code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
sys.path.append('../../pyutils')
import numpy as np
from scipy.linalg import solve_triangular
import metrics
# -
# # The Cholesky decomposition
#
# Let $A \in \mathbb{R}^{n*n}$ a symmetric positive definite matrix.
# $A$ can de decomposed as:
# $$A = LL^T$$
# with $L \in \mathbb{R}^{n*n}$ a lower triangular matrix with positive diagonal entries.
#
#
# $$L_{jj} = \sqrt{A_{jj} - \sum_{k=1}^{j-1} L_{jk}^2}$$
# $$L_{ij} = \frac{1}{L_{jj}}(A_{ij} - \sum_{k=1}^{j-1}L_{ik}L_{jk}) \text{ for } i > j$$
#
# The algorithm can be impleted row by row or column by column
# +
def cholesky(A):
n = len(A)
L = np.zeros((n, n))
for j in range(n):
L[j,j] = np.sqrt(A[j,j] - np.sum(L[j,:j]**2))
for i in range(j+1, n):
L[i,j] = (A[i,j] - np.sum(L[i, :j] * L[j, :j])) / L[j,j]
return L
A = np.random.randn(5, 5)
A = A @ A.T
L = cholesky(A)
print(metrics.tdist(L - np.tril(L), np.zeros(A.shape)))
print(metrics.tdist(L @ L.T, A))
# -
# # The LDL Decomposition
#
# Let $A \in \mathbb{R}^{n*n}$ a symmetric positive definite matrix.
# $A$ can de decomposed as:
# $$A = LDL^T$$
# with $L \in \mathbb{R}^{n*n}$ a lower unit triangular matrix and $D \in \mathbb{R}^{n*n}$ a diagonal matrix.
# This is a modified version of the Cholsky decomposition that doesn't need square roots.
#
# $$D_{jj} = A_{jj} - \sum_{k=1}^{j-1} L_{jk}^2D_{kk}$$
# $$L_{ij} = \frac{1}{D_{jj}}(A_{ij} - \sum_{k=1}^{j-1}L_{ik}L_{jk}D_{kk}) \text{ for } i > j$$
# +
def cholesky_ldl(A):
n = len(A)
L = np.eye(n)
d = np.zeros(n)
for j in range(n):
d[j] = A[j,j] - np.sum(L[j,:j]**2 * d[:j])
for i in range(j+1, n):
L[i,j] = (A[i,j] - np.sum(L[i,:j]*L[j,:j]*d[:j])) / d[j]
return L, d
A = np.random.randn(5, 5)
A = A @ A.T
L, d = cholesky_ldl(A)
print(metrics.tdist(L - np.tril(L), np.zeros(A.shape)))
print(metrics.tdist(np.diag(L), np.ones(len(L))))
print(metrics.tdist(L @ np.diag(d) @ L.T, A))
# -
# # Solve a linear system
#
# Find $x$ such that:
#
# $$Ax = b$$
#
# Compute the cholesky decomposition of $A$
#
# $$A = LL^T$$
# $$LL^Tx = b$$
#
# Solve the lower triangular system:
#
# $$Ly = b$$
#
# Solve the upper triangular system:
#
# $$L^Tx = y$$
# +
def cholesky_system(A, b):
L = cholesky(A)
y = solve_triangular(L, b, lower=True)
x = solve_triangular(L.T, y, lower=False)
return x
A = np.random.randn(5, 5)
A = A @ A.T
b = np.random.randn(5)
x = cholesky_system(A, b)
print(metrics.tdist(A @ x, b))
| courses/linear_algebra/cholesky_decomposition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# importacion general de librerias y de visualizacion (matplotlib y seaborn)
import pandas as pd
import numpy as np
import random
import re
pd.options.display.float_format = '{:20,.2f}'.format # suprimimos la notacion cientifica en los outputs
import warnings
warnings.filterwarnings('ignore')
# -
twt_data = pd.read_csv('~/Documents/Datos/DataSets/TP2/train_super_featured.csv')
twt_data.head()
pattern = re.compile("(?P<url>https?://[^\s]+)")
def remove_link(twt):
return pattern.sub("r ", twt)
# + active=""
# twt_data['text'] = twt_data['text'].map(remove_link)
# -
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.naive_bayes import MultinomialNB
from sklearn.pipeline import Pipeline
from sklearn.linear_model import SGDClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import TfidfVectorizer
train, test = train_test_split(twt_data, test_size=0.2)
count_vect = CountVectorizer(stop_words='english')
X_train_counts = count_vect.fit_transform(twt_data['super_clean_text'])
X_train_counts.shape
tfidf_transformer = TfidfTransformer()
X_train_tfidf = tfidf_transformer.fit_transform(X_train_counts)
X_train_tfidf.shape
count_vect.get_feature_names()
text_clf = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', MultinomialNB()),
])
text_clf = text_clf.fit(train.super_clean_text, train.target_relabeled)
predicted = text_clf.predict(test.super_clean_text)
np.mean(predicted == test.target_relabeled)
text_clf_svm = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf-svm', SGDClassifier(loss='hinge', penalty='l2',
alpha=1e-3, random_state=42)),
])
text_clf_svm = text_clf_svm.fit(train.super_clean_text, train.target_relabeled)
predicted_svm = text_clf_svm.predict(test.super_clean_text)
np.mean(predicted_svm == test.target_relabeled)
parameters = {'vect__ngram_range': [(1, 1), (1, 2)],
'tfidf__use_idf': (True, False),
'clf__alpha': (1e-2, 1e-3),
}
gs_clf = GridSearchCV(text_clf, parameters, n_jobs=-1)
gs_clf = gs_clf.fit(train.clean_text, train.target_label)
gs_clf.best_score_
gs_clf.best_params_
text_clf_improved = Pipeline([('vect', CountVectorizer()),
('clf', MultinomialNB()),
])
text_clf = text_clf_improved.fit(train.clean_text, train.target_label)
predicted = text_clf_improved.predict(test.clean_text)
np.mean(predicted == test.target_label)
vectorizer = TfidfVectorizer()
X = vectorizer.fit_transform(train.clean_text)
tf_idf_train = pd.DataFrame(data = X.toarray(), columns=vectorizer.get_feature_names())
tf_idf_train
test_data = pd.read_csv('~/Documents/Datos/DataSets/TP2/test_super_featured.csv')
test_data.head()
text_clf = text_clf.fit(twt_data.clean_text, twt_data.target_label)
predicted = text_clf.predict(test_data.clean_text)
predicted
test_data[test_data.clean_text != test_data.clean_text]
test_data['text_super_cleaned'].fillna(" ", inplace=True)
test_data[test_data.clean_text != test_data.clean_text]
test_data['target'] = predicted
test_data[['id_original', 'target']].rename(columns={'id_original': 'id'}).to_csv('~/Documents/Datos/DataSets/TP2/res_NB_1.csv', index=False)
| TP2/Alejo/1_NB_SUPERCLEAN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
os.chdir("..")
import pandas as pd
import scipy.stats as stats
# -
bio_merge_sample_df=\
pd.read_csv("constructed\\capstone\\bio_merge_sample.csv", sep=',')
bio_merge_sample_df=bio_merge_sample_df.loc[:, ~bio_merge_sample_df.columns.str.contains('^Unnamed')]
bio_merge_sample_df=bio_merge_sample_df.reset_index(drop=True)
bio_merge_sample_df.head(10)
bio_merge_sample_df.shape
#t-test to compare abnormal returns on sec filing days: Average return on days with SEC filings
#has more variance
bio_merge_sample_df['abs_abnormal_return']=abs(bio_merge_sample_df['arith_resid'])
returns_nosec=bio_merge_sample_df.loc[bio_merge_sample_df.path.isnull()]['abs_abnormal_return']
returns_sec=bio_merge_sample_df[bio_merge_sample_df.path.notnull()]['abs_abnormal_return']
t_stat, p_val = stats.ttest_ind(returns_nosec, returns_sec, equal_var=False)
t_stat
p_val
returns_nosec.describe()
returns_sec.describe()
# +
#compare Psychosocial Words and TFID-selected words for biotech sample
words_df= pd.read_csv("input files\\capstone\\capstone_sentiment.csv")
words_df=words_df.loc[:, ~words_df.columns.str.contains('^Unnamed')]
words_df=words_df.reset_index(drop=True)
words_df['word']=words_df['word'].str.lower()
#words_df.head(10)
bio_dataset_features= pd.read_csv("constructed\\bio_dataset_features.csv")
bio_dataset_features=bio_dataset_features.loc[:, ~bio_dataset_features.columns.str.contains('^Unnamed')]
bio_dataset_features=bio_dataset_features.reset_index(drop=True)
#bio_dataset_features.head(10)
comparison_df=pd.merge(words_df, bio_dataset_features, how='inner', \
on=['word'])
comparison_df.head(20)
# +
#compare Psychosocial Words and TFID-selected words for large sample:
all_features_features= pd.read_csv("constructed\\Large_dataset_features.csv")
all_features_features=all_features_features.loc[:, ~all_features_features.columns.str.contains('^Unnamed')]
all_features_features=all_features_features.reset_index(drop=True)
all_comparison_df=pd.merge(words_df, all_features_features, how='inner', \
on=['word'])
all_comparison_df.head(20)
# -
| python/capstone/Capstone Results Additional.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Study Designer Example (DNA sequencing)
from ipywidgets import (RadioButtons, VBox, HBox, Layout, Label, Checkbox, Text, IntSlider)
from qgrid import show_grid
label_layout = Layout(width='100%')
from isatools.create.models import *
from isatools.model import Investigation
from isatools.isatab import dump_tables_to_dataframes as dumpdf
import qgrid
qgrid.nbinstall(overwrite=True)
# ## Sample planning section
# ### Study design type
#
# Please specify if the study is an intervention or an observation.
rad_study_design = RadioButtons(options=['Intervention', 'Observation'], value='Intervention', disabled=False)
VBox([Label('Study design type?', layout=label_layout), rad_study_design])
# ### Intervention study
#
# If specifying an intervention study, please answer the following:
# - Are study subjects exposed to a single intervention or to multiple intervention?
# - Are there 'hard to change' factors, which restrict randomization of experimental unit?
#
# *Note: if you chose 'observation' as the study design type, the following choices will be disabled and you should skip to the Observation study section*
#
if rad_study_design.value == 'Intervention':
study_design = InterventionStudyDesign()
if rad_study_design.value == 'Observation':
study_design = None
intervention_ui_disabled = not isinstance(study_design, InterventionStudyDesign)
intervention_type = RadioButtons(options=['single', 'multiple'], value='single', disabled=intervention_ui_disabled)
intervention_type_vbox = VBox([Label('Single intervention or to multiple intervention?', layout=label_layout), intervention_type])
free_or_restricted_design = RadioButtons(options=['yes', 'no'], value='no', disabled=intervention_ui_disabled)
free_or_restricted_design_vbox = VBox([Label("Are there 'hard to change' factors?", layout=label_layout), free_or_restricted_design])
HBox([intervention_type_vbox, free_or_restricted_design_vbox])
hard_to_change_factors_ui_disabled = free_or_restricted_design.value == 'no'
hard_to_change_factors = RadioButtons(options=[1, 2], value=1, disabled=hard_to_change_factors_ui_disabled)
VBox([Label("If applicable, how many 'hard to change factors'?", layout=label_layout), hard_to_change_factors])
repeats = intervention_type.value != 'single'
factorial_design = free_or_restricted_design.value == 'no'
split_plot_design = (free_or_restricted_design.value == 'yes' and hard_to_change_factors.value == 1)
split_split_plot_design = (free_or_restricted_design.value == 'yes' and hard_to_change_factors.value == 2)
print('Interventions: {}'.format('Multiple interventions' if repeats else 'Single intervention'))
design_type = 'factorial' # always default to factorial
if split_plot_design:
design_type = 'split plot'
elif split_split_plot_design:
design_type = 'split split'
print('Design type: {}'.format(design_type))
# #### Factorial design - intervention types
#
# If specifying an factorial design, please list the intervention types here.
factorial_design_ui_disabled = not factorial_design
chemical_intervention = Checkbox(value=True, description='Chemical intervention', disabled=factorial_design_ui_disabled)
behavioural_intervention = Checkbox(value=False, description='Behavioural intervention', disabled=factorial_design_ui_disabled)
surgical_intervention = Checkbox(value=False, description='Surgical intervention', disabled=factorial_design_ui_disabled)
biological_intervention = Checkbox(value=False, description='Biological intervention', disabled=factorial_design_ui_disabled)
radiological_intervention = Checkbox(value=False, description='Radiological intervention', disabled=factorial_design_ui_disabled)
VBox([chemical_intervention, surgical_intervention, biological_intervention, radiological_intervention])
level_uis = []
if chemical_intervention:
agent_levels = Text(
value='calpol, no agent',
placeholder='e.g. cocaine, calpol',
description='Agent:',
disabled=False
)
dose_levels = Text(
value='low, high',
placeholder='e.g. low, high',
description='Dose levels:',
disabled=False
)
duration_of_exposure_levels = Text(
value='short, long',
placeholder='e.g. short, long',
description='Duration of exposure:',
disabled=False
)
VBox([Label("Chemical intervention factor levels:", layout=label_layout), agent_levels, dose_levels, duration_of_exposure_levels])
factory = TreatmentFactory(intervention_type=INTERVENTIONS['CHEMICAL'], factors=BASE_FACTORS)
for agent_level in agent_levels.value.split(','):
factory.add_factor_value(BASE_FACTORS[0], agent_level.strip())
for dose_level in dose_levels.value.split(','):
factory.add_factor_value(BASE_FACTORS[1], dose_level.strip())
for duration_of_exposure_level in duration_of_exposure_levels.value.split(','):
factory.add_factor_value(BASE_FACTORS[2], duration_of_exposure_level.strip())
print('Number of study groups (treatment groups): {}'.format(len(factory.compute_full_factorial_design())))
treatment_sequence = TreatmentSequence(ranked_treatments=factory.compute_full_factorial_design())
# Next, specify if all study groups of the same size, i.e have the same number of subjects? (in other words, are the groups balanced).
group_blanced = RadioButtons(options=['Balanced', 'Unbalanced'], value='Balanced', disabled=False)
VBox([Label('Are study groups balanced?', layout=label_layout), group_blanced])
# Provide the number of subject per study group:
group_size = IntSlider(value=5, min=0, max=100, step=1, description='Group size:', disabled=False, continuous_update=False, orientation='horizontal', readout=True, readout_format='d')
group_size
plan = SampleAssayPlan(group_size=group_size.value)
rad_sample_type = RadioButtons(options=['Blood', 'Sweat', 'Tears', 'Urine'], value='Blood', disabled=False)
VBox([Label('Sample type?', layout=label_layout), rad_sample_type])
# How many times each of the samples have been collected?
sampling_size = IntSlider(value=3, min=0, max=100, step=1, description='Sample size:', disabled=False, continuous_update=False, orientation='horizontal', readout=True, readout_format='d')
sampling_size
plan.add_sample_type(rad_sample_type.value)
plan.add_sample_plan_record(rad_sample_type.value, sampling_size.value)
isa_object_factory = IsaModelObjectFactory(plan, treatment_sequence)
# ## Generate ISA model objects from the sample plan and render the study-sample table
# *Check state of the Sample Assay Plan after entering sample planning information:*
import json
from isatools.create.models import SampleAssayPlanEncoder
print(json.dumps(plan, cls=SampleAssayPlanEncoder, sort_keys=True, indent=4, separators=(',', ': ')))
isa_investigation = Investigation(identifier='inv101')
isa_study = isa_object_factory.create_study_from_plan()
isa_study.filename = 's_study.txt'
isa_investigation.studies = [isa_study]
dataframes = dumpdf(isa_investigation)
sample_table = next(iter(dataframes.values()))
show_grid(sample_table)
print('Total rows generated: {}'.format(len(sample_table)))
# ## Assay planning
# ### Select assay technology type to map to sample type from sample plan
rad_assay_type = RadioButtons(options=['DNA microarray', 'DNA sequencing', 'Mass spectrometry', 'NMR spectroscopy'], value='DNA sequencing', disabled=False)
VBox([Label('Assay type to map to sample type "{}"?'.format(rad_sample_type.value), layout=label_layout), rad_assay_type])
if rad_assay_type.value == 'DNA sequencing':
assay_type = AssayType(measurement_type='genome sequencing', technology_type='nucleotide sequencing')
print('Selected measurement type "genome sequencing" and technology type "nucleotide sequencing"')
else:
raise Exception('Assay type not implemented')
# ### Topology modifications
technical_replicates = IntSlider(value=2, min=0, max=5, step=1, description='Technical repeats:', disabled=False, continuous_update=False, orientation='horizontal', readout=True, readout_format='d')
technical_replicates
inst_mod_454gs = Checkbox(value=True, description='Instrument: 454 GS')
inst_mod_454gsflx = Checkbox(value=True, description='Instrument: 454 GS FLX')
VBox([inst_mod_454gs, inst_mod_454gsflx])
instruments = set()
if inst_mod_454gs.value: instruments.add('454 GS')
if inst_mod_454gsflx.value: instruments.add('454 GS FLX')
top_mods = DNASeqAssayTopologyModifiers(technical_replicates=technical_replicates.value, instruments=instruments)
print('Technical replicates: {}'.format(top_mods.technical_replicates))
assay_type.topology_modifiers = top_mods
plan.add_assay_type(assay_type)
plan.add_assay_plan_record(rad_sample_type.value, assay_type)
assay_plan = next(iter(plan.assay_plan))
print('Added assay plan: {0} -> {1}/{2}'.format(assay_plan[0].value.term, assay_plan[1].measurement_type.term, assay_plan[1].technology_type.term))
if len(top_mods.instruments) > 0:
print('Instruments: {}'.format(list(top_mods.instruments)))
# ## Generate ISA model objects from the assay plan and render the assay table
# *Check state of Sample Assay Plan after entering assay plan information:*
print(json.dumps(plan, cls=SampleAssayPlanEncoder, sort_keys=True, indent=4, separators=(',', ': ')))
isa_investigation.studies = [isa_object_factory.create_assays_from_plan()]
for assay in isa_investigation.studies[-1].assays:
print('Assay generated: {0}, {1} samples, {2} processes, {3} data files'
.format(assay.filename, len(assay.samples), len(assay.process_sequence), len(assay.data_files)))
dataframes = dumpdf(isa_investigation)
show_grid(dataframes[next(iter(dataframes.keys()))])
| notebooks/Study Designer (DNA sequencing).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Predicing Housing Prices Using Linear Regression
#
#
# In our last exercises, we learned about Python, Matrices, Vectors, handing and plotting data. In this tutorial, we will use all that information and create a mathematical model to predict housing prices using Linear regression.
#
#
# #### What is Linear Regression
#
# Linear regression attempts to model the relationship between two variables by fitting a linear equation to observed data. One variable is considered to be an explanatory variable, and the other is considered to be a dependent variable.
#
# <img src="1.PNG" align="center"/>
#
# **Least Squares Methods**
#
# <img src="2.PNG" align="center"/>
from IPython.display import YouTubeVideo
YouTubeVideo('OxMNNjp-mDw', width=860, height=460)
# ### Predicting Housing prices using Linear Regression
#
# We will take the Housing dataset which contains information about different houses in Boston. We can access this data from the scikit-learn library. There are 506 samples and 13 feature variables in this dataset. The objective is to predict the value of prices of the house using the given features.
#
# So let’s get started.
# +
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
from sklearn.datasets import load_boston
boston_dataset = load_boston()
# -
print(boston_dataset.keys())
print (boston_dataset.DESCR)
boston = pd.DataFrame(boston_dataset.data, columns=boston_dataset.feature_names)
boston.head()
#- Median value of owner-occupied homes in $1000's
boston['MEDV'] = boston_dataset.target
sns.distplot(boston['MEDV'], bins=30)
plt.show()
# We will use the following features to train our Model.
#
# **RM:** It's an average number of rooms, so I would expect this to never be less than 1, while the maximum number of rooms could be quite high. Given the maximum price of $50,000, I would expect it to be either 1.385 or 5.609.
#
# **CRIM:** Per capita crime rate by town
#
# In machine learning, the variable that is being modeled is called the target variable; it's what you are trying to predict given the features. For this dataset, the suggested target is MEDV, the median house value in 1,000s of dollar
X = pd.DataFrame(np.c_[boston['CRIM'], boston['RM']], columns = ['CRIM','RM'])
Y = boston['MEDV']
# +
from sklearn.model_selection import train_test_split
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.2, random_state=5)
print(X_train.shape)
print(X_test.shape)
print(Y_train.shape)
print(Y_test.shape)
# +
from sklearn.linear_model import LinearRegression
lin_model = LinearRegression()
lin_model.fit(X_train, Y_train)
# -
Y_predict = lin_model.predict(X_test)
print (X_test.shape)
print (Y_predict)
# Scatter plots of Actual vs Predicted are one of the richest form of data visualization. You can tell pretty much everything from it. Ideally, all your points should be close to a regressed diagonal line.
# +
plt.scatter(Y_test, Y_predict , c = 'red')
# 100 percent accuracy line
plt.plot(Y_test, Y_test, c = 'blue')
plt.xlabel("Actual House Prices ($1000)")
plt.ylabel("Predicted House Prices: ($1000)")
plt.xticks(range(0, int(max(Y_test)),2))
plt.yticks(range(0, int(max(Y_predict)),2))
plt.title("Actual Prices vs Predicted prices")
| Implementation_Examples_Predicting_Something/Implementation_Examples_Predicting_Something.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas
data = pandas.read_csv('data/gapminder_gdp_europe.csv', index_col='country')
#Select value by position in tabel
#Select value in 1st row and 1st column
print(data.iloc[0, 0])
# -
#Select value by entry label, similar to using dictionary keys
print(data.loc["Albania", "gdpPercap_1952"])
#Using : to select all columns/rows
#Like slicing
#Would produce the same result without a 2nd index ( data.loc["Albania] )
print(data.loc["Albania", :])
# Would get the same result from
# data["gdpPercap_1952"]
# and data.gdpPercap_1952
print(data.loc[:, "gdpPercap_1952"])
#Select multiple columns/rows using a named slice
# loc is inclusive at both ends of the slice
# iloc includes everything but the final index
print(data.loc['Italy':'Poland', 'gdpPercap_1962':'gdpPercap_1972'])
#Calculate the max value in a slice
print(data.loc['Italy':'Poland', 'gdpPercap_1962':'gdpPercap_1972'].max())
#Calculate the min value in a slice
print(data.loc['Italy':'Poland', 'gdpPercap_1962':'gdpPercap_1972'].min())
#Use comparisons based on values
#Produces a boolean dataframe
subset = data.loc['Italy':'Poland', 'gdpPercap_1962':'gdpPercap_1972']
print('Subset of data:\n', subset)
# Which values were greater than 10000 ?
print('\nWhere are values large?\n', subset > 10000)
#Mask where false using NaN (Not a number)
mask = subset > 10000
print(subset[mask])
#NaN is ignored by stats operations
print(subset[subset > 10000].describe())
# # Select-Apply-Combine methods
#Split European contries by GDP
#Split into higher and lower than average GDP
#Estimate 'Wealthy score' by how many times a country has been higher or lower
mask_higher = data.apply(lambda x:x>x.mean())
wealth_score = mask_higher.aggregate('sum',axis=1)/len(data.columns)
wealth_score
#Sum financial contribution over the years
data.groupby(wealth_score).sum()
#Find per capita of Serbia in 2007
print(data.loc["Serbia", "gdpPercap_2007"])
#iloc includes all but last index: 0, 1
print(data.iloc[0:2, 0:2])
#loc includes all: 0, 1, 2
print(data.loc['Albania':'Belgium', 'gdpPercap_1952':'gdpPercap_1962'])
#first = Reads in data and assigns columns as row headings
#second = Selects data where continent=Americas
#third = From Americas data, remove Puerto Rico
#fourth = Remove the "continent" column - the 2nd column which has an index of 1
#Write to .csv called "result.csv"
first = pandas.read_csv('data/gapminder_all.csv', index_col='country')
second = first[first['continent'] == 'Americas']
third = second.drop('Puerto Rico')
fourth = third.drop('continent', axis = 1)
fourth.to_csv('result.csv')
#Return index value of column minimum
print(data.idxmin())
#Return index value of column maximum
print(data.idxmax())
#Select GDP per capita for all countries 1982
data['gdpPercap_1982']
#Select GDP per capita for Denmark in all years
data.loc['Denmark']
#Select GDP per capita for all countries for after 1985
data.loc[:,'gdpPercap_1985':]
#Select GDP per capita for each country in 2007 as a multiple of GDP
#per capita for that country in 1952
data['gdpPercap_2007']/data['gdpPercap_1952']
| Pandas/Pandas8_Dataframes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visualizing sequential Bayesian learning
#
# In this notebook we will examine the problem of estimation given observed data from a Bayesian perspective.
#
# We start by gathering a dataset $\mathcal{D}$ consisting of multiple observations. Each observation is independent and drawn from a parametric probability distribution with parameter $\mu$. We can thus write the probability of the dataset as $p(\mathcal{D}\mid\mu)$, which determines how likely it is to observe the dataset. For this reason, $p(\mathcal{D}\mid\mu)$ is known as the **likelihood** function.
#
# Furthermore, we believe that the parameter $\mu$ is itself a random variable with a known probability distribution $p(\mu)$ that encodes our prior belief. This distribution is known as the **prior** distribution.
#
# Now we might ask ourselves: at a moment posterior to observing the data $\mathcal{D}$, how does our initial belief on $\mu$ change? We would like to know what the probability $p(\mu\mid\mathcal{D})$ is, which is known as the **posterior** distribution. This probability, according to Bayes' theorem, is given by
#
# $$
# p(\mu\mid\mathcal{D}) = \frac{p(\mathcal{D}\mid\mu)p(\mu)}{p(\mathcal{D})}
# $$
#
# ## Flipping coins
#
# Suppose we have a strange coin for which we are not certain about the probability $\mu$ of getting heads with it. We can start flipping it many times, and as we record heads or tails in a dataset $\mathcal{D}$ we can modify our beliefs on $\mu$ given our observations. Given that we have flipped the coins $N$ times, the likelihood of observing $m$ heads is given by the Binomial distribution,
#
# $$\text{Bin}(m\mid N,\mu)={N\choose k}\mu^m(1-\mu)^{N-m}$$
#
# A mathematically convenient prior distribution for $\mu$ that works well with a binomial likelihood is the Beta distribution,
#
# $$\text{Beta}(\mu\mid a, b) = \frac{\Gamma(a+b)}{\Gamma(a)\Gamma(b)}\mu^{a-1}(1-\mu)^{b-1}$$
#
# We said $\mu$ would encode our *prior* belief on $\mu$, so why does it seem that we are choosing a prior for mathematical convenience instead? It turns out this prior, in addition to being convenient, will also allow us to encode a variety of prior beliefs as we might see fit. We can see this by plotting the distribution for different values of $a$ and $b$:
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import matplotlib.animation as animation
import numpy as np
from scipy.stats import beta
# The set of parameters of the prior to test
a_b_params = ((0.1, 0.1), (1, 1), (2, 3), (8, 4))
mu = np.linspace(0, 1, 100)
# Plot one figure per set of parameters
plt.figure(figsize=(13,3))
for i, (a, b) in enumerate(a_b_params):
plt.subplot(1, len(a_b_params), i+1)
prior = beta(a, b)
plt.plot(mu, prior.pdf(mu))
plt.xlabel(r'$\mu$')
plt.title("a = {:.1f}, b = {:.1f}".format(a, b))
plt.tight_layout()
# -
# We can see that the distribution is rather flexible. Since we don't know anything about the probability of the coin (which is also a valid prior belief), we can select $a=1$ and $b=1$ so that the probability is uniform in the interval $(0, 1)$.
#
# Now that we have the likelihood and prior distributions, we can obtain the posterior distribution $p(\mu\mid\mathcal{D})$: the revised beliefs on $\mu$ after flipping the coin many times and gathering a dataset $\mathcal{D}$. In general, calculating the posterior can be a very involved derivation, however what makes the choice of the prior mathematically convenient is that it makes the posterior distribution to be the same: a Beta distribution. A prior distribution is called a **conjugate prior** for a likelihood function if the resulting posterior has the same form of the prior. For our example, the posterior is
#
# $$p(\mu\mid m, l, a, b) = \frac{\Gamma(m+a+l+b)}{\Gamma(m+a)\Gamma{l+b}}\mu^{m+a-1}(1-\mu)^{l+b-1}$$
#
# where $m$ is the number of heads we observed and $l$ the number of tails (equal to $N-m$).
#
# That's it! Our prior belief is ready and the likelihood of our observations is determined, we can now start flipping coins and seeing how the posterior probability changes with each observation. We will set up a simulation where we flip a coin with probability 0.8 of landing on heads, which is the unknown probability that we aim to find. Via the process of bayesian learning we will discover the true value as we flip the coin one time after the other.
# +
def coin_flip(mu=0.8):
""" Returns True (heads) with probability mu, False (tails) otherwise. """
return np.random.random() < mu
# Parameters for a uniform prior
a = 1
b = 1
posterior = beta(a, b)
# Observed heads and tails
m = 0
l = 0
# A list to store posterior updates
updates = []
for toss in range(50):
# Store posterior
updates.append(posterior.pdf(mu))
# Get a new observation by flipping the coin
if coin_flip():
m += 1
else:
l += 1
# Update posterior
posterior = beta(a + m, b + l)
# -
# We now have a list, `updates`, containing the values of the posterior distribution after observing one coin toss. We can visualize the change in the posterior interactively using <a href="https://plot.ly/#/" target="_blank">Plotly</a>. Even though there are other options to do animations and use interactive widgets with Python, I chose Plotly because it is portable (Matplotlib's `FuncAnimation` requires extra components to be installed) and it looks nice. All we have to do is to define a dictionary according to the specifications, and we obtain an interactive plot that can be embedded in a notebook or a web page.
#
# In the figure below we will add *Play* and *Pause* buttons, as well as a slider to control playback, which will allow us to view the changes in the posterior distributions are new observations are made. Let's take a look at how it's done.
# +
# Set up and plot with Plotly
import plotly.offline as ply
ply.init_notebook_mode(connected=True)
figure = {'data': [{'x': [0, 1], 'y': [0, 1], 'mode': 'lines'}],
'layout':
{
'height': 400, 'width': 600,
'xaxis': {'range': [0, 1], 'autorange': False},
'yaxis': {'range': [0, 8], 'autorange': False},
'title': 'Posterior distribution',
'updatemenus':
[{
'type': 'buttons',
'buttons':
[
{
'label': 'Play',
'method': 'animate',
'args':
[
None,
{
'frame': {'duration': 500, 'redraw': False},
'fromcurrent': True,
'transition': {'duration': 0, 'easing': 'linear'}
}
]
},
{
'args':
[
[None],
{
'frame': {'duration': 0, 'redraw': False},
'mode': 'immediate',
'transition': {'duration': 0}
}
],
'label': 'Pause',
'method': 'animate'
}
]
}],
'sliders': [{
'steps': [{
'args':
[
['frame{}'.format(f)],
{
'frame': {'duration': 0, 'redraw': False},
'mode': 'immediate',
'transition': {'duration': 0}
}
],
'label': f,
'method': 'animate'
} for f, update in enumerate(updates)]
}]
},
'frames': [{'data': [{'x': mu, 'y': pu}], 'name': 'frame{}'.format(f)} for f, pu in enumerate(updates)]}
ply.iplot(figure, link_text='')
# -
# We can see how bayesian learning allows us to go from a uniform prior distribution on the parameter $\mu$, when there are no observations, and as the number of observations increases, our uncertainty is reduced (which can be seen from the reduced variance of the distribution) and the estimated value centers around the true value of 0.8.
#
# In my opinion, this is a very elegant and powerful method for inference that adequately adapts to a sequential setting as the one we have explored. Even more interesting is the fact that this simple idea can be applied to much more complicated models that deal with uncertainty. On the other hand, we have to keep in mind that the posterior was chosen for mathematical convenience, rather than a selection based on the nature of a particular problem, for which in some cases a conjugate prior might not be the best.
| 00-bayesian-learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # BuildSys 2020 Figures
# The point of this notebook is to generate good quality figures for the BuildSys 2020 conference.
import warnings
warnings.filterwarnings('ignore')
# # Package Import
# +
import os
import sys
sys.path.append('../')
from src.features import build_features
from src.visualization import visualize
from src.reports import make_report
import pandas as pd
import numpy as np
from datetime import datetime, timedelta
import matplotlib.pyplot as plt
import seaborn as sns
import matplotlib
import matplotlib.dates as mdates
from matplotlib.colors import ListedColormap, LinearSegmentedColormap
import matplotlib.animation as animation
# -
# # Overview of Figures
# There are few figures we want to create for the poster. In general, we want to create four figures:
# 1. Beacon
# 2. Fitbit
# 3. Beiwe
# 4. Combination of the previous three
#
# Zoltan is also very adamant about animating our figures, so we need to look into that. He sent a [link](https://towardsdatascience.com/how-to-create-animated-graphs-in-python-bb619cc2dec1) that has some promising information.
# ## Animation Test
# More information on animation can be found using these blog posts:
# - [Somewhat Helpful](https://towardsdatascience.com/how-to-create-animated-graphs-in-python-bb619cc2dec1)
# - [More Helpful](https://towardsdatascience.com/animations-with-matplotlib-d96375c5442c)
#
# And documentation on the main animate function is [here](https://matplotlib.org/3.3.2/api/_as_gen/matplotlib.animation.FuncAnimation.html)
beacon_data = pd.read_csv('../data/processed/bpeace2-beacon.csv',index_col=0,parse_dates=True)
# show it off:
beacon_data.head()
# +
beacon_bb = beacon_data[beacon_data['Beacon'] == 19][datetime(2020,7,6):datetime(2020,7,14)]
beacon_bb = beacon_bb.resample('60T').mean()
beacon_bb_pollutants = beacon_bb[['CO2','PM_C_2p5','NO2','CO','TVOC','Lux','T_CO']]
normalized_df = (beacon_bb_pollutants-beacon_bb_pollutants.min())/(beacon_bb_pollutants.max()-beacon_bb_pollutants.min())
# -
Writer = animation.writers['ffmpeg']
writer = Writer(fps=20, metadata=dict(artist='Me'), bitrate=1800)
# +
# #%matplotlib notebook
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from matplotlib import animation
fig = plt.figure()
frames = 50
def init():
initial = normalized_df.T
initial = initial.replace(initial, 0)
sns.heatmap(initial, vmax=1, cbar=False)
def animate(i):
data = normalized_df.T[normalized_df.T < i/frames]
sns.heatmap(data, vmax=1, cbar=False)
anim = animation.FuncAnimation(fig, animate, init_func=init, frames=frames, interval=1000, repeat = False)
anim.save('../test_heatmap.mp4', writer=writer)
# -
# # Beacon
# <div class="alert alert-block alert-success">
# We are looking for a good summary of a week's worth of data. Looks:
# <ul>
# <li>week of July 6 is a good place to start</li>
# <li>Beacon 19</li>
# </ul>
# </div>
def create_cmap(colors,nodes):
cmap = LinearSegmentedColormap.from_list("mycmap", list(zip(nodes, colors)))
return cmap
# ## Static Plot
# +
# Creating the dataframe
beacon_bb = beacon_data[beacon_data['Beacon'] == 19][datetime(2020,7,6):datetime(2020,7,14)]
beacon_bb = beacon_bb.resample('60T').mean()
beacon_bb_pollutants = beacon_bb[['CO2','PM_C_2p5','NO2','CO','TVOC','Lux','T_CO']]
fig, axes = plt.subplots(7,1,figsize=(16,7),sharex=True)
ylabels = ['CO$_2$',
'PM$_{2.5}$',
'NO$_2$',
'CO',
'TVOC',
'Light',
'T']
other_ylabels = ['ppm','$\mu$g/m$^3$','ppb','ppm','ppb','lux', '$^\circ$C']
cbar_ticks = [np.arange(400,1200,200),
np.arange(0,40,10),
np.arange(0,120,50),
np.arange(0,15,4),
[0,200,500,700],
np.arange(0,250,50),
np.arange(18,32,4)]
cmaps = [create_cmap(["green", "yellow", "orange", "red",],[0.0, 0.33, 0.66, 1]),
create_cmap(["green", "yellow", "orange", "red"],[0.0, 0.2, 0.4, 1]),
create_cmap(["green", "yellow", "orange", "red"],[0.0, 0.33, 0.66, 1]),
create_cmap(["green", "yellow", "orange", "red"],[0.0, 0.375, 0.75, 1]),
create_cmap(["green", "yellow", "orange", "red"],[0.0, 0.1, 0.31, 1]),
create_cmap(["black","purple","red","orange","yellow","green"],[0.0, 0.1, 0.16, 0.2, 0.64, 1]),
create_cmap(["cyan","blue","green","orange","red"],[0.0, 0.2, 0.4, 0.7, 1])]
for ax, var, low, high, ylabel, other_y, ticks, cmap in zip(axes,beacon_bb_pollutants.columns,[400,0,0,0,0,0,18],[1000,30,100,12,700,200,30],ylabels,other_ylabels,cbar_ticks,cmaps):
sns.heatmap(beacon_bb_pollutants[[var]].T,vmin=low,vmax=high,ax=ax,cbar_kws={'ticks':ticks},cmap=cmap)
ax.set_ylabel('')
ax.text(0,-0.1,f'{ylabel} ({other_y})')
ax.set_yticklabels([''])
ax.set_xlabel('')
xlabels = ax.get_xticklabels()
new_xlabels = []
for label in xlabels:
new_xlabels.append(label.get_text()[11:16])
ax.set_xticklabels(new_xlabels)
plt.subplots_adjust(hspace=0.6)
plt.savefig('../reports/BuildSys2020/beacon_heatmap.pdf',bbox_inches='tight')
plt.show()
plt.close()
# -
# ## Animation
# +
# #%matplotlib notebook
# Creating the dataframe
beacon_bb = beacon_data[beacon_data['Beacon'] == 19][datetime(2020,7,6):datetime(2020,7,14)]
beacon_bb = beacon_bb.resample('60T').mean()
beacon_bb_pollutants = beacon_bb[['CO2','PM_C_2p5','NO2','CO','TVOC','Lux','T_CO']]
# creating the figure
fig, axes = plt.subplots(7,1,figsize=(12,8),sharex=True)
ylabels = ['CO$_2$',
'PM$_{2.5}$',
'NO$_2$',
'CO',
'TVOC',
'Light',
'T']
other_ylabels = ['ppm','$\mu$g/m$^3$','ppb','ppm','ppb','lux', '$^\circ$C']
cbar_ticks = [np.arange(400,1200,200),
np.arange(0,40,10),
np.arange(0,120,50),
np.arange(0,15,4),
[0,200,500,700],
np.arange(0,250,50),
np.arange(18,32,4)]
cmaps = [create_cmap(["green", "yellow", "orange", "red",],[0.0, 0.33, 0.66, 1]),
create_cmap(["green", "yellow", "orange", "red"],[0.0, 0.2, 0.4, 1]),
create_cmap(["green", "yellow", "orange", "red"],[0.0, 0.33, 0.66, 1]),
create_cmap(["green", "yellow", "orange", "red"],[0.0, 0.375, 0.75, 1]),
create_cmap(["green", "yellow", "orange", "red"],[0.0, 0.1, 0.31, 1]),
create_cmap(["black","purple","red","orange","yellow","green"],[0.0, 0.1, 0.16, 0.2, 0.64, 1]),
create_cmap(["cyan","blue","green","orange","red"],[0.0, 0.2, 0.4, 0.7, 1])]
for ax, var, low, high, ylabel, other_y, ticks, cmap in zip(axes,beacon_bb_pollutants.columns,[400,0,0,0,0,0,18],[1000,30,100,12,700,200,30],ylabels,other_ylabels,cbar_ticks,cmaps):
sns.heatmap(beacon_bb_pollutants[[var]].T,vmin=low,vmax=high,ax=ax,cbar_kws={'ticks':ticks},cmap=cmap)
ax.set_ylabel('')
ax.text(0,-0.1,f'{ylabel} ({other_y})')
ax.set_yticklabels([''])
ax.set_xlabel('')
xlabels = ax.get_xticklabels()
new_xlabels = []
for label in xlabels:
new_xlabels.append(label.get_text()[11:16])
ax.set_xticklabels(new_xlabels)
plt.subplots_adjust(hspace=0.6)
# Animation
frames = 193
Writer = animation.writers['ffmpeg']
writer = Writer(fps=20, metadata=dict(artist='Me'), bitrate=1800)
def init():
initial = beacon_bb_pollutants
initial = initial.replace(initial, 0)
for ax, var in zip(axes,beacon_bb_pollutants.columns):
sns.heatmap(initial[[var]].T, cbar=False, ax=ax)
plt.subplots_adjust(hspace=0.6)
def animate(i):
ylabels = ['CO$_2$',
'PM$_{2.5}$',
'NO$_2$',
'CO',
'TVOC',
'Light',
'T']
other_ylabels = ['ppm','$\mu$g/m$^3$','ppb','ppm','ppb','lux', '$^\circ$C']
cbar_ticks = [np.arange(400,1200,200),
np.arange(0,40,10),
np.arange(0,120,50),
np.arange(0,15,4),
[0,200,500,700],
np.arange(0,250,50),
np.arange(18,32,4)]
cmaps = [create_cmap(["green", "yellow", "orange", "red",],[0.0, 0.33, 0.66, 1]),
create_cmap(["green", "yellow", "orange", "red"],[0.0, 0.2, 0.4, 1]),
create_cmap(["green", "yellow", "orange", "red"],[0.0, 0.33, 0.66, 1]),
create_cmap(["green", "yellow", "orange", "red"],[0.0, 0.375, 0.75, 1]),
create_cmap(["green", "yellow", "orange", "red"],[0.0, 0.1, 0.31, 1]),
create_cmap(["black","purple","red","orange","yellow","green"],[0.0, 0.1, 0.16, 0.2, 0.64, 1]),
create_cmap(["cyan","blue","green","orange","red"],[0.0, 0.2, 0.4, 0.7, 1])]
for ax, var, low, high, ylabel, other_y, ticks, cmap in zip(axes,beacon_bb_pollutants.columns,[400,0,0,0,0,0,18],[1000,30,100,12,700,200,30],ylabels,other_ylabels,cbar_ticks,cmaps):
df = beacon_bb_pollutants[[var]].T
#df = df[df <= np.nanmax(df)*((i+1)/frames)]
df = df.iloc[:,:i+1]
sns.heatmap(df,vmin=low,vmax=high,ax=ax,cbar=False,cmap=cmap)
ax.set_ylabel('')
ax.text(0,-0.1,f'{ylabel} ({other_y})')
ax.set_yticklabels([''])
ax.set_xlabel('')
xlabels = ax.get_xticklabels()
new_xlabels = []
for label in xlabels:
if label.get_text()[11:13] == '00':
new_xlabels.append(label.get_text()[5:10] + ' ' + label.get_text()[11:16])
else:
new_xlabels.append(label.get_text()[11:16])
ax.set_xticklabels(new_xlabels)
anim = animation.FuncAnimation(fig, animate, init_func=init, frames=frames, interval=500, repeat = False)
anim.save('../reports/BuildSys2020/beacon_heatmap.mp4', writer=writer)
# -
# # Fitbit
# In terms of purely Fitbit data, I was thinking of just showing something simple like daily steps with an overlay of the hourly numbers.
fb_hourly = pd.read_csv('../data/processed/bpeace2-fitbit-intraday.csv',
index_col=0,parse_dates=True,infer_datetime_format=True)
fb_hourly.head()
fb_daily = pd.read_csv('../data/processed/bpeace2-fitbit-daily.csv',
index_col=0,parse_dates=True,infer_datetime_format=True)
fb_daily.head()
# The participant that corresponds to beacon 19 is qh34m4r9. We will continue to use this participant, at least initially to make sure all the data overlap.
fb19_hourly = fb_hourly[fb_hourly['beiwe'] == 'qh34m4r9'][datetime(2020,7,6):datetime(2020,7,14)]
fb19_hourly = fb19_hourly.resample('60T').sum()
fb19_hourly.head()
fb19_daily = fb_daily[fb_daily['beiwe'] == 'qh34m4r9'][datetime(2020,7,6):datetime(2020,7,14)]
fb19_daily.head()
# ## Static Plot
# +
fig, ax1 = plt.subplots(figsize=(8,4.5))
ax2 = ax1.twinx()
# Daily Steps - Bar
ax1.stem(fb19_daily.index, fb19_daily['activities_steps'], linefmt='k-',markerfmt='ko',basefmt='w ')
# Formatting the x-axis
ax1.xaxis.set_tick_params(rotation=-30)
ax1.set_xlim([datetime(2020,7,5,20),datetime(2020,7,14,4)])
# Formatting the y-axis (daily)
ax1.set_yticks([0,2000,4000,6000,8000])
ax1.set_ylabel('Daily Steps')
# Hourly Steps - Time Series
scatter_color = '#bf5700'
ax2.scatter(fb19_hourly.index.values, fb19_hourly['activities_steps'].values,marker='s',s=5,color=scatter_color)
# Formatting the y-axis (hourly)
ax2.set_yticks([0,1000,2000,3000,4000])
ax2.spines['right'].set_color(scatter_color)
ax2.yaxis.label.set_color(scatter_color)
ax2.tick_params(axis='y', colors=scatter_color)
ax2.set_ylabel('Hourly Steps')
# Formatting the x-axis (again)
ax2.xaxis.set_major_locator(mdates.DayLocator())
myFmt = mdates.DateFormatter('%m/%d')
ax2.xaxis.set_major_formatter(myFmt)
# Saving
plt.savefig('../reports/BuildSys2020/fitbit_steps.pdf')
plt.show()
plt.close()
# -
# # Beiwe
# The best and simplest thing I can think to do with Beiwe is to include the submission times for the morning and weekly surveys.
ema_morning = pd.read_csv('../data/processed/bpeace2-morning-survey.csv',
index_col=0,parse_dates=True,infer_datetime_format=True)
ema_morning.sort_index(inplace=True)
ema_morning.head()
ema_evening = pd.read_csv('../data/processed/bpeace2-evening-survey.csv',
index_col=0,parse_dates=True,infer_datetime_format=True)
ema_evening.sort_index(inplace=True)
ema_evening.head()
morning_b19 = ema_morning[ema_morning['ID'] == 'qh34m4r9'][datetime(2020,7,6):datetime(2020,7,14)]
morning_b19.head()
evening_b19 = ema_evening[ema_evening['ID'] == 'qh34m4r9'][datetime(2020,7,6):datetime(2020,7,14)]
evening_b19.head()
# ## Static Plot
# ### First attempt
# Not well-received primarily because the way contentment is framed compared to the other moods.
# +
fig, ax = plt.subplots(figsize=(8,4.5))
# Plotting submission Times
for i in range(len(morning_b19)):
if i == 0:
ax.axvline(morning_b19.index[i],color='goldenrod',linestyle='dashed',zorder=1,label='Morning')
ax.axvline(evening_b19.index[i],color='indigo',linestyle='dashed',zorder=2,label='Evening')
else:
ax.axvline(morning_b19.index[i],color='goldenrod',linestyle='dashed',zorder=1)
ax.axvline(evening_b19.index[i],color='indigo',linestyle='dashed',zorder=2)
# Plotting mood scores
i = 0
for df in [morning_b19,evening_b19]:
for mood, color, offset in zip(['Content','Stress','Lonely','Sad'],['seagreen','firebrick','gray','navy'],[270,90,-90,-270]):
if i == 0:
ax.scatter(df.index+timedelta(minutes=offset),df[mood].values,color=color,zorder=10,label=mood)
else:
ax.scatter(df.index+timedelta(minutes=offset),df[mood].values,color=color,zorder=10)
i+= 1
# Formatting x-axis
ax.set_xlim([datetime(2020,7,6),datetime(2020,7,14,4)])
ax.xaxis.set_major_locator(mdates.DayLocator())
myFmt = mdates.DateFormatter('%m/%d')
ax.xaxis.set_major_formatter(myFmt)
# Formatting y-axis
ax.set_ylim([-0.5,3.5])
ax.set_yticks([0,1,2,3])
ax.set_yticklabels(['Not at all','A little bit','Quite a bit','Very much'])
ax.legend(loc='upper center',bbox_to_anchor=(1.1,1),ncol=1,frameon=False)
#plt.savefig('../reports/BuildSys2020/beiwe_mood.pdf')
plt.show()
plt.close()
# -
# ### Second Attempt
# Now for the second attempt where we keep the mood on the same axis and color the points based on the response
# +
fig, ax = plt.subplots(figsize=(8,4.5))
# Plotting submission Times
for i in range(len(morning_b19)):
if i == 0:
ax.axvline(morning_b19.index[i],color='gray',linestyle='dashed',zorder=1,label='Morning')
ax.axvline(evening_b19.index[i],color='black',linestyle='dashed',zorder=2,label='Evening')
else:
ax.axvline(morning_b19.index[i],color='gray',linestyle='dashed',zorder=1)
ax.axvline(evening_b19.index[i],color='black',linestyle='dashed',zorder=2)
# Plotting mood scores
i = 0
for df in [morning_b19,evening_b19]:
j = 0
for mood, color in zip(['Content','Stress','Lonely','Sad'],['seagreen','firebrick','gray','navy']):
ys = []
shades = []
for val in df[mood]:
if val == 0:
shades.append('red')
elif val == 1:
shades.append('orange')
elif val == 2:
shades.append('blue')
elif val == 3:
shades.append('green')
else:
shades.append('white')
ys.append(j)
if i == 0:
ax.scatter(df.index,ys,color=shades,edgecolor='gray',marker='s',s=200,zorder=10)
else:
ax.scatter(df.index,ys,color=shades,edgecolor='black',marker='s',s=200,zorder=10)
j += 1
i+= 1
# Formatting x-axis
ax.set_xlim([datetime(2020,7,6),datetime(2020,7,14,4)])
ax.xaxis.set_major_locator(mdates.DayLocator())
myFmt = mdates.DateFormatter('%m/%d')
ax.xaxis.set_major_formatter(myFmt)
# Formatting y-axis
ax.set_ylim([-0.5,3.5])
ax.set_yticks([0,1,2,3])
ax.set_yticklabels(['Content','Stress','Lonely','Sad'])
ax.legend(loc='upper center',bbox_to_anchor=(1.1,1),ncol=1,frameon=False)
plt.savefig('../reports/BuildSys2020/beiwe_mood.pdf')
plt.show()
plt.close()
# -
# Figure is decent and the iffy parts can be removed in illustrator.
| notebooks/archive/0.3.0-hef-BuildSys20-figures.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
vet = np.arange(50).reshape((5,10))
vet
vet.shape
vet[0:2][:1]
vet2 = vet[:4]
vet2
vet2[:] = 100
vet2
vet
vet2 = vet[:3].copy()
vet2
vet[1:4,5:7]
bol = vet > 50
bol
vet[bol]
vet3 = np.linspace(0,100,30)
vet3.shape
vet31 = vet3.reshape(3, 10)
vet31
vet31[0:2, 2]
| secao5 - Python para Analise de dados utizando Numpy/indexacao e fatiamento de arrays.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.2 64-bit
# metadata:
# interpreter:
# hash: 31f2aee4e71d21fbe5cf8b01ff0e069b9275f58929596ceb00d14d90e3e16cd6
# name: Python 3.8.2 64-bit
# ---
# We will use different algorithms to create models for predicting fraudulant transactions.
# Data source: https://www.kaggle.com/mlg-ulb/creditcardfraud
# 1. Let's try XGBoost. Tutorial: https://machinelearningmastery.com/develop-first-xgboost-model-python-scikit-learn/
# Importing classes that will be used later.
from numpy import loadtxt
from xgboost import XGBClassifier
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
# +
# load data
import pandas as pd
import numpy as np
import warnings
warnings.filterwarnings('ignore')
df = pd.read_csv('train/creditcard.csv')
df.head()
# -
# Split training data.
from sklearn.model_selection import train_test_split
X = df.drop('Class',axis=1)
y = df['Class']
seed = 7
test_size = 0.33
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=test_size, random_state=seed)
# Train the model.
model = XGBClassifier()
model.fit(X_train, y_train)
# Make predictions for test data.
y_pred = model.predict(X_test)
predictions = [round(value) for value in y_pred]
# + tags=[]
# Evaluate predictions.
accuracy = accuracy_score(y_test, predictions)
print("Accuracy: %.2f%%" % (accuracy * 100.0))
# Here we show the logic of accuracy_score: it basically counts the percentage of non-diff.
diff = 0
for i in range(len(y_test)):
if y_test[i] != predictions[i]: diff += 1
print("diff: {}, non-diff rate: {}%".format(diff,100-round(diff/len(y_test)*100,2)))
# + tags=[]
# Check out confusion matrix and calculate recall and accuracy.
from sklearn.metrics import confusion_matrix
data = confusion_matrix(y_test,predictions)
recall,precision = data[1][1]/sum(data[1]),data[1][1]/(data[0][1]+data[1][1])
print('recall is {}%, precision is {}%'.format(round(recall*100,1),round(precision*100,1)))
# +
# Let's try normalize training data with StandardScalar, which makes sure the standard deviation of the data is 1 and mean is 0. Then we run the training and testing again.
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
# Convert to numpy arrays for tensorflow.
y_train = y_train.values
y_test = y_test.values
# Transform to scaled.
X_train_scaled = scaler.fit_transform(X_train,y_train)
X_test_scaled = scaler.transform(X_test)
# -
# Train again.
model_scaled = XGBClassifier()
model_scaled.fit(X_train_scaled, y_train)
# Predict again.
y_pred_scaled = model_scaled.predict(X_test_scaled)
predictions_scaled = [round(value) for value in y_pred_scaled]
# + tags=[]
# Let's see recall and accuracy. It is the same as non-scaled version.
# It seems normalization is not needed for XGBoost: https://github.com/dmlc/xgboost/issues/2621
# Both decision trees and random forest are not sensitive to monotonic transformation: https://stats.stackexchange.com/questions/353462/what-are-the-implications-of-scaling-the-features-to-xgboost
data = confusion_matrix(y_test,predictions_scaled)
recall,precision = data[1][1]/sum(data[1]),data[1][1]/(data[0][1]+data[1][1])
print('recall is {}%, precision is {}%'.format(round(recall*100,1),round(precision*100,1)))
# + tags=[]
# Plot ROC curve.
import matplotlib.pyplot as plt
from sklearn.metrics import roc_curve, auc
fpr, tpr, _ = roc_curve(y_test, predictions)
roc_auc = auc(fpr, tpr)
#xgb.plot_importance(gbm)
#plt.show()
plt.figure()
lw = 2
plt.plot(fpr, tpr, color='darkorange',
lw=lw, label='ROC curve (area = %0.2f)' % roc_auc)
plt.plot([0, 1], [0, 1], color='navy', lw=lw, linestyle='--')
plt.xlim([-0.02, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC curve')
plt.legend(loc="lower right")
plt.show()
# -
| cc_fraud.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# import stuff
import os
import numpy as np
#import pandas as pd
from math import sqrt as sqrt
from itertools import product as product
import torch
import torch.utils.data as data
from torchvision import models
import torch.nn as nn
import torch.nn.init as init
import torch.nn.functional as F
from torch.autograd import Function
import pandas as pd
# -
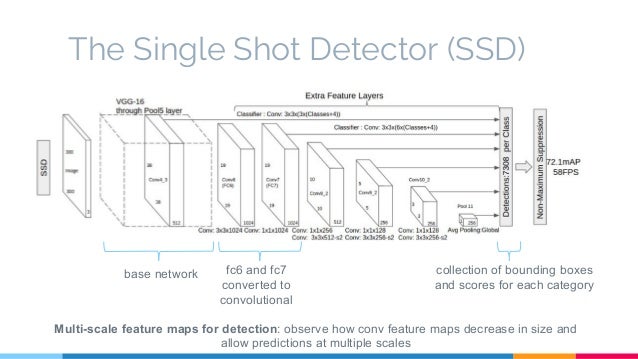
# # SSDメインモデルを構築
# VGGベースモデルを構築。
#
# TODO: Resnetベースへの改良?どこにつなげればよいか、正規化などを入れないといけないので参考文献などを読みたい。\
# 
# plot vgg model
vgg = models.vgg16(pretrained=False)
print(vgg)
# +
# VGG
# 転移学習なしで学習するのか?
def make_vgg():
layers = []
in_channels = 3
# VGGのモデル構造を記入
cfg = [64, 64, "M", 128, 128, "M", 256, 256, 256, "MC", 512, 512, 512, "M", 512, 512, 512]
for v in cfg:
if v == "M":
layers += [nn.MaxPool2d(kernel_size=2, stride=2)]
elif v == "MC":
layers += [nn.MaxPool2d(kernel_size=2, stride=2, ceil_mode=True)]
else:
conv2d = nn.Conv2d(in_channels, v, kernel_size=3, padding=1)
layers += [conv2d, nn.ReLU(inplace=True)] #メモリ節約
in_channels = v
pool5 = nn.MaxPool2d(kernel_size=3, stride=1, padding=1)
conv6 = nn.Conv2d(512, 1024, kernel_size=3, padding=6, dilation=6)
conv7 = nn.Conv2d(1024, 1024, kernel_size=1)
layers += [pool5, conv6, nn.ReLU(inplace=True), conv7, nn.ReLU(inplace=True)]
return nn.ModuleList(layers)
# 動作確認
vgg_test = make_vgg()
print(vgg_test)
# +
# 小さい物体のbbox検出用のextras moduleを追加
def make_extras():
layers = []
in_channels = 1024 # vgg module outputs
# extra modeule configs
cfg = [256, 512, 128, 256, 128, 256, 128, 256]
layers += [nn.Conv2d(in_channels, cfg[0], kernel_size=1)]
layers += [nn.Conv2d(cfg[0], cfg[1], kernel_size=3, stride=2, padding=1)]
layers += [nn.Conv2d(cfg[1], cfg[2], kernel_size=1)]
layers += [nn.Conv2d(cfg[2], cfg[3], kernel_size=3, stride=2, padding=1)]
layers += [nn.Conv2d(cfg[3], cfg[4], kernel_size=1)]
layers += [nn.Conv2d(cfg[4], cfg[5], kernel_size=3)]
layers += [nn.Conv2d(cfg[5], cfg[6], kernel_size=1)]
layers += [nn.Conv2d(cfg[6], cfg[7], kernel_size=3)]
return nn.ModuleList(layers)
extras_test = make_extras()
print(extras_test)
# -
# ## locとconfに対するモジュール。
# +
# locとconfモジュールを作成
def make_loc_conf(num_classes=21, bbox_aspect_num=[4, 6, 6, 6, 4, 4]):
loc_layers = []
conf_layers = []
# VGGの中間出力に対するレイヤ
loc_layers += [nn.Conv2d(512, bbox_aspect_num[0] * 4,
kernel_size=3, padding=1)]
conf_layers += [nn.Conv2d(512, bbox_aspect_num[0] * num_classes,
kernel_size=3, padding=1)]
# VGGの最終そうに対するCNN
loc_layers += [nn.Conv2d(1024, bbox_aspect_num[1] * 4,
kernel_size=3, padding=1)]
conf_layers += [nn.Conv2d(1024, bbox_aspect_num[1] * num_classes,
kernel_size=3, padding=1)]
# source3
loc_layers += [nn.Conv2d(512, bbox_aspect_num[2] * 4,
kernel_size=3, padding=1)]
conf_layers += [nn.Conv2d(512, bbox_aspect_num[2] * num_classes,
kernel_size=3, padding=1)]
# source4
loc_layers += [nn.Conv2d(256, bbox_aspect_num[3] * 4,
kernel_size=3, padding=1)]
conf_layers += [nn.Conv2d(256, bbox_aspect_num[3] * num_classes,
kernel_size=3, padding=1)]
# source5
loc_layers += [nn.Conv2d(256, bbox_aspect_num[4] * 4,
kernel_size=3, padding=1)]
conf_layers += [nn.Conv2d(256, bbox_aspect_num[4] * num_classes,
kernel_size=3, padding=1)]
# source6
loc_layers += [nn.Conv2d(256, bbox_aspect_num[5] * 4,
kernel_size=3, padding=1)]
conf_layers += [nn.Conv2d(256, bbox_aspect_num[5] * num_classes,
kernel_size=3, padding=1)]
return nn.ModuleList(loc_layers), nn.ModuleList(conf_layers)
loc_test, conf_test = make_loc_conf()
print(loc_test)
print(conf_test)
# -
# ## L2 normの実装
# ありなしで性能はどう変化するのか?
# 自作レイヤ
class L2Norm(nn.Module):
def __init__(self, input_channels=512, scale=20):
super(L2Norm, self).__init__()
self.weight = nn.Parameter(torch.Tensor(input_channels))
self.scale = scale
self.reset_parameters()
self.eps = 1e-10
def reset_parameters(self):
init.constant_(self.weight, self.scale) # weightの値が全てscaleになる
def forward(self, x):
"""
38x38の特徴量に対し、チャネル方向の和を求めそれを元に正規化する。
また正規化したあとに係数(weight)をかける(ロスが減るように学習してくれるみたい)
"""
norm = x.pow(2).sum(dim=1, keepdim=True).sqrt()+self.eps #チャネル方向の自乗和
x = torch.div(x, norm) # 正規化
weights = self.weight.unsqueeze(0).unsqueeze(2).unsqueeze(3).expand_as(x) # 学習させるパラメータ
out = weights * x
return out
# bbox
for i, j in product(range(3), repeat=2):
print(i, j)
# +
# binding boxを出力するクラス
class DBox(object):
def __init__(self, cfg):
super(DBox, self).__init__()
self.image_size = cfg["input_size"]
# 各sourceの特徴量マップのサイズ
self.feature_maps = cfg["feature_maps"]
self.num_priors = len(cfg["feature_maps"]) # number of sources
self.steps = cfg["steps"] #各boxのピクセルサイズ
self.min_sizes = cfg["min_sizes"] # 小さい正方形のサイズ
self.max_sizes = cfg["max_sizes"] # 大きい正方形のサイズ
self.aspect_ratios = cfg["aspect_ratios"]
def make_dbox_list(self):
mean = []
# feature maps = 38, 19, 10, 5, 3, 1
for k, f in enumerate(self.feature_maps):
for i, j in product(range(f), repeat=2):
# fxf画素の組み合わせを生成
f_k = self.image_size / self.steps[k]
# 300 / steps: 8, 16, 32, 64, 100, 300
# center cordinates normalized 0~1
cx = (j + 0.5) / f_k
cy = (i + 0.5) / f_k
# small bbox [cx, cy, w, h]
s_k = self.min_sizes[k] / self.image_size
mean += [cx, cy, s_k, s_k]
# larger bbox
s_k_prime = sqrt(s_k * (self.max_sizes[k]/self.image_size))
mean += [cx, cy, s_k_prime, s_k_prime]
# その他のアスペクト比のdefbox
for ar in self.aspect_ratios[k]:
mean += [cx, cy, s_k*sqrt(ar), s_k/sqrt(ar)]
mean += [cx, cy, s_k/sqrt(ar), s_k*sqrt(ar)]
# convert the list to tensor
output = torch.Tensor(mean).view(-1, 4)
# はみ出すのを防ぐため、大きさを最小0, 最大1にする
output.clamp_(max=1, min=0)
return output
# +
# test boxes
# SSD300の設定
ssd_cfg = {
'num_classes': 21, # 背景クラスを含めた合計クラス数
'input_size': 300, # 画像の入力サイズ
'bbox_aspect_num': [4, 6, 6, 6, 4, 4], # 出力するDBoxのアスペクト比の種類
'feature_maps': [38, 19, 10, 5, 3, 1], # 各sourceの画像サイズ
'steps': [8, 16, 32, 64, 100, 300], # DBOXの大きさを決める
'min_sizes': [30, 60, 111, 162, 213, 264], # DBOXの大きさを決める
'max_sizes': [60, 111, 162, 213, 264, 315], # DBOXの大きさを決める
'aspect_ratios': [[2], [2, 3], [2, 3], [2, 3], [2], [2]],
}
dbox = DBox(ssd_cfg)
dbox_list = dbox.make_dbox_list()
pd.DataFrame(dbox_list.numpy())
# -
# # SSDクラスを実装する
# +
class SSD(nn.Module):
def __init__(self, phase, cfg):
super(SSD, self).__init__()
self.phase = phase
self.num_classes = cfg["num_classes"]
# call SSD network
self.vgg = make_vgg()
self.extras = make_extras()
self.L2Norm = L2Norm()
self.loc, self.conf = make_loc_conf(self.num_classes, cfg["bbox_aspect_num"])
# make Dbox
dbox = DBox(cfg)
self.dbox_list = dbox.make_dbox_list()
# use Detect if inference
if phase == "inference":
self.detect = Detect()
# check operation
ssd_test = SSD(phase="train", cfg=ssd_cfg)
print(ssd_test)
# +
def decode(loc, dbox_list):
"""
DBox(cx,cy,w,h)から回帰情報のΔを使い、
BBox(xmin,ymin,xmax,ymax)方式に変換する。
loc: [8732, 4] [Δcx, Δcy, Δw, Δheight]
SSDのオフセットΔ情報
dbox_list: (cx,cy,w,h)
"""
boxes = torch.cat((
dbox_list[:, :2] + loc[:, :2] * 0.1 * dbox_list[:, :2],
dbox_list[:, 2:] * torch.exp(loc[:, 2:] * 0.2)), dim=1)
# convert boxes to (xmin,ymin,xmax,ymax)
boxes[:, :2] -= boxes[:, 2:] / 2
boxes[:, 2:] += boxes[:, :2]
return boxes
# test
dbox = DBox(ssd_cfg)
dbox_list = dbox.make_dbox_list()
print(dbox_list.size())
loc = torch.ones(8732, 4)
loc[0, :] = torch.tensor([-10, 0, 1, 1])
print(loc.size())
dbox_process = decode(loc, dbox_list)
pd.DataFrame(dbox_process.numpy())
# -
def nms(boxes, scores, overlap=0.45, top_k=200):
"""
overlap以上のディテクションに関して信頼度が高い方をキープする。
キープしないものは消去。
物体のクラス毎にnmsは実効する。
------------------
inputs:
scores: bboxの信頼度
bbox: bboxの座標情報
------------------
出力:
keep:
"""
# returnを定義
count = 0
keep = scores.new(scores.size(0)).zero_().long()
print(keep.size())
# keep: 確信度thresholdを超えたbboxの数
# 各bboxの面積を計算
x1 = boxes[:, 0]
x2 = boxes[:, 2]
y1 = boxes[:, 1]
y2 = boxes[:, 3]
area = torch.mul(x2 - x1, y2 - y1)
# copy boxes
tmp_x1 = boxes.new()
tmp_y1 = boxes.new()
tmp_x2 = boxes.new()
tmp_y2 = boxes.new()
tmp_w = boxes.new()
tmp_h = boxes.new()
# sort scores 高い信頼度のものを上に。
v, idx = scores.sort(0)
# topk個の箱のみ取り出す
idx = idx[-top_k:]
# indexの要素数が0でない限りループする。
while idx.numel() > 0:
i = idx[-1] # 一番高い信頼度のboxを指定
# keep の最後にconf最大のindexを格納
keep[count] = i
count += 1
# 最後の一つになったらbreak
if idx.size(0) == 1:
break
# indexをへらす
idx = idx[:-1]
# ------------------------------------
# このboxとiouの大きいboxを消していく。
# ------------------------------------
# torch.index_select(input, dim, index, out=None) → Tensor
torch.index_select(x1, 0, idx, out=tmp_x1)
torch.index_select(y1, 0, idx, out=tmp_y1)
torch.index_select(x2, 0, idx, out=tmp_x2)
torch.index_select(y2, 0, idx, out=tmp_y2)
# target boxの最小、最大にclamp
tmp_x1 = torch.clamp(tmp_x1, min=x1[i])
tmp_y1 = torch.clamp(tmp_y1, min=y1[i])
tmp_x2 = torch.clamp(tmp_x2, min=x2[i])
tmp_y2 = torch.clamp(tmp_y2, min=y2[i])
# wとhのテンソルサイズをindex一つ減らしたものにする
tmp_w.resize_as_(temp_x2)
tmp_h.resize_as_(temp_y2)
# clampした状態の高さ、幅を求める
tmp_w = tmp_x2 - tmp_x1
tmp_h = tmp_y2 - tmp_y1
# 幅や高さが負になっているものは0に
tmp_w = torch.clamp(tmp_w, min=0.0)
tmp_h = torch.clamp(tmp_h, min=0.0)
# clamp時の面積を導出
inter = tmp_w * tmp_h # オーバラップしている面積
# IoU の計算
# intersect=overlap
# IoU = intersect部分 / area(a) + area(b) - intersect
rem_areas = torch.index_select(area, 0, idx) # bbox元の面積
union = rem_areas + area[i] - inter
IoU = inter / union
# IoUがしきい値より大きいものは削除
idx = idx[IoU.le(overlap)] # leはless than or eqal to
return keep, count
# # 推論用のクラスDetectの実装
# +
class Detect(Function):
def __init__(self, conf_thresh=0.01, top_k=200, nms_thresh=0.45):
self.softmax = nn.Softmax(dim=-1)
self.conf_thresh = conf_thresh
self.top_k = top_k
self.nms_thresh = nms_thresh
def forward(self, loc_data, conf_data, dbox_list):
"""
SSDの推論結果を受け取り、bboxのデコードとnms処理を行う。
"""
#
num_batch = loc_data.size(0)
num_dbox = loc_data.size(1)
num_classes = conf_data.size(2)
# confをsoftmaxを使って正規化
conf_data = self.softmax(conf_data)
# 出力の方を作成する
# [batch, class, topk, 5]
output = torch.zeros(num_batch, num_classes, self.top_k, 5)
# conf_dataを[batch, 8732, classes]から[batch, classes, 8732]に変更
conf_preds = conf_data.tranpose(2, 1)
# batch毎にループ
for i in range(num_batch):
# 1. LocとDBoxからBBox情報に変換
decoded_boxes = decode(loc_data, dbox_list)
# confのコピー
conf_scores = conf_preds[i].clone()
# classごとにデコードとNMSを回す。
for cl in range(1, num_classes): # 背景は飛ばす。
# 2. 敷地を超えた結果を取り出す
c_mask = conf_scores[cl].gt(self.conf_thresh) # gt=greater than
# index maskを作成した。
# threshを超えると1, 超えなかったら0に。
# c_mask = [8732]
scores = conf_scores[cl][c_mask]
if scores.nelement() == 0:
continue
# 箱がなかったら終わり。
# cmaskをboxに適応できるようにサイズ変更
l_mask = c_mask.unsqueeze(1).expand_as(decoded_boxes)
# l_mask.size = [8732, 4]
boxes = decoded_boxes[l_mask].view(-1, 4) # reshape to [boxnum, 4]
# 3. NMSを適応する
ids, count = nms(boxes, scores, self.nms_thresh, self.top_k)
# torch.cat(tensors, dim=0, out=None) → Tensor
output[i, cl, :count] = torch.cat((scores[ids[:count]].unsqueeze(1), boxes[ids[:count]]),
1)
return output # torch.size([batch, 21, 200, 5])
# -
class SSD(nn.Module):
def __init__(self, phase, cfg):
super(SSD, self).__init__()
self.phase = phase
self.num_classes = cfg["num_classes"]
# call SSD network
self.vgg = make_vgg()
self.extras = make_extras()
self.L2Norm = L2Norm()
self.loc, self.conf = make_loc_conf(self.num_classes, cfg["bbox_aspect_num"])
# make Dbox
dbox = DBox(cfg)
self.dbox_list = dbox.make_dbox_list()
# use Detect if inference
if phase == "inference":
self.detect = Detect()
def forward(self, x):
sources = list()
loc = list()
conf = list()
# VGGのconv4_3まで計算
for k in range(23):
x = self.vgg[k](x)
# conv4_3の出力をL2Normに入力。source1をsourceに追加
source1 = self.L2Norm(x)
sources.append(source1)
# VGGを最後まで計算しsource2を取得
for k in range(23, len(self.vgg)):
x = self.vgg[k](x)
sources.append(x)
# extra層の計算を行う。
# source3-6に結果を格納。
for k, v in enumerate(self.extras):
x = F.relu(v(x), inplace = True)
if k % 2 == 1:
sources.append(x)
# source 1-6にそれぞれ対応するconvを適応しconfとlocを得る。
for (x, l, c) in zip(sources, self.loc, self.conf):
# Permuteは要素の順番を入れ替え
loc.append(l(x).permute(0, 2, 3, 1).contiguous())
conf.append(c(x).permute(0, 2, 3, 1).contiguous())
# convの出力は[batch, 4*anker, fh, fw]なので整形しなければならない。
# まず[batch, fh, fw, anker]に整形
# locとconfの形を変形
# locのサイズは、torch.Size([batch_num, 34928])
# confのサイズはtorch.Size([batch_num, 183372])になる
loc = torch.cat([o.view(o.size(0), -1) for o in loc], 1)
conf = torch.cat([o.view(o.size(0), -1) for o in conf], 1)
# さらにlocとconfの形を整える
# locのサイズは、torch.Size([batch_num, 8732, 4])
# confのサイズは、torch.Size([batch_num, 8732, 21])
loc = loc.view(loc.size(0), -1, 4)
conf = conf.view(conf.size(0), -1, self.num_classes)
# これで後段の処理につっこめるかたちになる。
output = (loc, conf, self.dbox_list)
if self.phase == "inference":
# Detectのforward
return self.detect(output[0], output[1], output[2])
else:
return output
| utils/ssd.ipynb |
/ ---
/ jupyter:
/ jupytext:
/ text_representation:
/ extension: .q
/ format_name: light
/ format_version: '1.5'
/ jupytext_version: 1.14.4
/ ---
/ + [markdown] cell_id="9703282e-b28b-4d15-8c0d-23b9e7bbd784" deepnote_cell_type="text-cell-h1" is_collapsed=false tags=[]
/ # Data exploration
/ + [markdown] cell_id="a7577f14-d78c-4fca-9a53-ea0a0c3e4e58" deepnote_cell_type="markdown" tags=[]
/ In this notebook we're analyzing the cleaned dataset as a first step in the project.
/
/ Here we'll plot the data as well as find summary statistics.
/ + cell_id="17ee1ebd-6222-4944-89ee-704dde66ee0c" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=3280 execution_start=1644074187803 source_hash="1643160e" tags=[]
#libs
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
import os
/ + [markdown] cell_id="de352d00-9bde-4115-9070-0644932caccf" deepnote_cell_type="markdown" tags=[]
/ Now, let's import our cleaned dataset. Remember that, because of raw data size, an auxiliar directory containing cleaned data was created.
/
/ ```
/ data/processed/cleaned_dataset.csv -> This won't be available on GitHub, only on Deepnote
/
/ data_sent_github/cleaned_dataset.csv -> This will be available on both GitHub and Deepnote
/ ```
/
/ For your convenience, I'll be working with the dataset which will be stored on GitHub.
/ + cell_id="5eae51e9-3f42-4df0-9c87-f9b5f4706345" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=5 execution_start=1644074195770 source_hash="add48eb4" tags=[]
DATA_CLEANED_DIR = os.path.join(os.getcwd(), os.pardir, 'data_sent_github')
print(DATA_CLEANED_DIR)
/ + cell_id="64d5004a-06e6-4fcd-919b-6cd903c6ede0" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=1267 execution_start=1644074199592 source_hash="e91f393c" tags=[]
df = pd.read_csv(DATA_CLEANED_DIR+'/cleaned_dataset.csv', index_col=[0])
/ + cell_id="a4081ae5-09f4-4737-96d0-000302cfeaae" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=40 execution_start=1644074203457 source_hash="f2eb8fb8" tags=[]
#Let's make sure there're no duplicates
df.drop_duplicates(subset=['CustomerId'], keep='first', inplace=True)
/ + cell_id="231b6516-fa24-4695-8269-36fae52eab0e" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=118 execution_start=1644022412198 source_hash="c085b6ba" tags=[]
df.head()
/ + cell_id="b4392250-f3ab-4db1-990d-2a635629be21" deepnote_cell_type="code" deepnote_output_heights=[520] deepnote_to_be_reexecuted=false execution_millis=8 execution_start=1644022421434 source_hash="52430027" tags=[]
df.dtypes
/ + cell_id="7f16af2c-9626-4ee8-a9a2-4b23fe8e528c" deepnote_cell_type="code" deepnote_output_heights=[232.3125] deepnote_to_be_reexecuted=false execution_millis=82 execution_start=1644074211341 source_hash="817d440e" tags=[]
#Transforming datetimes
df['application_date'] = pd.to_datetime(df['application_date'])
df['exit_date'] = pd.to_datetime(df['exit_date'])
df['birth_date'] = pd.to_datetime(df['birth_date'])
/ + cell_id="3a513dc4-a3b5-49e0-892c-045479b2ad74" deepnote_cell_type="code" deepnote_output_heights=[232] deepnote_to_be_reexecuted=false execution_millis=9 execution_start=1644075060525 source_hash="6159abf9" tags=[]
(df['exit_date'] - df['application_date']).dt.days
/ + [markdown] cell_id="482ad532-69f0-401c-8aa4-1f88503a4583" deepnote_cell_type="markdown" tags=[]
/ ### Caution
/
/ There was a mistake in the eligible column. So we're fixing it here. This will be crucial while training the classification model.
/
/ > Remember that clients whose exit date is a null datum are eligible. That is, they are still clients of the company.
/ + cell_id="b9c1c3e9-79a0-4bde-ba77-c80dc05c0dd0" deepnote_cell_type="code" deepnote_output_heights=[21] deepnote_to_be_reexecuted=false execution_millis=12 execution_start=1644075116561 source_hash="dc8e13f5" tags=[]
pd.isnull((df['exit_date'] - df['application_date']).dt.days[0])
/ + cell_id="cb70c1d1-5bad-4874-b769-c9cd88f02210" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=6 execution_start=1644075169353 source_hash="f3c2aeb2" tags=[]
eligible_days = (df['exit_date'] - df['application_date']).dt.days
/ + cell_id="6aa9fe54-c174-448f-8332-e1afd24eff05" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=3 execution_start=1644075218066 source_hash="3d91747b" tags=[]
is_eligible = []
/ + cell_id="01ae7757-4f90-48e5-99b7-5f62f2c2c69e" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=108 execution_start=1644075278183 source_hash="c820f66e" tags=[]
for i in eligible_days:
if i >= 2*365 or pd.isnull(i) == True:
is_eligible.append(1)
else:
is_eligible.append(0)
/ + cell_id="a7d7c64c-15d9-436c-8b46-f3ee73acb29d" deepnote_cell_type="code" deepnote_output_heights=[611] deepnote_to_be_reexecuted=false execution_millis=20 execution_start=1644075285648 source_hash="bf9db7f6" tags=[]
is_eligible
/ + cell_id="f69f1333-d16f-4184-ae3d-7472bdffca8e" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=36 execution_start=1644075560776 source_hash="4313bfff" tags=[]
df['eligible'] = is_eligible
/ + cell_id="6e2c094a-06d6-4b10-bff8-1d98fe5321a7" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=240 execution_start=1644075599662 source_hash="c085b6ba" tags=[]
df.head()
/ + [markdown] cell_id="0c73b4f9-a1e1-40be-af05-1c39a59d7596" deepnote_cell_type="markdown" tags=[]
/ **This dataframe will be exported again. This will be sent to both processed and auxiliar directory.**
/ + cell_id="d4f52d0b-da4c-4f49-9075-b7a8ce366653" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=1 execution_start=1644075865905 source_hash="91b23730" tags=[]
DATA_PROCESSED_DIR = os.path.join(os.getcwd(), os.pardir, 'data', 'processed')
print(DATA_PROCESSED_DIR)
/ + cell_id="264df02f-0064-4cfa-bfe8-106e383b752a" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=4744 execution_start=1644076107374 source_hash="82d57b5b" tags=[]
#Send to processed data dir
df.to_csv(DATA_PROCESSED_DIR+'/cleaned_dataset.csv')
/ + cell_id="1371590c-bf04-4283-abce-5680bfaa78a9" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=7 execution_start=1644076161552 source_hash="9f626637" tags=[]
DATA_AUX_DIR = os.path.join(os.getcwd(), os.pardir, 'data_sent_github')
print(DATA_AUX_DIR)
/ + cell_id="07718b81-dfd5-44ba-960b-864ebef863e7" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=4039 execution_start=1644076218843 source_hash="5b12965d" tags=[]
#Send to auxiliar dir
df.to_csv(DATA_AUX_DIR+'/cleaned_dataset.csv')
/ + [markdown] cell_id="0448e102-5b41-4dec-a10e-2c1752fedb5f" deepnote_cell_type="markdown" tags=[]
/ Remember that previous exports are ready to work with. Just be careful with datatypes (as always with a csv).
