.append(rmsep)
test_r2_list.append(test_r2)
test_adj_r2_list.append(test_adj_r2)
continue
rmsec = sqrt(mean_squared_error(temp.actual, temp.pred))
train_r2 = model.score(X_train,y_train)
train_adj_r2 = 1 - (1-train_r2)*(len(temp) - 1) / (len(temp) - (temp.shape[1] - 1) - 1)
# fill with <LOQ / >100 wt%
loq_df = train_pred_true[train_pred_true.pred < loq].copy(deep=True)
loq_df['pred'] = '<LOQ'
over_df = train_pred_true[train_pred_true.pred > 100].copy(deep=True)
if len(over_df) > 0:
over_df['pred'] = '>100 wt%'
train_pred_true = pd.concat([temp, over_df, loq_df], ignore_index=True)
else:
train_pred_true = pd.concat([temp, loq_df], ignore_index=True)
train_pred_true.to_csv(outpath+element+"_"+o+"_"+method+'_train_preds.csv', index=False)
#------------------------TEST--MODEL------------------------#
# test metadata
test_names = sorted(set(temp_test.Sample).intersection(df.columns)) # sorted
y_test = temp_test[temp_test.Sample.isin(test_names)][element].values # already alphabetized
# test spectra
X_test = df[test_names]
spec_list = []
for column in X_test.columns:
spectrum = list(X_test[column])
spec_list.append(spectrum)
X_test = np.array(spec_list)
# run predictions
test_pred = model.predict(X_test)
# get RMSE-P
test_pred_true = pd.DataFrame({
'sample' : test_names,
'actual' : y_test,
'pred' : test_pred.flatten().tolist()
})
temp = test_pred_true[(test_pred_true.pred < 100) &
(test_pred_true.pred > loq)].copy(deep=True)
n_test = len(temp)
if n_test < 2:
test_r2 = 'NA'
test_adj_r2 = 'NA'
rmsep = 'NA'
else:
# get RMSE-P
rmsep = sqrt(mean_squared_error(temp.actual, temp.pred))
# get R2
test_r2 = r2_score(temp.actual,temp.pred)
# adjusted r2
test_adj_r2 = 1 - (1-test_r2)*(len(temp) - 1) / (len(temp) - (temp.shape[1] - 1) - 1)
# fill with <LOQ / >100 wt%
loq_df = test_pred_true[test_pred_true.pred < loq].copy(deep=True)
loq_df['pred'] = '<LOQ'
over_df = test_pred_true[test_pred_true.pred > 100].copy(deep=True)
if len(over_df) > 0:
over_df['pred'] = '>100 wt%'
test_pred_true = pd.concat([temp, over_df, loq_df], ignore_index=True)
else:
test_pred_true = pd.concat([temp, loq_df], ignore_index=True)
test_pred_true.to_csv(outpath+element+"_"+o+"_"+method+'_test_preds.csv', index=False)
n_range_list.append(n_range)
outlier_list.append(o)
method_list.append(method)
element_list.append(element)
atm_list.append(atm)
inst_list.append(inst)
n_train_list.append(n_train)
rmsecv_list.append(rmsecv)
component_list.append(component)
lob_list.append(lob)
lod_list.append(lod)
loq_list.append(loq)
rmsec_list.append(rmsec)
train_r2_list.append(train_r2)
train_adj_r2_list.append(train_adj_r2)
n_test_list.append(n_test)
rmsep_list.append(rmsep)
test_r2_list.append(test_r2)
test_adj_r2_list.append(test_adj_r2)
# +
results = pd.DataFrame({
'element':element_list,
'outlier_defn':outlier_list,
'instrument':inst_list,
'atmosphere':atm_list,
'method':method_list,
'num_range':n_range_list,
'n_train':n_train_list,
'rmsecv':rmsecv_list,
'components':component_list,
'lob':lob_list,
'lod':lod_list,
'loq':loq_list,
'rmsec':rmsec_list,
'train_r2':train_r2_list,
'train_adj_r2':train_adj_r2_list,
'n_test':n_test_list,
'rmsep':rmsep_list,
'test_r2':test_r2_list,
'test_adj_r2':test_adj_r2_list
})
results.to_csv(fp+'results_011422.csv', index=False)
# -
| Step 2. Make PLS models, get LBDQ, test models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# coding: utf-8
# In[30]:
from keras.models import Sequential
import numpy as np
import pandas
from keras.layers import Dense, Activation, Dropout, LSTM
import wrangle as wr
from matplotlib import pyplot
from keras import callbacks
from livelossplot.keras import PlotLossesCallback
model = Sequential()
model.add(Dense(units=30, input_dim=9,kernel_initializer='normal', activation='elu'))
# self.model.add(Dropout(0.1))
model.add(Dense(units=30, kernel_initializer='normal', activation='elu'))
# self.model.add(Dropout(0.1))
model.add(Dense(units=4,kernel_initializer='normal', activation='elu'))
model.compile(loss='mse', optimizer='adam')
model.summary()
# serialize model to JSON
model_json = model.to_json()
with open("model.json", "w") as json_file:
json_file.write(model_json)
dataset = pandas.read_csv("/home/letrend/workspace/roboy_control/data0.log", delim_whitespace=True, header=1)
dataset = dataset.values[:500000,0:]
np.random.shuffle(dataset)
quaternion_set = np.array(dataset[:,0:4])
sensors_set = np.array(dataset[:,4:13])
sensors_set = wr.mean_zero(pandas.DataFrame(sensors_set)).values
data_in_train = sensors_set[:int(len(sensors_set)*0.7),:]
data_in_test = sensors_set[int(len(sensors_set)*0.7):,:]
data_out_train = quaternion_set[:int(len(sensors_set)*0.7),:]
data_out_test = quaternion_set[int(len(sensors_set)*0.7):,:]
train_X = data_in_train
test_X = data_in_test
train_y = data_out_train
test_y = data_out_test
print(train_X.shape, train_y.shape, test_X.shape, test_y.shape)
filepath="model_checkpoint-{epoch:02d}-{val_loss:.10f}.hdf5"
checkpoint = callbacks.ModelCheckpoint(filepath, monitor='val_loss', verbose=1, save_best_only=True, mode='min')
callbacks_list = [checkpoint, PlotLossesCallback()]
# fit network
history = model.fit(train_X, train_y, epochs=100, batch_size=20, validation_data=(test_X, test_y), verbose=2, shuffle=True, callbacks=callbacks_list)
# plot history
pyplot.plot(history.history['loss'], label='train')
pyplot.plot(history.history['val_loss'], label='test')
pyplot.legend()
pyplot.show()
# result = model.predict(train_X)
# print(result)
# serialize weights to HDF5
model.save("model.h5")
print("Saved model to disk")
# -
result = grid_result.best_estimator_.model.predict(x)
print(result[0])
print(y[0])
numpy.linalg.norm(result-y)
| python_old/shoulder_training_mlp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:datasci]
# language: python
# name: conda-env-datasci-py
# ---
# +
import bs4
import calendar
import re
import requests
import numpy as np
import xarray as xr
from tqdm import tqdm_notebook as tqdm
from tqdm import tqdm as TQDM
# -
import matplotlib.pyplot as plt
# %matplotlib inline
PARSER = 'lxml-html'
start = 'X-Body-of-Message'
end = 'X-Body-of-Message-End'
yr = 2017
mon = 8
num = 34
MONTH_INDEX_URL_MASK = 'https://www.lists.rdg.ac.uk/archives/met-jobs/{yr}-{mon:02d}/date.html'
MSG_URL_MASK = 'https://www.lists.rdg.ac.uk/archives/met-jobs/{yr}-{mon:02d}/{msg_href}'
# 'https://www.lists.rdg.ac.uk/archives/met-jobs/{yrmon:%Y-%m}/msg{num:05d}.html'
def get_msg_urls(year, month):
msg_regex = 'msg([0-9]+)\.html'
index_url = MONTH_INDEX_URL_MASK.format(yr=year, mon=month)
r = requests.get(index_url)
hrefs = []
if r.status_code == 200:
soup = bs4.BeautifulSoup(r.content, PARSER)
for i in soup.find_all('a', href=re.compile(msg_regex)):
hrefs += [i['href']]
return hrefs
def extract_text_between_comments(soup, comm1, comm2):
def _extract_text(node):
return ''.join(t.strip() for t in node(text=True,
recursive=False))
comm1_tag = soup.find_all(text=comm1)[0]
comm2_tag = soup.find_all(text=comm2)[0]
after_comm1 = [*filter(None, [_extract_text(i) for i in comm1_tag.find_all_next()])]
before_comm2 = [*filter(None, [_extract_text(i) for i in comm2_tag.find_all_previous()])]
# elements_between = set(a.find_all_next()).intersection(b.find_all_previous())
text = ''.join([s.strip() for s in after_comm1 if s in before_comm2 and len(s)>1])
return text
def count_trigger(text, kwd):
return kwd in text.lower()
year_range = range(2003, 2018)
month_range = range(1, 13)
kwds = ['python',
'fortran',
'matlab',
'javascript',
'perl',
'shell',
'bash',
'latex',
'c++',
'programming',
'scripting',
'netcdf',
'wrf',
'linux',
'unix',
'windows',
'microsoft']
data = np.zeros((len(year_range), len(kwds)))
result = []
for i, yr in tqdm(enumerate(year_range), desc='Year loop'):
counts = {k: 0 for k in kwds}
counts['msg_count'] = 0
for j, mon in tqdm(enumerate(month_range), desc='Month loop', leave=False):
hrefs = get_msg_urls(yr, mon)
for href in tqdm(hrefs, desc='Day loop', leave=False):
msg_url = MSG_URL_MASK.format(yr=yr, mon=mon, msg_href=href)
req = requests.get(msg_url)
if req.status_code == 200:
counts['msg_count'] += 1
soup = bs4.BeautifulSoup(req.content, PARSER)
try:
txt = extract_text_between_comments(soup, start, end)
for kwd in kwds:
if count_trigger(txt, kwd):
counts[kwd] += 1
except:
TQDM.write(f'Unable to parse {msg_url}')
data[i, j] = count
result.append(counts)
import pandas as pd
df = pd.DataFrame(result, index=year_range)
df
languages = ['bash', 'c++', 'fortran', 'javascript', 'matlab', 'perl', 'python', 'shell']
df['weights'] = df.msg_count.max() / df.msg_count
pure_df = df[[i for i in df.columns if i != 'msg_count' and i != 'weights']]
weigthed_by_msg_count = pure_df.multiply(df.weights, axis=0)
lang_df = weigthed_by_msg_count[languages]
lang_df = lang_df.multiply(1 / lang_df.sum(axis=1), axis=0)
fig, ax = plt.subplots(figsize=(15, 7))
lang_df.plot.area(ax=ax)
| _blog/_notebooks/scrape-metjobs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Deep learning with Python <NAME> 2018
#
# Train a convolution network to classify digits using the MNIST dataset with augmenting the training set using image generation.
# ## Load the train and test sets
#
# We start by loading the train and test sets consisting of 60000 and 10000 28x28 pixel grayscale images. Each pixel has a value between 0 and 255 stored in numpy uint8.
# +
from keras import datasets
(orig_train_images, orig_train_labels), (orig_test_images, orig_test_labels) = datasets.mnist.load_data()
print(f'The shape of the training set is: {orig_train_images.shape}')
print(f'The shape of the test set is: {orig_test_images.shape}')
print(f'The data type of the training set is: {orig_train_images.dtype}')
print(f'The data type of the test set is: {orig_test_images.dtype}')
# -
# ## Example images
#
# Let's plot first images from the training and test sets with their associated labels to see what they look like.
# +
# %matplotlib inline
def show_image(title, images, labels):
from matplotlib import pyplot
pyplot.rcParams["figure.figsize"] = (2,2)
pyplot.title(f'{title} set image (label {labels[0]})')
pyplot.imshow(images[0], cmap="gray")
pyplot.show()
show_image('training', orig_train_images, orig_train_labels)
show_image('test', orig_test_images, orig_test_labels)
# -
# ## Shape the training and test set images
#
# We need to shape the images to be suitable for training the network. Images are converted from 28x28 2D tensors with 0-255 numpy uint8s to 28*28 1D tensors with 0-1 numpy float32s.
# +
train_images = orig_train_images.reshape((orig_train_images.shape[0], 28, 28, 1)).astype('float32') / 255
test_images = orig_test_images.reshape((orig_test_images.shape[0], 28, 28, 1)).astype('float32') / 255
print(f'The shape of the original training set: {orig_train_images.shape}')
print(f'The shape of the actual training set: {train_images.shape}')
print(f'The shape of the original test set: {orig_test_images.shape}')
print(f'The shape of the actual test set: {test_images.shape}')
print(f'The data type of the original training set: {orig_train_images.dtype}')
print(f'The data type of the actual training set: {train_images.dtype}')
print(f'The data type of the original test set: {orig_test_images.dtype}')
print(f'The data type of the actual test set: {test_images.dtype}')
# -
# ## Augment the training set by generating additional images
# +
from keras.preprocessing.image import ImageDataGenerator
gen_train_images = ImageDataGenerator(
rotation_range=20,
shear_range=0.2)
# -
# ## Shape the training and test set labels
#
# We need to also shape the labels to be suitable for training the network. Scalar labels are replaced with 1D binary tensors with value 0 or 1 for each label. If the value is 1 then the image is associated with that label.
# +
from keras.utils import to_categorical
train_labels = to_categorical(orig_train_labels)
test_labels = to_categorical(orig_test_labels)
print(f'The shape of the original train labels: {orig_train_labels.shape}')
print(f'The shape of the actual train labels: {train_labels.shape}')
print(f'The shape of the original test labels: {orig_test_labels.shape}')
print(f'The shape of the actual test labels: {test_labels.shape}')
print(f'First original train label: {orig_train_labels[0]}')
print(f'First actual train label: {train_labels[0]}')
print(f'First original test label: {orig_test_labels[0]}')
print(f'First actual test label: {test_labels[0]}')
# -
# ## Define and train the network
#
# We use a sequential network. The input shape for the first layer must corresponds to the shape of the images (28*28 1D tensors). Once the network layers have been created we define the optimizer and loss function using the compile function. Finally, we train the network using the fit function and print out the loss and accuracy of the model.
# +
import math
from keras import models
from keras import layers
epochs = 7
network = models.Sequential()
network.add(layers.Conv2D(32, (3, 3), activation='relu', input_shape=(28, 28, 1)))
network.add(layers.MaxPooling2D(2, 2))
network.add(layers.Conv2D(64, (3, 3), activation='relu'))
network.add(layers.MaxPooling2D(2, 2))
network.add(layers.Conv2D(64, (3, 3), activation='relu'))
network.add(layers.Flatten())
network.add(layers.Dense(64, activation='relu'))
network.add(layers.Dense(10, activation='softmax'))
network.summary()
network.compile(optimizer='rmsprop', loss='categorical_crossentropy', metrics=['accuracy'])
history = network.fit_generator(gen_train_images.flow(train_images, train_labels, batch_size=32),
steps_per_epoch=len(train_images) / 32, epochs=epochs, validation_data=(test_images, test_labels))
loss, accuracy = network.evaluate(test_images, test_labels)
print(f'Loss: {loss}')
print(f'Accuracy: {accuracy}')
# -
# ## Model overfitting
#
# There is very little evidence of overfitting in our model.
# +
from matplotlib import pyplot
pyplot.rcParams["figure.figsize"] = (5,5)
train_accuracy = history.history['acc']
validation_accuracy = history.history['val_acc']
train_loss = history.history['loss']
validation_loss = history.history['val_loss']
epocs = range(1, len(train_loss) + 1)
pyplot.plot(epocs, train_accuracy, 'bo', label='Training accuracy')
pyplot.plot(epocs, validation_accuracy, 'b', label='Validation accuracy')
pyplot.title('Training and validation accuracy')
pyplot.xlabel('Epochs')
pyplot.xlabel('Accuracy')
pyplot.legend()
pyplot.show()
pyplot.plot(epocs, train_loss, 'bo', label='Training loss')
pyplot.plot(epocs, validation_loss, 'b', label='Validation loss')
pyplot.title('Training and validation loss')
pyplot.xlabel('Epochs')
pyplot.xlabel('Loss')
pyplot.legend()
pyplot.show()
# -
# ## Invalid prediction counts
#
# Lets show how may times each digit in the test set was incorrectly classified as one of the other digits.
# +
import numpy
from matplotlib import pyplot
predicted_labels = network.predict(test_images)
## Find invalid predictions.
invalid_prediction = []
invalid_prediction_cnt = numpy.zeros((10,10), dtype=int)
for i,v in enumerate(predicted_labels):
test_label = numpy.where(test_labels[i] == numpy.amax(test_labels[i]))[0].item(0)
predicted_test_label = numpy.where(predicted_labels[i] == numpy.amax(predicted_labels[i]))[0].item(0)
if test_label != predicted_test_label:
invalid_prediction_cnt[test_label][predicted_test_label] += 1
invalid_prediction.append((i, test_label, predicted_test_label))
total_invalid_prediction_cnt = numpy.sum(invalid_prediction_cnt)
## Plot invalid prediction counts.
x_test_labels = numpy.arange(10)
y_predicted_labels = numpy.arange(10)
fig, ax = pyplot.subplots(figsize=(8,8))
ax.imshow(invalid_prediction_cnt, cmap="coolwarm")
ax.set_title(f'Invalid predictions ({total_invalid_prediction_cnt})')
ax.set_xticks(x_test_labels)
ax.set_yticks(y_predicted_labels)
ax.set_xlabel('Test label')
ax.set_ylabel('Predicted label')
ax.invert_yaxis()
for i in range(len(x_test_labels)):
for j in range(len(y_predicted_labels)):
# Show number of invalid predictions.
text = ax.text(j, i, invalid_prediction_cnt[i, j],
ha="center", va="center", color="w")
fig.tight_layout()
pyplot.show()
# -
# ## Invalid predictions
#
# Finally lets show all the images in the test set that were categorized incorrectly.
# +
pyplot.rcParams["figure.figsize"] = (30,30)
ncols = 10
nrows = math.ceil(total_invalid_prediction_cnt / ncols)
for i in range(total_invalid_prediction_cnt):
idx, test_label, predicted_test_label = invalid_prediction[i]
row = math.floor(i / ncols)
col = i % ncols
ax = pyplot.subplot2grid((nrows,ncols),(row,col))
ax.axis('off')
ax.set_title(f'{test_label}->{predicted_test_label}')
ax.imshow(orig_test_images[idx], cmap="gray")
pyplot.show()
# -
| mnist_conv_gen.ipynb |
/ ---
/ jupyter:
/ jupytext:
/ text_representation:
/ extension: .q
/ format_name: light
/ format_version: '1.5'
/ jupytext_version: 1.14.4
/ kernelspec:
/ display_name: SQL
/ language: sql
/ name: SQL
/ ---
/ + [markdown] azdata_cell_guid="b2c71178-6846-423d-a1fa-f864fab7c58b"
/ INSTALL DEPENDENCIES
/ + azdata_cell_guid="a6b17618-9951-4cf0-aca9-39e39f3020bf"
!pip install pandas
!pip install scikit-learn
!pip install numpy
!pip install onnxmltools
!pip install onnxruntime
!pip install pyodbc
!pip install setuptools
!pip install skl2onnx
!pip install sqlalchemy
/ + [markdown] azdata_cell_guid="c455a0cb-048b-405d-ae6d-0095fae75161"
/ SET VALUES
/ + azdata_cell_guid="7c6e9aa7-c080-4e58-95bc-e846aab39984"
server = 'iotedgevm-be9601db-be9601db.eastus.cloudapp.azure.com' # SQL Server IP address
username = 'sa' # SQL Server username
password = '<PASSWORD>!' # SQL Server password
database = 'IoTEdgeDB'
data_table = 'OpcNodes'
models_table = 'Models'
features_table = 'Features'
target_table = 'Target'
master_connection_string = "Driver={ODBC Driver 17 for SQL Server};Server=" + server + ";Database=master;UID=" + username + ";PWD=" + password + ";"
db_connection_string = "Driver={ODBC Driver 17 for SQL Server};Server=" + server + ";Database=" + database + ";UID=" + username + ";PWD=" + password + ";"
/ + [markdown] azdata_cell_guid="bf2ef7fe-a9f0-479b-8e03-e34fed0a75df"
/ CREATE X TRAIN DATASET
/ + azdata_cell_guid="f52cbb89-8ac4-408d-b74c-2160b5691dc0"
import numpy as np
import onnxmltools
import onnxruntime as rt
import pandas as pd
import skl2onnx
import sklearn
import sklearn.datasets
import pyodbc
conn = pyodbc.connect(db_connection_string, autocommit=True)
query = 'SELECT ApplicationUri, DipData, SpikeData, RandomSignedInt32 FROM OpcNodes WHERE DipData IS NOT NULL AND SpikeData IS NOT NULL AND RandomSignedInt32 IS NOT NULL'
sql_query = pd.read_sql_query(query, conn)
x_train = pd.DataFrame(sql_query, columns=['DipData', 'SpikeData', 'RandomSignedInt32'])
print("\n*** Training dataset x\n")
print(x_train.head())
/ + [markdown] azdata_cell_guid="7f2af60d-bccd-494f-951a-1cc1e24f9c6a"
/ CREATE Y TRAIN DATASET
/ + azdata_cell_guid="297d66f3-8f21-409d-8068-b285c9425efe"
target_column = 'MEDV'
y_train = pd.DataFrame()
y_train[target_column] = np.random.randint(100, size=(len(x_train)))
print("\n*** Training dataset y\n")
print(y_train.head())
/ + [markdown] azdata_cell_guid="593b8da3-a05e-40d8-85b6-cc8ee56d2d95"
/ CREATE PIPELINE TO TRAIN THE LINEAR REGRESSION MODEL
/ + azdata_cell_guid="1220be84-0b79-4993-a151-b8a045ef2cfe"
from sklearn.compose import ColumnTransformer
from sklearn.linear_model import LinearRegression
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import RobustScaler
continuous_transformer = Pipeline(steps=[('scaler', RobustScaler())])
# All columns are numeric - normalize them
preprocessor = ColumnTransformer(
transformers=[
('continuous', continuous_transformer, [i for i in range(len(x_train.columns))])])
model = Pipeline(
steps=[
('preprocessor', preprocessor),
('regressor', LinearRegression())])
# Train the model
model.fit(x_train, y_train)
/ + [markdown] azdata_cell_guid="380624de-b936-4ecf-bee7-d5ba1d4d9659"
/ CHECK MODEL ACCURACY
/ + azdata_cell_guid="a628102e-fd8f-43d4-9723-444caa09cedf"
# Score the model
from sklearn.metrics import r2_score, mean_squared_error
y_pred = model.predict(x_train)
sklearn_r2_score = r2_score(y_train, y_pred)
sklearn_mse = mean_squared_error(y_train, y_pred)
print('*** Scikit-learn r2 score: {}'.format(sklearn_r2_score))
print('*** Scikit-learn MSE: {}'.format(sklearn_mse))
/ + [markdown] azdata_cell_guid="b83c76fe-ad20-43ca-af38-163c3750965d"
/ DEFINE FUNCTION TO CONVERT MODEL TO ONNX
/ + azdata_cell_guid="66d907db-56bf-4dcd-b742-cf2c9105d313"
from skl2onnx.common.data_types import FloatTensorType, Int64TensorType, DoubleTensorType
def convert_dataframe_schema(df, drop=None, batch_axis=False):
inputs = []
nrows = None if batch_axis else 1
for k, v in zip(df.columns, df.dtypes):
if drop is not None and k in drop:
continue
if v == 'int64':
t = Int64TensorType([nrows, 1])
elif v == 'float32':
t = FloatTensorType([nrows, 1])
elif v == 'float64':
t = DoubleTensorType([nrows, 1])
else:
raise Exception("Bad type")
inputs.append((k, t))
return inputs
/ + [markdown] azdata_cell_guid="2a9fc89a-b24d-47e1-8379-4aefd83b0397"
/ SAVE MODEL IN ONNX FORMAT
/ + azdata_cell_guid="95a37d5d-4fca-44b5-ab4a-757debe1a414"
# Convert the scikit model to onnx format
onnx_model = skl2onnx.convert_sklearn(model, 'OPC tags data', convert_dataframe_schema(x_train), final_types=[('variable1',FloatTensorType([1,1]))], target_opset=11)
# Save the onnx model locally
onnx_model_path = 'opcnodes.model.onnx'
onnxmltools.utils.save_model(onnx_model, onnx_model_path)
/ + [markdown] azdata_cell_guid="e7b75cf7-8a94-42b9-b012-beb0666fbe55"
/ TEST ONNX MODEL
/ + azdata_cell_guid="1ba02875-cb66-489c-9341-502e853be0f5"
import onnxruntime as rt
sess = rt.InferenceSession(onnx_model_path)
y_pred = np.full(shape=(len(x_train)), fill_value=np.nan)
for i in range(len(x_train)):
inputs = {}
for j in range(len(x_train.columns)):
inputs[x_train.columns[j]] = np.full(shape=(1,1), fill_value=x_train.iloc[i,j])
sess_pred = sess.run(None, inputs)
y_pred[i] = sess_pred[0][0][0]
onnx_r2_score = r2_score(y_train, y_pred)
onnx_mse = mean_squared_error(y_train, y_pred)
print()
print('*** Onnx r2 score: {}'.format(onnx_r2_score))
print('*** Onnx MSE: {}\n'.format(onnx_mse))
print('R2 Scores are equal' if sklearn_r2_score == onnx_r2_score else 'Difference in R2 scores: {}'.format(abs(sklearn_r2_score - onnx_r2_score)))
print('MSE are equal' if sklearn_mse == onnx_mse else 'Difference in MSE scores: {}'.format(abs(sklearn_mse - onnx_mse)))
print()
/ + [markdown] azdata_cell_guid="f1afba4f-18ea-4cfa-98c7-50cac1aad6f0"
/ <u>INSERT MODEL IN DATABASE</u>
/ + azdata_cell_guid="7903a3ec-b5c7-45e8-8d2b-3d4a93d0385a"
import pyodbc
# Insert the ONNX model into the models table for each value of ApplicationUri
query = f'SELECT DISTINCT ApplicationUri FROM {data_table}'
cursor = conn.cursor()
cursor.execute(query)
application_uris = cursor.fetchall()
for app in application_uris:
query = f"insert into {models_table} ([applicationUri], [description], [data]) values ('{app[0]}', 'Onnx Model', ?)"
model_bits = onnx_model.SerializeToString()
insert_params = (pyodbc.Binary(model_bits))
cursor.execute(query, insert_params)
conn.commit()
/ + [markdown] azdata_cell_guid="2beb33c4-1ab6-482e-91af-9e1e2ba331ff"
/ ENSURE MODEL HAS BEEN STORED IN TABLE
/ + azdata_cell_guid="061a59f9-69b2-499b-8131-4d355766e9f8"
conn = pyodbc.connect(db_connection_string, autocommit=True)
cursor = conn.cursor()
query = f'SELECT * FROM {models_table}'
cursor.execute(query)
for row in cursor.fetchall():
print(row)
/ + [markdown] azdata_cell_guid="38e9f5f6-2b9b-490e-8177-fe5b654c5211"
/ CREATE FEATURES AND TARGET TABLES
/ + azdata_cell_guid="3fe85731-6acf-48a7-9b7a-23961c76482c"
import sqlalchemy
from sqlalchemy import create_engine
import urllib
conn = pyodbc.connect(db_connection_string)
cursor = conn.cursor()
features_table_name = 'features'
# Drop the table if it exists
query = f'drop table if exists {features_table_name}'
cursor.execute(query)
conn.commit()
# Create the features table
query = \
f'create table {features_table_name} ( ' \
f' [CRIM] float, ' \
f' [ZN] float, ' \
f' [INDUS] float, ' \
f' [CHAS] float, ' \
f' [NOX] float, ' \
f' [RM] float, ' \
f' [AGE] float, ' \
f' [DIS] float, ' \
f' [RAD] float, ' \
f' [TAX] float, ' \
f' [PTRATIO] float, ' \
f' [B] float, ' \
f' [LSTAT] float, ' \
f' [id] int)'
cursor.execute(query)
conn.commit()
target_table_name = 'target'
query = f'drop table if exists {target_table_name}'
cursor.execute(query)
conn.commit()
# Create the target table
query = \
f'create table {target_table_name} ( ' \
f' [MEDV] float, ' \
f' [id] int)'
cursor.execute(query)
conn.commit()
x_train['id'] = range(1, len(x_train)+1)
y_train['id'] = range(1, len(y_train)+1)
print(x_train.head())
print(y_train.head())
/ + [markdown] azdata_cell_guid="45ce265f-81b9-4eac-9e4a-1b2483e8b69d"
/ INSERT FEATURES & TARGET DATASETS INTO THE DATABASE
/ + azdata_cell_guid="3853118d-cfc0-4876-9f58-aeccd1c61d74"
cursor = conn.cursor()
# for index, row in x_train.iterrows():
# # print(index)
# cursor.execute(f'INSERT INTO {features_table_name} (CRIM,ZN,INDUS,CHAS,NOX,RM,AGE,DIS,RAD,TAX,PTRATIO,B,LSTAT,id) VALUES ({row.CRIM},{row.ZN},{row.INDUS},{row.CHAS},{row.NOX},{row.RM},{row.AGE},{row.DIS},{row.RAD},{row.TAX},{row.PTRATIO},{row.B},{row.LSTAT},{row.id})')
# conn.commit()
for index, row in y_train.iterrows():
# print(index)
cursor.execute(f'INSERT INTO {target_table_name} (MEDV,id) VALUES ({row.MEDV},{row.id})')
conn.commit()
cursor.close()
/ + azdata_cell_guid="4a6c5c3c-87bd-4a1e-b08d-52981cbb8dce"
RUN PREDICT STATEMENT (SWITCH TO SQL KERNEL)
/ + azdata_cell_guid="a553facc-ef3d-4410-be76-8935c5c03972"
USE IoTEdgeDB
DECLARE @model VARBINARY(max) = (
SELECT DATA
FROM dbo.Models
WHERE applicationUri = 'urn:OpcPlc:opcserver1'
);
WITH predict_input
AS (
SELECT TOP (1000) [id]
, DipData
, SpikeData
, RandomSignedInt32
FROM [dbo].[OpcNodes]
)
SELECT predict_input.id
, p.variable1 AS MEDV
FROM PREDICT(MODEL = @model, DATA = predict_input, RUNTIME=ONNX) WITH (variable1 FLOAT) AS p;
| ML/Notebooks/train-opc-nodes-model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.2.0
# language: julia
# name: julia-1.2
# ---
# ## Construct a DataFrame
# +
using DataFrames
city = ["Corvallis", "Portland", "Eugene"]
pop = [57961, 647805, 168916]
df_cities = DataFrame(city=city, pop=pop) # cols are symbols
df_cities
# -
push!(df_cities, ["Bend", 94520])
df_cities
# ### Retreive a column
df_cities.pop
# ### Add Columns
df_cities[!, :state] = ["OR", "OR", "OR", "OR"] # ! = all rows
df_cities
size(df_cities)
# ### Delete row
deleterows!(df_cities, 3)
df_cities
# ## Iterate through the rows of a data frame
for row in eachrow(df_cities)
# row is like a dictionary
println("city = ", row[:city])
println("\tpop = ", row[:pop])
println("\tstate = ", row[:state])
end
| CHE599-IntroDataScience/lectures/dataframes/dataframes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: drlnd
# language: python
# name: drlnd
# ---
# # Collaboration and Competition
#
# ---
#
# In this notebook, you will learn how to use the Unity ML-Agents environment for the third project of the [Deep Reinforcement Learning Nanodegree](https://www.udacity.com/course/deep-reinforcement-learning-nanodegree--nd893) program.
#
# ### 1. Start the Environment
#
# We begin by importing the necessary packages. If the code cell below returns an error, please revisit the project instructions to double-check that you have installed [Unity ML-Agents](https://github.com/Unity-Technologies/ml-agents/blob/master/docs/Installation.md) and [NumPy](http://www.numpy.org/).
from Unity_Env_Wrapper import TennisEnv
from buffer import ReplayBuffer
from maddpg import MADDPG
import torch
import numpy as np
import os
from collections import deque
from ddpg import DDPGAgent
import torch
import torch.nn.functional as F
# Environments contain **_brains_** which are responsible for deciding the actions of their associated agents. Here we check for the first brain available, and set it as the default brain we will be controlling from Python.
main_tennis()
# +
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(1, len(scores)+1), scores)
plt.ylabel('Score')
plt.xlabel('Episode #')
# -
# When finished, you can close the environment.
| p3_collab-compet/Tennis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# https://jakevdp.github.io/blog/2017/12/05/installing-python-packages-from-jupyter/
from IPython.core.magic import register_line_magic
@register_line_magic
def pip(args):
"""Use pip from the current kernel"""
from pip import main
main(args.split())
# -
# %pip install matplotlib
import matplotlib.pyplot as plt
# +
plt.plot([1,2,3],[1,2,5], label="This is label")
plt.xlabel("Project Duration")
plt.ylabel("Cost")
plt.title("Nya")
ax = plt.gca()
ax.axes.xaxis.set_ticks([])
ax.axes.yaxis.set_ticks([])
plt.show()
| _posts/_media_working_copies/IDesign/Charts-Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/cascadet/PopGrid-SDG11.5-Tutorial/blob/main/1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="xE0t2RRpOHqG"
# # Cliping Step for Raster Based on Imported Country Shapefile
#
# # By <NAME>, Sep 2020 <br>
# # Notebook to clip rasters. <br>
# # **NOTE** Needs to be run for all geographies: Egypt forEGY.shp, Nepal gdam 0, M-M-Z0.shp, and ECU-clip0.shp
#
# + id="gy1msD_pJCyT"
# first step is to install some libraries 1. install rioxarray
# + colab={"base_uri": "https://localhost:8080/"} id="wnxoiPrdZF__" outputId="eb574361-68ba-47fb-dcff-2d4c7df7fa38"
pip install rioxarray
# + colab={"base_uri": "https://localhost:8080/"} id="bXnb0pRFBVSe" outputId="76deb1a7-91a5-4150-e069-0612c59ab9cb"
pip install geopandas
# + id="6RLPNmrGiKmL"
#### Dependencies
import numpy as np
import pandas as pd
import rasterio
import xarray as xr
import geopandas as gpd
import glob
import rioxarray as rio
from scipy.stats import variation
import rasterio.mask
# + id="zvU8PwHlOPfC"
#### File Paths & FNs
DATA_PATH = '/content/drive/MyDrive/PopGrid-Compare/'
DATA_PATH_OUT = '/content/drive/MyDrive/pop/'
# + id="eNW80iJYOR6_"
def raster_clip(rst_fn, polys, in_ext, out_ext):
""" function clips a raster and saves it out
args:
rst_fn = raster you want to clip
polys = polys you want to clip to
in_ext = tail of .tif file to clip off
out_ext = tail of .tif file for saving out
"""
# Get raster name
#rst_nm = rst_fn.split('interim\\')[1].split('.tif')[0]
#data = rst_fn.split(DATA_PATH+'interim/')[1].split(in_ext)[0]
data = rst_fn.split(DATA_PATH+'interim/')[1].split(in_ext)[0]
fn_out = DATA_PATH_OUT+'interim/'+data+out_ext
print(fn_out)
# clip raster
with rasterio.open(rst_fn) as src:
out_image, out_transform = rasterio.mask.mask(src, polys, crop=True)
out_meta = src.meta
# Update meta data
out_meta.update({"driver": "GTiff",
"height": out_image.shape[1],
"width": out_image.shape[2],
"transform": out_transform})
# write image
with rasterio.open(fn_out, "w", **out_meta) as dest:
dest.write(out_image)
# + colab={"base_uri": "https://localhost:8080/"} id="BSh0DeG8OUHP" outputId="e83e7e46-bdcb-4117-85af-95446e54e400"
#### Matched Rasters
rst_fns = glob.glob('/content/drive/MyDrive/PopGrid-Compare/interim/*_matched.tif')
rst_fns
# Open polys
country = '_EGY.tif'#'_NPL.tif'#'_ECU.tif'#'_MMZ.tif'#'_NPL.tif'
polys_fn = DATA_PATH_OUT+ 'interim/EGY.shp'#'interim/chile_quake_pop.shp' #'interim/M-M-Z-0.shp'
#hazem-polys_fn = DATA_PATH+ 'interim/M-M-Z-0.shp'#gwpv4_npl_admin4.shp'#'interim/ECU-clip0.shp' #'interim/M-M-Z-0.shp'
polys = gpd.read_file(polys_fn)
shapes = polys["geometry"]
in_ext = '_matched.tif'
out_ext = '_all'+country
for rst_fn in rst_fns:
raster_clip(rst_fn, shapes, in_ext, out_ext)
# + id="soaUxUkHOWEP"
#### Urban Rasters
rst_fns = glob.glob('/content/drive/MyDrive/PopGrid-Compare/interim/*_urban.tif')
rst_fns
in_ext = 'matched_urban.tif'
out_ext = 'urban'+country
for rst_fn in rst_fns:
raster_clip(rst_fn, shapes, in_ext, out_ext)
# + colab={"base_uri": "https://localhost:8080/"} id="hnUy-FbpOYQR" outputId="addd7016-7d87-457a-ecd6-b8b87785e7df"
#### Rural Rasters
rst_fns = glob.glob('/content/drive/MyDrive/pop/interim/*_rural.tif')
rst_fns
in_ext = 'matched_rural.tif'
out_ext = 'rural'+country
for rst_fn in rst_fns:
raster_clip(rst_fn, shapes, in_ext, out_ext)
# + colab={"base_uri": "https://localhost:8080/"} id="sp-ZtW_Cjuad" outputId="be2a585b-cf39-4b2b-f8ee-5039d0582a62"
#### Rural Rasters
rst_fns = glob.glob('/content/drive/MyDrive/pop/interim/*_rural.tif')
rst_fns
in_ext = 'matched_rural.tif'
out_ext = 'rural'+country
for rst_fn in rst_fns:
raster_clip(rst_fn, shapes, in_ext, out_ext)
# + colab={"base_uri": "https://localhost:8080/"} id="XUE32Q_9OaGI" outputId="353cc750-bd83-4516-cdad-a57aae60a858"
#### Urban Rasters
rst_fns = glob.glob('/content/drive/MyDrive/pop/interim/*_urban.tif')
rst_fns
in_ext = 'matched_urban.tif'
out_ext = 'urban'+country
for rst_fn in rst_fns:
raster_clip(rst_fn, shapes, in_ext, out_ext)
# + id="LJaZgOsKzUlf"
## Quake
# By <NAME>, June 2020
# Notebook finds zonal stats of populations for 2015 Nepal earthquakes by raster. <br><br>
# **NOTE** CRS should be epsg:4326 for everything!
# + colab={"base_uri": "https://localhost:8080/"} id="u4MP1amFjfEx" outputId="8ea80798-6179-482e-8f91-40d1c981f68a"
pip install rasterstats
# + id="i9qpq9WKj-HC"
#### Dependencies
import numpy as np
import pandas as pd
import rasterio
import geopandas as gpd
from rasterstats import zonal_stats, gen_zonal_stats
from glob import glob
from matplotlib import pyplot as plt
import matplotlib.patches as patches
Patch = patches.Patch
# + id="4Ptx8t-HkAcO"
#### Functions
def zone_loop(polys_in, rst_list, stats_type, col, split):
""" Function loops through rasters, calcs zonal_stats and returns stats as a data frame.
