code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import keras
from keras.layers import Dense, Embedding, LSTM, Input
import matplotlib.pyplot as plt
from keras.models import Model
from sklearn.model_selection import train_test_split
import seaborn as sns
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelEncoder
from keras.models import Model
from keras.layers import LSTM, Activation, Dense, Dropout, Input, Embedding
from keras.optimizers import RMSprop
from keras.preprocessing.text import Tokenizer
from keras.preprocessing import sequence
from keras.utils import to_categorical
from keras.callbacks import EarlyStopping
from keras.preprocessing.sequence import pad_sequences
import gensim
import pickle as pk
from io import StringIO
from scipy import spatial
from sklearn.feature_selection import chi2
from sklearn.preprocessing import LabelBinarizer
from keras.models import Model, Sequential
from keras.layers import LSTM,Dense,Input,Bidirectional, Embedding, Dropout
from keras import regularizers
from nltk.tokenize.treebank import TreebankWordTokenizer
from scipy import spatial
from random import shuffle
import pickle as pk
from keras.callbacks import ModelCheckpoint
from keras.preprocessing.text import one_hot
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.model_selection import train_test_split
from nltk.corpus import stopwords
df = pd.concat([pd.read_csv('fp.csv'), pd.read_csv('bc.csv'), pd.read_csv('chr.csv')])
len(df)
df.drop(columns=['Unnamed: 0','polarity'], inplace=True)
df = df[df['factuality'] != 'NOT_LABELED']
df.head()
len(df)
X_train, X_test, Y_train, Y_test = train_test_split(df.text, df.factuality, test_size = 0.15)
bn = LabelBinarizer()
bn.fit(Y_train)
Y_train_bin = bn.transform(Y_train)
Y_test_bin = bn.transform(Y_test)
Y_train_bin
tokenizer = Tokenizer()
tokenizer.fit_on_texts(df.text)
word_index = tokenizer.word_index
m = 0
for i in df.text.str.split():
m = max(m, len(i))
m
sequences_data = tokenizer.texts_to_sequences(X_train)
x_train = pad_sequences(sequences_data, maxlen= 100)
sequences_data_test = tokenizer.texts_to_sequences(X_test)
x_test = pad_sequences(sequences_data_test, maxlen= 100)
word2vec = gensim.models.KeyedVectors.load_word2vec_format('/Users/amin/Downloads/w2v_phrase3_npmi.bin')
word2vec.most_similar('sexy')
EMBEDDING_DIM = 300
MAX_SEQUENCE_LENGTH = 100
N_CLASSES = 3
embedding_dim = 300
# +
from keras.layers import Embedding
nb_words = len(word_index)+1
embedding_matrix = np.random.rand(nb_words, EMBEDDING_DIM)
for word, i in word_index.items():
if word in word2vec.vocab:
embedding_matrix[i] = word2vec.word_vec(word)
print('Null word embeddings: %d' % np.sum(np.sum(embedding_matrix, axis=1) == 0))
embedding_layer = Embedding(embedding_matrix.shape[0], # or len(word_index) + 1
embedding_matrix.shape[1], # or EMBEDDING_DIM,
weights=[embedding_matrix],
input_length=MAX_SEQUENCE_LENGTH,
trainable=False)
# -
embedding_matrix.shape
# +
from keras.layers import Dense, Input, Flatten, LSTM, Dropout, Bidirectional
from keras.layers import GlobalAveragePooling1D, Embedding , Reshape,Concatenate
from keras.models import Model
from keras.layers.embeddings import Embedding
from keras.layers.convolutional import Conv2D,MaxPooling2D
from keras.callbacks import ModelCheckpoint
from keras.optimizers import Adam
#filter_sizes = [2,3,4,5,6]
drop = 0.2
sequence_input = Input(shape=(MAX_SEQUENCE_LENGTH,), dtype='int32')
embedded_sequences = embedding_layer(sequence_input)
reshape = Reshape((MAX_SEQUENCE_LENGTH,EMBEDDING_DIM))(embedded_sequences)
lstm_0 = Bidirectional(LSTM(activation='tanh',units=40, return_sequences=True))(reshape)
lstm_1 = Bidirectional(LSTM(activation='tanh',units=40, return_sequences=True))(lstm_0)
lstm_2 = Bidirectional(LSTM(activation='tanh',units=40, dropout=drop))(lstm_1)
dense_0 = Dense(10,activation='tanh')(lstm_2)
out = Dense(N_CLASSES,activation='softmax')(dense_0)
model = Model(inputs=sequence_input, outputs=out)
#adam = Adam(lr=1e-4, beta_1=0.9, beta_2=0.999, epsilon=1e-08, decay=0.0)
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
model.summary()
# -
fp = './weights/factuality/lstm_bidir/weights_lstm_bidir-{epoch:02d}-{val_acc:.4f}.hdf'
checkpoint = ModelCheckpoint(fp, monitor='val_acc', verbose=1, save_best_only=True, mode='auto')
model.fit(x_train, Y_train_bin, validation_data=[x_test,Y_test_bin], nb_epoch=10, batch_size=16, callbacks=[checkpoint])
# +
from sklearn.metrics import classification_report
model.load_weights('./weights/factuality/lstm_bidir/weights_lstm_bidir-04-0.7591.hdf')
y_pred = model.predict(x_test)
output_test = np.argmax(y_pred, axis=1)
print(classification_report(np.argmax(Y_test_bin, axis=1), output_test))
# +
from keras.layers import Dense, Input, Flatten, LSTM, Dropout, Bidirectional
from keras.layers import GlobalAveragePooling1D, Embedding , Reshape,Concatenate
from keras.models import Model
from keras.layers.embeddings import Embedding
from keras.layers.convolutional import Conv2D,MaxPooling2D,Conv1D,MaxPooling1D
from keras.callbacks import ModelCheckpoint
from keras.optimizers import Adam
#filter_sizes = [2,3,4,5,6]
drop = 0.5
num_filters = 32
sequence_input = Input(shape=(MAX_SEQUENCE_LENGTH,), dtype='int32')
embedded_sequences = embedding_layer(sequence_input)
reshape = Reshape((MAX_SEQUENCE_LENGTH,EMBEDDING_DIM))(embedded_sequences)
conv_0 = Conv1D(num_filters, kernel_size=3, padding='valid', kernel_initializer='normal', activation='relu')(reshape)
maxpool_0 = MaxPooling1D(pool_size=2)(conv_0)
#print(maxpool_0.shape)
#reshape = Reshape((MAX_SEQUENCE_LENGTH,EMBEDDING_DIM))(embedded_sequences)
lstm_0 = Bidirectional(LSTM(activation='tanh',units=40, return_sequences=True))(maxpool_0)
lstm_1 = Bidirectional(LSTM(activation='tanh',units=40, return_sequences=True))(lstm_0)
lstm_2 = Bidirectional(LSTM(activation='tanh',units=40))(lstm_1)
dropout = Dropout(drop)(lstm_2)
dense_0 = Dense(20,activation='tanh')(dropout)
dense_1 = Dense(10,activation='tanh')(dense_0)
out = Dense(N_CLASSES,activation='softmax')(dense_0)
model_cnn_lstm = Model(inputs=sequence_input, outputs=out)
#adam = Adam(lr=1e-4, beta_1=0.9, beta_2=0.999, epsilon=1e-08, decay=0.0)
model_cnn_lstm.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
model_cnn_lstm.summary()
# -
fp = './weights/factuality/cnn_lstm/weights_cnn_lstm-{epoch:02d}-{val_acc:.4f}.hdf'
checkpoint = ModelCheckpoint(fp, monitor='val_acc', verbose=1, save_best_only=True, mode='auto')
model_cnn_lstm.load_weights('./weights/factuality/cnn_lstm/weights_cnn_lstm-01-0.7510.hdf')
model_cnn_lstm.fit(x_train, Y_train_bin, validation_data=[x_test,Y_test_bin], nb_epoch=10, batch_size=16, callbacks=[checkpoint])
# +
model_cnn_lstm.load_weights('./weights/factuality/cnn_lstm/weights_cnn_lstm-01-0.7510.hdf')
y_pred = model_cnn_lstm.predict(x_test)
output_test = np.argmax(y_pred, axis=1)
print(classification_report(np.argmax(Y_test_bin, axis=1), output_test))
# +
from keras.layers import Dense, Input, Flatten
from keras.layers import GlobalAveragePooling1D, Embedding , Reshape,Concatenate
from keras.models import Model
from keras.layers.embeddings import Embedding
from keras.layers.convolutional import Conv2D,MaxPooling2D
from keras.callbacks import ModelCheckpoint
from keras.optimizers import Adam
num_filters = 32
filter_sizes = [2,3,4,5]
drop = 0.4
sequence_input = Input(shape=(MAX_SEQUENCE_LENGTH,), dtype='int32')
embedded_sequences = embedding_layer(sequence_input)
# print(embedded_sequences.shape)
reshape = Reshape((MAX_SEQUENCE_LENGTH,EMBEDDING_DIM,1))(embedded_sequences)
# print(reshape.shape)
conv_0 = Conv2D(num_filters, kernel_size=(filter_sizes[0], embedding_dim), padding='valid', kernel_initializer='normal', activation='relu')(reshape)
conv_1 = Conv2D(num_filters, kernel_size=(filter_sizes[1], embedding_dim), padding='valid', kernel_initializer='normal', activation='relu')(reshape)
conv_2 = Conv2D(num_filters, kernel_size=(filter_sizes[2], embedding_dim), padding='valid', kernel_initializer='normal', activation='relu')(reshape)
conv_3 = Conv2D(num_filters, kernel_size=(filter_sizes[3], embedding_dim), padding='valid', kernel_initializer='normal', activation='relu')(reshape)
maxpool_0 = MaxPooling2D(pool_size=(MAX_SEQUENCE_LENGTH - filter_sizes[0] + 1, 1), strides=(1,1), padding='valid')(conv_0)
maxpool_1 = MaxPooling2D(pool_size=(MAX_SEQUENCE_LENGTH - filter_sizes[1] + 1, 1), strides=(1,1), padding='valid')(conv_1)
maxpool_2 = MaxPooling2D(pool_size=(MAX_SEQUENCE_LENGTH - filter_sizes[2] + 1, 1), strides=(1,1), padding='valid')(conv_2)
maxpool_3 = MaxPooling2D(pool_size=(MAX_SEQUENCE_LENGTH - filter_sizes[3] + 1, 1), strides=(1,1), padding='valid')(conv_3)
concatenated_tensor = Concatenate(axis=1)([maxpool_0, maxpool_1, maxpool_2,maxpool_3])
flatten = Flatten()(concatenated_tensor)
dropout = Dropout(drop)(flatten)
output = Dense(units= N_CLASSES, activation='softmax')(dropout)
# this creates a model that includes
model_cnn = Model(inputs=sequence_input, outputs=output)
checkpoint = ModelCheckpoint('weights_cnn_sentece.hdf5', monitor='val_acc', verbose=1, save_best_only=True, mode='auto')
adam = Adam(lr=1e-4, beta_1=0.9, beta_2=0.999, epsilon=1e-08, decay=0.0)
model_cnn.compile(optimizer=adam, loss='categorical_crossentropy', metrics=['accuracy'])
model_cnn.summary()
# -
fp = './weights/factuality/cnn/weights_cnn-{epoch:02d}-{val_acc:.4f}.hdf'
checkpoint = ModelCheckpoint(fp, monitor='val_acc', verbose=1, save_best_only=True, mode='auto')
model_cnn.fit(x_train, Y_train_bin, validation_data=[x_test,Y_test_bin], nb_epoch=3, batch_size=16, callbacks=[checkpoint])
# +
from sklearn.metrics import classification_report
model_cnn.load_weights('./weights/factuality/cnn/weights_cnn-03-0.7470.hdf')
y_pred = model_cnn.predict(x_test)
output_test = np.argmax(y_pred, axis=1)
print(classification_report(np.argmax(Y_test_bin, axis=1), output_test))
# -
| .ipynb_checkpoints/factuality-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import local_models.local_models
import numpy as np
import scipy.stats
import matplotlib.pyplot as plt
from importlib import reload
from ml_battery.utils import cmap
import matplotlib as mpl
import logging
import ml_battery.log
import time
import os
import functools
import collections
import itertools
import pymongo
import datetime
import pytz
import re
import local_models.loggin
import local_models.tf_w2v_models
import pickle
from nltk.tokenize import TweetTokenizer
logger = logging.getLogger(__name__)
np.random.seed(1)
#reload(local_models.loggin)
#reload(local_models.TLS_models)
np.warnings.filterwarnings('ignore')
# -
mpl.rcParams['figure.figsize'] = [8.0, 8.0]
RUN = 1
BANDWIDTH = 0.35
SEED = 1
np.random.seed(SEED)
KERNEL=local_models.local_models.GaussianKernel
root_dir = "/home/scott/local_w2v/"
project_dir = os.path.join(root_dir, "r{:03d}_k{}".format(RUN, KERNEL(bandwidth=BANDWIDTH)))
data_dir = os.path.join(root_dir, "data")
os.makedirs(project_dir, exist_ok=1)
# +
def remove_by_regex(text, regexp):
return regexp.sub("", text)
url_re = re.compile(r"http.?://[^\s]+[\s]?")
def remove_urls(text):
return remove_by_regex(text, url_re)
specialchar_re = re.compile("[" + ''.join(map(re.escape,
[",", ":", "\"", "=", "&", ";", "%", "$", "@", "%", "^", "*", "(", ")", "{", "}",
"[", "]", "|", "/", "\\", ">", "<", "-", "!", "?", ".", "'", "--", "---"])) + "]")
def remove_special_chars(text):
return remove_by_regex(text, specialchar_re)
username_re = re.compile(r"@[^\s]+[\s]?")
def remove_usernames(text):
return remove_by_regex(text, username_re)
number_re = re.compile(r"\s?[0-9]+\.?[0-9]*")
def remove_numbers(text):
return remove_by_regex(text, number_re)
def lower(text):
return text.lower()
cleanup = (remove_urls,
remove_usernames,
remove_special_chars,
remove_numbers,
lower)
# -
def get_cikm_tweet_column(line):
try:
return line.split("\t")[-2]
except Exception as e:
return None
def count(iterable, item):
tot = 0
for i in iterable:
if i == item:
tot += 1
return tot
pre_training_data = os.path.join(data_dir, "training_set_tweets.txt")
tknzr = TweetTokenizer(strip_handles=True, reduce_len=True)
# + active=""
# tweets = []
# with open(pre_training_data) as f:
# for line in f:
# try:
# tweet = get_cikm_tweet_column(line)
# for cleaner in cleanup:
# tweet = cleaner(tweet)
# tweets.append(tweet)
# except:
# continue
#
# print(len(tweets), count(tweets, None))
# tweets = [i for i in tweets if i is not None]
# print(len(tweets))
# toknized_tweets = [tknzr.tokenize(t) for t in tweets]
# -
class fuckit(object):
def __init__(self, message=None):
self.message = message
def __enter__(self): return self
def __exit__(self, *args):
if self.message is not None:
print(self.message)
return True
with fuckit("nothing to del"):
TFGlobalSesh.close()
with fuckit("nothing to del"):
del TFGlobalSesh
with fuckit("nothing to del"):
with TFGlobalGraph.as_default():
tf.reset_default_graph()
with fuckit("nothing to del"):
del TFGlobalGraph
with fuckit("nothing to del"):
del model
VOCAB_SIZE = 6000
BASE_MODEL_EPOCHS = 10000
BASE_MODEL_RUN = 2
try:
with open(os.path.join(root_dir, "base_model_v{:08d}_e{:08d}_r{:03d}.model".format(VOCAB_SIZE, BASE_MODEL_EPOCHS, BASE_MODEL_RUN)), 'rb') as f:
pmodel = pickle.load(f)
except FileNotFoundError as e:
pmodel = Word2Vec(vocabulary_size=VOCAB_SIZE, epochs=BASE_MODEL_EPOCHS, log_epochs=0.01)
pmodel.fit(toknized_tweets, sample_weight=np.ones(len(toknized_tweets)))
import pickle
with open(os.path.join(root_dir, "base_model_v{:08d}_e{:08d}_r{:03d}.model".format(VOCAB_SIZE, BASE_MODEL_EPOCHS, BASE_MODEL_RUN)), 'wb') as f:
pickle.dump(model, f)
pmodel.sort("sweet")
pmodel
pmodel.epochs = 10
pmodel.batch_size=1
rando_data = local_models.tf_w2v_models.build_dataset_predictionary(
["the quick brown fox jumps over the lazy dog".split(" ")],
pmodel.dictionary, pmodel.reverse_dictionary)
rando_data
pmodel.fit(rando_data, sample_weight=np.array([1]))
| examples/local_w2v_basemodel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <br>
# <font face="Helvetica" size="5"><b>VegMapper</b></font>
# <br><br>
# <font face="Helvetica" size="3">License Terms
#
# Copyright (c) 2019, California Institute of Technology ("Caltech"). U.S. Government sponsorship acknowledged.
#
# All rights reserved.
#
# Redistribution and use in source and binary forms, with or without modification, are permitted provided that the following conditions are met:
#
# * Redistributions of source code must retain the above copyright notice, this list of conditions and the following disclaimer.
# * Redistributions must reproduce the above copyright notice, this list of conditions and the following disclaimer in the documentation and/or other materials provided with the distribution.
# * Neither the name of Caltech nor its operating division, the Jet Propulsion Laboratory, nor the names of its contributors may be used to endorse or promote products derived from this software without specific prior written permission.
#
# THIS SOFTWARE IS PROVIDED BY THE COPYRIGHT HOLDERS AND CONTRIBUTORS "AS IS" AND ANY EXPRESS OR IMPLIED WARRANTIES, INCLUDING, BUT NOT LIMITED TO, THE IMPLIED WARRANTIES OF MERCHANTABILITY AND FITNESS FOR A PARTICULAR PURPOSE ARE DISCLAIMED. IN NO EVENT SHALL THE COPYRIGHT OWNER OR CONTRIBUTORS BE LIABLE FOR ANY DIRECT, INDIRECT, INCIDENTAL, SPECIAL, EXEMPLARY, OR CONSEQUENTIAL DAMAGES (INCLUDING, BUT NOT LIMITED TO, PROCUREMENT OF SUBSTITUTE GOODS OR SERVICES; LOSS OF USE, DATA, OR PROFITS; OR BUSINESS INTERRUPTION) HOWEVER CAUSED AND ON ANY THEORY OF LIABILITY, WHETHER IN CONTRACT, STRICT LIABILITY, OR TORT (INCLUDING NEGLIGENCE OR OTHERWISE) ARISING IN ANY WAY OUT OF THE USE OF THIS SOFTWARE, EVEN IF ADVISED OF THE POSSIBILITY OF SUCH DAMAGE.
#
# </font>
# <br>
#load in Python libraries
import ipywidgets as ipw
from ipyfilechooser import FileChooser
from IPython.display import Markdown, display
def printmd(string):
display(Markdown(string))
#set up Python/R dual functionality for notebook
# %load_ext rpy2.ipython
# + language="R"
# #initialize R cell and load in necessary libraries
# library(rgdal)
# library(arm)
# library(gdalUtils)
# library(raster)
# -
# <br><font face="Helvetica" size="3">User inputs:</font>
# +
print("\n")
##### FILES #####
printmd('<font face="Helvetica" size="4"><b>FILE INPUTS:<b>')
#all GIS data in lat lon WGS84 unless specified
#path to comma-delimited file, must have cols 'latitude', 'longitude', 'class'
fc_in_points = FileChooser()
fc_in_points.use_dir_icons = True
fc_in_points.title = '<font face="Helvetica" size="2"><b>Please select the validation points (csv file):</b>'
display(fc_in_points)
#remote sensing stack in ENVI flat binary format, get stack info
fc_in_stack = FileChooser()
fc_in_stack.use_dir_icons = True
fc_in_stack.title = '<font face="Helvetica" size="2"><b>Please select the remote sensing stack (NOT the .hdr file):</b>'
display(fc_in_stack)
#path to output comma-delimited file, same as in_points with appended remote sensing values
printmd('<font face="Helvetica" size="2"><b>Name of output validation csv appended remote sensing values (please include .csv):<b>')
printmd('<font face="Helvetica" size="2">NOTE: there is no need to press enter to confirm the name')
name_out_points = ipw.Text(
value='',
placeholder='Type something',
description='File name:',
disabled=False
)
display(name_out_points)
#name of the output map in ENVI flat binary format
printmd('<font face="Helvetica" size="2"><b>Name of output map in ENVI flat binary format:<b>')
printmd('<font face="Helvetica" size="2">NOTE: there is no need to press enter to confirm the name')
name_out_pred = ipw.Text(
value='',
placeholder='Type something',
description='File name:',
disabled=False
)
display(name_out_pred)
#name of output GeoTIFF
printmd('<font face="Helvetica" size="2"><b>Name of output GeoTIFF (please include .tif):<b>')
printmd('<font face="Helvetica" size="2">NOTE: there is no need to press enter to confirm the name')
name_out_tif = ipw.Text(
value='',
placeholder='Type something',
description='File name:',
disabled=False
)
display(name_out_tif)
#--------------------------------------------------------------#
print("\n")
##### PARAMETERS #####
printmd('<font face="Helvetica" size="4"><b>PARAMETER INPUTS:<b>')
printmd('<font face="Helvetica" size="2">NOTE: when probability is larger than threshold, we say that oil palm is present')
#buffer
value_buffer = ipw.BoundedIntText(
value=1,
min=0,
max=5,
step=1,
description='# of cells:',
disabled=False
)
#threshold
value_threshold = ipw.FloatSlider(
value=0.5,
min=0,
max=1.0,
step=0.05,
description='Float:',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='.1f',
)
#prior
status_use_prior = ipw.Checkbox(
value=False,
description='Use prior in model?',
disabled=False
)
accordion = ipw.Accordion(children=[value_buffer, value_threshold, status_use_prior])
accordion.set_title(0, 'Buffer size')
accordion.set_title(1, 'Threshold for logistic model')
accordion.set_title(2, 'Prior selection')
display(accordion)
# -
# <br><font face="Helvetica" size="3">Set inputs as variables:</font>
# +
in_points = fc_in_points.selected_filename
in_stack = fc_in_stack.selected_filename
out_points = name_out_points.value
out_pred = name_out_pred.value
out_tif = name_out_tif.value
buffer = value_buffer.value
threshold = value_threshold.value
use_prior = status_use_prior.value
print('User file inputs: ','\n', in_points, '\n', in_stack, '\n', out_points, '\n', out_pred, '\n', out_tif, '\n', buffer, '\n', threshold, '\n', use_prior)
#EDIT: could add exception handling here, other user input sections
# -
# <br><font face="Helvetica" size="3">Push file inputs to R:</font>
# %Rpush in_points in_stack out_points out_pred out_tif buffer threshold use_prior
# + language="R"
# #test push success
# push_file_variables = list(in_points, in_stack, out_points, out_pred, out_tif, buffer, threshold, use_prior)
# print(push_file_variables)
# -
# <br><font face="Helvetica" size="3">TEMP: OTHER USER INPUTS W/O WIDGET:</font>
# + language="R"
# ##### PARAMETERS #####
# hhBandIndex = 0 #NOTE: see Apply routine for information
# #RUN: priors (variable order corresponds to order in which bands are read), currently Costa Rica
# prior_mean = c(0.06491638, -26.63132179, 0.05590800, -29.64091620)
# prior_scale = c(0.02038204, 7.58200324, 0.01686930, 8.73995422)
# prior_mean_int = 1.99274801
# prior_scale_int = 7.22600112
#
# #lower and upper bounds for posterior credible intervals
# lp = 0.025
# up = 0.975
#
# #--------------------------------------------------------------#
#
# ##### BAND INFO #####
# stack_names = c("vcf", "c_rvi", "ndvi", "l_rvi_mosaic") #desired bands from the in_stack (names should match source)
# #NOTE: this is a DEVELOPER input and not a user input,
# # please do not change this unless you have been approved to do so
# bands = c(2,3,1,4) #index of each of the bands defined above (from in_stack)
# nodata = -9999 #NA value present in input bands: NEEDS TO BE A LIST OF NUMBERS IF NOT CONSISTENT ACROSS BANDS
# -
# <hr><br>
# <font face="Helvetica" size="5"><b>EXTRACT</b></font>
# <br><br>
# <font face="Helvetica" size="3">Objective: read remote sensing values at training points. The cell below creates a function to get intensity values from stack and executes extract routine:</font>
# + language="R"
# getPixel= function(gdalObject, X, Y, buffer, ulX, ulY, cellSize, bands){
# nrow = dim(gdalObject)[1]
# ncol = dim(gdalObject)[2]
# rowOffset = ((ulY-Y)/cellSize) - buffer
# if(rowOffset<0 | (rowOffset+buffer+2) > nrow){
# return(NA)
# }
# colOffset = ((X-ulX)/cellSize) - buffer
# if(colOffset<0 | (colOffset+buffer+2) > ncol){
# return(NA)
# }
# windowY = buffer+2
# windowX = windowY
# pixelValue = getRasterData(gdalObject, band=bands, offset=c(rowOffset, colOffset), region.dim=c(windowY, windowX))
# return(pixelValue)
# }
#
# #stack information
# r_stack = stack(in_stack)
# res = xres(r_stack)
# r_extent = r_stack@extent
# ulX = r_extent@xmin
# ulY = r_extent@ymax
# # s_info = GDALinfo(in_stack) also works, lacks ulY
#
# #grab stack
# gdalObj = new("GDALDataset", in_stack)
#
# #append remote sensing information to point table and write to cvs file
# inData <- read.csv(in_points, header=TRUE)
# numPoints <- nrow(inData)
#
# header <- c(colnames(inData), stack_names)
# write.table(x=t(header), file=out_points, append=FALSE, col.names=FALSE, row.names=FALSE, sep=",")
#
# print("Extracting values for...")
# for(i in 1:numPoints){
# allBands <- rep(NA, length(bands))
# for (j in 1:length(bands)){
# oneBand = getPixel(gdalObj, inData$longitude[i], inData$latitude[i], buffer, ulX, ulY, res, bands[j])
# w = which (oneBand == nodata[j])
# oneBand[w]<-NA
# allBands[j] = mean(oneBand, na.rm=TRUE)
# if(i==1) print(stack_names[j])
# }
# mydata <- data.frame(t(as.vector(allBands)))
# colnames(mydata) <- stack_names
# newRow = cbind(inData[i,],mydata)
# write.table(x=newRow, file=out_points, append=TRUE, col.names=FALSE, row.names=FALSE, sep=",")
# }
# -
# <hr><br>
# <font face="Helvetica" size="5"><b>RUN</b></font>
# <br><br>
# <font face="Helvetica" size="3">Objective: fit Bayesian model, calculate posteriors and confusion matrix. The cell below creates the model and executes the prediction, constructs a confusion matrix, calculates the prediction accuracy/posterior CI, calculates/builds posteriors for subsequent runs, and prints the result:</font>
# + language="R"
# #ADDITIONAL INPUTS
# #columns of the predictor variables to be used in this model (taken from pred csv)
# #column order indicated here is vcf, c_rvi, ndvi, l_rvi_mosaic, matching the priors above
# index = c(13,14,15,16)
# + language="R"
# #impute missing values by variable means
# data = read.csv(out_points)
# for (i in which(sapply(data, is.numeric))) {
# for (j in which(is.na(data[, i]))) {
# data[j, i] <- mean(data[data[, "my_class"] == data[j, "my_class"], i], na.rm = TRUE)
# }
# }
#
# #true_label: 1 for oil_palm and 0 for non oil_palm
# true_label = 1*(data$my_class == 'oil_palm') #EDIT: why this --> http://127.0.0.1:54098/notebooks/sample-run/example_analysis.ipynb
#
# #transform interested variables into a matrix which would be used
# x = as.matrix(data[, index])
# all_names = names(data)
# stack_names = all_names[index]
# colnames(x) = stack_names
#
# #build model by incorporating those variables
# formula = as.formula(paste("true_label ~ ", paste(stack_names, collapse="+"),sep = ""))
# use_data = as.data.frame(cbind(x, true_label))
#
# #to specify prior
# #if noninformative prior, use prior.mean=apply(x, 2, mean), prior.scale=Inf, prior.df=Inf
# #if having a prior, set prior.mean=c(....), prior.scale=c(.....)
# #length of prior mean and prior scale should be equal to the number of predictors
# if(! use_prior){
# model = bayesglm(formula, data=use_data, family=binomial(link='logit'), prior.mean=apply(x, 2, mean), prior.scale=Inf, scale=FALSE)
# }
# if(use_prior){
# model = bayesglm(formula, data=use_data, family=binomial(link='logit'),
# prior.mean=prior_mean,
# prior.scale=prior_scale,
# prior.mean.for.intercept=prior_mean_int,
# prior.scale.for.intercept=prior_scale_int,
# scale = FALSE)
# }
#
# #oil_palm prediction
# class_prediction = 1*(model$fitted.values >= threshold) #if the fitted value is above the threshold, value is changed to binary 1
# print(class_prediction)
#
# #used instead of na.remove to get rid of NA values in 2018 validation dataset
# true_label = true_label[!is.na(true_label)]
#
# #generate confusion matrix
# bayesian_conf_matrix = matrix(0,2,2)
# bayesian_conf_matrix[1,1] = sum(class_prediction + true_label == 0)
# bayesian_conf_matrix[2,2] = sum(class_prediction + true_label == 2)
# bayesian_conf_matrix[1,2] = sum((class_prediction == 0) & (true_label == 1))
# bayesian_conf_matrix[2,1] = sum((class_prediction == 1) & (true_label == 0))
# rownames(bayesian_conf_matrix) = c("Predicted non-oil-palm", "Predicted oil-palm")
# colnames(bayesian_conf_matrix) = c("Actual non-oil-palm", "Actual oil-palm")
# print(bayesian_conf_matrix)
#
# #overall accuracy of model
# accu_bayes = sum(class_prediction == true_label) / nrow(data)
# print("Overall accuracy:")
# print(accu_bayes)
#
# #EDIT: push values to Python and use numpy/matplotlib to display matrix
#
# # approach posterior distributions of coefficients
# # specify number of draws
# num_draw = 2000
# post_dist = sim(model, n.sims=num_draw)
# coef_matrix = coef(post_dist)
#
# # calculate posterior credible intervals for coefficients
# posterior_ci_coef = matrix(NA, ncol(x)+1, 2)
# for (i in 1:(ncol(x)+1)){
# posterior_ci_coef[i, ] = unname(quantile(coef_matrix[, i], probs=c(lp, up), na.rm=TRUE))
# }
#
# # calculate posterior credible intervals for every data point
# posterior_ci_data = matrix(NA, nrow(x), 2)
# for(i in 1:nrow(x)){
# temp = as.numeric()
# for(j in 1:num_draw){
# temp[j] = 1 / (1 + exp(-coef_matrix[j, 1] - sum(coef_matrix[j, 2:(length(index)+1)] * x[i, ])))
# }
# posterior_ci_data[i, ] = unname(quantile(temp, probs=c(lp, up), na.rm=TRUE))
# }
#
# # build posterior objects for next run
# posterior_mean = model$coefficients
# posterior_scale = apply(coef(post_dist), 2, sd)
#
# #print the posteriors, store them for later
# #EDIT: not sure if this works with combinations of other stack variables than current build (need to test in future)
# posterior_mean
# intercept = posterior_mean[["(Intercept)"]]
# numVars = length(posterior_mean)
# posteriors = posterior_mean[2:numVars] #EDIT: does this need to be transformed into c() variable?
# -
# <hr><br>
# <font face="Helvetica" size="5"><b>APPLY</b></font>
# <br><br>
# <font face="Helvetica" size="3">Objective: apply model fits to calculate OP3 for the area covered by the data stack. OP3 = oil palm probability presence, ranging between 0-1. The cell below applies the model to the stack and executes the predictive analysis, outputting the prediction surface in GeoTIFF and ENVI binary formats:</font>
# + language="R"
# #generate dummy for Docker
# #only use env variables if creation fails (will normally return NULL but still create object)
# # Sys.setenv(PROJ_LIB="/usr/bin/proj/")
# # Sys.getenv("PROJ_LIB")
# #dummy corresponds to one band from the original stack, used to save your output prediction map
# in_dummy = "temp_dummy"
# gdal_translate(src_dataset=in_stack, dst_dataset=in_dummy, of="ENVI", b=1)
#
# #create GIS objects
# gdalObjStack = new("GDALDataset", in_stack)
# gdalObjDummy = new("GDALDataset", in_dummy)
# rasterWidth = ncol(gdalObjStack)
# rasterRows = nrow(gdalObjStack)
#
# #calculate prediction for each pixel and save
# print("Checking a few values...")
# for(i in 1:rasterRows){
# oneRasterLine = getRasterData(gdalObjStack, offset=c(i-1,0), region.dim=c(1, rasterWidth))
# hhBand = hhBandIndex #PREVIOUSLY: which(bandNames == "alos2_hh")
# #NOTE: previous value was 0, bandNames/modelBands was removed for redundancy,
# # the above code has not been tested yet
# pred = rep(-9999, rasterWidth)
# for(j in 1:rasterWidth){
# #hh = (20*log10(oneRasterLine[j, 1, hhBand])) -83
# hh = oneRasterLine[j, 1, hhBand] #gets hh value at each of the pixels
# #open water mask
# #if(is.na(hh) | hh < -20){
# #pred[j] = 0
# #}
# #else{
# #select bands
# selectBands = oneRasterLine[j, 1, bands] #EDITED: changed modelBands to bands
# z = (intercept + sum(posteriors * selectBands))
# pred[j] = exp(z)/(1+ exp(z))
# #z = (intercept + sum(posteriors * scaledBands))
# #pred[j] = exp(z)/(1+ exp(z))
# if ((i/100 == i%/%100) & j == 1000) print(z) #reality check on the model fits
# #}
# }
# #write one row to file
# putRasterData(gdalObjDummy, pred, offset=c(i-1, 0)) #place predicted line in raster into dummy
# }
# saveDataset(gdalObjDummy, out_pred)
#
# #convert to GeoTiff
# gdal_translate(src_dataset=out_pred, dst_dataset=out_tif, of="GTiff")
| sample-run/example_analysis_WIDGETS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
import sys
sys.path.append("../")
# -
from korr import bootcorr, pearson
import numpy as np
# # Load Dataset
from sklearn.datasets import load_diabetes
X = load_diabetes().data
# # Correlation Estimate
cmat, _ = pearson(X)
print(cmat.round(2))
# # Bootstrap Estimation
r3, _, _ = bootcorr(X, n_draws=30, subsample=0.7, replace=True, corr_fn=pearson)
cmat_bmean = r3.mean(axis=0)
print(cmat_bmean.round(2))
# How stable are estimates
cmat_bstd = r3.std(axis=0)
print(cmat_bstd.round(2))
# How different is the Bootstrap Estimation from the Full-Sample Estimation
print(np.abs(cmat_bmean - cmat).round(2))
| examples/bootcorr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import random
import csv
import math
import numpy as np
import time
import operator
import matplotlib.pyplot as plt
import cPickle as pickle
import heapq
# # Load Stored Live Edges
# +
live_edges_200k_point001_point3 = pickle.load(open("live_edges_200k_point001_point3", "rb"))
print "ni hao"
live_edges_200k_point001_point5 = pickle.load(open("live_edges_200k_point001_point5", "rb"))
print "hello"
live_edges_200k_point005_point3 = pickle.load(open("live_edges_200k_point005_point3", "rb"))
print "hi"
live_edges_200k_point005_point5 = pickle.load(open("live_edges_200k_point005_point5", "rb"))
# -
# # CELF
def celf_population_live_edges(p_initial_infect, p_infect, lists_infected_nodes, num_detectors, seed=0, n=2029, debug=False):
runs_per_marginal_node = len(lists_infected_nodes)
print 'Running CELF to minimize population affected w/ live edges'
results = open("CELF_population_live_edge.txt", "w")
results.write("P_infect = " + str(p_infect) + " P_initial_infect = " + str(p_initial_infect) + " Runs per marginal node = " + str(runs_per_marginal_node))
results.write("\nNode ID\tReward so Far\tMarginal gain\tRuntime (minutes)\n")
final_population = []
final_runtimes = []
rewards_so_far = sum( [ len(infected_ids) for infected_ids in lists_infected_nodes ]) / float(runs_per_marginal_node) # population affected w/o detectors
marginals = [] # heap to track marginals
start_time = time.time()
# we are calculated population saved
# if the detector is in the infected nodes, we take the index of the detector b/c the infected nodes are in chronological order + 1 - length to represent how many we saved
# if not, we just take 0 b/c we didnt save any
# select the first node by exhuastively computing all marginal benefits
# our metric = (population affected w/ detector) - population affected w/o detector
# we are trying to minimize this metric
for potential_detector in range(1,n+1): # calculate marginals for each node
runs = [infected_ids.index(potential_detector)+1 if potential_detector in infected_ids else len(infected_ids) for infected_ids in lists_infected_nodes]
heapq.heappush(marginals, (sum(runs)/float(runs_per_marginal_node) - rewards_so_far, potential_detector) )
if debug: # gonna pop a couple
top_5 = []
for i in range(5):
top_5.append( heapq.heappop(marginals) )
for marginal in top_5:
print marginal
for i in range(5):
heapq.heappush(marginals, top_5[i])
best_tuple = heapq.heappop(marginals)
best_node = best_tuple[1]
marginal = best_tuple[0]
rewards_so_far = marginal + rewards_so_far
selected_detectors = [best_node] # we have found our first best node
print "We have our first best node: "
print selected_detectors
runtime = (time.time() - start_time) / 60
print "Runtime (minutes) = " + str( runtime )
final_population.append(rewards_so_far)
final_runtimes.append(runtime)
results.write(str(best_node) + '\t' + str(rewards_so_far) + '\t' + str(marginal) + '\t' + str(runtime) + '\n' )
for i in range(1, num_detectors): # select the next 19 detectors
if debug:
print "Trying to find the " + str(i+1) + "th detector"
# need to re-evaluate the top node once, at least
old_top_tuple = heapq.heappop(marginals)
old_top_node = old_top_tuple[1]
new_detectors = list(selected_detectors)
new_detectors.append(old_top_node)
# lazily evaluate for the top node
runs = [] # records the results for each run
for run in range(runs_per_marginal_node): # go thru the runs
infected_ids = lists_infected_nodes[run]
if bool(set(new_detectors) & set(infected_ids)): # detector is in the set of infected ids
for infected_node in infected_ids: # let's go thru the infected IDs to see which one is affected
if infected_node in new_detectors:
runs.append(infected_ids.index(infected_node) + 1)
break # since we found the earliest detection
else: # outbreak not detected, so everyone got affected
runs.append(len(infected_ids))
heapq.heappush(marginals, (sum(runs) / float(runs_per_marginal_node) - rewards_so_far, old_top_node))
if debug: # gonna pop a couple
top_5 = []
for i in range(5):
top_5.append( heapq.heappop(marginals) )
for marginal in top_5:
print marginal
for i in range(5):
heapq.heappush(marginals, top_5[i])
new_top_tuple = heapq.heappop(marginals)
new_top_node = new_top_tuple[1]
while new_top_node != old_top_node: # this is where we need to re-evaluate the next top node
old_top_node = new_top_node
new_detectors = list(selected_detectors)
new_detectors.append(old_top_node)
runs = [] # records number of runs
for run in range(runs_per_marginal_node): # go thru the runs
infected_ids = lists_infected_nodes[run]
if bool(set(new_detectors) & set(infected_ids)): # detector is in the set of infected ids
for infected_node in infected_ids: # let's go thru the infected IDs to see which one is affected
if infected_node in new_detectors:
runs.append(infected_ids.index(infected_node) + 1)
break
else: # outbreak not detected, so everyone got affected
runs.append(len(infected_ids))
heapq.heappush(marginals, (sum(runs) / float(runs_per_marginal_node) - rewards_so_far, old_top_node))
new_top_tuple = heapq.heappop(marginals)
new_top_node = new_top_tuple[1]
best_node = new_top_node
print "After doing CELF, we have found the " + str (i+1) + "th detector w/ best marginal gain = " + str(best_node)
selected_detectors.append(best_node)
runtime = (time.time() - start_time)/60
rewards_so_far = rewards_so_far + new_top_tuple[0] # update the rewards w/ the marginals gained by the current node
results.write(str(best_node) + '\t' + str(rewards_so_far) + '\t' + str(new_top_tuple[0]) + '\t' + str(runtime) + '\n')
final_population.append(rewards_so_far)
final_runtimes.append(runtime)
print "Population affected so far: " + str(rewards_so_far)
print "Runtime (minutes) = " + str(runtime)
results.close()
return final_population, final_runtimes, selected_detectors
# # Pure Social Score
# +
def social_population_live_edges(p_initial_infect, p_infect, lists_infected_nodes, num_detectors):
final_population = []
final_runtimes = []
with open("socialScore_modified_unpruned.txt") as social_file:
lines = social_file.readlines()
social_score_nodes = []
for line in lines:
social_score_nodes.append( int(line.split()[0]) )
runs_per_marginal_node = len(lists_infected_nodes)
rewards_so_far = 0
marginal = 0
start_time = time.time()
selected_detectors = []
for next_detector in range(num_detectors):
selected_detectors.append(social_score_nodes[next_detector]) # add the next detector
runs = [] # records number of runs
for run in range(runs_per_marginal_node): # go thru the runs
infected_ids = lists_infected_nodes[run]
if bool(set(selected_detectors) & set(infected_ids)): # detector is in the set of infected ids
for infected_node in infected_ids: # let's go thru the infected IDs to see which one is affected
if infected_node in selected_detectors:
runs.append(infected_ids.index(infected_node) + 1)
break
else: # outbreak not detected, so everyone got affected
runs.append(len(infected_ids))
population = sum(runs) / float(runs_per_marginal_node)
marginal = population - rewards_so_far
rewards_so_far = population
final_population.append(rewards_so_far)
print "After doing Social Score, we have found the " + str(next_detector+1) + "th detector = " + str(social_score_nodes[next_detector])
print "Population affected so far: " + str(rewards_so_far)
print "Marginal gain = " + str(marginal)
runtime = (time.time() - start_time) / 60
print "Runtime (minutes) = " + str(runtime)
final_runtimes.append(runtime)
return final_population, final_runtimes
# -
# # Speed Up
def speedup_population_live_edges(p_initial_infect, p_infect, lists_infected_nodes, num_detectors, social_score_file, seed=0, n=2029, debug=False):
runs_per_marginal_node = len(lists_infected_nodes)
final_population = []
final_runtimes = []
print 'Running Speedup to minimize population affected w/ live edges'
results = open("Speedup_population_live_edge.txt", "w")
results.write("P_infect = " + str(p_infect) + " P_initial_infect = " + str(p_initial_infect) + " Runs per marginal node = " + str(runs_per_marginal_node))
results.write("\nNode ID\tReward so Far\tMarginal gain\tRuntime (minutes)\n")
rewards_so_far = sum( [ len(infected_ids) for infected_ids in lists_infected_nodes ]) / float(runs_per_marginal_node) # population affected w/o detectors
marginals = [] # heap to track marginals
# how we keep track of the social score nodes
social_score_nodes = []
with open(social_score_file) as social_file:
lines = social_file.readlines()
for line in lines:
social_score_nodes.append( int(line.split()[0]) )
look_ahead = int(math.sqrt(n)) # how much we look ahead
start_time = time.time()
# we are calculated population saved
# if the detector is in the infected nodes, we take the index of the detector b/c the infected nodes are in chronological order + 1 - length to represent how many we saved
# if not, we just take 0 b/c we didnt save any
# select the first node by exhuastively computing all marginal benefits
# our metric = (population affected w/ detector) - population affected w/o detector
# we are trying to minimize this metric
for potential_detector in social_score_nodes[:look_ahead]: # calculate marginals for each node
runs = [infected_ids.index(potential_detector)+1 if potential_detector in infected_ids else len(infected_ids) for infected_ids in lists_infected_nodes]
heapq.heappush(marginals, (sum(runs)/float(runs_per_marginal_node) - rewards_so_far, potential_detector) )
best_tuple = heapq.heappop(marginals)
best_node = best_tuple[1]
marginal = best_tuple[0]
rewards_so_far = marginal + rewards_so_far
selected_detectors = [best_node] # we have found our first best node
print "We have our first best node: "
print selected_detectors
runtime = (time.time() - start_time) / 60
print "Runtime (minutes) = " + str( runtime )
final_population.append(rewards_so_far)
final_runtimes.append(runtime)
results.write(str(best_node) + '\t' + str(rewards_so_far) + '\t' + str(marginal) + '\t' + str(runtime) + '\n' )
for i in range(1, num_detectors): # select the next num_detectors-1 detectors
if debug:
print "Trying to find the " + str(i+1) + "th detector"
# add one new detector from social score
new_detectors=list(selected_detectors)
additional_detector = social_score_nodes[look_ahead - 1 + i]
new_detectors.append(additional_detector)
runs = [] # records the results for each run
for run in range(runs_per_marginal_node): # go thru the runs
infected_ids = lists_infected_nodes[run]
if bool(set(new_detectors) & set(infected_ids)): # detector is in the set of infected ids
for infected_node in infected_ids: # let's go thru the infected IDs to see which one is affected
if infected_node in new_detectors:
runs.append(infected_ids.index(infected_node) + 1)
break # since we found the earliest detection
else: # outbreak not detected, so everyone got affected
runs.append(len(infected_ids))
heapq.heappush(marginals, (sum(runs) / float(runs_per_marginal_node) - rewards_so_far, additional_detector))
# need to re-evaluate the top node once, at least
old_top_tuple = heapq.heappop(marginals)
old_top_node = old_top_tuple[1]
new_detectors = list(selected_detectors)
new_detectors.append(old_top_node)
# lazily evaluate for the top node
runs = [] # records the results for each run
for run in range(runs_per_marginal_node): # go thru the runs
infected_ids = lists_infected_nodes[run]
if bool(set(new_detectors) & set(infected_ids)): # detector is in the set of infected ids
for infected_node in infected_ids: # let's go thru the infected IDs to see which one is affected
if infected_node in new_detectors:
runs.append(infected_ids.index(infected_node) + 1)
break # since we found the earliest detection
else: # outbreak not detected, so everyone got affected
runs.append(len(infected_ids))
heapq.heappush(marginals, (sum(runs) / float(runs_per_marginal_node) - rewards_so_far, old_top_node))
new_top_tuple = heapq.heappop(marginals)
new_top_node = new_top_tuple[1]
while new_top_node != old_top_node: # this is where we need to re-evaluate the next top node
old_top_node = new_top_node
new_detectors = list(selected_detectors)
new_detectors.append(old_top_node)
runs = [] # records number of runs
for run in range(runs_per_marginal_node): # go thru the runs
infected_ids = lists_infected_nodes[run]
if bool(set(new_detectors) & set(infected_ids)): # detector is in the set of infected ids
for infected_node in infected_ids: # let's go thru the infected IDs to see which one is affected
if infected_node in new_detectors:
runs.append(infected_ids.index(infected_node) + 1)
break
else: # outbreak not detected, so everyone got affected
runs.append(len(infected_ids))
heapq.heappush(marginals, (sum(runs) / float(runs_per_marginal_node) - rewards_so_far, old_top_node))
new_top_tuple = heapq.heappop(marginals)
new_top_node = new_top_tuple[1]
best_node = new_top_node
print "After doing speed up, we have found the " + str (i+1) + "th detector w/ best marginal gain = " + str(best_node)
selected_detectors.append(best_node)
runtime = (time.time() - start_time)/60
rewards_so_far = rewards_so_far + new_top_tuple[0] # update the rewards w/ the marginals gained by the current node
results.write(str(best_node) + '\t' + str(rewards_so_far) + '\t' + str(new_top_tuple[0]) + '\t' + str(runtime) + '\n')
final_population.append(rewards_so_far)
final_runtimes.append(runtime)
print "Population affected so far: " + str(rewards_so_far)
print "Runtime (minutes) = " + str(runtime)
results.close()
return final_population, final_runtimes, selected_detectors
# # Comparisons
# P_infect = 0.3, p_initial = 0.001
celf_population_point001_point3, celf_population_runtimes_point001_point3, selected_detectors = celf_population_live_edges(0.001, 0.3, live_edges_200k_point001_point3[:100000], 50, debug=False )
social_population_point001_point3, social_population_runtimes_point001_point3 = social_population_live_edges(0.001, 0.3, live_edges_200k_point001_point3[:100000], 50 )
speedup_population_point001_point3, speedup_population_runtimes_point001_point3, selected_detectors = speedup_population_live_edges(0.001, 0.3, live_edges_200k_point001_point3[:100000], 50 , "socialScore_modified_unpruned.txt" )
plt.plot(range(1, 51), celf_population_point001_point3, label="CELF", marker="o", color="red")
plt.plot(range(1, 51), social_population_point001_point3, label="Social Score only", marker="^")
plt.plot(range(1, 51), speedup_population_point001_point3, label="Speed up", marker=".", color="green")
plt.plot(range(1,51), np.ones(50)*2, label="Initial number of infected nodes")
plt.title("100K runs, P_infect = 0.3, P_initial_infect = 0.001")
plt.ylabel("Population Affected")
plt.xlabel("# of Detectors")
plt.legend()
plt.plot(range(1,51), celf_population_runtimes_point001_point3, label="CELF", color="red")
plt.plot(range(1,51), [time + 7 for time in social_population_runtimes_point001_point3], label="Social Score only", color="blue")
plt.plot(range(1,51), [time + 7 for time in speedup_population_runtimes_point001_point3] , label="Speed up", color="green")
plt.title("100K runs, P_infect = 0.3, P_initial_infect = 0.001")
plt.ylabel("Runtime (minutes)")
plt.xlabel("# of Detectors")
plt.legend()
# p_initial = 0.005, p_infect = 0.5
celf_population_point005_point5, celf_population_runtimes_point005_point5, selected_detectors = celf_population_live_edges(0.005, 0.5, live_edges_200k_point005_point5[:100000], 50, debug=False )
social_population_point005_point5, social_population_runtimes_point005_point5 = social_population_live_edges(0.005, 0.5, live_edges_200k_point005_point5[:100000], 50 )
speedup_population_point005_point5, speedup_population_runtimes_point005_point5, selected_detectors = speedup_population_live_edges(0.005, 0.5, live_edges_200k_point005_point5[:100000], 50, "socialScore_modified_unpruned.txt", debug=False )
plt.plot(range(1, 51), celf_population_point005_point5, label="CELF", color="red", marker="o")
plt.plot(range(1, 51), social_population_point005_point5, label="Social Score only", marker="^")
plt.plot(range(1, 51), speedup_population_point005_point5, label="Speed up", marker=".", color="green")
plt.plot(range(1,51), np.ones(50)*10, label="Initial Number of Infected Nodes")
plt.title("100K runs, P_infect = 0.5, P_initial_infect = 0.005")
plt.ylabel("Population Affected")
plt.xlabel("# of Detectors")
plt.legend()
# +
plt.plot(range(1,51), celf_population_runtimes_point005_point5, label="CELF", color="red")
plt.plot(range(1,51), [time + 7 for time in social_population_runtimes_point005_point5], label="Social Score only", color="blue")
plt.plot(range(1,51), [time + 7 for time in speedup_population_runtimes_point005_point5], label="Speed up", color="green")
plt.title("100K runs, P_infect = 0.5, P_initial_infect = 0.005")
plt.ylabel("Runtime (minutes)")
plt.xlabel("# of Detectors")
plt.legend()
# -
# p_initial = 0.005, p_infect = 0.3
celf_population_point005_point3, celf_population_runtimes_point005_point3, selected_detectors = celf_population_live_edges(0.005, 0.3, live_edges_200k_point005_point3[:10000], 40, debug=False )
social_population_point005_point3, social_population_runtimes_point005_point3 = social_population_live_edges(0.005, 0.3, live_edges_200k_point005_point3[:100000], 40 )
speedup_population_point005_point3, speedup_population_runtimes_point005_point3, selected_detectors = speedup_population_live_edges(0.005, 0.3, live_edges_200k_point005_point3[:100000], 40, "socialScore_modified_unpruned.txt", debug=False )
plt.plot(range(1, 41), celf_population_point005_point3, label="CELF", marker = "s", color="red")
plt.plot(range(1, 41), social_population_point005_point3, label="Social Score only", marker = "^")
plt.plot(range(1, 41), speedup_population_point005_point3, label="Speed up", marker="o", color="green")
plt.plot(range(1,41), np.ones(40)*10, label="Initial Number of Infected Nodes")
plt.title("100K runs, P_infect = 0.3, P_initial_infect = 0.005")
plt.ylabel("Population Affected")
plt.xlabel("# of Detectors")
plt.legend()
celf_10k_runtime = celf_population_runtimes_point005_point3
plt.plot(range(1,41), celf_10k_runtime, label="CELF 10K runs", color="red")
plt.plot(range(1,41), [runtime + 7 for runtime in social_population_runtimes_point005_point3], label="Social Score only 100K runs", color="blue")
plt.plot(range(1,41), [runtime + 7 for runtime in speedup_population_runtimes_point005_point3], label="Speed up 100K runs", color="green")
plt.title("Runtime comparison for minimizing population affected:\nP_infect = 0.3, P_initial_infect = 0.005")
plt.ylabel("Runtime (minutes)")
plt.xlabel("# of Detectors")
plt.legend()
social_population_runtimes_point005_point3
# # Saving celf output
pickle.dump(celf_population_point001_point3, open("celf_population_point001_point3", "wb"))
pickle.dump(celf_population_runtimes_point001_point3, open("celf_population_runtimes_point001_point3", "wb"))
pickle.dump(celf_population_point005_point5, open("celf_population_point005_point5", "wb"))
pickle.dump(celf_population_runtimes_point005_point5, open("celf_population_runtimes_point005_point5", "wb"))
pickle.dump(celf_population_point005_point3, open("celf_population_point005_point3", "wb"))
pickle.dump(celf_population_runtimes_point005_point3, open("celf_population_runtimes_point005_point3", "wb"))
| sim_outbreak/Final Project - Population Affected.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
matplotlib.style.use('ggplot')#使用ggplot样式
# %matplotlib inline
# # 区域图
# 可以使用Series.plot.area()和DataFrame.plot.area()创建区域图。
#
# 默认情况下,区域图堆叠。 为了产生堆积区域图,每列必须是正值或全部负值。
#
# 当输入数据包含NaN时,它会自动填满0。
#
# 如果要删除缺失值或填充其他值,请在调用plot之前使用dataframe.dropna()或dataframe.fillna()。
df = pd.DataFrame(np.random.rand(10, 4), columns=['a', 'b', 'c', 'd'])
df.plot.area()
# 注:随机产生标准正态分布的df,四栏分别绘制堆积区域图
# +
# 为了产生一个未堆积的图,通过使参数stacked=False。
# 除非另有规定,否则透明度:Alpha值设置为0.5,即半透明:
df.plot.area(stacked=False)
# -
# ## 散点图
# 可以使用DataFrame.plot.scatter()方法绘制散点图。
#
# 散点图需要x和y轴的数字列。 这些可以由x和y关键字指定。
df = pd.DataFrame(np.random.rand(50, 4), columns=['a', 'b', 'c', 'd'])
df.plot.scatter(x='a', y='b')
# 将产生的a栏作为x轴数据,b栏作为y轴数据绘图
ax = df.plot.scatter(x='a', y='b', color='DarkBlue', label='Group 1')
df.plot.scatter(x='c', y='d', color='RED', label='Group 2', ax=ax)
# 注:要在单个轴上绘制多个列组,要重复指定目标轴的绘图方法,
# 建议指定颜色和标签关键字来区分每个组。
df.plot.scatter(x='a', y='b', c='c', s=50)
# 注:关键字c可以作为列的名称给出,以为每个点提供颜色
# +
# 你可以传递由matplotlib散点支持的其他关键字。
# 下面的示例显示使用数据框列值作为气泡大小的气泡图:
df.plot.scatter(x='a', y='b', s=df['c']*200)
# 注:增加c栏作为气泡(散点)大小值
# -
| notebook/procs-pandas-plot-area.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="YBHHl0MvFdBG" colab_type="code" colab={}
# + id="rGVc1OlVFinL" colab_type="code" colab={}
# Run this cell to mount your Google Drive.
from google.colab import drive
drive.mount('/content/drive')
| Colab Notebooks-20190911T131716Z-001/Colab Notebooks/Filtering(1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="Za8-Nr5k11fh"
# ##### Copyright 2018 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="Eq10uEbw0E4l"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="Ou0PGp_4icRo"
# # Time windows
# + [markdown] colab_type="text" id="93b0GzKph0jK"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/examples/blob/master/courses/udacity_intro_to_tensorflow_for_deep_learning/l08c04_time_windows.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/examples/blob/master/courses/udacity_intro_to_tensorflow_for_deep_learning/l08c04_time_windows.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="vidayERjaO5q"
# ## Setup
# + colab={} colab_type="code" id="gqWabzlJ63nL"
import tensorflow as tf
# + [markdown] colab_type="text" id="ViWVB9qd8OIR"
# ## Time Windows
#
# First, we will train a model to forecast the next step given the previous 20 steps, therefore, we need to create a dataset of 20-step windows for training.
# + colab={} colab_type="code" id="bgJkwtq88OIS"
dataset = tf.data.Dataset.range(10)
for val in dataset:
print(val.numpy())
# + colab={} colab_type="code" id="ad8C65JV8OIT"
dataset = tf.data.Dataset.range(10)
dataset = dataset.window(5, shift=1)
for window_dataset in dataset:
for val in window_dataset:
print(val.numpy(), end=" ")
print()
# + colab={} colab_type="code" id="AQtmODsi8OIU"
dataset = tf.data.Dataset.range(10)
dataset = dataset.window(5, shift=1, drop_remainder=True)
for window_dataset in dataset:
for val in window_dataset:
print(val.numpy(), end=" ")
print()
# + colab={} colab_type="code" id="kTRHiWxi8OIW"
dataset = tf.data.Dataset.range(10)
dataset = dataset.window(5, shift=1, drop_remainder=True)
dataset = dataset.flat_map(lambda window: window.batch(5))
for window in dataset:
print(window.numpy())
# + colab={} colab_type="code" id="iPsQbWHb8OIX"
dataset = tf.data.Dataset.range(10)
dataset = dataset.window(5, shift=1, drop_remainder=True)
dataset = dataset.flat_map(lambda window: window.batch(5))
dataset = dataset.map(lambda window: (window[:-1], window[-1:]))
for x, y in dataset:
print(x.numpy(), y.numpy())
# + colab={} colab_type="code" id="hzp7RD6_8OIY"
dataset = tf.data.Dataset.range(10)
dataset = dataset.window(5, shift=1, drop_remainder=True)
dataset = dataset.flat_map(lambda window: window.batch(5))
dataset = dataset.map(lambda window: (window[:-1], window[-1:]))
dataset = dataset.shuffle(buffer_size=10)
for x, y in dataset:
print(x.numpy(), y.numpy())
# + colab={} colab_type="code" id="y70nV0EI8OIZ"
dataset = tf.data.Dataset.range(10)
dataset = dataset.window(5, shift=1, drop_remainder=True)
dataset = dataset.flat_map(lambda window: window.batch(5))
dataset = dataset.map(lambda window: (window[:-1], window[-1:]))
dataset = dataset.shuffle(buffer_size=10)
dataset = dataset.batch(2).prefetch(1)
for x, y in dataset:
print("x =", x.numpy())
print("y =", y.numpy())
# + colab={} colab_type="code" id="1tl-0BOKkEtk"
def window_dataset(series, window_size, batch_size=32,
shuffle_buffer=1000):
dataset = tf.data.Dataset.from_tensor_slices(series)
dataset = dataset.window(window_size + 1, shift=1, drop_remainder=True)
dataset = dataset.flat_map(lambda window: window.batch(window_size + 1))
dataset = dataset.shuffle(shuffle_buffer)
dataset = dataset.map(lambda window: (window[:-1], window[-1]))
dataset = dataset.batch(batch_size).prefetch(1)
return dataset
| courses/udacity_intro_to_tensorflow_for_deep_learning/l08c04_time_windows.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Login
# # Features
# + [markdown] cell_style="split"
# **Description:** Feature Creation.
# + cell_style="split" hide_input=true
from IPython.lib.display import YouTubeVideo
YouTubeVideo('FbL2eaErCcA')
# -
from prediction.apis import data_munging_engine
data_munging_engine.generate_features(auth, database, collection, featureset, categoryfield, datefield, numfield, groupby, find)
data_munging_engine.get_categories(auth, database, collection, categoryfield, find, total)
data_munging_engine.foreign_key_lookup(auth, database, collection, attribute, search, mongodbf, collectionf, attributef, fields)
data_munging_engine.foreign_key_aggregator(auth, database, collection, attribute, search, mongodbf, collectionf, attributef, fields)
data_munging_engine.csv_import(auth, database, collection, csv_file)
data_munging_engine.get_data(auth, database, collection, field, limit, projections, skip)
data_munging_engine.get_data_aggregate(auth, database, collection, field, projections, aggregate, sort)
data_munging_engine.enrich_fragments(auth, database, collection, attribute, strings)
data_munging_engine.enrich_fragments2(auth, database, collection, attribute, strings, find)
data_munging_engine.enrich_mcc(auth, database, collection, attribute, find)
data_munging_engine.concat_columns(auth, databasename, collection, attribute)
data_munging_engine.concat_columns2(auth, database, collection, attribute, separator)
data_munging_engine.enrich_date(auth, database, collection, attribute)
data_munging_engine.enum_convert(auth, database, collection, attribute)
data_munging_engine.enrich_predictor(auth, database, collection, search, sort, predictor, predictor_label, attributes, skip, limit)
data_munging_engine.export_documents(auth, filename, filetype, database, collection, field, sort, projection, limit)
data_munging_engine.process_basket(auth, dbcust, colcust, searchcust, custfield, dbitem, colitem, itemfield, supportcount)
data_munging_engine.process_directed_graph(auth, graphMeta, graphParam)
| .ipynb_checkpoints/02_Feature Store Preparation-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import seaborn as sns
import os
cwd=os.getcwd()
import sys
os.chdir("../scripts")
import project_functions
os.chdir(cwd)
project_functions.RawDir()
# -
# Our original data set is given below
dfo=pd.read_csv('medical_expenses.csv')
project_functions.returnDir(cwd)
dfo
# This data set is describing expected medical costs in America based on other variables, it is from Machine Learning with R by <NAME>. This dataset is pretty clean but I will do some wrangling to it so it can be better used for analysis. From this dataset I would like to be able to answer how unhealthy life styles such as under or over average bmi and smoking effect medical charges of an individual and to what extent these factors play a roll in expected charges of an individual
# **Data Wrangling**
# ---
# ---
# First I would like to add some columns to this dataset and change some of the variable names to better suit the variable it is representing
df=project_functions.load_and_process(cwd)
df
# Here i dropped the children and the region column as they wont be used in further analysis so they are redundant and I rounded the bmi amd charges to values that are more commonly used.
project_functions.mean(df)
# Taking a mean of the charges column returns a value of 13270.42 which we used to determine if the patient is given excessive charges which means they pay out more than expected on average. Next i want to seperate the data sets into healthy and not healthy populations to see how excessive charges changes throughout the population
dfh = project_functions.Health(cwd)
dfun =project_functions.unHealth(cwd)
dfh
# Here is the data set for healthy people, the requirements for this was to be in healthy bmi range of 18.5-24.9 and to not be a smoker.
dfun
# Here is the data set for unhealthy people the requirments are the inverse of the healthy data set thus they have to be outside the healthy bmi range or be classified as a smoker. We can first look at some regression plots of age vs charges to see how having a unhealthy lifestyle effects the charges
# **Regression Plots**
# ---
# ---
project_functions.plotAvC(df)
# Regression plot for whole population
project_functions.plotAvC(dfh)
# Regression plot for healthy indivduals
project_functions.plotAvC(dfun)
# Regression plot for unhelathy individuals
# from a quick look we can clearly tell from the regression line that having a unhealthy habit leads to on average a higher charge at each age with only the outliers of the healthy individuals coming close to the average of the unhealthy ones.This is fairly ordinary as more unhealthy habits require more health check uos etc.However on closer inspection we see that in this unhealthy group there is a lot of spread of data points with there looking like there is actually 3 populations in this dataset, this makes sense as the requirement of this dataset was to be smoker OR outside the healthy bmi range which results in 3 different datsets more on that later.
#
# For now we can look at how more likely a indiviudal is to have excess charges if they have a unhealthy trait with frequency ar counts.
# **Bar plots**
# ---
# ---
project_functions.BrPltECD(df)
# Bar plot for whole population
project_functions.BrPltECD(dfh)
# frequency count of if an indivdual is to recieve excess charges in healthy population
project_functions.BrPltECD(dfun)
# frequency count of if an indivdual is to recieve excess charges in unhealthy population
# This further reinforces that a unhealthly life choice leads to a higher chance of an individual receiving a higher than average medical bill charge. We saw that there was outliers on the regression plots and we also couldnt see how much the data was spread for the healthy and unhealthy groups to better visualise this we can use box plots
# **Box plots**
# ---
# ---
project_functions.BoxPlt(df)
# This further shows us the spread of the unhealthy charges this is due to their being multiple distrubutions collected in there but from the analysis of healthy vs unhealthy it further shows how much less the people with a healthy life style pay less charges. We can draw many conclusions just from this graph, we can see that 75% of the healthy population pay less than 10,000 whilst only 50% of the unhealthy population pays below this figure.
# **different Unhealthy categories**
# ---
# ---
# We mentioned earlier how in the unhelathy regression plot there seemed to be 3 different distributions now lets look at these in further depth and how they effect the charges, the 3 categories for this are under bmi,over bmi and smokers. First lets create data frames for each of these categories with only one variable per person
dfs=project_functions.smoker(cwd)
dfs
# Data frame for smokers only
dfob=project_functions.overBmi(cwd)
dfob
# Data frame for over average bmi
dfub=project_functions.underBmi(cwd)
dfub
# Dataframe for under average bmi
# **Regression Plots**
# ---
project_functions.plotAvC(dfs)
# Regression plot for smokers only
#
# We see here that the smokers at the least pay around 14,000 this is larger than the average charge for all patients, this suggests that the leading factor in large charges is smoking
project_functions.plotAvC(dfob)
# Regression plot for over weight people
#
# Here is where we see the majority of our population and this looks very similar to the regression plot of healthy individuals with a similar frequency of outliers above the regression line to the healthy case this points towards elimination of over average bmi from being a leading factor in above average charges.
project_functions.plotAvC(dfub)
# Regression plot for underweight people only
#
# This regression plot actually shows a under average bmi leads towards a much lower charge per age as the lowest charge is around 2000 whilst the maxium is only around 13,000, however this is a very small population compared to the other cases of only 14 so we cannot say this is for certain but thsi evidence combined with the over average bmi helps eliminate bmi being a leading cause in excess medical charges
# These regression plots mostly show that the smokers have a large variety in the charges at each age and that bmi actually does not have too much of effect on the charges at each age and that a lower bmi actually looks like it causes a lower charge at each age group.
# **Bar Plots**
# ---
# ---
project_functions.BrPltECD(dfs)
# Bar plot for smokers only
#
# This further shows that smoking is a leading factor in excess medical charges as from a respectiable sample size we've shown that everyone who smokes will have excess charges.
project_functions.BrPltECD(dfob)
# Bar plot for over weight people only
#
# Here we see that over weight people are more likely to have excess charges as compared to healthy people as they appear to be charged excess charges around 0.15 where as healthy people around around 0.1. This shows that having a over average bmi leads towards a higher chance of being charged more than average costs this is further backed up as this population was quite large so it can be used to draw a better conclusion, there was some outliers but we can compare those with bar plots later.
project_functions.BrPltECD(dfub)
# Bar plot for under weight people only
#
# Here we see something interesting in that under weight people only get charged under the average charges. This is interesting but could be down to the small sample size of this population because of this i wont draw a conclusion that a lower bmi leads to lower medical fees but it can be used to rule it out in a leading factor for higher fees.
# **Box Plots**
# ---
# ---
project_functions.BoxPltob(df)
# Box plot of not over weight vs over weight
#
# Here we can see the over bmi population has a similar spread copared to healthy bmi but the over bmi is shifted upwards by a slight amount, we also see the over bmi has more outliers but this is expected due to a larger population size. This larger population size for over bmi could be due to the increasing obesity crisis in America thus more people are likely to be overweight as this is more normal this could be the reason that the frequency wasnt much higher for this group as its more likely to shift the mean of the dataset. So the higher frequency of excess charges and the more outliers in this case point towards the over bmi population having a larger medical charge on average as the outliers skew the mean thus it isnt a true representation of the mean which would need more investitgation.
project_functions.BoxPltub(df)
# Box plot of not under weight vs under weight
#
# Again we see that the average costs for under bmi is lower and the spread of the data seemes to be the same as the healthy case, there are no outliers in this under bmi population so the mean is more representative of the population.
project_functions.BoxPlts(df)
# Box plot of non smokers vs smokers
#
# The smoker population spread is clearly much larger and higher than the healthy population, we also see no outliers in this case so this spread is a good representation of the whole smoker population. It's obvious looking at this and the previous examples that smoking is the dominate factor in higher medical fees
# From the previous plots we saw that bmi doesnt have too much of an influence in charges, and in fact a lower bmi looked to cause lower fees but the population was too small to draw concrete conclusions from, but in the over bmi case we could look at the higher bmi which would be closer to the obese cases and would look at more extreme cases from the average population. From this we also saw that smoking was the dominant factor for the higher fees which we will investigate some more.
# **Smokers**
# ---
# ---
# We saw from the previous analysis that isolating only idividuals who only smoked showed a trend of excessive medical charges now I want to look at how people who smoke and have different bmi differ with their medical charges
dfas=project_functions.allsmoker(cwd)
dfas
# **Regression Plots**
# ---
# ---
project_functions.plotAvC(dfas)
# **Bar Plots**
# ---
# ---
project_functions.BrPltECD(dfas)
# **Box Plot**
# ---
project_functions.BoxPlts(dfas)
# Looking at these plots its clear to see that even when we extend the smokers case to all the people that do smoke that excess charges is always common between the population. Now we have a larger population we can draw a more concrete conclusion, the regression plot shows us that most of the above average charges are associated with smokers this is further backed up by the bar plot which shows almost all of the population of smokers have above average charges, the final nail for the argument that smoking is the leading factor in higher than average charges is the box plot which shows no outliers and the smallest fee being only just below the excessive charges threshold.
# **Obese**
# ---
# ---
# Now i want to look at how extreme obeseity effects the charges, this is classified as a value of 40 or higher
dfobs=project_functions.obese(cwd)
dfobs
# **Regression Plots**
# ---
# ---
project_functions.plotAvC(dfobs)
# **Bar plots**
# ---
# ---
project_functions.BrPltECD(dfobs)
# **Box plots**
# ---
project_functions.BoxPltob(dfobs)
# This is quite suprising in that the extrme case of obesity shows to have similar distributions to the healthy population with the frequency plot and box being almost similar and the regression plot showing nothing out of the ordinary for our other plots.This shows that even extreme obesity doesnt effect medical charges to a large degree, this is suprising as I thought this would of had a substancial impact on charges.
#
# All of this taken into acount we can say with confidence that the leading factor in excessive charges is smoking and not bmi of an individual.
| Analysis/Hugh Blakemore/analysis conduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/CodeGodZeus/MachineLearning_Python/blob/master/Untitled8.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="l4be6v1vJ1W4"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# + id="MiAQbT2_KUfG" outputId="1d9276f1-2318-4692-efca-8af733f43922" colab={"base_uri": "https://localhost:8080/", "height": 1000}
df=pd.read_csv("/content/IMDB-Movie-Data.csv")
df
# + id="4iW_NaEmKcUn" outputId="4c0c87eb-7cc1-4abe-f88a-bfd2869dc69e" colab={"base_uri": "https://localhost:8080/", "height": 255}
df.head
# + id="jt0sKfklKd5H" outputId="f967c859-415d-4ae0-8896-afd7239d7cd8" colab={"base_uri": "https://localhost:8080/", "height": 255}
print(df.head)
# + id="j0ESjJa0KgiX" outputId="0264fb0d-a972-43eb-e901-deb4acbb251c" colab={"base_uri": "https://localhost:8080/", "height": 153}
print(df.head())
# + id="Yr8av_F7Ky8G" outputId="36d4faf9-7799-4ca4-e7f3-481199c98291" colab={"base_uri": "https://localhost:8080/", "height": 238}
print(df.head(10))
# + id="AgOrwG88K1OG" outputId="79884ff3-06db-4431-8e5b-5a1d1876ea9f" colab={"base_uri": "https://localhost:8080/", "height": 119}
print(df.tail(3))
# + id="SdUJGF5eK9AN" outputId="13e651a2-c3c5-47b1-c3e6-89174d1283ba" colab={"base_uri": "https://localhost:8080/", "height": 357}
print(df.info())
# + id="Hoaty1MuLC6-" outputId="ed7a76f2-cbe0-4360-bcb2-0275530c2342" colab={"base_uri": "https://localhost:8080/", "height": 204}
print(df.describe())
# + id="-XlXSn7lLKNm"
df1=df['Genre']
# + id="RW2N35nwLKWV" outputId="35e662f3-2f3a-4a36-afd6-89aef47f09f6" colab={"base_uri": "https://localhost:8080/", "height": 34}
print(type(df1))
# + id="2c_6cW0gLKiX"
extracteddf=df[['Genre','Rating','Revenue (Millions)']]
# + id="hlkyEmSdLy5X" outputId="9b0011a6-6e75-45d9-802e-4fc4fdea930a" colab={"base_uri": "https://localhost:8080/", "height": 359}
extracteddf.head(10)
# + id="waMQkpXVL8eW" outputId="eb828f06-0c29-40c4-e225-d69f391df58c" colab={"base_uri": "https://localhost:8080/", "height": 465}
multiple_rows = df.loc['Guardians of the Galaxy':'Sing']
multiple_rows = df.iloc[0:4]
multiple_rows
# + id="2k-gkzvWNEM4" outputId="49d4a98c-55fc-4331-c10d-c4540831dc24" colab={"base_uri": "https://localhost:8080/", "height": 1000}
df[df['Year']==2015]
# + id="rjeCp0L0NEZ-" outputId="c4cf2fcd-d5e4-4774-cb7e-b00a7370f113" colab={"base_uri": "https://localhost:8080/", "height": 1000}
df[df['Rating']>8.0]
# + id="MboPGPDqNp8d" outputId="51635ac0-d0c0-4ee7-a4ae-ecb0ef916dca" colab={"base_uri": "https://localhost:8080/", "height": 450}
df.groupby('Director').sum()
# + id="pH-jO9M_NqD9" outputId="e0923be8-e211-4ae1-e57c-3cb73e68b492" colab={"base_uri": "https://localhost:8080/", "height": 421}
df.groupby("Year").sum()
# + id="oNH2MoCxOAw_" outputId="c1ea2069-6b72-471e-dc8a-d3122cdd7245" colab={"base_uri": "https://localhost:8080/", "height": 450}
df.groupby('Director')[['Rating']].sum()
# + id="2OmtvpxNOTK1" outputId="f3112094-fff3-4f80-e464-430cdfb982d3" colab={"base_uri": "https://localhost:8080/", "height": 450}
df.groupby('Director')[['Revenue (Millions)']].sum().sort_values(['Revenue (Millions)'],ascending=False)
# + id="rPIhlOlRPll8" outputId="419df0e6-f738-4090-a0ca-9ca1438de6eb" colab={"base_uri": "https://localhost:8080/", "height": 436}
df.isnull()
# + id="Ctf-ttumPlyP" outputId="a84f1fe4-382c-4345-acb8-c6458253e04f" colab={"base_uri": "https://localhost:8080/", "height": 238}
df.isnull().sum()
# + id="qgcZNQzOPtXk"
df_mean=df['Revenue (Millions)']
meana=df_mean.mean()
# + id="BD8rljcmP-yz"
df_mean.fillna(meana,inplace=True)
# + id="RwlS5BfCQP1a" outputId="f22aab87-80a4-4ab8-d0ab-e083ee252179" colab={"base_uri": "https://localhost:8080/", "height": 238}
df.isnull().sum()
| Untitled8.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Consumer segmentation -- mixed datatypes
# K-mean algorithm does not work well for segmenting consumers when there are both numerical and categorical variables. Instead, a modified method, **k-prototypes algorithm**, should be used to address the issue. Given the cluster number $k$, instead of minimizing SSE in k-means, k-prototypes minimizes the "clustering cost," which measures the clustering misfit for the mixed datatypes. You need to install the package "kmodes" from your anaconda prompt/cmd using `conda install -c conda-forge kmodes` or `pip install kmodes`.
# <br/>
#
#
#
# ## Importing packages and dataset
# + tags=[]
import numpy as np
import pandas as pd
#####################################
from kmodes.kprototypes import KPrototypes # We will use the k-prototypes algorithm
#####################################
# + [markdown] tags=[]
# We will use "MallCustomersAllVariables.csv" for analysis
# + tags=[]
url = "https://raw.githubusercontent.com/zoutianxin1992/MarketingAnalyticsPython/main/Marketing%20Analytics%20in%20Python/Segmentation/Datasets/MallCustomersAllVariables.csv"
df = pd.read_csv(url,index_col=0) # use the first column (customer id) as index
df.info()
# + [markdown] tags=[]
# Rename the variables to follow the naming conventions.
# + tags=[]
df = df.rename(columns = {"Gender":"gender",
"Age":"age",
"Annual Income (k$)":"annual_income",
"Spending Score (1-100)":"spending_score"})
df.head()
# -
# Note that gender (F, M) is a categorical variable, so k-mean algorithm should not be applied here. We need to use k-prototypes algorithm to accommodate categorical variables.
# + [markdown] tags=[]
# ## Segmenting consumers into three segments
# -
# ### Normalize the variables to a 0-1 scale (only for **numerical** variables)
# $$
# X_{transform} = \frac {X_{original} - X_{min}} {X_{max} - X_{min}}
# $$
# Since it makes no sense to normalize categorical variables to a 0-1 scale, the normalization should be applied **only to numerical variables**: `age`, `annual_income`, and `spending_score`.
# + tags=[]
df_normalized = df.copy() # create a copy of the orignial dataset
################################
df_normalized[['age','annual_income','spending_score']] = \
(df[['age','annual_income','spending_score']]-df[['age','annual_income','spending_score']].min()) \
/(df[['age','annual_income','spending_score']].max()-df[['age','annual_income','spending_score']].min())
df_normalized.head()
##################################
# -
# ### Applying k-prototypes algorithm to normalized data
# + tags=[]
#################################
kprotoSpec = KPrototypes(n_clusters = 3, n_init = 100) # setup the k-prototypes model specs
# apply the above method to normalized dataset
kproto_result3 = kprotoSpec.fit(df_normalized,categorical = [0]) # The categorical variable, gender, is in column 0
##############################
# + [markdown] tags=[]
# ## Post-segmentation analysis
# + [markdown] tags=[]
# ### Which segment does each consumer belong to?
# -
# Create a new column in the **original** dataframe for which segment a consumer belongs to.
# + tags=[]
###########################
df["segment"] = kproto_result3.labels_
df.head()
#############################
# + [markdown] tags=[]
# ### Summarizing segment characteristics
# -
# For each segment, summarize the mean/min/max's of all the **numerical** variable. (It makes no sense to calculate these stats for **categorical variables**.)
# For each segment, summarize the percentage of male/female customers in each segment.
# Also count the number of consumers in each segment.
#
# While Python has built-in functions for "mean", "min", "max", etc., the system has no ready functions to calculate "proportion of male." So, we need to define it by ourselves. The proportion can be calculated as `x[x == "M"].count()/x.count()`, where `x` is our `gender` data. </br>
# Specifically, `lambda x: x[x == "M"].count()/x.count() ` defines an inline function of `x` that calculates the proportion of male in `x`.
# +
summary_table = df.groupby("segment").aggregate({
############################
"gender": (lambda x: x[x == "M"].count()/x.count()), # lambda allows us to define an inline function
##################################
"age":["mean","min","max"],
"annual_income": ["mean","min","max"],
"spending_score": ["mean","min","max"],
"segment": "count"
}
)
summary_table
# -
#
#
#
| Marketing Analytics in Python/Segmentation/Notebooks/sgmt_mixvar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from align.spark.schemas import ARCHIVE_ORG_SCHEMA
import pyspark.sql.functions as F
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName('CC-BY-license').getOrCreate()
input_catalogue_path = "gs://the-peoples-speech-west-europe/archive_org/Mar_7_2021/EXPANDED_LICENSES_FILTERED_ACCESS.jsonl.gz"
df = spark.read.format('json').schema(ARCHIVE_ORG_SCHEMA).load(input_catalogue_path)
print(df.count())
df.show()
columns = [df.metadata.licenseurl, df.metadata.creator, df.metadata.title, df.metadata.credits]
licenses = ['https://creativecommons.org/licenses/by/4.0/',
'https://creativecommons.org/licenses/by/3.0/',
'http://creativecommons.org/licenses/by/2.0/',
'http://creativecommons.org/licenses/by/3.0/',
'https://creativecommons.org/licenses/by/3.0',
'http://creativecommons.org/licenses/by/2.5/',
'http://creativecommons.org/licenses/by/2.5/pl/',
'http://creativecommons.org/licenses/by/2.5/it/',
'http://creativecommons.org/licenses/by/4.0/',
'http://creativecommons.org/licenses/by/3.0/us/',
'http://creativecommons.org/licenses/by/2.5/ar/']
df = df.select(columns)
df.show()
c = '`metadata.licenseurl`'
df.agg(F.count(F.when(F.isnull(c), c)).alias('NULL_Count')).show()
# ### Look NA values
df_pd = df.toPandas()
df_pd.head()
df_pd.shape
df_pd.isna().sum()
df_pd['metadata.licenseurl'].value_counts(normalize=True) * 100
df_pd[df_pd['metadata.licenseurl'].isna()]
df_pd['metadata.licenseurl'].unique()
# ## Filter values by license
df = (df.withColumnRenamed('metadata.licenseurl','licenseurl').withColumnRenamed('metadata.creator', 'creator')
.withColumnRenamed('metadata.title', 'title').withColumnRenamed('metadata.credits', 'credits'))
df = df.dropna(subset=['licenseurl'])
df.show()
regexp = r"(http|https)://creativecommons.org/licenses/by/(1[.]0|2[.]0|2[.]5|3[.]0|4[.]0)"
df = df.filter(df['licenseurl'].rlike(regexp))
df.show()
len(df.columns)
# ## Types of licences
licenses_url = df.select('licenseurl').distinct().collect()
licenses_urls = []
for license in licenses_url:
licenses_urls.append(license[0])
licenses_urls
# ## Convert to SQL
df.createOrReplaceTempView("CC_BY_creators")
spark.sql('SELECT * FROM CC_BY_creators').show()
# ## Save as txt
# +
def myConcat(*cols):
concat_columns = []
for c in cols[:-1]:
concat_columns.append(F.coalesce(c, F.lit("*")))
concat_columns.append(F.lit(" "))
concat_columns.append(F.coalesce(cols[-1], F.lit("*")))
return F.concat(*concat_columns)
df = df.withColumn("credits", myConcat(*df.columns)).select("credits")
df.coalesce(1).write.format("text").option("header", "false").mode("append").save("credits.txt")
# -
| galvasr2/codelabs/CC-BY_filter #15.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
data_path = 'train.tfrecords' # address to save the hdf5 file
def init_weights(shape):
init_random_dist = tf.truncated_normal(shape, stddev=0.1)
return tf.Variable(init_random_dist)
def init_bias(shape):
init_bias_vals = tf.constant(0.1, shape=shape)
return tf.Variable(init_bias_vals)
def conv2d(x, W):
return tf.nn.conv2d(x, W, strides=[1, 1, 1, 1], padding='SAME')
def max_pool_2by2(x):
return tf.nn.max_pool(x, ksize=[1, 2, 2, 1],
strides=[1, 2, 2, 1], padding='SAME')
def convolutional_layer(input_x, shape):
W = init_weights(shape)
b = init_bias([shape[3]])
return tf.nn.relu(conv2d(input_x, W) + b)
def normal_full_layer(input_layer, size):
input_size = int(input_layer.get_shape()[1])
W = init_weights([input_size, size])
b = init_bias([size])
return tf.matmul(input_layer, W) + b
# +
x = tf.placeholder(tf.float32,shape=[None,30000])
y_true = tf.placeholder(tf.float32,shape=[None,2])
### Layers
x_image = tf.reshape(x,[-1,100,100,3])
# Using a 6by6 filter here, used 5by5 in video, you can play around with the filter size
# You can change the 32 output, that essentially represents the amount of filters used
# You need to pass in 32 to the next input though, the 1 comes from the original input of
# a single image.
convo_1 = convolutional_layer(x_image,shape=[6,6,3,32])
print(convo_1.get_shape())
convo_1_pooling = max_pool_2by2(convo_1)
print(convo_1_pooling.get_shape())
# Using a 6by6 filter here, used 5by5 in video, you can play around with the filter size
# You can actually change the 64 output if you want, you can think of that as a representation
# of the amount of 6by6 filters used.
convo_2 = convolutional_layer(convo_1_pooling,shape=[6,6,32,64])
print(convo_2.get_shape())
convo_2_pooling = max_pool_2by2(convo_2)
print(convo_2_pooling.get_shape())
# Why 7 by 7 image? Because we did 2 pooling layers, so (28/2)/2 = 7
# 64 then just comes from the output of the previous Convolution
convo_2_flat = tf.reshape(convo_2_pooling,[-1,7*7*64])
print(convo_2_flat.get_shape())
full_layer_one = tf.nn.relu(normal_full_layer(convo_2_flat,1024))
print(full_layer_one.get_shape())
# NOTE THE PLACEHOLDER HERE!
hold_prob = tf.placeholder(tf.float32)
full_one_dropout = tf.nn.dropout(full_layer_one,keep_prob=hold_prob)
print(full_one_dropout.get_shape())
y_pred = normal_full_layer(full_one_dropout,10)
print(y_pred.get_shape())
# +
cross_entropy = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(labels=y_true,logits=y_pred))
print(cross_entropy.get_shape())
### Optimizer
optimizer = tf.train.AdamOptimizer(learning_rate=0.0001)
train = optimizer.minimize(cross_entropy)
### Intialize Variables
init = tf.global_variables_initializer()
### Session
steps = 5000
# -
with tf.Session() as sess:
feature = {'train/image': tf.FixedLenFeature([], tf.string),
'train/label': tf.FixedLenFeature([], tf.int64)}
# Create a list of filenames and pass it to a queue
filename_queue = tf.train.string_input_producer([data_path], num_epochs=1)
# Define a reader and read the next record
reader = tf.TFRecordReader()
_, serialized_example = reader.read(filename_queue)
# Decode the record read by the reader
features = tf.parse_single_example(serialized_example, features=feature)
# Convert the image data from string back to the numbers
image = tf.decode_raw(features['train/image'], tf.float32)
# Cast label data into int32
label = tf.cast(features['train/label'], tf.int32)
# Reshape image data into the original shape
image = tf.reshape(image, [100, 100, 3])
# Any preprocessing here ...
# Creates batches by randomly shuffling tensors
images, labels = tf.train.shuffle_batch([image, label], batch_size=5, capacity=5, num_threads=1, min_after_dequeue=1)
# Initialize all global and local variables
init_op = tf.group(tf.global_variables_initializer(), tf.local_variables_initializer())
sess.run(init_op)
# Create a coordinator and run all QueueRunner objects
coord = tf.train.Coordinator()
threads = tf.train.start_queue_runners(coord=coord)
for batch_index in range(5):
img, lbl = sess.run([images, labels])
#sess.run(train,feed_dict={x:img,y_true:lbl,hold_prob:0.5})
img = img.astype(np.uint8)
for j in range(5):
plt.subplot(2, 3, j + 1)
plt.imshow(img[j, ...])
plt.title('apple' if lbl[j] == 1 else 'not apple')
plt.show()
# Stop the threads
coord.request_stop()
# Wait for threads to stop
coord.join(threads)
sess.close()
| reader.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="vVjGiHNkg8_7"
# <table align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/ShopRunner/collie/blob/main/tutorials/04_partial_credit_loss.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/ShopRunner/collie/blob/main/tutorials/04_partial_credit_loss.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a>
# </td>
# <td>
# <a target="_blank" href="https://raw.githubusercontent.com/ShopRunner/collie/main/tutorials/04_partial_credit_loss.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" /> Download notebook</a>
# </td>
# </table>
# + id="XyEpvflUg9AE"
# for Collab notebooks, we will start by installing the ``collie`` library
# !pip install collie --quiet
# + id="8BPm_KDPg9AG" outputId="52baa91c-5a52-407a-d353-c8f5c883c7a5"
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
# %env DATA_PATH data/
# + id="d8WsbqX8g9AI"
import os
import numpy as np
import pandas as pd
from pytorch_lightning.utilities.seed import seed_everything
from IPython.display import HTML
import joblib
import torch
from collie.metrics import mapk, mrr, auc, evaluate_in_batches
from collie.model import CollieTrainer, MatrixFactorizationModel
from collie.movielens import get_movielens_metadata, get_recommendation_visualizations
# + [markdown] id="44gLNrW-ghT_"
# ## Load Data From ``01_prepare_data`` Notebook
# If you're running this locally on Jupyter, you should be able to run the next cell quickly without a problem! If you are running this on Colab, you'll need to regenerate the data by running the cell below that, which should only take a few extra seconds to complete.
# + id="XKKPmYi_ghT_" outputId="23c16373-6747-4255-9f52-7091312e0b71"
try:
# let's grab the ``Interactions`` objects we saved in the last notebook
train_interactions = joblib.load(os.path.join(os.environ.get('DATA_PATH', 'data/'),
'train_interactions.pkl'))
val_interactions = joblib.load(os.path.join(os.environ.get('DATA_PATH', 'data/'),
'val_interactions.pkl'))
except FileNotFoundError:
# we're running this notebook on Colab where results from the first notebook are not saved
# regenerate this data below
from collie.cross_validation import stratified_split
from collie.interactions import Interactions
from collie.movielens import read_movielens_df
from collie.utils import convert_to_implicit, remove_users_with_fewer_than_n_interactions
df = read_movielens_df(decrement_ids=True)
implicit_df = convert_to_implicit(df, min_rating_to_keep=4)
implicit_df = remove_users_with_fewer_than_n_interactions(implicit_df, min_num_of_interactions=3)
interactions = Interactions(
users=implicit_df['user_id'],
items=implicit_df['item_id'],
ratings=implicit_df['rating'],
allow_missing_ids=True,
)
train_interactions, val_interactions = stratified_split(interactions, test_p=0.1, seed=42)
print('Train:', train_interactions)
print('Val: ', val_interactions)
# + [markdown] id="8hkoWyfVg9AK"
# # Partial Credit Loss
# Most of the time, we don't *only* have user-item interactions, but also side-data about our items that we are recommending. These next two notebooks will focus on incorporating this into the model training process.
#
# In this notebook, we're going to add a new component to our loss function - "partial credit". Specifically, we're going to use the genre information to give our model "partial credit" for predicting that a user would like a movie that they haven't interacted with, but is in the same genre as one that they liked. The goal is to help our model learn faster from these similarities.
# + [markdown] id="4iFhjr7eg9AK"
# ## Read in Data
# + id="_HsdIsXVg9AL" outputId="9bd787d0-7c38-44bc-ca93-31bd18bf8a88"
# read in the same metadata used in notebook ``03``
metadata_df = get_movielens_metadata()
metadata_df.head()
# + id="LSy_-Jsxg9AL" outputId="2a4bfca7-d216-40f5-a2c3-0ca5e99b5653"
# to do the partial credit calculation, we need this data in a slightly different form.
# Instead of the one-hot-encoded version above, we're going to make a ``1 x n_items`` tensor
# with a number representing the first genre associated with the film, for simplicity.
# Note that with Collie, we could instead make a metadata tensor for each genre and decade
genres = (
torch.tensor(metadata_df[[c for c in metadata_df.columns if 'genre' in c]].values)
.topk(1)
.indices
.view(-1)
)
genres
# + id="35eEIFu7g9AM" outputId="ae84ab70-9bc3-47a1-8350-65b568ac42ad"
# and, as always, set our random seed
seed_everything(22)
# + [markdown] id="LhaQLQQig9AM"
# ## Train a model with our new loss
# + id="Sysr04kSg9AN"
# now, we will pass in ``metadata_for_loss`` and ``metadata_for_loss_weights`` into the model
# ``metadata_for_loss`` should have a tensor containing the integer representations for metadata
# we created above for every item ID in our dataset
# ``metadata_for_loss_weights`` should have the weights for each of the keys in ``metadata_for_loss``
model = MatrixFactorizationModel(
train=train_interactions,
val=val_interactions,
embedding_dim=10,
lr=1e-2,
metadata_for_loss={'genre': genres},
metadata_for_loss_weights={'genre': 0.4},
)
# + colab={"referenced_widgets": ["", "44c30227b3964ef38c46726d4b7ea34b"]} id="ZAk1C815g9AN" outputId="f6f3778d-ccdb-48c2-9ac9-ec3e1025f199"
trainer = CollieTrainer(model=model, max_epochs=10, deterministic=True)
trainer.fit(model)
# + [markdown] id="NQdGTCPvg9AN"
# ## Evaluate the Model
# + [markdown] id="ISQzTUnVg9AO"
# Again, we'll evaluate the model and look at some particular users' recommendations to get a sense of what these recommendations look like using a partial credit loss function during model training.
# + colab={"referenced_widgets": ["77771450d4fe40ada165f6d9890bc4f8"]} id="6STWH4Ozg9AO" outputId="d216cd87-ec0a-4903-87ea-933314f5557f"
mapk_score, mrr_score, auc_score = evaluate_in_batches([mapk, mrr, auc], val_interactions, model)
print(f'MAP@10 Score: {mapk_score}')
print(f'MRR Score: {mrr_score}')
print(f'AUC Score: {auc_score}')
# + [markdown] id="Bk75mVQWg9AP"
# Broken record alert: we're not seeing as much performance increase here compared to the model in Tutorial ``02`` because MovieLens 100K has so few items. For a more dramatic difference, try training this model on a larger dataset, such as MovieLens 10M, adjusting the architecture-specific hyperparameters, or train longer.
# + id="dB6eeXWfg9AP" outputId="d1b03b63-9781-4139-8f7c-a4a20712eb81"
user_id = np.random.randint(0, train_interactions.num_users)
display(
HTML(
get_recommendation_visualizations(
model=model,
user_id=user_id,
filter_films=True,
shuffle=True,
detailed=True,
)
)
)
# + [markdown] id="ZoIh0oHfg9AQ"
# Partial credit loss is useful when we want an easy way to boost performance of any implicit model architecture, hybrid or not. When tuned properly, partial credit loss more fairly penalizes the model for more egregious mistakes and relaxes the loss applied when items are more similar.
#
# Of course, the loss function isn't the only place we can incorporate this metadata - we can also directly use this in the model (and even use a hybrid model combined with partial credit loss). The next tutorial will train a hybrid Collie model!
# + [markdown] id="WbSExVSTg9AQ"
# -----
| tutorials/04_partial_credit_loss.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 2D Nuclear Segmentation with Mask-RCNN
# +
# This notebook is for benchmarking the output of the original trained featurenet model
import os
import errno
import numpy as np
import deepcell
# +
# create folder for this set of experiments
experiment_folder = "featurenet_samir/"
MODEL_DIR = os.path.join("/data/analyses/", experiment_folder)
NPZ_DIR = "/data/npz_data/20201018_freeze/"
LOG_DIR = '/data/logs'
if not os.path.isdir(MODEL_DIR):
os.makedirs(MODEL_DIR)
# +
from tensorflow.keras.optimizers import SGD, Adam
from deepcell.utils.train_utils import rate_scheduler
from deepcell.utils.retinanet_anchor_utils import get_anchor_parameters
from deepcell.training import train_model_retinanet
from deepcell import model_zoo
from deepcell_toolbox.multiplex_utils import multiplex_preprocess
from timeit import default_timer
from skimage.measure import label
from deepcell_toolbox.deep_watershed import deep_watershed_mibi
def calc_jaccard_index_object(metric_predictions, true_labels, pred_labels):
jacc_list = []
for i in range(true_labels.shape[0]):
y_true = true_labels[i, :, :, 0]
y_pred = pred_labels[i, :, :, 0]
true_ids = metric_predictions[i][0]['correct']['y_true']
pred_ids = metric_predictions[i][0]['correct']['y_pred']
current_accum = []
for id in range(len(true_ids)):
true_mask = y_true == true_ids[id]
pred_mask = y_pred == pred_ids[id]
current_jacc = (np.sum(np.logical_and(true_mask, pred_mask)) /
np.sum(np.logical_or(true_mask, pred_mask)))
current_accum.append(current_jacc)
jacc_list.append(current_accum)
return jacc_list
model_splits = ['1', '2', '3']
metrics = {}
for split in model_splits:
print('loading data')
test_name = "20201018_multiplex_seed_{}_test_256x256.npz".format(split)
test_dict = np.load(NPZ_DIR + test_name)
X_test = test_dict['X'][..., :1]
#X_test = multiplex_preprocess(X_test)
y_test = test_dict['y']
model_name = 'featurenet_samir.h5'.format(split)
# start timing
time_start = default_timer()
print('creating model')
model = model_zoo.bn_feature_net_skip_2D(
receptive_field=61,
n_skips=3,
n_features=3,
norm_method='whole_image',
n_conv_filters=32,
n_dense_filters=128,
last_only=False,
input_shape=(256, 256, 1))
model.load_weights(MODEL_DIR + model_name)
print('predicting')
pixelwise = model.predict(X_test)[-1]
print('postprocessing')
labeled_images = deep_watershed_mibi({'inner-distance': pixelwise[:, :, :, 1:2],
'pixelwise-interior': pixelwise[:, :, :, 1:2]},
maxima_threshold=0.3, maxima_model_smooth=0,
interior_threshold=0.3, interior_model_smooth=0,
radius=3,
small_objects_threshold=10,
fill_holes_threshold=10,
pixel_expansion=3)
# end time
time_end = default_timer()
print("elapsed time is {}".format(time_end - time_start))
for i in range(labeled_images.shape[0]):
img = labeled_images[i, :, :, 0]
img = label(img)
labeled_images[i, :, :, 0] = img
for i in range(y_test.shape[0]):
img = y_test[i, :, :, 0]
img = label(img)
y_test[i, :, :, 0] = img
# calculating accuracy
print("calculating accuracy")
db = DatasetBenchmarker(y_true=y_test,
y_pred=labeled_images,
tissue_list=test_dict['tissue_list'],
platform_list=test_dict['platform_list'],
model_name='default_model')
tissue_stats, platform_stats = db.benchmark()
jacc = calc_jaccard_index_object(db.metrics.predictions, y_test, labeled_images)
jacc = np.concatenate(jacc)
jacc_mean = np.mean(jacc)
print(jacc_mean)
metrics[split] = {'tissue_stats':tissue_stats, 'platform_stats': platform_stats, 'jacc':jacc_mean}
# -
metrics['3']['tissue_stats']['all']['f1']
np.savez_compressed(os.path.join('/data/analyses/', 'featurenet_metrics_samir_jacc.npz'), **metrics)
# +
# Copyright 2016-2020 The Van Valen Lab at the California Institute of
# Technology (Caltech), with support from the Paul Allen Family Foundation,
# Google, & National Institutes of Health (NIH) under Grant U24CA224309-01.
# All rights reserved.
#
# Licensed under a modified Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.github.com/vanvalenlab/caliban-toolbox/LICENSE
#
# The Work provided may be used for non-commercial academic purposes only.
# For any other use of the Work, including commercial use, please contact:
# <EMAIL>
#
# Neither the name of Caltech nor the names of its contributors may be used
# to endorse or promote products derived from this software without specific
# prior written permission.
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
import numpy as np
from deepcell_toolbox.metrics import Metrics, stats_pixelbased
from scipy.stats import hmean
class DatasetBenchmarker(object):
"""Class to perform benchmarking across different tissue and platform types
Args:
y_true: true labels
y_pred: predicted labels
tissue_list: list of tissue names for each image
platform_list: list of platform names for each image
model_name: name of the model used to generate the predictions
metrics_kwargs: arguments to be passed to metrics package
Raises:
ValueError: if y_true and y_pred have different shapes
ValueError: if y_true and y_pred are not 4D
ValueError: if tissue_ids or platform_ids is not same length as labels
"""
def __init__(self,
y_true,
y_pred,
tissue_list,
platform_list,
model_name,
metrics_kwargs={}):
if y_true.shape != y_pred.shape:
raise ValueError('Shape mismatch: y_true has shape {}, '
'y_pred has shape {}. Labels must have the same'
'shape.'.format(y_true.shape, y_pred.shape))
if len(y_true.shape) != 4:
raise ValueError('Data must be 4D, supplied data is {}'.format(y_true.shape))
self.y_true = y_true
self.y_pred = y_pred
if len({y_true.shape[0], len(tissue_list), len(platform_list)}) != 1:
raise ValueError('Tissue_list and platform_list must have same length as labels')
self.tissue_list = tissue_list
self.platform_list = platform_list
self.model_name = model_name
self.metrics = Metrics(model_name, **metrics_kwargs)
def _benchmark_category(self, category_ids):
"""Compute benchmark stats over the different categories in supplied list
Args:
category_ids: list specifying which category each image belongs to
Returns:
stats_dict: dictionary of benchmarking results
"""
unique_ids = np.unique(category_ids)
# create dict to hold stats across each category
stats_dict = {}
for uid in unique_ids:
print("uid is {}".format(uid))
stats_dict[uid] = {}
category_idx = np.isin(category_ids, uid)
# sum metrics across individual images
for key in self.metrics.stats:
stats_dict[uid][key] = self.metrics.stats[key][category_idx].sum()
# compute additional metrics not produced by Metrics class
stats_dict[uid]['recall'] = \
stats_dict[uid]['correct_detections'] / stats_dict[uid]['n_true']
stats_dict[uid]['precision'] = \
stats_dict[uid]['correct_detections'] / stats_dict[uid]['n_pred']
stats_dict[uid]['f1'] = \
hmean([stats_dict[uid]['recall'], stats_dict[uid]['precision']])
pixel_stats = stats_pixelbased(self.y_true[category_idx] != 0,
self.y_pred[category_idx] != 0)
stats_dict[uid]['jaccard'] = pixel_stats['jaccard']
return stats_dict
def benchmark(self):
self.metrics.calc_object_stats(self.y_true, self.y_pred)
tissue_stats = self._benchmark_category(category_ids=self.tissue_list)
platform_stats = self._benchmark_category(category_ids=self.platform_list)
all_stats = self._benchmark_category(category_ids=['all'] * len(self.tissue_list))
tissue_stats['all'] = all_stats['all']
platform_stats['all'] = all_stats['all']
return tissue_stats, platform_stats
| 2021-Greenwald_Miller_et_al-Mesmer/notebooks/evaluation/Benchmark_metrics_featurenet_samir.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Solve the clustering problem using QAOA
# **Author: <NAME>**
# <br>
# <EMAIL>
#
# In this notebook we demonstrate a start-to-finish project workflow for using Quantum Approximate Optimization Algorithm to cluster a simple dataset. Along the way, we will explain the major concepts of QAOA and build intuition as to how QAOA can be used to solve clustering problems. This notebook will steer away from heavy mathematical explanations in favor of a higher level view of the algorithm's core components. It is mainly geared towards users that don't have physics background but come from computer science.