/ + [markdown] cell_id="dbfccc7b-861e-4f92-a87a-80db30711548" deepnote_cell_type="markdown" tags=[]
/ ### Continuing the analysis...
/ + cell_id="d7a14e1a-179d-4012-abab-440d4333633b" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=274 execution_start=1644075736853 source_hash="5d514141" tags=[]
#First summary stats
df.describe()
/ + [markdown] cell_id="8e90352c-87e4-4a85-8c72-1d8462b47ce7" deepnote_cell_type="markdown" tags=[]
/ Previous stats tell us that:
/
/ * The population is adult and most of them (at least 75%) are 45 year old or less.
/ * 25% of clients earn at least 61025.45.
/ * At least 50% of them have a credit card.
/
/ Let's visualize variables.
/ + cell_id="cb0e0328-11ce-4698-a848-1f8e3bc23920" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=1 execution_start=1644074218807 source_hash="f2f79983" tags=[]
sns.set_theme()
/ + [markdown] cell_id="6870ef6a-22ee-4389-b9ca-bdeb299def68" deepnote_cell_type="markdown" tags=[]
/ Geography:
/ + cell_id="1abc0b74-50e8-43f0-9a16-75b20388c95c" deepnote_cell_type="code" deepnote_output_heights=[79] deepnote_to_be_reexecuted=false execution_millis=13 execution_start=1643996256867 source_hash="e3fce688" tags=[]
df['Geography'].value_counts()
/ + cell_id="7ee51ae5-3cab-4907-9fb1-a0575d7ec4a2" deepnote_cell_type="code" deepnote_output_heights=[286] deepnote_to_be_reexecuted=false execution_millis=215 execution_start=1643996325371 source_hash="b3b41011" tags=[]
fig, ax = plt.subplots()
ax = df['Geography'].value_counts().plot.bar(rot=0)
plt.title('Geography')
plt.xlabel('Location')
plt.ylabel('Number of clients')
plt.show()
/ + [markdown] cell_id="e309c9b2-42bb-48d2-8998-a8eb2ca99bdf" deepnote_cell_type="markdown" tags=[]
/ Gender:
/ + cell_id="fa15c4f0-f716-4387-b38c-d3975a880259" deepnote_cell_type="code" deepnote_output_heights=[60] deepnote_to_be_reexecuted=false execution_millis=14 execution_start=1643996469494 source_hash="198459ac" tags=[]
df['Gender'].value_counts()
/ + cell_id="34154ece-7b08-49fc-92f3-3315befe2276" deepnote_cell_type="code" deepnote_output_heights=[286] deepnote_to_be_reexecuted=false execution_millis=221 execution_start=1643996966388 source_hash="c055471" tags=[]
fig, ax = plt.subplots()
ax = df['Gender'].value_counts().plot.bar(rot=0)
plt.title('Gender of clients')
plt.xlabel('Gender')
plt.ylabel('Number of clients')
plt.show()
/ + [markdown] cell_id="c69eaeeb-d111-4bd5-b22c-5c073aa3288f" deepnote_cell_type="markdown" tags=[]
/ Has credit card:
/ + cell_id="ec620058-817c-4b14-b530-933f26d7ab0a" deepnote_cell_type="code" deepnote_output_heights=[60] deepnote_to_be_reexecuted=false execution_millis=9 execution_start=1643995755064 source_hash="f00343e1" tags=[]
df['HasCrCard'].value_counts()
/ + cell_id="364fc350-dc19-451c-8cd3-971c2f579c93" deepnote_cell_type="code" deepnote_output_heights=[286] deepnote_to_be_reexecuted=false execution_millis=215 execution_start=1643995388508 source_hash="cb7a7778" tags=[]
fig, ax = plt.subplots()
ax = df['HasCrCard'].value_counts().plot.bar(rot=0)
plt.title('Clients with credit card')
plt.xlabel('1=True, 0=False')
plt.ylabel('Number of clients')
plt.show()
/ + [markdown] cell_id="1074a4fd-8f6e-41e5-86ee-fecafe69a894" deepnote_cell_type="markdown" tags=[]
/ Is active member:
/ + cell_id="e3035649-9e46-4eba-a727-300dd9077512" deepnote_cell_type="code" deepnote_output_heights=[60] deepnote_to_be_reexecuted=false execution_millis=3 execution_start=1643996155336 source_hash="67952625" tags=[]
df['IsActiveMember'].value_counts()
/ + cell_id="13906c9f-5480-4c1c-a19a-172692c8a33e" deepnote_cell_type="code" deepnote_output_heights=[286] deepnote_to_be_reexecuted=false execution_millis=270 execution_start=1643996182185 source_hash="2f21a359" tags=[]
fig, ax = plt.subplots()
ax = df['IsActiveMember'].value_counts().plot.bar(rot=0)
plt.title('Active members')
plt.xlabel('1=True, 0=False')
plt.ylabel('Number of clients')
plt.show()
/ + [markdown] cell_id="0d3e98ec-d1cc-4dc9-822b-d541b41a2573" deepnote_cell_type="markdown" tags=[]
/ Estimated salary:
/ + cell_id="79eade92-3d40-4232-ba1a-86c2abc3e92b" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=2 execution_start=1643997187980 source_hash="ff143fe" tags=[]
import math
/ + cell_id="b93e00ea-f99d-47ea-82b6-8d7dff733bcd" deepnote_cell_type="code" deepnote_output_heights=[21] deepnote_to_be_reexecuted=false execution_millis=6 execution_start=1643997244040 source_hash="ab8194ba" tags=[]
len(df['EstimatedSalary'])
/ + cell_id="217a383f-0cdb-4e49-ae32-5e5c7412806b" deepnote_cell_type="code" deepnote_output_heights=[286] deepnote_to_be_reexecuted=false execution_millis=420 execution_start=1643997710397 source_hash="c60d39dc" tags=[]
fig, ax = plt.subplots()
ax = sns.boxplot(data=df, x='EstimatedSalary')
plt.title('Boxplot of salaries')
plt.show()
/ + [markdown] cell_id="3ee56753-1173-4f09-a9c2-2f5cd416edf5" deepnote_cell_type="markdown" tags=[]
/ Products:
/ + cell_id="a7a0609b-0d56-41a2-a8d0-58d68552e6b7" deepnote_cell_type="code" deepnote_output_heights=[98] deepnote_to_be_reexecuted=false execution_millis=3 execution_start=1643997786761 source_hash="a354d396" tags=[]
df['Products'].value_counts()
/ + cell_id="e54af837-6266-41d9-8617-3b19f1c7fbdb" deepnote_cell_type="code" deepnote_output_heights=[286] deepnote_to_be_reexecuted=false execution_millis=285 execution_start=1643997829802 source_hash="640dc5b" tags=[]
fig, ax = plt.subplots()
ax = df['Products'].value_counts().plot.bar(rot=0)
plt.title('Products clients bought')
plt.xlabel('Products')
plt.ylabel('Number of clients')
plt.show()
/ + [markdown] cell_id="50877055-27ff-4b38-b543-e671a33edae3" deepnote_cell_type="markdown" tags=[]
/ Credit score at the moment of application:
/ + cell_id="8583116d-be3a-4d6c-b362-5bac4b7885a5" deepnote_cell_type="code" deepnote_output_heights=[286] deepnote_to_be_reexecuted=false execution_millis=320 execution_start=1644022593745 source_hash="2d2c570b" tags=[]
fig, ax = plt.subplots()
ax = sns.boxplot(data=df, x='Score')
plt.title('Boxplot of credit score at application moment')
plt.show()
/ + [markdown] cell_id="5924c0b2-639f-4e61-b7fd-73a79e19b75d" deepnote_cell_type="markdown" tags=[]
/ Transaction balance:
/ + cell_id="991041db-2895-4d4d-988e-4365b26f22b2" deepnote_cell_type="code" deepnote_output_heights=[340] deepnote_to_be_reexecuted=false execution_millis=349 execution_start=1644022856168 source_hash="28ec5522" tags=[]
fig, ax = plt.subplots(figsize=(7, 5))
ax = sns.boxplot(data=df, x='Value')
plt.title('Boxplot of transaction balance')
plt.show()
/ + [markdown] cell_id="00efb0d9-a52d-4553-8907-24a8bb055f4b" deepnote_cell_type="markdown" tags=[]
/ Eligible:
/ + cell_id="e3df0c72-074e-4f89-8ebb-4ceaa64593b2" deepnote_cell_type="code" deepnote_output_heights=[60] deepnote_to_be_reexecuted=false execution_millis=2 execution_start=1644075975549 source_hash="d4ce5bce" tags=[]
df['eligible'].value_counts()
/ + cell_id="efe6bc7e-de9c-4af2-957e-5ae7b863566c" deepnote_cell_type="code" deepnote_output_heights=[286] deepnote_to_be_reexecuted=false execution_millis=504 execution_start=1644075620992 source_hash="24e24a28" tags=[]
fig, ax = plt.subplots()
ax = df['eligible'].value_counts().plot.bar(rot=0)
plt.title('Eligible vs non-eligible clients')
plt.xlabel('0=False, 1=True')
plt.ylabel('Number of clients')
plt.show()
/ + [markdown] cell_id="9f514a68-a18f-4306-8c85-bee999a4beeb" deepnote_cell_type="markdown" tags=[]
/ Age of clients when they applied:
/ + cell_id="5140a028-1c1a-4ccd-a2d1-6465ddaac523" deepnote_cell_type="code" deepnote_output_heights=[286] deepnote_to_be_reexecuted=false execution_millis=313 execution_start=1644024050293 source_hash="fad608b9" tags=[]
fig, ax = plt.subplots()
ax = sns.boxplot(data=df, x='client_age')
plt.title('Boxplot of clients\' age when they applied')
plt.show()
/ + [markdown] cell_id="f3c64fbf-60c0-4d6b-8e52-1b735866f0ec" deepnote_cell_type="markdown" tags=[]
/ ## Impressions so far
/
/ Categorical variables:
/
/ * Locations are evenly distributed among France, Germany and Spain.
/ * There a bit more male clients, but the difference is very small. Same for clients who have a credit card and active members.
/ * There's no special preference for any product. All of them have almost the same number of sales.
/ * The number of eligible clients (those who have been in relationship with the company for at least 2 years) is smallaller than non-eligible ones (about 20% of total clients).
/
/ Numeric variables:
/
/ * It's more likely to find salaries from 0 to 250000. There're some outliers. Those may be high net worth people.
/ * Credit scores are more likely to be between 400 and 900. There're outliers at both extremes.
/ * The distribution of transactions is quite similar to salaries distribution.
/ * All clients are adults. They tend to be young adults. At least 50% of them are millennials. There some boomers as well. Outliers represent senior people (over 68 years old).
/ * Distributions don't follow a normal pattern.
/
/ + [markdown] created_in_deepnote_cell=true deepnote_cell_type="markdown" tags=[]
/ <a style='text-decoration:none;line-height:16px;display:flex;color:#5B5B62;padding:10px;justify-content:end;' href='https://deepnote.com?utm_source=created-in-deepnote-cell&projectId=fb2ec55e-ada8-4de4-93de-2d05f236c13b' target="_blank">
/ <img alt='Created in deepnote.com' style='display:inline;max-height:16px;margin:0px;margin-right:7.5px;' src='data:image/svg+xml;base64,<KEY> > </img>
/ Created in <span style='font-weight:600;margin-left:4px;'>Deepnote</span></a>
| notebooks/0.2-axel-exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Create Classifiers to Predict HEEM labels on Texts
#
# Use this notebook to generate the classifiers needed to predict HEEM labels for new texts. You only need to do this once. The notebook saves the classifiers on you hard disk.
#
# Usage:
#
# # %run embem/machinelearning/rakel_save_clf.py <train file> <output_dir>
#
# The train file is in `<embodied emotions data directory>/ml/all_spellingnormalized.txt` (or use `<embodied emotions data directory>/ml/all.txt` to train classifiers based on non spelling normalized data.)
#
# The classifier object is saved to `<output_dir>/classifier.pkl`
#
# Training the classifiers takes a couple of minutes.
# %run embem/machinelearning/rakel_save_clf.py /home/jvdzwaan/data/embem/ml/all_spellingnormalized.txt /home/jvdzwaan/data/tmp/classifier/
| 00_CreateClassifiers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import sys
sys.path.append('/home/jlinder2/multistrand')
import nupack
import pandas as pd
import numpy as np
def get_struct(seq, temperature=25) :
return nupack.mfe([seq], T=temperature, material='rna', sodium=0.05, magnesium=0.002)[0][0]
def get_mfe(seq, temperature=25) :
return float(nupack.mfe([seq], T=temperature, material='rna', sodium=0.05, magnesium=0.002)[0][1])
# -
seq_df = pd.read_csv('array_maxcut_snv_seqs.csv', sep=',')
# +
mfes = []
mfes_w_pas = []
structs = []
structs_w_pas = []
for index, row in seq_df.iterrows() :
if index % 1000 == 0 :
print('Folding sequence ' + str(index))
seq = row['master_seq']
fold_pos = row['fold_pos']
struct = get_struct(seq[56:fold_pos])
struct_w_pas = get_struct(seq[50:fold_pos])
mfe = get_mfe(seq[56:fold_pos])
mfe_w_pas = get_mfe(seq[50:fold_pos])
mfes.append(mfe)
mfes_w_pas.append(mfe)
structs.append(struct)
structs_w_pas.append(struct_w_pas)
seq_df['mfe_w_pas'] = mfes_w_pas
seq_df['mfe'] = mfes
seq_df['struct'] = structs
seq_df['struct_w_pas'] = structs_w_pas
# -
seq_df.to_csv('array_maxcut_snv_seqs_folded.csv', sep=',', index=False)
df_var = pd.read_csv('array_cut_var_seqs.csv', sep=',')
# +
mfes_var = []
structs_var = []
mfes_ref = []
structs_ref = []
for index, row in df_var.iterrows() :
if index % 1000 == 0 :
print('Folding sequence ' + str(index))
var_seq = row['master_seq']
ref_seq = row['wt_seq']
struct_var = get_struct(var_seq[50:130].replace('T', 'U'))
struct_ref = get_struct(ref_seq[50:130].replace('T', 'U'))
mfe_var = get_mfe(var_seq[50:130].replace('T', 'U'))
mfe_ref = get_mfe(ref_seq[50:130].replace('T', 'U'))
mfes_var.append(mfe_var)
mfes_ref.append(mfe_ref)
structs_var.append(struct_var)
structs_ref.append(struct_ref)
df_var['mfe_var'] = mfes_var
df_var['mfe_wt'] = mfes_ref
df_var['struct_var'] = structs_var
df_var['struct_wt'] = structs_ref
# -
df_var['delta_mfe'] = df_var['mfe_var'] - df_var['mfe_wt']
df_var.to_csv('array_cut_var_seqs_folded_long.csv', sep=',')
| analysis/folding/fold_array_seqs_snvs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import itertools
import math
import scipy
import matplotlib.pyplot as plt
import matplotlib
import matplotlib.patches as patches
from matplotlib import animation
from matplotlib import transforms
from mpl_toolkits.axes_grid1 import make_axes_locatable
import xarray as xr
import dask
from sklearn.cluster import KMeans
from sklearn.cluster import AgglomerativeClustering
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
import pandas as pd
import netCDF4
def latent_space_analysis(Images, title, iden):
mean_image = np.mean(Images, axis=0)
var_image = np.std(Images, axis=0)
cmap="RdBu_r"
fig, ax = plt.subplots(1,2, figsize=(16,2))
cs0 = ax[0].imshow(var_image, cmap=cmap)
ax[0].set_title("Image Standard Deviation")
cs1 = ax[1].imshow(mean_image, cmap=cmap)
ax[1].set_title("Image Mean")
ax[0].set_ylim(ax[0].get_ylim()[::-1])
ax[1].set_ylim(ax[1].get_ylim()[::-1])
ax[1].set_xlabel("CRMs")
ax[0].set_xlabel("CRMs")
ax[0].set_ylabel("Pressure")
ax[1].set_yticks([])
y_ticks = np.arange(1300, 0, -300)
ax[0].set_yticklabels(y_ticks)
ax[1].set_yticklabels(y_ticks)
divider = make_axes_locatable(ax[0])
cax = divider.append_axes("right", size="5%", pad=0.05)
fig.colorbar(cs0, cax=cax)
divider = make_axes_locatable(ax[1])
cax = divider.append_axes("right", size="5%", pad=0.05)
fig.colorbar(cs1, cax=cax)
plt.suptitle(title)
#plt.savefig("/fast/gmooers/gmooers_git/CBRAIN-CAM/MAPS/model_graphs/latent_space_components/"+iden+'_'+title+'.png')
# +
Train_Images = np.load("/fast/gmooers/Preprocessed_Data/Centered_50_50/Space_Time_W_Training.npy")
Test_Images = np.load("/fast/gmooers/Preprocessed_Data/W_Variable/Trackable_Space_Time_W_Test.npy")
Max_Scalar = np.load("/fast/gmooers/Preprocessed_Data/Centered_50_50/Space_Time_Max_Scalar.npy")
Min_Scalar = np.load("/fast/gmooers/Preprocessed_Data/Centered_50_50/Space_Time_Min_Scalar.npy")
Test_Images = np.interp(Test_Images, (0, 1), (Min_Scalar, Max_Scalar))
Train_Images = np.interp(Train_Images, (0, 1), (Min_Scalar, Max_Scalar))
# -
Test_Images = np.reshape(Test_Images, (len(Test_Images),30*128))
Train_Images = np.reshape(Train_Images, (len(Train_Images),30*128))
# +
sc = StandardScaler()
z_train_std = sc.fit_transform(Train_Images)
#z_train_std = sc.fit_transform(train_log_var)
z_test_std = sc.transform(Test_Images)
#z_test_std = sc.transform(test_log_var)
# Instantiate PCA
pca = PCA(n_components=2)
pca.fit(z_train_std)
z_test_tsne = pca.transform(z_test_std)
# -
Test_Images = np.reshape(Test_Images, (len(Test_Images),30,128))
Train_Images = np.reshape(Train_Images, (len(Train_Images),30,128))
plt.scatter(x=z_test_tsne[:, 0], y=z_test_tsne[:, 1], c="#3D9AD1", s=0.1)
plt.show()
horz_line = np.squeeze(np.argwhere(np.logical_and(z_test_tsne[:,1] > -1.03, z_test_tsne[:,1] < -0.970)))
vert_line = np.squeeze(np.argwhere(np.logical_and(z_test_tsne[:,0] > -0.008, z_test_tsne[:,0] < 0.008)))
#horz_line = np.squeeze(np.argwhere(np.logical_and(z_test_tsne[:,1] > -8.005, z_test_tsne[:,1] < -7.995)))
#vert_line = np.squeeze(np.argwhere(np.logical_and(z_test_tsne[:,0] > -12.025, z_test_tsne[:,0] < -11.975)))
# +
horz_line_images = Test_Images[horz_line,:,:]
horz_line_latent = z_test_tsne[horz_line,:]
vert_line_images = Test_Images[vert_line,:,:]
vert_line_latent = z_test_tsne[vert_line,:]
horz_line_images_sorted = np.empty(horz_line_images.shape)
horz_line_latent_sorted = np.empty(horz_line_latent.shape)
vert_line_images_sorted = np.empty(vert_line_images.shape)
vert_line_latent_sorted = np.empty(vert_line_latent.shape)
# +
count = 0
for i in range(len(horz_line_images_sorted)):
ind = np.nanargmin(horz_line_latent[:,0])
horz_line_images_sorted[count,:] = horz_line_images[ind,:]
horz_line_latent_sorted[count,:] = horz_line_latent[ind,:]
horz_line_latent[ind,:] = np.array([1000.0,1000.0])
#horz_line_images[ind,:] = np.array([1000.0,1000.0])
count = count+1
count = 0
for i in range(len(vert_line_images_sorted)):
ind = np.nanargmin(vert_line_latent[:,1])
vert_line_images_sorted[count,:] = vert_line_images[ind,:]
vert_line_latent_sorted[count,:] = vert_line_latent[ind,:]
vert_line_latent[ind,:] = np.array([10000.0,10000.0])
#vert_line_image[ind,:] = np.array([1000.0,1000.0])
count = count+1
# -
print(np.where(z_test_tsne == horz_line_latent_sorted[0]))
print(np.where(z_test_tsne == horz_line_latent_sorted[-1]))
print(np.where(z_test_tsne == vert_line_latent_sorted[0]))
print(np.where(z_test_tsne == vert_line_latent_sorted[-1]))
plt.scatter(x=z_test_tsne[:, 0], y=z_test_tsne[:, 1], c="#3D9AD1", s=2.0)
plt.scatter(x=horz_line_latent_sorted[:, 0], y=horz_line_latent_sorted[:, 1], c="Red", s=2.0)
plt.scatter(x=vert_line_latent_sorted[:, 0], y=vert_line_latent_sorted[:, 1], c="Purple", s=2.0)
plt.show()
print(horz_line_latent_sorted.shape)
print(vert_line_latent_sorted.shape)
# +
path = "/DFS-L/DATA/pritchard/gmooers/Workflow/MAPS/SPCAM/100_Days/New_SPCAM5/archive/TimestepOutput_Neuralnet_SPCAM_216/atm/hist/TimestepOutput_Neuralnet_SPCAM_216.cam.h1.2009-01-20-00000.nc"
extra_variables = xr.open_dataset(path)
ha = extra_variables.hyai.values
hb = extra_variables.hybi.values
PS = 1e5
Pressures_real = PS*ha+PS*hb
fz = 15
lw = 4
siz = 100
XNNA = 1.25 # Abscissa where architecture-constrained network will be placed
XTEXT = 0.25 # Text placement
YTEXT = 0.3 # Text placement
plt.rc('text', usetex=False)
matplotlib.rcParams['mathtext.fontset'] = 'stix'
matplotlib.rcParams['font.family'] = 'STIXGeneral'
#mpl.rcParams["font.serif"] = "STIX"
plt.rc('font', family='serif', size=fz)
matplotlib.rcParams['lines.linewidth'] = lw
others = netCDF4.Dataset("/fast/gmooers/Raw_Data/extras/TimestepOutput_Neuralnet_SPCAM_216.cam.h1.2009-01-01-72000.nc")
levs = np.array(others.variables['lev'])
new = np.flip(levs)
crms = np.arange(1,129,1)
Xs, Zs = np.meshgrid(crms, new)
# -
horz_line_latent_sorted = np.flip(horz_line_latent_sorted, axis=0)
vert_line_latent_sorted = np.flip(vert_line_latent_sorted, axis=0)
horz_line_images_sorted = np.flip(horz_line_images_sorted, axis=0)
vert_line_images_sorted = np.flip(vert_line_images_sorted, axis=0)
# +
# change vx/vy to location on sorted images
def mikes_latent_animation(h_coords, v_coords, h_const, v_const, latent_space, xdist, ydist, X, Z, hline, vline, h_images, v_images):
fig, ax = plt.subplots(2,2, figsize=(36,16))
feat_list = []
#the real total you need
num_steps = len(h_coords)
#num_steps = 20
cmap= "RdBu_r"
dummy_horz = np.zeros(shape=(30,128))
dummy_horz[:,:] = np.nan
dummy_vert = np.zeros(shape=(30,128))
dummy_vert[:,:] = np.nan
count = 29
for i in range(num_steps):
for j in range(len(dummy_horz)):
dummy_horz[count,:] = h_images[i,j,:]
if i <= len(v_coords) -1:
dummy_vert[count,:] = v_images[i,j,:]
else:
dummy_vert[count,:] = v_images[-1,j,:]
count = count-1
h_rect = patches.Rectangle((h_coords[i],h_const),xdist,ydist,linewidth=4,edgecolor='black',facecolor='none')
if i <= len(v_coords) -1:
v_rect = patches.Rectangle((v_const,v_coords[i]),xdist,ydist,linewidth=4,edgecolor='black',facecolor='none')
else:
v_rect = patches.Rectangle((v_const,v_coords[-1]),xdist,ydist,linewidth=4,edgecolor='black',facecolor='none')
ax[0,0].scatter(latent_space[:, 0], latent_space[:, 1], c="#3D9AD1", s=0.4, animated=True)
ax[0,0].scatter(x=hline[:, 0], y=hline[:, 1], c="Red", s=2.0, animated=True)
cs0 = ax[0,0].add_patch(h_rect)
cs2 = ax[1,0].scatter(latent_space[:, 0], latent_space[:, 1], c="#3D9AD1", s=0.4, animated=True)
ax[1,0].scatter(x=vline[:, 0], y=vline[:, 1], c="Green", s=2.0, animated=True)
cs2 = ax[1,0].add_patch(v_rect)
cs3 = ax[1,1].pcolor(X, Z, dummy_vert, cmap=cmap, animated=True, vmin = -1.0, vmax = 1.0)
ax[1,1].set_title("(y) Vertical Velocity", fontsize=fz*2.0)
cs1 = ax[0,1].pcolor(X, Z, dummy_horz, cmap=cmap, animated=True, vmin = -1.0, vmax = 1.0)
ax[0,1].set_title("(x) Vertical Velocity", fontsize=fz*2.0)
ax[0,1].set_xlabel("CRMs", fontsize=fz*1.5)
ax[1,1].set_xlabel("CRMs", fontsize=fz*1.5)
ax[0,1].set_ylabel("Pressure (hpa)", fontsize=fz*1.5)
ax[1,1].set_ylabel("Pressure (hpa)", fontsize=fz*1.5)
y_ticks = np.array([1000, 800, 600, 400, 200])
ax[1,1].set_yticklabels(y_ticks)
ax[0,1].set_yticklabels(y_ticks)
divider = make_axes_locatable(ax[1,1])
cax = divider.append_axes("right", size="5%", pad=0.05)
fig.colorbar(cs1, cax=cax)
divider = make_axes_locatable(ax[0,1])
cax = divider.append_axes("right", size="5%", pad=0.05)
fig.colorbar(cs1, cax=cax)
feat_list.append([cs2, cs3, cs1, cs0])
count = 29
ani = animation.ArtistAnimation(fig, feat_list, interval = 125, blit = False, repeat = True)
ani.save('/fast/gmooers/gmooers_git/CBRAIN-CAM/MAPS/Animations/Figures/31_PCA_Horz_Vert.mp4')
plt.show()
mikes_latent_animation(horz_line_latent_sorted[:,0], vert_line_latent_sorted[:,1], -8.0, -12.0, z_test_tsne, 0.2, 1, Xs, Zs, horz_line_latent_sorted, vert_line_latent_sorted, horz_line_images_sorted, vert_line_images_sorted)
| MAPS/Mooers_Logbook/Fully_Convolutional_W/PCA_Latent_Space_Animation_No_Average_Fully_Conv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# %reload_ext autoreload
import web
train_data, train_label, validation_data, validation_label, test_data, test_label = web.data_preparation_moe()
trained_model=web.main1()
predicted_indexes_moe=trained_model.predict(test_data)
predicted_indexes_moe = predicted_indexes_moe[0]
predicted_indexes_moe=predicted_indexes_moe.squeeze()
import pandas as pd
predicted_indexes_moe_df = pd.DataFrame({'predicted_index':predicted_indexes_moe,'actual_index':test_label[0]})
error_in_index_moe=predicted_indexes_moe-test_label[0]
# +
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(20,10))
ax = fig.add_subplot(411)
ax.plot( predicted_indexes_moe_df.actual_index, predicted_indexes_moe_df.predicted_index,'r.');
ax = fig.add_subplot(412)
ax.plot(train_label[0] , train_data)
ax = fig.add_subplot(413)
ax.plot(predicted_indexes_moe_df.predicted_index , train_data, 'b.')
ax = fig.add_subplot(414)
ax.plot(train_label[0], error_in_index_moe)
# -
| MOE_vs_BTREE_large.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:new]
# language: python
# name: conda-env-new-py
# ---
# # Final Processing
# The final processing stage requires:
# 1. Quick validation of catalogues and Bayesian Pvalue maps
# 2. Skewness level
# 3. Adding flag to catalogue
import seaborn as sns
from astropy.table import Table
# %matplotlib inline
import numpy as np
import pylab as plt
# ## Read tables
cat=Table.read('../data/Lockman-SWIRE/MIPS/dmu26_XID+MIPS_Lockman-SWIRE_cat.fits')
cat[0:10]
from astropy import units as u
u.degree
cat['RA'].unit=u.degree
cat['Dec'].unit=u.degree
cat['F_MIPS_24'].unit=u.uJy
cat['FErr_MIPS_24_l'].unit=u.uJy
cat['FErr_MIPS_24_u'].unit=u.uJy
# ## Look at Symmetry of PDFs to determine depth level of catalogue
skew=(cat['FErr_MIPS_24_u']-cat['F_MIPS_24'])/(cat['F_MIPS_24']-cat['FErr_MIPS_24_l'])
skew.name='(84th-50th)/(50th-16th) percentile'
#g=sns.jointplot(x=np.log10(cat['F_MIPS_24']),y=skew,ylim=(0.1,10))
g=sns.jointplot(x=np.log10(cat['F_MIPS_24'][0:5000]),y=skew[0:5000],ylim=(0.1,10))
# Both seem to have flux pdfs that become Gaussian at ~20$\mathrm{\mu Jy}$
# ## Add flag to catalogue
from astropy.table import Column
cat.add_column(Column(np.zeros(len(cat), dtype=bool),name='flag_mips_24'))
ind=(cat['Pval_res_24']>0.5) | (cat['F_MIPS_24'] < 20.0)
cat['flag_mips_24'][ind]=True
cat.write('../data/Lockman-SWIRE/MIPS/dmu26_XID+MIPS_Lockman-SWIRE_cat_20171214.fits', format='fits',overwrite=True)
| dmu26/dmu26_XID+MIPS_Lockman-SWIRE/XID+MIPS_Lockman-SWIRE_final_processing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="HxDJn2pEdms5"
#
#
# IMPORTING LIBRARIES
# + id="i0AmEo_KJ5Bk"
#for data analysis purposes use import libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
plt.style.use('fivethirtyeight')
# + id="qNYGFGW9J9xI"
#Needed for Linear Regression, Polinomial Regression, Ridge Regression, and Lasso Regression
from sklearn.preprocessing import PolynomialFeatures
from sklearn.pipeline import make_pipeline
from sklearn.linear_model import LinearRegression, Ridge, Lasso, Lars
from sklearn.metrics import mean_squared_error, mean_absolute_error
from math import sqrt
# + id="3gAYCnTaJ_sA" colab={"base_uri": "https://localhost:8080/"} outputId="9101e787-5914-40dd-95b2-f78f4d6c12c3"
#Mount Google Drive
from google.colab import drive
drive.mount('/content/drive')
# + id="Q5Orh78iKHOT"
#Link newFlight_Regg.csv to newFlights_Regg
newFlights_Regg = pd.read_csv('/content/drive/My Drive/BigDataProject_Group_1/BiggerAirline.csv')
# + id="ygXXGLLfZp-3" colab={"base_uri": "https://localhost:8080/"} outputId="f495e906-7c81-49ac-8dc2-9a3b51d11b8e"
# counts the number of rows that have 0s and 1s in DEP_DELAY
newFlights_Regg.DEP_DELAY.value_counts()
# + [markdown] id="lzkeEHXEdaji"
# FUNCTION TO MAKE CONFUSION MATRIX
# Function Retrived From : https://www.kaggle.com/agungor2/various-confusion-matrix-plots
# + id="-HBmL9njYtfq"
from sklearn.metrics import confusion_matrix
def plot_cm(y_true, y_pred, figsize=(10,10)):
cm = confusion_matrix(y_true, y_pred, labels=np.unique(y_true))
cm_sum = np.sum(cm, axis=1, keepdims=True)
cm_perc = cm / cm_sum.astype(float) * 100
annot = np.empty_like(cm).astype(str)
nrows, ncols = cm.shape
for i in range(nrows):
for j in range(ncols):
c = cm[i, j]
p = cm_perc[i, j]
if i == j:
s = cm_sum[i]
annot[i, j] = '%.1f%%\n%d/%d' % (p, c, s)
elif c == 0:
annot[i, j] = ''
else:
annot[i, j] = '%.1f%%\n%d' % (p, c)
cm = pd.DataFrame(cm, index=np.unique(y_true), columns=np.unique(y_true))
cm.index.name = 'Actual'
cm.columns.name = 'Predicted'
fig, ax = plt.subplots(figsize=figsize)
sns.heatmap(cm, cmap= "YlGnBu", annot=annot, fmt='', ax=ax)
# + id="DBL5meXRKc17" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="9eacb352-d976-41d4-86fb-ac7ad5874a49"
#head() displayes the first 5 rows of the dataset
newFlights_Regg.head()
# + id="LC8pQLZSKbyc"
#Trying with more features, not just 'DISTANCE'
features = ['SCHEDULED_ELAPSED_TIME', 'ACTUAL_ELAPSED_TIME', 'ARR_DELAY', 'DISTANCE','TAXI_OUT','TAXI_IN']
# + id="i846zGmmvsGT"
# split the data into train and test
# using random seed of 3333
np.random.seed(3333)
train = newFlights_Regg.sample(frac=0.8, random_state=3333)
test = pd.concat([newFlights_Regg, train]).drop_duplicates(keep=False)
# + [markdown] id="l5-3GT3yVvCi"
# LINEAR REGRESSION
# + id="nKT6xV6Uv0Gg" colab={"base_uri": "https://localhost:8080/"} outputId="d44dddc1-5ed0-4c09-93af-fc7819096481"
linearRegression = LinearRegression(normalize=True).fit(train[features], train[['DEP_DELAY']])
print("Training Score: ", linearRegression.score(train[features], train[['DEP_DELAY']]))
print("Testing Score: ", linearRegression.score(test[features], test[['DEP_DELAY']]))
print("Mean Squared Error: ", mean_squared_error(test['DEP_DELAY'], linearRegression.predict(test[features])))
print("Root Mean Squared Error: ", sqrt(mean_squared_error(test['DEP_DELAY'], linearRegression.predict(test[features]))))
print("Mean Absolute Error: ", mean_absolute_error(test['DEP_DELAY'], linearRegression.predict(test[features])))
# + [markdown] id="wpmsvUKEV41L"
# RIDGE REGRESSION
# + id="gOL9qKe-v7Tj" colab={"base_uri": "https://localhost:8080/"} outputId="12627f8a-fea5-4e3c-cbcd-5df632d90c2b"
ridgeRegression = Ridge(normalize=True).fit(train[features], train[['DEP_DELAY']])
print("Training Score: ", ridgeRegression.score(train[features], train[['DEP_DELAY']]))
print("Testing Score: ", ridgeRegression.score(test[features], test[['DEP_DELAY']]))
print("Mean Squared Error: ", mean_squared_error(test['DEP_DELAY'], ridgeRegression.predict(test[features])))
print("Root Mean Squared Error: ", sqrt(mean_squared_error(test['DEP_DELAY'], ridgeRegression.predict(test[features]))))
print("Mean Absolute Error: ", mean_absolute_error(test['DEP_DELAY'], ridgeRegression.predict(test[features])))
# + [markdown] id="x_EJrjNNV6az"
# LASSO REGRESSION
# + id="PLlR267Kv9FA" colab={"base_uri": "https://localhost:8080/"} outputId="19c783c7-b2c0-48ba-ccb4-352a4e6f1a0f"
lassoRegression = Lasso(normalize=True).fit(train[features], train[['DEP_DELAY']])
print("Training Score: ", lassoRegression.score(train[features], train[['DEP_DELAY']]))
print("Testing Score: ", lassoRegression.score(test[features], test[['DEP_DELAY']]))
print("Mean Squared Error: ", mean_squared_error(test['DEP_DELAY'], lassoRegression.predict(test[features])))
print("Root Mean Squared Error: ", sqrt(mean_squared_error(test['DEP_DELAY'], lassoRegression.predict(test[features]))))
print("Mean Absolute Error: ", mean_absolute_error(test['DEP_DELAY'], lassoRegression.predict(test[features])))
# + id="LH5E4Ku3m-vm"
#Since No of Delayed attribues and No of Non-Delayed attributes are not equal we are going to split the dataset into two and then make the no. of Non-Delayed attributes same as the Delayed
DelayedFlights=newFlights_Regg.loc[newFlights_Regg['DEP_DELAY']==1]
Non_DelayedFlights=newFlights_Regg.loc[newFlights_Regg['DEP_DELAY']==0]
# + id="_se1OS1UnXaX" colab={"base_uri": "https://localhost:8080/"} outputId="29e01ef2-512c-4333-8685-1af5e2428a45"
#len() shows no. of rows in the dataset
len(DelayedFlights)
# + id="4WXBMxNBoEXm"
#Re-adjusted the Non_DelayedFlights so that it has almost equal no. of rows as DelayedFlights
Non_DelayedFlights=Non_DelayedFlights.sample(frac=1)
Non_DelayedFlights =Non_DelayedFlights.iloc[:4500000]
# + id="Zs6QWWCLoYUZ"
DelayedFlights=DelayedFlights.sample(frac=1)
DelayedFlights= DelayedFlights.iloc[:1500000]
# + id="heFZ6U7VoaKx"
#Joining the previously seperated datasets
frames=[DelayedFlights,Non_DelayedFlights]
DF=pd.concat(frames)
# + id="MKHcYKUmoyrU" colab={"base_uri": "https://localhost:8080/"} outputId="e642de72-eb81-41cc-d010-dbb2cbba52b9"
#len() shows no. of rows in the dataset
len(DF)
# + [markdown] id="HZyE4eixdzKm"
# SPLITTING THE DATASET FOR LOGISTIC REGRESSION PURPOSE
# + id="Q-luTVq7MLHs"
# split the data into train and test
# using random seed of 3333
np.random.seed(3333)
train = DF.sample(frac=0.8, random_state=3333)
test = pd.concat([DF, train]).drop_duplicates(keep=False)
# + [markdown] id="JKhgBEDId11r"
# LOGISTIC REGRESSION
# + id="NBvytSDNdDgX"
#Import libraries for logistic regression, cross validation, kfold
from sklearn.model_selection import StratifiedKFold, cross_val_score
from sklearn.linear_model import LogisticRegressionCV
from sklearn.metrics import classification_report
# + id="lFq-U3BedHO2"
#to avoid overfitting we are using stratifiedKFold
kfold = StratifiedKFold(n_splits=3)
# + id="DkdCaNSjdJtX" colab={"base_uri": "https://localhost:8080/"} outputId="16468fc5-15e0-40d0-a60a-8ffbaeff3a6f"
#Logistic Regression Model with target variable DEP_DELAY and other attributes are used to train it
lregg_model = LogisticRegressionCV(cv=kfold,class_weight='balanced',random_state=888)
lregg_model.fit(train[features], train[['DEP_DELAY']])
# + id="TU93Al-cdPl-"
#predicting on test data
predict_lregg=lregg_model.predict(test[features])
# + id="Jd30mAghdRcA" colab={"base_uri": "https://localhost:8080/"} outputId="5f27f27d-cbfd-4946-904b-72905d41ba90"
#printing classification report
print(classification_report(test['DEP_DELAY'], predict_lregg))
# + id="tBin_Mc6dTc0" colab={"base_uri": "https://localhost:8080/"} outputId="c9df90e9-2c63-4cbe-ea4c-6deb8f425d4d"
#calculating accuracy on cross validation
accuracy_lregg=cross_val_score(lregg_model,test[features],test['DEP_DELAY'],cv=kfold,scoring='accuracy')
# + id="IvKPP6c2dVQX" colab={"base_uri": "https://localhost:8080/"} outputId="13b498e5-75aa-46bd-bc2c-0e27fe75d4b0"
print('Mean Accuracy: %s' % accuracy_lregg.mean())
# + id="u8t3xfBcdXhE" colab={"base_uri": "https://localhost:8080/", "height": 605} outputId="fe949202-2f54-4867-f28a-14974702601d"
#making confusion matrix
plot_cm(test["DEP_DELAY"],predict_lregg)
# + [markdown] id="ps6gPfvId74s"
# SVM CLASSIFYER
# + id="tXla_NrkKndT"
#now we are going to use SVC classifyer for DEP_DELAY and metrics from sklearn
from sklearn.svm import SVC
from sklearn import metrics
# + id="lOJESkaDK1WQ"
#making classifyer with hyperparameter of max_iter=100
clsfyer = SVC(kernel='rbf',max_iter=100)
# + id="lmbAeQ5XK3vA" colab={"base_uri": "https://localhost:8080/"} outputId="aaee5714-7939-498b-b6dd-b627084968c5"
#fitting the data
clsfyer.fit(train[features],train["DEP_DELAY"])
# + id="pFtUHHTRK5W3"
# predicting the test set results
clsfyer_predictoer=clsfyer.predict(test[features])
# + id="bqG82Lg7ZM21" colab={"base_uri": "https://localhost:8080/"} outputId="69c10f19-9022-4ac7-d1b9-5cbc46711172"
#printing the accuracy
print("Accuracy:", metrics.accuracy_score(test["DEP_DELAY"], clsfyer_predictoer))
# + id="2Wki7IznYxgh" colab={"base_uri": "https://localhost:8080/", "height": 605} outputId="6450369c-1b34-49cb-8fa2-38ee3dfe25e9"
#CONFUSION MATRIX
plot_cm(test["DEP_DELAY"],clsfyer_predictoer)
# + [markdown] id="Dgdd2lZJeCED"
# RANDOM FOREST CLASSIFIER
# + id="2wkc8o-PK8Zm"
#Import library for Random forest Classifier.
from sklearn.ensemble import RandomForestClassifier
from sklearn import metrics
# + id="3LWXZYCtK9yI"
# SCALING THE INPUT
from sklearn.preprocessing import StandardScaler
sc_X = StandardScaler()
train[features] = sc_X.fit_transform(train[features])
test[features] = sc_X.transform(test[features])
# + id="K8_Ouak3K_UL"
# making the Random Forest Classifier
RFC=RandomForestClassifier()
# + id="X1CRGOTKLAyk" colab={"base_uri": "https://localhost:8080/"} outputId="2656029f-bf0c-4bed-9630-9f3c375797e0"
#Fitting the data
RFC.fit(train[features],train["DEP_DELAY"])
# + id="TdSmOxR-LCYV"
# predicting the test set results
rfc_predictoer=RFC.predict(test[features])
# + id="ibNmQabRLESb" colab={"base_uri": "https://localhost:8080/"} outputId="b98796ac-86c6-41db-fb4d-d175ca7df3cf"
#printing the accuracy
print("Accuracy:",metrics.accuracy_score(test["DEP_DELAY"], rfc_predictoer))
# + id="URIbId62X0H0" colab={"base_uri": "https://localhost:8080/", "height": 605} outputId="a53f2ecb-c89d-4704-f1fe-55f7e02609ca"
#plotting Confusion Matrix
plot_cm(test['DEP_DELAY'],rfc_predictoer)
# + colab={"base_uri": "https://localhost:8080/"} id="cg2kx5iZhj_p" outputId="f0e5dd77-4b68-42ee-a2d9-93f3bc3c9eb8"
from sklearn.model_selection import RandomizedSearchCV
# Number of trees in random forest
n_estimators = [int(x) for x in np.linspace(start = 200, stop = 2000, num = 10)]
# Number of features to consider at every split
max_features = ['auto', 'sqrt']
# Maximum number of levels in tree
max_depth = [int(x) for x in np.linspace(10, 110, num = 11)]
max_depth.append(None)
# Minimum number of samples required to split a node
min_samples_split = [2, 5, 10]
# Minimum number of samples required at each leaf node
min_samples_leaf = [1, 2, 4]
# Method of selecting samples for training each tree
bootstrap = [True, False]
# Create the random grid
random_grid = {'n_estimators': n_estimators,
'max_features': max_features,
'max_depth': max_depth,
'min_samples_split': min_samples_split,
'min_samples_leaf': min_samples_leaf,
'bootstrap': bootstrap}
print(random_grid)
# + colab={"base_uri": "https://localhost:8080/"} id="GbbSfQdVhoDq" outputId="99daa979-82c7-4920-e245-b45e0bc6ef05"
# Use the random grid to search for best hyperparameters
# First create the base model to tune
rf = RandomForestClassifier()
# Random search of parameters, using 3 fold cross validation,
# search across 100 different combinations, and use all available cores
rf_random = RandomizedSearchCV(estimator = rf, param_distributions = random_grid, n_iter = 100, cv = 3, verbose=2, random_state=42, n_jobs = -1)
# Fit the random search model
rf_random.fit(train[features],train["DEP_DELAY"])
# + id="V1nfsGtuhydp" colab={"base_uri": "https://localhost:8080/"} outputId="e75889e6-0f4e-466f-a695-28cf592be582"
rf_random.best_params_
# + id="gWJD0FejgJcq"
from sklearn.model_selection import GridSearchCV
# Create the parameter grid based on the results of random search
param_grid = {
'bootstrap': [True],
'max_depth': [10, 20, 30, 40],
'max_features': [2, 3],
'min_samples_leaf': [3, 4, 5],
'min_samples_split': [4, 8, 10],
'n_estimators': [100, 200, 300, 1000]
}
# Create a based model
rf = RandomForestClassifier()
# Instantiate the grid search model
grid_search = GridSearchCV(estimator = rf, param_grid = param_grid,
cv = 3, n_jobs = -1, verbose = 2)
# + id="Ov_S45DOgXXC" colab={"base_uri": "https://localhost:8080/"} outputId="650a77bc-9dd8-4252-c1a1-bcb97fd47a30"
# Fit the grid search to the data
grid_search.fit(train[features],train["DEP_DELAY"])
grid_search.best_params_
# + id="OazLolX5c2i5"
RFC_1=RandomForestClassifier(bootstrap=True,
max_depth = 10,
max_features= 2,
min_samples_leaf= 4,
min_samples_split= 2,
n_estimators= 400)
# + id="X4sd8B6tdtbf"
RFC_1.fit(train[features],train["DEP_DELAY"])
# + colab={"base_uri": "https://localhost:8080/"} id="HqZQIaYoeC-Z" outputId="f287eb6f-7d90-49a3-fafe-7b0d5d292fdd"
print("Accuracy:",metrics.accuracy_score(test["DEP_DELAY"], RFC_1.predict(test[features])))
# + [markdown] id="37La5FoVoo9s"
# RANDOME FOREST REGRESSOR GOT 86%
# + id="aizsOL7TOhu4"
#plotting Confusion Matrix
plot_cm(test['DEP_DELAY'], RFC_1.predict(test[features]))
| PythonNotebooks/finalPredictionModels.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="9RxprXwt49SA"
import numpy as np
import matplotlib.pyplot as plt
import torch
from torch.autograd import Function
from torchvision import datasets, transforms
import torch.optim as optim
import torch.nn as nn
import torch.nn.functional as F
from torchsummary import summary
import qiskit
from qiskit.visualization import *
# + colab={"base_uri": "https://localhost:8080/"} id="YQgHj0PU56WP" outputId="b816663a-5b0d-4b28-ad13-1a6537b04326"
use_cuda = torch.cuda.is_available()
print('CUDA available:', use_cuda)
if use_cuda:
device = torch.device('cuda')
print('Training on GPU...')
else:
device = torch.device('cpu')
print('Training on CPU...')
# + id="qqZ4Bf7m6RoQ"
class QuantumCircuit:
"""
This class provides a simple interface for interaction
with the quantum circuit
"""
def __init__(self, n_qubits, backend, shots):
# --- Circuit definition ---
self._circuit = qiskit.QuantumCircuit(n_qubits)
all_qubits = [i for i in range(n_qubits)]
self.theta = qiskit.circuit.Parameter('theta')
self._circuit.h(all_qubits)
self._circuit.barrier()
self._circuit.ry(self.theta, all_qubits)
self._circuit.measure_all()
# ---------------------------
self.backend = backend
self.shots = shots
def run(self, thetas):
t_qc = transpile(self._circuit,
self.backend)
qobj = assemble(t_qc,
shots=self.shots,
parameter_binds = [{self.theta: theta} for theta in thetas])
job = self.backend.run(qobj)
result = job.result().get_counts()
counts = np.array(list(result.values()))
states = np.array(list(result.keys())).astype(float)
# Compute probabilities for each state
probabilities = counts / self.shots
# Get state expectation
expectation = np.sum(states * probabilities)
return np.array([expectation])
# + id="KAVZf72s_7l7"
import numpy as np
import matplotlib.pyplot as plt
import torch
from torch.autograd import Function
from torchvision import datasets, transforms
import torch.optim as optim
import torch.nn as nn
import torch.nn.functional as F
import qiskit
from qiskit import transpile, assemble
from qiskit.visualization import *
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="E_IJIC0V6jQ3" outputId="dbca9f70-72a7-41c1-e180-4b14c1b56523"
simulator = qiskit.Aer.get_backend('aer_simulator')
circuit = QuantumCircuit(1, simulator, 100)
print('Expected value for rotation pi {}'.format(circuit.run([np.pi])[0]))
circuit._circuit.draw()
# + id="wrq0Z2Wy6lf2"
class HybridFunction(Function):
""" Hybrid quantum - classical function definition """
@staticmethod
def forward(ctx, inputs, quantum_circuit, shift):
""" Forward pass computation """
ctx.shift = shift
ctx.quantum_circuit = quantum_circuit
expectation_z = []
for input in inputs:
expectation_z.append(ctx.quantum_circuit.run(input.tolist()))
result = torch.tensor(expectation_z).cuda()
ctx.save_for_backward(inputs, result)
return result
@staticmethod
def backward(ctx, grad_output):
""" Backward pass computation """
input, expectation_z = ctx.saved_tensors
input_list = np.array(input.tolist())
shift_right = input_list + np.ones(input_list.shape) * ctx.shift
shift_left = input_list - np.ones(input_list.shape) * ctx.shift
gradients = []
for i in range(len(input_list)):
expectation_right = ctx.quantum_circuit.run(shift_right[i])
expectation_left = ctx.quantum_circuit.run(shift_left[i])
gradient = torch.tensor([expectation_right]).cuda() - torch.tensor([expectation_left]).cuda()
gradients.append(gradient)
# gradients = np.array([gradients]).T
gradients = torch.tensor([gradients]).cuda()
gradients = torch.transpose(gradients, 0, 1)
# return torch.tensor([gradients]).float() * grad_output.float(), None, None
return gradients.float() * grad_output.float(), None, None
class Hybrid(nn.Module):
""" Hybrid quantum - classical layer definition """
def __init__(self, backend, shots, shift):
super(Hybrid, self).__init__()
self.quantum_circuit = QuantumCircuit(1, backend, shots)
self.shift = shift
def forward(self, input):
return HybridFunction.apply(input, self.quantum_circuit, self.shift)
# + id="c8nsTM-D7T2u"
n_samples = 512
batch_size = 256
X_train = datasets.MNIST(root='./data', train=True, download=True,
transform=transforms.Compose([transforms.ToTensor()]))
X_train.data = X_train.data[:n_samples]
X_train.targets = X_train.targets[:n_samples]
train_loader = torch.utils.data.DataLoader(X_train, batch_size=batch_size, shuffle=True)
# check MNIST data
# + colab={"base_uri": "https://localhost:8080/", "height": 130} id="8L2UsYYc7Vdq" outputId="8b34cc87-94dd-4a14-a2c8-5f6907701d76"
n_samples_show = 6
data_iter = iter(train_loader)
fig, axes = plt.subplots(nrows=1, ncols=n_samples_show, figsize=(10, 3))
images, targets = data_iter.__next__()
while n_samples_show > 0:
axes[n_samples_show - 1].imshow(images[n_samples_show].numpy().squeeze(), cmap='gray')
axes[n_samples_show - 1].set_xticks([])
axes[n_samples_show - 1].set_yticks([])
axes[n_samples_show - 1].set_title("Labeled: {}".format(int(targets[n_samples_show])))
n_samples_show -= 1
# + id="Hfyjc5Kf7W7q"
n_samples = 2048
X_test = datasets.MNIST(root='./data', train=False, download=True,
transform=transforms.Compose([transforms.ToTensor()]))
# idx = np.append(np.where(X_test.targets == 0)[0][:n_samples],
# np.where(X_test.targets == 1)[0][:n_samples])
X_test.data = X_test.data[:n_samples]
X_test.targets = X_test.targets[:n_samples]
test_loader = torch.utils.data.DataLoader(X_test, batch_size=batch_size, shuffle=True)
# + id="rMutExyM7ZXX"
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(1, 6, kernel_size=5)
self.conv2 = nn.Conv2d(6, 16, kernel_size=5)
self.dropout = nn.Dropout2d()
self.fc1 = nn.Linear(256, 64)
self.fc2 = nn.Linear(64, 10)
# self.hybrid = Hybrid(qiskit.Aer.get_backend('qasm_simulator'), 100, np.pi / 2)
self.hybrid = [Hybrid(qiskit.Aer.get_backend('qasm_simulator'), 100, np.pi / 2) for i in range(10)]
def forward(self, x):
x = F.relu(self.conv1(x))
x = F.max_pool2d(x, 2)
x = F.relu(self.conv2(x))
x = F.max_pool2d(x, 2)
x = self.dropout(x)
# x = x.view(-1, 256)
x = torch.flatten(x, start_dim=1)
x = F.relu(self.fc1(x))
x = self.fc2(x)
x = torch.chunk(x, 10, dim=1)
# x = self.hybrid(x)
x = tuple([hy(x_) for hy, x_ in zip(self.hybrid, x)])
return torch.cat(x, -1)
# + colab={"base_uri": "https://localhost:8080/"} id="DXFVRRZv7bBf" outputId="c2791d25-93da-405d-dfa4-f57bd5009fdd"
model = Net().cuda()
summary(model, (1, 28, 28))
# + colab={"base_uri": "https://localhost:8080/"} id="MhuFWDus7cae" outputId="9e1731ea-784c-44e8-c424-2a1acfa43ded"
optimizer = optim.Adam(model.parameters(), lr=0.001)
# loss_func = nn.NLLLoss()
loss_func = nn.CrossEntropyLoss().cuda()
epochs = 50
loss_list = []
model.train()
for epoch in range(epochs):
total_loss = []
for batch_idx, (data, target) in enumerate(train_loader):
optimizer.zero_grad()
data = data.cuda()
target = target.cuda()
# Forward pass
output = model(data).cuda()
# print("data ", data.size())
# print("output", output.size())
# print("target", target.size())
# Calculating loss
loss = loss_func(output, target)
# Backward pass
loss.backward()
# Optimize the weights
optimizer.step()
total_loss.append(loss.item())
loss_list.append(sum(total_loss)/len(total_loss))
print('Training [{:.0f}%]\tLoss: {:.4f}'.format(
100. * (epoch + 1) / epochs, loss_list[-1]))
# + colab={"base_uri": "https://localhost:8080/", "height": 312} id="5_ZPeOVl7eSH" outputId="9dcfbf34-9ea2-4091-cf02-c4489a478349"
plt.plot(loss_list)
plt.title('Hybrid NN Training Convergence')
plt.xlabel('Training Iterations')
plt.ylabel('Neg Log Likelihood Loss')
# + colab={"base_uri": "https://localhost:8080/"} id="XaGWji5s7gHF" outputId="ca49a13f-36df-4cf7-e9f8-4f14463c5c78"
model.eval()
with torch.no_grad():
correct = 0
for batch_idx, (data, target) in enumerate(test_loader):
data = data.cuda()
target = target.cuda()
output = model(data).cuda()
pred = output.argmax(dim=1, keepdim=True)
correct += pred.eq(target.view_as(pred)).sum().item()
loss = loss_func(output, target)
total_loss.append(loss.item())
print('Performance on test data:\n\tLoss: {:.4f}\n\tAccuracy: {:.1f}%'.format(
sum(total_loss) / len(total_loss),
correct / len(test_loader) * 100 / batch_size)
)
# + colab={"base_uri": "https://localhost:8080/", "height": 110} id="2F0Qn3T-7hqB" outputId="95305a51-68ca-4f78-9a74-38b6b183589a"
n_samples_show = 8
count = 0
fig, axes = plt.subplots(nrows=1, ncols=n_samples_show, figsize=(10, 3))
model.eval()
with torch.no_grad():
for batch_idx, (data, target) in enumerate(test_loader):
if count == n_samples_show:
break
data_cuda = data.cuda()
target_cuda = target.cuda()
output_cuda = model(data_cuda).cuda()
pred = output_cuda.argmax(dim=1, keepdim=True)
axes[count].imshow(data[count].numpy().squeeze(), cmap='gray')
axes[count].set_xticks([])
axes[count].set_yticks([])
axes[count].set_title('Predicted {}'.format(pred[count].item()))
count += 1
| Quantum Convolutional Neural Networks/QCNN_with_Pytorch_and_Qiskit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from skimage import io, img_as_ubyte
from skimage.color import rgb2gray
import cv2
import random
def show2imgs(im1, im2, title1='Obraz pierwszy', title2='Obraz drugi', size=(10,10)):
f, (ax1, ax2) = plt.subplots(1,2, figsize=size)
ax1.imshow(im1, cmap='gray')
ax1.axis('off')
ax1.set_title(title1)
ax2.imshow(im2, cmap='gray')
ax2.axis('off')
ax2.set_title(title2)
plt.show()
def segmentuj(img):
for i in range(img.shape[0]):
for j in range(img.shape[1]):
if img[i][j] != 255:
if img[i][j] == 0:
img[i][j] = random.randint(1, 100)
if img[i-1][j-1] != 0 and img[i-1][j-1] != 255:
img[i][j] = img[i-1][j-1]
elif img[i-1][j] != 0 and img[i-1][j] != 255:
img[i][j] = img[i-1][j]
elif img[i-1][j+1] != 0 and img[i-1][j+1] != 255:
img[i][j] = img[i-1][j+1]
elif img[i][j-1] != 0 and img[i][j-1] != 255:
img[i][j] = img[i][j-1]
elif img[i][j+1] != 0 and img[i][j+1] != 255:
img[i][j] = img[i][j+1]
elif img[i+1][j-1] != 0 and img[i+1][j-1] != 255:
img[i][j] = img[i-1][j-1]
elif img[i+1][j] != 0 and img[i+1][j] != 255:
img[i][j] = img[i+1][j]
elif img[i+1][j+1] != 0 and img[i+1][j+1] != 255:
img[i][j] = img[i+1][j+1]
else:
continue
#scalanie obiektów
if img[i][j] != img[i-1][j-1] and img[i-1][j-1] != 255:
img[i][j] = img[i-1][j-1]
elif img[i][j] != img[i-1][j] and img[i-1][j] != 255:
img[i][j] = img[i-1][j]
elif img[i][j] != img[i-1][j+1] and img[i-1][j+1] != 255:
img[i-1][j+1] = img[i][j]
elif img[i][j] != img[i][j-1] and img[i][j-1] != 255:
img[i][j-1] = img[i][j]
elif img[i][j] != img[i][j+1] and img[i][j+1] != 255:
img[i][j+1] = img[i][j]
elif img[i][j] != img[i+1][j-1] and img[i+1][j-1] != 255:
img[i][j] = img[i+1][j-1]
elif img[i][j] != img[i+1][j] and img[i+1][j] != 255:
img[i][j] = img[i+1][j]
elif img[i][j] != img[i+1][j+1] and img[i+1][j+1] != 255:
img[i][j] = img[i+1][j+1]
else:
continue
return img
def scal(img):
for i in range(img.shape[0]):
for j in range(img.shape[1]):
if img[i][j] != 255:
if img[i][j] != img[i-1][j-1] and img[i-1][j-1] != 255:
img[i][j] = img[i-1][j-1]
elif img[i][j] != img[i-1][j] and img[i-1][j] != 255:
img[i][j] = img[i-1][j]
elif img[i][j] != img[i-1][j+1] and img[i-1][j+1] != 255:
img[i-1][j+1] = img[i][j]
elif img[i][j] != img[i][j-1] and img[i][j-1] != 255:
img[i][j-1] = img[i][j]
elif img[i][j] != img[i][j+1] and img[i][j+1] != 255:
img[i][j+1] = img[i][j]
elif img[i][j] != img[i+1][j-1] and img[i+1][j-1] != 255:
img[i][j] = img[i+1][j-1]
elif img[i][j] != img[i+1][j] and img[i+1][j] != 255:
img[i][j] = img[i+1][j]
elif img[i][j] != img[i+1][j+1] and img[i+1][j+1] != 255:
img[i][j] = img[i+1][j+1]
else:
continue
return img
def licz_obiekty(img):
ile_obiektow = 0
obiekty = []
for i in range(img.shape[0]):
for j in range(img.shape[1]):
if img[i][j] != 255:
obiekty.append(img[i][j])
ile_obiektow = len(list(set(obiekty)))# zlicza niepowtarzające się elementy listy
return ile_obiektow
img = io.imread('pattern1.png')
th = 110
img = img_as_ubyte(rgb2gray(img))
th, bim = cv2.threshold(img, thresh=th, maxval=255, type=cv2.THRESH_BINARY)
bimCPY = bim;
plt.imshow(bimCPY, cmap="gray")
plt.axis('on')
plt.show()
# +
#najlepiej uruchomić segmentację 2 razy gdyż przez "niewielki" zakres możliwych kolorów mogą się one powtarzać
img = segmentuj(bim)
nimg = scal(img)
nimg = scal(nimg)
plt.imshow(nimg, cmap="hot")
plt.axis('on')
plt.show()
# -
ile = licz_obiekty(nimg)
print("Obiektów jest:\t", ile)
| Zadania Domowe/zadDomowe-2018-12-11.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
from numpy import array,zeros, sin,linspace,nanmin,where
from printSoln import *
import run_kut4 as runkut
from euler0 import *
import pylab as pl
import scipy
from scipy import integrate
import time
# # Question 1
# First part is done by replacing this bit:
#
# def integrate(F,x,y,xStop,h):
#
# def run_kut4(F,x,y,h):
#
# K0 = h*F(x,y)
#
# K1 = h*F(x + h/2.0, y + K0/2.0)
#
# K2 = h*F(x + h/2.0, y + K1/2.0)
#
# K3 = h*F(x + h, y + K2)
#
# return (K0 + 2.0*K1 + 2.0*K2 + K3)/6.0
#
# With this:
#
# def integrate0(F,x,y,xStop,h):
#
# def euler(F,x,y,h):
#
# K0 = h*F(x,y)
#
# return K0
#
# It's been saved as euler0.
#
# Honestly I have no idea how this was done but it might make sense if we look at the slides
# +
#Define the function by putting into an array as so:
def fb(x,y):
f=zeros(2)
f[0]=y[1]
f[1]=-y[0]
return f
# +
#x = lower limit of the integral
#xstop = upper limit of the integral
#y = array of the initial conditions ( y(0)=0 & y'(0)=1 )
#h = step size (I dunno, it's in the slides somewhere. I think.)
#freq = I dunno but it's usually 1 so just roll with it.
x = 0.0 # Start of integration
xStop = 20.0 # End of integration
y = array([0., 1.]) # Initial values of {y,y’}
h1 = 0.1 # Step size
freq = 1 # Printout frequency
# +
#This integrates using the Euler code but pretty much the same for RK4 too. Question is solved using both.
#There's also a timer whacked on to each to compare how long each takes as a higher h values means longer time to calculate.
#Euler
start0=time.time()
X0,Y0 = integrate0(fb,x,y,xStop,h1)
#printSoln(X0,Y0,freq)
end0=time.time()
errorEuler = sin(X0)-Y0[:,0] # Error is just the true value-the value given (both are arrays of numbers for the integral)
# +
#Runge-Kutta 4th order
start1=time.time()
X1,Y1=runkut.integrate(fb,x,y,xStop,h1)
#printSoln(X1,Y1,freq)
end1=time.time()
errorRK4 = sin(X0)-Y1[:,0]
# +
#Euler again with a smaller h
h2 = 0.001 # Step size
start2=time.time()
X2,Y2 = integrate0(fb,x,y,xStop,h2)
#printSoln(X2,Y2,freq)
end2=time.time()
errorEuler2 = sin(X2)-Y2[:,0]
# -
pl.plot(X0,Y0[:,0], label='Euler')
pl.plot(X1,Y1[:,0], label='RK4')
pl.plot(X2,Y2[:,0], label='Euler2')
pl.legend(loc='best')
pl.title('Integration Results')
pl.show()
pl.title('Errors')
pl.plot(X0,errorEuler, label='Euler')
pl.plot(X0,errorRK4, label='RK4')
pl.legend(loc='best')
pl.show()
# Clearly, the errors of the Euler method with step size of h = 0.1
# are unacceptably large. In order to decrease them to acceptable
# level (almost indistinguishable from the RK4 method errors) we
# need h = 0.001 as previously calculated.
# +
print 'Method Times:'
print 'Euler h=0.1 -', end0-start0, ' RK4 h=0.1 -', end1-start1, ' Euler h=0.001 -', end2-start2
pl.title('Errors')
pl.plot(X0,errorRK4, label='RK4')
pl.plot(X2,errorEuler2, label='Euler2')
pl.legend(loc='best')
pl.show()
# -
# However, we note that albeit the results are acceptable, it takes much longer to obtain the answer.
#
# The problem observed is that the Euler method, which is low-order continuously over-shoots the correct solution.
# # Question 2
#
# Comments:
# - Please give a comprehensive discussion that states your methods, motivates your choices, and makes a conclusion. Use text boxes surrouding your code.
# - What is the effect of changing initial conditions?
# In order for python to be able to solve the differential equations,
# we must first split the second order differential equation into
# two first order differential equations.
#
# y'' = g*(1-a*(y**3))
#
# y0 = 0.1 = x1
#
# y'0 = 0.0 = x2
#
# x1' = x2 = 0
#
# x2' = y'' = g*(1-a*(x1**3))
#define the constants a and g
a= 1.
g= 9.80665
#define the given function using the two previous calculated first order differentials
def fa(x,y):
f= zeros(2)
f[0]= (g*(1.0 - (a*(x[1])**3.0)))
f[1]= x[0]
return f
#generate arrays of numbers for the range of values of time to be used and the intial conditions of the function (initials velocity and displacement).
#the subscripts determine each part of the question, 1 is for y0 = 0.1 and 2 is for y0 = 0.9.
x= linspace(0, 2.5, 500)
y1= array([0, 0.1])
y2= array([0, 0.9])
#use scipy's inbuilt odeint function to solve the differential equation for both initial conditions.
z1= scipy.integrate.odeint(fa,y1,x)
z2= scipy.integrate.odeint(fa,y2,x)
#plot these results on a graph of displacement against time.
pl.plot(x, z1[:,1], label = '0.1m',color ='red')
pl.plot(x, z2[:,1], '--', label = '0.9m', color ='blue')
pl.title('Displacement against time for the conical float')
pl.xlabel('time (s)')
pl.ylabel('Displacement (m)')
pl.legend(loc='best')
pl.show()
# It can be estimated from the graph that, for the initial condition of y0 = 0.1m, the period and ampliteude are approximately 1.35s and 0.8m respectively.
# For the initial condition of y0 = 0.9m, the period and amplitude are approximately 1.2s and 0.09m respectively.
#repeat the same steps as previously using the new function.
def fb(x,y):
f= zeros(2)
f[0]= (g*(1.0 - (a*(x[1]))))
f[1]= x[0]
return f
z3= scipy.integrate.odeint(fb,y1,x)
z4= scipy.integrate.odeint(fb,y2,x)
pl.plot(x, z3[:,1], label = '0.1m',color ='red')
pl.plot(x, z4[:,1], '--', label = '0.9m', color ='blue')
pl.title('Displacement against time for the conical float')
pl.xlabel('time (s)')
pl.ylabel('Displacement (m)')
pl.legend(loc='best')
pl.show()
# It can be estimated from the graph that, for the initial condition of y0 = 0.1m, the period and ampliteude are approximately 2.0s and 0.8m respectively.
# For the initial condition of y0 = 0.9m, the period and amplitude are approximately 2.0s and 0.1m respectively.
#1)c)
#Plot the displacement against the velocity.
pl.plot(z1[:,1],z1[:,0],label='y0=0.1 for y^3')
pl.plot(z2[:,1],z2[:,0],label='y0=0.9 for y^3')
pl.plot(z3[:,1],z3[:,0],label='y0=0.1 for y')
pl.plot(z4[:,1],z4[:,0],label='y0=0.9 for y')
pl.title('Displacement against velocity.')
pl.xlabel('Displacement.')
pl.ylabel('Velocity.')
pl.legend(loc='best')
pl.show()
# # Past Paper Question 2013-14
#define a function of the two previously derived first order equations setting them in the same array.
def f(v,t):
f0 = v[1]
f1 = (g - ((Cd/m)*(v[1]**2)))
return [f0,f1]
#Define constants
g = 9.80665
Cd = 0.2028
m = 80
#generate our values for t to integrate over and our initial values for y and y' (displacement and velocity)
t = linspace(0,100,5000)
y0 = [0,0]
#integrate the function using odepack which generates an array of values for y and v.
#the last 2 lines just define y and v as each set of values in z.
z = integrate.odeint(f,y0,t)
y = z[:,0]
v = z[:,1]
pl.plot(t,-y+5000, label='Position')
pl.title('Displacement vs. time of skydiver')
pl.xlabel('time')
pl.ylabel('Displacement from starting position')
pl.ylim(0,5000)
pl.plot()
#finds the point at which y is more than 5000m in the array and gives the minimum value in the array t it is over this number.
#t_end is just calculating the actual value of t based on the parameters of the linspace code above (0-100 over 5000 points)
t0 = nanmin(where(y>5000))
t_end = (t0/5000.)*100
print 'time taken to fall 5000m =', t_end, 'seconds'
pl.plot(t,v, label='Velocity')
pl.title('Velocity vs. time of skydiver')
pl.xlabel('time')
pl.ylabel('Velocity')
pl.xlim(0,t_end)
pl.plot()
print 'terminal velocity =', max(v), 'm/s'
| ExamPrep/08 Differential Equations/_SC 08 - Differential Equations 1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import numpy as np
import matplotlib.pyplot as plt
import sys
sys.path.append('../')
import pyabeles as pa
# %matplotlib inline
data = np.genfromtxt('Miro data.csv',delimiter=',',skip_header=64)
x = data[:,0]
y = data[:,1]
scale = y[0]
for i in range(0,len(y)):
y[i] /= scale
x[i] /= 2
plt.plot(x,y)
plt.plot(x,np.log(y))
# ## DE for Fitting
np.random.seed(12345)
struct = pa.Surface()
STO = pa.Layer(0.,5.246,sigma=0.26,name="STO")
LSMO = pa.Layer(280.,6.6, sigma=5.85, name="LSMO")
for i in range(0,1):
struct.addLayer(LSMO)
struct.addLayer(STO)
exp = struct.doExperiment(x)
R = exp.genTheory(modify=False)
exp.theory = y
f = pa.Fitter(exp,method="de",cutoff_begin=20,cutoff_end=700)
# ### Easy code to modify when which vars are fixed
f.fixed = np.full(10,True)
for i in range(1,10):
f.set_free(i)
surf = f.fit()
mhm = exp.genTheory(modify=False)
plt.plot(x[0:],np.log10(mhm[0:]))
plt.plot(x[0:],np.log10(y[0:]))
f.error(exp.get_params_list())
de_params = np.copy(exp.get_params_list())
exp.get_params_list()
# ## Nelder-Mead for fitting
np.random.seed(12345)
struct = pa.Surface()
STO = pa.Layer(0.,5.246,sigma=0.26,name="STO")
LSMO = pa.Layer(280.,6.6, sigma=5.85, name="LSMO")
for i in range(0,1):
struct.addLayer(LSMO)
struct.addLayer(STO)
exp = struct.doExperiment(x)
R = exp.genTheory(modify=False)
exp.theory = y
f = pa.Fitter(exp,method="nm",cutoff_begin=20,cutoff_end=700)
# +
f.fixed = np.full(10,True)
for i in range(1,9):
f.set_free(i)
f.fixed
# -
surf = f.fit()
mhm = exp.genTheory(modify=False)
plt.plot(x[0:],np.log10(mhm[0:]))
plt.plot(x[0:],np.log10(y[0:]))
f.error(exp.get_params_list())
nm_params = np.copy(exp.get_params_list())
print de_params
print nm_params
[(0.75, 1.25),
(0.0, 1.9999999999999999e-06),
(-0.20000000000000001, 0.20000000000000001),
(0.1, 10.0),
(210.0, 350.0),
(0.0, 0.0),
(4.9499999999999993, 8.25),
(3.9345000000000003, 6.557500000000001),
(4.3874999999999993, 7.3125),
(0.19500000000000001, 0.32500000000000001)]
| notebooks/.ipynb_checkpoints/Untitled-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"name": "#%%\n"}
import pandas as pd
import nltk
nltk.download('stopwords')
import re
import string
from nltk import word_tokenize
#nltk.download('stopwords')
#nltk.download('punkt')
stopwords = nltk.corpus.stopwords.words('english')
from nltk.stem.snowball import SnowballStemmer
stemmer = SnowballStemmer("english")
# -
# # Load Data
# + pycharm={"name": "#%%\n"}
#---------------STARTING WITH R8----------------------------
##------------BUILDING THE DATASET
X_train = pd.read_csv('datasets/r8-train-all-terms.txt', sep="\t", header=None)
X_test = pd.read_csv('datasets/r8-test-all-terms.txt', sep="\t", header=None)
data_r8=pd.concat([X_train,X_test], ignore_index=True)
data_r8.columns = ["class", "text"]
#nb of classes
classes_count = data_r8.groupby('class').count().sort_values(by=['text'],ascending=False)
# -
# # I- PreProcessing
# + pycharm={"name": "#%%\n"}
# !pip install unidecode
# + pycharm={"name": "#%%\n"}
import nltk
nltk.download('stopwords')
nltk.download('punkt')
from unidecode import unidecode
from nltk import SnowballStemmer
import re
from nltk.tokenize import word_tokenize
# + pycharm={"name": "#%%\n"}
text = data_r8.text
#df to a list
rows = text.apply(''.join).tolist()
# + pycharm={"name": "#%%\n"}
# Import the wordcloud library
from wordcloud import WordCloud
# Join the different descriptions together.
long_string = ''.join(str(v) for v in text)
# Create a WordCloud object
wordcloud = WordCloud(background_color="white", max_words=5000, contour_width=3, contour_color='steelblue')
# Generate a word cloud
wordcloud.generate(long_string)
# Visualize the word cloud
wordcloud.to_image()
# + pycharm={"name": "#%%\n"}
# Load the library with the CountVectorizer method
from sklearn.feature_extraction.text import CountVectorizer
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style('whitegrid')
# %matplotlib inline
# Helper function
def plot_10_most_common_words(count_data, count_vectorizer):
import matplotlib.pyplot as plt
words = count_vectorizer.get_feature_names()
total_counts = np.zeros(len(words))
for t in count_data:
total_counts+=t.toarray()[0]
count_dict = (zip(words, total_counts))
count_dict = sorted(count_dict, key=lambda x:x[1], reverse=True)[0:10]
words = [w[0] for w in count_dict]
counts = [w[1] for w in count_dict]
x_pos = np.arange(len(words))
plt.figure(2, figsize=(15, 15/1.6180))
plt.subplot(title='10 most common words')
sns.set_context("notebook", font_scale=1.25, rc={"lines.linewidth": 2.5})
sns.barplot(x_pos, counts, palette='husl')
plt.xticks(x_pos, words, rotation=90)
plt.xlabel('words')
plt.ylabel('counts')
plt.show()
# Initialise the count vectorizer with the English stop words
count_vectorizer = CountVectorizer(max_df=0.95,stop_words='english')
# Fit and transform the processed descriptions
count_data = count_vectorizer.fit_transform(text)
# Visualise the 10 most common words
plot_10_most_common_words(count_data, count_vectorizer)
# -
# ## 1. Stop Words and Stemming¶
# + pycharm={"name": "#%%\n"}
# removes a list of words (ie. stopwords) from a tokenized list.
def removeWords(listOfTokens, listOfWords):
return [token for token in listOfTokens if token not in listOfWords]
# applies stemming to a list of tokenized words
def applyStemming(listOfTokens, stemmer):
return [stemmer.stem(token) for token in listOfTokens]
# removes any words composed of less than 2 or more than 21 letters
def twoLetters(listOfTokens):
twoLetterWord = []
for token in listOfTokens:
if len(token) <= 2 or len(token) >= 21:
twoLetterWord.append(token)
return twoLetterWord
def processCorpus(corpus, language):
stopwords = nltk.corpus.stopwords.words(language)
param_stemmer = SnowballStemmer(language)
for document in corpus:
index = corpus.index(document)
corpus[index] = corpus[index].replace(u'\ufffd', '8') # Replaces the ASCII '�' symbol with '8'
corpus[index] = corpus[index].replace(',', '') # Removes commas
corpus[index] = corpus[index].rstrip('\n') # Removes line breaks
corpus[index] = corpus[index].casefold() # Makes all letters lowercase
corpus[index] = re.sub('\W_',' ', corpus[index]) # removes specials characters and leaves only words
corpus[index] = re.sub("\S*\d\S*"," ", corpus[index]) # removes numbers and words concatenated with numbers IE h4ck3r. Removes road names such as BR-381.
corpus[index] = re.sub("\S*@\S*\s?"," ", corpus[index]) # removes emails and mentions (words with @)
corpus[index] = re.sub(r'http\S+', '', corpus[index]) # removes URLs with http
corpus[index] = re.sub(r'www\S+', '', corpus[index]) # removes URLs with www
listOfTokens = word_tokenize(corpus[index])
twoLetterWord = twoLetters(listOfTokens)
listOfTokens = removeWords(listOfTokens, stopwords)
listOfTokens = removeWords(listOfTokens, twoLetterWord)
listOfTokens = applyStemming(listOfTokens, param_stemmer)
corpus[index] = " ".join(listOfTokens)
corpus[index] = unidecode(corpus[index])
return corpus
# + pycharm={"name": "#%%\n"}
language = 'english'
rows_clean = processCorpus(rows, language)
rows_clean[13][0:460]
| exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JimKing100/nfl-test/blob/master/predictions-final/Offense_Predictions.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="E_J5wZaQFoPy" colab_type="code" colab={}
# Installs
# %%capture
# !pip install pmdarima
# !pip install category_encoders==2.0.0
# + id="DffZtLAbFvcW" colab_type="code" outputId="07ff05ec-c7dc-4fc3-86ce-d9d6343d83a3" colab={"base_uri": "https://localhost:8080/", "height": 71}
# Imports
import numpy as np
import pandas as pd
from statsmodels.tsa.arima_model import ARIMA
import pmdarima as pm
from sklearn import preprocessing
import category_encoders as ce
# + id="D-gz9-reFyBZ" colab_type="code" colab={}
# Import data
players_df = pd.read_csv('https://raw.githubusercontent.com/JimKing100/nfl-test/master/data-actuals/players_full.csv')
original_df = pd.read_csv('https://raw.githubusercontent.com/JimKing100/nfl-test/master/data-actuals/actuals_offense.csv')
kickers_df = pd.read_csv('https://raw.githubusercontent.com/JimKing100/nfl-test/master/data-revised/rookies_non_kicker.csv')
offense_df = pd.read_csv('https://raw.githubusercontent.com/JimKing100/nfl-test/master/data-revised/rookies_non_offense.csv')
bye_df = pd.read_csv('https://raw.githubusercontent.com/JimKing100/nfl-test/master/data-revised/bye.csv')
injury_df = pd.read_csv('https://raw.githubusercontent.com/JimKing100/nfl-test/master/data-revised/injury.csv')
player_df = pd.concat([kickers_df, offense_df], ignore_index=True)
# + id="m78bxZQ_GNtQ" colab_type="code" outputId="7f79093f-780e-407f-f938-fb59a56620f6" colab={"base_uri": "https://localhost:8080/", "height": 355}
# The dataframe of actual offensive points for each game from 2000-2019
original_df.head()
# + id="JaUeCnKAGTXY" colab_type="code" outputId="c7131802-c867-46a8-e2c5-1c6d709b08ca" colab={"base_uri": "https://localhost:8080/", "height": 204}
# The dataframe of all 2019 offensive players (kickers and offense)
player_df.head()
# + id="-tpuw1teGd-v" colab_type="code" outputId="ada764fd-774a-40ff-93fb-166fe3380104" colab={"base_uri": "https://localhost:8080/", "height": 204}
# The dataframe of bye weeks for 2019 teams
bye_df.head()
# + id="5rdO3zKnGhX3" colab_type="code" outputId="2aaacf21-8443-4450-f37e-f0fccdc4dc16" colab={"base_uri": "https://localhost:8080/", "height": 204}
# The dataframe of injuries for 2019 players
injury_df.head()
# + id="wrYthbyaGmkS" colab_type="code" colab={}
# Add a row to the final_df dataframe
# Each row represents the predicted points for each player
def add_row(df, p, f, l, n, pos, wc, wp, wa, cur, pred, act):
df = df.append({'player': p,
'first': f,
'last': l,
'name': n,
'position': pos,
wc: cur,
wp: pred,
wa: act
}, ignore_index=True)
return df
# + id="ZTGFQ4ClGwAP" colab_type="code" colab={}
# The main code for iterating through the player(offense and kicker) list, calculating the points and adding the rows
# to the final_df dataframe.
def main(w):
week_cname = 'week' + str(w) + '-cur'
week_pname = 'week' + str(w) + '-pred'
week_aname = 'week' + str(w) + '-act'
column_names = ['player',
'first',
'last',
'name',
'position',
week_cname,
week_pname,
week_aname
]
player_list = offense_df['player'].tolist()
end_col = 308 + w
week_df = pd.DataFrame(columns = column_names)
for player in player_list:
first = player_df['first'].loc[(player_df['player']==player)].iloc[0]
last = player_df['last'].loc[(player_df['player']==player)].iloc[0]
name = player_df['name'].loc[(player_df['player']==player)].iloc[0]
position1 = player_df['position1'].loc[(player_df['player']==player)].iloc[0]
start_year = player_df['start'].loc[(player_df['player']==player)].iloc[0]
row = original_df.index[(original_df['player']==player)][0]
team = players_df['cteam'].loc[(players_df['player']==player)].iloc[0]
injury_weeks = injury_df['week'].loc[(injury_df['player']==player)]
if (len(injury_weeks)==0):
injury_week = 18
else:
injury_week = min(injury_weeks)
bye_week = bye_df['bye-week'].loc[(bye_df['player']==team)].iloc[0]
if start_year < 2000:
start_year = 2000
col = ((start_year - 2000) * 16) + 5
if w > bye_week:
new_col = end_col - 1
else:
new_col = end_col
train_data = original_df.iloc[row, col:new_col]
if w == 1:
current = original_df.iloc[row, 293:309]
else:
current = original_df.iloc[row, 309:new_col]
cur_points = current.sum()
actuals = original_df.iloc[row, new_col:325]
act_points = actuals.sum()
print(player, train_data.sum())
if (start_year < 2016) & (player != 'GG-0310') & (train_data.sum() > 1):
# ARIMA model
model = pm.auto_arima(train_data, start_p=1, start_q=1,
test='adf', # use adftest to find optimal 'd'
max_p=3, max_q=3, # maximum p and q
m=1, # frequency of series
d=None, # let model determine 'd'
seasonal=False, # No Seasonality
start_P=0,
D=0,
trace=False,
error_action='ignore',
suppress_warnings=True,
stepwise=True)
# Forecast
if w > bye_week:
n_periods = 17 - w + 1
else:
n_periods = 17 - w
fc = model.predict(n_periods=n_periods, return_conf_int=False)
index_of_fc = np.arange(len(train_data), len(train_data)+n_periods)
fc_series = pd.Series(fc, index=index_of_fc)
if (injury_week <= w):
pred_points = 0
else:
pred_points = fc_series.sum()
else:
pred_points = train_data.mean() * (18 - w)
week_df = add_row(week_df, player, first, last, name, position1, week_cname,
week_pname, week_aname, cur_points, pred_points, act_points)
return week_df
# + id="qMBR3rxvJiw_" colab_type="code" outputId="56d1c305-9f8c-4954-d3fd-6b6b3c5bce9f" colab={"base_uri": "https://localhost:8080/", "height": 1000}
week = 1
final_df = main(week)
# + id="g0Z_Xwf5JnpW" colab_type="code" outputId="6ede5cc8-ba8f-4fbe-e90c-ff2fed35b5f9" colab={"base_uri": "https://localhost:8080/", "height": 1000}
final_df.head(50)
# + id="J6uBs2d2MZDq" colab_type="code" colab={}
week_pred = 'week' + str(week) + '-pred'
week_act = 'week' + str(week) + '-act'
week_diff = 'week' + str(week) + '-diff'
week_pct = 'week' + str(week) + '-pct'
# + id="saRhE3aSMfDH" colab_type="code" colab={}
# The final_df dataframe
final_df[week_diff] = final_df[week_pred] - final_df[week_act]
final_df[week_pct] = final_df[week_diff] / final_df[week_pred]
final_df[week_pred] = final_df[week_pred].astype(int)
# + id="mZ-SzgRTMjaV" colab_type="code" outputId="491d3b27-343b-4f92-aa63-fb72cb817506" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Calculate the metrics
pred_median_error = final_df[week_pct].median()
print('Median Error - %.4f%%' % (pred_median_error * 100))
# + id="11LISNgsMn-r" colab_type="code" outputId="76b257a1-cc24-402a-b328-22157a6b9ef4" colab={"base_uri": "https://localhost:8080/", "height": 1000}
final_df.head(50)
# + id="Oe0LqAy9LIcU" colab_type="code" colab={}
# Save the results to .csv file
file_name = '/content/week' + str(week) + '-pred-offense-norookies.csv'
final_df.to_csv(file_name, index=False)
| predictions-final/Offense_Predictions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:dl]
# language: python
# name: conda-env-dl-py
# ---
# <div class="alert alert-block alert-info">
# <font size="6"><b><center> Section 1</font></center>
# <br>
# <font size="6"><b><center> Introduction to PyTorch </font></center>
# </div>
# # Fundamental Building Blocks
# * Tensor and Tensor Operations
#
# * PyTorch’s Tensor Libraries
#
# * Computational Graph
#
# * Gradient Computation
#
# * Linear Mapping
#
# * PyTorch’s non-linear activation functions:
# * Sigmoid, tanh, ReLU, Leaky ReLU
#
# * Loss Function
#
# * Optimization algorithms used in training deep learning models
# # A 2-Layer Feed-Forward Neural Network Architecture
# ## Some notations on a simple feed-forward network
# $\bar{\mathbf{X}} = \{X_1, X_2, \dots, X_K, 1 \}$ is a $n \times K$ matrix of $K$ input features from $n$ training examples
#
# $X_k$ $ (k = 1,\dots,K) $ is a $n \times 1$ vector of n examples corresponding to feature $k$
#
# $\bar{\mathbf{W}}_{Xh} = \{w_1, w_2 \dots, w_p \}$
#
# $\bar{\mathbf{W}}_{Xh}$ of size $PK$
#
# where $P$ is the number of units in the hidden layer 1
#
# and K is the number of input features
#
# $\mathbf{b}$ bias
#
#
# ## A Simple Neural Network Architeture
# The input layer contains $d$ nodes that transmit the $d$ features $\mathbf{X} = \{x_1, \dots, x_d, 1 \}$ with edges of weights $\mathbf{W} = \{w_1, \dots, w_d, b \}$ to an output node.