Args:
polys_in = polygons
rst_list = list of paths & fns of rasters
stats_type = stats type for each poly gone (see zonal stats)
col = column to merge it all
split = where to split the file name string (e.g. _matched.tif)
"""
# copy polys to write out
polys_out = polys_in.copy()
for rst in rst_list:
# Get data name
data = rst.split(DATA_PATH+'interim/')[1].split(split)[0]
print('Started', data)
# Run zonal stats
zs_feats = zonal_stats(polys_in, rst, stats=stats_type, geojson_out=True)
zgdf = gpd.GeoDataFrame.from_features(zs_feats, crs=polys_in.crs)
# Rename columns and merge
zgdf = zgdf.rename(columns={stats_type: data+'_'+stats_type})
polys_out = polys_out.merge(zgdf[[col, data+'_'+stats_type]], on = col, how = 'inner')
return polys_out
# + id="K3Ez_EBykDLh"
def poly_prep(polys_fn, col):
"function opens earth quake polygons for zonal loop"
# open
polys = gpd.read_file(polys_fn)
# subset, be sure to check the admin level
polys = polys[['geometry', col]]
return polys
# + id="glwr137Y04Wq"
## Run on Shakemap Intensity Contours (MI) from USGS for this example: Nepal 2015
# + id="IhTFg9gw1GkX"
#### All
# + colab={"base_uri": "https://localhost:8080/"} id="Bhx7f9c0kInO" outputId="29bb5400-6d23-4d43-e687-0964cd2d1b3d"
# open polys
DATA_PATH = '/content/drive/MyDrive/pop/'
nepal_polys_fn = DATA_PATH+'interim/shakemap/mi.shp'
col = 'PARAMVALUE'
nepal_polys = poly_prep(nepal_polys_fn, col)
print (nepal_polys)
# + colab={"base_uri": "https://localhost:8080/"} id="qpNWPWKHkLy-" outputId="d9fc64de-be61-4373-85ae-b8fd441896ec"
# Git tif files
rst_fns = sorted(glob(DATA_PATH+'interim/*all_EGY.tif'))
rst_fns
# + colab={"base_uri": "https://localhost:8080/"} id="nIL2IicikNoO" outputId="6ea4321c-0878-4ac4-bbc8-f95c99b4ef96"
# Run zonal stats loop
nepal_polys_sum = zone_loop(nepal_polys, rst_fns, 'sum', col, '_all_EGY.tif')
# + id="FXKEbBFRkPST"
# Save the poly sums
nepal_polys_sum.to_file(DATA_PATH+'/interim/EGY_quake_pop.shp')
# + id="SaKqQtBk1Mj8"
#### Urban
# + id="K0DxT3BFkQv5"
# open polys
DATA_PATH = '/content/drive/MyDrive/pop/'
nepal_polys_fn = DATA_PATH+'interim/shakemap/mi.shp'
col = 'PARAMVALUE'
nepal_polys = poly_prep(nepal_polys_fn, col)
# + colab={"base_uri": "https://localhost:8080/"} id="1uH5U4JKkSTY" outputId="a5f3c3e8-5709-476c-9587-00f86465dde8"
# Git tif files
rst_fns = sorted(glob(DATA_PATH+'interim/*_urban_EGY.tif'))
rst_fns
# + colab={"base_uri": "https://localhost:8080/"} id="G7autbqDkT-a" outputId="be3fdfc4-46b5-43e5-862c-b634649e7660"
# Run zonal stats loop
nepal_polys_sum = zone_loop(nepal_polys, rst_fns, 'sum', col, '_urban_EGY.tif')
# + id="0WqARnEmkVfr"
#### Save the poly sums
nepal_polys_sum.to_file(DATA_PATH+'/interim/EGY_urban_quake_pop.shp')
# + id="yp7QSikm1Pt-"
#### Rural
# + id="No7g2-D_kW1y"
# open polys
DATA_PATH = '/content/drive/MyDrive/pop/'
nepal_polys_fn = DATA_PATH+'interim/shakemap/mi.shp'
col = 'PARAMVALUE'
nepal_polys = poly_prep(nepal_polys_fn, col)
# + colab={"base_uri": "https://localhost:8080/"} id="Xp3VY-2okYUx" outputId="3a4a491a-44c0-4290-87cc-1f8d83d61339"
# Git tif files
rst_fns = glob(DATA_PATH+'interim/*_rural_EGY.tif')
rst_fns
# + colab={"base_uri": "https://localhost:8080/"} id="bKJpW-tUkaKI" outputId="9674f7de-8c41-4fc2-c76b-c533b795470a"
# Run zonal stats loop
nepal_polys_sum = zone_loop(nepal_polys, rst_fns, 'sum', col, '_rural_EGY.tif')
# + id="0WSCvEyzkbzO"
#### Save the poly sums
nepal_polys_sum.to_file(DATA_PATH+'/interim/EGY_rural_quake_pop.shp')
# + id="qa9btLzQ1UtT"
# Check data
# + colab={"base_uri": "https://localhost:8080/", "height": 700} id="Sp_P0pTqkczM" outputId="951ff5ae-c5c3-46b5-9acd-ea6a48d52e6f"
# All
fn_in = DATA_PATH+'/interim/EGY_quake_pop.shp'
all_pop = gpd.read_file(fn_in)
all_pop
# + colab={"base_uri": "https://localhost:8080/", "height": 700} id="hEUYl2ttkd_W" outputId="8e07bb0b-03fe-4f53-9869-59b05413f7fb"
# All
fn_in = DATA_PATH+'/interim/EGY_quake_pop.shp'
all_pop = gpd.read_file(fn_in)
all_pop
# + colab={"base_uri": "https://localhost:8080/", "height": 700} id="DRTyU-Ezkndr" outputId="711bf5ab-825d-4fdf-91d8-e18b9f8e9ff6"
# Urban
fn_in = DATA_PATH+'/interim/EGY_urban_quake_pop.shp'
urban_pop = gpd.read_file(fn_in)
urban_pop
# + colab={"base_uri": "https://localhost:8080/", "height": 700} id="6S3XtwD5korC" outputId="901badf3-c102-4033-80f2-8e7f97099345"
# Urban
fn_in = DATA_PATH+'/interim/EGY_rural_quake_pop.shp'
rural_pop = gpd.read_file(fn_in)
rural_pop
# + id="OK2ROYvhkp7t"
x=urban_pop.iloc[:,1:6]
y=urban_pop.iloc[:,1:6]
z=all_pop.iloc[:,1:6]
# zz=x+y/z
# + id="9wcDHfyf1h7s"
#### Check that rural + urban = total
# + id="fxRmmuYtkrC3"
# Check the data
# (rural_pop.iloc[:,1:6] + urban_pop.iloc[:,1:6]) / all_pop.iloc[:,1:6]
# + id="xtNRaVqK1nK8"
# Final Plots
# + id="L1FDuPAjAnF9"
#### set colors
ESRI16_c = 'blue'
GHS15_c = 'indigo'
GPWv4_c = 'deeppink'
LS15_c = 'deepskyblue'
WP16_c = 'forestgreen'
# + id="Ej2bsdRNApjY"
npl_all_fn = DATA_PATH+'interim/EGY_quake_pop.shp'
npl_all = gpd.read_file(npl_all_fn)
# + id="KViDG-lOArMp"
npl_rural_fn = DATA_PATH+'interim/EGY_rural_quake_pop.shp'
npl_rural = gpd.read_file(npl_rural_fn)
# + id="ItwHzgbgAs4G"
npl_urban_fn = DATA_PATH+'interim/EGY_urban_quake_pop.shp'
npl_urban = gpd.read_file(npl_urban_fn)
# + id="ezxMoMWWAvM-"
test_a = npl_all[(npl_all['PARAMVALUE'] >= 4) & (npl_all['PARAMVALUE'] < 5)].iloc[:,1:6].sum(axis = 0)
# + id="NeWNcNkOAvO4"
test_b = npl_rural[(npl_rural['PARAMVALUE'] >= 4) & (npl_rural['PARAMVALUE'] < 5)].iloc[:,1:6].sum(axis = 0)
# + id="DgOih-ARAvRy"
test_c = npl_urban[(npl_urban['PARAMVALUE'] >= 4) & (npl_urban['PARAMVALUE'] < 5)].iloc[:,1:6].sum(axis = 0)
# + colab={"base_uri": "https://localhost:8080/"} id="TqbMv-xOAvUe" outputId="5c74e0d3-2c56-4766-97d6-7c9f12b33ee5"
(test_b + test_c) / test_a
# + id="84gEu5GiAvWz"
# Make data
def group(df):
" Group and sum population by MI ranges, args is df quake pop"
iv = df[(df['PARAMVALUE'] >= 4) & (df['PARAMVALUE'] < 5)].iloc[:,1:6].sum(axis = 0)
v = df[(df['PARAMVALUE'] >= 5) & (df['PARAMVALUE'] < 6)].iloc[:,1:6].sum(axis = 0)
vi = df[(df['PARAMVALUE'] >= 6) & (df['PARAMVALUE'] < 7)].iloc[:,1:6].sum(axis = 0)
vii = df[df['PARAMVALUE'] >= 7].iloc[:,1:6].sum(axis = 0)
out = pd.DataFrame()
out['iv'] = iv
out['v'] = v
out['vi'] = vi
out['vii'] = vii
out = out.transpose()
return out
# + id="Rr9s33duAvYw" colab={"base_uri": "https://localhost:8080/", "height": 173} outputId="368c8cea-c6ee-406e-93e4-31c49d73bde3"
# Double check urban + rural / total = 100
all_g = group(npl_all)
rural_g = group(npl_rural)
urban_g = group(npl_urban)
(rural_g+urban_g) / all_g
# + colab={"base_uri": "https://localhost:8080/", "height": 528} id="kUwQH3HqA2CJ" outputId="8f8fb06a-220d-42b7-cc94-1e5cf3c1c917"
# Make bar plot
fig, axs = plt.subplots(1, 1, figsize = (12, 8), sharex=True)
ws = 0.25
fig.subplots_adjust(wspace=ws)
scale = 10**6
# All Quake
data = group(npl_all)
# Bar locations
a = [1-.3,2-.3,3-.3, 4-.3]
b = [1-.15,2-.15,3-.15,4-.15]
c = [1,2,3,4]
d = [1+.15,2+.15,3+.15,4+.15]
e = [1+.3,2+.3,3+.3,4+.3]
# plots
# plt.bar(a, data.ESRI16_sum / scale, width=0.12, align='center', alpha = 0.5, color = ESRI16_c, ec = 'black')
# plt.bar(b, data.GHS15_sum / scale, width=0.12, align='center', alpha = 0.6, color = GHS15_c, ec = 'black')
plt.bar(c, data.GPWv4_sum / scale, width=0.12, align='center', alpha = 0.7, color = GPWv4_c, ec = 'black')
# plt.bar(d, data.LS15_sum / scale, width=0.12, align='center', alpha = 0.8, color = LS15_c, ec = 'black')
# plt.bar(e, data.WP16_sum / scale, width=0.12, align='center', alpha = 0.9, color = WP16_c, ec = 'black')
# Fake plot for rural hatch legend
# plt.bar(e, data.WP16_sum / scale, width=0.12, align='center', alpha = 0, color = 'white', ec = 'black',hatch = "///")
# rural floods
data = group(npl_rural)
# plt.bar(a, data.ESRI16_sum / scale, width=0.12, align='center', alpha = 0.5, color = ESRI16_c, ec = 'black', hatch = "///")
# plt.bar(b, data.GHS15_sum / scale, width=0.12, align='center', alpha = 0.6, color = GHS15_c, ec = 'black', hatch = "///")
plt.bar(c, data.GPWv4_sum / scale, width=0.12, align='center', alpha = 0.7, color = GPWv4_c, ec = 'black', hatch = "///")
# plt.bar(d, data.LS15_sum / scale, width=0.12, align='center', alpha = 0.8, color = LS15_c, ec = 'black', hatch = "///")
# plt.bar(e, data.WP16_sum / scale, width=0.12, align='center', alpha = 0.9, color = WP16_c, ec = 'black', hatch = "///")
# legend
legend_elements = [Patch(facecolor=ESRI16_c, alpha = 0.5, edgecolor=None, label='WPE-15'),
Patch(facecolor=GHS15_c, alpha = 0.6, edgecolor=None, label='GHSL-90'),
Patch(facecolor=GPWv4_c, alpha = 0.7, edgecolor=None, label='GPW-15'),
Patch(facecolor=LS15_c, alpha = 0.8, edgecolor=None, label='LS-15'),
Patch(facecolor= WP16_c, alpha = 0.9, edgecolor=None, label='WP-16'),
Patch(facecolor= 'white', alpha = 0.9, hatch = '///', edgecolor='black', label='rural pop')]
plt.legend(handles = legend_elements, bbox_to_anchor=(1, 1.02), loc='upper left', ncol=1, fontsize = 15);
# Labels / Titles
axs.set_title('Egypt 2015 Earthquake Simulation Impact', size = 20)
axs.set_xlabel('Instrumental Intesnity', fontsize = 15)
axs.set_ylabel('Total Population [millions]', fontsize = 15)
# Ticks
ticks_bar = ['>=4', ' >=5', '>=6', '>=7'];
plt.xticks([1,2,3,4], ticks_bar, fontsize = 15);
plt.yticks(fontsize = 15);
# save it out
fig_out = DATA_PATH+'FIGS/MS/Finalv1/Fig4.png'
# plt.savefig(fig_out, dpi = 300, facecolor = 'white', bbox_inches='tight')
| 1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Getting Started with pandas
import pandas as pd
from pandas import Series, DataFrame
import numpy as np
np.random.seed(12345) # fix a random seed, so results are reproducible.
import matplotlib.pyplot as plt
plt.rc('figure', figsize=(10, 6))
PREVIOUS_MAX_ROWS = pd.options.display.max_rows
pd.options.display.max_rows = 20
np.set_printoptions(precision=4, suppress=True)
# ## Introduction to pandas Data Structures
# ### Series
obj = pd.Series([4, 7, -5, 3])
obj
obj.values
obj.index # like range(4)
obj2 = pd.Series([4, 7, -5, 3], index=['d', 'b', 'a', 'c'])
obj2
obj2.index
obj2['a']
obj2['d'] = 6
obj2['d']
obj2[['c', 'a', 'd']]
obj2[obj2 > 3]
x = [-1.0, 2.5, 3]
#x > 1.1
obj2 * 2
obj2 / 2
np.exp(obj2)
np.exp(0)
'b' in obj2
x = ['red', 'blue']
'red' in x
'e' in obj2
# # from dictionary to Series
sdata = {'Ohio': 35000, 'Texas': 71000, 'Oregon': 16000, 'Utah': 5000}
obj3 = pd.Series(sdata)
obj3
states = ['Ohio', 'California', 'Oregon', 'Texas']
obj4 = pd.Series(sdata, index=states) #California leads a missing value
obj4
pd.isnull(obj4)
pd.notnull(obj4)
obj4.isnull()
print('\n-----obj3:\n', obj3)
print('\n-----obj4:\n', obj4)
print('\n-----obj3 + obj4:\n')
obj3 + obj4
obj4.name = 'population'
obj4.index.name = 'state'
print(obj4)
obj = pd.Series([4, 7, -5, 3])
obj.index = ['Bob', 'Steve', 'Jeff', 'Ryan']
obj
# ### DataFrame
# a dictionary with lists as values
data = {'state': ['Ohio', 'Ohio', 'Ohio', 'Nevada', 'Nevada', 'Nevada'],
'year': [2000, 2001, 2002, 2001, 2002, 2003],
'pop': [1.5, 1.7, 3.6, 2.4, 2.9, 3.2]}
# convert dictionary to dataframe.
frame = pd.DataFrame(data)
frame
frame.head()
frame.tail()
pd.DataFrame(data, columns=['year', 'state', 'pop'])
# # Second dataframe exaple, frame2
# +
# a dictionaruy
data = {'state': ['Ohio', 'Ohio', 'Ohio', 'Nevada', 'Nevada', 'Nevada'],
'year': [2000, 2001, 2002, 2001, 2002, 2003],
'pop': [1.5, 1.7, 3.6, 2.4, 2.9, 3.2]}
frame2 = pd.DataFrame(data, columns=['year', 'state', 'pop', 'debt'], # reorder columns
index=['one', 'two', 'three', 'four',
'five', 'six'])
print(frame2)
frame2.columns
# -
frame2['state']
frame2.year
frame2['year']
frame2.loc['three']
frame2.iloc[2]
frame2['debt'] = 16.5
print(frame2)
frame2['debt'] = np.arange(6.)
print(frame2)
val = pd.Series([-1.2, -1.5, -1.7], index=['two', 'four', 'five'])
frame2['debt'] = val
frame2
frame2['eastern'] = frame2.state == 'Ohio'
frame2
del frame2['eastern']
frame2.columns
frame2
# # Third example, frame3
pop = {'Nevada': {2001: 2.4, 2002: 2.9},
'Ohio': {2000: 1.5, 2001: 1.7, 2002: 3.6}} #ummatched dictionaries as input
frame3 = pd.DataFrame(pop)
frame3
frame3.T #transpose
frame3.T.T
pd.DataFrame(pop, index=[2001, 2002, 2003]) # reorder by rows
# +
pdata = {'Ohio': frame3['Ohio'][:-1],
'Nevada': frame3['Nevada'][:2]}
pd.DataFrame(pdata)
# +
pdata = {'Ohio': frame3['Ohio'][:3],
'Nevada': frame3['Nevada'][:3]}
pd.DataFrame(pdata)
# -
frame3.index.name = 'year'; frame3.columns.name = 'state'
frame3
frame3
frame3.values
frame2.values
# ### Index Objects
obj = pd.Series(range(3), index=['a', 'b', 'c'])
index = obj.index
index
index[1:]
index[1] = 'd' # TypeError
# # second obj2
labels = pd.Index(np.arange(3))
labels
obj2 = pd.Series([1.5, -2.5, 0], index=labels)
obj2
obj2.index is labels
frame3
frame3.columns
'Ohio' in frame3.columns
2003 in frame3.index
dup_labels = pd.Index(['foo', 'foo', 'bar', 'bar'])
dup_labels
# +
# We stop here Tue
# -
# ## Essential Functionality
# ### Reindexing
obj = pd.Series([4.5, 7.2, -5.3, 3.6], index=['d', 'b', 'a', 'c'])
obj
obj2 = obj.reindex(['a', 'b', 'c', 'd', 'e'])
obj2
obj3 = pd.Series(['blue', 'purple', 'yellow'], index=[0, 2, 4])
obj3
obj3.reindex(range(6), method='ffill')
frame = pd.DataFrame(np.arange(9).reshape((3, 3)),
index=['a', 'c', 'd'],
columns=['Ohio', 'Texas', 'California'])
frame
frame2 = frame.reindex(['a', 'b', 'c', 'd'])
frame2
states = ['Texas', 'Utah', 'California']
frame.reindex(columns=states)
frame.loc[['a', 'b', 'c', 'd'], states]
# ### Dropping Entries from an Axis
obj = pd.Series(np.arange(5.), index=['a', 'b', 'c', 'd', 'e'])
obj
new_obj = obj.drop('c')
new_obj
obj.drop(['d', 'c'])
data = pd.DataFrame(np.arange(16).reshape((4, 4)),
index=['Ohio', 'Colorado', 'Utah', 'New York'],
columns=['one', 'two', 'three', 'four'])
data
data.drop(['Colorado', 'Ohio'])
data.drop('two', axis=1)
data.drop(['two', 'four'], axis='columns')
obj.drop('c', inplace=True)
obj
# ### Indexing, Selection, and Filtering
obj = pd.Series(np.arange(4.), index=['a', 'b', 'c', 'd'])
print(obj)
obj['b']
obj[1]
obj[['b', 'a', 'd']]
obj[[1, 3]]
obj[obj < 2]
# +
obj[['b', 'a', 'd']]
# -
obj['b':'c']
obj['b':'c'] = 5
obj
data = pd.DataFrame(np.arange(16).reshape((4, 4)),
index=['Ohio', 'Colorado', 'Utah', 'New York'],
columns=['one', 'two', 'three', 'four'])
data
data['two']
data[['three', 'one']]
data[:2]
data[data['three'] > 5]
data < 5
data[data < 5] = 0
data
# #### Selection with loc and iloc
data.loc['Colorado', ['two', 'three']]
data.iloc[2, [3, 0, 1]]
data.iloc[2]
data.iloc[[1, 2], [3, 0, 1]]
data.loc[:'Utah', 'two']
data.iloc[:, :3]
data.iloc[:, :3][data.three > 9]
# ### Integer Indexes
ser = pd.Series(np.arange(3.))
ser
#ser[-1]
ser = pd.Series(np.arange(3.))
ser
ser2 = pd.Series(np.arange(3.), index=['a', 'b', 'c'])
ser2[-1]
ser[:1]
ser.loc[:1]
ser.iloc[:1]
# ### Arithmetic and Data Alignment
s1 = pd.Series([7.3, -2.5, 3.4, 1.5], index=['a', 'c', 'd', 'e'])
s2 = pd.Series([-2.1, 3.6, -1.5, 4, 3.1],
index=['a', 'c', 'e', 'f', 'g'])
s1
s2
s1 + s2
df1 = pd.DataFrame(np.arange(9.).reshape((3, 3)), columns=list('bcd'),
index=['Ohio', 'Texas', 'Colorado'])
df2 = pd.DataFrame(np.arange(12.).reshape((4, 3)), columns=list('bde'),
index=['Utah', 'Ohio', 'Texas', 'Oregon'])
df1
df2
df1 + df2
df1 = pd.DataFrame({'A': [1, 2]})
df2 = pd.DataFrame({'B': [3, 4]})
df1
df2
df1 - df2
# #### Arithmetic methods with fill values
df1 = pd.DataFrame(np.arange(12.).reshape((3, 4)),
columns=list('abcd'))
df2 = pd.DataFrame(np.arange(20.).reshape((4, 5)),
columns=list('abcde'))
df2.loc[1, 'b'] = np.nan
df1
df2
df1 + df2
df1.add(df2, fill_value=0)
1 / df1
df1.rdiv(1)
df1 = pd.DataFrame(np.arange(12.).reshape((3, 4)),
columns=list('abcd'))
df2 = pd.DataFrame(np.arange(20.).reshape((4, 5)),
columns=list('abcde'))
df2.loc[1, 'b'] = np.nan
print(df1)
print(df2)
df1.reindex(columns=df2.columns, fill_value=0)
df1.reindex(columns=df2.columns)
# #### Operations between DataFrame and Series
arr = np.arange(12.).reshape((3, 4))
arr
arr[0]
arr - arr[0]
frame = pd.DataFrame(np.arange(12.).reshape((4, 3)),
columns=list('bde'),
index=['Utah', 'Ohio', 'Texas', 'Oregon'])
series = frame.iloc[0]
series
frame
frame - series
series2 = pd.Series(range(3), index=['b', 'e', 'f'])
frame + series2
series3 = frame['d']
frame
series3
frame.sub(series3, axis='index')
# ### Function Application and Mapping
frame = pd.DataFrame(np.random.randn(4, 3), columns=list('bde'),
index=['Utah', 'Ohio', 'Texas', 'Oregon'])
frame
np.abs(frame)
f = lambda x: x.max() - x.min()
frame.apply(f)
frame.apply(f, axis='columns')
def f(x):
return pd.Series([x.min(), x.max()], index=['min', 'max'])
frame.apply(f)
format = lambda x: '%.2f' % x
frame.applymap(format)
frame['e'].map(format)
# ### Sorting and Ranking
obj = pd.Series(range(4), index=['d', 'a', 'b', 'c'])
obj.sort_index()
frame = pd.DataFrame(np.arange(8).reshape((2, 4)),
index=['three', 'one'],
columns=['d', 'a', 'b', 'c'])
frame.sort_index()
frame.sort_index(axis=1)
frame.sort_index(axis=1, ascending=False)
obj = pd.Series([4, 7, -3, 2])
obj.sort_values()
obj = pd.Series([4, np.nan, 7, np.nan, -3, 2])
obj.sort_values()
frame = pd.DataFrame({'b': [4, 7, -3, 2], 'a': [0, 1, 0, 1]})
frame
frame.sort_values(by='b')
frame.sort_values(by=['a', 'b'])
obj = pd.Series([7, -5, 7, 4, 2, 0, 4])
obj.rank()
obj.rank(method='first')
# Assign tie values the maximum rank in the group
obj.rank(ascending=False, method='max')
frame = pd.DataFrame({'b': [4.3, 7, -3, 2], 'a': [0, 1, 0, 1],
'c': [-2, 5, 8, -2.5]})
frame
frame.rank(axis='columns')
# ### Axis Indexes with Duplicate Labels
obj = pd.Series(range(5), index=['a', 'a', 'b', 'b', 'c'])
obj
obj.index.is_unique
obj['a']
obj['c']
df = pd.DataFrame(np.random.randn(4, 3), index=['a', 'a', 'b', 'b'])
df
df.loc['b']
# ## Summarizing and Computing Descriptive Statistics
df = pd.DataFrame([[1.4, np.nan], [7.1, -4.5],
[np.nan, np.nan], [0.75, -1.3]],
index=['a', 'b', 'c', 'd'],
columns=['one', 'two'])
df
df.sum()
df.sum(axis='columns')
df.mean(axis='columns', skipna=False)
df.idxmax()
df.cumsum()
df.describe()
obj = pd.Series(['a', 'a', 'b', 'c'] * 4)
obj.describe()
# ### Correlation and Covariance
# conda install pandas-datareader
price = pd.read_pickle('examples/yahoo_price.pkl')
volume = pd.read_pickle('examples/yahoo_volume.pkl')
# import pandas_datareader.data as web
# all_data = {ticker: web.get_data_yahoo(ticker)
# for ticker in ['AAPL', 'IBM', 'MSFT', 'GOOG']}
#
# price = pd.DataFrame({ticker: data['Adj Close']
# for ticker, data in all_data.items()})
# volume = pd.DataFrame({ticker: data['Volume']
# for ticker, data in all_data.items()})
returns = price.pct_change()
returns.tail()
returns['MSFT'].corr(returns['IBM'])
returns['MSFT'].cov(returns['IBM'])
returns.MSFT.corr(returns.IBM)
returns.corr()
returns.cov()
returns.corrwith(returns.IBM)
returns.corrwith(volume)
# ### Unique Values, Value Counts, and Membership
obj = pd.Series(['c', 'a', 'd', 'a', 'a', 'b', 'b', 'c', 'c'])
uniques = obj.unique()
uniques
obj.value_counts()
pd.value_counts(obj.values, sort=False)
obj
mask = obj.isin(['b', 'c'])
mask
obj[mask]
to_match = pd.Series(['c', 'a', 'b', 'b', 'c', 'a'])
unique_vals = pd.Series(['c', 'b', 'a'])
pd.Index(unique_vals).get_indexer(to_match)
data = pd.DataFrame({'Qu1': [1, 3, 4, 3, 4],
'Qu2': [2, 3, 1, 2, 3],
'Qu3': [1, 5, 2, 4, 4]})
data
result = data.apply(pd.value_counts).fillna(0)
result
# ## Conclusion
pd.options.display.max_rows = PREVIOUS_MAX_ROWS
| 5.wmch5-pands/wmch05-pands.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
"""
Generate Plot
"""
import pandas as pd
import matplotlib.pyplot as plt
TOKENS = 10000
BUCKET_WIDTH = 500
BINS = int(TOKENS / BUCKET_WIDTH)
BAR_WIDTH = 350
PLOT_COLOR = "r"
LOWER_BOUND = -1000
UPPER_BOUND = 11200
grifter_addresses = [
"0x3D9f185369eC7A6fD1be7C1B9f828C9C35A862Bd",
"0xCEFcDBfd67D70C091a4a7a4696ebBc94c06B832E",
"0xD9ADcc4b8d9aD089f262C0F322401B944cd43231",
"0x0D78579978566EE2aC<KEY>",
"0xB3879479eC82638e3c5E4247678eD6fCa771880b",
"0xfb8c9c42b9ba4502a72b0ba1bab35640f44c2dab",
]
for i in range(0, len(grifter_addresses)):
grifter_addresses[i] = grifter_addresses[i].lower()
minting = pd.read_csv("../data/superlativesecretsociety_minting.csv")
grifter_minting = minting[minting["to_account"].isin(grifter_addresses)]
grifter_mint_count = len(grifter_minting)
grifter_dict = {}
total_sales = 0
for i in range(0, BINS):
lower = i * BUCKET_WIDTH
upper = (i + 1) * BUCKET_WIDTH
temp_grifter_df = grifter_minting[
(grifter_minting["rank"] > lower) & (grifter_minting["rank"] < upper)
]
grifter_dict[lower] = len(temp_grifter_df) / grifter_mint_count * 100
plt.bar(
grifter_dict.keys(),
grifter_dict.values(),
width=BAR_WIDTH,
label='"Luckiest" Minters (Minted {} NFTs)'.format(grifter_mint_count),
color=PLOT_COLOR,
edgecolor="black",
)
plt.legend(loc="upper right")
plt.xlabel("Rarity Bucket")
plt.ylabel("Relative Frequency (%)")
plt.title("Tokens minted per Rarity bucket")
plt.xlim(LOWER_BOUND, UPPER_BOUND)
plt.savefig("../figures/luckiest_minters.png", dpi=300)
# -
| case_studies/sss/code/rare_tokens_per_bucket.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 with Spark
# language: python3
# name: python36
# ---
# # Quiz 5 - IBM Watson Studio
# TOTAL POINTS 5
# ## 1.Question 1
#
# ### What is the key difference between Skills Network Labs and IBM Watson Studio (Data Science Experience)?
#
#
# A. IBM Watson Studio is an enterprise-ready environment with scalability in mind, whereas Skills Network Labs is primarily a learning environment.
#
#
# B. IBM Watson Studio has a concept of "Projects" for Jupyter Notebooks, whereas Skills Netowrk Labs does not.
#
#
# C. There is no difference between IBM Watson Studio and Skills Network Labs
#
# ### Ans: A
#
# ## 2.Question 2
#
# ### True or False? The only way to add notebooks to your project on IBM Watson Studio is to either create an empty notebook, or upload a notebook from your computer.
#
#
# True
#
#
# False
#
# ### Ans: True
#
# ## 3.Question 3
#
# ### Which of the follow options are correct? Select all that apply:
#
#
# A. You can add collaborators to projects
#
#
# B. Collaborators on a project can add comments to notebooks.
#
#
# C. Projects are public and anyone can access the notebooks or data within.
#
#
# D. You can use Python 3, R or Scala with Jupyter Notebooks on Watson Studio
#
# ### Ans: A, B, D
#
# ## 4.Question 4
#
# ### In the upper right-hand corner of a Jupyter Notebook is the kernel interpreter (e.g., Python 3). Next to it is a circle. What does it mean if the circle is a full, black circle and what should you do?
#
#
# A. A black circle means that the kernel is not ready to execute more code. You can either wait, interrupt, or try to restart the kernel.
#
#
# B. A black circle means that the kernel is ready to execute code. You can run code as normal, which will be executed by the kernel as it is ready.
#
#
# C. A black circle means that the notebook is currently being saved, and you can wait a few seconds for it to finish saving.
