code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#3.1 Linear Regression
#3.1.2 Vectorization for Speed
#比較 使用for迴圈一個一個加 及 使用向量加法 的時間
import math
import time
import numpy as np
import torch
from d2l import torch as d2l
n = 10000
a = torch.ones(n)
b = torch.ones(n)
# -
class Timer:
"""Record multiple running times."""
def __init__(self):
self.times = []
self.start()
def start(self):
"""Start the timer."""
self.tik = time.time()
def stop(self):
"""Stop the timer and record the time in a list."""
self.times.append(time.time() - self.tik)
return self.times[-1]
def avg(self):
"""Return the average time."""
return sum(self.times) / len(self.times)
def sum(self):
"""Return the sum of time."""
return sum(self.times)
def cumsum(self):
"""Return the accumulated time."""
return np.array(self.times).cumsum().tolist()
# +
c = torch.zeros(n)
timer = Timer()
#用for迴圈一個一個加
for i in range(n):
c[i] = a[i] + b[i]
f'{timer.stop():.5f} sec'
# +
timer.start()
#用向量加法全部一起加
d = a + b
f'{timer.stop():.5f} sec'
# +
#3.1.3 The Normal Distribution and Squared Loss
def normal(x, mu, sigma): #正態分佈function
p = 1 / math.sqrt(2 * math.pi * sigma**2)
return p * np.exp(-0.5 / sigma**2 * (x - mu)**2)
# +
# Use numpy again for visualization
x = np.arange(-7, 7, 0.01)
# Mean and standard deviation pairs
params = [(0, 1), (0, 2), (3, 1)]
d2l.plot(x, [normal(x, mu, sigma) for mu, sigma in params], xlabel='x',
ylabel='p(x)', figsize=(4.5, 2.5),
legend=[f'mean {mu}, std {sigma}' for mu, sigma in params])
# -
| coookie89/Week1/ch3/3.1/example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Asserting Expectations
#
# In the previous chapters on [tracing](Tracer.ipynb) and [interactive debugging](Debugger.ipynb), we have seen how to observe executions. By checking our observations against our expectations, we can find out when and how the program state is faulty. So far, we have assumed that this check would be done by _humans_ – that is, us. However, having this check done by a _computer_, for instance as part of the execution, is infinitely more rigorous and efficient. In this chapter, we introduce techniques to _specify_ our expectations and to check them at runtime, enabling us to detect faults _right as they occur_.
# -
from bookutils import YouTubeVideo
YouTubeVideo("9mI9sbKFkwU")
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# **Prerequisites**
#
# * You should have read the [chapter on tracing executions](Tracer.ipynb).
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
import bookutils
# -
from bookutils import quiz
import Tracer
# + [markdown] slideshow={"slide_type": "skip"}
# ## Synopsis
# <!-- Automatically generated. Do not edit. -->
#
# To [use the code provided in this chapter](Importing.ipynb), write
#
# ```python
# >>> from debuggingbook.Assertions import <identifier>
# ```
#
# and then make use of the following features.
#
#
# This chapter discusses _assertions_ to define _assumptions_ on function inputs and results:
#
# ```python
# >>> def my_square_root(x): # type: ignore
# >>> assert x >= 0
# >>> y = square_root(x)
# >>> assert math.isclose(y * y, x)
# >>> return y
# ```
# Notably, assertions detect _violations_ of these assumptions at runtime:
#
# ```python
# >>> with ExpectError():
# >>> y = my_square_root(-1)
# Traceback (most recent call last):
# File "<ipython-input-158-b63177b4c2f7>", line 2, in <module>
# y = my_square_root(-1)
# File "<ipython-input-157-dc1d0082e740>", line 2, in my_square_root
# assert x >= 0
# AssertionError (expected)
#
# ```
# _System assertions_ help to detect invalid memory operations.
#
# ```python
# >>> managed_mem = ManagedMemory()
# >>> managed_mem
# ```
# |Address|<span style="color: blue">0</span>|<span style="color: blue">1</span>|<span style="color: lightgrey">2</span>|<span style="color: lightgrey">3</span>|<span style="color: lightgrey">4</span>|<span style="color: lightgrey">5</span>|<span style="color: lightgrey">6</span>|<span style="color: lightgrey">7</span>|<span style="color: lightgrey">8</span>|<span style="color: lightgrey">9</span>|
# |:---|:---|:---|:---|:---|:---|:---|:---|:---|:---|:---|
# |Allocated| | | | | | | | | | |
# |Initialized| | | | | | | | | | |
# |Content|-1|0| | | | | | | | |
#
# ```
# >>> with ExpectError():
# >>> x = managed_mem[2]
# Traceback (most recent call last):
# File "<ipython-input-160-89a206319ebb>", line 2, in <module>
# x = managed_mem[2]
# File "<ipython-input-97-d25e50073b38>", line 3, in __getitem__
# return self.read(address)
# File "<ipython-input-138-7b46abe4a8e1>", line 10, in read
# "Reading from unallocated memory"
# AssertionError: Reading from unallocated memory (expected)
#
# ```
#
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# ## Introducing Assertions
#
# [Tracers](Tracer.ipynb) and [Interactive Debuggers](Debugger.ipynb) are very flexible tools that allow you to observe precisely what happens during a program execution. It is still _you_, however, who has to check program states and traces against your expectations. There is nothing wrong with that – except that checking hundreds of statements or variables can quickly become a pretty boring and tedious task.
#
# Processing and checking large amounts of data is actually precisely what _computers were invented for_. Hence, we should aim to _delegate such checking tasks to our computers_ as much as we can. This automates another essential part of debugging – maybe even _the_ most essential part.
# -
# ### Assertions
#
# The standard tool for having the computer check specific conditions at runtime is called an _assertion_. An assertion takes the form
#
# ```python
# assert condition
# ```
#
# and states that, at runtime, the computer should check that `condition` holds, e.g. evaluates to True. If the condition holds, then nothing happens:
assert True
# If the condition evaluates to _False_, however, then the assertion _fails_, indicating an internal error.
from ExpectError import ExpectError
with ExpectError():
assert False
# A common usage for assertions is for _testing_. For instance, we can test a square root function as
def test_square_root() -> None:
assert square_root(4) == 2
assert square_root(9) == 3
...
# and `test_square_root()` will fail if `square_root()` returns a wrong value.
# Assertions are available in all programming languages. You can even go and implement assertions yourself:
def my_own_assert(cond: bool) -> None:
if not cond:
raise AssertionError
# ... and get (almost) the same functionality:
with ExpectError():
my_own_assert(2 + 2 == 5)
# ### Assertion Diagnostics
# In most languages, _built-in assertions_ offer a bit more functionality than what can be obtained with self-defined functions. Most notably, built-in assertions
#
# * frequently tell _which condition_ failed (`2 + 2 == 5`)
# * frequently tell _where_ the assertion failed (`line 2`), and
# * are _optional_ – that is, they can be turned off to save computation time.
# C and C++, for instance, provide an `assert()` function that does all this:
# ignore
open('testassert.c', 'w').write(r'''
#include <stdio.h>
#include "assert.h"
int main(int argc, char *argv[]) {
assert(2 + 2 == 5);
printf("Foo\n");
}
''');
# ignore
from bookutils import print_content
print_content(open('testassert.c').read(), '.h')
# If we compile this function and execute it, the assertion (expectedly) fails:
# !cc -g -o testassert testassert.c
# !./testassert
# How would the C `assert()` function be able to report the condition and the current location? In fact, `assert()` is commonly implemented as a _macro_ that besides checking the condition, also turns it into a _string_ for a potential error message. Additional macros such as `__FILE__` and `__LINE__` expand into the current location and line, which can then all be used in the assertion error message.
# A very simple definition of `assert()` that provides the above diagnostics looks like this:
# ignore
open('assert.h', 'w').write(r'''
#include <stdio.h>
#include <stdlib.h>
#ifndef NDEBUG
#define assert(cond) \
if (!(cond)) { \
fprintf(stderr, "Assertion failed: %s, function %s, file %s, line %d", \
#cond, __func__, __FILE__, __LINE__); \
exit(1); \
}
#else
#define assert(cond) ((void) 0)
#endif
''');
print_content(open('assert.h').read(), '.h')
# (If you think that this is cryptic, you should have a look at an [_actual_ `<assert.h>` header file](https://github.com/bminor/glibc/blob/master/assert/assert.h).)
# This header file reveals another important property of assertions – they can be _turned off_. In C and C++, defining the preprocessor variable `NDEBUG` ("no debug") turns off assertions, replacing them with a statement that does nothing. The `NDEBUG` variable can be set during compilation:
# !cc -DNDEBUG -g -o testassert testassert.c
# And, as you can see, the assertion has no effect anymore:
# !./testassert
# In Python, assertions can also be turned off, by invoking the `python` interpreter with the `-O` ("optimize") flag:
# !python -c 'assert 2 + 2 == 5; print("Foo")'
# !python -O -c 'assert 2 + 2 == 5; print("Foo")'
# In comparison, which language wins in the amount of assertion diagnostics? Have a look at the information Python provides. If, after defining `fun()` as
def fun() -> None:
assert 2 + 2 == 5
quiz("If we invoke `fun()` and the assertion fails,"
" which information do we get?",
[
"The failing condition (`2 + 2 == 5`)",
"The location of the assertion in the program",
"The list of callers",
"All of the above"
], '123456789 % 5')
# Indeed, a failed assertion (like any exception in Python) provides us with lots of debugging information, even including the source code:
with ExpectError():
fun()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ## Checking Preconditions
#
# Assertions show their true power when they are not used in a test, but used in a program instead, because that is when they can check not only _one_ run, but actually _all_ runs.
# -
# The classic example for the use of assertions is a _square root_ program, implementing the function $\sqrt{x}$. (Let's assume for a moment that the environment does not already have one.)
# We want to ensure that `square_root()` is always called with correct arguments. For this purpose, we set up an assertion:
def square_root(x): # type: ignore
assert x >= 0
... # compute square root in y
# This assertion is called the _precondition_. A precondition is checked at the beginning of a function. It checks whether all the conditions for using the function are met.
# So, if we call `square_root()` with an bad argument, we will get an exception. This holds for _any_ call, ever.
with ExpectError():
square_root(-1)
# For a dynamically typed language like Python, an assertion could actually also check that the argument has the correct type. For `square_root()`, we could ensure that `x` actually has a numeric type:
def square_root(x): # type: ignore
assert isinstance(x, (int, float))
assert x >= 0
... # compute square root in y
# And while calls with the correct types just work...
square_root(4)
square_root(4.0)
# ... a call with an illegal type will raise a revealing diagnostic:
with ExpectError():
square_root('4')
quiz("If we did not check for the type of `x`, "
"would the assertion `x >= 0` still catch a bad call?",
[
"Yes, since `>=` is only defined between numbers",
"No, because an empty list or string would evaluate to 0"
], '0b10 - 0b01')
# Fortunately (for us Python users), the assertion `x >= 0` would already catch a number of invalid types, because (in contrast to, say, JavaScript), Python has no implicit conversion of strings or structures to integers:
with ExpectError():
'4' >= 0 # type: ignore
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ## Checking Results
#
# While a precondition ensures that the _argument_ to a function is correct, a _postcondition_ checks the _result_ of this very function (assuming the precondition held in the first place). For our `square_root()` function, we can check that the result $y = \sqrt{x}$ is correct by checking that $y^2 = x$ holds:
# -
def square_root(x): # type: ignore
assert x >= 0
... # compute square root in y
assert y * y == x
# In practice, we might encounter problems with this assertion. What might these be?
quiz("Why could the assertion fail despite `square_root()` being correct?",
[
"We need to compute `y ** 2`, not `y * y`",
"We may encounter rounding errors",
"The value of `x` may have changed during computation",
"The interpreter / compiler may be buggy"
], '0b110011 - 0o61')
# Technically speaking, there could be many things that _also_ could cause the assertion to fail (cosmic radiation, operating system bugs, secret service bugs, anything) – but the by far most important reason is indeed rounding errors. Here's a simple example, using the Python built-in square root function:
import math
math.sqrt(2.0) * math.sqrt(2.0)
math.sqrt(2.0) * math.sqrt(2.0) == 2.0
# If you want to compare two floating-point values, you need to provide an _epsilon value_ denoting the margin of error.
def square_root(x): # type: ignore
assert x >= 0
... # compute square root in y
epsilon = 0.000001
assert abs(y * y - x) < epsilon
# In Python, the function `math.isclose(x, y)` also does the job, by default ensuring that the two values are the same within about 9 decimal digits:
math.isclose(math.sqrt(2.0) * math.sqrt(2.0), 2.0)
# So let's use `math.isclose()` for our revised postcondition:
def square_root(x): # type: ignore
assert x >= 0
... # compute square root in y
assert math.isclose(y * y, x)
# Let us try out this postcondition by using an actual implementation. The [Newton–Raphson method](https://en.wikipedia.org/wiki/Newton%27s_method) is an efficient way to compute square roots:
def square_root(x): # type: ignore
assert x >= 0 # precondition
approx = None
guess = x / 2
while approx != guess:
approx = guess
guess = (approx + x / approx) / 2
assert math.isclose(approx * approx, x)
return approx
# Apparently, this implementation does the job:
square_root(4.0)
# However, it is not just this call that produces the correct result – _all_ calls will produce the correct result. (If the postcondition assertion does not fail, that is.) So, a call like
square_root(12345.0)
# does not require us to _manually_ check the result – the postcondition assertion already has done that for us, and will continue to do so forever.
# ### Assertions and Tests
#
# Having assertions right in the code gives us an easy means to _test_ it – if we can feed sufficiently many inputs into the code without the postcondition ever failing, we can increase our confidence. Let us try this out with our `square_root()` function:
for x in range(1, 10000):
y = square_root(x)
# Note again that we do not have to check the value of `y` – the `square_root()` postcondition already did that for us.
# Instead of enumerating input values, we could also use random (non-negative) numbers; even totally random numbers could work if we filter out those tests where the precondition already fails. If you are interested in such _test generation techniques_, the [Fuzzing Book](fuzzingbook.org) is a great reference for you.
# Modern program verification tools even can _prove_ that your program will always meet its assertions. But for all this, you need to have _explicit_ and _formal_ assertions in the first place.
# For those interested in testing and verification, here is a quiz for you:
quiz("Is there a value for x that satisfies the precondition, "
"but fails the postcondition?",
[
"Yes",
"No"
], 'int("Y" in "Yes")')
# This is indeed something a test generator or program verifier might be able to find with zero effort.
# ### Partial Checks
#
# In the case of `square_root()`, our postcondition is _total_ – if it passes, then the result is correct (within the `epsilon` boundaries, that is). In practice, however, it is not always easy to provide such a total check. As an example, consider our `remove_html_markup()` function from the [Introduction to Debugging](Intro_Debugging.ipynb):
def remove_html_markup(s): # type: ignore
tag = False
quote = False
out = ""
for c in s:
if c == '<' and not quote:
tag = True
elif c == '>' and not quote:
tag = False
elif c == '"' or c == "'" and tag:
quote = not quote
elif not tag:
out = out + c
return out
remove_html_markup("I am a text with <strong>HTML markup</strong>")
# The precondition for `remove_html_markup()` is trivial – it accepts any string. (Strictly speaking, a precondition `assert isinstance(s, str)` could prevent it from being called with some other collection such as a list.)
# The challenge, however, is the _postcondition_. How do we check that `remove_html_markup()` produces the correct result?
#
# * We could check it against some other implementation that removes HTML markup – but if we already do have such a "golden" implementation, why bother implementing it again?
#
# * After a change, we could also check it against some earlier version to prevent _regression_ – that is, losing functionality that was there before. But how would we know the earlier version was correct? (And if it was, why change it?)
# If we do not aim for ensuring full correctness, our postcondition can also check for _partial properties_. For instance, a postcondition for `remove_html_markup()` may simply ensure that the result no longer contains any markup:
def remove_html_markup(s): # type: ignore
tag = False
quote = False
out = ""
for c in s:
if c == '<' and not quote:
tag = True
elif c == '>' and not quote:
tag = False
elif c == '"' or c == "'" and tag:
quote = not quote
elif not tag:
out = out + c
# postcondition
assert '<' not in out and '>' not in out
return out
# Besides doing a good job at checking results, the postcondition also does a good job in documenting what `remove_html_markup()` actually does.
quiz("Which of these inputs causes the assertion to fail?",
[
'`<foo>bar</foo>`',
'`"foo"`',
'`>foo<`',
'`"x > y"`'
], '1 + 1 -(-1) + (1 * -1) + 1 ** (1 - 1) + 1')
# Indeed. Our (partial) assertion does _not_ detect this error:
remove_html_markup('"foo"')
# But it detects this one:
with ExpectError():
remove_html_markup('"x > y"')
# ### Assertions and Documentation
#
# In contrast to "standard" documentation – "`square_root()` expects a non-negative number `x`; its result is $\sqrt{x}$" –, assertions have a big advantage: They are _formal_ – and thus have an unambiguous semantics. Notably, we can understand what a function does _uniquely by reading its pre- and postconditions_. Here is an example:
def some_obscure_function(x: int, y: int, z: int) -> int:
result = int(...) # type: ignore
assert x == y == z or result > min(x, y, z)
assert x == y == z or result < max(x, y, z)
return result
quiz("What does this function do?",
[
"It returns the minimum value out of `x`, `y`, `z`",
"It returns the middle value out of `x`, `y`, `z`",
"It returns the maximum value out of `x`, `y`, `z`",
], 'int(0.5 ** math.cos(math.pi))', globals())
# Indeed, this would be a useful (and bug-revaling!) postcondition for one of our showcase functions in the [chapter on statistical debugging](StatisticalDebugger.ipynb).
# ### Using Assertions to Trivially Locate Defects
#
# The final benefit of assertions, and possibly even the most important in the context of this book, is _how much assertions help_ with locating defects.
# Indeed, with proper assertions, it is almost trivial to locate the one function that is responsible for a failure.
# Consider the following situation. Assume I have
# * a function `f()` whose precondition is satisfied, calling
# * a function `g()` whose precondition is violated and raises an exception.
quiz("Which function is faulty here?",
[
"`g()` because it raises an exception",
"`f()`"
" because it violates the precondition of `g()`",
"Both `f()` and `g()`"
" because they are incompatible",
"None of the above"
], 'math.factorial(int(math.tau / math.pi))', globals())
# The rule is very simple: If some function `func()` is called with its preconditions satisfied, and the postcondition of `func()` fail, then the fault in the program state must have originated at some point between these two events. Assuming that all functions called by `func()` also are correct (because their postconditions held), the defect _can only be in the code of `func()`._
# What pre- and postconditions imply is actually often called a _contract_ between caller and callee:
#
# * The caller promises to satisfy the _precondition_ of the callee,
# whereas
# * the callee promises to satisfy its own _postcondition_, delivering a correct result.
#
# In the above setting, `f()` is the caller, and `g()` is the callee; but as `f()` violates the precondition of `g()`, it has not kept its promises. Hence, `f()` violates the contract and is at fault. `f()` thus needs to be fixed.
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ## Checking Data Structures
#
# Let us get back to debugging. In debugging, assertions serve two purposes:
#
# * They immediately detect bugs (if they fail)
# * They immediately rule out _specific parts of code and state_ (if they pass)
#
# This latter part is particularly interesting, as it allows us to focus our search on the lesser checked aspects of code and state.
#
# When we say "code _and_ state", what do we mean? Actually, assertions can not quickly check several executions of a function, but also _large amounts of data_, detecting faults in data _at the moment they are introduced_.
# -
# ### Times and Time Bombs
# Let us illustrate this by an example. Let's assume we want a `Time` class that represents the time of day. Its constructor takes the current time using hours, minutes, and seconds. (Note that this is a deliberately simple example – real-world classes for representing time are way more complex.)
class Time:
def __init__(self, hours: int = 0, minutes: int = 0, seconds: int = 0) -> None:
self._hours = hours
self._minutes = minutes
self._seconds = seconds
# To access the individual elements, we introduce a few getters:
class Time(Time):
def hours(self) -> int:
return self._hours
def minutes(self) -> int:
return self._minutes
def seconds(self) -> int:
return self._seconds
# We allow to print out time, using the [ISO 8601 format](https://en.wikipedia.org/wiki/ISO_8601):
class Time(Time):
def __repr__(self) -> str:
return f"{self.hours():02}:{self.minutes():02}:{self.seconds():02}"
# Three minutes to midnight can thus be represented as
t = Time(23, 57, 0)
t
# Unfortunately, there's nothing in our `Time` class that prevents blatant misuse. We can easily set up a time with negative numbers, for instance:
t = Time(-1, 0, 0)
t
# Such a thing _may_ have some semantics (relative time, maybe?), but it's not exactly conforming to ISO format.
# Even worse, we can even _construct_ a `Time` object with strings as numbers.
t = Time("High noon") # type: ignore
# and this will raise an error only at the moment we try to print it:
with ExpectError():
print(t)
# Note how cryptic this error message must be for the poor soul who has to debug this: Where on earth do we have an `'=' alignment` in our code? The fact that the error comes from an _invalid value_ is totally lost.
# In fact, what we have here is a _time bomb_ – a fault in the program state that can sleep for ages until someone steps on it. These are hard to debug, because one has to figure out when the time bomb was set – which can be thousands or millions of lines earlier in the program. Since in the absence of type checking, _any_ assignment to a `Time` object could be the culprit – so good luck with the search.
# This is again where assertions save the day. What you need is an _assertion that checks whether the data is correct_. For instance, we could revise our constructor such that it checks for correct arguments:
class Time(Time):
def __init__(self, hours: int = 0, minutes: int = 0, seconds: int = 0) -> None:
assert 0 <= hours <= 23
assert 0 <= minutes <= 59
assert 0 <= seconds <= 60 # Includes leap seconds (ISO8601)
self._hours = hours
self._minutes = minutes
self._seconds = seconds
# These conditions check whether `hours`, `minutes`, and `seconds` are within the right range. They are called _data invariants_ (or short _invariants_) because they hold for the given data (notably, the internal attributes) at all times.
#
# Note the unusual syntax for range checks (this is a Python special), and the fact that seconds can range from 0 to 60. That's because there's not only leap years, but also leap seconds.
# With this revised constructor, we now get errors as soon as we pass an invalid parameter:
with ExpectError():
t = Time(-23, 0, 0)
# Hence, _any_ attempt to set _any_ Time object to an illegal state will be immediately detected. In other words, the time bomb defuses itself at the moment it is being set.
# This means that when we are debugging, and search for potential faults in the state that could have caused the current failure, we can now rule out `Time` as a culprit, allowing us to focus on other parts of the state.
# The more of the state we have checked with invariants,
#
# * the less state we have to examine,
# * the fewer possible causes we have to investigate,
# * the faster we are done with determining the defect.
# ### Invariant Checkers
#
# For invariants to be effective, they have to be checked at all times. If we introduce a method that changes the state, then this method will also have to ensure that the invariant is satisfied:
class Time(Time):
def set_hours(self, hours: int) -> None:
assert 0 <= hours <= 23
self._hours = hours
# This also implies that state changes should go through methods, not direct accesses to attributes. If some code changes the attributes of your object directly, without going through the method that could check for consistency, then it will be much harder for you to a) detect the source of the problem and b) even detect that a problem exists.
# #### Excursion: Checked Getters and Setters in Python
# In Python, the `@property` decorator offers a handy way to implement checkers, even for otherwise direct accesses to attributes. It allows to define specific "getter" and "setter" functions for individual properties that would even be invoked when a (seemingly) attribute is accessed.
# Using `@property`, our `Time` class could look like this:
class MyTime(Time):
@property
def hours(self) -> int:
return self._hours
@hours.setter
def hours(self, new_hours: int) -> None:
assert 0 <= new_hours <= 23
self._hours = new_hours
# To access the current hour, we no longer go through a specific "getter" function; instead, we access a synthesized attribute that – behind the scenes – invokes the "getter" function marked with `@property`:
my_time = MyTime(11, 30, 0)
my_time.hours
# If we "assign" to the attribute, the "setter" function is called in the background;
my_time.hours = 12 # type: ignore
# We see this immediately when trying to assign an illegal value:
with ExpectError():
my_time.hours = 25 # type: ignore
# If you build large infrastructures in Python, you can use these features to implement
#
# * attributes that are _checked_ every time they are accessed or changed;
# * attributes that are easier to remember than a large slew of getter and setter functions.
#
# In this book, we do not have that many attributes, and we try to use not too many Python-specific features, so we usually go without `@property`. But for Python aficionados, and especially those who care about runtime checks, checked property accesses are a boon.
# #### End of Excursion
# If we have several methods that can alter an object, it can be helpful to factor out invariant checking into its own method. Such a method can also be called to check for inconsistencies that might have been introduced without going through one of the methods – e.g. by direct object access, memory manipulation, or memory corruption.
# By convention, methods that check invariants have the name `repOK()`, since they check whether the internal representation is okay, and return True if so.
# Here's a `repOK()` method for `Time`:
class Time(Time):
def repOK(self) -> bool:
assert 0 <= self.hours() <= 23
assert 0 <= self.minutes() <= 59
assert 0 <= self.seconds() <= 60
return True
# We can integrate this method right into our constructor and our setter:
class Time(Time):
def __init__(self, hours: int = 0, minutes: int = 0, seconds: int = 0) -> None:
self._hours = hours
self._minutes = minutes
self._seconds = seconds
assert self.repOK()
def set_hours(self, hours: int) -> None:
self._hours = hours
assert self.repOK()
with ExpectError():
t = Time(-23, 0, 0)
# Having a single method that checks everything can be beneficial, as it may explicitly check for more faulty states. For instance, it is still permissible to pass a floating-point number for hours and minutes, again breaking the Time representation:
Time(1.5) # type: ignore
# (Strictly speaking, ISO 8601 _does_ allow fractional parts for seconds and even for hours and minutes – but still wants two leading digits before the fraction separator. Plus, the comma is the "preferred" fraction separator. In short, you won't be making too many friends using times formatted like the one above.)
# We can extend our `repOK()` method to check for correct types, too.
class Time(Time):
def repOK(self) -> bool:
assert isinstance(self.hours(), int)
assert isinstance(self.minutes(), int)
assert isinstance(self.seconds(), int)
assert 0 <= self.hours() <= 23
assert 0 <= self.minutes() <= 59
assert 0 <= self.seconds() <= 60
return True
Time(14, 0, 0)
# This now also catches other type errors:
with ExpectError():
t = Time("After midnight") # type: ignore
# Our `repOK()` method can also be used in combination with pre- and postconditions. Typically, you'd like to make it part of the pre- and postcondition checks.
# Assume you want to implement an `advance()` method that adds a number of seconds to the current time. The preconditions and postconditions can be easily defined:
class Time(Time):
def seconds_since_midnight(self) -> int:
return self.hours() * 3600 + self.minutes() * 60 + self.seconds()
def advance(self, seconds_offset: int) -> None:
old_seconds = self.seconds_since_midnight()
... # Advance the clock
assert (self.seconds_since_midnight() ==
(old_seconds + seconds_offset) % (24 * 60 * 60))
# But you'd really like `advance()` to check the state before _and_ after its execution – again using `repOK()`:
class BetterTime(Time):
def advance(self, seconds_offset: int) -> None:
assert self.repOK()
old_seconds = self.seconds_since_midnight()
... # Advance the clock
assert (self.seconds_since_midnight() ==
(old_seconds + seconds_offset) % (24 * 60 * 60))
assert self.repOK()
# The first postcondition ensures that `advance()` produces the desired result; the second one ensures that the internal state is still okay.
# ### Large Data Structures
#
# Invariants are especially useful if you have a large, complex data structure which is very hard to track in a conventional debugger.
# Let's assume you have a [red-black search tree](https://en.wikipedia.org/wiki/Red–black_tree) for storing and searching data. Red-black trees are among the most efficient data structures for representing associative arrays (also known as mappings); they are self-balancing and guarantee search, insertion, and deletion in logarithmic time. They also are among the most ugly to debug.
# What is a red-black tree? Here is an example from Wikipedia:
# 
# As you can see, there are red nodes and black nodes (giving the tree its name). We can define a class `RedBlackTrees` and implement all the necessary operations.
class RedBlackTree:
RED = 'red'
BLACK = 'black'
...
# +
# ignore
# A few dummy methods to make the static type checker happy. Ignore.
class RedBlackNode:
def __init__(self) -> None:
self.parent = None
self.color = RedBlackTree.BLACK
pass
# +
# ignore
# More dummy methods to make the static type checker happy. Ignore.
class RedBlackTree(RedBlackTree):
def redNodesHaveOnlyBlackChildren(self) -> bool:
return True
def equalNumberOfBlackNodesOnSubtrees(self) -> bool:
return True
def treeIsAcyclic(self) -> bool:
return True
def parentsAreConsistent(self) -> bool:
return True
def __init__(self) -> None:
self._root = RedBlackNode()
self._root.parent = None
self._root.color = self.BLACK
# -
# However, before we start coding, it would be a good idea to _first_ reason about the invariants of a red-black tree. Indeed, a red-black tree has a number of important properties that hold at all times – for instance, that the root node be black or that the tree be balanced. When we implement a red-black tree, these _invariants_ can be encoded into a `repOK()` method:
class RedBlackTree(RedBlackTree):
def repOK(self) -> bool:
assert self.rootHasNoParent()
assert self.rootIsBlack()
assert self.redNodesHaveOnlyBlackChildren()
assert self.equalNumberOfBlackNodesOnSubtrees()
assert self.treeIsAcyclic()
assert self.parentsAreConsistent()
return True
# Each of these helper methods are checkers in their own right:
class RedBlackTree(RedBlackTree):
def rootHasNoParent(self) -> bool:
return self._root.parent is None
def rootIsBlack(self) -> bool:
return self._root.color == self.BLACK
...
# With all these helpers, our `repOK()` method will become very rigorous – but all this rigor is very much needed. Just for fun, check out the [description of red-black trees on Wikipedia](https://en.wikipedia.org/wiki/Red–black_tree). The description of how insertion or deletion work is 4 to 5 pages long (each!), with dozens of special cases that all have to be handled properly. If you ever face the task of implementing such a data structure, be sure to (1) write a `repOK()` method such as the above, and (2) call it before and after each method that alters the tree:
# ignore
from typing import Any, List
class RedBlackTree(RedBlackTree):
def insert(self, item: Any) -> None:
assert self.repOK()
... # four pages of code
assert self.repOK()
def delete(self, item: Any) -> None:
assert self.repOK()
... # five pages of code
assert self.repOK()
# Such checks will make your tree run much slower – essentially, instead of logarithmic time complexity, we now have linear time complexity, as the entire tree is traversed with each change – but you will find any bugs much, much faster. Once your tree goes in production, you can deactivate `repOK()` by default, using some debugging switch to turn it on again should the need ever arise:
class RedBlackTree(RedBlackTree):
def __init__(self, checkRepOK: bool = False) -> None:
...
self.checkRepOK = checkRepOK
def repOK(self) -> bool:
if not self.checkRepOK:
return True
assert self.rootHasNoParent()
assert self.rootIsBlack()
...
return True
# Just don't delete it – future maintainers of your code will be forever grateful that you have documented your assumptions and given them a means to quickly check their code.
# ## System Invariants
#
# When interacting with the operating system, there are a number of rules that programs must follow, lest they get themselves (or the system) in some state where they cannot execute properly anymore.
#
# * If you work with files, every file that you open also must be closed; otherwise, you will deplete resources.
# * If you create temporary files, be sure to delete them after use; otherwise, you will consume disk space.
# * If you work with locks, be sure to release locks after use; otherwise, your system may end up in a deadlock.
#
# One area in which it is particularly easy to make mistakes is _memory usage_. In Python, memory is maintained by the Python interpreter, and all memory accesses are checked at runtime. Accessing a non-existing element of a string, for instance, will raise a memory error:
with ExpectError():
index = 10
"foo"[index]
# The very same expression in a C program, though, will yield _undefined behavior_ – which means that anything can happen. Let us explore a couple of C programs with undefined behavior.
# ### The C Memory Model
#
# We start with a simple C program which uses the same invalid index as our Python expression, above. What does this program do?
# ignore
open('testoverflow.c', 'w').write(r'''
#include <stdio.h>
// Access memory out of bounds
int main(int argc, char *argv[]) {
int index = 10;
return "foo"[index]; // BOOM
}
''');
print_content(open('testoverflow.c').read())
# In our example, the program will read from a random chunk or memory, which may exist or not. In most cases, nothing at all will happen – which is a bad thing, because you won't realize that your program has a defect.
# !cc -g -o testoverflow testoverflow.c
# !./testoverflow
# To see what is going on behind the scenes, let us have a look at the C memory model.
# #### Excursion: A C Memory Model Simulator
# We build a little simulation of C memory. A `Memory` item stands for a block of continuous memory, which we can access by address using `read()` and `write()`. The `__repr__()` method shows memory contents as a string.
class Memory:
def __init__(self, size: int = 10) -> None:
self.size: int = size
self.memory: List[Any] = [None for i in range(size)]
def read(self, address: int) -> Any:
return self.memory[address]
def write(self, address: int, item: Any) -> None:
self.memory[address] = item
def __repr__(self) -> str:
return repr(self.memory)
mem: Memory = Memory()
mem
mem.write(0, 'a')
mem
mem.read(0)
# We introduce `[index]` syntax for easy read and write:
class Memory(Memory):
def __getitem__(self, address: int) -> Any:
return self.read(address)
def __setitem__(self, address: int, item: Any) -> None:
self.write(address, item)
mem_with_index: Memory = Memory()
mem_with_index[1] = 'a'
mem_with_index
mem_with_index[1]
# Here are some more advanced methods to show memory cntents. The `repr()` and `_repr_markdown_()` methods display memory as a table. In a notebook, we can simply evaluate the memory to see the table.
from IPython.display import display, Markdown, HTML
class Memory(Memory):
def show_header(self) -> str:
out = "|Address|"
for address in range(self.size):
out += f"{address}|"
return out + '\n'
def show_sep(self) -> str:
out = "|:---|"
for address in range(self.size):
out += ":---|"
return out + '\n'
def show_contents(self) -> str:
out = "|Content|"
for address in range(self.size):
contents = self.memory[address]
if contents is not None:
out += f"{repr(contents)}|"
else:
out += " |"
return out + '\n'
def __repr__(self) -> str:
return self.show_header() + self.show_sep() + self.show_contents()
def _repr_markdown_(self) -> str:
return repr(self)
mem_with_table: Memory = Memory()
for i in range(mem_with_table.size):
mem_with_table[i] = 10 * i
mem_with_table
# #### End of Excursion
# In C, memory comes as a single block of bytes at continuous addresses. Let us assume we have a memory of only 20 bytes (duh!) and the string "foo" is stored at address 5:
mem_with_table: Memory = Memory(20)
mem_with_table[5] = 'f'
mem_with_table[6] = 'o'
mem_with_table[7] = 'o'
mem_with_table
# When we try to access `"foo"[10]`, we try to read the memory location at address 15 – which may exist (or not), and which may have arbitrary contents, based on whatever previous instructions left there. From there on, the behavior of our program is undefined.
# Such _buffer overflows_ can also come as _writes_ into memory locations – and thus overwrite the item that happens to be at the location of interest. If the item at address 15 happens to be, say, a flag controlling administrator access, then setting it to a non-zero value can come handy for an attacker.
# ### Dynamic Memory
#
# A second source of errors in C programs is the use of dynamic memory – that is, memory allocated and deallocated at run-time. In C, the function `malloc()` returns a continuous block of memory of a given size; the function `free()` returns it back to the system.
#
# After a block has been `free()`'d, it must no longer be used, as the memory might already be in use by some other function (or program!) again. Here's a piece of code that violates this assumption:
# ignore
open('testuseafterfree.c', 'w').write(r'''
#include <stdlib.h>
// Access a chunk of memory after it has been given back to the system
int main(int argc, char *argv[]) {
int *array = malloc(100 * sizeof(int));
free(array);
return array[10]; // BOOM
}
''');
print_content(open('testuseafterfree.c').read())
# Again, if we compile and execute this program, nothing tells us that we have just entered undefined behavior:
# !cc -g -o testuseafterfree testuseafterfree.c
# !./testuseafterfree
# What's going on behind the scenes here?
# #### Excursion: Dynamic Memory in C
# `DynamicMemory` introduces dynamic memory allocation `allocate()` and deallocation `free()`, using a list of allocated blocks.
class DynamicMemory(Memory):
# Address at which our list of blocks starts
BLOCK_LIST_START = 0
def __init__(self, *args: Any) -> None:
super().__init__(*args)
# Before each block, we reserve two items:
# One pointing to the next block (-1 = END)
self.memory[self.BLOCK_LIST_START] = -1
# One giving the length of the current block (<0: freed)
self.memory[self.BLOCK_LIST_START + 1] = 0
def allocate(self, block_size: int) -> int:
"""Allocate a block of memory"""
# traverse block list
# until we find a free block of appropriate size
chunk = self.BLOCK_LIST_START
while chunk < self.size:
next_chunk = self.memory[chunk]
chunk_length = self.memory[chunk + 1]
if chunk_length < 0 and abs(chunk_length) >= block_size:
# Reuse this free block
self.memory[chunk + 1] = abs(chunk_length)
return chunk + 2
if next_chunk < 0:
# End of list - allocate new block
next_chunk = chunk + block_size + 2
if next_chunk >= self.size:
break
self.memory[chunk] = next_chunk
self.memory[chunk + 1] = block_size
self.memory[next_chunk] = -1
self.memory[next_chunk + 1] = 0
base = chunk + 2
return base
# Go to next block
chunk = next_chunk
raise MemoryError("Out of Memory")
def free(self, base: int) -> None:
"""Free a block of memory"""
# Mark block as available
chunk = base - 2
self.memory[chunk + 1] = -abs(self.memory[chunk + 1])
# In our table, we highlight free blocks in grey:
class DynamicMemory(DynamicMemory):
def show_header(self) -> str:
out = "|Address|"
color = "black"
chunk = self.BLOCK_LIST_START
allocated = False
# States and colors
for address in range(self.size):
if address == chunk:
color = "blue"
next_chunk = self.memory[address]
elif address == chunk + 1:
color = "blue"
allocated = self.memory[address] > 0
chunk = next_chunk
elif allocated:
color = "black"
else:
color = "lightgrey"
item = f'<span style="color: {color}">{address}</span>'
out += f"{item}|"
return out + '\n'
dynamic_mem: DynamicMemory = DynamicMemory(10)
dynamic_mem
dynamic_mem.allocate(2)
dynamic_mem
dynamic_mem.allocate(2)
dynamic_mem
dynamic_mem.free(2)
dynamic_mem
dynamic_mem.allocate(1)
dynamic_mem
with ExpectError():
dynamic_mem.allocate(1)
# #### End of Excursion
# Dynamic memory is allocated as part of our main memory. The following table shows unallocated memory in grey and allocated memory in black:
dynamic_mem: DynamicMemory = DynamicMemory(13)
dynamic_mem
# The numbers already stored (-1 and 0) are part of our dynamic memory housekeeping (highlighted in blue); they stand for the next block of memory and the length of the current block, respectively.
# Let us allocate a block of 3 bytes. Our (simulated) allocation mechanism places these at the first continuous block available:
p1 = dynamic_mem.allocate(3)
p1
# We see that a block of 3 items is allocated at address 2. The two numbers before that address (5 and 3) indicate the beginning of the next block as well as the length of the current one.
dynamic_mem
# Let us allocate some more.
p2 = dynamic_mem.allocate(4)
p2
# We can make use of that memory:
dynamic_mem[p1] = 123
dynamic_mem[p2] = 'x'
dynamic_mem
# When we free memory, the block is marked as free by giving it a negative length:
dynamic_mem.free(p1)
dynamic_mem
# Note that freeing memory does not clear memory; the item at `p1` is still there. And we can also still access it.
dynamic_mem[p1]
# But if, in the meantime, some other part of the program requests more memory and uses it...
p3 = dynamic_mem.allocate(2)
dynamic_mem[p3] = 'y'
dynamic_mem
# ... then the memory at `p1` may simply be overwritten.
dynamic_mem[p1]
# An even worse effect comes into play if one accidentally overwrites the dynamic memory allocation information; this can easily corrupt the entire memory management. In our case, such corrupted memory can lead to an endless loop when trying to allocate more memory:
from ExpectError import ExpectTimeout
dynamic_mem[p3 + 3] = 0
dynamic_mem
# When `allocate()` traverses the list of blocks, it will enter an endless loop between the block starting at address 0 (pointing to the next block at 5) and the block at address 5 (pointing back to 0).
with ExpectTimeout(1):
dynamic_mem.allocate(1)
# Real-world `malloc()` and `free()` implementations suffer from similar problems. As stated above: As soon as undefined behavior is reached, anything may happen.
# ### Managed Memory
#
# The solution to all these problems is to _keep track of memory_, specifically
#
# * which parts of memory have been _allocated_, and
# * which parts of memory have been _initialized_.
# To this end, we introduce two extra flags for each address:
#
# * The `allocated` flag tells whether an address has been allocated; the `allocate()` method sets them, and `free()` clears them again.
# * The `initialized` flag tells whether an address has been written to. This is cleared as part of `allocate()`.
# With these, we can run a number of checks:
#
# * When _writing_ into memory and _freeing_ memory, we can check whether the address has been _allocated_; and
# * When _reading_ from memory, we can check whether the address has been allocated and _initialized_.
#
# Both of these should effectively prevent memory errors.
# #### Excursion: Managed Memory
# We create a simulation of managed memory. `ManagedMemory` keeps track of every address whether it is intiialized and allocated.
class ManagedMemory(DynamicMemory):
def __init__(self, *args: Any) -> None:
super().__init__(*args)
self.initialized = [False for i in range(self.size)]
self.allocated = [False for i in range(self.size)]
# This allows memory access functions to run a number of extra checks:
class ManagedMemory(ManagedMemory):
def write(self, address: int, item: Any) -> None:
assert self.allocated[address], \
"Writing into unallocated memory"
self.memory[address] = item
self.initialized[address] = True
def read(self, address: int) -> Any:
assert self.allocated[address], \
"Reading from unallocated memory"
assert self.initialized[address], \
"Reading from uninitialized memory"
return self.memory[address]
# Dynamic memory functions are set up such that they keep track of these flags.
class ManagedMemory(ManagedMemory):
def allocate(self, block_size: int) -> int:
base = super().allocate(block_size)
for i in range(block_size):
self.allocated[base + i] = True
self.initialized[base + i] = False
return base
def free(self, base: int) -> None:
assert self.allocated[base], \
"Freeing memory that is already freed"
block_size = self.memory[base - 1]
for i in range(block_size):
self.allocated[base + i] = False
self.initialized[base + i] = False
super().free(base)
# Let us highlight these flags when printing out the table:
class ManagedMemory(ManagedMemory):
def show_contents(self) -> str:
return (self.show_allocated() +
self.show_initialized() +
DynamicMemory.show_contents(self))
def show_allocated(self) -> str:
out = "|Allocated|"
for address in range(self.size):
if self.allocated[address]:
out += "Y|"
else:
out += " |"
return out + '\n'
def show_initialized(self) -> str:
out = "|Initialized|"
for address in range(self.size):
if self.initialized[address]:
out += "Y|"
else:
out += " |"
return out + '\n'
# #### End of Excursion
# Here comes a simple simulation of managed memory. After we create memory, all addresses are neither allocated nor initialized:
managed_mem: ManagedMemory = ManagedMemory()
managed_mem
# Let us allocate some elements. We see that the first three bytes are now marked as allocated:
p = managed_mem.allocate(3)
managed_mem
# After writing into memory, the respective addresses are marked as "initialized":
managed_mem[p] = 10
managed_mem[p + 1] = 20
managed_mem
# Attempting to read uninitialized memory fails:
with ExpectError():
x = managed_mem[p + 2]
# When we free the block again, it is marked as not allocated:
managed_mem.free(p)
managed_mem
# And accessing any element of the free'd block will yield an error:
with ExpectError():
managed_mem[p] = 10
# Freeing the same block twice also yields an error:
with ExpectError():
managed_mem.free(p)
# With this, we now have a mechanism in place to fully detect memory issues in languages such as C.
# Obviously, keeping track of whether memory is allocated/initialized or not requires some extra memory – and also some extra computation time, as read and write accesses have to be checked first. During testing, however, such effort may quickly pay off, as memory bugs can be quickly discovered.
# To detect memory errors, a number of tools have been developed. The first class of tools _interprets_ the instructions of the executable code, tracking all memory accesses. For each memory access, they can check whether the memory accessed _exists_ and has been _initialized_ at some point.
# ### Checking Memory Usage with Valgrind
#
# The [Valgrind](https://www.valgrind.org) tool allows to _interpret_ executable code, thus tracking each and every memory access. You can use Valgrind to execute any program from the command-line, and it will check all memory accesses during execution.
#
# <!-- Installing ValGrind on macOS: https://github.com/LouisBrunner/valgrind-macos/ -->
#
# Here's what happens if we run Valgrind on our `testuseafterfree` program:
print_content(open('testuseafterfree.c').read())
# !valgrind ./testuseafterfree
# We see that Valgrind has detected the issue ("Invalid read of size 4") during execution; it also reported the current stack trace (and hence the location at which the error occurred). Note that the program continues execution even after the error occurred; should further errors occur, Valgrind will report these, too.
# Being an interpreter, Valgrind slows down execution of programs dramatically. However, it requires no recompilation and thus can work on code (and libraries) whose source code is not available.
# Valgrind is not perfect, though. For our `testoverflow` program, it fails to detect the illegal access:
print_content(open('testoverflow.c').read())
# !valgrind ./testoverflow
# This is because at compile time, the information about the length of the `"foo"` string is no longer available – all Valgrind sees is a read access into the static data portion of the executable that may be valid or invalid. To actually detect such errors, we need to hook into the compiler.
# ### Checking Memory Usage with Memory Sanitizer
#
# The second class of tools to detect memory issues are _address sanitizers_. An address sanitizer injects memory-checking code into the program _during compilation_. This means that every access will be checked – but this time, the code still runs on the processor itself, meaning that the speed is much less reduced.
# Here is an example of how to use the address sanitizer of the Clang C compiler:
# !cc -fsanitize=address -o testuseafterfree testuseafterfree.c
# At the very first moment we have an out-of-bounds access, the program aborts with a diagnostic message – in our case already during `read_overflow()`.
# !./testuseafterfree
# Likewise, if we apply the address sanitizer on `testoverflow`, we also immediately get an error:
# !cc -fsanitize=address -o testoverflow testoverflow.c
# !./testoverflow
# Since the address sanitizer monitors each and every read and write, as well as usage of `free()`, it will require some effort to create a bug that it won't catch. Also, while Valgrind runs the program ten times slower and more, the performance penalty for memory sanitization is much much lower. Sanitizers can also help in finding data races, memory leaks, and all other sorts of undefined behavior. As [<NAME> puts it](https://lemire.me/blog/2016/04/20/no-more-leaks-with-sanitize-flags-in-gcc-and-clang/):
#
# > Really, if you are using gcc or clang and you are not using these flags, you are not being serious.
# ## When Should Invariants be Checked?
#
# We have seen that during testing and debugging, invariants should be checked _as much as possible_, thus narrowing down the time it takes to detect a violation to a minimum. The easiest way to get there is to have them checked as _postcondition_ in the constructor and any other method that sets the state of an object.
# If you have means to alter the state of an object outside of these methods – for instance, by directly writing to memory, or by writing to internal attributes –, then you may have to check them even more frequently. Using the [tracing infrastructure](Tracer.ipynb), for instance, you can have the tracer invoke `repOK()` with each and every line executed, thereby again directly pinpointing the moment the state gets corrupted. While this will slow down execution tremendously, it is still better to have the computer do the work than you stepping backwards and forwards through an execution.
# Another question is whether assertions should remain active even in production code. Assertions take time, and this may be too much for production.
# ### Assertions are not Production Code
#
# First of all, assertions are _not_ production code – the rest of the code should not be impacted by any assertion being on or off. If you write code like
#
# ```python
# assert map.remove(location)
# ```
# your assertion will have a side effect, namely removing a location from the map. If one turns assertions off, the side effect will be turned off as well. You need to change this into
#
# ```python
# locationRemoved = map.remove(location)
# assert locationRemoved
# ```
# ### For System Preconditions, Use Production Code
#
# Consequently, you should not rely on assertions for _system preconditions_ – that is, conditions that are necessary to keep the system running. System input (or anything that could be controlled by another party) still has to be validated by production code, not assertions. Critical conditions have to be checked by production code, not (only) assertions.
#
# If you have code such as
# ```python
# assert command in {"open", "close", "exit"}
# exec(command)
# ```
# then having the assertion document and check your assumptions is fine. However, if you turn the assertion off in production code, it will only be a matter of time until somebody sets `command` to `'system("/bin/sh")'` and all of a sudden takes control over your system.
# ### Consider Leaving Some Assertions On
#
# The main reason for turning assertions off is efficiency. However, _failing early is better than having bad data and not failing._ Think carefully which assertions have a high impact on execution time, and turn these off first. Assertions that have little to no impact on resources can be left on.
# As an example, here's a piece of code that handles traffic in a simulation. The `light` variable can be either `RED`, `AMBER`, or `GREEN`:
#
# ```python
# if light == RED:
# traffic.stop()
# elif light == AMBER:
# traffic.prepare_to_stop()
# elif light == GREEN:
# traffic.go()
# else:
# pass # This can't happen!
# ```
# Having an assertion
#
# ```python
# assert light in [RED, AMBER, GREEN]
# ```
#
# in your code will eat some (minor) resources. However, adding a line
#
# ```python
# assert False
# ```
#
# in the place of the `This can't happen!` line, above, will still catch errors, but require no resources at all.
# If you have very critical software, it may be wise to actually pay the extra penalty for assertions (notably system assertions) rather than sacrifice reliability for performance. Keeping a memory sanitizer on even in production can have a small impact on performance, but will catch plenty of errors before some corrupted data (and even some attacks) have bad effects downstream.
# ### Define How Your Application Should Handle Internal Errors
#
# By default, failing assertions are not exactly user-friendly – the diagnosis they provide is of interest to the code maintainers only. Think of how your application should handle internal errors as discovered by assertions (or the runtime system). Simply exiting (as assertions on C do) may not be the best option for critical software. Think about implementing your own assert functions with appropriate recovery methods.
# ## Synopsis
# This chapter discusses _assertions_ to define _assumptions_ on function inputs and results:
def my_square_root(x): # type: ignore
assert x >= 0
y = square_root(x)
assert math.isclose(y * y, x)
return y
# Notably, assertions detect _violations_ of these assumptions at runtime:
with ExpectError():
y = my_square_root(-1)
# _System assertions_ help to detect invalid memory operations.
managed_mem = ManagedMemory()
managed_mem
with ExpectError():
x = managed_mem[2]
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# ## Lessons Learned
#
# * _Assertions_ are powerful tools to have the computer check invariants during execution:
# * _Preconditions_ check whether the arguments to a function are correct
# * _Postconditions_ check whether the result of a function is correct
# * _Data Invariants_ allow to check data structures for integrity
# * Since assertions can be turned off for optimization, they should
# * not _change correct operation_ in any way
# * not do _any work that your application requires for correct operation_
# * not be used as a _replacement for errors that can possibly happen_; create permanent checks (and own exceptions) for these
# * _System assertions_ are powerful tools to monitor the integrity of the runtime system (notably memory)
# * The more assertions,
# * the earlier errors are detected
# * the easier it is to locate defects
# * the better the guidance towards failure causes during debugging
# -
# ignore
import os
import shutil
for path in [
'assert.h',
'testassert',
'testassert.c',
'testassert.dSYM',
'testoverflow',
'testoverflow.c',
'testoverflow.dSYM',
'testuseafterfree',
'testuseafterfree.c',
'testuseafterfree.dSYM',
]:
if os.path.isdir(path):
shutil.rmtree(path)
else:
try:
os.remove(path)
except FileNotFoundError:
pass
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ## Next Steps
#
# In the next chapters, we will learn how to
#
# * [identify performance issues](PerformanceDebugger.ipynb)
# * [check flows and dependencies](Slicer.ipynb)
# -
# ## Background
#
# The usage of _assertions_ goes back to the earliest days of programming. In 1947, Neumann and Goldstine defined _assertion boxes_ that would check the limits of specific variables. In his 1949 talk ["Checking a Large Routine"](http://www.turingarchive.org/browse.php/B/8), Alan Turing suggested
#
# > How can one check a large routine in the sense of making sure that it's right? In order that the man who checks may not have too difficult a task, the programmer should make a number of definite _assertions_ which can be checked individually, and from which the correctness of the whole program easily follows.
#
# [Valgrind](https://valgrind.org) originated as an academic tool which has seen lots of industrial usage. A [list of papers](https://www.valgrind.org/docs/pubs.html) is available on the Valgrind page.
#
# The [Address Sanitizer](http://clang.llvm.org/docs/AddressSanitizer.html) discussed in this chapter was developed at Google; the [paper by Serebryany](https://www.usenix.org/system/files/conference/atc12/atc12-final39.pdf) discusses several details.
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# ## Exercises
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# ### Exercise 1 – Storage Assertions
# -
# The Python [`shelve`](https://docs.python.org/3/library/shelve.html) module provides a simple interface for permanent storage of Python objects:
import shelve
d = shelve.open('mydb')
d['123'] = 123
d['123']
d.close()
d = shelve.open('mydb')
d['123']
d.close()
# Based on `shelve`, we can implement a class `ObjectStorage` that uses a context manager (a `with` block) to ensure the shelve database is always closed - also in presence of exceptions:
# ignore
from typing import Sequence, Any, Callable, Optional, Type, Tuple, Any
from typing import Dict, Union, Set, List, FrozenSet, cast
from types import TracebackType
class Storage:
def __init__(self, dbname: str) -> None:
self.dbname = dbname
def __enter__(self) -> Any:
self.db = shelve.open(self.dbname)
return self
def __exit__(self, exc_tp: Type, exc_value: BaseException,
exc_traceback: TracebackType) -> Optional[bool]:
self.db.close()
return None
def __getitem__(self, key: str) -> Any:
return self.db[key]
def __setitem__(self, key: str, value: Any) -> None:
self.db[key] = value
with Storage('mydb') as storage:
print(storage['123'])
# #### Task 1 – Local Consistency
#
# Extend `Storage` with assertions that ensure that after adding an element, it also can be retrieved with the same value.
# **Solution.** One single assertion suffices:
class Storage(Storage):
def __setitem__(self, key: str, value: Any) -> None:
self.db[key] = value
assert self.db[key] == value
# #### Task 2 – Global Consistency
#
# Extend `Storage` with a "shadow dictionary" which holds elements in memory storage, too. Have a `repOK()` method that memory storage and `shelve` storage are identical at all times.
# **Solution.** Here is a possible implementation:
class ShadowStorage:
def __init__(self, dbname: str) -> None:
self.dbname = dbname
def __enter__(self) -> Any:
self.db = shelve.open(self.dbname)
self.memdb = {}
for key in self.db.keys():
self.memdb[key] = self.db[key]
assert self.repOK()
return self
def __exit__(self, exc_tp: Type, exc_value: BaseException,
exc_traceback: TracebackType) -> Optional[bool]:
self.db.close()
return None
def __getitem__(self, key: str) -> Any:
assert self.repOK()
return self.db[key]
def __setitem__(self, key: str, value: Any) -> None:
assert self.repOK()
self.memdb[key] = self.db[key] = value
assert self.repOK()
def repOK(self) -> bool:
assert self.db.keys() == self.memdb.keys(), f"{self.dbname}: Differing keys"
for key in self.memdb.keys():
assert self.db[key] == self.memdb[key], \
f"{self.dbname}: Differing values for {repr(key)}"
return True
with ShadowStorage('mydb') as storage:
storage['456'] = 456
print(storage['123'])
# ignore
try:
os.remove('mydb.db') # on macOS
except FileNotFoundError:
pass
# ignore
try:
os.remove('mydb') # on Linux
except FileNotFoundError:
pass
| notebooks/Assertions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
# !ls data/alex_hours_csv/
df = pd.read_csv('data/alex_hours_csv/17_5_2021.csv')
df["Time"] = '2021-05-17 ' + df['Time'].astype(str)
df["Time"] = pd.to_datetime(df["Time"])
df
df.plot(x="Time", y="Username")
| pandas_practice/.ipynb_checkpoints/sample_work_hours_practice-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# [View in Colaboratory](https://colab.research.google.com/github/ayan59dutta/Assignment-4/blob/ayan59dutta/ayan59dutta.ipynb)
# + [markdown] id="2XXfXed5YLbe" colab_type="text"
# # First Date with TensorFlow
#
# Hi all,<br/>
#
# You know what's important for understanding Deep Learning / Machine Learning?<br/>
# Intuition. Period.
#
# And Intuition comes when you run the code multiple times.
#
# So, today I can write a couple of defination and say this is this, this is that.<br/>
# You Google half of the things up. You find answers which you need to Google further.<br/>
# In the process, you probably won't even remember what's the first thing you started out with!
#
# # So?
#
# Hence on, I will execute cells with code. <br/>
# The neurons in your brain will optimize a function to get a hold of what each function is doing.<br/>
# **No Theory Just Code.**
#
# I will at max give a defination that extends for a line. That's it.<br/>
# Let's get started!
#
# <hr/>
#
# **RECOMMENDED!**<br/>
# Write the code in the cells using the signals sent by your brain to your fingers!<br/>
# Don't just `shift+enter` the cells.
#
# [Source](https://github.com/iArunava/TensorFlow-NoteBooks)
# + id="gYWUpE-bYKWP" colab_type="code" colab={}
# Essential imports
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
# + id="eKpz5NCIYMdi" colab_type="code" colab={}
# Let's define some tensors
t1 = tf.constant(2.0, dtype=tf.float32)
t2 = tf.constant([1.0, 2.0], dtype=tf.float32)
t3 = tf.constant([[[1.0, 9.0], [2.0, 3.0], [4.0, 5.0]],
[[1.0, 9.0], [2.0, 3.0], [4.0, 5.0]]])
# + id="vmMcjzTxbWzw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="97e61a64-612a-4e7b-89b4-755ba204b606"
# Let's print them out!
print (t1)
print (t2)
print (t3)
# + [markdown] id="10ahnfjYbcop" colab_type="text"
# Where's Waldo?<br/>
# I mean, the value?<br/>
#
# So, the thing is you can't print the value of tensors directly.<br/>
# You have to use `session`, so let's do that!
# + id="ol6O5I7Tb2nb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 219} outputId="3fd33587-4d6f-4c4d-88a0-060842eccf58"
sess = tf.Session()
print (sess.run(t1))
print ("=======================")
print (sess.run(t2))
print ("=======================")
print (sess.run(t3))
sess.close()
# + [markdown] id="rXKfVs_zb-kU" colab_type="text"
# Aaahaa!! Just printed those tensors!!!<br/>
# Feels good! <br/>
#
# For some of you, who are like, dude you got "No Theory Just Code" in bold <br/>
# And you are still using the markdown cells for the theory ?!
#
# I am just gonna say I am a unreasonable man.<br/>
#
#
# So, you are programming with tf.<br/>
# What ever you do is broken down to 2 basic steps:
# - Building the computational Graph!
# - Execute that graph using `session`!
#
# That's all!
#
# <hr/>
#
# Let's compare this 2 steps with what we did above!<br/>
# So, I defined 3 `tensor`s and these 3 `tensor`s formed my computational Graph.<br/>
# And then I executed each tensor in this graph using a `session`.
#
# That simple!
#
# <hr/>
#
# Now, let's define a few more computational graphs and execute them with sessions.
#
# Okay, to start with let's build this computational graph!
#
# 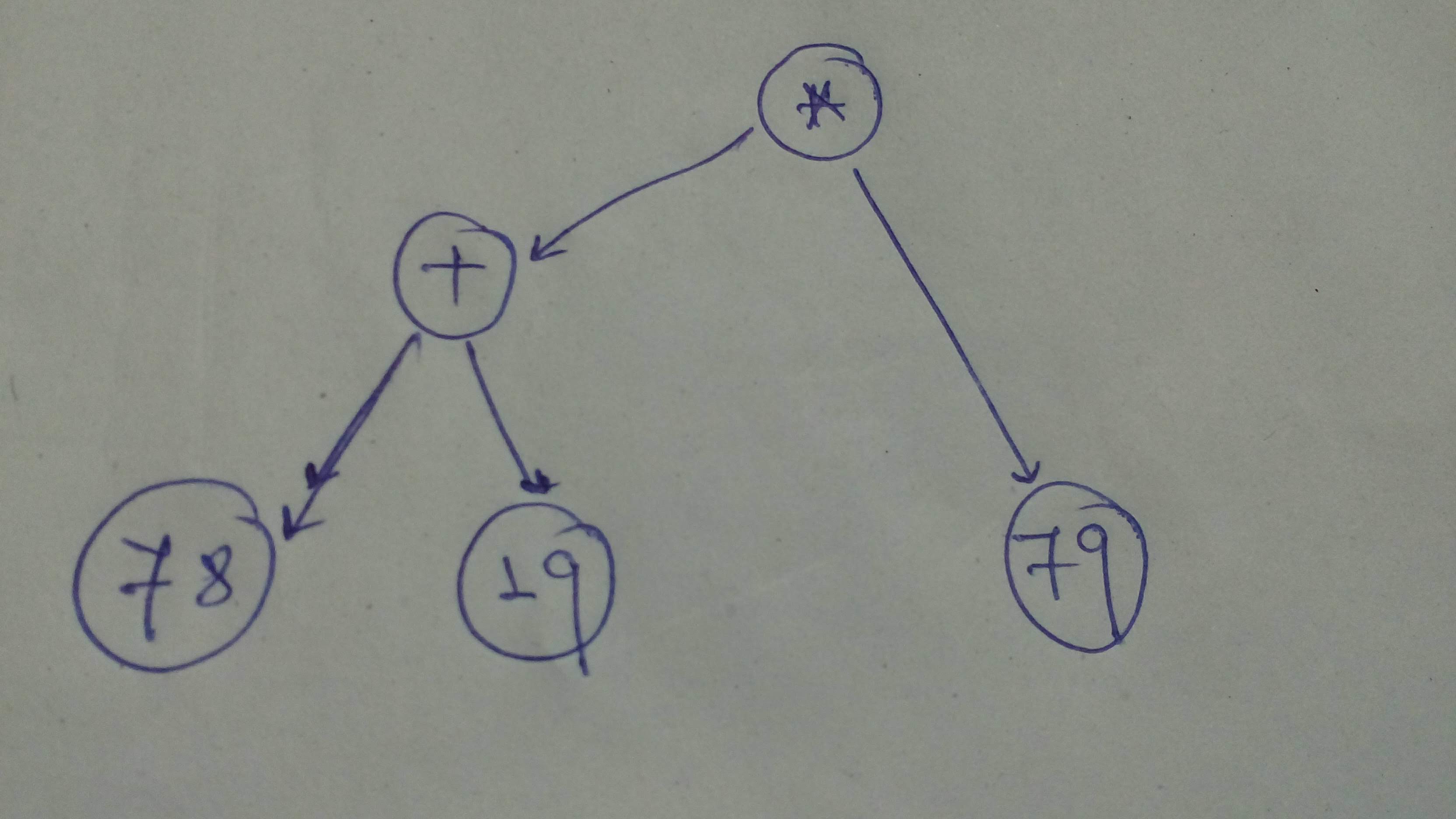
# + id="FyVz0GNqgreZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="df83c837-8f55-487f-b2d9-92466a952d6a"
# Let's define the graph
comp_graph_1 = tf.multiply(tf.add(78, 19), 79)
# Alternatively
comp_graph_1_alt = (tf.constant(78) + tf.constant(19)) * tf.constant(79)
# Let's execute using session
sess = tf.Session()
print ('Comp Graph 1 : ', sess.run(comp_graph_1))
print ('Comp Graph 1 Alt: ', sess.run(comp_graph_1_alt))
sess.close()
# + [markdown] id="SVMMtuFYhaQB" colab_type="text"
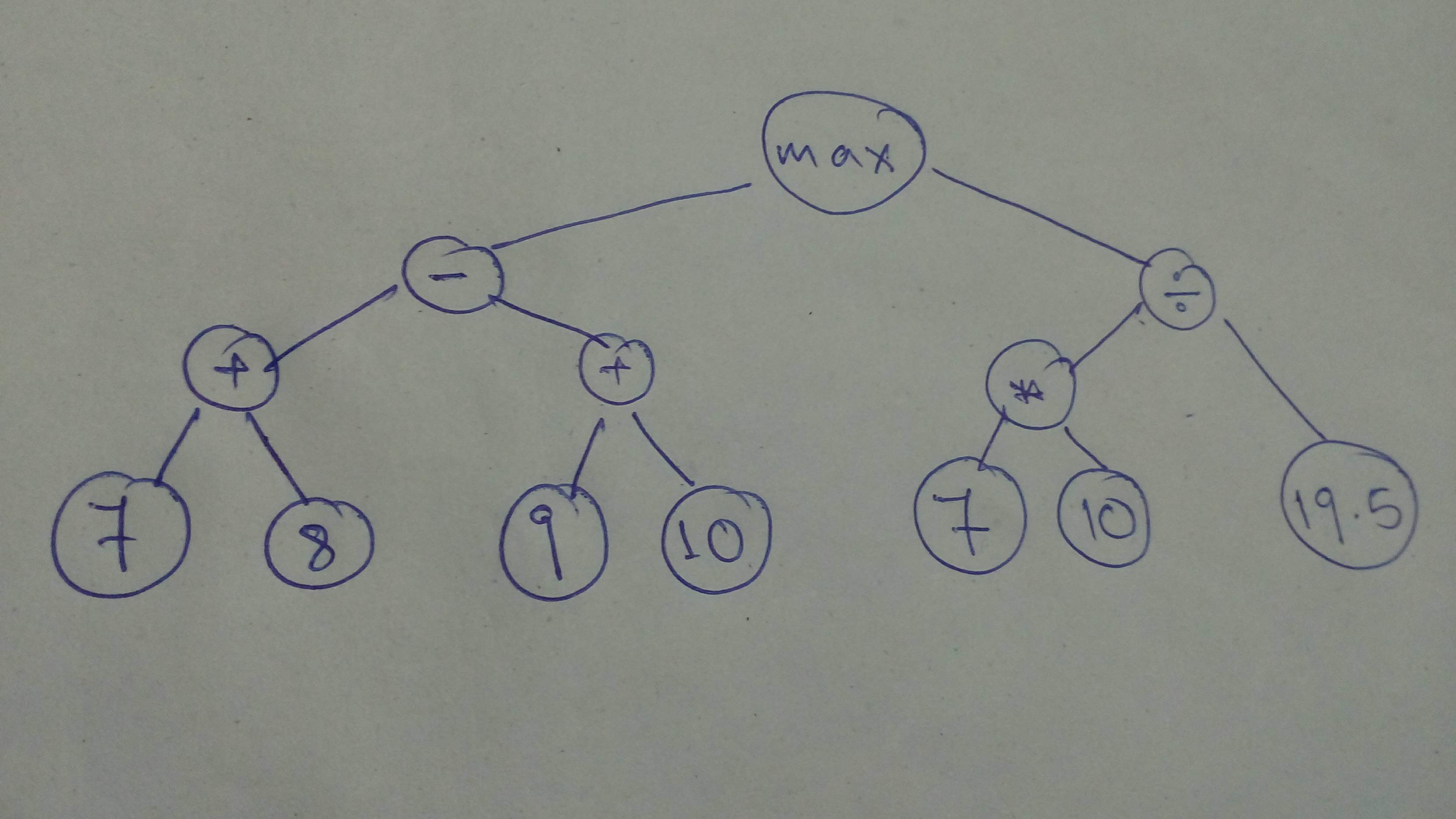
# Let's define a sligtly more involved graph!
#
# 
# + id="4856BTvRhiBb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="0dce8f47-b8f5-4b23-eb4c-313ac98034b6"
# Let build the graph
# We need to cast cause the tensors operated on should be of the same type
comp_graph_part_1 = tf.cast(tf.subtract(tf.add(7, 8), tf.add(9, 10)),
dtype=tf.float32)
comp_graph_part_2 = tf.divide(tf.cast(tf.multiply(7, 10), dtype=tf.float32), tf.constant(19.5))
comp_graph_complete = tf.maximum(comp_graph_part_1, comp_graph_part_2)
# Let's execute
sess = tf.Session()
part1_res, part2_res, total_res = sess.run([comp_graph_part_1, comp_graph_part_2, comp_graph_complete])
print ('Complete Result: ', total_res)
print ('Part 1 Result: ', part1_res)
print ('Part 2 Result: ', part2_res)
sess.close()
# + [markdown] id="B-_ZDtEbj4N0" colab_type="text"
# Cool! Let's go! Build another graph and execute it with sessions.<br/>
#
# But this time, it's all you!
#
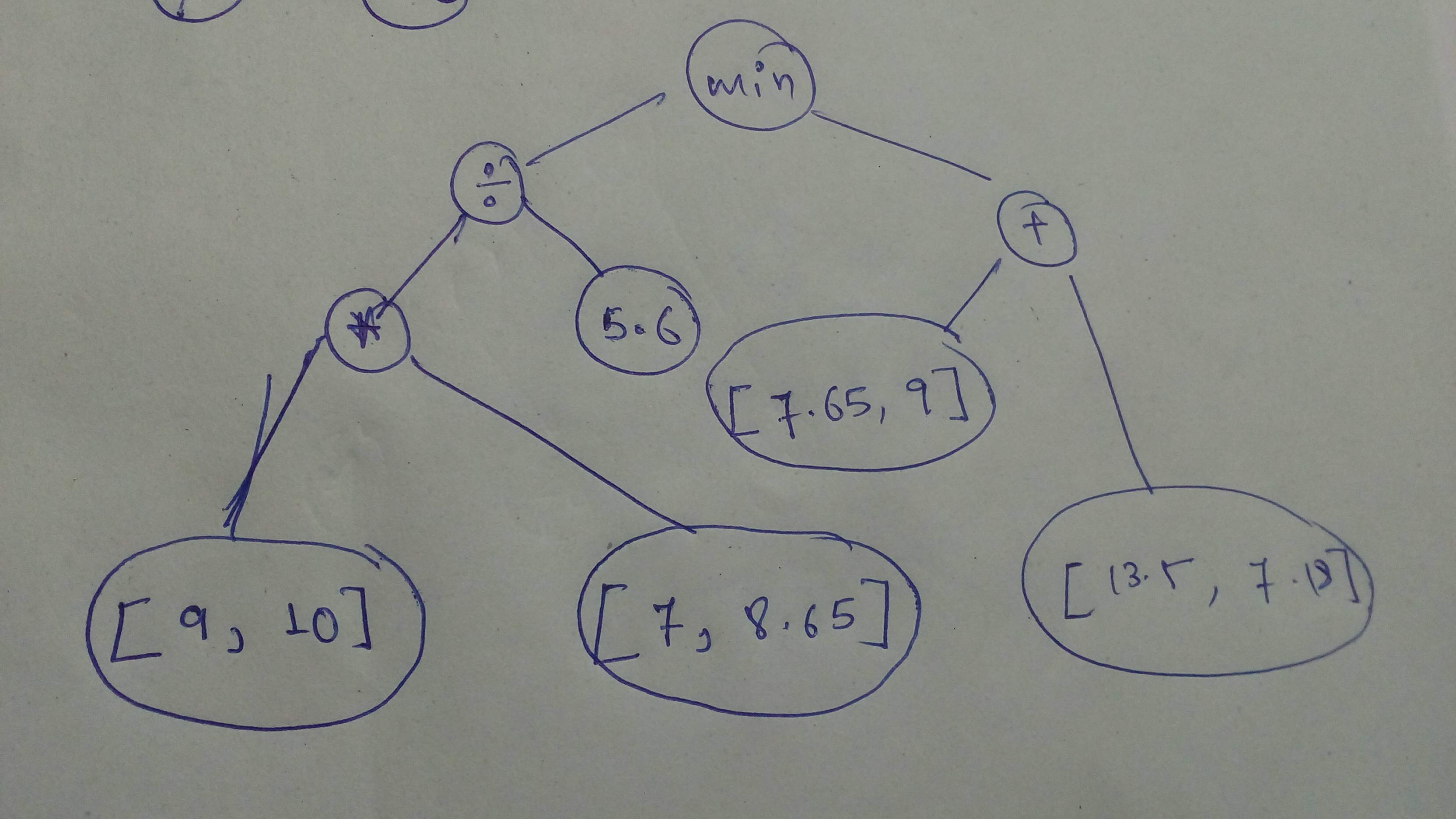
# Build this graph and execute it with `session`!
#
# 
#
# _Remember that `tensors` operated on should be of the same type!_<br/>
# _Search up errors and other help you need on Google_
# + id="-uHNe1BolJY0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="9faf40fc-22ec-4665-ffe1-0c060caf047f"
# Build the graph
# YOUR CODE HERE
n1 = tf.constant([9, 10], dtype=tf.float32)
n2 = tf.constant([7, 8.65], dtype=tf.float32)
n3 = tf.constant(5.6, dtype=tf.float32)
n4 = tf.constant([7.65, 9], dtype=tf.float32)
n5 = tf.constant([13.5, 7.18], dtype=tf.float32)
cg1 = tf.minimum(((n1 * n2) / n3), (n4 + n5))
# Execute
# YOUR CODE HERE
with tf.Session() as sess:
print(sess.run(cg1))
# + [markdown] id="qmap38WelREN" colab_type="text"
# Let's do another!<br/>
# It's fun! Isn't it?!
#
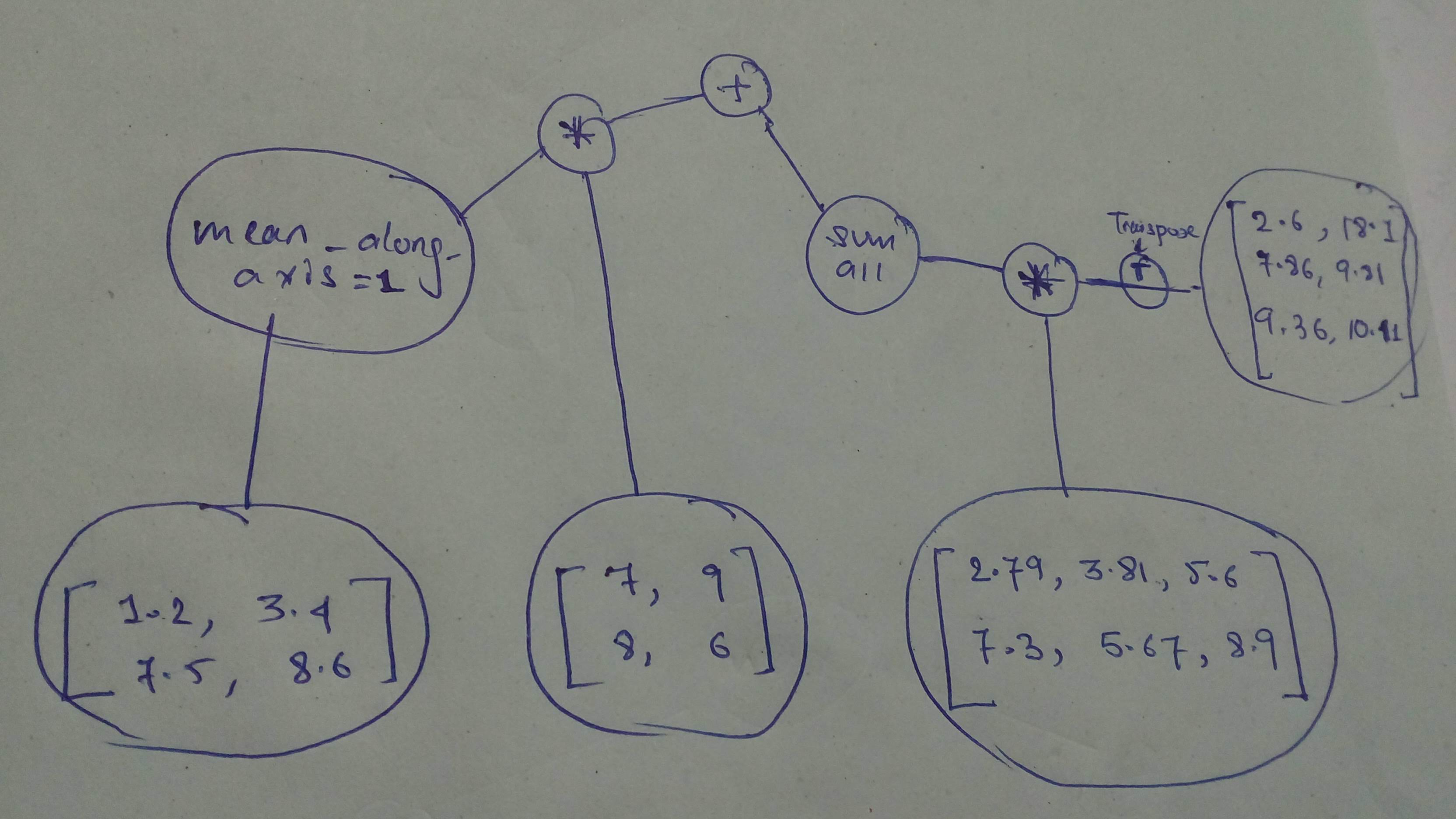
# Build and execute this one!
#
# 
# + id="0ZhYwAlLmEvB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="e2798e59-847e-400c-a1a7-ea8019f09419"
# Build the graph
# YOUR CODE HERE
n1 = tf.constant([[1.2, 3.4],
[7.5, 8.6]], dtype=tf.float32)
n2 = tf.constant([[7, 9],
[8, 6]], dtype=tf.float32)
n3 = tf.constant([[2.79, 3.81, 5.6],
[7.3, 5.67, 8.9]], dtype=tf.float32)
n4 = tf.constant([[2.6, 18.1],
[7.86, 9.81],
[9.36, 10.41]], dtype=tf.float32)
cg2 = (tf.reduce_mean(n1, axis=1) * n2) + tf.reduce_sum(n3 * tf.transpose(n4))
# Execute
# YOUR CODE HERE
with tf.Session() as sess:
print(sess.run(cg2))
# + [markdown] id="BnB0b6qCmGmg" colab_type="text"
# And a final one, before we move on to the next part!
#
# 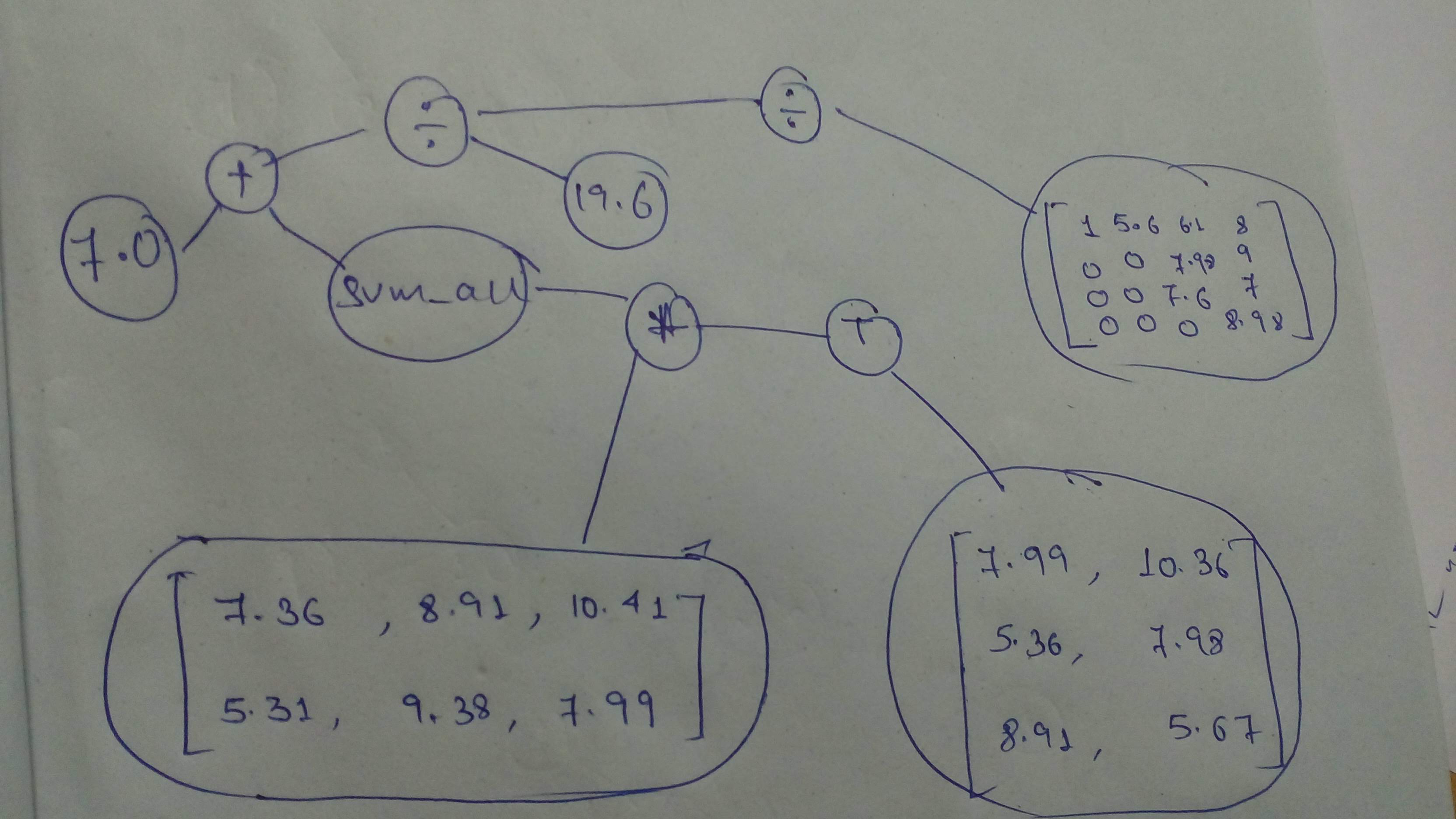
# + id="GQWyCvsQmMcL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 90} outputId="4b7e04ba-a97b-4f0d-bc60-aa7dd1adc5d0"
# Build the graph
# YOUR CODE HERE
n1 = tf.constant(7.0, dtype=tf.float32)
n2 = tf.constant([[7.36, 8.91, 10.41],
[5.31, 9.38, 7.99]], dtype=tf.float32)
n3 = tf.constant([[7.99, 10.36],
[5.36, 7.98],
[8.91, 5.67]], dtype=tf.float32)
n4 = tf.constant(19.6, dtype=tf.float32)
n5 = tf.constant([[1, 5.6, 6.1, 8],
[0, 0, 7.98, 9],
[0, 0, 7.6, 7],
[0, 0, 0, 8.98]], dtype=tf.float32)
cg3 = ((n1 + tf.reduce_sum(n2 * tf.transpose(n3))) / n4) / n5
# Execute
# YOUR CODE HERE
with tf.Session() as sess:
print(sess.run(cg3))
# + [markdown] id="12NC7XTPsJw7" colab_type="text"
# # Linear Regression
#
# Okay, now we will create a dummy dataset and perform linear regression on this dataset!
#
#
# To get you in the habit of looking up for the documentation, I am not providing what some of the following functions does, Google them up!
# + id="RTA72Bmol5tG" colab_type="code" colab={}
# Essential imports
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
# + id="hW31RZkjtNwI" colab_type="code" colab={}
# Create the dataset
X = np.linspace(-30.0, 300.0, 300)
Y = 2 * np.linspace(-30.0, 250.0, 300) + np.random.randn(*X.shape)
# Divide it into train and test
train_X = X[:250]
train_Y = Y[:250]
test_X = X[250:]
test_Y = Y[250:]
# + id="LQKy6U33y4lt" colab_type="code" colab={}
# Let's define the hyperparameters
learning_rate = 0.000005
n_epochs = 1000
interval = 50
# + id="1h1-D8K1uT48" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 347} outputId="0c6090c8-c4df-48ca-8702-f59966201ce6"
# let's viz the first 10 datapoints of the dataset
plt.plot(train_X[:10], train_Y[:10], 'g')
plt.show()
# + id="P3-iuxE4sjAf" colab_type="code" colab={}
# Let's define the placeholders
# # Placeholders?
# The input to the model changes on iteration
# So we cannot have a constant in the input as we did before
# And thus we need placeholders which we can change on each
# iteration of the training
x = tf.placeholder(tf.float32, name='x')
y = tf.placeholder(tf.float32, name='y')
# + id="8hPRkaoxvRyV" colab_type="code" colab={}
# Let's define the linear regression model
# # tf.Variable?
# We define the model parameters as tf.Variables
# as they get updated throghout the training.
# And variables denotes something which changes overtime.
W = tf.Variable(0.0, name='weight_1')
b = tf.Variable(0.0, name='bias_1')
pred_y = (W*x) + b
# + id="cSw1P8bkv96r" colab_type="code" colab={}
# Let's define the loss function
# We are going to use the mean squared loss
loss = tf.reduce_mean(tf.square(y - pred_y))
# + id="5G4uQqjsygNj" colab_type="code" colab={}
# Let's define the optimizer
# And specify the which value (i.e. loss) it has to minimize
optimizer = tf.train.GradientDescentOptimizer(learning_rate=learning_rate).minimize(loss)
# + id="ttI7ZT-ozAm1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 751} outputId="c7b50021-9628-4f9a-bdbc-6524b38bc362"
# So the graph is now built
# Now let's execute the graph using session
# i.e. lets train the model
# What it is to train a model?
# To update the paramters in the graph (i.e. tf.Variables)
# So that the loss is minimized
# Okay let's start!
with tf.Session() as sess:
# We need to initialize the variables in our graph
sess.run(tf.global_variables_initializer())
for epoch in range(n_epochs):
_, curr_loss = sess.run([optimizer, loss], feed_dict={x:train_X, y:train_Y})
if epoch % interval == 0:
print ('Loss after epoch', epoch, ' is ', curr_loss)
print ('Now testing the model in the test set')
final_preds, final_loss = sess.run([pred_y, loss], feed_dict={x:test_X, y:test_Y})
print ('The final loss is: ', final_loss)
# Plotting the final predictions against the true predictions
plt.plot(test_X[:10], test_Y[:10], 'g', label='True Function')
plt.plot(test_X[:10], final_preds[:10], 'r', label='Predicted Function')
plt.legend()
plt.show()
# + [markdown] id="jgmH3wwt1src" colab_type="text"
# Okay, so we are doing good!<br/>
#
# Now, let me just put everything here into one function so that you can tweak the hyperparameters easily!
#
# Or better, do it yourself!
# + id="OZ5TY7B_4E_v" colab_type="code" colab={}
def linear_regression(learning_rate=0.000005, n_epochs=100, interval=50):
# YOUR CODE HERE
x = tf.placeholder(tf.float32, name='x')
y = tf.placeholder(tf.float32, name='y')
W = tf.Variable(0.0, name='weight_1')
b = tf.Variable(0.0, name='bias_1')
pred_y = (W*x) + b
loss = tf.reduce_mean(tf.square(y - pred_y))
optimizer = tf.train.GradientDescentOptimizer(learning_rate=learning_rate).minimize(loss)
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
for epoch in range(n_epochs):
_, curr_loss = sess.run([optimizer, loss], feed_dict={x:train_X, y:train_Y})
if epoch % interval == 0:
print('Loss after epoch', epoch, ' is ', curr_loss)
print('Now testing the model in the test set')
final_preds, final_loss = sess.run([pred_y, loss], feed_dict={x:test_X, y:test_Y})
print('The final loss is: ', final_loss)
plt.plot(test_X[:10], test_Y[:10], 'g', label='True Function')
plt.plot(test_X[:10], final_preds[:10], 'r', label='Predicted Function')
plt.legend()
plt.show()
# + id="ufI5g3dzkl34" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 420} outputId="2512f030-91df-4b48-cfe3-2f88756466cb"
linear_regression()
# + id="6ivaSeXpmx4E" colab_type="code" colab={}
def linear_regression(learning_rate=0.000005, n_epochs=100, interval=50, weight=0.0, bias=0.0):
# YOUR CODE HERE
x = tf.placeholder(tf.float32, name='x')
y = tf.placeholder(tf.float32, name='y')
W = tf.Variable(weight, name='weight_1')
b = tf.Variable(bias, name='bias_1')
pred_y = (W*x) + b
loss = tf.reduce_mean(tf.square(y - pred_y))
optimizer = tf.train.GradientDescentOptimizer(learning_rate=learning_rate).minimize(loss)
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
for epoch in range(n_epochs):
_, curr_loss = sess.run([optimizer, loss], feed_dict={x:train_X, y:train_Y})
if epoch % interval == 0:
print('Loss after epoch', epoch, ' is ', curr_loss)
print('Now testing the model in the test set')
final_preds, final_loss = sess.run([pred_y, loss], feed_dict={x:test_X, y:test_Y})
print('The final loss is: ', final_loss)
plt.plot(test_X, test_Y, 'g', label='True Function')
plt.plot(test_X, final_preds, 'r', label='Predicted Function')
plt.legend()
plt.show()
# + id="A6MaclhK4rc6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 567} outputId="47f0ce71-b824-427b-a712-780e0478e0a4"
# Okay! Now let's tweak!
linear_regression(learning_rate=0.000034, n_epochs=500)
# + id="peoHmV2M40uU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 751} outputId="37439bf7-b27f-4d9f-d8f7-44387225e52b"
linear_regression(learning_rate=0.0000006, n_epochs=1000)
# + [markdown] id="KjY_KnlE5ClG" colab_type="text"
# ## Drive the loss to a minimum.
# + id="JKiHjGN15HPX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 751} outputId="9a5de3eb-af6a-4eea-b494-3976ea865974"
# YOUR CODE HERE
linear_regression(learning_rate=0.00005, n_epochs=100000, interval=5000, weight=1.0, bias=-7.0)
# + id="y8Sm9o0orWZ6" colab_type="code" colab={}
| ayan59dutta.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
#Generate BiPartites and save as Objects.
import itertools as it
import sys
import json
import re
import collections
import os
path='../../data'
filenames = os.listdir(path)
count=0
AllDataPidTitleBipartite={}
AllDataPidTrackListBipartite={}
AllDataAlbumTrackSetBipartite={}
AllDataArtistTrackSetBipartite={}
AllDataTrackArtistBipartite={}
AllDataTrackAlbumBipartite={}
AllDataTrackNameBipartite={}
AllDataAlbumNameBipartite={}
AllDataAritstNameBipartite={}
AllDataPidDescriptionBipartite={}
for filename in sorted(filenames):
if filename.startswith("mpd.slice.") and filename.endswith(".json"):
fullpath = os.sep.join((path, filename))
f = open(fullpath)
js = f.read()
f.close()
mpd_slice = json.loads(js)
for playlist in mpd_slice['playlists']:
playlistId=str(playlist['pid'])
playlistTracks=[]
playlistTitle=playlist['name']
for track in playlist['tracks']:
trackId=track['track_uri']
trackName=track['track_name']
trackArtistId=track['artist_uri']
trackArtistName=track['artist_name']
trackAlbumId=track['album_uri']
trackAlbumName=track['album_name']
playlistTracks.append(trackId)
AllDataAlbumTrackSetBipartite.setdefault(trackAlbumId,[]).append(trackId)
AllDataArtistTrackSetBipartite.setdefault(trackArtistId,[]).append(trackId)
AllDataTrackArtistBipartite[trackId]=trackArtistId
AllDataTrackAlbumBipartite[trackId]=trackAlbumId
AllDataTrackNameBipartite[trackId]=trackName
AllDataAlbumNameBipartite[trackAlbumId]=trackAlbumName
AllDataAritstNameBipartite[trackArtistId]=trackArtistName
AllDataPidTitleBipartite[playlistId]=playlistTitle
AllDataPidTrackListBipartite[playlistId]=playlistTracks
if 'description' in playlist:
AllDataPidDescriptionBipartite[playlistId]=playlist['description']
count=count+1
if count % 10000 ==0:
print 'processed' + str(count)
# -
'''
>>AllDataPidTitleBipartite={}
>>AllDataPidTrackListBipartite={}
>>AllDataAlbumTrackSetBipartite={}
AllDataArtistTrackSetBipartite={}
AllDataTrackArtistBipartite={}
AllDataTrackAlbumBipartite={}
AllDataTrackNameBipartite={}
AllDataAlbumNameBipartite={}
AllDataAritstNameBipartite={}
AllDataPidDescriptionBipartite={}
'''
import pickle
fileObj = open('./BiPartites/AllDataArtistTrackSetBipartite.pkl', 'w')
pickle.dump(AllDataArtistTrackSetBipartite, fileObj)
fileObj.close()
import pickle
fileObj = open('./BiPartites/AllDataTrackArtistBipartite.pkl', 'w')
pickle.dump(AllDataTrackArtistBipartite, fileObj)
fileObj.close()
import pickle
fileObj = open('./BiPartites/AllDataTrackAlbumBipartite.pkl', 'w')
pickle.dump(AllDataTrackAlbumBipartite, fileObj)
fileObj.close()
import pickle
fileObj = open('./BiPartites/AllDataTrackNameBipartite.pkl', 'w')
pickle.dump(AllDataTrackNameBipartite, fileObj)
fileObj.close()
import pickle
fileObj = open('./BiPartites/AllDataAlbumNameBipartite.pkl', 'w')
pickle.dump(AllDataAlbumNameBipartite, fileObj)
fileObj.close()
import pickle
fileObj = open('./BiPartites/AllDataAritstNameBipartite.pkl', 'w')
pickle.dump(AllDataAritstNameBipartite, fileObj)
fileObj.close()
import pickle
fileObj = open('./BiPartites/AllDataPidDescriptionBipartite.pkl', 'w')
pickle.dump(AllDataPidDescriptionBipartite, fileObj)
fileObj.close()
len(AllDataPidTitleBipartite), len(AllDataPidTrackListBipartite), len(AllDataAlbumTrackSetBipartite), len(AllDataArtistTrackSetBipartite), len(AllDataTrackArtistBipartite), len(AllDataTrackAlbumBipartite), len(AllDataTrackNameBipartite), len(AllDataAlbumNameBipartite), len(AllDataAritstNameBipartite)
# # Build Indri Ready Indexes
# # Load Background Pids
#BuildIndriIndexs
#Build750BackgroundIndexes/
import pickle
#backgroundPids=pickle.load(open('SplitsInformation/backgroundDataPids.pkl','rb'))
backgroundPids=AllDataPidTitleBipartite.keys()
len(backgroundPids)
# +
import codecs
def entry(docId, docContent):
entry='<DOC>\n<DOCNO>'+docId+'</DOCNO>\n<TEXT>'+docContent+'\n</TEXT>\n</DOC>'
return entry
def writeIndexToFile(indexDocsList,writePath):
count=0
errors=[]
with codecs.open(writePath,"w", encoding='utf-8') as f:
for line in indexDocsList:
try:
f.write(line +"\n")
count=count+1
except:
errors.append(line)
print count
return errors
# -
# # Build Index Ready Doc - Pid as Document and Tracks of Playlist as Terms
# +
#buildIndexPidAsDocTracksAsTerms
backgroundpidDocs=[]
for pid in backgroundPids:
trackList=AllDataPidTrackListBipartite[pid]
backgroundpidDocs.append(entry(str(pid), ' '.join([item.replace('spotify:track:','') for item in trackList])))
# -
writeIndexToFile(backgroundpidDocs,'./All1MDataIndexes/All1MIndexPidAsDocTracksAsTerms.txt')
len(backgroundpidDocs)
# +
import itertools as it
import sys
import json
import re
import collections
import os
path='../../data'
filenames = os.listdir(path)
count=0
pid750=[]
AlbumTrackSetBipartite750={}
ArtistTrackSetBipartite750={}
for filename in sorted(filenames[:750]):
if filename.startswith("mpd.slice.") and filename.endswith(".json"):
fullpath = os.sep.join((path, filename))
f = open(fullpath)
js = f.read()
f.close()
mpd_slice = json.loads(js)
for playlist in mpd_slice['playlists']:
playlistId=str(playlist['pid'])
pid750.append(playlistId)
for track in playlist['tracks']:
trackId=track['track_uri']
trackName=track['track_name']
trackArtistId=track['artist_uri']
trackArtistName=track['artist_name']
trackAlbumId=track['album_uri']
trackAlbumName=track['album_name']
AlbumTrackSetBipartite750.setdefault(trackAlbumId,[]).append(trackId)
ArtistTrackSetBipartite750.setdefault(trackArtistId,[]).append(trackId)
count=count+1
if count % 10000 ==0:
print 'processed' + str(count)
# -
len(pid750)
albumDocsNorm=[]
albumDocsNonNorm=[]
for albumid, tracks in AlbumTrackSetBipartite750.items():
normTracks=list(set(tracks))
albumDocsNonNorm.append(entry(albumid.strip(), ' '.join(list([item.replace('spotify:track:','') for item in tracks]))))
albumDocsNorm.append(entry(albumid.strip(), ' '.join(list([item.replace('spotify:track:','') for item in normTracks]))))
writeIndexToFile(albumDocsNorm,'./All1MDataIndexes/1MIndexAlbumAsDocNormTracksSetAsTerms.txt')
writeIndexToFile(albumDocsNonNorm,'./All1MDataIndexes/1MIndexAlbumAsDocNonNormTracksListAsTerms.txt')
artistDocsNorm=[]
artistDocsNonNorm=[]
for albumid, tracks in ArtistTrackSetBipartite750.items():
normTracks=list(set(tracks))
artistDocsNonNorm.append(entry(albumid.strip(), ' '.join(list([item.replace('spotify:track:','') for item in tracks]))))
artistDocsNorm.append(entry(albumid.strip(), ' '.join(list([item.replace('spotify:track:','') for item in normTracks]))))
len(ArtistTrackSetBipartite750.items())
writeIndexToFile(artistDocsNorm,'./All1MDataIndexes/1MIndexArtistsAsDocNormTracksSetAsTerms.txt')
writeIndexToFile(artistDocsNonNorm,'./All1MDataIndexes/1MIndexArtistsAsDocNonNormTracksListAsTerms.txt')
# # Normalize Text
# +
import codecs
stop='../stopwords.txt'
emojiSynonyms='../emojiWords.txt'
stopList=[]
emojiMap={}
emojis=[]
f=codecs.open(stop,'r', encoding='utf-8')
for line in f.readlines():
stopList.append(line.strip())
f=codecs.open(emojiSynonyms,'r', encoding='utf-8')
for line in f.readlines():
emoji, meaning= line.strip().split()
emojiMap[emoji.strip()]=' '.join(meaning.strip().split(','))
emojis.append(emoji.strip())
def NormalizeDates(title):
fullDecades=['1930','1940','1950','1960','1970','1980','1990','2000','2010']
truncDecades=['30','40','50','60','70','80','90','00','10']
year=['2001','2002','2003','2004','2005','2006','2007','2008','2009','2011','2012','2013','2014','2015','2016','2017']
truncYear=['02','03','04','05','06','07','08','09','11','12','13','14','15','16','17']
seasons=['spring', 'summer','fall','winter','sommer', 'autumn', 'verano' ]
months=['january','jan','february','feb','march','april','may','june','july','august','aug','spetember','sep','october','oct','novemeber','nov','december','dec']
monthPairs=[('january','jan'),('february','feb'),('august','aug'),('september','sept'), ('september','sep'),('october','oct'),('november','nov'),('december','dec')]
title=title.lower().strip()
for decade in fullDecades:
if decade in title:
newTitle=''
for word in title.split():
if decade in word:
word=word.replace(decade+"'",decade)
word=word.replace(decade+"s",decade)
word=word.replace(decade,' '+decade+'s ')
newTitle=newTitle+' '+word
#print title+'------'+newTitle
title=newTitle.strip()
for truncDecade in truncDecades:
if truncDecade in title and ('19'+truncDecade not in title) and ('20'+truncDecade not in title):
newTitle=''
for word in title.split():
if truncDecade+"'" in word or truncDecade+"s" in word or truncDecade+u"´s" in word or truncDecade+"'s" in word or truncDecade+" s " in word or truncDecade+"ies" in word:
word=word.replace(truncDecade+"'",truncDecade)
word=word.replace(truncDecade+"s",truncDecade)
word=word.replace(truncDecade+" s",truncDecade)
word=word.replace(truncDecade+u"´s",truncDecade)
word=word.replace(truncDecade+"ies",truncDecade)
if truncDecade in truncDecades[:7]:
century='19'
else:
century='20'
word=word.replace(truncDecade,' '+century+truncDecade+'s ')
newTitle=newTitle+' '+word
title=newTitle.strip()
if title.strip() in ['70-80','70,80,90','90/00', '70 80 90', '90 00', '80 90', '80-90','90-00', '60-70', '70 80']:
for truncDecade in truncDecades:
if truncDecade in truncDecades[:7]:
century='19'
else:
century='20'
title=title.replace(truncDecade,' '+century+truncDecade+'s ' )
title=title.strip()
for yr in year:
if yr in title:
title=title.replace(yr,' '+yr+' ')
for yr in truncYear:
if yr in title and '20'+yr not in title:
if "'"+yr in title:
#print title
title=title.replace("'"+yr,' 20'+yr+' ')
#print title
if yr+"'" in title:
#print title
title=title.replace(yr+"'",' 20'+yr+' ')
#print title
if '2k'+yr in title or '2K'+yr in title:
#print title
title=title.replace('2k'+yr,' 20'+yr+' ')
#print title
if '-'+yr in title :
#print title
title=title.replace('-'+yr,' 20'+yr+' ')
#print title
if yr in title and '20'+yr not in title:
for season in seasons:
if season in title:
#print title
title=title.replace(yr,' 20'+yr+' ')
#print title
for month in months:
if month in title and '20'+yr not in title:
#print title
title=title.replace(yr,' 20'+yr+' ')
#print '-'+title
for month,shortMonth in monthPairs:
newTitle=''
if shortMonth in title and month not in title:
#print title
for word in title.split():
if shortMonth== word:
newTitle=newTitle+' '+month
else:
newTitle=newTitle+' '+word
title=newTitle
#print '-'+title
title=title.strip()
title=' '.join(title.split())
return title
def handleEmojis(title):
for emo in emojis:
if emo in title:
title=title.replace(emo,' ')
title=title+' '+emojiMap[emo]+' '
title=title.replace(u'\U0001f3fc','')
title=title.replace(u'\U0001f3fd','')
title=title.replace(u'\U0001f3fb','')
title=title.replace(u'\U0001f3fe','')
title=title.replace(u'\u200d','')
title=title.replace(u'\ufe0f','')
title=title.replace(u'oshrug','shrug')
title=title.replace(u'\ufffd','')
title=title.replace(u'\U0001f37b','')
title=title.replace(u'\u200d','')
title=title.replace(u'\u2640','')
title=title.replace(u'\u2642','')
title=title.replace(u'\U0001f3b6','')
title=title.replace(u'\u2728','')
title=title.replace(u'\U0001f449','')
title=title.replace(u'\U0001f3ff','')
title=title.replace(u'\U0001f38a','')
title=title.replace(u'\U0001f445','')
title=title.replace(u'\U0001f608','')
title=title.replace(u'\U0001f381','')
title=title.replace(u'\U0001f60f','')
title=title.replace(u'\U0001f4a8','')
title=title.replace(u'�','')
title=title.replace('<3',' heart love ')
title=title.replace(':)',' smile happy ')
title=title.replace(';)',' smirk happy ')
title=title.replace(':-)',' smile happy ')
title=title.replace(': )',' smile happy ')
title=title.replace(u'😋',' smile happy ')
title=title.replace(u'\u263a\ufe0f',' smile happy ')
title=title.replace('r&b',' randb ')
title=title.replace('r & b',' randb ')
title=title.replace('rnb',' randb ')
title=title.replace(u'•',' ')
title=title.replace(u'\u263a\ufe0f',' death poison ')
title=title.replace(u'\u2615\ufe0f',' coffee tea morning ')
title=title.replace(u'💩',' poop ')
title=title.strip()
title=' '.join(title.split())
return title
# # ' & - $ . : ! / () * ,
import re
def normalize_nameTitle(name):
name = name.lower()
name = NormalizeDates(name)
name = handleEmojis(name)
name = name.replace('happysmile','happy smile')
name = re.sub(r"[.,\/#\'?\&\-!$%\^\*;:{}=\_`~()@]", ' ', name)
name = re.sub(r'\s+', ' ', name).strip()
return name
def normalize_name(name):
name = name.lower()
name = re.sub(r"[.,\/#\'?\&\-!$%\^\*;:{}=\_`~()@]", ' ', name)
name = re.sub(r'\s+', ' ', name).strip()
return name
# -
normalize_nameTitle('normalize2016 me? :) <3 .....')