# + [markdown] tags=["hide_all"]
# ## Contents
# 1. [Variational Hybrid Algorithms](#variational_hybrid_algorithms)
# 2. [The Maxcut Problem](#maxcut_problem)
# 3. [From Maxcut to QUBO](#maxcut_to_qubo)
# 4. [From QUBO to a Hamiltonian](#qubo_to_hamiltonian)
# 5. [Minimize the Hamiltonian with QAOA](#apply_qaoa)
# -
# <a id="variational_hybrid_algorithms"></a>
#
# ## Variational Hybrid Algorithms
#
# We often take for granted the many decades of progress that lead to today's widespread use of classical computers. As memory and compute power become ever cheapened by Moore's Law, the pressure to find optimal resource allocations for algorithms shrinks away. However, with quantum computers in their early stages, they still feel this daunting requirement. In response to this, a family of algorithms known as *variational hybrid quantum-classical algorithms* was created, with the notion that quantum resources can be made more useful when partnered with classical routines. The Quantum Approximate Optimization Algorithm (QAOA), belongs to the family of variatonal hybrid algorithms.
#
# We can infer a lot from merely unpacking this name. The presence of '*variational*' tells us these algorithms will follow an iterative approach, while '*hybrid*' tells us they will leverage the use of both quantum and classical computers. In fact, this describes the main flow of the algorithm, with all that needs be answered is *when* does this iteration stop and *what* information is passed between devices.
#
# 
# *A visual representation of a generic variational hybrid quantum-classical algorithm.*
#
# To answer the question of *what*, we note that the main goal of QAOA is optimize a set of **parameters**, which we denote as $\vec{\gamma}$ and $\vec{\beta}$. You'll notice that these symbols are vectors, as such they are $n-$length. We discuss later what aspects of our problem decide the value of $n$ in the second notebook.
#
# $\vec{\gamma}$ and $\vec{\beta}$ parameterize a **cost function** which is evaluated with our **Quantum Circuit** to produce a cost value. This output value is input to the optimizer, and is used to determine whether the nudging of our parameters is in a direction of lower cost. We will sometimes call the cost value an **expectation value**, represented by $\langle\psi|Cost|\psi\rangle$, which is the expected value of the cost function $Cost$ over the **wave function** $\psi$. If you were caught off guard by the term 'wave function', then it is equally as effective to think of $\langle\psi|Cost|\psi\rangle$ as the notion of cost as in the more traditional machine learning sense. The **Classical Optimizer** will return updated parameters to the quantum circuit for re-evaluation, and the cycle repeats.
#
# *When* does this algorithm stop? Well, once a stopping criterion is met of course. This criterion is often a pre-defined maximum number of iterations, or occurs after a repeat number of evaluations land within the same threshold of convergence (a tolerance for the cost value in which we consider numbers within an $\epsilon-$window the same). Once this criterion is met, the **optimized parameters** are returned and used to define the solution.
#
# 
# *A visual representation of QAOA in the format of a variational hybrid algorithm.*
#
# The above description should leave you with many questions.
# - How does the above process solve a clustering problem?
# - How exactly do $\vec{\gamma}$ and $\vec{\beta}$ define the solution?
# - How do we define a meaningful cost function for our problem?
# - What in the world is a wave function?
#
# We hope to answer these and more. For now, if you feel comfortable with the critical vocabulary of QAOA (the bolded words), then you'll be well prepared for the explanations below.
# ***
# ### Data Preparation
# Now let's get to the fun part! We will import our data and define the problem setting as a highly manicured example for this clustering demo.
#
# The dataset we will be using is the **Pokemon dataset**, which can be found on [Github](https://gist.github.com/armgilles/194bcff35001e7eb53a2a8b441e8b2c6). In our journey to Catch 'Em All, we will attempt to cluster Pokemon into Legendary and non-Legendary classes.
#
# **Import Libraries**
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# **Import Data**
df = pd.read_csv('./data/pokemon.csv')
df = df.set_index('#') #index pokemon by their ID number
df = df.rename_axis('ID') #rename axis to 'ID' instead of '#'
df = df.loc[~df.index.duplicated(keep='first')] #drop duplicates
df.head()
# To avoid the many bells and whistles of later iterations of Pokemon games, we'll stick to our roots and only consider Pokemon from the first three generations.
df = df.loc[df['Generation']<=3]
df.sample(frac=1).head() #sample the whole dataset (frac=1) to shuffle the arrangement
print('Percent of Non-Legendary Pokemon: %.2f' %((df.Legendary.count()-df.Legendary.sum())/df.Legendary.count()))
print('Percent of Legendary Pokemon: %.2f' %((df.Legendary.sum())/df.Legendary.count()))
# We can see that the classes are quite unevenly distributed. To remedy this, we will randomly select 5 Legendary and 5 Non-Legendary Pokemon to act as our samples to be clustered.
legendary = df.loc[df['Legendary'] == True].sample(5)
non_legendary = df.loc[df['Legendary'] == False].sample(5)
pokemon = pd.concat([legendary,non_legendary])
# To further simplify the problem, and not worry about the encoding of categorical data, we will only consider numerical values in our clustering of the data.
numerical_columns = ['Total','HP','Attack','Defense','Sp. Atk','Sp. Def','Speed']
labels = pokemon['Legendary']
data = pokemon[numerical_columns].copy()
data.head()
# We now have a dataset which is ready to be processed, but we may not be exactly clear on what to do with it. For that we must further understand how the QAOA process detailed above is actually used to solve a clustering problem.
# <a id="maxcut_problem"></a>
#
# ## The Maxcut Problem
#
# As laid out by [Rigetti's paper on QAOA](https://arxiv.org/pdf/1712.05771.pdf), there are a number of important steps that we must follow to map the problem of clustering into a format which QAOA can process. Broadly speaking, QAOA solves the **MAXCUT** problem, in which a graph of $n$ vertices is separated into two complementary subsets, $S$ and $S^{c}$, such that the number of edges between $S$ and $S^{c}$ is as large as possible.
#
#
# 
#
# *A depiction of the maxcut problem, displaying a cut which separates white and black vertices. Image credit:
# __[Wikipedia](https://en.wikipedia.org/wiki/Maximum_cut#/media/File:Max-cut.svg)__*
#
# This problem can be made more sophisticated by adding numerical values as <i>weights</i> to the edges, such that the best solution maximizes the sum of weights which separate $S$ and $S^{c}$. This is precisely the approach we take in using MAXCUT to cluster our data.
#
# We allow the weights associated to each edge to be some notion of distance between points. In this way, the sets dictated by our optimal cut, $S$ and $S^{c}$, separate the data into binary clusters which are maximally distant (and hence, maximally dissimilar) from one another.
#
# From our current understanding, we can already begin to formulate some first steps in preparing our data to fit this frameowrk.
#
# We can use the `distances_dataset` function from `entropica_qaoa.utilities` to easily turn this set of points into the desired matrix of pairwise distances.
# +
from entropica_qaoa.utilities import distances_dataset
dist = pd.DataFrame(distances_dataset(data.values),
index=data.index,columns=data.index)
dist.iloc[0:5, 0:5]
# -
df.loc[dist.index].head()
# <a id="maxcut_to_qubo"></a>
#
# ## From Maxcut to QUBO
# With an understanding of the Maxcut structure which produces our clustered output, we ask ourselves how we can turn what is effectively a graph problem into the setting of an optimization problem. The answer is to map our Maxcut interpretation into a **Quadratic Unconstrainted Binary Optimization** ([QUBO](https://en.wikipedia.org/wiki/Quadratic_unconstrained_binary_optimization)) problem. QUBO problems attempt to minimize a quadratic polynomial with binary variables. Luckily, MAXCUT already has a well-known QUBO cost function. This cost function is sophisticated enough to allow for our pairwise distanes to be meaningfully included, as well as to allow for the inclusion of bias terms on individual samples.
#
# $$
# Cost=-\sum_{\langle i j\rangle} J_{i j} \sigma_{i} \sigma_{j}-\mu \sum_{j} h_{j} \sigma_{j}
# $$
#
# To explain the notation:
# - $\sigma_{i}$ is the cluster class (-1 or 1) of sample $i$
# - $J_{i j}$ is the distance between sample $i$ and sample $j$
# - $h_{j}$ is a bias term on sample $j$
# - $\mu$ is a universal weight applied to all bias terms
#
# By convention, a negative sign is applied to the cost function, as above. In quantum mechanics we would denote thie function as $H(\sigma)$. The symbol $H$ stands for *Hamiltonian*, which is an operator which acts as a sum of the energies of the system. For the scope of this notebook, thinking of $Cost$ as any traditional cost function which we want to minimize will serve us equally as valuable.
# <a id="qubo_to_hamiltonian"></a>
#
# ## From QUBO to a Hamiltonian
# Now we must use our data to create the cost function defined above. To make a Hamiltonian that is recognizable by pyQuil, we must use the pyQuil `PauliTerm` object.
from pyquil.api import WavefunctionSimulator
from pyquil.paulis import PauliSum, PauliTerm
# A `PauliTerm` object can be quadratic or of order one. In the case of it being quadratic, it represents the relationship between any two samples of data. An order one `PauliTerm` would be an implementation of a bias term - a cost constraint which only affects one variable. Below we show some basic functionality of the `PauliTerm` object.
#Constructing a quadratic PauliTerm
i = 3
j = 6
print('Distance between samples %d and %d: %.3f' %(i,j,dist.values[i][j]))
# To create the quadratic term we multiply two Paulis together. Each `PauliTerm` has an accompanying coefficient which is also multiplied. For simplicity's sake, we include the pairwise distance as a coefficient of one factor, and make the other '1.0'.
term1 = PauliTerm("Z",i,dist.values[i][j])
term2 = PauliTerm("Z",j,1.0)
term = term1*term2
print(term)
# Feel free to play with the coefficient number of `term2` to see how it affects the output of the cell.
#
# For those new to quantum computing, you're likely wondering what the purpose of the letter 'Z' is. It indicates that this `PauliTerm` is a Z operator.
#
# You may also note that our sample numbers, $i=3$ and $j=6$, have found their way into the printed output. Including $i$ and $j$ in each `PauliTerm` tells pyQuil which samples or **qubits** the operation is applied to. That's right, in the QAOA setup we consider each datapoint to be mapped to a qubit. Thus, the above printed statement actually means _"apply a penalty of $Q$ should sample 3 and sample 6 be in the same class"_, where $Q$ is the coefficient of the operator product Z3*Z6. Said in a more quantum-intuitive sense: _"Apply a penalty of $Q$ should qubit 3 and qubit 6 both be found in the same spin state (spin up or spin down)"_.
#
# Thus, as QAOA tries to minimize the cost function, sample 3 and 6 will only appear in the same class if this configuration is optimal. The choice of our weights as the distances between the samples implies, that in a "good" configuration samples that lie far apart will end up in different classes.
#
# We can see now that to make the Hamiltonian for our system we must iterate over each distance in our distance matrix, and assign it within a `PauliTerm` as the interaction strength between the appropriate qubits. We can readily achieve this using the utility function `hamiltonian_from_distances`.
# +
from entropica_qaoa.utilities import hamiltonian_from_distances
hamiltonian = hamiltonian_from_distances(dist)
print(hamiltonian)
# -
# The above exercise brings up an important limitation to our present QAOA approach. The number of datapoints we are able to use is limited by the number of qubits we have available.
# <a id="apply_qaoa"></a>
#
# ## Minimize the Hamiltonian with QAOA
#
# Now that we have mapped the clustering problem to a Hamiltonian it is time to find the spin class assignments/spin configuration that minimizes our cost function. We do this using the QAOA algorithm. First we need to import the neccesary bits and pieces:
# +
# import the neccesary pyquil modules
from entropica_qaoa.qaoa.cost_function import QAOACostFunctionOnQVM, QAOACostFunctionOnWFSim
# import QAOAParameters
from entropica_qaoa.qaoa.parameters import ExtendedParams
# import an optimizer
from scipy.optimize import minimize
#Some utilities for time tracking and measuring our outcomes.
import time
from math import log
from entropica_qaoa.utilities import cluster_accuracy, max_probability_bitstring
# -
# Now we can set up the _hyperparameters_ (problem parameters that remain fixed for this problem instance):
timesteps = 3 # The QAOA p parameter
iters = 500 # Number of classical optimiser iterations
n_qubits = 10 #this number might be defined before your dataset - should equal the number of data points
#The hamiltonian is also a hyperparameter
# And of course also the parameters need to be chosen. In this QAOA run, we will use `ExtendedParameters`. This parameter class provides the most degrees of freedom for our optimizer to explore the energy landscape. Conversely, it also has the most parameters to optimize and thus will take longer to converge.
#
# To instantiate this parameter class, we need to pass in three separate lists of angles.
# - $\vec{\beta}$: every timestep requires $n_{qubits}$ beta rotations. Thus there are $n_{qubits}\times timesteps$ beta values.
# - $\vec{\gamma}_{pairs}$: there is a gamma rotation for every two-qubit interaction. A simple way to come up with this number is to measure the length of your Hamiltonian, subtracted by the number of single qubit bias terms in place.
# - $\vec{\gamma}_{singles}$: there is a gamma single rotation for each bias term included in the hamiltonian.
#
# We randomly generate these lists as their initial starting states are somewhat redunant. They will be optimized over 100s of iterations!
# +
betas = [round(val,1) for val in np.random.rand(timesteps*n_qubits)]
gammas_singles = [round(val,1) for val in np.random.rand(0)] #we don't want any bias terms
gammas_pairs = [round(val,1) for val in np.random.rand(timesteps*len(hamiltonian))]
hyperparameters = (hamiltonian, timesteps)
parameters = (betas, gammas_singles, gammas_pairs)
params = ExtendedParams(hyperparameters, parameters)
# -
# Before starting the simulator, make sure you are running Rigetti's QVM and Quil Compiler by running `qvm -S` and `quilc -S` in two open and disposable terminals
#
# Let's begin by running QAOA with $p=3$ timesteps, and a maximum of 500 optimiser Iterations.
# Set up the WavefunctionSimulator from pyQuil
sim = WavefunctionSimulator()
cost_function = QAOACostFunctionOnWFSim(hamiltonian,
params=params,
sim=sim,
enable_logging=True)
t0 = time.time()
res = minimize(cost_function, params.raw(), tol=1e-3, method='Cobyla',
options={"maxiter": iters})
print('Run complete!\n','Runtime:','{:.3f}'.format(time.time()-t0))
wave_func = cost_function.get_wavefunction(params.raw())
lowest = max_probability_bitstring(wave_func.probabilities())
true_clusters = [1 if val else 0 for val in labels]
acc = cluster_accuracy(lowest,true_clusters)
# We can analyze the optimizer to see whether or not our QAOA run converged. For the full message, run:
# ```python
# print(res)
# ```
print('Cost Function Value:', res.fun)
print('Converged?:',res.message)
# We can see we did not converge. Let's tighten up our operations by wrapping our QAOA runs in a function and increase the QAOA parameter $p$.
def run_qaoa(hamiltonian, params, timesteps, max_iters, init_state=None):
cost_function = QAOACostFunctionOnWFSim(hamiltonian,
params=params,
initial_state=init_state)
res = minimize(cost_function, params.raw(), tol=1e-3, method='Cobyla',
options={"maxiter" : max_iters})
return cost_function.get_wavefunction(params.raw()), res
# The cell below will take a couple of minutes to run:
t0 = time.time()
wave_func, res = run_qaoa(hamiltonian, params, timesteps=3, max_iters=1500)
print('Run complete\n','Runtime:','{:.3f}'.format(time.time()-t0))
lowest = max_probability_bitstring(wave_func.probabilities())
# +
true_clusters = [1 if val else 0 for val in labels]
acc = cluster_accuracy(lowest,true_clusters)
print('Cost Function Value:', res.fun)
print('Converged?:',res.message)
# -
# You should typically find that increasing the number of allowed iterations gives a more accurate answer. The precise numbers will depend on which Pokemons are randomly selected at the beginning.
| examples/6_ClusteringWithQAOA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7 (tensorflow)
# language: python
# name: tensorflow
# ---
# + [markdown] colab_type="text" id="klGNgWREsvQv"
# <a href="https://colab.research.google.com/github/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_12_04_atari.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="yzzbqc-JS2z9"
# # T81-558: Applications of Deep Neural Networks
# **Module 12: Reinforcement Learning**
# * Instructor: [<NAME>](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
# * For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# + [markdown] colab_type="text" id="pCrXGd_CS6eB"
# # Module 12 Video Material
#
# * Part 12.1: Introduction to the OpenAI Gym [[Video]](https://www.youtube.com/watch?v=_KbUxgyisjM&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_12_01_ai_gym.ipynb)
# * Part 12.2: Introduction to Q-Learning [[Video]](https://www.youtube.com/watch?v=A3sYFcJY3lA&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_12_02_qlearningreinforcement.ipynb)
# * Part 12.3: Keras Q-Learning in the OpenAI Gym [[Video]](https://www.youtube.com/watch?v=qy1SJmsRhvM&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_12_03_keras_reinforce.ipynb)
# * **Part 12.4: Atari Games with Keras Neural Networks** [[Video]](https://www.youtube.com/watch?v=co0SwPWoZh0&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_12_04_atari.ipynb)
# * Part 12.5: Application of Reinforcement Learning [[Video]](https://www.youtube.com/watch?v=1jQPP3RfwMI&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_12_05_apply_rl.ipynb)
#
# + [markdown] colab_type="text" id="pmDI-h7cI0tI"
# # Google CoLab Instructions
#
# The following code ensures that Google CoLab is running the correct version of TensorFlow, and has the necessary Python libraries installed.
# + colab={"base_uri": "https://localhost:8080/", "height": 136} colab_type="code" id="9KQhYThvTCQC" outputId="9106076b-1696-4878-8662-4eb4e40e28f2"
try:
from google.colab import drive
# %tensorflow_version 2.x
COLAB = True
print("Note: using Google CoLab")
except:
print("Note: not using Google CoLab")
COLAB = False
if COLAB:
# !sudo apt-get install -y xvfb ffmpeg
# !pip install -q 'gym==0.10.11'
# !pip install -q 'imageio==2.4.0'
# !pip install -q PILLOW
# !pip install -q 'pyglet==1.3.2'
# !pip install -q pyvirtualdisplay
# !pip install -q --upgrade tensorflow-probability
# !pip install -q tf-agents
# + [markdown] colab_type="text" id="lsaQlK8fFQqH"
# # Part 12.4: Atari Games with Keras Neural Networks
#
#
# The Atari 2600 is a home video game console from Atari, Inc. Released on September 11, 1977. It is credited with popularizing the use of microprocessor-based hardware and games stored on ROM cartridges instead of dedicated hardware with games physically built into the unit. The 2600 was bundled with two joystick controllers, a conjoined pair of paddle controllers, and a game cartridge: initially [Combat](https://en.wikipedia.org/wiki/Combat_(Atari_2600)), and later [Pac-Man](https://en.wikipedia.org/wiki/Pac-Man_(Atari_2600)).
#
# Atari emulators are popular and allow many of the old Atari video games to be played on modern computers. They are even available as JavaScript.
#
# * [Virtual Atari](http://www.virtualatari.org/listP.html)
#
# Atari games have become popular benchmarks for AI systems, particularly reinforcement learning. OpenAI Gym internally uses the [Stella Atari Emulator](https://stella-emu.github.io/). The Atari 2600 is shown in Figure 12.ATARI.
#
# **Figure 12.ATARI: The Atari 2600**
# 
#
# ### Actual Atari 2600 Specs
#
# * CPU: 1.19 MHz MOS Technology 6507
# * Audio + Video processor: Television Interface Adapter (TIA)
# * Playfield resolution: 40 x 192 pixels (NTSC). Uses a 20-pixel register that is mirrored or copied, left side to right side, to achieve the width of 40 pixels.
# * Player sprites: 8 x 192 pixels (NTSC). Player, ball, and missile sprites use pixels that are 1/4 the width of playfield pixels (unless stretched).
# * Ball and missile sprites: 1 x 192 pixels (NTSC).
# * Maximum resolution: 160 x 192 pixels (NTSC). Max resolution is only somewhat achievable with programming tricks that combine sprite pixels with playfield pixels.
# * 128 colors (NTSC). 128 possible on screen. Max of 4 per line: background, playfield, player0 sprite, and player1 sprite. Palette switching between lines is common. Palette switching mid line is possible but not common due to resource limitations.
# * 2 channels of 1-bit monaural sound with 4-bit volume control.
#
# ### OpenAI Lab Atari Pong
#
# OpenAI Gym can be used with Windows; however, it requires a special [installation procedure].(https://towardsdatascience.com/how-to-install-openai-gym-in-a-windows-environment-338969e24d30)
#
# This chapter demonstrates playing [Atari Pong](https://github.com/wau/keras-rl2/blob/master/examples/dqn_atari.py). Pong is a two-dimensional sports game that simulates table tennis. The player controls an in-game paddle by moving it vertically across the left or right side of the screen. They can compete against another player controlling a second paddle on the opposing side. Players use the paddles to hit a ball back and forth. The goal is for each player to reach eleven points before the opponent; you earn points when one fails to return it to the other. For the Atari 2600 version of Pong, a computer player (controlled by the 2600) is the opposing player.
#
# This section shows how to adapt TF-Agents to an Atari game. Some changes are necessary when compared to the pole-cart game presented earlier in this chapter. You can quickly adapt this example to any Atari game by simply changing the environment name. However, I tuned the code presented here for Pong, and it may not perform as well for other games. Some tuning will likely be necessary to produce a good agent for other games.
#
# We begin by importing the needed Python packages.
# + colab={} colab_type="code" id="sMitx5qSgJk1"
import base64
import imageio
import IPython
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
import PIL.Image
import pyvirtualdisplay
import tensorflow as tf
from tf_agents.agents.dqn import dqn_agent
from tf_agents.drivers import dynamic_step_driver
from tf_agents.environments import suite_gym, suite_atari
from tf_agents.environments import tf_py_environment, batched_py_environment
from tf_agents.eval import metric_utils
from tf_agents.metrics import tf_metrics
from tf_agents.networks import q_network
from tf_agents.policies import random_tf_policy
from tf_agents.replay_buffers import tf_uniform_replay_buffer
from tf_agents.trajectories import trajectory
from tf_agents.utils import common
from tf_agents.specs import tensor_spec
from tf_agents.trajectories import time_step as ts
# + colab={} colab_type="code" id="J6HsdS5GbSjd"
# Set up a virtual display for rendering OpenAI gym environments.
display = pyvirtualdisplay.Display(visible=0, size=(1400, 900)).start()
# + [markdown] colab_type="text" id="LmC0NDhdLIKY"
# ## Hyperparameters
#
# The hyperparameter names are the same as the previous DQN example; however, I tuned the numeric values for the more complex Atari game.
# + colab={} colab_type="code" id="HC1kNrOsLSIZ"
num_iterations = 250000
initial_collect_steps = 200
collect_steps_per_iteration = 10
replay_buffer_max_length = 100000
batch_size = 32
learning_rate = 2.5e-3
log_interval = 5000
num_eval_episodes = 5
eval_interval = 25000
# + [markdown] colab_type="text" id="XZUoWspvOJqB"
# The algorithm needs more iterations for an Atari game. I also found that increasing the number of collection steps helped the algorithm to train.
# + [markdown] colab_type="text" id="VMsJC3DEgI0x"
# ## Atari Environment's
#
# You must handle Atari environments differently than games like cart-poll. Atari games typically use their 2D displays as the environment state. AI Gym represents Atari games as either a 3D (height by width by color) state spaced based on their screens, or a vector representing the state of the gam's computer RAM. To preprocess Atari games for greater computational efficiency, we generally skip several frames, decrease the resolution, and discard color information. The following code shows how we can set up an Atari environment.
# + colab={} colab_type="code" id="pYEz-S9gEv2-"
#env_name = 'Breakout-v4'
env_name = 'Pong-v0'
#env_name = 'BreakoutDeterministic-v4'
#env = suite_gym.load(env_name)
# AtariPreprocessing runs 4 frames at a time, max-pooling over the last 2
# frames. We need to account for this when computing things like update
# intervals.
ATARI_FRAME_SKIP = 4
max_episode_frames=108000 # ALE frames
env = suite_atari.load(
env_name,
max_episode_steps=max_episode_frames / ATARI_FRAME_SKIP,
gym_env_wrappers=suite_atari.DEFAULT_ATARI_GYM_WRAPPERS_WITH_STACKING)
#env = batched_py_environment.BatchedPyEnvironment([env])
# + [markdown] colab_type="text" id="IIHYVBkuvPNw"
# We can now reset the environment and display one step. The following image shows how the Pong game environment appears to a user.
# + colab={"base_uri": "https://localhost:8080/", "height": 227} colab_type="code" id="RlO7WIQHu_7D" outputId="659aab33-6237-4ed1-8b94-ed6595fb7db1"
env.reset()
PIL.Image.fromarray(env.render())
# + [markdown] colab_type="text" id="b_lHcIcqUaqB"
# We are now ready to load and wrap the two environments for TF-Agents. The algorithm uses the first environment for evaluation, and the second to train.
# + colab={} colab_type="code" id="N7brXNIGWXjC"
train_py_env = suite_atari.load(
env_name,
max_episode_steps=max_episode_frames / ATARI_FRAME_SKIP,
gym_env_wrappers=suite_atari.DEFAULT_ATARI_GYM_WRAPPERS_WITH_STACKING)
eval_py_env = suite_atari.load(
env_name,
max_episode_steps=max_episode_frames / ATARI_FRAME_SKIP,
gym_env_wrappers=suite_atari.DEFAULT_ATARI_GYM_WRAPPERS_WITH_STACKING)
train_env = tf_py_environment.TFPyEnvironment(train_py_env)
eval_env = tf_py_environment.TFPyEnvironment(eval_py_env)
# + [markdown] colab_type="text" id="E9lW_OZYFR8A"
# ## Agent
#
# I used the following class, from TF-Agents examples, to wrap the regular Q-network class. The AtariQNetwork class ensures that the pixel values from the Atari screen are divided by 255. This division assists the neural network by normalizing the pixel values to between 0 and 1.
# + colab={} colab_type="code" id="EoLEdYvzUHeX"
class AtariQNetwork(q_network.QNetwork):
"""QNetwork subclass that divides observations by 255."""
def call(self,
observation,
step_type=None,
network_state=(),
training=False):
state = tf.cast(observation, tf.float32)
# We divide the grayscale pixel values by 255 here rather than storing
# normalized values beause uint8s are 4x cheaper to store than float32s.
state = state / 255
return super(AtariQNetwork, self).call(
state, step_type=step_type, network_state=network_state,
training=training)
# + [markdown] colab_type="text" id="l--Jj22eVRZD"
# Next, we introduce two hyperparameters that are specific to the neural network we are about to define.
# + colab={} colab_type="code" id="TgkdEPg_muzV"
fc_layer_params = (512,)
conv_layer_params=((32, (8, 8), 4), (64, (4, 4), 2), (64, (3, 3), 1))
q_net = AtariQNetwork(
train_env.observation_spec(),
train_env.action_spec(),
conv_layer_params=conv_layer_params,
fc_layer_params=fc_layer_params)
# + [markdown] colab_type="text" id="z62u55hSmviJ"
# Convolutional neural networks usually are made up of several alternating pairs of convolution and max-pooling layers, ultimately culminating in one or more dense layers. These layers are the same types as previously seen in this course. The QNetwork accepts two parameters that define the convolutional neural network structure.
#
# The more simple of the two parameters is **fc_layer_params**. This parameter specifies the size of each of the dense layers. A tuple specifies the size of each of the layers in a list.
#
# The second parameter, named **conv_layer_params**, is a list of convolution layers parameters, where each item is a length-three tuple indicating (filters, kernel_size, stride). This implementation of QNetwork supports only convolution layers. If you desire a more complex convolutional neural network, you must define your variant of the QNetwork.
#
# The QNetwork defined here is not the agent, instead, the QNetwork is used by the DQN agent to implement the actual neural network. This allows flexibility as you can set your own class if needed.
#
# Next, we define the optimizer. For this example, I used RMSPropOptimizer. However, AdamOptimizer is another popular choice. We also create the DQN and reference the Q-network we just created.
# + colab={} colab_type="code" id="jbY4yrjTEyc9"
optimizer = tf.compat.v1.train.RMSPropOptimizer(
learning_rate=learning_rate,
decay=0.95,
momentum=0.0,
epsilon=0.00001,
centered=True)
train_step_counter = tf.Variable(0)
observation_spec = tensor_spec.from_spec(train_env.observation_spec())
time_step_spec = ts.time_step_spec(observation_spec)
action_spec = tensor_spec.from_spec(train_env.action_spec())
target_update_period=32000 # ALE frames
update_period=16 # ALE frames
_update_period = update_period / ATARI_FRAME_SKIP
_global_step = tf.compat.v1.train.get_or_create_global_step()
agent = dqn_agent.DqnAgent(
time_step_spec,
action_spec,
q_network=q_net,
optimizer=optimizer,
epsilon_greedy=0.01,
n_step_update=1.0,
target_update_tau=1.0,
target_update_period=(
target_update_period / ATARI_FRAME_SKIP / _update_period),
td_errors_loss_fn=common.element_wise_huber_loss,
gamma=0.99,
reward_scale_factor=1.0,
gradient_clipping=None,
debug_summaries=False,
summarize_grads_and_vars=False,
train_step_counter=_global_step)
agent.initialize()
# + [markdown] colab_type="text" id="94rCXQtbUbXv"
# ## Metrics and Evaluation
#
# There are many different ways to measure the effectiveness of a model trained with reinforcement learning. The loss function of the internal Q-network is not a good measure of the entire DQN algorithm's overall fitness. The network loss function measures how close the Q-network was fit to the collected data and did not indicate how effective the DQN is in maximizing rewards. The method used for this example tracks the average reward received over several episodes.
# + colab={} colab_type="code" id="bitzHo5_UbXy"
def compute_avg_return(environment, policy, num_episodes=10):
total_return = 0.0
for _ in range(num_episodes):
time_step = environment.reset()
episode_return = 0.0
while not time_step.is_last():
action_step = policy.action(time_step)
time_step = environment.step(action_step.action)
episode_return += time_step.reward
total_return += episode_return
avg_return = total_return / num_episodes
return avg_return.numpy()[0]
# See also the metrics module for standard implementations of different metrics.
# https://github.com/tensorflow/agents/tree/master/tf_agents/metrics
# + [markdown] colab_type="text" id="NLva6g2jdWgr"
# ## Replay Buffer
#
# DQN works by training a neural network to predict the Q-values for every possible environment-state. A neural network needs training data, so the algorithm accumulates this training data as it runs episodes. The replay buffer is where this data is stored. Only the most recent episodes are stored, older episode data rolls off the queue as the queue accumulates new data.
# + colab={} colab_type="code" id="vX2zGUWJGWAl"
replay_buffer = tf_uniform_replay_buffer.TFUniformReplayBuffer(
data_spec=agent.collect_data_spec,
batch_size=train_env.batch_size,
max_length=replay_buffer_max_length)
# Dataset generates trajectories with shape [Bx2x...]
dataset = replay_buffer.as_dataset(
num_parallel_calls=3,
sample_batch_size=batch_size,
num_steps=2).prefetch(3)
# + [markdown] colab_type="text" id="rVD5nQ9ZGo8_"
# ## Random Collection
#
# The algorithm must prime the pump. Training cannot begin on an empty replay buffer. The following code performs a predefined number of steps to generate initial training data.
# + colab={} colab_type="code" id="wr1KSAEGG4h9"
random_policy = random_tf_policy.RandomTFPolicy(train_env.time_step_spec(),
train_env.action_spec())
def collect_step(environment, policy, buffer):
time_step = environment.current_time_step()
action_step = policy.action(time_step)
next_time_step = environment.step(action_step.action)
traj = trajectory.from_transition(time_step, action_step, next_time_step)
# Add trajectory to the replay buffer
buffer.add_batch(traj)
def collect_data(env, policy, buffer, steps):
for _ in range(steps):
collect_step(env, policy, buffer)
collect_data(train_env, random_policy, replay_buffer, steps=initial_collect_steps)
# + [markdown] colab_type="text" id="hBc9lj9VWWtZ"
# ## Training the agent
#
# We are now ready to train the DQN. This process can take many hours, depending on how many episodes you wish to run through. As training occurs, this code will update on both the loss and average return. As training becomes more successful, the average return should increase. The losses reported reflecting the average loss for individual training batches.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="0pTbJ3PeyF-u" outputId="4ddddecb-1297-4461-d907-b0cd12c9369d"
iterator = iter(dataset)
# (Optional) Optimize by wrapping some of the code in a graph using TF function.
agent.train = common.function(agent.train)
# Reset the train step
agent.train_step_counter.assign(0)
# Evaluate the agent's policy once before training.
avg_return = compute_avg_return(eval_env, agent.policy, num_eval_episodes)
returns = [avg_return]
for _ in range(num_iterations):
# Collect a few steps using collect_policy and save to the replay buffer.
for _ in range(collect_steps_per_iteration):
collect_step(train_env, agent.collect_policy, replay_buffer)
# Sample a batch of data from the buffer and update the agent's network.
experience, unused_info = next(iterator)
train_loss = agent.train(experience).loss
step = agent.train_step_counter.numpy()
if step % log_interval == 0:
print('step = {0}: loss = {1}'.format(step, train_loss))
if step % eval_interval == 0:
avg_return = compute_avg_return(eval_env, agent.policy, num_eval_episodes)
print('step = {0}: Average Return = {1}'.format(step, avg_return))
returns.append(avg_return)
# + [markdown] colab_type="text" id="68jNcA_TiJDq"
# ## Visualization
#
# The notebook can plot the average return over training iterations. The average return should increase as the program performs more training iterations.
# + colab={"base_uri": "https://localhost:8080/", "height": 300} colab_type="code" id="NxtL1mbOYCVO" outputId="a7387cb8-bf86-44ba-9e68-16c2fa48ceb6"
iterations = range(0, num_iterations + 1, eval_interval)
plt.plot(iterations, returns)
plt.ylabel('Average Return')
plt.xlabel('Iterations')
plt.ylim(top=10)
# + [markdown] colab_type="text" id="M7-XpPP99Cy7"
# ### Videos
#
# We now have a trained model and observed its training progress on a graph. Perhaps the most compelling way to view an Atari game's results is a video that allows us to see the agent play the game. The following functions are defined so that we can watch the agent play the game in the notebook.
# + colab={} colab_type="code" id="ULaGr8pvOKbl"
def embed_mp4(filename):
"""Embeds an mp4 file in the notebook."""
video = open(filename,'rb').read()
b64 = base64.b64encode(video)
tag = '''
<video width="640" height="480" controls>
<source src="data:video/mp4;base64,{0}" type="video/mp4">
Your browser does not support the video tag.
</video>'''.format(b64.decode())
return IPython.display.HTML(tag)
def create_policy_eval_video(policy, filename, num_episodes=5, fps=30):
filename = filename + ".mp4"
with imageio.get_writer(filename, fps=fps) as video:
for _ in range(num_episodes):
time_step = eval_env.reset()
video.append_data(eval_py_env.render())
while not time_step.is_last():
action_step = policy.action(time_step)
time_step = eval_env.step(action_step.action)
video.append_data(eval_py_env.render())
return embed_mp4(filename)
# + [markdown] colab_type="text" id="9c_PH-pX4Pr5"
# First, we will observe the trained agent play the game.
# + colab={"base_uri": "https://localhost:8080/", "height": 538} colab_type="code" id="owOVWB158NlF" outputId="177a310a-311d-4c90-bf5b-703fd517f941"
create_policy_eval_video(agent.policy, "trained-agent")
# + [markdown] colab_type="text" id="povaAOcZygLw"
# For comparison, we observe a random agent play. While the trained agent is far from perfect, it does outperform the random agent by a considerable amount.
# + colab={"base_uri": "https://localhost:8080/", "height": 538} colab_type="code" id="pJZIdC37yNH4" outputId="51282152-a92e-4296-c865-131c1bd6b0ca"
create_policy_eval_video(random_policy, "random-agent")
| t81_558_class_12_04_atari.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# import necessary packages
import os
import matplotlib.pyplot as plt
import geopandas as gpd
from descartes import PolygonPatch
import pandas as pd
import numpy as np
import seaborn as sns
import cbsodata
from mpl_toolkits.mplot3d import axes3d
buurten = gpd.read_file("data/neighborhoods/buurten.shp")
stations = gpd.read_file("data/ovhaltes/ovhaltes.shp")
buurten.plot()
stations.plot()
dfsjoin = gpd.sjoin(buurten,stations)
dfpivot = pd.pivot_table(dfsjoin,index='BUURTCODE',columns='TYPE',aggfunc={'TYPE':len})
dfpivot.columns = dfpivot.columns.droplevel()
dfpolynew = buurten.merge(dfpivot, how='left',on='BUURTCODE')
dfpolynew["bus_m"] = dfpolynew["bus"] / dfpolynew["OPPERVLAKT"] * 1000000
dfpolynew["bus_m"] = dfpolynew["bus_m"].fillna(0)
dfpolynew["trein_m"] = dfpolynew["trein"] / dfpolynew["OPPERVLAKT"] * 1000000
dfpolynew["trein_m"] = dfpolynew["trein_m"].fillna(0)
dfpolynew["metro_m"] = dfpolynew["metro"] / dfpolynew["OPPERVLAKT"] * 1000000
dfpolynew["metro_m"] = dfpolynew["metro_m"].fillna(0)
dfpolynew["tram_m"] = dfpolynew["tram"] / dfpolynew["OPPERVLAKT"] * 1000000
dfpolynew["tram_m"] = dfpolynew["tram_m"].fillna(0)
dfpolynew["total_m"] = dfpolynew["bus_m"] + dfpolynew["trein_m"] + dfpolynew["metro_m"] + dfpolynew["tram_m"]
dfpolynew.columns
df_short = dfpolynew[['BUURTCODE', 'BUURTNAAM','bus_m', 'trein_m', 'metro_m', 'tram_m', 'total_m']].set_index("BUURTCODE")
#The resulting dataset is a list of public transport stations per square kilometer of area for each of the buurten.
df_short
df_short.to_csv("output/ovstations.csv")
plt.plot(df_short["total_m"],"o")
| Python/Public Transport Stations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Network Traffic Dataset for malicious attack
# This dataset of network traffic flow is generated by CICFlowMeter, indicate whether the traffic is malicious attack (Bot) or not (Benign).
# CICFlowMeter - network traffic flow generator generates 69 statistical features such as Duration, Number of packets, Number of bytes, Length of packets, etc are also calculated separately in the forward and reverse direction.
# The output of the application is the CSV file format with two columns labeled for each flow, namely Benign or Bot. The dataset has been organized per day, for each day the raw data including the network traffic (Pcaps) and event logs (windows and Ubuntu event Logs) per machine are recorded. Download the dataset from the below wget command line provided and rename it as Network_Traffic.csv
# ! wget https://cse-cic-ids2018.s3.ca-central-1.amazonaws.com/Processed+Traffic+Data+for+ML+Algorithms/Friday-02-03-2018_TrafficForML_CICFlowMeter.csv
# ! mv Friday-02-03-2018_TrafficForML_CICFlowMeter.csv Network_Traffic.csv
# ## Create requirements.txt
# %%writefile requirements.txt
cloudpickle==1.1.1
pandas
scikit-learn==0.22.2
imblearn
joblib
numpy
seldon-core
tornado>=6.0.3
tensorflow==1.13.1
keras==2.2.4
google-cloud-storage
kubeflow-tfjob
azure==4.0.0
kubeflow-fairing
kubernetes==10.0.1
# ## Install the packages listed in requirements.txt using pip
# !pip install --user -r requirements.txt
# ## Restart the Kernal
from IPython.display import display_html
display_html("<script>Jupyter.notebook.kernel.restart()</script>",raw=True)
# # Configure docker credentials
#
# Get your docker registry user and password encoded in base64
#
# # echo -n USER:PASSWORD | base64
#
# Create a config.json file with your Docker registry url and the previous generated base64 string
# !echo -n USER:PASSWORD | base64
# %%writefile config.json
{
"auths": {
"https://index.docker.io/v1/": {
"auth": "<<Provide previous generated base64 string>>"
}
}
}
# ### Create a config-map in the namespace you're using with the docker config
# !kubectl create --namespace anonymous configmap docker-config --from-file=./config.json
# ## Dockerfile
# Update dockerfile base image with tensorflow-gpu image if device type is GPU
device_type="gpu" #Provide cpu or gpu
if device_type=="gpu":
# !sed -i "s/py3/gpu-py3/g" Dockerfile
# !cat Dockerfile
else:
# !cat Dockerfile
# ## Import Libraries
# +
from kubernetes.client import V1PodTemplateSpec
from kubernetes.client import V1ObjectMeta
from kubernetes.client import V1PodSpec
from kubernetes.client import V1Container
from kubernetes.client import V1VolumeMount
from kubernetes.client import V1Volume
from kubernetes.client import V1PersistentVolumeClaimVolumeSource
from kubernetes.client import V1ResourceRequirements
from kubeflow.tfjob import constants
from kubeflow.tfjob import utils
from kubeflow.tfjob import V1ReplicaSpec
from kubeflow.tfjob import V1TFJob
from kubeflow.tfjob import V1TFJobSpec
from kubeflow.tfjob import TFJobClient
import time
import re, os
import tensorflow as tf
import pandas as pd
import numpy as np
import logging
import sys
import importlib
# -
# ## Set up Kubeflow Fairing for training and predictions on On-premise
# Import the fairing library and configure the onprem environment that your training or prediction job will run in.
# +
from kubernetes import client as k8s_client
from kubernetes.client import rest as k8s_rest
from kubernetes import config as k8s_config
from kubernetes.client.rest import ApiException
from kubeflow import fairing
from kubeflow.fairing import utils as fairing_utils
from kubeflow.fairing import TrainJob
from kubeflow.fairing.preprocessors.function import FunctionPreProcessor
from kubeflow.fairing.preprocessors import base as base_preprocessor
from kubeflow.fairing.builders.cluster.cluster import ClusterBuilder
from kubeflow.fairing.cloud.k8s import MinioUploader
from kubeflow.fairing.builders.cluster.minio_context import MinioContextSource
from kubeflow.fairing import PredictionEndpoint
from kubeflow.fairing.kubernetes.utils import mounting_pvc
from kubeflow.fairing.kubernetes.utils import mounting_pvc
BackendClass = getattr(importlib.import_module('kubeflow.fairing.backends'), "KubernetesBackend")
namespace = fairing_utils.get_current_k8s_namespace()
print("Namespace : %s"%namespace)
# -
# ## Get minio-service cluster IP to upload docker build context
# #### Set DOCKER_REGISTRY
# The DOCKER_REGISTRY variable is used to push the newly built image.
# Please change the variable to the registry for which you've configured credentials.
# +
DOCKER_REGISTRY = "edward1723"
k8s_config.load_incluster_config()
api_client = k8s_client.CoreV1Api()
minio_service_endpoint = None
try:
minio_service_endpoint = api_client.read_namespaced_service(name='minio-service', namespace='kubeflow').spec.cluster_ip
except ApiException as e:
if e.status == 403:
logging.warning(f"The service account doesn't have sufficient privileges "
f"to get the kubeflow minio-service. "
f"You will have to manually enter the minio cluster-ip. "
f"To make this function work ask someone with cluster "
f"priveleges to create an appropriate "
f"clusterrolebinding by running a command.\n"
f"kubectl create --namespace=kubeflow rolebinding "
"--clusterrole=kubeflow-view "
"--serviceaccount=${NAMESPACE}:default-editor "
"${NAMESPACE}-minio-view")
logging.error("API access denied with reason: {e.reason}")
s3_endpoint = minio_service_endpoint
minio_endpoint = "http://"+s3_endpoint+":9000"
minio_username = "minio"
minio_key = "minio123"
minio_region = "us-east-1"
print(minio_endpoint)
minio_uploader = MinioUploader(endpoint_url=minio_endpoint, minio_secret=minio_username, minio_secret_key=minio_key, region_name=minio_region)
minio_context_source = MinioContextSource(endpoint_url=minio_endpoint, minio_secret=minio_username, minio_secret_key=minio_key, region_name=minio_region)
# -
# ## Build docker image
#
# Note: Upload dataset, Dockerfile, and network_model.py into notebook
# +
#output_map is a map of extra files to add to the notebook.
# It is a map from source location to the location inside the context.
output_map= {
"Dockerfile": "Dockerfile", #Dockerfile
"network_model.py":"network_model.py",
"Network_Traffic.csv": "Network_Traffic.csv"
}
preprocessor = base_preprocessor.BasePreProcessor(output_map=output_map)
preprocessor.preprocess()
builder = ClusterBuilder(registry=DOCKER_REGISTRY, preprocessor=preprocessor, context_source=minio_context_source)
builder.build()
# -
builder.image_tag
# ## Define TFJob Class to create training job
tfjob_name="network-fairing-tfjob"
class Tfjob(object):
def get_tfjob_params(self):
#Defining a Volume Mount
volume_mount = V1VolumeMount(name="nfsvolume", mount_path="/mnt/Model_Network")
#Defining a Persistent Volume Claim
persistent_vol_claim = V1PersistentVolumeClaimVolumeSource(claim_name="nfs1")
#Defining a Volume
volume = V1Volume(name="nfsvolume", persistent_volume_claim=persistent_vol_claim)
if device_type=="gpu":
#Defining a Container
container = V1Container(
name="tensorflow",
image=builder.image_tag,
volume_mounts=[volume_mount],
resources=V1ResourceRequirements(limits={"nvidia.com/gpu": 1})
)
else:
#Defining a Container
container = V1Container(
name="tensorflow",
image=builder.image_tag,
volume_mounts=[volume_mount]
)
return (volume_mount, persistent_vol_claim, volume, container)
def get_tfjob_nodes(self):
params = self.get_tfjob_params()
#Defining a Master
master = V1ReplicaSpec(replicas=1,
restart_policy="Never",
template=V1PodTemplateSpec(spec=V1PodSpec(
containers=[params[3]],
volumes=[params[2]])))
#Defining Worker Spec
worker = V1ReplicaSpec(replicas=1,
restart_policy="Never",
template=V1PodTemplateSpec(spec=V1PodSpec(
containers=[params[3]],
volumes=[params[2]],
)))
#Defining Parameter server(PS) Spec
ps = V1ReplicaSpec(replicas=1,
restart_policy="Never",
template=V1PodTemplateSpec(spec=V1PodSpec(
containers=[params[3]],
volumes=[params[2]])))
return (master,worker,ps)
def create_tfjob(self):
tfjob_node_spec = self.get_tfjob_nodes()
#Defining TFJob
tfjob = V1TFJob(
api_version="kubeflow.org/v1",
kind="TFJob",
metadata=V1ObjectMeta(name=tfjob_name,namespace=namespace),
spec=V1TFJobSpec(
clean_pod_policy="None",
tf_replica_specs={"PS":tfjob_node_spec[2],"Worker": tfjob_node_spec[1],"Master":tfjob_node_spec[0]}
)
)
#Creating TFJob
tfjob_client = TFJobClient()
tfjob_client.create(tfjob, namespace=namespace)
# ## Define Network class to be used by Kubeflow fairing
# ## ( Must necessarily contain train() and predict() methods)
class NetworkServe(object):
def __init__(self):
self.model=None
def train(self):
Tfjob().create_tfjob()
def predict(self,X,feature_names=None):
feature_col=['BwdIATMean', 'BwdIATTot', 'BwdPktLenMax', 'BwdPktLenMean', 'FlowDuration', 'FlowIATMean', 'FlowIATStd', 'FwdPSHFlags', 'FwdSegSizeMin', 'InitBwdWinByts']
model_input1=tf.train.Example()
for i in range(len(X)):
model_input1.features.feature[feature_col[i]].float_list.value.append(X[i])
path=os.path.join(os.getcwd(), "/mnt/Model_Network")
for dir in os.listdir(path):
if re.match('[0-9]',dir):
exported_path=os.path.join(path,dir)
break
# Open a Session to predict
with tf.Session() as sess:
tf.saved_model.loader.load(sess, [tf.saved_model.tag_constants.SERVING], exported_path)
model_input =model_input1
predictor= tf.contrib.predictor.from_saved_model(exported_path,signature_def_key='predict')
input_tensor=tf.get_default_graph().get_tensor_by_name("input_example_tensor:0")
model_input=model_input.SerializeToString()
output_dict= predictor({"examples":[model_input]})
sess.close()
response = output_dict.items()
print(response)
response1 = output_dict['class_ids']
return response1
# ## Train Network model remotely on Kubeflow
# Kubeflow Fairing packages the NetworkServe class, the training data, and the training job's software prerequisites as a Docker image. Then Kubeflow Fairing deploys and runs the training job on kubeflow.
train_job = TrainJob(NetworkServe, input_files=["Network_Traffic.csv", "requirements.txt"],
pod_spec_mutators = [mounting_pvc(pvc_name="nfs1", pvc_mount_path="/mnt/Model_Network")],
docker_registry=DOCKER_REGISTRY, backend=BackendClass(build_context_source=minio_context_source))
train_job.submit()
# ## Deploy the trained model to Kubeflow for predictions
# Kubeflow Fairing packages the NetworkServe class, the trained model, and the prediction endpoint's software prerequisites as a Docker image. Then Kubeflow Fairing deploys and runs the prediction endpoint on Kubeflow.
endpoint = PredictionEndpoint(NetworkServe, input_files=["Network_Traffic.csv", "requirements.txt"],
docker_registry=DOCKER_REGISTRY,
pod_spec_mutators = [mounting_pvc(pvc_name="nfs1", pvc_mount_path="/mnt/Model_Network")],
backend=BackendClass(build_context_source=minio_context_source))
endpoint.create()
# ## Wait for prediction pod ready state
# !kubectl get deploy -l fairing-deployer=serving -n anonymous
# ## Get prediction endpoint
endpoint.url
# ## Call the prediction endpoint
# Use the endpoint from previous cell
# ! curl -v http://fairing-service-pxnbw.anonymous.svc.cluster.local:5000/predict -H "Content-Type: application/x-www-form-urlencoded" -d 'json={"data":{"ndarray":[0.000000, 0.000000, 0.000000, 0.000000, 0.000005, 0.000000, 0.000000, 0.000000, 0.000000, 0.000004]}}'
# ## Clean up the prediction endpoint
# Delete the prediction endpoint created by this notebook.
endpoint.delete()
# ## Clean up TFjob
TFJobClient().delete(tfjob_name, namespace=namespace)
# ## Delete config.json and requirements.txt
# !rm -rf config.json requirements.txt
if device_type=="gpu":
# !sed -i "s/gpu-py3/py3/g" Dockerfile
# !cat Dockerfile
| apps/networking/network-traffic/onprem/fairing/Network-Classification-fairing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# # Kernels
#
# Some useful kernels.
# +
import numpy as np
import scipy.signal
import matplotlib.pyplot as plt
srate = 1000
spk_rate = 13.0 # Avg 30 spikes per second
tvec = np.arange(srate) / srate
spikeevents = (np.random.rand(srate) < (spk_rate / srate)).astype(np.float32)
spiketimes = tvec[spikeevents.nonzero()]
# +
from indl.misc.kernels import sskernel, Gauss
# Shimazaki et al. auto-kernel-width
kernel_width = sskernel(spiketimes - spiketimes[0], nbs=0)[2]
kernel_param = 1 / (2.0 * 2.7) * kernel_width
span_fac = 3.0
t_kern = np.arange(-span_fac * kernel_param, span_fac * kernel_param + (1 / srate), 1 / srate)
kernel = Gauss(t_kern, kernel_param)
spikerates = scipy.signal.convolve(spikeevents, kernel, 'same')
plt.subplot(3, 1, 1)
plt.plot(tvec, spikeevents)
plt.subplot(3, 1, 2)
plt.plot(t_kern, kernel)
plt.xlim([-0.5, 0.5])
plt.subplot(3, 1, 3)
plt.plot(tvec, spikerates)
# +
kernel_param = 0.050 # msec stdev
span_fac = 3.0 # How many stdevs wide the kernel should be. Too short will truncate kernel.
t_kern = np.arange(-span_fac * kernel_param, span_fac * kernel_param + (1 / srate), 1 / srate)
kernel = Gauss(t_kern, kernel_param)
spikerates = scipy.signal.convolve(spikeevents, kernel, 'same')
plt.subplot(3, 1, 1)
plt.plot(tvec, spikeevents)
plt.subplot(3, 1, 2)
plt.plot(t_kern, kernel)
plt.xlim([-0.5, 0.5])
plt.subplot(3, 1, 3)
plt.plot(tvec, spikerates)
# +
from indl.misc.kernels import Boxcar
kernel_param = 0.05 # The width of the rectangle in seconds
span_fac = np.sqrt(3.0)
kernel_param /= (2*np.sqrt(3.0))
t_kern = np.arange(-span_fac * kernel_param, span_fac * kernel_param + (1 / srate), 1 / srate)
kernel = Boxcar(t_kern, kernel_param)
spikerates = scipy.signal.convolve(spikeevents, kernel, 'same')
plt.subplot(3, 1, 1)
plt.plot(tvec, spikeevents)
plt.subplot(3, 1, 2)
plt.xlim([-0.5, 0.5])
plt.plot(t_kern, kernel)
plt.subplot(3, 1, 3)
plt.plot(tvec, spikerates)
# +
from indl.misc.kernels import Alpha
kernel_param = 0.03 # tau
kernel_param *= np.sqrt(2)
span_fac = 6.0
t_kern = np.arange(-span_fac * kernel_param, span_fac * kernel_param + (1 / srate), 1 / srate)
kernel = Alpha(t_kern, kernel_param)
spikerates = scipy.signal.convolve(spikeevents, kernel, 'same')
print(np.sum(spikeevents), np.mean(spikerates))
plt.subplot(3, 1, 1)
plt.plot(tvec, spikeevents)
plt.subplot(3, 1, 2)
plt.xlim([-0.5, 0.5])
plt.plot(t_kern, kernel)
plt.subplot(3, 1, 3)
plt.plot(tvec, spikerates)
# +
from indl.misc.kernels import Exponential
kernel_param = 0.05 # the time constant tau when the kernel reaches 1/e the maximum.
span_fac = 6.0
t_kern = np.arange(-span_fac * kernel_param, span_fac * kernel_param + (1 / srate), 1 / srate)
kernel = Exponential(t_kern, kernel_param)
spikerates = scipy.signal.convolve(spikeevents, kernel, 'same')
plt.subplot(3, 1, 1)
plt.plot(tvec, spikeevents)
plt.subplot(3, 1, 2)
plt.xlim([-0.5, 0.5])
plt.plot(t_kern, kernel)
plt.subplot(3, 1, 3)
plt.plot(tvec, spikerates)
| docs/Miscellaneous/kernels.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="8-duQOLXqtj9" colab_type="text"
# ## Probability Ratio Encoding
# + [markdown] id="KOEe_KX2qtkA" colab_type="text"
# 1. Probability of Survived based on Cabin --- **Categorical Feature**
# 2. Probability of Not Survived --- **1-prob(Survived)**
# 3. **prob(Survived)/prob(Not Survived)**
# 4. Dictonary to map cabin with probability.
# 5. Replace with the categorical feature.
# + id="DKZX-YWcqtkC" colab_type="code" colab={}
import pandas as pd
# + id="sE7tbNGEqtkQ" colab_type="code" colab={} outputId="204a2200-e4fa-4aa9-cca9-73dc5134ceb2"
df=pd.read_csv('titanic.csv',usecols=['Cabin','Survived'])
df.head()
# + [markdown] id="q1pF5T8KqtkY" colab_type="text"
# ### Replacing 'NAN' with 'Missing' Values
# + id="HYgbYu_zqtkb" colab_type="code" colab={} outputId="46ba9fcc-c802-4838-b321-eba13b6501a2"
df['Cabin'].fillna('Missing',inplace=True)
df.head()
# + id="QPVKyYoxqtkh" colab_type="code" colab={} outputId="8a3226c2-3aef-4eb6-d296-80e559f5406a"
df['Cabin'].unique()
# + id="qfx7jFqBqtko" colab_type="code" colab={} outputId="ef173022-018c-4374-88e3-fb09faa244b7"
df['Cabin']=df['Cabin'].astype(str).str[0]
df.head()
# + id="Y8LEUbd8qtkv" colab_type="code" colab={} outputId="35c37663-518d-4b0f-c48b-e3c9968beffa"
df.Cabin.unique()
# + id="I3csOTsKqtk1" colab_type="code" colab={}
prob_df=df.groupby(['Cabin'])['Survived'].mean()
prob_df = pd.DataFrame(prob_df)
# + id="qEtWq_coqtk7" colab_type="code" colab={} outputId="5dbfdb8f-5908-4bf5-a21a-3082c55f9077"
prob_df
# + id="T08MrRpmqtk_" colab_type="code" colab={}
prob_df['Died']= 1 - prob_df['Survived']
# + id="erViaohiqtlF" colab_type="code" colab={} outputId="90285664-babd-47f8-d7f0-edf9372fadb8"
prob_df.head()
# + id="2wMT2JZMqtlJ" colab_type="code" colab={} outputId="e54318fd-a890-465b-ff8c-20f96f686a24"
prob_df['Probability_ratio']=prob_df['Survived']/prob_df['Died']
prob_df.head()
# + id="Q7JxSdERqtlQ" colab_type="code" colab={} outputId="1bb0eb08-3120-40ed-c03d-e8fa7c5a3df7"
probability_encoded=prob_df['Probability_ratio'].to_dict()
probability_encoded
# + id="daPirTYEqtlV" colab_type="code" colab={} outputId="b2a971b4-3edf-4ca9-a310-1c7b4ebc8ef3"
df['Cabin_encoded']=df['Cabin'].map(probability_encoded)
df.head(15)
# + [markdown] id="Wu9CKiwyqtla" colab_type="text"
# ## Transformation of the Features.
#
# 1. Why Transformation of Features Are Required?
#
# * Linear Regression --- Gradient Descent --- **Global Minima**
# * Algorithms like KNN,K Means,Hierarichal Clustering --- **Eucledian Distance**
#
#
# 2. Every Point has some vector and direction.
#
#
# 3. Deep Learning Techniques(Standardization, Scaling --- 0-255 pixels)
#
# * ANN ---> Global Minima, Gradient Descent
# * CNN
# * RNN
# + [markdown] id="6iTm2AtEqtlb" colab_type="text"
# ### Types Of Transformation
#
# 1. Normalization And Standardization
#
#
# 2. Scaling to Minimum And Maximum values
#
#
# 3. Scaling To Median And Quantiles
#
#
# 4. Guassian Transformation
#
# * Logarithmic Transformation
# * Reciprocal Transformation
# * Square Root Transformation
# * Exponential Transformation
# * Box-Cox Transformation.
# + [markdown] id="qXMkfb4Dqtlc" colab_type="text"
# ### Standardization
#
# * We try to bring all the variables or features to a similar scale.
#
# * Standarization means centering the variable at zero.
#
# * **Z = (x-x_mean)/std**
#
# * Mean = 0, Standard Deviation = 1.
# + id="WDV4_haJqtle" colab_type="code" colab={} outputId="1da833a8-e09c-469d-e8f3-bea95dce4a93"
import pandas as pd
df=pd.read_csv('titanic.csv', usecols=['Pclass','Age','Fare','Survived'])
df.head()
# + id="-ivUXKHnqtlj" colab_type="code" colab={} outputId="1bc772e4-b67b-45df-f84e-6d5367f53345"
df.isnull().sum()
# + id="gqWQdEYIqtlo" colab_type="code" colab={}
df['Age'].fillna(df.Age.median(),inplace=True)
# + id="6vbaslZfqtlt" colab_type="code" colab={} outputId="1e47f432-5c14-4715-e77b-de399cc78c73"
df.isnull().sum()
# + [markdown] id="7XLeTi_Zqtlv" colab_type="text"
# #### Standarisation: We use the Standardscaler from sklearn Library.
# + id="4fwLAV4Cqtlx" colab_type="code" colab={}
from sklearn.preprocessing import StandardScaler
# + id="LnQvEjWUqtl0" colab_type="code" colab={}
scaler=StandardScaler()
# fit vs fit_transform
df_scaled=scaler.fit_transform(df)
# + id="Oc4XGhgNqtl6" colab_type="code" colab={} outputId="6102386d-7e7d-4f19-8745-b7a81e24e481"
df_scaled
# + id="9JpB7J42qtl-" colab_type="code" colab={} outputId="c9d3bf0f-5b92-48ea-d09c-7bb551b77fef"
pd.DataFrame(df_scaled)
# + id="tB0xkTWZqtmF" colab_type="code" colab={}
import matplotlib.pyplot as plt
# %matplotlib inline
# + id="sNaZF-ChqtmI" colab_type="code" colab={} outputId="8e98c4e3-dae0-4cf6-9c06-0c5cd2a3f7cd"
plt.hist(df_scaled[:,1],bins=20)
plt.xlabel('Pclass_Scaled')
plt.ylabel('Number of Points')
# + id="bq3JPsm2qtmN" colab_type="code" colab={} outputId="7131c04c-f71a-4fcc-98ef-95ce957f8b5e"
plt.hist(df_scaled[:,2],bins=20)
plt.xlabel('Age_Scaled')
plt.ylabel('Number of Points')
# + id="tDrrQIYCqtmR" colab_type="code" colab={} outputId="a6ca6436-889f-4487-f888-6ac51f3bc21b"
plt.hist(df_scaled[:,2],bins=20)
plt.xlabel('Fare_Scaled')
plt.ylabel('Number of Points')
# + [markdown] id="Sf6iRpVAqtmV" colab_type="text"
# * **If there are outliers, it will affect the standardization**
# + id="rmLwVzHhqtmW" colab_type="code" colab={} outputId="16d6fb53-8430-4fba-f91b-e429bfcc99b6"
plt.hist(df['Fare'],bins=20)
plt.xlabel('Fare_Not_Scaled')
plt.ylabel('Number of Points')
# + [markdown] id="oUcAvd0NqtmZ" colab_type="text"
# ### Min-Max Scaling (CNN) ---> Deep Learning Techniques
#
# * Min Max Scaling scales the values between 0 to 1.
#
#
# * X_scaled = (X - X.min / (X.max - X.min)
# + id="Y4EdhlX7qtma" colab_type="code" colab={} outputId="20486b68-1ed5-4493-d445-df39f807f87c"
from sklearn.preprocessing import MinMaxScaler
min_max=MinMaxScaler()
df_minmax=pd.DataFrame(min_max.fit_transform(df),columns=df.columns)
df_minmax.head()
# + id="S6GSBurUqtme" colab_type="code" colab={} outputId="beb8f215-829b-4fa9-caf2-fd7aa1bd2b56"
plt.hist(df_minmax['Pclass'],bins=20)
# + id="5yCIFogSqtmj" colab_type="code" colab={} outputId="52ed8374-3c90-4230-b682-45657f36ff60"
plt.hist(df_minmax['Fare'],bins=20)
# + id="Cya5RENMqtmp" colab_type="code" colab={} outputId="ebca6b22-5557-4351-9270-ff0b73f88d8f"
plt.hist(df_minmax['Age'],bins=20)
# + [markdown] id="fgBl2O2Qqtms" colab_type="text"
# ### Robust Scaler
#
# 1. It is used to scale the feature to median and quantiles.
#
#
# 2. Scaling using median and quantiles consists of substracting the median from all the observations, and then dividing by the interquantile difference.
#
#
# 3. The interquantile difference is the difference between the 75th and 25th quantile:
#
# * **IQR = 75th quantile - 25th quantile**
#
#
# 4. X_scaled = (X - X.median) / IQR
#
#
# 5. 0,1,2,3,4,5,6,7,8,9,10
#
# * **9** ---> 90 percentile ---> 90% of all values in this group is less than 9.
#
# * **1** ---> 10 precentile ---> 10% of all values in this group is less than 1.
# + id="PFnpuq8zqtmt" colab_type="code" colab={} outputId="6e90b3b5-ef14-421b-f613-27c416300d10"
from sklearn.preprocessing import RobustScaler
scaler=RobustScaler()
df_robust_scaler=pd.DataFrame(scaler.fit_transform(df),columns=df.columns)
df_robust_scaler.head()
# + id="Y7Ja-q-hqtmw" colab_type="code" colab={} outputId="6eae8cc1-f418-4262-adb3-6bb77b5e561b"
plt.hist(df_robust_scaler['Fare'],bins=20)
# + id="NanX0Ruhqtm0" colab_type="code" colab={} outputId="9c8f4eb6-dacc-4561-cf66-fb74aa3f5d57"
plt.hist(df_robust_scaler['Age'],bins=20)
# + id="labZ8ufJqtm4" colab_type="code" colab={} outputId="80589fd5-f364-444d-f55b-01dbe66c1d76"
plt.hist(df_robust_scaler['Pclass'],bins=20)
# + [markdown] id="oE4pNoZ_qtm9" colab_type="text"
# ### Guassian Transformation
#
# * Some machine learning algorithms like linear regression and logistic regression assume that the features are normally distributed ---> **Accuracy Performance increases when data is normally distributed**
#
# * Logarithmic transformation
# * Reciprocal transformation
# * Square Root transformation
# * Exponential transformation (more general, you can use any exponent)
# * BoxCox transformation
# + id="BeaMiJBOqtm9" colab_type="code" colab={} outputId="62d8e53d-8f90-4e62-af58-8fd8b6708dd3"
df=pd.read_csv('titanic.csv',usecols=['Age','Fare','Survived'])
df.head()
# + [markdown] id="5VX4WyRPqtnA" colab_type="text"
# #### Filling the missing "NAN" values with Median Values
# + id="V962iblHqtnA" colab_type="code" colab={}
df['Age']=df['Age'].fillna(df['Age'].median())
# + id="yYFhBPyCqtnD" colab_type="code" colab={} outputId="9a192584-66f5-4f0c-cc29-6145261d9f7f"
df.isnull().sum()
# + id="P5YZHBCoqtnR" colab_type="code" colab={}
import scipy.stats as stat
import pylab
# + [markdown] id="aSmnJTYkqtnT" colab_type="text"
# #### If you want to check whether feature is Guassian or Normal distributed we use ---> ***Q-Q plot***
# + id="hthz_30fqtnT" colab_type="code" colab={}
def plot_data(df, feature):
plt.figure(figsize=(10, 6))
plt.subplot(1, 2, 1) # ----> 1 row, 2 column and 1st index.
df[feature].hist()
plt.subplot(1, 2, 2) # ----> 1 row, 2 column and 2nd index.
stat.probplot(df[feature], dist='norm', plot=pylab)
plt.show()
# + id="6PwRHHCtqtnV" colab_type="code" colab={} outputId="c353659a-bc14-4254-8b66-edabb6519b93"
plot_data(df, 'Age')
# + [markdown] id="vDDFdiLqqtnY" colab_type="text"
# * If all the points are falling in the red line, then we can say that the feature is normally distributed.
# + id="ULVdFPREqtnY" colab_type="code" colab={} outputId="d9654a7e-5ab5-42ab-aa6f-ae5c54bc5934"
plot_data(df, 'Fare')
# + [markdown] id="Nss5iCRKqtnb" colab_type="text"
# #### Logarithmic Transformation
# + [markdown] id="YEt3paIpqtnb" colab_type="text"
# * **Logarithmic Transformation works best when your data is Right skewed or Left-skewed.**
# + id="S2MPgBmpqtnc" colab_type="code" colab={} outputId="3416984f-3621-4928-ecbd-d5e02d067461"
import numpy as np
df['Age_log'] = np.log(df['Age'])
plot_data(df, 'Age_log')
# + [markdown] id="IzYj4VK_qtnf" colab_type="text"
# #### Reciprocal Transformation
# + id="cgNdFIx6qtng" colab_type="code" colab={} outputId="8797f858-171b-4a64-ae43-93bc057c082f"
df['Age_reciprocal']=1/df.Age
plot_data(df,'Age_reciprocal')
# + [markdown] id="tRSsF79Bqtnj" colab_type="text"
# #### Square Root Transformation
# + id="L6pl2HjOqtnk" colab_type="code" colab={} outputId="3cf6cf20-68e8-45bf-f978-3d1a2e3f7a93"
df['Age_sqaure']=df.Age**(1/2)
plot_data(df,'Age_sqaure')
# + [markdown] id="W-rSksZFqtnm" colab_type="text"
# #### Exponential Transformation
# + id="epzONnegqtnm" colab_type="code" colab={} outputId="99c0a6ae-df9b-4e13-ac62-ff893fd84475"
df['Age_exponential']=df.Age**(1/1.2)
plot_data(df,'Age_exponential')
# + [markdown] id="3KxGMAQPqtnq" colab_type="text"
# #### Box-Cox Transformation
# + [markdown] id="Kll0lL0kqtnq" colab_type="text"
# * The Box-Cox transformation is defined as:
#
# * **T(Y)=(Y exp(λ)−1)/λ**
#
#
# * where Y is the response variable and λ is the transformation parameter. λ varies from -5 to 5. In the transformation, all values of λ are considered and the optimal value for a given variable is selected.
# + id="6QCOlDtLqtnr" colab_type="code" colab={} outputId="17cf8939-7204-46a0-9c15-b52a03a71c13"
stat.boxcox(df['Age'])
# + id="Mpi2et-iqtnv" colab_type="code" colab={}
df['Age_Boxcox'],parameters=stat.boxcox(df['Age'])
# + id="0sE1LYEIqtny" colab_type="code" colab={} outputId="41a9aad5-9c6a-4bb6-e560-187f4264d5b8"
parameters
# + id="l8WmPXsBqtn0" colab_type="code" colab={} outputId="7c6dfcc4-d40a-4789-8c28-8441bb7d38e7"
plot_data(df,'Age_Boxcox')
# + [markdown] id="yYNXfE4eqtn3" colab_type="text"
# #### 'Fare' Variable Plots
# + id="mhWAw3tWqtn3" colab_type="code" colab={} outputId="87c29da6-3ac7-4368-e298-0f7761fddfee"
plot_data(df,'Fare')
# + [markdown] id="j4vqAFnZqtn6" colab_type="text"
# #### Logarithmic Transformation of (x+1)
# + id="nxLzHC2Aqtn7" colab_type="code" colab={} outputId="b113a5c2-b98f-4fe0-94c9-506c3ab9e34f"
df['Fare_log']=np.log1p(df['Fare'])
plot_data(df,'Fare_log')
# + id="BhI-LaqZqtoA" colab_type="code" colab={} outputId="418bd587-0818-4adf-ae47-a43bd37089e6"
df['Fare_Boxcox'],parameters=stat.boxcox(df['Fare']+1)
plot_data(df,'Fare_Boxcox')
# + [markdown] id="Cn0Xos_gqtoD" colab_type="text"
# * **ANN require normalization as well.**
| Feature Engineering/Feature_Engineering_Part_5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Question 1:
# Make a regular expression to get all IP addresses from the below link and Extract the IP
# addresses.
# https://study-ccna.com/classes-of-ip-addresses/
# +
import requests, re
url = "https://study-ccna.com/classes-of-ip-addresses/"
r = requests.get(url)
data = r.text
ip = r'(\d{1,3}\.\d{1,3}\.\d{1,3}\.\d{1,3})'
list1_ip = re.findall(ip, data)
list1_ip = list(set(list1_ip))
for each in list1_ip:
print(each)
# -
| Day16 Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# <p>
# <img src="http://www.cerm.unifi.it/chianti/images/logo%20unifi_positivo.jpg"
# alt="UniFI logo" style="float: left; width: 20%; height: 20%;">
# <div align="right">
#
# <small>
# <NAME>, PhD.
# <br><br>
# May 27, 2020: init
# </small>
# </div>
# </p>
# <br>
# <br>
# <div align="center">
# <b>Abstract</b><br>
# Some tips, tricks and gotchas, in particular.
# </div>
# +
__AUTHORS__ = {'am': ("<NAME>",
"<EMAIL>",),
'mn': ("<NAME>",
"<EMAIL>",
"https://github.com/massimo-nocentini/",)}
__KEYWORDS__ = ['Python', 'Jupyter', 'gotchas', 'keynote',]
# -
# <center><img src="https://upload.wikimedia.org/wikipedia/commons/c/c3/Python-logo-notext.svg"></center>
# + [markdown] slideshow={"slide_type": "slide"}
# # Tips, tricks and gotchas
#
# This lecture addresses some gotchas that could arise in daily programming; moreover, at the beginning we will introduce some helpful objects that could make coding easier.
#
# First of all, some imports as usual:
# +
from collections import defaultdict, Counter
import matplotlib.pyplot as plt
# %matplotlib inline
plt.rcParams['figure.figsize'] = (8, 8)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## A grouping pattern, avoiding *quadratic* time
# -
# Assume to have two lists that have to be related in some way, namely using a predicate $P$. In the following example we want to build a list of all pairs (boy,girl) such that their names starts with the same letter. Here the input:
girls = ['alice', 'allie', 'bernice', 'brenda', 'clarice', 'cilly']
boys = ['chris', 'christopher', 'arald', 'arnold', 'bob']
# + [markdown] slideshow={"slide_type": "subslide"}
# the bad way, quadratic time:
# -
[(b, g) for b in boys for g in girls if b[0] == g[0]]
# + [markdown] slideshow={"slide_type": "subslide"}
# there is a better approach avoiding quadratic time, toward [`defaultdict`][dd]:
#
# [dd]:https://docs.python.org/3/library/collections.html#defaultdict-objects
# +
letterGirls = {}
for girl in girls:
letterGirls.setdefault(girl[0], []).append(girl)
[(b, g) for b in boys for g in letterGirls[b[0]]]
# + [markdown] slideshow={"slide_type": "subslide"}
# However there is an even better solution, as pointed out in the [example][e] subsection of the previous link: use `defaultdict` instead of repeating call `setdefault` method for each new key. From the official documentation:
#
# [e]:https://docs.python.org/3/library/collections.html#defaultdict-examples
# -
>>> s = [('yellow', 1), ('blue', 2), ('yellow', 3), ('blue', 4), ('red', 1)]
>>> d = defaultdict(list)
>>> for k, v in s:
... d[k].append(v)
...
>>> list(d.items())
[('blue', [2, 4]), ('red', [1]), ('yellow', [1, 3])]
# + [markdown] slideshow={"slide_type": "subslide"}
# ## The *Bunch* pattern
# -
# A very good book on algorithms implemented in Python is the one by <NAME>, https://www.apress.com/gp/book/9781484200568, with the companion Github repository https://github.com/apress/python-algorithms-14.
#
# Hetland, pag. 34, propose the following pattern to build a container of properties in order to avoid vanilla dict (adjusting from item 4.18 of <NAME>'s [*Python Cookbook*][cb]):
#
# [cb]:http://shop.oreilly.com/product/9780596007973.do
# + slideshow={"slide_type": "subslide"}
class Bunch(dict):
def __init__(self, *args, **kwds):
super(Bunch, self).__init__(*args, **kwds)
self.__dict__ = self
# + slideshow={"slide_type": "subslide"}
>>> T = Bunch
>>> t = T(left=T(left="a", right="b"), right=T(left="c"))
>>> t.left
# + slideshow={"slide_type": "subslide"}
>>> t.left.right
# -
>>> t['left']['right']
>>> "left" in t.right
"right" in t.right
# + [markdown] slideshow={"slide_type": "subslide"}
# However, inheriting from `dict` is discouraged by Alex:
#
# >A further tempting but not fully sound alternative is to have the Bunch class inherit
# from `dict`, and set attribute access special methods equal to the item access special
# methods, as follows:
#
# class DictBunch(dict):
# __getattr__ = dict.__getitem__
# __setattr__ = dict.__setitem__
# __delattr__ = dict.__delitem__
#
# >One problem with this approach is that, with this definition, an instance x of
# `DictBunch` has many attributes it doesn't really have, because it inherits all the
# attributes (methods, actually, but there's no significant difference in this context) of
# `dict`. So, you can’t meaningfully check `hasattr(x, someattr)` , as you could with the
# classes `Bunch` and `EvenSimplerBunch` (which sets the dictionary directly, without using `update`)
# previously shown, unless you can somehow rule
# out the value of someattr being any of several common words such as `keys` , `pop` ,
# and `get`. Python’s distinction between attributes and items is really a wellspring of clarity and
# simplicity. Unfortunately, many newcomers to Python wrongly believe that it would
# be better to confuse items with attributes, generally because of previous experience
# with JavaScript and other such languages, in which attributes and items are regularly
# confused. But educating newcomers is a much better idea than promoting item/
# attribute confusion.
# + [markdown] slideshow={"slide_type": "subslide"}
#
# Alex original definition reads as follows:
# -
class Bunch(object):
def __init__(self, **kwds):
self.__dict__.update(kwds)
# + [markdown] slideshow={"slide_type": "subslide"}
# It is interesting to observe that this idiom has been merged within the *standard library*, starting from Python **3.3**, as with the name of [`SimpleNamespace`][sn]:
#
# [sn]:https://docs.python.org/3/library/types.html#types.SimpleNamespace
# +
from types import SimpleNamespace
x, y = 32, 64
point = SimpleNamespace(datum=y, squared=y*y, coord=x)
point
# + slideshow={"slide_type": "subslide"}
point.datum, point.squared, point.coord
# -
[i for i in point]
# If you need `point` to be iterable use the structured object [`namedtuple`][nt] instead.
#
# [nt]:https://docs.python.org/3/library/collections.html#collections.namedtuple
# + [markdown] slideshow={"slide_type": "slide"}
# ## Python's `list.append` isn't Lisp's `cons`
# -
# Python `list` objects behave like `stack` objects, such that it is *cheap* to `append` and `pop` at the *top*, which is the *right* end. On the other hand, Lisp `pair` objects allows us to *easily* `cons` on the *beginning*, the very *opposite* direction.
# + slideshow={"slide_type": "subslide"}
def fast_countdown(count):
nums = []
for i in range(count):
nums.append(i)
nums.reverse()
return nums
def slow_countdown(count):
nums = []
for i in range(count):
nums.insert(0, i)
return nums
def printer(lst, chunk=10):
print("{}...{}".format(" ".join(map(str, lst[:chunk])),
" ".join(map(str, lst[-chunk:]))))
# + slideshow={"slide_type": "subslide"}
# %timeit nums = fast_countdown(10**5)
# -
# %timeit nums = slow_countdown(10**5)
# + [markdown] slideshow={"slide_type": "subslide"}
# Citing Hetland, pag 11:
#
# > Python lists aren’t really lists in the traditional computer science sense of the word, and that explains the puzzle of why append is so much more efficient than insert . A classical list - a so-called linked list - is implemented as a series of nodes, each (except for the last) keeping a reference to the next.
# The underlying implementation of Python’s list type is a bit different. Instead of several separate nodes
# referencing each other, a list is basically a single, contiguous slab of memory - what is usually known as an
# array. This leads to some important differences from linked lists. For example, while iterating over the contents
# of the list is equally efficient for both kinds (except for some overhead in the linked list), directly accessing an element at a given index is much more efficient in an array. This is because the position of the element can be
# calculated, and the right memory location can be accessed directly. In a linked list, however, one would have to
# traverse the list from the beginning.
# The difference we've been bumping up against, though, has to do with insertion. In a linked list, once you know
# where you want to insert something, insertion is cheap; it takes roughly the same amount of time, no matter how
# many elements the list contains. That's not the case with arrays: An insertion would have to move all elements
# that are to the right of the insertion point, possibly even moving all the elements to a larger array, if needed.
# A specific solution for appending is to use what’s often called a dynamic array, or vector. 4 The idea is to allocate an array that is too big and then to reallocate it in linear time whenever it overflows. It might seem that this makes the append just as bad as the insert. In both cases, we risk having to move a large number of elements.
# The main difference is that it happens less often with the append. In fact, if we can ensure that we always move
# to an array that is bigger than the last by a fixed percentage (say 20 percent or even 100 percent), the average
# cost, amortized over many appends, is constant.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### enhance with `deque` objects
# -
# `deque` implements *FIFO* queues: they are as cheap to append to the right as a normal `list`, but enhance it to *cheaply* insert on the *front* too.
# + slideshow={"slide_type": "subslide"}
from collections import deque
def enhanced_slow_countdown(count):
nums = deque()
for i in range(count):
nums.appendleft(i)
return nums
# -
# %timeit nums = enhanced_slow_countdown(10**5)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Hidden squares: concerning `list`s and `set`s
# +
from random import randrange
max_value = 10000
checks = 1000
L = [randrange(max_value) for i in range(checks)]
# + slideshow={"slide_type": "subslide"}
# %timeit [randrange(max_value) in L for _ in range(checks)]
# +
S = set(L) # convert the list to a set object.
# %timeit [randrange(max_value) in S for _ in range(checks)]
# + [markdown] slideshow={"slide_type": "subslide"}
# Hetland's words, pag. 35:
#
# >They're both pretty fast, and it might seem pointless to create a set from the list—unnecessary work, right? Well,
# it depends. If you're going to do many membership checks, it might pay off, because membership checks are linear
# for lists and constant for sets. What if, for example, you were to gradually add values to a collection and for each step check whether the value was already added? [...] Using a list would give you quadratic running time, whereas using a set would be linear. That’s a huge difference. **The lesson is that it's important to pick the right built-in data structure for the job.**
# + slideshow={"slide_type": "subslide"}
lists = [[1, 2], [3, 4, 5], [6]]
sum(lists, [])
# -
# Hetland, pag.36:
#
# >This works, and it even looks rather elegant, but it really isn't. You see, under the covers, the sum function doesn't know all too much about what you’re summing, and it has to do one addition after another. That way, you're right back at the quadratic running time of the += example for strings. Here's a better way: Just try timing both versions. As long as lists is pretty short, there won't be much difference, but it shouldn't
# take long before the sum version is thoroughly beaten.
# + slideshow={"slide_type": "subslide"}
res = []
for lst in lists:
res.extend(lst)
res
# -
# try to do that with more populated lists...
# + [markdown] slideshow={"slide_type": "subslide"}
# ### concerning `string`s
# -
def string_producer(length):
return ''.join([chr(randrange(ord('a'), ord('z'))) for _ in range(length)])
# +
# %%timeit
s = ""
for chunk in string_producer(10**5):
s += chunk
# + [markdown] slideshow={"slide_type": "-"}
# maybe some optimization is performed because `s` is a `string` object.
# + slideshow={"slide_type": "subslide"}
# %%timeit
chunks = []
for chunk in string_producer(10**5):
chunks.append(chunk)
s = ''.join(chunks)
# -
# a better approach using constant `append` to the top
# + slideshow={"slide_type": "subslide"}
# %timeit s = ''.join(string_producer(10**5))
# -
# maybe a little better since it doesn't loop with `for` explicitly.
# + [markdown] slideshow={"slide_type": "slide"}
# # Counting
# -
# ## Max permutation
#
# >Eight persons with very particular tastes have bought tickets to the movies. Some of them are happy with
# their seats, but most of them are not. Let’s say each of them has a favorite seat, and you want to find a way to let them switch seats to make as many people as possible happy with the result. However, all of them refuse to move to another seat if they can’t get their favorite.
#
# The following function `max_perm` computes the maximum permutation that can be applied given a desired one; namely,
# it produces a new permutation that moves as many elements as it can, in order to ensure the `one-to-one` property -- no one in the set points outside it, and each seat (in the set) is pointed
# to exactly once. It can be seen as a function that *fixes* a given permutation according to the required behavior.
# + slideshow={"slide_type": "subslide"}
def perm_isomorphism(M, domain):
iso = dict(enumerate(domain))
return [iso[M[i]] for i in range(len(M))]
def fix_perm(M, fix):
return [M[i] if i in fix else i for i in range(len(M))]
# + [markdown] slideshow={"slide_type": "subslide"}
# The following is a naive implementation, recursive but in $\mathcal{O}(n^{2})$, where $n$ is the permutation length.
# -
def naive_max_perm(M, A=None):
'''
Fix a permutation such that it is one-to-one and maximal, recursively.
consumes:
M - a permutation as a list of integers
A - a set of positions allowed to move
produces:
a set `fix` such that makes M maximal, ensuring to be one-to-one
'''
if A is None: A = set(range(len(M))) # init to handle first invocation, all elems can move
if len(A) == 1: return A # recursion base, unary perm can move, trivial
B = set(M[i] for i in A) # b in B iff b is desired by someone
C = A - B # c in C iff c isn't desired, so discard it
return naive_max_perm(M, A - C) if C else A # recur with desired position only
# + slideshow={"slide_type": "subslide"}
I = range(8) # the identity permutation
letters = "abcdefgh"
perm_isomorphism(I, letters)
# -
M = [2, 2, 0, 5, 3, 5, 7, 4]
perm_isomorphism(M, letters)
# + slideshow={"slide_type": "subslide"}
fix = naive_max_perm(M)
max_M = fix_perm(M, fix)
perm_isomorphism(max_M, letters)
# + [markdown] slideshow={"slide_type": "subslide"}
# Hetland, pag. 78:
#
# >The function `naive_max_perm` receives a set `A` of remaining people and creates a set `B` of seats that are pointed
# to. If it finds an element in `A` that is not in `B`, it removes the element and solves the remaining problem recursively. Let's use the implementation on our example, M = `[2, 2, 0, 5, 3, 5, 7, 4]`:
#
# -
naive_max_perm(M)
# + [markdown] slideshow={"slide_type": "subslide"}
# >So, a, c, and f can take part in the permutation. The others will have to sit in nonfavorite seats.
# The implementation isn't too bad. The handy set type lets us manipulate sets with ready-made high-level operations,
# rather than having to implement them ourselves. There are some problems, though. For one thing, we might want an
# iterative solution. [...] A worse problem, though, is that the algorithm is quadratic! (Exercise 4-10 asks you to show this.) The most wasteful operation is the repeated creation of the set B. If we could just keep track of which chairs are no longer pointed to, we could eliminate this operation entirely. One way of doing this would be to keep a count for each element. We could decrement the count for chair x when a person pointing to x is eliminated, and if x ever got a count of zero, both person and chair x would be out of the game.
# >>This idea of reference counting can be useful in general. It is, for example, a basic component in many systems
# for garbage collection (a form of memory management that automatically deallocates objects that are no longer useful). You'll see this technique again in the discussion of topological sorting.
# + [markdown] slideshow={"slide_type": "subslide"}
#
# >There may be more than one element to be eliminated at any one time, but we can just put any new ones we
# come across into a “to-do” list and deal with them later. If we needed to make sure the elements were eliminated in
# the order in which we discover that they’re no longer useful, we would need to use a first-in, first-out queue such as the deque class (discussed in Chapter 5). We don’t really care, so we could use a set, for example, but just appending to and popping from a list will probably give us quite a bit less overhead. But feel free to experiment, of course.
# + slideshow={"slide_type": "subslide"}
def max_perm(M):
n = len(M) # How many elements?
A = set(range(n)) # A = {0, 1, ... , n-1}
count = Counter(M) # desired positions by frequencies
Q = deque([i for i in A if not count[i]]) # useless elements
while Q: # While useless elts. left...
i = Q.pop() # get one of them
A.remove(i) # remove it from the maximal permutation
j = M[i] # get its desired position
count[j] -= 1 # and release it for someone else
if not count[j]: # if such position isn't desired anymore
Q.appendleft(j) # enqueue such position in order to discard it
return A
# + slideshow={"slide_type": "subslide"}
fix = max_perm(M)
max_M = fix_perm(M, fix)
perm_isomorphism(max_M, letters)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Counting Sort
# -
# Hetland, pag 85:
#
# >By default, I'm just sorting objects based on their values. By supplying a key function, you can sort by
# anything you’d like. Note that the keys must be integers in a limited range. If this range is $0\ldots k-1$, running time is then $\mathcal{O}(n + k)$. (Note that although the common implementation simply counts the elements and then figures out where to put them in `B`, Python makes it easy to just build value lists for each key and then
# concatenate them.) If several values have the same key, they'll end up in the original order with respect to
# each other. Sorting algorithms with this property are called *stable*.
# + slideshow={"slide_type": "subslide"}
def counting_sort(A, key=None, sort_boundary=None):
'''
Sorts the given collection A in linear time, assuming their elements are hashable.
This implementation implements a vanilla counting sort, working in linear time respect
iterable length and spacing between objects. It works best if elements are evenly, namely
*uniformly* distributed in the domain; on contrast, if they are sparse and concentrated
near accumulation points, traversing distances between them is time consuming.
If `sort_boundary` is instantiated to a float within [0,1], then the domain is ordered
using a classic loglinear algorithm before building the result.
'''
if key is None: key = lambda x: x
B, C = [], defaultdict(list)
for x in A:
C[key(x)].append(x)
domain = sorted(C) if sort_boundary and len(C) <= len(A)*sort_boundary \
else range(min(C), max(C)+1)
for k in domain:
B.extend(C[k])
return B
# + slideshow={"slide_type": "subslide"}
A = [randrange(50) for i in range(2*10**3)]
assert sorted(A) == counting_sort(A)
# + slideshow={"slide_type": "subslide"}
n, bins, patches = plt.hist(A, 10, facecolor='green', alpha=0.5)
plt.xlabel('elements'); plt.ylabel('frequencies'); plt.grid(True)
plt.show()
# + slideshow={"slide_type": "subslide"}
# %timeit counting_sort(A)
# -
# %timeit counting_sort(A, sort_boundary=1)
# + slideshow={"slide_type": "subslide"}
B = ([randrange(50) for i in range(10**3)] +
[10**4 + randrange(50) for i in range(10**3)])
# -
n, bins, patches = plt.hist(B, 100, facecolor='green', alpha=0.5)
plt.xlabel('elements'); plt.ylabel('frequencies'); plt.grid(True)
plt.show()
# + slideshow={"slide_type": "subslide"}
assert sorted(B) == counting_sort(B)
# -
# %timeit counting_sort(B)
# %timeit counting_sort(B, sort_boundary=1/8)
| UniFiCourseSpring2020/gotchas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.pyplot import imshow
import matplotlib.patches as patches
# %matplotlib inline
import json
import glob
import numpy as np
from PIL import Image
import rasterio
from rasterio.mask import mask
from rasterio.plot import reshape_as_image
#from scipy.misc import imsave
from imageio import imwrite
from sklearn.cluster import DBSCAN
from shapely.geometry import box
Image.MAX_IMAGE_PIXELS = None
# +
#To do: move these functions to .py files
def convert_coords(image, label, x, y):
'''
convert from geographic coordinates to image-scaled coordinates, relative to bottom left of image
'''
# get coordinates defining extent of input image and labels
bbox = image.bounds #features.bounds(label) #bounding box of image - this is different to bounding box of labels
x_min, y_min, x_max, y_max = bbox[0], bbox[1], bbox[2], bbox[3] #label['bbox'][0], label['bbox'][1], label['bbox'][2], label['bbox'][3]
# get distance of point x,y from image origin
x_relative = x - x_min
y_relative = y - y_min
# get scaling factors
width = x_max - x_min
height = y_max - y_min
#(width_pix, height_pix) = image.size
(height_pix, width_pix) = image.shape
x_pix = x_relative/width * width_pix
y_pix = y_relative/height * height_pix
return x_pix, y_pix
# -
def save_files(image, label, info, output_dir, input_name):
'''
for each cluster of bounding boxes, save a 512x512 image chip as .png
along with bounding box labels in YOLO format as .json
'''
# initialise a figure to visualise output tiles
fig = plt.figure(figsize=(20, 100))
n_tiles = len(info.keys())
for i,k in enumerate(info.keys()):
# get centre of each bounding box cluster
x, y = info[k]['centre'][0], info[k]['centre'][1] # in pixels, with origin in lower left
# define coordinates for cropping
# set limits since tile boundaries cannot exceed image boundaries
#width, height = image.getbbox()[2], image.getbbox()[3] #2,3 are right and lower bounds
width, height = image.bounds[2], image.bounds[1]
left, top, right, bottom = x-256, (height-y)-256, x+256, (height-y)+256 # in accordance with PIL library: in pix, origin top left
left_lim, top_lim = max(0,int(left)), max(0,int(top))
right_lim, bottom_lim = min(width, int(right)), min(height, int(bottom))
# crop and save image tiles
image_name = '{}/{}_{}.png'.format(output_dir, input_name, k)
#image_str = file_path + '/' + image_name
# it goes left-top-right-bottom where bottom=top+height (i.e. the origin is in the top left corner)
#image_tile = image.crop([left_lim, top_lim, right_lim, bottom_lim])
crop_bbox = [{'type': 'Polygon', 'coordinates': [[[left_lim, bottom_lim], [left_lim, top_lim], [right_lim, top_lim], [right_lim, bottom_lim], [left_lim, bottom_lim]]]}]
print(crop_bbox)
print(image.transform)
image_tile, image_tile_transform = mask(image, crop_bbox, crop=True)
#imwrite(image_name, image_tile)
ax = fig.add_subplot(n_tiles, 4, i+1)
ax.imshow(image_tile)
# save label file
#file = open(image_name.replace('.png', '.txt'), 'a')
for j, box in enumerate(info[k]['object_boxes']):
# get coordinates of lower left (x1) and upper right (x3) corner of bounding box
[[x1, y1], [x3, y3]] = box # in px, origin in lower left
# define bounding box coordinates relative to boundaries of current image tile
x1_rel, x3_rel = x1-left_lim, x3-left_lim
y1_rel, y3_rel = (height-y1)-top_lim, (height-y3)-top_lim
# get some scaling factors to convert from image to tile coordinates
tile_width = right_lim - left_lim
tile_height = top_lim - bottom_lim
# define bounding box centre & width
box_centre_x = (x1_rel+x3_rel)//2
box_centre_y = (y1_rel+y3_rel)//2
box_width = x3-x1
box_height = y1-y3
# add bounding boxes to tile subplot
rect = patches.Rectangle((x1_rel, y1_rel), box_width, box_height, linewidth=1, edgecolor='r', facecolor='none')
ax.add_patch(rect)
title = str(i+1)
ax.set_title(title)
# write label to .txt file
## 0 means first object i.e. whale
## (add a line here for multiple categoties - if info[k]['name'] = 'ship': lab = ...)
#
#lab = '0 {} {} {} {}\n'.format(abs(box_centre_x/tile_width), abs(box_centre_y/tile_height), abs(box_width/tile_width), abs(box_height/tile_height))
#file.write(lab)
#file.close()
plt.show()
# +
#Data paths:
liebre1_image_path = "/gws/nopw/j04/ai4er/users/kmgreen/data/liebrepansharp.tif"
liebre1_rgb_path = "/gws/nopw/j04/ai4er/users/kmgreen/data/liebre1pansharpRGB.tif"
liebre1_nostretch_image_path = "/gws/nopw/j04/ai4er/users/kmgreen/data/liebre1_nostretch.tif"
liebre1_points_shp_path = "/gws/nopw/j04/ai4er/users/kmgreen/data/liebre_points_2015_image.shp"
liebre1_boxes_shp_path = "/gws/nopw/j04/ai4er/users/kmgreen/data/liebre1_boxes.shp"
# -
#checking image path exists
import os.path
from os import path
print(path.exists(liebre1_image_path))
print(path.isfile(liebre1_image_path))
print(path.getsize(liebre1_image_path))
print(path.isabs(liebre1_image_path))
print(path.abspath(liebre1_image_path))
#Export shapefiles to a geojson with bounding box
import geopandas
#liebre_bbox = (778562.590800, 3063219.728100, 796312.956300, 3072412.944500)
image_bbox = (777470.0,3057298.0,797147.5,3073682.0)
liebre_box_shpfile = geopandas.read_file(liebre1_boxes_shp_path, bbox=image_bbox)
#liebre_box_shpfile.to_file('/gws/nopw/j04/ai4er/users/kmgreen/data/liebre1_boxes.geojson', driver='GeoJSON')
liebre1_box_path = "/gws/nopw/j04/ai4er/users/kmgreen/data/liebre1_boxes.geojson"
#look at geojson point file
boxes = geopandas.read_file(liebre1_box_path)
boxes.head()
boxes.crs
#checking
from rasterio import features
from rasterio.plot import show
with open(liebre1_box_path, 'r') as f:
label = json.load(f)
print(features.bounds(label))
#print(label)
first = label['features'][0]
print(first)
print(first['geometry']['coordinates'][0][0][0])
# visualise input image file + bounding boxes from label file
with open(liebre1_box_path, 'r') as f:
label = json.load(f)
#image = Image.open(liebre1_image_path)
image = rasterio.open(liebre1_image_path)
# create figure
fig1 = plt.figure(figsize=(16, 8))
ax1 = fig1.add_subplot(111, aspect='equal')
plt.imshow(np.flipud(image.read(1)), origin='lower') # flip because imshow defines upper left as origin
# plot bounding boxes
for object in label['features']:
# get origin and dimensions of each bounding box
bottom_left_unconv = object['geometry']['coordinates'][0][0]
top_right_unconv = object['geometry']['coordinates'][0][2]
bottom_left = convert_coords(image, label, bottom_left_unconv[0], bottom_left_unconv[1])
#print(bottom_left)
top_right = convert_coords(image, label, top_right_unconv[0], top_right_unconv[1])
width = top_right[0] - bottom_left[0]
height = top_right[1] - bottom_left[1]
# add bounding box to figure
rect = patches.Rectangle(bottom_left, width, height, linewidth=1, edgecolor='r', facecolor='none')
ax1.add_patch(rect)
plt.show
def read_coords(label):
coords, centres = [], []
for object in label['features']:
# get coordinates of lower left & upper right corners of each bounding box
[[x1, y1], [x3, y3]] = object['geometry']['coordinates'][0][0], object['geometry']['coordinates'][0][2]
coords.append([[x1, y1], [x3, y3]])
# store centre coordinate of each bounding box for clustering
centres.append([(x1+x3)//2, (y1+y3)//2])
return np.array(coords), centres
file_save_path = "/gws/nopw/j04/ai4er/users/kmgreen/data/cropped_images"
with open(liebre1_box_path, 'r') as f:
label = json.load(f)
#label['liebre1_image_path'] = image_file #What is this for?