#
#
# D. A black circle means that the notebook is now capable of running Python, R and Scala in the same notebook.
#
# ### Ans: C
#
# ## 5.Question 5
#
# ### True or False? RStudio is available on IBM Watson Studio
#
#
# True
#
#
# False
#
# ### Ans: True
| Open-Source-tools-for-Data-Science/Week 3/Quiz 5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="267d5504009eca2b809a2691131366baebe98436" id="B8nRWthodt0I" colab_type="code" colab={}
import os
import time
import tensorflow as tf
import numpy as np
from glob import glob
import datetime
import random
from PIL import Image
import matplotlib.pyplot as plt
from numpy import savetxt
import pandas as pd
import sys
from functools import reduce
# %matplotlib inline
array_sum = []
# + id="kvUj_IxLeZSO" colab_type="code" outputId="f180e2f0-c41f-468a-9ca7-98656377dc57" colab={"base_uri": "https://localhost:8080/", "height": 34}
from google.colab import drive
drive.mount('/content/drive')
# + _uuid="fc376df46433261bfeb643a95793718a9d969ed1" id="nXVubwxBdt0T" colab_type="code" colab={}
def generator(z, output_channel_dim, training):
with tf.variable_scope("generator", reuse= not training):
# 8x8x1024
fully_connected = tf.layers.dense(z, 8*8*1024)
fully_connected = tf.reshape(fully_connected, (-1, 8, 8, 1024))
fully_connected = tf.nn.leaky_relu(fully_connected)
# 8x8x1024 -> 16x16x512
trans_conv1 = tf.layers.conv2d_transpose(inputs=fully_connected,
filters=512,
kernel_size=[5,5],
strides=[2,2],
padding="SAME",
kernel_initializer=tf.truncated_normal_initializer(stddev=WEIGHT_INIT_STDDEV),
name="trans_conv1")
batch_trans_conv1 = tf.layers.batch_normalization(inputs = trans_conv1,
training=training,
epsilon=EPSILON,
name="batch_trans_conv1")
trans_conv1_out = tf.nn.leaky_relu(batch_trans_conv1,
name="trans_conv1_out")
# 16x16x512 -> 32x32x256
trans_conv2 = tf.layers.conv2d_transpose(inputs=trans_conv1_out,
filters=256,
kernel_size=[5,5],
strides=[2,2],
padding="SAME",
kernel_initializer=tf.truncated_normal_initializer(stddev=WEIGHT_INIT_STDDEV),
name="trans_conv2")
batch_trans_conv2 = tf.layers.batch_normalization(inputs = trans_conv2,
training=training,
epsilon=EPSILON,
name="batch_trans_conv2")
trans_conv2_out = tf.nn.leaky_relu(batch_trans_conv2,
name="trans_conv2_out")
# 32x32x256 -> 64x64x128
trans_conv3 = tf.layers.conv2d_transpose(inputs=trans_conv2_out,
filters=128,
kernel_size=[5,5],
strides=[2,2],
padding="SAME",
kernel_initializer=tf.truncated_normal_initializer(stddev=WEIGHT_INIT_STDDEV),
name="trans_conv3")
batch_trans_conv3 = tf.layers.batch_normalization(inputs = trans_conv3,
training=training,
epsilon=EPSILON,
name="batch_trans_conv3")
trans_conv3_out = tf.nn.leaky_relu(batch_trans_conv3,
name="trans_conv3_out")
# 64x64x128 -> 128x128x64
trans_conv4 = tf.layers.conv2d_transpose(inputs=trans_conv3_out,
filters=64,
kernel_size=[5,5],
strides=[2,2],
padding="SAME",
kernel_initializer=tf.truncated_normal_initializer(stddev=WEIGHT_INIT_STDDEV),
name="trans_conv4")
batch_trans_conv4 = tf.layers.batch_normalization(inputs = trans_conv4,
training=training,
epsilon=EPSILON,
name="batch_trans_conv4")
trans_conv4_out = tf.nn.leaky_relu(batch_trans_conv4,
name="trans_conv4_out")
# 128x128x64 -> 128x128x3
logits = tf.layers.conv2d_transpose(inputs=trans_conv4_out,
filters=3,
kernel_size=[5,5],
strides=[1,1],
padding="SAME",
kernel_initializer=tf.truncated_normal_initializer(stddev=WEIGHT_INIT_STDDEV),
name="logits")
out = tf.tanh(logits, name="out")
return out
# + _uuid="ba53a4bb09dbcd57d3e3392f74ccd054ecf23ecb" id="SSIfMkOEdt0Z" colab_type="code" colab={}
def discriminator(x, reuse):
with tf.variable_scope("discriminator", reuse=reuse):
# 128*128*3 -> 64x64x64
conv1 = tf.layers.conv2d(inputs=x,
filters=64,
kernel_size=[5,5],
strides=[2,2],
padding="SAME",
kernel_initializer=tf.truncated_normal_initializer(stddev=WEIGHT_INIT_STDDEV),
name='conv1')
batch_norm1 = tf.layers.batch_normalization(conv1,
training=True,
epsilon=EPSILON,
name='batch_norm1')
conv1_out = tf.nn.leaky_relu(batch_norm1,
name="conv1_out")
# 64x64x64-> 32x32x128
conv2 = tf.layers.conv2d(inputs=conv1_out,
filters=128,
kernel_size=[5, 5],
strides=[2, 2],
padding="SAME",
kernel_initializer=tf.truncated_normal_initializer(stddev=WEIGHT_INIT_STDDEV),
name='conv2')
batch_norm2 = tf.layers.batch_normalization(conv2,
training=True,
epsilon=EPSILON,
name='batch_norm2')
conv2_out = tf.nn.leaky_relu(batch_norm2,
name="conv2_out")
# 32x32x128 -> 16x16x256
conv3 = tf.layers.conv2d(inputs=conv2_out,
filters=256,
kernel_size=[5, 5],
strides=[2, 2],
padding="SAME",
kernel_initializer=tf.truncated_normal_initializer(stddev=WEIGHT_INIT_STDDEV),
name='conv3')
batch_norm3 = tf.layers.batch_normalization(conv3,
training=True,
epsilon=EPSILON,
name='batch_norm3')
conv3_out = tf.nn.leaky_relu(batch_norm3,
name="conv3_out")
# 16x16x256 -> 16x16x512
conv4 = tf.layers.conv2d(inputs=conv3_out,
filters=512,
kernel_size=[5, 5],
strides=[1, 1],
padding="SAME",
kernel_initializer=tf.truncated_normal_initializer(stddev=WEIGHT_INIT_STDDEV),
name='conv4')
batch_norm4 = tf.layers.batch_normalization(conv4,
training=True,
epsilon=EPSILON,
name='batch_norm4')
conv4_out = tf.nn.leaky_relu(batch_norm4,
name="conv4_out")
# 16x16x512 -> 8x8x1024
conv5 = tf.layers.conv2d(inputs=conv4_out,
filters=1024,
kernel_size=[5, 5],
strides=[2, 2],
padding="SAME",
kernel_initializer=tf.truncated_normal_initializer(stddev=WEIGHT_INIT_STDDEV),
name='conv5')
batch_norm5 = tf.layers.batch_normalization(conv5,
training=True,
epsilon=EPSILON,
name='batch_norm5')
conv5_out = tf.nn.leaky_relu(batch_norm5,
name="conv5_out")
flatten = tf.reshape(conv5_out, (-1, 8*8*1024))
logits = tf.layers.dense(inputs=flatten,
units=1,
activation=None)
out = tf.sigmoid(logits)
return out, logits
# + _uuid="8ef5bbb8e4d157577f1b15600aa64cb40289a754" id="zAB6e5Zcdt0f" colab_type="code" colab={}
def model_loss(input_real, input_z, output_channel_dim):
g_model = generator(input_z, output_channel_dim, True)
noisy_input_real = input_real + tf.random_normal(shape=tf.shape(input_real),
mean=0.0,
stddev=random.uniform(0.0, 0.1),
dtype=tf.float32)
d_model_real, d_logits_real = discriminator(noisy_input_real, reuse=False)
d_model_fake, d_logits_fake = discriminator(g_model, reuse=True)
d_loss_real = tf.reduce_mean(tf.nn.sigmoid_cross_entropy_with_logits(logits=d_logits_real,
labels=tf.ones_like(d_model_real)*random.uniform(0.9, 1.0)))
d_loss_fake = tf.reduce_mean(tf.nn.sigmoid_cross_entropy_with_logits(logits=d_logits_fake,
labels=tf.zeros_like(d_model_fake)))
d_loss = tf.reduce_mean(0.5 * (d_loss_real + d_loss_fake))
g_loss = tf.reduce_mean(tf.nn.sigmoid_cross_entropy_with_logits(logits=d_logits_fake,
labels=tf.ones_like(d_model_fake)))
return d_loss, g_loss
# + _uuid="01820b197de1b6d0ca39592043308557d941937a" id="R__LaaVKdt0k" colab_type="code" colab={}
def model_optimizers(d_loss, g_loss):
t_vars = tf.trainable_variables()
g_vars = [var for var in t_vars if var.name.startswith("generator")]
d_vars = [var for var in t_vars if var.name.startswith("discriminator")]
update_ops = tf.get_collection(tf.GraphKeys.UPDATE_OPS)
gen_updates = [op for op in update_ops if op.name.startswith('generator')]
with tf.control_dependencies(gen_updates):
d_train_opt = tf.train.AdamOptimizer(learning_rate=LR_D, beta1=BETA1).minimize(d_loss, var_list=d_vars)
g_train_opt = tf.train.AdamOptimizer(learning_rate=LR_G, beta1=BETA1).minimize(g_loss, var_list=g_vars)
return d_train_opt, g_train_opt
# + id="1BKiT1vDdt0o" colab_type="code" colab={}
def model_inputs(real_dim, z_dim):
inputs_real = tf.placeholder(tf.float32, (None, *real_dim), name='inputs_real')
inputs_z = tf.placeholder(tf.float32, (None, z_dim), name="input_z")
learning_rate_G = tf.placeholder(tf.float32, name="lr_g")
learning_rate_D = tf.placeholder(tf.float32, name="lr_d")
return inputs_real, inputs_z, learning_rate_G, learning_rate_D
# + _uuid="3a7808bacf25966ab5a8ca06e2e1075ac8eeb662" id="RwmTdg0hdt0t" colab_type="code" colab={}
def show_samples(sample_images, name, epoch):
figure, axes = plt.subplots(1, len(sample_images), figsize = (IMAGE_SIZE, IMAGE_SIZE))
for index, axis in enumerate(axes):
axis.axis('off')
#flatten() change 3d array into 1d
image_array = sample_images[index].flatten()
print("image_array:", image_array)
# !kill -9 -1
if epoch == 10:
# !kill -9 -1
array_sum = np.concatenate((image_array, array_sum), axis=0)
print("array_sum:", array_sum)
else:
print(epoch)
print("image_array_size:", image_array.size)
#print(image_array.shape)
#print("image_array:", image_array)
if epoch == 300:
#save to csv file
#pd.DataFrame(image_array).to_csv("/content/drive/My Drive/Colab Notebooks/image_generator/output.csv")
savetxt('/content/drive/My Drive/Colab Notebooks/image_generator/output.csv', array_sum, delimiter=' ')
#axis.imshow(image_array)
#实现array到image的转换
#image = Image.fromarray(image_array)
#image.save(name+"_"+str(epoch)+"_"+str(index)+".png")
#plt.savefig(name+"_"+str(epoch)+".png", bbox_inches='tight', pad_inches=0)
#plt.show()
#plt.close()
# + id="hXLGmyBUdt0x" colab_type="code" colab={}
def test(sess, input_z, out_channel_dim, epoch):
example_z = np.random.uniform(-1, 1, size=[SAMPLES_TO_SHOW, input_z.get_shape().as_list()[-1]])
samples = sess.run(generator(input_z, out_channel_dim, False), feed_dict={input_z: example_z})
samples = samples. flatten()
#sample_images = reduce(lambda z, y :z + y, samples)
#sample_images = [(sample + 1.0) for sample in samples]
#sample_images = [j for sub in sample_images for j in sub]
#sample_images = sample_images.flatten()
if epoch == 300:
print("samples_images:", samples)
print("samples_size:", samples.size)
savetxt('/content/output.csv', samples, delimiter=' ')
else:
print("epoch:", epoch)
print("samples_size:", samples.size)
print(samples)
#sample_images = [((sample + 1.0) * 127.5).astype(np.uint8) for sample in samples] #int8 字节(-128 ~ 127)
#show_samples(sample_images, OUTPUT_DIR + "samples", epoch)
# + _uuid="0ed8f1c378f936ac81ae89b35d5b6914cd6efbbb" id="7ecFv9S9dt01" colab_type="code" colab={}
def summarize_epoch(epoch, duration, sess, d_losses, g_losses, input_z, data_shape):
minibatch_size = int(data_shape[0]//BATCH_SIZE)
# print("Epoch {}/{}".format(epoch, EPOCHS),
# "\nDuration: {:.5f}".format(duration),
# "\nD Loss: {:.5f}".format(np.mean(d_losses[-minibatch_size:])),
# "\nG Loss: {:.5f}".format(np.mean(g_losses[-minibatch_size:])))
# fig, ax = plt.subplots()
# plt.plot(d_losses, label='Discriminator', alpha=0.6)
# plt.plot(g_losses, label='Generator', alpha=0.6)
# plt.title("Losses")
# plt.legend()
# plt.savefig(OUTPUT_DIR + "losses_" + str(epoch) + ".png")
# plt.show()
# plt.close()
#print(input_z)
#print("sess", sess, "d_losses:", d_losses, "input_z:", input_z)
test(sess, input_z, data_shape[3], epoch)
# + id="8FIugUZqdt05" colab_type="code" colab={}
def get_batches(data):
batches = []
for i in range(int(data.shape[0]//BATCH_SIZE)):
batch = data[i * BATCH_SIZE:(i + 1) * BATCH_SIZE]
augmented_images = []
for img in batch:
image = Image.fromarray(img)
if random.choice([True, False]):
image = image.transpose(Image.FLIP_LEFT_RIGHT)
augmented_images.append(np.asarray(image))
batch = np.asarray(augmented_images)
normalized_batch = (batch / 127.5) - 1.0
batches.append(normalized_batch)
return batches
# + id="I1emVMLFdt09" colab_type="code" colab={}
def train(get_batches, data_shape, checkpoint_to_load=None):
input_images, input_z, lr_G, lr_D = model_inputs(data_shape[1:], NOISE_SIZE)
d_loss, g_loss = model_loss(input_images, input_z, data_shape[3])
d_opt, g_opt = model_optimizers(d_loss, g_loss)
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
epoch = 0
iteration = 0
d_losses = []
g_losses = []
for epoch in range(EPOCHS):
epoch += 1
start_time = time.time()
for batch_images in get_batches:
iteration += 1
batch_z = np.random.uniform(-1, 1, size=(BATCH_SIZE, NOISE_SIZE))
_ = sess.run(d_opt, feed_dict={input_images: batch_images, input_z: batch_z, lr_D: LR_D})
_ = sess.run(g_opt, feed_dict={input_images: batch_images, input_z: batch_z, lr_G: LR_G})
d_losses.append(d_loss.eval({input_z: batch_z, input_images: batch_images}))
g_losses.append(g_loss.eval({input_z: batch_z}))
summarize_epoch(epoch, time.time()-start_time, sess, d_losses, g_losses, input_z, data_shape)
# + _uuid="d857345ed6c3ba7e4f614fa90b5ca42adb9917cf" id="haT4YG4_dt1B" colab_type="code" colab={}
# Paths
INPUT_DATA_DIR = "/content/drive/My Drive/Colab Notebooks/image_generator/" # Path to the folder with input images. For more info check simspons_dataset.txt
OUTPUT_DIR = './{date:%Y-%m-%d_%H:%M:%S}/'.format(date=datetime.datetime.now())
if not os.path.exists(OUTPUT_DIR):
os.makedirs(OUTPUT_DIR)
# + id="6JtZdl_adt1F" colab_type="code" colab={}
# Hyperparameters
IMAGE_SIZE = 128
NOISE_SIZE = 100
LR_D = 0.00004
LR_G = 0.0004
BATCH_SIZE = 64
EPOCHS = 300
BETA1 = 0.5
WEIGHT_INIT_STDDEV = 0.02
EPSILON = 0.00005
SAMPLES_TO_SHOW = 5
# + id="1GCmrEE3e2T4" colab_type="code" colab={}
def readcsv(filename):
data = pd.read_csv(filename) #Please add four spaces here before this line
return(np.array(data)) #Please add four spaces here before this line
# + _uuid="1ee6349afccb4d2f29f36d6605dd2f156350821a" id="wjcKFrFYdt1K" colab_type="code" outputId="265f828c-fe39-4046-99cb-595120c444e4" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# Training
#input_images = np.asarray([np.asarray(Image.open("/content/drive/My Drive/Colab Notebooks/AI_Lab/image_generator/1.png").resize((IMAGE_SIZE, IMAGE_SIZE))) for file in glob(INPUT_DATA_DIR + '*')])
#print (input_images.shape)
#print (input_images.size) #294912
# #!kill -9 -1
#np.random.shuffle(input_images)
#print ("==========================")
#sample_images = random.sample(list(input_images), SAMPLES_TO_SHOW)
#sample_images = list(input_images)
#print(sample_images)
Input_data = readcsv("/content/drive/My Drive/Colab Notebooks/image_generator/data02.csv") #data02 295098 data02(295098)-186=293912 这里data02已经减去了这么多,为了方便reshape矩阵
#Input_data = np.reshape(Input_data, (128, 3)) #这里还是要用它原来的维度 (6, 128, 128, 3)
#Input_data = np.reshape(Input_data, (128, 128, 3))
Input_data = np.reshape(Input_data, (294912,))
#print(Input_data.size)
#print(Input_data.shape)
#print(Input_data)
Input_data = np.reshape(Input_data, (6, 128, 128, 3))
#print(Input_data.size)
#Input_data = np.transpose(Input_data)
#print(Input_data)
#print (Input_data.shape)
print ("Input_data.size:", Input_data.size)
# #!kill -9 -1
#show_samples(Input_data, OUTPUT_DIR + "inputs", 0)
with tf.Graph().as_default():
train(get_batches(Input_data), Input_data.shape)
| ImageGeneratorDCGANmodify04.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import h5py as h5
import matplotlib.pyplot as plt
# +
thirty = '../db/rebus/30_rebus.hdf5'
three = '../db/rebus/3_rebus.hdf5'
f = h5.File(thirty, 'r')
k = h5.File(three, 'r')
keff_thirty = np.array(f['keff_BOC'])[0]
keff_three = np.array(k['keff_BOC'])[0]
for indx, x in enumerate(keff_three):
if x < 1.0:
kill_indx = indx
break
keff_three = keff_three[:kill_indx]
# +
thirty_x = np.arange(0, len(keff_thirty) * 30, 30)
three_x = np.arange(0, len(keff_three) * 3, 3)
print(len(thirty_x))
print(len(three_x))
# convert days to years
thirty_x = thirty_x / 365
three_x = three_x / 365
# -
plt.scatter(three_x, keff_three, label='3 days timestep', s=2)
plt.scatter(thirty_x, keff_thirty, label='30 days timestep', s=2)
plt.plot(thirty_x, [1.01]*len(thirty_x))
plt.xlabel('Operating time [Years]')
plt.ylabel('Keff after depletion')
plt.title('Keff values for different Saltproc timesteps')
plt.legend()
plt.grid()
plt.show()
| script/timestep_resolution_sensitivity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
t1 = tf.constant([[1, 2],
[3, 4],
[5, 6]])
tf.random.shuffle(t1, seed=42)
t2 = tf.constant([[11, 12],
[13, 14],
[15, 16]])
tf.random.set_seed(42)
tf.random.shuffle(t2)
# ## Other ways to create a tensor
tf.ones([1, 2, 3, 4])
tf.zeros(shape=(1, 2, 3, 4))
# +
# NumPy arrays into tensors
# -
import numpy as np
numpy_A = np.arange(1, 25, dtype=np.int32)
numpy_A
A = tf.constant(numpy_A, shape=(2, 3, 4))
B = tf.constant(numpy_A, shape=(3, 8))
A
B
| learning-notebooks/pracitce_tf_seed.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Miscellaneous notes
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import sklearn.datasets
data = sklearn.datasets.load_boston()
data.keys()
df = pd.DataFrame(data['data'], columns=data['feature_names'])
df.describe()
| assignments/misc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.0 64-bit
# language: python
# name: python3
# ---
# # Adafruit IO project docs
# You can find a more extensive documentation for the Adafruit_IO module [here](https://adafruit-io-python-client.readthedocs.io/en/latest/quickstart.html)
# #### Setting up adafruitIO client
# +
from Adafruit_IO import Client, MQTTClient, Feed, Data, RequestError
ioKey = "<KEY>",
c = "dafischnaller"
aio = Client(c, ioKey)
# -
# ## Send and recieve data from a feed
# +
# create a new feed with "test_feed" as name and key
test_feed = Feed(name="test_feed", key="test_feed")
aio.create_feed(test_feed)
# try accessing a feedtest via its key, else raise an error and create it
try:
feed = aio.feeds('feedtest')
except RequestError:
feed = aio.create_feed(Feed(name='feedtest', key='feedtest'))
aio.send('Foo', 100) # sends 100 to the feed Foo
data = aio.receive('Foo') # get's most reacent value in feed Foo
print(data.value) # prints value of the value of the retrieved data
# -
# #### Create, retrieve, delete Feeds
# +
# Create a feed called Foo
# Feed has the parameters name, ID, key
feed = Feed(name='Foo')
result = aio.create_feed(feed) # result will contain all the details about the feed
#===============================
# Retrieve feeds on IO
feeds = aio.feeds()
# Print out the feed names:
for f in feeds:
print('Feed: {0}'.format(f.name))
#===============================
# deletes feed with name foo
aio.delete_feed('Foo')
# -
# #### Data
# +
# creates data item with value 10
data = Data(value=10)
# adds data item to the Foo feed
aio.create_data('Foo', data)
#===============================
# Retrieve data
#Get array of all data items from feed Foo
data = aio.data('Foo')
#===============================
# Delete data
#Delete a data value from Foo 'Test' with ID 1.
data = aio.delete('Foo', 1)
# -
# # MQTT Client
# +
client = MQTTClient(c, ioKey)
def message(client, feed_id, payload):
# prints out the feed id and the payload
print('Feed {0} received new value: {1}'.format(feed_id, payload))
def connected(client):
# listens to feed Foo
client.subscribe('Foo')
def disconnected(client):
print('Disconnected from Adafruit IO!')
client.on_message = message
client.on_connect = connected
client.on_disconnect = disconnected
client.connect()
# loops to check for new data
client.loop_blocking()
# Alternative non blocking way
# client.loop_background()
| docs/AdafruitIO.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="vg-HOITYJAy0"
# #### Основы программирования в Python для социальных наук
#
# *Автор: <NAME>, НИУ ВШЭ*
#
#
# ## Cамостоятельная работа 3
#
# ## Вариант 1
#
# Самостоятельная работа по темам:
#
# * Web-scraping
#
# **Списывание и использование телефонов**
#
# Первое предупреждение, сделанное ассистентом или преподавателем: - 1 балл к вашей оценке.
#
# Второе предупреждение: работа аннулируется без права переписывания.
#
# **Все задачи, где не указано иного, должны принимать значения на ввод (решение для частного случая будет считаться неполным).**
# + [markdown] colab_type="text" id="S3g4tDtrMoxX"
# **Задание 1.**
# *5 баллов*
#
# 1. На странице в https://en.wikipedia.org/wiki/List_of_Rick_and_Morty_episodes нужно найти таблицу под названием "Season 1 (2013–14)".
# 2. С помощью поиска по тегам, нужно сохранить из таблицы следующие колонки: 'Title', 'Directed By', 'U.S. viewers (millions)'. Каждая колонка таблицы должна быть сохранена в отдельную переменную (см. ниже названия), внутри которой лежит список, где первое значение - название колонки.
#
# Обратите внимание, положение элемента в ряде с table headers (th) и в обыкновенных рядах (теги td) может не совпадать.
#
# Например, колонки 'Title' список будет выглядеть так:
# ['Title', "Pilot", ...остальные значения..., "Ricksy Business"]
#
# Значения для 'U.S. viewers (millions)' на этом этапе могут содержать ссылки (например, '1.10[7]')
#
# 3. Выведите эти три списка командой
# print(titles)
# print(directors)
# print(viewers)
# + colab={} colab_type="code" id="KRw05f_FKOXQ"
# Ваше решение. При необходимости создавайте новые ячейки под этой с помощью знака + wikiepisodetable
import requests
import re
from bs4 import BeautifulSoup
website_url = requests.get('https://en.wikipedia.org/wiki/List_of_Rick_and_Morty_episodes').text
soup = BeautifulSoup(website_url,'lxml')
My_table = soup.find_all('table',{'class':'wikitable plainrowheaders wikiepisodetable'})
rows = My_table[0].find_all('tr')
#for x in rows:
# print('______________________________________')
#print(x)
titles = []
for x in range(1, len(rows)):
titles.append(rows[x].find_all('td')[1].get_text())
directors = []
for x in range(1, len(rows)):
directors.append(rows[x].find_all('td')[2].get_text())
viewers = []
for x in range(1, len(rows)):
viewers.append(rows[x].find_all('td')[5].get_text())
print(titles)
print(directors)
print(viewers)
# + [markdown] colab_type="text" id="sZFN--3fNTt2"
# **Задание 2.**
# *5 баллов (каждый шаг 1 балл)*
#
# 1. Из списка viewers удалите то, что в wiki было ссылками (номера в квадратных строках) и переведите значения во float.
# Правильный список viewers будет выглядеть так:
# ['U.S. viewers(millions)', 1.1 ...]
#
# 2. Создайте словарь season1, в котором ключ - название эпизода, а значение - список из имени режиссера, количества зрителей (в формате float), и номера эпизода (не учитывая названия колонок в нулевом элементе). Например, пара ключ-значение для первого эпизода будет выглядеть так
# `'"Pilot"': ['<NAME>', 1.1, 1]`
# 2. Напишите функцию, которая берет аргументом название эпизода и печатает следующую строку: {номер эпизода} Episode {название эпизода} (directed by {имя режиссера} was watched by {количество зрителей} million viewers in the US.
#
# 3. Запустите вашу функцию в цикле для всех эпизодов (не учитывая названия колонки в нулевом элементе).
#
# 5. Сколько всего зрителей посмотрело эпизоды, которые режиссировал <NAME>?
# +
# Запустите эту ячейку
titles = ['Title', '"Pilot"', '"Lawnmower Dog"', '"Anatomy Park"', '"M. Night Shaym-Aliens!"', '"Meeseeks and Destroy"', '"Rick Potion #9"', '"Raising Gazorpazorp"', '"Rixty Minutes"', '"Something Ricked This Way Comes"', '"Close Rick-counters of the Rick Kind"', '"Ricksy Business"']
viewers = ['U.S. viewers(millions)', '1.10[5]', '1.51[6]', '1.30[7]', '1.32[8]', '1.61[9]', '1.75[10]', '1.76[11]', '1.48[12]', '1.54[13]', '1.75[14]', '2.13[15]']
directors = ['Directed by', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>']
print(titles)
print(viewers)
print(directors)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="pwLxGVugPVys" outputId="0eb3382d-eba5-4e68-8edb-2b3e15a57ac0"
# 1
import re
l = []
l.append('U.S. viewers(millions)')
for x in viewers:
if x != 'U.S. viewers(millions)':
l.append(re.sub('\[\d+\]', '', x))
viewers = l
print(viewers)
# +
# 2
ind = 0
season1 = {}
for x in titles:
if x != 'Title':
season1[x] = [directors[ind], viewers[ind], ind]
ind += 1
print(season1)
# -
# 3
def epinfo(name):
for x in season1:
if x == name:
print('{} Episode {}(directed by {} was watched by {}million viewers in the US'.format(
season1[name][2],
name,
season1[name][0],
season1[name][1]
))
epinfo('"Pilot"')
# 4
for x in titles:
if x != 'Title':
epinfo(x)
# 5
summ = 0
for x in season1.values():
if x[0] == '<NAME>':
summ += float(x[1])
print(summ)
| web_scraping_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.6 (via pyenv)
# language: python
# name: pyenv-3.7.6
# ---
# # Json file representation
# > Methods to work with `json_file` representation: load/dump from/to a file.
# +
# default_exp json_file
# -
# %load_ext autoreload
# %autoreload 2
# +
# export
import logging
import time
from dataclasses import dataclass
from typing import *
from pathlib import Path
import json
from cocorepr.utils import sort_dict, measure_time
from cocorepr.coco import *
# -
# export
logger = logging.getLogger()
# hide
logging.getLogger().setLevel(logging.DEBUG)
# +
#export
def load_json_file(annotations_json: Union[str, Path], *, kind: str = "object_detection") -> CocoDataset:
from_dict_function = get_dataset_class(kind).from_dict
annotations_json = Path(annotations_json)
logger.info(f"Loading json_file from: {annotations_json}")
ext = annotations_json.suffix
if ext != '.json':
raise ValueError(f'Expect .json file as input, got: {annotations_json}')
with measure_time() as timer:
with measure_time() as timer2:
D = json.loads(annotations_json.read_text())
logger.info(f" json file loaded: elapsed {timer2.elapsed}")
with measure_time() as timer2:
coco = from_dict_function(D)
logger.info(f" dataset constructed: elapsed {timer2.elapsed}")
logger.info(f"Loaded json_file: elapsed {timer.elapsed}: {coco.to_full_str()}")
return coco
# +
# hide
PATH = '../examples/coco_chunk/json_file/instances_train2017_chunk3x2.json'
d = load_json_file(PATH)
display(d.info)
display(d.annotations[0])
display(d.images[0])
display(d.categories[0])
assert isinstance(d.info, CocoInfo), type(d.info)
assert isinstance(d.annotations[0], CocoObjectDetectionAnnotation), type(d.annotations[0])
assert isinstance(d.images[0], CocoImage), type(d.images[0])
assert isinstance(d.categories[0], CocoObjectDetectionCategory), type(d.categories[0])
# +
# export
def dump_json_file(
coco: CocoDataset,
annotations_json: Union[str, Path],
*,
kind: str = "object_detection",
skip_nulls: bool = True,
overwrite: bool = False,
indent: Optional[int] = 4,
) -> None:
dataset_class = get_dataset_class(kind)
if skip_nulls:
to_dict_function = dataset_class.to_dict_skip_nulls
else:
to_dict_function = dataset_class.to_dict
annotations_json = Path(annotations_json)
if annotations_json.is_file() and not overwrite:
raise ValueError(f"Destination json_file already exists: {annotations_json}")
with measure_time() as timer:
raw = sort_dict(to_dict_function(coco))
logger.info(f"Sorted keys: elapsed {timer.elapsed}")
logger.info(f"Writing dataset {coco.to_full_str()} to json-file: {annotations_json}")
with measure_time() as timer:
annotations_json.parent.mkdir(parents=True, exist_ok=True)
annotations_json.write_text(json.dumps(raw, indent=indent, ensure_ascii=False))
logger.info(f"Dataset written to {annotations_json}: elapsed {timer.elapsed}")
# +
# hide
import tempfile
tmp = tempfile.mktemp()
dump_json_file(d, tmp)
# ! cat {tmp} | jq .info
# ! cat {tmp} | jq .images[0]
# ! cat {tmp} | jq .annotations[0]
# ! cat {tmp} | jq .categories[0]
# -
| nbs/02_json_file.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# Before reading this article, your PyTorch script probably looked like this:
# Load entire dataset
X, y = torch.load('some_training_set_with_labels.pt')
# Train model
for epoch in range(max_epochs):
for i in range(n_batches):
# Local batches and labels
local_X, local_y = X[i*n_batches:(i+1)*n_batches,], y[i*n_batches:(i+1)*n_batches,]
# Your model
[...]
# -
partition = {'train': ['id-1', 'id-2', 'id-3'], 'validation': ['id-4']}
labels = {'id-1': 0, 'id-2': 1, 'id-3': 2, 'id-4': 1}
def __init__(self, list_IDs, labels):
'Initialization'
self.labels = labels
self.list_IDs = list_IDs
| Code/006-data_loader.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Conceptos básicos de la estadística descriptiva
#
# La [estadística descriptiva](https://es.wikipedia.org/wiki/Estad%C3%ADstica_descriptiva) es una parte de las ciencias estadísticas que se encarga de **describir** los datos que tenemos en base a unos indicadores. Un ejemplo de estos son:
#
#
# * **Media aritmética**: La [media aritmética](https://es.wikipedia.org/wiki/Media_aritm%C3%A9tica) es el valor obtenido al sumar todos los valores y dividir el resultado entre el número total elementos. Se suele representar con la letra griega $\mu$. Si tenemos un conjunto de $n$ valores, $ \{x_i\}$, la *media aritmética*, $\mu$, es la suma de los valores divididos por el numero de elementos; en otras palabras:
#
# $$\mu = \frac{1}{n} \sum_{i}x_i$$
#
#
#
# * **Desviación respecto a la media**: La desviación respecto a la media es la diferencia en valor absoluto entre cada valor y la media aritmética de todos los valores. Se puede entender con la pregunta ¿cómo de lejos está el valor de un elemento con respecto a la media del conjunto de elementos?.
#
# $$D_i = |x_i - \mu|$$
#
#
# * **Varianza**: La [varianza](https://es.wikipedia.org/wiki/Varianza) es el sumatorio de las diferencias cuadráticas de cada valor menos la media artimética del conjunto de valores, entre la cantidad de valores que haya. Dicho de otra manera es el sumatorio del cuadrado de las desviaciones respecto a la media entre el n es la media aritmética del cuadrado de las desviaciones respecto a la media entre la cantidad de valores que haya. La varianza es una medida de **dispersión de los datos**. El cuadrado hace que los valores sean positivos para evitar que valores negativos modifiquen la varianza y además hace que diferencias altas impacten mucho en el valor final. Se representa como $\sigma^2$.
#
#
# $$\sigma^2 = \frac{\sum\limits_{i=1}^n(x_i - \mu)^2}{n} $$
#
#
# * **Desviación típica**: La [desviación típica](https://es.wikipedia.org/wiki/Desviaci%C3%B3n_t%C3%ADpica) es la raíz cuadrada de la varianza. Es una medida de la **dispersión** de los datos un poco más **robusta** dado que se elimina la componente cuadrática de la expresión. Se representa con la letra griega $\sigma$.
#
# $$\sigma = \sqrt{\frac{\sum\limits_{i=1}^n(x_i - \mu)^2}{n}} $$
#
#
# * **Moda**: La <a href="https://es.wikipedia.org/wiki/Moda_(estad%C3%ADstica)">moda</a> es el valor que tiene mayor frecuencia absoluta. Se representa con $M_0$
#
#
# * **Mediana**: La <a href="https://es.wikipedia.org/wiki/Mediana_(estad%C3%ADstica)">mediana</a> es el valor que ocupa el **lugar central** de todos los datos cuando éstos están ordenados de menor a mayor. Lo que equivaldría al valor que corresponde al 50 % de mis datos o aquel que me divide los datos en 50 % a la izquierda y 50 % a la derecha. Se representa con $\widetilde{x}$.
#
# * **Percentil n**: El [percentil enésimo](https://es.wikipedia.org/wiki/Percentil) es el valor que divide los valores, ordenados de menor a mayor, en n% a la izquierda y (1-n)% a la derecha. Los más importantes son el 25, y el 75 junto con el 50 que es la mediana.
#
# * **Correlación**: La [correlación](https://es.wikipedia.org/wiki/Correlaci%C3%B3n) trata de establecer la relación o **dependencia** que existe entre las dos variables que intervienen en una distribución bidimensional. Es decir, determinar si los cambios en una de las variables influyen en los cambios de la otra. En caso de que suceda, diremos que las variables están correlacionadas o que hay correlación entre ellas. La correlación es positiva cuando los valores de las variables aumenta juntos; y es negativa cuando un valor de una variable se reduce cuando el valor de la otra variable aumenta.
#
#
# * **Covarianza**: La [covarianza](https://es.wikipedia.org/wiki/Covarianza) es el equivalente de la varianza aplicado a una variable bidimensional. Es la media aritmética de los productos de las desviaciones de cada una de las variables respecto a sus medias respectivas.La covarianza indica el **sentido de la correlación** entre las variables; Si $\sigma_{xy} > 0$ la correlación es directa; Si $\sigma_{xy} < 0$ la correlación es inversa.
#
# $$\sigma_{xy} = \frac{\sum\limits_{i=1}^n(x_i - \mu_x)(y_i -\mu_y)}{n}$$
#
#
# * **Valor atípico**: Un [valor atípico](https://es.wikipedia.org/wiki/Valor_at%C3%ADpico) es una observación que se aleja demasiado de la moda; esta muy lejos de la tendencia principal del resto de los datos. Pueden ser causados por errores en la recolección de datos o medidas inusuales. Generalmente se recomienda **eliminarlos** del conjunto de datos
#
# ### Diferencia entre población y muestra
#
# La [población](https://es.wikipedia.org/wiki/Población_estad%C3%ADstica) en estadística es el conjunto de indivíduos, objetos o fenómenos sobre los cual queremos estudiar una o unas características y para ello haremos uso de un [muestra](https://es.wikipedia.org/wiki/Muestra_estad%C3%ADstica) de la población, que en función del estudio que queramos hacer podemos encontrarnos con muestras representativas, que eliminan el sesgo representacional, o muestras no representativas para las que existen maneras de tratar.
#
# # Ejemplos en Python
# +
# Ejemplos de estadistica descriptiva con python
import numpy as np # importando numpy
from scipy import stats # importando scipy.stats
np.random.seed(32) # para poder replicar el random
# + jupyter={"outputs_hidden": false}
# datos normalmente distribuidos en 1 fila con 50 columnas
# o un vector de 50 valores
datos_n = np.random.randn(1, 50)
datos_r = np.random.rand(1, 50) # datos aleatorios
# + jupyter={"outputs_hidden": false}
# media arítmetica usando la función de python
print(datos_n.mean())
print(datos_r.mean())
# + jupyter={"outputs_hidden": false}
# media arítmetica usando la función de numpy
print(np.mean(datos_n))
print(np.mean(datos_r))
# -
help(np.mean)
# + jupyter={"outputs_hidden": false}
# mediana
print(np.median(datos_n))
print(np.median(datos_r))
# -
help(np.median)
# + jupyter={"outputs_hidden": false}
# Desviación típica
print(np.std(datos_n))
print(np.std(datos_r))
# -
help(np.std)
# + jupyter={"outputs_hidden": false}
# varianza
print(np.var(datos_n))
print(np.var(datos_r))
# -
np.std(datos_n)**2
help(np.var)
# + jupyter={"outputs_hidden": false}
# Moda
# El segundo valor resultado es la cantidad de veces que se repite
print(stats.mode(datos_n))
print(stats.mode(datos_r))
# + jupyter={"outputs_hidden": false}
datos2 = np.array([1, 2, 3, 6, 6, 1, 2, 4, 2, 2, 6, 6, 8, 10, 6])
stats.mode(datos2) # aqui la moda es el 6 porque aparece 5 veces en el vector.
# -
help(stats.mode)
# + jupyter={"outputs_hidden": false}
# correlacion
print(np.corrcoef(datos_n))
print(np.corrcoef(datos_r))
# + jupyter={"outputs_hidden": false}
# calculando la correlación entre dos vectores.
np.corrcoef(datos_n, datos_r)
# -
help(np.corrcoef)
# + jupyter={"outputs_hidden": false}
# covarianza
print(np.cov(datos_n))
print(np.cov(datos_r))
# + jupyter={"outputs_hidden": false}
# covarianza de dos vectores
np.cov(datos_n, datos_r)
# -
help(np.cov)
# Percentil
print(np.percentile(datos_n, 25))
print(np.percentile(datos_r, 25))
help(np.percentile)
# # Introducción Probabilidad
# ### ¿Qué es la Probabilidad?
#
# La [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) mide la mayor o menor posibilidad de que se dé un determinado resultado (suceso o evento) cuando se realiza un experimento aleatorio. Toma valores entre 0 y 1 (o expresados en tanto por ciento, entre 0% y 100%).
#
# La [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) es un también complemento de la [estadística](http://es.wikipedia.org/wiki/Estad%C3%ADstica) cuando nos proporciona una sólida base para la [estadistica inferencial](https://es.wikipedia.org/wiki/Estad%C3%ADstica_inferencial). Cuando hay incertidumbre, no sabemos que puede pasar y hay alguna posibilidad de errores, utilizando [probabilidades](https://es.wikipedia.org/wiki/Probabilidad) podemos aprender formas de controlar la tasa de errores para reducirlos.
#
# ### Calculando probabilidades
#
# El cálculo de la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad), cambia dependiendo del tipo de evento que se está observando. Por ejemplo, no calcularíamos nuestras posibilidades de ganar la lotería de la misma manera que calcularíamos la probabilidad de que una maquina se estropee dado un cierto tiempo. Sin embargo, una vez que determinamos si los eventos son <a href="https://es.wikipedia.org/wiki/Independencia_(probabilidad)">independientes</a>, [condicionales](https://es.wikipedia.org/wiki/Probabilidad_condicionada) o mutuamente excluyentes, calcular su probabilidad es relativamente simple.
#
# #### Propiedades básicas de la probabilidad
#
# Antes de poder calcular las [probabilidades](https://es.wikipedia.org/wiki/Probabilidad), primero debemos conocer sus 3 propiedades fundamentales, ellas son:
#
# * La [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) se expresa como un ratio que será un valor positivo menor o igual a 1.
#
# $ 0 \le p(A) \le 1$
#
#
# * La [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) de un evento del que tenemos total certeza es 1.
#
# $ p(S) = 1 $
#
# * Si el evento $A$ y el evento $B$ son *mutuamente excluyentes*, entonces:
#
# $ p(A \cup B ) = p(A) + p(B) $
#
# A partir de estas propiedades básicas, se pueden derivar muchas otras propiedades.
#
# ### Teoría de conjuntos y probabilidades
#
# Vamos a ver como se pueden relacionar los eventos en términos de teoría de [teoría de conjuntos](https://es.wikipedia.org/wiki/Teor%C3%ADa_de_conjuntos).
#
# * **Unión:** La unión de varios eventos simples crea un evento compuesto. La unión de $E$ y $F$ se escribe $E \cup F$ y significa "Ya sea $E$ o $F$, o ambos $E$ y $F$."
#
# * **Intersección:** La intersección de dos o más eventos simples crea un evento compuesto. La intersección de $E$ y $F$ se escribe $E \cap F$ y significa "$E$ y $F$."
#
# * **Complemento:** El complemento de un evento significa todo en el [espacio de muestreo](https://es.wikipedia.org/wiki/Espacio_muestral) que no es ese evento. El complemento del evento $E$ se escribe varias veces como $\sim{E}$, $E^c$, o $\overline{E}$, y se lee como "no $E$" o "complemento $E$".
#
# * **Exclusión mutua:** Si los eventos no pueden ocurrir juntos, son *mutuamente excluyentes*. Siguiendo la misma línea de razonamiento, si dos conjuntos no tienen ningún evento en común, son mutuamente excluyentes.
#
# ### Calculando la probabilidad de múltiples eventos
#
# Ahora sí, ya podemos calcular las [probabilidades](https://es.wikipedia.org/wiki/Probabilidad) de los eventos. Recordemos que la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) de un solo evento se expresa como un ratio entre el número de resultados favorables sobre el número de los posibles resultados. Pero ¿qué pasa cuando tenemos múltiples eventos?
#
# #### Unión de eventos mutuamente excluyentes
# Si los eventos son *mutuamente excluyentes* entonces para calcular la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) de su unión, simplemente sumamos sus [probabilidades](https://es.wikipedia.org/wiki/Probabilidad) individuales.
#
# $p(E \cup F) = p(E) + p(F)$
#
# #### Unión de eventos que no son mutuamente excluyentes
# Si los eventos no son *mutuamente excluyentes* entonces debemos corregir la fórmula anterior para incluir el efecto de la superposición de los eventos. Esta superposición se da en el lugar de la *intersección* de los eventos; por lo tanto la formula para calcular la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) de estos eventos es:
#
# $p(E \cup F) = p(E) + p(F) - p(E \cap F)$
#
# #### Intersección de eventos independientes
# Para calcular la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) de que ocurran varios eventos (la intersección de varios eventos), se multiplican sus [probabilidades](https://es.wikipedia.org/wiki/Probabilidad) individuales. La fórmula específica utilizada dependerá de si los eventos son <a href="https://es.wikipedia.org/wiki/Independencia_(probabilidad)">independientes</a> o no.
# Si son <a href="https://es.wikipedia.org/wiki/Independencia_(probabilidad)">independientes</a>, la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) de $E$ y $F$ se calcula como:
#
# $p(E \cap F) = p(E) \times p(F)$
#
# #### Intersección de eventos no independientes
# Si dos eventos no son <a href="https://es.wikipedia.org/wiki/Independencia_(probabilidad)">independientes</a>, debemos conocer su [probabilidad condicional](https://es.wikipedia.org/wiki/Probabilidad_condicionada) para poder calcular la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) de que ambos se produzcan. La fórmula en este caso es:
#
# $p(E \cap F) = p(E) \times p(F|E)$
#
# ### La probabilidad condicional
#
# Con frecuencia queremos conocer la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) de algún evento, dado que otro evento ha ocurrido. Esto se expresa simbólicamente como $p(E | F)$ y se lee como "la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) de $E$ dado $F$". El segundo evento se conoce como la *condición* y el proceso se refiere a veces como "condicionamiento en F". La [probabilidad condicional](https://es.wikipedia.org/wiki/Probabilidad_condicionada) es un concepto importante de estadística, porque a menudo estamos tratando de establecer que un factor tiene una relación con un resultado, como por ejemplo, que las personas que fuman cigarrillos tienen más [probabilidades](https://es.wikipedia.org/wiki/Probabilidad) de desarrollar cáncer de pulmón. La [probabilidad condicional](https://es.wikipedia.org/wiki/Probabilidad_condicionada) también se puede usar para definir la <a href="https://es.wikipedia.org/wiki/Independencia_(probabilidad)">independencia</a>. Dos variables se dice que son <a href="https://es.wikipedia.org/wiki/Independencia_(probabilidad)">independientes</a> si la siguiente relación se cumple:
#
# $p(E | F) = p(E)$
#
# #### Calculando la probabilidad condicional
# Para calcular la probabilidad del evento $E$ dada la información de que el evento $F$ ha ocurrido utilizamos la siguiente formula:
#
# $p(E | F) = \frac{p(E \cap F)}{p(F)}$
#
#
#
# ### Distintas interpretaciones de la probabilidad
#
# Las [probabilidades](https://es.wikipedia.org/wiki/Probabilidad) pueden ser interpretadas generalmente de dos maneras distintas.
# La interpretación *clasica* de la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) es una perspectiva en la que las [probabilidades](https://es.wikipedia.org/wiki/Probabilidad) se consideran frecuencias relativas constantes a largo plazo. Este es el enfoque clásico de la [teoría de probabilidad](https://es.wikipedia.org/wiki/Teor%C3%ADa_de_la_probabilidad). La interpretación *Bayesiana* de la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) es una perspectiva en la que las [probabilidades](https://es.wikipedia.org/wiki/Probabilidad) son consideradas como *medidas de creencia* que pueden cambiar con el tiempo para reflejar nueva información. El *enfoque clásico* sostiene que los métodos *bayesianos* sufren de falta de objetividad, ya que diferentes individuos son libres de asignar diferentes [probabilidades](https://es.wikipedia.org/wiki/Probabilidad) al mismo evento según sus propias opiniones personales. Los *bayesianos* se oponen a los *clásicos* sosteniendo que la interpretación *frecuentista* de la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) tiene ya de por sí una subjetividad incorporada (por ejemplo, mediante la elección y el diseño del procedimiento de muestreo utilizado) y que la ventaja del *enfoque bayesiano* es que ya hace explícita esta subjetividad.