# +
import itertools as it
import sys
import json
import re
import collections
import os
path='../../data'
filenames = os.listdir(path)
count=0
pid750=[]
TrackIdTitle750={}
TitleTrackId750={}
TrackIdArtistName750={}
TrackIdAbumName750={}
TrackIdTrackName750={}
AlbumTrackSetBipartite750={}
ArtistTrackSetBipartite750={}
for filename in filenames:
if filename.startswith("mpd.slice.") and filename.endswith(".json"):
fullpath = os.sep.join((path, filename))
f = open(fullpath)
js = f.read()
f.close()
mpd_slice = json.loads(js)
for playlist in mpd_slice['playlists']:
playlistId=str(playlist['pid'])
pid750.append(playlistId)
pname=playlist['name']
normpName=normalize_nameTitle(pname).strip()
if normpName=='':
normpName='emptyTitle'
for track in playlist['tracks']:
trackId=track['track_uri']
trackName=track['track_name']
trackArtistId=track['artist_uri']
trackArtistName=track['artist_name']
trackAlbumId=track['album_uri']
trackAlbumName=track['album_name']
TrackIdTitle750.setdefault(trackId,[]).append(normpName)#--Done
TitleTrackId750.setdefault(normpName,[]).append(trackId)#--Done
TrackIdArtistName750[trackId]=trackArtistName#--meta2
TrackIdAbumName750[trackId]=trackAlbumName#--meta2
TrackIdTrackName750[trackId]=trackName#--meta2
AlbumTrackSetBipartite750.setdefault(trackAlbumId,[]).append(trackId)#done
ArtistTrackSetBipartite750.setdefault(trackArtistId,[]).append(trackId)#done
count=count+1
if count % 10000 ==0:
print 'processed' + str(count)
# -
len(TrackIdTitle750)
titleDocsNorm=[]
titletDocsNonNorm=[]
for title, tracks in TitleTrackId750.items():
normTracks=list(set(tracks))
titletDocsNonNorm.append(entry(title.strip(), ' '.join(list([item.replace('spotify:track:','') for item in tracks]))))
titleDocsNorm.append(entry(title.strip(), ' '.join(list([item.replace('spotify:track:','') for item in normTracks]))))
err1=writeIndexToFile(titleDocsNorm,'./All1MDataIndexes/1MIndexTitlesAsDocNormTracksSetAsTerms.txt')
err2=writeIndexToFile(titletDocsNonNorm,'./All1MDataIndexes/1MIndexTitlesAsDocNonNormTracksListAsTerms.txt')
import nltk
def titlePlusBigrams(title):
bigrm = list(nltk.bigrams(title.split()))
bis=''
for x1,x2 in bigrm:
bis=bis+' '+x1+x2
return title+' '+bis.strip()
len(titleDocsNorm)
# +
trackTitleDocs=[]
for trackId, titleList in TrackIdTitle750.items():
truncTrackId=trackId.replace('spotify:track:','')
concatTitle=''
for title in titleList:
concatTitle=concatTitle+' '+titlePlusBigrams(title)
trackTitleDocs.append(entry(truncTrackId.strip(), concatTitle))
# -
err3=writeIndexToFile(trackTitleDocs,'./All1MDataIndexes/1MIndexTracksAsDocTitlesAsTerms.txt')
# +
#TrackIdArtistName750[trackId]=trackArtistName#--meta2
#TrackIdAbumName750[trackId]=trackAlbumName#--meta2
#TrackIdTrackName750[trackId]=trackName#--meta2
meta2trackDocs=[]
for trackId, trackname in TrackIdTrackName750.items():
truncTrackId=trackId.replace('spotify:track:','')
normTrackName= normalize_nameTitle(trackname)
normAlbumName= normalize_nameTitle(TrackIdAbumName750[trackId])
normArtistName= normalize_nameTitle(TrackIdArtistName750[trackId])
meta2trackDocs.append(entry(truncTrackId,normTrackName+' '+normAlbumName+' '+normArtistName))
# -
err4=writeIndexToFile(meta2trackDocs,'./All1MDataIndexes/1MIndexTracksAsDocMeta2AsTerms.txt')
len(meta2trackDocs)
meta2trackDocs[:50]
| notebooks/10-Challenge-CreateBiPartites-Build_Indri_Index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Multiclass Reductions
# #### by <NAME> and Sö<NAME>
# This notebook demonstrates the reduction of a <a href="http://en.wikipedia.org/wiki/Multiclass_classification">multiclass problem</a> into binary ones using Shogun. Here, we will describe the built-in <a href="http://en.wikipedia.org/wiki/Multiclass_classification#one_vs_all">One-vs-Rest</a>, One-vs-One and Error Correcting Output Codes strategies.
#
#
# In `SHOGUN`, the strategies of reducing a multiclass problem to binary
# classification problems are described by an instance of
# `CMulticlassStrategy`. A multiclass strategy describes
#
# * How to train the multiclass machine as a number of binary machines?
# * How many binary machines are needed?
# * For each binary machine, what subset of the training samples are used, and how are they colored? In multiclass problems, we use *coloring* to refer partitioning the classes into two groups: $+1$ and $-1$, or black and white, or any other meaningful names.
# * How to combine the prediction results of binary machines into the final multiclass prediction?
#
# The user can derive from the virtual class [CMulticlassStrategy](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CMulticlassStrategy.html) to
# implement a customized multiclass strategy. But usually the built-in strategies
# are enough for general problems. We will describe the built-in *One-vs-Rest*,
# *One-vs-One* and *Error-Correcting Output Codes* strategies in this tutorial.
#
# The basic routine to use a multiclass machine with reduction to binary problems
# in shogun is to create a generic multiclass machine and then assign a particular
# multiclass strategy and a base binary machine.
# ## One-vs-Rest and One-vs-One
#
# The *One-vs-Rest* strategy is implemented in
# `MulticlassOneVsRestStrategy`. As indicated by the name, this
# strategy reduce a $K$-class problem to $K$ binary sub-problems. For the $k$-th
# problem, where $k\in\{1,\ldots,K\}$, the samples from class $k$ are colored as
# $+1$, and the samples from other classes are colored as $-1$. The multiclass
# prediction is given as
#
# $$
# f(x) = \operatorname*{argmax}_{k\in\{1,\ldots,K\}}\; f_k(x)
# $$
#
# where $f_k(x)$ is the prediction of the $k$-th binary machines.
#
# The One-vs-Rest strategy is easy to implement yet produces excellent performance
# in many cases. One interesting paper, [<NAME>. and <NAME>. (2004). *In defense of one-vs-all classification*. Journal of Machine
# Learning Research, 5:101–141](http://jmlr.org/papers/v5/rifkin04a.html), it was shown that the
# One-vs-Rest strategy can be
#
# > as accurate as any other approach, assuming that the underlying binary
# classifiers are well-tuned regularized classifiers such as support vector
# machines.
#
# Implemented in [MulticlassOneVsOneStrategy](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1MulticlassOneVsOneStrategy.html), the
# *One-vs-One* strategy is another simple and intuitive
# strategy: it basically produces one binary problem for each pair of classes. So there will be $\binom{K}{2}$ binary problems. At prediction time, the
# output of every binary classifiers are collected to do voting for the $K$
# classes. The class with the highest vote becomes the final prediction.
#
# Compared with the One-vs-Rest strategy, the One-vs-One strategy is usually more
# costly to train and evaluate because more binary machines are used.
#
# In the following, we demonstrate how to use `SHOGUN`'s One-vs-Rest and
# One-vs-One multiclass learning strategy on the USPS dataset. For
# demonstration, we randomly 200 samples from each class for training and 200
# samples from each class for testing.
#
# The [LibLinear](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1LibLinear.html) is used as the base binary classifier in a [LinearMulticlassMachine](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1LinearMulticlassMachine.html), with One-vs-Rest and One-vs-One strategies. The running time and performance (on my machine) is reported below:
# + active=""
# -------------------------------------------------
# Strategy Training Time Test Time Accuracy
# ------------- ------------- --------- --------
# One-vs-Rest 12.68 0.27 92.00%
# One-vs-One 11.54 1.50 93.90%
# -------------------------------------------------
# Table: Comparison of One-vs-Rest and One-vs-One multiclass reduction strategy on the USPS dataset.
# -
# First we load the data and initialize random splitting:
# +
# %pylab inline
# %matplotlib inline
import os
SHOGUN_DATA_DIR=os.getenv('SHOGUN_DATA_DIR', '../../../data')
import numpy as np
from numpy import random
from scipy.io import loadmat
import shogun as sg
import time
mat = loadmat(os.path.join(SHOGUN_DATA_DIR, 'multiclass/usps.mat'))
Xall = mat['data']
#normalize examples to have norm one
Xall = Xall / np.sqrt(sum(Xall**2,0))
Yall = mat['label'].squeeze()
# map from 1..10 to 0..9, since shogun
# requires multiclass labels to be
# 0, 1, ..., K-1
Yall = Yall - 1
N_train_per_class = 200
N_test_per_class = 200
N_class = 10
# to make the results reproducable
random.seed(0)
# index for subsampling
index = np.zeros((N_train_per_class+N_test_per_class, N_class), 'i')
for k in range(N_class):
Ik = (Yall == k).nonzero()[0] # index for samples of class k
I_subsample = random.permutation(len(Ik))[:N_train_per_class+N_test_per_class]
index[:, k] = Ik[I_subsample]
idx_train = index[:N_train_per_class, :].reshape(N_train_per_class*N_class)
idx_test = index[N_train_per_class:, :].reshape(N_test_per_class*N_class)
random.shuffle(idx_train)
random.shuffle(idx_test)
# -
# Convert features into SHOGUN format:
# +
feats_train = sg.create_features(Xall[:, idx_train])
feats_test = sg.create_features(Xall[:, idx_test])
lab_train = sg.create_labels(Yall[idx_train].astype('d'))
lab_test = sg.create_labels(Yall[idx_test].astype('d'))
# -
# define a helper function to train and evaluate multiclass machine given a strategy:
def evaluate(strategy, C):
bin_machine = sg.create_machine("LibLinear", liblinear_solver_type="L2R_L2LOSS_SVC",
use_bias=True, C1=C, C2=C)
mc_machine = sg.create_machine("LinearMulticlassMachine",
multiclass_strategy=strategy,
machine=bin_machine,
labels=lab_train)
t_begin = time.process_time()
mc_machine.train(feats_train)
t_train = time.process_time() - t_begin
t_begin = time.process_time()
pred_test = mc_machine.apply(feats_test)
t_test = time.process_time() - t_begin
evaluator = sg.create_evaluation("MulticlassAccuracy")
acc = evaluator.evaluate(pred_test, lab_test)
print("training time: %.4f" % t_train)
print("testing time: %.4f" % t_test)
print("accuracy: %.4f" % acc)
# Test on One-vs-Rest and One-vs-One strategies.
# +
print("\nOne-vs-Rest")
print("="*60)
evaluate(sg.create_multiclass_strategy("MulticlassOneVsRestStrategy"), 5.0)
print("\nOne-vs-One")
print("="*60)
evaluate(sg.create_multiclass_strategy("MulticlassOneVsOneStrategy"), 2.0)
# -
# LibLinear also has a true multiclass SVM implemenentation - so it is worthwhile to compare training time and accuracy with the above reduction schemes:
# +
mcsvm = sg.create_machine("MulticlassLibLinear", C=5.0,
labels=lab_train, use_bias=True)
t_begin = time.process_time()
mcsvm.train(feats_train)
t_train = time.process_time() - t_begin
t_begin = time.process_time()
pred_test = mcsvm.apply(feats_test)
t_test = time.process_time() - t_begin
evaluator = sg.create_evaluation("MulticlassAccuracy")
acc = evaluator.evaluate(pred_test, lab_test)
print("training time: %.4f" % t_train)
print("testing time: %.4f" % t_test)
print("accuracy: %.4f" % acc)
# -
# As you can see performance of all the three is very much the same though the multiclass svm is a bit faster in training. Usually training time of the true multiclass SVM is much slower than one-vs-rest approach. It should be noted that classification performance of one-vs-one is known to be slightly superior to one-vs-rest since the machines do not have to be properly scaled like in the one-vs-rest approach. However, with larger number of classes one-vs-one quickly becomes prohibitive and so one-vs-rest is the only suitable approach - or other schemes presented below.
# ## Error-Correcting Output Codes
#
# *Error-Correcting Output Codes* (ECOC) is a
# generalization of the One-vs-Rest and One-vs-One strategies. For example, we
# can represent the One-vs-Rest strategy with the following $K\times K$ coding
# matrix, or a codebook:
#
# $$
# \begin{bmatrix}
# +1 & -1 & -1 & \ldots & -1 & -1 \\\\
# -1 & +1 & -1 & \ldots & -1 & -1\\\\
# -1 & -1 & +1 & \ldots & -1 & -1\\\\
# \vdots & \vdots & \vdots & \ddots & \vdots & \vdots \\\\
# -1 & -1 & -1 & \ldots & +1 & -1 \\\\
# -1 & -1 & -1 & \ldots & -1 & +1
# \end{bmatrix}
# $$
#
# Denote the codebook by $B$, there is one column of the codebook associated with
# each of the $K$ classes. For example, the code for class $1$ is
# $[+1,-1,-1,\ldots,-1]$. Each row of the codebook corresponds to a binary
# coloring of all the $K$ classes. For example, in the first row, the class $1$
# is colored as $+1$, while the rest of the classes are all colored as $-1$.
# Associated with each row, there is a binary classifier trained according to the
# coloring. For example, the binary classifier associated with the first row is
# trained by treating all the examples of class $1$ as positive examples, and all
# the examples of the rest of the classes as negative examples.
#
# In this special case, there are $K$ rows in the codebook. The number of rows in
# the codebook is usually called the *code length*. As we can see, this
# codebook exactly describes how the One-vs-Rest strategy trains the binary
# sub-machines.
# +
OvR=-np.ones((10,10))
fill_diagonal(OvR, +1)
_=gray()
_=imshow(OvR, interpolation='nearest')
_=gca().set_xticks([])
_=gca().set_yticks([])
# -
# A further generalization is to allow $0$-values in the codebook. A $0$ for a
# class $k$ in a row means we ignore (the examples of) class $k$ when training
# the binary classifiers associated with this row. With this generalization, we
# can also easily describes the One-vs-One strategy with a $\binom{K}{2}\times K$
# codebook:
#
# $$
# \begin{bmatrix}
# +1 & -1 & 0 & \ldots & 0 & 0 \\\\
# +1 & 0 & -1 & \ldots & 0 & 0 \\\\
# \vdots & \vdots & \vdots & \ddots & \vdots & 0 \\\\
# +1 & 0 & 0 & \ldots & -1 & 0 \\\\
# 0 & +1 & -1 & \ldots & 0 & 0 \\\\
# \vdots & \vdots & \vdots & & \vdots & \vdots \\\\
# 0 & 0 & 0 & \ldots & +1 & -1
# \end{bmatrix}
# $$
#
# Here each of the $\binom{K}{2}$ rows describes a binary classifier trained with
# a pair of classes. The resultant binary classifiers will be identical as those
# described by a One-vs-One strategy.
#
# Since $0$ is allowed in the codebook to ignore some classes, this kind of
# codebooks are usually called *sparse codebook*, while the codebooks with
# only $+1$ and $-1$ are usually called *dense codebook*.
#
# In general case, we can specify any code length and fill the codebook
# arbitrarily. However, some rules should be followed:
#
# * Each row must describe a *valid* binary coloring. In other words, both $+1$ and $-1$ should appear at least once in each row. Or else a binary classifier cannot be obtained for this row.
# * It is good to avoid duplicated rows. There is generally no harm to have duplicated rows, but the resultant binary classifiers are completely identical provided the training algorithm for the binary classifiers are deterministic. So this can be a waste of computational resource.
# * Negative rows are also duplicated. Simply inversing the sign of a code row does not produce a "new" code row. Because the resultant binary classifier will simply be the negative classifier associated with the original row.
# Though you can certainly generate your own codebook, it is usually easier to
# use the `SHOGUN` built-in procedures to generate codebook automatically. There
# are various codebook generators (called *encoders*) in `SHOGUN`. However,
# before describing those encoders in details, let us notice that a codebook
# only describes how the sub-machines are trained. But we still need a
# way to specify how the binary classification results of the sub-machines can be
# combined to get a multiclass classification result.
#
# Review the codebook again: corresponding to each class, there is a column. We
# call the codebook column the (binary) *code* for that class. For a new
# sample $x$, by applying the binary classifiers associated with each row
# successively, we get a prediction vector of the same length as the
# *code*. Deciding the multiclass label from the prediction vector (called
# *decoding*) can be done by minimizing the *distance* between the
# codes and the prediction vector. Different *decoders* define different
# choices of distance functions. For this reason, it is usually good to make the
# mutual distance between codes of different classes large. In this way, even
# though several binary classifiers make wrong predictions, the distance of
# the resultant prediction vector to the code of the *true* class is likely
# to be still smaller than the distance to other classes. So correct results can
# still be obtained even when some of the binary classifiers make mistakes. This
# is the reason for the name *Error-Correcting Output Codes*.
#
# In `SHOGUN`, encoding schemes are described by subclasses of
# [CECOCEncoder](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CECOCEncoder.html), while decoding schemes are described by subclasses
# of [CECOCDecoder](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CECOCDecoder.html). Theoretically, any combinations of
# encoder-decoder pairs can be used. Here we will introduce several common
# encoder/decoders in shogun.
# * [CECOCRandomDenseEncoder](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CECOCRandomDenseEncoder.html): This encoder generate random dense ($+1$/$-1$) codebooks and choose the one with the largest *minimum mutual distance* among the classes. The recommended code length for this encoder is $10\log K$.
# * [CECOCRandomSparseEncoder](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CECOCRandomSparseEncoder.html): This is similar to the random dense encoder, except that sparse ($+1$/$-1$/$0$) codebooks are generated. The recommended code length for this encoder is $15\log K$.
# * [CECOCOVREncoder](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CECOCOVREncoder.html), [CECOCOVOEncoder](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CECOCOVOEncoder.html): These two encoders mimic the One-vs-Rest and One-vs-One strategies respectively. They are implemented mainly for demonstrative purpose. When suitable decoders are used, the results will be equivalent to the corresponding strategies, respectively.
# Using ECOC Strategy in `SHOGUN` is similar to ordinary one-vs-rest or one-vs-one. You need to choose an encoder and a decoder, and then construct a `ECOCStrategy`, as demonstrated below:
print("\nRandom Dense Encoder + Margin Loss based Decoder")
print("="*60)
evaluate(sg.ECOCStrategy(sg.ECOCRandomDenseEncoder(), sg.ECOCLLBDecoder()), 2.0)
# ### Using a kernel multiclass machine
# Expanding on the idea of creating a generic multiclass machine and then assigning a particular multiclass strategy and a base binary machine, one can also use the [KernelMulticlassMachine](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1KernelMulticlassMachine.html) with a kernel of choice.
#
# Here we will use a [GaussianKernel](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1GaussianKernel.html) with [LibSVM](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CLibSVM.html) as the classifer.
# All we have to do is define a new helper evaluate function with the features defined as in the above examples.
# +
def evaluate_multiclass_kernel(strategy):
width=2.1
epsilon=1e-5
kernel=sg.create_kernel("GaussianKernel", width=width)
kernel.init(feats_train, feats_train)
classifier = sg.create_machine("LibSVM", epsilon=epsilon)
mc_machine = sg.create_machine("KernelMulticlassMachine",
multiclass_strategy=strategy,
kernel=kernel,
machine=classifier,
labels=lab_train)
t_begin = time.process_time()
mc_machine.train()
t_train = time.process_time() - t_begin
t_begin = time.process_time()
pred_test = mc_machine.apply_multiclass(feats_test)
t_test = time.process_time() - t_begin
evaluator = sg.create_evaluation("MulticlassAccuracy")
acc = evaluator.evaluate(pred_test, lab_test)
print("training time: %.4f" % t_train)
print("testing time: %.4f" % t_test)
print("accuracy: %.4f" % acc)
print("\nOne-vs-Rest")
print("="*60)
evaluate_multiclass_kernel(sg.create_multiclass_strategy("MulticlassOneVsRestStrategy"))
# -
# So we have seen that we can classify multiclass samples using a base binary machine. If we dwell on this a bit more, we can easily spot the intuition behind this.
#
# The [MulticlassOneVsRestStrategy](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1MulticlassOneVsOneStrategy.html) classifies one class against the rest of the classes. This is done for each and every class by training a separate classifier for it.So we will have total $k$ classifiers where $k$ is the number of classes.
#
# Just to see this in action lets create some data using the gaussian mixture model class ([GMM](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CGMM.html)) from which we sample the data points.Four different classes are created and plotted.
# +
num=1000;
dist=1.0;
gmm=sg.GMM(4)
gmm.set_nth_mean(array([-dist*4,-dist]),0)
gmm.set_nth_mean(array([-dist*4,dist*4]),1)
gmm.set_nth_mean(array([dist*4,dist*4]),2)
gmm.set_nth_mean(array([dist*4,-dist]),3)
gmm.set_nth_cov(array([[1.0,0.0],[0.0,1.0]]),0)
gmm.set_nth_cov(array([[1.0,0.0],[0.0,1.0]]),1)
gmm.set_nth_cov(array([[1.0,0.0],[0.0,1.0]]),2)
gmm.set_nth_cov(array([[1.0,0.0],[0.0,1.0]]),3)
gmm.put('coefficients', array([1.0,0.0,0.0,0.0]))
x0=np.array([gmm.sample() for i in range(num)]).T
x0t=np.array([gmm.sample() for i in range(num)]).T
gmm.put('coefficients', array([0.0,1.0,0.0,0.0]))
x1=np.array([gmm.sample() for i in range(num)]).T
x1t=np.array([gmm.sample() for i in range(num)]).T
gmm.put('coefficients', array([0.0,0.0,1.0,0.0]))
x2=np.array([gmm.sample() for i in range(num)]).T
x2t=np.array([gmm.sample() for i in range(num)]).T
gmm.put('coefficients', array([0.0,0.0,0.0,1.0]))
x3=np.array([gmm.sample() for i in range(num)]).T
x3t=np.array([gmm.sample() for i in range(num)]).T
traindata=np.concatenate((x0,x1,x2,x3), axis=1)
testdata=np.concatenate((x0t,x1t,x2t,x3t), axis=1)
l0 = np.array([0.0 for i in range(num)])
l1 = np.array([1.0 for i in range(num)])
l2 = np.array([2.0 for i in range(num)])
l3 = np.array([3.0 for i in range(num)])
trainlab=np.concatenate((l0,l1,l2,l3))
testlab=np.concatenate((l0,l1,l2,l3))
# -
_=jet()
_=scatter(traindata[0,:], traindata[1,:], c=trainlab, s=100)
# Now that we have the data ready , lets convert it to shogun format features.
feats_tr=sg.create_features(traindata)
labels=sg.create_labels(trainlab)
# The [KernelMulticlassMachine](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1KernelMulticlassMachine.html) is used with [LibSVM](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CLibSVM.html) as the classifer just as in the above example.
#
# Now we have four different classes, so as explained above we will have four classifiers which in shogun terms are submachines.
#
# We can see the outputs of two of the four individual submachines (specified by the index) and of the main machine. The plots clearly show how the submachine classify each class as if it is a binary classification problem and this provides the base for the whole multiclass classification.
# +
width=2.1
epsilon=1e-5
kernel=sg.create_kernel("GaussianKernel", width=width)
kernel.init(feats_tr, feats_tr)
classifier=sg.create_machine("LibSVM", epsilon=epsilon)
mc_machine=sg.create_machine("KernelMulticlassMachine",
multiclass_strategy=sg.create_multiclass_strategy("MulticlassOneVsRestStrategy"),
kernel=kernel,
machine=classifier,
labels=labels)
mc_machine.train()
size=100
x1=linspace(-10, 10, size)
x2=linspace(-10, 10, size)
x, y=meshgrid(x1, x2)
grid=sg.create_features(np.array((ravel(x), ravel(y)))) #test features
out=mc_machine.apply_multiclass(grid) #main output
z=out.get("labels").reshape((size, size))
sub_out0=sg.as_machine(mc_machine.get("machines", 0)).apply_binary() #first submachine
sub_out1=sg.as_machine(mc_machine.get("machines", 1)).apply_binary() #second submachine
z0=sub_out0.get_labels().reshape((size, size))
z1=sub_out1.get_labels().reshape((size, size))
figure(figsize=(20,5))
subplot(131, title="Submachine 1")
c0=pcolor(x, y, z0)
_=contour(x, y, z0, linewidths=1, colors='black', hold=True)
_=colorbar(c0)
subplot(132, title="Submachine 2")
c1=pcolor(x, y, z1)
_=contour(x, y, z1, linewidths=1, colors='black', hold=True)
_=colorbar(c1)
subplot(133, title="Multiclass output")
c2=pcolor(x, y, z)
_=contour(x, y, z, linewidths=1, colors='black', hold=True)
_=colorbar(c2)
# -
# The `MulticlassOneVsOneStrategy` is a bit different with more number of machines.
# Since it trains a classifer for each pair of classes, we will have a total of $\frac{k(k-1)}{2}$ submachines for $k$ classes. Binary classification then takes place on each pair.
# Let's visualize this in a plot.
# +
C=2.0
bin_machine = sg.create_machine("LibLinear", liblinear_solver_type="L2R_L2LOSS_SVC",
use_bias=True, C1=C, C2=C)
mc_machine1 = sg.create_machine("LinearMulticlassMachine",
multiclass_strategy=sg.create_multiclass_strategy("MulticlassOneVsOneStrategy"),
machine=bin_machine,
labels=labels)
mc_machine1.train(feats_tr)
out1=mc_machine1.apply_multiclass(grid) #main output
z1=out1.get_labels().reshape((size, size))
sub_out10=sg.as_machine(mc_machine.get("machines", 0)).apply_binary() #first submachine
sub_out11=sg.as_machine(mc_machine.get("machines", 1)).apply_binary() #second submachine
z10=sub_out10.get_labels().reshape((size, size))
z11=sub_out11.get_labels().reshape((size, size))
no_color=array([5.0 for i in range(num)])
figure(figsize=(20,5))
subplot(131, title="Submachine 1") #plot submachine and traindata
c10=pcolor(x, y, z10)
_=contour(x, y, z10, linewidths=1, colors='black', hold=True)
lab1=concatenate((l0,l1,no_color,no_color))
_=scatter(traindata[0,:], traindata[1,:], c=lab1, cmap='gray', s=100)
_=colorbar(c10)
subplot(132, title="Submachine 2")
c11=pcolor(x, y, z11)
_=contour(x, y, z11, linewidths=1, colors='black', hold=True)
lab2=concatenate((l0, no_color, l2, no_color))
_=scatter(traindata[0,:], traindata[1,:], c=lab2, cmap="gray", s=100)
_=colorbar(c11)
subplot(133, title="Multiclass output")
c12=pcolor(x, y, z1)
_=contour(x, y, z1, linewidths=1, colors='black', hold=True)
_=colorbar(c12)
# -
# The first two plots help us visualize how the submachines do binary classification for each pair. The class with maximum votes is chosen for test samples, leading to a refined multiclass output as in the last plot.
| doc/ipython-notebooks/multiclass/multiclass_reduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CS207 Systems Development Final Project:
#
# ## Automatic Differentiation package: genericdiff
# ## TopCoderKitty-ML
#
# **Collaborators**: <NAME>, <NAME> and <NAME>
#
# <hr style="height:2pt">
# ## Introduction
#
# ### Overview
#
# Derivatives play an integral role in computational science, ranging from its use in gradient descent, Newton's method, to finding the posteriors of Bayesian models. We discuss numerical differentiation, symbolic differentiation, how both demonstrate limitations, and automatic differentiation, the focus of our software. We acknowledge its effectiveness in both its accuracy and efficiency when evaluating derivatives, provide real life applications in the biological context, as well as a guide to how to use our software.
#
# ### Motivation for Automatic Differentiation
#
# Because functions are often too complex to solve analytically, instead, we look to alternative methods that automatically calculate derivatives. There are three main ways to approach this issue: numerical differentiation from finding finite difference approximations, symbolic differentiation through expression manipulation, and automatic differentiation (AD or algorithmic differentiation). While numerical differentiation is easy to code, it is also subject to floating point errors; symbolic differentiation gives exact and accurate results, but is too computationally expensive. Thus, automatic differentiation proves to be the most effective method as it works to resolve both of these issues; AD is both exact/numerically stable and computationally efficient.
# ## Background
# ### What is AD?
# Conceptually straightforward, automatic differentiation can be defined as a family of techniques that evaluate the derivative through the use of elementary arithmetic operations (ie. addition, subtraction, etc.), elementary arithmetic functions (ie. exp, log, sin), and the chain rule. AD is a recursive process that involves repeatedly taking the chain rule to the elementary operations at hand, and allows us to calculate the individual components of the gradient (a list of partial derivatives in terms of each of the inputs) evaluations, to produce results that are automatic and precise. Because AD involves a specific family of techniques that compute derivatives through accumulation of values during code execution to generate numerical derivative evaluations rather than derivative expressions, it can attain machine precision. There are two modes in AD: the forward mode and reverse mode.
# ### Forward Mode
# The forward mode begins at the innermost portion of the function and repeatedly, or recursively, applies the chain rule while traversing out. Thus, the forward pass creates the evaluation trace, which is a composition of the finite set of elementary operations for which derivatives are known. This is then combined to evaluate the derivative of the overall composition. Notably, the derivative at subsequent steps are calculated based on the derivatives calculated in preceding steps. It is also important to note that the forward pass also finds the derivatives and the values of the partial derivatives at each step, thus requiring the setting of the seed vector, which indicates which input variable to the take the partial derivative in terms of. Taking the derivative of the m dependent output variables in terms of a single independent input variable make up one column of the jacobian matrix. Thus, the full jacobian matrix can be defined as the partial derivative of the m output variables in terms of the n input variables, or applying the forward pass across n evaluations. These recursive steps can be documented in a table, and visually represented through a computational graph.
#
# The forward mode can be simplified by utilizing another important component of automatic differentiation: dual numbers. Dual numbers are a type of number that uses and allows for simultaneously automatically differentiating a function while also evaluating the value of the function.
# The forward mode is efficient and straight- forward because it is able to compute all the derivatives in terms of one input with just one forward pass.
# ### Reverse mode
# In the backward mode, a forward pass creates the evaluation trace and indicates the partial derivatives at each step, but does not find the values of the partial derivatives. At the end of this process, the final node’s derivative is evaluated by using an arbitrary seed. Then, the values of the partial derivatives that constitute the end node’s derivative are found by performing a backward pass through the tree to get all the values.
# During both the forward and backward modes, all intermediate variables are evaluated, and their values are stored; these steps can be represented in a table, and further visualized in a computational graph. The graph (and table) essentially outlines this repeated chain rule process; it also serves as the basis of the logic behind our automatic differentiation software library.
#
# ### Jacobian product
# Auto differentiation can take in a vector of input functions with multiple variables. In this case, the autodifferentiation algorithm returns the Jacobian Product matrix which is just a matrix where each row represents the partial derivatives of the variables of a function within the vector. If the vector has m functions then the Jacobian product will have m rows. If the functions contain n variables, the Jacobian will contain n columns. The Jacobian product matrix is handy in first order derivative applications in the sciences when you are dealing with lots of different systems (functions) and unknowns in the systems (variables). We will see an application of this in the RNA velocity package we will be building.
#
# ### Application of AD
# AD can be applied to many branches of computational science, ranging from areas in biology to politics. We chose to focus our application in a biological context, namely RNA velocity. This involves a system of differential equations with pretty complicated solutions, that is able to predict the future state of individual cells through single cell RNA gene expression. Estimating this is important in aiding the analysis of developmental lineages and cellular dynamics, particularly in humans.
# ## Software organization
#
# Our implentation philosophy was to create an object such that all the elemental functions and constants could be arbritrarily combined and nested for differentiation.
#
# We created a ```genericdiff``` library that contains the following modules:
# - ```__init__.py``` : Initializes the genericdiff package
#
#
# - ```generic_diff.py``` : Contains GenericDiff class where any specialized classes such as sine or cos class were reduced to just value and derivative attributes such that we could use our overloaded operators. +, -, /, * , **, and negation were defined according to to the following differentiation rules: chain rule, generalized power rule, quotient rule and product rule. It also overloaded comparison operators, which include <, >, <=, >=, ==, and !=.
#
#
# - ```elemental_functions.py```: Specialized classes in the following modules that inherited from GenericDiff- exponential, sin, cos, tan, sinh, cosh, tanh, acos, asin, atan, exp, log, logit, sqrt classes were defined here.
#
#
# - ```vector_jacobian.py```: A class to handle taking in a vector of functions, multiple inputs and producing a jacobian product matrix
#
# We created a test package to test our generic_diff, elemental_functions, and vector_jacobian modules for all methods and error handling:
#
# - ```test_elemental_functions.py```
#
# - ```test_generic_diff.py```
#
# - ```test_generic_diff_comparisons.py```
#
# - ```test_vector_jacobian.py```
#
# The test suite is run through pytest and called through Travis CI using the travis yml file - it sends a report to codecov for code coverage.
#
# The driver scripts are located in /driverscripts is:
#
# - ```driver_univariate.py``` This solves for roots using newton's method - the univariate case is using GenericDiff class
# - ```driver_multiivariate.py``` This solves for roots using newton's method - the multivariate case is using JacobianProduct class
# - ```driver_jacobian_product.py``` Finds a jacobian product matrix - uses JacobianProduct class
#
# ## Implementation
#
# We wanted to keep our implementation lightweight. Therefore, we do not use any imports except python's standard math library for our single function, single input use case. In our jacobian vector product use case, we will use numpy.
#
# The core data structure is our GenericDiff class which has at its base, a value attribute and a derivative attribute. This can be combined or taken in by a more specialized class that differentiates specialized mathematical functions like sine, cosine, exp, etc. We have explained how this works based on the software organization above. Given this data structure, we can arbritrarily combine objects to represent extremely complex mathematical functions.
#
# In our jacobian product matrix we will utilize arrays to take care of multiple inputs.
#
# Our core classes are:
#
# - JacobianProduct: This class takes in function vectors and allows us to find partial derivatives and jacobian products given a set of input values. This is the user-facing class that takes care of multiple function, multiple input use cases.
# - GenericDiff: This is the underlying class that powers JacobianProduct. It takes care of uni-variate differentiation, all overloaded operator functions, variable and constant instantiation rules
# - Var: this is a variable class that inherits from GenericDiff. This identifies a variable in a function expression.
# - Constant: This is a constant class that inherits from GenericDiff. This converts any floats or integers in function expressions into a constant to assign constants with a derivative value of 0
# - Elemental functions: These classes are taken care of in the elemental_functions.py module
#
# The important attributes in our GenericDiff class are:
# - .val = value
# - .der = derivative
# - NOTE the end user does not need to interact directly with these, as the user can just use the JacobianProduct class and call methods in that class to find the values and derivatives.
#
# The external dependencies are:
# - math
# - numpy
#
# Elementary functions covered:
#
# - sin
# - cos
# - tan
# - sinh
# - cosh
# - tanh
# - acos
# - asin
# - atan
# - log
# - logit
# - sqrt
# - exp
# - powers
# - multiplication
# - division
# - addition
# - subtraction
# - unary negation
#
# ## How to use
#
# ```genericdiff``` differentiates by using the forward mode of automatic differentiation.
#
# To do this, one just has to instantiate the ```JacobianProduct``` object, which takes in vectors of functions with multiple inputs to return the jacobian product matrices. ```JacobianProduct``` classes can take in single function, single inputs as well.
#
# \* Developers looking to customize further, can use the single variable, single input ```GenericDiff``` class by instantiating a Var object. One can then combine Var objects with constants using various operations to arrive at derivative evaluations.
# ### Getting setup
#
# Installing the library is straight- forward.
#
# 1. Install the following from pip using the command line.
python -m pip install rnavelocity-genericdiff
# 2. Next, import the package:
import sys
sys.path.append("../")
from genericdiff import *
# ### Class and method examples
# ### JacobianProduct
# The JacobianProduct class is the user-facing tool that a layperson should use. This takes care of all single, multi input and multi function use cases. We will highlight some code showing how our underlying GenericDiff class (used for the single variate case and powers JacobianProduct) works later in this documentation but it is not something a user needs to call directly.
#
# The way in which this package can differentiate automatically is through the instantiation of a JacobianProduct class, which contains a vector of functions as its input. The partial method calculates the partial derivative with respect to a given variable, and the jacobian_product method gets the jacobian product matrix with respect to all variables. Here is a demo of the partial method being called with respect to x, as well as the jacobian_product method:
# +
f = lambda x, y: x**2 - y**3
h = lambda x, y: x**3 + y**3
function_vector = [f, h]
jp_object = JacobianProduct(function_vector)
# -
# getting partial (derivative) with respect to x (index 0 in lambdas)
inputs = [[1, 2, 3], 0]
partial_der_wrt = jp_object.partial_ders(wrt=0, inputs=inputs)
print(partial_der_wrt)
# getting partial (value) with respect to x (index 0 in lambdas)
inputs = [[1, 2, 3], 0]
partial_val_wrt = jp_object.partial_vals(wrt=0, inputs=inputs)
print(partial_val_wrt)
# +
f = lambda x, y: x**2 - y**3
h = lambda x, y: x**3 + y**3
function_vector = [f, h]
jp_object = JacobianProduct(function_vector)
# inputs are x = {1, 2, 3} and y = {1, 2, 3}
# this means we will calculate derivatives for
# (1, 1), (2, 2), and (3, 3)
inputs = [[1, 2, 3], [1, 2, 3]]
# getting jp matrix with respect to all variables
jp_matrix = jp_object.jacobian_product(inputs)
print(jp_matrix)
# -
# ### GenericDiff
#
# NOTE this class powers JacobianProduct and might be useful for developers, but all functionality of this class is captured in JacobianProduct in a more user friendly way.
#
# We can instantiate our GenericDiff objects for the uni-variate use case using the ```Var``` class to instantiate a variable. We can retrieve the .val and .der attributes from the GenericDiff object.
#
# Let's demonstrate this using our operator functions. We can combine two objects together (variable or constant) and get their values and derivatives using the mathematical operators:
#
# 1. subtraction -
# 2. addition +
# 3. division /
# 4. multiplication *
# 5. power **
# 6. negation -( )
#
# Here we demo the + operator for the single function single input use case:
h = Var(4) + 2
print("The value of x + 2 is:", h.val)
print("The derivative of x + 2 is:", h.der)
# Here we demo the power operator:
# +
g = Var(4) ** 2
print("The value of x ** 2 is:", g.val)
print("The derivative of x ** 2 is:", g.der)
# -
# We can also apply trigonometric and exp functions, which inherit from the GenericDiff class.
#
# The available functions are:
#
# We can also apply trigonometric and exp functions, which inherit from the GenericDiff class.
#
# The available functions are:
#
# 1. sine ```sin(x)```
# 2. cosine ```cos(x)```
# 3. tangent ```tan(x)```
# 4. $e^x$ ```exponential(x)```
# 5. hyperbolic sine ```sinh(x)```
# 6. hyperbolic cosine ```cosh(x)```
# 7. hyperbolic tangent ```tanh(x)```
# 8. arc sine ```asin(x)```
# 9. arc cosine ```acos(x)```
# 10. arc tangent ```atan(x)```
# 11. log ```log(x, base=e)``` numpy default base is e, but user can set any base they want
# 12. logit ```logit(x)``` $\frac{e^x}{1+e^x}$
# 13. square root ```sqrt(x)``` $\sqrt{x}$
#
#
# Here we demo the sinh function by applying it to the x + 2 function we created above:
#
#
# +
j = sinh(Var(4) + 2)
print("The value of sinh(x+2) is:", j.val)
print("The derivative of sinh(x+2) is:", j.der)
# -
# Note that the derivative in this case is ```j.der```. It is a scalar value since we are only doing single input single functions.
#
# We can further complicate this by exponentiating:
# +
g = exp(j)
print("The value of exp(sinh(x+2)) is:", g.val)
print("The derivative of exp(sinh(x+2)) is:", g.der)
# -
# We can use the comparison operators to compare the derivatives of the objects or compmare their derivatives to a value
print(j == g)
print(j != g)
print(j <= g)
print(j >= g)
print(j > g)
print(j < g)
print(j > 201)
# ## Advanced features
#
# We have applied our package to creating a biology package focused on ordering cells given gene expression values. Please read the documentation in the ```/rnavelocity folder``` for more information. Here is a brief primer.
#
# ### rnavelocity: derivative of single-cell gene expression state
#
# This package calculates the RNA velocities for a given set of genes and their cells using their expression values. We can think of RNA velocities as first order derivatives specified from a functional form. The applications of RNA velocities are:
#
# 1. To serve as important and powerful indicator of future state of individual cells
# 2. To distinguish between unspliced and spliced mRNAs in common single-cell RNA sequencing protocols
# 3. To predict future state of individual cells on a timescale of hours
# 4. To aid analysis of developmental lineages and cellular dynamics, particularly in humans
#
# In the rnavelocity package we specifically attack application 4 where we will optimize for parameters:
#
# 1. alpha $\alpha$ (transcription rate)
# 2. gamma $\gamma$ (decay rate of the product)
#
# We can then plug these optimized parameters into a time sorting algorithm to find the correct ordering of the lineage of cells given their genes. In simpler terms, we want to answer the chicken or egg question: "which cells were reactants that allowed other cells to develop down the line?"
#
# ## User Documentation
# ### JacobianProduct Class ###
# The JacobianProduct class contains the user facing methods for single variate, multi variate, and multi function use cases.
# #### Methods in JacobianProduct Class:
#
# ```JacobianProduct(function_vector)```
# This instantiates the JacobianProduct class with the user defined functions.
#
# Parameters:
#
# function_vector: list
# a list of functions on which to differentiate. For example, function_vector = [lambda x, y: x, lambda x, y: sin(x) + cos(y)]
#
#
# ```JacobianProduct.partial_ders(self, wrt, inputs, fun_idx=-1)```
# Returns the partial derivative with respect to a chosen variable
#
# Parameters:
#
# wrt: int
# index of variable partial derivative is taken with respect to. For example if x is the first variable in the function declaration, setting wrt=0 will differentiate with respect to x.
#
# inputs: list
# chosen input values for variables. Note that the inputs list is sensitive to the order in which you declared your variables in the function vector. The variable with which you are differentiating can take on multiple values. For example, if there are three variables x, y, z and wrt = 0, then you can specify multiple values as such [[1, 2, 3], 0, 0]. Note that variables held constant can only assume ONE value. In this case, y and z are held constant. If non-differentiating variables (in this case, y and z) are inputted with multiple values, this method will throw an error.
#
# fun_idx: int
# index of the functions in which partial derivative is applied. default is set to -1 differentiate for all functions in the function vector. For example, if the fun_idx = 0 then it will only differentiate against the first function in the function vector
#
# Output:
#
# returns a list of np arrays with partial DERIVATIVES for each function. For example, if we have two functions f and g with three x points: [[df/dx1, df/dx2, df/dx3], [dh/dx1, dh/dx2, dh/dx3]]
#
# ```JacobianProduct.partial_vals(self, wrt, inputs, fun_idx=-1)```
# Returns the partial derivative values with respect to a chosen variable. This is a convenience function for those looking to get the function values in the same format as the partial derivative lists.
#
# Parameters:
#
# wrt: int
# index of variable partial derivative is taken with respect to. For example if x is the first variable in the function declaration, setting wrt=0 will differentiate with respect to x.
#
# inputs: list
# chosen input values for variables. Note that the inputs list is sensitive to the order in which you declared your variables in the function vector. The variable with which you are differentiating can take on multiple values. For example, if there are three variables x, y, z and wrt = 0, then you can specify multiple values as such [[1, 2, 3], 0, 0]. Note that variables held constant can only assume ONE value. In this case, y and z are held constant. If non-differentiating variables (in this case, y and z) are inputted with multiple values, this method will throw an error.
#
# fun_idx: int
# index of the functions in which partial derivative is applied. default is set to -1 differentiate for all functions in the function vector. For example, if the fun_idx = 0 then it will only differentiate against the first function in the function vector
#
# Output:
#
# returns a list of np arrays with partial VALUES for each function. For example, if we have two functions f and g with three x points: [[f(x1), f(x2), f(x3)], [h(x1), h(x2), h(x3)]]
#
#
# ```JacobianProduct.jacobian_product(self, inputs, fun_idx=-1)```
# Returns list of jacobian products
#
# Parameters:
#
# inputs: list
# chosen inputs for variables. For exammple, if we have 3 inputs x, y and z. Single inputs can take the form [1, 2, 3] or [[1], [2], [3]]. Multiple inputs can take the form [[1, 2], [1, 2], [1, 2]]. If multiple inputs per variable, the length of inputs for each variable MUST be the same. If not, the method returns an error. The jacobian product will be found for all element-wise sets. In this example, a jacobian product is found for (1, 1, 1) and (2, 2, 2).
#
# fun_idx: int
# index of the functions in which jacobian product is applied. Default is set to -1 differentiate for all functions in the function vector. For example, if the fun_idx = 0 then it will only differentiate against the first function in the function vector
#
# Output:
#
# returns a list of np arrays which are jacobian products for all input sets. For example, if we have the input [[1, 2], [1, 2], [1, 2]], the output will be a list of two jacobian product matrices - one for set (1, 1, 1) and one for set (2, 2, 2).
#
# #### Example
# For more examples please see the vignette section
# +
##### Partial Derivative
f = lambda x, y: x**2 - cos(y)**3
h = lambda x, y: x**3 + cos(y)**3
function_vector = [f, h]
jp_object = JacobianProduct(function_vector)
# getting partial (derivative) with respect to x (index 0 in lambdas)
inputs = [[1, 2, 3], 0]
partial_der_wrt = jp_object.partial_ders(wrt=0, inputs=inputs)
print("======Partials wrt x======")
print(partial_der_wrt)
####### Jacobian Product
# inputs are x = {1, 2, 3} and y = {1, 2, 3}
# this means we will calculate jacobian products for
# (1, 1), (2, 2), and (3, 3)
inputs = [[1, 2, 3], [1, 2, 3]]
# getting jp matrix with respect to all variables
jp_matrix = jp_object.jacobian_product(inputs)
print("======Jacobian Product=======")
print(jp_matrix)
# -
# We've overloaded elemental functions such that a user can use these in their function definitions and they will be automatically converted to GenericDiff objects that are then differentiated in the JacobianProduct class. For examples, see genericdiff_vignette.ipynb. The following are available to use:
# #### Elemental Functions
#
# 1. sine ```sin(x)```
# 2. cosine ```cos(x)```
# 3. tangent ```tan(x)```
# 4. $e^x$ ```exponential(x)```
# 5. hyperbolic sine ```sinh(x)```
# 6. hyperbolic cosine ```cosh(x)```
# 7. hyperbolic tangent ```tanh(x)```
# 8. arc sine ```asin(x)```
# 9. arc cosine ```acos(x)```
# 10. arc tangent ```atan(x)```
# 11. log ```log(x, base=e)``` numpy default base is e, but user can set any base they want
# 12. logit ```logit(x)``` $\frac{e^x}{1+e^x}$
# 13. square root ```sqrt(x)``` $\sqrt{x}$
#
# For the JacobianProduct class function vector, the user can define a vector using a lambda function using the elemental functions in the following way:
f = lambda x: sin(x)
g = lambda x: cos(x)
function_vector = [f, g]
jp_object = JacobianProduct(function_vector)
# We've also overloaded operator functions to take care of any differential operations done to our GenericDiff class. The user does not need to call on these dunder methods explicitly but are here for reference.
# #### Methods in GenericDiff Class:
#
# ```GenericDiff.__add__(self, other)```
#
# Implements the binary arithmetic "+" operation
#
# Returns GenericDiff object with instances, addition value (self + other) and its derivative
#
# ```GenericDiff.__radd__(self, other)```
#
# Implements the binary arithmetic "+" operation
#
# Returns GenericDiff object with instances, addition value (other + self) and its derivative
#
# ```GenericDiff.__sub__(self, other)```
#
# Implements the binary arithmetic "-" operation
#
# Returns GenericDiff object with instances, subtraction value (self - other) and its derivative
#
# ```GenericDiff.__rsub__(self, other)```
#
# Implements the binary arithmetic "-" operation
#
# Returns GenericDiff object with instances, subtraction value (other - self) and its derivative
#
# ```GenericDiff.__mul__(self, other)```
#
# Implements the binary arithmetic "*" operation
#
# Returns GenericDiff object with instances, multiplication value (self * other) and its derivative
#
# ```GenericDiff.__rmul__(self, other)```
#
# Implements the binary arithmetic "*" operation
#
# Returns GenericDiff object with instances, multiplication value (other * self) and its derivative
#
# ```GenericDiff.__truediv__(self, other)```
#
# Implements the binary arithmetic "/" operation
#
# Returns GenericDiff object with instances, division value (self / other) and its derivative
#
# ```GenericDiff.__rtruediv__(self, other)```
#
# Implements the binary arithmetic "/" operation
#
# Returns GenericDiff object with instances, division value (other / self) and its derivative
#
# ```GenericDiff.__pow__(self, other)```
#
# Implements the binary arithmetic "**" operation
#
# Returns GenericDiff object with instances, power value (self ** other) and its derivative
#
# ```GenericDiff.__rpow__(self, other)```
#
# Implements the binary arithmetic "**" operation
#
# Returns GenericDiff object with instances, power value (other ** self) and its derivative
#
# ```GenericDiff.__neg__(self, other)```
#
# Implements the unary negation operation
#
# Returns GenericDiff object with instances, unary negation of value and derivative value
#
# ```GenericDiff.__gt__(self, other)```
# Returns a Boolean stating whether one derivative value is greater than another derivative value.
#
# ```GenericDiff.__lt__(self, other)```
# Returns a Boolean stating whether one derivative value is less than another derivative value.
#
# ```GenericDiff.__le__(self, other)```
# Returns a Boolean stating whether one derivative value is less than or equal another derivative value.
#
# ```GenericDiff.__ge__(self, other)```
# Returns a Boolean stating whether one derivative value is greater than or equal another derivative value.
#
# ```GenericDiff.__ne__(self, other)```
# Returns a boolean stating whether two derivative values are not equal.
#
# ```GenericDiff.__eq__(self, other)```
# Returns a boolean stating whether two derivative values are equal.
#
# Parameters:
#
# other: GenericDiff class or constant
| genericdiff/documentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Field {_field}
# ## Prepare data
# ### Load libraries
# +
import os
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
pd.set_option('display.max_columns', None)
# %matplotlib inline
# -
# ### Load data
# +
from pyspark.sql import SparkSession
from pyspark.sql.functions import *
spark = SparkSession.builder.appName('Python Spark SQL explorer')\
.config('spark.some.config.option', 'some-value')\
.getOrCreate()
data = spark.read.{_dataset_type}('{_dataset_source}')
# -
# ## General info
data.printSchema()
# ###Field [{_field}] was detected as ###categorical, so we perform some frequency exploration.
#
# ### Generate this column dataframe
col_1 = '{_field}'
# ### Describe
data.describe([col_1]).show()
# ### Type detection
data.schema[col_1]
# ### Show first values
data.select([col_1]).show()
# ### Null count
data.where(data[col_1].isNull()).count()
# ### Show Empties
data.where((data[col_1]) == '').show()
# ### Empty count
data.where((data[col_1]) == '').count()
# #### Value Count
data.groupBy([col_1]).count().show()
# #### Unique Value Count
data.groupBy([col_1]).count().count()
# #### Histogram Bar
data.groupBy([col_1]).count().toPandas().plot(kind='bar')
# #### Box
data.groupBy([col_1]).count().toPandas().plot.box()
# #### Area
data.groupBy([col_1]).count().toPandas().plot(kind='area')
# #### Pie
v = data.groupBy([col_1]).count().toPandas()
v.columns = ['Class', 'Count']
v.plot(kind='pie', labels=v['Class'], y = 'Count', autopct='%.2f', fontsize=20, figsize=(10,10), legend=False)
| jupyxplorer/notebooks/cat.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# ### Understand the CRF code and the feature_mapper code.
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
from lxmls import DATA_PATH
import lxmls
import lxmls.sequences.crf_online as crfo
import lxmls.readers.pos_corpus as pcc
import lxmls.sequences.id_feature as idfc
import lxmls.sequences.extended_feature as exfc
from lxmls.readers import pos_corpus
# Load data from the conll task
# +
corpus = lxmls.readers.pos_corpus.PostagCorpus()
train_seq = corpus.read_sequence_list_conll(DATA_PATH + "/train-02-21.conll",
max_sent_len=10, max_nr_sent=1000)
test_seq = corpus.read_sequence_list_conll(DATA_PATH + "/test-23.conll",
max_sent_len=10, max_nr_sent=1000)
dev_seq = corpus.read_sequence_list_conll(DATA_PATH + "/dev-22.conll",
max_sent_len=10, max_nr_sent=1000)
# -
print("There are", len(train_seq), "examples in train_seq")
print("First example:", train_seq[0])
# ### Feature generation
#
# Given a dataset, in order to build the features
#
# - An instance from **`lxmls.sequences.id_feature.IDFeatures(train_data)`** must be instanciated. We will call `feature_mapper` this instanciated object.
# - Then **`feature_mapper.build_features()`** must be executed
#
## Building features
feature_mapper = idfc.IDFeatures(train_seq)
feature_mapper.build_features()
#
#
# #### About feature_mappers
# A ```feature_mapper``` will contain the following attributes:
#
# - the dataset in ```.dataset```
# - if we instantiate the feature mapper with a dataset X then ```feature_mapper.dataset```will be a copy of X
#
#
# - a dictionary of features in ```.feature_dict```
# - this dictionary will default to ```{}```.
# - In order to build the features the feature mapper must call ```.build_features()``` function.
#
#
# - a list of features in ```.feature_list```
# - this list will default to ```[]```.
# - In order to build the list of features the feature mapper must call ```.build_features()``` function.
#
# A ```feature_mapper``` will contain the method
#
# - A method to generate features, ```.build_features```
# - this method will create features using the ```.dataset``.
# - This method will also fill ```.feature_dict``` and ```.feature_list``
#
#
#
len(feature_mapper.feature_list)
## Let's see the features for the first training example
feature_mapper.feature_list[0]
# +
## The previous features can be classified into:
print("\nInitial features:", feature_mapper.feature_list[0][0])
print("\nTransition features:", feature_mapper.feature_list[0][1])
print("\nFinal features:", feature_mapper.feature_list[0][2])
print("\nEmission features:", feature_mapper.feature_list[0][3])
# -
# #### An observation on the features for a given example
#
# All features for all the training examples in are saved in `train_seq` will be saved in ``feature_mapper.feature_list``.
#
# - If `feature_mapper.feature_list[m]` is our feature vector for training example `m`... why it's not a vector?
#
# - Good point! In order to make the algorithm fast, the code is written using dicts, so if we access only a few positions from the dict and compute substractions it will be much faster than computing the substraction of two huge weight vectors. Notice that there are `len(feature_mapper.feature_dict)` features.
len(train_seq), len(feature_mapper.feature_list)
# ### Codification of the features
#
#
# Features are identifyed by **init_tag:**, **prev_tag:**, **final_prev_tag:**, **id:**
#
# - **init_tag:** when they are Initial features
# - Example: **``init_tag:noun``** is an initial feature that describes that the first word is a noun
#
#
# - **prev_tag:** when they are transition features
# - Example: **``prev_tag:noun::noun``** is an transition feature that describes that the previous word was
# a noun and the current word is a noun.
# - Example: **``prev_tag:noun:.``** is an transition feature that describes that the previous word was
# a noun and the current word is a `.` (this is usually foud as the last transition feature since most phrases will end up with a dot)
#
#
#
# - **final_prev_tag:** when they are final features
# - Example: **``final_prev_tag:.``** is a final feature stating that the last "word" in the sentence was a dot.
#
#
# - **id:** when they are emission features
# - Example: **``id:plays::verb``** is an emission feature, describing that the current word is plays and the current hidden state is a verb.
# - Example: **``id:Feb.::noun``** is an emission feature, describing that the current word is "Feb." and the current hidden state is a noun.
#
#
#
#
inv_feature_dict = {word: pos for pos, word in feature_mapper.feature_dict.items()}
feature_mapper.feature_list[0][0]
[inv_feature_dict[x[0]] for x in feature_mapper.feature_list[0][0]]
[inv_feature_dict[x[0]] for x in feature_mapper.feature_list[0][1]]
[inv_feature_dict[x[0]] for x in feature_mapper.feature_list[0][2]]
len(train_seq.x_dict)
# ### Train a crf
## Train crf
crf_online = crfo.CRFOnline(corpus.word_dict, corpus.tag_dict, feature_mapper)
crf_online.num_epochs = 20
crf_online.train_supervised(train_seq)
# +
## You will receive feedback when each epoch is finished,
## note that running the 20 epochs might take a while. After training is done,
## evaluate the learned model on the training, development and test sets.
pred_train = crf_online.viterbi_decode_corpus(train_seq)
pred_dev = crf_online.viterbi_decode_corpus(dev_seq)
pred_test = crf_online.viterbi_decode_corpus(test_seq)
eval_train = crf_online.evaluate_corpus(train_seq, pred_train)
eval_dev = crf_online.evaluate_corpus(dev_seq, pred_dev)
eval_test = crf_online.evaluate_corpus(test_seq, pred_test)
# -
print("CRF - ID Features Accuracy Train: %.3f Dev: %.3f Test: %.3f" \
%(eval_train,eval_dev, eval_test))
| labs/notebooks/learning_structured_predictors/exercise_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **This notebook is an exercise in the [Data Cleaning](https://www.kaggle.com/learn/data-cleaning) course. You can reference the tutorial at [this link](https://www.kaggle.com/alexisbcook/inconsistent-data-entry).**
#
# ---
#
# In this exercise, you'll apply what you learned in the **Inconsistent data entry** tutorial.
#
# # Setup
#
# The questions below will give you feedback on your work. Run the following cell to set up the feedback system.
from learntools.core import binder
binder.bind(globals())
from learntools.data_cleaning.ex5 import *
print("Setup Complete")
# # Get our environment set up
#
# The first thing we'll need to do is load in the libraries and dataset we'll be using. We use the same dataset from the tutorial.
# +
# modules we'll use
import pandas as pd
import numpy as np
# helpful modules
import fuzzywuzzy
from fuzzywuzzy import process
import chardet
# read in all our data
professors = pd.read_csv("../input/pakistan-intellectual-capital/pakistan_intellectual_capital.csv")
# set seed for reproducibility
np.random.seed(0)
# -
# Next, we'll redo all of the work that we did in the tutorial.
# +
# convert to lower case
professors['Country'] = professors['Country'].str.lower()
# remove trailing white spaces
professors['Country'] = professors['Country'].str.strip()
# get the top 10 closest matches to "south korea"
countries = professors['Country'].unique()
matches = fuzzywuzzy.process.extract("south korea", countries, limit=10, scorer=fuzzywuzzy.fuzz.token_sort_ratio)
def replace_matches_in_column(df, column, string_to_match, min_ratio = 47):
# get a list of unique strings
strings = df[column].unique()
# get the top 10 closest matches to our input string
matches = fuzzywuzzy.process.extract(string_to_match, strings,
limit=10, scorer=fuzzywuzzy.fuzz.token_sort_ratio)
# only get matches with a ratio > 90
close_matches = [matches[0] for matches in matches if matches[1] >= min_ratio]
# get the rows of all the close matches in our dataframe
rows_with_matches = df[column].isin(close_matches)
# replace all rows with close matches with the input matches
df.loc[rows_with_matches, column] = string_to_match
# let us know the function's done
print("All done!")
replace_matches_in_column(df=professors, column='Country', string_to_match="south korea")
countries = professors['Country'].unique()
# -
# # 1) Examine another column
#
# Write code below to take a look at all the unique values in the "Graduated from" column.
professors.head()
# +
# TODO: Your code here
graduated = professors['Graduated from'].unique()
graduated.sort()
graduated
# -
# Do you notice any inconsistencies in the data? Can any of the inconsistencies in the data be fixed by removing white spaces at the beginning and end of cells?
#
# Once you have answered these questions, run the code cell below to get credit for your work.
#
# A: Yes there are a couple of places with spaces in the beginning.
# Check your answer (Run this code cell to receive credit!)
q1.check()
# +
# Line below will give you a hint
#q1.hint()
# -
# # 2) Do some text pre-processing
#
# Convert every entry in the "Graduated from" column in the `professors` DataFrame to remove white spaces at the beginning and end of cells.
# +
# TODO: Your code here
professors['Graduated from'] = professors['Graduated from'].str.strip()
# Check your answer
q2.check()
# +
graduated = professors['Graduated from'].unique()
graduated.sort()
graduated
# -
professors.head()
# +
# Lines below will give you a hint or solution code
#q2.hint()
#q2.solution()
# -
# # 3) Continue working with countries
#
# In the tutorial, we focused on cleaning up inconsistencies in the "Country" column. Run the code cell below to view the list of unique values that we ended with.
# +
# get all the unique values in the 'City' column
countries = professors['Country'].unique()
# sort them alphabetically and then take a closer look
countries.sort()
countries
# -
# Take another look at the "Country" column and see if there's any more data cleaning we need to do.
#
# It looks like 'usa' and 'usofa' should be the same country. Correct the "Country" column in the dataframe so that 'usofa' appears instead as 'usa'.
#
# **Use the most recent version of the DataFrame (with the whitespaces at the beginning and end of cells removed) from question 2.**
matches = fuzzywuzzy.process.extract("usa", countries, limit=10, scorer=fuzzywuzzy.fuzz.token_sort_ratio)
matches
# function to replace rows in the provided column of the provided dataframe
# that match the provided string above the provided ratio with the provided string
def replace_matches_in_column(df, column, string_to_match, min_ratio = 74):
# get a list of unique strings
strings = df[column].unique()
# get the top 10 closest matches to our input string
matches = fuzzywuzzy.process.extract(string_to_match, strings,
limit=10, scorer=fuzzywuzzy.fuzz.token_sort_ratio)
# only get matches with a ratio > 90
close_matches = [matches[0] for matches in matches if matches[1] >= min_ratio]
# get the rows of all the close matches in our dataframe
rows_with_matches = df[column].isin(close_matches)
# replace all rows with close matches with the input matches
df.loc[rows_with_matches, column] = string_to_match
# let us know the function's done
print("All done!")
# +
# get all the unique values in the 'Country' column
countries = professors['Country'].unique()
# sort them alphabetically and then take a closer look
countries.sort()
countries
# +
# TODO: Your code here!
replace_matches_in_column(df=professors, column='Country', string_to_match="usa")
# Check your answer
q3.check()
# +
# Lines below will give you a hint or solution code
#q3.hint()
#q3.solution()
# -
# # Congratulations!
#
# Congratulations for completing the **Data Cleaning** course on Kaggle Learn!
#
# To practice your new skills, you're encouraged to download and investigate some of [Kaggle's Datasets](https://www.kaggle.com/datasets).
# ---
#
#
#
#
# *Have questions or comments? Visit the [Learn Discussion forum](https://www.kaggle.com/learn-forum/172650) to chat with other Learners.*
| Kaggle/Courses/Data Cleaning/5-exercise-inconsistent-data-entry.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## anova
# +
import pandas as pd
from scipy import stats
data=pd.read_csv(r"C:\Users\sachi\Desktop\PlantGrowth.csv")
data.boxplot('weight',by='group',figsize=(12,8) )#weight is dependent and group is independent variables because it as 3 variales in it i.e ctrl,trt1 and trt2
ctrl=data['weight'][data.group=='ctrl']
grps=pd.unique(data.group.values)
d_data={grp:data['weight'][data.group==grp] for grp in grps}
k=len(pd.unique(data.group))#number of variables
N=len(data.values)#NUMBER OF items
n=data.groupby('group').size()[0]#unqiune means ctrl as 10 items ,same as trtl1 and trtl2 also as 10 items each
F,P=stats.f_oneway(d_data['ctrl'],d_data['trt1'],d_data['trt2'])
print("f value is:",F)
print("p value is:",P)
# -
| one_way_anova.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1 align="center"> Project Setup </h1>
# #### 1. Setup the project environnement
# #### 2. Install all requirements for the project
# # !pip install numpy
# # !pip install pandas
# # !pip install matplotlib
# # !pip install seaborn
# !pip install dateparser
# #### 3. Importing Dependencies
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import os
import sys
import dateparser
from datetime import datetime as dt
# -
# #### 4. Define some usefull variables
DOCS_PATH = 'docs/'
DATA_PATH = 'data/'
# #### 5. Importing Datasets
commune_df = pd.read_excel(DATA_PATH + "commune.xlsx")
enroll_df = pd.read_csv(DATA_PATH + "enroll.csv")
industry_df = pd.read_csv(DATA_PATH + "industry.csv")
ord_df = pd.read_csv(DATA_PATH + "ord.csv")
quest_df = pd.read_csv(DATA_PATH + "quest.csv")
studyd_df = pd.read_csv(DATA_PATH + "study_domain.csv")
tech_df = pd.read_csv(DATA_PATH + "technology.csv")
transaction_df = pd.read_csv(DATA_PATH + "transaction.csv")
# <br><br>
# <h1 align="center"> Data Processing </h1>
| data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import torch
import torch.fft as fft
import torch
import torch.nn as nn
import torch.nn.functional as F
import torchvision
import torchvision.transforms as transforms
import matplotlib.pyplot as plt
from matplotlib.offsetbox import OffsetImage, AnnotationBbox
import numpy as np
import warnings
warnings.filterwarnings('ignore')
def imscatter(X, images, zoom=2):
ax = plt.gca()
for i, img in enumerate(images):
x, y = X[i, :]
im = OffsetImage(img.numpy().T, zoom=zoom)
ab = AnnotationBbox(im, (x, y), xycoords='data', frameon=False)
ax.add_artist(ab)
ax.update_datalim(X)
ax.autoscale()
ax.set_xticks([])
ax.set_yticks([])
def plotMNIST(images, zoom = 2):
N = len(images)
dgrid = int(np.ceil(np.sqrt(N)))
ex = np.arange(dgrid)
x, y = np.meshgrid(ex, ex)
X = np.zeros((N, 2))
X[:, 0] = x.flatten()[0:N]
X[:, 1] = y.flatten()[0:N]
imscatter(X, images, zoom)
class TransformLowPass(object):
def __init__(self):
self.fft = True
self.lowpass = True
self.flatten = True
self.norm = False
rows, cols = 32,32
crow, ccol = int(rows / 2), int(cols / 2)
mask = np.zeros((rows, cols), np.uint8)
r = 6
center = [crow, ccol]
x, y = np.ogrid[:rows, :cols]
mask_area = (x - center[0]) ** 2 + (y - center[1]) ** 2 <= r*r
mask[mask_area] = 1
self.mask = torch.from_numpy(mask).bool()
# print(torch.masked_select(self.mask,self.mask).shape)
def __call__(self, img):
if self.fft:
img = fft.fft2(img)
if self.lowpass:
img = img*self.mask
if self.flatten:
img = torch.masked_select(img,self.mask)
if self.norm:
img = torch.abs(img)
return img
# +
epochs = 0
batch_size = 100
lr = .001
# ,transforms.Normalize((.5,.5,.5),(.5,.5,.5)),
transform = transforms.Compose([transforms.ToTensor(),TransformLowPass()])
train_dataset = torchvision.datasets.CIFAR10(root="./data",train=True, download=True,transform=transform)
test_dataset = torchvision.datasets.CIFAR10(root="./data",train=False, download=True,transform=transform)
train_loader = torch.utils.data.DataLoader(train_dataset, batch_size=batch_size, shuffle=True)
test_loader = torch.utils.data.DataLoader(test_dataset, batch_size=batch_size, shuffle=True)
classes = ('plane','car','bird','cat','deer','dog','frog','horse','ship','truck')
device = torch.device('cpu')
# -
images, labels = iter(train_loader).next()
images[0].shape
# +
# plotMNIST(images,zoom = 2)
# +
from complexPyTorch.complexLayers import ComplexBatchNorm2d, ComplexConv2d, ComplexLinear
from complexPyTorch.complexFunctions import complex_relu, complex_max_pool2d
class NeuralNet(nn.Module):
def __init__(self):
super(NeuralNet, self).__init__()
self.fc1 = ComplexLinear(339,10)
# self.fc2 = ComplexLinear(100,100)
# self.fc3 = ComplexLinear(100,10)
def forward(self,x):
x = complex_relu(self.fc1(x))
# x = self.fc3(x)
x = x.abs()
return x
model = NeuralNet().to(device)
print(sum(p.numel() for p in model.parameters() if p.requires_grad))
criterion = nn.CrossEntropyLoss()
im = model(images[0])
# +
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(model.parameters(),lr=lr)
for epoch in range(50):
for batch_idx, (images, labels) in enumerate(train_loader):
images = images.to(device)
labels = labels.to(device)
output = model(images)
loss = criterion(output, labels)
optimizer.zero_grad()
loss.backward()
optimizer.step()
if batch_idx % 100 == 0:
print('Train Epoch: {:3} [{:6}/{:6} ({:3.0f}%)]\tLoss: {:.6f}'.format(
epoch,
batch_idx * len(images),
len(train_loader.dataset),
100. * batch_idx / len(train_loader),
loss.item())
)
# Test
with torch.no_grad():
n_correct=0
n_samples = 0
n_class_correct = [0 for _ in range(10)]
n_class_samples = [0 for _ in range(10)]
for images, labels in test_loader:
images = images.to(device)
labels = labels.to(device)
output = model(images)
_,predicted = torch.max(output,1)
n_samples += labels.size(0)
n_correct += (predicted == labels).sum().item()
for i in range(batch_size):
label = labels[i]
pred = predicted[i]
if (label == pred):
n_class_correct[label]+=1
n_class_samples[label] += 1
acc = 100.0 *n_correct / n_samples
print(f"Accuracy of network: {acc}")
for i in range(10):
acc = 100.0 *n_class_correct[i] / n_class_samples[i]
print(f"Accuracy of class {classes[i]}: {acc}")
# -
| ImageAnalysis/CIFAR10-FNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Importing all the required libraries
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
from sklearn.preprocessing import LabelEncoder
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import BaggingClassifier, AdaBoostClassifier
from xgboost import XGBClassifier,XGBRFClassifier
# ### Reading the training dataset to Pandas DataFrame
data = pd.read_csv('train.csv')
data.head()
# ### Getting the target variables to Y variable
Y = data['Severity']
Y.shape
# ### Dropoing the irrelevent columns from training data
data = data.drop(columns=['Severity','Accident_ID','Accident_Type_Code'],axis=1)
data.head()
# ### creating the Label Encoder object which will encode the target severities to numerical form
label_encode = LabelEncoder()
y = label_encode.fit_transform(Y)
y.shape
# ### split the dataset for training and testing purpose
x_train,x_test,y_train,y_test = train_test_split(data,y,test_size = 0.3)
dtc = DecisionTreeClassifier(criterion='entropy',class_weight='balanced',presort=True,max_depth=10)
bag = BaggingClassifier(warm_start=True,n_estimators=150,base_estimator=dtc)
xgbclf = XGBClassifier()
xgbrgclf = XGBRFClassifier()
dtc.fit(x_train,y_train)
predictions = dtc.predict(x_test)
accuracy_score(y_test,predictions)
bag.fit(x_train,y_train)
predictions = bag.predict(x_test)
accuracy_score(y_test,predictions)
xgbclf.fit(x_train,y_train)
predictions = xgbclf.predict(x_test)
accuracy_score(y_test,predictions)
test_data = pd.read_csv('test.csv')
activity_id = test_data['Accident_ID']
test_data = test_data.drop(columns=['Accident_ID','Accident_Type_Code'])
new_pred = dtc.predict(test_data)
test_categories = label_encode.inverse_transform(new_pred)
result_df = pd.DataFrame({'Accident_ID':activity_id,'Severity':test_categories})
result_df.to_csv('Prediction.csv',index=False)
# ### Accuracy - 83.58
| Airplane_Accident - HackerEarth/3. HackerEarth ML - DecisionTree.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import tensorflow as tf
from tensorflow import keras
import pandas as pd
import seaborn as sns
from pylab import rcParams
import matplotlib.pyplot as plt
from matplotlib import rc
from sklearn.model_selection import train_test_split
from pandas.plotting import register_matplotlib_converters
# %matplotlib inline
# %config InlineBackend.figure_format='retina'
register_matplotlib_converters()
sns.set(style='whitegrid', palette='muted', font_scale=1.5)
rcParams['figure.figsize'] = 22, 10
RANDOM_SEED = 42
np.random.seed(RANDOM_SEED)
tf.random.set_seed(RANDOM_SEED)
# -
df=pd.read_csv("PDB_Load_History(2003).csv")
df.dtypes
df.info()
df
df["DateTime"] = pd.to_datetime(df.date.astype(str) + ' ' + (df.hour-1).astype(str), format='%m/%d/%Y %H')
df["DateTime"]=pd.to_datetime(df["DateTime"])
df.set_index("DateTime",inplace=True)
df
sns.lineplot(x=df.index, y="demand", data=df);
# +
df_by_month = df.resample('M').sum()
sns.lineplot(x=df_by_month.index, y="demand", data=df_by_month);
# -
from statsmodels.tsa.seasonal import seasonal_decompose
decomposed=seasonal_decompose(df["demand"],
model='additive'
)
trend=decomposed.trend
sesonal=decomposed.seasonal
residual=decomposed.resid
plt.figure(figsize=(20,16))
plt.subplot(411)
plt.plot(df["demand"],label="Orginal",color='red')
plt.legend(loc='upper left')
plt.subplot(412)
plt.plot(trend,label="Trend",color='red')
plt.legend(loc='upper left')
plt.subplot(413)
plt.plot(sesonal,label="Sesonal",color='red')
plt.legend(loc='upper left')
plt.subplot(414)
plt.plot(residual,label="Residual",color='red')
plt.legend(loc='upper left')
plt.show()
train_size = int(len(df) * 0.9)
test_size = len(df) - train_size
train, test = df.iloc[0:train_size], df.iloc[train_size:len(df)]
print(len(train), len(test))
# +
from sklearn.preprocessing import RobustScaler
f_columns = ['temperature']
f_transformer = RobustScaler()
cnt_transformer = RobustScaler()
f_transformer = f_transformer.fit(train[f_columns].to_numpy())
cnt_transformer = cnt_transformer.fit(train[['demand']])
train.loc[:, f_columns] = f_transformer.transform(train[f_columns].to_numpy())
train['demand'] = cnt_transformer.transform(train[['demand']])
test.loc[:, f_columns] = f_transformer.transform(test[f_columns].to_numpy())
test['demand'] = cnt_transformer.transform(test[['demand']])
# +
def create_dataset(X, y, time_steps=1):
Xs, ys = [], []
for i in range(len(X) - time_steps):
v = X.iloc[i:(i + time_steps)].values
Xs.append(v)
ys.append(y.iloc[i + time_steps])
return np.array(Xs), np.array(ys)
# +
time_steps = 10
# reshape to [samples, time_steps, n_features]
X_train, y_train = create_dataset(train, train.demand, time_steps)
X_test, y_test = create_dataset(test, test.demand, time_steps)
print(X_train.shape, y_train.shape)
# -
model = keras.Sequential()
model.add(
keras.layers.Bidirectional(
keras.layers.LSTM(
units=128,
input_shape=(X_train.shape[1], X_train.shape[2])
)
)
)
model.add(keras.layers.Dropout(rate=0.2))
model.add(keras.layers.Dense(units=1))
model.compile(loss='mean_squared_error', optimizer='adam')
history = model.fit(
X_train, y_train,
epochs=30,
batch_size=34,
validation_split=0.1,
shuffle=False
)
| Time Series/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 2021년 1월 16일 토요일
# ### leetCode - Smallest Range I (Python)
# ### 문제 : https://leetcode.com/problems/smallest-range-i/
# ### 블로그 : https://somjang.tistory.com/entry/leetCode-908-Smallest-Range-I-Python
# ### 첫번째 시도
class Solution:
def smallestRangeI(self, A: List[int], K: int) -> int:
return max(max(A) - min(A) - 2 * K, 0)
| DAY 201 ~ 300/DAY294_[leetCode] Smallest Range I (Python).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Interacting with JavaScript from Python
# For details, see
# - https://naked.readthedocs.io/toolshed_shell.html#javascript-node-js-execution-functions (documentation of the *Naked* package),<br>
# - http://sweetme.at/2014/02/17/a-simple-approach-to-execute-a-node.js-script-from-python (article on the *Naked* package that this notebook is based on),<br>
# - https://stackabuse.com/command-line-arguments-in-node-js (details on command line arguments that also apply to the `execute_js` and `muterun_js` commands),<br>
# - https://stackoverflow.com/questions/16311562/python-json-without-whitespaces (**important** details on whitespaces in JSON strings),<br>
# - https://docs.python.org/3/library/json.html#json.dump, (official Python documentation on the `json.dump` command),<br>
# - https://stackoverflow.com/questions/4351521/how-do-i-pass-command-line-arguments-to-a-node-js-program (illustration of the use of command line arguments within JavaScript), and<br>
# - https://www.json.org/json-en.html (details on JSON).
#
# Note that the execution of JavaScript from Python via the *naked* package depends on Node.js (https://nodejs.org).<br><br>
# There are many resources on the web that describe how to interface Python with JavaScript. Here, we want to run a JavaScript file within Python code. We do this with Node.js and the *Naked* package for Python. There are two relevant functions: `execute_js` and `muterun_js`. Both of them return output via `console.log()`. The former returns that output to the shell / terminal and the latter stores it in a variable that is accessible within Python. Either of them supports ES6 (ECMAScript 6). We start with the former (so check the shell / terminal to see the output).
from Naked.toolshed.shell import execute_js, muterun_js
import json
success = execute_js('./NodeJS_from_Python/testscript_1.js')
success
# This works well. Now, let's pass an argument to Node.js and run a file that does something with that argument. (Again, check the shell / terminal to see the output.)
success = execute_js('./NodeJS_from_Python/testscript_2.js', arguments="Karina")
success
# Let's move on to the command `muterun_js`, using the same JavaScript file. The output will not be passed to the terminal but returned to a variable that is accessible in Python.
response = muterun_js('./NodeJS_from_Python/testscript_2.js', arguments="Karina")
print(response.exitcode)
print(response.stderr)
response.stdout
# We do not only want to send variables to but also receive variables from JavaScript. For both directions, JSON – short for *JavaScript Object Notation* – is the right format. Moreover, JSON is implemented in many languages. Proper syntax (in terms of spaces, colons, and single / double quotations marks) can be very important. Indeed, it is important in the below example: we need to modify the default separators in order to make the JSON string readable for JavaScript. Then, JavaScript translates the transmitted data into variables, does something with it, encrypts it again in a JSON string, and sends the result back to Python. Python also translates it into variables, ...
myArray = [1, 3, -2, 15, 7.1]
myArray_json = json.dumps(myArray, separators=(',', ': ')) # or (',', ':')
print(myArray_json)
response = muterun_js('./NodeJS_from_Python/testscript_3.js', arguments=myArray_json)
if response.exitcode:
print(response.exitcode)
print(response.stderr)
result = 0
else:
result = json.loads(response.stdout)
result
# ... also does something with it, and prints the results.
for i in result:
print(i + 1)
# The file "processargv.js" is also very illustrative. In particular, it shows the first two arguments of the `execute_js` and `muterun_js` commands:
# 1. path to node and
# 2. path to called file.
#
# Try it out!
#
# <img src="./NodeJS_from_Python/stackabuse.png" width="90%">
| NodeJS_from_Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Plot Interaction of Categorical Factors
# In this example, we will visualize the interaction between categorical factors. First, we will create some categorical data. Then, we will plot it using the interaction_plot function, which internally re-codes the x-factor categories to integers.
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
from statsmodels.graphics.factorplots import interaction_plot
# -
np.random.seed(12345)
weight = pd.Series(np.repeat(['low', 'hi', 'low', 'hi'], 15), name='weight')
nutrition = pd.Series(np.repeat(['lo_carb', 'hi_carb'], 30), name='nutrition')
days = np.log(np.random.randint(1, 30, size=60))
fig, ax = plt.subplots(figsize=(6, 6))
fig = interaction_plot(x=weight, trace=nutrition, response=days,
colors=['red', 'blue'], markers=['D', '^'], ms=10, ax=ax)
| docs/source2/examples/notebooks/generated/categorical_interaction_plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
np.set_printoptions(threshold = np.nan, linewidth = 115)
import pickle
# Load with pickle instead of processing images again
training_img_1 = pickle.load(open('1vAll_img_res_Infiltration_1st_half.p', 'rb'))
training_img_2 = pickle.load(open('1vAll_img_res_Infiltration_2nd_half.p', 'rb'))
# -
training_img_one = np.append(training_img_1, training_img_2, axis=0)
training_img_3 = pickle.load(open('1vAll_img_res_Infiltration_3rd_half.p', 'rb'))
training_img_4 = pickle.load(open('1vAll_img_res_Infiltration_4th_half.p', 'rb'))
training_img_two = np.append(training_img_3, training_img_4, axis=0)
# +
training_img = np.append(training_img_one, training_img_two, axis=0)
training_img.shape
# -
val_img = training_img[:2269]
val_img = np.append(val_img, training_img[34029:], axis=0)
training_img = training_img[2269:34029]
print(len(val_img))
print(len(training_img))
print(len(training_img) + len(val_img))
labels_1 = pickle.load(open('1vAll_labels_res_Infiltration_1st_half.p', 'rb'))
labels_2 = pickle.load(open('1vAll_labels_res_Infiltration_2nd_half.p', 'rb'))
labels_3 = pickle.load(open('1vAll_labels_res_Infiltration_3rd_half.p', 'rb'))
labels_4 = pickle.load(open('1vAll_labels_res_Infiltration_4th_half.p', 'rb'))
training_labels = np.append(labels_1, np.append(labels_2, np.append(labels_3, labels_4, axis = 0), axis = 0), axis = 0)
val_labels = training_labels[:2269]
val_labels = np.append(val_labels, training_labels[34029:], axis=0)
training_labels = training_labels[2269:34029]
print(len(val_labels))
print(len(training_labels))
print(len(training_labels) + len(val_labels))
# +
test_img = pickle.load(open('1vAll_test_img.p', 'rb'))
test_labels = pickle.load(open('1vAll_test_labels.p', 'rb'))
print('Labels shape: ', training_labels.shape)
print('Length of test_labels: ', len(test_labels))
print('No. of Infiltration Diagnoses: ', sum(training_labels))
# +
import keras
from keras import models, optimizers, layers, regularizers, metrics, losses
from keras.layers.advanced_activations import LeakyReLU, PReLU, ELU, ReLU, ThresholdedReLU
from keras.layers.core import Dense, Dropout, SpatialDropout2D, Activation
from keras.layers.convolutional import Conv2D, SeparableConv2D
from keras.models import model_from_json, Sequential
from keras.callbacks import Callback
import tensorflow as tf
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
session = tf.Session(config = config)
import matplotlib.pyplot as plt
IMG_SIZE = 256
# Save Comparison model
def save_model(model_name, hist_str, model_str):
pickle.dump(model_name.history, open('Training Histories/'+ hist_str + '.p', 'wb'))
print("Saved " + hist_str + " to Training Histories folder")
# serialize model to JSON
model_name = model.to_json()
with open("CNN Models/" + model_str + ".json", "w") as json_file:
json_file.write(model_name)
# serialize weights to HDF5
model.save_weights("CNN Models/" + model_str + ".h5")
print("Saved " + model_str + " and weights to CNN Models folder")
# Load model architecture and weights NOTE: must compile again
def load_model():
model_str = str(input("Name of model to load: "))
# load json and create model
json_file = open('CNN Models/' + model_str + '.json', 'r')
loaded_model_json = json_file.read()
json_file.close()
loaded_model = model_from_json(loaded_model_json)
# load weights into new model
loaded_model.load_weights("CNN Models/" + model_str + ".h5")
print("Loaded " + model_str + " and weights from CNN Models folder")
return loaded_model
# Load history object
def load_history():
hist_str = str(input("Name of history to load: "))
loaded_history = pickle.load(open('Training Histories/' + hist_str + '.p', 'rb'))
print("Loaded " + hist_str + " from Training Histories folder")
return loaded_history
Infiltration_class_weight = [{0: 1, 1: 12.938}]
### Custom Loss function
#from keras import backend as K
#def LSEP(true_label, pred_label):
# return K.log(1 + K.sum(K.exp(true_label - pred_label)))
class True_Eval(Callback):
def __init__(self, validation_data):
self.validation_data = validation_data
self.total_accuracy = []
self.i_accuracy = []
self.e_accuracy = []
def ie_real_acc(self, prediction):
y_true = self.validation_data[1]
i_acc = 0
i_total = 0
e_acc = 0
e_total = 0
for i in range(0, len(prediction)):
if (y_true[i].round() == 0):
if (prediction[i].round() == y_true[i]):
i_acc += 1
i_total += 1
else:
if (prediction[i].round() == y_true[i]):
e_acc += 1
e_total += 1
return (i_acc/i_total), (e_acc/e_total)
def on_epoch_end(self, epoch, logs={}):
x_val = self.validation_data[0]
y_pred = self.model.predict(x_val)
i_real_acc, e_real_acc = self.ie_real_acc(y_pred)
print ("T Acc: %f" % i_real_acc)
print ("F Acc: %f" % e_real_acc)
self.i_accuracy.append(i_real_acc)
self.e_accuracy.append(e_real_acc)
class MultiLabel_Acc(Callback):
def __init__(self, validation_data):
self.validation_data = validation_data
self.accuracy = []
def getAccuracy(self, prediction):
y_true = self.validation_data[1]
correct = []
total = []
accuracy = []
for i in range(0, len(y_true[0])):
correct.append([0, 0])
total.append([0, 0])
for sample in range(0, len(prediction)):
for neuron in range(0, len(prediction[sample])):
if (y_true[sample][neuron] == 0.0):
if (round(prediction[sample][neuron]) == y_true[sample][neuron]):
correct[neuron][0] += 1
total[neuron][0] += 1
if (y_true[sample][neuron] == 1.0):
if (round(prediction[sample][neuron]) == y_true[sample][neuron]):
correct[neuron][1] += 1
total[neuron][1] += 1
for neuron in range(0, len(correct)):
accuracy.append((correct[neuron][0]/total[neuron][0],
correct[neuron][1]/total[neuron][1]))
return accuracy
def on_epoch_end(self, epoch, logs={}):
y_pred = self.model.predict(self.validation_data[0])
epoch_data = self.getAccuracy(y_pred)
self.accuracy.append(epoch_data)
for i in range(0, len(epoch_data)):
print(" Neuron: #"+ str(i + 1))
print("Zeroes Accuracy:", epoch_data[i][0])
print(" Ones Accuracy:", epoch_data[i][1])
from sklearn.metrics import confusion_matrix
import itertools
def plot_confusion_matrix(cm, classes,
normalize = False,
title = 'Confusion Matrix',
cmap=plt.cm.Blues):
_fontsize = 'xx-large'
if normalize:
cm = cm.astype('float') / cm.sum(axis = 1)[:, np.newaxis]
print("Normalized Confusion Matrix")
else:
print("Confusion Matrix without Normalization")
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title, fontsize=_fontsize)
cb = plt.colorbar()
cb.ax.tick_params(labelsize=15)
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation = 45, fontsize=_fontsize)
plt.yticks(tick_marks, classes, fontsize=_fontsize)
fmt = '.2f' if normalize else 'd'
thresh = cm.min() + 0.2
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment = 'center',
color='white' if cm[i, j] > thresh else 'black',
fontsize=_fontsize)
plt.tight_layout()
plt.ylabel('True Labels', fontsize=_fontsize)
plt.xlabel('Predicted Labels', fontsize=_fontsize)
# Metric Analysis
def _1vAll_accuracy(y_test, pred):
pred = np.squeeze(pred, axis = -1)
pred = np.round_(pred)
pred = pred.astype(dtype = 'uint8')
ft = pred == y_test
accuracy = sum(ft)/len(ft)
print('\t Complete Label Accuracy: %.2f' % round((accuracy * 100), 2), '%')
print('Sum of Fully Correct Predictions: ', sum(ft))
print('\t\t Total Labels: ', len(ft))
return accuracy
# -
# ### One Vs. All
# +
'''
Deep Residual Neural Network
v2: Changed cardinality (4 -> 16), I/O Channels in Residual Blocks,
Batch Normalization in 'add_common_layers()',
and included SpatialDropout2D to 'add_common_layers()'
'''
img_height = 256
img_width = 256
img_channels = 1
#
# network params
#
cardinality = 16
def residual_network(x):
"""
ResNeXt by default. For ResNet set `cardinality` = 1 above.
"""
def add_common_layers(y):
y = layers.BatchNormalization()(y)
y = layers.ReLU()(y)
y = layers.SpatialDropout2D(0.125)(y)
return y
def grouped_convolution(y, nb_channels, _strides):
# when `cardinality` == 1 this is just a standard convolution
if cardinality == 1:
return layers.Conv2D(nb_channels, kernel_size=(3, 3), strides=_strides, padding='same')(y)
assert not nb_channels % cardinality
_d = nb_channels // cardinality
# in a grouped convolution layer, input and output channels are divided into `cardinality` groups,
# and convolutions are separately performed within each group
groups = []
for j in range(cardinality):
group = layers.Lambda(lambda z: z[:, :, :, j * _d:j * _d + _d])(y)
groups.append(layers.Conv2D(_d, kernel_size=(3, 3), strides=_strides, padding='same')(group))
# the grouped convolutional layer concatenates them as the outputs of the layer
y = layers.concatenate(groups)
return y
def residual_block(y, nb_channels_in, nb_channels_out, _strides=(1, 1), _project_shortcut=False):
"""
Our network consists of a stack of residual blocks. These blocks have the same topology,
and are subject to two simple rules:
- If producing spatial maps of the same size, the blocks share the same hyper-parameters (width and filter sizes).
- Each time the spatial map is down-sampled by a factor of 2, the width of the blocks is multiplied by a factor of 2.
"""
shortcut = y
# we modify the residual building block as a bottleneck design to make the network more economical
y = layers.Conv2D(nb_channels_in, kernel_size=(1, 1), strides=(1, 1), padding='same')(y)
y = add_common_layers(y)
# ResNeXt (identical to ResNet when `cardinality` == 1)
y = grouped_convolution(y, nb_channels_in, _strides=_strides)
y = add_common_layers(y)
y = layers.Conv2D(nb_channels_out, kernel_size=(1, 1), strides=(1, 1), padding='same')(y)
# batch normalization is employed after aggregating the transformations and before adding to the shortcut
y = layers.BatchNormalization()(y)
# identity shortcuts used directly when the input and output are of the same dimensions
if _project_shortcut or _strides != (1, 1):
# when the dimensions increase projection shortcut is used to match dimensions (done by 1×1 convolutions)
# when the shortcuts go across feature maps of two sizes, they are performed with a stride of 2
shortcut = layers.Conv2D(nb_channels_out, kernel_size=(1, 1), strides=_strides, padding='same')(shortcut)
shortcut = layers.BatchNormalization()(shortcut)
y = layers.add([shortcut, y])
# relu is performed right after each batch normalization,
# expect for the output of the block where relu is performed after the adding to the shortcut
y = layers.ReLU()(y)
return y
# conv block 1
x = layers.Conv2D(32, kernel_size=(3, 3), strides=(1, 1), padding='same')(x)
x = add_common_layers(x)
x = layers.MaxPool2D(pool_size=(2, 2), strides=None, padding='same')(x)
# residual block
for i in range(3):
project_shortcut = (i == 0)
x = residual_block(x, 16, 32, _project_shortcut=project_shortcut)
# conv block 2
x = layers.Conv2D(32, kernel_size=(3, 3), strides=(1, 1), padding='same')(x)
x = add_common_layers(x)
x = layers.MaxPool2D(pool_size=(2, 2), strides=None, padding='same')(x)
# conv block 3
x = layers.Conv2D(32, kernel_size=(3, 3), strides=(1, 1), padding='same')(x)
x = add_common_layers(x)
x = layers.MaxPool2D(pool_size=(2, 2), strides=None, padding='same')(x)
for i in range(4):
# down-sampling is performed by conv3_1, conv4_1, and conv5_1 with a stride of 2
strides = (1, 1)
x = residual_block(x, 16, 32, _strides=strides)
# conv4
for i in range(6):
strides = (1, 1)
x = residual_block(x, 32, 64, _strides=strides)
# conv5
for i in range(3):
strides = (2, 2) if i == 0 else (1, 1)
x = residual_block(x, 64, 128, _strides=strides)
x = layers.GlobalAveragePooling2D()(x)
x = layers.Dense(16)(x)
x = layers.Dense(1)(x)
return x
image_tensor = layers.Input(shape=(img_height, img_width, img_channels))
network_output = residual_network(image_tensor)
model = models.Model(inputs=[image_tensor], outputs=[network_output])
print(model.summary())
# +
# Last model: RNN_Test
#model = load_model()
#model.summary()
model.compile(optimizer = optimizers.RMSprop(lr = 1e-4),
loss = 'binary_crossentropy',
metrics = ['accuracy'])
custom_metrics = MultiLabel_Acc((val_img, val_labels))
model_obj = model.fit(training_img, training_labels,
epochs = 50, initial_epoch = 25,
validation_data = (val_img, val_labels),
batch_size = 64, verbose = 1,
callbacks = [custom_metrics])
Predictions = model.predict(test_img)
Accuracy = _1vAll_accuracy(test_labels, Predictions)
history_str = 'ResNet_50e_history'
model_str = 'ResNet_50e'
save_model(model_obj, history_str, model_str)
acc = model_obj.history['acc']
val_acc = model_obj.history['val_acc']
loss = model_obj.history['loss']
val_loss = model_obj.history['val_loss']
epochs = range(1, len(acc) + 1)
plt.figure(figsize=(15, 10))
plt.plot(epochs, acc, 'bd', label='Training Accuracy')
plt.plot(epochs, val_acc, 'b', label='Validation Accuracy')
plt.title('Training and Validation Accuracy', fontsize='xx-large')
plt.xticks(fontsize='xx-large')
plt.xlabel('Epochs', fontsize='xx-large')
plt.yticks(fontsize='xx-large')
plt.ylabel('Accuracy', fontsize='xx-large')
plt.legend(fontsize='xx-large')
plt.show()
plt.figure(figsize=(15, 10))
plt.plot(epochs, loss, 'rd', label='Training loss')
plt.plot(epochs, val_loss, 'r', label='Validation loss')
plt.title('Training and Validation Loss', fontsize='xx-large')
plt.xticks(fontsize='xx-large')
plt.xlabel('Epochs', fontsize='xx-large')
plt.yticks(fontsize='xx-large')
plt.ylabel('Loss', fontsize='xx-large')
plt.legend(fontsize='xx-large')
plt.show()
# Plot Confusion Matrix
cm = confusion_matrix(test_labels, np.round_(Predictions))
cm_plot_labels = ['Not Infiltration', 'Infiltration']
plt.clf()
fig = plt.figure(figsize = (20, 10))
plot_confusion_matrix(cm, cm_plot_labels, title = model_str, normalize=True)
#save_plt(fig)
# -
rnn_test_history = load_history()
print(len(rnn_test_history['acc']))
| Neural_Network_Architectures.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# # R DESeq2 패키지 python으로 포팅
# > "rpy2를 활용하여 DESeq2 패키지 파이썬에서 사용하기"
#
# - toc: false
# - branch: master
# - badges: true
# - comments: true
# - author: 정진우
# - categories: [R]
# - image: images/2022-02-18-DESeq-example.png
# -
# DESeq2을 파이썬의 rpy2 라이브러리를 통해 포팅하는 방법은 [여기]("https://github.com/wckdouglas/diffexpr/blob/master/example/deseq_example.ipynb")를 참고하였다.