#image = Image.open(liebre1_image_path)
image = rasterio.open(liebre1_image_path)#.read()
#image = reshape_as_image(image1)
print(type(image))
coords, centres = read_coords(label)
# convert bounding box coordinates from geographic to image-scaled
centres_converted = np.array([convert_coords(image, label, point[0], point[1]) for point in centres])
coords_converted = np.array([[convert_coords(image, label, point[0][0], point[0][1]),
convert_coords(image, label, point[1][0], point[1][1])]
for point in coords
])
## DB-Scan algorithm for clustering ##
eps = 250 # threshold distance between two points to be in the same 'neighbourhood'
dbscan = DBSCAN(min_samples=1, eps=eps)
y = dbscan.fit_predict(centres_converted)
# storing coordinates of clusters, relative to boundaries of image (not tile)
info = {}
for i in range(y.max()+1):
# calculate the max and min coords of all the bounding boxes in the cluster
box_centres = centres_converted[np.where(y==i)[0]]
min_x, max_x = box_centres[:, 0].min(), box_centres[:, 0].max()
min_y, max_y = box_centres[:, 1].min(), box_centres[:, 1].max()
# assign each cluster of objects as an item
item = {}
item['centre'] = [(min_x+max_x)//2, (min_y+max_y)//2]
item['object_boxes'] = coords_converted[np.where(y==i)[0]].tolist()
item['name'] = "whale"
info[i] = item
# add a line here to generalize to multiple categories:
# if label['features']['NumShip'] == ... :
save_files(image, label, info, file_save_path, 'liebre1')
#from rasterio.transform import Affine
#print(image.transform)
#print(image.transform*(0,0))
#transform = Affine(0.5,0.0, 0,
#0.0 , -0.5, 0)
#print(transform*(0,0))
#with rasterio.open(
#liebre1_new_image_path,
#'w',
#driver = 'GTiff',
#height = image.shape[0],
#width = image.shape[1],
#counts = 4,
#transform = transform,
#) as dst:
# dst.write(image)
#Try loading in as an array and converting to PIL
image = rasterio.open(liebre1_image_path).read()
image_array = reshape_as_image(image)
image_array.shape
import cv2 as cv
#imageRGB = cv.cvtColor(image_array, cv.COLOR_BGR2RGB)
#imageRGB.shape
#PIL_image = Image.fromarray(np.uint8(image_array)).convert('RGB')
#PIL_image = Image.fromarray(np.uint8(image_array), 'RGBA')
PIL_image = Image.fromarray(np.uint8(imageRGB))
plt.imshow(PIL_image)
PIL_image.size
#Try loading with open cv and converting to PIL
image = cv.imread('/gws/nopw/j04/ai4er/users/kmgreen/data/liebrepansharp.tif', cv.IMREAD_UNCHANGED)
print(image)
image.shape
#from skimage import io
import tifffile as tiff
a = tiff.imread(liebre1_image_path)
print(a.dtype)
print(a)
#print(a)
plt.imshow(a, cmap='gray', vmin=0, vmax=65535)
PIL_image = Image.fromarray(np.uint8(a))
plt.imshow(PIL_image)
# ## Try rgb image
Image.open(r'/gws/nopw/j04/ai4er/users/kmgreen/data/liebre1pansharpRGB.tif')
#Try loading with open cv and converting to PIL
image = cv.imread('/gws/nopw/j04/ai4er/users/kmgreen/data/liebre1pansharpRGB.tif', cv.IMREAD_UNCHANGED)
print(image)
rgb = tiff.imread('/gws/nopw/j04/ai4er/users/kmgreen/data/liebre1pansharpRGB.tif')
print(rgb.dtype)
print(rgb.shape)
PIL_rgb = Image.fromarray(rgb)
plt.imshow(PIL_rgb)
| notebooks/exploratory/preprocessing0.1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Make a grid out of Image pixels
import ee
ee.Initialize()
from geetools import tools
from ipygee import *
# ### Geometry
p = ee.Geometry.Point([-71.33972167968751, -42.737619925503054])
aoi = p.buffer(8000).bounds()
# ### Image
i = ee.Image('COPERNICUS/S2/20181122T142749_20181122T143353_T18GYT').clip(aoi)
# ### Make Grid
grid = tools.image.toGrid(i, 3, geometry=aoi)
# ### Show on Map
Map = Map()
Map.show()
Map.addLayer(i, {'bands':['B2'], 'min':0, 'max':3000}, 'Image')
Map.addLayer(grid, None, 'Grid')
eprint(ee.Feature(grid.first()).geometry().projection())
eprint(i.select(0).projection())
Map.addLayer(ee.Feature(grid.first()).geometry().transform(i.select(0).projection(), 1), None, 'reprojected')
Map.addLayer(ee.Feature(grid.first()), None, 'first')
eprint(Map.getObject('reprojected'))
| notebooks/image/toGrid.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# __Objetivos__:
# - entender como o perceptron funciona intuitivamente, tanto em regressão quanto em classificação.
# # Sumário
# [Regressão](#Regressão)
#
# [Classificação](#Classificação)
# - [Porta AND](#Porta-AND)
# - [Porta OR](#Porta-OR)
# - [Porta XOR](#Porta-XOR)
# # Imports
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import ipywidgets as wg
from ipywidgets import interactive, fixed
# %matplotlib inline
# jupyter nbextension enable --py widgetsnbextension --sys-prefix
# restart jupyter notebook
# -
# # Regressão
df = pd.read_csv('data/medidas.csv')
print(df.shape)
df.head(10)
# +
x = df.Altura
y = df.Peso
plt.figure()
plt.scatter(x, y)
plt.xlabel('Altura')
plt.ylabel('Peso')
# -
def plot_line(w, b):
plt.figure(0, figsize=(20,4))
plt.subplot(1,3,3)
plt.scatter(x, y)
y_pred = x*w + b
plt.plot(x, y_pred, c='red')
plt.xlim(140, 210)
plt.ylim(40, 120)
plt.subplot(1,3,2)
x_ = np.array([0, x.max()])
y_ = x_*w + b
plt.scatter(x, y)
plt.plot(x_, y_, c='red')
plt.xlim(0, 210)
plt.ylim(-160, 120)
plt.subplot(1,3,1)
mse = np.mean((y - y_pred)**2)
loss.append(mse)
plt.plot(loss)
plt.title('Loss')
plt.show()
# +
loss = []
interactive_plot = interactive(plot_line, w=(1, 1.5, 0.01), b=(-200, 0, 1))
output = interactive_plot.children[-1]
output.layout_height = '350px'
interactive_plot
# +
from sklearn.linear_model import LinearRegression
reg = LinearRegression()
reg.fit(x.values.reshape(-1,1), y)
print("w: {:.2f} \nb: {:.2f}".format(reg.coef_[0], reg.intercept_))
# -
# # Classificação
def plot_line(w1, w2, b):
x1, x2 = np.meshgrid(np.linspace(0,1,100), np.linspace(0,1,100))
x_mesh = np.array([x1.ravel(), x2.ravel()]).T
plt.figure(0, figsize=(10,4))
plt.subplot(1,2,2)
plt.scatter(x[:,0], x[:,1], c=y.ravel(), s=100, cmap='bwr')
y_mesh = np.dot(x_mesh, np.array([w1, w2]).T) + b
y_mesh = np.where(y_mesh <= 0, 0, 1)
plt.contourf(x1, x2, y_mesh.reshape(x1.shape), cmap='bwr')
y_pred = np.dot(x, np.array([w1, w2]).T) + b
y_bin = np.where(y_pred <= 0, 0, 1)
print('{0} => {1}'.format(y_pred, y_bin))
plt.subplot(1,2,1)
mse = np.mean((y.ravel() - y_bin)**2)
loss.append(mse)
plt.plot(loss)
plt.title('Loss')
plt.show()
# ### Porta AND
# +
x = np.array([[0, 0], [0, 1], [1, 0], [1, 1]])
y = np.array([[0, 0, 0, 1]]).T
print(x, y, sep='\n')
# -
plt.scatter(x[:,0], x[:,1], c=y.ravel(), s=50, cmap='bwr')
# +
loss = []
interactive_plot = interactive(plot_line, w1=(-1,1,0.01), w2=(-1,1,0.01), b=(-1.5, 1.5, 0.01))
interactive_plot
# -
# ### Porta OR
# +
x = np.array([[0, 0], [0, 1], [1, 0], [1, 1]])
y = np.array([[0, 1, 1, 1]]).T
print(x, y, sep='\n')
# -
plt.scatter(x[:,0], x[:,1], c=y.ravel(), s=50, cmap='bwr')
# +
loss = []
interactive_plot = interactive(plot_line, w1=(-1,1,0.01), w2=(-1,1,0.01), b=(-1.5, 1.5, 0.01))
interactive_plot
# -
# ### Porta XOR
# +
x = np.array([[0, 0], [0, 1], [1, 0], [1, 1]])
y = np.array([[0, 1, 1, 0]]).T
print(x, y, sep='\n')
# -
plt.scatter(x[:,0], x[:,1], c=y.ravel(), s=50, cmap='bwr')
# +
loss = []
interactive_plot = interactive(plot_line, w1=(-1,1,0.01), w2=(-1,1,0.01), b=(-1.5, 1.5, 0.01))
interactive_plot
| Perceptron_Intuicao.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import molsysmt as msm
import openexplorer as oe
import numpy as np
from simtk import unit
from simtk.openmm import app
# # Explorer
# +
modeller = msm.convert('alanine_dipeptide.pdb', to_form='openmm.Modeller')
topology = modeller.topology
positions = modeller.positions
forcefield = app.ForceField('amber10.xml', 'amber10_obc.xml')
system = forcefield.createSystem(topology, constraints=app.HBonds, nonbondedMethod=app.NoCutoff)
# -
explorer = oe.Explorer(topology, system, platform='CUDA')
explorer.set_coordinates(positions)
explorer.get_energy()
explorer.get_gradient()
explorer.get_hessian()
explorer.quench(minimizer='L-BFGS', tolerance=1.0*unit.kilojoule_per_mole/unit.nanometer)
explorer.get_gradient()
explorer.quench(minimizer='FIRE', tolerance=1.0*unit.kilojoule_per_mole/unit.nanometer)
explorer.get_gradient()
explorer.quench(minimizer='gradient_descent', tolerance=0.1*unit.kilojoule_per_mole)
explorer.get_gradient()
| docs/_build/.doctrees/nbsphinx/contents/Explorer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Iris (Python 2)
# language: python
# name: iris_python2
# ---
# # Glider
# +
import iris
iris.FUTURE.netcdf_promote = True
url = ('http://tds.marine.rutgers.edu:8080/thredds/dodsC/'
'cool/glider/mab/Gridded/20130911T000000_20130920T000000_gp2013_modena.nc')
glider = iris.load(url)
lon = glider.extract_strict('Longitude').data
lat = glider.extract_strict('Latitude').data
glider = glider.extract_strict('Temperature')
depth = glider.coord('depth').points
# +
import numpy as np
import numpy.ma as ma
import seawater as sw
import matplotlib.pyplot as plt
from scipy.interpolate import interp1d
from mpl_toolkits.axes_grid1.inset_locator import inset_axes
from utilities import time_coord
# %matplotlib inline
def plot_glider(cube, mask_topo=False, track_inset=False, **kw):
"""Plot glider cube."""
cmap = kw.pop('cmap', plt.cm.rainbow)
data = ma.masked_invalid(cube.data.squeeze())
t = time_coord(cube)
#t = t.units.num2date(t.points.squeeze())
dist, pha = sw.dist(lat, lon, units='km')
dist = np.r_[0, np.cumsum(dist)]
dist, z = np.broadcast_arrays(dist[..., None], depth)
try:
z_range = cube.coord(axis='Z').attributes['actual_range']
except KeyError:
z_range = z.min(), z.max()
try:
data_range = cube.attributes['actual_range']
except KeyError:
data_range = data.min(), data.max()
condition = np.logical_and(data >= data_range[0], data <= data_range[1])
data = ma.masked_where(~condition, data)
condition = np.logical_and(z >= z_range[0], z <= z_range[1])
z = ma.masked_where(~condition, z)
fig, ax = plt.subplots(figsize=(9, 3.75))
cs = ax.pcolor(dist, z, data, cmap=cmap, snap=True, **kw)
if mask_topo:
h = z.max(axis=1)
x = dist[:, 0]
ax.plot(x, h, color='black', linewidth='0.5', zorder=3)
ax.fill_between(x, h, y2=h.max(), color='0.9', zorder=3)
#ax.set_title('Glider track from {} to {}'.format(t[0], t[-1]))
fig.tight_layout()
if track_inset:
axin = inset_axes(ax, width="25%", height="30%", loc=4)
axin.plot(lon, lat, 'k.')
start, end = (lon[0], lat[0]), (lon[-1], lat[-1])
kw = dict(marker='o', linestyle='none')
axin.plot(*start, color='g', **kw)
axin.plot(*end, color='r', **kw)
axin.axis('off')
return fig, ax, cs
# -
# # Models
# +
from utilities import CF_names, quick_load_cubes
models = dict(useast=('http://ecowatch.ncddc.noaa.gov/thredds/dodsC/'
'ncom_us_east_agg/US_East_Apr_05_2013_to_Current_best.ncd'),
hycom=('http://ecowatch.ncddc.noaa.gov/thredds/dodsC/'
'hycom/hycom_reg1_agg/HYCOM_Region_1_Aggregation_best.ncd'),
sabgom=('http://omgsrv1.meas.ncsu.edu:8080/thredds/dodsC/'
'fmrc/sabgom/SABGOM_Forecast_Model_Run_Collection_best.ncd'),
coawst=('http://geoport.whoi.edu/thredds/dodsC/'
'coawst_4/use/fmrc/coawst_4_use_best.ncd'))
name_list = CF_names['sea_water_temperature']
coawst = quick_load_cubes(models['coawst'], name_list, strict=True)
useast = quick_load_cubes(models['useast'], name_list, strict=True)
hycom = quick_load_cubes(models['hycom'], name_list, strict=True)
# +
from datetime import datetime
from utilities import proc_cube
# Glider info.
start = glider.coord(axis='T').attributes['minimum']
stop = glider.coord(axis='T').attributes['maximum']
start = datetime.strptime(start, '%Y-%m-%d %H:%M:%S')
stop = datetime.strptime(stop, '%Y-%m-%d %H:%M:%S')
bbox = lon.min(), lat.min(), lon.max(), lat.max()
# Subsetting the cube to the glider limits.
coawst = proc_cube(coawst, bbox=bbox, time=(start, stop), units=glider.units)
useast = proc_cube(useast, bbox=bbox, time=(start, stop), units=glider.units)
hycom = proc_cube(hycom, bbox=bbox, time=(start, stop), units=glider.units)
coawst, useast, hycom
# -
for aux in coawst.aux_factories:
coawst.remove_aux_factory(aux)
# +
from iris.analysis import trajectory
sample_points = [('latitude', lat),
('longitude', lon),
('time', glider.coord(axis='T').points)]
# -
depth = glider.coord('depth').points
fig, ax, cs = plot_glider(glider, mask_topo=False, track_inset=True)
# +
iuseast = trajectory.interpolate(useast, sample_points)
iuseast.transpose()
depth = -iuseast.coord(axis='Z').points
fig, ax, cs = plot_glider(iuseast, mask_topo=False, track_inset=True)
ax.set_ylim(-120, 0)
t = ax.set_title("USEAST")
# +
ihycom = trajectory.interpolate(hycom, sample_points)
ihycom.transpose()
depth = -ihycom.coord(axis='Z').points
fig, ax, cs = plot_glider(ihycom, mask_topo=False, track_inset=True)
ax.set_ylim(-120, 0)
t = ax.set_title("HYCOM")
# +
icoawst = trajectory.interpolate( coawst, sample_points)
icoawst.transpose()
depth = -icoawst.coord(axis='Z').points
fig, ax, cs = plot_glider(ihycom, mask_topo=False, track_inset=True)
ax.set_ylim(-120, 0)
t = ax.set_title("COAWST")
| notebooks/glider/glider_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Targeted Adversarial AI Attack Demo
# #### Constants and Flags
# +
import os
IMG_PATH = os.path.join(os.getcwd(), 'dataset/images/2153d9869fb5a9ef.png')
LABELS_PATH = os.path.join(os.getcwd(), 'dataset/ImageNet_labels.txt')
LABEL_DICT = eval(open(LABELS_PATH, 'r').read())
true_dict_keys = list(LABEL_DICT.keys())
true_dict_index_pos = list(LABEL_DICT.values()).index('matchstick')
TRUE_INDEX = true_dict_keys[true_dict_index_pos]
target_dict_keys = list(LABEL_DICT.keys())
target_dict_index_pos = list(LABEL_DICT.values()).index('goose')
TARGET_CLASS_INDEX = target_dict_keys[target_dict_index_pos]
print(TRUE_INDEX)
print(TARGET_CLASS_INDEX)
# ATTACK_TYPE = 'targeted_basic_iter'
ATTACK_TYPE = 'fgsm'
ATTACK_TYPE = 'targeted_fgsm'
ATTACK_TYPE = 'basic_iter'
ATTACK_TYPE = 'targeted_basic_iter'
# -
LABEL_DICT
# +
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import os
import warnings
warnings.filterwarnings("ignore")
from cleverhans.attacks import FastGradientMethod, BasicIterativeMethod, SaliencyMapMethod
import numpy as np
from PIL import Image
import tensorflow as tf
from tensorflow.contrib.slim.nets import inception
slim = tf.contrib.slim
from matplotlib.pyplot import imshow
# %matplotlib inline
tf.flags.DEFINE_string(
'master', '', 'The address of the TensorFlow master to use.')
tf.flags.DEFINE_string(
'checkpoint_path', './dataset/inception_v3.ckpt', 'Path to checkpoint for inception network.')
tf.flags.DEFINE_string(
'input_dir', './data/dev_images', 'Input directory with images.')
tf.flags.DEFINE_string(
'output_dir', './output/images', 'Output directory with images.')
tf.flags.DEFINE_float(
'max_epsilon', 16.0, 'Maximum size of adversarial perturbation.')
tf.flags.DEFINE_integer(
'image_width', 299, 'Width of each input images.')
tf.flags.DEFINE_integer(
'image_height', 299, 'Height of each input images.')
tf.flags.DEFINE_integer(
'batch_size', 1, 'How many images process at one time.')
tf.logging.set_verbosity(tf.logging.INFO)
FLAGS = tf.flags.FLAGS
BATCH_SHAPE = [FLAGS.batch_size, FLAGS.image_height, FLAGS.image_width, 3]
NUM_CLASSES = 1001
# Images for inception classifier are normalized to be in [-1, 1] interval,
# eps is a difference between pixels so it should be in [0, 2] interval.
# Renormalizing epsilon from [0, 255] to [0, 2].
EPSILON = 2.0 * FLAGS.max_epsilon / 255.0
# -
# #### Utility Functions
def load_image(image_path, batch_shape):
"""Read png images from input directory in batches.
Args:
image_path: path to image file (png)
batch_shape: shape of minibatch array, i.e. [batch_size, height, width, 3]
Yields:
filenames: list file names without path of each image
Length of this list could be less than batch_size, in this case only
first few images of the result are elements of the minibatch.
images: array with all images from this batch
"""
images = np.zeros(batch_shape)
filenames = []
batch_size = batch_shape[0]
with tf.gfile.Open(image_path) as f:
image = np.array(Image.open(f).convert('RGB')).astype(np.float) / 255.0 # [0,1]
# Images for inception classifier are normalized to be in [-1, 1] interval.
images[0, :, :, :] = image * 2.0 - 1.0
filenames.append(os.path.basename(image_path))
yield filenames, images
def show_images(imgs):
# Images for inception classifier are normalized to be in [-1, 1] interval,
# so rescale them back to [0, 1].
img = (((imgs[0, :, :, :] + 1.0) * 0.5) * 255.0).astype(np.uint8)
imshow(img)
# #### Define Defence Model
# +
class InceptionModel(object):
"""Model class for CleverHans library."""
def __init__(self, num_classes):
self.num_classes = num_classes
self.built = False
def __call__(self, x_input):
"""Constructs model and return probabilities for given input."""
reuse = True if self.built else None
with slim.arg_scope(inception.inception_v3_arg_scope()):
_, end_points = inception.inception_v3(
x_input, num_classes=self.num_classes, is_training=False,
reuse=reuse)
self.built = True
output = end_points['Predictions']
# Strip off the extra reshape op at the output
probs = output.op.inputs[0]
return probs
INCEPTION_MODEL = InceptionModel(NUM_CLASSES)
# -
# #### Setup Defence Model
# +
# Prepare defence graph
defence_graph = tf.Graph()
with defence_graph.as_default():
defence_x_input = tf.placeholder(tf.float32, shape=BATCH_SHAPE)
with slim.arg_scope(inception.inception_v3_arg_scope()):
_, end_points = inception.inception_v3(
defence_x_input, num_classes=NUM_CLASSES, is_training=False)
# Restore the checkpoint
defence_sess = tf.Session(graph=defence_graph)
saver = tf.train.Saver()
saver.restore(defence_sess, FLAGS.checkpoint_path)
defence_predicted_labels = tf.nn.top_k(end_points['Predictions'], k=5, sorted=True, name=None)
# Construct the scalar neuron tensor
logits = defence_graph.get_tensor_by_name('InceptionV3/Logits/SpatialSqueeze:0')
neuron_selector = tf.placeholder(tf.int32)
y = logits[0][neuron_selector]
# Construct tensor for predictions
prediction = tf.argmax(logits, 1)
# Prepare computation
# defence_saver = tf.train.Saver(slim.get_model_variables())
# defence_session_creator = tf.train.ChiefSessionCreator(
# scaffold=tf.train.Scaffold(saver=defence_saver),
# checkpoint_filename_with_path=FLAGS.checkpoint_path,
# master=FLAGS.master)
# defence_sess = tf.train.MonitoredSession(session_creator=defence_session_creator)
# -
# #### Setup Adversarial Attack
# Prepare attack graph
adv_graph = tf.Graph()
with adv_graph.as_default():
adv_x_input = tf.placeholder(tf.float32, shape=BATCH_SHAPE)
if ATTACK_TYPE == 'fgsm':
fgsm = FastGradientMethod(INCEPTION_MODEL)
x_adv = fgsm.generate(adv_x_input, eps=EPSILON, clip_min=-1., clip_max=1.)
if ATTACK_TYPE == 'targeted_fgsm':
fgsm = FastGradientMethod(INCEPTION_MODEL)
eps_iter = 2.0 * 1 / 255.0
one_hot_target = np.zeros((1, NUM_CLASSES), dtype=np.float32)
one_hot_target[0, TARGET_CLASS_INDEX] = 1
x_adv = fgsm.generate(adv_x_input, eps=EPSILON, eps_iter=eps_iter, nb_iter=10,
clip_min=-1., clip_max=1., y_target=one_hot_target)
# x_adv = fgsm.generate(adv_x_input, eps=EPSILON, clip_min=-1., clip_max=1.)
elif ATTACK_TYPE == 'basic_iter':
bim = BasicIterativeMethod(INCEPTION_MODEL)
eps_iter = 2.0 * 1 / 255.0
x_adv = bim.generate(adv_x_input, eps=EPSILON, eps_iter=eps_iter, nb_iter=10,
clip_min=-1., clip_max=1.)
elif ATTACK_TYPE == 'targeted_basic_iter':
bim = BasicIterativeMethod(INCEPTION_MODEL)
eps_iter = 2.0 * 1 / 255.0
# eps_iter = 2.0 * 1 / 255.0
# For infinity norm alpha = epsilon / num steps
# eps_iter = EPSILON / 10
one_hot_target = np.zeros((1, NUM_CLASSES), dtype=np.float32)
one_hot_target[0, TARGET_CLASS_INDEX] = 1
x_adv = bim.generate(adv_x_input, eps=EPSILON, eps_iter=eps_iter, nb_iter=10,
clip_min=-1., clip_max=1., y_target=one_hot_target)
# Prepare computation
adv_saver = tf.train.Saver(slim.get_model_variables())
adv_session_creator = tf.train.ChiefSessionCreator(
scaffold=tf.train.Scaffold(saver=adv_saver),
checkpoint_filename_with_path=FLAGS.checkpoint_path,
master=FLAGS.master)
adv_sess = tf.train.MonitoredSession(session_creator=adv_session_creator)
# ## ATTACK
# ### Classify Original Image
TARGET_CLASS_INDEX, LABEL_DICT[TARGET_CLASS_INDEX]
# +
for filenames, images in load_image(IMG_PATH, BATCH_SHAPE):
print('Querying defence model...\n')
labels = defence_sess.run(defence_predicted_labels, feed_dict={defence_x_input: images})
show_images(images)
accuracies_original, label_indices_original = list(labels[0][0]), list(labels[1][0])
print('Original Top-5 Predictions')
for i in range(len(accuracies_original)):
print("{0}:".format(i+1), LABEL_DICT[label_indices_original[i] - 1], accuracies_original[i])
# -
# ### Classify Adversarial Image
# +
for filenames, images in load_image(IMG_PATH, BATCH_SHAPE):
print('Generating adversarial image...')
adv_images = adv_sess.run(x_adv, feed_dict={adv_x_input: images})
show_images(adv_images)
ADV_IMAGE = adv_images[0]
print('Attacking defence model...\n')
adv_labels = defence_sess.run(defence_predicted_labels, feed_dict={defence_x_input: adv_images})
accuracies, label_indices = list(adv_labels[0][0]), list(adv_labels[1][0])
print('Adversarial Top-5 Predictions')
for i in range(len(accuracies)):
# Attack model
print("{0}:".format(i+1), LABEL_DICT[label_indices[i]], accuracies[i])
# -
# ## DEFENCE
# ### Gaussian Smooth Adversarial Images
from scipy.ndimage import gaussian_filter
# +
for filenames, images in load_image(IMG_PATH, BATCH_SHAPE):
print('Generating adversarial image...')
adv_images = adv_sess.run(x_adv, feed_dict={adv_x_input: images})
# gaussian filter to smooth image
print('Smoothing adversarial image...')
filt_images = np.zeros(BATCH_SHAPE)
filt_images[0] = gaussian_filter(adv_images[0], sigma=(1, 1, 0), order=0)
show_images(filt_images)
FILT_IMAGE = filt_images[0]
print('Attacking defence model...\n')
filt_labels = defence_sess.run(defence_predicted_labels, feed_dict={defence_x_input: filt_images})
filt_accuracies, filt_label_indices = list(filt_labels[0][0]), list(filt_labels[1][0])
print('Adversarial Top-5 Predictions after Gaussian Filter')
for i in range(len(filt_accuracies)):
# Attack model
print("{0}:".format(i+1), LABEL_DICT[filt_label_indices[i] - 1], filt_accuracies[i])
# -
# ## VISUALIZE GRAD-CAM
# +
import matplotlib.pyplot as plt
import saliency
from saliency import SaliencyMask
class GradCam(SaliencyMask):
"""A SaliencyMask class that computes saliency masks with Grad-CAM.
https://arxiv.org/abs/1610.02391
Example usage (based on Examples.ipynb):
grad_cam = GradCam(graph, sess, y, images, conv_layer = end_points['Mixed_7c'])
grad_mask_2d = grad_cam.GetMask(im, feed_dict = {neuron_selector: prediction_class},
should_resize = False,
three_dims = False)
The Grad-CAM paper suggests using the last convolutional layer, which would
be 'Mixed_5c' in inception_v2 and 'Mixed_7c' in inception_v3.
"""
def __init__(self, graph, session, y, x, conv_layer):
super(GradCam, self).__init__(graph, session, y, x)
self.conv_layer = conv_layer
self.gradients_node = tf.gradients(y, conv_layer)[0]
def GetMask(self, x_value, feed_dict={}, should_resize = True, three_dims = True):
"""
Returns a Grad-CAM mask.
Modified from https://github.com/Ankush96/grad-cam.tensorflow/blob/master/main.py#L29-L62
Args:
x_value: Input value, not batched.
feed_dict: (Optional) feed dictionary to pass to the session.run call.
should_resize: boolean that determines whether a low-res Grad-CAM mask should be
upsampled to match the size of the input image
three_dims: boolean that determines whether the grayscale mask should be converted
into a 3D mask by copying the 2D mask value's into each color channel
"""
feed_dict[self.x] = [x_value]
(output, grad) = self.session.run([self.conv_layer, self.gradients_node],
feed_dict=feed_dict)
output = output[0]
grad = grad[0]
weights = np.mean(grad, axis=(0,1))
grad_cam = np.ones(output.shape[0:2], dtype=np.float32)
# weighted average
for i, w in enumerate(weights):
grad_cam += w * output[:, :, i]
# pass through relu
grad_cam = np.maximum(grad_cam, 0)
# resize heatmap to be the same size as the input
if should_resize:
grad_cam = grad_cam / np.max(grad_cam) # values need to be [0,1] to be resized
with self.graph.as_default():
grad_cam = np.squeeze(tf.image.resize_bilinear(
np.expand_dims(np.expand_dims(grad_cam, 0), 3),
x_value.shape[:2]).eval(session=self.session))
# convert grayscale to 3-D
if three_dims:
grad_cam = np.expand_dims(grad_cam, axis=2)
grad_cam = np.tile(grad_cam,[1,1,3])
return grad_cam
def create_grad_cam_viz(img, label_index):
grad_mask = grad_cam.GetMask(img, feed_dict={neuron_selector: label_index},
should_resize=True, three_dims=True)
grad_mask = saliency.VisualizeImageGrayscale(grad_mask)
rgba_img = cmap(grad_mask)
rgb_img = np.delete(rgba_img, 3, 2)
return rgb_img
# +
def process_img(img):
'''For scaling back images to visualize gradients'''
img = img - np.min(img)
img = (img / np.max(img) * 255.).astype(np.uint8)
return img
def overlay(array1, array2, alpha=0.5):
"""Overlays `array1` onto `array2` with `alpha` blending.
Args:
array1: The first numpy array.
array2: The second numpy array.
alpha: The alpha value of `array1` as overlayed onto `array2`. This value needs to be between [0, 1],
with 0 being `array2` only to 1 being `array1` only (Default value = 0.5).
Returns:
The `array1`, overlayed with `array2` using `alpha` blending.
"""
if alpha < 0. or alpha > 1.:
raise ValueError("`alpha` needs to be between [0, 1]")
if array1.shape != array2.shape:
raise ValueError('`array1` and `array2` must have the same shapes')
return (array1 * alpha + array2 * (1. - alpha)).astype(array1.dtype)
# -
cmap = plt.get_cmap('jet')
grad_cam = GradCam(defence_graph, defence_sess, y, defence_x_input, conv_layer=end_points['Mixed_7c'])
# +
f, ax = plt.subplots(3,3, figsize=(12,12))
# original image
ax[0,0].imshow(process_img(images[0]))
ax[0,1].imshow(overlay(process_img(create_grad_cam_viz(images[0], TRUE_INDEX)),
process_img(images[0])))
ax[0,2].imshow(overlay(process_img(create_grad_cam_viz(images[0], TARGET_CLASS_INDEX)),
process_img(images[0])))
# adversarial image
ax[1,0].imshow(process_img(adv_images[0]))
ax[1,1].imshow(overlay(process_img(create_grad_cam_viz(adv_images[0], TRUE_INDEX)),
process_img(adv_images[0])))
ax[1,2].imshow(overlay(process_img(create_grad_cam_viz(adv_images[0], TARGET_CLASS_INDEX)),
process_img(adv_images[0])))
# gaussian filtered adversarial image
ax[2,0].imshow(process_img(filt_images[0]))
ax[2,1].imshow(overlay(process_img(create_grad_cam_viz(filt_images[0], TRUE_INDEX)),
process_img(filt_images[0])))
ax[2,2].imshow(overlay(process_img(create_grad_cam_viz(filt_images[0], TARGET_CLASS_INDEX)),
process_img(filt_images[0])))
ax[0,0].set_ylabel("original")
ax[1,0].set_ylabel("adversarial")
ax[2,0].set_ylabel("gaussian filtered adversarial")
ax[0,0].set_title("plain image")
ax[0,1].set_title("w.r.t. true label:\n {0}".format(LABEL_DICT[TRUE_INDEX]))
ax[0,2].set_title("w.r.t. target label:\n {0}".format(LABEL_DICT[TARGET_CLASS_INDEX]))
for i in range(3):
for j in range(3):
ax[i,j].set_xticklabels([])
ax[i,j].set_yticklabels([])
ax[i,j].set_xticks([])
ax[i,j].set_yticks([])
plt.suptitle("Grad-CAM Visualization");
# plt.savefig("sample_output/heatmap_{0}_to_{1}.png".format("_".join(LABEL_DICT[TRUE_INDEX].split()),
# "_".join(LABEL_DICT[TARGET_CLASS_INDEX].split())))
# -
| code/Targeted Adversarial Attack with Gaussian Filter Defense.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div class="alert alert-success">
# <b>Author</b>:
#
# <NAME>
# <EMAIL>
#
# </div>
# Sir only took CT1, didn't take any classes. [See class discussion](https://drive.google.com/file/d/1oMd6RQZ3OKaoqpeaogrQ1LB7q8AtrF9p/view)
| CSE_321_Software Engineering/Lecute_9_27.07.2020.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # R for Drug Development
# ## Pharmacokinetics
# ### [See orginal post](https://rstudio-pubs-static.s3.amazonaws.com/583410_576b7d609e8e490394ab3f4889b43e2e.html)
# 
# ### Example
# For example, consider a drug administered by infusion at a constant rate of ```I=20 mg/h```. The drug has a elimination constant of 0.02 /s and the volume of distribution of the patient ```Vd``` is estimated to be 20 l. The drug is administered for 300 h, and then stopped.
#
# Let us see how the plasma concentration varies with time.
library(tidyverse)
t=seq(0.01,600,0.01)
I=20
k=0.02
V=20
C=ifelse(t<300,(I/(k*V))*(1-exp(-k*t)),(I/(k*V))*exp(-k*(t-300)))
df<-tibble(t=t,C=C)
glimpse(df)
g<-ggplot(df,aes(x=t,y=C))+
geom_line()+
xlab('Time (h)')+
ylab('Concentration (mg/l)')
g
# ### Questions
# #### Determine steady state concentration
# What is the steady state concentration of this drug? Is it what you expect, given the values of I, k and Vd?
#
# #### Determine elimination constant
# Suppose you only had the data, how would you determine the value of k from the data? To do this, we should study the period after after 300 h, when infusion has stopped
#
# How would you plot the data to show only the data for `t > 300 h`?
# +
library(tidyverse)
df_elim<-filter(df,t>300)
df_elim<-mutate(df_elim,t=t-300)
g<-ggplot(df_elim,aes(x=t,y=C))+
geom_line()+
xlab('Time (h)')+
ylab('Concentration (mg/l)')
g
# -
# 
df_elim<-mutate(df_elim,logC=log(C))
glimpse(df_elim)
g<-ggplot(df_elim,aes(x=t,y=logC))+
geom_line()
g
fit_elim<-lm(logC~t,data=df_elim)
summary(fit_elim)
k=-coefficients(fit_elim)[2]
k
# ### Determine Half-life
# 
k=0.05
V=20
t_half=log(2)/k
t_half
| R for Drug Development - Pharmacokinetics.ipynb |
# <a href="https://colab.research.google.com/github/mottaquikarim/PYTH2/blob/master/src/PSETS/nb/hw1_psets.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# # BASIC DATA TYPES
#
# ## SHOPPING_LIST
#
#
# ### P1.PY
#
# +
"""
Shopping List Calculator I
"""
# Create five variables,
# set them to strings that represent 5 common shopping list items
item_name_1 = None
item_name_2 = None
item_name_3 = None
item_name_4 = None
item_name_5 = None
# Create five more variables,
# set them to floats that represent the prices of each of the items above
item_price_1 = None
item_price_2 = None
item_price_3 = None
item_price_4 = None
item_price_5 = None
# Create five more variables,
# set them to ints that represent the quantity of each of the items above
item_quant_1 = None
item_quant_2 = None
item_quant_3 = None
item_quant_4 = None
item_quant_5 = None
# Print to the console the name and price of each item defined above as follows:
# 1 Coco Puffs = $8.95.
# where:
# 1 would be item_quant_1
# Coco Puffs would be item_name_1
# 8.95 would be item_name_2
# -
#
#
# ### P2.PY
#
#
#
# +
"""
Shopping List Calculator II
"""
# Rewrite p1, but this time use the input() command to solicit user input for name, price, quantity. Here's how it works:
item_name_1 = input('Name your first item: ')
# ^ this will ask user to input value of item_name_1
# use input() function and ask user to name items
item_name_1 = None
item_name_2 = None
item_name_3 = None
item_name_4 = None
item_name_5 = None
# use input() function and ask user to name prices
item_price_1 = None
item_price_2 = None
item_price_3 = None
item_price_4 = None
item_price_5 = None
# use input() function and ask user to name quants
item_quant_1 = None
item_quant_2 = None
item_quant_3 = None
item_quant_4 = None
item_quant_5 = None
# Print to the console the name and price of each item defined above as follows:
# 1 Coco Puffs = $8.95.
# where:
# 1 would be item_quant_1
# Coco Puffs would be item_name_1
# 8.95 would be item_name_2
# JUST REMEMBER: now this will be defined by the user!!
# -
#
#
| src/PSETS/nb/hw1_psets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from pysrc.review.train.preprocess import parse_sents, standardize
from nltk.tokenize import sent_tokenize
def prepare_text(text):
text = parse_sents(sent_tokenize(text))
text = standardize(text)
text = ' '.join(text)
return text
# Prepare topic abstracts
#
topics_df = pd.read_csv("topic/topic_abstracts.csv")
topics_df = topics_df.groupby(by=['topic_id']).agg({'text': lambda x: ''.join(x)}).reset_index()
topics_df = topics_df.rename(columns={'topic_id':'id', 'text': 'paper_top50'})
topics_df
def create_dummy_dataframe(df):
df['paper_top50'] = df['paper_top50'].apply(lambda text: prepare_text(text))
df['abstract'] = 'dummy abstract'
df['gold_ids_top6'] = str([0])
return df
test_df = create_dummy_dataframe(topics_df)
test_df
test_df.to_csv("data/pubmedtop50_test_topic.csv")
| pysrc/review/train/topic_summarization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="Z9s6c9qUu3Pv"
import pandas as pd
# + [markdown] id="XZB2mSxAu3Pz"
# # 1.0 Limpieza base de datos ingresos
# + id="MHDPIPlxu3P0"
ingreso = pd.read_csv(r'conjunto_de_datos_ingresos_enigh_2018_ns.csv')
# + [markdown] id="-dPBdq_tu3P2"
# 1. Seleccionamos hogar principal
# + id="iC_qvHWRu3P3"
hogar_principal = ingreso['foliohog'] ==1
ingresos_hogarprincipal =ingreso[hogar_principal]
# + [markdown] id="FsWnKB9tu3P6"
# 1.1 limpiamos los espacios vacíos
# + id="qW24lbwou3P7"
df_updated = ingresos_hogarprincipal.replace(to_replace ='\s\s*', value = '0', regex = True)
# + [markdown] id="MNzjRO9ku3QA"
# 1.2 cambiamos ingresos a número
# + id="T2XcCgIcu3QA"
df_updated['ing_2'] = df_updated['ing_2'].astype(int)
df_updated['ing_3'] = df_updated['ing_3'].astype(int)
df_updated['ing_4'] = df_updated['ing_4'].astype(int)
df_updated['ing_5'] = df_updated['ing_5'].astype(int)
df_updated['ing_6'] = df_updated['ing_6'].astype(int)
# + [markdown] id="SIm-hetCu3QC"
# 1.3 sumo todos los ingresos del hogar: quiero el ingreso total de la vivienda
# + id="xCPy9DIWu3QD"
ingreso_todos=df_updated.groupby(
['folioviv']
).agg(
{
'ing_1':sum,
'ing_2':sum,
'ing_3':sum,
'ing_4':sum,
'ing_5':sum,
'ing_6':sum
}
)
# + id="5VbtxGuyu3QH" outputId="35a3c39e-b527-4cd4-e79c-b580d5e9a048" colab={"base_uri": "https://localhost:8080/", "height": 235}
ingreso_todos.head()
# + id="1z6nfuzlu3QK" outputId="d767729e-aa63-4380-beb3-20d7d68cf4fc" colab={"base_uri": "https://localhost:8080/", "height": 136}
ingreso_todos.dtypes
# + [markdown] id="OZf9zD47u3QM"
# 1.4 queremos el ingreso promedio de los 6 meses
# + id="I-2_of7tu3QM"
col = ingreso_todos.loc[: , "ing_1":"ing_6"]
ingreso_todos['mean_income'] = col.mean(axis=1)
# + [markdown] id="8G1RcuN2u3QO"
# 1.5 con el aggregate me indexo la columna folioviv la quiero como columna
# + id="0OGkrEysu3QP"
ingreso_todos=ingreso_todos.reset_index()
# + id="t1qDyxc5u3QR" outputId="b19c6220-f70c-43fe-80db-8574bb2f0edc" colab={"base_uri": "https://localhost:8080/", "height": 204}
ingreso_todos.head()
# + [markdown] id="Ec5p_urDu3QU"
# # 2.0 Limpieza base de datos gastos
# + id="kriQJjQPu3QU" outputId="5153ee03-3a30-497b-d830-65a0e4865bcc" colab={"base_uri": "https://localhost:8080/", "height": 71}
gastos_hogar = pd.read_csv(r'conjunto_de_datos_gastoshogar_enigh_2018_ns.csv')
# + [markdown] id="Qt3_pMZRu3QY"
# 2. Seleccionamos hogar principal
# + id="kx3JY-fdu3QZ"
hogar_principal = gastos_hogar['foliohog'] ==1
gastos_hogarprincipal =gastos_hogar[hogar_principal]
# + [markdown] id="4LsVHfutu3Qb"
# 2.1 sólo quiero el gasto mensual conservo sólo las columnas que necesito
# + id="lS2h5j6zu3Qc"
gastos_hogarprincipal_onlygastos = gastos_hogarprincipal[['folioviv', 'clave', 'gasto']]
# + [markdown] id="WuiKXuExu3Qh"
# 2.2 verifico que no haya espacios en blanco y tipos de variables
# + id="OWI4IAvPxJxo"
df_updated = gastos_hogarprincipal.replace(to_replace ='\s\s*', value = '0', regex = True)
# + id="eL806kQ0xQVN"
df_updated['folioviv'] = df_updated['folioviv'].astype(str)
df_updated['clave'] = df_updated['clave'].astype(str)
df_updated['gasto'] = df_updated['gasto'].astype(int)
# + id="karSR778yFDG"
df_updated2 = df_updated[['folioviv', 'clave', 'gasto']]
# + id="iwp7oiTpu3Qf"
df_updated2.to_csv(r'gastos_hogarprincipal_onlygastoos.csv')
# + id="xQQPV7jUu3Qk"
gastos_hogarprincipal_onlygastos = pd.read_csv(r'gastos_hogarprincipal_onlygastoos.csv', dtype={'folioviv':'str', 'clave':'str','gasto':'int32'})
# + id="fsKOtj1Iu3Qn" outputId="2f8807e9-893a-47e1-f6e4-f96046f24707" colab={"base_uri": "https://localhost:8080/", "height": 102}
gastos_hogarprincipal_onlygastos.dtypes
# + id="W_-oHopRu3Qp"
# por clave de gasto no voy a saber de que gasto hablo necesito unir el catalogo de gastos
# + [markdown] id="oXYgiDwfu3Qs"
# 2.3 uniendo el catalogo de gastos
# + id="BQu-JhQQu3Qs"
#ojo con el encoding o no voy a tener bien los acentos
catalogo_gastos=pd.read_csv(r'/content/gastos.csv', encoding='latin-1')
# + id="tKSoj19-u3Qu"
#tengo el mismo numero de hogares? cuales quiero conservar?
gastos_hogarprincipal = pd.merge(gastos_hogarprincipal_onlygastos, catalogo_gastos, left_on='clave', right_on='gastos', how='left')
# + id="wS5jJaJZywfq"
del(gastos_hogarprincipal['Unnamed: 0'])
# + id="Nqu28PdWu3Qw" outputId="cab32b7c-99fc-4fa1-d102-6a39330d61e5" colab={"base_uri": "https://localhost:8080/", "height": 204}
gastos_hogarprincipal.head()
# + id="qG8jGgzau3Q0" outputId="a216166a-435c-4f7f-a9a8-fa4cc0951285" colab={"base_uri": "https://localhost:8080/", "height": 170}
gastos_hogarprincipal['gasto'].describe()
# + id="KGrUQL5hu3Q3"
#ya no me interesa tener la clave de gasto
gastos_hogarprincipal=gastos_hogarprincipal[['folioviv','gasto', 'descripción']]
# + [markdown] id="soizaEp4u3Q7"
# 2.4 necesito una tabla con gasto como variable
# + id="ciEJ4ZFxu3Q8"
#quiero que los gastos sean columna no fila
result_hogar = gastos_hogarprincipal.pivot_table('gasto', ['folioviv'], 'descripción')
# + id="ngd0QI8Ku3Q-" outputId="f1440b40-9a12-4061-a832-b80fd3f67c5a" colab={"base_uri": "https://localhost:8080/", "height": 471}
result_hogar.head()
# + id="-U-UZnoJu3RB"
# que significan los NaN?
result_hogar=result_hogar.fillna(0)
# + id="_8F6oxhJu3RC" outputId="6079830a-6d80-49b0-8796-e8e0a4a23429" colab={"base_uri": "https://localhost:8080/", "height": 471}
result_hogar.head()
# + id="2ehMqQAwu3RF" outputId="35985ff5-de9a-4cb6-cb88-f660a8a202b7" colab={"base_uri": "https://localhost:8080/", "height": 34}
#verifico cuantos hogares tengo para ver que todo siga bien
gastos_hogarprincipal['folioviv'].nunique()
# + id="N3ckTAgyu3RH"
#necesito conservar el folioviv como variable no como index
result_hogar=result_hogar.reset_index()
# + id="H0m0eHQ2u3RK"
ingreso_todos['folioviv']=ingreso_todos['folioviv'].astype(str)
# + [markdown] id="xhVAbRSUu3RM"
# 2.5 unimos los ingresos
# + id="kfCBQUmYu3RM"
gastos_ingresos_hogar= pd.merge(result_hogar, ingreso_todos, on='folioviv')
# + id="BZePEzrRu3RO" outputId="c4adca72-37c9-4200-b713-c4993acb9819" colab={"base_uri": "https://localhost:8080/", "height": 34}
gastos_ingresos_hogar['folioviv'].nunique()
# + id="sQ3TraWKu3RS" outputId="d4f56d1e-eaaf-4d8a-9660-9873df0c1122" colab={"base_uri": "https://localhost:8080/", "height": 440}
gastos_ingresos_hogar.head()
# + id="QWcYktBnu3RW" outputId="2b27af95-6e4a-4b47-e787-15fea2f8df17" colab={"base_uri": "https://localhost:8080/", "height": 626}
gastos_ingresos_hogar.drop(['ing_1', 'ing_2', 'ing_3', 'ing_4', 'ing_5', 'ing_6'], axis=1)
# + id="z1eEfkM9u3RZ"
#quiero datos de la vivienda para hacer perfiles demograficos
# + [markdown] id="Kg8jnFSwu3Rb"
# # 3.0 datos vivienda
# + id="A20XEUXJu3Rc" outputId="68b22660-6916-4fae-fee3-e746f1c9fcc7" colab={"base_uri": "https://localhost:8080/", "height": 71}
datos_vivienda = pd.read_csv(r'conjunto_de_datos_viviendas_enigh_2018_ns.csv')
# + id="b4RAMzzYu3Re" outputId="3b5c5ba3-46a7-4245-a693-611bc4372529" colab={"base_uri": "https://localhost:8080/", "height": 34}
datos_vivienda['folioviv'].dtypes
# + id="4-hY8NuQu3Rg"
datos_vivienda['folioviv']=datos_vivienda['folioviv'].astype(str)
# + id="NTmSThJPu3Ri"
gastos_ingresos_vivienda= pd.merge(gastos_ingresos_hogar, datos_vivienda, on='folioviv', how='left')
# + id="EK7OIzkru3Rk" outputId="30063f72-e7ab-46e1-9170-9f874eb5c79e" colab={"base_uri": "https://localhost:8080/", "height": 34}
gastos_ingresos_vivienda['folioviv'].nunique()
# + id="XdRNTFNDu3Rr"
#tengo la columna ubicageo pero quisiera tenerlos por nombre
# + [markdown] id="sEKAB2NEu3Rt"
# # 4.0 catalogo de municipios
# + id="ClFzUdVSu3Rt"
ubica_geo=pd.read_csv(r'ubic_geo.csv', encoding='latin-1')
# + id="sSlqumX7u3Rv" outputId="59f4f0cd-0c18-4202-ca7a-445e07f39995" colab={"base_uri": "https://localhost:8080/", "height": 204}
ubica_geo.head()
# + id="TIOtWuPnu3Rx"
gastos_ingresos_vivienda= pd.merge(gastos_ingresos_vivienda, ubica_geo, left_on='ubica_geo', right_on='ubic_geo',how='left')
# + id="9rObuVO4u3R1" outputId="495bb0fe-64de-4fea-9804-d14d17465f8a" colab={"base_uri": "https://localhost:8080/", "height": 440}
gastos_ingresos_vivienda.head()
# + id="LaRAKdGe0Dsh" outputId="38d76ea2-a28f-4dc0-c28d-5db979ab68bd" colab={"base_uri": "https://localhost:8080/", "height": 1000}
gastos_ingresos_vivienda.sample(n=20)
# + id="MLxPQY0x0e9_"
df_final = gastos_ingresos_vivienda.replace(to_replace ='\s\s*', value = '0', regex = True)
# + id="T0J1h_X90mra" outputId="d3cf98b9-55b1-4698-b373-cfd6101b97ec" colab={"base_uri": "https://localhost:8080/", "height": 905}
df_final.sample(n=20)
# + id="KAQh8CPa0uZ3" outputId="9e7b557c-c9e7-4f8e-ff70-d8b3f90e7a6c" colab={"base_uri": "https://localhost:8080/", "height": 1000}
df_final.info(verbose =True)
# + id="QaWyZ49ku3R6"
# esta completa nuestra base la guardamos:
df_final.to_csv(r'gastos_ingresos_vivienda_geografico_enigh18_final.csv')
# + id="Vwdo8RJZu3R-"
/content/gastos_ingresos_vivienda_geografico_enigh18.csv
/content/gastos_ingresos_vivienda_geografico_enigh18_final.csv
| Notebooks/LimpiezaDatos/DF_FINAL_ENIGH.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Tensorflow introduction
# Tensorflow is an open-source software library for dataflow programming across a range of tasks. It is a symbolic math library, and also used for machine learning applications such as neural networks. Here we try to introduce it using some easy elementry examples.
# +
# %matplotlib inline
import numpy as np
import pylab as plt
import tensorflow as tf
params = {'axes.labelsize': 'x-large',
'axes.titlesize':'x-large',
'xtick.labelsize':'x-large',
'ytick.labelsize':'x-large'}
plt.rcParams.update(params)
# -
# You can define constant in different format like:
node1 = tf.constant(3.0, dtype=tf.float32)
node2 = tf.constant(4.0) # also tf.float32 implicitly
print(node1)
print(node2)
# As mentioned before, tnesorflow is a C basec packages which has python interface. For calculation you should open a tensorflow session like:
with tf.Session() as sess:
print(sess.run([node1, node2]))
# For example if you want to add two number using tensorflow, you can make the data flow through operations (here addition) then you have to open a session (here by with command) and run it like:
node3 = tf.add(node1, node2)
with tf.Session() as sess:
print(sess.run(node3))
a = tf.placeholder(tf.float32)
b = tf.placeholder(tf.float32)
# adder_node = tf.add(a, b)
adder_node = a + b
with tf.Session() as sess:
print(sess.run(adder_node, {a: 3, b:4.5}))
print(sess.run(adder_node, {a: [1,3], b: [2, 4]}))
# Or you can define more complicated precedures:
a = tf.placeholder(tf.float32)
b = tf.placeholder(tf.float32)
c = tf.placeholder(tf.float32)
adder_node = (a + b)*c
with tf.Session() as sess:
print(sess.run(adder_node, {a: 3, b:4.5, c:2}))
# Sinse almost every Machine learning problem is based on optimization, let's try to solve an optimization problem. For this pupose assume that there are an input set "x_train" which we want to find a simple linear model <br>
# $y=W\times x+b$ <br>
# to predict "y_train" vector.
# +
x_train = [1,2,3,4]
y_train = [0,-1,-2,-3]
W = tf.Variable([.3], dtype=tf.float32)
b = tf.Variable([-.3], dtype=tf.float32)
x = tf.placeholder(tf.float32)
linear_model = W * x + b
init = tf.global_variables_initializer()
with tf.Session() as sess:
sess.run(init)
print(sess.run(linear_model, {x:x_train}))
# +
W = tf.Variable([.3], dtype=tf.float32)
b = tf.Variable([-.3], dtype=tf.float32)
x = tf.placeholder(tf.float32)
linear_model = W * x + b
y = tf.placeholder(tf.float32)
squared_deltas = tf.square(linear_model - y)
loss = tf.reduce_sum(squared_deltas)
init = tf.global_variables_initializer()
with tf.Session() as sess:
sess.run(init)
print(sess.run(loss, {x:x_train, y:y_train}))
# +
W = tf.Variable([.3], dtype=tf.float32)
b = tf.Variable([-.3], dtype=tf.float32)
x = tf.placeholder(tf.float32)
linear_model = W * x + b
y = tf.placeholder(tf.float32)
squared_deltas = tf.square(linear_model - y)
loss = tf.reduce_sum(squared_deltas)
optimizer = tf.train.GradientDescentOptimizer(0.01)
train = optimizer.minimize(loss)
init = tf.global_variables_initializer()
with tf.Session() as sess:
sess.run(init)
for i in range(1000):
sess.run(train, {x:x_train, y:y_train})
print(sess.run([W, b]))
# -
# evaluate training accuracy
with tf.Session() as sess:
sess.run(init)
for i in range(1000):
sess.run(train, {x:x_train, y:y_train})
curr_W, curr_b, curr_loss = sess.run([W, b, loss], {x:x_train, y:y_train})
print("W: %s b: %s loss: %s"%(curr_W, curr_b, curr_loss))
# # Example
# +
with np.load('./datasets/mnist.npz', allow_pickle=True) as f:
x_train, y_train = f['x_train'], f['y_train']
x_test, y_test = f['x_test'], f['y_test']
print(x_train.shape,y_train.shape,x_test.shape,y_test.shape)
x_train, x_test = x_train / 255.0, x_test / 255.0
# -
plt.imshow(x_train[np.random.randint(len(x_train))])
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28)),
tf.keras.layers.Dense(512, activation=tf.nn.relu),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(10, activation=tf.nn.softmax)
])
model.summary()
# +
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
model.fit(x_train, y_train, epochs=5)
# -
model.evaluate(x_test, y_test)
# +
hist = model.history
fig,(ax1,ax2) = plt.subplots(1,2,figsize=(15,5))
ax1.plot(hist.epoch,hist.history['loss'])
ax1.set_xlabel('epochs')
ax1.set_ylabel('loss')
ax2.plot(hist.epoch,hist.history['acc'])
ax2.set_xlabel('epochs')
ax2.set_ylabel('acc')
# -
# +
n_train = 50000
with np.load('./datasets/Hoda_data.npz', allow_pickle=True) as f:
print (f['img'].shape,f['target'].shape)
x_train, y_train = f['img'][:n_train], f['target'][:n_train]
x_test, y_test = f['img'][n_train:], f['target'][n_train:]
print(x_train.shape,y_train.shape,x_test.shape,y_test.shape)
# -
plt.imshow(x_train[0])
| session_I-TF_intro/Inrtoduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import ipywidgets as widgets
from IPython.display import display, HTML
javascript_functions = {False: "hide()", True: "show()"}
button_descriptions = {False: "Show code", True: "Hide code"}
def toggle_code(state):
"""
Toggles the JavaScript show()/hide() function on the div.input element.
"""
output_string = "<script>$(\"div.input\").{}</script>"
output_args = (javascript_functions[state],)
output = output_string.format(*output_args)
display(HTML(output))
def button_action(value):
"""
Calls the toggle_code function and updates the button description.
"""
state = value.new
toggle_code(state)
value.owner.description = button_descriptions[state]
state = False
toggle_code(state)
button = widgets.ToggleButton(state, description = button_descriptions[state])
button.observe(button_action, "value")
display(button)
# -
from windows_widgets import CompileOutputOnly, CompileInputOuput, ShortAnswerQuestion, ChoiceQuestion, AchieveRate, add_link_buttons
# # Chapter 8 - 포인터
# ## 1.1 메모리 주서와 주소연산자 &
# ### 주소 개념
# **메모리 공간은 바이트마다 고유한 주소(address)가 있다.** 마치 아파트 각 세대마다 고유 번호가 있는 것과 같다. 아파트의 호수로 집을 찾듯이 주소를 이용하여 메모리의 위치를 파악할 수 있다. 메모리 주소는 0부터 바이트마다 1씩 증가한다. **메모리 주소는 저장 장소인 변수 이름과 함께 기억 장소를 참조하는 또 다른 방법이다.** 이 주소값을 이용하면 보다 편리하고 융통성 있는 프로그램을 만들 수 있다. 그러나 메모리 주소를 잘못 다루면 시스템에 심각한 문제를 일으킬 수 있다. 또한 메모리 주소를 처음 학습하는 사람에겐 좀 어려울 수 있다.
co1 = CompileOutputOnly('exer8_1')
cio1 = CompileInputOuput('exer8_9')
saq1 = ShortAnswerQuestion('(1) 메모리 공간은 바이트마다 고유한 ____(이)가 있다.', ['주소', '주소값', 'address', 'Address'], ' 주소를 말한다.', ' 주소를 이용하여 메모리의 위치를 파악할 수 있다.')
cq1 = ChoiceQuestion("""(2) 배열 선언
double a[] = {2, 4, 5, 7, 8, 9};
에서 *a와 *(a+2)의 참조값은 각각 무엇인가?
""", ['4, 7', '2, 5', '5, 8', '2, 4'], 1, ' 인덱스는 0부터 시작한다.', ' *a는 2, *(a+2)는 5이다.')
cq2 = ChoiceQuestion("""다음은 여러 포인터와 선언에 대한 설명이다. 다음 중에서 잘못 설명하고 있는 것은 무엇인가?""", ['double형 포인터 선언: double *pd;', 'int형 포인터 원소 4개인 배열 선언: int *p[4];', '일차원 배열 int a[3]의 배열 포인터 선언: int *p;', '이차원 배열 int b[3][4]의 배열 포인터 선언: int *p[3][4];'], 3, ' 이차원 배열 포인터는 *를 두개 붙여 선언한다.', ' int **p로 선언한다.')
rate = AchieveRate()
add_link_buttons(1, 'sample_windows2.ipynb')
| .ipynb_checkpoints/sample_windows3-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import boto3
import os
ACCESS_KEY = os.environ['S3_ACCESS_KEY']
SECRET_KEY = os.environ['S3_SECRET_KEY']
bucket=os.environ['S3_BUCKET']
s3 = boto3.client('s3', aws_access_key_id=ACCESS_KEY,
aws_secret_access_key=SECRET_KEY)
path='/data/ETLcache'
filelist= [file for file in os.listdir(path) if file.endswith('.png')]
filelist
for f in filelist:
s3.upload_file(path+'/'+f, bucket, f, ExtraArgs={'ACL':'public-read'})
| ETL/toS3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.1 64-bit
# language: python
# name: python38164bit9a8e3b63a08644a087459c5617ed8408
# ---
# # Waffle Charts <a id="6"></a>
#
#
# A `waffle chart` is an interesting visualization that is normally created to display progress toward goals. It is commonly an effective option when you are trying to add interesting visualization features to a visual that consists mainly of cells, such as an Excel dashboard.
#
# <hr>
# +
import numpy as np
import pandas as pd
df = pd.read_csv('Canada.csv')
df.drop(['Unnamed: 0'], axis='columns', inplace=True)
df.head()
# +
# Removing uncessary columns
df.drop(['AREA', 'REG', 'DEV', 'Type', 'Coverage'], axis='columns', inplace=True)
# Rename the columns so that they make sense
df.rename(columns={
'OdName':'Country', 'AreaName':'Continent','RegName':'Region'
}, inplace=True)
# Making all column labels of type string
df.columns = list(map(str, df.columns))
# Setting the country name as index
df.set_index('Country', inplace=True)
# Add total column
df['Total'] = df.sum(axis=1)
# +
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches # needed for waffle Charts
mpl.style.use('ggplot') # optional: for ggplot-like style
# +
# let's create a new dataframe for these three countries
df_dsn = df.loc[['Denmark', 'Norway', 'Sweden'], :]
# let's take a look at our dataframe
df_dsn
# -
# <hr>
#
# ## Creating own's waffle chart
# **Step 1.** The first step into creating a waffle chart is determing the proportion of each category with respect to the total.
# +
# compute the proportion of each category with respect to the total
total_values = sum(df_dsn['Total'])
category_proportions = [(float(value) / total_values) for value in df_dsn['Total']]
# print out proportions
for i, proportion in enumerate(category_proportions):
print (df_dsn.index.values[i] + ': ' + str(proportion))
# -
# **Step 2.** The second step is defining the overall size of the `waffle` chart.
# +
width = 40 # width of chart
height = 10 # height of chart
total_num_tiles = width * height # total number of tiles
print ('Total number of tiles is ', total_num_tiles)
# -
# **Step 3.** The third step is using the proportion of each category to determe it respective number of tiles
# +
# compute the number of tiles for each catagory
tiles_per_category = [round(proportion * total_num_tiles) for proportion in category_proportions]
# print out number of tiles per category
for i, tiles in enumerate(tiles_per_category):
print (df_dsn.index.values[i] + ': ' + str(tiles))
# -
# Based on the calculated proportions, Denmark will occupy 129 tiles of the `waffle` chart, Norway will occupy 77 tiles, and Sweden will occupy 194 tiles.
# **Step 4.** The fourth step is creating a matrix that resembles the `waffle` chart and populating it.
# +
# initialize the waffle chart as an empty matrix
waffle_chart = np.zeros((height, width))
# define indices to loop through waffle chart
category_index = 0
tile_index = 0
# populate the waffle chart
for col in range(width):
for row in range(height):
tile_index += 1
# if the number of tiles populated for the current category is equal to its corresponding allocated tiles...
if tile_index > sum(tiles_per_category[0:category_index]):
# ...proceed to the next category
category_index += 1
# set the class value to an integer, which increases with class
waffle_chart[row, col] = category_index
print ('Waffle chart populated!')
# -
waffle_chart
# As expected, the matrix consists of three categories and the total number of each category's instances matches the total number of tiles allocated to each category.
#
#
# **Step 5.** Map the `waffle` chart matrix into a visual.
# +
# instantiate a new figure object
fig = plt.figure()
# use matshow to display the waffle chart
colormap = plt.cm.coolwarm
plt.matshow(waffle_chart, cmap=colormap)
plt.colorbar()
# -
# **Step 6.** Prettify the chart.
# +
# instantiate a new figure object
fig = plt.figure()
# use matshow to display the waffle chart
colormap = plt.cm.coolwarm
plt.matshow(waffle_chart, cmap=colormap)
plt.colorbar()
# get the axis
ax = plt.gca()
# set minor ticks
ax.set_xticks(np.arange(-.5, (width), 1), minor=True)
ax.set_yticks(np.arange(-.5, (height), 1), minor=True)
# add gridlines based on minor ticks
ax.grid(which='minor', color='w', linestyle='-', linewidth=2)
plt.xticks([])
plt.yticks([])
# -
# **Step 7.** Create a legend and add it to chart.
# +
# instantiate a new figure object
fig = plt.figure()
# use matshow to display the waffle chart
colormap = plt.cm.coolwarm
plt.matshow(waffle_chart, cmap=colormap)
plt.colorbar()
# get the axis
ax = plt.gca()
# set minor ticks
ax.set_xticks(np.arange(-.5, (width), 1), minor=True)
ax.set_yticks(np.arange(-.5, (height), 1), minor=True)
# add gridlines based on minor ticks
ax.grid(which='minor', color='w', linestyle='-', linewidth=2)
plt.xticks([])
plt.yticks([])
# compute cumulative sum of individual categories to match color schemes between chart and legend
values_cumsum = np.cumsum(df_dsn['Total'])
total_values = values_cumsum[len(values_cumsum) - 1]
# create legend
legend_handles = []
for i, category in enumerate(df_dsn.index.values):
label_str = category + ' (' + str(df_dsn['Total'][i]) + ')'
color_val = colormap(float(values_cumsum[i])/total_values)
legend_handles.append(mpatches.Patch(color=color_val, label=label_str))
# add legend to chart
plt.legend(handles=legend_handles,
loc='lower center',
ncol=len(df_dsn.index.values),
bbox_to_anchor=(0., -0.2, 0.95, .1)
)
# -
# Now it would very inefficient to repeat these seven steps every time we wish to create a `waffle` chart. So let's combine all seven steps into one function called *create_waffle_chart*. This function would take the following parameters as input:
#
# > 1. **categories**: Unique categories or classes in dataframe.
# > 2. **values**: Values corresponding to categories or classes.
# > 3. **height**: Defined height of waffle chart.
# > 4. **width**: Defined width of waffle chart.
# > 5. **colormap**: Colormap class
# > 6. **value_sign**: In order to make our function more generalizable, we will add this parameter to address signs that could be associated with a value such as %, $, and so on. **value_sign** has a default value of empty string.
def create_waffle_chart(categories, values, height, width, colormap, value_sign=''):
# compute the proportion of each category with respect to the total
total_values = sum(values)
category_proportions = [(float(value) / total_values) for value in values]
# compute the total number of tiles
total_num_tiles = width * height # total number of tiles
print ('Total number of tiles is', total_num_tiles)
# compute the number of tiles for each catagory
tiles_per_category = [round(proportion * total_num_tiles) for proportion in category_proportions]
# print out number of tiles per category
for i, tiles in enumerate(tiles_per_category):
print (df_dsn.index.values[i] + ': ' + str(tiles))
# initialize the waffle chart as an empty matrix
waffle_chart = np.zeros((height, width))
# define indices to loop through waffle chart
category_index = 0
tile_index = 0
# populate the waffle chart
for col in range(width):
for row in range(height):
tile_index += 1
# if the number of tiles populated for the current category
# is equal to its corresponding allocated tiles...
if tile_index > sum(tiles_per_category[0:category_index]):
# ...proceed to the next category
category_index += 1
# set the class value to an integer, which increases with class
waffle_chart[row, col] = category_index
# instantiate a new figure object
fig = plt.figure()
# use matshow to display the waffle chart
colormap = plt.cm.coolwarm
plt.matshow(waffle_chart, cmap=colormap)
plt.colorbar()
# get the axis
ax = plt.gca()
# set minor ticks
ax.set_xticks(np.arange(-.5, (width), 1), minor=True)
ax.set_yticks(np.arange(-.5, (height), 1), minor=True)
# add dridlines based on minor ticks
ax.grid(which='minor', color='w', linestyle='-', linewidth=2)
plt.xticks([])
plt.yticks([])
# compute cumulative sum of individual categories to match color schemes between chart and legend
values_cumsum = np.cumsum(values)
total_values = values_cumsum[len(values_cumsum) - 1]
# create legend
legend_handles = []
for i, category in enumerate(categories):
if value_sign == '%':
label_str = category + ' (' + str(values[i]) + value_sign + ')'
else:
label_str = category + ' (' + value_sign + str(values[i]) + ')'
color_val = colormap(float(values_cumsum[i])/total_values)
legend_handles.append(mpatches.Patch(color=color_val, label=label_str))
# add legend to chart
plt.legend(
handles=legend_handles,
loc='lower center',
ncol=len(categories),
bbox_to_anchor=(0., -0.2, 0.95, .1)
)
# Now to create a waffle chart, all we have to do is call the function create_waffle_chart. Let's define the input parameters:
# +
width = 40 # width of chart
height = 10 # height of chart
categories = df_dsn.index.values # categories
values = df_dsn['Total'] # correponding values of categories
colormap = plt.cm.coolwarm # color map class
# -
create_waffle_chart(categories, values, height, width, colormap)
# <hr>
| uncommon-data-visualization/waffle-charts/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bubble Sort
# +
list_number = [5, 7, 3, 5, 2, 0, 6, 8]
list_sorted = [1,2,3,4,5,6,7,8,9]
sort = False
while sort is False:
sort = True
for i in range(len(list_number)-1):
if list_number[i] > list_number[i+1]:
list_number[i],list_number[i+1] = list_number[i+1], list_number[i]
sort = False
list_number
# -
# # Extracting the tax return from 2012 year
# +
import requests
link = "http://s3.amazonaws.com/mikeghen/costs_2012.txt"
f = requests.get(link)
raw_data = f.text
data = raw_data.split('\n')[:-1]
new_list = []
for d in data:
a = d.replace('$','')
new_list.append(float(a.replace(',','')))
average_of_2012 = (sum(new_list) + 348.43)/ len(data)
datalen_2012 = len(data)
average_of_2012
# -
# # Extracting the tax return from 2014 year
# +
import requests
link = "http://s3.amazonaws.com/mikeghen/costs_2014.txt"
f = requests.get(link)
raw_data = f.text
data = raw_data.split('\n')[:-1]
new_list = []
for d in data:
a = d.replace('$','')
new_list.append(float(a.replace(',','')))
average_of_2014 = (sum(new_list) + 948.38)/len(data)
datalen_2014 = len(data)
average_of_2014
# -
# # Extracting the tax return from 2016 year
# +
import requests
link = "http://s3.amazonaws.com/mikeghen/costs_2016.txt"
f = requests.get(link)
raw_data = f.text
data = raw_data.split('\n')[:-1]
new_list = []
for d in data:
a = d.replace('$','')
new_list.append(float(a.replace(',','')))
average_of_2016 = (sum(new_list) + 565.23)/ len(data)
datalen_2016 = len(data)
average_of_2016
# -
# # Percentage increase in average tax return from 2012 and 2014
per_change_12to14 = ((average_of_2014 - average_of_2012)/ average_of_2012)*100
per_change_12to14
# # Percentage increase in average tax return from 2014 and 2016
per_change_14to16 = ((average_of_2016 - average_of_2014)/average_of_2014)*100
per_change_14to16
# # Yearly Percentage increase
raised_value = per_change_14to16/per_change_12to14
raised_value
# # Predicted Percentage increase value from 2016 to 2018
per_value2018 = per_change_14to16 * raised_value
per_value2018
# # Predicted average tax return value for 2018
(average_of_2016 * (per_value2018/100)) + average_of_2016
| Class Assignments/class_02-12_Tax_Return_Problem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/opi-lab/stsiva-workshop/blob/main/notebooks/stsiva_workshop_notebook01.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="a8XrbGJa15jt"
# # Pinhole camera model fundamentals
#
# *STSIVA Workshop - 2021*
#
# **Speakers:**
# <NAME> - [@jhacsonmeza](https://github.com/jhacsonmeza)
# <NAME>, PhD - [@agmarrugo](https://github.com/agmarrugo)
# Universidad Tecnológica de Bolívar
#
# In this example, we will be exploring how to estimate the camera matrix and project world points from a known object. Adapted from [EGNN 512](http://inside.mines.edu/~whoff/) by <NAME>.
#
# *The source of this notebook is located at https://github.com/opi-lab/stsiva-workshop/*
#
# *Useful intro about [Colab](https://colab.research.google.com/notebooks/welcome.ipynb)*
#
# *Useful intro about [OpenCV](https://opencv.org/)*
# + [markdown] id="ADjH2PwCId-s"
# ### The image of a cube
#
# A cube has vertices in world coordinates: (0,0,0), (1,0,0),
# (1,1,0), (0,1,0), (0,0,1), (1,0,1), (1,1,1), (0,1,1). A camera is located at
# (X,Y,Z) = (3,-4,2) in world coordinates. The camera points directly
# at the origin and there is no roll about the axis (i.e., the +Z axis of the world points up in the image).
#
# Generate an image of a wireframe model of the cube as if were seen by the camera, as shown in the figure. Assume a pinhole camera model, with focal length = 600 pixels, where the image size is 640 pixels wide by 480 pixels high.
#
# 
# + [markdown] id="_EbFnb4pK9I4"
# ## Solution
#
# We use the fact that the +Z axis of the camera is the direction in which it points. We can get this by computing the vector from the camera’s location, to the point at which it is looking (the world
# origin): $\hat{\mathbf{z}} = -\mathbf{t}/|\mathbf{t}|$, where $\mathbf{t}$ is the location of the camera in the world.
#
# Next, we know that there is no roll about the axis (i.e., the +Z axis of the world points up in the image). This is equivalent to saying that the +X axis of the camera lies in the XY plane of the world. We can get this by doing a cross product of the camera’s Z axis with world Z axis.
#
# Finally, the Y axis of the camera is given by the cross product of the Z axis with the X axis.
# + [markdown] id="4IS4zpAnL89o"
# ## The code:
# + id="uuZJZXBvT1LU"
# Let's import the necessary libraries
import numpy as np
import cv2
import matplotlib.pylab as plt
# from google.colab.patches import cv2_imshow # for image display
# import pandas as pd
# from skimage import io
# from PIL import Image
# import urllib.request
# + colab={"base_uri": "https://localhost:8080/"} id="6a_ai8yIUPk9" outputId="b11e9e6d-66ce-414d-ca6d-b6955b42e77e"
# Create camera pose
tc_w = np.array([3, -4, 2]) # location of camera origin in world
# The z axis of camera is the unit vector from camera to the world origin
uz_w = -tc_w/np.linalg.norm(tc_w)
print('The camera z-axis:', uz_w)
# The x axis of camera is in xy plane of world.
ux_w = np.cross(uz_w, np.array([0, 0, 1]))
ux_w = ux_w/np.linalg.norm(ux_w)
print('The camera x-axis:', ux_w)
# finally, form the y axis
uy_w = np.cross(uz_w, ux_w);
print('The camera y-axis:', uy_w)
R_c_w = np.hstack((ux_w.reshape(3,1), uy_w.reshape(3,1), uz_w.reshape(3,1)))
print('The camera rotation matrix: \n', R_c_w)
H_c_w = np.hstack((R_c_w, tc_w.reshape(3,1)))
H_c_w = np.vstack((H_c_w, np.array([0, 0, 0, 1])))
H_w_c = np.linalg.inv(H_c_w)
# + colab={"base_uri": "https://localhost:8080/"} id="r3CxKwVdcAsg" outputId="78b9d5c8-a8d5-4fe2-c77b-2c175cee165d"
# Create camera projection matrices
# Here are the given parameters of the camera:
H = 480 # height of image in pixels
W = 640 # width of image in pixels
f = 600 # focal length in pixels
cx = W/2 # optical center
cy = H/2
# % Intrinsic camera parameter matrix
K = np.array([[f, 0, cx], [0, f, cy], [0, 0, 1]])
print('The camera intrinsic matrix:\n', K)
# Extrinsic camera parameter matrix
Mext = H_w_c[0:3,:]
print('The camera extrinsic matrix:\n', Mext)
# + id="WeyIL8Bic3xy"
# Define model in world coords
# The complete set of vertices is
P_w = np.array([[0, 1, 1, 0, 0, 1, 1, 0],
[0, 0, 1, 1, 0, 0, 1, 1],
[0, 0, 0, 0, 1, 1, 1, 1],
[1, 1, 1, 1, 1, 1, 1, 1]])
# print(P_w)
# P_w = P_w
# Define the lines to be drawn (indices of starting and ending points)
# lower 4 points form a square
# upper 4 points form a square
# connect lower square to upper square
Lines = np.array([
[0, 1],
[1, 2],
[2, 3],
[3, 0],
[4, 5],
[5, 6],
[6, 7],
[7, 4],
[0, 4],
[1, 5],
[2, 6],
[3, 7]])
# + id="uLNG1QaAeZYk" colab={"base_uri": "https://localhost:8080/"} outputId="bba56102-7352-4587-c320-752ab02dcdc7"
# Do perspective projection
p_img = K @ Mext @ P_w
# We convert p_img from homogenous to cartesian
p_img = p_img[:-1]/p_img[-1]
print('The projected points are:')
print(p_img)
# + colab={"base_uri": "https://localhost:8080/", "height": 269} id="y9qysiT6f31o" outputId="c5f40504-5bcb-430d-d59e-b1d807da52c0"
# Create image
I = np.zeros((H, W));
plt.imshow(I);
for i in range(0, p_img.shape[-1]):
plt.plot(p_img[0,i], p_img[1,i], 'w*')
plt.text(p_img[0,i], p_img[1,i], '{}'.format(i), fontsize=12, color='w')
for i in range(0, Lines.shape[0]):
i1 = Lines[i, 0] # index of starting point
i2 = Lines[i, 1] # index of ending point
plt.plot([p_img[0, i1], p_img[0, i2]], [p_img[1, i1], p_img[1, i2]], color='w')
plt.show()
# + [markdown] id="RUUqYOHoOevZ"
# ## TODO
#
# Modify the above code to move the camera's location closer to or further away from the cube. You can also scale or shift the cube's coordinates and check that the image also moves.
| notebooks/stsiva_workshop_notebook01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=false editable=false tags=[]
# ## Data Preprocessing
#
# *Prepared by:*
# **<NAME>**
# Faculty, Software Technology Department
# College of Computer Studies - De La Salle University
# -
# This notebook shows how to perform common preprocessing techniques in Python.
# ## Preliminaries
# ### Import libraries
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style('darkgrid')
sns.set_palette('Set2')
# sns.color_palette('Set2')
# -
# ### Load data
df = pd.read_csv('https://raw.githubusercontent.com/Cyntwikip/data-repository/main/titanic.csv')
df.head()
df.info()
# ## Preprocessing
# ### Text transformations
#
# By accessing the `str` attribute of an object feature/column in Pandas, we can use the methods under string data type / object.
df['Name'].str.lower()
df['Name'].str.upper()
df['Name'].str.title()
df['Name'].str.split(',')
# ### Encoding
#
# In many cases, we need our data to be in numerical format, so how should we deal with datasets with categorical data in it? We can use different encoding strategies for that. One of which is One-hot Encoder. This encoding strategy creates one column for each unique value in the original column. We use this when there is no hierarchy in our categories.
df[['Embarked']]
df['Embarked'].value_counts()
df['Embarked'].isnull().sum()
# #### Pandas get_dummies
#
# One approach for doing one-hot encoding is through Pandas' get_dummies function.
pd.get_dummies(df['Embarked'])
# #### Sklearn OneHotEncoder
#
# Another approach for doing one-hot encoding is through sklearn's OneHotEncoder class.
# +
from sklearn.preprocessing import OneHotEncoder
encoder = OneHotEncoder(handle_unknown='ignore')
df_encoded = encoder.fit_transform(df[['Embarked']]).toarray()
df_encoded = pd.DataFrame(df_encoded, columns=encoder.categories_)
df_encoded
# -
df_encoded.sum(axis=0)
# Notice that there are 4 columns here instead of 3. This is because it also creates a new column (the last one) for the null values.
#
# Additionally, we can transform it back to its original form.
pd.DataFrame(encoder.inverse_transform(df_encoded))
# ### Binning
#
# Binning converts a continuous feature into a categorical one by chunking/binning the values. This is somewhat like the opposite of one-hot encoding.
df['Fare']
df['Fare'].describe()
fig, ax = plt.subplots(1,1, figsize=(8,4), dpi=100)
df['Fare'].hist(bins=50, ax=ax)
plt.show()
# #### Manual cuts
# +
bins = [0, 50, 100, 200, 400]
# Create Group Names
group_names = ['0-49.99','50-99.99','100-199.99','200-399.99']
fare_binned = pd.cut(df['Fare'], bins, labels=group_names, include_lowest=True) # to include the leftmost value in the bins
fare_binned.head()
# -
fare_binned.value_counts()
fare_binned[fare_binned.isnull()]
df['Fare'][fare_binned.isnull()]
# #### Cuts with equal spacing
# +
fare_binned = pd.cut(df['Fare'], 5)
fare_binned.head()
# -
# ### Handling missing data
# #### Dropping
#
# If there is not much null values, we can simply drop them.
df.info()
df.dropna().info()
# Although in this example, many rows were omitted.
# #### Imputation
#
# Another approach is to impute or fill in missing values instead. We can change the imputation strategy to mean, median, most frequent, and constant. We will use most frequent since it can handle categorical data.
# +
from sklearn.impute import SimpleImputer
imputer = SimpleImputer(missing_values=np.nan, strategy='most_frequent')
df_imputed = pd.DataFrame(imputer.fit_transform(df), columns=df.columns)
df_imputed
# -
# No more missing values now!
df_imputed.info()
# ### Feature Selection
#
# Sometimes, we don't need some of the features, so we simply drop them.
df_selected = df.drop(['PassengerId', 'Name'], axis=1)
df_selected
# Here's one-liner code to remove categorical features which I believe will be very useful in many cases.
df_selected = df.loc[:, df.dtypes!='object']
df_selected
# ### Scaling
#
# There are times wherein we will have to rescale our data especially when dealing with Machine Learning.
# +
from sklearn.preprocessing import StandardScaler, MinMaxScaler
scaler = MinMaxScaler(feature_range=(0,1))
scaled = scaler.fit_transform(df_selected)
df_scaled = pd.DataFrame(scaled, columns=df_selected.columns)
df_scaled
# -
# Now our minimum and maximum values are 0 and 1.
df_scaled.describe()
# We can also use other scaler techniques such as standard scaler, which standardizes the data by scaling it based on its mean and standard deviation.
# +
scaler = StandardScaler()
scaled = scaler.fit_transform(df_selected)
df_scaled = pd.DataFrame(scaled, columns=df_selected.columns)
df_scaled
# -
# Now, the data is standardized.
df_scaled.describe()
# ### Grouping
#
# We can group by a specific feature/column and chain it with whatever aggregation function we would like to use.
df['Pclass'].value_counts()
df.groupby('Pclass').sum()
df.groupby('Pclass').mean()
# ### Using mathematical operations and functions
#
df_copy = df.copy()
df_copy['Fare_transformed'] = df['Fare']*2
df_copy[['Fare', 'Fare_transformed']]
df_copy['Fare_log'] = np.log(df_copy['Fare'])
df_copy[['Fare', 'Fare_log']]
# ### Using custom functions
# +
def func(x):
if x < 50:
return 'low'
elif x < 100:
return 'medium'
else:
return 'high'
df_copy['Fare_custom_func'] = df['Fare'].apply(func)
df_copy[['Fare', 'Fare_custom_func']]
# + [markdown] editable=false tags=[]
# ## End
# <sup>made by **<NAME>**</sup> <br>
# <sup>for comments, corrections, suggestions, please email:</sup><sup> <href><EMAIL></href> or <href><EMAIL></href></sup><br>
| preprocessing/basic-preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.5 ('base')
# language: python
# name: python3
# ---
# %matplotlib inline
import pandas as pd
import numpy as np
import os
import matplotlib.pyplot as plt
import seaborn as sns
from scipy.stats import poisson
from initial_data_processing import ProcessSoccerData
pd.options.display.max_columns = 15
pd.options.display.max_colwidth = 1000
soccer_data = ProcessSoccerData()
df = soccer_data.get_matches_df()
# dictionary_df = soccer_data.get_dictionary_df() #seperate df for each league/season
# +
df.groupby('League')['Home_Goals', 'Away_Goals'].mean().sort_values('Home_Goals').plot(kind='barh')
plt.ylabel('League')
plt.xlabel('Average goals')
plt.title('Average goals by league')
leg=plt.legend(loc='lower right', fontsize=13, ncol=1)
plt.show()
df.groupby('League')['Home_Goals', 'Away_Goals'].agg(['mean','var']).plot(kind='barh', width=1, figsize=(20,10))
plt.ylabel('League')
plt.xlabel('Average goals')
plt.title('Mean and Variance of goals by league')
leg=plt.legend(loc='lower left', fontsize=13, ncol=1)
plt.show()
# -
# Looking at the data in the 2nd plot, the means and variances are similar, which would be the case if the number of home goals and number of away goals are Poisson distributed(mean=var). However, in many leagues the variance is slightly larger than the mean, which is a sign of overdispersion in the Poisson case.
# +
home_away_goals = df[['Home_Goals', 'Away_Goals']]
# calculate Poisson probability for home goals and away goals
poisson_prob = np.column_stack([[poisson.pmf(i, home_away_goals.mean()[j]) for i in range(8)] for j in range(2)])
# plot histogram of actual goals
plt.hist(home_away_goals[['Home_Goals', 'Away_Goals']].values, range(9),
alpha=0.8, label=['Home', 'Away'],density=True, color=["steelblue", "coral"])
# add lines for the Poisson distributions
plt.plot([i-0.5 for i in range(1,9)], poisson_prob[:,0],
linestyle='-', marker='o',label="Home", color = 'steelblue')
plt.plot([i-0.5 for i in range(1,9)], poisson_prob[:,1],
linestyle='-', marker='o',label="Away", color = 'coral')
leg=plt.legend(loc='upper right', fontsize=13, ncol=2)
leg.set_title(" Actual Poisson ", prop = {'size':'14', 'weight':'bold'})
plt.xticks([i-0.5 for i in range(1,9)],[i for i in range(8)])
plt.xlabel("Goals per Match",size=13)
plt.ylabel("Proportion of Matches",size=13)
plt.title("Number of Goals per Match All Seasons and Leagues",size=14,fontweight='bold')
plt.ylim([-0.004, 0.4])
plt.tight_layout()
plt.show()
# -
print(df[['Home_goals', 'Away_goals']].mean())
print(df[['Home_goals', 'Away_goals']].var())
# small overdispersion but mean approx equal to var
# +
df.head()
df.shape
df.info()
df.describe().round(2)
#checking proportions
p1 = (df.Result.value_counts(normalize = True) *100).round(2).reset_index()
p1.columns = ['result', '%']
p1
p2 = (df.Match_Result.value_counts(normalize = True) *100).round(2).reset_index()
p2.columns = ['result', '%']
p2
# -
# Most of the time the home team wins as expected. 28% matches end in a draw.
number_cols = df.dtypes[df.dtypes != 'object'].index.tolist()
number_cols
# +
cols_to_drop = ['Season', 'Match_id','Home_Team', 'Away_Team']
cols_for_correlation = list(set(number_cols) - set(cols_to_drop))
cols_for_correlation
# +
df['H_win'] = np.where(df.Match_Result == 'H', 1, 0)
df['A_win'] = np.where(df.Match_Result == 'A', 1, 0)
df['Draw'] = np.where(df.Match_Result == 'D', 1, 0)
df[cols_for_correlation + ['H_win']].corr()['H_win'].sort_values(ascending = False).reset_index()
# -
df[cols_for_correlation + ['A_win']].corr()['A_win'].sort_values(ascending = False).reset_index()
df[cols_for_correlation + ['Draw']].corr()['Draw'].sort_values(ascending = False).reset_index()
# low correlation for draws - difficult to predict?
# Now do some analyis on calculated features:
#
# NEED TO DO!
#
# +
# for each feature col look at distribution see if normally dist'd
sns.histplot(df[column]).set(ylabel = None)
# -
| src/eda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# A tutorial exercise which uses cross-validation with linear models.
#
# This exercise is used in the [Cross-validated estimators](http://scikit-learn.org/stable/tutorial/statistical_inference/model_selection.html#cv-estimators-tut) part of the [Model selection: choosing estimators and their parameters](http://scikit-learn.org/stable/tutorial/statistical_inference/model_selection.html#model-selection-tut) section of the [A tutorial on statistical-learning for scientific data processing](http://scikit-learn.org/stable/tutorial/statistical_inference/index.html#stat-learn-tut-index).
# #### New to Plotly?
# Plotly's Python library is free and open source! [Get started](https://plot.ly/python/getting-started/) by downloading the client and [reading the primer](https://plot.ly/python/getting-started/).
# <br>You can set up Plotly to work in [online](https://plot.ly/python/getting-started/#initialization-for-online-plotting) or [offline](https://plot.ly/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plot.ly/python/getting-started/#start-plotting-online).
# <br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
# ### Version
import sklearn
sklearn.__version__
# ### Imports
# This tutorial imports [LassoCV](http://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LassoCV.html#sklearn.linear_model.LassoCV), [Lasso](http://scikit-learn.org/stable/modules/generated/sklearn.linear_model.Lasso.html#sklearn.linear_model.Lasso), [KFold](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.KFold.html#sklearn.model_selection.KFold) and [cross_val_score](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.cross_val_score.html#sklearn.model_selection.cross_val_score).
# +
from __future__ import print_function
print(__doc__)
import plotly.plotly as py
import plotly.graph_objs as go
import numpy as np
from sklearn import datasets
from sklearn.linear_model import LassoCV
from sklearn.linear_model import Lasso
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_val_score
# -
# ### Calculations
# +
diabetes = datasets.load_diabetes()
X = diabetes.data[:150]
y = diabetes.target[:150]
lasso = Lasso(random_state=0)
alphas = np.logspace(-4, -0.5, 30)
scores = list()
scores_std = list()
n_folds = 3
for alpha in alphas:
lasso.alpha = alpha
this_scores = cross_val_score(lasso, X, y, cv=n_folds, n_jobs=1)
scores.append(np.mean(this_scores))
scores_std.append(np.std(this_scores))
scores, scores_std = np.array(scores), np.array(scores_std)
# -
# ### Plot cross-validation with linear models
# +
p1 = go.Scatter(x=alphas, y=scores,
mode='lines',
line=dict(color='blue'),
fill='tonexty'
)
# plot error lines showing +/- std. errors of the scores
std_error = scores_std / np.sqrt(n_folds)
p2 = go.Scatter(x=alphas, y=scores + std_error,
mode='lines',
line=dict(color='blue', dash='dash'),
)
p3 = go.Scatter(x=alphas, y=scores - std_error,
mode='lines',
line=dict(color='blue', dash='dash'),
fill='tonexty')
line = go.Scatter(y=[np.max(scores), np.max(scores)],
x=[min(alphas), max(alphas)],
mode='lines',
line=dict(color='black', dash='dash',
width=1),
)
layout = go.Layout(xaxis=dict(title='alpha', type='log'),
yaxis=dict(title='CV score +/- std error'),
showlegend=False
)
fig = go.Figure(data=[p2, p1, p3, line], layout=layout)
# -
py.iplot(fig)
# ### Bonus Question
# Bonus: how much can you trust the selection of alpha?
# +
# To answer this question we use the LassoCV object that sets its alpha
# parameter automatically from the data by internal cross-validation (i.e. it
# performs cross-validation on the training data it receives).
# We use external cross-validation to see how much the automatically obtained
# alphas differ across different cross-validation folds.
lasso_cv = LassoCV(alphas=alphas, random_state=0)
k_fold = KFold(3)
print("Answer to the bonus question:",
"how much can you trust the selection of alpha?")
print()
print("Alpha parameters maximising the generalization score on different")
print("subsets of the data:")
for k, (train, test) in enumerate(k_fold.split(X, y)):
lasso_cv.fit(X[train], y[train])
print("[fold {0}] alpha: {1:.5f}, score: {2:.5f}".
format(k, lasso_cv.alpha_, lasso_cv.score(X[test], y[test])))
print()
print("Answer: Not very much since we obtained different alphas for different")
print("subsets of the data and moreover, the scores for these alphas differ")
print("quite substantially.")
# +
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
# ! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'Cross-validation on diabetes Dataset Exercise.ipynb', 'scikit-learn/plot-cv-diabetes/', 'Cross-validation on Diabetes Dataset Exercise | plotly',
' ',
title = 'Cross-validation on Diabetes Dataset Exercise | plotly',
name = 'Cross-validation on diabetes Dataset Exercise',
has_thumbnail='true', thumbnail='thumbnail/cv-diabetes.jpg',
language='scikit-learn', page_type='example_index',
display_as='tutorial_exercises', order=3,
ipynb= '~Diksha_Gabha/3069')
# -
| _posts/scikit/c-v-on-diabetes-dataset/Cross-validation on diabetes Dataset Exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **[Deep Learning Home Page](https://www.kaggle.com/learn/deep-learning)**
#
# ---
#
# # Exercise Introduction
#
# The cameraman who shot our deep learning videos mentioned a problem that we can solve with deep learning.
#
# He offers a service that scans photographs to store them digitally. He uses a machine that quickly scans many photos. But depending on the orientation of the original photo, many images are digitized sideways. He fixes these manually, looking at each photo to determine which ones to rotate.
#
# In this exercise, you will build a model that distinguishes which photos are sideways and which are upright, so an app could automatically rotate each image if necessary.
#
# If you were going to sell this service commercially, you might use a large dataset to train the model. But you'll have great success with even a small dataset. You'll work with a small dataset of dog pictures, half of which are rotated sideways.
#
# Specifying and compiling the model look the same as in the example you've seen. But you'll need to make some changes to fit the model.
#
# **Run the following cell to set up automatic feedback.**
# set up code checking
from learntools.core import binder
binder.bind(globals())
from learntools.deep_learning.exercise_4 import *
print("Setup is completed")
# # 1. Specify the Model
#
# Since this is your first time, we'll provide some starter code for you to modify. You will probably copy and modify code the first few times you work on your own projects.
#
# There are some important parts left blank in the following code.
#
# Fill in the blanks (marked with `____`) and run the cell
#
# + tags=["raises-exception"]
from tensorflow.keras.applications import ResNet50
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Flatten, GlobalAveragePooling2D
num_classes = 2
resnet_weights_path = '../input/resnet50/resnet50_weights_tf_dim_ordering_tf_kernels_notop.h5'
my_new_model = Sequential()
my_new_model.add(ResNet50(include_top=False, pooling='avg', weights=resnet_weights_path))
my_new_model.add(Dense(num_classes, activation='softmax'))
# indicate whether the first layer should be trained/changed or not.
my_new_model.layers[0].trainable = False
# check your answer
step_1.check()
# +
# step_1.hint()
# step_1.solution()
# -
# <hr/>
#
# # 2. Compile the Model
#
# You now compile the model with the following line. Run this cell.
my_new_model.compile(optimizer='sgd',
loss='categorical_crossentropy',
metrics=['accuracy'])
# That ran nearly instantaneously. Deep learning models have a reputation for being computationally demanding. Why did that run so quickly?
#
# After thinking about this, check your answer by uncommenting the cell below.
# check your answer (Run this code cell to receive credit!)
step_2.solution()
# <hr/>
#
# # 3. Review the Compile Step
# You provided three arguments in the compile step.
# - optimizer
# - loss
# - metrics
#
# Which arguments could affect the accuracy of the predictions that come out of the model? After you have your answer, run the cell below to see the solution.
# check your answer (Run this code cell to receive credit!)
step_3.solution()
# <hr/>
#
# # 4. Fit Model
#
# **Your training data is in the directory `../input/dogs-gone-sideways/images/train`. The validation data is in `../input/dogs-gone-sideways/images/val`**. Use that information when setting up `train_generator` and `validation_generator`.
#
# You have 220 images of training data and 217 of validation data. For the training generator, we set a batch size of 10. Figure out the appropriate value of `steps_per_epoch` in your `fit_generator` call.
#
# Fill in all the blanks (again marked as `____`). Then run the cell of code. Watch as your model trains the weights and the accuracy improves.
# + tags=["raises-exception"]
from tensorflow.keras.applications.resnet50 import preprocess_input
from tensorflow.keras.preprocessing.image import ImageDataGenerator
image_size = 224
data_generator = ImageDataGenerator(preprocess_input)
train_generator = data_generator.flow_from_directory(
directory="../input/dogs-gone-sideways/images/train",
target_size=(image_size, image_size),
batch_size=10,
class_mode='categorical')
validation_generator = data_generator.flow_from_directory(
directory="../input/dogs-gone-sideways/images/val",
target_size=(image_size, image_size),
class_mode='categorical')
# fit_stats below saves some statistics describing how model fitting went
# the key role of the following line is how it changes my_new_model by fitting to data
fit_stats = my_new_model.fit_generator(train_generator,
steps_per_epoch=22,
validation_data=validation_generator,
validation_steps=1)
# check your answer
step_4.check()
# +
# step_4.solution()
# -
#
# Can you tell from the results what fraction of the time your model was correct in the validation data?
#
# In the next step, we'll see if we can improve on that.
#
# # Keep Going
# Move on to learn about **[data augmentation](https://www.kaggle.com/dansbecker/data-augmentation/)**. It is a clever and easy way to improve your models. Then you'll apply data augmentation to this automatic image rotation problem.
#
# ---
# **[Deep Learning Home Page](https://www.kaggle.com/learn/deep-learning)**
#
#
#
#
#
# *Have questions or comments? Visit the [Learn Discussion forum](https://www.kaggle.com/learn-forum) to chat with other Learners.*
| deep-learning/exercise/transfer-learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + nbpresent={"id": "d3c1477a-770b-4903-8861-9d2f60e1a467"}
import os
from utils import load_caption, decode_caption, load_vocab
from tqdm import tqdm
import pickle
import numpy as np
from matplotlib import pyplot as plt
# %matplotlib inline
plt.style.use('ggplot')
# -
vocab = load_vocab('/home/spb61/coco2014_vocab.json')
# + nbpresent={"id": "ce7f4af1-bec5-485c-abab-f95d1f137f63"}
cap = load_caption(1, image_dir="/datadrive/val_beam_2_states/")
print(cap['total_prob'])
# + nbpresent={"id": "542e3243-fd17-41ea-bde3-6626618f24c6"}
# def total_probs(beam_size=2):
# image_dir = "/datadrive/val_beam_{}_states/".format(beam_size)
# total_probs = []
# for image in tqdm(os.listdir(image_dir)):
# cap = load_caption(image, image_dir=image_dir)
# total_probs.append(cap['total_prob'])
# if not image.endswith("json"):
# print(image)
# print("Images found: {}".format(len(total_probs)))
# return total_probs
# probs = {}
# for k in [1,2,10,100]:
# probs[k] = total_probs(beam_size=k)
# pickle.dump(probs, open( "probs.pickle", "wb" ) )
probs = pickle.load(open( "probs.pickle", "rb" ))
# +
mean_sizes = []
sizes_std = []
for k in sorted(probs.keys()):
mean_sizes.append(np.mean(probs[k]))
sizes_std.append(np.std(probs[k]))
print("Beam size {}: probability covered: {:.4f}\t{:.4f}".format(k, np.mean(probs[k]), np.std(probs[k])))
X = [str(x) for x in sorted(probs.keys())]
plt.bar(X, mean_sizes, yerr=sizes_std)
# + nbpresent={"id": "201a7cee-a750-42fa-a3d3-d9e6fac00609"}
base = plt.cm.get_cmap(plt.cm.jet)
color_list = base(np.linspace(0, 1, len(probs.keys())))
max_prob = np.max([np.max(ps) for ps in probs.values()]) + 0.03
for k, color in zip(sorted(probs.keys()), color_list):
plt.hist(probs[k], bins=100, label="k={}".format(k), color=color, histtype='step');
plt.hist(probs[k], bins=100, alpha=0.1, color=color);
plt.yscale('log')
plt.xlim([0, max_prob])
print(k)
plt.title("Distribution of total probabilty mass covered by the beam searches")
plt.legend()
plt.savefig('../outputs/figs/total_probs_distribution.png', bbox_inches='tight')
# + nbpresent={"id": "0c1f9e64-f147-4c9e-878d-af2b3a9c0fbf"}
# color_list = base(np.linspace(0, 1, len(probs.keys())))
# max_prob = np.max([np.max(ps) for ps in probs.values()]) + 0.03
# for k, color in zip(sorted(probs.keys()), color_list):
# plt.hist(probs[k], bins=100, label="k={}".format(k), color=color, histtype='step', cumulative=True);
# plt.hist(probs[k], bins=100, alpha=0.1, color=color, cumulative=True);
# # plt.yscale('log')
# plt.xlim([0, max_prob])
# print(k)
probs_100 = np.array(probs[100])
print(sum(probs_100 < 0.1)/ len(probs_100))
print(sum(probs_100 > 0.2) / len(probs_100))
# plt.hist(probs_100, bins=100, cumulative=True, histtype='step', color='red');
# plt.hist(probs_100, bins=100, alpha=0.1, cumulative=True, color='red');
# # plt.yscale('log')
| evaluation/Evaluate on server.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## First Example
#
# A football is kicked with an angle of 30 degrees with the ground. The mass of the ball is 0.43 kg. Consider the gravitational acceleration as 9.81 m/s$^2$. The initial speed of the ball is 30 m/s and the initial height is 0 m. Find the motion of the ball.
#
#
# The free-body diagram of the ball is depicted below:
#
# <figure><img src="../images/ballGrav.png\" width=700 />
import matplotlib.pyplot as plt
import numpy as np
# +
g = 9.81
t = np.arange(0, 3.5, 0.01)
x = 15*np.sqrt(3)*t
y = -g/2*t**2 + 15*t
plt.plot(x, y, 'r', linewidth = 5)
plt.ylim(0,13)
plt.yticks(np.arange(0,13,2), size=20)
plt.xticks(np.arange(0,90,20), size=20)
plt.xlabel('x (m)', size = 20)
plt.ylabel('y (m)', size = 20)
# +
dt = 0.01
x0 = 0
y0 = 0
vx0 = 15*np.sqrt(3)
vy0 = 15
x = x0
y = y0
vx = vx0
vy = vy0
rx = np.array([])
ry = np.array([])
rx = np.append(rx,x)
ry = np.append(ry,y)
# -
for i in t:
dvxdt = 0
dvydt = -g
dxdt = vx
dydt = vy
x = x + dt*dxdt
y = y + dt*dydt
vx = vx + dt*dvxdt
vy = vy + dt*dvydt
rx = np.append(rx, x)
ry = np.append(ry, y)
print('y='+ str(y))
print('x='+ str(x) + '\n')
plt.plot(rx,ry)
plt.ylim(0, 13)
# ### An example a little bit more complex
#
# Now, besides the gravity, we consider the air resistance ($b$ = 0.006 Ns/m). First we will consider the air resistance proportional to the speed and opposite direction of the velocity vector.
#
# <figure><img src="../images/ballGravLinearRes.png\" width=700 />
#
# ## Task for today
#
# Write a Jupyter notebook to find the trajectory of a ball considering the air drag proportional to the square root of the ball velocity.
#
# $\vec{F_b} = -\frac{b}{\sqrt{v}}\vec{v}$
#
# <figure><img src="../images/solP12018Q2_D.png\" width=200 />
| courses/modsim2018/renatowatanabe/.ipynb_checkpoints/Untitled2-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] origin_pos=0
# # 层和块
# :label:`sec_model_construction`
#
# 之前首次介绍神经网络时,我们关注的是具有单一输出的线性模型。
# 在这里,整个模型只有一个输出。
# 注意,单个神经网络
# (1)接受一些输入;
# (2)生成相应的标量输出;
# (3)具有一组相关 *参数*(parameters),更新这些参数可以优化某目标函数。
#
# 然后,当考虑具有多个输出的网络时,
# 我们利用矢量化算法来描述整层神经元。
# 像单个神经元一样,层(1)接受一组输入,
# (2)生成相应的输出,
# (3)由一组可调整参数描述。
# 当我们使用softmax回归时,一个单层本身就是模型。
# 然而,即使我们随后引入了多层感知机,我们仍然可以认为该模型保留了上面所说的基本架构。
#
# 对于多层感知机而言,整个模型及其组成层都是这种架构。
# 整个模型接受原始输入(特征),生成输出(预测),
# 并包含一些参数(所有组成层的参数集合)。
# 同样,每个单独的层接收输入(由前一层提供),
# 生成输出(到下一层的输入),并且具有一组可调参数,
# 这些参数根据从下一层反向传播的信号进行更新。
#
# 事实证明,研究讨论“比单个层大”但“比整个模型小”的组件更有价值。
# 例如,在计算机视觉中广泛流行的ResNet-152架构就有数百层,
# 这些层是由*层组*(groups of layers)的重复模式组成。
# 这个ResNet架构赢得了2015年ImageNet和COCO计算机视觉比赛
# 的识别和检测任务 :cite:`He.Zhang.Ren.ea.2016`。
# 目前ResNet架构仍然是许多视觉任务的首选架构。
# 在其他的领域,如自然语言处理和语音,
# 层组以各种重复模式排列的类似架构现在也是普遍存在。
#
# 为了实现这些复杂的网络,我们引入了神经网络*块*的概念。
# *块*(block)可以描述单个层、由多个层组成的组件或整个模型本身。
# 使用块进行抽象的一个好处是可以将一些块组合成更大的组件,
# 这一过程通常是递归的,如 :numref:`fig_blocks`所示。
# 通过定义代码来按需生成任意复杂度的块,
# 我们可以通过简洁的代码实现复杂的神经网络。
#
# 
# :label:`fig_blocks`
#
# 从编程的角度来看,块由*类*(class)表示。
# 它的任何子类都必须定义一个将其输入转换为输出的前向传播函数,
# 并且必须存储任何必需的参数。
# 注意,有些块不需要任何参数。
# 最后,为了计算梯度,块必须具有反向传播函数。
# 在定义我们自己的块时,由于自动微分(在 :numref:`sec_autograd` 中引入)
# 提供了一些后端实现,我们只需要考虑前向传播函数和必需的参数。
#
# 在构造自定义块之前,(**我们先回顾一下多层感知机**)
# ( :numref:`sec_mlp_concise` )的代码。
# 下面的代码生成一个网络,其中包含一个具有256个单元和ReLU激活函数的全连接隐藏层,
# 然后是一个具有10个隐藏单元且不带激活函数的全连接输出层。
#
# + origin_pos=3 tab=["tensorflow"]
import tensorflow as tf
net = tf.keras.models.Sequential([
tf.keras.layers.Dense(256, activation=tf.nn.relu),
tf.keras.layers.Dense(10),
])
X = tf.random.uniform((2, 20))
net(X)
# + [markdown] origin_pos=6 tab=["tensorflow"]
# 在这个例子中,我们通过实例化`keras.models.Sequential`来构建我们的模型,
# 层的执行顺序是作为参数传递的。
# 简而言之,`Sequential`定义了一种特殊的`keras.Model`,
# 即在Keras中表示一个块的类。
# 它维护了一个由`Model`组成的有序列表,
# 注意两个全连接层都是`Model`类的实例,
# 这个类本身就是`Model`的子类。
# 前向传播(`call`)函数也非常简单:
# 它将列表中的每个块连接在一起,将每个块的输出作为下一个块的输入。
# 注意,到目前为止,我们一直在通过`net(X)`调用我们的模型来获得模型的输出。
# 这实际上是`net.call(X)`的简写,
# 这是通过Block类的`__call__`函数实现的一个Python技巧。
#
# + [markdown] origin_pos=7
# ## [**自定义块**]
#
# 要想直观地了解块是如何工作的,最简单的方法就是自己实现一个。
# 在实现我们自定义块之前,我们简要总结一下每个块必须提供的基本功能:
#
# + [markdown] origin_pos=8 tab=["tensorflow"]
# 1. 将输入数据作为其前向传播函数的参数。
# 1. 通过前向传播函数来生成输出。请注意,输出的形状可能与输入的形状不同。例如,我们上面模型中的第一个全连接的层接收任意维的输入,但是返回一个维度256的输出。
# 1. 计算其输出关于输入的梯度,可通过其反向传播函数进行访问。通常这是自动发生的。
# 1. 存储和访问前向传播计算所需的参数。
# 1. 根据需要初始化模型参数。
#
# + [markdown] origin_pos=10
# 在下面的代码片段中,我们从零开始编写一个块。
# 它包含一个多层感知机,其具有256个隐藏单元的隐藏层和一个10维输出层。
# 注意,下面的`MLP`类继承了表示块的类。
# 我们的实现只需要提供我们自己的构造函数(Python中的`__init__`函数)和前向传播函数。
#
# + origin_pos=13 tab=["tensorflow"]
class MLP(tf.keras.Model):
# 用模型参数声明层。这里,我们声明两个全连接的层
def __init__(self):
# 调用MLP的父类Model的构造函数来执行必要的初始化。
# 这样,在类实例化时也可以指定其他函数参数,例如模型参数params(稍后将介绍)
super().__init__()
# Hiddenlayer
self.hidden = tf.keras.layers.Dense(units=256, activation=tf.nn.relu)
self.out = tf.keras.layers.Dense(units=10) # Outputlayer
# 定义模型的前向传播,即如何根据输入X返回所需的模型输出
def call(self, X):
return self.out(self.hidden((X)))
# + [markdown] origin_pos=14
# 我们首先看一下前向传播函数,它以`X`作为输入,
# 计算带有激活函数的隐藏表示,并输出其未规范化的输出值。
# 在这个`MLP`实现中,两个层都是实例变量。
# 要了解这为什么是合理的,可以想象实例化两个多层感知机(`net1`和`net2`),
# 并根据不同的数据对它们进行训练。
# 当然,我们希望它们学到两种不同的模型。
#
# 接着我们[**实例化多层感知机的层,然后在每次调用前向传播函数时调用这些层**]。
# 注意一些关键细节:
# 首先,我们定制的`__init__`函数通过`super().__init__()`
# 调用父类的`__init__`函数,
# 省去了重复编写模版代码的痛苦。
# 然后,我们实例化两个全连接层,
# 分别为`self.hidden`和`self.out`。
# 注意,除非我们实现一个新的运算符,
# 否则我们不必担心反向传播函数或参数初始化,
# 系统将自动生成这些。
#
# 我们来试一下这个函数:
#
# + origin_pos=17 tab=["tensorflow"]
net = MLP()
net(X)
# + [markdown] origin_pos=18
# 块的一个主要优点是它的多功能性。
# 我们可以子类化块以创建层(如全连接层的类)、
# 整个模型(如上面的`MLP`类)或具有中等复杂度的各种组件。
# 我们在接下来的章节中充分利用了这种多功能性,
# 比如在处理卷积神经网络时。
#
# ## [**顺序块**]
#
# 现在我们可以更仔细地看看`Sequential`类是如何工作的,
# 回想一下`Sequential`的设计是为了把其他模块串起来。
# 为了构建我们自己的简化的`MySequential`,
# 我们只需要定义两个关键函数:
#
# 1. 一种将块逐个追加到列表中的函数。
# 1. 一种前向传播函数,用于将输入按追加块的顺序传递给块组成的“链条”。
#
# 下面的`MySequential`类提供了与默认`Sequential`类相同的功能。
#
# + origin_pos=21 tab=["tensorflow"]
class MySequential(tf.keras.Model):
def __init__(self, *args):
super().__init__()
self.modules = []
for block in args:
# 这里,block是tf.keras.layers.Layer子类的一个实例
self.modules.append(block)
def call(self, X):
for module in self.modules:
X = module(X)
return X
# + [markdown] origin_pos=24
# 当`MySequential`的前向传播函数被调用时,
# 每个添加的块都按照它们被添加的顺序执行。
# 现在可以使用我们的`MySequential`类重新实现多层感知机。
#
# + origin_pos=27 tab=["tensorflow"]
net = MySequential(
tf.keras.layers.Dense(units=256, activation=tf.nn.relu),
tf.keras.layers.Dense(10))
net(X)
# + [markdown] origin_pos=28
# 请注意,`MySequential`的用法与之前为`Sequential`类编写的代码相同
# (如 :numref:`sec_mlp_concise` 中所述)。
#
# ## [**在前向传播函数中执行代码**]
#
# `Sequential`类使模型构造变得简单,
# 允许我们组合新的架构,而不必定义自己的类。
# 然而,并不是所有的架构都是简单的顺序架构。
# 当需要更强的灵活性时,我们需要定义自己的块。
# 例如,我们可能希望在前向传播函数中执行Python的控制流。
# 此外,我们可能希望执行任意的数学运算,
# 而不是简单地依赖预定义的神经网络层。
#
# 到目前为止,
# 我们网络中的所有操作都对网络的激活值及网络的参数起作用。
# 然而,有时我们可能希望合并既不是上一层的结果也不是可更新参数的项,
# 我们称之为*常数参数*(constant parameter)。
# 例如,我们需要一个计算函数
# $f(\mathbf{x},\mathbf{w}) = c \cdot \mathbf{w}^\top \mathbf{x}$的层,
# 其中$\mathbf{x}$是输入,
# $\mathbf{w}$是参数,
# $c$是某个在优化过程中没有更新的指定常量。
# 因此我们实现了一个`FixedHiddenMLP`类,如下所示:
#
# + origin_pos=31 tab=["tensorflow"]
class FixedHiddenMLP(tf.keras.Model):
def __init__(self):
super().__init__()
self.flatten = tf.keras.layers.Flatten()
# 使用tf.constant函数创建的随机权重参数在训练期间不会更新(即为常量参数)
self.rand_weight = tf.constant(tf.random.uniform((20, 20)))
self.dense = tf.keras.layers.Dense(20, activation=tf.nn.relu)
def call(self, inputs):
X = self.flatten(inputs)
# 使用创建的常量参数以及relu和matmul函数
X = tf.nn.relu(tf.matmul(X, self.rand_weight) + 1)
# 复用全连接层。这相当于两个全连接层共享参数。
X = self.dense(X)
# 控制流
while tf.reduce_sum(tf.math.abs(X)) > 1:
X /= 2
return tf.reduce_sum(X)
# + [markdown] origin_pos=32
# 在这个`FixedHiddenMLP`模型中,我们实现了一个隐藏层,
# 其权重(`self.rand_weight`)在实例化时被随机初始化,之后为常量。
# 这个权重不是一个模型参数,因此它永远不会被反向传播更新。
# 然后,神经网络将这个固定层的输出通过一个全连接层。
#
# 注意,在返回输出之前,模型做了一些不寻常的事情:
# 它运行了一个while循环,在$L_1$范数大于$1$的条件下,
# 将输出向量除以$2$,直到它满足条件为止。
# 最后,模型返回了`X`中所有项的和。
# 注意,此操作可能不会常用于在任何实际任务中,
# 我们只是向你展示如何将任意代码集成到神经网络计算的流程中。
#
# + origin_pos=34 tab=["tensorflow"]
net = FixedHiddenMLP()
net(X)
# + [markdown] origin_pos=35
# 我们可以[**混合搭配各种组合块的方法**]。
# 在下面的例子中,我们以一些想到的方法嵌套块。
#
# + origin_pos=38 tab=["tensorflow"]
class NestMLP(tf.keras.Model):
def __init__(self):
super().__init__()
self.net = tf.keras.Sequential()
self.net.add(tf.keras.layers.Dense(64, activation=tf.nn.relu))
self.net.add(tf.keras.layers.Dense(32, activation=tf.nn.relu))
self.dense = tf.keras.layers.Dense(16, activation=tf.nn.relu)
def call(self, inputs):
return self.dense(self.net(inputs))
chimera = tf.keras.Sequential()
chimera.add(NestMLP())
chimera.add(tf.keras.layers.Dense(20))
chimera.add(FixedHiddenMLP())
chimera(X)
# + [markdown] origin_pos=39
# ## 效率
#
# + [markdown] origin_pos=42 tab=["tensorflow"]
# 你可能会开始担心操作效率的问题。
# 毕竟,我们在一个高性能的深度学习库中进行了大量的字典查找、
# 代码执行和许多其他的Python代码。
# Python的问题[全局解释器锁](https://wiki.python.org/moin/GlobalInterpreterLock)
# 是众所周知的。
# 在深度学习环境中,我们担心速度极快的GPU可能要等到CPU运行Python代码后才能运行另一个作业。
#
# + [markdown] origin_pos=43
# ## 小结
#
# * 一个块可以由许多层组成;一个块可以由许多块组成。
# * 块可以包含代码。
# * 块负责大量的内部处理,包括参数初始化和反向传播。
# * 层和块的顺序连接由`Sequential`块处理。
#
# ## 练习
#
# 1. 如果将`MySequential`中存储块的方式更改为Python列表,会出现什么样的问题?
# 1. 实现一个块,它以两个块为参数,例如`net1`和`net2`,并返回前向传播中两个网络的串联输出。这也被称为平行块。
# 1. 假设你想要连接同一网络的多个实例。实现一个函数,该函数生成同一个块的多个实例,并在此基础上构建更大的网络。
#
# + [markdown] origin_pos=46 tab=["tensorflow"]
# [Discussions](https://discuss.d2l.ai/t/1826)
#
| tensorflow/chapter_deep-learning-computation/model-construction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Classes
#
# Classes allow you to define how to package data with functions to create objects. An object is an instance of a class, which contains its own data, and its own copy of functions that can operate on that data.