# En la actualidad, la mayoría de los problemas son abordados siguiendo un enfoque mixto entre ambas interpretaciones de la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad).
# ### Probabilidad y sentido común
#
# La incertidumbre constituye una pieza fundamental del mundo en que vivimos, en parte hace la vida mucho más interesante, ya que sería muy aburrido si todo fuera perfectamente predecible. Aun así, una parte de nosotros quisiera predecir el futuro y que las cosas sean mucho más predecibles. Para poder lidiar con la incertidumbre que nos rodea, solemos aplicar lo que llamamos nuestro "*sentido común*". Por ejemplo, si al levantarnos por la mañana vemos que el día se encuentra nublado, este hecho no nos da la **certeza** de que comenzará a llover más tarde; sin embargo, nuestro *sentido común* puede inducirnos a cambiar nuestros planes y a actuar como si *creyéramos* que fuera a llover si las nubes son los suficientemente oscuras o si escuchamos truenos, ya que nuestra experiencia nos dice que estos signos indicarían una mayor *posibilidad* de que el hecho de que fuera a llover más tarde realmente ocurra.
# Nuestro *sentido común* es algo tan arraigado en nuestro pensamiento, que lo utilizamos automáticamente sin siquiera ponernos a pensar en ello; pero muchas veces, el *sentido común* también nos puede jugar una mala pasada y hacernos elegir una respuesta incorrecta.
#
# Tomemos por ejemplo alguna de las siguiente situaciones...
#
# * **Situación 1 - La coincidencia de cumpleaños:** Vamos a una fiesta a la que concurren un total de 50 personas. Allí un amigo nos desafía afirmando que en la fiesta debe haber por lo menos 2 personas que cumplen años el mismo día y nos apuesta 100 pesos a que está en lo correcto. Es decir, que si él acierta deberíamos pagarle los 100 pesos; o en caso contrario, el nos pagará los 100 pesos. ¿Deberíamos aceptar la apuesta?
#
#
# * **Situación 2 - ¿Que puerta elegir?:** Estamos participando en un concurso en el cual se nos ofrece la posibilidad de elegir una entre tres puertas. Tras una de ellas se encuentra una ferrari ultimo modelo, y detrás de las otras dos hay una cabra; luego de elegir una puerta, el presentador del concurso abre una de las puertas restantes y muestra que hay una cabra (el presentador sabe que hay detrás de cada puerta). Luego de hacer esto, el presentador nos ofrece la posibilidad de cambiar nuestra elección inicial y quedarnos con la otra puerta que no habíamos elegido inicialmente. ¿Deberíamos cambiar o confiar en nuestra elección inicial?
#
# ¿Qué les diría su *sentido común* que deberían hacer en cada una de estas situaciones?
# # Distribuciones de probabilidad
#
# Las [variables aleatorias](https://es.wikipedia.org/wiki/Variable_aleatoria) han llegado a desempeñar un papel importante en casi todos los campos de estudio: en la [Física](https://es.wikipedia.org/wiki/F%C3%ADsica), la [Química](https://es.wikipedia.org/wiki/Qu%C3%ADmica) y la [Ingeniería](https://es.wikipedia.org/wiki/Ingenier%C3%ADa); y especialmente en las ciencias biológicas y sociales. Estas [variables aleatorias](https://es.wikipedia.org/wiki/Variable_aleatoria) son medidas y analizadas en términos
# de sus propiedades [estadísticas](https://es.wikipedia.org/wiki/Estad%C3%ADstica) y [probabilísticas](https://es.wikipedia.org/wiki/Probabilidad), de las cuales una característica subyacente es su [función de distribución](https://es.wikipedia.org/wiki/Funci%C3%B3n_de_distribuci%C3%B3n). A pesar de que el número potencial de [distribuciones](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad) puede ser muy grande, en la práctica, un número relativamente pequeño se utilizan; ya sea porque tienen características matemáticas que las hace fáciles de usar o porque se asemejan bastante bien a una porción de la realidad, o por ambas razones combinadas.
#
# ## ¿Por qué es importante conocer las distribuciones?
#
# Muchos resultados en las ciencias se basan en conclusiones que se extraen sobre una población general a partir del estudio de una [muestra](https://es.wikipedia.org/wiki/Muestra_estad%C3%ADstica) de esta población. Este proceso se conoce como **[inferencia estadística](https://es.wikipedia.org/wiki/Estad%C3%ADstica_inferencial)**; y este tipo de *inferencia* con frecuencia se basa en hacer suposiciones acerca de la forma en que los datos se distribuyen, o requiere realizar alguna transformación de los datos para que se ajusten mejor a alguna de las [distribuciones](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad) conocidas y estudiadas en profundidad.
#
# Las [distribuciones de probabilidad](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad) teóricas son útiles en la [inferencia estadística](https://es.wikipedia.org/wiki/Estad%C3%ADstica_inferencial) porque sus propiedades y características son conocidas. Si la [distribución](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad) real de un [conjunto de datos](https://es.wikipedia.org/wiki/Conjunto_de_datos) dado es razonablemente cercana a la de una [distribución de probabilidad](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad) teórica, muchos de los cálculos se pueden realizar en los datos reales utilizando hipótesis extraídas de la [distribución](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad) teórica.
#
# ## Graficando distribuciones
#
# ### Histogramas
#
# Una de las mejores maneras de describir una variable es representar los valores que aparecen en el [conjunto de datos](https://es.wikipedia.org/wiki/Conjunto_de_datos) y el número de veces que aparece cada valor. La representación más común de una [distribución](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad) es un [histograma](https://es.wikipedia.org/wiki/Histograma), que es un gráfico que muestra la frecuencia de cada valor.
#
# En [Python] podemos graficar de una manera sencilla usando las librerías de [matplotlib](https://matplotlib.org) o [seaborn](https://seaborn.pydata.org/index.html) entre otras. Por ejemplo, podríamos graficar el [histograma](https://es.wikipedia.org/wiki/Histograma) de una [distribución normal](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_normal) del siguiente modo.
# Huir de matplotlib (complicada), utilizar seaborn o, en su defecto, buscar una llamada Altair
# + hide_input=false
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
from scipy import stats
import seaborn as sns
np.random.seed(2016) # replicar random
# parametros esteticos de seaborn
sns.set_palette("deep", desat=.6)
sns.set_context(rc={"figure.figsize": (8, 4)})
sns.set()
# Graficando histograma
mu, sigma = 0, 0.2 # media y desvio estandar
datos = np.random.normal(mu, sigma, 10000) #creando muestra de datos
# histograma de distribución normal.
sns.distplot(datos, 20)
plt.ylabel('frequencia')
plt.xlabel('valores')
plt.title('Histograma')
plt.show()
# -
# ### Función de Masa de Probabilidad
#
# Otra forma de representar a las [distribuciones discretas](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad#Distribuciones_de_variable_discreta) es utilizando su [Función de Masa de Probabilidad](https://es.wikipedia.org/wiki/Funci%C3%B3n_de_probabilidad) o [FMP](https://es.wikipedia.org/wiki/Funci%C3%B3n_de_probabilidad), la cual relaciona cada valor con su [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) en lugar de su *frecuencia* como vimos anteriormente. Esta función es *normalizada* de forma tal que el valor total de [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) sea 1. La ventaja que nos ofrece utilizar la [FMP](https://es.wikipedia.org/wiki/Funci%C3%B3n_de_probabilidad) es que podemos comparar dos [distribuciones](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad) sin necesidad de ser confundidos por las diferencias en el tamaño de las [muestras](https://es.wikipedia.org/wiki/Muestra_estad%C3%ADstica). También debemos tener en cuenta que [FMP](https://es.wikipedia.org/wiki/Funci%C3%B3n_de_probabilidad) funciona bien si el número de valores es pequeño; pero a medida que el número de valores aumenta, la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) asociada a cada valor se hace cada vez más pequeña y el efecto del *ruido aleatorio* aumenta.
# Veamos un ejemplo con [Python](http://python.org/).
# Graficando FMP
n, p = 30, 0.4 # parametros de forma de la distribución binomial
n_1, p_1 = 20, 0.3 # parametros de forma de la distribución binomial
x = np.arange(stats.binom.ppf(0.01, n, p),
stats.binom.ppf(0.99, n, p))
x_1 = np.arange(stats.binom.ppf(0.01, n_1, p_1),
stats.binom.ppf(0.99, n_1, p_1))
fmp = stats.binom.pmf(x, n, p) # Función de Masa de Probabilidad
fmp_1 = stats.binom.pmf(x_1, n_1, p_1) # Función de Masa de Probabilidad
plt.plot(x, fmp, '--')
plt.plot(x_1, fmp_1)
plt.vlines(x, 0, fmp, colors='b', lw=5, alpha=0.5)
plt.vlines(x_1, 0, fmp_1, colors='g', lw=5, alpha=0.5)
plt.title('Función de Masa de Probabilidad')
plt.ylabel('probabilidad')
plt.xlabel('valores')
plt.show()
# ### Función de Distribución Acumulada
#
# Si queremos evitar los problemas que se generan con [FMP](https://es.wikipedia.org/wiki/Funci%C3%B3n_de_probabilidad) cuando el número de valores es muy grande, podemos recurrir a utilizar la [Función de Distribución Acumulada](https://es.wikipedia.org/wiki/Funci%C3%B3n_de_distribuci%C3%B3n) o [FDA](https://es.wikipedia.org/wiki/Funci%C3%B3n_de_distribuci%C3%B3n), para representar a nuestras [distribuciones](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad), tanto [discretas](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad#Distribuciones_de_variable_discreta) como [continuas](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad_continua). Esta función relaciona los valores con su correspondiente [percentil](https://es.wikipedia.org/wiki/Percentil); es decir que va a describir la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) de que una [variable aleatoria](https://es.wikipedia.org/wiki/Variable_aleatoria) X sujeta a cierta ley de [distribución de probabilidad](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad) se sitúe en la zona de valores menores o iguales a x.
# Graficando Función de Distribución Acumulada con Python
x_1 = np.linspace(stats.norm(10, 1.2).ppf(0.01),
stats.norm(10, 1.2).ppf(0.99), 100)
fda_binom = stats.binom.cdf(x, n, p) # Función de Distribución Acumulada
fda_normal = stats.norm(10, 1.2).cdf(x_1) # Función de Distribución Acumulada
plt.plot(x, fda_binom, '--', label='FDA binomial')
plt.plot(x_1, fda_normal, label='FDA nomal')
plt.title('Función de Distribución Acumulada')
plt.ylabel('probabilidad')
plt.xlabel('valores')
plt.legend(loc=4)
plt.show()
# ### Función de Densidad de Probabilidad
#
# Por último, el equivalente a la [FMP](https://es.wikipedia.org/wiki/Funci%C3%B3n_de_probabilidad) para [distribuciones continuas](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad_continua) es la [Función de Densidad de Probabilidad](https://es.wikipedia.org/wiki/Funci%C3%B3n_de_densidad_de_probabilidad) o [FDP](https://es.wikipedia.org/wiki/Funci%C3%B3n_de_densidad_de_probabilidad). Esta función es la [derivada](https://es.wikipedia.org/wiki/Derivada) de la [Función de Distribución Acumulada](https://es.wikipedia.org/wiki/Funci%C3%B3n_de_distribuci%C3%B3n).
# Por ejemplo, para la [distribución normal](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_normal) que graficamos anteriormente, su [FDP](https://es.wikipedia.org/wiki/Funci%C3%B3n_de_densidad_de_probabilidad) es la siguiente. La típica forma de campana que caracteriza a esta [distribución](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad).
# Graficando Función de Densidad de Probibilidad con Python
FDP_normal = stats.norm(10, 1.2).pdf(x_1) # FDP
plt.plot(x_1, FDP_normal, label='FDP nomal')
plt.title('Función de Densidad de Probabilidad')
plt.ylabel('probabilidad')
plt.xlabel('valores')
plt.show()
# ## Distribuciones Discretas
#
# Las [distribuciones discretas](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad#Distribuciones_de_variable_discreta) son aquellas en las que la variable puede tomar solo algunos valores determinados. Los principales exponentes de este grupo son las siguientes:
# ### Distribución Poisson
#
# La [Distribución Poisson](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_Poisson) esta dada por la formula:
#
# $$p(r; \mu) = \frac{\mu^r e^{-\mu}}{r!}$$
#
# En dónde $r$ es un [entero](https://es.wikipedia.org/wiki/N%C3%BAmero_entero) ($r \ge 0$) y $\mu$ es un [número real](https://es.wikipedia.org/wiki/N%C3%BAmero_real) positivo. La [Distribución Poisson](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_Poisson) describe la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) de encontrar exactamente $r$ eventos en un lapso de tiempo si los acontecimientos se producen de forma independiente a una velocidad constante $\mu$. Es una de las [distribuciones](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad) más utilizadas en [estadística](http://relopezbriega.github.io/tag/estadistica.html) con varias aplicaciones; como por ejemplo describir el número de fallos en un lote de materiales o la cantidad de llegadas por hora a un centro de servicios.
# +
# Graficando Poisson
mu = 7.4 # parametro de forma
poisson = stats.poisson(mu) # Distribución
print(poisson.ppf(0.01))
print(poisson.ppf(0.99))
x = np.arange(poisson.ppf(0.01),
poisson.ppf(0.99))
print(x)
fmp = poisson.pmf(x) # Función de Masa de Probabilidad
plt.plot(x, fmp, '--')
plt.vlines(x, 0, fmp, colors='b', lw=5, alpha=0.5)
plt.title('Distribución Poisson')
plt.ylabel('probabilidad')
plt.xlabel('valores')
plt.show()
# -
# histograma
aleatorios = poisson.rvs(100000) # genera aleatorios
cuenta, cajas, ignorar = plt.hist(aleatorios, 20)
plt.ylabel('frequencia')
plt.xlabel('valores')
plt.title('Histograma Poisson')
plt.show()
# ### Distribución Binomial
#
# La [Distribución Binomial](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_binomial) esta dada por la formula:
#
# $$p(r; N, p) = \left(\begin{array}{c} N \\ r \end{array}\right) p^r(1 - p)^{N - r}
# $$
#
#
# En dónde $r$ con la condición $0 \le r \le N$ y el parámetro $N$ ($N > 0$) son [enteros](https://es.wikipedia.org/wiki/N%C3%BAmero_entero); y el parámetro $p$ ($0 \le p \le 1$) es un [número real](https://es.wikipedia.org/wiki/N%C3%BAmero_real). La [Distribución Binomial](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_binomial) describe la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) de exactamente $r$ éxitos en $N$ pruebas si la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) de éxito en una sola prueba es $p$.
# +
# Graficando Binomial
N, p = 30, 0.5 # parametros de forma
binomial = stats.binom(N, p) # Distribución
# x = np.arange(binomial.ppf(0.01),
# binomial.ppf(0.99))
x=range(0,31)
fmp = binomial.pmf(x) # Función de Masa de Probabilidad
# plt.plot(x, fmp, '--')
plt.step(x, fmp, where='mid', color='blue')
#plt.vlines(x, 0, fmp, colors='b', lw=15, alpha=0.5)
plt.title('Distribución Binomial')
plt.ylabel('probabilidad')
plt.xlabel('valores')
plt.show()
# -
# histograma
aleatorios = binomial.rvs(100000) # genera aleatorios
cuenta, cajas, ignorar = plt.hist(aleatorios, 30)
plt.ylabel('frequencia')
plt.xlabel('valores')
plt.title('Histograma Binomial')
plt.show()
# ### Distribución Geométrica
#
# La [Distribución Geométrica](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_geom%C3%A9trica) esta dada por la formula:
#
# $$p(r; p) = p(1- p)^{r-1}
# $$
#
# En dónde $r \ge 1$ y el parámetro $p$ ($0 \le p \le 1$) es un [número real](https://es.wikipedia.org/wiki/N%C3%BAmero_real). La [Distribución Geométrica](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_geom%C3%A9trica) expresa la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) de tener que esperar exactamente $r$ pruebas hasta encontrar el primer éxito si la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) de éxito en una sola prueba es $p$. Por ejemplo, en un proceso de selección, podría definir el número de entrevistas que deberíamos realizar antes de encontrar al primer candidato aceptable.
# +
# Graficando Geométrica
p = 0.1666 # parametro de forma
geometrica = stats.geom(p) # Distribución
x = np.arange(geometrica.ppf(0.01),
geometrica.ppf(0.99))
print(x)
fmp = geometrica.pmf(x) # Función de Masa de Probabilidad
plt.plot(x, fmp, '--')
plt.vlines(x, 0, fmp, colors='b', lw=5, alpha=0.5)
plt.title('Distribución Geométrica')
plt.ylabel('probabilidad')
plt.xlabel('valores')
plt.show()
# -
# histograma
aleatorios = geometrica.rvs(1000) # genera aleatorios
cuenta, cajas, ignorar = plt.hist(aleatorios, 30)
plt.ylabel('frequencia')
plt.xlabel('valores')
plt.title('Histograma Geométrica')
plt.show()
# ### Distribución Hipergeométrica
#
# La [Distribución Hipergeométrica](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_hipergeom%C3%A9trica) esta dada por la formula:
#
# $$p(r; n, N, M) = \frac{\left(\begin{array}{c} M \\ r \end{array}\right)\left(\begin{array}{c} N - M\\ n -r \end{array}\right)}{\left(\begin{array}{c} N \\ n \end{array}\right)}
# $$
#
#
# En dónde el valor de $r$ esta limitado por $\max(0, n - N + M)$ y $\min(n, M)$ inclusive; y los parámetros $n$ ($1 \le n \le N$), $N$ ($N \ge 1$) y $M$ ($M \ge 1$) son todos [números enteros](https://es.wikipedia.org/wiki/N%C3%BAmero_entero). La [Distribución Hipergeométrica](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_hipergeom%C3%A9trica) describe experimentos en donde se seleccionan los elementos al azar *sin reemplazo* (se evita seleccionar el mismo elemento más de una vez). Más precisamente, supongamos que tenemos $N$ elementos de los cuales $M$ tienen un cierto atributo (y $N - M$ no tiene). Si escogemos $n$ elementos al azar *sin reemplazo*, $p(r)$ es la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) de que exactamente $r$ de los elementos seleccionados provienen del grupo con el atributo.
# Graficando Hipergeométrica
M, n, N = 30, 10, 12 # parametros de forma
hipergeometrica = stats.hypergeom(M, n, N) # Distribución
x = np.arange(0, n+1)
fmp = hipergeometrica.pmf(x) # Función de Masa de Probabilidad
plt.plot(x, fmp, '--')
plt.vlines(x, 0, fmp, colors='b', lw=5, alpha=0.5)
plt.title('Distribución Hipergeométrica')
plt.ylabel('probabilidad')
plt.xlabel('valores')
plt.show()
# histograma
aleatorios = hipergeometrica.rvs(1000) # genera aleatorios
cuenta, cajas, ignorar = plt.hist(aleatorios, 20)
plt.ylabel('frequencia')
plt.xlabel('valores')
plt.title('Histograma Hipergeométrica')
plt.show()
# ### Distribución de Bernoulli
#
# La [Distribución de Bernoulli](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_Bernoulli) esta dada por la formula:
#
# $$p(r;p) = \left\{
# \begin{array}{ll}
# 1 - p = q & \mbox{si } r = 0 \ \mbox{(fracaso)}\\
# p & \mbox{si } r = 1 \ \mbox{(éxito)}
# \end{array}
# \right.$$
#
#
# En dónde el parámetro $p$ es la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) de éxito en un solo ensayo, la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) de fracaso por lo tanto va a ser $1 - p$ (muchas veces expresada como $q$). Tanto $p$ como $q$ van a estar limitados al intervalo de cero a uno. La [Distribución de Bernoulli](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_Bernoulli) describe un experimento probabilístico en donde el ensayo tiene dos posibles resultados, éxito o fracaso. Desde esta [distribución](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad) se pueden deducir varias [Funciones de Densidad de Probabilidad](https://es.wikipedia.org/wiki/Funci%C3%B3n_de_densidad_de_probabilidad) de otras [distribuciones](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad) que se basen en una serie de ensayos independientes.
# Graficando Bernoulli
p = 0.5 # parametro de forma
bernoulli = stats.bernoulli(p)
x = np.arange(-1, 3)
fmp = bernoulli.pmf(x) # Función de Masa de Probabilidad
fig, ax = plt.subplots()
ax.plot(x, fmp, 'bo')
ax.vlines(x, 0, fmp, colors='b', lw=5, alpha=0.5)
ax.set_yticks([0., 0.2, 0.4, 0.6])
plt.title('Distribución Bernoulli')
plt.ylabel('probabilidad')
plt.xlabel('valores')
plt.show()
# +
# histograma
aleatorios = bernoulli.rvs(1000) # genera aleatorios
cuenta, cajas, ignorar = plt.hist(aleatorios, 20)
plt.ylabel('frequencia')
plt.xlabel('valores')
plt.title('Histograma Bernoulli')
plt.show()
# -
# ## Distribuciones continuas
#
# Ahora que ya conocemos las principales [distribuciones discretas](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad#Distribuciones_de_variable_discreta), podemos pasar a describir a las [distribuciones continuas](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad_continua); en ellas a diferencia de lo que veíamos antes, la variable puede tomar cualquier valor dentro de un intervalo específico. Dentro de este grupo vamos a encontrar a las siguientes:
#
# ### Distribución de Normal
#
# La [Distribución Normal](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_normal), o también llamada [Distribución de Gauss](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_normal), es aplicable a un amplio rango de problemas, lo que la convierte en la [distribución](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad) más utilizada en [estadística](http://relopezbriega.github.io/tag/estadistica.html); esta dada por la formula:
#
# $$p(x;\mu, \sigma^2) = \frac{1}{\sigma \sqrt{2 \pi}} e^{\frac{-1}{2}\left(\frac{x - \mu}{\sigma} \right)^2}
# $$
#
# En dónde $\mu$ es el parámetro de ubicación, y va a ser igual a la [media aritmética](https://es.wikipedia.org/wiki/Media_aritm%C3%A9tica) y $\sigma^2$ es el [desvío estándar](https://es.wikipedia.org/wiki/Desviaci%C3%B3n_t%C3%ADpica). Algunos ejemplos de variables asociadas a fenómenos naturales que siguen el modelo de la [Distribución Normal](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_normal) son:
# * características morfológicas de individuos, como la estatura;
# * características sociológicas, como el consumo de cierto producto por un mismo grupo de individuos;
# * características psicológicas, como el cociente intelectual;
# * nivel de ruido en telecomunicaciones;
# * errores cometidos al medir ciertas magnitudes;
# * etc.
# Graficando Normal
mu, sigma = 0, 0.2 # media y desvio estandar
normal = stats.norm(mu, sigma)
x = np.linspace(normal.ppf(0.01),
normal.ppf(0.99), 100)
fp = normal.pdf(x) # Función de Probabilidad
plt.plot(x, fp)
plt.title('Distribución Normal')
plt.ylabel('probabilidad')
plt.xlabel('valores')
plt.show()
# histograma
aleatorios = normal.rvs(1000) # genera aleatorios
cuenta, cajas, ignorar = plt.hist(aleatorios, 20)
plt.ylabel('frequencia')
plt.xlabel('valores')
plt.title('Histograma Normal')
plt.show()
# ### Distribución Uniforme
#
# La [Distribución Uniforme](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_uniforme_discreta) es un caso muy simple expresada por la función:
#
# $$f(x; a, b) = \frac{1}{b -a} \ \mbox{para} \ a \le x \le b
# $$
#
# Su [función de distribución](https://es.wikipedia.org/wiki/Funci%C3%B3n_de_distribuci%C3%B3n) esta entonces dada por:
#
# $$
# p(x;a, b) = \left\{
# \begin{array}{ll}
# 0 & \mbox{si } x \le a \\
# \frac{x-a}{b-a} & \mbox{si } a \le x \le b \\
# 1 & \mbox{si } b \le x
# \end{array}
# \right.
# $$
#
# Todos los valore tienen prácticamente la misma probabilidad.
# Graficando Uniforme
uniforme = stats.uniform()
x = np.linspace(uniforme.ppf(0.01),
uniforme.ppf(0.99), 100)
fp = uniforme.pdf(x) # Función de Probabilidad
fig, ax = plt.subplots()
ax.plot(x, fp, '--')
ax.vlines(x, 0, fp, colors='b', lw=5, alpha=0.5)
ax.set_yticks([0., 0.2, 0.4, 0.6, 0.8, 1., 1.2])
plt.title('Distribución Uniforme')
plt.ylabel('probabilidad')
plt.xlabel('valores')
plt.show()
# histograma
aleatorios = uniforme.rvs(100000) # genera aleatorios
cuenta, cajas, ignorar = plt.hist(aleatorios, 20)
plt.ylabel('frequencia')
plt.xlabel('valores')
plt.title('Histograma Uniforme')
plt.show()
#
# ### Distribución de Log-normal
#
# La [Distribución Log-normal](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_log-normal) esta dada por la formula:
#
# $$p(x;\mu, \sigma) = \frac{1}{ x \sigma \sqrt{2 \pi}} e^{\frac{-1}{2}\left(\frac{\ln x - \mu}{\sigma} \right)^2}
# $$
#
# En dónde la variable $x > 0$ y los parámetros $\mu$ y $\sigma > 0$ son todos [números reales](https://es.wikipedia.org/wiki/N%C3%BAmero_real). La [Distribución Log-normal](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_log-normal) es aplicable a [variables aleatorias](https://es.wikipedia.org/wiki/Variable_aleatoria) que están limitadas por cero, pero tienen pocos valores grandes. Es una [distribución](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad) con [asimetría positiva](https://es.wikipedia.org/wiki/Asimetr%C3%ADa_estad%C3%ADstica). Algunos de los ejemplos en que la solemos encontrar son:
# * El peso de los adultos.
# * La concentración de los minerales en depósitos.
# * Duración de licencia por enfermedad.
# * Distribución de riqueza
# * Tiempos muertos de maquinarias.
# Graficando Log-Normal
sigma = 0.6 # parametro
lognormal = stats.lognorm(sigma)
x = np.linspace(lognormal.ppf(0.01),
lognormal.ppf(0.99), 100)
fp = lognormal.pdf(x) # Función de Probabilidad
plt.plot(x, fp)
plt.title('Distribución Log-normal')
plt.ylabel('probabilidad')
plt.xlabel('valores')
plt.show()
# histograma
aleatorios = lognormal.rvs(1000) # genera aleatorios
cuenta, cajas, ignorar = plt.hist(aleatorios, 20)
plt.ylabel('frequencia')
plt.xlabel('valores')
plt.title('Histograma Log-normal')
plt.show()
# ### Distribución de Exponencial
#
# La [Distribución Exponencial](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_exponencial) esta dada por la formula:
#
# $$p(x;\alpha) = \frac{1}{ \alpha} e^{\frac{-x}{\alpha}}
# $$
#
# En dónde tanto la variable $x$ como el parámetro $\alpha$ son [números reales](https://es.wikipedia.org/wiki/N%C3%BAmero_real) positivos. La [Distribución Exponencial](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_exponencial) tiene bastantes aplicaciones, tales como la desintegración de un átomo radioactivo o el tiempo entre eventos en un proceso de [Poisson](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_Poisson) donde los acontecimientos suceden a una velocidad constante.
# Graficando Exponencial
exponencial = stats.expon()
x = np.linspace(exponencial.ppf(0.01),
exponencial.ppf(0.99), 100)
fp = exponencial.pdf(x) # Función de Probabilidad
plt.plot(x, fp)
plt.title('Distribución Exponencial')
plt.ylabel('probabilidad')
plt.xlabel('valores')
plt.show()
# histograma
aleatorios = exponencial.rvs(1000) # genera aleatorios
cuenta, cajas, ignorar = plt.hist(aleatorios, 20)
plt.ylabel('frequencia')
plt.xlabel('valores')
plt.title('Histograma Exponencial')
plt.show()
# ### Distribución Gamma
#
# La [Distribución Gamma](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_gamma) esta dada por la formula:
#
# $$p(x;a, b) = \frac{a(a x)^{b -1} e^{-ax}}{\Gamma(b)}
# $$
#
# En dónde los parámetros $a$ y $b$ y la variable $x$ son [números reales](https://es.wikipedia.org/wiki/N%C3%BAmero_real) positivos y $\Gamma(b)$ es la [función gamma](https://es.wikipedia.org/wiki/Funci%C3%B3n_gamma). La [Distribución Gamma](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_gamma) comienza en el *origen* de coordenadas y tiene una forma bastante flexible. Otras [distribuciones](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad) son casos especiales de ella.
# Graficando Gamma
a = 2.6 # parametro de forma.
gamma = stats.gamma(a)
x = np.linspace(gamma.ppf(0.01),
gamma.ppf(0.99), 100)
fp = gamma.pdf(x) # Función de Probabilidad
plt.plot(x, fp)
plt.title('Distribución Gamma')
plt.ylabel('probabilidad')
plt.xlabel('valores')
plt.show()
# histograma
aleatorios = gamma.rvs(1000) # genera aleatorios
cuenta, cajas, ignorar = plt.hist(aleatorios, 20)
plt.ylabel('frequencia')
plt.xlabel('valores')
plt.title('Histograma Gamma')
plt.show()
# ### Distribución Beta
#
# La [Distribución Beta](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_beta) esta dada por la formula:
#
# $$p(x;p, q) = \frac{1}{B(p, q)} x^{p-1}(1 - x)^{q-1}
# $$
#
# En dónde los parámetros $p$ y $q$ son [números reales](https://es.wikipedia.org/wiki/N%C3%BAmero_real) positivos, la variable $x$ satisface la condición $0 \le x \le 1$ y $B(p, q)$ es la [función beta](https://es.wikipedia.org/wiki/Funci%C3%B3n_beta). Las aplicaciones de la [Distribución Beta](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_beta) incluyen el modelado de [variables aleatorias](https://es.wikipedia.org/wiki/Variable_aleatoria) que tienen un rango finito de $a$ hasta $b$. Un
# ejemplo de ello es la distribución de los tiempos de actividad en las redes de proyectos. La [Distribución Beta](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_beta) se utiliza también con frecuencia como una [probabilidad a priori](https://es.wikipedia.org/wiki/Probabilidad_a_priori) para proporciones [binomiales]((https://es.wikipedia.org/wiki/Distribuci%C3%B3n_binomial) en el [análisis bayesiano](https://es.wikipedia.org/wiki/Inferencia_bayesiana).
# Graficando Beta
a, b = 2.3, 0.6 # parametros de forma.
beta = stats.beta(a, b)
x = np.linspace(beta.ppf(0.01),
beta.ppf(0.99), 100)
fp = beta.pdf(x) # Función de Probabilidad
plt.plot(x, fp)
plt.title('Distribución Beta')
plt.ylabel('probabilidad')
plt.xlabel('valores')
plt.show()
# histograma
aleatorios = beta.rvs(1000) # genera aleatorios
cuenta, cajas, ignorar = plt.hist(aleatorios, 20)
plt.ylabel('frequencia')
plt.xlabel('valores')
plt.title('Histograma Beta')
plt.show()
# ### Distribución Chi cuadrado
#
# La [Distribución Chi cuadrado](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_%CF%87%C2%B2) esta dada por la función:
#
# $$p(x; n) = \frac{\left(\frac{x}{2}\right)^{\frac{n}{2}-1} e^{\frac{-x}{2}}}{2\Gamma \left(\frac{n}{2}\right)}
# $$
#
# En dónde la variable $x \ge 0$ y el parámetro $n$, el número de grados de libertad, es un [número entero](https://es.wikipedia.org/wiki/N%C3%BAmero_entero) positivo. Una importante aplicación de la [Distribución Chi cuadrado](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_%CF%87%C2%B2) es que cuando un [conjunto de datos](https://es.wikipedia.org/wiki/Conjunto_de_datos) es representado por un modelo teórico, esta [distribución](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad) puede ser utilizada para controlar cuan bien se ajustan los valores predichos por el modelo, y los datos realmente observados.
# Graficando Chi cuadrado
df = 34 # parametro de forma.
chi2 = stats.chi2(df)
x = np.linspace(chi2.ppf(0.01),
chi2.ppf(0.99), 100)
fp = chi2.pdf(x) # Función de Probabilidad
plt.plot(x, fp)
plt.title('Distribución Chi cuadrado')
plt.ylabel('probabilidad')
plt.xlabel('valores')
plt.show()
# histograma
aleatorios = chi2.rvs(1000) # genera aleatorios
cuenta, cajas, ignorar = plt.hist(aleatorios, 20)
plt.ylabel('frequencia')
plt.xlabel('valores')
plt.title('Histograma Chi cuadrado')
plt.show()
# ### Distribución T de Student
#
# La [Distribución t de Student](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_t_de_Student) esta dada por la función:
#
# $$p(t; n) = \frac{\Gamma(\frac{n+1}{2})}{\sqrt{n\pi}\Gamma(\frac{n}{2})} \left( 1 + \frac{t^2}{2} \right)^{-\frac{n+1}{2}}
# $$
#
# En dónde la variable $t$ es un [número real](https://es.wikipedia.org/wiki/N%C3%BAmero_real) y el parámetro $n$ es un [número entero](https://es.wikipedia.org/wiki/N%C3%BAmero_entero) positivo. La [Distribución t de Student](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_t_de_Student) es utilizada para probar si la diferencia entre las *medias* de dos muestras de observaciones es estadísticamente significativa. Por ejemplo, las alturas de una muestra aleatoria de los jugadores de baloncesto podría compararse con las alturas de una muestra aleatoria de jugadores de fútbol; esta [distribución](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad) nos podría ayudar a determinar si un grupo es significativamente más alto que el otro.
# Graficando t de Student
df = 50 # parametro de forma.
t = stats.t(df)
x = np.linspace(t.ppf(0.01),
t.ppf(0.99), 100)
fp = t.pdf(x) # Función de Probabilidad
plt.plot(x, fp)
plt.title('Distribución t de Student')
plt.ylabel('probabilidad')
plt.xlabel('valores')
plt.show()
# histograma
aleatorios = t.rvs(1000) # genera aleatorios
cuenta, cajas, ignorar = plt.hist(aleatorios, 20)
plt.ylabel('frequencia')
plt.xlabel('valores')
plt.title('Histograma t de Student')
plt.show()
# ### Distribución de Pareto
#
# La [Distribución de Pareto](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_Pareto) esta dada por la función:
#
# $$p(x; \alpha, k) = \frac{\alpha k^{\alpha}}{x^{\alpha + 1}}
# $$
#
# En dónde la variable $x \ge k$ y el parámetro $\alpha > 0$ son [números reales](https://es.wikipedia.org/wiki/N%C3%BAmero_real). Esta [distribución](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad) fue introducida por su inventor, [<NAME>](https://es.wikipedia.org/wiki/Vilfredo_Pareto), con el fin de explicar la distribución de los salarios en la sociedad. La [Distribución de Pareto](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_Pareto) se describe a menudo como la base de la [regla 80/20](https://es.wikipedia.org/wiki/Principio_de_Pareto). Por ejemplo, el 80% de las quejas de los clientes con respecto al funcionamiento de su vehículo por lo general surgen del 20% de los componentes.
# Graficando Pareto
k = 2.3 # parametro de forma.
pareto = stats.pareto(k)
x = np.linspace(pareto.ppf(0.01),
pareto.ppf(0.99), 100)
fp = pareto.pdf(x) # Función de Probabilidad
plt.plot(x, fp)
plt.title('Distribución de Pareto')
plt.ylabel('probabilidad')
plt.xlabel('valores')
plt.show()
# histograma
aleatorios = pareto.rvs(1000) # genera aleatorios
cuenta, cajas, ignorar = plt.hist(aleatorios, 20)
plt.ylabel('frequencia')
plt.xlabel('valores')
plt.title('Histograma de Pareto')
plt.show()
# ### Distribuciones simetricas y asimetricas
#
# Una distribución es simétrica cuando moda, mediana y media coinciden aproximadamente en sus valores. Si una distribución es simétrica, existe el mismo número de valores a la derecha que a la izquierda de la media, por tanto, el mismo número de desviaciones con signo positivo que con signo negativo.
#
# Una distribución tiene [asimetria](https://es.wikipedia.org/wiki/Asimetr%C3%ADa_estad%C3%ADstica) positiva (o a la derecha) si la "cola" a la derecha de la media es más larga que la de la izquierda, es decir, si hay valores más separados de la media a la derecha. De la misma forma una distribución tiene [asimetria](https://es.wikipedia.org/wiki/Asimetr%C3%ADa_estad%C3%ADstica) negativa (o a la izquierda) si la "cola" a la izquierda de la media es más larga que la de la derecha, es decir, si hay valores más separados de la media a la izquierda.
#
# Las distribuciones asimétricas suelen ser problemáticas, ya que la mayoría de los métodos estadísticos suelen estar desarrollados para distribuciones del tipo [normal](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_normal). Para salvar estos problemas se suelen realizar transformaciones a los datos para hacer a estas distribuciones más simétricas y acercarse a la [distribución normal](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_normal).
# Dibujando la distribucion Gamma
x = stats.gamma(3).rvs(5000)
gamma = plt.hist(x, 70, histtype="stepfilled", alpha=.7)
# En este ejemplo podemos ver que la [distribución gamma](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_gamma) que dibujamos tiene una [asimetria](https://es.wikipedia.org/wiki/Asimetr%C3%ADa_estad%C3%ADstica) positiva.
# + jupyter={"outputs_hidden": false}
# Calculando la simetria con scipy
stats.skew(x)
# -
# ## Cuartiles y diagramas de cajas
#
# Los **[cuartiles](https://es.wikipedia.org/wiki/Cuartil)** son los tres valores de la variable estadística que dividen a un [conjunto de datos](https://es.wikipedia.org/wiki/Conjunto_de_datos) ordenados en cuatro partes iguales. Q1, Q2 y Q3 determinan los valores correspondientes al 25%, al 50% y al 75% de los datos. Q2 coincide con la <a href="https://es.wikipedia.org/wiki/Mediana_(estad%C3%ADstica)">mediana</a>.
#
# Los [diagramas de cajas](https://es.wikipedia.org/wiki/Diagrama_de_caja) son una presentación visual que describe varias características importantes al mismo tiempo, tales como la dispersión y simetría. Para su realización se representan los tres cuartiles y los valores mínimo y máximo de los datos, sobre un rectángulo, alineado horizontal o verticalmente. Estos gráficos nos proporcionan abundante información y son sumamente útiles para encontrar [valores atípicos](https://es.wikipedia.org/wiki/Valor_at%C3%ADpico) y comparar dos [conjunto de datos](https://es.wikipedia.org/wiki/Conjunto_de_datos).