# ### Dependencies
# 1. pandas
# 2. rpy2
# 3. tzlocal
# 4. biopython
# 5. ReportLab
# 6. pytest-cov
# 7. bioconductor-deseq2
# 8. codecov
# Install Guide
# ```python
# conda config --add channels defaults
# conda config --add channels bioconda
# conda config --add channels conda-forge
# conda create -q -n diffexpr python=3.6 \
# pandas tzlocal rpy2 biopython ReportLab pytest-cov \
# bioconductor-deseq2 codecov
# conda activate diffexpr # activate diffexpr environment
# Rscript setup.R #to install DESeq2 correctly
# python setup.py install
# ```
# > Note: 여기에서는 [MAQC 데이터의 샘플 A와 B](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3272078/)에서 [ERCC transcript](https://www.thermofisher.com/order/catalog/product/4456740)의 일부 예제를 사용하였다. 아래 실습에서 사용된 파일들은 [여기](https://github.com/wckdouglas/diffexpr/tree/master/test)에서 확인하실 수 있다.
# 필요한 패키지 로드...
# + tags=[]
# %load_ext autoreload
# %autoreload 2
import pandas as pd
import numpy as np
# + tags=[]
df = pd.read_table('ercc.txt')
df.head()
# -
# 그리고 여기에서는 유전자 발현의 정량화된 값이 포함된 테이블(count table)의 샘플을 기반으로 design matrix를 생성한다.
# 참고로, 샘플 이름은 pd.DataFrame의 인덱스로 사용해야 한다.
# + tags=[]
sample_df = pd.DataFrame({'samplename': df.columns}) \
.query('samplename != "id"')\
.assign(sample = lambda d: d.samplename.str.extract('([AB])_', expand=False)) \
.assign(replicate = lambda d: d.samplename.str.extract('_([123])', expand=False))
sample_df.index = sample_df.samplename
sample_df
# -
# DESeq2 패키지가 R에서 실행되는 방식과 유사하지만 count table의 유전자 ID인 `row.name` 대신에 어떤 열이 유전자 ID인지 알려야 한다.
# + tags=[]
from py_deseq import py_DESeq2
dds = py_DESeq2(count_matrix = df,
design_matrix = sample_df,
design_formula = '~ replicate + sample',
gene_column = 'id') # <- telling DESeq2 this should be the gene ID column
dds.run_deseq()
dds.get_deseq_result(contrast = ['sample','B','A'])
res = dds.deseq_result
res.head()
# + tags=[]
dds.normalized_count() #DESeq2 normalized count
# -
dds.comparison # show coefficients for GLM
# + tags=[]
# from the last cell, we see the arrangement of coefficients,
# so that we can now use "coef" for lfcShrink
# the comparison we want to focus on is 'sample_B_vs_A', so coef = 4 will be used
lfc_res = dds.lfcShrink(coef=4, method='apeglm')
lfc_res.head()
| _notebooks/2022-02-18-DESeq-example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="fk0evCZ7U9WO"
# # **Natural Language Processing**
# # **<NAME>**
#
# + [markdown] colab_type="text" id="Il_b_LFKXi8t"
# # NLTK Library
# + [markdown] colab_type="text" id="52_aJRSqaHgC"
# # Tokenizing words and Sentences using Nltk
# + [markdown] colab_type="text" id="o5_ElYaeaMbR"
# **Tokenization** is the process by which big quantity of text is divided into smaller parts called tokens. <br>It is crucial to understand the pattern in the text in order to perform various NLP tasks.These tokens are very useful for finding such patterns.<br>
#
# Natural Language toolkit has very important module tokenize which further comprises of sub-modules
#
# 1. word tokenize
# 2. sentence tokenize
# + colab={} colab_type="code" id="sby_OS3qZ_fz"
# Importing modules
import nltk
nltk.download('punkt') # For tokenizers
nltk.download('inaugural') # For dataset
from nltk.tokenize import word_tokenize,sent_tokenize
# + colab={} colab_type="code" id="ew9Aq5WHXSn-"
# Sample corpus.
from nltk.corpus import inaugural
corpus = inaugural.raw('1789-Washington.txt')
print(corpus)
# + [markdown] colab_type="text" id="V2wbXKzVW0GO"
# For the given corpus,
# 1. Print the number of sentences and tokens.
# 2. Print the average number of tokens per sentence.
# 3. Print the number of unique tokens
# 4. Print the number of tokens after stopword removal using the stopwords from nltk.
#
# + colab={} colab_type="code" id="jrtu9HcHXFe6"
sents = nltk.sent_tokenize(corpus)
print("The number of sentences is", len(sents)) #prints the number of sentences
words = nltk.word_tokenize(corpus)
print("The number of tokens is", len(words)) #prints the number of tokens
average_tokens = round(len(words)/len(sents))
print("The average number of tokens per sentence is", average_tokens) #prints the average number of tokens per sentence
unique_tokens = set(words)
print("The number of unique tokens are", len(unique_tokens)) #prints the number of unique tokens
from nltk.corpus import stopwords
stop_words = set(stopwords.words('english'))
final_tokens = []
for each in words:
if each not in stop_words:
final_tokens.append(each)
print("The number of total tokens after removing stopwords are", len((final_tokens))) #prints number of tokens after removing stopwords
# + [markdown] colab_type="text" id="UViYY9_3t2UE"
# # Stemming and Lemmatization with NLTK
# + [markdown] colab_type="text" id="g55XX9KDLgO7"
# **What is Stemming?** <br>
# Stemming is a kind of normalization for words. Normalization is a technique where a set of words in a sentence are converted into a sequence to shorten its lookup. The words which have the same meaning but have some variation according to the context or sentence are normalized.<br>
# Hence Stemming is a way to find the root word from any variations of respective word
#
# There are many stemmers provided by Nltk like **PorterStemmer**, **SnowballStemmer**, **LancasterStemmer**.<br>
#
# We will see differences between Porterstemmer and Snowballstemmer
# + colab={} colab_type="code" id="SS4Ij__XLfTB"
from nltk.stem import PorterStemmer
from nltk.stem import SnowballStemmer # Note that SnowballStemmer has language as parameter.
words = ["grows","leaves","fairly","cats","trouble","misunderstanding","friendships","easily", "rational", "relational"]
#Create instances of both stemmers, and stem the words using them.
stemmer_ps = PorterStemmer() #an instance of porter stemmer
stemmed_words_ps = [stemmer_ps.stem(word) for word in words]
print("Porter stemmed words: ", stemmed_words_ps)
stemmer_ss = SnowballStemmer("english") #an isntance of snowball stemmer
stemmed_words_ss = [stemmer_ss.stem(word) for word in words]
print("Snowball stemmed words: ", stemmed_words_ss)
# A function which takes a sentence/corpus and gets its stemmed version.
def stemSentence(sentence):
token_words=word_tokenize(sentence) #we need to tokenize the sentence or else stemming will return the entire sentence as is.
stem_sentence=[]
for word in token_words:
stem_sentence.append(stemmer_ps.stem(word))
stem_sentence.append(" ") #adding a space so that we can join all the words at the end to form the sentence again.
return "".join(stem_sentence)
stemmed_sentence = stemSentence("The circumstances under which I now meet you will acquit me from entering into that subject further than to refer to the great constitutional charter under which you are assembled, and which, in defining your powers, designates the objects to which your attention is to be given.")
print("The Porter stemmed sentence is: ", stemmed_sentence)
# + [markdown] colab_type="text" id="0JuE8CuDQSno"
# **What is Lemmatization?** <br>
# Lemmatization is the algorithmic process of finding the lemma of a word depending on their meaning. Lemmatization usually refers to the morphological analysis of words, which aims to remove inflectional endings. It helps in returning the base or dictionary form of a word, which is known as the lemma.<br>
#
# *The NLTK Lemmatization method is based on WorldNet's built-in morph function.*
# + colab={} colab_type="code" id="noyl1YNsQp98"
#imports
from nltk.stem import WordNetLemmatizer
nltk.download('wordnet') # Since Lemmatization method is based on WorldNet's built-in morph function.
# +
words = ["grows","leaves","fairly","cats","trouble","running","friendships","easily", "was", "relational","has"]
# Create an instance of the Lemmatizer and perform Lemmatization on above words
lemmatizer = WordNetLemmatizer() #an instance of Word Net Lemmatizer
lemmatized_words = [lemmatizer.lemmatize(word) for word in words]
print("The lemmatized words: ", lemmatized_words) #prints the lemmatized words
lemmatized_words_pos = [lemmatizer.lemmatize(word, pos = "v") for word in words]
print("The lemmatized words using a POS tag: ", lemmatized_words_pos) #prints POS tagged lemmatized words
# A function which takes a sentence/corpus and gets its lemmatized version.
def lemmatizeSentence(sentence):
token_words=word_tokenize(sentence) #we need to tokenize the sentence or else lemmatizing will return the entire sentence as is.
lemma_sentence=[]
for word in token_words:
lemma_sentence.append(lemmatizer.lemmatize(word))
lemma_sentence.append(" ")
return "".join(lemma_sentence)
lemma_sentence = lemmatizeSentence("The circumstances under which I now meet you will acquit me from entering into that subject further than to refer to the great constitutional charter under which you are assembled, and which, in defining your powers, designates the objects to which your attention is to be given.")
print("The lemmatized sentence is: ", lemma_sentence)
| nlp-preprocessing/NLP Preprocessing - Stemming and Lemmatization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 64-bit
# name: python_defaultSpec_1600198066860
# ---
# # Lab Two
# ---
#
# For this lab we're going to get into logic
#
# Our Goals are:
# - Using Conditionals
# - Using Loops
# - Creating a Function
# - Using a Class
# + tags=[]
# Create an if statement
X=55
Y=2
if X>Y:
print("X is greater than Y")
# + tags=[]
# Create an if else statement
X=2
Y=55
if X>Y:
print("X is greater than Y")
else:
print("Y is greater than X")
# + tags=[]
# Create an if elif else statement
X=2
Y=55
if X>Y:
print("X is greater than Y")
elif X==Y:
print("X and y are equal")
else:
print("Y is greater than X")
# + tags=[]
# Create a for loop using range(). Go from 0 to 9. Print out each number.
for X in range(10):
print(X)
# + tags=[]
# Create a for loop iterating through this list and printing out the value.
arr = ['Blue', 'Yellow', 'Red', 'Green', 'Purple', 'Magenta', 'Lilac']
for x in arr:
print(x)
# Get the length of the list above and print it.
# + tags=[]
# Create a while loop that ends after 6 times through. Print something for each pass.
count = 0
while count <= 5:
print(count)
count = count+1
# + tags=[]
# Create a function to add 2 numbers together. Print out the number
def add_function(X,Y):
print(X+Y)
add_function(5,5)
# + tags=[]
# Create a function that tells you if a number is odd or even and print the result.
def my_function(X):
if X % 2 ==0:
print("This number is even")
else:
print("This number is odd")
my_function(101)
# + tags=[]
# Initialize an instance of the following class. Use a variable to store the object and then call the info function to print out the attributes.
class Dog(object):
def __init__(self, name, height, weight, breed):
self.name = name
self.height = height
self.weight = weight
self.breed = breed
def info(self):
print("Name:", self.name)
print("Weight:", str(self.weight) + " Pounds")
print("Height:", str(self.height) + " Inches")
print("Breed:", self.breed)
| JupyterNotebooks/Labs/Lab 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
prefix = 'ens'
midx = '56'
import pandas as pd
import numpy as np
from sklearn.metrics import f1_score, confusion_matrix
# +
new = pd.read_csv('leak2d.csv')
new = new[new.Questionable != 1]
new.reset_index(inplace=True)
print(new.head())
print(new.shape)
ens = pd.read_csv('sub/'+prefix+midx+'.csv').fillna('')
ens = ens.set_index('Id')
print(ens.head())
print(ens.shape)
# +
changed = 0
unchanged = 0
for i, item in enumerate(new['Id']):
# print(test_preds.loc[i]['Target'])
if ens.loc[item]['Predicted'] != new.loc[i]['Target']:
print(ens.loc[item]['Predicted'],' => ', new.loc[i]['Target'])
changed += 1
ens.loc[item]['Predicted'] = new.loc[i]['Target']
else:
unchanged += 1
print('changed',changed,' unchanged',unchanged)
print()
fname = 'sub/'+prefix+midx+'d.csv'
ens.to_csv(fname)
print(fname)
# -
name_label_dict = {
0: "Nucleoplasm",
1: "Nuclear membrane",
2: "Nucleoli",
3: "Nucleoli fibrillar center",
4: "Nuclear speckles",
5: "Nuclear bodies",
6: "Endoplasmic reticulum",
7: "Golgi apparatus",
8: "Peroxisomes",
9: "Endosomes",
10: "Lysosomes",
11: "Intermediate filaments",
12: "Actin filaments",
13: "Focal adhesion sites",
14: "Microtubules",
15: "Microtubule ends",
16: "Cytokinetic bridge",
17: "Mitotic spindle",
18: "Microtubule organizing center",
19: "Centrosome",
20: "Lipid droplets",
21: "Plasma membrane",
22: "Cell junctions",
23: "Mitochondria",
24: "Aggresome",
25: "Cytosol",
26: "Cytoplasmic bodies",
27: "Rods & rings"
}
LABEL_MAP = name_label_dict
np.set_printoptions(precision=3, suppress=True, linewidth=100)
# +
# compute f1 score between two submission files
def f1_sub(csv0, csv1, num_classes=28):
c0 = pd.read_csv(csv0)
c1 = pd.read_csv(csv1)
assert c0.shape == c1.shape
s0 = [s if isinstance(s,str) else '' for s in c0.Predicted]
s1 = [s if isinstance(s,str) else '' for s in c1.Predicted]
p0 = [s.split() for s in s0]
p1 = [s.split() for s in s1]
y0 = np.zeros((c0.shape[0],num_classes)).astype(int)
y1 = np.zeros((c0.shape[0],num_classes)).astype(int)
# print(p0[:5])
for i in range(c0.shape[0]):
for j in p0[i]: y0[i,int(j)] = 1
for j in p1[i]: y1[i,int(j)] = 1
# print(y0[:5])
return f1_score(y0, y1, average='macro')
# computute confusion matrices between two submission files
def f1_confusion(csv0, csv1, num_classes=28):
c0 = pd.read_csv(csv0)
c1 = pd.read_csv(csv1)
assert c0.shape == c1.shape
s0 = [s if isinstance(s,str) else '' for s in c0.Predicted]
s1 = [s if isinstance(s,str) else '' for s in c1.Predicted]
p0 = [s.split() for s in s0]
p1 = [s.split() for s in s1]
y0 = np.zeros((c0.shape[0],num_classes)).astype(int)
y1 = np.zeros((c0.shape[0],num_classes)).astype(int)
# print(p0[:5])
for i in range(c0.shape[0]):
for j in p0[i]: y0[i,int(j)] = 1
for j in p1[i]: y1[i,int(j)] = 1
# print(y0[:5])
y0avg = np.average(y0,axis=0)
y1avg = np.average(y1,axis=0)
cm = [confusion_matrix(y0[:,i], y1[:,i]) for i in range(y0.shape[1])]
fm = [f1_score(y0[:,i], y1[:,i]) for i in range(y0.shape[1])]
for i in range(y0.shape[1]):
print(LABEL_MAP[i])
print(cm[i],' %4.2f' % fm[i],' %6.4f' % y0avg[i],' %6.4f' % y1avg[i],
' %6.4f' % (y0avg[i] - y1avg[i]))
print()
# print('y0avg')
# print(y0avg)
# print('y1avg')
# print(y1avg)
# print('y0avg - y1avg')
# print(y0avg-y1avg)
print('f1 macro')
print(np.mean(fm))
return f1_score(y0, y1, average='macro')
# -
f1_sub(fname,'sub/'+prefix+midx+'.csv')
f1_sub(fname,'sub/ens36c.csv')
f1_sub(fname,'sub/ens43c.csv')
f1_sub(fname,'sub/ens45c.csv')
f1_sub(fname,'sub/ens53c.csv')
f1_sub(fname,'sub/ens53d.csv')
f1_sub(fname,'sub/ens55d.csv')
f1_sub(fname,'sub/resnet15c.csv')
f1_sub(fname,'sub/leak_brian_tommy_en_res34swa_re50xt_re101xtswa_wrn_4.8.csv')
# +
# f1_confusion(fname,'sub/'+prefix+midx+'c.csv')
# -
| wienerschnitzelgemeinschaft/src/Russ/ens_sub56d.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## ECE 289: Homework 3
# ### Part (A): Networks Visualization
# +
# import required packages
# load results from HW1 to use. Otherwise, execute code from HW1 to produce results again.
# %time
#import useful packages, all of them are important but not necessarily used in this code
#enable inline plotting in Python Notebook
import warnings
warnings.filterwarnings('ignore')
#supress warnings
# %pylab inline
import networkx as nx
import numpy as np
import matplotlib
import scipy
import time
# -
# ### Use network datasets from the Public Data Folder.
# #### Please DO NOT copy dataset in your directory (or anywhere else!) because it will certainly result in 'timeout error' while submission.
# #### Now that we are familiar with Gephi and its usage, we will explore some built in tools from Gephi to improve our visualizations further. You will need results from the HW1. Only use results calculated using networkX here. This can be done by adding your results for node degree, centrality etc. as an attribute to the
# #### In this task, we will use Filter tool in Gephi to threshold available network data, using various properties.
# #### Visualise Facebook Network, Enron Emails Network, Collaboration (Erdos) Network applying following thesholds. Make sure to have all the visualizations labelled with appropriate node labels. This is a quiet open ended question, as you have a lot of scope to make your visualizations better trying different layouts, colors etc. So, turn in the best visualization that you get in each case. You should attach an image(.png, .jpg) for each visualization here in Notebook itself. Also, make sure that it is well readable.
# #### (1) Top ~50% nodes, thrsholded by Node Degree
# #### (2) Top ~10% nodes, thrsholded by Node Degree
# #### (3) Top ~5% nodes, thrsholded by Node Degree
# #### (4) Top ~1% nodes, thrsholded by Node Degree
# #### (5) Top ~50% nodes, thrsholded by Betweeness Centrality
# #### (6) Top ~10% nodes, thrsholded by Betweeness Centrality
# #### (7) Top ~5% nodes, thrsholded by Betweeness Centrality
# #### (8) Top ~1% nodes, thrsholded by Betweeness Centrality
# <img src = "SampleFilter.jpg">
# your response images here
# # FACEBOOK DATASET - By Degree
# ### Top ~1% nodes, thrsholded by Node Degree
# 
# ### Top ~5% nodes, thrsholded by Node Degree
# 
# ### Top ~10% nodes, thrsholded by Node Degree
# 
# ### Top ~50% nodes, thrsholded by Node Degree
# 
# # Facebook - By betweeness centrality
# ### Note : 10% and 50% node filtering wasn't possible in Gephi for this dataset, hence 5% and 100% variations are provided.
# ### Top ~1% nodes, thrsholded by Betweeness Centrality
# 
# ### Top ~5% nodes, thrsholded by Betweeness Centrality
# 
# ### Top ~100% nodes, thrsholded by Betweeness Centrality
# 
# # Enron - By node degree
# ### Top ~1% nodes, thrsholded by Node Degree
# 
# ### Top ~5% nodes, thrsholded by Node Degree
# 
# ### Top ~10% nodes, thrsholded by Node Degree
# 
# ### Top ~50% nodes, thrsholded by Node Degree
# 
# # Enron - By betweeness centrality
# ### Top ~1% nodes, thrsholded by betweeness centrality
# 
# ### Top ~5% nodes, thrsholded by betweeness centrality
# 
# ### Top ~10% nodes, thrsholded by betweeness centrality
# 
# ### Top ~50% nodes, thrsholded by betweeness centrality
# 
# # Erdos - By node degree -
# ### Note : 50% node filtering wasn't possible in Gephi for this dataset, hence 100% filtered variations are provided.
# ### Top ~1% nodes, thrsholded by Node Degree
# 
# ### Top ~5% nodes, thrsholded by Node Degree
# 
# ### Top ~10% nodes, thrsholded by Node Degree
# 
# ### Top ~100% nodes, thrsholded by Node Degree
# 
# # Erdos - By betweeness centrality -
# ### Note : 10%, 50% node filtering wasn't possible in Gephi for this dataset, hence 3.5% and 100% filtered variations are provided.
# ### Top ~1% nodes, thrsholded by betweeness centrality
# 
# ### Top ~3.5% nodes, thrsholded by betweeness centrality
# 
# ### Top ~100% nodes, thrsholded by betweeness centrality
# 
# ### Part (B): Community Detection
# #### In this task, we will try to find communities in the given netwok and explore more about them. NetworkX has built in functions for community detection (http://perso.crans.org/aynaud/communities/). Along with NetworkX, we will also use igraph library in this task, for community detection purposes.
# +
#install required packages and read their documentation to get used to them.
# #!pip install community
# #!pip install igraph
# -
# #### Community detection is a very common task for almost all networks. It helps us to understand network structure in much detail.
# #### More information on community detection: https://arxiv.org/abs/0906.0612
# ### There are multiple algorithms to detect communities. One of the commonly used algorithm is Louvain method. The method is a greedy optimization method that attempts to optimize the "modularity" of a partition of the network. 'community' library uses Lovain algorithm, and hence we get partitions based on optimized modularity. Implement a python code using 'community' library to find communities in the Citation network and Collaboration Network (Erdos). Write your code in the next cell and visualize your community detection results in Gephi for both the networks. Label the nodes in the visualization properly. Use largest connected components, if required. Include image(.jpg, .png) of the visualization here.
import community
#your code here
# # CitNet
# %%time
# load the network
file_name = "../ece289_public/Cit-HepTh.txt"
# it's a directed graph, so we should use nx.DiGraph to read
g1 = nx.read_edgelist(file_name, create_using=nx.DiGraph(), nodetype=int)
print nx.info(g1)
g2=g1.to_undirected()
partition_citnet = community.best_partition(g2)
g2=g1.to_undirected()
deg = g2.degree()
for k in g2.nodes():
g2.node[k]['comm'] = partition_citnet[k]
g2.node[k]['deg'] = deg[k]
# +
#import tarfile
#tarfile.open("../ECE289_HW_1/data/citNet/cit-HepTh-abstracts.tar.gz").extractall("../ECE289_HW_1/data/citNet/cit-HepTh-abstracts/")
# -
# extract the author name
def get_authors(l):
authors = reduce(list.__add__, [a.split(",") for a in l[9:].split("and")])
return [x.strip() for x in authors]
# attach the attribute author
#here G is your networkX graph
import os
for subdir, dirs, files in os.walk("../ECE289_HW_1/data/citNet/cit-HepTh-abstracts"):
for fl in files:
filepath = subdir + os.sep + fl
if filepath.endswith(".abs"):
node_num = int(fl[:-4])
name = ""
for l in open(filepath):
if l.startswith("Authors:"):
name = get_authors(l)[0]
if node_num in g2.nodes():
g2.node[node_num]['author'] = name
g2.node[9701151]
nx.write_gml(g2, "citNet_community.gml")
# # Erdos
# +
# build Collaboration Network
# undirected network
g1e = nx.Graph()
# add <NAME> into our network at first
dict_authors = {}
dict_authors['<NAME>'] = 0
g1e.add_node(0)
g1e.node[0]['author'] = '<NAME>'
# add the authors with Erdos number 1 and 2 from file
line_count = 1
skip_line = 24
skip_space = 1
is_new = False
author = ""
coauthor = ""
index = 1
ind_author = 1
ind_coauthor = 1
def parseLine(l, start):
end = start
while end < len(l) - 1 and not (l[end] == ' ' and l[end + 1] == ' '):
end += 1
return l[start:end]
def addAuthor(auth, ind):
if auth in dict_authors:
return ind
dict_authors[auth] = ind
return ind + 1
for l in open("../ece289_public/Erdos.html"):
if line_count >= skip_line:
if l == '\n':
is_new = True
elif is_new:
author = parseLine(l, 0)
index = addAuthor(author, index)
ind_author = dict_authors[author]
g1e.add_edge(0, ind_author)
g1e.node[ind_author]['author'] = author
is_new = False
elif l == '</pre>':
break
else:
coauthor = parseLine(l, skip_space)
index = addAuthor(coauthor, index)
ind_coauthor = dict_authors[coauthor]
g1e.add_edge(ind_author, ind_coauthor)
g1e.node[ind_coauthor]['author'] = coauthor
line_count += 1
print nx.info(g1e)
# -
deg=g1e.degree()
partition_erdos = community.best_partition(g1e)
for k in g1e.nodes():
g1e.node[k]['deg'] = deg[k]
g1e.node[k]['comm'] = partition_erdos[k]
nx.write_gml(g1e, "erdos_community.gml")
# +
# visualizations
# -
# # Citation network
# ## View 1
# 
# ## View 2
# 
# # Erdos network
# ## View 1
# 
# ## View 2
# 
# ### Compared to 'community' library, 'igraph' has more flexibilty to detect communities. igraph allows user to partition the network in the number of communities that user wishes. Obviously this number is bounded. Now, you will use this aspect to divide given network in '5' communities using 'igraph' and observe the results. Also, derive results for optimized modularity condition in igraph. Write a python code to implement above task for citation network & collaboration network (Erdos). Remember that unlike 'community', igraph has multiple approach for community detection. Obvious approach being greedy and it optimizes modularity. Visualize your community detection results in Gephi for both the networks. Label the nodes in the visualization properly. Use largest connected components, if required. Use different colors for nodes in every community. Include image(.jpg, .png) of the visualization here.
# +
from igraph import *
#your code here
#partition network using greedy approach. Note the number of communities
#partition network in 5 communities and see the difference in the visualization
# -
# # Erdos
gi1 = Graph.Read_GML('erdos_community.gml')
igraph.summary(gi1)
com_1 = gi1.simplify().community_fastgreedy().as_clustering()
mem = com_1.membership
for i in range(0,len(mem)):
gi1.vs[i]['igraph_com'] = mem[i]
print 'The numbers of clusters by optimizing modularity is for Erdos n/w is',len(com_1.subgraphs())
gi1.write_gml('gi1_erdos.gml')
# ## Optimized modularity - clusters
# 
# # Erdos - 5 clusters
com_e5 = gi1.simplify().community_fastgreedy().as_clustering(n=5)
mem = com_e5.membership
for i in range(0,len(mem)):
gi1.vs[i]['igraph_com'] = mem[i]
gi1.write_gml('gi1_clus5_erdos.gml')
# ## 5 clusters
# 
# # Citation
gic2 = Graph.Read_GML('citNet_community.gml')
com_c = gic2.simplify().community_fastgreedy().as_clustering()
mem = com_c.membership
#len(mem)
#len(gi1.degree())
#gi2.vs[0]
for i in range(0,len(mem)):
gic2.vs[i]['igraph_com'] = mem[i]
#gic2.vs[1000]
#summary(gic2)
print 'The numbers of clusters by optimizing modularity is for Citation n/w is',len(com_c.subgraphs())
gic2.write_gml('gic2_citnet.gml')
# ## Optimized modularity - clusters
# 
# # CitNet - 5 clusters
com_c = gic2.simplify().community_fastgreedy()
#mem = com_c.membership
#len(mem)
#len(gi1.degree())
#gi2.vs[0]
#for i in range(0,len(mem)):
# gic2.vs[i]['igraph_com'] = mem[i]
# ### The clustering for 5 clusters doesn't work. Error due to small merge matrices.
# ### So, we take larger clusters and trim it to 5.
# +
#comc1 = com_c.as_clustering(n=5)
# -
comc1 = com_c.as_clustering(n=150)
for i,c in enumerate(comc1):
if i<6:
for k in c:
gic2.vs[k]['igraph_com'] = i
gic2.write_gml('gic2_clus5_citnet.gml')
# ## 5 clusters
# 
# # visualizations
# ### Now that we have detected communities, we will further analyze our results. This task is only for Collaboration network (Erdos). Use results from communtiy detection using 'community'. Sort communities and get largest 5 communities. For each of these 5 communities, get 3 nodes with the highest node degree. So you will get 3 authors per community, for 5 communities. Now search the area of research for each of these authors and enlist them. Further, observe if there is any reason for those 3 authors to be in same community, for each community. State that reason in brief. Write all of your results in next cell. Also include any other interesting results that you may observe during process.
#your observations here
# 'g1e' contains the erdos graph as a networx graph
l = len(g1e.node)
c = []
for i in range(0,l):
c.append(g1e.node[i]['comm'])
print 'The top 5 communities in erdos graph are :'
print 'Community (Count)'
coms = []
for count, elem in sorted(((c.count(e), e) for e in set(c)), reverse=True)[:5]:
print '%s \t (%d)' % (elem, count)
coms.append(elem)
listofnodes = list([i for i in g1e.__iter__()])
for ci in coms:
nodes = list(filter(lambda k : g1e.node[k]['comm'] == ci,listofnodes))
lisofdeg = sorted([(g1e.node[i]['deg'],i) for i in nodes], reverse=True)[:3]
nodelists = [g1e.node[i[1]] for i in lisofdeg]
print 'The top 3 authors in the community ',ci,'are : '
for i in nodelists:
print '\t Author : %s with degree : %d' % (i['author'],i['deg'])
print '\n'
# ##### 1. Interesting observation : Nearly most of the authors have some relation with Erdos graph theory, and have either wrote papers with him or have proved his conjectures etc, which makes sense since it is a Erdos dataset we have.
# ##### 2. Also many people do publish in combinatrics, which is visible across clusters, and since they have max. degree; it means that a lot of papers are published in combinatorics, or atleast by the people associated with Erdos.
# # Community 11
#
# <NAME> : <NAME> was an American mathematician, who specialized in graph theory. He was widely recognized as one of the "fathers" of modern graph theory.
#
# <NAME> : <NAME> "Ron" Graham (born October 31, 1935) is a mathematician credited by the American Mathematical Society as being "one of the principal architects of the rapid development worldwide of discrete mathematics in recent years". He has done important work in scheduling theory, computational geometry, Ramsey theory, and quasi-randomness.
#
# <NAME> : <NAME> (Chinese: 金芳蓉; pinyin: Jīn Fāngróng; born October 9, 1949), known professionally as <NAME>, is a mathematician who works mainly in the areas of spectral graph theory, extremal graph theory and random graphs, in particular in generalizing the Erdős–Rényi model for graphs with general degree distribution (including power-law graphs in the study of large information networks).
#
# #### As we can see, all the 3 authors in the above community have worked in GRAPH THEORY, especially in discrete math.
# # Community 5
#
# <NAME> : <NAME> (born 16 March 1969 in Galați) is a Romanian mathematician who specializes in number theory with emphasis on Diophantine equations, linear recurrences and the distribution of values of arithmetic functions. He has made notable contributions to the proof that irrational automatic numbers are transcendental and the proof of a conjecture of Erdős on the intersection of the Euler function and the sum of divisors function.
#
# SHALLIT, <NAME> : <NAME> (born October 17, 1957) is a computer scientist, number theorist, a noted advocate for civil liberties on the Internet, and a noted critic of intelligent design. He is married to <NAME>, also a computer scientist.
#
# POMERANCE, <NAME> : <NAME> (born in 1944 in Joplin, Missouri) is an American number theorist. He attended college at Brown University and later received his Ph.D. from Harvard University in 1972 with a dissertation proving that any odd perfect number has at least seven distinct prime factors.[1] He immediately joined the faculty at the University of Georgia, becoming full professor in 1982. He subsequently worked at Lucent Technologies for a number of years, and then became a distinguished Professor at Dartmouth College.
#
# #### As we can see, all the 3 authors are working in number theory.
# # Community 14
#
# ALON, <NAME>. : Combinatorics, Graph Theory and their applications to Theoretical Computer Science. Combinatorial algorithms and circuit complexity. Combinatorial geometry and Combinatorial number theory. Algebraic and probabilistic methods in Combinatorics.
#
# <NAME> : <NAME> is a Hungarian mathematician, best known for his work in combinatorics, for which he was awarded the Wolf Prize and the Knuth Prize in 1999, and the Kyoto Prize in 2010. He is the current president of the Hungarian Academy of Sciences.
#
# SAKS, <NAME> : Saks research in computational complexity theory, combinatorics, and graph theory has contributed to the study of lower bounds in order theory, randomized computation, and space-time tradeoff.
#
# #### As we can see, all the 3 authors are working in combinatrics, graph theory and theortical computer science.
# # Community 0
#
# <NAME> : Erdős pursued and proposed problems in discrete mathematics, graph theory, number theory, mathematical analysis, approximation theory, set theory, and probability theory.
#
# STRAUS, ERNST GABOR* : Ernst Gabor Straus (February 25, 1922 – July 12, 1983) was a German-American mathematician of Jewish origin who helped found the theories of Euclidean Ramsey theory and of the arithmetic properties of analytic functions. His extensive list of co-authors includes <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, and <NAME>.
#
# CHOWLA, <NAME>.* : <NAME> (22 October 1907 – 10 December 1995) was a British-born Indian American mathematician, specializing in number theory. Among his contributions are a number of results which bear his name. These include the Bruck–Ryser–Chowla theorem, the Ankeny–Artin–Chowla congruence, the Chowla–Mordell theorem, and the Chowla–Selberg formula, and the Mian–Chowla sequence.
#
# #### As we can see, we have the main man, Dr. Erdos in this community, and he shares the community with other erdos'ists, who have worked with him in number theory, ramsey theory and discrete math.
# # Community 17 :
#
# TUZA, ZSOLT : Upenn guy. Graph theory, combinatrics, hypergraphs etc
#
# PACH, JANOS : <NAME> (born May 3, 1954)[2] is a mathematician and computer scientist working in the fields of combinatorics and discrete and computational geometry.
#
# ARONOV, BORIS : <NAME> is a computer scientist, currently a professor at the Tandon School of Engineering, New York University. His main area of research is computational geometry. He is a Sloan Research Fellow.
#
# #### As we can see we have computer scientists in the field of combinatorics in this cluster.
| ECE 289/ECE289_HW3/HW3Visualizations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # 1.4 Finite Difference Method - Simple Room
# Prepared by (C) <NAME>
# Let's step it up a little. Let's model a simple room with a window using 3 thermal nodes: the window, the room air and a node representing all the interior walls combined. For a more detailed analysis, the walls can be separated and the floor can be further discretized.
# ## Problem Statement
# A small room with a 9 m<sup>2</sup> window is to be simulated. In fact, the whole wall is a window! (Ridiculous!)
# All the interior surfaces of the room are lumped together and behave like a 100 mm thick 9 m<sup>2</sup> concrete slab. The air is assumed to have no thermal capacitance. The material properties are given in the script.
#
# The exterior temperature follows a sine with mean -15°C, amplitude 5°C peaking at 15:00.
#
# Just for fun, 500 W of heat is supplied as a pulse from 6:00 to 12:00.
# ------------
# ## FDM Implementation
# ### Load Dependencies
import numpy as np
import matplotlib.pylab as plt
import matplotlib as mpl
# Plot in notebook; comment out the line below for windowed plots
# %matplotlib inline
# mpl.rc('figure', figsize=(10, 10))
# import seaborn; seaborn.set() # Optional package: makes prettier plots
# ### Variable Names
# Uin: Conductance matrix input by user, upper triangle only, (nN x nN) (W/K)
# U: Conductance matrix (symmetrical) with added capacitance for diagonal term, (nN x nN) (W/K)
# C: Capacitance vector, (nN x 1) (J/K)
# F: Conductance matrix of nodes connected to a known temperature source, (nN x nM) (W/K)
# T: Temperature vector per timestep, (nT x nN) (degC)
# TK: Temperature vector of known temperatures per timestep, (nT x nM) (degC)
# Qin: Heat flow, only external sources, (nN x 1) (W)
# Q: Heat flow vector + external sources + capacitance from previous timestep (implicit only), (nN x 1) (W)
#
# nN: Number of nodes
# nM: Number of nodes with known temperatures
# nT: Number of timesteps
#
# Node Number: Object
# 0: window's interior surface, 9 m^2
# 1: room air
# 2: mass, 100 mm concrete, 9 m^2
# *Note that indexing starts at "0" in Python (unlike "1" in Matlab)
#
# Node Number with known temperatures: Object
# 0: exterior air
# ### Geometry and other Properties
Ac = 9. # m^2, concrete floor
Aw = 9. # m^2, window
A = [Aw, 0, Ac] # m^2, area vector
Uw = 4.03 # W/m^2K (ASHRAE Fund 2009 Ch 15.8 Table 4: Double glazed,
# 12.7 mm airspace, curtainwall, aluminum w/o thermal break)
h_out = 34. # W/m^2K
h_int = 8. # W/m^2K
k = 1.731 # W/(m*K)
rho = 2240. # kg/m^3
Cp = 840. # J/(kg*K) where J = kgm^2/s^2
dx = 0.10 # m
# ### Simulation Parameters
nN = 3 # number of nodes
nM = 1 # number of nodes with known temperatures
st = 24 # steps per hour
H = 24 # hr; number of hours simulated
nt = int(st*H) # number of timesteps-1
dt = 3600/st # s (3600 sec = 1 hour)
days = 10 # number of days simulated
# ### Known (Set) Temperatures & Nodal Connections
# +
# Declare variables
T = np.zeros((nt*days,nN)) # degC
TK = np.zeros((nt*days,nM)) # degC
Uin = np.zeros((nN,nN)) # K/W
F = np.zeros((nN,nM)) # K/W
C = np.zeros((nN,1)) # J/K
# Known temperature sources
T_out_m = -15. # degC, Exterior temperature
dT_out = 10. # degC, Temperature differential
T_out_theta = -5*np.pi/4 # Shift
w = 2*np.pi/86400 # Period
T_out = days*[T_out_m + dT_out/2*np.cos(w*t*dt + T_out_theta) for t in range(nt)]
TK[:,0] = T_out # degC; Exterior temperature
# How are the nodes connected?
Uin[0,1] = (1/(h_int*A[0]))**-1 # window surface to room air node
Uin[1,2] = (1/(h_int*A[2]) + dx/(2*k*A[2]))**-1 # room air to center mass
# Connected to temperature sources
# window's inside surface (node 0) is connected to the outdoor air node through
# the window's outside surface (combined conductances; series connection)
F[0,0] = (1/(h_out*A[0]) + 1/(Uw*A[0]))**-1
# Nodes with capacitance
# only node 2 has capacitance (concrete floor); no air capacitance
C[2] = rho*Cp*dx*A[2]
# -
# Plot known temperature sources
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
plotTK = TK[(days-1)*nt:(days)*nt:]
plt.plot(np.dot(dt/3600.,range(0,nt)),plotTK)
plt.xlim([0,24])
ax.set_xticks(np.arange(0,25,3))
plt.grid()
#plt.ylim([-20,-5])
labels = ['TK' + str(n) for n in range(nM)]
plt.legend(labels,loc='best', fontsize='medium')
plt.xlabel('Time, h')
plt.ylabel('Temperature, degC')
plt.title('Known (Set) Temperature Sources')
# #### Heat Sources
# +
# Assuming there's a pulse of 500 W of heat for 6 hours a day
q_in = np.concatenate( (np.zeros(nt*1/4), 500*np.ones(nt/4), np.zeros(nt*2/4)) , axis=0)
# Plot
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
plt.plot(np.dot(dt/3600.,range(0,nt)),q_in,'r')
plt.xlim([0,24])
ax.set_xticks(np.arange(0,25,3))
plt.ylim([0,600])
plt.grid()
#plt.legend(['Heating Input'], loc='best', fontsize='medium')
plt.xlabel('Time, h')
plt.ylabel('Power, W')
plt.title('Heating Input')
# -
# #### Initial Conditions
T[0,] = 10. # all temperatures set to 10 degC
# ------------
# ## Run Simulation (Implicit Scheme)
# Ideally, we should be able to reuse this part of the code applying only minimal changes. For cases where materials properties change over time or due to temperature, or when controls are present, the U-matrix completion should be done within the Main Loop.
# ### U-matrix completion
# Here, we complete the connections due to symmetry (?):
# Node 0 is connected to node 1; also, node 1 is connected to node 0.
U = -Uin - Uin.T # U is symmetrical, non-diagonals are -ve
s = -np.sum(U,1)
for i in range(0,nN):
U[i,i] = s[i] + np.sum(F[i,]) + C[i]/dt
del s
# ### Main Loop
# Loop for number of days, and for all timesteps in a day
for d in range(0,days):
for t in range(0,nt):
# Heat flow into the node, external sources
Qin = np.zeros((nN,1))
Qin[1] = q_in[t]
# Q-vector: Q = Qin + F*TM(t) + C/dt*T(t)
Q = Qin + np.reshape(np.dot(F, TK[d*nt+t,]),(nN,1)) + \
np.multiply(C/dt, np.reshape(T[d*nt+t,:],(nN,1)))
# Compute temperature
# This is where the magic happens. For a system where nothing changes
# (no controls, no moveable blinds, no PCM; we need only define the
# U-matrix once (done above) and use it here)
if (days*nt) > (d*nt+t+1):
T[d*nt+t+1,] = np.linalg.solve(U,Q).T
# ## Plotting
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
plt.hold(True)
plotT = T[(days-1)*nt:(days)*nt:]
plt.plot(np.dot(dt/3600.,range(0,nt)),plotT)
plt.xlim([0,24])
ax.set_xticks(np.arange(0,25,3))
plt.legend(['Window Inside Surface','Room Air','Concrete Floor'],loc='best')
plt.xlabel('Time, h')
plt.ylabel('Temperature, degC')
plt.title('Temperature Distribution on Day %s' %(days))
plt.grid()
plt.show()
# ------------
# ## Run Simulation (Explicit Scheme)
# +
# Declare variables
Te = np.zeros((nt*days,nN)) # degC, temperatures for explicit mode
Te[0,] = T[0,] # initial condition set to be the same as implicit case
# RHS matrix
Ue = Uin + Uin.T
s = -np.sum(Ue,1)
for i in range(0,nN):
Ue[i,i] = s[i] - np.sum(F[i,]) + C[i]/dt
del i, s
Und = Uin + Uin.T # Without the diagonals
# -
# ### Main Loop
# Loop for number of days, and for all timesteps in a day
for d in range(0,days):
for t in range(0,nt):
# Heat flow into the node
Qin = np.zeros((nN,1))
Qin[1] = q_in[t]
# Q-vector (explicit): Q = Qin + F*TM(t)
Q = Qin + np.reshape(np.dot(F, TK[d*nt+t,]),(nN,1))
# Compute temperature
# This is where the magic happens. Prepare to be amazed!
if (days*nt) > (d*nt+t+1):
for c in range(nN):
# Nodes with capacitance (diffusion nodes)
if (C[c] != 0.):
Te[d*nt+t+1,c] = dt/C[c] * (np.dot(Ue[c,],Te[d*nt+t,]) + Q[c])
# Nodes without capacitance (arithmetic nodes)
else:
Te[d*nt+t+1,c] = (np.dot(Und[c,],Te[d*nt+t,]) + np.dot(F[c,],TK[d*nt+t,]) + Qin[c]) \
/ (np.sum(Und[c,]) + np.sum(F[c,]))
# ### Plotting
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
plotTe = Te[(days-1)*nt:(days)*nt:]
plt.plot(np.dot(dt/3600.,range(0,nt)),plotTe)
plt.xlim([0,24])
ax.set_xticks(np.arange(0,25,3))
plt.legend(['Window Inside Surface','Room Air','Concrete Floor'],loc='best')
plt.xlabel('Time, h')
plt.ylabel('Temperature, degC')
plt.title('Temperature Distribution on Day %s' %(days))
plt.grid()
plt.show()
# ----------
# ## Comparative Plot
# Let's compare the Implicit and Explicit schemes. Play with the "steps per hour" term *st* and see its effect on the temperature profiles.
# +
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
plt.plot(np.dot(dt/3600.,range(0,nt)),plotT)
plt.gca().set_prop_cycle(None) # restart colouring order
plt.plot(np.dot(dt/3600.,range(0,nt)),plotTe,'--')
plt.xlim([0,24])
ax.set_xticks(np.arange(0,25,3))
plt.legend(['Window','Room','Concrete'],loc=1)
plt.xlabel('Time, h')
plt.ylabel('Temperature, degC')
plt.title('Comparative (Dashed is Explicit)')
plt.grid()
plt.show()
print "Chosen timestep: %.2f minutes" % (60./st)
| 1.4 Finite Difference Method - Simple Room.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # データ分析入門編:scikit-learnで交差検証!
# 以前作成したBoston House-Priceデータセットで作成したSVRのモデルを交差検証を使ってチューニングします。
# まずはデータをロードします。
import sklearn
from sklearn import datasets
boston = datasets.load_boston()
# SVRは事前にデータを正規化する必要がありますので、StandardScalr()関数を使って正規化を行います。
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(boston.data)
data = scaler.transform(boston.data)
# それでは、まずはチューニング前のデフォルトの設定の精度を見ましょう。精度の司法としては平均二乗誤差を使います。
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(data, boston.target, test_size=0.4, random_state=0)
from sklearn.svm import SVR
clf = SVR()
clf.fit(X_train, y_train)
from sklearn.metrics import mean_squared_error
mean_squared_error(y_test, clf.predict(X_test))
# 結果、平均二乗誤差は34となりました。これが今回のベンチマークとなります。これよりも少ない誤差を目指しましょう。
# 次に、交差検証とグリッドサーチを使って最適なハイパーパラメーターを探します。今回対象とするハイパーパラメーターは"C"です。デフォルトの値は1.0ですが、それを変えてみましょう。
from sklearn.model_selection import GridSearchCV
parameters = {'C':[0.001, 0.01, 0.1, 1.0, 10, 100, 1000, 10000]}
clf = GridSearchCV(SVR(), parameters, cv=10, scoring='neg_mean_squared_error')
clf.fit(data, boston.target)
# 一番良かったパラメーターはclf.best_estimator_で確認できます。
clf.best_estimator_
# 上記の通り、一番よいCの値は100となりました。では、実際にC=100と設定して、精度がどれほど変わるか見てみましょう。
from sklearn.svm import SVR
clf = SVR(C=100)
clf.fit(X_train, y_train)
from sklearn.metrics import mean_squared_error
mean_squared_error(y_test, clf.predict(X_test))
# 平均二乗誤差が34から14に減り、大分精度があがったことが分かります。
| Cross Validation with scikit-learn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="4v0ZWtfLkub3" outputId="8283df6d-0873-4596-ceaa-44d86d2291b5"
# # !pip install -Uqq fastai
# # !pip install voila
# # !jupyter serverextension enable voila —sys-prefix
# + colab={"base_uri": "https://localhost:8080/"} id="9bct3xnMkwC-" outputId="bdcd95c2-1775-43a0-eba7-9cee1b17ac2b"
from fastai.vision.all import *
from fastai.vision.widgets import *
import zipfile
# + id="Ty-p4BfI7CCn"
# Create a dummy label_func so that the trained model can be loaded without needing
def label_func():
return 0
# -
if not Path(Path.cwd()/'export.pkl').exists():
with zipfile.ZipFile('export.zip') as z:
z.extractall(Path.cwd())
# + id="3mNib2492ZYj"
# Load the model
learn = load_learner('export.pkl')
# + id="HwlNbzz23kGr"
# Create a button for uploading test images
btn_upload = widgets.FileUpload()
# + id="FDq-2iNI5Br6"
# Create an Output widget for displaying the loaded image
out_pl = widgets.Output()
# + id="Kro4_W805XHl"
# Create a Label widget for outputting the results of the classifier
lbl_pred = widgets.Label()
# + id="upfTNspM5kiZ"
# Creating a button used to run the classification
btn_run = widgets.Button(description='Classify')
# Defining a click event handler
def on_click_handler(o):
img = PILImage.create(btn_upload.data[-1])
out_pl.clear_output()
with out_pl: display(img.to_thumb(128,128))
pred,pred_idx,probs = learn.predict(img)
lbl_pred.value = f'Prediction: {pred}; Probability: {probs[pred_idx]:.04f}'
btn_run.on_click(on_click_handler)
# + colab={"base_uri": "https://localhost:8080/", "height": 238, "referenced_widgets": ["e487b5e5617444a7b93f5ae5cd533d6a", "442038ae159d4abf860ea3a034ba376b", "064e092ab3fd4442aa2eccbc5a3b7913", "e6f23d852e66496487ad0747c1420f87", "b6a23aa55a6f4b469b76cdbcee54e3a9", "7da5ea2c64804cff9c9f3f55cfde1855", "e58901b89f3b435b990a063dde077f1a", "1b8b42e39d524b4d813bf9ca09db350d", "ceb901d3f75f4e34b0a46686b2474f6c", "ad2be4bfb97347299f15e7095a02ae9b", "e3acc6dcaef244509dd23638fbb3b8ef", "<KEY>", "<KEY>", "0f65e97606454120a5264b9d2e07e954", "ee2edd7ca8fc44e8869372da48c5a895", "<KEY>"]} id="CnismduW8IKE" outputId="06289145-f94c-4bf2-fa62-366b4526b02e"
# Combining all widgets
VBox([widgets.Label('Select your car!'),
btn_upload, btn_run, out_pl, lbl_pred])
| app.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from gs_quant.common import Currency
from gs_quant.instrument import IRXccySwap
from gs_quant.session import Environment, GsSession
# external users should substitute their client id and secret; please skip this step if using internal jupyterhub
GsSession.use(Environment.PROD, client_id=None, client_secret=None, scopes=('run_analytics',))
xswap = IRXccySwap(payer_currency=Currency.EUR, receiver_currency=Currency.USD,
effective_date='3m', termination_date='10y')
print(xswap.price())
| gs_quant/documentation/02_pricing_and_risk/00_instruments_and_measures/examples/01_rates/000112_calc_xccy_swap_price.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import pandas as pd
from IPython.core.display import HTML
css = open('style-table.css').read() + open('style-notebook.css').read()
HTML('<style>{}</style>'.format(css))
sales1 = pd.read_csv('sales1.csv')
sales1
sales2 = pd.read_csv('sales2.csv')
sales2.fillna('')
# ### Challenge: first combine these sales together into a single dataframe, then compute how much money consumers spent on each book in each currency.
# +
# First: clean up data in table 1
sales1
# +
# rename columns to more uniform names
df1 = sales1.rename(columns = {
'Book title' : 'title',
'Number sold' : 'quantity_sold',
'Sales price' : 'sale_price',
'Royalty paid' : 'royalty'
}
)
df1
# +
# next: clean up data in table 2
t = sales2['Title'] # renaming so don't have to keep doing sales2.Title
t = t.where(t.str.endswith(')')).str.split().str[-1].str.strip('()')
sales2
# -
| Exercises-6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Quality Prediction in a Mining Process by using RNN
#
# In this notebook, we are going to predict how much impurity is in the ore concentrate. As this impurity is measured every hour, if we can predict how much silica (impurity) is in the ore concentrate, we can help the engineers, giving them early information to take actions. Hence, they will be able to take corrective actions in advance (reduce impurity, if it is the case) and also help the environment (reducing the amount of ore that goes to tailings as you reduce silica in the ore concentrate). To this end, we are going to use the dataset **Quality Prediction in a Mining Process Data** from [Kaggle](https://www.kaggle.com/edumagalhaes/quality-prediction-in-a-mining-process/home).
#
# In order to have a clean notebook, some functions are implemented in the file *utils.py* (e.g., plot_loss_and_accuracy).
#
# Summary:
# - [Data Pre-processing](#data_preprocessing)
# - [Data Visualisation](#data_viz)
# - [Data Normalisation](#normalisation)
# - [Building the Models](#models)
# - [Splitting the Data into Train and Test Sets](#split)
# - [Gated Recurrent Unit (GRU)](#gru)
# - [Long-short Term Memory (LSTM)](#lstm)
#
# __All the libraries used in this notebook are <font color='red'>Open Source</font>__.
# +
# Standard libraries - no deep learning yet
import numpy as np # written in C, is faster and robust library for numerical and matrix operations
import pandas as pd # data manipulation library, it is widely used for data analysis and relies on numpy library.
import matplotlib.pyplot as plt # for plotting
from datetime import datetime # supplies classes for manipulating dates and times in both simple and complex ways
from utils import *
# the following to lines will tell to the python kernel to alway update the kernel for every utils.py
# modification, without the need of restarting the kernel.
# Of course, for every motification in util.py, we need to reload this cell
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# -
# ## Data Pre-processing
# <a id='data_preprocessing'></a>
#
# **First download the dataset (click [here](https://www.kaggle.com/edumagalhaes/quality-prediction-in-a-mining-process/download)) unzip quality-prediction-in-a-mining-process.zip**
#
# The **Quality Prediction in a Mining Process Data** includes ([Kaggle](https://www.kaggle.com/edumagalhaes/quality-prediction-in-a-mining-process/home)):
#
# - The first column shows time and date range (from march of 2017 until september of 2017). Some columns were sampled every 20 second. Others were sampled on a hourly base. *This make the data processing harder, however, for this tutorial we will not re-sample the data*.
# - The second and third columns are quality measures of the iron ore pulp right before it is fed into the flotation plant.
# - Column 4 until column 8 are the most important variables that impact in the ore quality in the end of the process.
# - Column 9 until column 22, we can see process data level and air flow inside the flotation columns, which also impact in ore quality.
# - The last two columns are the final iron ore pulp quality measurement from the lab. Target is to predict the last column, which is the % of silica in the iron ore concentrate.
#
#
# We are going to use [Pandas](https://pandas.pydata.org/) for the data processing. The function [read_csv](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.read_csv.html) is going to be used to read the csv file.
# +
dataset = pd.read_csv('../data/MiningProcess_Flotation_Plant_Database.csv',index_col=0, decimal=",")
# Set the index name to 'date'
dataset.index.name = 'date'
dataset.head()
# -
# Given our time and computational resources restrictions, we are going to select the first 100,000 observations for this tutorial.
dataset = dataset.iloc[:100000,:]
# ### Data Visualisation
# <a id='data_viz'></a>
# +
# Ploting the Silica Concentrate
plt.figure(figsize = (15, 5))
plt.xlabel("x")
plt.ylabel("Silica Concentrate")
dataset['% Silica Concentrate'].plot()
# -
# ## Data Normalisation
# <a id='normalisation'></a>
#
# Here are going to normalise all the features and transform the data into a supervised learning problem. The features to be predicted are removed, as we would like to predict just the *Silica Concentrate* (last element in every feature array).
#
# ### Transforming the data into a supervised learning problem
#
# This step will involve framing the dataset as a **supervised learning problem**. As we would like to predict the "silica concentrate", we will set the corresponding column to be the output (label $y$).
#
# We would like to predict the silica concentrate ( $y_t$) at the current time ($t$) given the measurements at the prior time steps (lets say $t-1, t-2, \dots t-n$, in which $n$ is the number of past observations to be used to forcast $y_t$).
#
# The function **create_window** (see _utils.py_) converts the time-series to a supervised learning problem. The new dataset is constructed as a **DataFrame**, with each column suitably named both by variable number and time step, for example, $var1(t-1)$ for **%Iron Feed** at the previous observation ($t-1$). This allows you to design a variety of different time step sequence type forecasting problems from a given univariate or multivariate time series.
# +
# Scikit learn libraries
from sklearn.preprocessing import MinMaxScaler #Allows normalisation
# Convert the data to float
values = dataset.values.astype('float32')
# Normalise features
scaler = MinMaxScaler(feature_range=(0, 1))
scaled = scaler.fit_transform(values)
# Specify the number of lag
n_in = 5
n_features = 23
# Transform the time-series to a supervised learning problem representation
reframed = create_window(scaled, n_in = n_in, n_out = 1, drop_nan = True)
# Summarise the new frames (reframes)
print(reframed.head(1))
# -
# ## Building the Models
# <a id='models'></a>
# So far, we just preprocessed the dataset. Now, we are going to build the following sequential models:
#
# - [Gated Recurrent Unit (GRU)](#gru)
# - [Long Short-Term Memory (LSTM)](#lstm)
#
# The models consists in a **many_to_one** architecture, in which the input is a **sequence** of the past observations and the output is the predicted value (in this case with dimension equal 1).
# +
from keras.models import Sequential
from keras.layers import Dense, Dropout
from keras.layers import LSTM, GRU
from sklearn.metrics import mean_squared_error # allows compute the mean square error to performance analysis
# -
# ### Splitting the Data into Train and Test Sets
# <a id='split'></a>
# +
# split into train and test sets
values = reframed.values
# We will use 80% of the data for training and 20% for testing
n_train = round(0.8 * dataset.shape[0])
train = values[:n_train, :]
test = values[n_train:, :]
# Split into input and outputs
n_obs = n_in * n_features # the number of total features is given by the number of past
# observations * number of features. In this case we have
# 5 past observations and 23 features, so the number of total
# features is 115.
x_train, y_train = train[:, :n_obs], train[:, n_features-1] # note that fore y_train, we are removing
# just the last observation of the
# silica concentrate
x_test, y_test = test[:, :n_obs], test[:, n_features-1]
print('Number of total features (n_obs): ', x_train.shape[1])
print('Number of samples in training set: ', x_train.shape[0])
print('Number of samples in testing set: ', x_test.shape[0])
# Reshape input to be 3D [samples, timesteps, features]
x_train = x_train.reshape((x_train.shape[0], n_in, n_features))
x_test = x_test.reshape((x_test.shape[0], n_in, n_features))
# -
# ### Gated Recurrent Unit (GRU)
# <a id='gru'></a>
#
# To build the model, we are going use the following components from Keras:
#
# - [Sequencial](https://keras.io/models/sequential/): allows us to create models layer-by-layer.
# - [GRU](https://keras.io/layers/recurrent/): provides a GRU architecture
# - [Dense](https://keras.io/layers/core/): provides a regular fully-connected layer
# - [Activation](https://keras.io/activations/): defines the activation function to be used
#
# Basically, we can define the sequence of the model by using _Sequential()_:
#
# ```python
# model = Sequential()
# model.add(GRU(...))
# ...
# ```
# where the function _add(...)_ that stack the layers. Once created the model, we can configure the training by using the function [compile](https://keras.io/models/model/). Here we need to define the [loss](https://keras.io/losses/) function (mean squared error, mean absolute error, cosine proximity, among others.) and the [optimizer](https://keras.io/optimizers/) (Stochastic gradient descent, RMSprop, adam, among others), as follows:
#
# ```python
# model.compile(loss = "...",
# optimizer = "...")
# ```
#
# Also, we have the option to see a summary representation of the model by using the function [summary](https://keras.io/models/about-keras-models/#about-keras-models). This function summarises the model and tell us the number of parameters that we need to tune.
# LLPS-2022-02-03:
#
# * the ```model_gru.add(GRU...``` below rises *"NotImplementedError: Cannot convert a symbolic Tensor (gru_2/strided_slice:0) to a numpy array. This error may indicate that you're trying to pass a Tensor to a NumPy call, which is not supported"
#
# * problem seems to be related to numpy version in this Docker image rio_cs_p1 haridoop/cs_p1:latest, which is '1.21.2'. LLPS ```np.__version__``` is '1.20.3' and works!
#
# * People report that TF was built with 'numpy ~= 1.19.2' so, thats the reason! REF: https://github.com/tensorflow/tensorflow/issues/47242
np.__version__
# +
# Define the model.
model_gru = Sequential()
# the input_shape is the number of past observations (n_in) and the number of features
# per past observations (23)
model_gru.add(GRU(input_shape=(x_train.shape[1], x_train.shape[2]),
units = 128,
return_sequences = False))
model_gru.add(Dense(units=1))
# We compile the model by defining the mean absolute error (denoted by mae) as loss function and
# adam as optimizer
model_gru.compile(loss = "mae",
optimizer = "adam")
# just print the model
model_gru.summary()
# -
# ### Training the Model
# <a id=train></a>
#
# Once defined the model, we need to train it by using the function [fit](https://keras.io/models/model/). This function performs the optmisation step. Hence, we can define the following parameters such as:
#
# - batch size: defines the number of samples that will be propagated through the network
# - epochs: defines the number of times in which all the training set (x_train_scaled) are used once to update the weights
# - validation split: defines the percentage of training data to be used for validation
# - among others (click [here](https://keras.io/models/model/) for more information)
#
# This function return the _history_ of the training, that can be used for further performance analysis.
# Training
hist_gru = model_gru.fit(x_train, y_train,
epochs=50,
batch_size=256,
validation_split = 0.1,
verbose=1, # To not print the output, set verbose=0
shuffle=False)
# ### Prediction and Performance Analysis
# <a id='performance_gru'></a>
#
# Here we can see if the model overfits or underfits. First, we are going to plot the 'loss' and the 'Accuracy' in from the training step.
plot_loss(hist_gru)
# Once the model was trained, we can use the function [predict](https://keras.io/models/model/) for prediction tasks. We are going to use the function **inverse_transform** (see _utils.py_) to invert the scaling (transform the values to the original ones).
#
# Given the predictions and expected values in their original scale, we can then compute the error score for the model.
# +
yhat_gru = model_gru.predict(x_test)
# performing the inverse transform on test_X and yhat_rnn
inv_y_gru, inv_yhat_gru = inverse_transform_multiple(x_test, y_test, yhat_gru, scaler, n_in, n_features)
# calculate RMSE
mse_gru = mean_squared_error(inv_y_gru, inv_yhat_gru)
print('Test MSE: %.3f' % mse_gru)
# -
# ### Visualising the predicted Data
plot_comparison([inv_y_gru, inv_yhat_gru],
['Test_Y value','Predicted Value'],
title='Prediction Comparison')
plot_comparison([inv_y_gru[0:300], inv_yhat_gru[0:300]],
['Test_Y value', 'Predicted Value'],
title='Prediction Comparison of the first 300 Observations')
plot_comparison([inv_y_gru[5500:5800], inv_yhat_gru[5500:5800]],
['Test_Y value', 'Predicted Value'],
title='Prediction Comparison of a Farthest 300 Observations')
# ### Long-Short Term Memory (LSTM)
# <a id='lstm'></a>
#
# Now **you** are going to build the model based on LSTM. Like GRU, we are going use the following components from Keras:
#
# - [Sequencial](https://keras.io/models/sequential/): allows us to create models layer-by-layer.
# - [LSTM](https://keras.io/layers/recurrent/): provides a LSTM architecture
# - [Dense](https://keras.io/layers/core/): provides a regular fully-connected layer
# - [Activation](https://keras.io/activations/): defines the activation function to be used
#
# Basically, you are going to define the sequence of the model by using _Sequential()_:
#
# ```python
# model = Sequential()
# model.add(LSTM(...))
# ...
# ```
#
# and configure the training by using the function [compile](https://keras.io/models/model/):
#
# ```python
# model.compile(loss = "...",
# optimizer = "...")
# ```
#
# Follow the below steps for this task.
# **Step 1**: Create the model:
# 1) Define the number of layers (we suggest at this stage to use just one, but it is up to you)
# 2) Create the fully connected layer
#
# For example:
#
# ```python
# # Define the model.
# model_lstm = Sequential()
#
# # Stacking just one LSTM
# model_lstm.add(LSTM(input_shape=(train_X.shape[1], train_X.shape[2]),
# units = 128,
# return_sequences = False))
#
# # Fully connected layer
# model_lstm.add(Dense(units=1))
#
# ```
# **Step 2**: Configure the training:
# 1) Define the loss function (e.g., 'mae' for mean average error or 'mse' for mean squared error)
# 2) Define the optimiser (e.g., 'adam', 'rmsprop', 'sgd', 'adagrad, etc)
#
# For example:
#
# ```python
# model_lstm.compile(loss = "mae",
# optimizer = "adam")
#
# ```
# **Step 3:** Call the function
# ```python
# model_lstm.summary()
# ```
# to summarise the model.
# **Step 4:** Defined the number of epochs, validation_split and batch_size that best fit for you model and call the function fit to train the model.
#
# For example:
# ```python
# hist_lstm = model_lstm.fit(x_train, y_train,
# epochs=50,
# batch_size=256,
# validation_split = 0.1,
# verbose=1, shuffle=False)
# ```
# #### Using the history
#
# Here we can see if the model overfits or underfits
plot_loss(hist_lstm)
# +
yhat_lstm = model_lstm.predict(x_test)
# performing the inverse transform on test_X and yhat_rnn
inv_y_lstm, inv_yhat_lstm = inverse_transform_multiple(x_test, y_test, yhat_lstm, scaler, n_in, n_features)
# calculate RMSE
mse_lstm = mean_squared_error(inv_y_lstm, inv_yhat_lstm)
print('Test MSE: %.3f' % mse_lstm)
# -
# ### Visualising the predicted Data
plot_comparison([inv_y_lstm, inv_yhat_lstm],
['Test_Y value','Predicted Value'],
title='Prediction Comparison')
plot_comparison([inv_y_lstm[0:300], inv_yhat_lstm[0:300]],
['Test_Y value', 'Predicted Value'],
title='Prediction Comparison of first 300 Observations')
plot_comparison([inv_y_lstm[5500:5800], inv_yhat_lstm[5500:5800]],
['Test_Y value', 'Predicted Value'],
title='Prediction Comparison of a Farthest 300 Observations')
# ## Models comparison
#
# **Exercise**: run the code below and discuss the results.
plot_comparison([inv_y_lstm[0:300],
inv_yhat_gru[0:300], inv_yhat_lstm[0:300]],
['Original', 'GRU', 'LSTM'],
title='Prediction Comparison of the First 300 Observations')
plot_comparison([inv_y_lstm[5500:5800],
inv_yhat_gru[5500:5800], inv_yhat_lstm[5500:5800]],
['Original', 'GRU', 'LSTM'],
title='Prediction Comparison of a Farthest 300 Observations')
print('Comparing the MSE of the three models:')
print(' GRU: ', mse_gru)
print(' LSTM: ', mse_lstm)
| notebooks/pm2-2-quality-prediction-in-a-mining-process.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# +
# default_exp data/realData
# -
# +
# export
import numpy as np
import mat73
from scipy.io import loadmat
import h5py
# -
d = loadmat('/home/dzeiberg/ClassPriorEstimation/rawDatasets/anuran.mat')
d.keys()
# export
def getParams(nPDistr=lambda: np.random.poisson(25) + 1,
nUDistr=lambda: np.random.poisson(30) + 1,
alphaDistr=lambda: np.random.beta(2,10)):
nP = nPDistr()
nU = nUDistr()
alpha = alphaDistr()
numUnlabeledPos = max(1,int(alpha * nU))
numUnlabeledNeg = max(1,nU - numUnlabeledPos)
return nP, nU, alpha, numUnlabeledPos, numUnlabeledNeg
# export
def getBagDict(nP, nU, alpha, numUnlabeledPos, numUnlabeledNeg, d):
# get indices of all positives and negatives
posIdxs = np.where(d["y"] == 1)[0]
negIdxs = np.where(d["y"] == 0)[0]
# sample positives
posSampleIDXS = np.random.choice(posIdxs,replace=True,size=nP)
# sample unlabeled
unlabeledPosSampleIDXS = np.random.choice(posIdxs,replace=True,size=numUnlabeledPos)
unlabeledNegSampleIDXS = np.random.choice(negIdxs,replace=True,size=numUnlabeledNeg)
unlabeledSampleIDXS = np.concatenate((unlabeledPosSampleIDXS, unlabeledNegSampleIDXS))
posInstances = d["X"][posSampleIDXS]
unlabeledInstances = d["X"][unlabeledSampleIDXS]
hiddenLabels = np.concatenate((np.ones(numUnlabeledPos),
np.zeros(numUnlabeledNeg)))
return {"positiveInstances": posInstances,
"unlabeledInstances": unlabeledInstances,
"hiddenLabels": hiddenLabels,
"alpha_i": alpha,
"nP": nP,
"nU": nU}
# export
class Dataset:
def __init__(self, d):
self.positiveInstances = d["positiveInstances"]
self.unlabeledInstances = d["unlabeledInstances"]
self.trueAlphas = d["alpha_i"]
self.N = self.positiveInstances.shape[0]
self.numP = d["numP"]
self.numU = d["numU"]
self.hiddenLabels = d["hiddenLabels"]
def getBag(self,idx):
p = self.positiveInstances[idx, :self.numP[idx]]
u = self.unlabeledInstances[idx, :self.numU[idx]]
return p,u
def getAlpha(self,idx):
return self.trueAlphas[idx]
def __len__(self):
return self.N
# export
def buildDataset(dsPath, size,
nPDistr=lambda: np.random.poisson(25) + 1,
nUDistr=lambda: np.random.poisson(30) + 1,
alphaDistr=lambda: np.random.beta(2,10)):
try:
ds = loadmat(dsPath)
except:
ds= {}
for k,v in h5py.File(dsPath,"r").items():
ds[k] = np.array(v)
bags = []
for bag in range(size):
nP, nU, alpha, numUnlabeledPos, numUnlabeledNeg = getParams(nPDistr=nPDistr,
nUDistr=nUDistr,
alphaDistr=alphaDistr)
bagDict = getBagDict(nP, nU, alpha, numUnlabeledPos, numUnlabeledNeg, ds)
bags.append(bagDict)
# calculate max num Pos and Unlabeled to set sizes for matrices
maxP = np.max([d["nP"] for d in bags])
maxU = np.max([d["nU"] for d in bags])
dim = bags[0]["positiveInstances"].shape[1]
# init matrices
posMats = np.zeros((len(bags), maxP, dim))
unlabeledMats = np.zeros((len(bags), maxU, dim))
hiddenLabelMats = np.zeros((len(bags), maxU))
alphas = np.zeros((len(bags), 1))
numPos = np.zeros(len(bags),dtype=int)
numU = np.zeros(len(bags),dtype=int)
# fill matrices with bags
for bagNum,bag in enumerate(bags):
posPadding = maxP - bag["nP"]
unlabeledPadding = maxU - bag["nU"]
p_mat= np.concatenate((bag["positiveInstances"],
np.zeros((posPadding, dim))), axis=0)
posMats[bagNum] = p_mat
u_mat= np.concatenate((bag["unlabeledInstances"],
np.zeros((unlabeledPadding, dim))), axis=0)
unlabeledMats[bagNum] = u_mat
hiddenLabelMats[bagNum] = np.concatenate((bag["hiddenLabels"],
np.zeros(unlabeledPadding)))
alphas[bagNum] = bag["alpha_i"]
numPos[bagNum] = bag["nP"]
numU[bagNum] = bag["nU"]
dataset = Dataset({
"positiveInstances": posMats,
"unlabeledInstances": unlabeledMats,
"alpha_i": alphas,
"numP": numPos,
"numU": numU,
"hiddenLabels": hiddenLabelMats
})
return dataset
buildDataset('/home/dzeiberg/ClassPriorEstimation/rawDatasets/activity_recognition_s1.mat',100)
for k,v in h5py.File('/home/dzeiberg/ClassPriorEstimation/rawDatasets/20newsgroups.mat',"r").items():
keys = k
vals = np.array(v)
k,vals
np.array(v)
f["X
buildDataset('/home/dzeiberg/ClassPriorEstimation/rawDatasets/20newsgroups.mat',100)
dataset = buildDataset("/home/dzeiberg/ClassPriorEstimation/rawDatasets/abalone.mat", 100)
len(dataset)
import matplotlib.pyplot as plt
plt.hist(dataset.trueAlphas)
| 11_Construct_Datasets.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.4.2
# language: julia
# name: julia-1.4
# ---
#
# <a id='lucas-asset'></a>
# <div id="qe-notebook-header" style="text-align:right;">
# <a href="https://quantecon.org/" title="quantecon.org">
# <img style="width:250px;display:inline;" src="https://assets.quantecon.org/img/qe-menubar-logo.svg" alt="QuantEcon">
# </a>
# </div>
# # Asset Pricing II: The Lucas Asset Pricing Model
#
#
# <a id='index-0'></a>
# ## Contents
#
# - [Asset Pricing II: The Lucas Asset Pricing Model](#Asset-Pricing-II:-The-Lucas-Asset-Pricing-Model)
# - [Overview](#Overview)
# - [The Lucas Model](#The-Lucas-Model)
# - [Exercises](#Exercises)
# - [Solutions](#Solutions)
# ## Overview
#
# As stated in an [earlier lecture](markov_asset.html), an asset is a claim on a stream of prospective payments.
#
# What is the correct price to pay for such a claim?
#
# The elegant asset pricing model of Lucas [[Luc78]](../zreferences.html#lucas1978) attempts to answer this question in an equilibrium setting with risk averse agents.
#
# While we mentioned some consequences of Lucas’ model [earlier](markov_asset.html#mass-pra), it is now time to work through the model more carefully, and try to understand where the fundamental asset pricing equation comes from.
#
# A side benefit of studying Lucas’ model is that it provides a beautiful illustration of model building in general and equilibrium pricing in competitive models in particular.
#
# Another difference to our [first asset pricing lecture](markov_asset.html) is that the state space and shock will be continous rather than discrete.
# ## The Lucas Model
#
#
# <a id='index-1'></a>
# Lucas studied a pure exchange economy with a representative consumer (or household), where
#
# - *Pure exchange* means that all endowments are exogenous.
# - *Representative* consumer means that either
#
# - there is a single consumer (sometimes also referred to as a household), or
# - all consumers have identical endowments and preferences
#
#
#
# Either way, the assumption of a representative agent means that prices adjust to eradicate desires to trade.
#
# This makes it very easy to compute competitive equilibrium prices.
# ### Basic Setup
#
# Let’s review the set up.
# #### Assets
#
#
# <a id='index-2'></a>
# There is a single “productive unit” that costlessly generates a sequence of consumption goods $ \{y_t\}_{t=0}^{\infty} $.
#
# Another way to view $ \{y_t\}_{t=0}^{\infty} $ is as a *consumption endowment* for this economy.
#
# We will assume that this endowment is Markovian, following the exogenous process
#
# $$
# y_{t+1} = G(y_t, \xi_{t+1})
# $$
#
# Here $ \{ \xi_t \} $ is an iid shock sequence with known distribution $ \phi $ and $ y_t \geq 0 $.
#
# An asset is a claim on all or part of this endowment stream.
#
# The consumption goods $ \{y_t\}_{t=0}^{\infty} $ are nonstorable, so holding assets is the only way to transfer wealth into the future.
#
# For the purposes of intuition, it’s common to think of the productive unit as a “tree” that produces fruit.
#
# Based on this idea, a “Lucas tree” is a claim on the consumption endowment.
# #### Consumers
#
#
# <a id='index-3'></a>
# A representative consumer ranks consumption streams $ \{c_t\} $ according to the time separable utility functional
#
#
# <a id='equation-lt-uf'></a>
# $$
# \mathbb{E} \sum_{t=0}^\infty \beta^t u(c_t) \tag{1}
# $$
#
# Here
#
# - $ \beta \in (0,1) $ is a fixed discount factor
# - $ u $ is a strictly increasing, strictly concave, continuously differentiable period utility function
# - $ \mathbb{E} $ is a mathematical expectation
# ### Pricing a Lucas Tree
#
#
# <a id='index-4'></a>
# What is an appropriate price for a claim on the consumption endowment?
#
# We’ll price an *ex dividend* claim, meaning that
#
# - the seller retains this period’s dividend
# - the buyer pays $ p_t $ today to purchase a claim on
#
# - $ y_{t+1} $ and
# - the right to sell the claim tomorrow at price $ p_{t+1} $
#
#
#
# Since this is a competitive model, the first step is to pin down consumer
# behavior, taking prices as given.
#
# Next we’ll impose equilibrium constraints and try to back out prices.
#
# In the consumer problem, the consumer’s control variable is the share $ \pi_t $ of the claim held in each period.
#
# Thus, the consumer problem is to maximize [(1)](#equation-lt-uf) subject to
#
# $$
# c_t + \pi_{t+1} p_t \leq \pi_t y_t + \pi_t p_t
# $$
#
# along with $ c_t \geq 0 $ and $ 0 \leq \pi_t \leq 1 $ at each $ t $.
#
# The decision to hold share $ \pi_t $ is actually made at time $ t-1 $.
#
# But this value is inherited as a state variable at time $ t $, which explains the choice of subscript.
# #### The dynamic program
#
#
# <a id='index-5'></a>
# We can write the consumer problem as a dynamic programming problem.
#
# Our first observation is that prices depend on current information, and current information is really just the endowment process up until the current period.
#
# In fact the endowment process is Markovian, so that the only relevant
# information is the current state $ y \in \mathbb R_+ $ (dropping the time subscript).
#
# This leads us to guess an equilibrium where price is a function $ p $ of $ y $.
#
# Remarks on the solution method
#
# - Since this is a competitive (read: price taking) model, the consumer will take this function $ p $ as .
# - In this way we determine consumer behavior given $ p $ and then use equilibrium conditions to recover $ p $.
# - This is the standard way to solve competitive equilibrum models.
#
#
# Using the assumption that price is a given function $ p $ of $ y $, we write the value function and constraint as
#
# $$
# v(\pi, y) = \max_{c, \pi'}
# \left\{
# u(c) + \beta \int v(\pi', G(y, z)) \phi(dz)
# \right\}
# $$
#
# subject to
#
#
# <a id='equation-preltbe'></a>
# $$
# c + \pi' p(y) \leq \pi y + \pi p(y) \tag{2}
# $$
#
# We can invoke the fact that utility is increasing to claim equality in [(2)](#equation-preltbe) and hence eliminate the constraint, obtaining
#
#
# <a id='equation-ltbe'></a>
# $$
# v(\pi, y) = \max_{\pi'}
# \left\{
# u[\pi (y + p(y)) - \pi' p(y) ] + \beta \int v(\pi', G(y, z)) \phi(dz)
# \right\} \tag{3}
# $$
#
# The solution to this dynamic programming problem is an optimal policy expressing either $ \pi' $ or $ c $ as a function of the state $ (\pi, y) $.
#
# - Each one determines the other, since $ c(\pi, y) = \pi (y + p(y))- \pi' (\pi, y) p(y) $.
# #### Next steps
#
# What we need to do now is determine equilibrium prices.
#
# It seems that to obtain these, we will have to
#
# 1. Solve this two dimensional dynamic programming problem for the optimal policy.
# 1. Impose equilibrium constraints.
# 1. Solve out for the price function $ p(y) $ directly.
#
#
# However, as Lucas showed, there is a related but more straightforward way to do this.
# #### Equilibrium constraints
#
#
# <a id='index-6'></a>
# Since the consumption good is not storable, in equilibrium we must have $ c_t = y_t $ for all $ t $.
#
# In addition, since there is one representative consumer (alternatively, since
# all consumers are identical), there should be no trade in equilibrium.
#
# In particular, the representative consumer owns the whole tree in every period, so $ \pi_t = 1 $ for all $ t $.
#
# Prices must adjust to satisfy these two constraints.
# #### The equilibrium price function
#
#
# <a id='index-7'></a>
# Now observe that the first order condition for [(3)](#equation-ltbe) can be written as
#
# $$
# u'(c) p(y) = \beta \int v_1'(\pi', G(y, z)) \phi(dz)
# $$
#
# where $ v'_1 $ is the derivative of $ v $ with respect to its first argument.
#
# To obtain $ v'_1 $ we can simply differentiate the right hand side of
# [(3)](#equation-ltbe) with respect to $ \pi $, yielding
#
# $$
# v'_1(\pi, y) = u'(c) (y + p(y))
# $$
#
# Next we impose the equilibrium constraints while combining the last two
# equations to get
#
#
# <a id='equation-lteeq'></a>
# $$
# p(y) = \beta \int \frac{u'[G(y, z)]}{u'(y)} [G(y, z) + p(G(y, z))] \phi(dz) \tag{4}
# $$
#
# In sequential rather than functional notation, we can also write this as
#
#
# <a id='equation-lteeqs'></a>
# $$
# p_t = \mathbb{E}_t \left[ \beta \frac{u'(c_{t+1})}{u'(c_t)} ( y_{t+1} + p_{t+1} ) \right] \tag{5}
# $$
#
# This is the famous consumption-based asset pricing equation.
#
# Before discussing it further we want to solve out for prices.
# ### Solving the Model
#
#
# <a id='index-8'></a>
# Equation [(4)](#equation-lteeq) is a *functional equation* in the unknown function $ p $.
#
# The solution is an equilibrium price function $ p^* $.
#
# Let’s look at how to obtain it.
# #### Setting up the problem
#
# Instead of solving for it directly we’ll follow Lucas’ indirect approach, first setting
#
#
# <a id='equation-ltffp'></a>
# $$
# f(y) := u'(y) p(y) \tag{6}
# $$
#
# so that [(4)](#equation-lteeq) becomes
#
#
# <a id='equation-lteeq2'></a>
# $$
# f(y) = h(y) + \beta \int f[G(y, z)] \phi(dz) \tag{7}
# $$
#
# Here $ h(y) := \beta \int u'[G(y, z)] G(y, z) \phi(dz) $ is a function that
# depends only on the primitives.
#
# Equation [(7)](#equation-lteeq2) is a functional equation in $ f $.
#
# The plan is to solve out for $ f $ and convert back to $ p $ via [(6)](#equation-ltffp).
#
# To solve [(7)](#equation-lteeq2) we’ll use a standard method: convert it to a fixed point problem.
#
# First we introduce the operator $ T $ mapping $ f $ into $ Tf $ as defined by
#
#
# <a id='equation-lteeqt'></a>
# $$
# (Tf)(y) = h(y) + \beta \int f[G(y, z)] \phi(dz) \tag{8}
# $$
#
# The reason we do this is that a solution to [(7)](#equation-lteeq2) now corresponds to a
# function $ f^* $ satisfying $ (Tf^*)(y) = f^*(y) $ for all $ y $.
#
# In other words, a solution is a *fixed point* of $ T $.
#
# This means that we can use fixed point theory to obtain and compute the solution.
# #### A little fixed point theory
#
#
# <a id='index-9'></a>
# Let $ cb\mathbb{R}_+ $ be the set of continuous bounded functions $ f \colon \mathbb{R}_+ \to \mathbb{R}_+ $.
#
# We now show that
#
# 1. $ T $ has exactly one fixed point $ f^* $ in $ cb\mathbb{R}_+ $.
# 1. For any $ f \in cb\mathbb{R}_+ $, the sequence $ T^k f $ converges
# uniformly to $ f^* $.
#
#
# (Note: If you find the mathematics heavy going you can take 1–2 as given and skip to the [next section](#lt-comp-eg))
#
# Recall the [Banach contraction mapping theorem](https://en.wikipedia.org/wiki/Banach_fixed-point_theorem).
#
# It tells us that the previous statements will be true if we can find an
# $ \alpha < 1 $ such that
#
#
# <a id='equation-ltbc'></a>
# $$
# \| Tf - Tg \| \leq \alpha \| f - g \|,
# \qquad \forall \, f, g \in cb\mathbb{R}_+ \tag{9}
# $$
#
# Here $ \|h\| := \sup_{x \in \mathbb{R}_+} |h(x)| $.
#
# To see that [(9)](#equation-ltbc) is valid, pick any $ f,g \in cb\mathbb{R}_+ $ and any $ y \in \mathbb{R}_+ $.
#
# Observe that, since integrals get larger when absolute values are moved to the
# inside,
#
# $$
# \begin{aligned}
# |Tf(y) - Tg(y)|
# & = \left| \beta \int f[G(y, z)] \phi(dz)
# - \beta \int g[G(y, z)] \phi(dz) \right|
# \\
# & \leq \beta \int \left| f[G(y, z)] - g[G(y, z)] \right| \phi(dz)
# \\
# & \leq \beta \int \| f - g \| \phi(dz)
# \\
# & = \beta \| f - g \|
# \end{aligned}
# $$
#
# Since the right hand side is an upper bound, taking the sup over all $ y $
# on the left hand side gives [(9)](#equation-ltbc) with $ \alpha := \beta $.
#
#
# <a id='lt-comp-eg'></a>
# ### Computation – An Example
#
#
# <a id='index-10'></a>
# The preceding discussion tells that we can compute $ f^* $ by picking any arbitrary $ f \in cb\mathbb{R}_+ $ and then iterating with $ T $.
#
# The equilibrium price function $ p^* $ can then be recovered by $ p^*(y) = f^*(y) / u'(y) $.
#
# Let’s try this when $ \ln y_{t+1} = \alpha \ln y_t + \sigma \epsilon_{t+1} $ where $ \{\epsilon_t\} $ is iid and standard normal.
#
# Utility will take the isoelastic form $ u(c) = c^{1-\gamma}/(1-\gamma) $, where $ \gamma > 0 $ is the coefficient of relative risk aversion.
#
# Some code to implement the iterative computational procedure can be found below:
# ### Setup
# + hide-output=true
using InstantiateFromURL
# optionally add arguments to force installation: instantiate = true, precompile = true
github_project("QuantEcon/quantecon-notebooks-julia", version = "0.8.0")
# + hide-output=false
using LinearAlgebra, Statistics
using Distributions, Interpolations, Parameters, Plots, QuantEcon, Random
gr(fmt = :png);
# + hide-output=false
# model
function LucasTree(;γ = 2.0,
β = 0.95,
α = 0.9,
σ = 0.1,
grid_size = 100)
ϕ = LogNormal(0.0, σ)
shocks = rand(ϕ, 500)
# build a grid with mass around stationary distribution
ssd = σ / sqrt(1 - α^2)
grid_min, grid_max = exp(-4ssd), exp(4ssd)
grid = range(grid_min, grid_max, length = grid_size)
# set h(y) = β * int u'(G(y,z)) G(y,z) ϕ(dz)
h = similar(grid)
for (i, y) in enumerate(grid)
h[i] = β * mean((y^α .* shocks).^(1 - γ))
end
return (γ = γ, β = β, α = α, σ = σ, ϕ = ϕ, grid = grid, shocks = shocks, h = h)
end
# approximate Lucas operator, which returns the updated function Tf on the grid
function lucas_operator(lt, f)
# unpack input
@unpack grid, α, β, h = lt
z = lt.shocks
Af = LinearInterpolation(grid, f, extrapolation_bc=Line())
Tf = [ h[i] + β * mean(Af.(grid[i]^α .* z)) for i in 1:length(grid) ]
return Tf
end
# get equilibrium price for Lucas tree
function solve_lucas_model(lt;
tol = 1e-6,
max_iter = 500)
@unpack grid, γ = lt
i = 0
f = zero(grid) # Initial guess of f
error = tol + 1
while (error > tol) && (i < max_iter)
f_new = lucas_operator(lt, f)
error = maximum(abs, f_new - f)
f = f_new
i += 1
end
# p(y) = f(y) * y ^ γ
price = f .* grid.^γ
return price
end
# -
# An example of usage is given in the docstring and repeated here
# + hide-output=false
Random.seed!(42) # For reproducible results.
tree = LucasTree(γ = 2.0, β = 0.95, α = 0.90, σ = 0.1)
price_vals = solve_lucas_model(tree);
# -
# Here’s the resulting price function
# + hide-output=false
plot(tree.grid, price_vals, lw = 2, label = "p*(y)")
plot!(xlabel = "y", ylabel = "price", legend = :topleft)
# -
# The price is increasing, even if we remove all serial correlation from the endowment process.
#
# The reason is that a larger current endowment reduces current marginal
# utility.
#
# The price must therefore rise to induce the household to consume the entire endowment (and hence satisfy the resource constraint).
#
# What happens with a more patient consumer?
#
# Here the orange line corresponds to the previous parameters and the green line is price when $ \beta = 0.98 $.
#
#
# <a id='mass-lt-cb'></a>
# <img src="_static/figures/solution_mass_ex2.png" style="width:80%;">
#
#
# We see that when consumers are more patient the asset becomes more valuable, and the price of the Lucas tree shifts up.
#
# Exercise 1 asks you to replicate this figure.
# ## Exercises
#
#
# <a id='lucas-asset-ex1'></a>
# ### Exercise 1
#
# Replicate [the figure](#mass-lt-cb) to show how discount rates affect prices.
# ## Solutions
# + hide-output=false
plot()
for β in (.95, 0.98)
tree = LucasTree(;β = β)
grid = tree.grid
price_vals = solve_lucas_model(tree)
plot!(grid, price_vals, lw = 2, label = "beta = beta_var")
end
plot!(xlabel = "y", ylabel = "price", legend = :topleft)
| multi_agent_models/lucas_model.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .fs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: .NET (F#)
// language: F#
// name: .net-fsharp
// ---
// + [markdown] tags=[]
// <h2>--- Day 9: All in a Single Night ---</h2>
// -
// [](https://mybinder.org/v2/gh/oddrationale/AdventOfCode2015FSharp/master?urlpath=lab%2Ftree%2FDay09.ipynb)
// <p>Every year, Santa manages to deliver all of his presents in a single night.</p>
// <p>This year, however, he has some <span title="Bonus points if you recognize all of the locations.">new locations</span> to visit; his elves have provided him the distances between every pair of locations. He can start and end at any two (different) locations he wants, but he must visit each location exactly once. What is the <em>shortest distance</em> he can travel to achieve this?</p>
// <p>For example, given the following distances:</p>
// <pre><code>London to Dublin = 464
// London to Belfast = 518
// Dublin to Belfast = 141
// </code></pre>
// <p>The possible routes are therefore:</p>
// <pre><code>Dublin -> London -> Belfast = 982
// London -> Dublin -> Belfast = 605
// London -> Belfast -> Dublin = 659
// Dublin -> Belfast -> London = 659
// Belfast -> Dublin -> London = 605
// Belfast -> London -> Dublin = 982
// </code></pre>
// <p>The shortest of these is <code>London -> Dublin -> Belfast = 605</code>, and so the answer is <code>605</code> in this example.</p>
// <p>What is the distance of the shortest route?</p>
// + dotnet_interactive={"language": "fsharp"}
open System.Collections.Generic
// + dotnet_interactive={"language": "fsharp"} tags=[]
let input = File.ReadAllLines @"input/09.txt"
// + dotnet_interactive={"language": "fsharp"}
let createAdjacencyMatrix (input: string[]) =
let parseLine (line: string) =
let split1 = line.Split(" = ")
let split2 = split1.[0].Split(" ")
{| From = split2[0]; To = split2[2]; Distance = split1[1] |> int |}
let distances =
input
|> Array.map parseLine
let allDistances =
distances
|> Array.map (fun d -> {| From = d.To; To = d.From; Distance = d.Distance |})
|> Array.append distances
let locations =
allDistances
|> Array.map (fun d -> d.From)
|> Array.distinct
let findDistance f t =
if f = t then
0
else
(allDistances |> Array.find (fun d -> d.From = f && d.To = t)).Distance
dict [for f in locations -> f, dict [for t in locations -> t, findDistance f t]]
// + dotnet_interactive={"language": "fsharp"}
let graph = createAdjacencyMatrix input
// -
// Got the `permute` function [from StackOverflow](https://stackoverflow.com/a/3129136).
// + dotnet_interactive={"language": "fsharp"}
let rec distribute e = function
| [] -> [[e]]
| x::xs' as xs -> (e::xs)::[for xs in distribute e xs' -> x::xs]
let rec permute = function
| [] -> [[]]
| e::xs -> List.collect (distribute e) (permute xs)
// + dotnet_interactive={"language": "fsharp"}
let calculateDistance (graph: IDictionary<string,IDictionary<string,int>>) (route: seq<string>) =
route
|> Seq.windowed 2
|> Seq.map (fun pair -> graph.[pair.[0]].[pair.[1]])
|> Seq.sum
// + dotnet_interactive={"language": "fsharp"}
#!time
graph.Keys
|> List.ofSeq
|> permute
|> Seq.map (fun route -> calculateDistance graph route)
|> Seq.min
// -
// <h2 id="part2">--- Part Two ---</h2>
// <p>The next year, just to show off, Santa decides to take the route with the <em>longest distance</em> instead.</p>
// <p>He can still start and end at any two (different) locations he wants, and he still must visit each location exactly once.</p>
// <p>For example, given the distances above, the longest route would be <code>982</code> via (for example) <code>Dublin -> London -> Belfast</code>.</p>
// <p>What is the distance of the longest route?</p>
// + dotnet_interactive={"language": "fsharp"}
#!time
graph.Keys
|> List.ofSeq
|> permute
|> Seq.map (fun route -> calculateDistance graph route)
|> Seq.max
// -
// [Prev](Day08.ipynb) | [Next](Day10.ipynb)
| Day09.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.display import YouTubeVideo, HTML, Image
# +
# %%HTML
<div>
<span style="font-size:18px" align="right"><NAME> (Dajebbar)</span>
<br>
<span tyle="font-size:16px">AI Programming with Python</span>
<br>
<span><b>Licence CC BY-NC-ND</b></span>
</div>
# -
Image('Img/neural-python.jpg', width=150, height=150)
# # Coding, character sets and unicode
# +
YouTubeVideo(id='oXVmZGN6plY', width=900, height=400)
# -
# ##### A character is not a byte
# With Unicode, we broke the pattern *one character* == *one byte*. Also in Python 3, when it comes to manipulating data from various data sources:
#
# * the `byte` type is appropriate if you want to load raw binary data into memory, in the form of bytes therefore;
# * the `str` type is suitable for representing a string of characters - which again are not necessarily bytes;
# * switching from one of these types to the other by encoding and decoding operations, as illustrated below;
# * and for **all** encoding and decoding operations, it is necessary to know the encoding used.
# 
# You can call the `encode` and` decode` methods without specifying the encoding (in this case Python chooses the default encoding on your system). However, it is far better to be explicit and choose your encoding. If in doubt, it is recommended to **explicitly specify** `utf-8`, which generalizes to the detriment of older encodings like` cp1252` (Windows) and `iso8859- *`, than to leave the host system choose for you.
# ### What is an encoding?
# As you know, a computer's memory - or disk - can only store binary representations. So there is no "natural" way to represent a character like 'A', a quotation mark or a semicolon.
#
# An encoding is used for this, for example [the code `US-ASCII`] (http://www.asciitable.com/) stipulates, to make it simple, that an 'A' is represented by byte 65 which is written in binary 01000001. It turns out that there are several encodings, of course incompatible, depending on the system and language. You can find more details below.
#
# The important point is that in order to be able to open a file "properly", you must of course have the **contents** of the file, but you must also know the **encoding** that was used to write it.
# ### Precautions to be taken when encoding your source code
# Encoding is not just about string objects, but also your source code. **Python 3** assumes that your source code uses **by default the `UTF-8` encoding**. We advise you to keep this encoding which is the one that will offer you the most flexibility.
# You can still change the encoding **of your source code** by including in your files, **in the first or second line**, a declaration like this:
#
# ```python
# # -*- coding: <nom_de_l_encodage> -*-
#
# ```
# or more simply, like this:
#
#
# ```python
# # coding: <nom_de_l_encodage>
# ```
#
# Note that the first option is also interpreted by the _Emacs_ text editor to use the same encoding. Apart from the use of Emacs, the second option, simpler and therefore more pythonic, is to be preferred.
# The name **`UTF-8`** refers to **Unicode** (or to be precise, the most popular encoding among those defined in the Unicode standard, as we will see below). On some older systems you may need to use a different encoding. To determine the value to use in your specific case you can do in the interactive interpreter:
# ```python
# # this must be performed on your machine
# import sys
# print(sys.getdefaultencoding())
# ```
# For example with old versions of Windows (in principle increasingly rare) you may have to write:
# ```python
# # coding: cp1252
# ```
# The syntax of the `coding` line is specified in [this documentation] (https://docs.python.org/3/reference/lexical_analysis.html#encoding-declarations) and in [PEP 263] (https://www.python.org/dev/peps/pep-0263/).
# ### The great misunderstanding
# If I send you a file containing French encoded with, say, [ISO / IEC 8859-15 - a.k.a. `Latin-9`] (http://en.wikipedia.org/wiki/ISO/IEC_8859-15); you can see in the table that a '€' character will be materialized in my file by a byte '0xA4', that is 164.
#
# Now imagine that you are trying to open this same file from an old Windows computer configured for French. If it is not given any indication of the encoding, the program that will read this file on Windows will use the system default encoding, ie [CP1252] (http: //en.wikipedia. org / wiki / Windows-1252). As you can see in this table, the byte '0xA4' corresponds to the character ¤ and this is what you will see instead of €.
#
# Contrary to what one might hope, this type of problem cannot be solved by adding a tag `# coding: <name_of_encoding>`, which only acts on the encoding used * to read the source file in question * (the one that contains the tag).
#
# To correctly solve this type of problem, you must explicitly specify the encoding to be used to decode the file. And therefore have a reliable way to determine this encoding; which is not always easy, moreover, but unfortunately that is another discussion.
# This means that to be completely clean, you must be able to explicitly specify the `encoding` parameter when calling all the methods that are likely to need it.
# ### Why does it work locally?
# When the producer (the program that writes the file) and the consumer (the program that reads it) run on the same computer, everything works fine - in general - because both programs come down to the encoding defined as encoding. by default.