#
# Linear function (or linear mapping of data): $\mathbf{W} \cdot \mathbf{X} + b = b + \sum_{i=1}^d w_i x_i $
#
# $ y = b + \sum_{i=1}^d w_i x_i $ where $w$'s and $b$ are parameters to be learned
# # Tensor and Tensor Operations
# There are many types of tensor operations, and we will not cover all of them in this introduction. We will focus on operations that can help us start developing deep learning models immediately.
#
# The official documentation provides a comprehensive list: [pytorch.org](https://pytorch.org/docs/stable/torch.html#tensors)
#
#
# * Creation ops: functions for constructing a tensor, like ones and from_numpy
#
# * Indexing, slicing, joining, mutating ops: functions for changing the shape, stride or content a tensor, like transpose
#
# * Math ops: functions for manipulating the content of the tensor through computations
#
# * Pointwise ops: functions for obtaining a new tensor by applying a function to each element independently, like abs and cos
#
# * Reduction ops: functions for computing aggregate values by iterating through tensors, like mean, std and norm
#
# * Comparison ops: functions for evaluating numerical predicates over tensors, like equal and max
#
# * Spectral ops: functions for transforming in and operating in the frequency domain, like stft and hamming_window
#
# * Other operations: special functions operating on vectors, like cross, or matrices, like trace
#
# * BLAS and LAPACK operations: functions following the BLAS (Basic Linear Algebra Subprograms) specification for scalar, vector-vector, matrix-vector and matrix-matrix operations
#
# * Random sampling: functions for generating values by drawing randomly from probability distributions, like randn and normal
#
# * Serialization: functions for saving and loading tensors, like load and save
#
# * Parallelism: functions for controlling the number of threads for parallel CPU execution, like set_num_threads
#
#
# Import torch module
import torch
torch.version.__version__
# ## Creating Tensors and Examining tensors
# * `rand()`
#
# * `randn()`
#
# * `zeros()`
#
# * `ones()`
#
# * using a `Python list`
# ### Create a 1-D Tensor
# - PyTorch provides methods to create random or zero-filled tensors
# - Use case: to initialize weights and bias for a NN model
import torch
# `torch.rand()` returns a tensor of random numbers from a uniform [0,1) distribution
#
# [Source: Torch's random sampling](https://pytorch.org/docs/stable/torch.html#random-sampling)
# Draw a sequence of 10 random numbers
x = torch.rand(10)
type(x)
x.size()
print(x.min(), x.max(), x.mean(), x.std())
# Draw a matrix of size (10,3) random numbers
W = torch.rand(10,3)
type(W)
W.size()
W
# Another common random sampling is to generate random number from the standard normal distribution
# `torch.randn()` returns a tensor of random numbers from a standard normal distribution (i.e. a normal distribution with mean 0 and variance 1)
#
# [Source: Torch's random sampling](https://pytorch.org/docs/stable/torch.html#random-sampling)
W2 = torch.randn(10,3)
type(W2)
W2.dtype
W2.shape
W2
# **Note: Though it looks like it is similar to a list of number objects, it is not. A tensor stores its data as unboxed numeric values, so they are not Python objects but C numeric types - 32-bit (4 bytes) float**
# `torch.zeros()` can be used to initialize the `bias`
b = torch.zeros(10)
type(b)
b.shape
b
# Likewise, `torch.ones()` can be used to create a tensor filled with 1
a = torch.ones(3)
type(a)
a.shape
a
A = torch.ones((3,3,3))
A
# Convert a Python list to a tensor
A.shape
l = [1.0, 4.0, 2.0, 1.0, 3.0, 5.0]
torch.tensor(l)
# Subsetting a tensor: extract the first 2 elements of a 1-D tensor
torch.tensor([1.0, 4.0, 2.0, 1.0, 3.0, 5.0])[:2]
# ### Create a 2-D Tensor
a = torch.ones(3,3)
a
a.size()
b = torch.ones(3,3)
type(b)
# Simple addition
c = a + b
type(c)
c.type()
c.size()
# Create a 2-D tensor by passing a list of lists to the constructor
d = torch.tensor([[1.0, 4.0], [2.0, 1.0], [3.0, 5.0]])
d
d.size()
# We will obtain the same result by using `shape`
d.shape
# $[3,2]$ indicates the size of the tensor along each of its 2 dimensions
# Using the 0th-dimension index to get the 1st dimension of the 2-D tensor.
# Note that this is not a new tensor; this is just a different (partial) view of the original tensor
d[0]
d
d.storage()
e = torch.tensor([[[1.0, 3.0],
[5.0, 7.0]],
[[2.0, 4.0],
[6.0, 8.0]],
])
e.storage()
e.shape
e.storage_offset()
e.stride()
e.size()
points = torch.tensor([[1.0, 4.0], [2.0, 1.0], [3.0, 5.0]])
points
points.size()
points.stride()
points.storage()
# ## Subset a Tensor
points[2]
points[:2]
points[1:] # all rows but first, implicitly all columns
points[1:, :] # all rows but first, explicitly all columns
points[0,0]
points[0,1]
points[1,0]
points[0]
# **Note the changing the `sub-tensor` extracted (instead of cloned) from the original will change the original tensor**
second_points = points[0]
second_points
second_points[0] = 100.0
points
points[0,0]
# **If we don't want to change the original tensure when changing the `sub-tensor`, we will need to clone the sub-tensor from the original**
a = torch.tensor([[1.0, 4.0], [2.0, 1.0], [3.0, 5.0]])
b = a[0].clone()
b
b[0] = 100.0
a
b
# ## Transpose a Tensor
# ### Transposing a matrix
a
a_t = a.t()
a_t
a.storage()
a_t.storage()
# **Transposing a tensor does not change its storage**
id(a.storage()) == id(a_t.storage())
# ### Transposing a Multi-Dimensional Array
A = torch.ones(3, 4, 5)
A
# To transpose a multi-dimensional array, the dimension along which the tanspose is performed needs to be specified
A_t = A.transpose(0,2)
A.size()
A_t.size()
A.stride()
A_t.stride()
# ### Contiguous Tensors
# A tensor whose values are laid out in the storage starting from the **right-most dimension** onwards (i.e. moving along rows for a 2D tensor), is defined as contiguous
a
a.is_contiguous()
a.storage()
# ### Tensor Data Type
a.dtype
A.dtype
# ### Explicitly Specifying a Tensor Type During Creation
torch.zeros(3,2).double()
torch.zeros(2,3).to(torch.double)
# ### Type Casting
random_nums = torch.randn(10, 1)
random_nums_short = random_nums.type(torch.short)
random_nums.dtype
random_nums_short.dtype
# ### NumPy Interoperability
x = torch.ones(3,3)
x
x_np = x.numpy()
x.dtype
x_np.dtype
x2 = torch.from_numpy(x_np)
x2.dtype
# ### Tensor Serialization
import h5py
# **A very powerful feature of HDF5 file format is that it allows for indexing datasets on disk and only access the elements of interest. This feature lets you store huge amounts of numerical data, and easily manipulate that data from NumPy. For reference, see [h5py](https://www.h5py.org/)**
f = h5py.File('ourpoints.hdf5', 'w')
dset = f.create_dataset('coords', data=x2.numpy())
f.close()
f = h5py.File('ourpoints.hdf5', 'r')
dset = f['coords']
last_x2 = dset[1:]
last_x2
# +
# Importantly, we can pass the object and obtain a PyTorch tensor directly
f = h5py.File('ourpoints.hdf5', 'r')
dset = f['coords']
#last_x2 = dset[1:]
last_x2 = torch.from_numpy(dset[1:])
f.close()
last_x2
# -
# ### Tensors on GPU
# We will discuss more about this in the last section of the course
# ```python
# matrix_gpu = torch.tensor([[1.0, 4.0], [2.0, 1.0], [3.0, 4.0]], device='cuda')
# # transfer a tensor created on the CPU onto GPU using the to method
# x2_gpu = x2.to(device='cuda')
#
# points_gpu = points.to(device='cuda:0')
# ```
#
# CPU vs. GPU Performance Comparison
# ```python
# a = torch.rand(10000,10000)
# b = torch.rand(10000,10000)
# a.matmul(b)
#
# #Move the tensors to GPU
# a = a.cuda()
# b = b.cuda()
# a.matmul(b)
#
# ```
# # Gradient Computation
# Partial derivative of a function of several variables:
#
# $$ \frac{\partial f(x_1, x_2, \dots, x_p)}{\partial x_i} |_{\text{other variables constant}}$$
# * `torch.Tensor`
#
# * `torch.autograd` is an engine for computing vector-Jacobian product
#
# * `.requires_grad`
#
# * `.backward()`
#
# * `.grad`
#
# * `.detach()`
#
# * `with torch.no_grad()`
#
# * `Function`
#
# * `Tensor` and `Function` are connected and build up an acyclic graph, that encodes a complete history of computation.
# Let's look at a couple of examples:
# Example 1
# 1. Create a variable and set `.requires_grad` to True
# +
import torch
x = torch.ones(5,requires_grad=True)
#from torch.autograd import Variable
#x = Variable(torch.ones(5),requires_grad=True)
# -
x
x.type
x.grad
# Note that at this point, `x.grad` does not output anything because there is no operation performed on the tensor `x` yet. However, let's create another tensor `y` by performing a few operations (i.e. taking the mean) on the original tensor `x`.
y = x + 2
z = y.mean()
z.type
z
z.backward()
x.grad
x.grad_fn
x.data
y.grad_fn
z.grad_fn
# Example 2
x = torch.ones(2, 2, requires_grad=True)
y = x + 5
z = 2 * y * y # 2*(x+5)^2
h = z.mean()
z
z.shape
h.shape
h
h.backward()
print(x.grad)
x
# # Some notations on a simple feed-forward network
# ## A Simple Neural Network Architeture
# The input layer contains $d$ nodes that transmit the $d$ features $\mathbf{X} = \{x_1, \dots, x_d, 1 \}$ with edges of weights $\mathbf{W} = \{w_1, \dots, w_d, b \}$ to an output node.
#
# Linear function (or linear mapping of data): $\mathbf{W} \cdot \mathbf{X} + b = b + \sum_{i=1}^d w_i x_i $
#
# $ y = b + \sum_{i=1}^d w_i x_i $ where $w$'s and $b$ are parameters to be learned
# + [markdown] heading_collapsed=true
# ## Create data
# + hidden=true
def get_data():
train_X = np.asarray([3.3,4.4,5.5,6.71,6.93,4.168,9.779,6.182,7.59,2.167,
7.042,10.791,5.313,7.997,5.654,9.27,3.1])
train_Y = np.asarray([1.7,2.76,2.09,3.19,1.694,1.573,3.366,2.596,2.53,1.221,
2.827,3.465,1.65,2.904,2.42,2.94,1.3])
dtype = torch.FloatTensor
X = Variable(torch.from_numpy(train_X).type(dtype),requires_grad=False).view(17 ,1)
y = Variable(torch.from_numpy(train_Y).type(dtype),requires_grad=False)
return X, y, train_X, train_Y
# + [markdown] heading_collapsed=true
# ## Create (and Initialize) parameters
# + hidden=true
def get_weights():
w = Variable(torch.randn(1),requires_grad = True)
b = Variable(torch.randn(1),requires_grad=True)
return w,b
# + [markdown] heading_collapsed=true
# ## Visualize the data
# + hidden=true
# Get the data
X, y, X_np, y_np = get_data()
# + hidden=true
import matplotlib.pyplot as plt
from matplotlib.pyplot import figure
figure(num=None, figsize=(8, 6), dpi=100, facecolor='w', edgecolor='k')
plt.scatter(X_np,y_np)
plt.show()
# + [markdown] heading_collapsed=true
# ## Network Implementation
# + hidden=true
def simple_network(x):
y_pred = torch.matmul(x,w)+b
return y_pred
# + [markdown] hidden=true
# We do not need to implement a neural network model manually.
#
# PyTorch provides a higher-level abstraction in `torch.nn` called **layers**, which will take care of most of these underlying initialization and operations associated with most of the common techniques available in the neural network
# + [markdown] heading_collapsed=true
# ## Common Activation Functions
# + [markdown] hidden=true
# * **Sigmoid**: $\frac{1}{1 + exp(-x)}$
#
#
# * **Tanh**: $\frac{e^x - e^{-x}}{e^x + e^{-x}}$
#
#
# * **ReLU**: $max(0,x) + negative_slope∗min(0,x)$
#
#
# * **LeakyReLU(x)**: $max(0,x) + negative_slope∗min(0,x)$
#
#
# Reference: [Many more activation functions in PyTorch](https://pytorch.org/docs/stable/nn.html?highlight=activation%20function)
# + [markdown] heading_collapsed=true
# ## Define a loss function
# + hidden=true
def loss_fn(y,y_pred):
loss = (y_pred-y).pow(2).sum()
for param in [w,b]:
if not param.grad is None: param.grad.data.zero_()
loss.backward()
return loss.data[0]
# + [markdown] heading_collapsed=true
# ## Optimize the network
# + hidden=true
def optimize(learning_rate):
w.data -= learning_rate * w.grad.data
b.data -= learning_rate * b.grad.data
# + hidden=true
learning_rate = 0.0001
x,y,x_np,y_np = get_data() # x - represents training data,y - represents target variables
w,b = get_weights() # w,b - Learnable parameters
for i in range(500):
y_pred = simple_network(x) # function which computes wx + b
loss = loss_fn(y,y_pred) # calculates sum of the squared differences of y and y_pred
if i % 50 == 0:
print(loss)
optimize(learning_rate) # Adjust w,b to minimize the loss
# -
# # Lab 1
# Create a tensor of 20 random numbers from the uniform [0,1) distribution
# YOUR CODE HERE (1 line)
import torch
z = torch.rand(20)
z
print(z.min(),z.max())
# What is the mean of these numbers?
import numpy as np
# YOUR CODE HERE (1 line)
#np.mean(x.detach().numpy())
x.mean()
# Create a tensor of 5 zeros
# YOUR CODE HERE (1 line)
b = torch.zeros(5)
b
# Create a tensor of 5 ones
# YOUR CODE HERE (1 line)
a = torch.ones(5)
a
# Given the follow tensor, subset the first 2 rows and first 2 columns of this tensor.
A = torch.rand(4,4) # uniform[0,1)
# YOUR CODE HERE (1 line)
print(A)
print(A[:2,:2])
# What is the shape of the following tensor?
X = torch.randint(0, 10, (2, 5, 5))
# YOUR CODE HERE (1 line)
X.shape
X
# +
# Consider the following tensor.
# What are the gradients after the operations?
p = torch.ones(10, requires_grad=True)
q = p + 2
r = q.mean()
# YOUR CODE HERE (2 lines)
# -
r.backward()
p.grad
| Notebook/Section 1 - Introduction to PyTorch-v2-soln.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
from keras.models import Sequential
from keras.layers import Dense, LSTM, Conv1D, MaxPooling1D, Flatten, TimeDistributed, ConvLSTM2D, Reshape
import tensorflow as tf
import sklearn.metrics as sm
import keras
from keras import backend as K
# -
data = pd.read_csv("train.csv")
dtest = pd.read_csv("test.csv")
data.head()
data = data.drop(["ID","X0","X1","X2","X3","X4","X5","X6","X8" ], axis = 1)
data.head()
data.shape
dtest
dtest = dtest.drop(["ID","X0","X1","X2","X3","X4","X5","X6","X8" ], axis = 1)
dtest.head()
dtest.shape
X_train, y_train = data.values[0:3499,1:], data.values[0:3499, :1].ravel()
X_valid, y_valid = data.values[3500:4208,1:], data.values[3500:4208, :1].ravel()
X_test = dtest.values[:, :]
print("X_train----",X_train.shape)
print("y_train----",y_train.shape)
print("X_valid----",X_valid.shape)
print("y_valid----",y_valid.shape)
print("X_test-----",X_test.shape)
X_train[0]
y_train[0]
# y_train = min_max_scaler.fit_transform(y_train)
def mean_absolute_percentage_error(y_true, y_pred):
y_true, y_pred = np.array(y_true), np.array(y_pred)
return np.mean(np.abs((y_true - y_pred) / y_true)) * 100
def metrics(pred, y_test):
evs = sm.explained_variance_score(y_test, pred)
me = sm.max_error(y_test, pred)
mae = sm.mean_absolute_error(y_test, pred)
mse = sm.mean_squared_error(y_test, pred)
rmse = np.sqrt(mse)
#msle = sm.mean_squared_log_error(y_test, pred)
m_ae = sm.median_absolute_error(y_test, pred)
r2 = sm.r2_score(y_test, pred)
#mpd = sm.mean_poisson_deviance(y_test, pred)
#mgd = sm.mean_gamma_deviance(y_test, pred)
mape = mean_absolute_percentage_error(pred, y_test)
return({'Explained Variance Score': evs,
'Max Error': me,
'Mean Absolute Error': mae,
'Mean Squared Error': mse,
'Root Mean Squared Error': rmse,
#'Mean Squared Log Error': msle,
'Median Absolute Error': m_ae,
'R² Score': r2,
#'Mean Poisson Deviance': mpd,
#'Mean Gamma Deviance': mgd,
'Mean Absolute Percentage Error': mape
})
subsequences = 2
timesteps = X_train.shape[1]//subsequences
timesteps
X_train = X_train.reshape((X_train.shape[0], subsequences, 1, timesteps, 1))
X_valid = X_valid.reshape((X_valid.shape[0], subsequences, 1, timesteps, 1))
X_test = X_test.reshape((X_test.shape[0], subsequences, 1, timesteps, 1))
X_train.shape
early_stop = tf.keras.callbacks.EarlyStopping(monitor="loss", patience=20)
modelConvLSTM = Sequential()
modelConvLSTM.add(ConvLSTM2D(filters=32, kernel_size=(1,2), activation='relu', return_sequences=True,input_shape=(X_train.shape[1], 1, X_train.shape[3], X_train.shape[4])))
modelConvLSTM.add(ConvLSTM2D(filters=16, kernel_size=(1,2), activation='relu'))
modelConvLSTM.add(Flatten())
modelConvLSTM.add(Dense(1))
modelConvLSTM.compile(optimizer='adam', loss='mse')
history = modelConvLSTM.fit(X_train, y_train, batch_size=512, epochs=500, verbose=1, callbacks=[early_stop], validation_data = (X_valid, y_valid))
ConvLSTMpred = modelConvLSTM.predict(X_valid, verbose=0)
ConvLSTMpred = ConvLSTMpred.reshape((ConvLSTMpred.shape[0]))
ConvLSTMresults = metrics(ConvLSTMpred, y_valid)
ConvLSTMresults
ConvLSTMpred = modelConvLSTM.predict(X_test, verbose=0)
ConvLSTMpred = ConvLSTMpred.reshape((ConvLSTMpred.shape[0]))
| conlstm2d 2,184 simple.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Can git compress 'similar' (binary) files in repositories?
#
# ## Preparations
py348 = "https://www.python.org/ftp/python/3.4.8/Python-3.4.8.tgz"
py348rc1 = "https://www.python.org/ftp/python/3.4.8/Python-3.4.8rc1.tgz"
# !wget $py348
# !wget $py348rc1
# +
# strip off URL path
import os.path
py348 = os.path.split(py348)[1]
py348rc1 = os.path.split(py348rc1)[1]
# use letters A and B for convenience, and strip off file extension (.tgz)
A = py348[:-4]
B = py348rc1[:-4]
# -
# # Experiment 1: Adding gzipped tar files to git
Atgz = A + '.tgz'
# !rm -rf .git # clean start
# !git init .
# !git add $Atgz
# !git commit -m "file A"
# !du -hs .git
# Can we optimise disk usage further? (No:)
# Can we optimise git's storage?
# !git gc --aggressive
# !du -hs .git
Btgz = B + '.tgz'
# !git add $Btgz
# !git commit -m "file B"
# !du -hs .git
# !ls -lh Python*tgz
# Can we optimise git's storage?
# !git gc --aggressive
# !du -hs .git
# Conclusion 1:
# - Adding compressed files to git -> git cannot compress further
# - Adding two similar (but already compressed files) -> git cannot exploit similarity
# # Experiment 2: Adding (uncompressed) tar files to git
# !rm -f *.tar # tidy up
# !gunzip $Atgz $Btgz
# !ls -lh Python*tar
Atar = A + ".tar"
# !rm -rf .git # clean start
# !git init .
# !git add $Atar
# !git commit -m "file A"
# !du -hs .git
# Can we optimise git's storage?
# !git gc --aggressive
# !du -hs .git
# Yes, now git has achieved (approx) the same compression as gzip could do on the tarball.
Btar = B + ".tar"
# !git add $Btar
# !git commit -m "file B"
# !du -hs .git
# !git log --stat
# Can we optimise git's storage?
# !git gc --aggressive
# !du -hs .git
# Conclusion 2: Adding uncompressed files to git:
#
# - each file is compressed (although not efficiently as original gzip: 22MB instead of 19MB)
# - no apparent benefit in exploiting similarity of the two files
# - *until* 'git gc' is run; then very good compression of the two 'similar' files A and B.
# # Conclusion
#
# - Adding the two similar gzipped tar files to a git repository results in a repository size (i.e. size of ``.git`` of 38MB).
#
# - Adding the files as tar files, results in a repository size of 44MB. Once ``git gc`` is executed, the repository size shrinks to 19MB.
#
# So based on this, we should add tar files, not zipped archives.
#
# ## Acknowledgements
#
# Thanks to <NAME> who taught me about ``git gc`` and pointed to this study: https://gist.github.com/matthewmccullough/2695758
#
#
| How to add tar files to git.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Consider Scaling of the Upward-directed SWF for Clearsky
# We develop curves of SWF upwelling as function of SWF downwelling.
# ## Load Standard and User Libraries
# +
# %matplotlib inline
import os, sys, copy, glob
import numpy as np
import datetime
import seaborn as sns
import pylab as pl
import xarray as xr
# -
pl.rcParams['figure.figsize'] = (16.0, 8.0)
pl.rcParams['font.size'] = 18.0
pl.rcParams['lines.linewidth'] = 3
from tropy.standard_config import local_data_path
import tropy.analysis_tools.grid_and_interpolation as gi
import tropy.analysis_tools.statistics as stats
# +
import nawdex_analysis.analysis.ave_cre
import nawdex_analysis.io.collector
from nawdex_analysis.plot.exp_layout import get_exp_kws
# -
# ## Collect Data
rset = nawdex_analysis.io.collector.get_radflux4set( 'all', method = 'strict' )
# ## Scale Clearsky
scaling_factor = 0.88
nawdex_analysis.analysis.ave_cre.radflux2cre( rset, scaling = True, new_factor = scaling_factor, old_factor = 0.9 )
idlist = list( rset.idname.data )
#idlist.remove('msevi-scaled')
rclear = rset.sel(ct = 'clear_ocean', idname = idlist)
# ## Do Conditional Analysis
# +
cond = xr.Dataset()
bins = np.linspace(0,1000,11)
percentiles = [10, 25, 50, 75, 90]
mbins = gi.lmean(bins)
idlist.remove('msevi-scaled')
idlist.remove('msevi-not_scaled')
idlist.insert(0, 'msevi-scaled')
for i, idname in enumerate( idlist ):
v1 = -rclear.sel(idname = idname)['swf_down']
v2 = rclear.sel(idname = idname)['swf_up']
# calculate conditioned statistics
mbins = gi.lmean(bins)
v2p = stats.cond_perc_simple(v1, v2, bins, p=percentiles)
cond[idname] = xr.DataArray(v2p, coords=[percentiles, mbins], dims=['percentiles', 'bins'])
# +
idlist.insert(0, 'icon-scaled')
cond['icon-scaled'] = cond['10km_oneMom_Conv'].copy() * scaling_factor
# -
cond
# ## Plot Downwelling vs. Upwelling SWF
# +
pl.figure( figsize = (8,4))
nvars = len(idlist)
scal = 5
offset = -scal * nvars / 2.
for i, idname in enumerate( idlist ):
kws = get_exp_kws( idname, ptype = 'points' )
kws['linewidth'] = 2
if idname == 'icon-scaled':
kws['linestyle'] = '--'
kws['color'] = 'k'
# calculate conditioned statistics
mbins = cond['bins'].data
v2p = cond[idname].data
ymed = v2p[2]
yerr = np.abs( v2p[[1,3]] - v2p[2] )
# use shifted bins for visualization
sbins = mbins + scal * i + offset
ys = np.interp(sbins, mbins, ymed)
pl.errorbar(sbins, ys, yerr = yerr, label = idname, **kws)
sns.despine()
pl.xlabel('$|F_\mathrm{sw, down}|$ ($\mathrm{W\,m^{-2}}$)')
pl.ylabel('$F_\mathrm{sw, up}$ ($\mathrm{W\,m^{-2}}$)')
pl.subplots_adjust(bottom = 0.2)
pl.savefig('../pics/scaling_of_upwelling_clearsky.png', dpi = 300)
#pl.legend(loc = 'center right', bbox_to_anchor=(1.3, 0.5) )
# +
pl.figure( figsize = (8,4))
nvars = len(idlist)
scal = 5
offset = -scal * nvars / 2.
for i, idname in enumerate( idlist ):
kws = get_exp_kws( idname, ptype = 'points' )
kws['linewidth'] = 2
if idname == 'icon-scaled':
kws['linestyle'] = '--'
kws['color'] = 'k'
# calculate conditioned statistics
mbins = cond['bins'].data
v2p = cond[idname].data
ymed = v2p[2]
yerr = np.abs( v2p[[1,3]] - v2p[2] )
# use shifted bins for visualization
sbins = mbins + scal * i + offset
# no shifted bins
sbins = mbins
ys = np.interp(sbins, mbins, ymed)
if idname not in ['msevi-scaled', '10km_oneMom_Conv', 'icon-scaled']:
kws['color'] = 'gray'
kws['linestyle'] = '-'
kws['alpha'] = 0.1
kws['marker'] = None
yerr = 0
else:
print(idname)
print(kws)
pl.errorbar(sbins, ys, yerr = yerr, label = idname, **kws)
sns.despine()
pl.xlabel('$|F_\mathrm{sw, down}|$ ($\mathrm{W\,m^{-2}}$)')
pl.ylabel('$F_\mathrm{sw, up}$ ($\mathrm{W\,m^{-2}}$)')
pl.subplots_adjust(bottom = 0.2)
pl.savefig('../pics/scaling_of_upwelling_clearsky.png', dpi = 300)
#pl.legend(loc = 'center right', bbox_to_anchor=(1.3, 0.5) )
| nbooks/02-Scaling_of_the_upwarding_clearsky_SWF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <img style="float: left; padding-right: 10px; width: 45px" src="https://raw.githubusercontent.com/Harvard-IACS/2018-CS109A/master/content/styles/iacs.png"> CS109B Data Science 2: Advanced Topics in Data Science
#
# ### Lab 3 - Clustering
#
#
#
# **Harvard University**<br>
# **Spring 2020**<br>
# **Instructors:** <NAME>, <NAME>, and <NAME><br>
# **Lab Instructors:** <NAME> and <NAME><br>
# **Content:** <NAME> and <NAME>
#
# ---
## RUN THIS CELL TO PROPERLY HIGHLIGHT THE EXERCISES
import requests
from IPython.core.display import HTML
styles = requests.get("https://raw.githubusercontent.com/Harvard-IACS/2019-CS109B/master/content/styles/cs109.css").text
HTML(styles)
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
from sklearn import preprocessing
# %matplotlib inline
# -
# ## Learning Objectives
#
# By the end of this lab, you should be able to:
# * Explain what PCA is and know the differences between it and clustering
# * Understand the common distance metrics (e.g., Euclidean, Manhattan, Hamming)
# * Understand how different clustering algorithms work (e.g., k-means, Hierarchical, DBScan)
# * Explain the trade-offs between the clustering approaches
# * Quantitatively describe the quality clusters' fit, according to different metrics
# * Comfortably cluster any new data that you are presented with
#
# **This lab corresponds to Lectures #4 and #5 and maps to Homework #2.**
#
# ## Table of Contents
#
# 1. PCA Refresher
# 2. Distant Metrics
# 3. Clustering Algorithms and Measuring Quality of Clusters
#
# ## 1. PCA Refresher
#
# <br>
#
# <div class="discussion"><b>Discussion #1</b> What is PCA? How can it be useful?</div>
#
#
# ### How to use it ( via `sklearn`):
#
# \# assume we a DataFrame `df`
#
# #### <font color='blue'>a. Instantiate a new PCA object</font>:
# `pca_transformer = PCA()`
#
# #### <font color='blue'>b. Fit some data (learns the transformation based on this data)</font>:
# `fitted_pca = pca_transformer.fit(df)`
#
# #### <font color='blue'>c. Transform the data to the reduced dimensions</font>:
# `pca_df = fitted_pca.transform(df)`
#
# Using two distinct steps (i.e., (b) and (c)) to fit and transform our data allows one the flexibility to transform any dataset according to our learned `fit()`. Alternatively, if you know you only want to transform a single dataset, you can combine (b) and (c) into one step:
# #### <font color='blue'>Fit and transform</font>:
# `pca_df = pca_transformer.fit_transform(df)`
#
# ### Example:
ms_df = pd.read_csv("../data/multishapes.csv")[['x','y']] # loads x,y columns of a dataset
pca_transformer = PCA()
fitted_pca = pca_transformer.fit(ms_df)
pca_df = fitted_pca.transform(ms_df)
# **NOTE:** The above PCA transformation is a bit silly because we started with 2 dimensions and are transforming it to 2 dimensions -- no reduction. The data is still transforming the original data by applying a linear transformation so as to capture the most variance, but PCA is even more useful when the original data is high-dimensional. This example was just to remind you of the syntax.
#
# <br>
#
# <div class="discussion"><b>Discussion #2:</b> We didn't scale our data before applying PCA. Should we usually do so? Why or why not?</div>
#
# <br><br>
#
# ## 2. Distance Metrics
#
# In the picture below, we are concerned with measuring the distance between two points, **p** and **q**.
# <center>
# <img src="../fig/manhattan_distance.svg" width="250">(edited from Wikipedia.org)
# </center>
#
# ### Euclidean Distance:
#
# The Euclidean distance measures the shortest path between the two points, navigating through all dimensions:
#
# <center>
# <img src="../fig/euclidean_eq.png">
# </center>
#
#
# ### Manhattan Distance:
#
# The Manhattan distance measures the cumulative difference between the two points, across all dimensions.
#
# <center>
# <img src="../fig/manhattan_eq.png">
# </center>
#
# <div class="discussion"><b>Discussion #3:</b> Where have we seen something like this before in CS109A? What are the effects of using one versus another?</div>
#
# <br>
#
# ### Hamming Distance (extra credit):
#
# If our two elements of comparison can be represented a sequence of discrete items, it can be useful to measure how many of their elements differ.
#
# For example:
# - `Mahmoud` and `Mahmood` differ by just 1 character and thus have a hamming distance of 1.
# - `10101` and `01101` have a hamming distance of 2.
# - `Mary` and `Barry` have a hamming distance of 3 (m->b, y->r, null->y).
#
# Note: the last example may seem sub-optimal, as we could transform Mary to Barry by just 2 operations (substituting the M with a B, then adding an 'r'). The very related **Levenshtein distance** can handle this, and thus tends to be more appropriate for Strings.
#
# <br><br>
#
# ## 3. Clustering Algorithms
#
# <br>
#
# <div class="exercise"><b>Question:</b> Why do we care about clustering? How/why is it useful?</div>
#
# <br>
#
# <center>
# <img src="../fig/spotify_dailymix.png">
# </center>
#
# <br><br>
#
# We will now walk through three clustering algorithms, first discussing them at a high-level, then showing how to implement them with Python libraries. Let's first load and scale our data, so that particular dimensions don't naturally dominate in their contributions in the distant calculations:
# loads and displays our summary statistics of our data
multishapes = pd.read_csv("../data/multishapes.csv")
ms_df = multishapes[['x','y']]
ms_df.describe()
multishapes
ms_df
# scales our data
scaled_df = pd.DataFrame(preprocessing.scale(ms_df), index=multishapes['shape'], columns = ms_df.columns)
scaled_df.describe()
# plots our data
msplot = scaled_df.plot.scatter(x='x',y='y',c='Black',title="Multishapes data",figsize=(11,8.5))
msplot.set_xlabel("X")
msplot.set_ylabel("Y")
plt.show()
# ## 3a. k-Means clustering:
#
# <br>
#
# <div class="exercise" style="background-color:#b3e6ff"><b>Table Exercise #1</b>: With your table, collectively discuss how k-means works. Use a whiteboard, draw a bunch of dots, and walk through each step of how the algorithm works. When you're confident of your answer, speak with a TF to verify its correctness.</div>
#
# ### Code (via `sklearn`):
from sklearn.cluster import KMeans
ms_kmeans = KMeans(n_clusters=3, init='random', n_init=3, random_state=109).fit(scaled_df)
# That's it! Just 1 line of code!
#
# Now that we've run k-Means, we can look at various attributes of our clusters. Full documenation is [here](https://scikit-learn.org/stable/modules/generated/sklearn.cluster.KMeans.html).
display(ms_kmeans.cluster_centers_)
display(ms_kmeans.labels_[0:10])
# ### Plotting
# Take note of matplotlib's `c=` argument to color items in the plot, along with our stacking two different plotting functions in the same plot.
plt.figure(figsize=(10,10))
plt.scatter(scaled_df['x'],scaled_df['y'], c=ms_kmeans.labels_);
plt.scatter(ms_kmeans.cluster_centers_[:,0],ms_kmeans.cluster_centers_[:,1], c='r', marker='h', s=100);
# <div class="exercise"><b>Question</b>: Is this expected or did something go wrong? Should we always scale our data before clustering?</div>
# ### Lessons:
# - Initializations matter; run multiple times
# - Total Squared distance should never get worse during an update
# - k-Means can struggle with clusters that are close together; they can get lumped into one
# - There's no notion of 'not part of any cluster' or 'part of two clusters'
# - [Visualization here](http://web.stanford.edu/class/ee103/visualizations/kmeans/kmeans.html)
# ## Quality of Clusters: Inertia
# Inertia measures the total squared distance from points to their cluster's centroid. We obviously want this distance to be relatively small. If we increase the number of clusters, it will naturally make the average distance smaller. If every point has its own cluster, then our distance would be 0. That's obviously not an ideal way to cluster. One way to determine a reasonable number of clusters to simply try many different clusterings as we vary **k**, and each time, measure the overall inertia.
# +
wss = []
for i in range(1,11):
fitx = KMeans(n_clusters=i, init='random', n_init=5, random_state=109).fit(scaled_df)
wss.append(fitx.inertia_)
plt.figure(figsize=(11,8.5))
plt.plot(range(1,11), wss, 'bx-')
plt.xlabel('Number of clusters $k$')
plt.ylabel('Inertia')
plt.title('The Elbow Method showing the optimal $k$')
plt.show()
# -
# Look for the place(s) where distance stops decreasing as much (i.e., the 'elbow' of the curve). It seems that 4 would be a good number of clusters, as a higher *k* yields diminishing returns.
# ## Quality of Clusters: Silhouette
#
# Let's say we have a data point $i$, and the cluster it belongs to is referred to as $C(i)$. One way to measure the quality of a cluster $C(i)$ is to measure how close its data points are to each other (within-cluster) compared to nearby, other clusters $C(j)$. This is what `Silhouette Scores` provide for us. The range is [-1,1]; 0 indicates a point on the decision boundary (equal average closeness to points intra-cluster and out-of-cluster), and negative values mean that datum might be better in a different cluster.
#
# Specifically, let $a(i)$ denote the average distance data point $i$ is to the other points in the same cluster:
#
# <center>
# <img src="../fig/silhouette_intra.png">
# </center>
#
# Similarly, we can also compute the average distance that data point $i$ is to all **other** clusters. The cluster that yields the minimum distance is denoted by $b(i)$:
#
# <center>
# <img src="../fig/silhouette_outer.png">
# </center>
#
# Hopefully our data point $i$ is much closer, on average, to points within its own cluster (i.e., $a(i)$ than it is to its closest neighboring cluster $b(i)$). The silhouette score quantifies this as $s(i)$:
#
# <center>
# <img src="../fig/silhouette_eq.png">
# </center>
#
# **NOTE:** If data point $i$ belongs to its own cluster (no other points), then the silhouette score is set to 0 (otherwise, $a(i)$ would be undefined).
#
# The silhouette score plotted below is the **overall average** across all points in our dataset.
#
# The `silhouette_score()` function is available in `sklearn`. We can manually loop over values of K (for applying k-Means algorithm), then plot its silhouette score. This should allow us to make a reasonable choice for selecting the 'optimal' number of clusters.
# +
from sklearn.metrics import silhouette_score
scores = [0]
for i in range(2,11):
fitx = KMeans(n_clusters=i, init='random', n_init=5, random_state=109).fit(scaled_df)
score = silhouette_score(scaled_df, fitx.labels_)
scores.append(score)
plt.figure(figsize=(11,8.5))
plt.plot(range(1,11), np.array(scores), 'bx-')
plt.xlabel('Number of clusters $k$')
plt.ylabel('Average Silhouette')
plt.title('Silhouette Scores for varying $k$ clusters')
plt.show()
# -
# ### Visualizing all Silhoutte scores for a particular clustering
#
# Below, we borrow from an `sklearn` example. The second plot may be overkill.
# - The second plot is just the scaled data. It is *not* a PCA plot
# - If you only need the raw silhouette scores, use the `silhouette_samples()` function
# +
from sklearn.metrics import silhouette_samples, silhouette_score
import matplotlib.cm as cm
#modified code from http://scikit-learn.org/stable/auto_examples/cluster/plot_kmeans_silhouette_analysis.html
def silplot(X, clusterer, pointlabels=None):
cluster_labels = clusterer.labels_
n_clusters = clusterer.n_clusters
# Create a subplot with 1 row and 2 columns
fig, (ax1, ax2) = plt.subplots(1, 2)
fig.set_size_inches(11,8.5)
# The 1st subplot is the silhouette plot
# The silhouette coefficient can range from -1, 1 but in this example all
# lie within [-0.1, 1]
ax1.set_xlim([-0.1, 1])
# The (n_clusters+1)*10 is for inserting blank space between silhouette
# plots of individual clusters, to demarcate them clearly.
ax1.set_ylim([0, len(X) + (n_clusters + 1) * 10])
# The silhouette_score gives the average value for all the samples.
# This gives a perspective into the density and separation of the formed
# clusters
silhouette_avg = silhouette_score(X, cluster_labels)
print("For n_clusters = ", n_clusters,
", the average silhouette_score is ", silhouette_avg,".",sep="")
# Compute the silhouette scores for each sample
sample_silhouette_values = silhouette_samples(X, cluster_labels)
y_lower = 10
for i in range(0,n_clusters+1):
# Aggregate the silhouette scores for samples belonging to
# cluster i, and sort them
ith_cluster_silhouette_values = \
sample_silhouette_values[cluster_labels == i]
ith_cluster_silhouette_values.sort()
size_cluster_i = ith_cluster_silhouette_values.shape[0]
y_upper = y_lower + size_cluster_i
color = cm.nipy_spectral(float(i) / n_clusters)
ax1.fill_betweenx(np.arange(y_lower, y_upper),
0, ith_cluster_silhouette_values,
facecolor=color, edgecolor=color, alpha=0.7)
# Label the silhouette plots with their cluster numbers at the middle
ax1.text(-0.05, y_lower + 0.5 * size_cluster_i, str(i))
# Compute the new y_lower for next plot
y_lower = y_upper + 10 # 10 for the 0 samples
ax1.set_title("The silhouette plot for the various clusters.")
ax1.set_xlabel("The silhouette coefficient values")
ax1.set_ylabel("Cluster label")
# The vertical line for average silhouette score of all the values
ax1.axvline(x=silhouette_avg, color="red", linestyle="--")
ax1.set_yticks([]) # Clear the yaxis labels / ticks
ax1.set_xticks([-0.1, 0, 0.2, 0.4, 0.6, 0.8, 1])
# 2nd Plot showing the actual clusters formed
colors = cm.nipy_spectral(cluster_labels.astype(float) / n_clusters)
ax2.scatter(X[:, 0], X[:, 1], marker='.', s=200, lw=0, alpha=0.7,
c=colors, edgecolor='k')
xs = X[:, 0]
ys = X[:, 1]
if pointlabels is not None:
for i in range(len(xs)):
plt.text(xs[i],ys[i],pointlabels[i])
# Labeling the clusters
centers = clusterer.cluster_centers_
# Draw white circles at cluster centers
ax2.scatter(centers[:, 0], centers[:, 1], marker='o',
c="white", alpha=1, s=200, edgecolor='k')
for i, c in enumerate(centers):
ax2.scatter(c[0], c[1], marker='$%d$' % int(i), alpha=1,
s=50, edgecolor='k')
ax2.set_title("The visualization of the clustered data.")
ax2.set_xlabel("Feature space for the 1st feature")
ax2.set_ylabel("Feature space for the 2nd feature")
plt.suptitle(("Silhouette analysis for KMeans clustering on sample data "
"with n_clusters = %d" % n_clusters),
fontsize=14, fontweight='bold')
# +
# run k-means with 3 clusters
ms_kmeans = KMeans(n_clusters=3, init='random', n_init=3, random_state=109).fit(scaled_df)
# plot a fancy silhouette plot
silplot(scaled_df.values, ms_kmeans)
# -
# <div class="exercise"><b>Exercise #1</b>: </div>
#
# Using the silhouette scores' optimal number of clusters (per the elbow plot above):
# - Fit a new k-Means model with that many clusters
# - Plot the clusters like we originally did with k-means
# - Plot the silhouette scores just like the above cells
# - Which seems like a better clustering (i.e., 3 clusters or the number returned by the elbow plot above)?
# +
# # %load solutions/exercise1-solution.py
ms_kmeans = KMeans(n_clusters=4, init='random', n_init=3, random_state=109).fit(scaled_df)
plt.figure(figsize=(10,10))
plt.scatter(scaled_df['x'], scaled_df['y'], c=ms_kmeans.labels_);
plt.scatter(ms_kmeans.cluster_centers_[:,0],ms_kmeans.cluster_centers_[:,1], c='r', marker='h', s=100);
# plot a fancy silhouette plot
silplot(scaled_df.values, ms_kmeans)
# -
# ## Quality of Clusters: Gap Statistic
# The gap statistic compares within-cluster distances (like in silhouette), but instead of comparing against the second-best existing cluster for that point, it compares our clustering's overall average to the average we'd see if the data were generated at random (we'd expect randomly generated data to not necessarily have any inherit patterns that can be easily clustered). For full details, you can [read the original research paper.](https://statweb.stanford.edu/~gwalther/gap)
#
# In essence, the within-cluster distances (in the elbow plot) will go down just becuse we have more clusters. We additionally calculate how much they'd go down on non-clustered data with the same spread as our data and subtract that trend out to produce the plot below.
# +
from gap_statistic import OptimalK
from sklearn.datasets.samples_generator import make_blobs
gs_obj = OptimalK()
n_clusters = gs_obj(scaled_df.values, n_refs=50, cluster_array=np.arange(1, 15))
print('Optimal clusters: ', n_clusters)
# -
gs_obj.gap_df
gs_obj.plot_results() # makes nice plots
# If we wish to add error bars to help us decide how many clusters to use, the following code displays such:
# +
def display_gapstat_with_errbars(gap_df):
gaps = gap_df["gap_value"].values
diffs = gap_df["diff"]
err_bars = np.zeros(len(gap_df))
err_bars[1:] = diffs[:-1] - gaps[:-1] + gaps[1:]
plt.scatter(gap_df["n_clusters"], gap_df["gap_value"])
plt.errorbar(gap_df["n_clusters"], gap_df["gap_value"], yerr=err_bars, capsize=6)
plt.xlabel("Number of Clusters")
plt.ylabel("Gap Statistic")
plt.show()
display_gapstat_with_errbars(gs_obj.gap_df)
# -
# For more information about the `gap_stat` package, please see [the full documentation here](https://github.com/milesgranger/gap_statistic).
# ## 3b. Agglomerative Clustering
#
# <br>
#
# <div class="exercise" style="background-color:#b3e6ff"><b>Table Exercise #2</b>: With your table, collectively discuss how agglomerative clustering works. Use a whiteboard, draw a bunch of dots, and walk through each step of how the algorithm works. When you're confident of your answer, speak with a TF to verify its correctness.</div>
#
# ### Code (via `scipy`):
#
# There are many different cluster-merging criteria, one of which is Ward's criteria. Ward's optimizes having the lowest total within-cluster distances, so it merges the two clusters that will harm this objective least.
# `scipy`'s agglomerative clustering function implements Ward's method.
# +
import scipy.cluster.hierarchy as hac
from scipy.spatial.distance import pdist
plt.figure(figsize=(11,8.5))
dist_mat = pdist(scaled_df, metric="euclidean")
ward_data = hac.ward(dist_mat)
hac.dendrogram(ward_data);
# -
# <div class="discussion"><b>Discussion #4</b>: How do you read a plot like the above? What are valid options for number of clusters, and how can you tell? Are some more valid than others? Does it make sense to compute silhouette scores for an agglomerative clustering? If we wanted to compute silhouette scores, what would we need for this to be possible?</div>
#
# ### Lessons:
# - It's expensive: O(n^3) time complexity and O(n^2) space complexity.
# - Many choices for linkage criteria
# - Every node gets clustered (no child left behind)
# # %load solutions/discussion4-solution.py
labellings = hac.fcluster(ward_data, t=25, criterion='distance')
silhouette_score(scaled_df, labellings)
# ## 3c. DBscan Clustering
# DBscan uses an intuitive notion of denseness to define clusters, rather than defining clusters by a central point as in k-means.
#
# ### Code (via `sklearn`):
# DBscan is implemented in good 'ol sklearn, but there aren't great automated tools for searching for the optimal `epsilon` parameter. For full documentation, please [visit this page](https://scikit-learn.org/stable/modules/generated/sklearn.cluster.DBSCAN.html)
from sklearn.cluster import DBSCAN
plt.figure(figsize=(11,8.5))
fitted_dbscan = DBSCAN(eps=0.2).fit(scaled_df)
plt.scatter(scaled_df['x'],scaled_df['y'], c=fitted_dbscan.labels_);
# **Note:** the dark purple dots are not clustered with anything else. They are lone singletons. You can validate such by setting epsilon to a very small value, and increase the min_samples to a high value. Under these conditions, nothing would cluster, and yet all dots become dark purple.
#
# <br>
#
# <div class="exercise"><b>Exercise #2</b>: Experiment with the above code by changing its epsilon value and the min_samples (what is the default value for it, since the above code doesn't specify a value?)</div>
# <br><br>
# Instead of just empirically observing how the epsilon value affects the clustering (which would be very costly for large, high-dimensional data), we can also inspect how far each data point is to its $N^{th}$ closest neighbor:
# +
from sklearn.neighbors import NearestNeighbors
# x-axis is each individual data point, numbered by an artificial index
# y-axis is the distance to its 2nd closest neighbor
def plot_epsilon(df, min_samples):
fitted_neigbors = NearestNeighbors(n_neighbors=min_samples).fit(df)
distances, indices = fitted_neigbors.kneighbors(df)
dist_to_nth_nearest_neighbor = distances[:,-1]
plt.plot(np.sort(dist_to_nth_nearest_neighbor))
plt.xlabel("Index\n(sorted by increasing distances)")
plt.ylabel("{}-NN Distance (epsilon)".format(min_samples-1))
plt.tick_params(right=True, labelright=True)
# -
plot_epsilon(scaled_df, 3)
# ### Lessons:
# - Can cluster non-linear relationships very well; potential for more natural, arbritrarily shaped groupings
# - Does not require specifying the # of clusters (i.e., **k**); the algorithm determines such
# - Robust to outliers
# - Very sensitive to the parameters (requires strong knowledge of the data)
# - Doesn't guarantee that every (or ANY) item will be clustered
#
# <br>
#
# <div class="discussion"><b>Discussion #5</b>: </div>When should we prefer one type of clustering over another? Should we always just try all of them? Imagine you work at Spotify and you want to create personalized playlists for each person. One could imagine a dataset exists whereby each row is a particular song, and the columns are features (e.g., tempo (BPM), average vocal frequency, amount of bass, sentiment of lyrics, duration in seconds, etc). Let's use clustering to group one's catalog of favorite music, which will serve as disjoint starting points for suggesting future songs. Specifically, imagine that you've 'liked' 500 songs on Spotify so far, and your recommendation algorithm needs to cluster those 500 songs. Would you first experiment with k-Means, Agglomerative, or DBScan? Why?
| content/labs/lab03/notebook/cs109b_lab03_clustering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### CoreBx_island - Try to process entire N. Core Banks
#
# Interpolate the North Core Banks DEMs onto rotated 1-m grid and save each as a .nc file.
#
# New in v2
# * Now 4D maps, two made made during visit to Santa Cruz and two ftp'd from Andy
# * Apr. 9 - changed to _v3 for Sep map
# * Now does the interpolation without the loop
# * Apr. 21 - moved origin to SE to accomodate curvature in NE end of island. Add 400 m to size of array.
# * Watch file names, esp. underline (or not) after "1m_DEM"
#
# TODO: The alongshore/cross-shore names are switched.
import numpy as np
import matplotlib.pyplot as plt
import xarray as xr
# from dask.distributed import LocalCluster
from scipy import interpolate, signal
# %matplotlib inline
# define all of the functions by runnng this python file
# %run -i CoreBx_funcs
def make_grid(name=None,e0=None,n0=None,xlen=None,ylen=None,dxdy=None,theta=None):
nx = int((1./dxdy)*xlen)
ny = int((1./dxdy)*ylen)
xcoords = np.linspace(0.5*dxdy,xlen-0.5*dxdy,nx)
ycoords = np.linspace(0.5*dxdy,ylen-0.5*dxdy,ny)
# these will be the coordinates in rotated space
xrot, yrot = np.meshgrid(xcoords, ycoords ,sparse=False, indexing='xy')
print('Shape of xrot, yrot: ',np.shape(xrot),np.shape(yrot))
shp = np.shape(xrot)
xu, yu = box2UTMh(xrot.flatten(), yrot.flatten(), e0, n0, theta)
xu=np.reshape(xu,shp)
yu=np.reshape(yu,shp)
# write the UTM coords of the corners to an ASCII file
corners = np.asarray( [[xu[0][0],yu[0][0]],\
[xu[0][-1],yu[0][-1]],\
[xu[-1][-1],yu[-1][-1]],\
[xu[-1][0],yu[-1][0]],\
[xu[0][0],yu[0][0]]])
print(corners)
fn = name+'.csv'
np.savetxt(fn, corners, delimiter=",")
return xu, yu, xrot, yrot, xcoords, ycoords
# +
# April 9, 2020: Replaced "2019-09-12-13_1m_DEM_4D_crop.tif",\
# with _v3 and re-ran on my desktop
fdir = "C:/crs/proj/2019_DorianOBX/Santa_Cruz_Products/clipped_dems/"
#fdir = "D:/crs/proj/2019_DorianOBX/Santa_Cruz_Products/clipped_dems/"
fnames = (\
"2019-08-30_1m_DEM_4D_crop2.tif",\
"2019-09-12-13_1mDEM_4D_v3.tif",\
"2019-10-11_1m_DEM_4D_crop.tif",\
"2019-11-26_1m_DEM_4D_crop.tif")
titles = ([\
"8/30/2020 pre-Dorian",\
"9/12-13/2020 post-Dorian",\
"10/11/2020",\
"11/26 post-Nor'easter"])
nf = len(fnames)
fill_fnames = ('EBK_201909_YesLidar_Comb_Extent.tif')
fill_titles = ('Sep_fill')
# optional median-filter smoothing of original maps
smooth = False
# kernal size...this should be an odd number >= dxy/0.1
ksize = 3
# +
# Make an array of dicts, where analysis region is defined by:
# name
# e0 - UTM Easting of origin [m]
# n0 - UTM Northing of origin [m]
# xlen - Length of alongshore axis [m]
# ylen - Length of cross-shore axis [m]
# dxdy - grid size (must be isotropic right now) [m]
# theta - rotation CCW from x-axis [deg]
r = {'name':"ncorebx","e0": 378500.,"n0": 3856350.,"xlen": 36000.,"ylen": 1100.,"dxdy": 1.,"theta": 42.}
# +
# move the origin 400 m SE
xo,yo = xycoord(400.,42.+90)
print(xo,yo)
r['e0']=r['e0']+xo
r['n0']=r['n0']+yo
# add 400 m to ylen
r['ylen']=r['ylen']+400.
# -
r
print(r['name'])
xu,yu,xrot,yrot,xcoords,ycoords = make_grid(**r)
ny,nx = np.shape(xu)
print(ny,nx)
# +
# %%time
# Read in the fill map and make netcdf files
fn = fdir+fill_fnames
print(fn)
# open the tif with XArray as a DataArray
daf = xr.open_rasterio(fn)
print( np.shape(np.flipud(daf['y'].values)), np.shape(daf['x'].values), np.shape( np.flipud(daf.values)) )
x = daf['x'].values
y = np.flipud(daf['y'].values)
# Not sure how da.values got a singleton dimension, but squeeze gets rid of it.
# However, make sure to squeeze before flipping
z = np.flipud(np.squeeze(daf.values))
print(np.shape(x),np.shape(y),np.shape(z))
f = interpolate.RegularGridInterpolator( (y, x), z, method='linear')
# Array for interpolated elevations
zi=np.NaN*np.ones((ny,nx))
# this is a slow iteration through all of the points, but allows us to skip ones that are outside
for ij in np.ndindex(zi.shape):
try:
zi[ij]=f((yu[ij],xu[ij]))
except:
zi[ij]=np.NaN
# this is the fast technique.
# zi=f((yu,xu))
da = xr.DataArray(zi,dims=['Alongshore','Cross-shore'],coords={'Alongshore': ycoords, 'Cross-shore':xcoords })
da = da.chunk()
fno = r['name']+'_Sep_fill_v4.nc'
da.to_netcdf(fno)
# +
# %%time
dslist=[]
for i in range(nf):
fn = fdir+fnames[i]
print(i, fn)
# open the tif with XArray as a DataArray
da = xr.open_rasterio(fn)
print( np.shape(np.flipud(da['y'].values)), np.shape(da['x'].values), np.shape( np.flipud(da.values)) )
x = da['x'].values
y = np.flipud(da['y'].values)
# Not sure how da.values got a singleton dimension, but squeeze gets rid of it.
# However, make sure to squeeze before flipping
z = np.flipud(np.squeeze(da.values))
print(np.shape(x),np.shape(y),np.shape(z))
if(smooth):
# smooth with 2D running median
zs = signal.medfilt2d(z, kernel_size=ksize)
else:
zs = z
f = interpolate.RegularGridInterpolator( (y, x), zs, method='linear')
# Array for interpolated elevations
zi=np.NaN*np.ones((ny,nx))
# this is a slow iteration through all of the points, but allows us to skip ones that are outside
for ij in np.ndindex(zi.shape):
try:
zi[ij]=f((yu[ij],xu[ij]))
except:
zi[ij]=np.NaN
# this is the fast iteration, which only works when all of the source points fall inside the target box
# zi=f((yu,xu))
da = xr.DataArray(zi,dims=['Alongshore','Cross-shore'],coords={'Alongshore': ycoords, 'Cross-shore':xcoords })
da = da.chunk()
dslist.append(da)
dsa = xr.concat(dslist, dim='map')
fn = r['name']+'_v4.nc'
dsa.to_netcdf(fn)
# -
| CoreBx_island_v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 19. Knowledge in Learning
# **19.1** \[dbsig-exercise\]Show, by translating into conjunctive normal form and
# applying resolution, that the conclusion drawn on page [dbsig-page](#/)
# concerning Brazilians is sound.
#
# **19.2** For each of the following determinations, write down the logical
# representation and explain why the determination is true (if it is):
#
# 1. Design and denomination determine the mass of a coin.
#
# 2. For a given program, input determines output.
#
# 3. Climate, food intake, exercise, and metabolism determine weight gain
# and loss.
#
# 4. Baldness is determined by the baldness (or lack thereof) of one’s
# maternal grandfather.
#
# **19.3** For each of the following determinations, write down the logical
# representation and explain why the determination is true (if it is):
#
# 1. Zip code determines the state (U.S.).
#
# 2. Design and denomination determine the mass of a coin.
#
# 3. Climate, food intake, exercise, and metabolism determine weight gain
# and loss.
#
# 4. Baldness is determined by the baldness (or lack thereof) of one’s
# maternal grandfather.
#
# **19.4** Would a probabilistic version of determinations be useful? Suggest a
# definition.
#
# **19.5** \[ir-step-exercise\]Fill in the missing values for the clauses $C_1$ or
# $C_2$ (or both) in the following sets of clauses, given that $C$ is the
# resolvent of $C_1$ and $C_2$:
#
# 1. $C = {True} {\:\;{\Rightarrow}\:\;}P(A,B)$,
# $C_1 = P(x,y) {\:\;{\Rightarrow}\:\;}Q(x,y)$, $C_2
# = ??$.
#
# 2. $C = {True} {\:\;{\Rightarrow}\:\;}P(A,B)$, $C_1 = ??$,
# $C_2 = ??$.
#
# 3. $C = P(x,y) {\:\;{\Rightarrow}\:\;}P(x,f(y))$, $C_1 = ??$,
# $C_2 = ??$.
#
# If there is more than one possible solution, provide one example of each
# different kind.
#
# **19.6** \[prolog-ir-exercise\]Suppose one writes a logic program that carries
# out a resolution inference step. That is, let ${Resolve}(c_1,c_2,c)$
# succeed if $c$ is the result of resolving $c_1$ and $c_2$. Normally,
# ${Resolve}$ would be used as part of a theorem prover by calling it
# with $c_1$ and $c_2$ instantiated to particular clauses, thereby
# generating the resolvent $c$. Now suppose instead that we call it with
# $c$ instantiated and $c_1$ and $c_2$ uninstantiated. Will this succeed
# in generating the appropriate results of an inverse resolution step?
# Would you need any special modifications to the logic programming system
# for this to work?
#
# **19.7** \[foil-literals-exercise\]Suppose that is considering adding a literal
# to a clause using a binary predicate $P$ and that previous literals
# (including the head of the clause) contain five different variables.
#
# 1. How many functionally different literals can be generated? Two
# literals are functionally identical if they differ only in the names
# of the *new* variables that they contain.
#
# 2. Can you find a general formula for the number of different literals
# with a predicate of arity $r$ when there are $n$ variables
# previously used?
#
# 3. Why does not allow literals that contain no previously used
# variables?
#
# **19.8** Using the data from the family tree in
# Figure [family2-figure](#/), or a subset thereof, apply the
# algorithm to learn a definition for the ${Ancestor}$ predicate.
#
| Jupyter notebook/ilp-exercises.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pygame, sys
from pygame.locals import *
pygame.init()
DISPLAYSURF = pygame.display.set_mode((400, 300))
pygame.display.set_caption('Hello World!')
while True: # main game loop
for event in pygame.event.get():
if event.type == QUIT:
pygame.quit()
sys.exit()
pygame.display.update()
| python/games/.ipynb_checkpoints/Untitled1-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CSCI E7 Introduction to Programming with Python
# ## Lecture 07 Jupyter Notebook
# Fall 2021 (c) <NAME>
# # Topics
#
# - Code like a Pythonista
# - List Comprehensions
# - Program Practice
# - Slice is Forgiving
# - Tuples
# - Globals - Threat or Menace?
# - String Formatting
# - Tk File Picker
# ## Rules to live by
# ## Desiderata
#
# <NAME>
#
# Go placidly amid the noise and haste, and remember what peace there may be in silence.
#
# As far as possible, without surrender, be on good terms with all persons. Speak your truth quietly and clearly; and listen to others, even to the dull and the ignorant, they too have their story. Avoid loud and aggressive persons, they are vexations to the spirit.
#
# If you compare yourself with others, you may become vain and bitter; for always there will be greater and lesser persons than yourself. Enjoy your achievements as well as your plans. Keep interested in your own career, however humble; it is a real possession in the changing fortunes of time.
#
# Exercise caution in your business affairs, for the world is full of trickery. But let this not blind you to what virtue there is; many persons strive for high ideals, and everywhere life is full of heroism. Be yourself. Especially, do not feign affection. Neither be cynical about love, for in the face of all aridity and disenchantment it is perennial as the grass.
#
# Take kindly to the counsel of the years, gracefully surrendering the things of youth. Nurture strength of spirit to shield you in sudden misfortune. But do not distress yourself with dark imaginings. Many fears are born of fatigue and loneliness.
#
# Beyond a wholesome discipline, be gentle with yourself. You are a child of the universe, no less than the trees and the stars; you have a right to be here. And whether or not it is clear to you, no doubt the universe is unfolding as it should.
#
# Therefore be at peace with God, whatever you conceive Him to be, and whatever your labors and aspirations, in the noisy confusion of life, keep peace in your soul.
#
# With all its sham, drudgery and broken dreams, it is still a beautiful world.
#
# Be cheerful. Strive to be happy.
#
# https://en.wikipedia.org/wiki/Desiderata
# # Code Like a Pythonista!
#
# ### A guide to idiomatic Python
#
# ### http://www.omahapython.org/IdiomaticPython.html
#
# ### You know enough Python to start puzzling through this
# # List Comprehensions
# ### Iterating over something and returning a filtered list is a common operation.
# ### Common enough that there is an idiom for it.
# ### Before:
#
# ```python
# # Take the following fragment of pseudo-code
# new_list = []
# for item in collection:
# if condition(item):
# new_list.append(item)
# ```
#
# ### Rewrite as a List Comprehension
#
# ```python
# # and rewrite as the following pseudo-code
# new_list = [ item for item in collection
# if condition(item) ]
# ```
# ## We can use for filter, map and reduce
# - Filter: return those that match
# - Map: return a transformed list: Pounds to Kilos, or (Ht, Wt) -> BMI
# - Reduce: Take a list and summarize - take sum, average, stdev, etc...
#
# Our examples will focus on Filtering
# ## An Example of List Comprehension
lst = ['ship', 'set', 'mast']
# +
## Before
res = []
for word in lst:
if (len(word) == 4) and (word[-1] == 't'):
res.append(word)
# -
print(res)
# ## Replace with list comprehension
res = [ word for word in lst
if (len(word) == 4) and (word[-1] == 't') ]
print(res)
print([ word for word in lst
if (len(word) == 4) and (word[-1] == 't') ] )
# # Reversals: Before
s = ['abut', 'ant', 'fork', 'rat', 'tar', 'tuba', 'zap']
# +
def build_list(lst: list) -> list:
res = []
# Take each word in the list, and see if it's reverse is there as well
for word in lst:
rev = word[::-1]
# Don't include ('tuba', 'abut')
if (rev in lst) and (word <= rev):
res.append([word, rev])
return res
print(build_list(s))
# -
# ## Rewrite using List Comprehensions
# +
## List Comprehensions
def build_list(lst: list) -> list:
# lst has 113,809 words
# Find those with a reverse in the dictionary
lst1 = [wrd for wrd in lst
if (wrd[::-1] in lst)]
# 885 words
# [’tuba’,…,’aa’,…’yay’…’abut’,…]
# Filter out 'tuba' vs 'abut'
lst2 = [wrd for wrd in lst1
if (wrd[::-1] >= wrd)]
# 488 words - ‘tuba’ is gone
# Build a list of the pairs of words
return [[wrd, wrd[::-1]] for wrd in lst2]
# 488 pairs of words
print(build_list(s))
# -
# ## But we can write this as one list comprehension
# After
def build_list(lst: list) -> list:
return [[word, word[::-1]]
for word in lst
if (word <= word[::-1]) and (word[::-1] in lst)]
print(build_list(s))
# ## How about this?
# +
# After
def build_list(lst: list) -> list:
return [[word, word[::-1]] for word in lst if (word <= word[::-1]) and (word[::-1] in lst)]
print(build_list(s))
# -
# ## Violates PEP-8
# After
def build_list(lst: list) -> list:
return [[word, word[::-1]] for word in lst if (word <= word[::-1]) and (word[::-1] in lst)]
# 234567890123456789012345678901234567890123456789012345678901234567890123456789
# ## List Comprehensions are as fast as a hand coded loop
#
# ### No great speedup over finding reversals with a list
# ### The bulk of the time is the test for membership.
# # <NAME> on legibility
#
# “Programs must be written for people to read, and only incidentally for machines to execute.”
# # <NAME> on Notation
#
# "Of course, nothing is more incomprehensible than a symbolism which we do not understand....
#
# "[But by] relieving
# the brain of all unnecessary work, a good
# notation sets it free to concentrate on more
# advanced problems, and in effect increases
# [our] mental power"
# ## Pythonistas use list comprehension.
#
# ### You may not want to use them at first
# ### But to read code out there, you will need to be able to understand them.
# ## Here is another idiom you will encounter.
# ## What does this cell do?
[print(x) for x in range(3)]
# ## Document this: Catch the return value in variable 'garbage'
garbage = [print(x) for x in range(3)]
# ## What does this cell do?
#
# ## Idiom: Zero is False. Non zero integers are True.
#
# ## You will encounter this often.
#
# ## Pythonistas like Guido favor this idiom.
garbage = [print(x) for x in range(10) if x % 2]
# # Problem
# ## Find all numbers from 1-1000 that are divisible by 7 and have the digit 3 in them.
#
# ## We start by implementing the easy part to get going
# ## *Get it running, get it right, then make it fast*
# We start with the numbers less than 100
# Starting point
for i in range(100):
if (i % 7 ) == 0:
print(i)
# ### Start at 1, and simplify the test
#
# ### 0 is False, and other integers are not False
# Starting point
for i in range(1, 100):
if not (i % 7):
print(i)
# ## Rewrite as list comprehension
# Rewrite as list comprehension
[i for i in range(1, 1001)
if not (i % 7)]
# ## How can we test if a number contains the digit 3?
#
# ## We need to get the representations for the number
#
# str(num) returns a string
'3' in str(123)
for i in range(20):
if ('3' in str(i)):
print(i)
# Starting point
for i in range(1, 100):
if not (i % 7 ):
if '3' in str(i):
print(i)
# ## Rewrite using List Comprehension
#
# Another example of Filtering
[i for i in range(1, 1001)
if not (i % 7) and ('3' in str(i))]
# # Program Practice
# ## Long words begining with 'y'
# ## How many words of 10 letters or more begin with 'y'?
with open('../words.txt', 'r') as words:
result = []
for word in words:
if len(word) >= 10 and word[0] == 'y':
result.append(word)
print(result)
# ## Strip() the newline
with open('../words.txt', 'r') as words:
result = []
for word in words:
if len(word) >= 10 and word[0] == 'y':
result.append(word.strip())
print(result)
# ## Most of these words have 9 letters
# ### We are counting the \n
with open('../words.txt', 'r') as words:
result = []
for word in words:
word = word.strip()
if len(word) >= 10 and word[0] == 'y':
result.append(word)
print(result)
# ## Rewrite as a List Comprehension
with open('../words.txt', 'r') as words:
result = [word.strip() for word in words if len(word) > 10 and word[0] == 'y' ]
print(result)
# # Nested loops
# ## We can nest Loop Comprehensions to get the product
[(word, ch) for word in ['one', 'two', 'three', 'four']
for ch in 'aeiou']
# ## Add a condition: list the words, and only the vowels they contain
[(word, ch) for word in ['one', 'two', 'three', 'four']
for ch in 'aeiou'
if ch in word]
# ## Pythagorean Triples
#
# Looking for $(x, y, z)$ such that
#
# $x^2 + y^2 = z^2$
#
# Restrict it to integers under 30. Show each triple only once
[(x,y,z)
for x in range(1,30)
for y in range(x,30)
for z in range(y,30)
if x**2 + y**2 == z**2]
# ## *OK, maybe that was mathematics*
# # The Danger of a Short List
# +
lst = [1, 2, 3]
# Print the first 10 in the lst
for i in range(10):
print(lst[i])
# -
# ## Ways we could protect ourselves
# - Try-catch IndexError
# - For i in range(min(10, len(lst))):
#
# But there is a simpler way:
#
# ## "Slice is Forgiving"
# +
lst = [1, 2, 3]
# Print the first 10 in the lst
for val in lst[:10]:
print(val)
# -
# ## This allows us to write a one-line Rotation
#
# This works even on an empty list
# +
from typing import List
def rotate(lst: List) -> List:
return lst[1:] + lst[:1]
rotate([])
# -
# # Tuples
# +
t = 1,000,000
print(type(t))
print(f"{t = }")
# +
t = 'a', 'b', 'c', 'd'
print(type(t))
print(f"{t = }")
# -
s = (('a'))
print(type(s))
print(f"{s = }")
# ### Now add a trailing comma
s = 'a',
print(type(s))
print(f"{s = }")
# ### We need the trailing comma to tell that a singleton is a tuple
# +
s = 'a', 'b',
print(type(s))
print(f"{s = }")
# -
# ### We can index and slice
t = 'a', 'b', 'c', 'd'
print(type(t))
print(t[0])
print(t[1:3])
t = tuple('spam')
print(f"{t = }")
print(t[0])
print(t[1:3])
t[3] = 't'
# ## Tuples are immutable. We can use them as keys in a dictionary
# +
d = {(1, 2, 3): 'Let me be me'}
print(f"{d = }")
# -
# ### But we can replace an existing tuple with a new one
print(f"{t = }")
t = ('A',) + t[1:]
print(f"{t = }")
# ### Ordered lexicographically
print((0, 1, 2) < (0, 3, 4))
print((0, 1, 2000) < (0, 3, 4))
# ## Python uses Tuples to return values
for t in zip('ab', 'cd'):
print(f"{t = } {type(t) = }")
for t in enumerate('soap'):
print(f"{t = } {type(t) = }")
# # Tuples most famous party trick
# ### Before
a = 4
b = 1
print(f"{a = } {b = }")
temp = a
a = b
b = temp
print(f"{a = } {b = }")
# ### After, with Tuples
print(f"{a = } {b = }")
a, b = b, a
print(f"{a = } {b = }")
print(f"{a = } {b = }")
(a, b) = (b, a)
print(f"{a = } {b = }")
# ## Parsing e-mail addresses
addr = '<EMAIL>'
uname, domain = addr.split('@')
print(f"{uname = }")
print(f"{domain = }")
# ### We can nest tuples
t = ('a', ('b', 'c', ('d', 'e'), 'f'), 'g')
print(f"{t = }")
print(f"{t[1] = }")
print(f"{t[1][2] = }")
# ### Tuples and lists perform similar roles
#
# - Ordered lists of objects
# - Can nest both types of list
# - We can modify a list: delete, insert, modify
# - Tuples are immutable - can use as keys
#
#
# ## Global Scope
#
# ### All local scope
# +
def second():
val = 1;
print(f"\t\t{val = }")
def first():
val = 3;
print(f"\t{val = }")
second()
print(f"\t{val = }")
# Main program
val = 10
print(f"{val = }")
first()
print(f"{val = }")
# -
# ### Now use Global variable
# +
def second():
# Remove the local definition
# second now sees the global value
# val = 1
print(f"\t\t{val = }")
def first():
val = 3;
print(f"\t{val = }")
second()
print(f"\t{val = }")
# Main program
val = 10
print(f"{val = }")
first()
print(f"{val = }")
# -
# ## Globals considered Harmful
# +
def second():
print(f"\t\t{lst = }")
lst.pop()
print(f"\t\t{lst = }")
def first():
lst = ['a', 'b', 'c'];
print(f"\t{lst = }")
second()
print(f"\t{lst = }")
# Main program
lst = [1, 2, 3]
print(f"{lst = }")
first()
print(f"{lst = }")
# -
# ## Why is this better?
# +
def second(lst):
print(f"\t{lst = }")
lst.pop()
print(f"\t{lst = }")
# Main program
lst = [1, 2, 3]
print(f"{lst = }")
second(lst)
print(f"{lst = }")
# -
# ## It is better because we can identify a possible culprit
# # String Formatting
#
# ## Motivation
# +
shopping = {'milk':1, 'eggs':12, 'bread':2}
for w in shopping:
print(f'{shopping[w]} | {w}')
# -
# ## We would like to line things up
# Include a field width
# Field width of 3
for w in shopping:
print(f'{shopping[w]:3} | {w}')
# Integer in a field of width 3
#
# 3 Ordinary characters - ‘ | ‘
#
# String in variable field - %s
#
# 12 | eggs
# ## Stop and Think
#
# You have a string holding a phone number: 1234567890
#
# Wish to print in standard form: (123) 456-7890
#
# Write with f-string formatting
s = '1234567890'
# # WTF - File Picker
# +
# I have been able to use this in a program - it works well.
# See doublesPick.py
#
from tkinter import Tk
from tkinter.filedialog import askopenfilename
Tk().withdraw() # Don't need a full GUI
filename = askopenfilename() # show an "Open" dialog box
print(f"Filename: {filename}")
| introduction-to-programming-with-python/lectures/cscie7_lecture_07.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
# %matplotlib inline
df = pd.read_csv('kyphosis.csv')
df.head()
df.info()
sns.pairplot(df, hue='Kyphosis')
# + slideshow={"slide_type": "-"}
from sklearn.model_selection import train_test_split
X = df.drop('Kyphosis', axis=1)
y = df['Kyphosis']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3)
# +
from sklearn.tree import DecisionTreeClassifier
dtree = DecisionTreeClassifier()
dtree.fit(X_train, y_train)
# +
predictions = dtree.predict(X_test)
from sklearn.metrics import classification_report, confusion_matrix
print(confusion_matrix(y_test, predictions))
print(classification_report(y_test, predictions))
# +
from sklearn.ensemble import RandomForestClassifier
rfc = RandomForestClassifier(n_estimators=200)
rfc.fit(X_train, y_train)
# +
predictions = rfc.predict(X_test)
print(confusion_matrix(y_test, predictions))
print(classification_report(y_test, predictions))
# -
df['Kyphosis'].value_counts()
| python-for-data-science/Machine Learning/Decision Trees and Random Forests/Decision Trees and Random Forests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
from sklearn.model_selection import train_test_split
import os
import pandas as pd
data_dir = '../dataset'
fname = os.path.join(data_dir, 'data-02-stock_daily.csv')
# +
## 데이터 읽어오기.
df = pd.read_csv(fname)
dataset = df.values
ori_X = dataset[:,0:4]
ori_Y = dataset[:,4]
# -
seq_length=2
X_train, X_test, Y_train, Y_test = train_test_split(ori_X,ori_Y, test_size=0.2, shuffle=False)
data_gen=tf.keras.preprocessing.sequence.TimeseriesGenerator(X_train, Y_train,
length=seq_length, sampling_rate=1,stride=1,
batch_size=1,
start_index=0,reverse=True)
for i in range(1):
x, y = data_gen[i]
print(x, y)
| tensorflow/day6/practice/.ipynb_checkpoints/P_05_00_time_data-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
a = np.linspace(0,10,11)
b = a ** 4
x = np.arange(0, 10)
y = 2 * x
fig, axes = plt.subplots(nrows=1, ncols=2)
# axes[0].plot(x, y)
type(axes)
axes.shape
fig, axes = plt.subplots(nrows=1, ncols=2)
axes[0].plot(x, y)
# +
fig, axes = plt.subplots(nrows=2, ncols=2)
axes[0][0].plot(x, y)
axes[1][1].plot(a, b)
plt.tight_layout()
# -
| practical_ai/01_matplotlib/matplotlib_subplots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <small><small><i>
# All the IPython Notebooks in this lecture series by Dr. <NAME> are available @ **[GitHub](https://github.com/milaan9/05_Python_Files)**
# </i></small></small>
# # Python File I/O
#
# In this class, you'll learn about Python file operations. More specifically, opening a file, reading from it, writing into it, closing it, and various file methods that you should be aware of.
