code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Лабораторная работа №1.
#
# Данная лабораторная работа состоит из нескольких блоков. В каждом блоке вам предлагается произвести некоторые манипуляции с данными и сделать некоторые выводы.
# * Задавать вопросы можно и нужно.
# * Списывать не нужно. Работы, которые были списаны обнуляются.
# * Блоки выполняются последовательно и оцениваются отдельно.
# ## Part 2. SVM and kernels
# + [markdown] nbgrader={"grade": false, "grade_id": "cell-c7b8f71403aa9084", "locked": true, "schema_version": 2, "solution": false}
# Kernels concept get adopted in variety of ML algorithms (e.g. Kernel PCA, Gaussian Processes, kNN, ...).
#
# So in this task you are to examine kernels for SVM algorithm applied to rather simple artificial datasets.
#
# To make it clear: we will work with the classification problem through the whole notebook.
# + nbgrader={"grade": false, "grade_id": "cell-57f562bf4f554fae", "locked": true, "schema_version": 2, "solution": false}
from sklearn.datasets import make_moons
import matplotlib.pyplot as plt
import numpy as np
# + [markdown] nbgrader={"grade": false, "grade_id": "cell-1b128784928e8df1", "locked": true, "schema_version": 2, "solution": false}
# Let's generate our dataset and take a look on it.
# + nbgrader={"grade": false, "grade_id": "cell-ee8cf8e9cf114b9d", "locked": true, "schema_version": 2, "solution": false}
moons_points, moons_labels = make_moons(n_samples=500, noise=0.2, random_state=42)
plt.scatter(moons_points[:, 0], moons_points[:, 1], c=moons_labels)
# + [markdown] nbgrader={"grade": false, "grade_id": "cell-35b09404d22ab9f4", "locked": true, "schema_version": 2, "solution": false}
# ## 1.1 Pure models.
# First let's try to solve this case with good old Logistic Regression and simple (linear kernel) SVM classifier.
#
# Train LR and SVM classifiers (choose params by hand, no CV or intensive grid search neeeded) and plot their decision regions. Calculate one preffered classification metric.
#
# Describe results in one-two sentences.
#
# _Tip:_ to plot classifiers decisions you colud use either sklearn examples ([this](https://scikit-learn.org/stable/auto_examples/neural_networks/plot_mlp_alpha.html#sphx-glr-auto-examples-neural-networks-plot-mlp-alpha-py) or any other) and mess with matplotlib yourself or great [mlxtend](https://github.com/rasbt/mlxtend) package (see their examples for details)
#
# _Pro Tip:_ wirte function `plot_decisions` taking a dataset and an estimator and plotting the results cause you want to use it several times below
# + nbgrader={"grade": true, "grade_id": "cell-550546e70e191bc3", "locked": false, "points": 10, "schema_version": 2, "solution": true}
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
from mlxtend.plotting import plot_decision_regions
lr = LogisticRegression() # add some params
svm = SVC(kernel='linear') # here too
### YOUR CODE HERE
# -
# ## 1.2 Kernel tirck
#
# 
#
# Now use different kernels (`poly`, `rbf`, `sigmoid`) on SVC to get better results. Play `degree` parameter and others.
#
# For each kernel estimate optimal params, plot decision regions, calculate metric you've chosen eariler.
#
# Write couple of sentences on:
#
# * What have happenned with classification quality?
# * How did decision border changed for each kernel?
# * What `degree` have you chosen and why?
# + nbgrader={"grade": true, "grade_id": "cell-3a1681e6d52ed236", "locked": false, "points": 15, "schema_version": 2, "solution": true}
### YOUR CODE HERE
# + [markdown] nbgrader={"grade": false, "grade_id": "cell-ba9a59e3ec57f514", "locked": true, "schema_version": 2, "solution": false}
# ## 1.3 Simpler solution (of a kind)
# What is we could use Logisitc Regression to successfully solve this task?
#
# Feature generation is a thing to help here. Different techniques of feature generation are used in real life, couple of them will be covered in additional lectures.
#
# In particular case simple `PolynomialFeatures` ([link](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.PolynomialFeatures.html)) are able to save the day.
#
# Generate the set of new features, train LR on it, plot decision regions, calculate metric.
#
# * Comare SVM's results with this solution (quality, borders type)
# * What degree of PolynomialFeatures have you used? Compare with same SVM kernel parameter.
# + nbgrader={"grade": true, "grade_id": "cell-58a1e03cab2ca349", "locked": false, "points": 15, "schema_version": 2, "solution": true}
from sklearn.preprocessing import PolynomialFeatures
### YOUR CODE HERE
# + [markdown] nbgrader={"grade": false, "grade_id": "cell-868839a4a8358c59", "locked": true, "schema_version": 2, "solution": false}
# ## 1.4 Harder problem
#
# Let's make this task a bit more challenging via upgrading dataset:
# + nbgrader={"grade": false, "grade_id": "cell-86be614f32559cea", "locked": true, "schema_version": 2, "solution": false}
from sklearn.datasets import make_circles
circles_points, circles_labels = make_circles(n_samples=500, noise=0.06, random_state=42)
plt.figure(figsize=(5, 5))
plt.scatter(circles_points[:, 0], circles_points[:, 1], c=circles_labels)
# + [markdown] nbgrader={"grade": false, "grade_id": "cell-e7e5a8e0da66afbe", "locked": true, "schema_version": 2, "solution": false}
# And even more:
# + nbgrader={"grade": false, "grade_id": "cell-7a98ef8e43822e61", "locked": true, "schema_version": 2, "solution": false}
points = np.vstack((circles_points*2.5 + 0.5, moons_points))
labels = np.hstack((circles_labels, moons_labels + 2)) # + 2 to distinct moons classes
plt.figure(figsize=(5, 5))
plt.scatter(points[:, 0], points[:, 1], c=labels)
# + [markdown] nbgrader={"grade": false, "grade_id": "cell-7c2a785a2d63ce73", "locked": true, "schema_version": 2, "solution": false}
# Now do your best using all the approaches above!
#
# Tune LR with generated features, SVM with appropriate kernel of your choice. You may add some of your loved models to demonstrate their (and your) strength. Again plot decision regions, calculate metric.
#
# Justify the results in a few phrases.
# + nbgrader={"grade": true, "grade_id": "cell-e61b36ea61909c83", "locked": false, "points": 40, "schema_version": 2, "solution": true}
### YOUR CODE HERE
| homeworks_basic/Lab1_Ensembles_and_SVM/Lab1_part2_SVM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (cvxpy)
# language: python
# name: cvxpy
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Семинар 10
# # Оптимизация на множествах простой структуры
# + [markdown] slideshow={"slide_type": "slide"}
# ## На прошлом семинаре...
#
# - Метод Ньютона
# - Квазиньютоновские методы
# + [markdown] slideshow={"slide_type": "slide"}
# ## Методы решения каких задач уже известны или скоро будут известны
#
# - Безусловная минимизация: функция достаточно гладкая, но ограничений на аргумент нет.
# - Линейное программирование: линейная функция при линейных ограничениях - следующий семинар
#
# Следующий шаг:
#
# - произвольная достаточно гладкая функция на достаточно простом множестве - не обязательно полиэдральном.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Что такое "простое множество"?
#
# + [markdown] slideshow={"slide_type": "fragment"}
# - **Определение.** Множество будем называть *простым*, если проекцию на него можно найти существенно быстрее (чаще всего аналитически) по сравнению с решением исходной задачи минимизации.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Примеры простых множеств
#
# - Полиэдр $Ax = b, Cx \leq d$
# - аффинное множество
# - гиперплоскость
# - полупространство
# - отрезок, интервал, полуинтервал
# - симплекс
# - Конусы
# - положительный ортант
# - Лоренцев конус
# - $\mathbb{S}^n_{+}$
#
# **Замечание:** убедитесь, что Вы понимаете, что стоит за этими названиями и обозначениями!
# + [markdown] slideshow={"slide_type": "slide"}
# ## Что такое проекция?
# Для данной точки $y \in \mathbb{R}^n$ требуется решить следующую задачу
#
# $$
# \min_{x \in P} \|x - y \|_2
# $$
#
# Обозначение: $\pi_P(y)$ - проекция точки $y$ на множество $P$.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Примеры проекций
#
# - Отрезок $P = \{x | l \leq x \leq u \}$
#
# $$
# (\pi_P(y))_k =
# \begin{cases}
# u_k & y_k \geq u_k \\
# l_k & y_k \leq l_k \\
# y_k & \text{otherwise.}
# \end{cases}
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# - Аффинное множество $P = \{ x| Ax = b \}$
# $$
# \pi_P(y) = y - A^+(Ay - b),
# $$
# где $A^+$ - псевдообратная матрица. Если $A$ полного ранга и столбцы линейно-независимы, тогда $A^+ = (A^{\top}A)^{-1}A^{\top}$.
# + [markdown] slideshow={"slide_type": "slide"}
# - Конус положительно полуопределённых матриц $P = \mathbb{S}^n_+ = \{X \in \mathbb{R}^{n \times n} | X \succeq 0, \; X^{\top} = X \}$
# $$
# \pi_P(Y) = \sum_{i=1}^n (\lambda_i)_+ v_i v_i^{\top},
# $$
# где $(\lambda_i, v_i)$ - пары собственных значений и векторов матрицы $Y$.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Метод проекции градиента
# + [markdown] slideshow={"slide_type": "fragment"}
# $$
# \min_{x \in P} f(x)
# $$
# **Идея**: делать шаг градиентного спуска и проецировать полученную точку на допустимое множество $P$.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Псевдокод
#
# ```python
# def ProjectedGradientDescent(f, gradf, proj, x0, tol):
#
# x = x0
#
# while True:
#
# gradient = gradf(x)
#
# alpha = get_step_size(x, f, gradf, proj)
#
# x = proj(x - alpha * grad)
#
# if check_convergence(x, f, tol):
#
# break
#
# return x
#
# ```
# + [markdown] slideshow={"slide_type": "slide"}
# ## Поиск шага
#
# - Постоянный шаг: $\alpha_k = \alpha$, где $\alpha$ достаточно мало
# - Наискорейший спуск:
# $$
# \min_{\alpha > 0} f(x_k(\alpha))
# $$
# $x_k(\alpha) = \pi_P (x_k - \alpha f'(x_k))$
# - Линейный поиск: уменьшать шаг по правилу Армихо, пока не будет выполнено условие
# $$
# f(x_k(\alpha)) - f(x_k) \leq c_1 \langle f'(x_k), x_k(\alpha) - x_k \rangle
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# ## Теорема сходимости (<NAME> "Введение в оптимизацию", гл. 7, $\S$ 2)
#
# **Теорема.** Пусть $f$ выпуклая дифференцируемая функция и её градиент липшицев на $P$ с константой $L$. Пусть $P$ выпуклое и замкнутое множество и $0 < \alpha < 2 / L$.
#
# Тогда
# - $x_k \to x^*$
# - если $f$ сильно выпуклая, то $x_k \to x^*$ со скоростью геометрической прогрессии
# - если $f$ дважды дифференцируема и $f''(x) \succeq l\mathbf{I}, \; x \in P$, $l > 0$, то знаменатель прогрессии $q = \max \{ |1 - \alpha l|, |1 - \alpha L|\}$.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Критерии остановки
# - Сходимость по аргументу, то есть сходимость последовательности $x_k$ к предельной точке $x^*$
# - $x_k = x^*$ если $x_k = \pi_P(x_{k+1})$
#
# **Важное замечание:** проверять норму градиента бессмысленно, так как это условная оптимизация!
# + [markdown] slideshow={"slide_type": "slide"}
# ## Аффинная инвариантность
#
# **Упражнение.** Проверьте является ли метод проекции градиента аффинно инвариантным.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Pro & Contra
#
# Pro
# - часто можно аналитически вычислить проекцию
# - сходимость аналогична градиентному спуску в безусловной оптимизации
# - обобщается на негладкий случай - метод проекции субградиента
#
# Contra
# - при больших $n$ аналитическое вычисление проекции может быть слишком затратно: $O(n)$ для симплекса vs. решение задачи квадратичного программирования для полиэдрального множества
# - при обновлении градиента может теряться структура задачи: разреженность, малоранговость...
# + [markdown] slideshow={"slide_type": "slide"}
# ## Что такое "простое множество"?
# + [markdown] slideshow={"slide_type": "fragment"}
# - **Определение.** Множество $D$ будем называть *простым*, если можно найти решение следующей задачи
#
# $$
# \min_{x \in D} c^{\top}x
# $$
#
# существенно быстрее (чаще всего аналитически) по сравнению с
#
# решением исходной задачи минимизации.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Примеры простых множеств
#
# - Полиэдральное множество - задача линейного программирования вместо квадратичного программирования
# - Симплекс - $x^* = e_i$, где $c_i = \max\limits_{k = 1,\ldots, n} c_k$
# - Лоренцев конус - $x^* = -\frac{ct}{\| c\|_2}$
# - Все остальные множества из предыдущего определения
#
# - **Замечание 1:** отличие этого определения от предыдущего в линейности целевой функции (была квадратичная), поэтому простых множеств для этого определения больше.
#
# - **Замечание 2:** иногда на допустимое множество легко найти проекцию, но задача линейного программирования является неограниченной. Например, для множества
# $$
# D = \{ x \in \mathbb{R}^n \; | \; x_i \geq 0 \},
# $$
# проекция на которое очевидна, решение задачи линейного программирования равно $-\infty$, если есть хотя бы одна отрицательная компонента вектора $c$. Теорема с объяснением будет ниже.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Метод условного градиента <br> (aka Frank-Wolfe algorithm (1956))
# + [markdown] slideshow={"slide_type": "fragment"}
# $$
# \min_{x \in D} f(x)
# $$
#
# - **Идея**: делать шаг не по градиенту, а по направлению, которое точно не выведет из допустимого множества.
#
# Аналогия с градиентным спуском: линейная аппроксимация **на допустимом множестве**:
# $$
# f(x_k + s_k) = f(x_k) + \langle f'(x_k), s_k \rangle \to \min_{{\color{red}{s_k \in D}}}
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# ## Условный градиент
#
# **Определение** Направление $s_k - x_k$ называют *условным градиентом* функции $f$ в точке $x_k$ на допустимом множестве $D$.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Псевдокод
# ```python
# def FrankWolfe(f, gradf, linprogsolver, x0, tol):
# x = x0
#
# while True:
#
# gradient = gradf(x)
#
# s = linprogsolver(gradient)
#
# alpha = get_step_size(s, x, f)
#
# x = x + alpha * (s - x)
#
# if check_convergence(x, f, tol):
#
# break
#
#
# return x
# ```
# + [markdown] slideshow={"slide_type": "slide"}
# ## Выбор шага
#
# - Постоянный шаг: $\alpha_k = \alpha$
# - Убывающая последовательность, стандартный выбор $\alpha_k = \frac{2}{k + 2}$
# - Наискорейший спуск:
# $$
# \min_{{\color{red}{0 \leq \alpha_k \leq 1}}} f(x_k + \alpha_k(s_k - x_k))
# $$
# - Линейный поиск по правилу Армихо: должно выполняться условие
# $$
# f((x_k + \alpha_k(s_k - x_k)) \leq f(x_k) + c_1 \alpha_k \langle f'(x_k), s_k - x_k \rangle
# $$
# Начинать поиск нужно с $\alpha_k = 1$
# + [markdown] slideshow={"slide_type": "slide"}
# ## Критерий остановки
#
# - Так как показана сходимость к предельной точке $x^*$, то критерием остановки является сходимость по аргументу
# - Если $f(x)$ выпукла, то $f(s) \geq f(x_k) + \langle f'(x_k), s - x_k \rangle$ для любого вектора $s$, а значит и для любого $s \in D$. Следовательно
#
# $$
# f(x^*) \geq f(x) + \min_{s \in D} \langle f'(x), s - x\rangle
# $$
#
# или
#
# $$
# f(x) - f(x^*) \leq -\min_{s \in D} \langle f'(x), s - x\rangle = \max_{s \in D} \langle f'(x), x - s\rangle = g(x)
# $$
# Получили аналог зазора двойственности для контроля точности и устойчивости решения.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Аффинная инвариантность
#
# - Метод условного градиента является аффинно инвариантным относительно сюръективных отображений
# - Скорость сходимости и вид итерации не меняется
# + [markdown] slideshow={"slide_type": "slide"}
# ## Теорема сходимости
#
# Пусть $X$ - **выпуклый компакт** и $f(x)$ - дифференцируемая функция на $X$ с Липшицевым градиентом. Шаг выбирается по правилу Армихо. Тогда **для любого ${\color{red}{x_0 \in X}}$**
# - метод условного градиента генерирует последовательность $\{x_k\}$, которая имеет предельные точки
# - любая предельная точка $x^*$ является **стационарной**
# - если $f(x)$ выпукла на $X$, то $x^*$ - решение задачи
# + [markdown] slideshow={"slide_type": "slide"}
# ## Теоремы сходимости
#
# **Теорема (прямая).([Convex Optimization: Algorithms and Complexity, Th 3.8.](https://arxiv.org/abs/1405.4980))** Пусть $f$ выпуклая и дифференцируемая функция и её градиент Липшицев с константой $L$. Множество $X$ - выпуклый компакт диаметра $d > 0$.
# Тогда метод условного градиента с шагом $\alpha_k = \frac{2}{k + 1}$ сходится как
# $$
# f(x^*) - f(x_k) \leq \dfrac{2d^2L}{k + 2}, \quad k \geq 1
# $$
#
# **Теорема (двойственная) [см. эту статью](http://m8j.net/math/revisited-FW.pdf).** После выполнения $K$ итераций метода условного градиента для выпуклой и непрерывно дифференцируемой функции для функции $g$ и любого $k \leq K$ выполнено
# $$
# g(x_k) \leq \frac{2\beta C_f}{K+2} (1 + \delta),
# $$
# где $\beta \approx 3$, $\delta$ - точность решения промежуточных задач, $C_f$ - оценка кривизны $f$ на множестве $D$
# $$
# C_f = \sup_{x, s \in D; \gamma \in [0,1]} \frac{2}{\gamma^2}\left(f(x + \gamma(s - x)) - f(x) - \langle \gamma(s - x), f'(x)\rangle\right)
# $$
#
# Аргумент супремума так же известен как *дивергенция Брегмана*.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Как конструктивно задать "простые" множества?
# + [markdown] slideshow={"slide_type": "fragment"}
# **Определение**. Atomic norm называется следующая функция
#
# $$
# \|x\|_{\mathcal{D}} = \inf_{t \geq 0} \{ t | x \in t\mathcal{D} \}
# $$
#
# Она является нормой, если симметрична и $0 \in \mathrm{int}(\mathcal{D})$
# + [markdown] slideshow={"slide_type": "slide"}
# ### Сопряжённая atomic norm
# $$
# \|y\|^*_{\mathcal{D}} = \sup_{s \in \mathcal{D}} \langle s, y \rangle
# $$
#
# - Из определения выпуклой оболочки следует, что линейная функция достигает максимума в одной из "вершин" выпуклого множества
# - Следовательно, $\| y \|^*_{\mathcal{D}} = \| y \|^*_{\mathrm{conv}(\mathcal{D})}$
# - Это позволяет эффективно вычислять решение промежуточных задач для определения $s$
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="atomic_table.png">
#
# Таблица взята из [статьи](http://m8j.net/math/revisited-FW.pdf)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Разреженность vs. точность
#
# - Метод условного градиента на каждой итерации добьавляет к решению слагаемое, являющееся элементом множества $\mathcal{A}$
# - Решение моджет быть представлено в виде комбинации элементов $\mathcal{A}$
# - Теорема Каратеодори
# - Число элементов может быть существенно маньше требуемого теоремой Каратеодори
# + [markdown] slideshow={"slide_type": "slide"}
# ## Эксперименты
# + [markdown] slideshow={"slide_type": "slide"}
# ## Пример 1
#
# \begin{equation*}
# \begin{split}
# & \min \frac{1}{2}\|Ax - b \|^2_2\\
# \text{s.t. } & 0 \leq x_i \leq 1
# \end{split}
# \end{equation*}
# + slideshow={"slide_type": "slide"}
def func(x, A, b):
return 0.5 * np.linalg.norm(A.dot(x) - b)**2
f = lambda x: func(x, A, b)
def grad_f(x, A, b):
grad = -A.T.dot(b)
grad = grad + A.T.dot(A.dot(x))
return grad
grad = lambda x: grad_f(x, A, b)
# + slideshow={"slide_type": "slide"}
def linsolver(gradient):
x = np.zeros(gradient.shape[0])
pos_grad = gradient > 0
neg_grad = gradient < 0
x[pos_grad] = np.zeros(np.sum(pos_grad == True))
x[neg_grad] = np.ones(np.sum(neg_grad == True))
return x
# + slideshow={"slide_type": "slide"}
def projection(y):
return np.clip(y, 0, 1)
# + slideshow={"slide_type": "slide"}
import liboptpy.constr_solvers as cs
import liboptpy.step_size as ss
import numpy as np
from tqdm import tqdm
n = 200
m = 100
A = np.random.randn(m, n)
x_true = np.random.rand(n)
b = A.dot(x_true) + 0.01 * np.random.randn(m)
# + slideshow={"slide_type": "slide"}
def myplot(x, y, xlab, ylab, xscale="linear", yscale="log"):
plt.figure(figsize=(10, 8))
plt.xscale(xscale)
plt.yscale(yscale)
for key in y:
plt.plot(x[key], y[key], label=key)
plt.xticks(fontsize=24)
plt.yticks(fontsize=24)
plt.legend(loc="best", fontsize=24)
plt.xlabel(xlab, fontsize=24)
plt.ylabel(ylab, fontsize=24)
# + slideshow={"slide_type": "slide"}
x0 = np.random.rand(n)
cg = cs.FrankWolfe(f, grad, linsolver, ss.Backtracking(rule_type="Armijo", rho=0.5, beta=0.1, init_alpha=1.))
x_cg = cg.solve(x0=x0, max_iter=200, tol=1e-10, disp=1)
print("Optimal value CG =", f(x_cg))
# + slideshow={"slide_type": "slide"}
pg = cs.ProjectedGD(f, grad, projection, ss.Backtracking(rule_type="Armijo", rho=0.5, beta=0.1, init_alpha=1.))
x_pg = pg.solve(x0=x0, max_iter=200, tol=1e-10, disp=1)
print("Optimal value PG =", f(x_pg))
# + slideshow={"slide_type": "slide"}
import matplotlib.pyplot as plt
# %matplotlib inline
plt.rc("text", usetex=True)
y_hist_f_cg = [f(x) for x in cg.get_convergence()]
y_hist_f_pg = [f(x) for x in pg.get_convergence()]
myplot({"CG": range(1, len(y_hist_f_cg) + 1), "PG": range(1, len(y_hist_f_pg) + 1)},
{"CG": y_hist_f_cg, "PG": y_hist_f_pg}, "Number of iteration",
r"Objective function, $\frac{1}{2}\|Ax - b\|^2_2$")
# + slideshow={"slide_type": "slide"}
import cvxpy as cvx
x = cvx.Variable(n)
obj = cvx.Minimize(0.5 * cvx.norm(A * x - b, 2)**2)
constr = [x >= 0, x <= 1]
problem = cvx.Problem(objective=obj, constraints=constr)
value = problem.solve()
x_cvx = np.array(x.value).ravel()
print("CVX optimal value =", value)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Зависимость времени и числа итераций от точности
# + slideshow={"slide_type": "slide"}
eps = [10**(-i) for i in range(8)]
time_pg = np.zeros(len(eps))
time_cg = np.zeros(len(eps))
iter_pg = np.zeros(len(eps))
iter_cg = np.zeros(len(eps))
pg = cs.ProjectedGD(f, grad, projection, ss.Backtracking(rule_type="Armijo", rho=0.5, beta=0.1, init_alpha=1.))
cg = cs.FrankWolfe(f, grad, linsolver, ss.Backtracking(rule_type="Armijo", rho=0.5, beta=0.1, init_alpha=1.))
for i, tol in tqdm(enumerate(eps)):
# res = %timeit -o -q pg.solve(x0=x0, tol=tol, max_iter=100000)
time_pg[i] = res.average
iter_pg[i] = len(pg.get_convergence())
# res = %timeit -o -q cg.solve(x0=x0, tol=tol, max_iter=100000)
time_cg[i] = res.average
iter_cg[i] = len(cg.get_convergence())
# + slideshow={"slide_type": "slide"}
myplot({"CG":eps, "PG": eps}, {"CG": time_cg, "PG": time_pg}, r"Accuracy, $\varepsilon$", "Time, s", xscale="log")
# + slideshow={"slide_type": "slide"}
myplot({"CG":eps, "PG": eps}, {"CG": iter_cg, "PG": iter_pg}, r"Accuracy, $\varepsilon$", "Number of iterations", xscale="log")
# + [markdown] slideshow={"slide_type": "slide"}
# ## Пример 2
# Рассмотрим задачу:
# \begin{equation*}
# \begin{split}
# & \min \frac{1}{2}\|Ax - b \|^2_2 \\
# \text{s.t. } & \| x\|_1 \leq 1 \\
# & x_i \geq 0
# \end{split}
# \end{equation*}
# + slideshow={"slide_type": "slide"}
def linsolver(gradient):
x = np.zeros(gradient.shape[0])
idx_min = np.argmin(gradient)
if gradient[idx_min] > 0:
x[idx_min] = 0
else:
x[idx_min] = 1
return x
# + slideshow={"slide_type": "slide"}
def projection(y):
x = y.copy()
if np.all(x >= 0) and np.sum(x) <= 1:
return x
x = np.clip(x, 0, np.max(x))
if np.sum(x) <= 1:
return x
n = x.shape[0]
bget = False
x.sort()
x = x[::-1]
temp_sum = 0
t_hat = 0
for i in range(n - 1):
temp_sum += x[i]
t_hat = (temp_sum - 1.0) / (i + 1)
if t_hat >= x[i + 1]:
bget = True
break
if not bget:
t_hat = (temp_sum + x[n - 1] - 1.0) / n
return np.maximum(y - t_hat, 0)
# + slideshow={"slide_type": "slide"}
x0 = np.random.rand(n) * 10
x0 = x0 / x0.sum()
cg = cs.FrankWolfe(f, grad, linsolver, ss.Backtracking(rule_type="Armijo", rho=0.5, beta=0.1, init_alpha=1.))
x_cg = cg.solve(x0=x0, max_iter=200, tol=1e-10)
print("Optimal value CG =", f(x_cg))
# + slideshow={"slide_type": "slide"}
pg = cs.ProjectedGD(f, grad, projection, ss.Backtracking(rule_type="Armijo", rho=0.5, beta=0.1, init_alpha=1.))
x_pg = pg.solve(x0=x0, max_iter=200, tol=1e-10)
print("Optimal value PG =", f(x_pg))
# -
y_hist_f_cg = [f(x) for x in cg.get_convergence()]
y_hist_f_pg = [f(x) for x in pg.get_convergence()]
myplot({"CG": range(1, len(y_hist_f_cg) + 1), "PG": range(1, len(y_hist_f_pg) + 1)},
{"CG": y_hist_f_cg, "PG": y_hist_f_pg}, "Number of iteration",
r"Objective function, $\frac{1}{2}\|Ax - b\|^2_2$")
# + [markdown] slideshow={"slide_type": "slide"}
# ### Зависимость времени и числа итераций от точности
# + slideshow={"slide_type": "slide"}
eps = [10**(-i) for i in range(8)]
time_pg = np.zeros(len(eps))
time_cg = np.zeros(len(eps))
iter_pg = np.zeros(len(eps))
iter_cg = np.zeros(len(eps))
pg = cs.ProjectedGD(f, grad, projection, ss.Backtracking(rule_type="Armijo", rho=0.5, beta=0.1, init_alpha=1.))
cg = cs.FrankWolfe(f, grad, linsolver, ss.Backtracking(rule_type="Armijo", rho=0.5, beta=0.1, init_alpha=1.))
for i, tol in tqdm(enumerate(eps)):
# res = %timeit -o -q pg.solve(x0=x0, tol=tol, max_iter=100000)
time_pg[i] = res.average
iter_pg[i] = len(pg.get_convergence())
# res = %timeit -o -q cg.solve(x0=x0, tol=tol, max_iter=100000)
time_cg[i] = res.average
iter_cg[i] = len(cg.get_convergence())
# + slideshow={"slide_type": "slide"}
myplot({"CG":eps, "PG": eps}, {"CG": time_cg, "PG": time_pg},
r"Accuracy, $\varepsilon$", "Time, s", xscale="log")
# -
myplot({"CG":eps, "PG": eps}, {"CG": iter_cg, "PG": iter_pg},
r"Accuracy, $\varepsilon$", "Number of iterations", xscale="log")
# + slideshow={"slide_type": "slide"}
x = cvx.Variable(n)
obj = cvx.Minimize(0.5 * cvx.norm2(A * x - b)**2)
constr = [cvx.norm(x, 1) <= 1, x >= 0]
problem = cvx.Problem(objective=obj, constraints=constr)
value = problem.solve()
x_cvx = np.array(x.value).ravel()
print("CVX optimal value =", value)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Pro & Contra
#
# Pro
# - Оценка сходимости для функционала **не зависит** от размерности
# - Если множество - многоугольник, то $x_k$ - выпуклая комбинация $k$ вершин многоугольника - разреженная решение для $k \ll n$
# - Если множество выпуклая комбинация некоторых элементов, то решение - линейная комбинация подмножества этих элементов
# - Сходимость по функционалу не улучшаема даже для сильно выпуклых функций
# - Упрощение понятия "простое множество"
# - Существует подобие зазора двойственности и теоретические результаты о сходимости
#
# Contra
# - Сходимость по функционалу только сублинейная вида $\frac{C}{k}$
# - Не обобщается на негладкие задачи
# + [markdown] slideshow={"slide_type": "slide"}
# ## Резюме
# - Множество простой структуры
# - Проекция
# - Метод проекции градиента
# - Метод условного градиента
| 10-SimpleSets/Seminar10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Intel, 2019 update 1)
# language: python
# name: c009-intel_distribution_of_python_3_2019u1
# ---
# cd ~/fyp/code/
# +
import sys
sys.path.append('./log_helper')
sys.path.append('./model')
# -
from rl_train import run_rl_backtest
# Some more magic so that the notebook will reload external python modules;
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
# %load_ext autoreload
# %autoreload 2
df1, df2 = run_rl_backtest('AER', 'AAN', 3)
# df1, df2 = run_rl_backtest('NEWR', 'TYL', 3)
| model/evaluate_a_pair.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Dependencies
# %matplotlib notebook
from matplotlib import pyplot as plt
from scipy.stats import linregress
import numpy as np
from sklearn import datasets
import pandas as pd
import os
# # Dataset: winequality-red.csv
# Linear Regression - The predicted affect of 'pH' on red wine 'quality.
#
# **Source:**
# UCI Machine Learning Repository. https://archive.ics.uci.edu/ml/datasets/Wine+Quality
# Data source of origin: https://archive.ics.uci.edu/ml/machine-learning-databases/wine-quality/
# Link: https://archive.ics.uci.edu/ml/machine-learning-databases/wine-quality/winequality-red.csv
#
# **Description:**
# Winemakers use pH as a way to measure ripeness in relation to acidity. Low pH wines will taste tart and crisp, while higher pH wines are more susceptible to bacterial growth. Most wine pH's fall around 3 or 4; about 3.0 to 3.4 is desirable for white wines, while about 3.3 to 3.6 is best for reds.
#
# **Linear Regression:**
# A regression line is simply calculating a line that best fits the data. This is typically done through the least squares method where the line is chosen to have the smallest overall distance to the points.
# 𝑦=𝜃0+𝜃1𝑥y=θ0+θ1x
# • 𝑦y is the output response
# • 𝑥x is the input feature
# • 𝜃0θ0 is the y-axis intercept
# • 𝜃1θ1 is weight coefficient (slope)
#
# **Variables/Columns:**
# All Attribute Information:
#
# Independant Input variables (x) (based on physicochemical tests): 1 - fixed acidity 2 - volatile acidity 3 - citric acid 4 - residual sugar 5 - chlorides 6 - free sulfur dioxide 7 - total sulfur dioxide 8 - density 9 - pH 10 - sulphates 11 - alcohol
#
# Dependant Output variable (y) (based on sensory data): 12 - quality (score between 0 and 10)
#
# **Story - How do the physicochemical factors affect overall red wine quality?**
#
# *In this example we predicted affect of 'pH' on red wine 'quality'.*
#
# **Hypothesis**:
#
# Linear Regression: r-squared is: 0.22673436811123157
#
# **Conclusion**
# A good R2 Score will be close to 1. The above results individually the model does not predict a solid or good relationship between this factor and quality.
#
# Overall the individual factors (features) do not individually prove to affect the quality based on their scores. When we view the 'Combined' features they have a more high training and test score. This concludes that the quality is more a factor of the overall physicochemical mixture (formula) of factors than any one individual factors (features).
# This example compares different factors in the wine quality file
df = pd.read_csv(os.path.join('winequality-red.csv'))
df.head()
y = df["quality"]
y
X = df.drop("quality", axis=1)
X.head()
print(f"Labels: {y[:10]}")
print(f"Data: {X[:10]}")
X_arr = X.to_numpy()
X_arr
y_arr = y.to_numpy()
y_arr
X_arr[:,8]
# Plot out rooms versus median house price
x_arr = df['alcohol']
y_arr = df['quality']
plt.scatter(x_arr,y_arr)
plt.xlabel('Alcohol')
plt.ylabel('Quality')
plt.show()
# Add the linear regression equation and line to plot
x_arr = df['alcohol']
y_arr = df['quality']
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_arr, y_arr)
regress_values = x_arr * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_arr,y_arr)
plt.plot(x_arr,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('Alcohol')
plt.ylabel('Quality')
plt.show()
# Print out the r-squared value along with the plot.
x_arr = df['alcohol']
y_arr = df['quality']
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_arr, y_arr)
regress_values = x_arr * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_arr,y_arr)
plt.plot(x_arr,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
plt.xlabel('Alcohol')
plt.ylabel('Quality')
print(f"The r-squared is: {rvalue**2}")
plt.show()
| assets/linear-regression-wine-quality-pH.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import seaborn as sbn
datos = pd.read_csv('newdata.csv')
datos.iloc[0:3,0:4]
sbn.scatterplot(data=datos, x='Number of Comments', y='Cost Delivery')
sbn.scatterplot(data=datos, x='Number of Comments', y='Cost Delivery', hue='Moment')
datos[['Cost Delivery','Time(min)']].plot.kde()
# Observe que, aunque con la sintaxis anterior logramos sacar un gráfico de densidad para la variable "Time(min)" y la variable "Cost Delivery", dicho gráfico no tiene sentido. Ello es porque el gráfico está comparando dos variables que hacen referencia a cosas distintas (algo así como comparar peras con manzanas). La moraleja de este asunto es que en la analítica de datos aún cuando seamos capaces de lograr que la computadora nos obedezca y haga algo que le pidamos, el resultado no necesariamente tiene porque tener sentido lógico con la realidad.
| ScatterplotSeaborn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="fRxS371VhOp-" colab={"base_uri": "https://localhost:8080/", "height": 426} outputId="a469a08c-6de0-4223-9cb8-9f12870e8f78"
import pandas as pd
df = pd.read_excel('https://github.com/dayanandv/Data-Science/raw/main/dataset/survey.xls')
df
# + [markdown] id="Sfsz3HUY6qhd"
# # CrossTab
# + [markdown] id="fVjpcqfU6ZuI"
# ## Simple
# + id="Z-5fSsDO6ZOL" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="69990c51-a1ac-44bb-8620-542c0c64d371"
pd.crosstab(df.Nationality, df.Handedness)
# + id="RJ8eO_-R54r-" colab={"base_uri": "https://localhost:8080/", "height": 143} outputId="77351886-eff9-4b4d-eb45-136cc2a617bc"
pd.crosstab(df.Sex, df.Handedness)
# + [markdown] id="DLTmDAyw6dko"
# ## With Margins
# + id="GTwizSAQ6ir6" colab={"base_uri": "https://localhost:8080/", "height": 175} outputId="1e017e78-80e5-4ded-d912-cf3a2698ee29"
pd.crosstab(df.Sex, df.Handedness, margins=True)
# + [markdown] id="ZH13jjdD6wvA"
# ## Multi-Index Column and Rows
# + id="4O7Ut24a60Z1" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="cbc0bba4-d3c9-4646-e98b-231ae13ead2c"
pd.crosstab(df.Sex, [df.Handedness, df.Nationality], margins=True)
# + [markdown] id="_XULaxZj62Oq"
# ## Normalize
# + colab={"base_uri": "https://localhost:8080/", "height": 143} id="z0DN41iyVYXz" outputId="5bfaf957-16ac-4eb8-e6d8-c3e5698a0936"
pd.crosstab(df.Sex, df.Handedness, normalize='index')
# + [markdown] id="S_jlaCpf64Y0"
# ## Aggregate function
# + id="ufcWU_1D63Za" colab={"base_uri": "https://localhost:8080/", "height": 143} outputId="0d488fb8-ce73-48cd-ef57-afa856aa0c0c"
import numpy as np
pd.crosstab(df.Sex, df.Handedness, values=df.Age, aggfunc=np.average)
# + [markdown] id="cQ5HJZIP7r9s"
# # Automotive dataset example
# + [markdown] id="_oBffT2m9g0W"
# Define the headers since the data does not have any
# + id="e6mkT4oc8WQf"
headers = ["symboling", "normalized_losses", "make", "fuel_type", "aspiration","num_doors", "body_style", "drive_wheels",
"engine_location", "wheel_base", "length", "width", "height", "curb_weight", "engine_type", "num_cylinders",
"engine_size", "fuel_system", "bore", "stroke", "compression_ratio", "horsepower", "peak_rpm", "city_mpg",
"highway_mpg", "price"]
# + [markdown] id="Cvb3vXR89j5u"
# Read in the CSV file and convert "?" to NaN
# + id="so59u6ge9a8Y"
df_raw = pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/autos/imports-85.data', header=None, names=headers, na_values="?" )
# + [markdown] id="FkDO-SbI9qeN"
# Define a list of models that we want to review
# + id="f1vg6hWm9dh5"
models = ["toyota","nissan","mazda", "honda", "mitsubishi", "subaru", "volkswagen", "volvo"]
# + [markdown] id="hcEjQgLL9t-C"
# Create a copy of the data with only the top 8 manufacturers
# + id="nbBRNjmc9w4L"
df = df_raw[df_raw.make.isin(models)].copy()
# + [markdown] id="G2jFHPHC9xhI"
# CrossTab: make vs body_style
# + id="lGjp9jIf932T" colab={"base_uri": "https://localhost:8080/", "height": 332} outputId="28892111-eebc-43a6-b905-becbbea1f77e"
pd.crosstab(df.make, df.body_style)
# + [markdown] id="OAZnf5rQ95hn"
# Groupby
# + id="ItIVYYEF96fI" colab={"base_uri": "https://localhost:8080/", "height": 332} outputId="072cdaef-8187-4aef-8572-b2c80cc60346"
df.groupby(['make', 'body_style'])['body_style'].count().unstack().fillna(0)
# + [markdown] id="8-QwPoLK-3SS"
# Pivot table
# + id="Uerk10bV-5HC" colab={"base_uri": "https://localhost:8080/", "height": 363} outputId="0efdef6b-bdbb-49d8-f277-8ac014fb0fec"
df.pivot_table(index='make' , columns= 'body_style' , aggfunc={ 'body_style' :len}, fill_value=0)
# + [markdown] id="nU6fWoTe_Sny"
# Crosstab: make vs num_doors
# + id="iGxcT_uT_D7a" colab={"base_uri": "https://localhost:8080/", "height": 363} outputId="b958d032-8d84-4d03-de21-6ae3fffd895b"
pd.crosstab(df.make, df.num_doors, margins=True, margins_name="Total")
# + [markdown] id="CwE5foIh_u64"
# Crosstab: Multi-index
# + id="2_hmj5Nw_fCh" colab={"base_uri": "https://localhost:8080/", "height": 363} outputId="972b70af-7a7b-424d-c237-9cf5a631b53e"
pd.crosstab(df.make, [df.body_style, df.drive_wheels])
# + [markdown] id="hUVukVZYAHfQ"
# Crosstab: Normalize
# + id="Iqa86MIoAEbV" colab={"base_uri": "https://localhost:8080/", "height": 614} outputId="830ad1f8-76e9-401e-bb11-d54b5b41bea5"
pd.crosstab([df.make, df.num_doors], [df.body_style, df.drive_wheels], rownames=['Auto Manufacturer', "Doors"],
colnames=['Body Style', "Drive Type"], dropna=False)
# + [markdown] id="z_8p4fd2AoXC"
# A combination
# + id="Ro-gjUJbAM2G" colab={"base_uri": "https://localhost:8080/", "height": 363} outputId="56c16cda-6312-473d-882a-68fb6bbaa890"
pd.crosstab(df.make, [df.body_style, df.drive_wheels], values=df.curb_weight, aggfunc='mean').fillna('-')
# + [markdown] id="FHHDtQzDBHuB"
# ## Normalization
# + [markdown] id="kXM73zjUBPqk"
# All
# + id="d99um4NyA6At" colab={"base_uri": "https://localhost:8080/", "height": 332} outputId="535c4d25-7fe8-4f2d-eed4-1a100a0fe174"
pd.crosstab(df.make, df.body_style, normalize=True)
# + [markdown] id="p44sSjkxBXf1"
# Rows
# + id="cJgoGFSOBWFl" colab={"base_uri": "https://localhost:8080/", "height": 332} outputId="36aee484-9db4-43ec-ef7a-586c9cb7428b"
pd.crosstab(df.make, df.body_style, normalize='index')
# + [markdown] id="1XbFwL-5Bde8"
# Columns
# + id="OoKsm0ETBcOF" colab={"base_uri": "https://localhost:8080/", "height": 332} outputId="20a6ebdb-06cf-4a63-de80-15beb88cbb6d"
pd.crosstab(df.make, df.body_style, normalize='columns')
| Crosstab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
class Solution(object):
def licenseKeyFormatting(self, S, K):
"""
:type S: str
:type K: int
:rtype: str
"""
S = S.split('-')
S = ''.join(S).upper()
ret = ''
n = len(S)
while n > K:
ret = S[n-K:n] + '-' + ret
n -= K
if n > 0:
ret = S[0:n] + '-' + ret
return ret[:-1]
s = Solution()
s.licenseKeyFormatting("<KEY>",2)
| algorithms/482-license-key-formatting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
diamond =pd.read_csv('diamonds.csv')
diamond.head()
diamond.shape
diamond.iloc[:4,1]
diamond.loc[2, ['cut' , 'color']]
pd.crosstab(diamond['color'] , diamond['cut'])
diamond.groupby(['color'])[['cut' , 'clarity']].first()
pd.pivot_table(['color'])
| pandas/modern pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # How to test GACELA on your data
# ## First, set up the parameters of the network you trained or downloaded from our pre-trained networks
# +
import torch
from data.audioLoader import AudioLoader
from data.trainDataset import TrainDataset
from ganSystem import GANSystem
import logging
# logging.getLogger().setLevel(logging.DEBUG) # set root logger to debug
"""Just so logging works..."""
formatter = logging.Formatter('%(name)s:%(levelname)s:%(message)s')
console_handler = logging.StreamHandler()
console_handler.setLevel(logging.DEBUG)
console_handler.setFormatter(formatter)
logging.getLogger().addHandler(console_handler)
"""Just so logging works..."""
__author__ = 'Andres'
signal_split = [480, 64, 480]
md = 32
params_stft_discriminator = dict()
params_stft_discriminator['stride'] = [2, 2, 2, 2, 2]
params_stft_discriminator['nfilter'] = [md, 2 * md, 4 * md, 8 * md, 16 * md]
params_stft_discriminator['shape'] = [[5, 5], [5, 5], [5, 5], [5, 5], [5, 5]]
params_stft_discriminator['data_size'] = 2
params_mel_discriminator = dict()
params_mel_discriminator['stride'] = [2, 2, 2, 2, 2]
params_mel_discriminator['nfilter'] = [md//4, 2 * md//4, 4 * md//4, 8 * md//4, 16 * md//4]
params_mel_discriminator['shape'] = [[5, 5], [5, 5], [5, 5], [5, 5], [5, 5]]
params_mel_discriminator['data_size'] = 2
params_generator = dict()
params_generator['stride'] = [2, 2, 2, 2, 2]
params_generator['nfilter'] = [8 * md, 4 * md, 2 * md, md, 1]
params_generator['shape'] = [[4, 4], [4, 4], [8, 8], [8, 8], [8, 8]]
params_generator['padding'] = [[1, 1], [1, 1], [3, 3], [3, 3], [3, 3]]
params_generator['residual_blocks'] = 2
params_generator['full'] = 256 * md
params_generator['summary'] = True
params_generator['data_size'] = 2
params_generator['in_conv_shape'] = [16, 2]
params_generator['borders'] = dict()
params_generator['borders']['nfilter'] = [md, 2 * md, md, md / 2]
params_generator['borders']['shape'] = [[5, 5], [5, 5], [5, 5], [5, 5]]
params_generator['borders']['stride'] = [2, 2, 2, 2]
params_generator['borders']['data_size'] = 2
params_generator['borders']['border_scale'] = 1
# This does not work because of flipping, border 2 need to be flipped tf.reverse(l, axis=[1]), ask Nathanael
params_generator['borders']['width_full'] = None
# Optimization parameters inspired from 'Self-Attention Generative Adversarial Networks'
# - Spectral normalization GEN DISC
# - Batch norm GEN
# - TTUR ('GANs Trained by a Two Time-Scale Update Rule Converge to a Local Nash Equilibrium')
# - ADAM beta1=0 beta2=0.9, disc lr 0.0004, gen lr 0.0001
# - Hinge loss
# Parameters are similar to the ones in those papers...
# - 'PROGRESSIVE GROWING OF GANS FOR IMPROVED QUALITY, STABILITY, AND VARIATION'
# - 'LARGE SCALE GAN TRAINING FOR HIGH FIDELITY NATURAL IMAGE SYNTHESIS'
# - 'CGANS WITH PROJECTION DISCRIMINATOR'
params_optimization = dict()
params_optimization['batch_size'] = 64
params_stft_discriminator['batch_size'] = 64
params_mel_discriminator['batch_size'] = 64
params_optimization['n_critic'] = 1
params_optimization['generator'] = dict()
params_optimization['generator']['optimizer'] = 'adam'
params_optimization['generator']['kwargs'] = [0.5, 0.9]
params_optimization['generator']['learning_rate'] = 1e-4
params_optimization['discriminator'] = dict()
params_optimization['discriminator']['optimizer'] = 'adam'
params_optimization['discriminator']['kwargs'] = [0.5, 0.9]
params_optimization['discriminator']['learning_rate'] = 1e-4
# all parameters
params = dict()
params['net'] = dict() # All the parameters for the model
params['net']['generator'] = params_generator
params['net']['stft_discriminator'] = params_stft_discriminator
params['net']['mel_discriminator'] = params_mel_discriminator
params['net']['prior_distribution'] = 'gaussian'
params['net']['shape'] = [1, 512, 1024] # Shape of the image
params['net']['inpainting'] = dict()
params['net']['inpainting']['split'] = signal_split
params['net']['gamma_gp'] = 10 # Gradient penalty
# params['net']['fs'] = 16000//downscale
params['net']['loss_type'] = 'wasserstein'
params['optimization'] = params_optimization
params['summary_every'] = 250 # Tensorboard summaries every ** iterations
params['print_every'] = 50 # Console summaries every ** iterations
params['save_every'] = 1000 # Save the model every ** iterations
# params['summary_dir'] = os.path.join(global_path, name +'_summary/')
# params['save_dir'] = os.path.join(global_path, name + '_checkpoints/')
args = dict()
args['generator'] = params_generator
args['stft_discriminator_count'] = 2
args['mel_discriminator_count'] = 3
args['stft_discriminator'] = params_stft_discriminator
args['mel_discriminator'] = params_mel_discriminator
args['borderEncoder'] = params_generator['borders']
args['stft_discriminator_in_shape'] = [1, 512, 64]
args['mel_discriminator_in_shape'] = [1, 80, 64]
args['mel_discriminator_start_powscale'] = 2
args['generator_input'] = 1440
args['optimizer'] = params_optimization
args['split'] = signal_split
args['log_interval'] = 100
args['spectrogram_shape'] = params['net']['shape']
args['gamma_gp'] = params['net']['gamma_gp']
args['tensorboard_interval'] = 500
args['save_path'] = 'saved_results/'
args['experiment_name'] = 'fma_rock'
args['save_interval'] = 10000
args['fft_length'] = 1024
args['fft_hop_size'] = 256
args['sampling_rate'] = 22050
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
examples_per_file = 32
audioLoader = AudioLoader(args['sampling_rate'], args['fft_length'], args['fft_hop_size'], 50)
ganSystem = GANSystem(args)
# -
# ## Load the trained model
# +
start_at_step = 410000
start_at_epoch = 0
ganSystem.loadModel(start_at_step, start_at_epoch)
# -
# ## Load the data you want to inpaint and process the spectrograms
# +
from pathlib import Path
from data.audioLoader import AudioLoader
import numpy as np
audioLoader = AudioLoader(22050, 1024, 256, 50)
dataFolder = "rock/"
filenames = Path(dataFolder).rglob('*.mp3')
files = {}
for filename in filenames:
audio = audioLoader.loadSound(filename)
files[filename] = (audio, audioLoader.computeSpectrogram(audio))
audios = np.zeros([64, 256*1024])
spectrograms = np.zeros([64, 1, 512, 1024])
filenames = list(files.keys())
for index in range(64):
filename = np.random.choice(filenames)
audio = files[filename][0]
start = int((len(audio)-1024*256) * np.random.rand())
audios[index] = files[filename][0][start:start+1024*256]
spectrograms[index, 0] = files[filename][1][:-1, int(start/256):int(start/256)+1024]
# -
# ## Generate the gap with GACELA and concatenate it with the borders
# +
left_borders = torch.from_numpy(spectrograms[:, :, :, :args['split'][0]]).float().to(device)
right_borders = torch.from_numpy(spectrograms[:, :, :, args['split'][0] + args['split'][1]:]).float().to(device)
print('generate')
generated_spectrograms = ganSystem.generateGap([left_borders, right_borders])
fake_spectrograms = torch.cat((left_borders, generated_spectrograms, right_borders), 3)
# -
# ## Plot some of the results
# +
import matplotlib.pyplot as plt
plt.figure(figsize=(30, 14))
for i in range(8):
plt.subplot(2, 4, i+1)
plt.imshow(fake_spectrograms[3*i, 0, :, 200:-200].detach().cpu())
# -
# ## Invert the generated spectrograms using PGHI
# +
from tifresi.stft import GaussTruncTF, GaussTF
from tifresi.transforms import inv_log_spectrogram, log_spectrogram
import numpy as np
stft = GaussTruncTF(256, 1024)
fake_audios = np.zeros([len(audios), fake_spectrograms.shape[-1]*args['fft_hop_size']])
for index, (real_audio, spectrogram) in enumerate(zip(audios, fake_spectrograms)):
unprocessed_spectrogram = inv_log_spectrogram((spectrogram-1)*25).squeeze().detach().cpu().numpy()
unprocessed_spectrogram = np.concatenate([unprocessed_spectrogram,
np.ones_like(unprocessed_spectrogram)[0:1, :]*unprocessed_spectrogram.min()], axis=0) # Fill last column of freqs with zeros
audio = stft.invert_spectrogram(unprocessed_spectrogram)
fake_audios[index] = audio
# -
# ## Listen to the results
# +
from IPython.display import display, Audio
for generated_audio_signal in fake_audios:
display(Audio(generated_audio_signal[int(len(generated_audio_signal)*0.3):], rate=22050))
# -
# # Prepare and save the generated data
# +
import os
import numpy as np
import librosa
from tifresi.phase.modGabPhaseGrad import modgabphasegrad
from tifresi.phase.pghi_masked import pghi
from tifresi.stft import GaussTruncTF, GaussTF
hop_size = 256
stft_channels = 1024
stft = GaussTruncTF(256, 1024)
base_folder_name = "my_test/"
os.mkdir(base_folder_name)
real_folder = base_folder_name + 'real/'
fake_folder = base_folder_name + "GACELA/"
os.mkdir(real_folder)
os.mkdir(fake_folder)
for index, (real_audio, fake_audio) in enumerate(zip(audios, fake_audios)):
#Real
librosa.output.write_wav(real_folder + str(index) + '.wav', real_audio, sr=22050)
#Fake
librosa.output.write_wav(fake_folder + str(index) + '.wav', fake_audio, sr=22050)
# -
| Test GACELA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Python statistics essential training - 02_03_pandas
# Standard imports
# +
import numpy as np
import pandas as pd
import matplotlib
import matplotlib.pyplot as pp
# %matplotlib inline
# -
planets = pd.read_csv('Planets.csv')
planets
planets = pd.read_csv('Planets.csv',usecols=[0,1,2,3])
planets
planets['Mass']
planets.Mass
planets.index
planets.loc[0]
planets.set_index('Planet',inplace=True)
planets
planets.info()
len(planets)
planets.loc['MERCURY']
planets.loc['MERCURY':'EARTH']
planets.columns
planets = pd.read_csv('Planets.csv')
planets.set_index('Planet',inplace=True)
planets.FirstVisited['MERCURY']
planets.loc['MERCURY'].FirstVisited
planets.loc['MERCURY','FirstVisited']
type(planets.loc['MERCURY','FirstVisited'])
pd.to_datetime(planets.FirstVisited)
planets.FirstVisited = pd.to_datetime(planets.FirstVisited)
planets.FirstVisited.dt.year
2020 - planets.FirstVisited.dt.year
| Statistics Python Essential Training - Linkedin Learning/1). Importing and Cleaning Data/02_03/02_03_pandas_end.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from mllib.ensemble import RandomSeedAveragingRegressor
from mllib.model_selection import TPESearchCV
from mllib.utils import get_param_distributions
from optuna.distributions import *
from sklearn.datasets import load_boston
from sklearn.ensemble import RandomForestRegressor
from sklearn.externals.joblib import dump
from sklearn.model_selection import train_test_split
from yellowbrick.regressor import ResidualsPlot
# +
# TODO: use a dataset containing missing values
# TODO: use a dataset containing outliers
# TODO: use a dataset with mixed types of attributes
# TODO: plot rank features with yellowbrick
# TODO: plot a feature correlation with yellowbrick
# TODO: plot feature importances with yellowbrick
# -
rf = RandomForestRegressor(n_estimators=100, random_state=0)
param_distributions = get_param_distributions(rf.__class__.__name__)
tpe_search = TPESearchCV(
rf,
param_distributions,
n_jobs=-1,
random_state=0,
refit=False,
scoring='neg_mean_squared_error'
)
boston = load_boston()
feature_names = boston.feature_names
X = boston.data
y = boston.target
X_train, X_test, y_train, y_test = train_test_split(
X,
y,
random_state=0
)
X_train.shape
X_test.shape
tpe_search.fit(X_train, y_train)
tpe_search.trials_dataframe()
rf.set_params(**tpe_search.best_params_)
reg = RandomSeedAveragingRegressor(rf, random_state=0)
# +
viz = ResidualsPlot(reg)
viz.fit(X_train, y_train)
viz.score(X_test, y_test)
viz.poof(outpath='residuals_plot.png')
# -
dump(reg, 'regressor.pkl', compress=True)
| notebooks/tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # EJERCICIO 9
# La segmentación de imágenes digitales en regiones con apariencia de piel ha sido utilizado como un paso de pre-procesamiento en varias aplicaciones de detección y seguimiento de caras humanas y personas.
# Con el fin de generar un detector de piel/no-piel, se realizó un muestreo de fotografías digitales de caras de personas de diversas edades, géneros y razas al mismo tiempo que se hizo otro muestreo de imágenes que no incluyeran personas. El muestreo consistió en obtener mediciones del color de pixels de cada una de las clases requeridas (piel/no-piel). La medición del color se realizó en el espacio de colores RGB, obteniendo un total de 51444 muestras de las cuales 14654 son muestras de piel mientras que las restantes 36790 son muestras de otras texturas. Cada muestra está representada por el nivel de azul (B), verde (G) y rojo (R), siendo el nivel un valor entero entre 0 (mínimo nivel posible) y 255 (máximo nivel posible).
# Entrene una red neuronal artificial para tratar de predecir a partir del color de un pixel dado si se trata o no de piel humana. Utilice todas las muestras para el entrenamiento.
# Detalle la arquitectura empleada y los valores de los parámetros usados. Documente todos los intentos realizados. Informe los mejores resultados obtenidos. Utilice la imagen suministrada junto con el enunciado para validar la efectividad del clasificador.
import numpy as np
import matplotlib.pyplot as plt
import matplotlib as mpl
from matplotlib import pylab
import mpld3
# %matplotlib inline
mpld3.enable_notebook()
from cperceptron import Perceptron
from cbackpropagation import ANN #, Identidad, Sigmoide
import patrones as magia
def progreso(ann, X, T, y=None, n=-1, E=None):
if n % 20 == 0:
print("Pasos: {0} - Error: {1:.32f}".format(n, E))
def progresoPerceptron(perceptron, X, T, n):
y = perceptron.evaluar(X)
incorrectas = (T != y).sum()
print("Pasos: {0}\tIncorrectas: {1}\n".format(n, incorrectas))
# +
piel = np.load('piel.npy')
color = piel[:, :-1]
tipos = piel[:, -1]
# tipos == 1 --> Piel
# tipos == 2 --> No Piel
# -
#Armo Patrones
clases, patronesEnt, patronesTest = magia.generar_patrones(
magia.escalar(color),tipos,90)
X, T = magia.armar_patrones_y_salida_esperada(clases,patronesEnt)
Xtest, Ttest = magia.armar_patrones_y_salida_esperada(clases,patronesTest)
# ## Prueba con Perceptron
# +
# Esto es para poder usar Cython y que sea mas rapido
TT = T[:,0].copy(order='C')
TT = TT.astype(np.int8)
p1 = Perceptron(X.shape[1])
p1.reiniciar()
I1 = p1.entrenar(X, TT, max_pasos=10000, callback=progresoPerceptron, frecuencia_callback=5000)
print("Pasos:{0}".format(I1))
# -
print("Errores:{0} de {1}".format((p1.evaluar(Xtest) != Ttest[:,0]).sum(),Ttest.shape[0]))
# ## Prueba con BackPropagation
# Crea la red neuronal
ocultas = 8 #2,5,10(0.13),20(0.018),100(0.014),8(0.017)
entradas = X.shape[1]
salidas = T.shape[1]
ann = ANN(entradas, ocultas, salidas)
ann.reiniciar()
#Entreno
E, n = ann.entrenar_rprop(X, T, min_error=0, max_pasos=1000, callback=progreso, frecuencia_callback=100)
print("\nRed entrenada en {0} pasos con un error de {1:.32f}".format(n, E))
#Evaluo
Y = (ann.evaluar(Xtest) >= 0.89)
magia.matriz_de_confusion(Ttest,Y)
# Para probar el clasificador
imagen = np.load('imagen.npy')
pylab.imshow(imagen)
# La imagen ahora en una matriz donde en cada fila está el color de cada pixel.
imagen_lineal = imagen.reshape(-1, 3)[:, ::-1]
foto = imagen_lineal.copy(order='C').astype(np.float64)
#Evaluo
Y = (ann.evaluar(foto) >= 0.95).astype(np.int8)
fotoClasificada = Y[:,1].reshape(imagen.shape[:-1]).astype(np.int8)
# +
# Vuelve a darla las dimensiones de la imagen al resultado de la clasificación.
#clasificacion = Y.reshape(imagen.shape[:-1])
pylab.gray()
pylab.imshow(fotoClasificada)
pylab.show()
#LO BLANCO ES PIEL
# -
ann.guardar('')
| Argentina - Mondiola Rock - 90 pts/Practica/TP1/ejercicio 9/.ipynb_checkpoints/Ejercicio 9-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
import pandas as pd
from bs4 import BeautifulSoup
import bs4
import requests
club_data = []
for x in range(1, 33):
url = 'http://competitie.vttl.be/index.php?menu=1&cur_page='+str(x)
r = requests.get(url)
soup = BeautifulSoup(r.text, "lxml")
table = soup.find('table', attrs={'class':'DBTable'})
rows = table.find_all('tr')[2:-2]
for row in rows:
cols = row.find_all('td')
cols = [ele.text.strip() for ele in cols]
club_data.append([ele for ele in cols if ele])
club_df = pd.DataFrame(club_data)
club_df.columns = ['index', 'club', 'categorie', 'atl lokalen', 'site']
club_df.set_index('index', inplace=True)
club_df.index.name = None
club_df.head()
naamjuist_elo_df[(naamjuist_elo_df["naam_juist"]=="<NAME>") & (naamjuist_elo_df["club_y"]=="Merelbeke")].punten.values[0]
#Opmerking: om in deze rangschikking te worden opgenomen,
#moet de speler tenminste 24 individuele ontmoetingen in de laatste 12 maanden hebben gespeeld.
elo_data = []
for x in range(1, 191):
url = 'http://competitie.vttl.be/index.php?menu=5&perelo=1&club_id=0&cur_page='+str(x)
r = requests.get(url)
soup = BeautifulSoup(r.text, "lxml")
table = soup.find('table', attrs={'class':'DBTable'})
rows = table.find_all('tr')[2:-2]
for row in rows:
cols = row.find_all('td')
cols = [ele.text.strip() for ele in cols]
elo_data.append([ele for ele in cols if ele])
elo_df = pd.DataFrame(elo_data)
elo_df.columns = ['plaats', 'naam', 'klassement', 'club', 'punten']
elo_df.tail()
print("aantal dubbele namen: ",len(elo_df.naam[elo_df.naam.duplicated(keep=False)].index))
elo_df[elo_df["naam"]=="<NAME>"]
#er zijn toch wel redelijk wat dezelfde namen, wat doen we daar mee?
player_data = []
for x in range(1, 418):
url = 'http://competitie.vttl.be/index.php?menu=6&club_id=0&categ_id=1&cur_page='+str(x)
r = requests.get(url)
soup = BeautifulSoup(r.text, "lxml")
table = soup.find_all('table', attrs={'class':'DBTable'})[-1]
rows = table.find_all('tr')[2:-2]
for row in rows:
cols = row.find_all('td')
cols = [ele.text.strip() for ele in cols]
player_data.append([ele for ele in cols if ele])
player_df = pd.DataFrame(player_data)
player_df.columns = ['lidnummer', 'achternaam', 'voornaam', 'klassement', 'club']
def get_namen(row):
achternaam = str(row['achternaam'])
achternaam = achternaam.replace(u'\xa0', u' ')
voornaam = str(row['voornaam'])
voornaam = voornaam.replace(u'\xa0', u' ')
row['naam_juist']=voornaam+' '+achternaam
row['naam']=achternaam+' '+voornaam
return row
player_df = player_df.apply(get_namen, axis=1)
#join player_df en elo_df op naam_omgekeerd
naamjuist_elo_df = pd.merge(player_df, elo_df, on='naam')
club_match_df = pd.DataFrame(club_match_data)
club_match_df.columns = ['wedstrijd', 'thuis', 'bezoekers', 'score']
club_match_df.head()
ind_match_data = []
for x in range(1, 191):
url = 'http://competitie.vttl.be/index.php?menu=5&perelo=1&club_id=0&cur_page='+str(x)
r = requests.get(url)
soup = BeautifulSoup(r.text, "lxml")
table = soup.find('table', attrs={'class':'DBTable'})
for link in table.find_all('a'):
if link.get('href').startswith('http'):
name = str(link.contents)
r = requests.get(link.get('href'))
soup = BeautifulSoup(r.text, "lxml")
div = soup.find("div", {"id": "match_list"})
#table_2 = div.find('table', attrs={'class':'DBTable'})
rows = div.find_all('tr')[2:-2]
for row in rows:
cols = row.find_all('td')
cols = [ele.text.strip() for ele in cols]
cols.insert(0,name)
ind_match_data.append([ele for ele in cols if ele])
ind_match_df = pd.DataFrame(ind_match_data)
ind_match_df.columns = ['naam', 'datum', 'type', 'M/T', 'P/R', 'tegenstander', 'naam teg', 'klass teg', 'elo teg', 'sets', 'elo', 'diff']
ind_match_df.head()
#saving to a csv file as a backup
ind_match_df.to_csv('ind_match.csv')
#page crashed, loading it in again via csv, works perfectly
ind_match_df = pd.read_csv('ind_match.csv')
ind_match_df.head()
ind_match_df = ind_match_df.drop('Unnamed: 0', 1)
ind_match_df[['elo teg','elo','diff']] = ind_match_df[['elo teg','elo','diff']].apply(pd.to_numeric, errors='coerce')
len(ind_match_df.drop_duplicates(["naam"]).index)
ind_match_df.to_csv('ind_match_df.gz', compression='gzip')
import matplotlib.pyplot as plt
# %matplotlib inline
sample_df = ind_match_df[ind_match_df["naam"]=="['<NAME>']"]
sample_df_2 = ind_match_df[ind_match_df["naam"]=="['<NAME>']"]
sample_df_3 = ind_match_df[ind_match_df["naam"]=="['<NAME>']"]
ind_match_df[ind_match_df["naam"]=="['<NAME>']"].elo.tail(1).values[0]
def get_potential_ELO(row):
try:
tegenstander = str(row['tegenstander'])
naam = str(row['naam'])
naam = naam.replace(u'\xa0', u' ')
naam_teg = str(row['naam teg'])
naam_teg = naam_teg.replace(u'\xa0', u' ')
row['pot_elo']=ind_match_df[ind_match_df["naam"]==naam].elo.tail(1).values[0]
row['pot_teg_elo']=naamjuist_elo_df[(naamjuist_elo_df["naam_juist"]==naam_teg)].punten.values[0]
except:
row['pot_teg_elo']=row['elo teg']
return row
# +
plt.ylabel('Elo rating', fontsize=14)
plt.plot(sample_df_2.elo.values, label="<NAME>")
plt.plot(sample_df.elo.values, label="<NAME>")
plt.legend(loc="lower right")
#de vorm van de speler kunnen we misschien uit de rico halen van de ELO-grafiek, jonge spelers worden steeds beter,
#dus stijgende grafiek -> betere resultaten dan verwacht, oudere spelers kunnen ook afzwakken -> slechter scoren
#spelers die goed scoren tegen betere tegenstanders en slecht scoren tegen slechtere spelers kunnen we misschien
#herkennen aan het aantal grote sprongen (diff groter dan X) hun ELO maakt, daarvoor wel meer data nodig dan 1 seizoen
#of je neemt ELO op het einde van het jaar op, of max ELO -> potentieel
#maar ook die potentiele ELO van de tegenstander moet opgenomen worden, anders maar weinig effect
# -
#sets omzetten tot resultaten, 0/1 en een resultaat dat rekening houdt met het aantal sets
def transform_sets(row):
sets = str(row['sets'])
sets = sets.replace(u'\xa0', u' ')
if sets == '4 - 3':
row['sets_score']=0.8
row['score']=1
elif sets == '4 - 2':
row['sets_score']=0.85
row['score']=1
elif sets == '4 - 1':
row['sets_score']=0.9
row['score']=1
elif sets == '4 - 0':
row['sets_score']=1
row['score']=1
elif sets == '0 - 4':
row['sets_score']=0
row['score']=0
elif sets == '1 - 4':
row['sets_score']=0.1
row['score']=0
elif sets == '2 - 4':
row['sets_score']=0.15
row['score']=0
elif sets == '3 - 4':
row['sets_score']=0.2
row['score']=0
elif sets == '3 - 2':
row['sets_score']=0.8
row['score']=1
elif sets == '3 - 1':
row['sets_score']=0.9
row['score']=1
elif sets == '3 - 0':
row['sets_score']=1
row['score']=1
elif sets == '0 - 3':
row['sets_score']=0
row['score']=0
elif sets == '1 - 3':
row['sets_score']=0.1
row['score']=0
elif sets == '2 - 3':
row['sets_score']=0.2
row['score']=0
else:
row['sets_score']=None
row['score']=None
return row
test_df = ind_match_df.sample(5000).apply(transform_sets, axis=1)
test_df = test_df.apply(get_potential_ELO, axis=1)
#klasselabels encoden, deze labels hebben niet echt een toegevoegde waarde voor ons model
from sklearn import preprocessing
le = preprocessing.LabelEncoder()
le.fit(test_df['klass teg'])
test_df['klass teg'] = test_df['klass teg'].apply(le.transform)
test_df = test_df.drop(['naam', 'datum', 'type', 'M/T', 'P/R', 'tegenstander', 'naam teg', 'klass teg', 'sets','diff'], 1)
test_df = test_df.dropna()
test_df.head()
test_under100_df = test_df.loc[abs(test_df['elo'] - test_df['elo teg']) <= 100.0]
test_above100_df = test_df.loc[abs(test_df['elo'] - test_df['elo teg']) >= 101.0]
sample_count = 0.3*len(test_under100_df.index)
test_concat_df = pd.concat([test_under100_df, test_above100_df.sample(n=int(sample_count))])
test_concat_1 = test_df.loc[test_df['elo'] - test_df['elo teg'] > 0.0]
test_concat_2 = test_concat_1.loc[test_df['score'] > 0.0]
len(test_concat_2.index)/len(test_concat_1.index)
#Hiermee kan je checken wat de precisie is van het model "Elo > Elo teg -> winst" door deze te vergelijken
# +
from sklearn.linear_model import LogisticRegression
from sklearn.cross_validation import train_test_split
train_data, test_data = train_test_split(test_df, test_size=.2, random_state=42)
y_train = train_data['score']
x_train = train_data.drop(['sets_score','score'], 1)
y_test = test_data['score']
x_test = test_data.drop(['sets_score','score'], 1)
# -
lr_correct = LogisticRegression()
lr_correct.fit(x_train, y_train)
y_train_predict = lr_correct.predict(x_train)
# +
#lr_correct.predict_proba([1600,1300,1600,1500])
#dit kunnen we gebruiken om nieuwe matchen te voorspellen, op basis van beide ELO's en potentiële ELO's
# -
print(sum((y_train-y_train_predict)==0)/len(y_train))
y_test_predict = lr_correct.predict(x_test)
print(sum((y_test-y_test_predict)==0)/len(y_test))
from sklearn.metrics import precision_score
from sklearn.ensemble import RandomForestClassifier
rnd_clf = RandomForestClassifier(n_estimators=100)
rnd_clf.fit(x_train, y_train)
y_pred_rf = rnd_clf.predict(x_test)
print(precision_score(y_test, y_pred_rf), 'precision')
| Tabletennis_Project.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.4.0-dev
# language: julia
# name: julia-0.4
# ---
# # Integration tests for interleaved filter/smoother
#
# [<NAME> (2000)](http://onlinelibrary.wiley.com/doi/10.1111/1467-9892.00186/abstract) and <NAME> (2012) section 6.4 showed that a method of interleaving vector observations to create a univariate series avoided matrix inversion and resulted in a large computational speedup.
#
# **Important**: This method assumes that $\varepsilon_t \sim \mathcal{N}(0, H_t)$ with $H_t$ diagonal!
# +
using Distributions
using PyPlot
plt.style[:use]("ggplot")
srand(12345) # set random seed
# -
# # Generate data from a state space model
#
# **Note**: Observation noise is deliberately set low here so that recovered states line up *exactly* below (i.e., it's a sanity check).
# +
Zk = [1 -1. ; 0 1 ; 1 0] # observation matrix
Hk = 10 * [1 0 0 ; 0 1.2 0 ; 0 0 1.5]
Np, Nm = size(Zk)
th = 10 * pi / 180 # rotation angle
Tk = [cos(th) sin(th) ; -sin(th) cos(th)]
Rk = reshape([1.1 1], 2, 1)
Qk = reshape([0.25], 1, 1)
Nr = size(Rk, 2)
a_init = zeros(Nm)
P_init = 5 * eye(Nm)
# -
# ## Now generate some data
Nt = 100 # number of samples
Z = Array(Float64, size(Zk)..., Nt)
T = Array(Float64, size(Tk)..., Nt)
H = Array(Float64, size(Hk)..., Nt)
R = Array(Float64, size(Rk)..., Nt)
Q = Array(Float64, size(Qk)..., Nt)
for t in 1:Nt
Z[:, :, t] = Zk
T[:, :, t] = Tk + 0.05 * rand()
H[:, :, t] = (1 + 0.05 * rand()) * Hk
R[:, :, t] = (1 + 0.05 * rand()) * Rk
Q[:, :, t] = (1 + 0.15 * rand()) * Qk
end
# preallocate data arrays
α = Array(Float64, Nm, Nt)
y = Array(Float64, Np, Nt)
ϵ = Array(Float64, Np, Nt)
η = Array(Float64, Nr, Nt)
# +
# initialize
α[:, 1] = rand(MvNormal(a_init, P_init))
for t in 1:Nt
Z_t = Z[:, :, t]
T_t = T[:, :, t]
H_t = H[:, :, t]
R_t = R[:, :, t]
Q_t = Q[:, :, t]
ϵ[:, t] = rand(MvNormal(H_t))
η[:, t] = rand(MvNormal(Q_t))
y[:, t] = Z_t * α[:, t] + ϵ[:, t]
if t < Nt
α[:, t + 1] = T_t * α[:, t] + R_t * η[:, t]
end
end
# +
figure(figsize=(10, 4))
plot(y')
figure(figsize=(10, 4))
plot(α')
# -
# # Kalman filter
import SStools
v, K, Finv, a2, P = SStools.kalman_filter(y, a_init, P_init, Z, H, T, R, Q);
v, K, Finv, a, P = SStools.interleaved_kalman_filter(y, a_init, P_init, Z, H, T, R, Q);
figure(figsize=(10, 4))
plot(squeeze(a[:, 1, :], 2)', label="Interleaved")
plot(a2', label="Standard")
plot(α', label="Actual")
title("Actual and recovered states")
legend()
# #Smoother
ahat = SStools.interleaved_state_smoother(v, K, Finv, a_init, P_init, Z, T, R, Q);
v, K, Finv, a2, P = SStools.kalman_filter(y, a_init, P_init, Z, H, T, R, Q);
ahat2 = SStools.fast_state_smoother(v, K, Finv, a_init, P_init, Z, T, R, Q);
figure(figsize=(10, 4))
plot(ahat' - ahat2', label="difference")
title("Difference between interleaved and normal Kalman filter")
legend()
figure(figsize=(10, 4))
plot(ahat', label="smoothed")
plot(squeeze(a[:, 1, :], 2)', label="filtered")
plot(α', label="actual")
title("Actual and recovered states")
legend()
| julia/kalman_interleaved_tests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img align="left" src="https://lever-client-logos.s3.amazonaws.com/864372b1-534c-480e-acd5-9711f850815c-1524247202159.png" width=200>
# <br></br>
# <br></br>
#
# ## *Data Science Unit 4 Sprint 3 Assignment 2*
# # Convolutional Neural Networks (CNNs)
# + [markdown] colab_type="text" id="0lfZdD_cp1t5"
# # Assignment
#
# Load a pretrained network from Keras, [ResNet50](https://tfhub.dev/google/imagenet/resnet_v1_50/classification/1) - a 50 layer deep network trained to recognize [1000 objects](https://storage.googleapis.com/download.tensorflow.org/data/ImageNetLabels.txt). Starting usage:
#
# ```python
# import numpy as np
#
# from tensorflow.keras.applications.resnet50 import ResNet50
# from tensorflow.keras.preprocessing import image
# from tensorflow.keras.applications.resnet50 import preprocess_input, decode_predictions
#
# ResNet50 = ResNet50(weights='imagenet')
# features = model.predict(x)
#
# ```
#
# Next you will need to remove the last layer from the ResNet model. Here, we loop over the layers to use the sequential API. There are easier ways to add and remove layers using the Keras functional API, but doing so introduces other complexities.
#
# ```python
# # Remote the Last Layer of ResNEt
# ResNet50._layers.pop(0)
#
# # Out New Model
# model = Sequential()
#
# # Add Pre-trained layers of Old Model to New Model
# for layer in ResNet50.layers:
# model.add(layer)
#
# # Turn off additional training of ResNet Layers for speed of assignment
# for layer in model.layers:
# layer.trainable = False
#
# # Add New Output Layer to Model
# model.add(Dense(1, activation='sigmoid'))
# ```
#
# Your assignment is to apply the transfer learning above to classify images of Mountains (`./data/mountain/*`) and images of forests (`./data/forest/*`). Treat mountains as the postive class (1) and the forest images as the negative (zero).
#
# Steps to complete assignment:
# 1. Load in Image Data into numpy arrays (`X`) -> you do
# 2. Create a `y` for the labels -> you do
# 3. Train your model with pretrained layers from resnet
# 4. Report your model's accuracy
# -
pip install resnet
# +
import numpy as np
import pandas as pd
import requests
from tensorflow.keras.layers import Dense, GlobalAveragePooling2D
from tensorflow.keras.layers import Convolution2D, Flatten, MaxPooling2D
from tensorflow.keras.models import Model, Sequential # This is the functional API
# +
# instantiate the model, specifying pretrained weights
from tensorflow.keras.applications.resnet50 import ResNet50
from tensorflow.keras.preprocessing import image
from tensorflow.keras.applications.resnet50 import preprocess_input, decode_predictions
ResNet50 = ResNet50(weights='imagenet', include_top = False)
# -
# ## Restart
# +
from skimage.io import imread_collection
from skimage.transform import resize
from sklearn.model_selection import train_test_split
# image-data preprocessing
forests = 'data/forest/*.jpg'
mountains = 'data/mountain/*.jpg'
forests = imread_collection(forests).concatenate()
mountains = imread_collection(mountains).concatenate()
y_0 = np.zeros(forests.shape[0])
y_1 = np.ones(mountains.shape[0])
x = np.concatenate([forests,mountains])
#X = x/255
x = resize(x,(702,224,224,3))
y = np.concatenate([y_0, y_1])
x_train, x_test, y_train, y_test = train_test_split(x,y, test_size = .2)
# -
x_train.shape,x_test.shape,y_train.shape,y_test.shape
import matplotlib.pyplot as plt
plt.imshow(x[200])
# + jupyter={"outputs_hidden": true}
#help(ResNet50)
ResNet50 = ResNet50(input_shape=(256, 256, 3), weights='imagenet', include_top=False)
# +
for layer in ResNet50.layers:
layer.trainable = False
x = ResNet50.output
x = GlobalAveragePooling2D()(x)
x = Dense(1024, activation='relu')(x)
x = Dropout(0.25)(x)
x = Dense(512, activation='relu')(x)
x = Dropout(0.25)(x)
x = Dense(256, activation='relu')(x)
predictions = Dense(1, activation='sigmoid')(x)
model = Model(ResNet50.input, predictions)
# -
model.summary()
model.compile(optimizer='adam', loss='binary_crossentropy', metrics=['accuracy'])
model.fit(x_train, y_train, epochs=5, validation_split=.2)
# +
test_loss, test_acc = model.evaluate(x_test, y_test, verbose=2)
# --end of materials for the day--
# -
assert(2 == 1)
# +
resnet = ResNet50(inputs = x_train)
# error in top 2 lines; weights =,
for layer in resnet.layers:
layer.trainable = False
x = resnet.output
x = GlobalAveragePooling2D()(x)
x = Dense(1024, activation = 'relu')(x)
predictions = Dense(1, activation = 'sigmoid')(x)
model = Model(inputs = resnet.input, outputs = predictions)
model.compile(loss='binary_crossentropy',
optimizer='adam',
metrics=['accuracy'])
# -
resnet.summary()
# + [markdown] colab_type="text" id="uT3UV3gap9H6"
# # Resources and Stretch Goals
#
# Stretch goals
# - Enhance your code to use classes/functions and accept terms to search and classes to look for in recognizing the downloaded images (e.g. download images of parties, recognize all that contain balloons)
# - Check out [other available pretrained networks](https://tfhub.dev), try some and compare
# - Image recognition/classification is somewhat solved, but *relationships* between entities and describing an image is not - check out some of the extended resources (e.g. [Visual Genome](https://visualgenome.org/)) on the topic
# - Transfer learning - using images you source yourself, [retrain a classifier](https://www.tensorflow.org/hub/tutorials/image_retraining) with a new category
# - (Not CNN related) Use [piexif](https://pypi.org/project/piexif/) to check out the metadata of images passed in to your system - see if they're from a national park! (Note - many images lack GPS metadata, so this won't work in most cases, but still cool)
#
# Resources
# - [Deep Residual Learning for Image Recognition](https://arxiv.org/abs/1512.03385) - influential paper (introduced ResNet)
# - [YOLO: Real-Time Object Detection](https://pjreddie.com/darknet/yolo/) - an influential convolution based object detection system, focused on inference speed (for applications to e.g. self driving vehicles)
# - [R-CNN, Fast R-CNN, Faster R-CNN, YOLO](https://towardsdatascience.com/r-cnn-fast-r-cnn-faster-r-cnn-yolo-object-detection-algorithms-36d53571365e) - comparison of object detection systems
# - [Common Objects in Context](http://cocodataset.org/) - a large-scale object detection, segmentation, and captioning dataset
# - [Visual Genome](https://visualgenome.org/) - a dataset, a knowledge base, an ongoing effort to connect structured image concepts to language
| module2-convolutional-neural-networks/LS_DS_432_Convolution_Neural_Networks_Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/akankshawaghmare/Microspectra/blob/main/listDataType.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="SmS6-lNZzgDf" outputId="ec0b4683-da04-449a-a242-2e688f159e62"
#programs on list using range function
list1 = []
num1 = int(input('Enter size of list 1: '))
for n in range(num1):
x = int(input('Enter any number:'))
list1.append(x)
list2 = []
num2 = int(input('Enter size of list 2:'))
for n in range(num2):
y = int(input('Enter any number:'))
list2.append(y)
union = list(set().union(list1,list21))
print('The Union of two lists is:',union)
# + id="-sRm3agzkK06" colab={"base_uri": "https://localhost:8080/"} outputId="daf68830-231b-4b38-a0a1-52272974d483"
name=['vaishnavi','krutika','shivani','akanksha','archana','yogita','arpita']
for friend in range(len(name)):
print("my friend's name is", name[friend])
num=[]
sum=0
for i in range(1,10000000):
sum=sum + i
print(sum)
cubes=[]
for value in range(1,11):
c=value ** 3
cubes.append(c)
print(cubes)
cubes=[value**3 for value in range(1,11)]
print(cubes)
# + colab={"base_uri": "https://localhost:8080/"} id="bZYjTJ1d2OlR" outputId="c5c5fae9-c255-4398-b9d1-a9af1fd2d26b"
names=['akanksha','vaishnavi','krutika','archana']
x=input('Enter the name you want :')
if x in names :
print('you entered the name : ',x)
else:
print('The name you have entered is not present')
names.reverse()
print(names)
# + colab={"base_uri": "https://localhost:8080/"} id="YYrDNXGf33hw" outputId="ccdce959-d341-4977-8dd1-f96d08a6e449"
#program for addition of the list elements
list0=[]
sum=0
n=int(input("enter number of item in list"))
for i in range(0,n):
a=int(input())
list0.append(a)
print("list = ",list0)
for i in range(0,len(list0)):
sum=sum+list0[i]
print("sum of items in list is: ",sum)
# + colab={"base_uri": "https://localhost:8080/"} id="kdF4k85V48E-" outputId="b403d277-42ad-43f5-aea6-e61d6adabddf"
#program for even numbers
numbers=[1,2,3,4,5,6,7,8,9,10,11,12,13,14,15]
for i in numbers :
if i % 2 == 0 :
print(numbers[i])
# + colab={"base_uri": "https://localhost:8080/"} id="KOPPsYhF3Us8" outputId="c98d71a9-37ea-45a4-eeaa-e977c2623578"
num = [1,2,3,4,5,6,7,8,9,10]
eve=[]
for i in num:
if i%2==0:
print(i)
# eve.append(i)
#print(eve)
# + colab={"base_uri": "https://localhost:8080/"} id="hR9KO8MrfyMe" outputId="d883f07e-a099-4554-b15a-18ace3cabd8e"
list1 = [10, 21, 4, 45, 66, 93]
even_nos = [num for num in list1 if num % 2 == 0] #list comprehension for even numbers
print("Even numbers in the list: ", even_nos)
# + id="YFceY-z_f4rh"
eve=[i for i in range(0,11,2)] #list comprehension for even number in another way
print(eve)
| listDataType.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# %matplotlib inline
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
import scipy
from src import inception_v3_imagenet
from src import imagenet_labels
from src import utils
from src.diff_renderer import make_render_op
from src.utils import angles_to_matrix
# -
width, height = 1000, 1000
mesh = utils.load_obj('resources/dog.obj')
original_texture = mesh.texture_image.copy()
render_op = make_render_op(mesh, width, height)
trans = [0, 0, 2.6]
rotation = [0.4, np.pi+.7, 2.9]
fscale = 0.4
bgcolor = [0.9, 0.9, 0.9]
texture_image = mesh.texture_image
view_matrix = np.vstack((np.hstack((angles_to_matrix(rotation) , np.reshape(trans, (3, 1)) )) , np.array([0, 0, 0, 1])))
view_matrix
# +
pixel_center_offset = 0.5
near = 0.1
far = 100.
fmat = [float(width) / float(fscale), float(width) / float(fscale)]
f = 0.5 * (fmat[0] + fmat[1])
center = [width/2.,height/2.]
right = (width-(center[0]+pixel_center_offset)) * (near/f)
left = -(center[0]+pixel_center_offset) * (near/f)
top = -(height-(center[1]+pixel_center_offset)) * (near/f)
bottom = (center[1]+pixel_center_offset) * (near/f)
A = (right + left) / (right - left)
B = (top + bottom) / (top - bottom)
C = (far + near) / (far - near)
D = (2 * far * near) / (far - near)
camera_matrix = np.array([
[2 * near / (right - left), 0, A, 0],
[0, 2 * near / (top - bottom), B, 0],
[0, 0, C, D],
[0, 0, -1, 0]
])
# -
proj_matrix = camera_matrix.dot(view_matrix)
homo_v = np.hstack((mesh.v, np.ones((mesh.v.shape[0], 1) )))
homo_v
abnormal = proj_matrix.dot(homo_v.reshape((-1, 4, 1)))[:, :, 0]
XY = (abnormal[:,:] / abnormal[3,:]).T
XY
# +
# plt.set_autoscale_on(False)
plt.figure(figsize=(5,5))
plt.scatter(XY[:,0], XY[:, 1], c = XY[:, 2], s=3)
# plt.axes().set_aspect('equal', 'datalim')
plt.xlim([1, -1])
plt.ylim([1, -1])
# +
Z = XY[:,2]
face_depth = np.mean(np.take(Z, mesh.f.flatten()).reshape((-1, 3)), axis=1)
# -
face_depth
import matplotlib.cm as cm
# +
fXY = np.take(XY[:,0:2], mesh.f, axis=0)
Xmin, Xmax = np.min(fXY[:,:,0], axis=1), np.max(fXY[:,:,0], axis=1)
Ymin, Ymax = np.min(fXY[:,:,1], axis=1), np.max(fXY[:,:,1], axis=1)
# +
canvas = np.zeros((1000, 1000))
xmin = ((1. + Xmin) * 500.).astype(np.int)
xmax = ((1. + Xmax) * 500.).astype(np.int)
ymin = ((1. + Ymin) * 500.).astype(np.int)
ymax = ((1. + Ymax) * 500.).astype(np.int)
for i in range(len(mesh.f)):
canvas[ymin[i]:ymax[i], xmin[i]:xmax[i]] = i
plt.imshow(canvas)
# +
plt.figure(figsize=(5,5))
for i in range(len(mesh.f)):
tri = np.take(XY[:,0:2], mesh.f[i], axis=0)
circle = plt.Polygon(tri, fc=cm.hot( (1.1 + face_depth[i]) * 20))
plt.gca().add_patch(circle)
plt.xlim([1, -1])
plt.ylim([1, -1])
plt.show()
# -
np.array(np.meshgrid(np.linspace(0, 1, 1000), np.linspace(0, 1, 1000)))
| experiments/dr5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Lesson 2: Variables
# + [markdown] slideshow={"slide_type": "-"}
# In this lesson, we will learn about...
#
# # + Variable Assignment
# # + Primitive Data Types
# # + Type Casting
# + [markdown] slideshow={"slide_type": "slide"}
# # Variable Assignment
#
# **Assignment:** Giving a variable a value
# + slideshow={"slide_type": "-"}
a = 100
b = 1000.5
c = "Python"
# + [markdown] slideshow={"slide_type": "slide"}
# ## Identifiers
#
# # + An Identifier is a name to identify a variable, function, class, module, etc.
# # + Starts with a letter or an underscore, followed by one or more letters, underscores, digits
# + Class names start with a capital letter
# + All other identifiers start with a lowercase letter
# + Starting with an underscore is convention for a "private" variable
# # + No punctuation allowed
# # + Case sensitive
# # + Separate words with underscores
# + This is what Python's Style Guide suggests for readability
# + You may also see people using camelCase
# + [markdown] slideshow={"slide_type": "slide"}
# ## Reserved Words
#
# These are words with special meaning in Python, and cannot be used as identifiers. They usually change color in your IDE / editor.
# + slideshow={"slide_type": "-"}
help("keywords")
# + [markdown] slideshow={"slide_type": "slide"}
# ## Multiple Assignment
#
# Python allows you to assign values to multiple variables simulaneously.
# + slideshow={"slide_type": "-"}
d = e = 1
f,g,h = 2,3,"Programming"
# + [markdown] slideshow={"slide_type": "slide"}
# # Data Types
#
# Python has many built-in data types. We typically divide them into "primitives" and "non-primitives".
# + [markdown] slideshow={"slide_type": "slide"}
# ## Primitive Data Types
#
# These data types are the fundamental building blocks of a language. Python's primitives include...
#
# # + Text: `str`
# # + Numeric: `int`, `float`, `complex`
# # + Boolean: `bool`
#
# We'll learn about non-primitive data types in a later lesson.
# + slideshow={"slide_type": "subslide"}
this_is_a_string = "A String"
this_is_an_int = 5
this_is_a_float = 5.203
this_is_a_scientific_float = 1.5e6
this_is_complex = 5 + 3j
this_is_a_bool = True #(or False)
print(type(this_is_a_string))
print(type(this_is_an_int))
print(type(this_is_a_float))
print(type(this_is_a_scientific_float))
print(type(this_is_complex))
print(type(this_is_a_bool))
# + [markdown] slideshow={"slide_type": "notes"}
# # + **Strings** are literal text, enclosed in single, double, or triple quotes.
# # + **Integers** are whole numbers, positive or negative (or zero), without decimals
# # + **Floats** are positive or negative (or zero) numbers with decimals. Floats can be expressed in scientific notation as well.
# # + **Complex** numbers have a real component and an imaginary component (represented with `j`)
# # + **Booleans**, named after mathematician [George Bool](https://en.wikipedia.org/wiki/George_Boole), are binary values that are either `True` or `False`. The capitalization is important.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Type Casting
#
# # + Sometimes, you'll need to convert from one data type to another. This is called "type casting."
# # + Python will implicitly cast numbers when able to "larger" types to preserve data.
# + Ex: `int` + `float` yields a `float`
# + slideshow={"slide_type": "subslide"}
x = int(2.8)
y = float(3)
z = str(3)
w = int("7")
v = x + y
print(x)
print(type(x))
print(y)
print(type(y))
print(z)
print(type(z))
print(w)
print(type(w))
print(v)
print(type(v))
# + [markdown] slideshow={"slide_type": "slide"}
# ## String Special Features
#
# Strings represent a sequence of characters, and have some features we'll also see in the more complex data structures.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Get a Character
#
# # + Use brackets to get a character at a given position
# # + These **indexes** start with **the first character at 0**.
# + slideshow={"slide_type": "-"}
hello = "Hello, World!"
print(hello[1])
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Slicing / Substring
#
# # + Use the brackets with a colon to get a slice, or part, of the string from the start to end index (exclusive).
# # + `start` $\leq$ `end`
# # + **Tip:** `end` $-$ `start` $=$ length of slice
# # + You can also optionally leave out either the start or the end index
# + slideshow={"slide_type": "-"}
hello = "Hello, World!"
print(hello[2:5])
print(hello[:5])
print(hello[9:])
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Negative Indexing
#
# You can also use negative numbers to index from the end of the string! The last character is index `-1`.
# + slideshow={"slide_type": "-"}
hello = "Hello, World!"
print(hello[-5:-2])
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Length
#
# Use `len()` to get the length of, or number of characters in, a string.
# + slideshow={"slide_type": "-"}
hello = "Hello, World!"
print(len(hello))
# + [markdown] slideshow={"slide_type": "subslide"}
# ### String Methods
#
# There are various other methods you can run on a string. A full list can be found [here](https://www.w3schools.com/python/python_ref_string.asp).