#
#
# <img alt="diagrama de cajas" title="Diagrama de cajas" src="http://relopezbriega.github.io/images/diagCajas.png" width="600">
# El mínimo que aparece en la imagen previa es el mínimo intercuartílico, no el mínimo como tal. El cuartil es lo que se encuentra dentro de la caja roja, son los bombones del tema
# +
# Ejemplo de grafico de cajas en python
datos_1 = np.random.normal(125, 80, 200)
datos_2 = np.random.normal(80, 30, 200)
datos_3 = np.random.normal(90, 20, 200)
datos_4 = np.random.normal(70, 25, 200)
datos_graf = [datos_1, datos_2, datos_3, datos_4]
# Creando el objeto figura
fig = plt.figure(1, figsize=(9, 6))
# Creando el subgrafico
ax = fig.add_subplot(111)
# creando el grafico de cajas
bp = ax.boxplot(datos_graf)
# visualizar mas facile los atípicos
for flier in bp['fliers']:
flier.set(marker='o', markerfacecolor='red', alpha=1)
# los puntos aislados son valores atípicos
# -
# ## ¿Cómo elegir la distribución que mejor se ajusta a mis datos?
#
# Ahora ya tenemos un conocimiento general de las principales [distribuciones](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad) con que nos podemos encontrar; pero ¿cómo determinamos que [distribución](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad) debemos utilizar?
#
# Un modelo que podemos seguir cuando nos encontramos con datos que necesitamos ajustar a una [distribución](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad), es comenzar con los datos sin procesar y responder a cuatro preguntas básicas acerca de los mismos, que nos pueden ayudar a caracterizarlos. La **primer pregunta** se refiere a si los datos **pueden tomar valores [discretos](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad#Distribuciones_de_variable_discreta) o [continuos](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad_continua)**. **La segunda pregunta** que nos debemos hacer, hace referencia a la **[simetría](https://es.wikipedia.org/wiki/Asimetr%C3%ADa_estad%C3%ADstica) de los datos** y si hay asimetría, en qué dirección se encuentra; en otras palabras, son los [valores atípicos](https://es.wikipedia.org/wiki/Valor_at%C3%ADpico) positivos y negativos igualmente probables o es uno más probable que el otro. **La tercer pregunta** abarca los **límites superiores e inferiores en los datos**; hay algunos datos, como los ingresos, que no pueden ser inferiores a cero, mientras que hay otros, como los márgenes de operación que no puede exceder de un valor (100%). **La última pregunta** se refiere a la **posibilidad de observar valores extremos** en la [distribución](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad); en algunos casos, los valores extremos ocurren con muy poca frecuencia, mientras que en otros, se producen con mayor frecuencia.
# Este proceso, lo podemos resumir en el siguiente gráfico:
#
# <img alt="Distribuciones estadísticas" title="Distribuciones estadísticas" src="http://relopezbriega.github.io/images/distributions_choice.png
# " >
#
# Con la ayuda de estas preguntas fundamentales, más el conocimiento de las distintas [distribuciones](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_de_probabilidad) deberíamos estar en condiciones de poder caracterizar cualquier [conjunto de datos](https://es.wikipedia.org/wiki/Conjunto_de_datos).
#
# *Esta notebook fue creada originalmente como un blog post por [<NAME>](http://relopezbriega.com.ar/)
# # Independencia, la ley de grandes números y el teorema del límite central
#
# Una de las cosas más fascinantes sobre el estudio de la [teoría de probabilidad](https://es.wikipedia.org/wiki/Teor%C3%ADa_de_la_probabilidad) es que si bien el comportamiento de un evento individual es totalmente impredecible, el comportamiento de una cantidad suficientemente grande de eventos se puede predecir con un alto grado de certeza!.
# Si tomamos el caso clásico del lanzamiento de una moneda, no podemos predecir con exactitud cuantas caras podemos obtener luego de 10 tiradas, tal vez el azar haga que obtengamos 7, 10, o 3 caras, dependiendo de con cuanta suerte nos encontremos; pero si repetimos el lanzamiento un millón de veces, casi con seguridad que la cantidad de caras se aproximará a la verdadera [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) subyacente del experimento, es decir, al 50% de los lanzamientos. Este comportamiento es lo que en la [teoría de probabilidad](https://es.wikipedia.org/wiki/Teor%C3%ADa_de_la_probabilidad) se conoce con el nombre de [ley de grandes números](https://es.wikipedia.org/wiki/Ley_de_los_grandes_n%C3%BAmeros); pero antes de poder definir esta ley, primero debemos describir otro concepto también muy importante, la <a href="https://es.wikipedia.org/wiki/Independencia_(probabilidad)">independencia</a> de los [eventos](https://es.wikipedia.org/wiki/Evento_aleatorio) .
# ### El concepto de independencia
#
# En [teoría de probabilidad](https://es.wikipedia.org/wiki/Teor%C3%ADa_de_la_probabilidad), podemos decir que dos [eventos](https://es.wikipedia.org/wiki/Evento_aleatorio) son <a href="https://es.wikipedia.org/wiki/Independencia_(probabilidad)">independientes</a> cuando la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) de cada uno de ellos no se ve afecta porque el otro evento ocurra, es decir que no existe ninguna relación entre los [eventos](https://es.wikipedia.org/wiki/Evento_aleatorio). En el lanzamiento de la moneda; la moneda no sabe, ni le interesa saber si el resultado del lanzamiento anterior fue cara; cada lanzamiento es un suceso totalmente aislado el uno del otro y la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) del resultado va a ser siempre 50% en cada lanzamiento.
#
# ### Definiendo la ley de grandes números
#
# Ahora que ya conocemos el concepto de <a href="https://es.wikipedia.org/wiki/Independencia_(probabilidad)">independencia</a>, estamos en condiciones de dar una definición más formal de la [ley de grandes números](https://es.wikipedia.org/wiki/Ley_de_los_grandes_n%C3%BAmeros), que junto con el [Teorema del límite central](https://es.wikipedia.org/wiki/Teorema_del_l%C3%ADmite_central), constituyen los cimientos de la [teoría de probabilidad](https://es.wikipedia.org/wiki/Teor%C3%ADa_de_la_probabilidad). Podemos formular esta ley de la siguiente manera: **si se repite un [experimento aleatorio](https://es.wikipedia.org/wiki/Experimento_aleatorio), bajo las mismas condiciones, un número ilimitado de veces; y si estas repeticiones son <a href="https://es.wikipedia.org/wiki/Independencia_(probabilidad)">independientes</a> la una de la otra, entonces la frecuencia de veces que un evento $A$ ocurra, convergerá con [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) 1 a un número que es igual a la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) de que $A$ ocurra en una sola repetición del experimento.** Lo que esta ley nos enseña, es que la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) subyacente de cualquier suceso aleatorio puede ser aprendido por medio de la experimentación, simplemente tendríamos que repetirlo una cantidad suficientemente grande de veces!. Un error que la gente suele cometer y asociar a esta ley, es la idea de que un evento tiene más posibilidades de ocurrir porque ha o no ha ocurrido recientemente. Esta idea de que las chances de un evento con una [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) fija, aumentan o disminuyen dependiendo de las ocurrencias recientes del evento, es un error que se conoce bajo el nombre de la [falacia del apostador](https://es.wikipedia.org/wiki/Falacia_del_apostador).
#
# Para entender mejor la [ley de grandes números](https://es.wikipedia.org/wiki/Ley_de_los_grandes_n%C3%BAmeros), experimentemos con algunos ejemplos en [Python](https://www.python.org/). Utilicemos nuevamente el ejemplo del lanzamiento de la moneda, en el primer ejemplo, la moneda va a tener la misma posibilidad de caer en cara o seca; mientras que en el segundo ejemplo, vamos a modificar la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) de la moneda para que caiga cara solo en 1 de 6 veces.
# +
import matplotlib.pyplot as plt
import numpy as np # importando numpy
import pandas as pd # importando pandas
np.random.seed(2131982) # para poder replicar el random
# %matplotlib inline
# Ejemplo ley de grandes números
# moneda p=1/2 cara=1 seca=0
total_lanzamientos = 10000
resultados = []
for lanzamientos in range(1, total_lanzamientos):
lanzamientos = np.random.choice([0,1], lanzamientos)
caras = lanzamientos.mean()
resultados.append(caras)
# graficamente
df = pd.DataFrame({ 'lanzamientos' : resultados})
df.plot(title='Ley de grandes números',color='r',figsize=(8, 6))
plt.axhline(0.5)
plt.xlabel("Número de lanzamientos")
plt.ylabel("frecuencia caras")
plt.show()
# +
# moneda p=1/6 cara=1 seca=0
resultados = []
lanzamientos_totales = 10000
p0 = 5/6
p1 = 1/6
for lanzamientos in range(1, lanzamientos_totales):
lanzamientos = np.random.choice([0,1], lanzamientos, p=[p0, p1])
caras = lanzamientos.mean()
resultados.append(caras)
# graficamente
df = pd.DataFrame({ 'lanzamientos' : resultados})
df.plot(title='Ley de grandes números',color='r',figsize=(8, 6))
plt.axhline(p1)
plt.xlabel("Número de lanzamientos")
plt.ylabel("frecuencia caras")
plt.show()
# -
# Como estos ejemplos nos muestran, al comienzo, la frecuencia en que vamos obteniendo caras va variando considerablemente, pero a medida que aumentamos el número de repeticiones, la frecuencia de caras se va estabilizando en la [probabilidad](https://es.wikipedia.org/wiki/Probabilidad) subyacente el evento, 1 en 2 para el primer caso y 1 en 6 para el segundo ejemplo. En los gráficos podemos ver claramente el comportamiento de la ley.
#
# ### El Teorema del límite central
#
# El otro gran teorema de la [teoría de probabilidad](https://es.wikipedia.org/wiki/Teor%C3%ADa_de_la_probabilidad) es el [Teorema del límite central](https://es.wikipedia.org/wiki/Teorema_del_l%C3%ADmite_central). Este teorema establece que la suma o el promedio de casi cualquier conjunto de variables <a href="https://es.wikipedia.org/wiki/Independencia_(probabilidad)">independientes</a> generadas al azar se aproximan a la [Distribución Normal](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_normal). El [Teorema del límite central](https://es.wikipedia.org/wiki/Teorema_del_l%C3%ADmite_central) explica por qué la [Distribución Normal](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_normal) surge tan comúnmente y por qué es generalmente una aproximación excelente para la [media](https://es.wikipedia.org/wiki/Media_aritm%C3%A9tica) de casi cualquier colección de datos. Este notable hallazgo se mantiene verdadero sin importar la forma que adopte la [distribución de datos](http://relopezbriega.github.io/blog/2016/06/29/distribuciones-de-probabilidad-con-python/) que tomemos. Para ilustrar también este teorema, recurramos a un poco más de [Python](https://www.python.org/).
# Ejemplo teorema del límite central
for size in [10, 100, 1000, 10000]:
muestra_binomial = []
muestra_exp = []
muestra_possion = []
muestra_geometric = []
mu = .9
lam = 1.0
print(f'\t size={size} \n')
for i in range(1,size):
muestra = np.random.binomial(1, mu, size=size)
muestra_binomial.append(muestra.mean())
muestra = np.random.exponential(scale=2.0,size=size)
muestra_exp.append(muestra.mean())
muestra = np.random.geometric(p=.5, size=size)
muestra_geometric.append(muestra.mean())
muestra = np.random.poisson (lam=lam, size=size)
muestra_possion.append(muestra.mean())
df = pd.DataFrame({ 'binomial' : muestra_binomial,
'poission' : muestra_possion,
'geometrica' : muestra_geometric,
'exponencial' : muestra_exp})
fig, axes = plt.subplots(nrows=2, ncols=2, figsize=(10,10))
df['binomial'].plot(kind='hist', ax=axes[0,0], bins=1000, color='red')
df['exponencial'].plot(kind='hist', ax=axes[0,1],bins=1000, color='red')
df['poission'].plot(kind='hist', ax=axes[1,0],bins=1000, color='red')
df['geometrica'].plot(kind='hist', ax=axes[1,1],bins=1000, color='red')
axes[0,0].set_title('Binomial')
axes[0,1].set_title('Poisson')
axes[1,0].set_title('Geométrica')
axes[1,1].set_title('Exponencial')
plt.show()
# # La estadística bayesiana
#
# La [estadística bayesiana](https://es.wikipedia.org/wiki/Estad%C3%ADstica_bayesiana) es un subconjunto del campo de la [estadística](http://es.wikipedia.org/wiki/Estad%C3%ADstica) en la que la evidencia sobre el verdadero estado de las cosas se expresa en términos de grados de creencia. Esta filosofía de tratar a las creencias como probabilidad es algo natural para los seres humanos. Nosotros la utilizamos constantemente a medida que interactuamos con el mundo y sólo vemos verdades parciales; necesitando reunir pruebas para formar nuestras creencias.
#
# La diferencia fundamental entre la [estadística clásica](http://es.wikipedia.org/wiki/Estad%C3%ADstica) (frecuentista) y la [bayesiana](https://es.wikipedia.org/wiki/Estad%C3%ADstica_bayesiana) es el concepto de
# [probabilidad](https://es.wikipedia.org/wiki/Probabilidad). Para la [estadística clásica](http://es.wikipedia.org/wiki/Estad%C3%ADstica) es un concepto objetivo, que se encuentra en la naturaleza,
# mientras que para la [estadística bayesiana](https://es.wikipedia.org/wiki/Estad%C3%ADstica_bayesiana) se encuentra en el observador, siendo así un concepto subjetivo. De este modo, en [estadística clásica](http://es.wikipedia.org/wiki/Estad%C3%ADstica) solo se toma como fuente de información las muestras
# obtenidas. En el caso [bayesiano](https://es.wikipedia.org/wiki/Estad%C3%ADstica_bayesiana), sin embargo, además de la muestra también juega un papel fundamental la información previa o externa que se posee en relación a los fenómenos que se tratan de modelar.
#
# La [estadística bayesiana](https://es.wikipedia.org/wiki/Estad%C3%ADstica_bayesiana) está demostrando su utilidad en ciertas estimaciones basadas en el conocimiento subjetivo a priori y el hecho de permitir revisar esas estimaciones en función de la evidencia empírica es lo que está abriendo nuevas formas de hacer conocimiento. Una aplicación de esto son los [clasificadores bayesianos](https://es.wikipedia.org/wiki/Clasificador_bayesiano_ingenuo) que son frecuentemente usados en implementaciones de filtros de correo basura, que se adaptan con el uso.
# +
sns.set_context(rc={"figure.figsize": (11, 9)})
plt.figure(figsize=(20,10))
sns.set()
dist = stats.beta
n_trials = [0, 1, 2, 3, 4, 5, 8, 15, 50, 500]
data = stats.bernoulli.rvs(0.5, size=n_trials[-1])
x = np.linspace(0, 1, 100)
for k, N in enumerate(n_trials):
sx = plt.subplot(len(n_trials) / 2, 2, k + 1)
plt.xlabel("$p$, probabilidad de cara") \
if k in [0, len(n_trials) - 1] else None
plt.setp(sx.get_yticklabels(), visible=False)
heads = data[:N].sum()
y = dist.pdf(x, 1 + heads, 1 + N - heads)
plt.plot(x, y, label="lanzamientos observados %d,\n %d caras" % (N, heads))
plt.fill_between(x, 0, y, color="#348ABD", alpha=0.4)
plt.vlines(0.5, 0, 4, color="k", linestyles="--", lw=1)
leg = plt.legend()
leg.get_frame().set_alpha(0.4)
plt.autoscale(tight=True)
plt.suptitle("Actualizacion Bayesiana de probabilidades posterios",
y=1.02,
fontsize=15)
plt.tight_layout()
# -
# # El Teorema de Bayes
#
# [Thomas Bayes](https://es.wikipedia.org/wiki/Thomas_Bayes) fue un ministro presbiteriano y matemático inglés que estudió la relación íntima que existe entre la [probabilidad](http://relopezbriega.github.io/blog/2016/11/26/introduccion-a-la-teoria-de-probabilidad-con-python/), la predicción y el progreso científico. Su trabajo se centró principalmente en cómo formulamos nuestras creencias probabilísticas sobre el mundo que nos rodea cuando nos encontramos con nuevos datos o evidencias. El argumento de [Bayes](https://es.wikipedia.org/wiki/Thomas_Bayes) no es que el mundo es intrínsecamente probabilístico o incierto, ya que él era un creyente en la divina perfección; sino que aprendemos sobre el mundo a través de la aproximación, acercándonos cada vez más a la verdad a medida que recogemos más evidencias. Este argumento lo expresó matemáticamente a través de su famoso [teorema](https://es.wikipedia.org/wiki/Teorema_de_Bayes):
#
#
# $$P(H|D) = \frac{P(D|H)P(H)}{P(D)}
# $$
#
#
# En donde:
#
# * $P(H)$ es el **[a priori](https://es.wikipedia.org/wiki/Probabilidad_a_priori)**, la forma de introducir conocimiento previo sobre los valores que puede tomar la hipótesis. A veces cuando no sabemos demasiado se suelen usar *a prioris* que asignan igual probabilidad a todos los valores de la hipótesis; otras veces se puede elegir *a prioris* que restrinjan los valores a rangos razonables, por ejemplo solo valores positivos; y otras veces contamos con información mucho más precisa, como experimentos previos o límites impuesto por alguna teoría.
#
#
# * $P(D|H)$ es el **likelihood**, la forma de incluir nuestros datos en el análisis. Es una expresión matemática que especifica la plausibilidad de los datos. A medida que la cantidad de datos aumenta, el *likelihood* tiene cada vez más peso en los resultados. Debemos tener en cuenta que si bien el *likelihood* se asemeja a una *probabilidad*, en realidad no lo es; el *likelihood* de una hipótesis $H$, dados los datos $D$ va a ser proporcional a la *probabilidad* de obtener $D$ dado que $H$ es verdadera. Como el *likelihood* no es una *probabilidad* tampoco tiene que respetar las leyes de las probabilidades y por lo tanto no necesariamente tiene que sumar 1.
#
#
# * $P(H|D)$ es el **[a posteriori](https://es.wikipedia.org/wiki/Probabilidad_a_posteriori)**, la [distribución de probabilidad](http://relopezbriega.github.io/blog/2016/06/29/distribuciones-de-probabilidad-con-python/) final para la hipótesis. Es la consecuencia lógica de haber usado un conjunto de datos, un *likelihood* y un *a priori*. Se lo suele pensar como la versión actualizada del *a priori* luego de que hemos agregado los datos adicionales.
#
#
# * $P(D)$ es el **likelihood marginal** o **evidencia**, la probabilidad de observar los datos $D$ promediado sobre todas las posibles hipótesis $H$. En general, la *evidencia* puede ser vista como una simple constante de normalización que en la mayoría de los problemas prácticos puede omitirse sin demasiada perdida de generalidad.
#
# Si los fundamentos filosóficos del [teorema de Bayes](https://es.wikipedia.org/wiki/Teorema_de_Bayes) son sorprendentemente ricos, sus matemáticas son increíblemente simples. En su forma más básica, no es más que una expresión algebraica con tres variables conocidas y una incógnita; y que trabaja con [probabilidades condicionales](https://es.wikipedia.org/wiki/Probabilidad_condicionada); nos dice la probabilidad de que una hipótesis $H$ sea verdadera si algún evento $D$ ha sucedido. El [teorema de Bayes](https://es.wikipedia.org/wiki/Teorema_de_Bayes) es útil porque lo que normalmente sabemos es la [probabilidad](http://relopezbriega.github.io/blog/2016/11/26/introduccion-a-la-teoria-de-probabilidad-con-python/) de los efectos dados las causas, pero lo que queremos saber es la [probabilidad](http://relopezbriega.github.io/blog/2016/11/26/introduccion-a-la-teoria-de-probabilidad-con-python/) de las causas dadas los efectos. Por ejemplo, podemos saber cual es el porcentaje de pacientes con gripe que tiene fiebre, pero lo que realmente queremos saber es la probabilidad de que un paciente con fiebre tenga gripe. El [teorema de Bayes](https://es.wikipedia.org/wiki/Teorema_de_Bayes) nos permite ir de uno a otro con suma facilidad.
# bien1/2 -2: bien 5/6 | mal 1/6 || mal1/2 -2: mal 5/6 | bien 1/6
# P(bien/mal) = p(mal/bien) * p(bien) / p(mal) = 1/6
# ### La inferencia Bayesiana
#
# Toda forma de [inferencia](https://es.wikipedia.org/wiki/Inferencia) que realicemos sobre el mundo que nos rodea, debe indefectiblemente lidiar con la *incertidumbre*. Existen por lo menos, tres tipos de *incertidumbre* con la que nos debemos enfrentar:
#
# * **Ignorancia**, los límites de nuestro conocimiento nos llevan a ser ignorantes sobre muchas cosas.
# * **Aleatoriedad**, es imposible negar la influencia del azar en casi todo lo que nos rodea; incluso aunque podamos saber todo sobre una moneda y la forma de lanzarla, es imposible predecir con anterioridad si va a caer cara o seca.
# * **Vaguedad**, muchos de los conceptos que utilizamos en nuestro pensamiento tienen cierto grado de subjetividad en su definición. ¿cómo calificaríamos si una persona es valiente o no?. Cada uno de nosotros puede tener una apreciación diferente del concepto de valentía.
#
# La [inferencia bayesiana](https://es.wikipedia.org/wiki/Inferencia_bayesiana) es la filosofía que afirma que para entender la opinión humana como debe ser, limitada por la ignorancia y la incertidumbre; debemos utilizar al [cálculo de probabilidad](http://relopezbriega.github.io/blog/2016/11/26/introduccion-a-la-teoria-de-probabilidad-con-python/) como la herramienta más importante para representar la fortaleza de nuestras creencias.
#
# En esencia, la [inferencia bayesiana](https://es.wikipedia.org/wiki/Inferencia_bayesiana) combina nuestra experiencia previa, en la forma de la *probabilidad [a priori](https://es.wikipedia.org/wiki/Probabilidad_a_priori)*; con los datos observados, en la forma del *likelihood*; para interpretarlos y arribar a una *probabilidad [a posteriori](https://es.wikipedia.org/wiki/Probabilidad_a_posteriori)*. La [inferencia bayesiana](https://es.wikipedia.org/wiki/Inferencia_bayesiana) no nos va a garantizar que podamos alcanzar la respuesta correcta. En su lugar, nos va a proporcionar la [probabilidad](http://relopezbriega.github.io/blog/2016/11/26/introduccion-a-la-teoria-de-probabilidad-con-python/) de que cada una de un número de respuestas alternativas, sea verdadera. Y luego podemos utilizar esta información para encontrar la respuesta que más probablemente sea la correcta. En otras palabras, nos proporciona un mecanismo para hacer una especie de *adivinación basada en información*.
#
#
# ### Bayes en el diagnostico médico
#
# Para que quede más claro, ilustremos la aplicación de la [inferencia bayesiana](https://es.wikipedia.org/wiki/Inferencia_bayesiana) con un simple ejemplo del diagnostico médico, uno de los campos dónde más éxito ha tenido. Supongamos que nos hicimos un estudio y nos ha dado positivo para una rara enfermedad que solo el 0.3 % de la población tiene. La tasa de efectividad de este estudio es del 99 %, es decir, que solo da [falsos positivos](https://es.wikipedia.org/wiki/Errores_de_tipo_I_y_de_tipo_II) en el 1 % de los casos. ¿Cuán probable es que realmente tengamos la enfermedad?.
#
# En un principio, nos veríamos tentados a responder que hay un 99 % de probabilidad de que tengamos la enfermedad; pero en este caso nos estaríamos olvidando del concepto importante del [a priori](https://es.wikipedia.org/wiki/Probabilidad_a_priori). Sabemos con anterioridad que la enfermedad es extremadamente rara (solo el 0.3 % la tiene); si incluimos esta información previa en nuestro cálculo de probabilidad y aplicamos el [teorema de Bayes](https://es.wikipedia.org/wiki/Teorema_de_Bayes) podemos llegar a una conclusión totalmente distinta.
#
#
# $$ P(\text{ enfermedad | pos}) = \frac{P(\text{ pos | enfermedad})P( \text{enfermedad})}{P(\text{pos})}$$
# +
# Ejemplo simple teorema de Bayes aplicado a estimación de un sólo parámetro.
a_priori = 0.003
likelihood = 0.99
evidencia = 0.01
a_posteriori = likelihood * a_priori / evidencia
a_posteriori
# -
# Como vemos, luego de aplicar el [teorema de Bayes](https://es.wikipedia.org/wiki/Teorema_de_Bayes) llegamos a la conclusión de que en realidad nuestra probabilidad de estar realmente enfermo es de sólo 30 % y no de 99 %, ya que podemos ser uno de los [falsos positivos](https://es.wikipedia.org/wiki/Errores_de_tipo_I_y_de_tipo_II) del estudio y la enfermedad es realmente muy rara. Como este ejemplo demuestra, la inclusión del [a priori](https://es.wikipedia.org/wiki/Probabilidad_a_priori) es sumamente importante para la [inferencia bayesiana](https://es.wikipedia.org/wiki/Inferencia_bayesiana), por lo cual también debemos ser sumamente cuidadosos a la hora de elegirla. **Cuando nuestra [a priori](https://es.wikipedia.org/wiki/Probabilidad_a_priori) es fuerte, puede ser sorprendentemente resistente frente a nuevas evidencias.**
# # Redes Bayesianas
#
# El [teorema de Bayes](https://es.wikipedia.org/wiki/Teorema_de_Bayes) nos permite actualizar las probabilidades de variables cuyo estado no hemos observado dada una serie de nuevas observaciones. Las [redes bayesianas](https://es.wikipedia.org/wiki/Red_bayesiana) automatizan este proceso, permitiendo que el razonamiento avance en cualquier dirección a través de la red de variables. Las [redes bayesianas](https://es.wikipedia.org/wiki/Red_bayesiana) están constituidas por una estructura en forma de [grafo](https://es.wikipedia.org/wiki/Grafo), en la que cada <a href="https://es.wikipedia.org/wiki/V%C3%A9rtice_(teor%C3%ADa_de_grafos)">nodo</a> representa [variables aleatorias](https://es.wikipedia.org/wiki/Variable_aleatoria) (discretas o continuas) y cada <a href="https://es.wikipedia.org/wiki/Arista_(teor%C3%ADa_de_grafos)">arista</a> representa las conexiones directas entre ellas. Estas conexiones suelen representar relaciones de causalidad. Adicionalmente, las [redes bayesianas](https://es.wikipedia.org/wiki/Red_bayesiana) también modelan el peso cuantitativo de las conexiones entre las variables, permitiendo que las *creencias probabilísticas* sobre ellas se actualicen automáticamente a medida que se disponga de nueva información.
# Al construir una [red bayesiana](https://es.wikipedia.org/wiki/Red_bayesiana), los principales problemas de modelización que surgen son:
# 1. ¿Cuáles son las variables? ¿Cuáles son sus valores / estados?
# 2. ¿Cuál es la estructura del [grafo](https://es.wikipedia.org/wiki/Grafo)?
# 3. ¿Cuáles son los parámetros (probabilidades)?
#
# Profundicemos un poco en cada uno de estos puntos.
#
# ### Nodos y variables
# Lo primero que debemos hacer es identificar las variables de interés. Sus valores deben ser mutuamente excluyentes y exhaustivos. Los tipos de <a href="https://es.wikipedia.org/wiki/V%C3%A9rtice_(teor%C3%ADa_de_grafos)">nodos</a> discretos más comunes son:
#
# * **Nodos booleanos**, que representan proposiciones tomando los valores binarios Verdadero (V) y Falso (F). En el dominio del diagnóstico médico, por ejemplo, un <a href="https://es.wikipedia.org/wiki/V%C3%A9rtice_(teor%C3%ADa_de_grafos)">nodo</a> llamado "Cáncer" podría representar la proposición del que paciente tenga cáncer.
# * **Valores ordenados** Por ejemplo, un <a href="https://es.wikipedia.org/wiki/V%C3%A9rtice_(teor%C3%ADa_de_grafos)">nodo</a> "Contaminación" podría representar la exposición de un paciente a la contaminación del ambiente y tomar los valores {alta, baja}.
# * **Valores enteros**. Por ejemplo, un <a href="https://es.wikipedia.org/wiki/V%C3%A9rtice_(teor%C3%ADa_de_grafos)">nodo</a> llamado "Edad" puede representar la edad de un paciente y tener valores posibles de 1 a 120.
#
# Lo importante es elegir valores que representen el dominio de manera eficiente, pero con suficiente detalle para realizar el razonamiento requerido.
#
# ### Estructura
# La estructura o topología de la [red](https://es.wikipedia.org/wiki/Red_bayesiana) debe captar las relaciones cualitativas entre las variables. En particular, dos nodos deben conectarse directamente si uno afecta o causa al otro, con la <a href="https://es.wikipedia.org/wiki/Arista_(teor%C3%ADa_de_grafos)">arista</a> indicando la dirección del efecto. Por lo tanto, en nuestro ejemplo de diagnóstico médico, podríamos preguntarnos qué factores afectan la probabilidad de tener cáncer. Si la respuesta es "Contaminación y Fumar", entonces deberíamos agregar <a href="https://es.wikipedia.org/wiki/Arista_(teor%C3%ADa_de_grafos)">aristas</a> desde "Contaminación" y desde "Fumador" hacia el <a href="https://es.wikipedia.org/wiki/V%C3%A9rtice_(teor%C3%ADa_de_grafos)">nodo</a> "Cáncer". Del mismo modo, tener cáncer afectará la respiración del paciente y las posibilidades de tener un resultado positivo de rayos X. Por lo tanto, también podemos agregar <a href="https://es.wikipedia.org/wiki/Arista_(teor%C3%ADa_de_grafos)">aristas</a> de "Cáncer" a "Disnea" y "RayosX".
#
# Es deseable construir [redes bayesianas](https://es.wikipedia.org/wiki/Red_bayesiana) lo más compactas posibles por tres razones. Primero, mientras más compacto es el *modelo*, es más fácil de manejar. Segundo, cuando las redes se vuelven demasiado densas, fallan en representar la independencia en forma explícita. Y Tercero, las redes excesivamente densas no suelen representar las dependencias causales del dominio.
#
# ### Probabilidades condicionales
# Una vez que tenemos definida la estructura de la [red bayesiana](https://es.wikipedia.org/wiki/Red_bayesiana), el siguiente paso es cuantificar las relaciones entre los nodos interconectados; esto se hace especificando una [probabilidad condicional](https://es.wikipedia.org/wiki/Probabilidad_condicionada) para cada nodo.
# Primero, para cada nodo necesitamos mirar todas las posibles combinaciones de valores de los nodos padres.
# Por ejemplo, continuando con el ejemplo del diagnostico del cáncer, si tomamos el <a href="https://es.wikipedia.org/wiki/V%C3%A9rtice_(teor%C3%ADa_de_grafos)">nodo</a> "Cáncer" con sus dos <a href="https://es.wikipedia.org/wiki/V%C3%A9rtice_(teor%C3%ADa_de_grafos)">nodos</a> padres "Contaminación" y "Fumador" podemos calcular los posibles valores conjuntos { (A, V), (A, F), (B, V), (B, F)}. La tabla de probabilidad condicional especifica para cada uno de estos casos podría ser la siguiente: {0,05, 0,02, 0,03, 0,001}. Con estos datos, ya estamos en condiciones de representar el [grafo](https://es.wikipedia.org/wiki/Grafo) de la [red bayesiana](https://es.wikipedia.org/wiki/Red_bayesiana) de nuestro ejemplo.
#
# <img alt="Red Bayesiana" title="Red Bayesiana" src="http://relopezbriega.github.io/images/red_bayes.png" >
#
# ### Razonando con redes Bayesianas
# La tarea básica de cualquier sistema de [inferencia probabilística](https://es.wikipedia.org/wiki/Inferencia) es la de obtener la [distribución](http://relopezbriega.github.io/blog/2016/06/29/distribuciones-de-probabilidad-con-python/) [a posteriori](https://es.wikipedia.org/wiki/Probabilidad_a_posteriori) para cada conjunto de <a href="https://es.wikipedia.org/wiki/V%C3%A9rtice_(teor%C3%ADa_de_grafos)">nodos</a>. Esta tarea se llama *actualización de creencia* o [inferencia probabilística](https://es.wikipedia.org/wiki/Inferencia). En el caso de las [redes bayesianas](https://es.wikipedia.org/wiki/Red_bayesiana), el proceso de [inferencia](https://es.wikipedia.org/wiki/Inferencia) es muy flexible, nueva evidencia puede ser introducida en cualquiera de los <a href="https://es.wikipedia.org/wiki/V%C3%A9rtice_(teor%C3%ADa_de_grafos)">nodos</a> mientras que las *creencias* son actualizadas en cualquiera de los otros <a href="https://es.wikipedia.org/wiki/V%C3%A9rtice_(teor%C3%ADa_de_grafos)">nodos</a>. En la práctica, la velocidad del proceso de inferencia va a depender de la estructura y complejidad de la red.
# ## Clasificador Bayes ingenuo
#
# Uno de los clasificadores más utilizados en [Machine Learning](http://relopezbriega.github.io/category/machine-learning.html) por su simplicidad y rapidez, es el [Clasificador Bayes ingenuo](https://es.wikipedia.org/wiki/Clasificador_bayesiano_ingenuo). El cual es una técnica de clasificación supervisada basada en el [teorema de Bayes](https://es.wikipedia.org/wiki/Teorema_de_Bayes) que asume que existe una <a href="https://es.wikipedia.org/wiki/Independencia_(probabilidad)">independencia</a> entre los *atributos*. En términos simples, un [Clasificador Bayes ingenuo](https://es.wikipedia.org/wiki/Clasificador_bayesiano_ingenuo) asume que la presencia de una característica particular en una clase no está relacionada con la presencia de cualquier otra característica. Por ejemplo, una fruta puede considerarse como una manzana si es roja, redonda y de aproximadamente 9 cm de diámetro. Incluso si estas características dependen unas de otras o de la existencia de otras características, todas estas propiedades contribuyen independientemente a la probabilidad de que esta fruta sea una manzana. Se lo llama *ingenuo* ya que asumir <a href="https://es.wikipedia.org/wiki/Independencia_(probabilidad)">independencia</a> absoluta entre todos los atributos, no es algo que se suela dar en la realidad.
# El modelo [Bayes ingenuo](https://es.wikipedia.org/wiki/Clasificador_bayesiano_ingenuo) es fácil de construir y particularmente útil para conjuntos de datos muy grandes. A pesar de su simplicidad y de su *irealista* postulado de <a href="https://es.wikipedia.org/wiki/Independencia_(probabilidad)">independencia</a>, este clasificador se ha mostrado muy efectivo y se suele utilizar como el estándar para evaluar el rendimiento de otros modelos de [Machine Learning](http://relopezbriega.github.io/category/machine-learning.html).
#
# El [Clasificador Bayes ingenuo](https://es.wikipedia.org/wiki/Clasificador_bayesiano_ingenuo) se utiliza en múltiples escenarios de la vida real, tales como:
#
# * **Clasificación de texto:** Es uno de los algoritmos conocidos más exitosos cuando se trata de la clasificación de documentos de texto, es decir, si un documento de texto pertenece a una o más categorías (clases).
# * **Detección de spam:** Es un ejemplo de clasificación de texto. Se ha convertido en un mecanismo popular para distinguir el correo electrónico spam del correo electrónico legítimo.
# * **Análisis de sentimientos:** Puede ser utilizado para analizar el tono de tweets, comentarios y revisiones, ya sean negativos, positivos o neutrales.
# * **Sistema de Recomendaciones:** El algoritmo [Bayes ingenuo](https://es.wikipedia.org/wiki/Clasificador_bayesiano_ingenuo) en combinación con el filtrado colaborativo se utiliza para construir sistemas de recomendación híbridos que ayudan a predecir si un usuario desea un recurso determinado o no.
# En este sencillo ejemplo, podemos ver como el [Clasificador Bayes ingenuo](https://es.wikipedia.org/wiki/Clasificador_bayesiano_ingenuo) ha clasificado correctamente la mayoría de los casos del dataset [iris](https://es.wikipedia.org/wiki/Iris_flor_conjunto_de_datos), obteniendo un efectividad del 93 %.
#
# Debido a que los [clasificadores bayesianos ingenuos](https://es.wikipedia.org/wiki/Clasificador_bayesiano_ingenuo) hacen suposiciones tan estrictas acerca de los datos, generalmente no funcionarán tan bien con modelos más complicados. Dicho esto, tienen varias ventajas:
#
# * Son extremadamente rápidos tanto para entrenamiento como para predicción
# * Proporcionan una predicción probabilística directa
# * A menudo son muy fácilmente interpretables
# * Tienen muy pocos parámetros que necesiten optimizarse.
#
# Estas ventajas significan que un [clasificador bayesiano ingenuo](https://es.wikipedia.org/wiki/Clasificador_bayesiano_ingenuo) es a menudo una buena opción como un modelo de clasificación inicial. Si obtenemos resultados satisfactorios, entonces tenemos un clasificador muy rápido, y muy fácil de interpretar. Si no funciona bien, entonces podemos comenzar a explorar modelos más sofisticados.
#
# Aquí concluye esta introducción a la [inferencia bayesiana](https://es.wikipedia.org/wiki/Inferencia_bayesiana); como vemos es una teoría sumamente fascinante con serias implicancias filosóficas. La [teoría Bayesiana](https://es.wikipedia.org/wiki/Teorema_de_Bayes) es mucho más que un simple teorema de probabilidad, es una lógica para razonar sobre el amplio espectro de la vida que se encuentra en las áreas grises entre la *verdad absoluta* y la *incertidumbre total*. A menudo tenemos información sobre sólo una pequeña parte de lo que nos preguntamos.
# Sin embargo, todos queremos predecir algo basado en nuestras experiencias pasadas; y adaptamos nuestras creencias a medida que adquirimos nueva información. La [inferencia bayesiana](https://es.wikipedia.org/wiki/Inferencia_bayesiana) nos proporciona una forma de pensar racionalmente sobre el mundo que nos rodea.
# # Precisión, recall y p-valor
from IPython.display import SVG
SVG(filename='data/precreccal.svg')
# El p-value es la frecuencia con la que nuestro modelo podrá reproducir un resultado más extremo que el obervado de manera aleatoria. Para un científico de datos el p-valor es una métrica que te mide como de interesante puede ser tu modelo.
# * Un valor pequeño del p valor (≤ 0.05), invalida la hipótesis nula.
# * Un valor alto del p valor (> 0.05) significa que la hipótesis alternativa es vaga.
# H-^0 = hipotesis nula H-^1 = Hipotesis alternativa (siempre se mide la nula)
# # OPCIONAL
# ## Programación probabilística y PyMC3
#
# A pesar de que las [redes bayesianas](https://es.wikipedia.org/wiki/Red_bayesiana) y demás modelos de [inferencia bayesiana](https://es.wikipedia.org/wiki/Inferencia_bayesiana) son conceptualmente simples; a menudo los cálculos de sus probabilidades conducen a expresiones que no se pueden resolver en forma analítica. Durante muchos años, este fue un gran problema y fue probablemente una de las principales razones que obstaculizaron la adopción de los métodos bayesianos. La llegada de las computadoras y el desarrollo de métodos numéricos que se pueden aplicar para calcular la distribución *[a posteriori](https://es.wikipedia.org/wiki/Probabilidad_a_posteriori)* de casi cualquier modelo, junto con el avance en las técnicas de muestreo de los [métodos de Monte-Carlo](http://relopezbriega.github.io/blog/2017/01/10/introduccion-a-los-metodos-de-monte-carlo-con-python/); han transformado completamente la práctica del [análisis de datos](http://relopezbriega.github.io/category/analisis-de-datos.html) Bayesiano.