# ## Files
#
# Files are named locations on disk to store related information. They are used to permanently store data in a non-volatile memory (e.g. hard disk).
#
# Since Random Access Memory (RAM) is volatile (which loses its data when the computer is turned off), we use files for future use of the data by permanently storing them.
#
# When we want to read from or write to a file, we need to open it first. When we are done, it needs to be closed so that the resources that are tied with the file are freed.
#
# Hence, in Python, a file operation takes place in the following order:
# <b>
# 1. Open a file
# 2. Close the file
# 3. Read or write (perform operation)
# </b>
# ## 1. Opening Files in Python
#
# Python has a built-in **`open()`** function to open a file. This function returns a file object, also called a handle, as it is used to read or modify the file accordingly.
#
# ```python
# >>> f = open("test.txt") # open file in current directory
# >>> f = open("C:/Python99/README.txt") # specifying full path
# ```
#
# We can specify the mode while opening a file. In mode, we specify whether we want to read **`r`**, write **`w`** or append **`a`** to the file. We can also specify if we want to open the file in text mode or binary mode.
#
# The default is reading in text mode. In this mode, we get strings when reading from the file.
#
# On the other hand, binary mode returns bytes and this is the mode to be used when dealing with non-text files like images or executable files.
#
# | Mode | Description |
# |:----:| :--- |
# | **`r`** | Opens a file for reading only. The file pointer is placed at the beginning of the file. This is the default mode. |
# | **`t`** | Opens in text mode. (default). |
# | **`b`** | Opens in binary mode. |
# | **`x`** | Opens a file for exclusive creation. If the file already exists, the operation fails. |
# | **`rb`** | Opens a file for reading only in binary format. The file pointer is placed at the beginning of the file. This is the default mode. |
# | **`r+`** | Opens a file for both reading and writing. The file pointer placed at the beginning of the file. |
# | **`rb+`** | Opens a file for both reading and writing in binary format. The file pointer placed at the beginning of the file. |
# | **`w`** | Opens a file for writing only. Overwrites the file if the file exists. If the file does not exist, creates a new file for writing. |
# | **`wb`** | Opens a file for writing only in binary format. Overwrites the file if the file exists. If the file does not exist, creates a new file for writing. |
# | **`w+`** | Opens a file for both writing and reading. Overwrites the existing file if the file exists. If the file does not exist, creates a new file for reading and writing. |
# | **`wb+`** | Opens a file for both writing and reading in binary format. Overwrites the existing file if the file exists. If the file does not exist, creates a new file for reading and writing. |
# | **`a`** | Opens a file for appending. The file pointer is at the end of the file if the file exists. That is, the file is in the append mode. If the file does not exist, it creates a new file for writing. |
# | **`ab`** | Opens a file for appending in binary format. The file pointer is at the end of the file if the file exists. That is, the file is in the append mode. If the file does not exist, it creates a new file for writing. |
# | **`a+`** | Opens a file for both appending and reading. The file pointer is at the end of the file if the file exists. The file opens in the append mode. If the file does not exist, it creates a new file for reading and writing. |
# | **`ab+`** | Opens a file for both appending and reading in binary format. The file pointer is at the end of the file if the file exists. The file opens in the append mode. If the file does not exist, it creates a new file for reading and writing. |
f = open("test.txt") # equivalent to 'r' or 'rt'
f = open("test.txt",'w') # write in text mode
f = open("logo.png",'r+b') # read and write in binary mode
# Unlike other languages, the character **`a`** does not imply the number 97 until it is encoded using **`ASCII`** (or other equivalent encodings).
#
# Moreover, the default encoding is platform dependent. In windows, it is **`cp1252`** but **`utf-8`** in Linux.
#
# So, we must not also rely on the default encoding or else our code will behave differently in different platforms.
#
# Hence, when working with files in text mode, it is highly recommended to specify the encoding type.
f = open("test.txt", mode='r', encoding='utf-8')
# ## 2. Closing Files in Python
#
# When we are done with performing operations on the file, we need to properly close the file.
#
# Closing a file will free up the resources that were tied with the file. It is done using the **`close()`** method available in Python.
#
# Python has a garbage collector to clean up unreferenced objects but we must not rely on it to close the file.
f = open("test.txt", encoding = 'utf-8')
# perform file operations
f.close()
# This method is not entirely safe. If an exception occurs when we are performing some operation with the file, the code exits without closing the file.
#
# A safer way is to use a **[try-finally](https://github.com/milaan9/05_Python_Files/blob/main/004_Python_Exceptions_Handling.ipynb)** block.
try:
f = open("test.txt", encoding = 'utf-8')
# perform file operations
finally:
f.close()
# This way, we are guaranteeing that the file is properly closed even if an exception is raised that causes program flow to stop.
#
# The best way to close a file is by using the **`with`** statement. This ensures that the file is closed when the block inside the **`with`** statement is exited.
#
# We don't need to explicitly call the **`close()`** method. It is done internally.
#
# ```python
# >>>with open("test.txt", encoding = 'utf-8') as f:
# # perform file operations
# ```
# ### The file Object Attributes
#
# * **file.closed** - Returns true if file is closed, false otherwise.
# * **file.mode** - Returns access mode with which file was opened.
# * **file.name** - Returns name of the file.
# Open a file
data = open("data.txt", "wb")
print ("Name of the file: ", data.name)
print ("Closed or not : ", data.closed)
print ("Opening mode : ", data.mode)
data.close() #closed data.txt file
# ## 3. Writing to Files in Python
#
# In order to write into a file in Python, we need to open it in write **`w`**, append **`a`** or exclusive creation **`x`** mode.
#
# We need to be careful with the **`w`** mode, as it will overwrite into the file if it already exists. Due to this, all the previous data are erased.
#
# Writing a string or sequence of bytes (for binary files) is done using the **`write()`** method. This method returns the number of characters written to the file.
with open("test_1.txt",'w',encoding = 'utf-8') as f:
f.write("my first file\n")
f.write("This file\n\n")
f.write("contains three lines\n")
# This program will create a new file named **`test_1.txt`** in the current directory if it does not exist. If it does exist, it is overwritten.
#
# We must include the newline characters ourselves to distinguish the different lines.
with open("test_2.txt",'w',encoding = 'utf-8') as f:
f.write("This is file\n")
f.write("my\n")
f.write("first file\n")
# open a file in current directory
data = open("data_1.txt", "w") # "w" write in text mode,
data.write("Welcome to Dr. <NAME>'s Python Tutorial")
print("done")
data.close()
# <div>
# <img src="img/io1.png" width="1000"/>
# </div>
# ## 4. Reading Files in Python
#
# To read a file in Python, we must open the file in reading **`r`** mode.
#
# There are various methods available for this purpose. We can use the **`read(size)`** method to read in the **`size`** number of data. If the **`size`** parameter is not specified, it reads and returns up to the end of the file.
#
# We can read the **`text_1.txt`** file we wrote in the above section in the following way:
f = open("test_1.txt",'r',encoding = 'utf-8')
f.read(8) # read the first 8 data characters
f.read(5) # read the next 5 data characters
f.read() # read in the rest till end of file
f.read() # further reading returns empty sting
# We can see that the **`read()`** method returns a newline as **`'\n'`**. Once the end of the file is reached, we get an empty string on further reading.
#
# We can change our current file cursor (position) using the **`seek()`** method. Similarly, the **`tell()`** method returns our current position (in number of bytes).
f.tell() # get the current file position
f.seek(0) # bring file cursor to initial position
print(f.read()) # read the entire file
# We can read a file line-by-line using a **[for loop](https://github.com/milaan9/03_Python_Flow_Control/blob/main/005_Python_for_Loop.ipynb)**. This is both efficient and fast.
f = open("test_1.txt",'r',encoding = 'utf-8')
for line in f:
print(line, end = '')
# In this program, the lines in the file itself include a newline character **`\n`**. So, we use the end parameter of the **`print()`** function to avoid two newlines when printing.
# Alternatively, we can use the **`readline()`** method to read individual lines of a file. This method reads a file till the newline, including the newline character.
f.seek(0) # bring file cursor to initial position
f.readline()
f.readline()
f.readline()
f.readline()
# Lastly, the **`readlines()`** method returns a list of remaining lines of the entire file. All these reading methods return empty values when the end of file **(EOF)** is reached.
f.seek(0) # bring file cursor to initial position
f.readlines()
# Open a file
data = open("data_1.txt", "r+")
file_data = data.read(27) # read 3.375 byte only
full_data = data.read() # read all byte into file from last cursor
print(file_data)
print(full_data)
data.close()
# ## File Positions
#
# The **`tell()`** method tells you the current position within the file; in other words, the next read or write will occur at that many bytes from the beginning of the file.
#
# The **`seek(offset[, from])`** method changes the current file position. The offset argument indicates the number of bytes to be moved. The from argument specifies the reference position from where the bytes are to be moved.
#
# If from is set to 0, the beginning of the file is used as the reference position. If it is set to 1, the current position is used as the reference position. If it is set to 2 then the end of the file would be taken as the reference position.
# Open a file
data = open("data_1.txt", "r+")
file_data = data.read(27) # read 18 byte only
print("current position after reading 27 byte :",data.tell())
data.seek(0) #here current position set to 0 (starting of file)
full_data = data.read() #read all byte
print(file_data)
print(full_data)
print("position after reading file : ",data.tell())
data.close()
# ## Python File Methods
#
# There are various methods available with the file object. Some of them have been used in the above examples.
#
# Here is the complete list of methods in text mode with a brief description:
#
# | Method | Description |
# |:----| :--- |
# | **`close()`** | Closes an opened file. It has no effect if the file is already closed. |
# | **`detach()`** | Separates the underlying binary buffer from the **`TextIOBase`** and returns it. |
# | **`fileno()`** | Returns an integer number (file descriptor) of the file. |
# | **`flush()`** | Flushes the write buffer of the file stream. |
# | **`isatty()`** | Returns **`True`** if the file stream is interactive. |
# | **`read(n)`** | Reads at most `n` characters from the file. Reads till end of file if it is negative or `None`. |
# | **`readable()`** | Returns **`True`** if the file stream can be read from. |
# | **`readline(n=-1)`** | Reads and returns one line from the file. Reads in at most **`n`** bytes if specified. |
# | **`readlines(n=-1)`** | Reads and returns a list of lines from the file. Reads in at most **`n`** bytes/characters if specified. |
# | **`seek(offset,from=SEEK_SET)`** | Changes the file position to **`offset`** bytes, in reference to `from` (start, current, end). |
# | **`seekable()`** | Returns **`True`** if the file stream supports random access. |
# | **`tell()`** | Returns the current file location. |
# | **`truncate(size=None)`** | Resizes the file stream to **`size`** bytes. If **`size`** is not specified, resizes to current location.. |
# | **`writable()`** | Returns **`True`** if the file stream can be written to. |
# | **`write(s)`** | Writes the string **`s`** to the file and returns the number of characters written.. |
# | **`writelines(lines)`** | Writes a list of **`lines`** to the file.. |
| 001_Python_File_Input_Output.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Stochastic Gradient Descent
# +
# load libraries
import numpy as np
import pandas as pd
from sklearn import datasets
from sklearn.linear_model import LogisticRegression
from sklearn.linear_model import SGDClassifier
from sklearn.model_selection import KFold
from sklearn.metrics import accuracy_score
# +
# Note that the iris dataset is available in sklearn by default.
# This data is also conveniently preprocessed.
iris = datasets.load_iris()
X = iris["data"]
Y = iris["target"]
# set variables
numFolds = 10
kf = KFold(numFolds, shuffle=True, random_state=43)
# These are "Class objects". For each Class, find the AUC through
# Kfold cross validation.
Models = [LogisticRegression, SGDClassifier]
# Added n_iter here
params = [{}, {"loss": "log", "penalty": "l2", 'max_iter':1000}]
# -
# tuning
for param, Model in zip(params, Models):
total = 0
for train_indices, test_indices in kf.split(X):
train_X = X[train_indices, :]
train_Y = Y[train_indices]
test_X = X[test_indices, :]
test_Y = Y[test_indices]
reg = Model(**param)
reg.fit(train_X, train_Y)
predictions = reg.predict(test_X)
total += accuracy_score(test_Y, predictions)
accuracy = total / numFolds
print("Accuracy score of {0}: {1}".format(Model.__name__, accuracy))
import sklearn
sklearn.__version__
| deeplearning/sklearn/dl_python_sklearn_sgd_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # UAVSAR
#
# ```{admonition} Learning Objectives
# - find, visualize, analyze UAVSAR data
# ```
#
# Intro slide deck: https://uavsar.jpl.nasa.gov/education/what-is-uavsar.html
# ## Quick facts
#
# | frequency | resolution (rng x azi m) | swath width (km) |
# | - | - | - |
# | L-band | 1.8 | 16 |
#
# Documentation:
# * https://uavsar.jpl.nasa.gov/science/documents.html
# * https://asf.alaska.edu/data-sets/sar-data-sets/uavsar/
#
# Data Access:
#
# * [NASA Earthdata Suborbital Search](https://search.earthdata.nasa.gov/portal/suborbital/search?fi=UAVSAR&as[instrument][0]=UAVSAR)
# * [ASF Vertex SnowEx Grand Mesa Campaign]
#
# `https://search.asf.alaska.edu/#/?dataset=UAVSAR&mission=Grand%20Mesa,%20CO&resultsLoaded=true&granule=UA_grmesa_27416_21019-017_21021-005_0006d_s01_L090_01-AMPLITUDE_GRD&zoom=3¢er=-92.747866,10.530273&productTypes=AMPLITUDE_GRD`
# ## API data access
| book/tutorials/sar/uavsar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <p align="center">
# <img src="https://github.com/GeostatsGuy/GeostatsPy/blob/master/TCG_color_logo.png?raw=true" width="220" height="240" />
#
# </p>
#
# ## Interactive Confidence Interval Demonstration
#
# ### Boostrap and Analytical Confidence Intervals
#
# * we calculate the confidence interval in the mean with boostrap and compare to the analytical expression
#
# * with this workflow we all provide an interactive plot demonstration with matplotlib and ipywidget packages
#
# #### <NAME>, Associate Professor, University of Texas at Austin
#
# ##### [Twitter](https://twitter.com/geostatsguy) | [GitHub](https://github.com/GeostatsGuy) | [Website](http://michaelpyrcz.com) | [GoogleScholar](https://scholar.google.com/citations?user=QVZ20eQAAAAJ&hl=en&oi=ao) | [Book](https://www.amazon.com/Geostatistical-Reservoir-Modeling-Michael-Pyrcz/dp/0199731446) | [YouTube](https://www.youtube.com/channel/UCLqEr-xV-ceHdXXXrTId5ig) | [LinkedIn](https://www.linkedin.com/in/michael-pyrcz-61a648a1) | [GeostatsPy](https://github.com/GeostatsGuy/GeostatsPy)
#
# #### Confidence Intevals
#
# Confidence intervals are the uncertainty in a sample statistic or model parameter
#
# * for uncertainty in the sample mean we have:
#
# * center on the sample proportion, $\hat{p}$
#
# * the standard error in the proportion for the dispersion (spread)
#
# * Student's t distributed for small samples and Gaussian distributed for large sample sizes
#
# The analytical form for small samples ($n \lt 30$) is:
#
# \begin{equation}
# CI: \hat{p} \pm t_{\frac{\alpha}{2},n-1} \times \frac {\sqrt{p(1-p)}}{\sqrt{n}}
# \end{equation}
#
# where the sampling distribution of the proportion is student's t distributed with number of samples, $n$ - 1, degrees of freedom and $\alpha$ is the signficance level divded by 2 for the two tails.
#
# When the number of samples is large ($n \ge 30$) then the analytical form converges to Gaussian distributed:
#
# \begin{equation}
# CI: \hat{p} \pm N_{\frac{\alpha}{2}} \times \frac {\sqrt{p(1-p)}}{\sqrt{n}}
# \end{equation}
#
# #### Bootstrap
#
# Uncertainty in the sample statistics
# * one source of uncertainty is the paucity of data.
# * do 200 or even less wells provide a precise (and accurate estimate) of the mean? standard deviation? skew? P13?
#
# Would it be useful to know the uncertainty in these statistics due to limited sampling?
# * what is the impact of uncertainty in the mean porosity e.g. 20%+/-2%?
#
# **Bootstrap** is a method to assess the uncertainty in a sample statistic by repeated random sampling with replacement.
#
# Assumptions
# * sufficient, representative sampling, identical, idependent samples
#
# Limitations
# 1. assumes the samples are representative
# 2. assumes stationarity
# 3. only accounts for uncertainty due to too few samples, e.g. no uncertainty due to changes away from data
# 4. does not account for boundary of area of interest
# 5. assumes the samples are independent
# 6. does not account for other local information sources
#
# The Bootstrap Approach (Efron, 1982)
#
# Statistical resampling procedure to calculate uncertainty in a calculated statistic from the data itself.
# * Does this work? Prove it to yourself, for uncertainty in the mean solution is standard error:
#
# \begin{equation}
# \sigma^2_\overline{x} = \frac{\sigma^2_s}{n}
# \end{equation}
#
# Extremely powerful - could calculate uncertainty in any statistic! e.g. P13, skew etc.
# * Would not be possible access general uncertainty in any statistic without bootstrap.
# * Advanced forms account for spatial information and sampling strategy (game theory and Journel’s spatial bootstrap (1993).
#
# Steps:
#
# 1. assemble a sample set, must be representative, reasonable to assume independence between samples
#
# 2. optional: build a cumulative distribution function (CDF)
# * may account for declustering weights, tail extrapolation
# * could use analogous data to support
#
# 3. For $\ell = 1, \ldots, L$ realizations, do the following:
#
# * For $i = \alpha, \ldots, n$ data, do the following:
#
# * Draw a random sample with replacement from the sample set or Monte Carlo simulate from the CDF (if available).
#
# 6. Calculate a realization of the sammary statistic of interest from the $n$ samples, e.g. $m^\ell$, $\sigma^2_{\ell}$. Return to 3 for another realization.
#
# 7. Compile and summarize the $L$ realizations of the statistic of interest.
#
# This is a very powerful method. Let's try it out and compare the result to the analytical form of the confidence interval for the sample mean.
#
#
# #### Objective
#
# Provide an example and demonstration for:
#
# 1. interactive plotting in Jupyter Notebooks with Python packages matplotlib and ipywidgets
# 2. provide an intuitive hands-on example of confidence intervals and compare to statistical boostrap
#
# #### Getting Started
#
# Here's the steps to get setup in Python with the GeostatsPy package:
#
# 1. Install Anaconda 3 on your machine (https://www.anaconda.com/download/).
# 2. Open Jupyter and in the top block get started by copy and pasting the code block below from this Jupyter Notebook to start using the geostatspy functionality.
#
# #### Load the Required Libraries
#
# The following code loads the required libraries.
# %matplotlib inline
from ipywidgets import interactive # widgets and interactivity
from ipywidgets import widgets
from ipywidgets import Layout
from ipywidgets import Label
from ipywidgets import VBox, HBox
import matplotlib.pyplot as plt # plotting
from matplotlib.ticker import (MultipleLocator, AutoMinorLocator) # control of axes ticks
plt.rc('axes', axisbelow=True) # set axes and grids in the background for all plots
import numpy as np # working with arrays
import pandas as pd # working with DataFrames
import seaborn as sns # for matrix scatter plots
from scipy.stats import triang # parametric distributions
from scipy.stats import binom
from scipy.stats import norm
from scipy.stats import uniform
from scipy.stats import triang
from scipy.stats import t
from scipy import stats # statistical calculations
import random # random drawing / bootstrap realizations of the data
from matplotlib.gridspec import GridSpec # nonstandard subplots
import math # square root operator
# #### Make a Synthetic Dataset
#
# This is an interactive method to:
#
# * select a parametric distribution
#
# * select the distribution parameters
#
# * select the number of samples and visualize the synthetic dataset distribution
# +
# parameters for the synthetic dataset
bins = np.linspace(0,1000,1000)
# interactive calculation of the sample set (control of source parametric distribution and number of samples)
l = widgets.Text(value=' Simple Boostrap Demonstration, <NAME>, Associate Professor, The University of Texas at Austin',layout=Layout(width='950px', height='30px'))
a = widgets.IntSlider(min=0, max = 100, value = 2, step = 1, description = '$n_{red}$',orientation='horizontal',layout=Layout(width='400px', height='20px'),continuous_update=False)
a.style.handle_color = 'red'
b = widgets.IntSlider(min=0, max = 100, value = 3, step = 1, description = '$n_{green}$',orientation='horizontal',layout=Layout(width='400px', height='20px'),continuous_update=False)
b.style.handle_color = 'green'
c = widgets.IntSlider(min=1, max = 16, value = 3, step = 1, description = '$L$',orientation='horizontal',layout=Layout(width='400px', height='20px'),continuous_update=False)
c.style.handle_color = 'gray'
ui = widgets.HBox([a,b,c],) # basic widget formatting
ui2 = widgets.VBox([l,ui],)
def f_make(a, b, c): # function to take parameters, make sample and plot
red_freq = make_data(a, b, c)
labels = ['Red', 'Green']
nrows = np.round(np.sqrt(c)+0.4,0); ncols = np.round(c / nrows + 0.4,0)
plt.clf()
for i in range(0, c):
plt.subplot(ncols,nrows,i + 1)
draw = [red_freq[i],a + b - red_freq[i]]
plt.grid(zorder=0, color='black', axis = 'y', alpha = 0.2); plt.ylim(0,a + b);
plt.ylabel('Frequency'); plt.xlabel('Balls Drawn')
plt.yticks(np.arange(0,a + b + 1,max(1,round((a+b)/10))))
barlist = plt.bar(labels,draw,edgecolor = "black",linewidth = 1,alpha = 0.8); plt.title('Realization #' + str(i+1),zorder = 1)
barlist[0].set_color('r'); barlist[1].set_color('g')
plt.subplots_adjust(left=0.0, bottom=0.0, right=2.0, top=1.2 * nrows, wspace=0.2, hspace=0.2)
plt.show()
def make_data(a, b, c): # function to check parameters and make sample
prop_red = np.zeros(c)
for i in range(0, c):
prop_red[i] = np.random.multinomial(a+b,[a/(a+b),b/(a+b)], size = 1)[0][0]
return prop_red
# connect the function to make the samples and plot to the widgets
interactive_plot = widgets.interactive_output(f_make, {'a': a, 'b': b, 'c': c})
interactive_plot.clear_output(wait = True) # reduce flickering by delaying plot updating
# -
# ### Simple Bootstrap Demonstration - Drawing Red and Green Balls from a Virtual Cowboy Hat
#
# * drawing red and green balls from a hat with replacement to access uncertainty in the proportion
#
# * interactive plot demonstration with ipywidget, matplotlib packages
#
# #### <NAME>, Associate Professor, University of Texas at Austin
#
# ##### [Twitter](https://twitter.com/geostatsguy) | [GitHub](https://github.com/GeostatsGuy) | [Website](http://michaelpyrcz.com) | [GoogleScholar](https://scholar.google.com/citations?user=QVZ20eQAAAAJ&hl=en&oi=ao) | [Book](https://www.amazon.com/Geostatistical-Reservoir-Modeling-Michael-Pyrcz/dp/0199731446) | [YouTube](https://www.youtube.com/channel/UCLqEr-xV-ceHdXXXrTId5ig) | [LinkedIn](https://www.linkedin.com/in/michael-pyrcz-61a648a1) | [GeostatsPy](https://github.com/GeostatsGuy/GeostatsPy)
#
# ### The Problem
#
# Let's simulate bootstrap, resampling with replacement from a hat with $n_{red}$ and $n_{green}$ balls
#
# * **$n_{red}$**: number of red balls in the sample (placed in the hat)
#
# * **$n_{green}$**: number of green balls in the sample (placed in the hat)
#
# * **$L$**: number of bootstrap realizations
display(ui2, interactive_plot) # display the interactive plot
# #### Summarizing Bootstrap Uncertainty
#
# * Run more bootstrap realizations and evaluate the uncertianty model
#
# Now instead of looking at each bootstrap result, let's make many and summarize with:
#
# * **box and whisker plot** of the red and green ball frequencies
#
# * **histograms** of the red and green ball frequencies.
# +
# parameters for the synthetic dataset
bins = np.linspace(0,1000,1000)
# interactive calculation of the sample set (control of source parametric distribution and number of samples)
l2 = widgets.Text(value=' Confidence Interval for Proportions, Analytical and Bootstrap Demonstration, <NAME>, Associate Professor, The University of Texas at Austin',layout=Layout(width='950px', height='30px'))
a2 = widgets.IntSlider(min=0, max = 100, value = 20, step = 1, description = '$n_{red}$',orientation='horizontal',layout=Layout(width='400px', height='20px'),continuous_update=False)
a2.style.handle_color = 'red'
b2 = widgets.IntSlider(min=0, max = 100, value = 30, step = 1, description = '$n_{green}$',orientation='horizontal',layout=Layout(width='400px', height='20px'),continuous_update=False)
b2.style.handle_color = 'green'
c2 = widgets.IntSlider(min=5, max = 1000, value = 1000, step = 1, description = '$L$',orientation='horizontal',layout=Layout(width='400px', height='20px'),continuous_update=False)
c2.style.handle_color = 'gray'
alpha = widgets.FloatSlider(min=0.01, max = 0.40, value = 0.05, step = 0.01, description = r'$\alpha$',orientation='horizontal',layout=Layout(width='400px', height='20px'),continuous_update=False)
alpha.style.handle_color = 'gray'
uib = widgets.HBox([a2,b2,c2,alpha],) # basic widget formatting
uib2 = widgets.VBox([l2,uib],)
def s_make(a, b, c, alpha): # function to take parameters, make sample and plot
dof = a + b - 1
red_freq = make_data(a, b, c)
pred = red_freq/(a+b)
red_prop = (a / (a+b))
red_SE = math.sqrt((red_prop * (1.0 - red_prop)) / (a+b))
green_freq = (a + b) - red_freq
pgreen = green_freq/(a+b)
green_prop = (b / (a+b))
green_SE = math.sqrt((green_prop * (1.0 - green_prop)) / (a+b))
prop_red = red_freq / (a + b)
prop_green = green_freq / (a + b)
labels = ['Red Balls', 'Green Balls']
bins = np.linspace(0,a + b, a + b)
fig = plt.figure(constrained_layout=False)
gs = GridSpec(3, 2, figure=fig)
ax1 = fig.add_subplot(gs[:, 0])
boxplot = ax1.boxplot([pred,pgreen],labels = labels, notch = True, sym = '+',patch_artist=True)
colors = ['red','green']
for patch, color in zip(boxplot['boxes'], colors):
patch.set_facecolor(color)
for patch, color in zip(boxplot['medians'], colors):
patch.set_color('black')
ax1.set_ylim([0,1])
ax1.grid(zorder=0, color='black', axis = 'y', alpha = 0.2)
ax1.set_ylabel('Proportion of Balls'); ax1.set_xlabel('Ball Color');ax1.set_title('Bootstrap Uncertainty - Proportion Distributions')
ax1.grid(True, which='major',axis='y',linewidth = 1.0); ax1.grid(True, which='minor',axis='y',linewidth = 0.2) # add y grids
ax1.tick_params(which='major',length=7); ax1.tick_params(which='minor', length=4)
ax1.xaxis.set_minor_locator(AutoMinorLocator()); ax1.yaxis.set_minor_locator(AutoMinorLocator()) # turn on minor ticks
cumul_prob = np.linspace(0.0,1.0,100)
if a <= 30 or b <= 30:
red_prop_values = t.ppf(cumul_prob, dof)
red_lower = t.ppf(alpha/2, dof); red_upper = t.ppf(1-alpha/2, dof)
else:
red_prop_values = norm.ppf(cumul_prob)
red_lower = norm.ppf(alpha/2); red_upper = norm.ppf(1-alpha/2)
red_prop_values = red_prop_values * red_SE + red_prop
red_lower = red_lower * red_SE + red_prop
red_upper = red_upper * red_SE + red_prop
cumul_prob = np.linspace(0.01,0.99,100)
if a <= 30 or b <= 30:
green_prop_values = t.ppf(cumul_prob, dof)
green_lower = t.ppf(alpha/2, dof); green_upper = t.ppf(1-alpha/2, dof)
else:
green_prop_values = norm.ppf(cumul_prob)
green_lower = norm.ppf(alpha/2); green_upper = norm.ppf(1-alpha/2)
green_prop_values = green_prop_values * green_SE + green_prop
green_lower = green_lower * green_SE + green_prop
green_upper = green_upper * green_SE + green_prop
ax2 = fig.add_subplot(gs[0, 1])
ax2.hist(prop_red,cumulative = True, density = True, alpha=0.7,color="red",edgecolor="black",linewidth=2,bins = np.linspace(0,1,50), label = 'Bootstrap')
ax2.plot([red_lower,red_lower],[0,1],color='black',linewidth=2,linestyle='--',label='Lower/Upper')
ax2.plot([red_upper,red_upper],[0,1],color='black',linewidth=2,linestyle='--')
ax2.plot([red_prop,red_prop],[0,1],color='black',linewidth=3,label='Exp.')
ax2.set_title('Uncertainty in Proportion of Red Balls'); ax2.set_xlabel('Proportion of Red Balls'); ax2.set_ylabel('Cumulative Probability')
ax2.set_xlim([0,1]); ax2.set_ylim([0,1])
ax2.plot(red_prop_values, cumul_prob, color = 'black', linewidth = 2, label = 'Analytical')
ax2.legend()
ax3 = fig.add_subplot(gs[1, 1])
ax3.hist(prop_green,cumulative = True, density = True, alpha=0.7,color="green",edgecolor="black",linewidth=2,bins = np.linspace(0,1,50), label = 'Bootstrap')
ax3.plot([green_lower,green_lower],[0,1],color='black',linewidth=2,linestyle='--',label='Lower/Upper')
ax3.plot([green_upper,green_upper],[0,1],color='black',linewidth=2,linestyle='--')
ax3.plot([green_prop,green_prop],[0,1],color='black',linewidth=3,label='Exp.')
ax3.set_title('Uncertainty in Proportion of Green Balls'); ax3.set_xlabel('Proportion of Green Balls'); ax3.set_ylabel('Cumulative Probability')
ax3.set_xlim([0,1]); ax3.set_ylim([0,1])
ax3.plot(green_prop_values, cumul_prob, color = 'black', linewidth = 2, label = 'Analytical')
ax3.legend()
ax4 = fig.add_subplot(gs[2, 1])
ax4.hist(prop_green,cumulative = False, density = True, alpha=0.7,color="green",edgecolor="black",linewidth=2, bins = np.linspace(0,1,50), label = 'Bootstrap Prop. Green')
ax4.hist(prop_red,cumulative = False, density = True, alpha=0.7,color="red",edgecolor="black",linewidth=2, bins = np.linspace(0,1,50), label = 'Bootstrap Prop. Red')
ax4.set_title('Confidence Interval in Proportion of Red and Green Balls (Alpha = ' + str(alpha) + ')'); ax3.set_xlabel('Proportion of Green Balls')
ax4.set_xlabel('Proportion of Red and Green Balls'); ax4.set_ylabel('Frequency')
ax4.set_xlim([0,1])
prop_values = np.linspace(0.0,1.0,100)
if a <= 30 and b <= 30:
green_density = t.pdf(prop_values,loc = green_prop, df = dof, scale = green_SE)
else:
green_density = norm.pdf(prop_values,loc = green_prop, scale = green_SE)
ax4.plot(prop_values, green_density, color = 'black', linewidth = 5,zorder=99)
ax4.plot(prop_values, green_density, color = 'green', linewidth = 3, label = 'Analytical Prop. Green',zorder=100)
if a <= 30 and b <= 30:
red_density = t.pdf(prop_values,loc = red_prop, df = dof, scale = red_SE)
else:
red_density = norm.pdf(prop_values,loc = red_prop, scale = red_SE)
ax4.plot(prop_values, red_density, color = 'black', linewidth = 5,zorder=99)
ax4.plot(prop_values, red_density, color = 'red', linewidth = 3, label = 'Analytical Prop. Red',zorder=100)
ax4.fill_between(prop_values, 0, green_density, where = prop_values <= green_lower, facecolor='green', interpolate=True, alpha = 0.9,zorder=101)
ax4.fill_between(prop_values, 0, green_density, where = prop_values >= green_upper, facecolor='green', interpolate=True, alpha = 0.9,zorder=101)
ax4.fill_between(prop_values, 0, red_density, where = prop_values <= red_lower, facecolor='darkred', interpolate=True, alpha = 0.9,zorder=101)
ax4.fill_between(prop_values, 0, red_density, where = prop_values >= red_upper, facecolor='darkred', interpolate=True, alpha = 0.9,zorder=101)
ax4.legend()
plt.subplots_adjust(left=0.0, bottom=0.0, right=2.5, top=3.0, wspace=0.2, hspace=0.3)
plt.show()
# connect the function to make the samples and plot to the widgets
interactive_plot = widgets.interactive_output(s_make, {'a': a2, 'b': b2, 'c': c2, 'alpha': alpha})
interactive_plot.clear_output(wait = True) # reduce flickering by delaying plot updating
# -
# ### Simple Bootstrap and Analytical Confidence Interval Demonstration for Sample Proportions
#
# * drawing red and green balls from a hat with replacement to access uncertainty in the proportion
#
# * run many bootstrap realizations and summarize the results and compare to the analytical sampling distribution for the proportion
#
# * interactive plot demonstration with ipywidget, matplotlib packages
#
# #### <NAME>, Associate Professor, The University of Texas at Austin
#
# ##### [Twitter](https://twitter.com/geostatsguy) | [GitHub](https://github.com/GeostatsGuy) | [Website](http://michaelpyrcz.com) | [GoogleScholar](https://scholar.google.com/citations?user=QVZ20eQAAAAJ&hl=en&oi=ao) | [Book](https://www.amazon.com/Geostatistical-Reservoir-Modeling-Michael-Pyrcz/dp/0199731446) | [YouTube](https://www.youtube.com/channel/UCLqEr-xV-ceHdXXXrTId5ig) | [LinkedIn](https://www.linkedin.com/in/michael-pyrcz-61a648a1) | [GeostatsPy](https://github.com/GeostatsGuy/GeostatsPy)
#
# ### The Problem
#
# Let's simulate bootstrap, resampling with replacement from a hat with $n_{red}$ and $n_{green}$ balls
#
# * **$n_{red}$**: number of red balls in the sample (placed in the hat)
#
# * **$n_{green}$**: number of green balls in the sample (placed in the hat)
#
# * **$L$**: number of bootstrap realizations
#
# * **$\alpha$**: alpha level for the confidence interval (significance level)
#
# and then compare the uncertainty in the proportion of balls to the analytical expression.
# ### Confidence Interval Demonstration for Sample Proportions, Analytical and Bootstrap
#
# #### <NAME>, Associate Professor, University of Texas at Austin
#
# ##### [Twitter](https://twitter.com/geostatsguy) | [GitHub](https://github.com/GeostatsGuy) | [Website](http://michaelpyrcz.com) | [GoogleScholar](https://scholar.google.com/citations?user=QVZ20eQAAAAJ&hl=en&oi=ao) | [Book](https://www.amazon.com/Geostatistical-Reservoir-Modeling-Michael-Pyrcz/dp/0199731446) | [YouTube](https://www.youtube.com/channel/UCLqEr-xV-ceHdXXXrTId5ig) | [LinkedIn](https://www.linkedin.com/in/michael-pyrcz-61a648a1) | [GeostatsPy](https://github.com/GeostatsGuy/GeostatsPy)
display(uib2, interactive_plot) # display the interactive plot
# #### Observations
#
# Some observations:
#
# * sampling distribution for proportions become discrete with too few samples, as only $n$ cases are possible
#
# * enough bootstrap realizations are required for stable statistics
#
# * the analytical sampling distribution for the uncertainty in the sample proportion matches the results from bootstrap
#
#
# #### Comments
#
# This was a simple demonstration of interactive plots in Jupyter Notebook Python with the ipywidgets and matplotlib packages.
#
# I have many other demonstrations on data analytics and machine learning, e.g. on the basics of working with DataFrames, ndarrays, univariate statistics, plotting data, declustering, data transformations, trend modeling and many other workflows available at https://github.com/GeostatsGuy/PythonNumericalDemos and https://github.com/GeostatsGuy/GeostatsPy.
#
# I hope this was helpful,
#
# *Michael*
#
# #### The Author:
#
# ### <NAME>, Associate Professor, University of Texas at Austin
# *Novel Data Analytics, Geostatistics and Machine Learning Subsurface Solutions*
#
# With over 17 years of experience in subsurface consulting, research and development, Michael has returned to academia driven by his passion for teaching and enthusiasm for enhancing engineers' and geoscientists' impact in subsurface resource development.
#
# For more about Michael check out these links:
#
# #### [Twitter](https://twitter.com/geostatsguy) | [GitHub](https://github.com/GeostatsGuy) | [Website](http://michaelpyrcz.com) | [GoogleScholar](https://scholar.google.com/citations?user=QVZ20eQAAAAJ&hl=en&oi=ao) | [Book](https://www.amazon.com/Geostatistical-Reservoir-Modeling-Michael-Pyrcz/dp/0199731446) | [YouTube](https://www.youtube.com/channel/UCLqEr-xV-ceHdXXXrTId5ig) | [LinkedIn](https://www.linkedin.com/in/michael-pyrcz-61a648a1)
#
# #### Want to Work Together?
#
# I hope this content is helpful to those that want to learn more about subsurface modeling, data analytics and machine learning. Students and working professionals are welcome to participate.
#
# * Want to invite me to visit your company for training, mentoring, project review, workflow design and / or consulting? I'd be happy to drop by and work with you!
#
# * Interested in partnering, supporting my graduate student research or my Subsurface Data Analytics and Machine Learning consortium (co-PIs including Profs. Foster, Torres-Verdin and van Oort)? My research combines data analytics, stochastic modeling and machine learning theory with practice to develop novel methods and workflows to add value. We are solving challenging subsurface problems!
#
# * I can be reached at <EMAIL>exas.edu.
#
# I'm always happy to discuss,
#
# *Michael*
#
# <NAME>, Ph.D., P.Eng. Associate Professor The Hildebrand Department of Petroleum and Geosystems Engineering, Bureau of Economic Geology, The Jackson School of Geosciences, The University of Texas at Austin
#
# #### More Resources Available at: [Twitter](https://twitter.com/geostatsguy) | [GitHub](https://github.com/GeostatsGuy) | [Website](http://michaelpyrcz.com) | [GoogleScholar](https://scholar.google.com/citations?user=QVZ20eQAAAAJ&hl=en&oi=ao) | [Book](https://www.amazon.com/Geostatistical-Reservoir-Modeling-Michael-Pyrcz/dp/0199731446) | [YouTube](https://www.youtube.com/channel/UCLqEr-xV-ceHdXXXrTId5ig) | [LinkedIn](https://www.linkedin.com/in/michael-pyrcz-61a648a1)
#
| Interactive_Confidence_Interval.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# import relevant modules
import pandas as pd
# read grades dataset, save as a pandas dataframe
grades = pd.read_csv('grades.csv')
# display first few rows of grades
grades.head()
def mean_atleast_70(student_id):
"""Compute mean grade across all exams for student with given student_id.
Treat missing exam grades as zeros.
If mean grade is atleast 70, return True. Otherwise, return False."""
mean_grade = grades.loc[grades['student_id'] == student_id]['grade'].fillna(0).mean()
return mean_grade >= 70
# test mean_grade on student_id 1
assert mean_atleast_70(1) == False, 'test failed'
print('test passed')
# sequence containing all distinct student ids
student_ids = grades['student_id'].unique()
student_ids
list(filter(mean_atleast_70, student_ids))
| Ex_Files_Python_Data_Functions/Exercise Files/02_03_filter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# +
from fastai.imports import *
from fastai.structured import *
from pandas_summary import DataFrameSummary
from sklearn.ensemble import RandomForestRegressor, RandomForestClassifier
from IPython.display import display
from sklearn import metrics
# -
PATH = os.getcwd();
PATH
df_raw = pd.read_csv(f'{PATH}\\av_train_DaEJRFg.csv', low_memory=False, parse_dates=['incident_date'])
df_raw.head(1)
df_raw.criticality.value_counts()
df_raw.head(3)
add_datepart(test, 'incident_date')
test.head()
test.drop('incident_Elapsed', axis=1, inplace=True)
test['incident_location'].fillna('Dehradun',inplace=True)
train_cats(test)
os.makedirs('tmp', exist_ok=True)
df_raw.to_feather('tmp/av_rookree_raw')
test.head(2)
test.info()
y = df_raw.criticality
df_raw.drop('criticality',axis=1,inplace=True)
from sklearn.model_selection import train_test_split
X_train, X_validation, y_train, y_validation = train_test_split(df_raw, y, train_size=0.8, random_state=1234)
df_raw.info()
categorical_features_indices = np.where(df_raw.dtypes != (np.int64 or np.bool))[0]
categorical_features_indices
#importing library and building model
from catboost import CatBoostClassifier
model=CatBoostClassifier(iterations=1000, depth=13, learning_rate=0.01, loss_function='CrossEntropy',\
)
model.fit(X_train, y_train,cat_features=categorical_features_indices,eval_set=(X_validation, y_validation))
test = pd.read_csv(f'{PATH}\\av_test_TQDFDgg.csv', low_memory=False, parse_dates=['incident_date'])
test.head(3)
prediction_proba = model.predict_proba(test)
model.get_feature_importance(X_train,y_train,cat_features=categorical_features_indices)
prediction_proba[:,1]
def make_submission(probs):
sample = pd.read_csv(f'{PATH}//av_sample_submission_n2Tyn0h.csv')
submit = sample.copy()
submit['criticality'] = probs
return submit
submit = make_submission(prediction_proba[:,1])
submit.head(2)
submit.to_csv(PATH + 'av_cat_2.csv', index=False)
sub1 = pd.read_csv(PATH + 'av_cat_.csv')
sub2 = pd.read_csv(PATH + 'av_cat_2.csv')
sub3 = pd.read_csv(PATH)
sub1.head(1)
sub2.head(2)
pred1 = sub1['criticality'];
pred2 = sub2['criticality'];
pred3 = (.4*pred1 + .6*pred2)/2.
sub1['criticality'] = pred3
sub1.to_csv(PATH + 'av_cat_1_2_.csv', index=False)
# + [markdown] heading_collapsed=true
# ### gridcv
# + hidden=true
from sklearn.ensemble import GradientBoostingClassifier #GBM algorithm
from sklearn import cross_validation, metrics #Additional scklearn functions
from sklearn.grid_search import GridSearchCV #Perforing grid search
# + hidden=true
def modelfit(alg, dtrain, predictors, performCV=True, printFeatureImportance=True, cv_folds=5):
#Fit the algorithm on the data
alg.fit(dtrain[predictors], y)
#Predict training set:
dtrain_predictions = alg.predict(dtrain[predictors])
dtrain_predprob = alg.predict_proba(dtrain[predictors])[:,1]
#Perform cross-validation:
if performCV:
cv_score = cross_validation.cross_val_score(alg, dtrain[predictors], y, cv=cv_folds, scoring='roc_auc')
#Print model report:
print ("\nModel Report")
print ("Accuracy : %.4g" % metrics.accuracy_score(y , dtrain_predictions))
print ("AUC Score (Train): %f" % metrics.roc_auc_score(y , dtrain_predprob))
if performCV:
print ("CV Score : Mean - %.7g | Std - %.7g | Min - %.7g | Max - %.7g" % (np.mean(cv_score),np.std(cv_score),np.min(cv_score),np.max(cv_score)))
#Print Feature Importance:
if printFeatureImportance:
feat_imp = pd.Series(alg.feature_importances_, predictors).sort_values(ascending=False)
plt.figure(figsize=(20,20))
feat_imp.plot(kind='bar', title='Feature Importances')
plt.ylabel('Feature Importance Score')
# + hidden=true
#Choose all predictors except target & IDcols
predictors = df.columns
gbm0 = GradientBoostingClassifier(random_state=10)
modelfit(gbm0, df, predictors)
# + hidden=true
param_test1 = {'n_estimators':[20, 30, 40, 50, 60, 70, 80, 90]}
gsearch1 = GridSearchCV(estimator = GradientBoostingClassifier(learning_rate=0.05, min_samples_split=500,
min_samples_leaf=50,max_depth=8,max_features='sqrt',subsample=0.8,random_state=10),
param_grid = param_test1, scoring='roc_auc',n_jobs=4,iid=False, cv=5)
gsearch1.fit(df[predictors], y)
# + hidden=true
gsearch1.grid_scores_, gsearch1.best_params_, gsearch1.best_score_
# + hidden=true
## Test 2
param_test2 = {'max_depth':[5, 7, 9, 11, 13, 15] ,'min_samples_split': [200, 400, 600, 800, 1000]}
gsearch2 = GridSearchCV(estimator = GradientBoostingClassifier(learning_rate=0.05, n_estimators=90, max_features='sqrt', subsample=0.8, random_state=10),
param_grid = param_test2, scoring='roc_auc',n_jobs=4,iid=False, cv=5)
# + hidden=true
gsearch2.fit(df[predictors], y)
gsearch2.grid_scores_, gsearch2.best_params_, gsearch2.best_score_
# + hidden=true
#test 3
param_test3 = {'min_samples_split': [800, 1000, 1200, 1400, 1600] , 'min_samples_leaf': [30, 40, 50, 60, 70]}
gsearch3 = GridSearchCV(estimator = GradientBoostingClassifier(learning_rate=0.05, n_estimators=90,\
max_depth=7,max_features='sqrt', subsample=0.8, random_state=10),
param_grid = param_test3, scoring='roc_auc',n_jobs=4,iid=False, cv=5)
# + hidden=true
gsearch3.fit(df[predictors], y)
# + hidden=true
gsearch3.grid_scores_, gsearch3.best_params_, gsearch3.best_score_
# + hidden=true
modelfit(gsearch3.best_estimator_, df, predictors)
# + hidden=true
#test 4
param_test4 = {'max_features': [7, 9, 11, 13, 15, 17, 19, 21]}
gsearch4 = GridSearchCV(estimator = GradientBoostingClassifier(learning_rate=0.05, min_samples_split = 1000, n_estimators=70,max_depth=7,\
max_features='sqrt', subsample=0.8, random_state=10,min_samples_leaf = 50),
param_grid = param_test4, scoring='roc_auc',n_jobs=4,iid=False, cv=5)
# + hidden=true
gsearch4.fit(df[predictors], y )
gsearch4.grid_scores_, gsearch4.best_params_, gsearch4.best_score_
# + hidden=true
#test 5
param_test5 = {'subsample':[0.6,0.7,0.75,0.8,0.85,0.9]}
gsearch5 = GridSearchCV(estimator = GradientBoostingClassifier(learning_rate=0.05, min_samples_split = 1000, n_estimators=70,max_depth=7,\
subsample=0.8, \
random_state=10,min_samples_leaf = 50,max_features=17),
param_grid = param_test5, scoring='roc_auc',n_jobs=4,iid=False, cv=5)
# + hidden=true
gsearch5.fit(df[predictors], y )
gsearch5.grid_scores_, gsearch5.best_params_, gsearch5.best_score_
# + hidden=true
gbm_tuned_2 = GradientBoostingClassifier(learning_rate=0.05, min_samples_split = 1000, n_estimators=500,max_depth=10,\
subsample=0.8, random_state=10,min_samples_leaf = 50,max_features=17)
modelfit(gbm_tuned_2, df, predictors)
# + hidden=true
test.info()
# + hidden=true
prediction_proba_2 = gbm_tuned_2.predict_proba(test)
# + hidden=true
submit = make_submission(prediction_proba_2[:,1])
| AV/av_rookree.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#Consider the string given below and answer the following questions
string = "<KEY>"
print(len(string))
# #### Calculate the number of characters in the string
#
# - 193
# - 200
# - 300
# - 500
#
#
#
# +
# Type your code I
print(string.count('a'))
# -
# #### Count the number of occurences of 'a' in the string
#
# - 1
# - 2
# - 10
# - 5
# +
# Type your code here
print(string.startswith('if'))
# -
# #### Identify whether the string starts with 'if'.
#
# - True
# - False
# Type your code here
print(string[63:89])
# #### Return the substring starting at index 63 and ending at index 88 [Both the indexes are inclusive]
#
# - 'OQYWFClFhFGA'
# - '<KEY>'
# - '<KEY>'
# - '<KEY>'
# Type your code here
print(string[45])
# #### Identify the character at index 45
#
# - a
# - b
# - z
# - x
# +
# Type your code here
# +
arr = ['Pythons syntax is easy to learn', 'Pythons syntax is very clear']
','.join(arr)
# -
| Data Science/Python/Practice+Exercise+1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Composition
# ### Table of Contents
# - [Objectives](#Objectives)
# - [Composition](#What-is-Composition?)
# - [Advantages and Disadvantages of Inheritance](#Advantages-and-Disadvantages-of-Inheritance)
# - [Advantages and Disadvantages of Composition](#Advantages-and-Disadvantages-of-Composition)
# - [Code Example](#Code-Example)
# - [When to use Inheritance](#When-to-use-Inheritance)
# - [Takeaways](#Takeaways)
# ### Objectives
# * Knowledge of composition.
# * Advantages of composition over inheritance.
# ### What is Composition?
#
# Composition is another Object Oriented programming approach. We use it when we want to use some aspect of another class without promising all of the features of that other class.
# **Inheritance models a is-a relationship**. This means that when you have a Derived class that inherits from a Base class, you created a relationship where Derived is a specialized version of Base.
#
# **Composition models a has-a relationship**. It enables creating complex types by combining objects of other types. This means that a class Composite can contain an object of another class Component. This relationship means that a Composite has a Component.
# ### Object Oriented Design Principle: **Favour Composition Over Inheritance**
# +
# simple example showing inheritance vs composition
# Inheritance (white box reuse)
class Vehicle:
pass
class Bus(Vehicle):
pass
# Composition (black box reuse)
class Engine:
pass
class Bus:
def __init__(self):
self.engine = Engine()
# -
# ### Advantages and Disadvantages of Inheritance
#
# **Advantages**
# - It's well known.
# - Easy way to reuse code.
# - After inheriting, it's possible to add/change/modify the behaviour(s) of the inherited methods.
#
# **Disadvantages**
# - Inheritance supports weak encapsulation.
# - The derived class inherits everything including unneeded or unwanted stuff.
# - Changes in the base class will cause an impact on all the derived classes.
# - In inheritance, OOP can get unneccessarily complicated and full of ambiguity, confusion, and hierarchies.
# ### Advantages and Disadvantages of Composition
#
# **Advantages**
# - Supports great encapsulation.
# - Changes in one class has limited effect on the other classes.
# - In composition, one class(the composite class) can have relationships with many other classes(the component classes).
#
# **Disadvantages**
# - Actually requires more code than Inheritance.
# - Can be a bit more difficult to understand or read compared to inheritance.
# ### Code Example
# +
class Song:
def __init__(self, title, artist, album, track_number):
self.title = title
self.artist = artist
self.album = album
self.track_number = track_number
artist.add_song(self)
class Album:
def __init__(self, title, artist, year):
self.title = title
self.artist = artist
self.year = year
self.tracks = []
artist.add_album(self)
def add_track(self, title, artist=None):
if artist is None:
artist = self.artist
track_number = len(self.tracks)
song = Song(title, artist, self, track_number)
self.tracks.append(song)
class Artist:
def __init__(self, name):
self.name = name
self.albums = []
self.songs = []
def add_album(self, album):
self.albums.append(album)
def add_song(self, song):
self.songs.append(song)
class Playlist:
def __init__(self, name):
self.name = name
self.songs = []
def add_song(self, song):
self.songs.append(song)
def view_songs(self):
print(self.songs)
band = Artist("Bob's Awesome Band")
album = Album("<NAME>", band, 2013)
album.add_track("A Ballad about Cheese")
album.add_track("A Ballad about Cheese (dance remix)")
album.add_track("A Third Song to Use Up the Rest of the Space")
playlist = Playlist("My Favourite Songs")
for song in album.tracks:
print(song.album.year)
playlist.add_song(song)
print('\n')
for i in playlist.songs:
print(i.title)
# -
# ### When to use Inheritance
# - When the base class and the derived class(es) are under the control of the same programmers.
# - When the classes are well documented.
# - When there's clearly a "is-a" relationship.
# - When a new class REALLY defines a subtype of an existing class.
# - When extending classes specifically designed and documented for extension.
# #### => Classwork
# Using Composition, design a Developer class.
# ### Takeaways
| workshop/03-composition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <table width = "100%">
# <tr style="background-color:white;">
# <!-- QWorld Logo -->
# <td style="text-align:left;width:200px;">
# <a href="https://qworld.net/" target="_blank"><img src="../images/QWorld.png"> </a></td>
# <td style="text-align:right;vertical-align:bottom;font-size:16px;">
# Prepared by <a href="https://www.cmpe.boun.edu.tr/~ozlem.salehi/" target="_blank"> <NAME> </a> </td>
# </tr>
# </table>
#
# <hr>
# ## Quantum Annealing
# Quantum annealing (QA) is a heuristic method for solving optimization problems. It relies on the quantum adiabatic computation, which is an alternative to the gate based model. Unlike simulated annealing, it is a physical process, in which quantum fluctuations play the role of thermal fluctuations.
# <figure>
# <img src='../images/qa.jpg' width="350" />
# </figure>
# _Hamiltonian_ is the term used for describing the energies of any physical (classical) system. For many Hamiltonians, finding the lowest energy state is an NP-Hard problem, which is intractable for classical computers.
# For quantum systems, Hamiltonian is defined only for certain states, which are called the _eigenstates_. The idea behing quantum annealing is to create a Hamiltonian, whose lowest energy state (_ground state_) encodes the solution to the optimization problem we want to solve. Although finding the lowest energy state might be NP-Hard, quantum adiabatic theorem offers a new approach for solving the problem.
# Quantum adiabatic theorem guarantees that if a system is initially in ground state and evolves _slowly enough_ over the time, then it will be in ground state provided that the _spectral gap_ is always positive. Spectral gap is defined as the difference between the eigenvalues of the lowest energy state and the next lowest energy state (first excited state).
# Now consider the following Hamiltonian.
# $$H(t) = s(t) H_I + (1-s(t)) H_F$$
# $H_I$ is the initial Hamiltonian and $H_F$ is the final Hamiltonian. $s(t)$ decreases from 1 to 0, so that at the end of the procedure we have $H(t) = H_F$. Hence if we start with a Hamiltonian whose ground state is easy to prepare and evolve the system according to the above rules, then the system will end up with the ground state of Hamiltonian $H_F$. This gives the idea of picking $H_F$ as the Hamiltonian whose ground state is the solution we are looking for.
# For the initial state we pick equal superpoisiton of $|0\rangle$ and $|1\rangle$ for each qubit. Then $H_I$ can be expressed as
# $$
# H_I = \sum_{i=1}^n \frac{1}{2}(1- \sigma_i ^x)
# $$
# where $\sigma_i ^x$ denotes the Pauli-x operator applied on qubit $i$.
| notebooks/.ipynb_checkpoints/Quantum_Annealing-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Playground for exploring what gradient decent looks like on a 2D plane
#
# ##### Heavily influenced by https://lucidar.me/en/neural-networks/gradient-descent-example/
#
# # Find the minima of a simple 1D function:
# +
import numpy as np
import matplotlib.pyplot as plt
# Function to minimize
def function1D ( X ):
return (X-2)**2
# Numerical gradient
def numerical_gradient1D (XY, fn, delta):
X_plus = fn( XY + delta)
X_minus = fn( XY - delta)
return X_plus - X_minus
# Starting point
X = -30
# Step size multiplier
alpha=0.5
XHistory = np.zeros( (200,1) )
# Gradient descent (200 steps)
for x in range(0, 200):
X = X - alpha*numerical_gradient1D(X,function1D,0.05)
XHistory[x] = X
#print (XY)
# Print results
print ('X=', X)
print ('f=', function1D(X))
# -
# # Draw the path of the minimisation search in 1D
fig,ax=plt.subplots(1,1)
xlist = np.arange(-30, 30, 0.1)
ylist = function1D(xlist)
# Plot the funtion
ax.plot(xlist,function1D(xlist))
plt.show()
# # Plot again but with the path of the minima search
fig,ax=plt.subplots(1,1)
xlist = np.arange(-30, 30, 0.1)
ylist = function1D(xlist)
# Plot the funtion
ax.plot(xlist,function1D(xlist))
# Plot the history of the minima search
ax.scatter(XHistory,function1D(XHistory), color='r')
plt.show()
# ***
# # Find the minima of a 2D function
# +
# Function to minimize
def function ( XY ):
return (XY[0]-2)**2 + 2*(XY[1]-3)**2
# Gradient of the function
def gradient (XY):
return np.array([ 2*XY[0]-4 , 4*XY[1]-12 ])
# Numerical gradient
def numerical_gradient (XY, fn, delta):
deltax = np.array([delta,0])
deltay = np.array([0,delta])
X_plus = fn( XY + deltax)
X_minus = fn( XY - deltax)
Y_plus = fn( XY + deltay)
Y_minus = fn( XY - deltay)
return np.array ([ X_plus - X_minus, Y_plus - Y_minus])
# Starting point
XY = np.array([-30,-20])
# Step size multiplier
alpha=0.5
# Gradient descent (200 steps)
for x in range(0, 200):
XY = XY - alpha*numerical_gradient(XY,function,0.05)
#print (XY)
# Print results
print ('XY=', XY)
print ('f=', function(XY))
# Output expected:
# XY= [2.00000002 3. ]
# f= 3.9022927947211357e-16
# -
# # Plot the 2D plane of a simple function
# +
import numpy as np
import matplotlib.pyplot as plt
def DrawFunction( fn , range ):
xlist = np.linspace(-range, range, 300)
ylist = np.linspace(-range, range, 300)
X, Y = np.meshgrid(xlist, ylist)
Z = fn(np.array([X, Y]))
fig,ax=plt.subplots(1,1)
cp = ax.contour(X, Y, Z)
fig.colorbar(cp) # Add a colorbar to a plot
ax.clabel(cp, inline=True, fontsize=10)
ax.set_title('Contour Plot of Function')
#ax.set_xlabel('x (cm)')
ax.set_ylabel('y (cm)')
return fig, cp
myFig, ContourPlot = DrawFunction(function,30)
myFig.show()
# -
# # Plot the path of decent for a simple function
# +
fig, cp = DrawFunction(function,30)
# Starting point
XY = np.array([-30,-20])
# Step size multiplier
alpha=0.5
# Number of steps to take
steps = 200
# Gradient descent (200 steps)
XYHistory = np.zeros( (steps,2 ))
for x in range(0, 200):
XY = XY - alpha*numerical_gradient(XY,function,0.05)
XYHistory[x] = XY
#print (XY)
#print (XYHistory)
plt.scatter(XYHistory[:,0], XYHistory[:,1], color='r')
plt.show()
# -
# ***
# # Define and draw a more complex function with multiple local minima
# +
# Fancy function
def function2 ( XY ):
return 100*np.exp(np.cos(0.30*XY[0]) -np.sin(0.25*XY[1]))-100 +(XY[0]-2)**2 + 1*(XY[1]-3)**2
DrawFunction(function2,30)
# -
# # Find the minima through gradient decent, note how this starting location and step size doesn't give us the lowest minima
# +
DrawFunction(function2,30)
# Starting point
XY = np.array([-30,-20])
# Step size multiplier
alpha=0.5
# Number of steps to take
steps = 200
# Gradient descent (200 steps)
XYHistory = np.zeros( (steps,2 ))
for x in range(0, steps):
XY = XY - alpha*numerical_gradient(XY,function2,0.05)
XYHistory[x] = XY
#print (XY)
#print (XYHistory)
plt.scatter(XYHistory[:,0], XYHistory[:,1], color='r')
plt.show()
# Print results
print ('XY=', XY)
print ('f=', function(XY))
# -
# # Repeat above but add momentum like term with some damping... now the gradient decent can find a new minima
# +
DrawFunction(function2,30)
# Starting point
XY = np.array([-30,-20])
# Step size multiplier
alpha=0.5
# Number of steps to take
steps = 200
# Gradient descent (200 steps)
XYHistory = np.zeros( (steps,2 ))
Velocity = 0
for x in range(0, steps):
XY = XY - alpha*numerical_gradient(XY,function2,0.05) + Velocity* 0.7
XYHistory[x] = XY
Velocity = XYHistory[x] - XYHistory[x - 1]
#print (XY)
#print (XYHistory)
plt.scatter(XYHistory[:,0], XYHistory[:,1], color='r')
plt.show()
# Print results
print ('XY=', XY)
print ('f=', function(XY))
| GradientDecentPlayground.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install -r requirements.txt
from mindee import Client
import json
import cv2
from matplotlib import pyplot as plt
import numpy as np
import math
from IPython.lib.pretty import pretty
mindee_client = Client(
passport_token='<insert api key>',
raise_on_error=True
)
#get a free api key at platform.mindee.com
#load image - get height and width - show image
imagepath = 'images/fakepassport.jpg'
image = cv2.imread(imagepath)
#fix colour space
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
height, width, channels = image.shape
print(f"Image size: ({height}, {width})")
plt.imshow(image)
plt.show()
parsed_data = mindee_client.parse_passport(imagepath)
print(parsed_data.passport)
print(parsed_data.http_response)
given_names = parsed_data.passport.given_names
print("all given names: ")
for given_name in given_names:
# To get the name string
name = given_name.value
print(name)
surname = parsed_data.passport.surname.value
print("surname: ",surname)
# To get the passport's owner gender (string among {"M", "F"}
# Note that the fake passport in the example provides the incorrect value. The signature is also Bruce Lee's.
gender = parsed_data.passport.gender.value
print("gender: ", gender)
# To get the passport's owner full name (string)
full_name = parsed_data.passport.full_name.value
print("fullname: ",full_name)
# To get the passport's owner birth place (string)
birth_place = parsed_data.passport.birth_place.value
print("birthplace: ", birth_place)
# +
# To get the passport's owner date of birth (string)
birth_date = parsed_data.passport.birth_date.value
# To get the passport date of issuance (string)
issuance_date = parsed_data.passport.issuance_date.value
# To get the passport expiry date (string)
expiry_date = parsed_data.passport.expiry_date.value
print("DOB: ",birth_date)
print("issued: ", issuance_date)
print("expires: ",expiry_date)
# +
# To get the passport first line of machine readable zone (string)
mrz1 = parsed_data.passport.mrz1.value
# To get the passport seco d line of machine readable zone (string)
mrz2 = parsed_data.passport.mrz2.value
# To get the passport full machine readable zone (string)
mrz = parsed_data.passport.mrz.value
print(mrz1)
print(mrz2)
print(mrz)
# +
# To get the passport id number (string)
id_number = parsed_data.passport.id_number.value
# To get the passport country code (string)
country_code = parsed_data.passport.country.value
print("passport number: ", id_number)
print("passport country code: ", country_code)
| MindeePassport.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import warnings
warnings.filterwarnings("ignore")
from nilmtk.api import API
from nilmtk.disaggregate import Mean
from nilmtk_contrib.disaggregate import DAE,Seq2Point, Seq2Seq, RNN, WindowGRU
redd = {
'power': {
'mains': ['apparent','active'],
'appliance': ['apparent','active']
},
'sample_rate': 60,
'appliances': ['fridge'],
'methods': {
'WindowGRU':WindowGRU({'n_epochs':50,'batch_size':32}),
'RNN':RNN({'n_epochs':50,'batch_size':32}),
'DAE':DAE({'n_epochs':50,'batch_size':32}),
'Seq2Point':Seq2Point({'n_epochs':50,'batch_size':32}),
'Seq2Seq':Seq2Seq({'n_epochs':50,'batch_size':32}),
'Mean': Mean({}),
},
'train': {
'datasets': {
'Dataport': {
'path': 'C:/Users/Hp/Desktop/nilmtk-contrib/dataport.hdf5',
'buildings': {
10: {
'start_time': '2015-04-04',
'end_time': '2015-04-05'
},
# 56: {
# 'start_time': '2015-01-28',
# 'end_time': '2015-01-30'
# },
}
}
}
},
'test': {
'datasets': {
'Datport': {
'path': 'C:/Users/Hp/Desktop/nilmtk-contrib/dataport.hdf5',
'buildings': {
10: {
'start_time': '2015-04-05',
'end_time': '2015-04-06'
},
}
}
},
'metrics':['mae']
}
}
api_res = API(redd)
api_res.errors
api_res.errors_keys
| sample_notebooks/Using the API with NILMTK-CONTRIB.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# [View in Colaboratory](https://colab.research.google.com/github/nkcr/NETARCH/blob/master/Implementing%20a%20CNN%20for%20TC/Run%20Tensorboard.ipynb)
# + [markdown] id="A-bHuQhGSTVk" colab_type="text"
# ### Using TensorBoard
# + id="2skBXtLcRp4m" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="c60c78cc-b3ab-45a4-c50a-3a74f58b49f3"
import tensorflow as tf
#LOG_DIR = '/tmp/log'
LOG_DIR= '/content/runs/'
#LOG_DIR= '/content/runs/1526979864/checkpoints'
get_ipython().system_raw(
'tensorboard --logdir {} --host 0.0.0.0 --port 6006 &'
.format(LOG_DIR)
)
# ! wget -nc https://bin.equinox.io/c/4VmDzA7iaHb/ngrok-stable-linux-amd64.zip
# ! unzip -n ngrok-stable-linux-amd64.zip
get_ipython().system_raw('./ngrok http 6006 &')
# ! curl -s http://localhost:4040/api/tunnels | python3 -c \
# "import sys, json; print(json.load(sys.stdin)['tunnels'][0]['public_url'])"
# + id="hpKoj6vyo3M3" colab_type="code" colab={}
# ! cd runs && rm -rf *
# + id="UGkE7xt4iQMv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 125} outputId="1bf8c172-f0df-4631-a94b-fcfe2db835a6"
# ! ps -ef | grep 6006
# ! kill -9 23162 23190
# ! ps -ef | grep 6006
# + id="DWlZJfMJftSa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 53} outputId="ba754c4f-1c05-40ee-c4e9-3d9919214bf6"
# ! ls -l /content/runs/
# + id="DI-uBldlXkg7" colab_type="code" colab={}
# rm -rf runs/1527153935
# + [markdown] id="vSJ9qR5GXfrA" colab_type="text"
# ### Get public link
# + id="q0Pw5lGDpK_b" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 251} outputId="21713c69-cfb1-46a0-a61e-871e4c200001"
FILE="GoogleNews-vectors-negative300.bin"
# !wget -nc 'https://s3.amazonaws.com/dl4j-distribution/GoogleNews-vectors-negative300.bin.gz'
# !ls
get_ipython().system_raw(
'[ -f "{}" ] && echo "skipping {}" || gunzip "{}.gz"'
.format(FILE,FILE,FILE)
)
# !ls
| Implementing a CNN for TC/Run Tensorboard.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SPYN - III phase AC motor control
# <div class="alert bg-primary">This notebook will show control of a 3-phase AC motor using the EDPS (The Electric Drive Power Stage (EDPS) Board, a Trenz Electronic TEC0053, which is connected to the PYNQ-Z1 controller board for the evaluation..</div>
# 
# ## Objectives
#
# * [Access to Motor Control Parameters](#Step-2:-Instantiate-the-motor-control-object )
# * [Request Status Information of the Motor](#Step-4:-Read-status-registers)
# * [Programmatic Control of Motor](#Programmatic-Control-of-Motor)
# * [Continuous Status Capture from the Motor](#Step-5:-Allocate-DMA-for-stream-capture)
# * [Plots to Visualize Data Captured](#Plotting)
# * [Storing Captured Data for Analytics](#Dataframes-for-analytics)
# * [Live Interactive Plots to Investigate Data](#DASH-Demo)
#
# ### Step 1: Download the `EDDP` bitstream
# +
from pynq import Overlay
from pynq import MMIO
import numpy as np
overlay = Overlay(
"/opt/python3.6/lib/python3.6/site-packages/spyn/overlays/spyn/spyn.bit")
overlay.download()
# -
# ### Step 2: Instantiate the motor control object
# #### Set the motor mode
# +
from spyn.lib import *
motor = Motor_Controller()
print(f'Available motor modes : {motor.motor_modes}')
# -
motor.set_mode('reset_mode')
# #### Available control and status IP blocks
print(f'Memory mapped IO blocks : {motor.mmio_blocks}')
# ### Step 3: Set motor control mode and control using sliders
# +
from ipywidgets import interact, interactive, HBox, VBox
import ipywidgets as widgets
text = {'Motor': 'success', 'Forward': 'info', 'Reverse': 'warning'}
buttons, colors = list(text.keys()), list(text.values())
toggle = [
widgets.ToggleButton(description=f'{buttons[i]}',
button_style=f'{colors[i]}')
for i in range(3)]
mode = widgets.Dropdown(options=['Speed', 'Current'])
def clicked(toggle_0=toggle[0], mode=mode, toggle_1=toggle[1],
toggle_2=toggle[2], RPM=None, Torque=None):
if toggle_0:
if mode == 'Speed':
motor.set_mode('rpm_mode')
motor.set_rpm(RPM)
elif mode == 'Current':
motor.set_mode('torque_mode')
motor.set_torque(Torque)
else:
motor.stop()
w = interactive(clicked,
RPM=widgets.IntSlider(min=-5000, max=5000, step=1, value=1000),
Torque=widgets.IntSlider(min=-400, max=400, step=1, value=0))
VBox([HBox(w.children[:2]), w.children[2], w.children[3], w.children[4],
w.children[5]])
# -
# [Back to Objectives](#Objectives)
# ### Step 4: Read status registers
# +
motor_status = [(motor._read_controlreg(i + ANGLE.offset)) for i in
range(0, 16, 4)]
high_sp, low_sp = bytesplit(motor_status[1])
high_id, low_id = bytesplit(motor_status[2])
high_iq, low_iq = bytesplit(motor_status[3])
print(f'Angle in degrees : {motor_status[0] * 0.36}')
print(f'Angle in steps per thousand: {(motor_status[0])}')
print(f'Id : {np.int16(low_id) * 0.00039} Amp')
print(f'Iq : {np.int16(low_iq) * 0.00039} Amp')
print(f'Speed in RPM : {-(np.int16(low_sp))}')
# -
# [Back to Objectives](#Objectives)
# ### Programmatic Control of Motor
import time
motor.set_mode('rpm_mode')
for i in range(2):
motor.set_rpm(1000)
time.sleep(1)
motor.set_rpm(0)
time.sleep(2)
motor.set_rpm(-50)
time.sleep(2)
motor.set_rpm(0)
time.sleep(2)
motor.stop()
# [Back to Objectives](#Objectives)
# ### Set capture mode
print(f'Available stream capture modes : {motor.motor_capture_modes}')
motor.capture_mode('ia_ib_angle_rpm')
# ### Step 5: Allocate DMA for stream capture
# +
from pynq import Xlnk
xlnk = Xlnk()
input_buffer = xlnk.cma_array(shape=(256,), dtype=np.uint8)
capture_address = input_buffer.physical_address
# -
print(f'Physical Address of data stream capture: {hex(capture_address)}')
# ### Step 6: Log stream data as per control mode
# +
from pynq import MMIO
# capture_count = int(input('Enter capture count: '))
capture_count = 1000
def continuous_capture(capture_count):
mmio_stream = MMIO(capture_address, 256)
cap_list = [([]) for i in range(4)]
for _ in range(capture_count):
motor.stream_capture(capture_address)
for i in range(4, 260, 4):
stream = mmio_stream.read(i - 4, 4)
highbits, lowbits = bytesplit(stream)
if (i % 8 != 0):
cap_list[0].extend([(np.int16(lowbits))])
cap_list[1].extend([(np.int16(highbits))])
else:
cap_list[2].extend([(np.int16(lowbits))])
cap_list[3].extend([(np.int16(highbits))])
return cap_list
cap_list = continuous_capture(capture_count)
Ia, Ib, angle, rpm = cap_list[0], cap_list[1], cap_list[3], cap_list[2]
current_Ia = np.array(Ia) * 0.00039
current_Ib = np.array(Ib) * 0.00039
# -
# [Back to Objectives](#Objectives)
# ### Step 7: Plotting
# #### Ia vs Sample count
# %matplotlib inline
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(20, 10))
ax = fig.add_subplot(111)
ax.plot(current_Ia)
plt.show()
# #### Ib vs Sample count
fig = plt.figure(figsize=(20, 10))
ax = fig.add_subplot(111)
ax.plot(current_Ib)
plt.ylabel('Ib')
plt.show()
# #### Ia vs Ib
fig = plt.figure(figsize=(20, 10))
ax = fig.add_subplot(111)
ax.scatter(current_Ia, current_Ib)
plt.show()
# #### Angle vs Sample count
fig = plt.figure(figsize=(20, 10))
ax = fig.add_subplot(111)
ax.plot(angle)
plt.show()
# #### RPM vs Sample count
fig = plt.figure(figsize=(20, 10))
ax = fig.add_subplot(111)
ax.plot(rpm)
plt.show()
# [Back to Objectives](#Objectives)
# ### Step 8: Stop motor and reset xlnk
xlnk.xlnk_reset()
motor.stop()
# ### Dataframes for analytics
# +
import pandas as pd
data = {'Ia' : current_Ia,
'Ib' : current_Ib,
'angle':cap_list[3],
'rpm': cap_list[2]}
df = pd.DataFrame(data, columns = ['Ia', 'Ib', 'angle', 'rpm'])
df
# -
df.to_csv('motor_data.txt')
# [Back to Objectives](#Objectives)
| notebooks/spyn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# cd /Users/alex/Documents/data
# +
import os
import pandas as pd
def listdirInMac(path):
os_list = os.listdir(path)
for item in os_list:
if item.startswith('.') and os.path.isfile(os.path.join(path, item)):
os_list.remove(item)
os_list.sort()
return os_list
root='/Users/alex/Documents/data/1-15-15/'
newfile=pd.DataFrame()
pieces=[]
#all txt data will append on this variable
for file in listdirInMac(root):
if 'CSV' in file:
pathname=os.path.join(root,file) #file path <--add additional tab here
print(pathname)
temp=pd.read_csv(pathname,usecols=[1],header=None) # read csv <--add additional tab here
pieces.append(temp)
newfile = pd.concat(pieces, axis=1)
newfile.to_csv('/Users/alex/Documents/data/15.csv') #save as a newfile
# -
| 500CSV-1CSV.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="LHFPr4DxIFjY"
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.cluster import KMeans
from sklearn.decomposition import PCA
# + colab={"base_uri": "https://localhost:8080/"} id="KjZwVa72JhK9" outputId="381c1af1-4861-4a45-8a3f-5e0fb141030f"
df=pd.read_csv('f8.txt',sep=' ',index_col=None,header=None)
# + colab={"base_uri": "https://localhost:8080/"} id="lgXCcB1maOHf" outputId="fa8661a6-1a68-4442-8185-d6a45f966d09"
df.shape
# + [markdown] id="GWHiOiszZ3uz"
# We first normalize the data set
# + id="UcsXm4t8Z2qN"
df_norm = df.copy()
df_norm = df_norm.astype(float)
for j in range(df.shape[1]):
for i in range(len(df[j])):
df_norm[j][i] = (df_norm[j][i]-df[j].mean())/df[j].std()
# + id="be-m9OgdJ-sa"
X=df_norm.to_numpy()
# + [markdown] id="b0xooP52bAlT"
# Now we find the best choice of number of clusters
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="trs1A4HWbEiB" outputId="afb7f6ec-9305-4ca8-ad74-503b20d77791"
cost2=[]
for i in range(5,20):
KM2=KMeans(n_clusters=i)
KM2.fit(X)
cost2.append(KM2.inertia_)
plt.plot(range(5,20),cost2)
plt.xlabel('Value of K')
plt.ylabel('squared error(cost)')
plt.show
# + [markdown] id="OAjG35CmbZw_"
# We take no. of clusters as 16.
# + [markdown] id="jFzIo7YNgIYd"
# #Q:1
# + id="SXfODMZ_KGBy"
km=KMeans(n_clusters=16,random_state=0)
# + id="3n3hEjjmKSCa"
km_result=km.fit(X)
# + colab={"base_uri": "https://localhost:8080/"} id="UWMb617tlnLk" outputId="fca08320-078a-4b4e-83bf-034f841c63d7"
for i in range(len(km_result.labels_)):
print(km_result.labels_[i],', ', end='')
# + [markdown] id="DndhO3HngBFT"
# #Q:2
#
# Since, 'f8.txt' is a high dimensional dataset, therefore we first apply dimensionality reduction using 'PCA' and reduce that to 2-dimensions and then we visualise the clusters as before.
#
# Now, coming to PCA, it is a technique for reducing the number of dimensions in a dataset whilst retaining most information. It is using the correlation between some dimensions and tries to provide a minimum number of variables that keeps the maximum amount of variation or information about how the original data is distributed. It does not do this using guesswork but using hard mathematics and it uses the eigenvalues and eigenvectors of the data-matrix. These eigenvectors of the covariance matrix have the property that they point along the major directions of variation in the data. These are the directions of maximum variation in a dataset.