#
# You use classes to define objects that represent the concepts and things that your program will work with. For example, if your program managed exam results of students, then you may create one class that represents an Exam, and another that represents a Student.
class Exam:
def __init__(self, max_score=100):
self._max_score = max_score
self._actual_score = 0
def percent(self):
return 100.0 * self._actual_score / self._max_score
def setResult(self, score):
if (score < 0):
self._actual_score = 0
elif (score > self._max_score):
self._actual_score = self._max_score
else:
self._actual_score = score
def grade(self):
if (self._actual_score == 0):
return "U"
elif (self.percent() > 90.0):
return "A"
elif (self.percent() > 80.0):
return "B"
elif (self.percent() > 70.0):
return "C"
else:
return "F"
class Student:
def __init__(self):
self._exams = {}
def addExam(self, name, exam):
self._exams[name] = exam
def addResult(self, name, score):
self._exams[name].setResult(score)
def result(self, exam):
return self._exams[exam].percent()
def grade(self, exam):
return self._exams[exam].grade()
def grades(self):
g = {}
for exam in self._exams.keys():
g[exam] = self.grade(exam)
return g
# We can now create a student, and give them a set of exams that they need to complete.
s = Student()
s.addExam( "maths", Exam(20) )
s.addExam( "chemistry", Exam(75) )
# At this point, the student has not completed any exams, so the grades are all 'U'
s.grades()
# However, we can now add the results...
s.addResult("maths", 15)
s.addResult("chemistry", 62)
s.grades()
# Programming with classes makes the code easier to read, as the code more closely represents the concepts that make up the program. For example, here we have a class that represents a full school of students.
class School:
def __init__(self):
self._students = {}
self._exams = []
def addStudent(self, name):
self._students[name] = Student()
def addExam(self, exam, max_score):
self._exams.append(exam)
for key in self._students.keys():
self._students[key].addExam(exam, Exam(max_score))
def addResult(self, name, exam, score):
self._students[name].addResult(exam, score)
def grades(self):
g = {}
for name in self._students.keys():
g[name] = self._students[name].grades()
return g
# We can now create a whole school of students and manage the exams and results for all of them with some reasonably readable code :-)
school = School()
school.addStudent("Charlie")
school.addStudent("Matt")
school.addStudent("James")
school.addExam( "maths", 20 )
school.addExam( "physics", 50 )
school.addExam( "english literature", 30 )
school.grades()
# We can now add in the results of the exams, which have been returned to us by the exam markers...
englit_results = { "Charlie" : 10, "Matt" : 25, "James" : 3 }
phys_results = { "Matt" : 48, "James" : 3 }
maths_results = { "James" : 20, "Matt" : 18, "Charlie" : 4 }
# Indeed, we will do this by using a function...
def add_results(school, exam, results):
for student in results.keys():
school.addResult(student, exam, results[student])
add_results(school, "english literature", englit_results)
add_results(school, "physics", phys_results)
add_results(school, "maths", maths_results)
school.grades()
# # Exercise
#
# ## Exercise 1
#
# Here is a copy of the Morse class from the last section. Modify this class to add in a `decode` function that converts Morse code back to english. Check that this class works by seeing if `m.decode( m.encode(message) ) == message.lower()`.
class Morse:
def __init__(self):
self._letter_to_morse = {'a':'.-', 'b':'-...', 'c':'-.-.', 'd':'-..', 'e':'.', 'f':'..-.',
'g':'--.', 'h':'....', 'i':'..', 'j':'.---', 'k':'-.-', 'l':'.-..', 'm':'--',
'n':'-.', 'o':'---', 'p':'.--.', 'q':'--.-', 'r':'.-.', 's':'...', 't':'-',
'u':'..-', 'v':'...-', 'w':'.--', 'x':'-..-', 'y':'-.--', 'z':'--..',
'0':'-----', '1':'.----', '2':'..---', '3':'...--', '4':'....-',
'5':'.....', '6':'-....', '7':'--...', '8':'---..', '9':'----.',
' ':'/' }
self._morse_to_letter = {}
for letter in self._letter_to_morse.keys():
self._morse_to_letter[ self._letter_to_morse[letter] ] = letter
def encode(self, message):
morse = []
for letter in message:
morse.append( self._letter_to_morse[letter.lower()] )
return morse
def decode(self, morse):
message = []
for code in morse:
message.append( self._morse_to_letter[code] )
return "".join(message)
m = Morse()
message = "Hello World"
m.decode( m.encode(message) ) == message.lower()
# ## Exercise 2
#
# Below is a copy of the `School` class, together with a copy of the code needed to populate an object of that class with students and exam results. Edit the `School` class to add in the following functions:
#
# * `.resits()` : this should return the list of exams that each student should resit if they get a "F" or "U" grade.
# * `.prizeStudent()` : this should return the name of the student who scored the highest average percent across all of the exams.
# * `.reviseCourse(threshold)` : this should return the name of the exam that gets the lowest average score across all students, if the average score is below `threshold`.
#
# Use these functions to find out which students need to resit which exams, which student should be awarded the annual school prize, and which courses should be revised as the average mark is less than 50%.
class School:
def __init__(self):
self._students = {}
self._exams = []
def addStudent(self, name):
self._students[name] = Student()
def addExam(self, exam, max_score):
self._exams.append(exam)
for key in self._students.keys():
self._students[key].addExam(exam, Exam(max_score))
def addResult(self, name, exam, score):
self._students[name].addResult(exam, score)
def grades(self):
g = {}
for name in self._students.keys():
g[name] = self._students[name].grades()
return g
def resits(self):
r = {}
for name in self._students.keys():
student_resits = []
for exam in self._exams:
grade = self._students[name].grade(exam)
if (grade == "F" or grade == "U"):
student_resits.append(exam)
if len(student_resits) > 0:
r[name] = student_resits
return r
def prizeStudent(self):
prize_score = 0
prize_student = None
for name in self._students.keys():
avg_score = 0
for exam in self._exams:
avg_score += self._students[name].result(exam)
avg_score /= len(self._exams)
if avg_score > prize_score:
prize_score = avg_score
prize_student = name
return prize_student
def reviseCourse(self, threshold=50):
revise_course = {}
for exam in self._exams:
avg_score = 0
for name in self._students.keys():
avg_score += self._students[name].result(exam)
avg_score /= len(self._students)
if avg_score < threshold:
revise_course[exam] = avg_score
return revise_course
students = ["Charlie", "James", "Matt"]
exams = { "maths" : 20, "physics" : 50, "english literature" : 30 }
results = { "maths" : { "James" : 20, "Matt" : 18, "Charlie" : 4 },
"physics" : { "Matt" : 48, "James" : 3 },
"english literature" : { "Charlie" : 10, "Matt" : 25, "James" : 3 } }
school = School()
for student in students:
school.addStudent(student)
for exam in exams.keys():
school.addExam(exam, exams[exam])
for exam in results:
add_results(school, exam, results[exam])
school.grades()
school.resits()
school.prizeStudent()
school.reviseCourse(50)
| answers/06_classes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Helaena-Cambel/OOP-58001/blob/main/Fundamentals%20of%20Python%20(Edited).ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="UtIETdGDj2Z1"
# #Fundamentals of Python
# + [markdown] id="zrXeZuC4kYb5"
# **Python Variables**
# + colab={"base_uri": "https://localhost:8080/"} id="WyvrztfkkcwV" outputId="e9c157cc-6107-4a65-8dc3-28d14f3fef59"
x = float(1)
a, b = 0, -1
a, b, c = "Sally", "John", "Ana"
print('This is a sample')
print(a)
print(c)
# + [markdown] id="CHKfsH7VmUL2"
# **Casting**
# + colab={"base_uri": "https://localhost:8080/"} id="D0DxQ4h8mV6z" outputId="bc95119f-deac-4575-9a9e-dee3c81c3779"
print(x)
# + [markdown] id="iuMwFWxXmvQx"
# **Type() Function**
# + colab={"base_uri": "https://localhost:8080/"} id="VxrQUw6ZmzGL" outputId="2a5d710e-ef93-46cb-e856-ece995f42997"
y = "Johnny"
print(type(y))
print(type(x))
# + [markdown] id="QoJDfcKYn3_O"
# **Double Quotes and Single Quotes**
# + colab={"base_uri": "https://localhost:8080/"} id="nI58j4SQn78q" outputId="c556fab1-cfd4-49d7-888e-197609447185"
#h = "Maria"
h = 'Maria'
v = 1
V = 3
print(h)
print(v)
print(v+1)
print(V)
# + [markdown] id="spcjY6cQpJnx"
# **Multiple Variables with Different Value**
# + colab={"base_uri": "https://localhost:8080/"} id="PXlWSZk7pMx3" outputId="1c7b1690-6c74-4847-ebab-5020f3528582"
x, y, z = "one", "two", 'three'
print(x)
print(y)
print(z)
print(x, y, z)
# + [markdown] id="4RYWo2Msqr54"
# **One Value to Multiple Variables**
# + colab={"base_uri": "https://localhost:8080/"} id="eFSzqq6kqu9M" outputId="9cec7def-c29b-4527-f8ec-26c356e4493e"
x = y = z = "Stella"
print(x, y, z)
print(x)
print(y)
print(z)
# + [markdown] id="EhcwiaC3rck8"
# **Output Variables**
# + colab={"base_uri": "https://localhost:8080/"} id="nSTakKJ1rfVt" outputId="eb3b2017-c6ae-4168-911f-df39325c5bea"
x = "enjoying"
y = "Python is"
print("Python is " + x)
print(y + " " + x)
# + [markdown] id="yqa2eck1voRr"
# **Arithmetic Operations**
# + colab={"base_uri": "https://localhost:8080/"} id="XvFxG8lhvrur" outputId="c0833ec3-cb23-4db8-e98d-f9969249ef4f"
f = 2
g = 4
i = 6
print(f+g)
print(f-g)
print(f*i)
print(int(i/g))
print(3/g)
print(3%g)
print(3//g)
print(3**6)
# + [markdown] id="0YcItthPv5il"
# **Assignment Operators**
# + colab={"base_uri": "https://localhost:8080/"} id="yp6ysuTzv-CD" outputId="c4b6e5f6-348b-40a5-8a8d-e0d9a558a7aa"
k = 2
l = 3
k+=3 #same as k=k+3
print(k)
print(l>>1)
# + [markdown] id="PWYLzT4nxI9d"
# **Boolean Operators**
# + colab={"base_uri": "https://localhost:8080/"} id="FZk2n9NXxL1_" outputId="f1a5daff-17a4-423c-dd0d-9c93a53fcf15"
k = 5
l = 10
print(k>>2) #shift right twice
print(k<<2) #shift left twice
# + [markdown] id="cWzJ1HKiwec8"
# **Relational Operators**
# + colab={"base_uri": "https://localhost:8080/"} id="owTj6ygQwgwT" outputId="f17c4216-a3d7-472b-d3de-90720147c40f"
v=1
k=2
print(v>k)
print(v==k)
# + [markdown] id="O6DrV_cXxcO9"
# **Logical Operators**
# + colab={"base_uri": "https://localhost:8080/"} id="mMD4hU8_xeqe" outputId="4240d7b0-4c8f-472a-e583-452108bdfd96"
print(v<k and k==k)
print(v<k or k==v)
print(not (v<k or k==v))
# + [markdown] id="jxJU8EDqxzqK"
# **Identity Operators**
# + colab={"base_uri": "https://localhost:8080/"} id="nrc7jd8tx11i" outputId="1b8f82c9-c615-4999-86dc-805af04b63a7"
print(v is k)
print(v is not k)
| Fundamentals of Python (Edited).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:pySparkEnv]
# language: python
# name: conda-env-pySparkEnv-py
# ---
from __future__ import print_function
import csv
import datetime
import os
import json
import requests
import sys
import msgpack
from kafka import KafkaProducer
from kafka import SimpleProducer, KafkaClient
#basepath = '/root/sftp/volumes/saqn/
# +
import threading, logging, time
import multiprocessing
from kafka import KafkaConsumer, KafkaProducer
kafkaUrl = "swarm-node00.teco.edu"
kafkaUrl = "smartaqnet-dev.teco.edu"
#kafkaUrl = "172.16.31.103"#smartaqnet-dev.teco.edu"
class Producer(threading.Thread):
def __init__(self):
threading.Thread.__init__(self)
self.stop_event = threading.Event()
def stop(self):
self.stop_event.set()
def run(self):
print("Starting producer")
producer = KafkaProducer(bootstrap_servers=kafkaUrl+':9092')
print("Producer started")
while not self.stop_event.is_set():
print("someThing")
print(producer.send('Baka', "msg"))
time.sleep(2)
producer.close()
class Consumer(multiprocessing.Process):
def __init__(self):
multiprocessing.Process.__init__(self)
self.stop_event = multiprocessing.Event()
def stop(self):
self.stop_event.set()
def run(self):
print("Starting consumer")
consumer = KafkaConsumer(bootstrap_servers= kafkaUrl + ':49092',
auto_offset_reset='latest',
consumer_timeout_ms=1000)#,value_deserializer=msgpack.unpackb)
print("Consumer started")
consumer.subscribe(['Observations'])
while not self.stop_event.is_set():
for message in consumer:
print(message)
if self.stop_event.is_set():
break
consumer.close()
# +
tasks = [
#Producer(),
Consumer()
]
for t in tasks:
t.start()
# +
for t in tasks:
t.stop()
# -
for t in tasks:
t.stop()
docker run \
--net=host \
--rm confluentinc/cp-kafka \
bash -c "seq 42 | kafka-console-producer --broker-list localhost:29092 --topic bar && echo 'Produced 42 messages.'"
docker run \
--net=host \
--rm \
confluentinc/cp-kafka \
kafka-console-consumer --bootstrap-server localhost:29092 --topic bar --new-consumer --from-beginning --max-messages 42
| KrigingExample/3. PFF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import time
import random
from sklearn.preprocessing import MaxAbsScaler, StandardScaler, Normalizer, LabelEncoder, MinMaxScaler, RobustScaler, QuantileTransformer, PowerTransformer
from IPython import display
from pandas.plotting import scatter_matrix
import seaborn as sns
import matplotlib.pylab as pylab
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import sys
import warnings
warnings.filterwarnings('ignore')
# %matplotlib inline
sns.set_style('white')
pylab.rcParams['figure.figsize'] = 12, 8
# -
train = pd.read_csv('train.csv')
train['source'] = 'train'
test = pd.read_csv('test.csv')
test['source'] = 'test'
passngerID = test[['PassengerId']]
data = pd.concat([train, test], axis= 0, sort= False)
# # Data Exploring
data.head()
# ### Visualizing null values.
sns.heatmap(data.isnull(), yticklabels=False, cbar=False, cmap= 'viridis')
# - Fare column has only one null value.<br/>
# - Age column has many null values.<br/>
# - Cabin column has a majority of null values.<br/>
# - Survived column has null values for the test data.
data.info()
# ### Is data balanced?
sns.countplot(data = data, x= 'Survived')
# ### Which is the most survived gender?
sns.countplot(data = data, x= 'Survived', hue= 'Sex')
plt.legend(loc =(1.1,0.9)),
# ### Does first class have more survival rate?
sns.countplot(data = data, x='Survived', hue='Pclass')
# ### The distribution of passengers' age.
sns.distplot(data['Age'].dropna(), kde = False, bins = 35)
# ### The distribution of number of siblings.
sns.countplot(x = 'SibSp', data = data)
# ### Number of passenger's in each class.
sns.countplot(data= data.dropna(), x='Pclass')
# ### Proportion of each gender in different classes.
sns.countplot(data= data, x='Pclass', hue= 'Sex')
# ### Ticket fare for each class.
sns.boxplot(data= data.dropna(), x='Pclass', y= 'Fare')
data.describe()
# # Data cleaning
# ### Fill missing values in Age with the median age for the corresponding class
class_mean_age = data.pivot_table(values='Age', index='Pclass', aggfunc='median')
null_age = data['Age'].isnull()
data.loc[null_age,'Age'] = data.loc[null_age,'Pclass'].apply(lambda x: class_mean_age.loc[x] )
data.Age.isnull().sum()
# ### Fill the missing value in Fare with the median fare for the corresponding class.
class_mean_fare = data.pivot_table(values= 'Fare', index= 'Pclass', aggfunc='median')
null_fare = data['Fare'].isnull()
data.loc[null_fare, 'Fare'] = data.loc[null_fare, 'Pclass'].apply(lambda x: class_mean_fare.loc[x] )
data.Fare.isnull().sum()
# ### Fill the missing values in Embarked with the most common port for corresponding class.
data.Embarked.value_counts()
data['Embarked'] = data.Embarked.fillna('S')
data.Embarked.isnull().sum()
# # Feature Engineering
# ## Create New features
# ### Create a new feature with the title of each passenger.
data['Title'] = data.Name.apply(lambda x : x[x.find(',')+2:x.find('.')])
data.Title.value_counts()
# We can notice that only 4 titles have significant frequency and the others are repeated only 8 time or less.<br/> So, we will combine all titles with small frequency under one title (say, Other).
rare_titles = (data['Title'].value_counts() < 10)
data['Title'] = data['Title'].apply(lambda x : 'Other' if rare_titles.loc[x] == True else x)
# ### Create a new feature for the family size
# This feature combines the number of siblings and parents/children (SibSp and Parch) +1 (The passenger himself).
data['FamilySize'] = data['SibSp'] + data['Parch'] + 1
# ### Create a new feature to indicate whether the passenger was alone.
data['IsAlone'] = 0
data['IsAlone'].loc[ data['FamilySize'] == 1] = 1
# ### Create a new feature by discretizing Age into buckets/bins
# Age is discretized into 4 bins coresponding to 4 stages of human life:<br/>
# 1. Childhood.
# 2. Adolescence.
# 3. Adulthood.
# 4. Old Age. <br/>
# Check this link for more details: https://bit.ly/2LkPFPf
data['AgeBins'] = 0
data['AgeBins'].loc[(data['Age'] >= 11) & (data['Age'] < 20)] = 1
data['AgeBins'].loc[(data['Age'] >= 20) & (data['Age'] < 60)] = 2
data['AgeBins'].loc[data['Age'] >= 60] = 3
# ### Create new feature by discretizing Fare into 4 buckets/bins based on quantiles.
data['FareBins'] = pd.qcut(data['Fare'], 4)
# ### Drop unused columns from data.
# 1. Some features are expected to not have effect of the classification such as PassengerId, Name and Ticket. <br/>
# 2. Also some futures have too much missing values such as the Cabin which render it useless.
# 3. We'll also drop the original features we used to create the new features because there will be high correlation between these features which may confuse the model about feature importance.
data.columns
data.drop(columns=['PassengerId','Name','Ticket', 'Cabin', 'Age', 'Fare', 'SibSp', 'Parch'], inplace= True)
# ## Convert qualitative features into numeric form.
# ### Convert categorical features (Embarked, Sex, Title) to numerical features and drop one dummy variable for each.
data = pd.get_dummies(
data, columns=['Embarked', 'Sex', 'Title'], drop_first=True)
# ### Convert qualitative ordinal features (FareBins) into numeric form.
label = LabelEncoder()
data['FareBins'] = label.fit_transform(data['FareBins'])
data.head(7)
# ## Splitting Data back to train/test sets.
#Final train data
train = data[data.source == 'train'].drop(columns = ['source']).reset_index(drop=True)
test = data[data.source == 'test'].drop(columns = ['source','Survived']).reset_index(drop=True)
train['Survived'] = train.Survived.astype('int64')
# ## Rescaling features using different scalers
# ### Normalizing numeric features (Age, SibSp, Parch, FamilySize and Fare).
# We will try the following scalers and we'll select the best one:
# 1. MinMaxScaler
# 2. MaxAbsScaler
# 3. StandardScaler
# 4. RobustScaler
# 5. Normalizer
# 6. QuantileTransformer
# 7. PowerTransformer
feature_to_scale = ['FamilySize']
scalers = {}
for i in feature_to_scale:
scaler = RobustScaler()
scaler.fit(train[[i]])
train[i] = scaler.transform(train[[i]])
test[i] = scaler.transform(test[[i]])
scalers.update({i:scaler})
scalers
# ### Exporting modified train/test data to external file.
#Final Test data
train.to_csv('train_modified.csv', index = False)
test.to_csv('test_modified.csv', index = False)
passngerID.to_csv('ID.csv', index = False)
| Titanic-Classification-Problem-EDA&Preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# # Problem Set 3, due Wednesday, May 3rd, 5:30pm.
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ## Introduction to the assignment
#
# For this assignment, you will be using data from the [Progresa
# program](http://en.wikipedia.org/wiki/Oportunidades), a government
# social assistance program in Mexico. This program, as well as the
# details of its impact, are described in the paper "[School subsidies
# for the poor: evaluating the Mexican Progresa poverty
# program](http://www.sciencedirect.com/science/article/pii/S0304387803001858)",
# by <NAME> (available on Canvas). The data
# (progresa_sample.csv.gz) is available on canvas
# in files/data.
#
# Please read the paper to familiarize yourself with
# the PROGRESA program before beginning this problem set, so you have a
# rough sense of where the data come from and how they were
# generated. If you just proceed into the problem set without
# understanding Progresa or the data, it will be very difficult!
#
# The goal of this problem set is to implement some of the basic
# econometric techniques that you are learning in class to measure the
# impact of Progresa on secondary school enrollment rates, in particular
# differences-in-differences. Your task is to **estimate the impact of
# _progresa_ subsidies on the school attendance**. Note: this asks to
# estimate a causal effect.
#
# The timeline of the program was:
#
# * Baseline survey conducted in 1997
# * Intervention begins in 1998, "Wave 1" of data collected in 1998
# * "Wave 2 of data" collected in 1999
# * Evaluation ends in 2000, at which point the control villages were treated.
#
# When you are ready, download the progresa_sample.csv data from
# Canvas. The data are actual data collected to evaluate the impact of
# the Progresa program. In this file, each row corresponds to an
# observation taken for a given child for a given year. There are two
# years of data (1997 and 1998), and just under 40,000 children who are
# surveyed in both years. For each child-year observation, the
# following variables are collected:
#
# | Variable name | Description|
# |------|------|
# |year |year in which data is collected
# |sex |male = 1|
# |indig |indigenous = 1|
# |dist_sec |nearest distance to a secondary school|
# |sc |enrolled in school in year of survey (=1)|
# |grc |grade enrolled|
# |fam_n |family size|
# |min_dist | min distance to an urban center|
# |dist_cap | min distance to the capital|
# |poor | poor = "pobre", not poor = "no pobre"|
# |progresa |treatment = "basal", control = "0"|
# |hohedu |years of schooling of head of household|
# |hohwag |monthly wages of head of household|
# |welfare_index| welfare index used to classify poor|
# |hohsex |gender of head of household (male=1)|
# |hohage |age of head of household|
# |age |years old|
# |folnum |individual id|
# |village| village id|
# |sc97 |enrolled in school in 1997 (=1) |
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ## Part 1: Descriptive analysis
#
# ### 1.1 Summary Statistics
#
# Report summary statistics (mean, standard deviation, and number of missings) for all of
# the demographic variables in the dataset (i.e., everything except
# year, folnum, village). Present the actual levels for 'progresa' and
# 'poor'. Do these fit with the documentation above?
#
# Present these in a single table alphabetized
# by variable name. Do NOT simply expect the grader to scroll through
# your output!
# + ein.tags="worksheet-0" slideshow={"slide_type": "-"}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import statsmodels.formula.api as smf
import scipy.stats as stats
# +
# load it up
df = pd.read_csv('progresa_sample.csv')
# this changes the 'poor' column so that 'pobre' equals 1
df['poor'] = np.where(df['poor'] == "pobre", 1, 0)
# this changes the 'progresa' column so 'basal' equals 1
df['progresa'] = np.where(df['progresa'] == "basal", 1, 0)
# this produces summary stats and then reindexes alphabetically,
# keeping only count, mean, and std (for use later)
summary = df.describe().reindex(sorted(df.columns), axis=1)
summary = summary[0:3]
# this cleans up some stuff, dropping unnecessary columns
summary.drop(['year', 'folnum', 'village'], axis=1, inplace=True)
# and this collects the number of null values
summary.loc['null values'] = df.isnull().sum()
display = summary[1:].T
display
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ### 1.2 Differences at baseline?
# Now let's investigate the differences in baseline. Are the baseline (1997) demographic characteristics **for the poor**
# different in treatment and control villages?
#
# I suggest you to use a T-Test to
# determine whether there is a statistically significant difference in
# the average values of each of the variables in the dataset. Focus only
# on the data from 1997 for poor.
#
# Present your results in a single table with the following columns and 14 (or so) rows:
#
# | Variable name | Average value (Treatment villages) | Average value (Control villages) | Difference (Treat - Control) | p-value |
# |------|------|------|------|------|
# |Male|?|?|?|?|
#
# + ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# start by subset!
baseline = df[df.year == 97][df.poor == 1]
baseline_treat = baseline[baseline.progresa == 1]
baseline_control = baseline[baseline.progresa == 0]
# let's drop the same coumns we did above
# we can get rid of these because producing average values for them
# won't tell us anything at all
#
# we can also drop poor (=1) and sc97/grc97 (same as sc/grc in this context)
baseline.drop(['year', 'folnum', 'village', 'poor', 'sc97', 'grc97'], axis=1, inplace=True)
baseline = baseline.dropna().groupby('progresa').mean().T.reset_index()
baseline.columns = ['variable' ,'control', 'treatment']
baseline['difference (means)'] = baseline.treatment - baseline.control
# here's the actual t-test calculations... sorry for the loop ott, i know how you hate them!
varlist = list(baseline.variable)
test = [ stats.ttest_ind(baseline_control[i], baseline_treat[i], nan_policy='omit') for i in varlist ]
# and let's make it into a dataframe
ttest = pd.DataFrame(index = varlist, data = test)
# and add a column that tells us about significance
ttest['significance'] = ttest.pvalue < 0.05
# renaming the ttest dataframe so we can combine it with the baseline info
ttest = ttest.reset_index()
ttest.columns = ['variable', 't-stat', 'pvalue', 'significance']
result = pd.merge(baseline, ttest, on='variable')
# ...and final cleanup
result.columns = ['Variable Name' ,'Average Value (Control Villages)', \
'Average Value (Treatment Villages)', 'Difference (Treat - Ctrl)',\
'T-Statistic', 'P-Value', 'Significance?']
result
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ### 1.3 Interpretation
#
# * A: Are there statistically significant differences between treatment and control villages as baseline?
#
# *Yes, there are. See the 'Significance?' column above for entries labeled TRUE.*
#
# * B: Why does it matter if there are differences at baseline?
#
# *The goal of a t-test is to determine if there are differences between the means of two distributions, so that we can compare them adequately. As part of the t-test, we assume that the distributions are independent. If there are differences at baseline (i.e. if some statistical significance is shown), that means that it is harder to assert that some of the variables are independent.*
#
# * C: What does this imply about how to measure the impact of the treatment?
#
# *If we see statistical differences between populations at baseline, it means that the segregation method used to divide those two populations into different groups is flawed. If this is the case, this means we cannot assert that our experiment is truly random, and the outcomes may be called into question. We should probably use stratified random sampling in order to obtain a more representative sample, rather than just random. It may better address categories that "overlap" within our population.*
#
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ### 1.4 Graphical exploration, part 1
#
# For each level of household head education, compute the average
# enrollment rate in 1997. Create a scatterplot that shows this
# relationship. What do you notice?
# + ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# using the 'baseline' variable from above (which subsets df.year==97), we can
# group by head-of-household education, and then get the mean of enrollment
ed = df[df.year == 97][df.poor == 1]
education = ed.groupby(['hohedu']).sc.mean().reset_index()
# then plot it
plt.scatter(education.hohedu, education.sc)
plt.title("education vs. enrollment")
plt.show()
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# *It seems that, with a couple of exceptions, there's an upward trajectory from 0 years of head-of-household education until 18 years (the max), while the level of enrollment drops dramatically (by half!) at 20 years. I would guess this has something to do with school subsidies not being as necessary if the head of household has higher education (read: gotten that oh-so-important credential). If you have more education, you most likely are at a lower risk for poverty, and thus would not have to enroll your children in this particular social program.*
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ### 1.5 Graphical exploration, part 2
#
# Create a histogram of village enrollment rates **among poor households
# in treated villages**, before and after treatment. Specifically, for
# each village, calculate the average rate of enrollment of poor
# households in treated villages in 1997, then compute the average rate
# of enrollment of poor households in treated villages in 1998. Create
# two separate histograms showing the distribution of these average
# enrollments rates, one histogram for 1997 and one histogram for
# 1998. On each histogram, draw a vertical line that intersects the
# x-axis at the average value (across all households). Does there appear
# to be a difference? Is this difference statistically significant?
# + ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# let's make a new variable really quick, and group the important information
village = df.groupby(['village', 'year', 'poor', 'progresa']).sc.mean().reset_index()
# then let's do some smart subsetting, by year
ninetyseven = village[village.year == 97]
# then by poor and treatment
ninetyseven = ninetyseven[ninetyseven.poor == 1][ninetyseven.progresa == 1]
# same thing here...
ninetyeight = village[village.year == 98]
ninetyeight = ninetyeight[ninetyeight.poor == 1][ninetyeight.progresa == 1]
# and then we plot, with the '97 data first
plt.figure(figsize=(16, 9))
# we set the axes to make sure the graphs are the same aspect ratio and scale
plt.xlim([0.4,1])
plt.ylim([0,100])
plt.hist(ninetyseven.sc)
plt.title('enrollment before')
plt.axvline(x=ninetyseven.sc.mean(), alpha=0.5, color='red')
plt.show()
# and now the '98
plt.figure(figsize=(16, 9))
plt.xlim([0.4,1])
plt.ylim([0,100])
plt.hist(ninetyeight.sc)
plt.title('enrollment after')
plt.axvline(x=ninetyeight.sc.mean(), alpha=0.5, color='red')
plt.show()
difference = ninetyeight.sc.mean() - ninetyseven.sc.mean()
# we're trying to get a t-test for the means of two independent samples, so...
ttest = stats.ttest_ind(ninetyseven.sc, ninetyeight.sc, nan_policy='omit')
print("difference between means (pct):", difference\
, "\n\nt-statistic:", ttest[0],"\np-value:", ttest[1])
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# * Is there a difference? Is the difference statistically significant?
#
# *There appears to be a difference between the average values for households across years, of a couple percent (in actuality, 1.56%). If the null hypothesis is that there is no difference in enrollment, the alternative is that there would be greater enrollment. Calculating the t-test and observing the p-values lets us know that the means are really close to but outside of the critical range (-2, 2), and that the difference between the two of these means is statistically significant.*
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ## Part 2: Measuring Impact
#
# Our goal is to estimate the causal impact of the PROGRESA program on
# the social and economic outcomes of individuals in Mexico. We will
# focus on the impact of the program on the poor (those with
# poor=='pobre'), since only the poor were eligible to receive the
# PROGRESA assistance.
#
# ### 2.1 Cross-Sectional Estimator: T-test
#
# Begin by estimating the impact of Progresa using "simple differences."
# Restricting yourself to data from 1998 (after treatment), calculate
# the average enrollment rate among **poor** households in the Treatment
# villages and the average enrollment rate among **poor** households in
# the control villages. Use a t-test to determine if this difference is
# statistically significant. What do you conclude?
# + ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# let's reindex the dataframe a little bit, grouping by
# poor, year, and treatment
csecont = df[df.poor == 1][df.year == 98][df.progresa == 0]
csetreat = df[df.poor == 1][df.year == 98][df.progresa == 1]
# and then run a t-test, omitting NAs
csetstat = stats.ttest_ind(csetreat.sc, csecont.sc, nan_policy='omit')
print("the mean for the cse control group is",\
csecont.sc.mean(), "\nthe mean for the cse treatment group is", csetreat.sc.mean()\
,"\nthe t-statistic is", csetstat.statistic, "and the p-value is", csetstat.pvalue)
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# *After performing a cross-sectional estimator, we see that the t-statistic is outside of the critical range, and the p-value suggests significance. The difference between means is about 4%, and it appears to be a true difference (the treatment and control means are different). Practically we can conclude that Progresa appears to help poor people.*
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ### 2.2 Cross-Sectional Estimator: Regression
#
# Estimate the effects of Progresa on enrollment using a regression
# model, by regressing the 1998 enrollment rates **of the poor** on
# treatment assignment. Do not include any additional control
# variables.
# Discuss the following:
#
# * Based on this model, how much did Progresa increase the likelihood of a child enrolling?
# * How does your regression estimate compare to your t-test estimate from part 2.1?
# * Based on this regression model, can we reject the null hypothesis that the treatment effects are zero?
# * What is the counterfactual assumption underlying this regression?
#
# Note: I recommend to use statsmodels.formula.api as this allows to use
# convenient R-style formulas and provedes nice summaries of the results.
# + ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# Your code here
regress = df[df.year == 98][df.poor == 1]
m = smf.ols(formula = "sc~progresa", data=regress).fit()
print(m.summary())
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# * Based on this model, how much did Progresa increase the likelihood of a child enrolling?
#
# *Looking at the predicted slope coefficient for 'progresa', which is interpreted as the mean change of the response variable for one unit of change in the predictor variable while holding other predictors in the model constant (what a mouthful!), it appears that progresa creates a change of about 3.9% of that unit. It appears to be a statistically significant difference as well, which is good for us, and has minimal error.*
#
# * How does your regression estimate compare to your t-test estimate from part 2.1?
#
# *It is approximately the same for both the regression coefficients and the t-test estimate. We take the intercept and add in the effect size for progresa, and get ~0.84, with a t-statistic of 8.359.*
#
# * Based on this regression model, can we reject the null hypothesis that the treatment effects are zero?
#
# *Yes, we can reject the null hypothesis that treatment effects are zero. There is some statistical significance, in terms of the p-value threshold.*
#
# * What is the counterfactual assumption underlying this regression?
#
# *The counterfactual assumption underlying this regression is that if a member of the target population (i.e. a poor village person from 1998) recieves treatment (i.e. a stimulus that supports going to school) that school enrollment will change, and there would be no other external factors that could cause it to change.*
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ### 2.3 Multiple Regression
#
# Re-run the above regression estimated but this time include a set of
# relevant control variables. Include, for instance, age, distance to a secondary
# school, gender, education of household head, welfare index,
# indigenous, etc.
#
# * How do the controls affect the point estimate of treatment effect?
# * How do the controls affect the standard error on the treatment effect?
# * How do you interpret the differences (or similarities) between your estimates of 2.2 and 2.3?
# + ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# we can reuse the same data variable as above, but let's change model variable
n = smf.ols(formula = \
"sc~progresa + dist_sec + grc + fam_n + indig+ min_dist\
+ dist_cap + hohedu + hohwag + hohsex + hohage + age",\
data=regress).fit()
print(n.summary())
# -
# * How do the controls affect the point estimate of treatment effect?
#
# *We see that the addition of additional controls decreases the point estimate of the treatment effect slightly; and the t-statistic threshold increases slightly. Overall the results become more significant.*
#
# * How do the controls affect the standard error on the treatment effect?
#
# *The standard error decreases slightly between the two different models.*
#
# * How do you interpret the differences (or similarities) between your estimates of 2.2 and 2.3?
#
# *When you add a variable to a regression model, you are controlling for it, which means you are keeping it constant. If all of the independent variables are uncorrelated, then the coefficients (the betas) of the variable should not change. However, in real life, it is often hard to isolate variables to that degree in observations (especially like the real-world situation of the Progresa study), which suggests that you have to take into account intersectional effects.*
#
# *Simply, if you ask a different question, you get a different answer. Between questions 2.2 and 2.3 we are asking different questions--though they are quite similar, we come to different results.*
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ### 2.4 Difference-in-Difference, version 1 (tabular)
#
# Thus far, we have computed the effects of Progresa by estimating the
# difference in 1998 enrollment rates across villages. An alternative
# approach would be to compute the treatment effect using a
# difference-in-differences framework.
#
# Begin by estimating the average treatment effects of the program for poor households using data from 1997 and 1998. Specifically, calculate:
#
# a) the difference (between 1997 and 1998) in enrollment rates among poor households in treated villages
#
# b) the difference (between 1997 and 1998) in enrollment rates among poor households in control villages.
#
# c) The difference between these two differences is your estimate.
#
# * What is your estimate of the impact, and how does it compare to your earlier (simple difference) results?
#
# * What is the counterfactual assumption underlying this estimate?
#
# + ein.tags="worksheet-0" slideshow={"slide_type": "-"}
diff = df[df.poor == 1].groupby(['year', 'progresa']).sc.mean()
diff
treat = diff[3] - diff[1]
cont = diff[2] - diff[0]
diffindiff = treat - cont
print("a (treatment difference):",treat, "\nb (control difference) :",\
cont,"\nc (diff-in-diff) :", diffindiff)
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# * What is your estimate of the impact, and how does it compare to your earlier (simple difference) results?
#
# *Using the tabular method, it appears that the effect size should be around 0.031. This is really close to the multi-regression and significantly lower than the cross-sectional estimator regression. Diff-in-diff is more precise, after all. :)*
#
# * What is the counterfactual assumption underlying this estimate?
#
# *Generally, the logic underpinning the counterfactual is that we cannot observe both treatment and control in the same population. The counterfactual assumption underlying the diff-in-diff estimator is that there are separate populations (poor people that got treatment from progresa and those that didn't) that we care about across the two years, and that if it weren't for the treatment, we would not see the results that we did.*
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ### 2.5 Difference-in-Difference, version 2 (regression)
#
# Now use a regression specification to estimate the average treatment
# effects of the program in a difference-in-differences
# framework. Include at least 5 control variables.
#
# * What is your estimate of the impact of Progresa? Be very specific in
# interpreting your coefficients and standard errors, and make sure to
# specify exactly what units you are measuring and estimating.
#
# * How do these estimates of the treatment effect compare to the
# estimates based on the simple difference?
#
# * How do these estimates compare to the difference-in-difference
# estimates from 2.4 above? What accounts for these differences?
#
# * What is the counterfactual assumption underlying this regression?
# + ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# in order to do dif-in-diff we have to figure out a
# variable that addresses the intersection effectively.
# so lets use our good old dataframe from above
difregress = df
# and then a trick from ott in class, to resample year as T/F
difregress['time'] = (df.year > 97) + 0
# and then subset by poor
difregress = difregress[df.poor == 1]
p = smf.ols(formula = \
"sc~progresa + progresa*time + dist_sec\
+ min_dist + dist_sec*indig + indig + fam_n + hohedu + hohsex + hohage",\
data=difregress).fit()
print(p.summary())
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# * What is your estimate of the impact of Progresa? Be very specific in
# interpreting your coefficients and standard errors, and make sure to
# specify exactly what units you are measuring and estimating.
#
#
# * How do these estimates of the treatment effect compare to the
# estimates based on the simple difference?
#
# *The impact here is lower than the estimates based on simple difference. This is because simple difference is 'simple'. Diff-in-diff is a decidedly more precise way to answer the question, and allows us to introduce interaction terms which produce more granular results.
#
# * How do these estimates compare to the difference-in-difference
# estimates from 2.4 above? What accounts for these differences?
#
# *In this question, we operationalize treatment and time (year of treatment) differently, looking at the intersection between the two variables. The intersectional effect is almost the same as the diff-in-diff from above, changing only because of the other control variables we decide to add. The ones I picked, for example, bring the regression coefficients to almost exactly what we got for the diff-in-diff above.*
#
# * What is the counterfactual assumption underlying this regression?
#
# *The counterfactual assumption is similar to the question above--that is, the treatment and control groups would have been the same, if not for our inclusion of treatment, which produces the observed trends to the degree that we see them. In this case, we would assume normally that the factors relating to the head-of-household, the size of the family, whether or not a family is indigenous, the distance from a metro/capital center, and treatment over time would not have "done anything" to our population--our two separate populations would be the same, but for the inclusion of the 'progresa' treatment.*
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ### 2.6 Summary
#
# * Based on all the analysis you have undertaken to date, do you
# believe that Progresa had a causal impact on the enrollment rates of
# poor households in Mexico? Explain by referring to the methods and
# results above.
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# This is a complicated question to answer. I do believe that we can infer some kind of relationship between Progresa-as-treatment and the increase in enrollment of students in school. In that sense, there is no room to disagree: it has been effective on paper. We saw the p-values and the t-statistics above that suggest significance and results outside of the "normal" range. We also saw that, as we used more applied methods (specifically difference-in-difference), our answer for the effect size got more precise. Further, after actually reading the prospectus of the study, there are also some interesting "game theory" controls in order to prevent people from gaming the system and just taking the money. Also effective. But we have to consider other factors as well. Is what we see above enough to judge it effective on a social level, or to justify its existence as policy?
#
# Overall, I would want to look at the control and treatment groups with a more precise lens. We saw above that there were some statistically significant differences between control and treatment groups, which means that the baseline analysis may not have been as reliable as it could have been.The equally-large effect of 'indigenous-ness' seems to suggest that maybe there's other sorts of social inequity that Progresa doesn't effectively control for.
#
# The humanist in me believes that Progresa is a good thing. I believe that education is unquantifiably good for all people. However, the economist (or perhaps burgeoning econometricist) is concerned with the amount of money spent both by the government on this project, and the effect size. Is a 0.03 increase in enrollment good on a population level enough to justify the spending on the program? I assume so (since the government was probably pitched this project on a possible effect size range from the onset), but I cannot say for certain.
#
# The assumption undergirding this entire study is that direct monetary intervention is the thing that is affecting change--without Progresa, this kind of impact would not occur. What about the case that global poverty is decreasing and the standard of living is increasing, year-over-year for a long time? Perhaps we would have observed this effect over the '97-'98 period anyway, through some kind of external confounding effect (did NAFTA change financial arrangements and flows between rural poor and urban workers in Mexico--again, not sure?).
#
# It seems that in order to judge this program effective, we have to be students of statistics, history, politics, game theory, and maybe even Spanish culture. I'm not any/all of those things, so I would defer to the numbers and simply say it is effective. To what degree, I'm not sure.
| mccraney-kevin-ps03.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="0xys3CQ_JkK4" colab_type="text"
# **Table of Contents**
# 1. [Introduction to Python](#introduction-to-Python)
# 2. [First Steps](#first-steps)
# 3. [Python Variables and Data types](#python-variables-and-data-types)
# 4. [Python Operations](#python-operations)
# 5. [Strings and their Methods](#strings-and-their-methods)
# 6. [Collections](#collections)
# 7. [Conditions](#conditions)
# 8. [Functions](#functions)
# 9. [Let's Code Together](#lets-code-together)
# 10. [Further Reading](#further-reading)
#
# + [markdown] id="VLGQOXoJJkK5" colab_type="text"
# <a id='introduction-to-Python'></a>
# ## Introduction To Python
# + [markdown] id="mBI7mBUBJkK7" colab_type="text"
# **Python** is a programming language created by <NAME> in the late 1980s and is currently the [fastest-growing](https://insights.stackoverflow.com/survey/2019) major programming language.
#
# **Python** is Open Source and has a wide variety of applications such as:
# 1. Artificial Intelligence / Machine Learning
# - SciPi
# - NumPy
# - Pandas
# - PyTorch
# 2. Hardware and Micro-controllers
# - Raspberry Pi
# - MicroPython
# - CircuitPython
# 3. Web Development
# - Django
# - Flask
# 4. Scripting
# - Dev Ops Configuration scripts
# 5. Mathematics
# + [markdown] id="viNZNwHaJkK8" colab_type="text"
# <a id='first-steps'></a>
# ## First Steps
# + [markdown] id="iYpBMnXnJkK9" colab_type="text"
# **Printing**
#
# To print a python script we use the print command: print( )
# + id="wTk8EjngJkK-" colab_type="code" colab={}
print("Hello World. I am an awesome python programmer!")
# + id="pVcbJltWJkLE" colab_type="code" colab={}
print(1234567890)
# + [markdown] id="exqiW6V0JkLI" colab_type="text"
# **Comments** are complete sentences that are for you or anyone reading your code. Python's interpreter won't process your comments.
# + id="wyu_ShzmJkLJ" colab_type="code" colab={}
# I am a block comment
# + id="QNLOLaXQJkLN" colab_type="code" colab={}
"""
I am a docstring. I
can be oneline or multiline.
Unless I am used as a docstring, I am basically ignored!
Check me out!
"""
print(10)
# + [markdown] id="4F7-oL-cJkLQ" colab_type="text"
# Note:
# [PEP8](https://www.python.org/dev/peps/pep-0008/#block-comments), Python's official style guide, favours using block comments.
# + id="AqI4rfw7xQAI" colab_type="code" colab={}
#######################################################
################### Challenge 1 #######################
#######################################################
# Print the name of your city
# + [markdown] id="27UJaI72JkLT" colab_type="text"
# <a id='python-variables-and-data-**types**'></a>
# ## Python Variables and Data types
# + [markdown] id="sHWWSt8UJkLU" colab_type="text"
# ### Variables
# + [markdown] id="84OYS1kQJkLV" colab_type="text"
# Python Variables allow us to store information and give it a label that we can use to retrieve that information later. Unlike other programming languages, Python has no command for declaring a variable.
# + id="JjpmYugSJkLW" colab_type="code" colab={}
x = 7
y = "Netherlands"
print(x)
print(y)
# + [markdown] id="BfPenK-dJkLb" colab_type="text"
# Python variables are generally in lower case (Snake case) and separated by underscores.
# + id="ufcb5bkDJkLc" colab_type="code" colab={}
my_first_name = "Chidinma"
print(my_first_name)
# + [markdown] id="7agypCChJkLk" colab_type="text"
# ### Data Types
# + [markdown] id="zkB4ZnKDJkLm" colab_type="text"
# #### Numbers
# + [markdown] id="9sbzOqP2JkLo" colab_type="text"
# There are three different types of numbers in Python: **int** for Integer, **Float**, and **Complex**.
# + id="0kccXz-6JkLp" colab_type="code" colab={}
# These are integers
x = 10
y = -19
z = 0
# + id="b_0TBWC9JkLt" colab_type="code" colab={}
# These are floats
x = 3.141592653589793238462643383279502884197169
y = -10.223
z = 0.1
# anything with a "."
# + id="KTwh4uh2JkLy" colab_type="code" colab={}
# This is a complex number
x = 42j
# a + j*b
# in Python we use j instead of i
# + [markdown] id="D7zruVlPJkL3" colab_type="text"
# #### Boolean
# + [markdown] id="SY2hVN_yJkL3" colab_type="text"
# In Python, Booleans are of type bool. Booleans are True and False
# + id="nFcoSwZBJkL4" colab_type="code" colab={}
x = True
print(x)
# + id="7_L2O5kQJkL9" colab_type="code" colab={}
y = False
print(y)
# + [markdown] id="9efSgZ8uJkMA" colab_type="text"
# Surprisingly, the boolean types True and False are also numbers under the hood. So we can actually do things like add or subtract. We'll see this later in this lesson.
#
# * True is 1 under the hood.
# * False is 0 under the hood.
# + [markdown] id="eWItCyZqJkMC" colab_type="text"
# #### Strings
# + [markdown] id="SI7cBTCyJkME" colab_type="text"
# Strings in python are surrounded by either single quotation marks, or double quotation marks.
# + id="pCbdByAvJkMG" colab_type="code" colab={}
x = 'hello'
y = "hello"
print(x)
print(y)
# + [markdown] id="pkEeDByMJkML" colab_type="text"
# **Concatenating Strings**
#
# Strings can be concatenated using **'+'**
# + id="b7JfWMerJkMO" colab_type="code" colab={}
salutation = "Hello"
first_name = "Max"
last_name = "James"
greeting = salutation + first_name + last_name
print(greeting)
# + id="AzA-IrtBJkMU" colab_type="code" colab={}
greeting = salutation + ", " + first_name + " " + last_name + "! " + "It's nice to finally meet you."
print(greeting)
# + [markdown] id="tDeykMVuJkMb" colab_type="text"
# To use the same type of quote within a string, that quote needs to be escaped with a \ - backwards slash.
# + id="hSq42cgpJkMd" colab_type="code" colab={}
reminder = 'It\'s my mom\'s birthday tomorrow'
print(reminder)
# + [markdown] id="G_Zk3BAhJkMg" colab_type="text"
# Mixed quotes can be used instead in a Python string without escaping.
# + id="BS4TvHDTJkMh" colab_type="code" colab={}
reminder = "It's my mom's birthday tomorrow"
print(reminder)
# + [markdown] id="dZCoCjUOJkMl" colab_type="text"
# **Common String Errors**
# + id="7PoMgK2hJkMm" colab_type="code" colab={}
#######################################################
################### Challenge 2 #######################
#######################################################
# Come up with possible solutions for the error below
# Trying to concatenate a string with a number
error1 = "family" + 3
print(error1)
x = 3
error2 = "family" + x
print(error2)
# + id="73vcQRIdxsRd" colab_type="code" colab={}
#######################################################
################### Challenge 3 #######################
#######################################################
# Come up with possible solutions for the error below
# Trying to print a string with mismatching string quotes
error2 = "I love my family'
print(error2)
# + [markdown] id="x9wNnIlwJkMu" colab_type="text"
# **String Formatting**
# + [markdown] id="gEadrR9FJkMy" colab_type="text"
# <details>
# <summary><b>Pro Tip</b> (Click to Expand)</summary>
# You can find out the data type by using Python's built in function, type(). __type()__ tells you what an object’s type is, for example a string (str) or integer (int).
# </details>
# + id="rSqe5JINOKws" colab_type="code" colab={}
pi = 3.14
# + id="OWyUOemjJkMz" colab_type="code" colab={}
p = "Python"
q = 22
print("p: ", type(p))
print("q: ", type(q))
# + [markdown] id="MJbHBf3RJkM2" colab_type="text"
# #### Casting
# + id="GLxG-WzgJkM4" colab_type="code" colab={}
x = str(9)
print(x)
# + id="Q6MciQgyO_8k" colab_type="code" colab={}
x = '5'
y = int(x)
print(y)
print(type(y))
# + [markdown] id="pexGoy4pJkM8" colab_type="text"
# **Note:**
# 1. Python Variables can't start with a number.
# Python Variables must start with a letter or the underscore character.
# 2. Variable names can't contain special characters such as '!', '@', '#', '$', '%'.
# 3. Your Python variables shouldn't be named 'and', 'if', 'while', 'True', 'False' because Python uses these names for program control structure.
# 4. Don't give your variables the same name as our data types. For example: Don't name your variable int, dict, list, str, etc.
# Variable names are case sensitive. For example: King, king and KING are different variable names. Please beware.
# + [markdown] id="D3c9gQ0sJkNB" colab_type="text"
# <a id='python-operations'></a>
# ## Python Operations
# + [markdown] id="X0QfDR2IJkNC" colab_type="text"
# **Arithmetic Operators**
# 1. Addition +
# 2. Subtraction -
# 3. Multiplication *
# 4. Division /
# 5. Floor division //
# 6. Modulus (remainder) %
# 7. Exponentiation **
# 8. Matrix multiplication @
# + id="R6MRAYugJkND" colab_type="code" colab={}
s = 3 + 5
q = 5 - 3
w = 3 * 4
e = 13 / 2
f = 20 // 3
t = 21 % 3 # modulo used to check if a number is odd(not dividing to 2) or even
y = 2 ** 3
print(s)
print(q)
print(w)
print(e)
print(f)
print(t)
print(y)
# + id="bGX0e8WOTf8Q" colab_type="code" colab={}
round(20/3)
# + id="84ZpYXkeJkNH" colab_type="code" colab={}
#######################################################
################### Challenge 4 #######################
#######################################################
# Rewrite the code so that it accepts user input height and weight and uses that to calculate the BMI.
height = 1.74
weight = 80
bmi = float(weight) / (height) ** 2
print("BMI is: " + str(bmi))
# + [markdown] id="-B-eZ-hkJkNK" colab_type="text"
# **In-Place Operations**
# + id="zXaqkDEqJkNL" colab_type="code" colab={}
# *=
x = 10
x *= 2 # this is the same with x = x * 2
print(x) # x = 20
# # +=
x += 4 # this is the same with : x = x + 4
print(x) # x = 24
# -=
x -= 1 # this is the same with : x = x -1
print(x) # x = 23
# /=
x /= 5 # this is the same with : x = x / 5
print(x) # x = 4.6
# + [markdown] id="jxzf5KyBJkNR" colab_type="text"
# **Comparison Operators**
# 1. Equal ==
# 2. Not equal !=
# 3. Greater than >
# 4. Less than <
# 5. Greater than or equal to >=
# 6. Less than or equal to <=
# + id="BYN-o346JkNS" colab_type="code" colab={}
# a > b
# a < b
# a >= b
# a <= b
a = 10
b = 4.5
print(type(a) == type(b))
print(type(a))
print(type(b))
print(type(a) == int)
# + [markdown] id="Ihmw-Ss0JkNW" colab_type="text"
# **Logical Operators**
# 1. and
# 2. or
# 3. not
# + id="oc18t9C3JkNX" colab_type="code" colab={}
a = 5
b = 6
print((a == 5) and (b == 6))
print(4 <= a <= 6)
x = True
print(not(x))
# + [markdown] id="P5YiOg1mJkNb" colab_type="text"
# Python has other operators such as the **Bitwise**, **Membership** and **Identity** Operators.
# + [markdown] id="QSQXKzvGJkNc" colab_type="text"
# **TODO:**
# 1. Calculate your daily expenses, by taking your monthly expenses and diving it by 30. Given that your monthly expenses is €800.
# 2. What is the value of a in "a = -1 ** 2"?
# 3. What is the value of b in "b = 99 >= 3**4 and 70 == 7 * 10 or 12 != 3 * 4"?
# 4. Accept a number from a user, print whether it is divisible by ten and greater than hundred.
# + id="pPDyc8AOySEu" colab_type="code" colab={}
#######################################################
################### Challenge 5 #######################
#######################################################
# 1.Calculate your daily expenses, by taking your monthly expenses and diving it by 30. Given that your monthly expenses is €800.
# 2. What is the value of a in "a = -1 ** 2"?
# 3. What is the value of b in "b = 99 >= 3*4 and 70 == 7 * 10 or 12 != 3 * 4"?
# 4. Accept a number from a user, print whether it is divisible by ten and greater than hundred.
# + [markdown] id="ojDpNbmAJkNc" colab_type="text"
# <a id='strings-and-their-methods'></a>
# ## Strings & their Methods
# + [markdown] id="MDDmxw58JkNd" colab_type="text"
# A method is a function that is specific to a certain "object" in Python. Python has a set of built-in methods that you can use on strings.
# + id="pdgCDgrAJkNe" colab_type="code" colab={}
var = "let's code amsterdam"
upper_var = var.upper()
lower_var = var.lower()
capital_var = var.capitalize()
title_var = var.title()
swapcase_var = var.swapcase()
replace_var = var.replace('l', 'r')
# + [markdown] id="Lz3eO_pv04nO" colab_type="text"
# Length of a words/ sentence or a paragraph can be calculated by simply calling the built-in function len() -> which in fact gives the number of characters.
# + id="2lsT4KmG1JUI" colab_type="code" colab={}
# len(str) is the number of chars in a String
# len('I am excited to code in Python. This is awesome!')
# len('this has a lot of spaces')
# len('this ')
len('a ')
# + id="PxNM8CHv1msi" colab_type="code" colab={}
# str[i:j] extracts the substring starting at index i and running up to but not including index j.
var = 'I am amazing'
x = 'cuphjgusdhgusdicjdsolncjlsfnvsgfyvsjkxnck;amdolfndhnpdbcvnjbhb'
x[-4]
# + [markdown] id="AMGxeb-OJkNh" colab_type="text"
# Read More: [String Methods](https://docs.python.org/3/library/stdtypes.html#string-methods)
# + [markdown] id="ADxAjKsNJkNi" colab_type="text"
# **TODO:**
# 1. Take a user input, convert the first character of each word to upper case, split the string every whitespace and join all the strings with a plus character (+)
# + id="cLbLsLk9yjv0" colab_type="code" colab={}
#######################################################
################### Challenge 6 #######################
#######################################################
# Take a user input, convert the first character of each word to upper case,
# split the string every whitespace and join all the strings with a plus character (+)
# + [markdown] id="hGwG7xokJkNj" colab_type="text"
# <a id='collections'></a>
# ## Lists, Tuples, Sets, & Dictionaries
# + [markdown] id="LMVG7a69JkNk" colab_type="text"
# #### Lists
# + [markdown] id="ktoVJErGJkNk" colab_type="text"
# Lists are one of the most powerful data types in Python. Lists are used for storing similar items, and in cases where items need to be added or removed.
# An empty list can be created in 2 ways:
# 1. By calling the list method list()
# 2. By writing 2 empty brackets []
# + id="drRwJS9IJkNm" colab_type="code" colab={}
list_1 = [1,2,3]
list_2 = list(list_1)
print(list_2)
# + id="NfNjX0vhJkNr" colab_type="code" colab={}
# Searching a list
my_hobby_list = ["cooking", "travelling", "teaching"]
print("cooking" in my_hobby_list)
# appending to list (add to the end)
my_hobby_list.append("dancing")
print(my_hobby_list)
# inserting an item to a particular position in a list .insert(position, item)
my_hobby_list.insert(2, "sleeping")
print(my_hobby_list)
# accessing items by index
print(my_hobby_list[0])
# lists are mutable
my_hobby_list[0] = "eating"
print(my_hobby_list)
# sorting a list
my_hobby_list.sort() # from A to Z
print(my_hobby_list)
# reversing a list
my_hobby_list.reverse() # from Z to A
print(my_hobby_list)
# arranging a list in descending order
my_hobby_list.sort(reverse = True)
print(my_hobby_list)
# add items from another list (extend)
my_new_hobbies = ["Knitting", "Singing"]
my_hobby_list.extend(my_new_hobbies)
print(my_hobby_list)
# remove an item
my_hobby_list.remove("eating")
print(my_hobby_list)
# remove an item from the last position
my_hobby_list.pop()
print(my_hobby_list)
# remove an item from a particular index
my_hobby_list.pop(2)
print(my_hobby_list)
# + [markdown] id="akcx5kPQJkNu" colab_type="text"
# **Common List Errors**
# + id="jnGD0DRYJkN3" colab_type="code" colab={}
first_names = ["Jane", "Alex" "Sandra"]
print(first_names)
# + [markdown] id="NDjiGjpEJkN8" colab_type="text"
# <details>
# <summary><b>Pro Tip</b> (Click to Expand)</summary>
# To find out the built in methods for strings, lists, tuples, sets and dictionaries by typing dir(). dir() returns a list of valid attributes for an object, so you can quickly see what variables an object has or what functions you can call on it. Ignore the methods that start with underscores.
#
# help() brings up helpful documentation on any object. You can also type help() on its own to bring an interactive help console.
# </details>
# + id="M87PqReQyw0T" colab_type="code" colab={}
#######################################################
################### Challenge 7 #######################
#######################################################
# Print out the built in functions for lists.
# + [markdown] id="vO9NvpALJkN9" colab_type="text"
# #### Tuples
# + [markdown] id="2Xn7bdSPJkN-" colab_type="text"
# **Tuple** is a list that is immutable. It is used for storing a snapshot of related items when we don’t plan on modifying, adding, or removing data. Tuples can be used in other types of containers like sets and dictionaries
# Remember that because Tuples are immutable,
# 1. we can’t add or remove from tuples and
# 2. we can't sort tuples
# + id="qs6j8XQjJkN_" colab_type="code" colab={}
# Create a new tuple
my_tuple = ()
my_other_tuple = tuple()
student_tuple = ("Victoria", 24, "Computer Science major", 4.5)
# accessing by index
print(student_tuple[2])
# Tuple Unpacking
student_tuple = ("Victoria", 24, "Computer Science major", 4.5)
name, age, dept, grade = student_tuple
print(name, age, dept, grade)
# + [markdown] id="I8anqSjTfkNr" colab_type="text"
# **Common Tuple Errors**
# + id="mekxyWQYfELB" colab_type="code" colab={}
# reassigning tuples will result in an error
student_tuple[2] = "Maths major"
print(student_tuple)
# + id="CFeXeIFLy6TR" colab_type="code" colab={}
#######################################################
################### Challenge 8 #######################
#######################################################
# Print out the built in functions for tuples.
# + [markdown] id="u_9n8hFZJkOG" colab_type="text"
# #### Sets
# + [markdown] id="EvbCO4gNJkOI" colab_type="text"
# Sets are a datatype that allows you to store other immutable types in an unordered way. Sets ensure that there are no duplicates and all items are unique.
# + id="_OdQhzWMJkOJ" colab_type="code" colab={}
# Create a new set
my_set = set()
farming_set = {"tractor", "plants", "water", "sunlight", "plants", "water", "cutlass"}
print(farming_set) #notice that all duplicates are gone
# Also notice that sets don't have an order
# add to a set
farming_set.add("harvesters")
print(farming_set)
# remove from a set
farming_set.discard("cutlass")
print(farming_set)
# update a set with another set
other_farming_set = {"sprayer", "mower"}
farming_set.update(other_farming_set)
print(farming_set)
# + [markdown] id="-UrxKd5UJkOQ" colab_type="text"
# **Common Set Errors**
# + id="E3bXq74VJkOR" colab_type="code" colab={}
# 1. Using mutable data types will result in an error
friends = {"max", "john", "doe", ["ana", "loes"]}
print(friends)
# + id="BJess55gJkOU" colab_type="code" colab={}
# Trying to access set items by index will result in an error
print(farming_set[2])
# + id="08obTTPSJkOY" colab_type="code" colab={}
# updating a set with a string will give you results you might not be expecting.
farming_set.update("farm robots")
print(farming_set)
# + [markdown] id="2jzKzqOhJkOe" colab_type="text"
# #### Dictionaries
# + [markdown] id="pC0qU3KaJkOf" colab_type="text"
# Dictionary is used for storing data in key, value pairs. Keys used must be immutable data types.
# + id="z9PihbhHJkOg" colab_type="code" colab={}
# Create a new empty dictionary
my_dict = {}
my_other_dict = dict()
my_fitness_dict = {1: "football", 2: "cricket", 3: "table tennis", 4: "volleyball"}
print(my_fitness_dict)
# search for a key in the dictionary
print(1 in my_fitness_dict)
# get the items in a dictionary
print(my_fitness_dict.items())
# get the keys in a dictionary
print(my_fitness_dict.keys())
# get the values in a dictionary
print(my_fitness_dict.values())
# get the length of a dictionary
print(len(my_fitness_dict))
# add to a dictionary using the square notation
my_fitness_dict[5] = "hockey"
print(my_fitness_dict)
# updating a dictionary with another dictionary
new_fitness_dict = {6: "basketball", 7: "baseball"}
my_fitness_dict.update(new_fitness_dict)
print(my_fitness_dict)
# + [markdown] id="4QNTyg0yJkOk" colab_type="text"
# **Common Dictionary Errors**
# + id="5hXqmeMBJkOl" colab_type="code" colab={}
# trying to access an item by index
students = {"Jane": "History", "Ada": "Biology", "William":"Political Science"}
print(students[3])
# + id="GTV_duLbJkOr" colab_type="code" colab={}
# Using a mutable data type as a key
new_students = {"Emma": "Chemistry", ["Ava", "Logan"]: "Physics"}
print(new_students)
# + id="-Kf0VjrnzGQ1" colab_type="code" colab={}
#######################################################
################### Challenge 8 #######################
#######################################################
# Print out the built in functions for dictionaries.
# + [markdown] id="Hq4C1suGslvp" colab_type="text"
# <a id="conditions-and-loops"></a>
# ## Conditions And Loops
# + [markdown] id="lWlNAEqjKCXh" colab_type="text"
# ### Conditions
# + [markdown] id="zL43hgVrtGr-" colab_type="text"
# Conditions help us control the logical flow of our program.
# + [markdown] id="vnkf-9Bbtmhk" colab_type="text"
# "If the weather in Amsterdam is nice, I will go to the park. If not, I will see some movies on Netflix"
# + id="IBheMG1fs_2i" colab_type="code" colab={}
# Syntax
if <expression>:
<statement>
else:
<another statement>
# Question 1:
# - what is <expr> in our example ?
# - what is <statement> in our example ?
# + [markdown] id="aCME2xdruy0_" colab_type="text"
# **if-else branch**
#
# 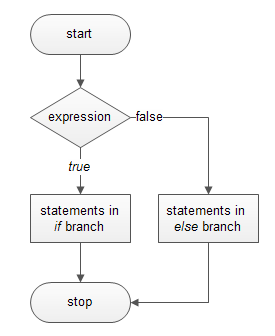
#
#
#
#
#
#
# **If-else multiple branches**
#
# 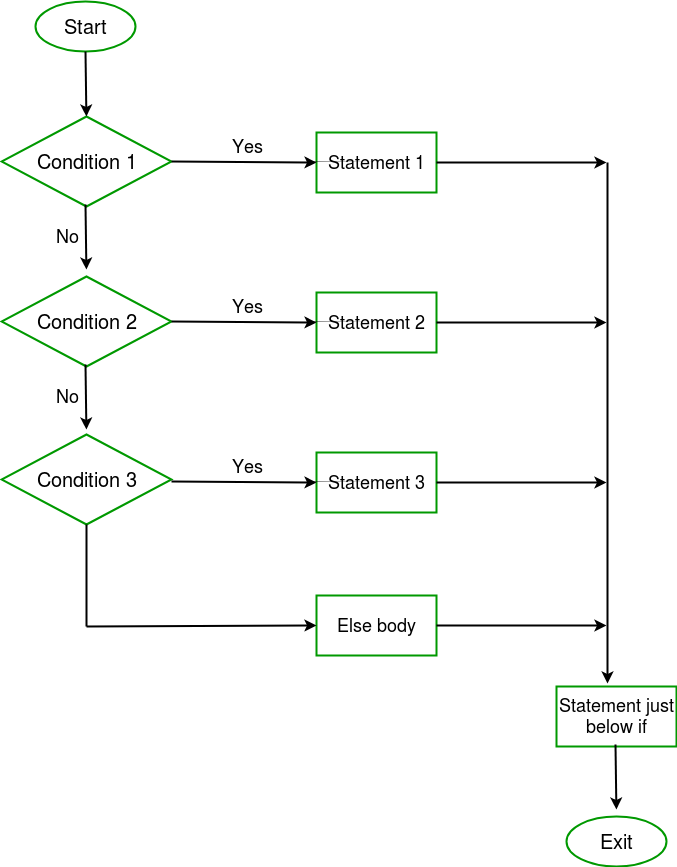
# + id="vd0k3tRZuthb" colab_type="code" colab={}
# Ana is 31 years old, Kate is 23 years old and David is 24 years old.
# Write a program which checks if Ana is the oldest or not the oldest of the group.
# Print the corresponding message.
ana_age = 31
kate_age = 23
david_age = 24
if (ana_age > david_age) and (ana_age > kate_age):
print("Ana is the oldest.")
else:
print("Ana is not the oldest.")
# + [markdown] id="y2T-q2ysyE_T" colab_type="text"
# ### Loops
# + [markdown] id="vY2VQUCoyKx-" colab_type="text"
# Looping in Python is simpler, cleaner process compared to other languages because the Python language prides itself on readability.
# + id="S2sdpN7zyeh6" colab_type="code" colab={}
# Syntax
for single_item in items:
body of the loop
# + [markdown] id="y4au6810yKdY" colab_type="text"
#
# * For loop
# * While loop
#
#
# The **while loop** constantly checks if a condition is satisfied, and continues to run as long as the condition remains satisfied. The **for loop** iterates over a list or a sequence.
# + id="UsPPTtX6zC6M" colab_type="code" colab={}
# for loop
seq = [1,2,3,4,5]
for i in seq:
print(i)
# + [markdown] id="kYijx67vzcQq" colab_type="text"
# **While loops** are a special type of loop in Python. Instead of running just once when a condition is met, like an if statement, they run forever until a condition is no longer met.
# + id="Ks0LjSQDziUv" colab_type="code" colab={}
counter = 0
max = 4
while counter < max:
print(f"The count is: {counter}")
counter = counter + 1
# + [markdown] id="StZlkbL7z4vl" colab_type="text"
# **Break and Continue** allow you to control the flow of your loops. They’re a concept that beginners to Python tend to misunderstand, so pay careful attention.
# + [markdown] id="Up_OLHMOz8s4" colab_type="text"
# #### Break
#
# The **break statement** will completely break out of the current loop, meaning it won’t run any more of the statements contained inside of it.
# + id="CmRy18lu0I22" colab_type="code" colab={}
names = []
for name in names:
print(f"Hello, {name}")
if name == "":
break
# + [markdown] id="jeAxxFEY0fwR" colab_type="text"
# #### Continue
#
# **continue** works a little differently. Instead, it goes back to the start of the loop, skipping over any other statements contained within the loop.
# + id="YO3HbeAX0sP2" colab_type="code" colab={}
names = []
for name in names:
if name != "":
continue
print(f"Hello, {name}")
# + [markdown] id="pDcs0qll1R36" colab_type="text"
# 
# + id="wKLUfDbB10_P" colab_type="code" colab={}
# print the names with a length of 4. After "Nina", break out of the loop
names = ["Jimmy", "Rose", "Max", "Nina", "Phillip"]
for name in names:
if len(name) != 4:
continue
print(f"Hello, {name}")
if name == "Nina":
break
print("Done!")
# + [markdown] id="NxyEyCmf1u_e" colab_type="text"
# Loop Control in While loops
# + id="XtbFW50-2bYb" colab_type="code" colab={}
count = 0
while True:
count += 1
print(count)
if count == 5:
print("Count reached")
break
# + [markdown] id="71ECHDWO2z57" colab_type="text"
# <a id="functions"></a>
# ## Functions
# + [markdown] id="j_3HVFCl27ah" colab_type="text"
# **Functions** simply take an input, do something with it, and then return the output.
#
# The purpose of functions in Python are to create reusable code. If we find ourselves copying and pasting the same code multiple times, that’s a good sign that a function might help!
#
#
# In Python we have 3 types of functions:
# * **Built-in Functions**: len(), min() and print() functions.
# * **User Defined Functions (UDF's) functions**: You can create these yourself.
# * **Anonymous or Lambda functions**: These are not declared with the standard def keyword
#
#
# **Watch**: [Function Video](https://www.youtube.com/watch?v=9Os0o3wzS_I)
#
#
# ---
#
#
# **Syntax**:
#
# def function_name(parameter):
#
# body_of_code
#
# The def keyword tells Python we’re about to start a function definition
# The
#
#
# **Remember that:**
# * a parameter is a placeholder for the actual value.
# * an argument is the actual value that is passed in
#
# [Python Docs](https://docs.python.org/3/faq/programming.html#what-is-the-difference-between-arguments-and-parameters)
#
# ---
#
#
#
# **==>** In the diagram below, which is the function name, the parameter and the argument
#
#
# 
#
#
#
#
# + id="lL1yTX9VBgNr" colab_type="code" colab={}
# A Basic Function that accepts no arguments
def hello_world():
print("Hello, World!")
# A Function that accepts an argument.
def euro_to_pounds(amount):
euros = amount*0.90
print(f'{amount} euros is {euros} pounds!')
return euros
# A Function that accepts two arguments, and returns the value of
# those numbers added together.
def add_numbers(x, y):
return x + y
# A function that has a default parameter
def my_function(country = "Norway"):
print("I am from " + country)
# + [markdown] id="NiKnJOJsEPUC" colab_type="text"
# ==> [Differences between 'print()' and 'return()'](https://stackoverflow.com/questions/7664779/what-is-the-formal-difference-between-print-and-return)
# + [markdown] id="i0z7JjOihYff" colab_type="text"
# <a id="lets-code-together"></a>
# ## Let's Code Together
# + [markdown] id="6FaIUwoZiGha" colab_type="text"
# We will code the game of Hangman together! The aim of the game is to guess a word or phrase by guessing letters. Each time you guess a letter that isn't part of the word, you lose an attempt. You have 7 attempts before the game is over.
# + id="gUJxGXZFNHPH" colab_type="code" colab={}
# We'll need this for what we are going to do next
from IPython.display import clear_output
import getpass
# + id="eSNcPOCQhzKp" colab_type="code" colab={}
# This function displays the blanks and letters
def display_word(gl, w, a=10):
# First clear the previous display
clear_output()
for character in w:
if character in gl:
#display the letters already guessed
print(character, end = '')
elif character == ' ':
#print spaces because we don't guess those
print(character, end = '')
else:
#otherwise print a dash
print('_', end = '')
print("\nAttempts left: " + str(a))
# Let's make a list to store the letters that have already been guessed
# We set the number of attempts to guess
# Now player 1 inputs a word or phrase
# Convert it to lower case to reduce complexity
# We make a set out of this word to keep track of all the letters to guess
# We also remove spaces because we don't guess those
# Start a loop where you keep track of attempts
# Player 2 guesses a letter
# Check if this letter has not been guessed before, in which case add it to the list
# Check if the letter is in the actual word or phrase. If not, player 2 loses an attempt.
# Otherwise remove this letter from the list of unique letters
# Use the function written above to display the guessed word
# Check for a win
# + [markdown] id="mzCGo1vPxuAD" colab_type="text"
# ## More challenges
# + id="esDP0K2qx1rK" colab_type="code" colab={}
#######################################################
################### Challenge 9 #######################
#######################################################
# Given a string of even length, return the first half. For example, if the string is given as "WooHoo" this will yield "Woo".
# + id="E0NaFvDfx4fG" colab_type="code" colab={}
#######################################################
################### Challenge 10#######################
#######################################################
# Write a sorting algorithm for a numerical dataset. Create the dataset with the use of list datatype.
# + id="ntym696AyAF9" colab_type="code" colab={}
########################################################
################### Challenge 11 #######################
########################################################
# Which one of these is floor division?
# a) /
# b) //
# c) %
# d) None of the mentioned
# Solution: c) %
# + id="MOmsQJU5yDA9" colab_type="code" colab={}
# Suppose list1 is [2, 33, 222, 14, 25], What is list1[-1]? How many elements does the list have?
# + id="XSXl09Q6rSqC" colab_type="code" colab={}
########################################################
################### Challenge 13 #######################
########################################################
# A taxi driver is calculating their profit over two weeks by adding up the fares they charge and subtracting the cost of gas.
# The price of gas changes over time - it was $3.52/gallon the first week and $3.57/gallon this second week.
# Their car gets 20 miles per gallon.
# For the first week the driver had a total of 23 passengers with average $29 fare each, and drove a total of 160 miles.
# For the second week they had 17 passengers with average 30 fare each, and drove a total of 220 miles.
# Assume that for both weeks they purchase all the gas needed during that week (i.e. they refuel every week to maintain a constant level of gas in the tank).
# Based on the above, answer the following questions:
# 1. What is their total profit over both weeks?
# 2. During which week was their average (mean) profit per passenger higher?
# + id="KyTDmel2wcpR" colab_type="code" colab={}
########################################################
################### Challenge 14 #######################
########################################################
# Ask the user for a number. Depending on whether the number is even or odd, print out an appropriate message to the user.
# Extras:
# 1. If the number is a multiple of 4, print out a different message.
# 2. Ask the user for two numbers: one number to check (call it num) and one number to divide by (check).
# If check divides evenly into num, tell that to the user. If not, print a different appropriate message.
# + id="7ba0V4PRGB6B" colab_type="code" colab={}
########################################################
################### Challenge 15 #######################
########################################################
# Complete filter_names function that takes a list of names and returns a filtered list of names using the following conditions:
# names that start with IGNORE_CHAR are ignored,
# names that have one or more digits are ignored,
# if a name starts with QUIT_CHAR it immediately exits the loop, so no more names are added/generated at this point (neither the QUIT_CHAR name),
# return up till MAX_NAMES names max.
IGNORE_CHAR = 'b'
QUIT_CHAR = 'q'
MAX_NAMES = 5
def filter_names(names):
filter_names(['dan', 'chidinma', 'joop', 'rihanna', 'queen', 'amber', 'bob', 'c1ndy', 'sara', 'molly', 'henry', 't2im', '1quinton', 'a3na', '4'])
# + [markdown] colab_type="text" id="nKbEFoQiYH7B"
# <a id='further-reading'></a>
# ## For Further Reading:
# + [markdown] id="b8My3tchJkOx" colab_type="text"
# - [Python Official Documentation](https://docs.python.org/3/)
# - [Jupyter Tips & Tricks 1](https://medium.com/ibm-data-science-experience/markdown-for-jupyter-notebooks-cheatsheet-386c05aeebed)
# - [Jupyter Tips & Tricks 2](https://www.dataquest.io/blog/jupyter-notebook-tips-tricks-shortcuts/)
# - [Python Wiki](https://wiki.python.org/moin/FrontPage)
# - [W3Schools](https://www.w3schools.com/python/default.asp)
# + [markdown] id="tPEqCONVI9x-" colab_type="text"
# **You Are Amazing!**
#
#
# @ChidinmaKO
# + colab_type="code" id="u2b3TAa7fx1o" colab={}
| Intro to Python/Lets_Code_Ams.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PythonData
# language: python
# name: pythondata
# ---
# !pip install psycopg2 sqlalchemy
# +
#Import pandas
import pandas as pd
#Import SQLAlchemy
from sqlalchemy import create_engine, inspect
#Import matplotlib
import matplotlib.pyplot as plt
#Import Numpy
import numpy as np
from config import pkey
# -
#Create Engine
url = "postgresql://postgres:" +pkey+"@localhost:5432/EmployeeSQL"
engine = create_engine(url)
connection = engine.connect()
#Query all record from database
salaries_data = pd.read_sql("SELECT * FROM salaries", connection)
employees_data = pd.read_sql("SELECT * FROM employees", connection)
title_data = pd.read_sql("SELECT * FROM titles", connection)
title_data.head()
# +
# Create a histogram to visualize the most common salary ranges for employees.
plt.hist(salaries_data["salary"], label="Salary")
plt.title("Common Salary Ranges")
plt.ylabel("Number of Employees")
plt.xlabel("Salary")
# +
# Create a bar chart of average salary by title.
join = pd.read_sql("SELECT * FROM salaries JOIN employees ON (salaries.emp_number=employees.emp_number) JOIN titles ON (titles.title_id=employees.emp_title_id)", connection)
group_join = join.groupby(["title"]).mean()["salary"]
title = group_join.index
avg_salary = group_join.values
x_axis = np.arange(len(title))
tick_locations = [value for value in x_axis]
plt.xticks(tick_locations, title, rotation = 90)
plt.bar(x_axis, avg_salary, color="b", align="center")
plt.title("Average Salary by Title")
plt.ylabel("Average Salary")
plt.xlabel("Title")
plt.show
# -
inspector=inspect(engine)
inspector.get_columns('salaries')
| Visualizations_Bonus.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# import the project1-prepareData notebook:
# !pip install ipynb
from ipynb.fs.full.data_analysis import *
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# +
berlinDf_final_xgboost = berlinDf_select.copy()
# berlinDf_final_linear.drop(['newlyConst', 'balcony','hasKitchen', 'lift', 'garden'], axis='columns', inplace=True)
df_full_train, df_train, df_val, df_test, y_full_train, y_train, y_val, y_test = split_dataFrame(berlinDf_final_xgboost)
berlinDf_final_linear
# berlinDf_final_linear.columns
berlinDf_final_xgboost.to_csv('immo_data_final.csv')
# +
#Model training with XGboost
# !pip install xgboost
import xgboost as xgb
from sklearn.feature_extraction import DictVectorizer
from sklearn.tree import DecisionTreeClassifier
def train(dataFrame, y, max_depth, eta):
# Hot Encoding
dicts = dataFrame.to_dict(orient="records")
dv = DictVectorizer(sparse=False)
X = dv.fit_transform(dicts)
features = dv.get_feature_names()
X = dv.fit_transform(dicts)
dtrain = xgb.DMatrix(X, label=y, feature_names=features)
# train
xgb_params = {
'eta': eta,
'max_depth': max_depth,
'min_child_weight': 1,
'objective': 'reg:squarederror',
'nthread': 8,
'seed':1,
'verbosity':0
}
model = xgb.train(xgb_params, dtrain, num_boost_round=10)
return dv, model
# +
# Predict
def predict(dataFrame, dv, model):
dicts = dataFrame.to_dict(orient="records")
X = dv.transform(dicts)
features = dv.get_feature_names()
dval = xgb.DMatrix(X, feature_names=features)
y_pred = model.predict(dval)
return y_pred, X
# -
model, dv = train(df_full_train, y_full_train, 20, 0.6)
| capstone-project/.ipynb_checkpoints/build_model-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # What's a neural network
# This video explains very well what is a neural network,
# and also basically how it works via the hand-written digits recognition example.
from IPython.display import YouTubeVideo
YouTubeVideo('aircAruvnKk')
# # The neural network we will build in this post
#
# <img src="./hand_written_digits_recognition_nn_model.png">
#
# We will again use the hand-written digits data to build a hand-written recognition neural network model in this post.
#
# As you can see from above NN model graph, our NN has 3 layers:
#
# - An input layer: recall that each of the input hand-written digit holds a 20 by 20 pixels, which gives us 400 input layer units plus 1 always `+1` bias unit;
#
# - A hidden layer: which has 25 units (not counting the extra bias unit which always outputs `+1`);
#
# - An output layer: which has 10 output units (corresponding to the 10 digit classes);
#
# That is:
#
# $$
# \begin{cases}
# a^{(1)}.shape &= (401, 1) \\
# \Theta^{(1)}.shape &= (25, 401) \\
# z^{(2)} = \Theta^{(1)} a^{(1)} = (25,401)@(401,1) &= (25, 1) \\
# \Theta^{(2)}.shape &= (10, 26) \\
# z^{(3)} = \Theta^{(2)} a^{(2)} = (10, 26)@(26, 1) &= (10, 1)
# \end{cases}
# $$
#
# ### Question: why the hidden layer has 25 units?
# # Hand-written digits recognition with neural network
# +
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# Sets the backend of matplotlib to the 'inline' backend.
#
# With this backend, the output of plotting commands is displayed inline within frontends like the Jupyter notebook,
# directly below the code cell that produced it.
# The resulting plots will then also be stored in the notebook document.
#
# More details: https://stackoverflow.com/questions/43027980/purpose-of-matplotlib-inline
# %matplotlib inline
from scipy.io import loadmat
data = loadmat(os.getcwd() + '/hand_written_digits.mat')
data
# +
X = data['X']
y = data['y']
X.shape, y.shape
# -
# ### Use [one-hot encoding](https://en.wikipedia.org/wiki/One-hot) to encode the classes labels
#
# [One-hot encoding](https://en.wikipedia.org/wiki/One-hot) projects class label $K_i$ to a $K$-length vector, which its component at index $i$ is 1, and all others components are 0.
# +
from sklearn.preprocessing import OneHotEncoder
onehot_encoder = OneHotEncoder(sparse=False)
y_onehot = onehot_encoder.fit_transform(y)
y_onehot.shape
# -
y[0], y_onehot[0, :]
def sigmoid(x):
return 1 / (1 + np.exp(-x))
# ### `forward_propagate` just simulates the process that all inputs run through the neural network we defined, then returns the intermediate results and the final output.
def forward_propagate(X, theta1, theta2):
a1 = np.insert(X, 0, values=np.ones(X.shape[0]), axis=1)
z2 = a1 @ theta1.T
a2 = np.insert(sigmoid(z2), 0, values=np.ones(X.shape[0]), axis=1)
z3 = a2 @ theta2.T
h = sigmoid(z3)
return a1, z2, a2, z3, h
# ### Define `cost` function (WITHOUT regularization item) to evaluate the loss of the network
#
# $$
# J(\theta) = -\frac{1}{n} \sum\limits_{i=1}^n \sum\limits_{k=1}^K \Big[ \
# y_k^{(i)}log\big( h_\theta(x^{(i)})_k \big) + \
# (1 - y_k^{(i)}) log\big( 1 - h_\theta(x^{(i)})_k \big) \
# \Big]
# $$
def cost(num_of_hidden_layer_units, num_of_labels, X, y, alpha):
theta1 = (
np.random.random(
size=(num_of_hidden_layer_units, X.shape[1] + 1)
) - 0.5
) * 0.25
theta2 = (
np.random.random(
size=(num_of_labels, num_of_hidden_layer_units + 1)
) - 0.5
) * 0.25
a1, z2, a2, z3, h = forward_propagate(X, theta1, theta2)
J = 0.
for i in range(X.shape[0]):
part0 = np.multiply(y[i,:], np.log(h[i,:]))
part1 = np.multiply(1 - y[i,:], np.log(1 - h[i,:]))
J += np.sum(part0 + part1)
return -J/X.shape[0]
cost(25, 10, X, y_onehot, 1)
# ### Define `cost` function (WITH regularization item) to evaluate the loss of the network
#
# $$
# J(\theta) = -\frac{1}{n} \sum\limits_{i=1}^n \sum\limits_{k=1}^K \Big[ \
# y_k^{(i)}log\big( h_\theta(x^{(i)})_k \big) + \
# (1 - y_k^{(i)}) log\big( 1 - h_\theta(x^{(i)})_k \big) \
# \Big] + \
# \frac{\alpha}{2n} \Big[ \
# \sum\limits_{j=1}^{25} \sum\limits_{k=1}^{400} (\Theta_{j,k}^{(1)})^2 + \
# \sum\limits_{j=1}^{10} \sum\limits_{k=1}^{25} (\Theta_{j,k}^{(2)})^2 \
# \Big]
# $$
#
# As you can see, we don't regularize the bias unit.
def cost(num_of_hidden_layer_units, num_of_labels, X, y, alpha):
theta1 = (
np.random.random(
size=(num_of_hidden_layer_units, X.shape[1] + 1)
) - 0.5
) * 0.25
theta2 = (
np.random.random(
size=(num_of_labels, num_of_hidden_layer_units + 1)
) - 0.5
) * 0.25
a1, z2, a2, z3, h = forward_propagate(X, theta1, theta2)
J = 0.
for i in range(X.shape[0]):
part0 = np.multiply(y[i,:], np.log(h[i,:]))
part1 = np.multiply(1 - y[i,:], np.log(1 - h[i,:]))
J += np.sum(part0 + part1)
regularization_item = float(alpha) / (2 * X.shape[0]) * (
np.sum(
np.power(theta1[:,1:], 2)
) +
np.sum(
np.power(theta2[:,1:], 2)
)
)
return -J/X.shape[0] + regularization_item
cost(25, 10, X, y_onehot, 1)
# ### Computes the gradient of the sigmoid function
def sigmoid_gradient(x):
return np.multiply(sigmoid(x), (1 - sigmoid(x)))
# ### Implement backpropagation algorithm (WITH cost regularization item and gradient regularization item)
#
# - Backpropagation computes the parameter updates that will reduce the error of the network on the training data.
#
#
# - Combine the [Chain rule](https://en.wikipedia.org/wiki/Chain_rule) and the below two graphs should be good enough to explain what is and what does backpropagation algorithm do.
#
# Lets take calculating the derivative of $e=(a+b)*(b+1)$ as example, and lets introduce in intermediate variables $c=a+b$ and $d=b+1$:
#
# <img src="./bp_0.png">
#
# For calculating the $d_e|_{a=2,b=1}$, with the [Chain rule](https://en.wikipedia.org/wiki/Chain_rule) we know:
# $$
# \begin{align*}
# d_e|_{a=2,b=1} &= \frac{\partial e}{\partial a} + \frac{\partial e}{\partial b} \\
# &= \frac{\partial e}{\partial c} \cdot \frac{\partial c}{\partial a} + \
# \frac{\partial e}{\partial c} \cdot \frac{\partial c}{\partial b} + \
# \frac{\partial e}{\partial d} \cdot \frac{\partial d}{\partial b}
# \end{align*}
# $$
#
# If we visualize the above chain rules in a tree, we get:
#
# <img src="./bp_1.png">
#
# We found that actually:
#
# 1. The value of $\frac{\partial e}{\partial a}$ is the product of all the derivatives on the path from node $a$ to node $e$;
#
# 2. The value of $\frac{\partial e}{\partial b}$ is the sum of the product of all the derivatives on the two different paths respectively from node $b$ to node $e$;
#
#
# That means: to upper node $p$ and lower node $q$, for calculating $\frac{\partial p}{\partial q}$ we need to find out all the paths from node $q$ to node $p$, then to each path we calculate the product of all the derivatives on that path, and then sum all the products from all the different paths!
#
# But maybe you already noticed: we visited certain paths multiple times, for example: path 'a-c-e' and 'b-c-e' both visited path 'c-e', this duplicated traversal cost to a huge neural network is significant!
#
#
# - And here is also where the backpropagation algorithm comes in: just like indicated in its name (back), it looks up the paths from the root node to the leaf nodes, and traverse each path eactly once, how it achieves this:
#
# 1. It starts from root node with initial value `1`, and processes the others nodes by layer from top to bottom;
#
# 2. To each node (lets say $p$), calculate the derivative of $p$ to each of its direct children (lets say $q$), that is: $\frac{\partial p}{\partial q}$, then store the product of the value that accumulated on node $p$ (for root node it is our initial value `1`) and the just calculated $\frac{\partial p}{\partial q}$ on node $q$;
#
# 3. After finished one layer, sum all the stored values on each node respectively, and store as its accumulated value;
#
# 4. Repeat step '2' and '3' until finish all the nodes, the value lastly accumulated on the leaf node (lets say $q$) is the derivative of the root node (lets say $p$) to this leaf node, that is: $\frac{\partial p}{\partial q}$!
#
# More clearly, still with above example, demonstrate the process with below graph:
#
# <img src="bp_demo.svg">
#
#
# - The computations required for backpropagation are a superset of those required in the cost function, so what we will do actually is extending the cost function to perform the backpropagation as well, and then return both the cost and the gradients.
#
#
# - And since we will use our `backprop` function with the `scipy.optimize.minimize` function, which means the `backprop` will be called upon each epoch of the training, so we cannot do the `theta1` and `theta2` random generation like our above `cost` function, but pass in through the `params`.
def backprop(params, num_of_hidden_layer_units, num_of_labels, X, y, alpha):
theta1 = np.reshape(
params[:num_of_hidden_layer_units * (X.shape[1] + 1)],
(num_of_hidden_layer_units, X.shape[1] + 1)
)
theta2 = np.reshape(
params[num_of_hidden_layer_units * (X.shape[1] + 1):],
(num_of_labels, num_of_hidden_layer_units + 1)
)
a1, z2, a2, z3, h = forward_propagate(X, theta1, theta2)
# Initializations.
J = 0.
delta1 = np.zeros(theta1.shape) # (25, 401)
delta2 = np.zeros(theta2.shape) # (10, 26)
# Compute the cost.
for i in range(X.shape[0]):
part0 = np.multiply(y[i,:], np.log(h[i,:]))
part1 = np.multiply(1 - y[i,:], np.log(1 - h[i,:]))
J += np.sum(part0 + part1)
J = -J/X.shape[0]
# Add the regularization item to cost.
cost_regularization_item = float(alpha) / (2 * X.shape[0]) * (
np.sum(
np.power(theta1[:,1:], 2)
) +
np.sum(
np.power(theta2[:,1:], 2)
)
)
J += cost_regularization_item
# Perform backpropagation.
for t in range(X.shape[0]):
a1t = a1[[t],:] # (1, 401)
z2t = z2[[t],:] # (1, 25)
a2t = a2[[t],:] # (1, 26)
ht = h[[t],:] # (1, 10)
yt = y[[t],:] # (1, 10)
d3t = ht - yt # (1, 10)
z2t = np.insert(z2t, 0, values=np.ones(z2t.shape[0]), axis=1) # (1, 26)
d2t = np.multiply(d3t @ theta2, sigmoid_gradient(z2t)) # (1, 26)
delta1 += d2t[:,1:].T @ a1t
delta2 += d3t.T @ a2t
delta1 /= X.shape[0]
delta2 /= X.shape[0]
# Add the regularization item to the gradient.
# Note:
# We never regularize the bias item.
delta1[:,1:] += theta1[:,1:] * alpha / X.shape[0]
delta2[:,1:] += theta2[:,1:] * alpha / X.shape[0]
# Unravel the gradient matrices into a single array.
# Note:
# The first parameter of `np.concatenate` needs to be a tuple.
grad = np.concatenate(
(np.ravel(delta1), np.ravel(delta2))
)
return J, grad
# +
num_of_labels = 10
num_of_hidden_layer_units = 25
params = (
np.random.random(
size=25 * (X.shape[1] + 1) + num_of_labels * (num_of_hidden_layer_units + 1)
) - 0.5
) * 0.25
J, grad = backprop(params, num_of_hidden_layer_units, num_of_labels, X, y_onehot, 1)
J, grad.shape
# -
# ### Finally we are ready to train our network
#
# We put a bound on the number of iterations since the objective function is not likely to completely converge. As you can see the total cost has dropped below to around 0.3 though, so that's a good indicator that the algorithm is working.
# +
from scipy.optimize import minimize
# Minimize the objective function.
fmin = minimize(
fun=backprop, x0=params, args=(num_of_hidden_layer_units, num_of_labels, X, y_onehot, 1),
method='TNC', jac=True, options={'maxiter': 250}
)
fmin
# -
# ### Let's use the parameters it found and forward-propagate them through the network to get some predictions, and evaluate the overall accuracy of our network
# +
theta1 = np.reshape(
fmin.x[:num_of_hidden_layer_units * (X.shape[1] + 1)],
(num_of_hidden_layer_units, X.shape[1] + 1)
)
theta2 = np.reshape(
fmin.x[num_of_hidden_layer_units * (X.shape[1] + 1):],
(num_of_labels, num_of_hidden_layer_units + 1)
)
a1, z2, a2, z3, h = forward_propagate(X, theta1, theta2)
y_pred = np.array(np.argmax(h, axis=1) + 1)
correct = [1 if a == b else 0 for (a, b) in zip(y_pred, y)]
accuracy = (sum(map(int, correct)) / float(len(correct)))
print('Total accuracy: {0:.2f}%'.format(accuracy * 100))
# -
# # References
#
# - [Programming Exercise 4: Neural Networks Learning](https://github.com/jdwittenauer/ipython-notebooks/blob/master/exercises/ML/ex4.pdf)
#
# - [机器学习练习(五)—— 神经网络](https://blog.csdn.net/and_w/article/details/53612320)
#
# - [One-hot](https://en.wikipedia.org/wiki/One-hot)
#
# - [如何直观地解释 backpropagation 算法?](https://www.zhihu.com/question/27239198/answer/89853077)
#
# - [Calculus on Computational Graphs: Backpropagation](http://colah.github.io/posts/2015-08-Backprop/)
#
# - [How the backpropagation algorithm works](http://neuralnetworksanddeeplearning.com/chap2.html)
#
# - [Chain rule](https://en.wikipedia.org/wiki/Chain_rule)
| ml_basics/rdm010_neural_network/neural_network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dataset generation
# Two types of projects generated into json files
from datasets.random_dataset_generator import random_dataset_generator
# ## Classic project datasets
# More stakeholders and requirements to plan for a release, less interaction between them, generic requirement estimations.
# Total costs were extracted from ISBSG 2015, using {A,B} values for "UFP rating", "New development" for "Development type" and "IFPUG 4+" for "Count approach". This procedure is used to generate percentile 25,50,75 of total FPs of a classic project, in order to generate a realistic sample of classic estimation of requirements, done by selecting randomly, for a given number of pbis, a list of costs that sums up to the percentile value.
# +
reqs=[50,200,500]
stkh=[15,100]
percentile_total_costs=[156,312,705]
range_stakeholder_importances=[1,3,5]
range_stakeholder_pbis_priorities=[1,3,5]
percentage_dependencies= 0.45
counter=0
perc=0
for r in reqs:
for s in stkh:
counter+=1
name=f"c{counter}"
random_dataset_generator(num_pbis = r, num_stakeholders=s, percentage_dependencies=percentage_dependencies,
total_pbi_costs=percentile_total_costs[perc],
range_stakeholder_importances=range_stakeholder_importances,
range_stakeholder_pbis_priorities=range_stakeholder_pbis_priorities, name=name)
perc+=1
# -
# ## Agile project datasets
# Less stakeholders and requirements to plan for a release, much more interaction between stakeholder interests, Fibonacci estimations.