#
# La posibilidad de automatizar la [inferencia probabilística](https://es.wikipedia.org/wiki/Inferencia) ha conducido al desarrollo de la [Programación probabilística](https://en.wikipedia.org/wiki/Probabilistic_programming_language), la cuál utiliza las ventajas de los lenguajes de programación modernos y nos permite realizar una clara separación entre la creación del modelo y el proceso de [inferencia](https://es.wikipedia.org/wiki/Inferencia). En [Programación probabilística](https://en.wikipedia.org/wiki/Probabilistic_programming_language), especificamos un modelo probabilístico completo escribiendo unos cuantos líneas de código y luego la inferencia se realiza en forma automática.
#
# ### PyMC3
#
# [PyMC3](https://pymc-devs.github.io/pymc3/index.html) es un paquete para [Programación probabilística](https://en.wikipedia.org/wiki/Probabilistic_programming_language) que utiliza el lenguaje de programación [Python](http://python.org/). [PyMC3](https://pymc-devs.github.io/pymc3/index.html) es lo suficientemente maduro para resolver muchos de los principales problemas estadísticos. Permite crear modelos probabilísticos usando una sintaxis intuitiva y fácil de leer que es muy similar a la sintaxis usada para describir modelos probabilísticos.
#
# Veamos algunos ejemplos:
#
# #### El problema de la moneda
# Los problemas de monedas son clásicos cuando hablamos de [probabilidad y estadística](http://relopezbriega.github.io/category/pobabilidad-y-estadistica.html), nos permiten ejemplificar conceptos abstractos de forma simple. Asimismo, pueden ser muchas veces conceptualmente similares a situaciones *reales*, de hecho cualquier problema en donde obtengamos resultados binarios, 0/1, enfermo/sano, spam/no-spam, puede ser pensado como si estuviéramos hablando de monedas.
# En este caso, la idea es utilizar un modelo [bayesiano](https://es.wikipedia.org/wiki/Inferencia_bayesiana) para inferir si la moneda se encuentra sesgada o no.
#
# Para este ejemplo, vamos a utilizar una [distribución binomial](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_binomial) como *likelihood* y una [distribución beta](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_beta) como *[a priori](https://es.wikipedia.org/wiki/Probabilidad_a_priori)*. Veamos como lo podemos modelar con [PyMC3](https://pymc-devs.github.io/pymc3/index.html).
# + jupyter={"outputs_hidden": true}
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import scipy.stats as stats
import seaborn as sns
import pymc3 as pm
import theano.tensor as tt
from sklearn import datasets
from sklearn.naive_bayes import GaussianNB
from sklearn.metrics import confusion_matrix
from sklearn.model_selection import train_test_split
np.random.seed(1984) #replicar random
# %matplotlib inline
# -
# El problema de la moneda
# de 100 lanzamientos 80 caras
n = 100
caras = 80
# Creación del modelo
niter = 2000
with pm.Model() as modelo_moneda:
# a priori
p = pm.Beta('p', alpha=2, beta=2)
# likelihood
y = pm.Binomial('y', n=n, p=p, observed=caras)
# Realizando el muestreo para la inferencia
with modelo_moneda:
trace = pm.sample(niter, njobs=4)
# Analizando los resultados
pm.traceplot(trace, varnames=['p'], lines={'p':.8})
pass
# Información resumen.
#Vemos que hay un 95% de probabilidades de que el valor de sesgo este entre
# .706 y .864
pm.summary(trace)
# Como vemos el modelo nos indica que la moneda parece tener un claro sesgo hacia cara.
#
# #### El problema de la hierba mojada
# Supongamos que hay dos eventos los cuales pueden causar que la hierba esté húmeda: que el rociador esté activado o que esté lloviendo. También supongamos que la lluvia tiene un efecto directo sobre el uso del rociador (usualmente cuando llueve el rociador se encuentra apagado). Entonces la situación puede ser modelada con la siguiente [red bayesiana](https://es.wikipedia.org/wiki/Red_bayesiana).
#
# <img alt="Red Bayesiana" title="Modelo Hierba mojada" src="https://upload.wikimedia.org/wikipedia/commons/9/97/Red_Bayesiana_Simple.png" >
#
# +
# Problema de la hierba mojada
# https://es.wikipedia.org/wiki/Red_bayesiana#Ejemplo
niter = 10000 # 10000
tune = 5000 # 5000
modelo = pm.Model()
with modelo:
tv = [1]
lluvia = pm.Bernoulli('lluvia', 0.2, shape=1, testval=tv)
rociador_p = pm.Deterministic('rociador_p',
pm.math.switch(lluvia, 0.01, 0.40))
rociador = pm.Bernoulli('rociador', rociador_p, shape=1, testval=tv)
hierba_mojada_p = pm.Deterministic('hierba_mojada_p',
pm.math.switch(lluvia, pm.math.switch(rociador, 0.99, 0.80),
pm.math.switch(rociador, 0.90, 0.0)))
hierba_mojada = pm.Bernoulli('hierba_mojada', hierba_mojada_p,
observed=np.array([1]), shape=1)
trace = pm.sample(20000,
step=[pm.BinaryGibbsMetropolis([lluvia, rociador])],
tune=tune, random_seed=124)
# pm.traceplot(trace)
dictionary = {
'lluvia': [1 if ii[0] else 0 for ii in trace['lluvia'].tolist() ],
'rociador': [1 if ii[0] else 0 for ii in trace['rociador'].tolist() ],
'rociador_p': [ii[0] for ii in trace['rociador_p'].tolist()],
'hierba_mojada_p': [ii[0] for ii in trace['hierba_mojada_p'].tolist()],
}
df = pd.DataFrame(dictionary)
p_lluvia = df[(df['lluvia'] == 1)].shape[0] / df.shape[0]
print("\nProbabilidad de que la hierba este mojada por la lluvia: {0}"
.format(p_lluvia))
p_rociador = df[(df['rociador'] == 1)].shape[0] / df.shape[0]
print("Probabilidad de que la hierba este mojada por el rociador: {0}"
.format(p_rociador))
# -
# De acuerdo a los resultados de la [red bayesiana](https://es.wikipedia.org/wiki/Red_bayesiana), si vemos que la hierba esta mojada, la probabilidad de que este lloviendo es alrededor del 38%.
| Precurso/03_Matematicas_estadistica_Git/Introduccion-estadistica.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# ## Source code for _Investigating particle size-flux relationships and the biological pump across a range of plankton ecosystem states from coastal to oligotrophic_
#
# Manuscript by: __<NAME>, <NAME>, <NAME>, <NAME>, <NAME>, and <NAME>__
#
# Code by: __<NAME> and <NAME> (FSU)__
#
# Manuscript submitted to Frontiers in Marine Science (March 2019)
#
#
# ### _Abstract_
#
# Sinking particles transport organic carbon produced in the surface ocean to the ocean interior, leading to net storage of atmospheric CO2 in the deep ocean. The rapid growth of in situ imaging technology has the potential to revolutionize our understanding of particle flux attenuation in the ocean; however, estimating particle flux from particle size and abundance (measured directly by in situ cameras) is challenging. Sinking rates are dependent on several factors, including particle excess density and porosity, which vary based on particle origin and type. Additionally, particle characteristics are transformed while sinking. We compare optically-measured particle size spectra profiles (Underwater Vision Profiler 5, UVP) with contemporaneous measurements of particle flux made using sediment traps and $^{234}Th:^{238}U$ disequilibrium on six process cruises from the California Current Ecosystem (CCE) LTER Program. These measurements allow us to assess the efficacy of using size-flux relationships to estimate fluxes from optical particle size measurements. We find that previously published parameterizations that estimate carbon flux from UVP profiles are a poor fit to direct flux measurements in the CCE. This discrepancy is found to result primarily from the important role of fecal pellets in particle flux. These pellets are primarily in a size range (i.e., 100 – 400 µm) that is not well-resolved as images by the UVP due to the resolution of the sensor. We develop a new, CCE-optimized algorithm for estimating carbon flux from UVP data in the southern California Current (Flux = $\sum_{i=1}^{x}{n_i A d_i^B ∆d_i }$), with A = 13.45, B = 1.35, d = particle diameter (mm) and Flux in units of $mg C m^{-2} d^{-1}$. We caution, however, that increased accuracy in flux estimates derived from optical instruments will require devices with greater resolution, the ability to differentiate fecal pellets from low porosity marine snow aggregates, and improved sampling of rapidly sinking fecal pellets. We also find that the particle size-flux relationships may be different within the euphotic zone than in the shallow twilight zone and hypothesize that the changing nature of sinking particles with depth must be considered when investigating the remineralization length scale of sinking particles in the ocean.
#
# [](https://zenodo.org/badge/latestdoi/178078252)
#
#
# ---
#
# ### Note on downloading the Raw Data
#
# An account with Ecotaxa will be necessary for the following steps.
#
# 1. From the home page, select the drop down window next to Action at the top right. Select the Particle Module.
# 2. Enter the relevant filters. For this study all CCELTER Particle Projects were selected.
# 3. Select Export Selection.
# 4. Download the RAW data format. Only this format will work with the provided code.
# 5. From the downloaded files, only the PAR files and the metadata summary file were used for analysis.
#
# ---
#
# ### Start by loading the source files
source('source.uvp.r') ## Functions to make uvp data objects
source('source.uvp.add.r') ## functions to add values to uvp data objects
source('source.r') ## General purpose functions (optional)
# Next we will define directories and load the UVP metadata files
# +
#### Load the Data
input.dir = '../UVP Data/All PAR files/'
input.dir.meta = '../UVP Data/All Metadata files/'
raw.files = list.files(input.dir)
meta.files = list.files(input.dir.meta)
## Reading in only first metadata file.
meta = as.data.frame(fread(paste0(input.dir.meta, meta.files[1])))
## Combining with the rest.
for (i in 2:length(meta.files)) {
meta = rbind(meta, as.data.frame(fread(paste0(input.dir.meta, meta.files[i]))))
}
# -
# _Optional:_ Here we load up the CTD and Sed Trap datasets and save them as rdata files for easy loading later.
# +
#### CTD and Sed Trap File section
#ctd.file = '../Other Data/CTD Downcast Data.xlsx'
#ctd = read.xlsx(ctd.file)
#ctd$Time = as.POSIXct(ctd$Datetime.GMT, tz = 'UTC') ## Get timestamps
#save(ctd, file = '../Other Data/CTD.rdata')
#sedtrap = read.xlsx('../Other Data/Sediment Trap.xlsx')
#sedtrap$Deployment.Time = as.POSIXct(sedtrap$Deployment.Time)
#sedtrap$Recovery.Datetime = as.POSIXct(sedtrap$Recovery.Datetime)
#save(sedtrap, file = '../Other Data/SedTrap.rdata')
# -
## Load saved datasets
load('../Other Data/CTD.rdata')
load('../Other Data/SedTrap.rdata')
# ### Build a UVP data object
# First we build one to make sure the system is working, and then we'll loop it for all the uvp cast files.
##Initialize empty data structure, set Files
MasterList = init.raw(Dir = input.dir, File = raw.files[269]) # Picked a random file.
MasterList$params$dz = 5
MasterList$params$A = 13.7
MasterList$params$b = 1.55
## Load our data: set data.raw, fill in meta
MasterList = load.raw(MasterList, meta, TRUE)
MasterList = filter.by.size(MasterList, d.min = 0.102, d.max = 1.5) # Filtering to 2nd smallest size bin
# Setup bins based on a logrithmic scale.
# +
k = 3 ## Our current bins (increase for higher resolution, Guidi = 3)
bins = c(2^(c((5*k):(11*k))/k))
bins = floor(bins)
delta.bins = diff(bins)
## Save bin values
MasterList$params$bins = bins
MasterList$params$delta.bins = delta.bins
MasterList$params$n = length(bins)
# +
## Make the bins
MasterList = make.bins(MasterList)
## Calculate the flux (5m bins)
MasterList = calc.flux(MasterList)
## Add export values (sed trap data)
MasterList = add.export(MasterList, thorium = thorium, sedtrap = sedtrap)
##Add ctd metadata
MasterList = add.ctd(MasterList, ctd, verbose = TRUE)
# -
# ### Done with test
#
# ---
#
# ### Full scale
# If that all works, then we can make a function to loop over all the files and make a list of each uvp cast object:
## See source.uvp.r for code on run.all.data(). This will need to be edited and provided as a template only.
cruises = run.all.data(raw.files, A = 13.45, b = 1.35, dz = 5, ctd = ctd, verbose = FALSE, k = 3)
# ### Save result
save(cruises, file = '../rstates/cruises.rdata')
| Main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"is_executing": false}
import matplotlib.pyplot as plt
from sklearn.manifold import TSNE
from util import getKaggleMNIST
# + pycharm={"metadata": false, "name": "#%%\n", "is_executing": false}
Xtrain, Ytrain, _, _ = getKaggleMNIST()
# + pycharm={"metadata": false, "name": "#%%\n", "is_executing": false}
sample_size = 1000
X = Xtrain[:sample_size]
Y = Ytrain[:sample_size]
# + pycharm={"metadata": false, "name": "#%%\n", "is_executing": false}
tsne = TSNE()
Z = tsne.fit_transform(X)
plt.scatter(Z[:, 0], Z[:, 1], s=100, c=Y, alpha=0.5)
# + pycharm={"metadata": false, "name": "#%%\n"}
| unsupervised2/tnse_mnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import io
import cv2
import matplotlib.pyplot as plt
import numpy as np
import requests
import torch
import torch.nn as nn
import torch.nn.functional as F
from PIL import Image
from torchvision import models, transforms
from exercise05.model import ConvNet
device = torch.device("cpu")
# -
np.random.seed(42)
torch.manual_seed(42)
# +
# load everything that we need
# here we load alexnet, an already trained neural network
# you can play with other models in models.*, but for some of the others it is harder
# to get the activations in the middle of the network
alexnet = models.alexnet(pretrained=True)
# define a 'layer' to normalize an image such that it is usable by the network
class Normalize(nn.Module):
def __init__(self):
super().__init__()
self.mean = torch.Tensor([0.485, 0.456, 0.406]).float().unsqueeze(0).unsqueeze(-1).unsqueeze(-1)
self.std = torch.Tensor([0.229, 0.224, 0.225]).float().unsqueeze(0).unsqueeze(-1).unsqueeze(-1)
def forward(self, x):
b, c, h, w = x.shape
mean = self.mean.expand(x.shape)
std = self.std.expand(x.shape)
return (x - self.mean)/self.std
# define a layer that flattens whatever it gets passed into a vector
class Flatten(nn.Module):
def forward(self, input):
return input.view(input.size(0), -1)
# function that takes our pre-processed image and computes a numpy matrix that we can plot as an image
def img2numpy(x):
x = x.clone().squeeze()
x = x.detach().numpy()
x = np.transpose(x, (1, 2, 0))
return x
# preprocess an image for the network
preprocess = transforms.Compose([
transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor()
])
# download an image from flickr
response = requests.get("http://c1.staticflickr.com/5/4070/5148597478_0c34ec0b7e_n.jpg")
image = Image.open(io.BytesIO(response.content))
image = preprocess(image).unsqueeze(0)
plt.imshow(img2numpy(image))
# -
# download a mapping of the imagenet class ids to text
# https://gist.github.com/yrevar/942d3a0ac09ec9e5eb3a
imagenet_classes_request = requests.get("https://gist.githubusercontent.com/yrevar/942d3a0ac09ec9e5eb3a/raw/c2c91c8e767d04621020c30ed31192724b863041/imagenet1000_clsid_to_human.txt")
# turn the downloaded id-to-text mapping into a dict
# before running this really inspect the output of the above comment
# you are about to run something downloaded from the internet
# run at your own risk
imagenet_classes = eval(imagenet_classes_request.content)
# try out the neural network
# we want to run our normalize layer first and then alexnet
model = nn.Sequential(Normalize(), alexnet)
logits = model(image).detach().numpy().ravel().argsort()[::-1][:5]
print('Top 5 classes for image:')
print(list(map(lambda x: imagenet_classes[x], logits.tolist())))
# We now take a closer look at the layers in the alexnet model.
# alexnet has two parts 'features' and 'classifier' where features is the convolutional part of the neural network.
alexnet.features
alexnet.classifier
# we see that we can use the individual parts of alexnet
# but we need to add a Flatten() layer between the two parts
model = nn.Sequential(Normalize(), alexnet.features, Flatten(), alexnet.classifier)
logits = model(image).detach().numpy().ravel().argsort()[::-1][:5]
print('Top 5 classes for image:')
print(list(map(lambda x: imagenet_classes[x], logits.tolist())))
def gradcam(model, image, layer, target, treshold=0.5):
"""
This method takes:
model - an alexnet
image - an input image
layer - an integer that indexes alexnet.features; this gives the layer that we use for the algorithm
target - the target class for visualization
treshold - how much of the heat map to show in the overlayed image
"""
assert 1 <= layer <= 12 # layer is valid index into alexnet.features
model.eval()
model.zero_grad()
# split model into two parts
model_part_1 = nn.Sequential(Normalize(), model.features[:layer+1])
model_part_2 = nn.Sequential(model.features[layer+1:], Flatten(), model.classifier)
# extract the activation at the requested layer
layer_activations = model_part_1(image).detach()
layer_activations.requires_grad_()
# compute forward pass through the rest of the network
logits = model_part_2(layer_activations.clone())
# compute gradient from target logit
logits[0, target].backward()
# compute weights alpha
G = layer_activations.grad
layer_shape = G.shape
alpha = G.reshape(layer_shape[:2] + (-1,)).mean(2)
# compute activation map
L = alpha.unsqueeze(-1).unsqueeze(-1).repeat((1,1) + layer_shape[2:]) * layer_activations
L = L.sum(1).relu().detach().numpy()[0, ...]
# create transparent heatmap for visualization
L_transparent = np.tile(L[:, :, None], (1, 1, 4))
L_transparent[:, :, 0] /= L_transparent[:, :, 0].max()
L_transparent[:, :, 1] /= L_transparent[:, :, 1].max()
L_transparent[:, :, 2] /= L_transparent[:, :, 2].max()
L_transparent[:, :, 3] = L_transparent[:, :, 3] > treshold * L_transparent[:, :, 3].max()
L_transparent = cv2.resize(L_transparent, dsize=image.shape[2:], interpolation=cv2.INTER_NEAREST)
# show the results
f, axarr = plt.subplots(1,3, figsize=(18, 6))
f.suptitle('Visualization for Class: ' + imagenet_classes[target], fontsize=16)
axarr[0].imshow(img2numpy(image))
axarr[0].set_title('Original Image')
axarr[1].imshow(L, interpolation='nearest')
axarr[1].set_title('Heatmap')
axarr[2].imshow(img2numpy(image))
axarr[2].imshow(L_transparent)
axarr[2].set_title('Overlay')
# ## Visualizing different classes
s = 163 #'bloodhound, sleuthhound'
gradcam(alexnet, image, 9, s, treshold=0.2)
s = 282 #tiger cat
gradcam(alexnet, image, 9, s, treshold=0.2)
s = 243 #'bull mastiff'
gradcam(alexnet, image, 9, s, treshold=0.2)
# ## Visualizing one class for various different layers
s = 163 #'bloodhound, sleuthhound'
gradcam(alexnet, image, 2, s, treshold=0.2)
s = 163 #'bloodhound, sleuthhound'
gradcam(alexnet, image, 3, s, treshold=0.2)
s = 163 #'bloodhound, sleuthhound'
gradcam(alexnet, image, 4, s, treshold=0.2)
s = 163 #'bloodhound, sleuthhound'
gradcam(alexnet, image, 5, s, treshold=0.2)
s = 163 #'bloodhound, sleuthhound'
gradcam(alexnet, image, 6, s, treshold=0.2)
s = 163 #'bloodhound, sleuthhound'
gradcam(alexnet, image, 7, s, treshold=0.2)
s = 163 #'bloodhound, sleuthhound'
gradcam(alexnet, image, 8, s, treshold=0.2)
s = 163 #'bloodhound, sleuthhound'
gradcam(alexnet, image, 9, s, treshold=0.2)
s = 163 #'bloodhound, sleuthhound'
gradcam(alexnet, image, 10, s, treshold=0.2)
s = 163 #'bloodhound, sleuthhound'
gradcam(alexnet, image, 11, s, treshold=0.2)
s = 163 #'bloodhound, sleuthhound'
gradcam(alexnet, image, 12, s, treshold=0.2)
| exercise08/gradcam.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] id="PDDqFd8hon3v" colab_type="text"
# <h1> Create and benchmark train, validation, and test datasets </h1>
#
# In this notebook, you will prepare the training, validation (sometimes called evaluation), and test datasets using the NYC taxi fare data. After that, you will benchmark against these datasets, in other words use your benchmark model to calculate the metric values for the training, validation, and test examples. In parallel, you will learn how to include various bash shell commands (e.g. ls, head, and others) in your Colab notebooks.
#
#
# ---
# Before you start, **make sure that you are logged in with your student account**. Otherwise you may incur Google Cloud charges for using this notebook.
#
# ---
#
# + id="Wq75B91eon3y" colab_type="code" cellView="form" colab={}
import numpy as np
import pandas as pd
import seaborn as sns
import shutil
from google.cloud import bigquery
#@markdown Copy-paste your GCP Project ID in the following field:
PROJECT = "" #@param {type: "string"}
#@markdown When running this cell you will need to **uncheck "Reset all runtimes before running"** as shown on the following screenshot:
#@markdown 
#@markdown Next, use Shift-Enter to run this cell and to complete authentication.
try:
from google.colab import auth
auth.authenticate_user()
print("AUTHENTICATED")
except:
print("FAILED to authenticate")
# + [markdown] id="quQPY9r8MzQ6" colab_type="text"
# Next, query for 1 out of 100,000 rows of the entire taxi fare dataset and apply the clean up pre-processing rules you have developed in the earlier lab. Based on the summary from the Pandas describe table, you can confirm that there are roughly 10,500 rows of cleaned-up data in the `tripsqc` variable.
# + id="YdiXbfDuI-LI" colab_type="code" colab={}
EVERY_N = 100000
bq = bigquery.Client(project=PROJECT)
trips = bq.query('''
SELECT
pickup_datetime,
pickup_longitude, pickup_latitude,
dropoff_longitude, dropoff_latitude,
passenger_count,
trip_distance,
tolls_amount,
fare_amount,
total_amount
FROM
`nyc-tlc.yellow.trips`
WHERE
MOD(ABS(FARM_FINGERPRINT(STRING(pickup_datetime))), %d) = 1
#note that that trips with zero distance or
#costing less than $2.50 are excluded
AND trip_distance > 0 AND fare_amount >= 2.5
''' % (EVERY_N)).to_dataframe()
def preprocess(trips_in):
trips = trips_in.copy(deep=True)
trips.fare_amount = trips.fare_amount + trips.tolls_amount
del trips['tolls_amount']
del trips['total_amount']
del trips['trip_distance']
del trips['pickup_datetime']
qc = np.all([\
trips['pickup_longitude'] > -78, \
trips['pickup_longitude'] < -70, \
trips['dropoff_longitude'] > -78, \
trips['dropoff_longitude'] < -70, \
trips['pickup_latitude'] > 37, \
trips['pickup_latitude'] < 45, \
trips['dropoff_latitude'] > 37, \
trips['dropoff_latitude'] < 45, \
trips['passenger_count'] > 0,
], axis=0)
return trips[qc]
tripsqc = preprocess(trips)
tripsqc.describe()
# + [markdown] id="bbsRzcqLon46" colab_type="text"
# <h3> Create ML datasets </h3>
#
# The next cell splits the cleaned up dataset randomly into training, validation and test sets. Since you are working with an in-memory dataset (for now), you will use a 70%-15%-15% split. Later you will learn about the benefits of allocating a larger percentage of the entire dataset for training.
# + id="9WZgUiO0on47" colab_type="code" colab={}
shuffled = tripsqc.sample(frac=1)
trainsize = int(len(shuffled['fare_amount']) * 0.70)
validsize = int(len(shuffled['fare_amount']) * 0.15)
df_train = shuffled.iloc[:trainsize, :]
df_valid = shuffled.iloc[trainsize:(trainsize+validsize), :]
df_test = shuffled.iloc[(trainsize+validsize):, :]
df_train.describe()
# + id="G60adT61on5C" colab_type="code" colab={}
df_valid.describe()
# + id="1i2mo1CAon5H" colab_type="code" colab={}
df_test.describe()
# + [markdown] id="ng1gbvDuon5K" colab_type="text"
# Let's write out the three dataframes to appropriately named csv files. The files will be useful for local training while you are developing your machine learning models. In future labs, you will scale out to a larger dataset using other serverless capabilities like Cloud Machine Learning Engine (Cloud MLE) and Dataflow.
# + id="yuQwBVVpon5L" colab_type="code" colab={}
def to_csv(df, filename):
outdf = df.copy(deep=False)
outdf.loc[:, 'key'] = np.arange(0, len(outdf)) # rownumber as key
# reorder columns so that target is first column
cols = outdf.columns.tolist()
cols.remove('fare_amount')
cols.insert(0, 'fare_amount')
print (cols) # new order of columns
outdf = outdf[cols]
outdf.to_csv(filename, header=False, index_label=False, index=False)
to_csv(df_train, 'taxi-train.csv')
to_csv(df_valid, 'taxi-valid.csv')
to_csv(df_test, 'taxi-test.csv')
# + [markdown] id="MOPzJGT7OtWu" colab_type="text"
# There are 2 ways to execute shell commands in the OS environment hosting this notebook:
#
# 1. You can prefix your `bash` command with an exclaimation mark as shown in the next code cell.
#
# 2. You can use the `%%bash` "magic" as the first line of a code cell. This approach is better suited for multi-line shell scripts.
#
# If you are interested in details about Jupyter "magics", you can learn more [here](https://nbviewer.jupyter.org/github/ipython/ipython/blob/1.x/examples/notebooks/Cell%20Magics.ipynb)
# + [markdown] id="RGZsBGRcon5T" colab_type="text"
# <h3> Verify that datasets exist </h3>
# + id="57XAd4O8on5U" colab_type="code" colab={}
# !ls -l *.csv
# + [markdown] id="f1AvPQIDon5Y" colab_type="text"
# There are 3 .csv files corresponding to train, valid, test datasets. The ratio of file sizes reflect the percentages in the split of the data.
# + id="Tcqtnc0Hon5Y" colab_type="code" colab={} language="bash"
# head -10 taxi-train.csv
# + [markdown] id="St5NHo1lon5b" colab_type="text"
# Looks good! Now the datasets are prepared so you will be ready to train machine learning models, evaluate, and test them.
# + [markdown] id="DBbYTPZAon5c" colab_type="text"
# <h3> Benchmark </h3>
#
# Before committing to a complex machine learning model, it is a good idea to come up with a very simple, heuristic model and use that as a benchmark.
#
# My model is going to simply divide the mean fare_amount by the mean trip_distance to come up with an average rate per kilometer and use that to predict. Let's compute the RMSE of such a model.
# + id="o4TuDR8Jon5c" colab_type="code" colab={}
def distance_between(lat1, lon1, lat2, lon2):
# haversine formula to compute distance "as the crow flies". Taxis can't fly of course.
dist = np.degrees(np.arccos(np.minimum(1,np.sin(np.radians(lat1)) * np.sin(np.radians(lat2)) + np.cos(np.radians(lat1)) * np.cos(np.radians(lat2)) * np.cos(np.radians(lon2 - lon1))))) * 60 * 1.515 * 1.609344
return dist
def estimate_distance(df):
return distance_between(df['pickuplat'], df['pickuplon'], df['dropofflat'], df['dropofflon'])
def compute_rmse(actual, predicted):
return np.sqrt(np.mean((actual-predicted)**2))
def print_rmse(df, rate, name):
print ("{1} RMSE = {0}".format(compute_rmse(df['fare_amount'], rate*estimate_distance(df)), name))
FEATURES = ['pickuplon','pickuplat','dropofflon','dropofflat','passengers']
TARGET = 'fare_amount'
columns = list([TARGET])
columns.extend(FEATURES) # in CSV, target is the first column, after the features
columns.append('key')
df_train = pd.read_csv('taxi-train.csv', header=None, names=columns)
df_valid = pd.read_csv('taxi-valid.csv', header=None, names=columns)
df_test = pd.read_csv('taxi-test.csv', header=None, names=columns)
rate = df_train['fare_amount'].mean() / estimate_distance(df_train).mean()
print ("Rate = ${0}/km".format(rate))
print_rmse(df_train, rate, 'Train')
print_rmse(df_valid, rate, 'Valid')
print_rmse(df_test, rate, 'Test')
# + [markdown] id="Z5TGIWkQon5i" colab_type="text"
# <h2>Benchmark on same dataset</h2>
#
# The RMSE depends on the dataset and for meaningful and reproducible comparisons it is critical to measure on the same dataset each time. The following query will continue to be reused in the later labs:
# + id="O9xVQv49on5j" colab_type="code" colab={}
def create_query(phase, EVERY_N):
"""
phase: 1=train 2=valid
"""
base_query = """
SELECT
(tolls_amount + fare_amount) AS fare_amount,
CONCAT( STRING(pickup_datetime),
CAST(pickup_longitude AS STRING),
CAST(pickup_latitude AS STRING),
CAST(dropoff_latitude AS STRING),
CAST(dropoff_longitude AS STRING)) AS key,
EXTRACT(DAYOFWEEK FROM pickup_datetime)*1.0 AS dayofweek,
EXTRACT(HOUR FROM pickup_datetime)*1.0 AS hourofday,
pickup_longitude AS pickuplon,
pickup_latitude AS pickuplat,
dropoff_longitude AS dropofflon,
dropoff_latitude AS dropofflat,
passenger_count*1.0 AS passengers
FROM
`nyc-tlc.yellow.trips`
WHERE
{}
AND trip_distance > 0
AND fare_amount >= 2.5
AND pickup_longitude > -78
AND pickup_longitude < -70
AND dropoff_longitude > -78
AND dropoff_longitude < -70
AND pickup_latitude > 37
AND pickup_latitude < 45
AND dropoff_latitude > 37
AND dropoff_latitude < 45
AND passenger_count > 0
"""
if EVERY_N == None:
if phase < 2:
# training
selector = "MOD(ABS(FARM_FINGERPRINT(STRING(pickup_datetime))), 4) < 2"
else:
selector = "MOD(ABS(FARM_FINGERPRINT(STRING(pickup_datetime))), 4) = 2"
else:
selector = "MOD(ABS(FARM_FINGERPRINT(STRING(pickup_datetime))), %d) = %d" % (EVERY_N, phase)
query = base_query.format(selector)
return query
sql = create_query(2, 100000)
df_valid = bq.query(sql).to_dataframe()
print_rmse(df_valid, 2.56, 'Final Validation Set')
# + [markdown] id="MT9dwy_ton5m" colab_type="text"
# The simple distance-based rule gives us a RMSE of <b>$7.42</b>. We have to beat this, of course, but you will find that simple rules of thumb like this can be surprisingly difficult to beat.
#
# Let's be ambitious, though, and make our goal to build ML models that have a RMSE of less than $6 on the test set.
# + [markdown] id="HVuX8oeMon5m" colab_type="text"
# Copyright 2019 CounterFactual.AI LLC. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
| mle1/create_datasets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] origin_pos=0
# # 权重衰减
# :label:`sec_weight_decay`
#
# 前一节我们描述了过拟合的问题,本节我们将介绍一些正则化模型的技术。
# 我们总是可以通过去收集更多的训练数据来缓解过拟合。
# 但这可能成本很高,耗时颇多,或者完全超出我们的控制,因而在短期内不可能做到。
# 假设我们已经拥有尽可能多的高质量数据,我们便可以将重点放在正则化技术上。
#
# 回想一下,在多项式回归的例子( :numref:`sec_model_selection`)中,
# 我们可以通过调整拟合多项式的阶数来限制模型的容量。
# 实际上,限制特征的数量是缓解过拟合的一种常用技术。
# 然而,简单地丢弃特征对于这项工作来说可能过于生硬。
# 我们继续思考多项式回归的例子,考虑高维输入可能发生的情况。
# 多项式对多变量数据的自然扩展称为*单项式*(monomials),
# 也可以说是变量幂的乘积。
# 单项式的阶数是幂的和。
# 例如,$x_1^2 x_2$和$x_3 x_5^2$都是3次单项式。
#
# 注意,随着阶数$d$的增长,带有阶数$d$的项数迅速增加。
# 给定$k$个变量,阶数$d$(即$k$多选$d$)的个数为
# ${k - 1 + d} \choose {k - 1}$。
# 即使是阶数上的微小变化,比如从$2$到$3$,
# 也会显著增加我们模型的复杂性。
# 因此,我们经常需要一个更细粒度的工具来调整函数的复杂性。
#
# ## 范数与权重衰减
#
# 在 :numref:`subsec_lin-algebra-norms`中,
# 我们已经描述了$L_2$范数和$L_1$范数,
# 它们是更为一般的$L_p$范数的特殊情况。
# (~~权重衰减是最广泛使用的正则化的技术之一~~)
# 在训练参数化机器学习模型时,
# *权重衰减*(weight decay)是最广泛使用的正则化的技术之一,
# 它通常也被称为$L_2$*正则化*。
# 这项技术通过函数与零的距离来衡量函数的复杂度,
# 因为在所有函数$f$中,函数$f = 0$(所有输入都得到值$0$)
# 在某种意义上是最简单的。
# 但是我们应该如何精确地测量一个函数和零之间的距离呢?
# 没有一个正确的答案。
# 事实上,函数分析和巴拿赫空间理论的研究,都在致力于回答这个问题。
#
# 一种简单的方法是通过线性函数
# $f(\mathbf{x}) = \mathbf{w}^\top \mathbf{x}$
# 中的权重向量的某个范数来度量其复杂性,
# 例如$\| \mathbf{w} \|^2$。
# 要保证权重向量比较小,
# 最常用方法是将其范数作为惩罚项加到最小化损失的问题中。
# 将原来的训练目标*最小化训练标签上的预测损失*,
# 调整为*最小化预测损失和惩罚项之和*。
# 现在,如果我们的权重向量增长的太大,
# 我们的学习算法可能会更集中于最小化权重范数$\| \mathbf{w} \|^2$。
# 这正是我们想要的。
# 让我们回顾一下 :numref:`sec_linear_regression`中的线性回归例子。
# 我们的损失由下式给出:
#
# $$L(\mathbf{w}, b) = \frac{1}{n}\sum_{i=1}^n \frac{1}{2}\left(\mathbf{w}^\top \mathbf{x}^{(i)} + b - y^{(i)}\right)^2.$$
#
# 回想一下,$\mathbf{x}^{(i)}$是样本$i$的特征,
# $y^{(i)}$是样本$i$的标签,
# $(\mathbf{w}, b)$是权重和偏置参数。
# 为了惩罚权重向量的大小,
# 我们必须以某种方式在损失函数中添加$\| \mathbf{w} \|^2$,
# 但是模型应该如何平衡这个新的额外惩罚的损失?
# 实际上,我们通过*正则化常数*$\lambda$来描述这种权衡,
# 这是一个非负超参数,我们使用验证数据拟合:
#
# $$L(\mathbf{w}, b) + \frac{\lambda}{2} \|\mathbf{w}\|^2,$$
#
# 对于$\lambda = 0$,我们恢复了原来的损失函数。
# 对于$\lambda > 0$,我们限制$\| \mathbf{w} \|$的大小。
# 这里我们仍然除以$2$:当我们取一个二次函数的导数时,
# $2$和$1/2$会抵消,以确保更新表达式看起来既漂亮又简单。
# 你可能会想知道为什么我们使用平方范数而不是标准范数(即欧几里得距离)?
# 我们这样做是为了便于计算。
# 通过平方$L_2$范数,我们去掉平方根,留下权重向量每个分量的平方和。
# 这使得惩罚的导数很容易计算:导数的和等于和的导数。
#
# 此外,你可能会问为什么我们首先使用$L_2$范数,而不是$L_1$范数。
# 事实上,这个选择在整个统计领域中都是有效的和受欢迎的。
# $L_2$正则化线性模型构成经典的*岭回归*(ridge regression)算法,
# $L_1$正则化线性回归是统计学中类似的基本模型,
# 通常被称为*套索回归*(lasso regression)。
# 使用$L_2$范数的一个原因是它对权重向量的大分量施加了巨大的惩罚。
# 这使得我们的学习算法偏向于在大量特征上均匀分布权重的模型。
# 在实践中,这可能使它们对单个变量中的观测误差更为稳定。
# 相比之下,$L_1$惩罚会导致模型将权重集中在一小部分特征上,
# 而将其他权重清除为零。
# 这称为*特征选择*(feature selection),这可能是其他场景下需要的。
#
# 使用与 :eqref:`eq_linreg_batch_update`中的相同符号,
# $L_2$正则化回归的小批量随机梯度下降更新如下式:
#
# $$
# \begin{aligned}
# \mathbf{w} & \leftarrow \left(1- \eta\lambda \right) \mathbf{w} - \frac{\eta}{|\mathcal{B}|} \sum_{i \in \mathcal{B}} \mathbf{x}^{(i)} \left(\mathbf{w}^\top \mathbf{x}^{(i)} + b - y^{(i)}\right).