#
# There is however a limit, if you are using a minimally configured Linux, it may default to the `US-ASCII` encoding - see below - which being very old does not" know "a simple é, nor a fortiori €. To write French, it is therefore necessary at least that the default encoding of your computer contains French characters, such as:
#
# * `ISO 8859-1` (` Latin-1`)
# * `ISO 8859-15` (` Latin-9`)
# * `UTF-8`
# * `CP1252`
#
# Again in this list, UTF-8 should be clearly preferred when possible.
# + language="html"
# <script src="https://cdn.rawgit.com/parente/4c3e6936d0d7a46fd071/raw/65b816fb9bdd3c28b4ddf3af602bfd6015486383/code_toggle.js"></script>
#
| Lecture 2 - Coding, character sets and unicode.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Worlddatascience/DataScienceCohort/blob/master/Project%2010_How_to_Create_a_Technical_Analysis_Interactive_Dashboard.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="z07xmDUDvFny"
# # <NAME> - Financial Data Scientist - [Linkedin](https://https://www.linkedin.com/in/anadedatascientist/)
# + [markdown] id="OKD97Gqu2Crg"
# In this Model I will create a Technical Analysis Dashboard
# + [markdown] id="QvW1-QYLvPhE"
# References:
#
# https://github.com/PacktPublishing/Python-for-Finance-Cookbook/blob/master/Chapter%2002/chapter_2.ipynb
#
# https://stackoverflow.com/questions/59031964/i-got-chart-studio-exceptions-plotlyrequesterror-authentication-credentials-we
#
#
# + id="DNEmf1i5vZO9" outputId="c942f62c-70d1-49dc-fafe-7441f1739291" colab={"base_uri": "https://localhost:8080/", "height": 454}
pip install yfinance #Code to install yfinance
# + id="ZOnA1vFix6F6" outputId="f8b685de-5f15-41cb-db3c-2bca1e60cb72" colab={"base_uri": "https://localhost:8080/", "height": 235}
pip install chart_studio
# + id="ulvD4gSyyFaz"
import chart_studio.plotly as py
# + id="urFjwb0h0sU7" outputId="04e719e8-b1a5-4f81-fd26-cdeed73cd0d8" colab={"base_uri": "https://localhost:8080/", "height": 270}
pip install plotly --upgrade
# + id="IdPXkWf8IeHo" outputId="1f797a41-2d59-441d-a651-b026daebcc64" colab={"base_uri": "https://localhost:8080/", "height": 353}
pip install cufflinks==0.8.2
# + id="OuLiRxd51X4T"
import pandas as pd
import yfinance as yf
import numpy as np
import ipywidgets as wd
from __future__ import absolute_import
import json
import copy
import pandas as pd
# %matplotlib inline
# + id="3DJuDvW7vey2"
Nasdaq100 = ['ADBE','PYPL','GOOGL', 'FB', 'TSLA', 'AMZN', 'NVDA']
indicators = ['Bollinger Bands', 'MACD', 'RSI']
#Moving Average Convergence Divergence (MACD)
# + id="Lyyb3wBzvTnh"
def ta_dashboard(asset, indicator, start_date, end_date,
bb_k, bb_n, macd_fast, macd_slow, macd_signal,
rsi_periods, rsi_upper, rsi_lower):
df = yf.download(asset,
start=start_date,
end=end_date,
progress=False,
auto_adjust=True)
if 'Bollinger Bands' in indicator:
add_Bollinger_Bands(periods=bb_n,
boll_std=bb_k)
if 'MACD' in indicator:
add_MACD(fast_period=macd_fast,
slow_period=macd_slow,
signal_period=macd_signal)
if 'RSI' in indicator:
add_rsi(periods=rsi_periods,
rsi_upper=rsi_upper,
rsi_lower=rsi_lower,
showbands=True)
return iplot()
# + id="d48BGBNoKGsP"
# + id="gHQgcWFrwc3P"
stocks_selector = wd.Dropdown(
options=Nasdaq100,
value=Nasdaq100[0],
description='Asset'
)
indicator_selector = wd.SelectMultiple(
description='Indicator',
options=indicators,
value=[indicators[0]]
)
start_date_selector = wd.DatePicker(
description='Start Date',
value=pd.to_datetime('2018-01-01'),
continuous_update=False
)
end_date_selector = wd.DatePicker(
description='End Date',
value=pd.to_datetime('2018-12-31'),
continuous_update=False
)
# + id="rZLfQtBmw6hP"
main_selector_label = wd.Label('Main parameters',
layout=wd.Layout(height='45px'))
main_selector_box = wd.VBox(children=[main_selector_label,
stocks_selector,
indicator_selector,
start_date_selector,
end_date_selector])
# + id="fl7L1CQCw-Nh"
bb_label = wd.Label('Bollinger Bands')
n_param = wd.IntSlider(value=20, min=1, max=40, step=1,
description='N:', continuous_update=False)
k_param = wd.FloatSlider(value=2, min=0.5, max=4, step=0.5,
description='k:', continuous_update=False)
bollinger_box = wd.VBox(children=[bb_label, n_param, k_param])
# + id="o8XsAdOZxCJr"
macd_label = wd.Label('MACD')
macd_fast = wd.IntSlider(value=12, min=2, max=50, step=1,
description='Fast avg:',
continuous_update=False)
macd_slow = wd.IntSlider(value=26, min=2, max=50, step=1,
description='Slow avg:',
continuous_update=False)
macd_signal = wd.IntSlider(value=9, min=2, max=50, step=1,
description='MACD signal:',
continuous_update=False)
macd_box = wd.VBox(children=[macd_label, macd_fast,
macd_slow, macd_signal])
# + id="D6Q1uq5AxPFn"
rsi_label = wd.Label('RSI')
rsi_periods = wd.IntSlider(value=14, min=2, max=50, step=1,
description='RSI periods:',
continuous_update=False)
rsi_upper = wd.IntSlider(value=70, min=1, max=100, step=1,
description='Upper Thr:',
continuous_update=False)
rsi_lower = wd.IntSlider(value=30, min=1, max=100, step=1,
description='Lower Thr:',
continuous_update=False)
rsi_box = wd.VBox(children=[rsi_label, rsi_periods,
rsi_upper, rsi_lower])
# + id="1gKIcmgXxUzZ"
sec_selector_label = wd.Label('Secondary parameters',
layout=wd.Layout(height='45px'))
blank_label = wd.Label('', layout=wd.Layout(height='45px'))
sec_box_1 = wd.VBox([sec_selector_label, bollinger_box, macd_box])
sec_box_2 = wd.VBox([blank_label, rsi_box])
secondary_selector_box = wd.HBox([sec_box_1, sec_box_2])
# + id="BlNLpcklxaRP"
controls_dict = {'asset':stocks_selector,
'indicator':indicator_selector,
'start_date':start_date_selector,
'end_date':end_date_selector,
'bb_k':k_param,
'bb_n':n_param,
'macd_fast': macd_fast,
'macd_slow': macd_slow,
'macd_signal': macd_signal,
'rsi_periods': rsi_periods,
'rsi_upper': rsi_upper,
'rsi_lower': rsi_lower}
ui = wd.HBox([main_selector_box, secondary_selector_box])
out = wd.interactive_output(ta_dashboard, controls_dict)
# + id="IVGt8IEPxgF7" outputId="00190acf-03c7-424a-ff4f-480bc0ec6647" colab={"base_uri": "https://localhost:8080/", "height": 620, "referenced_widgets": ["02f0b23eddde47d780502804a364e9d8", "50c2c60cd4bb44b5a8af7c185e023a18", "2902d957b9ed40d5b9bc773208a6bb9a", "f4015a452bd343029ebbc0cf7594d01d", "1bd5a556b034404a85055950fce2f5eb", "67721c80165e4f74a3da17633ca868e8", "7d8559e327344223b4651a9e2412ef84", "f9c0efcf6a834305be679d8f3bfdfdde", "6e1dbe3caa684148b9bc90a5783ffc65", "5c14484265334fe6965f2fe104cdd683", "8cefd201352d497e864e82d4626dda9d", "f8f85ed1413a4dcc9db9a2c6931f39f8", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "03ed0cfd5183448aa6169c34a284233b", "<KEY>", "162464b7ca294c83adafa07c6b6dd8a8", "6b80e684c8764f46b40b3b5a61dfa24d", "d985ccb3b2e24d948dcef1614bc03166", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "ff67f8c0e0f644fb8300973648d9f845", "<KEY>", "a82cfb3338be46d0b4684eec2cc95838", "<KEY>", "4d9ed44aeedd4db7a07b7c64e886e4bd", "<KEY>", "8f772efc271b4e1f949f40a4b4e48ffd", "<KEY>", "<KEY>", "4041eb6d84d648b1ac4738543184e18a", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "9b2e6fd872e74ead923f2023b71d2563", "<KEY>", "53f9976f87804740891f3e1650a0a401", "<KEY>", "<KEY>", "3d593683c3124af485125fc205ff7c5c", "e265d3051832494e8efa200081953774", "c8c9625007144c5ca922052ee877e5db", "430442ba293342ffad6e1b1d59dc00eb", "<KEY>", "f885bf731a0842e082ace7397ea1a9d5", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "ba17658bb7f2451aa011c43a4de18614", "<KEY>", "d35d990e501e43f0bdcedf5086adf31c", "<KEY>", "e8efba599d4e454db9ee93bf20dc7868", "9a9c98a67bad48b28db2ba2cf339a573", "<KEY>", "<KEY>", "fe49ba8ac03a4b04b4885d78bbcb7ed7", "<KEY>", "<KEY>", "c6ef1f2f70fe411e864fc7cdfedbb9f5"]}
display(ui, out)
# + id="xUmnnjC5A9Kp"
| Project 10_How_to_Create_a_Technical_Analysis_Interactive_Dashboard.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="ZMFqt2-EYnLI"
# Install captum package
# !pip install captum
# + id="-J9A0V63qLeW"
import pandas as pd
import numpy as np
import random
from utils import preprocessing, SlidingWindow, NegativeSampling, utils, model, explain
from sklearn.model_selection import train_test_split
import torch
import torch.nn as nn
import torch.optim as optim
from tqdm import tqdm
import time
import math
import os
from sklearn import metrics
from sklearn.metrics import precision_recall_fscore_support
from captum.attr import LayerIntegratedGradients
import collections
# + id="v7vbfwlP8IfJ"
DATASET_NAME = 'Thunderbird'
TRAIN_SIZE = 100000
WINDOW_SIZE = 100
STEP_SIZE = 20
RATIO = 0.1
SEED = 42
# + id="yerXXxqJqetY"
# Download dataset and parsing the dataset with Drain
preprocessing.parsing(DATASET_NAME)
# + id="ERsjmU1r5Zo9"
# Cut log data into sliding windows
# Split data into training normal dataset, test normal dataset, and test abnormal dataset
# Get the bigram from training normal dataset
# Train a Word2Vec model with the training normal data
random.seed(SEED)
np.random.seed(SEED)
train_normal, test_normal, test_abnormal, bigram, unique, weights, train_dict, w2v_dic = SlidingWindow.sliding(DATASET_NAME, WINDOW_SIZE, STEP_SIZE, TRAIN_SIZE)
# + id="CSsRoGViaEpG"
# # +1 for unknown
VOCAB_DIM = len(train_dict)+1
OUTPUT_DIM = 2
EMB_DIM = 8
HID_DIM = 256
N_LAYERS = 1
DROPOUT = 0.0
BATCH_SIZE = 16
TIMES = 20
# + id="L8HIfgwvblO5"
# Get negative samples and split into training data and val data
random.seed(SEED)
np.random.seed(SEED)
neg_samples = NegativeSampling.negative_sampling(train_normal, bigram, unique, TIMES, VOCAB_DIM)
df_neg = utils.get_dataframe(neg_samples, 1, w2v_dic)
df_pos = utils.get_dataframe(list(train_normal['EventId']), 0, w2v_dic)
df_pos.columns = df_pos.columns.astype(str)
df_train = pd.concat([df_pos, df_neg], ignore_index = True, axis=0)
df_train.reset_index(drop = True)
y = list(df_train.loc[:,'class_label'])
X = list(df_train['W2V_EventId'])
X_train, X_val, y_train, y_val = train_test_split(X, y, test_size=0.2, random_state=42)
X_train = torch.tensor(X_train,requires_grad=False).long()
X_val = torch.tensor(X_val,requires_grad=False).long()
y_train = torch.tensor(y_train).reshape(-1, 1).long()
y_val = torch.tensor(y_val).reshape(-1, 1).long()
train_iter = utils.get_iter(X_train, y_train, BATCH_SIZE)
val_iter = utils.get_iter(X_val, y_val, BATCH_SIZE)
# + id="zUEk54oerMa5"
random.seed(SEED)
np.random.seed(SEED)
torch.manual_seed(SEED)
torch.cuda.manual_seed(SEED)
torch.backends.cudnn.deterministic = True
device = torch.device( "cuda" if torch.cuda.is_available() else"cpu")
interpretableSAD = model.C_lstm(weights, VOCAB_DIM, OUTPUT_DIM, EMB_DIM, HID_DIM, N_LAYERS, DROPOUT, device, BATCH_SIZE).to(device)
print(f'The model has {model.count_parameters(interpretableSAD):,} trainable parameters')
print()
optimizer = optim.Adam(interpretableSAD.parameters())
criterion = nn.CrossEntropyLoss()
try:
os.makedirs('Model')
except:
pass
#Training interpretableSAD
N_EPOCHS = 10
CLIP = 1
best_test_loss = float('inf')
for epoch in tqdm(range(N_EPOCHS)):
start_time = time.time()
train_loss= model.train(interpretableSAD, train_iter, optimizer, criterion, CLIP, epoch, device)
val_loss = model.evaluate(interpretableSAD, val_iter, criterion, device)
end_time = time.time()
epoch_mins, epoch_secs = model.epoch_time(start_time, end_time)
if val_loss < best_test_loss:
best_test_loss = val_loss
torch.save(interpretableSAD.state_dict(), 'Model/interpretableSAD_Thunderbird.pt')
print(f'Epoch: {epoch+1:02} | Time: {epoch_mins}m {epoch_secs}s')
print(f'\tTrain Loss: {train_loss:.3f} | Train PPL: {math.exp(train_loss):7.3f}')
print(f'\t Val. Loss: {val_loss:.3f} | Val. PPL: {math.exp(val_loss):7.3f}')
# + id="pQi1_5zWWYZB"
# less or equal than 10% abnormal data
test_abnormal_ratio = model.ratio_abnormal_sequence(test_abnormal, WINDOW_SIZE, RATIO)
test_ab_X, test_ab_X_key_label = test_abnormal_ratio['W2V_EventId'], test_abnormal_ratio['Key_label']
test_n_X, test_n_X_key_label = test_normal['W2V_EventId'], test_normal['Key_label']
test_ab_y = test_abnormal_ratio['Label']
test_n_y = test_normal['Label']
y, y_pre = model.model_precision(interpretableSAD, device, test_n_X.values.tolist()[:int(len(test_n_X.values.tolist())*(len(test_abnormal_ratio)/len(test_abnormal)))], \
test_ab_X.values.tolist())
f1_acc = metrics.classification_report(y, y_pre, digits=5)
print(f1_acc)
# + id="8mHTspdaYq2w"
lig = LayerIntegratedGradients(interpretableSAD, interpretableSAD.embedding)
lst_train_keys = []
for i in train_normal.W2V_EventId.values:
lst_train_keys.extend(i)
dic_app = collections.Counter(lst_train_keys)
if w2v_dic[str(len(train_dict))] not in dic_app.keys():
dic_app[w2v_dic[str(len(train_dict))]] = 0
start = [w2v_dic[i] for i in unique]
lst_attr, lst_y, lst_dist, lst_keys, lst_baseline = explain.get_dataset(interpretableSAD, device, lig, test_ab_X, test_ab_X_key_label, dic_app, start, RATIO, WINDOW_SIZE)
# + id="jx0SnQgqYs0R"
exp_df = pd.DataFrame()
exp_df['key'] = lst_keys
exp_df['attr'] = lst_attr
exp_df['y'] = lst_y
exp_df['baseline'] = lst_baseline
best_inter = explain.get_mean_inter(exp_df)
# Zero as inter
mean_pred = explain.mean_inter(exp_df)
print("Accuracy for zero inter:",metrics.accuracy_score(lst_y, mean_pred))
print(metrics.classification_report(lst_y, mean_pred, digits=5))
# Best inter
mean_pred = explain.mean_inter(exp_df, interception=best_inter)
print("Accuracy for the best inter:",metrics.accuracy_score(lst_y, mean_pred))
print(metrics.classification_report(lst_y, mean_pred, digits=5))
| InterpretableSAD_Thunderbird.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="izaAYfDGveRj"
# # **Introduction to Tensorflow**
#
#
# <img src = "https://miro.medium.com/max/1025/1*vWsIxYG3EkR7C_lsziqPFQ.png">
#
# ## **What is Tensor?**
#
# A tensor is an N-dimensional array of data
#
# <img src = "https://raw.githubusercontent.com/mhuzaifadev/ml_zero_to_hero/master/tensor.png?token=<KEY>">
#
# + [markdown] id="fKH1OqniyoLU"
# ## **Components of TensorFlow: Tensors and Variables**
# + colab={"base_uri": "https://localhost:8080/"} id="YdiTAFMqy21a" outputId="078a9b90-55a7-4cff-9ebd-0ea89348ed8f"
#Importing Tensorflow
import tensorflow as tf
print(tf.__version__)
import numpy as np
print(np.__version__)
# + [markdown] id="QMi_8GEWzf7i"
# **Producing Constant Tensors**
# + colab={"base_uri": "https://localhost:8080/"} id="xRew53kesJXo" outputId="c3c51947-35f0-4ea0-9eda-b26bfcaaa4b1"
x = tf.constant([
[3,6,9],
[6,8,11]
])
x
# + [markdown] id="aWCJYXq1zlW5"
# **Tensors can be sliced**
# + colab={"base_uri": "https://localhost:8080/"} id="iaMgRjp4zlEP" outputId="0677546d-8ba8-43ab-fcc0-4a8cb1e5f1a0"
y = x[:,1]
y
# + [markdown] id="TvjyqmeqzyW_"
# **Tensors can be reshaped**
# + colab={"base_uri": "https://localhost:8080/"} id="LcXA-aNqz5qH" outputId="fe00d12a-7c6f-47a3-c697-8da8026230fd"
y = tf.reshape(x,[3,2])
y
# + [markdown] id="Q_QV4A-y6NVQ"
# **Here is a "scalar" or "rank-0" tensor . A scalar contains a single value, and no "axes".**
# + colab={"base_uri": "https://localhost:8080/"} id="agBm1YDH6NjN" outputId="b244af27-e8fa-479b-ef60-451c029dedd8"
# This will be an int32 tensor by default; see "dtypes" below.
rank_0_tensor = tf.constant(4)
print(rank_0_tensor)
# + [markdown] id="wPMnFMEq6U_n"
# **A "vector" or "rank-1" tensor is like a list of values. A vector has 1-axis:**
# + colab={"base_uri": "https://localhost:8080/"} id="aTJWv07b6VJB" outputId="d07bbf37-6a9e-465d-a578-96d79d0bf516"
# Let's make this a float tensor.
rank_1_tensor = tf.constant([2.0, 3.0, 4.0])
print(rank_1_tensor)
# + [markdown] id="Xberh3lo6bIw"
# **A "matrix" or "rank-2" tensor has 2-axes:**
# + colab={"base_uri": "https://localhost:8080/"} id="0il8RXrK6ghK" outputId="f0ca98bf-c319-4a80-85c4-53f3128c8492"
# If we want to be specific, we can set the dtype (see below) at creation time
rank_2_tensor = tf.constant([[1, 2],
[3, 4],
[5, 6]], dtype= float)
print(rank_2_tensor)
# + [markdown] id="C3TWseR87R19"
# <img src = "https://raw.githubusercontent.com/mhuzaifadev/ml_zero_to_hero/master/tens.png?token=<KEY>">
# + [markdown] id="JSQ-oXux7Yj_"
# **Tensors may have more axes, here is a tensor with 3-axes:**
# + colab={"base_uri": "https://localhost:8080/"} id="MmNA-C097dDD" outputId="cebb7580-92dd-4114-ab1a-6c411d39a732"
# There can be an arbitrary number of
# axes (sometimes called "dimensions")
rank_3_tensor = tf.constant([
[[0, 1, 2, 3, 4],
[5, 6, 7, 8, 9]],
[[10, 11, 12, 13, 14],
[15, 16, 17, 18, 19]],
[[20, 21, 22, 23, 24],
[25, 26, 27, 28, 29]],])
print(rank_3_tensor)
# + [markdown] id="SHQe4_4m8AOX"
# <img src ="https://raw.githubusercontent.com/mhuzaifadev/ml_zero_to_hero/master/3axistensor.png?token=<KEY>">
#
# + [markdown] id="trbjnRZK8mXf"
# **You can convert a tensor to a NumPy array either using np.array or the tensor.numpy method:**
# + colab={"base_uri": "https://localhost:8080/"} id="ad_kV2Hq8p_j" outputId="a91c7998-d4ea-4685-f369-45f9b9465d61"
# Convert a tensor to a NumPy array using `np.array` method
np.array(rank_2_tensor)
# + colab={"base_uri": "https://localhost:8080/"} id="UuHGLNID8qDx" outputId="fcdf8a0a-5757-4479-d551-106793f0e687"
# Convert a tensor to a NumPy array using `tensor.numpy` method
rank_2_tensor.numpy
# + [markdown] id="jpqj971898o5"
# **We can do basic math on tensors, including addition, element-wise multiplication, and matrix multiplication.**
# + colab={"base_uri": "https://localhost:8080/"} id="m3OMptRh97SA" outputId="31e3cf91-0698-49cd-f32b-8276e402db2f"
a = tf.constant([[1, 2],
[3, 4]])
b = tf.constant([[1, 1],
[1, 1]]) # Could have also said `tf.ones([2,2])`
print(tf.add(a, b), "\n")
print(tf.multiply(a, b), "\n")
print(tf.matmul(a, b), "\n")
# + colab={"base_uri": "https://localhost:8080/"} id="2phbMZjB-LMc" outputId="6d7bb41a-43f7-4a31-d527-847ca237fb14"
print(a + b, "\n") # element-wise addition
print(a * b, "\n") # element-wise multiplication
print(a @ b, "\n") # matrix multiplication
# + [markdown] id="Sw19Ep4k-8vV"
# **Tensors are used in all kinds of operations (ops).**
# + colab={"base_uri": "https://localhost:8080/"} id="Tf3bny3V-_Sv" outputId="04b7ee04-2ff7-4285-915d-88f05c24acfc"
c = tf.constant([[4.0, 5.0], [10.0, 1.0]])
# Find the largest value
print(tf.reduce_max(c))
# Find the index of the largest value
print(tf.math.argmax(c))
#Compute the Softmax
tf.nn.softmax(c)
# + [markdown] id="v1_d-cAWAT2D"
# ## **About Shapes**
#
# Tensors have shapes. Some vocabulary:
#
# * **Shape**: The length (number of elements) of each of the dimensions of a tensor.
# * **Rank**: Number of tensor dimensions. A scalar has rank 0, a vector has rank 1, a matrix is rank 2.
# * **Axis** or **Dimension**: A particular dimension of a tensor.
# * **Size**: The total number of items in the tensor, the product shape vector
#
# Note: Although you may see reference to a "tensor of two dimensions", a rank-2 tensor does not usually describe a 2D space.
#
# Tensors and `tf.TensorShape` objects have convenient properties for accessing these:
# + id="SEV8sI7kAZmV"
rank_4_tensor = tf.zeros([3, 2, 4, 5])
# + colab={"base_uri": "https://localhost:8080/"} id="BPBUQkpJBEjn" outputId="d065fb1a-47e6-4e90-fd0f-21c113cc1e73"
rank_4_tensor
# + [markdown] id="IW2KHCkIA3P1"
# <img src = "https://raw.githubusercontent.com/mhuzaifadev/ml_zero_to_hero/master/shapes.png?token=<KEY>">
#
# + colab={"base_uri": "https://localhost:8080/"} id="Nn7UdITqA8Dh" outputId="180ea7f1-6565-473b-cde4-cf6b8ba9b3d6"
print("Type of every element:", rank_4_tensor.dtype)
print("Number of dimensions:", rank_4_tensor.ndim)
print("Shape of tensor:", rank_4_tensor.shape)
print("Elements along axis 0 of tensor:", rank_4_tensor.shape[0])
print("Elements along the last axis of tensor:", rank_4_tensor.shape[-1])
print("Total number of elements (3*2*4*5): ", tf.size(rank_4_tensor).numpy())
# + [markdown] id="ZweEVM6WBPuk"
# While axes are often referred to by their indices, you should always keep track of the meaning of each. Often axes are ordered from global to local: The batch axis first, followed by spatial dimensions, and features for each location last. This way feature vectors are contiguous regions of memory.
#
# <img src = "https://raw.githubusercontent.com/mhuzaifadev/ml_zero_to_hero/master/rank4.png?token=<KEY>">
# + [markdown] id="fJ0Hiqg8B--a"
# ### **Single-axis indexing**
#
# TensorFlow follow standard python indexing rules, similar to indexing a list or a string in python, and the bacic rules for numpy indexing.
#
# indexes start at 0
# negative indices count backwards from the end
# colons, :, are used for slices start:stop:step
# + colab={"base_uri": "https://localhost:8080/"} id="kg4B58knCR5P" outputId="dd0c0a17-19a1-403a-dbb6-970b3df108bc"
rank_1_tensor = tf.constant([0, 1, 1, 2, 3, 5, 8, 13, 21, 34])
print(rank_1_tensor.numpy())
# + [markdown] id="obw5_VaNCVPI"
# **Indexing with a scalar removes the dimension:**
# + colab={"base_uri": "https://localhost:8080/"} id="jh6lkmGxCVZB" outputId="a001af75-871a-4cf5-ee4c-09e04b9f13dd"
print("First:", rank_1_tensor[0].numpy())
print("Second:", rank_1_tensor[1].numpy())
print("Last:", rank_1_tensor[-1].numpy())
# + [markdown] id="cR8noAQvCbXF"
# **Indexing with a : slice keeps the dimension:**
# + colab={"base_uri": "https://localhost:8080/"} id="aiKTcvkJCbeI" outputId="547e4da4-44c5-4017-9f83-4ea1ada59187"
print("Everything:", rank_1_tensor[:].numpy())
print("Before 4:", rank_1_tensor[:4].numpy())
print("From 4 to the end:", rank_1_tensor[4:].numpy())
print("From 2, before 7:", rank_1_tensor[2:7].numpy())
print("Every other item:", rank_1_tensor[::2].numpy())
print("Reversed:", rank_1_tensor[::-1].numpy())
# + [markdown] id="wRmICBN-CjbU"
# ### **Multi-axis indexing**
#
# Higher rank tensors are indexed by passing multiple indices.
#
# The single-axis exact same rules as in the single-axis case apply to each axis independently.
# + colab={"base_uri": "https://localhost:8080/"} id="F0kK7yxZCofw" outputId="b6023210-a54c-4f80-ff9c-b9977b2265a9"
print(rank_2_tensor.numpy())
# + [markdown] id="scCer4pZCtQQ"
# **Passing an integer for each index the result is a scalar.**
# + colab={"base_uri": "https://localhost:8080/"} id="WUUmKyx1CtYc" outputId="6d404ecb-dec7-420b-b0cf-75c32771a7b4"
# Pull out a single value from a 2-rank tensor
print(rank_2_tensor[1, 1].numpy())
# + [markdown] id="nD1iVpj7Ctf9"
# **You can index using any combination integers and slices:**
# + colab={"base_uri": "https://localhost:8080/"} id="XngWv5NzCtoC" outputId="33ea80ae-4929-4cd3-ca46-5c792d60037b"
# Get row and column tensors
print("Second row:", rank_2_tensor[1, :].numpy())
print("Second column:", rank_2_tensor[:, 1].numpy())
print("Last row:", rank_2_tensor[-1, :].numpy())
print("First item in last column:", rank_2_tensor[0, -1].numpy())
print("Skip the first row:")
print(rank_2_tensor[1:, :].numpy(), "\n")
# + [markdown] id="ftMQMpT2DBva"
# **Here is an example with a 3-axis tensor:**
# + colab={"base_uri": "https://localhost:8080/"} id="keum4kK2DFUf" outputId="1106ffe3-4d44-418b-fdd0-d1baf122e8ca"
print(rank_3_tensor[:, :, 4])
# + [markdown] id="rd7GhdYkDZtQ"
# <img src = "https://raw.githubusercontent.com/mhuzaifadev/ml_zero_to_hero/master/rank3.png?token=<KEY>">
# + [markdown] id="LOJEUbqZDiva"
# ## **Manipulating Shapes**
#
# Reshaping a tensor is of great utility.
#
# The `tf.reshape` operation is fast and cheap as the underlying data does not need to be duplicated.
# + colab={"base_uri": "https://localhost:8080/"} id="d558p1uGDmyk" outputId="bfe0e7d3-1b87-4467-dbc5-eb20d7a7a7e9"
# Shape returns a `TensorShape` object that shows the size on each dimension
var_x = tf.Variable(tf.constant([[1], [2], [3]]))
print(var_x.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="G_ocJiOzDtnw" outputId="a15ca846-d877-4dd6-ae38-2878e22c068e"
# You can convert this object into a Python list, too
print(var_x.shape.as_list())
# + [markdown] id="esBMCLPnDxrQ"
# **You can reshape a tensor into a new shape. Reshaping is fast and cheap as the underlying data does not need to be duplicated.**
# + id="aKsvSW7eDzNi"
# We can reshape a tensor to a new shape.
# Note that we're passing in a list
reshaped = tf.reshape(
var_x, [1,3]
)
# + colab={"base_uri": "https://localhost:8080/"} id="mr4NIh5_FTMq" outputId="581ce6bd-c482-4de3-d7fe-5de6dc432540"
print(var_x.shape)
print(reshaped.shape)
# + [markdown] id="7ZqAEYGVFYbc"
# The data maintains it's layout in memory and a new tensor is created, with the requested shape, pointing to the same data. TensorFlow uses C-style "row-major" memory ordering, where incrementing the right-most index corresponds to a single step in memory.
# + colab={"base_uri": "https://localhost:8080/"} id="1t2NKCQhFbYg" outputId="17d9d070-2586-4c32-b379-bad043f6d485"
print(rank_3_tensor)
# + [markdown] id="vOWQyNH3Fgp3"
# If you flatten a tensor you can see what order it is laid out in memory.
# + colab={"base_uri": "https://localhost:8080/"} id="NjffQkgCFihA" outputId="dd7a7e8c-e857-4211-8ec7-3d63e67c9cdb"
# A `-1` passed in the `shape` argument says "Whatever fits".
print(tf.reshape(rank_3_tensor, [-1]))
# + [markdown] id="yoJZvjvbFn4D"
# Typically the only reasonable uses of `tf.reshape` are to combine or split adjacent axes (or add/remove `1`s).
#
# For this 3x2x5 tensor, reshaping to (3x2)x5 or 3x(2x5) are both reasonable things to do, as the slices do not mix:
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="COeVVOb8Fpyd" outputId="5c595fc7-e9e5-40eb-8b1f-9c9fbd517190"
print(tf.reshape(rank_3_tensor, [3*2, 5]), "\n")
print(tf.reshape(rank_3_tensor, [3, -1]))
# + [markdown] id="PrSfJuVUF_F4"
# <img src = "https://raw.githubusercontent.com/mhuzaifadev/ml_zero_to_hero/master/concepts.png?token=<KEY>">
# + [markdown] id="rBgDJCKQGCq0"
# Reshaping will "work" for any new shape with the same total number of elements, but it will not do anything useful if you do not respect the order of the axes.
#
# Swapping axes in `tf.reshape` does not work, you need `tf.transpose` for that.
# + [markdown] id="WBRGwUl5HECc"
# ## **Broadcasting**
#
# Broadcasting is a concept borrowed from the equivalent feature in NumPy. In short, under certain conditions, smaller tensors are "stretched" automatically to fit larger tensors when running combined operations on them.
#
# The simplest and most common case is when you attempt to multiply or add a tensor to a scalar. In that case, the scalar is broadcast to be the same shape as the other argument.
# + colab={"base_uri": "https://localhost:8080/"} id="PPe6adxOGHSC" outputId="059dfe24-882d-48cf-9432-7fb19aa65a3c"
x = tf.constant([1, 2, 3])
y = tf.constant(2)
z = tf.constant([2, 2, 2])
# All of these are the same computation
print(tf.multiply(x, 2))
print(x * y)
print(x * z)
# + [markdown] id="nSUdF0lZHrDp"
# Likewise, 1-sized dimensions can be stretched out to match the other arguments. Both arguments can be stretched in the same computation.
#
# In this case a 3x1 matrix is element-wise multiplied by a 1x4 matrix to produce a 3x4 matrix. Note how the leading 1 is optional: The shape of y is [4].
# + colab={"base_uri": "https://localhost:8080/"} id="3UyCt6qMHuAz" outputId="4104286d-fc7a-4ef8-e4a4-c775010df08f"
# These are the same computations
x = tf.reshape(x,[3,1])
y = tf.range(1, 5)
print(x, "\n")
print(y, "\n")
print(tf.multiply(x, y))
# + [markdown] id="-V-eeRYWIPdb"
# <img src = "https://raw.githubusercontent.com/mhuzaifadev/ml_zero_to_hero/master/broadcast.png?token=AM<PASSWORD>">
#
#
#
# Here is the same operation without broadcasting:
# + colab={"base_uri": "https://localhost:8080/"} id="dqzkJDVEIUJy" outputId="fb5922b6-8abd-4a37-9214-5e4928e45508"
x_stretch = tf.constant([[1, 1, 1, 1],
[2, 2, 2, 2],
[3, 3, 3, 3]])
y_stretch = tf.constant([[1, 2, 3, 4],
[1, 2, 3, 4],
[1, 2, 3, 4]])
print(x_stretch * y_stretch) # Again, operator overloading
# + [markdown] id="A4DCP9khIceL"
# Most of the time, broadcasting is both time and space efficient, as the broadcast operation never materializes the expanded tensors in memory.
#
# You see what broadcasting looks like using `tf.broadcast_to.`
# + colab={"base_uri": "https://localhost:8080/"} id="pRUGKYf_Ijn0" outputId="0742de30-94d7-4a29-9af3-5e3f28cbb178"
print(tf.broadcast_to(tf.constant([1, 2, 3]), [3, 3]))
# + [markdown] id="5-xjjDdLIr-F"
# Unlike a mathematical op, for example, `broadcast_to` does nothing special to save memory. Here, you are materializing the tensor.
# + [markdown] id="S_Xki5rg2E36"
# **This is it for today**
| 02 Introduction_to_Tensorflow/02_Introduction_to_TensorFlow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="0vt2fPA4-vLs" colab={"base_uri": "https://localhost:8080/", "height": 36} outputId="8adfa825-9ecb-4e25-b84c-1c95febd5e8e"
import keras
keras.__version__
# + [markdown] id="zzJZJwplA09_"
# # CNN
#
# 05/2020
#
# # Resumo
#
# Neste notebook discutiremos o que é uma Rede Neural Convolucional, construiremos uma baseados nos conceitos aprendidos, e a utilizaremos para resolver um problema de classificação binária.
#
# # Sumário
#
# <ol>
# <li>1.Introdução</li>
# <li>2.Detalhes da Arquitetura</li>
# <li>3.Problema: Classificação de Imagens</li>
# <li>4.Bibliografia</li>
# </ol>
#
# ## 1.Introdução
#
# Apesar de todas as dificuldades provenientes de se treinar Redes Neurais com muitas camadas intermediárias, ao longo dos anos foram desenvolvidas algumas técnicas para que se pudesse treinar esse tipo de Rede. Na área de reconhecimento e processamento de imagens, o tipo de rede neural que tem dominado as recentes pesquisas é a Rede Neural Convolucional (ou CNN da sigla em inglês) cuja arquitetura é desenvolvida de modo que seja possível treinar as redes de maneira eficiente.
#
# ## 2.Detalhes da Arquitetura
#
#
# ### 2.1.Local Receptive Fields
#
# Diferente das Redes Neurais comuns, em que as camadas eram interpretadas como uma linha vertical de neurônios, nas Redes Neurais Convolucionais, as camadas de entrada devem ser interpretadas como um quadrado ou retângulo de neurônios como na imagem a seguir:
#
#
# 
#
# No problema do MNIST, por exemplo, todas as imagens estão em escala de cinza em dimensão 28x28, portanto a camada de entrada de uma CNN que resolva esse problema deve ser quadrada de dimensão também 28x28.
#
# Contudo, nem todo conjunto de imagens é tão uniforme, é comum que as imagens estejam em dimensões diferentes umas das outras. Por isso é uma prática comum re-escalar as imagens no pré-processamento de modo que todas fiquem com dimensões iguais às da camada de entrada da CNN.
#
# Diferente das camadas densas, onde todos os neurônios de uma camada são conectados com todos os neurônios da próxima camada, nas redes neurais convolucionais, cada neurônio da primeira camada intermediária conecta um número limitado de neurônios da camada de entrada. Para exemplificar será utilizada uma ilustração:
#
# 
#
# Podemos ver que um neurônio está conectado a um quadrado 5x5 de neurônios na entrada, essa pequena região que é apenas um pedaço da imagem original é chamado de *Campo Receptivo Local*. O neurônio da um peso para conexão que faz na entrada para aprendizagem, e ainda tem um bias. É possível interpretar isso como se o neurônio estivesse "aprendendo a analisar apenas seu próprio Campo Receptivo".
#
# Para completar a próxima camada da CNN é como se os Campos Receptivos Locais "deslizassem" pela imagem até completá-la, gerando assim um neurônio na primeira camada intermediária para cada Campo Receptivo, seguindo a próxima imagem:
#
# 
#
# Seguindo assim até que todos os Campos Receptivos da imagem sejam contemplados, podemos ver que para 28x28 neurônios de entrada teremos apenas 24x24 na próxima camada, isso se dá pelo fato de utilizarmos um tamanho para os Campos Receptivos de 5x5.
#
# ### 2.2.Feature Maps
#
# Um detalhe crucial para a arquitetura de uma CNN é que, por mais que cada neurônio de uma camada intermediária esteja conectado a apenas um pequeno pedaço da imagem, todos eles compartilham os mesmos pesos e bias. Sendo assim cada um dos 24x24 neurônios do exemplo que estamos construindo treinarão exatamente os mesmos pesos e bias. A saída de um neurônio qualquer dessa camada pode ser escrita por:
#
# $$ \sigma \left(b + \sum \limits _{l = 0} ^{4} \sum \limits _{m = 0} ^{4} w_{l, m} a_{j+l,k+m} \right) (1)$$
#
# Aqui seguindo a linguagem do Nielsen<sup>[1]</sup>, $\sigma$ é a função de ativação da rede neural, $b$ é o valor do bias e $w_{l, m}$ é conjunto 5x5 de pesos, todos compartilhados. Além disso, é utilizado $a_{x,y}$ para denotar a ativação do neurônio da posição $(x,y)$ da camada de entrada.
#
# Esse método de compartilhamento de pesos e bias leva a um atributo interessante, que é que todos os neurônios desta camada estão aprendendo a analisar o mesmo recurso (ou feature) apenas em locais diferentes da imagem.
#
# Por esse motivo, o mapeamento dos dados da entrada para a camada intermediária comummente recebe o nome de *Mapa de Recursos* (ou feature map). Como cada feature map só faz a detecção de um recurso da imagem, para fazer um reconhecimento eficiente de imagens, precisamos de mais de um feature map.
#
#
# 
#
# O exemplo acima tem 3 mapas de recursos, cada qual com 5x5 pesos compartilhados e 1 bias. Isso significa que essa Rede pode detectar 3 diferentes tipos de features, cada uma podendo ser identificada ao longo de toda imagem. Apesar da imagem mostrar apenas 3 features, frequentemente se utiliza muito mais feature maps em uma só camada da Rede Neural.
#
# ### 2.3.Pooling Layers
#
# Outro elemento presente na arquitetura das Redes Neurais Convolucionais são as chamadas *Camadas de Agrupamento* (ou Pooling Layer), essas camadas habitualmente estão presentes depois de uma camada de Mapa de Recursos, e seu papel é simplificar a informação presente na Camada Convolucional.
#
# Cada Camada de Agrupamento pega a saída de um determinado Feature Map e gera uma versão condensada do mesmo, como se sintetizasse suas informações em um Feature Map menor. Por exemplo, cada unidade da Pooling Layer pode sintetizar uma região de 2x2 ou 3x3 da camada anterior, transformando toda essa informação em apenas um neurônio.
#
# Existem várias maneiras de fazer essa síntese de informação, mas a maneira mais comum, que também sera utilizada neste notebook, é a chamada de Max-Pooling. Ela pega a região a ser condensada, e tem como saída o maior valor de ativação da mesma.
#
# Como existem vários Feature Maps em uma só CNN, se aplica uma Pooling Layer para cada Feature Map, como na imagem a seguir:
#
# 
#
# ### 2.4.Arquitetura Final
#
# Agora que sabemos todos os artefatos de uma Rede Neural Convolucional, podemos juntar todas as partes e verificar como é a estrutura de uma CNN completa. Para isso podemos simplesmente ligar todas as saídas da Max Pooling Layer e conectar densamente à camada de saída da nossa Rede Neural da seguinte forma:
#
# 
#
# A camada de saída da Rede Neural é idêntica às Redes Neurais rasas, pode ter ativação do tipo SOFTMAX para classificação por exemplo, ou sigmóide caso o problema seja de classificação binária.
#
# É comum ainda fazer adições a essa estrutura que criamos, como por exemplo adicionar mais Camadas Convolucionais seguidas de Pooling Layers, ou adicionar uma Camada Densa de Neurônios (completamente conectada), antes da camada de saída. Não há regra para essas alterações e elas devem se adequar ao problema atacado.
#
#
#
#
#
#
#
#
#
#
# + [markdown] id="dXqq_CEOXBCk"
# ## 3.Problema: Classificação de Imagens
#
# Para exemplificar a utilização das CNNs nesse notebook, resolveremos um problema que seria difícil de lidar de outra maneira. Esse problema é o de classificação de imagens de Cachorros e Gatos. O objetivo é aparentemente simples, devemos classificar imagens de cachorros e gatos, para isso utilizaremos o dataset "Dogs_Vs_Cats" presente no Kaggle, com mais de 25 mil imagens que são parecidas com as seguintes:
#
# 
#
#
# Vamos treinar uma Rede Neural Convolucional para que ela seja capaz de, quando receber uma imagem, classificar se nela está presente um cachorro, ou um gato. Será utilizada apenas uma parte do dataset, com 20 mil imagens de treino e 2 mil para validação.
# + [markdown] id="k6NRS0P_Z9BU"
# O primeiro passo é fazer a instalação do kaggle:
# + id="AsVhjkcqaJYA" colab={"base_uri": "https://localhost:8080/"} outputId="ebe62862-0234-422a-ed21-759941dcf5ba"
# !pip install kaggle
# + [markdown] id="VIBPDdkuaKHp"
# Depois devemos fazer o upload do API Token (obtida no site do kaggle), para o computador remoto, para isso podemos utilizar esse código:
# + id="gColD0n1zsM5" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": "OK"}}, "base_uri": "https://localhost:8080/", "height": 92} outputId="40dfe794-15ed-4052-d887-9fa83b2f5f53"
from google.colab import files
uploaded = files.upload()
for fn in uploaded.keys():
print('User uploaded file "{name}" with length {length} bytes'.format(
name=fn, length=len(uploaded[fn])))
# + [markdown] id="k4DwTp1paog2"
# Então podemos finalmente baixar o dataset direto do Kaggle, e descomprimí-lo com:
#
#
#
# + id="lIpJwBhjA30J" colab={"base_uri": "https://localhost:8080/"} outputId="5ed5a6b3-68b9-4bb3-bbc2-74b553acfc82"
# !mkdir $HOME/.kaggle
# !mv kaggle.json $HOME/.kaggle/kaggle.json
# !chmod 600 $HOME/.kaggle/kaggle.json
# ! kaggle competitions download -c 'dogs-vs-cats'
# + id="nlfSdtiaeuSr"
# !unzip -qq /content/train.zip -d /datalab/
# + [markdown] id="kZdAficfcR8y"
# Agora, com todas as imagens no computador remoto, vamos separá-las em pastas de modo que exista uma pasta para treino, uma para validação e uma para as imagens de teste.
#
# Nós pegaremos as 10 mil primeiras imagens de cachorros e de gatos e moveremos para a pasta de treino, e colocaremos mil imagens de cada tipo nas pastas de validação e teste. Em cada uma dessas pastas, cachorros e gatos serão separados em pastas diferentes.
#
#
# Podemos fazer isso com o código a seguir:
# + id="pkprCQO2fQPo"
import os, shutil
original_dataset_dir = '/datalab/train/'
dir_base = '/content/processed_datalab'
if not os.path.exists(dir_base):
os.mkdir(dir_base)
dir_treino = os.path.join(dir_base, 'train')
if not os.path.exists(dir_treino):
os.mkdir(dir_treino)
dir_valid = os.path.join(dir_base, 'validation')
if not os.path.exists(dir_valid):
os.mkdir(dir_valid)
dir_teste = os.path.join(dir_base, 'test')
if not os.path.exists(dir_teste):
os.mkdir(dir_teste)
train_cats_dir = os.path.join(dir_treino, 'cats')
if not os.path.exists(train_cats_dir):
os.mkdir(train_cats_dir)
train_dogs_dir = os.path.join(dir_treino, 'dogs')
if not os.path.exists(train_dogs_dir):
os.mkdir(train_dogs_dir)
validation_cats_dir = os.path.join(dir_valid, 'cats')
if not os.path.exists(validation_cats_dir):
os.mkdir(validation_cats_dir)
validation_dogs_dir = os.path.join(dir_valid, 'dogs')
if not os.path.exists(validation_dogs_dir):
os.mkdir(validation_dogs_dir)
test_cats_dir = os.path.join(dir_teste, 'cats')
if not os.path.exists(test_cats_dir):
os.mkdir(test_cats_dir)
test_dogs_dir = os.path.join(dir_teste, 'dogs')
if not os.path.exists(test_dogs_dir):
os.mkdir(test_dogs_dir)
fnames = ['cat.{}.jpg'.format(i) for i in range(10000)]
for fname in fnames:
src = os.path.join(original_dataset_dir, fname)
dst = os.path.join(train_cats_dir, fname)
shutil.copyfile(src, dst)
fnames = ['cat.{}.jpg'.format(i) for i in range(10000, 11000)]
for fname in fnames:
src = os.path.join(original_dataset_dir, fname)
dst = os.path.join(validation_cats_dir, fname)
shutil.copyfile(src, dst)
fnames = ['cat.{}.jpg'.format(i) for i in range(11000, 12000)]
for fname in fnames:
src = os.path.join(original_dataset_dir, fname)
dst = os.path.join(test_cats_dir, fname)
shutil.copyfile(src, dst)
fnames = ['dog.{}.jpg'.format(i) for i in range(10000)]
for fname in fnames:
src = os.path.join(original_dataset_dir, fname)
dst = os.path.join(train_dogs_dir, fname)
shutil.copyfile(src, dst)
fnames = ['dog.{}.jpg'.format(i) for i in range(10000, 11000)]
for fname in fnames:
src = os.path.join(original_dataset_dir, fname)
dst = os.path.join(validation_dogs_dir, fname)
shutil.copyfile(src, dst)
fnames = ['dog.{}.jpg'.format(i) for i in range(11000, 12000)]
for fname in fnames:
src = os.path.join(original_dataset_dir, fname)
dst = os.path.join(test_dogs_dir, fname)
shutil.copyfile(src, dst)
# + [markdown] id="SjZkggdqeZ4l"
# Mesmo com as imagens devidamente separadas, elas ainda não estão prontas para entrar em uma CNN. É necessário que se faça um pré-processamento das imagens para que todas fiquem na mesma escala e dimensão para que possam ser reconhecidas pela camada de entrada da nossa Rede Neural Convolucional. Mudaremos todas as imagens para escala de cinza de 8 bits, e deixaremos todas com medida 150x150.
#
# Utilizaremos o *ImageDataGenerator* do Keras.
# + id="MESbKg9ifUW_" colab={"base_uri": "https://localhost:8080/"} outputId="4779af1f-a183-4d62-cb6b-8dc0aae9c92b"
from keras.preprocessing.image import ImageDataGenerator
train_datagen = ImageDataGenerator(rescale=1./255)
test_datagen = ImageDataGenerator(rescale=1./255)
train_generator = train_datagen.flow_from_directory(
dir_treino,
target_size=(150, 150),
batch_size=20,
class_mode='binary')
validation_generator = test_datagen.flow_from_directory(
dir_valid,
target_size=(150, 150),
batch_size=20,
class_mode='binary')
# + [markdown] id="vN8kmwiag8Me"
# Com nossas imagens prontas, vamos desenvolver agora a nossa Rede Neural.
#
# Vamos adicionar 4 conjuntos Convolutional Layer e Max Pooling, com entrada 150x150, todas com Campos Receptivos Locais 5x5 e Região de Agrupamento 2x2. O número de feature maps irá aumentando ao longo das camadas, começando com 20, passando por 40 e tendo duas Camadas Convolucionais de 80 feature maps. A função de ativação utilizada será o Retificador Linear (ReLU).
#
# Podemos criar essa Rede Neural com o seguinte código:
# + id="xWSMYcpZgR2J"
from keras import layers
from keras import models
model = models.Sequential()
model.add(layers.Conv2D(16, (3, 3), activation='relu',
input_shape=(150, 150, 3)))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(32, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(32, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2)))
# + [markdown] id="O2hRKdmojoKT"
# Como as camadas até aqui tem duas dimensões, e as próximas camadas utilizadas tem apenas uma dimensão, precisaremos "achatar" os dados, de modo que eles fiquem com apenas uma dimensão, podemos fazer isso dessa maneira:
# + id="2-9lCju8j0gr"
model.add(layers.Flatten())
# + [markdown] id="-ef6zP9gkLlw"
# Agora adicionaremos uma camada com regularização do tipo dropout, para diminuir o efeito do overfitting, e adicionaremos uma camada densa de 100 neurônios, além da saída com a função de ativação sigmóide.
# + id="IDRrF-VFiITt"
model.add(layers.Dropout(0.5))
model.add(layers.Dense(512, activation='relu'))
model.add(layers.Dense(1, activation='sigmoid'))
# + [markdown] id="9vxKyIG3kdB3"
# Os hiper-parâmetros escolhidos neste notebook foram escolhidos baseados nos notebooks do Chollet<sup>[2]</sup>, no livro do Nielsen<sup>[1]</sup> e em testes empíricos.
#
# Com a função *summary* podemos ver como ficou nossa rede neural:
# + id="hD2EAAEbgUPS" colab={"base_uri": "https://localhost:8080/"} outputId="9055193e-de02-4099-b9bd-13ff2c345a9c"
model.summary()
# + [markdown] id="jAr5etXklGuC"
# Agora precisamos escolher nossos otimizadores:
# + id="UiBDuR9EVxbF" colab={"base_uri": "https://localhost:8080/"} outputId="689a2bf3-e8d2-4a74-b376-ed6c598f466a"
from keras import optimizers
model.compile(loss='binary_crossentropy',
optimizer=optimizers.RMSprop(lr=1e-4),
metrics=['acc'])
# + [markdown] id="cSbJN0JwlTH6"
# E enfim, podemos fazer o treino da nossa CNN, como os dados foram gerados pelo *DataGenerator*, nós teremos que utilizar a função fit_generator. Treinaremos para 50 épocas.
# + colab={"base_uri": "https://localhost:8080/"} id="9jSkqnfqXAYI" outputId="8c2d626e-8249-40a5-d4cb-9a66ab63e51a"
print(train_generator)
# + id="4xX-2OOsV2Xp" colab={"base_uri": "https://localhost:8080/"} outputId="6719532c-f9e7-4297-fa83-23bbe179cf27"
history = model.fit_generator(
train_generator,
steps_per_epoch=100,
epochs=100,
validation_data=validation_generator,
validation_steps=50)
# + id="LG1KivNlaGP2"
model.save('cats_and_dogs_test.h5')
# + [markdown] id="xzC2FqYMnNqr"
# Agora podemos avaliar os resultados:
# + id="mGB3Uid_lQ89"
model.save('cats_and_dogs_small_2.h5')
# + id="nl9KexkFV6XX"
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
# + id="21WdqZeXXRd0"
import matplotlib.pyplot as plt
epochs = range(len(acc))
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# + id="F_q6LwxmuUjq"
model.save('cats_and_dogs_small_2.h5')
# + id="vCDsXvMEfPVr" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="4a82d846-eac7-422f-b91c-cefd527d28db"
import numpy as np
np.shape(train_generator[0][0][0])
# + [markdown] id="PZZOz5eNnoHx"
# A instabilidade da função de custo mostra que ainda há espaço para melhoria, apesar disso, a CNN criada do zero conseguiu atingir ótimos resultados, uma acurácia de cerca de 87%.
# + [markdown] id="SCcN08zdpa50"
# ##5. Bibliografia
#
# [1] http://neuralnetworksanddeeplearning.com/index.html
#
# [2] https://github.com/fchollet/deep-learning-with-python-notebooks
#
# [3] https://colab.research.google.com/drive/19SVdlmnn6yRXCvNnE8PT1vbXrA8FrBo_#scrollTo=1HrGpk_b4dvL
#
# [4] https://www.kaggle.com/c/dogs-vs-cats
#
# [5] https://machinelearningmastery.com/how-to-develop-a-convolutional-neural-network-to-classify-photos-of-dogs-and-cats/
#
| Convolutional_Neural_Networks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Demo: VGG without Pre-trained Weights
# Below, you'll see how setting `weights=None` is equivalent to an un-trained network.
# ### Load example images and pre-process them
# +
# Load our images first, and we'll check what we have
from glob import glob
import matplotlib.image as mpimg
import matplotlib.pyplot as plt
from keras.preprocessing import image
from keras.applications.vgg16 import preprocess_input
import numpy as np
image_paths = glob('images/*.jpg')
i = 2 # Can change this to your desired image to test
img_path = image_paths[i]
img = image.load_img(img_path, target_size=(224, 224))
x = image.img_to_array(img)
x = np.expand_dims(x, axis=0)
x = preprocess_input(x)
# -
# ### Load VGG16 model, but without pre-trained weights
# This time, we won't use the pre-trained weights, so we'll likely get so wacky predictions.
# +
# Note - this will likely need to download a new version of VGG16
from keras.applications.vgg16 import VGG16, decode_predictions
# Load VGG16 without pre-trained weights
model = VGG16(weights=None)
# Perform inference on our pre-processed image
predictions = model.predict(x)
# Check the top 3 predictions of the model
print('Predicted:', decode_predictions(predictions, top=3)[0])
# -
# When we ran each image, we got a hand-blower, a guenon (a type of African monkey) and a mink. A little bit different than the elephant, labrador and zebra they are supposed to be!
| HelperProjects/Transfer-Learning/VGG/No_Pretraining.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# # Clustering menggunakan k-Means
#
# Notebook ini merupakan bagian dari buku **Machine Learning menggunakan Python** oleh **<NAME>**. Notebook ini berisi contoh kode untuk **BAB VIII - K-MEANS**
# +
# Mengimpor library
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.cm as cm
import matplotlib.pyplot as plt
from sklearn.cluster import KMeans
from sklearn.metrics import davies_bouldin_score, silhouette_samples, silhouette_score
# -
# ## Data Understanding
# Membaca data
df = pd.read_csv(r'../datasets/mall-customers.csv')
# + tags=[]
# Informasi dataset
df.info()
# -
# Statistik deskriptif
df.describe()
# Sampel data teratas
df.head()
# Agregasi data berdasarkan jenis kelamin
pd.pivot_table(df, index='sex')
# Scatter plot dan KDE berdasarkan jenis kelamin
sns.pairplot(data=df, hue="sex")
# ## Data Preparation
# Memisahkan features
X = df.iloc[:, 3:5].values
# +
# Elbow method
wcss = []
for i in range(1, 11):
kmeans = KMeans(n_clusters=i, random_state=42)
kmeans.fit(X)
wcss.append(kmeans.inertia_)
plt.plot(range(1, 11), wcss)
plt.xlabel('Number of clusters')
plt.ylabel('WCSS')
plt.show()
# -
# ## Modelling
# Membuat model k-means
kmeans = KMeans(n_clusters=5, random_state=42)
kmeans.fit(X)
# +
# Melakukan prediksi cluster
y_pred = kmeans.predict(X)
y_center = kmeans.cluster_centers_
df_cluster = df.copy()
df_cluster["cluster"] = y_pred
# -
# ## Evaluation
sns.scatterplot(data=df_cluster, x="annual-income", y="spending-score", hue="cluster", palette="viridis")
plt.scatter(y_center[:, 0], y_center[:, 1], s=300, c='yellow')
# Metrik
print("Davies-Bouldin Index:", davies_bouldin_score(X, y_pred))
print("Silhouette score:", silhouette_score(X, y_pred))
# + tags=[]
# Silhouette plot
# Sumber: https://scikit-learn.org/stable/auto_examples/cluster/plot_kmeans_silhouette_analysis.html
y_lower = 10
n_clusters = 5
fig, ax = plt.subplots()
ax.set_xlim([-0.1, 1])
ax.set_ylim([0, len(X) + (n_clusters + 1) * 10])
sample_silhouette_values = silhouette_samples(X, y_pred)
for i in range(n_clusters):
s_sample = sample_silhouette_values[y_pred == i]
s_sample.sort()
size_cluster_i = s_sample.shape[0]
y_upper = y_lower + size_cluster_i
color = cm.nipy_spectral(float(i) / n_clusters)
ax.fill_betweenx(np.arange(y_lower, y_upper), 0, s_sample, facecolor=color, edgecolor=color, alpha=0.7)
ax.text(-0.05, y_lower + 0.5 * size_cluster_i, str(i))
y_lower = y_upper + 10
silhouette_avg = silhouette_score(X, y_pred)
ax.axvline(x=silhouette_avg, color="red", linestyle="--")
ax.set_yticks([])
ax.set_xticks([-0.1, 0, 0.2, 0.4, 0.6, 0.8, 1])
ax.set_xlabel("Silhouette coefficient values")
ax.set_ylabel("Cluster label")
plt.show()
# -
# Box plot
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(10, 4))
sns.boxplot(data=df_cluster, x="cluster", y="spending-score", hue="sex", ax=ax1)
sns.boxplot(data=df_cluster, x="cluster", y="annual-income", hue="sex", ax=ax2)
# Pivot
pd.pivot_table(df_cluster, values=["annual-income", "spending-score"], columns="sex", index="cluster")
| jupyter/clustering_kmeans.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Dependencies and Setup
import pandas as pd
# File to Load (Remember to Change These)
school_data_to_load = "Resources/schools_complete.csv"
student_data_to_load = "Resources/students_complete.csv"
# Read School and Student Data File and store into Pandas DataFrames
school_data = pd.read_csv(school_data_to_load)
student_data = pd.read_csv(student_data_to_load)
# Combine the data into a single dataset.
school_data_df = pd.merge(student_data, school_data, how="left", on=["school_name", "school_name"])
school_data_df.head()
# -
school_data_df.count()
school_data_df.dtypes
#total schools
total_schools=len(school_data["school_name"].unique())
total_schools
#total students
total_students=student_data["student_name"].count()
total_students
#total budget
total_budget=school_data['budget'].sum()
total_budget
#avg math score
avg_m_score=student_data['math_score'].mean()
avg_m_score
#avg reading score
avg_r_score=student_data['reading_score'].mean()
avg_r_score
#70% math score
pass_math=student_data.loc[(student_data['math_score']>=70)]
count_math=pass_math['math_score'].count()
count_math
#70% reading score
pass_reading=student_data.loc[(student_data['reading_score']>=70)]
count_reading=pass_reading['reading_score'].count()
count_reading
#Passed Math %
pass_math_p=(count_math/total_students)*100
pass_math_p
#Passed Reading %
pass_reading_p=(count_reading/total_students)*100
pass_reading_p
# +
#Overall Passing % (passed math and reading)
overall_pass=(pass_math_p+pass_reading_p)/2
overall_pass
# +
#Summary Table
district_summary={"Total_Schools": total_schools,
"Total_Students": total_students,
"Total_Budget": total_budget,
"Average_Math_Score": avg_m_score,
"Average_Reading_Score": avg_r_score,
"%_Passing_Reading": pass_reading_p,
"%_Passing_Math": pass_math_p,
"%_Overall_Passing": overall_pass
}
district_summary_df=pd.DataFrame([district_summary])
district_summary_df=district_summary_df[["Total_Schools","Total_Students","Total_Budget","Average_Reading_Score","Average_Math_Score","%_Passing_Reading","%_Passing_Math","%_Overall_Passing"]]
district_summary_df
# -
# +
#SCHOOL SUMMARY PT 2
# -
school_copy=school_data
school_copy.columns
#per student budget
school_copy["per_student_budget"]=school_copy["budget"]/school_copy["size"]
school_copy.head()
average_ms_df=school_data_df.groupby(['school_name'])['math_score'].mean().reset_index()
average_ms_df.head()
average_rs_df=school_data_df.groupby(['school_name'])['reading_score'].mean().reset_index()
average_rs_df.head()
# +
averages_df=pd.merge(average_ms_df,average_rs_df,on="school_name",how="outer")
averages_df = averages_df.rename(columns={"math_score":"avg_math_score", "reading_score":"avg_reading_score"})
# -
school_copy=school_copy.merge(averages_df,on="school_name",how="outer")
school_copy.head()
#70% passing
passing_math_sum=school_data_df[school_data_df["math_score"]>=70]
passing_reading_sum=school_data_df[school_data_df["reading_score"]>=70]
#70% passing reading by school
passing_reading_count_sum=passing_reading_sum.groupby(["school_name"])["reading_score"].count().reset_index()
passing_reading_count_sum =passing_reading_count_sum.rename(columns={"reading_score":"reading_count"})
passing_reading_count_sum.head()
#70% passing math by school
passing_math_count_sum=passing_math_sum.groupby(["school_name"])["math_score"].count().reset_index()
passing_math_count_sum =passing_math_count_sum.rename(columns={"math_score":"math_count"})
passing_math_count_sum.head()
passing_count_totals=passing_math_count_sum.merge(passing_reading_count_sum, on="school_name", how="inner")
passing_count_totals.head()
school_copy=school_copy.merge(passing_count_totals,on="school_name",how="outer")
school_copy.head()
school_copy["%_passing_math"]=(school_copy["math_count"]/school_copy["size"])*100
school_copy["%_passing_reading"]=(school_copy["reading_count"]/school_copy["size"])*100
school_copy.head()
del school_copy["math_count"]
del school_copy["reading_count"]
school_copy.head()
school_copy["overall_passing_%"]=(school_copy['%_passing_math']+school_copy['%_passing_reading'])/2
school_copy.head()
# +
#TOP PERFORMING SCHOOLS PT 3
# -
top_performing_df=school_copy
top_performing_df=top_performing_df.sort_values(by=["overall_passing_%"],ascending=False)
top_performing_df.head()
# +
#BOTTOM PERFORMING SCHOOLS PT 4
# -
bottom_performing_df=school_copy
bottom_performing_df=bottom_performing_df.sort_values(by=["overall_passing_%"])
bottom_performing_df.head()
#MATH SCORES BY GRADE PT 5
student_data.head()
# +
math_score_9 = student_data.loc[student_data["grade"]=="9th"].groupby(["school_name"])["math_score"].mean()
math_score_10 = student_data.loc[student_data["grade"]=="10th"].groupby(["school_name"])["math_score"].mean()
math_score_11 = student_data.loc[student_data["grade"]=="11th"].groupby(["school_name"])["math_score"].mean()
math_score_12 = student_data.loc[student_data["grade"]=="12th"].groupby(["school_name"])["math_score"].mean()
math_scores_by_grade=pd.DataFrame({"9th_grade":math_score_9,"10th_grade":math_score_10,"11th_grade":math_score_11,"12th_grade":math_score_12})
math_scores_by_grade.head()
# -
# +
#READING SCORES BY GRADE 6
# +
reading_score_9 = student_data.loc[student_data["grade"]=="9th"].groupby(["school_name"])["reading_score"].mean()
reading_score_10 = student_data.loc[student_data["grade"]=="10th"].groupby(["school_name"])["reading_score"].mean()
reading_score_11 = student_data.loc[student_data["grade"]=="11th"].groupby(["school_name"])["reading_score"].mean()
reading_score_12 = student_data.loc[student_data["grade"]=="12th"].groupby(["school_name"])["reading_score"].mean()
reading_scores_by_grade=pd.DataFrame({"9th_grade":reading_score_9,"10th_grade":reading_score_10,"11th_grade":reading_score_11,"12th_grade":reading_score_12})
reading_scores_by_grade.head()
# -
# +
#SCORES BY SCHOOL SPENDING PT 7
# -
school_copy.head()
bins=[0,585,630,645,680]
groups=["<585","585-630","630-645","645-680"]
school_copy["spending_ranges"]=pd.cut(school_copy["budget"]/school_copy["size"],bins,labels=groups)
school_copy.head()
scores_by_school=school_copy.drop(columns=["School ID","type","size","budget","per_student_budget",])
scores_by_school
scores_by_school_df=scores_by_school[["spending_ranges","school_name","avg_math_score","avg_reading_score","%_passing_math","%_passing_reading","overall_passing_%"]]
scores_by_school_df
# +
#Scores by School Spending pt 2 Groupby
# -
scores_by_spending=scores_by_school_df.groupby(["spending_ranges"])
scores_by_spending.mean()
# +
#SCORES BY SCHOOL SIZE 8
# -
scores_by_size=school_copy
scores_by_size
size_bins=[0,1000,2000,5000]
size_labels=["small","medium","large"]
size_bins_df=pd.cut(scores_by_size["size"],size_bins,labels=size_labels)
size_bins_df=pd.DataFrame(size_bins_df)
school_copy["school_size"]=size_bins_df
scores_by_size=school_copy.groupby(["school_size","school_name"])["avg_math_score","avg_reading_score","%_passing_math","%_passing_reading","overall_passing_%"].mean()
scores_by_size
# +
#SCORES BY SCHOOL TYPE PT 9
# -
scores_by_type=school_copy
scores_by_type=pd.DataFrame(scores_by_type)
scores_by_type=school_copy.groupby(['type'])["avg_math_score","avg_reading_score","%_passing_math","%_passing_reading","overall_passing_%"].mean()
scores_by_type
# +
#Observataions
#There is no significant correlation between school spending and average math/reading scores
#Schools with a smaller population performed better than larger schools overall
#Charter schools outperformed district schools across th board
# -
| Pandas (School).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: IPython (Python 3)
# language: python
# name: python3
# ---
import glob
import os
import pandas as pd
import seaborn as sns
# +
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.use('Agg')
# %matplotlib inline
# -
# ! ls ../unused_reads/unmapped-final/results_summary/downsample_1000/
results = glob.glob('../unused_reads/unmapped-final/results_summary/downsample_1000/*.tsv')
results
results_dict = {}
for unmapped_result in results:
print(os.path.basename(unmapped_result))
sample_name = os.path.basename(unmapped_result).rstrip('.tsv')
results_dict[sample_name] = pd.read_csv(unmapped_result, sep = '\t', names=['reference', 'BLAST hits'])
forbidden_terms = ['ribosom', 'RNA', 'ynthetic']
# +
results_without_strings = {}
for result_name, result_df in results_dict.items():
print(result_name)
print('num rows before removal: {}'.format(result_df.shape))
# remove rows that have "ribosom", and "RNA"
for f_term in forbidden_terms:
# df.C.str.contains("XYZ") == False]
result_df = result_df[result_df['reference'].str.contains(f_term) == False]
print('num rows after removal: {}'.format(result_df.shape))
# append a column on that is the fraction of all BLAST hits.
hit_total = result_df['BLAST hits'].sum()
print('{} BLAST hits for {}'.format(hit_total, result_name))
result_df['BLAST hit fraction'] = result_df['BLAST hits']*1.0/hit_total
print(result_df.head(2))
results_dict[result_name] = result_df
print('==================')
# -
for df in results_dict.values():
print(df.shape)
plot_dir = './plots/160601_unused_read_histograms/'
if not os.path.exists(plot_dir):
os.makedirs(plot_dir)
for sample_name, result_df in results_dict.items():
fig, ax = plt.subplots()
result_df.head(10).plot.barh(x='reference', y='BLAST hit fraction',
stacked=True, ax=ax, legend=False,
title = sample_name,
figsize=(9,4))
#ax.set_yticklabels(['A', 'B', 'C', 'D', 'E', 'F'])
ax.invert_yaxis() # put big bars on top
ax.set_xlabel("fraction of BLAST hits \n(excluding RNA & synthetic constructs)")
plt.gcf().subplots_adjust(left=0.7)
plt.gcf().subplots_adjust(bottom=0.2)
plt.xticks(rotation=90)
#fig.tight_layout()
fig.savefig(plot_dir + '160601_{}_unmapped_reads.pdf'.format(sample_name))
fig.savefig(plot_dir + '160601_{}_unmapped_reads.svg'.format(sample_name))
| ipython_notebooks/160601_top_hits_for_unmapped_reads_excluding_RNA_and_synthetic_constructs.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# formats: ipynb,py:light
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
df = pd.read_csv('../../data/processed/LOLOracleDataWr.csv')
df.head()
df.columns
Red = ['RTop', 'RJng', 'RMid', 'RAdc','RSup']
Blue = ['BTop', 'BJng', 'BMid', 'BAdc', 'BSup']
for i in Red:
df[i]=-df[i]
df.head()
# +
#For all entries with Blue = 1 and Red = -1
for i in range(len(df)):
if df.loc[i,'Winner']==0:
df.loc[i,'Winner']=1
else:
df.loc[i,'Winner']=-1
df['Winner'] = df['Winner'].astype(int)
# -
df.head()
# ### Lets implement KNN model now
from sklearn.model_selection import train_test_split
X = df.drop('Winner',axis=1)
y = df['Winner']
X_train,X_test,y_train,y_test = train_test_split(X,y,test_size=0.3,random_state=101)
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier(n_neighbors=1)
knn.fit(X_train,y_train)
predictions = knn.predict(X_test)
from sklearn.metrics import confusion_matrix,classification_report
print(confusion_matrix(y_test,predictions))
print('\n')
print(classification_report(y_test,predictions))
# +
#Lets choose a K value now
# +
error_rate = []
for i in range(1,100):
knn = KNeighborsClassifier(n_neighbors=i)
knn.fit(X_train,y_train)
pred_i = knn.predict(X_test)
error_rate.append(np.mean(pred_i != y_test))
# -
plt.figure(figsize=(10,6))
plt.plot(range(1,100),error_rate,color='blue', linestyle='dashed', marker='o',
markerfacecolor='red', markersize=10)
plt.title('Error Rate vs. K Value')
plt.xlabel('K')
plt.ylabel('Error Rate')
knn = KNeighborsClassifier(n_neighbors=64)
knn.fit(X_train,y_train)
predictions = knn.predict(X_test)
from sklearn.metrics import confusion_matrix,classification_report
print(confusion_matrix(y_test,predictions))
print('\n')
print(classification_report(y_test,predictions))
| models/level2/KNN_ChampWr-Diffparam.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## [Experiments] Implementation of GP-Select
#
# Implement the optimisation criterion from the Paper "Selecting valuable items from massive data" with a custom kernel function.
# %matplotlib inline
from sklearn.gaussian_process import GaussianProcessRegressor
from sklearn.gaussian_process.kernels import (RBF, Matern, RationalQuadratic,
ExpSineSquared, DotProduct,
ConstantKernel)
import math
import numpy as np
from matplotlib import pyplot as plt
import sklearn.datasets as data
houses = data.load_boston()
X.shape
y = houses['target']
X = houses['data']
kernel = 1.0 * RBF(length_scale=1.0,length_scale_bounds=(1e-1,10.0))
gp = GaussianProcessRegressor(kernel=kernel)
# How does the std change when we use more data for prediction?
def fit_and_predict(using_examples):
gp.fit(X[:using_examples], y[:using_examples])
y_mean, y_std = gp.predict(X,return_std=True)
return y_std
# ## Random ordering
test_samples = range(1,506)
std = [fit_and_predict(i) for i in test_samples]
max_std = [item.max() for item in std]
mean_std = [item.mean() for item in std]
var_std = [item.var() for item in std]
std_y = range(len(test_samples))
plt.plot(std_y,max_std,color='b',)
plt.plot(std_y,mean_std,color='g')
plt.plot(std_y,var_std,color='y')
# ## Uncertainty Sampling, Greedy
def fit_and_predict_indexed(idx):
gp.fit(X[idx], y[idx])
y_mean, y_std = gp.predict(X,return_std=True)
return y_std
# +
current_idx = []
std_unSamp = []
# init model with random data point
start = np.random.choice(np.arange(506))
current_idx.append(start)
current_y_std = fit_and_predict_indexed(idx)
std_unSamp.append(current_y_std)
for index in range(2,100):
max_std_idx = np.unravel_index(np.argmax(current_y_std, axis=None), y_std.shape)
current_idx.append(max_std_idx)
current_y_std = fit_and_predict_indexed(idx)
std_unSamp.append(current_y_std)
# -
max_std_unSamp = [item.max() for item in std_unSamp]
mean_std_unSamp = [item.mean() for item in std_unSamp]
var_std_unSamp = [item.var() for item in std_unSamp]
std_y = range(len(std_unSamp))
plt.plot(std_y,max_std_unSamp,color='b')
plt.plot(std_y,mean_std_unSamp,color='g')
plt.plot(std_y,var_std_unSamp,color='y')
#=> Why do all the std's aquire the same value?
#=> If we compare this to the lower dimensional case we might be having the case of the curse of dim
#=> We have 15 dimensions and fit at most ~100 items
#=> How would this change if we changed the number of features?
def fit_and_predict_indexed_3features(idx):
gp.fit(X[idx,:3], y[idx])
y_mean, y_std = gp.predict(X[:,:3],return_std=True)
return y_std
# +
current_idx = []
std_unSampSmall = []
# init model with random data point
start = np.random.choice(np.arange(506))
current_idx.append(start)
current_y_std = fit_and_predict_indexed(idx)
std_unSampSmall.append(current_y_std)
for index in range(2,100):
max_std_idx = np.unravel_index(np.argmax(current_y_std, axis=None), y_std.shape)
current_idx.append(max_std_idx)
current_y_std = fit_and_predict_indexed(idx)
std_unSampSmall.append(current_y_std)
# -
max_std_unSampSmall = [item.max() for item in std_unSampSmall]
mean_std_unSampSmall = [item.mean() for item in std_unSampSmall]
var_std_unSampSmall = [item.var() for item in std_unSampSmall]
std_y = range(len(std_unSampSmall))
plt.plot(std_y,max_std_unSampSmall,color='b')
plt.plot(std_y,mean_std_unSampSmall,color='g')
plt.plot(std_y,var_std_unSampSmall,color='y')
#=> fewer features do not help!
#=> The argmax funtion takes always the first highest element
#=> so we are deterministically approximating our function space from the right side
#=> This is not a good idea
# When does a point in a function contain more information than its neighbour?
# maybe, if it was against several of our last hypothesises? Average the model over time
# Use the variational prediction at a point
# The question should specify information about **what**
# Maybe this is something about how muh local structure corresponds to global structure
# In active learning we would like to label the points that correspond best to the global structure, because this structure would give us the best *global* estimate
# Is this just a reformulation of the diversity term?
No
# The diversity term says we want to query terms that are very different from each other, but finding global structure implies that we do not
| notebooks/active_learning/fb-01-probabilistic approaches/03-GP_Select.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# **Url**
#
# https://boeing.mediaroom.com/news-releases-statements?l=100&o=200
# +
import os
os.environ['KMP_DUPLICATE_LIB_OK']='True'
import pandas as pd,requests,bs4,re,time,io,pytesseract,easyocr,random
from pdfminer.high_level import extract_text
from pathlib import Path
from pdf2image import convert_from_path
from selenium.webdriver.common.by import By
from goose3 import Goose
from datetime import datetime
from bs4 import BeautifulSoup
from selenium import webdriver
reader = easyocr.Reader(['en'])
import warnings
warnings.filterwarnings("ignore")
# %autosave 1
# +
SITE_NAME='Boeing'
DOMAIN = "https://boeing.mediaroom.com"
SITE_LINK="https://boeing.mediaroom.com/news-releases-statements?l=100&o=0"
# +
def parse_webpage_bs(search_url):
headers = {
"User-Agent": "Mozilla/5.0 (Windows NT 10.0; Win64; x64; rv:90.0) Gecko/20100101 Firefox/90.0"}
try:
site_request = requests.get(search_url, headers=headers, timeout=10)
except requests.exceptions.RequestException as e:
print(e)
site_request = None
if site_request != None and site_request.status_code==200:
site_soup = bs4.BeautifulSoup(site_request.content, "lxml")
else:
site_soup = None
return site_soup
def remove_esc_chars(text):
return text.replace("\n", " ").replace("\t", " ").replace("\r", " ")
# -
def get_text(link):
g = Goose()
article_extract = g.extract(url=link)
article = remove_esc_chars(article_extract.cleaned_text)
meta_data = remove_esc_chars(article_extract.meta_description)
whole_data = meta_data+article
text = whole_data.strip()
if len(text) < 10:
try:
response = requests.get(link)
text = remove_esc_chars(extract_text(io.BytesIO(response.content)))
if len(text) < 10:
texts = ""
r = requests.get(link)
filename = Path('temp.pdf')
filename.write_bytes(r.content)
pages = convert_from_path('temp.pdf', 500)
for x in pages:
x.save("temp.jpg")
output = reader.readtext("temp.jpg")
for o in output:
texts += o[1]
text = remove_esc_chars(texts)
except:
text = ""
return text
# +
article_list = []
pagination = 0
last_page = 11000
last_page = 200
while pagination < last_page:
url = f"https://boeing.mediaroom.com/news-releases-statements?l=100&o={pagination}"
soup = parse_webpage_bs(url)
if soup != None:
elements = soup.findAll('li',{'class' :'wd_item'})
for element in elements[:10]:
#title,published_date,link,thumbnail,author
title = element.find('div',{'class' :'wd_title'}).text.strip()
published_date = element.find('div',{'class' :'wd_date'}).text.strip()
link = element.find('div',{'class' :'wd_tooltip-trigger'}).find('a')['href']
author = SITE_NAME
try:
thumbnail = DOMAIN + element.find('div',{'class' :'wd_thumbnail'}).find('img')['src']
except:
thumbnail = "https://upload.wikimedia.org/wikipedia/commons/thumb/4/4f/Boeing_full_logo.svg/1920px-Boeing_full_logo.svg.png"
#text
text = get_text(link)
article = (published_date.strip(),title.strip(),text.strip(),link.strip(),thumbnail.strip(),author.strip())
article_list.append(article)
print(published_date,title)
pagination +=100
# -
temp_df = pd.DataFrame(article_list,columns=['date','title','article','url','thumbnail','author'])
temp_df.head()
# +
def see_data(iloc_no=random.randint(0,len(temp_df))-1):
print(temp_df.iloc[iloc_no]['date'],temp_df.iloc[iloc_no]['title'])
print(f"\n{temp_df.iloc[iloc_no]['author']} {temp_df.iloc[iloc_no]['url']}")
print(f"\n{temp_df.iloc[iloc_no]['article']}")
print(f"\n{temp_df.iloc[iloc_no]['thumbnail']}")
see_data()
# +
#to csv
temp_df.to_csv(f'{SITE_NAME} news.csv',index = False)
#to json
temp_df.to_json(f'{SITE_NAME} news.json')
# +
#to get rid of unwanteed trash created by the model use
def remove_trash():
try:
os.remove("temp.jpg")
os.remove("temp.pdf")
print("Trash removed successfully")
except:
print("No trash found")
remove_trash()
| boeing/boeing_bs4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import math
def factorial(n):
return(math.factorial(n))
num = int(input("Enter Your Number to find out The factorial: "))
print("Factorial of", num, "is",factorial(num))
# -
| Factorial-of-a-num.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import logging
import importlib
importlib.reload(logging) # see https://stackoverflow.com/a/21475297/1469195
log = logging.getLogger()
log.setLevel('INFO')
import sys
logging.basicConfig(format='%(asctime)s %(levelname)s : %(message)s',
level=logging.INFO, stream=sys.stdout)
# +
# %%capture
import os
import site
os.sys.path.insert(0, '/home/schirrmr/code/hyperoptim//')
os.sys.path.insert(0, '/home/schirrmr/code/reversible/')
os.sys.path.insert(0, '/home/schirrmr/braindecode/code/braindecode/')
# %load_ext autoreload
# %autoreload 2
import numpy as np
import logging
log = logging.getLogger()
log.setLevel('INFO')
import sys
logging.basicConfig(format='%(asctime)s %(levelname)s : %(message)s',
level=logging.INFO, stream=sys.stdout)
import matplotlib
from matplotlib import pyplot as plt
from matplotlib import cm
# %matplotlib inline
# %config InlineBackend.figure_format = 'png'
matplotlib.rcParams['figure.figsize'] = (12.0, 1.0)
matplotlib.rcParams['font.size'] = 14
import seaborn
seaborn.set_style('darkgrid')
# +
from hyperoptim.results import (load_data_frame,
remove_columns_with_same_value, mean_identical_exps, pairwise_compare_frame,
round_numeric_columns)
import pandas as pd
def load_df(folder):
df = load_data_frame(folder)
df = df.drop(['save_folder', 'seed', 'only_return_exp'], axis=1)
df = df[df.finished == 1]
df = df[df.debug == 0]
if len(df) > 0:
for key in [
'model_train_acc', 'model_test_acc', 'train_train_acc', 'train_test_acc',
'combined_train_acc', 'combined_test_acc', 'test_train_acc', 'test_test_acc',]:
if key in df.columns:
df.loc[:,key] = np.round(df.loc[:,key] * 100,1)
df = remove_columns_with_same_value(df)
df.runtime = pd.to_timedelta(np.round(df.runtime), unit='s')
if 'clf_loss' in df.columns:
df.clf_loss = df.clf_loss.replace([None], 'none')
df = df.fillna('-')
print(len(df))
return df
# +
result_cols = [
'model_train_acc', 'model_test_acc', 'train_train_acc', 'train_test_acc',
'combined_train_acc', 'combined_test_acc', 'test_train_acc', 'test_test_acc',
'traintime','train_NLL', 'test_NLL', 'train_OT', 'test_OT',
'lip_loss', 'runtime',]
interesting_result_cols = [
'model_train_acc', 'model_test_acc', 'train_train_acc', 'train_test_acc',
'train_NLL', 'test_NLL', 'train_OT', 'test_OT',]
# +
df = load_df('/data/schirrmr/schirrmr/reversible/experiments/dropout-weight-decay/')
df = mean_identical_exps(df, result_cols=result_cols).drop('n_exp', axis=1)
df.sort_values(by='train_test_acc', ascending=False)
# -
df[df.gauss_noise_factor!='-'].drop(np.setdiff1d(result_cols, interesting_result_cols), axis=1)
# +
df = load_df('/data/schirrmr/schirrmr/reversible/experiments/dropout-weight-decay-more-data//')
df = mean_identical_exps(df, result_cols=result_cols).drop('n_exp', axis=1)
df.sort_values(by='train_test_acc', ascending=False)
| notebooks/simpler-invnet-20-june-2019/.ipynb_checkpoints/DropoutWeightDecay-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="eTdCMVl9YAXw" colab_type="text"
# <a href="https://practicalai.me"><img src="https://raw.githubusercontent.com/practicalAI/images/master/images/rounded_logo.png" width="100" align="left" hspace="20px" vspace="20px"></a>
#
# <img src="https://raw.githubusercontent.com/practicalAI/images/master/basic_ml/12_Recurrent_Neural_Networks/simple_rnn.png" width="250" align="right">
#
# <div align="left">
# <h1>Recurrent Neural Networks (RNN) </h1>
#
# In this lesson we will learn how to process sequential data (sentences, time-series, etc.) with recurrent neural networks (RNNs). </div>
# + [markdown] id="xuabAj4PYj57" colab_type="text"
# <table align="center">
# <td>
# <img src="https://raw.githubusercontent.com/practicalAI/images/master/images/rounded_logo.png" width="25"><a target="_blank" href="https://practicalai.me"> View on practicalAI</a>
# </td>
# <td>
# <img src="https://raw.githubusercontent.com/practicalAI/images/master/images/colab_logo.png" width="25"><a target="_blank" href="https://colab.research.google.com/github/practicalAI/practicalAI/blob/master/notebooks/12_Recurrent_Neural_Networks.ipynb"> Run in Google Colab</a>
# </td>
# <td>
# <img src="https://raw.githubusercontent.com/practicalAI/images/master/images/github_logo.png" width="22"><a target="_blank" href="https://github.com/practicalAI/practicalAI/blob/master/notebooks/basic_ml/12_Recurrent_Neural_Networks.ipynb"> View code on GitHub</a>
# </td>
# </table>
# + [markdown] id="uXuhm14WaCxy" colab_type="text"
# So far we've processed inputs as whole (ex. applying filters across the entire input to extract features) but we can also processed our inputs sequentially. To illustrate what this looks like, suppose our inputs are sentences (like the news dataset inputs we've seen in previous lessons). Think of each word/puncutation (or token) in the sentence as a timestep. So a sentence with 8 tokens has 8 timesteps. We can process each timestep, one at a time, and predict the class after the last timestep (token) has been processed. This is very powerful because the model now has a meaningful way to account for the order in our sequence and predict accordingly.
# + [markdown] id="m5JJY0tl8StF" colab_type="text"
# # Overview
# + [markdown] id="JqxyljU18hvt" colab_type="text"
# * **Objective:** Process sequential data by accounting for the currend input and also what has been learned from previous inputs.
# * **Advantages:**
# * Account for order and previous inputs in a meaningful way.
# * Conditioned generation for generating sequences.
# * **Disadvantages:**
# * Each time step's prediction depends on the previous prediction so it's difficult to parallelize RNN operations.
# * Processing long sequences can yield memory and computation issues.
# * Interpretability is difficult but there are few [techniques](https://arxiv.org/abs/1506.02078) that use the activations from RNNs to see what parts of the inputs are processed.
# * **Miscellaneous:**
# * Architectural tweaks to make RNNs faster and interpretable is an ongoing area of research.
# + [markdown] id="oSOqIKzckhfc" colab_type="text"
# # Set up
# + id="vSPqM7rrkhE8" colab_type="code" outputId="38ef452c-e050-4d8a-d31b-e14b43d292bd" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Use TensorFlow 2.x
# %tensorflow_version 2.x
# + id="GuK4QHvikkpR" colab_type="code" outputId="128c7a2a-daa7-487e-ba04-2b7f957a0081" colab={"base_uri": "https://localhost:8080/", "height": 34}
import os
import numpy as np
import tensorflow as tf
print("GPU Available: ", tf.test.is_gpu_available())
# + id="0Zwz1m7mknyd" colab_type="code" colab={}
# Arguments
SEED = 1234
SHUFFLE = True
DATA_FILE = 'news.csv'
INPUT_FEATURE = 'title'
OUTPUT_FEATURE = 'category'
FILTERS = "!\"'#$%&()*+,-./:;<=>?@[\\]^_`{|}~"
LOWER = True
CHAR_LEVEL = False
TRAIN_SIZE = 0.7
VAL_SIZE = 0.15
TEST_SIZE = 0.15
NUM_EPOCHS = 10
BATCH_SIZE = 256
EMBEDDING_DIM = 100
RNN_HIDDEN_DIM = 128
RNN_DROPOUT_P = 0.1
NUM_LAYERS = 1
HIDDEN_DIM = 100
DROPOUT_P = 0.1
LEARNING_RATE = 1e-3
EARLY_STOPPING_CRITERIA = 3
# + id="HgEiBe6DkpXV" colab_type="code" colab={}
# Set seed for reproducibility
np.random.seed(SEED)
tf.random.set_seed(SEED)
# + [markdown] id="c69z9wpJ56nE" colab_type="text"
# # Data
# + [markdown] id="2V_nEp5G58M0" colab_type="text"
# We will download the [AG News dataset](http://www.di.unipi.it/~gulli/AG_corpus_of_news_articles.html), which consists of 120000 text samples from 4 unique classes ('Business', 'Sci/Tech', 'Sports', 'World')
# + id="y3qKSoEe57na" colab_type="code" colab={}
import pandas as pd
import re
import urllib
# + id="cGQo98566GIV" colab_type="code" colab={}
# Upload data from GitHub to notebook's local drive
url = "https://raw.githubusercontent.com/practicalAI/practicalAI/master/data/news.csv"
response = urllib.request.urlopen(url)
html = response.read()
with open(DATA_FILE, 'wb') as fp:
fp.write(html)
# + id="dG_Oltib6G-9" colab_type="code" outputId="eebcc768-99c4-4932-b3a2-bab6ea733920" colab={"base_uri": "https://localhost:8080/", "height": 204}
# Load data
df = pd.read_csv(DATA_FILE, header=0)
X = df[INPUT_FEATURE].values
y = df[OUTPUT_FEATURE].values
df.head(5)
# + [markdown] id="hxo6RKCQ71dl" colab_type="text"
# # Split data
# + id="eS6kCcfY6IHE" colab_type="code" colab={}
import collections
from sklearn.model_selection import train_test_split
# + [markdown] id="Mt3yeNsYXU_F" colab_type="text"
# ### Components
# + id="0gbh2TLBXVDu" colab_type="code" colab={}
def train_val_test_split(X, y, val_size, test_size, shuffle):
"""Split data into train/val/test datasets.
"""
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=test_size, stratify=y, shuffle=shuffle)
X_train, X_val, y_train, y_val = train_test_split(
X_train, y_train, test_size=val_size, stratify=y_train, shuffle=shuffle)
return X_train, X_val, X_test, y_train, y_val, y_test
# + [markdown] id="8XIdYU_n7536" colab_type="text"
# ### Operations
# + id="kqiQd2j_76gP" colab_type="code" outputId="7f1d72dd-11f1-4d31-cebd-9d00caf91524" colab={"base_uri": "https://localhost:8080/", "height": 119}
# Create data splits
X_train, X_val, X_test, y_train, y_val, y_test = train_val_test_split(
X=X, y=y, val_size=VAL_SIZE, test_size=TEST_SIZE, shuffle=SHUFFLE)
class_counts = dict(collections.Counter(y))
print (f"X_train: {X_train.shape}, y_train: {y_train.shape}")
print (f"X_val: {X_val.shape}, y_val: {y_val.shape}")
print (f"X_test: {X_test.shape}, y_test: {y_test.shape}")
print (f"X_train[0]: {X_train[0]}")
print (f"y_train[0]: {y_train[0]}")
print (f"Classes: {class_counts}")
# + [markdown] id="dIfmW7vJ8Jx1" colab_type="text"
# # Tokenizer
# + [markdown] id="JP4VCO0LAJUt" colab_type="text"
# Unlike the previous notebook, we will be processing our text at a word-level (as opposed to character-level).
# + id="DHPAxkKR7736" colab_type="code" colab={}
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.utils import to_categorical
# + [markdown] id="EKAXyjYHfEzU" colab_type="text"
# ### Components
# + id="bVgBSnFTfE5D" colab_type="code" colab={}
def untokenize(indices, tokenizer):
"""Untokenize a list of indices into string."""
return " ".join([tokenizer.index_word[index] for index in indices])
# + [markdown] id="_BD3XPKF8L84" colab_type="text"
# ### Operations
# + id="WcscM_vL8KvP" colab_type="code" colab={}
# Input vectorizer
X_tokenizer = Tokenizer(filters=FILTERS,
lower=LOWER,
char_level=CHAR_LEVEL,
oov_token='<UNK>')
# + id="xV2JgpOA8PwO" colab_type="code" outputId="b116f5dc-692f-4dfb-8d97-8dde0ceeb69d" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Fit only on train data
X_tokenizer.fit_on_texts(X_train)
vocab_size = len(X_tokenizer.word_index) + 1
print (f"# tokens: {vocab_size}")
# + id="ybb-YZSz8Qno" colab_type="code" outputId="742e501f-043e-4cb6-fac0-a8a8ea59cf2b" colab={"base_uri": "https://localhost:8080/", "height": 68}
# Convert text to sequence of tokens
original_text = X_train[0]
X_train = np.array(X_tokenizer.texts_to_sequences(X_train))
X_val = np.array(X_tokenizer.texts_to_sequences(X_val))
X_test = np.array(X_tokenizer.texts_to_sequences(X_test))
preprocessed_text = untokenize(X_train[0], X_tokenizer)
print (f"{original_text} \n\t→ {preprocessed_text} \n\t→ {X_train[0]}")
# + [markdown] id="ORGuhjCf8TKh" colab_type="text"
# # LabelEncoder
# + id="7aBBgzkW8Rxv" colab_type="code" colab={}
from sklearn.preprocessing import LabelEncoder
# + [markdown] id="z_jVCsl98U09" colab_type="text"
# ### Operations
# + id="ckM_MnQi8UTH" colab_type="code" colab={}
# Output vectorizer
y_tokenizer = LabelEncoder()
# + id="0-FkxqCT8WUk" colab_type="code" outputId="5d5dc244-95df-4bc6-b93f-5bbe3cf0be99" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Fit on train data
y_tokenizer = y_tokenizer.fit(y_train)
classes = list(y_tokenizer.classes_)
print (f"classes: {classes}")
# + id="yrLHd1i_8XAJ" colab_type="code" outputId="913fa40c-307e-4e98-d065-226a65311914" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Convert labels to tokens
y_train = y_tokenizer.transform(y_train)
y_val = y_tokenizer.transform(y_val)
y_test = y_tokenizer.transform(y_test)
print (f"y_train[0]: {y_train[0]}")
# + id="bJAR-FDUSP4q" colab_type="code" outputId="d8a30a83-5474-4e03-dc8b-cab3eb706dd3" colab={"base_uri": "https://localhost:8080/", "height": 51}
# Class weights
counts = np.bincount(y_train)
class_weights = {i: 1.0/count for i, count in enumerate(counts)}
print (f"class counts: {counts},\nclass weights: {class_weights}")
# + [markdown] id="eoWQk0hO9bK2" colab_type="text"
# # Generators
# + id="GVxnbzgW8X1V" colab_type="code" colab={}
import math
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tensorflow.keras.utils import Sequence
# + [markdown] id="IMtHyqex9gVI" colab_type="text"
# ### Components
# + id="1w6wVKJe9fxk" colab_type="code" colab={}
class DataGenerator(Sequence):
"""Custom data loader."""
def __init__(self, X, y, batch_size, shuffle=True):
self.X = X
self.y = y
self.batch_size = batch_size
self.shuffle = shuffle
self.on_epoch_end()
def __len__(self):
"""# of batches."""
return math.ceil(len(self.X) / self.batch_size)
def __str__(self):
return (f"<DataGenerator(" \
f"batch_size={self.batch_size}, " \
f"batches={len(self)}, " \
f"shuffle={self.shuffle})>")
def __getitem__(self, index):
"""Generate a batch."""
# Gather indices for this batch
batch_indices = self.epoch_indices[index*self.batch_size:(index+1)*self.batch_size]
# Generate batch data
inputs, outputs = self.create_batch(batch_indices=batch_indices)
return inputs, outputs
def on_epoch_end(self):
"""Create indices after each epoch."""
self.epoch_indices = np.arange(len(self.X))
if self.shuffle == True:
np.random.shuffle(self.epoch_indices)
def create_batch(self, batch_indices):
"""Generate data from indices."""
X = self.X[batch_indices]
y = self.y[batch_indices]
# Sequence lengths
seq_lengths = np.array([[i, len(x)-1] for i, x in enumerate(X)])
# Pad batch
max_seq_len = max([len(x) for x in X])
X = pad_sequences(X, padding="post", maxlen=max_seq_len)
return [X, seq_lengths], y
# + [markdown] id="u37JyFYV9ilS" colab_type="text"
# ### Operations
# + id="5T8mVj9d9hNI" colab_type="code" colab={}
# Dataset generator
training_generator = DataGenerator(X=X_train,
y=y_train,
batch_size=BATCH_SIZE,
shuffle=SHUFFLE)
validation_generator = DataGenerator(X=X_val,
y=y_val,
batch_size=BATCH_SIZE,
shuffle=False)
testing_generator = DataGenerator(X=X_test,
y=y_test,
batch_size=BATCH_SIZE,
shuffle=False)
# + id="drbY5WDX9kcL" colab_type="code" outputId="79462796-a13d-447f-9435-f67b6b83716d" colab={"base_uri": "https://localhost:8080/", "height": 68}
print (f"training_generator: {training_generator}")
print (f"validation_generator: {validation_generator}")
print (f"testing_generator: {testing_generator}")
# + [markdown] id="GN_ezwx7vBUo" colab_type="text"
# # Input
# + [markdown] id="EwyHfs4JvC7F" colab_type="text"
# Inputs to RNNs are sequential like text or time-series.
# + id="pSCOyGL_vPu5" colab_type="code" colab={}
from tensorflow.keras.layers import Input
# + id="JOs64ORxvBei" colab_type="code" outputId="22d20104-82af-4435-8c30-d93197c5eb5a" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Input
sequence_size = 8 # words per input
x = Input(shape=(sequence_size, EMBEDDING_DIM))
print (x)
# + [markdown] id="bUMibnFtuq_i" colab_type="text"
# # Simple RNN
# + [markdown] id="Ufxzi5G0DScl" colab_type="text"
# <img src="https://raw.githubusercontent.com/practicalAI/images/master/basic_ml/12_Recurrent_Neural_Networks/simple_rnn.png" width="500">
#
# RNN forward pass for a single time step $X_t$:
#
# $h_t = tanh(W_{hh}h_{t-1} + W_{xh}X_t+b_h)$
#
# *where*:
# * $W_{hh}$ = hidden units weights| $\in \mathbb{R}^{HXH}$ ($H$ is the hidden dim)
# * $h_{t-1}$ = previous timestep's hidden state $\in \mathbb{R}^{NXH}$
# * $W_{xh}$ = input weights| $\in \mathbb{R}^{EXH}$
# * $X_t$ = input at time step t | $\in \mathbb{R}^{NXE}$ ($N$ is the batch size, $E$ is the embedding dim)
# * $b_h$ = hidden units bias $\in \mathbb{R}^{HX1}$
# * $h_t$ = output from RNN for timestep $t$
#
# <div align="left">
# <img src="https://raw.githubusercontent.com/practicalAI/images/master/images/lightbulb.gif" width="45px" align="left" hspace="10px" vspace="10px">
# </div>
#
# At the first time step, the previous hidden state $h_{t-1}$ can either be a zero vector (unconditioned) or initialized (conditioned). If we are conditioning the RNN, the first hidden state $h_0$ can belong to a specific condition or we can concat the specific condition to the randomly initialized hidden vectors at each time step. More on this in the subsequent notebooks on RNNs.
# + id="_jJ5TmJ2ut3z" colab_type="code" colab={}
from tensorflow.keras.layers import SimpleRNN
# + id="z-O2Du78ut8J" colab_type="code" outputId="5f9fbd89-294f-4644-f0c1-09ea6d128b2a" colab={"base_uri": "https://localhost:8080/", "height": 51}
# RNN forward pass (many to one)
rnn = SimpleRNN(units=RNN_HIDDEN_DIM,
dropout=DROPOUT_P,
recurrent_dropout=RNN_DROPOUT_P,
return_sequences=False, # only get the output from the last sequential input
return_state=True)
output, hidden_state = rnn(x)
print (f"output {output.shape}")
print (f"hidden {hidden_state.shape}")
# + id="FqOtTdDx3Nz7" colab_type="code" outputId="7e99fcaf-aa6e-4911-8a04-44baa8950c22" colab={"base_uri": "https://localhost:8080/", "height": 51}
# RNN forward pass (many to many)
rnn = SimpleRNN(units=RNN_HIDDEN_DIM,
dropout=DROPOUT_P,
recurrent_dropout=RNN_DROPOUT_P,
return_sequences=True, # get outputs from every item in sequential input
return_state=True)
outputs, hidden_state = rnn(x)
print (f"output {outputs.shape}")
print (f"hidden {hidden_state.shape}")
# + [markdown] id="mnUh3iRpJ2j8" colab_type="text"
# There are many different ways to use RNNs. So far we've processed our inputs one timestep at a time and we could either use the RNN's output at each time step or just use the final input timestep's RNN output. Let's look at a few other possibilities.
#
# <img src="https://raw.githubusercontent.com/practicalAI/images/master/basic_ml/12_Recurrent_Neural_Networks/rnn_examples.png" width="1000">
# + [markdown] id="pfhjWZRD94hK" colab_type="text"
# # Model
# + [markdown] id="zVmJGm8m-KIz" colab_type="text"
# **Simple RNN**
# + id="T8oCutDJ-d1J" colab_type="code" colab={}
import tensorflow.keras.backend as K
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import Dropout
from tensorflow.keras.layers import Embedding
from tensorflow.keras.layers import Input
from tensorflow.keras.layers import Masking
from tensorflow.keras.losses import SparseCategoricalCrossentropy
from tensorflow.keras.models import Model
from tensorflow.keras.optimizers import Adam
# + [markdown] id="6B2MewCdCeKC" colab_type="text"
# ### Components
# + id="UPP5ROd69mXC" colab_type="code" colab={}
class TextClassificationRNNModel(Model):
def __init__(self, vocab_size, embedding_dim, rnn_cell,
hidden_dim, dropout_p, num_classes):
super(TextClassificationRNNModel, self).__init__()
# Embeddings
self.embedding = Embedding(input_dim=vocab_size,
output_dim=embedding_dim,
mask_zero=True,
trainable=True)
# Masking
self.mask = Masking(mask_value=0.)
# RNN
self.rnn = rnn_cell
# FC layers
self.fc1 = Dense(units=hidden_dim, activation='relu')
self.dropout = Dropout(rate=dropout_p)
self.fc2 = Dense(units=num_classes, activation='softmax')
def call(self, inputs, training=False):
"""Forward pass."""