# +
reqs=[50,200]
stkh=[5,15]
range_pbi_costs =[1,1,2,3,5,8,13,21,34]
range_stakeholder_importances=[1, 2, 3, 4, 5]
range_stakeholder_pbis_priorities=[1, 2, 3, 4, 5]
percentage_dependencies= 0.45
counter=0
for r in reqs:
for s in stkh:
counter+=1
name=f"a{counter}"
random_dataset_generator(num_pbis = r, num_stakeholders=s, percentage_dependencies=percentage_dependencies,
range_pbi_costs =range_pbi_costs, range_stakeholder_importances=range_stakeholder_importances,
range_stakeholder_pbis_priorities=range_stakeholder_pbis_priorities, name=name)
| datasets/dataset_generation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # # Appendix E - Avoid hyperparameters
# +
import pandas as pd
import networkx as nx
import numpy as np
import matplotlib.pyplot as plt
from sklearn.cluster import SpectralClustering
from sklearn.metrics import pairwise_distances
from sklearn.cluster import KMeans
from numba import jit, prange
import plotly.graph_objects as go
import plotly.express as px
import plotly.io as pio
pio.renderers.default = 'iframe'
# +
data = pd.read_csv('Mall_Customers.csv', usecols=['Annual Income (k$)', 'Spending Score (1-100)', 'Age']).dropna()
data = normalized_df=(data - data.mean()) / data.std()
data.head()
# -
fig = px.scatter_3d(data, x='Annual Income (k$)', y='Spending Score (1-100)', z='Age', opacity=1, size=0.1 * np.ones(len(data)))
fig.show()
# +
def find_communities(adj):
communities = np.zeros(adj.shape[0], dtype=np.int32)
label = 0
pool = set()
for idx, val in enumerate(communities):
if val == 0:
# Change the community
label += 1
communities[idx] = label
# Neighbours are in the same community
neighbours = np.nonzero(adj[idx])[0]
# Neighbours of neighbours are in the same community
pool |= set(neighbours)
while pool:
neigh_idx = pool.pop()
neigh_val = communities[neigh_idx]
# Don't look at previously used data
if neigh_val == 0:
communities[neigh_idx] = label
neighbours = np.nonzero(adj[neigh_idx])[0]
pool |= set(neighbours)
return communities
@jit(nopython=True, nogil=True, parallel=True, fastmath=True)
def compute_modularity(adj, communities):
n_edges_doubled = np.sum(adj)
k_all = np.sum(adj, axis=1)
out = np.zeros((adj.shape[0], adj.shape[0]), np.float32)
for row_i_idx in prange(adj.shape[0]):
for row_j_idx in prange(row_i_idx+1):
# Compute it only for nodes of the same community
if communities[row_i_idx] == communities[row_j_idx]:
A_ij = adj[row_i_idx, row_j_idx]
P_ij = (k_all[row_i_idx] * k_all[row_j_idx]) / n_edges_doubled
local_mod = A_ij - P_ij
out[row_i_idx][row_j_idx] = local_mod
out[row_j_idx][row_i_idx] = local_mod
out_sum = np.sum(out) / n_edges_doubled
return out_sum
# -
distances = pairwise_distances(data)
modularities = []
space = np.linspace(0.01, 0.99, 100)
for threshold in space:
binarized = (distances < threshold).astype(np.int32)
communities = find_communities(binarized)
modularity = compute_modularity(binarized, communities)
modularities.append(modularity)
fig = go.Figure(data=go.Scatter(x=space, y=modularities, mode='lines+markers'))
fig.update_layout(xaxis_title='Threshold',
yaxis_title='Modularity')
# +
# Binarize network
binarized = (distances < 0.41).astype(int)
plt.imshow(binarized)
plt.show()
# Compute the laplacian
D = np.diag(np.sum(binarized, axis=0))
laplacian = D - binarized
# Compute eigenvalues and eigenvectors
evals, evects = np.linalg.eigh(laplacian)
# -
# Plot eigenvalues
plt.figure(figsize=(20,10))
plt.bar(np.arange(60), evals[0:60])
plt.xticks(np.arange(0, 60), labels=np.arange(1, 60+1))
plt.xlabel('Eigenvalues')
plt.ylabel('Intensity')
plt.savefig('../figures/mall-eigens.eps')
plt.show()
kmeans = KMeans(n_clusters=57)
labeled_data = data.copy()
labeled_data['labels'] = kmeans.fit(evects[:,0:57]).labels_
fig = px.scatter_3d(labeled_data, x='Annual Income (k$)', y='Spending Score (1-100)', z='Age',
opacity=1, color='labels', size=0.1 * np.ones(len(data)))
fig.show()
clustering = SpectralClustering(n_clusters=5, assign_labels='kmeans').fit(data)
labeled_data = data.copy()
labeled_data['labels'] = clustering.labels_
fig = px.scatter_3d(labeled_data, x='Annual Income (k$)', y='Spending Score (1-100)', z='Age',
opacity=1, color='labels', size=0.1 * np.ones(len(data)))
fig.show()
| appendices/section-5-rwe.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"is_executing": false, "name": "#%%\n"}
import torch
# + pycharm={"is_executing": false, "name": "#%%\n"}
x = torch.ones(2, 2, requires_grad=True)
print(x)
print(x.grad_fn)
# + pycharm={"is_executing": false, "name": "#%%\n"}
y = x + 2
print(y)
print(y.grad_fn)
# + pycharm={"is_executing": false, "name": "#%%\n"}
print(x.is_leaf, y.is_leaf)
# + pycharm={"is_executing": false, "name": "#%%\n"}
z = y * y * 3
out = z.mean()
print(z, out)
# + pycharm={"is_executing": false, "name": "#%%\n"}
out.backward()
# + pycharm={"is_executing": false, "name": "#%%\n"}
print(x.grad)
# + pycharm={"is_executing": false, "name": "#%%\n"}
out2 = x.sum()
out2.backward()
print(x.grad)
# + pycharm={"is_executing": false, "name": "#%%\n"}
out3 = x.sum()
x.grad.data.zero_()
out3.backward()
print(x.grad)
# + pycharm={"is_executing": false, "name": "#%%\n"}
out3.backward()
x.grad
# + pycharm={"is_executing": false, "name": "#%%\n"}
x = torch.tensor([1.0, 2.0, 3.0, 4.0], requires_grad=True)
y = 2 * x
z = y.view(2, 2)
print(z)
# + pycharm={"is_executing": false, "name": "#%%\n"}
v = torch.tensor([[1.0, 0.1], [0.01, 0.001]], dtype=torch.float)
z.backward(v)
print(x.grad)
# + pycharm={"is_executing": false, "name": "#%%\n"}
x = torch.tensor(1.0, requires_grad=True)
y1 = x ** 2
# with torch.no_grad():
y2 = x ** 3
y3 = y1 + y2
print(x.requires_grad)
print(y1, y1.requires_grad)
print(y2, y2.requires_grad)
print(y3, y3.requires_grad)
# + pycharm={"is_executing": false, "name": "#%%\n"}
y3.backward()
print(x.grad)
# + pycharm={"is_executing": false, "name": "#%%\n"}
x = torch.ones(1, requires_grad=True)
print(x.data)
print(x.data.requires_grad)
y = 2 * x
x.data *= 100 # 只改变了值,不会记录在计算图,所以不会影响梯度传播
y.backward()
print(x)
print(x.grad)
# + pycharm={"is_executing": false, "name": "#%%\n"}
x = torch.tensor([[1., 2., 3.], [4., 5., 6.]], requires_grad=True)
y = x + 1
z = 2 * y * y
J = torch.mean(z)
J.backward()
print(x.grad)
# + pycharm={"name": "#%%\n"}
| Three_Part_Moudule/pytorch/data_operator/auto_gradient-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Optimizing polynomials with strange ways
#
# We'll be minimizing single and multivariable quartic functions (without using the knowledge it is one) in this file to see if we can make this stuff actually work. This leads up to attempting to use these methods on a machine learning model in the other file (ScikitModels.ipynb).
#
# Author: <NAME>
import random
import numpy as np
from scipy.optimize import minimize # checking goodness of result compared to scipy
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_theme(style="darkgrid")
# +
# Function to generate functions to test on
def quartic():
# default ranges for variables too
a, b = random.random() * 4.9 + 0.1, random.random() * 10 - 5
c, d = random.random() * 40 - 20, random.random() * 500 - 250
return lambda x: a * x**4 + b * x**3 + c * x**2 + d * x
# Making an nd-quartic function to test higher dimensionalities.
# For the sake of generality, this should be used for the single variable case as well.
# n - how many inputs the function takes
# outputs the sum of n random quartic functions
def quartic_n(n):
fs = [quartic() for _ in range(n)]
return lambda answers: sum(map(lambda pair: pair[0](pair[1]), zip(fs, answers)))
# Example scipy optimization of a 4d quartic function:
# minimize(quartic_n(4), (0,0,0,0))
# -
# # Implementations
# ### Differential Evolution
# +
# The main function that returns the best set of inputs it finds.
# f - the function.
# n - input dimensionality
# k - population size
# scaling - the scaling parameter when creating a new input set
# loops - how many loops it will do before giving up finding a better solution
# Outputs the minimum value of the function that it found and the necessary input for it
def diff_evo(f, n, k = 80, scaling = 0.5, loops=25):
# Create initial input set
pop = create_population(n, k)
fitnesses = calculate_fitness(f, pop)
best, bestval = getbest(pop, fitnesses) # pair of best input and its value
loops_since_improvement = 0 # Keep it going until it's not working anymore.
while loops_since_improvement < loops:
loops_since_improvement += 1
# Create next population by mutating previous one
newpop = create_next_population(pop, scaling)
newfitnesses = calculate_fitness(f, newpop)
nextbest, nextbestval = getbest(newpop, newfitnesses)
# Keep track of what the best outcome is
if nextbestval < bestval:
best, bestval = nextbest, nextbestval
loops_since_improvement = 0
# Always choose the better one of the two choices to represent the 'next generation'
for i in range(k):
if newfitnesses[i] < fitnesses[i]: # if something must be changed
pop[i], fitnesses[i] = newpop[i], newfitnesses[i]
# Return the best value and its inputs
return bestval, best
# Creates a population not knowing any previous information
# n - dimensionality of output
# k - population size
def create_population(n, k):
# Arbitrary range, but should be fine for the time being.
return [[random.random() * 200 - 100 for _ in range(n)] for _ in range(k)]
# Creates the next generation of input sets
# pop - the old population
# scaling - the scaling parameter
def create_next_population(pop, scaling = 0.5):
dim = len(pop[0])
n = len(pop)
newpop = [None] * n
for i in range(n):
a, b = random.randint(0, n-1), random.randint(0, n-1) # Indices of two random elements
diff = [(pop[a][d] - pop[b][d]) for d in range(dim)] # Difference of two random input vectors
newpop[i] = [pop[i][d] + diff[d] * scaling for d in range(dim)] # Mutated input has been created
return newpop
# Just makes a list of evaluation results
def calculate_fitness(f, pop):
return [f(inputs) for inputs in pop]
# Given a population and fitnesses, returns the best element and its fitness.
def getbest(pop, fitnesses):
best, bestfitness = pop[0], fitnesses[0]
for i in range(1, len(fitnesses)):
if fitnesses[i] < bestfitness:
best, bestfitness = pop[i], fitnesses[i]
return best, bestfitness
# -
# ### Particle Swarm Optimization
# +
# This is a super dodgy PSO that resets move speed to some fixed baseline at random times.
# f - the function.
# n - input dimensionality
# k - population size
# loops - how many loops it will do before giving up finding a better solution
# lr - how much velocity affects future velocity (between 0 and 1)
# c1 and c2 - weighting for personal and overall best when moving (between 0 and 1)
# res_speed - the speed we may reset to randomly
# res_speed_p - probability of reset
# Outputs the minimum value of the function that it found and the necessary input for it
def pso(f, n, k=25, loops=25, lr=1.0, c1=0.5, c2=0.5, res_speed=10, res_speed_p=0.1):
# Create initial population - including velocity, personal best locations
pop, velocity, pb_locs = create_pso(n, k)
# Also calculate personal best actual values.
pb_vals = calculate_fitness(f, pb_locs)
vals = pb_vals[::]
g_best_loc, g_best_val = getbest(pop, vals)
loops_since_improvement = 0
while loops_since_improvement < loops:
loops_since_improvement += 1
# Create the next generation - updates population, velocity
iterate_pso(pop, velocity, pb_locs, g_best_loc, lr, c1, c2, res_speed, res_speed_p)
# Now update values, personal bests, global bests
vals = calculate_fitness(f, pop)
update_personal_best(pb_vals, pb_locs, vals, pop)
next_best_loc, next_best_val = getbest(pop, vals)
if next_best_val < g_best_val:
loops_since_improvement = 0
g_best_loc, g_best_val = next_best_loc, next_best_val
return g_best_val, g_best_loc # best output and input
# n - input dimensionality
# k - population size
def create_pso(n, k):
pop = create_population(n, k) # Just use the same population init as DE
velocity = norm(create_population(n, k), 10)
pb_locs = pop[::]
return pop, velocity, pb_locs
# Iterates the PSO state.
# pop - current locations
# velocity - how fast we are moving and where
# pb_locs - the best positions value wise each element has been to
# best_loc - globally the best position that everybody also wants to move toward
# lr - how much velocity affects future velocity (between 0 and 1)
# c1 and c2 - weighting for personal and overall best when moving (between 0 and 1)
# res_speed - the speed we may reset to randomly
# res_speed_p - probability of reset
def iterate_pso(pop, velocity, pb_locs, best_loc, lr, c1, c2, res_speed, speed_res_p):
for i in range(len(pop)):
z1, z2 = random.random(), random.random()
velocity[i] = list(np.add(lr * np.array(velocity[i]),
np.add(c1*z1*np.subtract(pb_locs[i], pop[i]), c2*z2*np.subtract(best_loc, pop[i]))))
pop[i] = list(np.add(pop[i], velocity[i]))
if random.random() < speed_res_p:
norm(velocity, res_speed) # I will basically reset the speed my swarm moves at randomly - seems to help...
# does what it says
def update_personal_best(pb_vals, pb_locs, vals, locs):
for i in range(len(vals)):
if vals[i] < pb_vals[i]:
pb_vals[i], pb_locs[i] = vals[i], locs[i]
# updates a list of vectors in place such that they get a certain length
# vectors - list of vectors that all have the same dimensions
def norm(vectors, tolength=1):
if len(vectors) == 0:
return vectors
dim = len(vectors[0])
# length of vector is the sqrt of its dot product with itself
# simply divide each component with that value
for i in range(len(vectors)):
length = np.dot(vectors[i], vectors[i])**0.5
for j in range(dim):
vectors[i][j] /= length * tolength
return vectors
# -
# ### Genetic Algorithm
# +
# Does as the others do.
# f - the function.
# n - input dimensionality
# k - population size
# loops - how many loops it will do before giving up finding a better solution
def ga(f, n, k=100, loops=25):
# Create initial input set
pop = create_population(n, k)
fitnesses = calculate_fitness(f, pop)
sorted(list(zip(pop, fitnesses)))
best, bestval = getbest(pop, fitnesses) # pair of best input and its value
loops_since_improvement = 0 # Keep it going until it's not working anymore.
while loops_since_improvement < loops:
loops_since_improvement += 1
better_half = list(map(lambda x: x[0], sorted(list(zip(pop, fitnesses)), key=lambda x:x[1])[:k//2]))
next_half = crossover(better_half)
mutate(next_half) # chaotic
pop = better_half + next_half
fitnesses = calculate_fitness(f, pop)
nextbest, nextbestval = getbest(pop, fitnesses)
if nextbestval < bestval:
best, bestval = nextbest, nextbestval
loops_since_improvement = 0
# Return the best value and its inputs
return bestval, best
# random crossover between the better elements
def crossover(inputs):
next_inputs = []
for _ in range(len(inputs)):
next_input = []
a, b = random.choice(inputs), random.choice(inputs)
for i in range(len(a)): # for each dimension of input
next_input.append(random.choice([a[i], b[i]]))
next_inputs.append(next_input)
return next_inputs
# performs mutation
def mutate(inputs):
for x in inputs:
if random.random() < 0.5: # let some of them be
for i in range(len(x)): # the mutation here is scaling, s'all
x[i] *= random.random() * 1.5 + 0.5
# -
# # Sanity checking
# ### Testing Goodness of Differential Evolution
#
# I expect scipy and diff_evo to be pretty close overall by quality of result. Turns out, they are, at least in this test.
# +
# Generate a bunch of functions. Use scipy and diff_evo to find optimal solutions.
# Arbitrarily choose amount of functions for each dimensionality.
functions = 10
print('Measuring difference of scipy and diff_evo - the higher, the better for DE')
for dimensions in range(1,5):
print(f'Testing out {dimensions} dimensions...')
for _ in range(functions):
function = quartic_n(dimensions)
sp_ans = minimize(function, [0] * dimensions)
sp_bestval, sp_bestinput = sp_ans.fun, sp_ans.x
bestval, bestinput = diff_evo(function, dimensions)
print(f'Difference between scipy and diffevo: {sp_bestval - bestval}')
print()
# -
# ### Testing Goodness of Particle Swarm Optimization
#
# It's not bad. Sometimes gets a really good result in the end, thousands below the scipy default optimizer.
# +
functions = 10
print('Measuring difference of scipy and PSO - the higher, the better for PSO')
for dimensions in range(1,5):
print(f'Testing out {dimensions} dimensions...')
for _ in range(functions):
function = quartic_n(dimensions)
sp_ans = minimize(function, [0] * dimensions)
sp_bestval, sp_bestinput = sp_ans.fun, sp_ans.x
bestval, bestinput = pso(function, dimensions)
print(f'Difference between scipy and PSO: {sp_bestval - bestval}')
print()
# -
# ### Testing Goodness of Genetic Algorithm
#
# Surprisingly enough, it actually works. The default parameters might not be the best, though.
# +
functions = 10
print('Measuring difference of scipy and GA - the higher, the better for GA')
for dimensions in range(1,5):
print(f'Testing out {dimensions} dimensions...')
for _ in range(functions):
function = quartic_n(dimensions)
sp_ans = minimize(function, [0] * dimensions)
sp_bestval, sp_bestinput = sp_ans.fun, sp_ans.x
bestval, bestinput = ga(function, dimensions)
print(f'Difference between scipy and GA: {sp_bestval - bestval}')
print()
# -
# # Comparing
# ### Finding good parameters
#
# I'll be trying to find the fastest set of parameters for these methods such that they still get decent results most of the time. Say, better or within a few percentage points at least 90% of the time.
# We'll be using this function to generate tests. Parameters are all
# but the first two (function and dimensionality)
def test(f, parameters):
suitable = 0
n = 100
target = 90
dim = 4
full_params = [None, dim] + parameters
for _ in range(n):
q = quartic_n(dim)
full_params[0] = q
sp_ans = minimize(q, np.zeros(dim)).fun
f_ans = f(*full_params)[0]
# Better answer or very little absolute difference or small relative difference
abs_diff = abs(f_ans - sp_ans)
rel_diff = abs(f_ans) / abs(sp_ans)
if f_ans < sp_ans or abs_diff < 1 or rel_diff > 0.95 and rel_diff < 1.05:
suitable += 1
return suitable, suitable >= target
# Get some values with trial and error.
random.seed(42)
test(diff_evo, [30, 0.25, 25]) # Smaller jumps seem to help a lot and let us cut population quite a bit
# +
random.seed(42)
#test(pso, [10, 20, 1.0, 0.75, 0.75, 10, 0.2]) # Speed reset strangely helps a lot with bad params
#test(pso, [10, 20, 0.75, 0.75, 0.75, 10, 0.0]) # Without speed reset but decent learning rate picks, takes way longer
test(pso, [14, 20, 0.8, 0.75, 0.75, 10, 0.1]) # Doing both, seems okay, but doesn't help lower population anyway
# Increasing population seems to actually help converge faster - at least here, where fitness calculations are cheap.
# +
random.seed(42)
test(ga, [400, 25])
# The usual GA doesn't do the best job, but its performance could be changed by so many
# details in the implementation (how many children do you make, how many of the best you pick,
# how you do crossover and mutation).
# -
# ### Timing the methods
#
# As one might expect, these may not be the greatest methods to use for this task.
# %timeit -r 1 -n 1 test(diff_evo, [30, 0.25, 25])
# %timeit -r 1 -n 1 test(pso, [14, 20, 0.8, 0.75, 0.75, 10, 0.1])
# %timeit -r 1 -n 1 test(ga, [400, 25])
# Just for fun, see how much of this time scipy minimization would take
# %timeit -r 1 -n 1 [minimize(quartic_n(4), (0,0,0,0)) for _ in range(100)]
# # Graphs
#
# I'll now modify the main driving functions to have them return not the best thing it found in the end, but history of best fitnesses.
# +
def diff_evo(f, n, k = 80, scaling = 0.5, loops=25):
# Create initial input set
pop = create_population(n, k)
fitnesses = calculate_fitness(f, pop)
best, bestval = getbest(pop, fitnesses) # pair of best input and its value
his = [bestval]
loops_since_improvement = 0 # Keep it going until it's not working anymore.
while loops_since_improvement < loops:
loops_since_improvement += 1
newpop = create_next_population(pop, scaling)
newfitnesses = calculate_fitness(f, newpop)
nextbest, nextbestval = getbest(newpop, newfitnesses)
if nextbestval < bestval:
best, bestval = nextbest, nextbestval
loops_since_improvement = 0
for i in range(k):
if newfitnesses[i] < fitnesses[i]: # if something must be changed
pop[i], fitnesses[i] = newpop[i], newfitnesses[i]
his.append(bestval)
return his
def pso(f, n, k=25, loops=25, lr=1.0, c1=0.5, c2=0.5, res_speed=10, res_speed_p=0.1):
pop, velocity, pb_locs = create_pso(n, k)
pb_vals = calculate_fitness(f, pb_locs)
vals = pb_vals[::]
g_best_loc, g_best_val = getbest(pop, vals)
his = [g_best_val]
loops_since_improvement = 0
while loops_since_improvement < loops:
loops_since_improvement += 1
iterate_pso(pop, velocity, pb_locs, g_best_loc, lr, c1, c2, res_speed, res_speed_p)
vals = calculate_fitness(f, pop)
update_personal_best(pb_vals, pb_locs, vals, pop)
next_best_loc, next_best_val = getbest(pop, vals)
if next_best_val < g_best_val:
loops_since_improvement = 0
g_best_loc, g_best_val = next_best_loc, next_best_val
his.append(g_best_val)
return his
def ga(f, n, k=100, loops=25):
pop = create_population(n, k)
fitnesses = calculate_fitness(f, pop)
sorted(list(zip(pop, fitnesses)))
best, bestval = getbest(pop, fitnesses)
his = [bestval]
loops_since_improvement = 0
while loops_since_improvement < loops:
loops_since_improvement += 1
better_half = list(map(lambda x: x[0], sorted(list(zip(pop, fitnesses)), key=lambda x:x[1])[:k//2]))
next_half = crossover(better_half)
mutate(next_half)
pop = better_half + next_half
fitnesses = calculate_fitness(f, pop)
nextbest, nextbestval = getbest(pop, fitnesses)
if nextbestval < bestval:
best, bestval = nextbest, nextbestval
loops_since_improvement = 0
his.append(bestval)
return his
# -
# Now we may plot these very easily.
# +
function = quartic_n(4)
de_his = diff_evo(function, 4, 30, 0.25, 25)
ps_his = pso(function, 4, 14, 20, 0.8, 0.75, 0.75, 10, 0.1)
ga_his = ga(function, 4, 400, 25)
# -
def plot(data, method):
sns.lineplot(data = data)
plt.xlabel('Generation')
plt.ylabel('Best value')
plt.title(f'History of best value by generation using {method}')
plt.show()
plot(de_his, 'Differential Evolution')
plot(ps_his, 'Particle Swarm')
plot(ga_his, 'Genetic Algorithm (generic?)')
# Perhaps it would be a good idea to cut out the first some generations...
plot(de_his[10:], 'Differential Evolution (starting from the tenth gen)')
plot(ps_his[10:], 'Particle Swarm (starting from the tenth gen)')
plot(ga_his[10:], 'GA (generic?) (starting from the tenth gen)')
# It is interesting to see differential evolution, the fastest method so far, go through the most generations. For all of them we see that after a certain point that we reach very quickly results improve very little. Chances are these methods could be modified to stop once improvements are not large enough (as opposed to when there has been absolutely no improvement for a while) to gain some speed.
#
# Also of note is that even if DE takes the least time to run already, it also requires a relatively small *fraction* of its runtime to reach an acceptable point, so possibly it is also the most optimizable.
| clear/Polynomials.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercises for EBT617E - 2D Band structure
# See Homework \#2
# +
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
# +
# define 2d BZ for a square lattice
Nk = 128
kk, dk = np.linspace(-np.pi, np.pi, Nk, endpoint=True, retstep=True)
# +
# reciprocal lattice
nmax = 2
dimx = 2*nmax+1
nn = np.arange(-nmax,nmax+1)
mGx, mGy = np.meshgrid(nn,nn)
# -
mGx, mGy
# +
# let's enumerate states in the reciprocal state for each k: k + G
states = mGx+nmax + (mGy+nmax)*dimx
# -
states
# k in [-pi,pi]
def ϵkG(kx,ky,Gx,Gy):
return (kx+Gx)**2+(ky+Gy)**2
ϵkG(0,0,mGx,mGy)
ϵkG(0,0,mGx,mGy).flatten()
# +
# Let's do V(x,y) = Vm*( cos(2πx/a)/2 + cos(2πy/a) )
Vmax = 1
dF = 1
FCoeffs = np.zeros([2*dF+1,2*dF+1])
FCoeffs[-1+dF, 0+dF] = Vmax/2
FCoeffs[ 0+dF, 0+dF] = 0
FCoeffs[ 1+dF, 0+dF] = Vmax/2
FCoeffs[ 0+dF,-1+dF] = Vmax
FCoeffs[ 0+dF, 0+dF] = 0
FCoeffs[ 0+dF, 1+dF] = Vmax
# -
FCoeffs
for iFx, FCoeffx in enumerate(FCoeffs):
for iFy, FCoeff in enumerate(FCoeffx):
print(iFx,iFy,FCoeff)
# +
# let's plot this potential in real space
Nx = 101
xmax = 1.5
xx = np.linspace(-xmax,xmax,Nx,endpoint=True)
mx, my = np.meshgrid(xx,xx)
nF = np.arange(-dF,dF+1)
mGFx, mGFy = np.meshgrid(nF,nF)
expkR= np.exp( 1j * 2*np.pi * ( np.outer(mx,mGFx) + np.outer(my,mGFy) ) )
# -
Vr = expkR@FCoeffs.flatten()
plt.imshow(Vr.real.reshape(Nx,Nx))
plt.imshow(Vr.imag.reshape(Nx,Nx))
def FMat(FCoeffs, nmax):
dimx = 2*nmax+1
dB = dimx*dimx
MM = np.zeros([dB,dB])
dimF = len(FCoeffs)
dimFx = (dimF-1)//2
#print(dimFx,dimF)
for iFx, FCoeffx in enumerate(FCoeffs):
for iFy, FCoeff in enumerate(FCoeffx):
index_shift = iFx-dimFx + (iFy-dimFx)*dimF
#print(iFx-dimFx,iFy-dimFx,FCoeff,index_shift)
MM+=np.diag([FCoeff]*(dB-np.abs(index_shift)),index_shift)
return MM # +MM.T
plt.imshow(FMat(FCoeffs, nmax))
VFMat = FMat(FCoeffs, nmax)
def Hband(kx,ky,nmax,VFMat):
#dimx = 2*nmax+1
#dB = dimx*dimx
nn = np.arange(-nmax,nmax+1)
mGx, mGy = np.meshgrid(nn,nn)
HH = np.diag(ϵkG(kx,ky,mGx,mGy).flatten())
HH+= VFMat
return HH
kx=0.
ky=0.
Hb = Hband(kx,ky,nmax,VFMat)
plt.imshow(Hb)
# +
Ebands=[]
for ky in kk:
Eband=[]
for kx in kk:
Hb = Hband(kx,ky,nmax,VFMat)
ee, uu = np.linalg.eigh(Hb)
Eband.append(ee)
Ebands.append(Eband)
Ebands=np.array(Ebands)
# -
Ebands.shape
plt.imshow(Ebands[:,:,0])
from mpl_toolkits import mplot3d
# syntax for 3-D projection
ax = plt.axes(projection ='3d')
ax.plot_surface(kk,kk,Ebands[:,:,0])
# plot the first few bands for ky=0
for ni,n in enumerate(nn[:3]):
plt.plot(kk,Ebands[:,len(kk)//2,ni],'-')
# +
# plot eigenfunction in real-space
# -
| 2d_band_structure.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] deletable=true editable=true
# # Board Game Geek Data Analysis: Example 2
#
# ## Introduction
# For this data science project, I have created multiple python scripts that have scraped https://boardgamegeek.com
#
# As part of the web scraping/data mining, I have created a Postgres database where all of my board game data has been stored. A total of 10,000 board games were scraped from the website, using the Python packages "requests" and "BeautifulSoup", as well as the [XML formatted API](https://boardgamegeek.com/wiki/page/BGG_XML_API&redirectedfrom=XML_API#) that is provided by boardgamegeek.com.
#
# All of the data analysis is built around SQL queries to my boardgame database, for which I use the Python package "psycopg2" to communicate.
#
# The full list of data scraped for each boardgame is:
# * __id__: Unique ID assigned to the boardgame by boardgamegeek.com
# * __rank__: Current rank of the boardgame at the time of scraping
# * __name__: Name of boardgame
# * __href__: Part of URL that identifies this boardgame. Append it to https://boardgamegeek.com to go to game's page
# * __pub_year__: Year the boardgame was published
# * __geek_rating__: Current geek rating the boardgame has at the time of scraping. This is what the game is ranked by. Rating is weighted by the number of voters
# * __avg_rating__: Current average rating the boardgame has at the time of scraping. This is based off the people who have rated it
# * __num_voters__: Number of people who have rated the boardgame at the time of scraping
# * __min_players__: Minimum number of players for the boardgame
# * __max_players__: Maximum number of players for the boardgame
# * __play_time__: Estimated time it takes to play the boardgame
# * __sugg_age__: Suggested minimum age to play the boardgame
# * __complx_rating__: A rating based off votes by players for how difficult a boardgame is to learn and play
# * __designers__: List of designers who made the boardgame
# * __artists__: List of artists for the boardgame
# * __categories__: List of categories the boardgame was placed in
# * __mechanics__: List of gaming mechanics used in the boardgame
# * __family__: List of families the boardgame is grouped into
# * __type__: List of types the boardgame is classified under
#
# ## Project Proposal
#
# Boardgames are a very popular activity for people and boardgamegeek.com has a very active community. I believe it's possible to leverage the immense ammount of data on their website in order to investigate what makes a boardgame highly rated. There are many possibilites that could influence what makes a game well rated and it's likely that the data is very intertwined, where no one factor can really determine a game's success. This provides a difficult problem that machine learning/deep learning may be able to solve. At the very least there's plenty of data, including personal user data on the website that hasn't been tapped, which could help build a recommendation system that could recommend similar boardgames to a user, based on their interests.
#
# Project Ideas:
# * Use machine learning to learn what makes a highly rated game. Allow users to select game features and have the machine learning algorithm estimate a games rating.
# * Build a recommendation system. Recommend similar games to a user, based on their interests. Can leverage just boardgame data, boardgamegeek user data (requires more scraping), or both.
# * Build a chatbot that can talk with a user and help guide them through finding a game that they'd like to play/purchase. Use natural language processing and deep learning, combined with the data scraped from boardgamegeek.com.
#
# ## Exploratory Data Analysis: Game Designers
#
# An idea I wanted to investigate was if a boardgame's suggested player age trends with a boardgame's complexity rating, which users of the site vote on. More specifically, how does this distribution look for a specific element of a boardgame, such as its designer, artist, etc. The complexity rating is a measure of how complicated a game is to play and is on a scale from 1-5. For now I will focus on boardgame designers.
#
# Let's first look at some of the top game designers and how many boardgames they've designed. The function __count_column__ sends a simple SQL query to my database:
# ```
# SELECT designer, COUNT(bg_id) FROM designers GROUP BY designer ORDER BY COUNT(bg_id) DESC;
# ```
# + deletable=true editable=true
import data_analysis
des_count = data_analysis.count_column('designers')
for designer, num_games in des_count[:10]:
print '%20s %5s' %(designer, num_games)
# + [markdown] deletable=true editable=true
# We can see that 303 games did not have designers assigned to the boardgame on boardgamegeek.com. I'm personally amazed one man could design 191 games! Lets see how the top three most frequent boardgame designers compare with each other with respect to their games' complexity and suggested player age. During the SQL query to my database, I had to make sure I wasn't looking at games that were not not given a rating for boardgame complexity (i.e. rating of 0). This was easily achieved by using the WHERE statement in the SQL language. The querying required for the data in the plot below is found in the __comp_rating_sugg_age_for_item_in_column__ function in the [__data_analysis.py__](https://github.com/rolison/BoardGameGeek-DataScience/blob/master/data_analysis.py) script in my GitHub repo. The query looks like this:
# ```
# SELECT boardgames.complx_rating,
# boardgames.sugg_age
# FROM designers
# INNER JOIN boardgames
# ON boardgames.id=designers.bg_id
# WHERE designers.designer = %s
# AND boardgames.complx_rating > 0
# AND boardgames.sugg_age > 0;
# ```
# The ```%s``` option is a variable passed by the user indicating which specific designer to collect the data on.
#
# The __scatter_plot_complxr_vs_sugg_age__ function (and therefore the __comp_rating_sugg_age_for_item_in_column__ function) can make this plot for any column containing a list of text inside the boardgames table in my database. Those columns are: _designers_, _artists_, _categories_, _mechanics_, _family_, and _type_.
# + deletable=true editable=true
data_analysis.scatter_plot_complxr_vs_sugg_age('designers', ['<NAME>', '<NAME>','<NAME>'])
# + [markdown] deletable=true editable=true
# Overall, there does in fact seem to be a trend where the complexity rating of a boardgame increases with suggested player age. <NAME>, the German who has created 191 boardgames, seems to have designed games across the spectra, from simple children's games to more complex ones for an older audience. Wolfgang seems to mostly make games within the 8-10+ age group, where they're of normal difficulty. Richard, on the other hand, seems to make some very complex games that only teenagers and adults may have the attention span/capacity to learn and play. After quickly checking out Richard's biography on boardgamegeek.com, the data makes sense: he is a reknowned Wargame designer. Finding ways to accurately depict war and wartime strategy throughout the ages can create very complicated and nuanced rules for a game. You'd have to really love the stuff to want to play it.
#
# ### Conclusion
#
# For some boardgame designers, we can clearly see a trend in the kind of games they create, with respect to its complexity and age demographic. This is only feasible to predict when the designer has made many games, such conclusions can't be drawn when they've made only one or two. It may be possible to further filter the results above by singling out certain boardgame categories or mechanincs as well, to see what style of games a designer typically creates.
| bggDataAnalysis2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py36_Global] *
# language: python
# name: conda-env-py36_Global-py
# ---
# # Equilibrium urea denaturation curve
# CD spectra were recorded at 222 nm (minima for PpiA and PpiB) from 0 to 8M Urea
# %matplotlib inline
from symfit import parameters, Variable, Fit, exp
import matplotlib.pyplot as plt
import numpy as np
curve = np.genfromtxt('CD_PpiB_urea_curve.txt').T
curve.shape
# +
#plot raw data
xdata = curve[0]
Fdata = curve[1]
plt.xlabel('Urea')
plt.ylabel('Folded fraction')
plt.scatter(xdata,Fdata)
# +
# model formula for fitting with 2 parameters and one variable
# equation from pyfolding: https://github.com/quantumjot/PyFolding
# https://github.com/quantumjot/PyFolding/blob/3e343644f70d6cfe5e552e7c8ec5da76acb1d8c5/pyfolding/models.py#L78
m, d50 = parameters('m d50')
x = Variable('x')
RT=8.345*(273.15+25)
model = exp((m*(x-d50))/RT) / (1+exp((m*(x-d50))/RT))
model
# + pycharm={"name": "#%%\n"}
#fitting
fit = Fit(model, xdata, Fdata)
fit_result = fit.execute()
# -
#Printing the parameters from the fit with d50 the urea concentration for F=0.5 and m-value describing the slope
print(fit_result)
# Plotting the fit on a figure with good resolution (with x 100 steps between 0-8M Urea) and for Y the fit parameters
xmodel = np.linspace(0,8,num=100,endpoint=True)
ymodel = model(xmodel,**fit_result.params)
plt.plot(xmodel,ymodel)
plt.scatter(xdata, Fdata, color='k')
np.savetxt('CD_PpiB urea curve_fit2', ymodel)
| Circular_Dichroism/CD_Urea_denaturation_PpiB.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Exercise Week 6**
#
# **Author: <NAME>**
#
# **Email: <EMAIL>**
# +
def questionOne():
def dictionary(keys, values):
return dict(zip(keys, values))
## sample
keys = ['Phone', 'Tablet', 'Laptop']
values = ['iPhone 11', 'iPad Mini', 'Macbook Pro']
result = dictionary(keys, values)
return result
questionOne()
# +
from math import sqrt
## run to install humanize: pip install humanize
from humanize import number
def questionTwo():
def fibonacci(n):
if (n < 0):
return "Sorry, fibonacci start from 0"
if (n == 0):
return f"Next fibonacci number is {1}, and it is the 2nd Fibonacci number"
if (n == 1):
return f"Next fibonacci number is {1}, and it is the 3rd Fibonacci number"
n_2 = 1
n_1 = 1
position = 4
sums = n_2 + n_1
while (sums <= n):
position = position + 1
# Update the first
n_2 = n_1
# Update the second
n_1 = sums
# Update the third
sums = n_2 + n_1
return f"Next fibonacci number is {sums}, and it is the {number.ordinal(position)} Fibonacci number"
print(fibonacci(-1))
print(fibonacci(0))
print(fibonacci(14))
print(fibonacci(60))
print(fibonacci(55))
questionTwo()
# +
import numpy as np
def questionThree():
def scoring(input_array, approved, not_approved):
in_approved = [item for item in input_array if item in approved]
in_not_approved = [item for item in input_array if item in not_approved]
total_approved = len(in_approved)
total_not_approved = len(in_not_approved)
return total_approved - total_not_approved
print(scoring(
[1, 2, 3, 4, 10, 9, 8, 7],
[1, 10, 11],
[8, 5]
))
print(scoring(
[1, 1, 1, 5, 5, 2, 3, 10],
[1, 3],
[5]
))
questionThree()
| Exercise Week 6 - Ahmad Ichsan Baihaqi.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/HarvinderSinghDiwan/DEEP_LEARNING_SESSION/blob/master/DEEP_LEARNING.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="6ZuhdGD21p3f" colab_type="code" colab={}
### NEURAL NETWORKS ###
######ACTIVATION FUNCTION MAKES LEARN NON LINEAR RELATIONSHIP
###### OVERFITTING
###### (w.b)FORWARD PROPAGATION
####### loss function, cost function, objective function
##### GRADIENT DESCENT
### USE BACKWORD PROPAGATION
# + id="uXrM91vU3BgO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 678} outputId="4c3c1f2e-efba-447e-98cc-862d1cbe7b1e"
# #!/usr/bin/env python
# coding: utf-8
# ### Import all libraries
# In[1]:
#importing all the libraries
import pandas as pd
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
from sklearn.metrics import accuracy_score
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
# ### Load datasets and Normalize
# In[2]:
df = pd.read_csv('/content/wine.csv')
# print(df)
a = pd.get_dummies(df['Wine'])
df = pd.concat([df,a],axis=1)
X = df.drop([1, 2,3,'Wine'], axis = 1)
y = df[[1,2,3]].values
X_train, X_test, Y_train,Y_test = train_test_split(X, y, test_size=0.20,)
scaler = StandardScaler()
scaler.fit(X_train)
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
# Y_test,test
# ### Explore dataset
# In[3]:
print(df.head())
# In[5]:
print(df.tail())
# ### Forward Propagation function
# In[6]:
def forward_prop(model,a0):
# Load parameters from model
W1, b1, W2, b2, W3, b3 = model['W1'], model['b1'], model['W2'], model['b2'], model['W3'],model['b3']
# Do the first Linear step
# Z1 is the input layer x times the dot product of the weights + bias b
z1 = a0.dot(W1) + b1
# Put it through the first activation function
a1 = np.tanh(z1)
# Second linear step
z2 = a1.dot(W2) + b2
# Second activation function
a2 = np.tanh(z2)
#Third linear step
z3 = a2.dot(W3) + b3
#For the Third linear activation function we use the softmax function, either the sigmoid of softmax should be used for the last layer
a3 = softmax(z3)
#Store all results in these values
cache = {'a0':a0,'z1':z1,'a1':a1,'z2':z2,'a2':a2,'a3':a3,'z3':z3}
return cache
# ### Softmax Activation function
# In[7]:
def softmax(z):
#Calculate exponent term first
exp_scores = np.exp(z)
return exp_scores / np.sum(exp_scores, axis=1, keepdims=True)
# ### Backpropagation function
# In[8]:
def backward_prop(model,cache,y):
# Load parameters from model
W1, b1, W2, b2, W3, b3 = model['W1'], model['b1'], model['W2'], model['b2'],model['W3'],model['b3']
# Load forward propagation results
a0,a1, a2,a3 = cache['a0'],cache['a1'],cache['a2'],cache['a3']
# Get number of samples
m = y.shape[0]
# Calculate loss derivative with respect to output
dz3 = loss_derivative(y=y,y_hat=a3)
# Calculate loss derivative with respect to second layer weights
dW3 = 1/m*(a2.T).dot(dz3) #dW2 = 1/m*(a1.T).dot(dz2)
# Calculate loss derivative with respect to second layer bias
db3 = 1/m*np.sum(dz3, axis=0)
# Calculate loss derivative with respect to first layer
dz2 = np.multiply(dz3.dot(W3.T) ,tanh_derivative(a2))
# Calculate loss derivative with respect to first layer weights
dW2 = 1/m*np.dot(a1.T, dz2)
# Calculate loss derivative with respect to first layer bias
db2 = 1/m*np.sum(dz2, axis=0)
dz1 = np.multiply(dz2.dot(W2.T),tanh_derivative(a1))
dW1 = 1/m*np.dot(a0.T,dz1)
db1 = 1/m*np.sum(dz1,axis=0)
# Store gradients
grads = {'dW3':dW3, 'db3':db3, 'dW2':dW2,'db2':db2,'dW1':dW1,'db1':db1}
return grads
# ### Loss/Objective/Cost function
# In[9]:
def softmax_loss(y,y_hat):
# Clipping value
minval = 0.000000000001
# Number of samples
m = y.shape[0]
# Loss formula, note that np.sum sums up the entire matrix and therefore does the job of two sums from the formula
loss = -1/m * np.sum(y * np.log(y_hat.clip(min=minval)))
return loss
# ### Loss and activation derivative for backpropagation
# In[10]:
def loss_derivative(y,y_hat):
return (y_hat-y)
def tanh_derivative(x):
return (1 - np.power(x, 2))
# ### Randomly initialize all Neural Network parameters
# In[11]:
def initialize_parameters(nn_input_dim,nn_hdim,nn_output_dim):
# First layer weights
W1 = 2 *np.random.randn(nn_input_dim, nn_hdim) - 1
# First layer bias
b1 = np.zeros((1, nn_hdim))
# Second layer weights
W2 = 2 * np.random.randn(nn_hdim, nn_hdim) - 1
# Second layer bias
b2 = np.zeros((1, nn_hdim))
W3 = 2 * np.random.rand(nn_hdim, nn_output_dim) - 1
b3 = np.zeros((1,nn_output_dim))
# Package and return model
model = { 'W1': W1, 'b1': b1, 'W2': W2, 'b2': b2,'W3':W3,'b3':b3}
return model
# ### Update Parameters
# In[12]:
def update_parameters(model,grads,learning_rate):
# Load parameters
W1, b1, W2, b2,b3,W3 = model['W1'], model['b1'], model['W2'], model['b2'],model['b3'],model["W3"]
# Update parameters
W1 -= learning_rate * grads['dW1']
b1 -= learning_rate * grads['db1']
W2 -= learning_rate * grads['dW2']
b2 -= learning_rate * grads['db2']
W3 -= learning_rate * grads['dW3']
b3 -= learning_rate * grads['db3']
# Store and return parameters
model = { 'W1': W1, 'b1': b1, 'W2': W2, 'b2': b2, 'W3':W3,'b3':b3}
return model
# ### Predict function
# In[13]:
def predict(model, x):
# Do forward pass
c = forward_prop(model,x)
#get y_hat
y_hat = np.argmax(c['a3'], axis=1)
return y_hat
# ### Train function
# In[15]:
def train(model,X_,y_,learning_rate, iterations, print_loss=False):
# Gradient descent. Loop over epochs
for i in range(0, iterations):
# Forward propagation
cache = forward_prop(model,X_)
# Backpropagation
grads = backward_prop(model,cache,y_)
# Gradient descent parameter update
# Assign new parameters to the model
model = update_parameters(model=model,grads=grads,learning_rate=learning_rate)
# Pring loss & accuracy every 100 iterations
if print_loss and i % 100 == 0:
a3 = cache['a3']
print('Loss after iteration',i,':',softmax_loss(y_,a3))
y_hat = predict(model,X_)
y_true = y_.argmax(axis=1)
print('Accuracy after iteration',i,':',accuracy_score(y_pred=y_hat,y_true=y_true)*100,'%')
losses.append(accuracy_score(y_pred=y_hat,y_true=y_true)*100)
return model
# ### Initialize model parameters and train model on wine dataset
# In[16]:
model = initialize_parameters(nn_input_dim=13, nn_hdim= 5, nn_output_dim= 3)
model = train(model,X_train,Y_train,learning_rate=0.07,iterations=4500,print_loss=True)
plt.plot(losses)
# ### Calculate testing accuracy
# In[17]:
test = predict(model,X_test)
test = pd.get_dummies(test)
Y_test = pd.DataFrame(Y_test)
print("Testing accuracy is: ",str(accuracy_score(Y_test, test) * 100)+"%")
# + id="8Y_R8Ai9GZrL" colab_type="code" colab={}
def train(model,X_,y_,learning_rate, iterations, print_loss=False):
# Gradient descent. Loop over epochs
for i in range(0, iterations):
# Forward propagation
cache = forward_prop(model,X_)
# Backpropagation
grads = backward_prop(model,cache,y_)
# Gradient descent parameter update
# Assign new parameters to the model
model = update_parameters(model=model,grads=grads,learning_rate=learning_rate)
# Pring loss & accuracy every 100 iterations
if print_loss and i % 100 == 0:
a3 = cache['a3']
print('Loss after iteration',i,':',softmax_loss(y_,a3))
y_hat = predict(model,X_)
y_true = y_.argmax(axis=1)
print('Accuracy after iteration',i,':',accuracy_score(y_pred=y_hat,y_true=y_true)*100,'%')
losses.append(accuracy_score(y_pred=y_hat,y_true=y_true)*100)
return model
# + id="JpYvHEJOIJp9" colab_type="code" colab={}
# ### Predict function
# In[13]:
def predict(model, x):
# Do forward pass
c = forward_prop(model,x)
#get y_hat
y_hat = np.argmax(c['a3'], axis=1)
return y_hat
# + id="LXEtHaEOIOVl" colab_type="code" colab={}
# ### Backpropagation function
# In[8]:
def backward_prop(model,cache,y):
# Load parameters from model
W1, b1, W2, b2, W3, b3 = model['W1'], model['b1'], model['W2'], model['b2'],model['W3'],model['b3']
# Load forward propagation results
a0,a1, a2,a3 = cache['a0'],cache['a1'],cache['a2'],cache['a3']
# Get number of samples
m = y.shape[0]
# Calculate loss derivative with respect to output
dz3 = loss_derivative(y=y,y_hat=a3)
# Calculate loss derivative with respect to second layer weights
dW3 = 1/m*(a2.T).dot(dz3) #dW2 = 1/m*(a1.T).dot(dz2)
# Calculate loss derivative with respect to second layer bias
db3 = 1/m*np.sum(dz3, axis=0)
# Calculate loss derivative with respect to first layer
dz2 = np.multiply(dz3.dot(W3.T) ,tanh_derivative(a2))
# Calculate loss derivative with respect to first layer weights
dW2 = 1/m*np.dot(a1.T, dz2)
# Calculate loss derivative with respect to first layer bias
db2 = 1/m*np.sum(dz2, axis=0)
dz1 = np.multiply(dz2.dot(W2.T),tanh_derivative(a1))
dW1 = 1/m*np.dot(a0.T,dz1)
db1 = 1/m*np.sum(dz1,axis=0)
# Store gradients
grads = {'dW3':dW3, 'db3':db3, 'dW2':dW2,'db2':db2,'dW1':dW1,'db1':db1}
return grads
# + id="WS1JrF0vIkoD" colab_type="code" colab={}
# In[9]:
def softmax_loss(y,y_hat):
# Clipping value
minval = 0.000000000001
# Number of samples
m = y.shape[0]
# Loss formula, note that np.sum sums up the entire matrix and therefore does the job of two sums from the formula
loss = -1/m * np.sum(y * np.log(y_hat.clip(min=minval)))
return loss
# + id="_JIs4pe5JUtQ" colab_type="code" colab={}
def update_parameters(model,grads,learning_rate):
# Load parameters
W1, b1, W2, b2,b3,W3 = model['W1'], model['b1'], model['W2'], model['b2'],model['b3'],model["W3"]
# Update parameters
W1 -= learning_rate * grads['dW1']
b1 -= learning_rate * grads['db1']
W2 -= learning_rate * grads['dW2']
b2 -= learning_rate * grads['db2']
W3 -= learning_rate * grads['dW3']
b3 -= learning_rate * grads['db3']
# Store and return parameters
model = { 'W1': W1, 'b1': b1, 'W2': W2, 'b2': b2, 'W3':W3,'b3':b3}
return model
# + [markdown] id="LaoPPN03Jxj9" colab_type="text"
#
# + id="cbKNpyZLJdkg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="d576841f-1282-4f6f-f03a-da5676d6063b"
df.head()
# + id="bB4bP9DTJpg4" colab_type="code" colab={}
| DEEP_LEARNING.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Aula 02 Crash Course Pandas
#
# ## Objetivo:
#
# Objetivo desta aula é apresentar transformações usuais em base de dados usando pandas.
#
# ## Roteiro da aula:
#
# - Relembrando a aula passada
#
# - Dúvidas
#
# - O que é um dataframe
#
# - Importação
#
# - Indetificação de tipos de variáveis
#
# - Seleção de colunas e filtros
#
#
# - Dados Faltantes
#
# - O que fazer com NaNs
#
#
# - Variáveis Categóricas
#
# - Categorical dtype
#
# - pd.Categorical
#
# - .describe() em colunas categóricas
#
# - Método mutador (em inglês, 'accessors')
#
# - One Hot Encoder Pandas - WARNING: Categoria não é um array numpy
#
#
# - Metodologia Split-Apply-Combine
#
# - Grupos - Estrutura de MultiIndex
#
# - Usando apply em grupos
#
# - Função Aggregate
#
# - Apply em diferentes funções no grupo.
#
# - Função Transform
#
# - função filter
#
# - nlargest e nsmallest
#
# - Grupos por factor
#
# - Pipe: transformações em sequência
#
# - Tabelas Pivot
#
#
#
# ## Referências
#
# - [Extensões do Jupyter: artigo no Medium](https://medium.com/@seymatas/5-jupyter-notebook-extensions-for-a-fully-organized-data-science-project-d764195f91be)
#
# - [Instalando extensões do jupyter e possíveis erros - slackoverflow](https://stackoverflow.com/questions/37718907/variable-explorer-in-jupyter-notebook)
#
# - [Pandas](https://pandas.pydata.org/)
#
# - [Introdução em 10 minutos do Pandas](https://pandas.pydata.org/pandas-docs/stable/user_guide/10min.html)
#
# - [Documentação Pandas: Guia do usuário](https://pandas.pydata.org/docs/user_guide/index.html#user-guide)
#
# - [GitHub Cheat Sheet Markdown](https://github.com/adam-p/markdown-here/wiki/Markdown-Here-Cheatsheet)
#
# - [Numpy](https://numpy.org/)
#
# - [Kaggle Titanic Data](https://www.kaggle.com/c/titanic/data)
#
# - [Split-Apply-Combine Pandas](https://pandas.pydata.org/pandas-docs/stable/user_guide/groupby.html)
#
# - [Colunas Categóricas](https://pandas.pydata.org/pandas-docs/stable/user_guide/categorical.html)
#
# - [Selecionando colunas pelo tipo de variável](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.select_dtypes.html)
#
#
# ## Contato:
#
# - E-mails:
#
# - <EMAIL>
#
# - <EMAIL>
#
# - <EMAIL>
#
# - Redes Sociais:
#
# - [LinkedIn](https://www.linkedin.com/in/walterwsmf/)
#
# - [Twitter](https://twitter.com/walterwsmf)
# 1. Relembrando a aula passada:
# +
# %%time
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# -
# import dos dados
data = pd.read_csv('/Users/walter/Downloads/CrashCourse/titanic/train.csv')
data.head()
data.tail(10)
data.dtypes
# +
data = data.astype({'Name':'string',
'Sex':'category',
'Survived':'category',
'Pclass': 'category',
'Embarked':'category',
'Cabin':'string',
'Parch':'category',
'SibSp':'category'}
)
data.dtypes
# -
data.describe()
data._get_numeric_data()
# ### Ontem usamos seleção de colunas para identificar correlação
#
# O cuidado que devemos ter é transformar em coluna em categórica caso desejamos calcular algumas funções que dependam do numpy, pois o dtype 'category' não é um array.
# correlação somente funciona com colunas numéricas
data[["PassengerId","Survived","Pclass","Age","SibSp","Parch","Fare"]].corr()
data[["PassengerId","Survived","Pclass","Age","SibSp","Parch","Fare"]].astype({
'Survived': 'float',
'Pclass': 'float',
}).corr()
data.plot(x='Age',y='Fare',kind='scatter')
# # Filtro por classe
data.columns
data['Pclass'].unique()
# +
cor = ['red','green','blue']
plt.figure(figsize=(14,6))
for i in data['Pclass'].unique():
plt.scatter(data[data['Pclass']==i]['Age'],
data[data['Pclass']==i]['Fare'],
color=cor[i-1],
label=f"Classe {i}",
)
print(f"Estatísticas da classe {i}: {data[data['Pclass']==i].describe()}")
plt.legend()
# -
# ### - Faça o box plot equivalente aos dados filtrados por classe para coluna Fare.
# ### Dataset esta quebrado, será que desbalanço deve-se a isso?
#
# - NO caso do dataset do Titanic Kaggle, vemos que existem dois arquivos, Train e Test. Vamos combiná-los.
train = pd.read_csv('/Users/walter/Downloads/CrashCourse/titanic/train.csv')
test = pd.read_csv('/Users/walter/Downloads/CrashCourse/titanic/test.csv')
train.columns
test.columns
train.head()
test.head()
# Queremos concatená-los:
#
# ```python
# pd.concat
# ```
data = pd.concat([test,train],axis=0,sort=False)
data
pd.concat([test,train],axis=1,sort=False)# concat com axis=1 incluir as colunas ao lado
# ### Como concat não reinicia o Index, devemos fazer isso manualmente:
data.reset_index(drop=True,inplace=True)
data.head()
data.tail()
data.shape
del train, test # deletando dataframes (ou outras variáveis)
# É possível somente deletar uma coluna do dataframe, por exemplo:
#
# ```python
# del data['Age']
# ```
#
# deletaria a coluna 'Age'.
data.dtypes
# +
data = data.astype({'Name':'string',
'Sex':'category',
'Survived':'category',
'Pclass': 'category',
'Embarked':'category',
'Cabin':'string',
'Parch':'category',
'SibSp':'category'})
data.dtypes
# +
cor = ['red','green','blue']
plt.figure(figsize=(14,6))
for i in data['Pclass'].unique():
plt.scatter(data[data['Pclass']==i]['Age'],
data[data['Pclass']==i]['Fare'],
color=cor[i-1],
label=f"Classe {i}",
)
print(f"Estatísticas para classe {i}: {data[data['Pclass']==i].describe()}\n")
plt.legend()
# -
# # Podemos trabalhar nos dados faltantes:
#
# 1. Substituindo por valor
#
# 2. Removendo-os
data.isna().sum()
# ## Mas quando é cabível remoção ou a substituição?
#
# - Survived: provavelmente é devido o caso dos dados serem provenientes de uma competição, que deseja prever essa condição ( e por isto não é tabelada a resposta final). Dados faltantes nessa coluna devem-se a natureza de origem/aquisição
#
# - Cabin: é a maioria dos dados faltantes. Existe [debate sobre o quão relevante é a posição de cabine a sobrevivência durante o acidente do Titanic](https://www.encyclopedia-titanica.org/community/threads/which-passengers-most-likely-died-in-their-cabins.2296/) e caso tenham interesse, clique no link. Mas devido a terem muitos dados faltantes (muito mais do que metado dos casos), podemos dizer que a informação que possamos tirar deve ser pouca.
#
# - Age: São poucos dados, mas não é uma variável categórica. Talvez possamos estimar os dados por correlação com outras colunas, mas somente pela análise atual, não é clara.
help(data['Age'].fillna)
data
# # Colunas Categóricas
#
# - Podemos selecionar qualquer grupo de colunas pelo tipo dos elementos pela rotina **.select_dtypes**
help(data.select_dtypes)
data.select_dtypes(include=['category'])
data.select_dtypes(exclude=['category','string'])
# Uma coluna não categorica pode ser transformada pelo astype, como fizemos, ou forçando a classe categorical:
pd.Categorical(data['Sex'].astype('string'))
help(pd.Categorical)
# ### .describe em Categorias
#
# Para colunas categóricas, o **.describe** retorna quantos únicos existe, a classe com maior quantidade (top) e a quantidade amostral na classe com maior quantidade (freq).
data.select_dtypes(include=['category']).describe()
filtro = data['Sex'] == 'male'
filtro.sum()
# ### Método Mutador
#
# Método Mutador é o termo a um método que controla alterações.
#
# Pode ser usando no Pandas para identificar, como um filtro, condições.
#
#
# Mais usados são:
#
# - .str : para colunas categoricas incialmente com string
#
# - .dt : colunas categóricas de tempo
data['Sex'].str.contains("f") # sendo categoria um string incialmente, podemos usar condições de regex
data[data['Sex'].str.contains("f")]
data[~(data['Sex'].str.contains("f"))]
# Se usamos o mutador incorreto, o resultado é um erro:
data['Sex'].dt
# +
try:
z = data['Sex'].dt
except:
z = 0
z
# -
try:
z = data['Sex'].dt
except:
pass
# ### One Hot Encoder Pandas Categorical
#
# Tem vezes que queremos transformar a coluna categórica em uma matriz de zero e um, i.e., transformá-la em variáveis dummies.
#
# O pandas pode processar a coluna categorica de forma natural:
#
# ```python
# pd.get_dummies
# ```
data['Sex']
pd.get_dummies(data['Sex'])
# ### Em todas as colunas categóricas:
pd.get_dummies(data.select_dtypes(include=['category']))
# # Split-Apply-Combine
#
#
# A metodologia split-apply-combine possui a ideia de quebrar o dataset em grupos menores, com suas afinidades, para processá-los (apply/transform) e depois combinar todo resultado final.
data.head()
# Podemos imaginar agrupar pela classe e por intervalo de idade talvez.
#
# O primeiro grupo, por sexo, da-se da seguinte forma:
data.groupby(by=['Sex'])
# Cria um objeto pandas.core.groupby
g = data.groupby(by=['Sex'])
g,type(g)
g.count() # grupo cria sub-níveis de indexação para calculo, por exemplo, contagem de elementos.
g.apply(len) #podemos aplicar uma funcao diretamente nos novos indices do grupo
g['Age'].apply(np.array) #ou em coluna do grupo, neste cenário, o resultado é uma Série.
g['Age'].apply(np.array).reset_index()
# Outras formas de aplicar rotina é especuficar por exemplo as colunas:
def func(group):
return {
'Age media': group['Age'].mean(),
'Age std':group['Age'].std(),
'Age count':group['Age'].count(),
}
g.apply(func)
g.apply(func).apply(pd.Series)
g.describe()
g.first() #primeiros dos grupos
g.last() #últimos listados em cada grupo
g.ngroups #quantidade de grupos. Por que? porque podemos aumentar nivel de sub-indexacao
# # Trabalhando sem NaNs numa coluna específica
df = data.dropna(subset=['Survived']).copy()
df
df['Survived'] = df['Survived'].fillna(value=0).astype(bool)
df.head()
g = df.groupby(by=['Sex','Pclass'])
g.ngroups
g.first()
g.count()
df['Survived'].shape,df['Survived'].shape[0]
g.apply(lambda k: k['Survived'].sum()/df['Survived'].shape[0])
s = g.apply(lambda k: k['Survived'].sum()/df['Survived'].shape[0]).reset_index()
s
# +
#s.loc[0,0]
# -
s = s.rename(columns={0:'PercentageSurvived'},)
s
s.sort_values(by=['PercentageSurvived',],ascending=False)
# ### Podemos selecionar somente um grupo, sabendo as características do multiIndex esperado:
g.get_group(('female',1))
# ### Rotinas Aggregate
#
# Vamos considerar que não queremos incluir as colunas de grupo como indices, podemos usar o termo **as_index=False**, dentro do **groupby**
g = df.groupby(by=['Sex','Pclass'],as_index=False)
# g = df.groupby(by=['Sex','Pclass'],as_index=True)
g
g.sum()
g.aggregate(np.sum)
# ### .describe me grupos permite obter estat'siticas por cada grupo definido:
g = df.groupby(by=['Sex','Pclass'])
g.describe()
# Na prática, faz agregação das funções estaísticas por lista:
g['Age'].agg([np.sum, np.mean, np.std])
g['Age'].agg(
min_Age=pd.NamedAgg(column='min Age', aggfunc='min'),
max_Age=pd.NamedAgg(column='max Age', aggfunc='max'),
average_Age=pd.NamedAgg(column='mean Age', aggfunc=np.mean),
)
# Podemos aplicar as funções no grupo também por colunas em dicionário:
g.agg(
{'Age': np.mean, #na coluna Age aplique a media
'Fare': lambda x: np.std(x), #na coluna Fare aplique a funcao lambda x
})
help(np.std)
# ### Um grupo reage semelhane ao **zip** em python:
for i,j in zip(range(3),['A','B','C']):
print(i,j)
for name, group in g:
print(name)
print(group)
# ### Rotina Transform
#
# Assim como apply retorna novo dataframe com multindexação, o transforma aplica função no grupo,porém retorna a resposta como coluna para dataframe original do qual o grupo fora criado:
g['Age'].apply(np.mean)
df['mean age'] = g['Age'].transform(lambda x: np.mean(x))
df.head()
g['Age'].nlargest(3)
g['Age'].nsmallest(3)
# ### GroupBy por Fator
factor = pd.qcut(df['Age'], [0, .25, .5, .75, 1.])
factor
g2 = df.groupby(by=factor)
g2.mean()
# Um fator muito usado é **pd.Grouper** que permite agrupar uma coluna/série datetime por dias, meses, anos, quinzenas, etc, e desta forma, criar grupos com intervalos temporais regulares.
# # Pipe: re-uso do grupo
#
# Usado quando a funcao precisa chamar novamente grupo dentro dela:
g.pipe(lambda grp: grp['Age'].mean() / grp['Age'].std()).unstack().round(4)
# ### Pivot Table
df.pivot(index=df.index,columns='Sex')
df.pivot_table(index=df.index,columns=['Sex','Pclass'])
| Aula 02 - Crash Course Pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:adventofcode]
# language: python
# name: conda-env-adventofcode-py
# ---
# # Challenge 20
# ## Challenge 20.1
myinput = '/home/fmuinos/projects/adventofcode/2016/ferran/inputs/input20.txt'
# +
def min_func(myinput):
mylist = []
with open(myinput, 'rt') as f:
for line in f:
mylist.append([int(i) for i in line.rstrip().split('-')])
return sorted(mylist, key=lambda x: x[0])
def min_func(l):
c = 0
for i in range(len(l)):
a, b = l.pop(0)
if a <= c <= b:
c = b + 1
return c
# -
# ## Result
l = parse(myinput)
print(min_func(l))
# ## Challenge 20.2
upper_limit = 4294967295
def allowed(l):
c = -1
white = upper_limit + 1
for i in range(len(l)):
a, b = l.pop(0)
if a <= c <= b:
white -= b - c
c = b
elif c < a:
white -= b - a + 1
c = b
return white
l = parse(myinput)
print(allowed(l))
| 2016/ferran/day20.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
df = pd.read_csv('exemplo')
df
df.to_csv("exemplo.csv")
df = pd.read_excel("Exemplo_Excel.xlsx", sheetname='Sheet1')
df
df.to_excel("Exemplo_Excel.xlsx", sheet_name='Sheet1')
df = pd.read_html('http://www.fdic.gov/bank/individual/failed/banklist.html')
type(df)
df[0].loc[100]
| Utils/Pandas/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# # MF013: Introducción a Numpy
# Ing. <NAME>, MSF<br>
# MF-013 Análisis de Inversión<br>
# Clase del 5 de octubre 2021<br>
#
# Maestría de Finanzas, Facultad de Economía<br>
# UANL<br>
# + [markdown] tags=[]
# ## Imagina la siguiente situación
# -
precio_compra = [100, 98, 19, 50,3]
precio_venta = [10, 320, 25, 600, 1]
precio_venta - precio_compra
precio_venta + precio_compra
# ### Función `zip` en un for loop
# +
lista_utilidad = []
for compra, venta in zip(precio_compra, precio_venta):
utilidad = venta - compra
lista_utilidad.append(utilidad)
# -
lista_utilidad
# + [markdown] tags=[]
# ## 
# + [markdown] tags=[]
# ## Qué es NumPy?
# + [markdown] slideshow={"slide_type": "subslide"}
# > *"Numpy es __el__ paquete fundamental para la computación científica en Python. Contiene entre otras cosas:*
# >* _Un poderoso arreglo multidimensional_
# >* _Herramientas para integrar código de C/C++, Fortran_
# >* _Algebra lineal, transformaciones de Fourier, capacidad de generación de números aleatorios._<br><br>
# >
# >_A parte de su uso científico obvio, Numpy puede ser usado de manera eficiente como un contenedor multidimensional de datos, lo que le permite a Numpy integrarse se manera natural con una gran variedad de bases de datos._" www.numpy.org
# + [markdown] tags=[]
# ### Importar NumPy
# -
import numpy as np
# + [markdown] tags=[]
# ### Los arrays de NumPy
# -
precio_compra
precio_venta
# + [markdown] slideshow={"slide_type": "subslide"} tags=[]
# ### Crear un array con la función `array( )`
# -
# #### Crear array a partir de una lista
array_compra = np.array(precio_compra)
array_compra
array_compra[2:]
array_venta = np.array(precio_venta)
array_venta
array_venta - array_compra
array_venta * array_compra
# #### Crear array a partir de valores numéricos
compra = np.array([100, 10, 30])
venta = np.array([110, 20, 40])
compra
# ##### Número dimensiones
compra.ndim
# + [markdown] tags=[]
# #### Array de 2 dimensiones
# -
array_2d = np.array([[1,2,3],[10, 20, 30]])
array_2d
array_2d.ndim
# + [markdown] tags=[]
# #### Cambiar de dimensiones con la función `reshape( )`
# -
a = np.array([1, 2, 3, 4, 5, 6, 7, 8, 9, 10])
a
a.reshape((5,2))
b = a.reshape((5,2))
b.ndim
# + [markdown] tags=[]
# #### Poblar un array con un valor
# -
a
a.fill(1)
a
# #### Transponer un array
b
b = b.transpose()
b
# + [markdown] tags=[]
# #### Cambiar array de 2D a 1D
# -
b.ndim
b = b.flatten()
b
c = np.array([[[1,2,3],[1,2,3],[1,2,3]]])
c.ndim
c.flatten()
# + [markdown] slideshow={"slide_type": "subslide"} tags=[]
# ### Verificar un array
# -
# #### Función `type( )`
type(a)
# #### Verificar el tipo de datos que conforman el array con `.dtype`
c = a.dtype
c
# #### Verificar el número de dimensiones `.ndim`
b.ndim
# #### Ver la forma del array con `.shape`
array_2d.shape
# + [markdown] tags=[]
# ### Crear arrays con la función `arange( )`
# -
a = np.arange(1,11)
a
b = np.arange(10,101, 10)
b
c = np.array([np.arange(1,11), np.arange(10,101,10)])
c
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Multiplicar un array por un valor
# -
b * 10
b + 10
# + [markdown] slideshow={"slide_type": "subslide"} tags=[]
# ### Arrays especiales
# + [markdown] slideshow={"slide_type": "subslide"}
# #### "Zero" Arrays
# -
np.zeros((10,5), dtype='int')
# + [markdown] slideshow={"slide_type": "subslide"}
# #### "Ones" Arrays
# -
np.ones((5,3))
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Array (matriz) identidad
# -
a = np.identity(4)
a
# + [markdown] slideshow={"slide_type": "subslide"} tags=[]
# #### Linspace
# -
np.linspace(0,1,3)
np.linspace(1,10,4)
# + [markdown] tags=[]
# ### Operaciones matemáticas
# -
array_compra ** array_venta
# + [markdown] slideshow={"slide_type": "subslide"} tags=[]
# #### Operaciones matemáticas entre arrays de 2D
# -
# + [markdown] slideshow={"slide_type": "subslide"} tags=[]
# #### Algunas constantes
# -
np.pi
np.e * array_compra
ln = np.log(1)
log_10 = np.log10(1)
array_compra.sum()
# + [markdown] tags=[]
# ### Resumen operaciones matemáticas entre arrays
# + [markdown] slideshow={"slide_type": "slide"}
# Reglas:
# 1. Debe de coincidir los tamaños de los arrays.
# 2. Operadores matemáticos (+, *, exp, log,...) se aplican elemento por elemento.
# 3. Operaciones reductoras (sumar, promeido, std, skew, kurtosis, prod,...) se aplican a todo el arreglo almenos que definas un index.
# + [markdown] tags=[]
# ### Operaciones matemáticas en el arrays
# -
# + [markdown] tags=[]
# #### Sumar todos los elementos en el array
# -
array_compra
array_compra.sum()
# + [markdown] tags=[]
# #### Multiplicar todos los elementos en el array
# -
array_compra.prod()
# + [markdown] tags=[]
# #### Calcular el promedio de los elementos en el array
# -
array_compra.mean()
# + [markdown] tags=[]
# #### Encontrar los valores mínimos y máximos en el array
# -
array_compra.min()
array_compra.max()
# + [markdown] tags=[]
# #### Encontrar los index donde están los valores mínimos y máximos en un array
# -
array_compra
array_compra.argmin()
array_compra.argmax()
# + [markdown] slideshow={"slide_type": "subslide"} tags=[]
# ### Calcular retornos aritmétricos diarios de un activo financiero
# + [markdown] tags=[]
# #### Gameplan:
# -
# $$R_n=\frac{P_{n+1} - P_{n}}{P_{n}}$$
#
# 1. Importar datos de internet
# 1. Calcular diferencia entre P$_{n+1}$ y P$_{n}$
# 1. Dividir diferenia entre P$_n$
# 1. Calcular media de los retornos
# 1. Calcular $\sigma$
# 1. Graficar retornos
# 1. Números de retornos (eje x)
# 1. Titulo de la gráfica
# 1. Leyendas ejes X's y Y's
# 1. Graficar línea retorno promedios
# + [markdown] tags=[]
# ##### 1. Importar datos utilizando la función `loadtxt( )`
# -
url = 'https://raw.githubusercontent.com/carlosdeoncedos/analisisdeinversion2021/main/Datos/T5PreciosSimulados.csv'
precios = np.loadtxt(url, usecols=[0], delimiter=',')
np.shape(precios)
# Argumentos de `loadtxt( )`
# https://numpy.org/doc/stable/reference/generated/numpy.loadtxt.html
# ##### 2. Calcular diferencia entre $P_{n+1}$ y $P_{n}$
diferencia = precios[1:] - precios[:-1]
np.shape(diferencia)
# ##### 3. Dividir diferenia entre P$_n$
retornos = diferencia / precios[:-1]
retornos
# ##### 4. Calcular media de los retornos
promedio = retornos.mean()
promedio
# 5. Calcular $\sigma$
sigma = retornos.std()
sigma
# 6. Graficar retornos
# + [markdown] tags=[]
# ###### 6.a Contar el número de días
# -
dias = len(retornos)
dias
dias = np.arange(1, len(retornos)+1)
dias
# + [markdown] tags=[]
# ###### 6.b y 6.c Generar gráfica, incluir titulo y leyendas en los ejes X y Y
# -
import matplotlib.pyplot as plt
fig, ax = plt.subplots(figsize=(12,6))
ax.plot(dias, retornos, c='navy')
ax.set_title('Retornos ARITMETRICOS diarios');
fig, ax = plt.subplots(figsize=(12,6))
ax.plot(dias, retornos, c='navy')
ax.set_title('Retornos ARITMETRICOS diarios')
ax.axhline(promedio, c='r', ls=':');
# ##### Generar línea horizontal con el promedio de los retornos
# + [markdown] tags=[]
# ### Comparando arrays
# -
a = np.array([10,5,0])
b = np.array([3,5,8])
a>b
a == b
b > 5
# + [markdown] tags=[]
# #### Comparar utilizando la función `logical_and`
# -
# > "*`logical_and` calcula el valor verdadero de $x_1$ y $x_2$* con cada elemento del array". https://numpy.org/doc/stable/reference/generated/numpy.logical_and.html
np.logical_and(a>4, a<6)
# + [markdown] tags=[]
# #### Comparar utilizando la función `logical_or`
# -
# > "*`logical_and` calcula el valor verdadero de $x_1$ y $x_2$* con cada elemento del array". https://numpy.org/doc/stable/reference/generated/numpy.logical_and.html
a
np.logical_or(a<4, a>6)
# + [markdown] tags=[]
# #### Una especie del IF de excel en NumPy: `where( )`
# -
# > Sintaxis: where(evaluación, $respuesta_1$ si es verdadero, $respuesta_2$ si es falso)
a = np.array([4,6,8])
np.where(a==6, 1, 0)
# + [markdown] tags=[]
# ### Identificar volumen transacción mayor a 80 mil millones
# -
# Pregunta: _Entre mayor volumen de transacción diaria es un indicativo de que el activo está en una alza o una baja_?
# + [markdown] tags=[]
# #### Gameplan:
# -
# 1. Cargar precios BTC/USD.
# 1. Generar _filtro_.
# 1. Encontrar días que el volumen de transacción es mayor al filtro.
# 1. Visualizar resultados.
# + [markdown] tags=[]
# ##### 1. Cargar precios BTC/USD
# -
url = 'https://raw.githubusercontent.com/carlosdeoncedos/analisisdeinversion2021/main/Datos/BTC-USD.csv'
btc = np.loadtxt(url, delimiter=',')
btc
APERTURA = 0
MAXIMO = 1
MINIMO = 2
CIERRE = 3
VOLUMEN = 4
# + [markdown] tags=[]
# ##### 2. Generar _filtro_
# -
percentil = np.percentile(btc[:,VOLUMEN], 98)
percentil
filtro = btc[:,VOLUMEN]>=percentil
filtro
True + True
False + True
btc_dias_alto_volumen = np.sum(filtro)
btc_dias_alto_volumen
# + [markdown] tags=[]
# ##### 3. Encontrar días que el volumen de transacción es mayor al filtro
# -
index_alto_volumen = np.where(filtro)[0]
index_alto_volumen
# + [markdown] tags=[]
# ##### 4. Visualizar resultados
# -
fig, ax = plt.subplots(figsize=(12,6))
ax.plot(btc[:,CIERRE], c='navy');
fig, ax = plt.subplots(figsize=(12,6))
ax.plot(btc[:,CIERRE], c='navy')
ax.plot(index_alto_volumen, btc[index_alto_volumen, CIERRE], 'ro');
# + [markdown] tags=[]
# ### Estadísticas
# + [markdown] tags=[]
# #### Promedio
# -
np.shape(btc)
btc.mean()
btc.mean(axis=0)
btc.mean(axis=1)
# #### Varianza
btc.var(axis=0)
# #### Desviación standard
btc.std(axis=0)
# #### Mediana
# + [markdown] tags=[]
# ### Números aleatorios
# -
np.random.rand()
# + [markdown] tags=[]
# #### Números aleatorios con semilla
# -
np.random.seed(13)
np.random.rand()
np.random.seed(2021)
np.random.rand(4)
np.random.rand(2,3)
np.random.randint(5,10)
# # FINAL DE LA CLASE 5 DE OCTUBRE, 2021
# HASTA LA CELDA ANTERIOR LLEGAMOS EN LA CLASE DEL 5 DE OCTUBRE
# # CONTINUACIóN DE LA CLASE:
import numpy as np
# + [markdown] slideshow={"slide_type": "subslide"} tags=[]
# ## Trabajando con arrays
# + [markdown] slideshow={"slide_type": "fragment"}
# 
# -
a = np.array([[0,1,2,3],[10,11,12,13]])
a
a.shape
# + [markdown] slideshow={"slide_type": "subslide"}
# 
# -
a.size
# + [markdown] slideshow={"slide_type": "subslide"}
# 
# -
a.ndim
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Leer un elemento de un array
# -
# 
a[0,2]
variable = a[0,2]
variable
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Cambiar un elemento de un array
# + [markdown] slideshow={"slide_type": "subslide"}
# 
# -
a[1,3]
a[1,3] = -1
a
# + [markdown] slideshow={"slide_type": "subslide"} tags=[]
# ### Slicing
# -
# $$array[LimiteInferior:LimiteSuperior:Pasos]$$
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Slicing en array de 1D
# -
a = np.array([13, 14, 15, 16, 17])
a
# Seleccionar del 14 al 16:
a[1:4]
a[::2]
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Slicing en array de 2D
# -
# $$array[LimInfFila:LimSupFila:Pasos,LimInfColumna:LimSupColumna:Pasos]$$
# + [markdown] slideshow={"slide_type": "subslide"} tags=[]
# ### Ejericio Construir array 2D
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src="https://www.dropbox.com/s/ll7zezl88m3k5bz/08_array_2d.png?raw=1" width="400">
# -
# ### Pasos crear array 2D
# 1. Generar array del 0 al 59
# 1. Convertir array a 2D 6 x 10
# 1. Slice 0 al 5
# #### Paso 1
a = np.arange(60)
a
# #### Paso 2
a = a.reshape(6,10)
a
# #### Paso 3
a = a[:,:6]
a
# + [markdown] slideshow={"slide_type": "subslide"} tags=[]
# #### <font color='orange'>Hacer un slice en fila naranja
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src="https://www.dropbox.com/s/ll7zezl88m3k5bz/08_array_2d.png?raw=1" width="400">
# -
a[0,3:5]
# + [markdown] slideshow={"slide_type": "subslide"} tags=[]
# #### <font color='red'>Hacer un slice solo en la columna roja
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src="https://www.dropbox.com/s/ll7zezl88m3k5bz/08_array_2d.png?raw=1" width="400">
# -
a[:,2]
# + [markdown] slideshow={"slide_type": "subslide"} tags=[]
# #### <font color=#2B65EC>Partir elementos en color azul
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src="https://www.dropbox.com/s/ll7zezl88m3k5bz/08_array_2d.png?raw=1" width="400">
# -
a[4:,4:]
# + [markdown] slideshow={"slide_type": "subslide"} tags=[]
# #### <font color=#0020C2>Partir elementos en los recuadros azules
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src="https://www.dropbox.com/s/ll7zezl88m3k5bz/08_array_2d.png?raw=1" width="400">
# -
a[2::2,::2]
# ## Calcular retornos de los 99 los activos
url = 'https://raw.githubusercontent.com/carlosdeoncedos/analisisdeinversion2021/main/Datos/T5PreciosSimulados.csv'
precios = np.loadtxt(url, delimiter=',')
precios
np.shape(precios)
precios[:,14]
# +
# diferencia = precios[1:] - precio[:-1]
# retorno = diferencia / precio[:-1]
diferencia = precios[1:,:] - precios[:-1,:]
retornos = diferencia / precios[:-1,:]
# -
retornos
retornos.max(axis=0)
# + [markdown] slideshow={"slide_type": "subslide"} tags=[]
# ## NumPy Vs Python
# -
# %time lista_python = list(range(1000000000))
# %time arreglo_numpy = np.arange(1000000000)
# + [markdown] slideshow={"slide_type": "subslide"}
# Todas las imagenes son bajadas de internet
| Codigo/20211005Clase5.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.4.2
# language: julia
# name: julia-1.4
# ---
# # Holm-Hone equation
include("setup.jl")
# # Periodic boundary conditions
# +
function relaxation_functional(γ, unew, uold, param)
@unpack D, D2a, D4a, tmp1, tmp2, tmp3 = param
@. tmp3 = (1-γ)*uold + γ*unew
mul!(tmp2, D2a, tmp3)
mul!(tmp1, D4a, tmp3)
@. tmp1 = tmp3 * (4*tmp3 - 5*tmp2 + tmp1)
energy = integrate(tmp1, D)
end
function relaxation_functional(u, param)
@unpack D, D2a, D4a, tmp1, tmp2 = param
mul!(tmp2, D2a, u)
mul!(tmp1, D4a, u)
@. tmp1 = u * (4*u - 5*tmp2 + tmp1)
energy = integrate(tmp1, D)
end
function relaxation!(integrator)
told = integrator.tprev
uold = integrator.uprev
tnew = integrator.t
unew = integrator.u
γ = one(tnew)
terminate_integration = false
γlo = one(γ)/2
γhi = 3*one(γ)/2
energy_old = relaxation_functional(uold, integrator.p)
if (relaxation_functional(γlo, unew, uold, integrator.p)-energy_old) * (relaxation_functional(γhi, unew, uold, integrator.p)-energy_old) > 0
terminate_integration = true
else
γ = find_zero(g -> relaxation_functional(g, unew, uold, integrator.p)-energy_old, (γlo, γhi), Roots.AlefeldPotraShi())
end
if γ < eps(typeof(γ))
terminate_integration = true
end
@. unew = uold + γ * (unew - uold)
DiffEqBase.set_u!(integrator, unew)
if !(tnew ≈ top(integrator.opts.tstops))
tγ = told + γ * (tnew - told)
DiffEqBase.set_t!(integrator, tγ)
end
if terminate_integration
terminate!(integrator)
end
nothing
end
function save_func_hh_periodic(u, t, integrator)
@unpack D, D2a, D4a, tmp1, tmp2, tmp3, usol = integrator.p
mass = integrate(u, D)
mul!(tmp2, D2a, u)
mul!(tmp3, D4a, u)
@. tmp1 = 4*u - 5*tmp2 + tmp3
linear = integrate(tmp1, D)
@. tmp1 = u * tmp1
quadratic = integrate(tmp1, D)
SVector(mass, linear, quadratic)
end
function hh_periodic!(du, u, param, t)
@unpack D, inv4m5D2pD4, D2b, D4b, tmp1, tmp2, tmp3 = param
# conservative semidiscretization
mul!(tmp1, D2b, u)
mul!(tmp2, D4b, u)
@. tmp1 = -(4 * u - 5 * tmp1 + tmp2)
@. tmp2 = u * tmp1
mul!(tmp3, D, tmp2)
mul!(tmp2, D, u)
@. tmp3 = tmp3 + tmp2 * tmp1
ldiv!(du, inv4m5D2pD4, tmp3)
nothing
end
function solve_ode_hh_periodic(usol, D, D2a, D2b, D4a, D4b, tspan, alg, tol, dt, adaptive)
# lu instead of fatorize because of https://github.com/JuliaLang/julia/issues/30084
inv4m5D2pD4 = isa(D2a, AbstractMatrix) ? lu(4*I - 5*D2a + D4a) : 4*I - 5*D2a + D4a
x = grid(D)
u0 = usol.(tspan[1], x, x[1], -x[1])
tmp1 = similar(u0); tmp2 = similar(tmp1); tmp3 = similar(tmp1)
param = (D=D, D2a=D2a, D2b=D2b, D4a=D4a, D4b=D4b, inv4m5D2pD4=inv4m5D2pD4,
tmp1=tmp1, tmp2=tmp2, tmp3=tmp3, usol=usol)
ode = ODEProblem(hh_periodic!, u0, tspan, param)
saveat = range(tspan..., length=100)
saved_values_baseline = SavedValues(eltype(D), SVector{3,eltype(D)})
saving_baseline = SavingCallback(save_func_hh_periodic, saved_values_baseline, saveat=saveat)
saved_values_relaxation = SavedValues(eltype(D), SVector{3,eltype(D)})
saving_relaxation = SavingCallback(save_func_hh_periodic, saved_values_relaxation, saveat=saveat)
relaxation = DiscreteCallback((u,t,integrator) -> true, relaxation!, save_positions=(false,true))
cb_baseline = CallbackSet(saving_baseline)
cb_relaxation = CallbackSet(relaxation, saving_relaxation)
@time sol_relaxation = solve(ode, alg, abstol=tol, reltol=tol, dt=dt, adaptive=adaptive, save_everystep=false,
callback=cb_relaxation, tstops=saveat)
@time sol_baseline = solve(ode, alg, abstol=tol, reltol=tol, dt=dt, adaptive=adaptive, save_everystep=false,
callback=cb_baseline, tstops=saveat)
unum_baseline = sol_baseline[end]
unum_relaxation = sol_relaxation[end]
uana = usol.(tspan[end], x, x[1], -x[1])
@printf("Error in u (baseline): %.3e\n", integrate(u->u^2, unum_baseline - uana, D) |> sqrt)
@printf("Error in u (relaxation): %.3e\n", integrate(u->u^2, unum_relaxation - uana, D) |> sqrt)
@printf("Difference of baseline and relaxation in u: %.3e\n",
integrate(u->u^2, unum_baseline - unum_relaxation, D) |> sqrt)
sleep(0.1)
fig_u, ax = plt.subplots(1, 1)
plt.plot(x, u0, label=L"u^0")
plt.plot(x, uana, label=L"$u^\mathrm{ana}$")
plt.plot(x, unum_baseline, label=L"$u^\mathrm{num}$ (baseline)")
plt.plot(x, unum_relaxation, label=L"$u^\mathrm{num}$ (relaxation)")
plt.xlabel(L"x"); plt.ylabel(L"u")
plt.legend(loc="center left", bbox_to_anchor=(1.0, 0.5));
t_baseline = saved_values_baseline.t
t_relaxation = saved_values_relaxation.t
mass_baseline = map(x->x[1], saved_values_baseline.saveval)
mass_relaxation = map(x->x[1], saved_values_relaxation.saveval)
linear_baseline = map(x->x[2], saved_values_baseline.saveval)
linear_relaxation = map(x->x[2], saved_values_relaxation.saveval)
quadratic_baseline = map(x->x[3], saved_values_baseline.saveval)
quadratic_relaxation = map(x->x[3], saved_values_relaxation.saveval)
fig_invariants, ax = plt.subplots(1, 1)
ax.set_yscale("symlog", linthreshy=1.0e-14)
plt.plot(t_baseline, mass_baseline .- mass_baseline[1], label=L"$\int u$ (baseline)")
plt.plot(t_relaxation, mass_relaxation .- mass_relaxation[1], label=L"$\int u$ (relaxation)")
plt.plot(t_baseline, linear_baseline .- linear_baseline[1], label=L"$\int (4 - 5 \partial_x^2 + \partial_x^4) u$ (baseline)")
plt.plot(t_relaxation, linear_relaxation .- linear_relaxation[1], label=L"$\int (4 - 5 \partial_x^2 + \partial_x^4) u$ (relaxation)")
plt.plot(t_baseline, quadratic_baseline .- quadratic_baseline[1], label=L"$\int u (4 - 5 \partial_x^2 + \partial_x^4) u$ (baseline)")
plt.plot(t_relaxation, quadratic_relaxation .- quadratic_relaxation[1], label=L"$\int u (4 - 5 \partial_x^2 + \partial_x^4) u$ (relaxation)")
plt.xlabel(L"t"); plt.ylabel("Change of Invariants")
plt.legend(loc="center left", bbox_to_anchor=(1.0, 0.5))