#
# **Note:** All string methods return a *new* string, and don't replace the original.
# + slideshow={"slide_type": "-"}
hello = "Hello, World!"
print(hello.upper())
print(hello.lower())
print(hello.replace("H", "J"))
print(hello.split(","))
print("world" in hello)
# + [markdown] slideshow={"slide_type": "notes"}
# # + `upper()` and `lower()` transform the string to all uppercase and lowercase characters respecitively
# # + `replace()` will replace all instances of the first string with the second, case sensitive
# # + `split()` will split the string into a list of substrings at whatever string is specified. That separator is not included in any substring.
# # + `x in y` will return `True` if `x` is a substring of `y`, and `False` otherwise. We'll learn more about boolean operators later
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Concatenation
#
# You can concatenate two strings by adding them.
# + slideshow={"slide_type": "-"}
hello = "Hello, " + "World!"
print(hello)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Escape Characters
#
# Use escape sequences to insert invisible or otherwise illegal characters in strings. These include...
#
# |Escape|Result|
# |---:|:---|
# |`\'`|Single Quote|
# |`\"`|Double Quote|
# |`\n`|New Line|
# |`\r`|Carriage Return|
# |`\t`|Tab|
# |`\b`|Backspace|
# |`\\`|Backslash|
# + slideshow={"slide_type": "-"}
hello = "He\"ll\"o,\tWor\nld\\!\b"
print(hello)
| 02-variables/02-variables.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="fjPDduCLMPFZ" colab_type="code" colab={}
import os
import tensorflow as tf
import tensorflow_datasets as tfds
from tensorflow.keras import datasets, layers, models
# %matplotlib inline
import matplotlib.pyplot as plt
# + [markdown] id="VLbK_I05Mdbc" colab_type="text"
# # Gather Data
# + id="SqHSSX3WMl50" colab_type="code" outputId="38ad8e3f-2fde-4313-cdd5-d3149f3055f0" executionInfo={"status": "ok", "timestamp": 1588861650163, "user_tz": 240, "elapsed": 2632, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjAksu_ElVnuka7KuOLHjyaXiy4v9BA2P1AqagM2w=s64", "userId": "15195746376658990804"}} colab={"base_uri": "https://localhost:8080/", "height": 139}
# Download three different English translations of Homer's Illiad
# from three different translators.
url = 'https://storage.googleapis.com/download.tensorflow.org/data/illiad/'
names = ['cowper.txt', 'derby.txt', 'butler.txt']
for name in names:
text_dir = tf.keras.utils.get_file(name, origin=url+name)
# Note the data directory.
parent_dir = os.path.dirname(text_dir)
print(parent_dir)
# + id="y5Z0QWGwJ4tf" colab_type="code" outputId="7532cc63-eb74-4c19-9a6b-707a021e2890" executionInfo={"status": "ok", "timestamp": 1588861651476, "user_tz": 240, "elapsed": 3937, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjAksu_ElVnuka7KuOLHjyaXiy4v9BA2P1AqagM2w=s64", "userId": "15195746376658990804"}} colab={"base_uri": "https://localhost:8080/", "height": 86}
# See how many samples we have in each file.
# ! wc -l /root/.keras/datasets/*.txt
# + id="ucOL-ONt86SP" colab_type="code" outputId="a2201592-a420-47ab-fe69-c5710e9edf5b" executionInfo={"status": "ok", "timestamp": 1588861652611, "user_tz": 240, "elapsed": 5063, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjAksu_ElVnuka7KuOLHjyaXiy4v9BA2P1AqagM2w=s64", "userId": "15195746376658990804"}} colab={"base_uri": "https://localhost:8080/", "height": 191}
# Check out one of the translations.
# ! head /root/.keras/datasets/butler.txt
# + [markdown] id="eqc3TRfQZtjF" colab_type="text"
# # Pre-process the data
# + id="R8GwOfzQ9ip7" colab_type="code" colab={}
# labeler casts an input index to a new type (tf.int64).
def labeler(example, index):
return example, tf.cast(index, tf.int64)
# + id="W7HQT53ZSWGO" colab_type="code" colab={}
# This time we are going to use a couple convenient text-processing utilities
# built into TensorFlow to pre-process the data. PyTorch has similar utilies
# in TorchText.
# Create a list that will hold our labeled datasets.
labeled_data_sets = []
# Loop over the translation files and use TextLineDataset to create a dataset
# in which each sample is a line of text from the input file.
for i, name in enumerate(names):
lines_dataset = tf.data.TextLineDataset(os.path.join(parent_dir, name))
labeled_dataset = lines_dataset.map(lambda ex: labeler(ex, i))
labeled_data_sets.append(labeled_dataset)
# + id="laF8kcQo--ae" colab_type="code" colab={}
# Concatenate all the data together.
all_labeled_data = labeled_data_sets[0]
for labeled_dataset in labeled_data_sets[1:]:
all_labeled_data = all_labeled_data.concatenate(labeled_dataset)
# Shuffle the data. The 50000 number here is a buffer that TensorFlow fills as
# it shuffles the data. The buffer should be greater than or equal to the full
# numbers of samples in the dataset.
all_labeled_data = all_labeled_data.shuffle(
50000, reshuffle_each_iteration=False)
# + id="MpNUOqe9ASPF" colab_type="code" outputId="dca820e8-2b49-45b1-ac17-27b11fd58472" executionInfo={"status": "ok", "timestamp": 1588861663544, "user_tz": 240, "elapsed": 15977, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjAksu_ElVnuka7KuOLHjyaXiy4v9BA2P1AqagM2w=s64", "userId": "15195746376658990804"}} colab={"base_uri": "https://localhost:8080/", "height": 278}
# We can use the take method to print some of the data and see what it
# looks like.
for ex in all_labeled_data.take(5):
print('Text:', ex[0])
print('Label:', ex[1])
print('')
# + id="XzCoj0GZCYe5" colab_type="code" outputId="9b04ccdc-81d4-45a7-884b-90acc4831abb" executionInfo={"status": "ok", "timestamp": 1588861674301, "user_tz": 240, "elapsed": 26726, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjAksu_ElVnuka7KuOLHjyaXiy4v9BA2P1AqagM2w=s64", "userId": "15195746376658990804"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# Next we need to encode words into index values from a vocabulary, which also
# means that we need to build a vocabulary. First, let's create a tokenizer to
# split the text into words.
tokenizer = tfds.features.text.Tokenizer()
# Build our vocabulary of words.
vocabulary_set = set()
for text_tensor, _ in all_labeled_data:
tokens = tokenizer.tokenize(text_tensor.numpy())
vocabulary_set.update(tokens)
# See the size of our vocab.
vocab_size = len(vocabulary_set)
vocab_size
# + id="Smp9QD6BDIqt" colab_type="code" colab={}
# Finally let's encode each of our examples into a list of vocabulary index
# values using the vocabulary and the TokenTextEncoder function from TF.
encoder = tfds.features.text.TokenTextEncoder(vocabulary_set)
# encode_text encodes a text sample with the TokenTextEncoder.
def encode_text(text_tensor, label):
encoded_text = encoder.encode(text_tensor.numpy())
return encoded_text, label
# encode_sample encodes a labeled sample from the data set and outputs a tensor.
def encode_sample(text, label):
encoded_text, label = tf.py_function(encode_text, inp=[text, label],
Tout=(tf.int64, tf.int64))
encoded_text.set_shape([None])
label.set_shape([])
return encoded_text, label
# Encode all our data using the above functions.
all_encoded_data = all_labeled_data.map(encode_sample)
# + id="vxnMWF-iJWIc" colab_type="code" colab={}
# We will keep 5000 samples out for our test set, and we will use a batch size
# of 64 samples during training.
test_size = 5000
batch_size = 64
# Create our training and test batches. Note, we are using a function called
# padded_batch here which takes care of padding the samples due to their varied
# sequence length.
train_data = all_encoded_data.skip(test_size).shuffle(50000)
train_data = train_data.padded_batch(batch_size, padded_shapes=([None],[]))
test_data = all_encoded_data.take(test_size)
test_data = test_data.padded_batch(batch_size, padded_shapes=([None],[]))
# We have padded the dataset with a new token "0" so we need to increase
# the size of our vocab.
vocab_size += 1
# + [markdown] id="04EArt1TaKiY" colab_type="text"
# # Define the model
# + id="eGeSeGh_aC5P" colab_type="code" colab={}
# mynet initializes our neural network model
def mynet():
# This is a Sequential "stack of layers" model.
model = models.Sequential()
# Add an embedding layer with a size vocab_size x 64, because we are
# using a batch size of 64.
model.add(tf.keras.layers.Embedding(vocab_size, 64))
# Add an Bidirectional LSTM layer. Bidirectional means that forward looking
# and backward looking LSTM cells are both used.
model.add(tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(64)))
# Add 2 dense layers, each with 64 nodes.
for units in [64, 64]:
model.add(tf.keras.layers.Dense(units, activation='relu'))
# Create our output layer for our three classes.
model.add(tf.keras.layers.Dense(3))
return model
# + id="81f_tvdbcG3X" colab_type="code" colab={}
# Intialize the model
model = mynet()
# + id="yFWAza3WcJO7" colab_type="code" colab={}
# In this case we are going to update our optimization strategy to "adam," which
# is quite widely used. We are also going to use the SparseCategoricalCrossentropy
# loss which excepts our categories to NOT be one hot encoded and accounts for
# this internally.
model.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
# + [markdown] id="0FHj0HUAc92j" colab_type="text"
# # Train the model
# + id="k4ALEoCYc8Ve" colab_type="code" outputId="997913a9-0ef2-4c75-c2a3-d6444c4b271e" executionInfo={"status": "ok", "timestamp": 1588861771100, "user_tz": 240, "elapsed": 123505, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjAksu_ElVnuka7KuOLHjyaXiy4v9BA2P1AqagM2w=s64", "userId": "15195746376658990804"}} colab={"base_uri": "https://localhost:8080/", "height": 121}
# Train (aka fit) the model.
num_epochs = 3
history = model.fit(train_data, epochs=num_epochs)
# + id="QAg03DDBe8Sa" colab_type="code" outputId="562ff350-0369-4a01-9f3f-16175c303643" executionInfo={"status": "ok", "timestamp": 1588861771496, "user_tz": 240, "elapsed": 123897, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjAksu_ElVnuka7KuOLHjyaXiy4v9BA2P1AqagM2w=s64", "userId": "15195746376658990804"}} colab={"base_uri": "https://localhost:8080/", "height": 312}
# Plot the training loss for each epoch to see how the model converged.
history_dict = history.history
plt.plot(list(range(1,num_epochs+1)), history_dict['loss'], 'bo--', label='Training loss')
plt.title('Training loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
# + [markdown] id="hoaKKz5GfjJO" colab_type="text"
# # Evaluate the model
# + id="wXVw-Kn7dKfO" colab_type="code" outputId="7430d993-3120-441e-f43c-3203ecf3da74" executionInfo={"status": "ok", "timestamp": 1588861777088, "user_tz": 240, "elapsed": 129484, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjAksu_ElVnuka7KuOLHjyaXiy4v9BA2P1AqagM2w=s64", "userId": "15195746376658990804"}} colab={"base_uri": "https://localhost:8080/", "height": 69}
# Evaluate the model on the test set.
test_loss, test_acc = model.evaluate(test_data, verbose=2)
print('\nTest accuracy:', test_acc)
# + id="_ySrBUNtqkay" colab_type="code" colab={}
| day2/session3/example1.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.1
# language: julia
# name: julia-1.6.1
# ---
using Revise
using MDToolbox, PyPlot, Statistics, LinearAlgebra
PyPlot.plt.style.use("seaborn-colorblind")
ENV["COLUMNS"] = 110; #display width for MDToolbox
# -----
s1 = readlines("new1/run_1.rem")
s2 = readlines("new1/run_2.rem")
s3 = readlines("new1/run_3.rem")
s4 = readlines("new1/run_4.rem")
s5 = readlines("new1/run_5.rem")
s6 = readlines("new1/run_6.rem")
s7 = readlines("new1/run_7.rem")
s8 = readlines("new1/run_8.rem")
N = 100 #100000 エネルギーのアウトプット数 dihedralの場合10,000,000/1,000=10,000 10,000/100*12をした
STEP = zeros(Int,N)
rep_1 = zeros(Int32,N)
rep_2 = zeros(Int32,N)
rep_3 = zeros(Int32,N)
rep_4 = zeros(Int32,N)
rep_5 = zeros(Int32,N)
rep_6 = zeros(Int32,N)
rep_7 = zeros(Int32,N)
rep_8 = zeros(Int32,N)
temp_1 = 300
temp_2 = 345
temp_3 = 397
temp_4 = 452
temp_5 = 513
temp_6 = 585
temp_7 = 660
temp_8 = 740
k = 0
#for i = 1:length(s1) #iは全体のエネルギーの探索
#if startswith(s1[i]," 0") #ここからはじまり
for j = 1:N #N #jは10000000ステップの中での探索(エネルギーは100ステップおきに出力) このエネルギーを1000でストライドしたトラジェクトリに対応するように抽出していく
k = k + 1
#STEP[k] = parse(Int,s1[j*6000-1][3:10])
#rep_1[k] = parse(Int32,s1[j*6000-1][19:20])
#rep_2[k] = parse(Int32,s2[j*6000-1][19:20])
#rep_3[k] = parse(Int32,s3[j*6000-1][19:20])
#rep_4[k] = parse(Int32,s4[j*6000-1][19:20])
#rep_5[k] = parse(Int32,s5[j*6000-1][19:20])
#rep_6[k] = parse(Int32,s6[j*6000-1][19:20])
#rep_7[k] = parse(Int32,s7[j*6000-1][19:20])
#rep_8[k] = parse(Int32,s8[j*6000-1][19:20])
STEP[k] = parse(Int,s1[j][3:10])
rep_1[k] = parse(Int32,s1[j][19:20])
rep_2[k] = parse(Int32,s2[j][19:20])
rep_3[k] = parse(Int32,s3[j][19:20])
rep_4[k] = parse(Int32,s4[j][19:20])
rep_5[k] = parse(Int32,s5[j][19:20])
rep_6[k] = parse(Int32,s6[j][19:20])
rep_7[k] = parse(Int32,s7[j][19:20])
rep_8[k] = parse(Int32,s8[j][19:20])
end
#end
#end
STEP
# +
for i = 1:N #N #jは10000000ステップの中での探索(エネルギーは100ステップおきに出力) このエネルギーを1000でストライドしたトラジェクトリに対応するように抽出していく
if rep_1[i] == 1
rep_1[i] = temp_1
elseif rep_1[i] == 2
rep_1[i] = temp_2
elseif rep_1[i] == 3
rep_1[i] = temp_3
elseif rep_1[i] == 4
rep_1[i] = temp_4
elseif rep_1[i] == 5
rep_1[i] = temp_5
elseif rep_1[i] == 6
rep_1[i] = temp_6
elseif rep_1[i] == 7
rep_1[i] = temp_7
elseif rep_1[i] == 8
rep_1[i] = temp_8
end
if rep_2[i] == 1
rep_2[i] = temp_1
elseif rep_2[i] == 2
rep_2[i] = temp_2
elseif rep_2[i] == 3
rep_2[i] = temp_3
elseif rep_2[i] == 4
rep_2[i] = temp_4
elseif rep_2[i] == 5
rep_2[i] = temp_5
elseif rep_2[i] == 6
rep_2[i] = temp_6
elseif rep_2[i] == 7
rep_2[i] = temp_7
elseif rep_2[i] == 8
rep_2[i] = temp_8
end
if rep_3[i] == 1
rep_3[i] = temp_1
elseif rep_3[i] == 2
rep_3[i] = temp_2
elseif rep_3[i] == 3
rep_3[i] = temp_3
elseif rep_3[i] == 4
rep_3[i] = temp_4
elseif rep_3[i] == 5
rep_3[i] = temp_5
elseif rep_3[i] == 6
rep_3[i] = temp_6
elseif rep_3[i] == 7
rep_3[i] = temp_7
elseif rep_3[i] == 8
rep_3[i] = temp_8
end
if rep_4[i] == 1
rep_4[i] = temp_1
elseif rep_4[i] == 2
rep_4[i] = temp_2
elseif rep_4[i] == 3
rep_4[i] = temp_3
elseif rep_4[i] == 4
rep_4[i] = temp_4
elseif rep_4[i] == 5
rep_4[i] = temp_5
elseif rep_4[i] == 6
rep_4[i] = temp_6
elseif rep_4[i] == 7
rep_4[i] = temp_7
elseif rep_4[i] == 8
rep_4[i] = temp_8
end
if rep_5[i] == 1
rep_5[i] = temp_1
elseif rep_5[i] == 2
rep_5[i] = temp_2
elseif rep_5[i] == 3
rep_5[i] = temp_3
elseif rep_5[i] == 4
rep_5[i] = temp_4
elseif rep_5[i] == 5
rep_5[i] = temp_5
elseif rep_5[i] == 6
rep_5[i] = temp_6
elseif rep_5[i] == 7
rep_5[i] = temp_7
elseif rep_5[i] == 8
rep_5[i] = temp_8
end
if rep_6[i] == 1
rep_6[i] = temp_1
elseif rep_6[i] == 2
rep_6[i] = temp_2
elseif rep_6[i] == 3
rep_6[i] = temp_3
elseif rep_6[i] == 4
rep_6[i] = temp_4
elseif rep_6[i] == 5
rep_6[i] = temp_5
elseif rep_6[i] == 6
rep_6[i] = temp_6
elseif rep_6[i] == 7
rep_6[i] = temp_7
elseif rep_6[i] == 8
rep_6[i] = temp_8
end
if rep_7[i] == 1
rep_7[i] = temp_1
elseif rep_7[i] == 2
rep_7[i] = temp_2
elseif rep_7[i] == 3
rep_7[i] = temp_3
elseif rep_7[i] == 4
rep_7[i] = temp_4
elseif rep_7[i] == 5
rep_7[i] = temp_5
elseif rep_7[i] == 6
rep_7[i] = temp_6
elseif rep_7[i] == 7
rep_7[i] = temp_7
elseif rep_7[i] == 8
rep_7[i] = temp_8
end
if rep_8[i] == 1
rep_8[i] = temp_1
elseif rep_8[i] == 2
rep_8[i] = temp_2
elseif rep_8[i] == 3
rep_8[i] = temp_3
elseif rep_8[i] == 4
rep_8[i] = temp_4
elseif rep_8[i] == 5
rep_8[i] = temp_5
elseif rep_8[i] == 6
rep_8[i] = temp_6
elseif rep_8[i] == 7
rep_8[i] = temp_7
elseif rep_8[i] == 8
rep_8[i] = temp_8
end
end
# -
rep_1
# +
#fig, ax = subplots(figsize=(10, 5));
figure(figsize=(10,5),)
plot((1:size(rep_8,1))*0.025,rep_8, c="#7f7f7f", label="ID:8");
plot((1:size(rep_7,1))*0.025,rep_7, c="#e377c2", label="ID:7");
plot((1:size(rep_6,1))*0.025,rep_6, c="#8c564b", label="ID:6");
plot((1:size(rep_5,1))*0.025,rep_5, c="#9467bd", label="ID:5");
plot((1:size(rep_4,1))*0.025,rep_4, c="#d62728", label="ID:4");
plot((1:size(rep_3,1))*0.025,rep_3, c="#2ca02c", label="ID:3");
plot((1:size(rep_2,1))*0.025,rep_2, c="#ff7f0e", label="ID:2");
plot((1:size(rep_1,1))*0.025,rep_1, c="#1f77b4", label="ID:1");
legend(loc="upper right")
#xlim([0, 300])
#ylim([0, 130])
xlabel("time [ns]",fontsize=20);
ylabel("temperature [K]",fontsize=20);
xticks(fontsize=10);
yticks(fontsize=10);
savefig("param_4idlDIHED.png", dpi=350, bbox_inches="tight");
# -
| 4idl_dihedral/06_grest_production/.ipynb_checkpoints/parameter_4idlDIHED-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Task-1 Supervised Machine Learning
#
#
#
# # <NAME>
#Importing all the required libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# # Importing Data
url = "http://bit.ly/w-data"
s_data = pd.read_csv(url)
print("Data imported successfully")
s_data.head(10)
s_data.info()
s_data.describe()
s_data.isnull().sum()
# Let's plot our data points on 2-D graph to eyeball our dataset and see if we can manually find any relationship between the data. We can create the plot with the following script:
# Plotting the distribution of scores
x=s_data['Hours']
y=s_data['Scores']
plt.scatter(x,y,color='r',marker='o')
plt.grid(linewidth=1,linestyle="--",color='b')
plt.title("Hours vs percentage")
plt.xlabel('Hours')
plt.ylabel('Percentage score')
# # From the graph above, we can clearly see that there is a positive linear relation between the number of hours studied and percentage of score.
# # Preparing the data
# The next step is to divide the data into "attributes" (inputs) and "labels" (outputs).
X = s_data.iloc[:, :-1].values
y = s_data.iloc[:, 1].values
print(X)
print(y)
# Now that we have our attributes and labels, the next step is to split this data into training and test sets. We'll do this by using Scikit-Learn's built-in train_test_split() method:
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y,test_size=0.2, random_state=0)
# # Now Training the algorithm
# We have split our data into training and testing sets and now is finally the time to train our algorithm.
from sklearn.linear_model import LinearRegression
regressor = LinearRegression()
regressor.fit(X_train, y_train)
# +
line = regressor.coef_*X+regressor.intercept_
# Plotting for the test data
plt.scatter(X, y)
plt.plot(X, line);
plt.show()
# -
# # Making Predictions
print(X_test) # Testing data - In Hours
y_pred = regressor.predict(X_test) # Predicting the scores
s_data = pd.DataFrame({'Actual': y_test, 'Predicted': y_pred})
s_data
hours =[[9.25]]
own_pred = regressor.predict(hours)
print("No of Hours = {}".format(hours))
print("Predicted Score = {}".format(own_pred[0]))
# # Evaluating the Model
from sklearn import metrics
print('Mean absolute error:', metrics.mean_absolute_error(y_test,y_pred))
| Task 1- Supervised Machine Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# + language="javascript"
# IPython.OutputArea.prototype._should_scroll = function(lines) {
# return false;
# }
# -
import cv2
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
from glob import glob
# +
import lanelines
import roadplane
from compgraph import CompGraph, CompGraphRunner
import nxpd
nxpd.nxpdParams['show'] = 'ipynb'
# -
from newlanespipeline import computational_graph as cg
from newlanespipeline import parameters as cg_params
# ## Initialization
cm = np.load('serialize/camera_matrix.npy')
dc = np.load('serialize/dist_coefs.npy')
# +
CANVAS_SZ = (500, 1500)
OFFSET_X = 100
OFFSET_Y = 0
straight_images_files = ('test_images/straight_lines1.jpg', 'test_images/straight_lines2.jpg')
straight_images = [lanelines.open_image(f, convert_to_rgb=True) for f in straight_images_files]
straight_images_undist = [cv2.undistort(im, cm, dc) for im in straight_images]
warp_src = roadplane.define_flat_plane_on_road(straight_images_undist, x_offset=0)
warp_src[1, 0] += 8 # <- a hack
warp_dst = lanelines.get_rectangle_corners_in_image(CANVAS_SZ, offset_x=OFFSET_X, offset_y=OFFSET_Y)
M = cv2.getPerspectiveTransform(warp_src, warp_dst)
Minv = cv2.getPerspectiveTransform(warp_dst, warp_src)
# -
test_images = [lanelines.open_image(f, convert_to_rgb=True) for f in glob('test_images/*.jpg')]
test_images_undist = [cv2.undistort(im, cm, dc) for im in test_images]
# ## Pipeline
# +
runner = CompGraphRunner(cg, frozen_tokens=cg_params)
nxpd.draw(runner.token_manager.to_networkx())
# -
# ## Experiments
# +
last_y = CANVAS_SZ[1]
PIXELS_PER_METER = 50
plt.figure(figsize=(20, 5))
for i, im in enumerate(test_images_undist):
runner.run(image=im, M=M)
plt.subplot(1, 8, i+1)
plt.imshow( runner['all_thresholds'])
_ = plt.axis('off')
poly_y, poly_x_left, poly_x_right = lanelines.get_lane_polynomials_points(
runner['warped'],
runner['p_coefs_left'],
runner['p_coefs_right']
)
plt.plot(poly_x_left, poly_y, color='c')
plt.plot(poly_x_right, poly_y, color='c')
curv_left = lanelines.curvature_poly2(runner['p_coefs_left'], last_y)
curv_right = lanelines.curvature_poly2(runner['p_coefs_right'], last_y)
plt.title('{:.2f}, {:.2f}'.format(curv_left / PIXELS_PER_METER, curv_right / PIXELS_PER_METER))
# +
plt.figure(figsize=(10, 15))
plt.imshow(runner['warped'])
for i in range(60):
plt.axhline(i * 20, color='c')
for i in range(25):
plt.axvline(i * 20, color='c')
# +
pix_lane = 13.5 * 20
pix_dash = 9 * 20
pix_lane, pix_dash
# +
m_lane = 3.7
m_dash = 3.
p_x = m_lane / pix_lane
p_y = m_dash / pix_dash
p_x, p_y
# -
| 11_pixel_to_meter_ratios.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
3
2 7 4
2
# +
def func(n, input_val):
lst = list(map(int, input_val.split()))
lst.sort()
total_plants = 0
last_added = None
for index, value in enumerate(lst):
# check i-1
found = False
# check i+1
if index + 1 < n and lst[index+1] == value+1:
found = True
elif index - 1 >= 0 and lst[index-1] == value-1:
found = True
elif last_added != None and last_added + 1 == value:
found = True
if not found:
total_plants += 1
last_added = value+1
return total_plants
T = int(input())
for _ in range(T):
n = int(input())
lst = input()
print(func(n, lst))
# -
assert func(3, '2 4 7') == 2
assert func(1, '2') == 1
assert func(3, '1 2 3') == 0
| CodeChef/DEADEND.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Computer Vision models zoo
# + hide_input=true
from fastai.gen_doc.nbdoc import *
from fastai.vision.models.darknet import Darknet
from fastai.vision.models.wrn import wrn_22, WideResNet
# -
# On top of the models offered by [torchvision](https://pytorch.org/docs/stable/torchvision/models.html), the fastai library has implementations for the following models:
#
# - Darknet architecture, which is the base of [Yolo v3](https://pjreddie.com/media/files/papers/YOLOv3.pdf)
# - Unet architecture based on a pretrained model. The original unet is described [here](https://arxiv.org/abs/1505.04597), the model implementation is detailed in [`models.unet`](/vision.models.unet.html#vision.models.unet)
# - Wide resnets architectures, as introduced in [this article](https://arxiv.org/abs/1605.07146).
# + hide_input=true
show_doc(Darknet)
# -
# Create a Darknet with blocks of sizes given in `num_blocks`, ending with `num_classes` and using `nf` initial features. Darknet53 uses `num_blocks = [1,2,8,8,4]`.
# + hide_input=true
show_doc(WideResNet)
# -
# Each group contains `N` blocks. `start_nf` the initial number of features. Dropout of `drop_p` is applied in between the two convolutions in each block. The expected input channel size is fixed at 3.
# Structure: initial convolution -> `num_groups` x `N` blocks -> final layers of regularization and pooling
# The first block of each group joins a path containing 2 convolutions with filter size 3x3 (and various regularizations) with another path containing a single convolution with a filter size of 1x1. All other blocks in each group follow the more traditional res_block style, i.e., the input of the path with two convs is added to the output of that path.
#
# In the first group the stride is 1 for all convolutions. In all subsequent groups the stride in the first convolution of the first block is 2 and then all following convolutions have a stride of 1. Padding is always 1.
# + hide_input=true
show_doc(wrn_22)
# -
# This is a [`WideResNet`](/vision.models.wrn.html#WideResNet) with `num_groups=3`, `N=3`, `k=6` and `drop_p=0.`.
| docs_src/vision.models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Sky Brightness in LSST
# %pylab inline
import astropy.units as u
# ## Read Opsim Cadence
# See opsim db [summary table docs](http://lsst.org/scientists/simulations/opsim/summary-table-column-descriptions-v335).
import sqlite3
import astropy.table
conn = sqlite3.connect('/Data/DESC/minion_1016_sqlite.db')
c = conn.cursor()
# Fetch 1000 rows of r-band observations.
c.execute("""SELECT airmass,dist2Moon,moonAlt,moonPhase,filtSkyBrightness
FROM summary WHERE filter = 'r' LIMIT 1000""")
opsim = astropy.table.Table(
np.array(c.fetchall()),
names=('airmass', 'dist2Moon', 'moonAlt', 'moonPhase','filtSkyBrightness'))
conn.close()
# ## Predict Sky Brightness
# See [specsim atmosphere docs](http://specsim.readthedocs.org/en/latest/config.html#atmosphere)
# and [speclite filter docs](http://speclite.readthedocs.org/en/latest/filters.html). The moon's contribution to the sky brightness is calculated using Based on Krisciunas and Schaefer, “A model of the brightness of moonlight”, PASP, vol. 103, Sept. 1991, p. 1033-1039.
import speclite.filters
import specsim.atmosphere
import specsim.config
cfg = specsim.config.load_config('desi')
atm = specsim.atmosphere.initialize(cfg)
def predict(airmass, dist_to_moon, moon_altitude, moon_illumination, filter_name='sdss2010-r'):
"""Predict the sky brightness
Parameters
----------
airmass : float
Airmass at the field center of the visit.
dist_to_moon : float
Distance from the field center to the moon's center on the sky, in radians.
moon_altitude : float
Altitude of the Moon taking into account the elevation of the site, in radians.
moon_illumination : float
Percent illumination of the Moon (0=new, 100=full).
filter_name : str
Name of the filter to use for calculating the sky brightness.
Returns
-------
float
AB magnitude of the sky in the specified filter.
"""
atm.airmass = airmass
atm.moon.moon_zenith = (np.pi / 2 - moon_altitude) * u.rad
atm.moon.separation_angle = dist_to_moon * u.rad
# Convert illumination percentage to temporal phase in the range [0,1]
# with 0 = full moon, 1 = new moon.
atm.moon.moon_phase = np.arccos(2 * (moon_illumination / 100.) - 1) / np.pi
filt = speclite.filters.load_filter(filter_name)
wlen = cfg.wavelength
return filt.get_ab_magnitude(atm.surface_brightness * u.arcsec ** 2, wlen)
# ## Compare OpSim with SpecSim
r_predicted = []
for obs in opsim:
r_predicted.append(predict(obs['airmass'], obs['dist2Moon'], obs['moonAlt'], obs['moonPhase']))
r_predicted = np.array(r_predicted)
plt.plot(opsim['filtSkyBrightness'], r_predicted, 'x')
plt.xlim(20, 21.4);
# Hmm, I was expecting better agreement. It looks like there are subsets of observations with tight correlations. What opsim parameters define these different subsets? Try coloring points using additional opsim vars.
plt.scatter(opsim['filtSkyBrightness'], r_predicted, lw=0., s=15., c=opsim['airmass'])
plt.colorbar().set_label('airmass')
plt.xlim(20, 21.4);
plt.scatter(opsim['filtSkyBrightness'], r_predicted, lw=0., s=15., c=opsim['dist2Moon'])
plt.colorbar().set_label('moon separation (rad)')
plt.xlim(20, 21.4);
plt.scatter(opsim['filtSkyBrightness'], r_predicted, lw=0., s=15., c=opsim['moonAlt'])
plt.colorbar().set_label('moon alt (rad)')
plt.xlim(20, 21.4);
plt.scatter(opsim['filtSkyBrightness'], r_predicted, lw=0., s=15., c=opsim['moonPhase'])
plt.colorbar().set_label('moon phase')
plt.xlim(20, 21.4);
# It looks like the moon phase is the best predictor of discrepancies between specsim and opsim.
| examples/notebooks/SkyBrightnessCalculator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Observation in finite experimental time-window of processes of random onset and duration: Simulation
# ### by <NAME> and <NAME> (Last updated on 4/3/2019)
# Suppose that we got time trace of signal and we want to measure the mean dwell time (duration of events) to extract the kinetic information. The dynamic signal could be due to protein binding/unbinding or protein conformational changes. At the beginning or end of the time window, we might find incomplete (pre-existing or unfinished) events. If the time window is getting shorter, we have higher chances of finding those incomplete events. <br/>
#
# Considering these circumstances matter when you have a limited ovservation window due to issues such as, <br/>
# * if the kinetic rates governing the transition are very slow and you cannot obtain a long trajectory
# * time window is limited by technical issues including photo-bleaching, degredation of protein or surface chemistry at room temperature
# * you already got large amount data but it is not long enough, and you want to extract information out of it.
#
# We observe individual binding and unbinding events. <br/>
# $\Delta$t = duration of bound state time <br/>
# $t_b$ = time at binding occured <br/>
# $t_u$ = time at unbinding occured <br/>
# $\tau$ = an experimental time-window of fixed duration <br/>
#
# Consequently, we cannot observe $t_b$ and $t_u$ for every bound state we do observe. Thus, every observed bound state belongs to one of four classes: <br/>
# I. $t_b$ < 0 < $t_u$ < $\tau$ (Binding occured before we start observing. Unbinding occured before we finish observing) <br/>
# II. 0 < $t_b$ < $t_u$ < $\tau$ (Binding occured after we start observing. Unbinding occured before we finish observing) <br/>
# III. 0 < $t_b$ < $\tau$ < $t_u$ (Binding occured after we start observing. Unbinding occured after we finish observing) <br/>
# IV. $t_b$ < 0 < $\tau$ < $t_u$ (Binding occured before we start observing. Unbinding occured before we finish observing) <br/>
#
# Below, I run a simulation.
# ## Import libraries
# +
# Import libraries
from __future__ import division, print_function, absolute_import
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import sys
from scipy.stats import geom
sys.path.append("../APC/APC")
import config
from my_funcs import generate_trace, find_dwell
# -
# ## Below, I simulate traces that have only binary state signal (unbound or bound state). Real experimental data contain noise, and suppose that we processeded the data using a state determining algorithm.
# +
# User parameters
t_b = 5 # Mean bound time in frame (Poisson dwell time)
t_u = 5*t_b # Mean unbound time in frame (Poisson dwell time)
tau = 100 # Size of a window
n_window = 100000 # Number of window
t_total = tau * n_window # Total lenth of trace
trace_total = generate_trace(t_total, t_b, t_u)
traces = trace_total.reshape((n_window, tau))
# Plot a trace
fig, ((ax0, ax1, ax2, ax3)) = plt.subplots(1,4, figsize=(20, 5))
ax0.plot(traces[0], 'k')
ax1.plot(traces[1], 'k')
ax2.plot(traces[2], 'k')
ax3.plot(traces[3], 'k')
ax1.set_title('Window size = %d, Mean dwell = %.2f, # traces = %d' %(tau, t_b, n_window))
# -
# ## Dwell time distribution of each class
# +
#I have to figure out the binning issue. Data do not have value at zero which brings the exponential down
# List of dwell times for each class
dwell_class1 = []
dwell_class2 = []
dwell_class3 = []
dwell_class4 = []
for trace in traces:
t1, t2, t3, t4 = find_dwell(trace)
dwell_class1.extend(t1)
dwell_class2.extend(t2)
dwell_class3.extend(t3)
dwell_class4.extend(t4)
dwell_class1 = np.array(dwell_class1)
dwell_class2 = np.array(dwell_class2)
dwell_class3 = np.array(dwell_class3)
# Mean of class
mean_t1 = np.mean(dwell_class1)-0.5
mean_t2 = np.mean(dwell_class2)-0.5
mean_t3 = np.mean(dwell_class3)-0.5
# +
# Histogram
bins1 = np.linspace(0, max(dwell_class1), 20)
bins2 = np.linspace(0, max(dwell_class2), 20)
bins3 = np.linspace(0, max(dwell_class3), 20)
norm1 = len(dwell_class1)*(bins1[1]-bins1[0])
norm2 = len(dwell_class2)*(bins2[1]-bins2[0])
norm3 = len(dwell_class3)*(bins3[1]-bins3[0])
x1 = np.linspace(0, max(dwell_class1), 200)
x2 = np.linspace(0, max(dwell_class2), 200)
x3 = np.linspace(0, max(dwell_class3), 200)
y1 = np.exp(-x1/mean_t1)/(mean_t1)*norm1
y2 = np.exp(-x2/mean_t2)/(mean_t2)*norm2
y3 = np.exp(-x3/mean_t3)/(mean_t3)*norm3
y1_true = np.exp(-(x1)/t_b)/t_b*norm1
y2_true = np.exp(-(x2)/t_b)/t_b*norm2
y3_true = np.exp(-(x3)/t_b)/t_b*norm3
# Histogram of each class
fig, ((ax0, ax1, ax2), (ax3, ax4, ax5)) = plt.subplots(2,3, figsize=(20, 10))
ax0.hist(dwell_class1, bins1, color='k', histtype='step', lw=1)
ax0.plot(x1, y1, 'r')
ax0.plot(x1, y1_true, 'b')
ax0.set_title('Class 1, # event = %d' %(len(dwell_class1)))
ax1.hist(dwell_class2, bins2, color='k', histtype='step', lw=1)
ax1.plot(x2, y2, 'r')
ax1.plot(x2, y2_true, 'b')
ax1.set_title('Class 2, # event = %d' %(len(dwell_class2)))
ax2.hist(dwell_class3, bins3, color='k', histtype='step', lw=1)
ax2.plot(x3, y3, 'r')
ax2.plot(x3, y3_true, 'b')
ax2.set_title('Class 3, # event = %d' %(len(dwell_class3)))
ax3.hist(dwell_class1, bins1, color='k', histtype='step', lw=1, log=True)
ax3.plot(x1, y1, 'r')
ax3.plot(x1, y1_true, 'b')
ax3.set_title('Class 1, t_mean = %.2f (R), t_true = %.2f (B)' %(mean_t1, t_b))
ax4.hist(dwell_class2, bins2, color='k', histtype='step', lw=1, log=True)
ax4.plot(x2, y2, 'r')
ax4.plot(x2, y2_true, 'b')
ax4.set_title('Class 2, t_mean = %.2f (R), t_true = %.2f (B)' %(mean_t2, t_b))
ax5.hist(dwell_class3, bins3, color='k', histtype='step', lw=1, log=True)
ax5.plot(x3, y3, 'r')
ax5.plot(x3, y3_true, 'b')
ax5.set_title('Class 3, t_mean = %.2f (R), t_true = %.2f (B)' %(mean_t3, t_b))
# -
# ## Estimation of true mean from experimental mean and window size
# r = (r * tau - 2 + (r * tau + 2) * exp(-r * tau)) / (r * tau - 1 + exp(-r * tau)) / mean(t)
# +
dr = 0.0001
r = np.arange(dr, 2/mean_t2, dr)
LHS = r
RHS = 1/mean_t2 * (r*tau-2 + (r*tau+2)*np.exp(-r*tau))/(r*tau - 1 + np.exp(-r*tau))
r_correction = r[np.argmin(abs(LHS-RHS))]
# Plot a trace
fig, (ax0, ax1) = plt.subplots(1,2, figsize=(20,5))
ax0.plot(r, LHS, 'b')
ax0.plot(r, RHS, 'r')
ax0.set_title('LHS (B), RHS (R)')
ax0.set_xlabel('r')
ax1.plot(r, RHS-LHS, 'k')
ax1.axhline(y=0, ls=':', c='k')
ax1.set_xlabel('r')
ax1.set_ylabel('RHS-LHS')
ax1.set_title('r_correction = %.2f, t_correction = %.2f, t_true = %.2f, t_mean = %.2f' %(r_correction, 1/r_correction, t_b, mean_t2))
# -
# ## Estimation with varying window size
# +
# User parameters
t_b = 10 # Mean bound time in frame (Poisson dwell time)
t_u = 10*t_b # Mean unbound time in frame (Poisson dwell time)
tau_range = np.arange(4*t_b, 40*t_b, 2*t_b) # Size of a window
n_window = 1000 # Number of window
n_dataset = 100
est_mean1 = np.zeros((len(tau_range), n_dataset))
est_mean2 = np.zeros((len(tau_range), n_dataset))
est_mean3 = np.zeros((len(tau_range), n_dataset))
est_corr = np.zeros((len(tau_range), n_dataset))
for i, tau in enumerate(tau_range):
print(tau)
t_total = tau * n_window # Total lenth of trace
for j in range(n_dataset):
trace_total = generate_trace(t_total, t_b, t_u)
traces = trace_total.reshape((n_window, tau))
dwell_class1 = []
dwell_class2 = []
dwell_class3 = []
for trace in traces:
t1, t2, t3, t4 = find_dwell(trace)
dwell_class1.extend(t1)
dwell_class2.extend(t2)
dwell_class3.extend(t3)
# Mean estimation
est_mean1[i][j] = np.mean(dwell_class1)-0.5
est_mean2[i][j] = np.mean(dwell_class2)-0.5
est_mean3[i][j] = np.mean(dwell_class3)-0.5
# Correction estimation
dr = 0.0001
r = np.arange(dr, 2/est_mean2[i][j], dr)
LHS = r
RHS = 1/est_mean2[i][j] * (r*tau-2 + (r*tau+2)*np.exp(-r*tau))/(r*tau - 1 + np.exp(-r*tau))
r_corr = r[np.argmin(abs(LHS-RHS))]
est_corr[i][j] = 1/r_corr
# +
# Plot the result
fig, ((ax0, ax1, ax2, ax3), (ax4, ax5, ax6, ax7)) = plt.subplots(2,4, figsize=(20,10))
# Class2
ax0.errorbar(tau_range, np.mean(est_mean2, axis=1), yerr = np.std(est_mean2, axis=1), color='k', fmt='o')
ax0.axhline(y=t_b, color='k', linestyle='dotted', lw=1)
ax0.set_xticks(tau_range)
ax0.set_xlabel('Window size')
ax0.set_ylabel('Mean estimator2')
ax0.set_title('Class 2, Mean+/-SD (N = %d)' %(n_dataset))
# Class2_corrected
ax1.errorbar(tau_range, np.mean(est_corr, axis=1), yerr = np.std(est_corr, axis=1), color='k', fmt='o')
ax1.axhline(y=t_b, color='k', linestyle='dotted', lw=1)
ax1.set_xticks(tau_range)
ax1.set_xlabel('Window size')
ax1.set_ylabel('Corrected mean estimator2')
ax1.set_title('Class 2, Mean+/-SD (N = %d)' %(n_dataset))
# Class1
ax2.errorbar(tau_range, np.mean(est_mean1, axis=1), yerr = np.std(est_mean1, axis=1), color='k', fmt='o')
ax2.axhline(y=t_b, color='k', linestyle='dotted', lw=1)
ax2.set_xticks(tau_range)
ax2.set_xlabel('Window size')
ax2.set_ylabel('Mean estimator1')
ax2.set_title('Class 1, Mean+/-SD (N = %d)' %(n_dataset))
# Class3
ax3.errorbar(tau_range, np.mean(est_mean3, axis=1), yerr = np.std(est_mean3, axis=1), color='k', fmt='o')
ax3.axhline(y=t_b, color='k', linestyle='dotted', lw=1)
ax3.set_xticks(tau_range)
ax3.set_xlabel('Window size')
ax3.set_ylabel('Mean estimator3')
ax3.set_title('Class 3, Mean+/-SD (N = %d)' %(n_dataset))
# Class2
ax4.errorbar(tau_range/t_b, (np.mean(est_mean2, axis=1)-t_b)/t_b*100, yerr = (np.std(est_mean2, axis=1))/t_b*100, color='k', fmt='o')
ax4.axhline(y=0, color='k', linestyle='dotted', lw=1)
ax4.set_xticks(tau_range/t_b)
ax4.set_xlabel('Window size ratio')
ax4.set_ylabel('Mean estimator error (%)')
ax4.set_title('# window = %d' %(n_window))
# Class2_corrected
ax5.errorbar(tau_range/t_b, (np.mean(est_corr, axis=1)-t_b)/t_b*100, yerr = (np.std(est_corr, axis=1))/t_b*100, color='k', fmt='o')
ax5.axhline(y=0, color='k', linestyle='dotted', lw=1)
ax5.set_xticks(tau_range/t_b)
ax5.set_xlabel('Window size ratio')
ax5.set_ylabel('Corrected mean estimator error (%)')
ax5.set_title('# window = %d' %(n_window))
# Class1
ax6.errorbar(tau_range/t_b, (np.mean(est_mean1, axis=1)-t_b)/t_b*100, yerr = (np.std(est_mean1, axis=1))/t_b*100, color='k', fmt='o')
ax6.axhline(y=0, color='k', linestyle='dotted', lw=1)
ax6.set_xticks(tau_range/t_b)
ax6.set_xlabel('Window size ratio')
ax6.set_ylabel('Mean estimator error 1 (%)')
ax6.set_title('# window = %d' %(n_window))
# Class3
ax7.errorbar(tau_range/t_b, (np.mean(est_mean3, axis=1)-t_b)/t_b*100, yerr = (np.std(est_mean3, axis=1))/t_b*100, color='k', fmt='o')
ax7.axhline(y=0, color='k', linestyle='dotted', lw=1)
ax7.set_xticks(tau_range/t_b)
ax7.set_xlabel('Window size ratio')
ax7.set_ylabel('Mean estimator error 3 (%)')
ax7.set_title('# window = %d' %(n_window))
# -
| notebooks/.ipynb_checkpoints/01_dwell_finite_window_simul-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <NAME> (18/424191/PA/18296)
# Pada tugas ini, saya akan melakukan prediksi terhadap headline berita yang ada apakah tergolong clickbait atau tidak, kemudian dilanjutkan dengan perhitungan evaluation metrics untuk menilai performa dari model yang telah dibuat. Namun, berbeda dari biasanya dimana model di-generate menggunakan library, kali ini tugas ini akan menggunakan cara manual untuk melakukan prediksi dengan menggunakan metode (Multinomial) Naive Bayes.
import pandas as pd #meng-import library pandas untuk meng-handle dataframe
import re # meng-import library re untuk men-generate regex
# ## 1. Training Phase
#
# Pada tugas kali ini terdapat 2 dataset yaitu annotated_okezone.csv sebagai training set dan annotated_fimela.csv sebagai test set. Pada training set ini kita akan melakukan perhitungan probabilitas guna membantu prediksi class/label dari berita. Pertama-tama, agar dataframe lebih efektif, akan dilakukan drop terhadap kolom 'label'
train=pd.read_csv("annotated_okezone.csv")
train.head()
train=train.drop("label",axis=1)
train
# Di bawah ini merupakan fungsi untuk melakukan text pre-processing terhadap dataset. Di fungsi tersebut berisi penghapusan pattern (seperti digit, punctuation) menggunakan regex dan juga proses stopwords removal. Namun, setelah melakukan beberapa kali percobaan, didapatkan hasil yang tidak lebih baik dari dataset tanpa dilakukan pre-processing, sehingga bagian ini tidak akan saya jalankan.
# +
#stop words list
stop_words=set(['yang', 'untuk', 'pada', 'ke', 'para', 'namun', 'menurut', 'antara', 'dia', 'dua', 'ia', 'seperti', 'jika',
'sehingga', 'kembali', 'dan', 'tidak', 'ini', 'karena', 'kepada', 'oleh', 'saat', 'harus', 'sementara', 'setelah',
'belum', 'kami', 'sekitar', 'bagi', 'serta', 'di', 'dari', 'telah', 'sebagai', 'masih', 'hal', 'ketika', 'adalah',
'itu', 'dalam', 'bisa', 'bahwa', 'atau', 'hanya', 'kita', 'dengan', 'akan', 'juga', 'ada', 'mereka', 'sudah',
'saya', 'terhadap', 'secara', 'agar', 'lain', 'anda', 'begitu', 'mengapa', 'kenapa', 'yaitu', 'yakni', 'daripada',
'itulah', 'lagi', 'maka', 'tentang', 'demi', 'dimana', 'kemana', 'pula', 'sambil', 'sebelum', 'sesudah', 'supaya',
'guna', 'kah', 'pun', 'sampai', 'sedangkan', 'selagi', 'sementara', 'tetapi', 'apakah', 'kecuali', 'sebab',
'selain', 'seolah', 'seraya', 'seterusnya', 'tanpa', 'agak', 'boleh', 'dapat', 'dsb', 'dst', 'dll', 'dahulu',
'dulunya', 'anu', 'demikian', 'tapi', 'ingin', 'juga', 'nggak', 'mari', 'nanti', 'melainkan', 'oh', 'ok',
'seharusnya', 'sebetulnya', 'setiap', 'setidaknya', 'sesuatu', 'pasti', 'saja', 'toh', 'ya', 'walau', 'tolong',
'tentu', 'amat', 'apalagi', 'bagaimanapun'])
# pattern
pat_dig=r'[0-9]+'
pat_punc=r'[,\.%\"\'&:\[\]]+'
pattern=r'|'.join((pat_dig,pat_punc))
# +
from nltk.tokenize import WordPunctTokenizer
token=WordPunctTokenizer()
def preprocessing(df):
clean=[]
for index,row in df.iterrows():
pre_regex=re.sub(pat_punc,'',row.title)
pre_lower=pre_regex.lower()
pre_token=token.tokenize(pre_lower)
# pre_filter=[w for w in pre_token if not w in stop_words]
# clean.append(pre_lower.strip())
clean.append((" ".join(pre_token).strip()))
return clean
# proses=preprocessing(train)
# train.title=pd.DataFrame(proses)
# train
# -
# ### 1.1. Create Vocabulary
# Selanjutnya, untuk keperluan Naive Bayes nantinya, kita akan membangun vocabulary terlebih dahulu. Pertama-tama kita akan menggabungkan semua kalimat pada kolom 'title' dalam suatu list. Kemudian dengan fungsi vocab_count() kita akan membangun dictionary untuk menyimpan kata beserta frekuensi kemunculan. Selanjutnya list tadi akan displit untuk kemudian masuk ke perulangan untuk dihitung frekuensinya, jika kata tersebut belum ada di dictionary 'vocab_temp' maka akan dibuat dengan frekuensi 1, jika sudah dilakukan increment. Setelah itu return 'vocab_temp'.
# Kemudian, karena pada tugas ini akan diambil hanya 1000 kata dengan frekuensi tertinggi, maka dilakukan sorting terlebih dahulu pada dictionary 'count' dan kemudian pada 'vocab_list' akan diambil 1000 kata pertama yang dijadikan sebagai vocabulary
# +
bulk=" ".join(train['title'])
# fungsi untuk menghitung kata
def vocab_count(text):
vocab_temp = dict()
words = text.split()
for w in words:
if w in vocab_temp:
vocab_temp[w] += 1
else:
vocab_temp[w] = 1
return vocab_temp
count=vocab_count(bulk)
vocab={key: value for key, value in sorted(count.items(), reverse=True, key=lambda x: x[1])}
vocab_list=list(vocab)[:1000]
vocab_list
# -
# ### 1.2. Estimate Parameter for Multinomial Naive Bayes Model
#
# Proses selanjutnya adalah kita akan menghitung nilai conditional probability dari semua vocabulary.
#
# Pertama-tama, kita akan menghitung banyak nilai 0 dan 1 pada kolom 'label_score' dan kemudian diubah menjadi dictionary serta disimpan pada 'count_label'
count_label=dict(train['label_score'].value_counts())
count_label
# Kemudian, dari nilai count_label tersebut kita akan menghitung nilai P(c=0) dan P(c=1). Pertama kita akan menghitung banyak dokumen pada training set dengan perulangan dan memasukkan pada variabel 'jumlah'.
#
# Selanjutnya , akan dihitung P(c)=N(c)/N, dimana N(c) adalah banyaknya dokumen dengan kelas c dan N adalah variabel 'jumlah'
# +
prob_label={}; jumlah=0
for i in count_label.keys():
jumlah=jumlah+count_label[i]
for i in count_label.keys():
prob_label[i]=count_label[i]/jumlah
prob_label
# -
# Selanjutnya, kita akan menyiapkan suatu nested dictionary yang berisi 1000 vocabulary dari 'vocab_list' sebagai key dari dictionary pertama, dan untuk value dari dictionary pertama berisi dictionary kedua yang menunjukkan frekuensi/banyak kata di vocabulary pada masing-masing kelas, yaitu count(Wi | c=0) dan count(Wi | c=1). Namun sebelum menghitungnya, kita akan menginisialisasi dictionary seperti pada berikut untuk mengisi frekuensi semua kata pada setiap kelas adalah 0.
count_prob={i : {} for i in vocab_list}
for i in vocab_list:
for j in count_label.keys():
count_prob[i][j]=0
count_prob
# for i in range(len(train)):
# for j in vocab_list:
# count_prob[j][str(train.iloc[i]['label_score'])]=count_prob[j][str(train.iloc[i]['label_score'])]+train.iloc[i]['title'].count(j)
# count_prob
# Langkah selanjutnya adalah membagi dataset 'train' ke dua buah dataset berdasarkan nilai 'label_score'-nya. Ini dilakukan agar saat proses perhitungan frekuensi kata, waktu run cell tidak terlalu lama. Dataset dibagi menjadi dataframe 'class_zero' untuk headline dengan label=0 dan 'class_one' untuk label=1.
class_zero=train[train['label_score']==0]
class_zero
class_one=train[train['label_score']==1]
class_one
# Berikutnya adalah proses perhitungan frekuensi kata dari setiap kelas. Konsepnya adalah akan dilakukan iterasi untuk setiap document/row, dilanjutkan dengan melakukan increment frekuensi pada setiap kata di vocabulary jika ditemukan kata pada 'title' yang cocok dengan kata yang ada di 'vocab_list'. Untuk penyocokkan kata, dimanfaatkan regex re.findall() dan negative lookbehind (<?<!\S) guna mendapatkan pola kata yang benar-benar sesuai dengan yang dicari (pada tugas ini, konteksnya adalah kata pada vocabulary). Karena ini sifatnya melakukan iterasi untuk penyocokkan 1000 vocabulary pada setiap document, proses ini memakan waktu kurang lebih 5-10 menit. Proses ini dilakukan pada dua dataset 'class_zero' dan 'class_one' dan akan menghasilkan nilai untuk count(Wi | c)
for i in range(len(class_zero)):
for j in vocab_list:
count_prob[j][0]=count_prob[j][0]+len(re.findall(r'(?<!\S)'+re.escape(j)+r'(?!\S)',class_zero.iloc[i]['title']))
for i in range(len(class_one)):
for j in vocab_list:
count_prob[j][1]=count_prob[j][1]+len(re.findall(r'(?<!\S)'+re.escape(j)+r'(?!\S)',class_one.iloc[i]['title']))
count_prob
# Selanjutnya, kita akan menghitung total keseluruhan kata yang terdaftar dalam vocabulary untuk masing-masing kelas, yaitu nilai dari count(c=0) dan count(c=1) yang akan digunakan untuk menghitung probabilitas nantinya. Sama seperti sebelumnya, kita memanfaatkan dictionary 'count_words' untuk menyimpan nilainya. Untuk itu akan dilakukan iterasi untuk setiap label kelas. Guna menghitung jumlah, terdapat variabel 'jlh' sebagai penyimpan total kata per kelas. Setiap iterasi tersebut akan melakukan iterasi untuk setiap kata di vocabulary 'vocab_list'. Kemudian dengan memanfaatkan frekuensi 'count_prob' yang telah dihitung sebelumnya, akan dilakukan penjumlahan untuk setiap kata (yang ada di dalam 'vocab_list') kemudian disimpan dalam dictionary 'count_words' sebelum lanjut ke label berikutnya.
count_words={}
for j in count_label.keys():
jlh=0
for i in vocab_list:
jlh=jlh+count_prob[i][j]
count_words[j]=jlh
count_words
# Setelah mendapatkan semua elemen yang digunakan untuk perhitungan Multinomial Naive Bayes, kita akan langsung masuk ke rumus conditional probabilities. Sebelumnya, akan disiapkan terlebih dahulu nested dictionary 'cond_prob' untuk menyimpan probabilitas setiap kata untuk setiap kelasnya.
# Rumus conditional probabilities :
# P(Wi|C)=count(Wi|c)/count(c) dimana count(Wi|c) adalah frekuensi kemunculan kata Wi (pada vocab_list) untuk setiap kelas (count_prob) dan count(c) adalah total keseluruhan kata yang ada pada kelas c.
#Multinomial Naive Bayes
cond_prob={i : {} for i in vocab_list}
for i in count_prob.keys():
for j in count_label.keys():
cond_prob[i][j]=count_prob[i][j]/count_words[j]
cond_prob
# ### 1.3. Add-1 Smoothing
#
# Kita sudah menghitung conditional probability untuk semua kata di vocabulary untuk setiap kelasnya pada 'cond_prob'. Namun, sebagaimana kita tahu ada beberapa kata di vocab_list yang probabilitas nya 0 dimana nanti saat melakukan perhitungan probabilitas pada test set akan menghasilkan nilai 0. Oleh karena itu, kita akan mengaplikasikan konsep add-1 smoothing pada nilai probabilitas yang sudah kita hitung agar setidaknya semua kata dianggap muncul minimal 1 kali.
# Konsepnya sama seperti perhitungan sebelumnya, hanya rumusnya yang berbeda, yaitu :
# P(Wi|C)=count(Wi|c)+1/count(c)+len(V) dimana len(V) adalah banyaknya vocabulary pada vocab_list.
#Add-1 Smoothing
cond_prob={i : {} for i in vocab_list}
for i in count_prob.keys():
for j in count_label.keys():
cond_prob[i][j]=(count_prob[i][j]+1)/(count_words[j]+len(vocab_list))
cond_prob
# ## 2. Test Phase
#
# Kita sudah selesai menghitung conditional probability dari setiap kata di vocabulary kita dengan menggunakan konsep add-1 smoothing. Sekarang kita akan beralih ke dataset test untuk menguji performa dari perhitungan kita. Sama seperti training phase, kita akan menghapus kolom 'label' dan tidak melakukan text pre-processing karena akan menurunkan nilai akurasinya.
test=pd.read_csv("annotated_fimela.csv")
test.head()
test=test.drop("label",axis=1)
test.head()
# +
# proses_test=preprocessing(test)
# test.title=pd.DataFrame(proses_test)
# test
# -
# Selanjutnya kita akan mengekstrak kata-kata yang muncul di test set namun tidak ada di vocabulary 'vocab_list'. Ini dilakukan dengan menggabungkan semua kata di test set, dihitung frekuensinya dengan fungsi 'vocab_count' dan kemudian kata unik tersebut dicocokkan kemunculannya pada 'vocab_list' dan jika merupakan vocabulary baru akan disimpan di list 'vocab_test'. Karena kata-kata di 'vocab_test' merupakan kata yang tidak ada di 'vocab_list', maka probabilitasnya adalah 0 dan tentunya akan sangat mempengaruhi hasil prediksi nanti. Oleh karena itu, digunakan perhitungan add-1 smoothing pada kata-kata tersebut dan disimpan pada dictionary 'cond_prob_test'. Isi dari dictionary 'cond_prob_test' akan digabungkan dengan dictionary utama dari perhitungan 'cond_prob' dengan menggunakan method update()
bulk_test=" ".join(test['title'])
count_test=vocab_count(bulk_test)
vocab_test=[word for word in count_test.keys() if word not in vocab_list]
vocab_test
#Add-1 Smoothing
cond_prob_test={i : {} for i in vocab_test}
for i in vocab_test:
for j in count_label.keys():
cond_prob_test[i][j]=1/(count_words[j]+len(vocab_list))
cond_prob_test
cond_prob.update(cond_prob_test)
len(cond_prob)
# ### 2.1. Count Probabilities of Each Document in Both Labels
#
# Sebelum memprediksi label class sesuai perhitungan, kita akan menghitung terlebih dahulu probabilitas suatu document terhadap label c=0 dan c=1. Pertama kita akan memasukkan semua document tersebut ke dalam list 'temp_list' agar memudahkan kita dalam melakukan perhitungan. Di dalam list tersebut akan dibuat lagi list (nested list) untuk memecah setiap kata dalam document menjadi seperti token.
temp_list=[]
for j in range(len(test)):
temp_list.append(test.iloc[j]['title'].split())
temp_list
# Kita akan memulai perhitungan probabilitasnya. Konsepnya adalah kita melakukan perkalian probabilitas untuk setiap word di dalam document dan kemudian dikalikan dengan probabilitas dari label/classnya. Jadi kita melakukan iterasi untuk setiap kelasnya, kemudian untuk setiap kelas tersebut dilakukan iterasi lagi untuk akses ke setiap kata di 'temp_list'. Setelah itu, akan dilakukan perkalian probabilitas 'cond_prob' untuk setiap katanya hingga kata di document tersebut habis dan disimpan dalam 'prob_test'. Sebelum melanjutkan ke document berikutnya, nilai 'prob_test' akan dikalikan dengan 'prob_label'. Setelah itu nilai akhir 'prob_test' setiap document akan ditambahkan ke dalam list 'prob'. Nilai dari list 'prob' untuk setiap kelasnya akan dimasukkan ke test set dengan nama kolom 'prob_(0/1)'.
# +
for i in count_label.keys():
prob=[]
for j in temp_list:
prob_test=1
for k in j:
prob_test=prob_test*cond_prob[k][i]
#print(prob_test,cond_prob[k][i])
prob_test*=prob_label[i]
prob.append(prob_test)
#print(prob)
test["prob_"+str(i)]=prob
test.head()
# -
# ### 2.2. Predict Most Likely Class (CMAP)
#
# Kita sudah selesai menghitung probabilitas setiap document untuk setiap kelasnya. Saatnya kita memprediksi CMAP dari setiap document. Untuk memudahkan saya menginisiasi list 'label' untuk menyimpan nilai dari prediksi.
# Jadi, kita akan melakukan iterasi untuk setiap row/documentnya dan kemudian dibandingkan nilai antara probabilitas P(c=0|doc-x) dan P(c=1|doc-x). Jika nilai dari prob_1 > prob_0 maka, label akan ditambahkan dengan nilai 1, sebaliknya akan ditambahkan dengan nilai 0. Setelah itu dibuat kolom baru 'label_predicted' untuk menyimpan nilai list 'label'
# +
label=[]
for index,row in test.iterrows():
if row['prob_1']>row['prob_0']:
label.append(1)
else:
label.append(0)
test['label_predicted']=label
test.head()
# -
# ## 3. Evaluation Phase
# ### 3.1. Contigency Table Creation
#
# Kita sudah mendapatkan hasil prediksi kita pada tahap test. Selanjutnya kita akan melakukan evaluasi terhadap perhitungan model Naive Bayes kita. Salah satu perhitungan evaluasi adalah dengan membangun confusion matrix atau contigency table. Unuk menghitung tersebut kita akan menghitung nilai dari TP, FP, FN, dan TN terlebih dahulu. Setelah mendapatkan nilainya, kita membangun confusion matrix dengan menggunakan nested list.
# +
TP=0; FP=0; FN=0; TN=0;
for index,row in test.iterrows():
if (row['label_score']==0 and row['label_predicted']==0):
TP+=1
elif (row['label_score']==0 and row['label_predicted']==1):
FN+=1
elif (row['label_score']==1 and row['label_predicted']==0):
FP+=1
else:
TN+=1
confusion_matrix=[[TP,FP],[FN,TN]]
confusion_matrix
# -
# ### 3.2. Calculate Four Evaluation Metrics
#
# Setelah memperoleh nilai TP, FP, FN, TN kita dapat membangun 4 metriks evaluasi yaitu accuracy, precision, recall, dan F1 Score dengan rumus dibawah.
accuracy=(TP+TN)/(TP+FP+FN+TN)
precision=TP/(TP+FP)
recall=TP/(TP+FN)
F1=2*precision*recall/(precision+recall)
print("Accuracy: ",accuracy)
print("Precision: ",precision)
print("Recall: ",recall)
print("F1 Score: ",F1)
# ### 3.3. Compare Microaveraging vs Macroaveraging Results
#
# Tahap terakhir adalah menghitung nilai dari microaveraging dan macroaveraging. Sebelum memulai perhitungan, kita harus membangun tabel macroaverage dulu yaitu performa setiap kelasnya dan tabel microaverage.
# Untuk macroaverage dibuat tabel 'class_zero' untuk performa dari label 0 dan 'class_one' untuk performa dari label 1. Untuk microaverage merupakan penjumlahan dari dari baris-kolom yang bersesuaian antara 'class_zero' dan 'class_one'.
class_zero=[[TP,FP],[FN,TN]]
class_one=[[TN,FN],[FP,TP]]
micro_table=[[TP+TN,FP+FN],[FN+FP,TN+TP]]
print("Class-0: ",class_zero)
print("Class-1: ",class_one)
print("Micro Avg. Table : ",micro_table)
# Selanjutnya adalah perhitungan nilai microaveraging dan macroaveraging.
# Nilai dari microaveraging maupun macroaveraging akan dihitung untuk 4 metrik evaluasi.
# Perhitungan ini akan mengeluarkan hasil seperti di bawah.
# +
#Accuracy
acc_zero=(class_zero[0][0]+class_zero[1][1])/(sum(class_zero[0])+sum(class_zero[1]))
acc_one=(class_one[0][0]+class_one[1][1])/(sum(class_one[0])+sum(class_one[1]))
macro_avg_acc=(acc_zero+acc_one)/2
micro_avg_acc=(micro_table[0][0]+micro_table[1][1])/(sum(micro_table[0])+sum(micro_table[1]))
print("Accuracy Class-0 : ",acc_zero,"\nAccuracy Class-1 : ",acc_one)
print("Macroaveraged Accuracy: ",macro_avg_acc)
print("Microaveraged Accuracy: ",micro_avg_acc)
# +
#Precision
pre_zero=(class_zero[0][0]/sum(class_zero[0]))
pre_one=(class_one[0][0]/sum(class_one[0]))
macro_avg_precision=(pre_zero+pre_one)/2
micro_avg_precision=((micro_table[0][0]/sum(micro_table[0])))
print("Precision Class-0 : ",pre_zero,"\nPrecision Class-1 : ",pre_one)
print("Macroaveraged Precision: ",macro_avg_precision)
print("Microaveraged Precision: ",micro_avg_precision)
# +
#Recall
rec_zero=(class_zero[0][0]/(class_zero[0][0]+class_zero[1][0]))
rec_one=(class_one[0][0]/(class_one[0][0]+class_one[1][0]))
macro_avg_recall=(rec_zero+rec_one)/2
micro_avg_recall=((micro_table[0][0]/(micro_table[0][0]+micro_table[1][0])))
print("Recall Class-0 : ",rec_zero,"\nRecall Class-1 : ",rec_one)
print("Macroaveraged Recall: ",macro_avg_recall)
print("Microaveraged Recall: ",micro_avg_recall)
# +
#F1-Score
p_zero=(class_zero[0][0]/sum(class_zero[0]))
p_one=(class_one[0][0]/sum(class_one[0]))
r_zero=(class_zero[0][0]/(class_zero[0][0]+class_zero[1][0]))
r_one=(class_one[0][0]/(class_one[0][0]+class_one[1][0]))
f1_zero=2*p_zero*r_zero/(p_zero+r_zero)
f1_one=2*p_one*r_one/(p_one+r_one)
macro_avg_f1=(f1_zero+f1_one)/2
micro_avg_f1=2*micro_avg_precision*micro_avg_recall/(micro_avg_precision+micro_avg_recall)
print("F1 Class-0 : ",f1_zero,"\nF1 Class-1 : ",f1_one)
print("Macroaveraged F1-Score: ",macro_avg_f1)
print("Microaveraged F1-Score: ",micro_avg_f1)
# -
# ## Analisis
#
# Jika kita lihat dari hasil 4 evaluasi metriks diatas, semua microaveragenya sama yaitu 0.65. Namun, jika kita lihat satu per satu, nilai macroaveragenya bervariasi dan berbeda untuk setiap evaluasi metriks. Saya juga menampilkan perhitungan setiap evaluasi metriks per kelasnya sebelum dihitung rata-ratanya dalam macroaverage. Pada accuracy, semuanya sama bahkan dari perhitungan per kelasnya. Pada evaluation metrics lainnya seperti precision, recall, dan f1-score, antara macro-average dan micro-averagenya memiliki nilai yang hampir sama yaitu 60-70%. Tapi, jika kita lihat lebih dalam ke nilai setiap kelasnya, dapat kita lihat, misal pada class 0, presisinya sebesar 70% (dari 100 berita yang diprediksi tidak clickbait, 70 diantaranya benar) namun recall sebesar 34% (dari 100 berita yang tidak clickbait, 34 diantaranya benar). Ini dapat kita jelaskan bahwa model ini memberikan hasil yang baik ketika memprediksi class 1 (clickbait), namun tidak begitu baik ketika memprediksi class 0 (non clickbait). Ini juga sangat terlihat pada nilai macro- atau micro-averaged yang cukup kecil di label 0 dan cukup tinggi di label 1. <br>
# Namun, secara keseluruhan dapat dilihat bahwa nilai macro- dan micro-averaged saling mendekati dan dapat disimpulkan bahwa model Multinomial Naive Bayes ini cukup baik dalam melakukan prediksi pada dataset ini meskipun hasilnya masih belum maksimal, yaitu sekitar 60-70%. Selain itu dari nilai macro- dan micro-averaged-nya, dapat kita simpulkan bahwa data ini termasuk balance data.
| naivebayes_luis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="BzVbdvmd85ef" colab_type="text"
# ## **Neural network concept**
#
# ---
# ### **`Shallo neural network`**
# ---
# "shallow" neural networks is a term used to describe neural network that usually have a only one hidden layer as opposed to deep neural network which has several hidden layers, often of various types.
#
# [<img src="https://qphs.fs.quoracdn.net/main-qimg-257af1d7bfdc2d7c1c4f4c30366a3c77.webp" width="70%" />](https://qphs.fs.quoracdn.net/main-qimg-257af1d7bfdc2d7c1c4f4c30366a3c77.webp)
#
# ---
# ### **`Perceptron`**
# ---
# * Perceptron is the basic unit of a neural network.<br/>
#
# * The perceptron is a network takes a number of inputs,
# carries out some processing on these inputs and produces as output.
#
# * Artificial Neural Networks (ANN) are comprised of a large number of simple elements , called neurons, each of which makes simple decisions. Together, the neurons can provide accurate answers to some complex problems, such as natural language processing, computer vision, and AI.
#
# **Figure** :
#
# [<img src="https://qph.fs.quoracdn.net/main-qimg-1a057e476f5c069f825fa198780c211b.webp" width="50%"/>](https://qph.fs.quoracdn.net/main-qimg-1a057e476f5c069f825fa198780c211b.webp)
#
# * **`What are significance of hidden layer ?`**
#
# **Answer** : without a hidden layer . it's like single neuron i.e. perceptron or like logistic or svm
# single neuron can only implement a linear boundary.
# if we want a our network to classify a non linear boundary then we have to use hidden layer.
#
# * **`what are the hyper parameters ?`**
#
# **Answer** : number of the hidden layer and number of neurons in each hidden layer
# will be the hyper parameters
#
# * Increasing number of hidden layer leads to overfitting.
#
#
# ---
# ### **`Universal approximation theorem`**
# ---
#
# **_Defination_** :
# A feedforward network with a single layer is sufficient to represent any function, but the layer may be infeasibly large and may fail to learn and generalize correctly.
#
# That simply means that we can solve any non trivial function or continous function using only single hidden layer which contain n number of neurons in it with help of activation function.<br/>
#
# `Statement`:
# > Introducing non-linearity via an activation function allows us to approximate any function. It’s quite simple, really. — <NAME>
#
# It has been proven that non-trivial function (x³ + x² - x -1) using a single hidden layer and 6 neurons <br/>
#
# Type of activation function :
# 1. Linear or Identity Activation Function
# 2. Non-linear Activation Function
# * Logistic Function or sigmoid function
# * Tanh (hyperbolic tangent Activation) function
# * ReLu ( Rectified Linear Units )
# * Leaky ReLu
# * Softmax function
#
# We can use any activation function , now let us choose activation function as ReLu to represent the above non trivial function.
#
# [<img src="https://miro.medium.com/max/1400/1*6c0BjULsvVe5zqlkB_Vb3w.png
# " width="50%"/>](https://miro.medium.com/max/1400/1*6c0BjULsvVe5zqlkB_Vb3w.png)
#
# We chose x³+x²-x -1 as my target function. Using only ReLU max(0,x), we iteratively tried different combinations of ReLUs until we had an output that roughly resembled the target.
#
# Here are the results I achieved taking the weighted sum of 3 ReLUs.<br/>
#
# [<img src="https://miro.medium.com/max/1400/1*qt4SaoYphChAreRTDJewIw.png" width="50%"/>](https://miro.medium.com/max/1400/1*qt4SaoYphChAreRTDJewIw.png)
#
# So combining 3 ReLU functions is like training a network of 3 hidden neurons. Here are the equations I used to generate these charts.
#
# [<img src="https://miro.medium.com/max/1000/1*fdICiWJocvOTJPoRrhek_Q.png" width="50%"/>](https://miro.medium.com/max/1000/1*fdICiWJocvOTJPoRrhek_Q.png)
#
# Each neuron’s output equals ReLU wrapped around the weighted input wx + b.
#
# I found I could shift the ReLU function left and right by changing the bias and adjust the slope by changing the weight. I combined these 3 functions into a final sum of weighted inputs (Z) which is standard practice in most neural networks.
#
# The negative signs in Z represent the final layer’s weights which I set to -1 in order to “flip” the graph across the x-axis to match our concave target. After playing around a bit more I finally arrived at the following 7 equations that, together, roughly approximate x³+x²-x -1.
#
# [<img src="https://miro.medium.com/max/1400/1*lihbPNQgl7oKjpCsmzPDKw.png" width="50%"/>](https://miro.medium.com/max/1400/1*lihbPNQgl7oKjpCsmzPDKw.png).
#
# Hard-coding my weights into a real network
# Here is a diagram of a neural network initialized with my fake weights and biases. If you give this network a dataset that resembles x³+x²-x-1, it should be able approximate the correct output for inputs between -2 and 2.
#
# [<img src="https://miro.medium.com/max/1400/1*RWcNXtQSrIVoiw99bkcA8w.png" width="50%"/>](https://miro.medium.com/max/1400/1*RWcNXtQSrIVoiw99bkcA8w.png)
#
# That last statement, approximate the correct output for any input between -2 and 2, is key. The Universal Approximation Theorem states that a neural network with 1 hidden layer can approximate any continuous function for inputs within a specific range.
#
# Refer this : [https://towardsdatascience.com/can-neural-networks-really-learn-any-function-65e106617fc6](https://towardsdatascience.com/can-neural-networks-really-learn-any-function-65e106617fc6)
#
# ---
# ## **`Forward propagation in neural networks`**
# ---
# A learning algorithm/model finds out the parameters (weights and biases) with the help of forward propagation and backpropagation.
#
# The input data is fed in the forward direction through the network. Each hidden layer accepts the input data, processes it as per the activation function and passes to the successive layer.
#
# The feed-forward network helps in forward propagation.<br/>
#
# At each neuron in a hidden or output layer, the processing happens in two steps:
#
# * `Preactivation` : it is a weighted sum of inputs i.e. the linear transformation of weights w.r.t to inputs available. Based on this aggregated sum and activation function the neuron makes a decision whether to pass this information further or not.<br/>
# Eg : We compute F(x) = (x * wx + y * wy+b) at each neuron in hidden layer
#
# * `Activation` : the calculated weighted sum of inputs is passed to the activation function. An activation function is a mathematical function which adds non-linearity to the network. There are four commonly used and popular activation functions — sigmoid, hyperbolic tangent(tanh), ReLU and Softmax.
#
# [<img src="https://www.dspguide.com/graphics/F_26_6.gif" width="60%"/>](https://www.dspguide.com/graphics/F_26_6.gif)
#
# * we apply Activation function on these function (f(x)) , which will give a some output.
#
# if Activation function output is not equal to Target output then we apply weight updated.
#
# * `Weight update` :
#
# ---
#
# alpha : learning rate (Larget alpha that means we are making larger changes to the weight.)
# t : target output
# i : Data points
# p(i) : output of activation function
#
# w(new) = w(old) + alpha * (t - p(i)) * i <br/>
#
# ---
#
# Weights on the edge is decided based on the :<br/>
#
# number of input neuron i.e. (d) and
# number of neurons on hidden layer i.e.( n )
# hence we have (d * n) number of edge with corresponding weight.
#
# ---
#
# The data can be generated using make_moons() function of sklearn.datasets module. The total number of samples to be generated and noise about the moon’s shape can be adjusted using the function parameters.
#
# import numpy as np
# import matplotlib.pyplot as plt
# import matplotlib.colors
# from sklearn.datasets import make_moonsnp.random.seed(0)data, labels = make_moons(n_samples=200,noise = 0.04,random_state=0)
# print(data.shape, labels.shape)color_map = matplotlib.colors.LinearSegmentedColormap.from_list("", ["red","yellow"])
# plt.scatter(data[:,0], data[:,1], c=labels, cmap=my_cmap)
# plt.show()
#
# <br/>
#
# [<img src="https://miro.medium.com/max/924/1*F50x2COgQ8LySWV0LtdFLw.png" width="40%"/>](https://miro.medium.com/max/924/1*F50x2COgQ8LySWV0LtdFLw.png)
#
# Here, 200 samples are used to generate the data and it has two classes shown in red and green color.
#
# Now, let us see the neural network structure to predict the class for this binary classification problem. Here, I am going to use one hidden layer with two neurons, an output layer with a single neuron and sigmoid activation function.
#
# [<img src="https://miro.medium.com/max/1192/1*tp73P0isrrfpj8RG-5aH6w.png" width="40%"/>](https://miro.medium.com/max/1192/1*tp73P0isrrfpj8RG-5aH6w.png)
#
# During forward propagation at each node of hidden and output layer preactivation and activation takes place. For example at the first node of the hidden layer, a1(preactivation) is calculated first and then h1(activation) is calculated.
#
# a1 is a weighted sum of inputs. Here, the weights are randomly generated.
#
# a1 = w1 * x1 + w2 * x2 + b1 = 1.76 * 0.88 + 0.40 *(-0.49) + 0 = 1.37 approx and <br/> h1 is the value of activation function applied on a1.
#
# [<img src="https://miro.medium.com/max/576/1*WrkgXLQSjHpzmR_H3xsnCQ.png" width="30%"/>](https://miro.medium.com/max/576/1*WrkgXLQSjHpzmR_H3xsnCQ.png
# )
#
# Similarly
#
# a2 = w3*x1 + w4*x2 + b2 = 0.97 *0.88 + 2.24 *(- 0.49)+ 0 = -2.29 approx and
#
# [<img src="https://miro.medium.com/max/618/1*46xma79g8Gdew_LbT6x2aw.png" width="30%"/>](https://miro.medium.com/max/618/1*46xma79g8Gdew_LbT6x2aw.png)
#
# For any layer after the first hidden layer, the input is output from the previous layer.
#
# a3 = w5*h1 + w6*h2 + b3 = 1.86*0.8 + (-0.97)*0.44 + 0 = 1.1 approx
# and
#
# [<img src="https://miro.medium.com/max/598/1*lCVQROFldjILndKg-pKHxw.png" width="30%"/>](https://miro.medium.com/max/598/1*lCVQROFldjILndKg-pKHxw.png)
#
# So there are 74% chances the first observation will belong to class 1. Like this for all the other observations predicted output can be calculated.
#
# [<img src="https://miro.medium.com/max/1400/1*ts5LSdtkfSsMYS7M0X84Tw.gif" width="60%"/>](https://miro.medium.com/max/1400/1*ts5LSdtkfSsMYS7M0X84Tw.gif)
#
# Refer :
# [https://towardsdatascience.com/forward-propagation-in-neural-networks-simplified-math-and-code-version-bbcfef6f9250](https://towardsdatascience.com/forward-propagation-in-neural-networks-simplified-math-and-code-version-bbcfef6f9250)
#
# ---
# ## **`Activation function`**
# ---
#
# * While building a neural network, one of the mandatory choices we need to make is
# which activation function to use. In fact, it is an unavoidable choice because
# activation functions are the foundations for a neural network to learn and
# approximate any kind of complex and continuous relationship between variables.
# It simply adds non-linearity to the network.
#
# [<img src="https://image.ibb.co/gEmoSQ/mmm_act_function_1.png
# " width="50%"/>](https://image.ibb.co/gEmoSQ/mmm_act_function_1.png
# )
#
# * Activation functions reside within neurons, but not all neurons.
# Hidden and output layer neurons possess activation functions, but input layer neurons do not.
#
# * Activation functions perform a transformation on the input received,
# in order to keep values within a manageable range. Since values in the
# input layers are generally centered around zero and have already been appropriately scaled,
# they do not require transformation. However, these values, once multiplied by weights and
# summed, quickly get beyond the range of their original scale, which is where the activation
# functions come into play, forcing values back within this acceptable range and making them
# useful.
#
# * In order to be useful, activation functions must also be nonlinear and continuously differentiable.
# Nonlinearity allows the neural network to be a universal approximation; As we already discuss. A continuously differentiable function is necessary for gradient-based optimization methods,
# which is what allows the efficient back propagation of errors throughout the network.
#
# **`NOTE`**:
# Inside the neuron:
#
# * An activation function is assigned to the neuron or entire layer of neurons.
# * weighted sum of input values are added up.
# * the activation function is applied to weighted sum of input values and transformation takes place.
# * Activation functions also help normalize the output of each neuron to a
# range between 1 and 0 or between -1 and 1.
# * the output to the next layer consists of this transformed value.
# <br/>
#
# Reference :
#
# * [https://missinglink.ai/guides/neural-network-concepts/7-types-neural-network-activation-functions-right/](https://missinglink.ai/guides/neural-network-concepts/7-types-neural-network-activation-functions-right/)
#
# * [https://towardsdatascience.com/analyzing-different-types-of-activation-functions-in-neural-networks-which-one-to-prefer-e11649256209](https://towardsdatascience.com/analyzing-different-types-of-activation-functions-in-neural-networks-which-one-to-prefer-e11649256209)
#
# * [https://cs231n.github.io/neural-networks-1/#actfun](https://cs231n.github.io/neural-networks-1/#actfun)
#
# * [https://stats.stackexchange.com/questions/218542/which-activation-function-for-output-layer](https://stats.stackexchange.com/questions/218542/which-activation-function-for-output-layer)
#
# ---
# ## **`Types of Activation Functions`**
# ---
#
# The Activation Functions can be basically divided into 3 types-
#
# * Binary Step Function
# * Linear Activation Function
# * Non-linear Activation Functions
#
# <hr/>
#
# 1. #### **Binary Step Function**
#
# A binary step function is a threshold-based activation function. If the input value is above or below a certain threshold, the neuron is activated and sends exactly the same signal to the next layer.
#
# [<img src="https://missinglink.ai/wp-content/uploads/2018/11/binarystepfunction.png" width="40%"/>](https://missinglink.ai/wp-content/uploads/2018/11/binarystepfunction.png)
#
# The problem with a step function is that it does not allow multi-value outputs—for example, it cannot support classifying the inputs into one of several categories.
#
# 2. #### **Linear or Identity Activation Function**
#
# It takes the inputs, multiplied by the weights for each neuron,
# and creates an output signal proportional to the input. In one sense,
# a linear function is better than a step function because it allows multiple outputs,
# not just yes and no.
#
# A linear activation function takes the form: A = cx
#
# [<img src="https://missinglink.ai/wp-content/uploads/2018/11/graphsright.png" width="30%"/>](https://missinglink.ai/wp-content/uploads/2018/11/graphsright.png)
#
# However, a linear activation function has two major problems:
#
# * `Not possible to use backpropagation (gradient descent)` to train the model—the derivative of the function is a constant, and has no relation to the input, X. So it’s not possible to go back and understand which weights in the input neurons can provide a better prediction.
#
# * `All layers of the neural network collapse into one` —with linear activation functions, no matter how many layers in the neural network, the last layer will be a linear function of the first layer (because a linear combination of linear functions is still a linear function). So a linear activation function turns the neural network into just one layer.
#
# #### **`Example`**
# [<img src="https://miro.medium.com/max/1338/1*xcBdSYRndl6dhouE1y0KHg.png" width="50%"/>](https://miro.medium.com/max/1338/1*xcBdSYRndl6dhouE1y0KHg.png)
#
# <br/>
#
# Consider a case where no activation function is used in this network, then from the hidden layer 1 the calculated weighted sum of inputs will be directly passed to hidden layer 2 and it calculates a weighted sum of inputs and pass to the output layer and it calculates a weighted sum of inputs to produce the output. The output can be presented as
# <br/>
#
# [<img src="https://miro.medium.com/max/1068/1*9es-pAjxSJe3tN61B6p64A.png
# " width="50%"/>](https://miro.medium.com/max/1068/1*9es-pAjxSJe3tN61B6p64A.png)
#
# So the output is simply a linear transformation of weights and inputs and it is not adding any non-linearity to the network. Therefore, this network is similar to a linear regression model which can only address the linear relationship between variables i.e. a model with limited power and not suitable for complex problems like image classifications, object detections, language translations, etc.
#
#
# 3. **Non-Linear Activation Functions**
#
# Modern neural network models use non-linear activation functions.
# They allow the model to create complex mappings between the network’s inputs and outputs,
# which are essential for learning and modeling complex data, such as images, video, audio,
# and data sets which are non-linear or have high dimensionality.
#
# Non-linear functions address the problems of a linear activation function:
#
# 1. They allow backpropagation because they have a derivative function which is related to the inputs.
# 2. They allow “stacking” of multiple layers of neurons to create a deep neural network.
#
# ---
# ## **`Type of Nonlinear Activation Functions and How to Choose an Activation Function`**
# ---
#
# 1. Sigmoid / Logistic
# 2. tanh ( hyperbolic tangent)
# 3. ReLu (Rectified linear units)
# 4. Leaky ReLu
# 5. Softmax activation function
#
# * **Sigmoid / Logistic Activation function**
#
# It is a “S” shaped curve with equation : <br/>
#
# [<img src="https://miro.medium.com/max/152/1*2MoOSKaUQyj0_9Q-lnVkEA.png"/>](https://miro.medium.com/max/152/1*2MoOSKaUQyj0_9Q-lnVkEA.png)
#
# [<img src="https://miro.medium.com/max/920/1*qRS650xg0-JrXJPUD_E32w.png" width="30%"/>](https://miro.medium.com/max/920/1*qRS650xg0-JrXJPUD_E32w.png)
#
# **Advantages**
#
# Smooth gradient, preventing “jumps” in output values.
# Output values bound between 0 and 1, normalizing the output of each neuron.
# Clear predictions — For X above 2 or below -2, tends to bring the Y value (the prediction) to the edge of the curve, very close to 1 or 0. This enables clear predictions.
#
# **Disadvantages**
#
# Vanishing gradient—for very high or very low values of X, there is almost no change to the prediction, causing a vanishing gradient problem. This can result in the network refusing to learn further, or being too slow to reach an accurate prediction.
#
# 1. Outputs not zero centered.
# 2. Computationally expensive
#
# * **Tanh (Hyperbolic tangent) Function**
#
# It is similar to logistic activation function with a mathematical equation.<br/>
#
# [<img src="https://miro.medium.com/max/1230/1*ibDdAN-lHnSafuCG1EjP6g.png"/>](https://miro.medium.com/max/1230/1*ibDdAN-lHnSafuCG1EjP6g.png)
#
# [<img src="https://miro.medium.com/max/964/1*IrLb4Z_Mp-cbyCa6bBgsKg.png" width="50%"/>](https://miro.medium.com/max/964/1*IrLb4Z_Mp-cbyCa6bBgsKg.png)
#
# The output ranges from -1 to 1 and having an equal mass on both the sides of zero-axis so it is zero centered function. So tanh overcomes the non-zero centric issue of the logistic activation function. Hence optimization becomes comparatively easier than logistic and it is always preferred over logistic.
#
# But Still, a tanh activated neuron may lead to saturation and cause vanishing gradient problem.
#
# The derivative of tanh activation function.
#
# [<img src="https://miro.medium.com/max/1216/1*ZyQv9ma0lFipjC3vRwSRcw.png" width="50%"/>](https://miro.medium.com/max/1216/1*ZyQv9ma0lFipjC3vRwSRcw.png)
#
# Issues with tanh activation function:
#
# Saturated tanh neuron causes the gradient to vanish.
# Because of e^x, it is highly compute-intensive.
#
# * ReLu (Rectified linear units)
#
# It is the most commonly used function because of its simplicity. It is defined as <br/>
#
# [<img src="https://miro.medium.com/max/764/1*jyyzxadG8Sbqcgfv08ttAw.png" width="50%"/>](https://miro.medium.com/max/764/1*jyyzxadG8Sbqcgfv08ttAw.png)
#
# If the input is a positive number the function returns the number itself and if the input is a negative number then the function returns 0.
#
# [<img src="https://miro.medium.com/max/802/1*E9Az5dBreEwvI5JmlG1dTA.png" width="50%"/>](https://miro.medium.com/max/802/1*E9Az5dBreEwvI5JmlG1dTA.png)
#
# The derivative of ReLu activation function is given as : <br/>
#
# [<img src="https://miro.medium.com/max/636/1*vDV1QZKWD3MoS96Ht2GknA.png" width="50%"/>](https://miro.medium.com/max/636/1*vDV1QZKWD3MoS96Ht2GknA.png
# )
#
# Advantages of ReLu activation function:<br/>
#
# 1. Easy to compute.
# 2. Does not saturate for the positive value of the weighted sum of inputs.
#
# Because of its simplicity, ReLu is used as a standard activation function in CNN.
#
# But still, ReLu is not a zero-centered function.
#
# **`Issues with ReLu activation function`**
#
# ReLu is defined as max(0, w1x1 + w2x2 + …+b) <br/>
#
# Now Consider a case b(bias) takes on (or initialized to) a large negative value then the weighted sum of inputs is close to 0 and the neuron is not activated. <br/> That means the ReLu activation neuron dies now. Like this, up to 50% of ReLu activated neurons may die during the training phase.
#
# To overcome this problem, two solutions can be proposed
#
# 1. Initialize the bias(b) to a large positive value.
# 2. Use another variant of ReLu known as Leaky ReLu.
#
# * **Leaky ReLu**
#
# It was proposed to fix the dying neurons problem of ReLu. It introduces a small slope to keep the update alive for the neurons where the weighted sum of inputs is negative. It is defined as
#
# [<img src="https://miro.medium.com/max/924/1*sy8LauNPCdU6ycPFuqhKag.png" width="50%"/>](https://miro.medium.com/max/924/1*sy8LauNPCdU6ycPFuqhKag.png)
#
# If the input is a positive number the function returns the number itself and if the input is a negative number then it returns a negative value scaled by 0.01(or any other small value).
#
# [<img src="https://miro.medium.com/max/860/1*2U9o7ma_pp4rkalyMs2oXg.png" width="50%"/>](https://miro.medium.com/max/860/1*2U9o7ma_pp4rkalyMs2oXg.png)
#
# The derivative of LeakyReLu is given as:
#
# [<img src="https://miro.medium.com/max/712/1*nyX9El4tLjP2XdtaYsyxJQ.png" width="50%"/>](https://miro.medium.com/max/712/1*nyX9El4tLjP2XdtaYsyxJQ.png)
#
# **Advantages of LeakyReLu** <br/>
# 1. No saturation problem in both positive and negative region.
# 2. The neurons do not die because 0.01x ensures that at least a small gradient will flow through. Although the change in weight will be small but after a few iterations it may come out from its original value.
# 3. Easy to compute.
# 4. Close to zero-centered functions
#
#
# * **Softmax activation function**
#
# * Where other activation function get a input value and transform it.
# where softmax consider the information about the whole set of information that means
# it's a special function , where each element in output depends on the entire set
# of element of the input
#
# * Suppose we have input X , weight and bias
# now in hidden layer , each neuron compute
# 1. a = x*w + b <br/>
# 2. activation function(a)
#
# * Now we can apply any activation function let us apply a softmax function
#
# * Suppose we get value of a = [-0.21 , 0.47 , 1.72 ]
#
# <a href="https://ibb.co/dmnyS3R"><img src="https://i.ibb.co/2ZXBwLT/Screenshot-329.png" alt="Screenshot-329" width="70%" border="0"></a>
#
# * Now we apply softmax function on it.
# * Note : we could even normalize by dividing perticular number by sum of all number , so why are using exponent , so simple answer is exponent ensure positivity i.e. even negative number becomes positive.