# Inputs
x_in, seq_lengths = inputs
# Embed
x_emb = self.embedding(x_in)
# Masking
z = self.mask(x_emb)
# RNN
z, hidden_state = self.rnn(x_emb)
# Gather last relevant index
z = tf.gather_nd(z, K.cast(seq_lengths, 'int32'))
# FC
z = self.fc1(z)
if training:
z = self.dropout(z, training=training)
y_pred = self.fc2(z)
return y_pred
def sample(self, x_in_shape, seq_lengths_shape):
x_in = Input(shape=x_in_shape)
seq_lengths = Input(shape=seq_lengths_shape)
inputs = [x_in, seq_lengths]
return Model(inputs=inputs, outputs=self.call(inputs)).summary()
# + [markdown] id="ADmeEdSZj56Y" colab_type="text"
# In our model, we want to use the RNN's output after the last relevant token in the sentence is processed. The last relevant token doesn't refer the <PAD> tokens but to the last actual word in the sentence and its index is different for each input in the batch.
#
# This is why we included a `seq_lengths` variable in our `create_batch` function in the `Generator` class. The seq_lengths are passed in to the forward pass as an input and we use [tf.gather_nd](https://www.tensorflow.org/api_docs/python/tf/gather_nd) to gather the last relevant hidden state (before padding starts).
#
# ```
# def create_batch(self, batch_indices):
# ...
# # Sequence lengths
# seq_lengths = np.array([[i, len(x)-1] for i, x in enumerate(X)])
# ...
# return [X, seq_lengths], y
# ```
#
# Once we have the relevant hidden state extracted, we proceed to apply some fully-connected layers (with softmax) to generate the class probabilities.
# + [markdown] id="H39ggqTI5XfP" colab_type="text"
# # Inspection
# + [markdown] id="GJfY2QZbz2WL" colab_type="text"
# Let's say you want more transparency on the exact shapes of the inputs and outputs at every stage of your model. Well you could architect a model that just works and you can use model.sample() to see the summary but what if you have issues in the model in the first place?
# + id="1Rei-P5DRYFC" colab_type="code" colab={}
# Get the first data point
sample_inputs, sample_y = training_generator.create_batch([0])
# + id="aJOozQkzK5jA" colab_type="code" outputId="3c8f8342-8234-4752-bc6e-4c477477673a" colab={"base_uri": "https://localhost:8080/", "height": 68}
sample_X, sample_seq_length = sample_inputs
print (f"sample_X: {sample_X} ==> shape: {sample_X.shape}")
print (f"sample_seq_length: {sample_seq_length} ==> shape: {sample_seq_length.shape}")
print (f"sample_y: {sample_y} ==> shape: {sample_y.shape}")
# + [markdown] id="0_j0XG3h3DzZ" colab_type="text"
# Now we will create a smaller model to the point that we want to inspect. This is a great way to iteratively build your model, validating the output shapes at every step of the way.
# + id="2H6nmXq82GJo" colab_type="code" colab={}
class ModelToInspect(Model):
def __init__(self, vocab_size, embedding_dim, rnn_cell,
hidden_dim, dropout_p, num_classes):
super(ModelToInspect, self).__init__()
# Embeddings
self.embedding = Embedding(input_dim=vocab_size,
output_dim=embedding_dim,
mask_zero=True,
trainable=True)
# Masking
self.mask = Masking(mask_value=0.)
# RNN
self.rnn = rnn_cell
def call(self, inputs, training=False):
"""Forward pass."""
# Forward pass
x_in, seq_lengths = inputs
x_emb = self.embedding(x_in)
z = self.mask(x_emb)
z, hidden_state = self.rnn(x_emb)
return z
def sample(self, x_in_shape, seq_lengths_shape):
x_in = Input(shape=x_in_shape)
seq_lengths = Input(shape=seq_lengths_shape)
inputs = [x_in, seq_lengths]
return Model(inputs=inputs, outputs=self.call(inputs)).summary()
# + id="fkGjkJW02a2a" colab_type="code" colab={}
# RNN cell
simple_rnn = SimpleRNN(units=RNN_HIDDEN_DIM,
dropout=DROPOUT_P,
recurrent_dropout=RNN_DROPOUT_P,
return_sequences=True,
return_state=True)
# + id="dEOvGxPC2azp" colab_type="code" outputId="f07fcb89-5c6c-4083-bde8-8e63f46cdbcb" colab={"base_uri": "https://localhost:8080/", "height": 289}
model = ModelToInspect(vocab_size=vocab_size,
embedding_dim=EMBEDDING_DIM,
rnn_cell=simple_rnn,
hidden_dim=HIDDEN_DIM,
dropout_p=DROPOUT_P,
num_classes=len(classes))
model.sample(x_in_shape=(sample_X.shape[1],), seq_lengths_shape=(2,))
# + id="NJNTVi0dR8s-" colab_type="code" outputId="4eab615e-30b1-4cd9-85de-a5cbd27f1aa9" colab={"base_uri": "https://localhost:8080/", "height": 238}
z = model(sample_inputs)
print (f"z: {z} ==> shape: {z.shape}")
# + [markdown] id="QtqZKYks21ga" colab_type="text"
# The mode.sample() provided a decent summary of the shapes but with our close inspection strategy, we are able to see actual values for a specific input.
# + [markdown] id="S9ggYO6yHIm2" colab_type="text"
# # Training
# + id="geKOPVzVK6S9" colab_type="code" colab={}
from tensorflow.keras.callbacks import Callback
from tensorflow.keras.callbacks import EarlyStopping
from tensorflow.keras.callbacks import ReduceLROnPlateau
from tensorflow.keras.callbacks import TensorBoard
# %load_ext tensorboard
# + id="zsWl3U9UdiJ6" colab_type="code" colab={}
# RNN cell
simple_rnn = SimpleRNN(units=RNN_HIDDEN_DIM,
dropout=DROPOUT_P,
recurrent_dropout=RNN_DROPOUT_P,
return_sequences=True,
return_state=True)
# + id="wD4sRUS5_lwq" colab_type="code" outputId="7d2f8ab5-1b30-484b-821d-6cc516eaf857" colab={"base_uri": "https://localhost:8080/", "height": 442}
model = TextClassificationRNNModel(vocab_size=vocab_size,
embedding_dim=EMBEDDING_DIM,
rnn_cell=simple_rnn,
hidden_dim=HIDDEN_DIM,
dropout_p=DROPOUT_P,
num_classes=len(classes))
model.sample(x_in_shape=(sequence_size,), seq_lengths_shape=(2,))
# + id="Ucn3tYq1_sE1" colab_type="code" colab={}
# Compile
model.compile(optimizer=Adam(lr=LEARNING_RATE),
loss=SparseCategoricalCrossentropy(),
metrics=['accuracy'])
# + id="X5ac-uJXb-F7" colab_type="code" colab={}
# Callbacks
callbacks = [EarlyStopping(monitor='val_loss', patience=EARLY_STOPPING_CRITERIA, verbose=1, mode='min'),
ReduceLROnPlateau(patience=1, factor=0.1, verbose=0),
TensorBoard(log_dir='tensorboard/simple_rnn', histogram_freq=1, update_freq='epoch')]
# + id="qlqe2TVlCxvj" colab_type="code" outputId="12c59594-0934-4293-d953-3d7e545efe20" colab={"base_uri": "https://localhost:8080/", "height": 374}
# Training
training_history = model.fit_generator(generator=training_generator,
epochs=NUM_EPOCHS,
validation_data=validation_generator,
callbacks=callbacks,
shuffle=False,
class_weight=class_weights,
verbose=1)
# + id="fylHduxTK-0N" colab_type="code" outputId="656639a9-1e4e-46e3-dc85-e9112ba4f690" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Evaluation
testing_history = model.evaluate_generator(generator=testing_generator,
verbose=1)
# + id="gaJD2YGpcTfT" colab_type="code" outputId="ed0f00a4-2164-460d-94f0-57a0778953f2" colab={"base_uri": "https://localhost:8080/", "height": 17}
# %tensorboard --logdir tensorboard
# + [markdown] id="lFWSgL8E5_HU" colab_type="text"
# # Bidirectional RNN
# + id="sO_vP00z6BiJ" colab_type="code" colab={}
from tensorflow.keras.layers import Bidirectional
# + id="WqtGrVhm7914" colab_type="code" colab={}
class TextClassificationBiRNNModel(Model):
def __init__(self, vocab_size, embedding_dim, rnn_cell,
hidden_dim, dropout_p, num_classes):
super(TextClassificationBiRNNModel, self).__init__()
# Embeddings
self.embedding = Embedding(input_dim=vocab_size,
output_dim=embedding_dim,
mask_zero=True,
trainable=True)
# Masking
self.mask = Masking(mask_value=0.)
# RNN
self.rnn = Bidirectional(rnn_cell, merge_mode='sum')
# FC layers
self.fc1 = Dense(units=hidden_dim, activation='relu')
self.dropout = Dropout(rate=dropout_p)
self.fc2 = Dense(units=num_classes, activation='softmax')
def call(self, inputs, training=False):
"""Forward pass."""
# Inputs
x_in, seq_lengths = inputs
# Embed
x_emb = self.embedding(x_in)
# Masking
z = self.mask(x_emb)
# RNN
z, hidden_state_fw, hidden_state_bw = self.rnn(x_emb)
# Gather last relevant index
z = tf.gather_nd(z, K.cast(seq_lengths, 'int32'))
# FC
z = self.fc1(z)
if training:
z = self.dropout(z, training=training)
y_pred = self.fc2(z)
return y_pred
def sample(self, x_in_shape, seq_lengths_shape):
x_in = Input(shape=x_in_shape)
seq_lengths = Input(shape=seq_lengths_shape)
inputs = [x_in, seq_lengths]
return Model(inputs=inputs, outputs=self.call(inputs)).summary()
# + id="yzz9ctXv6BmZ" colab_type="code" colab={}
# Bidirectional RNN cell
simple_rnn = SimpleRNN(units=RNN_HIDDEN_DIM,
dropout=DROPOUT_P,
recurrent_dropout=RNN_DROPOUT_P,
return_sequences=True,
return_state=True)
# + id="FkqYRBWd6OCR" colab_type="code" outputId="14e5bdd0-b1fe-4906-842e-2c7c6ad055b8" colab={"base_uri": "https://localhost:8080/", "height": 442}
model = TextClassificationBiRNNModel(vocab_size=vocab_size,
embedding_dim=EMBEDDING_DIM,
rnn_cell=simple_rnn,
hidden_dim=HIDDEN_DIM,
dropout_p=DROPOUT_P,
num_classes=len(classes))
model.sample(x_in_shape=(sequence_size,), seq_lengths_shape=(2,))
# + id="O_XtqY6n6Qdh" colab_type="code" colab={}
# Compile
model.compile(optimizer=Adam(lr=LEARNING_RATE),
loss=SparseCategoricalCrossentropy(),
metrics=['accuracy'])
# + id="k07ilI7K6N_v" colab_type="code" colab={}
# Callbacks
callbacks = [EarlyStopping(monitor='val_loss', patience=EARLY_STOPPING_CRITERIA, verbose=1, mode='min'),
ReduceLROnPlateau(patience=1, factor=0.1, verbose=0),
TensorBoard(log_dir='tensorboard/simple_birnn', histogram_freq=1, update_freq='epoch')]
# + id="uuZwiGRt6X-w" colab_type="code" outputId="25bc5ff6-e534-41b3-aaa0-7efd5ff9f1f9" colab={"base_uri": "https://localhost:8080/", "height": 306}
# Training
training_history = model.fit_generator(generator=training_generator,
epochs=NUM_EPOCHS,
validation_data=validation_generator,
callbacks=callbacks,
shuffle=False,
class_weight=class_weights,
verbose=1)
# + [markdown] id="jR3trOIQcWY2" colab_type="text"
# # Gated RNNs: LSTMs & GRUs
# + [markdown] id="IqFsufNVch4W" colab_type="text"
# While our simple RNNs so far are great for sequentially processing our inputs, they have quite a few disadvantages. They commonly suffer from exploding or vanishing gradients as a result using the same set of weights ($W_{xh}$ and $W_{hh}$) with each timestep's input. During backpropagation, this can cause gradients to explode (>1) or vanish (<1). If you multiply any number greater than 1 with itself over and over, it moves towards infinity (exploding gradients) and similarly, If you multiply any number less than 1 with itself over and over, it moves towards zero (vanishing gradients). To mitigate this issue, gated RNNs were devised to selectively retrain information. If you're interested in learning more of the specifics, this [post](http://colah.github.io/posts/2015-08-Understanding-LSTMs/) is a must-read.
#
# There are two popular types of gated RNNs: Long Short-term Memory (LSTMs) units and Gated Recurrent Units (GRUs).
#
# <img src="https://raw.githubusercontent.com/practicalAI/images/master/basic_ml/12_Recurrent_Neural_Networks/gated_rnns.png" width="600"><br>
# <a href="http://colah.github.io/posts/2015-08-Understanding-LSTMs/">Understanding LSTM Networks</a> - Chris Olah
#
#
# <div align="left">
# <img src="https://raw.githubusercontent.com/practicalAI/images/master/images/lightbulb.gif" width="45px" align="left" hspace="10px">
# </div>
#
# When deciding between LSTMs and GRUs, empirical performance is the best factor but in genreal GRUs offer similar perforamnce with less complexity (less weights).
#
#
# + id="PqVCqWv2fz4N" colab_type="code" colab={}
from tensorflow.keras.layers import GRU
# + id="7tBHovnxcTwV" colab_type="code" colab={}
# RNN cell
gru = GRU(units=RNN_HIDDEN_DIM,
dropout=DROPOUT_P,
recurrent_dropout=RNN_DROPOUT_P,
return_sequences=True,
return_state=True)
# + id="n7BfnoAbcTtk" colab_type="code" outputId="0aa920b0-0e17-4257-929c-599d74882ac7" colab={"base_uri": "https://localhost:8080/", "height": 442}
model = TextClassificationBiRNNModel(vocab_size=vocab_size,
embedding_dim=EMBEDDING_DIM,
rnn_cell=gru,
hidden_dim=HIDDEN_DIM,
dropout_p=DROPOUT_P,
num_classes=len(classes))
model.sample(x_in_shape=(sequence_size,), seq_lengths_shape=(2,))
# + id="Vd8SL8UIcTqc" colab_type="code" colab={}
# Compile
model.compile(optimizer=Adam(lr=LEARNING_RATE),
loss=SparseCategoricalCrossentropy(),
metrics=['accuracy'])
# + id="c8dNn6VPcTkx" colab_type="code" colab={}
# Callbacks
callbacks = [EarlyStopping(monitor='val_loss', patience=EARLY_STOPPING_CRITERIA, verbose=1, mode='min'),
ReduceLROnPlateau(patience=1, factor=0.1, verbose=0),
TensorBoard(log_dir='tensorboard/gru', histogram_freq=1, update_freq='epoch')]
# + id="NbpjRVficTcY" colab_type="code" outputId="db96b684-6b21-415e-cebf-b755210cbcf2" colab={"base_uri": "https://localhost:8080/", "height": 340}
# Training
training_history = model.fit_generator(generator=training_generator,
epochs=NUM_EPOCHS,
validation_data=validation_generator,
callbacks=callbacks,
shuffle=False,
class_weight=class_weights,
verbose=1)
# + id="4ytG83xxcTZp" colab_type="code" outputId="5cb94c46-9189-4474-efa2-f9ec42e592e5" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Evaluation
testing_history = model.evaluate_generator(generator=testing_generator,
verbose=1)
# + id="0eXhh4rqcTXE" colab_type="code" outputId="c30a94d4-ac30-46d2-e711-7a8912408557" colab={"base_uri": "https://localhost:8080/", "height": 34}
# %tensorboard --logdir tensorboard
# + [markdown] id="vskwiiI3V3S6" colab_type="text"
# # Evaluation
# + id="Itq7lT9qV9Y8" colab_type="code" colab={}
import io
import itertools
import json
import matplotlib.pyplot as plt
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.metrics import precision_recall_fscore_support
# + [markdown] id="175J-ELiV-I6" colab_type="text"
# ### Components
# + id="VT9C8rUyWAWF" colab_type="code" colab={}
def plot_confusion_matrix(y_true, y_pred, classes, cmap=plt.cm.Blues):
"""Plot a confusion matrix using ground truth and predictions."""
# Confusion matrix
cm = confusion_matrix(y_true, y_pred)
cm_norm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
# Figure
fig = plt.figure()
ax = fig.add_subplot(111)
cax = ax.matshow(cm, cmap=plt.cm.Blues)
fig.colorbar(cax)
# Axis
plt.title("Confusion matrix")
plt.ylabel("True label")
plt.xlabel("Predicted label")
ax.set_xticklabels([''] + classes)
ax.set_yticklabels([''] + classes)
ax.xaxis.set_label_position('bottom')
ax.xaxis.tick_bottom()
# Values
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, f"{cm[i, j]:d} ({cm_norm[i, j]*100:.1f}%)",
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
# Display
plt.show()
# + id="OyguCCS3W6DO" colab_type="code" colab={}
def get_performance(y_true, y_pred, classes):
"""Per-class performance metrics."""
performance = {'overall': {}, 'class': {}}
y_pred = np.argmax(y_pred, axis=1)
metrics = precision_recall_fscore_support(y_true, y_pred)
# Overall performance
performance['overall']['precision'] = np.mean(metrics[0])
performance['overall']['recall'] = np.mean(metrics[1])
performance['overall']['f1'] = np.mean(metrics[2])
performance['overall']['num_samples'] = np.float64(np.sum(metrics[3]))
# Per-class performance
for i in range(len(classes)):
performance['class'][classes[i]] = {
"precision": metrics[0][i],
"recall": metrics[1][i],
"f1": metrics[2][i],
"num_samples": np.float64(metrics[3][i])
}
return performance
# + [markdown] id="yCAHPsyWWAar" colab_type="text"
# ### Operations
# + id="MHfGZh7JZGe6" colab_type="code" outputId="211b0e00-2ee5-4c6a-a49f-29fca1d908cd" colab={"base_uri": "https://localhost:8080/", "height": 68}
# Evaluation
test_history = model.evaluate_generator(generator=testing_generator, verbose=1)
y_pred = model.predict_generator(generator=testing_generator, verbose=1)
print (f"test history: {test_history}")
# + id="qdAj6KyCU88E" colab_type="code" outputId="fe62cde7-1111-4518-9002-45d4c700c4a6" colab={"base_uri": "https://localhost:8080/", "height": 595}
# Class performance
performance = get_performance(y_true=y_test,
y_pred=y_pred,
classes=classes)
print (json.dumps(performance, indent=4))
# + id="nRbPfqgZWaof" colab_type="code" outputId="4e1da3b1-fc78-48f6-a0c6-bf58cab626c1" colab={"base_uri": "https://localhost:8080/", "height": 598}
# Confusion matrix
plt.rcParams["figure.figsize"] = (7,7)
y_pred = np.argmax(y_pred, axis=1)
plot_confusion_matrix(y_test, y_pred, classes=classes)
print (classification_report(y_test, y_pred))
# + [markdown] id="yeiD1T_QZpdk" colab_type="text"
# # Inference
# + id="z7G7vuSTZHkQ" colab_type="code" colab={}
import collections
# + [markdown] id="Xeu952p4Zweb" colab_type="text"
# ### Components
# + id="1GiJpWeZZwkp" colab_type="code" colab={}
def get_probability_distributions(probabilities, classes):
"""Produce probability distributions with labels."""
probability_distributions = []
for i, y_prob in enumerate(probabilities):
probability_distribution = {}
for j, prob in enumerate(y_prob):
probability_distribution[classes[j]] = np.float64(prob)
probability_distribution = collections.OrderedDict(
sorted(probability_distribution.items(), key=lambda kv: kv[1], reverse=True))
probability_distributions.append(probability_distribution)
return probability_distributions
# + [markdown] id="6EZdo-fKZwo6" colab_type="text"
# ### Operations
# + id="CLP2Vzp3Zwth" colab_type="code" outputId="f582883f-36fc-41e7-a5a7-d094f0294bda" colab={"base_uri": "https://localhost:8080/", "height": 85}
# Inputs
texts = ["This weekend the greatest tennis players will fight for the championship."]
num_samples = len(texts)
X_infer = np.array(X_tokenizer.texts_to_sequences(texts))
print (f"{texts[0]} \n\t→ {untokenize(X_infer[0], X_tokenizer)} \n\t→ {X_infer[0]}")
print (f"len(X_infer[0]): {len(X_infer[0])} characters")
y_filler = np.array([0]*num_samples)
# + id="q1gFlI5MZ143" colab_type="code" colab={}
# Inference data generator
inference_generator = DataGenerator(X=X_infer,
y=y_filler,
batch_size=BATCH_SIZE,
shuffle=False)
# + id="UFE4sp_7aHTq" colab_type="code" outputId="8c403d7b-7359-4f81-898b-8d82fb3a6ea7" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Predict
probabilities = model.predict_generator(generator=inference_generator,
verbose=1)
# + id="bGi_NvbBaMap" colab_type="code" outputId="84ff5775-009a-45a3-81ea-f8aa9dc1bda4" colab={"base_uri": "https://localhost:8080/", "height": 238}
# Results
probability_distributions = get_probability_distributions(probabilities=probabilities,
classes=y_tokenizer.classes_)
results = []
for index in range(num_samples):
results.append({
'raw_input': texts[index],
'preprocessed_input': untokenize(indices=X_infer[index], tokenizer=X_tokenizer),
'tokenized_input': str(X_infer[index]),
'probabilities': probability_distributions[index]
})
print (json.dumps(results, indent=4))
# + [markdown] id="4xknbwa-bW3U" colab_type="text"
# <img width="45" src="http://bestanimations.com/HomeOffice/Lights/Bulbs/animated-light-bulb-gif-29.gif" align="left" vspace="5px" hspace="10px">
#
# We will learn how to get a little bit of interpretabiltiy with RNNs in the next lesson on attentional interfaces.
# + [markdown] id="2I4XMUfsILku" colab_type="text"
# ---
# <div align="center">
#
# Subscribe to our <a href="https://practicalai.me/#newsletter">newsletter</a> and follow us on social media to get the latest updates!
#
# <a class="ai-header-badge" target="_blank" href="https://github.com/practicalAI/practicalAI">
# <img src="https://img.shields.io/github/stars/practicalAI/practicalAI.svg?style=social&label=Star"></a>
# <a class="ai-header-badge" target="_blank" href="https://www.linkedin.com/company/madewithml">
# <img src="https://img.shields.io/badge/style--5eba00.svg?label=LinkedIn&logo=linkedin&style=social"></a>
# <a class="ai-header-badge" target="_blank" href="https://twitter.com/madewithml">
# <img src="https://img.shields.io/twitter/follow/madewithml.svg?label=Follow&style=social">
# </a>
# </div>
#
# </div>
| notebooks/basic_ml/12_Recurrent_Neural_Networks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import cv2
import numpy as np
import matplotlib.pyplot as plt
# Black and White (gray scale)
img = cv2.imread ('cat.jpg',0)
img1 = cv2.imread('cat.jpg',1)
resized_image = cv2.resize(img, (650,500))
resized_image1 = cv2.resize(img1, (650,500))
cv2.imshow('cat1',resized_image1)
cv2.imshow('cat', resized_image)
plt.imshow(resized_image)
plt.imshow(resized_image1)
# -
import numpy as np
import matplotlib.pyplot as plt
import cv2
# %matplotlib inline
#reading the image
image = cv2.imread('cat.jpg')
image = cv2.cvtColor(image,cv2.COLOR_BGR2GRAY)
#plotting the image
plt.imshow(image)
plt.imshow(img1)
plt.imshow(img)
plt.imshow(resized_image)
plt.imshow(resized_image1)
| MANOJKUMAR_ OPEN_CV.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 继续挑战
#
# ---
# ### 第8题地址[integrity.html](http://www.pythonchallenge.com/pc/def/integrity.html)
# * <img src="http://www.pythonchallenge.com/pc/def/integrity.jpg" alt="integrity.jpg" width="30%" height="30%">
# * 网页标题是`working hard?`,题目是`Where is the missing link?`,[源码](view-source:http://www.pythonchallenge.com/pc/def/integrity.html)中有两行隐藏信息:
# > un: 'BZh91AY&SYA\xaf\x82\r\x00\x00\x01\x01\x80\x02\xc0\x02\x00 \x00!\x9ah3M\x07<]\xc9\x14\xe1BA\x06\xbe\x084'<br>
# > pw: 'BZh91AY&SY\x94\$|\x0e\x00\x00\x00\x81\x00\x03\$ \x00!\x9ah3M\x13<]\xc9\x14\xe1BBP\x91\xf08'
# 图片上是以一个`<area>`区域画出来一个超链接,点击之后提示要输入用户名和密码,提示是`inflate`。这应该是题目的基本意思。<br>
# 而这个区域刚好描绘的是图中的蜜蜂,结合`bee`和网页标题`working hard`,应该意思是`busy`。<br>
# 再看看源码中的隐藏信息,应该指的是`username`和`password`,估计就是提示要输入的部分。
# 直接拿这一堆乱码去试肯定是行不通的<del>别说,我还真试了几次</del>。<br>
# 正确做法应该是要解读这个`busy`的意思,结合这个乱码(`BZh`开头),应该指的是`bzip2`压缩编码,python对应有`bz2`的包可以简易处理。
from bz2 import decompress
print('username:', decompress(b'BZh91AY&SYA\xaf\x82\r\x00\x00\x01\x01\x80\x02\xc0\x02\x00 \x00!\x9ah3M\x07<]\xc9\x14\xe1BA\x06\xbe\x084').decode())
print('password:', decompress(b'B<PASSWORD>\x94$|\x0e\x00\x00\x00\x81\x00\x03$ \x00!\x9ah3M\x13<]\xc9\x14\xe1BBP\x91\xf08').decode())
# 这个结果符合`inflate`的提示。将解码后的结果输入到提示框里面点确定,来到下一题[good.html](http://www.pythonchallenge.com/pc/return/good.html)
# ### 总结:题目跟图像处理无关,倒是又介绍了一种压缩编码和对应的python包。
# ###### 本题代码地址[8_integrity.ipynb](https://github.com/StevenPZChan/pythonchallenge/blob/notebook/nbfiles/8_integrity.ipynb)
| nbfiles/8_integrity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="images/logo.jpg" style="display: block; margin-left: auto; margin-right: auto;" alt="לוגו של מיזם לימוד הפייתון. נחש מצויר בצבעי צהוב וכחול, הנע בין האותיות של שם הקורס: לומדים פייתון. הסלוגן המופיע מעל לשם הקורס הוא מיזם חינמי ללימוד תכנות בעברית.">
# # <span style="text-align: right; direction: rtl; float: right; clear: both;"> תרגילים </span>
# ## <span style="text-align: right; direction: rtl; float: right; clear: both;">תרגיל מחשמל</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# בהנדסת חשמל, "שער לוגי" הוא רכיב בסיסי ולו כניסה אחת או יותר, ויציאה אחת בלבד.<br>
# הכניסות בשער הלוגי משמשות בתור הקלט שלו, והיציאה בתור הפלט.<br>
# כל כניסה יכולה לקבל את הערך "אמת" או "שקר".<br>
# השער הלוגי יבצע פעולה לוגית בהתאם לכניסות שלו, ויחזיר ערך חדש – שגם הוא אמת או שקר.
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# שער לוגי שוודאי יצא לכם לשמוע עליו הוא <dfn>AND</dfn>.<br>
# לשער הלוגי AND שתי כניסות. פלט השער יהיה <var>True</var> רק אם הקלט בשתי הכניסות הוא <var>True</var>, אחרת – יהיה הפלט <var>False</var>.<br>
# שער לוגי אחר שוודאי יצא לכם לשמוע עליו הוא <dfn>NOT</dfn>.<br>
# לשער הלוגי NOT כניסה אחת. פלט השער יהיה <var>True</var> אם הקלט הוא <var>False</var>, או <var>False</var> אם הקלט הוא <var>True</var>.
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# שערים עם כניסה אחת נקראים שערים אוּנָרִיִּים (unary), ושערים עם שתי כניסות נקראים שערים בִּינָרִיִּים (binary).<br>
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# חיבור של כמה שערים לוגיים אחד לשני ייצור רכיב לוגי.<br>
# בדוגמה הבאה, הקלטים (מלמעלה למטה) <var>True</var>, <var>False</var>, <var>False</var>, <var>True</var> יניבו <var>True</var>.
# </p>
# <figure>
# <img src="images/circuit_example.svg" style="width: 250px; margin-right: auto; margin-left: auto; text-align: center;" alt="דוגמה למעגל אלקטרוני. שער AND מצויר בצורת חצי אליפסה פחוסה לא סגורה, המחוברת משמאל למלבן שצלעו הימנית לא סגורה. שער OR מצויר כמו שער AND, רק שצלעו השמאלית נכנסת מעט פנימה בצורה מעוגלת. לשני השערים הלוגיים נכנסים 2 קווים משמאל, ומימינם יוצא קו אחד. המעגל נראה כך: בטור השמאלי, ישנו שער OR למעלה ושער AND למטה. בטור הימני ישנו שער OR. הפלט של העמודה השמאלית נכנס כקלט לעמודה הימנית."/>
# <figcaption style="margin-top: 2rem; text-align: center; direction: rtl;">
# דוגמה לרכיב לוגי.
# </figcaption>
# </figure>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# צרו את המחלקות <var>LogicGate</var>, <var>UnaryGate</var>, <var>BinaryGate</var> ו־<var>Connector</var>.<br>
# כמו כן, צרו את המחלקות <var>AndGate</var>, <var>NotGate</var>, <var>OrGate</var>, <var>NandGate</var> ו־<var>XorGate</var>.<br>
# במידת הצורך, קראו עוד על שערים לוגיים.
# </p>
# +
class OrGate:
def output(a,b):
return a or b
class AndGate:
def output(a,b):
return a and b
class NotGate:
def output(a):
return not a
class Connector:
def __init__(self, gate, input_gates):
self.gate = gate
self.input_gates = input_gates
def send_inputs(self,lst):
return self.gate.output(self.input_gates[0].output(lst[0],lst[1]) , self.input_gates[1].output(lst[2],lst[3]))
OrGate.output(True,False)
# -
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# לאחר שכתבתם את הקוד, ודאו שהקוד הבא, שמדמה את המעגל מהאיור למעלה, מדפיס רק True:
# </p>
circuit = Connector(gate=OrGate, input_gates=[OrGate, AndGate])
print(circuit.send_inputs([False, False, False, False]) == False)
print(circuit.send_inputs([False, False, False, True]) == False)
print(circuit.send_inputs([False, False, True, False]) == False)
print(circuit.send_inputs([False, False, True, True]) == True)
print(circuit.send_inputs([False, True, False, False]) == True)
print(circuit.send_inputs([False, True, False, True]) == True)
print(circuit.send_inputs([False, True, True, False]) == True)
print(circuit.send_inputs([False, True, True, True]) == True)
print(circuit.send_inputs([True, False, False, False]) == True)
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# בונוס: ודאו גם שהקוד הבא עובד ומדפיס <var>True</var>:
# </p>
# <figure>
# <img src="images/complex_circuit.svg" style="width: 400px; margin-right: auto; margin-left: auto; text-align: center;" alt="החידוש באיור זה הוא שער NOT, שמצויר כמשולש שבקודקודו הימני מצויר עיגול קטן."/>
# <figcaption style="margin-top: 2rem; text-align: center; direction: rtl;">
# דוגמה לרכיב לוגי מורכב יותר.
# </figcaption>
# </figure>
# +
class OrGate:
def output(lst):
res = lst[-1] or lst[-2]
lst.pop()
lst.pop()
lst.append(res)
return res
class AndGate:
def output(lst):
res = lst[-1] and lst[-2]
lst.pop()
lst.pop()
lst.append(res)
return res
class NotGate:
def output(lst):
res = not lst[-1]
lst.pop()
lst.append(res)
return res
class Connector:
def __init__(self, gate, input_gates):
self.gate = gate
self.input_gates = input_gates
def send_inputs(self,lst):
for gate in self.input_gates:
gate.output(lst)
res = self.gate.output(lst)
return res
def output(self,lst):
return self.send_inputs(lst)
OrGate.output([True,False])
# +
first_connector = Connector(
gate=OrGate, input_gates=[OrGate, AndGate],
)
second_connector = Connector(
gate=OrGate,
input_gates=[first_connector],
)
circuit = Connector(
gate=OrGate,
input_gates=[second_connector, NotGate],
)
inputs = [False, False, False, True, False, True]
print(circuit.send_inputs(inputs) == False)
# -
# ## <span style="text-align: right; direction: rtl; float: right; clear: both;">ימצי</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# כתבו מחלקה המייצגת קובייה.<br>
# לכל קובייה יש צבע, מספר פאות וערכים שמופיעים על הפאות.<br>
# הטלה של קובייה תבחר באופן אקראי ערך מאחת מפאות הקובייה ותחזיר אותו.
# </p>
#
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# על כל פאה של "קובייה מספרית" מופיע מספר שערכו בין 1 למספר הפאות בקובייה. כל מספר יופיע פעם אחת בלבד על הקובייה.<br>
# ב"קובייה לא מאוזנת" ישנו גם ערך שנקרא "סיכוי הטלה", שמפרט עבור כל פאה מה הסיכוי שתצא בהטלה.
# </p>
#
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# שם הצבע של הקובייה חייב להיות מורכב מאותיות בלבד, סיכויי ההטלה חייבים להיות חיוביים ומספר הפאות חייב להיות תואם למספר הערכים המופיעים על הפאות.<br>
# </p>
#
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# במשחק "Yamtzee" יש שק גדול שמכיל המון "קוביות אורך", ושק גדול נוסף של קוביות לא מאוזנות בצבעים שונים שנקראות "קוביות אות".<br>
# 10% מהקוביות בכל שק הן אדומות, 25% הן ירוקות ו־65% הן כחולות.<br>
# קוביית אורך היא קובייה מספרית עם 10 פאות. אם הקובייה מורה 1, זורקים אותה שוב עד שיוצא מספר אחר.<br>
# קוביית אות היא קובייה לא מאוזנת עם 26 פאות, כאשר בכל פאה אות מהא"ב האנגלי. סיכוי ההטלה עבור כל אחד מהערכים הוא לפי <a href="https://en.wikipedia.org/wiki/Letter_frequency">התדירות של האות</a>.<br>
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# בתחילת המשחק, המשתמש בוחר את מספר המשתתפים במשחק, ומה הניקוד שאליו צריך להגיע כדי שהמשחק יסתיים.<br>
# </p>
#
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# כל שחקן בתורו מטיל את קוביית האורך, ומטיל מספר קוביות אות הזהה למספר שהתקבל בקוביית האורך.<br>
# לדוגמה, אם קוביית האורך שהטלתי מורה 3, עליי להטיל 3 קוביות אות.
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# המשתתף צריך להשתמש בקוביות האות שיצאו לו כדי ליצור מילה תקנית בשפה האנגלית, שאורכה 2 אותיות לפחות.<br>
# הוא יכול לסדר את הקוביות מחדש ולבחור שלא להשתמש בחלק מהן, אבל הוא לא יכול להשתמש באותה קוביית אות פעמיים.<br>
# למרות זאת, יכול לקרות מצב שיותר מקוביית אות אחת תציג את אותה האות.
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# אם המשתמש הצליח להרכיב מילה, עבור כל אות שבה השתמש הוא זוכה ב־$\lfloor{\frac{12}{\lceil{f}\rceil^{\frac{3}{4}}}}\rfloor$ נקודות, כאשר f זו תדירות האות בא"ב האנגלי, מעוגלת כלפי מעלה.<br>
# במילים: 12 חלקי הביטוי הבא – הסיכוי שהאות תצא מעוגל כלפי מעלה, בחזקת 1.5 ואז להוציא מזה שורש. כל זה – מעוגל כלפי מטה.<br>
# לדוגמה, עבור המילה "zone" יזכה השחקן ב־17 נקודות לפי החישוב הזה:
# </p>
# | ניקוד | תדירות | אות |
# |:----------|:----------|------:|
# | 12 | 0.077% | z |
# | 2 | 7.507% | o |
# | 2 | 6.749% | n |
# | 1 | 12.702% | e |
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# קוביית אורך אדומה מעניקה למשתתף שהטיל אותה עוד קוביית אות במתנה.<br>
# קוביית אות אדומה מאפשרת למשתתף להשתמש באות שמופיעה על הקובייה כמה פעמים שירצה.<br>
# קוביות ירוקות נותנות למשתתף את האפשרות לבחור אם להטילן מחדש.<br>
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# ממשו את Yamtzee.<br>
# השתמשו ב־<a href="https://raw.githubusercontent.com/dwyl/english-words/master/words.txt">words.txt</a> כדי לוודא שהמילים שהכניס המשתמש תקינות.
# </p>
| week8/5_Summary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import math
cos=math.cos(0)
sen=math.sin(0)
a=0
b=0.00001
c=0.0001
d=0.001
e=0.01
f=0.1
g=1
h=10
x1=math.cos(0)
x2=math.cos(0.00001)
x3=math.cos(0.0001)
x4=math.cos(0.001)
x5=math.cos(0.01)
x6=math.cos(0.1)
x7=math.cos(1)
x8=math.cos(10)
a1=cos
b1=cos
c1=cos
d1=cos
e1=cos
f1=cos
g1=cos
h1=cos
a2=cos+sen
b2=cos+sen
c2=cos+sen
d2=cos+sen
e2=cos+sen
f2=cos+sen
g2=cos+sen
h2=cos+sen
a3=cos+sen-((a**2)/2)
b3=cos+sen-((b**2)/2)
c3=cos+sen-((c**2)/2)
d3=cos+sen-((d**2)/2)
e3=cos+sen-((e**2)/2)
f3=cos+sen-((f**2)/2)
g3=cos+sen-((g**2)/2)
h3=cos+sen-((h**2)/2)
a4=cos+sen-((a**2)/2)
b4=cos+sen-((b**2)/2)
c4=cos+sen-((c**2)/2)
d4=cos+sen-((d**2)/2)
e4=cos+sen-((e**2)/2)
f4=cos+sen-((f**2)/2)
g4=cos+sen-((g**2)/2)
h4=cos+sen-((h**2)/2)
a5=cos+sen-((a**2)/2)+0+((a**4)/24)
b5=cos+sen-((b**2)/2)+0+((b**4)/24)
c5=cos+sen-((c**2)/2)+0+((c**4)/24)
d5=cos+sen-((d**2)/2)+0+((d**4)/24)
e5=cos+sen-((e**2)/2)+0+((e**4)/24)
f5=cos+sen-((f**2)/2)+0+((f**4)/24)
g5=cos+sen-((g**2)/2)+0+((g**4)/24)
h5=cos+sen-((h**2)/2)+0+((h**4)/24)
print('x || Exacto ||Aprox.1 || Aprox.2 || Aprox.3 || Aprox.4 || Aprox.5 || ')
print('0 ||',x1,' ||',a1,' ||',a2,' ||',a3,' ||',a4,' ||',a5,' || ')
print('0.00001 ||',x2,' ||',b1,' ||',b2,' ||',b3,'||',b4,'||',b5,' || ')
print('0.0001 ||',x3,' ||',c1,' ||',c2,' ||',c3,' ||',c4,' ||',c5,' || ')
print('0.001 ||',x4,' ||',d1,' ||',d2,' ||',d3,' ||',d4,' ||',d5,' || ')
print('0.01 ||',x5,' ||',e1,' ||',e2,' ||',e3,' ||',e4,' ||',e5,' || ')
print('0.1 ||',x6,' ||',f1,' ||',f2,' ||',f3,' ||',f4,' ||',f5,' || ')
print('1 ||',x7,' ||',g1,' ||',g2,' ||',g3,' ||',g4,' ||',g5,' || ')
print('10 ||',x8,'||',h1,' ||',h2,' ||',h3,' ||',h4,' ||',h5,' || ')
x = float(input("Ingresar valor deseado de la tabla: "))
aproximacion = float(input("Ingresar aproximación deseada de la tabla: "))
if(aproximacion==1):
z=cos
print(z)
if(aproximacion==2):
z=cos+sen
print(z)
if(aproximacion==3):
z=cos+sen-((x**2)/2)
print(z)
if(aproximacion==4):
z=cos+sen-((x**2)/2)
print(z)
if(aproximacion==5):
z=cos+sen-((x**2)/2)+0+((x**4)/24)
print(z)
# -
# <NAME> - 161003313
# <NAME> - 161003307
| tarea2/tarea2-01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/pawan-nandakishore/image_emotion_recognition/blob/master/colab_notebooks/three_model_-ensemble2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="T0or5NdyMylK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e863dcf4-e9f1-4ef1-b5cc-a2437cf97f89"
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
from sklearn.metrics import accuracy_score
import tensorflow as tf
from tensorflow.keras.layers import *
from tensorflow.python.lib.io import file_io
from skimage.transform import rescale, resize
physical_devices = tf.config.experimental.list_physical_devices('GPU')
if physical_devices:
tf.config.experimental.set_memory_growth(physical_devices[0], True)
print(physical_devices)
# + id="Q3e1Mj7tQ15R" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 122} outputId="4c5aa2d9-3d03-4714-d648-d289d80cf7b9"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="Q4xTUVfHP2hP" colab_type="text"
#
# From Turker's ensemble training it seems like the top three models are:
#
# - ResNet-BEST-73.2.h5 : 73.5 % <br>
# - RESNET50-AUX-BEST-72.7.h5: 72.5 % <br>
# - SENET50-AUX-BEST-72.5.h5: 72.5 %
# The attempt will be to combine these three models and see what type of accuracies one can get.
# + id="wfCQilOGP9U3" colab_type="code" colab={}
# Function that reads the data from the csv file, increases the size of the images and returns the images and their labels
# dataset: Data path
IMAGE_SIZE = 48
def get_data(dataset):
file_stream = file_io.FileIO(dataset, mode='r')
data = pd.read_csv(file_stream)
data[' pixels'] = data[' pixels'].apply(lambda x: [int(pixel) for pixel in x.split()])
X, Y = data[' pixels'].tolist(), data['emotion'].values
X = np.array(X, dtype='float32').reshape(-1,IMAGE_SIZE, IMAGE_SIZE,1)
X = X/255.0
X_res = np.zeros((X.shape[0], Resize_pixelsize,Resize_pixelsize,3))
for ind in range(X.shape[0]):
sample = X[ind]
sample = sample.reshape(IMAGE_SIZE, IMAGE_SIZE)
image_resized = resize(sample, (Resize_pixelsize, Resize_pixelsize), anti_aliasing=True)
X_res[ind,:,:,:] = image_resized.reshape(Resize_pixelsize,Resize_pixelsize,1)
Y_res = np.zeros((Y.size, 7))
Y_res[np.arange(Y.size),Y] = 1
return X, X_res, Y_res
# + id="-hsrzMOCR2cF" colab_type="code" colab={}
Resize_pixelsize = 197
dev_dataset_dir = '/content/drive/My Drive/Personal projects/emotion_recognition_paper/data/fer_csv/dev.csv'
test_dataset_dir = '/content/drive/My Drive/Personal projects/emotion_recognition_paper/data/fer_csv/test.csv'
X_dev, X_res_dev, Y_dev = get_data(dev_dataset_dir)
X_test, X_res_test, Y_test = get_data(test_dataset_dir)
# + [markdown] id="8jjdFX6tTm5R" colab_type="text"
# ## Model 1: ACC 72.5%
# + id="-BsnKbpjQEHr" colab_type="code" colab={}
Senet_model_AUX = tf.keras.models.load_model('/content/drive/My Drive/Personal projects/emotion_recognition_paper/models/SENET50-AUX-BEST-72.5.h5')
# + id="rBVci8HPUTxw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 153} outputId="cd8cc471-7003-44fd-b1dd-6489652e164e"
print('\n# Evaluate on dev data')
results_dev =Senet_model_AUX.evaluate(X_res_dev,Y_dev)
print('dev loss, dev acc:', results_dev)
print('\n# Evaluate on test data')
results_test = Senet_model_AUX.evaluate(X_res_test,Y_test)
print('test loss, test acc:', results_test)
# + [markdown] id="gqQwjwg2XE28" colab_type="text"
# ## Model 2: ACC 73.2%
# + id="rVUlnnmQWNBv" colab_type="code" colab={}
Resnet_model = tf.keras.models.load_model('/content/drive/My Drive/Personal projects/emotion_recognition_paper/models/ResNet-BEST-73.2.h5')
# + id="Sgrj1HyyWP2p" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 153} outputId="610da938-df10-43a8-8c49-a031b2bb3723"
print('\n# Evaluate on dev data')
results_dev = Resnet_model.evaluate(X_res_dev,Y_dev)
print('dev loss, dev acc:', results_dev)
print('\n# Evaluate on test data')
results_test = Resnet_model.evaluate(X_res_test,Y_test)
print('test loss, test acc:', results_test)
# + [markdown] id="YRe12o9dXMBE" colab_type="text"
# ## Model 3: ACC 72.7%
# + id="HG7L3WL5XX7A" colab_type="code" colab={}
Resnet_model_AUX = tf.keras.models.load_model('/content/drive/My Drive/Personal projects/emotion_recognition_paper/models/RESNET50-AUX-BEST-72.7.h5')
# + id="TwAlE4uHXpxB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 153} outputId="f0e0f08f-14a8-4c84-988d-dfb0551fde5e"
print('\n# Evaluate on dev data')
results_dev = Resnet_model_AUX.evaluate(X_res_dev,Y_dev)
print('dev loss, dev acc:', results_dev)
print('\n# Evaluate on test data')
results_test = Resnet_model_AUX.evaluate(X_res_test,Y_test)
print('test loss, test acc:', results_test)
# + id="Hgy-dRl_XtAf" colab_type="code" colab={}
# make an ensemble prediction for multi-class classification
def ensemble_predictions( testX, models_TL, testresX):
# make predictions
yhats = np.zeros((len(models_TL),testX.shape[0],7))
for model_ind in range(len(models_TL)):
yhat = models_TL[model_ind].predict(testresX)
yhats[model_ind,:,:] = yhat
summed = np.sum(yhats, axis=0)
result = np.argmax(summed, axis=1)
return result
# evaluate a specific number of members in an ensemble
def evaluate_n_members(testX, models_TL, testresX, testy):
# select a subset of members
#subset = members[:n_members]
#print(len(subset))
# make prediction
yhat = ensemble_predictions(testX, models_TL, testresX)
# calculate accuracy
return accuracy_score(testy, yhat)
# + id="Ze0e6Ld9YIuT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="bb48ea7f-1479-45e3-9b87-35f7593053d4"
models_TL = [Resnet_model, Resnet_model_AUX, Senet_model_AUX]
ens_acc = evaluate_n_members(X_test, models_TL, X_res_test, np.argmax(Y_test, axis=1))
print(ens_acc)
# + id="l57e-KOwaFQ5" colab_type="code" colab={}
| colab_notebooks/three_model_-ensemble2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Resources for learnings the matplotlib OOP API.
# pyplot and OOP API: https://matplotlib.org/3.1.1/api/index.html
# matplotlib.figure.Figure module: https://matplotlib.org/3.1.1/api/_as_gen/matplotlib.figure.Figure.html#matplotlib.figure.Figure
# matplotlib.axes.Axes module: https://matplotlib.org/3.1.1/api/axes_api.html#matplotlib.axes.Axes
# other matplotlib modules: https://matplotlib.org/3.1.1/api/index.html#modules
import numpy as np
import pandas
import tool_belt # Connects to PostgreSQL (PostGIS) database
import matplotlib.pyplot as plt # State-based plotting.
from matplotlib.figure import Figure # The top level container for all the plot elements.
from matplotlib.axes import Axes # The Axes contains figure elements: Axis, Tick, Line2D, Text, Polygon, etc.
| notebooks/learning matplotlib OOP API.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # [Advent of Code 2020: Day 1](https://adventofcode.com/2020/day/1)
# ## \-\-\- Day 1: Report Repair \-\-\-
#
# After saving Christmas [five years in a row](https://adventofcode.com/events), you've decided to take a vacation at a nice resort on a tropical island. Surely, Christmas will go on without you.
#
# The tropical island has its own currency and is entirely cash\-only. The gold coins used there have a little picture of a starfish; the locals just call them **stars**. None of the currency exchanges seem to have heard of them, but somehow, you'll need to find fifty of these coins by the time you arrive so you can pay the deposit on your room.
#
# To save your vacation, you need to get all **fifty stars** by December 25th.
#
# Collect stars by solving puzzles. Two puzzles will be made available on each day in the Advent calendar; the second puzzle is unlocked when you complete the first. Each puzzle grants **one star**. Good luck!
#
# Before you leave, the Elves in accounting just need you to fix your **expense report** (your puzzle input); apparently, something isn't quite adding up.
#
# Specifically, they need you to **find the two entries that sum to `2020`** and then multiply those two numbers together.
#
# For example, suppose your expense report contained the following:
#
# ```
# 1721
# 979
# 366
# 299
# 675
# 1456
#
# ```
#
# In this list, the two entries that sum to `2020` are `1721` and `299`. Multiplying them together produces `1721 * 299 = 514579`, so the correct answer is **`514579`**.
#
# Of course, your expense report is much larger. **Find the two entries that sum to `2020`; what do you get if you multiply them together?**
# +
import unittest
from IPython.display import Markdown, display
from aoc_puzzle import AocPuzzle
class Report(AocPuzzle):
def parse_data(self, data):
self.data = list(map(int, data.split('\n')))
def run(self, output=False):
for index1, entry1 in enumerate(self.data):
for index2, entry2 in enumerate(self.data):
if index1 != index2:
if (entry1 + entry2) == 2020:
product = entry1 * entry2
if output:
display(Markdown(f'### Entries `{entry1}` and `{entry2}` sum to `2020` and their product is **`{product}`**'))
return product
class TestBasic(unittest.TestCase):
def test_report(self):
in_data = '1721\n979\n366\n299\n675\n1456'
exp_out = 514579
report = Report(in_data)
self.assertEqual(report.run(), exp_out)
unittest.main(argv=[""], exit=False)
# -
report = Report("input/d01.txt")
report.run(output=True)
# ## \-\-\- Part Two \-\-\-
#
# The Elves in accounting are thankful for your help; one of them even offers you a starfish coin they had left over from a past vacation. They offer you a second one if you can find **three** numbers in your expense report that meet the same criteria.
#
# Using the above example again, the three entries that sum to `2020` are `979`, `366`, and `675`. Multiplying them together produces the answer, **`241861950`**.
#
# In your expense report, **what is the product of the three entries that sum to `2020`?**
# +
class Report2(Report):
def run(self, output=False):
for i1, e1 in enumerate(self.data):
for i2, e2 in enumerate(self.data):
for i3, e3 in enumerate(self.data):
if i1 != i2 or i1 != i3:
if (e1 + e2 + e3) == 2020:
product = e1 * e2 * e3
if output:
display(Markdown(f'### Entries `{e1}`, `{e2}`, and `{e3}` sum to `2020` and their product is **`{product}`**'))
return product
class TestBasic(unittest.TestCase):
def test_report(self):
in_data = '2020\n979\n366\n299\n675\n1456'
exp_out = 241861950
report = Report2(in_data)
self.assertEqual(report.run(), exp_out)
unittest.main(argv=[""], exit=False)
# -
report = Report2("input/d01.txt")
report.run(output=True)
| AoC 2020/AoC 2020 - Day 01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from oda_api.api import DispatcherAPI
from oda_api.plot_tools import OdaImage,OdaLightCurve
from oda_api.data_products import BinaryData
import os
from astropy.io import fits
import numpy as np
from numpy import sqrt,exp
import matplotlib.pyplot as plt
# %matplotlib inline
# + tags=["parameters"]
source_name='3C 279'
ra=194.046527
dec=-5.789314
radius=15.
Tstart='2015-06-01T00:00:00'
Tstop='2015-06-30T00:00:00'
host='www.astro.unige.ch/cdci/astrooda/dispatch-data'
time_bin=10000
Nbins=7 # number of time bin in the long-term lightcurve
E1_keV=20.
E2_keV=100.
# -
try: input = raw_input
except NameError: pass
token=input() # token for restricted access server
cookies=dict(_oauth2_proxy=token)
disp=DispatcherAPI(host=host)
disp=DispatcherAPI(host=host)
import requests
url="https://www.astro.unige.ch/cdci/astrooda/dispatch-data/gw/timesystem/api/v1.0/scwlist/cons/"
def queryxtime(**args):
params=Tstart+'/'+Tstop+'?&ra='+str(ra)+'&dec='+str(dec)+'&radius='+str(radius)+'&min_good_isgri=100'
print(url+params)
return requests.get(url+params,cookies=cookies).json()
scwlist=queryxtime()
m=len(scwlist)
pointings_osa10=[]
pointings_osa11=[]
for i in range(m):
if scwlist[i][-2:]=='10':
if(int(scwlist[i][:4])<1626):
pointings_osa10.append(scwlist[i]+'.001')
else:
pointings_osa11.append(scwlist[i]+'.001')
#else:
# pointings=np.genfromtxt('scws_3C279_isgri_10deg.txt', dtype='str')
m_osa10=len(pointings_osa10)
m_osa11=len(pointings_osa11)
scw_lists_osa10=[]
scw_lists_osa11=[]
count=0
scw_string=''
for i in range(m_osa10):
if count<50:
scw_string=scw_string+str(pointings_osa10[i])+','
count+=1
else:
scw_lists_osa10.append(scw_string[:-1])
count=0
scw_string=str(pointings_osa10[i])+','
scw_lists_osa10.append(scw_string[:-1])
print(len(scw_lists_osa10))
count=0
scw_string=''
for i in range(m_osa11):
if count<50:
scw_string=scw_string+str(pointings_osa11[i])+','
count+=1
else:
scw_lists_osa11.append(scw_string[:-1])
count=0
scw_string=str(pointings_osa11[i])+','
scw_lists_osa11.append(scw_string[:-1])
print(len(scw_lists_osa11))
data=disp.get_product(instrument='isgri',
product='isgri_image',
scw_list=scw_lists_osa10[0],
E1_keV=E1_keV,
E2_keV=E2_keV,
osa_version='OSA10.2',
RA=ra,
DEC=dec,
detection_threshold=3,
product_type='Real')
data.dispatcher_catalog_1.table
FLAG=0
torm=[]
for ID,n in enumerate(data.dispatcher_catalog_1.table['src_names']):
if(n[0:3]=='NEW'):
torm.append(ID)
if(n==source_name):
FLAG=1
data.dispatcher_catalog_1.table.remove_rows(torm)
nrows=len(data.dispatcher_catalog_1.table['src_names'])
# +
if FLAG==0:
data.dispatcher_catalog_1.table.add_row((0,'3C 279',0,ra,dec,0,2,0,0))
data.dispatcher_catalog_1.table
# -
api_cat=data.dispatcher_catalog_1.get_api_dictionary()
# +
lc_results=[]
for i in range(len(scw_lists_osa10)):
print(i)
data=disp.get_product(instrument='isgri',
product='isgri_lc',
scw_list=scw_lists_osa10[i],
E1_keV=E1_keV,
E2_keV=E2_keV,
osa_version='OSA10.2',
RA=ra,
DEC=dec,
time_bin=time_bin,
selected_catalog=api_cat)
lc_results.append(data)
# +
t=[]
r=[]
err=[]
tot_counts=[]
backv=[]
backe=[]
i=0
for lc in lc_results:
for ID,s in enumerate(lc._p_list):
if s.meta_data['src_name']==source_name:
i=i+1
for tt in s.data_unit[1].data['TIME']:
t.append(tt)
for rr in s.data_unit[1].data['RATE']:
r.append(rr)
for ee in s.data_unit[1].data['ERROR']:
err.append(ee)
for tc in s.data_unit[1].data['TOT_COUNTS']:
tot_counts.append(tc)
for bv in s.data_unit[1].data['BACKV']:
backv.append(bv)
for be in s.data_unit[1].data['BACKE']:
backe.append(be)
t=np.array(t)
r=np.array(r)
err=np.array(err)
tot_counts=np.array(tot_counts)
backv=np.array(backv)
backe=np.array(backe)
# +
fig = plt.figure(figsize=(10,7))
Nbins=7
t0=51544
tmin=t0+t[0]
tmax=t0+t[-1]
time=np.linspace(tmin,tmax,Nbins)
dtime=time[1]-time[0]
time_av=time-dtime/2.
flux=np.zeros(Nbins)
error=np.zeros(Nbins)
j=0
sc=1.
k=0
while (j<len(time)):
while ((t[k]+t0<time[j])):
if (err[k]>0.):
flux[j]=flux[j]+r[k]/(err[k])**2
error[j]=error[j]+1./(err[k])**2
k=k+1
if (k==len(t)):
break
if (k==len(t)):
break
else:
j=j+1
plt.errorbar(t+t0-57188,r*exp((t+t0-52900)/40000.),yerr=err*exp((t+t0-52900)/40000.),linestyle='none',color='black',alpha=0.03,linewidth=2)
for i in range(len(flux)):
if(error[i]>0.):
flux[i]=flux[i]/(error[i])*exp((time_av[i]-52900)/40000.)
error[i]=1./sqrt(error[i])*exp((time_av[i]-52900)/40000.)
plt.errorbar(time_av-57188,flux,yerr=error,xerr=dtime/2.,linestyle='none',color='red',alpha=0.7,linewidth=4)
#plt.plot(time_av,exp((time_av-52900)/40000.)/2)
plt.tick_params(axis='both', which='major', labelsize=16)
ymax=1.1*max(flux+error)
plt.ylim(-0.1,ymax)
plt.xlabel('Time, MJD-57188',fontsize=16)
plt.ylabel('Rate, cts/s',fontsize=16)
#plt.text(tmin,0.9*ymax, str(E1_keV)+'-'+str(E2_keV)+' keV',fontsize=16)
#plt.text(tmin,0.8*ymax, source_name,fontsize=16)
name=source_name.replace(" ", "")
plt.savefig(name+'_lc_flare.pdf',format='pdf',dpi=100)
# + tags=["outputs"]
lc_3C279=name+'_lc_flare.pdf'
# -
| examples/3C279_lc_flare.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="95SYNAMCZYsB"
# ***Copyright 2020 Google LLC.***
#
# Licensed under the Apache License, Version 2.0 (the "License");
# + [markdown] id="aqZAWIB9Zc4a"
# Author: <NAME>
# Term: Summer 2020 Research Internship with Mixel/Brain
# Purpose: This notebook analyzes user history and time resolution within such histories.
# + id="Pirvljq39Gjw"
import tensorflow.compat.v1 as tf
from collections import defaultdict
from datetime import datetime
import copy
import random
import json
import string
import os
import numpy as np
import itertools
import matplotlib. pyplot as plt
# + id="oCxCrNvS-eKf"
""" Read data. For each label, we store a dictionary of users where
key:user_id and value: list of <item id, timestamp> tuples."""
base_dir = 'path/to/your/processed/data'
dataset_categories = ['Kindle_Store', 'CDs_and_Vinyl', 'Grocery_and_Gourmet_Food', 'Movies_and_TV', 'Video_Games', 'Pet_Supplies']
labels=['Kindle', 'CDs', 'Food', 'Movies', 'Games', 'Pets']
data = {label: [] for label in labels}
user_lists = {label: defaultdict(list) for label in labels}
for dataset_category, label in zip(dataset_categories, labels):
dataset_path = '{}{}_user_item_query_time_mapped.txt'.format(base_dir, dataset_category)
# Read user,item,query,time
with tf.gfile.Open(dataset_path, "r") as f:
for line in f:
u, i, _, t = [int(x) for x in line.rstrip().split(" ")] # Ignores query.
user_lists[label][u].append([i, t])
print("Category {} is done. Number of users = {}".format(dataset_category, len(user_lists[label])))
# + id="iUip6qyYB-qX"
""" Helper functions for timestamp."""
def convert_to_date(timestamp):
date_time = datetime.fromtimestamp(timestamp)
return date_time.strftime("%m/%d/%Y") # , %H:%M:%S
def time_delta(ts1, ts2):
if ts1 > ts2:
return (datetime.fromtimestamp(ts1) - datetime.fromtimestamp(ts2)).days
else:
return (datetime.fromtimestamp(ts2) - datetime.fromtimestamp(ts1)).days
# + id="flBVOWmn_i4X"
""" Sampling random users from Kindle category for visual inspection."""
for idx in random.sample(range(1, 1000), 10):
print([(i, convert_to_date(t), t) for i, t in user_lists['Kindle'][idx]])
# + id="u_ZGvaJHYf3s"
""" Plotting temporal resolution of the user history. """
sampled_users = []
num_users_sample = 100
user_len_sample = 20
while len(sampled_users) < num_users_sample:
idx = random.sample(range(1, len(user_lists['Kindle'])), 1)[0]
if len(user_lists['Kindle'][idx]) != user_len_sample:
continue
sampled_users.append(user_lists['Kindle'][idx])
num_users_plot = 5
plot_users = random.sample(sampled_users, num_users_plot)
bins = [int(i) for i in range(1, user_len_sample)]
idx = 0
plt.figure(figsize=(10, 8))
plt.xlabel('Time period (days)', fontsize=20)
plt.xticks(fontsize=20)
plt.ylabel('User interactions', fontsize=20)
plt.yticks(bins, fontsize=20)
for u in plot_users:
last_t = plot_users[-1][-1][1]
time_deltas = list(sorted([time_delta(last_t, t) for i, t in u[:-1]]))
plt.scatter(time_deltas, bins, label = 'user {}'.format(idx+1))
idx += 1
plt.legend()
# + id="1rq3YghkmVUj"
""" Compute the average time spent between two consecutive items. """
days_list = {label: [] for label in labels}
for label in labels:
for user, items in user_lists[label].items():
nxt_item = items[-1]
for item in reversed(items[:-1]):
time_d = time_delta(nxt_item[1], item[1])
days_list[label].append(time_d)
nxt_item = item
print("Mean time spent between two consecutive items {} for {} (in days)".format(np.mean(days_list[label]), label))
print("Median time spent between two consecutive items {} for {} (in days)".format(np.median(days_list[label]), label))
# + id="QyKzmR41uOui"
""" Compute the cumulative ratio of items vs time window (in days) with respect
to last item per user. """
max_day = 365*2 # Consider up to 2 years
days_dict = {label: defaultdict(lambda: []) for label in labels}
for label in labels:
for user, items in user_lists[label].items():
last_item = items[-1]
user_time_dict = {i:0 for i in range(max_day)}
scale = len(items) - 1
for item in reversed(items[:-1]):
time_d = time_delta(last_item[1], item[1])
if time_d not in user_time_dict:
break
user_time_dict[time_d] += 1/scale
sum_so_far = 0
for key in sorted(user_time_dict.keys()):
user_time_dict[key] += sum_so_far
sum_so_far = user_time_dict[key]
for key, value in user_time_dict.items():
days_dict[label][key].append(value)
# + id="-hzizWCIY1aO"
# Sanity check
len(days_dict['Pets'][0]) == len(days_dict['Pets'][100]) == len(days_dict['Pets'][500])
# + id="abDLSiWAakKB"
""" Compute the (non-overlapping) segment densities with respect to different
segment allocations.
"""
def _segment_boundary_exp(b, i):
""" Computes the end of segment i wrt to exponential with base b. The formula simply is:
t = b^i
"""
return b**i
def _segment_boundary_pow(b, i):
""" Computes the end of segment i wrt to power law with base b. The formula simply is:
t = i**b
"""
return i**b
def _segment_boundary_lin(b, i):
""" Computes the end of segment i wrt to linear with base b. The formula simply is:
t = b*i
"""
return b*i
# + id="g00kVpEzSieJ"
bases_exp = [3, 4, 5, 8]
num_segments = 10
max_day = 365*2 # Consider up to 2 years.
segment_analysis_dict_exp = {label:{} for label in labels}
for label in labels:
for base in bases_exp:
segment_boundaries = [_segment_boundary_exp(base, i) for i in range(num_segments)]
segment_densities = [0] * num_segments
for user, items in user_lists[label].items():
last_item = items[-1]
scale = len(items) - 1
active_segment_idx = 0
for item in reversed(items[:-1]):
time_d = time_delta(last_item[1], item[1])
if time_d > max_day:
break
while time_d > segment_boundaries[active_segment_idx] and active_segment_idx < (num_segments - 1):
active_segment_idx += 1
segment_densities[active_segment_idx] += 1/scale
segment_analysis_dict_exp[label][base] = np.array(segment_densities) /len(user_lists[label])
# + id="QD8blTEv_m1a"
| multi_resolution_rec/notebooks/user_history_time_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# 「Yahoo!ニュース」が配信するRSSからヘッドラインを抜き出す
import requests
from bs4 import BeautifulSoup # ①
xml = requests.get('http://news.yahoo.co.jp/pickup/science/rss.xml') # ②
soup = BeautifulSoup(xml.text, 'html.parser') # ③
for news in soup.findAll('item'): # ④
print(news.title.string) # ⑤
# -
| sample/Python_GOKUI/Python_GOKUI/chap08/sec03/WebScraping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Update Vaccination Data in ArcGIS Online from GitHub
# This script will help you automatically update the data within your ArcGIS Online account that we uploaded from the [Publishing Vaccination Data from GitHub to ArcGIS](Get to Publish World Vaccination Data.ipynb) tutorial.
#
# The only thing you need to change **if you followed** the Publishing tutorial is the `username` and the `password` for your account below.
#
# ### Credits
#
# This script was created with the help and documentation from the [ArcGIS API for Python](https://developers.arcgis.com/python/guide/accessing-and-creating-content/) within the [ArcGIS Developers](https://developers.arcgis.com/) platform.
# Import libraries
from arcgis.gis import GIS
from arcgis import features
import pandas as pd
# Connect to the GIS
gis = GIS(url='https://arcgis.com', username='surveyor_jr', password='')
# Search for the Vaccination Data
vaccination_data = gis.content.search(query="World COVID-19 Vaccination", item_type = "Feature Layer")
display(vaccination_data)
# +
# Assign variables to each independent layer
# --> For the fixed table
fixed_table = vaccination_data[1]
# Just to verify if True
display(fixed_table)
# -
# --> For the time series
time_series_table = vaccination_data[0]
# Verification
display(time_series_table)
# +
# URLs for updates
# for fixed data
fixed_updates = 'https://raw.githubusercontent.com/govex/COVID-19/master/data_tables/vaccine_data/global_data/vaccine_data_global.csv'
# for time_series data
time_series_updates = 'https://raw.githubusercontent.com/govex/COVID-19/master/data_tables/vaccine_data/global_data/time_series_covid19_vaccine_global.csv'
# -
# ### Fixed Table Data Overwrite
# Retrieve new data after sometime
# create another dataFrame
vaccine_fixed = pd.read_csv(fixed_updates)
# Just confirm to see if it works
vaccine_fixed.head(10)
# Write the DataFrame to file
import os
if not os.path.exists(os.path.join('data', 'updated_vaccination_data')):
os.mkdir(os.path.join('data','updated_vaccination_data'))
# Write the file into to new folder
vaccine_fixed.to_csv(os.path.join('data', 'updated_vaccination_data', 'vaccine_data_global.csv'))
# +
# Overwriting the Layer
from arcgis.features import FeatureLayerCollection
fixed_vaccination_data = FeatureLayerCollection.fromitem(fixed_table) # the fixed table-layer
#Finally overwriting the Layer
fixed_vaccination_data.manager.overwrite(os.path.join('data', 'updated_vaccination_data', 'vaccine_data_global.csv'))
# returns True if successful
# -
# ### Time Series Data Overwrite
# Create new data frame for the time-series data
vaccine_time_series = pd.read_csv(time_series_updates)
# Just confirm to see if it works
vaccine_time_series.head(10)
# Write the file into to the folder
vaccine_time_series.to_csv(os.path.join('data', 'updated_vaccination_data', 'time_series_covid19_vaccine_global.csv'))
# +
# Overwriting the Layer
from arcgis.features import FeatureLayerCollection
time_series_vaccination_data = FeatureLayerCollection.fromitem(time_series_table) # the fixed table-layer
#Finally overwriting the Layer
time_series_vaccination_data.manager.overwrite(os.path.join('data', 'updated_vaccination_data', 'time_series_covid19_vaccine_global.csv'))
# returns True if successful
# -
| automation_scripts/Overwrite and Update Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="_8SX62bL8dyp"
import pandas as pd
import numpy as np
# + [markdown] id="f9uHhwqTktd3"
# #Data Upload and Viewing
# + id="avPZ970bO1hK"
# load airbnb data
gz = pd.read_csv('http://data.insideairbnb.com/united-states/ca/los-angeles/2021-04-07/data/listings.csv.gz')
# + id="UJaTMEeiPApf"
print(gz.shape)
gz.head()
# + id="yASVdaQ-hPYX"
# remove unnecessary columns
cols = gz.columns
gz.columns
# + id="NdqBJzquhqQj"
# create a list of the columns to save
save = [5,6,27,29,30,35,36, 40,41]
cols_to_save = []
for i in save:
cols_to_save.append(cols[i])
print(type(cols_to_save[0]))
cols_to_save
# + id="23XoXtXIq9P6"
# create a new dataframe with only the needed columns
df = gz[cols_to_save].copy()
df.head()
# + id="A2mzvJb_f2aB"
# check data types for all columns
for i in df.columns:
des = type(df[i][1])
print(i)
print(des)
# + [markdown] id="pqWezrQdvZHv"
# # Data Pre-Processing
# + id="xxzPOfiHsz8b"
df.isnull().sum()
# + id="XMvg3lITvJsn"
# fill null values
df['description'] = df['description'].fillna("None")
df['neighborhood_overview'] = df['neighborhood_overview'].fillna("None")
df['bathrooms_text'] = df['bathrooms_text'].fillna("1")
df['bedrooms'] = df['bedrooms'].fillna(1)
# + id="B_P0C4W0vC3o"
# see if I missed any
df.isnull().sum()
# + id="759ty4-aXH_C"
import re
def fix_bathroom(text):
"""
Removes anything that is not a number
Converts and returns the remaining number from a str to float
"""
text = text.lower()
if text[-9:] == 'half-bath':
baths = float('0.5')
else:
baths = float(re.sub(r'[^0-9\.]', '', text))
return baths
def to_float(num):
return(float(num))
def fix_price(price_str):
"""
convert price column from string to int
"""
price = re.sub(r'[^0-9]', '', price_str)
price = price[:-2]
return int(price)
# + id="lcN0fkWMXUWC"
# adjust columns to appropriate data format
df['bathrooms'] = df['bathrooms_text'].apply(fix_bathroom)
df['minimum_nights'] = df['minimum_nights'].apply(to_float)
df['target'] = gz['price'].apply(fix_price)
# + id="lm48O7P1T30N"
# remove outliers
df = df[(df['target'] >= 25) & (df['target'] <= 2000) &
(df['bathrooms'] >=1) & (df['bathrooms'] < 4) &
(df['bedrooms'] >= 1) & (df['bedrooms'] <= 4) &
(df['minimum_nights']>=1) & (df['minimum_nights']<=30)]
df.shape
# + [markdown] id="r_pkPQFvcktS"
# #Model
# + id="e_h2bxI3z2Ix"
df.columns
# + id="dLfNssuxzu2V"
df.drop(columns = ['description', 'neighborhood_overview', 'bathrooms_text'], inplace=True)
# + id="sTZktRPJnBz8"
df['minimum_nights'].describe()
# + id="j0kbRo2Y_frg"
df.columns
# + id="aq1WgwUtXkGl"
X = df.drop(columns=['target','latitude','longitude', 'neighbourhood_cleansed'])
y = df['target']
# + id="afjOWhog0e75"
# !pip install category_encoders==2.*
# + id="l0TCVVDkzivK"
# one hot encode
import category_encoders as ce
encoder = ce.OneHotEncoder(use_cat_names = True)
X = encoder.fit_transform(X)
# + id="SB7mKAJTUmFu"
X.columns
# + id="R6yckqJZVlHW"
# # # define base model
# def baseline_model():
# # create model
# model = Sequential()
# model.add(Dense(5, input_dim=3, kernel_initializer='normal', activation='relu'))
# model.add(Dense(6, kernel_initializer='normal', activation='relu'))
# model.add(Dense(1, kernel_initializer='normal'))
# # Compile model
# model.compile(loss='mean_squared_error', optimizer='adam')
# return model
# # evaluate model with standardized dataset
# estimators = []
# estimators.append(('standardize', StandardScaler()))
# estimators.append(('mlp', KerasRegressor(build_fn=baseline_model, epochs=10, batch_size=5, verbose=0)))
# pipeline = Pipeline(estimators)
# kfold = KFold(n_splits=5)
# results = cross_val_score(pipeline, X, y, cv=kfold)
# print("Standardized: %.2f (%.2f) MSE" % (results.mean(), results.std()))
# + id="nTMqbzYA1lI-"
# + id="7cvyYjDSn2dl"
from keras.models import Sequential
from keras.layers import Dense
from keras.wrappers.scikit_learn import KerasRegressor
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import KFold
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import Pipeline
def create_model():
# create model
model = Sequential()
model.add(Dense(5, input_dim=3, kernel_initializer='normal', activation='relu'))
model.add(Dense(6, kernel_initializer='normal', activation='relu'))
model.add(Dense(1, kernel_initializer='normal'))
# Compile model
model.compile(loss='mean_squared_error', optimizer='adam')
return model
# # evaluate model with standardized dataset
# estimators = []
# estimators.append(('standardize', StandardScaler()))
# estimators.append(('mlp', KerasRegressor(build_fn=baseline_model, epochs=50, batch_size=5, verbose=0)))
# pipeline = Pipeline(estimators)
# kfold = KFold(n_splits=5)
# results = cross_val_score(pipeline, X, y, cv=kfold)
# print("Standardized: %.2f (%.2f) MSE" % (results.mean(), results.std()))
# + id="7fAUY6UL1_me"
model = create_model()
# + id="b1LjN7US5b31"
from keras.models import Sequential
from keras.layers import Dense
# from keras.wrappers.scikit_learn import KerasRegressor
# from sklearn.model_selection import cross_val_score
# from sklearn.model_selection import KFold
# from sklearn.preprocessing import StandardScaler
# from sklearn.pipeline import Pipeline
model = Sequential()
model.add(Dense(5, input_dim=4, kernel_initializer='normal', activation='relu'))
model.add(Dense(6, kernel_initializer='normal', activation='relu'))
model.add(Dense(1, kernel_initializer='normal'))
# Compile model
model.compile(loss='mean_squared_error', optimizer='adam')
# + id="WI20GdO8KK4F"
# + id="u8NRHQ836Rq1"
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.25, random_state=42)
# + id="ao804WVR6lc_"
type(y)
# + id="buhmB2ne2LkO"
model.fit(X, y,
epochs = 122,
batch_size=16,
validation_data=(X_test, y_test)
# metric='accuracy'
)
# + id="1ellBB-P4xHX"
y[0:5]
# + id="nAG1vsGAvl_H"
# + id="Sd-c5ZdR7N1z"
X.head(5)
# + id="sl9x3DkKMype"
model.predict([[2,30,730,2]])
# + id="4wkw1Vfg4m1D"
model.predict([[4, 30, 1, 2]])
# + id="Om82IUGVs3Kr"
# zip the model
# !zip -r ./nn.zip ./nn_model/
# + id="PreHfeD8sOeo"
# download to local machine
from google.colab import files
files.download("./nn.zip")
# + id="Qtm3ynG8uEmt"
# test uploading saved model
from tensorflow import keras
model = keras.models.load_model('./nn_model')
prediction = model.predict([[2,30,730,2]])
prediction[0][0].round()
# + [markdown] id="RRO4PSpTxwcQ"
# #Natural Language Processing
# + id="1UHYtBxJxzQc"
# Start NLP
from collections import Counter
# Plotting
import matplotlib.pyplot as plt
import seaborn as sns
# NLP Libraries
import re
from nltk.stem import PorterStemmer
import spacy
from spacy.tokenizer import Tokenizer
# + id="Wb8VTqa4x1LO"
# !python -m spacy download en_core_web_lg
# + id="wG8Oi6Gzx9UU"
# Initialize spacy model & tokenizer
nlp = spacy.load('en_core_web_lg')
tokenizer = Tokenizer(nlp.vocab)
# + id="Fn21F_uiyUg0"
# Create tokenize function
def tokenize(text):
tokens = re.sub(r'[^a-zA-Z ^0-9]', ',', text)
tokens = tokens.lower().replace(',', ' ')
tokens = tokens.split()
return tokens
def fix_bathroom(text):
"""
Removes anything that is not a number
Converts and returns the remaining number from a str to float
"""
text = text.lower()
if text[-9:] == 'half-bath':
baths = float('0.5')
else:
baths = float(re.sub(r'[^0-9\.]', '', text))
return baths
def fix_price(price_str):
"""
convert price column from string to int
"""
price = re.sub(r'[^0-9]', '', price_str)
price = price[:-2]
return int(price)
# + id="C3A71F2h96RH"
df.columns
# + id="YEWi25dMaWaV"
# convert str prices to int prices
gz['target'] = gz['price'].apply(fix_price)
# + id="YVhw6W4PFF3i"
token_cols
# + id="pAaAmxm2FNfh"
type(df['bathrooms_text'][0])
# + id="eFZaajTXF8-u"
df['bathrooms']
# + id="9MEsjoMryY4c"
# tokenize dataframe
token_cols = df.columns
# remove numerical columns
token_cols = token_cols.drop(['latitude','bedrooms','longitude','bathrooms', 'minimum_nights'])
token_cols
for i in token_cols:
if i == 'bathrooms_text':
# df['bathrooms'] = df[i].apply(fix_bathroom)
g=1+1
else:
df[i+"_token"] = df[i].apply(tokenize)
# + id="khiUonNHxf_A"
df[['bedrooms', 'bathrooms', 'latitude', 'longitude']].describe()
# + id="WA3fw_N4xf_C"
# remove outliers
df['target'] = gz['target']
# df = df[(df['target'] >= 25) & (df['target'] <= 2000)
# & (df['bathrooms'] <= 4) & (df['bathrooms'] >= 1)]
# # (df['bedrooms'] <= 4) & (df['bedrooms'] >= 1)]
df = df[(df['target'] >= 25) & (df['target'] <= 2000) &
(df['bathrooms'] >=1) & (df['bathrooms'] < 4) &
(df['bedrooms'] >= 1) & (df['bedrooms'] <= 4)]
# + id="yTSgwe6eIMZ6"
df.head()
# + id="cuhz8fbbJtin"
# Counter Function - takes a corpus of document and returns dataframe of word counts
from collections import Counter
word_counts = Counter()
def count(docs):
word_counts = Counter()
appears_in = Counter()
total_docs = len(docs)
for doc in docs:
word_counts.update(doc)
appears_in.update(set(doc))
temp = zip(word_counts.keys(), word_counts.values())
wc = pd.DataFrame(list(temp), columns = ['word', 'count'])
wc['rank'] = wc['count'].rank(method='first', ascending=False)
total = wc['count'].sum()
wc['pct_total'] = wc['count'].apply(lambda x: x / total)
wc = wc.sort_values(by='rank')
wc['cul_pct_total'] = wc['pct_total'].cumsum()
t2 = zip(appears_in.keys(), appears_in.values())
ac = pd.DataFrame(t2, columns=['word', 'appears_in'])
wc = ac.merge(wc, on='word')
wc['appears_in_pct'] = wc['appears_in'].apply(lambda x: x / total_docs)
return wc
# + id="FoxmTjpGJzLP"
wc_neighborhood = count(df['neighbourhood_cleansed_token'])
wc_neighborhood.sort_values(by='rank')
# + id="mBf7J8-tJ6Ls"
wc_neighborhood.shape
# + id="470lFlCQKFCO"
#Cumulative Distribution Plot
sns.lineplot(x='rank', y='cul_pct_total', data=wc_neighborhood);
# + id="usoayWrGKHCv"
# inspect some descriptions
df['description'].iloc[0]
# + id="km_FyRiuzYd9"
df['description'].iloc[1]
# + id="PnFU9H096bDr"
df['description'][0]
# + id="RPpfsHq4x4_9"
def clean(text):
text = (text
.str.replace('<br /><br />',' ')
.str.replace('<b>',' ')
.str.replace('</b><br />',' ')
# .str.replace('</b><br />',' ')
.str.replace('*','')
)
return text
def clean_description(text):
cleaned = re.sub(r'[^a-zA-Z]', ',', text)
cleaned = cleaned.lower().replace(',', ' ')
return cleaned
# + id="QJSJgOR92uyy"
df['cleaned_desc'] = df['description'].apply(clean_description)
# + id="Z5vX3TUWy4wx"
# inspect cleaned descriptions and look for stop words
df['cleaned_desc'].iloc[0]
# + id="gnyn6px60CGH"
# Vectorization
from sklearn.feature_extraction.text import TfidfVectorizer
# Instantiate Vectorizer
tfidf = TfidfVectorizer(stop_words='english',
max_features=5000)
# Create a vocabulary and tf-idf score per description
dtm = tfidf.fit_transform(df['cleaned_desc'])
# Get feature names to use as dataframe column headers
general_dtm = pd.DataFrame(dtm.todense(), columns=tfidf.get_feature_names())
general_dtm.head()
# + id="rHBuhiNiwzYP"
general_dtm.shape
# + id="uBtwPzJIDxKj"
df.shape
# + [markdown] id="E23AUiAZoFDJ"
# #Seasons
# + id="OaJZDYWv1fZp"
cal = pd.read_csv('http://data.insideairbnb.com/united-states/ca/los-angeles/2021-04-07/data/calendar.csv.gz')
# + id="ngv8DsT83Dpz"
print(cal.shape)
cal.head()
# + id="D2LzmnDo8q7G"
cal[cal['listing_id']==35922]
# + id="Qf4nO0duWFs9"
def date_to_season(date):
"""
Takes the string of date and returns what season it is in.
"""
season = ['Winter', 'Spring', 'Summer', 'Fall']
date_num = int(date[5:7])
# print(date_num)
if date_num <=2 or date_num ==12:
return season[0]
elif date_num<=5:
return season[1]
elif date_num<=8:
return season[2]
else:
return season[3]
return season[i]
# + id="n6wg4vcwX1Lk"
date_to_season(cal['date'][0])
# + id="2Ia0ganbZsaI"
cal.drop(labels = 'Season', axis = 1)
# + id="dsuODS1jYcK3"
cal['Season'] = cal['date'].apply(date_to_season)
cal.head()
# + id="IsZ1L6VFZqYK"
listed = cal.listing_id.unique()
len(listed)
# + id="dF_Ntnapahdk"
temp = cal[cal['listing_id']==35922]
temp
# + id="M1ufzewsbRQk"
seasons = temp.Season.unique()
season_str = np.array2string(seasons)
type(season_str)
# + id="BoYkqfEBgxXd"
new_season = list()
new_season.append(season_str)
new_season
# + id="_xUrvx4uhBtl"
def define_seasons(data):
"""
takes the dataframe and changes the
"""
| AirBnB_Price.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: dl-workshop
# language: python
# name: dl-workshop
# ---
# # Introduction
# ## Welcome to the course!
#
# Welcome to this course, **Deep learning fundamentals: from gradient descent to neural networks**.
# My name is Eric, and I'll be your instructor for the course.
#
# Deep learning is becoming more and more ubiquitous by the day.
# The hype, however, might stop us from thinking clearly and communicating intelligently
# about the models that we develop and use.
# If you're someone who knows a bit of programming in Python and NumPy,
# then this course will help you gain a firmer grasp of the fundamentals of deep learning.
# Whether you are a skilled executive or a front-line practitioner,
# come and leverage your programming skills
# to gain a fundamental understanding of deep learning
# and level up your knowledge!
# + [markdown] tags=[]
# ## Course Structure
#
# This course is designed to disambiguate one question:
# "What exactly is happening inside a neural network?"
# To answer this question, we will use a combination of code,
# prose,
# equations,
# and videos to show you answers to the questions:
#
# 1. What is gradient-based optimization?
# 2. How do we optimize parameter values to fit data in linear models?
# 3. How do the concepts from linear models transfer over to logistic models?
# 3. How does that same knowledge transfer conceptually to neural network models?
#
# Visually, these are the four notebooks that we have designed to answer those questions.
#
# <img src="../images/00-introduction/notebooks-overview.png">
# -
# ## Learning Objectives
#
# By the end of this course, I expect that you should be able to:
#
# 1. Explain gradient-based optimization in terms of an objective/loss function and parameters we are optimizing,
# 2. Articulate the connection between linear models and feed-forward neural networks, and
# 3. Identify situations where neural networks are useful in lieu of other computational models.
| notebooks/00-introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 教學目標
#
# 主要說明matplotlib 的基礎操作
#
#
#
# # 範例重點
#
# 如何使用亂數, 資料集來操作
#
# +
# 載入需要的...
import matplotlib.pyplot as plt
import numpy as np
# 準備數據 ... 假設我要畫一個sin波 從0~180度
# 設定要畫的的x,y數據list....
x = np.arange(0,180)
y = np.cos(x * np.pi / 180.0)
# 開始畫圖
plt.plot(x,y)
# 在這個指令之前,都還在做畫圖的動作
# 這個指令算是 "秀圖"
plt.show()
#畫出完整的 sin 圖形
x = np.arange(0, 3 * np.pi, 0.1)
y_sin = np.sin(x)
plt.plot(x, y_sin)
plt.show()
#畫出完整的 cos 圖形
x = np.arange(0, 3 * np.pi, 0.1)
y_cos = np.cos(x)
plt.plot(x, y_cos)
plt.show()
# -
# # 散點圖: Scatter Plots
#
# 顏色由(X,Y)的角度給出。
#
# 注意標記的大小,顏色和透明度。
# +
n = 1024
X = np.random.normal(0,1,n)
Y = np.random.normal(0,1,n)
T = np.arctan2(Y,X)
plt.scatter(X,Y, s=75, c=T, alpha=.5)
plt.xlim(-1.5,1.5)
plt.ylim(-1.5,1.5)
plt.show()
# -
| Solution/Day_18_Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Multiple linear regression and adjusted R-squared
# ## Import the relevant libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import statsmodels.api as sm
import seaborn
seaborn.set()
# ## Load the data
# Load the data from a .csv in the same folder
data = pd.read_csv('1.02. Multiple linear regression.csv')
# Let's check what's inside this data frame
data
# This method gives us very nice descriptive statistics.
data.describe()
# ## Create your first multiple regression
# Following the regression equation, our dependent variable (y) is the GPA
y = data ['GPA']
# Similarly, our independent variable (x) is the SAT score
x1 = data [['SAT','Rand 1,2,3']]
# Add a constant. Esentially, we are adding a new column (equal in lenght to x), which consists only of 1s
x = sm.add_constant(x1)
# Fit the model, according to the OLS (ordinary least squares) method with a dependent variable y and an idependent x
results = sm.OLS(y,x).fit()
# Print a nice summary of the regression.
results.summary()
dummy_data = pd.read_csv('1.03. Dummies.csv')
dummy_data
# #### Yes will become 1
# #### No will become 0
dummy_data_copy = dummy_data.copy()
dummy_data_copy['Attendance'] = dummy_data_copy['Attendance'].map({'Yes': 1, 'No': 0})
dummy_data_copy
dummy_data_copy.describe()
y = dummy_data_copy ['GPA']
# Similarly, our independent variable (x) is the SAT score
x1 = dummy_data_copy [['SAT','Attendance']]
# Add a constant. Esentially, we are adding a new column (equal in lenght to x), which consists only of 1s
x = sm.add_constant(x1)
# Fit the model, according to the OLS (ordinary least squares) method with a dependent variable y and an idependent x
results = sm.OLS(y,x).fit()
results.summary()
# +
# Create a scatter plot
plt.scatter(dummy_data_copy ['SAT'],y)
# Define the regression equation, so we can plot it later
#y_hat_yes = 0.6439 + 0.0014 + 0.2226*1 *SAT ##formula from above
y_hat_yes = 0.8665 + 0.0014*dummy_data_copy ['SAT']
#y_hat_no = 0.6439 + 0.0014 + 0.2226*0 *SAT ##formula from above
y_hat_no = 0.6439 + 0.0014*dummy_data_copy ['SAT']
# Plot the regression line against the independent variable (SAT)
fig = plt.plot(dummy_data_copy ['SAT'],y_hat_yes, lw=2, c='red', label ='Attended')
fig = plt.plot(dummy_data_copy ['SAT'],y_hat_no, lw=2, c='green', label ='Did Not Attend')
# Label the axes
plt.xlabel('SAT', fontsize = 25)
plt.ylabel('GPA', fontsize = 25)
plt.show()
# -
# # Make Predictions Based on the Regressions Created
x
# ## We will predict Bob who got 1700 on the SAT but did not attend class
# ## We will predict Alice who got a 1670 on the SAT and attended class
#create a new dataset with our prediction data
new_data = pd.DataFrame({'const': 1, 'SAT': [1700,1670], 'Attendance': [0, 1]})
new_data = new_data[['const', 'SAT', 'Attendance']]
new_data
#optionally rename columns
new_data.rename(index={0: 'Bob', 1: 'Alice'})
predictions = results.predict(new_data)
predictions
#Create a new df with predictions
predictions_df = pd.DataFrame({'Predictions': predictions})
#we will now join the data
joined_data = new_data.join(predictions_df)
joined_data.rename(index={0: 'Bob', 1: 'Alice'})
| Part_5_Advanced_Statistical_Methods_(Machine_Learning)/Multiple Linear Regression/Multiple Linear Regression - GPA Problem/Multiple linear regression and Adjusted R-squared - GPA Problem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.11 64-bit (''latest tf 101021'': conda)'
# name: python3
# ---
import numpy as np
import pandas as pd
from Evaluator import ModelEvaluator
import torch
import random
from matplotlib import pylab as plt
from IPython.display import clear_output
import copy
import math
class MalwareEnv():
def __init__(self,epoch,origin):
# Actions we can take, decrease, increse
self.action_space = 20
# Set start
self.state = self.generateWorld()
#model to evaluate the reward
self.evaluator=ModelEvaluator()
#units to move
self.block=self.calculateBlock(epoch, origin)
def generateWorld(self):
valuesToTest=np.array([])
for x in range(10):
valuesToTest=np.append(valuesToTest, random.randint(-100,100))
return valuesToTest
def calculateBlock(self,epoca,origin):
#return 10
return math.ceil(epoca/origin*60)
def step(self, action):
if(action<9):
self.state[action]+=self.block
elif(action>=9 and action<19):
self.state[action-9]-=self.block
def reward(self):
reward=-1
value=self.state.tolist()
result=self.evaluator.test(value)
result=np.array2string(result)
print(result)
if(result=="['malicious']"):
reward=+10
return reward
def render_np(self):
return self.state
# +
l1 = 10
l2 = 24
l3 = 72
l4 = 20
model = torch.nn.Sequential(
torch.nn.Linear(l1, l2),
torch.nn.ReLU(),
torch.nn.Linear(l2, l3),
torch.nn.ReLU(),
torch.nn.Linear(l3,l4)
)
#device=torch.device("cuda:0" if torch.cuda.is_available else "cpu")
model.cuda()
loss_fn = torch.nn.MSELoss()
learning_rate = 1e-3
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
gamma = 0.9
epsilon = 1.0
# +
initialeEpoch =500
epochs = copy.copy(initialeEpoch)
losses = []
resistance=0
effort=0
for i in range(epochs):
env = MalwareEnv(epochs,initialeEpoch)
state_ = env.render_np().reshape(1,10) + np.random.rand(1,10)/50.0
state1 = torch.from_numpy(state_).float()
status = 1
while(status == 1):
qval = model(state1.cuda())
qval_ = qval.data.cpu().numpy()
if (random.random() < epsilon):
action_ = np.random.randint(0,20)
else:
action_ = np.argmax(qval_)
env.step(action_)
state2_ = env.render_np().reshape(1,10) + np.random.rand(1,10)/50.0
state2 = torch.from_numpy(state2_).float()
reward = env.reward()
with torch.no_grad():
newQ = model(state2.reshape(1,10).cuda())
maxQ = torch.max(newQ)
if reward == -1:
Y = reward + (gamma * maxQ)
else:
Y = reward
Y = torch.Tensor([Y]).detach()
X = qval.squeeze()[action_]
loss = loss_fn(X.cuda(), Y.cuda())
print(resistance, i, loss.item())
clear_output(wait=True)
optimizer.zero_grad()
loss.backward()
if(effort<1000):
losses.append(loss.item())
optimizer.step()
state1 = state2
if reward != -1:
status = 0
resistance=0
effort=0
resistance+=1
effort=+1
if(resistance>120):
resistance=0
env = MalwareEnv(epochs,initialeEpoch)
if epsilon > 0.1:
epsilon -= (1/epochs)
| Android/SL and DRL for android malware detection(PDG)/code 2021/GrayBoxMLWR_gen_vol2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Spark Walmart Data Analysis Project Exercise
# Let's get some quick practice with your new Spark DataFrame skills, you will be asked some basic questions about some stock market data, in this case Walmart Stock from the years 2012-2017. This exercise will just ask a bunch of questions, unlike the future machine learning exercises, which will be a little looser and be in the form of "Consulting Projects", but more on that later!
#
# For now, just answer the questions and complete the tasks below.
# #### Use the walmart_stock.csv file to Answer and complete the tasks below!
# #### Start a simple Spark Session
# +
import findspark
findspark.init('/home/sbhople/spark-2.1.0-bin-hadoop2.7')
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName('walmart').getOrCreate()
# -
# #### Load the Walmart Stock CSV File, have Spark infer the data types.
df = spark.read.csv('walmart_stock.csv', inferSchema=True, header=True)
# #### What are the column names?
df.columns
# #### What does the Schema look like?
df.printSchema()
# #### Print out the first 5 columns.
for line in df.head(5):
print(line, '\n')
# #### Use describe() to learn about the DataFrame.
df.describe().show()
# ## Bonus Question!
# #### There are too many decimal places for mean and stddev in the describe() dataframe. Format the numbers to just show up to two decimal places. Pay careful attention to the datatypes that .describe() returns, we didn't cover how to do this exact formatting, but we covered something very similar. [Check this link for a hint](http://spark.apache.org/docs/latest/api/python/pyspark.sql.html#pyspark.sql.Column.cast)
#
# If you get stuck on this, don't worry, just view the solutions.
# +
'''
from pyspark.sql.types import (StructField, StringType,
IntegerType, StructType)
data_schema = [StructField('summary', StringType(), True),
StructField('Open', StringType(), True),
StructField('High', StringType(), True),
StructField('Low', StringType(), True),
StructField('Close', StringType(), True),
StructField('Volume', StringType(), True),
StructField('Adj Close', StringType(), True)
]
final_struc = StructType(fields=data_schema)
'''
df = spark.read.csv('walmart_stock.csv', inferSchema=True, header=True)
df.printSchema()
#The schema given below is wrong, as it is mostly from an older version.
#Spark is able to predict the schema correctly now
# +
from pyspark.sql.functions import format_number
summary = df.describe()
summary.select(summary['summary'],
format_number(summary['Open'].cast('float'), 2).alias('Open'),
format_number(summary['High'].cast('float'), 2).alias('High'),
format_number(summary['Low'].cast('float'), 2).alias('Low'),
format_number(summary['Close'].cast('float'), 2).alias('Close'),
format_number(summary['Volume'].cast('int'),0).alias('Volume')
).show()
# -
# #### Create a new dataframe with a column called HV Ratio that is the ratio of the High Price versus volume of stock traded for a day.
df_hv = df.withColumn('HV Ratio', df['High']/df['Volume']).select(['HV Ratio'])
df_hv.show()
# #### What day had the Peak High in Price?
df.orderBy(df['High'].desc()).select(['Date']).head(1)[0]['Date']
# #### What is the mean of the Close column?
# +
from pyspark.sql.functions import mean
df.select(mean('Close')).show()
# -
# #### What is the max and min of the Volume column?
# +
from pyspark.sql.functions import min, max
df.select(max('Volume'),min('Volume')).show()
# -
# #### How many days was the Close lower than 60 dollars?
df.filter(df['Close'] < 60).count()
# #### What percentage of the time was the High greater than 80 dollars ?
# #### In other words, (Number of Days High>80)/(Total Days in the dataset)
df.filter('High > 80').count() * 100/df.count()
# #### What is the Pearson correlation between High and Volume?
# #### [Hint](http://spark.apache.org/docs/latest/api/python/pyspark.sql.html#pyspark.sql.DataFrameStatFunctions.corr)
df.corr('High', 'Volume')
# +
from pyspark.sql.functions import corr
df.select(corr(df['High'], df['Volume'])).show()
# -
# #### What is the max High per year?
# +
from pyspark.sql.functions import (dayofmonth, hour,
dayofyear, month,
year, weekofyear,
format_number, date_format)
year_df = df.withColumn('Year', year(df['Date']))
year_df.groupBy('Year').max()['Year', 'max(High)'].show()
# -
# #### What is the average Close for each Calendar Month?
# #### In other words, across all the years, what is the average Close price for Jan,Feb, Mar, etc... Your result will have a value for each of these months.