# + [markdown] id="yjQCl7rbmq7z"
# #Q:3
# + [markdown] id="Px4yMrasp3gD"
# Applying dimensionality reduction using PCA to reduce the dimension to 2-Dimensions
# + id="-wPNvhwBKnU6"
pca=PCA(n_components=2)
pca.fit(X)
pca_data=pd.DataFrame(pca.transform(X))
# + [markdown] id="658zA1x0qLBb"
# Printing the reduced dataset
# + colab={"base_uri": "https://localhost:8080/"} id="QTdJKR-yLoen" outputId="21ca9b60-9865-46f6-ae19-f409e0b21b7d"
print(pca_data)
# + [markdown] id="3K0QYe7nqQaZ"
# Now we're going to apply KMeans on the reduced dataset for clustering
# + id="vgAzxMzdLvdl"
Z=pca_data.to_numpy()
# + id="SV16HVjLL2jm"
km16=KMeans(n_clusters=16,random_state=None)
# + id="vWNfToo9MDFH"
km16_result=km16.fit(Z)
# + [markdown] id="smE2yY_Fqny0"
# Plotting the clusters
# + colab={"base_uri": "https://localhost:8080/", "height": 434} id="ki8eTHhDmtKI" outputId="6ac24698-478e-4d28-abfb-7fcea828b390"
plt.figure(figsize=(16,10))
sns.scatterplot(x=Z[:,0],y=Z[:,1],hue=km16_result.labels_,palette=['red','green','blue','yellow','indigo','cyan','magenta','slategrey','thistle','olive','steelblue','deeppink','salmon','midnightblue','darkslategrey','maroon'])
# + [markdown] id="U1h5YBJFmnMT"
# #Q:4
# + [markdown] id="VP6OKY5jpvw4"
# Plotting the cluster centers in the plots thus prepared.
# + colab={"base_uri": "https://localhost:8080/", "height": 434} id="GoxOpzaxMHr0" outputId="38e05a00-746c-4f99-969d-e8dc608b1d3e"
plt.figure(figsize=(16,10))
sns.scatterplot(x=Z[:,0],y=Z[:,1],hue=km16_result.labels_,palette=['red','green','blue','yellow','indigo','cyan','magenta','slategrey','thistle','olive','steelblue','deeppink','salmon','midnightblue','darkslategrey','maroon'])
plt.scatter(x=km16_result.cluster_centers_[:,0],y=km16_result.cluster_centers_[:,1],marker="^",color='black')
| Clustering with more than 2 dimensions/Clustering with more than 2 dimensions, provided f8.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # Annual Key Performance Indicators for Strathcona County Library
#
# ## What you need to do
#
# The [Strathcona County Library](https://sclibrary.ca/) has many services, books, and programs. Now that they share a catalogue with [Fort Saskatchewan Public Library](https://fspl.ca/) and participate in [The Alberta Library](http://thealbertalibrary.ca/), they are considering expanding to a library location on Mars.
#
# So they need to know what they are doing well and should continue doing. The Strathcona County Library collects and provides open access to datasets, which can help answer this question. Performance indicators can provide valuable feedback and information for moving forward with this project.
#
# That’s where you as a data scientist come in. They’ve provided the data and you need to find out the library’s top-performing services, programs, and books, so they know what to bring to the red planet.
#
# ## Data Content
# This data set is provided by the Strathcona County Library, Strathcona County, Alberta, Canada. Source: https://data.strathcona.ca/Recreation-Culture/Library-Key-Performance-Indicators/ep8g-4kxs
#
# ### Key Performance Indicators (per year)
#
# | Indicator | Description |
# |---|---|
# | Answers | types of answers the library provides to staff and reference questions|
# | Awareness | number of awareness sessions and the number of County residents attending |
# | Borrowing | measure and classification of loans and borrowed items |
# | Collections | kinds and numbers of items the library manages |
# | Internet | number of patrons using WiFi or public computers |
# | Library use | number of cards purchased; number of people reporting use of the library |
# | Open days | total number of days the library was open |
# | Population | total number of served population |
# | Programs | number of offered program sessions and the number of participants |
# | Services | use of fax, printing, scanning, copying resumes (since 2011) |
# | Training | number of sessions and trainees on formal and informal technological sessions |
# | Visits | number of kinds of site visits |
# | Volunteers | number of volunteers |
# ## Downloading and parsing the data
#
# We'll begin by downloading the data from [the website](https://data.strathcona.ca/Recreation-Culture/Library-Key-Performance-Indicators/ep8g-4kxs) into a "dataframe" in this notebook.
#
# We selected the 'API' tag and chose CSV format on the top right side. Pressing the 'Copy' button gave us the URL to download the full dataset.
# +
# import the Pandas module that we will be using to manipulate the data
import pandas as pd
print("Importing Python library was sucessful!")
# this is the link we will get the data from
link = "https://data.strathcona.ca/resource/ep8g-4kxs.csv"
# Read and parse data as a Pandas CSV
rawData = pd.read_csv(link)
# Rename columns
rawData = rawData.rename(columns={"kpicategory": "category", "kpimeasure": "measure","kpiyear":"year","kpivalue":"value"})
# Look at the first five columns
rawData.head()
# -
# ## Selecting and Sorting Columns
#
# Now that we have downloaded the full dataset, it's time to select and sort columns using Pandas dataframes.
#
# We stored our data in a dataframe called `rawData`.
#
# Let's explore the `'category'` column and then look at the unique values.
# +
# Access the values under 'category' by using square brackets, followed by the column name in quotation marks
rawData["category"]
# Get unique values under 'category' column
rawData["category"].unique()
# possible category values are:
# 'Answers', 'Awareness', 'Borrowing', 'Collections', 'Internet',
# 'Library use', 'Open days', 'Population', 'Programs', 'Services',
# 'Training', 'Visits', 'Volunteers'
# -
# Let's look at the population size each year using the "Population" category.
#
# We can select the subset of data we are interested in by using a condition:
#
# `dataframe['column_name']=='Category_name'`
#
# and then passing that condition into the dataframe within square brackets:
#
# `dataframe[dataframe['column_name']=='Category_name']`
#
# Using our dataframe `rawData`, we can access the subset of the data whose `category` is `'Population'` as follows:
# Set up condition to get only those rows under 'category' whose value matches 'Population'
condition_1 = rawData["category"]=='Population'
# Pass the condition in square brackets to get subset of the data
rawData[condition_1]
# +
# You could also do this directly without declaring a condition_1 variable
# rawData[rawData["category"]=='Population']
# -
# We can also pass multiple conditions by encapsulating each condition in parenthesis, and separating each condition by the logical operator **and** `&` as well as the logical operator **or** `|`.
#
# In this dataset, any two values under `category` are [mutually exclusive](https://www.mathsisfun.com/data/probability-events-mutually-exclusive.html), so we would need to use the `|` operator.
#
# `category` and `year` are not mutually exclusive, so we can obtain specific data points by using the `&` symbol.
#
# Let's suppose we want to know how many people have used the library per year since 2012.
#
# ###### Condition 1:
#
# We want all those entries whose `'category'` is equal to `'Population'`
#
# `rawData["category"]=='Population'`
#
# ###### Condition 2:
#
# We want all those entries whose `'year'` is greater than or equal to `2012`
#
# `rawData["year"]>=2012`
# +
# set up the conditions
condition_1 = rawData["category"]=='Population'
condition_2 = rawData["year"]>=2012
# pass conditions - each in parenthesis and separated by |
rawData[(condition_1) & (condition_2)]
# -
# ---
# ### Challenge 1a
#
# Now that you know how to filter data using conditions, use code cells below to find how many "Volunteers" were there in 2015. (Hint: use the `year` and `category` as two conditions)
#
# ---
# ### Challenge 1b
#
# Like any good data scientist, explore the data by manipulating the dataframe to learn something about the data. You are free to choose any `category`, `measure`, `year` or `value` in the dataframe.
# ## Simple Summary Statistics
#
# We can use built-in functions in Python to find the average, maximum, and minimum values in the data set. Run the cell below to learn more about these values for the population.
# +
# We will then access the 'value' column within the subset containing the 'Population' category
# and compute the average size of popultion between 2001 and 2016 using the mean() method
print("The average served population size between 2001 and 2016 is: ")
print(rawData[condition_1]["value"].mean())
# Same thing, except print the maximum
print("The maximum served population size between 2001 and 2016 is: ")
print(rawData[condition_1]["value"].max())
# ... and minimum
print("The minimum served population size between 2001 and 2016 is: ")
print(rawData[condition_1]["value"].min())
# -
# ---
# ### Challenge 2
#
# Find the maximum, minimum, and average number of 'Open days' per year. The average is done for you to get you started :)
#
# ---
# +
# We will then access the 'value' column within the subset containing the 'Open days' category
# and compute the average size of popultion between 2001 and 2016 using the mean() method
condition_2 = rawData["category"]=='Open days'
print("The average number of days the library was open between 2001 and 2016 is: ")
print(rawData[condition_2]["value"].mean())
# -
# ## Simple Visualization
# It's not easy to see patterns and trends in data by looking at tables, so let's create visualizations (graphs) of our data. We'll use the `cufflinks` Python library.
# +
# If you get the error 'no module named cufflinks' then uncomment the follow line:
# #!pip install cufflinks ipywidgets
# load the Cufflinks library and call it by the short name "cf"
import cufflinks as cf
# command to display graphics correctly in Jupyter notebook
cf.go_offline()
def enable_plotly_in_cell():
import IPython
from plotly.offline import init_notebook_mode
display(IPython.core.display.HTML('''<script src="/static/components/requirejs/require.js"></script>'''))
init_notebook_mode(connected=False)
get_ipython().events.register('pre_run_cell', enable_plotly_in_cell)
print("Successfully imported Cufflinks and enabled Plotly!")
# -
# ### Statistics About Library Program Usage
#
# We want to know how many people have used library program services, and compare that to both the number of program services and the total population size.
#
# We can specify conditions to access that subset of the data:
#
# `condition_1 = rawData["category"]=='Population'`
#
# will let us access all data points containing the size of served population, and:
#
# `condition_3 = rawData["category"]=='Programs'`
#
# will let us access all data points containing the number of programs offered as well as the number of people who accessed the programs.
#
# Remember that these categories are mutually exclusive.
# We can then plot our subset that contains `condition_1` and `condition_3` to learn things from the data.
#
# In the plot below, we'll use a `scatter` plot, where the data are categorized by the `measure` column. On the x-axis is the `year` and the y-axis contains the `value`.
#
# We'll also add a `title` so we know what the graph represents.
# +
# set up conditions (type or paste from above)
condition_1 =
condition_3 =
rawData[ (condition_1)| (condition_3)].iplot(
kind="scatter",mode='markers',
y="value",x="year",text='measure',categories='measure',
title="Population served, compared against Program Participants and Number of Program Sessions")
# -
# #### Observations
#
# We picked this category to find out how the population size, number of program participants, and number of program sessions have changed over time.
#
# We observe an upward trend in the total population served per year.
#
# We also observe an upward trend in the number of program participants. This is interesting, given that the number of program sessions mostly didn't change.
#
# This suggests that the increase in number of participants is related to the served population size, not to the number of program sessions.
# ---
# ### Challenge 3
#
# Let's visualize multiple categories.
#
# 1. Run the cell below and look at the unique category values. Pick one that you are interested in.
# 2. In the next cell, look at the plot using the `'Collections'` category. Now substitute `'Collections'` for the value you chose, and change the `given_title` variable to give an appropriate name to your plot. Run that cell.
# 3. Hover over the plot. What do you observe?
#
# ---
rawData["category"].unique()
# +
# Change the input below
# Substitute the value to be selected under 'category'
condition_4 = rawData["category"]=="Collections"
# Give the plot an appropriate title
given_title = "Library Collections"
# Run this cell when you have an appropriate value under category and title for the plot
rawData[ (condition_4)].iplot(kind='scatter',mode='markers',y="value",
x="year",text='measure',categories='measure',
title=given_title)
# -
# #### Observations
#
# Double click this cell and state:
#
# 1. The category you picked
# 2. Why you were interested
# 3. What you learned from the graph
# ---
# ### Challenge 4
#
# Create visualizations of more categories.
#
# Think about how you would grade the library's performance based on the key indicators.
#
# ---
# # Conclusions
#
# Edit this cell to describe **what you would recommend for Strathcona County to bring and provide on Mars**, based on the dataset containing "Library Key Performance Indicators". Include any data filtering and sorting steps that you recommend, and why you would recommend them.
#
#
# ## Reflections
#
# Write about some or all of the following questions, either individually in separate markdown cells or as a group.
# - What is something you learned through this process?
# - How well did your group work together? Why do you think that is?
# - What were some of the hardest parts?
# - What are you proud of? What would you like to show others?
# - Are you curious about anything else related to this? Did anything surprise you?
# - How can you apply your learning to future activities?
# 
| StrathconaCountyOpenData/LibraryKeyPerformanceIndicators/Library-Challenge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
num = (int(input('Digite um valor: ')), int(input('Digite mais um valor: ')), int(input('Digite outro valor: ')), int(input('Digite mais um valor: ')))
print(f'Você digitou os valores {num}')
print(f'O valor 9 apareceu {num.count(9)} vezes')
if 3 in num:
print(f'O valor 3 apareceu na {num.index(3)+1}ª posição')
else:
print('Não foi encontrado o numero 3')
for n in num:
if n%2 ==0:
print(f'Os valores pares digitados foram ', end='')
print(n, end=' ')
print('\nNão tem numero par')
| .ipynb_checkpoints/EX075 - Análise de dados em uma Tupla-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tfs
# language: python
# name: tfs
# ---
# # Image classification: CIFAR dataset
# This notebooks experiments with image classification with a different ** model architecture** from those we've been using before
# +
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# -
labels = ['airplane', 'automobile', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck']
# initialize parameters
batch_size = 64
n_classes = 10
epochs = 100
lr = 1e-4
# ##### preprocess the dataset
# +
cfar = tf.keras.datasets.cifar10
(x_train, y_train), (x_test, y_test) = cfar.load_data()
x_train, x_test = x_train / 255., x_test / 255.
# -
# use one-hot encoding
y_train = tf.one_hot(y_train, depth=10)
y_test = tf.one_hot(y_test, 10)
# +
# create a function to help display our inputs
def show_images(images, y_labels=None):
"""
Displays an array of images on a visual plot
"""
fig=plt.figure(figsize=(8, 8))
columns = 6
rows = 4
for i in range(1, columns*rows +1):
img = np.reshape(images[i], [w, h])
ax = fig.add_subplot(rows, columns, i)
if y_labels is not None:
ax.set_title(labels[np.argmax(y_labels[i])])
plt.imshow(img, cmap='Greys')
plt.tight_layout()
plt.show()
# -
# ##### define the model architecture
model = tf.keras.Sequential()
# +
model.add(tf.keras.layers.Conv2D(filters=32,
kernel_size=3,
padding='same',
kernel_regularizer=tf.keras.regularizers.l2(lr),
input_shape=(28, 28, 1)))
model.add(tf.keras.layers.Activation('relu'))
model.add(tf.keras.layers.BatchNormalization())
model.add(tf.keras.layers.Conv2D(filters=32,
kernel_size=3,
padding='same',
kernel_regularizer=tf.keras.regularizers.l2(lr)))
model.add(tf.keras.layers.Activation('relu'))
model.add(tf.keras.layers.BatchNormalization())
model.add(tf.keras.layers.MaxPool2D())
model.add(tf.keras.layers.Dropout(.2))
model.add(tf.keras.layers.Conv2D(filters=64,
kernel_size=3,
padding='same',
kernel_regularizer=tf.keras.regularizers.l2(lr)))
model.add(tf.keras.layers.Activation('relu'))
model.add(tf.keras.layers.BatchNormalization())
model.add(tf.keras.layers.Conv2D(filters=64,
kernel_size=3,
padding='same',
kernel_regularizer=tf.keras.regularizers.l2(lr))
)
model.add(tf.keras.layers.Activation('relu'))
model.add(tf.keras.layers.BatchNormalization())
model.add(tf.keras.layers.MaxPool2D())
model.add(tf.keras.layers.Dropout(.3))
model.add(tf.keras.layers.Conv2D(filters=128,
kernel_size=3,
padding='same',
kernel_regularizer=tf.keras.regularizers.l2(lr)))
model.add(tf.keras.layers.Activation('relu'))
model.add(tf.keras.layers.BatchNormalization())
model.add(tf.keras.layers.Conv2D(filters=128,
kernel_size=3,
padding='same',
kernel_regularizer=tf.keras.regularizers.l2(lr)))
model.add(tf.keras.layers.Activation('relu'))
model.add(tf.keras.layers.BatchNormalization())
model.add(tf.keras.layers.MaxPool2D())
model.add(tf.keras.layers.Dropout(.4))
model.add(tf.keras.layers.Flatten())
model.add(tf.keras.layers.Dense(units=n_classes,
activation='softmax'))
# -
model.summary()
model.compile(optimizer=tf.keras.optimizers.Adam(lr),
loss=tf.keras.losses.categorical_crossentropy,
metrics=['accuracy'])
# #### implement real-time data augmentation
# This will apply various **transformations** to the images such as flips, shifts, rotations, which should enable the model **generalize** better
data_gen = tf.keras.preprocessing.image.ImageDataGenerator(
rotation_range=10, # randomly rotate images between 0 - 9 degrees
width_shift_range=0.1, # shift images horizontally for total width fraction,
height_shift_range=0.1, # shift vertically over height fraction
horizontal_flip=True,
vertical_flip=True,
zoom_range=1.0,
validation_split=.1
)
# +
# setup an early stopping callback
call_back = tf.keras.callbacks.EarlyStopping(monitor='val_loss', min_delta=1e-5, patience=3,
restore_best_weights=True)
# -
model.fit_generator(generator=data_gen.flow(x_train, y_train, batch_size=batch_size),
epochs=epochs, callbacks=[call_back])
# The fit generator method fits the model on the data returned by the `data_gen`
# ##### Save or load the model
model.save('.model.cifar.ckpt')
model.load('.model.cifar.ckpt')
# ##### Evaluate the model
model.evaluate(x_test, y_test)
# +
# show sample prediction
n_samples = 10
y_pred = model.predict(x_test[:n_samples])
show_images(x_test[:n_samples], y_pred)
| notebooks/3.0 Neural Networks/6.2 cifar classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 逻辑斯特回归示例
#
# - [逻辑斯特回归](#逻辑斯特回归)
# - [正则化后的逻辑斯特回归](#加正则化项的逻辑斯特回归)
# +
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib as mpl
import matplotlib.pyplot as plt
from scipy.optimize import minimize
from sklearn.preprocessing import PolynomialFeatures
pd.set_option('display.notebook_repr_html', False)
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', 150)
pd.set_option('display.max_seq_items', None)
sns.set_context('notebook')
sns.set_style('white')
# %matplotlib inline
# -
def load_data(file, delimeter=','):
data = np.loadtxt(file, delimiter=delimeter)
print('load_data: dimensions: ',data.shape)
print(data[1:6,:])
return data
def plot_data(data, label_x, label_y, label_pos, label_neg, axes=None):
if axes == None: axes = plt.gca()
# 获得正负样本的下标(即哪些是正样本,哪些是负样本)
neg = data[:,2] == 0
pos = data[:,2] == 1
axes.scatter(data[pos][:,0], data[pos][:,1], marker='+', c='k',
s=60, linewidth=2, label=label_pos)
axes.scatter(data[neg][:,0], data[neg][:,1], marker='o', c='y',
s=60, label=label_neg)
axes.set_xlabel(label_x)
axes.set_ylabel(label_y)
axes.legend(frameon= True, fancybox = True);
# ### 逻辑斯特回归
data = load_data('input/data1.txt', ',')
# +
# X = np.c_[np.ones((data.shape[0],1)), data[:,0:2]]
# y = np.c_[data[:,2]]
# -
plot_data(data, 'Exam 1 score', 'Exam 2 score', 'Pass', 'Fail')
# #### 逻辑斯特回归假设
# #### $$ h_{\theta}(x) = g(\theta^{T}x)$$
# #### $$ g(z)=\frac{1}{1+e^{−z}} $$
#定义sigmoid函数
def sigmoid(z):
return(1 / (1 + np.exp(-z)))
# 其实scipy包里有一个函数可以完成一样的功能:<BR>
# http://docs.scipy.org/doc/scipy/reference/generated/scipy.special.expit.html#scipy.special.expit
# #### 损失函数
# #### $$ J(\theta) = \frac{1}{m}\sum_{i=1}^{m}\big[-y^{(i)}\, log\,( h_\theta\,(x^{(i)}))-(1-y^{(i)})\,log\,(1-h_\theta(x^{(i)}))\big]$$
# #### 向量化的损失函数(矩阵形式)
# #### $$ J(\theta) = \frac{1}{m}\big((\,log\,(g(X\theta))^Ty+(\,log\,(1-g(X\theta))^T(1-y)\big)$$
#定义损失函数
def costFunction(theta, X, y):
m = y.size
h = sigmoid(X.dot(theta))
J = -1.0*(1.0/m)*(np.log(h).T.dot(y)+np.log(1-h).T.dot(1-y))
if np.isnan(J[0]):
return(np.inf)
return J[0]
# #### 求偏导(梯度)
#
# #### $$ \frac{\delta J(\theta)}{\delta\theta_{j}} = \frac{1}{m}\sum_{i=1}^{m} ( h_\theta (x^{(i)})-y^{(i)})x^{(i)}_{j} $$
# #### 向量化的偏导(梯度)
# #### $$ \frac{\delta J(\theta)}{\delta\theta_{j}} = \frac{1}{m} X^T(g(X\theta)-y)$$
#
#求解梯度
def gradient(theta, X, y):
m = y.size
h = sigmoid(X.dot(theta.reshape(-1,1)))
grad =(1.0/m)*X.T.dot(h-y)
return(grad.flatten())
initial_theta = np.zeros(X.shape[1])
cost = costFunction(initial_theta, X, y)
grad = gradient(initial_theta, X, y)
print('Cost: \n', cost)
print('Grad: \n', grad)
# #### 最小化损失函数
res = minimize(costFunction, initial_theta, args=(X,y), jac=gradient, options={'maxiter':400})
res
# #### 做一下预测吧
def predict(theta, X, threshold=0.5):
p = sigmoid(X.dot(theta.T)) >= threshold
return(p.astype('int'))
# #### 咱们来看看考试1得分45,考试2得分85的同学通过概率有多高
sigmoid(np.array([1, 45, 85]).dot(res.x.T))
# #### 画决策边界
plt.scatter(45, 85, s=60, c='r', marker='v', label='(45, 85)')
plotData(data, 'Exam 1 score', 'Exam 2 score', 'Admitted', 'Not admitted')
x1_min, x1_max = X[:,1].min(), X[:,1].max(),
x2_min, x2_max = X[:,2].min(), X[:,2].max(),
xx1, xx2 = np.meshgrid(np.linspace(x1_min, x1_max), np.linspace(x2_min, x2_max))
h = sigmoid(np.c_[np.ones((xx1.ravel().shape[0],1)), xx1.ravel(), xx2.ravel()].dot(res.x))
h = h.reshape(xx1.shape)
plt.contour(xx1, xx2, h, [0.5], linewidths=1, colors='b');
# ### 加正则化项的逻辑斯特回归
data2 = loaddata('input/data2.txt', ',')
# 拿到X和y
y = np.c_[data2[:,2]]
X = data2[:,0:2]
# 画个图
plotData(data2, 'Microchip Test 1', 'Microchip Test 2', 'y = 1', 'y = 0')
# #### 咱们整一点多项式特征出来(最高6阶)
poly = PolynomialFeatures(6)
XX = poly.fit_transform(data2[:,0:2])
# 看看形状(特征映射后x有多少维了)
XX.shape
# #### 正则化后损失函数
# #### $$ J(\theta) = \frac{1}{m}\sum_{i=1}^{m}\big[-y^{(i)}\, log\,( h_\theta\,(x^{(i)}))-(1-y^{(i)})\,log\,(1-h_\theta(x^{(i)}))\big] + \frac{\lambda}{2m}\sum_{j=1}^{n}\theta_{j}^{2}$$
# #### 向量化的损失函数(矩阵形式)
# #### $$ J(\theta) = \frac{1}{m}\big((\,log\,(g(X\theta))^Ty+(\,log\,(1-g(X\theta))^T(1-y)\big) + \frac{\lambda}{2m}\sum_{j=1}^{n}\theta_{j}^{2}$$
# 定义损失函数
def costFunctionReg(theta, reg, *args):
m = y.size
h = sigmoid(XX.dot(theta))
J = -1.0*(1.0/m)*(np.log(h).T.dot(y)+np.log(1-h).T.dot(1-y)) + (reg/(2.0*m))*np.sum(np.square(theta[1:]))
if np.isnan(J[0]):
return(np.inf)
return(J[0])
# #### 偏导(梯度)
#
# #### $$ \frac{\delta J(\theta)}{\delta\theta_{j}} = \frac{1}{m}\sum_{i=1}^{m} ( h_\theta (x^{(i)})-y^{(i)})x^{(i)}_{j} + \frac{\lambda}{m}\theta_{j}$$
# #### 向量化的偏导(梯度)
# #### $$ \frac{\delta J(\theta)}{\delta\theta_{j}} = \frac{1}{m} X^T(g(X\theta)-y) + \frac{\lambda}{m}\theta_{j}$$
# ##### $$\text{注意,我们另外自己加的参数 } \theta_{0} \text{ 不需要被正则化}$$
def gradientReg(theta, reg, *args):
m = y.size
h = sigmoid(XX.dot(theta.reshape(-1,1)))
grad = (1.0/m)*XX.T.dot(h-y) + (reg/m)*np.r_[[[0]],theta[1:].reshape(-1,1)]
return(grad.flatten())
initial_theta = np.zeros(XX.shape[1])
costFunctionReg(initial_theta, 1, XX, y)
# +
fig, axes = plt.subplots(1,3, sharey = True, figsize=(17,5))
# 决策边界,咱们分别来看看正则化系数lambda太大太小分别会出现什么情况
# Lambda = 0 : 就是没有正则化,这样的话,就过拟合咯
# Lambda = 1 : 这才是正确的打开方式
# Lambda = 100 : 卧槽,正则化项太激进,导致基本就没拟合出决策边界
for i, C in enumerate([0.0, 1.0, 100.0]):
# 最优化 costFunctionReg
res2 = minimize(costFunctionReg, initial_theta, args=(C, XX, y), jac=gradientReg, options={'maxiter':3000})
# 准确率
accuracy = 100.0*sum(predict(res2.x, XX) == y.ravel())/y.size
# 对X,y的散列绘图
plotData(data2, 'Microchip Test 1', 'Microchip Test 2', 'y = 1', 'y = 0', axes.flatten()[i])
# 画出决策边界
x1_min, x1_max = X[:,0].min(), X[:,0].max(),
x2_min, x2_max = X[:,1].min(), X[:,1].max(),
xx1, xx2 = np.meshgrid(np.linspace(x1_min, x1_max), np.linspace(x2_min, x2_max))
h = sigmoid(poly.fit_transform(np.c_[xx1.ravel(), xx2.ravel()]).dot(res2.x))
h = h.reshape(xx1.shape)
axes.flatten()[i].contour(xx1, xx2, h, [0.5], linewidths=1, colors='g');
axes.flatten()[i].set_title('Train accuracy {}% with Lambda = {}'.format(np.round(accuracy, decimals=2), C))
# -
| dev/statistical_ml/logistic_regression/ML001A_logistic_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # get predictions from final models
# +
import gc
import h5py
import numpy as np
import pandas as pd
import tensorflow as tf
from keras.layers import Input, Dense, Dropout
from keras import optimizers, regularizers, losses
from keras.models import Model
from keras import backend as K
from keras.callbacks import CSVLogger
from keras import metrics
import scipy
import datetime
import h5py
import keras
import sys
from sklearn.model_selection import ParameterGrid
import pickle
import os
sys.path.append("/home/nbbwang/bigdrive/AD_Project/")
from IntegratedGradients import *
# +
from matplotlib import pyplot as plt
# %matplotlib inline
import matplotlib
matplotlib.rcParams.update({'font.size': 18, 'font.family': "Times New Roman"})
plt.rcParams["font.family"] = "Times"
import IntegratedGradients as IG
import keras
from scipy import stats
path_to_configs = "../"
sys.path.append(path_to_configs)
from configs import *
from models import *
# -
MTL_final_final_model = pickle.load(open(path_to_configs+path_to_final_chosen_models + "MTL/final.p", "rb" ) )
baselines_final_final_model = pickle.load(open(path_to_configs+path_to_final_chosen_models + "MLP_baselines/final.p", "rb" ))
phens = ["CERAD", "BRAAK", "PLAQUES", "TANGLES", "ABETA_IHC", "TAU_IHC"]
path_to_ext_val_predictions = path_to_configs + path_to_ext_val_results
for dpath in ["", "_intersection"]:
for dset in os.listdir("%spredictions%s/"%(path_to_ext_val_predictions, dpath)):
if dpath=="_intersection":
dsets_to_check = ["train","test"]
else:
dsets_to_check = ["test"]
######### MD-AD AVERAGING ################
for dset_to_check in dsets_to_check:
all_pred_vals = []
if dset_to_check == "train":
path_to_preds = "%spredictions%s/%s/MTL/%s/train/"%(path_to_ext_val_predictions, dpath,dset, MTL_final_final_model)
else:
path_to_preds = "%spredictions%s/%s/MTL/%s/"%(path_to_ext_val_predictions, dpath,dset, MTL_final_final_model)
print("Saving consensus %s predictions to %s"%(dset_to_check,path_to_preds))
for f in os.listdir(path_to_preds):
if os.path.isdir(path_to_preds + f):
continue
pred_df = pd.read_csv("%s/%s"%(path_to_preds,f), index_col="Unnamed: 0")
all_pred_vals.append(pred_df.values)
final_preds = pd.DataFrame(np.mean(np.array(all_pred_vals),axis=0), columns=pred_df.columns)
if dset_to_check == "train":
final_preds.to_csv("%spredictions%s/%s/MTL/final_train.csv"%(path_to_ext_val_predictions,dpath,dset))
else:
final_preds.to_csv("%spredictions%s/%s/MTL/final.csv"%(path_to_ext_val_predictions, dpath,dset))
all_pred_vals = []
############## MLP averaging #######################
path_to_preds = "%spredictions%s/%s/MLP_baselines/"%(path_to_ext_val_predictions, dpath,dset)
print("Saving consensus predictions to %s"%path_to_preds)
for f in os.listdir(path_to_preds):
if f.split(".")[0] in [str(i) for i in range(100)]:
pred_df = pd.read_csv("%s/%s"%(path_to_preds,f), index_col="Unnamed: 0")
all_pred_vals.append(pred_df.values)
final_preds = pd.DataFrame(np.mean(np.array(all_pred_vals),axis=0), columns=pred_df.columns)
final_preds.to_csv(path_to_preds + "final.csv")
for f in os.listdir("%spredictions/"%(path_to_ext_val_predictions)):
for elt in os.listdir("%spredictions/%s/MLP_baselines/"%(path_to_ext_val_predictions,f)):
if "_" in elt:
print(subprocess.run(["rm", "-r", "%spredictions/%s/MLP_baselines/%s"%(path_to_ext_val_predictions,f, elt)]))
| Pipeline_Code/External_Validation/.ipynb_checkpoints/compute_final_model_predictions-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pygame
import numpy as np
import matplotlib.pyplot as plt
import random
pygame.init()
# -
def generate_danger_locations(number_of_locations):
Danger_location = []
for i in range(number_of_locations):
location =(random.randint(20,1180),random.randint(20,580))
Danger_location.append(location)
return Danger_location
Screen_width = 1200
Screen_height = 600
Start_position = (600,300)
Goal_position = (500,300)
number_of_danger_locations = 10
Danger_location = generate_danger_locations(number_of_danger_locations)
Actions = {0:"left",1:"right",2:"up",3:"down"}
game_over = False
exit_game = False
white = (255,255,255)
red = (255,0,0)
black = (0,0,0)
blue = (125,125,125)
agent_size = 20
danger_size = 20
fps = 30
jump_size = 5
class Enviornment:
def __init__(self):
self.danger_locations =Danger_location
self.start_pos =Start_position
self.goal_pos = Goal_position
self.actions =Actions
self.width = Screen_width
self.height = Screen_height
self.state_action_values = {}
self.current_position = Start_position
self.reward = 0
for x in range(self.width):
for y in range(self.height):
state = (x,y)
self.state_action_values[state] = [0.25 for i in range(len(self.actions))]
self.state_action_values[self.goal_pos] = [0.0 for i in range(len(self.actions))]
def nextState(self,action,current_state):
if self.actions[action] == "left":
newPossiblePos = (current_state[0]-jump_size,current_state[1])
if newPossiblePos[0]>0:
self.current_position = newPossiblePos
else:
pass
elif self.actions[action] == "right":
newPossiblePos = (current_state[0]+jump_size,current_state[1])
if newPossiblePos[0]<self.width:
self.current_position = newPossiblePos
else:
pass
elif self.actions[action] == "up":
newPossiblePos = (current_state[0],current_state[1]-jump_size)
if newPossiblePos[0]>0:
self.current_position = newPossiblePos
else:
pass
elif self.actions[action] == "down":
newPossiblePos = (current_state[0],current_state[1]+jump_size)
if newPossiblePos[0]<self.height:
self.current_position = newPossiblePos
else:
pass
return self.current_position
def giveReward(self):
if abs(self.current_position[0]-self.goal_pos[0])< agent_size or abs(self.current_position[1]-self.goal_pos[1])< agent_size:
self.reward = 100
elif self.current_position in self.danger_locations:
selfreward = -100
elif self.current_position not in self.danger_locations:
for state in self.danger_locations:
if abs(self.current_position[0]-state[0]) < agent_size or abs(self.current_position[1]-state[1])< agent_size:
self.reward = -10
else:
self.reward = 0
return self.reward
class Agent:
def __init__(self,discount_factor = 0.9, alpha = 0.1, epsilon = 0.2, max_steps = 200, num_episodes = 10):
self.enviornment = Enviornment()
self.epsilon = epsilon
self.num_episodes=num_episodes
self.alpha = alpha
self.max_steps = max_steps
self.gamma = discount_factor
self.TotalReward = 0
self.Totalreward_per_episode = []
self.Terminal = False
def chooseAction(self,state):
StateActionValues = self.enviornment.state_action_values
action_values = StateActionValues[state]
if self.epsilon < random.random():
action_values = np.asarray(action_values)
choosen_action = np.argmax(action_values)
else:
choosen_action = random.randint(0,len(self.enviornment.actions)-1)
return choosen_action
def play_game(self):
exit_game = False
game_over = False
clock = pygame.time.Clock()
gameWindow = pygame.display.set_mode((Screen_width,Screen_height))
pygame.display.set_caption("Life_runner")
pygame.display.update()
font = pygame.font.SysFont(None,55)
for i in range(self.num_episodes):
S = Start_position
agent_pos_x = Start_position[0]
agent_pos_y = Start_position[1]
steps = 0
self.Terminal = False
exit_game = False
while not exit_game and steps < self.max_steps and self.Terminal != True:
for event in pygame.event.get():
if event.type == pygame.QUIT:
exit_game = True
gameWindow.fill(white)#fill the window with white color
pygame.draw.rect(gameWindow,blue,[agent_pos_x,agent_pos_y,agent_size,agent_size])
pygame.draw.rect(gameWindow,black,[self.enviornment.goal_pos[0],self.enviornment.goal_pos[1],agent_size,agent_size])
for state in self.enviornment.danger_locations:
pygame.draw.rect(gameWindow,red,[state[0],state[1],danger_size,danger_size])
A = self.chooseAction(S)
S_prime = self.enviornment.nextState(A,S)
self.enviornment.current_position = S_prime
R = self.enviornment.giveReward()
agent_pos_x,agent_pos_y = S_prime[0],S_prime[1]
if S_prime == Goal_position:
self.Terminal = True
self.enviornment.state_action_values[S][A] += self.alpha*(R -self.enviornment.state_action_values[S][A])
break
else:
self.enviornment.state_action_values[S][A] += self.alpha*(R +self.gamma*max(self.enviornment.state_action_values[S_prime])-self.enviornment.state_action_values[S][A])
S = S_prime
steps += 1
pygame.display.update()
clock.tick(fps)
pygame.quit()
def return_stateActionValues(self):
return self.enviornment.state_action_values
agent = Agent()
| GridWorld_RL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from plotnine import *
from plotnine.data import *
# %matplotlib inline
# -
# ### Density Plot
mpg.head()
# The defaults are not exactly beautiful, but still quite clear.
(
ggplot(mpg, aes(x='cty'))
+ geom_density()
)
# Plotting multiple groups is straightforward, but as each group is plotted as an independent PDF summing to 1, the relative size of each group will be normalized.
(
ggplot(mpg, aes(x='cty', color='drv', fill='drv'))
+ geom_density(alpha=0.1)
)
# To plot multiple groups and scale them by their relative size, you can pass `stat(count)` to the mapping.
(
ggplot(mpg, aes(x='cty', color='drv', fill='drv'))
+ geom_density(
aes(y=after_stat('count')),
alpha=0.1
)
)
| plotnine_examples/examples/geom_density.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Applicazione del metodo del perceptron per la separazione lineare
import pandas as pd
import numpy as np
import seaborn as sns
# +
import matplotlib.pyplot as plt
import matplotlib.colors as mcolors
from matplotlib import cm
plt.style.use('fivethirtyeight')
plt.rcParams['font.family'] = 'sans-serif'
plt.rcParams['font.serif'] = 'Ubuntu'
plt.rcParams['font.monospace'] = 'Ubuntu Mono'
plt.rcParams['font.size'] = 10
plt.rcParams['axes.labelsize'] = 10
plt.rcParams['axes.labelweight'] = 'bold'
plt.rcParams['axes.titlesize'] = 10
plt.rcParams['xtick.labelsize'] = 8
plt.rcParams['ytick.labelsize'] = 8
plt.rcParams['legend.fontsize'] = 10
plt.rcParams['figure.titlesize'] = 12
plt.rcParams['image.cmap'] = 'jet'
plt.rcParams['image.interpolation'] = 'none'
plt.rcParams['figure.figsize'] = (16, 8)
plt.rcParams['lines.linewidth'] = 2
plt.rcParams['lines.markersize'] = 8
colors = ['xkcd:pale orange', 'xkcd:sea blue', 'xkcd:pale red', 'xkcd:sage green', 'xkcd:terra cotta', 'xkcd:dull purple', 'xkcd:teal', 'xkcd:goldenrod', 'xkcd:cadet blue',
'xkcd:scarlet']
cmap_big = cm.get_cmap('Spectral', 512)
cmap = mcolors.ListedColormap(cmap_big(np.linspace(0.7, 0.95, 256)))
# -
# normalizzazione dei valori delle features a media 0 e varianza 1
def normalizza(X):
mu = np.mean(X, axis=0)
sigma = np.std(X, axis=0, ddof=1)
return (X-mu)/sigma
# produce e restituisce una serie di statistiche
def statistics(theta,X,t):
# applica il modello
y = np.dot(X,theta)
# attribuisce i punti alle due classi
y = np.where(y>0, 1, 0)
# costruisce la confusion matrix
confmat = np.zeros((2, 2))
for i in range(2):
for j in range(2):
confmat[i,j] = np.sum(np.where(y==i,1,0)*np.where(t==j,1,0))
return confmat
# legge i dati in dataframe pandas
# +
data = pd.read_csv("../dataset/ex2data1.txt", header=0, delimiter=',', names=['x1','x2','t'])
# calcola dimensione dei dati
n = len(data)
# calcola dimensionalità delle features
nfeatures = len(data.columns)-1
X = np.array(data[['x1','x2']])
t = np.array(data['t']).reshape(-1,1)
# -
c=[colors[i] for i in np.nditer(t)]
deltax = max(X[:,0])-min(X[:,0])
deltay = max(X[:,1])-min(X[:,1])
minx = min(X[:,0])-deltax/10.0
maxx = max(X[:,0])+deltax/10.0
miny = min(X[:,1])-deltay/10.0
maxy = max(X[:,1])+deltay/10.0
fig = plt.figure(figsize=(10,10))
fig.patch.set_facecolor('white')
ax = fig.gca()
ax.scatter(X[:,0],X[:,1],s=40,c=c, marker='o', alpha=.7)
t1 = np.arange(minx, maxx,0.01)
plt.xlabel('x1', fontsize=10)
plt.ylabel('x2', fontsize=10)
plt.xticks(fontsize=10)
plt.yticks(fontsize=10)
plt.show()
X=normalizza(X)
# aggiunge una colonna di valori -1 alla matrice delle features
X= np.column_stack((-np.ones(n), X))
# +
# fissa il valore del parametro eta
eta = 0.25
# fissa una valore iniziale casuale per i coefficienti
theta = np.random.rand(nfeatures+1, 1) * 0.1 - 0.05
# -
deltax = max(X[:,1])-min(X[:,1])
deltay = max(X[:,2])-min(X[:,2])
minx = min(X[:,1])-deltax/10.0
maxx = max(X[:,1])+deltax/10.0
miny = min(X[:,2])-deltay/10.0
maxy = max(X[:,2])+deltay/10.0
t1 = np.arange(minx, maxx, 0.01)
u = np.linspace(minx, maxx, 100)
v = np.linspace(miny, maxy, 100)
#u, v = np.meshgrid(u, v)
z = np.zeros((len(u), len(v)))
plt.figure()
costs = []
# esegue il ciclo seguente per n volte
for k in range(2*n+1):
# determina la classificazione effettuata applicando la soglia 0-1 ai valori forniti
# dalla combinazione lineare delle features e dei coefficienti
y = np.where(np.dot(X, theta)>0,1,0)
cf=statistics(theta,X,t)
costs.append(round(np.trace(cf)/n,2))
# somma o sottrae eta da tutti i coefficienti per ogni osservazione mal classificata
if k in set([0,1,2,3]):
cf=statistics(theta,X,t)
theta1=-theta[1]/theta[2]
theta0=theta[0]/theta[2]
t2 = theta0+theta1*t1
for i in range(0, len(u)):
for j in range(0, len(v)):
z[i,j] = np.dot(theta.T, np.array([u[i],v[j],-1]).reshape(-1,1)).item()
plt.subplot(2,2,k+1)
#imshow_handle = plt.imshow(z, extent=(minx, maxx, miny, maxy), alpha=.5)
plt.scatter(X[:,1],X[:,2],s=20,c=c, marker='o', alpha=.7)
plt.plot(t1, theta0+theta1*t1, color=colors[3], linewidth=1.5, alpha=1)
plt.xlabel('x1', fontsize=10)
plt.ylabel('x2', fontsize=10)
plt.xticks(fontsize=10)
plt.yticks(fontsize=10)
plt.xlim(minx,maxx)
plt.ylim(miny,maxy)
plt.annotate(str(k)+': '+str(round(np.trace(cf)/n,2)), xy=(.03, .97), backgroundcolor='w', va='top', xycoords='axes fraction', fontsize=8)
plt.show()
theta += eta * np.dot(X.T, t-y)
costs
cf=statistics(theta,X,t)
print('Veri negativi (elementi di C0 classificati correttamente): {0:d}'.format(int(cf[0,0])))
print('Falsi negativi (elementi di C0 classificati in modo erroneo): {0:d}'.format(int(cf[0,1])))
print('Falsi positivi (elementi di C1 classificati in modo erroneo): {0:d}'.format(int(cf[1,0])))
print('Veri positivi (elementi di C1 classificati correttamente): {0:d}'.format(int(cf[1,1])))
plt.figure()
theta1=-theta[1]/theta[2]
theta0=theta[0]/theta[2]
t2 = theta0+theta1*t1
for i in range(0, len(u)):
for j in range(0, len(v)):
z[i,j] = np.dot(theta.T, np.array([u[i],v[j],-1]).reshape(-1,1)).item()
plt.scatter(X[:,1],X[:,2],s=40,c=c, marker='o', alpha=.7)
plt.plot(t1, theta0+theta1*t1, color=colors[3], linewidth=1.5, alpha=1)
plt.xlabel('x1', fontsize=10)
plt.ylabel('x2', fontsize=10)
plt.xticks(fontsize=10)
plt.yticks(fontsize=10)
plt.xlim(minx,maxx)
plt.ylim(miny,maxy)
plt.annotate(str(k)+': '+str(round(np.trace(cf)/n,2)), xy=(.03, .97), backgroundcolor='w', va='top', xycoords='axes fraction', fontsize=8)
plt.show()
| codici/old/Untitled2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ###### Content provided under a Creative Commons Attribution license, CC-BY 4.0; code under MIT license (c)2014 <NAME>, <NAME>. Thanks: NSF for support via CAREER award #1149784.
# ##### Version 0.3 -- March 2014
# # Source Sheet
# A source sheet is an infinite row of sources along one line. You have already created a vortex sheet by placing a [row of vortices](05_Lesson05_InfiniteRowOfVortices.ipynb) on one line, so you are ready for this.
#
# Make sure that you have studied [*AeroPython* Lesson 7](07_Lesson07_methodOfImages.ipynb), on the Method of Images and the use of classes in Python. From now on, classes will feature prominently!
#
# We start by importing our libraries and creating the grid of points, as we've done so many times before.
import numpy
import math
from matplotlib import pyplot
# embed the figures into the notebook
# %matplotlib inline
N = 100 # Number of points in each direction
x_start, x_end = -1.0, 1.0 # x-direction boundaries
y_start, y_end = -1.5, 1.5 # y-direction boundaries
x = numpy.linspace(x_start, x_end, N) # computes a 1D-array for x
y = numpy.linspace(y_start, y_end, N) # computes a 1D-array for y
X, Y = numpy.meshgrid(x, y) # generates a mesh grid
# We will include a uniform flow of magnitude $U_\infty = 1$ parallel to the horizontal axis. The arrays `u_freestream` and `v_freestream` contain the velocity components of the free stream. Let's fill these up:
# +
u_inf = 1.0 # free-stream speed
# calculate the free-stream velocity components
u_freestream = u_inf * numpy.ones((N, N), dtype=float)
v_freestream = numpy.zeros((N, N), dtype=float)
# -
# ### Finite number of sources along a line
# We consider first a finite number of sources arranged along a vertical line, normal to the freestream. The streamlines will come out of each source and be deflected by the streamlines coming from the freestream.
#
# From a computational point of view, the finite number of sources can be represented by a 1D-array containing objects, each one having been created by the class `Source`. This class will need as attributes the strength of a source, `strength`, and its location at (`x`,`y`). Its methods will compute the velocity components and stream function on a given mesh grid (`X`,`Y`), and of course, its *constructor*:
class Source:
"""
Contains information related to a source/sink.
"""
def __init__(self, strength, x, y):
"""
Sets the location and strength of the singularity.
Parameters
----------
strength: float
Strength of the source/sink.
x: float
x-coordinate of the source/sink.
y: float
y-coordinate of the source/sink.
"""
self.strength = strength
self.x, self.y = x, y
def velocity(self, X, Y):
"""
Computes the velocity field generated by the source/sink.
Parameters
----------
X: 2D Numpy array of floats
x-coordinate of the mesh points.
Y: 2D Numpy array of floats
y-coordinate of the mesh points.
"""
self.u = (self.strength / (2 * math.pi) *
(X - self.x) / ((X - self.x)**2 + (Y - self.y)**2))
self.v = (self.strength / (2 * math.pi) *
(Y - self.y) / ((X - self.x)**2 + (Y - self.y)**2))
def stream_function(self, X, Y):
"""
Computes the stream-function generated by the source/sink.
Parameters
----------
X: 2D Numpy array of floats
x-coordinate of the mesh points.
Y: 2D Numpy array of floats
y-coordinate of the mesh points.
"""
self.psi = (self.strength / (2 * math.pi) *
numpy.arctan2((Y - self.y), (X - self.x)))
# Now you'll really see how useful classes are! We will use our class `Source` to create as many sources as we want to place on a source sheet. Pick a number, `N_sources`, and call the class constructor that many times, moving along the $y$ axis for the location of the sources. Then call the function to calculate the velocity of each source.
# +
N_sources = 11 # number of sources
strength = 5.0 # sum of all source strengths
strength_source = strength / N_sources # strength of one source
x_source = numpy.zeros(N_sources, dtype=float) # horizontal position of all sources (1D array)
y_source = numpy.linspace(-1.0, 1.0, N_sources) # vertical position of all sources (1D array)
# create a source line (Numpy array of Source objects)
sources = numpy.empty(N_sources, dtype=object)
for i in range(N_sources):
sources[i] = Source(strength_source, x_source[i], y_source[i])
sources[i].velocity(X, Y)
# superposition of all sources to the free-stream flow
u = u_freestream.copy()
v = v_freestream.copy()
for source in sources:
u += source.u
v += source.v
# -
# Notice that the variable `sources` is a NumPy array, that is, a collection of items of the same type. What type? You're probably used to arrays of numbers, but we can also have arrays of objects of any type created by a class (in this case, of type `Source`).
#
# We are creating an empty NumPy array called `sources`, and telling Python that its items will be some object that is not a built-in data type, like `int` or `float` for integers or floating-point real numbers. The number of items contained in the array will be `N_sources`.
#
# In the first loop, we fill the `sources` array, calling the `Source`-class *constructor* for each item. We also go ahead and compute the velocity of each source (using the method `velocity()` of each source). In the second loop—after creating the velocity arrays `u` and `v` by copying the free-stream velocity components—, we're adding all the velocity contributions for every source `source` in the array `sources`.
#
# The final flow pattern corresponds to the superposition of a uniform flow and `N_sources` identical sources of strength `strength_source` equally spaced along a vertical line normal to the uniform flow. In our plot, we'll mark the location of the sources with red points and also define a filled contour of velocity magnitude to visualize the location of the stagnation point, defined by $u=0$ and $v=0$.
# plot the streamlines
width = 4
height = (y_end - y_start) / (x_end - x_start) * width
pyplot.figure(figsize=(width, height))
pyplot.grid()
pyplot.xlabel('x', fontsize=16)
pyplot.ylabel('y', fontsize=16)
pyplot.streamplot(X, Y, u, v,
density=2, linewidth=1, arrowsize=1, arrowstyle='->')
# plot the sources
pyplot.scatter(x_source, y_source,
color='#CD2305', s=80, marker='o')
# compute the velocity magniture and indices of the stagnation point
# note: the stagnation point is approximated as the point with the smallest velocity magnitude
magnitude = numpy.sqrt(u**2 + v**2)
j_stagn, i_stagn = numpy.unravel_index(magnitude.argmin(), magnitude.shape)
# plot the stagnation point
pyplot.scatter(x[i_stagn], y[j_stagn],
color='black', s=40, marker='D')
pyplot.xlim(x_start, x_end)
pyplot.ylim(y_start, y_end);
# Now try to change the total strength of the sources. What is the minimal total source strength so that the dividing streamline ($\psi = 0$) encloses all the singularities?
# ### Infinite line of sources
# By definition, a *source sheet* is an infinite collection of side-by-side sources of infinitesimal equal strength distributed along a given path.
#
# Consider $s$ to be the running coordinate along the sheet: we define $\sigma = \sigma(s)$ as the strength per unit length.
#
# Recall that the strength of a single source represents the volume flow rate per unit depth (i.e., per unit length in the $z$-direction).
# Therefore, $\sigma$ represents the volume flow rate per unit depth and per unit length (in the $s$-direction). Thus, $\sigma ds$ is the strength of an infinitesimal portion $ds$ of the source sheet. This infinitesimal portion is so small that it can be treated as a distinct source of strength $\sigma ds$.
#
# Following this, the stream-function at point $\left(r,\theta\right)$ of this infinitesimal portion is
#
# $$d\psi\left(r,\theta\right) = \frac{\sigma ds}{2\pi} \theta$$
#
# Integrating along $s$, we find the stream-function of the entire source sheet:
#
# $$\psi\left(r,\theta\right) = \frac{\sigma}{2\pi}\int_{\text{sheet}}\theta\left(s\right)ds$$
#
# In the previous section, we considered a vertical finite distribution of sources. Similarly, the stream function of a vertical source sheet at $x=0$ between $y_{\text{min}}$ and $y_{\text{max}}$ in Cartesian coordinates is
#
# $$\psi\left(x,y\right) = \frac{\sigma}{2\pi}\int_{y_{\text{min}}}^{y_{\text{max}}} \tan^{-1}\left(\frac{y-\xi}{x}\right)d\xi$$
#
# And the velocity components are
#
# $$u\left(x,y\right) = \frac{\sigma}{2\pi} \int_{y_{\text{min}}}^{y_{\text{max}}} \frac{x}{x^2+\left(y-\xi\right)^2}d\xi$$
#
# $$v\left(x,y\right) = \frac{\sigma}{2\pi} \int_{y_{\text{min}}}^{y_{\text{max}}} \frac{y-\xi}{x^2+\left(y-\xi\right)^2}d\xi$$
# ### Using SciPy
#
# We need to calculate the two integrals above to obtain the velocities. Bummer, you say. Calculating integrals is so passé. Don't worry! We have SciPy: a powerful collection of mathematical algorithms and functions. It includes the module [`integrate`](http://docs.scipy.org/doc/scipy/reference/tutorial/integrate.html), a fundamental set of functions for scientific computing including basic integration, quadrature rules and numerical integrators for ordinary differential equations. How helpful is that?
#
# Let's import the module we need from SciPy:
from scipy import integrate
# We are going to use the function [`quad(func,a,b)`](http://docs.scipy.org/doc/scipy/reference/generated/scipy.integrate.quad.html#scipy.integrate.quad) of the module `integrate` to compute the definite integral of a function of one variable between two limits:
#
# $$I=\int_a^b f(x)\ {\rm d}x$$
#
# The first argument to the function `quad` needs to be a Python function ... you already know how to create a Python function using a `def` statement, but you'll learn a new way to create Python functions: the `lambda` statement. Why do I need two ways to define functions, you may ask? The answer is rather subtle, and the subject of full blog posts! (See below)
#
# In short, a function created with a `lambda` statement is a single expression that returns a value (but without a `return` statement!). It is often called an *anonymous function*, because we don't need to give it a name.
# Let's see how we use a `lambda` statement to integrate a mathematical function. Suppose we want to integrate the function $f:x\rightarrow x^2$ from $0$ to $1$. You can manually do this one, right? It gives the value $\frac{1}{3}$.
#
# To use the function `quad()`, we give it as first parameter the expression `lambda x : x**2`:
print(integrate.quad(lambda x: x**2, 0.0, 1.0))
# You see, here we used the `lambda` statement to pass a function argument to `quad()`, without going through a function definition (in this sense, we talk of *anonymous* Python functions).
#
# Note that the function `quad()` returns a list: the first element is the result of the integral, and the second element is an estimate of the error. If you just want the value of the definite integral, you need to specify the index `[0]`, for the first element of the list!
#
# Note that the lambda function can accept several arguments:
a = 3.0
print(integrate.quad(lambda x, a: a * x**2, 0.0, 1.0, args=a))
b = 2.0
print(integrate.quad(lambda x, a, b: a * b * x**2, 0.0, 1.0, args=(a, b)))
# We are ready to use this for our velocity calculations of the source sheet.
#
# One last thing! The result of the integral, obtained with the function `quad()` is a float, so we would have to loop over our domain to compute the integral at each discrete point; this can become expensive as we refine our domain...
# Unless we use [`numpy.vectorize()`](https://docs.scipy.org/doc/numpy/reference/generated/numpy.vectorize.html) that will allow us to create a vectorized function that takes Numpy arrays as inputs and returns a Numpy array!
# +
sigma = 2.5 # strength of the source-sheet
# boundaries of the source-sheet
y_min, y_max = -1.0, 1.0
# create the anonymous functions
integrand_u = lambda s, x, y: x / (x**2 + (y - s)**2)
integrand_v = lambda s, x, y: (y - s) / (x**2 + (y - s)**2)
# create a dummy function to vectorize
def integration(x, y, integrand):
return integrate.quad(integrand, y_min, y_max, args=(x, y))[0]
vec_integration = numpy.vectorize(integration)
# compute the velocity field generated by the source-sheet
u_sheet = sigma / (2.0 * numpy.pi) * vec_integration(X, Y, integrand_u)
v_sheet = sigma / (2.0 * numpy.pi) * vec_integration(X, Y, integrand_v)
# superposition of the source-sheet to the uniform flow
u = u_freestream + u_sheet
v = v_freestream + v_sheet
# -
# Let's now visualize the streamlines (blue) on the mesh grid. We draw a red line to show the source sheet and a filled contour to visualize the location of the stagnation point.
# plot the streamlines
width = 4
height = (y_end - y_start) / (x_end - x_start) * width
pyplot.figure(figsize=(width, height))
pyplot.grid()
pyplot.xlabel('x', fontsize=16)
pyplot.ylabel('y', fontsize=16)
pyplot.streamplot(X, Y, u, v,
density=2, linewidth=1, arrowsize=1, arrowstyle='->')
# plot the source sheet
pyplot.axvline(0.0,
(y_min - y_start) / (y_end - y_start),
(y_max - y_start) / (y_end - y_start),
color='#CD2305', linewidth=4)
# compute the velocity magniture and indices of the stagnation point
# note: stagnation point approximated as point with smallest velocity magnitude
magnitude = numpy.sqrt(u**2 + v**2)
j_stagn, i_stagn = numpy.unravel_index(magnitude.argmin(), magnitude.shape)
# plot the stagnation point
pyplot.scatter(x[i_stagn], y[j_stagn],
color='black', s=40, marker='D')
pyplot.xlim(x_start, x_end)
pyplot.ylim(y_start, y_end);
# We said that the strength $\sigma$ represents a volume flow rate emanating from the source sheet. If you play with this parameter, you will see that the stagnation point comes closer and closer to the source sheet with decreasing strength.
#
# If we wanted to use multiple source sheets to represents the streamlines around a given body shape, we need to make each source sheet part of the dividing streamline. The question will be *What is the source strength to make this happen?*
#
# The volume flow rate on just the left side of the sheet is $\frac{\sigma}{2}$, flowing in the opposite direction of the freestream velocity $U_\infty$.
# Therefore, the flow-tangency boundary condition required is $\frac{\sigma}{2} = U_\infty$.
#
# Now go back to the code above an replace the `sigma` by the correct value. *Where is the stagnation point? Where is the dividing streamline?*
# ## Learn more
# The `lambda` statement can be especially confusing when you are starting out with Python. Here are a couple of places to dig deeper:
#
# * an interesting blog post that treats the subtleties of lambdas: [Yet Another Lambda Tutorial](http://pythonconquerstheuniverse.wordpress.com/2011/08/29/lambda_tutorial/) at the "Python Conquers the Universe" blog (29 August 2011)
# * the chapter on ["Anonymous functions: lambda"](https://www.inkling.com/read/learning-python-mark-lutz-4th/chapter-19/anonymous-functions-lambda) of the *Learning Python* book by <NAME>.
# ---
# + active=""
# Please ignore the cell below. It just loads our style for the notebook.
# -
from IPython.core.display import HTML
def css_styling(filepath):
styles = open(filepath, 'r').read()
return HTML(styles)
css_styling('../styles/custom.css')
| lessons/08_Lesson08_sourceSheet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# python 3.7
# matplotlib pyecharts
# 目的:分别采用matplotlib 和 pyecharts 绘图
# **思路**:
#
# 导入数据
#
# pandas清洗数据。
#
# 用matplotlib绘图
#
# 用pyecharts绘图
# ---
#
# # 海外数据的导入并简单清洗数据
# ### 导入
import pandas as pd
import numpy as np
# ls
foreign_df = pd.read_json('./data/foreign_df.json')
foreign_df.head()
# ### 清洗数据。
# foreign_df 按新增确诊排列
foreign_df=foreign_df.sort_values('新增确诊',ascending=False)
# +
#foreign_df 删除疫情列 foreign_df = foreign_df.iloc[:,0:len(foreign_df.columns)-1]
# -
# foreign_df 重置 index
foreign_df=foreign_df.reset_index(drop=True) # drop参数是问你是否删除之前的index
# 注意 reset_index() 和 reindex() 的区别
#
# foreign_df.reindex(index = list(np.arange(0,len(foreign_df.index),1)))
foreign_df.head()
# ---
# # china数据的导入与简单清理
# ### 导入
import pandas as pd
import numpy as np
china_df = pd.read_json('./data/china_df.json')
china_df.head()
# ### 清洗
# 重置index标签
china_df=china_df.reset_index(drop=True)
# 重命名colums标签
china_df = china_df.rename({0:'地区',1: '现有确诊', 2:'累计确诊',3: '治愈',4:'死亡',5:'其他'}, axis='columns')
china_df.head()
# ### china_df 拆分为 china_province_df 和 china_city_df
# 思路:如果最后一列是详情,则说明当前index是省数据。
# +
# 测试 结果是dataframe
print(china_df.iloc[0:1,5:6] ,'\n')
# 测试 结果是dataframe的元素的判断并赋值判断结果
print(china_df.iloc[0:1,5:6]=='详情' ,'\n')
# 测试 提取标量值
print(china_df.iloc[0,5],'\n')
# 测试 提取标量值
print(china_df['其他'][0] , '\n')
# -
index_list = list(china_df.index) # index标签列表
last_col_loc = len(china_df.columns)-1 #最后一列的位置
print(type(index_list[0]))
print(type(last_col_loc))
#省数据的idnex的列表推导式
# iloc[1,2]按位置提取标量值,注意这里不是iloc[ :,: ] 。
province_index = [ x for x in index_list if china_df.iloc[x,last_col_loc] ]
# 拿到china_province_df
china_province_df = china_df.loc[province_index,:] # 注意loc按标签提取 iloc按位置提取
# 拿到china_city_df
china_city_df = china_df.drop(province_index)
# 重置index
china_province_df = china_province_df.reset_index(drop=True)
china_city_df = china_city_df.reset_index(drop=True)
# +
# 删除最后一列 iloc[:,:]
# china_province_df = china_province_df.iloc[:,0:len(china_province_df.columns)-1]
# china_city_df = china_city_df.iloc[:,0:len(china_city_df.columns)-1]
# -
china_province_df.head()
china_city_df.head()
# ---
# # 可视化1:海外新增TOP10(matplotlib)
# ##### 数据清理
import pandas as pd
import numpy as np
foreign_df.head()
df = foreign_df
# 确保新增确诊是int类型(可以排序),我将强制转为int,然后生成一个新的df
dict_ = {'地区':df['地区'],'新增确诊':[ x for x in df['新增确诊'].values]}
df = pd.DataFrame(dict_)
# 按新增确诊排序
df = df.sort_values('新增确诊',ascending=False)
df.head()
# top
top=df.head(10)
# series转为list
x = list(top['地区'])
y = list(top['新增确诊'])
print(x)
print(y)
# ##### 可视化 matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
# 设置中文字体
plt.rcParams['font.family']=['sans-serif']
plt.rcParams['font.sans-serif']=['SimHei']
# +
plt.bar(x,y)
# 在每个柱上加数值标签
for a,b in zip(x,y):
# 参数 a,b+50 是文本标签的空间位置 ha是文本标签的内容居中
plt.text(a,b+50,'%d'%b,ha='center')
plt.title('海外新增确诊top')
# -
# ---
# # 可视化2:中国现有确诊top (matplotlib)
# #### 数据清理
import pandas as pd
import numpy as np
df = china_province_df
df.head()
# 为了确保新增确诊的元素是数值,这样的才能排序。
# 我们将新增确诊的元素强制为int类型,然后重新构建一个df
dict_ = {'地区':df['地区'],'现有确诊':pd.Series([ int(x) for x in df['现有确诊'].values ])}
df = pd.DataFrame(dict_)
# - 上述 dict_ ,pandas还有更简单的方法:
# - dict_ = {'地区':df['地区'],'现有确诊':pd.Series(list(df['现有确诊'].astype('int')))}
# 按照新增确诊排序 降序
df = df.sort_values('现有确诊',ascending=False)
# 重置index
df = df.reset_index(drop=True)
df.head()
# top
top = df.head(10)
x = list(top['地区'])
y = list(top['现有确诊'])
# test 注意看,a打印是不同于bc的,类型也不同。
a = top['地区'].values
b = list(top['地区'].values)
c = list(top['地区'])
print(a)
print(b)
print(c)
print(type(a))
print(type(b))
print(type(c))
# #### 可视化
import matplotlib.pyplot as plt
# %matplotlib inline
# 设置中文字体
plt.rcParams['font.family']=['sans-serif']
plt.rcParams['font.sans-serif']=['SimHei']
plt.bar(x,y)
# 设置标签
for a,b in zip(x,y):
plt.text(a,b+5,'%d'%b,ha='center')
plt.title('中国现有确诊(含港澳台)')
# # 可视化3:全球现有top 疫情地图(pyecharts)
# #### 导入与清洗数据
# 提取出海外国家的现有确诊数目
#
# 将中国(含港澳台)现有数据汇总
#
# 结合中国和海外的数据生成一个新的df。
import numpy as np
import pandas as pd
df_c = china_province_df
df_c.head(3)
# 对新增确诊的seriers并values求和 ,过程中将新增确诊的values强制转为int
sum_ = sum([ int(x) for x in df_c['现有确诊'] ])
dict_ = {'地区':pd.Series(['中国']),'现有确诊':pd.Series([sum_])}
df_c = pd.DataFrame(dict_)
# - 上述的求和,pandas有更简单的方式
# - sum( list(df_c['新增确诊'].astype(int)) )
df_c.head()
df_f = foreign_df
df_f.head(3)
# 将累计确诊、治愈、死亡的数据类型强制转为int,并生成list
cumulative_confirm = list(df_f['累计确诊'])
cure = list(df_f['治愈'])
death = list(df_f['死亡'])
# 现有确诊 = 累计确诊-治愈-死亡 注意: 列表没有减法,集合有减法但是集合会排序,所以都不行
existing_confirm= []
for i in range(len(cumulative_confirm)):
t = cumulative_confirm[i]-cure[i]-death[i]
existing_confirm.append(t)
# 生成df
dict_ = {'地区':df_f['地区'],'现有确诊':pd.Series(existing_confirm)}
df_f = pd.DataFrame(dict_)
# 按现有确诊排序,并重置index标签
df_f = df_f.sort_values('现有确诊',ascending=False)
df_f = df_f.reset_index(drop=True)
df_f.head()
# 合并df_c 和 df_f (追加的方式)
df = df_c.append(df_f)
# 对合并后的df,按新增确诊进行排序(降序)
df = df.sort_values('现有确诊',ascending=False)
# 重置index标签
df = df.reset_index(drop=True)
df.head()
# #### 可视化
# series 可以直接转为 list
countrys = list(df['地区'])
existing_confirm=list(df['现有确诊'])
# 注意:pycharts 的全球地图不可以用中文名。所以我们要进行国家名称的转化
# 我读入一个国家名称的中英映射文件,用于中英转化。
import json
with open('name_map.json','r',encoding='utf-8') as json_file:
# json.dump()和json.load()用于读写。 dumps()和loads()用于dict和json的转换
name_map = json.load(json_file)
# +
# 键值互换
# name_map = dict(zip(name_map.values(),name_map.keys()))
# -
# test
print( list(name_map.keys())[0] )
print( list(name_map.values())[0] )
# 将国家名称改为英文 注意用 dict.get()方法,因为不能保证一定有完整的映射关系。
countrys = [name_map.get(name) for name in countrys]
list_=[list(z) for z in zip(countrys,existing_confirm)]
# 剔除国家名称为空的数据
list_=[x for x in list_ if x[0]!=None]
# #### pyecharts
# https://pyecharts.readthedocs.io/projects/pyecharts-en/zh/latest/zh-cn/customize_map/
#
# pip install pyecharts
#
# pip install echarts-china-provinces-pypkg
#
# pip install echarts-countries-pypkg
from pyecharts.charts import Map
from pyecharts import options as opts
# +
# 数据划分等级
pieces = [
{'min':0,'max':99},
{'min':100,'max':999},
{'min':1000,'max':9999},
{'min':10000,'max':99999},
{'min':100000},
]
map_world = (
Map()
.add('世界疫情现有确诊',list_,'world')
.set_series_opts(label_opts=opts.LabelOpts(is_show=False))
.set_global_opts(visualmap_opts=opts.VisualMapOpts(is_show=True,is_piecewise=True,pieces=pieces))
)
# -
map_world.render_notebook()
| crawl and V/visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [<NAME>](http://sebastianraschka.com)
# <br>
# [Link to](https://github.com/rasbt/pattern_classification) the GitHub repository [pattern_classification](https://github.com/rasbt/pattern_classification)
# <br><br>
# %load_ext watermark
# %watermark -a '<NAME>' -d -u -v -p scikit-learn,numpy,matplotlib
# <hr>
# I would be happy to hear your comments and suggestions.
# Please feel free to drop me a note via
# [twitter](https://twitter.com/rasbt), [email](mailto:<EMAIL>), or [google+](https://plus.google.com/+SebastianRaschka).
# <hr>
# <br>
# <br>
# # About Feature Scaling: Z-Score Standardization and Min-Max Scaling
# <br>
# <br>
# ## Sections
# - [About standardization](#About-standardization)
# - [About Min-Max scaling / "normalization"](#About-Min-Max-scaling-normalization)
# - [Standardization or Min-Max scaling?](#Standardization-or-Min-Max-scaling?)
# - [Standardizing and normalizing - how it can be done using scikit-learn](#Standardizing-and-normalizing---how-it-can-be-done-using-scikit-learn)
# - [Bottom-up approaches](#Bottom-up-approaches)
# - [The effect of standardization on PCA in a pattern classification task](#The-effect-of-standardization-on-PCA-in-a-pattern-classification-task)
# <br>
# <br>
# ## About standardization
# [[back to top](#Sections)]
# The result of **standardization** (or **Z-score normalization**) is that the features will be rescaled so that they'll have the properties of a standard normal distribution with
#
# $\mu = 0$ and $\sigma = 1$
#
# where $\mu$ is the mean (average) and $\sigma$ is the standard deviation from the mean; standard scores (also called ***z*** scores) of the samples are calculated as follows:
#
# \begin{equation} z = \frac{x - \mu}{\sigma}\end{equation}
#
# Standardizing the features so that they are centered around 0 with a standard deviation of 1 is not only important if we are comparing measurements that have different units, but it is also a general requirement for many machine learning algorithms. Intuitively, we can think of gradient descent as a prominent example
# (an optimization algorithm often used in logistic regression, SVMs, perceptrons, neural networks etc.); with features being on different scales, certain weights may update faster than others since the feature values $x_j$ play a role in the weight updates
#
# $$\Delta w_j = - \eta \frac{\partial J}{\partial w_j} = \eta \sum_i (t^{(i)} - o^{(i)})x^{(i)}_{j},$$
#
# so that
#
# $$w_j := w_j + \Delta w_j,$$
# where $\eta$ is the learning rate, $t$ the target class label, and $o$ the actual output.
# Other intuitive examples include K-Nearest Neighbor algorithms and clustering algorithms that use, for example, Euclidean distance measures -- in fact, tree-based classifier are probably the only classifiers where feature scaling doesn't make a difference.
#
#
#
# To quote from the [`scikit-learn`](http://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html) documentation:
#
# *"Standardization of a dataset is a common requirement for many machine learning estimators: they might behave badly if the individual feature do not more or less look like standard normally distributed data (e.g. Gaussian with 0 mean and unit variance)."*
# <br>
# <br>
# <a id='About-Min-Max-scaling-normalization'></a>
# ## About Min-Max scaling
# [[back to top](#Sections)]
# An alternative approach to Z-score normalization (or standardization) is the so-called **Min-Max scaling** (often also simply called "normalization" - a common cause for ambiguities).
# In this approach, the data is scaled to a fixed range - usually 0 to 1.
# The cost of having this bounded range - in contrast to standardization - is that we will end up with smaller standard deviations, which can suppress the effect of outliers.
#
# A Min-Max scaling is typically done via the following equation:
#
# \begin{equation} X_{norm} = \frac{X - X_{min}}{X_{max}-X_{min}} \end{equation}
# <br>
# <br>
# ## Z-score standardization or Min-Max scaling?
# [[back to top](#Sections)]
# *"Standardization or Min-Max scaling?"* - There is no obvious answer to this question: it really depends on the application.
#
# For example, in clustering analyses, standardization may be especially crucial in order to compare similarities between features based on certain distance measures. Another prominent example is the Principal Component Analysis, where we usually prefer standardization over Min-Max scaling, since we are interested in the components that maximize the variance (depending on the question and if the PCA computes the components via the correlation matrix instead of the covariance matrix; [but more about PCA in my previous article](http://sebastianraschka.com/Articles/2014_pca_step_by_step.html)).
#
# However, this doesn't mean that Min-Max scaling is not useful at all! A popular application is image processing, where pixel intensities have to be normalized to fit within a certain range (i.e., 0 to 255 for the RGB color range). Also, typical neural network algorithm require data that on a 0-1 scale.
# <br>
# <br>
# ## Standardizing and normalizing - how it can be done using scikit-learn
# [[back to top](#Sections)]
# Of course, we could make use of NumPy's vectorization capabilities to calculate the z-scores for standardization and to normalize the data using the equations that were mentioned in the previous sections. However, there is an even more convenient approach using the preprocessing module from one of Python's open-source machine learning library [scikit-learn](http://scikit-learn.org ).
# <br>
# <br>
# %watermark -a '<NAME>' -d -p scikit-learn,numpy,pandas,matplotlib -v
# <font size="1.5em">[More information](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/ipython_magic/watermark.ipynb) about the `watermark` magic command extension.</font>
# <br>
# <br>
# For the following examples and discussion, we will have a look at the free "Wine" Dataset that is deposited on the UCI machine learning repository
# (http://archive.ics.uci.edu/ml/datasets/Wine).
#
# <br>
#
# <font size="1">
# **Reference:**
# <NAME> al, PARVUS - An Extendible Package for Data
# Exploration, Classification and Correlation. Institute of Pharmaceutical
# and Food Analysis and Technologies, Via Brigata Salerno,
# 16147 Genoa, Italy.
#
# <NAME>. & <NAME>. (2013). UCI Machine Learning Repository [http://archive.ics.uci.edu/ml]. Irvine, CA: University of California, School of Information and Computer Science.
#
# </font>
# The Wine dataset consists of 3 different classes where each row correspond to a particular wine sample.
#
# The class labels (1, 2, 3) are listed in the first column, and the columns 2-14 correspond to 13 different attributes (features):
#
# 1) Alcohol
# 2) Malic acid
# ...
# #### Loading the wine dataset
# +
import pandas as pd
import numpy as np
df = pd.io.parsers.read_csv(
'https://raw.githubusercontent.com/rasbt/pattern_classification/master/data/wine_data.csv',
header=None,
usecols=[0,1,2]
)
df.columns=['Class label', 'Alcohol', 'Malic acid']
df.head()
# -
# <br>
# <br>
# As we can see in the table above, the features **Alcohol** (percent/volumne) and **Malic acid** (g/l) are measured on different scales, so that ***Feature Scaling*** is necessary important prior to any comparison or combination of these data.
#
#
# <br>
# <br>
# #### Standardization and Min-Max scaling
# +
from sklearn import preprocessing
std_scale = preprocessing.StandardScaler().fit(df[['Alcohol', 'Malic acid']])
df_std = std_scale.transform(df[['Alcohol', 'Malic acid']])
minmax_scale = preprocessing.MinMaxScaler().fit(df[['Alcohol', 'Malic acid']])
df_minmax = minmax_scale.transform(df[['Alcohol', 'Malic acid']])
# -
print('Mean after standardization:\nAlcohol={:.2f}, Malic acid={:.2f}'
.format(df_std[:,0].mean(), df_std[:,1].mean()))
print('\nStandard deviation after standardization:\nAlcohol={:.2f}, Malic acid={:.2f}'
.format(df_std[:,0].std(), df_std[:,1].std()))
print('Min-value after min-max scaling:\nAlcohol={:.2f}, Malic acid={:.2f}'
.format(df_minmax[:,0].min(), df_minmax[:,1].min()))
print('\nMax-value after min-max scaling:\nAlcohol={:.2f}, Malic acid={:.2f}'
.format(df_minmax[:,0].max(), df_minmax[:,1].max()))
# <br>
# <br>
# #### Plotting
# %matplotlib inline
# +
from matplotlib import pyplot as plt
def plot():
plt.figure(figsize=(8,6))
plt.scatter(df['Alcohol'], df['Malic acid'],
color='green', label='input scale', alpha=0.5)
plt.scatter(df_std[:,0], df_std[:,1], color='red',
label='Standardized [$N (\mu=0, \; \sigma=1)$]', alpha=0.3)
plt.scatter(df_minmax[:,0], df_minmax[:,1],
color='blue', label='min-max scaled [min=0, max=1]', alpha=0.3)
plt.title('Alcohol and Malic Acid content of the wine dataset')
plt.xlabel('Alcohol')
plt.ylabel('Malic Acid')
plt.legend(loc='upper left')
plt.grid()
plt.tight_layout()
plot()
plt.show()
# -
# <br>
# <br>
# The plot above includes the wine datapoints on all three different scales: the input scale where the alcohol content was measured in volume-percent (green), the standardized features (red), and the normalized features (blue).
# In the following plot, we will zoom in into the three different axis-scales.
# <br>
# <br>
# +
fig, ax = plt.subplots(3, figsize=(6,14))
for a,d,l in zip(range(len(ax)),
(df[['Alcohol', 'Malic acid']].values, df_std, df_minmax),
('Input scale',
'Standardized [$N (\mu=0, \; \sigma=1)$]',
'min-max scaled [min=0, max=1]')
):
for i,c in zip(range(1,4), ('red', 'blue', 'green')):
ax[a].scatter(d[df['Class label'].values == i, 0],
d[df['Class label'].values == i, 1],
alpha=0.5,
color=c,
label='Class %s' %i
)
ax[a].set_title(l)
ax[a].set_xlabel('Alcohol')
ax[a].set_ylabel('Malic Acid')
ax[a].legend(loc='upper left')
ax[a].grid()
plt.tight_layout()
plt.show()
# -
# <br>
# <br>
# ## Bottom-up approaches
# [[back to top](#Sections)]
# Of course, we can also code the equations for standardization and 0-1 Min-Max scaling "manually". However, the scikit-learn methods are still useful if you are working with test and training data sets and want to scale them equally.
#
# E.g.,
# <pre>
# std_scale = preprocessing.StandardScaler().fit(X_train)
# X_train = std_scale.transform(X_train)
# X_test = std_scale.transform(X_test)
# </pre>
#
# Below, we will perform the calculations using "pure" Python code, and an more convenient NumPy solution, which is especially useful if we attempt to transform a whole matrix.
# <br>
# <br>
# Just to recall the equations that we are using:
#
# Standardization: \begin{equation} z = \frac{x - \mu}{\sigma} \end{equation}
#
# with mean:
#
# \begin{equation}\mu = \frac{1}{N} \sum_{i=1}^N (x_i)\end{equation}
#
# and standard deviation:
#
# \begin{equation}\sigma = \sqrt{\frac{1}{N} \sum_{i=1}^N (x_i - \mu)^2}\end{equation}
#
#
# Min-Max scaling: \begin{equation} X_{norm} = \frac{X - X_{min}}{X_{max}-X_{min}} \end{equation}
#
#
#
# ### Pure Python
# +
# Standardization
x = [1,4,5,6,6,2,3]
mean = sum(x)/len(x)
std_dev = (1/len(x) * sum([ (x_i - mean)**2 for x_i in x]))**0.5
z_scores = [(x_i - mean)/std_dev for x_i in x]
# Min-Max scaling
minmax = [(x_i - min(x)) / (max(x) - min(x)) for x_i in x]
# -
# <br>
# <br>
# ### NumPy
# +
import numpy as np
# Standardization
x_np = np.asarray(x)
z_scores_np = (x_np - x_np.mean()) / x_np.std()
# Min-Max scaling
np_minmax = (x_np - x_np.min()) / (x_np.max() - x_np.min())
# -
# <br>
# <br>
# ### Visualization
# Just to make sure that our code works correctly, let us plot the results via matplotlib.
# %matplotlib inline
# +
from matplotlib import pyplot as plt
fig, ((ax1, ax2), (ax3, ax4)) = plt.subplots(nrows=2, ncols=2, figsize=(10,5))
y_pos = [0 for i in range(len(x))]
ax1.scatter(z_scores, y_pos, color='g')
ax1.set_title('Python standardization', color='g')
ax2.scatter(minmax, y_pos, color='g')
ax2.set_title('Python Min-Max scaling', color='g')
ax3.scatter(z_scores_np, y_pos, color='b')
ax3.set_title('Python NumPy standardization', color='b')
ax4.scatter(np_minmax, y_pos, color='b')
ax4.set_title('Python NumPy Min-Max scaling', color='b')
plt.tight_layout()
for ax in (ax1, ax2, ax3, ax4):
ax.get_yaxis().set_visible(False)
ax.grid()
plt.show()
# -
# <br>
# <br>
# ## The effect of standardization on PCA in a pattern classification task
# [[back to top](#Sections)]
# Earlier, I mentioned the Principal Component Analysis (PCA) as an example where standardization is crucial, since it is "analyzing" the variances of the different features.