# * Property of softmax :
# 1. Normalize the value
# 2. Ranges between 0 to 1
# 3. sum of value = 1
# 4. Probability of each neuron
#
# <a href="https://ibb.co/kJNwHpC"><img src="https://i.ibb.co/0KH6Dkb/Screenshot-327.png" alt="Screenshot-327" width="70%" border="0"></a>
#
# * As we see we get value 0.1,0.2 and 0.7 after apply softmax
# * so now we can say that 0.7 would be highest probable value. As input was "horse" image
# so output probability of horse is 0.7.
#
# <a href="https://ibb.co/7VVn9Lj"><img src="https://i.ibb.co/wCCSvjB/Screenshot-328.png" alt="Screenshot-328" width="70%" border="0"></a>
#
# **Figure describing softmax funtion** :
#
# [<img src="https://developers.google.com/machine-learning/crash-course/images/SoftmaxLayer.svg" width="50%"/>](https://developers.google.com/machine-learning/crash-course/images/SoftmaxLayer.svg)
#
# **Note** :
#
# Sigmoid can be used in output layer , but it only ensure that value will be between 0 to 1 , but sum of these value would not be equal to 1 . which does not satisfying the principle of probability distribution.
#
# Softmax acivation function is mostly used in output layer only and not in hidden layer .
#
# Refer to this stack overflow for better explanation : <br/>
#
# [Why use softmax only in the output layer and not in hidden layers?](https://stackoverflow.com/questions/37588632/why-use-softmax-only-in-the-output-layer-and-not-in-hidden-layers)
#
# ---
#
# **Why derivative/differentiation is used ?**
#
# When updating the curve, to know in which direction and how much to change or update the curve depending upon the slope.That is why we use differentiation in almost every part of Machine Learning and Deep Learning.
#
# [<img src="https://miro.medium.com/max/1400/1*p_hyqAtyI8pbt2kEl6siOQ.png" width="50%"/>](https://miro.medium.com/max/1400/1*p_hyqAtyI8pbt2kEl6siOQ.png)
#
# [<img src="https://miro.medium.com/max/1400/1*n1HFBpwv21FCAzGjmWt1sg.png" width="50%" />](https://miro.medium.com/max/1400/1*n1HFBpwv21FCAzGjmWt1sg.png)
#
# ## **`End Notes: Now which one to prefer?`**
#
# * As a rule of thumb, you can start with ReLu as a general approximator and switch to other functions if ReLu doesn't provide better results.
#
# * For CNN, ReLu is treated as a standard activation function but if it suffers from dead neurons then switch to LeakyReLu.
#
# * Always remember ReLu should be only used in hidden layers.
# For classification, Sigmoid functions(Logistic, tanh, Softmax) and their combinations work well. But at the same time, it may suffer from vanishing gradient problem.
#
# * For RNN, the tanh activation function is preferred as a standard activation function.
#
# * For regression i.e. for real value <br/>
# output activation - Linear activation
# Loss function - MSE
#
# For classification i.e for probability <br/>
# output activation - softmax activation
# Loss function - Cross entropy
#
#
# ---
#
# **Reference** :
#
# 1. [https://www.youtube.com/watch?v=p-XCC0y8eeY](https://www.youtube.com/watch?v=p-XCC0y8eeY)
#
# 2. [https://towardsdatascience.com/activation-functions-neural-networks-1cbd9f8d91d6](https://towardsdatascience.com/activation-functions-neural-networks-1cbd9f8d91d6)
#
# 3. [https://stats.stackexchange.com/questions/218542/which-activation-function-for-output-layer?newreg=c9a3c1990ce2419785c5c368ecb77d2e](https://stats.stackexchange.com/questions/218542/which-activation-function-for-output-layer?newreg=c9a3c1990ce2419785c5c368ecb77d2e
# )
#
# ---
# ## **`So how neural network learn.`**
# ---
#
# * Suppose , consider the output layer where we get the probabillity of each classes to be predicted using softmax activation function.
#
# * Now we have label for perticular input.
# * we compute a cost based on : (predicted - desired output)^2
# * We keep minimizing the cost
#
# <a href="https://ibb.co/pzjv1rb"><img src="https://i.ibb.co/zRVhmHQ/Screenshot-331.png" alt="Screenshot-331" width="50%" border="0"></a>
#
# References:
#
# * [https://www.youtube.com/watch?v=IHZwWFHWa-w&list=PLZHQObOWTQDNU6R1_67000Dx_ZCJB-3pi&index=2](https://www.youtube.com/watch?v=IHZwWFHWa-w&list=PLZHQObOWTQDNU6R1_67000Dx_ZCJB-3pi&index=2)
#
# ---
# ## **`Epoch vs Batch Size vs Iterations vs Learning rate`**
# ---
#
# * ### **Epoch**
#
# One Epoch is when an ENTIRE dataset is passed forward and backward through the neural network only ONCE.
# OR
# Number of epochs is the number of times the whole training data is shown to the network while training.
#
# Since one epoch is too big to feed to the computer at once we divide it in several smaller batches.
#
# **`Why we use more than one Epoch?`**
#
# I know it doesn’t make sense in the starting that — passing the entire dataset through a neural network is not enough. And we need to pass the full dataset multiple times to the same neural network. But keep in mind that we are using a limited dataset and to optimise the learning and the graph we are using Gradient Descent which is an iterative process. So, updating the weights with single pass or one epoch is not enough.
#
# > One epoch leads to underfitting of the curve in the graph (below).
#
# [<img src="https://miro.medium.com/max/1400/1*i_lp_hUFyUD_Sq4pLer28g.png" width="50%" />](https://miro.medium.com/max/1400/1*i_lp_hUFyUD_Sq4pLer28g.png)
#
# As the number of epochs increases, more number of times the weight are changed in the neural network and the curve goes from underfitting to optimal to overfitting curve.
#
# **`So, what is the right numbers of epochs?`**
#
# Unfortunately, there is no right answer to this question. The answer is different for different datasets but you can say that the numbers of epochs is related to how diverse your data is...
#
# * ### **Batch Size**
#
# Total number of training examples present in a single batch.
#
# > Note: Batch size and number of batches are two different things.
#
# **`But What is a Batch?`** <br/>
#
# As I said, you can’t pass the entire dataset into the neural net at once. So, you divide dataset into Number of Batches or sets or parts.
# Just like you divide a big article into multiple sets/batches/parts
#
# * ### **Iterations**
#
# Iterations is the number of batches needed to complete one epoch.
#
# > **Note**: The number of batches is equal to number of iterations for one epoch.
#
# Let’s say we have 2000 training examples that we are going to use .
# We can divide the dataset of 2000 examples into batches of 500 then it will take 4 iterations to complete 1 epoch.<br/>
#
# **Where Batch Size is 500 and Iterations is 4, for 1 complete epoch.**
#
# * ### **Learning rate**
#
# **`Gradient Descent`** <br/>
# It is an iterative optimization algorithm used in machine learning to find the best results (minima of a curve).<br/>
# Gradient means the rate of inclination or declination of a slope.<br/>
# Descent means the instance of descending.<br/>
#
# The algorithm is iterative means that we need to get the results multiple times to get the most optimal result. The iterative quality of the gradient descent helps a under-fitted graph to make the graph fit optimally to the data.
#
# [<img src="https://miro.medium.com/max/1400/1*pwPIG-GWHyaPVMVGG5OhAQ.gif" width="70%"/>](https://miro.medium.com/max/1400/1*pwPIG-GWHyaPVMVGG5OhAQ.gif)
#
# [<img src="https://miro.medium.com/max/1276/0*FA9UmDXdzYzuOpeO.jpg" width="40%"/>](https://miro.medium.com/max/1276/0*FA9UmDXdzYzuOpeO.jpg)
#
# The Gradient descent has a parameter called learning rate. As you can see above (left), initially the steps are bigger that means the learning rate is higher and as the point goes down the learning rate becomes more smaller by the shorter size of steps. Also,the Cost Function is decreasing or the cost is decreasing
#
# **`What are hyperparameters?`**
#
# Hyperparameters are the variables which determines the network structure(Eg: Number of Hidden Units) and the variables which determine how the network is trained(Eg: Learning Rate).
# Hyperparameters are set before training(before optimizing the weights and bias).
#
# Reference :
#
# * [https://towardsdatascience.com/epoch-vs-iterations-vs-batch-size-4dfb9c7ce9c9](https://towardsdatascience.com/epoch-vs-iterations-vs-batch-size-4dfb9c7ce9c9)
#
# * [https://towardsdatascience.com/what-are-hyperparameters-and-how-to-tune-the-hyperparameters-in-a-deep-neural-network-d0604917584a](https://towardsdatascience.com/what-are-hyperparameters-and-how-to-tune-the-hyperparameters-in-a-deep-neural-network-d0604917584a)
#
# ---
# ## **`Back propagation`**
# ---
#
# Back-propagation is the essence of neural net training. It is the practice of fine-tuning the weights of a neural net based on the error rate (i.e. loss) obtained in the previous epoch (i.e. iteration). Proper tuning of the weights ensures lower error rates, making the model reliable by increasing its generalization.
#
# Back-propagation is all about feeding this loss backwards in such a way that we can fine-tune the weights based on which.
#
# The optimization function (Gradient Descent in our example) will help us find the weights that will — hopefully — yield a smaller loss in the next iteration.
#
# Backpropagation is basically minimize the difference between the
# target values we want to learn and output your network currently producing.
#
# #### **Explanation**:
#
# Suppose we have training dataset , which consist of input and label (target)
# Here
#
# Desire output is : t <br/>
# predicted output is : o <br/>
# Loss function : (o-t)^2 <br/>
#
# Step by step explanation :
# * Let us say , we got some output after training neural network
# * now we compute a value of loss function i.e. (predicted-target)^2
# * We use some optimizer like Gradient descent and SGD to minimize the lose or cost function.
# * now we modify the weight (in the region between output and last hidden layer) to get the
# Target output , with help partial derivative by applying chain rule.
# * similary we go one step backword at each step and update the weight.
# * Now again we feed forward a network with new weights , after updating weights
# it seems that error or loss is reducing and we reach the point where gradient is zero i.e. Global minima.
#
# ---
# **During gradient descent** :
#
# [<img src="https://miro.medium.com/max/1400/1*6sDUTAbKX_ICVVAjunCo3g.png" width="40%"/>](https://miro.medium.com/max/1400/1*6sDUTAbKX_ICVVAjunCo3g.png)
#
# Let’s check the derivative.
# - If it is positive, meaning the error increases if we increase the weights, then we should decrease the weight.
# - If it’s negative, meaning the error decreases if we increase the weights, then we should increase the weight.
# - If it’s 0, we do nothing, we reach our stable point.
#
# **Thus as a general rule of weight updates is the delta rule**:
#
# w(new) = w(old) - n*(dL/dw)
# New weight = old weight — learning rate * Derivative
#
# The learning rate is introduced as a constant (usually very small), in order to force the weight to get updated very smoothly and slowly (to avoid big steps and chaotic behaviour). (To remember: Learn slow and steady!)
#
# In a simple matter, we are designing a process that acts like gravity. No matter where we randomly initialize the ball on this error function curve, there is a kind of force field that drives the ball back to the lowest energy level of ground 0.
#
# [<img src="https://miro.medium.com/max/1360/1*dvgzK4beVXBGBELDXP9JpA.png" width="40%"/>](https://miro.medium.com/max/1360/1*dvgzK4beVXBGBELDXP9JpA.png)
#
# ---
#
# 1. Linear Regression is Neural network , then activation function is identity function. <br/>
# Loss function is Mean Squared Error Loss or Mean Squared Logarithmic Error Loss or Mean Absolute Error Loss
#
# 2. Logistic Regression is Neural network , then activation function is sigmoid function. <br/>
# Loss function is Binary Cross-Entropy or Hinge Loss or Squared Hinge Loss.
#
# 3. Multiclass classification is Neural network , then activation function is sigmoid function. <br/>
# Loss function is Multi-Class Cross-Entropy Loss or Sparse Multiclass Cross-Entropy Loss or Kullback Leibler Divergence Loss.
#
#
# **Overall picture**
#
# In order to summarize, Here is what the learning process on neural networks looks like (A full picture):
#
#
# [<img src="https://miro.medium.com/max/1400/1*mi-10dMgdMLQbIHkrG6-jQ.png" width="80%"/>](https://miro.medium.com/max/1400/1*mi-10dMgdMLQbIHkrG6-jQ.png)
#
#
#
#
# **Reference** :
#
# * [https://www.youtube.com/watch?v=GJXKOrqZauk](https://www.youtube.com/watch?v=GJXKOrqZauk)
#
# * [http://home.agh.edu.pl/~vlsi/AI/backp_t_en/backprop.html](http://home.agh.edu.pl/~vlsi/AI/backp_t_en/backprop.html)
#
# * [https://towardsdatascience.com/how-does-back-propagation-in-artificial-neural-networks-work-c7cad873ea7](https://towardsdatascience.com/how-does-back-propagation-in-artificial-neural-networks-work-c7cad873ea7)
#
# * [https://medium.com/datathings/neural-networks-and-backpropagation-explained-in-a-simple-way-f540a3611f5e](https://medium.com/datathings/neural-networks-and-backpropagation-explained-in-a-simple-way-f540a3611f5e)
#
# ## **`Reference`** :
#
# * [https://ml-cheatsheet.readthedocs.io/en/latest/forwardpropagation.html](https://ml-cheatsheet.readthedocs.io/en/latest/forwardpropagation.html)
#
# * [https://playground.tensorflow.org/#activation=tanh&batchSize=10&dataset=circle®Dataset=reg-plane&learningRate=0.03®ularizationRate=0&noise=0&networkShape=4,2&seed=0.58811&showTestData=false&discretize=false&percTrainData=50&x=true&y=true&xTimesY=false&xSquared=false&ySquared=false&cosX=false&sinX=false&cosY=false&sinY=false&collectStats=false&problem=classification&initZero=false&hideText=false](https://playground.tensorflow.org/#activation=tanh&batchSize=10&dataset=circle®Dataset=reg-plane&learningRate=0.03®ularizationRate=0&noise=0&networkShape=4,2&seed=0.58811&showTestData=false&discretize=false&percTrainData=50&x=true&y=true&xTimesY=false&xSquared=false&ySquared=false&cosX=false&sinX=false&cosY=false&sinY=false&collectStats=false&problem=classification&initZero=false&hideText=false)
#
# * [https://openai.com/blog/deep-double-descent/](https://openai.com/blog/deep-double-descent/)
#
# * [http://home.agh.edu.pl/~vlsi/AI/backp_t_en/backprop.html](http://home.agh.edu.pl/~vlsi/AI/backp_t_en/backprop.html)
#
# * [https://ai.googleblog.com/2019/10/learning-to-smell-using-deep-learning.html](https://ai.googleblog.com/2019/10/learning-to-smell-using-deep-learning.html)
#
# * [https://www.youtube.com/watch?v=tIeHLnjs5U8](https://www.youtube.com/watch?v=tIeHLnjs5U8)
#
# * [https://www.youtube.com/playlist?list=PLZHQObOWTQDNU6R1_67000Dx_ZCJB-3pi](https://www.youtube.com/playlist?list=PLZHQObOWTQDNU6R1_67000Dx_ZCJB-3pi)
#
# + id="-qK12Ul-s6UY" colab_type="code" colab={}
| Assignment_11/Neural_networks_concept_ipynb_txt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: rlawjdghek
# language: python
# name: rlawjdghek
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="6rNc_5dWKrcK" outputId="6346e5ca-0848-4da6-801b-549cf55fd0e0"
from google.colab import drive
drive.mount('/content/drive')
# + colab={} colab_type="code" id="lL0PIm37JjTA"
import cv2
import matplotlib.pyplot as plt
import os
import numpy as np
import torch
from torch.nn import Sequential, Conv2d, MaxPool2d, ReLU, BatchNorm2d, Linear
import torchvision
from unicodedata import normalize
from torch.utils.data import Dataset
import glob
import IPython
# -
NUMBER_EPOCHS = 40
HANGEUL_EPOCHS = 20
# + colab={} colab_type="code" id="EnlrEfbBgdkT"
#한글로 시작하는 파일은 모두 삭제
ROOT_PATH = "./data"
for i in os.listdir(ROOT_PATH):
if not i[0].isdigit():
os.remove(os.path.join(ROOT_PATH, i))
# + colab={} colab_type="code" id="lNoq11tcJjTO"
#파일명에 한글이 들어가서 사진을 읽을 때 이 함수로 읽는다
def my_cv_read(filepath):
ff = np.fromfile(filepath, dtype = np.uint8)
gray_img = cv2.imdecode(ff, cv2.IMREAD_GRAYSCALE)
(thresh, im_bw) = cv2.threshold(gray_img, 0, 255, cv2.THRESH_BINARY | cv2.THRESH_OTSU)
return im_bw
# + colab={} colab_type="code" id="SIkl0MHtJjTb"
#신식 번호판인지 구식 번호판인지 분류하는 함수
def version_classifier(image):
if image[10,10] ==0:
#옛날 번호판
return 0
else:
#신식 번호판
return 1
# + colab={} colab_type="code" id="SIfYdXtNJjTl"
#숫자 모델
class conv_block(torch.nn.Module):
def __init__(self, in_channels,out_channels, kernel_size = 3,stride = 1, padding = 1, pool_width = 2):
super(conv_block, self).__init__()
model = []
model.append(Conv2d(in_channels, out_channels, kernel_size = kernel_size, stride = stride, padding = padding))
model.append(BatchNorm2d(out_channels))
model.append(ReLU())
model.append(MaxPool2d((pool_width, pool_width)))
self.model = Sequential(*model)
def forward(self, input_):
return self.model(input_)
class number_classifier(torch.nn.Module):
def __init__(self):
super(number_classifier, self).__init__()
self.block1 = conv_block(1, 16)
self.block2 = conv_block(16, 32)
self.block3 = conv_block(32, 64)
self.block4 = conv_block(64,128)
self.block5 = conv_block(128,256)
self.fc = Linear(128*2*2, 10)
def features(self, input_):
x = self.block1(input_)
x = self.block2(x)
x = self.block3(x)
x = self.block4(x)
return x
def logits(self, features):
return self.fc(features)
def forward(self, input_):
x = self.features(input_)
x = x.reshape(x.shape[0], -1)
return self.logits(x)
class hangeul_classifier(torch.nn.Module):
def __init__(self):
super(hangeul_classifier, self).__init__()
self.block1 = conv_block(1, 16)
self.block2 = conv_block(16,32)
self.block3 = conv_block(32, 64)
self.block4 = conv_block(64,128)
self.fc = Linear(128*2*2,40)
def features(self, input_):
x= self.block1(input_)
x = self.block2(x)
x = self.block3(x)
x = self.block4(x)
return x
def logits(self, features):
return self.fc(features)
def forward(self, input_):
x = self.features(input_)
x = x.reshape(x.shape[0], -1)
return self.logits(x)
# + colab={} colab_type="code" id="K5FDCEPQJjTw"
transform = torchvision.transforms.Compose([torchvision.transforms.Resize(32),torchvision.transforms.ToTensor()])
train_data = torchvision.datasets.MNIST(root = "./", download = True, train=True, transform=transform)
test_data = torchvision.datasets.MNIST(root = "./", download = True, train = False, transform=transform)
# + colab={} colab_type="code" id="L3qU7rb_JjT7"
train_loader =torch.utils.data.DataLoader(train_data, batch_size = 512, shuffle=True)
test_loader = torch.utils.data.DataLoader(test_data, batch_size = 512, shuffle=False)
# + colab={} colab_type="code" id="WCcAokoUJjUH"
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
number_model = number_classifier()
number_model.to(device)
loss_func = torch.nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(number_model.parameters())
# + colab={"base_uri": "https://localhost:8080/", "height": 105} colab_type="code" id="sEykHnvpJjUQ" outputId="0b1ee21a-26c9-4566-f4d9-20e364473b4e"
#MNIST로 훈련시킨 모델
for i in range(NUMBER_EPOCHS):
for image, label in train_loader:
X_train = image.to(device)
y_train = label.to(device)
optimizer.zero_grad()
y_pred = number_model(X_train)
loss = loss_func(y_pred, y_train)
loss.backward()
optimizer.step()
print("epoch : {}".format(i))
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="LjRMSENOJjUa" outputId="d4ffbdbb-ff4c-4a72-cce7-c9818aa8ac06"
number_model.eval()
with torch.no_grad():
total = 0
correct = 0
for image, label in test_loader:
X_test =image.to(device)
y_test = label.to(device)
output = number_model(X_test)
_, output_idx = torch.max(output, 1)
total += output.shape[0]
correct += (output_idx == y_test).sum().float()
print("accuracy : {}".format(100 * correct / total))
# + [markdown] colab_type="text" id="1umR21SWJjUq"
# # 이미지 crop & 숫자 처리
# + colab={} colab_type="code" id="Axm6P1_OJjUr"
#버전에 맞게 이미지 자르는 함수
def crop_image(img):
version = version_classifier(img)
if version ==0:
print("옛날 번호판")
img = img/ 255.0
n1 = img[0:21, 50:87]
n2 = img[0:21, 87 : 112]
n3 = img[20:67, 3:50]
n4 = img[20:67, 50:100]
n5 = img[20:67, 100:150]
n6 = img[20:67, 150:194]
char = img[2:22, 112:150]
else:
print("신식 번호판")
img = img / 255.0
img = (1-img)
n1 = img[10:63,12:40]
n2 = img[10:63, 40:60]
n3 = img[10:63, 95:120]
n4 = img[10:63, 120:140]
n5 = img[10:63, 149:163]
n6 = img[10:63, 163:187]
char = img[10:63, 62:87]
# plt.imshow(n1)
# plt.show()
# plt.imshow(n2)
# plt.show()
# plt.imshow(n3)
# plt.show()
# plt.imshow(n4)
# plt.show()
# plt.imshow(n5)
# plt.show()
# plt.imshow(n6)
# plt.show()
# plt.imshow(char)
# plt.show()
return (n1,n2,n3,n4,n5,n6,char)
# + [markdown] colab_type="text" id="w6iVtLlpnlGu"
# # 한글 분류
# -
#한글 파일 imwrite에러를 방지한 코드
def my_imwrite(filename, img, params=None):
try:
ext = os.path.splitext(filename)[1]
result, n = cv2.imencode(ext, img, params)
if result:
with open(filename, mode='w+b') as f:
n.tofile(f)
return True
else:
return False
except Exception as e:
print(e)
return False
# + colab={} colab_type="code" id="LZixCNXMqLCT"
#한글만 자른 파일 만들기
HANGEUL_PATH = "./cropped_hangeul"
if not os.path.isdir(HANGEUL_PATH):
os.mkdir(HANGEUL_PATH)
if not os.path.isdir(os.path.join(HANGEUL_PATH, "train")):
os.mkdir(os.path.join(HANGEUL_PATH, "train"))
if not os.path.isdir(os.path.join(HANGEUL_PATH, "test")):
os.mkdir(os.path.join(HANGEUL_PATH, "test"))
# + colab={} colab_type="code" id="mP0KeAOfeo7G"
#한글 이미지만 뽑아서 분류후 폴더별로 저장
for file_name in os.listdir(ROOT_PATH):
file_path = os.path.join(ROOT_PATH, file_name)
img = my_cv_read(file_path)
new_file_name = normalize("NFC", file_name)
hangeul_char = new_file_name[2]
hangeul_image = crop_image(img)[-1]
#test폴더에 집어넣기
if np.random.randint(0,100) % 10 == 0:
if not os.path.isdir(os.path.join(HANGEUL_PATH,"test", hangeul_char)):
os.mkdir(os.path.join(HANGEUL_PATH,"test", hangeul_char))
my_imwrite(os.path.join(HANGEUL_PATH,"test", hangeul_char, file_name), hangeul_image)
#train폴더에 집어넣기
else:
if not os.path.isdir(os.path.join(HANGEUL_PATH , "train", hangeul_char)):
os.mkdir(os.path.join(HANGEUL_PATH, "train", hangeul_char))
my_imwrite(os.path.join(HANGEUL_PATH, "train", hangeul_char,file_name), hangeul_image)
IPython.display.clear_output()
# + colab={} colab_type="code" id="yos_no7Ln-2l"
#한글 분류하는 모델저장
class conv_block(torch.nn.Module):
def __init__(self, in_channels,out_channels, kernel_size = 3,stride = 1, padding = 1, pool_width = 2):
super(conv_block, self).__init__()
model = []
model.append(Conv2d(in_channels, out_channels, kernel_size = kernel_size, stride = stride, padding = padding))
model.append(BatchNorm2d(out_channels))
model.append(ReLU())
model.append(MaxPool2d((pool_width, pool_width)))
self.model = Sequential(*model)
def forward(self, input_):
return self.model(input_)
class hangeul_classifier(torch.nn.Module):
def __init__(self):
super(number_classifier, self).__init__()
self.block1 = conv_block(1, 16)
self.block2 = conv_block(16, 32)
self.block3 = conv_block(32, 64)
self.block4 = conv_block(64,128)
self.block5 = conv_block(128,256)
self.fc = Linear(128*2*2, 34)
def features(self, input_):
x = self.block1(input_)
x = self.block2(x)
x = self.block3(x)
x = self.block4(x)
return x
def logits(self, features):
return self.fc(features)
def forward(self, input_):
x = self.features(input_)
x = x.reshape(x.shape[0], -1)
return self.logits(x)
class hangeul_classifier(torch.nn.Module):
def __init__(self):
super(hangeul_classifier, self).__init__()
self.block1 = conv_block(1, 16)
self.block2 = conv_block(16,32)
self.block3 = conv_block(32, 64)
self.block4 = conv_block(64,128)
self.fc = Linear(128*2*2,40)
def features(self, input_):
x= self.block1(input_)
x = self.block2(x)
x = self.block3(x)
x = self.block4(x)
return x
def logits(self, features):
return self.fc(features)
def forward(self, input_):
x = self.features(input_)
x = x.reshape(x.shape[0], -1)
return self.logits(x)
# -
#resize한 한글 이미지를 새로저장
NEW_HANGEUL_PATH = "./new_cropped_hangeul"
HANGEUL_PATH = "./cropped_hangeul"
new_list = glob.glob(HANGEUL_PATH + "/train/" + "/*/" + "*")
if not os.path.isdir(NEW_HANGEUL_PATH):
os.mkdir(NEW_HANGEUL_PATH)
for i in new_list:
new_img = my_cv_read(i)
new_img = cv2.resize(new_img, (32,32))
new_char = i[-9]
if not os.path.isdir(os.path.join(NEW_HANGEUL_PATH, "train", new_char)):
os.makedirs(os.path.join(NEW_HANGEUL_PATH, "train", new_char))
my_imwrite(os.path.join(NEW_HANGEUL_PATH, "train", new_char, i[-11:]), new_img)
#test파일 저장
NEW_HANGEUL_PATH = "./new_cropped_hangeul"
HANGEUL_PATH = "./cropped_hangeul"
new_list = glob.glob(HANGEUL_PATH + "/test/" + "/*/" + "*")
if not os.path.isdir(NEW_HANGEUL_PATH):
os.mkdir(NEW_HANGEUL_PATH)
for i in new_list:
new_img = my_cv_read(i)
new_img = cv2.resize(new_img, (32,32))
new_char = i[-9]
if not os.path.isdir(os.path.join(NEW_HANGEUL_PATH, "test", new_char)):
os.makedirs(os.path.join(NEW_HANGEUL_PATH, "test", new_char))
my_imwrite(os.path.join(NEW_HANGEUL_PATH, "test", new_char, i[-11:]), new_img)
# + colab={} colab_type="code" id="zFdajkt9e1ER"
hangeul_model = hangeul_classifier()
hangeul_model.to(device)
loss_func = torch.nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(hangeul_model.parameters())
# + colab={} colab_type="code" id="AeyV5I--s7T1"
#한글을 레이블로, 레이블을 한글로 만드는 자료구조 및 커스텀 데이터 로드
hangeul_to_idx = {"가":0, "거":1, "고":2, "구":3, "나":4, "너":5, "노":6, "누":7, "다":8, "더":9, "도":10, "두":11, "라":12, "러":13, "로":14,
"루":15, "마":16, "머": 17,"모":18, "무":19, "버":20, "보":21, "부":22, "서":23, "소":24, "수":25, "어":26, "오":27, "우":28, "저":29,
"조":30, "주":31, "하":32, "호":33}
idx_to_hangeul = ["가","거", "고", "구", "나", "너", "노", "누", "다", "더", "도", "두", "라", "러", "로", "루", "마", "머","모", "무", "버", "보", "부",
"서", "소", "수", "어", "오", "우", "저","조", "주", "하", "호"]
class hangeulDataset(Dataset):
def __init__(self, root_dir, train = True, transform = None):
super(hangeulDataset, self).__init__()
self.root_dir = root_dir
self.transform = transform
if train:
self.image_paths = glob.glob(HANGEUL_PATH + "/train/" + "/*/" + "*")
else:
self.image_paths = glob.glob(HANGEUL_PATH + "/test/" + "/*/" + "*")
def __len__(self):
return len(self.image_paths)
def __getitem__(self, idx):
file_path = self.image_paths[idx]
image = my_cv_read(file_path)
image = cv2.resize(image, (32,32))
version = version_classifier(image)
image = transform(image)
label = hangeul_to_idx[file_path[-9]]
return (image, label)
# -
transform = torchvision.transforms.Compose([torchvision.transforms.ToPILImage(),torchvision.transforms.Grayscale(),torchvision.transforms.ToTensor()])
# +
train_data = hangeulDataset(HANGEUL_PATH, train = True, transform = transform)
test_data = hangeulDataset(HANGEUL_PATH, train = False, transform = transform)
train_loader = torch.utils.data.DataLoader(train_data, batch_size = 16, shuffle=True)
test_loader = torch.utils.data.DataLoader(test_data, batch_size = 16, shuffle = True)
# -
for epoch in range(HANGEUL_EPOCHS):
for image, label in train_loader:
X_train = image.to(device)
y_train = label.to(device)
optimizer.zero_grad()
y_pred = hangeul_model(X_train)
loss = loss_func(y_pred, y_train)
loss.backward()
optimizer.step()
print("epoch : {}".format(epoch))
# +
total = 0
correct = 0
hangeul_model.eval()
with torch.no_grad():
for image, label in test_loader:
X_test = image.to(device)
y_test = label.to(device)
output = hangeul_model(X_test)
_, output_idx = torch.max(output, 1)
total += output_idx.shape[0]
correct += (output_idx == y_test).sum().float()
print("accuracy : {}".format(100 * correct / total))
# -
# # 종합
idx = 8
file_name = os.listdir(ROOT_PATH)[idx]
test_image = my_cv_read(os.path.join(ROOT_PATH, file_name))
print("test image", end = " ")
version = version_classifier(test_image)
if version ==0:
print("옛날 번호판")
else:
print("신식 번호판")
plt.imshow(test_image)
plt.show()
# +
number_plate = []
number_model.eval()
hangeul_model.eval()
with torch.no_grad():
cropped_images = crop_image(test_image)
for image in cropped_images[:6]:
resized_number =cv2.resize(image, (32,32)).reshape(1,1,32,32)
resized_number = torch.from_numpy(resized_number).float().to(device)
number_plate.append(torch.max(number_model(resized_number), 1)[1].cpu().detach().numpy()[0])
resized_hangeul = cv2.resize(cropped_images[6], (32,32)).reshape(1,1,32,32)
resized_hangeul = torch.from_numpy(resized_hangeul).float().to(device)
hangeul = idx_to_hangeul[torch.max(hangeul_model(resized_hangeul), 1)[1].cpu().detach().numpy()[0]]
number_plate.insert(2, hangeul)
print("번호판 : ","".join(map(str, number_plate)))
# -
for i in [1,3,5,7,8]:
idx = i
file_name = os.listdir(ROOT_PATH)[idx]
test_image = my_cv_read(os.path.join(ROOT_PATH, file_name))
print("test image", end = " ")
version = version_classifier(test_image)
if version ==0:
print("옛날 번호판")
else:
print("신식 번호판")
plt.imshow(test_image)
plt.show()
number_plate = []
number_model.eval()
hangeul_model.eval()
with torch.no_grad():
cropped_images = crop_image(test_image)
for image in cropped_images[:6]:
resized_number =cv2.resize(image, (32,32)).reshape(1,1,32,32)
resized_number = torch.from_numpy(resized_number).float().to(device)
number_plate.append(torch.max(number_model(resized_number), 1)[1].cpu().detach().numpy()[0])
resized_hangeul = cv2.resize(cropped_images[6], (32,32)).reshape(1,1,32,32)
resized_hangeul = torch.from_numpy(resized_hangeul).float().to(device)
hangeul = idx_to_hangeul[torch.max(hangeul_model(resized_hangeul), 1)[1].cpu().detach().numpy()[0]]
number_plate.insert(2, hangeul)
print("번호판 : ","".join(map(str, number_plate)))
print("\n\n")
| hw5/[Jeongho_Kim]5_week.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.3.1
# language: julia
# name: julia-1.3
# ---
# # Solar-System15-Mixed-Precision
# <ul id="top">
# <li><a href="#Load-packages">
# Load Packages</a></li>
#
# <li><a href="#Computer-properties">
# Computer properties</a></li>
#
# <li><a href="#Initial-value-problem:-Solar-System15">
# Initial value problem: Solar-System15</a></li>
#
# <li><a href="#Ode-problem">
# Ode problem</a></li>
#
# <li><a href="#Error-in-energy">
# Error in energy</a></li>
#
# <li><a href="#Work-Precision-diagrams">
# Work-Precision diagrams</a></li>
#
# </ul>
# ## Load packages
using OrdinaryDiffEq,DiffEqDevTools,BenchmarkTools
using IRKGaussLegendre
using Plots,LinearAlgebra
using Dates
using RecursiveArrayTools
using JLD2, FileIO
setprecision(BigFloat, 108);
# +
#plotly()
# -
# <a href="#top">Back to the top</a>
#
# ## Computer properties
#export JULIA_NUM_THREADS=2
Threads.nthreads()
# +
#;cat /proc/cpuinfo # on Linux machines
# -
using Hwloc
Hwloc.num_physical_cores()
# <a href="#top">Back to the top</a>
#
# ## Initial value problem: Solar-System15
# +
include("../examples/Nbody.jl")
include("../examples/InitialNBody15.jl")
u0128, Gm128 =InitialNBody15(BigFloat)
q0128=u0128[2,:,:]
v0128=u0128[1,:,:]
dt=5
t0=0.0
t1=20*dt #1000.
mysaveat=1 #10
tspan128=(BigFloat(t0),BigFloat(t1))
prob128=ODEProblem(NbodyODE!,u0128,tspan128,Gm128);
# -
u064, Gm64 =InitialNBody15(Float64)
lpp=Gm64;
tspan128
# <a href="#top">Back to the top</a>
#
# ## Ode problem
# ### Test solution
prob128 = ODEProblem(NbodyODE!,u0128,tspan128,Gm128,;lpp=lpp);
#sol =solve(prob128,Vern9(),saveat=mysaveat,abstol=1e-24,reltol=1e-24,maxiters=100000);
#@save "./Data/solarsystem15small_test_solF128.jld2" sol
@load "./Data/solarsystem15small_test_solF128.jld2" sol
test_sol = TestSolution(sol);
# ### IRKGL16 (Fixed-Step)
sol1 =solve(prob128,IRKGL16(),dt=dt,saveat=mysaveat,adaptive=false,mixed_precision=false);
sol1.destats
sol2 = solve(prob128,IRKGL16(),dt=dt,saveat=mysaveat,adaptive=false,
mixed_precision=true, low_prec_type=Float64);
sol2.destats
# <a href="#top">Back to the top</a>
#
# ## Error in energy
# +
setprecision(BigFloat, 256)
u0128, Gm128 =InitialNBody15(BigFloat)
E0=NbodyEnergy(u0128,Gm128)
ΔE = map(x->NbodyEnergy(BigFloat.(x),Gm128), sol.u)./E0.-1
ΔE1 = map(x->NbodyEnergy(BigFloat.(x),Gm128), sol1.u)./E0.-1
ΔE2 = map(x->NbodyEnergy(BigFloat.(x),Gm128), sol2.u)./E0.-1
(Float32(maximum(abs.(ΔE))),Float32(maximum(abs.(ΔE1))),Float32(maximum(abs.(ΔE2))))
# -
ylimit1=-30
ylimit2=-18
plot(sol.t,log10.(abs.(ΔE)), label="Test solution",
ylims=(ylimit1,ylimit2),)
plot!(sol1.t,log10.(abs.(ΔE1)), label="IRKGL16-Float128",
ylims=(ylimit1,ylimit2),)
plot!(sol2.t,log10.(abs.(ΔE2)), label="IRKGL16-Mixed Precision: Float128/Float64",
ylims=(ylimit1,ylimit2),)
# <a href="#top">Back to the top</a>
#
# ## Work-Precision diagrams
Threads.nthreads()
abstols = 1.0 ./ 10.0 .^ (8:18)
reltols = 1.0 ./ 10.0 .^ (8:18)
dts=16*0.866.^(0:length(reltols)-1);
setups = [ Dict(:alg=>IRKGL16(),:adaptive=>false,:dts=>dts,:mixed_precision=>false)
Dict(:alg=>IRKGL16(),:adaptive=>false,:dts=>dts,:mixed_precision=>true,:low_prec_type=>Float64)
]
wp1 = WorkPrecisionSet(prob128,abstols,reltols,setups;appxsol=test_sol,save_everystep=false,numruns=1,maxiters=1000000)
plot(wp1)
# <a href="#top">Back to the top</a>
| Tutorials/.ipynb_checkpoints/Solar-System15-Mixed-Precision-mod2-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (dp_env)
# language: python
# name: dp_env
# ---
# ### You can also run the notebook in [COLAB](https://colab.research.google.com/github/deepmipt/DeepPavlov/blob/master/examples/gobot_tutorial.ipynb).
# !pip install deeppavlov
# # Goal-oriented bot in DeepPavlov
# The tutor is focused on building a goal-oriented dialogue system:
#
# 0. [Data preparation](#0.-Data-Preparation)
# 1. [Build database of items](#1.-Build-database-of-items)
# 2. [Build Slot Filler](#2.-Build-Slot-Filler)
# 3. [Train bot](#3.-Train-bot)
#
# An example of the final model served as a telegram bot is:
#
# 
# ## 0. Data Preparation
# The tutor's dialogue system will be on the domain of restaurant booking. [Dialogue State Tracking Challenge 2 (DSTC-2)](http://camdial.org/~mh521/dstc/) dataset provides dialogues of a human talking to a booking system labelled with slots and dialogue actions. The labels are will be used for training a dialogue policy network.
#
# See below a small chunk of the data.
# +
from deeppavlov.dataset_readers.dstc2_reader import SimpleDSTC2DatasetReader
data = SimpleDSTC2DatasetReader().read('my_data')
# -
# !ls my_data
# The training/validation/test data is stored in json files (`simple-dstc2-trn.json`, `simple-dstc2-val.json` and `simple-dstc2-tst.json`):
# !head -n 101 my_data/simple-dstc2-trn.json
# +
from deeppavlov.dataset_iterators.dialog_iterator import DialogDatasetIterator
iterator = DialogDatasetIterator(data)
# -
# You can now iterate over batches of preprocessed DSTC-2 dialogs:
# +
from pprint import pprint
for dialog in iterator.gen_batches(batch_size=1, data_type='train'):
turns_x, turns_y = dialog
print("User utterances:\n----------------\n")
pprint(turns_x[0], indent=4)
print("\nSystem responses:\n-----------------\n")
pprint(turns_y[0], indent=4)
break
# -
# !cp my_data/simple-dstc2-trn.json my_data/simple-dstc2-trn.full.json
# +
import json
NUM_TRAIN = 50
with open('my_data/simple-dstc2-trn.full.json', 'rt') as fin:
data = json.load(fin)
with open('my_data/simple-dstc2-trn.json', 'wt') as fout:
json.dump(data[:NUM_TRAIN], fout, indent=2)
print(f"Train set is reduced to {NUM_TRAIN} dialogues (out of {len(data)}).")
# -
# ## 1. Build database of items
#
# 
#
# For a valid goal-oriented bot there should be a `database` of relevant items. In the case of restaurant booking it will contain all available restaurants and their info.
#
# >> database([{'pricerange': 'cheap', 'area': 'south'}])
#
# Out[1]:
# [[{'name': 'the lucky star',
# 'food': 'chinese',
# 'pricerange': 'cheap',
# 'area': 'south',
# 'addr': 'cambridge leisure park clifton way cherry hinton',
# 'phone': '01223 244277',
# 'postcode': 'c.b 1, 7 d.y'},
# {'name': 'nandos',
# 'food': 'portuguese',
# 'pricerange': 'cheap',
# 'area': 'south',
# 'addr': 'cambridge leisure park clifton way',
# 'phone': '01223 327908',
# 'postcode': 'c.b 1, 7 d.y'}]]
#
# The dialogues in the training dataset should contain a `"db_result"` dictionary key. It is required for turns where system performs a special type of external action: an api call to the database of items. `"db_result"` should contain the result of the api call:
# !head -n 78 my_data/simple-dstc2-trn.json | tail +51
# +
from deeppavlov.core.data.sqlite_database import Sqlite3Database
database = Sqlite3Database(primary_keys=["name"],
save_path="my_bot/db.sqlite")
# -
# Set `primary_keys` to a list of slot names that have unique values for different items (common SQL term). For the case of DSTC-2, the primary slot is restaurant name.
#
# Let's find all `"db_result"` api call results and add it to our database of restaurants:
# +
db_results = []
for dialog in iterator.gen_batches(batch_size=1, data_type='all'):
turns_x, turns_y = dialog
db_results.extend(x['db_result'] for x in turns_x[0] if x.get('db_result'))
print(f"Adding {len(db_results)} items.")
if db_results:
database.fit(db_results)
# -
# ##### Interacting with database
# We can now play with the database and make requests to it:
database([{'pricerange': 'cheap', 'area': 'south'}])
# !ls my_bot
# ## 2. Build Slot Filler
#
# 
#
# Slot Filler is component that inputs text and outputs dictionary of slot names and their values:
#
# >> slot_filler(['I would like some chineese food'])
#
# Out[1]: [{'food': 'chinese'}]
#
# To implement a slot filler you need to provide
#
# - **slot types**
# - all possible **slot values**
# - optionally, it will be good to provide examples of mentions for every value of each slot
#
# The data should be in `slot_vals.json` file with the following format:
#
# {
# 'food': {
# 'chinese': ['chinese', 'chineese', 'chines'],
# 'french': ['french', 'freench'],
# 'dontcare': ['any food', 'any type of food']
# }
# }
#
#
# Let's use a simple non-trainable slot filler that relies on levenshtein distance:
# +
from deeppavlov.download import download_decompress
download_decompress(url='http://files.deeppavlov.ai/deeppavlov_data/dstc_slot_vals.tar.gz',
download_path='my_bot/slotfill')
# -
# !ls my_bot/slotfill
# !head -n 10 my_bot/slotfill/dstc_slot_vals.json
# ##### Metric scores on valid&test
# Let's check performance of our slot filler on DSTC-2 dataset:
# +
from deeppavlov import configs
from deeppavlov.core.common.file import read_json
slotfill_config = read_json(configs.ner.slotfill_simple_dstc2_raw)
# -
# We take [original DSTC2 slot-filling config](https://github.com/deepmipt/DeepPavlov/blob/master/deeppavlov/configs/ner/slotfill_dstc2_raw.json) and change variables determining data paths:
slotfill_config['metadata']['variables']['DATA_PATH'] = 'my_data'
slotfill_config['metadata']['variables']['SLOT_VALS_PATH'] = 'my_bot/slotfill/dstc_slot_vals.json'
# +
from deeppavlov import evaluate_model
slotfill = evaluate_model(slotfill_config);
# -
# We've got slot accuracy of **93% on valid** set and **94% on test** set.
# ##### Interacting with slot filler
# +
from deeppavlov import build_model
slotfill = build_model(slotfill_config)
# -
slotfill(['i want cheap chinee food'])
# ##### Dumping slot filler's config
# Saving slotfill config file to disk (we will require it's path later):
# +
import json
json.dump(slotfill_config, open('my_bot/slotfill_config.json', 'wt'))
# -
# !ls my_bot
# ## 3. Train bot
# Let's assemble all modules together and train the final module: dialogue policy network.
#
#
# 
#
#
# Policy network decides which action the system should take on each turn of a dialogue: should it say goodbye, request user's location or make api call to a database.
#
# The policy network is a recurrent neural network (recurrent over utterances represented as bags of words) and a dense layer with softmax function on top. The network classifies user utterance into one of predefined system actions.
#
#
# 
#
#
# All actions available for the system should be listed in a `simple-dstc2-templates.txt` file. Each action should be associated with a string of the corresponding system response.
#
# Templates should be in the format `<act>TAB<template>`, where `<act>` is a dialogue action and `<template>` is the corresponding response. Response text might contain slot type names, where every `#slot_type` will be filled with the slot value from a dialogue state.
# !head -n 10 my_data/simple-dstc2-templates.txt
# So, actions are actually classes we classify over. And `simple-dstc2-templates.txt` contains the set of classes.
#
# To train the dialogue policy network for classification task you need action label for each system utterance in training dialogues. The DSTC-2 contains `"act"` dictionary key that contains action associated with current response.
#
# The cell below provides an example of training data for the policy network.
# !head -n 24 my_data/simple-dstc2-trn.json
# Let's **construct the final pipeline** of a dialogue system.
#
# We take [default DSTC2 bot config](https://github.com/deepmipt/DeepPavlov/blob/master/deeppavlov/configs/go_bot/gobot_dstc2.json) ([more configs](https://github.com/deepmipt/DeepPavlov/blob/master/deeppavlov/configs/go_bot) are available) and change sections responsible for
# - templates,
# - database,
# - slot filler,
# - embeddings,
# - data and model load/save paths.
# +
from deeppavlov import configs
from deeppavlov.core.common.file import read_json
gobot_config = read_json(configs.go_bot.gobot_simple_dstc2)
# -
# **Configure** bot to use **templates**:
gobot_config['chainer']['pipe'][-1]['template_type'] = 'DefaultTemplate'
gobot_config['chainer']['pipe'][-1]['template_path'] = 'my_data/simple-dstc2-templates.txt'
# **Configure** bot to use our built **database**:
gobot_config['chainer']['pipe'][-1]['database'] = {
'class_name': 'sqlite_database',
'primary_keys': ["name"],
'save_path': 'my_bot/db.sqlite'
}
# **Configure** bot to use levenshtein distance based **slot filler**:
# +
gobot_config['chainer']['pipe'][-1]['slot_filler']['config_path'] = 'my_bot/slotfill_config.json'
gobot_config['chainer']['pipe'][-1]['tracker']['slot_names'] = ['pricerange', 'this', 'area', 'food']
# -
# You can use a simple **bag-of-words as embedder** (by default):
gobot_config['chainer']['pipe'][-1]['embedder'] = None
# Specify train/valid/test **data path** and **path to save** the final bot model:
# +
gobot_config['metadata']['variables']['DATA_PATH'] = 'my_data'
gobot_config['metadata']['variables']['MODEL_PATH'] = 'my_bot'
# -
# The whole dialogue system pipeline looks like this:
#
# 
# ##### Training policy network
# +
from deeppavlov import train_model
gobot_config['train']['batch_size'] = 8 # set batch size
gobot_config['train']['max_batches'] = 250 # maximum number of training batches
gobot_config['train']['val_every_n_batches'] = 40 # evaluate on full 'valid' split each 30 batches
gobot_config['train']['log_every_n_batches'] = 40 # evaluate on 20 batches of 'train' split every 30 batches
gobot_config['train']['log_on_k_batches'] = 20
train_model(gobot_config);
# -
# Training on 50 dialogues takes from 5 to 20 minutes depending on gpu/cpu. Training on full data takes 10-30 mins.
#
# See [config doc page](http://docs.deeppavlov.ai/en/master/intro/configuration.html) for advanced configuration of the training process.
# ##### Metric scores on valid&test
# Calculating **accuracy** of trained bot: whether predicted system responses match true responses (full string match).
# +
from deeppavlov import evaluate_model
evaluate_model(gobot_config);
# -
# With settings of `max_batches=200`, valid accuracy `= 0.5` and test accuracy is `~ 0.5`.
# ##### Chatting with bot
# +
from deeppavlov import build_model
bot = build_model(gobot_config)
# -
bot(['hi, i want to eat, can you suggest a place to go?'])
bot(['i want cheap food'])
bot(['chinese food'])
bot(['thanks, give me their address'])
bot(['i want their phone number too'])
bot(['bye'])
bot.reset()
bot(['hi, is there any cheap restaurant?'])
| examples/gobot_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.3
# language: python
# name: python-373
# ---
# # Rope
#
# [Rope](https://github.com/python-rope/rope) ist eine Python-Refactoring-Bibliothek.
# ## Installation
#
# Rope kann einfach installiert werden mit
#
# ```console
# $pipenv install rope
# ```
# ## Nutzung
#
# Nun importieren wir zunächst den `Project`-Typ und instanziieren ihn mit dem Pfad zum Projekt:
# +
from rope.base.project import Project
proj = Project('requests')
# -
# Dies erstellt dann einen Projektordner mit dem Namen `.ropeproject` in unserem Projekt.
[f.name for f in proj.get_files()]
# Die `proj`-Variable kann eine Reihe von Befehlen ausführen wie `get_files` und `get_file`. Im folgenden Beispiel nutzen wir dies um der Datei `api.py` die Variable `api` zuzuweisen.
# !cp requests/api.py requests/api_v1.py
api = proj.get_file('api.py')
# +
from rope.refactor.rename import Rename
change = Rename(proj, api).get_changes('api_v1')
proj.do(change)
# -
# !cd requests && git status
# !cd requests && git diff __init__.py
# Mit `proj.do(change)` ist also die Datei `requests/__init__.py` so geändert worden, dass von `new_api` anstatt von `api` importiert wird.
# Rope kann nicht nur zum Umbenennen von Dateien verwendet werden, sondern auch für hunderte anderer Fälle; siehe auch [Rope Refactorings](https://github.com/python-rope/rope/blob/master/docs/overview.rst#refactorings).
| docs/refactoring/qa/rope.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.10 64-bit (''gv2'': conda)'
# name: python3
# ---
# +
# Calcula los límites extremos de México
# y de la Cuenca del Valle de México.
import geopandas as gpd
# +
path = "../data/Cuencas/"
fname = "Regiones_Hidrologicas_Administrativas/rha250kgw.shp"
# Se cargan las regiones hidrológico administrativas.
gdf = gpd.read_file(path + fname)
# Se selecciona la Cuenca del Valle de México.
bounds = gdf[gdf["ORG_CUENCA"] ==
"Aguas del Valle de México"].bounds
# Límites extremos de la Cuenca del Valle de México.
print("lon1: " f"{bounds.iat[0, 0]:.3f}")
print("lon2: " f"{bounds.iat[0, 2]:.3f}")
print("lat1: " f"{bounds.iat[0, 1]:.3f}")
print("lat2: " f"{bounds.iat[0, 3]:.3f}")
# +
fname = "Contorno de México 1-4,000,000/conto4mgw.shp"
# Se carga el contorno de México.
gdf = gpd.read_file(path + fname)
bounds = gdf.bounds
# Límites extremos de México.
print("lon1: " f"{bounds.iloc[:, 0].min():.3f}")
print("lon2: " f"{bounds.iloc[:, 2].max():.3f}")
print("lat1: " f"{bounds.iloc[:, 1].min():.3f}")
print("lat2: " f"{bounds.iloc[:, 3].max():.3f}")
| code/bounds.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import seaborn as sb
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.metrics import classification_report,confusion_matrix
from sklearn.svm import SVC
from sklearn.model_selection import train_test_split
# %matplotlib inline
from sklearn.datasets import load_breast_cancer
cancer = load_breast_cancer()
cancer.keys()
df = pd.DataFrame(cancer['data'], columns = cancer['feature_names'])
df
x = df
y = cancer['target']
model = SVC()
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size = 0.3, random_state = 100)
from sklearn.model_selection import GridSearchCV
model.fit(x_train, y_train)
param_grid = {'C':[0.1,1,10,100,1000], 'gamma': [1, 0.1, 0.001, 0.0001]}
grid = GridSearchCV(SVC(), param_grid, verbose = 3)
grid.fit(x_train, y_train)
grid.best_params_
grid.fit(x_train,y_train)
grid.predictions = grid.predict(x_test)
print(classification_report(y_test, grid.predictions))
| GridSearchSVC2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ```python
# # #!/usr/bin/env python
# # coding: utf-8
#
# # This software component is licensed by ST under BSD 3-Clause license,
# # the "License"; You may not use this file except in compliance with the
# # License. You may obtain a copy of the License at:
# # https://opensource.org/licenses/BSD-3-Clause
#
#
# '''
# Training script of human activity recognition system (HAR), based on two different Convolutional Neural Network (CNN) architectures
# '''
# ```
# # Step by Step HAR Training STM32CubeAI
# This notebook provides a step by step demonstration of a simple <u>H</u>uman <u>A</u>ctivity <u>R</u>ecognition system (HAR), based on a convolutional networks (CNN). This script provides a simple data preperation script through `DataHelper` class and let user to preprocess, split, and segment the dataset to bring it into the form which can be used for training and validation of the HAR CNN. It also has a `CNNHandler` class which builds, trains and validate a CNN for a given set of input and output tensors. The `CNNHandler` can create one of the two provided CNN architectures namely, **IGN** and **GMP**.
#
# All the implementations are done in Python using [Keras](https:keras.io/) with [Tensorflow](https://www.tensorflow.org/) as backend.
#
# For demonstration purposes this script uses two datasets created for HAR using accelerometer sensor.
#
# * WISDM, a public dataset provided by <u>WI</u>reless <u>S</u>ensing <u>D</u>ata <u>M</u>ining group. The details of the dataset are available [here](http://www.cis.fordham.edu/wisdm/dataset.php).
#
# * AST our own propritery dataset.
#
# **Note**: We are not providing any dataset in the function pack. The user can download WISDM dataset from [here](http://www.cis.fordham.edu/wisdm/dataset.php), while AST is a private dataset and is not provided.
#
# Following figure shows the detailed workflow of HAR.
#
#
# <p align="center">
# <img width="760" height="400" src="workflow.png">
# </p>
#
# Let us start the implementation now.
# ## Step1 : Import necessary dependencies
# Following section imports all the required dependencies. This also sets seeds for random number generators in Numpy and Tensorflow environments to make the results reproducibile.
# +
import numpy as np
np.random.seed(611)
import argparse, os, logging, warnings
from os.path import isfile, join
from datetime import datetime
# private libraries
from PrepareDataset import DataHelper
from HARNN import ANNModelHandler
# for using callbacks to save the model during training and comparing the results at every epoch
from keras.callbacks import ModelCheckpoint
# disabling annoying warnings originating from Tensorflow
logging.getLogger('tensorflow').disabled = True
import tensorflow as tf
tf.compat.v1.set_random_seed(611)
# disabling annoying warnings originating from python
warnings.simplefilter("ignore")
# -
# ## Step2: Set environment variables
# Following section sets some user variables which will later be used for:
#
# * preparing the dataset.
# * preparing the neural networks.
# * training the neural networks.
# * validating the neural network.
# +
# data variables
dataset = 'WISDM'
merge = True
segmentLength = 24
stepSize = 24
dataDir = 'datasets/ai_logged_data'
preprocessing = True
# neural network variables
modelName = 'IGN'
# training variables
trainTestSplit = 0.6
trainValidationSplit = 0.7
nEpochs = 20
learningRate = 0.0005
decay = 1e-6
batchSize = 64
verbosity = 1
nrSamplesPostValid = 2
# -
# ## Step3: Result directory
# Each run can have different variables and to compare the results of different choices, such as different segment size for the window for data, different overlap settings etc, we need to save the results. Following section creates a result directory to save results for the current run. The name of the directory has following format. `Mmm_dd_yyyy_hh_mm_ss_dataset_model_seqLen_stepSize_epochs_results`, and example name for directory can be `Oct_24_2019_14_31_20_WISDM_IGN_24_16_20_results`, which shows the process was started at October 24, 2019, at 14:31:20, the dataset used was WISDM, with segment size = 24, segment step = 16, and Nr of epochs = 20.
# if not already exist create a parent directory for results.
if not os.path.exists( './results/'):
os.mkdir( './results/' )
resultDirName = 'results/{}/'.format(datetime.now().strftime( "%Y_%b_%d_%H_%M_%S" ) )
os.mkdir( resultDirName )
infoString = 'runTime : {}\nDatabase : {}\nNetwork : {}\nSeqLength : {}\nStepSize : {}\nEpochs : {}\n'.format( datetime.now().strftime("%Y-%b-%d at %H:%M:%S"), dataset, modelName, segmentLength, stepSize, nEpochs )
with open( resultDirName + 'info.txt', 'w' ) as text_file:
text_file.write( infoString )
# ## Step4: Create a `DataHelper` object
# The script in the following section creates a `DataHelper` object to preprocess, segment and split the dataset as well as to create one-hot-code labeling for the outputs to make the data training and testing ready using the choices set by the user in **Step2**.
myDataHelper = DataHelper( dataset = dataset, loggedDataDir = dataDir, merge = merge,
modelName = modelName, seqLength = segmentLength, seqStep = stepSize,
preprocessing = preprocessing, trainTestSplit = trainTestSplit,
trainValidSplit = trainValidationSplit, resultDir = resultDirName )
# ## Step5: Prepare the dataset
# Following section prepares the dataset and create six tensors namely `TrainX`, `TrainY`, `ValidationX`, `ValidationY`, `TestX`, `TestY`. Each of the variables with trailing `X` are the inputs with shape `[_, segmentLength, 3, 1 ]`and each of the variables with trailing `Y` are corresponding outputs with shape `[ _, NrClasses ]`. `NrClasses` for `WISDM` can be `4` or `6` and for `AST` is `5`.
TrainX, TrainY, ValidationX, ValidationY, TestX, TestY = myDataHelper.prepare_data()
# ## Step6: Create a `ANNModelHandler` object
# The script in the following section creates a `ANNModelHandler` object to create, train and validate the <u>A</u>rtificial <u>N</u>eural <u>N</u>etwork (ANN) using the variables created in **Step2**.
myHarHandler = ANNModelHandler( modelName = modelName, classes = myDataHelper.classes, resultDir = resultDirName,
inputShape = TrainX.shape, outputShape = TrainY.shape, learningRate = learningRate,
decayRate = decay, nEpochs = nEpochs, batchSize = batchSize,
modelFileName = 'har_' + modelName, verbosity = verbosity )
# ## Step6: Create a ANN model
# Following script creates the ANN and prints its summary to show the architecture and nr of parameters for ANN.
harModel = myHarHandler.build_model()
harModel.summary()
# ## Step7: Create a Checkpoint for ANN training
# The following script creates a check point for the training process of ANN to save the neural network as `h5` file. The settings are used in a way that the validation accuracy `val_acc` is maximized.
harModelCheckPoint = ModelCheckpoint( filepath = join(resultDirName, 'har_' + modelName + '.h5'),
monitor = 'val_acc', verbose = 0, save_best_only = True, mode = 'max' )
# ## Step7 : Train the created neural network
# The following script trains the created neural network with the provided checkpoint and created datasets.
harModel = myHarHandler.train_model( harModel, TrainX, TrainY, ValidationX, ValidationY, harModelCheckPoint )
# ## Step8: Validating the trained neural network
# The following section validates the created network and creates a confusion matrix for the test dataset to have a detailed picture of the errors.
myHarHandler.make_confusion_matrix( harModel, TestX, TestY )
# ## Step9: Create an npz file for validation after conversion from CubeAI.
myDataHelper.dump_data_for_post_validation( TestX, TestY, nrSamplesPostValid )
| SensorTile/STM32CubeFunctionPack_SENSING1_V4.0.2/Utilities/AI_Ressources/Training Scripts/HAR/CNN_HAR_KERAS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="HOAv4FLaAC3f"
# # Compute Log Loss for a Classification Model
# + colab={} colab_type="code" id="bN_083VRAC3h"
# import libraries
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="eMkdycnKAC3k" outputId="e4f4f1e3-9b87-4cd5-9539-4692a32407b6"
# data doesn't have headers, so let's create headers
_headers = ['buying', 'maint', 'doors', 'persons', 'lug_boot', 'safety', 'car']
# read in cars dataset
df = pd.read_csv('https://raw.githubusercontent.com/PacktWorkshops/The-Data-Science-Workshop/master/Chapter06/Dataset/car.data', names=_headers, index_col=None)
df.head()
# target column is 'car'
# + colab={"base_uri": "https://localhost:8080/", "height": 224} colab_type="code" id="EQvotJcIAC3o" outputId="ac75fe26-8ead-45a5-c7e2-7820f589f454"
# encode categorical variables
_df = pd.get_dummies(df, columns=['buying', 'maint', 'doors', 'persons', 'lug_boot', 'safety'])
_df.head()
# + colab={} colab_type="code" id="JcMh8CFzAC3q"
# target column is 'car'
features = _df.drop(['car'], axis=1).values
labels = _df[['car']].values
# split 80% for training and 20% into an evaluation set
X_train, X_eval, y_train, y_eval = train_test_split(features, labels, test_size=0.3, random_state=0)
# further split the evaluation set into validation and test sets of 10% each
X_val, X_test, y_val, y_test = train_test_split(X_eval, y_eval, test_size=0.5, random_state=0)
# + colab={"base_uri": "https://localhost:8080/", "height": 136} colab_type="code" id="-4nPRPmXAC3s" outputId="429ffe12-5d71-4048-c55d-5cb56162c398"
# train a Logistic Regression model
model = LogisticRegression()
model.fit(X_train, y_train)
# + colab={} colab_type="code" id="KziGroVtAC3u"
# make predictions for the validation dataset
y_pred = model.predict(X_val)
# + colab={} colab_type="code" id="YfpxsWw0AC3w"
# import libraries
from sklearn.metrics import log_loss
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="phVn4D7cAC3z" outputId="0f9a4e44-5e96-4ce0-8b78-08c5a91040a7"
_loss = log_loss(y_val, model.predict_proba(X_val))
print(_loss)
# + colab={} colab_type="code" id="MTlthhG0AC31"
| Chapter06/Old/Exercise6_10/Exercise6_10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/madhavjk/CP-Python/blob/main/SESSION12_(Collections).ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="AhEMAoWTnGGW"
from collections import Counter
# + id="CO1YDxjNnGGY" outputId="22911db9-d73d-45a8-aedd-922f742d6eb3"
lst = [1,2,2,2,2,3,3,3,1,2,1,12,3,2,32,1,21,1,223,1]
Counter(lst)
# + id="wIsVb2OdnGGa" outputId="16bb1c82-5ba3-4d2c-9df8-75e441f0d096"
Counter('aabsbsbsbhshhbbsbs')
# + id="G1vQJkBTnGGb" outputId="4d14a402-13fc-4f2d-f523-7ea687b23bb9"
s = 'How many times does each word show up in this sentence word times each each word'
words = s.split()
print(words)
Counter(words)
# + id="tdC3uugTnGGc" outputId="7f6ee658-cfe3-44e0-98b0-62a36d004deb"
c = Counter(words)
c.most_common(2)
# + [markdown] id="wUh5F5EwnGGc"
#
# + [markdown] id="6zytWjb9nGGd"
# # defaultdict
# + id="h9WCdmMKnGGd"
from collections import defaultdict
# + id="4nN1N6bCnGGe"
d = {1: "ABC",2: "XYZ"}
# + id="QT3ez1MxnGGf" outputId="e09e206a-8aaf-4c4e-fdf9-bcf831a5333b"
d[1]
# + id="oq_Ii28inGGf" outputId="aff40ecd-7698-48e1-cf21-4ea7037d8c9c"
d[2]
# + id="l-8h3z2knGGg" outputId="03a1ca1f-5141-446a-d2a8-1aa1fe5fff6b"
d[3]
# + id="inyzVpwsnGGg"
d1 = defaultdict(int)
# + id="qNnnGSDNnGGh"
d1['one'] = 1
d1['two'] = 2
# + id="jUKUaIWenGGh" outputId="e9d4cff9-ac1f-41de-e875-15bac2a50589"
d1['three']
# + id="zmGYiHcmnGGh"
d1 = defaultdict(str)
# + id="kJGjRiexnGGh" outputId="c4a83031-dfdc-4fd0-f9e4-75c366f4dbcc"
d1[1] = 'One'
d1[2] = 'Two'
d1[3]
# + [markdown] id="l41ZW7rdnGGi"
# # ordereddict
# + id="jQ3DxkQvnGGi"
from collections import OrderedDict
# + id="5q_ybz4WnGGi"
od = OrderedDict()
# + id="XDoe5wSXnGGi"
od['a'] = 1
od['b'] = 2
od['c'] = 3
# + id="gAfiC3wInGGj" outputId="a4691490-8800-4dea-f4da-45683a5d8658"
od
# + id="jQZ8LM3OnGGj" outputId="fba8f27f-b1bd-4712-b654-2a769e388bce"
od['a'] = 4
od
# + [markdown] id="ezNg2b6CnGGk"
# # deque
# + id="yIm-YRMvnGGk"
from collections import deque
# + id="MUVGrBYXnGGk"
l = ["a","b","c"]
# + id="00utH314nGGk"
deq = deque(l)
# + id="T5ITjvrGnGGl" outputId="223a7893-07ed-49f8-96c2-7112e8bf685b"
deq
# + id="U7jz_WIGnGGl" outputId="8d0eaa9d-35bb-42ad-ebde-41114c3497e0"
deq.append("d")
deq
# + id="rtpVl8wCnGGm" outputId="ca311880-1a2b-4c85-abef-3fc857c40b41"
deq.appendleft("x")
deq
# + id="tZujvh3QnGGm" outputId="b15b2947-35f1-4a38-f496-8698a238f4e6"
deq.pop()
# + id="X5m8Pfg5nGGn" outputId="b6eb1a0c-7c3c-4cbb-e711-bc9bce9d086f"
deq
# + id="voRqngRynGGn" outputId="b612c485-1e59-4fe7-fef7-d642bed10c86"
deq.popleft()
# + id="_4Ix6FQAnGGn" outputId="e729f7d8-a142-43d6-a135-eb474acd0b8b"
deq
# + id="rez1D4mvnGGo" outputId="daa1ad51-338d-4d26-c5ec-679edf5ab7eb"
deq.count("c")
# + id="jTKJlGp7nGGo"
deq.append("a")
# + id="qUNqCMvrnGGo" outputId="62de56ba-e9b2-4a41-be74-08c415cb48e0"
deq.count("a")
# + [markdown] id="1i9KYn8unGGp"
# # ChainMap
# + id="8eJ9zh5tnGGp"
d1 = {'a':1, 'b' : 2}
d2 = {'c': 3, 'd': 4}
# + id="Uh5aTyBZnGGp"
from collections import ChainMap
c_map = ChainMap(d1,d2)
# + id="P1rUMk-bnGGq" outputId="ada9cfb8-57c9-4e2d-849b-ed34c800b33f"
c_map.maps
# + id="xMf7E6cCnGGq" outputId="793f3c40-c475-4943-a539-f7e372b7586d"
c_map['a']
# + id="Cd-Dnuj1nGGr" outputId="9ebbc04f-7b32-49b3-c6ec-601dbf86ff66"
c_map['c']
# + id="81mUsDIanGGr" outputId="b4148287-0408-4383-d069-55e602c6f279"
list(c_map.keys())
# + id="zfpoW2cLnGGr" outputId="bb8ed6ff-ae36-4d7a-d8b1-97bc6655351a"
list(c_map.values())
# + id="wOQiu8I3nGGs"
d3 = {'e' : 5, 'f': 6}
c_map = c_map.new_child(d3)
# + id="SczJ3j87nGGs" outputId="571ab0b2-4f9c-4e1d-ca78-675e93bd561f"
c_map
# + id="D0JBipronGGs" outputId="801c1f4a-a13a-4d90-ea6c-57ca76cf6f14"
c_map['a']
# + id="jXwrvQRPnGGt" outputId="a766951b-3f8c-4957-de89-da4e0c1e5fe7"
c_map['e']
# + [markdown] id="NzMQEwZinGGt"
# # namedtuple
# + id="ts2gRj8CnGGu"
from collections import namedtuple
# + id="hU1EPsq_nGGu"
Student = namedtuple('Student', 'fname, lname, age')
# + id="e0ZgYnx9nGGu"
s1 = Student('Tom', 'Hanks', '60')
# + id="NlPg9AZwnGGu" outputId="f71ba24d-d9b0-4f84-8983-501179975ec0"
s1
# + id="rBgpVV66nGGv" outputId="806edd93-8de4-4091-c79b-ca8a9d3774db"
s1.fname
# + id="SqjIt3v3nGGv" outputId="85db35ec-91f7-4e07-9ff5-7d40a2ba41b9"
s1.age
# + id="s8I2mrjKnGGw"
l = ['John', 'Wick', '50']
s2 = Student._make(l)
# + id="rn9hiEpxnGGx" outputId="8ace2d30-7319-47a7-9ca6-ca429292aaa6"
s2
# + id="lOhHCvnDnGGx"
| SESSION12_(Collections).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# +
# REQUIRED INPUT
# data.norm.sub_phyloseqObject.RData: phyloseq object including
# Perturbations variable in sample_data
# +
###USER-INTERACTION###
# Cells with this header require user interaction, or adaptation
# of the code to the specific case study.
# -
# Load packages and functions
source("robust.clustering.metagenomics.functions.r")
###USER-INTERACTION###
# Define suffix for the current case study files
labelExp <- "CommunityExample" # For example: BifFae
setwd(labelExp)
###USER-INTERACTION###
# Define variables name from sampling table of phyloseq object
# Sample attribute for time points
stepVar <- "time"
# Sample attribute for subject
subjectVar <- "subject"
# Run robust clustering for up to 10 clusters
dir.create('RobustClustering')
file.copy('data.norm.sub_phyloseqObject.RData','RobustClustering/',copy.date=TRUE,overwrite=TRUE)
robust.clustering.all.steps('RobustClustering','data.norm.sub_phyloseqObject.RData',labelExp,stepVar,maxClus=10)
# Print state time serie, per subject
setwd(paste('RobustClustering/',labelExp,'_all/',sep=''))
fout <- paste('data.normAndDist_definitiveClustering_',labelExp,'.RData',sep='')
file.copy(fout,'../..',copy.date=TRUE,overwrite=TRUE)
load(fout)
table(sample_data(data.norm)$cluster)
tableSerie <- stateSerieTable(data.norm, stepVar, subjectId = subjectVar)
timeSerieHeatMap(tableSerie, "./", paste('statesSequence_all',labelExp,".pdf",sep=''))
setwd('../..')
###USER-INTERACTION###
# To compute the cluster, when you decide a fix number of them,
# to adapt better to the case study characteristics.
# Example with k=3
k=3
dir.create(paste('RobustClustering_k',k,sep=''))
file.copy('data.norm.sub_phyloseqObject.RData',paste('RobustClustering_k',k,sep=''),copy.date=TRUE,overwrite=TRUE)
robust.clustering.all.steps(paste('RobustClustering_k',k,sep=''),'data.norm.sub_phyloseqObject.RData',labelExp,'time',minClus=k,maxClus=k)
setwd(paste('RobustClustering_k',k,'/',labelExp,'_all/',sep=''))
fout <- paste('data.normAndDist_definitiveClustering_',labelExp,'.RData',sep='')
file.copy(fout,'../..',copy.date=TRUE,overwrite=TRUE)
load(fout)
table(sample_data(data.norm)$cluster)
tableSerie <- stateSerieTable(data.norm, stepVar, subjectId = subjectVar)
timeSerieHeatMap(tableSerie, "./", paste('statesSequence_all',labelExp,".pdf",sep=''))
setwd('../..')
# Starting MDPbiome
dir.create('MDPbiome')
file.copy('../MDPbiome_template/','.',copy.date=TRUE,recursive=TRUE,overwrite=TRUE)
file.rename('MDPbiome_template','MDPbiome')
file.copy(paste('data.normAndDist_definitiveClustering_',labelExp,'.RData',sep=''),'MDPbiome/Data/',copy.date=TRUE,overwrite=TRUE)
# Load MDPbiome sources
setwd('MDPbiome/Src/')
source("initMDPBiome.R")
dirdata <- "../Data/"
setwd(dirdata)
# Read OTU and mapping table
phyloObject_datafile <- paste('data.normAndDist_definitiveClustering_',labelExp,'.RData', sep = "")
load(phyloObject_datafile)
###USER-INTERACTION###
# Rename clusters for clarity
levels(sample_data(data.norm)$cluster) <- c("dysbiosis","risky","healthy")
# Rename perturbations field for clarity
sample_data(data.norm)$pert <- as.character(sample_data(data.norm)$Perturbations)
Perturbations <- c('pert') # Only 1 perturbation, with different values
# Associate perturbation to the sample it results in, rather than to
# the state where it is applied. To be the same than in microbiome
# sampling, where the mapping value of the perturbations is associated
# to the sample after the perturbation was applied.
for(subject in unique(get_variable(data.norm,subjectVar))){
subject.data <- phyloSubset(data.norm, subjectVar, subject)
vectPert <- NULL
vectPert <- get_variable(subject.data,'pert')
newVecPert <- c('NA',vectPert[1:(length(vectPert)-1)])
sample_data(data.norm)[sample_names(subject.data), "pert"] <- newVecPert
} # end-for move perturbation
# Generic functions that could be useful for compute Utility Function
# Mainly useful in MDPbiomeGEM (simulated data with GEM)
concMetabolite <- function(phyloObj,subjectVar, subject, met){
subject.data <- phyloSubset(phyloObj, subjectVar, subject)
# concentrations <- get_variable(subject.data,met) # vector
concentrations <- sample_data(subject.data)[,c(met)] # sample_data structure
return(concentrations)
}
# Compute mean of utility variable in all samples of a given cluster
clusterUtilityFunction <- function(clusterId){
cluster.data <- phyloSubset(data.norm, "cluster", clusterId)
scores <- get_variable(cluster.data,goalVar)
return(mean(scores))
}
###USER-INTERACTION###
# To define name utility function
# Ex: To maximize concentration of butyrate
goalVar <- "SCFAincrease"
# To compute utility function
# Each sample must have a <goalVar> variable in the sample_data phyloseq
# object (that clusterUtilityFunction() will use). It could be fill in
# following the next example.
# Example:
metName <- 'Butyrate_C4H8O2'
subjects <- unique(get_variable(data.norm,'subject'))
for (isubject in subjects){
vecConc <- concMetabolite(data.norm,subjectVar,isubject,metName)
newVec <- as.numeric(rep(0,nsamples(vecConc)))
for(pos in 2:(nsamples(vecConc))){
newVec[pos] <- get_variable(vecConc[pos],metName) - get_variable(vecConc[pos-1],metName)
} # end-for pos
sample_data(data.norm)[sample_names(vecConc), goalVar] <- newVec
} # end-for isubject
print('>> Utility Function by state:')
# Print computed values about change in utility function;
# here, a metabolite concentration
for (state in levels(sample_data(data.norm)$cluster)){
ss=subset_samples(subset_samples(data.norm,(cluster==state)),pert!='NA')
avg=mean(get_variable(ss,goalVar))
print(paste(goalVar,state,':',avg))
} # end-for state
print('>> Utility Function by perturbation:')
sample_data(data.norm)$pert=as.factor(sample_data(data.norm)$pert)
for (p in levels(sample_data(data.norm)$pert)){
ss=subset_samples(data.norm,(pert==p))
avg=mean(get_variable(ss,goalVar))
print(paste(p,'(',nsamples(ss), 'samples)',':',avg))
} # end-for perturbation
# +
###USER-INTERACTION###
# To assign cluster preferences if expert knowledge available
# cluster_preference <- c(0,0,1)
# -
# Compute vector of states (i.e. cluster) preferences
states <- levels(sample_data(data.norm)$cluster)
goal_preference <- sapply(states, clusterUtilityFunction)
cluster_preference <- goal_preference
save(data.norm,goal_preference,file='data.normAfterConfigMDPbiomePreprocess.RData')
tableSerie <- stateSerieTable(data.norm, stepVar, subjectId = subjectVar)
timeSerieHeatMap(tableSerie, "./", "statesSequence_allSamples.pdf")
createTreeDir(dirdata,Perturbations)
options(max.print = 9999999)
# Build model and compute stability evaluation
mdpBiomeBase(goalDiversity=FALSE, utilityVar=goalVar)
###USER-INTERACTION###
# Compute generality evaluation
# It is mandatory to define rewardType ("preferGood", "avoidBad" or "proportional")
titledata=labelExp
mdpBiomeLoocv(goal_preference=cluster_preference,rewardType="avoidBad",goalVar=goalVar)
# It usually takes a long time to finish (10-30 minutes), mainly depending
# on the number of subjects
# ALTERNATIVE RUN
### Re-run MDPbiome, when the preprocess R data was saved before
setwd('../Src')
source("initMDPBiome.R")
dirdata <- "../Data/"
labelExp <- "community1"
setwd(dirdata)
load('data.normAfterConfigMDPbiomePreprocess.RData')
Perturbations <- c('pert')
stepVar <- "time"
subjectVar <- "subject"
cluster_preference <- goal_preference
createTreeDir(dirdata,Perturbations)
options(max.print = 9999999)
mdpBiomeBase(goalDiversity=FALSE, utilityVar=goalVar)
| Src/DockerMDPbiome/config_MDPbiome_template.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 9.4
# language: sage
# name: sagemath
# ---
# # Graph Theory using SageMath
#
# In the following we are going to show a few useful ways of interacting with graphs using SageMath. The full reference manual can be found on the official webpage.
#
# [SageMath Reference Manual for Graph Theory](https://doc.sagemath.org/html/en/reference/graphs/index.html)
# +
# It is possible to define a graph directly using a python dictonary.
# 'v1' is the key and '['v2', 'v3', 'v4']' are its values. In graph theoretic language the values are the neighbors of 'v1'
g={'v1': ['v2', 'v3', 'v4']}
# Now, we can construct a Sage graph object as follows
G=Graph(g)
# We can plot it as follows.
G.show()
# -
# Show the graph in three dimensions
G.show3d()
# We can label the edges of a graph as well by nesting dictonaries.
# The key of the outer dictonary is still a vertex, while the nested dictonaries correspond to the neighbors mapped
# to the edge label.
g={0: {1 : 'e1', 2 : 'e2', 3 : 'e3'}, 1: {0 : 'e1', 2 : 'e4', 3: 'e5'}}
Graph(g).show(edge_labels = True)
# We can generate the above graph using the python builtin function 'dict' as follows.
g=dict([(0, dict([(1, ['e1']), (2, ['e2']), (3, ['e3'])])), (1, dict([(0, ['e1']), (2, ['e4']), (3, ['e5'])]))])
Graph(g).show(edge_labels = True)
# We can also get a dictonary from a given graph as follows.
g=graphs.HouseXGraph()
print(g.to_dictionary())
# Create a directed graph as follows.
g={0: {1 : 'e1', 2 : 'e2', 3 : 'e3'}, 1: {0 : 'e1', 2 : 'e4', 3: 'e5'}}
DiGraph(g).show(edge_labels = True)
# Show the directed graph in three dimensions. Notice that the edge lables cannot be displayed here.
DiGraph(g).show3d()
# Sage has some builtin graph which can be accessed by typing "graphs." followed by strg+space.
# We can use the question mark to show some more information about a special graph, for example:
graphs.HouseGraph?
# After we have created a graph, we can access a list of its vertices and a list of its edges as follows.
g=graphs.CompleteGraph(3) # generate the complete graph on 3 vertices using the builtin graphs of sage
g.show()
print(g.vertices())
print(g.edges())
g=Graph({0: {1 : 'e1', 2 : 'e2', 3 : 'e3'}, 1: {0 : 'e1', 2 : 'e4', 3: 'e5'}})
g.show()
# We can display the degree of a vertex as follows.
print(g.degree(0)) # degree of vertex 0
# We can access all the neighbors of a vertex as follows.
print(g.neighbors(0))
# We can access all paths between two vertex as follows.
print(g.all_paths(0, 1))
# We can find the shortest path between two vertices as follows.
print(g.shortest_path(0, 1))
# We can work with adjacency and incidence matrices as follows.
print(g.adjacency_matrix())
print(g.incidence_matrix())
# Creating a graph from a given matrix is also possible.
A = matrix([[0, 1, 1], [1, 0, 1], [1, 1, 0]])
Graph(A).show()
# If we want to use the incidence matrix, we need to specify this directly as follows.
A = matrix([[1, 1, 1, 0, 0], [1, 0, 0, 1, 1], [0, 1, 0, 1, 0], [0, 0, 1, 0, 1]])
Graph(A, format="incidence_matrix").show()
# +
# We can create graphs by using the "add_*" functions of sage graph objects.
g=Graph()
g.add_vertex(0)
g.add_vertex(1)
g.add_vertex(2)
g.show()
# -
# Now, add edges
g.add_edge(0, 2)
g.add_edge(0 ,1)
g.show()
# Notice, that we could use a shortcut here and just add the edges directly.
g=Graph()
g.add_edge(0, 1)
g.add_edge(0, 2)
g.show()
# We can add a list of vertices or edges at once.
g=Graph()
g.add_vertices([0 ,1, 2])
g.add_edges([(0, 1), (0, 2)])
g.show()
# We can iterate over all the neighbors of a vertex as follows.
g=Graph({0: {1 : 'e1', 2 : 'e2', 3 : 'e3'}, 1: {0 : 'e1', 2 : 'e4', 3: 'e5'}})
for v in g.neighbor_iterator(0):
print(v)
# +
# We can change the way Sage displays graph plots as follows.
from sage.graphs.graph_plot import GraphPlot
sage.graphs.graph_plot.DEFAULT_PLOT_OPTIONS["figsize"] = [4, 4] # instead of [4,4]
sage.graphs.graph_plot.DEFAULT_SHOW_OPTIONS["figsize"] = sage.graphs.graph_plot.DEFAULT_PLOT_OPTIONS["figsize"]
#sage.graphs.graph_plot.DEFAULT_PLOT_OPTIONS["vertex_size"] = 50 # instead of 200
#sage.graphs.graph_plot.DEFAULT_PLOT_OPTIONS["vertex_labels"] = False # instead of True
sage.graphs.graph_plot.DEFAULT_PLOT_OPTIONS["vertex_color"] = "white" #instead of salmon
# Notice that there are many, many more options.
graphs.CompleteGraph(4).show()
# -
# The chromatic number and the clique number are calculated straight forward.
g=graphs.HouseGraph()
print(g.chromatic_number())
print(g.clique_number())
# We can also easily check if a given graph is perfect.
print(g.is_perfect())
# We can iterate over the maximum clique of a given graph as follows.
g=graphs.HouseGraph()
g.show()
cliques=list(sage.graphs.cliquer.all_max_clique(g))
for c in cliques:
print(c)
# We can delete edges and vertices as follows.
g.delete_edge(2, 4)
g.show()
g.delete_vertex(4)
g.show()
# We can generate graphs using geng (for generation) and nauty (for isomorphism rejection) as follows.
for n in range(1,4):
print("Evaluating graphs of order {}".format(n))
for g in graphs.nauty_geng("{0} -c".format(n)):
g.show()
# There is a convenient way to tell whether two given graphs are isomorphic.
g=Graph({0: [1 , 2]})
h=Graph({1: [0, 2]})
g.show()
h.show()
print(g.is_isomorphic(h))
# +
# We can combine plots of graphs as follows.
from sage.plot.plot import plot
g=graphs.BarbellGraph(4, 4)
plot(g) + plot(sin, (-5, 5)) # returns a plot object which can be combined with other plot objects
# Of course, the above makes little to no sense
# -
# Print the maximal cliques of a graph
g=graphs.HouseXGraph()
g.show()
print(g.cliques_maximal())
# We can compute a coloring of a given graph as follows
g=graphs.BarbellGraph(3, 3)
P=g.coloring()
g.plot(partition=P) # this colors the given graph using the partition induced by P!