# +
#Create a new column Month from existing Date column
month_df = df.withColumn('Month', month(df['Date']))
#Group by month and take average of all other columns
month_df = month_df.groupBy('Month').mean()
#Sort by month
month_df = month_df.orderBy('Month')
#Display only month and avg(Close), the desired columns
month_df['Month', 'avg(Close)'].show()
# -
# # Thank you!
| Spark Projects/Spark MLlib/Spark_Walmart_Data_Analysis_Project/Spark Walmart Data Analysis Project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: BCM_pymc3
# language: python
# name: bcm_pymc3
# ---
# # Bayesian Cognitive Modeling in PyMC3
# PyMC3 port of <NAME> Wagenmakers' [Bayesian Cognitive Modeling - A Practical Course](http://bayesmodels.com)
#
# All the codes are in jupyter notebooks with the model explained in distributions (as in the book). Background information of the models please consult the book. You can also compare the result with the original code associated with the book ([WinBUGS and JAGS](https://webfiles.uci.edu/mdlee/Code.zip); [Stan](https://github.com/stan-dev/example-models/tree/master/Bayesian_Cognitive_Modeling))
#
# _All the codes are currently tested under PyMC3 v3.8 master with theano 1.0.4_
# ## Part II - PARAMETER ESTIMATION
#
# ### [Chapter 3: Inferences with binomials](./ParameterEstimation/Binomial.ipynb)
# [3.1 Inferring a rate](./ParameterEstimation/Binomial.ipynb#3.1-Inferring-a-rate)
# [3.2 Difference between two rates](./ParameterEstimation/Binomial.ipynb#3.2-Difference-between-two-rates)
# [3.3 Inferring a common rate](./ParameterEstimation/Binomial.ipynb#3.3-Inferring-a-common-rate)
# [3.4 Prior and posterior prediction](./ParameterEstimation/Binomial.ipynb#3.4-Prior-and-posterior-prediction)
# [3.5 Posterior prediction](./ParameterEstimation/Binomial.ipynb#3.5-Posterior-Predictive)
# [3.6 Joint distributions](./ParameterEstimation/Binomial.ipynb#3.6-Joint-distributions)
#
# ### [Chapter 4: Inferences with Gaussians](./ParameterEstimation/Gaussian.ipynb)
# [4.1 Inferring a mean and standard deviation](./ParameterEstimation/Gaussian.ipynb#4.1-Inferring-a-mean-and-standard-deviation)
# [4.2 The seven scientists](./ParameterEstimation/Gaussian.ipynb#4.2-The-seven-scientists)
# [4.3 Repeated measurement of IQ](./ParameterEstimation/Gaussian.ipynb#4.3-Repeated-measurement-of-IQ)
#
# ### [Chapter 5: Some examples of data analysis](./ParameterEstimation/DataAnalysis.ipynb)
# [5.1 Pearson correlation](./ParameterEstimation/DataAnalysis.ipynb#5.1-Pearson-correlation)
# [5.2 Pearson correlation with uncertainty](./ParameterEstimation/DataAnalysis.ipynb#5.2-Pearson-correlation-with-uncertainty)
# [5.3 The kappa coefficient of agreement](./ParameterEstimation/DataAnalysis.ipynb#5.3-The-kappa-coefficient-of-agreement)
# [5.4 Change detection in time series data](./ParameterEstimation/DataAnalysis.ipynb#5.4-Change-detection-in-time-series-data)
# [5.5 Censored data](./ParameterEstimation/DataAnalysis.ipynb#5.5-Censored-data)
# [5.6 Recapturing planes](./ParameterEstimation/DataAnalysis.ipynb#5.6-Recapturing-planes)
#
# ### [Chapter 6: Latent-mixture models](./ParameterEstimation/Latent-mixtureModels.ipynb)
# [6.1 Exam scores](./ParameterEstimation/Latent-mixtureModels.ipynb#6.1-Exam-scores)
# [6.2 Exam scores with individual differences](./ParameterEstimation/Latent-mixtureModels.ipynb#6.2-Exam-scores-with-individual-differences)
# [6.3 Twenty questions](./ParameterEstimation/Latent-mixtureModels.ipynb#6.3-Twenty-questions)
# [6.4 The two-country quiz](./ParameterEstimation/Latent-mixtureModels.ipynb#6.4-The-two-country-quiz)
# [6.5 Assessment of malingering](./ParameterEstimation/Latent-mixtureModels.ipynb#6.5-Assessment-of-malingering)
# [6.6 Individual differences in malingering](./ParameterEstimation/Latent-mixtureModels.ipynb#6.6-Individual-differences-in-malingering)
# [6.7 Alzheimer’s recall test cheating](./ParameterEstimation/Latent-mixtureModels.ipynb#6.7-Alzheimer's-recall-test-cheating)
# ## Part III - MODEL SELECTION
#
# ### [Chapter 8: Comparing Gaussian means](./ModelSelection/ComparingGaussianMeans.ipynb)
# [8.1 One-sample comparison](./ModelSelection/ComparingGaussianMeans.ipynb#8.1-One-sample-comparison)
# [8.2 Order-restricted one-sample comparison](./ModelSelection/ComparingGaussianMeans.ipynb#8.2-Order-restricted-one-sample-comparison)
# [8.3 Two-sample comparison](./ModelSelection/ComparingGaussianMeans.ipynb#8.3-Two-sample-comparison)
#
# ### [Chapter 9: Comparing binomial rates](./ModelSelection/ComparingBinomialRates.ipynb)
# [9.1 Equality of proportions](./ModelSelection/ComparingBinomialRates.ipynb#9.1-Equality-of-proportions)
# [9.2 Order-restricted equality of proportions](./ModelSelection/ComparingBinomialRates.ipynb#9.2-Order-restricted-equality-of-proportions)
# [9.3 Comparing within-subject proportions](./ModelSelection/ComparingBinomialRates.ipynb#9.3-Comparing-within-subject-proportions)
# [9.4 Comparing between-subject proportions](./ModelSelection/ComparingBinomialRates.ipynb#9.4-Comparing-between-subject-proportions)
# [9.5 Order-restricted between-subjects comparison](./ModelSelection/ComparingBinomialRates.ipynb#9.5-Order-restricted-between-subject-proportions)
# ## Part IV - CASE STUDIES
#
# ### [Chapter 10: Memory retention](./CaseStudies/MemoryRetention.ipynb)
# [10.1 No individual differences](./CaseStudies/MemoryRetention.ipynb#10.1-No-individual-differences)
# [10.2 Full individual differences](./CaseStudies/MemoryRetention.ipynb#10.2-Full-individual-differences)
# [10.3 Structured individual differences](./CaseStudies/MemoryRetention.ipynb#10.3-Structured-individual-differences)
#
# ### [Chapter 11: Signal detection theory](./CaseStudies/SignalDetectionTheory.ipynb)
# [11.1 Signal detection theory](./CaseStudies/SignalDetectionTheory.ipynb#11.1-Signal-detection-theory)
# [11.2 Hierarchical signal detection theory](./CaseStudies/SignalDetectionTheory.ipynb#11.2-Hierarchical-signal-detection-theory)
# [11.3 Parameter expansion](./CaseStudies/SignalDetectionTheory.ipynb#11.3-Parameter-expansion)
#
# ### [Chapter 12: Psychophysical functions](./CaseStudies/PsychophysicalFunctions.ipynb)
# [12.1 Psychophysical functions](./CaseStudies/PsychophysicalFunctions.ipynb#12.1-Psychophysical-functions)
# [12.2 Psychophysical functions under contamination](./CaseStudies/PsychophysicalFunctions.ipynb#12.2-Psychophysical-functions-under-contamination)
#
# ### [Chapter 13: Extrasensory perception](./CaseStudies/ExtrasensoryPerception.ipynb)
# [13.1 Evidence for optional stopping](./CaseStudies/ExtrasensoryPerception.ipynb#13.1-Evidence-for-optional-stopping)
# [13.2 Evidence for differences in ability](./CaseStudies/ExtrasensoryPerception.ipynb#13.2-Evidence-for-differences-in-ability)
# [13.3 Evidence for the impact of extraversion](./CaseStudies/ExtrasensoryPerception.ipynb#13.3-Evidence-for-the-impact-of-extraversion)
#
# ### [Chapter 14: Multinomial processing trees](./CaseStudies/MultinomialProcessingTrees.ipynb)
# [14.1 Multinomial processing model of pair-clustering](./CaseStudies/MultinomialProcessingTrees.ipynb#14.1-Multinomial-processing-model-of-pair-clustering)
# [14.2 Latent-trait MPT model](./CaseStudies/MultinomialProcessingTrees.ipynb#14.2-Latent-trait-MPT-model)
#
# ### [Chapter 15: The SIMPLE model of memory](./CaseStudies/TheSIMPLEModelofMemory.ipynb)
# [15.1 The SIMPLE model](./CaseStudies/TheSIMPLEModelofMemory.ipynb#15.1-The-SIMPLE-model)
# [15.2 A hierarchical extension of SIMPLE](./CaseStudies/TheSIMPLEModelofMemory.ipynb#15.2-A-hierarchical-extension-of-SIMPLE)
#
# ### [Chapter 16: The BART model of risk taking](./CaseStudies/TheBARTModelofRiskTaking.ipynb)
# [16.1 The BART model](./CaseStudies/TheBARTModelofRiskTaking.ipynb#16.1-The-BART-model)
# [16.2 A hierarchical extension of the BART model](./CaseStudies/TheBARTModelofRiskTaking.ipynb#16.2-A-hierarchical-extension-of-the-BART-model)
#
# ### [Chapter 17: The GCM model of categorization](./CaseStudies/TheGCMModelofCategorization.ipynb)
# [17.1 The GCM model](./CaseStudies/TheGCMModelofCategorization.ipynb#17.1-The-GCM-model)
# [17.2 Individual differences in the GCM](./CaseStudies/TheGCMModelofCategorization.ipynb#17.2-Individual-differences-in-the-GCM)
# [17.3 Latent groups in the GCM](./CaseStudies/TheGCMModelofCategorization.ipynb#17.3-Latent-groups-in-the-GCM)
#
# ### [Chapter 18: Heuristic decision-making](./CaseStudies/HeuristicDecisionMaking.ipynb)
# [18.1 Take-the-best](./CaseStudies/HeuristicDecisionMaking.ipynb#18.1-Take-the-best)
# [18.2 Stopping](./CaseStudies/HeuristicDecisionMaking.ipynb#18.2-Stopping)
# [18.3 Searching](./CaseStudies/HeuristicDecisionMaking.ipynb#18.3-Searching)
# [18.4 Searching and stopping](./CaseStudies/HeuristicDecisionMaking.ipynb#18.4-Searching-and-stopping)
#
# ### [Chapter 19: Number concept development](./CaseStudies/NumberConceptDevelopment.ipynb)
# [19.1 Knower-level model for Give-N](./CaseStudies/NumberConceptDevelopment.ipynb#19.1-Knower-level-model-for-Give-N)
# [19.2 Knower-level model for Fast-Cards](./CaseStudies/NumberConceptDevelopment.ipynb#19.2-Knower-level-model-for-Fast-Cards)
# [19.3 Knower-level model for Give-N and Fast-Cards](./CaseStudies/NumberConceptDevelopment.ipynb#19.3-Knower-level-model-for-Give-N-and-Fast-Cards)
# Python Environment and library version
# %load_ext watermark
# %watermark -v -n -u -w -p pymc3,theano,scipy,numpy,pandas,matplotlib,seaborn -m
| BCM/index.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# (PTDN)=
# # 1.6 Polinomios de Taylor y diferenciación numérica
# ```{admonition} Notas para contenedor de docker:
#
# Comando de docker para ejecución de la nota de forma local:
#
# nota: cambiar `<ruta a mi directorio>` por la ruta de directorio que se desea mapear a `/datos` dentro del contenedor de docker.
#
# `docker run --rm -v <ruta a mi directorio>:/datos --name jupyterlab_optimizacion -p 8888:8888 -d palmoreck/jupyterlab_optimizacion:2.1.4`
#
# password para jupyterlab: `<PASSWORD>`
#
# Detener el contenedor de docker:
#
# `docker stop jupyterlab_optimizacion`
#
# Documentación de la imagen de docker `palmoreck/jupyterlab_optimizacion:2.1.4` en [liga](https://github.com/palmoreck/dockerfiles/tree/master/jupyterlab/optimizacion).
#
# ```
# ---
# Nota generada a partir de la [liga1](https://www.dropbox.com/s/jfrxanjls8kndjp/Diferenciacion_e_Integracion.pdf?dl=0), [liga2](https://www.dropbox.com/s/mmd1uzvwhdwsyiu/4.3.2.Teoria_de_convexidad_Funciones_convexas.pdf?dl=0) e inicio de [liga3](https://www.dropbox.com/s/ko86cce1olbtsbk/4.3.1.Teoria_de_convexidad_Conjuntos_convexos.pdf?dl=0).
# ```{admonition} Al final de esta nota el y la lectora:
# :class: tip
#
# * Aprenderá que el método de diferenciación finita es un método inestable numéricamente respecto al redondeo.
#
# * Conocerá las expresiones de los polinomios de Taylor para funciones de varias variables.
#
# * Tendrá en su lista de programas del lenguaje *R* implementaciones para aproximar al gradiente y a la Hessiana de una función con los métodos de diferenciación finita.
#
# ```
# ## Problema: ¿Cómo aproximar una función $f$ en un punto $x_1$?
# Si $f$ es continuamente diferenciable en $x_0$ y $f^{(1)}, f^{(2)}$ existen y están acotadas en $x_0$ entonces:
#
# $$f(x_1) \approx f(x_0) + f^{(1)}(x_0)(x_1-x_0)$$
# y se nombra **aproximación de orden 1**. Ver {ref}`Definición de función, continuidad y derivada <FCD>` para definición de continuidad, diferenciabilidad y propiedades.
# ```{admonition} Comentarios
#
# * Lo anterior requiere de los valores: $x_0, x_1, f(x_0), f^{(1)}(x_0)$. Esta aproximación tiene un error de **orden** $2$ pues su error es **proporcional** al cuadrado del ancho del intervalo: $h=x_1-x_0$, esto es, si reducimos a la mitad $h$ entonces el error se reduce en una cuarta parte.
#
# * Otra aproximación más simple sería:
#
# $$f(x_1) \approx f(x_0)$$
#
# lo cual sólo requiere del conocimiento de $f(x_0)$ y se nombra aproximación de **orden** $0$, sin embargo esta aproximación tiene un error de **orden** $1$ pues este es proporcional a $h$ , esto es, al reducir a la mitad $h$ se reduce a la mitad el error.
#
# * Los errores anteriores los nombramos errores por **truncamiento**, ver {ref}`Fuentes del error <FuenErr>` y {ref}`Análisis del error <AnErr>` para un recordatorio de tal error. Utilizamos la notación "O grande" $\mathcal{O}(\cdot)$ para escribir lo anterior:
#
# $$f(x)-f(x_0)=\mathcal{O}(h)$$
#
# con la variable $h=x-x_0$. En este caso se representa a un error de orden $1$. Análogamente:
#
# $$f(x)-(f(x_0)+f^{(1)}(x_0)(x-x_0)) = \mathcal{O}(h^2)$$
#
# y se representa un error de orden $2$.
#
# ```
# ```{admonition} Observaciones
# :class: tip
#
# * No confundir órdenes de una aproximación con órdenes de error.
#
# * Otras aproximaciones a una función se pueden realizar con:
#
# * Interpoladores polinomiales (representación por Vandermonde, Newton, Lagrange).
# ```
# ## Aproximación a una función por el teorema de Taylor
# En esta sección se presenta el teorema de Taylor, el cual, bajo ciertas hipótesis nos proporciona una expansión de una función alrededor de un punto. Este teorema será utilizado en **diferenciación e integración numérica**. El teorema es el siguiente:
# ```{admonition} Teorema de Taylor
#
# Sea $f: \mathbb{R} \rightarrow \mathbb{R}$, $f \in \mathcal{C}^n([a,b])$ tal que $f^{(n+1)}$ existe en [a,b]. Si $x_0 \in [a,b]$ entonces $\forall x \in [a,b]$ se tiene: $f(x) = P_n(x) + R_n(x)$ donde:
#
# $$P_n(x) = \displaystyle \sum_{k=0}^n \frac{f^{(k)}(x_0)(x-x_0)^k}{k!} \quad (f^{(0)} = f)$$ y
#
# $$R_n(x) = \frac{f^{(n+1)}(\xi_x)(x-x_0)^{(n+1)}}{(n+1)!}$$
#
# con $\xi_x$ entre $x_0, x$ y $x_0$ se llama centro. Ver {ref}`Definición de función, continuidad y derivada <FCD>` para definición del conjunto $\mathcal{C}^n([a,b])$.
#
# ```
# ```{admonition} Comentarios
#
# * El teorema de Taylor nos indica que cualquier función suave (función en $\mathcal{C}^n)$ se le puede aproximar por un polinomio en el intervalo $[a,b]$, de hecho $f(x) \approx P_n(x)$.
#
# * Si el residuo no tiene una alta contribución a la suma $P_n(x) + R_n(x)$ entonces es una buena aproximación **local** (alta contribución y buena aproximación depende de factores como elección de la norma y la aplicación).
#
# * El teorema de Taylor es una generalización del [teorema del valor medio para derivadas](https://en.wikipedia.org/wiki/Mean_value_theorem).
#
# * $P_n(x)$ se le llama polinomio de Taylor alrededor de $x_0$ de orden $n$ y $R_n(x)$ es llamado residuo de Taylor alrededor de $x_0$ de orden $n+1$, tiene otras expresiones para representarlo y la que se utiliza en el enunciado anterior es en su forma de Lagrange (ver [liga](https://en.wikipedia.org/wiki/Taylor%27s_theorem) para otras expresiones del residuo).
#
# * $\xi_x$ es un punto entre $x_0, x$ desconocido y está en función de $x$ (por eso se le escribe un subíndice).
#
# * Una forma del teorema de Taylor es escribirlo definiendo a la variable $h=x-x_0$:
#
# $$f(x) = f(x_0+h) = P_n(h) + R_n(h) = \displaystyle \sum_{k=0}^n \frac{f^{(k)}(x_0)h^k}{k!} + \frac{f^{(n+1)}(\xi_h)h^{n+1}}{(n+1)!}$$
#
# y si $f^{(n+1)}$ es acotada, escribimos: $R_n(h) = \mathcal{O}(h^{n+1})$.
# ```
# ### Ejemplo
# Graficar la función y los polinomios de Taylor de grados $0,1,2,3$ y $4$ en una sola gráfica para el intervalo $[1,2]$ de la función $\frac{1}{x}$ con centro en $x_0=1.5$. ¿Cuánto es la aproximación de los polinomios en x=1.9?. Calcula el error relativo de tus aproximaciones.
# **Solución**
# Obtengamos los polinomios de Taylor de orden $n$ con $n \in \{0,1,2, 3\}$ y centro en $x_0=1.5$ para la función $\frac{1}{x}$ en el intervalo $[1,2]$. Los primeros tres polinomios de Taylor son:
# $$P_0(x) = f(x_0) = \frac{2}{3} \quad \text{(constante)}$$
# $$P_1(x) = f(x_0) + f^{(1)}(x_0)(x-x_0) = \frac{2}{3} - \frac{1}{x_0^2}(x-x_0) =\frac{2}{3} - \frac{1}{1.5^2}(x-1.5) \quad \text{(lineal)}$$
# $$
# \begin{eqnarray}
# P_2(x) &=& f(x_0) + f^{(1)}(x_0)(x-x_0) + \frac{f^{(2)}(x_0)(x-x_0)^2}{2} \nonumber \\
# &=& \frac{2}{3} - \frac{1}{x_0^2}(x-x_0) + \frac{1}{x_0^3}(x-x_0)^2 \nonumber \\
# &=& \frac{2}{3} -\frac{1}{1.5^2}(x-1.5) + \frac{1}{1.5^3}(x-1.5)^2 \quad \text{(cuadrático)} \nonumber
# \end{eqnarray}
# $$
library(ggplot2)
options(repr.plot.width=6, repr.plot.height=6) #esta línea sólo se ejecuta para jupyterlab con R
Taylor_approx <- function(x,c,n){
'
Taylor approximation for 1/x function. Will return Taylor polynomial of degree n with
center in c and evaluated in x.
Args:
x (double): numeric vector or scalar in which Taylor polynomial will be evaluated.
c (double): scalar which represents center of Taylor polynomial of degree n.
n (integer): scalar which represents degree of Taylor polynomial.
Returns:
sum (double): scalar evaluation of Taylor polynomial of degree n with center c in x.
'
length_x <- length(x)
sum <- vector("double", length_x)
for(j in 1:length_x){
mult <- c^(-1)
sum[j] <- mult
for(k in 1:n){
mult <- -1*c^(-1)*(x[j]-c)*mult
sum[j] <- sum[j] + mult
}
}
sum #accumulated sum
}
x0 <- 1.5
x <- seq(from=1,to=2,by=.005)
n <- c(0,1,2,3,4) #degrees of Taylor polynomials
f <- function(z)1/z
y <- f(x)
y_Taylor_0 <- f(x0)*(vector("double", length(x))+1)
y_Taylor_1 <- Taylor_approx(x,x0,1)
y_Taylor_2 <- Taylor_approx(x,x0,2)
y_Taylor_3 <- Taylor_approx(x,x0,3)
y_Taylor_4 <- Taylor_approx(x,x0,4)
gg <- ggplot()
print(gg+
geom_line(aes(x=x,y=y,color='f(x)')) +
geom_line(aes(x=x,y=y_Taylor_0,color='constante'))+
geom_line(aes(x=x,y=y_Taylor_1,color='lineal')) +
geom_line(aes(x=x,y=y_Taylor_2,color='grado 2')) +
geom_line(aes(x=x,y=y_Taylor_3,color='grado 3')) +
geom_line(aes(x=x,y=y_Taylor_4,color='grado 4')) +
geom_point(aes(x=x0, y=f(x0)), color='blue',size=3))
# ```{admonition} Observación
# :class: tip
#
# Para cualquier aproximación calculada siempre es una muy buena idea reportar el error relativo de la aproximación si tenemos el valor del objetivo. No olvidar esto :)
#
# ```
compute_error_point_wise<-function(obj, approx){
'
Relative or absolute error between approx and obj in a point wise fashion.
'
if (abs(obj) > .Machine$double.eps){
Err<-abs(obj-approx)/abs(obj)
}else
Err<-abs(obj-approx)
Err
}
# +
x_test_point <- 1.9
objective <- f(x_test_point)
#Approximations
p1_approx <- Taylor_approx(x_test_point, x0, 1)
p2_approx <- Taylor_approx(x_test_point, x0, 2)
p3_approx <- Taylor_approx(x_test_point, x0, 3)
p4_approx <- Taylor_approx(x_test_point, x0, 4)
print('error relativo polinomio constante')
print(compute_error_point_wise(objective, 1/x0))
print('error relativo polinomio lineal')
print(compute_error_point_wise(objective, p1_approx))
print('error relativo polinomio grado 2')
print(compute_error_point_wise(objective, p2_approx))
print('error relativo polinomio grado 3')
print(compute_error_point_wise(objective, p3_approx))
print('error relativo polinomio grado 4')
print(compute_error_point_wise(objective, p4_approx))
# -
# ```{admonition} Ejercicio
# :class: tip
#
# Aproximar $f(1)$ con polinomios de Taylor de orden $0,1,2,3,4$ si $f(x)=-0.1x^4-0.15x^3-0.5x^2-0.25x+1.2$ con centro en $x0=0$. Calcula los errores relativos de tus aproximaciones. Realiza las gráficas de cada polinomio en el intervalo $[0,1]$ con `ggplot2`. Observa que $R_5(x)$ es cero.
# ```
# (TEOTAYLORNVARIABLES)=
# ## Teorema de Taylor para una función $f: \mathbb{R}^n \rightarrow \mathbb{R}$
# Sea $f: \mathbb{R}^n \rightarrow \mathbb{R}$ diferenciable en $\text{dom}f$. Si $x_0, x \in \text{dom}f$ y $x_0+t(x-x_0) \in \text{dom}f, \forall t \in (0,1),$ entonces $\forall x \in \text{dom}f$ se tiene $f(x) = P_0(x) + R_0(x)$ donde:
#
# $$P_0(x) = f(x_0)$$
# $$R_0(x) = \nabla f(x_0+t_x(x-x_0))^T(x-x_0)$$
# para alguna $t_x \in (0,1)$ y $\nabla f(\cdot)$ gradiente de $f$, ver {ref}`Definición de función, continuidad y derivada <FCD>` para definición del gradiente de una función.
# ```{admonition} Observación
# :class: tip
#
# La aproximación anterior la nombramos **aproximación de orden $0$** para $f$ con centro en $x_0$. Si $\nabla f(\cdot)$ es acotado en $\text{dom}f$ entonces se escribe: $R_0(x)=\mathcal{O}(||x-x_0||)$.
# ```
# Si además $f$ es continuamente diferenciable en $\text{dom}f$(su derivada es continua, ver {ref}`Definición de función, continuidad y derivada <FCD>` para definición de continuidad), $f^{(2)}$ existe en $\text{dom}f$, se tiene $f(x) = P_1(x) + R_1(x)$ donde:
#
# $$P_1(x) = f(x_0) + \nabla f(x_0)^T(x-x_0)$$
# $$R_1(x) = \frac{1}{2}(x-x_0)^T \nabla ^2f(x_0+t_x(x-x_0))(x-x_0)$$
# para alguna $t_x \in (0,1)$ y $\nabla^2 f(\cdot)$ Hessiana de $f$ (ver {ref}`Definición de función, continuidad y derivada <FCD>` para definición de la matriz Hessiana).
# ```{admonition} Observación
# :class: tip
#
# La aproximación anterior la nombramos **aproximación de orden $1$** para $f$ con centro en $x_0$. Si $\nabla^2f(\cdot)$ es acotada en $\text{dom}f$ entonces se escribe: $R_1(x) = \mathcal{O}(||x-x_0||^2)$.
# ```
# Si $f^{(2)}$ es continuamente diferenciable y $f^{(3)}$ existe y es acotada en $\text{dom}f$, se tiene $f(x)=P_2(x) + R_2(x)$ donde:
#
# $$P_2(x) = f(x_0) + \nabla f(x_0)^T(x-x_0) + \frac{1}{2}(x-x_0)^T \nabla ^2f(x_0)(x-x_0)$$
# ```{admonition} Observación
# :class: tip
#
# * La aproximación anterior la nombramos **aproximación de orden $2$** para $f$ con centro en $x_0$. Para las suposiciones establecidas se tiene:
#
# $$R_2(x)= \mathcal{O}(||x-x_0||^3).$$
#
# * En este caso $f^{(3)}$ es un tensor.
#
# ```
# ```{admonition} Comentario
#
# Tomando $h=x-x_0$, se reescribe el teorema como sigue, por ejemplo para la aproximación de orden $1$ incluyendo su residuo:
#
# $$f(x) = f(x_0 + h) = \underbrace{f(x_0) + \nabla f(x_0)^Th}_{\textstyle P_1(h)} + \underbrace{\frac{1}{2}h^T \nabla ^2f(x_0+t_xh)h}_{\textstyle R_1(h)}.$$
#
# Si $f^{(2)}$ es acotada en $\text{dom}f$, escribimos: $R_1(h)=\mathcal{O}(||h||^2)$.
# ```
# (DIFNUMDIFFINITAS)=
# ## Diferenciación numérica por diferencias finitas
# ```{admonition} Comentario
#
# En esta sección se revisan métodos numéricos para aproximar las derivadas. Otros métodos para el cálculo de las derivadas se realizan con el cómputo simbólico o algebraico, ver {ref}` Definición de función, continuidad y derivada <FCD>` para ejemplos.
#
# ```
# Las fórmulas de aproximación a las derivadas por diferencias finitas pueden obtenerse con los polinomios de Taylor, presentes en el teorema del mismo autor, por ejemplo:
# Sea $f \in \mathcal{C}^1([a,b])$ y $f^{(2)}$ existe y está acotada $\forall x \in [a,b]$ entonces, si $x+h \in [a,b]$ con $h>0$ por el teorema de Taylor:
#
# $$f(x+h) = f(x) + f^{(1)}(x)h + f^{(2)}(\xi_{x+h})\frac{h^2}{2}$$ con $\xi_{x+h} \in [x,x+h]$ y al despejar $f^{(1)}(x)$ se tiene:
# $$f^{(1)}(x) = \frac{f(x+h)-f(x)}{h} - f^{(2)}(\xi_{x+h})\frac{h}{2}.$$
# y escribimos:
#
# $$f^{(1)}(x) = \frac{f(x+h)-f(x)}{h} + \mathcal{O}(h).$$
#
# La aproximación $\frac{f(x+h)-f(x)}{h}$ es una fórmula por diferencias hacia delante con error de orden $1$. Gráficamente se tiene:
#
# <img src="https://dl.dropboxusercontent.com/s/r1ypkxkwa9g3pmk/dif_hacia_delante.png?dl=0" heigth="500" width="500">
# Con las mismas suposiciones es posible obtener la fórmula para la aproximación por diferencias hacia atrás:
#
# $$f^{(1)}(x) = \frac{f(x)-f(x-h)}{h} + \mathcal{O}(h), h >0.$$
#
# <img src="https://dl.dropboxusercontent.com/s/mxmc8kohurlu9sp/dif_hacia_atras.png?dl=0" heigth="500" width="500">
# Considerando $f \in \mathcal{C}^2([a,b]), f^{(3)}$ existe y está acotada $\forall x \in [a,b]$ si $x-h, x+h \in [a,b]$ y $h>0$ entonces:
#
# $$f^{(1)}(x) = \frac{f(x+h)-f(x-h)}{2h} + \mathcal{O}(h^2), h >0.$$
#
# y el cociente $\frac{f(x+h)-f(x-h)}{2h}$ es la aproximación por diferencias centradas con error de orden $2$. Gráficamente:
#
# <img src="https://dl.dropboxusercontent.com/s/jwi0y1t5z58pydz/dif_centradas.png?dl=0" heigth="500" width="500">
# ```{admonition} Observaciones
# :class: tip
#
# * La aproximación por diferencias finitas a la primer derivada de la función tiene un error de orden $\mathcal{O}(h)$ por lo que una elección de $h$ igual a $.1 = 10^{-1}$ generará aproximaciones con alrededor de un dígito correcto.
#
# * La diferenciación numérica por diferencias finitas **no es un proceso con una alta exactitud** pues los problemas del redondeo de la aritmética en la máquina se hacen presentes en el mismo (ver nota {ref}`Sistema de punto flotante <SPF>`). Como ejemplo de esta situación realicemos el siguiente ejemplo.
#
# ```
# ### Ejemplo
# Realizar una gráfica de log(error relativo) vs log(h) (h en el eje horizontal) para aproximar la primera derivada de $f(x)=e^{-x}$ en $x=1$ con $h \in \{10^{-16}, 10^{-14}, \dots , 10^{-1}\}$ y diferencias hacia delante. Valor a aproximar: $f^{(1)}(1) = -e^{-1}$.
# **Definimos la función**
f <- function(x){
exp(-x)
}
# **Definimos la aproximación numérica por diferencias finitas a la primera derivada**
approx_first_derivative <- function(f, x, h){
'
Numerical differentiation by finite differences. Uses forward point formula
to approximate first derivative of function.
Args:
f (function): function definition.
x (float): point where first derivative will be approximated
h (float): step size for forward differences. Tipically less than 1
Returns:
res (float): approximation to first_derivative.
'
res <- (f(x+h)-f(x))/h
res
}
# **Puntos donde se evaluará la aproximación:**
# +
x<-1
h<-10^(-1*(1:16))
# -
# **Aproximación numérica:**
approx_df <- approx_first_derivative(f,x,h)
# **Derivada de la función:**
df<-function(x){
-exp(-x)
}
obj_df <- df(x)
# **Cálculo de errores:**
res_relative_error <- compute_error_point_wise(obj_df, approx_df)
# **Gráfica:**
gf <- ggplot()
print(gf+
geom_line(aes(x=log(h),y=log(res_relative_error)))+
ggtitle('Aproximación a la primera derivada por diferencias finitas'))
# ```{admonition} Ejercicio
# :class: tip
#
# Realizar una gráfica de log(error relativo) vs log(h) (h en el eje horizontal) con `ggplot2` para aproximar la segunda derivada de $f(x)=e^{-x}$ en $x=1$ con $h \in \{10^{-16}, 10^{-14}, \dots , 10^{-1}\}$ y diferencias hacia delante. Valor a aproximar: $f^{(2)}(1) = e^{-1}$. Usar:
#
# $$\frac{d^2f(x)}{dx} = \frac{f(x+2h)-2f(x+h)+f(x)}{h^2} + \mathcal{O}(h)$$
#
# Encontrar valor(es) de $h$ que minimiza(n) al error absoluto y relativo.
# ```
# ```{admonition} Comentario
#
# Aproximaciones a la segunda derivada de una función $f: \mathbb{R} \rightarrow \mathbb{R}$ se pueden obtener con las fórmulas:
#
# * $\frac{d^2f(x)}{dx} = \frac{f(x+2h)-2f(x+h)+f(x)}{h^2} + \mathcal{O}(h)$ por diferencias hacia delante.
#
# * $\frac{d^2f(x)}{dx} = \frac{f(x)-2f(x-h)+f(x-2h)}{h^2} + \mathcal{O}(h)$ por diferencias hacia atrás.
#
# * $\frac{d^2f(x)}{dx} = \frac{f(x+h)-2f(x)+f(x-h)}{h^2} + \mathcal{O}(h^2)$ por diferencias centradas.
#
# Estas fórmulas se obtienen con el teorema de Taylor bajo las suposiciones correctas.
#
# ```
#
# ## Análisis del error por redondeo y truncamiento en aproximación por diferencias finitas hacia delante
# El ejemplo anterior muestra (vía una gráfica) que el método numérico de diferenciación numérica no es estable numéricamente respecto al redondeo (ver nota {ref}`Condición de un problema y estabilidad de un algoritmo <CPEA>` para definición de estabilidad de un algoritmo) y también se puede corroborar realizando un análisis del error. En esta sección consideramos la aproximación a la primer derivada por diferencias finitas hacia delante:
#
# $$\frac{f(x+h)-f(x)}{h}$$
# Suponemos que $\hat{f}(x)$ aproxima a $f(x)$ y por errores de redondeo $\hat{f}(x) = f(x)(1 + \epsilon_{f(x)})$ con $|\epsilon_{f(x)}| \leq \epsilon_{maq}$ error de redondeo al evaluar $f$ en $x$. $\hat{f}(x)$ es la aproximación en un SPFN (ver nota {ref}`Sistema de punto flotante <SPF>`). Además supóngase que $x,x+h,h \in \mathcal{Fl}$ . Entonces en la aproximación a la primer derivada por diferencias hacia delante:
# $f^{(1)}(x) = \frac{f(x+h)-f(x)}{h} + \mathcal{O}(h)$ y calculando el error absoluto:
#
# $$
# \begin{eqnarray}
# \text{ErrAbs}\left ( \frac{\hat{f}(x+h)-\hat{f}(x)}{h} \right ) &=& \left |f^{(1)}(x) - \frac{\hat{f}(x+h)-\hat{f}(x)}{h} \right | \nonumber \\
# &=& \left | \frac{f(x+h)-f(x)}{h} + \mathcal{O}(h) - \left ( \frac{f(x+h)(1+\epsilon_{f(x+h)})-f(x)(1+\epsilon_{f(x)})}{h} \right ) \right | \nonumber \\
# &=& \left |\mathcal{O}(h) - \frac{f(x+h)\epsilon_{f(x+h)}-f(x)\epsilon_{f(x)}}{h} \right | \nonumber \\
# &\leq& \mathcal{O}(h) + \frac{C\epsilon_{maq}}{h}
# \end{eqnarray}
# $$
# suponiendo en el último paso que $|f(x+h)\epsilon_{f(x+h)} - f(x)\epsilon_{f(x)}| \leq C \epsilon_{maq}$ con $C>0$ constante que acota a la función $f$ en el intervalo $[a,b]$. Obsérvese que $\frac{\hat{f}(x+h)-\hat{f}(x)}{h}$ es la aproximación a la primer derivada por diferencias hacia delante que se obtiene en la computadora, por lo que la cantidad $\left |f^{(1)}(x) - \frac{\hat{f}(x+h)-\hat{f}(x)}{h} \right |$ es el error absoluto de la aproximación por diferencias hacia delante.
# El error relativo es:
# $$\text{ErrRel}\left (\frac{\hat{f}(x+h)-\hat{f}(x)}{h} \right) = \frac{\text{ErrAbs}\left ( \frac{\hat{f}(x+h)-\hat{f}(x)}{h} \right )}{|f^{(1)}(x)|} \leq \frac{\mathcal{O}(h) + \frac{C\epsilon_{maq}}{h}}{|f^{(1)}(x)|} = K_1h + K_2\frac{1}{h}$$
#
# con $K_1, K_2 >0$ constantes.
# Entonces la función $g(h) = \mathcal{O}(h) + \mathcal{O}\left (\frac{1}{h} \right)$ acota al error absoluto y al error relativo y se tiene:
#
# * Si $h \rightarrow 0$ la componente $\mathcal{O} \left ( \frac{1}{h} \right )$ domina a la componente $\mathcal{O}(h)$, la cual tiende a $0$.
#
# * Si $h \rightarrow \infty$ la componente $\mathcal{O}(h)$ domina a $\mathcal{O} \left ( \frac{1}{h} \right )$, la cual tiende a $0$.
#
# Por lo anterior, existe un valor de $h$ que minimiza a los errores. Tal valor se observa en las gráficas anteriores y es igual a:
print(h[which.min(res_relative_error)])
# ```{admonition} Ejercicio
# :class: tip
#
# Obtener de forma analítica el valor de $h$ que minimiza la función $g(h)$ anterior. Tip: utilizar criterio de primera y segunda derivada para encontrar mínimo global.
# ```
# ### Conclusiones y comentarios
#
# * La componente $\mathcal{O}(h)$ es el error por truncamiento, la cual resulta del teorema de Taylor. El teorema de Taylor nos indica que añadir términos en el polinomio de Taylor si la $x$ a aproximar es cercana al centro, las derivadas de $f$ son acotadas y $h \rightarrow 0$ entonces el error por truncamiento debe tender a $0$. Lo anterior no ocurre en la implementación numérica (corroborado de forma analítica y visual) del método por diferenciación numérica para la primer derivada por la presencia de la componente $\mathcal{O}\left(\frac{1}{h}\right)$ en los errores. Tal componente proviene del error por redondeo.
#
# * Obsérvese que el error relativo máximo es del $100\%$ lo que indica que no se tiene ninguna cifra correcta en la aproximación:
print(max(res_relative_error))
# y esto ocurre para un valor de $h$ igual a:
print(h[which.max(res_relative_error)])
# ```{admonition} Pregunta
# :class: tip
#
# ¿Por qué se alcanza el máximo error relativo en el valor de $h=10^{-16}$?.
# ```
# * Con lo anterior se tiene que la diferenciación numérica es un método **inestable numéricamente respecto al redondeo**. Ver nota {ref}`Condición de un problema y estabilidad de un algoritmo <CPEA>`.
# * Un análisis de error similar se utiliza para el método de diferencias finitas por diferencias centradas para aproximar la primera derivada. En este caso el valor de $h$ que minimiza a los errores es del orden $h^* = 10^{-6}$.
# ## Diferenciación numérica para una función $f: \mathbb{R}^n \rightarrow \mathbb{R}$
# Supongamos $f$ es dos veces diferenciable en $\text{intdom}f$. Si $f: \mathbb{R}^n \rightarrow \mathbb{R}$ entonces $\nabla f: \mathbb{R}^n \rightarrow \mathbb{R}^n$ y $\nabla ^2f: \mathbb{R}^n \rightarrow \mathbb{R}^{n\times n}$ (ver {ref}`Definición de función, continuidad y derivada <FCD>` para definición de derivadas en funciones $f: \mathbb{R}^n \rightarrow \mathbb{R}^m$). Ambas funciones al evaluarse resultan en un vector en $\mathbb{R}^n$ y en una matriz en $\mathbb{R}^{n\times n}$ respectivamente.
# Podemos utilizar las fórmulas de aproximación en diferenciación numérica con diferencias finitas para el caso $f: \mathbb{R} \rightarrow \mathbb{R}$ revisadas anteriormente para aproximar al gradiente y a la Hessiana.
#
# Para el caso del gradiente se tiene por **diferenciación hacia delante**:
#
# $$\nabla f(x) =
# \begin{array}{l}
# \left[ \begin{array}{c}
# \frac{\partial f(x)}{\partial x_1}\\
# \vdots\\
# \frac{\partial f(x)}{\partial x_n}
# \end{array}
# \right] \approx, \nabla \hat{f}(x) =\left[
# \begin{array}{c}
# \frac{f(x+he_1) - f(x)}{h}\\
# \vdots\\
# \frac{f(x+he_n) - f(x)}{h}
# \end{array}
# \right]
# \end{array} \in \mathbb{R}^n$$
#
# con $e_j$ $j$-ésimo vector canónico que tiene un número $1$ en la posición $j$ y $0$ en las entradas restantes para $j=1,\dots,n$. Se cumple $||\nabla f(x) - \nabla \hat{f}(x)|| = \mathcal{O}(h)$. Y para el caso de la Hessiana:
# $$\nabla^2f(x) = \left[\begin{array}{cccc}
# \frac{\partial^2f(x)}{\partial x_1^2} &\frac{\partial^2f(x)}{\partial x_2 \partial x_1}&\dots&\frac{\partial^2f(x)}{\partial x_n \partial x_1}\\
# \frac{\partial^2f(x)}{\partial x_1 \partial x_2} &\frac{\partial^2f(x)}{\partial x_2^2} &\dots&\frac{\partial^2f(x)}{\partial x_n \partial x_2}\\
# \vdots &\vdots& \ddots&\vdots\\
# \frac{\partial^2f(x)}{\partial x_1 \partial x_n} &\frac{\partial^2f(x)}{\partial x_2 \partial x_n}&\dots&\frac{\partial^2f(x)}{\partial x_n^2} \\
# \end{array}
# \right],
# $$
#
# $$
# \nabla^2 \hat{f}(x) = \left[\begin{array}{cccc}
# \frac{f(x+2he_1)-2f(x+he_1)+f(x)}{h^2} &\frac{f(x+he_1+he_2)-f(x+he_1)-f(x+he_2)+f(x)}{h^2}&\dots&\frac{f(x+he_1+he_n)-f(x+he_1)-f(x+he_n)+f(x)}{h^2}\\
# \frac{f(x+he_1+he_2)-f(x+he_2)-f(x+he_1)+f(x)}{h^2}&\frac{f(x+2he_2)-2f(x+he_2)+f(x)}{h^2} &\dots&\frac{f(x+he_2+he_n)-f(x+he_2)-f(x+he_n)+f(x)}{h^2}\\
# \vdots &\vdots& \ddots&\vdots\\
# \frac{f(x+he_1+he_n)-f(x+he_n)-f(x+he_1)+f(x)}{h^2}&\frac{f(x+he_2+he_n)-f(x+he_n)-f(x+he_2)+f(x)}{h^2}&\dots&\frac{f(x+2he_n)-2f(x+he_n)+f(x)}{h^2}\\
# \end{array}
# \right]
# $$
#
# Se cumple: $||\nabla^2f(x)-\nabla\hat{f}^2(x)||=\mathcal{O}(h)$.
# ### Ejemplo
#
# Aproximar $\nabla f(x), \nabla^2f(x)$ con diferencias hacia delante y $h \in \{10^{-16}, 10^{-14}, \dots , 10^{-1}\}$ para $f: \mathbb{R}^4 \rightarrow \mathbb{R}$, dada por $f(x) = (x_1^2-x_2^2)^2+x_1^2+(x_3^2-x_4^2)^2+x_3^2$ en el punto $x_0=(1.5,1.5,1.5,1.5)^T$. Realizar una gráfica de $\log($Err_rel) vs $\log($h)
#
# Para esta función se tiene:
#
# $$\nabla f(x) =
# \left[ \begin{array}{c}
# 4x_1(x_1^2-x_2^2) + 2x_1\\
# -4x_2(x_1^2-x_2^2)\\
# 4x_3(x_3^2-x_4^2)+2x_3\\
# -4x_4(x_3^2-x_4^2)
# \end{array}
# \right] ,
# $$
# $$\nabla^2f(x)=
# \left[\begin{array}{cccc}
# 12x_1^2-4x_2^2+2 &-8x_1x_2&0&0\\
# -8x_1x_2 &-4x_1^2+12x_2^2&0&0\\
# 0 &0&12x_3^2-4x_4^2+2&-8x_3x_4\\
# 0&0&-8x_3x_4&-4x_3^2+12x_4^2\\
# \end{array}
# \right]
# $$
# **Gradiente de f calculado de forma simbólica**
#
gf<-function(x){
c(4*x[1]*(x[1]^2-x[2]^2)+2*x[1],
-4*x[2]*(x[1]^2-x[2]^2),
4*x[3]*(x[3]^2-x[4]^2)+2*x[3],
-4*x[4]*(x[3]^2-x[4]^2))
}
# **Punto en el que se evaluará**
x_0<-c(1.5,1.5,1.5,1.5)
print(gf(x_0))
# $$\nabla f(x_0) =
# \left[ \begin{array}{c}
# 3\\
# 0\\
# 3\\
# 0
# \end{array}
# \right],
# $$
# **Hessiana de f calculada de forma simbólica**
#
gf2<-function(x){
matrix(c(12*x[1]^2-4*x[2]^2+2,-8*x[1]*x[2],0,0,
-8*x[1]*x[2],-4*x[1]^2+12*x[2]^2,0,0,
0,0,12*x[3]^2-4*x[4]^2+2,-8*x[3]*x[4],
0,0,-8*x[3]*x[4],-4*x[3]^2+12*x[4]^2),nrow=4,ncol=4)
}
# **Evaluación de la Hessiana**
print(gf2(x_0))
# $$\nabla^2f(x_0)=
# \left[\begin{array}{cccc}
# 20 &-18&0&0\\
# -18&18&0&0\\
# 0 &0&20&-18\\
# 0&0&-18&18\\
# \end{array}
# \right]
# $$
# **Definición de función y punto en el que se calculan las aproximaciones**
#
f <- function(x){
(x[1]^2-x[2]^2)^2+x[1]^2+(x[3]^2-x[4]^2)^2+x[3]^2
}
x0 <- rep(1.5,4)
# **Lo siguiente calcula el gradiente y la Hessiana de forma numérica con la aproximación por diferencias hacia delante**
inc_index<-function(vec,index,h){
'
Auxiliary function for gradient and Hessian computation.
Args:
vec (double): vector
index (int): index.
h (float): quantity that vec[index] will be increased.
Returns:
vec (double): vector with vec[index] increased by h.
'
vec[index]<-vec[index]+h
vec
}
gradient_approximation<-function(f,x,h=1e-8){
'
Numerical approximation of gradient for function f using forward differences.
Args:
f (expression): definition of function f.
x (double): vector that holds values where gradient will be computed.
h (float): step size for forward differences, tipically h=1e-8
Returns:
gf (array): numerical approximation to gradient of f.
'
n<-length(x)
gf<-vector("double",n)
for(i in 1:n){
gf[i]=(f(inc_index(x,i,h))-f(x))
}
gf/h
}
Hessian_approximation<-function(f,x,h=1e-6){
'
Numerical approximation of Hessian for function f using forward differences.
Args:
f (expression): definition of function f.
x (double): vector that holds values where Hessian will be computed.
h (float): step size for forward differences, tipically h=1e-6
Returns:
Hf (double): matrix of numerical approximation to Hessian of f.
'
n<-length(x)
Hf<-matrix(rep(0,n^2),nrow=n,ncol=n)
f_x<-f(x)
for(i in 1:n){
x_inc_in_i<-inc_index(x,i,h)
f_x_inc_in_i<-f(x_inc_in_i)
for(j in i:n){
dif<-f(inc_index(x_inc_in_i,j,h))-f_x_inc_in_i-f(inc_index(x,j,h))+f_x
Hf[i,j]<-dif
if(j!=i)
Hf[j,i]<-dif
}
}
Hf/h^2
}
# **Conjunto de valores de h para diferencias hacia delante**
h<-10^(-1*(1:16))
# **Funciones para cálculo del error:**
#
# $$\text{ErrRel(approx)} = \frac{||\text{approx}-\text{obj}||}{||\text{obj}||}$$
# +
Euclidian_norm<-function(vec){
'Compute Euclidian norm of vector'
sqrt(sum(vec*vec))
}
compute_error<-function(obj,approx){
'
Relative or absolute error between obj and approx based in Euclidian norm.
Approx is a numeric vector.
'
if (Euclidian_norm(obj) > .Machine$double.eps){
Err<-Euclidian_norm(obj-approx)/Euclidian_norm(obj)
}else
Err<-Euclidian_norm(obj-approx)
Err
}
# +
gf_numeric_approximations <- lapply(h,gradient_approximation,f=f,x=x0)
gf2_numeric_approximations <- lapply(h,Hessian_approximation,f=f,x=x0)
rel_err_gf <- sapply(gf_numeric_approximations,compute_error,obj=gf(x_0))
rel_err_gf2 <- sapply(gf2_numeric_approximations,compute_error,obj=gf2(x_0))
# -
gg<-ggplot()
print(gg+
geom_line(aes(x=log(h),y=log(rel_err_gf)))+
ggtitle('Aproximación al gradiente por diferencias finitas'))
print(h[which.min(rel_err_gf)])
print(gg+
geom_line(aes(x=log(h),y=log(rel_err_gf2)))+
ggtitle('Aproximación a la Hessiana por diferencias finitas'))
print(h[which.min(rel_err_gf2)])
# ```{admonition} Ejercicio
# :class: tip
#
# Aproximar $\nabla f(x), \nabla^2f(x)$ con diferencias hacia delante y $h \in \{10^{-16}, 10^{-14}, \dots , 10^{-1}\}$ para $f: \mathbb{R}^3 \rightarrow \mathbb{R}$, dada por $f(x) = x_1x_2\exp(x_1^2+x_3^2-5)$ en el punto $x_0=(1,3,-2)^T$ Realizar una gráfica de $\log($Err_rel) vs $\log($h).
# ```
# ```{admonition} Ejercicios
# :class: tip
#
# 1. Resuelve los ejercicios y preguntas de la nota.
# ```
# **Referencias**
#
# 1. <NAME>, <NAME>, Numerical Analysis, Brooks/Cole Cengage Learning, 2005.
#
# 2. <NAME>, Scientific Computing. An Introductory Survey, McGraw-Hill, 2002.
#
# 3. <NAME>, <NAME>, Convex Optimization. Cambridge University Press, 2004.
#
| libro_optimizacion/temas/I.computo_cientifico/1.6/Polinomios_de_Taylor_y_diferenciacion_numerica.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Riccati transformed inner problem
# This notebook will illustrate how to use solve the Riccati transformation of the inner problem to determine optimal shift $\ell$ and smoothing $\delta$ parameters for normalized mask functions $\Gamma$.
#
# Specifically, for a normalized mask function with support $-c < x < c$, we solve the Riccati equation
#
# $$ R' + R^2 = \delta^2 \Gamma,$$
#
# with initial condition
#
# $$ R(-c) = \delta,$$
#
# to find the optimal shift $\ell^*$ as
#
# $$\ell^*(\delta) = \left(\frac{1}{R(c)} - c\right) \delta.$$
#
# We will solve this problem numerically using python for a compact error function mask.
#
# It is possible to apply Newton iteration to the equation to efficiently determine the optimal smoothing that requires zero shift. This is done by differentiating the equations with respect to the parameter $\delta$.
# # Imports
import numpy as np
import scipy.integrate as spint
from scipy.special import erf
import matplotlib.pyplot as plt
from matplotlib import rc
rc('font',**{'family':'serif','serif':['Computer Modern Roman']})
rc('text', usetex=True)
# # Optimal shift
def Γ(z):
"""Define the normalized mask function you want."""
return 0.5*(1-erf(np.sqrt(np.pi)*z/np.sqrt(1-z**2)))
def Xt(X, t, δ, Γ):
"""The derivative for the coupled Riccati equations."""
return np.array([-X[0]**2 + δ**2*Γ(t),
-2*X[1]*X[0] + 2*δ*Γ(t)])
def solve_riccati(δ,Γ,ts=[-1,1]):
"""Solve the Riccati equation with mask K and damping scaling κ."""
X0 = np.array([δ,1.])
Xs = spint.odeint(Xt, X0, ts, tcrit=[-1,1],args=(δ,Γ))
return Xs
def shift(δ,Γ,ϵ=1e-16):
"""Calculate required shift given mask K and damping scaling κ."""
R1, dR1 = solve_riccati(δ,Γ)[-1,:]
return 1/R1 - 1, -dR1/R1**2
shift(3.14,Γ)
def ideal_shift(Γ,δ0,tol=1e-10,maxits=100):
"""Use Newton iteration to determine zero-shift smoothing δ."""
δ,dδ = np.array([δ0]), np.array([1.])
diff, its = 1, 0
while diff > tol and its < maxits:
li, dli = shift(δ[-1],Γ)
diff, its = - li/dli, its+1
δ, dδ = np.append(δ,δ[-1]+diff), np.append(dδ,diff)
return δ, dδ
ideal_shift(Γ,1)
| riccati.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Black Scholes Exercise 6: MPI implementation
# Use MPI to parallelize and distribute the work
# +
# Boilerplate for the example
import cProfile
import pstats
import numpy as np
try:
import numpy.random_intel as rnd
except:
import numpy.random as rnd
# make xrange available in python 3
try:
xrange
except NameError:
xrange = range
SEED = 7777777
S0L = 10.0
S0H = 50.0
XL = 10.0
XH = 50.0
TL = 1.0
TH = 2.0
RISK_FREE = 0.1
VOLATILITY = 0.2
TEST_ARRAY_LENGTH = 1024
###############################################
def gen_data(nopt):
return (
rnd.uniform(S0L, S0H, nopt),
rnd.uniform(XL, XH, nopt),
rnd.uniform(TL, TH, nopt),
)
nopt=100000
price, strike, t = gen_data(nopt)
call = np.zeros(nopt, dtype=np.float64)
put = -np.ones(nopt, dtype=np.float64)
# +
# Let's add a Black Scholes kernel (numpy) and an MPI wrapper
from numpy import log, invsqrt, exp, erf
# Black Scholes kernel
def black_scholes(nopt, price, strike, t, rate, vol):
mr = -rate
sig_sig_two = vol * vol * 2
P = price
S = strike
T = t
a = log(P / S)
b = T * mr
z = T * sig_sig_two
c = 0.25 * z
y = invsqrt(z)
w1 = (a - b + c) * y
w2 = (a - b - c) * y
d1 = 0.5 + 0.5 * erf(w1)
d2 = 0.5 + 0.5 * erf(w2)
Se = exp(b) * S
call = P * d1 - Se * d2
put = call - P + Se
return (call, put)
# MPI wrapper
from mpi4py import MPI
def black_scholes_mpi(nopt, price, strike, t, rate, vol):
comm = MPI.COMM_WORLD
nump = comm.size
noptpp = int(nopt/nump)
myprice = np.empty(noptpp, dtype=np.float64)
mystrike = np.empty(noptpp, dtype=np.float64)
myt = np.empty(noptpp, dtype=np.float64)
# Scatter data into arrays
comm.Scatter(price, myprice, root=0)
# TODO scatter strike and t
mycall, myput = black_scholes(noptpp, myprice, mystrike, myt, rate, vol)
comm.Gather(mycall, call)
#TODO gather put
return call, put
# -
# Running a single python function in MPI is possible with ipython but can be tricky to setup.
# Let's run the entire program with MPI.
# We need to add our own timing.
from time import clock
t1 = clock()
black_scholes_mpi(nopt, price, strike, t, RISK_FREE, VOLATILITY)
t2 = (clock()-t1)*1000
print("Time: {:.2f}ms".format(t2))
# Now we can safe the code to a file and run it with mpirun
# %save -f runme.py 1-3
# !mpirun -n 4 python ./runme.py
# ## 1. Let only rank 0 print the timing
# ## 2. How can we reduce the communication cost?
| 6_BlackScholes_mpi.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:lab]
# language: python
# name: conda-env-lab-py
# ---
# # Advanced NumPy usage
# +
from random import randint
gold = [randint(0, 100) for i in range(10000000)]
# -
# %%timeit
silver = []
for gold_value in gold:
silver.append(12 * gold_value)
# %timeit silver = [12 * gold_value for gold_value in gold]
import numpy as np
gold = np.random.randint(0, 100, (10000000,))
# %%timeit
silver = np.empty(gold.shape)
for i in range(gold.shape[0]):
silver[i] = 12 * gold[i]
# %timeit silver = 12 * gold
loots = np.random.randint(0, 100, (10,))
shares = np.array([.7, 1., 1.3, 2., 1.])
loots * shares
gold = np.random.randint(0, 100, (10,))
gold
ratio = np.array(12)
ratio_wide = ratio * np.ones(gold.shape)
ratio_wide
gold * ratio_wide
loots
loots.shape
loots[:, np.newaxis]
loots[:, np.newaxis].shape
loots[:, None]
np.newaxis is None
loots = np.random.randint(0, 100, (10,))
shares = np.array([.7, 1., 1.3, 2., 1.])
loots[:, np.newaxis] * shares
names = np.array(['bard', 'mage', 'leader', 'paladin', 'ranger'])
# character attributes = health, speed, strength, mana
attributes = np.array([[72, 15, 48, 22],
[68, 8, 44, 87],
[65, 12, 56, 28],
[82, 7, 78, 0],
[67, 11, 62, 14]])
attributes
sorter = attributes[:, 0].argsort()
sorter
attributes[sorter]
names[sorter]
contributions = np.array([.7, 1., 1.3, 2., 1.])
attributes.T * contributions
loots = np.random.randint(-100, 100, (10,))
loots
loots > 0
loots[loots > 0]
loots[np.logical_and(loots >= 5, loots <= 30)]
dungeon_difficulties = np.array([[ 7, 7, 5, 1, 5, 6, 9, 6, 10, 4, 2, 3],
[ 10, 16, 12, 12, 18, 0, 16, 20, 12, 4, 12, 12],
[ 3, 3, 12, 15, 6, 12, 27, 6, 3, 30, 15, 27],
[ 24, 12, 40, 32, 40, 12, 12, 16, 24, 4, 16, 4],
[ 25, 0, 35, 0, 40, 50, 40, 20, 35, 25, 25, 35],
[ 30, 24, 36, 48, 42, 6, 18, 54, 36, 54, 12, 24],
[ 49, 28, 42, 70, 21, 42, 7, 42, 49, 42, 0, 63],
[ 8, 16, 72, 16, 24, 40, 40, 40, 64, 24, 24, 8],
[ 81, 62, 18, 81, 81, 27, 27, 45, 18, 27, 90, 36],
[ 80, 90, 20, 100, 80, 90, 50, 80, 70, 90, 60, 40]])
dungeon_difficulties[[1, 4, 7], [0, 2, 3]]
dungeon_difficulties[np.ix_([1, 4, 7], [0, 2, 3])]
np.ones(dungeon_difficulties.shape).astype(int)
import numpy as np
objects_in_dungeons = np.array([[1, 1, 1, 2, 1, 3, 1, 5, 7, 1, 1, 9],
[1, 1, 1, 1, 2, 2, 2, 1, 1, 1, 1, 10],
[1, 1, 1, 5, 1, 1, 3, 1, 2, 1, 1, 8],
[1, 1, 1, 1, 10, 1, 1, 2, 2, 2, 1, 13],
[1, 1, 1, 2, 3, 3, 5, 8, 1, 2, 4, 10],
[2, 3, 2, 2, 1, 5, 6, 1, 9, 2, 7, 6],
[2, 2, 2, 1, 8, 2, 2, 1, 6, 1, 5, 12],
[3, 2, 3, 5, 1, 1, 4, 2, 1, 7, 1, 9],
[3, 3, 2, 2, 4, 3, 6, 2, 8, 1, 1, 17],
[2, 2, 2, 3, 3, 1, 8, 1, 1, 1, 1, 18]])
np.isin(objects_in_dungeons, 8).nonzero()
objects_in_dungeons[np.isin(objects_in_dungeons, np.arange(10, 20))]
dungeon_list = np.array([1, 4, 7])
floor_list = np.array([0, 2, 3])
np.isin(dungeon_difficulties, dungeon_list)
# %timeit dungeon_difficulties == 81
# %timeit np.isin(dungeon_difficulties, np.array([81]))
dungeon_difficulties[dungeon_difficulties]
N = 10000
large_dungeons = (np.arange(10, 110, N)[:, None] * np.round(np.random.rand(N, 12), decimals=1)).astype(int)
import numpy as np
loots_per_character_per_floor = np.random.randint(0, 100, (10, 5, 12))
loots_per_character_per_floor[0]
loots_per_character_per_floor[:, :, -1]
loots_per_character_per_floor[..., -1]
# # Indexing
import numpy as np
np.set_printoptions(suppress=True)
devises = np.array([20, 1.8, .5, 12])
loots = np.concatenate([np.random.randint(0, 4, (10,))[:, None],
np.random.randint(0, 100, (10,))[:, None]],
axis=1)
loots
loots[:, 1][:, None] * devises
(loots[:, 1][:, None] * devises)[np.arange(len(loots)), loots[:, 0]]
# # C-order vs F-order
a = np.random.rand(100, 1000000)
# %timeit a.sum(axis=0)
b = a.astype(dtype=a.dtype, order='F')
# %timeit b.sum(axis=0)
# # Einsum
# character attributes = health, speed, strength, mana
attributes = np.array([[72, 15, 48, 22],
[68, 8, 44, 87],
[65, 12, 56, 28],
[82, 7, 78, 1],
[67, 11, 62, 14]])
attributes.shape
loots = np.random.randint(0, 100, (3, 5))
loots
loots[:, :, None] / attributes
np.einsum('ij, jk -> ijk', loots, 1 / attributes)
# # View vs copy
a = np.arange(18).reshape(3, 6)
a
a[np.arange(3), [1, 2, 3]] = -1
a
a[np.mod(a, 3) == 0] = -1
a
np.arange(len(loots)), loots[:, 0]
import numpy as np
a = np.arange(27).reshape((9, 3))
a
# ## Under the hood
vectorization -> splitting loot part depending on character weight / contribution
# ## Vectorization
# The maze of indexing
normalization in one line
# ## Broadcasting
Crazy things with indexing
using weights / weighting samples
# ## Ufuncs
import numpy as np
# closest value from a reference list
ref_list = np.array([2, 5, 12])
def closest(a, ref):
return ref[(np.abs(ref - a)).argmin()]
x = np.random.randint(0, 20, size=(1000))
# %timeit np.apply_along_axis(closest, 1, x[:, None], ref_list)
closest_vect = np.vectorize(closest, excluded=['ref'])
# %timeit closest_vect(x, ref=ref_list)
def closest_numpy(arr, ref):
return ref[(np.abs(arr[:, None] - ref)).argmin(axis=1)]
ex = np.arange(5)
np.abs(ex[:, None] - ref_list)
# %timeit closest_numpy(x, ref=ref_list)
from numba import vectorize, int64
@vectorize([int64(int64)], nopython=True)
def closest_numba(a):
ref = ref_list
return ref[(np.abs(ref - a)).argmin()]
# %timeit closest_numba(x)
ex = np.arange(15)
closest_numba.at(ex, ex < 10)
ex
np.add.reduce(ex)
ex.sum()
# ## Numba and Numexpr
import numpy as np
np.set_printoptions(suppress=True)
# $ d_{euclid}(a, b) = \sqrt{||a-b||^2} = \sqrt{ (a_1 - b_1)^2 + (a_2 - b_2)^2} $
N = 1000
A = np.random.rand(N, 2) + 1
B = np.random.rand(N, 2) + 1
def euclid(a, b):
return np.sqrt(np.dot(a - b, (a - b).T))
def naive_euclid(X, Y):
res = np.empty((len(X), len(Y)))
for i in range(len(X)):
for j in range(len(Y)):
res[i, j] = euclid(X[i], Y[j])
return res
# %timeit naive_euclid(A, B)
def vectorized_euclid(X, Y):
diff = X[np.newaxis] - Y[:, np.newaxis]
return np.sqrt(np.einsum('ijk, ijk -> ji', diff, diff))
np.allclose(naive_euclid(A, B), vectorized_euclid(A, B))
# %timeit vectorized_euclid(A, B)
from sklearn.metrics import pairwise_distances
np.allclose(pairwise_distances(A, B), vectorized_euclid(A, B))
# %timeit pairwise_distances(A, B, metric='euclidean')
from numba import jit
@jit
def numba_euclid(X, Y):
res = np.empty((len(X), len(Y)))
for i in range(len(X)):
for j in range(len(Y)):
res[i, j] = np.sqrt(np.dot(X[i] - Y[j], X[i] - Y[j]))
return res
# %timeit numba_euclid(A, B)
@jit
def vectorized_euclid_numba(X, Y):
diff = X[np.newaxis] - Y[:, np.newaxis]
return np.sqrt(np.einsum('ijk, ijk -> ji', diff, diff))
# %timeit vectorized_euclid_numba(A, B)
# $d_{poinc}(a, b) = \mbox{arccosh} \left(1 + 2 \frac{||a - b ||^2}{(1 - ||a||^2) (1 - ||b|| ^2 )} \right)$
def poinc(a, b):
return np.arccosh(1+ 2* np.dot(a-b, a-b)/ ((1-np.dot(a,a))*(1-np.dot(b,b))))
def naive_poinc(X, Y):
res = np.empty((len(X), len(Y)))
for i in range(len(X)):
for j in range(len(Y)):
res[i, j] = poinc(X[i], Y[j])
return res
# %timeit naive_poinc(A, B)
def numpy_poinc(X, Y):
diff = X[np.newaxis] - Y[:, np.newaxis]
num = np.einsum('ijk, ijk -> ji', diff, diff)
den1 = 1 - np.einsum('ij, ji->i', X, X.T)
den2 = 1 - np.einsum('ij, ji->i', Y, Y.T)
den = den1[:, np.newaxis] * den2[np.newaxis]
return np.arccosh(1 + 2 * num / den)
# %timeit numpy_poinc(A, B)
@jit(nopython=True)
def numba_poinc(X, Y):
def poinc(a, b):
return np.arccosh(1+ 2* np.dot(a-b, a-b)/ ((1-np.dot(a,a))*(1-np.dot(b,b))))
res = np.empty((len(X), len(Y)))
i = 0
for i in range(len(X)):
for j in range(len(Y)):
res[i, j] = poinc(X[i], Y[j])
return res
# %timeit numba_poinc(A, B)
# # Strides
# # Windowed average
speed = np.random.rand(300000) * 75
def windowed_average(a, n):
av = np.empty(len(a) - n + 1)
for i in range(len(a) - n):
av[i] = a[i:i+n].sum() / n
return av
windowed_average(np.arange(30), 5)
# %timeit windowed_average(speed, 5)
# %timeit windowed_average_conv(speed, 5)
# %timeit windowed_average_cumsum(speed, 5)
def windowed_average_conv(a, n):
return np.convolve(a, np.ones((n,))/n, mode='valid')
windowed_average_conv(np.arange(30), 5)
def windowed_average_cumsum(a, n):
cumsum = np.cumsum(np.insert(a, 0, 0))
return (cumsum[n:] - cumsum[:-n]) / float(n)
windowed_average_cumsum(np.arange(30), 5)
# +
from numpy.lib.stride_tricks import as_strided
def windowed_median_strides(a, n):
stride = a.strides[0]
return np.median(as_strided(a, shape=[len(a) - n + 1, n], strides=[stride, stride]), axis=1)
# -
windowed_median_strides(np.arange(30), 5)
# %timeit windowed_median_strides(speed, 5)
from numba import guvectorize, float64, int64
@guvectorize(['void(float64[:], int64, float64[:])'], '(n),()->(n)', nopython=True)
def windowed_median(a, n, out):
for i in range(len(a) - n + 1):
out[i] = np.median(a[i: i + n])
windowed_median(np.arange(30), 5)
# %timeit windowed_median(speed, 5)
@guvectorize(['void(float64[:], int64, float64[:])'], '(n),()->(n)', nopython=True)
def windowed_average_numba(a, window_width, out):
asum = 0.0
count = 0
for i in range(window_width):
asum += a[i]
count += 1
out[i] = asum / count
for i in range(window_width, len(a)):
asum += a[i] - a[i - window_width]
out[i] = asum / count
windowed_average_numba(np.arange(30), 5)
# %timeit windowed_average_numba(speed, 5)
# ## Dask
| NumPy/code_samples.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.1.0
# language: julia
# name: julia-1.1
# ---
# # Gaussian mixture models
# Gaussian mixture models are popular for clustering data.
# Generally speaking, they are continuous random variables with a special probability density, namely
# $$
# \rho(x) = \sum_{i = 1}^{n} \frac{w_i}{\sqrt{2 \pi \sigma_i^2}} \exp \left( \frac{(x - \mu_i)^2}{2 \sigma_i^2} \right) \quad \text{with} \quad \sum_{i = 1}^n w_i = 1,
# $$
# where the pairs of means and standard deviations $(\mu_i, \sigma_i)$, and the weights $w_i$ for all $i \in \{ 1, \dots, n \}$ are given.
# Let's consider a simple example.
using Plots, LaTeXStrings
function f(x,μ,σ)
1/sqrt(2 *π*σ^2) * exp(-(x - μ)^2 / (2σ^2))
end
μ, σ = [1., 1.7], [0.2, 0.3]
ρ(x) = 0.5*f(x,μ[1],σ[1]) + 0.5*f(x,μ[2],σ[2])
x = 0:0.01:3
plot(x,ρ.(x))
xlabel!(L"x")
ylabel!(L"\rho(x)")
# This looks nice!
#
# What are now the polynomials that are orthogonal relative to this specific density?
using PolyChaos
deg = 4
meas = Measure("my_GaussMixture",ρ,(-Inf,Inf),false,Dict(:μ=>μ, σ=>σ)) # build measure
opq = OrthoPoly("my_op",deg,meas;Nquad = 100,Nrec = 2*deg) # construct orthogonal polynomial
showbasis(opq,digits=2) # in case you wondered
# Let's compute the square norms of the basis polynomials.
# opq = OrthoPolyQ(op) # add quadrature rule
T2 = Tensor(2,opq) # compute scalar products
T2num_1 = [ T2.get([i,j]) for i in 0:deg, j in 0:deg]
# This seems correct, but let's check against numerical integration.
using QuadGK
T2num_2 = [quadgk(x -> evaluate(i,x,opq)*evaluate(j,x,opq)*ρ(x),-Inf,Inf)[1] for i in 0:deg, j in 0:deg ]
T2num_1 - T2num_2
# Great!
# In case you run the code yourself, notice how much quicker `Tensor` is.
| examples/GaussianMixtureModel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
import sys
sys.version
# +
import pandas as pd
a3_path="/Users/mthipparthi/PycharmProjects/census/data/A-3_Vill/A-3 MDDS_Release.xls"
# Taking only first 8 columns as rest of them are mere derivatives of column 6,7,8
df = pd.read_excel(a3_path, header=1, parse_cols=8 )
# changing column names
df.columns = ['state_code','district_code', 'sub_district_code','level', 'area_name',
'inhabited_villages', 'rural_population', 'rural_male', 'rural_female']
# Removing invalid columns
df=df[df.area_name.notnull()]
df=df[df.level != 4]
df.head(8)
a3_clean_path="/Users/mthipparthi/PycharmProjects/census/data/cleaned_data/A-3_Vill/A-3 MDDS_Release_Clean.xls"
df.to_excel(a3_clean_path)
# +
import pandas as pd
b_DDW_0000B_02="/Users/mthipparthi/PycharmProjects/census/data/B-series/DDW-0000B-02.xlsx"
columns = ['table_name', 'state_code', 'district_code', 'type', 'area_name', 'religious_community', 'age_group',
'total_population', 'total_population_male', 'total_population_female',
'total_main_workers','total_main_workers_male', 'total_main_workers_female',
'total_marginal_workers','total_marginal_workers_male', 'total_marginal_workers_female',
'total_marginal_workers_seeking','total_marginal_workers_male_seeking', 'total_marginal_workers_female_seeking',
'total_non_workers','total_non_workers_male', 'total_non_workers_female',
'total_non_workers_seeking','total_non_workers_male_seeking', 'total_non_workers_female_seeking',
]
df = pd.read_excel(b_DDW_0000B_02, header=1)
# print(df.head(8))
df.columns = columns
df = df[df['table_name'].notnull()]
b_DDW_0000B_02_clean_path="/Users/mthipparthi/PycharmProjects/census/data/cleaned_data/B-series/DDW-0000B-02.xlsx"
df.to_excel(b_DDW_0000B_02_clean_path)
# +
import pandas as pd
b_DDW_0000B_12="/Users/mthipparthi/PycharmProjects/census/data/B-series/DDW-0000B-12.xlsx"
columns = [
'table_name', 'state_code', 'district_code', 'type', 'area_name', 'religious_community', 'age_group',
'total_marginal_workers','total_marginal_workers_male', 'total_marginal_workers_female',
'total_students','total_students_male', 'total_students_female',
'total_household_duties', 'total_household_duties_male', 'total_household_duties_female',
'total_dependents','total_dependents_male', 'total_dependents_female',
'total_pensioners','total_pensioners_male', 'total_pensioners_female',
'total_rentiers','total_rentiers_male', 'total_rentiers_female',
'total_beggers','total_beggers_male', 'total_beggers_female',
'total_others','total_others_male', 'total_others_female',
]
df = pd.read_excel(b_DDW_0000B_12, header=1)
df.columns = columns
df = df[df.table_name.notnull() & df.table_name.str.startswith('B1012')]
b_DDW_0000B_12_clean_path="/Users/mthipparthi/PycharmProjects/census/data/cleaned_data/B-series/DDW-0000B-12.xlsx"
df.to_excel(b_DDW_0000B_12_clean_path)
# +
b_DDW_0000B_14="/Users/mthipparthi/PycharmProjects/census/data/B-series/DDW-0000B-14.xlsx"
columns = [
'table_name', 'state_code', 'district_code', 'type', 'area_name', 'religious_community',
'total_non_workers','total_non_workers_male', 'total_non_workers_female',
'total_students','total_students_male', 'total_students_female',
'total_household_duties', 'total_household_duties_male', 'total_household_duties_female',
'total_dependents','total_dependents_male', 'total_dependents_female',
'total_pensioners','total_pensioners_male', 'total_pensioners_female',
'total_rentiers','total_rentiers_male', 'total_rentiers_female',
'total_beggers','total_beggers_male', 'total_beggers_female',
'total_others','total_others_male', 'total_others_female',
]
df = pd.read_excel(b_DDW_0000B_14, header=1)
df.columns = columns
df = df[df.table_name.notnull() & df.table_name.str.startswith('B1614')]
b_DDW_0000B_14_clean_path="/Users/mthipparthi/PycharmProjects/census/data/cleaned_data/B-series/DDW-0000B-14.xlsx"
df.to_excel(b_DDW_0000B_14_clean_path)
# +
import pandas as pd
b_DDWCITY_B_03="/Users/mthipparthi/PycharmProjects/census/data/B-series/DDWCITY-B-03-0000.xlsx"
columns = ['table_name', 'state_code', 'town_code', 'type', 'area_name', 'educational_level',
'total_population', 'total_population_male', 'total_population_female',
'total_main_workers','total_main_workers_male', 'total_main_workers_female',
'total_marginal_workers','total_marginal_workers_male', 'total_marginal_workers_female',
'total_marginal_workers_seeking','total_marginal_workers_male_seeking', 'total_marginal_workers_female_seeking',
'total_non_workers','total_non_workers_male', 'total_non_workers_female',
'total_non_workers_seeking','total_non_workers_male_seeking', 'total_non_workers_female_seeking',
]
df = pd.read_excel(b_DDWCITY_B_03, header=1)
# print(df.head(8))
df.columns = columns
df = df[df['table_name'].notnull() & df.table_name.str.startswith('B0103')]
b_DDWCITY_B_03_clean_path="/Users/mthipparthi/PycharmProjects/census/data/cleaned_data/B-series/DDWCITY-B-03-0000.xlsx"
df.to_excel(b_DDWCITY_B_03_clean_path)
# +
import pandas as pd
b_DDWCITY_B_09="/Users/mthipparthi/PycharmProjects/census/data/B-series/DDWCITY-B-09-0000.xlsx"
columns = ['table_name', 'state_code', 'town_code', 'type', 'area_name', 'educational_level',
'total_main_workers','total_main_workers_male', 'total_main_workers_female',
'total_main_workers_5_14','total_main_workers_5_14_male', 'total_main_workers_5_14_female',
'total_main_workers_15_19','total_main_workers_15_19_male', 'total_main_workers_15_19_female',
'total_main_workers_20_24','total_main_workers_20_24_male', 'total_main_workers_20_24_female',
'total_main_workers_25_29','total_main_workers_25_29_male', 'total_main_workers_25_29_female',
'total_main_workers_30_34','total_main_workers_30_34_male', 'total_main_workers_30_34_female',
'total_main_workers_35_39','total_main_workers_35_39_male', 'total_main_workers_35_39_female',
'total_main_workers_40_49','total_main_workers_40_49_male', 'total_main_workers_40_49_female',
'total_main_workers_50_59','total_main_workers_50_59_male', 'total_main_workers_50_59_female',
'total_main_workers_60_69','total_main_workers_60_69_male', 'total_main_workers_60_69_female',
'total_main_workers_70_plus','total_main_workers_70_plus_male', 'total_main_workers_70_plus_female',
'total_main_workers_age_not_stated','total_main_workers_age_not_stated_male', 'total_main_workers_age_not_stated_female',
]
df = pd.read_excel(b_DDWCITY_B_09, header=1)
# print(df.head(8))
df.columns = columns
df = df[df['table_name'].notnull() & df.table_name.str.startswith('B0409')]
b_DDWCITY_B_09_clean_path="/Users/mthipparthi/PycharmProjects/census/data/cleaned_data/B-series/DDWCITY-B-09-0000.xlsx"
df.to_excel(b_DDWCITY_B_09_clean_path)
# +
import pandas as pd
b_DDWCT_0000B_01="/Users/mthipparthi/PycharmProjects/census/data/B-series/DDWCT-0000B-01.xls"
columns = [
'table_name', 'state_code', 'town_code', 'town_name', 'type', 'age_group',
'total_population', 'total_population_male', 'total_population_female',
'total_main_workers','total_main_workers_male', 'total_main_workers_female',
'total_marginal_less_than_3_months','total_marginal_workers_less_than_3_months_male', 'total_marginal_workers_less_than_3_months_female',
'total_marginal_workers_3_to_6_months','total_marginal_workers_3_to_6_months_male', 'total_marginal_workers_3_to_6_months_female',
'total_marginal_workers_seeking','total_marginal_workers_male_seeking', 'total_marginal_workers_female_seeking',
'total_non_workers','total_non_workers_male', 'total_non_workers_female',
'total_non_workers_seeking','total_non_workers_male_seeking', 'total_non_workers_female_seeking',
]
df = pd.read_excel(b_DDWCT_0000B_01, header=1)
df.columns = columns
df = df[df.table_name.notnull() & df.table_name.str.startswith('B0101')]
b_b_DDWCT_0000B_01_clean_path="/Users/mthipparthi/PycharmProjects/census/data/cleaned_data/B-series/DDWCT-0000B-01.xls"
df.to_excel(b_b_DDWCT_0000B_01_clean_path)
# -
| data/Jupyter-Note Books/census_clean.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="1Ki5TPzdk-Jo" outputId="83f37003-272b-4db6-d915-34a8e14dd0b5"
import pandas as pd
import numpy as np
from tqdm import tqdm
from keras.preprocessing.text import Tokenizer
tqdm.pandas(desc="progress-bar")
from gensim.models import Doc2Vec
from sklearn import utils
from sklearn.model_selection import train_test_split
from keras.preprocessing.sequence import pad_sequences
import gensim
from sklearn.linear_model import LogisticRegression
from gensim.models.doc2vec import TaggedDocument
import re
import seaborn as sns
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/", "height": 205} id="lsnuTBIpjhDW" outputId="66df34e9-dc85-4eb6-c5db-d8b4eb5b4397"
df = pd.read_csv('all-data.csv',delimiter=',',encoding='latin-1')
df.head()
# + id="DGQ9CU2ak2Cm"
df = df.rename(columns={'neutral':'sentiment','According to Gran , the company has no plans to move all production to Russia , although that is where the company is growing .':'Message'})
# + colab={"base_uri": "https://localhost:8080/", "height": 205} id="MHf3D2susEK1" outputId="798679a3-e816-4218-ed1b-cdea4742b3a8"
# + colab={"base_uri": "https://localhost:8080/"} id="XPKGxPZos4H1" outputId="aadb2974-e2c6-4cc6-fa7d-79abe1b65e74"
df.index = range(4845)
df['Message'].apply(lambda x: len(x.split(' '))).sum()
# + [markdown] id="-mcRJDMHtI_z"
# **Lets visualize the Data**
# + colab={"base_uri": "https://localhost:8080/"} id="zdD68wWAwH2L" outputId="23380a74-0db3-47a0-ff5a-dc21984ff9a5"
sentiment_counts
# + colab={"base_uri": "https://localhost:8080/", "height": 341} id="ulEk5kbOtCMr" outputId="8d433059-b17e-4317-8311-ce8e23d2b279"
cnt_pro = df['sentiment'].value_counts()
sentiment_counts= cnt_pro.values
plt.figure(figsize=(12,4))
sns.barplot(cnt_pro.index, cnt_pro.values, alpha=0.8)
plt.ylabel('Number of Occurrences', fontsize=12)
plt.xlabel('sentiment', fontsize=12)
for i, v in enumerate(sentiment_counts):
plt.text( i, v ,s= str(v), fontweight= 'heavy')
plt.show();
# + colab={"base_uri": "https://localhost:8080/"} id="_nZVZC2etF9D" outputId="f78519b1-5a55-452d-e335-a92d88ca422d"
#Convert sting to numeric
sentiment = {'positive': 0,'neutral': 1,'negative':2}
df.sentiment = [sentiment[item] for item in df.sentiment]
print(df)
# + colab={"base_uri": "https://localhost:8080/"} id="Y217B5Zeyj3O" outputId="2e2a6c63-e34c-4e67-9119-0fe2746143b8"
def print_message(index):
example = df[df.index == index][['Message', 'sentiment']].values[0]
if len(example) > 0:
print(example[0])
print('Message:', example[1])
print_message(0)
# + [markdown] id="t9kiljPJzaPS"
# Now we convert text into lower case and remove punctuations
# + id="KcqY5FCby-yk"
from bs4 import BeautifulSoup
def cleanText(text):
text = BeautifulSoup(text, "lxml").text
text = re.sub(r'\|\|\|', r' ', text)
text = re.sub(r'http\S+', r'<URL>', text)
text = text.lower()
text = text.replace('x', '')
return text
df['Message'] = df['Message'].apply(cleanText)
# + [markdown] id="y-qFBTPb3bft"
# # Before Model building lets explore basic things about data!!
# + colab={"base_uri": "https://localhost:8080/"} id="m0osnwdRzors" outputId="cd378832-a05c-4683-f62d-26d07714b150"
import nltk
from nltk import sent_tokenize
from nltk import word_tokenize
nltk.download('punkt')
# + id="DNfohilp1Dtc"
text = df.Message
text = text.tolist()
# + colab={"base_uri": "https://localhost:8080/"} id="XSUd61170BNc" outputId="8e20a8ff-d3db-4ebd-e989-484d5319473e"
sentences= []
words= []
for i in range(100):
line = word_tokenize(text[i])
for i in line:
words.append(i)
print(len(words))
# + colab={"base_uri": "https://localhost:8080/"} id="9Y1SkMcn0Mq8" outputId="9fcb17cd-7124-4989-a437-556ce7ea0e49"
from nltk.probability import FreqDist
fdist= FreqDist(words)
fdist.most_common(10)
# + colab={"base_uri": "https://localhost:8080/"} id="cV3Y7YbQ21BA" outputId="a02d3710-2952-456b-cc9a-69a44eae3d0b"
#Lets remove some more symbols and puntuations
words_no_punc= []
for w in words:
if w.isalpha():
words_no_punc.append(w.lower())
print(words_no_punc)
print(len(words_no_punc))
# + colab={"base_uri": "https://localhost:8080/", "height": 495} id="Sfm8KEPY3t31" outputId="826e18c3-dc2f-416d-eb58-a93b4edd8c6d"
fdist= FreqDist(words_no_punc)
fdist.most_common(10), fdist.plot(10)
# + [markdown] id="mK5Yyc1j38Bd"
# **Lets remove the stopwords and then see the most frequent words**
# + colab={"base_uri": "https://localhost:8080/"} id="9s3HUd_F4NwV" outputId="cae180fa-0f05-456d-d821-aae4b0f8f5d5"
nltk.download('stopwords')
# + colab={"base_uri": "https://localhost:8080/"} id="RaMviHEx359e" outputId="607cee50-0e07-4f2b-a356-8263c9d38533"
from nltk.corpus import stopwords
stopwords= stopwords.words('english')
clean_words= []
for w in words_no_punc:
if w not in stopwords:
clean_words.append(w)
print('Now we see the length of words has become: ', len(clean_words))
# + colab={"base_uri": "https://localhost:8080/", "height": 324} id="f-sMDC_G4KsN" outputId="0ec753fb-3d95-45a9-dce3-a59abc9c3078"
fdist= FreqDist(clean_words)
fdist.plot(20)
# + [markdown] id="h-AwKa5-4-yN"
# Lets make some word cloud
# + id="lr_ZChtU5Zv3"
MyFile= open('Reviews.txt','w')
for element in text:
MyFile.write(element)
MyFile.write('\n')
MyFile.close()
# + id="qpN13Lbo6CrX"
text_file = open("Reviews.txt")
text = text_file.read()
# + colab={"base_uri": "https://localhost:8080/", "height": 366} id="13ndRw4_4mGm" outputId="3cd80ec5-24b5-4aaa-d917-764ff605d373"
from wordcloud import WordCloud
wordcloud= WordCloud().generate(text)
plt.figure(figsize = (12, 12))
plt.imshow(wordcloud)
plt.axis("off")
plt.show()
# + [markdown] id="0GFBBqNxPl49"
# Tokenization
# + colab={"base_uri": "https://localhost:8080/"} id="r1No3vM_5DEN" outputId="6d11045a-2eef-4cc2-f1a1-2602254aaa36"
train, test = train_test_split(df, test_size=0.000001 , random_state=42)
import nltk
from nltk.corpus import stopwords
def tokenize_text(text):
tokens = []
for sent in nltk.sent_tokenize(text):
for word in nltk.word_tokenize(sent):
if len(word) <= 0:
continue
tokens.append(word.lower())
return tokens
train_tagged = train.apply(lambda r: TaggedDocument(words=tokenize_text(r['Message']), tags=[r.sentiment]), axis=1)
test_tagged = test.apply(lambda r: TaggedDocument(words=tokenize_text(r['Message']), tags=[r.sentiment]), axis=1)
# The maximum number of words to be used. (most frequent)
max_fatures = 500000
# Max number of words in each complaint.
MAX_SEQUENCE_LENGTH = 50
#tokenizer = Tokenizer(num_words=max_fatures, split=' ')
tokenizer = Tokenizer(num_words=max_fatures, split=' ', filters='!"#$%&()*+,-./:;<=>?@[\]^_`{|}~', lower=True)
tokenizer.fit_on_texts(df['Message'].values)
X = tokenizer.texts_to_sequences(df['Message'].values)
X = pad_sequences(X ,maxlen= MAX_SEQUENCE_LENGTH)
print('Found %s unique tokens.' % len(X))
print('Shape of data tensor:', X.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="8DHep_DY8hWU" outputId="7a5bd36d-1b08-4d6a-bdad-c2a649ee1282"
train_tagged.values
# + [markdown] id="lRETN8M6Pwd0"
# # Doc2Vec :)
# + colab={"base_uri": "https://localhost:8080/"} id="COROsqCTB5qN" outputId="16618e92-7443-4197-f631-6bae6f58a1d7"
d2v_model = Doc2Vec(dm=1, dm_mean=1, size=20, window=8, min_count=1, workers=1, alpha=0.065, min_alpha=0.065)
d2v_model.build_vocab([x for x in tqdm(train_tagged.values)])
# + colab={"base_uri": "https://localhost:8080/"} id="x4XcK0JdCx4K" outputId="6b0e85fe-5c4e-45c5-c6c5-a63624897546"
# %%time
for epoch in range(30):
d2v_model.train(utils.shuffle([x for x in tqdm(train_tagged.values)]), total_examples=len(train_tagged.values), epochs=1)
d2v_model.alpha -= 0.002
d2v_model.min_alpha = d2v_model.alpha
# + colab={"base_uri": "https://localhost:8080/"} id="RYrMjz4pIGQx" outputId="eecd2425-988d-4bce-8143-c239bb502669"
print(d2v_model)
# + id="zAPKojW3IOdb"
# save the vectors in a new matrix
embedding_matrix = np.zeros((len(d2v_model.wv.vocab)+ 1, 20))
for i, vec in enumerate(d2v_model.docvecs.vectors_docs):
while i in vec <= 1000:
embedding_matrix[i]=vec
# + [markdown] id="xiHmf-sdIt4U"
# Measuring the most Similar Vectors
# + colab={"base_uri": "https://localhost:8080/"} id="gUBquCOXIYrE" outputId="afea47ba-419c-48a0-8fe5-e7e3c065e1f4"
d2v_model.wv.most_similar(positive=['profit'], topn=10)
# + [markdown] id="sy6_CYdiKNxU"
# # Building LSTM model
# + colab={"base_uri": "https://localhost:8080/"} id="3tBvx78QJpXD" outputId="b82dd8a4-9204-433a-c135-a15434a37b64"
from keras.models import Sequential
from keras.layers import LSTM, Dense, Embedding
model= Sequential()
#embed word vectors
model.add(Embedding(len(d2v_model.wv.vocab)+ 1,20 , input_length= X.shape[1], weights= [embedding_matrix], trainable= True))
def split_input(sequence):
return sequence[:-1], tf.reshape(sequence[1:], (-1,1))
model.add(LSTM(50,return_sequences=False))
model.add(Dense(3,activation="softmax"))
# output model skeleton
model.summary()
model.compile(optimizer="adam",loss="binary_crossentropy",metrics=['acc'])
# + colab={"base_uri": "https://localhost:8080/", "height": 369} id="AiwwRhX3Lqhf" outputId="4ccc0474-cff4-4a7a-ddca-c5456b0c70d0"
from keras.utils.vis_utils import plot_model
plot_model(model, to_file='model.png')
# + colab={"base_uri": "https://localhost:8080/"} id="fdkugqqrL3o8" outputId="85f0cc7d-120a-4910-ef7a-35f5baaad7c9"
Y = pd.get_dummies(df['sentiment']).values
X_train, X_test, Y_train, Y_test = train_test_split(X,Y, test_size = 0.15, random_state = 42)
print(X_train.shape,Y_train.shape)
print(X_test.shape,Y_test.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="qx4GzCaUMaRs" outputId="4c2df26e-7786-406c-f734-f58e81c5c5db"
batch_size = 32
history=model.fit(X_train, Y_train, epochs =50, batch_size=batch_size, verbose = 2)
# + colab={"base_uri": "https://localhost:8080/", "height": 573} id="ouYUfkTZMdOE" outputId="ca363d40-837a-44a5-a1c3-c0c385c72bfc"
plt.plot(history.history['acc'])
plt.title('model accuracy')
plt.ylabel('acc')
plt.xlabel('epochs')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
#plt.savefig('model_accuracy.png')
# summarize history for loss
plt.plot(history.history['loss'])
#plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epochs')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
#plt.savefig('model_loss.png')
# + colab={"base_uri": "https://localhost:8080/"} id="Ldj-nBQdNggf" outputId="18b004a6-d624-462a-828f-2eb8540fcf7b"
# evaluate the model
_, train_acc = model.evaluate(X_train, Y_train, verbose=2)
_, test_acc = model.evaluate(X_test, Y_test, verbose=2)
print('Train: %.3f, Test: %.4f' % (train_acc, test_acc))
# + colab={"base_uri": "https://localhost:8080/", "height": 406} id="iP_OpO1zNrXl" outputId="e469151e-d1b7-4df7-fc7a-bcd958e7e20b"
from sklearn.metrics import confusion_matrix
# predict probabilities for test set
yhat_probs = model.predict(X_test, verbose=0)
# predict classes for test set
yhat_classes = model.predict_classes(X_test, verbose=0)
rounded_labels=np.argmax(Y_test, axis=1)
lstm_val = confusion_matrix(rounded_labels, yhat_classes)
f, ax = plt.subplots(figsize=(5,5))
sns.heatmap(lstm_val, annot=True, linewidth=0.7, fmt='g', ax=ax, cmap="BuPu")
plt.title('LSTM Classification Confusion Matrix')
plt.xlabel('Y predict')
plt.ylabel('Y test (0= Neutral, 1= Positive, 2= Negative) ')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="zmewV8HWOV3F" outputId="19fb86a9-4d4a-4464-d8ee-7cc9319fea85"
validation_size = 610
X_validate = X_test[-validation_size:]
Y_validate = Y_test[-validation_size:]
X_test = X_test[:-validation_size]
Y_test = Y_test[:-validation_size]
score,acc = model.evaluate(X_test, Y_test, verbose = 1, batch_size = batch_size)
print("score: %.2f" % (score))
print("acc: %.2f" % (acc))
# + [markdown] id="iv36i_K4Q3yd"
# # Lets test it with different data
# + colab={"base_uri": "https://localhost:8080/"} id="AWV5jedHPHfY" outputId="c14b30f5-928a-43f2-8baf-8aafb8262497"
message = ['<NAME> has a responsibility and should not tweet to manipulate crypto market, says CoinDCX CEO']
seq = tokenizer.texts_to_sequences(message)
padded = pad_sequences(seq, maxlen=X.shape[1], dtype='int32', value=0)
pred = model.predict(padded)
labels = ['Neutral','Positive','Negative']
print(labels[np.argmax(pred)])
# + id="3Tdz3N4DSJzp"
| Sentiment_Analysis__for_Finanicial_News.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exploring Beat Frequencies
#
# This simple notebook will let you play with close frequencies and hear the beatings created by intermodulation. It's also a cute example of the interactivity you can achieve with notebooks.
# +
# standard bookkeeping
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
from IPython.display import Audio, display
# interactivity here:
from ipywidgets import interact, interactive, fixed, interact_manual
import ipywidgets as widgets
# -
plt.rcParams["figure.figsize"] = (14,4)
# Let's define a simple fuction that generates, plots and plays two sinusoids at the given frequencies:
def beat_freq(f1=220.0, f2=224.0):
# the clock of the system
LEN = 4 # seconds
Fs = 8000.0
n = np.arange(0, int(LEN * Fs))
s = np.cos(2*np.pi * f1/Fs * n) + np.cos(2*np.pi * f2/Fs * n)
# start from the first null of the beating frequency
K = int(Fs / (2 * abs(f2-f1)))
s = s[K:]
# play the sound
display(Audio(data=s, rate=Fs))
# display one second of audio
plt.plot(s[0:int(Fs)])
interact(beat_freq, f1=(200.0,300.0), f2=(200.0,300.0));
# +
# -
| FrequencyBeatings/FrequencyBeatings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import sys,os
from pathlib import Path
from lib_o_.rpaclss import *
sys.path.append(str(Path(os.getcwd()).parent.parent.absolute()))
zn = ['DHK_M','DHK_N','DHK_O','DHK_S','CTG_M','CTG_N','CTG_S','COM','NOA','BAR','KHL','KUS','MYM','RAJ','RANG','SYL']
def format_one(df = False):
if df == False:
df = pd.read_csv(os.getcwd() + "\\csv_o_\\sclick.csv")
print(df.shape[0])
xx = RPA(df)
xx.csvmap(os.getcwd() + "\\csv_o_\\vipsite.csv",'CUSTOMATTR15')
#xx.timecal('LASTOCCURRENCE')
#xx.timefmt('DUR','%H:%M:%S')
xx.add_cond('sZone','DHK_M')
xx.add_cond('CAT',['DL'])
xx.apply_cond(['Company'], condition='NA', operation='remove')
xx.msgitems(['CUSTOMATTR15','LASTOCCURRENCE','Company'])
return xx.rpagen()
format_one()
# -
| AOmPy_o_/test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# <img align="left" src="imgs/logo.jpg" width="50px" style="margin-right:10px">
# # Snorkel Workshop: Extracting Spouse Relations <br> from the News
# ## Part 4: Training our End Extraction Model
#
# In this final section of the tutorial, we'll use the noisy training labels we generated in the last tutorial part to train our end extraction model.
#
# For this tutorial, we will be training a fairly effective deep learning model. More generally, however, Snorkel plugs in with many ML libraries including [TensorFlow](https://www.tensorflow.org/), making it easy to use almost any state-of-the-art model as the end extractor!
# +
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
import os
import numpy as np
# Connect to the database backend and initalize a Snorkel session
from lib.init import *
from snorkel.annotations import load_marginals
from snorkel.models import candidate_subclass
Spouse = candidate_subclass('Spouse', ['person1', 'person2'])
# -
# ## I. Loading Candidates and Gold Labels
#
# +
from snorkel.annotations import load_gold_labels
train_cands = session.query(Spouse).filter(Spouse.split == 0).order_by(Spouse.id).all()
dev_cands = session.query(Spouse).filter(Spouse.split == 1).order_by(Spouse.id).all()
test_cands = session.query(Spouse).filter(Spouse.split == 2).order_by(Spouse.id).all()
L_gold_dev = load_gold_labels(session, annotator_name='gold', split=1, load_as_array=True, zero_one=True)
L_gold_test = load_gold_labels(session, annotator_name='gold', split=2, zero_one=True)
train_marginals = load_marginals(session, split=0)
# -
# ## II. Training a _Long Short-term Memory_ (LSTM) Neural Network
#
# [LSTMs](https://en.wikipedia.org/wiki/Long_short-term_memory) can acheive state-of-the-art performance on many text classification tasks. We'll train a simple LSTM model below.
#
# In deep learning, hyperparameter tuning is very important and computationally expensive step in training models. For purposes of this tutorial, we've pre-selected some settings so that you can train a model in under 10 minutes. Advanced users can look at our [Grid Search Tutorial](https://github.com/HazyResearch/snorkel/blob/master/tutorials/advanced/Hyperparameter_Search.ipynb) for more details on choosing these parameters.
#
# | Parameter | Definition |
# |---------------------|--------------------------------------------------------------------------------------------------------|
# | n_epochs | A single pass through all the data in your training set |
# | dim | Vector embedding (i.e., learned representation) dimension |
# | lr, | The learning rate by which we update model weights after,computing the gradient |
# | dropout | A neural network regularization techique [0.0 - 1.0] |
# | print_freq | Print updates every k epochs |
# | batch_size | Estimate the gradient using k samples. Larger batch sizes run faster, but may perform worse |
# | max_sentence_length | The max length of an input sequence. Setting this too large, can slow your training down substantially |
#
# ### Please Note !!!
# With the provided hyperparameters below, your model should train in about 9.5 minutes.
# +
from snorkel.learning.pytorch.rnn import LSTM
train_kwargs = {
'lr': 0.001,
'dim': 100,
'n_epochs': 10,
'dropout': 0.25,
'print_freq': 1,
'batch_size': 128,
'max_sentence_length': 100
}
lstm = LSTM(n_threads=1)
lstm.train(train_cands, train_marginals, X_dev=dev_cands, Y_dev=L_gold_dev, **train_kwargs)
# -
# Now, we get the precision, recall, and F1 score from the discriminative model:
p, r, f1 = lstm.score(test_cands, L_gold_test)
print("Prec: {0:.3f}, Recall: {1:.3f}, F1 Score: {2:.3f}".format(p, r, f1))
# We can also get the candidates returned in sets (true positives, false positives, true negatives, false negatives) as well as a more detailed score report:
tp, fp, tn, fn = lstm.error_analysis(session, test_cands, L_gold_test)
# Finally, let's save our model for later use.
lstm.save("spouse.lstm")
| tutorials/workshop/Workshop_4_Discriminative_Model_Training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:root] *
# language: python
# name: conda-root-py
# ---
# 
#
# +
# %matplotlib
import matplotlib.pyplot as plt
x = [1,2,3,4,5,6,7,8,9,10]
y = [1,2,1.5,3,5.4,6.3,7.2,7.9,8.7,10]
plt.scatter(x,y,label='scatter_plot',color='r',marker='*')
plt.xlabel('Age')
plt.ylabel('Num')
plt.title('The City')
plt.legend()
plt.show()
# +
import numpy as np
fig, ax = plt.subplots(2,2,figsize=(14,7))
month = [1,2,3,4,5,6]
A = [1,2,3,4,5,6]
B = [3,4,5,6,7,8]
ax[0,0].stackplot(month,A,B,labels=['1','2','3'])
ax[1,1].stackplot(month,A,B)
ax[0,0].legend(loc=10)
fig.show()
# +
x = np.linspace(0, 2, 100)
print(x)
# Note that even in the OO-style, we use `.pyplot.figure` to create the figure.
fig, ax = plt.subplots() # Create a figure and an axes.
ax.plot(x, x, label='linear') # Plot some data on the axes.
ax.plot(x, x**2, label='quadratic') # Plot more data on the axes...
ax.plot(x, x**3, label='cubic') # ... and some more.
ax.set_xlabel('x label') # Add an x-label to the axes.
ax.set_ylabel('y label') # Add a y-label to the axes.
ax.set_title("Simple Plot") # Add a title to the axes.
ax.legend() # Add a legend.
fig.show()
# -
| 007-数据可视化-绘制图表/Class_01_matplotlib_ex.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: bpt
# language: python
# name: myenv
# ---
# # Predicting Substance Dependence from Multi-Site Data
#
# This notebook explores an example using data from the ENIGMA Addiction Consortium. Within this notebook we will be trying to predict between participants with any drug dependence (alcohol, cocaine, etc...), vs. healthy controls. The data for this is sources from a number of individual studies from all around the world and with different scanners etc... making this a challenging problem with its own unique considerations. Structural FreeSurfer ROIs are used. The raw data cannot be made available due to data use agreements.
#
# The key idea explored in this notebook is a particular tricky problem introduced by case-only sites, which are subject's data from site's with only case's. This introduces a confound where you cannot easily tell if the classifier is learning to predict site or the dependence status of interest.