# Now, let us see how the standardization affects PCA and a following supervised classification on the whole wine dataset.
#
#
# In the following section, we will go through the following steps:
#
# - Reading in the dataset
# - Dividing the dataset into a separate training and test dataset
# - Standardization of the features
# - Principal Component Analysis (PCA) to reduce the dimensionality
# - Training a naive Bayes classifier
# - Evaluating the classification accuracy with and without standardization
# <br>
# <br>
# ### Reading in the dataset
# [[back to top](#Sections)]
# +
import pandas as pd
df = pd.io.parsers.read_csv(
'https://raw.githubusercontent.com/rasbt/pattern_classification/master/data/wine_data.csv',
header=None,
)
# -
# <br>
# <br>
# ### Dividing the dataset into a separate training and test dataset
# [[back to top](#Sections)]
# In this step, we will randomly divide the wine dataset into a training dataset and a test dataset where the training dataset will contain 70% of the samples and the test dataset will contain 30%, respectively.
# +
from sklearn.cross_validation import train_test_split
X_wine = df.values[:,1:]
y_wine = df.values[:,0]
X_train, X_test, y_train, y_test = train_test_split(X_wine, y_wine,
test_size=0.30, random_state=12345)
# -
# <br>
# <br>
# ### Feature Scaling - Standardization
# [[back to top](#Sections)]
# +
from sklearn import preprocessing
std_scale = preprocessing.StandardScaler().fit(X_train)
X_train_std = std_scale.transform(X_train)
X_test_std = std_scale.transform(X_test)
# -
# <br>
# <br>
# ### Dimensionality reduction via Principal Component Analysis (PCA)
# [[back to top](#Sections)]
# Now, we perform a PCA on the standardized and the non-standardized datasets to transform the dataset onto a 2-dimensional feature subspace.
# In a real application, a procedure like cross-validation would be done in order to find out what choice of features would yield a optimal balance between "preserving information" and "overfitting" for different classifiers. However, we will omit this step since we don't want to train a perfect classifier here, but merely compare the effects of standardization.
# +
from sklearn.decomposition import PCA
# on non-standardized data
pca = PCA(n_components=2).fit(X_train)
X_train = pca.transform(X_train)
X_test = pca.transform(X_test)
# om standardized data
pca_std = PCA(n_components=2).fit(X_train_std)
X_train_std = pca_std.transform(X_train_std)
X_test_std = pca_std.transform(X_test_std)
# -
# Let us quickly visualize how our new feature subspace looks like (note that class labels are not considered in a PCA - in contrast to a Linear Discriminant Analysis - but I will add them in the plot for clarity).
# %matplotlib inline
# +
from matplotlib import pyplot as plt
fig, (ax1, ax2) = plt.subplots(ncols=2, figsize=(10,4))
for l,c,m in zip(range(1,4), ('blue', 'red', 'green'), ('^', 's', 'o')):
ax1.scatter(X_train[y_train==l, 0], X_train[y_train==l, 1],
color=c,
label='class %s' %l,
alpha=0.5,
marker=m
)
for l,c,m in zip(range(1,4), ('blue', 'red', 'green'), ('^', 's', 'o')):
ax2.scatter(X_train_std[y_train==l, 0], X_train_std[y_train==l, 1],
color=c,
label='class %s' %l,
alpha=0.5,
marker=m
)
ax1.set_title('Transformed NON-standardized training dataset after PCA')
ax2.set_title('Transformed standardized training dataset after PCA')
for ax in (ax1, ax2):
ax.set_xlabel('1st principal component')
ax.set_ylabel('2nd principal component')
ax.legend(loc='upper right')
ax.grid()
plt.tight_layout()
plt.show()
# -
# <br>
# <br>
# ### Training a naive Bayes classifier
# [[back to top](#Sections)]
# We will use a naive Bayes classifier for the classification task. If you are not familiar with it, the term "naive" comes from the assumption that all features are "independent".
# All in all, it is a simple but robust classifier based on Bayes' rule
#
# Bayes' Rule:
#
#
# \begin{equation} P(\omega_j|x) = \frac{p(x|\omega_j) * P(\omega_j)}{p(x)} \end{equation}
#
# where
#
# - ω: class label
# - *P(ω|x)*: the posterior probability
# - *p(x|ω)*: prior probability (or likelihood)
#
# and the **decsion rule:**
#
# Decide $ \omega_1 $ if $ P(\omega_1|x) > P(\omega_2|x) $ else decide $ \omega_2 $.
# <br>
#
#
# \begin{equation}
# \Rightarrow \frac{p(x|\omega_1) * P(\omega_1)}{p(x)} > \frac{p(x|\omega_2) * P(\omega_2)}{p(x)}
# \end{equation}
#
#
# I don't want to get into more detail about Bayes' rule in this article, but if you are interested in a more detailed collection of examples, please have a look at the [Statistical Patter Classification](https://github.com/rasbt/pattern_classification#statistical-pattern-recognition-examples) in my pattern classification repository.
#
#
# +
from sklearn.naive_bayes import GaussianNB
# on non-standardized data
gnb = GaussianNB()
fit = gnb.fit(X_train, y_train)
# on standardized data
gnb_std = GaussianNB()
fit_std = gnb_std.fit(X_train_std, y_train)
# -
# <br>
# <br>
# ### Evaluating the classification accuracy with and without standardization
# [[back to top](#Sections)]
# +
from sklearn import metrics
pred_train = gnb.predict(X_train)
print('\nPrediction accuracy for the training dataset')
print('{:.2%}'.format(metrics.accuracy_score(y_train, pred_train)))
pred_test = gnb.predict(X_test)
print('\nPrediction accuracy for the test dataset')
print('{:.2%}\n'.format(metrics.accuracy_score(y_test, pred_test)))
# +
pred_train_std = gnb_std.predict(X_train_std)
print('\nPrediction accuracy for the training dataset')
print('{:.2%}'.format(metrics.accuracy_score(y_train, pred_train_std)))
pred_test_std = gnb_std.predict(X_test_std)
print('\nPrediction accuracy for the test dataset')
print('{:.2%}\n'.format(metrics.accuracy_score(y_test, pred_test_std)))
# -
# As we can see, the standardization prior to the PCA definitely led to an decrease in the empirical error rate on classifying samples from test dataset.
| pattern-classification/preprocessing/about_standardization_normalization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Episode 20
#
# In a desperate attempt to regain some composure following a cheese migration incident, we note the deprecation of `pgen` in the CPython implementation, and then continue to assume nothing is wrong. After all, using LL(1) pushdown automata is a perfectly fine method of parsing source code, and will continue to live in the heart and minds of countless pre-3.9 Pythonistas...
import parser
# %load_ext metamagics
def ignore_this_after_3_9(cell):
return parser.st2tuple(parser.suite(cell))
# %%mycell ignore_this_after_3_9
1+2-3
A=42
# +
import io
from lib2to3.pgen2 import pgen, driver
from typing import Callable, Dict
class Bindify:
class Binder:
def __init__(self, parent: 'Bindify', binding_name: str):
self.parent = parent
self.binding_name = binding_name
def __call__(self, cell, shell):
result = self.parent.magic_fun(cell, shell)
shell.user_global_ns[self.binding_name] = result
return result
def __init__(self, magic_fun: Callable):
self.binders: Dict[str, Binder] = {}
self.magic_fun = magic_fun
def __getitem__(self, name: str):
if name in self.binders:
return self.binders[name]
binder = Bindify.Binder(self, name)
self.binders[name] = binder
return binder
def __call__(self, cell, shell):
return self.magic_fun(cell, shell)
def _pgenify(cell, _):
generator = pgen.ParserGenerator('<cell-magic-string>', io.StringIO(cell))
return generator.make_grammar()
pgenify = Bindify(_pgenify)
# +
# %%mycell pgenify['calc_grammar']
statements: (statement NEWLINE)* ENDMARKER
statement: assign | expr
assign: NAME '=' expr
expr: atom [op expr]
atom: NAME | NUMBER
op: '+' | '-'
# +
# %%mycell pgenify['calc_grammar']
statements: (statement NEWLINE)* ENDMARKER
statement: expr ['=' expr]
expr: atom [op expr]
atom: NAME | NUMBER
op: '+' | '-'
# +
import nbimport
import Episode16
Tree = Episode16.Tree
from lib2to3 import pytree
calc_driver = driver.Driver(calc_grammar, pytree.convert)
calc_driver.parse_string('1 + 2 - 3\n')
# +
def pytree_to_tree(node):
if isinstance(node, pytree.Leaf):
return Tree(node, [])
else:
return Tree(node.type, [pytree_to_tree(child) for child in node.children])
def _calcify(cell, _):
return pytree_to_tree(calc_driver.parse_string(cell))
calcify = Bindify(_calcify)
# -
# %%mycell calcify['tree']
1 + 2 - 3
A = 42
Episode16.tree_to_tuple(tree)
calc_grammar.report()
# ## A Word of Warning
#
# There is a bug in the generated parser where the `convert` argument is optional, but when left unused, the raw representation will be a tuple and not an object.
#
# Therefore, this doesn't work:
bad_driver = driver.Driver(calc_grammar)
bad_driver.parse_string('1 + 2 - 3\n')
# +
class Tree2:
def __init__(self, contents, children):
self.contents = contents
self.children = children
def to_tuple(self):
return self.contents, [child.to_tuple() for child in self.children]
def ohop_convert(grammar, raw_node):
type, value, context, children = raw_node
if children or type in grammar.number2symbol:
return Tree2(type, children)
else:
return Tree2((type, value), [])
better_driver = driver.Driver(calc_grammar, ohop_convert)
better_driver.parse_string('1 + 2 - 3\n').to_tuple()
# -
def ohop_convert2(grammar, raw_node):
type, value, context, children = raw_node
if children or type in grammar.number2symbol:
return Tree2(type if type not in grammar.number2symbol else grammar.number2symbol[type], children)
else:
return Tree2((type, value), [])
yet_another_driver = driver.Driver(calc_grammar, ohop_convert2)
yet_another_driver.parse_string('1 + 2 - 3\nA = 42\n').to_tuple()
| ohop/Episode20.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import geopandas as gpd
import pandas as pd
import numpy as np
import glob, os
import salem
import matplotlib.pyplot as plt
import seaborn as sns
from oggm import utils
import cartopy.crs as ccrs
sns.set_context('paper')
sns.set_style('whitegrid')
# Get the RGI summary
mdf = pd.read_hdf(utils.file_downloader('https://cluster.klima.uni-bremen.de/~oggm/rgi/rgi62_stats.h5'))
mdf['O2Region'] = ['{:02d}-{:02d}'.format(int(d1), int(d2)) for d1, d2 in zip(mdf.O1Region, mdf.O2Region)]
df = salem.read_shapefile('/home/mowglie/disk/Data/GIS/SHAPES/RGI/RGI_V6/00_rgi60_regions/00_rgi60_O1Regions.shp')
dfsr = salem.read_shapefile('/home/mowglie/disk/Data/GIS/SHAPES/RGI/RGI_V6/00_rgi60_regions/00_rgi60_O2Regions.shp')
fac = 5
mdf['CenLonC'] = np.round(mdf.CenLon * fac).astype(int)
mdf['CenLatC'] = np.round(mdf.CenLat * fac).astype(int)
mdf['CenC'] = ['{}_{}'.format(lon, lat) for lon, lat in zip(mdf['CenLonC'], mdf['CenLatC'])]
selids = []
for i, g in mdf.groupby('CenC'):
dis = (g.CenLon - g.CenLonC/fac)**2 + (g.CenLat - g.CenLatC/fac)**2
selids.append(dis.idxmin())
smdf = mdf.loc[selids]
len(smdf)
# +
sns.set_context('talk')
f = plt.figure(figsize=(12, 7))
ax = plt.axes(projection=ccrs.PlateCarree())
ax.set_extent([-180, 180, -90, 90], crs=ccrs.PlateCarree())
ax.stock_img()
# ax.add_feature(cartopy.feature.COASTLINE
# ax.coastlines('50m')
ax.scatter(smdf.CenLon.values, smdf.CenLat.values, color='C3', marker='.', s=3)
# ax.scatter(dfr.lon.values, dfr.lat.values, edgecolor='k', facecolor='tab:cyan', s=24)
df.plot(ax=ax, transform=ccrs.PlateCarree(), edgecolor='k', facecolor='w', alpha=0.3);
dfsr.plot(ax=ax, transform=ccrs.PlateCarree(), edgecolor='k', linestyle=':', facecolor='none');
df.plot(ax=ax, transform=ccrs.PlateCarree(), edgecolor='k', facecolor='none', linewidth=2);
did1 = False
did10 = False
for i, r in df.iterrows():
tx = r.min_x + 2
ty = r.min_y + 2
fs = 12
t = r.RGI_CODE
if t == 1:
tx = r.min_x + 35
if did1:
continue
did1 = True
if t == 5:
tx = r.min_x + 19
if t == 6:
ty = r.min_y - 7.
if t == 8:
tx = r.min_x + 26
if t == 11:
tx = r.min_x - 8.5
if t == 10:
ty = r.min_y + 17
if not did10:
did10 = True
continue
if t == 13:
ty = r.min_y + 11
tx = r.min_x + 18
if t == 15:
tx = r.min_x + 21
if t == 17:
ty = r.min_y + 25
if t == 18:
ty = r.min_y + 17
# ax.text(tx, ty, t, transform=ccrs.PlateCarree(),
# ha='left', va='bottom', fontsize=fs,
# bbox=dict(facecolor='w', edgecolor='k', alpha=0.7))
# letkm = dict(color='black', ha='left', va='top', fontsize=16,
# bbox=dict(facecolor='white', edgecolor='black'))
# ax.text(-178, 88.2, 'a', **letkm)
plt.tight_layout();
plt.savefig('rgi_subreg_map.pdf', bbox_inches='tight');
# -
reg_names, subreg_names = utils.parse_rgi_meta(version='6')
area_per_reg = mdf[['Area', 'O2Region']].groupby('O2Region').sum()
area_per_reg['N Glaciers'] = mdf.groupby('O2Region').count().Name
area_per_reg['RegName'] = ['{}: {} '.format(i, subreg_names.loc[i].values[0]) for i in area_per_reg.index]
sns.set_context('paper')
f, ax = plt.subplots(1, 1, figsize=(8, 18))
sns.barplot(x="N Glaciers", y='RegName', data=area_per_reg, color='Grey');
plt.ylabel('');
plt.savefig('rgi_subreg_nglaciers.pdf', bbox_inches='tight');
sns.set_context('paper')
f, ax = plt.subplots(1, 1, figsize=(8, 18))
sns.barplot(x="Area", y='RegName', data=area_per_reg, color='Grey');
plt.ylabel('');
plt.savefig('rgi_subreg_area.pdf', bbox_inches='tight');
| rgi_subregs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JSDesai/NNDL_lab_2022/blob/main/bImplementation_of_Customer_churn_prediction_using_dataset_for_Bankipynb.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/", "height": 270} id="Gg4qyeycRWuG" outputId="eb1b02b4-c05f-4a3e-f4e2-3a6fad10179b"
import pandas as pd
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import train_test_split
import tensorflow as tf
from tensorflow import keras
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
df=pd.read_csv('/content/ChurnBank.csv')
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="T_QD3kzERcaE" outputId="6b3b0b31-c577-4001-f6f6-60612eeed691"
df.shape
# + colab={"base_uri": "https://localhost:8080/"} id="xyxFIFbYRffE" outputId="1b45ef1a-28c2-4127-dedf-18c06ca505ca"
df.dtypes
# + id="76rYse-sRiJP"
df = df.drop(['RowNumber', 'CustomerId', 'Surname'], axis=1)
# + [markdown] id="mtgs0pIxRkXl"
# Visualization
#
# + colab={"base_uri": "https://localhost:8080/", "height": 402} id="TJ1GsATwRnRQ" outputId="d44000e9-2314-49b7-b7b2-9116d434454e"
existed_yes = df[df['Exited']== 1].Tenure
existed_no = df[df['Exited']== 0].Tenure
plt.xlabel('Tenure')
plt.ylabel('no of customer')
plt.title('Churn prediction')
plt.hist([existed_no,existed_yes], color = ['red', 'blue'], label =(['Existed = no','Existed = yes']))
plt.legend()
# + colab={"base_uri": "https://localhost:8080/", "height": 402} id="bAVM2lTORqHI" outputId="6d7111d2-1388-4a9e-a5c1-e6479c70aa94"
salary_no_existed = df[df.Exited== 0].EstimatedSalary
salary_yes_existed = df[df.Exited== 1].EstimatedSalary
plt.xlabel('Estimad_salary')
plt.ylabel('no of customer')
plt.title('Churn prediction')
plt.hist([salary_no_existed,salary_yes_existed], color = ['green', 'blue'], label =(['Existed = no','Existed = yes']))
plt.legend()
# + colab={"base_uri": "https://localhost:8080/", "height": 270} id="s1bXYejzRr-M" outputId="63f3a006-28be-4513-e9fc-5c9a801e0671"
df.Gender = df.Gender.map({'Female':1, 'Male':0})
df = pd.get_dummies(data=df, columns=['Geography'])
scaler = MinMaxScaler()
df[['CreditScore', 'Age', 'Tenure', 'Balance', 'NumOfProducts', 'EstimatedSalary']] = scaler.fit_transform(df[['CreditScore', 'Age', 'Tenure', 'Balance', 'NumOfProducts', 'EstimatedSalary']])
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="lHeU2Cl2Rt6r" outputId="5e95a38e-6a90-42dc-841f-0b44ec00d4ef"
df.Exited.value_counts()
# + [markdown] id="HsL0NwwgRwK1"
# Training Model
# + colab={"base_uri": "https://localhost:8080/"} id="jcteoE-wRx75" outputId="ce23dd77-2332-4778-c7e3-a369ae3d931c"
x = df.drop('Exited', axis = 1)
y = df.Exited
x_train, x_test, y_train, y_test = train_test_split(x, y)
x_train.shape
# + colab={"base_uri": "https://localhost:8080/"} id="87roMtihR1Pw" outputId="49a7022c-3491-4d97-d3e3-b010c06a4244"
x_test.shape
# + colab={"base_uri": "https://localhost:8080/"} id="yQN_wafrR2MF" outputId="a7a540af-941c-4b85-e40b-46595efe42a3"
import tensorflow as tf
from tensorflow import keras
model = keras.Sequential([
keras.layers.Dense(12, input_shape = (12,), activation = 'relu'),
keras.layers.Dense(12, activation = 'relu'),
keras.layers.Dense(1, activation = 'sigmoid')])
model.compile(optimizer = 'adam', loss = 'binary_crossentropy', metrics = 'accuracy')
model.fit(x_train, y_train, epochs = 100)
# + colab={"base_uri": "https://localhost:8080/"} id="uG56VS6tR68M" outputId="965f3ff7-9e06-4a39-809e-9a467af1ac0c"
model.evaluate(x_test, y_test)
# + colab={"base_uri": "https://localhost:8080/"} id="5YvxnC8QR79p" outputId="47ea140e-23b1-4791-def6-f1d4f39b9824"
yp = model.predict(x_test)
yp[:5]
# + colab={"base_uri": "https://localhost:8080/"} id="CTKwxIG8R9bw" outputId="b2bd746f-386f-430d-b92b-b923a612a5b5"
y_pred = []
for element in yp:
if element > 0.5:
y_pred.append(1)
else:
y_pred.append(0)
from sklearn.metrics import confusion_matrix , classification_report
print(classification_report(y_test,y_pred))
# + colab={"base_uri": "https://localhost:8080/", "height": 459} id="lhhPZ-AFR_Y1" outputId="ab24d55d-ff6c-460e-d523-2e151bd5060c"
import seaborn as sn
from matplotlib import pyplot as plt
cm = tf.math.confusion_matrix(labels=y_test,predictions=y_pred)
plt.figure(figsize = (10,7))
sn.heatmap(cm, annot=True, fmt='d')
plt.xlabel('Predicted')
plt.ylabel('Truth')
# + [markdown] id="KHBC6v9jSEua"
# Accuracy
# + colab={"base_uri": "https://localhost:8080/"} id="7070yb91SCpO" outputId="7f3060a7-ffd2-4cbd-e876-afca277223f8"
round((1922+229)/(1922+229+275+74),2)
# + [markdown] id="2BgG8jFGSHBJ"
# Precision for class 0
# + colab={"base_uri": "https://localhost:8080/"} id="E0hpVbVkSDrh" outputId="cc07d0a4-a261-4557-c2ce-96935ef230d0"
round(1922/(1922+275),2)
# + [markdown] id="0bX49mjCSLvc"
# Precision for 1 class
# + colab={"base_uri": "https://localhost:8080/"} id="0Sat54FYSNxD" outputId="b5bc0d17-269d-4be4-ee6f-6ac81f50b314"
round(229/(229+74),2)
# + [markdown] id="9MCCaWP_SP-r"
# Recall for 0 class
# + colab={"base_uri": "https://localhost:8080/"} id="pgypvsWcSSPB" outputId="d81cf4f2-1b70-4780-e465-aae7c157752c"
round(1922/(1922+74),2)
# + colab={"base_uri": "https://localhost:8080/"} id="PItCJ1JtST11" outputId="d8f12808-4f89-4790-b575-8eb6292f40a6"
round(229/(229+275),2)
| 1b Implementation_of_Customer_churn_prediction_using_dataset_for_Bankipynb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Sheet 7: Reinforcement Learning I - <NAME> & <NAME>
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from IPython.display import set_matplotlib_formats
set_matplotlib_formats('png', 'pdf')
plt.rcParams.update({'font.size': 14})
plt.rcParams["figure.figsize"] = (14,7)
# ## Exercise 1: The 10-armed bandit
# ### 1. Generate an array $\bar{p}$ of 10 random lever reward probabilities $p_i ∈ [0, 0.9]$ and keep it fixed for the entire exercise sheet.
p = np.linspace(0, 0.9, 10)
print(p)
# ### 2. Write a function $generate\_reward$ that will behave as an N-armed bandit (given lever numbers corresponding to indices into $\bar{p}$, it returns rewards of 1 with a probability $p_i$ , otherwise zeros).
def generate_reward(p):
lever_num = len(p) # number of levers
reward = np.zeros(lever_num) # empty reward array
for i in range(lever_num):
reward[i] = np.random.choice([0,1], p=[1-p[i], p[i]])
return reward
# ### Confirm that your bandit works: Activate each lever 10000 times and compare the mean reward for each lever with its reward probability – they should be nearly identical.
total_reward = 0
trial_num = 10000
for i in range(trial_num):
total_reward += generate_reward(p)
mean_reward = total_reward/trial_num
print(mean_reward, ', mean reward for each lever')
print('%.4f, average difference between mean reward and reward probability' %(np.sum(p - mean_reward)/len(p)))
# Thus, the bandit works as the mean reward and reward probability are nearly identical for a sample size of 10,000.
# ### 3. Devise some method of storing and updating the $Q_i$ value associated with the action of pressing lever $i$ (take $η = 0.01$).
# ### $Q$ should be initialized with zeros, since the agent is pessimistic and initially associates zero value with each action.
def Q_update(Q, p, eta):
lever_num = len(p)
R = generate_reward(p)
Q_next = eta * (R - Q)
return Q_next
# ### 4. We will first implement the $\epsilon\_greedy$ policy: The agent will with probability $1-\epsilon$ press the lever which has the maximum associated $Q$-value (if there are several with that value, press them all), otherwise press a random lever. The agent starts by pressing all levers (since they all have the same $Q$-value initially), it updates the $Q$-values and then applies the described strategy iteratively. Write a function (or class method) $\epsilon\_greedy$ that implements this strategy. The function/method should accept as input the current $Q$ and should return the lever index (or indices) that are to be pressed, the $Q$-value of the chosen lever, as well as the largest $Q$-value.
def epsilon_greedy(Q_current, p, epsilon, eta):
initial_roll = np.random.choice([False, True], p=[epsilon, 1-epsilon]) # choose max Q or not
Q_max_idx = np.argmax(Q_current) # max Q value index
Q_max = Q_current[Q_max_idx] # max Q value
Q_num = len(Q_current) # size of Q
if np.all(Q_current == 0): # initial state
press = np.ones(Q_num) # press all levers
Q_chosen = 0
elif initial_roll: # press max Q lever
press = np.zeros(Q_num)
press[Q_max_idx] = 1
Q_chosen = Q_current[Q_max_idx]
else: # choose random lever
press = np.zeros(Q_num)
random_choice = np.random.randint(0, Q_num)
press[random_choice] = 1
Q_chosen = Q_current[random_choice]
return press, Q_chosen, Q_max
# ### 5. First we’ll test the pure-greedy algorithm by setting $epsilon = 0$ and running 1000 iterations of the algorithm:
# ### 1) Choose the next lever(s) ($eps\_greedy$).
# ### 2) Activate the chosen levers ($generate\_reward$).
# ### 3) Update the $Q$-values according to the received rewards.
def greedy_algorithm(trials, epsilon):
Q = np.zeros(len(p)) # pessimistic agent, initially Q=0 for all i
Q_expected = np.zeros(trials) # Q for current lever over trials (including initial state)
for i in range(trials): # loop over trials
press, Q_chosen, Q_max = epsilon_greedy(Q_current=Q, p=p, epsilon=epsilon, eta=0.01) # choose levers
reward = generate_reward(p=press) # press levers
Q += Q_update(Q=Q, p=reward, eta=0.01) # update Q
Q_expected[i] = Q_chosen # store chosen lever Q
return Q_expected, Q_max
# ### Plot the estimated expected reward (the $Q$-value of the current lever) against the trial number (each lever press constitutes one trial). Repeat the test 20 times and draw the resulting curves into a single plot.
# ### Does the agent consistently identify the levers with the largest reward probabilities?
iteration_num = 1000
[plt.plot(greedy_algorithm(trials=iteration_num, epsilon=0)[0], color='k', label=(i+1)) for i in range(20)]
plt.title('$Q$-value of Current Lever - Test # = 20')
plt.ylabel('Q')
plt.xlabel('trial #')
plt.grid()
plt.show()
# Running the test repeatedly shows that this agent does consistently identify the levers with the largest reward probabilities, as the curves overlay each other.
# ### 6. Run 100 repetitions (“lives”) of the $\epsilon-greedy$ strategy (with four different $\epsilon$-values 0.0, 0.01, 0.1 and 0.5) at 1000 iterations each and store the expected reward in each trial.
epsilons = [0.0, 0.01, 0.1, 0.5]
live_num = 100
Qs = np.zeros((len(epsilons), iteration_num)) # expected rewards
Q_maxes = np.zeros((len(epsilons), iteration_num)) # largest Q values
for i in range(live_num): # loop over lives
for j in range(len(epsilons)): # loop over epsilons
run_greedy = greedy_algorithm(trials=iteration_num, epsilon=epsilons[j])
Qs[j] += run_greedy[0] # take sum of the lives
Q_maxes[j] += run_greedy[1] # take sum of Q maxes
# ### Plot the average of the expected reward, and the average largest $Q$-value across lives against the trial number. Include a line in the plot that runs parallel to the x-axis and represents the maximum expected reward (the maximum reward probability).
# ### Why do the expected reward and largest $Q$-value curves look the way they do for the four cases?
# +
max_reward = 1 - np.transpose(epsilons) # the maximum reward probability
fig, axs = plt.subplots(2,1, sharex=True, sharey=True, figsize=(14,10))
color = iter(plt.cm.viridis(np.linspace(0, 1, np.shape(Qs)[0])))
for i in range(np.shape(Qs)[0]):
c = next(color)
axs[0].plot(Qs[i]/live_num, c=c, label='$\epsilon= %s$' %epsilons[i])
axs[0].axhline(max_reward[i], c=c, linestyle='--')
axs[0].set_title('Average of the Expected Reward')
axs[0].set_ylabel('average reward')
axs[0].legend()
axs[0].grid()
color = iter(plt.cm.viridis(np.linspace(0, 1, np.shape(Qs)[0])))
for i in range(np.shape(Q_maxes)[0]):
c = next(color)
axs[1].plot(Q_maxes[i]/live_num, c=c, label='$\epsilon= %s$' %epsilons[i])
axs[1].set_title('Average of the Largest $Q$-value')
axs[1].set_ylabel('average $Q_{max}$')
axs[1].set_xlabel('trial #')
axs[1].legend()
axs[1].grid()
plt.tight_layout()
plt.show()
# -
# Simply, as $\epsilon$ is increased, the randomness of chossing the lever is increased as well. For no $\epsilon$, the bandit only chooses the max reward (greedy). Otherwise, a bit of randomness is introduced with a magnitude corresponding to the size of epsilon. Also, the amount of reward max would be capped by this as the bandit will select lower rewards with a probability of $\epsilon-1$.
# Do think our plots of the largest Q-value are wrong, it's weird that it is just a straight line over all trials.
# ### 7. Finally, we program a SoftMax policy:
# ### The agent presses the levers with probabilities that depend on the Q-values:
# ## <font color='k'> $ P_i = exp(\beta Q_i) / \sum_j exp(\beta Q_j) $
# ### Write a function (or class method) $softmax$ that implements this strategy. The function/method should accept as input the current $Q$ and output the lever index (it will always be a single lever index!) that is to be pressed, as well as the maximum $Q$-value. Test this function/method by running 100 lives with 5000 iterations each and store the expected reward and maximum $Q$-value in each trial. Draw the same curves as in task 6 for different values of $β$ = 1, 5, 15, 50.
def softmax(Q, beta):
prob = np.exp(beta * Q) / np.sum(np.exp(beta * Q)) # probability for each lever
press_idx = np.random.choice(np.arange(0, len(Q), 1), p=prob) # lever index
Q_max = np.max(Q) # maximum Q value
return press_idx, Q_max
def softmax_algorithm(trials, beta):
Q = np.zeros(len(p)) # pessimistic agent, initially Q=0 for all i
Q_expected = np.zeros(trials) # Q for current lever over trials (including initial state)
for i in range(trials): # loop over trials
press_idx, Q_max = softmax(Q, beta) # choose lever
reward = generate_reward(p=p) # press levers
Q += Q_update(Q=Q, p=reward, eta=0.01) # update Q
Q_expected[i] = Q[press_idx] # expected Q
return Q_expected, Q_max
betas = [1, 5, 15, 50]
live_num = 100
iteration_num = 5000
Qs = np.zeros((len(betas), iteration_num)) # expected rewards
Q_maxes = np.zeros((len(betas), iteration_num)) # largest Q values
for i in range(live_num): # loop over lives
for j in range(len(betas)): # loop over betas
run_softmax = softmax_algorithm(trials=iteration_num, beta=betas[j])
Qs[j] += run_softmax[0] # take sum of the lives
Q_maxes[j] += run_softmax[1] # take sum of Q maxes
# +
fig, axs = plt.subplots(2,1, sharex=True, sharey=True, figsize=(14,10))
color = iter(plt.cm.viridis(np.linspace(0, 1, np.shape(Qs)[0])))
for i in range(np.shape(Qs)[0]):
c = next(color)
axs[0].plot(Qs[i]/live_num, c=c, label=r'$\beta= %i$' %betas[i])
axs[0].set_title('Average of the Expected Reward')
axs[0].set_ylabel('average reward')
axs[0].legend()
axs[0].grid()
color = iter(plt.cm.viridis(np.linspace(0, 1, np.shape(Qs)[0])))
for i in range(np.shape(Q_maxes)[0]):
c = next(color)
axs[1].plot(Q_maxes[i]/live_num, c=c, label=r'$\beta= %i$' %betas[i])
axs[1].set_title('Average of the Largest $Q$-value')
axs[1].set_ylabel('average $Q_{max}$')
axs[1].set_xlabel('trial #')
axs[1].legend()
axs[1].grid()
plt.tight_layout()
plt.show()
# -
# ### Do the results of $SoftMax$ show any improvements compared to the $\epsilon-greedy$ policies? How does the parameter $β$ relate to the exploration/exploitation-balance? And once again explain why for each $β$ the two curves (expected reward & largest $Q$-value) look the way they do.
# For a larger $\beta$, the probability of choosing a larger reward is higher. Thus, the average of the expected reward is higher as well.
# I would think that $\beta$ is an exploitation parameter, where the higher it is the more exploitation, and the lower it is the more exploration.
# ### 8. Modify the 10-armed bandit so that operating the last lever always gives a reward of -5000 (the agent is punished and has to return 5000 pieces of chocolate). Test both the $epsilon-greedy$ and $SoftMax$ policies on this modified bandit (use $\epsilon = 0.1, β = 15$) by running 500 lives with 1000 iterations each and plot the same curve as in task 6. Do you observe a difference? Give an intuition for the result! (After this task ditch the punishment and return to the original bandit.)
def generate_reward(p): # the modified reward function
lever_num = len(p) # number of levers
reward = np.zeros(lever_num) # empty reward array
for i in range(lever_num-1):
reward[i] = np.random.choice([0,1], p=[1-p[i], p[i]])
reward[-1] = -5000 # punishment lever
return reward
def softmax_algorithm(trials, beta):
Q = np.zeros(len(p)) # pessimistic agent, initially Q=0 for all i
Q_expected = np.zeros(trials) # Q for current lever over trials (including initial state)
for i in range(trials): # loop over trials
press_idx, Q_max = softmax(Q, beta) # choose lever
reward = generate_reward(p=p) # press levers
Q += Q_update(Q=Q, p=reward, eta=0.01) # update Q
Q_expected[i] = Q[press_idx] # expected Q
return Q_expected, Q_max
live_num = 500
iteration_num = 1000
Qs = np.zeros((2, iteration_num)) # expected rewards, both policies
Q_maxes = np.zeros((2, iteration_num)) # largest Q values, both policies
for i in range(live_num): # loop over lives
run_greedy = greedy_algorithm(trials=iteration_num, epsilon=0.1)
run_softmax = softmax_algorithm(trials=iteration_num, beta=15)
Qs[0] += run_greedy[0] # take sum of the lives
Q_maxes[0] += run_greedy[1] # take sum of Q maxes
Qs[1] += run_softmax[0] # take sum of the lives
Q_maxes[1] += run_softmax[1] # take sum of Q maxes
# +
fig, axs = plt.subplots(2,1, sharex=True, sharey=False, figsize=(14,10))
color = iter(plt.cm.viridis(np.linspace(0, 1, 2)))
axs[0].plot(Qs[0]/live_num, c=next(color), label=r'$\epsilon=%s$' %0.1)
axs[0].plot(Qs[1]/live_num, c=next(color), label=r'$\beta= %s$' %15)
axs[0].set_title('Average of the Expected Reward')
axs[0].set_ylabel('average reward')
axs[0].legend()
axs[0].grid()
color = iter(plt.cm.viridis(np.linspace(0, 1, 2)))
axs[1].plot(Q_maxes[0]/live_num, c=next(color), label=r'$\epsilon=%s$' %0.1)
axs[1].plot(Q_maxes[1]/live_num, c=next(color), label=r'$\beta= %s$' %15)
axs[1].set_title('Average of the Largest $Q$-value')
axs[1].set_ylabel('average $Q_{max}$')
axs[1].set_xlabel('trial #')
axs[1].legend()
axs[1].grid()
plt.tight_layout()
plt.show()
# -
# The epsilon-greedy policy is so much worse than the softmax as the randomness of choosing the punishment is larger than it being able to overcome it, as the punishment magnitude is so large.
# ### 9. Implement a modified SoftMax policy,
# ### where the parameter β increases linearly with the number of iterations $i$ starting at a value of 1:
# ## <font color='k'> $ \beta(i) = 1 + i/b $
# ### Rerun the simulations with 5 different $β$-slopes: $b$ = 0.1, 0.4, 1.6, 6.4, 25.6. Compare the results with the simple $SoftMax$ ($β$ = 5) by running 100 lives with 2000 iterations and plotting the same curves as in task 6.
def modified_softmax_algorithm(trials, b): # modified softmax policy
Q = np.zeros(len(p)) # pessimistic agent, initially Q=0 for all i
Q_expected = np.zeros(trials) # Q for current lever over trials (including initial state)
for i in range(trials): # loop over trials
beta = 1 + (1+i)/b
press_idx, Q_max = softmax(Q, beta) # choose lever
reward = generate_reward(p=p) # press levers
Q += Q_update(Q=Q, p=reward, eta=0.01) # update Q
Q_expected[i] = Q[press_idx] # expected Q
return Q_expected, Q_max
def generate_reward(p): # original reward function
lever_num = len(p) # number of levers
reward = np.zeros(lever_num) # empty reward array
for i in range(lever_num):
reward[i] = np.random.choice([0,1], p=[1-p[i], p[i]])
return reward
b = np.array([0.1, 0.4, 1.6, 6.4, 25.6]) * 100
live_num = 100
iteration_num = 2000
Qs = np.zeros((len(b), iteration_num)) # expected rewards
Q_maxes = np.zeros((len(b), iteration_num)) # largest Q values
for i in range(live_num): # loop over lives
for j in range(len(b)): # loop over betas
run_softmax = modified_softmax_algorithm(trials=iteration_num, b=b[j])
Qs[j] += run_softmax[0] # take sum of the lives
Q_maxes[j] += run_softmax[1] # take sum of Q maxes
# +
simple_Qs = np.zeros((len(b), iteration_num)) # expected rewards
simple_Q_maxes = np.zeros((len(b), iteration_num)) # largest Q values
for i in range(live_num): # loop over lives
run_simple = softmax_algorithm(trials=iteration_num, beta=5)
simple_Qs += run_simple[0]
simple_Q_maxes += run_simple[1]
fig, axs = plt.subplots(2,1, sharex=True, sharey=False, figsize=(14,10))
color = iter(plt.cm.viridis(np.linspace(0, 1, np.shape(Qs)[0])))
for i in range(np.shape(Qs)[0]):
c = next(color)
axs[0].plot(Qs[i]/live_num, c=c, label=r'$b= %s$' %b[i])
axs[0].plot(simple_Qs/live_num, color='k', label=r'simple, $\beta= %i$' %5) # simple softmax
axs[0].set_title('Average of the Expected Reward')
axs[0].set_ylabel('average reward')
axs[0].legend()
axs[0].grid()
color = iter(plt.cm.viridis(np.linspace(0, 1, np.shape(Qs)[0])))
for i in range(np.shape(Q_maxes)[0]):
c = next(color)
axs[1].plot(Q_maxes[i]/live_num, c=c, label=r'$b= %s$' %b[i])
axs[1].plot(simple_Q_maxes/live_num, color='k', label=r'simple, $\beta= %i$' %5) # simple softmax
axs[1].set_title('Average of the Largest $Q$-value')
axs[1].set_ylabel('average $Q_{max}$')
axs[1].set_xlabel('trial #')
axs[1].legend()
axs[1].grid()
plt.tight_layout()
plt.show()
# -
# ### In a different figure, plot the cumulative sum of true rewards (the sum of rewards until the current trial, averaged over lives) against trial number. What do you observe?
# Did not finish past this point.
# ### Furthermore, plot the cumulative reward at the end of the lifetime (2000 iterations) against the $β$-slope. Which of the five slopes generates the maximum overall reward? Explain the idea behind using a varying $β$, in particular regarding the exploration/exploitation-dilemma? Can you think of a similar modification to the $\epsilon$-greedy policy?
| MHBF/.ipynb_checkpoints/MHBF_CC7_Longren-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# + [markdown] id="cbOkgD3W22tk"
# # Exercise 1
# + id="vSepQCEM3wNq"
import numpy as np
import pandas as pd
# + [markdown] id="9DhSr8fJ22tw"
# ### Step 1. Go to https://www.kaggle.com/openfoodfacts/world-food-facts/data
# + [markdown] id="SUbW4_DO22tx"
# ### Step 2. Download the dataset to your computer and unzip it.
# + [markdown] id="MLEnLqrC22ty"
# ### Step 3. Use the tsv file and assign it to a dataframe called food
# + id="YrnTR2XW22tz" outputId="a6522cd5-4323-4750-f12a-27502111ba77" colab={"base_uri": "https://localhost:8080/"}
filepath = './en.openfoodfacts.org.products.tsv'
food = pd.read_csv(filepath, sep='\t')
# + [markdown] id="rLVOkqRL22t0"
# ### Step 4. See the first 5 entries
# + id="JrUzJn7x22t0" outputId="d3a34f61-a474-4981-ed1d-2afe0c566214" colab={"base_uri": "https://localhost:8080/", "height": 406}
food.head()
# + [markdown] id="tjJD1y6022t1"
# ### Step 5. What is the number of observations in the dataset?
# + id="lPvENR6X22t1" outputId="bba20a8b-b3a3-4293-9abf-e6a83048f2d4" colab={"base_uri": "https://localhost:8080/"}
food.shape[0]
# + [markdown] id="5Zr56IX622t2"
# ### Step 6. What is the number of columns in the dataset?
# + id="u8SW2bk522t2" outputId="dd04045e-2d85-4ff5-d0ad-c0fcc4715cac" colab={"base_uri": "https://localhost:8080/"}
food.shape[1]
# + [markdown] id="Fa7D8K2m22t3"
# ### Step 7. Print the name of all the columns.
# + id="qHGK31dT22t3" outputId="58a60799-8bac-4d33-eaba-71abfea0bf50" colab={"base_uri": "https://localhost:8080/"}
food.columns
# + [markdown] id="1o4ZiRKA22t3"
# ### Step 8. What is the name of 105th column?
# + id="uw-INiUR22t4" outputId="605c4059-566b-40cf-f83d-82636129ff97" colab={"base_uri": "https://localhost:8080/", "height": 35}
food.columns[104]
# + [markdown] id="UZUVV_qM22t4"
# ### Step 9. What is the type of the observations of the 105th column?
# + id="qjs1tALn22t5" outputId="cdcdb1a6-a51f-473f-a13a-a8700069d3e9" colab={"base_uri": "https://localhost:8080/"}
food.dtypes['-glucose_100g']
# + [markdown] id="nV5_PObs22t5"
# ### Step 10. How is the dataset indexed?
# + id="syflEmLl22t5" outputId="707568d5-3144-4ac4-961b-fa069799753c" colab={"base_uri": "https://localhost:8080/"}
food.index
# + [markdown] id="57A1Xe5w22t6"
# ### Step 11. What is the product name of the 19th observation?
# + id="raCyBMlF22t6" outputId="71057601-4476-40a7-fc17-a70f948d8bc8" colab={"base_uri": "https://localhost:8080/", "height": 35}
food.values[18][7]
| 01_Getting_&_Knowing_Your_Data/World Food Facts/Exercises[Solved].ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import sys
sys.path.insert(0, "../..")
# # Overview
#
# This notebook will continue showing how to define ConText modifiers with some more advanced options.
# +
import medspacy
from medspacy.context import ConTextRule, ConTextComponent
from medspacy.visualization import visualize_dep, visualize_ent
from spacy.tokens import Span
# -
nlp = medspacy.load()
# ## 1: **"TERMINATE"** rule
# As said before, the scope of a modifier is originally set to the entire sentence either before after a ConTextModifier, as defined by the ItemData's `direction` attribute. However, the scope can be modified by **termination points**, which is another ConTextModifier with the rule **"TERMINATE"**. For example, in "There is no evidence of pneumonia but there is CHF", the negation modifier should modify "pneumonia" but not "CHF". This can be achieved by defining a ConTextRule to terminate at the word "but".
text = "There is no evidence of pneumonia but there is CHF"
context_rules = [ConTextRule("no evidence of", "NEGATED_EXISTENCE", "FORWARD")]
context = ConTextComponent(nlp, rules=None)
context.add(context_rules)
doc = nlp(text)
doc.ents = (Span(doc, 5, 6, "CONDITION"), Span(doc, 9, 10, "CONDITION"))
doc.ents
context(doc)
# Here, you can see that both **"pneumonia"** and **"CHF"** are modified:
visualize_dep(doc)
modifier = doc._.context_graph.modifiers[0]
modifier
# The scope includes both "pneumonia" and "CHF", so both would be negated
modifier.scope
# Now let's add an additional ConTextRule with **"TERMINATE"**:
context_rules2 = [ConTextRule("but", "CONJ", "TERMINATE")]
context.add(context_rules2)
# doc = nlp(text)
context(doc)
doc._.context_graph.modifiers
modifier = doc._.context_graph.modifiers[0]
visualize_dep(doc)
# The scope now only encompasses "pneumonia"
modifier.scope
# ## 2: Pseudo-modifiers
# Sometimes, substrings of a phrase will incorrectly cause a modifier to be negated. For example:
#
# ---
# "There are no findings to explain the weakness"
#
# ---
#
# **"No"** is often a negation modifier, but **"no findings to explain"** should not **"weakness"**:
text = "There are no findings to explain the weakness"
doc = nlp(text)
doc.ents = (Span(doc, 7, 8, "CONDITION"),)
context = ConTextComponent(nlp, rules="default")
context(doc)
visualize_dep(doc)
# If we add **"no findings to explain"** as a **"PSEUDO"** modifier, it will supercede the substring **"no"** and will not modify the target concept:
pseudo = ConTextRule("no findings to explain", "PSEUDO", direction="PSEUDO")
context.add([pseudo])
context(doc)
visualize_dep(doc)
# ## 3: Pruned modifiers
# If two ConTextItems result in TagObjects where one is the substring of another, the modifiers will be pruned to keep **only** the larger span. We saw this above with **"no"** and **"no findings to explain"**.
#
# As another example, **"no history of"** is a negation modifier, while **"history of"** is a historical modifier. Both match the text "no history of afib", but only "no history of" should ultimately modify "afib".
item_data = [ConTextRule("no history of", "DEFINITE_NEGATED_EXISTENCE", "FORWARD"),
ConTextRule("history", "HISTORICAL", "FORWARD"),
]
text = "no history of"
context = ConTextComponent(nlp, rules=None)
context.add(item_data)
doc = nlp(text)
context(doc)
# Two overlapping modifiers
doc._.context_graph.modifiers
# ## 4: Manually limiting scope
# By default, the scope of a modifier is the **entire sentence** in the direction of the rule up until a termination point (see above). However, sometimes this is too much. In long sentences, this can cause a modifier to extend far beyond its location in the sentence. Some modifiers are really meant to be attached to a single concept, but they are instead distributed to all targets.
#
# To fix this, ConText allows optional attributes in `ItemData` to limit the scope: `max_scope` and `max_targets`. Both attributes are explained below.
# ### max_targets
# Some modifiers should really only attach to a single target. For example, in the sentence below:
#
# **"Pt presents with diabetes, pneumonia?"**
#
# **"?"** indicates uncertainty, but *only* with **"pneumonia"**. **"Diabetes"** should not be affected. We can achieve this by creating a bidirectional rule with a `max_targets` of **1**. This will limit the number of targets to 1 *on each side* of the tag object.
#
# Let's first see what this looks like *without* defining `max_targets`:
text = "Pt presents with diabetes, pneumonia?"
doc = nlp(text)
doc.ents = (doc[3:4], doc[5:6])
doc.ents
item = ConTextRule("?", category="UNCERTAIN",
direction="BACKWARD",
max_scope=None)
context = ConTextComponent(nlp, rules=None)
context.add([item])
context(doc)
# Both are modified
visualize_dep(doc)
# Now, let's start over and set `max_targets` to **1**:
doc = nlp(text)
doc.ents = (doc[3:4], doc[5:6])
rule = ConTextRule("?", category="UNCERTAIN",
direction="BACKWARD",
max_targets=1)
context = ConTextComponent(nlp, rules=None)
context.add([rule])
context(doc)
# Only "pneumonia" is modified
visualize_dep(doc)
# ### max_scope
# One limitation of using `max_targets` is that in a sentence like the example above, each concept has to be extracted as an entity in order for it to reduce the scope - if **"pneumonia"** was not extracted, then **"vs"** would still etend as far back as **"diabetes"**.
#
# We can address this by explicitly setting the scope to be no greater than a certain number of tokens using `max_scope`. For example, lab results may show up in a text document with many individual results:
#
# ---
# Adenovirus DETECTED<br>
# SARS NOT DETECTED<br>
# ...
# Cov HKU1 NOT DETECTED<br>
#
# ---
#
# Texts like this are often difficult to parse and they are often not ConText-friendly because many lines can be extracted as a single sentence. By default, a modifier like **"NOT DETECTED"** could extend far back to a concept such as **"Adenovirus"**, which we see returned positive. We may also not explicitly extract every virus tested in the lab, so `max_targets` won't work.
#
# With text formats like this, we can be fairly certain that **"Not Detected"** will only modify the single concept right before it. We can set `max_scope` to be so **only** a single concept will be modified.
text = """Adenovirus DETECTED Sars NOT DETECTED Pneumonia NOT DETECTED"""
doc = nlp(text)
doc.ents = (doc[0:1], doc[2:3], doc[5:6])
doc.ents
print([sent for sent in doc.sents])
# +
#assert len(list(doc.sents)) == 1
# -
rules = [ConTextRule("DETECTED", category="POSITIVE_EXISTENCE",
direction="BACKWARD",
max_scope=None),
ConTextRule("NOT DETECTED", category="DEFINITE_NEGATED_EXISTENCE",
direction="BACKWARD",
max_scope=None),
]
context = ConTextComponent(nlp, rules=None)
context.add(rules)
context(doc)
visualize_dep(doc)
# Let's now set `max_scope` to 1 and we'll find that only **"pneumonia"** and **"Sars"** are modified by **"NOT DETECTED"**:
doc = nlp(text)
doc.ents = (doc[0:1], doc[2:3], doc[5:6])
doc.ents
rules = [ConTextRule("DETECTED", category="POSITIVE_EXISTENCE",
direction="BACKWARD",
max_scope=1),
ConTextRule("NOT DETECTED", category="DEFINITE_NEGATED_EXISTENCE",
direction="BACKWARD",
max_scope=1),
]
context = ConTextComponent(nlp, rules=None)
context.add(rules)
context(doc)
visualize_dep(doc)
# ### Using `context_window`
# The default scope for context modifier is the sentence containing an entity. This means if the pipeline doesn't include a sentence splitting component, ConText will throw an error:
nlp_no_sents = medspacy.load(enable=[])
doc = nlp_no_sents("There is no evidence of pneumonia. Scheduled visit in two weeks.")
context = ConTextComponent(nlp)
try:
context(doc)
except ValueError as e:
print(e)
# However, sentence splitting is an expensive operation. To avoid that processing step, you can set context to only use the `max_scope` argument shown above. To do this, pass in the argument `use_context_window=True` and a value for `max_scope`:
nlp_no_sents = medspacy.load(enable=[])
context = ConTextComponent(nlp, use_context_window=True, max_scope=3)
doc = nlp_no_sents("There is no evidence of pneumonia. Scheduled visit in two weeks.")
context(doc)
for modifier in doc._.context_graph.modifiers:
print(modifier, modifier.scope)
# While this can allow faster processing, it also risks setting a less accurate modifying scope without knowing sentence boundries.
# ## 5: Filtering target types
# You may want modifiers to only modify targets with certain semantic classes. You can specify which types to be modified/not be modified through the `allowed_types` and `excluded_types` arguments.
#
# For example, in the sentence:
#
# ---
# "She is not prescribed any beta blockers for her hypertension."
#
# ---
#
# **"Beta blockers"** is negated by the phrase **not prescribed"**, but **"hypertension"** should not be negated. By default, a modifier will modify all concepts in its scope, regardless of semantic type:
from spacy.tokens import Span
# Let's write a function to create this manual example
def create_medication_example():
doc = nlp("She is not prescribed any beta blockers for her hypertension.")
# Manually define entities
medication_ent = Span(doc, 5, 7, "MEDICATION")
condition_ent = Span(doc, 9, 10, "CONDITION")
doc.ents = (medication_ent, condition_ent)
return doc
doc = create_medication_example()
doc
# Define our item data without any type restrictions
rules = [ConTextRule("not prescribed", "NEGATED_EXISTENCE", "FORWARD")]
context = ConTextComponent(nlp, rules="other", rule_list=rules)
context(doc)
# Visualize the modifiers
visualize_dep(doc)
# To change this, we can make sure that **"not prescribed"** only modifies **MEDICATION** entities by setting `allowed_types` to **"MEDICATION"**;
rules = [ConTextRule("not prescribed", "NEGATED_EXISTENCE", "FORWARD", allowed_types={"MEDICATION"})]
context = ConTextComponent(nlp, rules="other", rule_list=rules)
doc = create_medication_example()
context(doc)
# Now, only **"beta blockers"** will be negated:
visualize_dep(doc)
# The same can be achieved by setting `excluded_types` to `{"CONDITION"}`.
rules = [ConTextRule("not prescribed", "NEGATED_EXISTENCE", "FORWARD", excluded_types={"CONDITION"})]
# ## 7: Callbacks
# We can also define callback functions which can allow for flexible control of either which phrases are matched in the text or which concepts are modified. The two callback arguments are `on_match` and `on_modifies`.
#
# ### `on_match`
# This functionality is taken directly from spaCy's [rule-based matching](https://spacy.io/usage/rule-based-matching). When a match is found in the text, the `on_match` argument will run a callback function which can perform additional processing on the match, such as removing it if a certain condition is met.
#
# For example, let's say that we want the phrase **"Positive"** to be used as a positive modifier, but this is sometimes ambiguously used in the context of mental health (ie., **"positive thinking"**).
#
# Let's write a word-sense disambiguation function which will remove a match if mental health phrase is in the sentence. See the [spaCy documentation](https://spacy.io/usage/rule-based-matching) for more details:
def wsd_positive(matcher, doc, i, matches):
(_, start, end) = matches[i]
span = doc[start:end]
# Check if words related to mental health are in the sentence
sent = span.sent
for mh_phrase in ["resilience", "mental health", "therapy", "therapist", "feedback"]:
if mh_phrase in sent.text.lower():
print("Removing", span)
matches.pop(i)
return
# If not, keep the match
print("Keeping", span)
texts = [
"Therapist encouraging him to be positive during COVID-19 pandemic.",
"Positive for COVID-19.",
]
docs = list(nlp.pipe(texts))
docs[0].ents = (Span(docs[0], 7, 8, "CONDITION"),)
docs[1].ents = (Span(docs[1], 2, 3, "CONDITION"),)
context = ConTextComponent(nlp, rules=None)
rule = ConTextRule("positive", "POSITIVE_EXISTENCE", on_match=wsd_positive)
context.add([rule])
for doc in docs:
context(doc)
visualize_dep(docs[0])
visualize_dep(docs[1])
# ### `on_modifies`
# A modifier will usually modify a target concept so long as it is within the scope. `on_modifies` runs a one-time check between the modifier and a potential target concept to decide whether or not it will modify that concept.
#
# The function passed in for this argument should take these 3 arguments:
# - `target`: The entity Span
# - `modifier`: The modifying Span
# - `span_between`: The Span in between the target and modifier
#
# And it must return either `True`, which will cause the modifier to apply to the target, or `False`.
#
# For example, in the example below, "No evidence" may incorrectly modify both "Pneumonia" and "COVID". The phrase "post" might indicate that COVID is not negated, so this can prevent our modifier from applying to it.
text = "No evidence of pneumonia post COVID."
doc = nlp(text)
doc.ents = (Span(doc, 3, 4, "CONDITION"), Span(doc, 5, 6, "CONDITION"), )
context = ConTextComponent(nlp, rules=None)
def post_in_span_between(target, modifier, span_between):
print("Evaluating whether {0} will modify {1}".format(modifier, target))
if "post" in span_between.text.lower():
print("Will not modify")
print()
return False
print("Will modify")
print()
return True
rule = ConTextRule("no evidence of", "NEGATED_EXISTENCE", "FORWARD", on_modifies=post_in_span_between)
context.add([rule])
context(doc)
visualize_dep(doc)
# # Setting additional Span attributes
# As seen in an earlier notebook, ConText registers two new attributes for target Spans: `is_experienced` and `is_current`. These values are set to default values of True and changed if a target is modified by certain modifiers. This logic is set in the variable `DEFAULT_ATTRS`. This is a dictionary which maps modifier category names to the attribute name/value pair which should be set if a target is modified by that modifier type.
from medspacy.context import DEFAULT_ATTRS
DEFAULT_ATTRS
# ## Defining custom attributes
# Rather than using the logic shown above, you can set your own attributes by creating a dictionary with the same structure as DEFAULT_ATTRS and passing that in as the `add_attrs` parameter. If setting your own extensions, you must first call `Span.set_extension` on each of the extensions.
#
# If more complex logic is required, custom attributes can also be set manually outside of the ConTextComponent, for example as a post-processing step.
#
# Below, we'll create our own attribute mapping and have them override the default ConText attributes. We'll defined `is_experienced` and `is_family_history`. Because both a negated concept and a family history concept are not actually experienced by a patient, we'll specify both to set `is_experienced` to False. We'll also set the family history modifier to add a new attribute called `is_family_history`.
from spacy.tokens import Span
# Define modifiers and Span attributes
custom_attrs = {
'NEGATED_EXISTENCE': {'is_experienced': False},
'FAMILY_HISTORY': {'is_family_history': True,
'is_experienced': False},
}
# Register extensions - is_experienced should be True by default, `is_family_history` False
Span.set_extension("is_experienced", default=True)
Span.set_extension("is_family_history", default=False)
context = ConTextComponent(nlp, rules=None, add_attrs=custom_attrs)
context.context_attributes_mapping
# +
rules = [ConTextRule("no evidence of", "NEGATED_EXISTENCE", "FORWARD"),
ConTextRule("family history", "FAMILY_HISTORY", "FORWARD"),
]
context.add(rules)
# +
doc = nlp("There is no evidence of pneumonia. Family history of diabetes.")
doc.ents = doc[5:6], doc[-2:-1]
doc.ents
# -
context(doc)
# The new attributes are now available in `ent._`:
for ent in doc.ents:
print(ent)
print("is_experienced: ", ent._.is_experienced)
print("is_family_history: ", ent._.is_family_history)
print()
| notebooks/context/3-Advanced-Modifiers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/tanaymukherjee/Shapley-Value/blob/main/Attribution_Modeling_using_Shapley_value.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="pDlyYy64HUP8"
# ## Conversion ratio example
# Let’s walk through an example. Say that your company converted 100 opportunities at the end of a fiscal quarter. During that period, the marketing department advertised to the associated accounts using three channels:
#
# N = {Facebook, Google, LinkedIn}
#
# All 100 accounts were touched by one or more of the channels throughout their buyer journeys. In other words, the channels worked together by forming coalitions to increase the likelihood of opportunity conversion.
#
# The table below lists all possible channel coalitions and their conversion ratios:
# + colab={"base_uri": "https://localhost:8080/", "height": 264} id="T-Bbnku-HMQC" outputId="29f2ea3f-fc6a-4c24-fb20-a8bf4278cbe5"
import numpy as np
import pandas as pd
from itertools import combinations
def subsets(S):
'''Returns all possible subsets of the given set'''
s = []
for i in range(1, len(S)+1):
s.extend(map(list, combinations(S, i)))
return list(map('+'.join, s))
N = sorted({'Facebook', 'Google', 'LinkedIn'})
coalitions = subsets(N)
coalitions_lbl = ['S{}'.format(i) for i in range(1, len(coalitions)+1)]
# The coalition 'Facebook+Google' (S4) resulted in 10 conversions from 100 opportunities,
# so has a conversion ratio of 10/100 = 0.1 (10%)
IR = np.array([0.18, 0.04, 0.08, 0.1, 0.26, 0.07, 0.27])
pd.options.display.float_format = '{:,.2f}'.format
pd.DataFrame({
'Coalition': coalitions,
'Ratio': IR
}, dtype=np.float64, index=coalitions_lbl)
# + colab={"base_uri": "https://localhost:8080/", "height": 264} id="-mFKmEUxHeV7" outputId="f3ccf684-3a9b-4abd-a02f-f92ae97018fa"
# B is binary squared matrix that represents coalition membership.
# For example, coalition Facebook+LinkedIn (S5) includes members:
# - Facebook (S1)
# - LinkedIn (S3)
# - Facebook+LinkedIn (S5)
# Resulting in the coefficients: [1,0,1,0,1,0,0]
d = 2**len(N)-1
B = np.matrix(np.zeros((d, d)))
for i in range(0, d):
A = coalitions[i]
S = subsets(A.split('+'))
coef = [1 if c in S else 0 for c in coalitions]
B[i] = coef
pd.options.display.float_format = '{:,.0f}'.format
pd.DataFrame(data=B, index=coalitions, columns=coalitions)
# + [markdown] id="gEDQcIjNHihb"
# The worth of each coalition is determined by the characteristic function. In this example, the worth is represented as the sum of conversion ratios of each channel in a coalition:
#
# - Coalition S5 = Facebook+LinkedIn
# - v(S5) = Facebook (S1) + LinkedIn (S3) + Facebook+LinkedIn (S5)
# - v(S5) = 0.18 + 0.08 + 0.26
# - v(S5) = 0.52
#
# The coalition containing all players is known as the grand coalition v(N). The grand coalition's worth should be equal to the total payoff.
# + colab={"base_uri": "https://localhost:8080/", "height": 264} id="qxqtZ57RHg1S" outputId="f7690075-4569-49ae-aef1-9ef4d31f0551"
# The product of the matrices coalition membership and coalition ratios
# is the coalition worth - the result of the characteristic function 'v(S)'
vS = np.dot(B, IR)
vS = np.squeeze(np.asarray(vS))
vSx = ['v({})'.format(lbl) for lbl in coalitions_lbl]
pd.options.display.float_format = '{:,.2f}'.format
pd.DataFrame({
'Coalition': coalitions,
'Worth': vS
}, index=vSx)
# + [markdown] id="1iZQvZvCHvZz"
# ### Calculating the Shapley values
# Now that we know the worth of each coalition, the Shapley values can be calculated by taking the average of each channels marginal contribution to the game, accounting for all possible orderings. Specifically, the Shapley value gives us a way to distribute the worth of the grand coalition (total payoff) between the three channels.
#
# Facebook's Shapley value:
#
#
# | Order v(N) | Marginal contribution |
# | --------------------------- | --------------------- |
# | 1. Facebook+Google+LinkedIn | v(S1) = 0.18 |
# | 2. Facebook+LinkedIn+Google | v(S1) = 0.18 |
# | 3. Google+Facebook+LinkedIn | v(S4) - v(S2) = 0.28 |
# | 4. Google+LinkedIn+Facebook | v(S7) - v(S6) = 0.81 |
# | 5. LinkedIn+Facebook+Google | v(S5) - v(S3) = 0.44 |
# | 6. LinkedIn+Google+Facebook | v(S7) - v(S6) = 0.81 |
# | **Average contribution** | 0.45 |
#
#
# - In orders 1 and 2, Facebook is first to arrive so it receives its full contribution
# - In order 3, Facebook arrives after Google so its marginal contribution is the coalition containing both channels v(S4)
# minus the coalition without Facebook v(S2)
# - In orders 4 and 6, Facebook is last to arrive so its marginal contribution is the coalition containing all channels v(S7)
# minus the coalition without Facebook v(S6)
# - In order 5, Facebook arrives after LinkedIn so its marginal contribution is the coalition containing both channels v(S5)
# minus the coalition without Facebook v(S3)
#
# The Shapley values for all channels:
#
# | Order | Facebook | Google | LinkedIn |
# | --------------------------- | --------------------- | --------------------- | --------------------- |
# | 1. Facebook+Google+LinkedIn | v(S1) = 0.18 | v(S4) - v(S1) = 0.14 | v(S7) - v(S4) = 0.68 |
# | 2. Facebook+LinkedIn+Google | v(S1) = 0.18 | v(S7) - v(S5) = 0.48 | v(S5) - v(S1) = 0.34 |
# | 3. Google+Facebook+LinkedIn | v(S4) - v(S2) = 0.28 | v(S2) = 0.04 | v(S7) - v(S4) = 0.68 |
# | 4. Google+LinkedIn+Facebook | v(S7) - v(S6) = 0.81 | v(S2) = 0.04 | v(S6) - v(S2) = 0.15 |
# | 5. LinkedIn+Facebook+Google | v(S5) - v(S3) = 0.44 | v(S7) - v(S5) = 0.48 | v(S3) = 0.08 |
# | 6. LinkedIn+Google+Facebook | v(S7) - v(S6) = 0.81 | v(S6) - v(S3) = 0.11 | v(S3) = 0.08 |
# | **Average contribution** | 0.45 | 0.215 | 0.335 |
# + id="skT8PJoKIf0i"
from collections import defaultdict
from math import factorial
# Calculate the Shapley values - the average value of each channel's marginal contribution
# to the grand coalition, taking into account all possible orderings.
shapley = defaultdict(int)
n = len(N)
for i in N:
for A in coalitions:
S = A.split('+')
if i not in S:
k = len(S) # Cardinality of set |S|
Si = S
Si.append(i)
Si = '+'.join(sorted(Si))
# Weight = |S|!(n-|S|-1)!/n!
weight = (factorial(k) * factorial(n-k-1)) / factorial(n)
# Marginal contribution = v(S U {i})-v(S)
contrib = vS[coalitions.index(Si)] - vS[coalitions.index(A)]
shapley[i] += weight * contrib
shapley[i] += vS[coalitions.index(i)]/n
# + colab={"base_uri": "https://localhost:8080/", "height": 141} id="nYlyb3tVIkii" outputId="18ba3722-12ec-4e39-d186-1e942e686ff2"
pd.options.display.float_format = '{:,.3f}'.format
pd.DataFrame({
'Shapley value': list(shapley.values())
}, index=list(shapley.keys()))
| Attribution_Modeling_using_Shapley_value.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Use AIC to decide whether or not to include $\Delta C_{p}$ in model
# %matplotlib inline
import pytc
# ### Our results depend on our confidence in our measured heats.
#
# Origin does not return an uncertainty, so we have to estimate the uncertainty ourselves. This can be done by setting a constant uncertainty each measured for the experiment. The value of `HEAT_UNCERTAINTY` below lets us select between being able to resolve $\Delta C_{p}$ (`HEAT_UNCERTAINTY = 0.1`) or not (`HEAT_UNCERTAINTY = 1.0`).
HEAT_UNCERTAINTY = 0.1 #cal/mol; Favors dCp
#HEAT_UNCERTAINTY = 1.0 #cal/mol; Favors no dCp
# #### Fit to the VantHoff global connector to extract van't Hoff Enthalpy
# Titrations of recombinant human S100A14 with $ZnCl_{2}$ at $5$, $10$, $17$, $25$, $30$ and $35 ^{\circ}C$. Concentrations were: $[protein] = 110\ \mu M$ and $[ZnCl_{2}]=200\ mM$. Buffer was $25\ mM$ Trizma, $100\ mM$ NaCl, $pH\ 7.4$. Buffer was treated with chelex and filtered to $0.22\ \mu m$. Data collected using a GE ITC200.
# +
import numpy as np
# --------------------------------------------------------------------
# Create a global fitting instance
g1 = pytc.GlobalFit()
vh = pytc.global_connectors.VantHoff("vh")
#--------------------------------------------------------------------------------------------------
t5 = pytc.ITCExperiment("temp-dependence/5C.DH",pytc.indiv_models.SingleSite,shot_start=1,uncertainty=HEAT_UNCERTAINTY)
g1.add_experiment(t5)
g1.link_to_global(t5,"dH",vh.dH)
g1.link_to_global(t5,"K",vh.K)
g1.link_to_global(t5,"fx_competent","g_competent")
#--------------------------------------------------------------------------------------------------
t10 = pytc.ITCExperiment("temp-dependence/10C.DH",pytc.indiv_models.SingleSite,shot_start=1,uncertainty=HEAT_UNCERTAINTY)
g1.add_experiment(t10)
g1.link_to_global(t10,"dH",vh.dH)
g1.link_to_global(t10,"K",vh.K)
g1.link_to_global(t10,"fx_competent","g_competent")
#--------------------------------------------------------------------------------------------------
t17 = pytc.ITCExperiment("temp-dependence/17C.DH",pytc.indiv_models.SingleSite,shot_start=1,uncertainty=HEAT_UNCERTAINTY)
g1.add_experiment(t17)
g1.link_to_global(t17,"dH",vh.dH)
g1.link_to_global(t17,"K",vh.K)
g1.link_to_global(t17,"fx_competent","g_competent")
#--------------------------------------------------------------------------------------------------
t25 = pytc.ITCExperiment("temp-dependence/25C.DH",pytc.indiv_models.SingleSite,shot_start=1,uncertainty=HEAT_UNCERTAINTY)
g1.add_experiment(t25)
g1.link_to_global(t25,"dH",vh.dH)
g1.link_to_global(t25,"K",vh.K)
g1.link_to_global(t25,"fx_competent","g_competent")
#--------------------------------------------------------------------------------------------------
t30 = pytc.ITCExperiment("temp-dependence/30C.DH",pytc.indiv_models.SingleSite,shot_start=1,uncertainty=HEAT_UNCERTAINTY)
g1.add_experiment(t30)
g1.link_to_global(t30,"dH",vh.dH)
g1.link_to_global(t30,"K",vh.K)
g1.link_to_global(t30,"fx_competent","g_competent")
#--------------------------------------------------------------------------------------------------
t35 = pytc.ITCExperiment("temp-dependence/35C.DH",pytc.indiv_models.SingleSite,shot_start=1,uncertainty=HEAT_UNCERTAINTY)
g1.add_experiment(t35)
g1.link_to_global(t35,"dH",vh.dH)
g1.link_to_global(t35,"K",vh.K)
g1.link_to_global(t35,"fx_competent","g_competent")
g1.update_bounds("g_competent",(0.5,1.5))
g1.update_bounds("vh_K_ref",(1,np.inf))
# Do fit
g1.fit()
# Show the results
fit, ax = g1.plot()
c = g1.corner_plot()
print(g1.fit_as_csv)
# -
# #### Fit to the VantHoffExtended global connector to extract change in heat capacity (one more param)
# Titrations of recombinant human S100A14 with $ZnCl_{2}$ at $5$, $10$, $17$, $25$, $30$ and $35 ^{\circ}C$. Concentrations were: $[protein] = 110\ \mu M$ and $[ZnCl_{2}]=200\ mM$. Buffer was $25\ mM$ Trizma, $100\ mM$ NaCl, $pH\ 7.4$. Buffer was treated with chelex and filtered to $0.22\ \mu m$. Data collected using a GE ITC200.
# +
# --------------------------------------------------------------------
# Create a global fitting instance
g2 = pytc.GlobalFit()
vhe = pytc.global_connectors.VantHoffExtended("vhe")
#--------------------------------------------------------------------------------------------------
t5 = pytc.ITCExperiment("temp-dependence/5C.DH",pytc.indiv_models.SingleSite,shot_start=1,uncertainty=HEAT_UNCERTAINTY)
g2.add_experiment(t5)
g2.link_to_global(t5,"dH",vhe.dH)
g2.link_to_global(t5,"K",vhe.K)
g2.link_to_global(t5,"fx_competent","g_competent")
#--------------------------------------------------------------------------------------------------
t10 = pytc.ITCExperiment("temp-dependence/10C.DH",pytc.indiv_models.SingleSite,shot_start=1,uncertainty=HEAT_UNCERTAINTY)
g2.add_experiment(t10)
g2.link_to_global(t10,"dH",vhe.dH)
g2.link_to_global(t10,"K",vhe.K)
g2.link_to_global(t10,"fx_competent","g_competent")
#--------------------------------------------------------------------------------------------------
t17 = pytc.ITCExperiment("temp-dependence/17C.DH",pytc.indiv_models.SingleSite,shot_start=1,uncertainty=HEAT_UNCERTAINTY)
g2.add_experiment(t17)
g2.link_to_global(t17,"dH",vhe.dH)
g2.link_to_global(t17,"K",vhe.K)
g2.link_to_global(t17,"fx_competent","g_competent")
#--------------------------------------------------------------------------------------------------
t25 = pytc.ITCExperiment("temp-dependence/25C.DH",pytc.indiv_models.SingleSite,shot_start=1,uncertainty=HEAT_UNCERTAINTY)
g2.add_experiment(t25)
g2.link_to_global(t25,"dH",vhe.dH)
g2.link_to_global(t25,"K",vhe.K)
g2.link_to_global(t25,"fx_competent","g_competent")
#--------------------------------------------------------------------------------------------------
t30 = pytc.ITCExperiment("temp-dependence/30C.DH",pytc.indiv_models.SingleSite,shot_start=1,uncertainty=HEAT_UNCERTAINTY)
g2.add_experiment(t30)
g2.link_to_global(t30,"dH",vhe.dH)
g2.link_to_global(t30,"K",vhe.K)
g2.link_to_global(t30,"fx_competent","g_competent")
#--------------------------------------------------------------------------------------------------
t35 = pytc.ITCExperiment("temp-dependence/35C.DH",pytc.indiv_models.SingleSite,shot_start=1,uncertainty=HEAT_UNCERTAINTY)
g2.add_experiment(t35)
g2.link_to_global(t35,"dH",vhe.dH)
g2.link_to_global(t35,"K",vhe.K)
g2.link_to_global(t35,"fx_competent","g_competent")
g2.update_bounds("g_competent",(0.5,1.5))
g2.update_bounds("vhe_K_ref",(1,np.inf))
# Do fit
g2.fit()
# Show the results
fit, ax = g2.plot()
c = g2.corner_plot()
print(g2.fit_as_csv)
# -
# #### Use util.choose_model to select between the two models
pytc.util.compare_models(g1,g2)
| 11_use-aic-choose-model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + language="javascript"
# MathJax.Hub.Config({
# TeX: { equationNumbers: { autoNumber: "AMS" } }
# });
# -
# ## Exercise 2.15 MLE minimizes KL divergence to the empirical distribution
# Let $p_{emp}(x)$ be the empirical distribution, and let $q(x|\theta)$ be some model. Show that $\mathrm{argmin}_q\mathbb{KL}(p_{emp}||q)$
# is obtained by $q(x) = q(x; \theta)$, where $\theta$ is the MLE. Hint: use non-negativity of the KL divergence.
# ### Notes
# We will prove that the MLE minimises the KL divergence between the empirical distribution and some model. What does this mean? Suppose we get access to some data $\mathcal{D}$ generated by some unknown distribution and we are asked to estimate this distribution based on the data.
#
# In this situation, there are two approaches we could take: the **nonparametric** and the **parametric**. On the former approach, we could make an estimation based purely on the data we have at hand. This estimation could be the empirical distribution $p_{emp}(x)$, defined as
#
# $$
# p_{emp}(x) = \sum_{i=1}^N w_i\delta_{xi}(x)
# $$
#
# where we require $0\le w_i < 1$ and $\sum_{i=1}^N w_i = 1$.
#
# On the latter approach, we could make some reasonable assumptions about the data pattern and create a parametric model $q(x|\theta)$ for it. The model is based on the prior we have about the process behind the data and it dictates the general shape of the distribution (e.g. Gaussian). One of the most popular ways of estimating the parameters is the Maximum Likelihood Estimator (MLE). The MLE is the parameter value $\theta^*$ which solves the following optimization problem:
#
# \begin{equation}
# \theta^* = \mathrm{argmax}_\theta\prod_{x\in\mathcal{D}}p(x|\theta)
# \end{equation}
#
# The RHS of (1) is called the likelihood of the dataset and it is a function of $\theta$. Note that most of the times it's most convenient to optimize the log-likelihood, which gives the same result as optimizing the likelihood.
#
# $$
# \theta^* = \mathrm{argmax}_{\theta}\prod_{x\in\mathcal{D}}p(x|\theta) = \mathrm{argmax}_\theta\sum_{x\in\mathcal{D}}\log(p(x|\theta))
# $$
#
#
# Given two such different scenarios, this exercise proposes we find a bridge between them. We will find the parametric model $q(x|\theta)$ that is the most similar to the empirical distribution $p_{emp}(x)$. As you can imagine, one great way to do this is to express the similarity by the KL-divergence and minimize this value.
# ### Solution
#
# \begin{equation}
# \mathbb{KL}(p_{emp}||q) = \sum p_{emp}\log(\frac{p_{emp}}{q}) = \sum p_{emp}\log(p_{emp}) - \sum p_{emp}\log q
# \end{equation}
# Since $p_{emp}$ is fixed given the dataset $\mathcal{D}$, the first term of (1) is constant and we only need to focus on the second term.
#
# \begin{aligned}
# \mathrm{argmin}_\theta\mathbb{KL}(p_{emp}||q(\theta)) & = -\mathrm{argmin}_\theta\sum p_{emp}\log q \\
# & = \mathrm{argmax}_\theta\sum \log(q^{p_{emp}}) \\
# & = \mathrm{argmax}_\theta\log\left(\prod_{x\in\mathcal{D}} q^{p_{emp}}\right)
# \end{aligned}
# For the final step, note that the empirical distribution is defined as $p_{emp}(x_i) = N_i/N$, where $N_i$ is the number of occurrences of $x_i$ and $N$ is the size of our dataset.
#
# Thus:
#
# \begin{aligned}
# \mathrm{argmin}_\theta\mathbb{KL}(p_{emp}||q(\theta)) & = \mathrm{argmax}_\theta \log(\prod_{x\in\mathcal{D}}q^{p_{emp}}) \\
# & = \mathrm{argmax}_\theta\frac{1}{N}\log\left(\prod_{x\in\mathcal{D}}q(x_i|\theta)^{N_i}\right)
# \end{aligned}
#
# This is the same expression as the log-likelihood, with the exception of the $1/N$ term. Therefore the solution for this problem is the MLE parameter $\theta^*$.