# +
# Calculate the automorphism group of a given graph
g=graphs.HouseXGraph()
g.show()
print(g.automorphism_group())
# Notice that the permutations (in disjoint cyclic notation) (2 3)(0 1) are the generators of the automorphism group
# (2 3) and (0 1) present all the permutation of vertices of g such that the edge relations are preserved
G=PermutationGroup([[(2, 3)], [(0,1)]])
G.cayley_graph().show()
G.is_isomorphic(g.automorphism_group())
# -
print(G.cardinality())
# print all possbile ways of permuting the vertices of a graph such that the edge relations are preserved
print(list(G))
# Create a random regular graph on 20 vertices with degree 3 for each vertex
# Since every edge is incident to two vertices, n \times d must be even
g=graphs.RandomRegular(3, 20)
if g:
g.show()
# We can access a graph database of interesting graphs using a query language as follows
Q = GraphQuery(display_cols=['graph6', 'num_vertices', 'chromatic_number', 'clique_number'], num_edges=['<=', 3])
Q.show()
# The first column is an encoding of persisting graphs, called graph6
# We can easily create graphs from graph6 or convert a graph to graph6 as follows
g=Graph('F@??W')
g.show()
print(g.graph6_string())
# +
# We can form the disjoint union of two graphs as follows
G1=graphs.CycleGraph(4)
G2=graphs.HouseGraph()
H=G1.disjoint_union(G2)
H.show(vertex_size=600)
# -
# We can form the cartesian product of two graphs as follows
G1=graphs.CycleGraph(4)
G2=graphs.HouseGraph()
H=G1.cartesian_product(G2)
H.show(vertex_size=600)
# There is a nice function integrated in sage for pretty printing a list of graphs
upToFourEdges = list(graphs(5, lambda G: G.size() <= 4))
pretty_print(*upToFourEdges)
| graphs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/microprediction/microactors-causality/blob/main/Tigramite_Example.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="bBuMweKSIJ3A"
# !pip install tigramite
# + id="Ojf1dZluIerj"
# !pip install microprediction
# + [markdown] id="YLFuWRo6IPuY"
# Continuing to work down this long list https://www.microprediction.com/blog/popular-timeseries-packages and thought this looked interesting. Paper can be found at [link text](https://advances.sciencemag.org/content/5/11/eaau4996)
#
#
# + id="g92YQDrFIwKN"
import numpy as np
import matplotlib
from matplotlib import pyplot as plt
# %matplotlib inline
## use `%matplotlib notebook` for interactive figures
# plt.style.use('ggplot')
import sklearn
plt.rcParams["figure.figsize"] = (14,10)
import tigramite
from tigramite import data_processing as pp
from tigramite import plotting as tp
from tigramite.pcmci import PCMCI
from tigramite.independence_tests import ParCorr, GPDC, CMIknn, CMIsymb
from tigramite.models import LinearMediation, Prediction
# + colab={"base_uri": "https://localhost:8080/"} id="isG0yNDaI4hz" outputId="581d9795-fd4d-4f78-b9e8-3d521f9cf6cb"
from microprediction import MicroReader
mr = MicroReader()
NAMES = mr.get_stream_names()
NAMES[:2]
# + id="z5qfbrQzIWn6"
import pandas as pd
# + [markdown] id="Vtk3yaKRJJlO"
# First an example with three electricity time series
# + colab={"base_uri": "https://localhost:8080/", "height": 142} id="3hmYSpbkJOgk" outputId="6e58b7c0-3374-42bb-f4c5-d3f121599f11"
NUM=3
EXTRANEOUS = ['coin_a.json','coin_b.json']
COMMONALITY = 'electricity-lbmp-nyiso'
var_names = [ name for name in NAMES if COMMONALITY in name and '~' not in name and 'overall' not in name ][:NUM]+EXTRANEOUS
df = pd.DataFrame(columns=var_names)
for var_name in var_names:
df[var_name]=list(reversed(mr.get_lagged_values(var_name)))[:999]
df[:3]
# + colab={"base_uri": "https://localhost:8080/"} id="JiQdfzB9KcZB" outputId="7c6db42b-668d-44d7-ad3d-16c9a5d9e7d6"
short_var_names = [ n.replace(COMMONALITY,'').replace('.json','').replace('-','') for n in var_names]
short_var_names
# + id="44BSDtrVKfC3"
pp_frame = pp.DataFrame(data=df.values, var_names = short_var_names )
parcorr = ParCorr()
pcmci_parcorr = PCMCI(dataframe=pp_frame, cond_ind_test=parcorr,verbosity=0)
all_parents = pcmci_parcorr.run_pc_stable(tau_max=2, pc_alpha=0.2)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="3XW4J1XyMFtP" outputId="add8c01d-7700-4d9c-dd8c-ea1bea13451b"
results = pcmci_parcorr.run_pcmci(tau_max=2, pc_alpha=0.2)
pcmci_parcorr.print_significant_links( p_matrix = results['p_matrix'],
val_matrix = results['val_matrix'], alpha_level = 0.01)
link_matrix = pcmci_parcorr.return_significant_links(pq_matrix=results['p_matrix'],
val_matrix=results['val_matrix'], alpha_level=0.01)['link_matrix']
tp.plot_time_series_graph(
val_matrix=results['val_matrix'],
link_matrix=link_matrix,
var_names=var_names,
link_colorbar_label='MCI',
); plt.show()
# + id="wUP4O84XJ7wl"
| Tigramite_Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ANURAG-BALA/PythonLaboratory/blob/main/Apr_1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="jJPrvzq22dPN" outputId="0de41481-f224-4f8d-c0c7-74b0ca191917"
#declaring a varible
number=7
name='ANURAG'
print(number)
print(name)
# + colab={"base_uri": "https://localhost:8080/"} id="b9uIQFa33PaV" outputId="8e27b9be-17e6-4097-cf84-80bcd3081d92"
#assigning a value to multiple variables
x=y=z='python is a easy language'
print(x)
print(y)
print(z)
# + colab={"base_uri": "https://localhost:8080/"} id="dKft672J3liq" outputId="2353e4ab-7ea9-4d1d-e23b-8d86d9412c19"
#assigning multiple values to multiple variables
x,y,z='python',3,'value'
print(x)
print(y)
print(z)
# + colab={"base_uri": "https://localhost:8080/"} id="8Yw5K6kL4X_E" outputId="304434df-e200-4151-c839-97047e0e0928"
#datatype - number(float)
x=3.55
print(x, 'is type of',type(x))
# + colab={"base_uri": "https://localhost:8080/"} id="rFNqMAy45HXD" outputId="950c9408-ad85-4e50-d09a-7768405ef1f0"
#datatype - number(integer)
y=90
print(y, 'is type of',type(y))
# + colab={"base_uri": "https://localhost:8080/"} id="M2AZckt76Ftf" outputId="97ff3313-aa4a-4442-9071-1bddf18590ca"
#datatype - number(complex)
z=3+5j
print(z, 'is type of',type(z))
# + colab={"base_uri": "https://localhost:8080/"} id="sVWNhf0I6fNb" outputId="59bdb279-a4ab-4076-ff32-dfdbe2cfa079"
#datatype- string(concatenating a string)
string1='this is lab hour and '
string2='it is soon going to complete'
print(string1+string2)
# + colab={"base_uri": "https://localhost:8080/"} id="K-lq7CHs7q49" outputId="8834e9e2-04d3-4f06-bafa-3d21e4f4a741"
#datatype - string(repetiting a string)
string='python lab '
print(string*5)
| Apr_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Thickness budget in temperature space
# %load_ext autoreload
# %autoreload 2
import xarray as xr
import numpy as np
from matplotlib import pyplot as plt
import budgetcalcs as bc
import calc_wmt as wmt
import datetime
#import cftime
# +
rootdir = '/archive/gam/MOM6-examples/ice_ocean_SIS2/Baltic_OM4_025/1yr/'
averaging = '5daily'
filename = '19000101.ocean_'+averaging+'_native_*.nc'
# filename_snap = '19000101.ocean_'+averaging+'_snap_*.nc'
filename_grid = '19000101.ocean_static.nc'
ds = xr.open_mfdataset(rootdir+filename, combine='by_coords')
# ds = xr.open_dataset(rootdir+filename)
# ds_snap = xr.open_mfdataset(rootdir+filename_snap, combine='by_coords')
grid = xr.open_dataset(rootdir+filename_grid)
cp = 3992
rho0 = 1035
# -
# Load initial conditions, snapshop
filename_ic = 'MOM_IC.nc'
ds_ic = xr.open_dataset(rootdir+filename_ic)
terms = ['Temp','Salt','h']
ds_ic
ds_ic = ds_ic[terms].rename({'Temp':'temp','Salt':'salt','h':'thkcello',
'Time':'time','Layer':'zl','lath':'yh','lonh':'xh'}).squeeze()
time_new = xr.cftime_range(start='1900',end='1900',calendar='noleap')
ds_ic = ds_ic.assign_coords({'time':time_new})
ds_snap = xr.concat([ds_ic,ds_snap],dim='time')
# Budget terms
heat_terms = ['opottemptend','T_advection_xy','Th_tendency_vert_remap',
'boundary_forcing_heat_tendency','internal_heat_heat_tendency',
'opottempdiff','opottemppmdiff','frazil_heat_tendency']
salt_terms = ['osalttend','S_advection_xy','Sh_tendency_vert_remap',
'boundary_forcing_salt_tendency','osaltdiff','osaltpmdiff']
h_terms = ['dhdt','dynamics_h_tendency','vert_remap_h_tendency',
'boundary_forcing_h_tendency','internal_heat_h_tendency']
# Calculate material derivative and diffusive terms
# HEAT
signsLHS = [-1,1,1]
signsRHS = [1,1,1,1,1]
termsLHS = heat_terms[:3]
termsRHS = heat_terms[3:]
Dheat, dheat, error_heat = bc.calc_materialderivative(ds,termsLHS,signsLHS,termsRHS,signsRHS,plot=False)
def calc_refine(da,refineby,variable_type,vertc='zl'):
nk = len(da[vertc])
# Set vertical coordinate to layer index
da=da.assign_coords({vertc:np.linspace(1,nk,nk)})
# Assign a k-value for the interfaces
k_i = np.linspace(0.5,nk+0.5,nk+1)
# Develop the super grid, based on the interfaces
k_i_target = np.linspace(0.5,nk+0.5,nk*refineby+1)
# Get the value of the layers on the super grid
k_l_target = 0.5*(k_i_target[1:]+k_i_target[:-1])
# Refine the grid
if variable_type == 'intensive':
da_refined = da.interp({vertc:k_l_target},method='linear',kwargs={'fill_value':'extrapolate'})
if variable_type == 'extensive':
da_refined = xr.zeros_like(da.interp({vertc:k_l_target}))
for k in range(nk):
index = np.arange(k*refineby,(k+1)*refineby)
vertc_ones = xr.DataArray(np.ones(shape=(refineby)),dims=[vertc],coords={vertc:k_l_target[index]})
chunk = (da.isel({vertc:k})/refineby)*vertc_ones
# Input array must have same dimensional order as indexed array
### THERE MUST BE A MORE EFFICIENT WAY TO DO THIS ###
if len(da.dims)==1:
da_refined.loc[{vertc:k_l_target[index]}]=chunk
elif len(da.dims)==2:
da_refined.loc[{vertc:k_l_target[index]}]=chunk.transpose(list(da.dims)[0],list(da.dims)[1])
elif len(da.dims)==3:
da_refined.loc[{vertc:k_l_target[index]}]=chunk.transpose(list(da.dims)[0],list(da.dims)[1],list(da.dims)[2])
elif len(da.dims)==4:
da_refined.loc[{vertc:k_l_target[index]}]=chunk.transpose(list(da.dims)[0],list(da.dims)[1],list(da.dims)[2],list(da.dims)[3])
return da_refined
# +
# Time-mean : for evaluating dia-boundary transport and integrated process tendencies
l = ds['temp'] # Time-mean volume-defining tracer
l_name = l.name+'_bin' # Naming of binning variable as will be defined by xhistogram
dl = dheat/cp # Sum of diffusive tendencies for volume-defining tracer
c = xr.ones_like(ds['thkcello']) # Time-mean of budget tracer
h = ds['thkcello']
# Snapshots: for evaluating budget tracer content tendency
# NOTE: time-mean i corresponds to the snapshots at i and i-1
# so, for example, diff(snap[1]-snap[0])/dt = mean[1]
l_snap = ds_snap['temp'] # Snapshots of volume-defining tracer
c_snap = xr.ones_like(ds_snap['thkcello']) # Snapshots of budget tracer
h_snap = ds_snap['thkcello'] # Snapshots of layer thickness (for tracer content calculation)
# Grid dimensions
area = grid.areacello # Grid dimensions
# Time-mean tendencies of budget tracer due to different processes
f_c = ds['boundary_forcing_h_tendency'] # Boundary forcing
refine = False
refineby = 0
# -
# Refine vertical grid
if refine:
l = calc_refine(l, refineby=refineby, variable_type='intensive')
dl = calc_refine(dl, refineby=refineby, variable_type='extensive')
c = calc_refine(c, refineby=refineby, variable_type='intensive')
l_snap = calc_refine(l_snap, refineby=refineby, variable_type='intensive')
c_snap = calc_refine(c_snap, refineby=refineby, variable_type='intensive')
h_snap = calc_refine(h_snap, refineby=refineby, variable_type='extensive')
f_c = calc_refine(f_c, refineby=refineby, variable_type='extensive')
# +
# Binning variables
delta_l = 0.5
delta_l_E = 0.25
l_i_vals = np.arange(-4,36,delta_l)
l_i_vals_E = np.arange(-4,36,delta_l_E)
# Calculation of budget tracer content tendency, derived from snapshots
C_mean = wmt.calc_P(rho0*c*h,l,l_i_vals,area) # Binning
C = wmt.calc_P(rho0*c_snap*h_snap,l_snap,l_i_vals,area) # Binning at snapshots
dCdt = C.diff('time')/(C.time.diff('time').astype('float')*1E-9)
# dCdt = dCdt.isel(time=t)
# dCdt_alt = C.differentiate('time').isel(time=t) # Finite difference in time
dCdt = dCdt.rename({l_snap.name+'_bin':l_name}) # Rename dimension for consistency
# dCdt_alt = dCdt_alt.rename({l_snap.name+'_bin':l_name})
# Calculation of E : budget tracer tendency due to dia-boundary mass transport
E_c = wmt.calc_E(c,l,dl,l_i_vals_E,area,plot=False)#.isel(time=t)
E_c = E_c.rename({l_name:l_name+'_E'})
# Calculation of P^n : volume integrated budget tracer tendencies
F_c = wmt.calc_P(rho0*f_c,l,l_i_vals,area)#.isel(time=t)
dCdt = dCdt.assign_coords({'time':E_c.time})
# Residual
residual = dCdt-E_c-F_c
# -
t=1
fig,ax1 = plt.subplots(figsize=(12,6),ncols=1)
ax1.plot(dCdt[l_name],dCdt.isel(time=t))
ax1.plot(E_c[l_name+'_E'],E_c.isel(time=t))
fig,ax1 = plt.subplots(figsize=(12,6),ncols=1)
lw = 0.1
tt = 1
for t in range(tt,31):
# ax1.plot(dCdt[l_name],dCdt.isel(time=t),label='dCdt',color='tab:blue',linewidth=lw)
# ax1.plot(E_c[l_name],E_c.isel(time=t),label='E_c',color='tab:red',linewidth=lw)
# ax1.plot(F_c[l_name],F_c.isel(time=t),label='F_c',color='tab:orange',linewidth=lw)
ax1.plot(residual[l_name],residual.isel(time=t),linestyle='-',label='residual',color='k',linewidth=lw)
# ax1.plot(dCdt[l_name],dCdt.mean('time'),label='dCdt',color='tab:blue')
# ax1.plot(E_c[l_name],E_c.mean('time'),label='E_c',color='tab:red')
# ax1.plot(F_c[l_name],F_c.mean('time'),label='F_c',color='tab:orange')
ax1.plot(residual[l_name],residual.mean('time'),linestyle='-',label='residual',color='k')
# ax1.legend()
print(np.sqrt(np.mean(residual.isel(time=t)**2)))
# Total volume balance through time
ds_total=(ds.astype('float64')*grid['areacello']).sum(['xh','yh','zl'])
ds_total_timemean=ds_total.mean('time')
ds_snap_total=(ds_snap.astype('float64')*grid['areacello']).sum(['xh','yh','zl'])
ds_snap_total_timemean=ds_snap_total.mean('time')
dhdt_from_snap = ds_snap_total['thkcello'].diff('time')/86400
days=range(len(ds_total['time']))
days_snap=np.arange(len(ds_snap_total['time']))-0.5
fig,ax=plt.subplots(figsize=(12,6))
ax.plot(days,ds_total['dhdt'], label='dhdt')
ax.plot(days,ds_total['dynamics_h_tendency'])
ax.plot(days,ds_total['boundary_forcing_h_tendency'])
ax.plot(days,(ds_total['thkcello']-ds_total_timemean['thkcello'])*1E-6)
ax.plot(days_snap[1:],(ds_snap_total['thkcello']-ds_total_timemean['thkcello'])[1:]*1E-6)
ax.plot(days[1:],dhdt_from_snap[1:],'k--')
# +
# Now split the volume along some temperature contour and do the same thing
c = 4
ds_total=(ds.astype('float64')*grid['areacello']).where(ds['temp']>c).sum(['xh','yh','zl'])
ds_total_timemean=ds_total.mean('time')
ds_snap_total=(ds_snap.astype('float64')*grid['areacello']).where(ds_snap['temp']>c).sum(['xh','yh','zl'])
ds_snap_total_timemean=ds_snap_total.mean('time')
dhdt_from_snap = ds_snap_total['thkcello'].diff('time')/86400
days=range(len(ds_total['time']))
days_snap=np.arange(len(ds_snap_total['time']))-0.5
fig,ax=plt.subplots(figsize=(12,6))
ax.plot(days,ds_total['dhdt'],label='dhdt')
ax.plot(days,ds_total['dynamics_h_tendency'],label='dynamics')
ax.plot(days,ds_total['boundary_forcing_h_tendency'],label='boundary')
ax.plot(days,(ds_total['thkcello']-ds_total_timemean['thkcello'])*1E-6,label='volume anom *1E-6')
ax.plot(days_snap[1:],(ds_snap_total['thkcello']-ds_total_timemean['thkcello'])[1:]*1E-6,label='volume anom (snaps) *1E-6')
ax.plot(days[1:],dhdt_from_snap[1:],'k--',label='dhdt (from snaps)')
# Calculate E_c around at this contour
dc = 0.25
layer = (ds['temp']>c-dc/2) & (ds['temp']<c+dc/2)
E_c = (dheat*grid['areacello']/cp).where(layer).sum(['xh','yh','zl'])/dc
ax.plot(days,E_c/rho0,label='E_c',color='tab:brown')
ax.legend()
ax.plot(days[-1],dhdt_from_snap[1:].mean('time').values,'k.')
ax.plot(days[-1],(E_c/rho0)[1:].mean('time').values,'.',color='tab:brown')
# +
# How well does the time mean of the volume derived from snapshots match that of the volume from the timemean
c = 4
ds_total=(ds.astype('float64')*grid['areacello']).where(ds['temp']>c).sum(['xh','yh','zl'])
ds_snap_total=(ds_snap.astype('float64')*grid['areacello']).where(ds_snap['temp']>c).sum(['xh','yh','zl'])
days=np.arange(len(ds_total['time']))
days_snap=np.arange(len(ds_snap_total['time']))-0.5
fig,(ax,ax1)=plt.subplots(figsize=(12,6), nrows=2)
ax.plot(days,ds_total['thkcello'],label='volume from time-mean theta')
# ax.plot(days_snap,(ds_snap_total['thkcello']),label='volume (from snaps)')
h_snap_mean = 0.5*(ds_snap_total['thkcello'].values[1:]+ds_snap_total['thkcello'].values[:-1])
days_snap_mean = 0.5*(days_snap[1:]+days_snap[:-1])
ax.plot(days_snap_mean,h_snap_mean,label='time-mean volume from snapshot theta')
ax.set_ylabel('volume')
ax.legend()
# fig,ax=plt.subplots(figsize=(12,6))
ax1.plot(days,h_snap_mean-ds_total['thkcello'], label='difference')
ax1.set_xlabel('days')
ax1.set_ylabel('volume')
ax1.legend()
# -
# #### Timestep
# +
t=23
# Binning variables
delta_l = 0.5
l_i_vals = np.arange(-4,36,delta_l)
# Calculation of budget tracer content tendency
dCdt = wmt.calc_P(rho0*c*ds['dhdt'],l,l_i_vals,area).isel(time=t) # Binning at snapshots
# Calculation of E : budget tracer tendency due to dia-boundary mass transport
E_c = wmt.calc_E(c,l,dl,l_i_vals,area,plot=False).isel(time=t)
# Calculation of P^n : volume integrated budget tracer tendencies
F_c = wmt.calc_P(rho0*f_c,l,l_i_vals,area).isel(time=t)
# Residual
residual = dCdt-E_c-F_c
# -
fig,ax1 = plt.subplots(figsize=(12,6),ncols=1)
ax1.plot(dCdt['temp_bin'],dCdt)
ax1.plot(F_c['temp_bin'],F_c)
ax1.plot(E_c['temp_bin'],E_c)
ax1.plot(residual['temp_bin'],residual)
# ### Differences of two means
# +
# Time-mean : for evaluating dia-boundary transport and integrated process tendencies
l = ds['temp'] # Time-mean volume-defining tracer
l_name = l.name+'_bin' # Naming of binning variable as will be defined by xhistogram
dl = dheat/cp # Sum of diffusive tendencies for volume-defining tracer
c = xr.ones_like(ds['thkcello']) # Time-mean of budget tracer
h = ds['thkcello']
# Grid dimensions
area = grid.areacello # Grid dimensions
# dt
dt = ds['average_DT']
# Time-mean tendencies of budget tracer due to different processes
f_c = ds['boundary_forcing_h_tendency'] # Boundary forcing
refine = False
# +
# Volume
c = 2
V=(rho0*h*grid['areacello']).where(l>c).sum(['xh','yh','zl'])
dV = V.diff('time')
days=np.arange(len(V['time']))
days_mean = 0.5*(days[1:]+days[:-1])
# Calculate E_c around this contour
dc = 0.1
layer = (l>c-dc/2) & (l<c+dc/2)
G = ((dheat*grid['areacello']/cp).where(layer).sum(['xh','yh','zl'])/dc).load();
Gdt = G*(dt*1E-9)
# Gdt_mean = Gdt.rolling({'time':2},center=True).mean()
Gdt_mean = 0.5*(Gdt.values[1:]+Gdt.values[:-1]);
# # Calculate forcing terms
# F=((f_c*grid['areacello']).where(ds['temp']>c).sum(['xh','yh','zl'])).load();
# F_mean = 0.5*(F.values[1:]+F.values[:-1]);
fig,ax=plt.subplots(figsize=(12,3))
ax.plot(days_mean,dV,label='dV')
ax.plot(days_mean,Gdt_mean,label='Gdt_mean')
# ax.plot(days_mean,F_mean,label='F')
ax.legend()
# +
# Volume
c = 4
V=(h*grid['areacello']).where(l>c).sum(['xh','yh','zl'])
dVdt = V.diff('time')/86400
days=np.arange(len(V['time']))
days_mean = 0.5*(days[1:]+days[:-1])
# Calculate E_c around this contour
dc = 0.25
# Calculate it relative to the time mean contour
l_mean = l.rolling({'time':2},center=True).mean()
dheat_mean = 0.5*(dheat[1:,:,:,:].values+dheat[:-1,:,:,:].values)
layer = (l_mean>c-dc/2) & (l_mean<c+dc/2)
G = ((dheat*grid['areacello']/cp/rho0).where(layer).sum(['xh','yh','zl'])/dc).load();
G_mean = 0.5*(G.values[1:]+G.values[:-1]);
G_mean = G.rolling({'time':2},center=True).mean()
# Calculate forcing terms
F=((f_c*grid['areacello']).where(ds['temp']>c).sum(['xh','yh','zl'])).load();
F_mean = 0.5*(F.values[1:]+F.values[:-1]);
fig,ax=plt.subplots(figsize=(12,3))
ax.plot(days_mean,dVdt,label='dVdt')
ax.plot(days_mean,G_mean,label='G')
ax.plot(days_mean,F_mean,label='F')
ax.legend()
# +
# Coarsen to 5-day means
dt = 5
h_dt = h.coarsen(time=dt,boundary='trim').mean()
l_dt = l.coarsen(time=dt,boundary='trim').mean()
dheat_dt = dheat.coarsen(time=dt,boundary='trim').mean()
f_dt = f_c.coarsen(time=dt,boundary='trim').mean()
# Volume
c = 4
V=(rho0*h_dt*grid['areacello']).where(l_dt>c).sum(['xh','yh','zl'])
dVdt = V.diff('time')/(dt*86400)
days=np.arange(len(V['time']))
days_mean = 0.5*(days[1:]+days[:-1])
# Calculate E_c around this contour
dc = 0.25
layer = (l_dt>c-dc/2) & (l_dt<c+dc/2)
G = ((dheat_dt*grid['areacello']/cp).where(layer).sum(['xh','yh','zl'])/dc).load()
G_mean = 0.5*(G.values[1:]+G.values[:-1])
# Calculate forcing terms
F=((f_dt*grid['areacello']).where(l_dt>c).sum(['xh','yh','zl'])).load()
F_mean = 0.5*(F.values[1:]+F.values[:-1])
fig,ax=plt.subplots(figsize=(12,6))
ax.plot(days_mean,dVdt,label='dVdt')
ax.plot(days_mean,G_mean,label='G')
ax.plot(days_mean,F_mean,label='F')
ax.legend()
# -
| notebooks/calc_wmt_h.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # UBO COSC 301/DATA 301
# ## Milestone 2
# ## Task 3
# ## <NAME>
import seaborn as sns
import matplotlib.pyplot as plt
from matplotlib.pyplot import figure
import pandas as pd
import numpy as np
import sys
sys.path.append('..')
from project_scripts import project_fuctions as pf
# +
#DataFrame Loading for EDA
df_master = pf.load_and_process()
# +
#Interest rate vs Housing Prices
# -
pf.barPlotOverTime(df_master,'Prime Rate',30,25,12,'blue')
pf.barPlotOverTime(df_master,'BC New Housing Price Index',30,15,15,'green')
# +
pf.barPlotOverTime(df_master,'All-items',30,15,15,'red')
#This is a barplot of CPI
# -
df_master.plot(x="")
| analysis/AbirinderBrar/milestone2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Dashboard
#
# The dashboard is used to combine multiple `rubicon_ml` widgets into a sigle, interactive
# dashboard. When a `rubicon_ml.viz.ExperimentsTable` is present in the dashboard, it acts
# as a controller to select the experiments visible in the other widgets in the dashboard.
# +
import random
import numpy as np
import pandas as pd
from rubicon_ml import Rubicon
from rubicon_ml.viz import (
DataframePlot,
ExperimentsTable,
MetricCorrelationPlot,
MetricListsComparison,
)
from rubicon_ml.viz.dashboard import Dashboard
# -
# First, we'll create a few experiments and populate them full of parameters, metrics,
# and dataframes.
# +
dates = pd.date_range(start="1/1/2010", end="12/1/2020", freq="MS")
rubicon = Rubicon(persistence="memory", auto_git_enabled=True)
project = rubicon.get_or_create_project("dashboard composition")
for i in range(0, 10):
experiment = project.log_experiment()
experiment.log_parameter(
name="is_standardized",
value=random.choice([True, False]),
)
experiment.log_parameter(name="n_estimators", value=random.randrange(2, 10, 2))
experiment.log_parameter(
name="sample",
value=random.choice(["A", "B", "C", "D", "E"]),
)
experiment.log_metric(name="accuracy", value=random.random())
experiment.log_metric(name="confidence", value=random.random())
experiment.log_metric(
name="coefficients",
value=[random.random() for _ in range(0, 5)],
)
experiment.log_metric(
name="stderr",
value=[random.random() for _ in range(0, 5)],
)
data = np.array(
[
list(dates),
np.linspace(random.randint(0, 15000), random.randint(0, 15000), len(dates))
]
)
data_df = pd.DataFrame.from_records(
data.T,
columns=["calendar month", "open accounts"],
)
experiment.log_dataframe(data_df, name="open accounts")
# -
# Now, we can instantiate the `Dashboard` object with the experiments we just logged.
# By default, the dashboard will have an `ExperimentsTable` stacked on top of a
# `MetricCorrelationPlot`. Selecting an experiment in the experiments table will render
# it on the metric correlation plot.
#
# We can view the dashboard right in the notebook with `show`. The Dash application
# itself will be running on http://127.0.0.1:8050/ when running locally. Use the
# `serve` command to launch the server directly without rendering the widget in the
# current Python interpreter.
default_dashbaord = Dashboard(experiments=project.experiments())
default_dashbaord.show()
# 
# To customize a dashboard, we can pass in the widgets we want rendered. Arguments to `widgets`
# should be a list of lists of instantiated `rubicon_ml` widgets. Each inner list represents a row in the rendered dashboard. Note that the internal widgets are not instantiated with
# experiments. We provide the experiments to the dashboard itself so each widget shares the
# same experiments.
#
# Again, we can use `show` to see our four `rubicon_ml` widgets arranged into a 2x2 dashboard.
Dashboard(
experiments=project.experiments(),
widgets=[
[
ExperimentsTable(is_selectable=True),
MetricCorrelationPlot(selected_metric="accuracy"),
],
[
MetricListsComparison(column_names=[f"var_00{i}" for i in range(0, 5)]),
DataframePlot(dataframe_name="open accounts"),
],
],
).show()
# 
| notebooks/viz/dashboard.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# %config InlineBackend.figure_format = 'retina'
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
# + [markdown] hideCode=false hidePrompt=false slideshow={"slide_type": "slide"}
# # Product of 4 consecutive numbers is always 1 less than a perfect square
#
#
# <p>
# <center><NAME> (<a href="https://shubhanshu.com">shubhanshu.com</a>)</center>
#
#  
#
# </p>
# + [markdown] hideCode=false hidePrompt=false slideshow={"slide_type": "slide"}
# For every $n \in \mathbb{Z}$, we can have 4 consecutive numbers as follows:
#
# $
# n, n+1, n+2, n+3
# $
# + [markdown] hideCode=false hidePrompt=false slideshow={"slide_type": "fragment"}
# We can complete the proof, if we can show that there exists a $k \in \mathbb{Z}$, such that the following equation holds:
#
# $
# \begin{equation}
# n*(n+1)*(n+2)*(n+3) = (k^2 - 1)
# \end{equation}
# $
#
# + hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
i_max = 4
nums = np.arange(0, 50)+1
consecutive_nums = np.stack([
np.roll(nums, -i)
for i in range(i_max)
], axis=1)[:-i_max+1]
n_prods = consecutive_nums.prod(axis=1)
df = pd.DataFrame(consecutive_nums, columns=[f"n{i+1}" for i in range(i_max)])
df["prod"] = n_prods
df["k"] = np.sqrt(n_prods+1).astype(int)
df["k^2"] = df["k"]**2
df["k^2 - 1"] = df["k^2"] - 1
df
# + hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
fig, ax = plt.subplots(1,3, figsize=(18, 6))
ax[0].plot("n1", "prod", "bo-", data=df)
ax[0].set_xlabel("n", fontsize=20)
ax[0].set_ylabel(f"$y = \prod_{{i=0}}^{{i={i_max}}} (n+i)$", fontsize=20)
ax[1].plot(df["k"], df["prod"], "ko-")
ax[1].set_xlabel("$k = \sqrt{y + 1}$", fontsize=20)
ax[1].set_title("$y = k^2 - 1$", fontsize=20)
ax[2].plot(df["n1"], df["k"], "ko-")
ax[2].set_ylabel("$k = \sqrt{y + 1}$", fontsize=20)
ax[2].set_xlabel("$n$", fontsize=20)
fig.tight_layout()
# + [markdown] hideCode=false hidePrompt=false slideshow={"slide_type": "slide"}
# Let us look at the right hand side of the equation first, i.e. $k^2 - 1$.
# + [markdown] hideCode=false hidePrompt=false slideshow={"slide_type": "fragment"}
# This can be rewritten as $\textbf{(k-1)*(k+1)}$
# + [markdown] hideCode=false hidePrompt=false slideshow={"slide_type": "fragment"}
# **Now, this is where a hint lies.**
# + [markdown] hideCode=false hidePrompt=false slideshow={"slide_type": "slide"}
# What the right hand side means that it is a product of two integers ($k-1$ and $k+1$) which differ by 2.
# + [markdown] hideCode=false hidePrompt=false slideshow={"slide_type": "fragment"}
# We can see that this is the case:
#
# $
# \begin{equation}
# (k+1) - (k-1) \\
# = k + 1 - k - (-1) \\
# = k - k + 1 - (-1) \\
# = 0 + 1 + 1 \\
# = 2 \\
# \end{equation}
# $
# + [markdown] hideCode=false hidePrompt=false slideshow={"slide_type": "slide"}
# So, if we can somehow show that the left hand side of the original equation, i.e. $n*(n+1)*(n+2)*(n+3)$:
#
# * can be represented as a product of two numbers which differ by 2, then we are done,
# * as these numbers can then be mapped to $k-1$ and $k+1$ for some $k \in \mathbb{Z}$.
# + [markdown] hideCode=false hidePrompt=false slideshow={"slide_type": "slide"}
# We can group the numbers $\textbf{n, n+1, n+2, n+3}$ into pairs, with the hope of getting $k-1$ and $k+1$.
# + [markdown] slideshow={"slide_type": "fragment"}
# We can utilize following facts to choose the two pairs:
#
# * The difference of the products should be constant, and hence independent of $n$
# * Knowing that product of two factors of type $(n+i)*(n+j) = n^2 + (i+j)*n + i*j$,
# * We can observe that $i+j$ will be same for numbers which are equidistant from the middle of all numbers.
#
# Now we can select our pair of numbers.
# + [markdown] hideCode=false hidePrompt=false slideshow={"slide_type": "slide"}
# * The first pair is $n$ and $(n+3)$,
# * and their product is $\textbf{n * (n+3)}$
# * which can be expanded as $\color{red}{\textbf{n^2 + 3n}}$
# + [markdown] hideCode=false hidePrompt=false slideshow={"slide_type": "fragment"}
# * And, the second pair $(n+1)$ and $(n+2)$,
# * and their product is $\textbf{(n+1)*(n+2)}$
# * which can be expanded as $\color{red}{\textbf{n^2 + 3n}} + \textbf{2}$
# + [markdown] hideCode=false hidePrompt=false slideshow={"slide_type": "slide"}
# Based on the above pairing we can immediately see that the difference of these pair products is as follows:
#
# $
# \begin{equation}
# [(n+1)*(n+2)] - [n * (n+3)]\\
# = [\color{red}{n^2 + 3n} + 2] - [\color{red}{n^2 + 3n}]\\
# = n^2 + 3n + 2 - n^2 - 3n\\
# = (n^2 -n^2) + (3n - 3n) + 2\\
# = 0 + 0 + 2\\
# = 2
# \end{equation}
# $
# + [markdown] hideCode=false hidePrompt=false slideshow={"slide_type": "slide"}
# Hence, based on the above simplification, we can map:
#
# * $(\color{red}{n^2 + 3n} + 2) \rightarrow (k+1)$, and
# * $(\color{red}{n^2 + 3n}) \rightarrow (k-1)$.
# + [markdown] hideCode=false hidePrompt=false slideshow={"slide_type": "fragment"}
# Now, if we choose $\color{blue}{\textbf{k = (n^2 + 3n + 1)}}$, the following equations hold:
# + [markdown] hideCode=false hidePrompt=false slideshow={"slide_type": "fragment"}
# * $n^2 + 3n + 2 = \color{blue}{(n^2 + 3n + 1)} + 1 = \color{blue}{k} + 1$
# + [markdown] hideCode=false hidePrompt=false slideshow={"slide_type": "fragment"}
# * $n^2 + 3n = \color{blue}{(n^2 + 3n + 1)} - 1 = \color{blue}{k} - 1$
# + [markdown] hideCode=false hidePrompt=false slideshow={"slide_type": "slide"}
# Hence, we have proved the following:
#
# $
# \begin{equation}
# \forall n \in \mathbb{Z}, \\
# \exists k \in \mathbb{Z} \\
# n*(n+1)*(n+2)*(n+3) \\
# = [(n+3)*n]*[(n+1)*(n+2)]\\
# = [\color{red}{n^2 + 3n}]*[\color{red}{n^2 + 3n} + 2]\\
# = [\color{blue}{(n^2 + 3n + 1)} - 1]*[\color{blue}{(n^2 + 3n + 1)} + 1]\\
# = [\color{blue}{k} - 1]*[\color{blue}{k} + 1]\\
# = (k^2 - 1)
# \end{equation}
# $
# + [markdown] hideCode=false hidePrompt=false slideshow={"slide_type": "fragment"}
# And this equation can be solved by choosing $\color{blue}{\textbf{k = (n^2 + 3n + 1)}}$.
# + [markdown] hideCode=false hidePrompt=false slideshow={"slide_type": "fragment"}
# Hence, proved.
# + hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
df["k = n^2 + 3n + 1"] = (df["n1"]**2 + 3*df["n1"] + 1)
df
# + hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
fig, ax = plt.subplots(1,3, figsize=(12, 6))
ax[0].plot("n1", "prod", "bo-", data=df)
ax[0].set_xlabel("n", fontsize=20)
ax[0].set_ylabel(f"$y = \prod_{{i=0}}^{{i={i_max}}} (n+i)$", fontsize=20)
ax[1].plot(df["k"], df["prod"], "ko-")
ax[1].set_xlabel("$k = \sqrt{y + 1}$", fontsize=20)
ax[1].set_title("$y = k^2 - 1$", fontsize=20)
ax[2].plot(df["n1"], df["k"], "ko-", label="$k = \sqrt{y + 1}$")
ax[2].plot(df["n1"], df["k = n^2 + 3n + 1"], "r--", label="$k = n^2 + 3n + 1$")
ax[2].legend(fontsize=14)
ax[2].set_ylabel("$k = \sqrt{y + 1}$", fontsize=20)
ax[2].set_xlabel("$n$", fontsize=20)
fig.tight_layout()
# + [markdown] hideCode=false hidePrompt=false slideshow={"slide_type": "slide"}
# # More videos to come
#
#
# <p>
# <center><NAME> (<a href="https://shubhanshu.com">shubhanshu.com</a>)</center>
#
#  
#
# </p>
# + hideCode=false hidePrompt=true slideshow={"slide_type": "slide"}
fig, ax = plt.subplots(1,3, figsize=(12, 6))
fig.patch.set_facecolor('white')
ax[0].plot("n1", "prod", "bo-", data=df)
ax[0].set_xlabel("n", fontsize=20)
ax[0].set_ylabel(f"$y = \prod_{{i=0}}^{{i={i_max}}} (n+i)$", fontsize=20)
ax[1].plot(df["k"], df["prod"], "ko-")
ax[1].set_xlabel("$k = \sqrt{y + 1}$", fontsize=20)
ax[1].set_title("$y = k^2 - 1$", fontsize=20)
ax[2].plot(df["n1"], df["k"], "ko-", label="$k = \sqrt{y + 1}$")
ax[2].plot(df["n1"], df["k = n^2 + 3n + 1"], "r--", label="$k = n^2 + 3n + 1$")
ax[2].legend(fontsize=14)
ax[2].set_ylabel("$k = \sqrt{y + 1}$", fontsize=20)
ax[2].set_xlabel("$n$", fontsize=20)
fig.suptitle(f"Product of 4 consecutive integers is 1 less than a perfect square.", fontsize=20)
fig.tight_layout()
# -
# ## Related works
#
#
# * <NAME>. <NAME>. "The product of consecutive integers is never a power." Illinois J. Math. 19 (2) 292 - 301, June 1975. https://doi.org/10.1215/ijm/1256050816
# ## Visual Proof
nums = np.arange(10,10+4)
A = np.zeros((nums[0], nums[-1]))
A[:, nums[0]:] = 1
sns.heatmap(A, linewidth=2, cbar=False, vmin=0, vmax=4)
# +
nums = np.arange(10,10+4)
A = np.zeros((nums[1], nums[2]))
A[:, nums[0]:] = 2
A[nums[0]:, :] = 3
A[nums[0]:, nums[0]:] = 1
sns.heatmap(A, linewidth=2, cbar=False, vmin=0, vmax=4)
# -
import matplotlib.animation as animation
from IPython.display import HTML
# +
fig, ax = plt.subplots(1,1)
frames = []
nums = np.arange(10,10+4)
A = np.zeros((nums[1], nums[-1]))
im = ax.pcolormesh(A, cmap="inferno", vmin=0, vmax=4)
title = ax.set_title(f"Start")
ax.invert_yaxis()
ax.set_xticks(np.arange(A.shape[1]))
ax.set_yticks(np.arange(A.shape[0]))
ax.grid(which="major", color="w", linestyle='-', linewidth=3)
def init():
im.set_array(A)
title.set_text("")
return im, title
def animate(i):
text = ""
if i == 0:
A[:, nums[0]:] = 4
A[nums[0]:, :] = 4
text = "$n * n$"
if i == 1:
A[:, nums[0]:] = 2
A[nums[0]:, ] = 4
text = "$n * (n+3)$"
if i == 2:
A[:, nums[0]:] = 2
A[:, nums[2]:] = 3
A[nums[0]:, ] = 4
text = "$n * (n+3)$"
if i == 3:
A[:, nums[2]:] = 4
A[nums[0]:, :] = 3
A[nums[0]:, nums[0]:] = 4
A[nums[0]:, nums[0]:nums[2]] = 4
text = "$(n+1) * (n+2)$"
if i == 4:
A[nums[0]:, nums[0]:nums[2]] = 1
text = "$n * (n+3) = (n+1)*(n+2) - 2$"
# print(A)
im.set_array(A)
title.set_text(f"Step: {i} | {text}")
return im, title
# ax = sns.heatmap(A, linewidth=2, cbar=False, vmin=0, vmax=4)
fig.tight_layout()
ani = animation.FuncAnimation(fig,animate,frames=5,interval=2000,blit=True,repeat=True)
HTML(ani.to_html5_video())
# frames
# +
# ax.cla()
nums = np.arange(10,10+4)
A = np.zeros((nums[1], nums[-1]))
A[:, nums[0]:] = 2
A[nums[0]:, ] = 4
sns.heatmap(A, linewidth=2, cbar=False, vmin=0, vmax=4)
plt.show()
# plt.pause(1)
A[:, nums[2]:] = 4
sns.heatmap(A, linewidth=2, cbar=False, vmin=0, vmax=4)
plt.show()
# plt.pause(1)
A[nums[0]:, :] = 2
A[nums[0]:, nums[0]:] = 4
A[nums[0]:, nums[0]:nums[2]] = 1
sns.heatmap(A, linewidth=2, cbar=False, vmin=0, vmax=4)
plt.show()
# plt.pause(1)
# -
nums = np.arange(10,10+4)
A = np.zeros((nums[1], nums[-1]))
A[:, nums[0]:] = 2
A[nums[0]:, :] = 3
A[nums[0]:, nums[0]:] = 1
A[:, nums[2]:] = 4
sns.heatmap(A, linewidth=2, cbar=False, vmin=0, vmax=4)
| Slide Notebooks/Product of consecutive numbers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Getting Started with Boutiques
#
# As you've seen from our documentation, Boutiques is a flexible way to represent command line executables and distribute them across compute ecosystems consistently. A Boutiques tool descriptor is a JSON file that fully describes the input and output parameters and files for a given command line call (or calls, as you can include pipes(`|`) and ampersands (`&`)). There are several ways Boutiques helps you build a tool descriptor for your tool:
#
# - The [boutiques command-line utility](https://github.com/boutiques/boutiques/) contains a validator, simulator, and other tools which can help you either find an existing descriptor you wish to model yours after, or build and test your own.
# - The [examples](https://github.com/aces/cbrain-plugins-neuro/tree/master/cbrain_task_descriptors) provide useful references for development.
#
# To help you aid in this process, we will walk through the process of making an tool descriptor for [FSL's BET](http://fsl.fmrib.ox.ac.uk/fsl/fslwiki/BET) (finished product found [here](https://github.com/aces/cbrain-plugins-neuro/blob/master/cbrain_task_descriptors/fsl_bet.json)).
# ## Step 1: Describing the command line
#
# The first step in creating an tool descriptor for your command line call is creating a fully descriptive list of your command line options. If your tool was written in Python and you use the `argparse` library, then this is already done for you in large part. For many tools (bash, Python, or otherwise) this can be obtained by typing executing it with the `-h` flag. In the case of FSL's BET, we get the following:
# + language="bash"
# bet -h
# -
# Looking at all of these flags, we see a list of options which can be summarized by:
# ```
# bet [INPUT_FILE] [MASK] [FRACTIONAL_INTENSITY] [VERTICAL_GRADIENT] [CENTER_OF_GRAVITY] [OVERLAY_FLAG] [BINARY_MASK_FLAG] [APPROX_SKULL_FLAG] [NO_SEG_OUTPUT_FLAG] [VTK_VIEW_FLAG] [HEAD_RADIUS] [THRESHOLDING_FLAG] [ROBUST_ITERS_FLAG] [RES_OPTIC_CLEANUP_FLAG] [REDUCE_BIAS_FLAG] [SLICE_PADDING_FLAG] [MASK_WHOLE_SET_FLAG] [ADD_SURFACES_FLAG] [ADD_SURFACES_T2] [VERBOSE_FLAG] [DEBUG_FLAG]
# ```
#
# Now that we have summarized all command line options for our tool - some of which describe inputs and others, outputs - we can begin to craft our JSON Boutiques tool descriptor.
# ## Step 2: Understanding Boutiques + JSON
#
# For those unfamiliar with JSON, we recommend following this [3 minute JSON tutorial](http://www.secretgeek.net/json_3mins) to get you up to speed. In short, a JSON file is a dictionary object which contains *keys* and associated *values*. A *key* informs us what is being described, and a *value* is the description (which, importantly, can be arbitrarily typed). The Boutiques tool descriptor is a JSON file which requires the following keys, or, properties:
# - `name`
# - `description`
# - `schema-version`
# - `command-line`
# - `inputs`
# - `output-files`
#
# Some additional, optional, properties that a Boutiques fill will recognize are:
# - `groups`
# - `tool-version`
# - `suggested-resources`
# - `container-image`:
# - `type`
# - `image`
# - `index`
#
# In the case of BET, we will of course populate the required elements, but will also include `tool-version` and `groups`.
# ## Step 3: Populating the tool descriptor
#
# We will break-up populating the tool descriptor into two sections: adding meta-parameters (such as `name`, `description`, `schema-version`, `command-line`, `tool-version`, and `docker-image`, `docker-index` if we were to include them) and i/o-parameters (such as `inputs`, `output-files`, and `groups`).
#
# Currently, before adding any details, our tool descriptor should looks like this:
#
# ```
# {
# "name" : TODO,
# "tool-version": TODO,
# "description": TODO,
# "command-line": TODO,
# "scheme-version": TODO,
# "inputs": TODO,
# "output-files": TODO,
# }
# ```
# ### Step 3.1: Adding meta-parameters
#
# Many of the meta-parameters will be obvious to you if you're familiar with the tool, or extractable from the message received earlier when you passed the `-h` flag into your program. We can update our JSON to be the following:
#
# ```
# {
# "name" : "fsl_bet",
# "tool-version" : "1.0.0",
# "description" : "Automated brain extraction tool for FSL",
# "command-line" : "bet [INPUT_FILE] [MASK] [FRACTIONAL_INTENSITY] [VERTICAL_GRADIENT] [CENTER_OF_GRAVITY] [OVERLAY_FLAG] [BINARY_MASK_FLAG] [APPROX_SKULL_FLAG] [NO_SEG_OUTPUT_FLAG] [VTK_VIEW_FLAG] [HEAD_RADIUS] [THRESHOLDING_FLAG] [ROBUST_ITERS_FLAG] [RES_OPTIC_CLEANUP_FLAG] [REDUCE_BIAS_FLAG] [SLICE_PADDING_FLAG] [MASK_WHOLE_SET_FLAG] [ADD_SURFACES_FLAG] [ADD_SURFACES_T2] [VERBOSE_FLAG] [DEBUG_FLAG]",
# "schema-version" : "0.4",
# "inputs": TODO,
# "output-files": TODO,
# "groups": TODO
# }
# ```
# ### Step 3.2: Adding i/o parameters
#
# Inputs and outputs of many applications are complicated - outputs can be dependent upon input flags, flags can be mutually exclusive or require at least one option, etc. The way Boutiques handles this is with a detailed schema which consists of options for inputs and outputs, as well as optionally specifying groups of inputs which may add additional layers of input complexity.
#
# As you have surely noted, tools do only contain a single "name" or "version" being used, but may have many input and output parameters. This means that inputs, outputs, and groups, will be described as a list. Each element of these lists will be a dictionary following the input, output, or group schema, respectively. This means that our JSON actually looks more like this:
#
# ```
# {
# "name" : "fsl_bet",
# "tool-version" : "1.0.0",
# "description" : "Automated brain extraction tool for FSL",
# "command-line" : "bet [INPUT_FILE] [MASK] [FRACTIONAL_INTENSITY] [VERTICAL_GRADIENT] [CENTER_OF_GRAVITY] [OVERLAY_FLAG] [BINARY_MASK_FLAG] [APPROX_SKULL_FLAG] [NO_SEG_OUTPUT_FLAG] [VTK_VIEW_FLAG] [HEAD_RADIUS] [THRESHOLDING_FLAG] [ROBUST_ITERS_FLAG] [RES_OPTIC_CLEANUP_FLAG] [REDUCE_BIAS_FLAG] [SLICE_PADDING_FLAG] [MASK_WHOLE_SET_FLAG] [ADD_SURFACES_FLAG] [ADD_SURFACES_T2] [VERBOSE_FLAG] [DEBUG_FLAG]",
# "schema-version" : "0.4",
# "inputs": [
# {TODO},
# {TODO},
# ...
# ],
# "output-files": [
# {TODO},
# {TODO},
# ...
# ],
# }
# ```
#
# As the file is beginning to grow considerably in number of lines, we will no longer show you the full JSON at each step but will simply show you the dictionaries responsible for output, input, and group entries.
# #### Step 3.2.1: Specifying inputs
#
# The input schema contains several options, many of which can be ignored in this first example with the exception of `id`, `name`, and `type`. For BET, there are several input values we can choose to demonstrate this for you. We have chosen three with considerably different functionality and therefore schemas. In particular:
# - `[INPUT_FILE]`
# - `[FRACTIONAL_INTENSITY]`
# - `[CENTER_OF_GRAVITY]`
#
# **`[INPUT_FILE]`** The simplest of these in the `[INPUT_FILE]` which is a required parameter that simply expects a qualified path to a file. The dictionary entry is:
# ```
# {
# "id" : "infile",
# "name" : "Input file",
# "type" : "File",
# "description" : "Input image (e.g. img.nii.gz)",
# "optional": false,
# "value-key" : "[INPUT_FILE]"
# }
# ```
#
# **`[FRACTIONAL_INTENSITY]`** This parameter documents an optional flag that can be passed to the executable. Along with the flag, when it is passed, is a floating point value that can range from 0 to 1. We are able to validate at the level of Boutiques whether or not a valid input is passed, so that jobs are not submitted to the execution engine which will error, but they get flagged upon validation of inputs. This dictionary is:
# ```
# {
# "id" : "fractional_intensity",
# "name" : "Fractional intensity threshold",
# "type" : "Number",
# "description" : "Fractional intensity threshold (0->1); default=0.5; smaller values give larger brain outline estimates",
# "command-line-flag": "-f",
# "optional": true,
# "value-key" : "[FRACTIONAL_INTENSITY]",
# "integer" : false,
# "minimum" : 0,
# "maximum" : 1
# }
# ```
#
# **`[CENTER_OF_GRAVITY]`** The center of gravity value expects a triple (i.e. [X, Y, Z] position) if the flag is specified. Here we are able to set the condition that the length of the list received after this flag is 3, by specifying that the input is a list that has both a minimum and maximum length.
# ```
# {
# "id" : "center_of_gravity",
# "name" : "Center of gravity vector",
# "type" : "Number",
# "description" : "The xyz coordinates of the center of gravity (voxels, not mm) of initial mesh surface. Must have exactly three numerical entries in the list (3-vector).",
# "command-line-flag": "-c",
# "optional": true,
# "value-key" : "[CENTER_OF_GRAVITY]",
# "list" : true,
# "min-list-entries" : 3,
# "max-list-entries" : 3
# }
# ```
#
# For further examples of different types of inputs, feel free to explore [more examples](https://github.com/aces/cbrain-plugins-neuro/tree/master/cbrain_task_descriptors).
# #### Step 3.2.2: Specifying outputs
#
# The output schema also contains several options, with the only mandatory ones being `id`, `name`, and `path-template`. We again demonstrate an example from BET:
# - `outfile`
#
# **`outfile`** All of the output parameters in BET are similarly structured, and exploit the same core functionality of basing the output file, described by `path-template`, as a function of an input value on the command line, here given by `[MASK]`. The `optional` flag also describes whether or not a derivative should always be produced, and whether Boutiques should indicate an error if a file isn't found. The output descriptor is thus:
#
# ```
# {
# "id" : "outfile",
# "name" : "Output mask file",
# "description" : "Main default mask output of BET",
# "path-template" : "[MASK].nii.gz",
# "optional" : true
# }
# ```
#
# An extension of the feature of naming outputs based on inputs exists in newer versions of the schema than this example was originally developed, and enable stripping the extension of the input values used, as well. An example of this can be seen [here](https://github.com/neurodata/boutiques-tools/blob/master/cbrain_task_descriptors/ndmg.json#L158).
# #### Step 3.2.3: Specifying groups
#
# The group schema enables provides an additional layer of complexity when considering the relationships between inputs. For instance, if multiple inputs within a set are mutually exclusive, they may be grouped and a flag set indicating that only one can be selected. Alternatively, if at least one option within a group must be specified, the user can also set a flag indicating such. The following group from the BET implementation is used to illustrate this:
# - `variational_params_group`
#
# **`variational_params_group`** Many flags exist in BET, and each of them is represented in the command line we specified earlier. However, as you may have noticed when reading the output of `bet -h`, several of these options are mutually exclusive to one another. In order to again prevent jobs from being submitted to a scheduler and failing there, Boutiques enables grouping of inputs and forcing such mutual exclusivity so that the invalid inputs are flagged in the validation stage. This group dictionary is:
# ```
# {
# "id" : "variational_params_group",
# "name" : "Variations on Default Functionality",
# "description" : "Mutually exclusive options that specify variations on how BET should be run.",
# "members" : ["robust_iters_flag", "residual_optic_cleanup_flag", "reduce_bias_flag", "slice_padding_flag", "whole_set_mask_flag", "additional_surfaces_flag", "additional_surfaces_t2"],
# "mutually-exclusive" : true
# }
# ```
#
# Though an example of `one-is-required` input groups is not available in our BET example, you can investigate a validated tool descriptor [here](https://github.com/neurodata/boutiques-tools/blob/master/cbrain_task_descriptors/ndmg.json#L13) to see how it is implemented.
# ### Step 3.3: (optional) Extending the tool descriptor
#
# Now that the basic implementation of this tool has been done, you can check out the [schema](https://github.com/boutiques/boutiques/blob/master/tools/python/boutiques/schema/descriptor.schema.json) to explore deeper functionality of Boutiques. For example, if you have created a Docker or Singularity container, you can associate an image with your tool descriptor and any compute resource with Docker or Singularity installed will launch the executable through them (an example of using Docker can be found [here](https://github.com/neurodata/boutiques-tools/blob/master/cbrain_task_descriptors/ndmg.json#L6)).
# ## Step 4: Validating the tool descriptor
#
# Once you've completed your Boutiques tool descriptor, you should run the [validator](https://github.com/boutiques/boutiques#validation) to ensure that you have created it correctly. The `README.md` [here](https://github.com/boutiques/boutiques/) describes how to install and use the validator and remainder of the Boutiques shell (`bosh`) tools on your tool descriptor.
# ## Step 5: Using the tool descriptor
#
# Once the tool descriptor has been validated, your tool is now ready to be integrated in a platform that supports Boutiques. You can use the `localExec.py` tool described [here](https://github.com/boutiques/boutiques/tree/master/tools) to launch your container locally for preliminary testing. Once you feel comfortable with your tool, you can contact your system administrator and have them integrate it into their compute resources so you can test and use it to process your data.
| examples/Getting Started with Boutiques.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Tybalt latent space arithmetic with high grade serous ovarian cancer subtypes
#
# Recent applications of generative models (GANs and VAEs) in image processing has demonstrated the remarkable ability of the latent dimensions to capture a meaningful manifold representation of the input space. Here, we assess if the VAE learns a latent space that can be mathematically manipulated to reveal insight into the gene expression activation patterns of high grade serous ovarian cancer (HGSC) subtypes.
#
# Several previous studies have reported the presence of four gene expression based HGSC subtypes. However, we recently [published a paper](https://doi.org/10.1534/g3.116.033514) that revealed the inconsistency of subtype assignments across populations. We observed repeatable structure in the data transitioning between setting clustering algorithms to find different solutions. For instance, when setting algorithms to find 2 subtypes, the mesenchymal and immunoreactive and the proliferative and differentiated subtype consistently collapsed together. These observations may suggest that the subtypes exist on a gene expression continuum of differential activation patterns, and may only be artificially associated with "subtypes". Here, we test if the VAE can help to identify some differential patterns of expression.
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
plt.style.use('seaborn-notebook')
sns.set(style='white', color_codes=True)
sns.set_context('paper', rc={'font.size':12, 'axes.titlesize':15, 'axes.labelsize':20,
'xtick.labelsize':14, 'ytick.labelsize':14})
# Set seed for plotting
np.random.seed(123)
rnaseq_file = os.path.join('data', 'pancan_scaled_zeroone_rnaseq.tsv.gz')
rnaseq_df = pd.read_table(rnaseq_file, index_col=0)
rnaseq_df.shape
ov_file = os.path.join('data', 'ov_subtype_info.tsv')
ov_df = pd.read_table(ov_file, index_col=0)
ov_df.head(2)
encoded_file = os.path.join('data', "encoded_rnaseq_onehidden_warmup_batchnorm.tsv")
encoded_df = pd.read_table(encoded_file, index_col=0)
print(encoded_df.shape)
encoded_df.head(2)
# +
# Subset and merge the HGSC subtype info with the latent space feature activations
ov_samples = list(set(encoded_df.index) & (set(ov_df.index)))
ov_encoded = encoded_df.loc[ov_samples, ]
ov_encoded_subtype = pd.merge(ov_df.loc[:, ['SUBTYPE', 'SILHOUETTE WIDTH']], ov_encoded,
how='right', left_index=True, right_index=True)
ov_encoded_subtype = ov_encoded_subtype.assign(subtype_color = ov_encoded_subtype['SUBTYPE'])
ov_subtype_color_dict = {'Differentiated': 'purple',
'Immunoreactive': 'green',
'Mesenchymal': 'blue',
'Proliferative': 'red'}
ov_encoded_subtype = ov_encoded_subtype.replace({'subtype_color': ov_subtype_color_dict})
print(ov_encoded_subtype.shape)
ov_encoded_subtype.head(2)
# -
# Get the HGSC mean feature activation
ov_mean_subtypes = ov_encoded_subtype.groupby('SUBTYPE').mean()
ov_mean_subtypes
# ## HGSC Subtype Arithmetic
#
# Because of the relationship observed in the consistent clustering solutions, perform the following subtractions
#
# 1. Immunoreactive - Mesenchymal
# 2. Differentiated - Proliferative
#
# The goal is to observe the features with the largest difference between the aformentioned comparisons. The differences should be in absolute directions
# ### 1) Immunoreactive - Mesenchmymal
mes_mean_vector = ov_mean_subtypes.loc['Mesenchymal', [str(x) for x in range(1, 101)]]
imm_mean_vector = ov_mean_subtypes.loc['Immunoreactive', [str(x) for x in range(1, 101)]]
# +
high_immuno = (imm_mean_vector - mes_mean_vector).sort_values(ascending=False).head(2)
high_mesenc = (imm_mean_vector - mes_mean_vector).sort_values(ascending=False).tail(2)
print("Features with large differences: Immuno high, Mesenchymal low")
print(high_immuno)
print("Features with large differences: Mesenchymal high, Immuno low")
print(high_mesenc)
# -
# Select to visualize encoding 56 because it has high immuno and low everything else
ov_mean_subtypes.loc[:, ['87', '77', '56']]
# Node 87 has high mesenchymal, low immunoreactive
node87_file = os.path.join('figures', 'node87_distribution_ovsubtype.pdf')
g = sns.swarmplot(y = '87', x = 'SUBTYPE', data=ov_encoded_subtype,
order=['Mesenchymal', 'Immunoreactive', 'Proliferative', 'Differentiated']);
g.set(xlabel='', ylabel='encoding 87')
plt.xticks(rotation=0);
plt.tight_layout()
plt.savefig(node87_file)
# Node 77 has high immunoreactive, low mesenchymal
node77_file = os.path.join('figures', 'node77_distribution_ovsubtype.pdf')
g = sns.swarmplot(y = '77', x = 'SUBTYPE', data=ov_encoded_subtype,
order=['Mesenchymal', 'Immunoreactive', 'Proliferative', 'Differentiated']);
g.set(xlabel='', ylabel='encoding 77')
plt.xticks(rotation=0);
plt.tight_layout()
plt.savefig(node77_file)
# Node 56 has high immunoreactive, low mesenchymal (and prolif/diff)
node56_file = os.path.join('figures', 'node56_distribution_ovsubtype.pdf')
g = sns.swarmplot(y = '56', x = 'SUBTYPE', data=ov_encoded_subtype,
order=['Mesenchymal', 'Immunoreactive', 'Proliferative', 'Differentiated']);
g.set(xlabel='', ylabel='encoding 56')
plt.xticks(rotation=0);
plt.tight_layout()
plt.savefig(node56_file)
# ### 2) Differentiated - Proliferative
pro_mean_vector = ov_mean_subtypes.loc['Proliferative', [str(x) for x in range(1, 101)]]
dif_mean_vector = ov_mean_subtypes.loc['Differentiated', [str(x) for x in range(1, 101)]]
# +
high_differ = (dif_mean_vector - pro_mean_vector).sort_values(ascending=False).head(2)
high_prolif = (dif_mean_vector - pro_mean_vector).sort_values(ascending=False).tail(2)
print("Features with large differences: Differentiated high, Proliferative low")
print(high_differ)
print("Features with large differences: Proliferative high, Differentiated low")
print(high_prolif)
# -
# Select to visualize encoding 56 because it has high immuno and low everything else
ov_mean_subtypes.loc[:, ['38', '79']]
# Node 38 has high differentiated, low proliferative
node38_file = os.path.join('figures', 'node38_distribution_ovsubtype.pdf')
g = sns.swarmplot(y = '38', x = 'SUBTYPE', data=ov_encoded_subtype,
order=['Mesenchymal', 'Immunoreactive', 'Proliferative', 'Differentiated']);
g.set(xlabel='', ylabel='encoding 38')
plt.xticks(rotation=0);
plt.tight_layout()
plt.savefig(node38_file)
# Node 79 has high proliferative, low differentiated
node79_file = os.path.join('figures', 'node79_distribution_ovsubtype.pdf')
g = sns.swarmplot(y = '79', x = 'SUBTYPE', data=ov_encoded_subtype,
order=['Mesenchymal', 'Immunoreactive', 'Proliferative', 'Differentiated']);
g.set(xlabel='', ylabel='encoding 79')
plt.xticks(rotation=0);
plt.tight_layout()
plt.savefig(node79_file)
# ### Explore weights that explain the nodes
def get_high_weight(weight_matrix, node, high_std=2.5, direction='positive'):
"""
Determine high weight genes given a gene weight matrix and feature
Output tab separated file
"""
genes = weight_matrix.loc[node, :].sort_values(ascending=False)
if direction == 'positive':
node_df = (genes[genes > genes.std() * high_std])
abbrev = 'pos'
elif direction == 'negative':
node_df = (genes[genes < -1 * (genes.std() * high_std)])
abbrev = 'neg'
node_df = pd.DataFrame(node_df).reset_index()
node_df.columns = ['genes', 'weight']
node_base_file = 'hgsc_node{}genes_{}.tsv'.format(node, abbrev)
node_file = os.path.join('results', node_base_file)
node_df.to_csv(node_file, index=False, sep='\t')
return node_df
# +
# Obtain the decoder weights
weight_file = os.path.join('results', 'tybalt_gene_weights.tsv')
weight_df = pd.read_table(weight_file, index_col=0)
weight_df.head(2)
# +
# Output high weight genes for nodes representing mesenchymal vs immunoreactive
node87pos_df = get_high_weight(weight_df, node=87)
node87neg_df = get_high_weight(weight_df, node=87, direction='negative')
node77pos_df = get_high_weight(weight_df, node=77)
node77neg_df = get_high_weight(weight_df, node=77, direction='negative')
node56pos_df = get_high_weight(weight_df, node=56)
node56neg_df = get_high_weight(weight_df, node=56, direction='negative')
# +
# Output high weight genes for nodes representing proliferative vs differentiated
node79pos_df = get_high_weight(weight_df, node=79)
node79neg_df = get_high_weight(weight_df, node=79, direction='negative')
node38pos_df = get_high_weight(weight_df, node=38)
node38neg_df = get_high_weight(weight_df, node=38, direction='negative')
| hgsc_subtypes_tybalt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # hash_to_emoji
# Mitchell / Isthmus - July 2020
#
# Twitter recently applied a filter that appears to block any tweets containing alphanumeric strings longer than 26 characters. Unfortunately this includes hash digests (among many other use cases).
#
# This inspired the latest cryptographic stenographic innovation for censorship resistance: `hash_to_emoji`
#
# ### Example
#
# Input: `some prediction for the future`
#
# Output: 🐇🐈☁❄☃☃🌁🐕🌀💀☃🌁🎺🐕☃🐁✉👀🌁👀🌀🌀🐕🐁☁☃🌀☃🐈👀👍🐇☃🐈🎺🐕☂☃🐈🐇🐇❄🔔🐇❄💀☁🐇🐇☂👍☁🐕☁🔔💀🐈👍👍❄🐇🌀☃💀
#
# ### Notes
#
# - The 1:1 mapping from hex representation digit to emoji is painfully inefficient. Shorter final digests could be produced by using more characters from the large emoji set.
# - A possible extension would be an efficient (bidirectional) translation between arbitrary data blobs and emoji strings. (Silly example: can't access a p2p blockchain network to broadcast your transaction? Just convert it to an emoji string and tweet at @xyzGateway to be added to the main mempool)
# ## Import libraries
# #!pip install emoji
import emoji
import hashlib
# ## Inputs
message_to_hash = 'some prediction for the future'
# ## Process
# ### Calculate hash
# You can easily swap in different algorithms from hashlib
raw_hash = hashlib.sha256(message_to_hash.encode()).hexdigest()
# ### Convert alphanumeric hash to emoji set
mapping = {
"0":":skull:",
"1":":umbrella:",
"2":":cloud:",
"3":":snowflake:",
"4":":snowman:",
"5":":trumpet:",
"6":":cyclone:",
"7":":foggy:",
"8":":eyes:",
"9":":cat:",
"a":":dog:",
"b":":mouse:",
"c":":bell:",
"d":":rabbit:",
"e":":envelope:",
"f":":thumbs_up:"
}
output_vec = str()
for i in range(len(raw_hash)):
this_char = raw_hash[i]
output_vec = output_vec + mapping[this_char]
# ## Provide output
emoji_str = emoji.emojize(output_vec)
print(emoji.emojize('\nHash digest:\n\n' + emoji_str))
| hash_to_emoji.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## LOGISITC REGRESSION
# * typically a binary classification problem - Class 1 or Class 0
# * Project the group scores onto a sigmoid function that better covers both the classes than a straight line
# * Set a threshold to the best and classify into groups
#
#
# ### Mathematically
# * We have likelihood functions for x, y and beta.
# * Estimate the Beta - MLE. Select the beta value that maximizes the probability of observing data into the right class for the given vector of x points. For the given set of x values, a probability of y in class 1 or class 0 is projected. This is the likelihood of Beta. Now we maximize this.
# * The maximizing is done in two steps - Take the log likelihood, apply gradient descent. To apply the GD, we use the log loss function, that is the exact opposite of Log likelihood. We start by randomly choosing the beta value, keep iterating and arrive at the minimum error WHICH becomes the MLE of Beta
# * Use learning rate to control the gradient of the Betas
# +
## main function
def logistic_regression(x,y, iterations = 100, learning_rate = 0.01):
m,n = len(x), len(x[0])
beta_0, beta_other = initialize_params(n)
for i in range(iterations):
gradient_beta_0, gradient_beta_other = (compute_gradients(x,y,beta_0, beta_other,m,n, 50))
beta_0, beta_other = update_params(beta_0, beta_other, gradient_beta_0, gradient_beta_other, learning_rate)
return beta_0, beta_other
# +
#supporting functions
# initializing the beta parameters,random start for the gradient descent
def initialize_params(dimensions):
beta_0 = 0
beta_other = [random.random() for i in range(dimensions)]
return beta_0, beta_other
# +
# Compute functions
def compute_gradients(x,y,beta_0, beta_other, n,m):
gradient_beta_0 = 0
gradient_beta_other = [0]*n
for i , point in enumerate(x): # computing gradients for each data point in x
prediction = logistic_regression(point, beta_0, beta_other) # getting the prediction for that point
for j, features in enumerate(point): # compute the gradient for that single point
gradient_beta_other[j] += (pred - y[i])*features/m #similar to the function I wrote for Gradient at Betaj. Accumulate the data point from all data points and normalize them by /m
gradient_beta_0 += (pred - y[i])/m
return gradient_beta_0, gradient_beta_other
# +
#Mini - Batch gradient descent
def compute_gradients_minibatch(x,y,beta_0, beta_other, n,m, batch_size):
gradient_beta_0 = 0
gradient_beta_other = [0]*n
for i in range(batch_size): # we pick a batch size and perform validation. Something like cross validatipn
i = random.randint(0, m-1)
point = x[i]
prediction = logistic_regression(point, beta_0, beta_other) # getting the prediction for that point
for j, features in enumerate(point): # compute the gradient for that single point
gradient_beta_other[j] += (pred - y[i])*features/m #similar to the function I wrote for Gradient at Betaj. Accumulate the data point from all data points and normalize them by /m
gradient_beta_0 += (pred - y[i])/m
return gradient_beta_0, gradient_beta_other
# +
# Update the paramters
def update_params(beta_0, beta_other, gradient_beta_0, gradient_beta_other, learning_rate):
beta_0 -= gradient_beta_0 * learning_rate
for i in range(len(beta_other)):
beta_other[i] -= (gradient_beta_other[i])*learning_rate
return beta_0, beta_other
# -
| Logistic_Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Starbucks Capstone Challenge
#
# ### Introduction
#
# This data set contains simulated data that mimics customer behavior on the Starbucks rewards mobile app. Once every few days, Starbucks sends out an offer to users of the mobile app. An offer can be merely an advertisement for a drink or an actual offer such as a discount or BOGO (buy one get one free). **Some users might not receive any offer during certain weeks.**
#
# **Not all users receive the same offer, and that is the challenge to solve with this data set.**
#
# **Your task is to combine transaction, demographic and offer data to determine which demographic groups respond best to which offer type.** This data set is a simplified version of the real Starbucks app because the underlying simulator only has one product whereas Starbucks actually sells dozens of products.
#
# Every offer has a validity period before the offer expires. As an example, a BOGO offer might be valid for only 5 days. **You'll see in the data set that informational offers have a validity period even though these ads are merely providing information about a product;** for example, if an informational offer has 7 days of validity, you can assume the customer is feeling the influence of the offer for 7 days after receiving the advertisement.
#
# You'll be given transactional data showing user purchases made on the app including the timestamp of purchase and the amount of money spent on a purchase. This transactional data also has a record for each offer that a user **receives** as well as a record for when a user actually **views** the offer. There are also records for when a user **completes** an offer.
#
# Keep in mind as well that **someone using the app might make a purchase through the app without having received an offer or seen an offer.**
#
# ### Example
#
# To give an example, a user could receive a discount offer buy 10 dollars get 2 off on Monday. The offer is valid for 10 days from receipt. If the customer accumulates at least 10 dollars in purchases during the validity period, the customer completes the offer.
#
# However, there are a few things to watch out for in this data set. Customers do not opt into the offers that they receive; in other words, **a user can receive an offer, never actually view the offer, and still complete the offer.** For example, a user might receive the "buy 10 dollars get 2 dollars off offer", but the user never opens the offer during the 10 day validity period. The customer spends 15 dollars during those ten days. There will be an offer completion record in the data set; however, the customer was not influenced by the offer because the customer never viewed the offer.
#
# ### Cleaning
#
# This makes data cleaning especially important and tricky.
#
# You'll also want to take into account that some demographic groups will make purchases even if they don't receive an offer. From a business perspective, if a customer is going to make a 10 dollar purchase without an offer anyway, you wouldn't want to send a buy 10 dollars get 2 dollars off offer. **You'll want to try to assess what a certain demographic group will buy when not receiving any offers.**
#
# ### Final Advice
#
# Because this is a capstone project, you are free to analyze the data any way you see fit. For example, you could build a **machine learning model that predicts how much someone will spend based on demographics and offer type**. Or you could build **a model that predicts whether or not someone will respond to an offer**. Or, you don't need to build a machine learning model at all. You could develop a set of **heuristics that determine what offer you should send to each customer** (ie 75 percent of women customers who were 35 years old responded to offer A vs 40 percent from the same demographic to offer B, so send offer A).
# # Data Sets
#
# The data is contained in three files:
#
# * portfolio.json - containing offer ids and meta data about each offer (duration, type, etc.)
# * profile.json - demographic data for each customer
# * transcript.json - records for transactions, offers received, offers viewed, and offers completed
#
# Here is the schema and explanation of each variable in the files:
#
# **portfolio.json**
# * id (string) - offer id
# * offer_type (string) - type of offer ie BOGO, discount, informational
# * difficulty (int) - minimum required spend to complete an offer
# * reward (int) - reward given for completing an offer
# * duration (int) -
# * channels (list of strings)
#
# **profile.json**
# * age (int) - age of the customer
# * became_member_on (int) - date when customer created an app account
# * gender (str) - gender of the customer (note some entries contain 'O' for other rather than M or F)
# * id (str) - customer id
# * income (float) - customer's income
#
# **transcript.json**
# * event (str) - record description (ie transaction, offer received, offer viewed, etc.)
# * person (str) - customer id
# * time (int) - time in hours. The data begins at time t=0
# * value - (dict of strings) - either an offer id or transaction amount depending on the record
# ## I. Business Understanding
# Summarizing the above Introduction, we are going to:
# - Combine transaction, demographic and offer data to **analyze** which demographic groups respond(i.e.view&complete) best to which offer type;
# - Build a supervised learning model(specifically, a classification model) that predicts whether or not someone will respond to an offer
# ## II. Data Understanding and Data Engineering
# +
import pandas as pd
import numpy as np
import math
import json
% matplotlib inline
# read in the json files
portfolio = pd.read_json('data/portfolio.json', orient='records', lines=True)
profile = pd.read_json('data/profile.json', orient='records', lines=True)
transcript = pd.read_json('data/transcript.json', orient='records', lines=True)
# -
import matplotlib.pyplot as plt
import seaborn as sns
# ### 1. Portfolio Data Preprocessing
#create dummy columns
portfolio=portfolio.join(portfolio['channels'].str.join('|').str.get_dummies().add_prefix('channel_'))
portfolio=portfolio.drop('channels',axis=1)
#10 kinds of offers
portfolio
# ### 2. Profile Visualizations
profile.sample(5)
#Age Visualization
count_by_agegroup=profile.groupby(pd.cut(profile['age'], np.arange(0, 118+5, 5)))['id'].count()
plt.figure(figsize=(12,4))
plt.bar(np.arange(0, 118, 5),count_by_agegroup,width=3.5, align='edge')
plt.xticks(np.arange(0, 118, 5))
plt.xlabel('Age')
plt.ylabel('Persons Counts')
plt.title('The Persons Number of Different Age')
plt.show()
count_by_incomegroup=profile.groupby(pd.cut(profile['income'], np.arange(20000, 120000+10000, 10000)))['id'].count()
count_by_incomegroup.plot.pie(figsize=(6, 6))
# +
def make_autopct(values):
def my_autopct(pct):
total = sum(values)
val = int(round(pct*total/100.0))
return '{p:.2f}% ({v:d})'.format(p=pct,v=val)
return my_autopct
fig, ax = plt.subplots(figsize=(8, 8), subplot_kw=dict(aspect="equal"))
labels=['(20000, 30000]','(30000, 40000]','(40000, 50000]','(50000, 60000]','(60000, 70000]',\
'(70000, 80000]','(80000, 90000]','(90000, 100000]','(100000, 110000]','(110000, 120000]']
porcent = 100.*count_by_incomegroup/count_by_incomegroup.sum()
patches, texts, autotexts = ax.pie(count_by_incomegroup, autopct=make_autopct(count_by_incomegroup),startangle=90,counterclock=False)
labels = ['{0} - {1:1.2f} %'.format(i,j) for i,j in zip(labels, porcent)]
'''
sort_legend = True
if sort_legend:
patches, labels, dummy = zip(*sorted(zip(patches, labels, count_by_incomegroup),
key=lambda x: x[2],
reverse=True))
'''
ax.legend(patches, labels,
title='The Persons Number of Different Income',
loc="center left",
bbox_to_anchor=(1, 0, 0.5, 1))
plt.title('The Persons Number of Different Income')
#plt.legend(patches, labels, loc="lower right", bbox_transform=plt.gcf().transFigure)
#plt.subplots_adjust(left=0.0, bottom=0.1, right=0.45)
# -
# **Comments:** We can see that most people have income between 50000 and 80000.
# ### 3. Transcript Overview
transcript.head()
transcript[transcript.event=='offer received'].head(5)
transcript[transcript.event=='offer viewed'].head(5)
#Those purchased and have received an offer
transcript[transcript.event=='offer completed'].head(5)
transcript[transcript.event=='offer completed'].iloc[0].value
#Those purchased without an offer
transcript[transcript.event=='transaction'].head(5)
# ### 4. Data Engineering
# **Tables to be Generated:**
# - **transcript_new**: the transcript with offer id column name consolidated ((dup)rows are persons, columns are event, time, offer id, amount, reward)
# - **person_and_offer**: transcript_new joins portfolio((dup) adding offer info to transcript_new)
# - **person_offer_demographic**: person_and_offer joins parts of profile((dup) adding personal info to person_and_offer)
# - **offer**: (nodup)rows are persons, columns are counts of offers received, completed, viewed&completed, noviewed&completed, received bogo,received discount, received informational, v&c_bogo, v&c_discount, v&c_discount, v&c_informational, etc.
# - **offer_record**: (dup)rows are persons, columns are v&c offer id and time.
# - **offer_norec_comp**: (nodup)rows are persons, columns are counts of offers completed, completed bogo, completed discount, completed informational, received&completed, noreceived&completed, nr&c_bogo, nr&c_discount, nr&c_informational, etc.
# - **transaction_gen**: (nodup)rows are persons, columns are total amount of transactions(including those not completed), amount of transactions related to viewed&completed offers, amount of transactions related to noviewed&completed offers, etc.
# +
#convert dict into dummy
#transcript=pd.concat([transcript.drop(['value'], axis=1), transcript['value'].apply(pd.Series)], axis=1)
# -
transcript.head()
# Below are preprocessing code having run and saved as csv, **no need to run again.**
#it appears that those offer received and viewed have offer id in "offer id", while the offer completedc atogory have offer id in "offer_id".
#should consolidate these two columns
'''
transcript['consolidate_offer_id']=0
df_1=transcript[transcript.event=='offer received']
df_2=transcript[transcript.event=='offer viewed']
df_3=transcript[transcript.event=='offer completed']
df_4=transcript[transcript.event=='transaction']
df_1['consolidate_offer_id']=df_1['offer id']
df_2['consolidate_offer_id']=df_2['offer id']
df_3['consolidate_offer_id']=df_3['offer_id']
df_4['consolidate_offer_id']=df_4['offer_id']
transcript_new=pd.concat([df_1,df_2,df_3,df_4],axis=0)
'''
# +
#save it to read next time
#transcript_new.to_csv('data/transcript_new_2.csv')
# -
from tqdm import trange
# Read the saved output:
#transcript=pd.read_csv('data/transcript_new.csv')
transcript_new=pd.read_csv('data/transcript_new_2.csv')
del transcript_new['Unnamed: 0']
del transcript_new['offer id']
del transcript_new['offer_id']
transcript_new.sample(5)
#Visualizing those completed the offer
amount=transcript_new[transcript_new.event=='offer completed']['amount']
amount.isnull().mean()
# **Comments:** It seems that "offer completed" only has the offer id & reward; to see how it relates to transaction, it should join the "transactions"; or to see how is the difficulty, it should join the Portfolio table.
offer_completed=transcript_new[transcript_new.event=='offer completed']
transaction=transcript_new[transcript_new.event=='transaction']
#hour to day
transaction['time']=transaction['time']/24
transaction.rename(columns={'time':'transaction_time'}, inplace=True)
portfolio.rename(columns={'id':'consolidate_offer_id'}, inplace=True)
#map transcript with portfolio to see details of the offer
person_and_offer=transcript_new.merge(portfolio,on='consolidate_offer_id')
del person_and_offer['reward_x']
person_and_offer.rename(columns={'reward_y':'reward'}, inplace=True)
person_and_offer.columns
#reorder the columns
person_and_offer=person_and_offer[['event', 'person', 'consolidate_offer_id','offer_type','difficulty','amount','reward',\
'duration', 'time','channel_email',\
'channel_mobile', 'channel_social', 'channel_web']]
person_and_offer.rename(columns={'time':'offer_time'}, inplace=True)
#turn to days
person_and_offer['offer_time']=person_and_offer['offer_time']/24
person_and_offer.amount.isnull().mean()
#all nan in 'amount', hence can delete
del person_and_offer['amount']
transaction.reward.isnull().mean()
#all nan in 'reward', hence can delete
del transaction['reward']
del transaction['event']
transaction.consolidate_offer_id.isnull().mean()
#all nan in 'consolidate_offer_id', hence can delete
del transaction['consolidate_offer_id']
transaction.groupby('person').count().head()
np.unique(person_and_offer.event)
person_and_offer.reward.isnull().mean()
#sort by person and offer_time, which may help for the following transaction merge
person_and_offer=person_and_offer.sort_values(['person','offer_time'])
person_and_offer[person_and_offer.person=='003d66b6608740288d6cc97a6903f4f0']
transaction[transaction.person=='003d66b6608740288d6cc97a6903f4f0']
# **Comment:** The above cell shows that **person_and_offer** dataframe contains the tracking of the same offer.
#
# For example,
# offer "5a8bc65990b245e5a138643cd4eb9837" is received and viewed;
# offer "0b1e1539f2cc45b7b9fa7c272da2e1d7" is received and completed.
#
# Also,
# one may received a same offer **more than once**.
profile.rename(columns={'id':'person'}, inplace=True)
#merge person&offer with profile
person_offer_demographic=person_and_offer.merge(profile[['person','age','gender','income']],on='person')
person_and_offer.columns
#reorder columns
person_offer_demographic=person_offer_demographic[['event', 'person', 'age','gender','income','consolidate_offer_id', 'offer_type', 'difficulty',
'reward', 'duration', 'offer_time', 'channel_email', 'channel_mobile',
'channel_social', 'channel_web']]
person_offer_demographic.head()
#Save and reuse
person_offer_demographic.to_csv('data/person_offer_demographic.csv')
person_offer_demographic=pd.read_csv('data/person_offer_demographic.csv')
del person_offer_demographic['Unnamed: 0']
# **Comments:** In the above parts, we link person, offer and demographic information together.
#
#
# However, it is tricky to figure out **how many completed offers were from offers that the person viewed beforehand** because some of the offers were completed first and then viewed afterwards; Hence, we have to separate 'viewed and completed' offers from 'completed and then viewed' ones.
#
# Also, we are going to calculate the transactions associated with viewed and completed vs. completed but not viewed offers because for each transaction, we don't know if it was associated with a completed offer or not.
# We first conduct data engineering on offer information.
# +
#Loop through the unique persons one by one in the transcript. Each time:
#1.Extract all records for this person(using transcript['person id']==person id)
#2.Extract this person's 'offer received' records
#3.Loop through the received offers: For every loop, store the received offer time as "start", and "start" plus "duration" is the end of
#the offer valid time; A. find whether there are "offer viewed" in the time from "start" to "end", offer id equals this offer id;B. find
#whether there are "offer completed" in the time from "start" to "end", offer id equals this offer id; if A & B are satisfied, mark viewed
#&completed; if only B is satisfied, mark noviewed&completed; Use a list to keep track of viewed&completed offers person id, offer id, time
#4.After the loop in step 3, count the number of received, received bogo&discount&informational, completed, viewed&completed, noviewed&
#completed, viewed&completed bogo&discount&informational.
# -
def offer_analyzer(person_df,person,idx,offer,offer_record):
'''
This function takes in a person's events from transcript df and generate a df that includes the number and type
of offer the person received, completed, viewed and completed as well as not viewed but completed. The function
also generates a df of all viewed and completed offers by all users for future use.
inputs:
1. person_df - all events of this person
2. person - the person's id
3. idx - index to keep track of for generating 'viewed & completed offer dataframe'
4. offer - empty offer df
5. offer_record - empty offer_record df(if records viewed&completed offers)
outputs:
1. [final] - a list of all output variables including:
receive -- counts of offer received
comp -- counts of offer completed
view_comp -- counts of offer viewed and completed
noview_comp -- counts of offer completed without viewing
bogo -- counts of viewed & completed bogo offer
discount -- counts of viewed & completed discount offer
informational -- counts of viewed & completed informational offer and
10 columns for 10 types of offers-- how many times this specific offer was viewed & completed
2. idx - updated index to keep track of for generating 'viewed & completed offer dataframe'
'''
# select all offers the person received
offers = person_df[person_df.event=='offer received']
# start counting
comp = 0 # completed offers
view_comp = 0 # completed after view
noview_comp = 0 # completed without view
view_comp_offer_list = [] # keep track of offer id
view_comp_offer_type_list = [] # keep track of offer type
# loop through received offers and check if each offer was completed and/or viewed
# 'start->end' is the time window in which an offer can be viewed & completed
for i in range(len(offers)):
id = offers.iloc[i]['consolidate_offer_id'] # offer id
start = offers.iloc[i]['offer_time'] # time when this offer was received
end = offers.iloc[i]['duration'] + start # end-point of this offer
# now check if this offer was viewed and/or completed
viewed = 'offer viewed' in list(person_df[(person_df.offer_time>=start)&(person_df.offer_time<=end)&\
(person_df.consolidate_offer_id==id)]['event'])
completed = 'offer completed' in list(person_df[(person_df.offer_time>=start)&(person_df.offer_time<=end)&\
(person_df.consolidate_offer_id==id)]['event'])
if completed:
comp +=1
if viewed:
view_comp +=1
view_comp_offer_list.append(id)
view_comp_offer_type_list.append(person_df[person_df.consolidate_offer_id==id]['offer_type'].iloc[0])
idx_time = person_df[(person_df.offer_time>=start)&(person_df.offer_time<=end)&(person_df.consolidate_offer_id==id)&\
(person_df.event=='offer completed')].iloc[0]['offer_time']
offer_record.iloc[idx] = [person,id,idx_time] # keep track of viewed & completed offers in this df
idx +=1
else: noview_comp +=1
receive = len(offers)
receive_bogo = list(offers.offer_type).count('bogo')
receive_discount = list(offers.offer_type).count('discount')
receive_informational = list(offers.offer_type).count('informational')
bogo = view_comp_offer_type_list.count('bogo')
discount = view_comp_offer_type_list.count('discount')
informational = view_comp_offer_type_list.count('informational')
# count how many times each type of offer was viewed & completed
# loop through 10 different offers
counts = []
for off in list(portfolio.consolidate_offer_id):
counts.append(view_comp_offer_list.count(off))
final = [receive,receive_bogo,receive_discount,receive_informational,comp,view_comp,noview_comp,bogo,discount,informational] + counts
return final,idx,offer,offer_record
def get_offer_df(transcript):
'''
This function generates/modify 'offer' dataframe containing all offer history of all users,
and 'offer_record' dataframe containing all viewed & completed offers from all users.
inputs:
transcript - events df
outputs:
1. offer - a dataframe containing all offer summaries of users; columns represent:
[offers received; completed; viewed & completed; completed without viewing; viewed & completed bogo offer;
viewed & completed discount offer; counts for each offer id (viewed and completed counts)]
2. offer_record - a dataframe containing records (offer id, time and person id) for all
viewed & completed offers
'''
# create empty offer and offer_record dataframes
offer_record = pd.DataFrame(columns=['person','id','time'],index=range(len(transcript)))
offer = pd.DataFrame(columns = ['receive','rec_bogo','rec_discount','rec_informational','comp','view_comp',
'noview_comp','bogo','discount','informational',
'ae264e3637204a6fb9bb56bc8210ddfd','4d5c57ea9a6940dd891ad53e9dbe8da0',
'3f207df678b143eea3cee63160fa8bed','9b98b8c7a33c4b65b9aebfe6a799e6d9',
'0b1e1539f2cc45b7b9fa7c272da2e1d7','2298d6c36e964ae4a3e7e9706d1fb8c2',
'fafdcd668e3743c1bb461111dcafc2a4','5a8bc65990b245e5a138643cd4eb9837',
'f19421c1d4aa40978ebb69ca19b0e20d','2906b810c7d4411798c6938adc9daaa5']
,index=list(transcript.person.unique()))
persons = list(transcript.person.unique())
idx = 0
# loop through all users
for i in trange(len(persons)):
person=persons[i]
person_df = transcript[transcript.person==person]
# use above function to parse offers of a user, and save the result in offer df
final, idx, offer, offer_record = offer_analyzer(person_df,person,idx,offer,offer_record)
offer.loc[person] = final
offer = offer.reset_index()
return offer, offer_record
# +
# run function and get the dataframes modified!
offer, offer_record = get_offer_df(person_offer_demographic)
# take a look at the offer df
offer.sample(5)
# -
#Save and Reuse
#offer.to_csv('data/offer.csv')
#offer_record.to_csv('data/offer_record.csv')
offer=pd.read_csv('data/offer.csv')
offer_record=pd.read_csv('data/offer_record.csv')
del offer['Unnamed: 0']
del offer_record['Unnamed: 0']
offer.rename(columns={'index':'person','bogo':'vc_bogo','discount':'vc_discount','informational':'vc_informational'}, inplace=True)
offer.head(10)
# Below, we attempt to see whether there are offers completed without being received.
#
# Actually, we don't need to consider time comparison for this time, since if the offer is marked "completed", it **must** be completed in the valid time given!
def offer_analyzer_norec_comp(person_df,person,idx,offer,offer_record):
'''
This function takes in a person's events from transcript df and generate a df that includes the number and type
of offer the person completed, received, received and completed as well as not received but completed. The function
also generates a df of all not received but completed offers by all users for future use.
inputs:
1. person_df - all events of this person
2. person - the person's id
3. idx - index to keep track of for generating 'not received & completed offer dataframe'
4. offer - empty offer df
5. offer_record - empty offer_record df(if records not received&completed offers)
outputs:
1. [final] - a list of all output variables including:
receive -- counts of offer received
comp -- counts of offer completed
view_comp -- counts of offer viewed and completed
noview_comp -- counts of offer completed without viewing
bogo -- counts of viewed & completed bogo offer
discount -- counts of viewed & completed discount offer
informational -- counts of viewed & completed informational offer and
10 columns for 10 types of offers-- how many times this specific offer was viewed & completed
2. idx - updated index to keep track of for generating 'viewed & completed offer dataframe'
'''
# select all offers the person received
comp_offers = person_df[person_df.event=='offer completed']
# start counting
rec_comp = 0 # received&completed offers
norec_comp = 0 # completed without received
norec_comp_offer_list = [] # keep track of offer id
norec_comp_offer_type_list = [] # keep track of offer type
# loop through received offers and check if each offer was received/not recieved
for i in range(len(comp_offers)):
id = comp_offers.iloc[i]['consolidate_offer_id'] # offer id
comp_time = comp_offers.iloc[i]['offer_time'] # time when this offer was completed
# now check if this offer was received/not recieved
received = 'offer received' in list(person_df[(person_df.consolidate_offer_id==id)]['event'])
if received:
rec_comp +=1
else:
norec_comp +=1
norec_comp_offer_list.append(id)
norec_comp_offer_list.append(person_df[person_df.consolidate_offer_id==id]['offer_type'].iloc[0])
idx_time = person_df[(person_df.consolidate_offer_id==id)&\
(person_df.event=='offer completed')].iloc[0]['offer_time']
offer_record.iloc[idx] = [person,id,idx_time] # keep track of noreceived & completed offers in this df
idx +=1
complete = len(comp_offers)
complete_bogo = list(comp_offers.offer_type).count('bogo')
complete_discount = list(comp_offers.offer_type).count('discount')
complete_informational = list(comp_offers.offer_type).count('informational')
norec_comp_bogo = norec_comp_offer_type_list.count('bogo')
norec_comp_discount = norec_comp_offer_type_list.count('discount')
norec_comp_informational = norec_comp_offer_type_list.count('informational')
# count how many times each type of offer was viewed & completed
# loop through 10 different offers
counts = []
for off in list(portfolio.consolidate_offer_id):
counts.append(norec_comp_offer_list.count(off))
final = [complete,complete_bogo,complete_discount,complete_informational,rec_comp,norec_comp,norec_comp_bogo,norec_comp_discount,norec_comp_informational] + counts
return final,idx,offer,offer_record
def get_offer_df_norec_comp(transcript):
'''
This function generates/modify 'offer' dataframe containing all offer history of all users,
and 'offer_record' dataframe containing all noreceived & completed offers from all users.
inputs:
transcript - events df
outputs:
1. offer - a dataframe containing all offer summaries of users; columns represent:
[offers received; completed; viewed & completed; completed without viewing; viewed & completed bogo offer;
viewed & completed discount offer; counts for each offer id (viewed and completed counts)]
2. offer_record - a dataframe containing records (offer id, time and person id) for all
viewed & completed offers
'''
# create empty offer and offer_record dataframes
offer_record = pd.DataFrame(columns=['person','id','time'],index=range(len(transcript)))
offer = pd.DataFrame(columns = ['complete','complete_bogo','complete_discount','complete_informational','rec_comp','norec_comp',
'norec_comp_bogo','norec_comp_discount','norec_comp_informational',
'ae264e3637204a6fb9bb56bc8210ddfd','4d5c57ea9a6940dd891ad53e9dbe8da0',
'3f207df678b143eea3cee63160fa8bed','9b98b8c7a33c4b65b9aebfe6a799e6d9',
'0b1e1539f2cc45b7b9fa7c272da2e1d7','2298d6c36e964ae4a3e7e9706d1fb8c2',
'fafdcd668e3743c1bb461111dcafc2a4','5a8bc65990b245e5a138643cd4eb9837',
'f19421c1d4aa40978ebb69ca19b0e20d','2906b810c7d4411798c6938adc9daaa5']
,index=list(transcript.person.unique()))
persons = list(transcript.person.unique())
idx = 0
# loop through all users
for i in trange(len(persons)):
person=persons[i]
person_df = transcript[transcript.person==person]
# use above function to parse offers of a user, and save the result in offer df
final, idx, offer, offer_record = offer_analyzer_norec_comp(person_df,person,idx,offer,offer_record)
offer.loc[person] = final
offer = offer.reset_index()
return offer, offer_record
# +
from tqdm import trange
# run function and get the dataframes modified!
offer_norec_comp, offer_record_norec_comp = get_offer_df_norec_comp(person_offer_demographic)
# take a look at the offer df
offer_norec_comp.sample(5)
# -
offer_norec_comp.to_csv('data/offer_norec_comp.csv')
offer_record_norec_comp.to_csv('data/offer_record_norec_comp.csv')
offer_norec_comp=pd.read_csv('data/offer_norec_comp.csv')
offer_record_norec_comp=pd.read_csv('data/offer_record_norec_comp.csv')
del offer_norec_comp['Unnamed: 0']
offer_record_norec_comp.isnull().mean()
offer_norec_comp['norec_comp'].mean()
# **Comments:** It appears that no such offers(completed&noreceived) are in our records.
[1,4,5]+[1,2,3]
print(offer_record.shape)
offer_record.isnull().mean()#less than 0.17 in the offer received, offer viewed and offer completed records are viewed&completed
offer[offer['person']=='e1e614f30e9c45478d1c5aa8fe3c6dbb']
offer.shape
offer.head()
#viewed&completed offer record
offer_record.head()
# **Comments:** now the df "offer" can show clearly for each person,
# 1.how many offers received, within which how many received bogo&discount&informational;
# 2.how many offers completed, within which how many are viewed before completed and how many are completed before viewed;
# 3.within view&completed, how many are bogo&discount&informational;
# 4.within view&completed, the counts of each offer in the 10 choices.
#
# We then conduct data engineering on transaction information.
person_offer_demographic['offer_time'].sample()
transaction['transaction_time'].sample()
offer_record['time'][:100].sample()
# **Core logic of codes below:** If an offer is completed, then the transaction time should be the same as that of completed offer.