fig_u, fig_invariants
end
# +
# # single peakon solution
# xmin = -40.
# xmax = -xmin
# get_c() = 1.2
# function usol(t, x, xmin, xmax)
# c = get_c()
# x_t = mod(x - c*t - xmin, xmax - xmin) + xmin
# c * (2 * exp(-abs(x_t)) - exp(-2*abs(x_t)))
# end
# traveling wave solution obtained numerically
c, data = open("hh_traveling_wave_init_c12_ah01_l40_N65536.txt", "r") do io
line = readline(io)
line = readline(io)
line = readline(io)
c = parse(Float64, line[8:end])
data = readdlm(io, comments=true)
c, data
end
x = range(data[1,1], data[end,1], length=size(data,1))
xmin = x[1]; xmax = x[end]
u0 = data[:, 2]
u0itp = CubicSplineInterpolation((x,), u0, extrapolation_bc=Periodic())
get_c() = c
function usol(t, x, xmin, xmax)
c = get_c()
x_t = mod(x - c*t - xmin, xmax - xmin) + xmin
u0itp(x_t)
end
println("c = ", get_c())
println("xmin = ", xmin)
println("xmax = ", xmax)
@show usol(0., xmin, xmin, xmax)
@show usol(0., xmax, xmin, xmax)
@show N = 2^8
@show dt = 0.5 * (xmax - xmin) / (N * get_c())
# @show tspan = (0., (xmax-xmin)/(3*get_c()))
@show tspan = (0.0, (xmax-xmin)/(3*get_c()) + 1*(xmax-xmin)/get_c())
# @show tspan = (0.0, (xmax-xmin)/(3*get_c()) + 10*(xmax-xmin)/get_c())
# @show tspan = (0.0, (xmax-xmin)/(3*get_c()) + 100*(xmax-xmin)/get_c())
# @show tspan = (0.0, (xmax-xmin)/(3*get_c()) + 1000*(xmax-xmin)/get_c())
flush(stdout)
tol = 1.0e-7
adaptive = false
# D = fourier_derivative_operator(xmin, xmax, N)
# D2a = D2b = D^2; D4a = D4b = D^4
# D = periodic_derivative_operator(1, 6, xmin, xmax, N+1)
# D = periodic_derivative_operator(Holoborodko2008(), 1, 4, xmin, xmax, N+1)
# D2a = D2b = sparse(periodic_derivative_operator(2, 6, xmin, xmax, N+1))
# D4a = D4b = sparse(periodic_derivative_operator(4, 6, xmin, xmax, N+1))
# D2a = sparse(D)^2
# D2a = sparse(periodic_derivative_operator(2, 6, xmin, xmax, N+1))
# D2b = D^2
# D2b = periodic_derivative_operator(2, 6, xmin, xmax, N+1)
# D4a = sparse(D)^4
# D4a = sparse(periodic_derivative_operator(4, 6, xmin, xmax, N+1))
# D4b = D^4
# D4b = periodic_derivative_operator(4, 6, xmin, xmax, N+1)
p = 8; mesh = UniformPeriodicMesh1D(xmin, xmax, N÷p)
D = couple_continuosly(legendre_derivative_operator(-1., 1., p+1), mesh)
D2a = sparse(couple_continuosly(legendre_second_derivative_operator(-1., 1., p+1), mesh))
# D2a = sparse(D)^2
D2b = sparse(couple_continuosly(legendre_second_derivative_operator(-1., 1., p+1), mesh))
# D2b = sparse(D)^2
D4a = D2a^2; D4b = D2b^2
p = 7; Dop = legendre_derivative_operator(-1., 1., p+1); mesh = UniformPeriodicMesh1D(xmin, xmax, N÷(p+1))
D = couple_discontinuosly(Dop, mesh); D₊ = couple_discontinuosly(Dop, mesh, Val(:plus)); D₋ = couple_discontinuosly(Dop, mesh, Val(:minus))
# D2a = sparse(D₊) * sparse(D₋)
# D2a = sparse(D₋) * sparse(D₊)
D2a = sparse(D)^2
D2b = sparse(D)^2
D4a = D2a^2; D4b = D2b^2
fig_u, fig_invariants = solve_ode_hh_periodic(
usol, D, D2a, D2b, D4a, D4b, tspan, RK4(), tol, dt, adaptive)
# -
# # Convergence study with manufactured solutions
# +
import SymPy; sp = SymPy
function math_replacements(s)
s = replace(s, "cos(pi*" => "cospi(")
s = replace(s, "sin(pi*" => "sinpi(")
end
function usol(t, x)
exp(t/2) * sinpi(2*(x-t/2))
end
let (t, x) = sp.symbols("t, x", real=true)
u = usol(t, x)
println("u:")
4*sp.diff(u, t) - 5*sp.diff(u, x, 2, t, 1) + sp.diff(u, x, 4, t, 1) + u*sp.diff(u, x, 5) + 2*sp.diff(u, x)*sp.diff(u, x, 4) - 5*u*sp.diff(u, x, 3) - 10*sp.diff(u, x)*sp.diff(u, x, 2) + 12*u*sp.diff(u, x) |>
sp.simplify |> sp.string |> math_replacements |> println
end
# +
function save_func_hh_manufactured(u, t, integrator)
@unpack D, x, tmp1, usol = integrator.p
tmp1 .= ( usol.(t, x) .- u ).^2
error_u = integrate(tmp1, D) |> sqrt
error_u
end
function hh_periodic_manufactured!(du, u, param, t)
@unpack D, x, inv4m5D2pD4, D2b, D4b, tmp1, tmp2, tmp3 = param
mul!(tmp1, D2b, u)
mul!(tmp2, D4b, u)
@. tmp1 = -(4 * u - 5 * tmp1 + tmp2)
@. tmp2 = u * tmp1
mul!(tmp3, D, tmp2)
mul!(tmp2, D, u)
@. tmp3 = tmp3 + tmp2 * tmp1
@. tmp3 += -8*pi^4*(sinpi((t - 2*x)) + 2*pi*cospi((t - 2*x)))*exp(t/2) - 10*pi^2*(sinpi((t - 2*x)) + 2*pi*cospi((t - 2*x)))*exp(t/2) - 2*exp(t/2)*sinpi((t - 2*x)) - 4*pi*exp(t/2)*cospi((t - 2*x)) - 48*pi^5*exp(t)*sinpi((2*t - 4*x)) - 60*pi^3*exp(t)*sinpi((2*t - 4*x)) - 12*pi*exp(t)*sinpi((2*t - 4*x))
ldiv!(du, inv4m5D2pD4, tmp3)
nothing
end
function errors_hh_periodic_manufactured(usol, D, D2a, D2b, D4a, D4b, tspan, alg, tol, dt, adaptive)
inv4m5D2pD4 = isa(D2a, AbstractMatrix) ? lu(4*I - 5*D2a + D4a) : 4*I - 5*D2a + D4a
x = collect(grid(D))
u0 = usol.(tspan[1], x)
tmp1 = similar(u0); tmp2 = similar(tmp1); tmp3 = similar(tmp1)
param = (D=D, D2a=D2a, D2b=D2b, D4a=D4a, D4b=D4b, inv4m5D2pD4=inv4m5D2pD4,
x=x, tmp1=tmp1, tmp2=tmp2, tmp3=tmp3, usol=usol)
ode = ODEProblem(hh_periodic_manufactured!, u0, tspan, param)
saveat = range(tspan..., length=2)
saved_values_baseline = SavedValues(eltype(D), eltype(D))
saving_baseline = SavingCallback(save_func_hh_manufactured, saved_values_baseline, saveat=saveat)
cb_baseline = CallbackSet(saving_baseline)
sol_baseline = solve(ode, alg, abstol=tol, reltol=tol, dt=dt, adaptive=adaptive, save_everystep=false,
callback=cb_baseline, tstops=saveat)
error_u = saved_values_baseline.saveval[end]
end
xmin = 0.0
xmax = 1.0
tspan = (0.0, 1.0)
tol = 1.0e-6
# val_N = round.(Int, 2 .^ range(4, 5, length=2))
# val_N = round.(Int, 2 .^ range(4, 8, length=7)) |> evenodd_values
# val_N = round.(Int, 2 .^ range(3, 4.5, length=7)) |> evenodd_values
val_N = [2^2, 2^3, 2^4, 2^5]
# val_N = 2 .^ (2:7) .+ 1
val_error = Float64[]
for N in val_N
# D = fourier_derivative_operator(xmin, xmax, N)
# p = 6; D = periodic_derivative_operator(1, p, xmin, xmax, N+1)
# p = 4; D = periodic_derivative_operator(Holoborodko2008(), 1, p, xmin, xmax, N+1)
# D2 = D^2
# D2 = periodic_derivative_operator(2, p, xmin, xmax, N+1)
# D2 = periodic_derivative_operator(Holoborodko2008(), 1, p, xmin, xmax, N+1)^2
# D4 = periodic_derivative_operator(4, p, xmin, xmax, N+1)
p = 2; mesh = UniformPeriodicMesh1D(xmin, xmax, N)
D = couple_continuosly(legendre_derivative_operator(-1., 1., p+1), mesh)
D2 = sparse(couple_continuosly(legendre_second_derivative_operator(-1., 1., p+1), mesh))
# D2 = sparse(D)^2
p = 2; Dop = legendre_derivative_operator(-1., 1., p+1); mesh = UniformPeriodicMesh1D(xmin, xmax, N)
D = couple_discontinuosly(Dop, mesh); D₊ = couple_discontinuosly(Dop, mesh, Val(:plus)); D₋ = couple_discontinuosly(Dop, mesh, Val(:minus))
# D2 = sparse(D₊) * sparse(D₋)
# D2 = sparse(D₋) * sparse(D₊)
D2 = sparse(D)^2
D4 = D2^2
error_u = errors_hh_periodic_manufactured(usol, D, sparse(D2), D2, sparse(D4), D4, tspan, Tsit5(), tol, 1/N, true)
push!(val_error, error_u)
end
@show val_error
fig, ax = plt.subplots(1, 1)
ax.set_prop_cycle(marker_cycler)
plt.plot(val_N, val_error, label=L"$\| u - u_{\mathrm{ana}} \|$")
c0, c1 = linear_regression(log.(val_N), log.(val_error))
plt.plot(val_N, exp(c0) .* val_N.^c1, marker="", linestyle=":", color="gray", label=@sprintf("Order %.2f", -c1))
plt.xscale("log", basex=2)
plt.yscale("symlog", linthreshy=1.0e-12)
plt.legend(loc="center left", bbox_to_anchor=(1.0, 0.5))
ax.locator_params(axis="y", numticks=9)
# +
N = 2^6
p = 2
mesh = UniformPeriodicMesh1D(xmin, xmax, N)
D = couple_continuosly(legendre_derivative_operator(-1., 1., p+1), mesh)
# D2 = sparse(couple_continuosly(legendre_second_derivative_operator(-1., 1., p+1), mesh))
D2 = sparse(D)^2
# Dop = legendre_derivative_operator(-1., 1., p+1)
# D = couple_discontinuosly(Dop, mesh); D₊ = couple_discontinuosly(Dop, mesh, Val(:plus)); D₋ = couple_discontinuosly(Dop, mesh, Val(:minus))
# # D2 = sparse(D₊) * sparse(D₋)
# # D2 = sparse(D₋) * sparse(D₊)
# D2 = sparse(D)^2
D4 = D2^2
cond(Matrix(4*I - 5*D2 + D4))
# -
# ## Convergence study: Plots FD
# +
function do_stuff(p, val_N)
val_error = Float64[]
@show p
for N in val_N
@show N
flush(stdout)
D = periodic_derivative_operator(1, p, xmin, xmax, N+1)
# D2 = D^2
D2 = periodic_derivative_operator(2, p, xmin, xmax, N+1)
# D4 = D2^2
D4 = periodic_derivative_operator(4, p, xmin, xmax, N+1)
error_u = errors_hh_periodic_manufactured(usol, D, sparse(D2), D2, sparse(D4), D4, tspan, Tsit5(), tol, 1/N, true)
push!(val_error, error_u)
end
Dict("val_p$(p)_N" => val_N, "val_p$(p)_error" => val_error)
end
filename = "hh_periodic_manufactured_convergence_FD_narrow"
even_odd_values = evenodd_values
res = Dict{String,Any}()
p = 2
val_N = round.(Int, 2 .^ range(4, 10, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 4
val_N = round.(Int, 2 .^ range(4, 8.5, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 6
val_N = round.(Int, 2 .^ range(4, 7, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 8
val_N = round.(Int, 2 .^ range(4, 6.5, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
open("../data/" * filename * ".json", "w") do io
JSON.print(io, res, 2)
end
# +
fig, ax = plt.subplots(1, 1)
ax.set_prop_cycle(marker_cycler)
filename = "hh_periodic_manufactured_convergence_FD_narrow"
res = JSON.parsefile("../data/" * filename * ".json")
let p = 2
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("\$ p = %d \$, EOC %.2f", p, -c1), (val_N[4], 2.0*val_error_u[4]), color="gray")
end
let p = 4
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("\$ p = %d \$, EOC %.2f", p, -c1), (val_N[6], 2.0*val_error_u[6]), color="gray")
end
let p = 6
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("\$ p = %d \$, EOC %.2f", p, -c1), (0.6*val_N[end], 0.1*val_error_u[end]), color="gray")
end
let p = 8
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("\$ p = %d \$, EOC %.2f", p, -c1), (0.15*val_N[end], val_error_u[end]), color="gray")
end
# ax.set_ylim(1.0e-10, 1.5e0)
ax.set_xscale("log", basex=2)
ax.set_yscale("symlog", linthreshy=1.0e-13)
ax.set_xlabel(L"N")
ax.set_ylabel(L"\| u - u_{\mathrm{ana}} \|")
fig.savefig("../figures/" * filename * ".pdf", bbox_inches="tight")
# +
function do_stuff(p, val_N)
val_error = Float64[]
@show p
for N in val_N
@show N
flush(stdout)
D = periodic_derivative_operator(1, p, xmin, xmax, N+1)
# D2 = D^2
D2 = periodic_derivative_operator(2, p, xmin, xmax, N+1)
D4 = D2^2
# D4 = D^4
# D4 = periodic_derivative_operator(4, p, xmin, xmax, N+1)
error_u = errors_hh_periodic_manufactured(usol, D, sparse(D2), D2, sparse(D4), D4, tspan, Tsit5(), tol, 1/N, true)
push!(val_error, error_u)
end
Dict("val_p$(p)_N" => val_N, "val_p$(p)_error" => val_error)
end
filename = "hh_periodic_manufactured_convergence_FD_narrow_wide"
even_odd_values = evenodd_values
res = Dict{String,Any}()
p = 2
val_N = round.(Int, 2 .^ range(5.5, 8.5, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 4
val_N = round.(Int, 2 .^ range(5.5, 8, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 6
val_N = round.(Int, 2 .^ range(5, 6.4, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 8
val_N = round.(Int, 2 .^ range(4.5, 5.8, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
open("../data/" * filename * ".json", "w") do io
JSON.print(io, res, 2)
end
# +
fig, ax = plt.subplots(1, 1)
ax.set_prop_cycle(marker_cycler)
filename = "hh_periodic_manufactured_convergence_FD_narrow_wide"
res = JSON.parsefile("../data/" * filename * ".json")
let p = 2
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("\$ p = %d \$, EOC %.2f", p, -c1), (val_N[4], 2.0*val_error_u[4]), color="gray")
end
let p = 4
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("\$ p = %d \$, EOC %.2f", p, -c1), (val_N[6], 2.0*val_error_u[6]), color="gray")
end
let p = 6
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("\$ p = %d \$, EOC %.2f", p, -c1), (1.05val_N[end], 0.8*val_error_u[end]), color="gray")
end
let p = 8
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("\$ p = %d \$, EOC %.2f", p, -c1), (1.05*val_N[end], val_error_u[end]), color="gray")
end
ax.set_ylim(1.0e-10, 1.5e-1)
ax.set_xscale("log", basex=2)
ax.set_yscale("symlog", linthreshy=1.0e-13)
ax.set_xlabel(L"N")
ax.set_ylabel(L"\| u - u_{\mathrm{ana}} \|")
fig.savefig("../figures/" * filename * ".pdf", bbox_inches="tight")
# +
function do_stuff(p, val_N)
val_error = Float64[]
@show p
for N in val_N
@show N
flush(stdout)
D = periodic_derivative_operator(1, p, xmin, xmax, N+1)
D2 = D^2
# D2 = periodic_derivative_operator(2, p, xmin, xmax, N+1)
D4 = D^4
# D4 = periodic_derivative_operator(4, p, xmin, xmax, N+1)
error_u = errors_hh_periodic_manufactured(usol, D, sparse(D2), D2, sparse(D4), D4, tspan, Tsit5(), tol, 1/N, true)
push!(val_error, error_u)
end
Dict("val_p$(p)_N" => val_N, "val_p$(p)_error" => val_error)
end
filename = "hh_periodic_manufactured_convergence_FD_wide"
even_odd_values = evenodd_values
res = Dict{String,Any}()
p = 2
val_N = round.(Int, 2 .^ range(5.5, 8.5, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 4
val_N = round.(Int, 2 .^ range(5.5, 8, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 6
val_N = round.(Int, 2 .^ range(5, 7, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 8
val_N = round.(Int, 2 .^ range(4.5, 6, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
open("../data/" * filename * ".json", "w") do io
JSON.print(io, res, 2)
end
# +
fig, ax = plt.subplots(1, 1)
ax.set_prop_cycle(marker_cycler)
filename = "hh_periodic_manufactured_convergence_FD_wide"
res = JSON.parsefile("../data/" * filename * ".json")
let p = 2
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N[3:end]), log.(val_error_u[3:end]))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("\$ p = %d \$, EOC %.2f", p, -c1), (val_N[4], 2.0*val_error_u[4]), color="gray")
end
let p = 4
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N[3:end]), log.(val_error_u[3:end]))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("\$ p = %d \$, EOC %.2f", p, -c1), (val_N[6], 2.0*val_error_u[6]), color="gray")
end
let p = 6
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N[2:2:end]), log.(val_error_u[2:2:end]))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("\$ p = %d \$, EOC %.2f", p, -c1), (val_N[end], 0.3*val_error_u[end]), color="gray")
end
let p = 8
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("\$ p = %d \$, EOC %.2f", p, -c1), (0.3*val_N[end], val_error_u[end]), color="gray")
end
ax.set_ylim(1.0e-10, 1.5e-1)
ax.set_xscale("log", basex=2)
ax.set_yscale("symlog", linthreshy=1.0e-13)
ax.set_xlabel(L"N")
ax.set_ylabel(L"\| u - u_{\mathrm{ana}} \|")
fig.savefig("../figures/" * filename * ".pdf", bbox_inches="tight")
# -
# ## Convergence study: Plots CG
# +
function do_stuff(p, val_N)
val_error = Float64[]
@show p
for N in val_N
@show N
mesh = UniformPeriodicMesh1D(xmin, xmax, N)
D = couple_continuosly(legendre_derivative_operator(-1., 1., p+1), mesh)
D2 = sparse(D)^2
# D2 = sparse(couple_continuosly(legendre_second_derivative_operator(-1., 1., p+1), mesh))
D4 = D2^2
error_u = errors_hh_periodic_manufactured(usol, D, D2, D2, D4, D4, tspan, Tsit5(), tol, 1/N, true)
push!(val_error, error_u)
end
Dict("val_p$(p)_N" => val_N, "val_p$(p)_error" => val_error)
end
filename = "hh_periodic_manufactured_convergence_CG_D1c2"
even_odd_values = evenodd_values
res = Dict{String,Any}()
p = 1
val_N = round.(Int, 2 .^ range(5.5, 8.5, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 2
val_N = round.(Int, 2 .^ range(5.5, 7.25, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 3
val_N = round.(Int, 2 .^ range(2, 6.5, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 4
val_N = round.(Int, 2 .^ range(2, 6, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 5
val_N = round.(Int, 2 .^ range(2, 5.5, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 6
val_N = round.(Int, 2 .^ range(2, 5, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
open("../data/" * filename * ".json", "w") do io
JSON.print(io, res, 2)
end
# +
fig, ax = plt.subplots(1, 1)
ax.set_prop_cycle(marker_cycler)
filename = "hh_periodic_manufactured_convergence_CG_D1c2"
res = JSON.parsefile("../data/" * filename * ".json")
let p = 1
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("\$ p = %d \$, EOC %.2f", p, -c1), (val_N[3], 2.0*val_error_u[3]), color="gray")
end
let p = 2
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("\$ p = %d \$, EOC %.2f", p, -c1), (0.6*val_N[end], 0.2*val_error_u[end]), color="gray")
end
let p = 3
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("\$ p = %d \$, EOC %.2f", p, -c1), (val_N[end], 1.5*val_error_u[end]), color="gray")
end
let p = 4
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("\$ p = %d \$, EOC %.2f", p, -c1), (1.2*val_N[end], 0.5*val_error_u[end]), color="gray")
end
# let p = 5
# val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
# ax.plot(val_N, val_error_u, label="\$ p = $p \$")
# c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
# plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
# plt.annotate(@sprintf("EOC %.2f", -c1), (1.2*val_N[end], 0.2*val_error_u[end]), color="gray")
# end
# let p = 6
# val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
# ax.plot(val_N, val_error_u, label="\$ p = $p \$")
# c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
# plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
# plt.annotate(@sprintf("EOC %.2f", -c1), (0.2*val_N[end], 0.1*val_error_u[end]), color="gray")
# end
# ax.set_xlim(2^2-0.5, 2^13-1)
ax.set_ylim(1.0e-10, 1.5e-1)
ax.set_xscale("log", basex=2)
ax.set_yscale("symlog", linthreshy=1.0e-13)
ax.set_xlabel(L"N")
ax.set_ylabel(L"\| u - u_{\mathrm{ana}} \|")
fig.savefig("../figures/" * filename * ".pdf", bbox_inches="tight")
# plt.figure()
# handles, labels = ax.get_legend_handles_labels()
# plt.figlegend(handles, labels, loc="center", ncol=8)
# plt.savefig("../figures/Galerkin_legend_p1_p4.pdf", bbox_inches="tight")
# +
function do_stuff(p, val_N)
val_error = Float64[]
@show p
for N in val_N
@show N
mesh = UniformPeriodicMesh1D(xmin, xmax, N)
D = couple_continuosly(legendre_derivative_operator(-1., 1., p+1), mesh)
# D2 = sparse(D)^2
D2 = sparse(couple_continuosly(legendre_second_derivative_operator(-1., 1., p+1), mesh))
D4 = D2^2
error_u = errors_hh_periodic_manufactured(usol, D, D2, D2, D4, D4, tspan, Tsit5(), tol, 1/N, true)
push!(val_error, error_u)
end
Dict("val_p$(p)_N" => val_N, "val_p$(p)_error" => val_error)
end
filename = "hh_periodic_manufactured_convergence_CG_narrow"
even_odd_values = evenodd_values
res = Dict{String,Any}()
p = 1
val_N = round.(Int, 2 .^ range(5.5, 8.5, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 2
val_N = round.(Int, 2 .^ range(5.5, 7.25, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 3
val_N = round.(Int, 2 .^ range(2, 6.5, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 4
val_N = round.(Int, 2 .^ range(2, 6, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 5
val_N = round.(Int, 2 .^ range(2, 5.5, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 6
val_N = round.(Int, 2 .^ range(2, 5, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
open("../data/" * filename * ".json", "w") do io
JSON.print(io, res, 2)
end
# +
fig, ax = plt.subplots(1, 1)
ax.set_prop_cycle(marker_cycler)
filename = "hh_periodic_manufactured_convergence_CG_narrow"
res = JSON.parsefile("../data/" * filename * ".json")
let p = 1
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("EOC %.2f", -c1), (0.45*val_N[end], 0.2*val_error_u[end]), color="gray")
end
let p = 2
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("EOC %.2f", -c1), (val_N[2], 2.0*val_error_u[2]), color="gray")
end
let p = 3
val_N, val_error_u = res["val_p$(p)_N"][1:end-2], res["val_p$(p)_error"][1:end-2]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("EOC %.2f", -c1), (val_N[end], 1.5*val_error_u[end]), color="gray")
end
let p = 4
val_N, val_error_u = res["val_p$(p)_N"][1:end-2], res["val_p$(p)_error"][1:end-2]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("EOC %.2f", -c1), (1.2*val_N[end], 0.5*val_error_u[end]), color="gray")
end
# let p = 5
# val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
# ax.plot(val_N, val_error_u, label="\$ p = $p \$")
# c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
# plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
# plt.annotate(@sprintf("EOC %.2f", -c1), (1.2*val_N[end], 0.2*val_error_u[end]), color="gray")
# end
# let p = 6
# val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
# ax.plot(val_N, val_error_u, label="\$ p = $p \$")
# c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
# plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
# plt.annotate(@sprintf("EOC %.2f", -c1), (0.2*val_N[end], 0.1*val_error_u[end]), color="gray")
# end
# ax.set_xlim(2^2-0.5, 2^13-1)
# ax.set_ylim(1.0e-13, 1.5e0)
ax.set_xscale("log", basex=2)
ax.set_yscale("symlog", linthreshy=1.0e-13)
ax.set_xlabel(L"N")
ax.set_ylabel(L"\| u - u_{\mathrm{ana}} \|")
fig.savefig("../figures/" * filename * ".pdf", bbox_inches="tight")
# -
# ## Convergence study: Plots DG
# +
function do_stuff(p, val_N)
val_error = Float64[]
@show p
for N in val_N
@show N
mesh = UniformPeriodicMesh1D(xmin, xmax, N)
Dop = legendre_derivative_operator(-1., 1., p+1)
D = couple_discontinuosly(Dop, mesh); D₊ = couple_discontinuosly(Dop, mesh, Val(:plus)); D₋ = couple_discontinuosly(Dop, mesh, Val(:minus))
D2 = sparse(D)^2
# D2 = sparse(D₊) * sparse(D₋)
# D2 = sparse(D₋) * sparse(D₊)
D4 = D2^2
error_u = errors_hh_periodic_manufactured(usol, D, D2, D2, D4, D4, tspan, Tsit5(), 1.0e-6, 1/N, true)
push!(val_error, error_u)
end
Dict("val_p$(p)_N" => val_N, "val_p$(p)_error" => val_error)
end
filename = "hh_periodic_manufactured_convergence_DG_D1c2"
even_odd_values = evenodd_values
res = Dict{String,Any}()
p = 1
val_N = round.(Int, 2 .^ range(6, 8, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 2
val_N = round.(Int, 2 .^ range(6, 8, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 3
val_N = round.(Int, 2 .^ range(5, 7, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 4
val_N = round.(Int, 2 .^ range(2, 6, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
# p = 5
# val_N = round.(Int, 2 .^ range(2, 5.5, length=7)) |> even_odd_values
# res = merge(res, do_stuff(p, val_N))
# p = 6
# val_N = round.(Int, 2 .^ range(2, 5, length=7)) |> even_odd_values
# res = merge(res, do_stuff(p, val_N))
open("../data/" * filename * ".json", "w") do io
JSON.print(io, res, 2)
end
# +
fig, ax = plt.subplots(1, 1)
ax.set_prop_cycle(marker_cycler)
filename = "hh_periodic_manufactured_convergence_DG_D1c2"
res = JSON.parsefile("../data/" * filename * ".json")
let p = 1
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("EOC %.2f", -c1), (val_N[2], val_error_u[2]), color="gray")
end
let p = 2
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("EOC %.2f", -c1), (val_N[end], val_error_u[end]), color="gray")
end
let p = 3
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("EOC %.2f", -c1), (val_N[end], val_error_u[end]), color="gray")
end
let p = 4
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("EOC %.2f", -c1), (1.2*val_N[end], val_error_u[end]), color="gray")
end
# let p = 5
# val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
# ax.plot(val_N, val_error_u, label="\$ p = $p \$")
# c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
# plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
# plt.annotate(@sprintf("EOC %.2f", -c1), (1.2*val_N[end], 0.2*val_error_u[end]), color="gray")
# end
# let p = 6
# val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
# ax.plot(val_N, val_error_u, label="\$ p = $p \$")
# c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
# plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
# plt.annotate(@sprintf("EOC %.2f", -c1), (0.2*val_N[end], 0.1*val_error_u[end]), color="gray")
# end
# ax.set_xlim(2^2-0.5, 2^13-1)
# ax.set_ylim(1.0e-13, 1.5e0)
ax.set_xscale("log", basex=2)
ax.set_yscale("symlog", linthreshy=1.0e-13)
ax.set_xlabel(L"N")
ax.set_ylabel(L"\| u - u_{\mathrm{ana}} \|")
fig.savefig("../figures/" * filename * ".pdf", bbox_inches="tight")
# +
function do_stuff(p, val_N)
val_error = Float64[]
@show p
for N in val_N
@show N
mesh = UniformPeriodicMesh1D(xmin, xmax, N)
Dop = legendre_derivative_operator(-1., 1., p+1)
D = couple_discontinuosly(Dop, mesh); D₊ = couple_discontinuosly(Dop, mesh, Val(:plus)); D₋ = couple_discontinuosly(Dop, mesh, Val(:minus))
# D2 = sparse(D)^2
D2 = sparse(D₊) * sparse(D₋)
# D2 = sparse(D₋) * sparse(D₊)
D4 = D2^2
error_u = errors_hh_periodic_manufactured(usol, D, D2, D2, D4, D4, tspan, Tsit5(), tol, 1/N, true)
push!(val_error, error_u)
end
Dict("val_p$(p)_N" => val_N, "val_p$(p)_error" => val_error)
end
filename = "hh_periodic_manufactured_convergence_DG_D1pD1m"
even_odd_values = evenodd_values
res = Dict{String,Any}()
p = 1
val_N = round.(Int, 2 .^ range(4, 6.5, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 2
val_N = round.(Int, 2 .^ range(2, 4.5, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 3
val_N = round.(Int, 2 .^ range(2, 4.5, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 4
val_N = round.(Int, 2 .^ range(2, 4.5, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 5
val_N = round.(Int, 2 .^ range(2, 4.5, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
p = 6
val_N = round.(Int, 2 .^ range(2, 4.5, length=7)) |> even_odd_values
res = merge(res, do_stuff(p, val_N))
open("../data/" * filename * ".json", "w") do io
JSON.print(io, res, 2)
end
# +
fig, ax = plt.subplots(1, 1)
ax.set_prop_cycle(marker_cycler)
filename = "hh_periodic_manufactured_convergence_DG_D1pD1m"
res = JSON.parsefile("../data/" * filename * ".json")
let p = 1
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("EOC %.2f", -c1), (val_N[2], 2.0*val_error_u[2]), color="gray")
end
let p = 2
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("EOC %.2f", -c1), (val_N[end], 0.1*val_error_u[end]), color="gray")
end
let p = 3
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("EOC %.2f", -c1), (val_N[end], 1.5*val_error_u[end]), color="gray")
end
let p = 4
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("EOC %.2f", -c1), (1.2*val_N[end], 0.5*val_error_u[end]), color="gray")
end
let p = 5
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("EOC %.2f", -c1), (1.2*val_N[end], 0.2*val_error_u[end]), color="gray")
end
let p = 6
val_N, val_error_u = res["val_p$(p)_N"], res["val_p$(p)_error"]
ax.plot(val_N, val_error_u, label="\$ p = $p \$")
c0, c1 = linear_regression(log.(val_N), log.(val_error_u))
plt.plot(val_N, exp(c0).*val_N.^c1, marker="", linestyle=":", color="gray")
plt.annotate(@sprintf("EOC %.2f", -c1), (0.2*val_N[end], 0.1*val_error_u[end]), color="gray")
end
# ax.set_xlim(2^2-0.5, 2^13-1)
# ax.set_ylim(1.0e-13, 1.5e0)
ax.set_xscale("log", basex=2)
ax.set_yscale("symlog", linthreshy=1.0e-13)
ax.set_xlabel(L"N")
ax.set_ylabel(L"\| u - u_{\mathrm{ana}} \|")
fig.savefig("../figures/" * filename * ".pdf", bbox_inches="tight")
# -
| notebooks/hh.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# +
import json
import pydgraph
from pydgraph import DgraphClient, DgraphClientStub
from grapl_analyzerlib.schemas import *
from grapl_analyzerlib.schemas.schema_builder import ManyToMany
# +
def set_schema(client, schema, engagement=False):
op = pydgraph.Operation(schema=schema)
client.alter(op)
def drop_all(client):
op = pydgraph.Operation(drop_all=True)
client.alter(op)
def format_schemas(schema_defs):
schemas = "\n\n".join([schema.to_schema_str() for schema in schema_defs])
types = "\n\n".join([schema.generate_type() for schema in schema_defs])
return "\n".join([
" # Type Definitions",
types,
"\n # Schema Definitions",
schemas,
])
# -
mclient = DgraphClient(DgraphClientStub('alpha0.master-graph.grapl:9080'))
# +
# drop_all(mclient)
schemas = (
AssetSchema(),
ProcessSchema(),
FileSchema(),
IpConnectionSchema(),
IpAddressSchema(),
IpPortSchema(),
NetworkConnectionSchema(),
ProcessInboundConnectionSchema(),
ProcessOutboundConnectionSchema(),
RiskSchema(),
LensSchema(),
)
schema_str = format_schemas(schemas)
# print(schema_str)
set_schema(mclient, schema_str)
# + pycharm={"name": "#%%\n"}
import os
import string
from hashlib import pbkdf2_hmac, sha256
from random import randint, choice
import boto3
def hash_password(cleartext, salt) -> str:
hashed = sha256(cleartext).digest()
return pbkdf2_hmac(
'sha256',
hashed,
salt,
512000
).hex()
def create_user(username, cleartext):
assert cleartext
dynamodb = boto3.resource('dynamodb')
table = dynamodb.Table('user_auth_table')
# We hash before calling 'hashed_password' because the frontend will also perform
# client side hashing
cleartext += "f1dafbdcab924862a198deaa5b6bae29aef7f2a442f841da975f1c515529d254";
cleartext += username;
hashed = sha256(cleartext.encode('utf8')).hexdigest()
for i in range(0, 5000):
hashed = sha256(hashed.encode('utf8')).hexdigest()
salt = os.urandom(16)
password = hash_password(hashed.encode('utf8'), salt)
table.put_item(
Item={
'username': username,
'salt': salt,
'password': password
}
)
allchar = string.ascii_letters + string.punctuation + string.digits
password = "".join(choice(allchar) for x in range(randint(14, 16)))
print(f'your password is {password}')
username = ''
assert username, 'Replace the username with your desired username'
create_user(username, password)
password = ""
print("""Make sure to clear this cell and restart the notebook to ensure your password does not leak!""")
# + pycharm={"name": "#%%\n"}
# CLEAR CACHE
def clear_redis_caches():
from redis import Redis
def chunker(seq, size):
return [seq[pos:pos + size] for pos in range(0, len(seq), size)]
CACHE_ADDRS = [
]
CACHE_PORT = 6379
for CACHE_ADDR in CACHE_ADDRS:
r = Redis(host=CACHE_ADDR, port=CACHE_PORT, db=0, decode_responses=True)
for keys in chunker([k for k in r.keys()], 10000):
r.delete(*keys)
clear_redis_caches()
| etc/Grapl Provision.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp softwares.fastq.hisat2
# -
# # hisat2
# +
# export
from pybiotools4p.softwares.base import Base, modify_cmd
import os
# +
# export
class Hisat2(Base):
def __init__(self, software, fd):
super(Hisat2, self).__init__(software)
self._default = fd
if '/' in software:
bin = os.path.dirname(software) + '/'
else:
bin = ''
self._hisat2_build = bin + 'hisat2-build'
self._hisat2_extract_snps_haplotypes_UCSC = bin + 'hisat2_extract_snps_haplotypes_UCSC.py'
self._hisat2_align_l = bin + 'hisat2-align-l'
self._hisat2_extract_snps_haplotypes_VCF = bin + 'hisat2_extract_snps_haplotypes_VCF.py'
self._hisat2_align_s = bin + 'hisat2-align-s'
self._hisat2_extract_splice_sites = bin + 'hisat2_extract_splice_sites.py'
self._hisat2_inspect = bin + 'hisat2-inspect'
self._hisat2_build_l = bin + 'hisat2-build-l'
self._hisat2_inspect_l = bin + 'hisat2-inspect-l'
self._hisat2_build_s = bin + 'hisat2-build-s'
self._hisat2_inspect_s = bin + 'hisat2-inspect-s'
self._hisat2_extract_exons = bin + 'hisat2_extract_exons.py'
self._hisat2_simulate_reads = bin + 'hisat2_simulate_reads.py'
@modify_cmd
def cmd_version(self):
'''
:return:
'''
return 'echo {repr} ;{software} --version'.format(
repr=self.__repr__(),
software=self._software
)
@modify_cmd
def cmd_build_index(self, reference, genome_index_prefix, genome_ss=None, genome_exon=None,
genome_genotype=None, genome_snp=None):
'''
:param reference:
:param genome_ss:
:param genome_exon:
:param genome_genotype:
:param genome_snp:
:param genome_index_prefix:
:return:
'''
option = ''
if None is not genome_ss:
option += ' --ss ' + genome_ss
if None is not genome_exon:
option += ' --exon ' + genome_exon
if None is not genome_genotype:
option += ' --haplotype ' + genome_genotype
if None is not genome_snp:
option += ' --genome_snp ' + genome_snp
return r'''
{hisat_build} {build_index} {reference} {option} {genome_index_prefix}
'''.format(
hisat_build=self._hisat2_build,
build_index=self._default['build_index'],
reference=reference,
option=option,
genome_index_prefix=genome_index_prefix
)
@modify_cmd
def cmd_prepare_snp_ucsc(self,reference,snp_file,prefix):
'''
:param reference:
:param snp_file:
:param prefix:
:return:
'''
if None is snp_file:
return 'echo No snp_file'
return r'''
awk 'BEGIN{{OFS="\t"}} {{if($2 ~ /^chr/) {{$2 = substr($2, 4)}}; if($2 == "M") {{$2 = "MT"}} print}}' {snp_file} \
> {prefix}_snp.tmp
{software} {reference} {prefix}_snp.tmp {prefix}
'''.format(
software=self._hisat2_extract_snps_haplotypes_UCSC,
snp_file=snp_file,
prefix=prefix,
reference=reference
)
@modify_cmd
def cmd_prepare_exon_ss(self,gtf_file,prefix):
'''
:param gtf_file:
:param prefix:
:return:
'''
return r'''
{software1} {gtf_file} > {prefix}.ss
{software2} {gtf_file} > {prefix}.exon
'''.format(
software1=self._hisat2_extract_splice_sites,
software2=self._hisat2_extract_exons,
gtf_file=gtf_file,
prefix=prefix
)
def cmd_align(self, hisat2_idx, fq1, fq2, summary, samtools, samtools_idx, outbam):
'''
:param hisat2_idx:
:param fq1:
:param fq2:
:param summary:
:param samtools:
:param samtools_idx:
:param outbam:
:return:
'''
if None is fq2 or fq2 == '':
return r'''
{hisat2} {align_paras} -x {hisat2_idx} -U {fq1} --summary-file {summary} | {samtools_sam2bam} | {samtools_sort}
{samtools_index}
'''.format(
hisat2=self._software,
align_paras=self._default['align'],
samtools_sam2bam=samtools.cmd_sam2bam(samtools_idx, '-', bamfile=None),
samtools_sort=samtools.cmd_sort('-', sortbam=outbam),
samtools_index=samtools.cmd_index(outbam),
**locals()
)
else:
return r'''
{hisat2} {align_paras} -x {hisat2_idx} -1 {fq1} -2 {fq2} --summary-file {summary} | {samtools_sam2bam} | {samtools_sort}
{samtools_index}
'''.format(
hisat2=self._software,
align_paras=self._default['align'],
samtools_sam2bam=samtools.cmd_sam2bam(samtools_idx, '-', bamfile=None),
samtools_sort=samtools.cmd_sort('-', sortbam=outbam),
samtools_index=samtools.cmd_index(outbam),
**locals()
)
def __repr__(self):
return 'hisat2:' + self._software
def __str__(self):
return 'graph-based alignment of next generation sequencing reads to a population of genomes'
# +
import configparser
config=configparser.ConfigParser()
config.read('pybiotools4p/default.ini')
from pybiotools4p.softwares.bam.samtools import Samtools
samtools=Samtools('samtools',config['samtools'])
hisat2=Hisat2('hisat2',config['hisat2'])
# +
reference='biology-test-data/fasta/Homo_sapiens.GRCh38.dna.primary_assembly.chromosome22.fa'
gtf_file='biology-test-data/gtf/HS.22.gtf'
genome_index_prefix='./pybiotools/hisat2_index'
fq1='./biology-test-data/fastq/HS.22.r1.fq.gz'
fq2='./biology-test-data/fastq/HS.22.r2.fq.gz'
prefix='./pybiotools/HS_22'
snp_file='/path/to/snp'
prefix='./pybiotools/tmp'
genome_ss='./pybiotools/tmp.ss'
genome_exon='./pybiotools/tmp.exon'
summary_pe='./pybiotools/pe.summary.txt'
summary_se='./pybiotools/se.summary.txt'
samtools_idx='biology-test-data/fasta/Homo_sapiens.GRCh38.dna.primary_assembly.chromosome22.fa.fai'
outbam_pe='./pybiotools/hisat_pe.bam'
outbam_se='./pybiotools/hisat_se.bam'
# -
hisat2.cmd_version()
print(hisat2.cmd_build_index(reference, genome_index_prefix, genome_ss=None, genome_exon=None,
genome_genotype=None, genome_snp=None))
print(hisat2.cmd_prepare_snp_ucsc(reference,snp_file,prefix))
print(hisat2.cmd_prepare_exon_ss(gtf_file,prefix))
print(hisat2.cmd_build_index(reference, genome_index_prefix+'_ss_exon', genome_ss=genome_ss, genome_exon=genome_exon,
genome_genotype=None, genome_snp=None))
print(hisat2.cmd_align( genome_index_prefix, fq1, fq2, summary_pe, samtools, samtools_idx, outbam_pe))
print(hisat2.cmd_align( genome_index_prefix, fq1, '', summary_se, samtools, samtools_idx, outbam_se))
| 04_hisat2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # We will get started at 5 past the hour
# + [markdown] slideshow={"slide_type": "slide"}
# # Lecture 2 - Data Bootcamp
# ### Getting the goods on understanding data
# 
# - everything up until pandas we saw in 242, but now we will go a little deeper
# - obviously we don't have enough time to get into everthing in this class, but we will try to nerd out a little today
# + [markdown] slideshow={"slide_type": "slide"}
# ## Basic Data Types and Operations
# - what is a variable?
# - a variable is a location in memory
# - python takes care of this for us, on-the-fly and in a way that we never actually see
# - but if we take a peak beind the curtain, we can actually see how it actually works
# + slideshow={"slide_type": "fragment"}
x = 5
print (str(x)+ " and this variable's physical memory address is "+str(id(x)))
# + [markdown] slideshow={"slide_type": "slide"}
# - now, this is a good time to talk about _mutability_
# - mutability mean that variables can be changed without being erased.
# - some types of variables like integers are *immutable*
# - this means if we change them, they get a different memory address, and the old memory address gets recycled (thanks python!)
# + slideshow={"slide_type": "fragment"}
y = 5
print (str(y)+ " and this variable's physical memory address is "+str(id(y)))
y = "hello"
print (y+ " and this variable's physical memory address is "+str(id(y)))
# + [markdown] slideshow={"slide_type": "slide"}
# ### mutability continued
# - other variables can be changed in place, these are the mutable types
# - these are more complex data types, usually lists of information, dictionaries, data objects, and data frames
#
# + slideshow={"slide_type": "fragment"}
l = ["Hello","Goodbye"] # this is an array
print (str(l)+ " and this variables physical memory address is "+str(id(l)))
l.append("Coriander") # add this string to the array!
print (str(l)+ " and this variables physical memory address is "+str(id(l)))
# + [markdown] slideshow={"slide_type": "slide"}
# - mutability is important because of how memory management works
# - you can think about it in terms of the blockchain (Dodgecoin)
# - you don't want things to get overwritten
# - python does this for us without us seeing, which is great. in other languages you have to do it yourself!
# - if you don't use memory management well, you create 'leaks'. this causes the computer to crash!
# - in C you would have things called 'pointers' that are variables that track where memory exists. However, it used to be easy to get it wrong, and you could overwrite memory that is crtically important (like the memory that stores where the operating system exists!)
# - for us we only need to think about memory if we load data
# - lets say you have an array. then you add to it
# + slideshow={"slide_type": "slide"}
import sys # a systems command library
myArray = ["hello friends"] # simple array with one item
i =0 # simple integer counter
# to make this more understandable
byt = sys.getsizeof(myArray)
kb = byt/1024 #kilobytes
mb = kb/1024 #megabytes
# how much memory does this array take up?
print("myArray as bytes:"+str(byt)+"\n\tmyArray as kb:"+str(kb)+"\n\tmyArray as mb:"+str(mb))
# + slideshow={"slide_type": "fragment"}
# lets add some content to this array...
i =0
while i < 1:
myArray.append("adding item: "+str(i))
# what could go wrong ?
# use stop button after a bit
# + slideshow={"slide_type": "slide"}
print(str(len(myArray))) # print out the number of items in the array
# + slideshow={"slide_type": "fragment"}
import sys # we don't need to do this again, but in case this cell gets run out of sequence....
# to make this more understandable
byt = sys.getsizeof(myArray)
kb = byt/1024
mb = kb/1024
print("myArray as bytes:"+str(byt)+"\n\tmyArray as kb:"+str(kb)+"\n\tmyArray as mb:"+str(mb))
# + [markdown] slideshow={"slide_type": "fragment"}
# - in sum. memory management matters
# - mutable variables means we have to be careful about how we load and store data in variables
#
# ### but wait, what kinds of variables are there ?
# + [markdown] slideshow={"slide_type": "slide"}
# # simple variable types
# + [markdown] slideshow={"slide_type": "fragment"}
# ### basic data types and binary storage
# - int (1,2,3,4)
# - float (1.1,2.2,3.4)
# - bool
# - char/string "Hello World"
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### int
# - integer variables are whole numbers
# + slideshow={"slide_type": "fragment"}
# here is an int
x = 5
print("type of x:"+type(x).__name__)
print("size of x:"+str(sys.getsizeof(x)))
# + [markdown] slideshow={"slide_type": "slide"}
# ### float
# - floating point numbers are numbers with decimal values
# + slideshow={"slide_type": "fragment"}
# here is a float
y = 555555.533333
print("type of x:"+type(y).__name__)
print("size of x:"+str(sys.getsizeof(y)))
# + [markdown] slideshow={"slide_type": "slide"}
# ### ints are whole numbers. floats are decimal. but what happens when they mix?
# - when two types of variables mix, it is called casting
# - some types of casting are implicit.
# - this means that python seeing two type of data and makes a decision on how to match them for us
# + slideshow={"slide_type": "fragment"}
print(str(x/3))
# + slideshow={"slide_type": "fragment"}
y = x/3 #implicit casting
type(y)
# + slideshow={"slide_type": "fragment"}
y = int(x/3) #explicit casting
type (y)
y
# + [markdown] slideshow={"slide_type": "slide"}
# ### Bool
# - boolean variables are only True or false
# - they are used for conditional statements and flags
# - last week we used a boolean flag to set if out cake was done in our pseduocode
# + slideshow={"slide_type": "fragment"}
x = True
# + slideshow={"slide_type": "fragment"}
sys.getsizeof(x)
# funny, its about the size of an int....
# + [markdown] slideshow={"slide_type": "slide"}
# ### can you cast a boolean?
# + slideshow={"slide_type": "fragment"}
int(x)
# + slideshow={"slide_type": "fragment"}
y = False
int(y)
# + [markdown] slideshow={"slide_type": "slide"}
# ## what are bool good for?
# - booleans allow us to test logic
# - booleans give us the power of conditionals
# + slideshow={"slide_type": "fragment"}
x = 4
y = 5
x < 5
# + [markdown] slideshow={"slide_type": "fragment"}
# ### type of conditionals
# - there are many forms of conditionals
# - Equal to (==)
# - NOT (!)
# - OR (|) - inclusive
# - AND (&) -exclusive
# - exclusive OR (^) very exclusive. only if opposite
# - Greater Than (>)
# - Less Than (<)
# + slideshow={"slide_type": "slide"}
# equal to
x = 5
y = 4
x == y
# + slideshow={"slide_type": "fragment"}
print(True == False)
print(True == True)
print(False == False)
# + slideshow={"slide_type": "slide"}
# not
x = 5
y = 4
print(x != y)
# + slideshow={"slide_type": "fragment"}
print(True != False)
print(True != True)
# + slideshow={"slide_type": "slide"}
# or
x = True
y = False
print(x | y)
# + slideshow={"slide_type": "fragment"}
print(True | False)
print(True | True)
print(False | False)
# + slideshow={"slide_type": "slide"}
# and
# x & y
print(True & False)
print(True & True)
print(False & False)
# + slideshow={"slide_type": "slide"}
print(True ^ True)
print(False ^ False)
print(True ^ False)
print(False ^ True)
# + slideshow={"slide_type": "slide"}
#greater than
x > y
# + slideshow={"slide_type": "fragment"}
#less than
#x < y
x =5
y=5
x>=y
# + [markdown] slideshow={"slide_type": "slide"}
# ### char and strings
# - generally, chars (characters) are single items 'a' or 'b'
# - strings are sets of characters 'h','e','l','l','o'
# - in python, all chars are strings.
# + slideshow={"slide_type": "fragment"}
c = 'h'
type(c)
# + slideshow={"slide_type": "fragment"}
c = 'hello'
type(c)
# -
sys.getsizeof(c)
# + [markdown] slideshow={"slide_type": "slide"}
# ### strings have cool properties and methods
# - for example, we can use strings like they are arrays (more in these in a sec)
# + slideshow={"slide_type": "fragment"}
#here is a protery of the string c
c = 'hello'
c[3]
# + slideshow={"slide_type": "fragment"}
# we could also add two strings together
strA = "Hello"
strB = " Michael"
strA+strB
# + slideshow={"slide_type": "fragment"}
# here is a method we can call upon for var (variable) c
# c.upper()
#strings have lots of method/functions we can call
# for more of these, check out:
# c.capitalize()
c.count('l')
# + [markdown] slideshow={"slide_type": "slide"}
# ## lastly for now, escape sequences
# - escape sequences are how we are able to print out spceial items in a string
# - for example, how can you tell python to print items on a new line? how can you use a tab?
# - escape sequences are string modifiers. the usually begin with a "\\" and then a letter
# - a new line is \\n and a tab is \\t
# + slideshow={"slide_type": "fragment"}
strA = 'hello \nhow are you'
print(strA)
# + slideshow={"slide_type": "fragment"}
strB = "table layout \n1 \t2 \t3\n4 \t5 \t6\n7\t8\t9"
print(strB)
# + [markdown] slideshow={"slide_type": "slide"}
# ### but how do i print out an escape sequence?
# - add another "\\"
# + slideshow={"slide_type": "fragment"}
strC = "escape sequences are cool the the new line (\\n) and tab (\\t) escape \n\t see?\n\t\t...\n\t\t\t..."
print(strC)
# + [markdown] slideshow={"slide_type": "slide"}
# ### string casting
# - can you store the letter "s" on the hard disk?
# + [markdown] slideshow={"slide_type": "fragment"}
# - can you store the number 5 on the hard disk?
# + [markdown] slideshow={"slide_type": "fragment"}
# - if you can't store these letters and numbers, how are they stored?
# + [markdown] slideshow={"slide_type": "slide"}
# - strings are 'encoded' into memory
# - and different era's of computing used different kinds of encoding.
# - early encoding was ASCII
# - ascii gave us 256 values (8 bytes) to work with
#
# + [markdown] slideshow={"slide_type": "fragment"}
# 
# + [markdown] slideshow={"slide_type": "slide"}
# ### there are different types of encoding out there
# - ascii worked for a while
# - see:ascii art
#
# + [markdown] slideshow={"slide_type": "fragment"}
# 
# - courtesy wikipedia: https://en.wikipedia.org/wiki/ASCII_art
# + [markdown] slideshow={"slide_type": "slide"}
# ### clearly 256 characters isn't going to cut it.
# - more often now, we use unicode
# - unicode has multiple types of encoding for character sets (english and arabic for example)
# - unicode increases the size of characters from 256 to 143,859
# - see:https://en.wikipedia.org/wiki/Unicode
# - this matters because if you try display a string in one encoding type and it is another, you will have trouble
# + slideshow={"slide_type": "fragment"}
# let s be a string we want to convert to unicode character
s = "\U0001F590".encode("UTF-8")
print(s)
# + slideshow={"slide_type": "fragment"}
# okay, now decode the value again
print(s.decode())
# + [markdown] slideshow={"slide_type": "slide"}
# ## Arrays
# - arrays are lists of data
# - we have already seen them a couple times
# - we saw them in 242 last year of course
# - but we also saw them in lecture 1 with our names example, and today with strings
# - arrays are a fantastically lightweight way to store information
# + slideshow={"slide_type": "fragment"}
names = ["<NAME>","<NAME>","<NAME>","<NAME>",
"<NAME>","<NAME>","<NAME>","<NAME>",
"<NAME>","<NAME>","<NAME>","<NAME>","<NAME>",
"<NAME>","<NAME>","<NAME>","<NAME>","<NAME>",
"<NAME>","<NAME>","<NAME>","<NAME>","<NAME>",
"<NAME>","<NAME>","<NAME>","<NAME>","<NAME>"]
names
# + [markdown] slideshow={"slide_type": "slide"}
# - arrays are indexed
# - the index starts at 0
# - for the array \["hello","hi","gidday"\]
# - hello is index 0
# - hi is index 1
# - gidday is index 2
# - but the length of the array is 3
# - indexes are denoted by the \[\] characters
# + slideshow={"slide_type": "fragment"}
arr = ["hello","hi","gidday"]
arr[0]
# + slideshow={"slide_type": "fragment"}
# arr[len(arr)-1][len(arr[len(arr)-1]
i = arr[len(arr)-1]
i[len(i)-1]
#arr[len(arr)-1][len(arr[len(arr)-1])-1]
arr[-1][-1]
# + slideshow={"slide_type": "fragment"}
arr[3]
# + [markdown] slideshow={"slide_type": "slide"}
# ## arrays are lists of anything. including lists
# - this is where arrays really get useful
# - arrays are just collections of variables
# - you can mix variable types in arrays. I don't recommend it.
# + slideshow={"slide_type": "fragment"}
arr = ['hello',3,True]
sum(arr)
# + slideshow={"slide_type": "fragment"}
arr = [3,4,5]
sum(arr)
# + [markdown] slideshow={"slide_type": "slide"}
# ## arrays can have arrays as items
# - this is called a multi dimensional array
# + slideshow={"slide_type": "fragment"}
arr = [1,2]
arr2= [1,2,arr]
arr2
# + [markdown] slideshow={"slide_type": "fragment"}
# - a better idea is to make your dimensions consistent
# - this way you know you can expect elements in specific places. later in the course we are likely to have pretty crazy arrays, this will help a lot!
# + slideshow={"slide_type": "fragment"}
arr = [[1,2,3],[4,5,6],[7,8,9]]
arr
# + slideshow={"slide_type": "fragment"}
#is the same as
arr = [[1,2,3],
[4,5,6],
[7,8,9]]
# -
# ### arrays have a couple interesting functions you'll use regularly
# - len() is the number of items in an array
# - append() adds a new item to the array
# + [markdown] slideshow={"slide_type": "slide"}
# # Break!
#
# + [markdown] slideshow={"slide_type": "slide"}
# # patterns. if and loops
# - if, for, and while
# - these basic patterns get used over and over and over in programming. they allow us to process data relative to what we epect to see, and what we don't expect to see
# - introducing if, for, and while introduces us to encapulation and tab-indentation
# - anything that is tab-indented in something else belongs to it.
#
#
# - IF I GO TO THE STORE
# - THEN I SHOULD DO THIS THING
# - OTHERWISE
# - DO THIS OTHER THING
# - OH AND THIS OTHER THING TOO
# + [markdown] slideshow={"slide_type": "slide"}
# ### if
# - if is what changes the program flow based on data
# - there are three types of statements that we can use with if statements
# - if, else if,else
# - if statements are where boolean logic starts to get useful
# - if the logic is true, do this
# - else if, the logic is slightly different and true, do this
# - else, well fall back to this final thing
# + slideshow={"slide_type": "fragment"}
# if
x = 1
y = 2
if (x < y):
print(str(x)+" is less than "+str(y))
# + slideshow={"slide_type": "slide"}
# if-else
name = "<NAME>"
if (name == "<NAME>"):
print("you are the teaching leading this lecture")
else:
print ("you could be a teacher or a student")
# + slideshow={"slide_type": "fragment"}
# if-elif-else
name = "Amber"
# name = "Sila"
# name = "some other name"
if (name == "<NAME>"):
print("you are the teacher leading this lecture")
elif (name == "Sila"):
print ("ah, ok, you are also a teacher")
elif(name=="Amber"):
print("you are a tutor")
else:
print("you are not Michael or Sila, you must be a student")
# + [markdown] slideshow={"slide_type": "slide"}
# ### for loop
# - in python, for loops allow us to iterate through 'iterable' items
# - yes that is confusing, but it really means anything that has sub-items in it
# - an array is an iterable item
# + slideshow={"slide_type": "fragment"}
# for loop
arr = [1,2,3]
# for an item 'i' in the array 'arr'
for i in arr:
print ("item:"+str(i)) # print out the current item
# + [markdown] slideshow={"slide_type": "fragment"}
# - the for loop is deceptively useful.
# - you could use it to go through every polygon in a shapefile, for example
# + [markdown] slideshow={"slide_type": "slide"}
# ### while loop
# - while loops are like if statements, but far more dangerous.
# - remember at the beignning of this class when we intentionally made a memory leak?
# - imagine that, but you have no idea how you did it
# - that is what while loops do.
# + slideshow={"slide_type": "fragment"}
# while loops evaluate a condition at the top of every iteration
# they continue until that condition evaluates to false
while (true):
print("this loop will never end")
# + [markdown] slideshow={"slide_type": "slide"}
# - while loops are good if you have a non-iterable set of items that you need to get through
# - for example, you just want your array to count to 10
# + slideshow={"slide_type": "fragment"}
# a simple loop to count to 10
i = 0 # this is called a counter
while (i<=10): # keep going until i > 10
i = i+1 # increment counter by 1
print("i equals:"+str(i))
# what if we wanted it to actually go until it print i=10?
# -
# + [markdown] slideshow={"slide_type": "slide"}
# # Moving on to more complex data types
#
# - one last thing. Pandas.
#
# 
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Pandas
# 
#
# - for real though, what is pandas?
# - pandas is a _very_ powerful data handling and processing library for python.
# - it has a blazing fast ability to load and save data from a wide variety of formats (csv,json,excel, etc)
# - it can transform data very quickly, too.
# - ok, how do I get it?
# - run command prompt from anaconda
# - if you forget how, have a look at last week's lecture :)
# - type: pip install pandas
# + [markdown] slideshow={"slide_type": "slide"}
# ### Introducing, the dataframe
# - pandas is all organized around a concept called a DataFrame
# - the dataframe is like a crazzzzzzzy powerful 2D array
# - pandas brings data transformation, statistical analys, and plotting/visualization directly to you
# 
# + slideshow={"slide_type": "slide"}
# to get started, we need to import the pandas library
# if you get lost in class today, I highly recommend the pandas website
# the tutorials on the site are excellent!
# https://pandas.pydata.org/docs/getting_started
import pandas as pd
# to make a dataframe we can easily construct one ourselves
#creating a DataFrame using a dictionary
# we haven't covered dictionaries in detail yet, but we will be next week
# for now, all you need to know is that a dictionary is a
# type of data strcutre that stores information as "key":"value" pairs
# and like arrays the "value" can be any tpye of data, like in this case, an array
df = pd.DataFrame(
{
"Name":[
"Braund, Mr. <NAME>",
"Allen, Mr. <NAME>",
"<NAME>",
],
"Sex": ["male", "male", "female"],
"Age": [22, 35, 58],
})
df
# + [markdown] slideshow={"slide_type": "slide"}
# ### query the table, asking for a single column of information
# - in pandas, a column is a 'series'
# + slideshow={"slide_type": "fragment"}
# get a series (column) of data
df["Age"]
# + [markdown] slideshow={"slide_type": "slide"}
# ### pandas make summarizing data easy
# + slideshow={"slide_type": "fragment"}
# we can also simply as pandas for stats
# https://pandas.pydata.org/docs/getting_started/intro_tutorials/06_calculate_statistics.html
df.describe()
# + [markdown] slideshow={"slide_type": "slide"}
# ### pandas does plot the easy way!
# + slideshow={"slide_type": "fragment"}
# we can also ask pandas for a really simple graph of the data
# https://pandas.pydata.org/docs/getting_started/intro_tutorials/04_plotting.html
df.plot.bar(x='Name',y='Age')
# there are lots of different options for the type and styling of plots, far too much for today!
# for example these are all the types of plots!
# 'area','bar','barh','box','density',
# 'hexbin', 'hist', 'kde', 'line', 'pie', 'scatter'
# + [markdown] slideshow={"slide_type": "slide"}
# # lets make this interesting
# - over the weekend I got interested in harvesting Reddit.com data
# - reddit is an 'open api' meaning all the publically posted information is free to grab and play with
# - I'm going to gloss over how praw works a little over the next couple slides, but suffice to say it is pretty slick and makes getting data easy
# - if you want to install praw, just use
# - pip install praw
# + [markdown] slideshow={"slide_type": "slide"}
# ### First I'm going to collect login information
# - In the background I've created a developer account for reddit
# - I've written a file called reddit_app_login.json to sotre my login info
# - This is really important, as I don't want to broadcast my details with the code
# - A blank version is given with this week's zip file
# - you can create an account usingthe links in the code below
# + slideshow={"slide_type": "fragment"}
# useful links
# https://medium.com/@plog397/webscraping-reddit-python-reddit-api-wrapper-praw-tutorial-for-windows-a9106397d75e
# https://praw.readthedocs.io/en/latest/tutorials/comments.html
import praw # import the main module
from praw.models import MoreComments # a set of classes to handle comments
import datetime as dt # handling some time formats
# get account information from a JSON file
import json
with open("reddit_app_login.json","r") as read_file:
rKeys = json.load(read_file)
# rKeys.keys() # check to see if the right keys were collected from the JSON file
# + [markdown] slideshow={"slide_type": "slide"}
# ### getting the data from reddit
# - this is called 'calling' the API
# - we use this code to 'ask' reddit for the data we want
# + slideshow={"slide_type": "fragment"}
#reddit account information
reddit = praw.Reddit(
client_id=rKeys["client_id"],
client_secret=rKeys["client_secret"],
password=rKeys["password"],
user_agent=rKeys["user_agent"],
username=rKeys["username"],
)
# We are ready to start grabbing data from reddit
# lets grab from r/auckland
top_n = 1000 # how many posts would you like to get ?
r_auckland = reddit.subreddit('auckland')
new_auckland = r_auckland.hot(limit=top_n) # this time lets use the 'hot' posts
# Verify that we are recieving data
# this is a large amount of data so probably not going to do this.
# for i in new_auckland:
# print(i.title)
# + [markdown] slideshow={"slide_type": "slide"}
# ### getting dataframe ready
# - once we have the information, it isn't nessecarilly ready to be loaded directly in to a dataframe
# - its a simple matter of extracting the data in the new_auckland variable into a format pandas understands
# - in this case it is a dictionary
# +
# Now that we have the data from praw we want to format from its raw data
# into something we can stick into a Pandas Dataframe
# we make an empty dictionary called sub_post that can hold all data from the subreddit
sub_posts = {
"title":[],
"subreddit":[],
"score":[],
"id":[],
"url":[],
"comms_num": [],
"created_timestamp": [],
"created_datetime":[],
"body":[]}
# now, for each of the items in the subreddit that we found we are going
# to add the data from it into the dictionary keys as array items
for j in new_auckland:
sub_posts["title"].append(j.title) # note the append method. it is the same as we saw before for arrays
sub_posts['subreddit'].append(j.subreddit)
sub_posts["score"].append(j.score)
sub_posts["id"].append(j.id)
sub_posts["url"].append(j.url)
sub_posts["comms_num"].append(j.num_comments)
sub_posts["created_timestamp"].append(j.created)
sub_posts["created_datetime"].append(dt.datetime.fromtimestamp(j.created))
sub_posts["body"].append(j.selftext)
# now that the dictioary is in the shape pandas wants we
# convert our data into a pandas dataframe
rAuckland_df = pd.DataFrame(sub_posts)
# and now, lets write this dataframe to file
# in case anything happens to the data we can just reload it
filename = "top1000_rAuckland_posts.csv"
rAuckland_df.to_csv(filename)
# + [markdown] slideshow={"slide_type": "slide"}
# ### What did we just do?
# - if you look at the folder where this notbook is located, there will now be a file 'top1000_rAuckland_posts.csv'
# + slideshow={"slide_type": "fragment"}
# lets take a look at what pandas sees
# by defualt, pandas loads the top and bottom 10 items when we ask for information
filename="old_rAuckland_top_comments.csv"
rAuckland_df = pd.read_csv(filename,parse_dates=["created_datetime"]) # read the file and parse the datetime test as pandas datetime objets
# reddit_df
rAuckland_df
# + slideshow={"slide_type": "slide"}
# what are the stats of this?
rAuckland_df.describe()
# + [markdown] slideshow={"slide_type": "slide"}
# ### we can go deeper
# - getting all the posts was cool, but waht would be really cool would be to see all the comments posted in the posts
# - lets try and get every top level comment for every post ever made in r/Auckland
# + slideshow={"slide_type": "slide"}
# lets now put everything together into a master table of all the top posts from all the
# top 1000 hot posts on the subreddit
# create a dictionary object that will house all of our data for the time being,
# until we load it all into the Pandas Dataframe
# same as before
top_comments = {
"post_title":[],
"subreddit_name":[],
"subreddit_id":[],
"post_id":[],
"comment_id":[],
"comment_author":[],
"comment_body":[],
"created_utc":[],
"created_datetime":[],
"permalink":[],
"comment_score":[],
}
# -
# ### two layer deep loops
# - remember how we can have an array inside an array?
# - well we can have a loop inside a loop!
# - here we are just getting all the comments, for every post and adding it to a big master list
# - the code below has all been commented out to remind me not to run it.
# - it all works, it just takes a while and I don't want to waste out time waiting for all the comments to download
# + slideshow={"slide_type": "slide"}
# OK This one is going to be a bit of a doozey, but its not that hard really. we just need to follow the logic
# 1. for each of the top posts that we have gathered in the preceding dataframe
# 2. go through each of the top comments in the post
# 3. store the data in that top post as new data in the top_comments dictionary
# first, for each id in the rAuckland DataFrame
# for p in rAuckland_df["id"]:
# current_post = reddit.submission(id=p) # get the data from the submission/post
# # now, lets loop through the top level comments in the submission/post
# for top_level_comment in current_post.comments:
# if isinstance(top_level_comment, MoreComments): # this is a little helper to make sure data exists
# continue
# top_comments["post_title"].append(top_level_comment.submission.title)
# top_comments["subreddit_name"].append(top_level_comment.subreddit.display_name)
# top_comments["subreddit_id"].append(top_level_comment.subreddit_id)
# top_comments["post_id"].append(top_level_comment.parent_id)
# top_comments["comment_id"].append(top_level_comment.id)
# top_comments["comment_author"].append(top_level_comment.author)
# top_comments["comment_body"].append(top_level_comment.body)
# top_comments["created_utc"].append(top_level_comment.created_utc)
# top_comments["created_datetime"].append(dt.datetime.fromtimestamp(top_level_comment.created_utc))
# top_comments["permalink"].append(top_level_comment.permalink)
# top_comments["comment_score"].append(top_level_comment.score)
# # # And thats it! We have amde a master list of all the top comments for the top n posts of the reddit of interest!
# # create a pandas dataframe from the dictionary
# reddit_df = pd.DataFrame (top_comments)
# # and save for later
# filename="rAuckland_top_comments.csv"
# reddit_df.to_csv(filename)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Thats going to take a while...
# - actually its pretty fast given how much data its asking for
# - but in the meantime we can load the data that came with this folder for the lecture that I created ahead of time
# + slideshow={"slide_type": "fragment"}
# lets explore this dataset a little...
# load reddit_df from the csv we created.
filename="old_rAuckland_top_comments.csv"
reddit_df = pd.read_csv(filename,parse_dates=["created_datetime"]) # read the file and parse the datetime test as pandas datetime objets
reddit_df
# + [markdown] slideshow={"slide_type": "slide"}
# ### thats a lot of data. also a lot of posts with few comments.
# - lets investigate!
# + slideshow={"slide_type": "fragment"}
reddit_df["comment_score"].describe()
# + slideshow={"slide_type": "fragment"}
reddit_df["comment_score"].plot()
# + [markdown] slideshow={"slide_type": "slide"}
# ### investigation. When is the best time to post for max comments?
# + slideshow={"slide_type": "fragment"}
# https://pandas.pydata.org/docs/getting_started/intro_tutorials/09_timeseries.html
import matplotlib.pyplot as plt
fig,axs = plt.subplots(figsize=(12,4))
reddit_df.groupby(reddit_df["created_datetime"].dt.hour)["comment_score"].max().plot(kind='bar',rot=0,ax=axs)
plt.xlabel("hour of the day");
plt.ylabel("avg comment score");
# + [markdown] slideshow={"slide_type": "slide"}
# ### does comment length correlate to karma?
# + slideshow={"slide_type": "fragment"}
# https://pandas.pydata.org/docs/getting_started/intro_tutorials/10_text_data.html
reddit_df["comment_length"] = reddit_df["comment_body"].str.len()
reddit_df.plot.scatter(y="comment_score",x="comment_length",alpha=0.5,figsize=(12,6),logx=True) # Check out the axis!
# + [markdown] slideshow={"slide_type": "slide"}
# ### investigation. Who has the most single comment karma from r/auckland?
# - to answer this question we have to do something quite algorithmically taxing, groupby
# - to group all of the posts that are related to specific users, we have to sort, then summarize
# - in this groupby clause we use the .max() method that sums as it goes. Its kind of like the summary statistic you can add to a spatial join function in ArcGIS (*hiss*)
# + slideshow={"slide_type": "fragment"}
fig,axs = plt.subplots(figsize=(12,4))
# in this clause we use groupby. its a taxing operation
reddit_df.groupby(reddit_df["comment_author"])["comment_score"].max().plot(kind='bar',rot=0,ax=axs)
plt.xlabel("comment author");
plt.ylabel("avg comment score");
# + [markdown] slideshow={"slide_type": "slide"}
# ### final investigation. What user has the highest karma, from r/Auckland?
# + slideshow={"slide_type": "fragment"}
# create a new series 'k' by grouping name with summed karma
k = reddit_df.groupby(reddit_df["comment_author"])["comment_score"].sum()
#convert the series to a dataframe
l = pd.DataFrame(k)
# output a sorted version, and make permanent
l.sort_values(by=["comment_score"],ascending=False,inplace=True)
# the default version of this is too big a table, so lets just grab
# the top and bottom 5
overall_karma = pd.concat([l.head(5),l.tail(5)])
overall_karma
# + slideshow={"slide_type": "slide"}
overall_karma.plot()
# + [markdown] slideshow={"slide_type": "slide"}
# # Group Activity
# - in your groups, go through the /r/wallstreetbets dataset and see what you can come up with!
# - I've put a dataset in this week's zip file on Canvas called 'reddit_wsb.csv'
# - the data comes from Kaggle.com
# - Using we just looked at, plus using the pandas website, what can you hack together right now?
# - some ideas to get you started:
# - who posted the most?
# - when was the best time to get karma?
# +
# a dataset from r/wallstreetbets
# collected on kaggle using python lib praw
# dataset url: https://www.kaggle.com/gpreda/reddit-wallstreetsbets-posts
# here is some code to get you started
import pandas as pd
wsb = pd.read_csv("reddit_wsb.csv")
wsb.describe()
# -
| Lectures/Lecture 2/GISCI 343 - Lecture 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:pytorch-env]
# language: python
# name: conda-env-pytorch-env-py
# ---
# # For-Loops and List Comprehensions
# ## For Loops
#
# For-loops are an important *control-flow* construct - they allow you to repeatedly perform batches of similar operations. A for-loops needs an iterable to loop over; lists and their cousins are the most common iterables for this purpose.
# +
# sum the integers from 1 to 5
total =0
for j in [1,2,3,4,5]:
total = total + j
print(total)
# -
# A few points about this example.
#
# - The `in` keyword is used to specify the iterable over which we are looping.
# - The colon `:` begins the *body* of the loop.
# - **Indentation matters:** the same example would throw a syntax error if we omitted the indentation. In general indentation is very important when writing Python code and poor indentation can be a common cause of errors.
# The case of looping over integers up to `n` is so common that there is a dedicated function for achieving exactly this behavior: the `range()` function. To count from `0` to `n` inclusive, loop over `range(n+1)`:
# +
# Demonstrate use of the range: range(start,end,increment)
total =0
for j in range(1,101,2):
total = total + j
print(total)
# ---
# -
# ## Iterating over Strings
# Strings are also iterables:
# Print out elements of string (USE i)
s = "PIC16A is awesome"
for letter in s:
print(letter)
# ---
# A verbose way to achieve the same result, which is sometimes useful:
# Access elements of iterable (string) (use l)
for l in range(len(s)):
print(s[l])
# ---
# We can also use `str.split()` to loop over entire words:
# Splitting words use w
for w in s.split():
print(w)
# ---
# ## Indexing Variable
#
# In each case, the indexing variable is assigned in global scope (i.e. outside the context of the for loop), and can be used later if desired.
letter, l , w
# The indexing variable is reassigned with each iteration of the loop. This can occasionally be a source of mistakes. For example, take a moment to consider the following code: what is the value of `i` at the end of the loop?
# +
# Multiply all numbers 1-10 (INCORRECT)
i = 1
for i in range(1,11):
i = i*i
i
# -
# Compare to:
# +
# Multiply all numbers 1-10 (CORRECT)
i = 1
for j in range(1,11):
i = i*j
i
# -
# ## Creating Lists with For-Loops
#
# A versatile way to construct lists is by first initiating an empty list, and then incrementally adding to it. Suppose I wanted to make a list of all the squares of integers from 1 to 10. Here's a way to do this with a *for loop*:
# Generating a list of squares of numbers 1 to 10
squares = []
for i in range(1,11):
squares.append(i**2)
squares
# We can also create a list of the lengths of words in a string:
# +
# Get a list of the word lengths in a string
lengths = []
for word in s.split():
lengths.append(len(word))
lengths
# -
# ### List Comprehensions
#
# A much more compact and readable way to construct lists is provided by *list comprehensions.* List comprehensions are inspired by "set-builder" notation in mathematics. For example, we might write the `squares` list from above as
#
# $$\{i^2 \;|\; 1 \leq i \leq 10\}$$
#
# List comprehensions allow us to write very similar Python code, using the `for` keyword again.
# squares again
squares = [i**2 for i in range(1,11)]
squares
# We were able to condense the three lines of code from our for-loop into just one, readable line. Similarly,
# words in string
lengths = [len(words) for words in s.split()]
lengths
# We can also write *conditional* comprehensions to construct even more complex lists:
# comprehensions with conditions
squares = [i**2 for i in range(1,11) if i%2!=0 and i<9]
squares
# We can iterate over multiple indexing variables:
# comprehensions with multiple variables
new_list = [i*j for i in [1,2,3] for j in [1,2,3]]
new_list
# We can also easily construct lists of lists:
# Nest list comprehensions to get lists of lists
new_list = [[i*j for i in [1,2,3]] for j in [1,2,3]]
new_list
# Comprehensions are a powerful tool and should often be preferred to for-loops when constructing lists based on a simple set of rules. For more complicated list constructions, in which potentially multiple logic statements and calculations are required, the readability of a for loop is potentially the safer option.
| MM_material/lecture-materials/basic_objects/self_study_notebooks/blanks/for_loops_and_comprehensions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import json
import os
import warnings
os.chdir('..')
warnings.filterwarnings('ignore')
import numpy as np
import pandas as pd
import trane
# -
os.getcwd()
# +
# load a dataframe
df = pd.read_csv('Example/medical_no_show.csv', parse_dates=['appointment_day', 'scheduled_day'])
df = df.head(5000)
# load the table metadata
meta = trane.TableMeta(json.loads(open('Example/meta.json').read()))
# define a cutoff strategy
cutoff_fn = lambda rows, entity_id: np.datetime64('1980-02-25')
cutoff_strategy = trane.CutoffStrategy(generate_fn=cutoff_fn, description='with a fixed cutoff of 1980-02-25')
# define operations
filter_op = trane.ops.LessFilterOp(column_name='age'); filter_op.set_hyper_parameter(65)
row_op = trane.ops.IdentityRowOp(column_name='no_show')
transformation_op = trane.ops.IdentityTransformationOp(column_name='no_show')
aggregation_op = trane.ops.LastAggregationOp(column_name='no_show')
operations = [filter_op, row_op, transformation_op, aggregation_op]
# create the prediction problem
problem = trane.PredictionProblem(
operations=operations,
entity_id_col='appointment_id',
label_col='no_show',
table_meta=meta,
cutoff_strategy=cutoff_strategy)
# -
res = problem.execute(df)
res.head(10)
# +
# You can also generate questions
sampled_df = df.sample(frac=.2)
problem_generator = trane.PredictionProblemGenerator(
table_meta=meta, entity_col='appointment_id', label_col='no_show', filter_col='age')
problems = problem_generator.generate(sampled_df)
problems
# -
str(problems[0])
| Example/Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # http://tau-data.id/unhas/
# + [markdown] slideshow={"slide_type": "slide"}
# <center><img alt="" src="images/0_Cover.jpg"/></center>
#
# ## <center><font color="blue">Module 01: Pendahuluan Python</font></center>
# <b><center>(C) <NAME> - 2019</center>
# <center>tau-data Indonesia ~ https://tau-data.id</center>
# + [markdown] slideshow={"slide_type": "slide"}
# # <font color="blue">Workshop Schedule</font>
#
# <center><img alt="" src="images/Outline.jpeg"/></center>
# + [markdown] slideshow={"slide_type": "slide"}
# # <font color="blue"> Hardware Requirements:</font>
#
# Python Codes yang dibahas di workshop ini telah di verifikasi di lingkungan berikut:
#
# * Python 3.6.8 (**64 bit**) & OS Windows 10/Linux (Google Colab)
# * Program yang dijalankan di workshop ini membutuhkan **koneksi internet** saat di eksekusi.
# * Dianjurkan peserta mengupdate OS (Windows/Linux) sebelum mengikuti workshop.
# * Apple Macintosh **tidak disarankan** untuk digunakan untuk keperluan pengolahan data di Data Science/Machine Learning.
#
# Ekspektasi spesifikasi komputer ( **Online/Google Colab** ):
# * None
#
# Ekspektasi spesifikasi komputer ( **offline: WinPython/Anaconda** ):
# * **OS**: Windows/Linux **64bit with recent updates**
# * RAM 4Gb or More
# * CPU minimum 1 Ghz
#
# + [markdown] slideshow={"slide_type": "slide"}
# # <font color="blue">Instalasi
#
# 1. Menggunakan Google Colab seluruh link modul dapat diakses di link berikut: https://tau-data.id/unhas/
# 2. Untuk mengunduh seluruh materi workshop silahkan unduh dari link github berikut: https://github.com/taufikedys/unhas
# 3. Link untuk mengunduh program Python (WinPython) beserta modul workshop juga dapat dilakukan dari Link No. 1
# 4. Instalasi python offline di Linux harap dilakukan sendiri menggunakan Anaconda: https://problemsolvingwithpython.com/01-Orientation/01.05-Installing-Anaconda-on-Linux/
# + [markdown] slideshow={"slide_type": "slide"}
# ## <font color="blue">API Keys twitter (digunakan pada hari Ke-2)
#
# * Setelah workshop hari pertama selesai, peserta "disarankan" untuk mendaftar twitter API Keys.
# * https://www.youtube.com/watch?v=KPHC2ygBak4
# atau
# * https://www.youtube.com/watch?v=CvFRbHSvKAg
#
# ### Tips dan Catatan:
# * Silahkan lakukan langkah-langkah diatas. Jika ada masalah bisa ditanyakan setelah workshop hari ke-1.
# * Saat mendaftar API Keys twitter sampaikan bahwa alasan kita apply adalah untuk keperluan riset/academic. Bagi pekerja dapat memberikan alasan "mengembangkan App untuk client".
# * Karena keterbatasan waktu, di workshop ini kita hanya akan menggunakan twitter.
# -
# # <font color="blue">Resources Tambahan
#
# * Di setiap slide/cell biasanya terdapat link (tautan) pada istilah/konsep tertentu.
# * https://tau-data.id
#
# ## Daftar Pustaka
#
# * <NAME>., <NAME>., & <NAME>. (2016). <em>Mastering Natural Language Processing with Python. Packt Publishing Ltd.
# * <NAME>, <NAME>. (2017).Python Social Media Analytics. Packt Publishing Ltd.
# * <NAME>., & <NAME>. (2017). Natural language processing for social media. Synthesis Lectures on Human Language Technologies, 10(2), 1-195.
# * <NAME>., & <NAME>. (Eds.). (2007). Natural language processing and text mining. Springer Science Business Media.
# * <NAME>. (2014). Python 3 Text Processing with NLTK 3 Cookbook. Packt Publishing Ltd.
#
# * Referensi eBook gratis: https://www.learndatasci.com/free-data-science-books/
# # Outline Module
#
# <ol>
# <li><b>Pendahuluan:</b><br />
# * The Zen of Python<br />
# * Python distributions<br />
# * Python editors Jupyter & Spyder<br />
# * Python VS (R, Julia, Matlab, Java, C, PHP, etc)<br />
# * Strength & weakness of Python<br />
# * Google Colab</li>
# <li><b>Python Basics</b>:<br />
# * Syntax Format<br />
# (indenting, multiline, import, deklarasi/inisialisasi)<br />
# * Code descriptor & comments<br />
# * integer, float, bytes, boolean<br />
# * list, tuple, dictionary, (Frozen) Set<br />
# * types : beginner pitfall<br />
# * Slicing in Python</li>
# <li><b>Python Logic:</b><br />
# * (Nested – hierarchical) if Logic<br />
# * Looping For (& list comprehension)<br />
# * Iterator VS Iterable<br />
# * Looping while<br />
# * Breaking loop<br />
# * Python exception<br />
# * TQDM</li>
# </ol>
#
# <h1 id="Why-Python">Why Python (Interpreter and-or compiler)</h1>
#
# * Free
# * Community support
# * Growing in needs & popularity
# * Portable - Multi Platform
# * “Fast” http://s.id/4lP
# * Rich libraries/Modules (termasuk Matematika, Statistika, Data Science/Machine Learning, Big Data, etc)
# * Easier to learn (learning curve not steep)
# * OOP, etc
#
# <p><img alt="" src="images/1_why_python.jpg" style="width: 800px; height: 516px;" /></p>
#
# <h2 id="Filosofi-&-Sejarah-singkat-Python">Filosofi & Sejarah singkat Python</h2>
#
# <p>Python diperkenalkan di sekitar tahun 1980 oleh <strong><NAME></strong>.</p>
#
# <p>Python memiliki system yang "mirip" seperti Java (multi platform dan memiliki kernel), namun lebih mudah (tanpa deklarasi variabel seperti matlab). Python juga ringan sehingga supported di IoT/SBC devices.</p>
# <h1 id="Python-Benchmarks">Python Benchmarks</h1>
#
# <img alt="" src="images/1_Python_VS_TheRest.png" />
# <h2 id="Beberapa-kelemahan-Python:">Beberapa kelemahan Python:</h2>
#
# <ol>
# <li>Reference to Variabel (can be a good thing - older version)</li>
# <li>Dependensi dan Versi Module</li>
# <li>Dependensi Compiler C yang spesific dan terkadang tidak konsisten</li>
# <li>Message Parsing (process locked) - di pemrograman paralel</li>
# </ol>
#
# # Python Distro
#
# * "Vanilla" Python: https://www.python.org/
# * Anaconda : https://www.anaconda.com/distribution/#download-section
# * WinPython: https://sourceforge.net/projects/winpython/files/ ==> **Recommended** for academics
#
# Catt: Kita akan menggunakan **Python 3.6**
# # Python IDE: Jupyter VS Spyder
#
# * Jupyter: Not just Codes
# * Spyder: IDE - Integrated development environment
# * PyCharm
# * NotedPad++
# * Python hanya membutuhkan (sembarang) text + Terminal (misal command Prompt)
# # Google Colab
#
# * Cocok untuk komputer yang memiliki spesifikasi yang relatif minim.
# * Free with GPU support (penting saat nanti belajar Data Science/Machine Learning, terutama Deep Learning)
# * Google Colab dapat digunakan untuk memudahkan instalasi dan menyiapkan environment dalam menjalankan scripts Python untuk berbagai kegiatan seperti workshop, presentasi, kuliah, dsb:
#
# * https://colab.research.google.com
# <h1><font color="blue">Sebelum dimulai - Resources tambahan yang sangat baik untuk belajar Python</font></h1>
#
# * Python basic: https://www.python-course.eu/python3_history_and_philosophy.php
# * Data Science Basic: https://scikit-learn.org/stable/tutorial/index.html
# * Advance Python: http://andy.terrel.us/blog/2012/09/27/starting-with-python/
# * Visualisasi di Python: https://matplotlib.org/gallery.html
# # Syntax Python (Mohon untuk di coba di jupyter dan-atau Spyder)
#
# * indenting tanpa tanda "{}"
# * akhir code tanpa tanda ";", tapi bisa juga iya
# +
# Python tidak membutuhkan tanda semi colon ";" di akhir setiap baris seperti R, Java, atau C
n = 5
b = 3
b, n
# +
# tapi semi-colon bisa juga digunakan di Python untuk mengakhiri "satu instruksi"
n = 5; b = 3
print(b, n)
# Atau
a, b = 5, 3
a, b # di Jupyter perintah PRINT tidak dibutuhkan jika variabel yang ingin dilihat nilainya berada di baris terakhir.
# -
a = 5
b = 3
a, b = b,a
a, b
# +
# Python menggunakan "indenting" dan bukan tanda kurung seperti banyak bahasa pemrograman lain
if True:
print('ini menggunakan tab')
print('ini menggunakan spasi x4')
print('teks ini sudah diluar "IF"')
# inden dinyatakan dengan "tab" atau "spasi 4x"
# +
# Teks di python bisa menggunakan ' atau "
print(' "hi", text enclosed by \' ')
print(" 'hi', text enclosed by \" ")
# tanda "\" digunakan untuk merubah character khusus di python menjadi character biasa
# Sangat berguna bagi Data Scientist saat menangani data dari database
# -
# # Syntax Python (Mohon untuk di coba di jupyter dan Spyder)
#
# * "import" yang baik (misal math/numpy)
# * import alias
# * Deklarasi
# * Deklarasi multiline (it's a matter of preference)
sin(0)
# +
import math
import numpy
print(numpy.cos(0))
math.sin(0)
# +
from numpy import cos # import parsial
from numpy import sin
print(sin(3.14))
cos(0)
# +
# bisa menggunakan "alias" dan multi columns
import numpy as np, math
math.log10(100)
# -
# Python (seperti matlab) tidak memerlukan deklarasi variabel
a = 2.0
# Untuk mengetahui tipe suatu variabel kita bisa menggunakan perintah "type"
print(a)
type(a)
# +
# Tapi hati-hati "juggling variable"
# Analogi kaleng biscuit VS rengginang VS alat jahit
a = 3.0
print(type(a))
type(a/3)
# Solusi/Saran - untuk Data Science - atau Pemrograman secara umum
# +
# ini komentar satu baris
"""
ini komentar beberapa baris.
Sangat baik untuk memberi keterangan ttg suatu "fungsi" di Python
Atau keterangan tambahan lainnya
Komentar tidak di eksekusi oleh Python
"""
# -
# # Struktur data dasar di Python
#
# Di Python variabel tidak perlu deklarasi (seperti matlab)
a = 2
b = 3.4
c = 'teks'
d = True
# + slideshow={"slide_type": "slide"}
# Untuk mengetahui tipe suatu variabel kita bisa menggunakan perintah "type"
type(a)
# -
a = 3.4
# "Array" dasar di Python bisa berupa List, Tuple, Set atau Dictionary
A = [2,4,1,6,4,9,7,10]
A
A.sort()
A[::-1]
# Semua variable di Python adalah object
A
A.append(33)
print(A)
# Hati-hati
A.append([10,11])
A
# Gunakan "Extend"
A = [2,4,1,6,4,9,7,10]
A.extend([99,33])
A
dir(A)
# Alternatif
A = [2,4,1,6,4,9,7,10]
B = [99, 33]
A+B
A.pop()
A
# ## List VS Tuple VS SET VS Dictionary
A = [9,2,4,1] # List - weight - parameter/weight
T = (9,2,4,1) # Tuple - imutable - data
H = set([3,4,5,6,99, 99, 99]) # No order/urutan
D = {'a':33, 56:'xyz', 8:[3,4], 9:True}
# +
# Slicing & Modifying
# -
# # Di data science kapan sebaiknya menggunakan List, Tuple, Set, dan dictionary?
# # hati-hati Boolean Trap di Python
#
# Unfortunately it is not as easy in real life as it is in Python to differentiate between **true and false**:
# The following objects are evaluated by Python as **False**:
#
# - numerical zero values (0, 0.0, 0.0+0.0j),
# - the Boolean value False,
# - empty strings,
# - empty lists and empty tuples,
# - empty dictionaries.
# - special value "None".
#
# **All other values are considered to be True**
x = 0
if x:
print('x dianggap BENAR oleh Python')
else:
print('x dianggap SALAH oleh Python')
# # Usecase logic diatas
#
# * Pengolahan data media sosial
# * Pengolahan data dari database
# * Not so useful pada data terstruktur yang preprocessed/tidak noisy
# <h1 id="Pendahuluan-Logika-Percabangan-(Conditional-Statement)---IF-"><font color="blue">Pendahuluan Logika Percabangan (Conditional Statement) - IF </font></h1>
#
# <ul>
# <li>Bentuk (unit) paling dasar dari kecerdasan buatan atau pembelajaran komputer, dahulu disebut sebagai <em>explicit rules machine learning</em> (but of course it's not AI ... :)).</li>
# <li>Bentuk umumnya:<br />
# <strong>IF</strong> (<em>kondisi</em>) <strong>MAKA</strong> (<em>tindakan</em>)</li>
# <li>Syarat utamanya adalah "kondisi/syarat" harus bernilai Boolean (True/False), sedangkan tindakan bisa apa saja.</li>
# </ul>
# <p><img alt="" src="images/4_Cond_Statement_as_ML0.png" style="width: 800px ; height: 300px" /></p>
#
# <h1><font color="blue">Telaah Komponen Percabangan - IF </font></h1>
#
# <p><font color="blue"><img alt="" src="images/4_IF_Components_.png" style="width: 800px; height: 424px;" /></font></p>
# +
# Bentuk paling sederhana
x = input('Masukkan nilai x = ') # input by default dikenali sebagai "string/characters"
if int(x)<0: # sehingga perlu dirubah ke "integer" dengan perintah "int"
print('x negatif')
else:
print('x bukan negatif')
# +
# Boolean Statements in Python
a = False
b = True
print("AND: ", a & b, ", ATAU: ", a and b)
print("OR: ", a | b, ", ATAU: ", a or b)
# +
n = int(input('n = '))
if n%2==0:
print('genap')
elif(n%2==1):
print('ganjil')
else:
print('neither')
# -
# ## <font color="blue"> Exceptions (Try-Except): One of the reason why Python is awesome!</font>
#
# * Secara umum sebuah program sebaiknya harus memiliki "precaution", just in case input yang tidak diinginkan masuk ke sistem dan mengakibatkan error.
# * Istilah formal untuk hal ini adalah "Domain Constraint".
# * Pada dasarnya hampir semua bahasa pemrograman (dan database) mampu melakukan domain constraint. Namun di Python dapat dilakukan dengan jauuuuhhh lebih sederhana (simple)
n = 5
try:
print( 'nilai fungsi = ', (n**2-1) / ( (n-1)*(n-5)*(n-7) ) )
except:
print("I can't divide by 0 ... please try other integer")
# # <font color="blue"> Pendahuluan Loop di Python </font>
#
# <p><font color="blue"><img alt="" src="images/3_Looping_Components.png" style="width: 800px; height: 424px;" /></font></p>
#
# # <font color="blue"> Loop in Python and why it looks like an algorithm :) </font>
#
# * But first, let's talk about "**range function**" in Python
# Fungsi Range di Python
list( range(7) )
print(list(range(2)))
print(list(range(1, 5)))
print(list(range(1, 5, 2)))
print(list(range(5, -3, -2)))
for i in range(7):
print(i)
# +
A = [45, 60, 75, 99, 30]
for i, a in enumerate(A):
A[i] = 20 +a
A
# -
# <h1><font color="blue">Kasus sederhana data science: Skewness dan Transformasi Data </font></h1>
#
# <img alt="" src="images/3_data_transformation.png" style="width: 769px; height: 404px;" />
#
# <h1><font color="blue">Contoh Inverse Transformation : Cara 1 </font></h1>
# Contoh 1 bagaimana mentransform suatu data
A = [99,89,80,50,25,12,11,10,9,8,7,6,5,4,3,2,1, -1]
B = [] # inisialisasi dengan array (list) kosong
for datum in A:
B.append(1/datum) # APPEND adalah perintah untuk menambahkan elemen ke suatu array (list)
print(A)
print(B)
[1/a for a in A]
# <h1><font color="blue">Contoh Inverse Transformation : Cara 2 </font></h1>
#
# * Cara diatas memiliki kelemahan yang cukup serius ketika datanya besar. Apakah itu?
# * *Copy Memory*
# Contoh 2 bagaimana mentransform suatu data
# Kita akan menelaah lebih jauh di pertemuan setelah UTS
A = [99,89,80,50,25,12,11,10,9,8,7,6,5,4,3,2,1]
N = len(A)
for i in range(N):
A[i] = 1/A[i] # ini disebut in-place processing. Jauh lebih hemat memory
print(A)
#print(N)
# Contoh 3 bagaimana mentransform suatu data
# Cara yg spesifik untuk Python (recommended)
A = [99,89,80,50,25,12,11,10,9,8,7,6,5,4,3,2,1]
for i, a in enumerate(A): # perintah ENUMERATE di Python akan menghasilkan variable "index" dari "a"
A[i] = 1/a # tetap in-place processing
print(A)
# # <font color="blue"> List Comprehension: It's a Python thing</font>
# +
# Contoh 4 bagaimana mentransform suatu data
# Cara yg spesifik untuk Python (Highly recommended)
A = [99,89,80,50,25,12,11,10,9,8,7,6,5,4,3,2,1]
[1/x for x in A] # List comprehesion secara umum lebih cepat dan sederhana
# -
# <h1 id="-Pendahuluan-Indefinite-Loop---While-"><font color="blue">Pendahuluan Indefinite Loop - While </font></h1>
#
# <ol>
# <li>While loop digunakan saat kita hendak melakukan perulangan/iterasi, namun tidak mengetahui secara pasti jumlah perulangannya.</li>
# <li>Kondisi di perulangan = pencabangan IF, yaitu bernilai boolean</li>
# <li>Kondisi di While biasa disebut Stopper</li>
# <li>Stopper dapat diletakkan di awal atau akhir (Repeat-Until)</li>
# <li>Di dalam "loop body" wajib memuat suatu mekanisme yang mengakibatkan suatu saat Stopper bernilai True.</li>
# <li>Precautions sangat disarankan di While loop dengan memasukkan jumlah iterasi maksimum</li>
# </ol>
#
# <p><img alt="" src="images/5_while_loop_Schema.png" style="width: 280px; height: 225px;" /></p>
#
# Contoh Looping While di Python
n=7
while n>0:
print(n, end = ', ')
n=n-1
# # TQDM awesome modul untuk loop
#
# * Sangat disarankan di data science terutama untuk proses yg kompleks atau data yang besar
# +
from tqdm import tqdm # juga berfungsi di terminal unix/command prompt
from time import sleep
for i in tqdm(range(10)):
sleep(1) # menunggu 1 detik
# -
# # Print to File in Python
f = open("Output_Python.txt","w") # W = write new file, a='append existing file, new file if file does not exist'
print("ini output pertama", file=f)
f.close() # kalau tidak di close, maka bagaikan file Doc yg sedang dibuka microsoft word
# lihat di folder "notebook" akan ada file baru dengan nama file yg ditetapkan diatas
# Perhatikan perintah ini tidak mengeluarkan output langsung di notebook/terminal (silenced)
f = open("Output_Python.txt","w") # W = write new file, a='append existing file, new file if file does not exist'
print("ini output pertama lagi karena opennya pakai 'w'", file=f)
f.close()
f = open("Output_Python.txt","a") # W = write new file, a='append existing file, new file if file does not exist'
print("ini baru output kedua karena opennya pakai 'a'", file=f)
f.close()
# Jangan lupa utk mengingat "print" by default akan ganti baris, sehingga di akhir setiap print tidak perlu penambahan "\n"
# # Python is not perfect, but if we know its weakness we can use it perfectly
#
# ## Pointer to Reference
# +
# Hati-hati dengan copy of reference (Pointer to Variabel) yg implisit di Python
A =[2,3,4]
B = A
A.append(7) # Kita tidak melakukan apa-apa ke B
print('A = ', A)
print('B = ', B)
# Mengapa outputnya seperti itu???...
# Python sebisa mungkin menghindari "copy of memory" yg cocok utk DS/ML
# menyebalkannya behavious semacam ini terkadang berbeda untuk versi python yg berbeda.
# Data Scientist perlu informasi perubahan yg terjadi ketika memakai Python versi yg lebih terbaru
# # Solusi?
# -
A =[2,3,4]
B = A.copy() # ini yang mirip dgn bahasa pemrograman lain B=A
A.append(7)
print('A = ', A)
print('B = ', B)
# ## Zipping Lists dalam data science
# ### Akan sering ditemukan proses di DS dimana 2 buah vector/array harus di olah bersamaan pada setiap elemennya tapi dalam index yang bersesuaian.
# +
# Zipping List
A = ['Minggu','Senin','Jumat']
B = ['Libur', 'Kerja', 'Kerja tapi ingin Libur']
for a,b in zip(A,B):
print('{} {}'.format(a,b))
# Apa yang terjadi jika listnya beda ukuran???... ==> hati-hati
# sering juga digunakan di Parallel Processing di Python ==> dibahas di PDS3
# -
# ### Tidak hanya list comprehension
D = {hari:kegiatan for hari, kegiatan in zip(A,B)}
D
type(D)
D['Minggu']
# <h1 id="Fungsi-di-Matematika-VS-Pemrograman">Fungsi di Matematika VS Pemrograman</h1>
#
# <p><img alt="" src="images/7_Fungsi_di_Pemrograman_vs_Math.png" /></p>
#
# <h1 id="Fungsi-di-Python">Fungsi di Python (Structured/Procedural) - Pure Functions</h1>
#
# <p><img alt="" src="images/7_fungsi_py.png" /></p>
#
# +
# Fungsi di Python cukup sederhana dan
# parameter fungsi adalah variabel to reference
# Sebaiknya diberikan multiple lines of metaFunction (keterangan tentang fungsi)
def fungsi(A,B):
"""
ini adalah contoh fungsi sederhana penjumlahan 2 bilangan
input: a,b sembarang 2 bilangan (int/float)
output: a+b
"""
C = A+B
return C**2
fungsi(1, 2)
# +
# default Value sangat berguna di DS/ML
def fungsi(A, B=5):
C = B+A**2
return C
fungsi(7)
# -
# # Python untuk Komputasi Numerik
# +
import numpy as np
s = [2.1, 2.8, 1.9, 2.5, 2.7, 2.3, 1.8, 1.2, 0.9, 0.1]
C = np.array(s)
print(C)
C.shape
# -
# elemen wise operations
print(C * 2+1)
C = C.tolist()
type(C)
print(s * 2+1)
# Error : tidak bisa dilakukan di List
A = np.array([2,3])
B = np.array([1,5])
print(A*B)
print(np.dot(A,B)) # Jarak Euclidean di Data Science, misal k-Means
# # Exploratory Data Analysis (EDA)
#
# * Diperkenalkan oleh <NAME> 1961: " _Procedures for analyzing data, techniques for interpreting the results of such procedures, ways of planning the gathering of data to make its analysis easier, more precise or more accurate, and all the machinery and results of (mathematical) statistics which apply to analyzing data._"
# * Tukey promoted the use of five number summary of numerical data—the two extremes (maximum and minimum), the median, and the quartiles.
# * EDA refers to the critical process of performing **initial investigations on data** so as to discover patterns,to spot **anomalies**,to test hypothesis and to **check assumptions** with the help of **summary statistics and graphical representations**.
# * Tools: Python, R, S-Plus, etc
#
# <p><img alt="" src="images/edamethods.png" /></p>
# # Data Visualizations
# <p><img alt="" src="images/Purpose_Visualize_Data.jpg" /></p>
# # Tujuan EDA
#
# * **Suggest hypotheses** about the causes of observed phenomena
# * **Assess assumptions** on which statistical inference will be based
# * Support the **selection of appropriate statistical techniques**
# * Provide a basis for further data collection
# # Some References
#
# * Tukey, <NAME> (1977). Exploratory Data Analysis. Addison-Wesley. ISBN 978-0-201-07616-5.
# * <NAME>. <NAME>. and <NAME>. (2006) Visual Statistics: Seeing your data with Dynamic Interactive Graphics. Wiley ISBN 978-0-471-68160-1
# * <NAME> & <NAME> (2005) Exploratory Analysis of Spatial and Temporal Data. A Systematic Approach. Springer. ISBN 3-540-25994-5
# !mkdir data
# !wget -P data/ https://raw.githubusercontent.com/taufikedys/UnHas/master/data/price.csv
# +
# Importing Some Python Modules
import scipy, itertools, pandas as pd, matplotlib.pyplot as plt, seaborn as sns, numpy as np
import warnings; warnings.simplefilter('ignore')
from scipy import stats
from sklearn.preprocessing import StandardScaler, MinMaxScaler
# %matplotlib inline
plt.style.use('bmh'); sns.set()
# -
# # Studi Kasus
#
# * Misal seorang Data Scientist ditugaskan untuk menentukan investasi properti terbaik.
# * Tujuan analisa di modul ini adalah menemukan harga rumah yang lebih rendah dari harga pasaran
# * Asumsikan kita memiliki data harga rumah yang ditawarkan dan variabel-variabel terkait lainnya.
# * Untuk membuat keputusan investasi, kita akan melakukan EDA dan membuat pada data yang ada.
#
# <p><img alt="" src="images/Regression-model.jpg" /></p>
# # Contoh Kasus Data Harga Property Rumah
#
# * Sumber Data: http://byebuyhome.com/
# * Objective: menemukan harga rumah yang berada di bawah pasaran.
# * Variable:
# - **Dist_Taxi** – distance to nearest taxi stand from the property
# - **Dist_Market** – distance to nearest grocery market from the property
# - **Dist_Hospital** – distance to nearest hospital from the property
# - **Carpet** – carpet area of the property in square feet
# - **Builtup** – built-up area of the property in square feet
# - **Parking** – type of car parking available with the property
# - **City_Category** – categorization of the city based on the size
# - **Rainfall** – annual rainfall in the area where property is located
# - **House_Price** – price at which the property was sold
# Importing CSV data https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.read_csv.html
price = pd.read_csv('data/price.csv')
# Prefer XLS atau CSV di Data Science/Machine Learning ... Mengapa?
# Tipe Datanya : DataFrame (df)
type(price)
# Ukuran Data
N, P = price.shape
print('baris = ', N, ', Kolom = ', P)
# tipe data di setiap kolom
# Wajib di periksa apakah tipe datanya sudah tepat?
# Perhatikan df sebagaimana semua variable di Python diperlakukan seperti object
price.info()
# chosen at random
price.sample(8)
# # Removing a variable
# Drop kolom pertama karena tidak berguna (hanya index)
price.drop("Observation", axis=1, inplace=True)
price.sample(3)
# Cek tipe variabel
price.dtypes
# dataframe types: https://pbpython.com/pandas_dtypes.html
price['Parking'] = price['Parking'].astype('category')
price['City_Category'] = price['City_Category'].astype('category')
price.dtypes
# # Statistika Deskriptif
# Statistika Sederhana dari data "Numerik"-nya
price.describe(include='all')
# ini adalah parameter tambahan jika kita juga ingin mendapatkan statistik sederhana seluruh datanya
# (termasuk data kategorik)
price[['Dist_Taxi','Parking']].describe(include='all')
# # Variable Selection
# Memilih hanya variable dengan tipe tertentu
price_num = price.select_dtypes(include = ['float64', 'int64'])
price_num.head()
# Perhatikan price_num adalah variable df baru! ... (hati-hati di data yang besar)
# Memilih hanya variable dengan tipe tertentu
price_cat = price.select_dtypes(include = ['category'])
price_cat.head()
df = pd.get_dummies(price['Parking'], prefix='Park')
df.head()
# # Menggabungkan dengan data awal (concat)
df2 = pd.concat([price, df], axis = 1)
df2.head()
# # Distribusi nilai pada setiap variabel kategorik
# get all unique values of a variable/column
for col in price_cat.columns:
print(col,': ', set(price[col].unique()))
plt.figure(figsize=(8,6)) # https://matplotlib.org/api/_as_gen/matplotlib.pyplot.figure.html#matplotlib.pyplot.figure
p = sns.countplot(x="Parking", data=price)
# ini dilakukan jika kita ingin menyimpan plotnya ke dalam suatu file
p.figure.savefig('barChart.png')
# lihat di folder ipynb-nya akan muncul file baru.
# PieChart
plot = price.Parking.value_counts().plot(kind='pie')
# ## Apakah ada kecenderungan perbedaan harga rumah akibat dari tipe tempat parkir?
p = sns.catplot(x="Parking", y="House_Price", data=price)
# Apa yang bisa dilihat dari hasil ini?
# # Outlier using Normality Assumption
#
# <p><img alt="" src="images/XII_normal_CI.png" style="width: 800px; height: 374px;" /></p>
# Distributions
p = sns.distplot(price['House_Price'], kde=True, rug=True)
# +
# Misal dengan asumsi data berdistribusi normal
# dan menggunakan 99% confidence interval di sekitar variabel "harga"
df = np.abs(price.House_Price - price.House_Price.mean())<=(2*price.House_Price.std())
# mu-2s<x<mu+2s
print(df.shape)
df.head()
# -
price2 = price[df] # Data tanpa outliers
print(price2.shape, price.shape)
price2.head
# Perhatikan disini sengaja data yang telah di remove outliernya
# disimpan dalam variabel baru "Price2"
# Jika datanya besar hati-hati melakukan hal ini
# Distributions
p = sns.distplot(price2['House_Price'], kde=True, rug=True)
# Plot lagi setelah outlier removal
p= sns.catplot(x="Parking", y="House_Price", data=price2)
# Apakah ada kecenderungan perbedaan harga rumah akibat dari tipe tempat parkir?
# +
# Bisa juga plot dengan informasi dari 3 variabel sekaligus
# (untuk melihat kemungkinan faktor interaksi)
p= sns.catplot(x="Parking", y="House_Price", hue="City_Category", kind="swarm", data=price2)
# -
# # Ada "dugaan" apakah dari hasil diatas?
# # Missing Values
# General Look at the Missing Values
print(price2.isnull().sum())
# Simplest solution, if the MV is not a lot
# drop rows with missing values : Ada berbagai cara
X = price.dropna() # jika ada MV minimal satu di salah satu kolom, maka baris di hapus
price2.dropna(how='all') # jika ada MV di semua kolom, maka baris di hapus
price2.dropna(thresh=2) # jika ada MV minimal di salah 2 kolom, maka baris di hapus
price2.dropna(subset=['Dist_Hospital'])[:7] # jika ada MV minimal satu di salah kolom Dist_Hospital
# inplace=True if really really sure
price2.dropna(inplace=True)
print(price2.isnull().sum())
#mengecek apakah ada duplikat data?
print(price2.shape)
price2.duplicated().sum()
#menghapus entri yang memiliki data duplikat
price2.drop_duplicates(inplace=True)
print(price2.duplicated().sum()) # no more duplicates
print(price2.shape) # re-check by printing data size
# # (PairWise) Scatter Plot
# Scatter Plots; https://seaborn.pydata.org/generated/seaborn.pairplot.html
p = sns.pairplot(price2, hue="City_Category")
# Warning agak lambat (variabel cukup banyak)
# Coba kita perhatikan sebagiannya saja dulu dan coba kelompokkan berdasarkan "Parking"
p = sns.pairplot(price[['House_Price','Builtup','Dist_Hospital','Dist_Taxi','Parking','City_Category']], hue="City_Category")
# Ada pola menarik?
# # BoxPlot
# BoxPlots
p = sns.boxplot(x="House_Price", y="Parking", data=price2)
# Apa makna pola yang terlihat di data oleh BoxPlot ini?
# Jika ada outlier grafiknya menjadi tidak jelas (data = price, bukan price2)
p = sns.boxplot(x="House_Price", y="Parking", data=price)
# # Boxplot dapat juga dipisahkan berdasarkan suatu kategori
p = sns.catplot(x="Parking", y="House_Price", hue="City_Category", kind="box", data=price2)
# # Ada dugaan/interpretasi (baru) apakah dari boxPlot diatas?
# # Checking Correlations
price2.corr()
# +
# HeatMap untuk menyelidiki korelasi
corr2 = price2.corr() # We already examined SalePrice correlations
plt.figure(figsize=(12, 10))
sns.heatmap(corr2[(corr2 >= 0.5) | (corr2 <= -0.4)],
cmap='viridis', vmax=1.0, vmin=-1.0, linewidths=0.1,
annot=True, annot_kws={"size": 8}, square=True);
# + [markdown] slideshow={"slide_type": "slide"}
# # <center><font color="blue"> End of Module 01
#
# <hr />
# <p><img alt="" src="images/meme_4.png" /></p>
| UnHas_01_Pendahuluan_Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: venv_playground
# language: python
# name: venv_playground
# ---
# # **How to handle Exceptions?**
# Not all exceptions are exceptions.
# +
x = 10
if x > 5:
raise "x deve ser menor que 5"
# -
# There is an error hierarchy.
#
# - BaseException:
# - SystemExit
# - KeyboardInterrupt
# - GeneratorExit
# - <font color="blue">**Exception**</font>
# +
class BigNumbersError(BaseException):
pass
x = 10
if x > 5:
raise BigNumbersError("x deve ser menor que 5")
# -
# > **The built-in exception classes can be subclassed to define new exceptions; <font color="orange">programmers are encouraged to derive new exceptions from the Exception class or one of its subclasses, and not from BaseException.** <a href="https://docs.python.org/3/library/exceptions.html">Source</a></font>
# ## **Exceptions**
# + [markdown] tags=[]
# There are many exceptions available - <a href="https://docs.python.org/3/library/exceptions.html#exception-hierarchy">Exception hierarchy</a>
# <details>
#
# ```
# Exception
# +-- StopIteration
# +-- StopAsyncIteration
# +-- ArithmeticError
# | +-- FloatingPointError
# | +-- OverflowError
# | +-- ZeroDivisionError
# +-- AssertionError
# +-- AttributeError
# +-- BufferError
# +-- EOFError
# +-- ImportError
# | +-- ModuleNotFoundError
# +-- LookupError
# | +-- IndexError
# | +-- KeyError
# +-- MemoryError
# +-- NameError
# | +-- UnboundLocalError
# +-- OSError
# | +-- BlockingIOError
# | +-- ChildProcessError
# | +-- ConnectionError
# | | +-- BrokenPipeError
# | | +-- ConnectionAbortedError
# | | +-- ConnectionRefusedError
# | | +-- ConnectionResetError
# | +-- FileExistsError
# | +-- FileNotFoundError
# | +-- InterruptedError
# | +-- IsADirectoryError
# | +-- NotADirectoryError
# | +-- PermissionError
# | +-- ProcessLookupError
# | +-- TimeoutError
# +-- ReferenceError
# +-- RuntimeError
# | +-- NotImplementedError
# | +-- RecursionError
# +-- SyntaxError
# | +-- IndentationError
# | +-- TabError
# +-- SystemError
# +-- TypeError
# +-- ValueError
# | +-- UnicodeError
# | +-- UnicodeDecodeError
# | +-- UnicodeEncodeError
# | +-- UnicodeTranslateError
# +-- Warning
# +-- DeprecationWarning
# +-- PendingDeprecationWarning
# +-- RuntimeWarning
# +-- SyntaxWarning
# +-- UserWarning
# +-- FutureWarning
# +-- ImportWarning
# +-- UnicodeWarning
# +-- BytesWarning
# +-- ResourceWarning
# ```
#
# </details>
#
# -
# ### **AttributeError**
# > **<a href="https://docs.python.org/3/library/exceptions.html#AttributeError">Attribute Error</a>: Raised when an attribute reference or assignment fails.**
# obs: Also raises an error for a missing method
# +
class Dummy:
def __init__(self):
self.x = 1
pass
def do_stuff(self):
pass
try:
dummy = Dummy()
dummy.do_stuff2()
except Exception as e:
print(type(e),e)
pass
# -
# ### **LookupError**
# - > **<a href="https://docs.python.org/3/library/exceptions.html#IndexError">Index Error</a>: Raised when a sequence subscript is out of range.**
#
# - > **<a href="https://docs.python.org/3/library/exceptions.html#KeyError">Key Error</a>: Raised when a mapping (dictionary) key is not found in the set of existing keys.**
#
# +
lst = [1,2,3]
try:
lst[4]
except LookupError as e:
print(type(e),e,"\n")
dic = {"a":1,"b":2}
try:
dic["c"]
except LookupError as e:
print(type(e),e)
# -
lst = [1,2,3]
dic = {"a":1,"b":2}
try:
dic["c"]
except IndexError as e:
print(f"Index out of range: your list has {len(lst)} elements.")
except KeyError as e:
print(f"The selected key does not exist.")
# ### **TypeError**
# - > **<a href="https://docs.python.org/3/library/exceptions.html#TypeError">Type Error</a>: Raised when an operation or function is applied to an object of inappropriate type.**
try:
() + ""
except Exception as e:
print(type(e), e)
try:
len(1)
except Exception as e:
print(type(e), e)
# ### **NameError & UnboundLocalError**
# - > **<a href="https://docs.python.org/3/library/exceptions.html#NameError">Nmae Error</a>: Raised when a local or global name is not found.**
#
# - > **<a href="https://docs.python.org/3/library/exceptions.html#UnboundLocalError">UnboundLocal Error</a>: Raised when a reference is made to a local variable in a function or method, but no value has been bound to that variable.**
# +
def do():
print(var)
def do2():
print(var)
var=2
try:
do()
except NameError as e:
print(type(e),e)
try:
do2()
except NameError as e:
print(type(e),e)
# +
#One of the many reasons to avoid global variables.
var = 10
try:
do2()
except Exception as e:
print(type(e), e)
# -
# <font color="blue">A variable is determined to be free or local at compile time.</font> (important to better understand when creating decorators).
#
# <a href="https://www.youtube.com/watch?v=9v8eu4MOet8">Nice explanation</a>
# ## HTT
#
# Obs
#
# All assert statements will actually be removed from your code automatically when the interpreter is run with the -O or -OO flags to optimize the bytecode.
#
# Assert statemens should be used only in tests or during development phase.
# ## **Releasing resources**
# + tags=[]
"""
- What if file does not exist?
- What if an error occurs in the exception clause?
"""
def profund_math_to_solve_exception(x,y):
return x/y
#path = 'data/file.txt'
path = 'data/file2.txt'
x, y = 2,2
try:
#Consume resource
file = open(path, "r")
#Processing
text = file.read()
print(f"Content: {text}")
except FileNotFoundError as e:
print(f"An exception {type(e)} occured")
print("Solving exception ...")
_=profund_math_to_solve_exception(x,y)
print("Done")
print("Closing file ...")
file.close()
print("Done")
# +
"""
- What if the file does not exist?
- What if an error occurs in the exception clause?
"""
def profund_math_to_solve_exception(x,y):
return x/y
#path = 'data/file.txt'
path = 'data/file2.txt'
x, y = 0,0
try:
#Consume resource
file = open(path, "r")
#Processing
text = file.read()
print(f"Content: {text}")
except FileNotFoundError as e:
print(f"An exception {type(e)} occured")
print("Solving exception ...")
valor=profund_math_to_solve_exception(x,y)
print("Done")
finally:
print("Closing file ...")
file.close()
print("Done")
# -
# > In real world applications, the finally clause is useful for releasing external resources (such as files or network connections), regardless of whether the use of the resource was successful. https://docs.python.org/3/tutorial/errors.html#tut-userexceptions
# +
"""
Example 1
- What if file does not exist?
"""
def profund_math_to_solve_exception(x,y):
return x/y
path = 'data/file.txt'
try:
#Consume resource
file = open(path, "r")
text = file.read()
print(f"Content: {text}")
except:
valor=profund_math_to_solve_exception(2,2)
print("Closing file ...")
file.close()
print("Done")
# +
"""
Horrible
"""
path = "data/manager.txt"
try:
file = open(path, "r")
texts = file.read().splitlines()
print(texts)
except:
pass
# +
"""
If the finally clause executes a break, continue or return statement, exceptions are not re-raised.
"""
def my_test():
try:
raise KeyboardInterrupt
finally:
print('Goodbye, world!')
return 42
my_test()
# +
"""
If the finally clause executes a break, continue or return statement, exceptions are not re-raised.
"""
def my_test():
try:
return "try"
finally:
return 'Finnaly'
my_test()
# + tags=[]
try:
with open("test.txt", "r") as f:
f.read()
except Exception as e:
raise "rodrigo"
# + tags=[]
try:
with open("test.txt", "r") as f:
f.read()
except Exception as e:
print(type(e))
raise e("rodrigo")
# -
try:
x=0/0
except Exception as e:
print(type(e))
raise e("rodrigo")
# +
x = 10
if x > 5:
raise Exception("x deve ser menor que 5")
# -
# ## **Nem toda exceção é uma exceção**
#
# - If some code path simply must broadly catch all exceptions - for example, the top-level loop for some long-running persistent process - then each such caught exception must write the full stack trace to a log or file, along with a timestamp. Not just the exception type and message, but the full stack trace.
#
# - For all other except clauses - which really should be the vast majority - the caught exception type must be as specific as possible. Something like KeyError, or ConnectionTimeout, etc.
#
| advanced/error/error.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Exercise 2.14 A measure of correlation (normalized mutual information)
# (Source: (Cover and Thomas 1991, Q2.20).) Let $X$ and $Y$ be discrete random variables which are identically distributed (so $H(X) = H(Y))$ but not necessarily independent. Define
#
# $$
# r = 1-\frac{H(Y|X)}{H(X)}
# $$
#
# * a) Show $r = \frac{I(X, Y)}{H(X)}$
# * b) Show $0 \le r \le 1$
# * c) When is $r=0$?
# * d) When is $r=1$?
# In this question, we are going to analyse the meaning of $r$, a function defined in terms of entropies. As the text hints, we should expect that $r$ behaves like a information theory correlation function. In other words, $r$ should be a normalized value that tells how dependent two random variables are from each other.
# ### Solution
# #### (a)
# We must show that $r=\frac{I(X, Y)}{H(X)}$:
#
# $$
# r = 1 - \frac{H(Y|X)}{H(X)} = \frac{H(X) - H(Y|X)}{H(X)} = \frac{I(X, Y)}{H(X)}
# $$
#
# #### (b)
#
# $$
# H(p) = \sum p(-\log p)\ge 0
# $$
#
# Using this, we can write
#
# $$
# \frac{H(Y|X)}{H(X)} \ge 0 \implies 1- \frac{H(Y|X)}{H(X)} \le 1
# $$
#
# Also,
#
# $$
# I(X, Y) = H(X) - H(Y|X)\ge 0 \implies H(Y|X) \le H(X) \implies 1-\frac{H(Y|X)}{H(X)} \ge 0
# $$
# Thus $0\le r \le 1$
#
# #### (c)
# $$
# r = \frac{I(X, Y)}{H(X)} = 0 \iff I(X, Y) = 0
# $$
#
# #### (d)
# $r = 1 \iff I(X, Y) = H(X).$ Which is the same thing as saying $r=1 \iff H(Y|X) = 0$. Let's see what $H(Y|X) = 0$ means:
#
# \begin{aligned}
# H(Y|X) = \sum_xp(x)H(Y|X=x) & = \sum_xp(x)\sum_yp(y|x)(-\log(p(y|x)) \\
# & = \sum_x\sum_y p(x)p(y|x)(-\log(p(y|x))
# \end{aligned}
#
# Observe that each term of the summation is non negative. Thus every term must be 0 in order for the above equation to be true. Thus $p(y|x)=0$ or $\log p(y|x) = 0$. The second condition only happens when $p(y|x) = 1$.