#
# Featured in this notebook as well are some helpful
# code snippets for converting from BPt versions earlier than BPt 2.0 to valid BPt 2.0+ code.
# +
import pandas as pd
import BPt as bp
import numpy as np
import matplotlib.pyplot as plt
from warnings import simplefilter
simplefilter("ignore", category=FutureWarning)
# -
# ### Loading / Preparing Data
# As a general tip it can be useful to wrap something like a series of steps that load multiple DataFrames and merge them into a function. That said, it is often useful when first writing the function to try it taking advantage of the interactive-ness of the jupyter-notebook.
def load_base_df():
'''Loads and merges a DataFrame from multiple raw files'''
na_vals = [' ', ' ', 'nan', 'NaN']
# Load first part of data
d1 = pd.read_excel('/home/sage/Downloads/e1.xlsx', na_values=na_vals)
d2 = pd.read_excel('/home/sage/Downloads/e2.xlsx', na_values=na_vals)
df = pd.concat([d1, d2])
df['Subject'] = df['Subject'].astype('str')
df.rename({'Subject': 'subject'}, axis=1, inplace=True)
df.set_index('subject', inplace=True)
# Load second part
df2 = pd.read_excel('/home/sage/Downloads/e3.xlsx', na_values=na_vals)
df2['Subject ID'] = df2['Subject ID'].astype('str')
df2.rename({'Subject ID': 'subject'}, axis=1, inplace=True)
df2.set_index('subject', inplace=True)
# Merge
data = df2.merge(df, on='subject', how='outer')
# Rename age and sex
data = data.rename({'Sex_y': 'Sex', 'Age_y': 'Age'}, axis=1)
# Remove subject name to obsficate
data.index = list(range(len(data.index)))
return data
df = load_base_df()
df.shape
# +
# Cast to dataset
data = bp.Dataset(df)
data.verbose = 1
# Drop non relevant columns
data.drop_cols_by_nan(threshold=.5, inplace=True)
data.drop_cols(scope='Dependent', inclusions='any drug', inplace=True)
data.drop_cols(exclusions=['Half', '30 days', 'Site ',
'Sex_', 'Age_', 'Primary Drug', 'ICV.'], inplace=True)
# Set binary vars as categorical
data.auto_detect_categorical(inplace=True)
data.to_binary(scope='category', inplace=True)
print('to binary cols:', data.get_cols('category'))
# Set target and drop any NaNs
data.set_role('Dependent any drug', 'target', inplace=True)
data.drop_nan_subjects('target', inplace=True)
# Save this set of vars under scope covars
data = data.add_scope(['ICV', 'Sex', 'Age', 'Education', 'Handedness'], 'covars')
print('scope covars = ', data.get_cols('covars'))
# Set site as non input
data = data.set_role('Site', 'non input')
data = data.ordinalize(scope='non input')
# Drop subjects with too many NaN's and big outliers
data.drop_subjects_by_nan(threshold=.5, scope='all', inplace=True)
data.filter_outliers_by_std(n_std=10, scope='float', inplace=True)
# -
# **Legacy Equivilent pre BPt 2.0 loading code**
#
# ```
# ML = BPt_ML('Enigma_Alc',
# log_dr = None,
# n_jobs = 8)
#
# ML.Set_Default_Load_Params(subject_id = 'subject',
# na_values = [' ', ' ', 'nan', 'NaN'],
# drop_na = .5)
#
# ML.Load_Data(df=df,
# drop_keys = ['Unnamed:', 'Site', 'Half', 'PI', 'Dependent',
# 'Surface Area', 'Thickness', 'ICV', 'Subcortical',
# 'Sex', 'Age', 'Primary Drug', 'Education', 'Handedness'],
# inclusion_keys=None,
# unique_val_warn=None,
# clear_existing=True)
#
# ML.Load_Targets(df=df,
# col_name = 'Dependent any drug',
# data_type = 'b')
#
# ML.Load_Covars(df=df,
# col_name = ['ICV', 'Sex', 'Age'],
# drop_na = False,
# data_type = ['f', 'b', 'f'])
#
# ML.Load_Covars(df = df,
# col_name = ['Education', 'Handedness'],
# data_type = ['f', 'b'],
# drop_na = False,
# filter_outlier_std = 10)
#
# ML.Load_Strat(df=df,
# col_name=['Sex', 'Site'],
# binary_col=[True, False]
# )
#
# ML.Prepare_All_Data()
# ```
# Let's take a look at what we prepared. We can see that visually the Dataset is grouped by role.
# Let's plot some variables of interest
data.plot('target')
data.plot('Site')
data.plot_bivar('Site', 'Dependent any drug')
# That bi-variate plot is a little hard to read... we can make it bigger though easy enough. Let's say we also wanted to save it (we need to add show=False)
plt.figure(figsize=(10, 10))
data.plot_bivar('Site', 'Dependent any drug', show=False)
plt.savefig('site_by_drug.png', dpi=200)
# Okay next, we are going to define a custom validation strategy to use, which is essentially going to preserve subjects within the same train and test fold based on site.
#
#
# **Legacy Code**
# ```
# from BPt import CV
# group_site = CV(groups='Site')
# ```
group_site = bp.CVStrategy(groups='Site')
group_site
# We then can use this CV strategy as an argument when defining the train test split.
#
# **Legacy Code**
# ```
# ML.Train_Test_Split(test_size =.2, cv=group_site)
# ```
data = data.set_test_split(size=.2, cv_strategy=group_site, random_state=5)
data.plot('target', subjects='train')
data.plot('target', subjects='test')
# ### Running ML
#
# Next, we are going to evaluate some different machine learning models on the problem we have defined. Notably not much has changed here with respect to the old version, except some cosmetic changes like Problem_Spec to ProblemSpec. Also we have a new `evaluate` function instead of calling Evaluate via the ML object (`ML.Evaluate`).
# This just holds some commonly used values
ps = bp.ProblemSpec(subjects='train',
scorer=['matthews', 'roc_auc', 'balanced_accuracy'],
n_jobs=16)
ps
# Define a ModelPipeline to use with imputation, scaling and an elastic net
pipe = bp.ModelPipeline(imputers=[bp.Imputer(obj='mean', scope='float'),
bp.Imputer(obj='median', scope='category')],
scalers=bp.Scaler('standard'),
model=bp.Model('elastic', params=1),
param_search=bp.ParamSearch(
search_type='DiscreteOnePlusOne', n_iter=64))
pipe.print_all()
# ### 3-fold CV random splits
#
# Let's start by evaluating this model is a fairly naive way, just 3 folds of CV.
results = bp.evaluate(pipeline=pipe, dataset=data, mute_warnings=True,
problem_spec=ps, cv=bp.CV(splits=3))
results
# Notably this is likely going to give us overly optimistic results. Let's look at just the Site's drawn from the first evaluation fold to get a feel for why.
data.plot_bivar('Site', 'Dependent any drug', subjects=results.train_subjects[0])
data.plot_bivar('Site', 'Dependent any drug', subjects=results.val_subjects[0])
# ### Preserving Groups by Site
#
# We can see that for example the 149 subjects from site 1 in the validation set had 269 subjects also from site 1 to potentially memorize site effects from! The confound is a site with only cases. One way we can account for this is through something we've already hinted at, using the group site cv from earlier. We can create a new CV object with this attribute.
site_cv = bp.CV(splits=3, cv_strategy=group_site)
site_cv
results = bp.evaluate(pipeline=pipe, dataset=data, mute_warnings=True,
problem_spec=ps, cv=site_cv)
results
# We can now make the same plots as before, confirming now that the sites are different.
data.plot_bivar('Site', 'Dependent any drug', subjects=results.train_subjects[0])
data.plot_bivar('Site', 'Dependent any drug', subjects=results.val_subjects[0])
# Another piece we might want to look at in this case is the weighted_mean_scores. That is, each of the splits has train and test sets of slightly different sizes, so how does the metric change if we weight it by the number of validation subjects in that fold
results.weighted_mean_scores
results = bp.evaluate(pipeline=pipe, dataset=data, mute_warnings=True,
problem_spec=ps, cv=3)
results
# ### Other options?
#
# What other options do we have to handle site with CV besides just the group splitting?
#
# 1. Explicit leave-out-site CV: Where every train set is composed of all sites but one, and the validation set is the left out site.
#
# 2. Use the train-only parameter in CV.
#
# 3. Only evaluate subjects from sites with both cases and controls.
#
# Let's try option 1 first.
# ### Leave-out-site CV
# +
# All we need to do for leave-out-site is specify that splits = Site
leave_out_site_cv = bp.CV(splits='Site')
# This will actually raise an error
try:
results = bp.evaluate(pipeline=pipe, dataset=data, mute_warnings=True,
problem_spec=ps, cv=leave_out_site_cv)
except Exception as exc:
print(exc)
# -
# Uh oh looks like this option actually failed. What happened? Well basically since we have sites with only one class, we are trying to compute metrics that require two classes. Let's try again this time just using scorers that can work in the case of only one class.
# +
results = bp.evaluate(pipeline=pipe, dataset=data, scorer='accuracy', mute_warnings=True,
problem_spec=ps, cv=leave_out_site_cv)
results.mean_scores, results.std_scores, results.weighted_mean_scores
# -
# The problem here is accuracy is likely a pretty bad metric, especially for the sites which have both cases and controls, but a skewed distribution. In general using the group preserving by Site is likely a better option.
# ### Train-only subjects
#
# One optional way of handling these subjects from sites with only cases or only controls is to just never evaluate them. To instead always treat them as train subjects. We can notably then either use random splits or group preserving by CV.
# +
def check_imbalanced(df):
as_int = df['Dependent any drug'].astype('int')
return np.sum(as_int) == len(as_int)
# Apply function to check if each site has all the same target variable
is_imbalanced = data.groupby('Site').apply(check_imbalanced)
# Note these are internal values, not the original ones
imbalanced_sites = np.flatnonzero(is_imbalanced)
# We can specify just these subjects using the ValueSubset wrapper
imbalanced_subjs = bp.ValueSubset('Site', values=imbalanced_sites, decode_values=False)
# +
# Now define our new version of the group site cv
site_cv_tr_only = bp.CV(splits=3, cv_strategy=group_site,
train_only_subjects=imbalanced_subjs)
# And evaluate
results = bp.evaluate(pipeline=pipe, dataset=data, mute_warnings=True,
problem_spec=ps, cv=site_cv_tr_only)
results
# -
# The take-away here is maybe a little bleak. Essentially it is saying that some of the high scores we got earlier might have been in large part due to scores from the imbalanced sites.
# +
# And we can also run a version with just random splits
tr_only_cv = bp.CV(splits=3, train_only_subjects=imbalanced_subjs)
# And evaluate
results = bp.evaluate(pipeline=pipe, dataset=data, mute_warnings=True,
problem_spec=ps, cv=tr_only_cv)
results
# -
# While this version with random splits does better, we are still notably ignoring the distribution of case to control in the sites with both cases and controls. Based on the group preserving results what it seems like is going on is that when subjects from the same site are split across validation folds, the classifier might be able to just memorize Site instead of the effect of interest. Lets use Site 29 as an example.
#
# Let's look at just fold 1 first.
# +
# The train and val subjects from fold 0
tr_subjs = results.train_subjects[0]
val_subjs = results.val_subjects[0]
# An object specifying just site 29's subjects
site29 = bp.ValueSubset('Site', 29, decode_values=True)
# Get the intersection w/ intersection obj
site_29_tr = bp.Intersection([tr_subjs, site29])
site_29_val = bp.Intersection([val_subjs, site29])
# Plot
data.plot_bivar('Site', 'Dependent any drug', subjects=bp.Intersection([tr_subjs, site29]))
data.plot_bivar('Site', 'Dependent any drug', subjects=bp.Intersection([val_subjs, site29]))
# Grab just the subjects as actual index
val_subjs = data.get_subjects(site_29_val)
# Get a dataframe with the predictions made for just fold0
preds_df_fold0 = results.get_preds_dfs()[0]
# Let's see predictions for just these validation subjects
preds_df_fold0.loc[val_subjs]
# -
# We can see that it has learned to just predict 0 for every subject, which because of the imbalance is right most of the time, but really doesn't tell us if it has learned to predict site or substance dependence.
# ### Drop imbalanced subjects
# The last option was just don't use the imbalanced subjects at all. We will do this by indexing only the balanced sites
# +
# Note the ~ before the previously defined is_imbalanced
balanced_sites = np.flatnonzero(~is_imbalanced)
# We can specify just these subjects using the ValueSubset wrapper
balanced_subjs = bp.ValueSubset('Site', values=balanced_sites, decode_values=False)
# +
# Evaluate with site preserving cv from earlier but now using just the balanced train subjects
train_just_balanced = bp.Intersection(['train', balanced_subjs])
results = bp.evaluate(pipeline=pipe, dataset=data, mute_warnings=True,
subjects=train_just_balanced,
problem_spec=ps, cv=site_cv)
results
# -
# This actually seems to show that the imbalanced site confound with doesn't just corrupt cross validated results, but also actively impedes the model learning the real relationship. That we are better off just not using subjects with from sites with only cases. Yikes.
#
# Likewise, we can do the random split strategy again too.
results = bp.evaluate(pipeline=pipe, dataset=data, mute_warnings=True,
subjects=train_just_balanced,
problem_spec=ps, cv=bp.CV(splits=3))
results
# Also, we could run it with a leave-out-site CV. Now we can use the normal metrics.
results = bp.evaluate(pipeline=pipe, dataset=data, mute_warnings=True,
subjects=train_just_balanced,
problem_spec=ps, cv=leave_out_site_cv)
results
# ### Results Summary
#
# To summarize, of all of the above different iterations. Only the results from the versions where subjects from case-only sites are treated as train-only or dropped and evaluated group-preserving by site are likely un-biased! The leave-out-site CV results should be okay too, but let's skip those for now.
#
# Train Only Scores:
# mean_scores = {'matthews': 0.024294039432900597, 'roc_auc': 0.5292721521624003, 'balanced_accuracy': 0.5116503104881627}
#
# Dropped Scores:
# mean_scores = {'matthews': 0.23079311975258734, 'roc_auc': 0.6494071534373412, 'balanced_accuracy': 0.6141868842892168}
#
# By these interm results we can with some confidence conclude dropping subjects is the way to go, unless we want to dig deeper...
# ### Disentangle Target and Site?
#
# Can information on dependance from case-only sites be some how disentangled from site?
#
# This actually leads to a follow up a question. Namely is there any way to extract usable information from case-only site subjects? This question I actually investigated more deeply in https://onlinelibrary.wiley.com/doi/full/10.1002/hbm.25248 The idea of train-only subjects is actually from this paper, though it still has some important differences then the way we formulated it.
#
# The key piece in the paper which we haven't tried to do yet is centered around the problem of the classifier learning site related information instead of dependence related information in the train-only setup. We essentially need a way of forcing the classifier to learn this. The way proposed in the paper proposes that maybe we can by restricting the features provided to the classifier, give it only features related to dependence and not those related to site. In order to do this, we need to add a feature selection step.
#
# There are different ways to compose the problem, but we will specify the feature selection step as a seperate layer of nesting, importantly one that is evaluated according to the same site_cv_tr_only that we will be evaluating with. Remember the goal here is to select features which help us learn to predict useful information from case only sites, i.e., with feature only related to Dependence and not site!
# +
# Same imputer and scaler
imputers = [bp.Imputer(obj='mean', scope='float'),
bp.Imputer(obj='median', scope='category')]
scaler = bp.Scaler('standard')
# This object will be used to set features as an array of binary hyper-parameters
feat_selector = bp.FeatSelector('selector', params=1)
# This param search is responsible for optimizing the selected features from feat_selector
param_search = bp.ParamSearch('RandomSearch',
n_iter=16,
cv=site_cv_tr_only)
# We create a nested elastic net model to optimize - no particular CV should be okay
random_search = bp.ParamSearch('RandomSearch', n_iter=16)
elastic_search = bp.Model('elastic', params=1,
param_search=random_search)
# Put it all together in a pipeline
fs_pipe = bp.Pipeline(imputers + [scaler, feat_selector, elastic_search],
param_search=param_search)
# -
# Note: This may take a while to evaluate as we now have two levels of parameters to optimize, e.g., for each choice
# of features, we need to train a nested elastic net.
results = bp.evaluate(pipeline=fs_pipe, dataset=data, mute_warnings=True,
problem_spec=ps, cv=site_cv_tr_only)
results
# Unfortunetly this doesn't seem to work. Notably we still arn't quite following the method from the paper, which performs a meta-analysis on the results from the feature search to find overlapping subsets of features. As of the time of writing this example, a simmilar strategy has not been able to be easily integrated into BPt.
#
# The above method still has a few problems, one being severe overfitting. Namely, because a number of subjects are reserved in train-only roles, the same small number of subjects are being used over and over in cross-validation.
#
# We can confirm this intutiton by comparing the best score achieved internally for each set of features w/ how they score in the outer cross validation.
-results.estimators[0].best_score_, results.scores['matthews'][0]
-results.estimators[1].best_score_, results.scores['matthews'][1]
-results.estimators[2].best_score_, results.scores['matthews'][2]
# These values are quite different to say the least. Part of the problem again is that such a small amount of data is being used to determine generalization. What if instead of a the 3-fold group preserving train only feature selection CV, we instead used a leave-out-site scheme on site with train only? I doubt it will help, but let's see...
# +
leave_out_site_tr_only_cv = bp.CV(splits='Site',
train_only_subjects=imbalanced_subjs)
fs_pipe.param_search = bp.ParamSearch(
'RandomSearch', n_iter=16,
cv=leave_out_site_tr_only_cv)
results = bp.evaluate(pipeline=fs_pipe, dataset=data, mute_warnings=True,
problem_spec=ps, cv=site_cv_tr_only)
results
# -
# This does a little better. But only slightly. What if we also used this leave-out site strategy for the outer loop? This would be to address the issue that the outer CV has the same problem as the inner, where the validation set is overly specific and small in some folds.
results = bp.evaluate(pipeline=fs_pipe,
dataset=data,
problem_spec=ps,
mute_warnings=True,
cv=leave_out_site_tr_only_cv,
eval_verbose=1)
results
# Seeing the results for each site seperately with the verbose option really makes some of the problem clear, namely that a number of small sites don't seem to generalize well. This is simmilar to what we saw in the case of using leave-out-site with case-only subjects dropped, where we had a huge std in roc auc between folds.
#
# Another potential problem with doing leave-out-site is that the way the underlying data was collected, different sites will have subjects with dependence to different substances. This can certainly cause issues, as say our classifier only learns to classify alcohol dependence (the strongest signal), then it will do poorly on sites for other substances. Which is good in one way, that it lets us know the classifier is not generalizing, but it doesn't solve the generalization problem.
#
# Anyways, that will end this example for now. Leaving the problem of how to best exploit information from case-only sites as an open question (though if interested, a more faithful reproduction of the method presented in https://onlinelibrary.wiley.com/doi/full/10.1002/hbm.25248 might prove to be fruitful. Specifically, in the end only using a few, maybe 3-4, features, which is notably quite different then what we are trying above, i.e., trying just 16 random sets of features).
| Examples/Full_Examples/Predicting Substance Dependence from Multi-Site Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import warnings
warnings.filterwarnings("ignore")
import torchaudio as ta
ta.set_audio_backend("sox_io")
import torch
from torch.utils.data import DataLoader
import torch.nn as nn
import torch.nn.functional as F
import torch.autograd.profiler as profiler
# import pytorch_lightning as pl
import numpy as np
import os
import IPython.display as ipd
import numpy as np
import math
import glob
from tqdm.auto import tqdm
from python_files.Noise_Reduction_Datagen_fp16 import Signal_Synthesis_DataGen
import warnings
warnings.filterwarnings("ignore")
import gc
# from numba import jit
# -
torch.__version__
# +
noise_dir = "./dataset/UrbanSound8K-Resampled/audio/"
signal_dir = "./dataset/cv-corpus-5.1-2020-06-22-Resampled/en/clips/"
signal_nums_save = "./dataset_loader_files/signal_paths_nums_save.npy"
num_noise_samples=100
num_signal_samples = 100
noise_save_path = "./dataset_loader_files/noise_paths_resampled_save.npy"
n_fft=1024
win_length=n_fft
hop_len=n_fft//4
create_specgram = False
perform_stft = False
default_sr = 16000
sec = 6
augment=True
device_datagen = "cpu"
signal_synthesis_dataset = Signal_Synthesis_DataGen(noise_dir, signal_dir, \
signal_nums_save=signal_nums_save, num_noise_samples=num_noise_samples, \
num_signal_samples=num_signal_samples, noise_path_save=noise_save_path, \
n_fft=n_fft, win_length=win_length, hop_len=hop_len, create_specgram=create_specgram, \
perform_stft=perform_stft, normalize=True, default_sr=default_sr, sec=sec, epsilon=1e-5, augment=False, device=device_datagen)
# +
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
BATCH_SIZE = 4
shuffle = True
num_workers = 0
pin_memory = False
# data_loader = DataLoader(signal_synthesis_dataset, batch_size=BATCH_SIZE, shuffle=shuffle, num_workers=num_workers)
data_loader = DataLoader(signal_synthesis_dataset, batch_size=BATCH_SIZE, shuffle=shuffle, num_workers=num_workers, pin_memory=pin_memory)
# data_loader.to(device)
# -
signal_synthesis_dataset.__len__()
# %%time
# data_loader_iter = iter(data_loader)
for index, i in enumerate(data_loader):
# i = next(data_loader)
if index < 32-1:
pass
else:
break
print(i[1].shape,i[0].min(), i[0].max(), index)
# +
# %%time
stft_sig = torch.stft(i[0], n_fft=n_fft, hop_length=hop_len, win_length=win_length)
i[1].max()
nan_sig = signal_synthesis_dataset.__getitem__(119)
# -
nan_sig[0].max(), nan_sig[1].min()
stft_sig.max()
def normalize(tensor):
tensor_minusmean = tensor - tensor.min()
return tensor_minusmean/tensor_minusmean.abs().max()
# +
aud = i[0][0]
aud.dtype
aud.max(), aud.min()
# -
x = aud.t().to("cpu").numpy()
ipd.Audio(x, rate=default_sr)
# +
sig1 = i[0].unsqueeze(dim=1)
sig2 = i[0].unsqueeze(dim=1)
stacked_sig = torch.cat((sig1, sig2), dim=1)
sig2 = i[0].unsqueeze(dim=1)
sig2.shape
torch.sum(stacked_sig, dim=1).shape
# -
np.floor(((default_sr*sec) - (win_length - 1) - 1)/ hop_len + 5)
# +
n_fft // 2 + 1
n_fft // 2 + 1
# -
stft_sig.shape
# +
class InstantLayerNormalization(nn.Module):
def __init__(self, in_shape, out_shape):
self.in_shape = in_shape
self.out_shape = out_shape
self.epsilon = 1e-7
self.gamma = None
self.beta = None
super(InstantLayerNormalization, self).__init__()
self.gamma = torch.ones(out_shape)
self.gamma = nn.Parameter(self.gamma)
self.beta = torch.zeros(out_shape)
self.beta = nn.Parameter(self.beta)
def forward(self, inps):
mean = inps.mean(-1, keepdim=True)
variance = torch.mean(torch.square(inps - mean), dim=-1, keepdim=True)
std = torch.sqrt(variance + self.epsilon)
outs = (inps - mean) / std
print(outs.shape, self.gamma.shape)
outs = outs * self.gamma
outs = outs + self.beta
return outs
class Multiply():
def __init__(self):
super(Multiply, self).__init__()
def forward(self, ten1, ten2):
mul_out = torch.mul(ten1, ten2)
return mul_out
class NoiseReducer(nn.Module):
def __init__(self, default_sr, n_fft, win_length, hop_len, sec, dropout=0.5, batch_first=True, stride=2, normalized=False, bidir=False):
self.default_sr = default_sr
self.n_fft = n_fft
self.win_length = win_length
self.hop_len = hop_len
self.sec = sec
self.normalized = normalized
self.conv_filters = 256
# Universal LSTM Units
self.batch_first = True
self.dropout = 0.25
self.bidir = bidir
# LSTM 1 UNITS
self.rnn1_dims = n_fft // 2 + 1
self.hidden_size_1 = 256
self.num_layers = 2
# LSTM 2 UNITS
self.rnn2_dims = self.conv_filters
self.hidden_size_2 = self.hidden_size_1
# Conv1d Layer Units
self.conv1_in = 1
self.conv1_out = self.conv_filters
# InstanceNorm Layer Units
self.instance1_in = self.rnn1_dims
self.instance2_in = self.conv1_out
# Dense1 Layer Units
self.dense1_in = self.hidden_size_1
self.dense1_out = self.rnn1_dims #int(np.floor(((default_sr*sec) - (win_length - 1) - 1)/ hop_len + 5))#3))
# Dense2 Layer Units
self.dense2_in = self.hidden_size_2
self.dense2_out = self.conv1_out
# Dense3 Layer Units
self.dense3_in = self.hidden_size_1
self.dense3_out = self.rnn1_dims
# Conv2d Layer Units
self.conv2_in = self.dense2_out
self.conv2_out = self.conv_filters
super(NoiseReducer, self).__init__()
self.lstm1 = nn.LSTM(input_size=self.rnn1_dims, hidden_size=self.hidden_size_1, num_layers=self.num_layers, batch_first=self.batch_first, dropout=self.dropout, bidirectional=self.bidir)
self.lstm3 = nn.LSTM(input_size=self.rnn1_dims, hidden_size=self.hidden_size_1, num_layers=self.num_layers, batch_first=self.batch_first, dropout=self.dropout, bidirectional=self.bidir)
print(self.rnn2_dims)
self.lstm2 = nn.LSTM(input_size=self.rnn2_dims, hidden_size=self.hidden_size_2, num_layers=self.num_layers*2, batch_first=self.batch_first, dropout=self.dropout, bidirectional=self.bidir)
self.instancenorm1 = nn.InstanceNorm1d(self.rnn1_dims)
self.instancenorm2 = nn.InstanceNorm1d(self.rnn2_dims)
self.instancenorm3 = nn.InstanceNorm1d(self.rnn1_dims)
self.dense1 = nn.Linear(self.dense1_in, self.dense1_out)
self.dense2 = nn.Linear(self.dense2_in, self.dense2_out)
self.dense3 = nn.Linear(self.dense3_in, self.dense3_out)
self.conv1 = nn.Conv1d(self.conv1_in, self.conv1_out, kernel_size=3, stride=1, padding=1)
self.conv2 = nn.Conv1d(self.conv2_in, self.conv2_out, kernel_size=3, stride=1, padding=1)
self.prelu_conv1 = nn.PReLU(self.conv1_out)
self.prelu_conv2 = nn.PReLU(self.conv2_out)
@torch.jit.export
def stft_layer(self, sig):
sig_stft = torch.stft(sig, n_fft=self.n_fft, hop_length=self.hop_len, win_length=self.win_length)
sig_cplx = torch.view_as_complex(sig_stft)
mag = sig_cplx.abs().permute(0, 2, 1)
angle = sig_cplx.angle().permute(0, 2, 1)
return [mag, angle]
@torch.jit.export
def istft_layer(self, mag, angle):
mag = mag.permute(0, 2, 1)
angle = angle.permute(0, 2, 1)
mag = torch.unsqueeze(mag, dim=-1)
angle = torch.unsqueeze(angle, dim=-1)
pre_stft = torch.cat((mag, angle), dim=-1)
stft_sig = torch.istft(pre_stft, n_fft=self.n_fft, win_length=self.win_length, hop_length=self.hop_len)
return stft_sig
@torch.cuda.amp.autocast()
def forward(self, inp_tensor):
mag, angle = self.stft_layer(inp_tensor)
mag_norm = self.instancenorm1(mag)
angle_norm = self.instancenorm3(angle)
x_mag, hidden_states_mag = self.lstm1(mag_norm)
x_angle, hidden_states_angle = self.lstm3(angle_norm)
mask_mag = F.sigmoid(self.dense1(x_mag))
estimated_mag = torch.mul(mag, mask_mag)
mask_angle = F.sigmoid(self.dense3(x_angle))
estimated_angle = torch.mul(angle, mask_angle)
signal = self.istft_layer(estimated_mag, estimated_angle)
signal = signal.unsqueeze(dim=1)
feature_rep = self.conv1(signal)
feature_rep = self.prelu_conv2(feature_rep)
feature_norm = self.instancenorm2(feature_rep)
feature_norm = feature_norm.permute(0, 2, 1)
x, hidden_states = self.lstm2(feature_norm)
mask = self.dense2(x)
feature_mask = F.sigmoid(mask)
feature_mask = feature_mask.permute(0, 2, 1)
estimate_feat = torch.mul(feature_rep, feature_mask)
estimate_frames = (self.conv2(estimate_feat))
estimate_frames = self.prelu_conv2(estimate_frames)
estimate_sig = torch.sum(estimate_frames, dim=1)
return estimate_sig
# -
class Negative_SNR_Loss(nn.Module):
def __init__(self):
super(Negative_SNR_Loss, self).__init__()
def forward(self, sig_pred, sig_true):
sig_true_sq = torch.square(sig_true)
sig_pred_sq = torch.square(sig_true - sig_pred)
sig_true_mean = torch.mean(sig_true_sq)
sig_pred_mean = torch.mean(sig_pred_sq)
snr = sig_true_mean / sig_pred_mean + 1e-7
loss = -1*torch.log10(snr)
return loss
# +
use_scripted_model = False
if not use_scripted_model:
print("Using Primary model")
model = NoiseReducer(default_sr=default_sr, n_fft=n_fft, win_length=win_length, hop_len=hop_len, sec=sec).to(device)
model.to(device)
else:
print("Using Scripted Model")
model = scripted_model
optimizer = torch.optim.AdamW(model.parameters(), lr=0.0001)
criterion = Negative_SNR_Loss()
n_epochs=100
model.train()
scaler = torch.cuda.amp.GradScaler()
# +
# fake_inputs = torch.randn(BATCH_SIZE, int(default_sr*sec)).type(torch.float32).to(device)
# outs = model(fake_inputs)
# outs.shape
# +
scaler = torch.cuda.amp.GradScaler()
for epoch in range(1, n_epochs+1):
loop = tqdm(enumerate(data_loader), leave=True, total=len(data_loader))
train_loss = np.zeros((len(data_loader)))
for index, (data, target) in loop:
data = data.type(torch.float32).to(device)
target = target.type(torch.float32).to(device)
optimizer.zero_grad(set_to_none=True)
with torch.cuda.amp.autocast():
output = model(data)
loss = criterion(output, target)
scaler.scale(loss).backward()
scaler.step(optimizer)
scaler.update()
train_loss[index] = loss.item()
if np.isnan(loss.item()) or np.isnan(np.sum(train_loss)/index+1e-5):
print(f"Data shape = {data.shape}\nTarget Shape = {target.shape}, \nindex = {index}")
loop.set_description(f"Epoch: [{epoch}/{n_epochs}]\t")
loop.set_postfix(loss = np.sum(train_loss)/index+1e-5)
# -
| .ipynb_checkpoints/Training_model_2-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
import torch2trt
import matplotlib.pyplot as plt
import cv2
model = torch.load('/home/brian/models/weights.pt')
model = model.cuda().eval().half()
ino = 858
# Read a sample image and mask from the data-set
img = cv2.imread(f'/home/brian/mycar/data/tub_47_20-10-15/{ino:03d}_cam-image1_.jpg')
def equalize(im):
gray = cv2.cvtColor(im,cv2.COLOR_RGB2GRAY)
# create a CLAHE object
clahe = cv2.createCLAHE(clipLimit=2.0, tileGridSize=(8,8))
cli = clahe.apply(gray)
im2 = cv2.cvtColor(cli,cv2.COLOR_GRAY2RGB)
return im2
img2 = equalize(img)
crop_img = img2[250:550, 125:775].copy()
img3 = crop_img.transpose(2,0,1).reshape(1,3,300,650)
input = torch.from_numpy(img3).type(torch.cuda.HalfTensor)/255
# +
import time
torch.cuda.current_stream().synchronize()
t0 = time.time()
with torch.no_grad():
output = model(input)
torch.cuda.current_stream().synchronize()
t1 = time.time()
print(1.0 / (t1 - t0))
# -
output['out'].shape
plt.hist(output['out'].data.cpu().numpy().flatten())
plt.figure(figsize=(10,10));
plt.subplot(131);
plt.imshow(img3[0,...].transpose(1,2,0));
plt.title('Image')
plt.axis('off');
plt.subplot(132);
plt.imshow(output['out'].cpu().detach().numpy()[0][0]>0.4);
plt.title('Segmentation Output')
plt.axis('off');
| torcheval.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # A Schema of Groups
# ---
#
# In this notebook we present a typology for modelling groups from natural language.
#
# The schema is structured using the "five group problems" identified by Martin which any group must resolve to be successful. As group attributes of the schema they are applied in seven different calssifications of ideology. At the highest level of abstraction these features are:
#
# 1. Identity:the identification of who is in the group and who is out of the group to determine who should, versus who should not, benefit from the advantages of group living.
# * identity: named groups
# * ingroup: groups identifying an ingroup
# * outgroup: groups identifying an outgroup
# * entities: entities identifying the group context
# 2. Hierarchy: a system of governance to establish group leadership and resolve problems caused by status seeking to enable resource distribution.
# * Title: titles given to people within a particular context
# * People: who are the figures a group reveres
# 3. Disease control: the control of diseases for the maintenance of group health.
# * this feature is considered as a social category rather than a feature
# 4. Trade: independent of hierarchy, systems constituting the fair terms of exchange between individuals towards developing an underpinning concept of altruism in a society.
# * Good: terms of exchange considered to be good
# * Bad: terms of exchange considered to be bad
# 5. Punishment: the group jurisprudence for moderating the systems seeking to resolve the other group problems.
# * Right: considered to be just
# * Wrong: considered to be unjust and worthy of punishment
#
# We identified sets of seed words as linguistic representations of religion and ideology. These sets were classified by a named concept then placed in schema according to attribute and group ideology.
#
# This process identified eight classifications for group ideologies referred to by each orator: social, academia, medical, geopolitics, religion, economic, justice and military.
# ## The Schema
#
# The next cell contains the schema and seed terms classified by named concept
# +
## Create a json object of the group typology
import json
import os
from datetime import datetime
group_typology = {
"social" : {
"identity" : {
"SOCGROUP" : ["ladies", "gentlemen", "men", "women", "Women", "boys", "girls", "youth", "society", "people", "children", "minority", \
"mankind", "passengers", "stranger", "group", "community", "organization", "brotherhood", "network", "alliance", "brethren", \
"tribe", "population", "ummah", "Ummah", "human", "personnel", "person", "man", "Woman", "woman", "boy", "girl", "child", \
"humankind", "countryman", "volunteer", "individual", "freeman", "humanity", "society"]},
"ingroup" : {
"SELF" : ["we", "our", "i"],
"FAMILY" : ["family", "parent", "children", "forefather", "spouse", "mother", "father", "husband", "wife", "mom", "mum", "dad", "son", \
"daughter", "brother", "sister", "grandson", "granddaughter", "descendent", "ancestor"],
"AFFILIATE" : ["ourselves", "collaborator", "friend", "ally", "associate", "partner", "companion", "fellow", "kinship"]},
"outgroup" : {
"OUTCAST": ["imbecile", "critic", "wolf", "snake", "dog", "hypocrite"]},
"entity" : {
"CAUSE" : ["goal", "cause", "struggle"],
"CREDO" : ["philosophy", "Philosophy", "ideology", "Belief", "belief", "creed"],
"LOCATION" : ["ground", "homeland", "sanctuary", "safe haven", "land", "sea", "site", "underworld"],
"SOCFAC" : ["installation", "home", "shelter", "institution", "facility", "infrastructure", "refuge", "tower"],
"SOCWORKOFART" : ["poetry", "song", "picture", "art"]},
"hierarchy" : {
"TITLE" : ["mr", "mrs", "miss", "ms", "leader"]},
"LEADERSHIP" : ["leadership"]
"trade" : {
"BENEVOLANCE" : ["love", "kinship", "honesty", "tolerance", "patience", "decency", "sympathy", "peace", "good", "best", "great", \
"goodness", "hope", "courage", "resolve", "friendship", "loving", "peaceful", "rightness", "brave", \
"strong", "peaceful", "fierce", "honesty", "kind", "generous", "resourceful", "truth", "pride", "defiance", "strength", \
"comfort", "solace", "respect", "dignity", "honor", "danger", "freedom", "honorable", "grateful", "compassion", "condolence", \
"sympathy", "fulfillment", "dedication", "dignity", "noble", "truthfulness", "happy", "enthusiasm", "perseverance", \
"persistence", "toughness", "beauty", "beautiful", "commendable", "praiseworthy", "destiny", "generosity", "supremecy", \
"obedience", "superior"],
"MALEVOLANCE" : ["grief", "sorrow", "tradegy", "damage", "bad", "misinformation", "confusion", "falsehood", "humiliation", "catastrophe", \
"terror", "fear", "threat", "cruelty", "danger", "anger", "harm", "suffering", "harrassment", "deceit", "death", "anger", \
"hate", "hatred", "adversity", "chaos", "loneliness", "sadness", "misery", "prejudice", "horrifying", "cynicism", "despair", \
"rogue", "hostile", "dangerous", "tears", "peril", "unfavourable", "vile", "sad", "cowardly", "grieve", "shame", "ugly", \
"insanity", "arrogance", "hypocrisy", "horror", "monstrous", "suspicious", "disaster", "malice", "menace", "repressive", \
"malicious", "nightmare", "isolation", "debauchery", "greedy"]},
"punishment" : {
"PERMITTED" : ["acceptable", "right", "laugh", "deed"],
"FORBIDDEN" : ["unacceptable", "wrong", "catastrophic", "disastrous", "catastrophe", "mischief", "disappoint", "humiliate", "deceive", "lie"]}
},
"academia" : {
"identity" : {
"ACADEMICGROUP" : ["students", "graduates", "graduate", "scholar"]},
"ingroup" : {},
"outgroup" : {},
"entity" : {
"ACADEMICENTITY" : ["school", "university"]
},
"hierarchy" : {
"ACADEMICTITLE" : ["teacher", "dr", "professor"]
},
"trade" : {},
"punishment" : {}
},
"medical" : {
"identity" : {
"MEDICALGROUP" : ["blood donors", "disabled", "injured"]},
"ingroup" : {},
"outgroup" : {
"VERMIN" : ["vermin", "parasite"]},
"entity" : {
"MEDICALENT" : ["heart", "soul", "skin graft", "tomb", "palm", "limb", "drug", "chemical", "biological", "heroin", "vaccine", "health", \
"blood", "body", "medicine", "remedy", "organ", "intestine", "tongue"],
"SEXUALITY" : ["fornication", "homosexuality", "sex"],
"MEDICALFAC" : ["hospital"],
"INTOXICANT" : ["intoxicant", "sarin", "anthrax", "nerve agent", "nerve gas"],
"DISEASE" : ["AIDS"]},
"hierarchy" : {
"MEDTITLE" : ["doctor", "nurse"]},
"trade" : {
"HEALTHY" : ["life"],
"UNHEALTHY" : ["death", "unhealed", "illness", "disease", "filthy", "wound", "injury", "scar", "suffocation"]},
"punishment" : {
"CLEANSE" : ["cure", "remedy", "cleanse"],
"POISON" : ["poison", "pollute", "bleed"]}
},
"geopolitics" : {
"identity" : {
"GPEGROUP" : ["ministry", "Ministry", "government", "civilian", "nation", "Union", "civilization", "congress", "Congress", "alliance", \
"patriot", "citizen", "journalist", "diplomat", "agency", "delegate", "coalition", "axis", "compatriots", "administration", \
"monarchy", "political party", "communist"]},
"ingroup" : {},
"outgroup" : {
"GPEOUTGROUP" : ["regime", "opponent"]},
"entity" : {
"GPEENTITY" : ["human rights", "unity", "diplomatic", "citizenship", "legislation", "senate", "secretary", "elect", "election", "reign", \
"embassy", "policy", "diplomacy", "media", "power", "edict"],
"TERRITORY" : ["homeland", "territory", "planet", "land", "peninsula", "city", "country", "neighborhood", "home", "region", "place", \
"area", "peninsula", "continent", "reform", "kingdom", "empire"],
"GPEFAC" : [],
"GPEWORKOFART" : []},
"hierarchy" : {
"GPETITLE" : ["president", "minister", "speaker", "prime minister", "senator", "mayor", "governor", "President", "Minister", "Prime Minister", \
"Speaker", "Senator", "Mayor", "Governor", "King", "king", "Prince", "Leader", "commander-in-chief", "ruler", "chairman", \
"congressman", "amir", "pharaoh"]},
"trade" : {
"GPEIDEOLOGY" : ["liberty", "sovereignty", "pluralism", "patriotism", "democracy", "communism", "bipartisanship"],
"AUTHORITARIANISM" : ["nationalism", "fascism", "nazism", "totalitarianism", "nazi", "tyranny", "sectarianism", "anti-semitism"],
"CONFRONTATION" : ["jihad", "confrontation", "feud", "partisanship", "division", "dissociation"]},
"punishment" : {
"JUST" : [],
"UNJUST" : ["inequality", "usury", "poverty", "slavery", "injustice", "oppression", "subdual", "misrule", "occupation", "usurpation", \
"starvation", "starving", "servitude", "surpression"]}
},
"religion" : {
"identity" : {
"RELGROUP" : ["ulamah", "Ulamah", "ulema", "moguls", "Moguls", "Sunnah", "Seerah", "Christian", "sunnah", "christian", "muslims", "islamic", \
"sunnah", "seerah", "polytheist", "houries", "the people of the book", "merciful"]},
"ingroup" : {
"BELIEVER" : ["believer"]},
"outgroup" : {
"APOSTATE" : ["kufr", "Kufr", "kuffaar", "infidel", "evildoer", "cult", "mushrik", "unbeliever", "disbeliever", "mushrikeen", "pagan", \
"idolater", "apostate"]},
"entity" : {
"RELENTITY" : ["faith", "pray", "prayer", "mourn", "vigil", "prayer", "remembrance", "praise", "bless", "last rites", "angel", "soul", \
"memorial", "revelation", "sanctify", "grace", "religion", "repentance", "exalted", "repent", "seerah", "confession", \
"exaltation", "praise", "commandment", "wonderment", "supplication", "worship", "testament"],
"RELIGIOUSLAW" : ["shari'ah", "Shari'ah", "shari'a", "Shari'a", "fatawa", "Fatawa", "fatwa", "Fatwas"],
"FAITH" : ["da'ees", "piety", "creationism"],
"RELFAC" : ["Mosque", "mosque", "sanctity", "cathedral", "Cathedral"],
"RELWORKOFART" : ["sheerah"],
"RELPLACE" : ["heaven", "paradise"]},
"hierarchy" : {
"DEITY" : ["all-wighty"],
"RELFIGURE" : ["Apostle", "Prophet", "apostle", "prophet", "lord", "priest"],
"RELTITLE" : ["priest", "cleric", "Immam", "immam", "saint", "st.", "sheikh"]},
"trade" : {
"HOLY" : ["pious", "devout", "holy", "righteous", "serve", "sacrifice", "forgive", "martyrdom", "piety", "polytheism", "divine", "miracle"],
"UNHOLY" : ["hell", "hellish", "unholy", "unrighteous", "evil", "devil", "devilish", "demon", "demonic", "evildoe", "satan", "immoral", \
"immorality", "non-righteous", "blasphemy"]},
"punishment" : {
"VIRTUE" : ["Grace", "grace", "halal", "forgiveness", "mercy", "righteous", "mercy", "righteousness", "purity"],
"SIN" : ["iniquity", "haram", "adultery", "sin", "blashpeme"]}
},
"economic" : {
"identity" : {
"ECONGROUP" : ["merchant", "employee", "economist", "worker", "entrepreneur", "shopkeeper", "servant", "company", "shareholder", \
"contractor", "merchant", "consumer", "rich", "passenger", "corporation"]},
"ingroup" : {},
"outgroup" : {
"COMPETITOR" : ["competitor"]},
"entity" : {
"ECONENTITY" : ["economic", "tax", "trade", "work", "currency", "bank", "business", "economy", "asset", "fund", "sponsor", "shop", \
"financing", "innovation", "micromanage", "export", "job", "budget", "spending", "paycheck", "market", "growth", \
"investment", "factory", "welfare", "pension", "accounting", "retirement", "industry", "agriculture", "income", \
"spending", "expensive", "purchase", "wealth", "economical", "booty", "buy", "sell", "imprisonment", "denunciation", \
"price", "monie", "gambling", "production", "advertising", "tool", "cargo","reconstruction", "contract", "tax", \
"financial", "wealth", "power", "prosperity"],
"COMMODITY" : ["money", "oil", "water", "energy", "currency", "jewel", "jewellery", "gold", "industry", "agriculture", "inflation", "debt", "income", "price"],
"ECONFAC" : ["airport", "subway", "farm", "charity"],
"ECONWORKOFART" : []},
"hierarchy" : {
"ECONTITLE" : ["ceo"],
"ECONFIGURE" : []},
"trade" : {
"ECONOMICAL" : ["capitalism", "communism", "economical", "economic", "tourism", "commercial"],
"UNECONOMICAL" : ["uncommercial", "boycott", "bankruptcy"]},
"punishment" : {
"EQUITABLE" : ["reward"],
"UNEQUITABLE" : ["recession", "unemployment"]}
},
"justice" : {
"identity" : {
"SECGROUP" : ["police", "officers", "policeman", "law enforcement", "firefighter", "rescuer", "lawyer", "agent", "authority", \
"protector", "guardian", "captive", "marshal"],
"VICTIM" : ["victim", "dead", "casualty", "innocent", "persecuted", "slave"]},
"ingroup" : {},
"outgroup" : {
"CRIMEGROUP" : ["criminal", "mafia", "prisoner", "murderer", "terrorist", "hijacker", "outlaw", "violator", "killer", "executioner", "thief"]},
"entity": {
"SECENTITY" : ["trial", "security", "law", "law-enforcement", "intelligence", "arrest", "decree", "sailor", "surveillance", "warrant", \
"penalty", "statute", "investigate", "enforce", "attorney", "treaty", "duty", "jail", "imprisonment", "justification", \
"judge", "prohibit", "custody", "shield"],
"LAW" : [],
"SECFAC" : ["prison", "sanctuary"],
"LEGALWORKOFART" : []},
"hierarchy" : {
"LEGALPERSON" : [],
"SECTITLE" : []},
"trade" : {
"LAWFUL" : ["duty", "justice", "morality"],
"UNLAWFUL" : ["crime", "terrorism", "extremism", "murderous", "imtimidation", "harrassment", "trafficking", "criminal", "suicide", "plot", \
"brutality", "coercion", "subversion", "bioterrorism", "propaganda", "corrupt", "corruption", "scandal", "betray", "betrayal", \
"misappropriation"]},
"punishment" : {
"LEGAL" : ["legal", "protect", "legitimate", "liberation", "liberate"],
"ILLEGAL" : ["illegal", "counterfeit", "money-laundering", "guilty", "blackmail", "threaten", "punishment", "conspiracy", "illegitimate", \
"infringement", "adultery", "wicked", "tricked", "harm", "incest"],
"PHYSICALVIOLENCE" : ["murder", "hijack", "kill"]}
},
"military" : {
"identity" : {
"ARMEDGROUP" : ["commander", "vetran", "Vetran", "occupier", "guard", "invader", "military", "Mujahideen", "mujahideen", "army", "navy", \
"air force", "troops", "defender", "recruit", "guerrilla", "knight", "special forces", "fatality", "martyr", "vanguard"],
"BELIGERENT" : ["aggressor", "troop", "fighter", "soldier", "warrior", "Mujahid", "mujahid", "soldier"]},
"ingroup" : {},
"outgroup" : {
"ENEMY" : ["traitor", "oppressor", "enemy", "crusader", "aggressor", "invader", "occupier"]},
"entity" : {
"MILENTITY" : ["battlefield", "beachead", "campaign", "military", "training camp", "armed", "force", "uniform", "chain of command", "target", \
"defense", "nuclear", "mission", "battleship", "aircraft carriers", "infantry", "tank", "air power", "shooter", "cavalry", \
"arming", "enlist", "conquest", "base", "buy", "surrender", "soldier", "airman", "marine", "biodefense"],
"WEAPON" : ["weapon", "weaponry", "bomb", "missile", "munition", "explosive", "arms", "bullet", "sword", "spear", "gun", "rocket"],
"MILFAC" : ["fortress"],
"MILWORKOFART" : []},
"hierarchy" : {
"MILRANK" : ["lieutenant", "commander", "adjutant", "mujahid"],
"MILFIGURE" : []},
"trade" : {
"WARFARE" : ["victory", "war", "warfare", "battle", "blockade"],
"MILACTION" : ["destruction", "violence", "conflict", "slain", "besiege", "massacre", "atrocity", "aggression", "attack", "assault", "fight",\
"explosion", "combat", "invasion", "ruin", "bombardment", "expel", "fighting", "defeat", "expel", "ambush", "overthrow", "destabilize"]},
"punishment" : {
"BARBARY" : ["genocide", "brutalize", "holocaust", "torture", "slaughter"]}
}
}
filepath = "C:/Users/Steve/OneDrive - University of Southampton/CulturalViolence/KnowledgeBases/data/"
with open(os.path.join(filepath, "group_typology.json"), "wb") as f:
f.write(json.dumps(group_typology).encode("utf-8"))
print("complete at: ", datetime.now().strftime("%d/%m/%Y - %H:%M:%S"))
# -
# ## Structure of the Schema
#
# The following cell displays the structure for the schema and how each named concept has been classified.
# +
## https://www.datacamp.com/community/tutorials/joining-dataframes-pandas
## Display a DataFrame of the Typology
import pandas as pd
labels = []
typology = dict()
typology_chart = dict()
## create a list of keys
typology = {ideology: {subcat: ', '.join(list(terms.keys())) for (subcat, terms) in value.items()}
for (ideology, value) in group_typology.items()}
keys = [list(cat.keys()) for cat in list(typology.values())][0]
## Create frames for table
frames = []
typology = {ideology: {subcat: list(terms.keys()) for (subcat, terms) in value.items()}
for (ideology, value) in group_typology.items()}
for frame in [list(cat.keys()) for cat in list(typology.values())][0]:
frames.append(pd.DataFrame.from_dict({k : v[frame] for k, v in list(typology.items())}, orient = 'index').fillna("").T)
# display table
display(pd.concat(frames , keys = keys))
# -
# ## Results for Each Orator
#
# The following cell shows how each of the ideologies are represented over each orators texts.
#
# From the total number of concepts of each speech, these infographics shows the percentage of concepts used by each orator for each ideology.
#
# In all speeches they are addressing "the people" which is why "social" scores most highly.
#
# "Justice" scores most highly in Bush's speech on the 26/10/2001 in which he addresses the signing of the US Patriot Act. As might be expected, in his speech on the 14/09/2001 at the Episcopal National Cathedral for a day of Prayer and Remembrance, "religion" features most highly.
#
# For <NAME> in his second and third speeches, religion, military and geopolitics feature highly. In his speech following 9/11, religion features most highly in how he confers a divine legitimacy to the attacks.
#
#
# Using these terms, this annotation framework could be extended to create a new topic modelling schema specific for cultural violence.
# +
import os
import json
import pandas as pd
filepath = r"C:/Users/Steve/OneDrive - University of Southampton/CulturalViolence/data/"
files = ['bushideologiesfile.json', 'binladenideologiesfile.json']
cmp = "Reds"
for file in files:
with open(os.path.join(filepath, file), "r") as f:
table = json.load(f)
display(pd.DataFrame.from_dict(table, orient = 'index').fillna("0").T \
.style.background_gradient(cmap=cmp).format("{:.0%}"))
| Obj 0 - pre-processing/Experiment 0.2 - Developing the Markup Schema/.ipynb_checkpoints/Group Schema-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.linear_model import Perceptron
from sklearn.pipeline import Pipeline
from sklearn.datasets import load_files
from sklearn.model_selection import train_test_split
from sklearn import metrics
# The training data folder must be passed as first argument
languages_data_folder = './data/languages/paragraphs'
dataset = load_files(languages_data_folder)
# Split the dataset in training and test set:
docs_train, docs_test, y_train, y_test = train_test_split(dataset.data, dataset.target, test_size=0.5)
# Build a vectorizer that splits strings into sequence of 1 to 3 characters instead of word tokens
vectorizer = TfidfVectorizer(ngram_range=(1, 3), analyzer='char', use_idf=False)
# Build a vectorizer / classifier pipeline using the previous analyzer
# the pipeline instance should stored in a variable named clf
clf = Pipeline([('vec', vectorizer),('clf', Perceptron()),])
# Fit the pipeline on the training set
clf.fit(docs_train, y_train)
# Predict the outcome on the testing set in a variable named y_predicted
y_predicted = clf.predict(docs_test)
# Print the classification report
print(metrics.classification_report(y_test, y_predicted, target_names=dataset.target_names))
# Plot the confusion matrix
cm = metrics.confusion_matrix(y_test, y_predicted)
print(cm)
# Predict the result on some short new sentences:
sentences = [
u'I am <NAME> and i am a student.',
u'Soy <NAME> una chica.',
u'Je suis <NAME> ça va.',
u'Я <NAME>.',
u'私はです.',
u'Meu nome é <NAME>.',
u'Ich heiße <NAME> und ich komme aus Brasilien.',
]
predicted = clf.predict(sentences)
for s, p in zip(sentences, predicted):
print(u'The language of "%s" is "%s"' % (s, dataset.target_names[p]))
| lab02/text_analytics/language_train_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.3 64-bit (''base'': conda)'
# language: python
# name: python373jvsc74a57bd0210f9608a45c0278a93c9e0b10db32a427986ab48cfc0d20c139811eb78c4bbc
# ---
# ## Load the data
# ## Clean the data
# ## Modelling
import pandas as pd
data = pd.read_csv('./data.csv')
X,y = data.drop('target',axis=1),data['target']
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test = train_test_split(X,y,test_size=0.25)
import torch
import torch.nn as nn
import numpy as np
X_train = torch.from_numpy(np.array(X_train).astype(np.float32))
y_train = torch.from_numpy(np.array(y_train).astype(np.float32))
X_test = torch.from_numpy(np.array(X_test).astype(np.float32))
y_test = torch.from_numpy(np.array(y_test).astype(np.float32))
X_train.shape
X_test.shape
y_train.shape
y_test.shape
# #### Modelling
import torch.nn.functional as F
class Test_Model(nn.Module):
def __init__(self):
super().__init__()
self.fc1 = nn.Linear(13,64)
self.fc2 = nn.Linear(64,128)
self.fc3 = nn.Linear(128,256)
self.fc4 = nn.Linear(256,512)
self.fc5 = nn.Linear(512,1024)
self.fc6 = nn.Linear(1024,512)
self.fc7 = nn.Linear(512,1)
def forward(self,X):
preds = self.fc1(X)
preds = F.relu(preds)
preds = self.fc2(preds)
preds = F.relu(preds)
preds = self.fc3(preds)
preds = F.relu(preds)
preds = self.fc4(preds)
preds = F.relu(preds)
preds = self.fc5(preds)
preds = F.relu(preds)
preds = self.fc6(preds)
preds = F.relu(preds)
preds = self.fc7(preds)
return F.sigmoid(preds)
device = torch.device('cuda')
X_train = X_train.to(device)
y_train = y_train.to(device)
X_test = X_test.to(device)
y_test = y_test.to(device)
PROJECT_NAME = 'Heart-Disease-UCI'
def get_loss(criterion,X,y,model):
model.eval()
with torch.no_grad():
preds = model(X.float().to(device))
preds = preds.view(len(preds),).to(device)
y = y.view(len(y),).to(device)
loss = criterion(preds,y)
model.train()
return loss.item()
def get_accuracy(preds,y):
correct = 0
total = 0
for real,pred in zip(y_train,preds):
if real == pred:
correct += 1
total += 1
return round(correct/total,3)
import wandb
from tqdm import tqdm
EPOCHS = 212
# EPOCHS = 100
# +
# model = Test_Model().to(device)
# optimizer = torch.optim.SGD(model.parameters(),lr=0.25)
# criterion = nn.L1Loss()
# wandb.init(project=PROJECT_NAME,name='baseline')
# for _ in tqdm(range(EPOCHS)):
# preds = model(X_train.float().to(device))
# preds = preds.view(len(preds),)
# preds.to(device)
# loss = criterion(preds,y_train)
# optimizer.zero_grad()
# loss.backward()
# optimizer.step()
# wandb.log({'loss':loss.item(),'val_loss':get_loss(criterion,X_test,y_test,model),'accuracy':get_accuracy(X_train,y_train,model),'val_accuracy':get_accuracy(X_test,y_test,model)})
# wandb.finish()
# +
# preds[:10]
# +
# preds = torch.round(preds)
# +
# correct = 0
# total = 0
# for real,pred in zip(y_train,preds):
# if real == pred:
# correct += 1
# # total += 1
# +
# round(correct/total,3)
# +
## Testing Modelling
# -
import torch
import torch.nn as nn
class Test_Model(nn.Module):
def __init__(self,num_of_layers=1,activation=F.relu,input_shape=13,fc1_output=32,fc2_output=64,fc3_output=128,fc4_output=256,output_shape=1):
super().__init__()
self.num_of_layers = num_of_layers
self.activation = activation
self.fc1 = nn.Linear(input_shape,fc1_output)
self.fc2 = nn.Linear(fc1_output,fc2_output)
self.fc3 = nn.Linear(fc2_output,fc3_output)
self.fc4 = nn.Linear(fc3_output,fc4_output)
self.fc5 = nn.Linear(fc4_output,fc3_output)
self.fc6 = nn.Linear(fc3_output,fc3_output)
self.fc7 = nn.Linear(fc3_output,output_shape)
def forward(self,X,activation=False):
preds = self.fc1(X)
if activation:
preds = self.activation(preds)
preds = self.fc2(preds)
if activation:
preds = self.activation(preds)
preds = self.fc3(preds)
if activation:
preds = self.activation(preds)
preds = self.fc4(preds)
if activation:
preds = self.activation(preds)
preds = self.fc5(preds)
if activation:
preds = self.activation(preds)
for _ in range(self.num_of_layers):
preds = self.fc6(preds)
if activation:
preds = self.activation(preds)
preds = self.fc7(preds)
preds = F.sigmoid(preds)
return preds
device = torch.device('cuda')
# +
# preds = torch.round(preds)
# +
# num_of_layers = 1
# fc1_output = 256
# fc2_output =
# fc3_output =
# fc4_output =
# optimizer = torch.optim.SGD
# criterion = nn.MSELoss
# lr = 0.125
# activtion = nn.Tanh()
# -
fc3_outputs = [16,32,64,128,256,512,1024]
for fc3_output in fc3_outputs:
model = Test_Model(num_of_layers=1,activation=nn.Tanh(),fc1_output=256,fc2_output=fc2_output,fc3_output=fc3_output).to(device)
model.to(device)
optimizer = torch.optim.SGD(model.parameters(),lr=0.125)
criterion = nn.MSELoss()
wandb.init(project=PROJECT_NAME,name=f'fc3_output-{fc3_output}')
for _ in tqdm(range(212)):
preds = model(X_train.float().to(device),True)
preds = preds.view(len(preds),)
preds.to(device)
loss = criterion(preds,y_train)
optimizer.zero_grad()
loss.backward()
optimizer.step()
wandb.log({'loss':loss.item(),'val_loss':get_loss(criterion,X_test,y_test,model),'accuracy':get_accuracy(preds,y_train)})
wandb.finish()
| wandb/run-20210520_093748-32zi0bkr/tmp/code/00-main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Alookapola/pytorch-tutorial/blob/master/Regression.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="y1TRdOtdw8_w" outputId="0c1411a4-7f3c-44b7-8d61-33b2b529d141"
# 设置路径
tr_path = 'covid.train.csv' # path to training data
tt_path = 'covid.test.csv' # path to testing data
# !gdown --id '19CCyCgJrUxtvgZF53vnctJiOJ23T5mqF' --output covid.train.csv
# !gdown --id '1CE240jLm2npU-tdz81-oVKEF3T2yfT1O' --output covid.test.csv
# + colab={"base_uri": "https://localhost:8080/"} id="W0Fb4DLO7zwn" outputId="aa244175-7e9d-4497-fd9e-5780fae8567c"
# test
with open(tr_path, 'r') as fp:
data = list(csv.reader(fp))
data = np.array(data[1:])[:, 1:].astype(float)
data.shape
# (2700, 94)
# + [markdown] id="oSf_AZC77zBo"
#
# + id="ngjJjUEwxYfd"
# Pytorch
import torch
import torch.nn as nn
from torch.utils.data import Dataset, DataLoader ##
# For data processing
import numpy as np
import csv
import os
# Plotting
import matplotlib.pyplot as plt
from matplotlib.pyplot import figure
myseed = 42069
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
np.random.seed(myseed)
torch.manual_seed(myseed)
if torch.cuda.is_available():
torch.cuda.manual_seed_all(myseed)
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="DvH3bZXU4xE7" outputId="7fc61e92-1050-4942-fe11-7f2b247c573d"
def get_device():
return 'cuda' if torch.cuda.is_available() else 'cpu'
get_device()
def plot_learning_curve(loss, title=''):
total_steps = len(loss['train'])
def plot_pred(dv_set, model, device, lim=35., preds=None, targets=None):
# + colab={"base_uri": "https://localhost:8080/", "height": 136} id="3JbEXdT440_R" outputId="1743f75a-d216-43a8-eec3-830562063579"
# 给dataset建模一个class
class COVID19Dataset(Dataset):
def __init__(self, path, mode='train', target_only=False):
self.mode = mode
with open(path, 'r') as fp:
data = list(csv.reader(fp)) #$$
data = np.array(data[1:])[:, 1:].astype(float) # 排除第一列之后读取
if not target_only:
feats = list(range(93)) # 全部列的序号
else:
feats = list(range(40))
feats.append(57)
feats.append(75)
if mode == 'test':
# Testing data
# data: 893 x 93 (40 states + day 1 (18) + day 2 (18) + day 3 (17))
data = data[:, feats]
self.data = torch.FloatTensor(data)
else:
# Training data (train/dev sets)
# data: 2700 x 94 (40 states + day 1 (18) + day 2 (18) + day 3 (18))
target = data[:, -1]
data= data[:, feats]
if mode == 'train':
idx = [i for i in range(len(data)) if i % 10 != 0]
elif mode == 'dev':
idx = [i for i in range(len(data)) if i % 10 == 0] # 验证集
# tensors转化
self.data = torch.FloatTensor(data[idx])
self.target = torch.FloatTensor(target[idx])
# Normalize features (you may remove this part to see what will happen)特征标准化 z-score标准化 同一个feature不同人之间
self.data[:, 40:] = \
(self.data[:, 40:] - self.data[:, 40:].mean(dim=0, keepdim=True))/self.data[:, 40:].std(dim=0, keepdim = True)
self.dim = self.data.shape[1] #横向的编号是1 这个方向的维度
print('Finished reading the {} set of COVID19 Dataset ( {} samples found, each dim = {})'
.format(mode, len(self.data), self.dim))
#
def __getitem__(self, index):
if self.mode in ['train', 'dev']:
return
# + id="AFWz2NjjSN6w"
a = list(range(40))
a.append(57)
a.append(75)
#a.append(57).append(75)
# + [markdown] id="7tbw6CvvZBt0"
# **DATALOADER**
# + id="mRgL4zMiZAeZ"
def prep_dataloader(path, mode, batch_size, n_jobs=0, target_only=False):
''' Generates a dataset, then is put into a dataloader. '''
dataset = COVID19Dataset(path, mode=mode, target_only=target_only) # Construct dataset
dataloader = DataLoader(
dataset, batch_size,
shuffle=(mode == 'train'), drop_last=False, # 只在training时shuffle
num_workers=n_jobs, pin_memory=True) # Construct dataloader
return dataloader
# + id="czs9GHoamLz2"
class NeuralNet(nn.Nodule):
def __init__(self, input_dim):
super(NeuralNet, self).__init__() # 利用父类initialization
# Define the Network
self.net = nn.Sequential(
nn.Linear(input_dim, 64),
nn.ReLU(),
nn.Linear(64, 1)
)
# Mean squared error loss
self.criterion = nn.MSELoss(reduction = 'mean') # torch.nn中的功能
def forward(self, x):
return self.net(x).squeeze(1)
def cal_loss(self, pred, target):
return self.criterion(pred, target)
# + [markdown] id="vKM-Rd4hoj13"
# Training
# + colab={"base_uri": "https://localhost:8080/", "height": 136} id="RmgG7Yw3olYw" outputId="4f9fee69-e54d-493c-fe22-f945fefd5910"
def train(tr_set, dv_set, model, config, device): # 注意这里的model作为输入
n_epochs = config['n_epochs'] # 总共的epochs数
optimizer = getattr(torch.optim, config['optimizer'])(
model.parameters(), **config['optim_hparas']) # 优化器 默认SGD
min_mse = 1000.
loss_record = {'train':[], 'dev':[]}
early_stop_cnt = 0
epoch = 0
while epoch < n_epochs:
model.train() # 设置为训练模式
for x, y in tr_set:
optimizer.zero_grad()
x, y = x.to(device), y.to(device) # 将data移动到device中
pred = model(x)
mse_loss = model.cal_loss(pred, y)
mse_loss.backward()
optimizer.step() # update model with optimizer
loss_record['train'].append(mse_loss.detach().cpu().item())
# detach阻断反向传播 也可以用mse_loss().detach().cpu().numpy()
# After each epoch, test your model on the validation (development) set.
dev_mse = dev(dev_set, model, device) # 主义这里的dev函数, 在后面定义
if dev_mse < min_mse:
# 保留改进之后的模型
min_mse = dev_mse
torch.save(model.state_dict(), config['save_path'])
early_stop_cut = 0 # 重新计数
else:
early_stop_cnt += 1
epoch += 1
loss_record['dev'].apppend(dev_mse)
if early_stop_cnt > config['early_stop']: # 很多步没有改进之后,跳出循环
break
print('Finished training after {} epochs'.format(epoch))
return min_mse, loss_record # 优化过程中最小的损失函数值 and 训练中loss_function的值变化
# + [markdown] id="Q8_YwSTO6b8D"
# Validation
# + id="TTK4O2Lh6ZEW"
def dev(dv_set, model, device):
model.eval() # 调整到evaluation模式
total_loss = 0
for x, y in dv_set:
x, y = x.to(device), y.to(device)
with torch.no_grad(): # 不计算梯度
pred = model(x)
mse_loss = model.cal_loss(pred,y)
total_loss += mse_loss.detach().cpu().item()
# + id="t82tqu-co2L7"
# 一系列超参数
device = get_device() # get the current available device ('cpu' or 'cuda')
os.makedirs('models', exist_ok=True) # The trained model will be saved to ./models/
target_only = False # TODO: Using 40 states & 2 tested_positive features
# TODO: How to tune these hyper-parameters to improve your model's performance?
config = {
'n_epochs': 3000, # maximum number of epochs
'batch_size': 270, # mini-batch size for dataloader
'optimizer': 'SGD', # optimization algorithm (optimizer in torch.optim)
'optim_hparas': { # hyper-parameters for the optimizer (depends on which optimizer you are using)
'lr': 0.001, # learning rate of SGD
'momentum': 0.9 # momentum for SGD
},
'early_stop': 200, # early stopping epochs (the number epochs since your model's last improvement)
'save_path': 'models/model.pth' # your model will be saved here
}
| Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# +
# %%capture
import numpy as np
import math
import random
import os
import cv2
import glob
import tensorflow as tf
from tensorflow.keras.callbacks import *
from tensorflow.keras.optimizers import *
from tensorflow.keras.models import Model
from tensorflow.keras.layers import *
from tensorflow.keras import backend as K
import tensorflow_addons as tfa
from dd_data_generator import DDDataGenerator
import sys
sys.path.insert(1, '../')
from constants import *
# -
# # Create DD-Net model
batch_size = 32
# + code_folding=[0]
## triu
def get_triu_indicies(batch_size, num_frames, num_joints):
triu_idxs = np.array(np.triu_indices(num_joints))
num_member = len(triu_idxs[0])
triu_idxs = np.tile(triu_idxs, batch_size*num_frames).transpose()
grid = np.mgrid[0:batch_size,0:NUM_FRAME_SAMPLES,0:num_member].reshape(3,-1).transpose()
return np.concatenate([grid, triu_idxs], axis=1)[:,[0,1,3,4]]
# Half of distance matrix indicies.
GATHER_IDXS_POSE = get_triu_indicies(batch_size, num_frames=NUM_FRAME_SAMPLES, num_joints=NUM_SELECTED_POSENET_JOINTS)
GATHER_IDXS_FACE = get_triu_indicies(batch_size, num_frames=NUM_FRAME_SAMPLES, num_joints=NUM_SELECTED_FACE_JOINTS)
GATHER_IDXS_HAND = get_triu_indicies(batch_size, num_frames=NUM_FRAME_SAMPLES, num_joints=NUM_HAND_JOINTS)
# + code_folding=[0]
## utils.
def get_JCD(frames_batched, gather_idxs):
"""
Get batched half-distance matrix.
"""
# distance matrix.
d_m = batch_frames_cdist(frames_batched, frames_batched)
# gather only upper-right half of distance matrix.
d_m = tf.gather_nd(d_m, gather_idxs, batch_dims=0)
d_m = tf.reshape(d_m, (batch_size, NUM_FRAME_SAMPLES, -1))
return d_m
def batch_frames_cdist(a, b):
return tf.sqrt(tf.reduce_sum(tf.square(tf.expand_dims(a, 2) - tf.expand_dims(b, 3)), axis=-1))
def pose_motion(raw_poses):
diff_slow = poses_diff(raw_poses)
# flatten last 2 dims.
diff_slow = tf.reshape(diff_slow, (-1, diff_slow.shape[1], diff_slow.shape[2]*diff_slow.shape[3]))
# jump frame
fast = raw_poses[:, fd00:a516:7c1b:17cd:6d81:2137:bd2a:2c5b, :, :]
diff_fast = poses_diff(fast)
# flatten last 2 dims.
diff_fast = tf.reshape(diff_fast, (-1, diff_fast.shape[1], diff_fast.shape[2]*diff_fast.shape[3]))
return diff_slow, diff_fast
def poses_diff(x):
# frame t - frame(t-1)
x = x[:, 1:, :, :] - x[:, :-1, :, :]
x_d = tf.expand_dims(x[:, 0, :, :], 1)
x_d = tf.concat([x_d, x], axis=1)
return x_d
# + code_folding=[0]
## nural blocks.
def c1D(x, filters, kernel):
x = Conv1D(filters, kernel_size=kernel, padding='same', use_bias=False)(x)
x = BatchNormalization()(x)
x = PReLU(shared_axes=[1])(x)
return x
def block(x, filters):
x = c1D(x, filters, 3)
x = c1D(x, filters, 3)
return x
def d1D(x, filters):
x = Dense(filters, use_bias=False)(x)
x = BatchNormalization()(x)
x = PReLU(shared_axes=[1])(x)
return x
#pose encoder
def encode_jcds(x, filters, drop_out=0.4):
x = c1D(x, filters*2, 1)
x = SpatialDropout1D(0.1)(x)
x = c1D(x, filters, 3)
x = SpatialDropout1D(0.1)(x)
x = c1D(x, filters, 1)
x = MaxPooling1D(2)(x)
x = SpatialDropout1D(0.1)(x)
x = Dropout(drop_out)(x)
return x
# hands encoder
def joints_encoder(filters, drop_out=0.4):
encoder_input = Input(shape=(NUM_FRAME_SAMPLES, 231))
x = c1D(encoder_input, filters*2, 1)
x = SpatialDropout1D(0.1)(x)
x = c1D(x, filters, 3)
x = SpatialDropout1D(0.1)(x)
x = c1D(x, filters, 1)
x = MaxPooling1D(2)(x)
x = SpatialDropout1D(0.1)(x)
x = Dropout(drop_out)(x)
return Model(inputs=encoder_input, outputs=x)
def encode_diff_slow(diff_slow, filters):
x_d_slow = c1D(diff_slow, filters*2, 1)
x_d_slow = SpatialDropout1D(0.1)(x_d_slow)
x_d_slow = c1D(x_d_slow, filters, 3)
x_d_slow = SpatialDropout1D(0.1)(x_d_slow)
x_d_slow = c1D(x_d_slow, filters, 1)
x_d_slow = MaxPool1D(2)(x_d_slow)
x_d_slow = SpatialDropout1D(0.1)(x_d_slow)
return x_d_slow
def encode_diff_fast(diff_fast, filters):
x_d_fast = c1D(diff_fast, filters*2, 1)
x_d_fast = SpatialDropout1D(0.1)(x_d_fast)
x_d_fast = c1D(x_d_fast, filters, 3)
x_d_fast = SpatialDropout1D(0.1)(x_d_fast)
x_d_fast = c1D(x_d_fast, filters, 1)
x_d_fast = SpatialDropout1D(0.1)(x_d_fast)
return x_d_fast
# + code_folding=[0]
## backbone
def build_backbone(
pose_frames,
diff_slow, diff_fast,
face_frames,
left_hand_frames, right_hand_frames,
filters):
# pose
pose_encoded = encode_jcds(pose_frames, filters//2, drop_out=0.4)
pose_encoded = c1D(pose_encoded, 256, 3)
pose_encoded = MaxPooling1D(4)(pose_encoded)
pose_encoded = c1D(pose_encoded, 256, 3)
pose_encoded = MaxPooling1D(2)(pose_encoded)
pose_encoded= Flatten()(pose_encoded)
pose_encoded = Dense(256)(pose_encoded)
pose_encoded = tf.math.l2_normalize(pose_encoded, axis=-1)
# face
face_encoded = encode_jcds(face_frames, filters//2, drop_out=0.3)
face_encoded = c1D(face_encoded, 128, 3)
face_encoded = MaxPooling1D(4)(face_encoded)
face_encoded = c1D(face_encoded, 128, 3)
face_encoded = MaxPooling1D(2)(face_encoded)
face_encoded= Flatten()(face_encoded)
face_encoded = Dense(64)(face_encoded)
face_encoded = tf.math.l2_normalize(face_encoded, axis=-1)
# hands
diff_slow_encoded = encode_diff_slow(diff_slow, filters)
diff_fast_encoded = encode_diff_fast(diff_fast, filters)
hand_encoder = joints_encoder(int(filters*4), drop_out=0.4)
left_hands_encoded = hand_encoder(left_hand_frames)
right_hands_encoded = hand_encoder(right_hand_frames)
hands = concatenate([diff_slow_encoded, diff_fast_encoded, left_hands_encoded, right_hands_encoded])
hands = c1D(hands, 256, 3)
hands = MaxPooling1D(4)(hands)
hands = c1D(hands, 512, 3)
hands = MaxPooling1D(2)(hands)
hands = Flatten()(hands)
hands = Dense(512)(hands)
hands = tf.math.l2_normalize(hands, axis=-1)
# all feats
x = concatenate([pose_encoded, face_encoded, hands])
return x
# + code_folding=[0]
## build.
def build_DD_Net():
# input layers.
pose_frames_input = Input(shape=(NUM_FRAME_SAMPLES, NUM_SELECTED_POSENET_JOINTS, POSENET_JOINT_DIMS), name='pose_frames_input')
face_frames_input = Input(shape=(NUM_FRAME_SAMPLES, NUM_SELECTED_FACE_JOINTS, FACE_JOINT_DIMS), name='face_frames_input')
left_hand_frames_input = Input(shape=(NUM_FRAME_SAMPLES, NUM_HAND_JOINTS, HAND_JOINT_DIMS), name='left_hand_frames_input')
right_hand_frames_input = Input(shape=(NUM_FRAME_SAMPLES, NUM_HAND_JOINTS, HAND_JOINT_DIMS), name='right_hand_frames_input')
# poses.
pose_jcd = get_JCD(pose_frames_input, GATHER_IDXS_POSE)
# faces.
face_jcd = get_JCD(face_frames_input, GATHER_IDXS_FACE)
# hands.
left_hand_jcd = get_JCD(left_hand_frames_input, GATHER_IDXS_HAND)
right_hand_jcd = get_JCD(right_hand_frames_input, GATHER_IDXS_HAND)
# hands diff.
hand_cat = concatenate([left_hand_frames_input, right_hand_frames_input], axis=-2)
diff_slow, diff_fast = pose_motion(hand_cat)
# embed and backbone.
x = build_backbone(pose_jcd,
diff_slow, diff_fast,
face_jcd,
left_hand_jcd, right_hand_jcd,
filters=NUM_START_FILTERS)
model = Model(inputs=[pose_frames_input, face_frames_input, left_hand_frames_input, right_hand_frames_input], outputs=x)
return model
# + code_folding=[0]
## input
pose_frames_input = Input(batch_shape=(batch_size, NUM_FRAME_SAMPLES, NUM_SELECTED_POSENET_JOINTS, POSENET_JOINT_DIMS), name='pose_frames_input')
face_frames_input = Input(batch_shape=(batch_size, NUM_FRAME_SAMPLES, NUM_SELECTED_FACE_JOINTS, FACE_JOINT_DIMS), name='face_frames_input')
left_hand_frames_input = Input(batch_shape=(batch_size, NUM_FRAME_SAMPLES, NUM_HAND_JOINTS, HAND_JOINT_DIMS), name='left_hand_frames_input')
right_hand_frames_input = Input(batch_shape=(batch_size, NUM_FRAME_SAMPLES, NUM_HAND_JOINTS, HAND_JOINT_DIMS), name='right_hand_frames_input')
embedder_model = build_DD_Net()
# embed.
feats_out = embedder_model([pose_frames_input, face_frames_input, left_hand_frames_input, right_hand_frames_input])
cls_out = Dense(NUM_CLASSES, activation="softmax", name="cls_out")(feats_out)
model = Model(inputs=[pose_frames_input, face_frames_input,
left_hand_frames_input, right_hand_frames_input],
outputs=[feats_out,cls_out])
# -
model.summary()
# # Data Generator
# + code_folding=[]
batch_size = 32
train_generator = DDDataGenerator('my_preprocessed_dataset/train',
batch_size=batch_size, use_augment=True)
val_generator = DDDataGenerator('my_preprocessed_dataset/val',
batch_size=batch_size, use_augment=True)
# -
# ## Train
model.compile(loss=tfa.losses.triplet_hard_loss, optimizer="Adam")
# +
filepath = ("checkpoints/{epoch:02d}-{loss:.4f}.h5")
checkpoint = ModelCheckpoint(filepath, monitor='loss', save_best_only=False,
mode='auto', save_weights_only=False)
history = model.fit_generator(train_generator,
steps_per_epoch=200,
epochs=200,
initial_epoch=0,
callbacks=[checkpoint],
validation_data=val_generator,
validation_steps=500,
validation_freq=3,
workers=3, use_multiprocessing=False
)
# -
| classifier/train_dd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Nonhardening Plasticity
#
# ## Overview
#
# In contrast to elasticity, plasticity describes the deformation of bodies that undergo nonreversible changes of shape in response to applied load. When a body transitions from elastic to plastic deformation it is said to "yield". The yield transition point, also known as the **yield strength** is a property of the material and generally changes in response to continued loading. When the yield strength increases with continued loading it is said to *harden* and the response is described by hardening theories of plasticity. The simplified nonhardening theory of plasticity describes materials whose yield strength does not change in response to continued loading. This notebook
#
# - presents an introduction to nonhardening plasticity,
# - implements a nonhardening "$J_2$" plasticity model in Matmodlab, and
# - verifies the $J_2$ model against analytic solutions.
# ## See Also
#
# - [User Defined Materials: Introduction](UserMaterials.ipynb)
# - [Linear Elastic Material](LinearElastic.ipynb)
# <a name='contents'></a>
# ## Contents
#
# 1. <a href='#plastic'>Fundamental Equations</a>
# 2. <a href='#j2'>$J_2$ Plasticity</a>
# 3. <a href='#umat.std'>Standard Model Implementation</a>
# 4. <a href='#umat.ver'>Model Verification</a>
# <a name='plastic'></a>
# ## Fundamental Equations
#
# The mechanical response of a nonhardening plastic material is predicated on the assumption that there exists an **elastic limit**, beyond which stress states are not achievable through normal processes. The elastic limit is defined by a **yield surface** satisfying
#
# $$
# f\left(\pmb{\sigma}\right) = 0
# $$
#
# where $f$ is the **yield function** and $\pmb{\sigma}$ the mechanical stress, defined as
#
# $$
# \dot{\pmb{\sigma}} = \mathbb{C}{:}\dot{\pmb{\epsilon}}^{\rm e}
# $$
#
# where $\mathbb{C}$ is the elastic stiffness and $\dot{\pmb{\epsilon}}^{\rm e}$ the rate of elastic strain. The rate of strain is commonly regarded as the sum of elastic and plastic parts, giving for the mechanical response
#
# $$
# \dot{\pmb{\sigma}} = \mathbb{C}{:}\left(\dot{\pmb{\epsilon}} - \dot{\pmb{\epsilon}}^{\rm p}\right)
# $$
#
# where $\dot{\pmb{\epsilon}}^{\rm p}$ is the rate of plastic strain. Replacing $\dot{\pmb{\epsilon}}^{\rm p}$ with $\dot{\lambda}\pmb{m}$, $\dot{\lambda}$ being the magnitude of $\dot{\pmb{\epsilon}}^{\rm p}$ and $\pmb{m}$ its direction, the mechanical response of the material is
#
# $$
# \dot{\pmb{\sigma}} = \mathbb{C}{:}\left(\dot{\pmb{\epsilon}} - \dot{\lambda}\pmb{m}\right)
# $$
#
# The solution to the plasticity problem is reduced to determining $\dot{\lambda}$ such that $f\left(\pmb{\sigma}(t)\right)\leq 0 \ \forall t>0$
# ### Solution Process
#
# Given the current state of stress $\pmb{\sigma}_n$, the solution to the plasticity problem begins with the hypothesis that the entire strain increment is elastic:
#
# $$
# \pmb{\sigma}_{n+1} \stackrel{?}{=} \pmb{\sigma}_{n} + \mathbb{C}{:}\dot{\pmb{\epsilon}}dt = \pmb{\sigma}^{\rm test}
# $$
#
# where the descriptor "test" is used to signal the fact that at this point $\pmb{\sigma}^{\rm test}$ is merely a hypothesized solution. The hypothesis is validated if $\pmb{\sigma}^{\rm test}$ satisfies the **yield condition**
#
# $$f\left(\pmb{\sigma}^{\rm test}\right)\leq 0$$
#
# so that $\pmb{\sigma}_{n+1}=\pmb{\sigma}^{\rm test}$.
#
# If instead the hypothesis is *falsefied*, i.e., the predicted test stress falls outside of the yield surface defined by $f=0$, the plasticity problem,
#
# $$\begin{align}
# \pmb{\sigma}_{n+1} = \pmb{\sigma}_{n} + \mathbb{C}{:}\left(\dot{\pmb{\epsilon}} - \dot{\lambda}\pmb{m}\right)dt &= \pmb{\sigma}^{\rm trial} - \dot{\lambda}\pmb{A}dt\\
# f\left(\pmb{\sigma}^{\rm trial} - \dot{\lambda}\pmb{A}dt\right) &= 0
# \end{align}$$
#
# where $\pmb{A}=\mathbb{C}{:}\pmb{m}$, is solved. $\dot{\pmb{\sigma}}^{\rm trial}=\mathbb{C}{:}\dot{\pmb{\epsilon}}$ is distinguished from $\dot{\pmb{\sigma}}^{\rm test}$ in that for stress driven problems $\dot{\pmb{\sigma}}^{\rm trial}$ is not necessarily known because the strain rates $\dot{\epsilon}$ are not known.
#
# The unknown scalar $\dot{\lambda}$ is determined by noting the following observation: if $f\left(\pmb{\sigma}_{n}\right)=0$ and, after continued loading, $f\left(\pmb{\sigma}_{n+1}\right)=0$, the rate of change of $f$ itself must be zero. Thus, by the chain rule,
#
# $$\begin{align}
# \dot{f}{\left(\pmb{\sigma}\right)}
# &=\frac{df}{d\pmb{\sigma}}{:}\dot{\pmb{\sigma}}\\
# &=\frac{df}{d\pmb{\sigma}}{:}\left(\mathbb{C}{:}\dot{\epsilon}-\dot{\lambda}\pmb{A}dt\right)=0
# \end{align}$$
#
# which is known as the **consistency condition** and $\dot{\lambda}$ the **consistency parameter**. The consistency condition is equivalent to the statement that $\dot{\lambda}\dot{f}=0$. The consistency conditions can be shown to be equivalent to the Karush-Kuhn-Tucker conditions
#
# $$
# \dot{\lambda} \ge 0 ~,~~ f \le 0~,~~ \dot{\lambda}\,f = 0 \,.
# $$
# Letting
#
# $$
# \pmb{n} = \frac{df}{d\pmb{\sigma}}\Big/\Big\lVert\frac{df}{d\pmb{\sigma}}\Big\rVert
# $$
#
# the preceding equation can be solved $\dot{\lambda}$, giving
#
# $$
# \dot{\lambda}
# = \frac{\pmb{n}{:}\mathbb{C}{:}\dot{\epsilon}}{\pmb{n}{:}\pmb{A}}
# $$
#
# Substituting $\dot{\lambda}$ in to the expression for stress rate gives
#
# $$\begin{align}
# \dot{\pmb{\sigma}}
# &= \mathbb{C}{:}\dot{\pmb{\epsilon}} - \frac{\pmb{n}{:}\mathbb{C}{:}\dot{\epsilon}}{\pmb{n}{:}\pmb{A}}\pmb{A}\\
# &= \mathbb{C}{:}\dot{\pmb{\epsilon}} - \frac{1}{\pmb{n}{:}\pmb{A}}\pmb{Q}\pmb{A}{:}\dot{\pmb{\epsilon}}\\
# &= \mathbb{D}{:}\dot{\pmb{\epsilon}}
# \end{align}$$
#
# where
#
# $$
# \mathbb{D} = \mathbb{C} - \frac{1}{\pmb{n}{:}\pmb{A}}\pmb{Q}\pmb{A}
# $$
#
# The stress rate is then integrated through time to determine $\pmb{\sigma}$
# ### Integration Procedure
#
# It can be shown that there exists a scalar $\Gamma$ such that
#
# $$\pmb{\sigma}_{n+1} = \pmb{\sigma}^{\rm trial} - \Gamma\pmb{A}$$
#
# $\Gamma$ is determined by satisfying the yield condition. In other words, $\Gamma$ is the solution to
#
# $$f\left(\pmb{\sigma}^{\rm trial} - \Gamma\pmb{A}\right)=0$$
#
# The unknown $\Gamma$ is found such that $f\left(\pmb{\sigma}(\Gamma)\right)=0$. The solution can be found by solving the preceding equation iteratively by applying Newton's method so that
#
# $$
# \Gamma^{i+1} = \Gamma^i + \frac{f\left(\pmb{\sigma}(\Gamma^{n})\right)}{(df/d\pmb{\sigma})\big|_{\Gamma^i}{:}\pmb{A}}
# $$
#
# When $\Gamma^{i+1}-\Gamma^i<\epsilon$, where $\epsilon$ is a small number, the iterations are complete and the updated stress can be determined.
#
# Note that the scalar $\Gamma$ is also equal to $\Gamma=\dot{\lambda}dt$, but since $\dot{\lambda}=0$ for elastic loading, $\Gamma=\dot{\lambda}dt^p$, where $dt^p$ is the plastic part of the time step. This gives $\Gamma$ the following physical interpretation: it is the magnigute of the total plastic strain increment.
# #### Alternative Iterative method
#
# Brannon argues that redefining $\Gamma$ such that
#
# $$\pmb{\sigma}_{n+1} = \pmb{\sigma}^{\rm trial} + \Gamma\pmb{A}$$
#
# can lead to fast convergence for yield surfaces with considerable curvature [2002, Radial Return document]. To update the stress, begin Newton iterations with $\Gamma=0$ and compute the improved estimate as
#
# $$
# \Gamma^{i+1} = -\frac{f\left(\pmb{\sigma}(\Gamma^{i})\right)}{(df/d\pmb{\sigma})\big|_{\Gamma^i}{:}\pmb{A}}
# $$
#
# when $\Gamma^{i+1}<\epsilon$ the Newton procedure is complete.
# ## $J_2$ Plasticity
#
# The equations developed thus far are general in the sense that they apply to any material that can be modeled by nonhardening plasticity. The equations are now specialized to the case of isotropic hypoelasticity and $J_2$ plasticity by defining
#
# $$\begin{align}
# \dot{\pmb{\sigma}} &= 3K\,\mathrm{iso}\dot{\pmb{\epsilon}}^{\rm e} + 2G\,\mathrm{dev}\dot{\pmb{\epsilon}}^{\rm e} \\
# f\left(\pmb{\sigma}\right) &= \sqrt{J_2} - k
# \end{align}
# $$
#
# where $K$ and $G$ are the bulk and shear moduli, respectively, and $J_2$ is the second invariant of the stress deviator, defined as
#
# $$J_2 = \frac{1}{2}\pmb{s}{:}\pmb{s}, \quad \pmb{s} = \pmb{\sigma} - \frac{1}{3}\mathrm{tr}\left(\pmb{\sigma}\right)\pmb{I}$$
#
# Additionally, we adopt the assumption of an **associative flow rule** wherein $\pmb{m}=\pmb{n}$. Accordingly,
#
# $$\begin{align}
# \frac{df}{d\pmb{\sigma}}&=\frac{1}{2\sqrt{J_2}}\pmb{s}, &\pmb{n}=\frac{1}{\sqrt{2J_2}}\pmb{s} \\
# \pmb{A}&=\frac{2G}{\sqrt{2J_2}}\pmb{s}, &\pmb{Q}=\frac{2G}{\sqrt{2J_2}}\pmb{s}
# \end{align}$$
#
# ### Required Parameters
#
# The model as described above requires at minimum 3 parameters: 2 independent elastic moduli and a yield strength measure. Commonly, the yield strength in tension $Y$ is chosen in lieu of the yield strength in shear $k$. The conversion from $Y$ to $k$, as required by the model development, is found from evaluating the yield criterion for the case of uniaxial tension, where the stress and its deviator are
#
# $$
# \pmb{\sigma} = \begin{bmatrix}\sigma_{\rm ax}&&\\& 0&\\&& 0\end{bmatrix}, \quad
# \pmb{s} = \frac{1}{3}\begin{bmatrix}2\sigma_{\rm ax}&&\\& -\sigma_{\rm ax}&\\&& -\sigma_{\rm ax}\end{bmatrix}
# $$
#
# Accordingly,
#
# $$J_2 = \frac{1}{2}\pmb{s}{:}\pmb{s} = \frac{1}{3}\sigma_{\rm ax}^2$$
#
# evaluating the yield function leads to
#
# $$\sqrt{J_2} = \frac{1}{\sqrt{3}}Y = k$$
#
# ### Optional Solution Dependent Variables
#
# The simple $J_2$ plasticity model described does not require the storage and tracking of solution dependent variables. However, the equivalent plastic strain is often stored for output purposes. The equivalent plastic strain is defined so that the plastic work
#
# $$W^p = \pmb{\sigma}{:}\dot{\pmb{\epsilon}}^p=\sigma_{eq}\dot{\epsilon}^p_{eq}$$
#
# where $\sigma_{eq}$ is the equivalent von Mises stress $\sigma_{eq} = \sqrt{\frac{3}{2}\pmb{s}{:}\pmb{s}}$, requiring that
#
# $$\dot{\epsilon}^p_{eq} = \sqrt{\frac{2}{3}}\lVert{\rm dev}\left(\dot{\pmb{\epsilon}}^p\right)\rVert$$
# <a name='umat.std'></a>
# ## Model Implementation
#
# The plastic material described above is implemented as `NonhardeningPlasticMaterial` in `matmodlab2/materials/plastic2.py`. `NonhardeningPlasticMaterial` is implemented as a subclass of the `matmodlab2.core.material.Material` class. `NonhardeningPlasticMaterial` defines
#
# - `name`: *class attribute*
#
# Used for referencing the material model in the `MaterialPointSimulator`.
#
# - `eval`: *instance method*
#
# Updates the material stress, stiffness (optional), and state dependent variables to the end of the time increment.
#
# In the example below, in addition to some standard functions imported from `Numpy`, several helper functions are imported from various locations in Matmodlab:
#
# - `matmodlab.utils.tensor`
#
# - `iso`, `dev`: computes the isotropic and deviatoric parts of a second-order tensor
# - `mag`: computes the magnitude $\left(\lVert x_{ij} \rVert=\sqrt{x_{ij}x_{ij}}\right)$ of a second-order tensors
# - `ddot`, `dyad`: computes the double dot product and dyadic product of second-order tensors
# - `matmodlab.constants`
# - `VOIGT`: mulitplier for converting tensor strain components to engineering strain components
# - `ROOT3`: $\sqrt{3}$
# - `TOLER`: A small number, used as a tolerance
#
#
# ### A Note on Implementation
#
# For the simplistic $J_2$ plasticity model, it is a simple task to determine the analytic response and avoid Newton iterations. In the model implemented, Newton iterations are used to verify the algorithm. The source of the file can be viewed by executing the following cell.
# %pycat ../matmodlab2/materials/plastic3.py
# %pylab inline
from matmodlab2 import *
import pandas as pd
pd.set_option('precision', 18)
# <a name='umat.ver'></a>
# ## Verification
#
# Exercising the elastic model through a path of uniaxial stress should result in the slope of axial stress vs. axial strain being equal to the input parameter `E` for the elastic portion. The maximum stress should be equal to the input parameter `Y`.
# +
E = 10e6
Nu = .1
Y = 40e3
K = E / (3. * (1. - 2. * Nu))
G = E / 2.0 / (1. + Nu)
#mps1 = MaterialPointSimulator('uplastic')
#mps1.material = NonhardeningPlasticMaterial(E=E, Nu=Nu, Y=Y)
#mps1.run_step('EEE', (.02, 0, 0), frames=50)
mps2 = MaterialPointSimulator('Job-2')
mps2.material = HardeningPlasticMaterial(E=E, Nu=Nu, Y0=Y)
#mps2.run_step('EEE', (.005, 0, 0), frames=100)
#mps1.df.plot('E.XX', 'S.XX')
#mps2.df.plot('E.XX', 'S.XX')
#print(mps2.df.keys())
#mps2.df[['F.XX', 'S.XX', 'S.YY', 'EP_Equiv']]
time = 0
dtime = 1
temp = 0
dtemp = 0
F0 = eye(3).flatten()
F1 = eye(3).flatten()
d = zeros(6)
stran = zeros(6)
X = zeros(10)
stress=zeros(6)
t,x,d = mps2.material.eval(time, dtime, temp, dtemp, F0, F1,
stran, d, stress, X)
t
# +
def gen_uniax_strain_path(Y, YF, G, LAM):
EPSY = Y / (2.0 * G) # axial strain at yield
SIGAY = (2.0 * G + LAM) * EPSY # axial stress at yield
SIGLY = LAM * EPSY # lateral stress at yield
EPSY2 = 2.0 * Y / (2.0 * G) # final axial strain
SIGAY2 = ((2 * G + 3 * LAM) * EPSY2 - YF ) / 3.0 + YF # final axial stress
SIGLY2 = ((2 * G + 3 * LAM) * EPSY2 - YF ) / 3.0 # final lateral stress
return [[EPSY, 0.0, 0.0], [EPSY2, 0.0, 0.0]]
def copper_params():
E = 0.1100000E+12
NU = 0.3400000
LAM = 0.87220149253731343283582089552238805970149253731343E+11
G = 0.41044776119402985074626865671641791044776119402985E+11
K = 0.11458333333333333333333333333333333333333333333333E+12
USM = 0.16930970149253731343283582089552238805970149253731E+12
Y = 70.0E+6
return NU, E, K, G, LAM, Y
def test_j2_1():
jobid = 'j2_plast'
mps = MaterialPointSimulator(jobid)
NU, E, K, G, LAM, Y = copper_params()
YF, H, BETA = Y, 0, 0
parameters = {'K': K, 'G': G, 'Y0': YF, 'H': H, 'BETA': 0}
print(E, NU, YF)
material = VonMisesMaterial(**parameters)
mps.assign_material(material)
path = gen_uniax_strain_path(Y, YF, G, LAM)
for row in path:
mps.run_step('E', row, increment=1.0, frames=25)
mps.plot('E.XX', 'S.XX')
print(mps.df[['F.XX', 'S.XX', 'S.YY']])
print(mps.df['EQPS'])
print(mps.df.keys())
test_j2_1()
# -
print(mps)
| notebooks/NonhardeningJ2Plasticity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Stop words
#
# ## What are stop words?
#
# A stop word is a word that has very little meaning by itself, such as `the`,`a`, `and`, `an`,...
# Most search engines remove these "stop words" when you do a search.
#
# 
#
# ## How to remove these stop words?
#
# You could remove them by hand with the `replace()` function, but if you want to go faster, you can use libraries like `SpaCy`, `NLTK`, `Gensim`, and more. Each library will behave slightly differently, but not enough to make big changes to your model.
#
# ## Practice time!
#
# Using the library of your choice, remove all the stop words of this text:
# +
# Remove all my stop words
text = "At BeCode, we like to learn. Sometime, we play games not win a price but to have fun!"
# You can use any library!
# -
# The result should be something like this:
# ```
# ['BeCode', ',', 'like', 'learn', '.', ',', 'play', 'games', 'win', 'price', 'fun', '!']
# ```
#
# So as you can see, depending on what kind of information you want to extract, you will be able to exclude stop words. For document classification or semantic search, you will not need those stop words for example.
#
# ## Customize your stop words
#
# You can also add or remove stop words from the list that the libraries uses for stop words. If there is a specific word in your document that should not be considered as a stop word, or one that should absolutely be given to the model, you can do so.
#
#
# ## Additional resources
# * [NLP Essentials: Removing stopwords and performing Text Normalization using NLTK and spaCy in Python](https://www.analyticsvidhya.com/blog/2019/08/how-to-remove-stopwords-text-normalization-nltk-spacy-gensim-python/)
# * [Removing stop words from strings in Python](https://stackabuse.com/removing-stop-words-from-strings-in-python/#usingpythonsnltklibrary)
# * [Dropping common terms: stop words](https://nlp.stanford.edu/IR-book/html/htmledition/dropping-common-terms-stop-words-1.html)
| Content/2.basic_preprocessing/3.stop_words.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:devil]
# language: python
# name: conda-env-devil-py
# ---
# # Predicting the Output Electrical Power of Combined Cycle Power Plant with the help of 6 Years(2006-2011) data of the plant.
# ## Authour : <NAME>
# ## Importing revelent packages
import tensorflow as tf
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# ## Importing Datasets
raw_data = pd.read_excel('Combined Cycle Power Plant Dataset.xlsx')
data = raw_data.copy()
data.head()
# ## Data Preprocessing
data.rename(columns={'AT':'Temp', 'V':'Exhaust_Vaccum', 'AP':'Pressure', 'RH':'Humidity', 'PE':'Power_Output'})
data.shape
x = data.iloc[:,:-1].values
y = data.iloc[:, -1].values
from sklearn.model_selection import train_test_split as tts
x_train, x_test, y_train, y_test = tts(x, y, random_state = 143, test_size=0.2)
# ## Feature Scaling
from sklearn.preprocessing import StandardScaler
ss = StandardScaler()
x_train = ss.fit_transform(x_train)
x_test = ss.transform(x_test)
# ## Buliding the Neural Network
ann_r = tf.keras.models.Sequential()
ann_r.add(tf.keras.layers.Dense(units=50, activation = 'relu'))
ann_r.add(tf.keras.layers.Dense(units=50, activation = 'relu'))
ann_r.add(tf.keras.layers.Dense(units=50, activation = 'relu'))
ann_r.add(tf.keras.layers.Dense(units=1))
ann_r.compile(optimizer='adam', loss='MeanSquaredError')
ann_r.fit(x_train, y_train, batch_size=32, epochs=100)
ann_r.evaluate(x_train, y_train)
# ## Testing our network on unseen test dataset
y_pred = ann_r.predict(x_test)
# ### Comparing the predicted and actual outputs side-by-side
Evaluate = pd.DataFrame(data = y_pred, columns=['Predicted_Values'])
Evaluate['Targets'] = y_test
Evaluate
from sklearn.metrics import *
r2_score(y_pred, y_test)
# # Our Artifical Brain is 94% accurate on the unseen dataset
| Predict Power Output.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <iframe width="1000" height="1000" align="center" src="https://www.falstad.com/qmatom/" frameborder="0" allow="accelerometer; autoplay; encrypted-media; gyroscope; picture-in-picture" allowfullscreen></iframe>
#
#
#
| CHEM1035_Chemistry_For_Engineers/.ipynb_checkpoints/test-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# VEZBE 2.