# Hence we first extract the transaction time, then see if any completed offer time of this person matches ->transaction associated with viewed and completed offers;
# Else->transaction associated with completed but not viewed offers
from tqdm import trange
# +
#Loop through the unique persons one by one in the transaction data. Each time:
#1.Extract all transaction records for this person(using transaction['person id']==person id)
#2.Sum the transaction amount of this person and mark as "total"
#3.Loop through the transaction records: For every loop, store the transaction time and amount; find in the transcript(non-transaction
#records) satisfying "offer completed" and the same time as transaction time; Judge whether the offer(s) falls in the viewed&completed list;
#if YES, add the transaction amount to viewed&completed transaction amount; if NO, add it to noviewed&completed transaction amount;
# -
def transaction_calculator(trans_ori,person,person_df,offer_record,transaction):
'''
This function takes in a person's events and calculate the total transaction, transaction associated
with viewed & completed offers and transaction associated with not viewed but completed offers.
inputs:
1. trans_ori - transaction dataframe
2. person - person id
3. person_df - all events of the person
4. offer_record - record of all viewed & completed offers, will be used to assess if a certain amount of
transaction is associated with viewed & completed offer
outputs:
1. [final] - a list including all output variables:
total - total transaction made by this person
view_comp - transaction associated with viewed and completed offers
noview_comp - transaction associated with completed but not viewed offers
'''
# calculate total transaction made by this person
trans = trans_ori[trans_ori['person']==person][['person','transaction_time','amount']] # all transactions
total = trans['amount'].sum()
# start calculating
view_comp = 0
noview_comp = 0
view = False
comp = False
# loop through transactions to see if they are associated with viewed & completed offer
for i in range(len(trans)):
time = trans.iloc[i]['transaction_time'] # time of this transaction
amount = trans.iloc[i]['amount'] # amount of this transaction
# check if there's any completed offer(s) at this transaction time
comp_off = person_df[(person_df.offer_time==time)&(person_df.event=='offer completed')] # completed offers df
if len(comp_off) > 0:
comp = True
# check if the completed offer was viewed
# if more than one offers were completed simultaneously, check if ANY of them were viewed
for j in range(len(comp_off)):
if ((offer_record.person==comp_off.iloc[j]['person'])&(offer_record.id==comp_off.iloc[j]['consolidate_offer_id'])&
(offer_record.time==comp_off.iloc[j]['offer_time'])).any(): #offer record is a df of viewed&completed offers
view = True
# update transactions for viewed & completed offers as well as not viewed but completed offers
if comp and view:
view_comp += amount
else:
noview_comp += amount
# reset the value for next transaction
view = False
comp = False
final = [total,view_comp,noview_comp]
return final,transaction
def get_transaction_df(trans_ori,transcript):
'''
This function generates a transaction dataframe containing all purchase behavior of all users.
inputs:
1. trans_ori - transaction dataframe
2. transcript - events df
3. transaction - empty transaction df
outputs:
1. transaction_df - a dataframe contains all purchase behavior of all users including total transaction
amount, transaction associated with viewed & compelted offers as well as transaction associated with
not viewed but completed offers
'''
# create an empty transaction dataframe
transaction = pd.DataFrame(columns = ['total','view_complete_tran','noview_complete_tran'], index=list(transcript.person.unique()))
# loop through all users in transcript (which is also the overall users in profile)
persons = list(transcript.person.unique())
for i in trange(len(persons)):
person=persons[i]
person_df = transcript[transcript.person==person]
# use above function to parse transaction and save the result in transaction df
final,transaction = transaction_calculator(trans_ori,person,person_df,offer_record,transaction)
transaction.loc[person] = final
transaction = transaction.reset_index()
return transaction
# +
# run the function and get the transaction df modified!
transaction_gen = get_transaction_df(transaction,person_offer_demographic)
# take a look at the transaction df
transaction_gen.sample(5)
# -
transaction_gen.to_csv('data/transaction_gen.csv')
transaction_gen=pd.read_csv('data/transaction_gen.csv')
del transaction_gen['Unnamed: 0']
transaction_gen.head()
# ### 5. Customer Behavior Analysis
# #### 5.1 Customer Response Demographic Analysis
# Combine customer information&offer&transaction altogether:
import datetime
person_all_information=transaction_gen.merge(profile,on='person')
person_all_information=person_all_information.merge(offer,on='person')
person_all_information['became_member_on'] = person_all_information.became_member_on.apply(lambda x: datetime.datetime.strptime(str(x),'%Y%m%d').date())
person_all_information.became_member_on=person_all_information.became_member_on.apply(lambda x: x.toordinal())
person_all_information.to_csv('data/person_all_information.csv')
person_all_information=pd.read_csv('data/person_all_information.csv')
del person_all_information['Unnamed: 0']
person_all_information.columns
person_all_information=person_all_information[['person','age','gender','income','became_member_on','total','view_complete_tran','noview_complete_tran','receive','rec_bogo','rec_discount','rec_informational','comp','view_comp',
'noview_comp','vc_bogo','vc_discount','vc_informational',
'ae264e3637204a6fb9bb56bc8210ddfd','4d5c57ea9a6940dd891ad53e9dbe8da0',
'3f207df678b143eea3cee63160fa8bed','9b98b8c7a33c4b65b9aebfe6a799e6d9',
'0b1e1539f2cc45b7b9fa7c272da2e1d7','2298d6c36e964ae4a3e7e9706d1fb8c2',
'fafdcd668e3743c1bb461111dcafc2a4','5a8bc65990b245e5a138643cd4eb9837',
'f19421c1d4aa40978ebb69ca19b0e20d','2906b810c7d4411798c6938adc9daaa5']]
person_all_information.head()
person_all_information['responded'] = person_all_information.view_comp.apply(lambda x: 'T' if x!=0 else 'F')
person_all_information['gender']=person_all_information.gender.apply(lambda x: 1 if x=='M' else (0 if x=='F' else 2))
import warnings
warnings.filterwarnings('ignore') # turn off warning on missing values
sns.pairplot(person_all_information[['age','gender','income','became_member_on','view_complete_tran','responded']].fillna(0),hue='responded',hue_order=['T','F'],plot_kws=dict(alpha=0.2),dropna=True);
# **Comments:** From the above graph, we can clearly see that the response has positive relations with age, income and became_member_on(concluded from the last row of the graph).
#
# Also, we can see there are orange lines in the 3 plots(age, income, became_member_on) of the last row. Those customers might be those who newly registered the mobile app and have low income.
#
# From the gender-gender plot, we can see that female tend to respond to offer compared with male and those input "Other" or did not input.
#
# Furthermore, those who didn't fully provide their personal information tend not to respond to the offer.
portfolio
person_and_offer.head()
person_all_information.head()
# #### 5.2 Analysis Based on Offer Information
# Next, we make analysis based on the offer information, e.g. offer type, channel, etc.
# **Offer Type**
complete_rate=person_all_information.comp.sum()/person_all_information.receive.sum()
complete_rate
bogo_complete_rate=offer_norec_comp.complete_bogo.sum()/person_all_information.rec_bogo.sum()
bogo_complete_rate
discount_complete_rate=offer_norec_comp.complete_discount.sum()/person_all_information.rec_discount.sum()
discount_complete_rate
informational_complete_rate=offer_norec_comp.complete_informational.sum()/person_all_information.rec_informational.sum()
informational_complete_rate
vc_rate=person_all_information.view_comp.sum()/person_all_information.receive.sum()
vc_rate
bogo_vc_rate=person_all_information.vc_bogo.sum()/person_all_information.rec_bogo.sum()
bogo_vc_rate
discount_vc_rate=person_all_information.vc_discount.sum()/person_all_information.rec_bogo.sum()
discount_vc_rate
informational_vc_rate=person_all_information.vc_informational.sum()/person_all_information.rec_bogo.sum()
informational_vc_rate
# From above we can see that the complete rate is about 44%, and it appears that discount offer is more preferrable than bogo and informational. Especially, for informational offer, the complete rate is 0, which shows that this type of offer might be simply providing information and hardlly inspires people to buy the products.
# **Channel**
email_rec=person_and_offer[person_and_offer.event=='offer received'].channel_email.sum()
email_comp=person_and_offer[person_and_offer.event=='offer completed'].channel_email.sum()
email_comp/email_rec
mobile_rec=person_and_offer[person_and_offer.event=='offer received'].channel_mobile.sum()
mobile_comp=person_and_offer[person_and_offer.event=='offer completed'].channel_mobile.sum()
mobile_comp/mobile_rec
social_rec=person_and_offer[person_and_offer.event=='offer received'].channel_social.sum()
social_comp=person_and_offer[person_and_offer.event=='offer completed'].channel_social.sum()
social_comp/social_rec
web_rec=person_and_offer[person_and_offer.event=='offer received'].channel_web.sum()
web_comp=person_and_offer[person_and_offer.event=='offer completed'].channel_web.sum()
web_comp/web_rec
# From above, it appears that web might be the most efficient channel, which contributing the highest complete rate.
# ## III. Data Modeling
# In this part, we are going to build a machine learning model that predicts whether or not someone will respond to an offer.
# From above analysis, we know that age, gender, income, membership date, offer type can affect whether a customer respond to an offer. Hence, we make them as explanatory variable and make 'responded' as binary response variable.
#Import all useful packages
import datetime
#from sklearn.pipeline import Pipeline
from sklearn.preprocessing import MinMaxScaler,StandardScaler
from sklearn.model_selection import train_test_split,GridSearchCV
from sklearn.svm import SVC
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.ensemble import AdaBoostClassifier
from sklearn.metrics import accuracy_score,f1_score
#make the time data available to extract year&month
profile['became_member_on'] = profile.became_member_on.apply(lambda x: datetime.datetime.strptime(str(x),'%Y%m%d').date())
# Note that, below we are going to mark viewed&completed as responded instead of mark "completed" as responded, because the latter is actually a random behavior without **knowing** he/she has such an offer.
def transform_and_tts(profile,offer):
'''
This function takes in profile and offer dataframes and returns training and test datasets for ML.
inputs:
1. profile - profile dataframe
2. offer - offer dataframe
outputs:
1. X_train, X_test, y_train, y_test - input data for training and test, target label for training and test
2. age_interval, income_interval - interval index for age and income variables
'''
# transform features and label
prof = profile.copy()
prof = prof[prof.person.isin(list(offer[offer.receive==0]['person']))==False] # exclude people never received an offer
prof['member_year'] = prof.became_member_on.apply(lambda x: x.year)
prof['member_month'] = prof.became_member_on.apply(lambda x: x.month)
prof.drop('became_member_on',axis=1,inplace=True)
# create 'offer' and 'label' columns: offer col has two values (bogo or discount) and label col shows whether
# the user responded to the offer or not
bogo = offer[offer.rec_bogo!=0][['person','vc_bogo']]
bogo['label'] = bogo.vc_bogo.apply(lambda x: 0 if x==0 else 1)
bogo.drop('vc_bogo',axis=1,inplace=True)
bogo = prof.merge(bogo,on='person').drop('person',axis=1)
bogo['offer'] = 'bogo'
discount = offer[offer.rec_discount!=0][['person','vc_discount']]
discount['label'] = discount.vc_discount.apply(lambda x: 0 if x==0 else 1)
discount.drop('vc_discount',axis=1,inplace=True)
discount = prof.merge(discount,on='person').drop('person',axis=1)
discount['offer'] = 'discount'
# concat bogo and discount df
df = pd.concat([bogo,discount])
df.age.replace(118,np.NaN,inplace=True)
df=df.dropna()#drop nan for convenience
# creat dummy variables
df = pd.get_dummies(df,columns=['gender','member_year','member_month','offer'],dummy_na=True)
# assign X and y
X = df.drop('label',axis=1)
y = df['label']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42)
return X_train, X_test, y_train, y_test
X_train, X_test, y_train, y_test = transform_and_tts(profile,offer)
X_train.head()
y_train.value_counts()
y_test.value_counts()
# From above, we can see that the labels are **imbalanced**, approximately 2:1 for Positive vs. Negative labels. Hence, we should use F1_score instead of accuracy, since F1_score is a metric to balance recall&precision and to deal with imbalanced labels.
# \begin{equation*}
# F1=2\times \frac{Precision\times Recall}{Precision+Recall}
# \end{equation*}
# \begin{equation*}
# Precision=\frac{True Positive}{True Positive+False Positive}
# \end{equation*}
# \begin{equation*}
# Recall=\frac{True Positive}{True Positive+False Negative}
# \end{equation*}
# +
# Let's check out which classifier will work best in our case
classifiers = [SVC(),
DecisionTreeClassifier(),
RandomForestClassifier(),
GaussianNB(),
AdaBoostClassifier()]
X_train, X_test, y_train, y_test = transform_and_tts(profile,offer)
#Scale the X_train & X_test to [0,1]
#scaler=MinMaxScaler()
#X_train_scaled = scaler.fit_transform(X_train)
#X_test_scaled = scaler.fit_transform(X_test)
performance = pd.DataFrame(columns=["Classifier", "F1_Score"])
acc_dict = {}
for clf in classifiers:
name = clf.__class__.__name__
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
acc = f1_score(y_test, y_pred)
if name not in acc_dict:
acc_dict[name] = acc
else:
acc_dict[name] += acc
for clf in acc_dict:
performance_record = pd.DataFrame([[clf, acc_dict[clf]]], columns=["Classifier", "F1_Score"])
performance = performance.append(performance_record)
plt.xlabel('F1_Score')
plt.title('Classifier F1_Score')
sns.barplot(x='F1_Score', y='Classifier', data=performance);
# -
# It appears that AdaBoost is the best classifier. Therefore, we conduct GridSearch on it to find the best parameters:
# +
param_grid = {"base_estimator__criterion" : ["gini", "entropy"],
"base_estimator__splitter" : ["best", "random"],
"n_estimators": [5, 10, 20,50],
"learning_rate":[0.001, 0.01, 0.1, 1],
'base_estimator__max_depth':[1,2,3,4]
}
DTC = DecisionTreeClassifier(random_state=42)
ADA = AdaBoostClassifier(base_estimator = DTC)
grid_ada = GridSearchCV(estimator=ADA,param_grid=param_grid,scoring='f1',cv=5)
grid_ada.fit(X_train,y_train)
print('Training F1_score is:', grid_ada.score(X_train,y_train))
print('Test F1_score is:', grid_ada.score(X_test,y_test))
# -
from sklearn.externals import joblib
joblib.dump(grid_ada.best_estimator_, 'filename4.pkl')
from sklearn.externals import joblib
model=joblib.load('filename4.pkl')
model
# Next, we want to make a function such that we take in customer info, offer type, etc, then we can transform all these information into format which can be taken by the classifier like the test data, then predict whether the customer respond or not based on the raw profile&offer information:
X_test.columns
def predict_engine(model,customer,offer_type):
'''
This function takes in a customers info and offer type and transform it to the same format as test data, then returns a
predicted result(respond or not).
inputs:
1. model - the best classifier
2. customer - customer's info, the same format as original profile df
3. offer_type - 'bogo' or 'discount'
outputs:
prediction - whether the customer would respond to given offer or not
'''
# First, let's check whether the customer provided demographic info or not
if customer['age']==118 or customer.isnull().any():
flag = False
else:
flag = True
# Creat new customer df and tranform datetime col
cols = transform_and_tts(profile,offer)[1].columns#takes the X_test
customer_df = pd.DataFrame(columns=cols,index=[0])
year = customer['became_member_on'].year
month = customer['became_member_on'].month
customer_df['member_year_' + str(str(float(year)))] = 1
customer_df['member_month_' + str(str(float(month)))] = 1
if flag:
# transform profile info if provided
gender = customer['gender']
customer_df['gender_' + str(gender)] = 1
customer_df['age']=customer['age']
customer_df['income']=customer['income']
customer_df['offer_' + offer_type] = 1
else:
customer_df['gender_nan'] = 1
customer_df['offer_' + offer_type] = 1
customer_df.fillna(0,inplace=True)
pred = model.predict(customer_df)[0]
if pred==0:
print('Not respond!')
else:
print('Respond!')
return customer_df
predict_engine(model, profile.sample(1).iloc[0], 'bogo')
predict_engine(model, profile.sample(1).iloc[0], 'discount')
# ## IV. Evaluation of Results
# To conclude, in this notebook, we:
# - Cleanse the offer data such that we can separate the completed offer data into A.viewed&completed offer and B.noviewed&completed offer. The reason to do this is that even though the customers made the transaction and completed the offer, he/she may not be aware of the offer. In other words, his/her action may not be offer-oriented, hence should not be counted as **responded** to the offer;
# - Cleanse the offer data such that we can separate the completed offer data into A.received&completed offer and B.noreceived&completed offer to see whether there is anyone who completed the offer without actually receiving the offer. However, it appears that there is no record for **noreceived&completed** offer in our case/data;
# - Compare the transaction time and the offer completed time in order to determine whether the transaction is associated with a completed offer(more specifically, viewed&completed offer or noviewed&completed offer or other offer), because in the raw data, there are only person, amounts and time given, without telling us whether it is related to any offer;
# - Next, we have demographic, offer type and channel analysis on the complete rate;
# - Finally, we build a machine learning model to enable the prediction of the customer's response given a customer with age, gender, income, offer type and other information.
# Possible Enhancement in the Future:
# - For convenience, I drop all the NaNs; in the future, other skills can be used, such as using mean, median, etc; or just simply keep it as a value, since in the future, there will still be part of customers who won't fill in their information;
# - This is a classification model; Alternatively, regression model can be built to predict how much someone will spend(i.e. transaction) based on demographics and offer type;
# - A web app can be built such that when inputting the customer information, the prediction of response/transaction amount can be output.
| Starbucks_Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Support Vector Machines
# In the first part of this exercise, you will build support vector machines
# (SVMs) for solving binary classification problems. You will experiment with your
# classifier on three example 2D datasets. Experimenting with these datasets
# will help you gain intuition into how SVMs work and how to use a Gaussian
# kernel with SVMs.
# +
import random
import numpy as np
import matplotlib.pyplot as plt
# This is a bit of magic to make matplotlib figures appear inline in the notebook
# rather than in a new window.
# %matplotlib inline
plt.rcParams['figure.figsize'] = (10.0, 8.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# Some more magic so that the notebook will reload external python modules;
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
# %load_ext autoreload
# %autoreload 2
# -
# ## Data set 1
# We will begin with a 2D example dataset which can be separated by a
# linear boundary. In
# this dataset, the positions of the positive examples (green circles) and the
# negative examples (indicated with red circles) suggest a natural separation indicated
# by the gap. However, notice that there is an outlier positive example on
# the far left at about (0.1, 4.1). As part of this exercise, you will also see how
# this outlier affects the SVM decision boundary.
# +
import linear_svm
import utils
from sklearn import preprocessing, metrics
from linear_classifier import LinearSVM_twoclass
############################################################################
# Part 0: Loading and Visualizing Data #
# We start the exercise by first loading and visualizing the dataset. #
# The following code will load the dataset into your environment and plot #
# the data. #
############################################################################
# load ex4data1.mat
X,y = utils.load_mat('data/ex4data1.mat')
utils.plot_twoclass_data(X,y,'x1', 'x2',['neg','pos'])
plt.show()
# -
# ## The hinge loss function and gradient
# Now you will implement the hinge loss cost function and its gradient for support vector machines.
# Complete the **binary\_svm\_loss** function in **linear\_svm.py** to return the cost and gradient for the hinge loss function.
# Recall that the hinge loss function is
# $$ J(\theta) = \frac{1}{2m} \sum_{j=0}^{d} {\theta_j}^2 + \frac{C}{m} \sum_{i=1}^{m} max(0, 1 -y^{(i)}h_\theta(x^{(i)})) $$
#
# where $h_{\theta}(x) = \theta^ T x$ with $x_0 = 1$. $C$ is the penalty factor which measures how much misclassifications are penalized. If $y^{(i)}h_\theta(x^{(i)})) > 1$, then $x^{(i)}$ is correctly classified and the loss associated with that example is zero. If $y^{(i)}h_\theta(x^{(i)})) < 1$, then $x^{(i)}$ is not within the appropriate margin (positive or negative) and the loss associated with that example is greater than zero. The gradient of the hinge loss
# function is a vector of the same length as $\theta$ where the $j^{th}$ element, $j=0,1,\ldots,d$ is defined as follows:
#
# \begin{eqnarray*} \frac{\partial J(\theta)}{\partial \theta_j} & = &
# \left \{
# \begin{array}{l l}
# \frac{1}{m} \theta_j + \frac{C}{m} \sum_{i=1}^{m} -y^{(i)}x_j^{(i)}& \mbox{ if } y^{(i)}h_\theta(x^{(i)})) < 1\\
# \frac{1}{m} \theta_j & \mbox{ if } y^{(i)}h_\theta(x^{(i)})) >= 1\\
# \end{array} \right.
# \end{eqnarray*}
#
# Once you are done, the cell below will call your **binary\_svm\_loss** function with a zero vector $\theta$.
# You should see that the cost $J$ is 1.0. The gradient of the loss function with respect to an all-zeros $\theta$ vector is also computed and should be $[-0.12956186 -0.00167647]^T$.
# +
############################################################################
# Part 1: Hinge loss function and gradient #
############################################################################
C = 1
theta = np.zeros((X.shape[1],))
yy = np.ones(y.shape)
yy[y==0] = -1
J,grad = linear_svm.binary_svm_loss(theta,X,yy,C)
print("J = %.4f grad = %s" %(J,grad))
# -
# ## Impact of varying C
# In this part of the exercise, you will try using different values of the C
# parameter with SVMs. Informally, the C parameter is a positive value that
# controls the penalty for misclassified training examples. A large C parameter
# tells the SVM to try to classify all the examples correctly. C plays a role
# similar to $\frac{1}{\lambda}$, where $\lambda$ is the regularization parameter that we were using
# previously for logistic regression.
#
# The SVM training function is in **linear\_classifier.py** -- this is a gradient descent algorithm that uses your loss and gradient functions.
# The cell below will train an SVM on the example data set 1 with C = 1. It first scales the data to have zero mean and unit variance, and adds the intercept term to the data matrix.
# When C = 1, you should find that the SVM puts the decision boundary in
# the gap between the two datasets and misclassifies the data point on the far
# left.
#
# Your task is to try different values of C on this dataset. Specifically, you
# should change the value of C in the cell below to C = 100 and run the SVM
# training again. When C = 100, you should find that the SVM now classifies
# every single example correctly, but has a decision boundary that does not
# appear to be a natural fit for the data. Include a plot of the decision boundary for C = 100 in writeup.pdf.
# +
############################################################################
# Scale the data and set up the SVM training #
############################################################################
# scale the data
scaler = preprocessing.StandardScaler().fit(X)
scaleX = scaler.transform(X)
# add an intercept term and convert y values from [0,1] to [-1,1]
XX = np.array([(1,x1,x2) for (x1,x2) in scaleX])
yy = np.ones(y.shape)
yy[y == 0] = -1
yy[y == 0] = -1
############################################################################
# Part 2: Training linear SVM #
# We train a linear SVM on the data set and the plot the learned #
# decision boundary #
############################################################################
############################################################################
# You will change this line below to vary C. #
############################################################################
C = 100
svm = LinearSVM_twoclass()
svm.theta = np.zeros((XX.shape[1],))
svm.train(XX,yy,learning_rate=1e-4,reg=C,num_iters=50000,verbose=True,batch_size=XX.shape[0])
# classify the training data
y_pred = svm.predict(XX)
print("Accuracy on training data = %.3f" %metrics.accuracy_score(yy,y_pred))
# visualize the decision boundary
utils.plot_decision_boundary(scaleX,y,svm,'x1','x2',['neg','pos'])
# -
# ## SVMs with Gaussian kernels
# In this part of the exercise, you will be using SVMs to do non-linear classification.
# In particular, you will be using SVMs with Gaussian kernels on
# datasets that are not linearly separable.
#
# To find non-linear decision boundaries with the SVM, we need to first implement
# a Gaussian kernel. You can think of the Gaussian kernel as a similarity
# function that measures the distance between a pair of examples,
# $(x^{(i)}, x^{(j)})$. The Gaussian kernel is also parameterized by a bandwidth parameter,
# $\sigma$, which determines how fast the similarity metric decreases (to 0)
# as the examples are further apart.
# You should now complete the function **gaussian\_kernel** in **utils.py** to compute
# the Gaussian kernel between two examples. The Gaussian kernel
# function is defined as:
#
# $$ k(x^{(i)},x^{(j)}) = exp\left(- \frac{{||x^{(i)}-x^{(j)}||}^2}{2\sigma^2}\right) $$
#
# When you have completed the function, the cell below
# will test your kernel function on two provided examples and you should expect
# to see a value of 0.324652.
# +
############################################################################
# Part 3: Training SVM with a kernel #
# We train an SVM with an RBF kernel on the data set and the plot the #
# learned decision boundary #
############################################################################
# test your Gaussian kernel implementation
x1 = np.array([1,2,1])
x2 = np.array([0,4,-1])
sigma = 2
print("Gaussian kernel value (should be around 0.324652) = %.5f" %utils.gaussian_kernel(x1,x2,sigma))
# -
# ## SVMs with Gaussian kernels on Dataset 2
# The next cell will load and plot dataset 2. From
# the plot, you can observe that there is no linear decision boundary that
# separates the positive and negative examples for this dataset. However, by
# using the Gaussian kernel with the SVM, you will be able to learn a non-linear
# decision boundary that can perform reasonably well for the dataset.
# If you have correctly implemented the Gaussian kernel function, the cell below
# will proceed to train the SVM with the Gaussian kernel on this dataset.
#
# The decision boundary found by the SVM with C = 1 and a Gaussian
# kernel with $\sigma = 0.01$ will be plotted. The decision boundary is able to separate most of the positive and
# negative examples correctly and follows the contours of the dataset well.
#
# +
# load ex4data2.mat
X,y = utils.load_mat('data/ex4data2.mat')
# visualize the data
utils.plot_twoclass_data(X,y,'', '',['neg','pos'])
# convert X to kernel form with the kernel function
sigma = 0.02
# compute the kernel (slow!)
K = np.array([utils.gaussian_kernel(x1,x2,sigma) for x1 in X for x2 in X]).reshape(X.shape[0],X.shape[0])
# scale the kernelized data matrix
scaler = preprocessing.StandardScaler().fit(K)
scaleK = scaler.transform(K)
# add the intercept term
KK = np.vstack([np.ones((scaleK.shape[0],)),scaleK.T]).T
# transform y from [0,1] to [-1,1]
yy = np.ones(y.shape)
yy[y == 0] = -1
# set up the SVM and learn the parameters
svm = LinearSVM_twoclass()
svm.theta = np.zeros((KK.shape[1],))
C = 1
svm.train(KK,yy,learning_rate=1e-4,reg=C,num_iters=20000,verbose=True,batch_size=KK.shape[0])
# visualize the boundary
utils.plot_decision_kernel_boundary(X,y,scaler,sigma,svm,'','',['neg','pos'])
# -
# ## Selecting hyperparameters for SVMs with Gaussian kernels
# In this part of the exercise, you will gain more practical skills on how to use
# a SVM with a Gaussian kernel. The next cell will load and display
# a third dataset.
# In the provided dataset, **ex4data3.mat**, you are given the variables **X**,
# **y**, **Xval**, **yval**. You will be using the SVM with the Gaussian
# kernel with this dataset. Your task is to use the validation set **Xval**, **yval** to determine the
# best C and $\sigma$ parameter to use. You should write any additional code necessary
# to help you search over the parameters C and $\sigma$. For both C and $\sigma$, we
# suggest trying values in multiplicative steps (e.g., 0.01, 0.03, 0.1, 0.3, 1, 3, 10, 30).
# Note that you should try all possible pairs of values for C and $\sigma$ (e.g., C = 0.3
# and $\sigma$ = 0.1). For example, if you try each of the 8 values listed above for C
# and for $\sigma$, you would end up training and evaluating (on the validation
# set) a total of $8^2 = 64$ different models.
#
# When selecting the
# best C and $\sigma$ parameter to use, you train on {\tt X,y} with a given C and $\sigma$, and then evaluate the error of the model on the
# validation set. Recall that for classification, the error is defined as the
# fraction of the validation examples that were classified incorrectly.
# You can use the **predict** method of the SVM classifier to generate the predictions for the
# validation set.
#
# After you have determined the best C and $\sigma$ parameters to use, you
# should replace the assignments to **best\_C** and **best\_sigma** in the cell below.
# +
############################################################################
# Part 4: Training SVM with a kernel #
# We train an SVM with an RBF kernel on the data set and the plot the #
# learned decision boundary #
############################################################################
# load ex4data3.mat
X,y,Xval,yval = utils.loadval_mat('data/ex4data3.mat')
# transform y and yval from [0,1] to [-1,1]
yy = np.ones(y.shape)
yy[y == 0] = -1
yyval = np.ones(yval.shape)
yyval[yval == 0] = -1
# visualize the data
utils.plot_twoclass_data(X,y,'x1', 'x2',['neg','pos'])
############################################################################
# TODO #
# select hyperparameters C and sigma for this dataset using #
# Xval and yval #
############################################################################
Cvals = [0.01,0.03,0.1,0.3,1,3,10,30]
sigma_vals = [0.01,0.03,0.1,0.3,1,3,10,30]
# TODO
# select hyperparameters here; loopover all Cvals and sigma_vals.
# About 8-10 lines of code here
max_acc = 0;
best_C = 0;
best_sigma = 0;
for sigma in sigma_vals:
K = np.array([utils.gaussian_kernel(x1,x2,sigma) for x1 in X for x2 in X]).reshape(X.shape[0],X.shape[0])
# scale the kernelized data matrix
scaler = preprocessing.StandardScaler().fit(K)
scaleK = scaler.transform(K)
# add the intercept term
KK = np.vstack([np.ones((scaleK.shape[0],)),scaleK.T]).T
Kval = np.array([utils.gaussian_kernel(x1,x2,sigma) for x1 in Xval for x2 in X]).reshape(Xval.shape[0],X.shape[0])
# scale the kernelized data matrix
scale_Kval = scaler.transform(Kval)
# add the intercept term
KK_val = np.vstack([np.ones((scale_Kval.shape[0],)),scale_Kval.T]).T
svm = LinearSVM_twoclass()
for C in Cvals:
svm.theta = np.zeros((KK.shape[1],))
svm.train(KK,yy,learning_rate=1e-4,reg=C,num_iters=20000,verbose=False,batch_size=KK.shape[0])
pred_val = svm.predict(KK_val)
accuracy = np.sum((pred_val == yyval)*1)/len(yyval)
print("current accuracy: " + str(accuracy),"max accuracy: " + str(max_acc))
if (accuracy > max_acc):
max_acc = accuracy;
best_C = C;
best_sigma = sigma;
print(max_acc,best_C,best_sigma)
############################################################################
# END OF YOUR CODE #
############################################################################
# +
# TODO: make sure you put in the best_C and best_sigma from the analysis above!
best_C = 0.3
best_sigma = 0.1
# train an SVM on (X,y) with best_C and best_sigma
best_svm = LinearSVM_twoclass()
############################################################################
# TODO: construct the Gram matrix of the data with best_sigma, scale it, add the column of ones
# Then use svm_train to train best_svm with the best_C parameter. Use 20,000 iterations and
# a learning rate of 1e-4. Use batch_size of the entire training data set.
# About 5-6 lines of code expected here.
############################################################################
K = np.array([utils.gaussian_kernel(x1,x2,best_sigma) for x1 in X for x2 in X]).reshape(X.shape[0],X.shape[0])
scaler = preprocessing.StandardScaler().fit(K)
scaleK = scaler.transform(K)
KK = np.vstack([np.ones((scaleK.shape[0],)),scaleK.T]).T
best_svm.theta = np.zeros((KK.shape[1],))
best_svm.train(KK,yy,learning_rate=1e-4,reg=best_C,num_iters=20000,verbose=False,batch_size=KK.shape[0])
############################################################################
# END OF YOUR CODE #
############################################################################
# visualize the boundary (uncomment this line after you learn the best svm)
utils.plot_decision_kernel_boundary(X,y,scaler,best_sigma,best_svm,'','',['neg','pos'])
# -
# %run -i grader
| hw4/binary_svm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: workshop
# name: workshop
# ---
# ### Function
def hello(): #'def'+'name of function'+'():'
print('Function Hello')
hello()
def print_max(a,b): #Add parameter -> 'def'+'name of function'+'(para1,para2,...):'
if a > b:
print(a, 'is maximum')
elif a == b:
print(a, 'is equil to', b)
else:
print(b, 'is maximum')
print_max(4,8)
# ### Local variable
# +
x = 50 #Global variable
def func(x):
print('Global x is', x)
x = 2
print('Change local x to', x)
func(x)
print ('x not in function still is', x)
# -
# ### Global variable
# +
x = 30 #global variable
def func_global():
global x
print ('call global x is', x)
x = 10
print('change to global x is', x)
func_global()
print('current x is',x)
# -
# ### Default value
def say(message, times=1): #define default value to time
print(message * times)
say('hello')
say('five_',5)
# ### keyword
# +
def func(a, b=3, c=5): #give b and c have default value
print('a is', a, 'and b is', b, 'and c is', c)
func(3, 8)
func(25, c=50)
func(c=1, a=9) #not relative order, we get it assign
# -
# ### VarArgs Parameters
# 除了一般變數外 (Var), 還有 *args 與 **kwargs
#*args function
def args_func(*args):
for item in args:
print('single item:', item)
args_func(1,2,3,4)
# +
#**kwargs function
def kwargs_func(**kwargs):
for key in kwargs:
print('key index is {0}; value is {1}'.format(key,kwargs[key]))
kwargs_func(a=10,b=20)
# -
#**kwargs function used item()
def kwargs_item(**kwargs):
for k, v in kwargs.items():
print('key is {0}, value is {1}'.format(k, v))
kwargs_item(c=30,d=40)
# ### Return
def maximum_return(x, y):
if x > y:
return x
elif x == y:
return 'Ther number is equal'
else:
return y
maximum_return(3,3)
# ### DocStrings
# +
def print_max(x, y):
'''Print out maxium document content.
two integer'''
x = int(x)
y = int(y)
if x > y:
print(x, 'is maximum')
else:
print(y, 'is maximum')
print_max(5,7)
print(print_max.__doc__)
# -
help(print_max)
| Function.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="mt9dL5dIir8X"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="ufPx7EiCiqgR"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="ucMoYase6URl"
# # Load images
# + [markdown] colab_type="text" id="_Wwu5SXZmEkB"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/load_data/images"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/load_data/images.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/load_data/images.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/load_data/images.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="Oxw4WahM7DU9"
# This tutorial provides a simple example of how to load an image dataset using `tf.data`.
#
# The dataset used in this example is distributed as directories of images, with one class of image per directory.
# + [markdown] colab_type="text" id="hoQQiZDB6URn"
# ## Setup
# + colab={} colab_type="code" id="3vhAMaIOBIee"
import tensorflow as tf
# + colab={} colab_type="code" id="KT6CcaqgQewg"
AUTOTUNE = tf.data.experimental.AUTOTUNE
# + colab={} colab_type="code" id="gIksPgtT8B6B"
import IPython.display as display
from PIL import Image
import numpy as np
import matplotlib.pyplot as plt
import os
# + colab={} colab_type="code" id="ZJ20R66fzktl"
tf.__version__
# + [markdown] colab_type="text" id="wO0InzL66URu"
# ### Retrieve the images
#
# Before you start any training, you will need a set of images to teach the network about the new classes you want to recognize. You can use an archive of creative-commons licensed flower photos from Google.
#
# Note: all images are licensed CC-BY, creators are listed in the `LICENSE.txt` file.
# + colab={} colab_type="code" id="rN-Pc6Zd6awg"
import pathlib
data_dir = tf.keras.utils.get_file(origin='https://storage.googleapis.com/download.tensorflow.org/example_images/flower_photos.tgz',
fname='flower_photos', untar=True)
data_dir = pathlib.Path(data_dir)
# + [markdown] colab_type="text" id="rFkFK74oO--g"
# After downloading (218MB), you should now have a copy of the flower photos available.
#
# The directory contains 5 sub-directories, one per class:
# + colab={} colab_type="code" id="QhewYCxhXQBX"
image_count = len(list(data_dir.glob('*/*.jpg')))
image_count
# + colab={} colab_type="code" id="sJ1HKKdR4A7c"
CLASS_NAMES = np.array([item.name for item in data_dir.glob('*') if item.name != "LICENSE.txt"])
CLASS_NAMES
# + [markdown] colab_type="text" id="IVxsk4OW61TY"
# Each directory contains images of that type of flower. Here are some roses:
# + colab={} colab_type="code" id="crs7ZjEp60Ot"
roses = list(data_dir.glob('roses/*'))
for image_path in roses[:3]:
display.display(Image.open(str(image_path)))
# + [markdown] colab_type="text" id="6jobDTUs8Wxu"
# ## Load using `keras.preprocessing`
# + [markdown] colab_type="text" id="ehhW308g8soJ"
# A simple way to load images is to use `tf.keras.preprocessing`.
# + colab={} colab_type="code" id="syDdF_LWVrWE"
# The 1./255 is to convert from uint8 to float32 in range [0,1].
image_generator = tf.keras.preprocessing.image.ImageDataGenerator(rescale=1./255)
# + [markdown] colab_type="text" id="lAmtzsnjDNhB"
# Define some parameters for the loader:
# + colab={} colab_type="code" id="1zf695or-Flq"
BATCH_SIZE = 32
IMG_HEIGHT = 224
IMG_WIDTH = 224
STEPS_PER_EPOCH = np.ceil(image_count/BATCH_SIZE)
# + colab={} colab_type="code" id="Pw94ajOOVrWI"
train_data_gen = image_generator.flow_from_directory(directory=str(data_dir),
batch_size=BATCH_SIZE,
shuffle=True,
target_size=(IMG_HEIGHT, IMG_WIDTH),
classes = list(CLASS_NAMES))
# + [markdown] colab_type="text" id="2ZgIZeXaDUsF"
# Inspect a batch:
# + colab={} colab_type="code" id="nLp0XVG_Vgi2"
def show_batch(image_batch, label_batch):
plt.figure(figsize=(10,10))
for n in range(25):
ax = plt.subplot(5,5,n+1)
plt.imshow(image_batch[n])
plt.title(CLASS_NAMES[label_batch[n]==1][0].title())
plt.axis('off')
# + colab={} colab_type="code" id="suh6Sjv68rY3"
image_batch, label_batch = next(train_data_gen)
show_batch(image_batch, label_batch)
# + [markdown] colab_type="text" id="AxS1cLzM8mEp"
# ## Load using `tf.data`
# + [markdown] colab_type="text" id="Ylj9fgkamgWZ"
# The above `keras.preprocessing` method is convienient, but has three downsides:
#
# 1. It's slow. See the performance section below.
# 1. It lacks fine-grained control.
# 1. It is not well integrated with the rest of TensorFlow.
# + [markdown] colab_type="text" id="IIG5CPaULegg"
# To load the files as a `tf.data.Dataset` first create a dataset of the file paths:
# + colab={} colab_type="code" id="lAkQp5uxoINu"
list_ds = tf.data.Dataset.list_files(str(data_dir/'*/*'))
# + colab={} colab_type="code" id="coORvEH-NGwc"
for f in list_ds.take(5):
print(f.numpy())
# + [markdown] colab_type="text" id="91CPfUUJ_8SZ"
# Write a short pure-tensorflow function that converts a file path to an `(img, label)` pair:
# + colab={} colab_type="code" id="arSQzIey-4D4"
def get_label(file_path):
# convert the path to a list of path components
parts = tf.strings.split(file_path, os.path.sep)
# The second to last is the class-directory
return parts[-2] == CLASS_NAMES
# + colab={} colab_type="code" id="MGlq4IP4Aktb"
def decode_img(img):
# convert the compressed string to a 3D uint8 tensor
img = tf.image.decode_jpeg(img, channels=3)
# Use `convert_image_dtype` to convert to floats in the [0,1] range.
img = tf.image.convert_image_dtype(img, tf.float32)
# resize the image to the desired size.
return tf.image.resize(img, [IMG_WIDTH, IMG_HEIGHT])
# + colab={} colab_type="code" id="-xhBRgvNqRRe"
def process_path(file_path):
label = get_label(file_path)
# load the raw data from the file as a string
img = tf.io.read_file(file_path)
img = decode_img(img)
return img, label
# + [markdown] colab_type="text" id="S9a5GpsUOBx8"
# Use `Dataset.map` to create a dataset of `image, label` pairs:
# + colab={} colab_type="code" id="3SDhbo8lOBQv"
# Set `num_parallel_calls` so multiple images are loaded/processed in parallel.
labeled_ds = list_ds.map(process_path, num_parallel_calls=AUTOTUNE)
# + colab={} colab_type="code" id="kxrl0lGdnpRz"
for image, label in labeled_ds.take(1):
print("Image shape: ", image.numpy().shape)
print("Label: ", label.numpy())
# + [markdown] colab_type="text" id="vYGCgJuR_9Qp"
# ### Basic methods for training
# + [markdown] colab_type="text" id="wwZavzgsIytz"
# To train a model with this dataset you will want the data:
#
# * To be well shuffled.
# * To be batched.
# * Batches to be available as soon as possible.
#
# These features can be easily added using the `tf.data` api.
# + colab={} colab_type="code" id="uZmZJx8ePw_5"
def prepare_for_training(ds, cache=True, shuffle_buffer_size=1000):
# This is a small dataset, only load it once, and keep it in memory.
# use `.cache(filename)` to cache preprocessing work for datasets that don't
# fit in memory.
if cache:
if isinstance(cache, str):
ds = ds.cache(cache)
else:
ds = ds.cache()
ds = ds.shuffle(buffer_size=shuffle_buffer_size)
# Repeat forever
ds = ds.repeat()
ds = ds.batch(BATCH_SIZE)
# `prefetch` lets the dataset fetch batches in the background while the model
# is training.
ds = ds.prefetch(buffer_size=AUTOTUNE)
return ds
# + colab={} colab_type="code" id="-YKnrfAeZV10"
train_ds = prepare_for_training(labeled_ds)
image_batch, label_batch = next(iter(train_ds))
# + colab={} colab_type="code" id="UN_Dnl72YNIj"
show_batch(image_batch.numpy(), label_batch.numpy())
# + [markdown] colab_type="text" id="UMVnoBcG_NlQ"
# ## Performance
#
# Note: This section just shows a couple of easy tricks that may help performance. For an in depth guide see [Input Pipeline Performance](../../guide/performance/datasets).
# + [markdown] colab_type="text" id="oNmQqgGhLWie"
# To investigate, first here's a function to check the performance of our datasets:
# + colab={} colab_type="code" id="_gFVe1rp_MYr"
import time
default_timeit_steps = 1000
def timeit(ds, steps=default_timeit_steps):
start = time.time()
it = iter(ds)
for i in range(steps):
batch = next(it)
if i%10 == 0:
print('.',end='')
print()
end = time.time()
duration = end-start
print("{} batches: {} s".format(steps, duration))
print("{:0.5f} Images/s".format(BATCH_SIZE*steps/duration))
# + [markdown] colab_type="text" id="TYiOr4vdLcNX"
# Let's compare the speed of the two data generators:
# + colab={} colab_type="code" id="85Yc-jZnVjvm"
# `keras.preprocessing`
timeit(train_data_gen)
# + colab={} colab_type="code" id="IjouTJadRxyp"
# `tf.data`
timeit(train_ds)
# + [markdown] colab_type="text" id="ZB2TjJR62BJ3"
# A large part of the performance gain comes from the use of `.cache`.
# + colab={} colab_type="code" id="Oq1V854E2Nh4"
uncached_ds = prepare_for_training(labeled_ds, cache=False)
timeit(uncached_ds)
# + [markdown] colab_type="text" id="-JCHymejWSPZ"
# If the dataset doesn't fit in memory use a cache file to maintain some of the advantages:
# + colab={} colab_type="code" id="RqHFQFwxWNbu"
filecache_ds = prepare_for_training(labeled_ds, cache="./flowers.tfcache")
timeit(filecache_ds)
| site/en/tutorials/load_data/images.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from scipy.interpolate import interpn
from constant import *
from multiprocessing import Pool
from functools import partial
import warnings
warnings.filterwarnings("ignore")
np.printoptions(precision=2)
# ### The value of renting
# Assuming we obtain the value: $\tilde{V}_{t+1}(x_{t+1})$ where:
# $x_{t+1} = [w_{t+1}, n_{t+1}, M_{t+1}, e_{t+1}, \hat{S}_{t+1}, z_{t+1}, (H)]$ from interpolation. We know $H$ and $M_t$ from the action taken and we could calculate mortgage payment $m$ and $rh$ (now treated as constant) is observed from the market.
# * Housing choice is limited: $H_{\text{choice}} = \{750, 1000, 1500, 2000\}$
# * Mortgage choice is also limitted to discrete values $M_{t} = [0.2H, 0.5H, 0.8H]$
# * State: continue to rent: $x = [w, n, e, s, z]$ switch to owning a house: $x = [w,n,M,e,s,z]$
# * Action: continue to rent: $a = (c, b, k, h)$ switch to owning a house: $a = (c, b, k, M, H)$
# * Buying house activities can only happend during the age of 20 and age of 45.
# +
#Define the utility function
def u(c):
# shift utility function to the left, so it only takes positive value
return (np.float_power(c, 1-gamma) - 1)/(1 - gamma)
#Define the bequeath function, which is a function of wealth
def uB(tb):
return B*u(tb)
#Calculate TB_rent
def calTB_rent(x):
# change input x as numpy array
# w, n, e, s, z = x
TB = x[:,0] + x[:,1]
return TB
#Calculate TB_own
def calTB_own(x):
# change input x as numpy array
# transiton from (w, n, e, s, z) -> (w, n, M, e, s, z, H)
TB = x[:,0] + x[:,1] + x[:,6]*pt - x[:,2]
return TB
#Reward function for renting
def u_rent(a):
'''
Input:
action a: c, b, k, h = a
Output:
reward value: the length of return should be equal to the length of a
'''
c = a[:,0]
h = a[:,3]
C = np.float_power(c, alpha) * np.float_power(h, 1-alpha)
return u(C)
#Reward function for owning
def u_own(a):
'''
Input:
action a: c, b, k, M, H = a
Output:
reward value: the length of return should be equal to the length of a
'''
c = a[:,0]
H = a[:,4]
C = np.float_power(c, alpha) * np.float_power((1+kappa)*H, 1-alpha)
return u(C)
# +
def transition_to_rent(x,a,t):
'''
imput: a is np array constains all possible actions
output: from x = [w, n, e, s, z] to x = [w, n, e, s, z]
'''
w, n, e, s, z = x
s = int(s)
e = int(e)
nX = len(x)
aSize = len(a)
# actions
b = a[:,1]
k = a[:,2]
h = a[:,3]
# transition of z
z_next = np.ones(aSize)
if z == 0:
z_next[k==0] = 0
# transition before T_R and after T_R
if t >= T_R:
future_states = np.zeros((aSize*nS,nX))
n_next = gn(t, x, (r_k+r_b)/2)
future_states[:,0] = np.repeat(b*(1+r_b[s]), nS) + np.repeat(k, nS)*(1+np.tile(r_k, aSize))
future_states[:,1] = np.tile(n_next,aSize)
future_states[:,2] = 0
future_states[:,3] = np.tile(range(nS),aSize)
future_states[:,4] = np.repeat(z_next,nS)
future_probs = np.tile(Ps[s],aSize)
else:
future_states = np.zeros((2*aSize*nS,nX))
n_next = gn(t, x, (r_k+r_b)/2)
future_states[:,0] = np.repeat(b*(1+r_b[s]), 2*nS) + np.repeat(k, 2*nS)*(1+np.tile(r_k, 2*aSize))
future_states[:,1] = np.tile(n_next,2*aSize)
future_states[:,2] = np.tile(np.repeat([0,1],nS), aSize)
future_states[:,3] = np.tile(range(nS),2*aSize)
future_states[:,4] = np.repeat(z_next,2*nS)
# employed right now:
if e == 1:
future_probs = np.tile(np.append(Ps[s]*Pe[s,e], Ps[s]*(1-Pe[s,e])),aSize)
else:
future_probs = np.tile(np.append(Ps[s]*(1-Pe[s,e]), Ps[s]*Pe[s,e]),aSize)
return future_states, future_probs
def transition_to_own(x,a,t):
'''
imput a is np array constains all possible actions
from x = [w, n, e, s, z] to x = [w, n, M, e, s, z, H]
'''
w, n, e, s, z = x
s = int(s)
e = int(e)
nX = len(x)+2
aSize = len(a)
# actions
b = a[:,1]
k = a[:,2]
M = a[:,3]
M_next = M*(1+rh)
H = a[:,4]
# transition of z
z_next = np.ones(aSize)
if z == 0:
z_next[k==0] = 0
# transition before T_R and after T_R
if t >= T_R:
future_states = np.zeros((aSize*nS,nX))
n_next = gn(t, x, (r_k+r_b)/2)
future_states[:,0] = np.repeat(b*(1+r_b[s]), nS) + np.repeat(k, nS)*(1+np.tile(r_k, aSize))
future_states[:,1] = np.tile(n_next,aSize)
future_states[:,2] = np.repeat(M_next,nS)
future_states[:,3] = 0
future_states[:,4] = np.tile(range(nS),aSize)
future_states[:,5] = np.repeat(z_next,nS)
future_states[:,6] = np.repeat(H,nS)
future_probs = np.tile(Ps[s],aSize)
else:
future_states = np.zeros((2*aSize*nS,nX))
n_next = gn(t, x, (r_k+r_b)/2)
future_states[:,0] = np.repeat(b*(1+r_b[s]), 2*nS) + np.repeat(k, 2*nS)*(1+np.tile(r_k, 2*aSize))
future_states[:,1] = np.tile(n_next,2*aSize)
future_states[:,2] = np.repeat(M_next,2*nS)
future_states[:,3] = np.tile(np.repeat([0,1],nS), aSize)
future_states[:,4] = np.tile(range(nS),2*aSize)
future_states[:,5] = np.repeat(z_next,2*nS)
future_states[:,6] = np.repeat(H,2*nS)
# employed right now:
if e == 1:
future_probs = np.tile(np.append(Ps[s]*Pe[s,e], Ps[s]*(1-Pe[s,e])),aSize)
else:
future_probs = np.tile(np.append(Ps[s]*(1-Pe[s,e]), Ps[s]*Pe[s,e]),aSize)
return future_states, future_probs
# +
class Approxy(object):
def __init__(self, pointsRent, Vrent, Vown, t):
self.Vrent = Vrent
self.Vown = Vown
self.Prent = pointsRent
self.t = t
def predict(self, xx):
if xx.shape[1] == 5:
# x = [w, n, e, s, z]
pvalues = np.zeros(xx.shape[0])
for e in [0,1]:
for s in range(nS):
for z in [0,1]:
index = (xx[:,2] == e) & (xx[:,3] == s) & (xx[:,4] == z)
pvalues[index]=interpn(self.Prent, self.Vrent[:,:,e,s,z], xx[index][:,:2],
bounds_error = False, fill_value = None)
return pvalues
else:
# x = w, n, M, e, s, z, H
pvalues = np.zeros(xx.shape[0])
for i in range(len(H_options)):
H = H_options[i]
# Mortgage amount, * 0.25 is the housing price per unit
Ms = np.array([0.01*H,0.05*H,0.1*H,0.2*H,0.3*H,0.4*H,0.5*H,0.8*H]) * pt
points = (ws,ns,Ms)
for e in [0,1]:
for s in range(nS):
for z in [0,1]:
index = (xx[:,3] == e) & (xx[:,4] == s) & (xx[:,5] == z) & (xx[:,6] == H)
pvalues[index]=interpn(points, self.Vown[i][:,:,:,e,s,z,self.t], xx[index][:,:3],
method = "nearest",bounds_error = False, fill_value = None)
return pvalues
# used to calculate dot product
def dotProduct(p_next, uBTB, t):
if t >= T_R:
return (p_next*uBTB).reshape((len(p_next)//(nS),(nS))).sum(axis = 1)
else:
return (p_next*uBTB).reshape((len(p_next)//(2*nS),(2*nS))).sum(axis = 1)
# Value function is a function of state and time, according to the restriction transfer from renting to ownning can only happen
# between the age: 0 - 25
def V(x, t, NN):
w, n, e, s, z = x
yat = yAT(t,x)
# first define the objective function solver and then the objective function
def obj_solver_rent(obj_rent):
# a = [c, b, k, h]
# Constrain: yat + w = c + b + k + pr*h
actions = []
for hp in np.linspace(0.001,0.999,20):
budget1 = yat + w
h = budget1 * hp/pr
budget2 = budget1 * (1-hp)
for cp in np.linspace(0.001,0.999,11):
c = budget2*cp
budget3 = budget2 * (1-cp)
#.....................stock participation cost...............
for kp in np.linspace(0,1,11):
# If z == 1 pay for matainance cost Km = 0.5
if z == 1:
# kk is stock allocation
kk = budget3 * kp
if kk > Km:
k = kk - Km
b = budget3 * (1-kp)
else:
k = 0
b = budget3
# If z == 0 and k > 0 payfor participation fee Kc = 5
else:
kk = budget3 * kp
if kk > Kc:
k = kk - Kc
b = budget3 * (1-kp)
else:
k = 0
b = budget3
#..............................................................
actions.append([c,b,k,h])
actions = np.array(actions)
values = obj_rent(actions)
fun = np.max(values)
ma = actions[np.argmax(values)]
return fun, ma
def obj_solver_own(obj_own):
# a = [c, b, k, M, H]
# possible value of H = {750, 1000, 1500, 2000} possible value of [0.2H, 0.5H, 0.8H]]*pt
# (M, t, rh) --> m
# Constrain: yat + w = c + b + k + (H*pt - M) + ch
actions = []
for H in H_options:
for mp in M_options:
M = mp*H*pt
m = M/D[T_max - t]
# 5 is the welfare income which is also the minimum income
if (H*pt - M) + c_h <= yat + w and m < pr*H + 5:
budget1 = yat + w - (H*pt - M) - c_h
for cp in np.linspace(0.001,0.999,11):
c = budget1*cp
budget2 = budget1 * (1-cp)
#.....................stock participation cost...............
for kp in np.linspace(0,1,11):
# If z == 1 pay for matainance cost Km = 0.5
if z == 1:
# kk is stock allocation
kk = budget2 * kp
if kk > Km:
k = kk - Km
b = budget2 * (1-kp)
else:
k = 0
b = budget2
# If z == 0 and k > 0 payfor participation fee Kc = 5
else:
kk = budget2 * kp
if kk > Kc:
k = kk - Kc
b = budget2 * (1-kp)
else:
k = 0
b = budget2
#..............................................................
actions.append([c,b,k,M,H])
if len(actions) == 0:
return -np.inf, [0,0,0,0,0]
else:
actions = np.array(actions)
values = obj_own(actions)
fun = np.max(values)
ma = actions[np.argmax(values)]
return fun, ma
if t == T_max-1:
# The objective function of renting
def obj_rent(actions):
# a = [c, b, k, h]
x_next, p_next = transition_to_rent(x, actions, t)
uBTB = uB(calTB_rent(x_next))
return u_rent(actions) + beta * dotProduct(uBTB, p_next, t)
fun, action = obj_solver_rent(obj_rent)
return np.array([fun, action])
# If the agent is older that 25 or if the agent is unemployed then keep renting
elif t > 30 or t < 10 or e == 0:
# The objective function of renting
def obj_rent(actions):
# a = [c, b, k, h]
x_next, p_next = transition_to_rent(x, actions, t)
V_tilda = NN.predict(x_next) # V_rent_{t+1} used to approximate, shape of x is [w,n,e,s]
uBTB = uB(calTB_rent(x_next))
return u_rent(actions) + beta * (Pa[t] * dotProduct(V_tilda, p_next, t) + (1 - Pa[t]) * dotProduct(uBTB, p_next, t))
fun, action = obj_solver_rent(obj_rent)
return np.array([fun, action])
# If the agent is younger that 45 and agent is employed.
else:
# The objective function of renting
def obj_rent(actions):
# a = [c, b, k, h]
x_next, p_next = transition_to_rent(x, actions, t)
V_tilda = NN.predict(x_next) # V_rent_{t+1} used to approximate, shape of x is [w,n,e,s]
uBTB = uB(calTB_rent(x_next))
return u_rent(actions) + beta * (Pa[t] * dotProduct(V_tilda, p_next, t) + (1 - Pa[t]) * dotProduct(uBTB, p_next, t))
# The objective function of owning
def obj_own(actions):
# a = [c, b, k, M, H]
x_next, p_next = transition_to_own(x, actions, t)
V_tilda = NN.predict(x_next) # V_own_{t+1} used to approximate, shape of x is [w, n, M, e, s, H]
uBTB = uB(calTB_own(x_next))
return u_own(actions) + beta * (Pa[t] * dotProduct(V_tilda, p_next, t) + (1 - Pa[t]) * dotProduct(uBTB, p_next, t))
fun1, action1 = obj_solver_rent(obj_rent)
fun2, action2 = obj_solver_own(obj_own)
if fun1 > fun2:
return np.array([fun1, action1])
else:
return np.array([fun2, action2])
# +
# wealth discretization
ws = np.array([10,25,50,75,100,125,150,175,200,250,500,750,1000,1500,3000])
w_grid_size = len(ws)
# 401k amount discretization
ns = np.array([1, 5, 10, 15, 25, 50, 100, 150, 400, 1000])
n_grid_size = len(ns)
pointsRent = (ws, ns)
# dimentions of the state
dim = (w_grid_size, n_grid_size, 2, nS, 2)
dimSize = len(dim)
xgrid = np.array([[w, n, e, s, z]
for w in ws
for n in ns
for e in [0,1]
for s in range(nS)
for z in [0,1]
]).reshape(dim + (dimSize,))
xs = xgrid.reshape((np.prod(dim),dimSize))
Vgrid = np.zeros(dim + (T_max,))
cgrid = np.zeros(dim + (T_max,))
bgrid = np.zeros(dim + (T_max,))
kgrid = np.zeros(dim + (T_max,))
hgrid = np.zeros(dim + (T_max,))
# Policy function of buying a house
Mgrid = np.zeros(dim + (T_max,))
Hgrid = np.zeros(dim + (T_max,))
# # Define housing choice part: Housing unit options and Mortgage amount options
V1000 = np.load("Vgrid1000.npy")
V1500 = np.load("Vgrid1500.npy")
V2000 = np.load("Vgrid2000.npy")
V750 = np.load("Vgrid750.npy")
H_options = [750, 1000, 1500, 2000]
M_options = [0.2, 0.5, 0.8]
Vown = [V750, V1000, V1500, V2000]
# +
# %%time
# value iteration part
pool = Pool()
for t in range(T_max-1,T_min, -1):
print(t)
if t == T_max - 1:
f = partial(V, t = t, NN = None)
results = np.array(pool.map(f, xs))
else:
approx = Approxy(pointsRent,Vgrid[:,:,:,:,:,t+1], Vown, t+1)
f = partial(V, t = t, NN = approx)
results = np.array(pool.map(f, xs))
Vgrid[:,:,:,:,:,t] = results[:,0].reshape(dim)
cgrid[:,:,:,:,:,t] = np.array([r[0] for r in results[:,1]]).reshape(dim)
bgrid[:,:,:,:,:,t] = np.array([r[1] for r in results[:,1]]).reshape(dim)
kgrid[:,:,:,:,:,t] = np.array([r[2] for r in results[:,1]]).reshape(dim)
# if a = [c, b, k, h]
hgrid[:,:,:,:,:,t] = np.array([r[3] if len(r) == 4 else r[4] for r in results[:,1]]).reshape(dim)
# if a = [c, b, k, M, H]
Mgrid[:,:,:,:,:,t] = np.array([r[3] if len(r) == 5 else 0 for r in results[:,1]]).reshape(dim)
Hgrid[:,:,:,:,:,t] = np.array([r[4] if len(r) == 5 else 0 for r in results[:,1]]).reshape(dim)
pool.close()
np.save("Vgrid_renting",Vgrid)
np.save("cgrid_renting",cgrid)
np.save("bgrid_renting",bgrid)
np.save("kgrid_renting",kgrid)
np.save("hgrid_renting",hgrid)
np.save("Mgrid_renting",Mgrid)
np.save("Hgrid_renting",Hgrid)
# -
| 20201120/simpleRenting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbsphinx="hidden"
# This notebook is part of https://github.com/AudioSceneDescriptionFormat/splines, see also http://splines.readthedocs.io/.
# -
# # Uniform Kochanek-Bartels Splines (TCB Splines)
#
# Kochanek-Bartels splines are a superset of Cardinal splines which themselves are a superset of [Catmull-Rom splines](catmull-rom-uniform.ipynb).
# They have three parameters per vertex (of course they can also be chosen to be the same values for the whole spline).
#
# The parameters are called
# $T$ for "tension",
# $C$ for "continuity" and
# $B$ for "bias".
# With the default values of $C = 0$ and $B = 0$, a Kochanek-Bartels spline is identical with a cardinal spline.
# If the "tension" parameter also has its default value $T = 0$ it is identical with a Catmull-Rom spline.
# Starting point: tangent vector from Catmull-Rom splines:
#
# \begin{equation}
# \boldsymbol{\dot{x}}_0 = \frac{
# (\boldsymbol{x}_0 - \boldsymbol{x}_{-1}) +
# (\boldsymbol{x}_1 - \boldsymbol{x}_0)
# }{2}
# \end{equation}
# ## Parameters
# ### Tension
#
# \begin{equation}
# \boldsymbol{\dot{x}}_0 = (1 - T_0) \frac{
# (\boldsymbol{x}_0 - \boldsymbol{x}_{-1}) +
# (\boldsymbol{x}_1 - \boldsymbol{x}_0)
# }{2}
# \end{equation}
#
# TODO: comparison with "tension" parameter of cardinal splines
#
# TODO: images
# ### Continuity
#
# Up to now, the goal was having a continuous second derivative at the control points, i.e. the incoming and outgoing tangent vectors are identical:
#
# \begin{equation}
# \boldsymbol{\dot{x}}_0 = \boldsymbol{\dot{x}}_0^{(-)} = \boldsymbol{\dot{x}}_0^{(+)}
# \end{equation}
#
# The "continuity" parameter allows us to break this continuity if we so desire:
#
# \begin{align}
# \boldsymbol{\dot{x}}_0^{(-)} &= \frac{
# (1 - C_0) (\boldsymbol{x}_0 - \boldsymbol{x}_{-1}) +
# (1 + C_0) (\boldsymbol{x}_1 - \boldsymbol{x}_0)
# }{2}\\
# \boldsymbol{\dot{x}}_0^{(+)} &= \frac{
# (1 + C_0) (\boldsymbol{x}_0 - \boldsymbol{x}_{-1}) +
# (1 - C_0) (\boldsymbol{x}_1 - \boldsymbol{x}_0)
# }{2}
# \end{align}
#
# When $C_0 = 0$, we are back at a Catmull-Rom spline.
# When $C_0 = -1$, we get a tangent like in a piecewise linear curve.
# When $C_0 = 1$, we get some weird "inverse corners".
#
# TODO: Example: compare $T_0 = 1$ and $C_0 = -1$: similar shape (a.k.a. "image"), different timing
# ### Bias
#
# \begin{equation}
# \boldsymbol{\dot{x}}_0 = \frac{
# (1 + B_0) (\boldsymbol{x}_0 - \boldsymbol{x}_{-1}) +
# (1 - B_0) (\boldsymbol{x}_1 - \boldsymbol{x}_0)
# }{2}
# \end{equation}
# ### All Three Combined
#
# \begin{align}
# \boldsymbol{\dot{x}}_0^{(+)} &= \frac{
# (1 - T_0) (1 + C_0) (1 + B_0) (\boldsymbol{x}_0 - \boldsymbol{x}_{-1}) +
# (1 - T_0) (1 - C_0) (1 - B_0) (\boldsymbol{x}_1 - \boldsymbol{x}_0)
# }{2}\\
# \boldsymbol{\dot{x}}_1^{(-)} &= \frac{
# (1 - T_1) (1 - C_1) (1 + B_1) (\boldsymbol{x}_1 - \boldsymbol{x}_0) +
# (1 - T_1) (1 + C_1) (1 - B_1) (\boldsymbol{x}_2 - \boldsymbol{x}_1)
# }{2}
# \end{align}
# TODO: cite Kochanek and Bartels, equation 9
# TODO: cite Kochanek and Bartels, equation 8
# Note: There is an error in eq (6.11) of Ian Millington's paper (all subscripts of $x$ are wrong, most likely copy-pasted from the preceding equation).
# To simplify the result we will get later, we introduce the following shorthands (as suggested in Millington's paper):
#
# \begin{align}
# a &= (1 - T_0) (1 + C_0) (1 + B_0)\\
# b &= (1 - T_0) (1 - C_0) (1 - B_0)\\
# c &= (1 - T_1) (1 - C_1) (1 + B_1)\\
# d &= (1 - T_1) (1 + C_1) (1 - B_1)
# \end{align}
#
# This leads to the simplified equations
#
# \begin{align}
# \boldsymbol{\dot{x}}_0^{(+)} &= \frac{
# a (\boldsymbol{x}_0 - \boldsymbol{x}_{-1}) +
# b (\boldsymbol{x}_1 - \boldsymbol{x}_0)
# }{2}\\
# \boldsymbol{\dot{x}}_1^{(-)} &= \frac{
# c (\boldsymbol{x}_1 - \boldsymbol{x}_0) +
# d (\boldsymbol{x}_2 - \boldsymbol{x}_1)
# }{2}
# \end{align}
# ## Calculation
# %matplotlib inline
from IPython.display import display
import sympy as sp
sp.init_printing()
from utility import NamedExpression, NamedMatrix
# Same control values as Catmull-Rom ...
x_1, x0, x1, x2 = sp.symbols('xbm_-1 xbm:3')
control_values_KB = sp.Matrix([x_1, x0, x1, x2])
control_values_KB
# ... but three additional parameters per vertex.
# In our calculation, the parameters belonging to $\boldsymbol{x}_0$ and $\boldsymbol{x}_1$ are relevant:
T0, T1 = sp.symbols('T:2')
C0, C1 = sp.symbols('C:2')
B0, B1 = sp.symbols('B:2')
a = NamedExpression('a', (1 - T0) * (1 + C0) * (1 + B0))
b = NamedExpression('b', (1 - T0) * (1 - C0) * (1 - B0))
c = NamedExpression('c', (1 - T1) * (1 - C1) * (1 + B1))
d = NamedExpression('d', (1 - T1) * (1 + C1) * (1 - B1))
display(a, b, c, d)
xd0 = NamedExpression('xdotbm0', sp.S.Half * (a.name * (x0 - x_1) + b.name * (x1 - x0)))
xd1 = NamedExpression('xdotbm1', sp.S.Half * (c.name * (x1 - x0) + d.name * (x2 - x1)))
display(xd0, xd1)
display(xd0.subs([a, b]))
display(xd1.subs([c, d]))
# Same as with Catmull-Rom, try to find a transformation from cardinal control values to Hermite control values.
# This can be used to get the full characteristic matrix.
control_values_H = sp.Matrix([x0, x1, xd0.name, xd1.name])
control_values_H
# From the [notebook about uniform Hermite splines](hermite-uniform.ipynb):
M_H = NamedMatrix(
r'{M_\text{H}}',
sp.S('Matrix([[2, -2, 1, 1], [-3, 3, -2, -1], [0, 0, 1, 0], [1, 0, 0, 0]])'))
M_H
M_KBtoH = NamedMatrix(r'{M_\text{KB$\to$H}}', 4, 4)
M_KB = NamedMatrix(r'{M_\text{KB}}', M_H.name * M_KBtoH.name)
M_KB
sp.Eq(control_values_H, M_KBtoH.name * control_values_KB)
# If we substitute the above definitions of $\boldsymbol{\dot{x}}_0$ and $\boldsymbol{\dot{x}}_1$, we can directly read off the matrix elements:
M_KBtoH.expr = sp.Matrix([[expr.coeff(cv) for cv in control_values_KB]
for expr in control_values_H.subs([xd0.args, xd1.args]).expand()])
M_KBtoH
M_KBtoH.pull_out(sp.S.Half)
M_KB = M_KB.subs([M_H, M_KBtoH]).doit()
M_KB
M_KB.pull_out(sp.S.Half)
# And for completeness' sake, its inverse:
M_KB.I
t = sp.symbols('t')
b_KB = NamedMatrix(r'{b_\text{KB}}', sp.Matrix([t**3, t**2, t, 1]).T * M_KB.expr)
b_KB.T
# To be able to plot the basis functions, let's substitute $a$, $b$, $c$ and $d$ back in (which isn't pretty):
b_KB = b_KB.subs([a, b, c, d]).simplify()
b_KB.T
sp.plot(*b_KB.expr.subs({T0: 0, T1: 0, C0: 0, C1: 1, B0: 0, B1: 0}), (t, 0, 1));
sp.plot(*b_KB.expr.subs({T0: 0, T1: 0, C0: 0, C1: -0.5, B0: 0, B1: 0}), (t, 0, 1));
# TODO: plot some example curves
| doc/kochanek-bartels-uniform.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# name: python2
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/SuyashSonawane/python_workshop/blob/master/Student_database.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="oY9qjPAWNapA" colab_type="code" colab={}
class Student:
def __init__(self, name,branch,phone,age,roll_no):
self.name = name
self.branch=branch
self.phone=phone
self.age=age
self.roll_no=roll_no
def print_info(self):
print('Name of the student is : ' + self.name + ' and is in ' + self.branch + ' barnch' + ' his roll number is '+ str(self.roll_no)+ ', his phone number is '+ str(self.phone)+"\n\n" )
# + id="Km301DyHN-_k" colab_type="code" outputId="b6c51e2b-1099-4726-ddba-d2865e35fbc8" colab={"base_uri": "https://localhost:8080/", "height": 1037}
def phone_check():
ph_number=int(raw_input('Enter the mobile number'))
return ph_number
if len(str(ph_number)) > 11:
print("Invaild mobile number\nEnter correct mobile number")
phone_check()
students_dict={}
def create_student():
stu_name=str(raw_input('Enter name of the student '))
stu_branch =raw_input('Enter the branch of the student ')
stu_age=int(raw_input('whats his age '))
stu_phone=phone_check()
stu_roll_no=int(input('Enter the roll number'))
students_dict[stu_roll_no]=(Student(stu_name,stu_branch,stu_phone,stu_age,stu_roll_no))
students_dict[stu_roll_no].print_info()
def search_student():
q=input('Enter the roll no of the student ')
if(q in students_dict):
students_dict[q].print_info()
else:
print("Student doesn't exist in database ")
def del_student():
r=int(raw_input('Enter the roll number of the student '))
if r in students_dict:
del students_dict[r]
print('student deleted')
if(len(students_dict)==0):
print('the database has no students now')
else:
print(str(len(students_dict)) + ' students are there in database now')
else:
print('No such student in database')
print('Welcome to student database center')
while True:
print('What do you want to do :\n1.Add student\n2.Search Students\n3.Delete Students\n99.Exit program ')
ans=int(raw_input())
if(ans==1):
n=int(input('How many students do you want to add '))
i=0
while i<n:
create_student();
i+=1
elif(ans==2):
search_student()
elif(ans==3):
del_student()
elif(ans==99):
print("thank you ")
break
else:
print('Enter correct option')
# + id="huSmLSNINYaQ" colab_type="code" colab={}
3
| Student_database.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# (02:System-setup)=
# # System setup
# <hr style="height:1px;border:none;color:#666;background-color:#666;" />
# If you intend to follow along with the code presented in this book, we recommend you follow these setup instructions so that you will run into fewer technical issues.
# ## The command-line interface
# A command-line interface\index{command-line interface} (CLI) is a text-based interface used to interact with your computer. We'll be using a CLI for various tasks throughout this book. We'll assume Mac and Linux users are using the "Terminal"\index{Terminal} and Windows users are using the "Anaconda Prompt"\index{Anaconda Prompt} (which we'll install in the next section) as a CLI.
# ## Installing software
# (02:Installing-Python)=
# ### Installing Python
# We recommend installing the latest version of Python via the Miniconda\index{Miniconda} distribution by following the instructions in the Miniconda [documentation](https://docs.conda.io/en/latest/miniconda.html). Miniconda is a lightweight version of the popular Anaconda\index{Anaconda} distribution. If you have previously installed the Anaconda or Miniconda distribution feel free to skip to **{numref}`02:Install-packaging-software`**.
#
# If you are unfamiliar with Miniconda and Anaconda, they are distributions of Python that also include the `conda`\index{conda} package and environment manager, and a number of other useful packages. The difference between Anaconda and Miniconda is that Anaconda installs over 250 additional packages (many of which you might never use), while Miniconda is a much smaller distribution that comes bundled with just a few key packages; you can then install additional packages as you need them using the command `conda install`.
#
# `conda` is a piece of software that supports the process of installing and updating software (like Python packages). It is also an environment manager, which is the key function we'll be using it for in this book. An environment manager helps you create "virtual environments\index{virtual environment}" on your machine, where you can safely install different packages and their dependencies in an isolated location. Installing all the packages you need in the same place (i.e., the system default location) can be problematic because different packages often depend on different versions of the same dependencies; as you install more packages, you'll inevitably get conflicts between dependencies, and your code will start to break. Virtual environments help you compartmentalize and isolate the packages you are using for different projects to avoid this issue. You can read more about virtual environments in the `conda` [documentation](https://conda.io/projects/conda/en/latest/user-guide/concepts/environments.html). While alternative package and environment managers exist, we choose to use `conda` in this book because of its popularity, ease-of-use, and ability to handle any software stack (not just Python).
# (02:Install-packaging-software)=
# ### Install packaging software
# Once you've installed the Miniconda\index{Miniconda} distribution, ensure that Python and `conda`\index{conda} are up to date by running the following command at the command line:
#
# ```{prompt} bash \$ auto
# $ conda update --all
# ```
#
# Now we'll install the two main pieces of software we'll be using to help us create Python packages in this book:
#
# 1. [`poetry`\index{poetry}](https://python-poetry.org/): software that will help us build our own Python packages. `poetry` is under active development, thus we recommend referring to the official [`poetry` documentation](https://python-poetry.org/docs/) for detailed installation instructions and support.
#
# 2. [`cookiecutter`\index{cookiecutter}](https://github.com/cookiecutter/cookiecutter): software that will help us create packages from pre-made templates. It can be installed with `conda` as follows:
#
# ```{prompt} bash \$ auto
# $ conda install -c conda-forge cookiecutter
# ```
# (02:Register-for-a-PyPI-account)=
# ## Register for a PyPI account
# The Python Package Index (PyPI)\index{PyPI} is the official online software repository for Python. A software repository\index{software repository} is a storage location for downloadable software, like Python packages. In this book we'll be publishing a package to PyPI. Before publishing packages to PyPI, it is typical to "test drive" their publication on TestPyPI\index{TestPyPI}, which is a test version of PyPI. To follow along with this book, you should register for a TestPyPI account on the [TestPyPI website](https://test.pypi.org/account/register/) and a PyPI account on the [PyPI website](https://pypi.org/account/register/).
# (02:Set-up-Git-and-GitHub)=
# ## Set up Git and GitHub
# If you're not using a version control\index{version control} system, we highly recommend you get into the habit! A version control system tracks changes to the file(s) of your project in a clear and organized way (no more "document_1.doc", "document_1_new.doc", "document_final.doc", etc.). As a result, a version control system contains a full history of all the revisions made to your project, which you can view and retrieve at any time. You don't *need* to use or be familiar with version control to read this book, but if you're serious about creating Python packages, version control will become an invaluable part of your workflow, so now is a good time to learn!
#
# There are many version control systems available, but the most common is Git\index{Git} and we'll be using it throughout this book. You can download Git by following the instructions in the [Git documentation](https://git-scm.com/book/en/v2/Getting-Started-Installing-Git). Git helps track changes to a project on a local computer, but what if we want to collaborate with others? Or, what happens if your computer crashes and you lose all your work? That's where GitHub\index{GitHub} comes in. GitHub is one of many online services for hosting Git-managed projects. GitHub helps you create an online copy of your local Git repository, which acts as a backup of your local work and allows others to easily and transparently collaborate on your project. You can sign up for a free GitHub account on the [GitHub website](https://www.github.com).
#
# We assume that those who choose to follow the optional version control sections of this book have basic familiarity with Git and GitHub (or equivalent). Two excellent learning resources are [*Happy Git and GitHub for the useR*](https://happygitwithr.com){cite:p}`bryan2021` and [*Research Software Engineering with Python*](https://merely-useful.tech/py-rse/git-cmdline.html){cite:p}`rsep2021`.
# ## Python integrated development environments
# A Python integrated development environment\index{integrated development environment} (IDE) will make the process of creating Python packages significantly easier. An IDE is a piece of software that provides advanced functionality for code development, such as directory and file creation and navigation, autocomplete, debugging, and syntax highlighting, to name a few. An IDE will save you time and help you write better code. Commonly used free Python IDEs include [Visual Studio Code\index{Visual Studio Code}](https://code.visualstudio.com/), [Atom](https://atom.io/), [Sublime Text](https://www.sublimetext.com/), [Spyder](https://www.spyder-ide.org/), and [PyCharm Community Edition](https://www.jetbrains.com/pycharm/). For those more familiar with the Jupyter\index{Jupyter} ecosystem, [JupyterLab](https://jupyter.org/) is a suitable browser-based IDE. Finally, for the R\index{R} community, the [RStudio\index{RStudio} IDE](https://rstudio.com/products/rstudio/download/) also supports Python.
#
# You'll be able to follow along with the examples presented in this book regardless of what IDE you choose to develop your Python code in. If you don't know which IDE to use, we recommend starting with Visual Studio Code. Below we briefly describe how to set up Visual Studio Code, JupyterLab, and RStudio as Python IDEs (these are the IDEs we personally use in our day-to-day work).
# ### Visual Studio Code
# You can download Visual Studio Code\index{Visual Studio Code} (VS Code) from the Visual Studio Code [website](https://code.visualstudio.com/). Once you've installed VS Code, you should install the "Python" extension from the VS Code Marketplace. To do this, follow the steps listed below and illustrated in {numref}`02-vscode-1-fig`:
#
# 1. Open the Marketplace by clicking the *Extensions* tab on the VS Code activity bar.
# 2. Search for "Python" in the search bar.
# 3. Select the extension named "Python" and then click *Install*.
#
# ```{figure} images/02-vscode-1.png
# ---
# width: 100%
# name: 02-vscode-1-fig
# alt: Installing the Python extension in Visual Studio Code.
# ---
# Installing the Python extension in Visual Studio Code.
# ```
#
# Once this is done, you have everything you need to start creating packages! For example, you can create files and directories from the *File Explorer* tab on the VS Code activity bar, and you can open up an integrated CLI by selecting *Terminal* from the *View* menu. {numref}`02-vscode-2-fig` shows an example of executing a Python *.py* file from the command line in VS Code.
#
# ```{figure} images/02-vscode-2.png
# ---
# width: 100%
# name: 02-vscode-2-fig
# alt: Executing a simple Python file called hello-world.py from the integrated terminal in Visual Studio Code.
# ---
# Executing a simple Python file called *hello-world.py* from the integrated terminal in Visual Studio Code.
# ```
#
# We recommend you take a look at the VS Code [Getting Started Guide](https://code.visualstudio.com/docs) to learn more about using VS Code. While you don't need to install any additional extensions to start creating packages in VS Code, there are many extensions available that can support and streamline your programming workflows in VS Code. Below are a few we recommend installing to support the workflows we use in this book (you can search for and install these from the "Marketplace" as we did earlier):
#
# - [Python Docstring Generator](https://marketplace.visualstudio.com/items?itemName=njpwerner.autodocstring): an extension to quickly generate documentation strings (docstrings\index{docstring}) for Python functions.
# - [Markdown All in One](https://marketplace.visualstudio.com/items?itemName=yzhang.markdown-all-in-one): an extension that provides keyboard shortcuts, automatic table of contents, and preview functionality for Markdown\index{Markdown} files. [Markdown](https://www.markdownguide.org) is a plain-text markup language that we'll use and learn about in this book.
# ### JupyterLab
# For those comfortable in the Jupyter\index{Jupyter} ecosystem feel free to stay there to create your Python packages! JupyterLab is a browser-based IDE that supports all of the core functionality we need to create packages. As per the JupyterLab [installation instructions](https://jupyterlab.readthedocs.io/en/stable/getting_started/installation.html), you can install JupyterLab with:
#
# ```{prompt} bash \$ auto
# $ conda install -c conda-forge jupyterlab
# ```
#
# Once installed, you can launch JupyterLab from your current directory by typing the following command in your terminal:
#
# ```{prompt} bash \$ auto
# $ jupyter lab
# ```
#
# In JupyterLab, you can create files and directories from the *File Browser* and can open up an integrated terminal from the *File* menu. {numref}`02-jupyterlab-fig` shows an example of executing a Python *.py* file from the command line in JupyterLab.
#
# ```{figure} images/02-jupyterlab.png
# ---
# width: 100%
# name: 02-jupyterlab-fig
# alt: Executing a simple Python file called hello-world.py from a terminal in JupyterLab.
# ---
# Executing a simple Python file called *hello-world.py* from a terminal in JupyterLab.
# ```
#
# We recommend you take a look at the JupyterLab [documentation](https://jupyterlab.readthedocs.io/en/stable/index.html) to learn more about how to use Jupyterlab. In particular, we'll note that, like VS Code, JupyterLab supports an ecosystem of extensions that can add additional functionality to the IDE. We won't install any here, but you can browse them in the JupyterLab *Extension Manager* if you're interested.
# ### RStudio
# Users with an R\index{R} background may prefer to stay in the RStudio\index{RStudio} IDE. We recommend installing the most recent version of the IDE from the RStudio [website](https://rstudio.com/products/rstudio/download/preview/) (we recommend installing at least version ^1.4) and then installing the most recent version of R from [CRAN](https://cran.r-project.org/). To use Python in RStudio, you will need to install the [reticulate\index{reticulate}](https://rstudio.github.io/reticulate/) R package by typing the following in the R console inside RStudio:
#
# ```r
# install.packages("reticulate")
# ```
#
# When installing reticulate, you may be prompted to install the Anaconda distribution. We already installed the Miniconda distribution of Python in **{numref}`02:Installing-Python`**, so answer "no" to this prompt. Before being able to use Python in RStudio, you will need to configure `reticulate`. We will briefly describe how to do this for different operating systems below, but we encourage you to look at the `reticulate` [documentation](https://rstudio.github.io/reticulate/) for more help.
# **Mac and Linux**
# 1. Find the path to the Python interpreter installed with Miniconda by typing `which python` at the command line.
# 2. Open (or create) an `.Rprofile` file in your HOME directory and add the line `Sys.setenv(RETICULATE_PYTHON = "path_to_python")`, where `"path_to_python"` is the path identified in step 1.
# 3. Open (or create) a `.bash_profile` file in your HOME directory and add the line `export PATH="/opt/miniconda3/bin:$PATH"`, replacing `/opt/miniconda3/bin` with the path you identified in step 1 but without the `python` at the end.
# 4. Restart R.
# 5. Try using Python in RStudio by running the following in the R console:
#
# ```r
# library(reticulate)
# repl_python()
# ```
# **Windows**
# 1. Find the path to the Python interpreter installed with Miniconda by opening an Anaconda Prompt from the Start Menu and typing `where python` in a terminal.
# 2. Open (or create) an `.Rprofile` file in your HOME directory and add the line `Sys.setenv(RETICULATE_PYTHON = "path_to_python")`, where `"path_to_python"` is the path identified in step 1. Note that in Windows, you need `\\` instead of `\` to separate the directories; for example your path might look like: `C:\\Users\\miniconda3\\python.exe`.
# 3. Open (or create) a `.bash_profile` file in your HOME directory and add the line `export PATH="/opt/miniconda3/bin:$PATH"`, replacing `/opt/miniconda3/bin` with the path you identified in step 1 but without the `python` at the end.