#
# So $H(Y|X) = 0$ says to us that given $X$, we know exactly which $Y$ is going to occur. In other words, $Y = f(X)$. Since the problem is symmetrical, $f$ is invertible and we can also express this as $X = f^{-1}(X)$.
# ### Conclusion
#
# In this section we study the parameter $r$ defined as $r=1-\frac{H(Y|X)}{H(X)}$. We discovered it behaves very similarly to the classical correlation. Being more specific, we found out four things:
#
# 1. $r$ is a ratio between mutual information and entropy, much like the classical correlation is a ratio between covariance and variances (Note that since we are assuming $H(X) = H(Y)$ then $r=\frac{I(X,Y)}{\sqrt{H(x)}\sqrt{H(Y)}}$.
# 2. The ratio is normalised (i.e. between 0 and 1)
# 3. $r=0\iff X$ and $Y$ are independent
# 4. $r=1\iff Y = f(X)$, where $f$ is invertible.
#
# Overall, we see that the parameter $r$ has the same goal as $\rho$, but it is more general, because it uses entropies instead of covariances. The first advantage is that it captures linear and non-linear functions between $X$ and $Y$. The second advantage is that $r=0 \implies X$ and $Y$ are independent.
| murphy-book/chapter02/q14.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import os
import regex as re
from collections import Counter, defaultdict
import sys
CONST_A = 0
CONST_C = 1
CONST_G = 2
CONST_T = 3
CONST_NT_MAP = ['A', 'C', 'G', 'T']
def remove_duplicates_round(df,hamm_thres=4,merge_counts=False):
seqs = list(df.Seq.values)
counts = list(df.Counts.values)
c = 0
while c<(len(counts)-1):
if(distance(seqs[c],seqs[c+1]))<hamm_thres:
if(counts[c]>counts[c+1]):
if(merge_counts):
counts[c]+=counts[c+1]
del counts[c+1],seqs[c+1]
else:
if(merge_counts):
counts[c+1]+=counts[c]
del counts[c],seqs[c]
else:
c+=1
return pd.DataFrame({'Seq':seqs,'Counts':counts})
def remove_all_duplicates(sequences,counts,hamming_thresh=4,merge_counts=False):
df = pd.DataFrame({'Seq':sequences,'Counts':counts})
seq_len = len(sequences[0])
print('Removing hamming neighbors on dimension:')
for i in range(seq_len):
df = df.ix[(df.Seq.str.slice(seq_len-i)+df.Seq.str.slice(i)).sort_values().index]
df = remove_duplicates_round(df,hamm_thres=hamming_thresh,merge_counts=merge_counts)
print(i)
return df
def distance(astring, bstring) :
distance = 0
limit = len(astring)
diff = len(bstring) - len(astring)
if len(bstring) < len(astring) :
limit = len(bstring)
diff = len(astring) - len(bstring)
for i in range(limit) :
if astring[i] != bstring[i] :
distance += 1
return distance + diff
def increment_bp_map(seq, bp_map, magnitude=1) :
for i in range(0, len(seq)) :
if seq[i] == 'A' :
bp_map[i][CONST_A] += magnitude
elif seq[i] == 'C' :
bp_map[i][CONST_C] += magnitude
elif seq[i] == 'G' :
bp_map[i][CONST_G] += magnitude
elif seq[i] == 'T' :
bp_map[i][CONST_T] += magnitude
return bp_map
def get_consensus_sequence(bp_map) :
seq = ''
for i in range(0, len(bp_map)) :
max_count = 0
max_j = 0
for j in range(0, 4) :
if bp_map[i][j] > max_count :
max_count = bp_map[i][j]
max_j = j
seq += CONST_NT_MAP[max_j]
return seq
def get_hamming_neighbor_1(seq, seq_map, start_r, end_r) :
for i in range(start_r, end_r) :
for base1 in CONST_NT_MAP :
mut_seq = seq[:i] + base1 + seq[i+1:]
if mut_seq in seq_map :
return mut_seq
return None
def get_hamming_neighbor_2(seq, seq_map, start_r, end_r) :
for i in range(start_r, end_r) :
for j in range(i + 1, end_r) :
for base1 in CONST_NT_MAP :
for base2 in CONST_NT_MAP :
mut_seq = seq[:i] + base1 + seq[i+1:j] + base2 + seq[j+1:]
if mut_seq in seq_map :
return mut_seq
return None
# +
#a = [[0] * 4] * 6
a = []
for i in range(0, 6) :
a.append([])
for j in range(0, 4) :
a[i].append(0)
at_seq = 'ATATAT'
a = increment_bp_map(at_seq, a)
print(a)
# -
r1 = 'C:/Users/Johannes/Desktop/apa_analysis/r1_apa_fr.fq'
r2 = 'C:/Users/Johannes/Desktop/apa_analysis/r2_apa_fr.fq'
# +
dna_regex = re.compile(r"(CAGACACAGCTC){s<=1}") # Sequence found only in plasmid amplicon on read 2
upstream_regex = re.compile(r"(CAATTCTGCT[ACGTN]{40}CTAAAATATA){s<=2}") # 12x2 bp sequences flanking upstream randomized region
downstream_regex = re.compile(r"(AGTATGAAAC[ACGTN]{20}ACCCTTATCC){s<=2}") # 12x2 bp sequences flanking dnstream randomized region
seq_regex = re.compile(r"(CAATTCTGCT[ACGTN]{40}CTAAAATATA){s<=2}.*(AGTATGAAAC[ACGTN]{20}ACCCTTATCC){s<=2}")
is_dna = 'CAGACACAGCTC'
n10_a = re.compile(r"(TGTTAAGAAC[ACGTN]{10}CTGGTAACTGACCTTCAAAG){s<=3}")
n10_b = re.compile(r"(TGTTAAGAACAAGTT[ACGTN]{10}AACTGACCTTCAAAG){s<=3}")
n10_c = re.compile(r"(TGTTAAGAACAAGTTTGGCT[ACGTN]{10}ACCTTCAAAG){s<=3}")
n20_a = re.compile(r"([ACGTN]{20}CTGGTAACTGACCTTCAAAG){s<=3}")
n20_b = re.compile(r"(TGTTAAGAAC[ACGTN]{20}ACCTTCAAAG){s<=3}")
n20_c = re.compile(r"(TGTTAAGAACAAGTTTGGCT[ACGTN]{20}){s<=3}")
wt20_down = re.compile(r"(GATGTCTCGTGATCTGGTGT){s<=3}")
# +
f = {}
f[0] = open(r1,'r')
f[1] = open(r2,'r')
head, seq, pr, q = ({} for i in range(4))
n_seqs, n_qual, xyseqs = ({} for i in range(3))
count = 0
dna_test_limit = 1
dna_count_map = {}
dna_quality_map = {}
dna_seq_map = {}
dna_lib_map = {}
valid_dna_seq_length = 113
valid_seq_count = 0
n20_a_wt_down_count = 0
n20_a_n20_down_count = 0
n20_b_wt_down_count = 0
n20_b_n20_down_count = 0
n20_c_wt_down_count = 0
n20_c_n20_down_count = 0
dna_bp_map = {}
dna_upstream_map = {}
dna_downstream_map = {}
dna_pas_map = {}
dna_seq_map = {}
matched_on_dict_count = 0
matched_on_hamming1_count = 0
matched_on_hamming2_count = 0
while True:
for i in range(2):
head[i] = f[i].readline()[:-1]
seq[i] = f[i].readline()[:-1]
pr[i] = f[i].readline()[:-1]
q[i] = f[i].readline()[:-1]
if len(q[1]) == 0:
break # End of File
dna_test = seq[1][12:24]
dna_test_distance = distance(dna_test, is_dna)
if dna_test_distance <= dna_test_limit :
upstream_flank = re.search(upstream_regex, seq[0][12:200])
downstream_flank = re.search(downstream_regex, seq[0][100:220])
both_flank = re.search(seq_regex, seq[0][12:220])
if upstream_flank != None and downstream_flank != None and both_flank != None:
valid_seq_count += 1
upstream_flank_seq = upstream_flank.group()
downstream_flank_seq = downstream_flank.group()
both_flank_seq = both_flank.group()
n10_match = False
if re.search(n10_a, upstream_flank_seq) or re.search(n10_b, upstream_flank_seq) or re.search(n10_c, upstream_flank_seq) :
n10_match = True
n20_match = False
if re.search(n20_a, upstream_flank_seq) or re.search(n20_b, upstream_flank_seq) or re.search(n20_c, upstream_flank_seq) :
n20_match = True
if n10_match == False and n20_match == True and len(both_flank_seq) == valid_dna_seq_length :
is_n20_down = True
if re.search(wt20_down, downstream_flank_seq) :
is_n20_down = False
upstream_seq = upstream_flank_seq[10:50]
downstream_seq = downstream_flank_seq[10:30]
pas_seq = both_flank_seq[50:65]
full_seq = both_flank_seq[10:103]
dna_lib = -1
start_r = 0
end_r = 40
if re.search(n20_a, upstream_flank_seq) and is_n20_down == False :
n20_a_wt_down_count += 1
upstream_seq = upstream_seq[0:20] + 'CTGGTAACTGACCTTCAAAG'
dna_lib = 0
start_r = 0
end_r = 20
elif re.search(n20_a, upstream_flank_seq) and is_n20_down == True :
n20_a_n20_down_count += 1
upstream_seq = upstream_seq[0:20] + 'CTGGTAACTGACCTTCAAAG'
dna_lib = 3
start_r = 0
end_r = 20
elif re.search(n20_b, upstream_flank_seq) and is_n20_down == False :
n20_b_wt_down_count += 1
upstream_seq = 'TGTTAAGAAC' + upstream_seq[10:30] + 'ACCTTCAAAG'
dna_lib = 1
start_r = 10
end_r = 30
elif re.search(n20_b, upstream_flank_seq) and is_n20_down == True :
n20_b_n20_down_count += 1
upstream_seq = 'TGTTAAGAAC' + upstream_seq[10:30] + 'ACCTTCAAAG'
dna_lib = 4
start_r = 10
end_r = 30
elif re.search(n20_c, upstream_flank_seq) and is_n20_down == False :
n20_c_wt_down_count += 1
upstream_seq = 'TGTTAAGAACAAGTTTGGCT' + upstream_seq[20:40]
dna_lib = 2
start_r = 20
end_r = 40
elif re.search(n20_c, upstream_flank_seq) and is_n20_down == True :
n20_c_n20_down_count += 1
upstream_seq = 'TGTTAAGAACAAGTTTGGCT' + upstream_seq[20:40]
dna_lib = 5
start_r = 20
end_r = 40
dx, xd = upstream_flank.start(), upstream_flank.end()
qualityscore = np.array([ord(i) - 33 for i in q[0][dx:xd]])
new_member = True
upstream_seq_key = upstream_seq
if upstream_seq in dna_count_map :
new_member = False
matched_on_dict_count += 1
else :
upstream_seq_h1 = get_hamming_neighbor_1(upstream_seq, dna_count_map, start_r, end_r)
if upstream_seq_h1 != None :
new_member = False
upstream_seq_key = upstream_seq_h1
matched_on_hamming1_count += 1
else :
upstream_seq_h2 = get_hamming_neighbor_2(upstream_seq, dna_count_map, start_r, end_r)
if upstream_seq_h2 != None :
new_member = False
upstream_seq_key = upstream_seq_h2
matched_on_hamming2_count += 1
if new_member == True :
dna_count_map[upstream_seq_key] = 0
dna_quality_map[upstream_seq_key] = 0
dna_lib_map[upstream_seq_key] = dna_lib
dna_upstream_map[upstream_seq_key] = {}
dna_downstream_map[upstream_seq_key] = {}
dna_pas_map[upstream_seq_key] = {}
dna_seq_map[upstream_seq_key] = {}
dna_count_map[upstream_seq_key] += 1
dna_quality_map[upstream_seq_key] += qualityscore
if upstream_seq not in dna_upstream_map[upstream_seq_key] :
dna_upstream_map[upstream_seq_key][upstream_seq] = 1
else :
dna_upstream_map[upstream_seq_key][upstream_seq] += 1
if pas_seq not in dna_pas_map[upstream_seq_key] :
dna_pas_map[upstream_seq_key][pas_seq] = 1
else :
dna_pas_map[upstream_seq_key][pas_seq] += 1
if downstream_seq not in dna_downstream_map[upstream_seq_key] :
dna_downstream_map[upstream_seq_key][downstream_seq] = 1
else :
dna_downstream_map[upstream_seq_key][downstream_seq] += 1
if full_seq not in dna_seq_map[upstream_seq_key] :
dna_seq_map[upstream_seq_key][full_seq] = 1
else :
dna_seq_map[upstream_seq_key][full_seq] += 1
#if upstream_flank_seq not in dna_seq_map :
# dna_seq_map[upstream_flank_seq] = both_flank_seq
if (count % 100000) == 0:
print(count)
print(str(valid_seq_count) + ' valid DNA reads extracted.')
print('N20 A WT DOWN: ' + str(n20_a_wt_down_count))
print('N20 A N20 DOWN: ' + str(n20_a_n20_down_count))
print('N20 B WT DOWN: ' + str(n20_b_wt_down_count))
print('N20 B N20 DOWN: ' + str(n20_b_n20_down_count))
print('N20 C WT DOWN: ' + str(n20_c_wt_down_count))
print('N20 C N20 DOWN: ' + str(n20_c_n20_down_count))
print('Number of unique DNA members: ' + str(len(dna_count_map)))
print('Matched on dictionary: ' + str(matched_on_dict_count))
print('Matched on hamming 1: ' + str(matched_on_hamming1_count))
print('Matched on hamming 2: ' + str(matched_on_hamming2_count))
count += 1
print('COMPLETE')
print(str(valid_seq_count) + ' valid DNA reads extracted.')
print('N20 A WT DOWN: ' + str(n20_a_wt_down_count))
print('N20 A N20 DOWN: ' + str(n20_a_n20_down_count))
print('N20 B WT DOWN: ' + str(n20_b_wt_down_count))
print('N20 B N20 DOWN: ' + str(n20_b_n20_down_count))
print('N20 C WT DOWN: ' + str(n20_c_wt_down_count))
print('N20 C N20 DOWN: ' + str(n20_c_n20_down_count))
print('Number of unique DNA members: ' + str(len(dna_count_map)))
print('Matched on dictionary: ' + str(matched_on_dict_count))
print('Matched on hamming 1: ' + str(matched_on_hamming1_count))
print('Matched on hamming 2: ' + str(matched_on_hamming2_count))
f[0].close()
f[1].close()
# +
upstream_seq_map = {}
pas_seq_map = {}
downstream_seq_map = {}
seq_map = {}
for upstream_key in dna_upstream_map :
upstream_bp_map = []
pas_bp_map = []
downstream_bp_map = []
seq_bp_map = []
for i in range(0, 40) :
upstream_bp_map.append([])
for j in range(0, 4) :
upstream_bp_map[i].append(0)
for i in range(0, 15) :
pas_bp_map.append([])
for j in range(0, 4) :
pas_bp_map[i].append(0)
for i in range(0, 20) :
downstream_bp_map.append([])
for j in range(0, 4) :
downstream_bp_map[i].append(0)
for i in range(0, 93) :
seq_bp_map.append([])
for j in range(0, 4) :
seq_bp_map[i].append(0)
upstream_list = list(dna_upstream_map[upstream_key].keys())
pas_list = list(dna_pas_map[upstream_key].keys())
downstream_list = list(dna_downstream_map[upstream_key].keys())
seq_list = list(dna_seq_map[upstream_key].keys())
if dna_count_map[upstream_key] > 2 :
for upstream in dna_upstream_map[upstream_key] :
upstream_bp_map = increment_bp_map(upstream, upstream_bp_map, magnitude=dna_upstream_map[upstream_key][upstream])
upstream_seq_map[upstream_key] = get_consensus_sequence(upstream_bp_map)
else :
upstream_seq_map[upstream_key] = upstream_list[0]
if dna_count_map[upstream_key] > 2 :
for pas in dna_pas_map[upstream_key] :
pas_bp_map = increment_bp_map(pas, pas_bp_map, magnitude=dna_pas_map[upstream_key][pas])
pas_seq_map[upstream_key] = get_consensus_sequence(pas_bp_map)
else :
pas_seq_map[upstream_key] = pas_list[0]
if dna_count_map[upstream_key] > 2 :
for downstream in dna_downstream_map[upstream_key] :
downstream_bp_map = increment_bp_map(downstream, downstream_bp_map, magnitude=dna_downstream_map[upstream_key][downstream])
downstream_seq_map[upstream_key] = get_consensus_sequence(downstream_bp_map)
else :
downstream_seq_map[upstream_key] = downstream_list[0]
if dna_count_map[upstream_key] > 2 :
for seq in dna_seq_map[upstream_key] :
seq_bp_map = increment_bp_map(seq, seq_bp_map, magnitude=dna_seq_map[upstream_key][seq])
seq_map[upstream_key] = get_consensus_sequence(seq_bp_map)
else :
seq_map[upstream_key] = seq_list[0]
# +
dna_upstream_list = []
dna_pas_list = []
dna_downstream_list = []
dna_seq_list = []
dna_upstream_count_list = []
dna_pas_count_list = []
dna_downstream_count_list = []
dna_seq_count_list = []
dna_count_list = []
dna_lib_list = []
for upstream in dna_count_map :
dna_upstream_list.append(upstream_seq_map[upstream])
dna_pas_list.append(pas_seq_map[upstream])
dna_downstream_list.append(downstream_seq_map[upstream])
dna_seq_list.append(seq_map[upstream])
dna_upstream_count_list.append(len(dna_upstream_map[upstream]))
dna_pas_count_list.append(len(dna_pas_map[upstream]))
dna_downstream_count_list.append(len(dna_downstream_map[upstream]))
dna_seq_count_list.append(len(dna_seq_map[upstream]))
dna_count_list.append(dna_count_map[upstream])
dna_lib_list.append(dna_lib_map[upstream])
df = pd.DataFrame({'upstream_seq': dna_upstream_list,
'pas_seq': dna_pas_list,
'downstream_seq': dna_downstream_list,
'seq': dna_seq_list,
'unique_upstream_seq_count': dna_upstream_count_list,
'unique_pas_seq_count': dna_pas_count_list,
'unique_downstream_seq_count': dna_downstream_count_list,
'unique_seq_count': dna_seq_count_list,
'library' : dna_lib_list,
'read_count': dna_count_list})
df = df.sort_values('read_count')
new_columns = ['upstream_seq', 'pas_seq', 'downstream_seq', 'seq', 'unique_upstream_seq_count', 'unique_downstream_seq_count', 'unique_pas_seq_count', 'unique_seq_count', 'library', 'read_count']
df.to_csv('apa_nextseq_v2_dna_20160922.csv', sep=',', header=True, columns=new_columns, index=False)
lib_summary = [0, 0, 0, 0, 0, 0]
for lib in dna_lib_list :
lib_summary[lib] += 1
for i in range(0, len(lib_summary)) :
print('Member count for library ' + str(i) + ': ' + str(lib_summary[i]))
# +
dna_upstream_key_list = list(dna_count_map.keys())
print(len(dna_upstream_key_list))
print(len(dna_count_list))
hamming_thresh = 4
filtered_dna_df = remove_all_duplicates(dna_upstream_key_list, dna_count_list, hamming_thresh, merge_counts=False)
hamming_upstream_list = list(filtered_dna_df.Seq.values)
hamming_count_list = list(filtered_dna_df.Counts.values)
print(len(hamming_upstream_list))
print(len(hamming_count_list))
print('{:,}'.format(len(hamming_upstream_list)) + ' sequences with levenshtein d >= ' + str(hamming_thresh))
# +
filtered_upstream_list = []
filtered_pas_list = []
filtered_downstream_list = []
filtered_seq_list = []
filtered_upstream_count_list = []
filtered_pas_count_list = []
filtered_downstream_count_list = []
filtered_seq_count_list = []
filtered_count_list = []
filtered_lib_list = []
for upstream in hamming_upstream_list :
if dna_count_map[upstream] > 2 or (dna_count_map[upstream] == 2 and len(dna_pas_map[upstream]) == 1 and len(dna_downstream_map[upstream]) == 1) :
filtered_upstream_list.append(upstream_seq_map[upstream])
filtered_pas_list.append(pas_seq_map[upstream])
filtered_downstream_list.append(downstream_seq_map[upstream])
filtered_seq_list.append(seq_map[upstream])
filtered_upstream_count_list.append(len(dna_upstream_map[upstream]))
filtered_pas_count_list.append(len(dna_pas_map[upstream]))
filtered_downstream_count_list.append(len(dna_downstream_map[upstream]))
filtered_seq_count_list.append(len(dna_seq_map[upstream]))
filtered_count_list.append(dna_count_map[upstream])
filtered_lib_list.append(dna_lib_map[upstream])
print(len(filtered_upstream_list))
df = pd.DataFrame({'upstream_seq': filtered_upstream_list,
'pas_seq': filtered_pas_list,
'downstream_seq': filtered_downstream_list,
'seq': filtered_seq_list,
'unique_upstream_seq_count': filtered_upstream_count_list,
'unique_pas_seq_count': filtered_pas_count_list,
'unique_downstream_seq_count': filtered_downstream_count_list,
'unique_seq_count': filtered_seq_count_list,
'library' : filtered_lib_list,
'read_count': filtered_count_list})
df = df.sort_values('read_count')
new_columns = ['upstream_seq', 'pas_seq', 'downstream_seq', 'seq', 'unique_upstream_seq_count', 'unique_pas_seq_count', 'unique_downstream_seq_count', 'unique_seq_count', 'library', 'read_count']
df.to_csv('apa_nextseq_v2_dna_filtered_20160922.csv', sep=',', header=True, columns=new_columns, index=False)
lib_summary = [0, 0, 0, 0, 0, 0]
for lib in filtered_lib_list :
lib_summary[lib] += 1
for i in range(0, len(lib_summary)) :
print('Member count for library ' + str(i) + ': ' + str(lib_summary[i]))
# -
| data/random_mpra/individual_library/tomm5/unprocessed_data/tomm5_dna_processing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Network with Applications
#
# In this file, we'll demonstrate the simulation of a more complicated network topology with randomized applications. These applications will act on each node, first choosing a random other node from the network and then requesting a random number of entangled pairs between the local and distant nodes. The network topology, including hardware components, is shown below:
#
# <img src="./notebook_images/star_network.png" width="700"/>
# ## Example
#
# In this example, we construct the network described above and add the random request app included in SeQUeNCe. We'll be building the topology from an external json file `star_network.json`.
#
# ### Imports
# We must first import the necessary tools from SeQUeNCe.
# - `Timeline` is the main simulation tool, providing an interface for the discrete-event simulation kernel.
# - `Topology` is a powerful class for creating and managing complex network topologies. We'll be using it to build our network and intefrace with specific nodes and node types.
# - `RandomRequestApp` is an example application included with SeQUeNCe. We will investigate its behavior when we add applications to our network.
import pandas as pd
from ipywidgets import interact
import time
from sequence.kernel.timeline import Timeline
from sequence.topology.topology import Topology
from sequence.app.random_request import RandomRequestApp
# ### Building the Simulation
#
# We'll now construct the network and add our applications. This example follows the usual process to ensure that all tools function properly:
# 1. Create the timeline for the simulation
# 2. Create the simulated network topology. In this case, we are using an external JSON file to specify nodes and their connectivity.
# - This includes specifying hardware parameters in the `set_parameters` function, defined later
# 3. Install custom protocols/applications and ensure all are set up properly
# 4. Initialize and run the simulation
# 5. Collect and display the desired metrics
#
# The JSON file specifies that network nodes should be of type `QuantumRouter`, a node type defined by SeQUeNCe. This will automatically create all necessary hardware and protocol instances on the nodes, and the `Topology` class will automatically generate `BSMNode` instances between such nodes.
#
# To construct an application, we need:
# - The node to attach the application to
# - The names (given as strings) of other possible nodes to generate links with
# - A seed for the internal random number generator of the application
#
# We can get a list of all desired application nodes, in this case routers, from the `Topology` class with the `get_nodes_by_type` method. We then set an application on each one, with the other possible connections being every other node in the network. We also give a unique random seed `i` to each application.
def test(sim_time, qc_atten):
"""
sim_time: duration of simulation time (ms)
qc_atten: quantum channel attenuation (dB/km)
"""
network_config = "star_network.json"
# construct the simulation timeline; the constructor argument is the simulation time (in ps)
tl = Timeline(sim_time * 1e9)
tl.seed(0)
# here, we make a new topology with name "network_topo" and attached to timeline tl
network_topo = Topology("network_topo", tl)
network_topo.load_config(network_config)
set_parameters(network_topo, qc_atten)
# construct random request applications
node_names = [node.name for node in network_topo.get_nodes_by_type("QuantumRouter")]
apps = []
for i, name in enumerate(node_names):
other_nodes = node_names[:] # copy node name list
other_nodes.remove(name)
# create our application
# arguments are the host node, possible destination node names, and a seed for the random number generator.
app = RandomRequestApp(network_topo.nodes[name], other_nodes, i)
apps.append(app)
app.start()
# run the simulation
tl.init()
tick = time.time()
tl.run()
print("execution time %.2f sec" % (time.time() - tick))
for app in apps:
print("node " + app.node.name)
print("\tnumber of wait times: ", len(app.get_wait_time()))
print("\twait times:", app.get_wait_time())
print("\treservations: ", app.reserves)
print("\tthroughput: ", app.get_throughput())
# create a table to showcase information about the reservations
print("\nReservations Table:\n")
node_names = []
start_times = []
end_times = []
memory_sizes = []
for node in network_topo.get_nodes_by_type("QuantumRouter"):
node_name = node.name
for reservation in node.network_manager.protocol_stack[1].accepted_reservation:
s_t, e_t, size = reservation.start_time, reservation.end_time, reservation.memory_size
if reservation.initiator != node.name and reservation.responder != node.name:
size *= 2
node_names.append(node_name)
start_times.append(s_t)
end_times.append(e_t)
memory_sizes.append(size)
log = {"Node": node_names, "Start_time": start_times, "End_time": end_times, "Memory_size": memory_sizes}
df = pd.DataFrame(log)
print(df)
# ### Setting parameters
#
# Here we define the `set_parameters` function we used earlier. This function will take a `Topology` as input and change many parameters to desired values.
#
# Quantum memories and detectors are hardware elements, and so parameters are changed by accessing the hardware included with the `QuantumRouter` and `BSMNode` node types. Many complex hardware elements, such as bsm devices or memory arrays, have methods to update parameters for all included hardware elements. This includes `update_memory_params` to change all memories in an array or `update_detector_params` to change all detectors.
#
# We will also set the success probability and swapping degradation of the entanglement swapping protocol. This will be set in the Network management Module (specifically the reservation protocol), as this information is necessary to create and manage the rules for the Resource Management module.
#
# Lastly, we'll update some parameters of the quantum channels. Quantum channels (and, similarly, classical channels) can be accessed from the `Topology` class as the `qchannels` field. Since these are individual hardware elements, we will set the parameters directly.
def set_parameters(topology, attenuation):
# set memory parameters
MEMO_FREQ = 2e3
MEMO_EXPIRE = 0
MEMO_EFFICIENCY = 1
MEMO_FIDELITY = 0.9349367588934053
for node in topology.get_nodes_by_type("QuantumRouter"):
node.memory_array.update_memory_params("frequency", MEMO_FREQ)
node.memory_array.update_memory_params("coherence_time", MEMO_EXPIRE)
node.memory_array.update_memory_params("efficiency", MEMO_EFFICIENCY)
node.memory_array.update_memory_params("raw_fidelity", MEMO_FIDELITY)
# set detector parameters
DETECTOR_EFFICIENCY = 0.9
DETECTOR_COUNT_RATE = 5e7
DETECTOR_RESOLUTION = 100
for node in topology.get_nodes_by_type("BSMNode"):
node.bsm.update_detectors_params("efficiency", DETECTOR_EFFICIENCY)
node.bsm.update_detectors_params("count_rate", DETECTOR_COUNT_RATE)
node.bsm.update_detectors_params("time_resolution", DETECTOR_RESOLUTION)
# set entanglement swapping parameters
SWAP_SUCC_PROB = 0.90
SWAP_DEGRADATION = 0.99
for node in topology.get_nodes_by_type("QuantumRouter"):
node.network_manager.protocol_stack[1].set_swapping_success_rate(SWAP_SUCC_PROB)
node.network_manager.protocol_stack[1].set_swapping_degradation(SWAP_DEGRADATION)
# set quantum channel parameters
ATTENUATION = attenuation
QC_FREQ = 1e11
for qc in topology.qchannels:
qc.attenuation = ATTENUATION
qc.frequency = QC_FREQ
# ### Running the Simulation
#
# All that is left is to run the simulation with user input. Note that different hardware parameters or network topologies may cause the simulation to run for a very long time.
interact(test, sim_time=50e3, qc_atten=[0, 1e-5, 2e-5])
| example/random_request_network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="u1nPXqSteG5O" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} outputId="c73aac8a-a670-4cf4-c85b-bfce83b2f1bd" executionInfo={"status": "ok", "timestamp": 1524098558754, "user_tz": 240, "elapsed": 1274, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "117867794654112750077"}}
import os
import numpy as np
import pandas as pd
import cv2
import matplotlib.pyplot as plt
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
from sklearn.model_selection import train_test_split
import itertools
from keras.preprocessing import image
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import ReduceLROnPlateau
from keras.models import Sequential, Model
from keras.layers import Dense, Activation, Flatten, Dropout, concatenate, Input, Conv2D, MaxPooling2D
from keras.optimizers import Adam, Adadelta
from keras.layers.advanced_activations import LeakyReLU
from keras.utils.np_utils import to_categorical
# + id="RygTVXJcfcOF" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
import scipy.io as sio
My_data = sio.loadmat('drive/Plant Classification Using C-CNN/train/Image_Processed_1data.mat')
x_train = My_data['train']
labels = My_data["train_labels"]
# + id="M3cWdhcYfkIm" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
#x_train, x_val, y_train, y_val = train_test_split(x_train, labels, test_size = 0.1, random_state=10, stratify=labels)
#print(len(x_train), len(x_val), len(y_train), len(y_val))
# + id="sF76LRdZfpm9" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
#x_train_dummy = x_train
x_train, x_val, y_train, y_val = train_test_split(x_train, labels, test_size = 0.1, random_state=10, stratify=labels)
x_train, x_test, y_train, y_test = train_test_split(x_train, y_train, test_size = 0.1, random_state=10, stratify =y_train)
# + id="n64GH5f7XsdX" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
#print('Train data:', len(x_train), ', Val data:', len(x_val), ', Test data:', len(x_test), ', Train labels:', len(y_train), ', Val labels:', len(y_val), ', Test labels:', len(y_test))
# + id="BZIs6zhrgAAR" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} outputId="13b85fa9-40a2-4904-de83-2962a9502d44" executionInfo={"status": "ok", "timestamp": 1524098632065, "user_tz": 240, "elapsed": 207, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "117867794654112750077"}}
input_shape = x_train[1].shape
print('Input Shape is :', input_shape)
# + id="LCW1WnP_gDTo" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 4607} outputId="d98e0269-a306-4cf2-bd6e-379d61174913" executionInfo={"status": "ok", "timestamp": 1524098637595, "user_tz": 240, "elapsed": 5466, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "117867794654112750077"}}
from keras.layers import MaxPooling2D
from keras.layers import Add
from keras.layers import BatchNormalization
def Pyramidnet(x):
#ResNet1, Number of filters =16
x= Conv2D(16, (3,3), padding='same')(x)
x= BatchNormalization(axis=-1, momentum=0.99, epsilon=0.001)(x)
x= LeakyReLU(alpha=0.15)(x)
x_in = Conv2D(16, (3,3), padding='same')(x)
x_in = BatchNormalization(axis=-1, momentum=0.99, epsilon=0.001)(x_in)
x_in =LeakyReLU(alpha=0.15)(x_in)
x_in = Conv2D(16, (3,3), padding='same')(x_in)
x_out = Add()([x, x_in])
x_out = BatchNormalization(axis=-1, momentum=0.99, epsilon=0.001)(x_out)
x_out = LeakyReLU(alpha=0.15)(x_out)
#ResNet2, Number of filters =32
x= Conv2D(32, (3,3), padding='same')(x_out)
x= BatchNormalization(axis=-1, momentum=0.99, epsilon=0.001)(x)
x= LeakyReLU(alpha=0.15)(x)
x_in = Conv2D(32, (3,3), padding='same')(x)
x_in = BatchNormalization(axis=-1, momentum=0.99, epsilon=0.001)(x_in)
x_in =LeakyReLU(alpha=0.15)(x_in)
x_in = Conv2D(32, (3,3), padding='same')(x_in)
x_out = Add()([x, x_in])
x_out = BatchNormalization(axis=-1, momentum=0.99, epsilon=0.001)(x_out)
x_out = LeakyReLU(alpha=0.15)(x_out)
#ResNet3, Number of filters =48
x= Conv2D(48, (3,3), padding='same')(x_out)
x= BatchNormalization(axis=-1, momentum=0.99, epsilon=0.001)(x)
x= LeakyReLU(alpha=0.15)(x)
x_in = Conv2D(48, (3,3), padding='same')(x)
x_in = BatchNormalization(axis=-1, momentum=0.99, epsilon=0.001)(x_in)
x_in =LeakyReLU(alpha=0.15)(x_in)
x_in = Conv2D(48, (3,3), padding='same')(x_in)
x_out = Add()([x, x_in])
x_out = BatchNormalization(axis=-1, momentum=0.99, epsilon=0.001)(x_out)
x_out = LeakyReLU(alpha=0.15)(x_out)
return x_out
def fire_incept(x, fire=16, intercept=64):
x = Conv2D(fire, (5,5), strides=(2,2))(x)
x = BatchNormalization(axis=-1, momentum=0.99, epsilon=0.001)(x)
x = LeakyReLU(alpha=0.15)(x)
left = Conv2D(intercept, (3,3), padding='same')(x)
left = BatchNormalization(axis=-1, momentum=0.99, epsilon=0.001)(left)
left = LeakyReLU(alpha=0.15)(left)
right = Conv2D(intercept, (5,5), padding='same')(x)
right = BatchNormalization(axis=-1, momentum=0.99, epsilon=0.001)(right)
right = LeakyReLU(alpha=0.15)(right)
x = concatenate([left, right], axis=3)
return x
def fire_squeeze(x, fire=16, intercept=64):
x = Conv2D(fire, (1,1))(x)
x= BatchNormalization(axis=-1, momentum=0.99, epsilon=0.001)(x)
x = LeakyReLU(alpha=0.15)(x)
left = Conv2D(intercept, (1,1))(x)
left = BatchNormalization(axis=-1, momentum=0.99, epsilon=0.001)(left)
left = LeakyReLU(alpha=0.15)(left)
right = Conv2D(intercept, (3,3), padding='same')(x)
right = BatchNormalization(axis=-1, momentum=0.99, epsilon=0.001)(right)
right = LeakyReLU(alpha=0.15)(right)
x = concatenate([left, right], axis=3)
return x
image_input=Input(shape=input_shape)
ip = Pyramidnet(image_input)
ip = MaxPooling2D(pool_size=(2, 2), strides=None, padding='valid', data_format=None)(ip)
ip = Pyramidnet(ip)
ip = MaxPooling2D(pool_size=(2, 2), strides=None, padding='valid', data_format=None)(ip)
ip = Pyramidnet(ip)
ip = fire_incept(ip, fire=32, intercept=32)
ip = fire_squeeze(ip, fire=32, intercept=32)
ip = Conv2D(64, (3,3))(ip)
ip = BatchNormalization(axis=-1, momentum=0.99, epsilon=0.001)(ip)
ip = LeakyReLU(alpha=0.1)(ip)
ip = Flatten()(ip)
ip = Dense(512)(ip)
ip = BatchNormalization(axis=-1, momentum=0.99, epsilon=0.001)(ip)
ip = LeakyReLU(alpha=0.1)(ip)
ip = Dropout(0.5)(ip)
ip = Dense(256)(ip)
ip = BatchNormalization(axis=-1, momentum=0.99, epsilon=0.001)(ip)
ip = LeakyReLU(alpha=0.1)(ip)
ip = Dropout(0.2)(ip)
out = Dense(12, activation='softmax')(ip)
model_new = Model(image_input, out)
model_new.summary()
# + id="iEkd7xmeo3_-" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
model_new.compile(optimizer = Adam(lr=.00025) , loss = 'categorical_crossentropy', metrics=['accuracy'])
# + id="O53JWB0Uo6Jm" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 598} outputId="d5c86d43-89dc-4957-b48f-72c1a8391484" executionInfo={"status": "ok", "timestamp": 1524105239815, "user_tz": 240, "elapsed": 6589855, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "117867794654112750077"}}
# %%time
history = model_new.fit(x_train, y_train,validation_split=0.1, epochs=15, batch_size=25)
# + id="VCY8zTvGK3uZ" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
y_val_pred = model_new.evaluate(x_val, y_val, batch_size=32, verbose=1, sample_weight=None)
print()
print ("Validation Loss = " + str(y_val_pred[0]))
print ("Validation Accuracy = " + str(y_val_pred[1]))
# + id="0W0T9g9upW5e" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 85} outputId="15116b63-fc74-4126-bdb6-c99969c2ed77" executionInfo={"status": "ok", "timestamp": 1524105346759, "user_tz": 240, "elapsed": 16905, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "117867794654112750077"}}
y_test_pred = model_new.evaluate(x_test, y_test, batch_size=32, verbose=1, sample_weight=None)
print()
print ("Test Loss = " + str(y_test_pred[0]))
print ("Test Accuracy = " + str(y_test_pred[1]))
# + id="eC-B_rA5PmS7" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 51} outputId="e9b0c10e-fc3d-45d7-fe05-bef28e145f2c" executionInfo={"status": "ok", "timestamp": 1524105496323, "user_tz": 240, "elapsed": 149537, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "117867794654112750077"}}
y_train_pred = model_new.evaluate(x_train, y_train, batch_size=32, verbose=1, sample_weight=None)
# + id="t4-y1KYRQPbZ" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 51} outputId="62cda3e3-7e01-4ec5-fac9-b71852d0149f" executionInfo={"status": "ok", "timestamp": 1524105496553, "user_tz": 240, "elapsed": 208, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "117867794654112750077"}}
print ("Train Loss = " + str(y_train_pred[0]))
print ("Train Accuracy = " + str(y_train_pred[1]))
# + id="KhEEphc43HDv" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} outputId="83e7bad1-2cd2-4d0d-bda2-eabd36d60bc9" executionInfo={"status": "ok", "timestamp": 1524105522465, "user_tz": 240, "elapsed": 25833, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "117867794654112750077"}}
y_train_pred =model_new.predict(x_train, batch_size=64, verbose=1, steps=None)
y_test_pred =model_new.predict(x_test, batch_size=64, verbose=1, steps=None)
y_val_pred =model_new.predict(x_val, batch_size=64, verbose=1, steps=None)
# + id="da-jK_fqH6ny" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} outputId="3bffc0d7-987c-4626-c93c-90108b139b61" executionInfo={"status": "ok", "timestamp": 1524105673458, "user_tz": 240, "elapsed": 150974, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "117867794654112750077"}}
y_train_pred = np.argmax(y_train_pred, axis=1)
y_test_pred = np.argmax(y_test_pred, axis=1)
y_val_pred = np.argmax(y_val_pred, axis=1)
# + id="b-_TAYKZLZAK" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
y_train_x = np.argmax(y_train, axis=1)
y_test_x = np.argmax(y_test, axis=1)
y_val_x = np.argmax(y_val, axis=1)
# + id="5yLP1J7YLyCf" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
#y_val_pred = np.argmax(y_val_pred, axis=1)
#y_val = np.argmax(y_val, axis=1)
# + id="wz9YDRXHDdEP" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 1047} outputId="f0e12ab0-c6d7-4854-ad7c-a5a018d71e14" executionInfo={"status": "ok", "timestamp": 1522569516872, "user_tz": 240, "elapsed": 1951, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "107126697910026170785"}}
from sklearn.metrics import confusion_matrix
SPECIES = ['Black-grass', 'Charlock', 'Cleavers', 'Common Chickweed', 'Common wheat', 'Fat Hen',
'Loose Silky-bent', 'Maize', 'Scentless Mayweed', 'Shepherds Purse',
'Small-flowered Cranesbill', 'Sugar beet']
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Confusion matrix")
else:
print('Classification Matrix')
print(cm)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
# Compute confusion matrix for Train
cnf_matrix = confusion_matrix(y_train_x, y_train_pred)
np.set_printoptions(precision=2)
# Plot non-normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=SPECIES,
title='Classification matrix')
# Plot normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=SPECIES, normalize=True,
title='Confusion matrix')
plt.show()
# + id="ovS6mDQiEC9d" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 1047} outputId="87463026-5278-4502-d328-179d83a4695a" executionInfo={"status": "ok", "timestamp": 1522570104496, "user_tz": 240, "elapsed": 2147, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "107126697910026170785"}}
# Compute confusion matrix
cnf_matrix = confusion_matrix(y_test_x, y_test_pred)
np.set_printoptions(precision=2)
# Plot non-normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=SPECIES,
title='Confusion matrix')
# Plot normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=SPECIES, normalize=True,
title='Normalized confusion matrix')
plt.show()
# + id="Q1JfK6SHQTUI" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 1047} outputId="10626b8a-2b59-4cca-f3aa-a13d6f42733d" executionInfo={"status": "ok", "timestamp": 1522570195201, "user_tz": 240, "elapsed": 2050, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "107126697910026170785"}}
# Compute confusion matrix
cnf_matrix = confusion_matrix(y_val_x, y_val_pred)
np.set_printoptions(precision=2)
# Plot non-normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=SPECIES,
title='Confusion matrix')
# Plot normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=SPECIES, normalize=True,
title='Normalized confusion matrix')
plt.show()
# + id="UsccW7PpAYtQ" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} outputId="341aa8ca-aa24-42ee-f09d-b4f8f27867d7" executionInfo={"status": "ok", "timestamp": 1522570420395, "user_tz": 240, "elapsed": 340, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "107126697910026170785"}}
print(history.history.keys())
# + id="_AH_r98bAt7r" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 571} outputId="fd25df70-1ed8-4ed4-c7cb-beaba38838f8" executionInfo={"status": "ok", "timestamp": 1522571432625, "user_tz": 240, "elapsed": 572, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "107126697910026170785"}}
from matplotlib import axes as plt2
from matplotlib import pyplot as plt
# summarize history for accuracy
plt.plot(history.history['acc'])
#plt.plot(history.history['val_acc'])
#plt.plot(history.history['loss'])
plt.title('Model accuracy graph')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Accuracy'], loc='upper centre')
plt.show()
# summarize history for loss
plt.plot(history.history['loss'])
#plt.plot(history.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# + id="UnPWzSmqRuCF" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
| Tejas_Plant_classification CODE files/Pyramid_Unit_and_Narrow_wide_Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1><b><font color = black><a id='division_ID0'>GLOBAL TERRORISM ANALYSIS & PREDICTION</a></font></b></h1><br>
# <b>Built by <NAME>, <NAME>, <NAME>, <NAME>, <NAME></b>
# <br><b>Guidance - Indranil Das, Technical Analyst, Webskitters Academy</b>
# <br><b>In Association with Webskitters Academy</b>
# <br><b>Hooghly Engineering & Technology College</b>
#
# <p><b>This data is collected from - <a href = "https://gtd.terrorismdata.com/register-type/?type=non-commercial">Global Terrorism Database</a></b></p>
#
# <p><b>Global Terrorism Code Book - <a href = "https://www.start.umd.edu/gtd/downloads/Codebook.pdf">Click Here</a></b></p>
#
# <p>The Global Terrorism Database (GTD) is an open-source database including information on domestic and international terrorist attacks around the world from 1970 through 2019, and now includes more than 200,000 cases. For each event, information is available on the date and location of the incident, the weapons used and nature of the target, the number of casualties, and–when identifiable–the group or individual responsible.<br>
# The National Consortium for the Study of Terrorism and Responses to Terrorism (START) makes the GTD available via this online interface in an effort to increase understanding of terrorist violence so that it can be more readily studied and defeated.</p>
#
# <b>Charactertistics</b>
# - Contains information on over 200,000 terrorist attacks
# - Currently the most comprehensive unclassified database on terrorist attacks in the world
# - Includes information on more than 95,000 bombings, 20,000 assassinations, and 15,000 kidnappings and hostage events since 1970
# - Includes information on at least 45 variables for each case, with more recent incidents including information on more than 120 variables
# - More than 4,000,000 news articles and 25,000 news sources were reviewed to collect incident data from 1998 to 2019 alone
#
# <h3><b>Contents</b></h3>
#
# [Data Collection](#division_ID1) | [Data Preprocessing](#division_ID2) | [Feature Extraction](#division_ID3) | [Data Visualization](#division_ID4) | [Model Training](#division_ID5)| [Model Evaluation](#division_ID6) | [Model Testing](#division_ID7) | [Model Saving](#division_ID8) | [User Interactive](#division_ID9)
# <h2><a id='division_ID1'>Data Collection</a></h2>
#
# [MENU](#division_ID0)
import pandas as pd
df = pd.read_csv('global_terrorism.csv', encoding = 'ISO-8859-1', low_memory=False)
df
# <h2><a id='division_ID2'>Data Preprocessing</a></h2>
#
# [MENU](#division_ID0)
# check the null values for every column
j = 0
data1 = []
data2 = []
data3 = []
data4 = []
for i in df.columns:
data1.append(j)
data2.append(i)
data3.append(df[i].isnull().sum())
data4.append((df[i].isnull().sum()/len(df))*100)
j = j + 1
data = list(zip(data1, data2, data3, data4))
col_stats = pd.DataFrame(data, columns = ['Column Index', 'Column Name', 'Null Values', 'NA %'])
col_stats
# plot the null value statistics from the columns
import matplotlib.pyplot as plt
x = col_stats[col_stats.iloc[:, 2] != 0].iloc[:, 1]
y = col_stats[col_stats.iloc[:, 2] != 0].iloc[:, 2]
plt.figure(figsize = (20, 30))
plt.barh(x, y, .3)
plt.title('Statistics of Null Value Contating Columns', bbox={'facecolor':'0.8', 'pad':5}, loc = 'center')
plt.ylabel('Column Names')
plt.xlabel('No. of Null Values')
plt.show()
# find the groups those who did 5 or more attacks
groups = list(df['gname'].unique())
print("Discarded Group Names:")
for i in groups:
if len(df[df['gname'] == i]) < 5:
print(i)
df = df[df['gname'] != i].copy()
df
# label encode countries
from sklearn.preprocessing import LabelEncoder
enc = LabelEncoder()
df['country'] = enc.fit_transform(df['country'])
# create a dictionary of country names from the dataset
country_dict = {}
for i in range(len(df)):
country_dict.update({df.iloc[i, list(df.columns).index('country')]: df.iloc[i, list(df.columns).index('country_txt')]})
# label encode gnames
gnames = list(df['gname'].copy())
df['gname'] = enc.fit_transform(df['gname'])
# create a dictionary of gnames from the dataset
gname_dict = {}
for i in range(len(gnames)):
gname_dict.update({df.iloc[i, list(df.columns).index('gname')]: gnames[i]})
# <h2><a id='division_ID3'>Feature Extraction</a></h2>
#
# [MENU](#division_ID0)
# filter most nessecary columns
terror = df[['iyear', 'crit1', 'crit2', 'crit3', 'doubtterr', 'country', 'attacktype1',
'weaptype1', 'targtype1', 'gname', 'individual']].copy()
terror
# check null values in the existing dataframe
terror.isnull().sum()
# drop all null values in the existing dataframe
terror.dropna(inplace = True)
# change the columns datatypes into integer
for i in terror.columns:
terror[i] = terror[i].astype(int)
# <h2><a id='division_ID4'>Data Visualization</a></h2>
#
# [MENU](#division_ID0)
# lets have a look on dataset
terror
# year wise terrorist attack plot
import matplotlib.pyplot as plt
import seaborn as sns
sns.color_palette("rocket", as_cmap=True)
sns.set_style('dark')
timeline = terror['iyear'].unique()
attacks = []
for i in timeline:
attacks.append(len(terror[terror['iyear'] == i]))
plt.figure(figsize = (16, 9))
plt.title("1970 - 2018 Terrorist Attacks per year", bbox={'facecolor':'0.8', 'pad':5}, loc = 'center')
sns.barplot(y = attacks, x = timeline, palette = 'magma')
plt.xticks(rotation = 90)
plt.xlabel('Year Timeline')
plt.ylabel('No. of Terrorist Attacks')
plt.show()
# top 20 countries where terrorist attacks happened in last 50 years
areas = terror['country'].unique()
con_attacks = []
for i in areas:
con_attacks.append(len(terror[terror.iloc[:, list(terror.columns).index('country')] == i]))
for i in range(len(con_attacks) - 1):
for j in range(len(con_attacks) - i - 1):
if con_attacks[j] <= con_attacks[j+1]:
temp = con_attacks[j]
con_attacks[j] = con_attacks[j+1]
con_attacks[j+1] = temp
temp = areas[j]
areas[j] = areas[j+1]
areas[j+1] = temp
con_attacks = con_attacks[: 20]
areas = areas[: 20]
print(con_attacks)
print(areas)
# change the country numbers with country names from the GTD codebook
country = ['Iraq', 'Pakistan', 'Afghanistan', 'India', 'Colombia', 'Philippines', 'Peru', 'El Salvador', 'United Kingdom',
'Turkey', 'Somalia', 'Nigeria', 'Thailand', 'Yemen', 'Spain', 'Sri Lanka', 'Algeria', 'United States', 'France', 'Egypt']
sns.color_palette("flare", as_cmap=True)
sns.set_style("darkgrid")
plt.figure(figsize = (16, 9))
plt.title("Top 20 countries in Terrorist Attack", bbox={'facecolor':'0.8', 'pad':5}, loc = 'center')
sns.barplot(x = con_attacks, y = country)
plt.xlabel("No. of Terrorist Attacks")
plt.show()
# top 20 terrorist groups
terrorist_groups = terror['gname'].unique()
top_groups = []
for i in terrorist_groups:
top_groups.append(len(terror[terror.iloc[:, list(terror.columns).index('gname')] == i]))
for i in range(len(top_groups) - 1):
for j in range(len(top_groups) - i - 1):
if top_groups[j] <= top_groups[j+1]:
temp = top_groups[j]
top_groups[j] = top_groups[j+1]
top_groups[j+1] = temp
temp = terrorist_groups[j]
terrorist_groups[j] = terrorist_groups[j+1]
terrorist_groups[j+1] = temp
terrorist_groups = terrorist_groups[1: 21]
top_groups = top_groups[1: 21]
gnames = []
for i in terrorist_groups:
gnames.append(gname_dict[i])
print(top_groups)
print(gnames)
# plot no. of attacks by top 20 terrorist organizations
plt.figure(figsize = (16, 9))
plt.title("Top 20 Terrorist Organizations by attacks", bbox={'facecolor':'0.8', 'pad':5}, loc = 'center')
sns.barplot(x = top_groups, y = gnames, palette = "crest")
plt.xlabel("No. of Attacks")
plt.show()
# +
# terrorist attack target types
types = list(terror['targtype1'].unique())
freq = []
for i in types:
freq.append(len(terror[terror.iloc[:, list(terror.columns).index('targtype1')] == i]))
type_names = {
1: 'Business',
2: 'Government (General)',
3: 'Police',
4: 'Military',
5: 'Abortion related',
6: 'Airport & Aircraft',
7: 'Government (Diplomatic)',
8: 'Educational Institution',
9: 'Food or Water Supply',
10: 'Journalists & Media',
11: 'Maritime',
12: 'NGO',
13: 'Other',
14: 'Private Citizens & Property',
15: 'Religious Figures / Institutions',
16: 'Telecommunication',
17: 'Terrorists / Non State Militias',
18: 'Tourists',
19: 'Transportation',
20: 'Unknown',
21: 'Utilities',
22: 'Violent Political Parties'
}
for i in types:
types[types.index(i)] = type_names[i]
colors = ['lightseagreen', 'gold', 'red', 'green', 'blue', 'violet', 'yellow', 'slateblue', 'indigo', 'plum',
'darkslategray', 'tomato', 'olive', 'paleturquoise', 'darkorange', 'crimson', 'cyan', 'purple', 'powderblue', 'chartreuse']
plt.figure(figsize = (16, 9))
plt.title('Target types of Terrorist Attacks', bbox={'facecolor':'0.8', 'pad':10}, loc = 'center')
plt.pie(freq, startangle=90, colors = colors)
my_circle=plt.Circle( (0,0), 0.7, color='white')
p=plt.gcf()
p.gca().add_artist(my_circle)
plt.legend(types, loc='best', bbox_to_anchor=(-0.1, 1.), fontsize=15)
plt.show()
# +
# terrorist weapon types
weaps = list(terror['weaptype1'].unique())
freq = []
for i in weaps:
freq.append(len(terror[terror.iloc[:, list(terror.columns).index('weaptype1')] == i]))
type_weaps = {
1: 'Biological',
2: 'Chemical',
3: 'Radiological',
4: 'Nuclear',
5: 'Firearms',
6: 'Explosives',
7: 'Fake Weapons',
8: 'Incendiary',
9: 'Melee',
10: 'Vehicle',
11: 'Sabotage Equipment',
12: 'Other',
13: 'Unknown'
}
for i in weaps:
weaps[weaps.index(i)] = type_weaps[i]
colors = ['lightseagreen', 'gold', 'red', 'green', 'blue', 'violet', 'yellow', 'slateblue', 'indigo', 'plum',
'darkslategray', 'tomato', 'olive']
plt.figure(figsize = (14, 8))
plt.title('Weapons used in Terrorist Attacks', bbox={'facecolor':'0.8', 'pad':10}, loc = 'center')
plt.pie(freq, startangle=90, colors = colors)
my_circle=plt.Circle( (0,0), 0.7, color='white')
p=plt.gcf()
p.gca().add_artist(my_circle)
plt.legend(weaps, loc='best', bbox_to_anchor=(-0.1, 1.), fontsize=20)
plt.show()
# +
# terrorist attack types
attacks = list(terror['attacktype1'].unique())
freq = []
for i in attacks:
freq.append(len(terror[terror.iloc[:, list(terror.columns).index('attacktype1')] == i]))
type_attacks = {
1: 'Assasination',
2: 'Armed Assault',
3: 'Bombing / Explosion',
4: 'Hijacking',
5: 'Hostage (Barricade)',
6: 'Hostage (Kidnap)',
7: 'Facility Attack',
8: 'Unarmed Assault',
9: 'Unknown'
}
for i in attacks:
attacks[attacks.index(i)] = type_attacks[i]
colors = ['lightseagreen', 'gold', 'red', 'green', 'blue', 'violet', 'yellow', 'slateblue', 'indigo']
plt.figure(figsize = (12, 7))
plt.title('Attack Types in various Terrorist Attacks', bbox={'facecolor':'0.8', 'pad':10}, loc = 'center')
plt.pie(freq, colors = colors)
my_circle=plt.Circle( (0,0), 0.7, color='white')
p=plt.gcf()
p.gca().add_artist(my_circle)
plt.legend(attacks, loc='best', bbox_to_anchor=(-0.1, 1.), fontsize=25)
plt.show()
# +
# plot terrorist attack hot zone
import geopandas
from descartes import PolygonPatch
def plotCountryPatch( axes, country_name, fcolor ):
# plot a country on the provided axes
nami = world[world.name == country_name]
namigm = nami.__geo_interface__['features'] # geopandas's geo_interface
namig0 = {'type': namigm[0]['geometry']['type'], \
'coordinates': namigm[0]['geometry']['coordinates']}
axes.add_patch(PolygonPatch( namig0, fc=fcolor, ec="black", alpha=0.85, zorder=2 ))
world = geopandas.read_file(geopandas.datasets.get_path('naturalearth_lowres'))
world = world[world.continent != 'Antarctica']
cities = geopandas.read_file(geopandas.datasets.get_path('naturalearth_cities'))
fig, ax = plt.subplots(1, 1, figsize=(16, 12))
ax.set_title("Terrorist Attack Hot Zone", bbox={'facecolor':'0.8', 'pad':10}, loc = 'center')
country = ['Iraq', 'Pakistan', 'India', 'Afghanistan', 'Colombia', 'Philippines', 'Peru', 'United Kingdom', 'Turkey', 'Somalia',
'Kenya', 'Nigeria', 'Thailand', 'Yemen', 'Spain', 'Sri Lanka', 'Algeria', 'United States of America', 'France', 'Egypt']
for i in range(len(country)):
plotCountryPatch(ax, country[i], 'red')
stats = pd.DataFrame(con_attacks, columns = ['attacks'])
world.boundary.plot(ax = ax)
# -
# <h2><a id='division_ID5'>Model Training</a></h2>
#
# [MENU](#division_ID0)
# extract dependent and independent features
x1 = terror.drop(['country'], axis = 1)
y1 = terror.iloc[:, 5]
x2 = terror.drop(['gname'], axis = 1)
y2 = terror.iloc[:, -2]
x3 = terror.drop(['attacktype1'], axis = 1)
y3 = terror.iloc[:, 6]
x4 = terror.drop(['weaptype1'], axis = 1)
y4 = terror.iloc[:, 7]
x5 = terror.drop(['targtype1'], axis = 1)
y5 = terror.iloc[:, -3]
# apply train test split
from sklearn.model_selection import train_test_split
x1_train, x1_test, y1_train, y1_test = train_test_split(x1, y1, test_size = 0.2, random_state = 60) # for country
x2_train, x2_test, y2_train, y2_test = train_test_split(x2, y2, test_size = 0.2, random_state = 60) # for gnames
x3_train, x3_test, y3_train, y3_test = train_test_split(x3, y3, test_size = 0.2, random_state = 60) # for attacktype
x4_train, x4_test, y4_train, y4_test = train_test_split(x4, y4, test_size = 0.2, random_state = 60) # for weapontype
x5_train, x5_test, y5_train, y5_test = train_test_split(x5, y5, test_size = 0.2, random_state = 60) # for targettype
# import nessecary algorithms
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier, BaggingClassifier, AdaBoostClassifier, ExtraTreesClassifier
# - <h3><b>Prediction Model for Target Country</b></h3>
# apply decision tree classifier
dtc_model1 = DecisionTreeClassifier()
dtc_model1.fit(x1_train, y1_train)
dtc_model1_pred = dtc_model1.predict(x1_test)
m1s1 = dtc_model1.score(x1_test, y1_test)
# apply random forest classifier
rfc_model1 = RandomForestClassifier(n_estimators = 10)
rfc_model1.fit(x1_train, y1_train)
rfc_model1_pred = rfc_model1.predict(x1_test)
m1s2 = rfc_model1.score(x1_test, y1_test)
# - <h3><b>Prediction Model for Terrorist Group Name</b></h3>
# apply decision tree classifier
dtc_model2 = DecisionTreeClassifier()
dtc_model2.fit(x2_train, y2_train)
dtc_model2_pred = dtc_model2.predict(x2_test)
m2s1 = dtc_model2.score(x2_test, y2_test)
# apply random forest classifier
rfc_model2 = RandomForestClassifier(n_estimators = 10)
rfc_model2.fit(x2_train, y2_train)
rfc_model2_pred = rfc_model2.predict(x2_test)
m2s2 = rfc_model2.score(x2_test, y2_test)
# - <h3><b>Prediction Model for Terrorist Attack Type</b></h3>
# apply decision tree classifier
dtc_model3 = DecisionTreeClassifier()
dtc_model3.fit(x3_train, y3_train)
dtc_model3_pred = dtc_model3.predict(x3_test)
m3s1 = dtc_model3.score(x3_test, y3_test)
# apply random forest classifier
rfc_model3 = RandomForestClassifier(n_estimators = 10)
rfc_model3.fit(x3_train, y3_train)
rfc_model3_pred = rfc_model3.predict(x3_test)
m3s2 = rfc_model3.score(x3_test, y3_test)
# - <h3><b>Prediction Model for Terrorist Attack Weapon Type</b></h3>
# apply decision tree classifier
dtc_model4 = DecisionTreeClassifier()
dtc_model4.fit(x4_train, y4_train)
dtc_model4_pred = dtc_model4.predict(x4_test)
m4s1 = dtc_model4.score(x4_test, y4_test)
# apply random forest classifier
rfc_model4 = RandomForestClassifier(n_estimators = 10)
rfc_model4.fit(x4_train, y4_train)
rfc_model4_pred = rfc_model4.predict(x4_test)
m4s2 = rfc_model4.score(x4_test, y4_test)
# - <h3><b>Prediction Model for Terrorist Attack Target Type</b></h3>
# apply decision tree classifier
dtc_model5 = DecisionTreeClassifier()
dtc_model5.fit(x5_train, y5_train)
dtc_model5_pred = dtc_model5.predict(x5_test)
m5s1 = dtc_model5.score(x5_test, y5_test)
# apply random forest classifier
rfc_model5 = RandomForestClassifier(n_estimators = 10)
rfc_model5.fit(x5_train, y5_train)
rfc_model5_pred = rfc_model5.predict(x5_test)
m5s2 = rfc_model5.score(x5_test, y5_test)
# <h2><a id='division_ID6'>Model Evaluation</a></h2>
#
# [MENU](#division_ID0)
# +
# plot a bar graph to check scores of every model
import numpy as np
from matplotlib import style
style.use('classic')
r = np.arange(5)
w = .2
algo = ['Decision Tree Classifier', 'Random Forest Classifier']
plt.figure(figsize = (16, 9))
models = ['Country\nPrediction\nModel', 'Terrorist\nGroup Name\nPrediction\nModel',
'Terrorist\nAttack Type\nPrediction\nModel', 'Attack\nWeapon Type\nPrediction Model',
'Attack\nTarget Type\nPrediction Model']
plt.bar(r, [m1s1, m2s1, m3s1, m4s1, m5s1], width = w)
plt.bar(r+w, [m1s2, m2s2, m3s2, m4s2, m5s2], width = w)
plt.xticks(r+w/2, models)
plt.ylim(0, 1)
plt.ylabel('Model Score')
plt.title("Model Score Study for Decision Tree and Random Forest Classifier", bbox={'facecolor':'0.8', 'pad':10}, loc = 'center')
plt.legend(algo, loc = 2)
# -
# - <h3><b>Evaluation of Terrorist Attack Country Prediction Model</b></h3>
# classification report
from sklearn.metrics import classification_report
print(classification_report(y1_test, dtc_model1_pred, zero_division = 1))
# confusion matrix
from sklearn.metrics import confusion_matrix
confusion_matrix(y1_test, dtc_model1_pred)
# - <h3><b>Evaluation of Terrorist Group Name Prediction Model</b></h3>
# classification report
print(classification_report(y2_test, rfc_model2_pred, zero_division = 1))
# confusion matrix
confusion_matrix(y2_test, rfc_model2_pred)
# - <h3><b>Evaluation of Terrorist Attack Type Prediction Model</b></h3>
# classification report
print(classification_report(y3_test, rfc_model3_pred, zero_division = 1))
# confusion matrix
confusion_matrix(y3_test, rfc_model3_pred)
# - <h3><b>Evaluation of Terrorist Attack Weapon Type Prediction Model</b></h3>
# classification report
print(classification_report(y4_test, rfc_model4_pred, zero_division = 1))
# confusion matrix
confusion_matrix(y4_test, rfc_model4_pred)
# - <h3><b>Evaluation of Terrorist Attack Target Type Prediction Model</b></h3>
# classification report
print(classification_report(y5_test, rfc_model5_pred, zero_division = 1))
# confusion matrix
confusion_matrix(y5_test, rfc_model5_pred)
# <h2><a id='division_ID7'>Model Testing</a></h2>
#
# [MENU](#division_ID0)
iyear = 2012
crit1 = 1
crit2 = 1
crit3 = 1
doubtterr = 0
country = 92
attacktype1 = 3
weaptype1 = 6
targtype1 = 4
gname = 911
individual = 0
# - <h3><b>Customized I/O for Terrorist Attack Country Prediction Model</b></h3>
val = dtc_model1.predict([[iyear, crit1, crit2, crit3, doubtterr, attacktype1, weaptype1, targtype1, gname, individual]])
val = int(val[0])
if val in country_dict.keys():
print("Country Name: ", country_dict[val])
else:
print("Country Name: Unknown")
# - <h3><b>Customized I/O for Terrorist Group Name Prediction Model</b></h3>
val = rfc_model2.predict([[iyear, crit1, crit2, crit3, doubtterr, country, attacktype1, weaptype1, targtype1, individual]])
val = int(val[0])
if val in gname_dict.keys():
print("Group Name: ", gname_dict[val])
else:
print("Group Name: Unknown")
# - <h3><b>Customized I/O for Terrorist Attack Type Prediction Model</b></h3>
val = rfc_model3.predict([[iyear, crit1, crit2, crit3, doubtterr, country, weaptype1, targtype1, gname, individual]])
val = int(val[0])
if val in type_attacks.keys():
print("Attack Type: ", type_attacks[val])
else:
print("Attack Type: Unknown")
# - <h3><b>Customized I/O for Terrorist Attack Weapon Type Prediction Model</b></h3>
val = rfc_model4.predict([[iyear, crit1, crit2, crit3, doubtterr, country, attacktype1, targtype1, gname, individual]])
val = int(val[0])
if val in type_weaps.keys():
print("Weapon Type: ", type_weaps[val])
else:
print("Weapon Type: Unknown")
# - <h3><b>Customized I/O for Terrorist Attack Target Type Prediction Model</b></h3>
val = rfc_model4.predict([[iyear, crit1, crit2, crit3, doubtterr, country, weaptype1, attacktype1, gname, individual]])
val = int(val[0])
if val in type_names.keys():
print("Target Type: ", type_names[val])
else:
print("Target Type: Unknown")
# <h2><a id='division_ID8'>Model Saving</a></h2>
#
# [MENU](#division_ID0)
# save these 4 models for future prediction
import joblib
joblib.dump(dtc_model1, 'GT_Country_Prediction')
joblib.dump(rfc_model2, 'GT_Gname_Prediction')
joblib.dump(rfc_model3, 'GT_Attacktype_Prediction')
joblib.dump(rfc_model4, 'GT_Weapontype_Prediction')
joblib.dump(rfc_model5, 'GT_Targettype_Prediction')
# <h2><a id='division_ID9'>User Interactive</a></h2>
#
# [MENU](#division_ID0)
# +
def usr_choice(choice):
arr = []
arr.append(int(input('Year: ')))
arr.append(int(input('Criteria 1 [0/1]: ')))
arr.append(int(input('Criteria 2 [0/1]: ')))
arr.append(int(input('Criteria 3 [0/1]: ')))
arr.append(int(input('Doubt [0/1]: ')))
if choice != 1:
arr.append(int(input('Country [Only Country Code is accepted]: ')))
if choice != 2:
arr.append(int(input('Terrorist Group [Only Group Code is accepted]: ')))
if choice != 3:
arr.append(int(input('Attack Type [Only Attack Code is accepted]: ')))
if choice != 4:
arr.append(int(input('Weapon Type [Only Weapon Code is accepted]: ')))
if choice != 5:
arr.append(int(input('Target Type [Only Target Code is accepted]: ')))
arr.append(int(input('Individual [0/1]: ')))
return arr
while True:
print('<<< GLOBAL TERRORISM PREDICTION PROJECT >>>')
print('Press 0 ---> Exit')
print('Press 1 ---> Predict Target Country')
print('Press 2 ---> Predict Terrorist Group Name')
print('Press 3 ---> Predict Terrorist Attack Type')
print('Press 4 ---> Predict Terrorist Weapon Type')
print('Press 5 ---> Predict Terrorist Attack Target Type')
ch = int(input('Enter your choice: '))
if ch == 0:
print('UI is closed!')
break
elif ch == 1:
model = joblib.load('GT_Country_Prediction')
val = model.predict([usr_choice(ch)])
val = int(val[0])
if val in country_dict.keys():
print("Country Name: ", country_dict[val])
else:
print("Country Name: Unknown")
elif ch == 2:
model = joblib.load('GT_Gname_Prediction')
val = model.predict([usr_choice(ch)])
val = int(val[0])
if val in gname_dict.keys():
print("Group Name: ", gname_dict[val])
else:
print("Group Name: Unknown")
elif ch == 3:
model = joblib.load('GT_Attacktype_Prediction')
val = model.predict([usr_choice(ch)])
val = int(val[0])
if val in type_attacks.keys():
print("Attack Type: ", type_attacks[val])
else:
print("Attack Type: Unknown")
elif ch == 4:
model = joblib.load('GT_Weapontype_Prediction')
val = model.predict([usr_choice(ch)])
val = int(val[0])
if val in type_weaps.keys():
print("Weapon Type: ", type_weaps[val])
else:
print("Weapon Type: Unknown")
elif ch == 5:
model = joblib.load('GT_Targettype_Prediction')
val = model.predict([usr_choice(ch)])
val = int(val[0])
if val in type_names.keys():
print("Target Type: ", type_names[val])
else:
print("Target Type: Unknown")
else:
print('You select a wrong choice :(')
print('----------------------------------------------------------------------')
# -
# <center>For further information or query mail us at <a href = "mailto: <EMAIL>"><EMAIL></a></center>
#
#
# <br><center>© All rights reserved by Webskitters Academy</center>
| Project Global Terrorism Prediction/PROJECT_GTP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### To test:
# 1. Create a folder ../data/luna16/
# 2. Create a folder ../data/luna16/subset2
# -Under this folder copy one scan for testing (script will process all the scan at this location)
# 1.3.6.1.4.1.14519.5.2.1.6279.6001.100621383016233746780170740405.mhd & raw file
# (Google drive https://drive.google.com/drive/u/1/folders/13wmubTgm-7sh3MxPGxqmVZuoqi0G3ufW
# 3. Create a folder ../data/luna16/hdf5
# -Under this copy UNET_weights_H2.h5 (download from google drive)
# +
import pandas as pd
import numpy as np
import h5py
import pandas as pd
import argparse
import SimpleITK as sitk
from PIL import Image
import os, glob
import os, os.path
import tensorflow as tf
import keras
from ipywidgets import interact
import json
import pickle
from datetime import datetime
from tqdm import tqdm, trange
from UNET_utils import *
# %matplotlib inline
# +
# import argparse
# parser = argparse.ArgumentParser(description='Prediction on HOLDOUT subset',add_help=True)
# parser.add_argument("--holdout", type=int, default=0, help="HOLDOUT subset for predictions")
# args = parser.parse_args()
# HOLDOUT = args.holdout
# -
HOLDOUT = 5
HO_dir = 'HO{}/'.format(HOLDOUT)
data_dir = '/home/tony/data/luna16/'
model_wghts = 'hdf5/UNET_weights_modelB_H{}.h5'.format(HOLDOUT)
TILE_HSIZE = 64
TILE_WSIZE = 64
TILE_DSIZE = 64
def model_create_loadWghts_Model_B():
input_shape=(None, None, None, 1)
model = unet3D_modelB(input_shape, use_upsampling=True)
model.load_weights(data_dir + model_wghts)
return model
def model_create_loadWghts_Model_A():
input_shape=(None, None, None, 1)
model = create_unet3D_Model_A(input_shape, use_upsampling=True)
model.load_weights(data_dir + model_wghts)
return model
def model_create_loadWghts():
input_shape=(None, None, None, 1)
model = create_UNET3D(input_shape, use_upsampling=True)
model.load_weights(data_dir + model_wghts)
return model
# +
def find_mask(model, img):
height, width, depth = img.shape
# Need to do ceiling to get the right prediction mask size
# This way prediction mask will be a multiple of the tile size.
# So just need to pad whole image to that multiple.
# At the end we crop back to the original image size.
pred_height = int(TILE_HSIZE * np.ceil(1.0*height/TILE_HSIZE))
pred_width = int(TILE_WSIZE * np.ceil(1.0*width/TILE_WSIZE))
pred_depth = int(TILE_DSIZE * np.ceil(1.0*depth/TILE_DSIZE))
# Prediction mask is now a multiple of the TILE_SIZE.
prediction_mask = np.zeros((pred_height, pred_width, pred_depth, 1))
idxH = 0
for startH_idx in range(0, height, TILE_HSIZE):
stopH_idx = startH_idx + TILE_HSIZE
if stopH_idx > height:
stopH_idx = height
idxW = 0
for startW_idx in range(0, width, TILE_WSIZE):
stopW_idx = startW_idx + TILE_WSIZE
if stopW_idx > width:
stopW_idx = width
idxD = 0
for startD_idx in range(0, depth, TILE_DSIZE):
stopD_idx = startD_idx + TILE_DSIZE
if stopD_idx > depth:
stopD_idx = depth
snippet = img[startH_idx:stopH_idx, startW_idx:stopW_idx, startD_idx:stopD_idx]
tile = np.zeros([TILE_HSIZE,TILE_WSIZE,TILE_DSIZE])
snippet_width = stopW_idx - startW_idx
snippet_height = stopH_idx - startH_idx
snippet_depth = stopD_idx - startD_idx
tile[:snippet_height,:snippet_width,:snippet_depth] = snippet
tile = np.expand_dims(tile, 0)
tile = np.expand_dims(tile, -1)
tile_mask = model.predict(tile, verbose=0)
prediction_mask[idxH:(idxH+TILE_HSIZE), idxW:(idxW+TILE_WSIZE), idxD:(idxD+TILE_DSIZE), :] = tile_mask[0]
idxD += TILE_DSIZE
idxW += TILE_WSIZE
idxH += TILE_HSIZE
return prediction_mask[:height, :width, :depth, :] # Truncate to original image size
# +
# %%time
t0 = datetime.now()
predictions_dict = {}
size_dict = {}
model = model_create_loadWghts_Model_B()
fileCount = len(glob.glob(data_dir + 'subset5/' + '*.mhd'))
for f in tqdm(glob.glob(data_dir + 'subset5/' + '*.mhd'), total=fileCount, unit="files") :
print ("\n Processing scan file: {}".format(os.path.basename(f)))
seriesuid = os.path.splitext(os.path.basename(f))[0]
# Step-1
itk_img = sitk.ReadImage(f)
img_np_array = sitk.GetArrayFromImage(itk_img)
original_size = img_np_array.shape
print ("Original-Size of loaded image : {}".format(original_size))
# Step-2
itk_img_norm = normalize_img(itk_img)
img_np_array_norm = sitk.GetArrayFromImage(itk_img_norm)
normalized_size = img_np_array_norm.shape
# Step-3
img = img_np_array_norm.copy()
# img = normalize_HU(img_np_array_norm)
img = np.swapaxes(img, 0,2) ##needed as SITK swaps axis
print ("Normalized input image size: {}".format(img.shape))
predicted_mask = find_mask(model, img)
predictions_dict[seriesuid] = (img.shape, img, predicted_mask)
size_dict[seriesuid] = img.shape
print('Predicted Mask sum for entire scan: {}'.format(np.sum(predicted_mask)))
pickle.dump(predictions_dict, open('Model_B_noHU_entire_predictions_{}.dat'.format(seriesuid), 'wb'))
pickle.dump(size_dict, open('Model_B_noHU_entire_size_{}.dat'.format(seriesuid), 'wb'))
print('Processing runtime: {}'.format(datetime.now() - t0))
# +
def displaySlice(sliceNo):
plt.figure(figsize=[10,10]);
plt.subplot(1,2,1)
plt.title("Predicted Mask")
plt.imshow(np.round(predicted_mask[:, :, sliceNo,0]), cmap='bone');
plt.subplot(1,2,2)
plt.title("Overlay Mask")
plt.imshow(img[:, :, sliceNo], cmap="bone");
plt.imshow(predicted_mask[:, :, sliceNo,0]>0.01, alpha=0.5, cmap='Reds');
plt.show()
interact(displaySlice, sliceNo=(1,img.shape[2],1)); #172
# -
# ###### Following sections for reference & WIP code snippets -AL
# +
## Multiple tile test....performance hog, so exploiting the GPU for entire slice without compromising predictions
##and for better performance -AL
# slices = 16
# predicted_img = np.zeros(padded_size)
# for i in range(368//slices):
# tile_1 = padded_img[:224, :224, (i*slices) : slices*(i+1)]
# tile_2 = padded_img[224:, 224:, (i*slices) : slices*(i+1) ]
# +
# slices = 8
# predicted_mask = np.zeros(PADDED_SIZE)
# for i in range(24//SLICES):
# tile = padded_img[:, :, (i*SLICES) : SLICES*(i+1)]
# tile = tile.reshape(tuple([1] + list (tile.shape) + [1]))
# # print(tile.shape)
# tile_predictions = model.predict(tile, verbose=2)
# tile_mask = tile_predictions[0].reshape(448, 448, 8)
# print (tile_mask.shape)
# predicted_mask[:, :, (i*SLICES) : SLICES*(i+1)] = tile_mask
# +
# slices = 8
# test_slice = padded_img[:, :, :slices]
# print(test_slice.shape)
# model = model_create_loadWghts(test_slice.shape)
# # slice_predictions = model.predict(test_slice, verbose=2)
# +
# print ("Shape of predicted mask or segmented image : {}".format(predictions_small_img[0].shape))
# print ("Shape of predicted class : {}".format(predictions_small_img[1].shape))
# predictions_small_img[0] [:, 25 : 26, :]
# +
# ## AL - TEST : making an image of size 48,48,48 with random 0 or 1
# ### Case 2 : As a test created an input image of size (1, 48,48,48,1)
# # with random 0 or 1; this works fine and able to create predictions successfully
# t2 = np.random.choice(2,(48,48,48))
# t2 = t2.reshape(tuple([1] + list (t2.shape) + [1]))
# print ("Shape of test input image : {}".format(t2.shape))
# predictions = model.predict(t2, verbose=2)
# print ("Shape of predicted mask or segmented image : {}".format(predictions[0].shape))
# print ("Shape of predicted class : {}".format(predictions[1].shape))
# # predictions[0] [:, 25 : 26, :]
# +
# padded_img[225:232, 225:232, 175]
# predicted_mask[225:232, 225:232, 175]
| src/predictions/UNET_Prediction_EntireScan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + tags=["hide-cell"]
import numpy as np
import matplotlib.pyplot as plt
from scipy.integrate import odeint
plt.style.use('fivethirtyeight')
import numpy as np
import matplotlib.pyplot as plt
from scipy.optimize import fsolve
plt.style.use('fivethirtyeight')
from matplotlib import rc
plt.rc('text', usetex=True)
plt.rc('font', family='sans')
# -
# # Module 05 - project _build an example_
#
# This is your chance to build something important to you and/or your career. If you're interested in cars you can create a drivetrain model. If you want to get into biomechanics you can model an orthopedic joint or build a 2D model of someone walking. If you love toys (_I know Prof. Cooper does!_) you can model a yoyo spinning on a cord or a spinning top or measure the coefficient of restitution of a bouncy ball on concrete.
#
# Some tips:
# - __start simple__ its often easier to _add_ complexity rather than _remove_
# complexity if you're considering a 6-DOF system, try a 1-DOF part, then add
# components incrementally
# - __use what you have__ you have working models for four-bar linkages,
# pendulums, and a yoyo despinning mechanism can you take your system and model
# a piece of it as a pendulum? is angular momentum conserved?
# - __communicate__ let us know what you're working on simple sketches are great!
# don't get hung up on a final result until you have some sketches and
# conversations
# - __always draw a FBD__ this step is _so often_ skipped, but its really where
# the bulk of your engineering work is decided. The FBD is your main
# contribution to any engineering project, everything else is a combination of
# puzzle-solving and applied mathematics
#
l1 = 0.25 #m length of my forearm
l2 = 0.3 #m length of my bicep and shoulder
l3 = 0.55 #m length of both my forearm and bicep and shoulder
a1 = np.pi/2
dy = 0
dx = 0.41 #the measured distance between my two shoulders, meaning l1 and l3
Fbar = lambda a1,x: np.array([l1*np.sin(a1)+l2*np.sin(x[0])-l3*np.sin(x[1])-dy,
l1*np.cos(a1)+l2*np.cos(x[0])-l3*np.cos(x[1])-dx])
a1 = np.linspace(0, 2*np.pi)
a2 = np.zeros(len(a1))
a3 = np.zeros(len(a1))
xsol = np.array([0, np.pi/4])
for i in range(len(a1)):
xsol = fsolve(lambda x: Fbar(a1[i], x), xsol)
a2[i] = xsol[0]
a3[i] = xsol[1]
plt.plot(a1, a2, label = r'$\theta_2$')
plt.plot(a1, a3, label = r'$\theta_3$')
plt.xlabel(r'$\theta_1$ (radian)')
plt.ylabel('output angle (radian)')
plt.legend();
#this looks like the swing I am trying to achieve since the links are limited in their range
rA = l1*np.vstack([np.cos(a1), np.sin(a1)])
rB = rA + l2*np.vstack([np.cos(a2), np.sin(a2)])
rC = rB - l3*np.vstack([np.cos(a3), np.sin(a3)])
rP = rA + l2/2*np.vstack([np.cos(a2), np.sin(a2)])
links_x_locations = np.vstack([np.zeros(len(a1)),
rA[0, :],
rB[0, :],
rC[0, :]])
links_y_locations = np.vstack([np.zeros(len(a1)),
rA[1, :],
rB[1, :],
rC[1, :]])
i = 10
plt.plot(links_x_locations[:, i],
links_y_locations[:, i], 'k-o')
plt.plot(rA[0,:], rA[1,:], label = 'hinge A')
plt.plot(rB[0,:], rB[1,:], label = 'hinge B')
plt.plot(rC[0,:], rC[1,:], label = 'hinge C')
plt.plot(rP[0,:], rP[1,:], label = 'midpoint AB')
plt.legend()
plt.title('Paths and orientation for\n'+
r'$\theta_1$ = {:.1f}, $\theta_2$ = {:.1f}, $\theta_3$ = {:.1f}'.format(a1[i], a2[i], a3[i]))
plt.axis('equal');
# +
drive_rate = 10 #rad/s
dFbar = lambda a1, a2, a3, dx: np.array([l1*drive_rate*np.sin(a1)+\
l2*dx[0]*np.sin(a2)-\
l3*dx[1]*np.sin(a3),\
l1*drive_rate*np.cos(a1)+\
l2*dx[0]*np.cos(a2)-\
l3*dx[1]*np.cos(a3)])
da1 = np.ones(len(a1))*10
da2 = np.zeros(len(a1))
da3 = np.zeros(len(a1))
xsol = np.array([0, 0])
for i in range(len(a1)):
xsol = fsolve(lambda dx: dFbar(a1[i], a2[i], a3[i], dx), xsol)
da2[i] = xsol[0]
da3[i] = xsol[1]
plt.plot(a1, da1, label = r'$\dot{\theta}_1$')
plt.plot(a1, da2, label = r'$\dot{\theta}_2$')
plt.plot(a1, da3, label = r'$\dot{\theta}_3$')
plt.legend()
plt.xlabel(r'$\theta_1$ (radian)')
plt.ylabel('output angular speed (radian/s)')
# -
# ## Proposed Outline:
#
# ### Background
#
# - What are you trying to model?
# I am trying to simulate a good golf swing with the circular shape of the shoulders. The model is simplified because it acts in a four-bar linkage design. The bicep and forearm of one arm (made into two links, l1 and l2) and the other entire arm (l3). Free body diagram and motion of golf swing is shown in the jpeg (l1 and l2 should be switched). [IMG_5C3DE768BA33-1.jpeg](attachment:IMG_5C3DE768BA33-1.jpeg)
#
# I used my model from the Module 2, the four bar linkage design. It still could use work, but the general idea is there. Firstly, I defined my variable values l1 is the length of my forearm, l2 is the length of bicep, l3 is the length of the other arm, and dx is the length between my two shoulders (the model of the graph called 'Path and Orientation For...' shows this in birds-eye-view of my model). I then defined the motion of the hinges and their angles, plotted in the first graph 'output angle vs. theta 1' and second graph 'Paths and Orientation for.' I then found the angular velocity plotted in third graph, 'output angular speed vs theta.'
# My assumptions are that the model returns back to the same place as the start of the swing right before hitting the ball otherwise it cannot hit the ball. My constraints are that one arm has no hinge while the other does; this allows for a good and extended backswing but limits the front swing because it cannot move passed the full extension of the arm containing l1 and l2. This is due to the fact that l3 cannot bend to allow for the extension of the back arm (l1 and l2). The equations are used are in my code above. I used my Newton-Euler equations and constraint vectors to define postions of linkages. My analysis is shown above and graphs of my work are displayed above to demonstrate the angles, angular speed, and hinge movement.
#
#
# Originally I set up my linkage to run just the back arm: one link was the entire arm (including forearm and bicep), one link was the stationary collor bone area, and one link represented the spine. This model was not suitable to demonstrate the swing and professor helped me to change my model to include both arms and making one arm represented as a whole and and the second arm split at the elbow. This allowed me to meet my goal because it allowed for a fuller and more complete demonstration and analysis of the rotation of the golf swing.I have played golf my whole life and always knew the importance of the back swing but never realized how important the front swing is. Without full extension the ball cannot go its full potential. Since beginning this project I have focused more on my front swing and I have seen improvement in my game and reduced problems such as slicing (if you are a righty you hit the ball to the right and vice versa), as I thought my problem was my backswing but it was jut the lazy follow-through of my front swing that made me push the ball. I could improve the analysis of this device by splitting my front arm (l3) to have a hinge to allow for full extension of the front swing.
#
| M05_submissions/project-6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Python basics for exploring a PDB file
#
# See [here](https://github.com/fomightez/Python_basics_on_PDB_file) for information about this notebook.
#
# ------
#
# <div class="alert alert-block alert-warning">
# <p>If you haven't used one of these notebooks before, they're basically web pages in which you can write, edit, and run live code. They're meant to encourage experimentation, so don't feel nervous. Just try running a few cells and see what happens!.</p>
#
# <p>
# Some tips:
# <ul>
# <li>Code cells have boxes around them.</li>
# <li>To run a code cell either click the Play icon on the menu bar above, or click on the cell and then hit <b>Shift+Enter</b>. The <b>Shift+Enter</b> combo will also move you to the next cell, so it's a quick way to work through the notebook.</li>
# <li>While a cell is running a <b>*</b> appears in the square brackets next to the cell. Once the cell has finished running the asterisk will be replaced with a number.</li>
# <li>In most cases you'll want to start from the top of notebook and work your way down running each cell in turn. Later cells might depend on the results of earlier ones.</li>
# <li>To edit a code cell, just click on it and type stuff. Remember to run the cell once you've finished editing.</li>
# </ul>
# </p>
# </div>
#
#
# These sessions are temporary and will time out after ten minutes of inactivity. However, a safety net is built in if you start doing serious work in these. Even if it times out, you can save the notebook and upload it again later. See the notebook [safety net demo](safety%20net%20demo.ipynb) so that you are prepared for when it happens. You won't be able to open the demo if it has already happened.
#
# ----
#
#
# -
# ## Starting simple
#
#
# ## Pre-processing this PDB file to look more typical
#
# Use a file derived from [PDB entry for 3hyd](https://files.rcsb.org/view/3hyd.pdb) to see about columns.
#
# 3hyd we saw in the electron density cloud to isomesh demo tosay.
#
# [PDB entry for 3hyd](https://files.rcsb.org/view/3hyd.pdb) was modified to remove the anisotropic temperature factor lines. They begin with `ANISOU`.
#
# To remove those lines from the original [3hyd.pdb](3hyd.pdb) file, I used regular expressions in a text editor. (Not Microsoft Word). This is like fancy 'Find and replace'.
# We could use Python for this data pre-procesing step, but sometimes there is an easier way.
#
# The specifics are spelled out in [How_made_without_ANISOU_lines.md](How_made_without_ANISOU_lines.md).
#
# Let's use a fragment of the PDB file to see how it is done. We'll use [REGEX101](https://regex101.com/) to demonstrate right now.
#
# Performing that process on `3hyd.pdb` results in [3hydWITHOUTanisouLINES.pdb](3hydWITHOUTanisouLINES.pdb).
#
# We'll use the ATOM section of that that to define `t`.
t='''ATOM 1 N LEU A 1 1.149 1.920 3.550 1.00 5.65 N
ATOM 2 CA LEU A 1 2.138 2.288 4.580 1.00 5.04 C
ATOM 3 C LEU A 1 3.461 1.638 4.282 1.00 3.88 C
ATOM 4 O LEU A 1 3.527 0.405 4.165 1.00 4.79 O
ATOM 5 CB LEU A 1 1.635 1.889 5.948 1.00 6.19 C
ATOM 6 CG LEU A 1 2.444 2.344 7.182 1.00 10.41 C
ATOM 7 CD1 LEU A 1 1.603 2.227 8.438 1.00 18.81 C
ATOM 8 CD2 LEU A 1 3.699 1.583 7.375 1.00 10.45 C
ATOM 9 H1 LEU A 1 1.127 0.953 3.458 1.00 5.40 H
ATOM 10 H2 LEU A 1 0.274 2.239 3.813 1.00 5.15 H
ATOM 11 H3 LEU A 1 1.404 2.323 2.704 1.00 5.24 H
ATOM 12 HA LEU A 1 2.249 3.258 4.575 1.00 4.84 H
ATOM 13 HB2 LEU A 1 0.742 2.251 6.048 1.00 6.49 H
ATOM 14 HB3 LEU A 1 1.585 0.920 5.978 1.00 6.58 H
ATOM 15 HG LEU A 1 2.680 3.278 7.071 1.00 10.84 H
ATOM 16 HD11 LEU A 1 2.094 2.588 9.181 1.00 14.99 H
ATOM 17 HD12 LEU A 1 1.404 1.298 8.594 1.00 15.61 H
ATOM 18 HD13 LEU A 1 0.787 2.722 8.316 1.00 15.30 H
ATOM 19 HD21 LEU A 1 3.547 0.650 7.120 1.00 10.08 H
ATOM 20 HD22 LEU A 1 3.964 1.634 8.317 1.00 10.06 H
ATOM 21 HD23 LEU A 1 4.396 1.980 6.816 1.00 9.93 H
ATOM 22 N VAL A 2 4.521 2.434 4.230 1.00 3.54 N
ATOM 23 CA VAL A 2 5.866 1.930 4.058 1.00 3.13 C
ATOM 24 C VAL A 2 6.790 2.592 5.064 1.00 3.68 C
ATOM 25 O VAL A 2 6.806 3.837 5.179 1.00 4.26 O
ATOM 26 CB VAL A 2 6.425 2.168 2.650 1.00 3.78 C
ATOM 27 CG1 VAL A 2 7.834 1.667 2.541 1.00 5.01 C
ATOM 28 CG2 VAL A 2 5.522 1.554 1.584 1.00 5.05 C
ATOM 29 H VAL A 2 4.482 3.290 4.290 1.00 3.28 H
ATOM 30 HA VAL A 2 5.875 0.967 4.219 1.00 3.38 H
ATOM 31 HB VAL A 2 6.447 3.134 2.489 1.00 3.90 H
ATOM 32 HG11 VAL A 2 8.026 1.465 1.622 1.00 4.53 H
ATOM 33 HG12 VAL A 2 7.933 0.873 3.073 1.00 4.63 H
ATOM 34 HG13 VAL A 2 8.436 2.348 2.851 1.00 4.60 H
ATOM 35 HG21 VAL A 2 5.433 0.614 1.755 1.00 4.78 H
ATOM 36 HG22 VAL A 2 5.916 1.693 0.720 1.00 4.65 H
ATOM 37 HG23 VAL A 2 4.662 1.976 1.619 1.00 4.78 H
ATOM 38 N GLU A 3 7.567 1.777 5.771 1.00 2.92 N
ATOM 39 CA AGLU A 3 8.674 2.248 6.587 0.50 3.42 C
ATOM 40 CA BGLU A 3 8.670 2.254 6.575 0.50 3.37 C
ATOM 41 C GLU A 3 9.920 1.636 5.964 1.00 2.87 C
ATOM 42 O GLU A 3 10.041 0.405 5.901 1.00 3.28 O
ATOM 43 CB AGLU A 3 8.541 1.813 8.049 0.50 3.31 C
ATOM 44 CB BGLU A 3 8.513 1.848 8.037 0.50 3.20 C
ATOM 45 CG AGLU A 3 9.625 2.367 8.970 0.50 3.86 C
ATOM 46 CG BGLU A 3 9.663 2.264 8.926 0.50 4.43 C
ATOM 47 CD AGLU A 3 9.581 1.798 10.378 0.50 4.30 C
ATOM 48 CD BGLU A 3 9.372 2.079 10.393 0.50 7.27 C
ATOM 49 OE1AGLU A 3 9.326 0.609 10.555 0.50 4.93 O
ATOM 50 OE1BGLU A 3 8.211 1.826 10.769 0.50 8.58 O
ATOM 51 OE2AGLU A 3 9.823 2.595 11.317 0.50 5.24 O
ATOM 52 OE2BGLU A 3 10.301 2.252 11.187 0.50 5.38 O
ATOM 53 H GLU A 3 7.465 0.924 5.796 1.00 3.13 H
ATOM 54 HA AGLU A 3 8.739 3.223 6.557 0.50 3.34 H
ATOM 55 HA BGLU A 3 8.736 3.228 6.532 0.50 3.32 H
ATOM 56 HB2AGLU A 3 7.685 2.119 8.386 0.50 3.51 H
ATOM 57 HB2BGLU A 3 7.707 2.257 8.388 0.50 3.63 H
ATOM 58 HB3AGLU A 3 8.581 0.845 8.093 0.50 3.50 H
ATOM 59 HB3BGLU A 3 8.438 0.883 8.085 0.50 3.60 H
ATOM 60 HG2AGLU A 3 10.496 2.159 8.600 0.50 3.92 H
ATOM 61 HG2BGLU A 3 10.441 1.727 8.712 0.50 4.83 H
ATOM 62 HG3AGLU A 3 9.518 3.329 9.035 0.50 4.00 H
ATOM 63 HG3BGLU A 3 9.855 3.203 8.775 0.50 4.81 H
ATOM 64 N ALA A 4 10.842 2.465 5.490 1.00 2.75 N
ATOM 65 CA ALA A 4 12.006 2.016 4.750 1.00 3.09 C
ATOM 66 C ALA A 4 13.270 2.689 5.248 1.00 2.94 C
ATOM 67 O ALA A 4 13.284 3.899 5.529 1.00 3.58 O
ATOM 68 CB ALA A 4 11.833 2.220 3.258 1.00 4.15 C
ATOM 69 H ALA A 4 10.814 3.320 5.588 1.00 2.76 H
ATOM 70 HA ALA A 4 12.112 1.056 4.889 1.00 3.11 H
ATOM 71 HB1 ALA A 4 11.060 1.734 2.964 1.00 3.79 H
ATOM 72 HB2 ALA A 4 12.615 1.896 2.807 1.00 3.83 H
ATOM 73 HB3 ALA A 4 11.717 3.157 3.082 1.00 3.98 H
ATOM 74 N LEU A 5 14.334 1.918 5.332 1.00 2.99 N
ATOM 75 CA LEU A 5 15.634 2.369 5.796 1.00 3.30 C
ATOM 76 C LEU A 5 16.689 1.821 4.849 1.00 3.28 C
ATOM 77 O LEU A 5 16.716 0.605 4.614 1.00 3.43 O
ATOM 78 CB LEU A 5 15.875 1.877 7.209 1.00 4.50 C
ATOM 79 CG LEU A 5 17.250 2.151 7.852 1.00 9.40 C
ATOM 80 CD1 LEU A 5 17.789 3.495 7.677 1.00 9.16 C
ATOM 81 CD2 LEU A 5 17.128 1.821 9.337 1.00 11.72 C
ATOM 82 H LEU A 5 14.330 1.085 5.118 1.00 2.86 H
ATOM 83 HA LEU A 5 15.679 3.347 5.792 1.00 3.38 H
ATOM 84 HB2 LEU A 5 15.205 2.260 7.785 1.00 4.85 H
ATOM 85 HB3 LEU A 5 15.765 0.916 7.196 1.00 4.98 H
ATOM 86 HG LEU A 5 17.893 1.532 7.474 1.00 8.68 H
ATOM 87 HD11 LEU A 5 18.580 3.588 8.214 1.00 8.30 H
ATOM 88 HD12 LEU A 5 17.128 4.128 7.958 1.00 8.71 H
ATOM 89 HD13 LEU A 5 18.004 3.632 6.753 1.00 8.61 H
ATOM 90 HD21 LEU A 5 16.648 2.528 9.774 1.00 10.72 H
ATOM 91 HD22 LEU A 5 18.009 1.743 9.711 1.00 10.33 H
ATOM 92 HD23 LEU A 5 16.655 0.991 9.440 1.00 10.20 H
ATOM 93 N TYR A 6 17.519 2.699 4.281 1.00 3.06 N
ATOM 94 CA TYR A 6 18.548 2.342 3.321 1.00 3.08 C
ATOM 95 C TYR A 6 19.876 2.878 3.804 1.00 3.48 C
ATOM 96 O TYR A 6 20.024 4.082 3.994 1.00 4.60 O
ATOM 97 CB TYR A 6 18.275 2.935 1.947 1.00 3.85 C
ATOM 98 CG TYR A 6 16.873 2.775 1.400 1.00 3.42 C
ATOM 99 CD1 TYR A 6 15.855 3.614 1.819 1.00 3.67 C
ATOM 100 CD2 TYR A 6 16.556 1.796 0.450 1.00 3.75 C
ATOM 101 CE1 TYR A 6 14.558 3.509 1.303 1.00 3.54 C
ATOM 102 CE2 TYR A 6 15.281 1.695 -0.071 1.00 3.30 C
ATOM 103 CZ TYR A 6 14.283 2.550 0.357 1.00 3.37 C
ATOM 104 OH TYR A 6 13.017 2.418 -0.175 1.00 3.64 O
ATOM 105 H TYR A 6 17.497 3.543 4.445 1.00 2.94 H
ATOM 106 HA TYR A 6 18.611 1.369 3.230 1.00 3.22 H
ATOM 107 HB2 TYR A 6 18.454 3.887 1.978 1.00 3.50 H
ATOM 108 HB3 TYR A 6 18.879 2.517 1.315 1.00 3.43 H
ATOM 109 HD1 TYR A 6 16.041 4.275 2.443 1.00 3.43 H
ATOM 110 HD2 TYR A 6 17.223 1.227 0.139 1.00 3.37 H
ATOM 111 HE1 TYR A 6 13.891 4.088 1.592 1.00 3.38 H
ATOM 112 HE2 TYR A 6 15.090 1.043 -0.705 1.00 3.24 H
ATOM 113 HH TYR A 6 12.808 3.103 -0.576 1.00 3.44 H
ATOM 114 N LEU A 7 20.857 2.006 3.973 1.00 4.38 N
ATOM 115 CA LEU A 7 22.208 2.473 4.312 1.00 6.08 C
ATOM 116 C LEU A 7 23.293 1.644 3.744 1.00 7.46 C
ATOM 117 O LEU A 7 23.018 0.697 3.000 1.00 13.45 O
ATOM 118 CB LEU A 7 22.356 2.753 5.793 1.00 11.09 C
ATOM 119 CG LEU A 7 22.263 1.578 6.717 1.00 11.34 C
ATOM 120 CD1 LEU A 7 22.913 1.965 8.038 1.00 18.81 C
ATOM 121 CD2 LEU A 7 20.853 1.118 7.009 1.00 11.10 C
ATOM 122 OXT LEU A 7 24.470 1.913 3.995 1.00 9.60 O
ATOM 123 H LEU A 7 20.779 1.153 3.901 1.00 4.08 H
ATOM 124 HA LEU A 7 22.322 3.342 3.880 1.00 6.70 H
ATOM 125 HB2 LEU A 7 23.219 3.171 5.933 1.00 9.43 H
ATOM 126 HB3 LEU A 7 21.660 3.376 6.052 1.00 9.61 H
ATOM 127 HG LEU A 7 22.755 0.830 6.347 1.00 11.80 H
ATOM 128 HD11 LEU A 7 23.845 2.140 7.890 1.00 14.43 H
ATOM 129 HD12 LEU A 7 22.815 1.242 8.663 1.00 14.59 H
ATOM 130 HD13 LEU A 7 22.483 2.753 8.379 1.00 14.87 H
ATOM 131 HD21 LEU A 7 20.870 0.474 7.720 1.00 10.59 H
ATOM 132 HD22 LEU A 7 20.484 0.716 6.220 1.00 10.51 H
ATOM 133 HD23 LEU A 7 20.323 1.876 7.268 1.00 10.64 H '''
#Let's parse the text on line breaks
for line in t.split("\n"):
print(line)
#Let's examine the 17th column
for line in t.split("\n"):
print(line[16])
#Let's only show those that have a 17th column
for line in t.split("\n"):
if line[16] != ' ':
print(line[16])
# Note `line` isn't magical. We could have used any variable.
#Let's only show those that have a 17th column
for x in t.split("\n"):
if x[16] != ' ':
print(x[16])
# Python is simply assigning the `x` to each individual element in the list made by `t.split("\n")`.
# By itself that isn't that shocking but that patter of Python automatically taking the individual elements from an iterable object is useful. We'll see it soon where we read a file item and the lines are actually specified without us needing ti split on the end of line representation `\n` in that case.
#
# But why are we using the number `16` when we talked about the 17th column?
# Lets explore indexing in Python.
#
# Let's look at what numbers in prints if we iterate on a `range` of integers. Note that `range` is green because it is a special Python object whereas `x` and `line` we used above weren't green.
for x in range(5):
print (x)
# Okay. We get five numbers. However, the first one is zero and not `1` as we saw at [PDB ATOMIC COORDINATE FILE FORMAT](https://zhanglab.ccmb.med.umich.edu/BindProfX/pdb_atom_format.html).
#
# This is because Python is zero indexed. Not the normal way but if you recall sometimes you'll see in grade school number lines beginning with zero. Mathematics branches and computer science often use zero as the index of the first item in a list. This is what Python uses. (The statistics language R using 1 indexing.)
#
# We'll see zero indexing has some advantages as we go along. It may take until session #5 to really see the benefits.
#
# Let's see if `line[16]` corresponing to the 17th column makes more sense now.
#
# One way to look at it:
#Let's only show those that have a 17th column
for line in t.split("\n"):
if line[17-1] != ' ':
print(line[17-1]) # because to account for zero index, we want one less than number 17
print(line[0:17])
# ` print(line[0:17])` is slicing the line and not prining a single character at a specified index.
#
# Note the syntax is a little different than you might expect, it is saying start with zero index (number on the right) and go up to **BUT NOT INCLUDE** the character at the index on the right. We'll cover slicing some more in Session #5 as it can be a little odd at first and since there are shortcuts you can use. However, you should see it easily allows you to access items spanning particular numbered columns as we saw [the PDB specification](https://zhanglab.ccmb.med.umich.edu/BindProfX/pdb_atom_format.html).
# ## File reading
#
# That was slightly tedious getting the PDB file and pasting it in above to make the text string `t`. Normally you'd read stright from the file.
#
# First let's use the shell command line utility `curl` to get (You may be more familiar with `wget` which acts similarly?)
#
# Note we put an `!` (exclamation point at the beginning of the curl command to specify that is special and instead of running as Python (which is the kernel this notebook is based on, see upper left just above the notabook), we want to run it as a command bash shell command. (Another way to think about it: If you were in a terminal, you'd leave off the exclamation point.)
#
# Feel free to change it to your favorite PDB file id accession at the end. In other words replace the `1avw` with whatver you'd like.
# !curl -OL https://files.rcsb.org/download/1p3v.pdb
# Running that gets another PDB file.
#
# Let's parse that straight from the file.
# read in the PDB file
with open("1p3v.pdb", 'r') as input:
lines_read = 0
for line in input:
lines_read = lines_read + 1
print (lines_read)
# Note to get that to work we didn't have to split the string on the new line character `/n` this time. Python knows to split a file object on the line. Or even more general the iterable assigned to a file object is a line.