# \end{aligned}
# $$
#
# 根据之前章节所讲的,我们根据估计值与观测值之间的差异来更新$\mathbf{w}$。
# 然而,我们同时也在试图将$\mathbf{w}$的大小缩小到零。
# 这就是为什么这种方法有时被称为*权重衰减*。
# 我们仅考虑惩罚项,优化算法在训练的每一步*衰减*权重。
# 与特征选择相比,权重衰减为我们提供了一种连续的机制来调整函数的复杂度。
# 较小的$\lambda$值对应较少约束的$\mathbf{w}$,
# 而较大的$\lambda$值对$\mathbf{w}$的约束更大。
#
# 是否对相应的偏置$b^2$进行惩罚在不同的实践中会有所不同,
# 在神经网络的不同层中也会有所不同。
# 通常,网络输出层的偏置项不会被正则化。
#
# ## 高维线性回归
#
# 我们通过一个简单的例子来演示权重衰减。
#
# + origin_pos=3 tab=["tensorflow"]
# %matplotlib inline
import tensorflow as tf
from d2l import tensorflow as d2l
# + [markdown] origin_pos=4
# 首先,我们[**像以前一样生成一些数据**],生成公式如下:
#
# (**$$y = 0.05 + \sum_{i = 1}^d 0.01 x_i + \epsilon \text{ where }
# \epsilon \sim \mathcal{N}(0, 0.01^2).$$**)
#
# 我们选择标签是关于输入的线性函数。
# 标签同时被均值为0,标准差为0.01高斯噪声破坏。
# 为了使过拟合的效果更加明显,我们可以将问题的维数增加到$d = 200$,
# 并使用一个只包含20个样本的小训练集。
#
# + origin_pos=5 tab=["tensorflow"]
n_train, n_test, num_inputs, batch_size = 20, 100, 200, 5
true_w, true_b = tf.ones((num_inputs, 1)) * 0.01, 0.05
train_data = d2l.synthetic_data(true_w, true_b, n_train)
train_iter = d2l.load_array(train_data, batch_size)
test_data = d2l.synthetic_data(true_w, true_b, n_test)
test_iter = d2l.load_array(test_data, batch_size, is_train=False)
# + [markdown] origin_pos=6
# ## 从零开始实现
#
# 下面我们将从头开始实现权重衰减,只需将$L_2$的平方惩罚添加到原始目标函数中。
#
# ### [**初始化模型参数**]
#
# 首先,我们将定义一个函数来随机初始化模型参数。
#
# + origin_pos=9 tab=["tensorflow"]
def init_params():
w = tf.Variable(tf.random.normal(mean=1, shape=(num_inputs, 1)))
b = tf.Variable(tf.zeros(shape=(1, )))
return [w, b]
# + [markdown] origin_pos=10
# ### (**定义$L_2$范数惩罚**)
#
# 实现这一惩罚最方便的方法是对所有项求平方后并将它们求和。
#
# + origin_pos=13 tab=["tensorflow"]
def l2_penalty(w):
return tf.reduce_sum(tf.pow(w, 2)) / 2
# + [markdown] origin_pos=14
# ### [**定义训练代码实现**]
#
# 下面的代码将模型拟合训练数据集,并在测试数据集上进行评估。
# 从 :numref:`chap_linear`以来,线性网络和平方损失没有变化,
# 所以我们通过`d2l.linreg`和`d2l.squared_loss`导入它们。
# 唯一的变化是损失现在包括了惩罚项。
#
# + origin_pos=17 tab=["tensorflow"]
def train(lambd):
w, b = init_params()
net, loss = lambda X: d2l.linreg(X, w, b), d2l.squared_loss
num_epochs, lr = 100, 0.003
animator = d2l.Animator(xlabel='epochs', ylabel='loss', yscale='log',
xlim=[5, num_epochs], legend=['train', 'test'])
for epoch in range(num_epochs):
for X, y in train_iter:
with tf.GradientTape() as tape:
# 增加了L2范数惩罚项,
# 广播机制使l2_penalty(w)成为一个长度为batch_size的向量
l = loss(net(X), y) + lambd * l2_penalty(w)
grads = tape.gradient(l, [w, b])
d2l.sgd([w, b], grads, lr, batch_size)
if (epoch + 1) % 5 == 0:
animator.add(epoch + 1, (d2l.evaluate_loss(net, train_iter, loss),
d2l.evaluate_loss(net, test_iter, loss)))
print('w的L2范数是:', tf.norm(w).numpy())
# + [markdown] origin_pos=18
# ### [**忽略正则化直接训练**]
#
# 我们现在用`lambd = 0`禁用权重衰减后运行这个代码。
# 注意,这里训练误差有了减少,但测试误差没有减少,
# 这意味着出现了严重的过拟合。
#
# + origin_pos=19 tab=["tensorflow"]
train(lambd=0)
# + [markdown] origin_pos=20
# ### [**使用权重衰减**]
#
# 下面,我们使用权重衰减来运行代码。
# 注意,在这里训练误差增大,但测试误差减小。
# 这正是我们期望从正则化中得到的效果。
#
# + origin_pos=21 tab=["tensorflow"]
train(lambd=3)
# + [markdown] origin_pos=22
# ## [**简洁实现**]
#
# 由于权重衰减在神经网络优化中很常用,
# 深度学习框架为了便于我们使用权重衰减,
# 将权重衰减集成到优化算法中,以便与任何损失函数结合使用。
# 此外,这种集成还有计算上的好处,
# 允许在不增加任何额外的计算开销的情况下向算法中添加权重衰减。
# 由于更新的权重衰减部分仅依赖于每个参数的当前值,
# 因此优化器必须至少接触每个参数一次。
#
# + [markdown] origin_pos=25 tab=["tensorflow"]
# 在下面的代码中,我们使用权重衰减超参数`wd`创建一个$L_2$正则化器,
# 并通过`kernel_regularizer`参数将其应用于网络层。
#
# + origin_pos=28 tab=["tensorflow"]
def train_concise(wd):
net = tf.keras.models.Sequential()
net.add(tf.keras.layers.Dense(
1, kernel_regularizer=tf.keras.regularizers.l2(wd)))
net.build(input_shape=(1, num_inputs))
w, b = net.trainable_variables
loss = tf.keras.losses.MeanSquaredError()
num_epochs, lr = 100, 0.003
trainer = tf.keras.optimizers.SGD(learning_rate=lr)
animator = d2l.Animator(xlabel='epochs', ylabel='loss', yscale='log',
xlim=[5, num_epochs], legend=['train', 'test'])
for epoch in range(num_epochs):
for X, y in train_iter:
with tf.GradientTape() as tape:
# tf.keras需要为自定义训练代码手动添加损失。
l = loss(net(X), y) + net.losses
grads = tape.gradient(l, net.trainable_variables)
trainer.apply_gradients(zip(grads, net.trainable_variables))
if (epoch + 1) % 5 == 0:
animator.add(epoch + 1, (d2l.evaluate_loss(net, train_iter, loss),
d2l.evaluate_loss(net, test_iter, loss)))
print('w的L2范数:', tf.norm(net.get_weights()[0]).numpy())
# + [markdown] origin_pos=29
# [**这些图看起来和我们从零开始实现权重衰减时的图相同**]。
# 然而,它们运行得更快,更容易实现。
# 对于更复杂的问题,这一好处将变得更加明显。
#
# + origin_pos=30 tab=["tensorflow"]
train_concise(0)
# + origin_pos=31 tab=["tensorflow"]
train_concise(3)
# + [markdown] origin_pos=32
# 到目前为止,我们只接触到一个简单线性函数的概念。
# 此外,由什么构成一个简单的非线性函数可能是一个更复杂的问题。
# 例如,[再生核希尔伯特空间(RKHS)](https://en.wikipedia.org/wiki/Reproducing_kernel_Hilbert_space)
# 允许在非线性环境中应用为线性函数引入的工具。
# 不幸的是,基于RKHS的算法往往难以应用到大型、高维的数据。
# 在这本书中,我们将默认使用简单的启发式方法,即在深层网络的所有层上应用权重衰减。
#
# ## 小结
#
# * 正则化是处理过拟合的常用方法:在训练集的损失函数中加入惩罚项,以降低学习到的模型的复杂度。
# * 保持模型简单的一个特别的选择是使用$L_2$惩罚的权重衰减。这会导致学习算法更新步骤中的权重衰减。
# * 权重衰减功能在深度学习框架的优化器中提供。
# * 在同一训练代码实现中,不同的参数集可以有不同的更新行为。
#
# ## 练习
#
# 1. 在本节的估计问题中使用$\lambda$的值进行实验。绘制训练和测试精度关于$\lambda$的函数。你观察到了什么?
# 1. 使用验证集来找到最佳值$\lambda$。它真的是最优值吗?这有关系吗?
# 1. 如果我们使用$\sum_i |w_i|$作为我们选择的惩罚($L_1$正则化),那么更新方程会是什么样子?
# 1. 我们知道$\|\mathbf{w}\|^2 = \mathbf{w}^\top \mathbf{w}$。你能找到类似的矩阵方程吗(见 :numref:`subsec_lin-algebra-norms` 中的Frobenius范数)?
# 1. 回顾训练误差和泛化误差之间的关系。除了权重衰减、增加训练数据、使用适当复杂度的模型之外,你还能想出其他什么方法来处理过拟合?
# 1. 在贝叶斯统计中,我们使用先验和似然的乘积,通过公式$P(w \mid x) \propto P(x \mid w) P(w)$得到后验。如何得到带正则化的$P(w)$?
#
# + [markdown] origin_pos=35 tab=["tensorflow"]
# [Discussions](https://discuss.d2l.ai/t/1809)
#
| tensorflow/chapter_multilayer-perceptrons/weight-decay.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: myDL
# language: python
# name: mydl
# ---
# + [markdown] colab_type="text" id="QAILTZn5AWjr"
# ## Learning Generalized Quasi-Geostrophic Models Using Deep Neural Numerical Models
#
# Lguensat et al. https://arxiv.org/abs/1911.08856
# -
# This notebook presents the PyTorch written one-layer quasi geostrophic model used for the paper, this code is adapted for GPU use and could serve for training neural networks as done in the paper. To run properly you must use double precision (float64). For the moment it does not deal with land pixels, any help/collaboration is appreciated.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="KnV1ZGiRAWjs"
# %matplotlib inline
# standard imports
import numpy as np
#import xarray as xr
#import netCDF4
import time
import math
from math import pi
#import seaborn as sns
import matplotlib.pyplot as plt
# + [markdown] colab_type="text" id="hB1riIIEAWj7"
# ### loading datasets (region near the Gulf stream, NATL60 data)
# https://meom-group.github.io/swot-natl60/access-data.html
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 37664, "status": "ok", "timestamp": 1531728024107, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-9SK_Y43vRwM/AAAAAAAAAAI/AAAAAAAATS8/VfJspu_Ywhk/s50-c-k-no/photo.jpg", "userId": "104960468215040209517"}, "user_tz": -120} id="njGHPCXHaXm_" outputId="f4d60cb4-92f9-4ac5-b335-124adb558a56"
SSH_train = np.expand_dims(np.load('./Data/SSH_365Train_GS.npy'),1)[:,:,::-1,:].astype('float64')
lon_train = np.tile(np.load('./Data/LON_GS.npy')[::-1,:],(365,1,1,1)).astype('float64')
lat_train = np.tile(np.load('./Data/LAT_GS.npy')[::-1,:],(365,1,1,1)).astype('float64')
SSH_train.shape, lon_train.shape, lat_train.shape
# + [markdown] colab_type="text" id="2di4psmaAWkO"
# ### import PyTorch
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 10739, "status": "ok", "timestamp": 1531728060319, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-9SK_Y43vRwM/AAAAAAAAAAI/AAAAAAAATS8/VfJspu_Ywhk/s50-c-k-no/photo.jpg", "userId": "104960468215040209517"}, "user_tz": -120} id="Pd7jbAeoPgG2" outputId="51181b07-52c9-4014-f2f0-f44b07d7d5e9"
import torch
torch.manual_seed(42)
import torch.nn as nn
import torch.nn.functional as F
import torch.utils.data as Data
# -
torch.backends.cudnn.deterministic = True
torch.__version__
# !nvidia-smi
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
torch.cuda.set_device(0)
print(device)
# ## One-layer QG in PyTorch (written for GPU use)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 2040} colab_type="code" executionInfo={"elapsed": 956, "status": "ok", "timestamp": 1531728062998, "user": {"displayName": "<NAME>", "photoUrl": "//lh5.googleusercontent.com/-9SK_Y43vRwM/AAAAAAAAAAI/AAAAAAAATS8/VfJspu_Ywhk/s50-c-k-no/photo.jpg", "userId": "104960468215040209517"}, "user_tz": -120} id="8iy0jxph3xZZ" outputId="4e412437-d000-4c57-cd58-e33e9e9d3532"
class sshtoqRegression(nn.Module):
def __init__(self, qg, nbiter):
super(sshtoqRegression, self).__init__()
self.qg = qg
self.nbiter = nbiter
def forward(self, x):
self.gradient_x_filter = torch.cuda.DoubleTensor([-0.5, +0.5]).view([1, 1, 1, 2])
self.gradient_x = torch.cuda.DoubleTensor([[-0.25, -0.25], [0, 0], [0.25, 0.25]]).view([1, 1, 3, 2])
self.gradient_y_filter = torch.transpose(self.gradient_x_filter,2,3)
self.gradient_y = torch.transpose(self.gradient_x,2,3)
self.laplacian_x = torch.cuda.DoubleTensor([1, -2, 1]).view([1, 1, 1, 3])
self.laplacian_y = torch.transpose(self.laplacian_x,2,3)
self.diff_x_filter = torch.cuda.DoubleTensor([-1, +1]).view([1, 1, 1, 2])
self.diff_y_filter = torch.transpose(self.diff_x_filter,2,3)
self.gr1=torch.cuda.DoubleTensor([0.5, 0.5]).view([1, 1, 1, 2])
self.gr2=torch.transpose(self.gr1,2,3)
self.qf1=torch.cuda.DoubleTensor([1/6, -1, 1/2, 1/3]).view([1, 1, 1, 4])
self.qf2=torch.transpose(self.qf1,2,3)
self.qf1bis=torch.cuda.DoubleTensor([1/3, 1/2, -1, 1/6]).view([1, 1, 1, 4])
self.qf2bis=torch.transpose(self.qf1bis, 2,3)
self.diffv2_y_filter= torch.cuda.DoubleTensor([-1,0,+1]).view([1, 1, 3, 1])
#########
ssh_tensor,lon_input,lat_input = x
q = self.sshtoq((ssh_tensor,lon_input,lat_input))
q10 = self.dynqhardcoded((q,ssh_tensor,lon_input,lat_input))
hh = self.CGblockZERO((q10,ssh_tensor,lon_input,lat_input)) #, rn, dn
#######
hb = ssh_tensor#hh
for iterat in range(self.nbiter):
hguess = 2*hh-hb
hb = hh
q10 = self.dynqhardcoded((q10,hh,lon_input,lat_input))
hh = self.CGblockZERO((q10,hguess,lon_input,lat_input))#, rn, dn
return hh
def sshtoq(self, x):
ssh_tensor,lon_input,lat_input = x
dx,dy,f0 = self.gridvar([lon_input,lat_input])
laplaciansshx = F.conv2d(ssh_tensor, self.laplacian_x, padding=0)[:,:,2:-3,1:-2]*((9.81/(f0*(dx**2)))[:,:,2:-3,2:-3])
laplaciansshy = F.conv2d(ssh_tensor, self.laplacian_y, padding=0)[:,:,1:-2,2:-3]*((9.81/(f0*(dy**2)))[:,:,2:-3,2:-3])
zp = nn.ZeroPad2d((2, 3, 2, 3))
if self.qg:
return (-9.81*f0/(1.5**2))*ssh_tensor+ zp(laplaciansshx)+ zp(laplaciansshy) ##1.5 is the constant Rossby first baroclinic phase speed set to constant value in m/s
else:
return ###put here your neural net
def Geostroph(self, x):
ssh_tensor,lon_input,lat_input = x
dx,dy,f0 = self.gridvar([lon_input,lat_input])
zp1 = nn.ZeroPad2d((1,1,1,0))
zp2 = nn.ZeroPad2d((1,0,1,1))
vvv_QG = zp1(F.conv2d(ssh_tensor,self.gradient_y,padding=0))
uuu_QG = zp2(F.conv2d(ssh_tensor,self.gradient_x,padding=0))
return -9.81*uuu_QG/(f0*dy),9.81*vvv_QG/(f0*dx)
def dynqhardcoded(self, x):
q_tensor,ssh_tensor,lon_input,lat_input = x
dx,dy,f0 = self.gridvar([lon_input,lat_input])
zp1 = nn.ZeroPad2d((1,1,1,0))
zp2 = nn.ZeroPad2d((1,0,1,1))
vvv_QG = zp1(F.conv2d(ssh_tensor,self.gradient_y,padding=0))
uuu_QG = zp2(F.conv2d(ssh_tensor,self.gradient_x,padding=0))
if self.qg:
v = 9.81*vvv_QG/(f0*dx)
u = -9.81*uuu_QG/(f0*dy)
else:
v # put your neural net
u # put your neural net
#####
utmp = F.conv2d(u, self.gr1, padding=0)[:,:,2:-2,2:-1]
uplus = F.relu(utmp)
uminus = -1*F.relu(-1*utmp)
vtmp = F.conv2d(v, self.gr2, padding=0)[:,:,2:-1,2:-2]
vplus = F.relu(vtmp)
vminus = -1*F.relu(-1*vtmp)
#########
q1= F.conv2d(q_tensor[:,:,2:-2,0:-1], self.qf1)
q2= F.conv2d(q_tensor[:,:,2:-2,1:], self.qf1bis)
q3= F.conv2d(q_tensor[:,:,:-1,2:-2], self.qf2)
q4= F.conv2d(q_tensor[:,:,1:,2:-2], self.qf2bis)
#########
adding1 = (uminus*q2-uplus*q1)/(dx[:,:,2:-2,2:-2])
adding2 = (vminus*q4-vplus*q3)/(dy[:,:,2:-2,2:-2])
f0tmp = F.conv2d(f0[:,:,1:-1,2:-2], self.diffv2_y_filter)
vtmp2 = ((-1.*f0tmp)/(2.*dy[:,:,2:-2,2:-2]))*vtmp
rq = (adding1+adding2+vtmp2)[:,:,:-1,:-1]
########
zp3 = nn.ZeroPad2d((2,3,2,3))
return q_tensor + 600.*zp3(rq) #_______Attention______________dt=10min=600s
def CGblockZERO(self, x):
## one iteration of the conjugate gradient
q_tensor, map_tensor, lon_input, lat_input = x
Fssh = self.sshtoq((map_tensor, lon_input, lat_input))
rr = q_tensor-Fssh
Frr = self.sshtoq((rr, lon_input, lat_input))
########## alpha
alpha = torch.sum(rr*rr, dim=(1,2,3), keepdim=True)/torch.sum(rr*Frr, dim=(1,2,3), keepdim=True)
##############
mapnext = map_tensor + alpha*rr
#rrnext = q_tensor - self.sshtoq((mapnext, lon_input, lat_input))
########## beta
#beta = torch.sum(rrnext*rrnext, dim=(1,2,3), keepdim=True)/torch.sum(rr*rr, dim=(1,2,3), keepdim=True)
##############
#dir1 = rrnext + beta*rr
return mapnext#, rrnext, dir1
def gridvar(self, x):
lon_tensor,lat_tensor = x
f0= 2.*2.*pi/86164.*torch.sin(lat_tensor*pi/180.)
dlony=F.conv2d(lon_tensor[:,:,1:,1:-1], self.diff_y_filter, padding=0)
dlaty=F.conv2d(lat_tensor[:,:,1:,1:-1], self.diff_y_filter, padding=0)
dlonx=F.conv2d(lon_tensor[:,:,1:-1,1:], self.diff_x_filter, padding=0)
dlatx=F.conv2d(lat_tensor[:,:,1:-1,1:], self.diff_x_filter, padding=0)
tmp=torch.cos(lat_tensor[:,:,1:-1,1:-1]*pi/180.)
dx=torch.sqrt((dlonx*111000.*tmp)**2 + (dlatx*111000.)**2)
dy=torch.sqrt((dlony*111000.*tmp)**2 + (dlaty*111000.)**2)
return F.pad(dx,(1,1,1,1),'replicate'), F.pad(dy,(1,1,1,1),'replicate'), f0
# -
# ## Test
randindex = np.random.randint(365)
randindex
# #### the code already does one iteration, so for a 24h SSH forecasting with a 10min time step, 144-1=143 steps are needed
# +
netqg = sshtoqRegression(qg=True, nbiter=143).double().to(device)
netqg.eval()
res_qg = netqg([torch.from_numpy(SSH_train[None,randindex,:,:,:]).to(torch.float64).to(device),
torch.from_numpy(lon_train[None,randindex,:,:,:]).to(torch.float64).to(device),
torch.from_numpy(lat_train[None,randindex,:,:,:]).to(torch.float64).to(device)])
# -
# ## Comparison with the original Python code
import sys
sys.path.insert(0,'/home/lguensar/deepocean/QGmodel')
from moddyn import h2uv,qrhs
from modelliptic import pv2h, h2pv
import modgrid
#####
# +
c=SSH_train[0,0,:,:]*0. + 1.5 # Set constant Rossby first baroclinic phase speed to constant value in m/s
dt=600 ##10min propagator time step
h = SSH_train[randindex,0,:,:]
grdtst=modgrid.grid(h, c, 0., lon_train[randindex,0,:,:], lat_train[randindex,0,:,:])
qQG,= h2pv(h,grdtst)
hb=+h
for step in range(144): # 144 because 10min
#init
hguess=2*h-hb
hb=+h
qb=+qQG
#forecasting
# 1/
u,v, = h2uv(h,grdtst)
# 2/
rq, = qrhs(u,v,qb,grdtst,+1)
# 3/
qQG = qb + dt*rq
# 4/
h,=pv2h(qQG,hguess,grdtst,1) ##### one CG iteration
# +
plt.figure(figsize=(20, 10))
plt.subplot(141)
plt.contour(SSH_train[randindex,0,:,:], cmap='viridis')
plt.imshow(SSH_train[randindex,0,:,:], cmap='coolwarm')
plt.colorbar(extend='both', fraction=0.042, pad=0.04)
#plt.clim(-0.5,0.5)
plt.title('SSH_0_True')
plt.subplot(142)
plt.contour((res_qg[0,0,:,:]).detach().cpu().numpy(), cmap='viridis')
plt.imshow((res_qg[0,0,:,:]).detach().cpu().numpy(), cmap='coolwarm')
plt.colorbar(extend='both', fraction=0.042, pad=0.04)
#plt.clim(-0.5,0.5)
plt.title('SSH_1days_QG_pytorch')
plt.subplot(143)
plt.contour(h, cmap='viridis')
plt.imshow(h, cmap='coolwarm')
plt.colorbar(extend='both', fraction=0.042, pad=0.04)
#plt.clim(-0.5,0.5)
plt.title('SSH_1days_QG_original')
plt.subplot(144)
plt.imshow(h-(res_qg[0,0,:,:]).detach().cpu().numpy(), cmap='seismic')
plt.colorbar(extend='both', fraction=0.042, pad=0.04)
#plt.clim(-0.2,0.2)
plt.title('qg_pytorch error')
# +
from sklearn.metrics import mean_squared_error
mean_squared_error(h,res_qg[0,0,:,:].detach().cpu().numpy())
# -
# ## test the stability, check forecasting at 20 days
# +
netqg = sshtoqRegression(qg=True, nbiter=20*144-1).double().to(device)
netqg.eval()
res_qg = netqg([torch.from_numpy(SSH_train[None,randindex,:,:,:]).to(torch.float64).to(device),
torch.from_numpy(lon_train[None,randindex,:,:,:]).to(torch.float64).to(device),
torch.from_numpy(lat_train[None,randindex,:,:,:]).to(torch.float64).to(device)])
# +
c=SSH_train[0,0,:,:]*0. + 1.5 # Set constant Rossby first baroclinic phase speed to constant value in m/s
dt=600 ##10min propagator time step
h = SSH_train[randindex,0,:,:]
grdtst=modgrid.grid(h, c, 0., lon_train[randindex,0,:,:], lat_train[randindex,0,:,:])
qQG,= h2pv(h,grdtst)
hb=+h
for step in range(144*20): # 144 because 10min
#init
hguess=2*h-hb
hb=+h
qb=+qQG
#forecasting
# 1/
u,v, = h2uv(h,grdtst)
# 2/
rq, = qrhs(u,v,qb,grdtst,+1)
# 3/
qQG = qb + dt*rq
# 4/
h,=pv2h(qQG,hguess,grdtst,1) ##### one CG iteration
# +
plt.figure(figsize=(20, 10))
plt.subplot(141)
plt.contour(SSH_train[randindex,0,:,:], cmap='viridis')
plt.imshow(SSH_train[randindex,0,:,:], cmap='coolwarm')
plt.colorbar(extend='both', fraction=0.042, pad=0.04)
#plt.clim(-0.5,0.5)
plt.title('SSH_0_True')
plt.subplot(142)
plt.contour((res_qg[0,0,:,:]).detach().cpu().numpy(), cmap='viridis')
plt.imshow((res_qg[0,0,:,:]).detach().cpu().numpy(), cmap='coolwarm')
plt.colorbar(extend='both', fraction=0.042, pad=0.04)
#plt.clim(-0.5,0.5)
plt.title('SSH_20days_QG_pytorch')
plt.subplot(143)
plt.contour(h, cmap='viridis')
plt.imshow(h, cmap='coolwarm')
plt.colorbar(extend='both', fraction=0.042, pad=0.04)
#plt.clim(-0.5,0.5)
plt.title('SSH_20days_QG_original')
plt.subplot(144)
plt.imshow(h-(res_qg[0,0,:,:]).detach().cpu().numpy(), cmap='seismic')
plt.colorbar(extend='both', fraction=0.042, pad=0.04)
#plt.clim(-0.2,0.2)
plt.title('qg_pytorch error')
# -
mean_squared_error(h,res_qg[0,0,:,:].detach().cpu().numpy())
| QGNET/QG_PyTorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Experiment 1
#
# - Fine-tuning of the pre-trained TAPE model in a progressively specialized manner while extending the frozen layers
# - Performance evaluation using two external datasets, Shomuradova and ImmuneCODE datatsets, which contain SARS-CoV-2 S-protein <sub>269~-277</sub>(YLQPRTFLL) epitope-specific TCR CDR3βs
# ## Training process and model architecture
#
#
# <img src='res/fig1.png' width='70%'>
#
# Training process for the proposed model. The initial model is cloned from pre-trained Tasks Assessing Protein Embeddings (TAPE) model, adding a classification layer at the end. The pre-trained model is fine-tuned in two rounds in a progressively specialized manner while extending the frozen layers between rounds.
#
# <img src='res/fig2.png' width='30%'>
#
# The proposed model architecture. Input amino acid sequences concatenated by epitope and CDR3β sequences are first encoded into tokens using a tokenizer. Each token is then embedded into a 768-dimensional vector in the pre-trained Tasks Assessing Protein Embeddings (TAPE) model which has 12 encoding layers with 12 self-attention heads in each layer. The final classifier, a 2-layer feed forward network, is then used to predict either binder or not from the output of the TAPE model.
# ## Global configurations
# +
import logging
import logging.config
import os
import sys
import warnings
from enum import auto
import pandas as pd
import numpy as np
from IPython.core.display import display
rootdir = '/home/hym/trunk/TCRBert'
workdir = '%s/notebook' % rootdir
datadir = '%s/data' % rootdir
srcdir = '%s/tcrbert' % rootdir
outdir = '%s/output' % rootdir
os.chdir(workdir)
sys.path.append(rootdir)
sys.path.append(srcdir)
from tcrbert.exp import Experiment
from tcrbert.predlistener import PredResultRecoder
# Display
pd.set_option('display.max.rows', 2000)
pd.set_option('display.max.columns', 2000)
# Logger
warnings.filterwarnings('ignore')
logging.config.fileConfig('../config/logging.conf')
logger = logging.getLogger('tcrbert')
logger.setLevel(logging.INFO)
# Target experiment
exp_key = 'exp1'
experiment = Experiment.from_key(exp_key)
exp_conf = experiment.exp_conf
display(exp_conf)
# -
# ## Fine-tuning results
# +
# %pylab inline
def show_train_result(train_result=None, ax=None, title=None):
df = pd.DataFrame({
'train_loss': train_result['train.score']['loss'],
'val_loss': train_result['val.score']['loss'],
'train_accuracy': train_result['train.score']['accuracy'],
'val_accuracy': train_result['val.score']['accuracy']
})
ax = df.loc[:, ['train_accuracy', 'val_accuracy']].plot(ax=ax, title=title)
bs = train_result['best_score']
n_epochs = train_result['stopped_epoch']
ax.axhline(y=bs, ls='--', c='red')
ax.xaxis.set_major_locator(MaxNLocator(integer=True))
ax.legend(["Train accuracy", "Validation accuracy", "Best accuracy = %0.3f" % bs], prop={"size": 11})
print('##############################')
print('n_epochs: %s' % train_result['n_epochs'])
print('stopped_epoch: %s' % train_result['stopped_epoch'])
print('best_epoch: %s' % train_result['best_epoch'])
print('best_score: %s' % train_result['best_score'])
print('best_chk: %s' % train_result['best_chk'])
fig, axes = plt.subplots(nrows=1, ncols=2)
fig.set_figwidth(10)
fig.set_figheight(4)
plt.tight_layout(h_pad=2, w_pad=2)
show_train_result(train_result=experiment.get_train_result(0),
ax=axes[0])
show_train_result(train_result=experiment.get_train_result(1),
ax=axes[1])
# -
# ## Loading final fine-tuned model
# +
model = experiment.load_eval_model()
display(model)
# Eval result recoder
eval_recoder = PredResultRecoder(output_attentions=True, output_hidden_states=True)
model.add_pred_listener(eval_recoder)
# -
# ## Model evaluations
# ### Shomuradova <i>et al.</i>
#
# - Performance evaluation for the dataset containing COVID-19 S-protein<sub>266-277</sub>(YLQPRTFLL) epitope-specific CDR3βs from Shomuradova <i>et al.</i>
# #### Loading eval dataset
# +
from torch.utils.data import DataLoader
from tcrbert.dataset import TCREpitopeSentenceDataset
import numpy as np
from tcrbert.dataset import CN
from tcrbert.bioseq import write_fa
epitope = 'YLQPRTFLL'
sh_ds = TCREpitopeSentenceDataset.from_key('shomuradova')
sh_df = sh_ds.df_enc
print(sh_ds.name)
display(sh_df.head(), sh_df.shape)
summary_df(sh_df)
# -
# #### Performance evaluation for Shomuradova dataset
# +
# %pylab inline
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.metrics import roc_curve, auc
import numpy as np
metrics = ['accuracy', 'f1', 'roc_auc']
data_loader = DataLoader(sh_ds, batch_size=len(sh_ds), shuffle=False, num_workers=2)
model.predict(data_loader=data_loader, metrics=metrics)
print('score_map: %s' % eval_recoder.result_map['score_map'])
input_labels = np.array(eval_recoder.result_map['input_labels'])
output_labels = np.array(eval_recoder.result_map['output_labels'])
output_probs = np.array(eval_recoder.result_map['output_probs'])
fpr, tpr, _ = roc_curve(input_labels, output_probs)
score = auc(fpr, tpr)
plt.plot(fpr, tpr, color='darkorange', label='ROC curve (AUROC = %0.3f)' % score)
plt.plot([0, 1], [0, 1], color='navy', linestyle='--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.legend(loc="lower right")
# Epitope-specific ROC curves
# epitopes = sh_df[CN.epitope].unique()
# fig, axes = plt.subplots(nrows=epitopes.shape[0], ncols=1)
# fig.set_figwidth(5)
# fig.set_figheight(4*len(epitopes))
# plt.tight_layout(h_pad=5, w_pad=3)
# for i, epitope in enumerate(epitopes):
# indices = np.where(sh_df[CN.epitope] == epitope)[0]
# # title = '%s-specific CDR3βs' % epitope
# print(indices, len(indices))
# title = None
# ax = axes[i] if epitopes.shape[0] > 1 else axes
# show_roc_curve(input_labels[indices], output_probs[indices], title=title, ax=ax)
# -
# #### Investigating position-wise attention weights
#
# ##### Selecting target CDR3β sequences with most common lengths and predicted as a binder
# +
# %pylab inline
import matplotlib.pyplot as plt
import matplotlib.image as image
import seaborn as sns
from collections import Counter, OrderedDict
pos_indices = np.where(output_labels == 1)[0]
# print('pos_indices: %s(%s)' % (pos_indices, str(pos_indices.shape)))
pos_cdr3b = sh_df[CN.cdr3b].values[pos_indices]
##########################################
lens, cnts = zip(*sorted(Counter(map(lambda x: len(x), pos_cdr3b)).items()))
lens = np.array(lens)
cnts = np.array(cnts)
ax = sns.barplot(x=lens, y=cnts, color='blue')
ax.set_title('Length distribution of postives')
ax.set_xlabel('CDR3β sequence length')
ax.set_ylabel('# of positives')
# Select target indices by cdr3b sequence lenghts
target_index_map = OrderedDict()
order = np.argsort(cnts)[::-1]
cum_cnt = 0
for cur_len, cur_cnt in zip(lens[order], cnts[order]):
cum_cnt += cur_cnt
cum_ratio = cum_cnt/pos_indices.shape[0]
print('cum_ratio: %s/%s=%s' % (cum_cnt, pos_indices.shape[0], cum_ratio))
if cum_ratio < 0.9:
target_indices = np.where((output_labels == 1) & (sh_df[CN.cdr3b].map(lambda x: len(x) == cur_len)))[0]
print('target_indices for %s: %s(%s)' % (cur_len, target_indices, target_indices.shape[0]))
target_index_map[cur_len] = target_indices
# -
# ##### Marginalized position-wise attention weights for target YLQPRTFLL-CDR3b sequences
#
# - The output attention weights have the dimension $(L, N, H, S, S)$, where $L$ is the number of encoding layers, $N$ is the number of YLQPRTFLL-CDR3β sequence pairs, $H$ is the number of attention heads, and $S$ is the fixed-length of the sequences. The attention weights were marginalized into a one-dimensional vector of length of $S$. A value of the vector at the position $m$, $Am$ is given by the following equation:
#
# <img src='res/eq1.png' width='50%'>
# +
# %pylab inline
# n_layers = model.config.num_hidden_layers
# n_data = len(eval_ds)
# n_heads = model.config.num_attention_heads
# max_len = eval_ds.max_len
epitope_len = len(epitope)
attentions = eval_recoder.result_map['attentions']
# attentions.shape: (n_layers, n_data, n_heads, max_len, max_len)
print('attentions.shape: %s' % str(attentions.shape))
# target_attn_map = OrderedDict()
fig, axes = plt.subplots(nrows=3, ncols=1)
fig.set_figwidth(7)
fig.set_figheight(12)
plt.tight_layout(h_pad=3, w_pad=3)
for i, (cur_len, cur_indices) in enumerate(target_index_map.items()):
attns = attentions[:, cur_indices]
sent_len = epitope_len + cur_len
# Position-wise marginal attentions by mean
attns = np.mean(attns, axis=(0, 1, 2, 3))[1:sent_len+1]
print('Attention weights for %s: %s' % (cur_len, attns))
ax = pd.Series(attns).plot(kind='bar', ax=axes[i], rot=0)
ax.set_title('%smer CDR3β sequences' % cur_len)
ticks = list(epitope) + list(range(1, cur_len + 1))
mark_ratio = 0.1
mark_pos = []
for rank, pos in enumerate(np.argsort(attns[:epitope_len])[::-1]):
if rank < (epitope_len*mark_ratio):
ticks[pos] = '%s\n•' % (ticks[pos])
mark_pos.append(pos)
for rank, pos in enumerate(np.argsort(attns[epitope_len:])[::-1]):
if rank < (cur_len*mark_ratio):
ticks[epitope_len+pos] = '%s\n•' % (ticks[epitope_len+pos])
mark_pos.append(epitope_len+pos)
ax.set_xticklabels(ticks)
for i, tick in enumerate(ax.get_xticklabels()):
if i < epitope_len:
tick.set_color('green')
else:
tick.set_color('black')
if i in mark_pos:
tick.set_color('darkred')
# -
# ### ImmuneCODE dataset
#
# - The dataset contained 718 COVID-19 S-protein<sub>269-277</sub>-specific TCRs from the ImmuneRACE study launched by Adaptive Biotechnologies and Microsoft (https://immunerace.adaptivebiotech.com, June 10, 2020 dataset, from hereafter referred to as ImmuneCODE dataset
# #### Loading eval dataset
# +
im_ds = TCREpitopeSentenceDataset.from_key('immunecode')
display(im_ds.name, len(im_ds))
# Remove duplicated CDR3beta seqs with Shomuradova
im_ds.df_enc = im_ds.df_enc[
im_ds.df_enc[CN.cdr3b].map(lambda seq: seq not in sh_df[CN.cdr3b].values)
]
im_df = im_ds.df_enc
print('After removing duplicates, len(im_ds): %s' % len(im_ds))
display(summary_df(im_df), im_df.shape)
# -
# #### Performance evaluation for the ImmuneCODE dataset
# +
# %pylab inline
from sklearn.metrics import roc_curve, auc
data_loader = DataLoader(im_ds, batch_size=len(im_ds), shuffle=False, num_workers=2)
model.predict(data_loader=data_loader, metrics=metrics)
input_labels = np.array(eval_recoder.result_map['input_labels'])
output_labels = np.array(eval_recoder.result_map['output_labels'])
output_probs = np.array(eval_recoder.result_map['output_probs'])
fpr, tpr, _ = roc_curve(input_labels, output_probs)
score = auc(fpr, tpr)
plt.plot(fpr, tpr, color='darkorange', label='ROC curve (AUROC = %0.3f)' % score)
plt.plot([0, 1], [0, 1], color='navy', linestyle='--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.legend(loc="lower right")
print('score_map: %s' % eval_recoder.result_map['score_map'])
# -
# #### Investigating position-wise attention weights
#
# ##### Selecting target CDR3β sequences with most common lengths and predicted as a binder
# +
# %pylab inline
import matplotlib.pyplot as plt
import matplotlib.image as image
import seaborn as sns
from collections import Counter, OrderedDict
pos_indices = np.where(output_labels == 1)[0]
# print('pos_indices: %s(%s)' % (pos_indices, str(pos_indices.shape)))
pos_cdr3b = im_df[CN.cdr3b].values[pos_indices]
##########################################
lens, cnts = zip(*sorted(Counter(map(lambda x: len(x), pos_cdr3b)).items()))
lens = np.array(lens)
cnts = np.array(cnts)
ax = sns.barplot(x=lens, y=cnts, color='blue')
ax.set_title('Length distribution of postives')
ax.set_xlabel('Length')
# Select target indices by cdr3b sequence lenghts
target_index_map = OrderedDict()
order = np.argsort(cnts)[::-1]
cum_cnt = 0
for cur_len, cur_cnt in zip(lens[order], cnts[order]):
cum_cnt += cur_cnt
cum_ratio = cum_cnt/pos_indices.shape[0]
print('cum_ratio: %s/%s=%s' % (cum_cnt, pos_indices.shape[0], cum_ratio))
if cum_ratio < 0.85:
target_indices = np.where((output_labels == 1) & (im_df[CN.cdr3b].map(lambda x: len(x) == cur_len)))[0]
print('target_indices for %s: %s(%s)' % (cur_len, target_indices, target_indices.shape[0]))
target_index_map[cur_len] = target_indices
# -
# ##### Marginalized position-wise attention weights for target YLQPRTFLL-CDR3b sequences
# +
# %pylab inline
# n_layers = model.config.num_hidden_layers
# n_data = len(eval_ds)
# n_heads = model.config.num_attention_heads
# max_len = eval_ds.max_len
epitope_len = len(epitope)
attentions = eval_recoder.result_map['attentions']
# attentions.shape: (n_layers, n_data, n_heads, max_len, max_len)
print('attentions.shape: %s' % str(attentions.shape))
# target_attn_map = OrderedDict()
fig, axes = plt.subplots(nrows=3, ncols=1)
fig.set_figwidth(7)
fig.set_figheight(12)
plt.tight_layout(h_pad=3, w_pad=3)
for i, (cur_len, cur_indices) in enumerate(target_index_map.items()):
attns = attentions[:, cur_indices]
sent_len = epitope_len + cur_len
# Position-wise marginal attentions by mean
attns = np.mean(attns[-10:], axis=(0, 1, 2, 3))[1:sent_len+1]
print('Current attns for %s: %s' % (cur_len, attns))
ax = pd.Series(attns).plot(kind='bar', ax=axes[i], rot=0)
ax.set_title('%smer CDR3β sequences' % cur_len)
ticks = list(epitope) + list(range(1, cur_len + 1))
mark_ratio = 0.1
mark_pos = []
for rank, pos in enumerate(np.argsort(attns[:epitope_len])[::-1]):
if rank < (epitope_len*mark_ratio):
ticks[pos] = '%s\n•' % (ticks[pos])
mark_pos.append(pos)
for rank, pos in enumerate(np.argsort(attns[epitope_len:])[::-1]):
if rank < (cur_len*mark_ratio):
ticks[epitope_len+pos] = '%s\n•' % (ticks[epitope_len+pos])
mark_pos.append(epitope_len+pos)
ax.set_xticklabels(ticks)
for i, tick in enumerate(ax.get_xticklabels()):
if i < epitope_len:
tick.set_color('darkgreen')
else:
tick.set_color('black')
if i in mark_pos:
tick.set_color('darkred')
# -
# ### <NAME>. et al
#
# - Performance evaluation for independent test dataset from the recent pMHC-TCR interaction study[Lu, T. et al, 2021]
# - The dataset contains 619 pMHC-TCR sequence pairs compiled from VDJdb
# - The same number of negative examples were added
# - <NAME>., <NAME>., <NAME>. et al. Deep learning-based prediction of the T cell receptor–antigen binding specificity. Nat Mach Intell 3, 864–875 (2021). https://doi.org/10.1038/s42256-021-00383-2
#
#
# + pycharm={"name": "#%%\n"}
pm_ds = TCREpitopeSentenceDataset.from_key('pTMnet')
display(pm_ds.name, len(pm_ds))
pm_df = pm_ds.df_enc
display(pm_df.head(), pm_df.shape)
# + pycharm={"name": "#%%\n"}
# %pylab inline
from sklearn.metrics import roc_curve, auc
data_loader = DataLoader(pm_ds, batch_size=len(pm_ds), shuffle=False, num_workers=2)
model.predict(data_loader=data_loader, metrics=metrics)
input_labels = np.array(eval_recoder.result_map['input_labels'])
output_labels = np.array(eval_recoder.result_map['output_labels'])
output_probs = np.array(eval_recoder.result_map['output_probs'])
fpr, tpr, _ = roc_curve(input_labels, output_probs)
score = auc(fpr, tpr)
plt.plot(fpr, tpr, color='darkorange', label='ROC curve (AUROC = %0.3f)' % score)
plt.plot([0, 1], [0, 1], color='navy', linestyle='--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.legend(loc="lower right")
print('score_map: %s' % eval_recoder.result_map['score_map'])
| notebook/exp1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/iyengaraditya/iyengaraditya.github.io/blob/master/MA_214_Code_Snippets.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="I3Lq48ZVzuB-" colab_type="text"
# [RUNGE KUTTA METHODS](https://en.wikipedia.org/wiki/Runge%E2%80%93Kutta_methods)
#
# Solving Initial Value Problems in a Closed Interval
# + id="MHacYUTizzbJ" colab_type="code" colab={}
def funct(x, y):
#define function here where y' = f(x, y), y(x0) = y0
return 1
# + id="l59oGIAkzBoU" colab_type="code" colab={}
import numpy as np
def exact_soln(x):
#define exact solution here if required, not needed by default
return ((x+1)*(x+1) - 0.5*np.exp(x))
# + id="WEW42--axZ00" colab_type="code" colab={}
#RUNGE KUTTA ORDER 4
import numpy as np
#enter initial conditions here
#the ivp is of the type y' = f(x, y), y(x0) = y0
#solution is sought in the interval [x0, xn]
#number of steps is N
y0 = 4
x0 = 2
xn = 4
N = 100
y_precision = 6
x_precision = 2
#end of initial conditions
x = x0
i = 0
y = y0
f = 0
h = (xn - x0)/N
k1 = 0
k2 = 0
k3 = 0
k4 = 0
print("x_n" + "\t" + "RK-4 Approximation")
print(str(round(x0, x_precision)) + "\t" + str(round(y0, y_precision)))
for i in range(1, N+1):
#y_exact = round(exact_soln(x+h), y_precision)
k1 = h*funct(x, y)
k2 = h*funct(x+(h/2), y+(k1/2))
k3 = h*funct(x+(h/2), y + (k2/2))
k4 = h*funct(x+h, y+k3)
y = round(y + ((k1 + (2*k2) + (2*k3) + k4))/6, y_precision)
x = round(x + h, x_precision)
#error = (np.abs(y-y_exact))*100/y_exact
#print(str(y_exact) + " " + str(y) + " " + str(error))
print(str(x) + "\t" + str(y))
# + id="IaVc-p-sA1hG" colab_type="code" colab={}
#RUNGE KUTTA ORDER 2
import numpy as np
#enter initial conditions here
#the ivp is of the type y' = f(x, y), y(x0) = y0
#solution is sought in the interval [x0, xn]
#number of steps is N
y0 = 4
x0 = 2
xn = 4
N = 100
y_precision = 6
x_precision = 2
#end of initial conditions
x = x0
i = 0
y = y0
f = 0
h = (xn - x0)/N
k1 = 0
k2 = 0
print("x_n" + "\t" + "RK-4 Approximation")
print(str(round(x0, x_precision)) + "\t" + str(round(y0, y_precision)))
for i in range(1, N+1):
#y_exact = round(exact_soln(x+h), y_precision)
k1 = h*funct(x, y)
k2 = h*funct(x+h, y+k1)
y = round(y + (k1 + k2)/2, y_precision)
x = round(x + h, x_precision)
#error = (np.abs(y-y_exact))*100/y_exact
#print(str(y_exact) + " " + str(y) + " " + str(error))
print(str(x) + "\t" + str(y))
| MA_214_Code_Snippets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Self-Driving Car Engineer Nanodegree
#
# ## Deep Learning
#
# ## Project: Build a Traffic Sign Recognition Classifier
#
# In this notebook, a template is provided for you to implement your functionality in stages, which is required to successfully complete this project. If additional code is required that cannot be included in the notebook, be sure that the Python code is successfully imported and included in your submission if necessary.