# 4. Restart R.
# 5. Try using Python in RStudio by running the following in the R console:
#
# ```r
# library(reticulate)
# repl_python()
# ```
#
# {numref}`02-rstudio-fig` shows an example of executing Python code interactively within the RStudio console.
#
# ```{figure} images/02-rstudio.png
# ---
# width: 100%
# name: 02-rstudio-fig
# alt: Executing Python code in the RStudio.
# ---
# Executing Python code in RStudio.
# ```
| py-pkgs/02-setup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.8 64-bit (''base'': conda)'
# name: python3
# ---
# +
# https://dplyr.tidyverse.org/reference/rename.html
from datar.datasets import iris
from datar.all import *
# %run nb_helpers.py
nb_header(rename, rename_with)
# -
rename(iris, petal_length='Petal_Length')
rename_with(iris, str.upper)
iris >> rename_with(str.upper, starts_with("Petal"))
iris >> rename_with(lambda x: x.replace('_', '.').lower())
# names can be selected by indexes
iris >> rename(Sp=5)
iris >> rename(Sp=4, base0_=True)
| docs/notebooks/rename.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (tensorflow_cpu)
# language: python
# name: tensorflow_cpu
# ---
# +
import tensorflow as tf
from tensorflow.keras import backend as K
from tensorflow.keras.layers import TimeDistributed
from tensorflow.keras.layers import LSTM
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import RepeatVector
from tensorflow.keras.layers import Input
from tensorflow.keras.layers import concatenate
from tensorflow.keras.models import Model
import matplotlib as mpl
import pickle
import matplotlib.pyplot as plt
import matplotlib
import numpy as np
import os
# Init NuScenes. Requires the dataset to be stored on disk.
from nuscenes.nuscenes import NuScenes
from nuscenes.map_expansion.map_api import NuScenesMap
matplotlib.rcParams['figure.figsize'] = (24, 18)
matplotlib.rcParams['figure.facecolor'] = 'white'
matplotlib.rcParams.update({'font.size': 20})
TRAIN_SIZE = 9800
TRAIN_TIME = 6
BATCH_SIZE = 32
BUFFER_SIZE = 500
# +
total_ped_matrix = np.load("../details/new_ped_matrix.npy")
with open("../details/ped_dataset.pkl", 'rb') as f:
ped_dataset = pickle.load(f)
with open("../details/scene_info.pkl", 'rb') as handle:
scene_info = pickle.load(handle)
# +
nusc = NuScenes(version='v1.0-trainval', \
dataroot='../../../../../data/', \
verbose=False)
so_map = NuScenesMap(dataroot='../../../../../data/', \
map_name='singapore-onenorth')
bs_map = NuScenesMap(dataroot='../../../../../data/', \
map_name='boston-seaport')
sh_map = NuScenesMap(dataroot='../../../../../data/', \
map_name='singapore-hollandvillage')
sq_map = NuScenesMap(dataroot='../../../../../data/', \
map_name='singapore-queenstown')
# dict mapping map name to map file
map_files = {'singapore-onenorth': so_map,
'boston-seaport': bs_map,
'singapore-hollandvillage': sh_map,
'singapore-queenstown': sq_map}
# +
# # calculating the values for standardization for every feature
# mean_values = np.mean(total_ped_matrix[:TRAIN_SIZE, :TRAIN_TIME, :], axis=(0,1))
# std_values = np.std(total_ped_matrix[:TRAIN_SIZE, :TRAIN_TIME, :], axis=(0,1))
# # standardization
# total_ped_matrix = (total_ped_matrix - mean_values) / std_values
# +
# train_test split
x_train = total_ped_matrix[:TRAIN_SIZE, :TRAIN_TIME, :]
y_train = total_ped_matrix[:TRAIN_SIZE, TRAIN_TIME:, :2]
x_test = total_ped_matrix[TRAIN_SIZE:, :TRAIN_TIME, :]
y_test = total_ped_matrix[TRAIN_SIZE:, TRAIN_TIME:, :2]
# +
train_data = tf.data.Dataset.from_tensor_slices((x_train, y_train))
train_data = train_data.cache().shuffle(BUFFER_SIZE).batch(BATCH_SIZE).repeat()
val_data = tf.data.Dataset.from_tensor_slices((x_test, y_test))
val_data = val_data.batch(BATCH_SIZE).repeat()
# +
# defining the custom rmse loss function
def model_loss(gt, pred):
'''
calculates custom rmse loss between every time point
'''
l2_x = K.square(gt[:,:,0] - pred[:,:,0])
l2_y = K.square(gt[:,:,1] - pred[:,:,1])
# log(sigma^2)
logs_x = pred[:,:,2] * 0.5
logs_y = pred[:,:,3] * 0.5
# sigma^2
s_x = K.exp(pred[:,:,2])
s_y = K.exp(pred[:,:,3])
# weight for aleatoric loss
w = 0.5
r = (l2_x/(2*w*s_x)) + (l2_y/(2*w*s_y))
return K.mean(r) + w*logs_x + w*logs_y
def euc_dist(gt, pred):
# custom metric to monitor rmse
gt_path = gt
pred_path = pred[:,:,:2]
gt_x = gt_path[:,:,0]
gt_y = gt_path[:,:,1]
pred_x = pred_path[:,:,0]
pred_y = pred_path[:,:,1]
rmse = K.mean(K.sqrt(K.sum(K.square(gt_path - pred_path), axis=1)))
return rmse
# +
traj_input = Input(shape=(x_train.shape[-2:]))
x = LSTM(16, activation='relu')(traj_input)
x = RepeatVector(10)(x)
x = LSTM(8, return_sequences=True, activation='relu')(x)
x_reg = TimeDistributed(Dense(2, activation='linear'))(x)
x_sig = TimeDistributed(Dense(2, activation='tanh'))(x)
combined_output = concatenate([x_reg, x_sig], axis=-1)
ul_model = Model(inputs=[traj_input], outputs=[combined_output])
ul_model.compile(optimizer='adam',
loss=model_loss, metrics=[euc_dist])
# -
ul_model.summary()
# +
# checkpoint for saving the best model
# filepath="../checkpoints/uncertain_lstm_best.hdf5"
# checkpoint = tf.keras.callbacks.ModelCheckpoint(filepath, monitor='val_euc_dist',
# verbose=1, save_best_only=True, mode='min')
# callbacks_list = [checkpoint]
train_history = ul_model.fit(train_data, epochs=30,
verbose=2, callbacks=None,
validation_data=val_data,
steps_per_epoch=400,
validation_steps=70
)
# -
def plot_train_history(history, title):
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(loss))
plt.figure()
plt.plot(epochs, loss, 'b', label='Training loss')
plt.plot(epochs, val_loss, 'r', label='Validation loss')
plt.title(title)
plt.legend()
plt.show()
plot_train_history(train_history, "MLP train and validation loss")
# +
# undo normalization for plotting
def move_from_origin(l, origin):
x0, y0 = origin
return [[x + x0, y + y0] for x, y in l]
def rotate_from_y(l, angle):
theta = -angle
return [(x*np.cos(theta) - y*np.sin(theta),
x*np.sin(theta) + y*np.cos(theta)) for x, y in l]
# loss calculation for test prediction
def rmse_error(l1, l2):
loss = 0.0
if len(np.array(l1).shape) < 2:
return ((l1[0] - l2[0])**2 + (l1[1] - l2[1])**2)**0.5
for p1, p2 in zip(l1, l2):
loss += ((p1[0] - p2[0])**2 + (p1[1] - p2[1])**2)**0.5
return (loss / float(len(l1)))
# +
# loading the model
ulstm_model = tf.keras.models.load_model("../checkpoints/uncertain_lstm_best.hdf5", compile=False)
ulstm_model.compile(optimizer=tf.keras.optimizers.RMSprop(clipvalue=1.0),
loss=model_loss,
metrics=[euc_dist])
# +
indexes = [9804, 9805, 11662, 12201, 11984, 11980, 11334, 11682, 11232,
11179, 9821, 10245, 10369]
for test_idx in indexes:
for i in np.linspace(0,9,4):
i = int(i)
# color setting for plot
alphas = np.linspace(1, 0.3, 10)
# for red the first column needs to be one
red_colors = np.zeros((10,4))
red_colors[:,0] = 1.0
# the fourth column needs to be your alphas
red_colors[:, 3] = alphas
test_data = total_ped_matrix[test_idx:test_idx+1,i:i+6,:]
predictions = ulstm_model.predict(test_data)[:,:,:2].reshape(-1, 2)
predictions = move_from_origin(rotate_from_y(predictions, ped_dataset[test_idx]["angle"]),
ped_dataset[test_idx]["origin"])
n_scene = ped_dataset[test_idx]["scene_no"]
ego_poses = map_files[scene_info[str(n_scene)]["map_name"]].render_pedposes_on_fancy_map(
nusc, scene_tokens=[nusc.scene[n_scene]['token']],
ped_path=np.array(ped_dataset[test_idx]["translation"])[:,:2],
verbose=False,
render_egoposes=True, render_egoposes_range=False,
render_legend=True)
plt.scatter(*zip(*np.array(ped_dataset[test_idx]["translation"])[i:i+6,:2]), c='k', s=5, zorder=2)
plt.scatter(*zip(*np.array(ped_dataset[test_idx]["translation"])[i+6:,:2]),
c='b', s=5, zorder=3)
plt.scatter(*zip(*predictions),
color=red_colors, s=5, zorder=4)
plt.savefig(f"../images/diffsteps_lstm/{test_idx}_{i}steps.png", bbox_inches='tight', pad_inches=0)
plt.close()
# loss = rmse_error(predictions,
# np.array(ped_dataset[test_idx]["translation"])[6:,:2])
# final_loss = rmse_error(predictions[-1],
# np.array(ped_dataset[test_idx]["translation"])[-1,:2])
# print(f"Loss in m is {loss}")
# print(f"Loss of final position in m is {final_loss}")
# -
print(ul_model.predict(total_ped_matrix[9800:9801,:6,:]))
| python-sdk/nuscenes/map_expansion/training/LSTM_uncertainty.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp utils
# -
# # utils.py
#
# > utilities + helpers
# hide
# %load_ext autoreload
# %autoreload 2
# hide
from nbdev.showdoc import *
from nbdev.export import notebook2script
# export
import json
import numpy as np
from mpl_toolkits.mplot3d import Axes3D
from pathlib import Path
# ## constants
# +
# exports
TRAIN_SUBJECTS = [1, 5, 6, 7, 8]
TEST_SUBJECTS = [9, 11]
H36M_NAMES = ['']*32
H36M_NAMES[0] = 'Hip'
H36M_NAMES[1] = 'RHip'
H36M_NAMES[2] = 'RKnee'
H36M_NAMES[3] = 'RFoot'
H36M_NAMES[6] = 'LHip'
H36M_NAMES[7] = 'LKnee'
H36M_NAMES[8] = 'LFoot'
H36M_NAMES[12] = 'Spine'
H36M_NAMES[13] = 'Thorax'
H36M_NAMES[14] = 'Neck/Nose'
H36M_NAMES[15] = 'Head'
H36M_NAMES[17] = 'LShoulder'
H36M_NAMES[18] = 'LElbow'
H36M_NAMES[19] = 'LWrist'
H36M_NAMES[25] = 'RShoulder'
H36M_NAMES[26] = 'RElbow'
H36M_NAMES[27] = 'RWrist'
N_CAMERAS = 4
N_JOINTS = 32
PLOT_RADIUS = 300
# -
# export
Path.ls = lambda x: list(x.iterdir())
# exports
data_path = Path('data')
data_path.ls()
# ## actions
# export
def get_actions(action):
"""
"""
actions = ['Directions',
'Discussion',
'Eating',
'Greeting',
'Phoning',
'Photo',
'Posing',
'Purchases',
'Sitting',
'SittingDown',
'Smoking',
'Waiting',
'WalkDog',
'Walking',
'WalkTogether']
if action == 'All' or action == 'all':
return actions
if action not in actions:
raise (ValueError, f'{action} is not found in {x for x in actions}')
return [action]
assert get_actions('all') == ['Directions', 'Discussion', 'Eating', 'Greeting', 'Phoning', 'Photo', 'Posing', 'Purchases', 'Sitting', 'SittingDown', 'Smoking', 'Waiting', 'WalkDog', 'Walking', 'WalkTogether']
assert get_actions('All') == ['Directions', 'Discussion', 'Eating', 'Greeting', 'Phoning', 'Photo', 'Posing', 'Purchases', 'Sitting', 'SittingDown', 'Smoking', 'Waiting', 'WalkDog', 'Walking', 'WalkTogether']
assert get_actions('Smoking') == ['Smoking']
# ## data_utils
# export
def normalize_data(unnormalized, mean, std, dim_use):
normalized = {}
for key in unnormalized.keys():
unnormalized[key] = unnormalized[key][:, dim_use]
m = mean[dim_use]
s = std[dim_use]
normalized[key] = np.divide((unnormalized[key] - m), s)
return normalized
# export
def unnormalize_data(normalized, mean, std, dim_ignore):
T = normalized.shape[0]
D = mean.shape[0]
orig = np.zeros((T, D), dtype=np.float32)
dim_use = np.array([dim for dim in range(D) if dim not in dim_ignore])
orig[:, dim_use] = normalized
std_m = std.reshape((1, D))
std_m = np.repeat(std_m, T, axis=0)
mean_m = mean.reshape((1, D))
mean_m = np.repeat(mean_m, T, axis=0)
orig = np.multiply(orig, std_m) + mean_m
return orig
# export
def normalize_kp(kp, mean, std, dim_use):
m = mean[dim_use]
s = std[dim_use]
return np.divide((kp - m), s)
# export
def get_kp_from_json(fname):
with open(fname) as f:
kp = json.load(f)
kpl = np.array(kp['people'][0]['pose_keypoints_2d'])
return kpl
# export
def coco_to_skel(s):
s = s.reshape(-1, 2)
hip = (s[8] + s[11]) / 2
rhip = s[8]
rknee = s[9]
rfoot = s[10]
lhip = s[11]
lknee = s[12]
lfoot = s[13]
spine = (s[1] + hip) / 2
thorax = s[1]
head = (s[16] + s[17]) / 2 # TODO: kurang tinggi
lshoulder = s[5]
lelbow = s[6]
lwrist = s[7]
rshoulder = s[2]
relbow = s[3]
rwrist = s[4]
return np.array([hip, rhip, rknee, rfoot, lhip, lknee, lfoot,
spine, thorax, head,
lshoulder, lelbow, lwrist,
rshoulder, relbow, rwrist ]).reshape(1, -1)
# ## cameras
# export
def get_cam_rt(key, rcams):
subj, _, sname = key
cname = sname.split('.')[1] # <-- camera name
scams = {(subj,c+1): rcams[(subj,c+1)] for c in range(N_CAMERAS)} # cams of this subject
scam_idx = [scams[(subj,c+1)][-1] for c in range(N_CAMERAS)].index( cname ) # index of camera used
the_cam = scams[(subj, scam_idx+1)] # <-- the camera used
R, T, f, c, k, p, name = the_cam
assert name == cname
return R, T
# export
def camera_to_world_frame(P, R, T):
X_cam = R.T.dot( P.T ) + T
return X_cam.T
# export
def cam_to_world_centered(data, key, rcams):
R, T = get_cam_rt(key, rcams)
data_3d_worldframe = camera_to_world_frame(data.reshape((-1, 3)), R, T)
data_3d_worldframe = data_3d_worldframe.reshape((-1, N_JOINTS*3))
# subtract root translation
return data_3d_worldframe - np.tile( data_3d_worldframe[:,:3], (1,N_JOINTS) )
# ## viz
# export
def show_2d_pose(skel, ax, lcolor='#094e94', rcolor='#940909'):
kps = np.reshape(skel, (len(H36M_NAMES), -1))
start = np.array([1,2,3,1,7,8,1, 13,14,14,18,19,14,26,27])-1 # start points
end = np.array([2,3,4,7,8,9,13,14,16,18,19,20,26,27,28])-1 # end points
left_right = np.array([1,1,1,0,0,0,0, 0, 0, 0, 0, 0, 1, 1, 1], dtype=bool)
for i in range(len(start)):
x, y = [np.array( [kps[start[i], j], kps[end[i], j]] ) for j in range(2)]
ax.plot(x, y, lw=2, c=lcolor if left_right[i] else rcolor)
ax.scatter(x, y, c=lcolor if left_right[i] else rcolor)
xroot, yroot = kps[0,0], kps[0,1]
ax.set_xlim(-PLOT_RADIUS+xroot, PLOT_RADIUS+xroot)
ax.set_ylim(-PLOT_RADIUS+yroot, PLOT_RADIUS+yroot)
# export
def show_3d_pose(skel, ax, lcolor='#094e94', rcolor='#940909'):
kps = np.reshape(skel, (len(H36M_NAMES), -1))
start = np.array([1,2,3,1,7,8,1, 13,14,15,14,18,19,14,26,27])-1 # start points
end = np.array([2,3,4,7,8,9,13,14,15,16,18,19,20,26,27,28])-1 # end points
left_right = np.array([1,1,1,0,0,0,0, 0, 0, 0, 0, 0, 0, 1, 1, 1], dtype=bool)
for i in np.arange( len(start) ):
x, y, z = [np.array( [kps[start[i], j], kps[end[i], j]] ) for j in range(3)]
ax.plot(x, y, z, lw=2, c=lcolor if left_right[i] else rcolor)
ax.scatter(x, y, z, c=lcolor if left_right[i] else rcolor)
xroot, yroot, zroot = kps[0,0], kps[0,1], kps[0,2]
ax.set_xlim3d([-PLOT_RADIUS*2+xroot, PLOT_RADIUS*2+xroot])
ax.set_zlim3d([-PLOT_RADIUS*2+zroot, PLOT_RADIUS*2+zroot])
ax.set_ylim3d([-PLOT_RADIUS*2+yroot, PLOT_RADIUS*2+yroot])
ax.set_xlabel("x")
ax.set_ylabel("y")
ax.set_zlabel("z")
white = (1.0, 1.0, 1.0, 0.0)
ax.w_xaxis.set_pane_color(white)
ax.w_yaxis.set_pane_color(white)
ax.w_xaxis.line.set_color(white)
ax.w_yaxis.line.set_color(white)
ax.w_zaxis.line.set_color(white)
# hide
notebook2script()
| 00_utils.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
from util.performance import plot_moderation_performance
from tqdm import tqdm
from kneed import DataGenerator, KneeLocator
import matplotlib.pyplot as plt
plt.style.use('seaborn-whitegrid')
# -
# load df
names = ['CNN2_BL', 'CNN2_MCD', 'CNN2_BBB', 'CNN2_EN']
label = ['KimCNN-BL', 'KimCNN-MCD', 'KimCNN-BBB', 'KimCNN-EN']
colours = ['b', 'r', 'y', 'g']
dfs_all = []
for name in names:
dfs = []
for i in range(5):
df = pd.read_pickle(f"../pickle/newsGroups/{name}_{i}.pkl")
dfs.append(df)
i = i+1
dfs_all.append(dfs)
plt.rc('font', size=17)
plt.rcParams["figure.figsize"] = (6,4)
# +
for i in range(len(dfs_all)):
plot_moderation_performance(dfs_all[i], 'u_lc', names[i], colours[i], average='micro')
handles, labels = plt.gca().get_legend_handles_labels()
order = [0,1,2,3]
plt.legend([handles[idx] for idx in order],[label[idx] for idx in order])
plt.xlim((0, 1))
plt.ylim((0.869, 1))
plt.xticks([0, .25, .50, .75, 1], ['0%', '25%', '50%', '75%', '100%'])
plt.yticks([0.88, .9, .92, .94, .96, .98, 1], ['0.88', '0.90', '0.92', '0.94', '0.96', '0.98', '1.00'])
plt.xlabel('Moderation Effort')
plt.ylabel('F1-Score')
plt.text(0.01, .990, '20NewsGroups')
plt.savefig('cnn2_20n_y.pdf', bbox_inches='tight')
# +
# Knee LC
values = []
for i in tqdm(range(len(dfs_all))):
a, b = plot_moderation_performance(dfs_all[i], 'u_lc', names[i], colours[i], average='micro', eps=1000)
values.append((a, b))
i = 0
for (a, b) in tqdm(values):
x = a
y = b.values
kneedle = KneeLocator(x, y, S=1, curve='concave', direction='increasing', interp_method="polynomial")
print('\n' + names[i])
print('Effort (%): ', round(kneedle.knee, 3))
print('F1_Score : ', y[int(kneedle.knee*1000)])
kneedle.plot_knee_normalized()
i+= 1
| evaluation/NewsGroups_CCN2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
ZAPIER_EMAIL_HOOK =
GMAPS_API_TOKEN =
start = '8925 Melrose Ave, West Hollywood, CA 90069'
destination = '3747 South La Brea Ave, Los Angeles, CA 90016'
from IPython.display import display, HTML
import googlemaps
from datetime import datetime
import requests
from urllib.parse import urlparse
import urllib.request as req
# +
gmaps = googlemaps.Client(key=GMAPS_API_TOKEN)
# Request directions via driving
now = datetime.now()
directions_result = gmaps.directions(start, destination, mode="driving", departure_time=now)
# -
routes = len(directions_result)
for route in directions_result:
print(route.keys())
duration = route['legs'][0]['duration']['text']
distance = route['legs'][0]['distance']['text']
steps = route['legs'][0]['steps']
warnings = route['warnings']
warnings
steps_to_get_there = ''
i = 0
for step in steps:
i+=1
steps_to_get_there += str(i) +' - '+ step['html_instructions'] + '<br>'
display(HTML(step['html_instructions']))
display(HTML(steps_to_get_there))
message = "Fastest route from {} to {} will take you {}. <br><br> {}".format(start, destination, duration, steps_to_get_there)
to_address = '<EMAIL>'
map_traffic = """<img width="600" src="https://maps.googleapis.com/maps/api/staticmap?size=600x400&path=enc%3A{}">""".format(req.pathname2url(directions_result[0]['overview_polyline']['points']))
body = {"text": message, "email": to_address, "traffic_map":map_traffic}
requests.post(ZAPIER_EMAIL_HOOK, data=body)
display(HTML(map_traffic))
| norush.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="bnIbwiK7Ohv2"
# %tensorflow_version 2.x
# %load_ext tensorboard
import tensorflow as tf
from tensorflow.keras import layers, models
import matplotlib.pyplot as plt
from os import path, walk
import numpy as np
import datetime
device_name = tf.test.gpu_device_name()
if device_name != '/device:GPU:0':
raise SystemError('GPU device not found')
print('Found GPU at: {}'.format(device_name))
# + id="5nL7VJBlkL0c"
ca_data_dir = path.join(path.curdir, "databases", "cajus-amarelos")
ca_train_ds = tf.keras.preprocessing.image_dataset_from_directory(
ca_data_dir,
validation_split=0.2,
subset="training",
image_size=(512, 512),
batch_size=12,
seed=234)
ca_val_ds = tf.keras.preprocessing.image_dataset_from_directory(
ca_data_dir,
validation_split=0.2,
subset="validation",
image_size=(512, 512),
batch_size=12,
seed=234)
# + id="Y2HYvq8QTlqb"
cv_data_dir = path.join(path.curdir, "databases", "cajus-vermelhos")
cv_train_ds = tf.keras.preprocessing.image_dataset_from_directory(
cv_data_dir,
validation_split=0.2,
subset="training",
image_size=(512, 512),
batch_size=12,
seed=123)
cv_val_ds = tf.keras.preprocessing.image_dataset_from_directory(
cv_data_dir,
validation_split=0.2,
subset="validation",
image_size=(512, 512),
batch_size=12,
seed=123)
# + id="oG5EAsiBTt1V"
train_ds = ca_train_ds.concatenate(cv_train_ds)
val_ds = ca_val_ds.concatenate(cv_val_ds)
# + id="lkBUOsIM2Rq0"
AUTOTUNE = tf.data.experimental.AUTOTUNE
train_ds = train_ds.shuffle(1000).cache().prefetch(buffer_size=AUTOTUNE)
val_ds = val_ds.shuffle(1000).cache().prefetch(buffer_size=AUTOTUNE)
# + id="MIcM_pZ2lWNV"
model = tf.keras.models.Sequential()
model.add(tf.keras.layers.experimental.preprocessing.RandomFlip("horizontal_and_vertical", input_shape=(512, 512, 3)))
model.add(tf.keras.layers.experimental.preprocessing.RandomRotation(0.4, fill_mode="nearest"))
# model.add(tf.keras.layers.Conv2D(16, (3, 3), activation="swish"))
# model.add(tf.keras.layers.MaxPooling2D((2, 2)))
model.add(tf.keras.layers.Conv2D(32, (3, 3), activation="swish"))
model.add(tf.keras.layers.MaxPooling2D((2, 2)))
model.add(tf.keras.layers.Conv2D(32, (3, 3), activation="swish"))
model.add(tf.keras.layers.MaxPooling2D((2, 2)))
model.add(tf.keras.layers.Conv2D(64, (3, 3), activation="swish"))
model.add(tf.keras.layers.MaxPooling2D((2, 2)))
model.add(tf.keras.layers.Conv2D(64, (3, 3), activation="swish"))
model.add(tf.keras.layers.MaxPooling2D((2, 2)))
model.add(tf.keras.layers.SpatialDropout2D(0.5))
model.add(tf.keras.layers.GlobalAveragePooling2D())
model.add(tf.keras.layers.Dense(128, activation="elu"))
model.add(tf.keras.layers.Dense(64, activation="elu"))
model.add(tf.keras.layers.Dense(32, activation="elu"))
model.add(tf.keras.layers.Dense(16, activation="elu"))
model.add(tf.keras.layers.Dropout(0.1))
model.add(tf.keras.layers.Dense(3, activation="softmax"))
model.summary()
# + id="L_tLzBdtld75"
model.compile(
optimizer=tf.keras.optimizers.Adam(learning_rate=0.0005),
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=["accuracy"]
)
# + id="rQXYFZm3V87Y"
earlyStopping = tf.keras.callbacks.EarlyStopping(monitor='val_loss', patience=50, verbose=1)
log_dir = "./logs/cajus-vermelhos-e-amarelos/rgb/" + datetime.datetime.now().strftime("%Y%m%d-%H%M%S")
tensorboard_callback = tf.keras.callbacks.TensorBoard(log_dir=log_dir, histogram_freq=1, profile_batch=0)
# + id="bxPYdIhBljsJ"
history = model.fit(
train_ds,
epochs=1000,
validation_data=val_ds,
callbacks=[earlyStopping, tensorboard_callback]
)
# + id="wlxRroBt-9F2"
loss, acc = model.evaluate(val_ds)
# + id="_DY5ce42FUim"
plt.figure()
plt.ylabel("Loss (training and validation)")
plt.xlabel("Training Steps")
plt.ylim([0,2])
plt.plot(history.history["loss"])
plt.plot(history.history["val_loss"])
plt.figure()
plt.ylabel("Accuracy (training and validation)")
plt.xlabel("Training Steps")
plt.ylim([0,1])
plt.plot(history.history["accuracy"])
plt.plot(history.history["val_accuracy"])
# + id="wYbVJIrwQW8O"
model.save('./models/cajus-vermelhos-e-amarelos/rgb')
# + id="0g7lmZx18pK9"
# %tensorboard --logdir ./logs/cajus-vermelhos-e-amarelos/rgb/
# + id="qUShrlskAfNq"
# !tensorboard dev upload \
# --logdir ./logs/cajus-vermelhos-e-amarelos/rgb/20210225-181939 \
# --name "cajus-vermelhos-e-amarelos-rgb" \
# --description "cnn model on cajus-vermelhos-e-amarelos rbg images" \
# --one_shot
# + id="u4gAfpd7TjY1"
loaded_model = tf.keras.models.load_model('./models/cajus-vermelhos-e-amarelos/rgb')
# + id="bxNkv3AWM8NJ"
loaded_model.to_json()
| notebooks/cajus-vermelhos-e-amarelos/regular/cajus-vermelhos-e-amarelos-rgb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import numpy as np
import pandas as pd
from avgn.utils.paths import DATA_DIR, most_recent_subdirectory, ensure_dir
from sklearn.preprocessing import scale
import matplotlib.pyplot as plt
from tqdm.autonotebook import tqdm
import seaborn as sns
from avgn.dataset_names import species_dict
hopkins_dfs = list(DATA_DIR.glob('clusterability/convex_sample_indvs/*.pickle'))
hopkins_dfs[:3], len(hopkins_dfs)
clusterability_df = pd.concat([pd.read_pickle(i) for i in tqdm(hopkins_dfs)])
clusterability_df['species'] = [species_dict[ds]['species'] for ds in clusterability_df['dataset'].values]
clusterability_df['family'] = [species_dict[ds]['group'] for ds in clusterability_df['dataset'].values]
pd.set_option('display.max_rows', 100)
clusterability_df['single_indv'] = True
grouped_indvs = ['giant_otter', 'gibbon_morita_segmented']
clusterability_df.loc[clusterability_df.dataset.isin(grouped_indvs), 'single_indv'] = False
clusterability_df[:3]
# %load_ext rpy2.ipython
# + magic_args="-i clusterability_df" language="R"
# library('lme4')
# glmer(data = clusterability_df)
# -
import pandas as pd
df = pd.DataFrame({
'cups_of_coffee': [0, 1, 2, 3, 4, 5, 6, 7, 8, 9],
'productivity': [2, 5, 6, 8, 9, 8, 0, 1, 0, -1]
})
# + magic_args="-i df -w 5 -h 5 --units in -r 200" language="R"
# # import df from global environment
# # make default figure size 5 by 5 inches with 200 dpi resolution
#
# install.packages("ggplot2", repos='http://cran.us.r-project.org', quiet=TRUE)
# library(ggplot2)
# ggplot(df, aes(x=cups_of_coffee, y=productivity)) + geom_line()
# -
clusterability_df[:3]
import statsmodels.api as sm
import statsmodels.formula.api as smf
md = smf.ols(
formula="umap_hopkins_10 ~ C(family)",
groups=clusterability_df["species"],
data=clusterability_df,
).fit()
print(md.summary())
md = smf.ols(
formula="umap_hopkins_10 ~ C(family) + (1 | C(species)*C(single_indv))",
groups=clusterability_df["species"],
data=clusterability_df,
).fit()
clusterability ~ family + (exemplars | (species*single_indv))
| notebooks/09.0-clusterability/delete-make-clusterability-statistic-linear-model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # OpenHPC環境の削除
#
# ---
#
# 構築したOpenHPC環境を削除します。
# ## パラメータの指定
#
# 削除を行うのに必要となるパラメータを入力します。
# ### VCCアクセストークンの入力
# VCノード, VCディスクを削除するためにVC Controller(VCC)のアクセストークンが必要となります。
# 次のセルを実行すると表示される入力枠にVCCのアクセストークンを入力してください。
#
# > アクセストークン入力後に Enter キーを押すことで入力が完了します。
# + tags=["vcp:skip", "vcp:parameters"]
from getpass import getpass
vcc_access_token = getpass()
# -
# 入力されたアクセストークンが正しいことを、実際にVCCにアクセスして確認します。
# + tags=["vcp:rewrite:vcpsdk"]
from common import logsetting
from vcpsdk.vcpsdk import VcpSDK
vcp = VcpSDK(vcc_access_token)
# -
# 上のセルの実行結果がエラーとなり以下のようなメッセージが表示されている場合は、入力されたアクセストークンに誤りがあります。
#
# ```
# 2018-09-XX XX:XX:XX,XXX - ERROR - config vc failed: http_status(403)
# 2018-09-XX XX:XX:XX,XXX - ERROR - 2018/XX/XX XX:XX:XX UTC: VCPAuthException: xxxxxxx:token lookup is failed: permission denied
# ```
#
# エラーになった場合はこの節のセルを全て `unfreeze` してから、もう一度アクセストークンの入力を行ってください。
# ### グループ名
# OpenHPCのUnitGroup名を指定してください。
#
# > VCノードを起動した際に指定したものと同じ名前を指定してください。
# + tags=["vcp:parameters"]
ugroup_name = 'OpenHPC'
# -
# ## 構築環境の削除
#
# 構築したOpenHPC環境を削除します。
# ### VCノードの削除
#
# 起動したVCノードを削除します。
# 現在のUnitGroupの一覧を確認します。
vcp.df_ugroups()
# 現在のVCノードの状態を確認します。
ug = vcp.get_ugroup(ugroup_name)
ug.df_nodes()
# まず計算ノード用VCノードを削除します。
ug.delete_units('compute', force=True)
# マスターノードと UnitGroup の削除を行います。
ug.cleanup()
# 削除後の UnitGroupの一覧を確認します。
vcp.df_ugroups()
# ### VCディスクの削除
# NFS用のVCディスクを削除します。
#
# > VCディスクを作成していない場合は、何もしません。
# 現在の状態を確認します。
from IPython.display import display
ug_disk = vcp.get_ugroup(ugroup_name + '_disk')
if ug_disk:
display(ug_disk.df_nodes())
# VCディスクを削除します。
if ug_disk:
ug_disk.cleanup()
# 削除後のUnitGroupの一覧を確認します。
vcp.df_ugroups()
# ## Ansible設定のクリア
#
# 削除した環境に対応するAnsibleの設定をクリアします。
# ### group_varsファイル
#
# group_varsファイルをリネームします。
# !mv group_vars/{ugroup_name}.yml group_vars/{ugroup_name}.yml.bak
# ### インベントリ
#
# インベントリから UnitGroup に対応するグループを削除します。
# +
import yaml
from pathlib import Path
inventory_path = Path('inventory.yml')
# !cp {str(inventory_path)} {str(inventory_path)}.bak
with inventory_path.open() as f:
inventory = yaml.safe_load(f)
if ugroup_name in inventory['all']['children']:
del(inventory['all']['children'][ugroup_name])
with inventory_path.open(mode='w') as f:
f.write(yaml.safe_dump(inventory))
# !cat {str(inventory_path)}
| OpenHPC-v1/notebooks/920-OpenHPC環境の削除.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # Commodity price forecasting - Baseline models
#
# - Naive
# - Seasonal naive
# - Random walk with drift
library(readr)
library(dplyr)
library(ggplot2)
library(gridExtra)
library(xts)
library(fpp2)
library(forecast)
library(DMwR)
library(Metrics)
library(hydroGOF)
options(repr.plot.width=8, repr.plot.height=3)
# ## 1. Data import and analysis
data <- read.csv("data/gold-silver.csv", sep=';')
head(data)
gold <- ts(data$gold, start = c(1993, 11), frequency = 12)
silver <- ts(data$silver, start = c(1993, 11), frequency = 12)
# +
p1 <- ggplot2::autoplot(gold) + ggtitle('gold price') + ylab('$')
p2 <- ggplot2::autoplot(silver) + ggtitle('silver price') + ylab('$')
gridExtra::grid.arrange(p1, p2, ncol=2)
# +
p3 <- ggplot2::autoplot(log(gold)) + ggtitle('gold price') + ylab('log($)')
p4 <- ggplot2::autoplot(log(silver)) + ggtitle('silver price') + ylab('log($)')
gridExtra::grid.arrange(p3, p4, ncol=2)
# -
nrow(data)
# ## 2. Train test split
# +
test_size <- as.numeric(12)
train_size <- length(gold) - test_size
train_gold <- head(gold, train_size)
test_gold <- tail(gold, test_size)
train_silver <- head(silver, train_size)
test_silver <- tail(silver, test_size)
# -
# ## 3. Evaluation function
model_evaluation <- function(model, frcst, train, test){
train_pred <- fitted(model)
test_pred <- frcst$mean
train_rmse <- sqrt(mse(train, train_pred))
train_mae <- mae(train, train_pred)
train_nrmse <- train_rmse/sd(train)
test_rmse <- sqrt(mse(test, test_pred))
test_mae <- mae(test, test_pred)
test_nrmse <- test_rmse/sd(test)
print(paste0('Training NRMSE :', round(train_nrmse, 3)))
print(paste0('Training MAE :', round(train_mae, 3)))
print(paste0('Test NRMSE :', round(test_nrmse, 3)))
print(paste0('Test MAE :', round(test_mae, 3)))
}
# ## 4. Naive forecasting
# +
## naive forecasting for gold
naive_model <- Arima(train_gold, order = c(0, 1, 0))
# multi-step forecasting
naive_frcst <- forecast(naive_model, h = 12)
model_evaluation(naive_model, naive_frcst, train_gold, test_gold)
autoplot(gold, series = 'actual data') +
autolayer(fitted(naive_model), series = 'train prediction') +
autolayer(naive_frcst$mean, series = 'test prediction') +
xlab('Year') +
ylab('$') +
ggtitle('Naive method forecasting for gold')
# +
## naive forecasting for gold
naive_model <- Arima(train_silver, order = c(0, 1, 0))
# multi-step forecasting
naive_frcst <- forecast(naive_model, h = 12)
model_evaluation(naive_model, naive_frcst, train_silver, test_silver)
autoplot(silver, series = 'actual data') +
autolayer(fitted(naive_model), series = 'train prediction') +
autolayer(naive_frcst$mean, series = 'test prediction') +
xlab('Year') +
ylab('$') +
ggtitle('Naive method forecasting for silver')
# -
# ## 5. Seasonal naive forecasting
# +
## seasonal naive forecasting for gold
snaive_model <- Arima(train_gold, order = c(0, 0, 0), seasonal = list(order = c(0, 1, 0)))
# multi-step forecasting
snaive_frcst <- forecast(snaive_model, h = 12)
model_evaluation(snaive_model, snaive_frcst, train_gold, test_gold)
autoplot(gold, series = 'actual data') +
autolayer(fitted(snaive_model), series = 'train prediction') +
autolayer(snaive_frcst$mean, series = 'test prediction') +
xlab('Year') +
ylab('$') +
ggtitle('Seasonal naive method forecasting for gold')
# +
## seasonal naive forecasting for silver
snaive_model <- Arima(train_silver, order = c(0, 0, 0), seasonal = list(order = c(0, 1, 0)))
# multi-step forecasting
snaive_frcst <- forecast(snaive_model, h = 12)
model_evaluation(snaive_model, snaive_frcst, train_silver, test_silver)
autoplot(silver, series = 'actual data') +
autolayer(fitted(snaive_model), series = 'train prediction') +
autolayer(snaive_frcst$mean, series = 'test prediction') +
xlab('Year') +
ylab('$') +
ggtitle('Seasonal naive method forecasting for silver')
# -
# ## 6. Random walk with drift
# +
## rwd forecasting for gold
rwd_model <- Arima(train_gold, order = c(0, 1, 0), include.drift = TRUE)
# multi-step forecasting
rwd_frcst <- forecast(rwd_model, h = 12)
model_evaluation(rwd_model, rwd_frcst, train_gold, test_gold)
autoplot(gold, series = 'actual data') +
autolayer(fitted(rwd_model), series = 'train prediction') +
autolayer(rwd_frcst$mean, series = 'test prediction') +
xlab('Year') +
ylab('$') +
ggtitle('Random walk with drift method forecasting for gold')
# +
## rwd forecasting for silver
rwd_model <- Arima(train_silver, order = c(0, 1, 0), include.drift = TRUE)
# multi-step forecasting
rwd_frcst <- forecast(rwd_model, h = 12)
model_evaluation(rwd_model, rwd_frcst, train_silver, test_silver)
autoplot(silver, series = 'actual data') +
autolayer(fitted(rwd_model), series = 'train prediction') +
autolayer(rwd_frcst$mean, series = 'test prediction') +
xlab('Year') +
ylab('$') +
ggtitle('Random walk with drift method forecasting for silver')
# -
| 04.gold_silver/01.commodity-forecasting-Baseline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: TensorFlow-1.13.1
# language: python
# name: tensorflow-1.13.1
# ---
# # 学习率与优化器
# 在本例中,通过比较**同样结构**,**同样数据集**,**相同训练轮数**下**不同优化器**以及**不同初始学习率**下训练的值得出优化器与初始学习率对训练的影响。
# 在模型训练过程中涉及到一个关键概念——**学习率**。学习率代表着模型学习的速率,学习率的值越大,损失函数的变化速度越快。较高的学习率大小可以使模型快速收敛,但是可能会导致模型在局部极小值周围震荡。较小的学习率虽然可以让模型在局部极小值周围收敛,但是收敛速度很慢。所以合理的学习率是在底部使用大学习率,在顶部使用小学习率来进行梯度下降。
#
# 对于学习率这样十分重要但是调整难度很大的参数,有几种自适应学习率算法进行学习率的调整。
# 下面我们采用不同的优化器,对相同的模型结构和数据集训练情况下分别进行训练。我们选择的不同的优化器为:
#
# - rmsprop
# - adam
# - SGD(stochastic gradient descent)
#
# 这些自适应学习率算法不需要开发者进行学习率的设置而是模型在训练过程中进行学习率的衰减。
# #### 注意:本次实验每一个训练都是只包含5个轮次的训练,每一个5轮次训练大约耗时20分钟。
# 实验之前我们进行keras,keras_applications版本配置以及数据集下载。
# !pip install --upgrade keras_applications==1.0.6 keras==2.2.4
import os
if os.path.exists('./data') == False:
from modelarts.session import Session
session = Session()
if session.region_name == 'cn-north-1':
bucket_path="modelarts-labs/end2end/image_recognition/dog_and_cat_25000.tar.gz"
elif session.region_name == 'cn-north-4':
bucket_path="modelarts-labs-bj4/end2end/image_recognition/dog_and_cat_25000.tar.gz"
else:
print("请更换地区到北京一或北京四")
session.download_data(
bucket_path=bucket_path,
path="./dog_and_cat_25000.tar.gz")
# 使用tar命令解压资源包
# !tar xf ./dog_and_cat_25000.tar.gz
# 清理压缩包
# !rm -f ./dog_and_cat_25000.tar.gz
# ## 引入相关的包
# +
from keras.applications.vgg16 import VGG16
from keras.preprocessing import image
from keras.applications.resnet50 import preprocess_input, decode_predictions
import numpy as np
from keras.applications.mobilenetv2 import MobileNetV2
from keras.preprocessing import image
from keras.models import Model
from keras.layers import Dense, GlobalAveragePooling2D
from keras import backend as K
from keras.models import load_model
from keras.preprocessing.image import ImageDataGenerator
import os
from PIL import Image
# -
# ## 加载数据
def load_data():
dirname = "./data"
path = "./data"
num_train_samples = 25000
x_train = np.empty((num_train_samples, 224,224,3), dtype='uint8')
y_train = np.empty((num_train_samples,1), dtype='uint8')
index = 0
for file in os.listdir("./data"):
image = Image.open(os.path.join(dirname,file)).resize((224,224))
image = np.array(image)
x_train[index,:,:,:] = image
if "cat" in file:
y_train[index,0] =1
elif "dog" in file:
y_train[index,0] =0
index += 1
return (x_train, y_train)
(x_train, y_train) = load_data()
print(x_train.shape)
print(y_train.shape)
# ## 数据处理
from keras.utils import np_utils
def process_data(x_train,y_train):
x_train = x_train.astype(np.float32)
x_train /= 255
n_classes = 2
y_train = np_utils.to_categorical(y_train, n_classes)
return x_train,y_train
x_train,y_train= process_data(x_train,y_train)
print(x_train.shape)
print(y_train.shape)
# ## 构建模型
def build_model(base_model):
x = base_model.output
x = GlobalAveragePooling2D()(x)
predictions = Dense(2, activation='softmax')(x)
model = Model(inputs=base_model.input, outputs=predictions)
print(type(model))
return model
base_model = VGG16(weights=None, include_top=False)
model = build_model(base_model)
model.summary()
# ## 定义优化器与训练
#
# ### rmsprop
# 在下面的测试中,我们使用优化器为**rmsprop**,训练轮数为5,可以看到训练初期的模型指数变化情况。
import keras
opt = keras.optimizers.rmsprop(lr=0.0001, decay=1e-6)
model.compile(loss='categorical_crossentropy',
optimizer=opt,
metrics=['accuracy'])
from keras.callbacks import ModelCheckpoint, EarlyStopping
es = EarlyStopping(monitor='val_acc', baseline=0.9, patience=30, verbose=1, mode='auto')
callbacks = [es]
# 开始训练
history_rmsprop = model.fit(x=x_train,
y=y_train,
batch_size=32,
epochs=5,
verbose=1,
callbacks=callbacks,
validation_split=0.25,
shuffle=True,
initial_epoch=0,
)
# +
import matplotlib.pyplot as plt
# 绘制训练 & 验证的准确率值
plt.plot(history_rmsprop.history['acc'])
plt.plot(history_rmsprop.history['val_acc'])
plt.title('Model accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# -
# 绘制训练 & 验证的损失值
plt.plot(history_rmsprop.history['loss'])
plt.plot(history_rmsprop.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# ### Adam
# 在下面的例子中,加载新的模型,使用**优化器Adam**,训练轮数为5,可以看到训练初期的模型指数变化情况。
base_model = VGG16(weights=None, include_top=False)
model_adam = build_model(base_model)
opt = keras.optimizers.Adam(lr=0.0001, decay=1e-6)
model_adam.compile(loss='categorical_crossentropy',
optimizer=opt,
metrics=['accuracy'])
# 开始训练
history_adam = model_adam.fit(x=x_train,
y=y_train,
batch_size=32,
epochs=5,
verbose=1,
callbacks=callbacks,
validation_split=0.25,
shuffle=True,
initial_epoch=0
)
# +
import matplotlib.pyplot as plt
# 绘制训练 & 验证的准确率值
plt.plot(history_adam.history['acc'])
plt.plot(history_adam.history['val_acc'])
plt.title('Model accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# -
# 绘制训练 & 验证的损失值
plt.plot(history_adam.history['loss'])
plt.plot(history_adam.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# ### SGD
# 在下面的例子中,加载新的模型,使用**优化器SGD**,训练轮数为5,可以看到训练初期的模型指数变化情况。可以看到SGD作为随机梯度下降模型,在训练初期较少的轮数下,表现并不稳定。
base_model = VGG16(weights=None, include_top=False)
model_sgd = build_model(base_model)
opt = keras.optimizers.SGD(lr=0.0001, decay=1e-6)
model_sgd.compile(loss='categorical_crossentropy',
optimizer=opt,
metrics=['accuracy'])
# 开始训练
history_sgd = model_sgd.fit(x=x_train,
y=y_train,
batch_size=32,
epochs=5,
verbose=1,
callbacks=callbacks,
validation_split=0.25,
shuffle=True,
initial_epoch=0,
)
# +
import matplotlib.pyplot as plt
# 绘制训练 & 验证的准确率值
plt.plot(history_sgd.history['acc'])
plt.plot(history_sgd.history['val_acc'])
plt.title('Model accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# -
# 绘制训练 & 验证的损失值
plt.plot(history_sgd.history['loss'])
plt.plot(history_sgd.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# **下面的训练中学习率的初始赋值与上一次训练相比由0.0001变为0.001**
# 下面的训练结果并不出色,对比我们之前的Adam优化器,这次的训练几乎没有收敛。因为过大的学习率,导致模型在局部最优附近震荡,无法获得好的结果。
base_model = VGG16(weights=None, include_top=False)
model_large_lr = build_model(base_model)
opt = keras.optimizers.Adam(lr=0.001, decay=1e-6)
model_large_lr.compile(loss='categorical_crossentropy',
optimizer=opt,
metrics=['accuracy'])
# 开始训练
history_large_lr = model_large_lr.fit(x=x_train,
y=y_train,
batch_size=32,
epochs=5,
verbose=1,
callbacks=callbacks,
validation_split=0.25,
shuffle=True,
initial_epoch=0,
)
# 绘制训练 & 验证的准确率值
plt.plot(history_large_lr.history['acc'])
plt.plot(history_large_lr.history['val_acc'])
plt.title('Model accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
plt.plot(history_large_lr.history['loss'])
plt.plot(history_large_lr.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# # 思考
#
# 以上的优化器对比都是在训练初期轮数较少的情况下进行,但是在轮数较多或者出现鞍点等情况时,不同的优化器都有不一样的表现。可以尝试用更多的轮数对比不同的优化器,重新审视各个模型在训练各个阶段的表现。
| notebook/DL_image_hyperparameter_tuning/01_lr_opt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Python for Epis
#
# All these files are here: https://github.com/kialio/py4Epis You should be able to install python and run them after this. Feel free to ask me questions now or later.
#
# * I'm going to give some background and then some high level examples as fast as I can...
# * There are many examples on the web.
# * There are even some for SAS Users. Here's a good one: https://github.com/RandyBetancourt/PythonForSASUsers
# + [markdown] slideshow={"slide_type": "slide"}
# # Objectives
# * Introduce you to the Python language
# * Show its utility in your research life
# * (I'm not going to show how to install python or get it going on your machine, if you want to get going quickly, check out conda: https://docs.conda.io/en/latest/)
#
# ## Credits
# * Borrowed heavily from https://github.com/profjsb/python-bootcamp
# + [markdown] slideshow={"slide_type": "slide"}
# # Who I Am
#
# ## <NAME>
# [@oldmanperkins](https://twitter.com/oldmanperkins)
#
# https://github.com/kialio
#
# I work at NASA/GSFC (here as a private citizen) and work on developing next generation gamma-ray instrumentation ([AMEGO](https://asd.gsfc.nasa.gov/amego/), [BurstCube](https://asd.gsfc.nasa.gov/burstcube/)). I use python to analyze data, control hardware, figure out budgets (I try to get data out of the excel spreadsheets my financial people give me as fast as possible), make pretty plots...
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Introduction
# * What is Python?
# * Why Python?
# * Getting Started...
# + [markdown] slideshow={"slide_type": "slide"}
# # What is Python?
#
# >Python is an interpreted, object-oriented, high-level programming language with dynamic semantics. Its high-level built in data structures, combined with dynamic typing and dynamic binding, make it very attractive for Rapid Application Development, as well as for use as a scripting or glue language to connect existing components together. Python's simple, easy to learn syntax emphasizes readability and therefore reduces the cost of program maintenance. Python supports modules and packages, which encourages program modularity and code reuse. The Python interpreter and the extensive standard library are available in source or binary form without charge for all major platforms, and can be freely distributed.
#
# https://www.python.org/doc/essays/blurb/
# + [markdown] slideshow={"slide_type": "subslide"}
# # What is Python?
# <table>
# <tr style="border-color: white;">
# <td style="border-color: white;">interpreted</td>
# <td style="border-color: white;">no need for a compiling stage</td>
# </tr>
# <tr style="background-color: #D8D8D8; border-color: white;">
# <td style="border-color: white;">object-oriented</td>
# <td style="border-color: white;">programming paradigm that uses objects (complex data structures with methods)</td>
# </tr>
# <tr style="border-color: white;">
# <td style="border-color: white;">high level</td>
# <td style="border-color: white;">abstraction from the way machine interprets & executes</td>
# </tr>
# <tr style="background-color: #D8D8D8; border-color: white;">
# <td style="border-color: white;">dynamic semantics</td>
# <td style="border-color: white;">can change meaning on-the-fly</td>
# </tr>
# <tr style="border-color: white;">
# <td style="border-color: white;">built in</td>
# <td style="border-color: white;">core language (not external)</td>
# </tr>
# <tr style="background-color: #D8D8D8; border-color: white;">
# <td style="border-color: white;">data structures</td>
# <td style="border-color: white;">ways of storing/manipulating data</td>
# </tr>
# <tr style="border-color: white;">
# <td style="border-color: white;">script/glue</td>
# <td style="border-color: white;">programs that control other programs</td>
# </tr>
# <tr style="background-color: #D8D8D8; border-color: white;">
# <td style="border-color: white;">typing</td>
# <td style="border-color: white;">the sort of variable (int, string)</td>
# </tr>
# <tr style="border-color: white;">
# <td style="border-color: white;">syntax</td>
# <td style="border-color: white;">grammar which defines the language</td>
# </tr>
# <tr style="background-color: #D8D8D8; border-color: white;">
# <td style="border-color: white;">library</td>
# <td style="border-color: white;">reusable collection of code</td>
# </tr>
# <tr style="border-color: white;">
# <td style="border-color: white;">binary</td>
# <td style="border-color: white;">a file that you can run/execute</td>
# </tr>
# </table>
# + [markdown] slideshow={"slide_type": "subslide"}
# # Development History
#
# * Started over the Christmas break 1989, by <NAME>
# * Developed in the early 1990s
# * Name comes from Monty Python’s Flying Circus
# * Guido is the Benevolent Dictator for Life (BDFL), meaning that he continues to oversee Python’s development.
# + [markdown] slideshow={"slide_type": "subslide"}
# # Development History
#
# * Open-sourced development from the start (BSD licensed now)
# * http://www.opensource.org/licenses/bsd-license.php
# * Relies on large community input (bugs, patches) and 3rd party add-on software
# * Version 2.0 (2000), 2.6 (2008), 2.7 (2010).
# * Version 2.7.X is reaching end of life this year.
# * Version 3.X (2008) is not backward compatible with 1.X & 2.X. If you're starting now, use 3.X.
# + [markdown] slideshow={"slide_type": "slide"}
# # Why Python
# ## Some of the Alternatives
# I've used almost all of these at some point
# ### C, C++, Fortran
# *Pros: great performance, backbone of legacy scientific computing codes*
#
# `Cons: syntax not optimized for causal programming, no interactive facilities, difficult visualization, text processing, etc. `
#
# ### Mathmatica, Maple, Matlab, IDL (and I guess SAS, SPSS,...)
# *Pros: interactive, great visuals, extensive libraries*
#
# `Cons: costly, proprietary, unpleasant for large-scale programs and non-mathematical tasks.`
#
# ### Perl
# http://strombergers.com/python/
# + [markdown] slideshow={"slide_type": "subslide"}
# # Why Python
# * **Free** (BSD license), highly portable (Linux, OSX, Windows, lots...)
# * **Interactive** interpreter provided.
# * Extremely readable syntax (**“executable pseudo-code”**).
# * **Simple**: non-professional programmers can use it effectively
# * great documentation
# * total abstraction of memory management
# * Clean object-oriented model, but **not mandatory**.
# * Rich built-in types: lists, sets, dictionaries (hash tables), strings, ...
# * Very comprehensive standard library (**batteries included**)
# * Standard libraries for IDL/Matlab-like arrays (NumPy)
# * Easy to wrap existing C, C++ and FORTRAN codes.
# + [markdown] slideshow={"slide_type": "subslide"}
# # Why Python
# ## Amazingly Scalable
# * Interactive experimentation
# * build small, self-contained scripts or million-lines projects.
# * From occasional/novice to full-time use (try that with C++).
# * Large community of open source packages
#
# ## The Kitchen Sink (in a good way)
# * really can do anything you want, with impressive simplicity
#
# ## Performance, if you need it
# * As an interpreted language, Python is slow.
# * But...if you need speed you can do the heavy lifting in C or FORTRAN <br/>...or you can use a Python compiler (e.g., Cython)
# + [markdown] slideshow={"slide_type": "subslide"}
# # My Group Uses Python For
# ## Providing a comprehensive analysis framework for Fermi LAT data
#
# (I was forced into using python...)
#
# * Interface to the low-level (c++) code - Interactive data analysis
# * Scripting
# * Developing new analysis techniques
# * Adding features to static code quickly
# * Providing high-level analysis tools (data selection, statistical testing, simulation development, plot making, and so on and so forth)
# * Validation and Testing
#
#
# # What I Use Python For
# * Data reduction & Analysis
# * processing FITS images quickly
# * wrapping around 3rd party software
# * A Handy & Quick Calculator
# * Prototyping new algorithms/ideas
# * Making plots for papers
# * Notebooking (i.e. making me remember stuff)
# * see the iPython sessions later
# * Writing Presentations (these slides)
# * Controling hardware
# + [markdown] slideshow={"slide_type": "subslide"}
# # Python is everywhere
#
# https://wiki.python.org/moin/OrganizationsUsingPython
#
# # Applications are Numerous
#
# * Scripting and Programing
# * GUI's
# * Web Development
# * Interactive Notebooks (see later)
# * Visualization
# * Parralelization
# * Animation
# * And so on...
# + [markdown] slideshow={"slide_type": "slide"}
# # Firing up the interpreter in OSX
#
# ## Go to Utilities->Terminal
#
# ***
# `[pyuser@pymac ~]$ python`<br/>
# `Python 3.6.7 | packaged by conda-forge | (default, Jul 2 2019, 02:07:37)`<br/>
# `[GCC 4.2.1 Compatible Clang 4.0.1 (tags/RELEASE_401/final)] on darwin`<br/>
# `Type "help", "copyright", "credits" or "license" for more information.`<br/>
# `>>>`<br/>
# ***
# The details might be different (different version, different compiler). You could also use iPython:
# ***
# `[pyuser@pymac ~]$ ipython `<br/>
# `Python 3.6.7 | packaged by conda-forge | (default, Jul 2 2019, 02:07:37)`<br/>
# `Type 'copyright', 'credits' or 'license' for more information`<br/>
# `IPython 7.8.0 -- An enhanced Interactive Python. Type '?' for help.`<br/>
# <br/>
# `In [1]:`<br/>
# ***
# -
# ## Firing it up in other OS's like Windows
#
# Install python via Conda and follow the directions.
# + [markdown] slideshow={"slide_type": "slide"}
# # Creating Python Programs and Scripts
#
# * Basically, any raw text editor will do
# * Lot's of the basic ones will do syntax highlighting (reccomended)
# * You create a python program or script file in the text editor and usually save it with a *.py extension
# * There are lots of programs out there that can do this and have fancy markup.
# * I'm still using emacs
# * List: https://wiki.python.org/moin/PythonEditors
# * Make sure it saves as raw text (and not rich text or something else)
# + [markdown] slideshow={"slide_type": "slide"}
# # Last Thing: The Notebook
#
# * The jupyter Notebook is a powerful tool
# * You **will** want to use it.
# * To start it up from the terminal type
#
# `jupyter notebook`
#
# and a browser window should open that looks like this
#
# 
#
# + slideshow={"slide_type": "slide"}
# %matplotlib inline
from matplotlib import pyplot as plt
import numpy as np
# + slideshow={"slide_type": "fragment"}
plt.xkcd()
plt.figure(figsize=(16,8))
x = np.arange(10)
plt.plot(x,x+0.5*x*x)
plt.xlabel('Years Since Release')
plt.ylabel('Interest in Python')
plt.show()
# -
| 00_Intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Chapter 19 - Metric Predicted Variable with One Nominal Predictor
# - [19.3 - Hierarchical Bayesian Approach](#19.3---Hierarchical-Bayesian-Approach)
# - [19.4 - Adding a Metric Predictor](#19.4---Adding-a-Metric-Predictor)
# - [19.5 - Heterogeneous Variances and Robustness against Outliers](#19.5---Heterogeneous-Variances-and-Robustness-against-Outliers)
# ### Review of Tradition ANOVA
# - ANOVA: Analysis of Variance
# - Analysis means here 'separation': total variance = within-group variance + between-group variance
# - Assumptions:
# 1. normality of each group
# 2. homogeneity of variance: the same variance for all groups
import sys
sys.version
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import pymc3 as pm
import arviz as az
import theano.tensor as tt
import warnings
warnings.filterwarnings("ignore", category=FutureWarning)
from scipy.stats import norm
from IPython.display import Image
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error
# %matplotlib inline
plt.style.use('seaborn-white')
color = '#87ceeb'
# -
# %load_ext watermark
# %watermark -p pandas,numpy,pymc3,theano,matplotlib,seaborn,scipy
# +
def gammaShRaFromModeSD(mode, sd):
"""Calculate Gamma shape and rate from mode and sd."""
rate = (mode + np.sqrt( mode**2 + 4 * sd**2 ) ) / ( 2 * sd**2 )
shape = 1 + mode * rate
return(shape, rate)
def plot_mustache(var, sd, j, axis, width=.75):
for i in np.arange(start=0, stop=len(var), step=int(len(var)*.1)):
rv = norm(loc=var[i], scale=sd[i])
yrange = np.linspace(rv.ppf(0.01), rv.ppf(0.99), 100)
xrange = rv.pdf(yrange)
# When the SD of a group is large compared to others, then the top of its mustache is relatively
# low and does not plot well together with low SD groups.
# Scale the xrange so that the 'height' of the all mustaches is 0.75
xrange_scaled = xrange*(width/xrange.max())
# Using the negative value to flip the mustache in the right direction.
axis.plot(-xrange_scaled+j, yrange, color=color, alpha=.6)
def plot_cred_lines(b0, bj, bcov, x, ax):
"""Plot credible posterior distribution lines for model in section 19.4"""
B = pd.DataFrame(np.c_[b0, bj, bcov], columns=['beta0', 'betaj', 'betacov'])
# Credible posterior prediction lines
# hpd_interval = pm.hpd(B.values, alpha=0.05)
hpd_interval = az.hdi(B.values, hdi_prob=1. - 0.05)
B_hpd = B[B.beta0.between(*hpd_interval[0,:]) &
B.betaj.between(*hpd_interval[1,:]) &
B.betacov.between(*hpd_interval[2,:])]
xrange = np.linspace(x.min()*.95, x.max()*1.05)
for i in np.random.randint(0, len(B_hpd), 10):
ax.plot(xrange, B_hpd.iloc[i,0]+B_hpd.iloc[i,1]+B_hpd.iloc[i,2]*xrange, c=color, alpha=.6, zorder=0)
# -
# ### 19.3 - Hierarchical Bayesian Approach
df = pd.read_csv('data/FruitflyDataReduced.csv', dtype={'CompanionNumber':'category'})
df.info()
df.groupby('CompanionNumber').head(2)
# Count the number of records per nominal group
df.CompanionNumber.value_counts()
# #### Model (Kruschke, 2015)
Image('images/fig19_2.png')
# +
x = df.CompanionNumber.cat.codes.values
y = df.Longevity
yMean = y.mean()
ySD = y.std()
NxLvl = len(df.CompanionNumber.cat.categories)
agammaShRa = gammaShRaFromModeSD(ySD/2, 2*ySD)
with pm.Model() as model1:
aSigma = pm.Gamma('aSigma', agammaShRa[0], agammaShRa[1])
a0 = pm.Normal('a0', yMean, tau=1/(ySD*5)**2)
a = pm.Normal('a', 0.0, tau=1/aSigma**2, shape=NxLvl)
ySigma = pm.Uniform('ySigma', ySD/100, ySD*10)
y = pm.Normal('y', a0 + a[x], tau=1/ySigma**2, observed=y)
# Convert a0,a to sum-to-zero b0,b
m = pm.Deterministic('m', a0 + a)
b0 = pm.Deterministic('b0', tt.mean(m))
b = pm.Deterministic('b', m - b0)
pm.model_to_graphviz(model1)
# -
with model1:
trace1 = pm.sample(3000)
az.plot_trace(trace1);
# #### Figure 19.3 (top)
# +
# Here we plot the metric predicted variable for each group. Then we superimpose the
# posterior predictive distribution
None0 = trace1['m'][:,0]
Pregnant1 = trace1['m'][:,1]
Pregnant8 = trace1['m'][:,2]
Virgin1 = trace1['m'][:,3]
Virgin8 = trace1['m'][:,4]
scale = trace1['ySigma'][:]
fig, ax = plt.subplots(1,1, figsize=(8,5))
ax.set_title('Data with Posterior Predictive Distribution')
sns.swarmplot('CompanionNumber', 'Longevity', data=df, ax=ax);
ax.set_xlim(xmin=-1)
for i, grp in enumerate([None0, Pregnant1, Pregnant8, Virgin1, Virgin8]):
plot_mustache(grp, scale, i, ax)
# -
# #### Contrasts
# +
fig, axes = plt.subplots(2,4, figsize=(15,6))
contrasts = [np.mean([Pregnant1, Pregnant8], axis=0)-None0,
np.mean([Pregnant1, Pregnant8, None0], axis=0)-Virgin1,
Virgin1-Virgin8,
np.mean([Pregnant1, Pregnant8, None0], axis=0)-np.mean([Virgin1, Virgin8], axis=0)]
contrast_titles = ['Pregnant1.Pregnant8 \n vs \n None0',
'Pregnant1.Pregnant8.None0 \n vs \n Virgin1',
'Virgin1 \n vs \n Virgin8',
'Pregnant1.Pregnant8.None0 \n vs \n Virgin1.Virgin8']
for contr, ctitle, ax_top, ax_bottom in zip(contrasts, contrast_titles, fig.axes[:4], fig.axes[4:]):
az.plot_posterior(contr, ref_val=0, color=color, ax=ax_top)
az.plot_posterior(contr/scale, ref_val=0, color=color, ax=ax_bottom)
ax_top.set_title(ctitle)
ax_bottom.set_title(ctitle)
ax_top.set_xlabel('Difference')
ax_bottom.set_xlabel('Effect Size')
fig.tight_layout()
# -
# ### 19.4 - Adding a Metric Predictor
# #### Model (Kruschke, 2015)
Image('images/fig19_4.png')
# +
y = df.Longevity
yMean = y.mean()
ySD = y.std()
xNom = df.CompanionNumber.cat.categories
xMet = df.Thorax
xMetMean = df.Thorax.mean()
xMetSD = df.Thorax.std()
NxNomLvl = len(df.CompanionNumber.cat.categories)
X = pd.concat([df.Thorax, pd.get_dummies(df.CompanionNumber, drop_first=True)], axis=1)
lmInfo = LinearRegression().fit(X, y)
residSD = np.sqrt(mean_squared_error(y, lmInfo.predict(X)))
agammaShRa = gammaShRaFromModeSD(ySD/2, 2*ySD)
with pm.Model() as model2:
aSigma = pm.Gamma('aSigma', agammaShRa[0], agammaShRa[1])
a0 = pm.Normal('a0', yMean, tau=1/(ySD*5)**2)
a = pm.Normal('a', 0.0, tau=1/aSigma**2, shape=NxNomLvl)
aMet = pm.Normal('aMet', 0, tau=1/(2*ySD/xMetSD)**2)
ySigma = pm.Uniform('ySigma', residSD/100, ySD*10)
mu = a0 + a[x] + aMet*(xMet - xMetMean)
y = pm.Normal('y', mu, tau=1/ySigma**2, observed=y)
# Convert a0,a to sum-to-zero b0,b
b0 = pm.Deterministic('b0', a0 + tt.mean(a) + aMet*(-xMetMean))
b = pm.Deterministic('b', a - tt.mean(a))
pm.model_to_graphviz(model2)
# -
with model2:
trace2 = pm.sample(3000)
az.plot_trace(trace2);
# #### Figure 19.5
# +
# Here we plot, for every group, the predicted variable and the metric predictor.
# Superimposed are are the posterior predictive distributions.
fg = sns.FacetGrid(df, col='CompanionNumber', despine=False)
fg.map(plt.scatter, 'Thorax', 'Longevity', facecolor='none', edgecolor='r')
plt.suptitle('Data with Posterior Predictive Distribution', y=1.10, fontsize=15)
for i, ax in enumerate(fg.axes.flatten()):
plot_cred_lines(trace2['b0'],
trace2['b'][:,i],
trace2['aMet'][:],
xMet, ax)
ax.set_xticks(np.arange(.6, 1.1, .1));
# -
# #### Contrasts
# +
None0 = trace2['b'][:,0]
Pregnant1 = trace2['b'][:,1]
Pregnant8 = trace2['b'][:,2]
Virgin1 = trace2['b'][:,3]
Virgin8 = trace2['b'][:,4]
scale = trace2['ySigma']
fig, axes = plt.subplots(2,4, figsize=(15,6))
contrasts = [np.mean([Pregnant1, Pregnant8], axis=0)-None0,
np.mean([Pregnant1, Pregnant8, None0], axis=0)-Virgin1,
Virgin1-Virgin8,
np.mean([Pregnant1, Pregnant8, None0], axis=0)-np.mean([Virgin1, Virgin8], axis=0)]
for contr, ctitle, ax_top, ax_bottom in zip(contrasts, contrast_titles, fig.axes[:4], fig.axes[4:]):
az.plot_posterior(contr, ref_val=0, color=color, ax=ax_top)
az.plot_posterior(contr/scale, ref_val=0, color=color, ax=ax_bottom)
ax_top.set_title(ctitle)
ax_bottom.set_title(ctitle)
ax_top.set_xlabel('Difference')
ax_bottom.set_xlabel('Effect Size')
fig.tight_layout()
# -
# ### 19.5 - Heterogeneous Variances and Robustness against Outliers
df2 = pd.read_csv('data/NonhomogVarData.csv', dtype={'Group':'category'})
df2.info()
df2.groupby('Group').head(3)
# #### Model (Kruschke, 2015)
Image('images/fig19_6.png')
# +
y = df2.Y
x = df2.Group.cat.codes.values
xlevels = df2.Group.cat.categories
NxLvl = len(xlevels)
yMean = y.mean()
ySD = y.std()
aGammaShRa = gammaShRaFromModeSD(ySD/2, 2*ySD)
medianCellSD = df2.groupby('Group').std().dropna().median()
with pm.Model() as model3:
aSigma = pm.Gamma('aSigma', aGammaShRa[0], aGammaShRa[1])
a0 = pm.Normal('a0', yMean, tau=1/(ySD*10)**2)
a = pm.Normal('a', 0.0, tau=1/aSigma**2, shape=NxLvl)
ySigmaSD = pm.Gamma('ySigmaSD', aGammaShRa[0], aGammaShRa[1])
ySigmaMode = pm.Gamma('ySigmaMode', aGammaShRa[0], aGammaShRa[1])
ySigmaRa = (ySigmaMode + np.sqrt(ySigmaMode**2 + 4*ySigmaSD**2))/2*ySigmaSD**2
ySigmaSh = ySigmaMode*ySigmaRa
sigma = pm.Gamma('sigma', ySigmaSh, ySigmaRa, shape=NxLvl)
ySigma = pm.Deterministic('ySigma', tt.maximum(sigma, medianCellSD/1000))
nu_minus1 = pm.Exponential('nu_minus1', 1/29.)
nu = pm.Deterministic('nu', nu_minus1+1)
like = pm.StudentT('y', nu=nu, mu=a0 + a[x], sd=ySigma[x], observed=y)
# Convert a0,a to sum-to-zero b0,b
m = pm.Deterministic('m', a0 + a)
b0 = pm.Deterministic('b0', tt.mean(m))
b = pm.Deterministic('b', m - b0)
pm.model_to_graphviz(model3)
# -
with model3:
# Initializing NUTS with advi since jitter seems to create a problem in this model.
# https://github.com/pymc-devs/pymc3/issues/2897
trace3 = pm.sample(3000, target_accept=0.95, tune=2000)
az.plot_trace(trace3);
# #### Model that assumes equal variances
# +
y = df2.Y
x = df2.Group.cat.codes.values
xlevels = df2.Group.cat.categories
NxLvl = len(xlevels)
yMean = y.mean()
ySD = y.std()
aGammaShRa = gammaShRaFromModeSD(ySD/2, 2*ySD)
with pm.Model() as model3b:
aSigma = pm.Gamma('aSigma', agammaShRa[0], agammaShRa[1])
a0 = pm.Normal('a0', yMean, tau=1/(ySD*5)**2)
a = pm.Normal('a', 0.0, tau=1/aSigma**2, shape=NxLvl)
ySigma = pm.Uniform('ySigma', ySD/100, ySD*10)
y = pm.Normal('y', a0 + a[x], tau=1/ySigma**2, observed=y)
# Convert a0,a to sum-to-zero b0,b
m = pm.Deterministic('m', a0 + a)
b0 = pm.Deterministic('b0', tt.mean(m))
b = pm.Deterministic('b', m - b0)
pm.model_to_graphviz(model3b)
# -
with model3b:
trace3b = pm.sample(3000, cores=4, target_accept=0.95)
pm.traceplot(trace3b);
# #### Figure 19.7
# +
group_a = trace3b['m'][:,0]
group_b = trace3b['m'][:,1]
group_c = trace3b['m'][:,2]
group_d = trace3b['m'][:,3]
scale = trace3b['ySigma']
fig, ax = plt.subplots(1,1, figsize=(8,6))
ax.set_title('Data with Posterior Predictive Distribution\n(Heterogeneous variances)')
sns.swarmplot('Group', 'Y', data=df2, size=5, ax=ax)
ax.set_xlim(xmin=-1);
for i, grp, in enumerate([group_a, group_b, group_c, group_d]):
plot_mustache(grp, scale, i, ax)
# +
fig, axes = plt.subplots(2,2, figsize=(8,6))
contrasts = [group_d-group_a,
group_c-group_b]
contrast_titles = ['D vs A',
'C vs B']
for contr, ctitle, ax_top, ax_bottom in zip(contrasts, contrast_titles, fig.axes[:2], fig.axes[2:]):
az.plot_posterior(contr, ref_val=0, color=color, ax=ax_top)
az.plot_posterior(contr/scale, ref_val=0, color=color, ax=ax_bottom)
ax_top.set_title(ctitle)
ax_bottom.set_title(ctitle)
ax_top.set_xlabel('Difference')
ax_bottom.set_xlabel('Effect Size')
fig.tight_layout()
# -
# #### Figure 19.8
# +
group_a = trace3['m'][:,0]
group_b = trace3['m'][:,1]
group_c = trace3['m'][:,2]
group_d = trace3['m'][:,3]
scale_a = trace3['ySigma'][:,0]
scale_b = trace3['ySigma'][:,1]
scale_c = trace3['ySigma'][:,2]
scale_d = trace3['ySigma'][:,3]
fig, ax = plt.subplots(1,1, figsize=(8,6))
ax.set_title('Data with Posterior Predictive Distribution\n(Heterogeneous variances)')
sns.swarmplot('Group', 'Y', data=df2, size=5, ax=ax)
ax.set_xlim(xmin=-1);
for i, (grp, scale) in enumerate(zip([group_a, group_b, group_c, group_d],
[scale_a, scale_b, scale_c, scale_d])):
plot_mustache(grp, scale, i, ax)
# -
# #### Contrasts
# +
fig, axes = plt.subplots(2,2, figsize=(8,6))
contrasts = [group_d-group_a,
group_c-group_b]
scales = [scale_d**2 + scale_a**2,
scale_c**2 + scale_b**2]
contrast_titles = ['D vs A',
'C vs B']
for contr, scale, ctitle, ax_top, ax_bottom in zip(contrasts, scales, contrast_titles, fig.axes[:2], fig.axes[2:]):
az.plot_posterior(contr, ref_val=0, color=color, ax=ax_top)
az.plot_posterior(contr/(np.sqrt(scale/2)), ref_val=0, color=color, ax=ax_bottom)
ax_top.set_title(ctitle)
ax_bottom.set_title(ctitle)
ax_top.set_xlabel('Difference')
ax_bottom.set_xlabel('Effect Size')
fig.tight_layout()
# -
| Notebooks/Chapter 19.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pyanitools as pyt
import numpy as np
import os
from sklearn import linear_model
import hdnntools as hdt
# -
datadir = '/home/jsmith48/scratch/ccsd_extrapolation/h5files_holdout_split/trainset/delta/'
sae_out = '/home/jsmith48/scratch/ccsd_extrapolation/learning_cases/delta_DZ_retrain/delta_dft_1/sae_linfit.dat'
smap = {'H':0,'C':1,'N':2,'O':3}
Na = len(smap)
files = os.listdir(datadir)
np.random.shuffle(files)
X = []
y = []
for f in files[0:20]:
print(f)
adl = pyt.anidataloader(datadir+f)
for data in adl:
#print(data['path'])
S = data['species']
E = data['energies']
unique, counts = np.unique(S, return_counts=True)
x = np.zeros(Na, dtype=np.float64)
for u,c in zip(unique,counts):
x[smap[u]]=c
for e in E:
X.append(np.array(x))
y.append(np.array(e))
X = np.array(X)
y = np.array(y).reshape(-1,1)
print(X.shape)
print(y.shape)
lin = linear_model.LinearRegression(fit_intercept=False)
lin.fit(X,y)
coef = lin.coef_
print(coef)
# +
sae = open(sae_out,'w')
for i,c in enumerate(coef[0]):
sae.write(next(key for key, value in smap.items() if value == i)+','+str(i)+'='+str(c)+'\n')
sae.close()
# -
| datatools/linear_fitter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# +
import pandas as pd
import numpy as np
from evosim import evosim
# -
import imp
imp.reload(evosim)
# agents = [pd.DataFrame([[0.4, 0.6], [0.6, 0.4]]), pd.DataFrame([[0.4, 0.6], [0.5, 0.5]]), pd.DataFrame([[0.7, 0.3], [0.6, 0.4]])]
agents = [pd.DataFrame([[0.4, 0.4, 0.2], [0.6, 0.4, np.nan]]), pd.DataFrame([[0.4, 0.6], [0.5, 0.5]])]
population = evosim.SimplePopulation(agents)
environment = evosim.MapEnvironment(population, 10)
population.agents[1]
population.mutate()
np.random.choice(population.agents[1][1], p=np.array([.4,.6,np.nan]))
actions = population.draw_actions()
population.agents[1][1]
environment.get_utilities(actions)
np.random.normal(scale=population.agents, size=population.agents.shape)
np.random.normal(scale=population.agents, size=population.agents.shape)
pd.Panel(np.array([[[1,2], [3,4]], [[5,6], [7,8]], [[1,2], [4,5]]]))
s=pd.DataFrame(10, index=np.arange(3), columns=np.arange(4))
s.iloc[1,2]
| evosim.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + Collapsed="false"
import altair as alt
alt.__version__
# + [markdown] Collapsed="false"
# ## Default Altair Renderer
# + Collapsed="false"
import altair as alt
from vega_datasets import data
source = data.movies.url
chart = alt.Chart(source).mark_bar().encode(
alt.X("IMDB_Rating:Q", bin=True),
alt.Y('count()'),
).properties(title = 'IMDB Ratings and Counts').configure_axis(
labelFontSize=20,
titleFontSize=20
).interactive()
chart
# -
# ## Switch to `mimetype`
alt.renderers.enable('mimetype')
chart
# + [markdown] Collapsed="false"
# ## Switch to PNG renderer
# + Collapsed="false"
alt.renderers.enable('png')
chart
# + [markdown] Collapsed="false"
# ## Switch to html renderer
# + Collapsed="false"
alt.renderers.enable('html')
chart
# + [markdown] Collapsed="false"
# ## Switch to Other renderers
# + Collapsed="false"
alt.renderers.enable('zeppelin')
chart
# + Collapsed="false"
alt.renderers.enable('nteract')
chart
# + Collapsed="false"
alt.renderers.enable('svg')
chart
# + Collapsed="false"
alt.renderers.enable('jupyterlab')
chart
# + Collapsed="false"
alt.renderers.enable('kaggle')
chart
# + Collapsed="false"
alt.renderers.enable('colab')
chart
# + Collapsed="false"
| altair_renderer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/BeatHubmann/legendary-giggle/blob/main/PyTorch_Uncertainty_Estimation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="xaodMrOfUiIb"
import torch
import torch.nn as nn
import numpy as np
import matplotlib.pyplot as plt
from sklearn.preprocessing import StandardScaler
# + id="g-fgANGnU94U"
# data will come from a known distribution
# so we can check our answer, duh!
def generate_batch(batch_size=32):
# x in (-5, +5)
x = np.random.random(batch_size)*10 - 5
# sd is a function of x
sd = 0.05 + 0.1 * (x + 5)
# target = mean + noise * sd
y = np.cos(x) - 0.3 * x + np.random.randn(batch_size) * sd
return x, y
# + id="QyAi_ahaWUT4" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="d72c186f-52e4-4d12-96d3-e22b8eb02c28"
# visualize the data
x, y = generate_batch(1024)
plt.scatter(x, y, alpha=0.5);
# + id="DllkLhrPaf7R"
class Model(nn.Module):
def __init__(self):
super().__init__()
self.ann1 = nn.Sequential(
nn.Linear(1, 10),
nn.Tanh(),
nn.Linear(10, 1),
)
self.ann2 = nn.Sequential(
nn.Linear(1, 10),
nn.Tanh(),
nn.Linear(10, 1),
)
def forward(self, inputs):
# returns (mean, log-variance)
return self.ann1(inputs), self.ann2(inputs)
# + id="6onIwzs2aqFr"
model = Model()
# + id="BEDINlYhasAO"
def criterion(outputs, targets):
mu = outputs[0]
v = torch.exp(outputs[1])
# coefficient term
c = torch.log(torch.sqrt(2 * np.pi * v))
# exponent term
f = 0.5 / v * (targets - mu)**2
# mean log-likelihood
nll = torch.mean(c + f)
return nll
# + id="KlamGAs1a1Bz"
optimizer = torch.optim.Adam(model.parameters())
# + id="GAylP2iDa5TY" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="28697fee-3ac8-42ec-ecdd-a3b3fd53f2e2"
n_epochs = 5000
batch_size = 128
losses = np.zeros(n_epochs)
for i in range(n_epochs):
x, y = generate_batch(batch_size)
# conver to torch tensor
inputs = torch.from_numpy(x).float()
targets = torch.from_numpy(y).float()
# reshape data
inputs, targets = inputs.view(-1, 1), targets.view(-1, 1)
# zero grad
optimizer.zero_grad()
# forward pass
outputs = model(inputs)
loss = criterion(outputs, targets)
# store loss
losses[i] = loss.item()
# print loss
if i % 1000 == 0:
print(i, losses[i])
# optimize
loss.backward()
optimizer.step()
# + id="zAsm5ZHsbIPa" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="49a5b198-366b-4cfc-8bd5-134fef5057b6"
plt.plot(losses);
# + id="K5mHeCmZbIxV" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="521c4ce2-90b4-42a2-ca69-cb8ee0931808"
# plot the model predictions
x, y = generate_batch(1024)
plt.scatter(x, y, alpha=0.5)
# conver to torch tensor
inputs = torch.from_numpy(x).float()
targets = torch.from_numpy(y).float()
# reshape data
inputs, targets = inputs.view(-1, 1), targets.view(-1, 1)
with torch.no_grad():
outputs = model(inputs)
yhat = outputs[0].numpy().flatten()
sd = np.exp(outputs[1].numpy().flatten() / 2) # since encoded variance by default
idx = np.argsort(x)
plt.plot(x[idx], yhat[idx], linewidth=3, color='red')
plt.fill_between(x[idx], yhat[idx] - sd[idx], yhat[idx] + sd[idx], color='red', alpha=0.3)
plt.show()
| PyTorch_Uncertainty_Estimation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
## import libaries
import pandas as pd
import numpy as np
import cv2
import os, sys
from tqdm import tqdm
## load data
train = pd.read_csv('train.csv')
test = pd.read_csv('test.csv')
# function to read image
def read_img(img_path):
img = cv2.imread(img_path, cv2.IMREAD_COLOR)
img = cv2.resize(img, (256,256))
return img
## set path for images
TRAIN_PATH = 'train_img/'
TEST_PATH = 'test_img/'
# +
# load data
train_img, test_img = [],[]
for img_path in tqdm(train['image_id'].values):
train_img.append(read_img(TRAIN_PATH + img_path + '.png'))
for img_path in tqdm(test['image_id'].values):
test_img.append(read_img(TEST_PATH + img_path + '.png'))
# -
# normalize images
x_train = np.array(train_img, np.float32) / 255.
x_test = np.array(test_img, np.float32) / 255.
# target variable - encoding numeric value
label_list = train['label'].tolist()
Y_train = {k:v+1 for v,k in enumerate(set(label_list))}
y_train = [Y_train[k] for k in label_list]
y_train = np.array(y_train)
from keras import applications
from keras.models import Model
from keras import optimizers
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
from keras.layers import Conv2D, MaxPooling2D
from keras.layers.normalization import BatchNormalization
from keras.metrics import categorical_accuracy
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import EarlyStopping
from keras.utils import to_categorical
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import ModelCheckpoint
y_train = to_categorical(y_train)
#Transfer learning with Inception V3
base_model = applications.VGG16(weights='imagenet', include_top=False, input_shape=(256, 256, 3))
# +
## set model architechture
add_model = Sequential()
add_model.add(Flatten(input_shape=base_model.output_shape[1:]))
add_model.add(Dense(256, activation='relu'))
add_model.add(Dense(y_train.shape[1], activation='softmax'))
model = Model(inputs=base_model.input, outputs=add_model(base_model.output))
model.compile(loss='categorical_crossentropy', optimizer=optimizers.SGD(lr=1e-4, momentum=0.9),
metrics=['accuracy'])
model.summary()
# +
batch_size = 32 # tune it
epochs = 5 # increase it
train_datagen = ImageDataGenerator(
rotation_range=30,
width_shift_range=0.1,
height_shift_range=0.1,
horizontal_flip=True)
train_datagen.fit(x_train)
# -
history = model.fit_generator(
train_datagen.flow(x_train, y_train, batch_size=batch_size),
steps_per_epoch=x_train.shape[0] // batch_size,
epochs=epochs,
callbacks=[ModelCheckpoint('VGG16-transferlearning.model', monitor='val_acc', save_best_only=True)]
)
## predict test data
predictions = model.predict(x_test)
# get labels
predictions = np.argmax(predictions, axis=1)
rev_y = {v:k for k,v in Y_train.items()}
pred_labels = [rev_y[k] for k in predictions]
## make submission
sub = pd.DataFrame({'image_id':test.image_id, 'label':pred_labels})
sub.to_csv('sub_vgg.csv', index=False) ## ~0.59
| challenge-September/keras_vggmodel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Fitting model to data. We start with fitting to a line. ie Linear Regression
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import pearsonr
from scipy.stats import linregress
import seaborn as sns; sns.set()
# +
data = pd.DataFrame(
[
[5,7,12,16,20],
[4,12,18,21,24]
],
index=['x','y'])
# use numpy to take the transform of data
data_T = data.T
data.T
# -
sns.scatterplot(x='x', y='y', data=data_T)
plt.show()
# +
df_x=data_T.iloc[:,0]
df_y=data_T.iloc[:,1]
slope, intercept, r_value, p_value, std_err = linregress(df_x,df_y)
print(f'The equation of regression line is y={slope:.3f}x+{intercept:.3f}')
# +
X_plot = np.linspace(0,20,100)
Y_plot = slope*X_plot+intercept
sns.scatterplot(x='x', y='y', data=data_T)
plt.plot(X_plot, Y_plot, color='r')
y1 = slope*df_x+intercept
for i in range(len(df_x)):
xx = [df_x[i],df_x[i]]
yy = [df_y[i],y1[i]]
plt.plot(xx,yy, c='g')
plt.show()
# -
# ## How good is the fit? How do we measure goodness of fit?
# +
def mse(actual, pred):
actual, pred = np.array(actual), np.array(pred)
return np.square(np.subtract(actual,pred)).mean()
print(f'mean square error: {mse(df_y, slope*df_x +intercept)}')
# -
# ## mean and standard deviation of the data
data_T.mean()
data_T.std(ddof=1)
# Covariance - std.std
data_T.cov()
# +
## use st.linregress to calculate regression line
# +
import scipy.stats as st
import statsmodels.api as sm
x = data_T['x']
y = data_T['y']
distribution = getattr(st, 'linregress')
params = distribution(x,y)
params
# -
| 04-Linear_Regresion_Python/regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 3D analysis
#
# This tutorial shows how to run a 3D map-based analysis using three example observations of the Galactic center region with CTA.
# ## Setup
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import astropy.units as u
from astropy.coordinates import SkyCoord
from gammapy.extern.pathlib import Path
from gammapy.data import DataStore
from gammapy.irf import EnergyDispersion
from gammapy.maps import WcsGeom, MapAxis, Map
from gammapy.cube import MapMaker, PSFKernel, MapFit
from gammapy.cube.models import SkyModel
from gammapy.spectrum.models import PowerLaw
from gammapy.image.models import SkyGaussian, SkyPointSource
from regions import CircleSkyRegion
# !gammapy info --no-envvar --no-dependencies --no-system
# ## Prepare modeling input data
#
# ### Prepare input maps
#
# We first use the `DataStore` object to access the CTA observations and retrieve a list of observations by passing the observations IDs to the `.obs_list()` method:
# Define which data to use
data_store = DataStore.from_dir("$GAMMAPY_DATA/cta-1dc/index/gps/")
obs_ids = [110380, 111140, 111159]
obs_list = data_store.obs_list(obs_ids)
# Now we define a reference geometry for our analysis, We choose a WCS based gemoetry with a binsize of 0.02 deg and also define an energy axis:
energy_axis = MapAxis.from_edges(
np.logspace(-1., 1., 10), unit="TeV", name="energy", interp="log"
)
geom = WcsGeom.create(
skydir=(0, 0),
binsz=0.02,
width=(10, 8),
coordsys="GAL",
proj="CAR",
axes=[energy_axis],
)
# The `MapMaker` object is initialized with this reference geometry and a field of view cut of 4 deg:
# %%time
maker = MapMaker(geom, offset_max=4. * u.deg)
maps = maker.run(obs_list)
# The maps are prepare by calling the `.run()` method and passing the observation list `obs_list`. The `.run()` method returns a Python `dict` containing a `counts`, `background` and `exposure` map:
print(maps)
# This is what the summed counts image looks like:
counts = maps["counts"].sum_over_axes()
counts.smooth(width=0.1 * u.deg).plot(stretch="sqrt", add_cbar=True, vmax=6);
# And the background image:
background = maps["background"].sum_over_axes()
background.smooth(width=0.1 * u.deg).plot(
stretch="sqrt", add_cbar=True, vmax=6
);
# We can also compute an excess image just with a few lines of code:
excess = Map.from_geom(geom.to_image())
excess.data = counts.data - background.data
excess.smooth(5).plot(stretch="sqrt");
# ### Prepare IRFs
#
# To estimate the mean PSF across all observations at a given source position `src_pos`, we use the `obs_list.make_mean_psf()` method:
# +
# mean PSF
src_pos = SkyCoord(0, 0, unit="deg", frame="galactic")
table_psf = obs_list.make_mean_psf(src_pos)
# PSF kernel used for the model convolution
psf_kernel = PSFKernel.from_table_psf(table_psf, geom, max_radius="0.3 deg")
# -
# To estimate the mean energy dispersion across all observations at a given source position `src_pos`, we use the `obs_list.make_mean_edisp()` method:
# +
# define energy grid
energy = energy_axis.edges * energy_axis.unit
# mean edisp
edisp = obs_list.make_mean_edisp(
position=src_pos, e_true=energy, e_reco=energy
)
# -
# ### Save maps and IRFs to disk
#
# It is common to run the preparation step independent of the likelihood fit, because often the preparation of maps, PSF and energy dispersion is slow if you have a lot of data. We first create a folder:
path = Path("analysis_3d")
path.mkdir(exist_ok=True)
# And the write the maps and IRFs to disk by calling the dedicated `.write()` methods:
# +
# write maps
maps["counts"].write(str(path / "counts.fits"), overwrite=True)
maps["background"].write(str(path / "background.fits"), overwrite=True)
maps["exposure"].write(str(path / "exposure.fits"), overwrite=True)
# write IRFs
psf_kernel.write(str(path / "psf.fits"), overwrite=True)
edisp.write(str(path / "edisp.fits"), overwrite=True)
# -
# ## Likelihood fit
#
# ### Reading maps and IRFs
# As first step we read in the maps and IRFs that we have saved to disk again:
# +
# read maps
maps = {
"counts": Map.read(str(path / "counts.fits")),
"background": Map.read(str(path / "background.fits")),
"exposure": Map.read(str(path / "exposure.fits")),
}
# read IRFs
psf_kernel = PSFKernel.read(str(path / "psf.fits"))
edisp = EnergyDispersion.read(str(path / "edisp.fits"))
# -
# Let's cut out only part of the maps, so that we the fitting step does not take so long:
cmaps = {
name: m.cutout(SkyCoord(0, 0, unit="deg", frame="galactic"), 2 * u.deg)
for name, m in maps.items()
}
cmaps["counts"].sum_over_axes().plot(stretch="sqrt");
# ### Fit mask
#
# To select a certain spatial region and/or energy range for the fit we can create a fit mask:
# +
mask = Map.from_geom(cmaps["counts"].geom)
region = CircleSkyRegion(center=src_pos, radius=0.6 * u.deg)
mask.data = mask.geom.region_mask([region])
mask.get_image_by_idx((0,)).plot();
# -
# In addition we also exclude the range below 0.3 TeV for the fit:
coords = mask.geom.get_coord()
mask.data &= coords["energy"] > 0.3
# ### Model fit
#
# No we are ready for the actual likelihood fit. We first define the model as a combination of a point source with a powerlaw:
spatial_model = SkyPointSource(lon_0="0.01 deg", lat_0="0.01 deg")
spectral_model = PowerLaw(
index=2.2, amplitude="3e-12 cm-2 s-1 TeV-1", reference="1 TeV"
)
model = SkyModel(spatial_model=spatial_model, spectral_model=spectral_model)
# Now we set up the `MapFit` object by passing the prepared maps, IRFs as well as the model:
fit = MapFit(
model=model,
counts=cmaps["counts"],
exposure=cmaps["exposure"],
background=cmaps["background"],
mask=mask,
psf=psf_kernel,
edisp=edisp,
)
# No we run the model fit:
# %%time
result = fit.run(optimize_opts={"print_level": 1})
# ### Check model fit
#
# Finally we check the model fit by cmputing a residual image. For this we first get the number of predicted counts from the fit evaluator:
npred = fit.evaluator.compute_npred()
# And compute a residual image:
residual = Map.from_geom(cmaps["counts"].geom)
residual.data = cmaps["counts"].data - npred.data
residual.sum_over_axes().smooth(width=0.05 * u.deg).plot(
cmap="coolwarm", vmin=-3, vmax=3, add_cbar=True
);
# Apparently our model should be improved by adding a component for diffuse Galactic emission and at least one second point
# source (see exercises at the end of the notebook).
#
# We can also plot the best fit spectrum:
spec = result.model.spectral_model
energy_range = [0.3, 10] * u.TeV
spec.plot(energy_range=energy_range, energy_power=2)
ax = spec.plot_error(energy_range=energy_range, energy_power=2)
# ## Exercises
#
# * Analyse the second source in the field of view: G0.9+0.1
# * Run the model fit with energy dispersion (pass edisp to MapFit)
| tutorials/analysis_3d.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: cafe
# language: python
# name: cafe
# ---
# # Desafio 2
#
# Fazer o gráfico abaixo utilizando um ***loop for***.
#
# <br>
# <br>
#
# 
# ## Dados
# +
x = [1.59, 4.53, 8.16, 10.2, 10.88, 22.68, 22.68, 30.62, 37.42, 45.36, 68.04] # peso em kilos
y = [19.05, 25.4, 26.67, 29.21, 36.83, 52.07, 54.61, 57.15, 60.96, 62.23, 67.31] # altura em em centímetros
raca_cachorro = ['Chihuahua', 'Poodle Toy', 'Pug', 'French Bulldog', 'Beagle', 'Chow Chow', 'Siberian Husky',
'Labrador Retriever', 'German Shepherd Dog', 'Rottweiler', 'Saint Bernard'] # nome da respectiva raça
# -
# ## Importações
import matplotlib.pyplot as plt
import matplotlib as mpl
# ## Constantes
face_color = ['none', 'none', 'none', 'none', 'none', 'none', 'none', 'none', 'k', 'none', 'none']
marker = ['o', 's', 'p', '*', 'v', '^', '<', '>', 'x', 'D', 'H']
mpl.rc('font', family = 'Arial', size=14)
# ## Gráfico de dispersão
plt.figure(figsize=(8,6))
for i in range(len(x)):
plt.scatter(x[i], y[i], label=raca_cachorro[i], edgecolor='k', facecolor=face_color[i],
marker=marker[i], s = 80)
plt.legend(fontsize=12)
plt.xlabel("Peso (kg)", labelpad=15)
plt.ylabel("Altura (cm)", labelpad=15)
plt.title("Relação entre o peso e a altura de diversas raças de cachorros", pad=15)
plt.show()
| curso/grafico-dispersao/desafio-2/Desafio-2-final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbsphinx="hidden"
# This notebook is part of the `nbsphinx` documentation: https://nbsphinx.readthedocs.io/.
# -
# # Prolog and Epilog
#
# When including notebooks in your Sphinx documentation, you can choose to add some generic content before and after each notebook.
# This can be done with the configuration values `nbsphinx_prolog` and `nbsphinx_epilog` in the file `conf.py`.
#
# The prolog and epilog strings can hold arbitrary [reST](https://www.sphinx-doc.org/rest.html) markup.
# Particularly, the [only](https://www.sphinx-doc.org/en/master/usage/restructuredtext/directives.html#directive-only) and [raw](https://docutils.sourceforge.io/docs/ref/rst/directives.html#raw-data-pass-through) directives can be used to have different content for HTML and LaTeX output.
#
# Those strings are also processed by the [Jinja2](https://jinja.palletsprojects.com/) templating engine.
# This means you can run Python-like code within those strings.
# You have access to the current [Sphinx build environment](https://www.sphinx-doc.org/en/master/extdev/envapi.html) via the variable `env`.
# Most notably, you can get the file name of the current notebook with
#
# {{ env.doc2path(env.docname, base=None) }}
#
# Have a look at the [Jinja2 template documentation](https://jinja.palletsprojects.com/templates/) for more information.
#
# <div class="alert alert-warning">
#
# Warning
#
# If you use invalid syntax, you might get an error like this:
#
# jinja2.exceptions.TemplateSyntaxError: expected token ':', got '}'
#
# This is especially prone to happen when using raw LaTeX, with its abundance of braces.
# To avoid clashing braces you can try to insert additional spaces or LaTeX macros that don't have a visible effect, like e.g. `\strut{}`.
# For example, you can avoid three consecutive opening braces with something like that:
#
# \texttt{\strut{}{{ env.doc2path(env.docname, base=None) }}}
#
# NB: The three consecutive closing braces in this example are not problematic.
#
# An alternative work-around would be to surround LaTeX braces with Jinja braces like this:
#
# {{ '{' }}
#
# The string within will not be touched by Jinja.
#
# Another special Jinja syntax is `{%`, which is also often used in fancy TeX/LaTeX code.
# A work-around for this situation would be to use
#
# {{ '{%' }}
#
# </div>
# ## Examples
#
# You can include a simple static string, using [reST](https://www.sphinx-doc.org/rest.html) markup if you like:
#
# ```python
# nbsphinx_epilog = """
# ----
#
# Generated by nbsphinx_ from a Jupyter_ notebook.
#
# .. _nbsphinx: https://nbsphinx.readthedocs.io/
# .. _Jupyter: https://jupyter.org/
# """
# ```
#
# Using some additional Jinja2 markup and the information from the `env` variable, you can create URLs that point to the current notebook file, but located on some other server:
#
# ```python
# nbsphinx_prolog = """
# Go there: https://example.org/notebooks/{{ env.doc2path(env.docname, base=None) }}
#
# ----
# """
# ```
#
# You can also use separate content for HTML and LaTeX output, e.g.:
#
# ```python
# nbsphinx_prolog = r"""
# {% set docname = env.doc2path(env.docname, base=None) %}
#
# .. only:: html
#
# Go there: https://example.org/notebooks/{{ docname }}
#
# .. raw:: latex
#
# \nbsphinxstartnotebook{The following section was created from
# \texttt{\strut{}{{ docname }}}:}
# """
#
# nbsphinx_epilog = r"""
# .. raw:: latex
#
# \nbsphinxstopnotebook{\hfill End of notebook.}
# """
# ```
#
# Note the use of the `\nbsphinxstartnotebook` and `\nbsphinxstopnotebook` commands.
# Those make sure there is not too much space between the "prolog" and the beginning of the notebook and, respectively, between the end of the notebook and the "epilog".
# They also avoid page breaks, in order for the "prolog"/"epilog" not to end up on the page before/after the notebook.
#
# For a more involved example for different HTML and LaTeX versions, see the file [conf.py](conf.py) of the `nbsphinx` documentation.
| doc/prolog-and-epilog.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sqlite3
con = sqlite3.connect('mydatabase.db')
cursor = con.cursor()
# #### Two ways of passing code:
#
# 1. Manually as an integer
# 2. Authomatically as an entity
cursor.execute("CREATE TABLE employees(id integer PRIMARY KEY, name text, salary real, department text, position text, hireDate text)")
con.commit()
cursor.execute("INSERT INTO employees VALUES(5, 'Wieto',600,'IT','Manager','2021-06-22')")
con.commit()
cursor.execute("INSERT INTO employees VALUES(6, 'Mayo',4000,'Data Scientist','Manager','2021-06-22')")
con.commit()
# #### Add information automatically
entities = (9, '<NAME>',1200,'HR ','Assistant Manager','2021-06-22')
cursor.execute("INSERT INTO employees(id,name,salary,department,position,hireDate) VALUES(?,?,?,?,?,?)",entities)
con.commit()
user_id = int(input('Enter your ID: '))
name = input('Enter your name: ')
salary = float(input('Enter your salary: '))
department = input('Enter your department: ')
position = input('Enter your position: ')
date = input('Enter date "YYYY-M-D": ')
entities = (user_id,name,salary,department,position,date)
cursor.execute("INSERT INTO employees(id,name,salary,department,position,hireDate) VALUES(?,?,?,?,?,?)",entities)
con.commit()
print('Data has been inserted!!!')
# +
#user_id = int(input('Enter your ID: '))
#name = input('Enter your name: ')
#salary = float(input('Enter your salary: '))
#department = input('Enter your department: ')
#position = input('Enter your position: ')
#date = input('Enter date "YYYY-M-D": ')
#entities = (user_id,name,salary,department,position,date)
#cursor.execute("INSERT INTO employees(id,name,salary,department,position,hireDate) VALUES(?,?,?,?,?,?)",entities)
#con.commit()
#print('Data has been inserted!!')