#
# If we interated on the line now, we'd get characters. This unit drill down nature of Python is one of the things that make it so useful. The other is that it looks much like if you wrote out what you want to do. In other words it is a higher level language closer to English than some other programming languages.
# If you click to open your PDB file and scroll down, that number reported above should match although it may be off by one as the last line showing a line number in the editor view isn't counted as a line by Python as it contains nothing.
#
# We can confirm that by printing the last known value of `line` and see it corresponds to the content on last line where there is nothing. (Line 2119 if you are using 1p3v.)
print(line)
# Let's use that to do something more useful that will let us explore the PDB file more.
#
# We'll remove the counting and trying looking at how many cysteines this protein (or structure if yours has more chains) has.
with open("1p3v.pdb", 'r') as input:
for line in input:
if 'CYS' in line:
print(line)
# Well we got our answer. But it could be cleaner. The first two lines come from the section of the PDB file called the header. It would be worse if we had a lot of cysteines. Those first two lines all the parts above the ATOM coordinates.
#
# Let's see how much worse it could get by trying with `HIS`.
# read in the PDB file
with open("1p3v.pdb", 'r') as input:
for line in input:
if 'HIS' in line:
print(line)
# Now we are getting lines with `THIS` in the header, too.
#
# Let's skip past the header.
#
# We didn't talk about it above but we can use another conditional like the `if something` we used above a few times.
with open("1p3v.pdb", 'r') as input:
for line in input:
if line.startswith("ATOM"):
if 'CYS' in line:
print(line)
# That is cleaner. And it will work better with `HIS` now.
with open("1p3v.pdb", 'r') as input:
# prepare to give feeback later or allow skipping to certain start
for line in input:
if line.startswith("ATOM"):
if 'HIS' in line:
print(line)
# But hard to read how many since several of them. Let's focus on the alpha-carbons in the PDB file.
with open("1p3v.pdb", 'r') as input:
# prepare to give feeback later or allow skipping to certain start
for line in input:
if line.startswith("ATOM") and "CA" in line:
if 'HIS' in line:
print(line)
# So there are nine histidines if you are using 1p3v example.
#
# So this was using Python interactively in Jupyter. And you could save this notebook to easily save your result of the code above looking at the alpha-carbons of the histides for a report or something later. A lot of people prefer this way to work now. We'll explore more of the advatanges in Session #5.
#
#
# ----
#
# ## Running a script
#
# But you may have heard reference to running a Python script or running a script. Let's finish by running a script version of our histidine alpha-carbon listing code.
#
# To do that, copy the code in the cell above to your clipboard. It will be similar to the cod below but may be slightly different if you are using a differnet PDB file:
#
# ```python
# with open("1p3v.pdb", 'r') as input:
# # prepare to give feeback later or allow skipping to certain start
# for line in input:
# if line.startswith("ATOM") and "CA" in line:
# if 'HIS' in line:
# print(line)
# ```
#
# Next select `File` > `New` > `Text File` from the main menu at the top above the panels.
#
# Paste the code into the text file and then right-click choose `Rename File` and name the file `script.py`
#
# **It is important you remove `.txt`.** The highlighting should indicate code now if you did it correctly.
#
# Choose `File` > `Save Python File.`
#
#
# Next, select `File` > `New` > `Terminal` from the main menu at the top above the panels.
#
# A terminal will open. There type the following to run your script:
#
# ```shell
# python script.py
# ```
#
# You'll see the same result you saw in the notebook. However, it isn't as useful as saving the notebook automatically. You'd have to add extra handling to save those results to a file. It is easy but you can already see one advantage to using a Jupyter notebook from that. Of course, you could copy and paste the result out of terminal but doing that more than a few times is tedious and not good practice to working reproducibly. The most direct way is to add a shell redirect. The ouput to standard out in the terminal will be sent to a file named `results.txt` with the following addition of `> results.txt` at the end of the call to the script. Like so:
#
# ```shell
# python script.py > results.txt
# ```
#
# You won't see any ouput but a file named `results.txt` will be made with the outout.
#
# -----
| session3_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#copied from https://colab.research.google.com/drive/1pTuQhug6Dhl9XalKB0zUGf4FIdYFlpcX#scrollTo=Z474sSC6oe7A
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="DEfSbAA4QHas" outputId="a6536544-6bd1-462a-d946-ec6bdb8a1a2d"
# import tensorflow as tf
# # Get the GPU device name.
# device_name = tf.test.gpu_device_name()
# # The device name should look like the following:
# if device_name == '/device:GPU:0':
# print('Found GPU at: {}'.format(device_name))
# else:
# raise SystemError('GPU device not found')
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="oYsV4H8fCpZ-" outputId="47d8fbc6-39b6-4067-a37b-c792342c54b8"
import torch
# tasks = ['cola', 'MRPC']
TASK = 'MRPC'
import pdb
def get_device():
# If there's a GPU available...
if torch.cuda.is_available():
# Tell PyTorch to use the GPU.
device = torch.device("cuda")
print('There are %d GPU(s) available.' % torch.cuda.device_count())
print('We will use the GPU:', torch.cuda.get_device_name(0))
# If not...
else:
print('No GPU available, using the CPU instead.')
device = torch.device("cpu")
return device
device = get_device()
# + colab={"base_uri": "https://localhost:8080/", "height": 406} colab_type="code" id="0NmMdkZO8R6q" outputId="81878b39-ed58-4dce-cf0a-24ef4a470dfd"
# !pip install transformers
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="5m6AnuFv0QXQ" outputId="dad7440c-0ec0-4d80-b80d-a6f96efa7787"
# !pip install wget
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="pMtmPMkBzrvs" outputId="8923f4fa-e40c-412d-a644-f65793c3cd9c"
import wget
import os
print('Downloading dataset...')
def download_data(task):
task_to_data = {
'cola': ('https://nyu-mll.github.io/CoLA/cola_public_1.1.zip', './cola_public_1.1.zip', './cola_public/')
}
if task != 'cola':
return
url, download_file, unzip_file = task_to_data[task]
# Download the file (if we haven't already)
if not os.path.exists(download_file):
wget.download(url, download_file)
# Unzip the dataset (if we haven't already)
if not os.path.exists(unzip_file):
# !unzip $unzip_file
download_data("cola")
# +
def create_examples(lines, set_type):
"""Creates examples for the training and dev sets."""
examples = []
for (i, line) in enumerate(lines):
if i == 0:
continue
guid = "%s-%s" % (set_type, i)
text_a = convert_to_unicode(line[3])
text_b = convert_to_unicode(line[4])
if set_type == "test":
label = "0"
else:
label = convert_to_unicode(line[0])
examples.append(
{'guid':guid, 'text_a':text_a, 'text_b':text_b, 'label':label})
return pd.DataFrame(examples)
def read_tsv(input_file, quotechar=None):
"""Reads a tab separated value file."""
with open(input_file, "r", encoding="utf8") as f:
reader = csv.reader(f, delimiter="\t", quotechar=quotechar)
lines = []
for line in reader:
lines.append(line)
return lines
def convert_to_unicode(text):
if isinstance(text, str):
return text
elif isinstance(text, bytes):
return text.decode("utf-8", "ignore")
# + colab={"base_uri": "https://localhost:8080/", "height": 377} colab_type="code" id="_UkeC7SG2krJ" outputId="47ae0de6-4c95-4b82-ad0f-e3fb27b8ef76"
import pandas as pd
import csv
def load_data(sample_data = False):
if TASK == 'cola':
# Load the dataset into a pandas dataframe.
df = pd.read_csv("./cola_public/raw/in_domain_train.tsv", delimiter='\t', header=None,
names=['sentence_source', 'label', 'label_notes', 'sentence'])
if TASK == 'MRPC':
return pd.concat((
[create_examples(read_tsv('glue_data/MRPC/' + name + '.tsv'), name)
for name in ['train', 'dev', 'test']]), axis=0)
# Report the number of sentences.
print('Number of training sentences: {:,}\n'.format(df.shape[0]))
if sample_data:
df = df.head(sample_data)
return df
df = load_data()
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="Z474sSC6oe7A" outputId="7e93e7c9-9edf-473b-dbe9-eb78cdab6996"
from transformers import BertTokenizer
# Load the BERT tokenizer.
print('Loading BERT tokenizer...')
tokenizer = BertTokenizer.from_pretrained('bert-base-uncased', do_lower_case=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="cKsH2sU0OCQA" outputId="0a95553f-5911-419f-b12f-f72b8f56a235"
# # max_len = 0
# # # For every sentence...
# # for sent in sentences:
# # # Tokenize the text and add `[CLS]` and `[SEP]` tokens.
# # input_ids = tokenizer.encode(sent, add_special_tokens=True)
# # # Update the maximum sentence length.
# # max_len = max(max_len, len(input_ids))
# # print('Max sentence length: ', max_len)
# max_len = 30
# + colab={"base_uri": "https://localhost:8080/", "height": 151} colab_type="code" id="2bBdb3pt8LuQ" outputId="b34143b4-dac4-4240-ee92-f3694fd566f9"
# def get_features_old(task, df = None):
# if task == 'cola':
# return get_features_cola(df)
def get_features(df):
# Get the lists of sentences and their labels.
if TASK == 'cola':
sentences = df.sentence.values
if TASK == 'MRPC':
sentences = df[['text_a', 'text_b']].values.tolist()
labels = df.label.values.astype(int)
# Tokenize all of the sentences and map the tokens to thier word IDs.
input_ids = []
attention_masks = []
token_type_ids = []
# For every sentence...
for sent in sentences:
# `encode_plus` will:
# (1) Tokenize the sentence.
# (2) Prepend the `[CLS]` token to the start.
# (3) Append the `[SEP]` token to the end.
# (4) Map tokens to their IDs.
# (5) Pad or truncate the sentence to `max_length`
# (6) Create attention masks for [PAD] tokens.
if TASK == 'MRPC':
text_b = sent[1]
sent = sent[0]
else:
text_b = None
encoded_dict = tokenizer.encode_plus(
text = sent,
text_pair = text_b,
add_special_tokens = True, # Add '[CLS]' and '[SEP]'
max_length = 64, # Pad & truncate all sentences.
pad_to_max_length = True,
return_attention_mask = True, # Construct attn. masks.
return_tensors = 'pt',
truncation = True # Return pytorch tensors.
)
if TASK == 'MRPC':
token_type_ids.append(encoded_dict['token_type_ids'])
# Add the encoded sentence to the list.
input_ids.append(encoded_dict['input_ids'])
# And its attention mask (simply differentiates padding from non-padding).
attention_masks.append(encoded_dict['attention_mask'])
# Convert the lists into tensors.
input_ids = torch.cat(input_ids, dim=0)
attention_masks = torch.cat(attention_masks, dim=0)
labels = torch.tensor(labels)
# Print sentence 0, now as a list of IDs.
print('Original: ', sentences[0])
print('Token IDs:', input_ids[0])
ret = [input_ids, attention_masks, labels]
if TASK == 'MRPC':
token_type_ids = torch.cat(token_type_ids, dim=0)
ret += [token_type_ids]
else:
ret += [None]
return ret
input_ids, attention_masks, labels, token_type_ids = get_features(df)
# +
import numpy as np
from torch.utils.data import TensorDataset, random_split
if token_type_ids is not None:
token_type_ids = [token_type_ids]
else:
token_type_ids = []
dataset = TensorDataset(input_ids, attention_masks, labels, *token_type_ids)
def get_kfold(input_ids, attention_masks, labels, token_type_ids, k=3):
dataset = TensorDataset(input_ids, attention_masks, labels, *token_type_ids)
idx = np.arange(len(input_ids))
np.random.shuffle(idx)
fold_length = len(input_ids)//k
data = []
for i in range(k):
start_idx = i*fold_length
end_idx = (i+1)*fold_length
if i == k-1:
end_idx = len(input_ids)
validation_idx = idx[start_idx: end_idx]
train_idx = np.concatenate((idx[0: start_idx], idx[end_idx: len(input_ids)]))
validation_set = TensorDataset(*dataset[validation_idx])
training_set = TensorDataset(*dataset[train_idx])
data.append((training_set, validation_set))
return data
data = get_kfold(input_ids, attention_masks, labels, token_type_ids, k=3)
full_dataset = TensorDataset(input_ids, attention_masks, labels, *token_type_ids)
# -
# +
from torch.utils.data import DataLoader, RandomSampler, SequentialSampler
from transformers import BertForSequenceClassification, AdamW, BertConfig
from transformers import get_linear_schedule_with_warmup
import numpy as np
batch_size = 32
from torch import optim
import random
import numpy as np
import pdb
# Function to calculate the accuracy of our predictions vs labels
def flat_accuracy(preds, labels):
pred_flat = np.argmax(preds, axis=1).flatten()
labels_flat = labels.flatten()
return np.sum(pred_flat == labels_flat) / len(labels_flat)
import time
import datetime
def check_mcc(model, prediction_dataloader):
predictions, true_labels = make_predictions(model, prediction_dataloader)
true_labels = np.concatenate(true_labels)
preds = np.concatenate(predictions)
preds = preds.argmax(1)
return matthews_corrcoef(true_labels, preds)
from sklearn.metrics import matthews_corrcoef
def make_predictions(model, prediction_dataloader):
# Prediction on test set
model.eval()
predictions , true_labels = [], []
for batch in prediction_dataloader:
batch = tuple(t.to(device) for t in batch)
b_input_ids, b_input_mask, b_labels = batch
with torch.no_grad():
outputs = model(b_input_ids, token_type_ids=None,
attention_mask=b_input_mask)
logits = outputs[0]
logits = logits.detach().cpu().numpy()
label_ids = b_labels.to('cpu').numpy()
predictions.append(logits)
true_labels.append(label_ids)
model.train()
return predictions, true_labels
def format_time(elapsed):
'''
Takes a time in seconds and returns a string hh:mm:ss
'''
# Round to the nearest second.
elapsed_rounded = int(round((elapsed)))
# Format as hh:mm:ss
return str(datetime.timedelta(seconds=elapsed_rounded))
def train_model(
epochs, train_dataloader, validation_dataloader,
verbose = False, release = False, prediction_dataloader = None,
lr = 2e-5, lrbase = 1e-5, lrclass = 1e-4):
model = BertForSequenceClassification.from_pretrained(
"bert-base-uncased", # Use the 12-layer BERT model, with an uncased vocab.
num_labels = 2,
output_attentions = False, # Whether the model returns attentions weights.
output_hidden_states = False, # Whether the model returns all hidden-states.
)
mcc_arr = []
model = model.to(device)
optimizer = AdamW(model.classifier.parameters(),
lr = lr, # args.learning_rate - default is 5e-5, our notebook had 2e-5
eps = 1e-8 # args.adam_epsilon - default is 1e-8.
)
# optimizer = optim.Adam([
# # {'params': model.bert.parameters(), 'lr': lrbase},
# {'params': model.classifier.parameters(), 'lr': lrclass}],
# lr = lr/10, # args.learning_rate - default is 5e-5, our notebook had 2e-5
# eps = 1e-8 # args.adam_epsilon - default is 1e-8.
# )
# optim.SGD([
# {'params': model.base.parameters()},
# {'params': model.classifier.parameters(), 'lr': 1e-3}
# ], lr=1e-2, momentum=0.9)
total_steps = len(train_dataloader) * epochs
# scheduler = get_linear_schedule_with_warmup(optimizer,
# num_warmup_steps = 0, # Default value in run_glue.py
# num_training_steps = total_steps)
# initval = [v.cpu().detach().numpy() for v in model.parameters()]
# We'll store a number of quantities such as training and validation loss,
# validation accuracy, and timings.
training_stats = []
total_t0 = time.time()
for epoch_i in range(0, epochs):
if verbose:
print("")
print('======== Epoch {:} / {:} ========'.format(epoch_i + 1, epochs))
print('Training...')
t0 = time.time()
total_train_loss = 0
model.train()
for step, batch in enumerate(train_dataloader):
if step % 40 == 0 and not step == 0:
elapsed = format_time(time.time() - t0)
print(' Batch {:>5,} of {:>5,}. Elapsed: {:}.'.format(
step, len(train_dataloader), elapsed))
if release:
mcc = check_mcc(model, prediction_dataloader)
print('mcc: ', mcc)
b_input_ids = batch[0].to(device)
b_input_mask = batch[1].to(device)
b_labels = batch[2].to(device).long()
if len(batch) > 3:
b_token_type_ids = batch[3].to(device)
else:
b_token_type_ids = None
model.zero_grad()
loss, logits = model(b_input_ids,
attention_mask=b_input_mask,
labels=b_labels,
token_type_ids = b_token_type_ids)
total_train_loss += loss.item()
loss.backward()
torch.nn.utils.clip_grad_norm_(model.parameters(), 1.0)
# tempval = [v.cpu().detach().numpy() for v in model.parameters()]
optimizer.step()
# scheduler.step()
# newval = [v.cpu().detach().numpy() for v in model.parameters()]
avg_train_loss = total_train_loss / len(train_dataloader)
training_time = format_time(time.time() - t0)
if verbose:
print("")
print(" Average training loss: {0:.2f}".format(avg_train_loss))
print(" Training epcoh took: {:}".format(training_time))
print("")
print("Running Validation...")
if epoch_i == 0:
optimizer = AdamW(model.parameters(),
lr = lr, # args.learning_rate - default is 5e-5, our notebook had 2e-5
eps = 1e-8 # args.adam_epsilon - default is 1e-8.
)
# optimizer = optim.Adam([
# {'params': model.bert.parameters(), 'lr': lrbase},
# {'params': model.classifier.parameters(), 'lr': lrclass}],
# lr = lr/10, # args.learning_rate - default is 5e-5, our notebook had 2e-5
# eps = 1e-8 # args.adam_epsilon - default is 1e-8.
# )
t0 = time.time()
if release:
mcc = check_mcc(model, prediction_dataloader)
print('mcc: ', mcc)
mcc_arr.append(mcc)
continue
model.eval()
total_eval_accuracy = 0
total_eval_loss = 0
nb_eval_steps = 0
for batch in validation_dataloader:
b_input_ids = batch[0].to(device)
b_input_mask = batch[1].to(device)
b_labels = batch[2].to(device).long()
if len(batch) > 3:
b_token_type_ids = batch[3].to(device)
else:
b_token_type_ids = None
with torch.no_grad():
(loss, logits) = model(b_input_ids,
token_type_ids=b_token_type_ids,
attention_mask=b_input_mask,
labels=b_labels)
total_eval_loss += loss.item()
logits = logits.detach().cpu().numpy()
label_ids = b_labels.to('cpu').numpy()
total_eval_accuracy += flat_accuracy(logits, label_ids)
avg_val_accuracy = total_eval_accuracy / len(validation_dataloader)
print(" Accuracy: {0:.2f}".format(avg_val_accuracy))
avg_val_loss = total_eval_loss / len(validation_dataloader)
# Measure how long the validation run took.
validation_time = format_time(time.time() - t0)
print(" Validation Loss: {0:.2f}".format(avg_val_loss))
print(" Validation took: {:}".format(validation_time))
# Record all statistics from this epoch.
training_stats.append(
{
'epoch': epoch_i + 1,
'Training Loss': avg_train_loss,
'Valid. Loss': avg_val_loss,
'Valid. Accur.': avg_val_accuracy,
'Training Time': training_time,
'Validation Time': validation_time
}
)
if release:
return _, _, mcc_arr, model
print("")
print("Training complete!")
print("Total training took {:} (h:mm:ss)".format(format_time(time.time()-total_t0)))
return avg_val_accuracy, avg_val_loss, training_stats, model
# + colab={} colab_type="code" id="XGUqOCtgqGhP"
# avg_val_accuracies, avg_val_losses = [], []
# for train_dataset, val_dataset in data:
# train_dataloader = DataLoader(
# train_dataset, # The training samples.
# sampler = RandomSampler(train_dataset), # Select batches randomly
# batch_size = batch_size # Trains with this batch size.
# )
# validation_dataloader = DataLoader(
# val_dataset, # The validation samples.
# sampler = SequentialSampler(val_dataset), # Pull out batches sequentially.
# batch_size = batch_size # Evaluate with this batch size.
# )
# epochs = 2
# avg_val_accuracy, avg_val_loss, training_stats, model = train_model(
# epochs, train_dataloader, validation_dataloader)
# avg_val_accuracies.append(avg_val_accuracy)
# avg_val_losses.append(avg_val_loss)
# val_loss, val_acc = [sum(arr) / len(arr) for arr in [avg_val_losses, avg_val_accuracies]]
# + colab={"base_uri": "https://localhost:8080/", "height": 195} colab_type="code" id="6O_NbXFGMukX" outputId="6686e966-46a5-4445-f899-d1f4f79a2927"
# import pandas as pd
# # Display floats with two decimal places.
# pd.set_option('precision', 2)
# # Create a DataFrame from our training statistics.
# df_stats = pd.DataFrame(data=training_stats)
# # Use the 'epoch' as the row index.
# df_stats = df_stats.set_index('epoch')
# # A hack to force the column headers to wrap.
# #df = df.style.set_table_styles([dict(selector="th",props=[('max-width', '70px')])])
# # Display the table.
# df_stats
# + colab={"base_uri": "https://localhost:8080/", "height": 427} colab_type="code" id="68xreA9JAmG5" outputId="265c6738-9f28-4817-a350-a63a1ac01934"
# import matplotlib.pyplot as plt
# # %matplotlib inline
# import seaborn as sns
# # Use plot styling from seaborn.
# sns.set(style='darkgrid')
# # Increase the plot size and font size.
# sns.set(font_scale=1.5)
# plt.rcParams["figure.figsize"] = (12,6)
# # Plot the learning curve.
# plt.plot(df_stats['Training Loss'], 'b-o', label="Training")
# plt.plot(df_stats['Valid. Loss'], 'g-o', label="Validation")
# # Label the plot.
# plt.title("Training & Validation Loss")
# plt.xlabel("Epoch")
# plt.ylabel("Loss")
# plt.legend()
# plt.xticks([1, 2, 3, 4])
# plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="mAN0LZBOOPVh" outputId="e97be795-f802-4c2e-8f2b-d2ac5cd2eead"
import pandas as pd
if TASK == 'cola':
df = pd.read_csv("./cola_public/raw/out_of_domain_dev.tsv", delimiter='\t',
header=None, names=['sentence_source', 'label', 'label_notes', 'sentence'])
sentences = df.sentence.values
labels = df.label.values
if TASK == 'MRPC':
sentences = df[['text_a', 'text_b']].values.tolist()
labels = df.label.values.astype(int)
# Tokenize all of the sentences and map the tokens to thier word IDs.
input_ids = []
attention_masks = []
# For every sentence...
for sent in sentences:
# `encode_plus` will:
# (1) Tokenize the sentence.
# (2) Prepend the `[CLS]` token to the start.
# (3) Append the `[SEP]` token to the end.
# (4) Map tokens to their IDs.
# (5) Pad or truncate the sentence to `max_length`
# (6) Create attention masks for [PAD] tokens.
if TASK == 'MRPC':
text_b = sent[1]
sent = sent[0]
else:
text_b = None
encoded_dict = tokenizer.encode_plus(
text = sent,
text_pair = text_b,
add_special_tokens = True, # Add '[CLS]' and '[SEP]'
max_length = 64, # Pad & truncate all sentences.
pad_to_max_length = True,
return_attention_mask = True, # Construct attn. masks.
return_tensors = 'pt',
truncation = True # Return pytorch tensors.
)
# Add the encoded sentence to the list.
input_ids.append(encoded_dict['input_ids'])
# And its attention mask (simply differentiates padding from non-padding).
attention_masks.append(encoded_dict['attention_mask'])
# Convert the lists into tensors.
input_ids = torch.cat(input_ids, dim=0)
attention_masks = torch.cat(attention_masks, dim=0)
labels = torch.tensor(labels)
# Set the batch size.
# Create the DataLoader.
prediction_data = TensorDataset(input_ids, attention_masks, labels)
prediction_sampler = SequentialSampler(prediction_data)
prediction_dataloader = DataLoader(prediction_data, sampler=prediction_sampler, batch_size=batch_size)
# + [markdown] colab_type="text" id="16lctEOyNFik"
# ## 5.2. Evaluate on Test Set
#
# + [markdown] colab_type="text" id="rhR99IISNMg9"
#
# With the test set prepared, we can apply our fine-tuned model to generate predictions on the test set.
# +
dfhyper = pd.DataFrame()
epochs = 12
mccs = []
full_dataloader = DataLoader(
full_dataset, # The training samples.
sampler = RandomSampler(full_dataset), # Select batches randomly
batch_size = batch_size # Trains with this batch size.
)
if TASK == 'cola':
lrs = [4e-5, 2e-5, 1e-5]
if TASK == 'MRPC':
lrs = [1e-3, 3e-4, 1e-4, 3e-5, 1e-5]
for lr in lrs:
_, _, mcc_arr, model = train_model(
epochs, full_dataloader, None, release = True, lr=lr,
prediction_dataloader = prediction_dataloader)
del model
dfhyper = dfhyper.append({'lr':lr, 'mcc_arr':mcc_arr}, ignore_index=True)
mccs.append(mcc_arr)
print(dfhyper)
# for lrbase in [3e-4, 1e-4, 3e-5, 1e-5][::-1]:
# for lrclass in [3e-4, 1e-4, 3e-5, 1e-5]:
# epochs = 6
# _, _, mcc_arr, model = train_model(
# epochs, full_dataloader, None, release = True, lrclass=lrclass, lrbase=lrbase,
# prediction_dataloader = prediction_dataloader)
# del model
# dfhyper.append({'lrbase':lrbase, 'lrclass':lrclass, 'mcc_arr':mcc_arr}, ignore_index=True)
# -
[print(f) for f in zip(dfhyper['lr'],dfhyper['mcc_arr'].values)]
sdfsdf
# +
# full_dataloader = DataLoader(
# full_dataset, # The training samples.
# sampler = RandomSampler(full_dataset), # Select batches randomly
# batch_size = batch_size # Trains with this batch size.
# )
# epochs = 6
# _, _, mcc_arr, model = train_model(
# epochs, full_dataloader, None, release = True, lr=1e-4)
# -
torch.cuda.empty_cache()
mcc_arr
# + colab={"base_uri": "https://localhost:8080/", "height": 84} colab_type="code" id="6ulTWaOr8QNY" outputId="a5517081-2e05-4244-c8df-77a9558ff75a"
import os
# Saving best-practices: if you use defaults names for the model, you can reload it using from_pretrained()
output_dir = './model_save/'
# Create output directory if needed
if not os.path.exists(output_dir):
os.makedirs(output_dir)
print("Saving model to %s" % output_dir)
# Save a trained model, configuration and tokenizer using `save_pretrained()`.
# They can then be reloaded using `from_pretrained()`
model_to_save = model.module if hasattr(model, 'module') else model # Take care of distributed/parallel training
model_to_save.save_pretrained(output_dir)
tokenizer.save_pretrained(output_dir)
# Good practice: save your training arguments together with the trained model
# torch.save(args, os.path.join(output_dir, 'training_args.bin'))
# + [markdown] colab_type="text" id="Z-tjHkR7lc1I"
# Let's check out the file sizes, out of curiosity.
# + colab={"base_uri": "https://localhost:8080/", "height": 118} colab_type="code" id="mqMzI3VTCZo5" outputId="6df0b283-6458-4d95-8455-2e7537193d1b"
# !ls -l --block-size=K ./model_save/
# + [markdown] colab_type="text" id="fr_bt2rFlgDn"
# The largest file is the model weights, at around 418 megabytes.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="-WUFUIQ8Cu8D" outputId="70780762-7790-474f-e5c2-304a066945ae"
# !ls -l --block-size=M ./model_save/pytorch_model.bin
# + [markdown] colab_type="text" id="dzGKvOFAll_e"
# To save your model across Colab Notebook sessions, download it to your local machine, or ideally copy it to your Google Drive.
# + colab={} colab_type="code" id="Trr-A-POC18_"
# Mount Google Drive to this Notebook instance.
# from google.colab import drive
# drive.mount('/content/drive')
# + colab={} colab_type="code" id="NxlZsafTC-V5"
# Copy the model files to a directory in your Google Drive.
# !cp -r ./model_save/ "./drive/Shared drives/ChrisMcCormick.AI/Blog Posts/BERT Fine-Tuning/"
# + [markdown] colab_type="text" id="W0vstijw85SZ"
# The following functions will load the model back from disk.
# + colab={} colab_type="code" id="nskPzUM084zL"
# Load a trained model and vocabulary that you have fine-tuned
model = model_class.from_pretrained(output_dir)
tokenizer = tokenizer_class.from_pretrained(output_dir)
# Copy the model to the GPU.
model.to(device)
# + colab={} colab_type="code" id="QxSMw0FrptiL"
# This code is taken from:
# https://github.com/huggingface/transformers/blob/5bfcd0485ece086ebcbed2d008813037968a9e58/examples/run_glue.py#L102
# Don't apply weight decay to any parameters whose names include these tokens.
# (Here, the BERT doesn't have `gamma` or `beta` parameters, only `bias` terms)
no_decay = ['bias', 'LayerNorm.weight']
# Separate the `weight` parameters from the `bias` parameters.
# - For the `weight` parameters, this specifies a 'weight_decay_rate' of 0.01.
# - For the `bias` parameters, the 'weight_decay_rate' is 0.0.
optimizer_grouped_parameters = [
# Filter for all parameters which *don't* include 'bias', 'gamma', 'beta'.
{'params': [p for n, p in param_optimizer if not any(nd in n for nd in no_decay)],
'weight_decay_rate': 0.1},
# Filter for parameters which *do* include those.
{'params': [p for n, p in param_optimizer if any(nd in n for nd in no_decay)],
'weight_decay_rate': 0.0}
]
# Note - `optimizer_grouped_parameters` only includes the parameter values, not
# the names.
| BERT_Fine_Tuning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
from PIL import Image
from IPython.display import display
import random
import json
import os
import pprint
# +
# path = "./trait-layers"
path = "./high-res-traits"
trait_directories = os.listdir(path)
trait_directories.remove(".DS_Store")
trait_directories.sort()
print(trait_directories)
trait_files = {}
for file in trait_directories:
if file in trait_files:
continue
else:
trait_files[file] = os.listdir(path + "/" + file)
for t in trait_files:
trait_files[t].sort()
print(t)
print(trait_files[t])
# +
# Each image is made up a series of traits
# The weightings for each trait drive the rarity and add up to 100%
traits = {}
background = ['01.png', '01b.png', '01c.png', '02a.png', '02b.png', '02c.png', '03a.png', '03b.png', '03c.png', '04a.png', '04b.png', '04c.png', '04d.png', '05a.png', '05b.png', '05c.png', '05d.png', '06a.png', '06b.png', '06c.png', '06d.png']
background_weights = [5, 5, 5, 5, 5, 5, 5, 5, 5, 5, 5, 5, 5, 5, 5, 5, 4, 4, 4, 4, 4]
base = ['01a.png', '01b.png', '01c.png', '01d.png', '01e.png', '02a.png', '02b.png', '02c.png', '02d.png', '02e.png', '03a.png', '03b.png', '03c.png', '03d.png', '03e.png', '04a.png', '04b.png', '04c.png', '04d.png', '04e.png']
base_weights = [25, 5, 5, 5, 5, 5, 5, 5, 5, 5, 4, 4, 4, 4, 4, 4, 4, 4, 4, 4]
eyebrow = ['01.png', '02.png', '03.png', '04.png']
eyebrow_weights = [25, 25, 25, 25]
eye = ['01.png', '02.png', '03.png', '04.png', '05.png']
eye_weights = [25, 20, 20, 20, 5]
lashes = ['', '01a.png', '01b.png', '01c.png', '01d.png', '01e.png']
lashes_weights = [30, 14, 14, 14, 14, 14]
nose = ['', '01.png']
nose_weights = [50, 50]
makeup = ['', '01.png', '02.png', '03.png', '04.png', '05.png', '06.png', '07.png', '08.png', '09.png', '10a.png', '10b.png', '10c.png', '10d.png', '11a.png']
makeup_weights = [40, 4, 4, 4, 4, 4, 5, 5, 5, 4, 4, 4, 4, 4, 4]
jewelry = ['', '01a.png', '01b.png', '01c.png', '01d.png', '01e.png', '02a.png', '02b.png', '02c.png', '02d.png', '02e.png', '03a.png', '03b.png', '03c.png', '03d.png', '04a.png', '04b.png', '04c.png', '04d.png', '04e.png', '05a.png', '05b.png', '05c.png', '05d.png', '05e.png', '06a.png', '06b.png', '06c.png', '06d.png', '06e.png']
jewelry_over = ['05a.png', '05b.png', '05c.png', '05d.png', '05e.png']
jewelry_weights = [10, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3]
clothes = ['', '01a.png', '01b.png', '01c.png', '01d.png', '01e.png', '02a.png', '02b.png', '02c.png', '02d.png', '03a.png', '03b.png', '03c.png', '03d.png', '03e.png', '04a.png', '04b.png', '04c.png', '04d.png', '04e.png', '05a.png', '05b.png', '05c.png', '05d.png', '05e.png', '06a.png', '06b.png', '06c.png', '06d.png', '06e.png', '07a.png', '07b.png', '07c.png', '07d.png', '07e.png', '07f.png', '08a.png', '08b.png', '09a.png', '09b.png', '09c.png', '10a.png', '10b.png', '10c.png', '10d.png', '11a.png', '11b.png', '11c.png', '11d.png', '12a.png', '12b.png', '12c.png', '12d.png', '13a.png', '13b.png', '13c.png', '14a.png', '14b.png', '14c.png', '14d.png', '14e.png', '14f.png', '15a.png', '15b.png', '15c.png', '15d.png', '15e.png']
clothes_no_hat = ['09a.png', '09b.png', '09c.png', '14a.png', '14b.png', '14c.png', '14d.png', '14e.png', '14f.png']
clothes_no_sunglasses = ['09a.png', '09b.png', '09c.png']
clothes_necklace = ['07a.png', '07b.png', '07c.png', '07d.png', '07e.png', '07f.png', '08a.png', '08b.png', '09a.png', '09b.png', '09c.png', '11a.png', '11b.png', '11c.png', '11d.png', '12a.png', '12b.png', '12c.png', '12d.png']
clothes_weights = [1, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1]
necklace = ['', '01.png', '02.png', '03a.png', '03b.png', '03c.png', '03d.png', '03e.png', '04a.png', '04b.png', '04c.png', '04d.png', '05a.png', '05b.png', '05c.png', '05d.png', '06a.png', '06b.png', '06c.png', '06d.png', '08d.png', '08e.png']
necklace_no_clothes = ['01.png']
necklace_weights = [60, 0.2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2]
hair = ['', '01a.png', '01b.png', '01c.png', '01d.png', '01e.png', '02a.png', '02b.png', '02c.png', '02d.png', '02e.png', '03a.png', '04a.png', '04b.png', '04c.png', '04d.png', '04e.png', '05a.png', '06a.png', '06b.png', '06c.png', '06d.png']
hair_no_hat = ['03a.png', '04a.png', '04b.png', '04c.png', '04d.png', '04e.png', '05a.png', '06a.png', '06b.png', '06c.png', '06d.png']
hair_no_necklace = ['05a.png']
hair_weights = [65, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 2, 1.5, 1.5, 1.5, 1.5, 1.5, 4.5, 1.5, 1.5, 1.5, 1.5]
hat = ['', '01a.png', '01b.png', '01c.png', '01d.png', '02.png', '03a.png', '03b.png', '03c.png', '03d.png', '03e.png', '04a.png', '04b.png', '04c.png', '04d.png', '04e.png', '05a.png', '05b.png', '05c.png', '05d.png', '06a.png', '06b.png', '06c.png', '06d.png', '06e.png', '07a.png', '07b.png', '07c.png', '07d.png', '07e.png', '07f.png', '08a.png', '08b.png', '08c.png', '08d.png', '09a.png', '09b.png', '09c.png', '09d.png', '10a.png', '10b.png', '10c.png', '10d.png', '11a.png', '11b.png', '11c.png', '11d.png', '12a.png', '12b.png', '12c.png', '12d.png']
hat_no_glasses = ['08a.png', '08b.png', '08c.png', '08d.png', '10a.png', '10b.png', '10c.png', '10d.png', '12a.png', '12b.png', '12c.png', '12d.png']
hat_no_har = []
hat_weights = [40, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1.5, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1]
rings = ['', '01a.png', '01b.png', '01c.png', '02a.png', '02b.png', '02c.png', '03a.png', '03b.png', '03c.png', '04a.png', '04b.png', '04c.png', '05a.png', '05b.png', '05c.png', '06a.png', '06b.png', '06c.png', '06d.png', '06e.png']
rings_weights = [40, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3]
sunglasses = ['', '01.png', '02a.png', '02b.png', '02c.png', '02d.png', '02e.png', '03.png', '04.png', '05a.png', '05b.png', '05c.png', '05d.png', '05e.png', '06a.png', '06b.png', '06c.png', '06d.png', '07a.png', '07b.png', '07c.png', '08a.png', '08b.png', '08c.png', '08d.png', '08e.png', '09a.png', '09b.png', '09c.png', '09d.png', '09e.png', '10a.png', '10b.png', '10c.png', '10d.png', '10e.png', '11a.png', '11b.png', '11c.png', '11d.png', '12a.png', '12b.png', '12c.png', '12d.png', '13a.png', '13b.png', '13c.png', '13d.png', '14a.png', '14b.png', '14c.png', '14d.png']
sunglasses_no_hat = ['07a.png', '07b.png', '07c.png', '11a.png', '11b.png', '11c.png', '11d.png']
sunglasses_mask_no_hair = ['07a.png', '07b.png', '07c.png']
sunglasses_weights = [52, 1, 1, 1, 1, 1, 1, 1, 0.5, 1, 1, 1, 1, 1, 1, 1, 1, 1, 0.2, 0.2, 0.2, 0.2, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1]
others = ['', '01a.png', '01b.png', '01c.png', '01d.png', '01e.png']
others_weights = [95, 1, 1, 1, 1, 1]
# Dictionary variable for each trait.
# Each trait corresponds to its file name
traits['background'] = background
traits['base'] = base
traits['eyebrow'] = eyebrow
traits['eye'] = eye
traits['lashes'] = lashes
traits['nose'] = nose
traits['makeup'] = makeup
traits['jewelry'] = jewelry
traits['clothes'] = clothes
traits['necklace'] = necklace
traits['hair'] = hair
traits['hat'] = hat
traits['rings'] = rings
traits['sunglasses'] = sunglasses
traits['others'] = others
print(traits)
directory_names = ['01_background', '02_base', '03_eyebrow', '04_eye', '05_lashes', '06_nose', '07_makeup', '08_jewelry', '09_clothes', '10_hair', '11_hat', '12_necklace', '13_rings', '14_sunglasses', '15_others']
directory_mappings = {}
for i, s in enumerate(directory_names):
for t in traits.keys():
if t in s:
directory_mappings[t] = directory_names[i]
print(directory_mappings)
# +
# Custom image
traits = {}
image = {}
image['background'] = '01c.png'
image['base'] = '04e.png'
image['eyebrow'] = '03.png'
image['eye'] = '02.png'
image['lashes'] = '01c.png'
# image['nose'] = '01.png'
image['makeup'] = '01.png'
image['jewelry'] = '01d.png'
# image['necklace'] = '06b.png'
image['clothes'] = '07f.png'
image['hair'] = '04d.png'
# image['hat'] = '04d.png'
# image['rings'] = '05b.png'
image['sunglasses'] = '11c.png'
# image['others'] = ''
imgs = []
for trait in image:
img = Image.open(f'./high-res-traits/{directory_mappings[trait]}/{image[trait]}').convert('RGBA')
imgs.append(img)
#Create each composite
start_image = Image.alpha_composite(imgs[0], imgs[1])
next_image = start_image
for idx, val in enumerate(imgs):
if idx < 2:
continue
else:
next_image = Image.alpha_composite(start_image, imgs[idx])
start_image = next_image
#Convert to RGB
rgb_im = next_image.convert('RGB')
# resize images for sampling. Remove below line for final images
smaller_img = rgb_im.resize((1024,1024),Image.ANTIALIAS)
file_name = 'thetechie.png'
smaller_img.save("./custom_images/" + file_name)
# +
## Generate Traits
TOTAL_IMAGES = 100 # Number of random unique images we want to generate
all_images = []
# A recursive function to generate unique image combinations
def create_new_image():
new_image = {} #
# For each trait category, select a random trait based on the weightings
new_image ["background"] = random.choices(background, background_weights)[0]
new_image ["base"] = random.choices(base, base_weights)[0]
new_image ["makeup"] = random.choices(makeup, makeup_weights)[0]
new_image ["eye"] = random.choices(eye, eye_weights)[0]
new_image ["hair"] = random.choices(hair, hair_weights)[0]
new_image ["hat"] = random.choices(hat, hat_weights)[0]
if new_image["hat"] in hat_special:
new_image ["necklace"] = necklace[0]
else:
new_image ["necklace"] = random.choices(necklace, necklace_weights)[0]
new_image ["rings"] = random.choices(rings, rings_weights)[0]
new_image ["sunglasses"] = random.choices(sunglasses, sunglasses_weights)[0]
if new_image in all_images:
return create_new_image()
else:
return new_image
# Generate the unique combinations based on trait weightings
for i in range(TOTAL_IMAGES):
new_trait_image = create_new_image()
all_images.append(new_trait_image)
# -
# +
# Returns true if all images are unique
def all_images_unique(all_images):
seen = list()
return not any(i in seen or seen.append(i) for i in all_images)
print("Are all images unique?", all_images_unique(all_images))
# -
# Add token Id to each image
i = 200
for item in all_images:
item["tokenId"] = i
i = i + 1
# + tags=[]
print(all_images)
# +
# Calculate Trait Counts
trait_stats = {}
sum_all_traits = 0
for trait in traits:
sum_all_traits += len(traits[trait])
trait_stats['sum_all_traits'] = sum_all_traits
print("Total items in collection: " + str(sum_all_traits))
trait_group_counts = {}
for trait in traits:
trait_group_counts[trait] = 0
trait_counts = {}
for trait_value in traits[trait]:
trait_counts[trait_value] = 0
trait_group_counts[trait] = trait_counts
for image in all_images:
for trait in trait_group_counts:
trait_group_counts[trait][image[trait]] += 1
print(trait_group_counts)
# +
## Calculate rarity scores
rarity_scores = {}
for trait, values in trait_group_counts.items():
rarity_scores_traits = {}
for val in values:
if values[val] == 0:
next
else:
print(trait)
print(values)
rarity_score = 1 / (values[val] / TOTAL_IMAGES)
rarity_scores_traits[val] = { 'rarity_score': rarity_score, 'count': values[val]}
rarity_scores[trait] = rarity_scores_traits
print(rarity_scores)
## Save meta stats
with open('./metadata/all-stats.json', 'w') as outfile:
json.dump(rarity_scores, outfile, indent=4)
# -
#### Generate Metadata for all Traits
METADATA_FILE_NAME = './metadata/all-traits.json';
with open(METADATA_FILE_NAME, 'w') as outfile:
json.dump(all_images, outfile, indent=4)
# +
#### Generate Images
for item in all_images:
print(item)
imgs = []
for trait in item:
if trait == 'tokenId':
continue
# don't create images which have a blank option
if item[trait] == '':
continue
# img = Image.open(f'./trait-layers/{directory_mappings[trait]}/{item[trait]}').convert('RGBA')
img = Image.open(f'./high-res-traits/{directory_mappings[trait]}/{item[trait]}').convert('RGBA')
# img = img.resize((400,400),Image.ANTIALIAS)
imgs.append(img)
#Create each composite
start_image = Image.alpha_composite(imgs[0], imgs[1])
next_image = start_image
for idx, val in enumerate(imgs):
if idx < 2:
continue
else:
next_image = Image.alpha_composite(start_image, imgs[idx])
start_image = next_image
#Convert to RGB
rgb_im = next_image.convert('RGB')
# resize images for sampling. Remove below line for final images
smaller_img = rgb_im.resize((1024,1024),Image.ANTIALIAS)
file_name = str(item["tokenId"]) + ".png"
smaller_img.save("./images/" + file_name)
# +
#### Generate Metadata for each Image
f = open('./metadata/all-traits.json',)
data = json.load(f)
IMAGES_BASE_URI = "ADD_IMAGES_BASE_URI_HERE"
PROJECT_NAME = "ADD_PROJECT_NAME_HERE"
def getAttribute(key, value):
return {
"trait_type": key,
"value": value
}
for i in data:
token_id = i['tokenId']
token = {
"image": IMAGES_BASE_URI + str(token_id) + '.png',
"tokenId": token_id,
"name": PROJECT_NAME + ' ' + str(token_id),
"attributes": []
}
for attr in i:
token["attributes"].append(getAttribute(attr, i[attr]))
print(token)
with open('./metadata/' + str(token_id), 'w') as outfile:
json.dump(token, outfile, indent=4)
f.close()
# -
| .ipynb_checkpoints/custom-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="cVD54YAodIu3" colab_type="text"
# # try...except...else
# + [markdown] id="n5v6L8TgfgDs" colab_type="text"
# else bloğu eğer herhangi bir hata olmazsa şu kodu çalıştır anlamında kullanılır çoğunlukla
# + id="RkFd2sC4fpnS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="6a15484f-76ed-4006-dbc2-51e30e044f3b"
try:
print("Hello")
except:
print("something went wrong")
else:
print("Nothing went wrong")
# + [markdown] id="j7r5dN6Gf5lC" colab_type="text"
# veya kodları parçalarak ayrı ayrı hata mesajı verdirebilir
# + id="bVZve2mkgClk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="76e9184e-8024-43fb-9d4f-ca1ae820b2b9"
try:
bolunen = int(input("Bolunecek sayıyı giriniz: "))
bolen = int(input("Bölecek sayıyı giriniz: "))
except ValueError:
print("Lütfen sadece sayı değerini giriniz: ")
else:
try:
print(bolunen/bolen)
except ZeroDivisionError:
print("Bir sayıyı 0'a bölemezsiniz!")
# + [markdown] id="Lv_Kcypcg6wQ" colab_type="text"
# # + hata vereceğini düşündüğümüz kodları try bloğun içine yazdık
# # + beklediğimiz hata ise ValueError olduğu için except bloğu sayesinde, eğer bu hataya ulaşırsan şunu yap dedik
# # + daha sonra else: print(bolunen/bolen) yazdık böylece eğer hata almazsan iki sayıyı böl dedik
# # + fakat bir hata daha olduğu için ayrı bir blog daha oluştururak else: bloğunda yazılan kodu try bloğun içine aldık
# # + daha sonra şu hataya karşılaşınca şunu yazdır dedik
# + [markdown] id="cnPqtDdZhqTO" colab_type="text"
# Yukarıdaki kodu şu şekil yazabilirdik:
# + id="ILYf-v7Shs-q" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="03cff300-906e-490a-f97c-eee7e5dddd26"
try:
bolunen = int(input("Bölünecek sayıyı giriniz: "))
bolen = int(input("Bölecek sayıyı giriniz: "))
print(bolunen/bolen)
except ValueError:
print("Sadece sayı giriniz!")
except ZeroDivisionError:
print("bir sayıyı 0 ile bölemezsiniz!")
| hataYakalama/tryExceptElse.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import trustedanalytics as ta
ta.connect()
# ### Create a frame with data that we'll use to train the ARX model
#
# The frame has columns for the observed value "y" and several other columns that contain exogenous variables (visitors, weekends, seasonality, etc).
#
schema = [("y", ta.float64),("visitors", ta.float64),("wkends", ta.float64),("seasonality", ta.float64),("incidentRate", ta.float64), ("holidayFlag", ta.float64),("postHolidayFlag", ta.float64),("mintemp", ta.float64)]
csv = ta.CsvFile("train_atk.csv", schema=schema, skip_header_lines=1)
frame = ta.Frame(csv)
frame.inspect()
# ### Create and train the model
#
# Create an ARX model, and then train the model by providing the frame of data, the "y" column, a list of "x" columns, y max lag, x max lag, and a boolean flag indicating if the intercept should be dropped.
#
# The ARX model train() return 'c' (an intercept term, or 0 for no intercept) and a list of coefficients (one for each "x" column).
#
arx = ta.ArxModel()
y_column = "y"
x_columns = ["visitors","wkends","seasonality","incidentRate","holidayFlag","postHolidayFlag","mintemp"]
y_max_lag = 0
x_max_lag = 0
no_intercept = True
arx.train(frame, y_column, x_columns, y_max_lag, x_max_lag, no_intercept)
# So, in this example the coefficients are:
#
# | x | coefficient |
# |----------------|------------------------|
# | vistors | -1.136026484226831e-08 |
# |wkends | 8.637677568908233e-07 |
# |seasonality | 15238.143039368977 |
# |incidentRate | -7.993535860373772e-09 |
# |holidayFlag | -5.198597570089805e-07 |
# |postHolidayFlag | 1.5691547009557947e-08|
# |mintemp: | 7.409621376205488e-08 |
#
#
# ### Create a frame that contains test data
#
# The test data is in test_atk.csv and has the same schema that we used for training.
test_csv = ta.CsvFile("test_atk.csv", schema=schema, skip_header_lines=1)
test_frame = ta.Frame(test_csv)
test_frame.inspect()
# ### Predict
#
# Using the frame of test data, run ARX predict().
p = arx.predict(test_frame, y_column, x_columns)
p.inspect(n=p.row_count,columns=["y", "predicted_y"])
arx.publish()
| python-client/ARX Demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="HXCjXrtz19MK"
import pandas as pd
import numpy as np
import time
import concurrent.futures
# + id="IP6GLtL_pQ-M"
# !pip install -U -q PyDrive
# + id="H_Vas2Ovpowk"
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
# + id="GyXbPTmmpqAd"
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
# + id="HfLihr06pq_V"
drive.CreateFile({'id':'1YAAvPUaPeBIddkuVzWgw7fhPPcT2uJTY'}).GetContentFile('billboard_dataset_unique.csv')
df_billboard = pd.read_csv("billboard_dataset_unique.csv").drop('Unnamed: 0',axis=1)
drive.CreateFile({'id':'1kEYc4W4-rPhWtFJYpkDAWaAOGGO-eB3B'}).GetContentFile('dataset2.csv')
df_songs = pd.read_csv("dataset2.csv")
# + id="CbKmFBMdD9uf"
df_songs_bak = df_songs.copy()
# + id="VfPM3gF7_VD4"
# elimino tracce precedenti al 1960
mask = df_songs.year > 1959
df_songs = df_songs[mask]
# converto colonna 'release_date' in datetime
df_songs.release_date = pd.to_datetime(df_songs.release_date,format="%Y-%m-%d",exact=False)
# ordino il df per data
df_songs = df_songs.sort_values('release_date',ascending=False)
# elimino eventuali id ripetuti
df_songs = df_songs.drop_duplicates(subset='id',keep='last') # keep='last' considerando che ho ordinato il df in senso discendente
# elimino ripubblicazioni (tenendo solo traccia più vecchia)
df_songs = df_songs.drop_duplicates(subset=['artists','name'],keep='last') # keep='last' considerando che ho ordinato il df in senso discendente
# converto colonna artists da stringa a lista di stringhe
df_songs.artists = df_songs.artists.apply(eval)
# + id="WYllj5iRH1OQ"
# inserisco i due target 'hit' e 'weeks' nel dataframe delle canzoni
# 1) creo 2 nuove colonne nel dataframe delle canzoni (df_songs)
hit_array = np.zeros(df_songs.id.count())
weeks_array = np.zeros(df_songs.id.count())
df_songs.insert(19,'hit',hit_array)
df_songs.insert(20,'weeks',weeks_array)
# + id="lFT-2Ol4AMEs"
def hit_match(i,df_billboard,df_songs):
entry_title = df_billboard.title[i]
entry_weeks = df_billboard.weeks[i]
entry_artist = df_billboard.artist[i]
output = {}
print("Checking %s by %s ..." % (entry_title,entry_artist))
mask = df_songs.name.str.lower() == entry_title.lower()
sub_df = df_songs[mask] # creo sotto-dataframe (di df_songs) con le canzoni con lo stesso titolo della hit in esame
for k in range(sub_df.shape[0]): # per ogni traccia del sotto-dataframe (sub_df)
artist_list = sub_df.artists.iloc[k]
id = sub_df.id.iloc[k]
for j in range(len(artist_list)):
if(artist_list[j] in entry_artist):
match = True
else:
match = False
break
if(match == True):
print("*** %s by %s it's a HIT! ***" % (entry_title,entry_artist))
output.update({id:entry_weeks})
return output
# + id="yxRQJbXZ1qXt" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1611095808300, "user_tz": -60, "elapsed": 10880311, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhsE3N_0rvi4i7PTUy9okHWPKo4CtX-Vq2O3tR8K8M=s64", "userId": "14558454588425913231"}} outputId="b260fd0b-38e0-4ab4-95bf-b1438cd266e6"
time_0 = time.perf_counter()
with concurrent.futures.ProcessPoolExecutor() as executor:
results = [executor.submit(hit_match, i,df_billboard,df_songs) for i in range(df_billboard.shape[0])]
hit_dict = {}
for f in concurrent.futures.as_completed(results):
hit_dict.update(f.result())
print("Completato in %.4f secondi" % (time.perf_counter()-time_0))
# + id="FIVzkFbs0EPG"
df_songs_bak = df_songs.copy()
# + id="8MpTpz-TX43K"
for key in list(hit_dict):
mask = df_songs.id == key
df_songs[mask] = df_songs[mask].apply(lambda x: 1 if x.name == 'hit' else x)
df_songs[mask] = df_songs[mask].apply(lambda x: hit_dict[key] if x.name == 'weeks' else x)
# + id="r4fyeOr_5_w_" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1611101431091, "user_tz": -60, "elapsed": 868670, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhsE3N_0rvi4i7PTUy9okHWPKo4CtX-Vq2O3tR8K8M=s64", "userId": "14558454588425913231"}} outputId="fefa5ffa-513a-4381-f549-98281b2095a1"
from google.colab import drive
# mounts the google drive to Colab Notebook
drive.mount('/content/drive',force_remount=True)
# + id="WoBvYLdr6GtI"
df_songs.to_csv('/content/drive/My Drive/Colab Notebooks/datasets/dataset2_X_billboard.csv')
# + colab={"base_uri": "https://localhost:8080/"} id="TeS5dlhbE6a-" executionInfo={"status": "ok", "timestamp": 1611101571122, "user_tz": -60, "elapsed": 673, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhsE3N_0rvi4i7PTUy9okHWPKo4CtX-Vq2O3tR8K8M=s64", "userId": "14558454588425913231"}} outputId="3d9f6540-15f0-4289-844f-d88dfea01256"
df_songs.id.count()
# + id="PSzAArMCImkJ"
| dataset_merging.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# name: python3
# ---
# - https://www.six-sigma-material.com/Proportions-Tests.html
# - https://online.stat.psu.edu/statprogram/reviews/statistical-concepts/proportions
# - each pValue < 0.01 meaning we are even 99% sure that null hypo must not be true
# - I have tried all methods but all reject null hypo
import pandas as pd
from scipy import stats
import numpy as np
# for population data
cols=['State', 'Level', 'Name', 'TRU', 'No_HH', 'TOT_P', 'TOT_M', 'TOT_F']
census=pd.read_excel('datasets/census.xlsx',engine='openpyxl',usecols=cols)
censusIndia=census.iloc[0,:]
census=census.loc[census.Level=='STATE']
census=census.loc[census.TRU=='Total']
census=census.append(censusIndia,ignore_index=True)
census.sort_values(by=['State'],axis=0,inplace=True)
census.reset_index(drop=True,inplace=True)
# def pTester(ratio_1,ratio_2,totalPop):
# p0=(ratio_2+ratio_1)/2 # under null hypo
# zStat=(ratio_1-p0)/np.sqrt(p0*(1-p0)/totalPop)
# pValue=stats.distributions.norm.cdf(-np.abs(zStat))
# return pValue
# def pTester(ratio_1,ratio_2,totalPop):
# p0=0.5 # under null hypo
# zStat=(ratio_1-p0)/np.sqrt(p0*(1-p0)/totalPop)
# pValue=stats.distributions.norm.cdf(-np.abs(zStat))
# return pValue
# #===================
# # # a function that just takes state code that does
# # manipulates and finds appropriate sum
# # finds total pop fro that state from census data
# # then returns a nice ratio dictionary
# #===================
#
# def ratioFinder(stateCode):
# # 5 is specific to TOT_P
# statePopMale=census.iloc[stateCode,6]
# statePopFemale=census.iloc[stateCode,7]
# # modify state code
# stateCode=str(stateCode).zfill(2)
# # read df from file
# df=pd.read_excel(f'datasets/C-17/{stateCode}.xlsx',skiprows=6,header=None,engine='openpyxl')
# df.fillna(value=0,inplace=True)
# # find total no
# # note: these col numbers are specific to df #
# male=sum(df.iloc[:,15].tolist())
# female=sum(df.iloc[:,16].tolist())
# #========
# # should I use statePops or (male+female) for 3+ langs to devide
# # right now I am doing thid with male+female for 3+ langs
# # in fact I feel why I am including whole state pops to devide my study pop that is persons 3+
# #========
# totalPop=male+female
# #totalPop=statePopFemale+statePopMale
# ratio_1=(male/totalPop) # no rounding off
# ratio_2=(female/totalPop)
# #pValue=pTester(ratio_1,ratio_2,totalPop) # func that gives pValue for ratios
# pValue=pTester(male,female,totalPop)
# #store ratios in a dict
# stateRatios={
# 'StateCode':stateCode,
# 'male-percentage':ratio_1,
# 'female-percentage':ratio_2,
# 'p-value':pValue
# }
# #print('Processed state -- ',stateCode)
# return stateRatios
# #===================
# # # a function that just takes state code that does
# # manipulates and finds appropriate sum
# # finds total pop fro that state from census data
# # then returns a nice ratio dictionary
# #===================
# ####based on: https://online.stat.psu.edu/stat415/lesson/9/9.4####
# def ratioFinder(stateCode):
# # 5 is specific to TOT_P
# statePopMale=census.iloc[stateCode,6]
# statePopFemale=census.iloc[stateCode,7]
# # modify state code
# stateCode=str(stateCode).zfill(2)
# # read df from file
# df=pd.read_excel(f'datasets/C-17/{stateCode}.xlsx',skiprows=6,header=None,engine='openpyxl')
# df.fillna(value=0,inplace=True)
# # find total no
# # note: these col numbers are specific to df #
# male=sum(df.iloc[:,15].tolist())
# female=sum(df.iloc[:,16].tolist())
# #========
# # should I use statePops or (male+female) for 3+ langs to devide
# # right now I am doing thid with male+female for 3+ langs
# # in fact I feel why I am including whole state pops to devide my study pop that is persons 3+
# #========
# totalPop=male+female
# #totalPop=statePopFemale+statePopMale
# ratio_1=(male/statePopMale) # no rounding off
# ratio_2=(female/statePopFemale)
# #pValue=pTester(ratio_1,ratio_2,totalPop) # func that gives pValue for ratios
# #pValue=pTester(male,female,totalPop)
# ratio=totalPop/(statePopFemale+statePopMale)
# zStat=(ratio_1-ratio_2)/(np.sqrt((ratio*(1-ratio))*((1/statePopFemale)+(1/statePopMale))))
# pValue=2*stats.distributions.norm.cdf(-np.abs(zStat))
# #store ratios in a dict
# stateRatios={
# 'StateCode':stateCode,
# 'male-percentage':ratio_1,
# 'female-percentage':ratio_2,
# 'p-value':pValue
# }
# #print('Processed state -- ',stateCode)
# return stateRatios
#===================
# # a function that just takes state code that does
# manipulates and finds appropriate sum
# finds total pop fro that state from census data
# then returns a nice ratio dictionary
#===================
####based on chi quare test with df=1
def ratioFinder(stateCode):
# 5 is specific to TOT_P
statePopMale=census.iloc[stateCode,6]
statePopFemale=census.iloc[stateCode,7]
# modify state code
stateCode=str(stateCode).zfill(2)
# read df from file
df=pd.read_excel(f'datasets/C-17/{stateCode}.xlsx',skiprows=6,header=None,engine='openpyxl')
df.fillna(value=0,inplace=True)
# find total no
# note: these col numbers are specific to df #
male=sum(df.iloc[:,15].tolist())
female=sum(df.iloc[:,16].tolist())
#========
# should I use statePops or (male+female) for 3+ langs to devide
# right now I am doing thid with male+female for 3+ langs
# in fact I feel why I am including whole state pops to devide my study pop that is persons 3+
#========
totalPop=male+female
#totalPop=statePopFemale+statePopMale
ratio_1=(male/statePopMale) # no rounding off
ratio_2=(female/statePopFemale)
#pValue=pTester(ratio_1,ratio_2,totalPop) # func that gives pValue for ratios
#pValue=pTester(male,female,totalPop)
ratio=(ratio_1+ratio_2)/2
zStat=((ratio_1-ratio)**2+(ratio_2-ratio)**2)/ratio
pValue=1-stats.distributions.chi2.sf(np.abs(zStat),1)
#store ratios in a dict
stateRatios={
'state-code':stateCode,
'male-percentage':ratio_1,
'female-percentage':ratio_2,
'p-value':pValue
}
#print('Processed state -- ',stateCode)
return stateRatios
# %%time
ratioList=[ratioFinder(i) for i in range(0,36)]
ratioDF=pd.DataFrame(ratioList)
ratioDF
#based on chi-square
# but it is not for this
| Q2_asgn2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
plt.rcParams["figure.figsize"] = (16, 9)
plt.close("all")
# -
bikes = pd.read_csv("../data/bikesharing/data.csv")
bikes["timestamp"] = pd.to_datetime(bikes["timestamp"], format="%Y-%m-%d %H:%M:%S")
bikes.head()
# Column description:
#
# - "timestamp" - timestamp field for grouping the data
# - "cnt" - the count of a new bike shares
# - "t1" - real temperature in C
# - "t2" - temperature in C "feels like"
# - "hum" - humidity in percentage
# - "windspeed" - wind speed in km/h
# - "weathercode" - category of the weather
# - "isholiday" - boolean field - 1 holiday / 0 non holiday
# - "isweekend" - boolean field - 1 if the day is weekend
# - "season" - category field meteorological seasons: 0-spring ; 1-summer; 2-fall; 3-winter.
# # Exercise 1) Scatter plot
# a) create a figure with one axis instance. You can either use plt.figure() or use plt.subplots(). Get comfortable with both methods
# b) create a scatter plot where you plot the humidity (hum) vs the real temperature (t1) column. HINT: you can show the figure by calling the figure instance at the end of the cell
# c) label the x and y axis with their respective names. HINT: use the .set_xlabel(x_name) and .set_ylabel(x_name) label methods
# d) give the axis instance a title. HINT: use .set_title(axis_title)
# e) save your figure as a .png file. HINT: have a look at the matplotlib documentation (https://matplotlib.org/stable/api/_as_gen/matplotlib.pyplot.savefig.html)
# # Exercise 2) Line plot
# a) use a lineplot to visualize the count of new bike shares (cnt column) before April 2015. HINT: you need to subset the dataframe by using the timestamp column
# b) place a red marker on each point where the respective timestamp belongs to the weekend (is_weekend == 1)
# 1. identify the rows where is_holiday == 1 (i.e. create a mask)
# 2. use a scatter plot to visualize the points that are observed on the weekend. HINT: (x=timestamp, y=cnt) and for controlling the marker size and color see (https://matplotlib.org/stable/api/_as_gen/matplotlib.pyplot.scatter.html)
# c) what happens if you use .plot(...) instead of .scatter(...) as in b). HINT: recreate the figure that you created in exercise 2a)
| lesson05/exercises/exercises_visualisation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Load the data set
# %run datasetup.py
# import statements
import pandas as pd
# %matplotlib notebook
import zipfile
# Let's inspect the zipfile
fname = 'data/household_power_consumption.zip'
with zipfile.ZipFile(fname, mode = 'r') as fpzip:
print(fpzip.infolist())
print(fpzip.namelist())
with fpzip.open(fpzip.namelist()[0],'r') as fp:
counter = 0
for line in fp:
print(line)
counter += 1
if counter == 5:
break
# There is only one file in the zip archive so Let's try to read it with pandas
df = pd.read_csv(fname, sep=';', compression='zip', nrows=10000)
df.head()
# Let's compute the memory of the full dataset
dftmp = pd.read_csv(fname, sep=';', compression='zip')
print(f'Size of data in memory: {dftmp.memory_usage(deep=True).sum() * 1.e-9} GB')
print(dftmp.shape)
del dftmp
# Let's improve the csv reading
df = pd.read_csv(
fname,
sep=';',
compression='zip',
nrows=10000,
parse_dates={'ts':['Date','Time']},
index_col='ts',
dayfirst=True,
na_values='?'
)
df.head()
df.index
df.isnull().any()
df.dtypes
# How is the voltage distribution hourly?
df.groupby(df.index.hour)['Voltage'].agg(['min', 'max', 'mean']).plot.bar()
# +
# write to a CSV it's as easy as this
# df.to_csv('my_first_csv_with_pandas.csv')
| talks/Feb2019-Intro-Pandas-Dask/Example-CSV-time-series.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline #this is a formatting statement for jupyter notebooks
#
# # Transitioning from using IDL to using Python for your science
#
#
# This is an introduction to moving your science from an IDL environment to Python. If you havne't done so yet, please download and install the the latest versions of anaconda (www.continuum.io/downloads) and sunpy (docs.sunpy.org/en/stable/guide/installation/).
#
# Let's make sure that your instillation of anaconda is up to date. From the terminal, type:
#
# + active=""
# conda update conda
# -
#
# + active=""
# conda update anaconda
# -
# Now, lets launch an ipython (interactive python) session
# + active=""
# ipython
# -
# # Getting started with Python
# The first thing to note is that by itself, Python is just a platform. You can think of Python as an engine - it is very powerful but unless you connect it to a transmission, axle, and wheels, it isn't very effective at getting you down the road.
#
# Fortunatly in most cases you don't need to reinvent the wheel and you can just find some that someone else created that will fit your needs.
#
# The Anaconda package comes with literally thousands of tools to help you work with data.
#
# Let's explore some basic tools. NumPy is the fundamental package for scientific computing with Python. Let's load it in to our current session.
import numpy as np
# Notice the syntax we are using: import "the library" as "what we want to call it."
# Now whenever we want to call a tool in numpy, we can use "np."
#
# Next we will want to plot our data, so lets grab some plotting tools. We will use a package called matplotlib. Matplotlib is a 2D plotting library, which produces all kinds figures. We don't need everything that matplotlib can do, so let's import just the tool that produces line plots. This tool is called pyplot.
import matplotlib.pyplot as plt
# Again notice the syntax we are using: import "library.tool" as "what we want to call it."
#
# Okay, we have the tools 'np' and 'plt' defined. Loading in the tools and libraries you want to use is something you have to do every time you write a program or load a python session.
#
# Now lets put our tools to use:
dt = 0.1
Fs = 1/dt
t=np.arange(0,Fs,dt)
# We can dynamically define variables (i.e. we don't have to specify the type) and use conventional assignments to make new variables.
#
# We also use a tool within np to create t: np.arange. Np.arange is similar to IDL's FINDGEN( ). It creates an evenly spaced array with the syntax of output=np.arange(start,stop,step).
#
# So what does t look like? We just type the variable or use a print statement:
t
print(t)
# But what are the dimensions of t? How many elements are in it?
#
# In IDL we would have to call a function to return this information, such as MAX( ). In Python, since it is an object oriented programming language, the variable t is actually more than just a simple array. It is self-aware! So let's ask t about itself.
t.min
# Um... what just happened here? Python just said "yep, that is a valid function of t" but didn't actually return the value. To get the value out, we need to use a different syntax:
t.min()
# Also we can ask:
t.max()
t.mean()
t.var()
# These are all "t.something( )" - "something( )" is a function of the object "t".
# To find out how many elements are the array
t.size
# To see the shape of the array:
t.shape
# If you want to see all of the functions already available in t, type "t." and then 'tab'. This will show you dynamically all of the functions that exist. Go ahead, try it!
#
# You'll note that .size and .shape don't have brackets after them, but .min and .var do. This is because the size and the shape of the array are attributes of the array. The reason .min (and other methods) have brackets is that these functions have different options that you may want to use.
#
# Let's create a 2d random array
#
rand_nn = np.random.randn(t.size, t.size)
# We can find the minimum values along one dimension of the array...
rand_nn.min(axis=0)
# or the other one...
rand_nn.min(axis=1)
# Or the whole array.
rand_nn.min(), rand_nn.min(axis=None)
# The min( ) function with the axis option finds the minimum value along different dimensions of the array rand_nn.
# We used "axis=None". There is a special value in Python called **None**, which indicates **no value**. In this case, this means that axis has no assigned value. The axis function then resorts to a default behavior, which is to give you the minimum value over the entire array.
#
# Now let's create an array of random numbers with the same number of elements as t. We will use the Numpy tool 'random', but there are lots of ways to generate random numbers. In IDL we might use RANDOMN( ) or RANDOMU( ) depending on the distribution we want to sample from. Numpy.random had many ways of calculating random numbers too.
#
# Try typing "np.random." and then 'tab'.
#
# Let's sample from a normal distribution:
rand_n = np.random.randn(t.size)
# Notice that we can use the t.size attribute of 't' without having to define another variable or call a separate function.
#
# Let's plot our variables. To do this we will use the matplotlib's plot function.
plt.plot(t,rand_n)
plt.show()
# Matplotlib generates a plot and then in a separate command you have to display it to the screen. Yeah, it's a pain but you get use to it.
#
# Let's now generate some more complex data:
r = np.exp(-t/0.05)
r_dat = np.convolve(rand_n, r)*dt
r_dat.size
# The convolution returned an array that is twice the size of our other working data sets. We should trim that down. The syntax for segmenting an array uses square brackets:
r_dat=r_dat[:t.size]
# This means we want r_dat from the beginning element to t.size (which is 100). Let's create one more data set to work with using some more of NumPy's functions (sin and pi).
ss = 0.1*np.sin(2*np.pi*t) + r_dat
# What does all of this generated data look like? Why don't we make five different plots to highlight some different plotting capabilities.
#
# First, we will define a plotting space, but instead of showing the plot this time, we are going to plot some more values. First, we will plot the magnitude spectrum of the generated data with a linear scale and a dB (logarithmic) scale. This can all be done within matplotlib. Next, We can easily show the wrapped and unwrapped phase spectrum.
# The 'subplot' command means that we want a 3 by 2 grid of plots and we want to operate on the first sextant.
# +
plt.subplot(3, 2, 1)
plt.plot(t, ss)
plt.subplot(3, 2, 3)
plt.magnitude_spectrum(ss, Fs=Fs)
plt.subplot(3, 2, 4)
plt.magnitude_spectrum(ss, Fs=Fs, scale='dB')
plt.subplot(3, 2, 5)
plt.angle_spectrum(ss, Fs=Fs)
plt.subplot(3, 2, 6)
plt.phase_spectrum(ss, Fs=Fs)
plt.show()
# -
# Using matplotlib, we can showcase our generated data set in several different ways. Of course this is just an example and there are many more types of plots that matplotlib can generate.
# # Basic Programming Techniques in Python
#
# To showcase some other basic programming techniques using the skills we just learned, lets analytically calculate $\pi$.
#
# The easiest way of doing this is using the Monty Carlo Method through finding random points inside a circle inscribed in a square.
#
import numpy as np #always good to be explicit about our libraries used
# Next, we will initialize our counting variables
total = 0
inside = 0
# Now we will set up a for-loop to calculate our points. Loops in Python are a bit different than in IDL. First, Notice that in Python loops are tab delimited. Unlike IDL there is no need for 'begin' and 'end' statements since the tabbing indicates where the loop starts and ends.
for ii in range(10000):
# Generate two coordinates in the interval 0-1:
x_coord = np.random.uniform()
y_coord = np.random.uniform()
# Calculate the distance to the origin
r = np.sqrt(x_coord**2 + y_coord**2)
# Count this if it is inside the circle.
if r< 1:
inside += 1
total += 1
# You will see somethig else too: the variable we are iterating over, 'ii', is nowhere to be found in the loop. This is because the iteration of 'ii' over the list that the range( ) function produces is implicit. We can make the interation explicit using a slightly different syntax.
print('Pi=',4.0*inside/(1.0*total))
# In this case, 'iteration' now contains the iteration number (0,1,...,9999) while ii contains what ever is in the list at that iteration. This could be any data type. To drive this point home, let's show a trivial example:
for value in 'Jack':
print(value)
print
for iteration, value in enumerate('Jack'):
print(iteration, value)
# A few other things to notice: we are using np.random again, as well as np.sqrt. To define an exponent python uses a double astrix. We also encounter an if statement. For conditionals, python drops the 'then' from the statement and uses tabbing to denote the begining and ending.
# # Writing A Function
#
# Lets take this same python script and make it into a simple function we can iterate over.
import numpy as np #always good to be explicit about our libraries used
def pitest():
# Create the coordinate
xtemp = np.random.uniform()
ytemp = np.random.uniform()
# Calculate the distance to the origin
r = np.sqrt(xtemp**2 + ytemp**2)
# Count this if it is inside the circle.
if r< 1:
return True
return False
# The Python syntax for functions use the same tabbing to start and end the function that loops and conditional statemetns do. This particular function pitest( ) does not have any variable input, but if it did, it would go in the parenthesis. Now let's put the new function in a loop and see how it works:
# +
count = 0
inside = 0
for i in range(10000):
is_inside = pitest()
inside += is_inside
count += 1.
print('Pi=',((inside/count)*4.))
# -
# Python has boolean data types, **True** and **False**. These are also understood to have the integer values 1 and 0 respectively.
# # Running Your Own Function
# So now I have writen a script, how do I run it from the command line? Let's say that the function we wrote, __'pitest,'__ is saved with the name __'Pifunction.py'__. First, change to the directory where your program is located.
# + active=""
# cd /Users/mkirk/code/python/
# -
# Then import the name of your script.
# + active=""
# import Pifunction as Pifunc
# -
# And then run your function like we have all along:
# + active=""
# Pifunc.pitest()
# -
| SDO2016_Py1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 2d Plotting
#
# - [Download the lecture notes](https://philchodrow.github.io/PIC16A/content/np_plt/plt_2.ipynb).
#
# Often we want to visualize data with two independent variables. In this case, your "go-to" function should usually be `plt.imshow()`. Let's see how it works.
import numpy as np
from matplotlib import pyplot as plt
# ## Mathematical Functions
# Here's how we'll plot the function
#
# $$z = \sin(x) + \cos(y)$$
# horizontal and vertical axes
x = np.linspace(0, 2*np.pi, 101)
y = np.linspace(0, 2*np.pi, 101)
# +
# meshgrid turns each into a 2d array
X,Y = np.meshgrid(x,y)
fig, ax = plt.subplots(1, 2)
ax[0].imshow(X)
ax[1].imshow(Y)
# -
# compute the function values
Z = np.sin(X) + np.cos(Y)
# +
fig, ax = plt.subplots(1)
im = ax.imshow(Z)
# add a colorbar to help interpret the scales.
plt.colorbar(im)
# -
# ## Example 2: the Mandelbrot Set
# +
# create the background grid, higher resolution this time
x = np.linspace(-1, 1, 1001)
y = np.linspace(-1.5, .5, 1001)
X, Y = np.meshgrid(x, y)
# create the Mandelbrot set: start with a 2d grid of
# complex constants C:
C = X*1j + Y
# initialize a dummy value of Z
Z = 0
# repeatedly square Z and add constant C to it
# Mandelbrot set is the set of values of C such that the
# resulting value of Z is finite.
for i in range(200):
Z = Z**2 + C
# -
fig, ax = plt.subplots(1, figsize = (10, 10))
ax.imshow(abs(Z), interpolation = "none")
ax.axis('off') # turn off the surrounding frame
# ## Plotting Images
#
# The `ax.imshow()` method is actually strikingly versatile, and can plot RGB color data when used with `numpy` arrays of the correct data type. This allows us to use `imshow()` to inspect images, for example, those that we read in from files.
# +
import matplotlib.image as mpimg
img=mpimg.imread('semicolons.jpg')
# This is a 735x500 image. Each pixel has an RGB value, represented as 3 numbers
img.shape
# -
fig, ax = plt.subplots(1, figsize = (6, 4.5))
ax.imshow(img)
fig.suptitle("C++ programmers be like")
| content/np_plt/plt_2.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.6.0
# language: julia
# name: julia-0.6
# ---
# # Setup environment and load data
# +
using ClobberingReload
@ausing GERDADeepLearning
env = DLEnv();
data = get(env, "preprocessed")
detector_names = detectors(env, "coax")
filter!(data, :detector_name, det -> det ∈ detector_names)
filter!(data, :isMuVetoed, 0)
filter!(data, :isLArVetoed, 0)
filter!(data, :multiplicity, 1)
filter!(data, :E, E -> (E>500)&&(E<9999))
filter!(data, :FailedPreprocessing, 0)
# -
# # Calculate efficiencies for coax
# +
use_gpus(env)
after_cut_90 = EventLibrary[]
for i in 1:length(detector_names)
ch_data = data[:detector_name=>detector_names[i]]
ch_data = scale_waveforms(ch_data, 256)
new_properties!(identity, env, "autoencoder", "autoencoder-det-$(detector_names[i])")
net = autoencoder(env, ch_data; id="autoencoder-det-$(detector_names[i])", action=:load)
compact = encode(ch_data, net)
new_properties!(identity, env, "latent-dnn-classifier", "latent-dnn-classifier-det-$(detector_names[i])")
dnn = dnn_classifier(env, compact; id="latent-dnn-classifier-det-$(detector_names[i])", action=:load, train_key=nothing, xval_key=nothing, evaluate=["phy"])
ctest = flatten(compact)
label_energy_peaks(ctest)
effs = load_effs(env, detector_names[i]);
ind, cut_value, sig_eff, bkg_rej = background_rejection_at(0.9, effs)
cut_events = filter(ctest, :psd, psd->psd>cut_value)
push!(after_cut_90, cut_events)
println("Detector $(detector_names[i]): $(eventcount(cut_events)) / $(eventcount(ch_data)) (cut value $cut_value)")
end
after_cut_90 = cat_events(after_cut_90...);
# -
# # Plot physics spectrum
# +
using Plots, StatsBase, LaTeXStrings
bins = linspace(500, 5000, 91)
x_axis = bins[1:end-1]
hist_before = fit(Histogram, data[:E], bins, closed=:left)
hist_after_90 = fit(Histogram, after_cut_90[:E], bins, closed=:left)
hist_ANN = fit(Histogram, data[:ANN_mse_class=>0][:E], bins, closed=:left)
hist_all = plot(size=(15 * 39.37, 8 * 39.37))
plot!(hist_before, line=0, fill=(:grey), label="AC + Mu + LAr")
plot!(hist_after_90, line=0, fill=(:blue), label="Presented approach")
plot!(x_axis, hist_ANN.weights, line=(2, :steppost, :lightblue), label="Current implementation")
yaxis!("Events", :log10, (0.5, maximum(hist_before.weights)))
xaxis!("Energy ($(Int64(round(bins[2]-bins[1]))) keV per bin)")
title!("Physics spectrum (semi-coaxial)")
savefig(resolvepath(env, "plots", "Phy-spectrum-coax.pdf"))
savefig(resolvepath(env, "plots", "Phy-spectrum-coax.png"))
hist_all
# +
bins = linspace(500, 1800, 41)
x_axis = bins[1:end-1]
hist_before = fit(Histogram, data[:E], bins, closed=:left)
hist_after_90 = fit(Histogram, after_cut_90[:E], bins, closed=:left)
hist_ANN = fit(Histogram, data[:ANN_mse_class=>0][:E], bins, closed=:left)
hist_surv_90 = 1 - hist_after_90.weights ./ hist_before.weights
hist_surv_ANN = 1 - hist_ANN.weights ./ hist_before.weights
fig_fraction = plot(size=(600, 300))
plot!(x_axis, hist_surv_ANN, line=(:steppost, :lightblue), label="Current implementation")
plot!(x_axis, hist_surv_90, line=(:steppost, :blue), label="Presented approach")
xaxis!("Energy (keV)")
yaxis!("Rejected fraction", (0,1))
title!(L"Rej. on $2\nu\beta\beta$", legend=:none)
savefig(resolvepath(env, "plots", "Rejection-2vbb-spectrum-coax.pdf"))
savefig(resolvepath(env, "plots", "Rejection-2vbb-spectrum-coax.png"))
fig_fraction
# -
fig = plot(hist_all, fig_fraction, layout=@layout([a b{0.25w}]), size=(15 * 39.37, 7 * 39.37))
savefig("plots/Physics Spectrum coax.pdf")
fig
# # Calculate efficiencies
N_2vbb = eventcount(filter(data, :E, E -> (E>=1000)&&(E<=1300)))
N_PNN = eventcount(filter(after_cut_90, :E, E -> (E>=1000)&&(E<=1300)))
N_AoE = eventcount(filter(data[:ANN_mse_class=>0], :E, E -> (E>=1000)&&(E<=1300)))
println("PNN: $(100*N_PNN/N_2vbb) ($N_PNN)")
println("ANN: $(100*N_AoE/N_2vbb) ($N_AoE)")
| examples/Physics Spectrum - coax.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Import Required Libraries
import string
import re
from numpy import array, argmax, random, take
import pandas as pd
from keras.models import Sequential
from keras.layers import Dense, LSTM, Embedding, Bidirectional, RepeatVector, TimeDistributed
from keras.preprocessing.text import Tokenizer
from keras.callbacks import ModelCheckpoint
from keras.preprocessing.sequence import pad_sequences
from keras.models import load_model
from keras import optimizers
import matplotlib.pyplot as plt
% matplotlib inline
pd.set_option('display.max_colwidth', 200)
# ### Read Data
# Our data is a text file of English-German sentence pairs. First we will read the file using the function defined below.
# function to read raw text file
def read_text(filename):
# open the file
file = open(filename, mode='rt', encoding='utf-8')
# read all text
text = file.read()
file.close()
return text
# Now let's define a function to split the text into English-German pairs separated by '\n' and then split these pairs into English sentences and German sentences.
# split a text into sentences
def to_lines(text):
sents = text.strip().split('\n')
sents = [i.split('\t') for i in sents]
return sents
# __Download the data from [here.](http://www.manythings.org/anki/deu-eng.zip)__ and extract "deu.txt" in your working directory.
data = read_text("deu.txt")
deu_eng = to_lines(data)
deu_eng = array(deu_eng)
# The actual data contains over 150,000 sentence-pairs. However, we will use the first 50,000 sentence pairs only to reduce the training time of the model. You can change this number as per you system computation power.
deu_eng = deu_eng[:50000,:]
# ### Text Pre-Processing
# #### Text Cleaning
#
# Let's take a look at our data, then we will decide which pre-processing steps to adopt.
deu_eng
# We will get rid of the punctuation marks, and then convert the text to lower case.
# Remove punctuation
deu_eng[:,0] = [s.translate(str.maketrans('', '', string.punctuation)) for s in deu_eng[:,0]]
deu_eng[:,1] = [s.translate(str.maketrans('', '', string.punctuation)) for s in deu_eng[:,1]]
deu_eng
# convert to lowercase
for i in range(len(deu_eng)):
deu_eng[i,0] = deu_eng[i,0].lower()
deu_eng[i,1] = deu_eng[i,1].lower()
deu_eng
# #### Text to Sequence Conversion
#
# To feed our data in a Seq2Seq model, we will have to convert both the input and the output sentences into integer sequences of fixed length. Before that, let's visualise the length of the sentences. We will capture the lengths of all the sentences in two separate lists for English and German, respectively.
# +
# empty lists
eng_l = []
deu_l = []
# populate the lists with sentence lengths
for i in deu_eng[:,0]:
eng_l.append(len(i.split()))
for i in deu_eng[:,1]:
deu_l.append(len(i.split()))
# -
length_df = pd.DataFrame({'eng':eng_l, 'deu':deu_l})
length_df.hist(bins = 30)
plt.show()
# The maximum length of the German sentences is 11 and that of the English phrases is 8.
# Let's vectorize our text data by using Keras's Tokenizer() class. It will turn our sentences into sequences of integers. Then we will pad those sequences with zeros to make all the sequences of same length.
# function to build a tokenizer
def tokenization(lines):
tokenizer = Tokenizer()
tokenizer.fit_on_texts(lines)
return tokenizer
# +
# prepare english tokenizer
eng_tokenizer = tokenization(deu_eng[:, 0])
eng_vocab_size = len(eng_tokenizer.word_index) + 1
eng_length = 8
print('English Vocabulary Size: %d' % eng_vocab_size)
# +
# prepare Deutch tokenizer
deu_tokenizer = tokenization(deu_eng[:, 1])
deu_vocab_size = len(deu_tokenizer.word_index) + 1
deu_length = 8
print('Deutch Vocabulary Size: %d' % deu_vocab_size)
# -
# Given below is a function to prepare the sequences. It will also perform sequence padding to a maximum sentence length as mentioned above.
# encode and pad sequences
def encode_sequences(tokenizer, length, lines):
# integer encode sequences
seq = tokenizer.texts_to_sequences(lines)
# pad sequences with 0 values
seq = pad_sequences(seq, maxlen=length, padding='post')
return seq
# ### Model Building
# We will now split the data into train and test set for model training and evaluation, respectively.
from sklearn.model_selection import train_test_split
train, test = train_test_split(deu_eng, test_size=0.2, random_state = 12)
# It's time to encode the sentences. We will encode German sentences as the input sequences and English sentences as the target sequences. It will be done for both train and test datasets.
# prepare training data
trainX = encode_sequences(deu_tokenizer, deu_length, train[:, 1])
trainY = encode_sequences(eng_tokenizer, eng_length, train[:, 0])
# prepare validation data
testX = encode_sequences(deu_tokenizer, deu_length, test[:, 1])
testY = encode_sequences(eng_tokenizer, eng_length, test[:, 0])
# Now comes the exciting part! Let us define our Seq2Seq model architecture. We are using an Embedding layer and an LSTM layer as our encoder and another LSTM layer followed by a Dense layer as the decoder.
# build NMT model
def build_model(in_vocab, out_vocab, in_timesteps, out_timesteps, units):
model = Sequential()
model.add(Embedding(in_vocab, units, input_length=in_timesteps, mask_zero=True))
model.add(LSTM(units))
model.add(RepeatVector(out_timesteps))
model.add(LSTM(units, return_sequences=True))
model.add(Dense(out_vocab, activation='softmax'))
return model
# We are using RMSprop optimizer in this model as it is usually a good choice for recurrent neural networks.
model = build_model(deu_vocab_size, eng_vocab_size, deu_length, eng_length, 512)
rms = optimizers.RMSprop(lr=0.001)
model.compile(optimizer=rms, loss='sparse_categorical_crossentropy')
# Please note that we have used __'sparse_categorical_crossentropy'__ as the loss function because it allows us to use the target sequence as it is instead of one hot encoded format. One hot encoding the target sequences with such a huge vocabulary might consume our system's entire memory.
# It seems we are all set to start training our model. We will train it for 30 epochs and with a batch size of 512. You may change and play these hyperparameters. We will also be using __ModelCheckpoint()__ to save the best model with lowest validation loss. I personally prefer this method over early stopping.
# +
filename = 'model.h1.24_jan_19'
checkpoint = ModelCheckpoint(filename, monitor='val_loss', verbose=1, save_best_only=True, mode='min')
history = model.fit(trainX, trainY.reshape(trainY.shape[0], trainY.shape[1], 1),
epochs=30, batch_size=512,
validation_split = 0.2,
callbacks=[checkpoint], verbose=1)
# -
# Let's compare the training loss and the validation loss.
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.legend(['train','validation'])
plt.show()
# ### Make Predictions
# Let's load the saved model to make predictions.
model = load_model('model.h1.24_jan_19')
preds = model.predict_classes(testX.reshape((testX.shape[0],testX.shape[1])))
def get_word(n, tokenizer):
for word, index in tokenizer.word_index.items():
if index == n:
return word
return None
# convert predictions into text (English)
preds_text = []
for i in preds:
temp = []
for j in range(len(i)):
t = get_word(i[j], eng_tokenizer)
if j > 0:
if (t == get_word(i[j-1], eng_tokenizer)) or (t == None):
temp.append('')
else:
temp.append(t)
else:
if(t == None):
temp.append('')
else:
temp.append(t)
preds_text.append(' '.join(temp))
pred_df = pd.DataFrame({'actual' : test[:,0], 'predicted' : preds_text})
pd.set_option('display.max_colwidth', 200)
pred_df.head(15)
pred_df.tail(15)
pred_df.tail(15)
pred_df.sample(15)
| german_to_english.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Intro
# This notebook will go over variable importance metrics for uplift models. It will go over:
#
# - Data Generating Process
# - Model Building
# - Variable Importance Metric
#
# For an introduction on uplift models please see [example on single responses](https://github.com/Ibotta/ibotta_uplift/blob/master/examples/ibotta_uplift_multiple_response_example.ipynb)
#
#
# ### Data Generating Process
#
# Below is the data generating process of the data we have.
#
# \begin{equation}
# x_1 \sim runif(0,1)
# \end{equation}
#
# \begin{equation}
# x_2 \sim runif(0,1)
# \end{equation}
#
# \begin{equation}
# x_3 \sim runif(0,1)
# \end{equation}
#
#
# \begin{equation}
# e_1 \sim rnorm(0,1)
# \end{equation}
#
# \begin{equation}
# e_2 \sim rnorm(0,1)
# \end{equation}
#
# \begin{equation}
# t \sim rbinom(.5)
# \end{equation}
#
# \begin{equation}
# noise \sim rnorm(0,1)
# \end{equation}
#
# \begin{equation}
# revenue = x_1*t + e_1
# \end{equation}
#
# \begin{equation}
# costs = x_2*t + e_2
# \end{equation}
#
# \begin{equation}
# profit = revenue - costs
# \end{equation}
#
#
#
# +
import numpy as np
import pandas as pd
from mr_uplift.dataset.data_simulation import get_simple_uplift_data
from mr_uplift.mr_uplift import MRUplift
from ggplot import *
num_obs = 10000
y, x, t = get_simple_uplift_data(num_obs)
y = pd.DataFrame(y)
y.columns = ['revenue','cost', 'noise']
y['profit'] = y['revenue'] - y['cost']
#include noise explanatory variable
x = pd.DataFrame(x)
x.columns = ['x_1', 'x_2']
x['x_3'] = np.random.normal(0, 1, num_obs)
# +
#build model
uplift_model = MRUplift()
param_grid = dict(num_nodes=[8], dropout=[.1,.5], activation=[
'relu'], num_layers=[1,2], epochs=[25], batch_size=[30])
uplift_model.fit(x, y, t.reshape(-1,1), param_grid = param_grid, n_jobs = 1)
# -
# ### Variable Importance Uplift
#
# The variable importance metrice described here is a variation on permutation importance; shuffle a column and measure how much the output disagrees with output of the original data.
#
# Continuing the notation from the [multiple response example](https://github.com/Ibotta/ibotta_uplift/blob/master/examples/ibotta_uplift_multiple_response_example.ipynb) we have policy assignment $\pi(x_i, W)$ as a function of weights $W$, explanatory variables $X$, and estimated model $E[]$.
#
#
# \begin{equation}
# \pi(x_i, W) =argmax \:_{t \in T} \sum_j w_j *E[y_{j,i} | X=x_i, T=t]
# \end{equation}
#
#
# To obtain a variable importance for a particular explanatory variable $p$ a permutation is performed on that column and a new dataset for each user is generated $x_{i,permuted_p}$. The variable importance is a disagreement between original decision for all $n$ observations :
#
#
# \begin{equation}
# variableimportance_p = 1 - 1/n \sum_{i=1}^{n} I(\pi(x_i, W) = \pi(x_{i,permuted_p}, W))
# \end{equation}
#
# Intuitively, if the decisions of the permuted data is the same as the unpermuted data then we can conclude it is not an important variable. Alternatively, if the decisions are very different then we can conclude that variable is very important.
#
# Below is the variable importance from the fitted model. Note that the noise variable $x_3$ has very low importance relative to the other two. This makes sense since $x_3$ does not effect the response variables.
#
#
uplift_model.permutation_varimp(weights = np.array([.6,-.4,0,0]).reshape(1,-1))
| examples/mr_uplift_variable_importance_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="wWvi-NkazeQC"
# # 必要なライブラリを読み込む
# + colab={} colab_type="code" id="6sp_8vI9vKKZ"
from qore_sdk.client import WebQoreClient
from qore_sdk.featurizer import Featurizer
import qore_sdk.utils
from sklearn import model_selection
from sklearn.metrics import accuracy_score, f1_score
from sklearn.neural_network import MLPClassifier
from sklearn.linear_model import LogisticRegression
import time
import numpy as np
import os
import matplotlib.pyplot as plt
# + [markdown] colab_type="text" id="Y4i401tUtUBH"
# # データの読み込み
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="F6GQoYD-zsG6" outputId="79e7c379-34de-4c19-f8d2-85fccd7e8094"
def load_xyz(str_dir):
with open(os.path.join(str_dir, 'x.txt'), 'r') as f:
x = np.loadtxt(f, delimiter=',', usecols=1)
with open(os.path.join(str_dir, 'y.txt'), 'r') as f:
y = np.loadtxt(f, delimiter=',', usecols=1)
with open(os.path.join(str_dir, 'z.txt'), 'r') as f:
z = np.loadtxt(f, delimiter=',', usecols=1)
return np.stack([x, y, z], 1) # 2D-array
# +
list_data = [
'./data/control',
'./data/udetate',
'./data/hukkin',
'./data/squat',
'./data/roller']
list_X = []
list_y = []
j_label = 0 # incremental label for each data
for i_data in list_data:
print('loading: ' + i_data)
array_loaded = load_xyz(i_data)
array_label = np.repeat(j_label, array_loaded.shape[0])
j_label += 1
plt.figure()
plt.plot(array_loaded)
# 時系列を複数の小時系列に分割する。
# https://qcore-info.github.io/advent-calendar-2019/index.html#qore_sdk.utils.sliding_window
print(array_loaded.shape)
X, y = qore_sdk.utils.sliding_window(array_loaded, width=100, stepsize=1, axis=0, y=array_label, y_def='mode')
print(X.shape, y.shape)
list_X.append(X)
list_y.append(y)
X_all = np.concatenate(list_X, 0)
y_all = np.concatenate(list_y, 0)
# -
# # n_samples_per_classでクラス当たりのサンプル数を揃える。
#
# https://qcore-info.github.io/advent-calendar-2019/index.html#qore_sdk.utils.under_sample
# +
_, counts = np.unique(y_all, return_counts=True)
print(counts)
# サンプル数が一番少ないデータの数に合わせる
X, y = qore_sdk.utils.under_sample(X_all, y_all.flatten(), n_samples_per_class=counts.min())
_, counts = np.unique(y, return_counts=True)
print(counts)
# -
# # QoreSDKのFeaturizerを使って特徴抽出
n_filters = 40
featurizer = Featurizer(n_filters)
X = featurizer.featurize(X, axis=2)
print('X.shape:', X.shape)
# # 学習データとテストデータに分割
# +
X_train, X_test, y_train, y_test = model_selection.train_test_split(
X, y, test_size=0.2, random_state=1
)
print(X_train.shape)
print(y_train.shape)
print(X_test.shape)
print(y_test.shape)
# -
# # アカウント情報の入力
#
# 実際のアカウント情報を載せることはできないので、インタラクティブに入力するようにした
#
# 事前に発行されたユーザーネーム、パスワード、Endpointが必要
# 詳しくは[Advent Calenderの公式Github](https://github.com/qcore-info/advent-calendar-2019)を参照
# +
import getpass
username = getpass.getpass(prompt='username: ', stream=None)
password = getpass.getpass(prompt='password: ', stream=None)
endpoint = getpass.getpass(prompt='endpoint: ', stream=None)
# + colab={} colab_type="code" id="DHXYqKqc05Zx"
# authentication
client = WebQoreClient(username=username,
password=password,
endpoint=endpoint)
# + [markdown] colab_type="text" id="gEJUGAqm8c0M"
# 学習を行う
# +
# サンプル数が多すぎると 505 bad gateway エラーになるので、必要であれば、データ数を減らす
# qoresdk の制限: N*T*V < 150,000 && N*T < 10,000
# 制限は、厳密ではないようであるが、これぐらいに抑えないとエラーになる
n_samples_per_class = 200
X_train, y_train = qore_sdk.utils.under_sample(X_train, y_train.flatten(), n_samples_per_class=n_samples_per_class)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="jWbZJtH32ADt" outputId="2250c28f-46a9-4eb7-aedd-8891af0616a0"
start = time.time()
res = client.classifier_train(X=X_train, Y=y_train)
print(res)
# + [markdown] colab_type="text" id="EkJFSL7w8fTj"
# `
# classifier_test
# `を用いると、精度が簡単に求められて便利
#
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="DwOZ8ZGd2M--" outputId="40520ca9-7758-4a2e-a1c5-070d78deefdd"
res = client.classifier_test(X=X_test, Y=y_test)
print(res)
# + [markdown] colab_type="text" id="uefLZ3wd8rZd"
# 最後には推論もしてみる
# + colab={"base_uri": "https://localhost:8080/", "height": 105} colab_type="code" id="PH5l5IPF2S8w" outputId="e860a970-2928-4094-be20-67ee410ce0ea"
res = client.classifier_predict(X=X_test)
print("acc=", accuracy_score(y_test.tolist(), res["Y"]))
print("f1=", f1_score(y_test.tolist(), res["Y"], average="weighted"))
elapsed_time = time.time() - start
print("elapsed_time:{0}".format(elapsed_time) + "[sec]")
print(res['Y'])
# + [markdown] colab_type="text" id="szVXRg3k3thc"
# # 参考
# 単純な線形回帰、簡単な深層学習と比較する
# + colab={"base_uri": "https://localhost:8080/", "height": 241} colab_type="code" id="rWuixZTz2toL" outputId="1270efce-72cb-45c9-b6a4-0af8be0209ae"
X_train = X_train.reshape(len(X_train), -1).astype(np.float64)
X_test = X_test.reshape(len(X_test), -1).astype(np.float64)
y_train = np.ravel(y_train)
y_test = np.ravel(y_test)
print("===LogisticRegression(Using Sklearn)===")
start = time.time()
lr_cls = LogisticRegression(C=9.0)
lr_cls.fit(X_train, y_train)
elapsed_time = time.time() - start
print("elapsed_time:{0}".format(elapsed_time) + "[sec]")
res = lr_cls.predict(X=X_test)
print("acc=", accuracy_score(y_test.tolist(), res))
print("f1=", f1_score(y_test.tolist(), res, average="weighted"))
print("===MLP(Using Sklearn)===")
start = time.time()
mlp_cls = MLPClassifier(hidden_layer_sizes=(100, 100, 100, 10))
mlp_cls.fit(X_train, y_train)
elapsed_time = time.time() - start
print("elapsed_time:{0}".format(elapsed_time) + "[sec]")
res = mlp_cls.predict(X=X_test)
print("acc=", accuracy_score(y_test.tolist(), res))
print("f1=", f1_score(y_test.tolist(), res, average="weighted"))
# -
# # 新規取得テストデータで検証
# ## 腕立て 12 回の後、少し休憩(ほんとんど動作なし)を行ったデータに対して推論
# +
# 新規取得データの読み込み
list_data = [
'./data/test01_udetate',
]
list_X = []
list_y = []
# j_label = 0 # incremental label for each data
for i_data in list_data:
print('loading: ' + i_data)
array_loaded = load_xyz(i_data)
array_label = np.repeat(j_label, array_loaded.shape[0])
j_label += 1
plt.figure()
plt.plot(array_loaded)
# 時系列を複数の小時系列に分割する。
# https://qcore-info.github.io/advent-calendar-2019/index.html#qore_sdk.utils.sliding_window
print(array_loaded.shape)
# X, y = qore_sdk.utils.sliding_window(array_loaded, width=100, stepsize=1, axis=0, y=array_label, y_def='mode')
X = qore_sdk.utils.sliding_window(array_loaded, width=100, stepsize=1, axis=0)
# print(X.shape, y.shape)
list_X.append(X)
# list_y.append(y)
X_all = np.concatenate(list_X, 0)
# y_all = np.concatenate(list_y, 0)
# +
# 検証データの Featurizer による特徴抽出
print(X_all.shape)
n_filters = 40
featurizer = Featurizer(n_filters)
X = featurizer.featurize(X_all, axis=2)
print('X.shape:', X.shape)
# +
# 推論
res = client.classifier_predict(X=X)
# print("acc=", accuracy_score(y_test.tolist(), res["Y"]))
# print("f1=", f1_score(y_test.tolist(), res["Y"], average="weighted"))
elapsed_time = time.time() - start
print("elapsed_time:{0}".format(elapsed_time) + "[sec]")
print(res['Y'])
# -
# 推論結果の可視化
plt.figure()
plt.plot(res['Y'])
# データラベル
#
# 0 --> 筋トレしていない
# 1 --> 腕立て
# 2 --> 腹筋
# 3 --> スクワット
# 4 --> 腹筋ローラー
#
# データ数 200 あたりで腕立てを終えて、休憩している。
# 腕立てをしている間は、ほぼ正解だが、休憩中は当たっていない。
# これは、学習に利用した、筋トレしていない状態のデータに問題があると思われる。
# 以下、改善案
#
# - 筋トレしていない状態のデータに、もっと様々な状況のデータを追加する
# - 静止、休止状態のクラスを追加する
# ## 新規取得した腹筋ローラーデータに対して推論
# +
# 新規取得データの読み込み
list_data = [
'./data/test02_roller',
]
list_X = []
list_y = []
j_label = 4 # incremental label for each data
for i_data in list_data:
print('loading: ' + i_data)
array_loaded = load_xyz(i_data)
array_label = np.repeat(j_label, array_loaded.shape[0])
# j_label += 1
plt.figure()
plt.plot(array_loaded)
# 時系列を複数の小時系列に分割する。
# https://qcore-info.github.io/advent-calendar-2019/index.html#qore_sdk.utils.sliding_window
print(array_loaded.shape)
X, y = qore_sdk.utils.sliding_window(array_loaded, width=100, stepsize=1, axis=0, y=array_label, y_def='mode')
# X = qore_sdk.utils.sliding_window(array_loaded, width=100, stepsize=1, axis=0)
print(X.shape, y.shape)
list_X.append(X)
list_y.append(y)
X_all = np.concatenate(list_X, 0)
y_all = np.concatenate(list_y, 0)
# +
# 検証データの Featurizer による特徴抽出
print(X_all.shape)
n_filters = 40
featurizer = Featurizer(n_filters)
X = featurizer.featurize(X_all, axis=2)
print('X.shape:', X.shape)
# +
# 推論
res = client.classifier_predict(X=X)
print("acc=", accuracy_score(y_all.tolist(), res["Y"]))
print("f1=", f1_score(y_all.tolist(), res["Y"], average="weighted"))
elapsed_time = time.time() - start
print("elapsed_time:{0}".format(elapsed_time) + "[sec]")
print(res['Y'])
# -
# 推論結果の可視化
plt.figure()
plt.plot(res['Y'])
# +
X_test = X.reshape(len(X), -1).astype(np.float64)
y_test = np.ravel(y_all)
print("===LogisticRegression(Using Sklearn)===")
res = lr_cls.predict(X=X_test)
print("acc=", accuracy_score(y_test.tolist(), res))
print("f1=", f1_score(y_test.tolist(), res, average="weighted"))
print("===MLP(Using Sklearn)===")
res = mlp_cls.predict(X=X_test)
print("acc=", accuracy_score(y_test.tolist(), res))
print("f1=", f1_score(y_test.tolist(), res, average="weighted"))
# -
| muscle_QoreSDK_v2.ipynb |