# +
#7. zadatak
#Za funkciju prenosa G(s) modela motora jednosmerne struje nacrtati impulsne i odskocne odzive
#sistema za Km=1 Tm1=1, Tm2=5 i Tm3=10.
#G(s)= Km/(Tms+1)
# -
import numpy as np;
import scipy as sp;
import matplotlib.pyplot as plt
from scipy import signal
# +
#Km=1
Gs1=signal.lti([1],[1,1]) #Tm1=1
Gs2=signal.lti([1],[5,1]) #Tm2=5
Gs3=signal.lti([1],[10,1]) #Tm3=10
# +
#odskocni odziv
t1,u1=Gs1.step()
t2,u2=Gs2.step()
t3,u3=Gs3.step()
plt.title("Одскочни одзив система")
plt.ylabel("брзина $\omega$ [rad/s]")
plt.xlabel("време t [s]")
plt.plot(t1,u1,'r')
plt.plot(t2,u2,'b')
plt.plot(t3,u3,'g')
plt.grid()
plt.show()
#Sa grafika mozemo da zakljucimo da se brzina odziva sistema smanjuje sa povecanjem vremenske konstante
# +
#impulsni odziv
t1,u1=Gs1.impulse()
t2,u2=Gs2.impulse()
t3,u3=Gs3.impulse()
plt.title("Импулсни одзив система")
plt.ylabel("брзина $\omega$ [rad/s]")
plt.xlabel("време t [s]")
plt.plot(t1,u1,'r')
plt.plot(t2,u2,'b')
plt.plot(t3,u3,'g')
plt.grid()
plt.show()
#Sa grafika mozemo da zakljucimo da se brzina odziva sistema smanjuje sa povecanjem vremenske konstante
# +
#8. zadatak
#Neka je data prenosna fja Gs(s) sistema drugog reda: Gs(s) = Wn^2/(s^2+2*c*Wn+Wn^2)
#pri cemu je sa Wn oznacena neprigusena prirodna ucestanost
#a sa c oznacen faktor relativnog prigusenja, c je iz skupa (0,1).
#Na sistem deluje pobuda oblika Hevisajdove funkcije
#a) Za Wn>0 odrediti odziv, oceniti da li je sistem stabilan i odrediti u kojoj poluravni se nalaze polovi sistema, c=0.1
#b) Za Wn<0 odrediti odziv, oceniti da li je sistem stabilan i odrediti u kojoj poluravni se nalaze polovi sistema, c=0.1
# +
#a)
#Prvo unosimo polinom funkcije prenosa u promenljivu Gs4.
c4=0.1
Wn4=1
Gs4=signal.lti([Wn4*Wn4],[1,2*c4*Wn4,Wn4*Wn4])
# +
#Odredjujemo odziv sistema na jedinicnu pobudu (Hevisajdova fja).
t4,u4 = Gs4.step()
plt.plot(t4,u4,"g")
#Jednostavnom proverom zakljucujemo da za priblizno u4[87]=1.00 u t4[87]=61.51 nastaje stacionarno stanje.
plt.axvline(x=61.51, color='r', linestyle='-')
plt.xlim(0,100)
plt.title("Одзив система Gs4(s) на одскочну побуду")
plt.ylabel("Одскочни одзив система y(t)")
plt.xlabel("време t [s]")
plt.grid()
plt.show()
#Zakljucujemo da je za Wn>0 sistem stabilan jer sistem posle odredjenog vremena ulazi u stacionarno stanje.
# +
#Sada proveravamo polozaj polova sistema.
#Smestamo koordinate imaginarnog dela polova u jw4 a realnog u sigma4.
jw4 = Gs4.poles.imag
sigma4 = Gs4.poles.real
# +
#Zatim iscrtavamo polozaj polova u s ravni sistema.
plt.scatter(sigma4,jw4,marker="x")
plt.xlim(-0.5,0.5)
plt.ylim(-1.5,1.5)
plt.axvline(x=0, color='r', linestyle='-')
plt.axhline(y=0, color='r', linestyle='-')
plt.title("s раван система")
plt.xlabel(r'Реална оса $\sigma$')
plt.ylabel(r'Имагинарна оса j$\omega$')
plt.grid()
plt.show()
#Polovi sistema su u levoj s poluravni, sto je ujedno i uslov da sistem bude stabilan.
# -
#b)
c5=0.1
Wn5=-1
Gs5=signal.lti([Wn5*Wn5],[1,2*c5*Wn5,Wn5*Wn5])
# +
t5,u5 = Gs5.step()
plt.plot(t5,u5,"r")
plt.xlim(0,100)
plt.title("Одзив система Gs5(s) на одскочну побуду")
plt.ylabel("Одскочни одзив система y(t)")
plt.xlabel("време t [s]")
plt.grid()
plt.show()
plt.show()
#Kako se sa grafika odziva funkcije moze videti intuitivno zakljucujemo da je sistem
#nestabilan i da sistem nikada nece izaci iz prelaznog stanja.
#Zakljucujemo da je za Wn<0 sistem nestabilan.
# -
jw5 = Gs5.poles.imag
sigma5 = Gs5.poles.real
# +
plt.scatter(sigma5,jw5,marker="x")
plt.xlim(-0.5,0.5)
plt.ylim(-1.5,1.5)
plt.axvline(x=0, color='r', linestyle='-')
plt.axhline(y=0, color='r', linestyle='-')
plt.title("s раван система")
plt.xlabel(r'Реална оса $\sigma$')
plt.ylabel(r'Имагинарна оса j$\omega$')
plt.grid()
plt.show()
#Polozaj polova ukazuje na cinjenicu da je sistem nestabilan jer se polovi nalaze
#u desnoj s poluravni.
| Jupyter/Exercise 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# $
# % START OF MACRO DEF
# % DO NOT EDIT IN INDIVIDUAL NOTEBOOKS, BUT IN macros.py
# %
# \newcommand{\Reals}{\mathbb{R}}
# \newcommand{\Expect}[0]{\mathbb{E}}
# \newcommand{\NormDist}{\mathcal{N}}
# %
# \newcommand{\DynMod}[0]{\mathscr{M}}
# \newcommand{\ObsMod}[0]{\mathscr{H}}
# %
# \newcommand{\mat}[1]{{\mathbf{{#1}}}}
# %\newcommand{\mat}[1]{{\pmb{\mathsf{#1}}}}
# \newcommand{\bvec}[1]{{\mathbf{#1}}}
# %
# \newcommand{\trsign}{{\mathsf{T}}}
# \newcommand{\tr}{^{\trsign}}
# \newcommand{\tn}[1]{#1}
# \newcommand{\ceq}[0]{\mathrel{≔}}
# %
# \newcommand{\I}[0]{\mat{I}}
# \newcommand{\K}[0]{\mat{K}}
# \newcommand{\bP}[0]{\mat{P}}
# \newcommand{\bH}[0]{\mat{H}}
# \newcommand{\bF}[0]{\mat{F}}
# \newcommand{\R}[0]{\mat{R}}
# \newcommand{\Q}[0]{\mat{Q}}
# \newcommand{\B}[0]{\mat{B}}
# \newcommand{\C}[0]{\mat{C}}
# \newcommand{\Ri}[0]{\R^{-1}}
# \newcommand{\Bi}[0]{\B^{-1}}
# \newcommand{\X}[0]{\mat{X}}
# \newcommand{\A}[0]{\mat{A}}
# \newcommand{\Y}[0]{\mat{Y}}
# \newcommand{\E}[0]{\mat{E}}
# \newcommand{\U}[0]{\mat{U}}
# \newcommand{\V}[0]{\mat{V}}
# %
# \newcommand{\x}[0]{\bvec{x}}
# \newcommand{\y}[0]{\bvec{y}}
# \newcommand{\z}[0]{\bvec{z}}
# \newcommand{\q}[0]{\bvec{q}}
# \newcommand{\br}[0]{\bvec{r}}
# \newcommand{\bb}[0]{\bvec{b}}
# %
# \newcommand{\bx}[0]{\bvec{\bar{x}}}
# \newcommand{\by}[0]{\bvec{\bar{y}}}
# \newcommand{\barB}[0]{\mat{\bar{B}}}
# \newcommand{\barP}[0]{\mat{\bar{P}}}
# \newcommand{\barC}[0]{\mat{\bar{C}}}
# \newcommand{\barK}[0]{\mat{\bar{K}}}
# %
# \newcommand{\D}[0]{\mat{D}}
# \newcommand{\Dobs}[0]{\mat{D}_{\text{obs}}}
# \newcommand{\Dmod}[0]{\mat{D}_{\text{obs}}}
# %
# \newcommand{\ones}[0]{\bvec{1}}
# \newcommand{\AN}[0]{\big( \I_N - \ones \ones\tr / N \big)}
# %
# % END OF MACRO DEF
# $
#
# # Data assimilation (DA) & the ensemble Kalman filter (EnKF)
# *Copyright (c) 2020, <NAME>
# ### Jupyter notebooks is:
# the format used for these tutorials. Notebooks combine **cells** of code (Python) with cells of text (markdown).
# The exercises in these tutorials only require light Python experience.
# For example, edit the cell below (double-click it),
# insert your name,
# and run it (press "Run" in the toolbar).
name = "Batman"
print("Hello world! I'm " + name)
for i,c in enumerate(name):
print(i,c)
# You will likely be more efficient if you know these **keyboard shortcuts**:
#
# | Navigate | Edit | Exit | Run | Run & go to next |
# | ------------- | : ------------- : | ------------- | : --- : | : ------------- : |
# | <kbd>↓</kbd> and <kbd>↑</kbd> | <kbd>Enter</kbd> | <kbd>Esc</kbd> | <kbd>Ctrl</kbd>+<kbd>Enter</kbd> | <kbd>Shift</kbd>+<kbd>Enter</kbd> |
#
# When you open a notebook it starts a **session (kernel/runtime)** of Python in the background. All of the Python code cells (in a given notebook) are connected (they use the same Python kernel and thus share variables, functions, and classes). Thus, the **order** in which you run the cells matters. For example:
#
# <mark><font size="-1">
# The 1st two code cells in each tutorial (from now on) will be the following, which <em>you must run before any other</em>:
# </font></mark>
from resources.workspace import *
# One thing you must know is how to **restart** the Python session, which clears all of your variables, functions, etc, so that you can start over. Test this now by going through the top menu bar: `Kernel` → `Restart & Clear Output`. But remember to run the above cell again!
# There is a huge amount of libraries available in **Python**, including the popular `scipy` (with `numpy` at its core) and `matplotlib` packages. These were (implicitly) imported (and abbreviated) as `sp`, `np`, and `mpl` and `plt` in the previous cell. Try them out by running the following:
# +
# Use numpy's arrays for vectors and matrices. Example constructions:
a = np.arange(10) # Alternatively: np.array([0,1,2,3,4,5,6,7,8,9])
I = 2*np.eye(10) # Alternatively: np.diag(2*np.ones(10))
print("Indexing examples:")
print("a =", a)
print("a[3] =", a[3])
print("a[0:3] =", a[0:3])
print("a[:3] =", a[:3])
print("a[3:] =", a[3:])
print("a[-1] =", a[-1])
print("I[:3,:3] =", I[:3,:3], sep="\n")
print("\nLinear algebra examples:")
print("100+a =", 100+a)
print("I@a =", I@a)
print("I*a =", I*a, sep="\n")
plt.title("Plotting example")
plt.ylabel("i $x^2$")
for i in range(4):
plt.plot(i * a**2, label="i = %d"%i)
plt.legend();
# -
# ### Data assimilation (DA) is:
#
# * the calibration of big models with big data;
# * the fusion of forecasts with observations.
#
# This illustrated on the right (source: Data Assimilation Research Team, <a href="http://www.aics.riken.jp">www.aics.riken.jp</a>) as a "bridge" between data and models.
# <img align="right" width="400" src="./resources/DA_bridges.jpg" alt='DA "bridges" data and models.'/>
#
# The problem of ***DA*** fits well within the math/stats theory of ***state estimation*** and ***sequential inference***. A concise theoretical overview of DA is given by <NAME> Berliner: [A Bayesian tutorial for data assimilation](http://web-static-aws.seas.harvard.edu/climate/pdf/2007/Wikle_Berliner_InPress.pdf)
#
# Modern DA builds on state estimation techniques such as the ***Kalman filter (KF)***, which is an algorithm that recursively performs a form of least-squares regression. It was developed to steer the Apollo mission rockets to the moon, but also has applications outside of control systems, such as speech recognition, video tracking, and finance. An [introduction by pictures](http://www.bzarg.com/p/how-a-kalman-filter-works-in-pictures/) is provided by <NAME>. An [interactive tutorial](https://github.com/rlabbe/Kalman-and-Bayesian-Filters-in-Python) has been made by <NAME>.
#
# When it was first proposed to apply the KF to DA (specifically, weather forecasting), the idea sounded ludicrous because of some severe **technical challenges in DA (vs. "classic" state estimation)**:
# * size of data and models;
# * nonlinearity of models;
# * sparsity and inhomogeneous-ness of data.
#
# Some of these challenges may be recognized in the video below.
envisat_video()
# ### The EnKF is:
# an ensemble (Monte-Carlo) formulation of the KF
# that manages (fairly well) to deal with the above challenges in DA.
#
# For those familiar with the method of 4D-Var, **further advantages of the EnKF** include it being:
# * Non-invasive: the models are treated as black boxes, and no explicit Jacobian is required.
# * Bayesian:
# * provides ensemble of possible realities;
# - arguably the most practical form of "uncertainty quantification";
# - ideal way to initialize "ensemble forecasts";
# * uses "flow-dependent" background covariances in the analysis.
# * Embarrassingly parallelizable:
# * distributed across realizations for model forecasting;
# * distributed across local domains for observation analysis.
#
# The rest of this tutorial provides an EnKF-centric presentation of DA; it also has a [theoretical companion](./resources/companion/DA_tut.pdf).
# ### DAPPER example
# This tutorial builds on the underlying package, DAPPER, made for academic research in DA and its dissemination. For example, the code below is taken from `DAPPER/example_1.py`. It illustrates DA on a small toy problem.
#
# Run the cells in order and try to interpret the output.
#
# <mark><font size="-1">
# <em>Don't worry</em> if you can't understand what's going on -- we will discuss it later throughout the tutorials.
# </font></mark>
#
from dapper.mods.Lorenz63.sak12 import HMM
HMM.t.T = 30
# print(HMM)
xx,yy = simulate(HMM)
config = EnKF_N(N=4, store_u=True)
stats = config.assimilate(HMM,xx,yy)
avrgs = stats.average_in_time()
# print(avrgs)
print_averages(config,avrgs,[],['rmse_a','rmv_a'])
replay(stats,figlist="all")
# ### Exercises
#
# <mark><font size="-1">
# Exercises marked with an asterisk (*) are <em>optional.</em>
# </font></mark>
#
# **Exc 1.2:** Word association.
# Fill in the `X`'s in the table to group the words according to meaning.
#
# `Filtering, Sample, Random, Measurements, Kalman filter (KF), Monte-Carlo, Observations, Set of draws, State estimation, Data fusion`
#
# ```
# Data Assimilation (DA) Ensemble Stochastic Data
# ------------------------------------------------------------
# X X X X
# X X X X
# X
# X
# ```
# +
#show_answer('thesaurus 1')
# -
# * "The answer" is given from the perspective of DA. Do you agree with it?
# * Can you describe the (important!) nuances between the similar words?
# **Exc 1.3*:** Word association (advanced).
# Group these words:
#
# `Inverse problems, Sample point, Probability, Particle, Sequential, Inversion, Realization, Relative frequency, Iterative, Estimation, Single draw, Serial, Approximation, Regression, Fitting`
#
#
# Statistical inference Ensemble member Quantitative belief Recursive
# -----------------------------------------------------------------------------
# X X X X
# X X X X
# X X X
# X X
# X
# X
# +
#show_answer('thesaurus 2')
# -
# **Exc 1.5*:** Prepare to discuss the following questions. Use any tool at your disposal.
# * (a) What is DA?
# * (b) What are "state variables"? How do they differ from parameters?
# * (c) What is a "dynamical system"?
# * (d) Is DA a science, an engineering art, or a dark art?
# * (e) What is the point of "Hidden Markov Models"?
# +
#show_answer('Discussion topics 1')
# -
# ### Next: [Bayesian inference](T2%20-%20Bayesian%20inference.ipynb)
| notebooks/T1 - DA & EnKF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/unburied/DS-Unit-1-Sprint-1-Dealing-With-Data/blob/master/LS_DS_114_Making_Data_backed_Assertions.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Okfr_uhwhS1X" colab_type="text"
# # Lambda School Data Science - Making Data-backed Assertions
#
# This is, for many, the main point of data science - to create and support reasoned arguments based on evidence. It's not a topic to master in a day, but it is worth some focused time thinking about and structuring your approach to it.
# + [markdown] id="9dtJETFRhnOG" colab_type="text"
# ## Lecture - generating a confounding variable
#
# The prewatch material told a story about a hypothetical health condition where both the drug usage and overall health outcome were related to gender - thus making gender a confounding variable, obfuscating the possible relationship between the drug and the outcome.
#
# Let's use Python to generate data that actually behaves in this fashion!
# + id="WiBkgmPJhmhE" colab_type="code" outputId="4da96497-5895-471e-d544-26b253e33c10" colab={"base_uri": "https://localhost:8080/", "height": 1054}
import random
dir(random) # Reminding ourselves what we can do here
# + id="Ks5qFtpnq-q5" colab_type="code" outputId="6bbcce65-7a8a-49de-fcb4-3ba5f41a1484" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Let's think of another scenario:
# We work for a company that sells accessories for mobile phones.
# They have an ecommerce site, and we are supposed to analyze logs
# to determine what sort of usage is related to purchases, and thus guide
# website development to encourage higher conversion.
# The hypothesis - users who spend longer on the site tend
# to spend more. Seems reasonable, no?
# But there's a confounding variable! If they're on a phone, they:
# a) Spend less time on the site, but
# b) Are more likely to be interested in the actual products!
# Let's use namedtuple to represent our data
from collections import namedtuple
# purchased and mobile are bools, time_on_site in seconds
User = namedtuple('User', ['purchased','time_on_site', 'mobile'])
example_user = User(False, 12, False)
print(example_user)
# + id="lfPiHNG_sefL" colab_type="code" outputId="109c6e36-3d3f-462c-ac9a-17b4a3738a9d" colab={"base_uri": "https://localhost:8080/", "height": 54}
# And now let's generate 1000 example users
# 750 mobile, 250 not (i.e. desktop)
# A desktop user has a base conversion likelihood of 10%
# And it goes up by 1% for each 15 seconds they spend on the site
# And they spend anywhere from 10 seconds to 10 minutes on the site (uniform)
# Mobile users spend on average half as much time on the site as desktop
# But have three times as much base likelihood of buying something
users = []
for _ in range(250):
# Desktop users
time_on_site = random.uniform(10, 600)
purchased = random.random() < 0.1 + (time_on_site / 1500)
users.append(User(purchased, time_on_site, False))
for _ in range(750):
# Mobile users
time_on_site = random.uniform(5, 300)
purchased = random.random() < 0.3 + (time_on_site / 1500)
users.append(User(purchased, time_on_site, True))
random.shuffle(users)
print(users[:10])
# + id="9gDYb5qGuRzy" colab_type="code" outputId="8799f14d-ee92-4eb1-dab8-1c43cce3af91" colab={"base_uri": "https://localhost:8080/", "height": 204}
# Let's put this in a dataframe so we can look at it more easily
import pandas as pd
user_data = pd.DataFrame(users)
user_data.head()
# + id="sr6IJv77ulVl" colab_type="code" outputId="32eb7fae-73da-4f95-f606-89a690197000" colab={"base_uri": "https://localhost:8080/", "height": 191}
# Let's use crosstabulation to try to see what's going on
pd.crosstab(user_data['purchased'], user_data['time_on_site'])
# + id="hvAv6J3EwA9s" colab_type="code" outputId="91a20912-13be-4679-fda3-6b16fd96e44a" colab={"base_uri": "https://localhost:8080/", "height": 340}
# OK, that's not quite what we want
# Time is continuous! We need to put it in discrete buckets
# Pandas calls these bins, and pandas.cut helps make them
time_bins = pd.cut(user_data['time_on_site'], 5) # 5 equal-sized bins
pd.crosstab(user_data['purchased'], time_bins)
# + id="pjcXnJw0wfaj" colab_type="code" colab={}
# We can make this a bit clearer by normalizing (getting %)
pd.crosstab(user_data['purchased'], time_bins, normalize='columns')
# + id="C3GzvDxlvZMa" colab_type="code" colab={}
# That seems counter to our hypothesis
# More time on the site can actually have fewer purchases
# But we know why, since we generated the data!
# Let's look at mobile and purchased
pd.crosstab(user_data['purchased'], user_data['mobile'], normalize='columns')
# + id="KQb-wU60xCum" colab_type="code" colab={}
# Yep, mobile users are more likely to buy things
# But we're still not seeing the *whole* story until we look at all 3 at once
# Live/stretch goal - how can we do that?
# + [markdown] id="lOqaPds9huME" colab_type="text"
# ## Assignment - what's going on here?
#
# Consider the data in `persons.csv` (already prepared for you, in the repo for the week). It has four columns - a unique id, followed by age (in years), weight (in lbs), and exercise time (in minutes/week) of 1200 (hypothetical) people.
#
# Try to figure out which variables are possibly related to each other, and which may be confounding relationships.
# + id="xRy7-aEhACJh" colab_type="code" colab={}
import pandas as pd
# + id="8LfuNdZzAHiV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="cdb59df2-7de8-4bcf-ce53-b558216ca732"
df = pd.read_csv('persons.csv')
df.head()
# + id="3nNx5OZVAcfH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="84e89fcd-f00e-41fd-c361-556ddfca7df8"
df.dtypes
# + id="bjJsBJ1BBKnU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="09db66d7-e83d-4aa6-828e-3f94fef6d92d"
df.drop(labels = 'Unnamed: 0', axis = 1, inplace = True)
df.head()
# + id="8FGZLQgdKzy3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="470a3b7e-7423-4f45-b16e-9f26295aa015"
df['age'].hist();
# + id="EdZP1tArLGM5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="0fe884dc-fed6-4fcc-8eb6-1ae7ee9ece73"
df['weight'].hist();
# + id="AMhoK1weLTR8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="754324c0-e7cf-474e-e577-275ef97c11af"
df['exercise_time'].hist();
# + id="hchUPg9jLfpO" colab_type="code" colab={}
import matplotlib.pyplot as plt
# + id="d8pqMFiyLpe4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="12ec2aa8-3d62-47b7-9497-9df5bba4713c"
plt.scatter(df['age'], df['weight']);
# + id="7WWnRtoxL0n_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="aff08621-4a27-42af-fb78-4f6d9e4d8d0e"
plt.scatter(df['age'], df['exercise_time']);
# + id="av1j1e4MM51E" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="94c69408-263b-4c27-a279-e20b16bbc8fe"
plt.scatter(df['weight'], df['exercise_time']);
# + id="jzyjSVqePEoi" colab_type="code" colab={}
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
# + id="KdQ58un2PWpn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="065f5fa4-3f91-4d9e-d9fe-c1b9b163135f"
train, test = train_test_split(df)
train.shape, test.shape
# + id="9bOF-J9OP3QM" colab_type="code" colab={}
from sklearn.metrics import mean_absolute_error
def error():
y_true = train[target]
y_pred = model.predict(train[features])
train_error = mean_absolute_error(y_true, y_pred)
y_true = test[target]
y_pred = model.predict(test[features])
test_error = mean_absolute_error(y_true, y_pred)
print('Trained Weight Error: ', round(train_error))
print('Test Weight Error: ', round(test_error))
# + id="R1j_BBzARdv3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="aaa7f177-8488-4b76-f644-fd8fc76ef44f"
features = ['exercise_time']
target = 'weight'
model = LinearRegression()
model.fit(train[features], train[target])
error()
# + id="VDZ4_b9RSVdq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="a6357ef9-f3ef-42b1-81ab-09de08e49a89"
df['weight'].describe()
# + id="_NNbHR8pS7Ht" colab_type="code" colab={}
b = model.intercept_
m = model.coef_
x = df['exercise_time']
y = m * x + b
# + id="yb32kMzyUHWH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="e7d9614a-ec19-49fd-a2ec-5578e917954d"
plt.plot(x,y);
# + [markdown] id="BT9gdS7viJZa" colab_type="text"
# ### Assignment questions
#
# After you've worked on some code, answer the following questions in this text block:
#
#
#
# 1. What are the variable types in the data?
#
# All of the variables are integers, and couold also be considered semi-continuous
#
#
#
# 2. What are the relationships between the variables?
#
#
#
# Weight seems to be a dependant variable here. I used a model to better outline what the relationships entailed. Age and Excercise time appear to be independant, and unrelated to each other.
#
#
#
# 3. Which relationships are "real", and which spurious?
#
#
# Age and Excercise can be considered spurious. You cant claim you get older because you exercise more, or vice versa. There would be other factors involved. Time being one. I initially thought there would be a relationship between Age and Weight, but upon closer inspection, there was none. People of all ages can come in all sizes. There 'real' relationship came from Weight and Exercise. As you can see from the model above, the more you exercise, the less you weigh. The model appears to be pretty accurate, with the degree of error being less than one standard deviation of the weight distribution.
# + [markdown] id="_XXg2crAipwP" colab_type="text"
# ## Stretch goals and resources
#
# Following are *optional* things for you to take a look at. Focus on the above assignment first, and make sure to commit and push your changes to GitHub.
#
# - [Spurious Correlations](http://tylervigen.com/spurious-correlations)
# - [NIH on controlling for confounding variables](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4017459/)
#
# Stretch goals:
#
# - Produce your own plot inspired by the Spurious Correlation visualizations (and consider writing a blog post about it - both the content and how you made it)
# - Pick one of the techniques that NIH highlights for confounding variables - we'll be going into many of them later, but see if you can find which Python modules may help (hint - check scikit-learn)
| LS_DS_114_Making_Data_backed_Assertions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc="true"
# # Table of Contents
# <p><div class="lev1 toc-item"><a href="#Dataset" data-toc-modified-id="Dataset-1"><span class="toc-item-num">1 </span>Dataset</a></div><div class="lev1 toc-item"><a href="#Imports" data-toc-modified-id="Imports-2"><span class="toc-item-num">2 </span>Imports</a></div><div class="lev1 toc-item"><a href="#Read-Tags" data-toc-modified-id="Read-Tags-3"><span class="toc-item-num">3 </span>Read Tags</a></div><div class="lev1 toc-item"><a href="#Read-Posts" data-toc-modified-id="Read-Posts-4"><span class="toc-item-num">4 </span>Read Posts</a></div><div class="lev1 toc-item"><a href="#Read-post-parquet" data-toc-modified-id="Read-post-parquet-5"><span class="toc-item-num">5 </span>Read post parquet</a></div><div class="lev1 toc-item"><a href="#Read-Tags-xml" data-toc-modified-id="Read-Tags-xml-6"><span class="toc-item-num">6 </span>Read Tags xml</a></div><div class="lev1 toc-item"><a href="#Join-post-and-tag" data-toc-modified-id="Join-post-and-tag-7"><span class="toc-item-num">7 </span>Join post and tag</a></div>
# + [markdown] deletable=true editable=true
# # Dataset
#
# stackoverflow datascience 7zip stored at [Archive.org](https://archive.org/download/stackexchange/datascience.stackexchange.com.7z)
#
# Download the 7zip file and decompress the xml files inside the folder called `datascience`.
# + [markdown] deletable=true editable=true
# # Imports
# + deletable=true editable=true
import numpy as np
import pandas as pd
# plotting
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
sns.set(color_codes=True)
sns.set()
sns.set_style('darkgrid')
# + deletable=true editable=true
import dask
import dask.bag as db
import dask.dataframe as dd
from dask.dot import dot_graph
from dask.diagnostics import ProgressBar
# + deletable=true editable=true
import re
import html
from datetime import datetime
# + deletable=true editable=true
# %load_ext watermark
# %watermark --iversions
# + [markdown] deletable=true editable=true
# # Read Tags
# + deletable=true editable=true
tags_xml = db.read_text('datascience//Tags.xml', encoding='utf-8')
# + deletable=true editable=true
lst = [attr for attr in dir(tags_xml) if attr[0].islower() ]
lst = np.array_split(lst,5)
pd.DataFrame(lst).T
# + deletable=true editable=true
tags_xml.take(10)
# + deletable=true editable=true
# we need only lines with word rows
# + deletable=true editable=true
tags_rows = tags_xml.filter(lambda line: line.find('<row') >= 0)
tags_rows.take(10)
# + deletable=true editable=true
def extract_column_value(line, col_name, cast_type=str):
# TagName=".net" ==> .net when column is TagName
pattern_tpl = r'{col}="([^"]*)"'
pattern = pattern_tpl.format(col=col_name)
match = re.search(pattern, line)
if cast_type == int:
null_value = 0 # numpy does not have nans for ints, make it zero
else:
null_value = None
return cast_type(match[1]) if match is not None else null_value
# + deletable=true editable=true
test_tags_row = '<row Id="1" TagName=".net" Count="257092" ExcerptPostId="3624959" WikiPostId="3607476" />'
print(extract_column_value(test_tags_row, 'TagName'))
print(extract_column_value(test_tags_row, 'TagName'))
print(extract_column_value(test_tags_row, 'NotHere', str))
print(extract_column_value(test_tags_row, 'NotHere', int))
# + deletable=true editable=true
def extract_tags_columns(line):
row = {
'id': extract_column_value(line, 'Id', int),
'tag_name': extract_column_value(line, 'TagName', str),
'count': extract_column_value(line, 'Count', int),
}
return row
# + deletable=true editable=true
test_tags_row = '<row Id="1" TagName=".net" Count="257092" ExcerptPostId="3624959" WikiPostId="3607476" />'
extract_tags_columns(test_tags_row)
# + deletable=true editable=true
# now, get dask dataframes
# to_dataframe makes pandas dataframe.
tags = tags_rows.map(extract_tags_columns).to_dataframe()
tags
# + deletable=true editable=true
tags.visualize()
# + deletable=true editable=true
# after building graph, compute
# now, its panads! enjoy the calculations.
tags_df = tags.compute()
print(type(tags_df))
tags_df.head()
# + [markdown] deletable=true editable=true
# # Read Posts
# + deletable=true editable=true
posts_xml = db.read_text('datascience/Posts.xml', encoding='utf-8', blocksize=1e8)
posts_xml
# + deletable=true editable=true
posts_xml.take(5)
# + deletable=true editable=true
# filter only the rows
posts_rows = posts_xml.filter(lambda line: line.find('<row') >= 0)
posts_rows.take(5)
# + deletable=true editable=true
# deal with timestamp
def str_to_datetime(timestamp):
f = '%Y-%m-%dT%H:%M:%S.%f'
return datetime.strptime(timestamp, f)
str_to_datetime('2008-07-31T23:55:37.967')
# + deletable=true editable=true
def extract_posts_columns(line):
l = html.unescape(line)
row = {
'id': extract_column_value(l, 'Id', int),
'creation_date': extract_column_value(l, 'CreationDate', str_to_datetime),
'score': extract_column_value(l, 'Score', int),
'view_count': extract_column_value(l, 'ViewCount', int),
'tags': extract_column_value(l, 'Tags', str),
}
return row
# + deletable=true editable=true
test_posts_row = ' <row Id="4" PostTypeId="1" AcceptedAnswerId="7" CreationDate="2008-07-31T21:42:52.667" Score="506" ViewCount="32399" Body="<p>I want to use a track-bar to change a form\'s opacity.</p>

<p>This is my code:</p>

<pre><code>decimal trans = trackBar1.Value / 5000;
this.Opacity = trans;
</code></pre>

<p>When I build the application, it gives the following error:</p>

<blockquote>
 <p>Cannot implicitly convert type \'decimal\' to \'double\'.</p>
</blockquote>

<p>I tried using <code>trans</code> and <code>double</code> but then the control doesn\'t work. This code worked fine in a past VB.NET project. </p>
" OwnerUserId="8" LastEditorUserId="126970" LastEditorDisplayName="<NAME>" LastEditDate="2017-03-10T15:18:33.147" LastActivityDate="2017-03-10T15:18:33.147" Title="While applying opacity to a form should we use a decimal or double value?" Tags="<c#><winforms><type-conversion><decimal><opacity>" AnswerCount="13" CommentCount="5" FavoriteCount="37" CommunityOwnedDate="2012-10-31T16:42:47.213" />\n'
extract_posts_columns(test_posts_row)
# + deletable=true editable=true
# skip all the posts without tags.
posts = posts_rows.map(extract_posts_columns)\
.filter(lambda r: r['tags'] is not None)\
.to_dataframe()
posts
# + deletable=true editable=true
posts.head()
# + deletable=true editable=true
# after cleaning xml, write data to a more suitable parquet format.
with ProgressBar():
posts.repartition(npartitions=100)\
.to_parquet('datascience/posts_tags.parq', engine='fastparquet', compression='GZIP')
# + [markdown] deletable=true editable=true
# # Read post parquet
# + deletable=true editable=true
posts = dd.read_parquet('datascience/posts_tags.parq', engine='fastparquet')
posts
# + deletable=true editable=true
# yearly posts
with ProgressBar():
posts_count = posts.groupby(posts.creation_date.dt.year)\
.id.count()\
.to_frame()\
.rename(columns={'id': 'count'})\
.compute()
posts_count
# + deletable=true editable=true
posts_count.plot.bar();
# + deletable=true editable=true
# faster method
with ProgressBar():
posts_count = posts.creation_date.dt.year.value_counts()\
.to_frame()\
.rename(columns={'creation_date': 'count'})\
.compute()
posts_count
# + deletable=true editable=true
# 1. since parquet files are much much smaller than csv we can use pandas.
# 2. dask gives multiple parquet files, one per partition, we need to combine
# them to use in pandas since pandas load only one parquet.
#
# 3. we can also create pandas df directly from xml above, without writing to
# parquet and then reading from parquet, but resulting pandas dataframe
# will be much large (maybe 2GB) and jupyter kernels may die because of
# large object. e.g. when code completion.
#
# WHEN USING BIG OBJECTS, do not use CODE-COMPLETION!!!
# + deletable=true editable=true
posts = posts.set_index('id')
posts.head(2)
# + deletable=true editable=true
type(posts)
# + deletable=true editable=true
posts_count = posts.creation_date.dt.date.value_counts()
posts_count
# + deletable=true editable=true
with ProgressBar():
posts_count_df = posts_count.compute()
posts_count_df.head()
# + deletable=true editable=true
type(posts_count_df)
# + deletable=true editable=true
posts_count_df.plot();
# + deletable=true editable=true
# daily data looks messy, make it smoother using monthly resampling
# + deletable=true editable=true
# to deal with timeseries, alway make index timeseries
# + deletable=true editable=true
posts_count_df.index = pd.DatetimeIndex(posts_count_df.index)
# + deletable=true editable=true
posts_by_month = posts_count_df.resample('MS').sum()\
.rename('count').to_frame()
posts_by_month.head()
# + deletable=true editable=true
posts_by_month.plot();
# + deletable=true editable=true
# yearly plot
posts_count_df.groupby(posts_count_df.index.year).sum()\
.rename('count').to_frame().plot.bar();
# + [markdown] deletable=true editable=true
# # Read Tags xml
# + deletable=true editable=true
tags = db.read_text('datascience/Tags.xml')\
.filter(lambda l: l.find('<row') > 0)\
.map(extract_tags_columns)\
.to_dataframe()\
.set_index('tag_name')\
.compute()
print(tags.shape)
tags.head(10)
# + deletable=true editable=true
langs = ['c', 'c++', 'c#', 'java', 'javascript', 'python', 'r']
tags.loc[tags.index.isin(langs), :]
# + deletable=true editable=true
posts.head()
# + deletable=true editable=true
post_tags = posts.tags.str.lower()\
.str.extractall('<([^>]*)>')\
.rename(columns={0: 'tag'})
post_tags.head()
# + deletable=true editable=true
# persist keep data in RAM
with ProgressBar():
post_known_tags = post_tags.where(post_tags.tag.isin(langs))\
.dropna().persist()
# + deletable=true editable=true
post_known_tags.head()
# + deletable=true editable=true
# since we have data in RAM, this takes much small time.
with ProgressBar():
known_tags_count = post_known_tags.tag.value_counts().to_frame().compute()
# + deletable=true editable=true
known_tags_count
# + [markdown] deletable=true editable=true
# # Join post and tag
# + deletable=true editable=true
posts.head(2)
# + deletable=true editable=true
post_known_tags.head(2)
# + deletable=true editable=true
post_tag_join = (posts.join(post_known_tags, how='inner')
# timestamp 2014-05-14 11:15:40.907 ==> 2014-05-14 string format
.assign(creation_date = lambda df: df.creation_date.dt.date)
# object ==> category
.assign(tag = lambda df: df.tag.astype('category')))
post_tag_join
# + deletable=true editable=true
with ProgressBar():
post_tag_join = post_tag_join.persist()
# + deletable=true editable=true
post_tag_join.head()
# + deletable=true editable=true
post_tag_join.dtypes
# + deletable=true editable=true
# languages vs creation date pivot table
post_tag_join['tag'] = post_tag_join['tag'].cat.as_known()
post_pivot = post_tag_join.pivot_table(
index='creation_date',
columns='tag',
values='score',
aggfunc='count')
post_pivot
# + deletable=true editable=true
with ProgressBar():
post_pivot_df = post_pivot.compute()
post_pivot_df.head()
# + deletable=true editable=true
post_pivot_df.plot();
# + deletable=true editable=true
post_pivot_df.index = pd.DatetimeIndex(post_pivot_df.index)
# + deletable=true editable=true
tags_by_month = post_pivot_df.resample('MS').sum()
tags_by_month.head(10)
# weird!! no c tagged question for 2014 may to 2015 ???
# + deletable=true editable=true
tags_by_month.loc[tags_by_month.c >0 ,:].head()
# + deletable=true editable=true
tags_by_month.loc[tags_by_month.javascript >0 ,:].head()
# + deletable=true editable=true
tags_by_month.plot();
# + deletable=true editable=true
| a03_Dask/stackovr_datascience/stackovr_datascience_analysis_using_dask.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append('/opt/anaconda3/envs/pytorch/lib/python3.6')
sys.path.append('/opt/anaconda3/envs/pytorch/lib/python3.6/site-packages')
sys.path.append('/opt/anaconda3/envs/pytorch/lib/python3.6/site-packages/appv1')
import random
import time
import numpy as np
from tqdm import tqdm
import torch
from torch import nn
import torch.optim as optim
import torchtext
from utils.dataloader import get_chABSA_DataLoaders_and_TEXT
from utils.bert import BertTokenizer
# JUMANの動作確認
from pyknp import Juman
juman = Juman()
result = juman.analysis("今夜の月は綺麗ですね。")
for mrph in result.mrph_list(): # 各形態素にアクセス
print("見出し:%s, 読み:%s, 原形:%s, 品詞:%s, 品詞細分類:%s, 活用型:%s, 活用形:%s, 意味情報:%s, 代表表記:%s" \
% (mrph.midasi, mrph.yomi, mrph.genkei, mrph.hinsi, mrph.bunrui, mrph.katuyou1, mrph.katuyou2, mrph.imis, mrph.repname))
from config import *
from predict import predict, create_vocab_text, build_bert_model
from IPython.display import HTML, display
TEXT = create_vocab_text()
# +
from utils.bert import get_config, BertModel,BertForchABSA, set_learned_params
from utils.bert import get_config, BertModel,BertForchABSA, set_learned_params
# The setting parameter has been saved in config.json
config = get_config(file_path="/opt/anaconda3/envs/pytorch/lib/python3.6/site-packages/appv1/weights/bert_config.json")
# build a BERT model
net_bert = BertModel(config)
# Trained BERT model params setting
net_bert = set_learned_params(
net_bert, weights_path="/opt/anaconda3/envs/pytorch/lib/python3.6/site-packages/appv1/weights/pytorch_model.bin")
# +
# load the finetuning module
net_trained = BertForchABSA(net_bert)
save_path = '/opt/anaconda3/envs/pytorch/lib/python3.6/site-packages/appv1/weights/bert_fine_tuning_chABSA.pth'
net_trained.load_state_dict(torch.load(save_path, map_location='cpu'))
net_trained.eval()
# +
from utils.predict import predict2
input_text = "当社の業績は多岐にわたる変動要因の影響を受ける可能性があります。有価証券報告書に記載した事業の状況、経理の状況等に関する事項のうち、投資者の判断に重要な影響を及ぼす可能性のある事項には、以下のようなものがあります。なお、文中における将来に関する事項は、当事業年度末現在において当社が判断したものであります。(1)主要市場での需要の急激な変動について当社は、主に半導体業界及びFPD業界を対象として、その生産ラインで用いられる各種生産設備部品の製造・販売を行っていますが、半導体業界におきましてシリコンサイクル、FPD業界におきましてクリスタルサイクルと呼ばれる業界特有の好不況の波が存在します。当社におきましては、メーカーの設備投資動向に左右されない消耗品などの安定的な販売が見込める分野の受注に注力するなどの対策を行い、業績への影響を最小限にすべく努力しております。しかしながら、これらの景気変動によって、当社の業績及び財務状況に影響を与える可能性があります。"
#net_trained = build_bert_model()
net_trained.eval()
html_output = predict(input_text, net_trained)
print("======================推論結果の表示======================")
print(input_text)
display(HTML(html_output))
pred_output = predict2(input_text, net_trained)
print(pred_output)
# +
from utils.predict import predict2
df = pd.read_csv("/Users/chenying/testsentiment.csv", encoding="utf-8-sig")
df["PREDICT"] = np.nan #予測列を追加
net_trained.eval() #推論モードに。
for index, row in df.iterrows():
df.at[index, "PREDICT"] = predict2(row['text'], net_trained.numpy()[0] # GPU環境の場合は「.cpu().numpy()」としてください。
#print(df)
df.to_csv("/Users/chenying/predicted_test .csv", encoding="utf-8-sig", index=False)
# -
| chABSA_Predict.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.1 64-bit ('venv')
# metadata:
# interpreter:
# hash: a081492f977d15fe2cc5c47b1fe307f337ae9e296dcaffa5436ed2570310f973
# name: python3
# ---
# # Lab 1: MNIST
#
# Overview
# ## Get data
#
# The first step is to download the dataset from the sklearn collection.
#
# Then we are going to split the dataset into an `X` matrix with atributes, and an array `y` with labels.
# +
import sklearn
assert sklearn.__version__ >= "0.20"
from sklearn.datasets import fetch_openml
mnist = fetch_openml('mnist_784', version=1)
mnist.keys()
# -
X, y = mnist['data'], mnist['target']
X.shape
y.shape
# ## Data pre-processing
#
# We are going to take a look at the dataset.
#
# Then we're going to make some transformations.
# +
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
any_number = X.iloc[42].to_numpy()
any_number_image = any_number.reshape(28, 28)
plt.imshow(any_number_image, cmap = 'binary')
plt.axis("off")
plt.show()
# -
y[42]
# ### Cast label string to int
import numpy as np
y = y.astype(np.uint8)
y[42]
# ### Split dataset
X_train, X_test, y_train, y_test = X[:60000], X[60000:], y[:60000], y[60000:]
# ### Scaling inputs
# +
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train.astype(np.float64))
X_test_scaled = scaler.fit_transform(X_test.astype(np.float64))
# -
# ## Stochastic gradient descent learning
#
# ### Model creation
# +
from sklearn.linear_model import SGDClassifier
sgd_clf = SGDClassifier(random_state = 42)
# -
# ### Model training (with historical data)
sgd_clf.fit(X_train_scaled, y_train)
sgd_clf.predict([any_number])
# ### Model evaluation
# +
from sklearn.metrics import accuracy_score
# Accuracy
sgd_predictions = sgd_clf.predict(X_test_scaled)
accuracy_score(y_test, sgd_predictions)
# -
# ### Cross-Validation
# out of the box
from sklearn.model_selection import cross_val_score
cross_val_score(sgd_clf, X_test_scaled, y_test, cv=3, scoring='accuracy')
# ### Confusion Matrix
# +
# Get predictions
from sklearn.model_selection import cross_val_predict
y_test_pred = cross_val_predict(sgd_clf, X_test_scaled, y_test, cv=3)
# +
# Let's create a Confussion matrix
from sklearn.metrics import confusion_matrix
sgd_conf_mx = confusion_matrix(y_test, y_test_pred)
plt.matshow(sgd_conf_mx, cmap = plt.cm.gray)
plt.show()
# -
# ## Logistic regression learning
#
# ### Model creation
# +
from sklearn.linear_model import LogisticRegression
lr_clf = LogisticRegression(random_state = 42)
# -
# ### Model training (with historical data)
lr_clf.fit(X_train_scaled, y_train)
lr_clf.predict([any_number])
# ### Model evaluation
# +
# Accuracy
lr_predictions = lr_clf.predict(X_test_scaled)
accuracy_score(y_test, lr_predictions)
# -
# ### Cross-Validation
cross_val_score(lr_clf, X_test_scaled, y_test, cv=3, scoring='accuracy')
# ### Confusion Matrix
y_test_pred_lr = cross_val_predict(lr_clf, X_test_scaled, y_test, cv=3)
lr_conf_mx = confusion_matrix(y_test, y_test_pred_lr)
plt.matshow(lr_conf_mx, cmap = plt.cm.gray)
plt.show()
# ## Passive Aggressive Classifier learning
#
# ### Model creation
# +
from sklearn.linear_model import PassiveAggressiveClassifier
pa_clf = PassiveAggressiveClassifier(random_state = 42)
# -
# ### Model training (with historical data)
# pa_clf.fit(X_train_scaled, y_train) -> this did not work well here!
pa_clf.fit(X_train, y_train)
pa_clf.predict([any_number])
# ### Model evaluation
# +
# Accuracy
pa_predictions = pa_clf.predict(X_test)
accuracy_score(y_test, pa_predictions)
# -
# ### Cross-Validation
cross_val_score(pa_clf, X_test, y_test, cv=3, scoring='accuracy')
# ### Confusion Matrix
y_test_pred_pa = cross_val_predict(pa_clf, X_test, y_test, cv=3)
pa_conf_mx = confusion_matrix(y_test, y_test_pred_pa)
plt.matshow(pa_conf_mx, cmap = plt.cm.gray)
plt.show()
# ## Perceptron learning
#
# ### Model creation
# +
from sklearn.linear_model import Perceptron
per_clf = Perceptron(random_state = 42)
# -
# ### Model training (with historical data)
per_clf.fit(X_train_scaled, y_train)
per_clf.predict([any_number])
# ### Model evaluation
# +
# Accuracy
per_predictions = per_clf.predict(X_test_scaled)
accuracy_score(y_test, per_predictions)
# -
# ### Cross-Validation
cross_val_score(per_clf, X_test_scaled, y_test, cv=3, scoring='accuracy')
# ### Confusion Matrix
y_test_pred_per = cross_val_predict(per_clf, X_test_scaled, y_test, cv=3)
per_conf_mx = confusion_matrix(y_test, y_test_pred_per)
plt.matshow(per_conf_mx, cmap = plt.cm.gray)
plt.show()
# ## Ridge Classifier learning
#
# ### Model creation
# +
from sklearn.linear_model import RidgeClassifier
rid_clf = RidgeClassifier(random_state = 42)
# -
# ### Model training (with historical data)
rid_clf.fit(X_train_scaled, y_train)
rid_clf.predict([any_number])
# ### Model evaluation
# +
# Accuracy
rid_predictions = rid_clf.predict(X_test_scaled)
accuracy_score(y_test, rid_predictions)
# -
# ### Cross-Validation
cross_val_score(rid_clf, X_test_scaled, y_test, cv=3, scoring='accuracy')
# ### Confusion Matrix
y_test_pred_rid = cross_val_predict(rid_clf, X_test_scaled, y_test, cv=3)
rid_conf_mx = confusion_matrix(y_test, y_test_pred_rid)
plt.matshow(rid_conf_mx, cmap = plt.cm.gray)
plt.show()
# ## Results
# Results | SGDClassifier | LogisticRegression | PassiveAggressiveClassifier | Perceptron | RidgeClassifier |
# :--------------------| :------------------------------------| :-------------------------------------| :-------------------------------------| :-------------------------------------| :-------------------------|
# Accuracy | 0.8971 | 0.9232 | 0.8797 | 0.8782 | 0.8598 |
# Cross-Validation | 0.86412717, 0.89708971, 0.89528953 | 0.86172765, 0.89138914, 0.89258926 | 0.82633473, 0.86918692, 0.87488749 | 0.82633473, 0.86918692, 0.87488749 | 0.78824235, 0.82928293, 0.83888389 |
| labs/lab-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## HackerEarth Competition notebook
# ### Import modules
# +
% matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import style
style.use('fivethirtyeight')
# for prediction, use machine learning
from sklearn import preprocessing, cross_validation
from sklearn.preprocessing import LabelEncoder
from sklearn.ensemble import RandomForestClassifier, GradientBoostingClassifier
# -
inputData = pd.read_csv('ign.csv')
inputData.head()
# ### Listing the platforms with the most "Editor's Choice" awards
# +
res = []
# we groupby platform
# groupby -> platform as first and rest as dataframe
# data contains rows grouped
for platform, data in inputData.groupby('platform'):
c = data[data['editors_choice'] == 'Y'].shape[0]
res.append((platform, c))
res.sort(key=lambda x: x[1], reverse=True)
res = pd.DataFrame.from_records(res, columns=('Platform', 'Number of EC Awards'))
res.sort()
# -
# #### Determine if the number of games by a platform in a year have any effect on the editors choice awards?
# +
# we can check how many games per platform are released in a year and how many gets award, calculate percentage
# can use above code, nah lets write again (remove comment)
calcPer = []
for platform, data in inputData.groupby('platform'):
c = data[data['editors_choice'] == 'Y'].shape[0]
calcPer.append((platform, c, data.shape[0], float(c)/data.shape[0]))
calcPer = pd.DataFrame.from_records(calcPer, columns=('Platform', 'Total Awards', 'Total Games', 'Award Rate'))
calcPer
# -
# ** Well I think that the total number of games released on a platform in a year have no effect on the number of editor's choice awards presented to games for a platform. **
# ### Calculate Macintosh's average award count?
# +
# number of awards given to Macintosh
number_of_awards_for_macintosh = res.loc[res['Platform'] == 'Macintosh']['Number of EC Awards']
# total number of games released on Macintosh platform
macplatform = inputData.groupby('platform')['platform']
total_mac_games = macplatform.get_group('Macintosh').count()
# Macintosh average award count
c = float(number_of_awards_for_macintosh) / total_mac_games
print c
# -
# ** So from above we can infer that for every two Mac games released and reviewed, one gets the Editor's Choice award. Total number of Macintosh games is 81 and total number of awards awarded are 40 (roughly half) **
# ### Finding optimal month for releasing a game?
# +
# we'll just group by total games released in a given month, months are in number (1-12)
# better to have a list of months for better view
import calendar
# inputData['release_month'] = inputData['release_month'].apply(lambda m: calendar.month_abbr[m])
# # inputData.head()
# d = inputData.groupby('release_month')['release_month']
# d.count()
res_months = []
for month, data in inputData.groupby('release_month'):
res_months.append((calendar.month_abbr[month], data.shape[0]))
res_months = pd.DataFrame.from_records(res_months, columns=('month', 'count'))
res_months
# -
# plotting for better view (just cause we can)
res_months[['count']].plot.bar(x = res_months['month'], legend=True)
# ** So from the above graph we can infer that the optimal month for releasing game is November followed by October. Meaning companies have been releasing most of the games in October-November period. **
# ### Analyse the percentage growth in the gaming industry over the years
# +
# we can check for the total number of games released in the years
tgy = []
for year, data in inputData.groupby('release_year'):
tgy.append((year, data.shape[0]))
tgy = pd.DataFrame.from_records(tgy, columns=('Year', 'Total Games'))
tgy
# -
# *** It would seem that the release of video games has decreased over the years, lets plot a graph to better view it. ***
# let plot this one too for better view
tgy[['Total Games']].plot.bar(x = tgy['Year'], legend=True)
# ** We can confirm that the growth rate of the games industry is decreasing with the years as less and less games are released every year. Also it can be that the data is not proper or is incomplete with more older games. Also the data doesn't consist of revenue, maybe companies are releasing less games but the each game is earning a lot of revenue. **
# ## Lets build a predictive model for predicting which games will win editor's choice awards in a given year?
# +
inputData.describe()
# target column
target_col = ['editors_choice']
# numerical variables
num_cols = ['score', 'release_year', 'release_month', 'release_day']
# categorical variables
cat_cols = ['score_phrase', 'platform', 'genre']
other_col = ['Unnamed: 0']
extra_cols = ['url', 'title']
# combine num_cols and cat_cols
num_cat_cols = num_cols + cat_cols
# -
# check missing values
inputData.isnull().any()
# +
# create label encoders for categorical features
for item in cat_cols:
num = LabelEncoder()
inputData[item] = num.fit_transform(inputData[item].astype('str'))
# target variable is also categorical, so convert it
inputData['editors_choice'] = num.fit_transform(inputData['editors_choice'].astype('str'))
# -
# create features
features=list(set(list(inputData.columns))-set(target_col)-set(other_col)-set(extra_cols))
features
X = inputData[list(features)].values
y = inputData['editors_choice'].values
# split the data into train data and test data (70-30)
X_train, X_test, y_train, y_test = cross_validation.train_test_split(X, y, test_size=0.3)
# lets train
clf = RandomForestClassifier(n_estimators=1000)
clf.fit(X_train, y_train)
print clf.score(X_test, y_test)
| HackerEarthComp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install tweepy
# ## Scraping Twitter
#
# In this notebook, we make use of [Tweepy](https://www.tweepy.org/) to download tweets from [Twitter](https://twitter.com/).
# Do note that at the point of creating this notebook, Twitter API is transiting to v2.
# +
import datetime
import time
import pandas as pd
import tweepy
# -
tweepy.__version__
# Below are the data that we are going to capture from the downloaded tweets.
def get_df():
return pd.DataFrame(
columns=[
"tweet_id",
"name",
"screen_name",
"retweet_count",
"text",
"mined_at",
"created_at",
"favourite_count",
"hashtags",
"status_count",
"followers_count",
"location",
"source_device",
]
)
# ### Twitter developer account and authentication
#
# Before starting out, remember to get a [Twitter developer account](https://developer.twitter.com/en/docs/apps/overview) from its [Developer portal](https://developer.twitter.com/en) if you haven't.
# Refer to the [Twitter API documentation](https://developer.twitter.com/en/docs/authentication/oauth-1-0a) on how to get the access tokens to be set under the `twitter_keys` below:
# The [standard search API](https://developer.twitter.com/en/docs/twitter-api/v1/tweets/search/overview) from Twitter API v1.1 searches against sampling of recent Tweets published in the past 7 days. This will be replaced by the [recent search](https://developer.twitter.com/en/docs/twitter-api/tweets/search/introduction) endpoint in v2.
#
# This search is not exhausive. Alternatively, if you have the tweet id, you can always pass the array of id to [`api.statuses_lookup()`](http://docs.tweepy.org/en/v3.5.0/api.html#API.statuses_lookup) to retrieve the historical tweets. You can find the list of tweets used in this series of notebook [here](https://s3.eu-west-3.amazonaws.com/data.atoti.io/notebooks/twitter/tweets_sentiments.csv) alongside with the sentiments at the point of data collection.
# Remember to set `wait_on_rate_limit` to true so that exception won't be thrown when the rate limits are hit.
class TweetMiner(object):
result_limit = 20
data = []
api = False
twitter_keys = {
"consumer_key": "<To be replace>",
"consumer_secret": "<To be replace>",
"access_token_key": "<To be replace>",
"access_token_secret": "<To be replace>",
}
def __init__(self, keys_dict=twitter_keys, api=api):
self.twitter_keys = keys_dict
auth = tweepy.OAuthHandler(
keys_dict["consumer_key"], keys_dict["consumer_secret"]
)
auth.set_access_token(
keys_dict["access_token_key"], keys_dict["access_token_secret"]
)
self.api = tweepy.API(
auth, wait_on_rate_limit=True, wait_on_rate_limit_notify=True
)
self.twitter_keys = keys_dict
def mine_crypto_currency_tweets(self, query="BTC"):
last_tweet_id = False
page_num = 1
data = get_df()
cypto_query = f"#{query}"
print(" ===== ", query, cypto_query)
for page in tweepy.Cursor(
self.api.search,
q=cypto_query,
lang="en",
tweet_mode="extended",
count=200, # max_id=1295144957439690000
).pages():
print(" ...... new page", page_num)
page_num += 1
for item in page:
mined = {
"tweet_id": item.id,
"name": item.user.name,
"screen_name": item.user.screen_name,
"retweet_count": item.retweet_count,
"text": item.full_text,
"mined_at": datetime.datetime.now(),
"created_at": item.created_at,
"favourite_count": item.favorite_count,
"hashtags": item.entities["hashtags"],
"status_count": item.user.statuses_count,
"followers_count": item.user.followers_count,
"location": item.place,
"source_device": item.source,
}
try:
mined["retweet_text"] = item.retweeted_status.full_text
except:
mined["retweet_text"] = "None"
last_tweet_id = item.id
data = data.append(mined, ignore_index=True)
if page_num % 180 == 0:
date_label = datetime.datetime.now().strftime("%Y%m%d_%H%M%S")
print("....... outputting to csv", page_num, len(data))
data.to_csv(f"{query}_{page_num}_{date_label}.csv", index=False)
print(" ..... resetting df")
data = get_df()
date_label = datetime.datetime.now().strftime("%Y%m%d_%H%M%S")
data.to_csv(f"{query}_{page_num}_{date_label}.csv", index=False)
miner = TweetMiner()
# Below are the cryptocurrency hashtags that will be used to query. We have the option of combining query but in this case, we download the tweets for each cryptocurrency separately.
handle_list = [
"BTC",
"ETH",
"USDT",
"XRP",
"BCH",
"ADA",
"BSV",
"LTC",
"LINK",
"BNB",
"EOS",
"TRON",
]
# We created threads to queue the mining of the cryptocurrency so that it can run unmanned for 10 iterations.
# Since the tweets are sampled, we figured we should repeat the mining to gather as much data as possible.
# +
import queue
import threading
should_publish = threading.Event()
update_queue = queue.Queue()
def start_publisher():
global handle_list
starttime = time.time()
print("Start polling", starttime)
poll_iteration = 1
for i in range(10):
for name in handle_list:
print(i, poll_iteration, "\rpublishing update ", end="")
update_queue.put((poll_iteration, name))
poll_iteration += 1
time.sleep(900)
print("\rawaiting for publishing update", end="")
should_publish.wait()
update_queue.join()
def start_update_listener():
while True:
poll_iteration, name = update_queue.get()
print(" --- ", name)
try:
miner.mine_crypto_currency_tweets(query=name)
update_queue.task_done()
except Exception as e: # work on python 3.x
print("Failed to upload to ftp: " + str(e))
listener_thread = threading.Thread(target=start_update_listener, daemon=True)
publisher_thread = threading.Thread(target=start_publisher, daemon=True)
# -
publisher_thread.start()
listener_thread.start()
# start publishing
should_publish.set()
# If you would like to stop the data polling before the 10 iterations end, run the below cell.
# pause publishing
should_publish.clear()
| notebooks/twitter/01_tweets_mining.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Script for extracting product list for a seller
#
# The script takes as input a .txt file containing a list of brands offered by a seller, and then iterates through all the brands in the list.
#
# For each brand in the brand list, it finds all the products offered by the given seller. The details are stored in a .jsonl file.
import urllib.request # FOR URL ENCODING
import requests # For making requests to download a webpage content
from selectorlib import Extractor # For extracting specific fileds from downloaded webpage
import json
import random
from time import sleep
import os
import jsonlines
import pandas as pd
import datetime
import re
# #### Step 1: Read Brand List
#
# **NOTE:** Before running this, change the path variable 'brands' to point to the Brand List file. The brand list should be in .txt format with each line containing a brand name.
#
# The following code loads a brand list file, and reads all its brands into a list.
# !ls ../DATASET/BrandLists/
# +
brands = open('./../DATASET/BrandLists/Appario_Brand_List.txt', 'r')
# brands = open('./../DATASETS/BrandLists/CloudtailBrandListTop.txt', 'r')
brand_list = []
for b in brands:
# Removing (\n) from the end of each brand name read
b = b.strip(" \n")
b = b.strip("\n")
brand_list.append(b)
print('Brand List: ', brand_list[:10])
print('Brand Count: ', len(brand_list))
# -
# #### Step 2: Define Headers
#
# Each header is a unique user agent which will be used to request the data from the website to be scraped. We use multiple user agents to ensure that if our request is rejected, we can retry.
#
# To create more headers, simply copy any one of the old headers and replace the 'user-agent' string with a new 'user-agent' string, which can be found online. (Eg. https://developer.chrome.com/multidevice/user-agent)
headers = [
{
'dnt': '1',
'upgrade-insecure-requests': '1',
'user-agent': 'Mozilla/5.0 (Windows NT 10.0; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/42.0.2311.135 Safari/537.36 Edge/12.246',
'accept': 'text/html,application/xhtml+xml,application/xml;q=0.9,image/webp,image/apng,*/*;q=0.8,application/signed-exchange;v=b3;q=0.9',
'sec-fetch-site': 'same-origin',
'sec-fetch-mode': 'navigate',
'sec-fetch-user': '?1',
'sec-fetch-dest': 'document',
'referer': 'https://www.amazon.com/',
'accept-language': 'en-GB,en-US;q=0.9,en;q=0.8',
},
{
'dnt': '1',
'upgrade-insecure-requests': '1',
'user-agent': 'Mozilla/5.0 (X11; CrOS x86_64 8172.45.0) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/51.0.2704.64 Safari/537.36',
'accept': 'text/html,application/xhtml+xml,application/xml;q=0.9,image/webp,image/apng,*/*;q=0.8,application/signed-exchange;v=b3;q=0.9',
'sec-fetch-site': 'same-origin',
'sec-fetch-mode': 'navigate',
'sec-fetch-user': '?1',
'sec-fetch-dest': 'document',
'referer': 'https://www.amazon.com/',
'accept-language': 'en-GB,en-US;q=0.9,en;q=0.8',
},
{
'dnt': '1',
'upgrade-insecure-requests': '1',
'user-agent': 'Mozilla/5.0 (Macintosh; Intel Mac OS X 10_11_2) AppleWebKit/601.3.9 (KHTML, like Gecko) Version/9.0.2 Safari/601.3.9',
'accept': 'text/html,application/xhtml+xml,application/xml;q=0.9,image/webp,image/apng,*/*;q=0.8,application/signed-exchange;v=b3;q=0.9',
'sec-fetch-site': 'same-origin',
'sec-fetch-mode': 'navigate',
'sec-fetch-user': '?1',
'sec-fetch-dest': 'document',
'referer': 'https://www.amazon.com/',
'accept-language': 'en-GB,en-US;q=0.9,en;q=0.8',
},
{
'dnt': '1',
'upgrade-insecure-requests': '1',
'user-agent': 'Mozilla/5.0 (Windows NT 6.1; WOW64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/47.0.2526.111 Safari/537.36',
'accept': 'text/html,application/xhtml+xml,application/xml;q=0.9,image/webp,image/apng,*/*;q=0.8,application/signed-exchange;v=b3;q=0.9',
'sec-fetch-site': 'same-origin',
'sec-fetch-mode': 'navigate',
'sec-fetch-user': '?1',
'sec-fetch-dest': 'document',
'referer': 'https://www.amazon.com/',
'accept-language': 'en-GB,en-US;q=0.9,en;q=0.8',
},
{
'dnt': '1',
'upgrade-insecure-requests': '1',
'user-agent': 'Mozilla/5.0 (X11; Ubuntu; Linux x86_64; rv:15.0) Gecko/20100101 Firefox/15.0.1',
'accept': 'text/html,application/xhtml+xml,application/xml;q=0.9,image/webp,image/apng,*/*;q=0.8,application/signed-exchange;v=b3;q=0.9',
'sec-fetch-site': 'same-origin',
'sec-fetch-mode': 'navigate',
'sec-fetch-user': '?1',
'sec-fetch-dest': 'document',
'referer': 'https://www.amazon.com/',
'accept-language': 'en-GB,en-US;q=0.9,en;q=0.8',
},
{
'dnt': '1',
'upgrade-insecure-requests': '1',
'user-agent': 'Mozilla/5.0 (Macintosh; Intel Mac OS X 10_15_4) AppleWebKit/605.1.15 (KHTML, like Gecko) Version/13.1 Safari/605.1.15',
'accept': 'text/html,application/xhtml+xml,application/xml;q=0.9,image/webp,image/apng,*/*;q=0.8,application/signed-exchange;v=b3;q=0.9',
'sec-fetch-site': 'same-origin',
'sec-fetch-mode': 'navigate',
'sec-fetch-user': '?1',
'sec-fetch-dest': 'document',
'referer': 'https://www.amazon.com/',
'accept-language': 'en-GB,en-US;q=0.9,en;q=0.8',
},
{
'dnt': '1',
'upgrade-insecure-requests': '1',
'user-agent': 'Mozilla/5.0 (Windows NT 10.0; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/58.0.3029.110 Safari/537.36 Edge/16.16299',
'accept': 'text/html,application/xhtml+xml,application/xml;q=0.9,image/webp,image/apng,*/*;q=0.8,application/signed-exchange;v=b3;q=0.9',
'sec-fetch-site': 'same-origin',
'sec-fetch-mode': 'navigate',
'sec-fetch-user': '?1',
'sec-fetch-dest': 'document',
'referer': 'https://www.amazon.com/',
'accept-language': 'en-GB,en-US;q=0.9,en;q=0.8',
},
{
'dnt': '1',
'upgrade-insecure-requests': '1',
'user-agent': 'Mozilla/5.0 (Macintosh; Intel Mac OS X 10_15) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/78.0.3904.70 Safari/537.36',
'accept': 'text/html,application/xhtml+xml,application/xml;q=0.9,image/webp,image/apng,*/*;q=0.8,application/signed-exchange;v=b3;q=0.9',
'sec-fetch-site': 'same-origin',
'sec-fetch-mode': 'navigate',
'sec-fetch-user': '?1',
'sec-fetch-dest': 'document',
'referer': 'https://www.amazon.com/',
'accept-language': 'en-GB,en-US;q=0.9,en;q=0.8',
},
{
'dnt': '1',
'upgrade-insecure-requests': '1',
'user-agent': 'Mozilla/5.0 (Windows NT 10.0; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/78.0.3904.70 Safari/537.36',
'accept': 'text/html,application/xhtml+xml,application/xml;q=0.9,image/webp,image/apng,*/*;q=0.8,application/signed-exchange;v=b3;q=0.9',
'sec-fetch-site': 'same-origin',
'sec-fetch-mode': 'navigate',
'sec-fetch-user': '?1',
'sec-fetch-dest': 'document',
'referer': 'https://www.amazon.com/',
'accept-language': 'en-GB,en-US;q=0.9,en;q=0.8',
},
{
'dnt': '1',
'upgrade-insecure-requests': '1',
'user-agent': 'Mozilla/5.0 (Macintosh; Intel Mac OS X 10.15; rv:70.0) Gecko/20100101 Firefox/70.0',
'accept': 'text/html,application/xhtml+xml,application/xml;q=0.9,image/webp,image/apng,*/*;q=0.8,application/signed-exchange;v=b3;q=0.9',
'sec-fetch-site': 'same-origin',
'sec-fetch-mode': 'navigate',
'sec-fetch-user': '?1',
'sec-fetch-dest': 'document',
'referer': 'https://www.amazon.com/',
'accept-language': 'en-GB,en-US;q=0.9,en;q=0.8',
},
{
'dnt': '1',
'upgrade-insecure-requests': '1',
'user-agent': 'Mozilla/5.0 (Windows NT 10.0; Win64; x64; rv:70.0) Gecko/20100101 Firefox/70.0',
'accept': 'text/html,application/xhtml+xml,application/xml;q=0.9,image/webp,image/apng,*/*;q=0.8,application/signed-exchange;v=b3;q=0.9',
'sec-fetch-site': 'same-origin',
'sec-fetch-mode': 'navigate',
'sec-fetch-user': '?1',
'sec-fetch-dest': 'document',
'referer': 'https://www.amazon.com/',
'accept-language': 'en-GB,en-US;q=0.9,en;q=0.8',
},
{
'dnt': '1',
'upgrade-insecure-requests': '1',
'user-agent': 'Mozilla/5.0 (Windows NT 10.0; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/70.0.3538.77 Safari/537.36',
'accept': 'text/html,application/xhtml+xml,application/xml;q=0.9,image/webp,image/apng,*/*;q=0.8,application/signed-exchange;v=b3;q=0.9',
'sec-fetch-site': 'same-origin',
'sec-fetch-mode': 'navigate',
'sec-fetch-user': '?1',
'sec-fetch-dest': 'document',
'referer': 'https://www.amazon.com/',
'accept-language': 'en-GB,en-US;q=0.9,en;q=0.8',
},
{
'dnt': '1',
'upgrade-insecure-requests': '1',
'user-agent': 'Mozilla/5.0 (Macintosh; Intel Mac OS X 10_15_5) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/83.0.4103.97 Safari/537.36',
'accept': 'text/html,application/xhtml+xml,application/xml;q=0.9,image/webp,image/apng,*/*;q=0.8,application/signed-exchange;v=b3;q=0.9',
'sec-fetch-site': 'same-origin',
'sec-fetch-mode': 'navigate',
'sec-fetch-user': '?1',
'sec-fetch-dest': 'document',
'referer': 'https://www.amazon.com/',
'accept-language': 'en-GB,en-US;q=0.9,en;q=0.8',
},
{
'dnt': '1',
'upgrade-insecure-requests': '1',
'user-agent': 'Mozilla/5.0 (Windows NT 10.0; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/83.0.4103.97 Safari/537.36 OPR/68.0.3618.165',
'accept': 'text/html,application/xhtml+xml,application/xml;q=0.9,image/webp,image/apng,*/*;q=0.8,application/signed-exchange;v=b3;q=0.9',
'sec-fetch-site': 'same-origin',
'sec-fetch-mode': 'navigate',
'sec-fetch-user': '?1',
'sec-fetch-dest': 'document',
'referer': 'https://www.amazon.com/',
'accept-language': 'en-GB,en-US;q=0.9,en;q=0.8',
},
{
'dnt': '1',
'upgrade-insecure-requests': '1',
'user-agent': 'Mozilla/5.0 (Windows NT 10.0; Trident/7.0; rv:11.0) like Gecko',
'accept': 'text/html,application/xhtml+xml,application/xml;q=0.9,image/webp,image/apng,*/*;q=0.8,application/signed-exchange;v=b3;q=0.9',
'sec-fetch-site': 'same-origin',
'sec-fetch-mode': 'navigate',
'sec-fetch-user': '?1',
'sec-fetch-dest': 'document',
'referer': 'https://www.amazon.com/',
'accept-language': 'en-GB,en-US;q=0.9,en;q=0.8',
},
{
'dnt': '1',
'upgrade-insecure-requests': '1',
'user-agent': 'Mozilla/5.0 (Macintosh; Intel Mac OS X 10_15_5) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/83.0.4103.97 Safari/537.36 Edg/83.0.478.37',
'accept': 'text/html,application/xhtml+xml,application/xml;q=0.9,image/webp,image/apng,*/*;q=0.8,application/signed-exchange;v=b3;q=0.9',
'sec-fetch-site': 'same-origin',
'sec-fetch-mode': 'navigate',
'sec-fetch-user': '?1',
'sec-fetch-dest': 'document',
'referer': 'https://www.amazon.com/',
'accept-language': 'en-GB,en-US;q=0.9,en;q=0.8',
}
]
# #### Step 3: Read Extractor Files
#
# The extractor (.yml) files contain *css id* information about the fields which we intend to extract from the scarped website. Here, the two extractor files are:
# ##### 1. product_list.yml
# From the scraped webpage, this extractor file extracts the main *css division* which contains all the individual (child) products. Once the main div is scraped, it extracts all the child divisions (products) contained in it.
# ##### 2. nextpg.yml
# Extracts the 'next' button from the website, to check if its disabled. If it is disabled, it means that we have reached the end of the product list for the current brand. We then move onto the next brand to continue our scraping.
e = Extractor.from_yaml_file('./Extractor/product_list.yml')
l = Extractor.from_yaml_file('./Extractor/nextpg.yml')
# #### Step 4: Define scrape function
# **NOTE:** Set the variables MAX_TRIALS & ERROR_THRESHHOLD according to your preferences.
#
# A high MAX_TRIALS will slow down the scraping as it will scrape those pages without actually any data multiple times too, but it will reduce the chances of error.
# A low ERROR_THRESHHOLD will also slow down the scraping, as VPN will need to changed multiple times. However, it will reduce the chances missing data due to errors.
#
# The function scrape(url) downloads the webpage at the given url (here: product list pages) using requests module, and looks for products on the page. If it finds any product, it extracts the required fields and returns the data. If no product is found, it continues to randomly select a new header and retry scraping untill the limit MAX_TRIALS is reached, where it concludes that the page does not contain any data.
#
# These multiple trials are required, as amazon often blocks a user for repeqatedly making requests using the same user agent.
# +
MAX_TRIALS = 15 # Set the max number of trials to perform here.
ERROR_COUNT = 1 # Used for keeping a count of errors, if the count exceeds threshhold, the user is asked to
# change the vpn
ERROR_THRESHHOLD = 10 # Number of pages with missed information after which vpn change is required
def scrape(url):
global MAX_TRIALS
global ERROR_COUNT
global ERROR_THRESHHOLD
'''
This function downloads the webpage at the given url using requests module.
Parameters:
url (string): URL of webpage to scrape
Returns:
string: If the URL contains products, returns the html of the webpage as text, else returns 'False'.
'''
# Download the page using requests
print("Downloading %s"%url)
trial = 0
while(True):
if ERROR_COUNT % ERROR_THRESHHOLD == 0:
_ = input('Please Change VPN and press any key to continue')
ERROR_COUNT += 1
if trial == MAX_TRIALS:
print("Max trials exceeded yet no Data found on this page!")
ERROR_COUNT += 1
return 'False'
trial = trial + 1
print("Trial no:", trial)
# Get the html data from the url
while True:
try:
r = requests.get(url, headers=random.choice(headers), timeout = 15)
# We use product_list.yml extractor to extract the product details from the html data text
data = e.extract(r.text)
# If the products div in the scraped html is not empty, return html text.
#If the products div in the scraped html is empty, retry with new user agent.
if (data['products'] != None):
return r.text
else:
print("Retrying with new user agent!")
break
except requests.exceptions.RequestException as err:
print('Error Detected: ', err)
print('Retrying after 30 seconds')
sleep(30)
continue
except requests.exceptions.HTTPError as err:
print('Error Detected: ', err)
print('Retrying after 30 seconds')
sleep(30)
continue
except requests.exceptions.ConnectionError as err:
print('Error Detected: ', err)
print('Retrying after 30 seconds')
sleep(30)
continue
except requests.exceptions.Timeout as err:
print('Error Detected: ', err)
print('Retrying after 30 seconds')
sleep(30)
continue
# -
# #### Step 5: Initialise path of output file
#
# **NOTE:** Set the File Name accoring to what is being scraped here
#
# Eg: SCRAPED_PRODUCT_LIST_APPARIO or SCRAPED_PRODUCT_LIST_CLOUDTAIL
# +
FileName = input('Enter a Filename for output file!\n')
outfile_path = str('./ScriptOutput/DATASET/' + str(FileName) + '.jsonl')
# -
# #### Step 6: Enter Seller Name
#
# Enter Seller Name which the brand list is associated with.
# Eg: Cloudtail India or Appario Retail Pvt Ltd
seller = input('Enter Seller Name!\n')
# #### Step 7: Defining Functions to clean the data.
# +
def CleanRating(s):
'''
Here, the input is rating in a string format, eg: "3.3 out of 5 stars".
The function converts it to a float, eg: '3.3'
'''
if s is not None:
try:
return float(s.split(' ')[0])
except ValueError:
return None
except AttributeError:
return None
else:
return None
def CleanRatingCount(s):
'''
Here, the input is RatingCount in a string format, eg: "336 ratings".
The function converts it to a float, eg: '336'
'''
if s is not None:
return float(s.split(' ')[0].replace(',', ''))
else:
return float(0)
def CleanAmazonPrice(s):
'''
Here, the input is AmazonPrice in a string format, eg: "₹ 336.00".
The function converts it to a float, eg: '336'
'''
if s is not None:
print(s)
s = s.replace('₹', '').replace(',', '').replace(r'\x', '').replace('a', '')
return float(s.strip().split(' ')[0])
else:
return s
# -
# #### Step 8: Begin Main Scraping
#
# ##### NOTE: CHANGE THE URL BASED ON THE SELLER
#
# EG:
#
# Cloudtail URLs-> https://www.amazon.in/s?i=merchant-items&me=AT95IG9ONZD7S&rh=p_4%3AAmazon
#
# Appario URLS-> https://www.amazon.in/s?i=merchant-items&me=A14CZOWI0VEHLG&rh=p_4%3Amazon
#
# Note that using both these urls, we are searching for 'Amazon' brand products, but first url searches for Amazon brand products on Cloudtail Storefront, and second one on Appario's.
with open(outfile_path,'a') as outfile:
for b in brand_list:
pg_number = 1
while True:
# To account for differnt urls based on page number
if pg_number == 1:
url = str("https://www.amazon.in/s?i=merchant-items&me=A14CZOWI0VEHLG&rh=p_4%3A"+str(b))
else:
url = str("https://www.amazon.in/s?i=merchant-items&me=A14CZOWI0VEHLG&rh=p_4%3A"+str(b)+"&dc&page="+str(pg_number))
data_text = scrape(url)
# Case 1: Scraped page does not contain any products
if data_text == 'False':
pass
# Case 2: Scraped page contains products
else:
# Extract all product details in a dict 'data' using the extractor file
data = e.extract(data_text)
# Save html text to file
html_files_path = str('./ScriptOutput/HTML/'+ str(FileName) + '/' + str(b) +'/Page_'+str(pg_number)+'.html')
os.makedirs(os.path.dirname(html_files_path), exist_ok=True) # Create file to save our html data
with open(html_files_path, 'w') as file:
file.write(data_text)
# data['products'] is a dict which contains details of all products present on the scraped page
for product in data['products']:
product['Rating'] = CleanRating(product['Rating'])
product['RatingCount'] = CleanRatingCount(product['RatingCount'])
product['AmazonPrice'] = CleanAmazonPrice(product['AmazonPrice'])
product['SearchUrl'] = url
product['Brand'] = b
product['Seller'] = seller
date = datetime.datetime.now()
product['Timestamp'] = date.strftime("%c")
product['ProductPageUrl'] = str('https://www.amazon.in' + str(product['ProductPageUrl']))
print("Saving Product: %s"%product['Title'])
print(product)
json.dump(product,outfile)
outfile.write("\n")
# If next page is not available, break and go to next brand
if l.extract(data_text)['last'] == 'Next →':
break
elif data_text == 'False':
break
else:
pg_number += 1 # Incrementing page numbe
# #### Step 9: Read .jsonl File
# +
# ProductListFile = open('./ScriptOutput/DATASET/test.jsonl', 'r')
ProductListFile = open(outfile_path)
ProductList = []
reader = jsonlines.Reader(ProductListFile)
for item in reader.iter():
ProductList.append(item)
df = pd.DataFrame(ProductList)
print(df.count())
df.head()
# -
| SCRIPTS/ScrapeProductListBySeller.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sos
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SoS
# language: sos
# name: sos
# ---
# + [markdown] kernel="SoS"
# # How to avoid or force the re-execution of executed steps
# + [markdown] kernel="SoS"
# * **Difficulty level**: easy
# * **Time need to lean**: 10 minutes or less
# * **Key points**:
# * Runtime signatures avoids repeated execution of steps
# * Option `-s` controls the behavior of signatures
#
# + [markdown] kernel="SoS"
# ## Runtime signature
# + [markdown] kernel="SoS"
# One of the most annonying problems with the development and execution of workflows is that it can take very long times to execute then. What makes things worse is that we frequently need to re-run the workflow with different paremeters and even different tools -- it can be really time-consuming to re-execute the whole workflow repeatedly, but it is also very error-prone to repeat selected steps of a workflow.
#
# SoS addresses this problem by using <font color='red'>runtime signatures</font> to keep track of <font color='red'>execution units</font>, namely the input, output, and dependent targets, and related SoS variables of a piece of workflow. SoS tracks execution of statements at the step level for each [input group](../documentation/SoS_Step.html) and saves runtime signatures at a folder called `.sos` under the project directory. The runtime signatures are used to
#
# 1. Avoid repeated execution of identical units, and
# 2. Keep track of workflow related files for project management
#
# This tutorial focuses on the first usage. The second one would be described in detail in [Project Management](Project_Management.html).
# + [markdown] kernel="SoS"
# ### `ignore` mode
# + [markdown] kernel="SoS"
# SoS workflows can be executed in batch mode and in interactive mode using the SoS kernel in Jupyter notebook or qtconsole. Because the SoS kernel is mostly used to execute short statements in SOS and other kernels, runtime signatures are by default set to `ignore` in interactive mode (and to `default` in batch mode).
# + [markdown] kernel="SoS"
# Let us create a temporary directory and execute a workflow that take a bit of time to execute. This is done in the default `ignore` signature mode of the Jupyter notebook
# + kernel="SoS"
# %sandbox --dir tmp
!rm -rf .sos/.runtime
![ -d temp ] || mkdir temp
# + kernel="SoS"
# %sandbox --dir tmp
# %run
parameter: size=1000
[10]
output: "temp/result.txt"
sh: expand=True
dd if=/dev/urandom of={_output} count={size}
[20]
output: 'temp/size.txt'
with open(_output[0], 'w') as sz:
sz.write(f"{_input}: {os.path.getsize(_input[0])}\n")
# + [markdown] kernel="SoS"
# Now, if we re-run the last script, nothing changes and it takes a bit of time to execute the script.
# + kernel="SoS"
# %sandbox --dir tmp
# %rerun
# + [markdown] kernel="SoS"
# ### `default` mode
# + [markdown] kernel="SoS"
# Now let us switch to `default` mode of signature by running the script with option `-s default`. When you run the script for the first time, it would execute normally and save runtime signature of the steps.
# + kernel="SoS"
# %sandbox --dir tmp
# %rerun -s default
# + [markdown] kernel="SoS"
# but both steps would be ignored. Here we use `-v2` to show the `ignored` message. This time we use magic `%set` to make option `-s default` persistent so that we do not have to specify it each time.
# + kernel="SoS"
# %sandbox --dir tmp
# %set -s default
# %rerun -v2
# + [markdown] kernel="SoS"
# However, if you use a different parameter (not the default `size=1000`), the steps would be rerun
# + kernel="SoS"
# %sandbox --dir tmp
# %rerun -v2 --size 2000
# + [markdown] kernel="SoS"
# The signature is at the step level so if you change the second step of the script, the first step would still be skipped. Note that the step is independent of the script executed so a step would be skipped even if its signature was saved by the execution of another workflow. The signature is clever enough to allow minor changes such as addition of spaces and comments.
# + kernel="SoS"
# %sandbox --dir tmp
# %run --size 2000 -v2
parameter: size=1000
[10]
output: "temp/result.txt"
# added comment
sh: expand=True
dd if=/dev/urandom of={_output} count={size}
[20]
output: 'temp/size.txt'
with open(_output[0], 'w') as sz:
sz.write(f"Modified {_input}: {os.path.getsize(_input[0])}\n")
# + [markdown] kernel="SoS"
# ### `assert` mode
# + [markdown] kernel="SoS"
# The `assert` mode is used to detect if anything has been changed after the execution of a workflow. For example,
# + kernel="SoS"
# %sandbox --dir tmp
# %set -s assert
# %rerun --size 2000 -v2
# + [markdown] kernel="SoS"
# Now if you change one of the output files, sos would fail with an error message.
# + kernel="sos"
# %sandbox --expect-error --dir tmp
!echo "aaa" >> temp/result.txt
# %rerun --size 2000 -v2
# + [markdown] kernel="SoS"
# ### `force` mode
# + [markdown] kernel="SoS"
# The `force` signature mode ignores existing signatures and re-run the workflow. This is needed when you would like to forcefully re-run all the steps to generate another set of output if outcome of some steps is random, or to re-run the workflow because of changes that is not tracked by SoS, for example after you have installed a new version of a program.
# + kernel="SoS"
# %sandbox --dir tmp
# %set
# %rerun --size 2000 -s force
# + [markdown] kernel="SoS"
# ### `build` mode
# + [markdown] kernel="SoS"
# The `build` mode is somewhat opposite to the `force` mode in that it creates (or overwrite existing signature if exists) with existing output files. It is useful, for example, if you are adding a step to a workflow that you have tested outside of SoS (without signature) but do not want to rerun it, or if for some reason you have lost your signature files and would like to reconstruct them from existing outputs.
# + kernel="SoS"
# %sandbox --dir tmp
# %rerun --size 2000 -s build -v2
# + [markdown] kernel="SoS"
# This mode can introduce erraneous files to the signatures because it does not check the validity of the incorporated files. For example, SoS would not complain if you change parameter and replace `temp/result.txt` with something else.
# + kernel="SoS"
# %sandbox --dir tmp
!echo "something else" > temp/result.txt
# %rerun -s build -v2
# + kernel="SoS"
# cleanup
!rm -rf tmp
# + [markdown] kernel="SoS"
# ## Further reading
#
# *
| src/user_guide/signature.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # treant-js
#
# [treant-js](http://fperucic.github.io/treant-js/) is a javascript library to plot diagram and trees. The goal is to wrap it as a library for python.
# [documentation](http://fperucic.github.io/treant-js/) [source](https://github.com/fperucic/treant-js) [tutorial](http://help.plot.ly/tutorials/) [gallery](http://fperucic.github.io/treant-js/#orgchart-examples)
from jyquickhelper import add_notebook_menu
add_notebook_menu()
# ## Javacript in the notebook
#
# Let's take one of the example and look at the source: [tennis draw](http://fperucic.github.io/treant-js/examples/tennis-draw/). I took the source of the example and I changed relative to absolute paths. There are two parts:
#
# * HTML: defines styles, references external script, new HTML and section div
# * Javascript: defines a tree structure as Json and calls *treant-js* to convert it into SVG.
# + language="html"
#
# <style>
# .chart { height: 600px; width: 900px; margin: 5px; margin: 15px auto; border: 3px solid #DDD; border-radius: 3px; }
#
# .tennis-draw {
# font-size: 10px;
# width: 100px;
# }
#
# .tennis-draw.winner { height: 38px; }
# .tennis-draw.winner:hover { background: url('http://fperucic.github.io/treant-js/examples/tennis-draw/trophy.png') right 0 no-repeat; }
# .tennis-draw.winner .node-name { padding-left: 10px; margin-top: 1px; display: block; }
#
# .tennis-draw .node-name { padding: 2px; white-space: pre; color: #00AFF0; }
# .tennis-draw .node-desc { padding: 2px; color: #999; }
#
# .tennis-draw.first-draw .node-title,
# .tennis-draw.first-draw .node-name,
# .tennis-draw.first-draw img { position: absolute; top: -8px; }
# .tennis-draw.first-draw:hover img { width: 20px; top: -12px; }
#
# .tennis-draw.first-draw { width: 165px; height: 20px; }
# .tennis-draw.first-draw img { margin: 3px 4px 0 0; left: 25px; }
# .tennis-draw.first-draw .node-title { margin-top: 3px; }
# .tennis-draw.first-draw .node-name { width: 113px; padding-left: 50px; }
# .tennis-draw.first-draw.bye .node-name { color: #999; }
# </style>
#
# <link rel="stylesheet" href="http://fperucic.github.io/treant-js/Treant.css">
# <script src="http://fperucic.github.io/treant-js/vendor/raphael.js"></script>
# <script src="http://fperucic.github.io/treant-js/Treant.js"></script>
# <div class="chart" id="OrganiseChart6"></div>
# + language="javascript"
#
# // source:
# // http://www.atpworldtour.com/Share/Event-Draws.aspx?EventId=410&Year=2013
#
# var tree_structure = {
# chart: {
# container: "#OrganiseChart6",
# levelSeparation: 20,
# siblingSeparation: 15,
# subTeeSeparation: 15,
# rootOrientation: "EAST",
#
# node: {
# HTMLclass: "tennis-draw",
# drawLineThrough: true
# },
# connectors: {
# type: "straight",
# style: {
# "stroke-width": 2,
# "stroke": "#ccc"
# }
# }
# },
#
# nodeStructure: {
# text: {
# name: {val: "<NAME>",
# href: "http://www.atpworldtour.com/Tennis/Players/Top-Players/Novak-Djokovic.aspx"}
# },
# HTMLclass: "winner",
# children: [
# {
# text: {
# name: "<NAME>",
# desc: "4-6, 6-2, 6-2"
# },
# children: [
# {
# text: {
# name: "<NAME>",
# desc: "4-6, 6-1, 6-4"
# },
# children: [
# {
# text: {
# name: "<NAME>",
# desc: "4-6, 6-1, 6-4"
# },
# children: [
# {
# text: {
# name: "<NAME>",
# title: 1
# },
# image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/srb.jpg",
# HTMLclass: "first-draw",
# },
# {
# text: {
# name: "Bye",
# title: 2
# },
# HTMLclass: "first-draw bye"
# }
# ]
# },
# {
# text: {
# name: "<NAME>",
# desc: "6-4, 6-0"
# },
# children: [
# {
# text: {
# name: "<NAME>",
# title: 3
# },
# image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/rus.jpg",
# HTMLclass: "first-draw"
# },
# {
# text: {
# name: "<NAME>",
# title: 4
# },
# image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/esp.jpg",
# HTMLclass: "first-draw"
# }
# ]
# }
# ]
# },
# {
# text: {
# name: "<NAME>",
# desc: "6-0, 3-6, 6-3"
# },
# children: [
# {
# text: {
# name: "<NAME>",
# desc: "4-6, 6-2, 6-3"
# },
# children: [
# {
# text: {
# name: "<NAME>",
# title: 5
# },
# image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/lat.jpg",
# HTMLclass: "first-draw"
# },
# {
# text: {
# name: "<NAME>",
# title: 6
# },
# image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/usa.jpg",
# HTMLclass: "first-draw"
# }
# ]
# },
# {
# text: {
# name: "<NAME>",
# desc: "6-4, 6-0"
# },
# children: [
# {
# text: {
# name: "<NAME>",
# title: 7
# },
# image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/slo.jpg",
# HTMLclass: "first-draw"
# },
# {
# text: {
# name: "<NAME>",
# title: 8
# },
# image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/arg.jpg",
# HTMLclass: "first-draw"
# }
# ]
# }
# ]
# }
# ]
# },
# {
# text: {
# name: "<NAME>",
# desc: "6-3, 1-6, 7-6(3)"
# },
# children: [
# {
# text: {
# name: "<NAME>",
# desc: "4-6, 6-1, 6-4"
# },
# children: [
# {
# text: {
# name: "<NAME>",
# desc: "6-1, 6-4"
# },
# children: [
# {
# text: {
# name: "<NAME>",
# title: 9
# },
# image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/can.jpg",
# HTMLclass: "first-draw"
# },
# {
# text: {
# name: "<NAME>",
# title: 10
# },
# image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/fra.jpg",
# HTMLclass: "first-draw"
# }
# ]
# },
# {
# text: {
# name: "<NAME>",
# desc: "6-1, 6-2"
# },
# children: [
# {
# text: {
# name: "<NAME>",
# title: 11
# },
# image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/fin.jpg",
# HTMLclass: "first-draw"
# },
# {
# text: {
# name: "<NAME>",
# title: 12
# },
# image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/srb.jpg",
# HTMLclass: "first-draw"
# }
# ]
# }
# ]
# },
# {
# text: {
# name: "<NAME>, <NAME>",
# desc: "6-2, 6-4"
# },
# children: [
# {
# text: {
# name: "<NAME>",
# desc: "4-6, 6-2, 6-3"
# },
# children: [
# {
# text: {
# name: "<NAME>",
# title: 13
# },
# image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/ukr.jpg",
# HTMLclass: "first-draw"
# },
# {
# text: {
# name: "<NAME>",
# title: 14
# },
# image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/aus.jpg",
# HTMLclass: "first-draw"
# }
# ]
# },
# {
# text: {
# name: "<NAME>, <NAME>",
# desc: "6-4, 6-0"
# },
# children: [
# {
# text: {
# name: "Bye",
# title: 15
# },
# HTMLclass: "first-draw bye"
# },
# {
# text: {
# name: "<NAME>, <NAME>",
# title: 16
# },
# image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/arg.jpg",
# HTMLclass: "first-draw"
# }
# ]
# }
# ]
# }
# ]
# }
# ]
# }
# };
#
# new Treant( tree_structure );
# -
# ## Wrap treant-js into a Python library
#
# We would to produce a function with the following signature:
def display_treant(json, css):
pass
# First issue: the javascript is hosted on a website which means the notebook will not work offline unless we import the javascript at a specific location. See [Jupyter notebook extensions](https://github.com/ipython-contrib/IPython-notebook-extensions) to see where to place them.
# +
css = """
.chart { height: 600px; width: 900px; margin: 5px; margin: 15px auto; border: 3px solid #DDD; border-radius: 3px; }
.tennis-draw {
font-size: 10px;
width: 100px;
}
.tennis-draw.winner { height: 38px; }
.tennis-draw.winner:hover {
background: url('http://fperucic.github.io/treant-js/examples/tennis-draw/trophy.png') right 0 no-repeat;
}
.tennis-draw.winner .node-name { padding-left: 10px; margin-top: 1px; display: block; }
.tennis-draw .node-name { padding: 2px; white-space: pre; color: #00AFF0; }
.tennis-draw .node-desc { padding: 2px; color: #999; }
.tennis-draw.first-draw .node-title,
.tennis-draw.first-draw .node-name,
.tennis-draw.first-draw img { position: absolute; top: -8px; }
.tennis-draw.first-draw:hover img { width: 20px; top: -12px; }
.tennis-draw.first-draw { width: 165px; height: 20px; }
.tennis-draw.first-draw img { margin: 3px 4px 0 0; left: 25px; }
.tennis-draw.first-draw .node-title { margin-top: 3px; }
.tennis-draw.first-draw .node-name { width: 113px; padding-left: 50px; }
.tennis-draw.first-draw.bye .node-name { color: #999; }
"""
classname = "chart"
# this part should be part of a nice API (to avoid trick like this __DIVID__)
json_tree = """{
container: "#__DIVID__",
levelSeparation: 20,
siblingSeparation: 15,
subTeeSeparation: 15,
rootOrientation: "EAST",
node: {
HTMLclass: "tennis-draw",
drawLineThrough: true
},
connectors: {
type: "straight",
style: {
"stroke-width": 2,
"stroke": "#ccc"
}
}
}"""
# there should a nice API to define that
json_data = """{
text: {
name: {val: "<NAME>",
href: "http://www.atpworldtour.com/Tennis/Players/Top-Players/Novak-Djokovic.aspx"}
},
HTMLclass: "winner",
children: [
{
text: {
name: "<NAME>",
desc: "4-6, 6-2, 6-2"
},
children: [
{
text: {
name: "<NAME>",
desc: "4-6, 6-1, 6-4"
},
children: [
{
text: {
name: "<NAME>",
desc: "4-6, 6-1, 6-4"
},
children: [
{
text: {
name: "<NAME>",
title: 1
},
image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/srb.jpg",
HTMLclass: "first-draw",
},
{
text: {
name: "Bye",
title: 2
},
HTMLclass: "first-draw bye"
}
]
},
{
text: {
name: "<NAME>",
desc: "6-4, 6-0"
},
children: [
{
text: {
name: "<NAME>",
title: 3
},
image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/rus.jpg",
HTMLclass: "first-draw"
},
{
text: {
name: "<NAME>",
title: 4
},
image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/esp.jpg",
HTMLclass: "first-draw"
}
]
}
]
},
{
text: {
name: "<NAME>",
desc: "6-0, 3-6, 6-3"
},
children: [
{
text: {
name: "<NAME>",
desc: "4-6, 6-2, 6-3"
},
children: [
{
text: {
name: "<NAME>",
title: 5
},
image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/lat.jpg",
HTMLclass: "first-draw"
},
{
text: {
name: "<NAME>",
title: 6
},
image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/usa.jpg",
HTMLclass: "first-draw"
}
]
},
{
text: {
name: "<NAME>",
desc: "6-4, 6-0"
},
children: [
{
text: {
name: "<NAME>",
title: 7
},
image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/slo.jpg",
HTMLclass: "first-draw"
},
{
text: {
name: "<NAME>",
title: 8
},
image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/arg.jpg",
HTMLclass: "first-draw"
}
]
}
]
}
]
},
{
text: {
name: "<NAME>",
desc: "6-3, 1-6, 7-6(3)"
},
children: [
{
text: {
name: "<NAME>",
desc: "4-6, 6-1, 6-4"
},
children: [
{
text: {
name: "<NAME>",
desc: "6-1, 6-4"
},
children: [
{
text: {
name: "<NAME>",
title: 9
},
image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/can.jpg",
HTMLclass: "first-draw"
},
{
text: {
name: "<NAME>",
title: 10
},
image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/fra.jpg",
HTMLclass: "first-draw"
}
]
},
{
text: {
name: "<NAME>",
desc: "6-1, 6-2"
},
children: [
{
text: {
name: "<NAME>",
title: 11
},
image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/fin.jpg",
HTMLclass: "first-draw"
},
{
text: {
name: "<NAME>",
title: 12
},
image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/srb.jpg",
HTMLclass: "first-draw"
}
]
}
]
},
{
text: {
name: "<NAME>, <NAME>",
desc: "6-2, 6-4"
},
children: [
{
text: {
name: "<NAME>",
desc: "4-6, 6-2, 6-3"
},
children: [
{
text: {
name: "<NAME>",
title: 13
},
image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/ukr.jpg",
HTMLclass: "first-draw"
},
{
text: {
name: "<NAME>",
title: 14
},
image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/aus.jpg",
HTMLclass: "first-draw"
}
]
},
{
text: {
name: "<NAME>, <NAME>",
desc: "6-4, 6-0"
},
children: [
{
text: {
name: "Bye",
title: 15
},
HTMLclass: "first-draw bye"
},
{
text: {
name: "<NAME>, <NAME>",
title: 16
},
image: "http://fperucic.github.io/treant-js/examples/tennis-draw/flags/arg.jpg",
HTMLclass: "first-draw"
}
]
}
]
}
]
}
]
}"""
# -
from jupytalk.talk_examples.treant_wrapper import display_treant
from IPython.core.display import display
display(display_treant(json_tree, json_data, css, classname))
# ## What's next?
#
# * Add local support
# * Make the graph stick when the notebook is display again (reload dependencies)
# * Change parameter ``json_tree`` for a dictionary with options
# * Create an API to create the data in Python and not in Json.
| _doc/notebooks/2016/pydata/jsonly_treant.ipynb |