# -
# #### Update Name/ Add/ Edit Name and Information in the DataBase
cursor.execute('UPDATE employees SET name = "George" where id = 3')
con.commit()
cursor.execute('UPDATE employees SET salary = 650 where id = 3')
con.commit()
cursor.execute('SELECT * FROM employees')
cursor.execute('SELECT * FROM employees')
rows = cursor.fetchall()
for row in rows:
print(row)
# #### Change more than 1 data in a row
cursor.execute("UPDATE employees SET salary=700,name='Peter' where id = 4")
con.commit()
cursor.execute('SELECT * FROM employees')
rows = cursor.fetchall()
for row in rows:
print(row)
cursor.execute('SELECT salary,hireDate FROM employees')
rows = cursor.fetchall()
for row in rows:
print(row)
# #### Select unique value i.e remove duplicates
cursor.execute('SELECT DISTINCT salary,hireDate FROM employees')
rows = cursor.fetchall()
for row in rows:
print(row)
cursor.execute('SELECT salary,hireDate FROM employees WHERE salary = 700')
rows = cursor.fetchall()
for row in rows:
print(row)
# #### Applying logical Operators
cursor.execute("UPDATE employees SET hireDate='2021-06-20' where id = 1")
con.commit()
cursor.execute('SELECT salary,hireDate FROM employees WHERE salary = 700')
rows = cursor.fetchall()
for row in rows:
print(row)
cursor.execute('SELECT salary,hireDate FROM employees WHERE salary = 700 AND hireDate ="2021-06-20"')
rows = cursor.fetchall()
for row in rows:
print(row)
cursor.execute('SELECT * FROM employees WHERE salary = 700 AND hireDate ="2021-06-20"')
rows = cursor.fetchall()
for row in rows:
print(row)
# #### Sorting/ Arranging and filtering
cursor.execute('SELECT * FROM employees ORDER BY id')
rows = cursor.fetchall()
rows
cursor.execute('SELECT * FROM employees ORDER BY salary')
rows = cursor.fetchall()
rows
cursor.execute('SELECT * FROM employees ORDER BY name')
rows = cursor.fetchall()
rows
# #### Arranging from DESCENDING TO ASCENDING
cursor.execute('SELECT * FROM employees ORDER BY id DESC')
rows = cursor.fetchall()
rows
cursor.execute('SELECT * FROM employees ORDER BY salary DESC')
rows = cursor.fetchall()
rows
cursor.execute('SELECT * FROM employees ORDER BY name DESC')
rows = cursor.fetchall()
rows
# #### How to Delete Data, Name, etc from your system
cursor.execute('DELETE FROM employees WHERE id=9')
con.commit()
cursor.execute('SELECT * FROM employees ORDER BY id DESC')
rows = cursor.fetchall()
rows
# #### Counting Individual records in the Database
cursor.execute('SELECT COUNT(id),name FROM employees')
rows = cursor.fetchall()
rows
cursor.execute('SELECT COUNT(id),department FROM employees GROUP BY department')
rows = cursor.fetchall()
rows
cursor.execute('SELECT * FROM employees WHERE department = "IT" ')
rows = cursor.fetchall()
for row in rows:
print(row)
cursor.execute('SELECT * FROM employees WHERE department = "Data Scientist" ')
rows = cursor.fetchall()
for row in rows:
print(row)
# #### How to delete Table from Database
# +
#cursor.execute('drop table if exists employees')
#con.commit()
# -
# #### Close the database. Once closed, it's unassessible till opened.
con.close()
# #### To continue working on the table file, Open the existing database you are using.
con = sqlite3.connect('mydatabase.db')
| SQL Class.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.3 64-bit ('venv')
# metadata:
# interpreter:
# hash: c1fe9593d27870d231fe1da4a80aa11de54923dcde2f41fcae4a2d40a3260c02
# name: python3
# ---
# +
# Regular Imports
import torch
import matplotlib.pyplot as plt
# Local Imports
import sys
sys.path.insert(0, '..')
from relu_nn.relu_nn import FFReLUNet
# -
| examples/jnotebook_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="Ik0iJ9Rxj0Gm"
import pandas as pd
import numpy as np
import ir_datasets
import math
import logging
from datetime import datetime
import sys
import os
import gzip
import csv
import random
from pathlib import Path
from typing import List, Dict, Tuple, Iterable, Type, Union, Callable
import transformers
from sentence_transformers import models, losses, datasets
from sentence_transformers import LoggingHandler, SentenceTransformer, util, InputExample
from sentence_transformers.evaluation import EmbeddingSimilarityEvaluator, SentenceEvaluator
import torch
from torch import nn, Tensor, device
from torch.optim import Optimizer
from torch.utils.data import DataLoader
from sklearn.model_selection import train_test_split
from fastcore.basics import store_attr
import pytorch_lightning as pl
from pytorch_lightning import Trainer, seed_everything
seed = 0
seed_everything(seed, workers=True)
from pytorch_lightning.loggers import WandbLogger
import config
# -
dataset_name = config.DATASET
save_path = Path(f"data/{dataset_name}")
df = pd.read_pickle(save_path/"data.pkl")
df.head()
df.relevance.value_counts()
# + colab={"base_uri": "https://localhost:8080/", "height": 233} executionInfo={"elapsed": 465, "status": "ok", "timestamp": 1622960820208, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjlU8zpRNrvsulni8P4zzsD1TFpmA3QoAMVIOlZwA=s64", "userId": "06219224739467266869"}, "user_tz": -330} id="xki-ddd7-jny" outputId="f04391e9-3351-414c-ccef-eed8896d1157"
train_size = 0.8
print(len(df))
df_train, df_val = train_test_split(df, train_size=train_size, stratify=df.relevance, random_state=seed)
print(len(df_train), len(df_val))
df_val.head()
# +
train_samples = []
for row in df_train.itertuples():
train_samples.append(InputExample(texts=[row.query_text, row.doc_text], label=row.relevance))
test_samples = []
for row in df_val.itertuples():
test_samples.append(InputExample(texts=[row.query_text, row.doc_text], label=row.relevance))
print(test_samples[0])
# + [markdown] id="McmZsPE3-H-X"
# ## Training
# + id="TQVVRH-GD5ca"
model_name = config.MODEL
train_batch_size = 128 #The larger you select this, the better the results (usually). But it requires more GPU memory
val_batch_size = 128
max_seq_length = 128
num_epochs = 1
# + id="C-qg3IBiGwoZ"
model_name = config.MODEL
model = SentenceTransformer(model_name)
model_name
# + id="71eLlooq8JO9"
class DataModule(pl.LightningDataModule):
def __init__(self, train_batch_size=32, val_batch_size=32):
super().__init__()
self.train_batch_size = train_batch_size
self.val_batch_size = val_batch_size
def prepare_data(self):
self.train_data = train_samples
self.val_data = test_samples
def setup(self, stage=None):
pass
def train_dataloader(self):
train_dataloader = datasets.NoDuplicatesDataLoader(self.train_data, batch_size=self.train_batch_size)
return train_dataloader
def val_dataloader(self):
val_dataloader = datasets.NoDuplicatesDataLoader(self.val_data, batch_size=self.val_batch_size)
return val_dataloader
# + id="tTo2Cc6aEBb3"
class SentenceTransformerModel(pl.LightningModule):
def __init__(self,
loss_model,
max_seq_length: int = 128,
evaluator: SentenceEvaluator = None,
epochs: int = 1,
steps_per_epoch = None,
scheduler: str = 'WarmupLinear',
warmup_steps: int = 10000,
optimizer_class: Type[Optimizer] = transformers.AdamW,
optimizer_params : Dict[str, object]= {'lr': 2e-5},
weight_decay: float = 0.01,
):
super(SentenceTransformerModel, self).__init__()
self.save_hyperparameters()
store_attr("loss_model, epochs, weight_decay, optimizer_class, optimizer_params, steps_per_epoch, scheduler, warmup_steps")
self.loss_model.max_seq_length = max_seq_length
# def on_epoch_start(self):
# print('\n')
def forward(self, features, labels):
loss = self.loss_model(features, labels)
return loss
def training_step(self, data, batch_idx):
features, labels = self.loss_model.model.smart_batching_collate(data)
loss = self.forward(features, labels)
self.log("train_loss", loss, prog_bar=True)
return loss
def validation_step(self, data, batch_idx):
#TODO: dev_evaluator = EmbeddingSimilarityEvaluator.from_input_examples(dev_samples, batch_size=train_batch_size, name='sts-dev')
features, labels = self.loss_model.model.smart_batching_collate(data)
loss = self.forward(features, labels)
# _, preds = torch.max(logits, dim=1)
# val_acc = accuracy_score(preds.cpu(), batch["label"].cpu())
# val_acc = torch.tensor(val_acc)
self.log("val_loss", loss, prog_bar=True)
# self.log("val_acc", val_acc, prog_bar=True)
# return loss
def configure_optimizers(self):
# return torch.optim.Adam(self.parameters(), lr=self.hparams["lr"])
param_optimizer = list(self.loss_model.named_parameters())
no_decay = ['bias', 'LayerNorm.bias', 'LayerNorm.weight']
optimizer_grouped_parameters = [
{'params': [p for n, p in param_optimizer if not any(nd in n for nd in no_decay)], 'weight_decay': self.weight_decay},
{'params': [p for n, p in param_optimizer if any(nd in n for nd in no_decay)], 'weight_decay': 0.0}
]
optimizer = self.optimizer_class(optimizer_grouped_parameters, **self.optimizer_params)
# if self.steps_per_epoch is None or self.steps_per_epoch == 0:
# self.steps_per_epoch = min([len(dataloader) for dataloader in dataloaders])
# num_train_steps = int(self.steps_per_epoch * self.epochs)
# scheduler_obj = self.loss_model.model._get_scheduler(optimizer, scheduler=self.scheduler, warmup_steps=self.warmup_steps, t_total=num_train_steps)
# return [[optimizer], [scheduler_obj]]
return optimizer
# +
from pytorch_lightning.callbacks import ProgressBar, ModelCheckpoint
class LitProgressBar(ProgressBar):
def on_train_epoch_end(self, *args, **kwargs):
super().on_train_epoch_end(*args, **kwargs)
print()
checkpoint_callback = ModelCheckpoint(dirpath="./models", monitor="val_loss", mode="min")
early_stop_callback = EarlyStopping(monitor='val_loss', min_delta=0.00, patience=5, verbose=True, mode='auto')
# -
wandb_logger = WandbLogger(project="lightning-sentence-transformers", name="test", reinit=True)
# + id="w_ckceLKb8Lx"
pl_data = DataModule()
loss_model = losses.MultipleNegativesRankingLoss(model)
steps_per_epoch = 476
stl_model = SentenceTransformerModel(loss_model, steps_per_epoch=steps_per_epoch,)
#TODO: Add learning rate scheduler
trainer = pl.Trainer(
default_root_dir="logs",
gpus=(1 if torch.cuda.is_available() else 0),
max_epochs=10,
fast_dev_run=False,
gradient_clip_val=1.0,
amp_backend='native',
amp_level='O2',
precision=16,
auto_lr_find=True,
auto_scale_batch_size=False,
auto_select_gpus=True,
# callbacks=[LitProgressBar()],
# logger=pl.loggers.TensorBoardLogger("logs/", name=model_name, version=1),
logger=wandb_logger,
deterministic=True,
)
trainer.fit(stl_model, pl_data)
# -
| notebooks/3. train-lightning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Solving Captcha using Tensorflow
# +
# Import all the packages
import cv2
import pickle
import os.path
import time
import matplotlib.pyplot as plt
import numpy as np
import imutils
from imutils import paths
from sklearn.preprocessing import LabelBinarizer
import tensorflow as tf
from tensorflow.python.framework import ops
from helpers import resize_to_fit
train_graph = tf.Graph()
# -
# Store all file names and folder names
LETTER_IMAGES_FOLDER = "extracted_letter_images"
MODEL_LABELS_FILENAME = "model_labels.dat"
TEST_DATA_FOLDER = 'test_captcha'
CHECKPOINT = "./train_model.ckpt"
# ## Getting preprocessed train images and it's labels
# +
# Initialize the data and labels
data = []
labels = []
# loop over the input images
for image_file in paths.list_images(LETTER_IMAGES_FOLDER):
# Load the image and convert it to grayscale
image = cv2.imread(image_file)
image = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
# Resize the letter so it fits in a 20x20 pixel box
image = resize_to_fit(image, 20, 20)
# Add a third channel dimension to the image
image = np.expand_dims(image, axis=2)
# Grab the name of the letter based on the folder it was in
label = image_file.split(os.path.sep)[-2]
# Add the letter image and it's label to our training data
data.append(image)
labels.append(label)
# +
# Scale the raw pixel intensities to the range [0, 1] (this improves training)
data = np.array(data, dtype="float") / 255.0
labels = np.array(np.expand_dims(labels, axis=1))
print(data.shape)
print(labels.shape)
# +
# Convert the labels (letters) into one-hot encodings
lb = LabelBinarizer().fit(labels)
labels = lb.transform(labels)
print(labels.shape)
# -
# Save the mapping from labels to one-hot encodings
# We'll need this later when we use the model to decode what it's predictions mean
with open(MODEL_LABELS_FILENAME, "wb") as f:
pickle.dump(lb, f)
# +
m = data.shape[0] # Number of training examples
n_H = data.shape[1] # Images' height
n_W = data.shape[2] # Images' width
n_C = data.shape[3] # number of channels
n_cls = labels.shape[1] # number of classes
# Create placeholders for the train data and label
with train_graph.as_default():
X = tf.placeholder(tf.float32, [None, n_H, n_W, n_C], name = 'input')
Y = tf.placeholder(tf.float32, [None, n_cls], name = 'output')
# -
# Initialize the weights for the convolution layers
# shape = [filter_size, filter_size, num_input_channels, num_filters]
with train_graph.as_default():
W1 = tf.get_variable("W1", [5, 5, 1, 20], initializer = tf.contrib.layers.xavier_initializer(seed=0))
W2 = tf.get_variable("W2", [5, 5, 20, 50], initializer = tf.contrib.layers.xavier_initializer(seed=0))
# ## CNN Architecture
# Create convolutional neural network
with train_graph.as_default():
# Layer1 - Convolutional
conv_layer1 = tf.nn.conv2d(X, W1, strides=[1, 1, 1, 1], padding='SAME', name = 'conv1')
relu_layer1 = tf.nn.relu(conv_layer1, name = 'relu1')
max_pool_layer1 = tf.nn.max_pool(relu_layer1, ksize = [1, 2, 2, 1], strides = [1, 2, 2, 1], padding='SAME', name = 'pool1')
# Layer2 - Convolutional
conv_layer2 = tf.nn.conv2d(max_pool_layer1, W2, strides=[1, 1, 1, 1], padding='SAME', name = 'conv2')
relu_layer2 = tf.nn.relu(conv_layer2, name = 'relu2')
max_pool_layer2 = tf.nn.max_pool(relu_layer2, ksize = [1, 2, 2, 1], strides = [1, 2, 2, 1], padding='SAME', name = 'pool2')
# Layer3 - Fully_Connected (Don't forget to flatten the previous layer)
flatten_layer3 = tf.contrib.layers.flatten(max_pool_layer2)
fc_layer3 = tf.contrib.layers.fully_connected(flatten_layer3, 500, activation_fn=tf.nn.relu, scope = 'fc1')
# Layer4 - Fully_Connected
fc_layer4 = tf.contrib.layers.fully_connected(fc_layer3, n_cls, activation_fn=None, scope = 'fc2')
print(fc_layer4)
# Use cross entropy cost function
with train_graph.as_default():
cross_entropy = tf.nn.softmax_cross_entropy_with_logits_v2(logits=fc_layer4, labels=Y)
cost = tf.reduce_mean(cross_entropy)
# Use adam optimizer
optimizer = tf.train.AdamOptimizer(learning_rate=1e-4).minimize(cost)
# Funcion: To pick random minibatches to train the model
def random_mini_batches(train, labels, batch_size, seed):
# Always change the seed so that we randomize in different order
np.random.seed(seed)
# Make sure we shuffle both the train data and the label in the same order
p = np.random.permutation(len(train))
train = train[p]
labels = labels[p]
train_batches = []
label_batches = []
# Dividing the train data into minibatches
for batch_i in range(0, len(train)//batch_size):
start_i = batch_i * batch_size
train_batch = train[start_i:start_i + batch_size]
label_batch = labels[start_i:start_i + batch_size]
train_batches.append(train_batch)
label_batches.append(label_batch)
return train_batches, label_batches
# ## Training the model
# +
ops.reset_default_graph()
tf.set_random_seed(1)
# Initialize all the hyperparameters
seed = 3
num_epochs=10
minibatch_size=64
costs = []
# Training the model
with tf.Session(graph=train_graph) as sess:
# Initialize all variables
sess.run(tf.global_variables_initializer())
# If we want to continue training a previous session
# loader = tf.train.import_meta_graph("./" + CHECKPOINT + '.meta')
# loader.restore(sess, CHECKPOINT)
# Loop over number of epochs
for epoch in range(num_epochs):
start_time = time.time()
minibatch_cost = 0
num_minibatches = int(m / minibatch_size)
seed = seed + 1
# Calling the random_mini_batches function to get the batches
train_batches, label_batches = random_mini_batches(data, labels, minibatch_size, seed)
# Now train the model for each of that batches and calculate the minibatch cost
for batch_i in range(num_minibatches):
# Choose the minibatches
minibatch_X = train_batches[batch_i]
minibatch_Y = label_batches[batch_i]
_ , temp_cost = sess.run([optimizer, cost], feed_dict={X:minibatch_X, Y:minibatch_Y})
minibatch_cost += temp_cost / num_minibatches
# Print the cost every 2 epoch
if epoch % 2 == 0:
print("Epoch "+str(epoch)+" completed : Time usage "+str(int(time.time()-start_time))+" seconds")
print("\t- Cost after epoch %i: %f" % (epoch, minibatch_cost))
# Don't forget to save the model
saver = tf.train.Saver()
saver.save(sess, CHECKPOINT)
if epoch % 1 == 0:
costs.append(minibatch_cost)
# plot the cost
plt.plot(np.squeeze(costs))
plt.ylabel('cost')
plt.xlabel('iterations (per tens)')
plt.show()
# Calculate the correct predictions
predict_op = tf.argmax(fc_layer4, 1)
correct_prediction = tf.equal(predict_op, tf.argmax(Y, 1))
# Calculate accuracy for the training data
accuracy = tf.reduce_mean(tf.cast(correct_prediction, "float"))
train_accuracy = accuracy.eval({X: data, Y: labels})
print("Train Accuracy:", train_accuracy)
# -
# Let's check the model on few tesrt data
test_data_files = list(paths.list_images(TEST_DATA_FOLDER))
print(test_data_files)
# ## Preprocessing the test images and making predicitons
# +
# Load up the model labels (so we can translate model predictions to actual letters)
with open(MODEL_LABELS_FILENAME, "rb") as f:
lb = pickle.load(f)
# Ignoring the INFO from the tensorflow
tf.logging.set_verbosity(tf.logging.ERROR)
loaded_graph = tf.Graph()
# loop over the image paths
for image_file in test_data_files:
# Name of the image file is the ground truth for our predictions.
filename = os.path.basename(image_file)
captcha_correct_text = os.path.splitext(filename)[0]
# Load the image and convert it to grayscale
image = cv2.imread(image_file)
image = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
# Add some extra padding around the image
image = cv2.copyMakeBorder(image, 20, 20, 20, 20, cv2.BORDER_REPLICATE)
# threshold the image (convert it to pure black and white)
thresh = cv2.threshold(image, 0, 255, cv2.THRESH_BINARY_INV | cv2.THRESH_OTSU)[1]
# find the contours (continuous blobs of pixels) the image
contours = cv2.findContours(thresh.copy(), cv2.RETR_EXTERNAL, cv2.CHAIN_APPROX_SIMPLE)
# Hack for compatibility with different OpenCV versions
contours = contours[0] if imutils.is_cv2() else contours[1]
letter_image_regions = []
# Now we can loop through each of the four contours and extract the letter
# inside of each one
for contour in contours:
# Get the rectangle that contains the contour
(x, y, w, h) = cv2.boundingRect(contour)
# Compare the width and height of the contour to detect letters that
# are conjoined into one chunk
if w / h > 1.25:
# This contour is too wide to be a single letter!
# Split it in half into two letter regions!
half_width = int(w / 2)
letter_image_regions.append((x, y, half_width, h))
letter_image_regions.append((x + half_width, y, half_width, h))
else:
# This is a normal letter by itself
letter_image_regions.append((x, y, w, h))
# If we found more or less than 6 letters in the captcha, our letter extraction
# didn't work correcly. Skip the image.
if len(letter_image_regions) != 6:
continue
# Sort the detected letter images based on the x coordinate to make sure
# we are processing them from left-to-right so we match the right image
# with the right letter
letter_image_regions = sorted(letter_image_regions, key=lambda x: x[0])
# Create an output image and a list to hold our predicted letters
output = cv2.merge([image] * 3)
predictions = []
# loop over the letters
for n,letter_bounding_box in enumerate(letter_image_regions):
# Grab the coordinates of the letter in the image
x, y, w, h = letter_bounding_box
# Extract the letter from the original image with a 2-pixel margin around the edge
letter_image = image[y - 2:y + h + 2, x - 2:x + w + 2]
# Re-size the letter image to 20x20 pixels to match training data
letter_image = resize_to_fit(letter_image, 20, 20)
# Turn the single image into a 4d list of images so that the Tensorflow can handle
letter_image = np.expand_dims(letter_image, axis=2)
letter_image = np.expand_dims(letter_image, axis=0)
# Load the Tensorflow session
with tf.Session(graph=loaded_graph) as sess:
# Load the saved model
loader = tf.train.import_meta_graph(CHECKPOINT + '.meta')
loader.restore(sess, CHECKPOINT)
# Load the required parameters from the graph
final_layer = loaded_graph.get_tensor_by_name('fc2/BiasAdd:0')
input_layer = loaded_graph.get_tensor_by_name('input:0')
# Making the predicitons
predict = tf.argmax(final_layer, 1)
output = predict.eval({input_layer: letter_image})
# Append the correct letters to a list
predictions.append(lb.classes_[output[0]])
# Let's print our results and determine if it's correct or not
print("Original Captcha - " + captcha_correct_text)
print("Predicted Captcha - " + ''.join(predictions))
if captcha_correct_text == ''.join(predictions):
print("---CORRECT---")
else:
print("---WRONG---")
# Plotting the captcha image as well
plt.imshow(image)
plt.show()
| notebooks/train_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.core.display import HTML
with open('style.css', 'r') as file:
css = file.read()
HTML(css)
# # Refutational Completeness of the Cut Rule
# This notebook implements a number of procedures that are needed in our proof of the <em style="color:blue">refutational completeness</em> of the cut rule.
# The function $\texttt{complement}(l)$ computes the <em style="color:blue">complement</em> of a literal $l$.
# If $p$ is a propositional variable, we have the following:
# <ol>
# <li>$\texttt{complement}(p) = \neg p$,
# </li>
# <li>$\texttt{complement}(\neg p) = p$.
# </li>
# </ol>
def complement(l):
"Compute the complement of the literal l."
if isinstance(l, str): # l is a propositional variable
return ('¬', l)
else: # l = ('¬', 'p')
return l[1] # l[1] = p
complement('p')
complement(('¬', 'p'))
# The function $\texttt{extractVariable}(l)$ extracts the propositional variable from the literal $l$.
# If $p$ is a propositional variable, we have the following:
# <ol>
# <li>$\texttt{extractVariable}(p) = p$,
# </li>
# <li>$\texttt{extractVariable}(\neg p) = p$.
# </li>
# </ol>
def extractVariable(l):
"Extract the variable of the literal l."
if isinstance(l, str): # l is a propositional variable
return l
else: # l = ('¬', 'p')
return l[1]
extractVariable('p')
extractVariable(('¬', 'p'))
# The function $\texttt{collectsVariables}(M)$ takes a set of clauses $M$ as its input and computes the set of all propositional variables occurring in $M$. The clauses in $M$ are represented as sets of literals.
def collectVariables(M):
"Return the set of all variables occurring in M."
return { extractVariable(l) for C in M
for l in C
}
C1 = frozenset({ 'p', 'q', 'r' })
C2 = frozenset({ ('¬', 'p'), ('¬', 'q'), ('¬', 's') })
collectVariables({C1, C2})
# Given two clauses $C_1$ and $C_2$ that are represented as sets of literals, the function `cutRule`$(C_1, C_2)$ computes all clauses that can be derived from $C_1$ and $C_2$ using the *cut rule*. In set notation, the cut rule is the following rule of inference:
# $$
# \frac{\displaystyle \;C_1\cup \{l\} \quad C_2 \cup \bigl\{\overline{\,l\,}\bigr\}}{\displaystyle C_1 \cup C_2}
# $$
def cutRule(C1, C2):
"Return the set of all clauses that can be deduced by the cut rule from C1 and C2."
return { C1 - {l} | C2 - {complement(l) } for l in C1
if complement(l) in C2
}
C1 = frozenset({ 'p', 'q' })
C2 = frozenset({ ('¬', 'p'), ('¬', 'q') })
cutRule(C1, C2)
# In the expression `saturate(Clauses)` below, `Clauses` is a set of *clauses*, where each clause is a set of *literals*. The call `saturate(Clauses)` computes the set of all clauses that can be derived from clauses in the set `Clauses` using the *cut rule*. The function keeps applying the cut rule until either no new clauses can be derived, or the empty clause $\{\}$ is derived. The resulting set of clauses is *saturated* in the following sense: If $C_1$ and $C_2$ are clauses from the set `Clauses` and the clause $D$ can be derived from $C_1$ and $C_2$ via the cut rule, then $D \in \texttt{Clauses}$ or $\{\} \in \texttt{Clauses}$.
def saturate(Clauses):
while True:
Derived = { C for C1 in Clauses
for C2 in Clauses
for C in cutRule(C1, C2)
}
if frozenset() in Derived:
return { frozenset() } # This is the set notation of ⊥.
Derived -= Clauses # remove clauses that were present before
if Derived == set(): # no new clauses have been found
return Clauses
Clauses |= Derived
C1 = frozenset({ 'p', 'q' })
C2 = frozenset({ ('¬', 'p') })
C3 = frozenset({ ('¬', 'p'), ('¬', 'q') })
saturate({C1, C2, C3})
# The function $\texttt{findValuation}(\texttt{Clauses})$ takes a set of clauses as input. The function tries to compute a variable interpretation that makes all of the clauses true. If this is successful, a set of literals is returned. This set of literals does not contain any complementary literals and therefore corresponds to a variable assignment satisfying all clauses. If $\texttt{Clauses}$ is unsatisfiable, <tt>False</tt> is returned.
def findValuation(Clauses):
"Given a set of Clauses, find a propositional valuation satisfying all of these clauses."
Variables = collectVariables(Clauses)
Clauses = saturate(Clauses)
if frozenset() in Clauses: # The set Clauses is inconsistent.
return False
Literals = set()
for p in Variables:
if any(C for C in Clauses
if p in C and C - {p} <= { complement(l) for l in Literals }
):
Literals |= { p }
else:
Literals |= { ('¬', p) }
return Literals
C1 = frozenset({ 'r', 'p', 's' })
C2 = frozenset({ 'r', 's' })
C3 = frozenset({ 'p', 'q', 's' })
C4 = frozenset({ ('¬', 'p'), ('¬', 'q') })
C5 = frozenset({ ('¬', 'p'), 's', ('¬', 'r') })
C6 = frozenset({ 'p', ('¬', 'q'), 'r'})
C7 = frozenset({ ('¬', 'r'), ('¬', 's'), 'q' })
C8 = frozenset({ ('¬', 'p'), ('¬', 's')})
C9 = frozenset({ 'p', ('¬', 'r'), ('¬', 'q') })
C0 = frozenset({ ('¬', 'p'), 'r', 'q', ('¬', 's') })
Clauses = { C0, C1, C2, C3, C4, C5, C6, C7, C8, C9 }
findValuation(Clauses)
| Python/Completeness.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:PythonData]
# language: python
# name: conda-env-PythonData-py
# ---
# +
# Dependencies
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import requests
from census import Census
# Census API Key
from config import api_key
c = Census(api_key, year=2013)
# +
# Run Census Search to retrieve data on all zip codes (2013 ACS5 Census)
# See: https://github.com/CommerceDataService/census-wrapper for library documentation
# See: https://gist.github.com/afhaque/60558290d6efd892351c4b64e5c01e9b for labels
census_data = c.acs5.get(("NAME", "B19013_001E", "B01003_001E", "B01002_001E",
"B19301_001E",
"B17001_002E"), {'for': 'zip code tabulation area:*'})
# Convert to DataFrame
census_pd = pd.DataFrame(census_data)
# Column Reordering
census_pd = census_pd.rename(columns={"B01003_001E": "Population",
"B01002_001E": "Median Age",
"B19013_001E": "Household Income",
"B19301_001E": "Per Capita Income",
"B17001_002E": "Poverty Count",
"NAME": "Name", "zip code tabulation area": "Zipcode"})
# Add in Poverty Rate (Poverty Count / Population)
census_pd["Poverty Rate"] = 100 * \
census_pd["Poverty Count"].astype(
int) / census_pd["Population"].astype(int)
# Final DataFrame
census_pd = census_pd[["Zipcode", "Population", "Median Age", "Household Income",
"Per Capita Income", "Poverty Count", "Poverty Rate"]]
# Visualize
print(len(census_pd))
census_pd.head()
# -
# Save as a csv
# Note to avoid any issues later, use encoding="utf-8"
census_pd.to_csv("census_data.csv", encoding="utf-8", index=False)
| 3/Activities/08-Ins_Census/Solved/Census_Demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Exploring the THREDDS catalog with Unidata's Siphon
#
# Created: 2017-01-18
#
# [Siphon](http://siphon.readthedocs.io/en/latest/) is a Python module for accessing data hosted on a THREDDS data server.
# Siphon works by parsing the catalog XML and exposing it with higher level functions.
#
# In this notebook we will explore data available on the Central & Northern California Ocean Observing System (CeNCOOS) THREDDS. The cell below extracts the catalog information
# +
from siphon.catalog import TDSCatalog
catalog = TDSCatalog("https://thredds.cencoos.org/thredds/catalog.xml")
info = """
Catalog information
-------------------
Base THREDDS URL: {}
Catalog name: {}
Catalog URL: {}
Metadata: {}
""".format(
catalog.base_tds_url, catalog.catalog_name, catalog.catalog_url, catalog.metadata
)
print(info)
# -
# Unfortunately this catalog has no metadata. So let's check what kind of services are available.
for service in catalog.services:
print(service.name)
# And what datasets are there?
print("\n".join(catalog.datasets.keys()))
# It looks like model runs as well as satellite and HFR data. One can also check the catalog refs for more information
print("\n".join(catalog.catalog_refs.keys()))
# +
ref = catalog.catalog_refs["Global"]
[value for value in dir(ref) if not value.startswith("__")]
# +
info = """
Href: {}
Name: {}
Title: {}
""".format(
ref.href, ref.name, ref.title
)
print(info)
# -
# The `follow` method navigates to that catalog `ref` and returns a new `siphon.catalog.TDSCatalog` object for that part of the THREDDS catalog.
# +
cat = ref.follow()
print(type(cat))
# -
# That makes it easier to explore a small subset of the datasets available in the catalog.
# Here are the data from the *Global* subset.
print("\n".join(cat.datasets.keys()))
# Let's extract the `Global 1-km Sea Surface Temperature` dataset from the global `ref`.
# +
dataset = "Global 1-km Sea Surface Temperature (G1SST)"
ds = cat.datasets[dataset]
ds.name, ds.url_path
# -
# Siphon has a `ncss` (NetCDF subset service) access, here is a quote from the documentation:
#
# > This module contains code to support making data requests to
# the NetCDF subset service (NCSS) on a THREDDS Data Server (TDS). This includes
# forming proper queries as well as parsing the returned data.
#
# Let's check if the catalog offers the `NetcdfSubset` in the `access_urls`.
for name, ds in catalog.datasets.items():
if ds.access_urls:
print(name)
# All `access_urls` returned empty.... Maybe that is just a metadata issue because there is `NetcdfSubset` access when navigating in the webpage.
# +
from IPython.display import HTML
iframe = (
'<iframe src="{src}" width="800" height="550" style="border:none;"></iframe>'.format
)
url = "https://thredds.cencoos.org/thredds/catalog.html?dataset=G1_SST_US_WEST_COAST"
HTML(iframe(src=url))
# -
# To finish the post let's check if there is any WMS service available and overlay the data in a slippy (interactive) map.
# +
services = [service for service in catalog.services if service.name == "wms"]
services
# -
# Found only one, let's tease that out and check the URL.
# +
service = services[0]
url = service.base
url
# -
# OWSLib helps to inspect the available layers before plotting. Here we will get the first layer that has G1_SST_US_WEST_COAST on it.
#
# Note, however, we are skipping the discovery step of the `wms` information and hard-coding it instead.
# That is to save time because parsing the URL [http://pdx.axiomalaska.com/ncWMS/wms](http://pdx.axiomalaska.com/ncWMS/wms) takes ~ 10 minutes. See [this](https://github.com/ioos/notebooks_demos/pull/171#issuecomment-271705056) issue for more information.
# +
from owslib.wms import WebMapService
if False:
web_map_services = WebMapService(url)
layer = [
key for key in web_map_services.contents.keys() if "G1_SST_US_WEST_COAST" in key
][0]
wms = web_map_services.contents[layer]
title = wms.title
lon = (wms.boundingBox[0] + wms.boundingBox[2]) / 2.0
lat = (wms.boundingBox[1] + wms.boundingBox[3]) / 2.0
time = wms.defaulttimeposition
else:
layer = "G1_SST_US_WEST_COAST/analysed_sst"
title = "Sea Surface Temperature"
lon, lat = -122.50, 39.50
time = "undefined"
# +
import folium
m = folium.Map(location=[lat, lon], zoom_start=4)
folium.WmsTileLayer(
name="{} at {}".format(title, time),
url=url,
layers=layer,
fmt="image/png",
transparent=True,
).add_to(m)
folium.LayerControl().add_to(m)
m
# -
# Last but not least a static image for the page thumbnail.
# +
from IPython.display import Image
Image(m._to_png())
| jupyterbook/content/code_gallery/data_access_notebooks/2017-01-18-siphon-explore-thredds.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/badcortex/opt4ds/blob/master/rna_folding_opt4ds.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="sd0DKJ04Ukxb"
import shutil
import sys
import os.path
if not shutil.which("pyomo"):
# !pip install -q pyomo
assert(shutil.which("pyomo"))
if not (shutil.which("glpk") or os.path.isfile("glpk")):
if "google.colab" in sys.modules:
# !apt-get install -y -qq glpk-utils
else:
try:
# !conda install -c conda-forge glpk
except:
pass
# + id="cHsSPqxIUtEB"
from timeit import default_timer as timer
import matplotlib.pyplot as plt
from pyomo.environ import *
import pandas as pd
import numpy as np
# + [markdown] id="icajhHbiXASO"
# # The RNA Folding problem
#
# **The Simple RNA Folding problem** : Given the nucleotide sequence *s* of a RNA molecule, find a *nested pairing* that pairs the *maximum* number of nucleotides, compared to any other nested pairing.
#
# ### Integer Linear Programming formulation
# The ILP formulation for the Simple RNA Folding problem will have one binary variable, called $P(i,j)$, for each pair $(i,j)$ of positions in *s*, where $i < j$. The value $P(i,j)$ given by a feasible solution to the ILP formulation whether or not the nucleotide in $i$ of *s* will be paired with the nucleotide in position $j$ of *s*.
#
# 1. **The objective funtion** : $$ Maximize \sum_{i<j} P(i,j) $$
# 2. **The inequalities** : For every pair $(i,j)$ of positions in *s* that do not have complementary characters $$ P(i,j) = 0 $$
# For each position $j$ in *s* $$ \sum_{k < j} P(k,j) + \sum_{k > j} P(j,k) \leq 1 $$ For every choice of four positions $i < i'< j < j'$
# $$ P(i,j)+P(i',j') \leq 1 $$
#
# See more: *Chapter 6, Integer Linear Programming in Computational and Systems Biology: An entry-level text and course, <NAME>*
# + id="pK6yeYAgUv4n"
def solve_ilp_model(rna_sequence):
rna_length = len(rna_sequence)
# Model
model = ConcreteModel()
# Indexes
model.I = RangeSet(1, rna_length-1)
model.J = RangeSet(1, rna_length)
# Decision variables
model.P = Var(model.I, model.J, within=Binary)
# Objective function
model.Obj = Objective(expr=sum(model.P[i,j] for i in model.I for j in model.J if i<j),
sense=maximize)
# Complementary pairs constraint
model.ComPairs = ConstraintList()
for j in model.J:
for i in model.I:
if i < j:
if (rna_sequence[i-1], rna_sequence[j-1]) not in [("A","U"),("U","A"),("C","G"),("G","C")]:
model.ComPairs.add(model.P[i,j] == 0)
# Pairing constraint
model.PairNucs = ConstraintList()
for j in model.J:
expr = 0
for k in range(1, j):
expr += model.P[k,j]
for k in range(j+1, rna_length+1):
expr += model.P[j,k]
model.PairNucs.add(expr <= 1)
# Nesting contraint
model.NestPair = ConstraintList()
for h in range(1, rna_length-2):
for i in range(h+1, rna_length-1):
for j in range(i+1, rna_length):
for k in range(j+1, rna_length+1):
model.NestPair.add(model.P[h,j] + model.P[i,k] <= 1)
# Solve the model
sol = SolverFactory("glpk").solve(model)
time = sol["Solver"].Time
# Print P(i,j)
# for j in model.J:
# for i in model.I:
# if i < j:
# print("P({},{}) = {}".format(i, j, model.P[i,j]()))
# Optimal solution with dot-bracket notation
rna_db_sequence = ["." for i in rna_sequence]
for i in model.I:
for j in model.J:
if i < j:
if model.P[i,j]() == 1:
rna_db_sequence[i-1] = "("
rna_db_sequence[j-1] = ")"
solution = {"Time": time, "ObjValue": model.Obj(), "DBSeq": rna_db_sequence}
return solution
# + [markdown] id="V9XPQypFXfZk"
# ### Dynamic programming: the Nussinov Algorithm
#
# Similarly to the ILP approach, The Nussinov algorithm solves the problem of predicting secondary RNA structures by maximizing base pairs.
#
# "This is achieved by assigning a score to our input structure within an $L×L$ matrix, $N_{ij}$. To do this, for every paired set of nucleotides, we give it a score of $+1$, and for others, $0$. We then attempt to maximize the scores and backtrack on the nucleotides which maximize our overall score. To maximize our base pairs, Nussinov states only 4 possible rules we may use when comparing nucleotides".
#
# **Algorithm:**
# 1. Add unpaired position $i$ onto best substructure for subsequence $i+1,j$
# 2. Add unpaired position $j$ onto best substructure for subsequence $i,j–1$
# 3. Add paired bases $i–j$ to the best substructure for the subsequence $i+j,j–1$
# 4. Combine two optimal substructures $i,k$ and $k+1,j$
#
# $$\gamma(i, j) \;
# max \begin{cases} \gamma(i + 1, j), \\ \gamma(i, j – 1), \\ \gamma(i + 1, j – 1) + \delta(r_i, r_j), \\ max \{\gamma(i, k) + \gamma(k + 1, j) \} \end{cases}$$
#
# where:
#
# $$\delta(i, j) = \begin{cases} 1, & \text{if } x_i – x_j \text{is a pair} \\ 0, & \text{else.} \end{cases}$$
#
#
# See more: [Nussinov algorithm to predict secondary RNA fold structures](https://bayesianneuron.com/2019/02/nussinov-predict-2nd-rna-fold-structure-algorithm/)
#
#
# + id="PBZ0ftFQUz7r"
def couple(pair):
"""
Return True if RNA nucleotides are Watson-Crick base pairs
"""
pairs = {"A":"U", "U":"A", "G":"C", "C":"G"}
# check if pair is couplable
if pair in pairs.items():
return True
return False
def fill(nm, rna):
"""
Fill the matrix as per the Nussinov algorithm
"""
minimal_loop_length = 0
for k in range(1, len(rna)):
for i in range(len(rna) - k):
j = i + k
if j - i >= minimal_loop_length:
down = nm[i+1][j] # 1st rule
left = nm[i][j-1] # 2nd rule
diag = nm[i+1][j-1] + couple((rna[i], rna[j])) # 3rd rule
rc = max([nm[i][t] + nm[t+1][j] for t in range(i, j)]) # 4th rule
nm[i][j] = max(down, left, diag, rc) # max of all
else:
nm[i][j] = 0
return nm
def traceback(nm, rna, fold, i, L):
"""
Traceback through complete Nussinov matrix to find optimial RNA secondary structure solution through max base-pairs
"""
j = L
if i < j:
if nm[i][j] == nm[i+1][j]: # 1st rule
traceback(nm, rna, fold, i + 1, j)
elif nm[i][j] == nm[i][j-1]: # 2nd rule
traceback(nm, rna, fold, i, j - 1)
elif nm[i][j] == nm[i+1][j-1] + couple((rna[i], rna[j])): # 3rd rule
fold.append((i, j))
traceback(nm, rna, fold, i + 1, j - 1)
else:
for k in range(i+1, j-1):
if nm[i][j] == nm[i, k] + nm[k+1][j]: # 4th rule
traceback(nm, rna, fold, i, k)
traceback(nm, rna, fold, k + 1, j)
break
return fold
def dot_write(rna, fold):
dot = ["." for i in range(len(rna))]
for s in fold:
#print(min(s), max(s))
dot[min(s)] = "("
dot[max(s)] = ")"
return "".join(dot)
def init_matrix(rna):
M = len(rna)
# init matrix
nm = np.empty([M, M])
nm[:] = np.NAN
# init diaganols to 0
# few ways to do this: np.fill_diaganol(), np.diag(), nested loop, ...
nm[range(M), range(M)] = 0
nm[range(1, len(rna)), range(len(rna) - 1)] = 0
return nm
# + [markdown] id="swyATOQME_PB"
# Run the following cell to fold RNA sequences with the two methods above. At the end there are graph illustrating how the times and objective values vary with changing string lengths.
#
# + id="PU-xBHyOU6Yg"
if __name__ == "__main__":
# RNA sequence
rna_sequences = [["A","C","U","G","U"],
["A","C","U","G","U","A","C","U","G","U"],
["A","C","U","G","U","A","C","U","G","U","A","C","U","G","U"],
["A","C","U","G","U","A","C","U","G","U","A","C","U","G","U","A","C","U","G","U"]]
obj_value_ilp = []
db_seq_ilp = []
time_ilp = []
obj_value_dp = []
db_seq_dp = []
time_dp = []
for i, rna_sequence in enumerate(rna_sequences):
# Solve ILP model
ilp_sol = solve_ilp_model(rna_sequence)
obj_value_ilp.append(ilp_sol["ObjValue"])
db_seq_ilp.append("".join(ilp_sol["DBSeq"]))
time_ilp.append(ilp_sol["Time"])
print("#{} RNA sequence: {}\nRNA sequence length: {}\n".format(i+1,"".join(rna_sequence), len(rna_sequence)))
print("+ ILP Model")
print("ObjValue: {}\nDBSeq: {}\nTime: {}\n".format(obj_value_ilp[i], db_seq_ilp[i], time_ilp[i]))
# Solve with Nussinov Algorithm
start_time = timer()
nm = init_matrix("".join(rna_sequence))
nm = fill(nm, "".join(rna_sequence))
fold = []
sec = traceback(nm, "".join(rna_sequence), fold, 0, len(rna_sequence) - 1)
end_time = timer()
res = dot_write("".join(rna_sequence), fold)
names = [_ for _ in "".join(rna_sequence)]
df = pd.DataFrame(nm, index=names, columns=names)
obj_value_dp.append(np.nanmax(nm))
db_seq_dp.append(res)
time_dp.append(end_time - start_time)
print("+ Nussinov Algorithm")
print("ObjValue: {}\nDBSeq: {}\nTime: {}\n".format(obj_value_dp[i], db_seq_dp[i], time_dp[i]))
print(df, "\n")
# + id="Ib78wH6DcZ4H" outputId="94c33f44-5eaa-49a2-86c6-92abb96440c7" colab={"base_uri": "https://localhost:8080/", "height": 312}
fig, ax = plt.subplots()
ax.plot([len(rna_sequence) for rna_sequence in rna_sequences], obj_value_ilp,
label='ILP Model', marker=".")
ax.plot([len(rna_sequence) for rna_sequence in rna_sequences], obj_value_dp,
label='Nussinov Algorithm', marker=".")
ax.set_xlabel('RNA sequence length')
ax.set_ylabel('Number of pairs')
ax.set_title("RNA sequence length vs Number of P(i,j)")
ax.legend()
# + id="88hGHv-gnjYd" outputId="1cc7f16d-97c8-43e6-e3fa-e9d0c065fe30" colab={"base_uri": "https://localhost:8080/", "height": 312}
fig, ax = plt.subplots()
ax.plot([len(rna_sequence) for rna_sequence in rna_sequences], time_ilp,
label='ILP Model', marker=".")
ax.plot([len(rna_sequence) for rna_sequence in rna_sequences], time_dp,
label='Nussinov Algorithm', marker=".")
ax.set_xlabel('RNA sequence length')
ax.set_ylabel('Time to solve [ms]')
ax.set_title("RNA sequence length vs Time")
ax.legend()
# + id="_1HxsDZeWMiq" outputId="d02b7dfe-e26a-48ac-8122-11bca4f40c92" colab={"base_uri": "https://localhost:8080/", "height": 166}
from graphviz import Digraph
# Visualize #1 RNA sequence optimal fold
rna_sequence = rna_sequences[0]
db_seq = db_seq_ilp[0]
# Graph creation
ss = Digraph(comment='RNA Secondary structure')
ss.attr(rankdir='LR')
for i in range(len(rna_sequence)):
ss.node(str(i), rna_sequence[i])
for i in range(len(rna_sequence)-1):
ss.edge(str(i), str(i+1))
max_length = []
for i in range(len(db_seq)-1):
for j in range(i+1, len(db_seq)):
if i < j:
if db_seq[i] == "(" and db_seq[j] == ")":
max_length.append((i,j))
for i in range(len(db_seq)-1):
if db_seq[i] == "(" and db_seq[len(db_seq)-i-1] == ")":
ss.edge(str(i), str(len(db_seq)-i-1))
# print(ss.source)
ss
# + [markdown] id="cdVo3VQoMFLx"
# ### Integer Linear Programming formulation with biological enhancements
# The central assumption in RNA folding prediction is that the most *stable* fold is the most *likely fold*. *Base stacking* contributes significantly to the stability of an RNA fold so a more complex model (based on maximum stability) should *encourage* paired nucleotides to be organized into stacks as much as possible.
#
# A matched pair $(i,j)$ in a nested pairing is called *stacked pair* if either $(i+1,j-1)$ or $(i-1,j+1)$ is also a matched pair in the nested pairing. If $(i,j)$ and $(i+1,j-1)$ are stacked pairs, the four positions $(i,i+1,j-1,j)$ is called a *stacked quartet*.
#
# A new binary variable $Q(i,j)$ is introduced to indicate wheter the pair $(i,j)$ is the first pair in a stacked quartet.
#
# 1. **The objective funtion** : $$ Maximize \sum_{i<j} P(i,j) + Q(i,j)$$
# 2. **The inequalities** : For every pair $(i,j)$ of positions in *s* that do not have complementary characters $$ P(i,j) = 0 $$
# For each position $j$ in *s* $$ \sum_{k < j} P(k,j) + \sum_{k > j} P(j,k) \leq 1 $$ For every choice of four positions $i < i'< j < j'$
# $$ P(i,j)+P(i',j') \leq 1 $$
# For each pair $(i,j)$ where $j>i$
# $$ P(i,j) + P(i+1,j-1) - Q(i,j) \leq 1$$
# $$ 2Q(i,j) - P(i,j) - P(i+1,j-1) \leq 0$$
# + id="UeUy8yGasR3Y" outputId="3ac01b58-d780-488a-c067-de7a61bd3352" colab={"base_uri": "https://localhost:8080/", "height": 252}
from pyomo.environ import *
# RNA sequence
rna_sequence = ["A","C","C","A","G","A","G","C","C","U"]
rna_length = len(rna_sequence)
minD = 2
# Print RNA details
print("RNA sequence: {} \nRNA sequence length: {}".format("".join(rna_sequence), rna_length))
# Model
model = ConcreteModel()
# Indexes
model.I = RangeSet(1, rna_length-1)
model.J = RangeSet(1, rna_length)
# Print indexes
# model.I.pprint()
# model.J.pprint()
# Decision variables
model.P = Var(model.I, model.J, within=Binary)
model.Q = Var(model.I, model.J, within=Binary)
# Print decision variables
# model.P.pprint()
# model.Q.pprint()
# Objective function
model.Obj = Objective(expr=sum(model.P[i,j]+model.Q[i,j] for i in model.I for j in model.J if j-i>minD),
sense=maximize)
# Print objective function
# model.Obj.pprint()
# Complementary pairs constraint
model.ComPairs = ConstraintList()
for j in model.J:
for i in model.I:
if j-1 > minD:
if (rna_sequence[i-1], rna_sequence[j-1]) not in [("A","U"),("U","A"),("C","G"),("G","C")]:
model.ComPairs.add(model.P[i,j] == 0)
# Print complementary pairs constraint
# model.ComPairs.pprint()
# Pairing constraint
model.PairNucs = ConstraintList()
for j in model.J:
expr = 0
for k in range(1, j):
expr += model.P[k,j]
for k in range(j+1, rna_length+1):
expr += model.P[j,k]
model.PairNucs.add(expr <= 1)
# Print paired nucleotides constraints
# model.PairNucs.pprint()
# Nesting contraint
model.NestPair = ConstraintList()
for h in range(1, rna_length-2):
for i in range(h+1, rna_length-1):
for j in range(i+1, rna_length):
for k in range(j+1, rna_length+1):
model.NestPair.add(model.P[h,j] + model.P[i,k] <= 1)
# Quartets detection
model.CountQuartet = ConstraintList()
model.IfQuartet = ConstraintList()
model.ThenQuartet = ConstraintList()
for j in model.J:
for i in model.I:
if j-i > minD:
model.IfQuartet.add(model.P[i,j]+model.P[i+1,j-1]-model.Q[i,j] <= 1)
model.ThenQuartet.add(2*model.Q[i,j]-model.P[i+1,j-1]-model.P[i,j] <= 0)
else:
model.CountQuartet.add(model.Q[i,j] == 0)
# Print nesting constraints
# model.NestPair.pprint()
# Solve the model
sol = SolverFactory("glpk").solve(model)
# Basic info about the solution process
for info in sol["Solver"]:
print(info)
# Report solution value
print("Optimal solution value: {}".format(model.Obj()))
# Optimal solution with dot-bracket notation
rna_db_sequence = ["." for i in rna_sequence]
for i in model.I:
for j in model.J:
if i < j:
if model.P[i,j]() == 1:
rna_db_sequence[i-1] = "("
rna_db_sequence[j-1] = ")"
print("Optimal fold dot-bracket notation: {}".format("".join(rna_db_sequence)))
# + [markdown] id="7iW3m9xYWfjg"
# <a rel="license" href="http://creativecommons.org/licenses/by/4.0/"><img alt="Creative Commons License" style="border-width:0" src="https://i.creativecommons.org/l/by/4.0/88x31.png" /></a><br /><span xmlns:dct="http://purl.org/dc/terms/" property="dct:title"><b>The RNA Folding Problem: An ILP Approach,</b></span> by <NAME> is licensed under a <a rel="license" href="http://creativecommons.org/licenses/by/4.0/">Creative Commons Attribution 4.0 International License</a>.<br />Based on a work at <a xmlns:dct="http://purl.org/dc/terms/" href="https://github.com/mathcoding/opt4ds" rel="dct:source">https://github.com/mathcoding/opt4ds</a>.
| rna_folding_opt4ds.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py27]
# language: python
# name: conda-env-py27-py
# ---
# # MIDS - w261 Machine Learning At Scale
# __Course Lead:__ Dr <NAME> (__email__ Jimi via James.Shanahan _AT_ gmail.com)
#
# ## Assignment - HW5 Phase 2
#
#
# ---
# __Name:__ <NAME>, Stan..., <NAME>
# __Class:__ MIDS w261 Winter 2018, Section 1
# __Email:__ <EMAIL>; <EMAIL>
# __StudentId__ 303218617 __End of StudentId__
# __Week:__ 5.5
#
# __NOTE:__ please replace `1234567` with your student id above
# __Due Time:__ HW is due the Thursday of the following week by 8AM (West coast time).
#
# * __HW5 Phase 1__
# This can be done on a local machine (with a unit test on the cloud such as Altiscale's PaaS or on AWS) and is due Thursday, Week 6 by 8AM (West coast time). It will primarily focus on building a unit/systems and for pairwise similarity calculations pipeline (for stripe documents)
#
# * __HW5 Phase 2__
# This will require the Altiscale cluster and will be due Thursday of the following week by 8AM (West coast time).
# The focus of HW5 Phase 2 will be to scale up the unit/systems tests to the Google 5 gram corpus.
# # Datasets
#
# For Phase 2 you will first use the small datasets from phase 1 to systems test your code in the cloud. Then you will test your code on 1 file and then 20 files before running the full (191 file) Google n-gram dataset.
# __Small data for systems tests__
# %%writefile atlas-boon-systems-test.txt
atlas boon 50 50 50
boon cava dipped 10 10 10
atlas dipped 15 15 15
# %%writefile googlebooks-eng-all-5gram-20090715-0-filtered-first-10-lines.txt
A BILL FOR ESTABLISHING RELIGIOUS 59 59 54
A Biography of General George 92 90 74
A Case Study in Government 102 102 78
A Case Study of Female 447 447 327
A Case Study of Limited 55 55 43
A Child's Christmas in Wales 1099 1061 866
A Circumstantial Narrative of the 62 62 50
A City by the Sea 62 60 49
A Collection of Fairy Tales 123 117 80
A Collection of Forms of 116 103 82
# SETUP: __Paths to Main data in HDFS on Altiscale AND OTHER SETTINGS__
TEST_1 = "/user/winegarj/data/1_test"
TEST_20 = "/user/winegarj/data/20_test"
FULL_DATA = "/user/winegarj/data/full"
import os
# USER = !whoami
USER = USER[0]
OUTPUT_PATH_BASE = '/user/{USER}'.format(USER=USER)
# # Set - Up for Phase 2
# Before you can run your simlarity analysis on the full Google n-gram dataset you should confirm that the code your wrote in Phase 1 works on the cloud. In the space below, copy the code for your three jobs from Phase 1 (`buildStripes.py`, `invertedIndex.py`, `similarity.py`) and rerun your atlas-boon systems tests on Altiscale (i.e. ** the cloud**). NOTE: _you may end up modifying this code when you get to 5.7, that's fine._
# ### `buildStripes.py` Note: changed to `buildStripes_v2.py`
# +
# %%writefile buildStripes_v2.py
# #!~/opt/anaconda2/bin/python
# -*- coding: utf-8 -*-
from __future__ import division
import re
import mrjob
import json
from mrjob.protocol import RawProtocol
from mrjob.job import MRJob
from mrjob.step import MRStep
import itertools
class MRbuildStripes(MRJob):
SORT_VALUES = True
def mapper(self, _, line):
fields = line.lower().strip("\n").split("\t")
words = fields[0].split(" ")
occurrence_count = int(fields[1])
for subset in itertools.combinations(sorted(set(words)), 2):
yield subset[0], (subset[1], occurrence_count)
yield subset[1], (subset[0], occurrence_count)
def reducer(self, word, occurrence_counts):
stripe = {}
for other_word, occurrence_count in occurrence_counts:
stripe[other_word] = stripe.get(other_word,0)+occurrence_count
yield word, stripe
if __name__ == '__main__':
MRbuildStripes.run()
# -
# ### `invertedIndex.py` Note: changed to `invertedIndex_v2.py`
# +
# %%writefile invertedIndex_v2.py
# #!/usr/bin/env python
# -*- coding: utf-8 -*-
from __future__ import division
import collections
import sys
import re
import json
import math
import numpy as np
import itertools
import mrjob
from mrjob.protocol import RawProtocol
from mrjob.job import MRJob
from mrjob.step import MRStep
class MRinvertedIndex(MRJob):
#START SUDENT CODE531_INV_INDEX
SORT_VALUES = True
def steps(self):
JOBCONF_STEP = {
'mapreduce.job.output.key.comparator.class': 'org.apache.hadoop.mapred.lib.KeyFieldBasedComparator',
'mapreduce.partition.keycomparator.options': '-k1'
}
return [
MRStep(jobconf=JOBCONF_STEP,
mapper=self.mapper,
reducer=self.reducer)
]
def mapper(self, _, line):
sys.stderr.write("reporter:counter:Mapper Counters,Calls,1\n")
tokens = line.strip().split('\t')
value_dict = json.loads(tokens[1])
term_len = len(value_dict)
for key in value_dict.keys():
yield key, [tokens[0], term_len]
def reducer(self, key, values):
sys.stderr.write("reporter:counter:Reducer Counters,Calls,1\n")
out = []
for value_dict in values:
value_dict[0] = value_dict[0].replace('"','')
out.append(value_dict)
yield key, out
#END SUDENT CODE531_INV_INDEX
if __name__ == '__main__':
MRinvertedIndex.run()
# -
# ### `similarity.py` Note: changed to `similarity_v2.py`
# +
# %%writefile similarity_v2.py
# #!/usr/bin/env python
# -*- coding: utf-8 -*-
from __future__ import division
import sys
import collections
import re
import json
import math
import numpy as np
import itertools
import mrjob
from mrjob.protocol import RawProtocol
from mrjob.protocol import JSONProtocol
from mrjob.job import MRJob
from mrjob.step import MRStep
class MRsimilarity(MRJob):
SORT_VALUES = True
OUTPUT_PROTOCOL = RawProtocol
def steps(self):
JOBCONF_STEP_2 = {
'mapreduce.job.output.key.comparator.class': 'org.apache.hadoop.mapred.lib.KeyFieldBasedComparator',
'mapreduce.partition.keycomparator.options': '-k1,1nr',
"mapreduce.job.reduces": "1",
"SORT_VALUES":True
}
JOBCONF_STEP_1 = {
"mapreduce.job.reduces": "64",
"mapreduce.job.maps": "64",
}
return [
MRStep(jobconf=JOBCONF_STEP_1,
mapper=self.mapper_pair_sim,
reducer=self.reducer_pair_sim
),
MRStep(jobconf=JOBCONF_STEP_2,
reducer=self.reducer_pair_sim2
)
]
def mapper_pair_sim(self, _, line):
sys.stderr.write("reporter:counter:Mapper Counters,Calls,1\n")
line = line.strip()
index, posting = line.split('\t')
posting = json.loads(posting)
posting = dict(posting)
for docs in itertools.combinations(sorted(posting.keys()), 2):
yield (docs, posting[docs[0]], posting[docs[1]]), 1
def reducer_pair_sim(self, key, values):
sys.stderr.write("reporter:counter:Reducer Counters,Calls,1\n")
total = sum(values)
cosine = total/(np.sqrt(key[1])*np.sqrt(key[2]))
jacard = total/(key[1]+key[2]-total)
overlap = total/min(key[1],key[2])
dice = 2*total/(key[1]+key[2])
yield np.mean([cosine, jacard, overlap, dice]), (key[0][0]+' - '+key[0][1],cosine,jacard,overlap,dice)
def reducer_pair_sim2(self, key, values):
sys.stderr.write("reporter:counter:Intermediate Reducer Counters,Calls,1\n")
for value in values:
yield str(key), json.dumps(value)
if __name__ == '__main__':
MRsimilarity.run()
# -
# #### atlas-boon systems test
# !python buildStripes_v2.py \
# -r local atlas-boon-systems-test.txt
# +
# Run in Hadoop
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'tests')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !python buildStripes_v2.py \
# -r hadoop atlas-boon-systems-test.txt \
# --output-dir={OUTPUT_PATH} \
# --no-output
# -
# !hadoop fs -cat {OUTPUT_PATH}/*
# Save into file for processing
# !hadoop fs -cat {OUTPUT_PATH}/* > test_stripes_1
# Testing inverted index
# !python invertedIndex_v2.py \
# -r local test_stripes_1
# +
# Run in Hadoop
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'tests')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !python invertedIndex_v2.py \
# -r hadoop test_stripes_1 \
# --output-dir={OUTPUT_PATH} \
# --no-output
# -
# Save into file for processing
# !hadoop fs -cat {OUTPUT_PATH}/* > test_index_1
# !cat test_index_1
# +
##########################################################
# Pretty print systems tests for generating Inverted Index
##########################################################
import json
for i in range(1,2):
print "—"*100
print "Systems test ",i," - Inverted Index"
print "—"*100
with open("test_index_"+str(i),"r") as f:
lines = f.readlines()
for line in lines:
line = line.strip()
word,stripe = line.split("\t")
stripe = json.loads(stripe)
stripe.extend([["",""] for _ in xrange(3 - len(stripe))])
print "{0:>16} |{1:>16} |{2:>16} |{3:>16}".format((word),
stripe[0][0]+" "+str(stripe[0][1]), stripe[1][0]+" "+str(stripe[1][1]), stripe[2][0]+" "+str(stripe[2][1]))
# -
# Testing similarity metrics
# !python similarity_v2.py \
# -r local test_index_1
# +
# Run in Hadoop
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'tests')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !python similarity_v2.py \
# -r hadoop test_index_1 \
# --output-dir={OUTPUT_PATH} \
# --no-output
# -
# Save into file for processing
# !hadoop fs -cat {OUTPUT_PATH}/* > test_similarities_1
# !cat test_similarities_1
# +
############################################
# Pretty print systems tests
# Note: adjust print formatting if you need to
############################################
import json
for i in range(1,2):
print '—'*110
print "Systems test ",i," - Similarity measures"
print '—'*110
print "{0:>15} |{1:>15} |{2:>15} |{3:>15} |{4:>15} |{5:>15}".format(
"average", "pair", "cosine", "jaccard", "overlap", "dice")
print '-'*110
with open("test_similarities_"+str(i),"r") as f:
lines = f.readlines()
for line in lines:
line = line.strip()
avg,stripe = line.split("\t")
stripe = json.loads(stripe)
print "{0:>15f} |{1:>15} |{2:>15f} |{3:>15f} |{4:>15f} |{5:>15f}".format(float(avg),
stripe[0], float(stripe[1]), float(stripe[2]), float(stripe[3]), float(stripe[4]))
# -
#
#
# #### 10-line systems test
# +
# Build Stripes
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'tests')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !time python buildStripes_v2.py \
# -r hadoop googlebooks-eng-all-5gram-20090715-0-filtered-first-10-lines.txt \
# --output-dir={OUTPUT_PATH} \
# --no-output
# !hadoop fs -cat {OUTPUT_PATH}/* > test_stripes_2
# -
# !cat test_stripes_2
# +
# Build Inverted Index
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'tests')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !time python invertedIndex_v2.py \
# -r hadoop test_stripes_2 \
# --output-dir={OUTPUT_PATH} \
# --no-output
# !hadoop fs -cat {OUTPUT_PATH}/* > test_index_2
# +
# Calculate Similarity
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'tests')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !time python similarity_v2.py \
# -r hadoop test_index_2 \
# --output-dir={OUTPUT_PATH} \
# --no-output
# !hadoop fs -cat {OUTPUT_PATH}/* > test_similarities_2
# +
############################################
# Pretty print systems tests
# Note: adjust print formatting if you need to
############################################
import json
for i in range(1,3):
print '—'*110
print "Systems test ",i," - Similarity measures"
print '—'*110
print "{0:>15} |{1:>15} |{2:>15} |{3:>15} |{4:>15} |{5:>15}".format(
"average", "pair", "cosine", "jaccard", "overlap", "dice")
print '-'*110
with open("test_similarities_"+str(i),"r") as f:
lines = f.readlines()
for line in lines:
line = line.strip()
avg,stripe = line.split("\t")
stripe = json.loads(stripe)
print "{0:>15f} |{1:>15} |{2:>15f} |{3:>15f} |{4:>15f} |{5:>15f}".format(float(avg),
stripe[0], float(stripe[1]), float(stripe[2]), float(stripe[3]), float(stripe[4]))
# -
# # HW5.6 -Google n-grams EDA
#
# Do some EDA on this dataset using mrjob, e.g.,
#
# - A. Longest 5-gram (number of characters)
# - B. Top 10 most frequent words (please use the count information), i.e., unigrams
# - C. 20 Most/Least densely appearing words (count/pages_count) sorted in decreasing order of relative frequency
# - D. Distribution of 5-gram sizes (character length). E.g., count (using the count field) up how many times a 5-gram of 50 characters shows up. Plot the data graphically using a histogram.
# ### HW5.6.1 - A. Longest 5-gram (number of characters)
# +
# %%writefile longest5gram.py
# #!/opt/anaconda2/bin/python
# -*- coding: utf-8 -*-
import re
from datetime import datetime
import sys
import mrjob
from mrjob.protocol import RawProtocol
from mrjob.job import MRJob
from mrjob.step import MRStep
class longest5gram(MRJob):
# SORT_VALUES = True
def mapper(self, _, line):
fields = line.strip("\n").split("\t")
yield len(fields[0]), fields[0]
def reducer_init(self):
self.longest_ngrams = []
self.longest_size = 0
def reducer(self, key, values):
if int(key)> self.longest_size:
self.longest_size = int(key)
self.longest_ngrams = list(values)
elif int(key) == self.longest_size:
self.longest_ngrams = list(self.longest_ngrams)+list(values)
def reducer_final(self):
yield self.longest_size, ";".join(list(self.longest_ngrams))
def reducer_2_init(self):
self.longest_2_ngrams = []
self.longest_2_size = 0
def reducer_2(self, key, values):
if int(key)> self.longest_2_size:
self.longest_2_size = int(key)
self.longest_2_ngrams = list(values)
elif int(key) == self.longest_2_size:
self.longest_2_ngrams = list(self.longest_2_ngrams)+list(values)
def reducer_2_final(self):
yield self.longest_2_size, ";".join(list(self.longest_2_ngrams))
def steps(self):
return [
MRStep(
mapper = self.mapper,
reducer_init = self.reducer_init,
reducer_final = self.reducer_final,
reducer = self.reducer,
jobconf={
"mapreduce.job.reduces": "32",
"stream.num.map.output.key.fields": 1,
"mapreduce.job.output.key.comparator.class" : "org.apache.hadoop.mapred.lib.KeyFieldBasedComparator",
"mapreduce.partition.keycomparator.options":"-k1,1nr",
}
),
MRStep(
reducer_init = self.reducer_2_init,
reducer_final = self.reducer_2_final,
reducer = self.reducer_2,
jobconf={
"mapreduce.job.reduces": "1"
}
)
]
if __name__ == '__main__':
start_time = datetime.now()
longest5gram.run()
end_time = datetime.now()
elapsed_time = end_time - start_time
sys.stderr.write(str(elapsed_time))
# -
# __On test data set:__
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'longest_ngram_10lines')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !time python longest5gram.py \
# -r hadoop googlebooks-eng-all-5gram-20090715-0-filtered-first-10-lines.txt \
# --output-dir={OUTPUT_PATH} \
# --no-output
# !hadoop fs -cat {OUTPUT_PATH}/*
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'longest_ngram_test1')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !time python longest5gram.py \
# -r hadoop hdfs://{TEST_1} \
# --output-dir={OUTPUT_PATH} \
# --no-output
# !hdfs dfs -cat {OUTPUT_PATH}/*
# __ On the 20 files dataset: __
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'longest_ngram_test20')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !time python longest5gram.py \
# -r hadoop hdfs://{TEST_20} \
# --output-dir={OUTPUT_PATH} \
# --no-output
# !hdfs dfs -cat {OUTPUT_PATH}/*
# __On full data set:__
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'longest_full')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !time python longest5gram.py \
# -r hadoop hdfs://{FULL_DATA} \
# --output-dir={OUTPUT_PATH} \
# --no-output
# !hadoop fs -cat {OUTPUT_PATH}/*
# ### Longest 5grams MR stats
#
# ec2_instance_type: m3.xlarge
# num_ec2_instances: 15
#
# __Step 1:__
#
# RUNNING for 107.0s ~= 2 minutes
# Reduce tasks = 16
#
# __Step 2:__
#
# RUNNING for 108.8s ~= 2 minutes
# Reduce tasks = 1
# ### HW5.6.1 - B. Top 10 most frequent words
# +
# %%writefile mostFrequentWords.py
# #!~/anaconda2/bin/python
# -*- coding: utf-8 -*-
import re
import mrjob
from mrjob.protocol import RawProtocol
from mrjob.job import MRJob
from mrjob.step import MRStep
class mostFrequentWords(MRJob):
# START STUDENT CODE 5.6.1.B
SORT_VALUES = True
def steps(self):
JOBCONF_STEP1 = {'mapreduce.job.reduces': '10',
}
JOBCONF_STEP2 = {
'mapreduce.job.output.key.comparator.class': 'org.apache.hadoop.mapred.lib.KeyFieldBasedComparator',
'stream.num.map.output.key.fields':'2',
'stream.map.output.field.separator':'\t',
'mapreduce.partition.keycomparator.options': '-k1,1nr',
'mapreduce.job.reduces': '1',
}
return [
MRStep(#jobconf=JOBCONF_STEP1,
mapper=self.mapper,
combiner=self.combiner,
reducer=self.reducer,
),
MRStep(jobconf=JOBCONF_STEP2,
mapper=self.mapper2,
),
]
def mapper(self, _, line):
words = re.findall(r'[a-z\']+', line.lower())
for word in words:
yield word, 1
def combiner(self, word, counts):
yield word, sum(counts)
def reducer(self, word, counts):
yield word, sum(counts)
def mapper2(self, word, counts):
yield counts, word
# END STUDENT CODE 5.6.1.B
if __name__ == '__main__':
mostFrequentWords.run()
# -
# __On the test data set:__
# !python mostFrequentWords.py \
# -r local googlebooks-eng-all-5gram-20090715-0-filtered-first-10-lines.txt
# +
# Find top 10 most frequent words
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'tests')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !python mostFrequentWords.py \
# -r hadoop hdfs://{TEST_1}/* \
# --output-dir={OUTPUT_PATH} \
# --no-output
# -
# !hadoop fs -cat {OUTPUT_PATH}/* | head -n 10
# __ On the 20 files dataset: __
# +
# Find top 10 most frequent word
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'tests')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !python mostFrequentWords.py \
# -r hadoop hdfs://{TEST_20}/* \
# --output-dir={OUTPUT_PATH} \
# --no-output
# -
# !hadoop fs -cat {OUTPUT_PATH}/* | head -n 10
# __On the full data set:__
# +
# Find top 10 most frequent word
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'tests')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !python mostFrequentWords.py \
# -r hadoop hdfs://{FULL_DATA}/* \
# --output-dir={OUTPUT_PATH} \
# --no-output
# -
# !hadoop fs -cat {OUTPUT_PATH}/* | head -n 10000 > test.output
# !cat test.output | head -n 100
# !cat test.output | tail -n 10
# **Version that excludes stop words**
# +
# %%writefile mostFrequentWords_v2.py
# #!/opt/anaconda2/bin/python
# -*- coding: utf-8 -*-
import re
import mrjob
from mrjob.protocol import RawProtocol
from mrjob.job import MRJob
from mrjob.step import MRStep
class mostFrequentWords(MRJob):
# START STUDENT CODE 5.6.1.B
SORT_VALUES = True
def steps(self):
JOBCONF_STEP1 = {'mapreduce.job.reduces': '10',
}
JOBCONF_STEP2 = {
'mapreduce.job.output.key.comparator.class': 'org.apache.hadoop.mapred.lib.KeyFieldBasedComparator',
'stream.num.map.output.key.fields':2,
'stream.map.output.field.separator':'\t',
'mapreduce.partition.keycomparator.options': '-k1,1nr',
'mapreduce.job.reduces': '1',
}
return [
MRStep(jobconf=JOBCONF_STEP1,
mapper_init=self.mapper_init,
mapper=self.mapper,
combiner=self.combiner,
reducer=self.reducer,
),
MRStep(jobconf=JOBCONF_STEP2,
mapper=self.mapper2,
),
]
def mapper_init(self):
self.stopwords = ['i', 'me', 'my', 'myself', 'we', 'our', 'ours',
'ourselves', 'you', 'your', 'yours', 'yourself',
'yourselves', 'he', 'him', 'his', 'himself', 'she',
'her', 'hers', 'herself', 'it', 'its', 'itself',
'they', 'them', 'their', 'theirs', 'themselves',
'what', 'which', 'who', 'whom', 'this', 'that',
'these', 'those', 'am', 'is', 'are', 'was', 'were',
'be', 'been', 'being', 'have', 'has', 'had', 'having',
'do', 'does', 'did', 'doing', 'a', 'an', 'the', 'and',
'but', 'if', 'or', 'because', 'as', 'until', 'while',
'of', 'at', 'by', 'for', 'with', 'about', 'against',
'between', 'into', 'through', 'during', 'before',
'after', 'above', 'below', 'to', 'from', 'up', 'down',
'in', 'out', 'on', 'off', 'over', 'under', 'again',
'further', 'then', 'once', 'here', 'there', 'when',
'where', 'why', 'how', 'all', 'any', 'both', 'each',
'few', 'more', 'most', 'other', 'some', 'such', 'no',
'nor', 'not', 'only', 'own', 'same', 'so', 'than',
'too', 'very', 's', 't', 'can', 'will', 'just',
'don', 'should', 'now']
def mapper(self, _, line):
words = re.findall(r'[a-z\']+', line.lower())
for word in words:
if word not in self.stopwords:
yield word, 1
def combiner(self, word, counts):
yield word, sum(counts)
def reducer(self, word, counts):
yield word, sum(counts)
def mapper2(self, word, counts):
yield counts, word
# END STUDENT CODE 5.6.1.B
if __name__ == '__main__':
mostFrequentWords.run()
# -
# !python mostFrequentWords_v2.py \
# -r local googlebooks-eng-all-5gram-20090715-0-filtered-first-10-lines.txt
# +
# Find top 10 most frequent words
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'tests')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !python mostFrequentWords_v2.py \
# -r hadoop hdfs://{TEST_1}/* \
# --output-dir={OUTPUT_PATH} \
# --no-output
# -
# !hadoop fs -cat {OUTPUT_PATH}/* | head -n 100
# ### Most frequent words MR stats
#
# ec2_instance_type: m3.xlarge
# num_ec2_instances: 15
#
# __Step 1:__
#
# RUNNING for 590.7s ~= 10 minutes
# Launched map tasks=191
# Launched reduce tasks=57
#
# __Step 2:__
#
# RUNNING for 76.6s
# Launched map tasks=110
# Launched reduce tasks=16
#
# ### HW5.6.1 - C. 20 Most/Least densely appearing words
# +
# %%writefile mostLeastDenseWords.py
# #!/opt/anaconda2/bin/python
# -*- coding: utf-8 -*-
from __future__ import division
import re
import numpy as np
import mrjob
import json
import sys
from mrjob.protocol import RawProtocol
from mrjob.job import MRJob
from mrjob.step import MRStep
class mostLeastDenseWords(MRJob):
# START STUDENT CODE 5.6.1.C
OUTPUT_PROTOCOL = RawProtocol
SORT_VALUES = True
total_page_count = 0
def steps(self):
JOBCONF_STEP1 = {'mapreduce.job.reduces': '10'}
JOBCONF_STEP2 = {
'mapreduce.job.output.key.comparator.class': 'org.apache.hadoop.mapred.lib.KeyFieldBasedComparator',
'stream.num.map.output.key.fields':'2',
'stream.map.output.field.separator':'\t',
'mapreduce.partition.keycomparator.options': '-k2,2nr',
'mapreduce.job.reduces': '1',
}
return [MRStep(jobconf=JOBCONF_STEP1,
mapper=self.mapper,
reducer=self.reducer
),
MRStep(jobconf=JOBCONF_STEP2,
reducer=self.reducer_output)
]
def mapper(self, _, line):
data = line.split("\t")
words = data[0].lower().split()
count = int(data[1])
page_count = int(data[2])
for w in words:
yield w, count
yield "!Total", page_count
def reducer(self, key, data):
yield key, sum(data)
def reducer_output(self, key, data):
if key == "!Total":
self.total_page_count = sum(data)
else:
yield key, str(sum(data)/self.total_page_count)
# END STUDENT CODE 5.6.1.C
if __name__ == '__main__':
mostLeastDenseWords.run()
# -
# __On the test data set:__
# !python mostLeastDenseWords.py \
# -r local googlebooks-eng-all-5gram-20090715-0-filtered-first-10-lines.txt
# +
# Density for 1 file
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'tests')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !time python mostLeastDenseWords.py \
# -r hadoop hdfs://{TEST_1}/* \
# --output-dir={OUTPUT_PATH} \
# --no-output
# -
# !hdfs dfs -cat {OUTPUT_PATH}/*
# __ On the 20 files dataset: __
# __On the full data set:__
# ### Word density MR stats
#
# ec2_instance_type: m3.xlarge
# num_ec2_instances: 15
#
# __Step 1:__
#
# RUNNING for 649.2s ~= 10 minutes
# Launched map tasks=190
# Launched reduce tasks=57
#
# __Step 2:__
#
# RUNNING for 74.4s ~= 1 minute
# Launched map tasks=110
# Launched reduce tasks=20
# ### HW5.6.1 - D. Distribution of 5-gram sizes (character length)
# +
# %%writefile distribution.py
# #!/opt/anaconda2/bin/python
# -*- coding: utf-8 -*-
import mrjob
from mrjob.protocol import RawProtocol
from mrjob.job import MRJob
from mrjob.step import MRStep
class distribution(MRJob):
#### TODO: divide the counts by 1000s to make the graph more readable
#### TODO: split the lengths into buckets <10, <25, <50, <75, <100
def mapper(self, _, line):
fields = line.strip("\n").split("\t")
yield len(fields[0]), int(fields[1])
def combiner(self,length, counts):
yield length, sum(counts)
def reducer(self,length, counts):
yield length, sum(counts)
def reducer_sort(self, key, values):
yield key, list(values)[0]
def steps(self):
return [
MRStep(
mapper = self.mapper,
combiner = self.combiner,
reducer = self.reducer,
jobconf = {
"mapreduce.job.reduces": "4",
}
),
MRStep(
reducer = self.reducer_sort,
jobconf = {
"SORT_VALUES":True,
"mapreduce.job.reduces": "1",
"stream.num.map.output.key.fields": 1,
"mapreduce.job.output.key.comparator.class" : "org.apache.hadoop.mapred.lib.KeyFieldBasedComparator",
"mapreduce.partition.keycomparator.options":"-k1,1nr",
}
)
]
if __name__ == '__main__':
distribution.run()
# -
# __On the test data set:__
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'distributions_10lines')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !time python distribution.py \
# -r hadoop googlebooks-eng-all-5gram-20090715-0-filtered-first-10-lines.txt \
# --output-dir={OUTPUT_PATH} \
# --no-output
# !hadoop fs -cat {OUTPUT_PATH}/* > distributions_10lines.txt
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'distributions_1file')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !python distribution.py \
# -r hadoop hdfs://{TEST_1} \
# --output-dir={OUTPUT_PATH} \
# --no-output
# !hadoop fs -cat {OUTPUT_PATH}/* > distributions_1file.txt
# __ On the 20 files dataset: __
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'distributions_20files')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !time python distribution.py \
# -r hadoop hdfs://{TEST_20} \
# --output-dir={OUTPUT_PATH} \
# --no-output
# !hadoop fs -cat {OUTPUT_PATH}/* > distributions_20files.txt
# __On the full data set:__
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'distributions_full')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !time python distribution.py \
# -r hadoop hdfs://{FULL_DATA} \
# --output-dir={OUTPUT_PATH} \
# --no-output
# !hadoop fs -cat {OUTPUT_PATH}/* > distributions_full.txt
# ### Distribution MRJob stats
#
# __Step 1:__
#
# RUNNING for 157.8s ~= 2.6 minutes
# Launched map tasks=191
# Launched reduce tasks=16
#
# __Step 2:__
#
# RUNNING for 115.0s ~= 2 minutes
# Launched map tasks=139
# Launched reduce tasks=1
# +
# %matplotlib inline
import numpy as np
import pylab as pl
results_A = []
for line in open("distributions_10lines.txt").readlines():
line = line.strip()
X,Y = line.split("\t")
results_A.append([int(X),int(Y)])
items = (np.array(results_A)[::-1].T)
fig = pl.figure(figsize=(17,7))
ax = pl.subplot(111)
width=0.8
ax.bar(range(len(items[0])), items[1], width=width)
ax.set_xticks(np.arange(len(items[0])) + width/2)
ax.set_xticklabels(items[0], rotation=90)
pl.title("Distributions of 5 Gram lengths using 10-line sample")
pl.show()
# +
# %matplotlib inline
import numpy as np
import pylab as pl
results_A = []
for line in open("distributions_1file.txt").readlines():
line = line.strip()
X,Y = line.split("\t")
results_A.append([int(X),int(Y)])
items = (np.array(results_A)[::-1].T)
fig = pl.figure(figsize=(17,7))
ax = pl.subplot(111)
width=0.8
ax.bar(range(len(items[0])), items[1], width=width)
ax.set_xticks(np.arange(len(items[0])) + width/2)
ax.set_xticklabels(items[0], rotation=90)
pl.title("Distributions of 5 Gram lengths using 1 file")
pl.show()
# +
# %matplotlib inline
import numpy as np
import pylab as pl
results_A = []
for line in open("distributions_20files.txt").readlines():
line = line.strip()
X,Y = line.split("\t")
results_A.append([int(X),int(Y)])
items = (np.array(results_A)[::-1].T)
fig = pl.figure(figsize=(17,7))
ax = pl.subplot(111)
width=0.8
ax.bar(range(len(items[0])), items[1], width=width)
ax.set_xticks(np.arange(len(items[0])) + width/2)
ax.set_xticklabels(items[0], rotation=90)
pl.title("Distributions of 5 Gram lengths using 20 files")
pl.show()
# +
# %matplotlib inline
import numpy as np
import pylab as pl
results_A = []
for line in open("distributions_full.txt").readlines():
line = line.strip()
X,Y = line.split("\t")
results_A.append([int(X),int(Y)])
items = (np.array(results_A)[::-1].T)
fig = pl.figure(figsize=(17,7))
ax = pl.subplot(111)
width=0.8
ax.bar(range(len(items[0])), items[1], width=width)
ax.set_xticks(np.arange(len(items[0])) + width/2)
ax.set_xticklabels(items[0], rotation=90)
pl.title("Distributions of 5 Gram lengths using all files")
pl.show()
# -
# ### HW5.6.2 - OPTIONAL: log-log plots (PHASE 2)
#
# Plot the log-log plot of the frequency distributuion of unigrams. Does it follow power law distribution?
#
# For more background see:
# - https://en.wikipedia.org/wiki/Log%E2%80%93log_plot
# - https://en.wikipedia.org/wiki/Power_law
# # HW5.7 - Synonym detection over 2Gig of Data with extra Preprocessing steps (HW5.3-5 plus some preprocessing)
#
# For the remainder of this assignment please feel free to eliminate stop words from your analysis (see stopWords in the cell below)
#
# __A large subset of the Google n-grams dataset as was described above__
#
# For each HW 5.6 - 5.7.1 Please unit test and system test your code with respect
# to SYSTEMS TEST DATASET and show the results.
# Please compute the expected answer by hand and show your hand calculations for the
# SYSTEMS TEST DATASET. Then show the results you get with your system.
#
# In this part of the assignment we will focus on developing methods for detecting synonyms, using the Google 5-grams dataset. At a high level:
#
#
# 1. remove stopwords
# 2. get 10,000 most frequent
# 3. get 1000 (9001-10000) features
# 3. build stripes
#
# To accomplish this you must script two main tasks using MRJob:
#
#
# __TASK (1)__ Build stripes for the most frequent 10,000 words using cooccurence information based on
# the words ranked from 9001,-10,000 as a basis/vocabulary (drop stopword-like terms),
# and output to a file in your bucket on s3 (bigram analysis, though the words are non-contiguous).
#
#
# __TASK (2)__ Using two (symmetric) comparison methods of your choice
# (e.g., correlations, distances, similarities), pairwise compare
# all stripes (vectors), and output to a file in your bucket on s3.
#
#
# For this task you will have to determine a method of comparison.
# Here are a few that you might consider:
#
# - Jaccard
# - Cosine similarity
# - Spearman correlation
# - Euclidean distance
# - Taxicab (Manhattan) distance
# - Shortest path graph distance (a graph, because our data is symmetric!)
# - Pearson correlation
# - Kendall correlation
#
# However, be cautioned that some comparison methods are more difficult to
# parallelize than others, and do not perform more associations than is necessary,
# since your choice of association will be symmetric.
#
# Please report the size of the cluster used and the amount of time it takes to run for the index construction task and for the synonym calculation task. How many pairs need to be processed (HINT: use the posting list length to calculate directly)? Report your Cluster configuration!
stopwords = ['i', 'me', 'my', 'myself', 'we', 'our', 'ours',
'ourselves', 'you', 'your', 'yours', 'yourself',
'yourselves', 'he', 'him', 'his', 'himself', 'she',
'her', 'hers', 'herself', 'it', 'its', 'itself',
'they', 'them', 'their', 'theirs', 'themselves',
'what', 'which', 'who', 'whom', 'this', 'that',
'these', 'those', 'am', 'is', 'are', 'was', 'were',
'be', 'been', 'being', 'have', 'has', 'had', 'having',
'do', 'does', 'did', 'doing', 'a', 'an', 'the', 'and',
'but', 'if', 'or', 'because', 'as', 'until', 'while',
'of', 'at', 'by', 'for', 'with', 'about', 'against',
'between', 'into', 'through', 'during', 'before',
'after', 'above', 'below', 'to', 'from', 'up', 'down',
'in', 'out', 'on', 'off', 'over', 'under', 'again',
'further', 'then', 'once', 'here', 'there', 'when',
'where', 'why', 'how', 'all', 'any', 'both', 'each',
'few', 'more', 'most', 'other', 'some', 'such', 'no',
'nor', 'not', 'only', 'own', 'same', 'so', 'than',
'too', 'very', 's', 't', 'can', 'will', 'just',
'don', 'should', 'now']
# __STEP 1: Code and Steps for Preprocessing__
# +
# Remove stop words, get 10,000 most frequent words
# Find top 10,000 most frequent words
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'ten_thousand_1')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !python mostFrequentWords_v2.py \
# -r hadoop hdfs://{TEST_1}/* \
# --output-dir={OUTPUT_PATH} \
# --no-output
# -
# !hadoop fs -cat {OUTPUT_PATH}/* | head -n 10000 > ten_thousand_1.dat
# +
# Find top 10,000 most frequent words
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'ten_thousand_20')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !python mostFrequentWords_v2.py \
# -r hadoop hdfs://{TEST_20}/* \
# --output-dir={OUTPUT_PATH} \
# --no-output
# -
# !hadoop fs -cat {OUTPUT_PATH}/* | head -n 10000 > ten_thousand_20.dat
# !cat ten_thousand_20.dat | head
# !cat ten_thousand_20.dat | tail
# +
# Find top 10,000 most frequent words
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'ten_thousand_FULL')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !python mostFrequentWords_v2.py \
# -r hadoop hdfs://{FULL_DATA}/* \
# --output-dir={OUTPUT_PATH} \
# --no-output
# -
# !hadoop fs -cat {OUTPUT_PATH}/* | head -n 10000 > ten_thousand_FULL.dat
# !cat ten_thousand_FULL.dat | head
# !cat ten_thousand_FULL.dat | tail
# Also create features
# !cat ten_thousand_1.dat |tail -n 999 > features_1.dat
# !cat ten_thousand_20.dat |tail -n 999 > features_20.dat
# !cat ten_thousand_FULL.dat |tail -n 999 > features_FULL.dat
# **We now have files with 10K words => `ten_thousand_*.dat` AND
# features with words from 9,001 to 10,000 in => `features_*.dat` **
### Some extra preprocessing to load files faster
import json
files = ['features_1','features_20', 'features_FULL', 'ten_thousand_1', 'ten_thousand_20','ten_thousand_FULL']
for fileName in files:
with open(fileName+'.dat') as f:
words = []
for line in f:
word = line.split("\t")[1].strip('"')
words.append(word)
with open(fileName+'.json', 'w') as outfile:
json.dump(words, outfile)
# __STEP 2: MODIFY BUILD STRIPES TO ONLY INCLUDE 10K words with 1K as FEATURES__
# +
# %%writefile buildStripes_stopwords_1.py
# #!~/opt/anaconda2/bin/python
# -*- coding: utf-8 -*-
from __future__ import division
import re
import mrjob
import json
from mrjob.protocol import RawProtocol
from mrjob.job import MRJob
from mrjob.step import MRStep
import itertools
class MRbuildStripes(MRJob):
SORT_VALUES = True
def steps(self):
return [
MRStep(
mapper_init=self.mapper_init,
mapper=self.mapper,
reducer=self.reducer,
jobconf = {
"mapreduce.job.reduces": "64",
"mapreduce.job.maps": "64",
# "SORT_VALUES":True
}
),
MRStep(
reducer=self.reducer_2,
jobconf = {
"mapreduce.job.reduces": "1",
"SORT_VALUES":True
}
)
]
def mapper_init(self):
self.idx = 9001 # To define when feature set starts
self.filename = 'ten_thousand_1.json'
self.top_words = []
self.features = []
#with open('features_20.json', 'r') as infile:
# self.features = json.loads(infile.read())
with open(self.filename, 'r') as infile:
self.top_words = json.loads(infile.read())
self.features =self.top_words[self.idx:]
def mapper(self, _, line):
fields = line.lower().strip("\n").split("\t")
words = fields[0].split(" ")
occurrence_count = int(fields[1])
filtered_words = [word.decode('utf-8', 'ignore') for word in words if word.decode('utf-8', 'ignore') in self.top_words]
for subset in itertools.combinations(sorted(set(filtered_words)), 2):
if subset[0] in self.top_words and subset[1] in self.features:
yield subset[0], (subset[1], occurrence_count)
if subset[1] in self.top_words and subset[0] in self.features:
yield subset[1], (subset[0], occurrence_count)
def reducer(self, word, occurrence_counts):
stripe = {}
for other_word, occurrence_count in occurrence_counts:
stripe[other_word] = stripe.get(other_word,0)+occurrence_count
yield word, stripe
def reducer_2(self, key, values):
yield str(key), list(values)[0]
if __name__ == '__main__':
MRbuildStripes.run()
# -
# ### HW5.7.1 Running on 1 file
# +
TEST_1 = "/user/winegarj/data/1_test"
TEST_20 = "/user/winegarj/data/20_test"
FULL_DATA = "/user/winegarj/data/full"
import os
# USER = !whoami
USER = USER[0]
OUTPUT_PATH_BASE = '/user/{USER}'.format(USER=USER)
# +
# Run in Hadoop
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'tests')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !time python buildStripes_stopwords_1.py \
# -r hadoop hdfs://{TEST_1}/* \
# --file ten_thousand_1.json \
# --output-dir={OUTPUT_PATH} \
# --no-output
# +
####NOT USED, BUT KEEP JUST IN CASE
import os
from os import listdir
from os.path import isfile, join
def totalOrderSort(myPath, outFileName):
wordsFiles = {}
words = []
for f in listdir(myPath):
if isfile(join(myPath,f)) and "part" in f:
with open(join(myPath,f)) as openFile:
word = f.readline().split("\t")[0]
words.append(word)
wordsFiles[word]= join(mypath,f)
print wordsFiles
print words
# -
# Look at data
# # !hadoop fs -cat {OUTPUT_PATH}/*
# Save into file for processing
# !hadoop fs -cat {OUTPUT_PATH}/* > google_stripes_1
# +
# Run in Hadoop
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'tests')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !time python invertedIndex_v2.py \
# -r hadoop google_stripes_1 \
# --output-dir={OUTPUT_PATH} \
# --no-output
# -
# Save into file for processing
# !hadoop fs -cat {OUTPUT_PATH}/* > google_index_1
# !cat google_index_1 | head
# +
##########################################################
# Pretty print systems tests for generating Inverted Index
##########################################################
import json
for i in range(1,2):
print "—"*100
print "Systems test ",i," - Inverted Index"
print "—"*100
with open("google_index_"+str(i)+"","r") as f:
lines = f.readlines()
for line in lines:
line = line.strip()
word,stripe = line.split("\t")
stripe = json.loads(stripe)
stripe.extend([["",""] for _ in xrange(3 - len(stripe))])
print "{0:>16} |{1:>16} |{2:>16} |{3:>16}".format((word),
stripe[0][0]+" "+str(stripe[0][1]), stripe[1][0]+" "+str(stripe[1][1]), stripe[2][0]+" "+str(stripe[2][1]))
# +
# Run in Hadoop
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'tests')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !time python similarity_v2.py \
# -r hadoop google_index_1 \
# --output-dir={OUTPUT_PATH} \
# --no-output
# -
# Save into file for processing
# !hadoop fs -cat {OUTPUT_PATH}/* > google_similarities_1
# !cat google_similarities_1 | head
# +
############################################
# Pretty print systems tests
# Note: adjust print formatting if you need to
############################################
import json
for i in range(1,2):
print '—'*110
print "Systems test ",i," - Similarity measures"
print '—'*110
print "{0:>21} | {1:>15} |{2:>15} | {3:>15} | {4:>15} | {5:>15}".format("pair",
"cosine", "jaccard", "overlap", "dice", "average")
print '-'*110
with open("google_similarities_"+str(i),"r") as f:
lines = f.readlines()
for line in lines:
line = line.strip()
avg,stripe = line.split("\t")
stripe = json.loads(stripe)
print "{0:>21} | {1:>15f} |{2:>15f} |{3:>15f} | {4:>15f} | {5:>15f} ".format(stripe[0],
float(stripe[1]), float(stripe[2]), float(stripe[3]), float(stripe[4]), float(avg))
# +
# ADD CELLS AS NEEDED
# -
# ### HW5.7.2 Running on 20 test files
# +
# %%writefile buildStripes_stopwords_20.py
# #!~/opt/anaconda2/bin/python
# -*- coding: utf-8 -*-
from __future__ import division
import re
import mrjob
import json
from mrjob.protocol import RawProtocol
from mrjob.job import MRJob
from mrjob.step import MRStep
import itertools
class MRbuildStripes(MRJob):
SORT_VALUES = True
def steps(self):
return [
MRStep(
mapper_init=self.mapper_init,
mapper=self.mapper,
reducer=self.reducer,
jobconf = {
"mapreduce.job.reduces": "64",
"mapreduce.job.maps": "64",
# "SORT_VALUES":True
}
),
MRStep(
reducer=self.reducer_2,
jobconf = {
"mapreduce.job.reduces": "1",
"SORT_VALUES":True
}
)
]
def mapper_init(self):
self.idx = 9001 # To define when feature set starts
self.filename = 'ten_thousand_20.json'
self.top_words = []
self.features = []
#with open('features_20.json', 'r') as infile:
# self.features = json.loads(infile.read())
with open(self.filename, 'r') as infile:
self.top_words = json.loads(infile.read())
self.features =self.top_words[self.idx:]
def mapper(self, _, line):
fields = line.lower().strip("\n").split("\t")
words = fields[0].split(" ")
occurrence_count = int(fields[1])
filtered_words = [word.decode('utf-8', 'ignore') for word in words if word.decode('utf-8', 'ignore') in self.top_words]
for subset in itertools.combinations(sorted(set(filtered_words)), 2):
if subset[0] in self.top_words and subset[1] in self.features:
yield subset[0], (subset[1], occurrence_count)
if subset[1] in self.top_words and subset[0] in self.features:
yield subset[1], (subset[0], occurrence_count)
def reducer(self, word, occurrence_counts):
stripe = {}
for other_word, occurrence_count in occurrence_counts:
stripe[other_word] = stripe.get(other_word,0)+occurrence_count
yield word, stripe
def reducer_2(self, key, values):
yield str(key), list(values)[0]
if __name__ == '__main__':
MRbuildStripes.run()
# +
# Run in Hadoop
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'tests')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !time python buildStripes_stopwords_20.py \
# -r hadoop hdfs://{TEST_20}/* \
# --file ten_thousand_20.json \
# --output-dir={OUTPUT_PATH} \
# --no-output
# -
# !hadoop fs -cat {OUTPUT_PATH}/* > google_stripes_20
# +
# Run in Hadoop
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'tests')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !time python invertedIndex_v2.py \
# -r hadoop google_stripes_20 \
# --output-dir={OUTPUT_PATH} \
# --no-output
# -
# !hadoop fs -cat {OUTPUT_PATH}/* > google_index_20
# +
# Run in Hadoop
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'tests')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !time python similarity_v2.py \
# -r hadoop google_index_20 \
# --output-dir={OUTPUT_PATH} \
# --no-output
# -
# !hadoop fs -cat {OUTPUT_PATH}/* > google_similarities_2
# !cat google_similarities_2 | head
# +
############################################
# Pretty print systems tests
# Note: adjust print formatting if you need to
############################################
import json
for i in range(2,3):
print '—'*110
print "Systems test ",i," - Similarity measures"
print '—'*110
print "{0:>15} |{1:>15} |{2:>15} |{3:>15} |{4:>15} |{5:>15}".format(
"average", "pair", "cosine", "jaccard", "overlap", "dice")
print '-'*110
with open("google_similarities_"+str(i),"r") as f:
lines = f.readlines()
for line in lines:
line = line.strip()
avg,stripe = line.split("\t")
stripe = json.loads(stripe)
print "{0:>15f} |{1:>15} |{2:>15f} |{3:>15f} |{4:>15f} |{5:>15f}".format(float(avg),
stripe[0], float(stripe[1]), float(stripe[2]), float(stripe[3]), float(stripe[4]))
# -
# ### HW5.7.3 Running the full dataset on Altiscale
#
# Please contact the TAs for approval after obtaining results from 5.7.2. We have ran into issues in the past where the clusters froze because people did not test their code on a smaller dataset.