# ### Conclusion
# In this exercise, we saw that the MLE of a parametric model is the most similar distribution to the empirical distribution, amongst all the models considered $(\theta\in\Theta)$. This makes a lot of sense - as the name tells, the maximum likelihood estimator is the parameter that maximizes the likelihood that our given dataset could occur. So its optimization is strongly based on the data (not taking into account a probability prior, like Bayesian models. The only prior is the shape of the distribution, like a Gaussian, which is given by the model). Therefore, as both the MLE and $p_{emp}$ have such a strong relationship with the empirical data, it makes sense that they have a strong relationship between themselves.
| murphy-book/chapter02/q15.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: torch0.4
# language: python
# name: torch0.4
# ---
# +
# Import packages
from unityagents import UnityEnvironment
import numpy as np
import random
import torch
from collections import deque
import matplotlib.pyplot as plt
# %matplotlib inline
# Set enviroment
env = UnityEnvironment(file_name="/media/riley/Work/PersonalProjects/AI/Deep_Reinforcement_Learning/Project1/Banana_Linux/Banana.x86_64")
# -
# Get the default brain
brain_name = env.brain_names[0]
brain = env.brains[brain_name]
# +
# reset the environment
env_info = env.reset(train_mode=True)[brain_name]
# number of agents in the environment
print('Number of agents:', len(env_info.agents))
# number of actions
action_size = brain.vector_action_space_size
print('Number of actions:', action_size)
# examine the state space
state = env_info.vector_observations[0]
print('States look like:', state)
state_size = len(state)
print('States have length:', state_size)
# -
# # DQN
# +
#from dqn_agent import Agent
from agent import DQNPRBAgent
from agent import DQNAgent
agent = DQNAgent(state_size=37, action_size=4, seed=0, use_dueling=True, use_double=True)
# +
def dqn(n_episodes=2000, eps_start=1.0, eps_end=0.01, eps_decay=0.995):
"""Train the Deep Q network."""
highest_score = 0
scores = []
scores_windows = deque(maxlen=100)
eps = eps_start
solved = False
for i_episode in range(1, n_episodes + 1):
env_info = env.reset(train_mode=True)[brain_name]
state = env_info.vector_observations[0]
score = 0
while True:
action = agent.act(state, eps)
# send the action to the environment
env_info = env.step(action)[brain_name]
# get the next state
next_state = env_info.vector_observations[0]
# get the reward
reward = env_info.rewards[0]
# see if episode has finished
done = env_info.local_done[0]
agent.step(state, action, reward, next_state, done)
# update the score
score += reward
# roll over the state to next time step
state = next_state
if done:
break
scores_windows.append(score)
scores.append(score)
eps = max(eps_end, eps * eps_decay)
if np.mean(scores_windows) > highest_score:
highest_score = np.mean(scores_windows)
print('\rEpisode {}\tAverage Score: {:.2f}'.format(i_episode, np.mean(scores_windows)), end="")
if i_episode % 100 == 0:
print('\rEpisode {}\tAverage Score: {:.2f}'.format(i_episode, np.mean(scores_windows)))
if np.mean(scores_windows)>=13.0 and not solved:
print('\nEnvironment solved in {:d} episodes!\tAverage Score: {:.2f}'.format(i_episode-100, np.mean(scores_windows)))
solved = True
if np.mean(scores_windows)>=13.0 and solved:
if np.mean(scores_windows) == highest_score:
print('\tModel Saved')
torch.save(agent.qnetwork_local.state_dict(), 'checkpoint.pth')
return scores
scores = dqn()
# plot the scores
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(len(scores[:2000])), scores[:2000])
plt.ylabel('Score')
plt.xlabel('Episode #')
plt.show()
# -
# ## Load and observe the trained agent
# +
import time
checkpoint = torch.load('checkpoint.pth')
agent = DQNAgent(state_size=37, action_size=4, seed=0, use_dueling=True, use_double=True)
agent.qnetwork_local.load_state_dict(checkpoint)
env_info = env.reset(train_mode=True)[brain_name]
state = env_info.vector_observations[0]
score = 0
while True:
time.sleep(0.025)
action = agent.act(state, 0)
env_info = env.step(action)[brain_name]
next_state = env_info.vector_observations[0]
reward = env_info.rewards[0]
done = env_info.local_done[0]
score += reward
state = next_state
if done:
break
print("Score: {}".format(score))
# -
env.close()
| Navigation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Detection Algorithm Results
# Read and analyze the results from running the detection algorithm on all APF data, which has already been deblazed, normalized, and velocity shifted by Anna Zuckerman using specmatch-emp.
import pandas as pd
import numpy as np
import matplotlib.pylab as plt
import os, glob #glob.glob(DIR_NAME + '/*fits')
from astropy.io import fits
# ## Read in Reduced Data from Anna Zuckerman's repo
# Read in all the spectra that was fed into the detection algorithm.
# +
directory = '/mnt_home/azuckerman/BL_APF_DAP/APF_spectra/NDR_corrected_wl_scale'
# every file in Anna's NDR_corrected_wl_scale folder
list_of_files = []
for filename in os.listdir(directory):
if filename.endswith(".fits"):
file_path = os.path.join(directory, filename)
list_of_files = np.append(list_of_files, file_path)
# -
# ## Look at total detections tables
# The spectra were divided into six bins to speed up the runtime of the detection algorithm. Combine all the individual tables into one table with all the results.
dir = '/mnt_home/zoek/code/APF-BL-DAP/Zoe/MyStuff/LaserDetectionResults/SNRBinningResults/'
results1000 = pd.read_csv(dir + 'LaserDetectionResults1/all_results.csv', index_col = 'Index')
results2000 = pd.read_csv(dir + 'LaserDetectionResults2/all_results.csv', index_col = 'Index')
results3000 = pd.read_csv(dir + 'LaserDetectionResults3/all_results.csv', index_col = 'Index')
results4000 = pd.read_csv(dir + 'LaserDetectionResults4/all_results.csv', index_col = 'Index')
results5000 = pd.read_csv(dir + 'LaserDetectionResults5/all_results.csv', index_col = 'Index')
results6000 = pd.read_csv(dir + 'LaserDetectionResults6/all_results.csv', index_col = 'Index')
total_results = results1000.append([results2000, results3000, results4000, results5000, results6000])
total_results
# ## Add SNR data to table
# Use SNR.ipynb to calculate the S/N ratios for all the spectra.
# read or create an array containing the SNRs of all the targets
SNR = np.load('SNR.npy')
# Select SNR values corresponding to the targets we ran the algorithm on.
# This step is only necessary if we omitted certain targets from the list.
indicies = total_results.index
actual_SNR = []
for i in indicies:
s = SNR[i]
actual_SNR = np.append(actual_SNR, s)
total_results['SNR'] = actual_SNR
# +
# Save results
# total_results.to_csv('/home/zoek/code/APF-BL-DAP/Zoe/LaserDetectionResults.csv')
# -
# ## Create a histogram of all the detections across all the spectra
num_detections = total_results['# Wider than PSF']
# plt.rcParams["figure.figsize"] = (18,12)
# plt.rcParams.update({'font.size': 50})
plt.hist(num_detections.tolist(), range=[0, 20])
plt.xlabel('Number of Detections')
plt.ylabel('Number of Spectra')
plt.title('Number of Detections across all Spectra')
plt.show()
# total number of detections
sum(num_detections)
# ## Create a histogram of all the detections across all unique stars
aggregation_functions = {'# Wider than PSF': 'median'}
total_results_by_star = total_results.groupby(total_results['Star']).aggregate(aggregation_functions)
stars = total_results_by_star.index.to_numpy()
print(len(stars))
print('there are ' + str(total_results_by_star.shape[0]) + ' unique stars')
num_detections_by_star = total_results_by_star['# Wider than PSF']
max_num_detections = max(num_detections_by_star)
print('max # of detections: ' + str(max_num_detections))
# plt.rcParams["figure.figsize"] = (10,5)
# plt.rcParams.update({'font.size': 20})
plt.hist(num_detections_by_star, range=[0, 20])
plt.xlabel('Number of Detections')
plt.ylabel('Number of Stars')
plt.title('Detection Counts Histogram')
plt.show()
# ## Find percentage of stars that have a certain # of detections
num_stars = total_results_by_star.shape[0]
zero_detections = total_results_by_star[total_results_by_star['# Wider than PSF'] < 0.5]
num_zero_detections = zero_detections.shape[0]
print(str(num_zero_detections * 100 / num_stars) + '% of the stars have 0 detections' + ' (' + str(num_zero_detections) + ' stars)')
less_than_10_detections = total_results_by_star[total_results_by_star['# Wider than PSF'] < 10]
num_less_than_10_detections = less_than_10_detections.shape[0]
print(str(num_less_than_10_detections * 100 / num_stars) + '% of the stars have under 10 detections' + ' (' + str(num_less_than_10_detections) + ' stars)')
test = total_results[total_results[['# Wider than PSF']] == 0]
test.shape
# ## Observe the relationship between SNR and number of detections
num_detections = total_results['# Wider than PSF'].to_numpy()
SNR = total_results['SNR'].to_numpy()
# +
plt.rcParams["figure.figsize"] = (18,12)
plt.rcParams.update({'font.size': 30})
plt.scatter(SNR, num_detections) # S/N 225 and above,
plt.axvline(x=10, c='gray')
plt.axvline(x=150, c='gray')
plt.xlabel('SNR')
plt.ylabel('# Detections')
plt.title('Relationship between SNR and # Detections')
plt.show()
# -
all_s = []
for i in np.arange(len(SNR)):
s = SNR[i]
n = num_detections.tolist()[i]
for j in np.arange(n):
all_s = np.append(all_s, s)
plt.hist(all_s)
plt.xlabel('SNR')
plt.ylabel('# of Detections')
plt.title('SNR vs. Number of Detections')
plt.show()
# +
fig, ax = plt.subplots()
# plot the cumulative histogram
ax.hist(all_s, cumulative=True)
plt.axvline(x=150, color = 'black')
plt.xlabel('SNR')
plt.ylabel('Cumulative # of Detections')
plt.title('SNR vs. Cumulative Number of Detections')
plt.show()
# -
# ## IGNORE BELOW
max_detections = total_results[total_results.SNR > 150]
max_detections
# max_detections.sort_values(by=['ndetections'], ascending=True)
high_SNR = total_results[total_results.index == 415]
high_SNR.sort_values(by=['SNR'], ascending=False)
# +
exposure_times = []
for file in list_of_files:
file = fits.open(file)
header = file[0].header
exposure_time = header['EXPTIME']
exposure_times = np.append(exposure_times, exposure_time)
# print('File path: ' + APF_flux_path)
# print('Star: ' + star)
# print(exposure_time)
# plt.figure()
# plt.plot(wl, flux)
# plt.ylim(0, 1.5)
# plt.xlim(5880, 5920)
# plt.show()
# 14, 21, 24, 26, 27
# 20, 28
# weird 38
# spect = flux
# -
print(len(exposure_times))
new1 = np.delete(exposure_times, 1762)
new2 = np.delete(new1, 1760)
final_exposure_times = new2.tolist()
plt.scatter(final_exposure_times, num_detections)
plt.axvline(x=20, c='gray')
plt.axhline(y=20, c='gray')
# plt.ylim(1199, 1201)
plt.xlabel('Exposure Times')
plt.ylabel('# Detections')
plt.title('Relationship between Exposure Time and # Detections')
plt.show()
| Zoe/AnalyzeResults.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Functions
# ### Keyword and positional arguments
#
# * default argument values can be passed, even unordered, using keyword arguments (named arguments)
# * keyword arguments are passed as a dictionary {key:value,...}
# * not keyword arguments are called positional arguments
# * positional arguments should be starred if after a keyword argument
# * \*expression must evaluate an iterable
def foo(*positional, **keywords):
print("Positional:", positional, end='\t')
print("Keywords:", keywords)
foo('1st', '2nd', '3rd')
foo(par1='1st', par2='2nd', par3='3rd')
foo('1st', par2='2nd', par3='3rd')
foo(par1='1st_key',*('tuple','unpacking'), par2='2nd_key')
foo(par1='1st_key',*['1st_pos'], par2='2nd_key',*['2st_pos','3rd_pos'])
# ### more on tuple unpacking
a,b,*rest = range(5)
a,b,rest
a,*body,c,d = range(5)
print(a,body,c,d)
*head, a,b, *wrong = range(5) # only one * is allowed
*_, last = range(5)
print(last)
# ### nested tuples
# +
cities = [('Tokyo', 'JP', 'un', 'important', 'fields',(35.689,139.692)),
('San Paulo', 'BR','not', 'relevant', 'fields',(-23.547,-46.6358))]
for city, *_, latitude, longitude in cities:
print(city, latitude, longitude)
| lectures/lectures/python/01_intro/07_functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data cleaning - American dream
# +
#Modules
import pandas as pd
import sqlalchemy
import mysql.connector
import sys
sys.path.insert(0, "/home/apprenant/PycharmProjects/American_dream")
from src.d00_utils.mysql_func import mysql_connect, save_to_mysql
connect = mysql_connect()
# -
# ## 1 - Importation des données
df1 = pd.read_sql("survey_1",con=connect)
dfk = pd.read_sql("survey_k",con=connect)
# ## Modifications
# ### Vérifications des doublons
print(df1.duplicated().value_counts())
df1.shape
df1 = df1.drop(['index'], axis=1)
print(df1.duplicated(subset=df1.columns.difference(['Timestamp'])))
duplicated_rows = df1[df1.duplicated(subset = df1.columns.difference(['Timestamp']), keep=False)]
duplicated_rows.head(10)
df1 = df1.drop(['Timestamp'], axis=1)
print(df1.duplicated().value_counts())
# ### Tri pour sélectionner seulement les données des Etats-Unis
df1.shape
df1 = df1.loc[df1['Country'] == 'United States',:]
df1.shape
# ## 2 - Sélection des colonnes
# ### Database 1
print(df1.head())
df1.shape
print(df1.columns)
# J'ai choisi les colonnes qui me semblaient utiles pour la visualisation des données.
df1=df1[['SalaryUSD', 'Country', 'PostalCode', 'EmploymentStatus', 'JobTitle', 'ManageStaff', 'YearsWithThisTypeOfJob', 'HowManyCompanies', 'OtherPeopleOnYourTeam', 'CompanyEmployeesOverall', 'EmploymentSector', 'LookingForAnotherJob', 'CareerPlansThisYear', 'Gender']]
# ### Database 2
print(dfk.head())
dfk.shape
print(dfk.columns)
# J'ai ensuite choisi les colonnes qui semblaient utiles pour la suite.
dfk=dfk[['Job Title', 'Salary Estimate', 'Company Name', 'Location', 'Size', 'Sector']]
# ## 3 - Valeurs manquantes
# ### Database 1
missing_values_count = df1.isnull().sum()
print(missing_values_count)
# Il manque 1390 valeurs pour PostalCode. On ne peut pas assigner de valeur de code postal arbitraire dans cette colonne. Je garde donc les lignes où les valeurs sont manquantes, mais je garde en mémoire qu'il manque des valeurs si je travaille sur les codes postaux.
# ### Database 2
print(dfk.isnull().sum())
# Il y a une seule valeur manquante pour 'Company Name'.
# J'ai choisi de la remplacer par 'Unknown'
dfk.loc[dfk['Company Name'].isnull(), "Company Name"] = "Unknown"
print(dfk.isnull().sum())
# ## 4 - Colonnes à transformer en datetime
# ### Database 1
print(df1.dtypes)
# Aucune de mes colonnes ne correspond à une date.
# ### Database 2
print(dfk.dtypes)
# Aucune de mes colonnes ne correspond à une date.
# ## 5 - Doublons
# ### Database 1
# Fait plus haut
print(df1.shape)
# ### Database 2
print(dfk.duplicated().value_counts())
# Cette database ne semble pas avoir de doublons.
# ## 6 - Vérifier la consistence des données
# ### Database 1
# #### get all the unique values in the 'JobTitle' column
job_titles = df1['JobTitle'].unique()
print(len(job_titles))
job_titles.sort()
print(job_titles)
# Il y a 21 job titles différents.
# ### Modifications : regrouper les catégories de Job Title
df1["JobTitle"] = df1["JobTitle"].apply(
lambda x: "Database Admin" if x.startswith('DBA') else x
)
df1["JobTitle"] = df1["JobTitle"].apply(
lambda x: "Developer" if x.startswith('Developer') else x
)
job_titles = df1['JobTitle'].unique()
print(len(job_titles))
job_titles.sort()
print(job_titles)
s = df1["JobTitle"].value_counts()
df1.loc[df1["JobTitle"].isin(s[(s < 5)].index), "JobTitle"] = "Other"
job_titles = df1['JobTitle'].unique()
print(len(job_titles))
job_titles.sort()
print(job_titles)
# #### get all the unique values in 'HowManyCompanies' column
nb_companies = df1['HowManyCompanies'].unique()
print(len(nb_companies))
nb_companies.sort()
print(nb_companies)
df1["HowManyCompanies"] = df1["HowManyCompanies"].apply(
lambda x: "1" if str(x).startswith('1') else "2" if str(x).startswith('2') else x
)
nb_companies = df1['HowManyCompanies'].unique()
print(len(nb_companies))
nb_companies.sort()
print(nb_companies)
# Les valeurs pour HowManyCompanies sont classées dans 7 catégories. Il n'y a pas de catégorie redondante.
# #### get all the unique values in the 'OtherPeopleOnYourTeam' column
team = df1['OtherPeopleOnYourTeam'].unique()
print(len(team))
team.sort()
print(team)
# Les valeurs pour OtherPeopleOnYourTeam sont classées dans 7 catégories. Il n'y a pas de catégorie redondante.
# #### get all the unique values in the 'Gender' column
genders = df1['Gender'].unique()
print(len(genders))
# Il semble y avoir beaucoup de réponses différentes dans la colonne 'Gender
genders.sort()
print(genders)
print(df1['Gender'].value_counts())
# Sur la totalité des réponses dans la colonne Genre, on préférerait pouvoir les classer en 5 catégories : Male, Female, Non-binary/third gender, Prefer not to say et Not Asked.
# Il y a donc 29 réponses à modifier.
valid_genders = ["Male", "Female", "Not Asked", "Prefer not to say", "Non-binary/third gender"]
df1.loc[~df1['Gender'].isin(valid_genders), "Gender"] = "other"
print(df1['Gender'].value_counts())
# ### Database 2
titles = dfk['Job Title'].unique()
print(len(titles))
print(len(dfk['Salary Estimate'].unique()))
# ## 7 - Recherche de valeurs aberrantes
# ### Database 1
import seaborn as sns
sns.boxenplot(x = df1["SalaryUSD"])
sns.boxenplot(x = df1["YearsWithThisTypeOfJob"])
# ### Database 2
# ### Sauvegarde des databases dans mysql
save_to_mysql(db_connect=connect,df_to_save=df1,df_name='survey_1_v1.0')
save_to_mysql(db_connect=connect,df_to_save=df1,df_name='survey_1_v2.0')
| Notebook/Data Cleaning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # From one dataset to many
# **Load the data**
import pandas as pd
data = pd.read_csv('../data/fake_beiersdorf_data.csv')
data.shape
data.head()
# ## Split large dataset into many small ones based on the variable `Name`
data.Name
data.Name.unique()
data.Name.nunique()
for name in data.Name.unique():
print(f'Computing {name} ...')
subset = data.loc[data.Name == name]
subset.sample(10)
import os
for name in data.Name.unique():
print(f'Computing {name} ...')
subset = data.loc[data.Name == name]
fname = f'subset_{name.replace(" ", "_")}.csv'
filepath = os.path.join('..', 'data', 'interim', fname)
subset.to_csv(filepath, index=False)
# ***
# # From many to one
# !ls ../data/interim/
import glob
filelist = glob.glob(os.path.join('..', 'data', 'interim', '*.csv'))
filelist
dfs = []
for f in filelist:
data = pd.read_csv(f)
dfs.append(data)
fulldata = pd.concat(dfs)
fulldata.shape
fulldata.sample(10)
# ***
| 03_automate_the_boring_stuff/notebooks/01a_from_one_2_many_and_many_to_one.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import keras
import os
#from params import get_params
from sklearn import preprocessing
import sklearn.preprocessing
import numpy as np
import matplotlib.pyplot as plt
from keras.models import Model
from keras.preprocessing import image
from keras.applications.vgg19 import VGG19
from keras.applications.vgg19 import preprocess_input
from PIL import Image, ImageOps
import pickle
# -
descriptors_val = pickle.load(open("save_val.p", "rb"))
descriptors_train = pickle.load(open("save_train.p", "rb"))
imagen = pickle.load(open("save_img.p", "rb"))
imagen3 = pickle.load(open("save_img3.p", "rb"))
dic_val = pickle.load(open("save_dic_val.p", "rb"))
dic_train = pickle.load(open("save_dic_train.p", "rb"))
x_val = np.reshape(descriptors_val, (477,4096))
x_train = np.reshape(descriptors_train, (1194,4096))
x_val = sklearn.preprocessing.normalize(x_val, norm='l2', axis=1, copy=True, return_norm=False)
x_train = sklearn.preprocessing.normalize(x_train, norm='l2', axis=1, copy=True, return_norm=False)
descriptors_traint = x_train.transpose()
similarities=np.matmul(x_val,descriptors_traint)
ranks = np.argsort(similarities, axis=1)[:,::-1]
# get the original images for visualization
x_val_images = []
x_train_images = []
v = 0
for v in range(477):
x_val_images.append(np.array(imagen[v]))
b = 0
for b in range(1194):
x_train_images.append(np.array(imagen3[b]))
h,w = (224, 224)
new_image= Image.new('RGB', (h*6,w*13))
# Visualize ranks of the 10 queries
offset = 0 # it will show results from query #'offset' to #offset+10
relnotrel = []
for q in range(477):
ranks_q = ranks[q*(offset+1),:]
relevant = dic_val[q*(offset+1)]
rel_help = []
for i in range(1194):
if relevant == dic_train[ranks_q[i]] and relevant != "desconegut":
new_image.paste(ImageOps.expand(Image.fromarray(x_train_images[ranks_q[i]]), border=10, fill='green'), (h*(1+i),w*q))
rel_help.append(1)
else:
new_image.paste(ImageOps.expand(Image.fromarray(x_train_images[ranks_q[i]]), border=10, fill='red'), (h*(1+i),w*q))
rel_help.append(0)
relnotrel.append(np.asarray(rel_help))
# visualize query
ima_q = Image.fromarray(x_val_images[q*(offset+1)])
ima_q = ImageOps.expand(ima_q, border=10, fill='blue')
new_image.paste(ima_q, (0,w*q))
plt.imshow(new_image)
plt.axis('off')
plt.show()
# +
#new_image.save('val1.png')
# -
guarda = []
accu_array =[]
AP = []
numRel_tot = []
for t in range(len(relnotrel)):
numRel = 0
accu = 0
graphic = []
for k in range(len(relnotrel[t])):
# If the value is 1
if relnotrel[t][k] == 1:
# We add 1 to the number of correct instances
numRel = numRel + 1
# We calculate the precision at k (+1 because we start at 0)
# and we accumulate it
accu += float(numRel)/ float(k+1)
graphic.append(float( numRel )/ float(k+1))
if numRel != 0:
numRel_tot.append(numRel)
accu_array.append(accu)
guarda.append(np.asarray(graphic))
AP = []
for h in range(len(numRel_tot)):
AP.append(float(accu_array[h])/float(numRel_tot[h]))
MAP = np.sum(AP)/len(numRel_tot)
MAP * 100
plt.plot(guarda[0])
plt.plot(guarda[10])
guarda2 = []
for t2 in range(len(relnotrel)):
graphic2 = []
len_final = 0
numRel2 = 0
for k2 in range(len(relnotrel[t2])):
# If the value is 1
if relnotrel[t2][k2] == 1:
# We add 1 to the number of correct instances
numRel2 = numRel2 + 1
graphic2.append(float( numRel2 )/ float(numRel))
guarda2.append(np.asarray(graphic2))
plt.plot(guarda2[0])
plt.plot(guarda2[10])
p = 0
guarda[p][0]= 1.0
guarda2[p][0]= 0.0
plt.plot(guarda2[p],guarda[p])
plt.title('Precision-Recall')
plt.xlabel('Recall')
plt.ylabel('Precision')
plt.show()
| Layers/VGG19/cerca_visual_fc2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Import the necessary packages and data
from collections import defaultdict
from datetime import datetime
import matplotlib.pyplot as plt
import numpy as np
import os
import pandas as pd
import seaborn as sns
sns.set()
path = os.path.expanduser('~/Projects/capstone-two/data/interim/Clean_Harbor_Water_Quality.csv', )
wq_df = pd.read_csv(path, parse_dates=['Sample Date'], index_col=0)
row_counts = wq_df.notnull().sum(axis=1)
_ = plt.bar(x=row_counts.value_counts().index,
height=row_counts.value_counts().values)
# I want to use only rows with at least 30 columns complete. This is to prevent my data from being overloaded with imputed data.
wq_df['row_sums'] = wq_df.notnull().sum(axis=1)
wq_df[wq_df['row_sums'] < 30]
wq_df = wq_df.drop(columns=['row_sums'])
# Next I will fill in blank values
ffill_df = pd.DataFrame()
for site in wq_df['Sampling Location'].unique():
site_df = wq_df[wq_df['Sampling Location'] == site].sort_values(by='Sample Date')
site_df = site_df.fillna(method='ffill')
ffill_df = ffill_df.append(site_df)
final_df = pd.DataFrame()
for site in ffill_df['Sampling Location'].unique():
site_df = ffill_df[ffill_df['Sampling Location'] == site].sort_values(by='Sample Date')
site_df = site_df.fillna(method='bfill')
final_df = final_df.append(site_df)
final_df = final_df.sort_values('Sample Date')
final_df = final_df.reset_index(drop=True)
final_df = final_df.dropna()
# Make dummy columns for dry/wet
final_df = pd.get_dummies(final_df, columns=['Weather Condition (Dry or Wet)'])
final_df = final_df.drop(columns=['Weather Condition (Dry or Wet)_D'])
# I noticed something weird with the pH values, so I need to fix them.
final_df.nsmallest(10, 'Top PH')['Top PH']
final_df = final_df.drop(index=[29360])
final_df = final_df.reset_index(drop=True)
# Different parameters will serve different purposes in my analysis. Some measurements have standard limits that must not be exceeded. The rest of the columns will be potential indicators of poor water quality. I have outlined each column's limit/indicator status below:
#
# - Water Temperature = Indicator
#
# - Salinity = Indicator
#
# - Conductivity = Indicator
#
# - Dissolved Oxygen = 3.0 mg/L
#
# - Light Transparency = Indicator
#
# - Sigma-T = Indicator
#
# - Secchi Depth = Indicator
#
# - pH = 6.0 - 9.5
#
# - Fecal Coliform = 2000 cells/100mL
#
# - Enterococcus = 104 cells/100mL
#
# - Ammonia = 2.0 mg/L
#
# - Ortho-phosphate = 0.20 mg/L
#
# - Kjeldahl Nitrogen = 10.0 mg/L
#
# - Silica = Indicator
#
# - Total Suspended Solids = Indicator
#
# - Chlorophyll a = 30
#
# - Dissolved Organic Carbon = Indicator
#
# +
# Make functions to
def check_do(measure):
'''Function checks Dissolved Oxygen
concentration against established
standards and returns 1 if test
fails (ie below 3.0).'''
if measure < 3.0:
return 1
else:
return 0
def check_pH(measure):
'''Function checks pH levels
against established standards.
Returns 1 if test fails (ie less
than 6.5 or greater than 9.5)'''
if measure > 9.5 or measure < 6.5:
return 1
else:
return 0
def check_fecal(measure):
'''Function checks Fecal Coliform
concentration agains established
standards. Returns 1 if test
fails (ie above 2000)'''
if measure > 2000:
return 1
else:
return 0
def check_enter(measure):
'''Function checks Enterococci
concentration against established
standards. Returns 1 if test fails
(ie above 104)'''
if measure > 104:
return 1
else:
return 0
def check_nitrate(measure):
'''Function checks Nitrate
concentration against established
standards. Returns 1 if test fails
(ie above 10)'''
if measure > 10:
return 1
else:
return 0
def check_ammonia(measure):
'''Function checks Ammonia
concentration against established
standards. Returns 1 if test fails
(ie above 2)'''
if measure > 2:
return 1
else:
return 0
def check_phosphate(measure):
'''Function checks Phosphate
concentration against established
standards. Returns 1 if test fails
(ie above 0.20)'''
if measure > 0.20:
return 1
else:
return 0
def check_chloro(measure):
'''Function checks Chlorophyll A
concentration against established
standards. Returns 1 if test fails
(ie above 30)'''
if measure > 30:
return 1
else:
return 0
# Make a dictionary of each parameter with limits and assign its check function
check_funcs = {'Top Ammonium (mg/L)':check_ammonia,
'Top Active Chlorophyll \'A\' (µg/L)':check_chloro,
'CTD (conductivity, temperature, depth profiler) Top Dissolved Oxygen (mg/L)':check_do,
'Top Enterococci Bacteria (Cells/100mL)':check_enter,
'Top Fecal Coliform Bacteria (Cells/100mL)':check_fecal,
'Top Total Kjeldhal Nitrogen (mg/L)':check_nitrate,
'Top PH':check_pH,
'Top Ortho-Phosphorus (mg/L)':check_phosphate
}
# -
# Create columns that signify whether a measurement is outside limits
fail_col = []
for col, func in check_funcs.items():
new_col = col + ' Failure'
fail_col.append(new_col)
final_df[new_col] = final_df[col].apply(func)
final_df.loc[:, fail_col].sum()
# # Correlations
# Since a lot of my data have varying scales, I am using Spearman's Correlation rather than Pearson's.
corr_df = final_df.corr('spearman')[['Top Active Chlorophyll \'A\' (µg/L)',
'Top Enterococci Bacteria (Cells/100mL)',
'Top Ortho-Phosphorus (mg/L)']]
corr_df
# # Strong Correlations
# ## Chlorophyll A
# - Top pH
# - Top Silica
#
# ## Enterococci
# - Top Salinity
# - Bottom Salinity
# - Top Conductivity
# - Bottom Conductivity
# - Top Sigma-T
# - Fecal Coliform
# - Top Silica
# - Weather Condition
# - Fecal Coliform Failure
#
# ## Ortho-phosphorus
# - Top pH
# - Ammonium
# - Total Kjeldhal Nitrogen
# - Top Silica
# # Chlorophyll A
chloro_corr_df = final_df.corr('spearman')['Top Active Chlorophyll \'A\' (µg/L)']
# +
chloro_corr_col = chloro_corr_df[(chloro_corr_df.abs() > 0.3)].index
chloro_corr_col = chloro_corr_col.drop(labels=['Top Active Chlorophyll \'A\' (µg/L)',
'Top Active Chlorophyll \'A\' (µg/L) Failure'])
#sub = 211
for col in chloro_corr_col:
# _ = plt.subplot(sub)
_ = plt.figure(figsize=(16,10))
_ = plt.scatter(x=final_df['Top Active Chlorophyll \'A\' (µg/L)'],
y=final_df[col], alpha=0.1)
_ = plt.xlabel('Top Active Chlorophyll A')
_ = plt.ylabel(col)
_ = plt.xscale('log')
_ = plt.title(col)
# sub += 1
_ = plt.show()
# -
# # Enterococci
ent_corr_df = final_df.corr('spearman')['Top Enterococci Bacteria (Cells/100mL)']
ent_corr_col = ent_corr_df[(ent_corr_df.abs() > 0.3)].index
ent_corr_col = ent_corr_col.drop(labels=['Top Enterococci Bacteria (Cells/100mL)',
'Top Enterococci Bacteria (Cells/100mL) Failure'])
# +
sub = 521
_ = plt.figure(figsize=(12,16))
for col in ent_corr_col:
_ = plt.subplot(sub)
_ = plt.scatter(x=final_df['Top Enterococci Bacteria (Cells/100mL)'],
y=final_df[col], alpha=0.4)
_ = plt.xscale('log')
_ = plt.xlim((10e0, 10e6))
_ = plt.title(col)
sub += 1
_ = plt.show()
# -
# # Ortho-Phosphorus
pho_corr_df = final_df.corr('spearman')['Top Ortho-Phosphorus (mg/L)']
pho_corr_col = pho_corr_df[(pho_corr_df.abs() > 0.3)].index
pho_corr_col = pho_corr_col.drop(labels=['Top Ortho-Phosphorus (mg/L)',
'Top Ortho-Phosphorus (mg/L) Failure'
]
)
sub = 221
_ = plt.figure(figsize=(16,12))
for col in pho_corr_col:
if np.dtype(final_df[col]) != object:
_ = plt.subplot(sub)
_ = plt.scatter(x=final_df['Top Ortho-Phosphorus (mg/L)'],
y=final_df[col])
_ = plt.title('Ortho-Phosphorus vs. ' + col)
_ = plt.xlabel('Top Ortho-Phosphorus (mg/L)')
_ = plt.ylabel(col)
sub += 1
_ = plt.show()
# Taking a look at the totals for each Failure column, the most common pollutant is Chlorophyll A by a followed by Enterococci and Ortho-Phosphorus. I'm going to investigate these three parameters in greater depth.
# +
_ = plt.figure(figsize=(12, 12))
for location in final_df['Sampling Location'].unique():
num = final_df[final_df['Sampling Location'] == location].loc[:, fail_col].sum().sum()
den = final_df[final_df['Sampling Location'] == location].shape[0]
ht = num / den / 8 * 100
_ = plt.bar(x = location,
height = ht)
_ = plt.title('Percent Test Failure at Each Sample Site',
fontsize=30)
_ = plt.xlabel('Sampling Site', fontsize=18)
_ = plt.ylabel('Percentage of Failed Tests', fontsize=18)
plt.xticks(rotation='vertical')
plt.tight_layout()
plt.show()
# -
# I want to see if there are significant differences between samples that fail to meet EPA limits.
#
# - The null hypothesis is that there is no significant difference between the mean values for samples that exceed limits and those that don't.
# - The alternate hypothesis is that there is a significant difference between the means of both.
#
# My original $\alpha$ value is 0.05, but since I am testing 39 columns simultaneously, I need to perform a Bonferroni correction on my $\alpha$ by dividing 0.05 by 39. After doing so, my new $\alpha$ = 0.00128.
#
# I will be performing this for Top Enterococci Bacteria, Top Active Chlorophyll A, and Top Ortho-Phosphate.
# # Compare failures to non failed starting with Enterococci
# +
failed_e = final_df[final_df['Top Enterococci Bacteria (Cells/100mL) Failure'] == 1]
passed_e = final_df[final_df['Top Enterococci Bacteria (Cells/100mL) Failure'] == 0]
failed_e = failed_e.drop(columns=['Sampling Location',
'Sample Date',
'Sample Time',
'Lat', 'Long'])
passed_e = passed_e.drop(columns=['Sampling Location',
'Sample Date',
'Sample Time',
'Lat', 'Long'])
# +
from statsmodels.stats import weightstats
h_o = zip(failed_e.columns, weightstats.ztest(passed_e, failed_e)[1])
def sig(value, alpha):
if value < alpha:
return 'Significant'
else:
return 'Not significant'
for i in list(h_o):
if sig(i[1], 0.00128) == 'Significant':
print(i[0])
# -
# # Chlorophyll A Differences
# +
failed_c = final_df[final_df['Top Active Chlorophyll \'A\' (µg/L) Failure'] == 1]
passed_c = final_df[final_df['Top Active Chlorophyll \'A\' (µg/L) Failure'] == 0]
failed_c = failed_c.drop(columns=['Sampling Location',
'Sample Date',
'Sample Time',
'Lat', 'Long'])
passed_c = passed_c.drop(columns=['Sampling Location',
'Sample Date',
'Sample Time',
'Lat', 'Long'])
# +
from statsmodels.stats import weightstats
h_o = zip(failed_c.columns, weightstats.ztest(passed_c, failed_c)[1])
def sig(value, alpha):
if value < alpha:
return 'Significant'
else:
return 'Not significant'
for i in list(h_o):
if sig(i[1], 0.00128) == 'Significant':
print(i[0])
# -
# # Ortho-Phosphate Differences
# +
failed_p = final_df[final_df['Top Enterococci Bacteria (Cells/100mL) Failure'] == 1]
passed_p = final_df[final_df['Top Enterococci Bacteria (Cells/100mL) Failure'] == 0]
failed_p = failed_p.drop(columns=['Sampling Location',
'Sample Date',
'Sample Time',
'Lat', 'Long'])
passed_p = passed_p.drop(columns=['Sampling Location',
'Sample Date',
'Sample Time',
'Lat', 'Long'])
# +
from statsmodels.stats import weightstats
h_o = zip(failed_p.columns, weightstats.ztest(passed_p, failed_p)[1])
def sig(value, alpha):
if value < alpha:
return 'Significant'
else:
return 'Not significant'
for i in list(h_o):
if sig(i[1], 0.00128) == 'Significant':
print(i[0])
# -
# I am going to look at monthly failures
final_df['Month'] = final_df['Sample Date'].map(lambda x: x.month)
pivot = final_df.pivot_table(values=['Top Active Chlorophyll \'A\' (µg/L) Failure',
'Top Enterococci Bacteria (Cells/100mL) Failure',
'Top Ortho-Phosphorus (mg/L) Failure'
],
index='Month', aggfunc='sum')
_ = pivot.plot(kind='bar', figsize=(14, 12))
_ = plt.xticks(np.arange(0, 12),
labels=['Jan', 'Feb', 'Mar', 'Apr',
'May', 'Jun', 'Jul', 'Aug',
'Sep', 'Oct', 'Nov', 'Dec'],
rotation=45)
_ = plt.title('Number of Failed Tests By Month', fontsize=25)
_ = plt.show()
# There are clearly temportal trends in failed tests.
#
# Now that I have a sense of my data, I can begin performing analysis.
outpath = os.path.expanduser('~/Projects/capstone-two/data/processed/Final_Clean_Harbor_Water_Quality.csv')
final_df.to_csv(outpath)
| notebooks/2.0-dra-data-exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tokenisation
#
# The notebook contains three types of tokenisation techniques:
# 1. Word tokenisation
# 2. Sentence tokenisation
# 3. Tweet tokenisation
# 4. Custom tokenisation using regular expressions
# ### 1. Word tokenisation
document = "At nine o'clock I visited him myself. It looks like religious mania, and he'll soon think that he himself is God."
print(document)
# Tokenising on spaces using python
print(document.split())
# Tokenising using nltk word tokeniser
import nltk
nltk.download('punkt')
# + tags=[]
from nltk.tokenize import word_tokenize
words = word_tokenize(document)
# -
print(words)
# NLTK's word tokeniser not only breaks on whitespaces but also breaks contraction words such as he'll into "he" and "'ll". On the other hand it doesn't break "o'clock" and treats it as a separate token.
# ### 2. Sentence tokeniser
# Tokenising based on sentence requires you to split on the period ('.'). Let's use nltk sentence tokeniser.
# + tags=[]
from nltk.tokenize import sent_tokenize
sentences = sent_tokenize(document)
# -
print(sentences)
# ### 3. Tweet tokeniser
# A problem with word tokeniser is that it fails to tokeniser emojis and other complex special characters such as word with hashtags. Emojis are common these days and people use them all the time.
# + tags=[]
message = "i recently watched this show called mindhunters:). i totally loved it 😍. it was gr8 <3. #bingewatching #nothingtodo 😎"
# -
print(word_tokenize(message))
# The word tokeniser breaks the emoji '<3' into '<' and '3' which is something that we don't want. Emojis have their own significance in areas like sentiment analysis where a happy face and sad face can salone prove to be a really good predictor of the sentiment. Similarly, the hashtags are broken into two tokens. A hashtag is used for searching specific topics or photos in social media apps such as Instagram and facebook. So there, you want to use the hashtag as is.
#
# Let's use the tweet tokeniser of nltk to tokenise this message.
# + tags=[]
from nltk.tokenize import TweetTokenizer
tknzr = TweetTokenizer()
# -
tknzr.tokenize(message)
# As you can see, it handles all the emojis and the hashtags pretty well.
#
# Now, there is a tokeniser that takes a regular expression and tokenises and returns result based on the pattern of regular expression.
#
# Let's look at how you can use regular expression tokeniser.
# + tags=[]
from nltk.tokenize import regexp_tokenize
message = "i recently watched this show called mindhunters:). i totally loved it 😍. it was gr8 <3. #bingewatching #nothingtodo 😎"
pattern = "#[\w]+"
# -
regexp_tokenize(message, pattern)
from nltk.tokenize import ToktokTokenizer
tb = ToktokTokenizer()
tb.tokenize(message)
| 9. NLP/1. Lexical Processing/2. Basic Lexical Processing/2. tokenisation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Toy model for estimation of connectivity in MOU process
#
# This Python notebook estimates the connectivity matrix from the observed activity generated by a MOU process.
import os
import numpy as np
import scipy.linalg as spl
import scipy.stats as stt
from pymou import MOU
import pymou.tools as tools
import matplotlib.pyplot as pp
# ## Simulation
# +
N = 30 # number of nodes
d = 0.3 # density of connectivity
# generate random matrix
C_orig = tools.make_rnd_connectivity(N, density=d, w_min=0.5/N/d, w_max=1.0/N/d)
# create MOU process
mou_orig = MOU(C_orig)
T = 1000 # time in seconds
# simulate
ts_sim = mou_orig.simulate(T)
# plots
pp.figure()
pp.plot(range(T),ts_sim)
pp.xlabel('time')
pp.ylabel('activity')
pp.title('simulated MOU signals')
D = np.linalg.eigvals(C_orig)
pp.figure()
pp.scatter(np.real(D),np.imag(D))
pp.plot([1,1],[-1,1],'--k')
pp.xlabel('real part of eigenvalue')
pp.ylabel('imag part of eigenvalue')
pp.title('spectrum of original C')
pp.show()
# +
Q_sim = np.tensordot(ts_sim,ts_sim,axes=(0,0)) / (T-1)
J = -np.eye(N) + C_orig
Sigma = np.eye(N)
Q_th = spl.solve_continuous_lyapunov(J,-Sigma)
# plots
pp.figure()
pp.imshow(Q_sim)
pp.colorbar()
pp.xlabel('target ROI')
pp.ylabel('source ROI')
pp.title('covariance matrix (functional connectivity)')
pp.figure()
pp.plot([0,Q_th.max()],[0,Q_th.max()],'--k')
pp.plot(Q_sim,Q_th,'.b')
pp.xlabel('simulated covariances')
pp.ylabel('theoretical covariances')
pp.show()
# -
# ## Connectivity estimation
# +
# Lyapunov optimization
mou_est = MOU()
if True:
# estimate of weights without knowledge of the topology of existing weights in C_orig
# regularization may be helpful here to "push" small weights to zero here
mou_est.fit(ts_sim, i_tau_opt=1, regul_C=1.)
else:
# estimate of weights knowing the topology of existing weights in C_orig
mou_est.fit(ts_sim, i_tau_opt=1, mask_C=C_orig>0)
C_est = mou_est.get_C()
# plots
pp.figure()
pp.imshow(C_orig,vmin=0)
pp.colorbar()
pp.xlabel('target ROI')
pp.ylabel('source ROI')
pp.title('original connectivity')
pp.figure()
pp.imshow(C_est,vmin=0)
pp.colorbar()
pp.xlabel('target ROI')
pp.ylabel('source ROI')
pp.title('estimated connectivity')
pp.figure()
pp.plot([0,C_orig.max()],[0,C_orig.max()],'--k')
pp.plot(C_est,C_orig,'xr')
pp.xlabel('estimated connectivity')
pp.ylabel('original connectivity')
pp.show()
# +
# moments method
mou_est2 = MOU()
mou_est2.fit(ts_sim,method='moments')
C_est2 = mou_est2.get_C()
# plots
pp.figure()
pp.imshow(C_orig,vmin=0)
pp.colorbar()
pp.xlabel('target ROI')
pp.ylabel('source ROI')
pp.title('original connectivity')
pp.figure()
pp.imshow(C_est2,vmin=0)
pp.colorbar()
pp.xlabel('target ROI')
pp.ylabel('source ROI')
pp.title('estimated connectivity')
pp.figure()
pp.plot([0,C_orig.max()],[0,C_orig.max()],'--k')
pp.plot(C_est2,C_orig,'xr')
pp.xlabel('estimated connectivity')
pp.ylabel('original connectivity')
pp.show()
# +
print('model fit for Lyapunov:',mou_est.d_fit['correlation'])
print('model fit for moments:',mou_est2.d_fit['correlation'])
print('C_orig fit for Lyapunov',stt.pearsonr(C_orig.flatten(),C_est.flatten()))
print('C_orig fit for moments',stt.pearsonr(C_orig.flatten(),C_est2.flatten()))
print('C_orig fit for moments + positive constraints',stt.pearsonr(C_orig.flatten(),np.maximum(C_est2,0).flatten()))
min_weight = min(C_est.min(),C_est2.min())
max_weight = max(C_est.max(),C_est2.max())
bins = np.linspace(min_weight,max_weight,40)
pp.figure()
pp.subplot(211)
pp.hist(C_est[C_orig>0], bins=bins, histtype='step', color='g')
pp.hist(C_est[C_orig==0], bins=bins, histtype='step', color='k')
pp.ylabel('distributions estimates')
pp.title('green = true; black = false')
pp.subplot(212)
pp.hist(C_est2[C_orig>0], bins=bins, histtype='step', color='g')
pp.hist(C_est2[C_orig==0], bins=bins, histtype='step', color='k')
pp.ylabel('distributions estimates')
pp.figure()
pp.subplot(211)
pp.hist(C_est[C_orig>0], bins=bins, histtype='step', cumulative=True, density=True, color='g')
pp.hist(C_est[C_orig==0], bins=bins, histtype='step', cumulative=True, density=True, color='k')
pp.ylabel('cumulative density')
pp.title('green = true; black = false')
pp.subplot(212)
pp.hist(C_est2[C_orig>0], bins=bins, histtype='step', cumulative=True, density=True, color='g')
pp.hist(C_est2[C_orig==0], bins=bins, histtype='step', cumulative=True, density=True, color='k')
pp.ylabel('cumulative density')
pp.figure()
pp.plot([0,C_orig.max()],[0,C_orig.max()],'--k')
pp.plot(C_est,C_est2,'xr')
pp.xlabel('LO estimate')
pp.ylabel('moment estimate')
pp.show()
| examples/MOU_Simulation_Estimation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Django Shell-Plus
# language: python
# name: python3
# ---
# # Merge land cover for publication figure
# This notebook merges the high-resolution LISA land cover with the lower-resolution CORINE land cover to create a land cover raster that covers a large extent to depict the surrounding landscape around the study area in a publication figure. <br />
#
# The 1.07 version of [basemap](https://github.com/matplotlib/basemap) is not compatible with the 0.36 version of [rasterio](https://github.com/mapbox/rasterio), which is required to merge the rasters. Therefore, this notebook must be run in a different environment than the other notebooks.
# #### Import statements
from geo.models import Raster
import rasterio
from rasterio.merge import merge
import os.path
# #### Load input raster objects
landcover_study_area = Raster.objects.get(name='landcover study area')
landcover_background = Raster.objects.get(name='landcover corine mapped lisa')
# #### Merge land cover
# s1 = rasterio.open(landcover_study_area.filepath, mode='r')
# s2 = rasterio.open(landcover_background.filepath, mode='r')
# landcover, transform = merge(sources=[s1, s2], res=(5., 5.), nodata=255)
# #### Write merged raster to disk
output_filepath = os.path.join(os.path.dirname(landcover_background.filepath), "landcover_merged_publication.tif")
with rasterio.open(path=output_filepath, mode='w', driver='GTiff',
width=landcover.shape[2], height=landcover.shape[1], count=landcover.shape[0],
crs="+init=epsg:31254", transform=transform, dtype=landcover.dtype, nodata=int(s1.nodata)) as dest:
dest.write(landcover[0], 1)
# #### Close input raster files
s1.close()
s2.close()
# #### Save new raster object to database
Raster(name='landcover merged publication', filepath=output_filepath).save()
| utilities/Merge landcover for publication figure.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/BDML-Workshop/DataFrames/blob/main/Spark_DataFrames_AutoEval_Exercise.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="pimQRInyvEkk"
# # Spark DataFrames Auto eval Exercise
# + [markdown] id="PMjoEy6HvEkl"
# Let's get some quick practice with your new Spark DataFrame skills, you will be asked some basic questions about some stock market data, in this case Walmart Stock from the years 2012-2017. This exercise will just ask a bunch of questions, unlike the future machine learning exercises, which will be a little looser and be in the form of "Consulting Projects", but more on that later!
#
# For now, just answer the questions and complete the tasks below.
# + [markdown] id="cLhVMdMfvEkl"
# #### Use the walmart_stock.csv file to Answer and complete the tasks below!
# + [markdown] id="etJ3HCySvEkl"
# #### Start a simple Spark Session in Google Collab
# + jupyter={"outputs_hidden": true} id="KxCsvEu2vEkl"
# + [markdown] id="y1iRsEChwJtK"
# Get the WalMart CSV from https://github.com/BDML-Workshop/DataFrames to the Collab session
# + [markdown] id="higLzv4LwXPh"
#
# + [markdown] id="IyNrUoelvEkl"
# #### Load the Walmart Stock CSV File, have Spark infer the data types.
# + jupyter={"outputs_hidden": true} id="XqYq7R2svEkl"
# + [markdown] id="W1Ku7LYhvEkl"
# #### What are the column names?
# + jupyter={"outputs_hidden": false} id="4c534Qv_vEkl" outputId="b6564159-b0f4-4034-f563-4adfd0711633"
# + [markdown] id="Z0mPXdlAvEkn"
# #### What does the Schema look like?
# + jupyter={"outputs_hidden": false} id="qQAev3sSvEkn" outputId="f618316e-3431-4d95-df11-d4db299e43cd"
# + [markdown] id="oQmSGfJOvEkn"
# #### Print out the first 5 columns.
# + jupyter={"outputs_hidden": false} id="oJGOOnsovEkn" outputId="fcf34275-1a53-4b98-90d5-c7adea54d369"
# + [markdown] id="K9Yu9ou3vEkn"
# #### Use describe() to learn about the DataFrame.
# + jupyter={"outputs_hidden": false} id="PxpbWaiqvEkn" outputId="5eac3b23-5528-456c-a42b-435db091af7f"
# + [markdown] id="9cDTbKOgvEkn"
# ## Bonus Question!
# #### There are too many decimal places for mean and stddev in the describe() dataframe. Format the numbers to just show up to two decimal places. Pay careful attention to the datatypes that .describe() returns, we didn't cover how to do this exact formatting, but we covered something very similar. [Check this link for a hint](http://spark.apache.org/docs/latest/api/python/pyspark.sql.html#pyspark.sql.Column.cast)
#
# If you get stuck on this, don't worry, just view the solutions.
# + jupyter={"outputs_hidden": false} id="bRtwtd-QvEkn" outputId="2501aa36-5ede-4601-ec90-99290414a42a"
# + jupyter={"outputs_hidden": true} id="kYVDZEOIvEkn"
# + jupyter={"outputs_hidden": false} id="HyBaxRsnvEkn" outputId="1e7b2ee2-8021-4997-a3df-890b94bb18f8"
# + [markdown] id="VuF7KRKXvEkn"
# #### Create a new dataframe with a column called HV Ratio that is the ratio of the High Price versus volume of stock traded for a day.
# + jupyter={"outputs_hidden": false} id="900za-TQvEkn" outputId="5cb0d7d5-535e-47b4-ac52-cfd0e4187bda"
# + [markdown] id="EfUBTrYyvEkn"
# #### What day had the Peak High in Price?
# + jupyter={"outputs_hidden": false} id="VMjdNUqsvEkn" outputId="20cb48c7-5601-4387-e618-acd219635bd3"
# + [markdown] id="XwKyr0KUvEko"
# #### What is the mean of the Close column?
# + jupyter={"outputs_hidden": false} id="6u8_I69evEko" outputId="fd624474-6738-4a2e-8b13-5afb62f0135a"
# + [markdown] id="AVLr2gkzvEko"
# #### What is the max and min of the Volume column?
# + jupyter={"outputs_hidden": true} id="FhlZFGMHvEko"
# + jupyter={"outputs_hidden": false} id="pybD6r8WvEko" outputId="1bda7bcf-a631-45ea-bf79-427e788a3c5b"
# + [markdown] id="0zpxjFZmvEko"
# #### How many days was the Close lower than 60 dollars?
# + jupyter={"outputs_hidden": false} id="fGIV315UvEko" outputId="4177dd54-c860-4d80-a523-3bc7617459ed"
# + [markdown] id="KtC2LmHwvEko"
# #### What percentage of the time was the High greater than 80 dollars ?
# #### In other words, (Number of Days High>80)/(Total Days in the dataset)
# + jupyter={"outputs_hidden": false} id="HtPQUusPvEko" outputId="e4b52467-6e85-4a34-d91d-ab0fd6e5428f"
# + [markdown] id="MQlThS50vEko"
# #### What is the Pearson correlation between High and Volume?
# #### [Hint](http://spark.apache.org/docs/latest/api/python/pyspark.sql.html#pyspark.sql.DataFrameStatFunctions.corr)
# + jupyter={"outputs_hidden": false} id="42I9zmVwvEko" outputId="f0f3de81-d2e5-4d23-adb1-266c5fac1b52"
# + [markdown] id="NimDsaRpvEkp"
# #### What is the max High per year?
# + jupyter={"outputs_hidden": false} id="vsVZNrP8vEkp" outputId="da8ded15-158e-45ec-b232-d8ecf934d2aa"
# + [markdown] id="hEq5zwFevEkp"
# #### What is the average Close for each Calendar Month?
# #### In other words, across all the years, what is the average Close price for Jan,Feb, Mar, etc... Your result will have a value for each of these months.
# + jupyter={"outputs_hidden": false} id="CyiDFza3vEkp" outputId="7da82b8b-1dd3-46c4-f070-840fa616ef07"
# + [markdown] id="caDPcklnvEkp"
# # Great Job!
| Spark_DataFrames_AutoEval_Exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# # Welcome to jupyter notebooks!
# ### Congratulations, the hardest step is always the first one. This exercise is designed to help you get personal with the format of jupyter notebooks, as well as learn how data is accessed and manipulated in python.
#
name = "Liz" #type your name before the pound sign, make sure you put it in quotes!
age = "135" #type your age before the pound sign, make sure you put it in quotes! More on this later.
address = "462C Link Hall" #type your address before the pound sign, make sure you put it in quotes!
story = "My name is " + name + ". I am " + age + " years old. I am a data scientist! Find me at " + address + "."
print(story)
# ## Variable types:
# Variables are names in your python environment where you store values. A variable can be a number, a character string, or a more complex data structure (such as a pandas spatial dataframe...more later).
#
# At the most basic level, variables representing single values can either be numbers or strings.
#
# Numeric values can be stored as intergers, floating point numbers, or complex numbers.
#
# Strings represent strings of characters, like writing.
#
# We find out what type of variable we're dealing with by using the type() function.
#
# More info here: https://mindmajix.com/python-variable-types
from IPython.display import Image
Image("https://cdn.mindmajix.com/blog/images/Screenshot_12-460x385.png")
print(type(name))
print(type(age))
print(type(address))
print(type(story))
#We can convert numeric strings to numeric formats using either float() or int()
age = float(age)
print(age)
print(type(age))
age=int(age)
print(age)
print(type(age))
#But we cannot input a numeric value into a string:
story = "My name is " + name + ". I am " + age + " years old. Find me at " + address + "."
print(story)
# ## TASK 1:
# Edit the code in the above cell so that it will print your story without adding any additional lines of code.
#Type story here
story = "My name is " + name + ". I am " + str(age) + " years old. Find me at " + address + "."
print(story)
# ## Mathematical operators in python:
# We're learning Python because it is a really powerful calculator. Below is a summary of common mathematical operators in Python.
Image("https://i0.wp.com/makemeanalyst.com/wp-content/uploads/2017/06/Python-Operators.png")
Image("https://i0.wp.com/makemeanalyst.com/wp-content/uploads/2017/06/Relational-Operators-in-Python.png")
Image("https://i0.wp.com/makemeanalyst.com/wp-content/uploads/2017/06/Bitwise-Operators_python.png")
Image("https://i1.wp.com/makemeanalyst.com/wp-content/uploads/2017/06/Assignment-Operator.png")
# We can apply mathematical operators to variables by writing equations, just like we do with a calculator.
#
# ## TASK 2:
# Using your "age" variable. how old will you be in five years (call this variable "a")? How many months have you been alive (we're going to estimate this, based on the fact that there are 12 months in a year, call this variable "b")? Are you older than 100 (call this variable "c")?
# +
#first, convert "age" back to a numeric (float) variable
age = float(age)
#Complete your equations in python syntax below. Remember to use "age" as a variable.
a = age + 5
b = age * 12
c = age >= 100
print(a,b,c)
# -
# ## Functions
# Often in life, we want to repeat the same sequence of operations on multiple values. To do this, we use functions. Functions are blocks of code that will repeat the same task over and over again (return some outputs) on different variables (inputs). https://docs.python.org/3/tutorial/controlflow.html#defining-functions
# +
#For example, this function squares the input
def fun1 (x):
return x*x # note the indentation
fun1(2)
# what happens if you delete the indentation?
# +
#Functions can have more than one input:
def fun2 (x, y):
return x * y
fun2(2,3) #experiment with different values
# -
# ### TASK 3:
#
# To simplify your future restaurant interactions, write a function that calculates a 15% tip and adds it to any check amount. Name your function tip15. How much will you owe in total on a $42.75 check?
# +
#write tip15 function here
def tip15 (check):
# check is the bill amount
return check * 1.15
tip15(42.75)
# -
# ### TASK 4:
#
# To customize your future restaurant interactions to future exemplary service, write a function that calculates a variable tip and adds it to any check amount. Name your function tip. How much will you owe in total if you're tipping 25% on a $42.75 check?
# +
#write tip function here
def tip15 (check, rate):
# check is the bill amount
# rate is the tipping rate expressed as a decimal, for example a 15% tip is .15
return check * (1 + rate)
tip15(42.75, .25)
# -
# ## Lists
# Using base python, you can store multiple variables as a list by wrapping them in square brackets. A list can accept any kind of variable: float, integer, or string:
names = ["Homer", "Marge", "Bart", "Maggie", "Lisa"]
ages = [40,39,11,1,9]
print(names)
print(ages)
type(names)
type(ages)
# We can make lists of lists, which are more complex data objects:
names_and_ages = [names, ages]
print(names_and_ages)
type(names_and_ages)
# We can extract individual values from the list using **indexing**, which in python starts with zero. We'd use our python mathematical operator on this single variable the same way we would use a calculator. For example, the age of the first person in the list is:
first = ages[0]
print(first)
# And we can ask: How old will the first numeric element on our list be in five years?
ages[0]+5
# ### TASK 5:
# Find the name and age of the third person in your list
# type answer here
print(names[2]+ ' is ' + str(ages[2]) + " years old.")
# ### TASK 6:
# Add an extra person to the list, named Grandpa, who is 67 years old. *Hint: Google "add values to a list python" if you're not sure what to do!!!!
# type answer here
names.append("Grandpa")
ages.append(67)
print(names_and_ages)
# ### TASK 7:
# Maggie just had a birthday! Change her age accordingly in the list.
# type answer here
names_and_ages[1][3] = names_and_ages[1][3]+1
print(names_and_ages)
# But because lists can be either string or numeric, we cannot apply a mathematical operator to a list. For example, how old will everyone be in five years?
ages + 5
#this returns an error. But hey! Errors are opportunities! Google your error message, see if you can find a workaround!
# ## For loops
# One way that we can run an entire operation on a list is to use a for loop. A for loop iterates through every element of a sequence, and completes a task or a set of tasks. Example:
#Measure some strings
words = ['cat', 'window', 'defenestrate']
for w in words:
print(w, len(w))
# A very handy function that folks use a lot when write for loops is the range function. range() creates a range object, which generates list of intergers for each element in a series.
#
print(len(ages))
# What does len() do?
print(range(len(ages)))
for y in range(len(ages)):
print(ages[y])
# ### TASK 8:
# Use the range function in a for loop to add 1 year to everyone's age in the names_and_ages list and print the result to the screen.
# +
# type your for loop here
for y in range(len(ages)):
ages[y] = ages[y] + 1
print(names_and_ages)
# -
# ## if/else statements
# Another thing that is commonly used in for loops is an if/else statement. This allows us to select certain attributes that we want to use to treat elements that we're iterating over differently.
for y in range(len(ages)):
if ages[y]>=18:
print(names[y] + " is an adult")
elif ages[y] >= 10:
print(names[y]+" is an adolescent")
else:
print(names[y] + " is a child")
# The above statement reads: if the age of the person is greater than or equal to 18, the person is an adult, or else if the age of the person is greater than or equal to 10, the person is an adolescent, or else the person is a child.
#
# Pay attention to the use of indentation in the above statement!
# ### TASK 9:
# Using if, elif, and/or else statements, write a for loop that adds 1 year to everyone's life except Maggie...since we already gave her an extra birthday.
# +
# Write for loop here
for y in range(len(ages)):
if names[y] != "Maggie":
ages[y] = ages[y] + 1
print(names_and_ages)
# -
# ## numpy arrays and pandas dataframes
# For loops can take a lot of time on large datasets. To deal with this issue, we can rely on two other packages for data structures, called numpy and pandas.
#
# *Packages* are collections of *modules*. *Modules* contain executables, functions, and object types that someone else has already written for you.
#
# Numpy arrays are standard gridded data formats (lists, matrices, and multi-dimensional arrays) where all items are of one data type, and have implicit indexing. Pandas data frames allow for mixed data types to be stored in different columns, and have explicit indicing. Pandas data frames work just like excel spreadsheets. Check out [https://www.earthdatascience.org/courses/intro-to-earth-data-science/scientific-data-structures-python/numpy-arrays/] and [https://www.earthdatascience.org/courses/intro-to-earth-data-science/scientific-data-structures-python/pandas-dataframes/] to learn more. Either numpy arrays or pandas data frames allow mathematical operators to be applied to all numeric elements.
#
# We're going to start off by loading the numpy and pandas packages. We're going to short their names to "np" and "pd", just so we don't have to type as many characters. These packages should have been downloaded automatically in Anaconda, but if you can't load them, raise your hand and share you're screen. We'll talk about installing packages and everyone will be all the wiser.
#
# Note: if importing python packages is new to you, please check out [https://www.earthdatascience.org/courses/intro-to-earth-data-science/python-code-fundamentals/use-python-packages/] to learn more.
import numpy as np #import the numpy package, and call it np
ages = np.array(ages)
ages + 5
#Wait! Did you get an error? Read it carefull...and if it doesn't make sense, try Googling it.
import pandas as pd #import the pandas pacakge, and call it pd
names_and_ages = pd.DataFrame({'names':names,'ages':ages})
print(names_and_ages)
type(names_and_ages)
#How does the "type" of variable change?
# The cell bellow shows how to add a new column to a pandas data frame called 'ages_5yr', and how to calculate values for the new column:
names_and_ages['ages_5yr']=names_and_ages.ages + 5
print(names_and_ages)
# There are three(?) ways to select a column in a pandas dataframe:
#
# *? = probably many more!
print(names_and_ages['names']) #index by column name
print(names_and_ages.names) #call column name as an object attribute
print(names_and_ages.iloc[:,0]) #use pandas built-in indexing function
# The last option, iloc, is a pandas function. The first slot (before the comma) tells us which *rows* to pick. The second slot (after the comma) tells us which *columns* to pick. To pick find the value "ages_5yrs" of the first row, for example, we could use iloc like this:
print(names_and_ages.iloc[0,2])
# ### TASK 10:
# Create a new column in names_and_ages called "age_months", and populate it with each indiviuals age in months (approximately age in years times 12).
#Complete task ten in the space below
names_and_ages['age_months'] = names_and_ages.ages*12
names_and_ages
# ### TASK 11:
# Create a new column in names_and_ages called "maturity", and populate it with "adult", "adolescent", and "child", based on the age cutoffs mentioned above.
# +
# Create new column, fill it with empty values (np.NaN)
names_and_ages['maturity']=np.NaN
# Write for loop here
for i in range(names_and_ages.shape[0]):
if names_and_ages.ages[i]>= 18:
names_and_ages.maturity.iloc[i] = "adult"
elif names_and_ages.ages[i]>=10:
names_and_ages.maturity.iloc[i] = "adolescent"
else:
names_and_ages.maturity.iloc[i] = "child"
print(names_and_ages)
# -
# # Read in data.
# You will often be using python to process files or data stored locally on your computer or server. Here will learn how to use the package os to set a path to the files you want, called your working directory.
import os
os.listdir()
# This .csv file is a record of daily discharge from a USGS stream gage near campus for the past year. It contains two columns, "Date" and "Discharge". It was downloaded with some modifications from [https://maps.waterdata.usgs.gov/mapper/index.html]. This is a great source of data for surface water measurements!
#
# Now that you've got the filepath set up so that python can see your stream discharge data, let's open it as a pandas data frame using the pandas.read_csv() function. This is similar to what we would do opening new data in Excel
stream = pd.read_csv('USGS_04240105_SYRACUSE_NY.csv', infer_datetime_format=True)
print(type(stream))
#the "head" function is built into a pandas.DataFrame object, and it allows you to specify the number of rows you want to see
stream.head(5)
# ##### Here, we see the date (as "%m/%d/%Y" format) and the mean daily discharge in cubic feet/second measured by the streamgage for each day.
#
# ## Check documentation on an object
# The pandas dataframe is an object that is associated with many attributes and functions. How can we tell what other people have enabled their modules to do, and learn how to use them?
#
# Remember, we just read in our data (USGS_04240105_SYRACUSE_NY.csv") as a 'pandas.core.frame.DataFrame'. What does that mean?
#We can ask a question:
# ?stream
#We can ask a deeper queston:
??stream
#This gives you full documentation
#Or we can use the 'dot' 'tab' magic.
stream.
#Don't click Enter!!! Click 'Tab'. What happens?
#We can use 'dot' 'tab' and question marks together!
# ?stream.min
#
# ### TASK 12: use pandas.DataFrame built in functions to determine the minimum, maximum, and mean daily discharge over the record for the station.
#use pandas.DataFrame functions to assign appropriate values to the variables here.
min_dis = stream.Discharge.min()
max_dis = stream.Discharge.max()
mean_dis = stream.Discharge.mean()
print(min_dis, max_dis, mean_dis)
# ### TASK 13: create a new column called "Discharge_total" where cumulative daily discharge is calculated (hint: there are 24 x 60 x 60 seconds in a day)
#enter calculations here
stream['Discharge_total'] = stream.Discharge*24*60*60
stream.head()
# ## EXTRA CREDIT: Use indexing to determine the DATE on which the maximum daily discharge occured.
# (Hint! check out ?stream.iloc and ?stream.argmax, then try Googling "Find row where value of column is maximum in pandas DataFrame" or something similar)
#
# (Still stuck? Enter the last thing you tried in this box, and then post your unsolved code on the Questions and Answers page on blackboard).
#Type your answer here, see if you can get it in one line of code!
stream.Date.iloc[stream['Discharge'].idxmax()]
# # You're almost done for today! Last steps:
# ## * save your .ipynb
# ## * download it as .HTML
# ## * upload .HTML to today's assignment on Blackboard
#
# ## If this was brand new to you, please take time to work through Sections 4,6, and 7 in [https://www.earthdatascience.org/courses/intro-to-earth-data-science/]
| CodeSprints/Python101_21_Solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DevNet NETCONF/YANG on Nexus Learning Lab Walkthrough
#
# ## Hands-On Practice Environment for the DevNet Learning Lab Exercises
#
# ---
# ### Overview
#
# This Notebook provides a place to enter and test the commands found in the [DevNet NETCONF/YANG for Nexus Learning Lab](https://developer.cisco.com/learning/tracks/nxos-programmability/netconf-yang-nexus/yang_devnet-format_part1/step/1 "DevNet NETCONF/YANG for Nexus Learning Lab").
#
# You may use the NX-OS switch to run the lab exercises. The device connection information below connects to the [DevNet Reservable Open NX-OS Sandbox](https://devnetsandbox.cisco.com/RM/Diagram/Index/0e22761d-f813-415d-a557-24fa0e17ab50?diagramType=Topology "DevNet Reservable Open NX-OS Sandbox").
# - The use of this specific DevNet lab requires a reservation plus connectivity via VPN.
#
# To drive the strongest learning retention, try to enter the code/payloads for each task in the Learning Lab guide on your own, without copying and pasting the code samples from the Learning Lab guide.
# - Yes, this means you should become comfortable manyally building the NETCONF XML payloads themselves.
# - Try to build your skills to the point that you can determine what the XML body of a NETCONF request should look like, based on the target XPath or information in the Learning Lab scenarios.
# - The point of this exercise is not to memorize XML payloads or complex Python code. The point is to become familiar enough with the `ncclient` syntax and NETCONF XML payload format that you feel comfortable building the code responses on your own.
#
# Although all of the code to successully complete these exercises is available in the Learning Lab Guide, there is a [solution document available](solutions/nx_os_ncclient_devnet_lab_hands_on_solution.ipynb "DevNet NETCONF/YANG Sample Code") which provides quick access to sample code.
#
# ---
# ### Establish Imports & Constants to support connectivity to a Nexus switch, using the Python ncclient
# Import required Python modules
from ncclient import manager
from xmltodict import parse, unparse
from lxml.etree import fromstring
from pprint import pprint as pp
# Constant variable to store `ncclient` connection information
DEVICE = {
'host': '10.10.20.58',
'port': 830,
'username': 'admin',
'password': '<PASSWORD>',
'hostkey_verify': False,
'device_params': {
'name': 'nexus'
}
}
# ### Get NETCONF Capabilities and the Serial Number for Nexus Switch
#
# ---
# +
# Get NETCONF Capabilities
# Enter your commands here
# +
# Print NETCONF Capabilities
# Enter your commands here
# +
# Construct filter to get serial number
# Enter your commands here
# +
# Get the serial number
# Enter your commands here
# +
# Enter your commands here
# -
# ### Use NETCONF to Configure Interfaces
#
# ---
# +
# Create a NETCONF XML playload which will create and configure a loopback interface
# Enter your commands here
# -
# ** Optional **
# Strictly for testing, use of the lxml.etree.fromstring method, to create an XML document from a string
fromstring(config)
# +
# Send the NETCONF payload to create the loopback interface
# Enter your commands here
# +
# Display the NETCONF response
# Enter your commands here
# -
# ### Use NETCONF to Configure BGP
#
# ---
# +
# Create the NETCONF payload to configure BGP
# Enter your commands here
# Stictly for testing, use of the lxml.etree.fromstring method, to create an XML document from a string
fromstring(config)
# +
# Send the NETCONF payload to configure BGP
# Enter your commands here
# +
# Create a NETCONF payload to add a BGP prefix to the switch
# Enter your commands here
# ** Optional **
# Strictly for testing, use of the lxml.etree.fromstring method, to create an XML document from a string
# Enter your commands here
# +
# Send the NETCONF payload to implement the BGP prefix config
# Enter your commands here
# +
# Display the NETCONF response
# Enter your commands here
# +
# Create a NETCONF payload to get the BGP prefix list
# Enter your commands here
# ** Optional **
# Strictly for testing, use of the lxml.etree.fromstring method, to create an XML document from a string
# Enter your commands here
# +
# Send the NETCONF payload to get the serial number
# Enter your commands here
# +
# Create a NETCONF to delete a BGP prefix
# Add the following attribute to the <prefix-items> tag:
# xmlns:nc="urn:ietf:params:xml:ns:netconf:base:1.0" nc:operation="delete"
# This abbreviated version also works:
# operation="delete"
# Enter your commands here
# ** Optional **
# Strictly for testing, use of the lxml.etree.fromstring method, to create an XML document from a string
# Enter your commands here
# +
# Send NETCONF payload to remove the BGP prefix config
# Enter your commands here
# +
# Display the NETCONF response
# Enter your commands here
# -
# ---
# ---
# +
# Create an OpenConfig payload to get loopback interface configurations
# Enter your commands here
# ** Optional **
# Strictly for testing, use of the lxml.etree.fromstring method, to create an XML document from a string
# Enter your commands here
# +
# Send the payload to get interface details
# Enter your commands here
# +
# Create an OpenConfig NETCONF payload to add a loopback interface
# Enter your commands here
# ** Optional **
# Strictly for testing, use of the lxml.etree.fromstring method, to create an XML document from a string
# Enter your commands here
# +
# Send the NETCONF configuration payload
# Enter your commands here
# -
| resources/nexus/netconf/devnet_learning_lab_hands_on/nx_os_ncclient_devnet_lab_hands_on.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/NoCodeProgram/CodingTest/blob/main/sorting/heapSort.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="ZRv6MhTA-af9"
# Title : Heap Sort
#
# Chapter : sort
#
# Link :
#
# ChapterLink :
#
# 문제: 주어진 array를 heapSort 하여라
# + id="5fw6pVSD-YrI" colab={"base_uri": "https://localhost:8080/"} outputId="389a78b2-1098-4f4f-ae7d-1721467d25be"
from typing import List
import heapq
#in place memory sorting
def heapSort(nums:List[int])->List[int]:
#python does not have maxHeap. multiply by -1
nums = [-1*n for n in nums]
heapq.heapify(nums)
sorted = [0] * len(nums)
for i in range(len(nums)-1,-1,-1):
largest = -1 * heapq.heappop(nums)
sorted[i] = largest
return sorted
print(heapSort(nums=[3, 5, 7, 9, 4, 2]))
# + id="ERHYvbm7OxeZ"
| sorting/heapSort.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import sys
import os.path as op
basename = "/home/adrian/PhD/Tweet-Classification-Diabetes-Distress/"
path_utils = op.join(basename , "utils")
sys.path.insert(0, path_utils)
from sys_utils import load_library
from tweet_utils import tweet_vectorizer
from defines import Patterns, Emotions
from emotion_codes import UNICODE_EMOJI, EMOJI_TO_CATEGORY
from preprocess import Preprocess
prep = Preprocess()
data_path = "/home/adrian/PhD/Data/Tweets20210128/Tweets_per_months_personal_noJokes/matching-tweets_diab_noRT-noDupl_20210128_personal_noJokes_withFullText.parquet"
# -
# load data
data = pd.read_parquet(data_path)#.sample(n=40, random_state=33)
print(data.columns)
print("Total count:", data.shape[0])
data.head()
# +
def check_emoticonEmoji_in_tweet(tweet):
""" Check if in each tweet occurs either an emoji or emoticon """
tweet = prep.replace_hashtags_URL_USER(tweet, mode_URL="delete", mode_Mentions="delete")
tweet = prep.tokenize(tweet)
for word in tweet:
match_emoticon = Patterns.EMOTICONS_PATTERN.findall(word)
# if emoticon found
if match_emoticon:
if match_emoticon[0] is not ":" and match_emoticon[0] is not ")": return True
# if emoji found
if word in UNICODE_EMOJI:
emot_cat = EMOJI_TO_CATEGORY[UNICODE_EMOJI[word]]
if emot_cat != "": return True
return False
def check_emotKeywords(tweet):
tweet = prep.replace_hashtags_URL_USER(tweet, mode_URL="delete", mode_Mentions="delete")
tweet = tweet.replace("nerve pain", "")
tweet = tweet.replace("nerve damage", "")
tweet = tweet.replace("pain of neuropathy", "")
tweet = prep.tokenize(tweet)
tweet = prep.remove_punctuation(tweet)
tweet = prep.to_lowercase(tweet)
tweet = [word for word in tweet if word not in ["healthcare"]]
for word in tweet:
for emot in Emotions.emotions_full_list:
if emot == word: return True
return False
data["emotion"] = data.full_text.map(lambda tweet: check_emoticonEmoji_in_tweet(tweet) or check_emotKeywords(tweet))
emot_tweets = data[data["emotion"] == True]
print(emot_tweets.shape)
data[["full_text", "emotion"]].head(200)
# -
for i, row in data[["full_text", "emotion"]][0:200].iterrows():
print(row["full_text"])
print(row["emotion"])
print()
Emotions.emotions_full_list
emot_tweets.head()
emot_tweets.to_parquet("/home/adrian/PhD/Data/Tweets20210128/Tweets_per_months_personal_noJokes/matching-tweets_diab_noRT-noDupl_20210128_personal_noJokes_withFullText_emotional.parquet")
| Emotional tweets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + papermill={"duration": 5.605641, "end_time": "2021-03-16T14:44:11.004872", "exception": false, "start_time": "2021-03-16T14:44:05.399231", "status": "completed"} tags=[]
import os
import sys
sys.path.append('../input/pytorch-images-seresnet')
import os
import math
import time
import random
import shutil
from pathlib import Path
from contextlib import contextmanager
from collections import defaultdict, Counter
import scipy as sp
import numpy as np
import pandas as pd
from tqdm.auto import tqdm
from functools import partial
import cv2
from PIL import Image
from matplotlib import pyplot as plt
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.optim import Adam, SGD
import torchvision.models as models
from torch.nn.parameter import Parameter
from torch.utils.data import DataLoader, Dataset
import albumentations
from albumentations import *
from albumentations.pytorch import ToTensorV2
import timm
from torch.cuda.amp import autocast, GradScaler
import warnings
warnings.filterwarnings('ignore')
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 0.01503, "end_time": "2021-03-16T14:44:11.029839", "exception": false, "start_time": "2021-03-16T14:44:11.014809", "status": "completed"} tags=[]
BATCH_SIZE = 64
# BATCH_SIZE_b5 = 32
TEST_PATH = '../input/ranzcr-clip-catheter-line-classification/test'
# + papermill={"duration": 0.03996, "end_time": "2021-03-16T14:44:11.078014", "exception": false, "start_time": "2021-03-16T14:44:11.038054", "status": "completed"} tags=[]
test = pd.read_csv('../input/ranzcr-clip-catheter-line-classification/sample_submission.csv')
# + papermill={"duration": 0.019015, "end_time": "2021-03-16T14:44:11.105942", "exception": false, "start_time": "2021-03-16T14:44:11.086927", "status": "completed"} tags=[]
class TestDataset(Dataset):
def __init__(self, df, transform=None):
self.df = df
self.file_names = df['StudyInstanceUID'].values
self.transform = transform
def __len__(self):
return len(self.df)
def __getitem__(self, idx):
file_name = self.file_names[idx]
file_path = f'{TEST_PATH}/{file_name}.jpg'
image = cv2.imread(file_path, cv2.IMREAD_GRAYSCALE)
mask = image > 0
image = image[np.ix_(mask.any(1), mask.any(0))]
image = cv2.cvtColor(image, cv2.COLOR_GRAY2RGB)
# image = cv2.imread(file_path)
# image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
if self.transform:
augmented = self.transform(image=image)
image = augmented['image']
return image
# + papermill={"duration": 0.016995, "end_time": "2021-03-16T14:44:11.131310", "exception": false, "start_time": "2021-03-16T14:44:11.114315", "status": "completed"} tags=[]
def get_transforms(image_size=684):
return Compose([
Resize(image_size, image_size),
# RandomResizedCrop(image_size, image_size, scale=(0.85, 1.0)),
# HorizontalFlip(p=0.5),
Normalize(),
ToTensorV2(),
])
# + papermill={"duration": 0.029914, "end_time": "2021-03-16T14:44:11.169895", "exception": false, "start_time": "2021-03-16T14:44:11.139981", "status": "completed"} tags=[]
class ResNet200D(nn.Module):
def __init__(self, model_name='resnet200d_320'):
super().__init__()
self.model = timm.create_model(model_name, pretrained=False)
n_features = self.model.fc.in_features
self.model.global_pool = nn.Identity()
self.model.fc = nn.Identity()
self.pooling = nn.AdaptiveAvgPool2d(1)
self.fc = nn.Linear(n_features, 11)
def forward(self, x):
bs = x.size(0)
features = self.model(x)
pooled_features = self.pooling(features).view(bs, -1)
output = self.fc(pooled_features)
return output
class SeResNet152D(nn.Module):
def __init__(self, model_name='seresnet152d'):
super().__init__()
self.model = timm.create_model(model_name, pretrained=False)
n_features = self.model.fc.in_features
self.model.global_pool = nn.Identity()
self.model.fc = nn.Identity()
self.pooling = nn.AdaptiveAvgPool2d(1)
self.fc = nn.Linear(n_features, 11)
def forward(self, x):
bs = x.size(0)
features = self.model(x)
pooled_features = self.pooling(features).view(bs, -1)
output = self.fc(pooled_features)
return output
class RANZCRResNet200D(nn.Module):
def __init__(self, model_name='resnet200d', out_dim=11):
super().__init__()
self.model = timm.create_model(model_name, pretrained=False)
n_features = self.model.fc.in_features
self.model.global_pool = nn.Identity()
self.model.fc = nn.Identity()
self.pooling = nn.AdaptiveAvgPool2d(1)
self.fc = nn.Linear(n_features, out_dim)
def forward(self, x):
bs = x.size(0)
features = self.model(x)
pooled_features = self.pooling(features).view(bs, -1)
output = self.fc(pooled_features)
return output
class CustomEffNet(nn.Module):
def __init__(self, model_name='tf_efficientnet_b5_ns', out_dim=11):
super().__init__()
self.model = timm.create_model(model_name, pretrained=False)
n_features = self.model.classifier.in_features
self.model.global_pool = nn.Identity()
self.model.classifier = nn.Identity()
self.pooling = nn.AdaptiveAvgPool2d(1)
self.classifier = nn.Linear(n_features, 11)
def forward(self, x):
bs = x.size(0)
features = self.model(x)
pooled_features = self.pooling(features).view(bs, -1)
output = self.classifier(pooled_features)
return output
# + papermill={"duration": 0.021872, "end_time": "2021-03-16T14:44:11.200794", "exception": false, "start_time": "2021-03-16T14:44:11.178922", "status": "completed"} tags=[]
def inference(models, test_loader, device):
tk0 = tqdm(enumerate(test_loader), total=len(test_loader))
probs = []
for i, (images) in tk0:
images = images.to(device)
avg_preds = []
for model in models:
with torch.no_grad():
y_preds1 = model(images)
y_preds2 = model(images.flip(-1))
y_preds = (y_preds1.sigmoid().to('cpu').numpy() + y_preds2.sigmoid().to('cpu').numpy()) / 2
avg_preds.append(y_preds)
avg_preds = np.mean(avg_preds, axis=0)
probs.append(avg_preds)
probs = np.concatenate(probs)
return probs
def tta_inference(model, loader, tta_steps=5):
all_probs = []
for i, step in enumerate(range(tta_steps)):
probs = []
for step, (images) in tqdm(enumerate(loader), total=len(loader)):
images = images.to(device)
with torch.no_grad():
y_preds = model(images)
y_preds = y_preds.sigmoid().to('cpu').numpy()
probs.append(y_preds)
all_probs.append(np.concatenate(probs))
avg_probs = np.mean(all_probs, axis=0)
return avg_probs
# + papermill={"duration": 29.767921, "end_time": "2021-03-16T14:44:40.978397", "exception": false, "start_time": "2021-03-16T14:44:11.210476", "status": "completed"} tags=[]
models200d_320_tuned_LowestLoss = []
model = ResNet200D()
model.load_state_dict(torch.load('../input/stage3-res320-f2e1-9734/Stage3_resnet200d_320_fold_2_epoch_1_97.34.pth'))
model.eval() # loss 0.0936
model.to(device)
models200d_320_tuned_LowestLoss.append(model)
model = ResNet200D()
model.load_state_dict(torch.load('../input/stage3-resnet320-f3-96969738/Stage3_resnet200d_320_fold_3_epoch_2_97.38.pth'))
model.eval() # loss 0.0984
model.to(device)
models200d_320_tuned_LowestLoss.append(model)
models200d_320_tuned = []
model = ResNet200D()
model.load_state_dict(torch.load("../input/stage3-resnet200d-320-f0e2-9718/Stage3_resnet200d_320_fold_3_epoch_2_97.18.pth"))
model.eval() # loss 0.1034
model.to(device)
models200d_320_tuned.append(model)
# model = ResNet200D()
# model.load_state_dict(torch.load('../input/stage3-res320-f1e3-9726/Stage3_resnet200d_320_fold_1_epoch_3_97.26.pth'))
# model.eval() # loss 0.1092
# model.to(device)
# models200d_320_tuned.append(model)
model = ResNet200D()
model.load_state_dict(torch.load('../input/stage3-resnet320-f3-96969738/Stage3_resnet200d_320_fold_3_epoch_1_96.96000000000001.pth'))
model.eval() # loss 0.1031
model.to(device)
models200d_320_tuned.append(model)
models200d_320_general = []
model = ResNet200D()
model.load_state_dict(torch.load('../input/stage3-general-res320-f3-96719702/Stage3_resnet200d_320_fold_3_epoch_11_97.02.pth'))
model.eval() # loss 0.1087
model.to(device)
models200d_320_general.append(model)
# + papermill={"duration": 25.277625, "end_time": "2021-03-16T14:45:06.265236", "exception": false, "start_time": "2021-03-16T14:44:40.987611", "status": "completed"} tags=[]
models200D_2_tuned = []
model = RANZCRResNet200D()
model.load_state_dict(torch.load('../input/resent200d-all-folds-fine-tuned/Stage3_resnet200d_fold_0_epoch_3_96.47.pth'))
model.eval()
model.to(device)
models200D_2_tuned.append(model)
model = RANZCRResNet200D()
model.load_state_dict(torch.load('../input/resent200d-all-folds-fine-tuned/Stage3_resnet200d_fold_1_epoch_2_95.88.pth'))
model.eval()
model.to(device)
models200D_2_tuned.append(model)
model = RANZCRResNet200D()
model.load_state_dict(torch.load('../input/resent200d-all-folds-fine-tuned/Stage3_resnet200d_fold_2_epoch_2_96.16.pth'))
model.eval()
model.to(device)
models200D_2_tuned.append(model)
model = RANZCRResNet200D()
model.load_state_dict(torch.load('../input/resent200d-all-folds-fine-tuned/Stage3_resnet200d_fold_3_epoch_4_96.26.pth'))
model.eval()
model.to(device)
models200D_2_tuned.append(model)
model = RANZCRResNet200D()
model.load_state_dict(torch.load('../input/resent200d-all-folds-fine-tuned/Stage3_resnet200d_fold_4_epoch_4_96.61999999999999.pth'))
model.eval()
model.to(device)
models200D_2_tuned.append(model)
# + [markdown] papermill={"duration": 0.009241, "end_time": "2021-03-16T14:45:06.284560", "exception": false, "start_time": "2021-03-16T14:45:06.275319", "status": "completed"} tags=[]
# SeResNet152D
# + papermill={"duration": 4.92469, "end_time": "2021-03-16T14:45:11.218846", "exception": false, "start_time": "2021-03-16T14:45:06.294156", "status": "completed"} tags=[]
models152D = []
# model = SeResNet152D()
# model.load_state_dict(torch.load('../input/seresnet152d-cv9615/seresnet152d_320_CV96.15.pth')['model'])
# model.eval()
# model.to(device)
# models152D.append(model)
# # model152d_96_16 = []
# model = SeResNet152D()
# model.load_state_dict(torch.load('../input/stage3-seresnet-f0e2-9616/Stage3_seresnet152d_fold_0_epoch_2_96.16.pth'))
# model.eval()
# model.to(device)
# models152D.append(model)
model = SeResNet152D()
model.load_state_dict(torch.load('../input/stage3-seresnet-f4e2-9669/Stage3_seresnet152d_fold_4_epoch_2_96.69.pth'))
model.eval()
model.to(device)
models152D.append(model)
# + [markdown] papermill={"duration": 0.00927, "end_time": "2021-03-16T14:45:11.237717", "exception": false, "start_time": "2021-03-16T14:45:11.228447", "status": "completed"} tags=[]
# ## MAIN
# + papermill={"duration": 5251.975438, "end_time": "2021-03-16T16:12:43.223535", "exception": false, "start_time": "2021-03-16T14:45:11.248097", "status": "completed"} tags=[]
test_dataset_864 = TestDataset(test, transform=get_transforms(image_size=864))
test_loader_864 = DataLoader(test_dataset_864, batch_size=BATCH_SIZE, shuffle=False, num_workers=4 , pin_memory=True)
predictions200d_320_tuned_LowestLoss = inference(models200d_320_tuned_LowestLoss, test_loader_864, device)
predictions200d_320_tuned = inference(models200d_320_tuned, test_loader_864, device)
predictions200d_320_general = inference(models200d_320_general, test_loader_864, device)
predictions200d_2_tuned = inference(models200D_2_tuned, test_loader_864, device)
predictions152d = inference(models152D, test_loader_864, device)
predictions = (1.20 * predictions200d_320_tuned_LowestLoss + predictions200d_320_tuned +
0.70 * predictions200d_320_general + 0.75 * predictions200d_2_tuned + 0.75 * predictions152d) / 4.40
# + papermill={"duration": 1.052294, "end_time": "2021-03-16T16:12:44.288627", "exception": false, "start_time": "2021-03-16T16:12:43.236333", "status": "completed"} tags=[]
target_cols = test.iloc[:, 1:12].columns.tolist()
test[target_cols] = predictions
test[['StudyInstanceUID'] + target_cols].to_csv('submission.csv', index=False)
test.head()
| Ranzcr CLIP/Inference Notebooks/2-tuned-97-02-864-1-2-1-0-7-0-75-1-0-75.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# You can write up to 20GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