#
# > **Note**: Once you have completed all of the code implementations, you need to finalize your work by exporting the iPython Notebook as an HTML document. Before exporting the notebook to html, all of the code cells need to have been run so that reviewers can see the final implementation and output. You can then export the notebook by using the menu above and navigating to \n",
# "**File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
#
# In addition to implementing code, there is a writeup to complete. The writeup should be completed in a separate file, which can be either a markdown file or a pdf document. There is a [write up template](https://github.com/udacity/CarND-Traffic-Sign-Classifier-Project/blob/master/writeup_template.md) that can be used to guide the writing process. Completing the code template and writeup template will cover all of the [rubric points](https://review.udacity.com/#!/rubrics/481/view) for this project.
#
# The [rubric](https://review.udacity.com/#!/rubrics/481/view) contains "Stand Out Suggestions" for enhancing the project beyond the minimum requirements. The stand out suggestions are optional. If you decide to pursue the "stand out suggestions", you can include the code in this Ipython notebook and also discuss the results in the writeup file.
#
#
# >**Note:** Code and Markdown cells can be executed using the **Shift + Enter** keyboard shortcut. In addition, Markdown cells can be edited by typically double-clicking the cell to enter edit mode.
# ---
# ## Step 0: Load The Data
# +
# Load pickled data
import pickle
# TODO: Fill this in based on where you saved the training and testing data
#add new rotated images in a training file to help with processing time (turn this off for submission)
use_pre_rotate = 0
if use_pre_rotate:
new_training_file = 'data.p'
with open(new_training_file, mode='rb') as f:
train = pickle.load(f)
X_train, y_train = train['Training'], train['Labels']
else:
training_file = 'train.p'
with open(training_file, mode='rb') as f:
train = pickle.load(f)
X_train, y_train = train['features'], train['labels']
validation_file='valid.p'
testing_file = 'test.p'
with open(validation_file, mode='rb') as f:
valid = pickle.load(f)
with open(testing_file, mode='rb') as f:
test = pickle.load(f)
X_valid, y_valid = valid['features'], valid['labels']
X_test, y_test = test['features'], test['labels']
# -
# ---
#
# ## Step 1: Dataset Summary & Exploration
#
# The pickled data is a dictionary with 4 key/value pairs:
#
# - `'features'` is a 4D array containing raw pixel data of the traffic sign images, (num examples, width, height, channels).
# - `'labels'` is a 1D array containing the label/class id of the traffic sign. The file `signnames.csv` contains id -> name mappings for each id.
# - `'sizes'` is a list containing tuples, (width, height) representing the original width and height the image.
# - `'coords'` is a list containing tuples, (x1, y1, x2, y2) representing coordinates of a bounding box around the sign in the image. **THESE COORDINATES ASSUME THE ORIGINAL IMAGE. THE PICKLED DATA CONTAINS RESIZED VERSIONS (32 by 32) OF THESE IMAGES**
#
# Complete the basic data summary below. Use python, numpy and/or pandas methods to calculate the data summary rather than hard coding the results. For example, the [pandas shape method](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.shape.html) might be useful for calculating some of the summary results.
# ### Provide a Basic Summary of the Data Set Using Python, Numpy and/or Pandas
# +
### Replace each question mark with the appropriate value.
### Use python, pandas or numpy methods rather than hard coding the results
#we know that the images have all been resized, so grab the shape of an image
h_image, w_image, layer_image = X_train[0].shape
# TODO: Number of training examples
n_train = len(X_train)
# TODO: Number of validation examples
n_validation = len(X_valid)
# TODO: Number of testing examples.
n_test = len(X_test)
# TODO: What's the shape of an traffic sign image?
image_shape = (h_image, w_image)
# TODO: How many unique classes/labels there are in the dataset.
n_classes = len(set(y_train))
print("Number of training examples =", n_train)
print("Number of validation examples =", n_validation)
print("Number of testing examples =", n_test)
print("Image data shape =", image_shape)
print("Number of classes =", n_classes)
# -
# ### Include an exploratory visualization of the dataset
# Visualize the German Traffic Signs Dataset using the pickled file(s). This is open ended, suggestions include: plotting traffic sign images, plotting the count of each sign, etc.
#
# The [Matplotlib](http://matplotlib.org/) [examples](http://matplotlib.org/examples/index.html) and [gallery](http://matplotlib.org/gallery.html) pages are a great resource for doing visualizations in Python.
#
# **NOTE:** It's recommended you start with something simple first. If you wish to do more, come back to it after you've completed the rest of the sections. It can be interesting to look at the distribution of classes in the training, validation and test set. Is the distribution the same? Are there more examples of some classes than others?
# +
### Data exploration visualization code goes here.
### Feel free to use as many code cells as needed.
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
# Visualizations will be shown in the notebook.
# %matplotlib inline
#Plot a few of the traffic signs in a 3x3 output. randomize to get an idea of various signs
f, (ax1, ax2, ax3) = plt.subplots(1, 3)
ax1.imshow(X_train[np.random.randint(0,n_train)])
ax2.imshow(X_train[np.random.randint(0,n_train)])
ax3.imshow(X_train[np.random.randint(0,n_train)])
f, (ax1, ax2, ax3) = plt.subplots(1, 3)
ax1.imshow(X_train[np.random.randint(0,n_train)])
ax2.imshow(X_train[np.random.randint(0,n_train)])
ax3.imshow(X_train[np.random.randint(0,n_train)])
f, (ax1, ax2, ax3) = plt.subplots(1, 3)
ax1.imshow(X_train[np.random.randint(0,n_train)])
ax2.imshow(X_train[np.random.randint(0,n_train)])
ax3.imshow(X_train[np.random.randint(0,n_train)])
#f, (ax1, ax2, ax3) = plt.subplots(1, 3)
#print(X_train[found[0][0]])
#ax1.imshow(X_train[found[0][0]])
#ax2.imshow(X_train[np.random.randint(0,n_train)])
#ax3.imshow(X_train[np.random.randint(0,n_train)])
# +
#make a histogram of each class of sign
#if the data are loaded from the new pickle file, this histogram is affected
bins = np.arange(0,43,1)
plt.xlim([-5, 48])
plt.hist(y_train, bins = bins)
plt.title("Train Class Histogram")
plt.xlabel("Class")
plt.ylabel("Count")
# +
#make a histogram of each class of sign
bins = np.arange(0,43,1)
plt.xlim([-5, 48])
plt.hist(y_valid, bins = bins)
plt.title("Valid Class Histogram")
plt.xlabel("Class")
plt.ylabel("Count")
# -
# ----
#
# ## Step 2: Design and Test a Model Architecture
#
# Design and implement a deep learning model that learns to recognize traffic signs. Train and test your model on the [German Traffic Sign Dataset](http://benchmark.ini.rub.de/?section=gtsrb&subsection=dataset).
#
# The LeNet-5 implementation shown in the [classroom](https://classroom.udacity.com/nanodegrees/nd013/parts/fbf77062-5703-404e-b60c-95b78b2f3f9e/modules/6df7ae49-c61c-4bb2-a23e-6527e69209ec/lessons/601ae704-1035-4287-8b11-e2c2716217ad/concepts/d4aca031-508f-4e0b-b493-e7b706120f81) at the end of the CNN lesson is a solid starting point. You'll have to change the number of classes and possibly the preprocessing, but aside from that it's plug and play!
#
# With the LeNet-5 solution from the lecture, you should expect a validation set accuracy of about 0.89. To meet specifications, the validation set accuracy will need to be at least 0.93. It is possible to get an even higher accuracy, but 0.93 is the minimum for a successful project submission.
#
# There are various aspects to consider when thinking about this problem:
#
# - Neural network architecture (is the network over or underfitting?)
# - Play around preprocessing techniques (normalization, rgb to grayscale, etc)
# - Number of examples per label (some have more than others).
# - Generate fake data.
#
# Here is an example of a [published baseline model on this problem](http://yann.lecun.com/exdb/publis/pdf/sermanet-ijcnn-11.pdf). It's not required to be familiar with the approach used in the paper but, it's good practice to try to read papers like these.
# ### Pre-process the Data Set (normalization, grayscale, etc.)
# Minimally, the image data should be normalized so that the data has mean zero and equal variance. For image data, `(pixel - 128)/ 128` is a quick way to approximately normalize the data and can be used in this project.
#
# Other pre-processing steps are optional. You can try different techniques to see if it improves performance.
#
# Use the code cell (or multiple code cells, if necessary) to implement the first step of your project.
# +
### Preprocess the data here. It is required to normalize the data. Other preprocessing steps could include
### converting to grayscale, etc.
### Feel free to use as many code cells as needed.
import cv2
#convert to grayscale
def convert2Gray(image):
gray_im = np.copy(image)
gray_im = cv2.cvtColor(gray_im, cv2.COLOR_RGB2GRAY)
# https://medium.com/computer-car/intricacies-of-traffic-sign-classification-with-tensorflow-8f994b1c8ba
gray_im = cv2.equalizeHist(gray_im)
gray_im.shape = (32,32,-1)
return gray_im
#https://www.pyimagesearch.com/2017/01/02/rotate-images-correctly-with-opencv-and-python/
#used this as a template but did not implement the 'correctly' part (translation commpensation)
def rotate_bound(image, angle):
# grab the dimensions of the image and then determine the
# center
(h, w) = image.shape[:2]
(cX, cY) = (w // 2, h // 2)
# grab the rotation matrix (applying the negative of the
# angle to rotate clockwise), then grab the sine and cosine
# (i.e., the rotation components of the matrix)
M = cv2.getRotationMatrix2D((cX, cY), -angle, 1.0)
cos = np.abs(M[0, 0])
sin = np.abs(M[0, 1])
# compute the new bounding dimensions of the image
nW = 32#int((h * sin) + (w * cos))
nH = 32#int((h * cos) + (w * sin))
# adjust the rotation matrix to take into account translation
#M[0, 2] += (nW / 2) - cX
#M[1, 2] += (nH / 2) - cY
# perform the actual rotation and return the image
return cv2.warpAffine(image, M, (nW, nH))
# +
#Add rotation augmentation
#this part of the code runs long, so the data are saved in a pickle file for further model development
use_gray = 1
use_augment = 1
bins = np.arange(0,43,1)
hist_val, hist_val_ind = np.histogram(y_train,bins)
#find the categories with low image count
if not use_pre_rotate:
low_category = []
for ii in range(len(hist_val)):
if hist_val[ii]<500:#500 is arbitrary
low_category.append(hist_val_ind[ii])
#append augmented images
if use_augment:
augment_images = X_train[0]
augment_images.shape = (1,32,32,3)
augment_labels = y_train[0]
for ii in range(len(X_train)):
if(y_train[ii] in low_category):
#Do two random rotations
#rotate the image
#append to augment
temp_image = rotate_bound(X_train[ii], np.random.randint(-6,6))
temp_image.shape = (1,32,32,3)
augment_images = np.append(augment_images,temp_image,0)
augment_labels = np.append(augment_labels,y_train[ii])
#rotate the image one more time
#append to augment
temp_image = rotate_bound(X_train[ii], np.random.randint(-6,6))
temp_image.shape = (1,32,32,3)
augment_images = np.append(augment_images,temp_image,0)
augment_labels = np.append(augment_labels,y_train[ii])
# -
#Plot a few of the traffic signs in a 3x3 output. randomize to get an idea of various signs
if not use_pre_rotate:
f, (ax1, ax2, ax3) = plt.subplots(1, 3)
ax1.imshow(augment_images[np.random.randint(0,len(augment_images))])
ax2.imshow(augment_images[np.random.randint(0,len(augment_images))])
ax3.imshow(augment_images[np.random.randint(0,len(augment_images))])
#Append the augmented parts to the original arrays for training
if not use_pre_rotate:
X_train = np.append(X_train,augment_images,0)
y_train = np.append(y_train,augment_labels,0)
#Save the data into a pickle file for further model development
if not use_pre_rotate:
data = {
'Training': X_train,
'Labels': y_train
}
with open('data.p', 'wb') as f:
# Pickle the 'data' dictionary using the highest protocol available.
pickle.dump(data, f, pickle.HIGHEST_PROTOCOL)
# +
#Normalize and grayscale
X_train_normal = []
for ii in range(len(y_train)):
if use_gray:
X_train_normal.append(convert2Gray(X_train[ii]))
X_train_normal[ii] = (X_train_normal[ii]-128.0)/128.0
else:
X_train_normal.append((np.array(X_train[ii])-128.0)/128.0)
h_train_image, w__train_image, layer_train_image = X_train_normal[0].shape
# -
#Normalize and grayscale
X_valid_normal = []
for ii in range(len(y_valid)):
if use_gray:
X_valid_normal.append(convert2Gray(X_valid[ii]))
X_valid_normal[ii] = (X_valid_normal[ii]-128.0)/128.0
else:
X_valid_normal.append((np.array(X_valid[ii])-128.0)/128.0)
#Normalize and grayscale
X_test_normal = []
for ii in range(len(y_test)):
if use_gray:
X_test_normal.append(convert2Gray(X_test[ii]))
X_test_normal[ii] = (X_test_normal[ii]-128.0)/128.0
else:
X_test_normal.append((np.array(X_test[ii])-128.0)/128.0)
#Plot the new histogram
plt.xlim([-5, 48])
plt.hist(y_train, bins = bins)
plt.title("Train Class Histogram")
plt.xlabel("Class")
plt.ylabel("Count")
# +
#Plot the sequence used for augmenting 'underrepresented image'
image_num = np.random.randint(0,n_train)
f, (ax1, ax2, ax3,ax4) = plt.subplots(1, 4)
ax1.imshow(X_train[image_num])
ax2.imshow(cv2.cvtColor(X_train[image_num], cv2.COLOR_RGB2GRAY),cmap='gray')
ax3.imshow(cv2.equalizeHist(cv2.cvtColor(X_train[image_num], cv2.COLOR_RGB2GRAY)),cmap='gray')
ax4.imshow(rotate_bound(cv2.equalizeHist(cv2.cvtColor(X_train[image_num], cv2.COLOR_RGB2GRAY)), np.random.randint(-6,6)),cmap='gray')
# -
# ### Model Architecture
# +
### Define your architecture here.
### Feel free to use as many code cells as needed.
from sklearn.utils import shuffle
#More preprocess - randomize the order since we appended a bunch of images
X_train_normal, y_train = shuffle(X_train_normal, y_train)
# -
# ### Set up TensorFlow
# +
import tensorflow as tf
EPOCHS = 5
BATCH_SIZE = 128
from tensorflow.contrib.layers import flatten
# +
#Implement LeNet from the LeNet-Lab
def LeNet(x, keep_dropout1, keep_dropout2):
# Hyperparameters
mu = 0
sigma = 0.1
#Layer 1: Convolutional. Input = 32x32x1. Output = 28x28x6.
conv1_W = tf.Variable(tf.truncated_normal(shape=(5, 5, 1, 6), mean = mu, stddev = sigma))
conv1_b = tf.Variable(tf.zeros(6))
conv1 = tf.nn.conv2d(x, conv1_W, strides=[1, 1, 1, 1], padding='VALID') + conv1_b
#Activation.
conv1 = tf.nn.relu(conv1)
#Layer 2: Convolutional Input = 28x28x6. Output = 24x24x10
convNew1_W = tf.Variable(tf.truncated_normal(shape=(5, 5, 6, 10), mean = mu, stddev = sigma))
convNew1_b = tf.Variable(tf.zeros(10))
convNew1 = tf.nn.conv2d(conv1, convNew1_W, strides=[1, 1, 1, 1], padding='VALID') + convNew1_b
#Activation.
convNew1 = tf.nn.relu(convNew1)
#Pooling. Input = 24x24x10. Output = 12x12x10.
convNew1 = tf.nn.max_pool(convNew1, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='VALID')
#Layer 3: Convolutional. Output = 8x8x28.
conv2_W = tf.Variable(tf.truncated_normal(shape=(5, 5, 10, 28), mean = mu, stddev = sigma))
conv2_b = tf.Variable(tf.zeros(28))
conv2 = tf.nn.conv2d(convNew1, conv2_W, strides=[1, 1, 1, 1], padding='VALID') + conv2_b
#Activation.
conv2 = tf.nn.relu(conv2)
#Pooling. Input = 8x8x28. Output = 4x4x28.
conv2 = tf.nn.max_pool(conv2, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='VALID')
#Flatten. Input = 4x4x28. Output = 384.
fc0 = flatten(conv2)
#Layer 4: Fully Connected. Input = 448. Output = 240.
fc1_W = tf.Variable(tf.truncated_normal(shape=(448, 240), mean = mu, stddev = sigma))
fc1_b = tf.Variable(tf.zeros(240))
fc1 = tf.matmul(fc0, fc1_W) + fc1_b
#Activation.
fc1 = tf.nn.relu(fc1)
fc1 = tf.nn.dropout(fc1, keep_dropout1)
#Layer 5: Fully Connected. Input = 240. Output = 84.
fc2_W = tf.Variable(tf.truncated_normal(shape=(240, 84), mean = mu, stddev = sigma))
fc2_b = tf.Variable(tf.zeros(84))
fc2 = tf.matmul(fc1, fc2_W) + fc2_b
#Activation.
fc2 = tf.nn.relu(fc2)
fc2 = tf.nn.dropout(fc2, keep_dropout2)
#Layer 6: Fully Connected. Input = 84. Output = 43.
fc3_W = tf.Variable(tf.truncated_normal(shape=(84, n_classes), mean = mu, stddev = sigma))
fc3_b = tf.Variable(tf.zeros(n_classes))
logits = tf.matmul(fc2, fc3_W) + fc3_b
return logits
# -
# ### Features and Labels
# +
# placeholder for a batch of input images
x = tf.placeholder(tf.float32, (None, h_image, w_image, layer_train_image))
# placeholder for a batch of output labels
y = tf.placeholder(tf.int32, (None))
#placeholder for keep value on dropout
keep_dropout1 = tf.placeholder(tf.float32)
keep_dropout2 = tf.placeholder(tf.float32)
#one hot encoding
one_hot_y = tf.one_hot(y, n_classes)
# -
# ### Train, Validate and Test the Model
# A validation set can be used to assess how well the model is performing. A low accuracy on the training and validation
# sets imply underfitting. A high accuracy on the training set but low accuracy on the validation set implies overfitting.
# +
### Train your model here.
### Calculate and report the accuracy on the training and validation set.
### Once a final model architecture is selected,
### the accuracy on the test set should be calculated and reported as well.
### Feel free to use as many code cells as needed.
# Training Pipeline
rate = 0.001
logits = LeNet(x,keep_dropout1, keep_dropout2)
cross_entropy = tf.nn.softmax_cross_entropy_with_logits(labels = one_hot_y, logits = logits)
loss_operation = tf.reduce_mean(cross_entropy)
optimizer = tf.train.AdamOptimizer(learning_rate = rate)
training_operation = optimizer.minimize(loss_operation)
# -
# ### Model Evaluation
# +
correct_prediction = tf.equal(tf.argmax(logits, 1), tf.argmax(one_hot_y, 1)) #find where the logits and the one-hot are matching
accuracy_operation = tf.reduce_mean(tf.cast(correct_prediction, tf.float32)) #sum all the correct predictions / length of correct predictions
saver = tf.train.Saver()
def evaluate(X_data, y_data):
num_examples = len(X_data)
total_accuracy = 0
sess = tf.get_default_session()
for offset in range(0, num_examples, BATCH_SIZE):
batch_x, batch_y = X_data[offset:offset+BATCH_SIZE], y_data[offset:offset+BATCH_SIZE]
accuracy = sess.run(accuracy_operation, feed_dict = {x: batch_x, y: batch_y, keep_dropout1: 1.0, keep_dropout2: 1.0})
total_accuracy += (accuracy * len(batch_x))
return total_accuracy / num_examples
# -
# ### Train the Model
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
num_examples = len(X_train_normal)
print("Training...")
print()
validation_accuracy = []
training_accuracy = []
for i in range(EPOCHS):
X_train_normal, y_train = shuffle(X_train_normal, y_train)
for offset in range(0, num_examples, BATCH_SIZE):
end = offset+BATCH_SIZE
batch_x, batch_y = X_train_normal[offset:end], y_train[offset:end]
sess.run(training_operation, feed_dict={x: batch_x, y: batch_y, keep_dropout1: 0.5, keep_dropout2: 0.5})
validation_accuracy.append(evaluate(X_valid_normal, y_valid))
training_accuracy.append(evaluate(X_train_normal, y_train))
print("EPOCH {} ...".format(i+1))
print("Validation Accuracy = {:.3f}".format(validation_accuracy[i]))
print("Training Accuracy = {:.3f}".format(training_accuracy[i]))
print()
saver.save(sess, './lenet')
print("Model saved")
# ---
#
# ## Step 3: Test a Model on New Images
#
# To give yourself more insight into how your model is working, download at least five pictures of German traffic signs from the web and use your model to predict the traffic sign type.
#
# You may find `signnames.csv` useful as it contains mappings from the class id (integer) to the actual sign name.
# ### Load and Output the Images
# +
### Load the images and plot them here.
### Feel free to use as many code cells as needed.
from PIL import Image
test_image = []
test_image.append(Image.open('family_crossing.jpg'))
test_image.append(Image.open('right_turn.jpg'))
test_image.append(Image.open('speed_30.jpg'))
test_image.append(Image.open('stop.jpg'))
test_image.append(Image.open('truck.jpg'))
f, (ax1, ax2, ax3, ax4, ax5) = plt.subplots(1, 5)
ax1.imshow(test_image[0])
ax2.imshow(test_image[1])
ax3.imshow(test_image[2])
ax4.imshow(test_image[3])
ax5.imshow(test_image[4])
# -
# crop the images and plot them to check
from PIL import Image
test_image[0] = test_image[0].crop((70,0,250,250-70))
test_image[1] = test_image[1].crop((50,0,230,230-50))
test_image[2] = test_image[2].crop((50,0,220,220-50))
test_image[3] = test_image[3].crop((70,0,260,260-70))
test_image[4] = test_image[4].crop((0 ,0,220,220-0 ))
f, (ax1, ax2, ax3, ax4, ax5) = plt.subplots(1, 5)
ax1.imshow(test_image[0])
ax2.imshow(test_image[1])
ax3.imshow(test_image[2])
ax4.imshow(test_image[3])
ax5.imshow(test_image[4])
for ii in range(5):
test_image[ii] = test_image[ii].resize((32,32))
f, (ax1, ax2, ax3, ax4, ax5) = plt.subplots(1, 5)
ax1.imshow(test_image[0])
ax2.imshow(test_image[1])
ax3.imshow(test_image[2])
ax4.imshow(test_image[3])
ax5.imshow(test_image[4])
#process the 5 images like the test and validation images
X_test5_normal = []
for ii in range(5):
if use_gray:
X_test5_normal.append(convert2Gray(test_image[ii]))
X_test5_normal[ii] = (X_test5_normal[ii]-128.0)/128.0
else:
X_test5_normal.append((np.array(test_image[ii])-128.0)/128.0)
# +
#plot the preprocessed image
#plt.imshow(X_test5_normal[4])
# -
#define the labels for the web images
y_test5 = []
y_test5.append(28)
y_test5.append(33)
y_test5.append(1)
y_test5.append(14)
y_test5.append(16)
# ### Predict the Sign Type for Each Image
# +
### Run the predictions here and use the model to output the prediction for each image.
### Make sure to pre-process the images with the same pre-processing pipeline used earlier.
### Feel free to use as many code cells as needed.
prediction_operation = tf.argmax(logits,1)
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
#for ii in range(5):
prediction = sess.run(prediction_operation , feed_dict={x: X_test5_normal, y: y_test5, keep_dropout1: 1.0, keep_dropout2: 1.0})
logit_out = sess.run(tf.nn.softmax(logits) , feed_dict={x: X_test5_normal, y: y_test5, keep_dropout1: 1.0, keep_dropout2: 1.0})
top_5_prob = sess.run(tf.nn.top_k(tf.nn.softmax(logits),k = 5), feed_dict={x: X_test5_normal, y: y_test5, keep_dropout1: 1.0, keep_dropout2: 1.0})
#print(prediction)
for ii in range(5):
print("Predicted Label = {:d}".format(prediction[ii]))
print("Actual Label = {:d}".format(y_test5[ii]))
print()
# -
# ### Analyze Performance
# +
### Calculate the accuracy for these 5 new images.
### For example, if the model predicted 1 out of 5 signs correctly, it's 20% accurate on these new images.
prediction_operation = tf.argmax(logits,1)
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
print("Accuracy for the five images = {:.1f}%".format(evaluate(X_test5_normal, y_test5)*100))
# -
# ### Output Top 5 Softmax Probabilities For Each Image Found on the Web
# For each of the new images, print out the model's softmax probabilities to show the **certainty** of the model's predictions (limit the output to the top 5 probabilities for each image). [`tf.nn.top_k`](https://www.tensorflow.org/versions/r0.12/api_docs/python/nn.html#top_k) could prove helpful here.
#
# The example below demonstrates how tf.nn.top_k can be used to find the top k predictions for each image.
#
# `tf.nn.top_k` will return the values and indices (class ids) of the top k predictions. So if k=3, for each sign, it'll return the 3 largest probabilities (out of a possible 43) and the correspoding class ids.
#
# Take this numpy array as an example. The values in the array represent predictions. The array contains softmax probabilities for five candidate images with six possible classes. `tf.nn.top_k` is used to choose the three classes with the highest probability:
#
# ```
# # (5, 6) array
# a = np.array([[ 0.24879643, 0.07032244, 0.12641572, 0.34763842, 0.07893497,
# 0.12789202],
# [ 0.28086119, 0.27569815, 0.08594638, 0.0178669 , 0.18063401,
# 0.15899337],
# [ 0.26076848, 0.23664738, 0.08020603, 0.07001922, 0.1134371 ,
# 0.23892179],
# [ 0.11943333, 0.29198961, 0.02605103, 0.26234032, 0.1351348 ,
# 0.16505091],
# [ 0.09561176, 0.34396535, 0.0643941 , 0.16240774, 0.24206137,
# 0.09155967]])
# ```
#
# Running it through `sess.run(tf.nn.top_k(tf.constant(a), k=3))` produces:
#
# ```
# TopKV2(values=array([[ 0.34763842, 0.24879643, 0.12789202],
# [ 0.28086119, 0.27569815, 0.18063401],
# [ 0.26076848, 0.23892179, 0.23664738],
# [ 0.29198961, 0.26234032, 0.16505091],
# [ 0.34396535, 0.24206137, 0.16240774]]), indices=array([[3, 0, 5],
# [0, 1, 4],
# [0, 5, 1],
# [1, 3, 5],
# [1, 4, 3]], dtype=int32))
# ```
#
# Looking just at the first row we get `[ 0.34763842, 0.24879643, 0.12789202]`, you can confirm these are the 3 largest probabilities in `a`. You'll also notice `[3, 0, 5]` are the corresponding indices.
# +
### Print out the top five softmax probabilities for the predictions on the German traffic sign images found on the web.
### Feel free to use as many code cells as needed.
#print(top_5_prob.values)
softmax_probs = []
for ii in range(5):
print("Top Softmax Probability for image number {:d}".format(ii+1))
print(top_5_prob.values[ii][:6])
softmax_probs.append(top_5_prob.values[ii][0])
print()
objects = ('Children Xing', 'Turn Right', '30 km/hr', 'Stop', '3.5 Tons')
y_pos = np.arange(len(objects))
plt.bar(y_pos, softmax_probs, align='center', alpha=0.5)
plt.xticks(y_pos, objects)
plt.ylabel('Probability')
plt.title('SoftMax Probabilities')
plt.show()
# +
### Calculate the accuracy for these 5 new images.
### For example, if the model predicted 1 out of 5 signs correctly, it's 20% accurate on these new images.
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
print("Testing...")
print()
print("Test Accuracy = {:.3f}".format(evaluate(X_test_normal, y_test)))
# -
#Determine the predictions
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
prediction = sess.run(prediction_operation , feed_dict={x: X_test_normal, y: y_test, keep_dropout1: 1.0, keep_dropout2: 1.0})
# +
#Count the number of predictions and actual
bins = np.arange(0,44,1)
hist_label, hist_label_ind = np.histogram(y_test,bins)
hist_test, hist_test_ind = np.histogram(prediction,bins)
# -
#Plot a histogram of labels from test set
plt.xlim([-5, 48])
plt.hist(y_test, bins = bins)
plt.title("Test Class Histogram")
plt.xlabel("Class")
plt.ylabel("Count")
print(bins)
#Plot a histogram of labels from predition set (should look similar to above)
plt.xlim([-5, 48])
plt.hist(prediction, bins = bins)
plt.title("Test Class Histogram")
plt.xlabel("Class")
plt.ylabel("Count")
# +
#Determine the number of true positives
true_positives = []
for ii in range(len(prediction)):
if prediction[ii]==y_test[ii]:
true_positives.append(y_test[ii])
else:
true_positives.append(-1)
hist_tp, hist_tpl_ind = np.histogram(true_positives,bins)
plt.xlim([-5, 48])
plt.hist(true_positives, bins = bins)
plt.title("True Positives Histogram")
plt.xlabel("Class")
plt.ylabel("Count")
# -
#Calculate the Precision
precision_test = []
recall_test = []
for ii in range(len(hist_tp)):
precision_test.append(hist_tp[ii]/hist_test[ii])
recall_test.append(hist_tp[ii]/hist_label[ii])
#Plot the precision
plt.figure(figsize=(12,3))
plt.bar(np.arange(0,43,1), precision_test, align='center', alpha=0.5)
plt.xticks(bins[:len(bins)-1], bins[:len(bins)-1])
plt.ylabel('Precision')
plt.title('Model Precision')
#Plot the recall
plt.figure(figsize=(12,3))
plt.bar(np.arange(0,43,1), recall_test, align='center', alpha=0.5)
plt.xticks(bins[:len(bins)-1], bins)
plt.ylabel('Recall')
plt.title('Model Recall')
# + active=""
# ### Project Writeup
#
# Once you have completed the code implementation, document your results in a project writeup using this [template](https://github.com/udacity/CarND-Traffic-Sign-Classifier-Project/blob/master/writeup_template.md) as a guide. The writeup can be in a markdown or pdf file.
# -
# > **Note**: Once you have completed all of the code implementations and successfully answered each question above, you may finalize your work by exporting the iPython Notebook as an HTML document. You can do this by using the menu above and navigating to \n",
# "**File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
# ---
#
# ## Step 4 (Optional): Visualize the Neural Network's State with Test Images
#
# This Section is not required to complete but acts as an additional excersise for understaning the output of a neural network's weights. While neural networks can be a great learning device they are often referred to as a black box. We can understand what the weights of a neural network look like better by plotting their feature maps. After successfully training your neural network you can see what it's feature maps look like by plotting the output of the network's weight layers in response to a test stimuli image. From these plotted feature maps, it's possible to see what characteristics of an image the network finds interesting. For a sign, maybe the inner network feature maps react with high activation to the sign's boundary outline or to the contrast in the sign's painted symbol.
#
# Provided for you below is the function code that allows you to get the visualization output of any tensorflow weight layer you want. The inputs to the function should be a stimuli image, one used during training or a new one you provided, and then the tensorflow variable name that represents the layer's state during the training process, for instance if you wanted to see what the [LeNet lab's](https://classroom.udacity.com/nanodegrees/nd013/parts/fbf77062-5703-404e-b60c-95b78b2f3f9e/modules/6df7ae49-c61c-4bb2-a23e-6527e69209ec/lessons/601ae704-1035-4287-8b11-e2c2716217ad/concepts/d4aca031-508f-4e0b-b493-e7b706120f81) feature maps looked like for it's second convolutional layer you could enter conv2 as the tf_activation variable.
#
# For an example of what feature map outputs look like, check out NVIDIA's results in their paper [End-to-End Deep Learning for Self-Driving Cars](https://devblogs.nvidia.com/parallelforall/deep-learning-self-driving-cars/) in the section Visualization of internal CNN State. NVIDIA was able to show that their network's inner weights had high activations to road boundary lines by comparing feature maps from an image with a clear path to one without. Try experimenting with a similar test to show that your trained network's weights are looking for interesting features, whether it's looking at differences in feature maps from images with or without a sign, or even what feature maps look like in a trained network vs a completely untrained one on the same sign image.
#
# <figure>
# <img src="visualize_cnn.png" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> Your output should look something like this (above)</p>
# </figcaption>
# </figure>
# <p></p>
#
# +
### Visualize your network's feature maps here.
### Feel free to use as many code cells as needed.
# image_input: the test image being fed into the network to produce the feature maps
# tf_activation: should be a tf variable name used during your training procedure that represents the calculated state of a specific weight layer
# activation_min/max: can be used to view the activation contrast in more detail, by default matplot sets min and max to the actual min and max values of the output
# plt_num: used to plot out multiple different weight feature map sets on the same block, just extend the plt number for each new feature map entry
def outputFeatureMap(image_input, tf_activation, activation_min=-1, activation_max=-1 ,plt_num=1):
# Here make sure to preprocess your image_input in a way your network expects
# with size, normalization, ect if needed
# image_input =
# Note: x should be the same name as your network's tensorflow data placeholder variable
# If you get an error tf_activation is not defined it may be having trouble accessing the variable from inside a function
activation = tf_activation.eval(session=sess,feed_dict={x : image_input})
featuremaps = activation.shape[3]
plt.figure(plt_num, figsize=(15,15))
for featuremap in range(featuremaps):
plt.subplot(6,8, featuremap+1) # sets the number of feature maps to show on each row and column
plt.title('FeatureMap ' + str(featuremap)) # displays the feature map number
if activation_min != -1 & activation_max != -1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmin =activation_min, vmax=activation_max, cmap="gray")
elif activation_max != -1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmax=activation_max, cmap="gray")
elif activation_min !=-1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmin=activation_min, cmap="gray")
else:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", cmap="gray")
# -
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
#print([tensor.name for tensor in tf.get_default_graph().as_graph_def().node])
conv2 = tf.get_default_graph().get_tensor_by_name("Conv2D_2:0")
outputFeatureMap(X_test_normal,conv2)
| Traffic_Sign_Classifier.ipynb |