# +
# %%writefile buildStripes_stopwords.py
# #!~/opt/anaconda2/bin/python
# -*- coding: utf-8 -*-
from __future__ import division
import re
import mrjob
import json
from mrjob.protocol import RawProtocol
from mrjob.job import MRJob
from mrjob.step import MRStep
import itertools
class MRbuildStripes(MRJob):
SORT_VALUES = True
def steps(self):
return [
MRStep(
mapper_init=self.mapper_init,
mapper=self.mapper,
reducer=self.reducer,
jobconf = {
"mapreduce.job.reduces": "64",
"mapreduce.job.maps": "64",
# "SORT_VALUES":True
}
),
MRStep(
reducer=self.reducer_2,
jobconf = {
"mapreduce.job.reduces": "1",
"SORT_VALUES":True
}
)
]
def mapper_init(self):
self.idx = 9001 # To define when feature set starts
self.filename = 'ten_thousand_FULL.json'
self.top_words = []
self.features = []
#with open('features_20.json', 'r') as infile:
# self.features = json.loads(infile.read())
with open(self.filename, 'r') as infile:
self.top_words = json.loads(infile.read())
self.features =self.top_words[self.idx:]
def mapper(self, _, line):
fields = line.lower().strip("\n").split("\t")
words = fields[0].split(" ")
occurrence_count = int(fields[1])
filtered_words = [word.decode('utf-8', 'ignore') for word in words if word.decode('utf-8', 'ignore') in self.top_words]
for subset in itertools.combinations(sorted(set(filtered_words)), 2):
if subset[0] in self.top_words and subset[1] in self.features:
yield subset[0], (subset[1], occurrence_count)
if subset[1] in self.top_words and subset[0] in self.features:
yield subset[1], (subset[0], occurrence_count)
def reducer(self, word, occurrence_counts):
stripe = {}
for other_word, occurrence_count in occurrence_counts:
stripe[other_word] = stripe.get(other_word,0)+occurrence_count
yield word, stripe
def reducer_2(self, key, values):
yield str(key), list(values)[0]
if __name__ == '__main__':
MRbuildStripes.run()
# +
# Run in Hadoop
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'tests')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !time python buildStripes_stopwords.py \
# -r hadoop hdfs://{FULL_DATA}/* \
# --file ten_thousand_FULL.json \
# --output-dir={OUTPUT_PATH} \
# --no-output
# -
# !hadoop fs -cat {OUTPUT_PATH}/* > google_stripes_FULL
# +
# Run in Hadoop
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'tests')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !time python invertedIndex_v2.py \
# -r hadoop google_stripes_FULL \
# --output-dir={OUTPUT_PATH} \
# --no-output
# -
# !hadoop fs -cat {OUTPUT_PATH}/* > google_index_FULL
# +
# Run in Hadoop
OUTPUT_PATH = os.path.join(OUTPUT_PATH_BASE,'tests')
# !hadoop fs -rm -r {OUTPUT_PATH}
# !time python similarity_v2.py \
# -r hadoop google_index_FULL \
# --output-dir={OUTPUT_PATH} \
# --no-output
# -
# !hadoop fs -cat {OUTPUT_PATH}/* > google_similarities_3
# !cat google_similarities_3 | head
# +
############################################
# Pretty print systems tests
# Note: adjust print formatting if you need to
############################################
import json
for i in range(2,3):
print '—'*110
print "Systems test ",i," - Similarity measures"
print '—'*110
print "{0:>15} |{1:>15} |{2:>15} |{3:>15} |{4:>15} |{5:>15}".format(
"average", "pair", "cosine", "jaccard", "overlap", "dice")
print '-'*110
with open("google_similarities_"+str(i),"r") as f:
lines = f.readlines()
for line in lines:
line = line.strip()
avg,stripe = line.split("\t")
stripe = json.loads(stripe)
print "{0:>15f} |{1:>15} |{2:>15f} |{3:>15f} |{4:>15f} |{5:>15f}".format(float(avg),
stripe[0], float(stripe[1]), float(stripe[2]), float(stripe[3]), float(stripe[4]))
# -
# #### Pretty print results
# NOTE: depending on how you processed the stop words your results may differ from the table provided.
# +
print "\nTop/Bottom 20 results - Similarity measures - sorted by cosine"
print "(From the entire data set)"
print '—'*117
print "{0:>30} |{1:>15} |{2:>15} |{3:>15} |{4:>15} |{5:>15}".format(
"pair", "cosine", "jaccard", "overlap", "dice", "average")
print '-'*117
for stripe in sortedSims[:20]:
print "{0:>30} |{1:>15f} |{2:>15f} |{3:>15f} |{4:>15f} |{5:>15f}".format(
stripe[0], float(stripe[1]), float(stripe[2]), float(stripe[3]), float(stripe[4]), float(stripe[5]) )
print '—'*117
for stripe in sortedSims[-20:]:
print "{0:>30} |{1:>15f} |{2:>15f} |{3:>15f} |{4:>15f} |{5:>15f}".format(
stripe[0], float(stripe[1]), float(stripe[2]), float(stripe[3]), float(stripe[4]), float(stripe[5]) )
# -
Top/Bottom 20 results - Similarity measures - sorted by cosine
(From the entire data set)
—————————————————————————————————————————————————————————————————————————————————————————————————————————————————————
pair | cosine | jaccard | overlap | dice | average
---------------------------------------------------------------------------------------------------------------------
cons - pros | 0.894427 | 0.800000 | 1.000000 | 0.888889 | 0.895829
forties - twenties | 0.816497 | 0.666667 | 1.000000 | 0.800000 | 0.820791
own - time | 0.809510 | 0.670563 | 0.921168 | 0.802799 | 0.801010
little - time | 0.784197 | 0.630621 | 0.926101 | 0.773473 | 0.778598
found - time | 0.783434 | 0.636364 | 0.883788 | 0.777778 | 0.770341
nova - scotia | 0.774597 | 0.600000 | 1.000000 | 0.750000 | 0.781149
hong - kong | 0.769800 | 0.615385 | 0.888889 | 0.761905 | 0.758995
life - time | 0.769666 | 0.608789 | 0.925081 | 0.756829 | 0.765091
time - world | 0.755476 | 0.585049 | 0.937500 | 0.738209 | 0.754058
means - time | 0.752181 | 0.587117 | 0.902597 | 0.739854 | 0.745437
form - time | 0.749943 | 0.588418 | 0.876733 | 0.740885 | 0.738995
infarction - myocardial | 0.748331 | 0.560000 | 1.000000 | 0.717949 | 0.756570
people - time | 0.745788 | 0.573577 | 0.923875 | 0.729010 | 0.743063
angeles - los | 0.745499 | 0.586207 | 0.850000 | 0.739130 | 0.730209
little - own | 0.739343 | 0.585834 | 0.767296 | 0.738834 | 0.707827
life - own | 0.737053 | 0.582217 | 0.778502 | 0.735951 | 0.708430
anterior - posterior | 0.733388 | 0.576471 | 0.790323 | 0.731343 | 0.707881
power - time | 0.719611 | 0.533623 | 0.933586 | 0.695898 | 0.720680
dearly - install | 0.707107 | 0.500000 | 1.000000 | 0.666667 | 0.718443
found - own | 0.704802 | 0.544134 | 0.710949 | 0.704776 | 0.666165
—————————————————————————————————————————————————————————————————————————————————————————————————————————————————————
arrival - essential | 0.008258 | 0.004098 | 0.009615 | 0.008163 | 0.007534
governments - surface | 0.008251 | 0.003534 | 0.014706 | 0.007042 | 0.008383
king - lesions | 0.008178 | 0.003106 | 0.017857 | 0.006192 | 0.008833
clinical - stood | 0.008178 | 0.003831 | 0.011905 | 0.007634 | 0.007887
till - validity | 0.008172 | 0.003367 | 0.015625 | 0.006711 | 0.008469
evidence - started | 0.008159 | 0.003802 | 0.012048 | 0.007576 | 0.007896
forces - record | 0.008152 | 0.003876 | 0.011364 | 0.007722 | 0.007778
primary - stone | 0.008146 | 0.004065 | 0.009091 | 0.008097 | 0.007350
beneath - federal | 0.008134 | 0.004082 | 0.008403 | 0.008130 | 0.007187
factors - rose | 0.008113 | 0.004032 | 0.009346 | 0.008032 | 0.007381
evening - functions | 0.008069 | 0.004049 | 0.008333 | 0.008065 | 0.007129
bone - told | 0.008061 | 0.003704 | 0.012346 | 0.007380 | 0.007873
building - occurs | 0.008002 | 0.003891 | 0.010309 | 0.007752 | 0.007489
company - fig | 0.007913 | 0.003257 | 0.015152 | 0.006494 | 0.008204
chronic - north | 0.007803 | 0.003268 | 0.014493 | 0.006515 | 0.008020
evaluation - king | 0.007650 | 0.003030 | 0.015625 | 0.006042 | 0.008087
resulting - stood | 0.007650 | 0.003663 | 0.010417 | 0.007299 | 0.007257
agent - round | 0.007515 | 0.003289 | 0.012821 | 0.006557 | 0.007546
afterwards - analysis | 0.007387 | 0.003521 | 0.010204 | 0.007018 | 0.007032
posterior - spirit | 0.007156 | 0.002660 | 0.016129 | 0.005305 | 0.007812
# # HW5.8 - Evaluation of synonyms that your discovered
#
# In this part of the assignment you will evaluate the success of you synonym detector. Take the top 1,000 closest/most similar/correlative pairs of words as determined by your measure in HW5.7, and use the synonyms function from the wordnet synonnyms list from the nltk package (see provided code below).
#
# For each (word1,word2) pair, check to see if word1 is in the list,
# synonyms(word2), and vice-versa. If one of the two is a synonym of the other,
# then consider this pair a 'hit', and then report the precision, recall, and F1 measure of
# your detector across your 1,000 best guesses. Report the macro averages of these measures.
# ### Calculate performance measures:
# $$Precision (P) = \frac{TP}{TP + FP} $$
# $$Recall (R) = \frac{TP}{TP + FN} $$
# $$F1 = \frac{2 * ( precision * recall )}{precision + recall}$$
#
#
# We calculate Precision by counting the number of hits and dividing by the number of occurances in our top1000 (opportunities)
# We calculate Recall by counting the number of hits, and dividing by the number of synonyms in wordnet (syns)
#
#
# Other diagnostic measures not implemented here: https://en.wikipedia.org/wiki/F1_score#Diagnostic_Testing
# +
''' Performance measures '''
from __future__ import division
import numpy as np
import json
import nltk
from nltk.corpus import wordnet as wn
import sys
#print all the synset element of an element
def synonyms(string):
syndict = {}
for i,j in enumerate(wn.synsets(string)):
syns = j.lemma_names()
for syn in syns:
syndict.setdefault(syn,1)
return syndict.keys()
hits = []
TP = 0
FP = 0
TOTAL = 0
flag = False # so we don't double count, but at the same time don't miss hits
top1000sims = []
with open("sims2/top1000sims","r") as f:
for line in f.readlines():
line = line.strip()
avg,lisst = line.split("\t")
lisst = json.loads(lisst)
lisst.append(avg)
top1000sims.append(lisst)
measures = {}
not_in_wordnet = []
for line in top1000sims:
TOTAL += 1
pair = line[0]
words = pair.split(" - ")
for word in words:
if word not in measures:
measures[word] = {"syns":0,"opps": 0,"hits":0}
measures[word]["opps"] += 1
syns0 = synonyms(words[0])
measures[words[1]]["syns"] = len(syns0)
if len(syns0) == 0:
not_in_wordnet.append(words[0])
if words[1] in syns0:
TP += 1
hits.append(line)
flag = True
measures[words[1]]["hits"] += 1
syns1 = synonyms(words[1])
measures[words[0]]["syns"] = len(syns1)
if len(syns1) == 0:
not_in_wordnet.append(words[1])
if words[0] in syns1:
if flag == False:
TP += 1
hits.append(line)
measures[words[0]]["hits"] += 1
flag = False
precision = []
recall = []
f1 = []
for key in measures:
p,r,f = 0,0,0
if measures[key]["hits"] > 0 and measures[key]["syns"] > 0:
p = measures[key]["hits"]/measures[key]["opps"]
r = measures[key]["hits"]/measures[key]["syns"]
f = 2 * (p*r)/(p+r)
# For calculating measures, only take into account words that have synonyms in wordnet
if measures[key]["syns"] > 0:
precision.append(p)
recall.append(r)
f1.append(f)
# Take the mean of each measure
print "—"*110
print "Number of Hits:",TP, "out of top",TOTAL
print "Number of words without synonyms:",len(not_in_wordnet)
print "—"*110
print "Precision\t", np.mean(precision)
print "Recall\t\t", np.mean(recall)
print "F1\t\t", np.mean(f1)
print "—"*110
print "Words without synonyms:"
print "-"*100
for word in not_in_wordnet:
print synonyms(word),word
# -
# ### Sample output
——————————————————————————————————————————————————————————————————————————————————————————————————————————————
Number of Hits: 31 out of top 1000
Number of words without synonyms: 67
——————————————————————————————————————————————————————————————————————————————————————————————————————————————
Precision 0.0280214404967
Recall 0.0178598869579
F1 0.013965517619
——————————————————————————————————————————————————————————————————————————————————————————————————————————————
Words without synonyms:
----------------------------------------------------------------------------------------------------
[] scotia
[] hong
[] kong
[] angeles
[] los
[] nor
[] themselves
[]
.......
# # HW5.9 - OPTIONAL: using different vocabulary subsets
#
# Repeat HW5 using vocabulary words ranked from 8001,-10,000; 7001,-10,000; 6001,-10,000; 5001,-10,000; 3001,-10,000; and 1001,-10,000;
# Dont forget to report you Cluster configuration.
#
# Generate the following graphs:
# -- vocabulary size (X-Axis) versus CPU time for indexing
# -- vocabulary size (X-Axis) versus number of pairs processed
# -- vocabulary size (X-Axis) versus F1 measure, Precision, Recall
#
# # HW5.10 - OPTIONAL
#
# There are many good ways to build our synonym detectors, so for this optional homework,
# measure co-occurrence by (left/right/all) consecutive words only,
# or make stripes according to word co-occurrences with the accompanying
# 2-, 3-, or 4-grams (note here that your output will no longer
# be interpretable as a network) inside of the 5-grams.
# # HW5.11 - OPTIONAL
#
# Once again, benchmark your top 10,000 associations (as in 5.7), this time for your
# results from 5.8. Has your detector improved?
| HW5/MIDS-W261-HW-05-PHASE2-TEMPLATE-V2.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Scala (2.12)
// language: scala
// name: scala212
// ---
// # Sttp
// A minimal setup of sttp
import $ivy.`com.softwaremill.sttp.client::core:2.0.0-RC2`
import sttp.client.quick.{quickRequest, UriContext, backend}
quickRequest.get(uri"http://httpbin.org/ip").send()
// ## Async sttp
// Use the async http client backend of sttp, which uses the netty behind the scenes.
// +
import $ivy.`com.softwaremill.sttp.client::async-http-client-backend-monix:2.0.0-RC2`
import sttp.client.asynchttpclient.monix._
implicit val sttpBackend = AsyncHttpClientMonixBackend()
import sttp.client.basicRequest
val r = basicRequest.get(uri"http://httpbin.org/ip")
// -
import monix.execution.Scheduler.Implicits.global
val task = for {
backend <- sttpBackend
res <- r.send()(backend, implicitly)
} yield res
task.foreach(println)
// # Armeria
// +
import $ivy.`com.linecorp.armeria:armeria:0.96.0`
import com.linecorp.armeria.client.WebClient
val client = WebClient.of("http://httpbin.org/")
val future = client.get("/ip").aggregate()
val res = future.join()
// -
val body = res.contentUtf8
| notebooks/http.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import networkx as nx
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# ## 1.建立Graph
#
# 如下图所示建立一个图。
# 
# +
plt.figure(figsize=(10,5))
G = nx.DiGraph()
G.add_nodes_from(['Robin_Yao_Wenbin','Wei','Yang','Yuan','Xiao','Chen','Peng']) #加点集合
G.add_edge('Wei' ,'Robin_Yao_Wenbin' ) #一次添加一条边
G.add_edge('Yang' ,'Robin_Yao_Wenbin' ) #一次添加一条边
G.add_edge('Yuan' ,'Robin_Yao_Wenbin' ) #一次添加一条边
G.add_edge('Chen' ,'Robin_Yao_Wenbin' ) #一次添加一条边
G.add_edge('Peng' ,'Chen' ) #一次添加一条边
G.add_edge('Xiao' ,'Wei' ) #一次添加一条边
G.add_edge('Chen' ,'Wei' ) #一次添加一条边
G.add_edge('Yuan' ,'Yang' ) #一次添加一条边
nx.draw(G, with_labels=True , node_size = 2000)
plt.show()
# -
# ## 2.计算Pagerank
#
# **PR=alpha * (A * PR+dangling分配)+(1-alpha) * 平均分配**
#
# **也就是三部分,A*PR其实是我们用图矩阵分配的,dangling分配则是对dangling node的PR值进行分配,(1-alpha)分配则是天下为公大家一人一份分配的**
#
# **dangling node 也就是悬空结点,它的出度为0,也就是无法从它到任何其他结点,解决办法是增加一定的随机性,dangling分配其实就是加上一个随机向量,也就是无法从这个结点去往任何其他结点,但是可能会随机重新去一个结点,也可以这么理解,到了一个网站,这个网站不连接到任何网站,但是浏览者可能重新随机打开一个页面。**
#
# **其实通俗的来说,我们可以将PageRank看成抢夺大赛,有三种抢夺机制。**
#
# **1,A*PR这种是自由分配,大家都愿意参与竞争交流的分配**
#
# **2,dangling是强制分配,有点类似打倒土豪分田地的感觉,你不参与自由市场,那好,我们就特地帮你强制分。**
#
# **3,平均分配,其实就是有个机会大家实现共产主义了,不让spider trap这种产生rank sink的节点捞太多油水,其实客观上也是在帮dangling分配。**
#
# **从图和矩阵的角度来说,可以这样理解,我们这个矩阵可以看出是个有向图**
#
# **矩阵要收敛-->矩阵有唯一解-->n阶方阵对应有向图是强连通的-->两个节点相互可达,1能到2,2能到1**
#
# **如果是个强连通图,就是我们上面说的第1种情况,自由竞争,那么我们可以确定是收敛的**
#
# **不然就会有spider trap造成rank sink问题**
#
#
# 具体可见下述链接:
# https://blog.csdn.net/a_31415926/article/details/40510175
pr = nx.pagerank(G, alpha=0.85, personalization=None,
max_iter=100, tol=1.0e-6, nstart=None, weight='weight',
dangling=None)
pr
# ## 3.检验理解的正确性
# +
plt.figure(figsize=(4,1))
G2 = nx.DiGraph()
G2.add_nodes_from(['A','B']) #加点集合
G2.add_edge('A' ,'B' ) #一次添加一条边
# G2.add_edge('B' ,'A' ) #一次添加一条边
nx.draw(G2, with_labels=True , node_size = 2000)
plt.show()
pr = nx.pagerank(G2, alpha=0.85, personalization=None,
max_iter=100, tol=1.0e-6, nstart=None, weight='weight',
dangling=None)
pr
# -
# 验证方法:
#
# 将上述算法计算得到的PR(A)和PR(B)代入到PR(A)和PR(B)的计算公式中,观察结果是否稳定,即达到达到pagerank的收敛条件。
#
# PR(A)=0.15/2+0.85 * 0.5 * 0.649123=0.350877
#
# PR(B)=0.350877+0.85 * 0.35087=0.649123
# 得证,说明源码解析中的说法是正确的。
#
# 同时我还发现有意思的是用网上传统的那些说法,归一化后也和上述计算值是一模一样的。
# 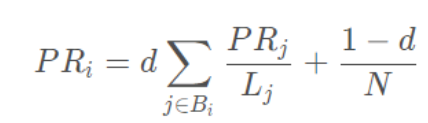
# +
plt.figure(figsize=(4,1))
G3 = nx.DiGraph()
G3.add_nodes_from(['A','B','C']) #加点集合
G3.add_edge('A' ,'B' ) #一次添加一条边
G3.add_edge('B' ,'C' ) #一次添加一条边
nx.draw(G3, with_labels=True , node_size = 2000)
plt.show()
pr = nx.pagerank(G3, alpha=0.85, personalization=None,
max_iter=300, tol=1.0e-6, nstart=None, weight='weight',
dangling=None)
pr
| python业务代码/networkx-pagerank/networkx-pagerank.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
# Add the path to system, local or mounted S3 bucket, e.g. /dbfs/mnt/<path_to_bucket>
sys.path.append('./secrets.py')
import logging
import math
import os
from influxdb import DataFrameClient
import numpy as np
import matplotlib.mlab as mlab
import pandas as pd
import matplotlib.pyplot as plt
from tabulate import tabulate
from tqdm import tqdm
# %matplotlib inline
logging.basicConfig(level=logging.INFO)
LOGGER = logging.getLogger(__name__)
# Need to ssh tunnel for this to work
# ssh -L 8086:localhost:8086 aq.byu.edu -N
influx = DataFrameClient(
host=HOST,
port=PORT,
username=USERNAME,
password=PASSWORD,
database=DATABASE,
)
def large_query(influx, measurement, query, total=None, limit=100_000):
if total is not None:
total = math.ceil(total / limit)
with tqdm(total=total) as pbar:
offset = 0
while True:
new_query = query + " LIMIT {} OFFSET {}".format(limit, offset)
data = influx.query(new_query)
data = data[measurement]
received = len(data)
pbar.update(1)
yield data
offset += limit
if received != limit:
break
def load_data(filename):
if os.path.exists(filename):
LOGGER.info("Loading cached data...")
return pd.read_hdf(filename)
LOGGER.info("Downloading data...")
result = influx.query(
"SELECT COUNT(sequence) FROM air_quality_sensor WHERE time > '2019-10-01' AND time <= now()"
)
count = result["air_quality_sensor"].values[0][0]
queries = large_query(
influx,
"air_quality_sensor",
"SELECT * FROM air_quality_sensor WHERE time > '2019-10-01' AND time <= now()",
count,
)
all_data = pd.concat(list(queries), sort=False)
all_data.to_hdf(filename, "data")
return all_data
data = load_data("aq_data.h5")
gold_data = load_data("aq_data.h5")
LOGGER.info("Done loading data...")
# +
# Day - This is the working boxplot for hybrid only Mongolia deployed sensors
# https://stackoverflow.com/questions/22800079/converting-time-zone-pandas-dataframe
# https://pandas.pydata.org/pandas-docs/version/0.23.4/generated/pandas.DataFrame.between_time.html
from IPython.core.debugger import set_trace
# https://matplotlib.org/3.1.3/gallery/statistics/boxplot_color.html
data = gold_data
print(data.index[1])
# data = data.tz_convert(None)
# data.index = data.index.tz_localize('GMT')
data.index = data.index.tz_convert('Asia/Ulaanbaatar')
print(data.index[1])
labels = ['Mon', 'Tue', 'Wed', 'Thu', 'Fri', 'Sat', 'Sun']
plt.xlabel('Dates')
plt.ylabel('PM 2.5 Value')
plt.title('Week PM 2.5 for sensors')
plt.grid(True)
days = ['05', '06', '07', '08', '09', '10', '11']
data = data[data.pm2_5 >= 0]
data = data[data.location_name == 'Mongolia']
heater_only_modified_gers = ['GA', 'ND', 'NE']
data = data[(data.name == heater_only_modified_gers[0]) | (data.name == heater_only_modified_gers[1]) | (data.name == heater_only_modified_gers[2])]
all_sensors_names = []
for name, sensor_data in data.groupby("name"):
all_sensors_names.append(name)
print("All Sensors names:", all_sensors_names)
day_names = data.index.day_name()
print(type(day_names))
print(day_names[:10])
monday = data[(data.index.day_name() == 'Monday')].between_time('15:00:00', '01:00:00')['pm2_5']
tuesday = data[data.index.day_name() == 'Tuesday'].between_time('15:00:00', '01:00:00')['pm2_5']
wednesday = data[data.index.day_name() == 'Wednesday'].between_time('15:00:00', '01:00:00')['pm2_5']
thursday = data[data.index.day_name() == 'Thursday'].between_time('15:00:00', '01:00:00')['pm2_5']
friday = data[data.index.day_name() == 'Friday'].between_time('15:00:00', '01:00:00')['pm2_5']
saturday = data[data.index.day_name() == 'Saturday'].between_time('15:00:00', '01:00:00')['pm2_5']
sunday = data[data.index.day_name() == 'Sunday'].between_time('15:00:00', '01:00:00')['pm2_5']
all_days = [monday, tuesday, wednesday, thursday, friday, saturday, sunday]
results = plt.boxplot(all_days, showfliers=False, labels=labels, showmeans=True, meanline=True)
plt.savefig("./weely_hybrid_day_boxplot_pm_2_5/weely_hybrid_day_boxplot_pm_2_5.png", format='png')
# print(results)
print('whiskers: ', [item.get_ydata()[1] for item in results['whiskers']])
print('caps: ', [item.get_ydata()[1] for item in results['caps']])
print('boxes: ', [item.get_ydata()[1] for item in results['boxes']])
print('medians: ', [item.get_ydata()[1] for item in results['medians']])
print('means: ', [item.get_ydata()[1] for item in results['means']])
print('fliers: ', [item.get_ydata()[1] for item in results['fliers']])
data = gold_data
# +
# Day each sensors - This is the working boxplot for hybrid only Mongolia deployed sensors
# https://stackoverflow.com/questions/22800079/converting-time-zone-pandas-dataframe
# https://pandas.pydata.org/pandas-docs/version/0.23.4/generated/pandas.DataFrame.between_time.html
import numpy as np
from IPython.core.debugger import set_trace
# https://matplotlib.org/3.1.3/gallery/statistics/boxplot_color.html
data = gold_data
print(data.index[1])
# data = data.tz_convert(None)
# data.index = data.index.tz_localize('GMT')
data.index = data.index.tz_convert('Asia/Ulaanbaatar')
print(data.index[1])
labels = ['AK', 'AL', 'AR', 'AZ', 'CA', 'CO', 'CT', 'DE', 'FL', 'IA', 'KS', 'KY', 'LA', 'MD', 'ME', 'MI', 'MN', 'MS', 'MT', 'NC', 'NH']
plt.xlabel('Sensors')
plt.ylabel('PM 2.5 Value')
plt.title('Week PM 2.5 for sensors')
plt.grid(True)
days_of_week = ['Monday', 'Tuesday', 'Wednesday', 'Thursday', 'Friday', 'Saturday', 'Sunday']
data = data[data.pm2_5 >= 0]
data = data[data.location_name == 'Mongolia']
heater_only_modified_gers = ['GA', 'ND', 'NE']
data = data[(data.name == heater_only_modified_gers[0]) | (data.name == heater_only_modified_gers[1]) | (data.name == heater_only_modified_gers[2])]
all_sensors_names = []
for name, sensor_data in data.groupby("name"):
all_sensors_names.append(name)
print("All Sensors names:", all_sensors_names)
day_names = data.index.day_name()
print(type(day_names))
print(day_names[:10])
# for name, sensor_data in data.groupby("name"):
for day in days_of_week:
plt.xlabel('Sensors for {0}'.format(day))
plt.ylabel('PM 2.5 Value')
plt.title('Week PM 2.5 for sensors for {0}'.format(day))
plt.grid(True)
sensors_data = list()
sensors_name = list()
temp = data[data.index.day_name() == day]
# temp = temp.groupby("name")
for name, sensor_data in temp.groupby("name"):
sensors_name.append(name)
sensors_data.append(temp.groupby("name").get_group(name)["pm2_5"].between_time('15:00:00', '01:00:00').to_numpy().tolist())
results = plt.boxplot(sensors_data, showfliers=False, labels=sensors_name, showmeans=True, meanline=True)
print('whiskers: ', [item.get_ydata()[1] for item in results['whiskers']])
print('caps: ', [item.get_ydata()[1] for item in results['caps']])
print('boxes: ', [item.get_ydata()[1] for item in results['boxes']])
print('medians: ', [item.get_ydata()[1] for item in results['medians']])
print('means: ', [item.get_ydata()[1] for item in results['means']])
print('fliers: ', [item.get_ydata()[1] for item in results['fliers']])
plt.show()
# monday = data[(data.index.day_name() == 'Monday')]
# monday = monday['AK'].between_time('15:00:00', '01:00:00')['pm2_5']
# tuesday = data[data.index.day_name() == 'Tuesday'].between_time('15:00:00', '01:00:00')['pm2_5']
# wednesday = data[data.index.day_name() == 'Wednesday'].between_time('15:00:00', '01:00:00')['pm2_5']
# thursday = data[data.index.day_name() == 'Thursday'].between_time('15:00:00', '01:00:00')['pm2_5']
# friday = data[data.index.day_name() == 'Friday'].between_time('15:00:00', '01:00:00')['pm2_5']
# saturday = data[data.index.day_name() == 'Saturday'].between_time('15:00:00', '01:00:00')['pm2_5']
# sunday = data[data.index.day_name() == 'Sunday'].between_time('15:00:00', '01:00:00')['pm2_5']
# all_days = [monday, tuesday, wednesday, thursday, friday, saturday, sunday]
# results = plt.boxplot(all_days, showfliers=False, labels=labels, showmeans=True, meanline=True)
# plt.savefig("./weely_hybrid_day_boxplot_pm_2_5/weely_hybrid_day_boxplot_pm_2_5.png", format='png')
# print(results)
# print('whiskers: ', [item.get_ydata()[1] for item in results['whiskers']])
# print('caps: ', [item.get_ydata()[1] for item in results['caps']])
# print('boxes: ', [item.get_ydata()[1] for item in results['boxes']])
# print('medians: ', [item.get_ydata()[1] for item in results['medians']])
# print('means: ', [item.get_ydata()[1] for item in results['means']])
# print('fliers: ', [item.get_ydata()[1] for item in results['fliers']])
data = gold_data
# +
# Night - This is the working boxplot for hybrid only Mongolia deployed sensors
# https://stackoverflow.com/questions/22800079/converting-time-zone-pandas-dataframe
# https://pandas.pydata.org/pandas-docs/version/0.23.4/generated/pandas.DataFrame.between_time.html
from IPython.core.debugger import set_trace
# https://matplotlib.org/3.1.3/gallery/statistics/boxplot_color.html
data = gold_data
print(data.index[1])
# data = data.tz_convert(None)
# data.index = data.index.tz_localize('GMT')
data.index = data.index.tz_convert('Asia/Ulaanbaatar')
print(data.index[1])
# data.tz_localize('UTC', level=8)
labels = ['Mon', 'Tue', 'Wed', 'Thu', 'Fri', 'Sat', 'Sun']
plt.xlabel('Dates')
plt.ylabel('PM 2.5 Value')
plt.title('Week PM 2.5 for sensors')
plt.grid(True)
days = ['05', '06', '07', '08', '09', '10', '11']
data = data[data.pm2_5 >= 0]
data = data[data.location_name == 'Mongolia']
heater_only_modified_gers = ['GA', 'ND', 'NE']
data = data[(data.name == heater_only_modified_gers[0]) | (data.name == heater_only_modified_gers[1]) | (data.name == heater_only_modified_gers[2])]
all_sensors_names = []
for name, sensor_data in data.groupby("name"):
all_sensors_names.append(name)
print("All Sensors names:", all_sensors_names)
day_names = data.index.day_name()
print(type(day_names))
print(day_names[:10])
monday = data[(data.index.day_name() == 'Monday')].between_time('01:01', '07:59')['pm2_5']
tuesday = data[data.index.day_name() == 'Tuesday'].between_time('01:01', '07:59')['pm2_5']
wednesday = data[data.index.day_name() == 'Wednesday'].between_time('01:01', '07:59')['pm2_5']
thursday = data[data.index.day_name() == 'Thursday'].between_time('01:01', '07:59')['pm2_5']
friday = data[data.index.day_name() == 'Friday'].between_time('01:01', '07:59')['pm2_5']
saturday = data[data.index.day_name() == 'Saturday'].between_time('01:01', '07:59')['pm2_5']
sunday = data[data.index.day_name() == 'Sunday'].between_time('01:01', '07:59')['pm2_5']
all_days = [monday, tuesday, wednesday, thursday, friday, saturday, sunday]
results = plt.boxplot(all_days, showfliers=False, labels=labels, showmeans=True, meanline=True)
plt.savefig("./weely_hybrid_night_boxplot_pm_2_5/weely_hybrid_night_boxplot_pm_2_5.png", format='png')
# print(results)
print('whiskers: ', [item.get_ydata()[1] for item in results['whiskers']])
print('caps: ', [item.get_ydata()[1] for item in results['caps']])
print('boxes: ', [item.get_ydata()[1] for item in results['boxes']])
print('medians: ', [item.get_ydata()[1] for item in results['medians']])
print('means: ', [item.get_ydata()[1] for item in results['means']])
print('fliers: ', [item.get_ydata()[1] for item in results['fliers']])
data = gold_data
# +
# Night each sensors - This is the working boxplot for hybrid only Mongolia deployed sensors
# https://stackoverflow.com/questions/22800079/converting-time-zone-pandas-dataframe
# https://pandas.pydata.org/pandas-docs/version/0.23.4/generated/pandas.DataFrame.between_time.html
from IPython.core.debugger import set_trace
# https://matplotlib.org/3.1.3/gallery/statistics/boxplot_color.html
data = gold_data
print(data.index[1])
# data = data.tz_convert(None)
# data.index = data.index.tz_localize('GMT')
data.index = data.index.tz_convert('Asia/Ulaanbaatar')
print(data.index[1])
# data.tz_localize('UTC', level=8)
labels = ['Mon', 'Tue', 'Wed', 'Thu', 'Fri', 'Sat', 'Sun']
plt.xlabel('Dates')
plt.ylabel('PM 2.5 Value')
plt.title('Week PM 2.5 for sensors')
plt.grid(True)
days_of_week = ['Monday', 'Tuesday', 'Wednesday', 'Thursday', 'Friday', 'Saturday', 'Sunday']
data = data[data.pm2_5 >= 0]
data = data[data.location_name == 'Mongolia']
heater_only_modified_gers = ['GA', 'ND', 'NE']
data = data[(data.name == heater_only_modified_gers[0]) | (data.name == heater_only_modified_gers[1]) | (data.name == heater_only_modified_gers[2])]
all_sensors_names = []
for name, sensor_data in data.groupby("name"):
all_sensors_names.append(name)
print("All Sensors names:", all_sensors_names)
day_names = data.index.day_name()
print(type(day_names))
print(day_names[:10])
for day in days_of_week:
plt.xlabel('Sensors for {0}'.format(day))
plt.ylabel('PM 2.5 Value')
plt.title('Week PM 2.5 for sensors for {0}'.format(day))
plt.grid(True)
sensors_data = list()
sensors_name = list()
temp = data[data.index.day_name() == day]
# temp = temp.groupby("name")
for name, sensor_data in temp.groupby("name"):
sensors_name.append(name)
sensors_data.append(temp.groupby("name").get_group(name)["pm2_5"].between_time('01:01', '07:59').to_numpy().tolist())
results = plt.boxplot(sensors_data, showfliers=False, labels=sensors_name, showmeans=True, meanline=True)
print('whiskers: ', [item.get_ydata()[1] for item in results['whiskers']])
print('caps: ', [item.get_ydata()[1] for item in results['caps']])
print('boxes: ', [item.get_ydata()[1] for item in results['boxes']])
print('medians: ', [item.get_ydata()[1] for item in results['medians']])
print('means: ', [item.get_ydata()[1] for item in results['means']])
print('fliers: ', [item.get_ydata()[1] for item in results['fliers']])
plt.show()
# monday = data[(data.index.day_name() == 'Monday')].between_time('01:01', '07:59')['pm2_5']
# tuesday = data[data.index.day_name() == 'Tuesday'].between_time('01:01', '07:59')['pm2_5']
# wednesday = data[data.index.day_name() == 'Wednesday'].between_time('01:01', '07:59')['pm2_5']
# thursday = data[data.index.day_name() == 'Thursday'].between_time('01:01', '07:59')['pm2_5']
# friday = data[data.index.day_name() == 'Friday'].between_time('01:01', '07:59')['pm2_5']
# saturday = data[data.index.day_name() == 'Saturday'].between_time('01:01', '07:59')['pm2_5']
# sunday = data[data.index.day_name() == 'Sunday'].between_time('01:01', '07:59')['pm2_5']
# all_days = [monday, tuesday, wednesday, thursday, friday, saturday, sunday]
# results = plt.boxplot(all_days, showfliers=False, labels=labels, showmeans=True, meanline=True)
# plt.savefig("./weely_hybrid_night_boxplot_pm_2_5/weely_hybrid_night_boxplot_pm_2_5.png", format='png')
# # print(results)
# print('whiskers: ', [item.get_ydata()[1] for item in results['whiskers']])
# print('caps: ', [item.get_ydata()[1] for item in results['caps']])
# print('boxes: ', [item.get_ydata()[1] for item in results['boxes']])
# print('medians: ', [item.get_ydata()[1] for item in results['medians']])
# print('means: ', [item.get_ydata()[1] for item in results['means']])
# print('fliers: ', [item.get_ydata()[1] for item in results['fliers']])
data = gold_data
# -
| day_vs_night_heater_only_modified/day_vs_night.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dimuon spectrum
#
# This code is a columnar adaptation of [a ROOT tutorial](https://root.cern.ch/doc/master/df102__NanoAODDimuonAnalysis_8py.html) showcasing the awkward array toolset, and utilizing FCAT histograms.
# This also shows the analysis object syntax implemented by FCAT `JaggedCandidateArray`, and the usage of an accumulator class provided by FCAT.
# +
import time
import uproot
import awkward
# %matplotlib inline
from coffea import hist
from coffea.analysis_objects import JaggedCandidateArray
from coffea.processor import defaultdict_accumulator
# -
# uproot supports xrootd, but its nicer to have them local (about 7 GB)
# !mkdir -p data
# !xrdcp root://eospublic.cern.ch//eos/root-eos/cms_opendata_2012_nanoaod/Run2012B_DoubleMuParked.root data/
# !xrdcp root://eospublic.cern.ch//eos/root-eos/cms_opendata_2012_nanoaod/Run2012C_DoubleMuParked.root data/
# +
tstart = time.time()
files = [
'data/Run2012B_DoubleMuParked.root',
'data/Run2012C_DoubleMuParked.root',
]
masshist = hist.Hist("Counts", hist.Bin("mass", r"$m_{\mu\mu}$ [GeV]", 30000, 0.25, 300))
cutflow = defaultdict_accumulator(lambda: 0)
branches = ['nMuon', 'Muon_pt', 'Muon_eta', 'Muon_phi', 'Muon_mass', 'Muon_charge']
for chunk in uproot.iterate(files, 'Events', branches=branches, entrysteps=500000, namedecode='ascii'):
muons = JaggedCandidateArray.candidatesfromcounts(chunk['nMuon'],
pt=chunk['Muon_pt'].content,
eta=chunk['Muon_eta'].content,
phi=chunk['Muon_phi'].content,
mass=chunk['Muon_mass'].content,
charge=chunk['Muon_charge'].content,
)
cutflow['all events'] += muons.size
twomuons = (muons.counts == 2)
cutflow['two muons'] += twomuons.sum()
opposite_charge = twomuons & (muons['charge'].prod() == -1)
cutflow['opposite charge'] += opposite_charge.sum()
dimuons = muons[opposite_charge].distincts()
masshist.fill(mass=dimuons.mass.flatten())
elapsed = time.time() - tstart
print(dict(cutflow))
# -
ax = hist.plot1d(masshist)
ax.set_xscale('log')
ax.set_yscale('log')
ax.set_ylim(0.1, 1e6)
print("Events/s:", cutflow['all events']/elapsed)
| binder/muonspectrum_v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# Import Librerie
import pandas as pd
import numpy as np
import os
from sqlalchemy import create_engine
import nltk
from nltk.tokenize import word_tokenize
import re
from collections import Counter
from nltk.corpus import stopwords
import string
import datetime
config = {}
config_path = os.path.join(os.path.abspath('../../'))
config_name = 'config.py'
config_file = os.path.join(config_path,config_name)
exec(open(config_file).read(),config)
# +
# get database connection
db=config['DATABASE_ELE']
schema=config['SCHEMA_ELE']
engine = create_engine(db)
user1 = config['USER1']
user2 = config['USER2']
user3 = config['USER3']
# -
#Leggo il file messo a disposizione da Istat per i comuni
url="http://www.istat.it/storage/codici-unita-amministrative/Elenco-comuni-italiani.csv"
df_comuni = pd.read_csv(url,sep = ';',encoding='latin-1')
def calc_prov(x):
prov = x['Denominazione provincia']
citt = x[u'Denominazione Città metropolitana']
if(prov=='-'):
prov = citt
return prov
df_comuni['provincia_new'] = df_comuni.apply(lambda x: calc_prov(x), axis=1)
comuni = df_comuni[u'Denominazione in italiano']
province = df_comuni[['Denominazione in italiano','provincia_new']]
# get today's date
todays_date = datetime.datetime.now()
# Leggo le ultime news
cur = engine.execute(
'''
SELECT
"desc" as msg
,to_char ("pubAt"::timestamp at time zone 'UTC', 'YYYY-MM-DD') as dt
,fonte as fonte
,"user" as user
FROM ''' + schema + '''."news"
WHERE dt_rif=(select max(dt_rif) from ''' + schema + '''."news")
''')
f_news = cur.fetchall()
header = ['msg','dt','fonte','user']
df_news = pd.DataFrame(f_news, columns=header)
# Leggo gli ultime social
cur = engine.execute(
'''
SELECT
msg as msg
,to_char(dt::timestamp, 'YYYY-MM-DD') as dt
,sorgente as fonte
,"user" as user
FROM ''' + schema + '''."timeline"
WHERE
sorgente in ('twitter','facebook') and
dt_rif=(select max(dt_rif) from ''' + schema + '''."timeline")
''')
f_social = cur.fetchall()
header = ['msg','dt','fonte','user']
df_social = pd.DataFrame(f_social, columns=header)
df = df_social.append(df_news)
df.reset_index(drop=True, inplace=True)
df.head(2)
# +
emoticons_str = r"""
(?:
[:=;] # Eyes
[oO\-]? # Nose (optional)
[D\)\]\(\]/\\OpP] # Mouth
)"""
regex_str = [
emoticons_str,
r'<[^>]+>', # HTML tags
r'(?:@[\w_]+)', # @-mentions
r'http[s]?://(?:[a-z]|[0-9]|[$-_@.&+]|[!*\(\),]|(?:%[0-9a-f][0-9a-f]))+', # URLs
r'(?:(?:\d+,?)+(?:\.?\d+)?)', # numbers
r"(?:[a-z][a-z'\-_]+[a-z])", # words with - and '
r'(?:[\w_]+)', # other words
r'(?:\S)', # anything else
]
tokens_re = re.compile(r'('+'|'.join(regex_str)+')', re.VERBOSE | re.IGNORECASE)
emoticon_re = re.compile(r'^'+emoticons_str+'$', re.VERBOSE | re.IGNORECASE)
def tokenize(s):
return tokens_re.findall(s)
def preprocess(s, lowercase=False):
tokens = tokenize(s)
if lowercase:
tokens = [token if emoticon_re.search(token) else token.lower() for token in tokens]
return tokens
# -
punctuation = list(string.punctuation)
stop = punctuation
mappe = []
for i, row in df['msg'].iteritems():
terms_only = [term for term in preprocess(row) if term not in stop and not term.startswith((':/'))]
for term in terms_only:
d = {}
# check comune
if not(comuni[comuni.isin([term])].empty):
d['comune'] = term
d['fonte'] = df['fonte'][i]
d['dt_post'] = df['dt'][i]
d['user'] = df['user'][i]
d['dt_rif'] = todays_date
d['provincia'] = province.loc[province['Denominazione in italiano'] == term]['provincia_new'].values[0]
mappe.append(d)
df_mappe = pd.DataFrame(mappe)
df_mappe
# write to db
df_mappe.to_sql('mappe', engine, schema=schema, if_exists='append')
| Mappe.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeClassifier
from sklearn import tree
from sklearn.metrics import classification_report
from sklearn import preprocessing
data = pd.read_csv('Company_Data.csv')
data
data.info()
data.describe()
import seaborn as sns
sns.pairplot(data)
corr = data.corr()
fig, ax = plt.subplots(figsize=(10, 6))
sns.heatmap(corr, cmap='magma', annot=True, fmt=".2f")
plt.xticks(range(len(corr.columns)), corr.columns);
plt.yticks(range(len(corr.columns)), corr.columns)
plt.show()
# +
sns.countplot(data['ShelveLoc'])
plt.show()
sns.countplot(data['Urban'])
plt.show()
sns.countplot(data['US'])
plt.show()
# -
data = pd.get_dummies(data)
data.head()
data['Sales'] = pd.cut(x=data['Sales'],bins=[0, 6, 12, 18], labels=['Low','Medium', 'High'], right = False)
data['Sales']
data['Sales'].value_counts()
data.head()
dataset = data.values
X = dataset[:, 1:]
Y = dataset[:,0]
from numpy import set_printoptions
from sklearn.feature_selection import SelectKBest
from sklearn.feature_selection import chi2
test = SelectKBest(score_func=chi2, k=5)
fit = test.fit(X, Y)
# +
scores = fit.scores_
features = fit.transform(X)
# -
scores
data.columns
col_names = ['CompPrice', 'Income', 'Advertising', 'Population', 'Price',
'Age', 'Education', 'ShelveLoc_Bad', 'ShelveLoc_Good',
'ShelveLoc_Medium', 'Urban_No', 'Urban_Yes', 'US_No', 'US_Yes']
score_df = pd.DataFrame(list(zip(scores, col_names)),
columns =['Score', 'Feature'])
score_df
data_model = data[['Sales', 'Price', 'Advertising', 'Income', 'Age', 'ShelveLoc_Bad', 'ShelveLoc_Good', 'ShelveLoc_Medium']]
data_model.head()
X = data_model.iloc[:, 1:]
Y = data['Sales']
x_train, x_test, y_train, y_test = train_test_split(X, Y, test_size=0.33, random_state=42)
print(x_train.shape)
print(x_test.shape)
print(y_train.shape)
print(y_test.shape)
# +
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_val_score
from sklearn.ensemble import RandomForestClassifier
num_trees = 100
max_features = 'auto'
kfold = KFold(n_splits=10, random_state=42)
model = RandomForestClassifier(n_estimators=num_trees, max_features=max_features)
# -
model.fit(x_train, y_train)
results = cross_val_score(model, x_train, y_train, cv=kfold)
print(results.mean())
# +
from sklearn.ensemble import BaggingClassifier
kfold = KFold(n_splits=10, random_state=42)
cart = DecisionTreeClassifier()
num_trees = 100
model = BaggingClassifier(base_estimator=cart, n_estimators=num_trees, random_state=42)
results = cross_val_score(model, x_train, y_train, cv=kfold)
print(results.mean())
# +
from sklearn.ensemble import AdaBoostClassifier
kfold = KFold(n_splits=10, random_state=42)
model = AdaBoostClassifier(n_estimators=10, random_state=42)
results = cross_val_score(model, x_train, y_train, cv=kfold)
print(results.mean())
# -
| Assignment No-15( Random Forest, Company Data).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
df = pd.read_parquet("../data/brunch/session.parquet")
# +
import json
from tqdm import tqdm
with open("../data/dictionary.json") as fp:
dictionary = json.load(fp)
def make_label(x):
session = x["history"]
session = [dictionary[elem] for elem in session if elem in dictionary]
x["session"] = session
return x
tqdm.pandas()
df = df.progress_apply(make_label, axis=1)
# +
def make_mask(x):
session_length = len(x["session"])
# generating session mask
session_mask = [1.0] * (session_length - 1)
session_mask = [0.0] + session_mask
# generating user mask
user_mask = [0.0] * (session_length - 1)
user_mask = user_mask + [1.0]
x["session_mask"] = session_mask
x["user_mask"] = user_mask
return x
df = df.progress_apply(make_mask, axis=1)
# -
#000127ad0f1981cae1292efdb228f0e9
df_train = pd.read_parquet("../data/train.parquet")
sample = df_train[df_train.id == "#000549d84169355d490b029755f99381"]
session_mask = sample["session_mask"].values
session_mask
# +
import numpy as np
elements = [elem for elem in session_mask]
array = np.concatenate(elements, axis=None)
array
# -
sample
| notebook/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import nltk
from collections import Counter
from sklearn.metrics import log_loss
from scipy.optimize import minimize
import multiprocessing
import difflib
import time
import xgboost as xgb
from sklearn.cross_validation import train_test_split
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
# -
def get_train():
keras_q1 = np.load('../../data/transformed/keras_tokenizer/train_q1_transformed.npy')
keras_q2 = np.load('../../data/transformed/keras_tokenizer/train_q2_transformed.npy')
xgb_feats = pd.read_csv('../../data/features/the_1owl/owl_train.csv')
abhishek_feats = pd.read_csv('../../data/features/abhishek/train_features.csv',
encoding = 'ISO-8859-1').iloc[:, 2:]
text_feats = pd.read_csv('../../data/features/other_features/text_features_train.csv',
encoding = 'ISO-8859-1')
img_feats = pd.read_csv('../../data/features/other_features/img_features_train.csv')
srk_feats = pd.read_csv('../../data/features/srk/SRK_grams_features_train.csv')
xgb_feats.drop(['z_len1', 'z_len2', 'z_word_len1', 'z_word_len2'], axis = 1, inplace = True)
y_train = xgb_feats['is_duplicate']
xgb_feats = xgb_feats.iloc[:, 8:]
X_train2 = np.concatenate([xgb_feats, abhishek_feats], axis = 1)
#X_train2 = np.concatenate([keras_q1, keras_q2, xgb_feats, abhishek_feats, text_feats], axis = 1)
for i in range(X_train2.shape[1]):
if np.sum(X_train2[:, i] == y_train.values) == X_train2.shape[0]:
print('LEAK FOUND')
X_train2 = X_train2.astype('float32')
X_train2 = pd.DataFrame(X_train2)
X_train2['is_duplicate'] = y_train
print('Training data shape:', X_train2.shape)
return X_train2, y_train
train = pd.read_csv('../../data/features/the_1owl/owl_train.csv')
pos_train = train[train['is_duplicate'] == 1]
neg_train = train[train['is_duplicate'] == 0]
p = 0.165
scale = ((len(pos_train) / (len(pos_train) + len(neg_train))) / p) - 1
while scale > 1:
neg_train = pd.concat([neg_train, neg_train])
scale -=1
neg_train = pd.concat([neg_train, neg_train[:int(scale * len(neg_train))]])
train = pd.concat([pos_train, neg_train])
# +
x_train, x_valid, y_train, y_valid = train_test_split(train.iloc[:, 8:], train['is_duplicate'],
test_size=0.2, random_state=0)
params = {}
params["objective"] = "binary:logistic"
params['eval_metric'] = 'logloss'
params["eta"] = 0.05
params["subsample"] = 0.7
params["min_child_weight"] = 1
params["colsample_bytree"] = 0.7
params["max_depth"] = 4
params["silent"] = 1
params["seed"] = 1632
params['nthread'] = 6
d_train = xgb.DMatrix(x_train, label=y_train)
d_valid = xgb.DMatrix(x_valid, label=y_valid)
watchlist = [(d_train, 'train'), (d_valid, 'valid')]
bst = xgb.train(params, d_train, 10000, watchlist, early_stopping_rounds=100, verbose_eval=100)
print(log_loss(train.is_duplicate, bst.predict(xgb.DMatrix(train[col]))))
# + active=""
# pos_train = X_train[X_train['is_duplicate'] == 1]
# neg_train = X_train[X_train['is_duplicate'] == 0]
# p = 0.165
# scale = ((len(pos_train) / (len(pos_train) + len(neg_train))) / p) - 1
# while scale > 1:
# neg_train = pd.concat([neg_train, neg_train])
# scale -=1
# neg_train = pd.concat([neg_train, neg_train[:int(scale * len(neg_train))]])
# X_train2 = pd.concat([pos_train, neg_train])
#
# +
def kappa(preds, y):
score = []
a = 0.165 / 0.37
b = (1 - 0.165) / (1 - 0.37)
for pp,yy in zip(preds, y.get_label()):
score.append(a * yy * np.log (pp) + b * (1 - yy) * np.log(1-pp))
score = -np.sum(score) / len(score)
return 'kappa', score
params = {
'seed': 1337,
'colsample_bytree': 0.7,
'silent': 1,
'subsample': 0.7,
'eta': 0.05,
'objective': 'binary:logistic',
'eval_metric': 'logloss',
'max_depth': 4,
'min_child_weight': 1,
'nthread': 6,
}
X_tr, X_val, y_tr, y_val = train_test_split(X_train2.iloc[:, 8:], X_train2['is_duplicate'],
test_size = 0.2, random_state = 111)
dtrain = xgb.DMatrix(X_tr, label = y_tr)
dval = xgb.DMatrix(X_val, label = y_val)
watchlist = [(dtrain, 'train'), (dval, 'valid')]
bst = xgb.train(params, dtrain, 100000, watchlist, early_stopping_rounds=100, verbose_eval=50)
# feval = kappa)
# -
| models/.ipynb_checkpoints/19.04 - XGB Debugging-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Labels And Colors
#
#
# Draw a graph with matplotlib, color by degree.
#
# You must have matplotlib for this to work.
#
# +
# Author: <NAME> (<EMAIL>)
import matplotlib.pyplot as plt
import networkx as nx
G = nx.cubical_graph()
pos = nx.spring_layout(G) # positions for all nodes
# nodes
nx.draw_networkx_nodes(G, pos,
nodelist=[0, 1, 2, 3],
node_color='r',
node_size=500,
alpha=0.8)
nx.draw_networkx_nodes(G, pos,
nodelist=[4, 5, 6, 7],
node_color='b',
node_size=500,
alpha=0.8)
# edges
nx.draw_networkx_edges(G, pos, width=1.0, alpha=0.5)
nx.draw_networkx_edges(G, pos,
edgelist=[(0, 1), (1, 2), (2, 3), (3, 0)],
width=8, alpha=0.5, edge_color='r')
nx.draw_networkx_edges(G, pos,
edgelist=[(4, 5), (5, 6), (6, 7), (7, 4)],
width=8, alpha=0.5, edge_color='b')
# some math labels
labels = {}
labels[0] = r'$a$'
labels[1] = r'$b$'
labels[2] = r'$c$'
labels[3] = r'$d$'
labels[4] = r'$\alpha$'
labels[5] = r'$\beta$'
labels[6] = r'$\gamma$'
labels[7] = r'$\delta$'
nx.draw_networkx_labels(G, pos, labels, font_size=16)
plt.axis('off')
plt.show()
| NoSQL/NetworkX/plot_labels_and_colors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Calculate numbers for subject selection flow chart
# +
import numpy as np
import pandas as pd
import src.data.preprocess_data as prep
import src.data.var_names as var_names
from definitions import REPO_ROOT, PROCESSED_DATA_DIR, RAW_DATA_DIR
# -
# Seed for one child selection was 77 in our study
seed = 77
# +
binary_diagnoses_df = prep.create_binary_diagnoses_df(RAW_DATA_DIR)
sri24_df = prep.load_sri24_df(RAW_DATA_DIR)
freesurfer_df = prep.load_freesurfer_df(RAW_DATA_DIR)
sociodem_df = prep.load_sociodem_df(RAW_DATA_DIR)
print(f"Number of subjects with...")
print(f" ...complete SRI24 data: {len(sri24_df.dropna())}")
abcd_data_df = sri24_df.merge(
right=freesurfer_df, how='inner', left_index=True, right_index=True
)
print(f" ...complete SRI24 and Freesurfer data: {len(abcd_data_df.dropna())}")
abcd_data_df = abcd_data_df.merge(
right=sociodem_df, how='inner', left_index=True, right_index=True
)
print(f" ...complete SRI24, Freesurfer, and sociodemographic data: {len(abcd_data_df.dropna())}")
abcd_data_df = abcd_data_df.merge(
right=binary_diagnoses_df, how='inner', left_index=True, right_index=True
)
print(f" ...complete SRI24, Freesurfer, sociodemographic, and KSADS data: {len(abcd_data_df.dropna())}")
abcd_data_df = prep.select_one_child_per_family(
abcd_data_path=RAW_DATA_DIR,
abcd_df=abcd_data_df,
random_state=seed
)
print(f" ...with only one subject per family: {len(abcd_data_df.dropna())}")
| notebooks/reports/0.2-rg-subject_selection_flowchart.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="zewQ8m6diTS8"
# ## Tutorial 3: Demonstration of developing original *Agent* with DRL
# This tutorial demonstrate how to develop *Agent* with DRL algorithm by using ***KSPDRLAgent*** .
#
# *Agent* base classes are as follows:
#
# - `Agent`(used in **Tutorial 2**)
# - `KSPAgent`(used in **Tutorial 2**)
# - `PrioritizedKSPAgent`(used in **Tutorial 2**)
# - `KSPDRLAgent`
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} executionInfo={"elapsed": 48939, "status": "ok", "timestamp": 1606444394375, "user": {"displayName": "\u4e0b\u7530\u5c06\u4e4b", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhBSV0yzOkNvYKgj70Klrh7A9Vq3AJsnlWlPn1e=s64", "userId": "06013661169560345566"}, "user_tz": -540} id="P8UGHoZ6iVV2" outputId="960c37fe-e824-4574-a452-225aa73713be"
# !pip install git+https://github.com/Optical-Networks-Group/rsa-rl.git
# + [markdown] id="X__XTKCciTS8"
# ## Evaluation Settings
# For evaluation, prepare *Environment* and evaluation function.
# Please see **Tutorial 1** if you have not seen it.
# + executionInfo={"elapsed": 54422, "status": "ok", "timestamp": 1606444399868, "user": {"displayName": "\u4e0b\u7530\u5c06\u4e4b", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhBSV0yzOkNvYKgj70Klrh7A9Vq3AJsnlWlPn1e=s64", "userId": "06013661169560345566"}, "user_tz": -540} id="89r63g3KiTS9"
import functools
import numpy as np
from rsarl.envs import DeepRMSAEnv, make_multiprocess_vector_env
from rsarl.requester import UniformRequester
from rsarl.networks import SingleFiberNetwork
from rsarl.evaluator import batch_warming_up, batch_evaluation, batch_summary
# + executionInfo={"elapsed": 54421, "status": "ok", "timestamp": 1606444399873, "user": {"displayName": "\u4e0b\u7530\u5c06\u4e4b", "photoUrl": "https://lh3.googleusercontent.com/a-/AO<KEY>e=s64", "userId": "06013661169560345566"}, "user_tz": -540} id="l1v4_3VuiTS9"
# Set the device id to use GPU. To use CPU only, set it to -1.
gpu = -1
# + executionInfo={"elapsed": 828, "status": "ok", "timestamp": 1606444612598, "user": {"displayName": "\u4e0b\u7530\u5c06\u4e4b", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhBSV0yzOkNvYKgj70Klrh7A9Vq3AJsnlWlPn1e=s64", "userId": "06013661169560345566"}, "user_tz": -540} id="YPCJzzqPiTS9"
# exp settings
n_requests = 100
n_envs, seed = 2, 0
# build network
net = SingleFiberNetwork("nsf", n_slot=60, is_weight=True)
# build requester
requester = UniformRequester(
net.n_nodes,
avg_service_time=10,
avg_request_arrival_rate=12)
# build env
env = DeepRMSAEnv(net, requester)
# envs for training and evaluation
envs = make_multiprocess_vector_env(env, n_envs, seed, test=False)
test_envs = make_multiprocess_vector_env(env, n_envs, seed, test=True)
# + executionInfo={"elapsed": 669, "status": "ok", "timestamp": 1606444613797, "user": {"displayName": "\u4e0b\u7530\u5c06\u4e4b", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhBSV0yzOkNvYKgj70Klrh7A9Vq3AJsnlWlPn1e=s64", "userId": "06013661169560345566"}, "user_tz": -540} id="ixJW1uumiTS9"
def _evaluation(envs, agent, n_requests):
# start simulation
envs.reset()
#
batch_warming_up(envs, agent, n_requests=3000)
# evaluation
experiences = batch_evaluation(envs, agent, n_requests=n_requests)
# calc performance
blocking_probs, avg_utils, total_rewards = batch_summary(experiences)
for env_id, (blocking_prob, avg_util, total_reward) in enumerate(zip(blocking_probs, avg_utils, total_rewards)):
print(f'[{env_id}-th ENV]Blocking Probability: {blocking_prob}')
print(f'[{env_id}-th ENV]Avg. Slot-utilization: {avg_util}')
print(f'[{env_id}-th ENV]Total Rewards: {total_reward}')
# evaluation with test environments
evaluation = functools.partial(_evaluation, envs=test_envs, n_requests=n_requests)
# + [markdown] id="MQd2PXftiTS9"
# ## Step1: Select DRL algorithm from PFRL
# *RSA-RL* assumes that DRL algorithm provided by [PFRL](https://github.com/pfnet/pfrl) library is used.
# ***PFRL*** is a DRL library that implements various state-of-the-art deep reinforcement algorithms in Python using[PyTorch](https://github.com/pytorch/pytorch).
# Discrete action algorithms are as follows:
#
# - ***DQN(Double DQN)***
# - ***Rainbow***
# - ***IQN***
# - ***A3C***, ***A2C***
# - ***ACER***
# - ***PPO***
# - ***TRPO***
#
# In this tutorial, we try to reproduct the prior [DeepRMSA](https://ieeexplore.ieee.org/document/8386173) that applies DRL to ***routing algorithm*** that selects one from the *k* shortest paths.
# This tutorial call it ***DeepRMSAv1***, and implement it by using ***Double DQN (DDQN)***.
# In the case of using DDQN, there are three steps:
#
# 1. Build deep neural network (DNN) model
# 2. Specify ***Explore*** and ***Replay Buffer***, e.g., epsilon greedy and prioritized replay buffer, respectively
# 3. Build DDQN
#
# First, you develop a DNN that the number of outputs is *k*.
# + executionInfo={"elapsed": 54408, "status": "ok", "timestamp": 1606444399876, "user": {"displayName": "\u4e0b\u7530\u5c06\u4e4b", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhBSV0yzOkNvYKgj70Klrh7A9Vq3AJsnlWlPn1e=s64", "userId": "06013661169560345566"}, "user_tz": -540} id="P7ExKbXSiTS9"
import pfrl
import torch
import torch.nn as nn
# + executionInfo={"elapsed": 54404, "status": "ok", "timestamp": 1606444399877, "user": {"displayName": "\u4e0b\u7530\u5c06\u4e4b", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhBSV0yzOkNvYKgj70Klrh7A9Vq3AJsnlWlPn1e=s64", "userId": "06013661169560345566"}, "user_tz": -540} id="fiBR68HUiTS9"
class DeepRMSAv1_DNN(torch.nn.Module):
def __init__(self, SLOT: int, ICH: int, K: int, n_edges: int):
super().__init__()
self.SLOT = SLOT
# CNN
self.conv = nn.Sequential(*[
nn.Conv2d(ICH, 1, kernel_size=(1,1), stride=(1, 1)),
nn.ReLU(),
# 2 conv layers with16 filters
nn.Conv2d(1, 16, kernel_size=(n_edges,1), stride=(1, 1)),
nn.ReLU(),
nn.Conv2d(16, 16, kernel_size=(1,1), stride=(1, 1)),
nn.ReLU(),
# 2 depthwise conv layers with 1 filter
nn.ZeroPad2d((1, 0, 0, 0)), # left, right, top, bottom
nn.Conv2d(16, 16, kernel_size=(1,2), stride=(1, 1), groups=16),
nn.ReLU(),
nn.ZeroPad2d((1, 0, 0, 0)),
nn.Conv2d(16, 16, kernel_size=(1,2), stride=(1, 1), groups=16),
nn.ReLU(),
])
# fc
self.fc = nn.Sequential(*[
nn.Linear(SLOT*16, 128),
nn.ReLU(),
nn.Linear(128, 50),
nn.ReLU(),
nn.Linear(50, K),
])
def forward(self, x):
h = x
h = self.conv(h)
h = h.view(-1, self.SLOT*16)
h = self.fc(h)
return pfrl.action_value.DiscreteActionValue(h)
# + executionInfo={"elapsed": 54401, "status": "ok", "timestamp": 1606444399877, "user": {"displayName": "\u4e0b\u7530\u5c06\u4e4b", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhBSV0yzOkNvYKgj70Klrh7A9Vq3AJsnlWlPn1e=s64", "userId": "06013661169560345566"}, "user_tz": -540} id="RJICX7dIiTS9"
# Experimental Settings
K = 5
# slot-table(1) + one-hot-node * 2 + bandwidth(1)
ICH = 1 + 2 * net.n_nodes + 1
# build DNN for Q-function
q_func = DeepRMSAv1_DNN( net.n_slot, ICH, K, net.n_edges)
# Specify optimizer
optimizer = torch.optim.Adam(q_func.parameters(), eps=1e-2)
# + [markdown] id="LeNbJNARiTS9"
# ### Specify *Explore* and *Replay Buffer*
# This tutorial selects ConstantEpsilonGreedy.
# If you want to use others, please refere *PFRL*'s documentation:
# - [explore](https://pfrl.readthedocs.io/en/latest/explorers.html)
# - [replay buffer](https://pfrl.readthedocs.io/en/latest/replay_buffers.html)
# + executionInfo={"elapsed": 54398, "status": "ok", "timestamp": 1606444399878, "user": {"displayName": "\u4e0b\u7530\u5c06\u4e4b", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhBSV0yzOkNvYKgj70Klrh7A9Vq3AJsnlWlPn1e=s64", "userId": "06013661169560345566"}, "user_tz": -540} id="HvlobeuOiTS9"
def _action_sampler(k):
return np.random.randint(0, k)
# random action function
action_sampler = functools.partial(_action_sampler, k=K)
# + executionInfo={"elapsed": 54395, "status": "ok", "timestamp": 1606444399879, "user": {"displayName": "\u4e0b\u7530\u5c06\u4e4b", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhBSV0yzOkNvYKgj70Klrh7A9Vq3AJsnlWlPn1e=s64", "userId": "06013661169560345566"}, "user_tz": -540} id="IQCjTnKeiTS-"
# Set the discount factor that discounts future rewards.
gamma = 0.99
# Use epsilon-greedy for exploration
explorer = pfrl.explorers.ConstantEpsilonGreedy(
epsilon=0.1, random_action_func=action_sampler)
# DQN uses Experience Replay.
# Specify a replay buffer and its capacity.
replay_buffer = pfrl.replay_buffers.ReplayBuffer(capacity=10 ** 6, num_steps=50)
# + [markdown] id="I8OSpz80iTS-"
# ### Build DDQN
# NOTE that since DeepRMSAv1 does not show sufficient information of hyper parameter,
# we cannot reproduct it precisely.
# + executionInfo={"elapsed": 65056, "status": "ok", "timestamp": 1606444410544, "user": {"displayName": "\u4e0b\u7530\u5c06\u4e4b", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhBSV0yzOkNvYKgj70Klrh7A9Vq3AJsnlWlPn1e=s64", "userId": "06013661169560345566"}, "user_tz": -540} id="1f1xA2nliTS-"
# Now create an agent that will interact with the environment.
DDQN = pfrl.agents.DQN(
q_func,
optimizer,
replay_buffer,
gamma,
explorer,
minibatch_size=50,
update_interval=1,
replay_start_size=500,
target_update_interval=100,
gpu=gpu,
)
# + [markdown] id="3iyucmBniTS-"
# ## Step 2: Develop your algorithm by using *KSPDRLAgent*
# *RSA-RL* provides ***KSPDRLAgent*** that is based on *KSPAgent* class, which means that ***k-shortest path table*** can be used.
# You need to override two methods:
# - `preprocess`: create *feature vector* from *observation*
# - `map_drlout_to_action`: map outputs of DRL algorithms to *Action*
# + executionInfo={"elapsed": 65058, "status": "ok", "timestamp": 1606444410549, "user": {"displayName": "\u4e0b\u7530\u5c06\u4e4b", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhBSV0yzOkNvYKgj70Klrh7A9Vq3AJsnlWlPn1e=s64", "userId": "06013661169560345566"}, "user_tz": -540} id="dZMJKGKciTS-"
import numpy as np
import networkx as nx
from rsarl.data import Action
from rsarl.agents import KSPDRLAgent
from rsarl.utils import cal_slot, sort_tuple
from rsarl.algorithms import SpectrumAssignment
# + executionInfo={"elapsed": 65056, "status": "ok", "timestamp": 1606444410550, "user": {"displayName": "\u4e0b\u7530\u5c06\u4e4b", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhBSV0yzOkNvYKgj70Klrh7A9Vq3AJsnlWlPn1e=s64", "userId": "06013661169560345566"}, "user_tz": -540} id="8xmSD5xhiTS-"
def vectorize(n_nodes: int, node_id: int):
mp = np.eye(n_nodes, dtype=np.float32)[node_id].reshape(-1, 1, 1)
return mp
class DRLAgent(KSPDRLAgent):
def preprocess(self, obs):
"""
"""
net = obs.net
source, destination, bandwidth, duration = obs.request
# slot table
whole_slot = np.array(list(nx.get_edge_attributes(net.G, name="slot").values()))
whole_slot = whole_slot.reshape(1, net.n_edges, net.n_slot).astype(np.float32)
# source, destination, bandwidth map
smap = np.ones_like(whole_slot) * vectorize(net.n_nodes, source)
dmap = np.ones_like(whole_slot) * vectorize(net.n_nodes, destination)
bmap = np.ones_like(whole_slot) * bandwidth
# concate: (1, ICH, #edges, #slots)
fvec = np.concatenate([whole_slot, smap, dmap, bmap], axis=0)
return fvec.astype(np.float32, copy=False)
def map_drlout_to_action(self, obs, out):
net = obs.net
s, d, bandwidth, duration = obs.request
paths = self.path_table[sort_tuple((s, d))]
# map
path = paths[out]
#required slots
path_len = net.distance(path)
n_req_slot = cal_slot(bandwidth, path_len)
#FF
path_slot = net.path_slot(path)
slot_index = SpectrumAssignment.first_fit(path_slot, n_req_slot)
if slot_index is None:
return None
else:
return Action(path, slot_index, n_req_slot, duration)
# + executionInfo={"elapsed": 65053, "status": "ok", "timestamp": 1606444410550, "user": {"displayName": "\u4e0b\u7530\u5c06\u4e4b", "photoUrl": "https://lh3.googleusercontent.com/a-/<KEY>e=s64", "userId": "06013661169560345566"}, "user_tz": -540} id="BHzndvTQiTS-"
agent = DRLAgent(k=5, drl=DDQN)
# prepare path table
agent.prepare_ksp_table(net)
# + [markdown] id="Kho8Y5DJiTS-"
# ## Step 3: Training and Evaluate *DRL Agent*
# Finally, let's training and evaluation!
# Interaction between *Agent* with *Environment* automatically trains *Agent*.
# NOTE that before evaluation, you should change DRL model to ***evaluation mode*** by `eval_mode` method that *explore* does not run.
# + colab={"base_uri": "https://localhost:8080/"} id="haDhbU40iTS-" outputId="8aed44be-799c-48d4-97cf-d4c1592e3049"
# Batch act
obses = envs.reset()
resets = [False for _ in range(len(obses))]
for train_cnt in range(200000):
acts = agent.batch_act(obses)
obses, rews, dones, infos = envs.step(acts)
agent.batch_observe(obses, rews, dones, resets)
# Make mask(not_end). 0 if done/reset, 1 if pass
not_end = np.logical_not(dones)
obses = envs.reset(not_end)
if train_cnt % 20000 == 0:
print(f'[{train_cnt}-th EVAL]')
test_envs.reset()
with agent.drl.eval_mode():
evaluation(agent=agent)
# + [markdown] id="XI9wtQr4iTS_"
# ## Conclusion
# + [markdown] id="tYEZwr7biTS_"
# That's all!
# This tutorial demonstrates how to develop DRL *Agent*.
# Next tutorial demonstrate how to develop your own ***Environment***.
# + executionInfo={"elapsed": 150595, "status": "ok", "timestamp": 1606444496103, "user": {"displayName": "\u4e0b\u7530\u5c06\u4e4b", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhBSV0yzOkNvYKgj70Klrh7A9Vq3AJsnlWlPn1e=s64", "userId": "06013661169560345566"}, "user_tz": -540} id="2gkjt9KIiTS_"
| 03_DRL_Agent_en.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DataDictionary_RDF_Data_Cube2
#
# This Notebook steps through the development of a method to convert a UKDS DataDictionary .rtf file to a RDF file using the Data Cube vocabulary https://www.w3.org/TR/vocab-data-cube/
# ## Initial setup
# ### Import packages
import os, ukds
import pandas as pd
from fairly import CreateRDF, Fuseki
# ### Set filepaths
#
# This sets a filepath to an example data dictionary on a local file system, in this case the 'uktus15_household_ukda_data_dictionary.rtf' file.
base_dir=os.path.join(*[os.pardir]*4,r'_Data\United_Kingdom_Time_Use_Survey_2014-2015\UKDA-8128-tab')
dd_fp=os.path.join(base_dir,r'mrdoc\allissue\uktus15_household_ukda_data_dictionary.rtf')
# ### Create DataDictionary
#
# A ukds.DataDictionary instance is created and the .rtf file is read into it.
dd=ukds.DataDictionary()
dd.read_rtf(dd_fp)
dd.variable_list[0:1]
# ## Discussion
# ### Aim
#
# The aim of this notebook is to develop a method to convert the information in UKDS data dictionary files into RDF format based on RDF Data Cube vocabulary https://www.w3.org/TR/vocab-data-cube/#schemes.
#
# Once converted this RDF data can be combined with RDF Data Cube for the UKDS data table files.
# ### Sample call
#
# Sample code could look like:
#
# ```python
# t=dd.to_rdf_data_cube() # dd is a DataDictionary instance
# ```
#
# where t is a string of a turtle file.
# ## Method development
# ### Setup
c=CreateRDF()
c.add_data_cube_prefixes()
c.add_skos_prefixes()
c.add_prefix('ukds8128','<http://purl.org/berg/ukds8128/>')
c.add_prefix('ukds8128-concept','<http://purl.org/berg/ukds8128/concept/>')
# ### Inputs
row_dimension_property_uri='ukds:household'
row_dimension_property_label='"the Household serial number"'
row_concept_uri='ukds8128-concept:household'
row_concept_prefLabel='"A household"'
# ### Dimensions
#
# In Data Cube, each observation has an associated set of dimensions and measures.
#
# Here, each single value (or cell) in a UKDS data table is converted to a single qb:Observation. This is the 'measure dimension' appraoch.
#
# What are the dimensions and measures for each observation?
# - one dimension is the table row. In UKDS data tables each row normally represents an entity (such as individual, household etc). The entity can be specified with a uri and the observation linked to it.
# - a second dimension is the table column. In UKDS data tables the columns are the type of observation made, i.e. what is measured. This can correspond to a measure dimension for each observation.
# #### Table row dimension
#
# If each row is an entity of a class (such as individuals or households), then this class should be defined as a skos:Concept.
#
# In our example, each row of the 'uktus15_household.tab' data table is a household.
# +
def add_skos_table_row_concept(self,
c,
row_concept_uri,
row_concept_prefLabel):
"""Adds a skos:Concept for the table row dimension
"""
return c.add_skos_concept(
concept_uri=row_concept_uri,
topConceptOf_uri=None,
prefLabel=row_concept_prefLabel,
notation=None,
inScheme_uri=None,
predicate_object_list=None
)
print(add_skos_table_row_concept(dd,c,row_concept_uri,row_concept_prefLabel))
# -
# This concept is then used as a reference for the dimension property
# +
def add_data_cube_table_row_dimension_property(self,
c,
row_dimension_property_uri,
row_dimension_property_label,
row_concept_uri,
):
"""Adds a qb:DimensionProperty for the tabel row dimension
"""
return c.add_data_cube_dimension_property(
dimension_property_uri=row_dimension_property_uri,
label=row_dimension_property_label,
subPropertyOf_uri=None,
range_uri=None,
concept_uri=row_concept_uri,
code_list_uri=None,
predicate_object_list=None)
print(add_data_cube_table_row_dimension_property(dd,
c,
row_dimension_property_uri,
row_dimension_property_label,
row_concept_uri))
# -
# #### Table column dimensions (measure dimension)
# +
def add_skos_table_column_concept(self,
c,
column_concept_uri,
column_concept_prefLabel,
column_concept_notation,
column_concept_comment
):
"""Adds a skos:Concept for the table row dimension
"""
return c.add_skos_concept(
concept_uri=column_concept_uri,
topConceptOf_uri=None,
prefLabel=column_concept_prefLabel,
notation=column_concept_notation,
inScheme_uri=None,
predicate_object_list=[('rdfs:comment',column_concept_comment)]
)
column_concept_uri='ukds-concept:serial'
column_concept_prefLabel='"Household number"'
column_concept_notation='"serial"'
column_concept_comment='"%s"' % str({'pos': '1',
'variable': 'serial',
'variable_label': 'Household number',
'variable_type': 'numeric',
'SPSS_measurement_level': 'SCALE',
'SPSS_user_missing_values': '',
'value_labels': ''})
print(add_skos_table_column_concept(dd,
c,
column_concept_uri,
column_concept_prefLabel,
column_concept_notation,
column_concept_comment))
# -
# ### Sample RDF file
#
# The RDF Data Cube webpage gives, in Example 14, an example of a concept (or variable) and it's code list using the `skos` vocabulary.
#
# The proposal is the RDF file would look as below. This shows the data in turtle (.ttl) format for the variable *serial*:
#
# ```turtle
# @prefix qb: <http://purl.org/linked-data/cube#> .
# @prefix skos: <http://www.w3.org/2004/02/skos/core#> .
# @prefix rdf: <http://www.w3.org/1999/02/22-rdf-syntax-ns#> .
# @prefix rdfs: <http://www.w3.org/2000/01/rdf-schema#> .
# @prefix xsd: <http://www.w3.org/2001/XMLSchema#> .
# @prefix ukds8128-code: <http://purl.org/berg/ukds8128/code/> . # a prefix for the UKDS Time Use Survey 2014-2015 dataset
# @prefix ukds8128-measure: <http://purl.org/berg/ukds8128/measure/> .
#
# ukds8128-measure:serial a rdf:Property, qb:MeasureProperty ;
# rdfs:label "serial"@en ;
# rdfs:subPropertyOf sdmx-measure:obsValue ;
# rdfs:range xsd:decimal .
#
# ukds8128-code:serial a skos:ConceptScheme ;
# skos:prefLabel "serial"@en ; # the 'variable' value
# rdfs:label "serial"@en ; # the 'variable' value
# skos:notation "serial" ; # the 'variable' value
# skos:note "Household number."@en ; # the 'variable_label' value
# skos:definition <ukds8128:uktus15_household_ukda_data_dictionary> ; # a uri based on the file name
# .
# ```
# This shows the data in turtle (.ttl) format for the variable *strata*:
#
# ```turtle
# @prefix qb: <http://purl.org/linked-data/cube#> .
# @prefix skos: <http://www.w3.org/2004/02/skos/core#> .
# @prefix rdfs: <http://www.w3.org/2000/01/rdf-schema#> .
# @prefix ukds8128: <http://purl.org/berg/ukds8128/> .
#
# ukds8128-code:strata a skos:ConceptScheme ;
# skos:prefLabel "strata"@en ;
# rdfs:label "strata"@en ;
# skos:notation "strata" ;
# skos:note "Strata"@en ;
# skos:definition <ukds8128:uktus15_household_ukda_data_dictionary> ;
# rdfs:seeAlso ukds8128-code:Strata ;
# skos:hasTopConcept ukds8128-code:strata_code_-2.0 .
#
# ukds8128-code:Strata a rdfs:Class, owl:Class;
# rdfs:subClassOf skos:Concept ;
# rdfs:label "strata"@en;
# rdfs:comment "Strata"@en;
# rdfs:seeAlso ukds8128-code:strata .
#
# ukds8128-code:strata_code_-2.0 a skos:Concept, ukds8128-code:Strata;
# skos:topConceptOf ukds8128-code:strata;
# skos:prefLabel "Schedule not applicable"@en ;
# skos:notation -2.0 ;
# skos:inScheme ukds8128-code:strata .
#
# ```
#
# Here this includes a single 'value label' code.
#
# ## Developing the method
# ### to_data_structure_definition
# +
def to_data_structure_definition(self,prefix,dataset_name):
"""Returns a RDF Turtle string of the qb:DataStructureDefinition using the Data Cube and skos vocabulary
Arguments:
- self: the DataDictionary instance
- prefix (str): the prefix to use for the data dictionary uris
- dataset_name (str): the name of the dataset
"""
l=['%s:%s-dsd a qb:DataStructureDefinition' % (prefix,dataset_name)]
for i,variable in enumerate(dd.get_variable_names()):
l.append('qb:component [ qb:measure %s-measure:%s; qb:order %s ]' % (prefix,variable,i+1))
st=' ;\n\t'.join(l) + ' .\n\n'
return st
print(to_data_structure_definition(dd,'ukds8128','uktus15_household'))
# -
# ### to_measure_property
# +
def to_measure_property(self,variable,prefix,):
"""Returns a RDF Turtle string of the qb:MeaureProperty using the Data Cube and skos vocabulary
Arguments:
- self: the DataDictionary instance
- variable (str): the variable to convert to RDF
- prefix (str): the prefix to use for the data dictionary uris
"""
d=self.get_variable_dict(variable)
if d['value_labels']:
x=', qb:CodedProperty'
else:
x=''
l=[
'ukds8128-measure:%s a rdf:Property, qb:MeasureProperty%s' % (variable,x) ,
'rdfs:label "serial"@en' ,
'rdfs:subPropertyOf sdmx-measure:obsValue',
]
if d['value_labels']:
l.append('qb:CodeList %s-code:%s' % (prefix,variable))
l.append('rdfs:range %s-code:%s' % (prefix,variable[0].upper()+variable[1:]))
else:
if d['variable_type']=='numeric':
l.append('rdfs:range xsd:decimal')
st=' ;\n\t'.join(l) + ' .\n\n'
return st
print(to_measure_property(dd,'serial','ukds8128'))
print(to_measure_property(dd,'strata','ukds8128'))
# -
# ### to_codelist
# +
def to_codelist(self,variable,prefix,filename_no_ext):
"""Returns a RDF Turtle string of the codelist using the Data Cube and skos vocabulary
Arguments:
- self: the DataDictionary instance
- variable (str): the variable to convert to RDF
- filename_no_ext (str): the filename of the Data Dictionary file with no extension included.
- prefix (str): the prefix to use for the data dictionary uris
"""
d=self.get_variable_dict(variable)
variable_lower=d['variable'][0].lower() + d['variable'][1:]
variable_upper=d['variable'][0].upper() + d['variable'][1:]
# ConceptScheme
l=[]
l+=[
'%s-code:%s a skos:ConceptScheme' % (prefix,variable_lower),
'skos:prefLabel "%s"@en' % d['variable'],
'rdfs:label "%s"@en' % d['variable'],
'skos:notation "%s"' % d['variable'],
'skos:note "%s"@en' % d['variable_label'],
'skos:definition <%s:%s>' % (prefix,filename_no_ext),
]
if d['value_labels']:
l.append('rdfs:seeAlso %s-code:%s' % (prefix,variable_upper))
for k,v in d['value_labels'].items():
l.append('skos:hasTopConcept %s:%s_code_%s' % (prefix,variable_lower,k))
st=' ;\n\t'.join(l) + ' .\n\n'
# Code
if d['value_labels']:
l=[
'%s-code:%s a rdfs:Class, owl:Class ' % (prefix,variable_upper),
'rdfs:subClassOf skos:Concept ',
'rdfs:label "%s"@en ' % d['variable'],
'rdfs:comment "%s"@en ' % d['variable_label'],
'rdfs:seeAlso %s-code:%s ' % (prefix,variable_lower),
]
st+=' ;\n\t'.join(l) + ' .\n\n'
for k,v in d['value_labels'].items():
l=[
'%s-code:%s_code_%s a skos:Concept, %s-code:%s' % (prefix,variable_lower,k,prefix,variable_upper),
'skos:topConceptOf %s-code:%s' % (prefix,variable_lower),
'skos:prefLabel "%s"@en' % v,
'skos:notation %s' % k,
'skos:inScheme %s-code:%s' % (prefix,variable_lower),
]
st+=' ;\n\t'.join(l) + ' .\n\n'
return st
print(to_codelist(dd,'serial','ukds8128','uktus15_household_ukda_data_dictionary'))
print(to_codelist(dd,'strata','ukds8128','uktus15_household_ukda_data_dictionary'))
# -
def to_rdf_data_cube(self,prefix,base_uri,filename_no_ext,dataset_name):
"""Returns a RDF Turtle string using the Data Cube vocabulary
Arguments:
- self: the DataDictionary instance
- filename_no_ext (str): the filename of the Data Dictionary file with no extension included.
- prefix (str): the prefix to use for the data dictionary base uri
- base_uri (str): the data dictionary base uri
- dataset_name (str): the name of the dataset
"""
st="""
@prefix qb: <http://purl.org/linked-data/cube#> .
@prefix skos: <http://www.w3.org/2004/02/skos/core#> .
@prefix rdf: <http://www.w3.org/1999/02/22-rdf-syntax-ns#> .
@prefix rdfs: <http://www.w3.org/2000/01/rdf-schema#> .
@prefix xsd: <http://www.w3.org/2001/XMLSchema#> .
@prefix %s: %s> .
@prefix %s-code: %scode#> .
@prefix %s-measure: %smeasure#> .
""" % (prefix,base_uri,prefix,base_uri,prefix,base_uri)
st+=to_data_structure_definition(self,prefix,dataset_name)
for variable in dd.get_variable_names():
st+=to_measure_property(self,variable,prefix)
st+=to_codelist(self,variable,prefix,filename_no_ext)
break
return st
print(to_rdf_data_cube(dd,'ukds8128','<http://purl.org/berg/ukds8128/','uktus15_household_ukda_data_dictionary','uktus15_household'))
| development/DataDictionary_RDF_Data_Cube2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ML Course, Bogotá, Colombia (© <NAME>; June 2019)
# %run ../talktools.py
# ## Generative Adversarial Networks
#
# One of the downsides of VAEs is that the generated samples are interpolated between real samples as you walk through the latent space. This can lead to unrealistic looking images (what's half way between a shoe and a sweater?).
#
# <img src="imgs/gans.png">
# Source: <NAME>, DeepMind
#
# Another way to create non-parametric generative models--with more realistic imagined samples--is with GANs. GANs are the result of a competition between a network which tries to generate realistic data (generator) and a network which tries to get learn how to distinguish between fake data and real data from the sample (discriminator).
#
# A really good set of tutorials on GANs: https://sites.google.com/view/cvpr2018tutorialongans/
# ### The basic idea
#
# <img src="https://lilianweng.github.io/lil-log/assets/images/GAN.png">
#
# **Let's Check out an Online Demo**: https://poloclub.github.io/ganlab/
#
# <img src="https://raw.githubusercontent.com/hindupuravinash/the-gan-zoo/master/cumulative_gans.jpg">
# Source: https://github.com/hindupuravinash/the-gan-zoo
#
# <img src="imgs/faces.png">
# Source: Ian Goodfellow
#
#
# By Dec 2018, extremely photorealistic images were able to be generated. An example of the high-quality results from the latest GAN work, the following pictures are not of real people but instead are generated:
#
# <img src="https://cdn.technologyreview.com/i/images/screen-shot-2018-12-14-at-10.13.53-am.png?sw=2544&cx=0&cy=0&cw=594&ch=262">
#
# https://www.technologyreview.com/s/612612/these-incredibly-real-fake-faces-show-how-algorithms-can-now-mess-with-us/
#
# **Check this out**: https://thispersondoesnotexist.com/
# ## So What Good Can You Do with GANs?
#
# (We'll note some of the less ethical uses in practice today in the last Lecture)
#
# ** Useful to create inputs to simulations **
#
# <img src="http://www.yaronhadad.com/wp-content/uploads/2017/03/WEB_lead_Nguyen-and-Mandelbaum.jpg">
# Source: AI-generated images of galaxies (left, lower of each pair) and volcanoes. Left: Figure: S. Ravanbakhsh/data: arxiv.org/abs/1609.05796; Right: Nguyen et al./arxiv.org/abs/1612.00005
# ** Creativity and Marketing **
#
# <img src="https://github.com/maxorange/pix2vox/raw/master/img/sample.gif">
# <img src="https://github.com/maxorange/pix2vox/raw/master/img/single-category-generation.png">
# Pix2VOX: https://github.com/maxorange/pix2vox
#
# <img src="imgs/fashion.png">
# ** Inhancement / Superresolution **
#
# <img src="imgs/super.png">
| Lectures/6_GenerativeAndCompressiveModels/01_GenerativeAdversarialNetworks.ipynb |