# -
# # Importing all required modules.
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.metrics import confusion_matrix,classification_report
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import SVC
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
import scipy
# ## Importing dataset.
# + jupyter={"outputs_hidden": true}
df = pd.read_csv('/kaggle/input/10000-amazon-products-dataset/Amazon_Products.csv')
# -
df.head()
# ## Checking for null values and repeated data in the dataset.
df.isnull().sum()
# It shows that it contains come null values and some unnamed values.
df[df.duplicated()].count()
# #### There's no duplicate values in the dataset.
df.columns
# # Plotting graph for all columns.
df.shape
sns.catplot(x="manufacturer", y="price", data=df.head(10),height=5, aspect=27/9,)
# #### Visualizing the range of products make by the manufacturer. 👆 (only for 10 rows)
sns.catplot(x="number_available_in_stock", y="number_of_reviews", data=df.head(10004),height=5, aspect=27/9,)
# # describing the given dataset
df.describe()
df.corr()
# ## Plotting relational graphs.
plt.plot(df.number_available_in_stock, df.number_of_reviews)
| Amazon Dataset/amazon-data-visualisations.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Ruby 2.4.1
# language: ruby
# name: ruby
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc" style="margin-top: 1em;"><ul class="toc-item"><li><span><a href="#Name" data-toc-modified-id="Name-1"><span class="toc-item-num">1 </span>Name</a></span></li><li><span><a href="#Summary" data-toc-modified-id="Summary-2"><span class="toc-item-num">2 </span>Summary</a></span></li><li><span><a href="#Installation" data-toc-modified-id="Installation-3"><span class="toc-item-num">3 </span>Installation</a></span></li><li><span><a href="#Usage" data-toc-modified-id="Usage-4"><span class="toc-item-num">4 </span>Usage</a></span></li><li><span><a href="#Uninstall" data-toc-modified-id="Uninstall-5"><span class="toc-item-num">5 </span>Uninstall</a></span></li><li><span><a href="#Development" data-toc-modified-id="Development-6"><span class="toc-item-num">6 </span>Development</a></span></li><li><span><a href="#Contributing" data-toc-modified-id="Contributing-7"><span class="toc-item-num">7 </span>Contributing</a></span></li><li><span><a href="#License" data-toc-modified-id="License-8"><span class="toc-item-num">8 </span>License</a></span></li><li><span><a href="#Code-of-Conduct" data-toc-modified-id="Code-of-Conduct-9"><span class="toc-item-num">9 </span>Code of Conduct</a></span></li></ul></div>
# -
# # Name
# nb_util
# # Summary
# nb_util allows you to generate latex(tex) format from your jupyter notebook(ipynb) format.
# This gem supplies to help when you use jupyter notebook.
#
# what can it help me?
#
# 1. Convert [my_help](https://github.com/daddygongon/my_help) to jupyter notebook(ipynb)
# 1. Combine multiple jupyter notebook into one jupyter note book
# 1. Extract data from jupyter notebook, then convert to the file format you extracted data.
# 1. To see jupyter notbook's contents
# 1. Convert jupyter notebook(ipynb) to latex(tex) format(thesis and handout(A4))
#
# # Installation
# Add this line to your application's Gemfile:
#
# ```ruby
# gem 'nb_util'
# ```
#
# And then execute:
#
# $ bundle
#
# Or install it yourself as:
#
# $ gem install nb_util
#
# To use this library:
#
# ```ruby
# require 'nb_util'
# ```
#
# On your local system, run the following in your command line:
#
# $ git clone <EMAIL>:EAGLE12/nb_util.git
# $ cd nb_util
# $ bundle update
#
# Run the following in your command line:
#
# $ bundle exec exe/nb_util
#
# ```ruby
# nb_util says hello, EAGLE !!
# Commands:
# nb_util combine [input file1] [input file2] [output filename] # combine file1 and file2
# nb_util getcode [filename] # save in ruby format
# nb_util help [COMMAND] # Describe available commands or one specific command
# nb_util iputs [filename] # display ipynb file contents
# nb_util ipynb2tex [filename] [option] # convert ipynb to tex's thiesis format
# nb_util red WORD [OPTION] # red words print.
# nb_util yaml2ipynb [input filename] # convert yaml to ipynb
# ```
#
# Have a good life with nb_util!
# # Usage
# ```
# $ nb_util
# ```
#
# ```ruby
# nb_util says hello, EAGLE !!
# Commands:
# nb_util combine [input file1] [input file2] [output filename] # combine file1 and file2
# nb_util getcode [filename] # save in ruby format
# nb_util help [COMMAND] # Describe available commands or one specific command
# nb_util iputs [filename] # display ipynb file contents
# nb_util ipynb2tex [filename] [option] # convert ipynb to tex's thiesis format
# nb_util red WORD [OPTION] # red words print.
# nb_util yaml2ipynb [input filename] # convert yaml to ipynb
# ```
# # Uninstall
# + [markdown] cell_style="center"
# ```
# $ gem uninstall nb_util
# ```
# And then
#
# You want to uninstall by filling number on it
#
# ```ruby
# Select gem to uninstall:
# 1. nb_util-0.3.4
# 2. nb_util-0.3.5
# 3. nb_util-0.3.6
# 4. nb_util-0.3.7
# 5. nb_util-0.3.8
# 6. nb_util-0.4.0
# 7. All versions
# > 7
# Successfully uninstalled nb_util-0.3.4
# Successfully uninstalled nb_util-0.3.5
# Successfully uninstalled nb_util-0.3.6
# Successfully uninstalled nb_util-0.3.7
# Successfully uninstalled nb_util-0.3.8
# Remove executables:
# nb_util
#
# in addition to the gem? [Yn] y
# Removing nb_util
# Successfully uninstalled nb_util-0.4.0
# ```
#
# -
# # Development
# After checking out the repo, run `bin/setup` to install dependencies. Then, run `rake spec` to run the tests. You can also run `bin/console` for an interactive prompt that will allow you to experiment.
#
# To install this gem onto your local machine, run `bundle exec rake install`.
# To release a new version, update the version number in `version.rb`, and then run `bundle exec rake release`, which will create a git tag for the version, push git commits and tags, and push the `.gem` file to [rubygems.org](https://rubygems.org/gems/nb_util).
# # Contributing
# Bug reports and pull requests are welcome on GitHub at https://github.com/EAGLE12/nb_util. This project is intended to be a safe, welcoming space for collaboration, and contributors are expected to adhere to the [Contributor Covenant](http://contributor-covenant.org) code of conduct.
#
# 1. Fork it (https://github.com/EAGLE12/nb_util/fork)
# 1. Create your feature branch (git checkout -b my-new-feature)
# 1. Commit your changes (git commit -am 'Added some feature')
# 1. Push to the branch (git push origin my-new-feature)
# 1. Create new Pull Request
# # License
# The gem is available as open source under the terms of the [MIT License](https://opensource.org/licenses/MIT).
# # Code of Conduct
# Everyone interacting in the NbUtil project’s codebases, issue trackers, chat rooms and mailing lists is expected to follow the [code of conduct](https://github.com/EAGLE12/nb_util/blob/master/CODE_OF_CONDUCT.md).
| README.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/LuisFelipeUrena/DS-Unit-2-Linear-Models/blob/master/Assignments/DS13_TL_BTR_LUIS_URENA_UNIT2_1_4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="V1cSKhNaKDy6" colab_type="text"
# Lambda School Data Science
#
# *Unit 2, Sprint 1, Module 4*
#
# ---
# + [markdown] colab_type="text" id="7IXUfiQ2UKj6"
# # Logistic Regression
#
#
# ## Assignment 🌯
#
# You'll use a [**dataset of 400+ burrito reviews**](https://srcole.github.io/100burritos/). How accurately can you predict whether a burrito is rated 'Great'?
#
# > We have developed a 10-dimensional system for rating the burritos in San Diego. ... Generate models for what makes a burrito great and investigate correlations in its dimensions.
#
# - [ ] Do train/validate/test split. Train on reviews from 2016 & earlier. Validate on 2017. Test on 2018 & later.
# - [ ] Begin with baselines for classification.
# - [ ] Use scikit-learn for logistic regression.
# - [ ] Get your model's validation accuracy. (Multiple times if you try multiple iterations.)
# - [ ] Get your model's test accuracy. (One time, at the end.)
# - [ ] Commit your notebook to your fork of the GitHub repo.
#
#
# ## Stretch Goals
#
# - [ ] Add your own stretch goal(s) !
# - [ ] Make exploratory visualizations.
# - [ ] Do one-hot encoding.
# - [ ] Do [feature scaling](https://scikit-learn.org/stable/modules/preprocessing.html).
# - [ ] Get and plot your coefficients.
# - [ ] Try [scikit-learn pipelines](https://scikit-learn.org/stable/modules/compose.html).
# + colab_type="code" id="o9eSnDYhUGD7" colab={}
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Linear-Models/master/data/'
# !pip install category_encoders==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# + id="7aWfL-9iKDzG" colab_type="code" colab={}
# Load data downloaded from https://srcole.github.io/100burritos/
import pandas as pd
df = pd.read_csv(DATA_PATH+'burritos/burritos.csv')
# + id="qopYVvQ2KDzK" colab_type="code" colab={}
# Derive binary classification target:
# We define a 'Great' burrito as having an
# overall rating of 4 or higher, on a 5 point scale.
# Drop unrated burritos.
df = df.dropna(subset=['overall'])
df['Great'] = df['overall'] >= 4
# + id="VFC1JoErKDzO" colab_type="code" colab={}
# Clean/combine the Burrito categories
df['Burrito'] = df['Burrito'].str.lower()
california = df['Burrito'].str.contains('california')
asada = df['Burrito'].str.contains('asada')
surf = df['Burrito'].str.contains('surf')
carnitas = df['Burrito'].str.contains('carnitas')
df.loc[california, 'Burrito'] = 'California'
df.loc[asada, 'Burrito'] = 'Asada'
df.loc[surf, 'Burrito'] = 'Surf & Turf'
df.loc[carnitas, 'Burrito'] = 'Carnitas'
df.loc[~california & ~asada & ~surf & ~carnitas, 'Burrito'] = 'Other'
# + id="21WchimMKDzR" colab_type="code" colab={}
# Drop some high cardinality categoricals
df = df.drop(columns=['Notes', 'Location', 'Reviewer', 'Address', 'URL', 'Neighborhood'])
# + id="GlJTFS6hKDzV" colab_type="code" colab={}
# Drop some columns to prevent "leakage"
df = df.drop(columns=['Rec', 'overall'])
# + id="enFdFV_YKDzY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 232} outputId="5eb02214-e5df-4842-cc5c-c1e29c4008e9"
df.head()
# + id="h5PhtoaNK8V2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 232} outputId="60e955a4-02c3-4a69-d98e-da62ecc75717"
df.tail()
# + id="Q5-VH_47LZVU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="a285dc18-82a2-484a-f261-a5f5c98ac1a0"
df.dtypes
# + id="QjkRfGLaNOVj" colab_type="code" colab={}
#changing the date feature from object to Datetime format
df['Date'] = pd.to_datetime(df['Date'])
# + id="1772iW4rNj0Q" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="46228be8-25c0-4b08-be60-32fe20f6d289"
df.dtypes
# + id="NKEr9LZ2Nlbm" colab_type="code" colab={}
# separating test train and validate data
train = df[df['Date'].dt.year <=2016]
test = df[df['Date'].dt.year >= 2018]
val = df[df['Date'].dt.year == 2017]
# + id="Fz1TbZsuN4s3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 218} outputId="3d2a8436-0893-4b02-e83c-034d0c7782ac"
#making sure the dates selected are there
val['Date']
# + id="pQN2XsgvOmbr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 672} outputId="adce29d5-f439-4456-afaf-bc78edada313"
#we are good here
test['Date']
# + id="AVh_tRLYOooJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 218} outputId="c4292c33-5170-483e-bf22-0ed58efe3ee1"
#everything is in order
train['Date']
# + id="p3GlxPC3OtHs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="35fcafa2-a3dd-4906-dc26-a1af4f293bc6"
BL_count = df['Great'].value_counts(normalize=True)
BL_count
# + id="6jiJzrYgc6Jg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0db23c16-2e07-4135-a06d-2b74001262ed"
Baseline = BL_count[0]
print('The Baseline for our model is:',Baseline)
# + id="dygMqevOSGIA" colab_type="code" colab={}
#Getting the target and features
target = 'Great'
features = ['Tortilla','Wrap','Fillings','Burrito']
#getting train subsets
X_train = train[features]
Y_train = train[target]
#getting validate subsets
X_val = val[features]
Y_val = val[target]
#getting test subsets
X_test = test[features]
Y_test = test[target]
# + id="7h_7G2ssTvk3" colab_type="code" colab={}
#The libraries
import category_encoders as ce
from sklearn.impute import SimpleImputer
from sklearn.linear_model import LogisticRegression
from sklearn.preprocessing import StandardScaler
# + id="1aoJcc_Xx1cM" colab_type="code" colab={}
# One Hot encoding to the categorical feature of the burrito type
encoder = ce.OneHotEncoder(use_cat_names=True)
X_train_encoded = encoder.fit_transform(X_train)
X_val_encoded = encoder.transform(X_val)
# + id="sIzVbO4IVGeA" colab_type="code" colab={}
# first let's use the simple imputer
imputer = SimpleImputer(strategy='mean')
X_train_imputed = imputer.fit_transform(X_train_encoded)
X_val_imputed = imputer.transform(X_val_encoded)
# + id="Ppzks1yZYZDK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="bfec67ed-99ee-4ed3-f3c9-71a6f01ee35d"
#now lets fit the Logistical Regression
log_reg = LogisticRegression(solver='lbfgs')
log_reg.fit(X_train_imputed,Y_train)
#Validation accuracy
print('Validation Accuracy:',log_reg.score(X_val_imputed,Y_val))
# + id="CpkNQR8iZAI0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="dab8f043-cf29-4a2e-8068-aa5cdd5f1a40"
#test accuracy
x_test_encoded = encoder.fit_transform(X_test)
X_test_imputed = imputer.fit_transform(x_test_encoded)
print('Test Accuracy:',log_reg.score(X_test_imputed,Y_test))
# + id="iPzN2YBvZ9xp" colab_type="code" colab={}
#scalar
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train_imputed)
X_val_scaled = scaler.fit_transform(X_val_imputed)
# + id="iLMNPX1x1L6Y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="28545444-d964-47ac-a96b-c1eef07baa24"
from sklearn.linear_model import LogisticRegressionCV
model = LogisticRegressionCV
model.fit(X_train_scaled, Y_train)
# + id="R3G4BODx1bAP" colab_type="code" colab={}
| Assignments/DS13_TL_BTR_LUIS_URENA_UNIT2_1_4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:nlp]
# language: python
# name: conda-env-nlp-py
# ---
# Importing the libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# Importing the dataset
dataset = pd.read_csv('Restaurant_Reviews.tsv', delimiter = '\t', quoting = 3)
#quoting=3 ignore double quotes
dataset
# Cleaning the texts
import re
import nltk
#nltk.download('stopwords')
from nltk.corpus import stopwords
from nltk.stem.porter import PorterStemmer
def rem_stop_word(review):
review_processed =[]
for word in review:
if word not in stopwords.words('english'):
review_processed.append(word)
return review_processed
review = re.sub('[^a-zA-Z]', ' ', dataset['Review'][0])
print(review)
review = review.lower() # change to lower case letters
print(review)
review = review.split() # convert to list of words
print(review)
review = rem_stop_word(review)
print(review)
ps = PorterStemmer()
review = [ps.stem(word) for word in review]
print(review)
review = re.sub('[^a-zA-Z]', ' ', dataset['Review'][0])
print('Remove special characters - \t',review)
review = review.lower() # change to lower case letters
print('To lower -\t\t\t',review)
review = review.split() # convert to list of words
print('Split - \t\t\t', review)
ps = PorterStemmer()
review = [ps.stem(word) for word in review if not word in stopwords.words('english')]
print('Stopword Removal and Stemming -\t', review)
review = ' '.join(review)
print('Join\t\t\t\t', review)
corpus = []
for i in range(0, len(dataset)):
review = re.sub('[^a-zA-Z]', ' ', dataset['Review'][i])
#Keep only the characters from a to z
#[^a-zA-Z] to specify what is required in the review
# ' ' to replace removed characters with space
review = review.lower() # change to lower case letters
review = review.split() # convert to list of words
ps = PorterStemmer()
review = [ps.stem(word) for word in review if not word in set(stopwords.words('english'))]
# remove the stopwords ie. irrelevant words in the sentence for sentiment analysis
review = ' '.join(review) # join the words separated by space
corpus.append(review)
corpus
# Creating the Bag of Words model
from sklearn.feature_extraction.text import CountVectorizer
cv = CountVectorizer(max_features = 1500) #most frequent 1500 words
X = cv.fit_transform(corpus).toarray()
y = dataset.iloc[:, 1].values
X.shape
y
# Splitting the dataset into the Training set and Test set
from sklearn.cross_validation import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.20, random_state = 0)
# Fitting Naive Bayes to the Training set
from sklearn.naive_bayes import GaussianNB
classifier = GaussianNB()
classifier.fit(X_train, y_train)
# Predicting the Test set results
y_pred = classifier.predict(X_test)
# Making the Confusion Matrix
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, y_pred)
cm
# ### Accuracy
(cm[0][0]+cm[1][1])/np.sum(cm)
| NLP_Bag_of_Words-Detailed.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Enrico-Call/RL-AKI/blob/modeling-notebook/Modeling.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Cb_YT0vk_rLr"
# <img src="https://github.com/AmsterdamUMC/AmsterdamUMCdb/blob/master/img/logo_amds.png?raw=1" alt="Logo" width=128px/>
#
# # VUmc Research Project - Reinforcement Learning for Sepsis Prevention
# # Data Extraction
#
# AmsterdamUMCdb version 1.0.2 March 2020
# Copyright © 2003-2022 Amsterdam UMC - Amsterdam Medical Data Science
# + [markdown] id="WP9iu5etAvuv"
# ## 1. Set Up Environment and Load Data
# + id="wxUQAhuAAXdo"
from google.colab import drive
import os
import pandas as pd
drive.mount('/content/drive', force_remount=True)
os.chdir('/content/drive/MyDrive/MLRFH')
state_space = pd.read_csv('.csv')
action_space = pd.read_csv('.csv')
# + [markdown] id="eME6SrFs_t3k"
# ## 2. Modeling
#
# Create a Q-Learning Agent to iterate over the dataset.
# + id="rG68gt6yBI70"
# # Transition Matrix
# # Counting number of unique clusters (now we know it, but for later if we add more might be useful)
# numbers = sorted(final['cluster'].unique())
# # Find how many times a state is followed by another
# groups = final.groupby(['cluster', 'next'])
# counts = {i[0]:(len(i[1]) if i[0][0] != i[0][1] else 0) for i in groups}
# # Build a matrix based on the counts just performed
# matrix = pd.DataFrame()
# for x in numbers:
# matrix[x] = pd.Series([counts.get((x,y), 0) for y in numbers], index=numbers)
# matrix_normed = matrix / matrix.sum(axis=0)
# matrix = matrix_normed.to_numpy()
# print(matrix)
# + id="g0BMU2M1HU1B"
# Check how many times a state is associated with a certain reward
# For now, it might make sense to model the reward of each state based on the highest number of occurrences of a certain reward
# rewards = final.groupby(['cluster', 'reward']).size()
# print(rewards)
rewards = final.groupby(['cluster'], sort=False)['reward'].max()
rewards.sort_index(inplace=True)
rewards = rewards.to_numpy()
# + id="L4beaKx5HX3j"
# rew = final.groupby(['cluster', 'action', 'next']).size()
# # rew.sort_index(inplace=True)
# print(rew)
# # rew = rew.to_numpy()
# + id="gnOrYn1OHa9N"
# alphas = range(0, 85, 5) # Learning Rate
# gamma = 0.99 # Discount Factor
# for alpha in alphas:
# alpha = alpha/100
# val = []
# # Q(s, a) matrix
# for i in range(10):
# Q = np.zeros((50, 4))
# from sklearn.model_selection import GroupShuffleSplit
# train_inds, test_inds = GroupShuffleSplit(test_size=.20, n_splits=2, random_state = 7).split(final, groups=final['admissionid'])
# train = final.iloc[train_inds[0]]
# test = final.iloc[test_inds[0]]
# iterations = 1000000
# train = train[['cluster', 'action', 'next', 'reward']].sample(n = iterations, replace = True)
# test = test[['cluster', 'action', 'next', 'reward']].sample(n = iterations, replace = True)
# i = 0
# for row in train.loc[:,['cluster', 'action', 'next', 'reward']].itertuples():
# index, curr, act, next, rew = row
# delta = rew+gamma*np.max(Q[int(next), :])- Q[int(curr), int(act)]
# Q[int(curr), int(act)] += alpha*delta
# i += 1
# # if the update is smaller than a certain threshold, stop the iteration
# # if delta < 1e-10: break
# p_optim = 0.9
# actions = np.argmax(Q, axis = 1)
# pi = np.full((50, 4), ((1-p_optim)/(50-1)))
# for i, j in enumerate(actions):
# pi[i,j] = p_optim
# Q_1 = np.zeros((50, 4))
# for row in test.loc[:,['cluster', 'action', 'next', 'reward']].itertuples():
# index, curr, act, next, rew = row
# Q_1[int(curr), int(act)] += 1
# Q_1 = Q_1/Q_1.sum(axis=1, keepdims=True)
# best_act_pi = np.argmax(Q, axis=1)
# best_act_test = np.argmax(Q_1, axis=1)
# val.append(np.count_nonzero(best_act_pi == best_act_test))
# val = np.array(val)
# print(alpha, val.mean())
# + [markdown] id="8a_cO30EIiRF"
# ### Parameter Tuning
# + id="emsTuJRuIeTu"
alphas = range(0, 85, 5) # Learning Rate
gamma = 0.99 # Discount Factor
for alpha in alphas:
alpha = alpha/100
Q = np.zeros((50, 4))
from sklearn.model_selection import GroupShuffleSplit
train_inds, test_inds = GroupShuffleSplit(test_size=.20, n_splits=2, random_state = 7).split(final, groups=final['patientid'])
train = final.iloc[train_inds[0]]
test = final.iloc[test_inds[0]]
iterations = 1000000
train = train[['cluster', 'action', 'next', 'reward']].sample(n = iterations, replace = True)
test = test[['cluster', 'action', 'next', 'reward']].sample(n = iterations, replace = True)
i = 0
for row in train.loc[:,['cluster', 'action', 'next', 'reward']].itertuples():
index, curr, act, next, rew = row
delta = rew+gamma*np.max(Q[int(next), :])- Q[int(curr), int(act)]
Q[int(curr), int(act)] += alpha*delta
i += 1
# if the update is smaller than a certain threshold, stop the iteration
# if delta < 1e-10: break
p_optim = 0.9
actions = np.argmax(Q, axis = 1)
pi = np.full((50, 4), ((1-p_optim)/(50-1)))
for i, j in enumerate(actions):
pi[i,j] = p_optim
Q_1 = np.zeros((50, 4))
for row in test.loc[:,['cluster', 'action', 'next', 'reward']].itertuples():
index, curr, act, next, rew = row
Q_1[int(curr), int(act)] += 1
Q_1 = Q_1/Q_1.sum(axis=1, keepdims=True)
best_act_pi = np.argmax(Q, axis=1)
best_act_test = np.argmax(Q_1, axis=1)
print(alpha, np.count_nonzero(best_act_pi == best_act_test))
# + [markdown] id="a_X0dzsMJzy5"
# ### Run Model
# + id="On6x2ZjRI9to"
from sklearn.model_selection import GroupShuffleSplit
train_inds, test_inds = GroupShuffleSplit(test_size=.20, n_splits=2, random_state = 7).split(final, groups=final['admissionid'])
train = final.iloc[train_inds[0]]
test = final.iloc[test_inds[0]]
train = train[['cluster', 'action', 'next', 'reward']].sample(n = iterations, replace = True)
test = test[['cluster', 'action', 'next', 'reward']].sample(n = iterations, replace = True)
alphas = 0.75 # Learning Rate
gamma = 0.99 # Discount Factor
avg_Q = []
avg_Q1 = []
for i in range(50):
Q = np.zeros((50, 4))
iterations = 1000000
i = 0
for row in train.loc[:,['cluster', 'action', 'next', 'reward']].itertuples():
index, curr, act, next, rew = row
delta = rew+gamma*np.max(Q[int(next), :])- Q[int(curr), int(act)]
Q[int(curr), int(act)] += alphas*delta
i += 1
# if the update is smaller than a certain threshold, stop the iteration
# if delta < 1e-10: break
p_optim = 0.9
actions = np.argmax(Q, axis = 1)
pi = np.full((50, 4), ((1-p_optim)/(50-1)))
for i, j in enumerate(actions):
pi[i,j] = p_optim
avg_Q.append(np.average(pi, axis=0))
# + id="kUIvbnZFJ5Fd"
avg_Q1 = []
for i in range(10):
Q_1 = np.zeros((50, 4))
for row in test.loc[:,['cluster', 'action', 'next', 'reward']].itertuples():
index, curr, act, next, rew = row
Q_1[int(curr), int(act)] += 1
Q_1 = Q_1/Q_1.sum(axis=1, keepdims=True)
best_act_pi = np.argmax(Q, axis=1)
best_act_test = np.argmax(Q_1, axis=1)
avg_Q1.append(np.average(Q_1, axis = 0))
# + id="QOp_40ojJ8yZ"
Q_plot = np.average(pi, axis=0)
Q_1_plot = np.average(Q_1, axis = 0)
Q_plot, Q_1_plot
# + id="GNETZ8HtJ_Xi"
np.count_nonzero(best_act_pi == best_act_test)
# + id="mni3k_qJKCYP"
labels = ['0', '1', '2', '3']
x = np.arange(len(labels)) # the label locations
width = 0.35 # the width of the bars
fig, ax = plt.subplots()
rects1 = ax.bar(x - width/2, Q_plot, width, label='Algorithm Policy')
rects2 = ax.bar(x + width/2, Q_1_plot, width, label='Test Set Actions')
# Add some text for labels, title and custom x-axis tick labels, etc.
ax.set_ylabel('Probability of Taking Action')
ax.set_title('Action')
ax.set_xticks([0, 1, 2, 3])
ax.legend()
fig.tight_layout()
plt.show()
| Modeling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import scipy.stats as sts
import numpy as np
from scipy.stats import linregress
import seaborn as sns
# Study data files
Lifestyle_data = "../Row-2-Group-Project/cdc_npao.csv"
Incidence_data = "../Row-2-Group-Project/heighest_lowest_Incidencerate_data.csv"
# Read CSV
raw_data = pd.read_csv(Lifestyle_data)
cancer_data = pd.read_csv(Incidence_data)
#Cleanup of LifeStyle Data
raw_data_df = pd.DataFrame(raw_data).rename(columns = {'YearStart':"Year", 'LocationDesc':"State", "Data_Value":"Percentage", "Class":"Category"})
del raw_data_df['YearEnd']
del raw_data_df['LocationAbbr']
del raw_data_df['Datasource']
del raw_data_df['Topic']
del raw_data_df['Data_Value_Unit']
del raw_data_df['Data_Value_Type']
del raw_data_df['Data_Value_Alt']
del raw_data_df['Data_Value_Footnote_Symbol']
del raw_data_df['Data_Value_Footnote']
del raw_data_df['Low_Confidence_Limit']
del raw_data_df['High_Confidence_Limit ']
del raw_data_df['Education']
del raw_data_df['Gender']
del raw_data_df['Income']
del raw_data_df['Race/Ethnicity']
del raw_data_df['GeoLocation']
del raw_data_df['ClassID']
del raw_data_df['TopicID']
del raw_data_df['Sample_Size']
del raw_data_df['Total']
del raw_data_df['Age(years)']
del raw_data_df['QuestionID']
del raw_data_df['DataValueTypeID']
del raw_data_df['LocationID']
del raw_data_df['StratificationCategoryId1']
del raw_data_df['StratificationCategory1']
del raw_data_df['Stratification1']
del raw_data_df['StratificationID1']
raw_data_df
# Merge our two data frames together
combined_data = pd.merge(raw_data_df, cancer_data, on="State")
combined_data=combined_data.dropna()
export_csv = cat_data.to_csv (r'../Row-2-Group-Project/combined_lifestyle_data.csv', header=True)
combined_data.head()
# +
#Top States Category Overall Averages
years_selected = combined_data["Year"].isin(['2012','2013','2014','2015','2016'])
filter_topstates_data = combined_data[years_selected]
grouped_topstates_data = filter_topstates_data.groupby(["Category",])
Avg_topstates_Percent = (grouped_topstates_data["Percentage"]).mean()
cat_topstates_data = pd.DataFrame({"Top States Overall Averages":Avg_topstates_Percent})
export_csv = cat_topstates_data.to_csv (r'../Row-2-Group-Project/cat_topstates_data.csv', header=True)
cat_topstates_data
# -
#plotting the bar graph - Nationwide
ax = cat_topstates_data.plot(kind='bar' ,figsize=(10,5), fontsize=13,width=0.75)
plt.legend(loc='best')
plt.title('Top States Overall Lifestyle Habits')
plt.xlabel('Lifestyle Habit')
plt.ylabel('Percentage')
plt.savefig('../Row-2-Group-Project/Top_States_Overall_Lifestyle_Bar_Chart.png',bbox_inches='tight')
# +
#Creating a dataframe to gather data for bar chart
years_selected = combined_data["Year"].isin(['2012','2013','2014','2015','2016'])
#get physical activity data
cat_physical_chosen= raw_data_df["Category"].isin(['Physical Activity'])
filter_physical_data = combined_data[years_selected&cat_physical_chosen]
grouped_physical_data = filter_physical_data.groupby(["State",])
Avg_physical_Percent = (grouped_physical_data["Percentage"]).mean()
#get obesity data
cat_obesity_chosen= raw_data_df["Category"].isin(['Obesity / Weight Status'])
filter_obesity_data = combined_data[years_selected&cat_obesity_chosen]
grouped_obesity_data = filter_obesity_data.groupby(["State",])
Avg_obesity_Percent = (grouped_obesity_data["Percentage"]).mean()
#get nutrition data
cat_nutrition_chosen= raw_data_df["Category"].isin(['Fruits and Vegetables'])
filter_nutrition_data = combined_data[years_selected&cat_nutrition_chosen]
grouped_nutrition_data = filter_nutrition_data.groupby(["State",])
Avg_nutrition_Percent = (grouped_nutrition_data["Percentage"]).mean()
#Setting up the dataframe
cat_data = pd.DataFrame({"Physical Activity":Avg_physical_Percent,
"Obesity / Weight Status":Avg_obesity_Percent,
"Fruits and Vegetables":Avg_nutrition_Percent})
export_csv = cat_data.to_csv (r'../Row-2-Group-Project/cat_data.csv', header=True)
cat_data.dropna()
# -
#plotting the bar graph
ax = cat_data.plot(kind='barh' ,figsize=(20,20), fontsize=13,width=0.75)
plt.legend(loc='best')
plt.title('Top States Lifestyle Habits')
plt.xlabel('Percentage')
plt.savefig('../Row-2-Group-Project/Top_States_Lifestyle_Bar_Chart.png',bbox_inches='tight')
# +
#Creating a dataframe to gather data to calculate correlations
Incidence_rate = combined_data.groupby(["State"])["Incidence Rate"].mean()
corr_cat_data = pd.DataFrame({"Physical Activity":Avg_physical_Percent,
"Obesity / Weight Status":Avg_obesity_Percent,
"Fruits and Vegetables" :Avg_nutrition_Percent,
"Incidence Rate": Incidence_rate})
export_csv = corr_cat_data.to_csv (r'../Row-2-Group-Project/corr_cat_data.csv', header=True)
corr_cat_data.dropna()
# +
#Top States Overall Correlations (Top States with data)
fig, ax = plt.subplots(figsize=(10,8))
akws = {"ha": 'left',"va": 'bottom'}
sns.heatmap(corr_cat_data.corr(method='pearson'), annot=False, annot_kws = akws,fmt='.2f',
cmap=plt.get_cmap('Blues'), cbar=True, ax=ax,linewidths = 0.5,)
ax.set_yticklabels(ax.get_yticklabels(), rotation="horizontal")
plt.title("Correlations between Lifestyle Habits and Incidence Rate")
plt.savefig('../Row-2-Group-Project/Correlation_Chart.png',bbox_inches='tight')
plt.show()
# +
#Scatter plot of Physical Activity V Incidence Rate
x_values = cat_data["Physical Activity"]
y_values = corr_cat_data["Incidence Rate"]
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(10,5),fontsize=15,color="red")
plt.title('Comparison of Physical Activity and Incidence Rate')
plt.xlabel('Physical Activity')
plt.ylabel('Incidence Rate')
plt.savefig('../Row-2-Group-Project/Physical_Activity_Incidence_Scatter_Chart.png')
plt.show()
# +
#Scatter plot of Obesity / Weight Status V Incidence Rate
x_values = cat_data["Obesity / Weight Status"]
y_values = corr_cat_data["Incidence Rate"]
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(10,5),fontsize=15,color="red")
plt.title('Comparison of Obesity/Weight and Incidence Rate')
plt.xlabel('Obesity / Weight Status')
plt.ylabel('Incidence Rate')
plt.savefig('../Row-2-Group-Project/Obesity_Incidence_Scatter_Chart.png')
plt.show()
# +
#Scatter plot of Fruits and Vegetables V Incidence Rate
x_values = cat_data["Fruits and Vegetables"]
y_values = corr_cat_data["Incidence Rate"]
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(10,5),fontsize=15,color="red")
plt.title('Comparison of Nutrition and Incidence Rate')
plt.xlabel('Fruits and Vegetables')
plt.ylabel('Incidence Rate')
plt.savefig('../Row-2-Group-Project/Nutrition_Incidence_Scatter_Chart.png')
plt.show()
# -
| Lifestyle Data Code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import scipy
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from scipy import stats
import warnings
warnings.filterwarnings('ignore')
pd.set_option('display.max_colwidth', -1)
pd.set_option('display.float_format', lambda x: '%.3f' % x)
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1000)
plt.style.use('classic')
# %matplotlib inline
s=12
# +
# sns.set(font_scale=2)
# sns.set_style("white")
fig = plt.figure(figsize=(12,4), dpi=300)
# Momentum
plt.errorbar(x=1.074863, y=4.0, xerr=np.true_divide(1.213658-0.936068,3.92), fmt='o', c="red") # Lending
plt.errorbar(x=0.477487, y=4.2, xerr=np.true_divide(0.984142-(-0.029168),3.92), fmt='o', c="blue") # Equity
plt.errorbar(x=0.522143, y=3.8, xerr=np.true_divide(1.823640-0.220645,3.92), fmt='o', c="green") # Charity
# Appeal
plt.errorbar(x=1.790959, y=5.0, xerr=np.true_divide(1.991322-1.590595,3.92), fmt='o', c="blue") # Lending - Done
plt.errorbar(x=1.441204, y=5.2, xerr=np.true_divide(2.491060-0.391347,3.92), fmt='o', c="red") #Equity
plt.errorbar(x=0.864068, y=4.8, xerr=np.true_divide(1.462422-0.265714,3.92), fmt='o', c="green") # Charity
# Variation
plt.errorbar(x=1.236335, y=3.0, xerr=np.true_divide(1.367220-1.105450,3.92), fmt='o', c="red") # Lending
plt.errorbar(x=0.584149, y=3.2, xerr=np.true_divide(1.083733-0.084565,3.92), fmt='o', c="blue") # Equity
plt.errorbar(x=0.966268, y=2.8, xerr=np.true_divide(1.284117-0.648419,3.92), fmt='o', c="green") # Charity
# Latency
plt.errorbar(x=-1.535896, y=2.0, xerr=np.true_divide(-1.453612-(-0.618180),3.92), fmt='o', c="red") # Lending
plt.errorbar(x=-0.594363, y=2.2, xerr=np.true_divide(-0.136442-(-1.052284),3.92), fmt='o', c="blue") # Equity
plt.errorbar(x=-0.025478, y=1.8, xerr=np.true_divide(0.131920-(-0.182876),3.92), fmt='o', c="green") # Charity
# Engagement
plt.errorbar(x=-0.680682, y=1.0, xerr=np.true_divide(-0.613640-(-0.747724),3.92), fmt='o', c="red") # Lending
plt.errorbar(x=0.141986, y=1.2, xerr=np.true_divide(0.605570-(-0.321599),3.92), fmt='o', c="blue") # Equity
plt.errorbar(x=-0.956404, y=0.8, xerr=np.true_divide(-0.610256-(-1.302551),3.92), fmt='o', c="green") # Charity
plt.legend(["Lending", "Equity", "Charity"],bbox_to_anchor=(0., .95, 1., .1), loc=2, ncol=3, mode="expand", borderaxespad=0)
plt.barh(np.arange(0,6,1),
[-2]*6,
height=1.1,
color= ["#FFFFFF","#E5E4E4"]*6,
edgecolor = "none",
align="center")
plt.barh(np.arange(0,6,1),
[2]*6,
height=1.1,
color= ["#FFFFFF","#E5E4E4"]*6,
edgecolor = "none",
align="center")
plt.yticks(np.arange(6), ('', 'Engagement', 'Latency', 'Variation', 'Momentum', 'Appeal'), fontsize=12)
plt.ylim(0,6)
plt.xlim(-1.9,1.9)
ax = plt.vlines(0, 0, 1000, colors='k', linestyles='dashed')
plt.xlabel("Sample Average Treatment Effect on the Treated (SATT)", fontsize=12)
# plt.tight_layout()
plt.show()
# fig.savefig('cem.png', dpi=300)
# -
| CSCW21-Fundraising/Multiplatform/Notebooks/CEM SATT Plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import builtwith
from urllib.request import urlopen as uReq
from bs4 import BeautifulSoup
import pandas as pd
import re
builtwith.parse('https://www.netleaseadvisor.com')
html=uReq('https://www.netleaseadvisor.com/tenant/popeyes/')
soup=BeautifulSoup(html,'lxml')
print(soup.prettify())
links=soup.findAll('a')
t=1
for i in links:
print(t,i)
t+=1
QSR_links=[]
for link in links[61:86]:
QSR_links.append(link['href'])
print(link['href'])
QSR_links
QSR_links[1]
html2=uReq(QSR_links[1])
soup2=BeautifulSoup(html2,'lxml')
print(soup2.prettify())
def Average_Cap_Rate(soup):
return soup.find(string='Average Cap Rate').findNext('div').get_text()
Average_Cap_Rate(soup2)
def Average_Sale_Price(soup):
return soup.find(string='Average Sale Price').findNext('td').get_text()
Average_Sale_Price(soup2)
def NOI_f(soup):
return soup.find(string='NOI').findNext('td').get_text()
NOI_f(soup2)
def d_Square_Foot(soup):
return soup.find(string='$/Square Foot').findNext('td').get_text()
d_Square_Foot(soup2)
def Building_SF(soup):
return soup.find(string='Building SF').findNext('td').get_text()
Building_SF(soup2)
def Lot_Size(soup):
return soup.find(string='Building SF').findNext('td').get_text()
Lot_Size(soup2)
def Lease_Term(soup):
return soup.find(string='Lease Term').findNext('td').get_text()
Lease_Term(soup2)
def Escalations_f(soup):
return soup.find(string='Escalations').findNext('td').get_text()
Escalations_f(soup2)
def Stock_Symbol(soup):
return soup.find(string='Stock Symbol').findNext('td').get_text()
Stock_Symbol(soup2)
def S_P(soup):
return soup.find(string='S&P').findNext('td').get_text()
S_P(soup2)
def Moodys_f(soup):
return soup.find(string='Moody\'s').findNext('td').get_text()
Moodys_f(soup2)
def Trend_1Y(soup):
return soup.findAll('div',{'class':'trend-rate'})[0].get_text()
Trend_1Y(soup2)
def Trend_2Y(soup):
return soup.findAll('div',{'class':'trend-rate'})[1].get_text()
Trend_2Y(soup2)
QSR_links
AverageCapRates=[]
AverageSalePrices=[]
NOI=[]
DollarSquareFoot=[]
BuildingSF=[]
LotSize=[]
LeaseTerm=[]
Escalations=[]
StockSymbol=[]
SP=[]
Moodys=[]
Trend1Y=[]
Trend2Y=[]
def QSR_f(links):
for link in links:
html=uReq(link)
soup=BeautifulSoup(html,'lxml')
AverageCapRates.append(Average_Cap_Rate(soup))
AverageSalePrices.append(Average_Sale_Price(soup))
NOI.append(NOI_f(soup))
DollarSquareFoot.append(d_Square_Foot(soup))
BuildingSF.append(Building_SF(soup))
LotSize.append(Lot_Size(soup))
LeaseTerm.append(Lease_Term(soup))
Escalations.append(Escalations_f(soup))
StockSymbol.append(Stock_Symbol(soup))
SP.append(S_P(soup))
Moodys.append(Moodys_f(soup))
Trend1Y.append(Trend_1Y(soup))
Trend2Y.append(Trend_2Y(soup))
QSR_f(QSR_links)
QSR_df=pd.DataFrame({'Average Cap Rates' : AverageCapRates,
'Average Sale Prices' : AverageSalePrices,
'NOI':NOI,'$/Square Foot' : DollarSquareFoot,
'Building SF' : BuildingSF,
'Lot Size':LotSize,'Lease Term' : LeaseTerm,
'Escalations' : Escalations,
'Stock Symbol':StockSymbol,'S&P' : SP,
'Moodys' : Moodys,
'Trend1Y':Trend1Y,'Trend2Y':Trend2Y,'QSR_links':QSR_links
})
QSR_df
QSR_df.to_csv('QSR.csv', sep=',')
| QSR_Project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/pragmalingu/experiments/blob/master/00_Data/MedlineCorpus.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="ZOrLXNr5DVER"
# # Setup an Elasticsearch Instance in Google Colab
# + [markdown] id="V5AzHO1FDVEU"
# Everthing to connect to Elasticsearch, for detailed explaination see [this Notebook.](https://)
# Download:
# + id="bBnu7OFEDVEX"
import os
from subprocess import Popen, PIPE, STDOUT
# download elasticsearch
# !wget https://artifacts.elastic.co/downloads/elasticsearch/elasticsearch-7.9.1-linux-x86_64.tar.gz -q
# !tar -xzf elasticsearch-7.9.1-linux-x86_64.tar.gz
# !chown -R daemon:daemon elasticsearch-7.9.1
# + [markdown] id="bxgS1p4_-zun"
# Start a local server:
# + id="L8rAj0Ti-pOC"
# start server
es_server = Popen(['elasticsearch-7.9.1/bin/elasticsearch'],
stdout=PIPE, stderr=STDOUT,
preexec_fn=lambda: os.setuid(1) # as daemon
)
# client-side
# !pip install elasticsearch -q
from elasticsearch import Elasticsearch
from datetime import datetime
es = Elasticsearch()
es.ping() # got True
# + [markdown] id="LW6zgF8_JG4e"
# # Parsing Medline Corpus
# + [markdown] id="coEJjiebJG4j"
# You can get the corpus from [this link](http://ir.dcs.gla.ac.uk/resources/test_collections/medl/). <br>
# For detailed information about the format of the files, see the PragmaLingu [Data Sets](https://pragmalingu.de/docs/guides/data-comparison).
# You can learn about parsing in general by reading our [parsing guide](https://pragmalingu.de/docs/guides/how-to-parse).
# + [markdown] id="LDNzoWupJG4m"
# ### Dowlnoad Data
# + [markdown] id="30J3Uno8JG4o"
# Download and unzip data.
# + id="6Yo-Qv89JG4q"
# !wget http://ir.dcs.gla.ac.uk/resources/test_collections/medl/med.tar.gz
# !tar -xf med.tar.gz
# + [markdown] id="Q4za9iN6JG4y"
# Set paths to the dowloaded data as variables:
# + id="1pRBe5nzJG41"
PATH_TO_MED_TXT = '/content/MED.ALL'
PATH_TO_MED_QRY = '/content/MED.QRY'
PATH_TO_MED_REL = '/content/MED.REL'
# + [markdown] id="UaetDYMrJG47"
# ### Imports
# + [markdown] id="0RR_-4FMJG49"
# Make all the imports:
# + id="bA6ajOZ4JG4-"
from collections import defaultdict
import re
import json
from io import StringIO
import numpy as np
# + [markdown] id="r2TDnrAiJG5E"
# ### Process Data
# + [markdown] id="N5bS8EKJJG5F"
# Get the text entries from the text and query file preprocessed as a list:
# + id="Eex42pVGJG5G"
ID_marker = re.compile('^\.I',re.MULTILINE)
def get_data(PATH_TO_FILE, marker):
"""
Reads file and spilts text into entries at the ID marker '.I'.
First entry is empty, so it's removed.
'marker' contains the regex at which we want to split
"""
with open (PATH_TO_FILE,'r') as f:
text = f.read()
lines = re.split(marker,text)
lines.pop(0)
return lines
med_txt_list = get_data(PATH_TO_MED_TXT, ID_marker)
med_qry_list = get_data(PATH_TO_MED_QRY, ID_marker)
# + [markdown] id="XG02EfYUJG5M"
# Process the list of the texts and queries into nested dictionaries which can be saved as json:
# + id="Jyk8FD8dJG5M"
qry_chunk_start = re.compile('\.W')
# process the document data
med_txt_data = defaultdict(dict)
med_qry_data = defaultdict(dict)
def fill_dictionary(dictionary, chunk_list, marker, key_name):
for n in range(0,len(chunk_list)):
line = chunk_list[n]
_ , chunk = re.split(marker,line)
dictionary[n+1][key_name] = chunk.strip()
fill_dictionary(med_txt_data, med_txt_list, qry_chunk_start, 'text')
fill_dictionary(med_qry_data, med_qry_list, qry_chunk_start, 'question')
# + [markdown] id="a3IA3Hb-JG5X"
# Relevance assesments are saved as numpy and parsed to dictionary:
# + id="kN9Xp8bnJG5Y"
# process relevance assesments with rating
med_rel_data = open(PATH_TO_MED_REL)
med_np = np.loadtxt(med_rel_data, dtype=int)
med_rel_rat = defaultdict(list)
for row in med_np:
med_rel_rat[row[0]].append(row[2])
# process relevance assesments without rating
med_rel = defaultdict(list)
with open (PATH_TO_MED_REL,'r') as f:
for line in f:
line = re.split(' ',line)
med_rel[int(line[0])].append(line[2])
# + [markdown] id="fiA5r3R3JG5e"
# ### Create index for Medline corpus
# + [markdown] id="11sdTkFhpy5T"
# Create an index for the medline corpus. This is only possible if it isn't created yet.
#
# (For more information see the [Elasticsearch documentation](https://elasticsearch-py.readthedocs.io/en/master/api.html#elasticsearch.client.IndicesClient.create))
# + id="cXBubnBUJG5f"
#create index, see https://elasticsearch-py.readthedocs.io/en/master/api.html#elasticsearch.client.IndicesClient.create
es.indices.create("medline-corpus")
#print new index list
create_response = es.cat.indices()
print(create_response)
# + [markdown] id="EOarkQK0p5mI"
# Index all the documents that are processed to the created index in elasticsearch:
#
# (For more information see the [Elasticsearch documentation](https://elasticsearch-py.readthedocs.io/en/master/#example-usage))
# + id="7UMoCAEdJG5j"
#index document, see https://elasticsearch-py.readthedocs.io/en/master/#example-usage
med_index = "medline-corpus"
for ID, doc_data in txt_data.items():
es.index(index=med_index, id=ID, body=doc_data)
# + [markdown] id="3NnzjwaHp-9I"
# Verify if everthing went right by printing it:
# + id="egkB6CTaqBPg" colab={"base_uri": "https://localhost:8080/", "height": 459} outputId="acb30a69-e785-47e5-da22-f5e8766fb810"
create_response = es.cat.indices()
print(create_response)
# + [markdown] id="DfDLyAs3JG5n"
# ### Use Corpus in Ranking API
# + [markdown] id="TCYE0zm5qFa_"
# Use the ranking evaluation API from elasticsearch to evaluate the corpus:
#
# (For more information see the [python documentation](https://elasticsearch-py.readthedocs.io/en/master/api.html?highlight=_rank_eval#elasticsearch.Elasticsearch.rank_eval) and the [Elasticsearch documentation](https://www.elastic.co/guide/en/elasticsearch/reference/current/search-rank-eval.html#search-rank-eval))
# + id="csXCyPM1JG5p"
#use rank eval api, see https://elasticsearch-py.readthedocs.io/en/master/api.html?highlight=_rank_eval#elasticsearch.Elasticsearch.rank_eval
#and https://www.elastic.co/guide/en/elasticsearch/reference/current/search-rank-eval.html#search-rank-eval
from collections import defaultdict
med_index = "medline-corpus"
def create_query_body(query_dict, rel_dict, index_name):
"""
The function creates a request for every query in query_dict and rates the relevant documents with rel_dict to 1.
The index name has to be the same as from the documents your looking at.
An evaluation body for the elasticsearch ranking API is returned.
"""
eval_body = {
"requests":'',
"metric": {
"recall": {
"relevant_rating_threshold": 1,
"k": 20
}
}
}
requests = []
current_request = defaultdict(lambda: defaultdict())
current_rel = {"_index": index_name, "_id": '', "rating": int}
for query_ID, query_txt in query_dict.items():
current_query = {"query": { "multi_match": { "query": '' , "fields" : ["title","text"]}}}
current_query["query"]["multi_match"]["query"] = query_txt['question']
current_request["id"] = 'Query_'+str(query_ID)
current_request["request"] = current_query.copy()
current_request["ratings"] = [{"_index": index_name, "_id": str(el), "rating": 1} for el in rel_dict[query_ID]]
requests.append(current_request.copy())
eval_body["requests"] = requests
return eval_body
med_create = create_query_body(med_qry_data, med_rel, med_index)
med_eval_body = json.dumps(med_create)
# + [markdown] id="GHQ27azrJG5s"
# Print results of Ranking API:
# + id="IxStXr31JG5u"
med_res = es.rank_eval(med_eval_body, med_index)
print(json.dumps(med_res, indent=4, sort_keys=True))
| 00_Data/MedlineCorpus.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W1D3-ModelFitting/student/W1D3_Tutorial1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="sSukjRujSzSt"
# # Neuromatch Academy: Week 1, Day 3, Tutorial 1
# # Model Fitting: Linear regression with MSE
#
# + [markdown] colab_type="text" id="EBYoWocQ2hDw"
# #Tutorial Objectives
#
# This is Tutorial 1 of a series on fitting models to data. We start with simple linear regression, using least squares optimization (Tutorial 1) and Maximum Likelihood Estimation (Tutorial 2). We will use bootstrapping to build confidence intervals around the inferred linear model parameters (Tutorial 3). We'll finish our exploration of linear models by generalizing to multiple linear regression (Tutorial 4). We then move on to polynomial regression (Tutorial 5). We end by learning how to choose between these various models. We discuss the bias-variance trade-off (Tutorial 6) and two common methods for model selection, AIC and Cross Validation (Tutorial 7).
#
# In this tutorial, we will learn how to fit simple linear models to data.
# - Learn how to calculate the mean-squared error (MSE)
# - Explore how model parameters (slope) influence the MSE
# - Learn how to find the optimal model parameter using least-squares optimization
#
# ---
#
# **acknowledgements:**
# - we thank <NAME>, much of this tutorial is inspired by exercises asigned in his mathtools class.
# + [markdown] colab_type="text" id="kUp2TAyO1yzv"
# # Setup
# + cellView="form" colab={} colab_type="code" id="7lFaCKBThezP"
# @title Imports
import numpy as np
import matplotlib.pyplot as plt
import ipywidgets as widgets
# + cellView="form" colab={} colab_type="code" id="DN-998MpYd4t"
#@title Figure Settings
# %matplotlib inline
fig_w, fig_h = (8, 6)
plt.rcParams.update({'figure.figsize': (fig_w, fig_h)})
# %config InlineBackend.figure_format = 'retina'
# + [markdown] colab_type="text" id="Ph9hfNoYZnF4"
# # Mean Squared Error (MSE)
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 518} colab_type="code" id="bZeldtxEZkHQ" outputId="63d9ebb8-a795-4c7f-bad8-32aa6f2c7fe3"
#@title Video: Linear Regression & Mean Squared Error
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="8e9Hvc8zaUQ", width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
# + [markdown] colab_type="text" id="Y5n3kKMXi9QA"
# Linear least squares regression is an old but gold optimization procedure that we are going to use for data fitting. Least squares (LS) optimization problems are those in which the objective function is a quadratic function of the
# parameter(s) being optimized.
#
# Suppose you have a set of measurements, $y_{n}$ (the "dependent" variable) obtained for different input values, $x_{n}$ (the "independent" or "explanatory" variable). Suppose we believe the measurements are proportional to the input values, but are corrupted by some (random) measurement errors, $\epsilon_{n}$, that is:
#
# $$y_{n}= \theta x_{n}+\epsilon_{n}$$
#
# for some unknown slope parameter $\theta.$ The least squares regression problem uses mean squared error (MSE) as its objective function, it aims to find the value of the parameter $\theta$ by minimizing the average of squared errors:
#
# \begin{align}
# \min _{\theta} \frac{1}{N}\sum_{n=1}^{N}\left(y_{n}-\theta x_{n}\right)^{2}
# \end{align}
# + [markdown] colab_type="text" id="wZXaDXvhkYOb"
# We will now explore how MSE is used in fitting a linear regression model to data. For illustrative purposes, we will create a simple synthetic dataset where we know the true underlying model. This will allow us to see how our estimation efforts compare in uncovering the real model (though in practice we rarely have this luxury).
#
# First we will generate some noisy samples $x$ from [0, 10) along the line $y = 1.2x$ as our dataset we wish to fit a model to.
# + colab={"base_uri": "https://localhost:8080/", "height": 386} colab_type="code" id="aFnZzv3rlklT" outputId="2045c362-8a85-49a3-b178-1ee99b76c034"
# setting a fixed seed to our random number generator ensures we will always
# get the same psuedorandom number sequence
np.random.seed(121)
theta = 1.2
n_samples = 30
x = 10*np.random.rand(n_samples) # sample from a uniform distribution over [0,10)
noise = np.random.randn(n_samples) # sample from a standard normal distribution
y = theta*x + noise
fig, ax = plt.subplots()
ax.scatter(x, y) # produces a scatter plot
ax.set(xlabel='x', ylabel='y');
# + [markdown] colab_type="text" id="aNXpFHRZ24zD"
# Now that we have our suitably noisy dataset, we can start trying to estimate the underlying model that produced it. We use MSE to evaluate how successful a particular slope estimate $\hat{\theta}$ is for explaining the data, with the closer to 0 the MSE is, the better our estimate fits the data.
# + [markdown] colab_type="text" id="Vd6prQstiBue"
# ### Exercise: Compute MSE
#
# In this exercise you will implement a method to compute the mean squared error for a set of inputs $x$, measurements $y$, and slope estimate $\hat{\theta}$.
# + colab={} colab_type="code" id="gNdJYx4Fi9Q5"
def mse(x, y, theta_hat):
"""Compute the mean squared error
Args:
x (ndarray): An array of shape (samples,) that contains the input values.
y (ndarray): An array of shape (samples,) that contains the corresponding
measurement values to the inputs.
theta_hat (float): An estimate of the slope parameter.
Returns:
float: The mean squared error of the data with the estimated parameter.
"""
####################################################
## TODO for students: compute the mean squared error
####################################################
# comment this out when you've filled
raise NotImplementedError("Student excercise: compute the mean squared error")
return mse
# + [markdown] colab_type="text" id="6Xql92Togvad"
# Now that we have our MSE method, let's see what errors we observe for different values of $\hat{\theta}$.
#
#
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 70} colab_type="code" id="7xVUmJIpeyDk" outputId="8fa93136-0110-410d-e847-365c72761768"
theta_hats = [0.75, 1.0, 1.5]
for theta_hat in theta_hats:
print(f"theta_hat of {theta_hat} has an MSE of {mse(x, y, theta_hat):.2f}")
# + [markdown] colab_type="text" id="v-hIQWZPd-mA"
# We should see that $\hat{\theta} = 1.0$ is our best estimate from the three we tried. Looking just at the raw numbers, however, isn't always satisfying, so let's visualize what our estimated model looks like over the data.
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 294} colab_type="code" id="35Z-7MbAi9RP" outputId="4611880f-ac68-486d-935e-ea42439b3918"
fig, axes = plt.subplots(ncols=3, figsize=(18,4))
for theta_hat, ax in zip(theta_hats, axes):
ax.scatter(x, y) # our data scatter plot
y_hat = theta_hat * x
ax.plot(x, y_hat, color='C1') # our estimated model
ax.set(
title=f"theta_hat = {theta_hat}, MSE = {mse(x, y, theta_hat):.2f}",
xlabel='x',
ylabel='y'
);
# + [markdown] colab_type="text" id="E7CYoA8s3oRI"
# ### Data Exploration
#
# Using an interactive widget, we can easily see how changing our slope estimate changes our model fit. Note that we also display the residuals as segments between observed responses (the data point), and predicted responses (on the slope).
# + cellView="both" colab={"base_uri": "https://localhost:8080/", "height": 434, "referenced_widgets": ["0fb1aa78cc764fe18be5a6a4e6c3a7f0", "f2dedb93e6114ea5bbdd8081c7b689a5", "<KEY>", "<KEY>", "af4caf82c9ac426e98a885d9962a6fef", "b1395dd66f7e48008a541e920b2da441", "90d39e1d04ea4c5480e9b0b91b638bec"]} colab_type="code" id="klWaQnZ7tA0D" outputId="12f69002-70ef-4566-cfb4-3b90ed656591"
#@title MSE Explorer
@widgets.interact(theta_hat=widgets.FloatSlider(1.0, min=0.0, max=2.0))
def plot_data_estimate(theta_hat):
fig, ax = plt.subplots()
ax.scatter(x, y) # our data scatter plot
y_hat = theta_hat * x
ax.plot(x, y_hat, color='C1', label='fit') # our estimated model
# plot residuals
ymin = np.minimum(y, y_hat)
ymax = np.maximum(y, y_hat)
ax.vlines(x, ymin, ymax, 'g', alpha=0.5, label='residuals')
ax.set(
title=f"theta_hat = {theta_hat}, MSE = {mse(x, y, theta_hat):.2f}",
xlabel='x',
ylabel='y'
)
ax.legend()
# + [markdown] colab_type="text" id="CEAIahS3i9Ri"
# While exploring single estimates by hand can be instructive, it's not the most efficient for finding the best estimate to fit our data. Another technique we can use is to evaluate the MSE across a reasonable range of values and plot the resulting error surface.
# + colab={"base_uri": "https://localhost:8080/", "height": 403} colab_type="code" id="_YQqDw0g4Gnd" outputId="53407341-ba51-4622-adfe-8e831709464b"
theta_hats = np.linspace(-2.0, 4.0)
errors = np.zeros(len(theta_hats))
for i, theta_hat in enumerate(theta_hats):
errors[i] = mse(x, y, theta_hat)
fig, ax = plt.subplots()
ax.plot(theta_hats, errors, label='mse', c='C1')
ax.axvline(theta, color='r', ls='--', label="theta")
best_error = np.min(errors)
best_fit = theta_hats[np.argmin(errors)]
ax.set(
title=f"Best fit: theta_hat = {best_fit:.2f}, MSE = {best_error:.2f}",
xlabel='theta_hat',
ylabel='MSE')
ax.legend();
# + [markdown] colab_type="text" id="jy039uYb4FBE"
# We can see that our best fit is $\hat{\theta}=1.18$ with an MSE of 1.45. This is quite close to the original true value $\theta=1.2$!
#
# While this approach quickly got us to a good estimate, it still relied on evaluating the MSE value across a grid of hand-specified values. If we didn't pick a good range to begin with, or with enough granularity, we might end up in a local minima and not find the best possible estimator. Let's go one step further, and instead of finding the minimum MSE from a set of candidate estimates, let's solve for it analytically.
#
# We can do this by minimizing the cost function. Mean squared error is a convex objective function, therefore we can compute its minimum using calculus.
#
# We set the derivative of the error expression with respect to $\theta$ equal to zero,
#
# \begin{align}
# \frac{d}{d\theta}\frac{1}{N}\sum_{i=1}^N(\hat{y}_i-y_i)^2 = 0
# \end{align}
#
# solving for $\theta$, we obtain an optimal value of:
#
# \begin{align}
# \hat\theta = \sum_{i=1}^N \frac{x_i y_i}{x_i^2}
# \end{align}
#
# This is known as solving the *normal equations*. For different ways of obtaining the solution, see the notes on [Least Squares Optimization](https://www.cns.nyu.edu/~eero/NOTES/leastSquares.pdf) by <NAME>.
# + [markdown] colab_type="text" id="fq4jThOV5zIH"
# ### Exercise: Solve for the Optimal Estimator
#
# In this exercise, you will write a function that finds the optimal $\hat{\theta}$ value using the least squares optimization approach (the previous equation above) to solving MSE minimization. It shoud take as arguments $x$ and $y$, and return the solution $\hat{\theta}$.
# + colab={} colab_type="code" id="YArCcg6Di9R6"
def solve_normal_eqn(x, y):
"""Solve the normal equations to produce the value of theta_hat that minimizes
MSE.
Args:
x (ndarray): An array of shape (samples,) that contains the input values.
y (ndarray): An array of shape (samples,) that contains the corresponding
measurement values to the inputs.
Returns:
float: the value for theta_hat arrived from minimizing MSE
"""
######################################################################
## TODO for students: solve for the best parameter using least squares
######################################################################
# comment this out when you've filled
raise NotImplementedError("Student excercise: solve for theta_hat using least squares")
return theta_hat
# + [markdown] colab_type="text" id="AXAqq1Ht54H7"
# Use your function to compute an estimate $\hat{\theta}$ and plot the resulting prediction on top of the data.
# + colab={"base_uri": "https://localhost:8080/", "height": 403} colab_type="code" id="bu_pnb8ti9SI" outputId="8d6f41f0-a9bc-4c86-829b-765ff2d9cdda"
theta_hat = solve_normal_eqn(x, y)
y_hat = theta_hat * x
fig, ax = plt.subplots()
ax.scatter(x, y) # our data scatter plot
ax.plot(x, y_hat, color='C1', label='fit') # our estimated model
# plot residuals
ymin = np.minimum(y, y_hat)
ymax = np.maximum(y, y_hat)
ax.vlines(x, ymin, ymax, 'g', alpha=0.5, label='residuals')
ax.set(
title=f"theta_hat = {theta_hat:0.2f}, MSE = {mse(x, y, theta_hat):.2f}",
xlabel='x',
ylabel='y'
)
ax.legend();
# + [markdown] colab_type="text" id="ThUTocisKLwX"
# We see that the analytic solution produces an even better result than our grid search from before, producing $\hat{\theta} = 1.21$ with MSE = 1.43!
# + [markdown] colab_type="text" id="K15UOyozi9SY"
# ## Summary
#
# - Linear least squares regression is an optimisation procedure that can be used for data fitting:
# - Task: predict a value for $y$ given $x$
# - Performance measure: $\textrm{MSE}$
# - Procedure: minimize $\textrm{MSE}$ by solving the normal equations
# - **Key point**: We fit the model by defining an *objective function* and minimizing it.
# - **Note**: In this case, there is an *analytical* solution to the minimization problem and in practice, this solution can be computed using *linear algebra*. This is *extremely* powerful and forms the basis for much of numerical computation throughout the sciences.
| tutorials/W1D3-ModelFitting/student/W1D3_Tutorial1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pulp
import numpy as np
import networkx as nx
from numpy import genfromtxt
import pandas as pd
import matplotlib.pyplot as plt
import time
import seaborn as sns
from numpy import genfromtxt
from numpy import linalg as LA
import scipy as sp
import sympy
import sys
import networkx.algorithms.approximation as nxaa
mydata = genfromtxt('1hmk.csv', delimiter=',')
mydata = mydata[1:,1:]
m = len(mydata)
print(m)
# +
for i in range(0,m):
for j in range(0,m):
if i ==j:
mydata[i][j] = 1
print(mydata)
# -
plt.figure(figsize=(12,8))
plt.imshow(mydata, cmap='spring', interpolation='nearest')
plt.show()
# ### creat the graph using networkX
G = nx.Graph(mydata)
# ## removing isolated nodes
"""
for component in list(nx.connected_components(G)):
if len(component)<= 3:
for node in component:
G.remove_node(node)
m = len(G.nodes)
print(m)
"""
# # MIP package
from mip import Model, xsum, maximize, BINARY
from itertools import product
from sys import stdout as out
from mip import Model, xsum, minimize, BINARY
# +
m = Model("minimum_dominating_set")
I = range(len(G.nodes()))
x = [m.add_var(var_type=BINARY) for i in I]
m.objective = minimize(xsum(x[i] for i in I))
for j in G.nodes():
m += x[j]+ xsum(x[u] for u in G.neighbors(j)) >= 1
m.optimize()
selected=[]
number_of_mds=0
for i in I:
if x[i].x >= 0.99:
selected = np.append(selected,i)
number_of_mds += 1
print("selected items: {}".format(selected))
print(number_of_mds)
# -
# # PuLp package
# +
# define the problem
prob = pulp.LpProblem("minimum_dominating_set", pulp.LpMinimize)
# define the variables
x = pulp.LpVariable.dicts("x", G.nodes(), cat=pulp.LpBinary)
# define the objective function
start_time = time.time()
for (v,u) in G.edges():
prob += pulp.lpSum(x)
# define the constraints
for v in G.nodes():
prob += x[v] + pulp.lpSum([x[u] for u in G.neighbors(v)]) >= 1
color_map = []
# solve
prob.solve()
end_time = time.time()
print("time = %s seconds" % (end_time - start_time))
# display solution
for v in G.nodes():
if pulp.value(x[v]) > 0.99:
color_map.append('red')
print(v,end=',')
else:
color_map.append('dodgerblue')
plt.figure(3,figsize=(10,10))
nx.draw(G, node_color=color_map, node_size=25,width=0.55, with_labels=True,font_size=8)
plt.savefig("1b9c_center.pdf")
plt.show()
# -
i = 0
for v in prob.variables():
if v.varValue == 1:
#print(v.name, "=", v.varValue)
i = i+1
print(i)
def nodes_connected(u, v):
return u in G.neighbors(v)
nodes_connected(40,213)
| full data mds_final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] cell_id="5c855f28-ce38-4df2-b4cb-58e24c6bcc8b" deepnote_cell_type="markdown" tags=[]
# # Week 8 Video notebooks
# + cell_id="74bf1174-6047-4a8a-9ca4-bb128b14839c" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=4 execution_start=1644968158257 source_hash="aaa7822b" tags=[]
import numpy as np
import torch
from torch import nn
from torchvision.transforms import ToTensor
# + [markdown] cell_id="c4c9b9b7-9fdd-4af7-a90a-e65b99680c44" deepnote_cell_type="markdown" tags=[]
# ## A little neural network
#
# 
# + [markdown] cell_id="2d85b966-b29b-4e00-b5a0-392115b5e749" deepnote_cell_type="markdown" tags=[]
# ## Neural network for logical or
# + cell_id="8b1be2b3-3cb0-435d-a3a6-69158258cc2f" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=0 execution_start=1644969299973 source_hash="9d191a80" tags=[]
X = torch.tensor([[0,0],[0,1],[1,0],[1,1]]).to(torch.float)
y = torch.tensor([0,1,1,1]).to(torch.float).reshape(-1,1)
# + cell_id="01c1249a-b7f8-4ec8-9f9f-7ef69807061a" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=0 execution_start=1644968630637 source_hash="33d863b7" tags=[]
class LogicOr(nn.Module):
def __init__(self):
super().__init__()
self.layers = nn.Sequential(
nn.Linear(2,1),
nn.Sigmoid()
)
def forward(self,x):
return self.layers(x)
# + cell_id="21cd842b-41be-4809-b7f6-783d7637a2fe" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=3 execution_start=1644968634355 source_hash="2b0613f4" tags=[]
model = LogicOr()
# + cell_id="c3e6768e-941b-41d2-9c2a-8be1da673844" deepnote_cell_type="code" deepnote_output_heights=[78.78125] deepnote_to_be_reexecuted=false execution_millis=168 execution_start=1644968645304 source_hash="c38c1e75" tags=[]
model(X)
# + cell_id="834405d7-2af3-400e-9691-84765e07adc4" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=6 execution_start=1644968671208 source_hash="37e1d2f6" tags=[]
for p in model.parameters():
print(p)
# + [markdown] cell_id="8fad4e7d-eb32-463f-8e1e-ceac0c6d9cf5" deepnote_cell_type="markdown" tags=[]
# ## Evaluating our neural network
# + cell_id="2f44c910-21a9-4e99-940f-b9c5ef70ab28" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=2 execution_start=1644969222213 source_hash="7d63c4c4" tags=[]
loss_fn = nn.BCELoss()
# + cell_id="25ae4952-6448-4d0b-bb79-51f49010f66e" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=7 execution_start=1644969306452 source_hash="57b8ea20" tags=[]
y.shape
# + cell_id="f7c83d83-0c62-4b4e-a2c9-daea1fd79a91" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=10 execution_start=1644969312995 source_hash="792aba57" tags=[]
loss_fn(model(X),y)
# + [markdown] cell_id="17d4d716-4a9d-4767-8b74-4e0999d06733" deepnote_cell_type="markdown" tags=[]
# ## Optimizer
# + cell_id="ba0f4523-e302-411a-8959-01aa3e373d6f" deepnote_cell_type="code" deepnote_output_heights=[155.59375] deepnote_to_be_reexecuted=false execution_millis=13 execution_start=1644969554983 source_hash="c2972089" tags=[]
optimizer = torch.optim.SGD(model.parameters(), lr=0.1)
# + cell_id="6e81db50-f5ad-4d3c-9a75-a0037b9ada01" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=10 execution_start=1644969583063 source_hash="418181e6" tags=[]
for p in model.parameters():
print(p)
print(p.grad)
print("")
# + cell_id="828bc4ae-650f-4889-be15-290644099100" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=1 execution_start=1644969626940 source_hash="2997554f" tags=[]
loss = loss_fn(model(X),y)
# + cell_id="1bf4d5cf-6ea5-4297-b57f-438a75d2a21b" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=8 execution_start=1644969663962 source_hash="ff19e473" tags=[]
optimizer.zero_grad()
loss.backward()
# + cell_id="2a0b48e3-94c0-45bf-bf0d-8b55b8751649" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=3 execution_start=1644969685145 source_hash="418181e6" tags=[]
for p in model.parameters():
print(p)
print(p.grad)
print("")
# + cell_id="4fded704-ec98-41bb-a2f3-eb077e72739f" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=1989 execution_start=1644969702952 source_hash="8a03d43" tags=[]
optimizer.step()
# + cell_id="ea1846c7-dc01-40ee-bedd-471642333466" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=8 execution_start=1644969722307 source_hash="418181e6" tags=[]
for p in model.parameters():
print(p)
print(p.grad)
print("")
# + [markdown] cell_id="97109ce9-fb42-4727-a93c-35a758afdb7f" deepnote_cell_type="markdown" tags=[]
# ## Training the model
# + cell_id="44217d4b-eb23-4f34-9892-0130a2b4258f" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=5 execution_start=1644970100898 source_hash="fabc3f51" tags=[]
100%50
# + cell_id="e5f463da-f27a-43dd-ac45-8207611667cb" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=266 execution_start=1644970111352 source_hash="3bb4db40" tags=[]
epochs = 1000
for i in range(epochs):
loss = loss_fn(model(X),y)
optimizer.zero_grad()
loss.backward()
optimizer.step()
if i%50 == 0:
print(loss)
# + cell_id="9a2d4c47-a751-4d07-a897-b49ea9ceca53" deepnote_cell_type="code" tags=[]
epochs = 1000
for i in range(epochs):
loss = loss_fn(model(X),y)
optimizer.zero_grad()
loss.backward()
optimizer.step()
if i%100 == 0:
print(loss)
# + cell_id="860500ee-facb-4d95-a92a-301b2d3b8343" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=289 execution_start=1644970238417 source_hash="7a2c19a9" tags=[]
epochs = 1000
for i in range(epochs):
loss = loss_fn(model(X),y)
optimizer.zero_grad()
loss.backward()
optimizer.step()
if i%100 == 0:
print(loss)
# + cell_id="7b30549a-3a16-4fbf-9c64-850d9e3cfd92" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=190 execution_start=1644970242128 source_hash="418181e6" tags=[]
for p in model.parameters():
print(p)
print(p.grad)
print("")
# + cell_id="748c3ffe-af9a-431f-8dc9-cc6d87633c27" deepnote_cell_type="code" deepnote_output_heights=[78.78125] deepnote_to_be_reexecuted=false execution_millis=2 execution_start=1644970260113 source_hash="c38c1e75" tags=[]
model(X)
# + [markdown] cell_id="b336dc8b-d766-4eb7-99b4-3feeb3307a0b" deepnote_cell_type="markdown" tags=[]
# ## A little neural network: results
#
# 
| _build/jupyter_execute/Week8/Week8-Video-notebooks.ipynb |